Alpha-amylase Combinatorial Variants
Cascao-Pereira; Luis G. ; et al.
U.S. patent application number 16/292057 was filed with the patent office on 2020-03-19 for alpha-amylase combinatorial variants. The applicant listed for this patent is DANISCO US INC. Invention is credited to Pieter Augustinus, Richard R Bott, Luis G. Cascao-Pereira, Dina Finan, Roel Hermant, Marc Kolkman, Monica Ocha Ruiz, Dewy Van Tol, David Edward Wildes.
| Application Number | 20200087644 16/292057 |
| Document ID | / |
| Family ID | 50680117 |
| Filed Date | 2020-03-19 |











View All Diagrams
| United States Patent Application | 20200087644 |
| Kind Code | A1 |
| Cascao-Pereira; Luis G. ; et al. | March 19, 2020 |
ALPHA-AMYLASE COMBINATORIAL VARIANTS
Abstract
Disclosed are compositions and methods relating to variant alpha-amylases. The variant alpha-amylases are useful, for example, for starch liquefaction and saccharification, for cleaning starchy stains in laundry, dishwashing, and other applications, for textile processing (e.g., desizing), in animal feed for improving digestibility, and for baking and brewing.
| Inventors: | Cascao-Pereira; Luis G.; (Redwood City, CA) ; Finan; Dina; (Palo Alto, CA) ; Wildes; David Edward; (Palo Alto, CA) ; Augustinus; Pieter; (Palo Alto, CA) ; Hermant; Roel; (Palo Alto, CA) ; Ruiz; Monica Ocha; (Palo Alto, CA) ; Van Tol; Dewy; (Palo Alto, CA) ; Bott; Richard R; (Palo Alto, CA) ; Kolkman; Marc; (Palo Alto, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 50680117 | ||||||||||
| Appl. No.: | 16/292057 | ||||||||||
| Filed: | March 4, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14775595 | Sep 11, 2015 | |||
| PCT/US14/23458 | Mar 11, 2014 | |||
| 16292057 | ||||
| 61776699 | Mar 11, 2013 | |||
| 61906617 | Nov 20, 2013 | |||
| 61907131 | Nov 21, 2013 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 9/2414 20130101; C12P 19/14 20130101; A23L 2/382 20130101; C12N 9/2417 20130101; C11D 3/386 20130101; C11D 3/38681 20130101; A23V 2002/00 20130101; C12P 19/02 20130101 |
| International Class: | C12N 9/26 20060101 C12N009/26; C12N 9/28 20060101 C12N009/28; C11D 3/386 20060101 C11D003/386; A23L 2/38 20060101 A23L002/38; C12P 19/02 20060101 C12P019/02; C12P 19/14 20060101 C12P019/14 |
Claims
1. A recombinant variant of a parent .alpha.-amylase comprising: a mutation at an amino acid residue corresponding to E187 or S241; and at least one mutation at an amino acid residue corresponding to an amino acid residue selected from the group consisting of N126, Y150, F153, L171, T180, and, I203; wherein the variant .alpha.-amylase or the parent .alpha.-amylase has at least 60% amino acid sequence identity relative to SEQ ID NO: 1, which is used for numbering; and wherein the variant has increased thermostability, detergent stability, starch liquifaction activity, and/or cleaning performance compared to the parent .alpha.-amylase or a reference .alpha.-amylase differing from the variant .alpha.-amylase only by the absence of the mutations.
2. The variant .alpha.-amylase of claim 1, comprising at least two mutations at amino acid residues corresponding N126, Y150, F153, L171, and, I203, using SEQ ID NO: 1 for numbering.
3. The variant .alpha.-amylase of claim 1, further comprising a deletion of at least one amino acid residue corresponding to R178, G179, T180, and G181, using SEQ ID NO: 1 for numbering.
4. The variant .alpha.-amylase of claim 1, further comprising deletions of amino acid residues corresponding to R178 and G179, or T180 and G181.
5. The variant .alpha.-amylase of claim 1, further comprising a mutation at an amino acid residue corresponding to G476 and/or G477, using SEQ ID NO: 1 for numbering.
6. The variant .alpha.-amylase of claim 1, further comprising a mutation in an amino acid residue corresponding to an amino acid residue selected from the group consisting of E132, Q167, T180, and A277, using SEQ ID NO: 1 for numbering.
7. The variant .alpha.-amylase of claim 1, further comprising a mutation in an amino acid residue corresponding to an amino acid residue selected from the group consisting of R458, T459, and D460, using SEQ ID NO: 1 for numbering.
8. The variant .alpha.-amylase of claim 1, further comprising a mutation in an amino acid residue corresponding to T180, using SEQ ID NO: 1 for numbering.
9. The variant .alpha.-amylase of claim 1, further comprising a mutation in an amino acid residue corresponding to N205, using SEQ ID NO: 3 for numbering.
10. The variant .alpha.-amylase of claim 1, further comprising a mutation in an amino acid residue corresponding to an amino acid residue selected from the group consisting of T333G, A335S, and Q337E, using SEQ ID NO: 3 for numbering.
11. The variant .alpha.-amylase of claim 1, further comprising a mutation in an amino acid residue corresponding to an amino acid residue position selected from the group consisting of 6, 7, 8, 11, 14, 15, 20, 21, 23, 26, 27, 28, 37, 38, 39, 40, 42, 45, 46, 48, 49, 50, 51, 52, 53, 54, 58, 61, 62, 68, 70, 71, 72, 73, 79, 80, 81, 82, 84, 85, 87, 88, 89, 92, 93, 94, 95, 96, 97, 98, 101, 108, 111, 112, 113, 114, 115, 116, 117, 118, 120, 122, 123, 124, 126, 127, 129, 130, 131, 132, 133, 134, 136, 137, 138, 140, 142, 143, 144, 147, 148, 149, 150, 151, 152, 153, 154, 155, 156, 158, 159, 165, 167, 168, 170, 171, 172, 175, 176, 177, 180, 181, 182, 187, 190, 191, 193, 199, 200, 201, 203, 206, 208, 210, 211, 212, 214, 215, 216, 219, 221, 223, 225, 226, 227, 235, 238, 239, 240, 241, 242, 243, 245, 246, 247, 248, 249, 250, 252, 253, 254, 256, 257, 258, 260, 261, 262, 266, 267, 268, 269, 270, 271, 273, 276, 277, 279, 280, 282, 284, 285, 286, 288, 296, 299, 300, 301, 302, 303, 304, 307, 308, 310, 311, 312, 313, 316, 317, 318, 320, 321, 325, 327, 335, 338, 342, 348, 349, 352, 356, 357, 360, 362, 363, 368, 369, 377, 381, 382, 383, 384, 385, 388, 390, 392, 394, 395, 396, 397, 398, 400, 401, 402, 403, 404, 405, 407, 408, 410, 414, 415, 416, 418, 419, 420, 421, 422, 423, 424, 426, 428, 429, 430, 431, 434, 435, 436, 439, 441, 442, 444, 445, 446, 447, 448, 449, 450, 451, 454, 455, 457, 460, 461, 462, 463, 464, 465, 466, 467, 469, 470, 471, 473, 474, 475, 476, 477, 479, 480, 481, 482, 483, and 484, using SEQ ID NO: 1 for numbering.
12. The variant .alpha.-amylase of claim 1, comprising a combinations of mutations corresponding to mutations selected from the group consisting of: E187P+I203Y+G476K, E187P+I203Y+G476K+R458N+T459S+D460T, T180D+E187P+I203Y+G476K, N126Y+T180D+E187P+I203Y+G476K, N126Y+T180D+E187P+I203Y+Y303D+G476T+G477E, N126Y+T180D+E187P+I203Y+Y303D+N475E+G477Q, N126Y+T180D+E187P+I203Y+Y303R+N475E+G476T+G477R, T038N+N088H+N126Y+T129I+N134M+F153W+L171R+T180D+E187P+I203Y+G476K+G477E, N126Y+E132H+T180D+E187P+I203Y+Y303D+G476T+G477E, N126Y+E187P+G476K, N126Y+F153W+E187P+G476K, N126Y+F153W+E187P+G4726+G477R, N126Y+E187P+I203Y, N126Y+I203Y+S241Q, N126Y+T180H+E187P+I203Y, N126Y+T180H+I203Y+S241Q, N126Y+F153W+T180H+E187P+I203Y, N126Y+F153W+T180H+I203Y+S241Q, N126Y+Y150H+F153W+L171N+E187P+I203Y, N126Y+Y150H+F153W+L171N+I203Y+S241Q, N126Y+Y150H+F153W+L171N+T180H+E187P+I203Y, N126Y+Y150H+F153W+L171N+T180H+I203Y+S241Q, and N126Y+F153W+T180D+I203Y+S241Q; wherein the variant has increased thermostability, detergent stability, stability starch liquifaction activity, or cleaning performance compared to the parent; wherein the variant or the parent has at least 60% amino acid sequence identity relative to SEQ ID NO: 1, which is used for numbering.
13. The variant amylase of claim 1, comprising the combinations of mutations corresponding to N126Y+F153W+T180D+I203Y+S241Q and one or more mutations corresponding to mutations selected from the group consisting of E132H, Q167E, A277F, and T400K.
14. The variant amylase of claim 13, comprising the combinations of mutations corresponding to mutations selected from the group consisting of: N126Y+E132H+F153W+T180D+I203Y+S241Q+A277F, N126Y+E132H+F153W+Q167E+T180D+I203Y+S241Q+A277F, and N126Y+E132H+F153W+Q167E+T180D+I203Y+S241Q+A277F+T400K.
15. The variant amylase of claim 1, wherein the parental .alpha.-amylase is from a Cytophaga species.
16. The variant amylase of claim 1, wherein the parental .alpha.-amylase is from a Paenibacillus species.
17. The variant amylase of claim 1, wherein the parental .alpha.-amylase is not from a Bacillus species.
18. The variant amylase of claim 1, wherein the parental .alpha.-amylase or the variant .alpha.-amylase has at least 70% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5.
19. The variant amylase of claim 1, wherein the parental .alpha.-amylase or the variant .alpha.-amylase has at least 70% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1 or SEQ ID NO: 3.
20. The variant amylase of claim 1, wherein the parental .alpha.-amylase or the variant .alpha.-amylase has at least 80% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5.
21. The variant amylase of claim 1, wherein the parental .alpha.-amylase or the variant .alpha.-amylase has at least 90% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5.
22. A composition comprising the variant .alpha.-amylase of any of the preceding claims.
23. The composition of claim 22, wherein the composition is effective for removing starchy stains from laundry, dishes, or textiles.
24. The composition of claim 22, further comprising a surfactant.
25. The composition of claim 22, wherein the composition is a detergent composition.
26. The composition of claim 22, wherein the composition is a laundry detergent or a laundry detergent additive.
27. The composition of claim 22, wherein the composition is a manual or automatic dishwashing detergent.
28. The composition of claim 22, further comprising one or more additional enzymes selected from the group consisting of protease, hemicellulase, cellulase, peroxidase, lipolytic enzyme, metallolipolytic enzyme, xylanase, lipase, phospholipase, esterase, perhydrolase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, .beta.-glucanase, arabinosidase, hyaluronidase, chondroitinase, laccase, metalloproteinase, amadoriase, glucoamylase, arabinofuranosidase, phytase, isomerase, transferase, and an amylase other than the amylase of claim 1.
29. The composition of claim 21, wherein the composition is for liquifying starch.
30. The composition of claim 21, wherein the composition is for saccharifying a composition comprising starch, for SSF post liquefaction, or for direct SSF without prior liquefaction.
31. The composition of claim 21, wherein the composition is for producing a fermented beverage.
32. The composition of claim 21, wherein the composition is for producing a baked food product.
33. The composition of claim 21, wherein the composition is for textile desizing.
34-56. (canceled)
Description
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This application claims benefit of priority from US provisional applications U.S. Ser. No. 61/776,699, filed 11 Mar. 2013, U.S. Ser. No. 61/906,617, filed 20 Nov. 2013, and U.S. Ser. No. 61/907,131, filed 21 Nov. 2013, and are incorporated herein by reference in their entirety.
FIELD OF THE INVENTION
[0002] Disclosed are compositions and methods relating to variant .alpha.-amylases containing a plurality of combinable mutations. The variant .alpha.-amylases are useful, for example, for starch liquefaction and saccharification, cleaning starchy stains, textile desizing, baking, and brewing.
BACKGROUND
[0003] Starch consists of a mixture of amylose (15-30% w/w) and amylopectin (70-85% w/w). Amylose consists of linear chains of .alpha.-1,4-linked glucose units having a molecular weight (MW) from about 60,000 to about 800,000. Amylopectin is a branched polymer containing .alpha.-1,6 branch points every 24-30 glucose units; its MW may be as high as 100 million.
[0004] Sugars from starch, in the form of concentrated dextrose syrups, are currently produced by an enzyme catalyzed process involving: (1) liquefaction (or viscosity reduction) of solid starch with an .alpha.-amylase into dextrins having an average degree of polymerization of about 7-10, and (2) saccharification of the resulting liquefied starch (i.e. starch hydrolysate) with amyloglucosidase (also called glucoamylase or GA). The resulting syrup has a high glucose content. Much of the glucose syrup that is commercially produced is subsequently enzymatically isomerized to a dextrose/fructose mixture known as isosyrup. The resulting syrup also may be fermented with microorganisms, such as yeast, to produce commercial products including ethanol, citric acid, lactic acid, succinic acid, itaconic acid, monosodium glutamate, gluconates, lysine, other organic acids, other amino acids, and other biochemicals, for example. Fermentation and saccharification can be conducted simultaneously (i.e., an SSF process) to achieve greater economy and efficiency.
[0005] .alpha.-amylases hydrolyze starch, glycogen, and related polysaccharides by cleaving internal .alpha.-1,4-glucosidic bonds at random. .alpha.-amylases, particularly from Bacilli, have been used for a variety of different purposes, including starch liquefaction and saccharification, textile desizing, starch modification in the paper and pulp industry, brewing, baking, production of syrups for the food industry, production of feedstocks for fermentation processes, and in animal feed to increase digestability. These enzymes can also be used to remove starchy soils and stains during dishwashing and laundry washing.
[0006] Numerous publications have described mutations in .alpha.-amylases. However, not all mutations produce the same effect in different molecules and not all mutation can be combined. In addition, many mutations produce molecules that have certain desirable qualities at the expense of other properties. The need exists for robust engineered .alpha.-amylases molecules.
SUMMARY
[0007] The present compositions and methods relate to variant amylase polypeptides, and methods of use, thereof. Aspects and embodiments of the present compositions and methods are summarized in the following separately-numbered paragraphs:
[0008] 1. In one aspect, a recombinant variant of a parent .alpha.-amylase is provided, comprising: a mutation at an amino acid residue corresponding to E187 or S241; and at least one mutation at an amino acid residue corresponding to an amino acid residue selected from the group consisting of N126, Y150, F153, L171, T180, and, I203; wherein the variant .alpha.-amylase or the parent .alpha.-amylase has at least 60% amino acid sequence identity relative to SEQ ID NO: 1, which is used for numbering; and wherein the variant has increased thermostability, detergent stability, starch liquifaction activity, and/or cleaning performance compared to the parent .alpha.-amylase or a reference .alpha.-amylase differing from the variant .alpha.-amylase only by the absence of the mutations.
[0009] 2. In some embodiments, the variant .alpha.-amylase of paragraph 1 comprises at least two mutations at amino acid residues corresponding N126, Y150, F153, L171, and, I203, using SEQ ID NO: 1 for numbering.
[0010] 3. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a deletion of at least one amino acid residue corresponding to R178, G179, T180, and G181, using SEQ ID NO: 1 for numbering.
[0011] 4. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprise deletions of amino acid residues corresponding to R178 and G179, or T180 and G181.
[0012] 5. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a mutation at an amino acid residue corresponding to G476 and/or G477, using SEQ ID NO: 1 for numbering.
[0013] 6. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a mutation in an amino acid residue corresponding to an amino acid residue selected from the group consisting of E132, Q167, T180, and A277, using SEQ ID NO: 1 for numbering.
[0014] 7. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a mutation in an amino acid residue corresponding to an amino acid residue selected from the group consisting of R458, T459, and D460, using SEQ ID NO: 1 for numbering.
[0015] 8. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a mutation in an amino acid residue corresponding to T180, using SEQ ID NO: 1 for numbering.
[0016] 9. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a mutation in an amino acid residue corresponding to N205, using SEQ ID NO: 3 for numbering.
[0017] 10. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a mutation in an amino acid residue corresponding to an amino acid residue selected from the group consisting of T333G, A335S, and Q337E, using SEQ ID NO: 3 for numbering.
[0018] 11. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs further comprises a mutation in an amino acid residue corresponding to an amino acid residue position selected from the group consisting of 6, 7, 8, 11, 14, 15, 20, 21, 23, 26, 27, 28, 37, 38, 39, 40, 42, 45, 46, 48, 49, 50, 51, 52, 53, 54, 58, 61, 62, 68, 70, 71, 72, 73, 79, 80, 81, 82, 84, 85, 87, 88, 89, 92, 93, 94, 95, 96, 97, 98, 101, 108, 111, 112, 113, 114, 115, 116, 117, 118, 120, 122, 123, 124, 126, 127, 129, 130, 131, 132, 133, 134, 136, 137, 138, 140, 142, 143, 144, 147, 148, 149, 150, 151, 152, 153, 154, 155, 156, 158, 159, 165, 167, 168, 170, 171, 172, 175, 176, 177, 180, 181, 182, 187, 190, 191, 193, 199, 200, 201, 203, 206, 208, 210, 211, 212, 214, 215, 216, 219, 221, 223, 225, 226, 227, 235, 238, 239, 240, 241, 242, 243, 245, 246, 247, 248, 249, 250, 252, 253, 254, 256, 257, 258, 260, 261, 262, 266, 267, 268, 269, 270, 271, 273, 276, 277, 279, 280, 282, 284, 285, 286, 288, 296, 299, 300, 301, 302, 303, 304, 307, 308, 310, 311, 312, 313, 316, 317, 318, 320, 321, 325, 327, 335, 338, 342, 348, 349, 352, 356, 357, 360, 362, 363, 368, 369, 377, 381, 382, 383, 384, 385, 388, 390, 392, 394, 395, 396, 397, 398, 400, 401, 402, 403, 404, 405, 407, 408, 410, 414, 415, 416, 418, 419, 420, 421, 422, 423, 424, 426, 428, 429, 430, 431, 434, 435, 436, 439, 441, 442, 444, 445, 446, 447, 448, 449, 450, 451, 454, 455, 457, 460, 461, 462, 463, 464, 465, 466, 467, 469, 470, 471, 473, 474, 475, 476, 477, 479, 480, 481, 482, 483, and 484, using SEQ ID NO: 1 for numbering.
[0019] 12. In some embodiments, the variant .alpha.-amylase of any of the preceding paragraphs comprises a combinations of mutations corresponding to mutations selected from the group consisting of: [0020] E187P+I203Y+G476K, [0021] E187P+I203Y+G476K+R458N+T459S+D460T, [0022] T180D+E187P+I203Y+G476K, [0023] N126Y+T180D+E187P+I203Y+G476K, [0024] N126Y+T180D+E187P+I203Y+Y303D+G476T+G477E, [0025] N126Y+T180D+E187P+I203Y+Y303D+N475E+G477Q, [0026] N126Y+T180D+E187P+I203Y+Y303R+N475E+G476T+G477R, [0027] T038N+N088H+N126Y+T129I+N134M+F153W+L171R+T180D+E187P+I203Y+G476K+G477E, [0028] N126Y+E132H+T180D+E187P+I203Y+Y303D+G476T+G477E, [0029] N126Y+E187P+G476K, [0030] N126Y+F153W+E187P+G476K, [0031] N126Y+F153W+E187P+G4726+G477R, [0032] N126Y+E187P+I203Y, [0033] N126Y+I203Y+S241Q, [0034] N126Y+T180H+E187P+I203Y, [0035] N126Y+T180H+I203Y+S241Q, [0036] N126Y+F153W+T180H+E187P+I203Y, [0037] N126Y+F153W+T180H+I203Y+S241Q, [0038] N126Y+Y150H+F153W+L171N+E187P+I203Y, [0039] N126Y+Y150H+F153W+L171N+I203Y+S241Q, [0040] N126Y+Y150H+F153W+L171N+T180H+E187P+I203Y, [0041] N126Y+Y150H+F153W+L171N+T180H+I203Y+S241Q, and [0042] N126Y+F153W+T180D+I203Y+S241Q;
[0043] wherein the variant has increased thermostability, detergent stability, stability starch liquifaction activity, or cleaning performance compared to the parent; and wherein the variant or the parent has at least 60% amino acid sequence identity relative to SEQ ID NO: 1, which is used for numbering.
[0044] 13. In some embodiments, the variant amylase of any of paragraphs 1-12 comprises the combinations of mutations corresponding to N126Y+F153W+T180D+I203Y+S241Q and one or more mutations corresponding to mutations selected from the group consisting of E132H, Q167E, A277F, and T400K.
[0045] 14. In some embodiments, the variant amylase of paragraph 13 comprises the combinations of mutations corresponding to mutations selected from the group consisting of: [0046] N126Y+E132H+F153W+T180D+I203Y+S241Q+A277F, [0047] N126Y+E132H+F153W+Q167E+T180D+I203Y+S241Q+A277F, and [0048] N126Y+E132H+F153W+Q167E+T180D+I203Y+S241Q+A277F+T400K.
[0049] 15. In some embodiments, the variant amylase of any of the preceding paragraphs is from a Cytophaga species.
[0050] 16. In some embodiments, the variant amylase of any of the preceding paragraphs is from a Paenibacillus species.
[0051] 17. In some embodiments, the variant amylase of any of the preceding paragraphs is not from a Bacillus species.
[0052] 18. In some embodiments, the variant amylase of any of the preceding paragraphs has at least 70% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5.
[0053] 19. In some embodiments, the variant amylase of any of the preceding paragraphs has at least 70% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1 or SEQ ID NO: 3.
[0054] 20. In some embodiments, the variant amylase of any of paragraphs 1-18 has at least 80% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5.
[0055] 21 In some embodiments, the variant amylase of any of paragraphs 1-18 has at least 90% amino acid sequence identity to the amino acid sequence of SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5.
[0056] 22. In another aspect, a composition comprising the variant .alpha.-amylase of any of the preceding paragraphs is provided.
[0057] 23. In some embodiments, the composition of paragraph 22 is effective for removing starchy stains from laundry, dishes, or textiles.
[0058] 24. In some embodiments, the composition of paragraph 22 or 23 further comprises a surfactant.
[0059] 25. In some embodiments, the composition of any of paragraphs 22-24 is a detergent composition.
[0060] 26. In some embodiments, the composition of any of paragraphs 22-24 is a laundry detergent or a laundry detergent additive.
[0061] 27. In some embodiments, the composition of any of paragraphs 22-24 is a manual or automatic dishwashing detergent.
[0062] 28. In some embodiments, the composition of any of paragraphs 22-24 further comprises one or more additional enzymes selected from the group consisting of protease, hemicellulase, cellulase, peroxidase, lipolytic enzyme, metallolipolytic enzyme, xylanase, lipase, phospholipase, esterase, perhydrolase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, .beta.-glucanase, arabinosidase, hyaluronidase, chondroitinase, laccase, metalloproteinase, amadoriase, glucoamylase, arabinofuranosidase, phytase, isomerase, transferase, and an amylase other than the amylase of any one of paragraphs 1-21.
[0063] 29. In some embodiments, the composition of paragraph 21 is for liquifying starch.
[0064] 30. In some embodiments, the composition of paragraph 21 is for saccharifying a composition comprising starch, for SSF post liquefaction, or for direct SSF without prior liquefaction.
[0065] 31. In some embodiments, the composition of paragraph 21 is for producing a fermented beverage.
[0066] 32. In some embodiments, the composition of paragraph 21 is for producing a baked food product.
[0067] 33. In some embodiments, the composition of paragraph 21 is for textile desizing.
[0068] 34. In another aspect, a method for removing a starchy stain or soil from a surface is provided, comprising: contacting the surface in the presence of a composition comprising an effective amount of the variant amylase of any of the paragraphs 1-21, and allowing the polypeptide to hydrolyze starch components present in the starchy stain to produce smaller starch-derived molecules that dissolve in the aqueous composition, thereby removing the starchy stain from the surface.
[0069] 35. In some embodiments of the method of paragraph 34 the aqueous composition further comprises a surfactant.
[0070] 36. In some embodiments of the method of any of paragraphs 34 or 35 the surface is a textile surface or a surface on dishes.
[0071] 37. In some embodiments of the method of any of paragraphs 34-36 the composition further comprises at least one additional enzymes selected from the group consisting of protease, hemicellulase, cellulase, peroxidase, lipolytic enzyme, metallolipolytic enzyme, xylanase, lipase, phospholipase, esterase, perhydrolase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, .beta.-glucanase, arabinosidase, hyaluronidase, chondroitinase, laccase, metalloproteinase, amadoriase, glucoamylase, arabinofuranosidase, phytase, isomerase, transferase, and an amylase other than the amylase of any one of paragraphs 1-21.
[0072] 38. In another aspect, a method for saccharifying a composition comprising starch to produce a composition comprising glucose is provided, wherein the method comprises: [0073] (i) contacting the solution comprising starch with effective amount of the variant amylase of any of the paragraphs 1-21; and [0074] (ii) saccharifying the solution comprising starch to produce the composition comprising glucose; wherein the variant amylase catalyzes the saccharification of the starch solution to glucose or other enriched carbohydrate syrups.
[0075] 39. In some embodiments of the method of paragraph 38 the composition comprising starch comprises liquefied starch, gelatinized starch, granular starch, or starch heat-treated below its gelatinization temperature.
[0076] 40. In some embodiments of the method of paragraph 38 or 39 the fermentation is a simultaneous saccharification and fermentation (SSF) reaction.
[0077] 41. In some embodiments of the method of any of paragraphs 38-40 the method further comprises contacting a mash and/or a wort with an amylase.
[0078] 42. In some embodiments, the method of any one of paragraphs 38-41 further comprises adding glucoamylase, hexokinase, xylanase, glucose isomerase, xylose isomerase, phosphatase, phytase, pullulanase, .beta.-amylase, .alpha.-amylase that is not the variant .alpha.-amylase, protease, cellulase, hemicellulase, lipase, cutinase, isoamylase, redox enzyme, esterase, transferase, pectinase, alpha-glucosidase, beta-glucosidase, or a combination thereof, to the starch solution.
[0079] 43. In some embodiments of the method of any one of paragraphs 38-42 the amylase is expressed and secreted by a host cell.
[0080] 44. In some embodiments of the method of paragraph 43 the composition comprising starch is contacted with the host cell.
[0081] 45. In some embodiments of the method of paragraph 43 or 44 the host cell further expresses and secretes one or more enzymes selected from the group consisting of glucoamylase, hexokinase, xylanase, glucose isomerase, xylose isomerase, phosphatase, phytase, pullulanase, .beta.-amylase, .alpha.-amylase that is not the variant .alpha.-amylase, protease, cellulase, hemicellulase, lipase, cutinase, isoamylase, redox enzyme, esterase, transferase, pectinase, alpha-glucosidase, and beta-glucosidase.
[0082] 46. In some embodiments of the method of any one of paragraphs 43-45 the host cell further expresses and secretes a glucoamylase.
[0083] 47. In some embodiments of the method of any one of paragraphs 43-46 the host cell is capable of fermenting the composition.
[0084] 48. In another aspect, a composition comprising glucose produced by the method of any one of paragraphs 38-47 is provided.
[0085] 49. In another aspect, a liquefied starch produced by the method of any one of paragraphs 38-47 is provided.
[0086] 50. In another aspect, a fermented beverage produced by the method of any one of paragraphs 38-47 is provided.
[0087] 51. In another aspect, use of an amylase of any of paragraphs 1-21 in the production of a composition comprising glucose, in the production of a liquefied starch, in the production of a fermented beverage, in cleaning starchy stains, or in textile desizing, is provided.
[0088] 52. In another aspect, a method of desizing a textile is provided, comprising contacting a desizing composition with a sized textile for a time sufficient to desize the textile, wherein the desizing composition comprises a variant .alpha.-amylase of any one of paragraphs 1-21.
[0089] 53. In another aspect, an isolated polynucleotide encoding a polypeptide of any of paragraphs 1-21 is provided.
[0090] 54. In another aspect, an expression vector comprising the polynucleotide of paragraph 53 is provided.
[0091] 55. In another aspect, a host cell comprising the expression vector of paragraph 54 is provided
[0092] 56. In another aspect, a polypeptide according to any one of paragraphs 1-21 encoded by a polynucleotide that hybridizes under stringent conditions to a polynucleotide complementary to the full-length of the polynucleotide of SEQ ID NO: 7, SEQ ID NO: 33, or SEQ ID NO: 38 is provided.
[0093] These and other aspects and embodiments of the compositions and methods will be apparent from the present description and drawings.
BRIEF DESCRIPTION OF THE DRAWINGS
[0094] FIG. 1 shows an amino acid sequence alignment of CspAmy2 .alpha.-amylase (SEQ ID NO: 1), PcuAmy1 .alpha.-amylase (SEQ ID NO: 3), and BASE .alpha.-amylase, using Clustal W with default parameters.
[0095] FIG. 2 is a graph showing the cleaning benefit of different doses of CspAmy2-v5 and CspAmy2-v6 on CS-28 rice starch at pH 8.
[0096] FIG. 3 is a graph showing the thermal stability of CspAmy2-v5 and CspAmy2-v6 in buffer.
[0097] FIG. 4 is a graph showing the thermal stability of CspAmy2-v5 and CspAmy2-v6 in buffer with calcium.
[0098] FIG. 5 is a graph showing the detergent stability of CspAmy2-v5 and CspAmy2-v6 in OMO.TM. Color detergent.
[0099] FIG. 6 is a graph showing the detergent stability of CspAmy2-v5 and CspAmy2-v6 in EPSIL.TM. Perfect detergent.
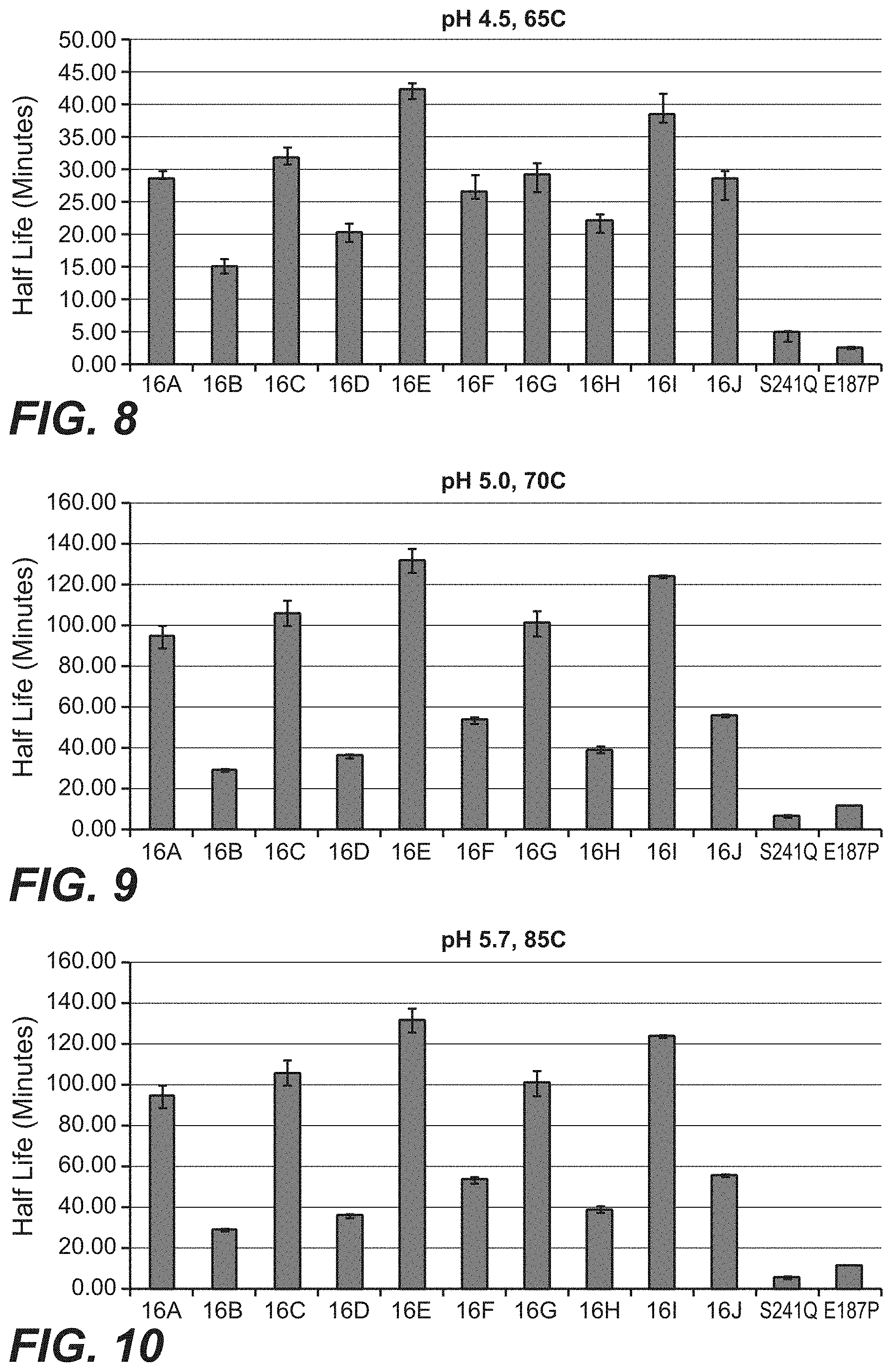
[0100] FIG. 7 is a table showing the relative half-lives and performance indexes of the C16 variants and reference molecules CspAmy2-v1-E187P and CspAmy2-v1-S241Q.
[0101] FIG. 8 is a graph showing the thermal stability of the C16 variants and reference molecules CspAmy2-v1-E187P and CspAmy2-v1-S241Q at pH 4.5 and 65.degree. C.
[0102] FIG. 9 is a graph showing the thermal stability of the C16 variants and reference molecules CspAmy2-v1-E187P and CspAmy2-v1-S241Q at pH 5.0 and 70.degree. C.
[0103] FIG. 10 is a graph showing the thermal stability of the C16 variants and reference molecules CspAmy2-v1-E187P and CspAmy2-v1-S241Q at pH 5.7 and 85.degree. C.
[0104] FIG. 11 is a graph showing the detergent stability of CspAmy2-v5, CspAmy2-v171, CspAmy2-v172, and ACE-QK.
[0105] FIG. 12 is a graph showing the relative cleaning performance of CspAmy2-v5 and STAINZYME.RTM. in a hand dishwashing application.
[0106] FIG. 13 FIGS. 13A and 13B, includes are tables showing the compositions of WfK B citrate-based detergent (FIG. 13A) and WfK C phospate-based detergent (FIG. 13B).
[0107] FIGS. 14 and 15 show the cleaning performance of CspAmy2-v6 (squares) compared to POWERASE.RTM. (diamonds), dosed at 0, 1, 2, 4, or 8 ppm in WfK B detergent against the mixed starch stain (FIG. 14) and the pasta stain (FIG. 15).
[0108] FIGS. 16 and 17 show the cleaning performance of CspAmy2-v6 (squares) compared to STAINZYME.RTM. (circles), dosed at 0, 1, 2, 4, or 8 ppm WfK B detergent against the mixed starch stain (FIG. 16) and the pasta stain (FIG. 17).
[0109] FIGS. 18 and 19 show the cleaning performance of CspAmy2-v6 (squares) compared to POWERASE.RTM. (diamonds), dosed at 0, 1, 2, 4, or 8 ppm in WfK C detergent against the mixed starch stain (FIG. 18) and the pasta stain (FIG. 19). CspAmy2-v6 clearly outperformed POWERASE.RTM. against both stains.
[0110] FIG. 20 is a graph showing examples of C18P variants demonstrating improved hydrolysis of corn starch at high temperatures. CspAmy2-C18P (N126Y+F153W+T180D+I203Y+S241Q) is shown as a reference.
[0111] FIG. 21 is a graph showing examples of C18P variants demonstrating improved hydrolysis of amylopectin from corn. C18P is shown as a reference.
[0112] FIG. 22 is a graph showing examples of variants demonstrating improved generation of reducing sugars from starch. C18P is shown as a reference.
[0113] FIG. 23 is a graph showing examples of C18P variants demonstrating improved release of iodine staining material from starch. C18P is shown as a reference.
[0114] FIG. 24 is a graph showing the viscosity reduction of corn flour slurry produced by three C18P variants reported as fluidity (1/viscosity) versus dose of the variants (in .mu.g). C18P and C16F are shown as references.
[0115] FIG. 25 is a table showing the PI values for C16F variants having different pairwise combinations of mutations at positions G476 and G477, relative to a C16F control, in a corn starch microswatch assay. PI values for revertants (i.e., G476G and G477G) are empirically determined.
[0116] FIG. 26 is a table showing the PI values for C16F variants having different pairwise combinations of mutations at positions G476 and G477, relative to a C16F control, in a corn amylose hydrolysis assay. PI values for revertants (i.e., G476G and G477G) are empirically determined.
[0117] FIG. 27 is a graph showing the relative liquefaction performance of CspAmy2-C25F, B, and A compared to C16F.
[0118] FIG. 28 is a graph showing the results of cleaning assays performed at 0.015 ppm with CspAmy2-v179, v186, and v191 compared to STAINZYME.RTM. and ACE-QK.
[0119] FIG. 29 is a graph showing the relative thermostability of CspAmy2 variants v5, v179, v186, and v191 compared to STAINZYME.RTM. and ACE-QK at temperatures ranging from 77.degree. C. to 97.degree. C.
[0120] FIG. 30 is a graph showing the relative in-detergent storage stability of CspAmy2 variants v5 and v179 compared to STAINZYME.RTM. and ACE-QK in TIDE.RTM. regular HDL.
[0121] FIG. 31 is a graph showing the relative in-detergent storage stability of CspAmy2 variants v5 and v179 compared to STAINZYME.RTM. and ACE-QK in US TIDE.RTM. PODS.TM..
[0122] FIG. 32 is a graph showing the relative in-detergent storage stability of CspAmy2 variants v5 and v179 compared to STAINZYME.RTM. and ACE-QK in European ARIEL.TM. HDL.
[0123] FIG. 33 is a graph showing the relative in-detergent storage stability of CspAmy2 variants v5 and v179 compared to STAINZYME.RTM. and ACE-QK in European OMO.TM. Color HDL.
[0124] FIG. 34 is a graph showing the relative in-detergent storage stability of CspAmy2 variants v5 and v179 compared to STAINZYME.RTM. and ACE-QK in Chinese OMO.TM. Color HDL.
[0125] FIG. 35 is a graph showing the relative in-detergent storage stability of CspAmy2 variants v5 and v179 compared to STAINZYME.RTM. and ACE-QK in Chinese LIBY.TM. HDL.
[0126] FIG. 36 is a graph showing the relative cleaning performance of PcuAmy1 variants v1, v6, v8, and v16 compared to STAINZYME.RTM. and ACE-QK in buffer at pH 8.0. Enzyme doses are noted on the x-axis.
[0127] FIG. 37 is a graph showing the relative thermal stability of PcuAmy1 variants v1, v6, v8, and v16 compared to STAINZYME.RTM. in buffer at the temperatures indicated 5 ppm of PcuAmy1 variants and 10 ppm of STAINZYME.RTM. were used.
[0128] FIG. 38 is a graph showing the relative thermal stability of .DELTA.RG BASE variants incubated for the indicated amounts of time at 95.degree. C.
[0129] FIG. 39 shows a portion of the three-dimensional structure of CspAmy2-v1 highlighting the potential for interaction between a glutamate at position 132 and a threonine at position 180.
[0130] FIG. 40 shows a portion of the three-dimensional structure of CspAmy2-v1 highlighting the potential for interaction between a glutamate at position 132 and a histidine at position 180.
[0131] FIG. 41 shows a portion of the three-dimensional structure of CspAmy2-v1 highlighting the potential for interaction between a glutamate at position 132 and an aspartate at position 180.
[0132] FIG. 42 shows a portion of the three-dimensional structure of CspAmy2-v1 highlighting the potential for interaction between a histidine at position 132 and an aspartate at position 180.
[0133] FIG. 43 is an image of an SDS/PAGE gel showing the cleavage of PcuAmy1-v1 in the presence of increasing amounts of GG36 protease. The letters on the right side of the gel indicate (A) intact full-length PcuAmy1-v1, (B) a first cleavage product of PcuAmy1-v1, (C) GG36 protease, (D) a contaminant in the GG36 protein preparation, and (E) a second cleavage product of PcuAmy1-v1.
[0134] FIG. 44 is a graph showing the residual .alpha.-amylase activity of PcuAmy1 and several engineered variants following incubation with GG36 protease.
[0135] FIG. 45 is an image of an SDS/PAGE gel showing the proteolytic cleavage of PcuAmy1 and several engineered variants following incubation with GG36 protease.
[0136] FIG. 46 is a graph showing the stability of PcuAmy1-v1 and several engineered variants following incubation with GG36 protease in MIFA Total detergent for up to 14 days at 37.degree. C.
[0137] FIG. 47 is a graph showing the stability of PcuAmy1-v1 and several engineered variants following incubation with GG36 protease in MIFA Total detergent for 3 or 14 days at 37.degree. C.
[0138] FIG. 48 is a graph showing the stability of PcuAmy1-v1 and several engineered variants following incubation with GG36 protease in Unilever Omo detergent for up to 14 days at 37.degree. C.
[0139] FIG. 49 is a graph showing the stability of PcuAmy1-v1 and several engineered variants following incubation with GG36 protease in Unilever Omo detergent for 3 or 14 days at 37.degree. C.
[0140] FIG. 50 is a graph showing the dose-dependent cleaning performance of PcuAmy1-3B and PcuAmy1-3L in buffer at pH 8.0 compared to two commercial benchmarks.
[0141] FIG. 51 is a graph showing the stability of PcuAmy1-3B and PcuAmy1-3L in Persil Universal Gel Gold detergent compared to two commercial benchmarks.
[0142] FIG. 52 is a graph showing the stability of PcuAmy1-v1 and several engineered variants following incubation with GG36 protease in MIFA Total detergent for 3 or 14 days at 37.degree. C.
DETAILED DESCRIPTION
[0143] Described are compositions and methods relating to variant amylase enzymes. The variants were discovered by a combination of experimental approaches, as detailed in the appended Examples. The approaches include the use of site evaluation libraries (SELs) and structure-based analysis. Exemplary applications for the variant amylase enzymes are for starch liquefaction and saccharification, for cleaning starchy stains in laundry, dishwashing, and other applications, for textile processing (e.g., desizing), in animal feed for improving digestibility, and for baking and brewing. These and other aspects of the compositions and methods are described in detail, below.
[0144] Prior to describing the various aspects and embodiments of the present compositions and methods, the following definitions and abbreviations are described.
1. Definitions and Abbreviations
[0145] In accordance with this detailed description, the following abbreviations and definitions apply. Note that the singular forms "a," "an," and "the" include plural referents unless the context clearly dictates otherwise. Thus, for example, reference to "an enzyme" includes a plurality of such enzymes, and reference to "the dosage" includes reference to one or more dosages and equivalents thereof known to those skilled in the art, and so forth.
[0146] The present document is organized into a number of sections for ease of reading; however, the reader will appreciate that statements made in one section may apply to other sections. In this manner, the headings used for different sections of the disclosure should not be construed as limiting.
[0147] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art. The following terms are provided below.
1.1. Abbreviations and Acronyms
[0148] The following abbreviations/acronyms have the following meanings unless otherwise specified:
[0149] ABTS 2,2-azino-bis-3-ethylbenzothiazoline-6-sulfonic acid
[0150] AE or AEO alcohol ethoxylate
[0151] AES or AEOS alcohol ethoxysulfate
[0152] AkAA Aspergillus kawachii .alpha.-amylase
[0153] AnGA Aspergillus niger glucoamylase
[0154] AOS .alpha.-olefinsulfonate
[0155] AS alkyl sulfate
[0156] cDNA complementary DNA
[0157] ct/kg cents/kg (US currency)
[0158] CMC carboxymethylcellulose
[0159] DE dextrose equivalent
[0160] DNA deoxyribonucleic acid
[0161] DPn degree of saccharide polymerization having n subunits
[0162] ds or DS dry solids
[0163] DTMPA diethylenetriaminepentaacetic acid
[0164] EC Enzyme Commission
[0165] EDTA ethylenediaminetetraacetic acid
[0166] EO ethylene oxide (polymer fragment)
[0167] EOF end of fermentation
[0168] FH French hardness
[0169] GA glucoamylase
[0170] GAU/g ds glucoamylase activity unit/gram dry solids
[0171] GH general hardness
[0172] HDL high density liquid detergent
[0173] HDD heavy duty powder detergent
[0174] HSG high suds granular detergent
[0175] HFCS high fructose corn syrup
[0176] HgGA Humicola grisea glucoamylase
[0177] IPTG isopropyl .beta.-D-thiogalactoside
[0178] IRS insoluble residual starch
[0179] kDa kiloDalton
[0180] LAS linear alkylbenzenesulfonate
[0181] LAT, BLA B. licheniformis amylase
[0182] MW molecular weight
[0183] MWU modified Wohlgemuth unit; 1.6.times.10.sup.-5 mg/MWU=unit of activity
[0184] NCBI National Center for Biotechnology Information
[0185] NOBS nonanoyloxybenzenesulfonate
[0186] NTA nitriloacetic acid
[0187] OxAm Purastar HPAM 5000L (Danisco US Inc.)
[0188] PAHBAH p-hydroxybenzoic acid hydrazide
[0189] PEG polyethyleneglycol
[0190] pI isoelectric point
[0191] PI performance index
[0192] ppm parts per million, e.g., .mu.g protein per gram dry solid
[0193] PVA poly(vinyl alcohol)
[0194] PVP poly(vinylpyrrolidone)
[0195] RCF relative centrifugal/centripetal force (i.e., x gravity)
[0196] RNA ribonucleic acid
[0197] SAS alkanesulfonate
[0198] SDS-PAGE sodium dodecyl sulfate polyacrylamide gel electrophoresis
[0199] SSF simultaneous saccharification and fermentation
[0200] SSU/g solid soluble starch unit/gram dry solids
[0201] sp. species
[0202] TAED tetraacetylethylenediamine
[0203] Tm melting temperature
[0204] TrGA Trichoderma reesei glucoamylase
[0205] w/v weight/volume
[0206] w/w weight/weight
[0207] v/v volume/volume
[0208] wt % weight percent
[0209] .degree. C. degrees Centigrade
[0210] H.sub.2O water
[0211] dH.sub.2O or DI deionized water
[0212] dIH.sub.2O deionized water, Milli-Q filtration
[0213] g or gm grams
[0214] micrograms
[0215] mg milligrams
[0216] kg kilograms
[0217] .mu.L and .mu.l microliters
[0218] mL and ml milliliters
[0219] mm millimeters
[0220] micrometer
[0221] M molar
[0222] mM millimolar
[0223] .mu.M micromolar
[0224] U units
[0225] sec seconds
[0226] min(s) minute/minutes
[0227] hr(s) hour/hours
[0228] DO dissolved oxygen
[0229] Ncm Newton centimeter
[0230] ETOH ethanol
[0231] eq. equivalents
[0232] N normal
[0233] uPWA variant .alpha.-amylase derived from Pyrococcus woesei
[0234] PWA .alpha.-amylase from Pyrococcus woesei
[0235] MWCO molecular weight cut-off
[0236] SSRL Stanford Synchrotron Radiation Lightsource
[0237] PDB Protein Database
[0238] CAZy Carbohydrate-Active Enzymes database
[0239] Tris-HCl tris(hydroxymethyl)aminomethane hydrochloride
[0240] HEPES 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
1.2. Definitions
[0241] The terms "amylase" or "amylolytic enzyme" refer to an enzyme that is, among other things, capable of catalyzing the degradation of starch. .alpha.-Amylases are hydrolases that cleave the .alpha.-D-(1.fwdarw.4) 0-glycosidic linkages in starch. Generally, .alpha.-amylases (EC 3.2.1.1; .alpha.-D-(1.fwdarw.4)-glucan glucanohydrolase) are defined as endo-acting enzymes cleaving .alpha.-D-(1.fwdarw.4) 0-glycosidic linkages within the starch molecule in a random fashion yielding polysaccharides containing three or more (1-4)-.alpha.-linked D-glucose units. In contrast, the exo-acting amylolytic enzymes, such as .beta.-amylases (EC 3.2.1.2; .alpha.-D-(1.fwdarw.4)-glucan maltohydrolase) and some product-specific amylases like maltogenic .alpha.-amylase (EC 3.2.1.133) cleave the polysaccharide molecule from the non-reducing end of the substrate. .beta.-amylases, .alpha.-glucosidases (EC 3.2.1.20; .alpha.-D-glucoside glucohydrolase), glucoamylase (EC 3.2.1.3; .alpha.-D-(1.fwdarw.4)-glucan glucohydrolase), and product-specific amylases like the maltotetraosidases (EC 3.2.1.60) and the maltohexaosidases (EC 3.2.1.98) can produce malto-oligosaccharides of a specific length or enriched syrups of specific maltooligosaccharides.
[0242] "Enzyme units" herein refer to the amount of product formed per time under the specified conditions of the assay. For example, a "glucoamylase activity unit" (GAU) is defined as the amount of enzyme that produces 1 g of glucose per hour from soluble starch substrate (4% DS) at 60.degree. C., pH 4.2. A "soluble starch unit" (SSU) is the amount of enzyme that produces 1 mg of glucose per minute from soluble starch substrate (4% DS) at pH 4.5, 50.degree. C. DS refers to "dry solids."
[0243] The term "starch" refers to any material comprised of the complex polysaccharide carbohydrates of plants, comprised of amylose and amylopectin with the formula (C6H10O5)x, wherein X can be any number. The term includes plant-based materials such as grains, cereal, grasses, tubers and roots, and more specifically materials obtained from wheat, barley, corn, rye, rice, sorghum, brans, cassava, millet, milo, potato, sweet potato, and tapioca. The term "starch" includes granular starch. The term "granular starch" refers to raw, i.e., uncooked starch, e.g., starch that has not been subject to gelatinization.
[0244] The terms, "wild-type," "parental," or "reference," with respect to a polypeptide, refer to a naturally-occurring polypeptide that does not include a man-made substitution, insertion, or deletion at one or more amino acid positions. Similarly, the terms "wild-type," "parental," or "reference," with respect to a polynucleotide, refer to a naturally-occurring polynucleotide that does not include a man-made nucleoside change. However, note that a polynucleotide encoding a wild-type, parental, or reference polypeptide is not limited to a naturally-occurring polynucleotide, and encompasses any polynucleotide encoding the wild-type, parental, or reference polypeptide.
[0245] Reference to the wild-type polypeptide is understood to include the mature form of the polypeptide. A "mature" polypeptide or variant, thereof, is one in which a signal sequence is absent, for example, cleaved from an immature form of the polypeptide during or following expression of the polypeptide.
[0246] The term "variant," with respect to a polypeptide, refers to a polypeptide that differs from a specified wild-type, parental, or reference polypeptide in that it includes one or more naturally-occurring or man-made substitutions, insertions, or deletions of an amino acid. Similarly, the term "variant," with respect to a polynucleotide, refers to a polynucleotide that differs in nucleotide sequence from a specified wild-type, parental, or reference polynucleotide. The identity of the wild-type, parental, or reference polypeptide or polynucleotide will be apparent from context.
[0247] In the case of the present .alpha.-amylases, "activity" refers to .alpha.-amylase activity, which can be measured as described, herein.
[0248] The term "performance benefit" refers to an improvement in a desirable property of a molecule. Exemplary performance benefits include, but are not limited to, increased hydrolysis of a starch substrate, increased grain, cereal or other starch substrate liquifaction performance, increased cleaning performance, increased thermal stability, increased detergent stability, increased storage stability, increased solubility, an altered pH profile, decreased calcium dependence, increased specific activity, modified substrate specificity, modified substrate binding, modified pH-dependent activity, modified pH-dependent stability, increased oxidative stability, and increased expression. In some cases, the performance benefit is realized at a relatively low temperature. In some cases, the performance benefit is realized at relatively high temperature.
[0249] The terms "protease" and "proteinase" refer to an enzyme protein that has the ability to perform "proteolysis" or "proteolytic cleavage" which refers to hydrolysis of peptide bonds that link amino acids together in a peptide or polypeptide chain forming the protein. This activity of a protease as a protein-digesting enzyme is referred to as "proteolytic activity." Many well-known procedures exist for measuring proteolytic activity (See e.g., Kalisz, "Microbial Proteinases," In: Fiechter (ed.), Advances in Biochemical Engineering/Biotechnology, (1988)). For example, proteolytic activity may be ascertained by comparative assays which analyze the respective protease's ability to hydrolyze a commercial substrate. Exemplary substrates useful in the analysis of protease or proteolytic activity, include, but are not limited to, di-methyl casein (Sigma C-9801), bovine collagen (Sigma C-9879), bovine elastin (Sigma E-1625), and bovine keratin (ICN Biomedical 902111). Colorimetric assays utilizing these substrates are well known in the art (See e.g., WO 99/34011 and U.S. Pat. No. 6,376,450, both of which are incorporated herein by reference). The pNA assay (See e.g., Del Mar et al., Anal. Biochem. 99:316-320 [1979]) also finds use in determining the active enzyme concentration for fractions collected during gradient elution. This assay measures the rate at which p-nitroaniline is released as the enzyme hydrolyzes a soluble synthetic peptide substrate, such as succinyl-alanine-alanine-proline-phenylalanine-p-nitroanilide (suc-AAPF-pNA), and cleavage occurs between the C-terminal amino acid (phenylalanine) and the p-NA, causing the production of yellow color from the hydrolysis reaction, which is measured at 410 nm on a spectrophotometer and is proportional to the active enzyme concentration. Measurement of the color change allows calculation of the rate of the reaction. In addition, absorbance measurements at 280 nanometers (nm) can be used to determine the total protein concentration. The active enzyme/total protein ratio gives the enzyme purity when a reference standard is used.
[0250] The terms "serine protease" refers to enzymes that cleave peptide bonds in proteins, in which enzymes serine serves as the nucleophilic amino acid at the enzyme active site. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like. Most commonly used in laundry and dishwashing detergents are serine protease, particularly subtlisins.
[0251] The term "TIM barrel" refers to a three dimensional polypeptide structure that include eight .alpha.-helices and eight parallel .beta.-strands that alternate along the peptide backbone.
[0252] The term "surface-exposed" with respect to an amino acid residue in a polypeptide refers to a residue that is on the exterior surface of a polypeptide when the polypeptide is intact and properly folded, i.e., not denatured or fragmented. In the case of an .alpha.-amylase, the structure is referred to as a TIM barrel.
[0253] The term "non-canonical" with reference to an amino acid residue in a polypeptide refers to a residue that is not normally found at a given position based on amino acid sequence alignments of similar molecules using Clustal W with default parameter. In some cases, the particular residue is found at a given position in only 1 in 10, 1 in 20, 1 in 30, 1 in 50, or even 1 in 100 similar molecules.
[0254] "Combinatorial variants" are variants comprising two or more mutations, e.g., 2, 3, 4, 5, 6, 7, 8, 9, 10, or more, substitutions, deletions, and/or insertions.
[0255] "Combinable mutations" are mutations at any amino acid position that can be used to make combinatorial variants. Combinable mutations improve at least one desired property of the molecule (in this case, an amylase), while not significantly decreasing either expression, activity, or stability.
[0256] Terms, such as "a remaining non-G residue in the calcium-binding loop," "a non-G amino acid residue remaining in the calcium-binding loop," and similar terms, refer to an amino acid residue in the calcium-binding loop of a variant .alpha.-amylase, which remains in the variant following a deletion of at least one amino acid residue in the calcium-binding loop of a parent .alpha.-amylases, and which is not a glycine residue. The non-G residue may be a member of an "XG" pair, of which there are two in most .alpha.-amylases, and may be the remaining non-G residue following a pair-wise deletion of one of the two XG residue pairs in the calcium binding loop of a parent .alpha.-amylase.
[0257] A "stabilizing interaction" between the residue at position 132 (using SEQ ID NO: 1 for numbering) and the remaining non-G residue in the X.sub.1G/S.sub.1X.sub.2G.sub.2 motif (corresponding to residues at positions 178-181 of SEQ ID NO: 1) refers to a hydrogen bond or a salt bridge formed between the side chains of the subject amino acid residues. The stabilization can result from charge balancing the interacting residues, such that if one residue is positively charged at a preselected pH, the other is negatively charged, and the overall charge is zero.
[0258] The term "recombinant," when used in reference to a subject cell, nucleic acid, protein or vector, indicates that the subject has been modified from its native state. Thus, for example, recombinant cells express genes that are not found within the native (non-recombinant) form of the cell, or express native genes at different levels or under different conditions than found in nature. Recombinant nucleic acids differ from a native sequence by one or more nucleotides and/or are operably linked to heterologous sequences, e.g., a heterologous promoter in an expression vector. Recombinant proteins may differ from a native sequence by one or more amino acids and/or are fused with heterologous sequences. A vector comprising a nucleic acid encoding an amylase is a recombinant vector.
[0259] The terms "recovered," "isolated," and "separated," refer to a compound, protein (polypeptides), cell, nucleic acid, amino acid, or other specified material or component that is removed from at least one other material or component with which it is naturally associated as found in nature. An "isolated" polypeptides, thereof, includes, but is not limited to, a culture broth containing secreted polypeptide expressed in a heterologous host cell.
[0260] The term "purified" refers to material (e.g., an isolated polypeptide or polynucleotide) that is in a relatively pure state, e.g., at least about 90% pure, at least about 95% pure, at least about 98% pure, or even at least about 99% pure.
[0261] The term "enriched" refers to material (e.g., an isolated polypeptide or polynucleotide) that is in about 50% pure, at least about 60% pure, at least about 70% pure, or even at least about 70% pure.
[0262] The terms "thermostable" and "thermostability," with reference to an enzyme, refer to the ability of the enzyme to retain activity after exposure to an elevated temperature. The thermostability of an enzyme, such as an amylase enzyme, is measured by its half-life (t1/2) given in minutes, hours, or days, during which half the enzyme activity is lost under defined conditions. The half-life may be calculated by measuring residual .alpha.-amylase activity following exposure to (i.e., challenge by) an elevated temperature.
[0263] A "pH range," with reference to an enzyme, refers to the range of pH values under which the enzyme exhibits catalytic activity.
[0264] The terms "pH stable" and "pH stability," with reference to an enzyme, relate to the ability of the enzyme to retain activity over a wide range of pH values for a predetermined period of time (e.g., 15 min., 30 min., 1 hour).
[0265] The term "amino acid sequence" is synonymous with the terms "polypeptide," "protein," and "peptide," and are used interchangeably. Where such amino acid sequences exhibit activity, they may be referred to as an "enzyme." The conventional one-letter or three-letter codes for amino acid residues are used, with amino acid sequences being presented in the standard amino-to-carboxy terminal orientation (i.e., N.fwdarw.C).
[0266] The term "nucleic acid" encompasses DNA, RNA, heteroduplexes, and synthetic molecules capable of encoding a polypeptide. Nucleic acids may be single stranded or double stranded, and may be chemical modifications. The terms "nucleic acid" and "polynucleotide" are used interchangeably. Because the genetic code is degenerate, more than one codon may be used to encode a particular amino acid, and the present compositions and methods encompass nucleotide sequences that encode a particular amino acid sequence. Unless otherwise indicated, nucleic acid sequences are presented in 5'-to-3' orientation.
[0267] "Hybridization" refers to the process by which one strand of nucleic acid forms a duplex with, i.e., base pairs with, a complementary strand, as occurs during blot hybridization techniques and PCR techniques. Stringent hybridization conditions are exemplified by hybridization under the following conditions: 65.degree. C. and 0.1.times.SSC (where 1.times.SSC=0.15 M NaCl, 0.015 M Na3 citrate, pH 7.0). Hybridized, duplex nucleic acids are characterized by a melting temperature (Tm), where one half of the hybridized nucleic acids are unpaired with the complementary strand. Mismatched nucleotides within the duplex lower the Tm. A nucleic acid encoding a variant .alpha.-amylase may have a Tm reduced by 1.degree. C.-3.degree. C. or more compared to a duplex formed between the nucleotide of SEQ ID NO: 2 and its identical complement.
[0268] A "synthetic" molecule is produced by in vitro chemical or enzymatic synthesis rather than by an organism.
[0269] The terms "transformed," "stably transformed," and "transgenic," used with reference to a cell means that the cell contains a non-native (e.g., heterologous) nucleic acid sequence integrated into its genome or carried as an episome that is maintained through multiple generations.
[0270] The term "introduced" in the context of inserting a nucleic acid sequence into a cell, means "transfection", "transformation" or "transduction," as known in the art.
[0271] A "host strain" or "host cell" is an organism into which an expression vector, phage, virus, or other DNA construct, including a polynucleotide encoding a polypeptide of interest (e.g., an amylase) has been introduced. Exemplary host strains are microorganism cells (e.g., bacteria, filamentous fungi, and yeast) capable of expressing the polypeptide of interest and/or fermenting saccharides. The term "host cell" includes protoplasts created from cells.
[0272] The term "heterologous" with reference to a polynucleotide or protein refers to a polynucleotide or protein that does not naturally occur in a host cell.
[0273] The term "endogenous" with reference to a polynucleotide or protein refers to a polynucleotide or protein that occurs naturally in the host cell.
[0274] The term "expression" refers to the process by which a polypeptide is produced based on a nucleic acid sequence. The process includes both transcription and translation.
[0275] A "selective marker" or "selectable marker" refers to a gene capable of being expressed in a host to facilitate selection of host cells carrying the gene. Examples of selectable markers include but are not limited to antimicrobials (e.g., hygromycin, bleomycin, or chloramphenicol) and/or genes that confer a metabolic advantage, such as a nutritional advantage on the host cell.
[0276] A "vector" refers to a polynucleotide sequence designed to introduce nucleic acids into one or more cell types. Vectors include cloning vectors, expression vectors, shuttle vectors, plasmids, phage particles, cassettes and the like.
[0277] An "expression vector" refers to a DNA construct comprising a DNA sequence encoding a polypeptide of interest, which coding sequence is operably linked to a suitable control sequence capable of effecting expression of the DNA in a suitable host. Such control sequences may include a promoter to effect transcription, an optional operator sequence to control transcription, a sequence encoding suitable ribosome binding sites on the mRNA, enhancers and sequences which control termination of transcription and translation.
[0278] The term "operably linked" means that specified components are in a relationship (including but not limited to juxtaposition) permitting them to function in an intended manner. For example, a regulatory sequence is operably linked to a coding sequence such that expression of the coding sequence is under control of the regulatory sequences.
[0279] A "signal sequence" is a sequence of amino acids attached to the N-terminal portion of a protein, which facilitates the secretion of the protein outside the cell. The mature form of an extracellular protein lacks the signal sequence, which is cleaved off during the secretion process.
[0280] "Biologically active" refer to a sequence having a specified biological activity, such an enzymatic activity.
[0281] The term "specific activity" refers to the number of moles of substrate that can be converted to product by an enzyme or enzyme preparation per unit time under specific conditions. Specific activity is generally expressed as units (U)/mg of protein.
[0282] As used herein, "water hardness" is a measure of the minerals (e.g., calcium and magnesium) present in water.
[0283] A "swatch" is a piece of material such as a fabric that has a stain applied thereto. The material can be, for example, fabrics made of cotton, polyester or mixtures of natural and synthetic fibers. The swatch can further be paper, such as filter paper or nitrocellulose, or a piece of a hard material such as ceramic, metal, or glass. For amylases, the stain is starch based, but can include blood, milk, ink, grass, tea, wine, spinach, gravy, chocolate, egg, cheese, clay, pigment, oil, or mixtures of these compounds.
[0284] A "smaller swatch" is a section of the swatch that has been cut with a single hole punch device, or has been cut with a custom manufactured 96-hole punch device, where the pattern of the multi-hole punch is matched to standard 96-well microtiter plates, or the section has been otherwise removed from the swatch. The swatch can be of textile, paper, metal, or other suitable material. The smaller swatch can have the stain affixed either before or after it is placed into the well of a 24-, 48- or 96-well microtiter plate. The smaller swatch can also be made by applying a stain to a small piece of material. For example, the smaller swatch can be a stained piece of fabric 5/8'' or 0.25'' in diameter. The custom manufactured punch is designed in such a manner that it delivers 96 swatches simultaneously to all wells of a 96-well plate. The device allows delivery of more than one swatch per well by simply loading the same 96-well plate multiple times. Multi-hole punch devices can be conceived of to deliver simultaneously swatches to any format plate, including but not limited to 24-well, 48-well, and 96-well plates. In another conceivable method, the soiled test platform can be a bead made of metal, plastic, glass, ceramic, or another suitable material that is coated with the soil substrate. The one or more coated beads are then placed into wells of 96-, 48-, or 24-well plates or larger formats, containing suitable buffer and enzyme.
[0285] "A cultured cell material comprising an amylase" or similar language, refers to a cell lysate or supernatant (including media) that includes an amylase as a component. The cell material may be from a heterologous host that is grown in culture for the purpose of producing the amylase.
[0286] "Percent sequence identity" means that a particular sequence has at least a certain percentage of amino acid residues identical to those in a specified reference sequence, when aligned using the CLUSTAL W algorithm with default parameters. See Thompson et al. (1994) Nucleic Acids Res. 22:4673-4680. Default parameters for the CLUSTAL W algorithm are: [0287] Gap opening penalty: 10.0 [0288] Gap extension penalty: 0.05 [0289] Protein weight matrix: BLOSUM series [0290] DNA weight matrix: IUB [0291] Delay divergent sequences %: 40 [0292] Gap separation distance: 8 [0293] DNA transitions weight: 0.50 [0294] List hydrophilic residues: GPSNDQEKR [0295] Use negative matrix: OFF [0296] Toggle Residue specific penalties: ON [0297] Toggle hydrophilic penalties: ON [0298] Toggle end gap separation penalty OFF.
[0299] Deletions are counted as non-identical residues, compared to a reference sequence. Deletions occurring at either termini are included. For example, a variant with five amino acid deletions of the C-terminus of the mature CspAmy2 polypeptide of SEQ ID NO: 1 would have a percent sequence identity of 99% (612/617 identical residues.times.100, rounded to the nearest whole number) relative to the mature polypeptide. Such a variant would be encompassed by a variant having "at least 99% sequence identity" to a mature amylase polypeptide.
[0300] "Fused" polypeptide sequences are connected, i.e., operably linked, via a peptide bond between two subject polypeptide sequences.
[0301] The term "filamentous fungi" refers to all filamentous forms of the subdivision Eumycotina, particulary Pezizomycotina species.
[0302] The term "degree of polymerization" (DP) refers to the number (n) of anhydro-glucopyranose units in a given saccharide. Examples of DP1 are the monosaccharides glucose and fructose. Examples of DP2 are the disaccharides maltose and sucrose. The term "DE," or "dextrose equivalent," is defined as the percentage of reducing sugar, i.e., D-glucose, as a fraction of total carbohydrate in a syrup.
[0303] The term "dry solids content" (ds) refers to the total solids of a slurry in a dry weight percent basis. The term "slurry" refers to an aqueous mixture containing insoluble solids.
[0304] The phrase "simultaneous saccharification and fermentation (SSF)" refers to a process in the production of biochemicals in which a microbial organism, such as an ethanologenic microorganism, and at least one enzyme, such as an amylase, are present during the same process step. SSF includes the contemporaneous hydrolysis of starch substrates (granular, liquefied, or solubilized) to saccharides, including glucose, and the fermentation of the saccharides into alcohol or other biochemical or biomaterial in the same reactor vessel.
[0305] An "ethanologenic microorganism" refers to a microorganism with the ability to convert a sugar or oligosaccharide to ethanol.
[0306] The term "fermented beverage" refers to any beverage produced by a method comprising a fermentation process, such as a microbial fermentation, e.g., a bacterial and/or fungal fermentation. "Beer" is an example of such a fermented beverage, and the term "beer" is meant to comprise any fermented wort produced by fermentation/brewing of a starch-containing plant material. Often, beer is produced exclusively from malt or adjunct, or any combination of malt and adjunct. Examples of beers include: full malted beer, beer brewed under the "Reinheitsgebot," ale, India pale ale, lager, pilsner, bitter, Happoshu (second beer), third beer, dry beer, near beer, light beer, low alcohol beer, low calorie beer, porter, bock, dopplebock, stout, porter, malt liquor, non-alcoholic beer, non-alcoholic malt liquor and the like, but also alternative cereal and malt beverages such as fruit flavored malt beverages, e.g., citrus flavored, such as lemon-, orange-, lime-, or berry-flavored malt beverages, liquor flavored malt beverages, e.g., vodka-, rum-, or tequila-flavored malt liquor, or coffee flavored malt beverages, such as caffeine-flavored malt liquor, and the like.
[0307] The term "malt" refers to any malted cereal grain, such as malted barley or wheat.
[0308] The term "adjunct" refers to any starch and/or sugar containing plant material that is not malt, such as barley or wheat malt. Examples of adjuncts include common corn grits, refined corn grits, brewer's milled yeast, rice, sorghum, refined corn starch, barley, barley starch, dehusked barley, wheat, wheat starch, torrified cereal, cereal flakes, rye, oats, potato, tapioca, cassava and syrups, such as corn syrup, sugar cane syrup, inverted sugar syrup, barley and/or wheat syrups, and the like.
[0309] The term "mash" refers to an aqueous slurry of any starch and/or sugar containing plant material, such as grist, e.g., comprising crushed barley malt, crushed barley, and/or other adjunct or a combination thereof, mixed with water later to be separated into wort and spent grains.
[0310] The term "wort" refers to the unfermented liquor run-off following extracting the grist during mashing.
[0311] "Iodine-positive starch" or "IPS" refers to (1) amylose that is not hydrolyzed after liquefaction and saccharification, or (2) a retrograded starch polymer. When saccharified starch or saccharide liquor is tested with iodine, the high DPn amylose or the retrograded starch polymer binds iodine and produces a characteristic blue color. The saccharide liquor is thus termed "iodine-positive saccharide," "blue saccharide," or "blue sac."
[0312] The terms "retrograded starch" or "starch retrogradation" refer to changes that occur spontaneously in a starch paste or gel on ageing.
[0313] The term "about" refers to .+-.15% to the referenced value.
2. .alpha.-Amylase Variants
[0314] An aspect of the present compositions and methods is variant amylase enzymes that include combinations of mutations that improve their performance in industrial applications. The combinatorial variants were initially discovered using an .alpha.-amylase from Cytophaga sp. (herein, "CspAmy2 amylase"), which was previously described by Jeang, C-L et al. ((2002) Applied and Environmental Microbiology, 68:3651-54). The amino acid sequence of the mature form of the CspAmy2 .alpha.-amylase polypeptide is shown below as SEQ ID NO: 1:
TABLE-US-00001 AATNGTMMQY FEWYVPNDGQ QWNRLRTDAP YLSSVGITAV WTPPAYKGTS QADVGYGPYD LYDLGEFNQK GTVRTKYGTK GELKSAVNTL HSNGIQVYGD VVMNHKAGAD YTENVTAVEV NPSNRNQETS GEYNIQAWTG FNFPGRGTTY SNFKWQWFHF DGTDWDQSRS LSRIFKFRGT GKAWDWEVSS ENGNYDYLMY ADIDYDHPDV VNEMKKWGVW YANEVGLDGY RLDAVKHIKF SFLKDWVDNA RAATGKEMFT VGEYWQNDLG ALNNYLAKVN YNQSLFDAPL HYNFYAASTG GGYYDMRNIL NNTLVASNPT KAVTLVENHD TQPGQSLEST VQPWFKPLAY AFILTRSGGY PSVFYGDMYG TKGTTTREIP ALKSKIEPLL KARKDYAYGT QRDYIDNPDV IGWTREGDST KAKSGLATVI TDGPGGSKRM YVGTSNAGEI WYDLTGNRTD KITIGSDGYA TFPVNGGSVS VWVQQ
[0315] In SEQ ID NO: 1, R178 and G179 are underlined. A variant of the Cytophaga sp. .alpha.-amylase having a deletion of both R178 and G179 (herein, "CspAmy2-v1") has also been described (Shiau, R-J. et al. (2003) Applied and Environmental Microbiology, 69:2383-85). The amino acid sequence of the mature CspAmy2-v1 .alpha.-amylase polypeptide is shown below as SEQ ID NO: 2:
TABLE-US-00002 AATNGTMMQY FEWYVPNDGQ QWNRLRTDAP YLSSVGITAV WTPPAYKGTS QADVGYGPYD LYDLGEFNQK GTVRTKYGTK GELKSAVNTL HSNGIQVYGD VVMNHKAGAD YTENVTAVEV NPSNRNQETS GEYNIQAWTG FNFPGRGTTY SNFKWQWFHF DGTDWDQSRS LSRIFKFTGK AWDWEVSSEN GNYDYLMYAD IDYDHPDVVN EMKKWGVWYA NEVGLDGYRL DAVKHIKFSF LKDWVDNARA ATGKEMFTVG EYWQNDLGAL NNYLAKVNYN QSLFDAPLHY NFYAASTGGG YYDMRNILNN TLVASNPTKA VTLVENHDTQ PGQSLESTVQ PWFKPLAYAF ILTRSGGYPS VFYGDMYGTK GTTTREIPAL KSKIEPLLKA RKDYAYGTQR DYIDNPDVIG WTREGDSTKA KSGLATVITD GPGGSKRMYV GTSNAGEIWY DLTGNRTDKI TIGSDGYATF PVNGGSVSVW VQQ
[0316] Using SEQ ID NO: 2 as a starting point, a number of combinatorial CspAmy2 variants were initially made and tested as described in the Examples section. The best performing variants generally included a stabilizing mutation at an amino acid position corresponding to either E187 or S241, but not both positions, and at least one additional performance-enhancing mutation at amino acid position selected from the group consisting of N126, Y150, F153, L171, T180, and, I203 (using SEQ ID NO: 1 for numbering).
[0317] It is known that many bacterial (and other) .alpha.-amylases share the same fold, often share significant amino acid sequence identity, and sometimes benefit from the same mutations. In the present case, corresponding amino acid positions were identified by amino acid sequence alignment in an .alpha.-amylase from Paenibacillus curdlanolyticus (i.e., PcuAmy1; SEQ ID NO: 3) and a C-terminal-truncated version of the Bacillus sp. TS-23 .alpha.-amylase (i.e., "BASE;" SEQ ID NO: 5; see, e.g., US20120045817 and WO2010/115028).
[0318] The amino acid sequence of the mature form of the PcuAmy1 .alpha.-amylase polypeptide is shown below as SEQ ID NO: 3:
TABLE-US-00003 ADNGTIMQYF EWYLPNDGAH WNRLNNDAQN LKNVGITAVW IPPAYKGGSS ADVGYGVYDT YDLGEFNQKG TVRTKYGTKS ELISAVNNLH AKGIAVYGDV VLNHRMNADA TELVDAVEVD PNNRNVETTS TYQIQAWTQY DFPGRGNTYS SFKWRWYHFD GVDWDQSRGL NRIYKLRGDG KDWDWEVDSE YGNYDYLMGA DLDFNHPDVV NETKTWGKWF VNTVNLDGVR LDAVKHIKFD FMRDWVNNVR STTGKNLFAV GEYWHYDVNK LNSYITKTNG TMSLFDVPLH FRFYDASNGG GGYDMRNLLN NTLMSSNPMK AVTFVENHDT QPTQALQSTV QSWFKPLAYA TILTREQGYP CVFYGDYYGT SDGKISSYKP IMDKLLNARK VYAYGTQRDY FDHPDIVGWT REGDAAHAGS GLATLITDGP GGSKWMYVGT SKAGQVWTDK TGNRSGTVTI DANGWGNFWV NGGSVSVWAK
[0319] In SEQ ID NO: 3, R177 and R178 are underlined. The amino acid sequence of a variant form of PcuAmy1 .alpha.-amylase having a deletion of both R177 and R178 (herein, "PcuAmy1-v1") is shown below as SEQ ID NO: 4:
TABLE-US-00004 ADNGTIMQYF EWYLPNDGAH WNRLNNDAQN LKNVGITAVW IPPAYKGGSS ADVGYGVYDT YDLGEFNQKG TVRTKYGTKS ELISAVNNLH AKGIAVYGDV VLNHRMNADA TELVDAVEVD PNNRNVETTS TYQIQAWTQY DFPGRGNTYS SFKWRWYHFD GVDWDQSRGL NRIYKLDGKD WDWEVDSEYG NYDYLMGADL DFNHPDVVNE TKTWGKWFVN TVNLDGVRLD AVKHIKFDFM RDWVNNVRST TGKNLFAVGE YWHYDVNKLN SYITKTNGTM SLFDVPLHFR FYDASNGGGG YDMRNLLNNT LMSSNPMKAV TFVENHDTQP TQALQSTVQS WFKPLAYATI LTREQGYPCV FYGDYYGTSD GKISSYKPIM DKLLNARKVY AYGTQRDYFD HPDIVGWTRE GDAAHAGSGL ATLITDGPGG SKWMYVGTSK AGQVWTDKTG NRSGTVTIDA NGWGNFWVNG GSVSVWAK
[0320] The amino acid sequence of a C-terminal-truncated version of the Bacillus sp. TS-23 .alpha.-amylase (herein, "BASE;" see, e.g., US20120045817 and WO2010/115028) is shown, below as SEQ ID NO: 5:
TABLE-US-00005 NTAPINETMM QYFEWDLPND GTLWTKVKNE AANLSSLGIT ALWLPPAYKG TSQSDVGYGV YDLYDLGEFN QKGTIRTKYG TKTQYIQAIQ AAKAAGMQVY ADVVFNHKAG ADGTEFVDAV EVDPSNRNQE TSGTYQIQAW TKFDFPGRGN TYSSFKWRWY HFDGTDWDES RKLNRIYKFR STGKAWDWEV DTENGNYDYL MFADLDMDHP EVVTELKNWG TWYVNTTNID GFRLDAVKHI KYSFFPDWLT YVRNQTGKNL FAVGEFWSYD VNKLHNYITK TNGSMSLFDA PLHNNFYTAS KSSGYFDMRY LLNNTLMKDQ PSLAVTLVDN HDTQPGQSLQ SWVEPWFKPL AYAFILTRQE GYPCVFYGDY YGIPKYNIPG LKSKIDPLLI ARRDYAYGTQ RDYIDHQDII GWTREGIDTK PNSGLAALIT DGPGGSKWMY VGKKHAGKVF YDLTGNRSDT VTINADGWGE FKVNGGSVSI WVAK
[0321] In SEQ ID NO: 5, R180 and S181 are underlined. The amino acid sequence of a variant form of BASE .alpha.-amylase having a deletion of both R180 and S181 (herein, "ACE") is shown, below as SEQ ID NO: 6:
TABLE-US-00006 NTAPINETMM QYFEWDLPND GTLWTKVKNE AANLSSLGIT ALWLPPAYKG TSQSDVGYGV YDLYDLGEFN QKGTIRTKYG TKTQYIQAIQ AAKAAGMQVY ADVVFNHKAG ADGTEFVDAV EVDPSNRNQE TSGTYQIQAW TKFDFPGRGN TYSSFKWRWY HFDGTDWDES RKLNRIYKFT GKAWDWEVDT ENGNYDYLMF ADLDMDHPEV VTELKNWGTW YVNTTNIDGF RLDAVKHIKY SFFPDWLTYV RNQTGKNLFA VGEFWSYDVN KLHNYITKTN GSMSLFDAPL HNNFYTASKS SGYFDMRYLL NNTLMKDQPS LAVTLVDNHD TQPGQSLQSW VEPWFKPLAY AFILTRQEGY PCVFYGDYYG IPKYNIPGLK SKIDPLLIAR RDYAYGTQRD YIDHQDIIGW TREGIDTKPN SGLAALITDG PGGSKWMYVG KKHAGKVFYD LTGNRSDTVT INADGWGEFK VNGGSVSIWV AK
[0322] An amino acid sequence alignment of CspAmy2 (SEQ ID NO: 1), PcuAmy1 (SEQ ID NO: 3), and BASE (SEQ ID NO: 5), using Clustal W with default parameters, is shown in FIG. 1. Positions N126, Y150, F153, L171, R178, G179, T180, E187, 1203, and S241 in CspAmy2 correspond to positions N125, Y149, F152, L170, R177, G178, D179, E186, L202, and D240 in PcuAmy1, respectively, and positions N128, Y152, F155, L173, R180, S181, T182, E189, L205, and S243, respectively in BASE. Numbering for other positions through out the molecules can be determined using the alignment and information, herein.
[0323] Based on experimental data obtained using the aforementioned three parent .alpha.-amylases, embodiments of the present variant .alpha.-amylases include variants having a mutation at an amino acid position corresponding to E187 or S241 in combination with at least one mutation at an amino acid position corresponding to a position selected from N126, Y150, F153, L171, T180, and I203 (using SEQ ID NO: 1 for numbering), wherein the mutations provide at least one performance benefit to the resulting variant.
[0324] Referring to SEQ ID NO: 1 for numbering, exemplary mutations at amino acid position E187 include E187V and E187P. Exemplary mutations at amino acid position S241 include S241Q and S241A. In some embodiments, mutations are made in only one of these positions. Exemplary mutations at amino acid position N126 includes N126Y. Exemplary mutations at amino acid position Y150 include Y150F, Y150H, and Y150W. Exemplary mutations at amino acid position F153 include F153H, F153W, and F153Y. Exemplary mutations at amino acid position L171 include L171F, L171G, L171I, L171M, L171R, L171V, L171W, L171Y, L171H, L171K, L171N, L171Q, and L171S. Exemplary mutations at amino acid position T180 include T180D and T180H. Exemplary mutations at amino acid position I203 includes I203C, I203V, I203F, I203L, I203M, and I203Y.
[0325] In some embodiments, the variant .alpha.-amylases further include a mutation in amino acid residues corresponding to E132, Q167, A277, and/or T400, using SEQ ID NO: 1 for numbering. Exemplary mutations at amino acid position E132 include E132A, E132C, E132D, E132F, E132G, E132H, E132I, E132K, E132L, E132M, E132N, E132P, E132Q, E132R, E132V, and E132W. Exemplary mutations at amino acid position Q167 include Q167A, Q167D, Q167E, Q167G, Q167H, Q167K, Q167M, Q167N, Q167P, Q167S, Q167T, and Q167V. Exemplary mutations at amino acid position A277 include A277C, A277D, A277E, A277F, A277G, A277I, A277K, A277L, A277M, A277N, A277Q, A277R, A277S, A277T, A277V, A277W, and A277Y. Exemplary mutations at amino acid position T400 include T400A, T400C, T400D, T400F, T400G, T400I, T400K, T400L, T400M, T400N, T400Q, T400R, T400W, and T400Y.
[0326] In some embodiments, the variant .alpha.-amylases further include a mutation in an amino acid residue corresponding to G476, using SEQ ID NO: 1 for numbering. Exemplary mutations at amino acid position G476 include G476A, G476C, G476H, G476K, G476N, G476P, G476Q, G476R, G476S, G476T, G476V, and G476Y.
[0327] In some embodiments, the variant .alpha.-amylases are those that include, or further include, mutations in both amino acid residues corresponding to G476 and G477, using SEQ ID NO: 1 for numbering. Surprisingly, experimental evidence suggests that any combination of residues at these positions, other than two adjacent glycines as are present in many naturally occurring .alpha.-amylases, increases improve starch hydrolysis activity, particularly in cleaning applications.
[0328] In some embodiments, the variant .alpha.-amylases further include mutations in amino acid residues corresponding to R458, T459, and/or D460. Exemplary mutations are R458N, T459S, and D460T, respectively.
[0329] In some embodiments, the variant .alpha.-amylases further include a deletion in the X.sub.1G/S.sub.1X.sub.2G.sub.2 motif adjacent to the calcium-binding loop corresponding to R178, G179, T180, and G181, using SEQ ID NO: 1 for numbering. In some embodiments, the variant .alpha.-amylases include adjacent, pair-wise deletions of amino acid residues corresponding to R178 and G179, or T180 and G181. A deletion in amino acid residues corresponding to R178 and G179 may be referred to as ".DELTA.RG," while a deletion in amino acid residues corresponding to T180 and G181 ".DELTA.TG." This nomenclature will naturally change depending on the amino acid residues originally present in the parent molecule.
[0330] In some embodiments, the variant .alpha.-amylases include mutations at positions corresponding to E132 and/or T180 (using SEQ ID NO: 1 for numbering), in combination with an RG-deletion or a TG-deletion (or equivalent deletion based on the sequence of the parent .alpha.-amylases), such that a stabilizing interaction can occur between the remaining non-G residue in the X.sub.1G/S.sub.1X.sub.2G.sub.2 motif and the residue at position 132. In some embodiments, the residue at position 132 is negatively charged (i.e., D or E) and the remaining non-G residue is positively charged (i.e., H, R, or K). In some embodiments, the residue at position 132 is positively charged (i.e., H, R, or K) and the remaining non-G residue is negatively charged (i.e., D or E).
[0331] Exemplary combinations of mutation (using SEQ ID NO: 1 for numbering) are shown, below:
E187P+I203Y+G476K (i.e., CspAmy2-v5);
E187P+I203Y+G476K+R458N+T459S+D460T (i.e., CspAmy2-v6);
[0332] T180D+E187P+I203Y+G476K (i.e., CspAmy2 v171); N126Y+T180D+E187P+I203Y+G476K (i.e., CspAmy2 v172); N126Y+T180D+E187P+I203Y+Y303D+G476T+G477E, (i.e., CspAmy2 v179); N126Y+T180D+E187P+I203Y+Y303D+N475E+G477Q, (i.e., CspAmy2 v180); N126Y+T180D+E187P+I203Y+Y303R+N475E+G476T+G477R, (i.e., CspAmy2 v181); T038N+N88H+N126Y+T129I+N134M+F153W+L171R+T180D+E187P+I203Y+G476K+G477E, (i.e., CspAmy2 v186); N126Y+E132H+T180D+E187P+I203Y+Y303D+G476T+G477E, (i.e., CspAmy2 v191);
N126Y+E187P+I203Y (i.e., CspAmy2-vC16A);
N126Y+I203Y+S241Q (i.e., CspAmy2-vC16B);
N126Y+T180H+E187P+I203Y (i.e., CspAmy2-vC16C);
N126Y+T180H+I203Y+S241Q (i.e., CspAmy2-vC16D);
N126Y+F153W+T180H+E187P+I203Y (i.e., CspAmy2-vC16E);
N126Y+F153W+T180H+I203Y+S241Q (i.e., CspAmy2-vC16F);
N126Y+Y150H+F153W+L171N+E187P+I203Y (i.e., CspAmy2-vC16G);
N126Y+Y150H+F153W+L171N+I203Y+S241Q (i.e., CspAmy2-vC16H);
N126Y+Y150H+F153W+L171N+T180H+E187P+I203Y (i.e., CspAmy2-vC16I);
N126Y+Y150H+F153W+L171N+T180H+I203Y+S241Q (i.e., CspAmy2-vC16J);
N126Y+F153W+T180D+I203Y+S241Q, (i.e., CspAmy2-v C18P);
N126Y+E132H+F153W+T180D+I203Y+S241Q+A277F (i.e., CspAmy2-C25F);
N126Y+E132H+F153W+Q167E+T180D+I203Y+S241Q+A277F (i.e., CspAmy2-C25B); and
N126Y+E132H+F153W+Q167E+T180D+I203Y+S241Q+A277F+T400K (i.e., CspAmy2-C25A).
[0333] All the above combinations of mutation are contemplated for use in conjunction with the aforementioned deletions at positions corresponding to R178, G179, T180, and/or G181. Such deletions may be naturally occurring, as in the case of Bacillus licheniformis .alpha.-amylase.
[0334] In addition to the aforementioned mutations, the PcuAmy1 variants may further include mutations at position T333, A335, and Q337E (using SEQ ID NO: 3 numbering). These positions are in a surface-exposed loop and mutations at these positions, particularly at T333, impart protease resistance to PcuAmy1 but do not otherwise affect performance. These mutations also appear to be fully compatible with other mutations. A further mutation in PcuAmy1 variants is N205, which mutation changes the wild-type N residue to D. D is the residue that typically occupies this position in .alpha.-amylases.
[0335] Accordingly, the present .alpha.-amylases include the all the exemplary combinations of mutations shown above in the context of CspAmy2, as well as the following exemplary combinations (using SEQ ID NO: 3 numbering) are shown, below:
N125Y+E186P+T333G+A335S+Q337E+G472K (i.e., PcuAmy1-v1A);
N125Y+F152W+E186P+T333G+A335S+Q337E+G472K (i.e., PcuAmy1-v6);
N125Y+F152W+E186P+T333G+A335S+Q337E+G472R+G473R (i.e., PcuAmy1-v8); and
[0336] N125Y+F152W+E186P+N205D+T333G+A335S+Q337E+G472K (i.e., PcuAmy1-v16).
[0337] All the above combinations of mutations are contemplated for use in conjunction with deletions at positions corresponding to R177, G178, D179, and/or G180, and such deletions may be naturally occurring, as in the case of Bacillus licheniformis .alpha.-amylase.
[0338] The present variants can also be described in terms of BASE numbering (i.e., SEQ ID NO: 5), for example,
N128Y+E189P+G475R (i.e., BASE-V28);
F155W+E189P+G475R (i.e., BASE-V29);
T134E+T182H+E189P+G475R (i.e., BASE-V30);
N128Y+T134E+T182H+E189P+G475R (i.e., BASE-V31);
N128Y+F155W+E189P+G475R (i.e., BASE-V32);
T134E+F155W+T182H+E189P+G475R (i.e., BASE-V33);
N128Y+T134E+F155W+T182H+E189P+G475R (i.e., BASE-V34);
N128Y+T134H+F155W+T182D+E189P+G475R (i.e., BASE-V35); and
N128Y+T134E+F155W+T182G+E189P+G457R (i.e., BASE-V36).
[0339] All the above combinations of mutations are contemplated for use in conjunction with deletions at positions corresponding to R180, S181, T182, and/or G183, and such deletions may be naturally occurring, as in the case of Bacillus licheniformis .alpha.-amylase.
[0340] Corresponding amino acid positions in other .alpha.-amylases be identified by amino acid sequence alignment using CspAmy2 (SEQ ID NO: 1), PcuAmy1 (SEQ ID NO: 3), or BASE (SEQ ID NO: 5), using Clustal W with default parameters. .alpha.-amylases in which the foregoing mutations are likely to produce a performance benefit include those having a similar fold and/or having 60% or greater amino acid sequence identity to any of the well-known Bacillus amylases (e.g., from B. lichenifomis (i.e., BLA and LAT), B. stearothermophilus (i.e., BSG), and B. amyloliquifaciens (i.e., P00692, BACAM, and BAA)), Carbohydrate-Active Enzymes database (CAZy) Family 13 amylases, or any amylase that has heretofore been referred to by the descriptive term, "Termamyl-like." Exemplary .alpha.-amylases include but are not limited to those from Bacillus sp. SG-1, Bacillus sp. 707, Bacillus sp. DSM12368 (i.e., A7-7), Bacillus sp. DSM 12649 (i.e., AA560), Bacillus sp. SP722, Bacillus megaterium (DSM90 14), and KSM AP1378.
[0341] The reader will appreciate that where an .alpha.-amylase naturally has a mutation listed above (i.e., where the wild-type .alpha.-amylase already comprised a residue identified as a mutation), then that particular mutation does not apply to that .alpha.-amylase. However, other described mutations may work in combination with the naturally occurring residue at that position.
[0342] In some embodiments, the present .alpha.-amylase variants have the indicated combinations of mutations and a defined degree of amino acid sequence homology/identity to SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5, for example, at least 60%, at least 65%, at least 70%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or even at least 99% amino acid sequence homology/identity.
[0343] In some embodiments, the present .alpha.-amylase variants have the indicated combinations of mutations and are derived from a parental amylase having a defined degree of amino acid sequence homology/identity to SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5, for example, at least 60%, at least 65%, at least 70%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or even at least 99% amino acid sequence homology/identity.
[0344] In some embodiments, in addition to the mutations described above, the present .alpha.-amylases further include one or more mutations that provide a further performance benefit. Additional mutations that were experimentally determined to provide at least one performance advantage when combined with the aforementioned combinatorial variants include mutations at positions corresponding to 6, 7, 8, 11, 14, 15, 20, 21, 23, 26, 27, 28, 37, 38, 39, 40, 42, 45, 46, 48, 49, 50, 51, 52, 53, 54, 58, 61, 62, 68, 70, 71, 72, 73, 79, 80, 81, 82, 84, 85, 87, 88, 89, 92, 93, 94, 95, 96, 97, 98, 101, 108, 111, 112, 113, 114, 115, 116, 117, 118, 120, 122, 123, 124, 126, 127, 129, 130, 131, 132, 133, 134, 136, 137, 138, 140, 142, 143, 144, 147, 148, 149, 150, 151, 152, 153, 154, 155, 156, 158, 159, 165, 167, 168, 170, 171, 172, 175, 176, 177, 180, 181, 182, 187, 190, 191, 193, 199, 200, 201, 203, 206, 208, 210, 211, 212, 214, 215, 216, 219, 221, 223, 225, 226, 227, 235, 238, 239, 240, 241, 242, 243, 245, 246, 247, 248, 249, 250, 252, 253, 254, 256, 257, 258, 260, 261, 262, 266, 267, 268, 269, 270, 271, 273, 276, 277, 279, 280, 282, 284, 285, 286, 288, 296, 299, 300, 301, 302, 303, 304, 307, 308, 310, 311, 312, 313, 316, 317, 318, 320, 321, 325, 327, 335, 338, 342, 348, 349, 352, 356, 357, 360, 362, 363, 368, 369, 377, 381, 382, 383, 384, 385, 388, 390, 392, 394, 395, 396, 397, 398, 400, 401, 402, 403, 404, 405, 407, 408, 410, 414, 415, 416, 418, 419, 420, 421, 422, 423, 424, 426, 428, 429, 430, 431, 434, 435, 436, 439, 441, 442, 444, 445, 446, 447, 448, 449, 450, 451, 454, 455, 457, 460, 461, 462, 463, 464, 465, 466, 467, 469, 470, 471, 473, 474, 475, 476, 477, 479, 480, 481, 482, 483, and 484, using SEQ ID NO: 1 for numbering. Specific mutations are T6A, T6D, T6E, T6G, T6K, T6M, T6N, T6Q, T6S, M7A, M7V, MBC, MBF, M8I, M8L, M8Y, F11V, Y14A, Y14I, Y14Q, Y14T, Y14V, V15C, V15D, V15I, V15N, V15T, Q20A, Q20C, Q20D, Q20H, Q20K, Q20M, Q20N, Q20R, Q205, Q20Y, Q21F, Q21W, N23A, N23C, N23D, N23E, N23H, N23K, N23Q, N23R, N23S, N23T, N23V, R26C, R26E, R26G, R26K, R26M, R26S, R26T, T27A, T27C, T27D, T27E, T27F, T27H, T27I, T27K, T27L, T27M, T27N, T27Q, T27R, T27S, T27V, T27Y, D28A, D28C, D28T, I37F, I37V, T38D, T38N, A39L, V40A, V40C, V40D, V40G, V40H, V40I, V40K, V40M, V40P, V40Q, V40R, V405, V40T, V40W, V40Y, T42A, T42C, T42I, T42M, T42V, A45C, A45G, Y46F, G48A, T49I, S50A, S50C, S50E, S50G, S50K, S50M, S50N, S50Q, S50R, S50T, S50Y, Q51C, Q51D, Q51E, Q51S, Q51V, A52C, A52D, A52E, A52F, A52G, A52H, A52K, A52L, A52M, A52R, A52S, A52T, A52V, A52Y, D53E, V54N, V54T, P58A, P58C, P58H, P58I, P58S, P58T, P58V, L61M, L61V, Y62A, Y62C, Y62D, Y62F, Y62G, Y62H, Y62I, Y62K, Y62L, Y62M, Y62N, Y62P, Y62Q, Y62R, Y62S, Y62V, N68C, N68D, N68E, N68F, N68H, N68L, N68P, N68Q, N68R, N68S, N68V, N68W, N68Y, K70R, G71A, G71C, G71D, G71E, G71K, G71R, G71S, T72G, T72S, V73S, T79F, T791, T79L, T79M, T79N, T79S, T79Y, K80A, K80C, K80D, K80F, K80H, K80I, K80M, K80N, K80Q, K80R, K80S, K80T, K80V, K80Y, G81A, G81D, G81E, G81F, G81H, G81I, G81K, G81N, G81P, G81R, G81S, G81T, E82A, E82D, E82M, E82Q, K84A, K84C, K84E, K84I, K84Q, K84R, K84S, K84T, K84Y, S85A, S85C, S85D, S85E, S85G, S85H, S851, S85L, S85M, S85N, S85Q, S85R, S85T, S85V, S85Y, V871, V87T, N88C, N88D, N88E, N88G, N88H, N88I, N88K, N88L, N88M, N88Q, N88R, N88S, N88T, N88V, N88W, N88Y, T89A, T89D, T89E, T89F, T89H, T891, T89K, T89L, T89M, T89N, T89Q, T89R, T89S, T89Y, S92A, S92C, S92D, S92E, S92F, S92G, S92H, S92L, S92M, S92N, S92Q, S92R, S92T, S92W, S92Y, N93A, N93C, N93E, N93F, N93H, N93I, N93K, N93L, N93Q, N93S, N93T, N93Y, G94A, G94C, G94N, I95M, Q96A, Q96E, Q96H, Q961, Q96K, Q96L, Q96M, Q96N, Q96R, Q96V, Q96Y, V971, V97T, Y98F, Y981, Y98L, Y98V, V101C, V101T, G108A, G108S, Y111D, Y111E, Y111L, Y111N, Y111S, Y111T, Y111V, T112A, T112F, T112H, T112I, T112K, T112L, T112M, T112N, T112P, T112R, T112V, T112W, E113D, E113N, E113Q, E113T, N114A, N114C, N114D, N114E, N114F, N114G, N114H, N114I, N114L, N114P, N114Q, N114R, N114S, N114T, N114V, N114W, N114Y, V115A, V115I, T116A, T116C, T116D, T116G, T116H, T116I, T116K, T116N, T116P, T116Q, T116R, T116S, A117C, A117I, A117S, A117V, V118A, V118C, V118E, V118F, V118H, V118I, V118L, V118M, V118N, V118Q, V118R, V118S, V118W, V118Y, V120A, V120C, V120I, V120M, V120T, P122A, P122D, P122E, P122G, P122H, P122N, P122R, P122S, P122T, P122W, S123A, S123C, S123G, S123K, S123L, N124A, N124D, N124F, N124G, N124L, N124R, N124S, N124V, Y126A, Y126C, Y126D, Y126E, Y126G, Y126H, Y126I, Y126K, Y126L, Y126M, Y126N, Y126Q, Y126R, Y126S, Y126T, Y126V, Y126W, Q127A, Q127C, Q127D, Q127E, Q127F, Q127H, Q127I, Q127K, Q127L, Q127M, Q127N, Q127R, Q127S, Q127T, Q127V, Q127W, Q127Y, T129A, T129C, T129D, T129E, T129F, T129G, T129I, T129K, T129L, T129M, T129N, T129Q, T129R, T129S, T129V, T129W, T129Y, S130A, S130G, S130I, S130K, S130L, S130M, S130N, S130P, S130R, S130T, S130V, S130W, G131A, G131C, G131D, G131F, G131H, G131I, G131K, G131L, G131M, G131P, G131Q, G131V, G131W, G131Y, E132A, E132C, E132D, E132F, E132G, E132H, E132I, E132K, E132L, E132M, E132N, E132P, E132Q, E132R, E132V, E132W, Y133A, Y133D, Y133E, Y133G, Y133K, Y133L, Y133M, Y133R, Y133S, Y133T, Y133W, N134A, N134D, N134F, N134G, N134H, N134I, N134K, N134L, N134M, N134R, N134S, N134V, N134W, N134Y, Q136A, Q136C, Q136D, Q136E, Q136F, Q136H, Q136K, Q136L, Q136M, Q136N, Q136R, Q136S, Q136T, Q136V, Q136W, Q136Y, A137S, A137T, A137V, W138F, W138Y, G140A, G140M, N142F, N142K, N142L, N142M, N142P, N142V, N142W, N142Y, F143H, F143Y, P144C, P144D, P144E, P144G, P144H, P1441, P144K, P144L, P144M, P144N, P144Q, P144S, P144T, P144Y, G147A, G147C, G147H, G147K, G147L, G147M, G147N, G147Q, G147R, T148A, T148D, T148E, T148F, T1481, T148K, T148L, T148R, T148S, T148V, T148W, T149A, T149C, T149D, T149E, T149F, T149H, T149K, T149L, T149M, T149N, T149Q, T149V, T149W, T149Y, Y150H, S151A, S151E, S151F, S151H, S1511, S151K, S151L, S151M, S151Q, S151R, S151V, N152A, N152C, N152D, N152E, N152G, N152H, N152K, N152M, N152P, N152Q, N152R, N152S, N152T, W153F, W153H, W153Q, W153R, W153T, W153Y, K154A, K154C, K154D, K154E, K154G, K1541, K154L, K154M, K154N, K154R, K154T, K154Y, W155P, Q156A, Q156D, Q156E, Q156F, Q156G, Q156H, Q1561, Q156L, Q156M, Q156N, Q156R, Q156S, Q156Y, F158C, F158D, F158K, F158L, F158M, F158N, F158P, F158Q, F158R, F158S, F158V, H159M, H159Y, W165C, W165D, W165E, W165F, W165H, W1651, W165K, W165L, W165M, W165Q, W165R, W165T, W165Y, Q167A, Q167D, Q167E, Q167G, Q167H, Q167K, Q167M, Q167N, Q167P, Q167S, Q167T, Q167V, S168A, S168D, S168F, S168H, S1681, S168K, S168M, S168N, S168Q, S168R, S168T, S168V, S168W, S168Y, S170A, S170C, S170D, S170E, S170F, S170L, S170M, S170N, S170Q, S170R, S170T, L171A, L171C, L171F, L171G, L171H, L171I, L171N, L171Q, L171R, L171T, L171V, L171W, L171Y, S172A, S172D, S172H, S172R, S172T, F175M, F175Y, K176I, K176T, F177L, F177V, F177W, D180A, D180C, D180F, D180G, D180H, D180I, D180L, D180M, D180N, D180Q, D180R, D180S, D180T, D180V, D180W, D180Y, G181A, G181C, G181D, G181E, G181F, G181H, G181K, G181L, G181M, G181N, G181Q, G181R, G181S, G181T, G181V, G181Y, K182A, K182P, E187D, E187I, E187K, E187M, E187N, E187P, E187R, E187S, E187T, E187V, E187Y, S190A, S190C, S190D, S190F, S190L, S190N, S190P, S190Q, E191A, E191C, E191F, E191G, E191H, E191I, E191K, E191N, E191Q, E191R, E191S, E191T, E191W, E191Y, G193A, G193C, G193D, G193E, G193F, G193H, G193K, G193L, G193M, G193N, G193Q, G193R, G193S, G193T, G193V, G193W, M199L, Y200F, A201M, Y2031, Y203L, Y203V, D206A, D206E, D206G, D206H, D206M, D206N, D206Q, D206R, D206S, D206T, P208A, P208E, P208F, P2081, P208K, P208L, P208T, P208V, P208Y, V210A, V210E, V210H, V210K, V210N, V210Q, V210R, V210S, V210T, V211A, V211E, V211H, V2111, V211Q, V211R, N212E, N212F, N212G, N212L, N212M, N212R, N212V, M2141, M214L, K215C, K215E, K215F, K215M, K215N, K215R, K215Y, K216F, V2191, V219T, Y221F, Y2211, Y221L, N223C, N223E, N223I, N223K, N223Q, N223R, N223S, N223T, N223V, N223W, N223Y, V225A, V2251, V225L, V225M, G226D, G226M, G226Q, G226R, G226S, L227F, L227I, L227W, L227Y, V235A, I238A, I238L, I238M, K239D, K239E, K239P, K239Q, K239R, K239S, K239T, F240K, F240L, F240M, F240Q, F240R, Q241A, Q241C, Q241D, Q241E, Q241G, Q241H, Q241K, Q241L, Q241M, Q241N, Q241P, Q241R, Q241S, Q241T, Q241V, Q241W, Q241Y, F242V, L243C, L243Y, D245A, D245C, D245E, D245G, D245L, D245M, D245N, W246F, V247I, V247L, D248E, D248H, D248N, D248T, D248V, N249A, N249E, N249G, N249H, N249Q, N249Y, A250M, A250S, A250V, A252C, A252D, A252E, A252G, A252H, A252I, A252K, A252L, A252M, A252N, A252Q, A252S, A252V, A252W, A252Y, A253E, A253I, A253K, A253L, A253M, A253Q, A253S, A253T, A253V, A253Y, T254F, T254K, T254S, K256A, K256M, K256N, K256S, E257Q, E257S, M258L, T260A, T260C, T260S, T260V, V261I, V261W, G262A, Q266A, Q266D, Q266E, Q266H, Q266I, Q266M, Q266N, Q266S, Q266T, Q266V, Q266Y, N267H, N267I, N267Q, N267R, N267S, N267T, N267V, N267Y, D268G, D268N, L269C, L269D, L269I, L269K, L269Q, L269S, L269T, L269Y, G270A, G270D, G270E, G270F, G270H, G270I, G270L, G270M, G270Q, G270T, G270V, G270Y, A271C, A271D, A271E, A271H, A271K, A271M, A271Q, A271R, A271S, A271T, A271V, A271Y, N273C, N273G, N273H, N273I, N273K, N273R, L276I, L276M, A277C, A277D, A277E, A277F, A277G, A277I, A277K, A277L, A277M, A277N, A277Q, A277R, A277S, A277T, A277V, A277W, A277Y, V279C, V279T, N280A, N282S, N282T, S284T, S284Y, L285A, L285C, L285I, L285V, F286M, A288C, A296C, A296D, A296E, A296F, A296G, A296H, A296I, A296L, A296M, A296N, A296Q, A296R, A296S, A296V, A296W, A296Y, T299A, T299D, T299E, T299F, T299G, T299K, T299L, T299M, T299R, T299S, T299V, T299W, G300A, G300C, G300D, G300E, G300F, G300H, G300K, G300M, G300Q, G300R, G300V, G300W, G301A, G301C, G301D, G301E, G301F, G301H, G301K, G301L, G301M, G301Q, G301R, G301S, G301T, G301V, G301W, G301Y, G302S, Y303A, Y303C, Y303D, Y303E, Y303F, Y303G, Y303H, Y3031, Y303K, Y303L, Y303M, Y303N, Y303Q, Y303R, Y303S, Y303T, Y303V, Y303W, Y304F, Y304K, Y304W, R307A, R307C, R307E, R307G, R307H, R307K, R307M, R307N, R307Q, R307S, R307T, N308C, N308D, N308E, N308G, N308L, N308M, N308T, N308V, L310A, L310C, L310D, L310E, L310H, L310I, L310M, L310P, L310W, L310Y, N311C, N311E, N311G, N311H, N311K, N311Q, N311R, N311S, N311V, N311W, N311Y, N312D, N312F, N312G, N312H, N312K, N312Q, N312R, T313A, T313S, A316D, A316E, A316G, A316H, A316K, A316Q, A316R, A316Y, S317C, S317D, S317G, S317H, S317K, S317L, S317N, S317Q, S317R, S317T, S317W, S317Y, N318A, N318C, N318F, N318G, N318I, N318K, N318L, N318M, N318Q, N318R, N318S, N318T, N318V, N318W, T320A, T320C, T320D, T320E, T320G, T320H, T3201, T320K, T320N, T320P, T320Q, T320R, T320V, T320W, T320Y, K321C, K321F, K321H, K321N, K321S, K321Y, L325A, L325C, L325F, L3251, L325M, L325Q, L325V, E327D, E327L, Q335C, Q335E, E338A, E338D, E338F, E338G, E338H, E3381, E338K, E338P, E338Q, E338R, E338T, E338V, E338Y, Q342A, Q342C, Q342G, Q342L, Q342M, Q342R, Q342S, Q342T, Q342V, Q342W, L348A, L348C, L348H, L3481, L348M, L348Q, L348S, L348T, A349G, A349R, F3521, F352L, F352M, F352T, F352V, R356Q, S357A, S357C, S357D, S357E, S357F, S357H, S3571, S357K, S357L, S357N, S357Q, S357T, S357V, S357W, S357Y, Y360F, Y3601, Y360L, Y360M, Y360V, S362A, S362C, S362E, S3621, S362T, S362V, V3631, V363L, M368F, M3681, M368L, M368Y, Y369A, Y369E, Y3691, Y369L, Y369N, Y369V, R377A, R377C, R377D, R377E, R377F, R377G, R377H, R3771, R377K, R377L, R377N, R377Q, R377S, R377T, R377V, R377W, R377Y, A381C, A381E, A381G, A381H, A381K, A381L, A381M, A381N, A381P, A381Q, A381V, A381W, A381Y, L382F, L382H, L382Q, L382S, K383A, K383C, K383D, K383E, K383H, K3831, K383L, K383M, K383N, K383Q, K383R, K383S, K383W, K383Y, S384A, S384C, S384D, S384F, S384G, S384H, S3841, S384L, S384N, S384R, S384V, S384Y, K385A, K385D, K385E, K385F, K385G, K385H, K385L, K385M, K385Q, K385R, K385T, K385V, K385Y, P388A, P388C, P388D, P388I, P388L, P388N, P388R, P388S, P388T, P388V, L390M, L390V, A392C, A392S, K394A, K394C, K394E, K394F, K394G, K394H, K3941, K394L, K394Q, K394R, K394S, K394T, K394V, K394W, K394Y, D395A, D395E, D395N, D395S, D395T, Y396A, Y396D, Y396F, Y396K, Y396M, Y396N, Y396Q, Y396T, Y396V, Y396W, A397D, A397E, A397H, A397K, A397M, A397N, A397Q, A397V, Y398A, Y398C, Y398F, Y398H, Y3981, Y398L, Y398W, T400A, T400C, T400D, T400F, T400G, T400I, T400K, T400L, T400M, T400N, T400Q, T400R, T400W, T400Y, Q401A, Q401C, Q401F, Q401H, Q401I, Q401L, Q401M, Q401N, R402C, R402D, R402F, R402K, R402L, R402M, R402N, R402Q, R402S, R402W, D403E, D403N, D403S, Y404E, Y404G, Y404K, Y404M, Y404N, Y404R, Y404W, I405C, I405F, I405L, I405M, I405V, N407A, N407C, N407D, N407E, N407G, N407K, N407M, N407Q, N407R, P408A, P408C, P408D, P4081, P408K, P408L, P408M, P408N, P408Q, P408V, P408W, V410A, V410C, V410D, V410E, V410F, V410H, V4101, V410L, V410M, V410N, V410Q, V410S, V410T, T414A, T4141, T414S, T414V, R415M, E416C, E416D, E416F, E416H, E4161, E416K, E416L, E416M, E416N, E416Q, E416R, E416T, E416V, E416W, E416Y, D418A, D418E, D418G, D418H, D418I, D418K, D418L, D418M, D418N, D418Q, D418S, D418T, D418V, D418W, S419A, S419C, S419E, S419G, S419L, S419M, S419N, S419R, S419V, S419W, S419Y, T420A, T420C, T420D, T420E, T420G, T420H, T420I, T420K, T420M, T420P, T420S, T420V, T420W, T420Y, K421A, K421D, K421E, K421H, K421I, K421L, K421M, K421N, K421P, K421Q, K421R, K421T, K421V, K421W, K421Y, A422C, A422D, A422E, A422F, A422G, A422I, A422L, A422N, A422P, A422Q, A422R, A422S, A422Y, K423A, K423D, K423E, K423F, K423H, K423I, K423L, K423M, K423N, K423Q, K423R, K423S, K423T, K423V, K423W, K423Y, S424A, S424C, S424G, S424K, S424N, S424Q, S424R, S424T, L426S, L426T, L426V, T428G, T428V, V429A, V429C, V429I, V429L, I430C, I430G, I430L, I430M, I430Q, I430V, T431A, T431C, T431S, P434A, P434C, P434D, P434E, P434F, P434H, P434I, P434K, P434L, P434M, P434N, P434Q, P434R, P434S, P434V, P434Y, G435A, G435C, G435D, G435E, G435F, G435H, G435I, G435K, G435M, G435N, G435P, G435Q, G435R, G435S, G435T, G435W, G436F, G436I, G436M, G436N, G436Q, G436S, G436V, R439A, R439D, R439G, R439H, R439K, R439M, R439N, R439P, R439Q, R439S, R439V, R439W, R439Y, Y441A, Y441C, Y441D, Y441F, Y441G, Y441H, Y441K, Y441L, Y441M, Y441N, Y441P, Y441R, Y441S, Y441T, Y441W, V442A, V442C, V442I, V442T, T444C, T444D, T444E, T444F, T444G, T444H, T444I, T444K, T444L, T444M, T444N, T444P, T444R, T444S, T444W, S445A, S445C, S445E, S445G, S445H, S445K, S445L, S445M, S445N, S445T, S445V, N446A, N446C, N446H, N446K, A447C, A447D, A447F, A447H, A447L, A447M, A447N, A447Q, A447R, A447S, A447Y, G448A, G448C, G448D, G448E, G448H, G448K, G448L, G448M, G448N, G448Q, G448R, G448S, G448T, G448W, E449D, E449H, E449K, E449T, I450A, I450C, I450D, I450E, I450G, I450K, I450L, I450M, I450N, I450Q, I450S, I450T, I450W, I450Y, W451Y, L454A, L454I, L454K, L454M, L454W, T455A, T455I, T455L, T455S, N457H, N457K, N457R, N457T, N457V, N457Y, D460A, D460E, D460G, D460M, D460N, D460Q, D460S, D460V, K461C, K461H, K461L, K461M, K461N, K461Q, K461T, K461Y, I462A, I462L, I462M, I462Q, I462T, I462V, T463D, T463E, T463H, T463P, T463Q, T463R, T463V, T463Y, I464P, I464T, G465A, G465C, G465D, G465E, G465K, G465L, G465M, G465N, G465Q, G465W, G465Y, S466A, S466C, S466D, S466G, S466H, S466K, S466L, S466M, S466N, S466T, S466W, S466Y, D467E, D467G, D467L, Y469A, Y469D, Y469E, Y469I, Y469M, Y469N, Y469R, Y469S, Y469T, Y469V, Y469W, A470S, A470V, T471A, T471C, T471E, T471G, T471H, T471I, T471L, T471M, T471N, T471S, T471V, T471W, P473C, P473D, P473E, P473G, P473I, P473K, P473L, P473R, P473T, P473W, V474A, V474C, V474L, V474S, N475A, N475C, N475E, N475F, N475H, N475K, N475L, N475M, N475P, N475Q, N475R, N475S, N475T, N475V, G476A, G476D, G476E, G476F, G476H, G4761, G476L, G476M, G476N, G476P, G476Q, G476R, G476S, G476T, G476V, G476W, G476Y, G477A, G477D, G477F, G477H, G4771, G477K, G477L, G477M, G477Q, G477S, G477T, G477V, G477W, G477Y, V479C, V479D, V479E, V479F, V479H, V4791, V479N, V479P, V479Y, S480A, S480C, S480H, V481A, V481C, V481N, W482Y, V483A, V483G, V4831, V483K, V483L, V483M, V483R, V483Y, Q484A, Q484C, Q484F, Q484G, Q484H, Q484K, Q484L, Q484M, Q484P, Q484R, Q484T, and Q484Y.
[0345] Furthermore, the present amylases may include any number of conservative amino acid substitutions. Exemplary conservative amino acid substitutions are listed in Table 1 Table 1. Conservative amino acid substitutions
TABLE-US-00007 For Amino Acid Code Replace with any of Alanine A D-Ala, Gly, beta-Ala, L-Cys, D-Cys Arginine R D-Arg, Lys, D-Lys, homo-Arg, D-homo-Arg, Met, Ile, D-Met, D-Ile, Orn, D-Orn Asparagine N D-Asn, Asp, D-Asp, Glu, D-Glu, Gln, D-Gln Aspartic Acid D D-Asp, D-Asn, Asn, Glu, D-Glu, Gln, D-Gln Cysteine C D-Cys, S-Me-Cys, Met, D-Met, Thr, D-Thr Glutamine Q D-Gln, Asn, D-Asn, Glu, D-Glu, Asp, D-Asp Glutamic Acid E D-Glu, D-Asp, Asp, Asn, D-Asn, Gln, D-Gln Glycine G Ala, D-Ala, Pro, D-Pro, b-Ala, Acp Isoleucine I D-Ile, Val, D-Val, Leu, D-Leu, Met, D-Met Leucine L D-Leu, Val, D-Val, Leu, D-Leu, Met, D-Met Lysine K D-Lys, Arg, D-Arg, homo-Arg, D-homo-Arg, Met, D-Met, Ile, D-Ile, Orn, D-Orn Methionine M D-Met, S-Me-Cys, Ile, D-Ile, Leu, D-Leu, Val, D-Val Phenylalanine F D-Phe, Tyr, D-Thr, L-Dopa, His, D-His, Trp, D-Trp, Trans-3,4, or 5-phenylproline, cis-3,4, or 5-phenylproline Proline P D-Pro, L-I-thioazolidine-4-carboxylic acid, D-or L-1-oxazolidine-4-carboxylic acid Serine S D-Ser, Thr, D-Thr, allo-Thr, Met, D-Met, Met(O), D-Met(O), L-Cys, D-Cys Threonine T D-Thr, Ser, D-Ser, allo-Thr, Met, D-Met, Met(O), D-Met(O), Val, D-Val Tyrosine Y D-Tyr, Phe, D-Phe, L-Dopa, His, D-His Valine V D-Val, Leu, D-Leu, Ile, D-Ile, Met, D-Met
[0346] The reader will appreciate that some of the above mentioned conservative mutations can be produced by genetic manipulation, while others are produced by introducing synthetic amino acids into a polypeptide by genetic or other means.
[0347] The present amylase may be "precursor," "immature," or "full-length," in which case they include a signal sequence, or "mature," in which case they lack a signal sequence. Mature forms of the polypeptides are generally the most useful. Unless otherwise noted, the amino acid residue numbering used herein refers to the mature forms of the respective amylase polypeptides. The present amylase polypeptides may also be truncated to remove the N or C-termini, so long as the resulting polypeptides retain amylase activity.
[0348] The present amylase may be a "chimeric" or "hybrid" polypeptide, in that it includes at least a portion of a first amylase polypeptide, and at least a portion of a second amylase polypeptide (such chimeric amylases have recently been "rediscovered" as domain-swap amylases). The present amylases may further include heterologous signal sequence, an epitope to allow tracking or purification, or the like. Exemplary heterologous signal sequences are from B. licheniformis amylase (LAT), B. subtilis (AmyE or AprE), and Streptomyces CelA.
2.5. Nucleotides Encoding Variant Amylase Polypeptides
[0349] In another aspect, nucleic acids encoding a variant amylase polypeptide are provided. The nucleic acid may encode a particular amylase polypeptide, or an amylase having a specified degree of amino acid sequence identity to the particular amylase.
[0350] In one example, the nucleic acid encodes an amylase having at least 60%, at least 65%, at least 70%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or even at least 99% homology/identity to SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5 (excluding the portion of the nucleic acid that encodes the signal sequence). It will be appreciated that due to the degeneracy of the genetic code, a plurality of nucleic acids may encode the same polypeptide.
[0351] In another example, the nucleic acid hybridizes under stringent or very stringent conditions to a nucleic acid encoding (or complementary to a nucleic acid encoding) an amylase having at least 60%, at least 65%, at least 70%, at least 75%, at least 76%, at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or even at least 99% homology/identity to SEQ ID NO: 1, SEQ ID NO: 3, or SEQ ID NO: 5 (excluding the portion of the nucleic acid that encodes the signal sequence).
[0352] In some embodiments, the nucleic acid hybridizes under stringent or very stringent conditions to the nucleic acid of SEQ ID NO: 7, SEQ ID NO: 33, or SEQ ID NO: 38, or to a nucleic acid complementary to these nucleic acids.
[0353] Nucleic acids may encode a "full-length" ("fl" or "FL") amylase, which includes a signal sequence, only the mature form of an amylase, which lacks the signal sequence, or a truncated form of an amylase, which lacks the N or C-terminus of the mature form.
[0354] A nucleic acid that encodes a .alpha.-amylase can be operably linked to various promoters and regulators in a vector suitable for expressing the .alpha.-amylase in host cells. Exemplary promoters are from B. licheniformis amylase (LAT), B. subtilis (AmyE or AprE), and Streptomyces CelA. Such a nucleic acid can also be linked to other coding sequences, e.g., to encode a chimeric polypeptide.
3. Production of Variant Amylases
[0355] The present variant amylases can be produced in host cells, for example, by secretion or intracellular expression. A cultured cell material (e.g., a whole-cell broth) comprising a variant amylase can be obtained following secretion of the variant amylase into the cell medium. Optionally, the variant amylase can be isolated from the host cells, or even isolated from the cell broth, depending on the desired purity of the final variant amylase. A gene encoding a variant amylase can be cloned and expressed according to methods well known in the art. Suitable host cells include bacterial, fungal (including yeast and filamentous fungi), and plant cells (including algae). Particularly useful host cells include Aspergillus niger, Aspergillus oryzae or Trichoderma reesei. Other host cells include bacterial cells, e.g., Bacillus subtilis or B. licheniformis, as well as Streptomyces.
[0356] The host cell further may express a nucleic acid encoding a homologous or heterologous glucoamylase, i.e., a glucoamylase that is not the same species as the host cell, or one or more other enzymes. The glucoamylase may be a variant glucoamylase, such as one of the glucoamylase variants disclosed in U.S. Pat. No. 8,058,033 (Danisco US Inc.), for example. Additionally, the host may express one or more accessory enzymes, proteins, peptides. These may benefit liquefaction, saccharification, fermentation, SSF, etc processes. Furthermore, the host cell may produce biochemicals in addition to enzymes used to digest the various feedstock(s). Such host cells may be useful for fermentation or simultaneous saccharification and fermentation processes to reduce or eliminate the need to add enzymes.
3.1. Vectors
[0357] A DNA construct comprising a nucleic acid encoding variant amylases can be constructed to be expressed in a host cell. Representative nucleic acids that encode variant amylases include SEQ ID NO: 4. Because of the well-known degeneracy in the genetic code, variant polynucleotides that encode an identical amino acid sequence can be designed and made with routine skill. It is also well-known in the art to optimize codon use for a particular host cell. Nucleic acids encoding variant amylases can be incorporated into a vector. Vectors can be transferred to a host cell using well-known transformation techniques, such as those disclosed below.
[0358] The vector may be any vector that can be transformed into and replicated within a host cell. For example, a vector comprising a nucleic acid encoding a variant amylase can be transformed and replicated in a bacterial host cell as a means of propagating and amplifying the vector. The vector also may be transformed into an expression host, so that the encoding nucleic acids can be expressed as a functional amylase. Host cells that serve as expression hosts can include filamentous fungi, for example. The Fungal Genetics Stock Center (FGSC) Catalogue of Strains lists suitable vectors for expression in fungal host cells. See FGSC, Catalogue of Strains, University of Missouri, at www.fgsc.net (last modified Jan. 17, 2007). A representative vector is pJG153, a promoterless Cre expression vector that can be replicated in a bacterial host. See Harrison et al. (June 2011) Applied Environ. Microbiol. 77: 3916-22. pJG153 can be modified with routine skill to comprise and express a nucleic acid encoding an amylase variant.
[0359] A nucleic acid encoding a variant amylase can be operably linked to a suitable promoter, which allows transcription in the host cell. The promoter may be any DNA sequence that shows transcriptional activity in the host cell of choice and may be derived from genes encoding proteins either homologous or heterologous to the host cell. Exemplary promoters for directing the transcription of the DNA sequence encoding a variant amylase, especially in a bacterial host, are the promoter of the lac operon of E. coli, the Streptomyces coelicolor agarase gene dagA or celA promoters, the promoters of the Bacillus licheniformis .alpha.-amylase gene (amyL), the promoters of the Bacillus stearothermophilus maltogenic amylase gene (amyM), the promoters of the Bacillus amyloliquefaciens .alpha.-amylase (amyQ), the promoters of the Bacillus subtilis xylA and xylB genes etc. For transcription in a fungal host, examples of useful promoters are those derived from the gene encoding Aspergillus oryzae TAKA amylase, Rhizomucor miehei aspartic proteinase, Aspergillus niger neutral .alpha.-amylase, A. niger acid stable .alpha.-amylase, A. niger glucoamylase, Rhizomucor miehei lipase, A. oryzae alkaline protease, A. oryzae triose phosphate isomerase, or A. nidulans acetamidase. When a gene encoding an amylase is expressed in a bacterial species such as E. coli, a suitable promoter can be selected, for example, from a bacteriophage promoter including a T7 promoter and a phage lambda promoter. Examples of suitable promoters for the expression in a yeast species include but are not limited to the Gal 1 and Gal 10 promoters of Saccharomyces cerevisiae and the Pichia pastoris AOX1 or AOX2 promoters. cbh1 is an endogenous, inducible promoter from T. reesei. See Liu et al. (2008) "Improved heterologous gene expression in Trichoderma reesei by cellobiohydrolase I gene (cbh1) promoter optimization," Acta Biochim. Biophys. Sin (Shanghai) 40(2): 158-65.
[0360] The coding sequence can be operably linked to a signal sequence. The DNA encoding the signal sequence may be the DNA sequence naturally associated with the amylase gene to be expressed or from a different Genus or species. A signal sequence and a promoter sequence comprising a DNA construct or vector can be introduced into a fungal host cell and can be derived from the same source. For example, the signal sequence is the cbh1 signal sequence that is operably linked to a cbh1 promoter.
[0361] An expression vector may also comprise a suitable transcription terminator and, in eukaryotes, polyadenylation sequences operably linked to the DNA sequence encoding a variant amylase. Termination and polyadenylation sequences may suitably be derived from the same sources as the promoter.
[0362] The vector may further comprise a DNA sequence enabling the vector to replicate in the host cell. Examples of such sequences are the origins of replication of plasmids pUC19, pACYC177, pUB110, pE194, pAMB1, and pIJ702.
[0363] The vector may also comprise a selectable marker, e.g., a gene the product of which complements a defect in the isolated host cell, such as the dal genes from B. subtilis or B. licheniformis, or a gene that confers antibiotic resistance such as, e.g., ampicillin, kanamycin, chloramphenicol or tetracycline resistance. Furthermore, the vector may comprise Aspergillus selection markers such as amdS, argB, niaD and xxsC, a marker giving rise to hygromycin resistance, or the selection may be accomplished by co-transformation, such as known in the art. See e.g., International PCT Application WO 91/17243.
[0364] Intracellular expression may be advantageous in some respects, e.g., when using certain bacteria or fungi as host cells to produce large amounts of amylase for subsequent enrichment or purification. Extracellular secretion of amylase into the culture medium can also be used to make a cultured cell material comprising the isolated amylase.
[0365] The expression vector typically includes the components of a cloning vector, such as, for example, an element that permits autonomous replication of the vector in the selected host organism and one or more phenotypically detectable markers for selection purposes. The expression vector normally comprises control nucleotide sequences such as a promoter, operator, ribosome binding site, translation initiation signal and optionally, a repressor gene or one or more activator genes. Additionally, the expression vector may comprise a sequence coding for an amino acid sequence capable of targeting the amylase to a host cell organelle such as a peroxisome, or to a particular host cell compartment. Such a targeting sequence includes but is not limited to the sequence, SKL. For expression under the direction of control sequences, the nucleic acid sequence of the amylase is operably linked to the control sequences in proper manner with respect to expression.
[0366] The procedures used to ligate the DNA construct encoding an amylase, the promoter, terminator and other elements, respectively, and to insert them into suitable vectors containing the information necessary for replication, are well known to persons skilled in the art (see, e.g., Sambrook et al., MOLECULAR CLONING: A LABORATORY MANUAL, 2.sup.nd ed., Cold Spring Harbor, 1989, and 3.sup.rd ed., 2001).
3.2. Transformation and Culture of Host Cells
[0367] An isolated cell, either comprising a DNA construct or an expression vector, is advantageously used as a host cell in the recombinant production of an amylase. The cell may be transformed with the DNA construct encoding the enzyme, conveniently by integrating the DNA construct (in one or more copies) in the host chromosome. This integration is generally considered to be an advantage, as the DNA sequence is more likely to be stably maintained in the cell. Integration of the DNA constructs into the host chromosome may be performed according to conventional methods, e.g., by homologous or heterologous recombination. Alternatively, the cell may be transformed with an expression vector as described above in connection with the different types of host cells.
[0368] Examples of suitable bacterial host organisms are Gram positive bacterial species such as Bacillaceae including Bacillus subtilis, Bacillus licheniformis, Bacillus lentus, Bacillus brevis, Geobacillus (formerly Bacillus) stearothermophilus, Bacillus alkalophilus, Bacillus amyloliquefaciens, Bacillus coagulans, Bacillus lautus, Bacillus megaterium, and Bacillus thuringiensis; Streptomyces species such as Streptomyces murinus; lactic acid bacterial species including Lactococcus sp. such as Lactococcus lactis; Lactobacillus sp. including Lactobacillus reuteri; Leuconostoc sp.; Pediococcus sp.; and Streptococcus sp. Alternatively, strains of a Gram negative bacterial species belonging to Enterobacteriaceae including E. coli, or to Pseudomonadaceae can be selected as the host organism.
[0369] A suitable yeast host organism can be selected from the biotechnologically relevant yeasts species such as but not limited to yeast species such as Pichia sp., Hansenula sp., or Kluyveromyces, Yarrowinia, Schizosaccharomyces species or a species of Saccharomyces, including Saccharomyces cerevisiae or a species belonging to Schizosaccharomyces such as, for example, S. pombe. A strain of the methylotrophic yeast species, Pichia pastoris, can be used as the host organism. Alternatively, the host organism can be a Hansenula species. Suitable host organisms among filamentous fungi include species of Aspergillus, e.g., Aspergillus niger, Aspergillus oryzae, Aspergillus tubigensis, Aspergillus awamori, or Aspergillus nidulans. Alternatively, strains of a Fusarium sp., e.g., Fusarium oxysporum or of a Rhizomucor sp. such as Rhizomucor miehei can be used as the host organism. Other suitable strains include Thermomyces and Mucor sp. In addition, Trichoderma sp. can be used as a host. A suitable procedure for transformation of Aspergillus host cells includes, for example, that described in EP238023. An amylase expressed by a fungal host cell can be glycosylated, i.e., will comprise a glycosyl moiety. The glycosylation pattern can be the same or different as present in the wild-type amylase. The type and/or degree of glycosylation may impart changes in enzymatic and/or biochemical properties.
[0370] It is advantageous to delete genes from expression hosts, where the gene deficiency can be cured by the transformed expression vector. Known methods may be used to obtain a fungal host cell having one or more inactivated genes. Gene inactivation may be accomplished by complete or partial deletion, by insertional inactivation or by any other means that renders a gene nonfunctional for its intended purpose, such that the gene is prevented from expression of a functional protein. Any gene from a Trichoderma sp. or other filamentous fungal host that has been cloned can be deleted, for example, cbh1, cbh2, egl1, and egl2 genes. Gene deletion may be accomplished by inserting a form of the desired gene to be inactivated into a plasmid by methods known in the art.
[0371] Introduction of a DNA construct or vector into a host cell includes techniques such as transformation; electroporation; nuclear microinjection; transduction; transfection, e.g., lipofection mediated and DEAE-Dextrin mediated transfection; incubation with calcium phosphate DNA precipitate; high velocity bombardment with DNA-coated microprojectiles; and protoplast fusion. General transformation techniques are known in the art. See, e.g., Sambrook et al. (2001), supra. The expression of heterologous protein in Trichoderma is described, for example, in U.S. Pat. No. 6,022,725. Reference is also made to Cao et al. (2000) Science 9:991-1001 for transformation of Aspergillus strains. Genetically stable transformants can be constructed with vector systems whereby the nucleic acid encoding an amylase is stably integrated into a host cell chromosome. Transformants are then selected and purified by known techniques.
[0372] The preparation of Trichoderma sp. for transformation, for example, may involve the preparation of protoplasts from fungal mycelia. See Campbell et al. (1989) Curr. Genet. 16: 53-56. The mycelia can be obtained from germinated vegetative spores. The mycelia are treated with an enzyme that digests the cell wall, resulting in protoplasts. The protoplasts are protected by the presence of an osmotic stabilizer in the suspending medium. These stabilizers include sorbitol, mannitol, potassium chloride, magnesium sulfate, and the like. Usually the concentration of these stabilizers varies between 0.8 M and 1.2 M, e.g., a 1.2 M solution of sorbitol can be used in the suspension medium.
[0373] Uptake of DNA into the host Trichoderma sp. strain depends upon the calcium ion concentration. Generally, between about 10-50 mM CaCl2 is used in an uptake solution. Additional suitable compounds include a buffering system, such as TE buffer (10 mM Tris, pH 7.4; 1 mM EDTA) or 10 mM MOPS, pH 6.0 and polyethylene glycol. The polyethylene glycol is believed to fuse the cell membranes, thus permitting the contents of the medium to be delivered into the cytoplasm of the Trichoderma sp. strain. This fusion frequently leaves multiple copies of the plasmid DNA integrated into the host chromosome.
[0374] Usually transformation of Trichoderma sp. uses protoplasts or cells that have been subjected to a permeability treatment, typically at a density of 10.sup.5 to 10.sup.7/mL, particularly 2.times.10.sup.6/mL. A volume of 100 .mu.L of these protoplasts or cells in an appropriate solution (e.g., 1.2 M sorbitol and 50 mM CaCl.sub.2)) may be mixed with the desired DNA. Generally, a high concentration of PEG is added to the uptake solution. From 0.1 to 1 volume of 25% PEG 4000 can be added to the protoplast suspension; however, it is useful to add about 0.25 volumes to the protoplast suspension. Additives, such as dimethyl sulfoxide, heparin, spermidine, potassium chloride and the like, may also be added to the uptake solution to facilitate transformation. Similar procedures are available for other fungal host cells. See, e.g., U.S. Pat. No. 6,022,725.
3.3. Expression
[0375] A method of producing an amylase may comprise cultivating a host cell as described above under conditions conducive to the production of the enzyme and recovering the enzyme from the cells and/or culture medium.
[0376] The medium used to cultivate the cells may be any conventional medium suitable for growing the host cell in question and obtaining expression of an amylase. Suitable media and media components are available from commercial suppliers or may be prepared according to published recipes (e.g., as described in catalogues of the American Type Culture Collection).
[0377] An enzyme secreted from the host cells can be used in a whole broth preparation. In the present methods, the preparation of a spent whole fermentation broth of a recombinant microorganism can be achieved using any cultivation method known in the art resulting in the expression of an .alpha.-amylase. Fermentation may, therefore, be understood as comprising shake flask cultivation, small- or large-scale fermentation (including continuous, batch, fed-batch, or solid state fermentations) in laboratory or industrial fermenters performed in a suitable medium and under conditions allowing the amylase to be expressed or isolated. The term "spent whole fermentation broth" is defined herein as unfractionated contents of fermentation material that includes culture medium, extracellular proteins (e.g., enzymes), and cellular biomass. It is understood that the term "spent whole fermentation broth" also encompasses cellular biomass that has been lysed or permeabilized using methods well known in the art.
[0378] An enzyme secreted from the host cells may conveniently be recovered from the culture medium by well-known procedures, including separating the cells from the medium by centrifugation or filtration, and precipitating proteinaceous components of the medium by means of a salt such as ammonium sulfate, followed by the use of chromatographic procedures such as ion exchange chromatography, affinity chromatography, or the like.
[0379] The polynucleotide encoding an amylase in a vector can be operably linked to a control sequence that is capable of providing for the expression of the coding sequence by the host cell, i.e. the vector is an expression vector. The control sequences may be modified, for example by the addition of further transcriptional regulatory elements to make the level of transcription directed by the control sequences more responsive to transcriptional modulators. The control sequences may in particular comprise promoters.
[0380] Host cells may be cultured under suitable conditions that allow expression of an amylase. Expression of the enzymes may be constitutive such that they are continually produced, or inducible, requiring a stimulus to initiate expression. In the case of inducible expression, protein production can be initiated when required by, for example, addition of an inducer substance to the culture medium, for example dexamethasone or IPTG or Sophorose. Polypeptides can also be produced recombinantly in an in vitro cell-free system, such as the TNT.TM. (Promega) rabbit reticulocyte system.
[0381] An expression host also can be cultured in the appropriate medium for the host, under aerobic conditions. Shaking or a combination of agitation and aeration can be provided, with production occurring at the appropriate temperature for that host, e.g., from about 25.degree. C. to about 75.degree. C. (e.g., 30.degree. C. to 45.degree. C.), depending on the needs of the host and production of the desired variant amylase. Culturing can occur from about 12 to about 100 hours or greater (and any hour value there between, e.g., from 24 to 72 hours). Typically, the culture broth is at a pH of about 4.0 to about 8.0, again depending on the culture conditions needed for the host relative to production of an amylase.
3.4. Identification of Amylase Activity
[0382] To evaluate the expression of an amylase in a host cell, assays can measure the expressed protein, corresponding mRNA, or .alpha.-amylase activity. For example, suitable assays include Northern blotting, reverse transcriptase polymerase chain reaction, and in situ hybridization, using an appropriately labeled hybridizing probe. Suitable assays also include measuring amylase activity in a sample, for example, by assays directly measuring reducing sugars such as glucose in the culture media. For example, glucose concentration may be determined using glucose reagent kit No. 15-UV (Sigma Chemical Co.) or an instrument, such as Technicon Autoanalyzer. .alpha.-amylase activity also may be measured by any known method, such as the PAHBAH or ABTS assays, described below.
3.5. Methods for Enriching and Purifying Variants Amylases
[0383] Fermentation, separation, and concentration techniques are well known in the art and conventional methods can be used in order to prepare a concentrated a variant .alpha.-amylase polypeptide-containing solution.
[0384] After fermentation, a fermentation broth is obtained, the microbial cells and various suspended solids, including residual raw fermentation materials, are removed by conventional separation techniques in order to obtain an amylase solution. Filtration, centrifugation, microfiltration, rotary vacuum drum filtration, ultrafiltration, centrifugation followed by ultra-filtration, extraction, or chromatography, or the like, are generally used.
[0385] It is desirable to concentrate a variant .alpha.-amylase polypeptide-containing solution in order to optimize recovery. Use of unconcentrated solutions requires increased incubation time in order to collect the enriched or purified enzyme precipitate.
[0386] The enzyme containing solution is concentrated using conventional concentration techniques until the desired enzyme level is obtained. Concentration of the enzyme containing solution may be achieved by any of the techniques discussed herein. Exemplary methods of enrichment and purification include but are not limited to rotary vacuum filtration and/or ultrafiltration.
[0387] The enzyme solution is concentrated into a concentrated enzyme solution until the enzyme activity of the concentrated variant .alpha.-amylase polypeptide-containing solution is at a desired level.
[0388] Concentration may be performed using, e.g., a precipitation agent, such as a metal halide precipitation agent. Metal halide precipitation agents include but are not limited to alkali metal chlorides, alkali metal bromides and blends of two or more of these metal halides. Exemplary metal halides include sodium chloride, potassium chloride, sodium bromide, potassium bromide and blends of two or more of these metal halides. The metal halide precipitation agent, sodium chloride, can also be used as a preservative.
[0389] The metal halide precipitation agent is used in an amount effective to precipitate an amylase. The selection of at least an effective amount and an optimum amount of metal halide effective to cause precipitation of the enzyme, as well as the conditions of the precipitation for maximum recovery including incubation time, pH, temperature and concentration of enzyme, will be readily apparent to one of ordinary skill in the art, after routine testing.
[0390] Generally, at least about 5% w/v (weight/volume) to about 25% w/v of metal halide is added to the concentrated enzyme solution, and usually at least 8% w/v. Generally, no more than about 25% w/v of metal halide is added to the concentrated enzyme solution and usually no more than about 20% w/v. The optimal concentration of the metal halide precipitation agent will depend, among others, on the nature of the specific variant .alpha.-amylase polypeptide and on its concentration in the concentrated enzyme solution.
[0391] Another alternative way to precipitate the enzyme is to use organic compounds. Exemplary organic compound precipitating agents include: 4-hydroxybenzoic acid, alkali metal salts of 4-hydroxybenzoic acid, alkyl esters of 4-hydroxybenzoic acid, and blends of two or more of these organic compounds. The addition of the organic compound precipitation agents can take place prior to, simultaneously with or subsequent to the addition of the metal halide precipitation agent, and the addition of both precipitation agents, organic compound and metal halide, may be carried out sequentially or simultaneously.
[0392] Generally, the organic precipitation agents are selected from the group consisting of alkali metal salts of 4-hydroxybenzoic acid, such as sodium or potassium salts, and linear or branched alkyl esters of 4-hydroxybenzoic acid, wherein the alkyl group contains from 1 to 12 carbon atoms, and blends of two or more of these organic compounds. The organic compound precipitation agents can be, for example, linear or branched alkyl esters of 4-hydroxybenzoic acid, wherein the alkyl group contains from 1 to 10 carbon atoms, and blends of two or more of these organic compounds. Exemplary organic compounds are linear alkyl esters of 4-hydroxybenzoic acid, wherein the alkyl group contains from 1 to 6 carbon atoms, and blends of two or more of these organic compounds. Methyl esters of 4-hydroxybenzoic acid, propyl esters of 4-hydroxybenzoic acid, butyl ester of 4-hydroxybenzoic acid, ethyl ester of 4-hydroxybenzoic acid and blends of two or more of these organic compounds can also be used. Additional organic compounds also include but are not limited to 4-hydroxybenzoic acid methyl ester (named methyl PARABEN), 4-hydroxybenzoic acid propyl ester (named propyl PARABEN), which also are both amylase preservative agents. For further descriptions, see, e.g., U.S. Pat. No. 5,281,526.
[0393] Addition of the organic compound precipitation agent provides the advantage of high flexibility of the precipitation conditions with respect to pH, temperature, variant amylase concentration, precipitation agent concentration, and time of incubation.
[0394] The organic compound precipitation agent is used in an amount effective to improve precipitation of the enzyme by means of the metal halide precipitation agent. The selection of at least an effective amount and an optimum amount of organic compound precipitation agent, as well as the conditions of the precipitation for maximum recovery including incubation time, pH, temperature and concentration of enzyme, will be readily apparent to one of ordinary skill in the art, in light of the present disclosure, after routine testing.
[0395] Generally, at least about 0.01% w/v of organic compound precipitation agent is added to the concentrated enzyme solution and usually at least about 0.02% w/v. Generally, no more than about 0.3% w/v of organic compound precipitation agent is added to the concentrated enzyme solution and usually no more than about 0.2% w/v.
[0396] The concentrated polypeptide solution, containing the metal halide precipitation agent, and the organic compound precipitation agent, can be adjusted to a pH, which will, of necessity, depend on the enzyme to be enriched or purified. Generally, the pH is adjusted at a level near the isoelectric point of the amylase. The pH can be adjusted at a pH in a range from about 2.5 pH units below the isoelectric point (pI) up to about 2.5 pH units above the isoelectric point.
[0397] The incubation time necessary to obtain an enriched or purified enzyme precipitate depends on the nature of the specific enzyme, the concentration of enzyme, and the specific precipitation agent(s) and its (their) concentration. Generally, the time effective to precipitate the enzyme is between about 1 to about 30 hours; usually it does not exceed about 25 hours. In the presence of the organic compound precipitation agent, the time of incubation can still be reduced to less about 10 hours and in most cases even about 6 hours.
[0398] Generally, the temperature during incubation is between about 4.degree. C. and about 50.degree. C. Usually, the method is carried out at a temperature between about 10.degree. C. and about 45.degree. C. (e.g., between about 20.degree. C. and about 40.degree. C.). The optimal temperature for inducing precipitation varies according to the solution conditions and the enzyme or precipitation agent(s) used.
[0399] The overall recovery of enriched or purified enzyme precipitate, and the efficiency with which the process is conducted, is improved by agitating the solution comprising the enzyme, the added metal halide and the added organic compound. The agitation step is done both during addition of the metal halide and the organic compound, and during the subsequent incubation period. Suitable agitation methods include mechanical stirring or shaking, vigorous aeration, or any similar technique.
[0400] After the incubation period, the enriched or purified enzyme is then separated from the dissociated pigment and other impurities and collected by conventional separation techniques, such as filtration, centrifugation, microfiltration, rotary vacuum filtration, ultrafiltration, press filtration, cross membrane microfiltration, cross flow membrane microfiltration, or the like. Further enrichment or purification of the enzyme precipitate can be obtained by washing the precipitate with water. For example, the enriched or purified enzyme precipitate is washed with water containing the metal halide precipitation agent, or with water containing the metal halide and the organic compound precipitation agents.
[0401] During fermentation, a variant .alpha.-amylase polypeptide accumulates in the culture broth. For the isolation, enrichment, or purification of the desired variant .alpha.-amylase, the culture broth is centrifuged or filtered to eliminate cells, and the resulting cell-free liquid is used for enzyme enrichment or purification. In one embodiment, the cell-free broth is subjected to salting out using ammonium sulfate at about 70% saturation; the 70% saturation-precipitation fraction is then dissolved in a buffer and applied to a column such as a Sephadex G-100 column, and eluted to recover the enzyme-active fraction. For further enrichment or purification, a conventional procedure such as ion exchange chromatography may be used.
[0402] Enriched or purified enzymes are useful for laundry and cleaning applications. For example, they can be used in laundry detergents and spot removers. They can be made into a final product that is either liquid (solution, slurry) or solid (granular, powder).
[0403] A more specific example of enrichment or purification, is described in Sumitani et al. (2000) "New type of starch-binding domain: the direct repeat motif in the C-terminal region of Bacillus sp. 195 .alpha.-amylase contributes to starch binding and raw starch degrading," Biochem. J. 350: 477-484, and is briefly summarized here. The enzyme obtained from 4 liters of a Streptomyces lividans TK24 culture supernatant was treated with (NH.sub.4).sub.2SO.sub.4 at 80% saturation. The precipitate was recovered by centrifugation at 10,000.times.g (20 min. and 4.degree. C.) and re-dissolved in 20 mM Tris/HCl buffer (pH 7.0) containing 5 mM CaCl.sub.2). The solubilized precipitate was then dialyzed against the same buffer. The dialyzed sample was then applied to a Sephacryl S-200 column, which had previously been equilibrated with 20 mM Tris/HCl buffer, (pH 7.0), 5 mM CaCl2, and eluted at a linear flow rate of 7 mL/hr with the same buffer. Fractions from the column were collected and assessed for activity as judged by enzyme assay and SDS-PAGE. The protein was further purified as follows. A Toyopearl HW55 column (Tosoh Bioscience, Montgomeryville, Pa.; Cat. No. 19812) was equilibrated with 20 mM Tris/HCl buffer (pH 7.0) containing 5 mM CaCl2 and 1.5 M (NH4)2SO4. The enzyme was eluted with a linear gradient of 1.5 to 0 M (NH.sub.4).sub.2SO.sub.4 in 20 mM Tris/HCL buffer, pH 7.0 containing 5 mM CaCl.sub.2). The active fractions were collected, and the enzyme precipitated with (NH.sub.4).sub.2SO.sub.4 at 80% saturation. The precipitate was recovered, re-dissolved, and dialyzed as described above. The dialyzed sample was then applied to a Mono Q HR5/5 column (Amersham Pharmacia; Cat. No. 17-5167-01) previously equilibrated with 20 mM Tris/HCl buffer (pH 7.0) containing 5 mM CaCl.sub.2), at a flow rate of 60 mL/hour. The active fractions are collected and added to a 1.5 M (NH.sub.4).sub.2SO.sub.4 solution. The active enzyme fractions were re-chromatographed on a Toyopearl HW55 column, as before, to yield a homogeneous enzyme as determined by SDS-PAGE. See, e.g., Sumitani et al. (2000) Biochem. J. 350: 477-484, for general discussion of the method and variations thereon.
[0404] For production scale recovery, variant .alpha.-amylase polypeptides can be enriched or partially purified as generally described above by removing cells via flocculation with polymers. Alternatively, the enzyme can be enriched or purified by microfiltration followed by concentration by ultrafiltration using available membranes and equipment. However, for some applications, the enzyme does not need to be enriched or purified, and whole broth culture can be lysed and used without further treatment. The enzyme can then be processed, for example, into granules.
4. Compositions and Uses of Variant Amylases
[0405] Variants amylases are useful for a variety of industrial applications. For example, variant amylases are useful in a starch conversion process, particularly in a saccharification process of a starch that has undergone liquefaction. The desired end-product may be any product that may be produced by the enzymatic conversion of the starch substrate. For example, the desired product may be a syrup rich in glucose and maltose, which can be used in other processes, such as the preparation of HFCS, or which can be converted into a number of other useful products, such as ascorbic acid intermediates (e.g., gluconate; 2-keto-L-gulonic acid; 5-keto-gluconate; and 2,5-diketogluconate); 1,3-propanediol; aromatic amino acids (e.g., tyrosine, phenylalanine and tryptophan); organic acids (e.g., lactate, pyruvate, succinate, isocitrate, and oxaloacetate); amino acids (e.g., serine and glycine); antibiotics; antimicrobials; enzymes; vitamins; and hormones.
[0406] The starch conversion process may be a precursor to, or simultaneous with, a fermentation process designed to produce alcohol for fuel or drinking (i.e., potable alcohol). One skilled in the art is aware of various fermentation conditions that may be used in the production of these end-products. Variant amylases are also useful in compositions and methods of food preparation. These various uses of variant amylases are described in more detail below.
4.1. Preparation of Starch Substrates
[0407] Those of general skill in the art are well aware of available methods that may be used to prepare starch substrates for use in the processes disclosed herein. For example, a useful starch substrate may be obtained from tubers, roots, stems, legumes, cereals or whole grain. More specifically, the granular starch may be obtained from corn, cobs, wheat, barley, rye, triticale, milo, sago, millet, cassava, tapioca, sorghum, rice, peas, bean, banana, or potatoes. Corn contains about 60-68% starch; barley contains about 55-65% starch; millet contains about 75-80% starch; wheat contains about 60-65% starch; and polished rice contains 70-72% starch. Specifically contemplated starch substrates are corn starch and wheat starch. The starch from a grain may be ground or whole and includes corn solids, such as kernels, bran and/or cobs. The starch may also be highly refined raw starch or feedstock from starch refinery processes. Various starches also are commercially available. For example, corn starch is available from Cerestar, Sigma, and Katayama Chemical Industry Co. (Japan); wheat starch is available from Sigma; sweet potato starch is available from Wako Pure Chemical Industry Co. (Japan); and potato starch is available from Nakaari Chemical Pharmaceutical Co. (Japan).
[0408] The starch substrate can be a crude starch from milled whole grain, which contains non-starch fractions, e.g., germ residues and fibers. Milling may comprise either wet milling or dry milling or grinding. In wet milling, whole grain is soaked in water or dilute acid to separate the grain into its component parts, e.g., starch, protein, germ, oil, kernel fibers. Wet milling efficiently separates the germ and meal (i.e., starch granules and protein) and is especially suitable for production of syrups. In dry milling or grinding, whole kernels are ground into a fine powder and often processed without fractionating the grain into its component parts. In some cases, oils from the kernels are recovered. Dry ground grain thus will comprise significant amounts of non-starch carbohydrate compounds, in addition to starch. Dry grinding of the starch substrate can be used for production of ethanol and other biochemicals. The starch to be processed may be a highly refined starch quality, for example, at least 90%, at least 95%, at least 97%, or at least 99.5% pure.
4.2. Gelatinization and Liquefaction of Starch
[0409] As used herein, the term "liquefaction" or "liquefy" means a process by which starch is converted to less viscous and shorter chain dextrins. Generally, this process involves gelatinization of starch simultaneously with or followed by the addition of an .alpha.-amylase, although additional liquefaction-inducing enzymes optionally may be added. In some embodiments, the starch substrate prepared as described above is slurried with water. The starch slurry may contain starch as a weight percent of dry solids of about 10-55%, about 20-45%, about 30-45%, about 30-40%, or about 30-35%. .alpha.-amylase may be added to the slurry, with a metering pump, for example. The .alpha.-amylase typically used for this application is a thermally stable, bacterial .alpha.-amylase, such as a Geobacillus stearothermophilus .alpha.-amylase. The .alpha.-amylase is usually supplied, for example, at about 1500 units per kg dry matter of starch. To optimize .alpha.-amylase stability and activity, the pH of the slurry typically is adjusted to about pH 5.5-6.5 and about 1 mM of calcium (about 40 ppm free calcium ions) can also be added. Bacterial .alpha.-amylase remaining in the slurry following liquefaction may be deactivated via a number of methods, including lowering the pH in a subsequent reaction step or by removing calcium from the slurry in cases where the enzyme is dependent upon calcium.
[0410] The slurry of starch plus the .alpha.-amylase may be pumped continuously through a jet cooker, which is steam heated to 105.degree. C. Gelatinization occurs rapidly under these conditions, and the enzymatic activity, combined with the significant shear forces, begins the hydrolysis of the starch substrate. The residence time in the jet cooker is brief. The partly gelatinized starch may be passed into a series of holding tubes maintained at 105-110.degree. C. and held for 5-8 min. to complete the gelatinization process ("primary liquefaction"). Hydrolysis to the required DE is completed in holding tanks at 85-95.degree. C. or higher temperatures for about 1 to 2 hours ("secondary liquefaction"). These tanks may contain baffles to discourage back mixing. As used herein, the term "minutes of secondary liquefaction" refers to the time that has elapsed from the start of secondary liquefaction to the time that the Dextrose Equivalent (DE) is measured. The slurry is then allowed to cool to room temperature. This cooling step can be 30 minutes to 180 minutes, e.g., 90 minutes to 120 minutes. The liquefied starch typically is in the form of a slurry having a dry solids content (w/w) of about 10-50%; about 10-45%; about 15-40%; about 20-40%; about 25-40%; or about 25-35%.
[0411] Liquefaction with variant amylases advantageously can be conducted at low pH, eliminating the requirement to adjust the pH to about pH 5.5-6.5. Variants amylases can be used for liquefaction at a pH range of 2 to 7, e.g., pH 3.0-7.5, pH 4.0-6.0, or pH 4.5-5.8. Variant amylases can maintain liquefying activity at a temperature range of about 85.degree. C.-95.degree. C., e.g., 85.degree. C., 90.degree. C., or 95.degree. C. For example, liquefaction can be conducted with 800 .mu.g an amylase in a solution of 25% DS corn starch for 10 min at pH 5.8 and 85.degree. C., or pH 4.5 and 95.degree. C., for example. Liquefying activity can be assayed using any of a number of known viscosity assays in the art.
[0412] In particular embodiments using the present amylase variants, starch liquifaction is performed at a temperature range of 90-115.degree. C., for the purpose of producing high-purity glucose syrups, HFCS, maltodextrins, etc.
4.3. Saccharification
[0413] The liquefied starch can be saccharified into a syrup rich in lower DP (e.g., DP1+DP2) saccharides, using variant amylases, optionally in the presence of another enzyme(s). The exact composition of the products of saccharification depends on the combination of enzymes used, as well as the type of granular starch processed. Advantageously, the syrup obtainable using the provided variant amylases may contain a weight percent of DP2 of the total oligosaccharides in the saccharified starch exceeding 30%, e.g., 45%-65% or 55%-65%. The weight percent of (DP1+DP2) in the saccharified starch may exceed about 70%, e.g., 75%-85% or 80%-85%. The present amylases also produce a relatively high yield of glucose, e.g., DP1>20%, in the syrup product.
[0414] Whereas liquefaction is generally run as a continuous process, saccharification is often conducted as a batch process. Saccharification typically is most effective at temperatures of about 60-65.degree. C. and a pH of about 4.0-4.5, e.g., pH 4.3, necessitating cooling and adjusting the pH of the liquefied starch. Saccharification may be performed, for example, at a temperature between about 40.degree. C., about 50.degree. C., or about 55.degree. C. to about 60.degree. C. or about 65.degree. C. Saccharification is normally conducted in stirred tanks, which may take several hours to fill or empty. Enzymes typically are added either at a fixed ratio to dried solids as the tanks are filled or added as a single dose at the commencement of the filling stage. A saccharification reaction to make a syrup typically is run over about 24-72 hours, for example, 24-48 hours. When a maximum or desired DE has been attained, the reaction is stopped by heating to 85.degree. C. for 5 min., for example. Further incubation will result in a lower DE, eventually to about 90 DE, as accumulated glucose re-polymerizes to isomaltose and/or other reversion products via an enzymatic reversion reaction and/or with the approach of thermodynamic equilibrium. When using an amylase, saccharification optimally is conducted at a temperature range of about 30.degree. C. to about 75.degree. C., e.g., 45.degree. C.-75.degree. C. or 47.degree. C.-74.degree. C. The saccharifying may be conducted over a pH range of about pH 3 to about pH 7, e.g., pH 3.0-pH 7.5, pH 3.5-pH 5.5, pH 3.5, pH 3.8, or pH 4.5.
[0415] An .alpha.-amylase may be added to the slurry in the form of a composition. An .alpha.-amylase can be added to a slurry of a granular starch substrate in an amount of about 0.6-10 ppm ds, e.g., 2 ppm ds. An .alpha.-amylase can be added as a whole broth, clarified, enriched, partially purified, or purified enzyme. The specific activity of the amylase may be about 300 U/mg of enzyme, for example, measured with the PAHBAH assay. The .alpha.-amylase also can be added as a whole broth product.
[0416] An .alpha.-amylase may be added to the slurry as an isolated enzyme solution. For example, an .alpha.-amylase can be added in the form of a cultured cell material produced by host cells expressing an amylase. An .alpha.-amylase may also be secreted by a host cell into the reaction medium during the fermentation or SSF process, such that the enzyme is provided continuously into the reaction. The host cell producing and secreting amylase may also express an additional enzyme, such as a glucoamylase. For example, U.S. Pat. No. 5,422,267 discloses the use of a glucoamylase in yeast for production of alcoholic beverages. For example, a host cell, e.g., Trichoderma reesei or Aspergillus niger, may be engineered to co-express an .alpha.-amylase and a glucoamylase, e.g., HgGA, TrGA, or a TrGA variant, during saccharification. The host cell can be genetically modified so as not to express its endogenous glucoamylase and/or other enzymes, proteins or other materials. The host cell can be engineered to express a broad spectrum of various saccharolytic enzymes. For example, the recombinant yeast host cell can comprise nucleic acids encoding a glucoamylase, an alpha-glucosidase, an enzyme that utilizes pentose sugar, an .alpha.-amylase, a pullulanase, an isoamylase, and/or an isopullulanase. See, e.g., WO 2011/153516 A2.
4.4. Isomerization
[0417] The soluble starch hydrolysate produced by treatment with amylase can be converted into high fructose starch-based syrup (HFSS), such as high fructose corn syrup (HFCS). This conversion can be achieved using a glucose isomerase, particularly a glucose isomerase immobilized on a solid support. The pH is increased to about 6.0 to about 8.0, e.g., pH 7.5 (depending on the isomerase), and Ca.sup.2+ is removed by ion exchange. Suitable isomerases include SWEETZYME.RTM., IT (Novozymes A/S); G-ZYME.RTM. IMGI, and G-ZYME.RTM. G993, KETOMAX.RTM., G-ZYME.RTM. G993, G-ZYME.RTM. G993 liquid, and GENSWEET.RTM. IGI. Following isomerization, the mixture typically contains about 40-45% fructose, e.g., 42% fructose.
4.5. Fermentation
[0418] The soluble starch hydrolysate, particularly a glucose rich syrup, can be fermented by contacting the starch hydrolysate with a fermenting organism typically at a temperature around 32.degree. C., such as from 30.degree. C. to 35.degree. C. for alcohol-producing yeast. The temperature and pH of the fermentation will depend upon the fermenting organism. EOF products include metabolites, such as citric acid, lactic acid, succinic acid, monosodium glutamate, gluconic acid, sodium gluconate, calcium gluconate, potassium gluconate, itaconic acid and other carboxylic acids, glucono delta-lactone, sodium erythorbate, lysine and other amino acids, omega 3 fatty acid, butanol, isoprene, 1,3-propanediol and other biomaterials.
[0419] Ethanologenic microorganisms include yeast, such as Saccharomyces cerevisiae and bacteria, e.g., Zymomonas moblis, expressing alcohol dehydrogenase and pyruvate decarboxylase. The ethanologenic microorganism can express xylose reductase and xylitol dehydrogenase, which convert xylose to xylulose. Improved strains of ethanologenic microorganisms, which can withstand higher temperatures, for example, are known in the art and can be used. See Liu et al. (2011) Sheng Wu Gong Cheng Xue Bao 27(7): 1049-56. Commercial sources of yeast include ETHANOL RED.RTM. (LeSaffre); Thermosacc.RTM. (Lallemand); RED STAR.RTM. (Red Star); FERMIOL.RTM. (DSM Specialties); and SUPERSTART.RTM. (Alltech). Microorganisms that produce other metabolites, such as citric acid and lactic acid, by fermentation are also known in the art. See, e.g., Papagianni (2007) "Advances in citric acid fermentation by Aspergillus niger: biochemical aspects, membrane transport and modeling," Biotechnol. Adv. 25(3): 244-63; John et al. (2009) "Direct lactic acid fermentation: focus on simultaneous saccharification and lactic acid production," Biotechnol. Adv. 27(2): 145-52.
[0420] The saccharification and fermentation processes may be carried out as an SSF process. Fermentation may comprise subsequent enrichment, purification, and recovery of ethanol, for example. During the fermentation, the ethanol content of the broth or "beer" may reach about 8-18% v/v, e.g., 14-15% v/v. The broth may be distilled to produce enriched, e.g., 96% pure, solutions of ethanol. Further, CO2 generated by fermentation may be collected with a CO2 scrubber, compressed, and marketed for other uses, e.g., carbonating beverage or dry ice production. Solid waste from the fermentation process may be used as protein-rich products, e.g., livestock feed.
[0421] As mentioned above, an SSF process can be conducted with fungal cells that express and secrete amylase continuously throughout SSF. The fungal cells expressing amylase also can be the fermenting microorganism, e.g., an ethanologenic microorganism. Ethanol production thus can be carried out using a fungal cell that expresses sufficient amylase so that less or no enzyme has to be added exogenously. The fungal host cell can be from an appropriately engineered fungal strain. Fungal host cells that express and secrete other enzymes, in addition to amylase, also can be used. Such cells may express glucoamylase and/or a pullulanase, phytase, alpha-glucosidase, isoamylase, beta-amylase cellulase, xylanase, other hemicellulases, protease, beta-glucosidase, pectinase, esterase, redox enzymes, transferase, or other enzyme.
[0422] A variation on this process is a "fed-batch fermentation" system, where the substrate is added in increments as the fermentation progresses. Fed-batch systems are useful when catabolite repression may inhibit the metabolism of the cells and where it is desirable to have limited amounts of substrate in the medium. The actual substrate concentration in fed-batch systems is estimated by the changes of measurable factors such as pH, dissolved oxygen and the partial pressure of waste gases, such as CO2. Batch and fed-batch fermentations are common and well known in the art.
[0423] Continuous fermentation is an open system where a defined fermentation medium is added continuously to a bioreactor, and an equal amount of conditioned medium is removed simultaneously for processing. Continuous fermentation generally maintains the cultures at a constant high density where cells are primarily in log phase growth. Continuous fermentation permits modulation of cell growth and/or product concentration. For example, a limiting nutrient such as the carbon source or nitrogen source is maintained at a fixed rate and all other parameters are allowed to moderate. Because growth is maintained at a steady state, cell loss due to medium being drawn off should be balanced against the cell growth rate in the fermentation. Methods of optimizing continuous fermentation processes and maximizing the rate of product formation are well known in the art of industrial microbiology.
4.6. Compositions Comprising Variants Amylases
[0424] Variant amylases may be combined with a glucoamylase (EC 3.2.1.3), e.g., a Trichoderma glucoamylase or variant thereof. An exemplary glucoamylase is Trichoderma reesei glucoamylase (TrGA) and variants thereof that possess superior specific activity and thermal stability. See U.S. Published Applications Nos. 2006/0094080, 2007/0004018, and 2007/0015266 (Danisco US Inc.). Suitable variants of TrGA include those with glucoamylase activity and at least 80%, at least 90%, or at least 95% sequence identity to wild-type TrGA. Variant amylases advantageously increase the yield of glucose produced in a saccharification process catalyzed by TrGA.
[0425] Alternatively, the glucoamylase may be another glucoamylase derived from plants (including algae), fungi, or bacteria. For example, the glucoamylases may be Aspergillus niger G1 or G2 glucoamylase or its variants (e.g., Boel et al. (1984) EMBO J. 3: 1097-1102; WO 92/00381; WO 00/04136 (Novo Nordisk A/S)); and A. awamori glucoamylase (e.g., WO 84/02921 (Cetus Corp.)). Other contemplated Aspergillus glucoamylase include variants with enhanced thermal stability, e.g., G137A and G139A (Chen et al. (1996) Prot. Eng. 9: 499-505); D257E and D293E/Q (Chen et al. (1995) Prot. Eng. 8: 575-582); N182 (Chen et al. (1994) Biochem. J. 301: 275-281); A246C (Fierobe et al. (1996) Biochemistry, 35: 8698-8704); and variants with Pro residues in positions A435 and 5436 (Li et al. (1997) Protein Eng. 10: 1199-1204). Other contemplated glucoamylases include Talaromyces glucoamylases, in particular derived from T. emersonii (e.g., WO 99/28448 (Novo Nordisk A/S), T. leycettanus (e.g., U.S. Pat. No. RE 32,153 (CPC International, Inc.)), T. duponti, or T. thermophilus (e.g., U.S. Pat. No. 4,587,215). Contemplated bacterial glucoamylases include glucoamylases from the genus Clostridium, in particular C. thermoamylolyticum (e.g., EP 135,138 (CPC International, Inc.) and C. thermohydrosulfuricum (e.g., WO 86/01831 (Michigan Biotechnology Institute)). Suitable glucoamylases include the glucoamylases derived from Aspergillus oryzae, such as a glucoamylase shown in SEQ ID NO:2 in WO 00/04136 (Novo Nordisk A/S). Also suitable are commercial glucoamylases, such as AMG 200L; AMG 300 L; SAN.TM. SUPER and AMG.TM. E (Novozymes); OPTIDEX.RTM. 300 and OPTIDEX L-400 (Danisco US Inc.); AMIGASE.TM. and AMIGASE.TM. PLUS (DSM); G-ZYME.RTM. G900 (Enzyme Bio-Systems); and G-ZYME.RTM. G990 ZR (A. niger glucoamylase with a low protease content). Still other suitable glucoamylases include Aspergillus fumigatus glucoamylase, Talaromyces glucoamylase, Thielavia glucoamylase, Trametes glucoamylase, Thermomyces glucoamylase, Athelia glucoamylase, or Humicola glucoamylase (e.g., HgGA). Glucoamylases typically are added in an amount of about 0.1-2 glucoamylase units (GAU)/g ds, e.g., about 0.16 GAU/g ds, 0.23 GAU/g ds, or 0.33 GAU/g ds.
[0426] Other suitable enzymes that can be used with amylase include a phytase, protease, pullulanase, .beta.-amylase, isoamylase, a different .alpha.-amylase, alpha-glucosidase, cellulase, xylanase, other hemicellulases, beta-glucosidase, transferase, pectinase, lipase, cutinase, esterase, redox enzymes, or a combination thereof. For example, a debranching enzyme, such as an isoamylase (EC 3.2.1.68), may be added in effective amounts well known to the person skilled in the art. A pullulanase (EC 3.2.1.41), e.g., PROMOZYME.RTM., is also suitable. Pullulanase typically is added at 100 U/kg ds. Further suitable enzymes include proteases, such as fungal and bacterial proteases. Fungal proteases include those obtained from Aspergillus, such as A. niger, A. awamori, A. oryzae; Mucor (e.g., M. miehei); Rhizopus; and Trichoderma.
[0427] .beta.-Amylases (EC 3.2.1.2) are exo-acting maltogenic amylases, which catalyze the hydrolysis of 1,4-.alpha.-glucosidic linkages into amylopectin and related glucose polymers, thereby releasing maltose. .beta.-Amylases have been isolated from various plants and microorganisms. See Fogarty et al. (1979) in Progress in Industrial Microbiology, Vol. 15, pp. 112-115. These (3-Amylases have optimum temperatures in the range from 40.degree. C. to 65.degree. C. and optimum pH in the range from about 4.5 to about 7.0. Contemplated .beta.-amylases include, but are not limited to, .beta.-amylases from barley SPEZYME.RTM. BBA 1500, SPEZYME.RTM. DBA, OPTIMALT.TM. ME, OPTIMALT.TM. BBA (Danisco US Inc.); and NOVOZYM.TM. WBA (Novozymes A/S).
[0428] Compositions comprising the present amylases may be aqueous or non-aqueous formulations, granules, powders, gels, slurries, pastes, etc., which may further comprise any one or more of the additional enzymes listed, herein, along with buffers, salts, preservatives, water, co-solvents, surfactants, and the like. Such compositions may work in combination with endogenous enzymes or other ingredients already present in a slurry, water bath, washing machine, food or drink product, etc, for example, endogenous plant (including algal) enzymes, residual enzymes from a prior processing step, and the like.
5. Compositions and Methods for Baking and Food Preparation
[0429] The present invention also relates to a "food composition," including but not limited to a food product, animal feed and/or food/feed additives, comprising an amylase, and methods for preparing such a food composition comprising mixing variant amylase with one or more food ingredients, or uses thereof.
[0430] Furthermore, the present invention relates to the use of an amylase in the preparation of a food composition, wherein the food composition is baked subsequent to the addition of the polypeptide of the invention. As used herein the term "baking composition" means any composition and/or additive prepared in the process of providing a baked food product, including but not limited to bakers flour, a dough, a baking additive and/or a baked product. The food composition or additive may be liquid or solid.
[0431] As used herein, the term "flour" means milled or ground cereal grain. The term "flour" also may mean Sago or tuber products that have been ground or mashed. In some embodiments, flour may also contain components in addition to the milled or mashed cereal or plant matter. An example of an additional component, although not intended to be limiting, is a leavening agent. Cereal grains include wheat, oat, rye, and barley. Tuber products include tapioca flour, cassava flour, and custard powder. The term "flour" also includes ground corn flour, maize-meal, rice flour, whole-meal flour, self-rising flour, tapioca flour, cassava flour, ground rice, enriched flower, and custard powder.
[0432] For the commercial and home use of flour for baking and food production, it is important to maintain an appropriate level of .alpha.-amylase activity in the flour. A level of activity that is too high may result in a product that is sticky and/or doughy and therefore unmarketable. Flour with insufficient .alpha.-amylase activity may not contain enough sugar for proper yeast function, resulting in dry, crumbly bread, or baked products. Accordingly, an amylase, by itself or in combination with another .alpha.-amylase(s), may be added to the flour to augment the level of endogenous .alpha.-amylase activity in flour.
[0433] An amylase can further be added alone or in a combination with other amylases to prevent or retard staling, i.e., crumb firming of baked products. The amount of anti-staling amylase will typically be in the range of 0.01-10 mg of enzyme protein per kg of flour, e.g., 0.5 mg/kg ds. Additional anti-staling amylases that can be used in combination with an amylase include an endo-amylase, e.g., a bacterial endo-amylase from Bacillus. The additional amylase can be another maltogenic .alpha.-amylase (EC 3.2.1.133), e.g., from Bacillus. NOVAMYL.RTM. is an exemplary maltogenic .alpha.-amylase from B. stearothermophilus strain NCIB 11837 and is described in Christophersen et al. (1997) Starch 50: 39-45. Other examples of anti-staling endo-amylases include bacterial .alpha.-amylases derived from Bacillus, such as B. licheniformis or B. amyloliquefaciens. The anti-staling amylase may be an exo-amylase, such as .beta.-amylase, e.g., from plant sources, such as soy bean, or from microbial sources, such as Bacillus.
[0434] The baking composition comprising an amylase further can comprise a phospholipase or enzyme with phospholipase activity. An enzyme with phospholipase activity has an activity that can be measured in Lipase Units (LU). The phospholipase may have A1 or A2 activity to remove fatty acid from the phospholipids, forming a lysophospholipid. It may or may not have lipase activity, i.e., activity on triglyceride substrates. The phospholipase typically has a temperature optimum in the range of 30-90.degree. C., e.g., 30-70.degree. C. The added phospholipases can be of animal origin, for example, from pancreas, e.g., bovine or porcine pancreas, snake venom or bee venom. Alternatively, the phospholipase may be of microbial origin, e.g., from filamentous fungi, yeast or bacteria, for example.
[0435] The phospholipase is added in an amount that improves the softness of the bread during the initial period after baking, particularly the first 24 hours. The amount of phospholipase will typically be in the range of 0.01-10 mg of enzyme protein per kg of flour, e.g., 0.1-5 mg/kg. That is, phospholipase activity generally will be in the range of 20-1000 LU/kg of flour, where a Lipase Unit is defined as the amount of enzyme required to release 1 .mu.mol butyric acid per minute at 30.degree. C., pH 7.0, with gum arabic as emulsifier and tributyrin as substrate.
[0436] Compositions of dough generally comprise wheat meal or wheat flour and/or other types of meal, flour or starch such as corn flour, cornstarch, rye meal, rye flour, oat flour, oatmeal, soy flour, sorghum meal, sorghum flour, potato meal, potato flour or potato starch. The dough may be fresh, frozen or par-baked. The dough can be a leavened dough or a dough to be subjected to leavening. The dough may be leavened in various ways, such as by adding chemical leavening agents, e.g., sodium bicarbonate or by adding a leaven, i.e., fermenting dough. Dough also may be leavened by adding a suitable yeast culture, such as a culture of Saccharomyces cerevisiae (baker's yeast), e.g., a commercially available strain of S. cerevisiae.
[0437] The dough may also comprise other conventional dough ingredients, e.g., proteins, such as milk powder, gluten, and soy; eggs (e.g., whole eggs, egg yolks or egg whites); an oxidant, such as ascorbic acid, potassium bromate, potassium iodate, azodicarbonamide (ADA) or ammonium persulfate; an amino acid such as L-cysteine; a sugar; or a salt, such as sodium chloride, calcium acetate, sodium sulfate or calcium sulfate. The dough further may comprise fat, e.g., triglyceride, such as granulated fat or shortening. The dough further may comprise an emulsifier such as mono- or diglycerides, diacetyl tartaric acid esters of mono- or diglycerides, sugar esters of fatty acids, polyglycerol esters of fatty acids, lactic acid esters of monoglycerides, acetic acid esters of monoglycerides, polyoxyethylene stearates, or lysolecithin. In particular, the dough can be made without addition of emulsifiers.
[0438] The dough product may be any processed dough product, including fried, deep fried, roasted, baked, steamed and boiled doughs, such as steamed bread and rice cakes. In one embodiment, the food product is a bakery product. Typical bakery (baked) products include bread--such as loaves, rolls, buns, bagels, pizza bases etc. pastry, pretzels, tortillas, cakes, cookies, biscuits, crackers etc.
[0439] Optionally, an additional enzyme may be used together with the anti-staling amylase and the phospholipase. The additional enzyme may be a second amylase, such as an amyloglucosidase, a .beta.-amylase, a cyclodextrin glucanotransferase, or the additional enzyme may be a peptidase, in particular an exopeptidase, a transglutaminase, a lipase, a cellulase, a xylanase, a protease, a protein disulfide isomerase, e.g., a protein disulfide isomerase as disclosed in WO 95/00636, for example, a glycosyltransferase, a branching enzyme (1,4-.alpha.-glucan branching enzyme), a 4-.alpha.-glucanotransferase (dextrin glycosyltransferase) or an oxidoreductase, e.g., a peroxidase, a laccase, a glucose oxidase, an amadoriase, a metalloproteinase, a pyranose oxidase, a lipooxygenase, an L-amino acid oxidase or a carbohydrate oxidase. The additional enzyme(s) may be of any origin, including mammalian and plant, and particularly of microbial (bacterial, yeast or fungal) origin and may be obtained by techniques conventionally used in the art.
[0440] The xylanase is typically of microbial origin, e.g., derived from a bacterium or fungus, such as a strain of Aspergillus. Xylanases include PENTOPAN.RTM. and NOVOZYM 384.RTM., for example, which are commercially available xylanase preparations produced from Trichoderma reesei. The amyloglucosidase may be an A. niger amyloglucosidase (such as AMG.RTM.). Other useful amylase products include GRINDAMYL.RTM. A 1000 or A 5000 (Grindsted Products, Denmark) and AMYLASE H.TM. or AMYLASE P.TM. (DSM). The glucose oxidase may be a fungal glucose oxidase, in particular an Aspergillus niger glucose oxidase (such as GLUZYME.RTM.). An exemplary protease is NEUTRASE.RTM..
[0441] The process may be used for any kind of baked product prepared from dough, either of a soft or a crisp character, either of a white, light or dark type. Examples are bread, particularly white, whole-meal or rye bread, typically in the form of loaves or rolls, such as, but not limited to, French baguette-type bread, pita bread, tortillas, cakes, pancakes, biscuits, cookies, pie crusts, crisp bread, steamed bread, pizza and the like.
[0442] An amylase may be used in a pre-mix, comprising flour together with an anti-staling amylase, a phospholipase, and/or a phospholipid. The pre-mix may contain other dough-improving and/or bread-improving additives, e.g., any of the additives, including enzymes, mentioned above. An amylase can be a component of an enzyme preparation comprising an anti-staling amylase and a phospholipase, for use as a baking additive.
[0443] The enzyme preparation is optionally in the form of a granulate or agglomerated powder. The preparation can have a narrow particle size distribution with more than 95% (by weight) of the particles in the range from 25 to 500 .mu.m. Granulates and agglomerated powders may be prepared by conventional methods, e.g., by spraying an amylase onto a carrier in a fluid-bed granulator. The carrier may consist of particulate cores having a suitable particle size. The carrier may be soluble or insoluble, e.g., a salt (such as NaCl or sodium sulfate), a sugar (such as sucrose or lactose), a sugar alcohol (such as sorbitol), starch, rice, corn grits, or soy.
[0444] Enveloped particles, i.e., .alpha.-amylase particles, can comprise an amylase. To prepare enveloped .alpha.-amylase particles, the enzyme is contacted with a food grade lipid in sufficient quantity to suspend all of the .alpha.-amylase particles. Food grade lipids, as used herein, may be any naturally organic compound that is insoluble in water but is soluble in non-polar organic solvents such as hydrocarbon or diethyl ether. Suitable food grade lipids include, but are not limited to, triglycerides either in the form of fats or oils that are either saturated or unsaturated. Examples of fatty acids and combinations thereof which make up the saturated triglycerides include, but are not limited to, butyric (derived from milk fat), palmitic (derived from animal and plant fat), and/or stearic (derived from animal and plant fat). Examples of fatty acids and combinations thereof which make up the unsaturated triglycerides include, but are not limited to, palmitoleic (derived from animal and plant fat), oleic (derived from animal and plant fat), linoleic (derived from plant oils), and/or linolenic (derived from linseed oil). Other suitable food grade lipids include, but are not limited to, monoglycerides and diglycerides derived from the triglycerides discussed above, phospholipids and glycolipids.
[0445] The food grade lipid, particularly in the liquid form, is contacted with a powdered form of the .alpha.-amylase particles in such a fashion that the lipid material covers at least a portion of the surface of at least a majority, e.g., 100% of the .alpha.-amylase particles. Thus, each .alpha.-amylase particle is individually enveloped in a lipid. For example, all or substantially all of the .alpha.-amylase particles are provided with a thin, continuous, enveloping film of lipid. This can be accomplished by first pouring a quantity of lipid into a container, and then slurrying the .alpha.-amylase particles so that the lipid thoroughly wets the surface of each .alpha.-amylase particle. After a short period of stirring, the enveloped .alpha.-amylase particles, carrying a substantial amount of the lipids on their surfaces, are recovered. The thickness of the coating so applied to the particles of .alpha.-amylase can be controlled by selection of the type of lipid used and by repeating the operation in order to build up a thicker film, when desired.
[0446] The storing, handling and incorporation of the loaded delivery vehicle can be accomplished by means of a packaged mix. The packaged mix can comprise the enveloped .alpha.-amylase. However, the packaged mix may further contain additional ingredients as required by the manufacturer or baker. After the enveloped .alpha.-amylase has been incorporated into the dough, the baker continues through the normal production process for that product.
[0447] The advantages of enveloping the .alpha.-amylase particles are two-fold. First, the food grade lipid protects the enzyme from thermal denaturation during the baking process for those enzymes that are heat labile. Consequently, while the .alpha.-amylase is stabilized and protected during the proving and baking stages, it is released from the protective coating in the final baked good product, where it hydrolyzes the glucosidic linkages in polyglucans. The loaded delivery vehicle also provides a sustained release of the active enzyme into the baked good. That is, following the baking process, active .alpha.-amylase is continually released from the protective coating at a rate that counteracts, and therefore reduces the rate of, staling mechanisms.
[0448] In general, the amount of lipid applied to the .alpha.-amylase particles can vary from a few percent of the total weight of the .alpha.-amylase to many times that weight, depending upon the nature of the lipid, the manner in which it is applied to the .alpha.-amylase particles, the composition of the dough mixture to be treated, and the severity of the dough-mixing operation involved.
[0449] The loaded delivery vehicle, i.e., the lipid-enveloped enzyme, is added to the ingredients used to prepare a baked good in an effective amount to extend the shelf-life of the baked good. The baker computes the amount of enveloped .alpha.-amylase, prepared as discussed above, that will be required to achieve the desired anti-staling effect. The amount of the enveloped .alpha.-amylase required is calculated based on the concentration of enzyme enveloped and on the proportion of .alpha.-amylase to flour specified. A wide range of concentrations has been found to be effective, although, as has been discussed, observable improvements in anti-staling do not correspond linearly with the .alpha.-amylase concentration, but above certain minimal levels, large increases in .alpha.-amylase concentration produce little additional improvement. The .alpha.-amylase concentration actually used in a particular bakery production could be much higher than the minimum necessary to provide the baker with some insurance against inadvertent under-measurement errors by the baker. The lower limit of enzyme concentration is determined by the minimum anti-staling effect the baker wishes to achieve.
[0450] A method of preparing a baked good may comprise: a) preparing lipid-coated .alpha.-amylase particles, where substantially all of the .alpha.-amylase particles are coated; b) mixing a dough containing flour; c) adding the lipid-coated .alpha.-amylase to the dough before the mixing is complete and terminating the mixing before the lipid coating is removed from the .alpha.-amylase; d) proofing the dough; and e) baking the dough to provide the baked good, where the .alpha.-amylase is inactive during the mixing, proofing and baking stages and is active in the baked good.
[0451] The enveloped .alpha.-amylase can be added to the dough during the mix cycle, e.g., near the end of the mix cycle. The enveloped .alpha.-amylase is added at a point in the mixing stage that allows sufficient distribution of the enveloped .alpha.-amylase throughout the dough; however, the mixing stage is terminated before the protective coating becomes stripped from the .alpha.-amylase particle(s). Depending on the type and volume of dough, and mixer action and speed, anywhere from one to six minutes or more might be required to mix the enveloped .alpha.-amylase into the dough, but two to four minutes is average. Thus, several variables may determine the precise procedure. First, the quantity of enveloped .alpha.-amylase should have a total volume sufficient to allow the enveloped .alpha.-amylase to be spread throughout the dough mix. If the preparation of enveloped .alpha.-amylase is highly concentrated, additional oil may need to be added to the pre-mix before the enveloped .alpha.-amylase is added to the dough. Recipes and production processes may require specific modifications; however, good results can generally be achieved when 25% of the oil specified in a bread dough formula is held out of the dough and is used as a carrier for a concentrated enveloped .alpha.-amylase when added near the end of the mix cycle. In bread or other baked goods, particularly those having a low fat content, e.g., French-style breads, an enveloped .alpha.-amylase mixture of approximately 1% of the dry flour weight is sufficient to admix the enveloped .alpha.-amylase properly with the dough. The range of suitable percentages is wide and depends on the formula, finished product, and production methodology requirements of the individual baker. Second, the enveloped .alpha.-amylase suspension should be added to the mix with sufficient time for complete mixture into the dough, but not for such a time that excessive mechanical action strips the protective lipid coating from the enveloped .alpha.-amylase particles.
[0452] In a further aspect of the invention, the food composition is an oil, meat, lard, composition comprising an amylase. In this context the term "[oil/meat/lard] composition" means any composition, based on, made from and/or containing oil, meat or lard, respectively. Another aspect the invention relates to a method of preparing an oil or meat or lard composition and/or additive comprising an amylase, comprising mixing the polypeptide of the invention with a oil/meat/lard composition and/or additive ingredients.
[0453] In a further aspect of the invention, the food composition is an animal feed composition, animal feed additive and/or pet food comprising an amylase and variants thereof. The present invention further relates to a method for preparing such an animal feed composition, animal feed additive composition and/or pet food comprising mixing an amylase and variants thereof with one or more animal feed ingredients and/or animal feed additive ingredients and/or pet food ingredients. Furthermore, the present invention relates to the use of an amylase in the preparation of an animal feed composition and/or animal feed additive composition and/or pet food.
[0454] The term "animal" includes all non-ruminant and ruminant animals. In a particular embodiment, the animal is a non-ruminant animal, such as a horse and a mono-gastric animal. Examples of mono-gastric animals include, but are not limited to, pigs and swine, such as piglets, growing pigs, sows; poultry such as turkeys, ducks, chicken, broiler chicks, layers; fish such as salmon, trout, tilapia, catfish and carps; and crustaceans such as shrimps and prawns. In a further embodiment the animal is a ruminant animal including, but not limited to, cattle, young calves, goats, sheep, giraffes, bison, moose, elk, yaks, water buffalo, deer, camels, alpacas, llamas, antelope, pronghorn and nilgai.
[0455] In the present context, it is intended that the term "pet food" is understood to mean a food for a household animal such as, but not limited to dogs, cats, gerbils, hamsters, chinchillas, fancy rats, guinea pigs; avian pets, such as canaries, parakeets, and parrots; reptile pets, such as turtles, lizards and snakes; and aquatic pets, such as tropical fish and frogs.
[0456] The terms "animal feed composition," "feedstuff" and "fodder" are used interchangeably and may comprise one or more feed materials selected from the group comprising a) cereals, such as small grains (e.g., wheat, barley, rye, oats and combinations thereof) and/or large grains such as maize or sorghum; b) by products from cereals, such as corn gluten meal, Distillers Dried Grain Solubles (DDGS) (particularly corn based Distillers Dried Grain Solubles (cDDGS), wheat bran, wheat middlings, wheat shorts, rice bran, rice hulls, oat hulls, palm kernel, and citrus pulp; c) protein obtained from sources such as soya, sunflower, peanut, lupin, peas, fava beans, cotton, canola, fish meal, dried plasma protein, meat and bone meal, potato protein, whey, copra, sesame; d) oils and fats obtained from vegetable and animal sources; e) minerals and vitamins.
6. Textile Desizing Compositions and Use
[0457] Also contemplated are compositions and methods of treating fabrics (e.g., to desize a textile) using an amylase. Fabric-treating methods are well known in the art (see, e.g., U.S. Pat. No. 6,077,316). For example, the feel and appearance of a fabric can be improved by a method comprising contacting the fabric with an amylase in a solution. The fabric can be treated with the solution under pressure.
[0458] An amylase can be applied during or after the weaving of a textile, or during the desizing stage, or one or more additional fabric processing steps. During the weaving of textiles, the threads are exposed to considerable mechanical strain. Prior to weaving on mechanical looms, warp yarns are often coated with sizing starch or starch derivatives to increase their tensile strength and to prevent breaking. An amylase can be applied during or after the weaving to remove these sizing starch or starch derivatives. After weaving, an amylase can be used to remove the size coating before further processing the fabric to ensure a homogeneous and wash-proof result.
[0459] An amylase can be used alone or with other desizing chemical reagents and/or desizing enzymes to desize fabrics, including cotton-containing fabrics, as detergent additives, e.g., in aqueous compositions. An amylase also can be used in compositions and methods for producing a stonewashed look on indigo-dyed denim fabric and garments. For the manufacture of clothes, the fabric can be cut and sewn into clothes or garments, which are afterwards finished. In particular, for the manufacture of denim jeans, different enzymatic finishing methods have been developed. The finishing of denim garment normally is initiated with an enzymatic desizing step, during which garments are subjected to the action of amylolytic enzymes to provide softness to the fabric and make the cotton more accessible to the subsequent enzymatic finishing steps. An amylase can be used in methods of finishing denim garments (e.g., a "bio-stoning process"), enzymatic desizing and providing softness to fabrics, and/or finishing process.
7. Cleaning Compositions
[0460] An aspect of the present compositions and methods is a cleaning composition that includes an amylase as a component. An amylase polypeptide can be used as a component in detergent compositions for, e.g., hand washing, laundry washing, dishwashing, and other hard-surface cleaning. Such compositions include heavy duty liquid (HDL), heavy duty dry (HDD), and hand (manual) laundry detergent compositions, including unit dose format laundry detergent compositions, and automatic dishwashing (ADW) and hand (manual) dishwashing compositions, including unit dose format dishwashing compositions.
7.1. Overview
[0461] Preferably, an amylase is incorporated into detergents at or near a concentration conventionally used for amylase in detergents. For example, an amylase polypeptide may be added in amount corresponding to 0.00001-1 mg (calculated as pure enzyme protein) of amylase per liter of wash/dishwash liquor. Exemplary formulations are provided herein, as exemplified by the following:
[0462] An amylase polypeptide may be a component of a detergent composition, as the only enzyme or with other enzymes including other amylolytic enzymes. As such, it may be included in the detergent composition in the form of a non-dusting granulate, a stabilized liquid, or a protected enzyme. Non-dusting granulates may be produced, e.g., as disclosed in U.S. Pat. Nos. 4,106,991 and 4,661,452 and may optionally be coated by methods known in the art. Examples of waxy coating materials are poly(ethylene oxide) products (polyethyleneglycol, PEG) with mean molar weights of 1,000 to 20,000; ethoxylated nonylphenols having from 16 to 50 ethylene oxide units; ethoxylated fatty alcohols in which the alcohol contains from 12 to 20 carbon atoms and in which there are 15 to 80 ethylene oxide units; fatty alcohols; fatty acids; and mono- and di- and triglycerides of fatty acids. Examples of film-forming coating materials suitable for application by fluid bed techniques are given in, for example, GB 1483591. Liquid enzyme preparations may, for instance, be stabilized by adding a polyol such as propylene glycol, a sugar or sugar alcohol, lactic acid or boric acid according to established methods. Other enzyme stabilizers are known in the art. Protected enzymes may be prepared according to the method disclosed in for example EP 238 216. Polyols have long been recognized as stabilizers of proteins, as well as improving protein solubility.
[0463] The detergent composition may be in any useful form, e.g., as powders, granules, pastes, bars, or liquid. A liquid detergent may be aqueous, typically containing up to about 70% of water and 0% to about 30% of organic solvent. It may also be in the form of a compact gel type containing only about 30% water.
[0464] The detergent composition comprises one or more surfactants, each of which may be anionic, nonionic, cationic, or zwitterionic. The detergent will usually contain 0% to about 50% of anionic surfactant, such as linear alkylbenzenesulfonate (LAS); .alpha.-olefinsulfonate (AOS); alkyl sulfate (fatty alcohol sulfate) (AS); alcohol ethoxysulfate (AEOS or AES); secondary alkanesulfonates (SAS); .alpha.-sulfo fatty acid methyl esters; alkyl- or alkenylsuccinic acid; or soap. The composition may also contain 0% to about 40% of nonionic surfactant such as alcohol ethoxylate (AEO or AE), carboxylated alcohol ethoxylates, nonylphenol ethoxylate, alkylpolyglycoside, alkyldimethylamineoxide, ethoxylated fatty acid monoethanolamide, fatty acid monoethanolamide, or polyhydroxy alkyl fatty acid amide (as described for example in WO 92/06154).
[0465] The detergent composition may additionally comprise one or more other enzymes, such as proteases, another amylolytic enzyme, cutinase, lipase, cellulase, pectate lyase, perhydrolase, xylanase, peroxidase, and/or laccase in any combination.
[0466] The detergent may contain about 1% to about 65% of a detergent builder or complexing agent such as zeolite, diphosphate, triphosphate, phosphonate, citrate, nitrilotriacetic acid (NTA), ethylenediaminetetraacetic acid (EDTA), diethylenetriaminepentaacetic acid (DTMPA), alkyl- or alkenylsuccinic acid, soluble silicates or layered silicates (e.g., SKS-6 from Hoechst). The detergent may also be unbuilt, i.e. essentially free of detergent builder. The enzymes can be used in any composition compatible with the stability of the enzyme. Enzymes generally can be protected against deleterious components by known forms of encapsulation, for example, by granulation or sequestration in hydro gels. Enzymes, and specifically amylases, either with or without starch binding domains, can be used in a variety of compositions including laundry and dishwashing applications, surface cleaners, as well as in compositions for ethanol production from starch or biomass.
[0467] The detergent may comprise one or more polymers. Examples include carboxymethylcellulose (CMC), poly(vinylpyrrolidone) (PVP), polyethyleneglycol (PEG), poly(vinyl alcohol) (PVA), polycarboxylates such as polyacrylates, maleic/acrylic acid copolymers and lauryl methacrylate/acrylic acid copolymers.
[0468] The detergent may contain a bleaching system, which may comprise a H2O2 source such as perborate or percarbonate, which may be combined with a peracid-forming bleach activator such as tetraacetylethylenediamine (TAED) or nonanoyloxybenzenesulfonate (NOBS).
[0469] Alternatively, the bleaching system may comprise peroxyacids (e.g., the amide, imide, or sulfone type peroxyacids). The bleaching system can also be an enzymatic bleaching system, for example, perhydrolase, such as that described in International PCT Application WO 2005/056783.
[0470] The enzymes of the detergent composition may be stabilized using conventional stabilizing agents, e.g., a polyol such as propylene glycol or glycerol; a sugar or sugar alcohol; lactic acid; boric acid or a boric acid derivative such as, e.g., an aromatic borate ester; and the composition may be formulated as described in, e.g., WO 92/19709 and WO 92/19708.
[0471] The detergent may also contain other conventional detergent ingredients such as e.g., fabric conditioners including clays, foam boosters, suds suppressors, anti-corrosion agents, soil-suspending agents, anti-soil redeposition agents, dyes, bactericides, tarnish inhibitors, optical brighteners, or perfumes.
[0472] The pH (measured in aqueous solution at use concentration) is usually neutral or alkaline, e.g., pH about 7.0 to about 11.0.
[0473] Particular forms of detergent compositions for inclusion of the present .alpha.-amylase are described, below. Many of these composition can be provided in unit dose format for ease of use. Unit dose formulations and packaging are described in, for example, US20090209445A1, US20100081598A1, U.S. Pat. No. 7,001,878B2, EP1504994B1, WO2001085888A2, WO2003089562A1, WO2009098659A1, WO2009098660A1, WO2009112992A1, WO2009124160A1, WO2009152031A1, WO2010059483A1, WO2010088112A1, WO2010090915A1, WO2010135238A1, WO2011094687A1, WO2011094690A1, WO2011127102A1, WO2011163428A1, WO2008000567A1, WO2006045391A1, WO2006007911A1, WO2012027404A1, EP1740690B1, WO2012059336A1, U.S. Pat. No. 6,730,646B1, WO2008087426A1, WO2010116139A1, and WO2012104613A1.
7.2. Heavy Duty Liquid (HDL) Laundry Detergent Composition
[0474] Exemplary HDL laundry detergent compositions includes a detersive surfactant (10%-40% wt/wt), including an anionic detersive surfactant (selected from a group of linear or branched or random chain, substituted or unsubstituted alkyl sulphates, alkyl sulphonates, alkyl alkoxylated sulphate, alkyl phosphates, alkyl phosphonates, alkyl carboxylates, and/or mixtures thereof), and optionally non-ionic surfactant (selected from a group of linear or branched or random chain, substituted or unsubstituted alkyl alkoxylated alcohol, for example a C8-C18 alkyl ethoxylated alcohol and/or C6-C12 alkyl phenol alkoxylates), wherein the weight ratio of anionic detersive surfactant (with a hydrophilic index (HIc) of from 6.0 to 9) to non-ionic detersive surfactant is greater than 1:1. Suitable detersive surfactants also include cationic detersive surfactants (selected from a group of alkyl pyridinium compounds, alkyl quarternary ammonium compounds, alkyl quarternary phosphonium compounds, alkyl ternary sulphonium compounds, and/or mixtures thereof); zwitterionic and/or amphoteric detersive surfactants (selected from a group of alkanolamine sulpho-betaines); ampholytic surfactants; semi-polar non-ionic surfactants and mixtures thereof.
[0475] The composition may optionally include, a surfactancy boosting polymer consisting of amphiphilic alkoxylated grease cleaning polymers (selected from a group of alkoxylated polymers having branched hydrophilic and hydrophobic properties, such as alkoxylated polyalkylenimines in the range of 0.05 wt %-10 wt %) and/or random graft polymers (typically comprising of hydrophilic backbone comprising monomers selected from the group consisting of: unsaturated C1-C6 carboxylic acids, ethers, alcohols, aldehydes, ketones, esters, sugar units, alkoxy units, maleic anhydride, saturated polyalcohols such as glycerol, and mixtures thereof and hydrophobic side chain(s) selected from the group consisting of: C4-C25 alkyl group, polypropylene, polybutylene, vinyl ester of a saturated C1-C6 mono-carboxylic acid, C1-C6 alkyl ester of acrylic or methacrylic acid, and mixtures thereof.
[0476] The composition may include additional polymers such as soil release polymers (include anionically end-capped polyesters, for example SRP1, polymers comprising at least one monomer unit selected from saccharide, dicarboxylic acid, polyol and combinations thereof, in random or block configuration, ethylene terephthalate-based polymers and co-polymers thereof in random or block configuration, for example Repel-o-tex SF, SF-2 and SRP6, Texcare SRA100, SRA300, SRN100, SRN170, SRN240, SRN300 and SRN325, Marloquest SL), anti-redeposition polymers (0.1 wt % to 10 wt %, include carboxylate polymers, such as polymers comprising at least one monomer selected from acrylic acid, maleic acid (or maleic anhydride), fumaric acid, itaconic acid, aconitic acid, mesaconic acid, citraconic acid, methylenemalonic acid, and any mixture thereof, vinylpyrrolidone homopolymer, and/or polyethylene glycol, molecular weight in the range of from 500 to 100,000 Da); cellulosic polymer (including those selected from alkyl cellulose, alkyl alkoxyalkyl cellulose, carboxyalkyl cellulose, alkyl carboxyalkyl cellulose examples of which include carboxymethyl cellulose, methyl cellulose, methyl hydroxyethyl cellulose, methyl carboxymethyl cellulose, and mixtures thereof) and polymeric carboxylate (such as maleate/acrylate random copolymer or polyacrylate homopolymer).
[0477] The composition may further include saturated or unsaturated fatty acid, preferably saturated or unsaturated C12-C24 fatty acid (0 wt % to 10 wt %); deposition aids (examples for which include polysaccharides, preferably cellulosic polymers, poly diallyl dimethyl ammonium halides (DADMAC), and co-polymers of DAD MAC with vinyl pyrrolidone, acrylamides, imidazoles, imidazolinium halides, and mixtures thereof, in random or block configuration, cationic guar gum, cationic cellulose such as cationic hydoxyethyl cellulose, cationic starch, cationic polyacylamides, and mixtures thereof.
[0478] The composition may further include dye transfer inhibiting agents, examples of which include manganese phthalocyanine, peroxidases, polyvinylpyrrolidone polymers, polyamine N-oxide polymers, copolymers of N-vinylpyrrolidone and N-vinylimidazole, polyvinyloxazolidones and polyvinylimidazoles and/or mixtures thereof; chelating agents, examples of which include ethylene-diamine-tetraacetic acid (EDTA), diethylene triamine penta methylene phosphonic acid (DTPMP), hydroxy-ethane diphosphonic acid (HEDP), ethylenediamine N,N'-disuccinic acid (EDDS), methyl glycine diacetic acid (MGDA), diethylene triamine penta acetic acid (DTPA), propylene diamine tetracetic acid (PDT A), 2-hydroxypyridine-N-oxide (HPNO), or methyl glycine diacetic acid (MGDA), glutamic acid N,N-diacetic acid (N,N-dicarboxymethyl glutamic acid tetrasodium salt (GLDA), nitrilotriacetic acid (NTA), 4,5-dihydroxy-m-benzenedisulfonic acid, citric acid and any salts thereof, N-hydroxyethylethylenediaminetri-acetic acid (HEDTA), triethylenetetraaminehexaacetic acid (TTHA), N-hydroxyethyliminodiacetic acid (HEIDA), dihydroxyethylglycine (DHEG), ethylenediaminetetrapropionic acid (EDTP), and derivatives thereof.
[0479] The composition preferably included enzymes (generally about 0.01 wt % active enzyme to 0.03 wt % active enzyme) selected from proteases, amylases, lipases, cellulases, choline oxidases, peroxidases/oxidases, pectate lyases, mannanases, cutinases, laccases, phospholipases, lysophospholipases, acyltransferases, perhydrolases, arylesterases, and any mixture thereof. The composition may include an enzyme stabilizer (examples of which include polyols such as propylene glycol or glycerol, sugar or sugar alcohol, lactic acid, reversible protease inhibitor, boric acid, or a boric acid derivative, e.g., an aromatic borate ester, or a phenyl boronic acid derivative such as 4-formylphenyl boronic acid).
[0480] The composition optionally include silicone or fatty-acid based suds suppressors; hueing dyes, calcium and magnesium cations, visual signaling ingredients, anti-foam (0.001 wt % to about 4.0 wt %), and/or structurant/thickener (0.01 wt % to 5 wt %, selected from the group consisting of diglycerides and triglycerides, ethylene glycol distearate, microcrystalline cellulose, cellulose based materials, microfiber cellulose, biopolymers, xanthan gum, gellan gum, and mixtures thereof).
[0481] The composition can be any liquid form, for example a liquid or gel form, or any combination thereof. The composition may be in any unit dose form, for example a pouch.
7.3. Heavy Duty Dry/Solid (HDD) Laundry Detergent Composition
[0482] Exemplary HDD laundry detergent compositions includes a detersive surfactant, including anionic detersive surfactants (e.g., linear or branched or random chain, substituted or unsubstituted alkyl sulphates, alkyl sulphonates, alkyl alkoxylated sulphate, alkyl phosphates, alkyl phosphonates, alkyl carboxylates and/or mixtures thereof), non-ionic detersive surfactant (e.g., linear or branched or random chain, substituted or unsubstituted C8-C18 alkyl ethoxylates, and/or C6-C12 alkyl phenol alkoxylates), cationic detersive surfactants (e.g., alkyl pyridinium compounds, alkyl quaternary ammonium compounds, alkyl quaternary phosphonium compounds, alkyl ternary sulphonium compounds, and mixtures thereof), zwitterionic and/or amphoteric detersive surfactants (e.g., alkanolamine sulpho-betaines), ampholytic surfactants, semi-polar non-ionic surfactants, and mixtures thereof; builders including phosphate free builders (for example zeolite builders examples which include zeolite A, zeolite X, zeolite P and zeolite MAP in the range of 0 wt % to less than 10 wt %), phosphate builders (for example sodium tri-polyphosphate in the range of 0 wt % to less than 10 wt %), citric acid, citrate salts and nitrilotriacetic acid, silicate salt (e.g., sodium or potassium silicate or sodium meta-silicate in the range of 0 wt % to less than 10 wt %, or layered silicate (SKS-6)); carbonate salt (e.g., sodium carbonate and/or sodium bicarbonate in the range of 0 wt % to less than 80 wt %); and bleaching agents including photobleaches (e.g., sulfonated zinc phthalocyanines, sulfonated aluminum phthalocyanines, xanthenes dyes, and mixtures thereof) hydrophobic or hydrophilic bleach activators (e.g., dodecanoyl oxybenzene sulfonate, decanoyl oxybenzene sulfonate, decanoyl oxybenzoic acid or salts thereof, 3,5,5-trimethy hexanoyl oxybenzene sulfonate, tetraacetyl ethylene diamine-TAED, nonanoyloxybenzene sulfonate-NOBS, nitrile quats, and mixtures thereof), sources of hydrogen peroxide (e.g., inorganic perhydrate salts examples of which include mono or tetra hydrate sodium salt of perborate, percarbonate, persulfate, perphosphate, or persilicate), preformed hydrophilic and/or hydrophobic peracids (e.g., percarboxylic acids and salts, percarbonic acids and salts, perimidic acids and salts, peroxymonosulfuric acids and salts, and mixtures thereof), and/or bleach catalysts (e.g., imine bleach boosters (examples of which include iminium cations and polyions), iminium zwitterions, modified amines, modified amine oxides, N-sulphonyl imines, N-phosphonyl imines, N-acyl imines, thiadiazole dioxides, perfluoroimines, cyclic sugar ketones, and mixtures thereof, and metal-containing bleach catalysts (e.g., copper, iron, titanium, ruthenium, tungsten, molybdenum, or manganese cations along with an auxiliary metal cations such as zinc or aluminum and a sequestrate such as ethylenediaminetetraacetic acid, ethylenediaminetetra(methylenephosphonic acid), and water-soluble salts thereof).
[0483] The composition preferably includes enzymes, e.g., proteases, amylases, lipases, cellulases, choline oxidases, peroxidases/oxidases, pectate lyases, mannanases, cutinases, laccases, phospholipases, lysophospholipases, acyltransferase, perhydrolase, arylesterase, and any mixture thereof.
[0484] The composition may optionally include additional detergent ingredients including perfume microcapsules, starch encapsulated perfume accord, hueing agents, additional polymers, including fabric integrity and cationic polymers, dye-lock ingredients, fabric-softening agents, brighteners (for example C.I. Fluorescent brighteners), flocculating agents, chelating agents, alkoxylated polyamines, fabric deposition aids, and/or cyclodextrin.
7.4. Automatic Dishwashing (ADW) Detergent Composition
[0485] Exemplary ADW detergent composition includes non-ionic surfactants, including ethoxylated non-ionic surfactants, alcohol alkoxylated surfactants, epoxy-capped poly(oxyalkylated) alcohols, or amine oxide surfactants present in amounts from 0 to 10% by weight; builders in the range of 5-60% including phosphate builders (e.g., mono-phosphates, di-phosphates, tri-polyphosphates, other oligomeric-poylphosphates, sodium tripolyphosphate-STPP) and phosphate-free builders (e.g., amino acid-based compounds including methyl-glycine-diacetic acid (MGDA) and salts and derivatives thereof, glutamic-N,N-diacetic acid (GLDA) and salts and derivatives thereof, iminodisuccinic acid (IDS) and salts and derivatives thereof, carboxy methyl inulin and salts and derivatives thereof, nitrilotriacetic acid (NTA), diethylene triamine penta acetic acid (DTPA), B-alaninediacetic acid (B-ADA) and their salts, homopolymers and copolymers of poly-carboxylic acids and their partially or completely neutralized salts, monomeric polycarboxylic acids and hydroxycarboxylic acids and their salts in the range of 0.5% to 50% by weight; sulfonated/carboxylated polymers in the range of about 0.1% to about 50% by weight to provide dimensional stability; drying aids in the range of about 0.1% to about 10% by weight (e.g., polyesters, especially anionic polyesters, optionally together with further monomers with 3 to 6 functionalities--typically acid, alcohol or ester functionalities which are conducive to polycondensation, polycarbonate-, polyurethane- and/or polyurea-polyorganosiloxane compounds or precursor compounds, thereof, particularly of the reactive cyclic carbonate and urea type); silicates in the range from about 1% to about 20% by weight (including sodium or potassium silicates for example sodium disilicate, sodium meta-silicate and crystalline phyllosilicates); inorganic bleach (e.g., perhydrate salts such as perborate, percarbonate, perphosphate, persulfate and persilicate salts) and organic bleach (e.g., organic peroxyacids, including diacyl and tetraacylperoxides, especially diperoxydodecanedioc acid, diperoxytetradecanedioc acid, and diperoxyhexadecanedioc acid); bleach activators (i.e., organic peracid precursors in the range from about 0.1% to about 10% by weight); bleach catalysts (e.g., manganese triazacyclononane and related complexes, Co, Cu, Mn, and Fe bispyridylamine and related complexes, and pentamine acetate cobalt(III) and related complexes); metal care agents in the range from about 0.1% to 5% by weight (e.g., benzatriazoles, metal salts and complexes, and/or silicates); enzymes in the range from about 0.01 to 5.0 mg of active enzyme per gram of automatic dishwashing detergent composition (e.g., proteases, amylases, lipases, cellulases, choline oxidases, peroxidases/oxidases, pectate lyases, mannanases, cutinases, laccases, phospholipases, lysophospholipases, acyltransferase, perhydrolase, arylesterase, and mixtures thereof); and enzyme stabilizer components (e.g., oligosaccharides, polysaccharides, and inorganic divalent metal salts).
7.5. Additional Detergent Compositions
[0486] Additional exemplary detergent formulations to which the present amylase can be added are described, below, in the numbered paragraphs.
[0487] 1) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising linear alkylbenzenesulfonate (calculated as acid) about 7% to about 12%; alcohol ethoxysulfate (e.g., C12-18 alcohol, 1-2 ethylene oxide (EO)) or alkyl sulfate (e.g., C16-18) about 1% to about 4%; alcohol ethoxylate (e.g., C14-15 alcohol, 7 EO) about 5% to about 9%; sodium carbonate (e.g., Na2CO3) about 14% to about 20%; soluble silicate (e.g., Na2O, 2SiO2) about 2 to about 6%; zeolite (e.g., NaAlSiO4) about 15% to about 22%; sodium sulfate (e.g., Na2SO4) 0% to about 6%; sodium citrate/citric acid (e.g., C6H5Na3O7/C6H8O7) about 0% to about 15%; sodium perborate (e.g., NaBO3H2O) about 11% to about 18%; TAED about 2% to about 6%; carboxymethylcellulose (CMC) and 0% to about 2%; polymers (e.g., maleic/acrylic acid, copolymer, PVP, PEG) 0-3%; enzymes (calculated as pure enzyme) 0.0001-0.1% protein; and minor ingredients (e.g., suds suppressors, perfumes, optical brightener, photobleach) 0-5%.
[0488] 2) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising linear alkylbenzenesulfonate (calculated as acid) about 6% to about 11%; alcohol ethoxysulfate (e.g., C12-18 alcohol, 1-2 EO) or alkyl sulfate (e.g., C16-18) about 1% to about 3%; alcohol ethoxylate (e.g., C14-15 alcohol, 7 EO) about 5% to about 9%; sodium carbonate (e.g., Na2CO3) about 15% to about 21%; soluble silicate (e.g., Na2O, 2SiO2) about 1% to about 4%; zeolite (e.g., NaAlSiO4) about 24% to about 34%; sodium sulfate (e.g., Na2SO4) about 4% to about 10%; sodium citrate/citric acid (e.g., C6H5Na3O7/C6H8O7) 0% to about 15%; carboxymethylcellulose (CMC) 0% to about 2%; polymers (e.g., maleic/acrylic acid copolymer, PVP, PEG) 1-6%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; minor ingredients (e.g., suds suppressors, perfume) 0-5%.
[0489] 3) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising linear alkylbenzenesulfonate (calculated as acid) about 5% to about 9%; alcohol ethoxylate (e.g., C12-15 alcohol, 7 EO) about 7% to about 14%; Soap as fatty acid (e.g., C16-22 fatty acid) about 1 to about 3%; sodium carbonate (as Na2CO3) about 10% to about 17%; soluble silicate (e.g., Na2O, 2SiO2) about 3% to about 9%; zeolite (as NaAlSiO4) about 23% to about 33%; sodium sulfate (e.g., Na2SO4) 0% to about 4%; sodium perborate (e.g., NaBO3H2O) about 8% to about 16%; TAED about 2% to about 8%; phosphonate (e.g., EDTMPA) 0% to about 1%; carboxymethylcellulose (CMC) 0% to about 2%; polymers (e.g., maleic/acrylic acid copolymer, PVP, PEG) 0-3%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; minor ingredients (e.g., suds suppressors, perfume, optical brightener) 0-5%.
[0490] 4) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising linear alkylbenzenesulfonate (calculated as acid) about 8% to about 12%; alcohol ethoxylate (e.g., C12-15 alcohol, 7 EO) about 10% to about 25%; sodium carbonate (as Na2CO3) about 14% to about 22%; soluble silicate (e.g., Na2O, 2SiO2) about 1% to about 5%; zeolite (e.g., NaAlSiO4) about 25% to about 35%; sodium sulfate (e.g., Na2SO4) 0% to about 10%; carboxymethylcellulose (CMC) 0% to about 2%; polymers (e.g., maleic/acrylic acid copolymer, PVP, PEG) 1-3%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., suds suppressors, perfume) 0-5%.
[0491] 5) An aqueous liquid detergent composition comprising linear alkylbenzenesulfonate (calculated as acid) about 15% to about 21%; alcohol ethoxylate (e.g., C12-15 alcohol, 7 EO or C12-15 alcohol, 5 EO) about 12% to about 18%; soap as fatty acid (e.g., oleic acid) about 3% to about 13%; alkenylsuccinic acid (C12-14) 0% to about 13%; aminoethanol about 8% to about 18%; citric acid about 2% to about 8%; phosphonate 0% to about 3%; polymers (e.g., PVP, PEG) 0% to about 3%; borate (e.g., B407) 0% to about 2%; ethanol 0% to about 3%; propylene glycol about 8% to about 14%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., dispersants, suds suppressors, perfume, optical brightener) 0-5%.
[0492] 6) An aqueous structured liquid detergent composition comprising linear alkylbenzenesulfonate (calculated as acid) about 15% to about 21%; alcohol ethoxylate (e.g., C12-15 alcohol, 7 EO, or C12-15 alcohol, 5 EO) 3-9%; soap as fatty acid (e.g., oleic acid) about 3% to about 10%; zeolite (as NaAlSiO4) about 14% to about 22%; potassium citrate about 9% to about 18%; borate (e.g., B407) 0% to about 2%; carboxymethylcellulose (CMC) 0% to about 2%; polymers (e.g., PEG, PVP) 0% to about 3%; anchoring polymers such as, e.g., lauryl methacrylate/acrylic acid copolymer; molar ratio 25:1, MW 3800) 0% to about 3%; glycerol 0% to about 5%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., dispersants, suds suppressors, perfume, optical brighteners) 0-5%.
[0493] 7) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising fatty alcohol sulfate about 5% to about 10%; ethoxylated fatty acid monoethanolamide about 3% to about 9%; soap as fatty acid 0-3%; sodium carbonate (e.g., Na2CO3) about 5% to about 10%; Soluble silicate (e.g., Na2O, 2SiO2) about 1% to about 4%; zeolite (e.g., NaAlSiO4) about 20% to about 40%; Sodium sulfate (e.g., Na2SO4) about 2% to about 8%; sodium perborate (e.g., NaBO3H2O) about 12% to about 18%; TAED about 2% to about 7%; polymers (e.g., maleic/acrylic acid copolymer, PEG) about 1% to about 5%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., optical brightener, suds suppressors, perfume) 0-5%.
[0494] 8) A detergent composition formulated as a granulate comprising linear alkylbenzenesulfonate (calculated as acid) about 8% to about 14%; ethoxylated fatty acid monoethanolamide about 5% to about 11%; soap as fatty acid 0% to about 3%; sodium carbonate (e.g., Na2CO3) about 4% to about 10%; soluble silicate (Na2O, 2SiO2) about 1% to about 4%; zeolite (e.g., NaAlSiO4) about 30% to about 50%; sodium sulfate (e.g., Na2SO4) about 3% to about 11%; sodium citrate (e.g., C6H5Na3O7) about 5% to about 12%; polymers (e.g., PVP, maleic/acrylic acid copolymer, PEG) about 1% to about 5%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., suds suppressors, perfume) 0-5%.
[0495] 9) A detergent composition formulated as a granulate comprising linear alkylbenzenesulfonate (calculated as acid) about 6% to about 12%; nonionic surfactant about 1% to about 4%; soap as fatty acid about 2% to about 6%; sodium carbonate (e.g., Na2CO3) about 14% to about 22%; zeolite (e.g., NaAlSiO4) about 18% to about 32%; sodium sulfate (e.g., Na2SO4) about 5% to about 20%; sodium citrate (e.g., C6H5Na3O7) about 3% to about 8%; sodium perborate (e.g., NaBO3H2O) about 4% to about 9%; bleach activator (e.g., NOBS or TAED) about 1% to about 5%; carboxymethylcellulose (CMC) 0% to about 2%; polymers (e.g., polycarboxylate or PEG) about 1% to about 5%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., optical brightener, perfume) 0-5%.
[0496] 10) An aqueous liquid detergent composition comprising linear alkylbenzenesulfonate (calculated as acid) about 15% to about 23%; alcohol ethoxysulfate (e.g., C12-15 alcohol, 2-3 EO) about 8% to about 15%; alcohol ethoxylate (e.g., C12-15 alcohol, 7 EO, or C12-15 alcohol, 5 EO) about 3% to about 9%; soap as fatty acid (e.g., lauric acid) 0% to about 3%; aminoethanol about 1% to about 5%; sodium citrate about 5% to about 10%; hydrotrope (e.g., sodium toluensulfonate) about 2% to about 6%; borate (e.g., B407) 0% to about 2%; carboxymethylcellulose 0% to about 1%; ethanol about 1% to about 3%; propylene glycol about 2% to about 5%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., polymers, dispersants, perfume, optical brighteners) 0-5%.
[0497] 11) An aqueous liquid detergent composition comprising linear alkylbenzenesulfonate (calculated as acid) about 20% to about 32%; alcohol ethoxylate (e.g., C12-15 alcohol, 7 EO, or C12-15 alcohol, 5 EO) 6-12%; aminoethanol about 2% to about 6%; citric acid about 8% to about 14%; borate (e.g., B407) about 1% to about 3%; polymer (e.g., maleic/acrylic acid copolymer, anchoring polymer such as, e.g., lauryl methacrylate/acrylic acid copolymer) 0% to about 3%; glycerol about 3% to about 8%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., hydrotropes, dispersants, perfume, optical brighteners) 0-5%.
[0498] 12) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising anionic surfactant (linear alkylbenzenesulfonate, alkyl sulfate, .alpha.-olefinsulfonate, .alpha.-sulfo fatty acid methyl esters, alkanesulfonates, soap) about 25% to about 40%; nonionic surfactant (e.g., alcohol ethoxylate) about 1% to about 10%; sodium carbonate (e.g., Na2CO3) about 8% to about 25%; soluble silicates (e.g., Na2O, 2SiO2) about 5% to about 15%; sodium sulfate (e.g., Na2SO4) 0% to about 5%; zeolite (NaAlSiO4) about 15% to about 28%; sodium perborate (e.g., NaBO3.4H2O) 0% to about 20%; bleach activator (TAED or NOBS) about 0% to about 5%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; minor ingredients (e.g., perfume, optical brighteners) 0-3%.
[0499] 13) Detergent compositions as described in compositions 1)-12) supra, wherein all or part of the linear alkylbenzenesulfonate is replaced by (C12-C18) alkyl sulfate.
[0500] 14) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising (C12-C18) alkyl sulfate about 9% to about 15%; alcohol ethoxylate about 3% to about 6%; polyhydroxy alkyl fatty acid amide about 1% to about 5%; zeolite (e.g., NaAlSiO4) about 10% to about 20%; layered disilicate (e.g., SK56 from Hoechst) about 10% to about 20%; sodium carbonate (e.g., Na2CO3) about 3% to about 12%; soluble silicate (e.g., Na2O, 2SiO2) 0% to about 6%; sodium citrate about 4% to about 8%; sodium percarbonate about 13% to about 22%; TAED about 3% to about 8%; polymers (e.g., polycarboxylates and PVP) 0% to about 5%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., optical brightener, photobleach, perfume, suds suppressors) 0-5%.
[0501] 15) A detergent composition formulated as a granulate having a bulk density of at least 600 g/L comprising (C12-C18) alkyl sulfate about 4% to about 8%; alcohol ethoxylate about 11% to about 15%; soap about 1% to about 4%; zeolite MAP or zeolite A about 35% to about 45%; sodium carbonate (as Na2CO3) about 2% to about 8%; soluble silicate (e.g., Na2O, 2SiO2) 0% to about 4%; sodium percarbonate about 13% to about 22%; TAED 1-8%; carboxymethylcellulose (CMC) 0% to about 3%; polymers (e.g., polycarboxylates and PVP) 0% to about 3%; enzymes (calculated as pure enzyme protein) 0.0001-0.1%; and minor ingredients (e.g., optical brightener, phosphonate, perfume) 0-3%.
[0502] 16) Detergent formulations as described in 1)-15) supra, which contain a stabilized or encapsulated peracid, either as an additional component or as a substitute for already specified bleach systems.
[0503] 17) Detergent compositions as described supra in 1), 3), 7), 9), and 12), wherein perborate is replaced by percarbonate.
[0504] 18) Detergent compositions as described supra in 1), 3), 7), 9), 12), 14), and 15), which additionally contain a manganese catalyst. The manganese catalyst for example is one of the compounds described in "Efficient manganese catalysts for low-temperature bleaching," Nature 369: 637-639 (1994).
[0505] 19) Detergent composition formulated as a non-aqueous detergent liquid comprising a liquid nonionic surfactant such as, e.g., linear alkoxylated primary alcohol, a builder system (e.g., phosphate), an enzyme(s), and alkali. The detergent may also comprise anionic surfactant and/or a bleach system.
[0506] As above, the present amylase polypeptide may be incorporated at a concentration conventionally employed in detergents. It is at present contemplated that, in the detergent composition, the enzyme may be added in an amount corresponding to 0.00001-1.0 mg (calculated as pure enzyme protein) of amylase polypeptide per liter of wash liquor.
[0507] The detergent composition may also contain other conventional detergent ingredients, e.g., deflocculant material, filler material, foam depressors, anti-corrosion agents, soil-suspending agents, sequestering agents, anti-soil redeposition agents, dehydrating agents, dyes, bactericides, fluorescers, thickeners, and perfumes.
[0508] The detergent composition may be formulated as a hand (manual) or machine (automatic) laundry detergent composition, including a laundry additive composition suitable for pre-treatment of stained fabrics and a rinse added fabric softener composition, or be formulated as a detergent composition for use in general household hard surface cleaning operations, or be formulated for manual or automatic dishwashing operations.
[0509] Any of the cleaning compositions described, herein, may include any number of additional enzymes. In general the enzyme(s) should be compatible with the selected detergent, (e.g., with respect to pH-optimum, compatibility with other enzymatic and non-enzymatic ingredients, and the like), and the enzyme(s) should be present in effective amounts. The following enzymes are provided as examples.
[0510] Proteases: Suitable proteases include those of animal, vegetable or microbial origin. Chemically modified or protein engineered mutants are included, as well as naturally processed proteins. The protease may be a serine protease or a metalloprotease, an alkaline microbial protease, a trypsin-like protease, or a chymotrypsin-like protease. Examples of alkaline proteases are subtilisins, especially those derived from Bacillus, e.g., subtilisin Novo, subtilisin Carlsberg, subtilisin 309, subtilisin 147, and subtilisin 168 (see, e.g., WO 89/06279). Additional examples include those mutant proteases described in U.S. Pat. Nos. RE 34,606, 5,955,340, 5,700,676, 6,312,936, and 6,482,628, all of which are incorporated herein by reference. Examples of trypsin-like proteases are trypsin (e.g., of porcine or bovine origin), and Fusarium proteases (see, e.g., WO 89/06270 and WO 94/25583). Examples of useful proteases also include but are not limited to the variants described in WO 92/19729, WO 98/20115, WO 98/20116, and WO 98/34946. Commercially available protease enzymes include but are not limited to: Alcalase.RTM., Savinase.RTM., Primase.TM., Duralase.TM., Esperase.RTM., BLAZE.TM. POLARZYME.RTM., OVOZYME.RTM., KANNASE.RTM., LIQUANASE.RTM., NEUTRASE.RTM., RELASE.RTM., and ESPERASE.RTM. (Novo Nordisk A/S and Novozymes A/S), Maxatase.RTM., Maxacal.TM. Maxapem.TM., Properase.RTM., Purafect.RTM., Purafect OxP.TM., Purafect Prime.TM., FNA.TM., FN2.TM. FN3.TM., OPTICLEAN.RTM., OPTIMASE.RTM., PURAMAX.TM., EXCELLASE.TM., and PURAFAST.TM. (Danisco US Inc./DuPont Industrial Biosciences, Palo Alto, Calif., USA), BLAP.TM. and BLAP.TM. variants (Henkel Kommanditgesellschaft auf Aktien, Duesseldorf, Germany), and KAP (B. alkalophilus subtilisin; Kao Corp., Tokyo, Japan). Another exemplary proteases NprE from Bacillus amyloliquifaciens and ASP from Cellulomonas sp. strain 69B4 (Danisco US Inc./DuPont Industrial Biosciences, Palo Alto, Calif., USA). Various proteases are described in WO95/23221, WO 92/21760, WO 09/149200, WO 09/149144, WO 09/149145, WO 11/072099, WO 10/056640, WO 10/056653, WO 11/140364, WO 12/151534, U.S. Pat. Publ. No. 2008/0090747, and U.S. Pat. Nos. 5,801,039, 5,340,735, 5,500,364, 5,855,625, US RE 34,606, 5,955,340, 5,700,676, 6,312,936, and 6,482,628, and various other patents. In some further embodiments, metalloproteases find use in the present invention, including but not limited to the neutral metalloprotease described in WO 07/044993. Suitable proteases include naturally occurring proteases or engineered variants specifically selected or engineered to work at relatively low temperatures.
[0511] Lipases: Suitable lipases include those of bacterial or fungal origin. Chemically modified, proteolytically modified, or protein engineered mutants are included. Examples of useful lipases include but are not limited to lipases from Humicola (synonym Thermomyces), e.g., from H. lanuginosa (T. lanuginosus) (see e.g., EP 258068 and EP 305216), from H. insolens (see e.g., WO 96/13580); a Pseudomonas lipase (e.g., from P. alcaligenes or P. pseudoalcaligenes; see, e.g., EP 218 272), P. cepacia (see e.g., EP 331 376), P. stutzeri (see e.g., GB 1,372,034), P. fluorescens, Pseudomonas sp. strain SD 705 (see e.g., WO 95/06720 and WO 96/27002), P. wisconsinensis (see e.g., WO 96/12012); a Bacillus lipase (e.g., from B. subtilis; see e.g., Dartois et al. Biochemica et Biophysica Acta, 1131: 253-360 (1993)), B. stearothermophilus (see e.g., JP 64/744992), or B. pumilus (see e.g., WO 91/16422). Additional lipase variants contemplated for use in the formulations include those described for example in: WO 92/05249, WO 94/01541, WO 95/35381, WO 96/00292, WO 95/30744, WO 94/25578, WO 95/14783, WO 95/22615, WO 97/04079, WO 97/07202, EP 407225, and EP 260105. Some commercially available lipase enzymes include Lipolase.RTM. and Lipolase Ultra.TM. (Novo Nordisk A/S and Novozymes A/S).
[0512] Polyesterases: Suitable polyesterases can be included in the composition, such as those described in, for example, WO 01/34899, WO 01/14629, and U.S. Pat. No. 6,933,140.
[0513] Amylases: The present compositions can be combined with other amylases, including other .alpha.-amylases. Such a combination is particularly desirable when different .alpha.-amylases demonstrate different performance characteristics and the combination of a plurality of different .alpha.-amylases results in a composition that provides the benefits of the different .alpha.-amylases. Other amylases include commercially available amylases, such as but not limited to STAINZYME.RTM., NATALASE.RTM., DURAMYL.RTM., TERMAMYL.RTM., FUNGAMYL.RTM. and BAN.TM. (Novo Nordisk A/S and Novozymes A/S); RAPIDASE.RTM., POWERASE.RTM., PURASTAR.RTM., and PREFERENZ.TM. (from DuPont Industrial Biosciences).
[0514] Cellulases: Cellulases can be added to the compositions. Suitable cellulases include those of bacterial or fungal origin. Chemically modified or protein engineered mutants are included. Suitable cellulases include cellulases from the genera Bacillus, Pseudomonas, Humicola, Fusarium, Thielavia, Acremonium, e.g., the fungal cellulases produced from Humicola insolens, Myceliophthora thermophila and Fusarium oxysporum disclosed for example in U.S. Pat. Nos. 4,435,307; 5,648,263; 5,691,178; 5,776,757; and WO 89/09259. Exemplary cellulases contemplated for use are those having color care benefit for the textile. Examples of such cellulases are cellulases described in for example EP 0495257, EP 0531372, WO 96/11262, WO 96/29397, and WO 98/08940. Other examples are cellulase variants, such as those described in WO 94/07998; WO 98/12307; WO 95/24471; PCT/DK98/00299; EP 531315; U.S. Pat. Nos. 5,457,046; 5,686,593; and 5,763,254. Commercially available cellulases include CELLUZYME.RTM. and CAREZYME.RTM. (Novo Nordisk A/S and Novozymes A/S); CLAZINASE.RTM. and PURADAX HA.RTM. (DuPont Industrial Biosciences); and KAC-500(B).TM. (Kao Corporation).
[0515] Peroxidases/Oxidases: Suitable peroxidases/oxidases contemplated for use in the compositions include those of plant, bacterial or fungal origin. Chemically modified or protein engineered mutants are included. Examples of useful peroxidases include peroxidases from Coprinus, e.g., from C. cinereus, and variants thereof as those described in WO 93/24618, WO 95/10602, and WO 98/15257. Commercially available peroxidases include for example GUARDZYME.TM. (Novo Nordisk A/S and Novozymes A/S).
[0516] The detergent composition can also comprise 2,6-.beta.-D-fructan hydrolase, which is effective for removal/cleaning of biofilm present on household and/or industrial textile/laundry.
[0517] The detergent enzyme(s) may be included in a detergent composition by adding separate additives containing one or more enzymes, or by adding a combined additive comprising all of these enzymes. A detergent additive, i.e. a separate additive or a combined additive, can be formulated e.g., as a granulate, a liquid, a slurry, and the like. Exemplary detergent additive formulations include but are not limited to granulates, in particular non-dusting granulates, liquids, in particular stabilized liquids or slurries.
[0518] Non-dusting granulates may be produced, e.g., as disclosed in U.S. Pat. Nos. 4,106,991 and 4,661,452 and may optionally be coated by methods known in the art. Examples of waxy coating materials are poly(ethylene oxide) products (e.g., polyethyleneglycol, PEG) with mean molar weights of 1,000 to 20,000; ethoxylated nonylphenols having from 16 to 50 ethylene oxide units; ethoxylated fatty alcohols in which the alcohol contains from 12 to 20 carbon atoms and in which there are 15 to 80 ethylene oxide units; fatty alcohols; fatty acids; and mono- and di- and triglycerides of fatty acids. Examples of film-forming coating materials suitable for application by fluid bed techniques are given in, for example, GB 1483591. Liquid enzyme preparations may, for instance, be stabilized by adding a polyol such as propylene glycol, a sugar or sugar alcohol, lactic acid or boric acid according to established methods. Protected enzymes may be prepared according to the method disclosed in EP 238,216.
[0519] The detergent composition may be in any convenient form, e.g., a bar, a tablet, a powder, a granule, a paste, or a liquid. A liquid detergent may be aqueous, typically containing up to about 70% water, and 0% to about 30% organic solvent. Compact detergent gels containing about 30% or less water are also contemplated. The detergent composition can optionally comprise one or more surfactants, which may be non-ionic, including semi-polar and/or anionic and/or cationic and/or zwitterionic. The surfactants can be present in a wide range, from about 0.1% to about 60% by weight.
[0520] When included therein the detergent will typically contain from about 1% to about 40% of an anionic surfactant, such as linear alkylbenzenesulfonate, .alpha.-olefinsulfonate, alkyl sulfate (fatty alcohol sulfate), alcohol ethoxysulfate, secondary alkanesulfonate, .alpha.-sulfo fatty acid methyl ester, alkyl- or alkenylsuccinic acid, or soap.
[0521] When included therein, the detergent will usually contain from about 0.2% to about 40% of a non-ionic surfactant such as alcohol ethoxylate, nonylphenol ethoxylate, alkylpolyglycoside, alkyldimethylamineoxide, ethoxylated fatty acid monoethanolamide, fatty acid monoethanolamide, polyhydroxy alkyl fatty acid amide, or N-acyl-N-alkyl derivatives of glucosamine ("glucamides").
[0522] The detergent may contain 0% to about 65% of a detergent builder or complexing agent such as zeolite, diphosphate, triphosphate, phosphonate, carbonate, citrate, nitrilotriacetic acid, ethylenediaminetetraacetic acid (EDTA), diethylenetriaminepentaacetic acid, alkyl- or alkenylsuccinic acid, soluble silicates or layered silicates (e.g., SKS-6 from Hoechst).
[0523] The detergent may comprise one or more polymers. Exemplary polymers include carboxymethylcellulose (CMC), poly(vinylpyrrolidone) (PVP), poly(ethylene glycol) (PEG), poly(vinyl alcohol) (PVA), poly(vinylpyridine-N-oxide), poly(vinylimidazole), polycarboxylates e.g., polyacrylates, maleic/acrylic acid copolymers), and lauryl methacrylate/acrylic acid copolymers.
[0524] The enzyme(s) of the detergent composition may be stabilized using conventional stabilizing agents, e.g., as polyol (e.g., propylene glycol or glycerol), a sugar or sugar alcohol, lactic acid, boric acid, or a boric acid derivative (e.g., an aromatic borate ester), or a phenyl boronic acid derivative (e.g., 4-formylphenyl boronic acid). The composition may be formulated as described in WO 92/19709 and WO 92/19708.
[0525] It is contemplated that in the detergent compositions, in particular the enzyme variants, may be added in an amount corresponding to about 0.01 to about 100 mg of enzyme protein per liter of wash liquor (e.g., about 0.05 to about 5.0 mg of enzyme protein per liter of wash liquor or 0.1 to about 1.0 mg of enzyme protein per liter of wash liquor).
[0526] Numerous exemplary detergent formulations to which the present amylases can be added (or is in some cases are identified as a component of) are described in WO2013063460. These include commercially available unit dose detergent formulations/packages such as PUREX.RTM. UltraPacks (Henkel), FINISH.RTM. Quantum (Reckitt Benckiser), CLOROX.TM. 2 Packs (Clorox), OxiClean Max Force Power Paks (Church & Dwight), TIDE.RTM. Stain Release, CASCADE.RTM. ActionPacs, and TIDE.RTM. Pods.TM. (Procter & Gamble), PS.
7.6. Methods of Assessing Amylase Activity in Detergent Compositions
[0527] Numerous .alpha.-amylase cleaning assays are known in the art, including swatch and micro-swatch assays. The appended Examples describe only a few such assays.
[0528] In order to further illustrate the compositions and methods, and advantages thereof, the following specific examples are given with the understanding that they are illustrative rather than limiting.
8. Brewing Compositions
[0529] The present variant amylase may be a component of a brewing composition used in a process of brewing, i.e., making a fermented malt beverage. Non-fermentable carbohydrates form the majority of the dissolved solids in the final beer. This residue remains because of the inability of malt amylases to hydrolyze the alpha-1,6-linkages of the starch. The non-fermentable carbohydrates contribute about 50 calories per 12 ounces of beer. an amylase, in combination with a glucoamylase and optionally a pullulanase and/or isoamylase, assist in converting the starch into dextrins and fermentable sugars, lowering the residual non-fermentable carbohydrates in the final beer.
[0530] The principal raw materials used in making these beverages are water, hops and malt. In addition, adjuncts such as common corn grits, refined corn grits, brewer's milled yeast, rice, sorghum, refined corn starch, barley, barley starch, dehusked barley, wheat, wheat starch, torrified cereal, cereal flakes, rye, oats, potato, tapioca, and syrups, such as corn syrup, sugar cane syrup, inverted sugar syrup, barley and/or wheat syrups, and the like may be used as a source of starch.
[0531] For a number of reasons, the malt, which is produced principally from selected varieties of barley, has the greatest effect on the overall character and quality of the beer. First, the malt is the primary flavoring agent in beer. Second, the malt provides the major portion of the fermentable sugar. Third, the malt provides the proteins, which will contribute to the body and foam character of the beer. Fourth, the malt provides the necessary enzymatic activity during mashing. Hops also contribute significantly to beer quality, including flavoring. In particular, hops (or hops constituents) add desirable bittering substances to the beer. In addition, the hops act as protein precipitants, establish preservative agents and aid in foam formation and stabilization.
[0532] Grains, such as barley, oats, wheat, as well as plant components, such as corn, hops, and rice, also are used for brewing, both in industry and for home brewing. The components used in brewing may be unmalted or may be malted, i.e., partially germinated, resulting in an increase in the levels of enzymes, including .alpha.-amylase. For successful brewing, adequate levels of .alpha.-amylase enzyme activity are necessary to ensure the appropriate levels of sugars for fermentation. an amylase, by itself or in combination with another .alpha.-amylase(s), accordingly may be added to the components used for brewing.
[0533] As used herein, the term "stock" means grains and plant components that are crushed or broken. For example, barley used in beer production is a grain that has been coarsely ground or crushed to yield a consistency appropriate for producing a mash for fermentation. As used herein, the term "stock" includes any of the aforementioned types of plants and grains in crushed or coarsely ground forms. The methods described herein may be used to determine .alpha.-amylase activity levels in both flours and stock.
[0534] Processes for making beer are well known in the art. See, e.g., Wolfgang Kunze (2004) "Technology Brewing and Malting," Research and Teaching Institute of Brewing, Berlin (VLB), 3rd edition. Briefly, the process involves: (a) preparing a mash, (b) filtering the mash to prepare a wort, and (c) fermenting the wort to obtain a fermented beverage, such as beer. Typically, milled or crushed malt is mixed with water and held for a period of time under controlled temperatures to permit the enzymes present in the malt to convert the starch present in the malt into fermentable sugars. The mash is then transferred to a mash filter where the liquid is separated from the grain residue. This sweet liquid is called "wort," and the left over grain residue is called "spent grain." The mash is typically subjected to an extraction, which involves adding water to the mash in order to recover the residual soluble extract from the spent grain. The wort is then boiled vigorously to sterilizes the wort and help develop the color, flavor and odor. Hops are added at some point during the boiling. The wort is cooled and transferred to a fermentor.
[0535] The wort is then contacted in a fermentor with yeast. The fermentor may be chilled to stop fermentation. The yeast flocculates and is removed. Finally, the beer is cooled and stored for a period of time, during which the beer clarifies and its flavor develops, and any material that might impair the appearance, flavor and shelf life of the beer settles out. The beer usually contains from about 2% to about 10% v/v alcohol, although beer with a higher alcohol content, e.g., 18% v/v, may be obtained. Prior to packaging, the beer is carbonated and, optionally, filtered and pasteurized.
[0536] The brewing composition comprising an amylase, in combination with a glucoamylase and optionally a pullulanase and/or isoamylase, may be added to the mash of step (a) above, i.e., during the preparation of the mash. Alternatively, or in addition, the brewing composition may be added to the mash of step (b) above, i.e., during the filtration of the mash. Alternatively, or in addition, the brewing composition may be added to the wort of step (c) above, i.e., during the fermenting of the wort.
[0537] A fermented beverage, such as a beer, can be produced by one of the methods above. The fermented beverage can be a beer, such as full malted beer, beer brewed under the "Reinheitsgebot," ale, IPA, lager, bitter, Happoshu (second beer), third beer, dry beer, near beer, light beer, low alcohol beer, low calorie beer, porter, bock beer, stout, malt liquor, non-alcoholic beer, non-alcoholic malt liquor and the like, but also alternative cereal and malt beverages such as fruit flavored malt beverages, e.g., citrus flavored, such as lemon-, orange-, lime-, or berry-flavored malt beverages, liquor flavored malt beverages, e.g., vodka-, rum-, or tequila-flavored malt liquor, or coffee flavored malt beverages, such as caffeine-flavored malt liquor, and the like.
9. Reduction of Iodine-Positive Starch
[0538] Variant amylases may reduce the iodine-positive starch (IPS), when used in a method of liquefaction and/or saccharification. One source of IPS is from amylose that escapes hydrolysis and/or from retrograded starch polymer. Starch retrogradation occurs spontaneously in a starch paste, or gel on ageing, because of the tendency of starch molecules to bind to one another followed by an increase in crystallinity. Solutions of low concentration become increasingly cloudy due to the progressive association of starch molecules into larger articles. Spontaneous precipitation takes place and the precipitated starch appears to be reverting to its original condition of cold-water insolubility. Pastes of higher concentration on cooling set to a gel, which on ageing becomes steadily firmer due to the increasing association of the starch molecules. This arises because of the strong tendency for hydrogen bond formation between hydroxy groups on adjacent starch molecules. See J. A. Radley, ed., Starch and its Derivatives 194-201 (Chapman and Hall, London (1968)).
[0539] The presence of IPS in saccharide liquor negatively affects final product quality and represents a major issue with downstream processing. IPS plugs or slows filtration system, and fouls the carbon columns used for purification. When IPS reaches sufficiently high levels, it may leak through the carbon columns and decrease production efficiency. Additionally, it may results in hazy final product upon storage, which is unacceptable for final product quality. The amount of IPS can be reduced by isolating the saccharification tank and blending the contents back. IPS nevertheless will accumulate in carbon columns and filter systems, among other things. The use of variant amylases is expected to improve overall process performance by reducing the amount of IPS.
[0540] All references cited herein are herein incorporated by reference in their entirety for all purposes. In order to further illustrate the compositions and methods, and advantages thereof, the following specific examples are given with the understanding that they are illustrative rather than limiting.
EXAMPLES
Example 1
Assays
[0541] Various assays used herein are set forth, below, for ease in reading. Any deviations from the protocols in later Examples are indicated in the relevant sections. In these experiments, a spectrophotometer was used to measure the absorbance of the products formed after the completion of the reactions.
A. Chelex Bead Treatment of Culture Supernatants
[0542] 96-well microtiter plates (MTPs) containing growing cultures were removed from incubators and Enzyscreen lids were replaced with disposable plastic sealers (Nunc cat. #236366; Rochester, N.Y., USA). Cells were separated from culture supernatant via centrifugation (1118 RCF, 5 minutes). 150 .mu.L supernatant was removed from each well and transferred to filter plates (Millipore Multiscreen HTS, Billerica, Mass., USA) containing Chelex beads prepared as described below. Plates were shaken vigorously for 5 minutes and supernatant from 3 replicate growth plates were collected into a single deep-well microtiter plate (Axygen, PDW-11-C) using a vacuum manifold device. Plates containing supernatants were sealed and stored at 4.degree. C.
[0543] Chelex-100 beads, 200-400 mesh (BioRad, Hercules, Calif., USA) were washed twice with 2 bed-volumes of 1 M HCl followed by 5 bed-volumes of ultrapure water on a sintered glass filter apparatus. 2 bed-volumes of 1 M KOH were used to wash the beads followed by another 5 bed-volume wash with ultrapure water. Filtered beads were transferred to a beaker and suspended with enough ultrapure water to produce slurry capable of mixing. The pH of the slurry was adjusted to 8-8.5 using HCl. The liquid was removed and the beads were dried using a scintered glass filter. A slurry of beads (40% w/v) was prepared in ultra pure water and its pH was adjusted to 8.0 using KOH/HCl. A slurry having a constant consistency was maintained by vigorous mixing. A bubble paddle reservoir device (V&P Scientific, San Diego, Calif., USA) was used to transfer 100 .mu.L of slurry to all wells of filter plates. Liquid was removed using a vacuum manifold device.
B. Protein Purification
[0544] Bacillus strains expressing amylase variants were grown in 2.5 L flasks in cultivation medium (enriched semi-defined media based on MOPs buffer, with urea as the major nitrogen source, glucose as the main carbon source, and supplemented with 1% soytone for robust cell growth) for 60-72 hours at 37.degree. C. or in 14 L tanks using a fed batch fermentation process with a medium of corn steep and soy flour supplemented with mineral salts and glucose as carbon source for 100 hours at 36.degree. C. Following incubation, the cells were separated from the fermentation medium by centrifugation and the supernatants were concentrated by ultra-filtration. Ammonium sulphate was added to the concentrate to a final concentration of 0.5M. The proteins were purified using hydrophobic interaction chromatography using a phenyl sepharose column on the AKTA Explorer FPLC system (GE Healthcare). The column was equilibrated with 50 mM HEPES, pH 8, with 2 mM CaCl.sub.2) and 0.5 M ammonium sulfate, and the proteins were eluted with 50 mM HEPES, pH 8, with 2 mM CaCl.sub.2) and 50% propylene glycol. After each HPLC run, liquid fractions associated with the peak of interest were pooled, and absorbance measurements of the pooled fractions were taken to estimate initial concentrations. Protein concentration of concentrated samples was determined by averaging the result from three different measurements: absorbance measurements at 280 nm, SDS-PAGE densitometry of acid-treated samples compared to a known standard, and by running the proteins on an HPLC system and taking absorbance measurements at 215 nm and 280 nm.
C. Protein Determination Assay
[0545] Protein determination assays were performed using chelex bead-treated culture supernatant from cultures grown in 96-well micro-titer plates (MTPs) over 3 days at 37.degree. C. with shaking at 300 rpm and 80% humidity. A fresh 96-well round-bottom MTP containing 25 supernatant per well was used for the High Performance Liquid Chromatography (HPLC) protein determination method. Supernatants were diluted four fold into 25 mM sodium acetate pH 5.5, and 10 .mu.L of each diluted sample was analyzed. An Agilent 1200 (Hewlett Packard) HPLC equipped with a Poroshell 300SB-C8 (Agilent Technologies Santa Clara, Calif., USA) column was used. Sample was bound to the column using 25 mM sodium acetate pH 5.5 and eluted over a gradient up to 70% acetonitrile. Absorbance was measured at 220 nm, integrated using ChemStation software (Agilent Technologies) and the protein concentration of samples was determined based on a standard curve of purified CspAmy2-v1 protein.
D. Ceralpha .alpha.-Amylase Activity Assay
[0546] The Ceralpha .alpha.-amylase assay was performed using the Ceralpha Kit (Megazyme, Wicklow, Ireland). The assay involves incubating culture supernatant with a substrate mixture under defined conditions, and the reaction is terminated (and color developed) by the addition of borate buffer (200 mM Boric acid/NaOH buffer, pH 10). The substrate is a mixture of the defined oligosaccharide "nonreducing-end blocked p-nitrophenyl maltoheptaoside" (BPNPG7) and excess levels of .alpha.-glucosidase (which has no action on the native substrate due to the presence of the "blocking group"). On hydrolysis of the oligosaccharide by endoacting .alpha.-amylase, the excess quantities of .alpha.-glucosidase present in the mixture give instantaneous and quantitative hydrolysis of the p-nitrophenyl maltosaccharide fragment to glucose and free p-nitrophenol. The absorbance at 405 nm was measured, which relates directly to the level of amylase in the sample analyzed.
[0547] The equipment used for this assay included a Biomek FX Robot (Beckman Coulter Brea, Calif., USA); a SpectraMAX MTP Reader (type 340-Molecular Devices, Sunnyvale, Calif., USA) and iEMS incubator/shaker (Thermo Scientific, Rockford, Ill., USA). The reagent and solutions used were: [0548] 1) p-nitrophenyl maltoheptaoside (BPNPG7) substrate (Megazyme Ceralpha HR kit); [0549] 2) 50 mM Malate buffer, 0.005% TWEEN.RTM. 80, pH 5.6 or 50 mM MOPS, 0.005% TWEEN.RTM. 80, pH 7 (dilution buffers); and [0550] 3) 200 mM Boric acid/NaOH buffer, pH 10 (STOP buffer).
[0551] A vial containing 54.5 mg BPNPG7 substrate was dissolved in 10 mL of MilliQ water and then diluted into 30 mL of dilution buffer to make up 40 mL of the working substrate (1.36 mg/mL). The amylase samples (fermentation supernatant) were diluted 40.times. with dilution buffer. The assay was performed by adding 54, of diluted amylase solution into the wells of a MTP followed by the addition of 55 .mu.L of diluted BPNPG7 working substrate solution. The solutions were mixed and the MTP was sealed with a plate seal and placed in an incubator/shaker (iEMS--Thermo Scientific) for 4 minutes at 25.degree. C. The reaction was terminated by adding 70 .mu.L STOP buffer and the absorbance was read at wavelength 400 nm in an MTP-Reader. A non-enzyme control was used to correct for background absorbance values.
E. Thermostability Assay
[0552] The thermostability of CspAmy2-v1 and variants was measured by determining the amylase activity using the Ceralpha .alpha.-amylase assay. The equipment used for this assay included a Biomek FX Robot (Beckman Coulter); a SpectraMAX MTP Reader (type 340-Molecular Devices), a Tetrad2DNA Engine PCR machine (Biorad), and iEMS incubator/shaker (Thermo Scientific). The reagent solutions used were (* not in all assays): [0553] 1) Heat stress buffers [0554] a) 50 mM KOAc pH 4.5 (5 ppm CaCl.sub.2), 50 ppm NaCl)*, [0555] b) 50 mM KOAc pH 5.0 (10 ppm CaCl.sub.2), 10 mM NaCl) [0556] c) 50 mM KOAc pH 5.7 (5 ppm CaCl.sub.2), 50 ppm NaCl), [0557] d) 50 mM KOAc pH 5.7 (no salt condition)*, [0558] 2) p-nitrophenyl maltoheptaoside (BPNPG7) substrate (Megazyme Ceralpha HR kit): [0559] 3) 50 mM Malate buffer, 0.005% TWEEN.RTM. 80, pH 5.6 (dilution buffer); and [0560] 4) 200 mM Boric acid/NaOH, pH 10 (STOP buffer). [0561] 5) Amylase culture supernatant: 1:10 master dilution enzyme plates were diluted 1:10 in each of the four heat stress buffers in a PCR plate
[0562] 5 .mu.L of the diluted enzyme samples were added to a 96-well PCR plate containing 55 .mu.L of diluted BPNPG7 working substrate solution and the initial amylase activity of the samples was determined using the Ceralpha .alpha.-amylase assay as described in Section C. The samples were subjected to heat stress for 3-6 minutes in a PCR thermocycler as follows: Buffers (a) 50.degree. C., (b) 59.degree.-60.degree. C., (c) 65.degree.-70.degree. C., and (d) 65.degree. C. The heat stressed samples were cooled immediately to room temperature and 5 .mu.L aliquots were assayed for amylase activity using the Ceralpha .alpha.-amylase assay as described in Section C. For each variant, the ratio of the initial and residual amylase activities was used to calculate the thermostability as follows: Thermostability=[t.sub.residual value]/[t.sub.initial value], so the heat stability activity ratio was calculated based on enzyme activity after heat incubation divided by enzyme activity before heat incubation. For each sample (variants) the performance index (PI) is calculated. The performance index for thermostability stability is determined by comparing the thermostability of the variant enzyme with that of a similarly treated reference enzyme.
F. Starch Hydrolysis Assays (Corn Flour and Corn Starch Application Assays)
[0563] Starch hydrolysis of corn flour and corn starch were used to measure specific activity of CspAmy2-v1 and variants. Activity was measured as reducing ends generated by the enzymatic breakdown of corn flour or corn starch. The reducing ends generated during the incubation with either substrate were quantified using a PAHBAH (p-hydroxybenzoic acid hydrazide) assay. The equipment used for the assay included a Biomek FX Robot (Beckman Coulter); a SpectraMAX MTP Reader (type 340-Molecular Devices), a Tetrad2DNA Engine PCR machine (Biorad), and iEMS incubator/shaker (Thermo Scientific), and a Bubble Paddle Reservoir.
[0564] Azure Farms Organic Corn Flour (Norco, CA) was ground to a fine powder using a consumer coffee grinder and then sifted to obtain a <250 micron fraction. The sifted corn flour was washed extensively with MilliQ water by repeated suspension and centrifugation. Cargill Farms Organic Corn Starch material was also washed extensively with MilliQ water by repeated suspension and centrifugation.
[0565] Both corn flour and corn starch washed fractions were suspended in MilliQ water containing 0.005% sodium azide as 20% (w/w) stock solutions. The stock solutions were further diluted with a 20.times. stock buffer solution to 10.9% w/v corn flour and corn starch solutions (final buffer concentration: 55 mM KOAc, pH 5).
[0566] 55 .mu.L of the diluted corn flour and corn starch substrates were added to PCR microtiter plates along with 5 .mu.L of 1:10 diluted enzyme samples using a bubble paddle reservoir. The plates were sealed and placed at 83.degree. C. for 5 minutes followed by a ramp down to 45.degree. C. The starch hydrolysis reaction was terminated by addition of 70 .mu.L 0.1 N NaOH. The plates were sealed and centrifuged for 3 minutes at 1610 RCF. The starch hydrolysis reaction products from both reactions were analyzed by the PAHBAH assay as described below.
[0567] PAHBAH Assay:
[0568] Aliquots of 80 .mu.L of 0.5 N NaOH were added to all wells of an empty PCR plate (a "PAHBAH reaction plate"), followed by 20 .mu.L of PAHBAH reagent (5% w/v p-hydroxybenzoic acid hydrazide (Sigma # H9882, St. Louis, Mo.), dissolved in 0.5 N HCl). The solutions were mixed by pipetting up and down. 20 .mu.L of the starch hydrolysis reaction supernatants were added to each well of the PAHBAH reaction plate. The plates were sealed and placed in a thermocycler, programmed for 2 minutes at 95.degree. C. to develop color, and then cooled to 20.degree. C. Samples of 80 .mu.L of the developed PAHBAH reaction mixtures were transferred to a fresh plate, and absorbance was measured at 450 nm in a spectrophotometer.
G. CS-28 Rice Starch Microswatch Assay
[0569] The principle of this amylase assay is the liberation of an orange dye due to the hydrolysis of rice starch incorporated in a cotton microswatch. The absorbance at 488 nm of the wash liquid is measured and this relates to the level of amylase activity in the sample analyzed at the desired conditions (pH, temperature, and buffer).
[0570] The equipment used for this assay included a Biomek FX Robot (Beckman Coulter), a SpectraMAX MTP Reader (type 340-Molecular Devices) and iEMS incubator/shaker (Thermo Scientific). The reagent and solutions used were: [0571] 1)CS-28 Microswatches (rice starch, colored); [0572] 2) 10 mM HEPES, 2 mM CaCl.sub.2), 0.005% TWEEN 80 buffer, pH 8.0, conductivity 1 mS/cm; [0573] 3) 25 mM CAPS, 2 mM CaCl.sub.2), 0.005% TWEEN 80 buffer, pH 10.0; conductivity 5 mS/cm (adjusted with 5M NaCl); and [0574] 4) 10 mM NaCl, 0.1 mM CaCl.sub.2), 0.005% TWEEN 80. [0575] 5) 50 mM MOPS pH7.15, 0.1 mM CaCl.sub.2).
[0576] CS-28 microswatches of 5.5 mm circular diameter were provided by the Center for Testmaterials (CFT, Vlaardingen, The Netherlands). Two microswatches were placed in each well of a 96-well Corning 9017 flat bottomed polystyrene MTP. The culture supernatants were diluted eight fold in 50 mM MOPS pH7.15, 0.1 mM CaCl.sub.2), and subsequently in 10 mM NaCl, 0.1 mM CaCl.sub.2), 0.005% TWEEN.RTM. 80 solution to approximately 1 ppm, final enzyme concentration.
[0577] The incubator/shaker was set at the desired temperature, 25.degree. C. (ambient temperature) or 50.degree. C. 174 .mu.L or 177 .mu.L of either HEPES or CAPS buffer, respectively, was added to each well of microswatch containing MTP and subsequently 6 .mu.L or 3 .mu.L of diluted enzyme solution was added to each well resulting in a total volume of 180 .mu.L/well. The MTP was sealed with a plate seal and placed in the iEMS incubator/shaker and incubated for 15 minutes at 1150 rpm at 25.degree. C. for cleaning at pH 8, low conductivity (1 mS/cm), or 15 minutes at 1150 rpm at 50.degree. C. for cleaning at pH 10, high conductivity (5 mS/cm). Following incubation under the appropriate conditions, 100 .mu.L of solution from each well was transferred to a new MTP, and the absorbance at 488 nm was measured using a MTP-spectrophotometer. Controls containing two microswatches and buffer but no enzyme were included for subtraction of background cleaning performance.
[0578] Each absorbance value was corrected by subtracting the blank (obtained after incubation of microswatches in the absence of enzyme), and the resulting absorbance provided a measure of the hydrolytic activity. A performance index (PI) was calculated for each sample.
[0579] For calculation of the wash performance indices (PI), the Langmuir equation was used to fit the data based on the reference enzyme control. Using the protein concentration of the variants, the expected performance based on the curve-fit was calculated. The observed performance was divided by the calculated performance. This value was then divided by the performance of the reference enzyme.
H. Detergent Stability Assay
[0580] The stability of the reference amylase and variants thereof was determined by measuring their activity after incubation under defined conditions, in the presence of a 10% detergent mixture (commercially purchased Persil Color Gel detergent, Henkel (Dusseldorf, Germany), purchased in 2011). The detergent was heat-inactivated before use, and the initial and residual amylase activities were determined using the Ceralpha .alpha.-amylase assay as described in section C, above.
[0581] The equipment used for this assay included a Biomek FX Robot (Beckman Coulter); a SpectraMAX MTP Reader (type 340-Molecular Devices), a Tetrad2DNA Engine PCR machine (Biorad), and iEMS incubator/shaker (Thermo Scientific). The reagent solutions used were: [0582] 1) p-nitrophenyl maltoheptaoside (BPNPG7) substrate (Megazyme Ceralpha HR kit): [0583] 2) Liquid detergent (Persil color gel, enzyme inactivated by heating for 4 hrs at 90.degree. C.); [0584] 3) 50 mM MOPS, 0.1 mM CaCl.sub.2), 0.005% TWEEN.RTM.80 buffer, pH 7 (dilution buffer); [0585] 4) 10% detergent solution diluted in dilution buffer; [0586] 5) 200 mM Boric acid/NaOH buffer, pH 10 (STOP buffer); and [0587] 6) Amylase culture supernatants diluted eight fold in 50 mM MOPS pH 7.15, 0.1 mM CaCl.sub.2) containing 0-100 .mu.g/mL protein.
[0588] 85 .mu.L of a 10% detergent solution was added to a 96-well PCR plate and mixed with 15 .mu.L of the diluted culture supernatant. A sample from the PCR plate was diluted 3.times. in dilution buffer and a 5 .mu.L aliquot of this dilution was used to determine initial amylase activity. The PCR plate was incubated in a Tetrad PCR block at 80.5.degree. C. for 5 minutes. After incubation, detergent-enzyme mix was diluted 3.times. in dilution buffer and residual activity was measured. Initial (t.sub.initial) and residual (t.sub.residual) amylase activity was determined by the Ceralpha .alpha.-amylase assay as described above in Section C using a 5 .mu.L sample.
[0589] For each variant, the ratio of the residual and initial amylase activities was used to calculate the detergent stability as follows: Detergent stability=[t.sub.residual value]/[t.sub.initial value].
[0590] For each sample (variants) the performance index (PI) was calculated. The performance index for detergent stability is determined by comparing the detergent stability of the variant enzyme with that of the similarly treated reference enzyme.
I. Performance Index
[0591] The performance index (PI) compares the performance or stability of the variant and the reference enzyme at the same protein concentration. In addition, the theoretical values can be calculated, using the parameters of the Langmuir equation of the standard enzyme. A performance index (PI) that is greater than 1 (PI>1) indicates improved performance by a variant as compared to the reference enzyme, while a PI of 1 (PI=1) identifies a variant that performs the same as the reference enzyme, and a PI that is less than 1 (PI<1) identifies a variant that performs worse than the reference enzyme.
Example 2
Generation of Bacillus Strains Expressing CspAmy2-v1
[0592] In this example, the construction of Bacillus subtilis strains expressing CspAmy2-v1 (SEQ ID NO: 2) amylase and variants, thereof, is described. CspAmy2-v1 amylase is a variant of CspAmy2 (SEQ ID NO: 1) amylase having a deletion of both R178 and G179 (i.e., .DELTA.RG). CspAmy2 is an amylase from a Cytophaga sp., for which the nucleotide sequence was described by Chii-Ling et al. (2002) Appl. Environ. Microbiol. 68: 3651-3654. CspAmy2-v1 was described as having increased thermostability over the CspAmy2 by Rong-Jen et al. (2003) Appl. Environ. Microbiol. 69: 2383-85.
[0593] A synthetic DNA fragment (SEQ ID NO: 7) encoding CspAmy2-v1 was produced by GeneArt AG (Regensburg, Germany) and served as template DNA for the construction of Bacillus subtilis strains expressing CspAmy2-v1 amylase and variants, thereof. The DNA fragment includes a codon-modified nucleotide sequence encoding the mature form of CspAmy2-v1 amylase adjacent to a sequence encoding the LAT signal peptide (underlined):
TABLE-US-00008 ATGAAACAACAAAAACGGCTTTACGCCCGATTGCTGACGCTGTTATTTGC GCTCATCTTCTTGCTGCCTCATTCTGCAGCTAGCGCAGCAGCGACAAACG GAACAATGATGCAGTATTTCGAGTGGTATGTACCTAACGACGGCCAGCAA TGGAACAGACTGAGAACAGATGCCCCTTACTTGTCATCTGTTGGTATTAC AGCAGTATGGACACCGCCGGCTTATAAGGGCACGTCTCAAGCAGATGTGG GGTACGGCCCGTACGATCTGTATGATTTAGGCGAGTTTAATCAAAAAGGT ACAGTCAGAACGAAGTATGGCACAAAAGGAGAACTTAAATCTGCTGTTAA CACGCTGCATTCAAATGGAATCCAAGTGTATGGTGATGTCGTGATGAATC ATAAAGCAGGTGCTGATTATACAGAAAACGTAACGGCGGTGGAGGTGAAT CCGTCTAATAGAAATCAGGAAACGAGCGGCGAATATAATATTCAGGCATG GACAGGCTTCAACTTTCCGGGCAGAGGAACAACGTATTCTAACTTCAAAT GGCAGTGGTTCCATTTTGATGGAACGGATTGGGACCAGAGCAGAAGCCTC TCTAGAATCTTCAAATTCACGGGAAAGGCGTGGGACTGGGAGGTTTCTTC AGAAAACGGAAATTATGACTATCTGATGTACGCGGACATTGATTATGACC ATCCGGATGTCGTGAATGAAATGAAAAAGTGGGGCGTCTGGTATGCCAAC GAAGTTGGGTTAGATGGATACAGACTTGACGCGGTCAAACATATTAAATT TAGCTTTCTCAAAGACTGGGTGGATAACGCAAGAGCAGCGACGGGAAAAG AAATGTTTACGGTTGGCGAATATTGGCAAAATGATTTAGGGGCCCTGAAT AACTACCTGGCAAAGGTAAATTACAACCAATCTCTTTTTGATGCGCCGTT GCATTACAACTTTTACGCTGCCTCAACAGGGGGTGGATATTACGATATGA GAAATATTCTTAATAACACGTTAGTCGCAAGCAATCCGACAAAGGCTGTT ACGTTAGTTGAGAATCATGACACACAGCCTGGACAATCACTGGAATCAAC AGTCCAACCGTGGTTTAAACCGTTAGCCTACGCGTTTATTCTCACGAGAA GCGGAGGCTATCCTTCTGTATTTTATGGAGATATGTACGGTACAAAAGGA ACGACAACAAGAGAGATCCCTGCTCTTAAATCTAAAATCGAACCTTTGCT TAAGGCTAGAAAAGACTATGCTTATGGAACACAGAGAGACTATATTGATA ACCCGGATGTCATTGGCTGGACGAGAGAAGGGGACTCAACGAAAGCCAAG AGCGGTCTGGCCACAGTGATTACAGATGGGCCGGGCGGTTCAAAAAGAAT GTATGTTGGCACGAGCAATGCGGGTGAAATCTGGTATGATTTGACAGGGA ATAGAACAGATAAAATCACGATTGGAAGCGATGGCTATGCAACATTTCCT GTCAATGGGGGCTCAGTTTCAGTATGGGTGCAGCAA
[0594] The mature form of the CspAmy2-v1 polypeptide produced from the pHPLT02-CspAmy2-v1 vector is shown, below, as SEQ ID NO: 2.
TABLE-US-00009 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRNQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWEVSSENGNYDYLMYAD IDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0595] To express CspAmy2-v1, the CspAmy2-v1-encoding DNA fragment was cloned into the pHPLT02 vector, a modified version of the pHPLT vector (Solingen et al. (2001) Extremophiles 5:333-341) by GeneArt and fused in-frame to the AmyL (LAT) signal peptide using the unique NheI and XhoI restriction sites, resulting in plasmid pHPLT02-CspAmy2-v1. The pHPLT expression vector contains the B. licheniformis LAT promoter (Plat) and additional elements from pUB110 (McKenzie et al. (1986) Plasmid, 15: 93-103) including a replicase gene (reppUB), a neomycin/kanamycin resistance gene (neo) and a bleomycin resistance marker (bleo). Site-directed mutagenesis (Stratagene) was used to change the nucleotides 5'-TCA-3' of Serine 28 of the AmyL signal peptide to nucleotides 5'-AGC-3' in order to introduce the unique NheI restriction site.
[0596] A suitable B. subtilis strain was transformed with pHPLT02-CspAmy2-v1 plasmid DNA using a method known in the art (WO 02/14490). The B. subtilis transformants were selected on agar plates containing heart infusion agar (Difco, Catalog No. 244400, Lawrence, Kans., USA and 10 mg/L neomycin sulfate (Sigma, Catalog No. N-1876; contains 732 .mu.g neomycin per mg, St. Louis, Mo., USA). Selective growth of B. subtilis transformants harboring the pHPLT02-CspAmy2-v1 plasmid was performed in shake flasks at 37.degree. C. for .about.65h in MBD medium (enriched semi-defined medium based on MOPs buffer, with urea as major nitrogen source, glucose as the main carbon source, and supplemented with 1% soytone for robust cell growth) containing 5 mM CaCl.sub.2) and 10 ppm neomycin. Growth resulted in the production of secreted CspAmy2-v1 amylase with starch hydrolyzing activity.
Example 3
Generation of Bacillus Strains Expressing CspAmy2 Combinatorial Variants
[0597] In this example, the construction of Bacillus subtilis and Bacillus licheniformis strains expressing two combinatorial CspAmy2 variants (CspAmy2-v5 and CspAmy2-v6) is described. CspAmy2-v5 is a variant of CspAmy2 with the mutations E187P, I203Y, G476K, and lacking R178 and G179. CspAmy2-v6 is a variant of CspAmy2 with the mutations E187P, I203Y, G476K, R458N, T459S, D460T, and lacking R178 and G179. For expression of CspAmy2-v5 and CspAmy2-v6, expression constructs were created by a combination of gene synthesis and fusion PCR, using standard techniques.
[0598] The CspAmy2-v5 and CspAmy2-v6 expression constructs were ligated into vector pICatH (described in, e.g., EP2428572 and U.S. Pat. No. 7,968,691) and transformed into competent B. subtilis cells (amyE negative) as known in the art (WO 2002/014490). In these clones, the DNA fragments encoding CspAmy2-v5 and CspAmy2-v6 are fused in-frame to a sequence encoding the B. licheniformis amylase (amyL) signal peptide which leads to secretion of the amylase variants into the growth medium. Expression of CspAmy2-v5 and CspAmy2-v6 in these clones is driven by the B. licheniformis amylase (amyL) promoter (FIGS. 1A and 1B).
[0599] B. subtilis transformants were selected on agar plates containing heart infusion agar (Difco) and 10 mg/L neomycin sulfate, 5 mg/L chloramphenicol and starch azure (Sigma). For each construct, one transformant with starch hydrolyzing activity was selected and the amylase expression constructs in pICatH were sequence verified by DNA sequencing (BaseClear, the Netherlands). The resulting plasmids, pICatH-CspAmy2-v5 and pICatH-CspAmy2-v6, were isolated from the B. subtilis clones and transformed into B. licheniformis cells (amyL and catH negative) and integrated into the genome as described in EP2428572 and U.S. Pat. No. 7,968,691. After excision of vector sequences, expression constructs were amplified by subjecting the strains to a stepwise increase in chloramphenicol concentration up to 50 .mu.g/ml. The B. licheniformis strains showed halos on starch azure plates, indicating secretion of active CspAmy2-v5 and CspAmy2-v6 amylase.
[0600] The amino acid sequence of the mature form of CspAmy2-v5 is shown, below, as SEQ ID NO: 8:
TABLE-US-00010 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRNQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNKGSVSVWVQQ
[0601] The amino acid sequence of the mature form of CspAmy2-v6 is shown, below, as SEQ ID NO: 9:
TABLE-US-00011 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRNQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNNSTKITIGSDGYATFPVNKGSVSVWVQQ
Example 4
Cleaning Performance of CspAmy2 Combinatorial Variants
[0602] The cleaning performance of purified CspAmy2-v5 and CspAmy2-v6 was analyzed in a microswatch cleaning assay. CFT CS-28 rice starch on cotton swatches or EMPA 161 maize starch on cotton cretonne (Center for Testmaterials, BV, Vlaardingen, Netherlands) containing an indicator dye bound to the starch were punched to form discs measuring 5.5 mm in diameter. Two discs were placed in each well of 3 flat-bottom non-binding 96-well assay plates.
[0603] Both variants (i.e., CspAmy2-v5 and CspAmy2-v6), the parent molecule (CspAmy2), and a commercially-available benchmark detergent .alpha.-amylase, i.e., PURASTAR.RTM. (AmyL; B. licheniformis .alpha.-amylase; Danisco US Inc.), were diluted to 0.5 mg/mL in dilution buffer (50 mM MOPS, pH 7.2, 0.005% Tween), and then further diluted to 2 ppm in a microtiter plate. 200 .mu.L of these samples were transferred into the first row of each of the swatch plates. 100 .mu.L of HEPES buffer (25 mM HEPES, pH 8.0, with 2 mM CaCl.sub.2) and 0.005% Tween-80) was then added to each well of the next 5 rows of the swatch plates. 100 .mu.L of the diluted enzyme samples were then transferred from the first row into the next row, mixed well, and serial dilutions were continued until before the last row, which served as blanks. Once all rows contained 100 .mu.L of solution, 100 .mu.L of buffer were added to each well of the plate to result in final volumes of 200 .mu.L per well, and final enzyme concentrations of 1, 0.5, 0.25, 0.125, 0.0625, and 0 ppm.
[0604] Plates were incubated at 25.degree. C. with agitation at 1150 rpm for 15 minutes. Enzyme performance was assessed by the amount of color released into the wash liquor, which was quantified spectrophotometrically at 488 nm by the transfer of 150 .mu.L of the final wash solution to fresh medium-binding microtiter plates. Triplicate reads were blank-subtracted and averaged. The results of the soil removal assay are shown in FIG. 2. CspAmy2-v5 and CspAmy2-v6 demonstrated similar performance to each other and are significantly better than the parent molecule, CspAmy2, and much better than PURASTAR.RTM..
Example 5
Thermostability Assessment of CspAmy2 Combinatorial Variants
A. Thermostability in Buffer
[0605] 0.5 mg/mL stocks of purified enzymes (i.e., CspAmy2-v5, CspAmy2-v6, CspAmy2), PURASTAR.RTM. (Bacillus licheniformis .alpha.-amylase), and ACE-QK (variant of Bacillus sp. TS-23 .alpha.-amylase described in US20120045817 and WO2010/115028) were further diluted to 5 ppm in dilution buffer (50 mM MOPS, pH 7.2, 0.005% Tween). 50 .mu.L of each enzyme were added to each of 9 strips of PCR tubes and sealed. One "unstressed" sample of each enzyme was incubated at room temperature throughout the duration of the experiment. The other 8 samples were incubated in a thermocycler at 45, 55, 65, 70, 75, 80, 85, or 90.degree. C. for 15 minutes. The samples were then transferred to microtiter plates in triplicate, and .alpha.-amylase activity was for the unstressed and stressed samples using the Ceralpha reagent (Megazyme, Inc.). Residual activity was calculated by dividing the activity of each amylase after the thermal stress by the activity of that unstressed amylase. The results are shown in FIG. 3. CspAmy2-v5 and CspAmy2-v6 demonstrated increased thermostability at low calcium concentrations compared to all other tested enzymes.
B. Thermostability in Buffer with Added Calcium
[0606] As above, 0.5 mg/mL stocks of purified enzymes were further diluted to 5 ppm in dilution buffer, in this case supplemented with 5 mM calcium chloride (i.e., 50 mM MOPS, pH 7.2, 0.005% Tween, 5 mM calcium chloride). The experiment was carried out as described above. The results are shown in FIG. 4. CspAmy2-v5 and CspAmy2-v6 demonstrated similar thermostability compared to the other tested enzymes.
C. Thermostability in Liquid Detergent
[0607] Commercial detergents EPSIL.TM. Perfect (McBride) and OMO.TM. Color (Unilever) were heat inactivated at 90.degree. C. for 4 hours to eliminate existing enzyme activities. Following inactivation, the activity CspAmy2-v5, CspAmy2-v6, CspAmy2, PURASTAR.RTM., and ACE-QK in the heat-inactivated detergents was measured using the Suc-AAPF-pNA and Ceralpha assays to ensure that any protease and amylase activities, respectively, had been abolished. 10% solutions of both of these liquid detergents were made in water.
[0608] 100 .mu.L of each of the 0.5 mg/mL purified enzyme stocks at were added to 400 .mu.L of the 10% detergent solutions. 50 .mu.L of these samples were added to strips of PCR tubes and incubated at the aforementioned temperatures for 15 minutes each, as described above. When samples were removed from the thermocycler, and before .alpha.-amylase activity was measured, an additional 1:20 dilution was made in dilution buffer. As above, residual activity was calculated by dividing the activity of each amylase after the detergent and thermal stress by the activity of that unstressed amylase. The results are shown in FIG. 5 (OMO.TM.) and FIG. 6 (EPSIL.TM.) CspAmy2-v5 and CspAmy2-v6 demonstrated superior thermostability compared to the parent enzyme and other tested enzymes.
Example 6
Generation of Bacillus Strains Expressing CspAmy2 Combinatorial Variants
A. Design of Combinatorial Variants
[0609] Synthetic DNA encoding CspAmy2-v1 variants having various combinations of substitutions at positions N126, Y150, F153, L171, T180, E187, 1203, and S241 (referring to SEQ ID NO: 1) were constructed by GeneArt and delivered as plasmids transformed in a B. subtilis host, as described for CspAmy2-v5 and CspAmy2-v6 in Example 2. With single substitutions at each of eight sites, 256 combinations were possible. The specific substitutions N126Y, Y150H, F153W, L171N, T180H, E187P, I203Y, and S241Q, were selected, and ten of the possible combinations were made and tested. The names of the variants, and the amino acid residues present at each of the eight positions, is shown in Table 2. For clarity, the full names of the variants in the Table are CspAmy2-C16A-CspAmy2-C16J. In some table and figure the names of the variants are abbreviated but always clear from the description. Further variants of CspAmy2-v1 additionally having the mutations E187P or S241Q were made, and designated CspAmy2-v1-E187P or CspAmy2-v1-S241Q, respectively.
TABLE-US-00012 TABLE 2 Combinations of mutations tested Name N126 Y150 F153 L171 T180 E187 I203 S241 C16A Y Y F L T P Y S C16B Y Y F L T E Y Q C16C Y Y W L T P Y S C16D Y Y W L T E Y Q C16E Y Y W L H P Y S C16F Y Y W L H E Y Q C16G Y H W N T P Y S C16H Y H W N T E Y Q C16I Y H W N H P Y S C16J Y H W N H E Y Q
B. Construction of Bacillus Strains Expressing CspAmy2 Combinatorial Variants
[0610] The pHPLT02-CspAmy2-v1 plasmid DNA (encoding CspAmy2-v1, see Example 2) served as template to produce the additional combinatorial libraries at pre-selected sites in the mature region. The pHPLT02 expression vector was derived from the pHPLT vector. The pHPLT expression vector contains the B. licheniformis LAT promoter (Plat) and additional elements from pUB110 (McKenzie et al. (1986) Plasmid, 15: 93-103) including a replicase gene (reppUB), a neomycin/kanamycin resistance gene (neo) and a bleomycin resistance marker (bleo). GeneArt AG (Regensburg, Germany) created combinatorial libraries at the positions described using their standard protocols. The corresponding codons for each site of interest were substituted with codons for at least one non-wild-type amino acid. The codon-mutagenized pHPLT02-CspAmy2-v1 mixes were used to transform competent B. subtilis cells as known in the art (WO 2002/014490) to generate the CspAmy2-v1 combinatorial libraries. Transformation mixes were plated on Heart Infusion (HI) agar plates containing 10 mg/L neomycin sulfate. For each library, single bacterial colonies were picked and grown in TSB (tryptone and soy-based broth) liquid medium with 10 mg/ml neomycin selection for subsequent DNA isolation and gene sequence analysis. Variants were generated and identified as members of this combinatorial library. Selective growth of the variants was performed in 96-well MTPs at 37.degree. C. for 68 hours in MBD medium (enriched semi-defined medium based on MOPs buffer, with urea as major nitrogen source, glucose as the main carbon source, and supplemented with 1% soytone for robust cell growth).
[0611] The amino acid sequence of mature CspAmy2-16A is shown, below, as SEQ ID NO: 10:
TABLE-US-00013 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0612] The amino acid sequence of mature CspAmy2-16B is shown, below, as SEQ ID NO: 11:
TABLE-US-00014 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWEVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFQFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0613] The amino acid sequence of mature CspAmy2-16C is shown, below, as SEQ ID NO: 12:
TABLE-US-00015 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNWKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0614] The amino acid sequence of mature CspAmy2-16D is shown, below, as SEQ ID NO: 13:
TABLE-US-00016 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNWKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWEVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFQFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0615] The amino acid sequence of the mature CspAmy2-16E is shown, below, as SEQ ID NO: 14:
TABLE-US-00017 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNWKWQWFHFDGTDWDQSRSLSRIFKFHGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0616] The amino acid sequence of mature CspAmy2-16F is shown, below, as SEQ ID NO: 15:
TABLE-US-00018 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNWKWQWFHFDGTDWDQSRSLSRIFKFHGKAWDWEVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFQFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0617] The amino acid sequence of mature CspAmy2-16G is shown, below, as SEQ ID NO: 16:
TABLE-US-00019 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTH SNWKWQWFHFDGTDWDQSRSNSRIFKFTGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0618] The amino acid sequence of mature CspAmy2-16H is shown, below, as SEQ ID NO: 17:
TABLE-US-00020 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTH SNWKWQWFHFDGTDWDQSRSNSRIFKFTGKAWDWEVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFQFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0619] The amino acid sequence of mature CspAmy2-16I is shown, below, as SEQ ID NO: 18:
TABLE-US-00021 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTH SNWKWQWFHFDGTDWDQSRSNSRIFKFHGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0620] The amino acid sequence of mature CspAmy2-16J is shown, below, as SEQ ID NO: 19:
TABLE-US-00022 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTH SNWKWQWFHFDGTDWDQSRSNSRIFKFHGKAWDWEVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFQFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0621] The amino acid sequence of mature CspAmy2-v1-E187P is shown, below, as SEQ ID NO: 20:
TABLE-US-00023 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRNQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWPVSSENGNYDYLMYAD IDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
[0622] The amino acid sequence of mature CspAmy2-v1-S241Q is shown, below, as SEQ ID NO: 21:
TABLE-US-00024 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRNQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFTGKAWDWEVSSENGNYDYLMYAD IDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFQFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNGGSVSVWVQQ
Example 7
Thermostability Assessment of CspAmy2 Variants
[0623] Combination variants CspAmy2-C16A-CspAmy2-C16J from Example 5 (i.e., "C16A"-"C16G" or "16A-16G," respectively) were tested for thermal stability under the following conditions:
[0624] 1. 50 mM potassium acetate, pH 5.7, 0.125 mM CaCl.sub.2), 2.2 mM NaCl, 85.degree. C.
[0625] 2. 50 mM potassium acetate, pH 5.0, 0.125 mM CaCl.sub.2), 2.2 mM NaCl, 70.degree. C.
[0626] 3. 50 mM potassium acetate, pH 4.5, 0.125 mM CaCl.sub.2), 2.2 mM NaCl, 65.degree. C.
[0627] Stock solutions of each variant were prepared by diluting purified variants to a final protein concentration of 1 mg/mL in milli-Q water, then each variant was further diluted (200-fold) in each of the above buffers (final enzyme dose is 5 .mu.g/mL). The diluted enzyme solutions were pre-heated to appropriate temperature for two minutes and then cooled on ice to disrupt any protein aggregates. 50 .mu.L of each enzyme solution was transferred to 0.2 mL PCR strip tubes, which were heated to the appropriate temperature (based on buffer pH) and allowed to incubate over a two-hour period. The samples were then placed in an ice-water bath to end the heat-stress period.
[0628] Once all time points were collected for each buffer, residual activity was determined using the Ceralpha assay, as described in Example 1. Two independent inactivation time-course experiments were performed for each variant. Plots of residual activity vs. time were modeled with a single exponential decay equation to determine a rate constant (k) for decay. The half-life of decay was defined as ln(2)/k. These experiments were performed in duplicate for each variant.
[0629] The performance index (PI) for each variant was defined as the ratio of the variant half-life to the half-life of a reference parent molecule. For variants C16A, C16C, C16E, C16G, and C16I, the reference molecule was CspAmy2-v1-E187P. For variants C16B, C16D, C16F, C16H, and C16J, the reference molecule was CspAmy2-v1-S241Q. The relative half-lives and performance indexes are shown in the Table in FIG. 7. Other experiments demonstrated that the stability of C16D was similar to that of STAINZYME.RTM., while the stability of C16F was greater than that of STAINZYME.RTM. (not shown). The relative performance of the combination variants at pH 4.5 and 65.degree. C., pH 5.0 at 70.degree. C., and pH 5.7 ay 85.degree. C., is shown graphically in FIGS. 8-10, respectively. All C16A-J variants were more stable than their respective parent molecules.
Example 8
Viscosity Reduction Using CspAmy2 Variants
[0630] Combination variants CspAmy2-C16A-CspAmy2-C16J from Example 5 were tested for their ability to reduce the viscosity of a starch solution, as a measure of starch hydrolysis activity. Viscosity experiments were performed using a Rapid Visco Analyser (Newport Scientific; herein "RVA") with 38 mm.times.68 mm plain washed cans (Part No. AA0384001) and a double-skirted paddle (Part No. NS101783). Data acquisition and analysis was performed with Thermocline for Windows (version 3.11; (Newport Scientific)). Immediately before each run, 33 g of 25% dry solids (ds) corn flour slurry was prepared in an RVA can as follows: 9.17 g corn flour (11.8% moisture content) was weighed out on an analytical balance and mixed with 23.83 g deionized water. Sample pH was adjusted with 0.0828 ml 1N sulfuric acid (for pH 5.8) or 0.285 ml 1N sulfuric acid (for pH 5.0). Enzyme was added at appropriate dose, and the can was placed in the viscometer. All runs were 10 minutes in length, with a temperature ramp to 85.degree. C. over 80 seconds followed by a temperature hold at 85.degree. C. for the remainder of the run. Each enzyme was assessed at three doses. Dose responses were calculated for two parts of the viscosity curve. Peak viscosity is defined as the maximum viscosity reached during the experiment, and final viscosity is defined as the viscosity reading at the end of the experiment. Because viscosity has a reciprocal relationship with enzyme dose, dose response curves were linearized by plotting 1/viscosity (also called fluidity) vs. enzyme dose. Slopes of these curves were used as a quantitative measure of the specific activity of each variant. The performance index (PI) for each variant is defined as the ratio of the variant specific activity to the specific activity of a reference parent molecule. For variants CspAmy2-C16A, CspAmy2-C16C, CspAmy2-C16E, CspAmy2-C16G, and CspAmy2-C16I, the reference is CspAmy2-v1-E187P. For variants CspAmy2-C16B, CspAmy2-C16D, CspAmy2-C16F, CspAmy2-C16H, and CspAmy2-C16J, the reference is CspAmy2-v1-S241Q. Viscosity reduction performance is shown in Tables 3 and 4.
TABLE-US-00025 TABLE 3 Improvements in viscosity reduction at pH 5.8 Peak Final Fluidity/mg Fluidity/mg enzyme Peak PI enzyme Final PI C16A 0.0044 1.10 0.17 1.99 C16B 0.0047 1.18 0.17 1.47 C16C 0.0048 1.20 0.17 1.96 C16D 0.0049 1.22 0.18 1.53 C16E 0.0043 1.08 0.18 2.12 C16F 0.0044 1.10 0.18 1.56 C16G 0.0051 1.28 0.17 1.99 C16H 0.0045 1.11 0.17 1.44 C16I 0.0047 1.19 0.19 2.16 C16J 0.0050 1.25 0.19 1.65 E187P 0.0040 1.00 0.09 1.00 S241Q 0.0040 1.00 0.12 1.00
TABLE-US-00026 TABLE 4 Improvements in viscosity reduction at pH 5.0 Peak Final Fluidity/mg Fluidity/mg enzyme Peak PI enzyme Final PI C16A 0.0180 1.27 0.17 1.96 C16B 0.0065 1.27 0.14 1.80 C16C 0.0061 1.24 0.19 2.08 C16D 0.0062 1.20 0.16 1.97 C16E 0.0065 1.33 0.19 2.16 C16F 0.0061 1.18 0.17 2.16 C16G 0.0072 1.46 0.19 2.10 C16H 0.0063 1.23 0.16 1.98 C16I 0.0067 1.35 0.20 2.23 C16J 0.0069 1.35 0.19 2.36 E187P 0.0049 1.00 0.09 1.00 S241Q 0.0051 1.00 0.08 1.00
Example 9
Cleaning Performance and Detergent Stability of CspAmy2 Variants
A. Cleaning Performance of CspAmy2 Variants
[0631] The cleaning performance of combination variants CspAmy2-C16A-CspAmy2-C16J from Example 5 was analyzed in a microswatch cleaning assay using CFT CS-28 rice starch on cotton swatches. The assay was performed using culture supernatants. Protein concentration for the supernatants was quantified using HPLC. The equipment used for this assay included a Biomek FX Robot (Beckman Coulter), a SpectraMAX MTP Reader (type 340-Molecular Devices) and iEMS incubator/shaker (Thermo Scientific). The reagent and solutions used were: [0632] 1)CS-28 Microswatches (rice starch, colored); [0633] 2) 10 mM HEPES, 2 mM CaCl.sub.2), 0.005% TWEEN 80 buffer, pH 8.0, conductivity 1 mS/cm; [0634] 3) 50 mM MOPS pH7.15, 0.005% TWEEN 80.
[0635] CS-28 microswatches of 5.5 mm circular diameter were provided by the Center for Testmaterials (CFT, Vlaardingen, The Netherlands). Two microswatches were placed in each well of a 96-well Corning 9017 flat bottomed polystyrene MTP. The culture supernatants were diluted twenty-fold in 50 mM MOPS pH7.15, 0.005% TWEEN 80.
[0636] The incubator/shaker was set at 25.degree. C. (ambient temperature). 178.5 .mu.L of HEPES buffer, respectively, was added to each well of microswatch containing MTP and subsequently 1.5 .mu.L of diluted enzyme solution was added to each well resulting in a total volume of 180 .mu.L/well. The MTP was sealed with a plate seal and placed in the iEMS incubator/shaker and incubated for 15 minutes at 1150 rpm at 25.degree. C. Following incubation under the appropriate conditions, 100 .mu.L of solution from each well was transferred to a new MTP, and the absorbance at 488 nm was measured using a MTP-spectrophotometer. Controls containing two microswatches and buffer but no enzyme were included for subtraction of background cleaning performance.
[0637] Each absorbance value was corrected by subtracting the blank (obtained after incubation of microswatches in the absence of enzyme), and the resulting absorbance provided a measure of the hydrolytic activity. A performance index (PI) was calculated for each sample compared to CspAmy2-v1, assuming a linear response in the range of the assay used. The results are shown in Table 4.1.
B. Detergent Stability of CspAmy2 Variants
[0638] The detergent stability of the additional CspAmy2 variants was determined by measuring their activity after incubation under defined conditions, in the presence of a 10% detergent mixture (commercially purchased Persil Color Gel detergent, Henkel (Dusseldorf, Germany), purchased in 2011). The detergent was heat-inactivated before use, and the initial and residual amylase activities were determined using the Ceralpha .alpha.-amylase assay. The equipment used for this assay included a Biomek FX Robot (Beckman Coulter); a SpectraMAX MTP Reader (type 340-Molecular Devices), a Tetrad2DNA Engine PCR machine (Biorad), and iEMS incubator/shaker (Thermo Scientific). The reagent solutions used were as follows: [0639] 1) p-nitrophenyl maltoheptaoside (BPNPG7) substrate (Megazyme Ceralpha HR kit): [0640] 2) Liquid detergent (Persil color gel, enzyme inactivated by heating for 4 hrs at 90.degree. C.); [0641] 3) 50 mM MOPS, 0.1 mM CaCl.sub.2), 0.005% TWEEN.RTM.80 buffer, pH 7 (dilution buffer); [0642] 4) 10% detergent solution diluted in dilution buffer; [0643] 5) 200 mM Boric acid/NaOH buffer, pH 10 (STOP buffer) [0644] 6) Amylase culture supernatants diluted eight fold in 50 mM MOPS pH7.15, 0.1 mM CaCl.sub.2) containing 0-100 .mu.g/mL protein
[0645] 80 .mu.L of a 10% detergent solution was added to a 96-well PCR plate and mixed with 20 .mu.L of the diluted culture supernatant. A sample from the PCR plate was diluted 10.times. in dilution buffer and a 5 .mu.L aliquot of this dilution was used to determine initial amylase activity. The PCR plate was incubated in a Tetrad PCR block at 80.5.degree. C. for 5 minutes. After incubation, detergent-enzyme mix was diluted 10.times. in dilution buffer and residual activity was measured. Initial (t.sub.initial) and residual (t.sub.residual) amylase activity was determined in triplicate by the Ceralpha .alpha.-amylase assay using 5 .mu.L samples.
[0646] For each variant, the ratio of the residual and initial amylase activities was used to calculate the detergent stability as follows: Detergent stability=[t.sub.residual value]/[t.sub.initial value]. For each sample (variants) the performance index (PI) was calculated. The performance index for detergent stability is determined by comparing the detergent stability of the variant enzyme with that of the similarly treated CspAmy2-v1 enzyme. The results are shown in Table 5.
TABLE-US-00027 TABLE 5 Cleaning performance and stability of CspAmy2 variants Performance Index Variant Cleaning Stability CspAmy2-v1 1 1 CspAmy2-C16B 1.02 2.51 CspAmy2-C16D 1.32 2.37 CspAmy2-C16F 1.25 2.41 CspAmy2-C16H 1.2 1.17 CspAmy2-C16J 1.06 0.83
Example 10
Generation of Bacillus Strains Expressing CspAmy2 Combinatorial Variants
A. Design of Combinatorial Variants
[0647] Synthetic DNA encoding two additional CspAmy2 variants were constructed by GeneArt and delivered as plasmids transformed in a B. subtilis host, as described for CspAmy2-v5 and CspAmy2-v6 in Example 2. CspAmy2 v171 is a variant of CspAmy2 having the mutations T180D, E187P, I203Y, G476K, and lacking R178 and G179. CspAmy2 v172 is a variant of CspAmy2 having the mutations N126Y, T180D, E187P, I203Y, G476K, and lacking R178 and G179.
[0648] The amino acid sequence of mature CspAmy2-v171 is shown, below, as SEQ ID NO: 22:
TABLE-US-00028 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRNQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFDGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNKGSVSVWVQQ
[0649] The amino acid sequence of mature CspAmy2-v172 is shown, below, as SEQ ID NO: 23:
TABLE-US-00029 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFDGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNKGSVSVWVQQ
Example 11
Thermostability Assessment of CspAmy2 Combinatorial Variants
[0650] The cleaning detergent stability of purified CspAmy2-v171 and CspAmy2-v172 was compared to that of CspAmy2-v5, and ACE-QK, as described in Example 5. The results are shown in FIG. 11. The stability of CspAmy2-v172 was greater than all other tested enzymes.
Example 12
Cleaning Benefit of a CspAmy2 Combinatorial Variant in Hand Dishwashing
[0651] The cleaning benefit of a CspAmy2 variant was compared to that of a commercially-available .alpha.-amylase [STAINZYME.RTM. (Novozymes A/S)] in a hand dishwashing application. In an assay intended to simulate hand washing, 0.01% (wt/wt) active enzyme (CspAmy2-v6, STAINZYME.RTM., or no enzyme as a control) was added to European Dreft laundry detergent (1 g/L in water; Procter & Gamble, Cincinnati, Ohio, USA) and incubated in the presence of DM77 starch monitor (i.e., ADW melamine tiles from Center for Test Materials CFT BV in Vlaardingen, the Netherlands) at 40.degree. C. with agitation in Laundr-o-meter. The results are shown in FIG. 12. CspAmy2-v5 was far superior to STAINZYME.RTM. in cleaning performance in terms of the rate of cleaning observed and the end point reached. Importantly, the rate of cleaning achieved by CspAmy2-v5 was so rapid that benefits were observed within a period of time relevant to hand dishwahing applications (i.e., about 30 seconds). These results indicate that CspAmy2 variants have potential as additives for hand dishwashing compositions.
Example 13
Cleaning Benefit of a CspAmy2 Combinatorial Variant in Automatic Dishwashing
[0652] The cleaning benefit of a CspAmy2 variant was compared to that of two benchmark .alpha.-amylases (i.e., STAINZYME.RTM. and POWERASE.RTM.) in an automatic dishwashing (ADW) application. The cleaning assays were performed in a standard Miele 6382 dishwasher using a normal cycle (50.degree. C. and 60 minutes) with water having a hardness of 21 GH and 37.5 FH. 20 g of ADW powder detergent was used for each dishwashing cycle. The detergent was either WfK Type B or Type C (Testgewebe GmbH, Bruggen, Deutschland), which are described in Tables A and B, respectively of FIG. 13.
[0653] The test samples were either pasta or mixed-starch stained dishes. The pasta samples were prepared by mixing 150 g of strained pasta cooked in 17 GH water with 200 mL distilled water in a blender for 5 minutes to obtain a chewing gum-like suspension. About 3 g of the suspension was brushed onto the surface of each dish to be included in the cleaning assay, allowed to dry for 24 hours, baked onto the dishes at 120.degree. C. for 2 hours, and allowed to cool. Cleaning performance was evaluated by adding iodine to the washed dishes and using a photo scale rating of 1-10. The mixed starch samples were prepared by mixing 26 g each of potato, corn, rice, and wheat starch in about 4 L g 16 GH water and heating to 95.degree. C. for 10 minutes. After cooling to room temperature, about 30 mL of the mixture was applied to the surface of each dish to be included in the cleaning assay, allowed to dry on the surface of the dishes for 48 hours at room temperature, baked onto the dishes at 80.degree. C. for 1 hour, and allowed to cool. Cleaning performance was determined by weighing the dishes before and after the cleaning assay to determine the amount of starch removed.
[0654] FIGS. 14 and 15 show the cleaning performance of CspAmy2-v6 (squares) compared to POWERASE.RTM. (diamonds), dosed at 0, 1, 2, 4, or 8 ppm in about 2.5 cents/kg (ct/kg) WfK B detergent against the mixed starch stain (FIG. 14) and the pasta stain (FIG. 15). CspAmy2-v6 clearly outperformed POWERASE.RTM., particularly against the mixed starch stain. FIGS. 16 and 17 show the cleaning performance of CspAmy2-v6 (squares) compared to STAINZYME.RTM. (circles), dosed at 0, 1, 2, 4, or 8 ppm in about 2.5 ct/kg WfK B detergent against the mixed starch stain (FIG. 16) and the pasta stain (FIG. 17). CspAmy2-v6 clearly outperformed STAINZYME.RTM., especially at low doses. FIGS. 18 and 19 show the cleaning performance of CspAmy2-v6 (squares) compared to POWERASE.RTM. (diamonds), dosed at 0, 1, 2, 4, or 8 ppm in about 2.5 ct/kg WfK C detergent against the mixed starch stain (FIG. 18) and the pasta stain (FIG. 19). CspAmy2-v6 clearly outperformed POWERASE.RTM. against both stains.
Example 14
Site Evaluation Library Screen of C18P and Identification of Activity-Enhancing Mutations
[0655] Site evaluation libraries (SELs) were constructed and screened at 283 of 485 positions in variant CspAmy2-C18P (i.e., CspAmy2 with the mutations N126Y, F153W, T180D, I203Y, and S241Q, and lacking R178 and G179 (referring to SEQ ID NO: 1 for numbering) to identify additional variants with enhanced activity on one or more of the following substrates: corn amylopectin, swelled corn starch, granular corn starch, and corn starch-stained microswatches. Mutations were discovered throughout the molecule that improved activity. A subset of these variants was re-tested at different dose responses to confirm enhanced activity.
[0656] The amino acid sequence of the mature CspAmy2-C18P amylase polypeptide is shown, below, as SEQ ID NO: 24 (the substitutions are underlined):
TABLE-US-00030 AATNGTMMQY FEWYVPNDGQ QWNRLRTDAP YLSSVGITAV WTPPAYKGTS QADVGYGPYD LYDLGEFNQK GTVRTKYGTK GELKSAVNTL HSNGIQVYGD VVMNHKAGAD YTENVTAVEV NPSNRYQETS GEYNIQAWTG FNFPGRGTTY SNWKWQWFHF DGTDWDQSRS LSRIFKFDGK AWDWEVSSEN GNYDYLMYAD YDYDHPDVVN EMKKWGVWYA NEVGLDGYRL DAVKHIKFQF LKDWVDNARA ATGKEMFTVG EYWQNDLGAL NNYLAKVNYN QSLFDAPLHY NFYAASTGGG YYDMRNILNN TLVASNPTKA VTLVENHDTQ PGQSLESTVQ PWFKPLAYAF ILTRSGGYPS VFYGDMYGTK GTTTREIPAL KSKIEPLLKA RKDYAYGTQR DYIDNPDVIG WTREGDSTKA KSGLATVITD GPGGSKRMYV GTSNAGEIWY DLTGNRTDKI TIGSDGYATF PVNGGSVSVW VQQ
Corn Amylopectin Assay
[0657] Hydrolysis of soluble corn amylopectin was used to measure specific activity of amylase variants. Activity was measured as reducing ends generated by the enzymatic hydrolysis of amylopectin polymers as quantified using a bicinchoninic acid (BCA) assay. The equipment used for the assay included a Biomek FX Robot (Beckman Coulter), a SpectraMAX MTP Reader (type 340-Molecular Devices), and a Tetrad2DNA Engine PCR machine (Biorad).
[0658] Purified corn amylopectin (MP Biomedicals, LLC, cat. #195048) was solubilized by boiling for 5 minutes while stirring as a 1.5% (w/w) suspension in water. The material was allowed to cool to room temperature and water and concentrated stock solutions were added to obtain a final amylopectin substrate with 1.25% (w/w) amylopectin, 6.25 ppm calcium, 62.5 ppm sodium, 62.5 mM potassium acetate (pH 5.0), and 0.005% (v/v) Tween 80.
[0659] 40 .mu.l of the amylopectin substrate was added to 3 PCR microtiter plates. 10 .mu.l of culture supernatant diluted 1:2000 in water/0.005% tween 80 was added to plates followed by through mixing by up and down pipetting. The plates were sealed and placed at 70.degree. C. for 5 minutes followed by a ramp down to 25.degree. C. Amylopectin hydrolysis was terminated by immediate addition/mixing of 10 .mu.L 0.5 N NaOH.
[0660] The starch hydrolysis reaction products were analyzed by the BCA assay. Briefly, a reagent provided in a commercial kit (Pierce Chemical, cat. no. 23225) was prepared as per manufacturer's instructions. 90 .mu.l was aliquoted to PCR plates followed by 10 .mu.l of the terminated enzyme reaction described above. After through mixing of components the plates were sealed and placed in a thermocycler, programmed for 3 minutes at 95.degree. C. to develop color, and then cooled to 30.degree. C. Samples of 70 .mu.L of the developed BCA reaction mixtures were transferred to a fresh plate and absorbance was measured at 562 nm in a spectrophotometer. Data acquired for three replicate plates were averaged.
Corn Starch Assay
[0661] Cargill Corn Starch material (flour or starch) was washed extensively with MilliQ water by repeated suspension and centrifugation prior to use in the assay. The washed corn starch was suspended in MilliQ water containing 0.005% sodium azide to obtain 20% (w/w) stock solutions, which were further diluted with a 20.times. stock buffer solution to 10.9% w/v corn flour and corn starch solutions (final buffer concentration of 55 mM KOAc, pH 5).
[0662] 55 .mu.L of the diluted corn flour or corn starch substrates were added to PCR microtiter plates along with 5 .mu.L of 1:10 diluted enzyme samples using a bubble paddle reservoir. The plates were sealed and placed at 86.degree. C. for 5 minutes followed by a ramp down to 45.degree. C. The starch hydrolysis reaction was terminated by addition of 70 .mu.L 0.1 N NaOH. The plates were sealed and centrifuged for 3 minutes at 1,610 RCF. The starch hydrolysis reaction products were analyzed by the BCA assay as described above.
Swelled Corn Starch (SCS) Assay
[0663] The SCS assay measures alpha-amylase activity on hydrated but intact ("swelled") starch granules. The overall number of enzymatic turnovers is assessed using the BCA reducing sugar assay, while the tendency of the enzyme to release large starch fragments into solution is assessed using iodine staining.
[0664] 2% (w/w) swelled corn starch substrate was prepared by suspending 2 g of corn starch (Cargill Farms Organic Corn Starch) in 90 g of MilliQ water and heating in a submerged water bath set to 80.degree. C. for 1 hour with regular swirling. After overnight cooling at room temperature, the volume was brought up to 100 mL with the addition of potassium acetate buffer, calcium chloride, sodium chloride and Tween-80 to give final concentrations of 50 mM KOAc (pH5.5), 0.125 mM CaCl.sub.2), 2 mM NaCl, and 0.005% Tween-80.
[0665] 55 .mu.L of swelled corn starch substrate was dispensed into NUNC V-bottom polystyrene microtiter plates using a bubble paddle reservoir. The reaction was initiated by adding 5 .mu.L of 12.times. enzyme solution to the plate to give a final enzyme concentration of -0.03 ppm in reaction. The reaction plate was sealed with a Nunc seal and immediately placed in an iEMS incubator. Incubation occurred for 10 min at 25.degree. C., or for 4 min at 60.degree. C., while shaking at 1,150 RPM. After incubation, the reaction was terminated with the addition of 70 .mu.L of 0.1 N NaOH. The plates were sealed and spun for 3 min at 3,000 RPM. Swelled corn starch reactions were done as a single plate, but assayed in triplicate for subsequent Iodine and BCA assays.
[0666] The BCA assay is described above. For the iodine reaction, 95 .mu.L of Lugol's reagent (freshly diluted by 12-fold in water; Sigma-Aldrich L6146-1L) was added to 96-well Costar 9017 microtiter plates. 5 .mu.L of supernatant sample was added to the plates and mixed six times by pipetting up and down. The plates were then shaken on a table top microtiter plate mixer for 1 minute at speed 6-7. Absorbance was read at 530 nm using a SpectraMax M5 spectrophotometer.
CS-26 Corn Starch Microswatch Assay
[0667] The assays were performed as described, above, except that 170 .mu.l of potassium acetate buffer was added to each well of microswatch-containing MTP and subsequently 10 .mu.L of diluted enzyme solution was added to each well resulting in a total volume of 180 .mu.L/well. The MTP was sealed and incubated for 20 minutes at 1,150 rpm at 25.degree. C.
Viscosity Analysis
[0668] Viscosity experiments were performed using a Rapid Visco Analyser (Newport Scientific) with 38 mm.times.68 mm plain washed cans (Part No. AA0384001) and a double-skirted paddle (Part no. NS101783). Data acquisition and analysis was performed with Thermocline for Windows (version 3.11). Immediately before each run, 33 g of 25% dry solids (ds) corn flour slurry was prepared in an RVA can as follows: 9.17 g corn flour (11.8% moisture content) was weighed out on an analytical balance and mixed with 23.83 g deionized water. The sample pH was adjusted with 0.0828 mL 1N sulfuric acid (for pH 5.7) or 0.285 mL 1N sulfuric acid (for pH 5.2). Enzyme was added at appropriate dose, and the can was placed in the viscometer. All runs were 10 minutes in length, with a temperature ramp to 85.degree. C. or 95.degree. C. over 80 seconds followed by a temperature hold at 85.degree. C. or 95.degree. C. for the remainder of the run.
Initial Screening Results
Mutations resulting in variants with expression greater than 250 ug/mL and at least 110% of the performance of CspAmy2-C18P in any one of the indicated assays in each assay were classified as performance-enhancing mutations. CspAmy2-C18P SEL variants with mutations at positions 6, 7, 8, 11, 14, 15, 20, 21, 23, 26, 27, 28, 37, 38, 39, 40, 42, 45, 46, 48, 49, 50, 51, 52, 53, 54, 58, 61, 62, 68, 70, 71, 72, 73, 79, 80, 81, 82, 84, 85, 87, 88, 89, 92, 93, 94, 95, 96, 97, 98, 101, 108, 111, 112, 113, 114, 115, 116, 117, 118, 120, 122, 123, 124, 126, 127, 129, 130, 131, 132, 133, 134, 136, 137, 138, 140, 142, 143, 144, 147, 148, 149, 150, 151, 152, 153, 154, 155, 156, 158, 159, 165, 167, 168, 170, 171, 172, 175, 176, 177, 180, 181, 182, 187, 190, 191, 193, 199, 200, 201, 203, 206, 208, 210, 211, 212, 214, 215, 216, 219, 221, 223, 225, 226, 227, 235, 238, 239, 240, 241, 242, 243, 245, 246, 247, 248, 249, 250, 252, 253, 254, 256, 257, 258, 260, 261, 262, 266, 267, 268, 269, 270, 271, 273, 276, 277, 279, 280, 282, 284, 285, 286, 288, 296, 299, 300, 301, 302, 303, 304, 307, 308, 310, 311, 312, 313, 316, 317, 318, 320, 321, 325, 327, 335, 338, 342, 348, 349, 352, 356, 357, 360, 362, 363, 368, 369, 377, 381, 382, 383, 384, 385, 388, 390, 392, 394, 395, 396, 397, 398, 400, 401, 402, 403, 404, 405, 407, 408, 410, 414, 415, 416, 418, 419, 420, 421, 422, 423, 424, 426, 428, 429, 430, 431, 434, 435, 436, 439, 441, 442, 444, 445, 446, 447, 448, 449, 450, 451, 454, 455, 457, 460, 461, 462, 463, 464, 465, 466, 467, 469, 470, 471, 473, 474, 475, 476, 477, 479, 480, 481, 482, 483, and 484 met this criteria. Specific performance-enhancing mutations included T6A, T6D, T6E, T6G, T6K, T6M, T6N, T6Q, T6S, M7A, M7V, MBC, MBF, M8I, M8L, M8Y, F11V, Y14A, Y14I, Y14Q, Y14T, Y14V, V15C, V15D, V15I, V15N, V15T, Q20A, Q20C, Q20D, Q20H, Q20K, Q20M, Q2ON, Q20R, Q205, Q20Y, Q21F, Q21W, N23A, N23C, N23D, N23E, N23H, N23K, N23Q, N23R, N23S, N23T, N23V, R26C, R26E, R26G, R26K, R26M, R26S, R26T, T27A, T27C, T27D, T27E, T27F, T27H, T27I, T27K, T27L, T27M, T27N, T27Q, T27R, T27S, T27V, T27Y, D28A, D28C, D28T, I37F, I37V, T38D, T38N, A39L, V40A, V40C, V40D, V40G, V40H, V401, V40K, V40M, V40P, V40Q, V40R, V405, V40T, V40W, V40Y, T42A, T42C, T421, T42M, T42V, A45C, A45G, Y46F, G48A, T49I, S50A, S50C, S50E, S50G, S50K, S50M, S50N, S50Q, 550R, S50T, S50Y, Q51C, Q51D, Q51E, Q51S, Q51V, A52C, A52D, A52E, A52F, A52G, A52H, A52K, A52L, A52M, A52R, A52S, A52T, A52V, A52Y, D53E, V54N, V54T, P58A, P58C, P58H, P581, P58S, P58T, P58V, L61M, L61V, Y62A, Y62C, Y62D, Y62F, Y62G, Y62H, Y62I, Y62K, Y62L, Y62M, Y62N, Y62P, Y62Q, Y62R, Y62S, Y62V, N68C, N68D, N68E, N68F, N68H, N68L, N68P, N68Q, N68R, N68S, N68V, N68W, N68Y, K70R, G71A, G71C, G71D, G71E, G71K, G71R, G71S, T72G, T72S, V73S, T79F, T791, T79L, T79M, T79N, T79S, T79Y, K80A, K80C, K80D, K80F, K80H, K80I, K80M, K80N, K80Q, K80R, K80S, K80T, K80V, K80Y, G81A, G81D, G81E, G81F, G81H, G81I, G81K, G81N, G81P, G81R, G81S, G81T, E82A, E82D, E82M, E82Q, K84A, K84C, K84E, K84I, K84Q, K84R, K84S, K84T, K84Y, S85A, S85C, S85D, S85E, S85G, S85H, 5851, S85L, S85M, S85N, S85Q, S85R, S85T, S85V, S85Y, V871, V87T, N88C, N88D, N88E, N88G, N88H, N88I, N88K, N88L, N88M, N88Q, N88R, N88S, N88T, N88V, N88W, N88Y, T89A, T89D, T89E, T89F, T89H, T891, T89K, T89L, T89M, T89N, T89Q, T89R, T89S, T89Y, S92A, S92C, S92D, S92E, S92F, S92G, S92H, S92L, S92M, S92N, S92Q, S92R, S92T, S92W, S92Y, N93A, N93C, N93E, N93F, N93H, N93I, N93K, N93L, N93Q, N93S, N93T, N93Y, G94A, G94C, G94N, I95M, Q96A, Q96E, Q96H, Q961, Q96K, Q96L, Q96M, Q96N, Q96R, Q96V, Q96Y, V971, V97T, Y98F, Y981, Y98L, Y98V, V101C, V101T, G108A, G108S, Y111D, Y111E, Y111L, Y111N, Y111S, Y111T, Y111V, T112A, T112F, T112H, T1121, T112K, T112L, T112M, T112N, T112P, T112R, T112V, T112W, E113D, E113N, E113Q, E113T, N114A, N114C, N114D, N114E, N114F, N114G, N114H, N114I, N114L, N114P, N114Q, N114R, N114S, N114T, N114V, N114W, N114Y, V115A, V115I, T116A, T116C, T116D, T116G, T116H, T116I, T116K, T116N, T116P, T116Q, T116R, T116S, A117C, A117I, A117S, A117V, V118A, V118C, V118E, V118F, V118H, V118I, V118L, V118M, V118N, V118Q, V118R, V118S, V118W, V118Y, V120A, V120C, V1201, V120M, V120T, P122A, P122D, P122E, P122G, P122H, P122N, P122R, P122S, P122T, P122W, S123A, S123C, S123G, S123K, S123L, N124A, N124D, N124F, N124G, N124L, N124R, N124S, N124V, Y126A, Y126C, Y126D, Y126E, Y126G, Y126H, Y1261, Y126K, Y126L, Y126M, Y126N, Y126Q, Y126R, Y126S, Y126T, Y126V, Y126W, Q127A, Q127C, Q127D, Q127E, Q127F, Q127H, Q1271, Q127K, Q127L, Q127M, Q127N, Q127R, Q127S, Q127T, Q127V, Q127W, Q127Y, T129A, T129C, T129D, T129E, T129F, T129G, T129I, T129K, T129L, T129M, T129N, T129Q, T129R, T129S, T129V, T129W, T129Y, S130A, S130G, 51301, S130K, S130L, S130M, S130N, S130P, S130R, S130T, S130V, S130W, G131A, G131C, G131D, G131F, G131H, G131I, G131K, G131L, G131M, G131P, G131Q, G131V, G131W, G131Y, E132A, E132C, E132D, E132F, E132G, E132H, E132I, E132K, E132L, E132M, E132N, E132P, E132Q, E132R, E132V, E132W, Y133A, Y133D, Y133E, Y133G, Y133K, Y133L, Y133M, Y133R, Y133S, Y133T, Y133W, N134A, N134D, N134F, N134G, N134H, N134I, N134K, N134L, N134M, N134R, N134S, N134V, N134W, N134Y, Q136A, Q136C, Q136D, Q136E, Q136F, Q136H, Q136K, Q136L, Q136M, Q136N, Q136R, Q136S, Q136T, Q136V, Q136W, Q136Y, A137S, A137T, A137V, W138F, W138Y, G140A, G140M, N142F, N142K, N142L, N142M, N142P, N142V, N142W, N142Y, F143H, F143Y, P144C, P144D, P144E, P144G, P144H, P144I, P144K, P144L, P144M, P144N, P144Q, P144S, P144T, P144Y, G147A, G147C, G147H, G147K, G147L, G147M, G147N, G147Q, G147R, T148A, T148D, T148E, T148F, T1481, T148K, T148L, T148R, T148S, T148V, T148W, T149A, T149C, T149D, T149E, T149F, T149H, T149K, T149L, T149M, T149N, T149Q, T149V, T149W, T149Y, Y150H, S151A, S151E, S151F, S151H, S151I, S151K, S151L, S151M, S151Q, S151R, S151V, N152A, N152C, N152D, N152E, N152G, N152H, N152K, N152M, N152P, N152Q, N152R, N152S, N152T, W153F, W153H, W153Q, W153R, W153T, W153Y, K154A, K154C, K154D, K154E, K154G, K1541, K154L, K154M, K154N, K154R, K154T, K154Y, W155P, Q156A, Q156D, Q156E, Q156F, Q156G, Q156H, Q1561, Q156L, Q156M, Q156N, Q156R, Q156S, Q156Y, F158C, F158D, F158K, F158L, F158M, F158N, F158P, F158Q, F158R, F158S, F158V, H159M, H159Y, W165C, W165D, W165E, W165F, W165H, W1651, W165K, W165L, W165M, W165Q, W165R, W165T, W165Y, Q167A, Q167D, Q167E, Q167G, Q167H, Q167K, Q167M, Q167N, Q167P, Q167S, Q167T, Q167V, S168A, S168D, S168F, S168H, 51681, S168K, S168M, S168N, S168Q, S168R, S168T, S168V, S168W, S168Y, S170A, S170C, S170D, S170E, S170F, S170L, S170M, S170N, S170Q, S170R, S170T, L171A, L171C, L171F, L171G, L171H, L171I, L171N, L171Q, L171R, L171T, L171V, L171W, L171Y, S172A, S172D, S172H, S172R, S172T, F175M, F175Y, K1761, K176T, F177L, F177V, F177W, D180A, D180C, D180F, D180G, D180H, D1801, D180L, D180M, D180N, D180Q, D180R, D180S, D180T, D180V, D180W, D180Y, G181A, G181C, G181D, G181E, G181F, G181H, G181K, G181L, G181M, G181N, G181Q, G181R, G181S, G181T, G181V, G181Y, K182A, K182P, E187D, E1871, E187K, E187M, E187N, E187P, E187R, E187S, E187T, E187V, E187Y, S190A, S190C, S190D, S190F, S190L, S190N, S190P, S190Q, E191A, E191C, E191F, E191G, E191H, E1911, E191K, E191N, E191Q, E191R, E191S, E191T, E191W, E191Y, G193A, G193C, G193D, G193E, G193F, G193H, G193K, G193L, G193M, G193N, G193Q, G193R, G193S, G193T, G193V, G193W, M199L, Y200F, A201M, Y2031, Y203L, Y203V, D206A, D206E, D206G, D206H, D206M, D206N, D206Q, D206R, D206S, D206T, P208A, P208E, P208F, P2081, P208K, P208L, P208T, P208V, P208Y, V210A, V210E, V210H, V210K, V210N, V210Q, V210R, V210S, V210T, V211A, V211E, V211H, V2111, V211Q, V211R, N212E, N212F, N212G, N212L, N212M, N212R, N212V, M2141, M214L, K215C, K215E, K215F, K215M, K215N, K215R, K215Y, K216F, V2191, V219T, Y221F, Y2211, Y221L, N223C, N223E, N223I, N223K, N223Q, N223R, N223 S, N223T, N223V, N223W, N223Y, V225A, V2251, V225L, V225M, G226D, G226M, G226Q, G226R, G226S, L227F, L227I, L227W, L227Y, V235A, I238A, I238L, I238M, K239D, K239E, K239P, K239Q, K239R, K239S, K239T, F240K, F240L, F240M, F240Q, F240R, Q241A, Q241C, Q241D, Q241E, Q241G, Q241H, Q241K, Q241L, Q241M, Q241N, Q241P, Q241R, Q241S, Q241T, Q241V, Q241W, Q241Y, F242V, L243C, L243Y, D245A, D245C, D245E, D245G, D245L, D245M, D245N, W246F, V2471, V247L, D248E, D248H, D248N, D248T, D248V, N249A, N249E, N249G, N249H, N249Q, N249Y, A250M, A250S, A250V, A252C, A252D, A252E, A252G, A252H, A2521, A252K, A252L, A252M, A252N, A252Q, A252S, A252V, A252W, A252Y, A253E, A2531, A253K, A253L, A253M, A253Q, A253S, A253T, A253V, A253Y, T254F, T254K, T254S, K256A, K256M, K256N, K256S, E257Q, E257S, M258L, T260A, T260C, T260S, T260V, V261I, V261W, G262A, Q266A, Q266D, Q266E, Q266H, Q2661, Q266M, Q266N, Q266S, Q266T, Q266V, Q266Y, N267H, N267I, N267Q, N267R, N267S, N267T, N267V, N267Y, D268G, D268N, L269C, L269D, L269I, L269K, L269Q, L269S, L269T, L269Y, G270A, G270D, G270E, G270F, G270H, G270I, G270L, G270M, G270Q, G270T, G270V, G270Y, A271C, A271D, A271E, A271H, A271K, A271M, A271Q, A271R, A271S, A271T, A271V, A271Y, N273C, N273G, N273H, N273I, N273K, N273R, L2761, L276M, A277C, A277D, A277E, A277F, A277G, A277I, A277K, A277L, A277M, A277N, A277Q, A277R, A277S, A277T, A277V, A277W, A277Y, V279C, V279T, N280A, N282S, N282T, S284T, S284Y, L285A, L285C, L2851, L285V, F286M, A288C, A296C, A296D, A296E, A296F, A296G, A296H, A2961, A296L, A296M, A296N, A296Q, A296R, A296S, A296V, A296W, A296Y, T299A, T299D, T299E, T299F, T299G, T299K, T299L, T299M, T299R, T299S, T299V, T299W, G300A, G300C, G300D, G300E, G300F, G300H, G300K, G300M, G300Q, G300R, G300V, G300W, G301A, G301C, G301D, G301E, G301F, G301H, G301K, G301L, G301M, G301Q, G301R, G301S, G301T, G301V, G301W, G301Y, G302S, Y303A, Y303C, Y303D, Y303E, Y303F, Y303G, Y303H, Y3031, Y303K, Y303L, Y303M, Y303N, Y303Q, Y303R, Y303S, Y303T, Y303V, Y303W, Y304F, Y304K, Y304W, R307A, R307C, R307E, R307G, R307H, R307K, R307M, R307N, R307Q, R307S, R307T, N308C, N308D, N308E, N308G, N308L, N308M, N308T, N308V, L310A, L310C, L310D, L310E, L310H, L310I, L310M, L310P, L310W, L310Y, N311C, N311E, N311G, N311H, N311K, N311Q, N311R, N311S, N311V, N311W, N311Y, N312D, N312F, N312G, N312H, N312K, N312Q, N312R, T313A, T313S, A316D, A316E, A316G, A316H, A316K, A316Q, A316R, A316Y, S317C, S317D, S317G, S317H, S317K, S317L, S317N, S317Q, S317R, S317T, S317W, S317Y, N318A, N318C, N318F, N318G, N318I, N318K, N318L, N318M, N318Q, N318R, N318S, N318T, N318V, N318W, T320A, T320C, T320D, T320E, T320G, T320H, T320I, T320K, T320N, T320P, T320Q, T320R, T320V, T320W, T320Y, K321C, K321F, K321H, K321N, K321S, K321Y, L325A, L325C, L325F, L3251, L325M, L325Q, L325V, E327D, E327L, Q335C, Q335E, E338A, E338D, E338F, E338G, E338H, E338I, E338K, E338P, E338Q, E338R, E338T, E338V, E338Y, Q342A, Q342C, Q342G, Q342L, Q342M, Q342R, Q342S, Q342T, Q342V, Q342W, L348A, L348C, L348H, L3481, L348M, L348Q, L348S, L348T, A349G, A349R, F3521, F352L, F352M, F352T, F352V, R356Q, S357A, S357C, S357D, S357E, S357F, S357H, 53571, S357K, S357L, S357N, S357Q, S357T, S357V, S357W, S357Y, Y360F, Y3601, Y360L, Y360M, Y360V, S362A, S362C, S362E, 53621, S362T, S362V, V363I, V363L, M368F, M368I, M368L, M368Y, Y369A, Y369E, Y369I, Y369L, Y369N, Y369V, R377A, R377C, R377D, R377E, R377F, R377G, R377H, R3771, R377K, R377L, R377N, R377Q, R377S, R377T, R377V, R377W, R377Y, A381C, A381E, A381G, A381H, A381K, A381L, A381M, A381N, A381P, A381Q, A381V, A381W, A381Y, L382F, L382H, L382Q, L382S, K383A, K383C, K383D, K383E, K383H, K3831, K383L, K383M, K383N, K383Q, K383R, K383S, K383W, K383Y, S384A, S384C, S384D, S384F, S384G, S384H, S3841, S384L, S384N, S384R, S384V, S384Y, K385A, K385D, K385E, K385F, K385G, K385H, K385L, K385M, K385Q, K385R, K385T, K385V, K385Y, P388A, P388C, P388D, P388I, P388L, P388N, P388R, P388S, P388T, P388V, L390M, L390V, A392C, A392S, K394A, K394C, K394E, K394F, K394G, K394H, K3941, K394L, K394Q, K394R, K394S, K394T, K394V, K394W, K394Y, D395A, D395E, D395N, D395S, D395T, Y396A, Y396D, Y396F, Y396K, Y396M, Y396N, Y396Q, Y396T, Y396V, Y396W, A397D, A397E, A397H, A397K, A397M, A397N, A397Q, A397V, Y398A, Y398C, Y398F, Y398H, Y3981, Y398L, Y398W, T400A, T400C, T400D, T400F, T400G, T400I, T400K, T400L, T400M, T400N, T400Q, T400R, T400W, T400Y, Q401A, Q401C, Q401F, Q401H, Q401I, Q401L, Q401M, Q401N, R402C, R402D, R402F, R402K, R402L, R402M, R402N, R402Q, R402S, R402W, D403E, D403N, D403S, Y404E, Y404G, Y404K, Y404M, Y404N, Y404R, Y404W, I405C, I405F, I405L, I405M, I405V, N407A, N407C, N407D, N407E, N407G, N407K, N407M, N407Q, N407R, P408A, P408C, P408D, P4081, P408K, P408L, P408M, P408N, P408Q, P408V, P408W, V410A, V410C, V410D, V410E, V410F, V410H, V4101, V410L, V410M, V410N, V410Q, V410S, V410T, T414A, T4141, T414S, T414V, R415M, E416C, E416D, E416F, E416H, E4161, E416K, E416L, E416M, E416N, E416Q, E416R, E416T, E416V, E416W, E416Y, D418A, D418E, D418G, D418H, D4181, D418K, D418L, D418M, D418N, D418Q, D418S, D418T, D418V, D418W, S419A, S419C, S419E, S419G, S419L, S419M, S419N, S419R, S419V, S419W, S419Y, T420A, T420C, T420D, T420E, T420G, T420H, T420I, T420K, T420M, T420P, T420S, T420V, T420W, T420Y, K421A, K421D, K421E, K421H, K4211, K421L, K421M, K421N, K421P, K421Q, K421R, K421T, K421V, K421W, K421Y, A422C, A422D, A422E, A422F, A422G, A4221, A422L, A422N, A422P, A422Q, A422R, A422S, A422Y, K423A, K423D, K423E, K423F, K423H, K4231, K423L, K423M, K423N, K423Q, K423R, K423S, K423T, K423V, K423W, K423Y, S424A, S424C, S424G, S424K, S424N, S424Q, S424R, S424T, L426S, L426T, L426V, T428G, T428V, V429A, V429C, V4291, V429L, I430C, I430G, I430L, I430M, I430Q, I430V, T431A, T431C, T431S, P434A, P434C, P434D, P434E, P434F, P434H, P4341, P434K, P434L, P434M, P434N, P434Q, P434R, P434S, P434V, P434Y, G435A, G435C, G435D, G435E, G435F, G435H, G4351, G435K, G435M, G435N, G435P, G435Q, G435R, G435S, G435T, G435W, G436F, G4361, G436M, G436N, G436Q, G436S, G436V, R439A, R439D, R439G, R439H, R439K, R439M, R439N, R439P, R439Q, R439S, R439V, R439W, R439Y, Y441A, Y441C, Y441D, Y441F, Y441G, Y441H, Y441K, Y441L, Y441M, Y441N, Y441P, Y441R, Y441S, Y441T, Y441W, V442A, V442C, V4421, V442T, T444C, T444D, T444E, T444F, T444G, T444H, T4441, T444K, T444L, T444M, T444N, T444P, T444R, T444S, T444W, S445A, S445C, S445E, S445G, S445H, S445K, S445L, S445M, S445N, S445T, S445V, N446A, N446C, N446H, N446K, A447C, A447D, A447F, A447H, A447L, A447M, A447N, A447Q, A447R, A447S, A447Y, G448A, G448C, G448D, G448E, G448H, G448K, G448L, G448M, G448N, G448Q, G448R, G448S, G448T, G448W, E449D, E449H, E449K, E449T, I450A, I450C, I450D, I450E, I450G, I450K, I450L, I450M, I450N, I450Q, I450S, I450T, I450W, I450Y, W451Y, L454A, L4541, L454K, L454M, L454W, T455A, T4551, T455L, T455S, N457H, N457K, N457R, N457T, N457V, N457Y, D460A, D460E, D460G, D460M, D460N, D460Q, D460S, D460V, K461C, K461H, K461L, K461M, K461N, K461Q, K461T, K461Y, I462A, I462L, I462M, I462Q, I462T, I462V, T463D, T463E, T463H, T463P, T463Q, T463R, T463V, T463Y, I464P, I464T, G465A, G465C, G465D, G465E, G465K, G465L, G465M, G465N, G465Q, G465W, G465Y, S466A, S466C, S466D, S466G, S466H, S466K, S466L, S466M, S466N, S466T, S466W, S466Y, D467E, D467G, D467L, Y469A, Y469D, Y469E, Y4691, Y469M, Y469N, Y469R, Y469S, Y469T, Y469V, Y469W, A470S, A470V, T471A, T471C, T471E, T471G, T471H, T4711, T471L, T471M, T471N, T471S, T471V, T471W, P473C, P473D, P473E, P473G, P4731, P473K, P473L, P473R, P473T, P473W, V474A, V474C, V474L, V474S, N475A, N475C, N475E, N475F, N475H, N475K, N475L, N475M, N475P, N475Q, N475R, N475S, N475T, N475V, G476A, G476D, G476E, G476F, G476H, G4761, G476L, G476M, G476N, G476P, G476Q, G476R, G476S, G476T, G476V, G476W, G476Y, G477A, G477D, G477F, G477H, G4771, G477K, G477L, G477M, G477Q, G477S, G477T, G477V, G477W, G477Y, V479C, V479D, V479E, V479F, V479H, V4791, V479N, V479P, V479Y, S480A, S480C, S480H, V481A, V481C, V481N, W482Y, V483A, V483G, V4831, V483K, V483L, V483M, V483R, V483Y, Q484A, Q484C, Q484F, Q484G, Q484H, Q484K, Q484L, Q484M, Q484P, Q484R, Q484T, and Q484Y.
[0669] Dose Response Screening Results
[0670] A subset of the variants with performance-enhancing mutations were tested more rigorously to demonstrate improved activity relative to wild type at multiple protein concentrations (i.e., improved specific activity). FIG. 20 shows examples of variants with improved hydrolysis of corn starch at high temperature. FIG. 21 shows examples of variants with improved hydrolysis of amylopectin from corn. FIG. 22 shows examples of variants with improved generation of reducing sugars from starch. FIG. 23 shows examples of variants with improved release of iodine staining material from starch. In FIGS. 20-23, the legends adjacent to the graphs indicate the mutations present in addition to those present to the control variant, CspAmy2-C18P. FIG. 24 shows examples of variants that demonstrate improved reduction of corn slurry viscosity compared to the CspAmy2-C18P control. CspAmy2-C16F is also included for comparison.
Example 15
Pair-Wise Combinations of Mutations at Positions 476 and 477
[0671] Site evaluation libraries were constructed and screened to determined the effect of pair-wise combinations of mutations at positions 476 and 477 in variant CspAmy2-C16F (CspAmy2 with the mutations N126Y, F153W, T180H, I203Y, and S241Q, and lacking R178 and G179). The matrix in FIG. 25 shows the PI values for CspAmy2-C16F position 476/477 variants compared to a CspAmy2-C16F control (with a PI value set at 1) in a CS-26 corn starch microswatch assay, as described, above. The amino acid residues at positions 476 and 477 are indicated on the left-side and top of the matrix, respectively. A "wild-type" revertant, i.e., G476G/G477G had an experimentally obtained PI score of 1.01, suggesting that the assay produces very reliable results. The absence of a number at a position in the matrix indicates poor expression of the particular variant or that the variant was not present among those tested. The matrix in FIG. 26 shows the PI values for the same variants in an amylose hydrolysis assay, as described, above. Again, a "wild-type" revertant, i.e., G476G/G477G had an experimentally obtained PI score corresponding to that of the control.
[0672] Remarkably, the results suggest that almost any combination of residues at positions 476 and 477, other the glycine pair that occurs in naturally in CspAmy2 (and in many .alpha.-amylases), increases the performance of the .alpha.-amylase in terms of improving the hydrolysis of insoluble amylopectin and improving the release of iodine staining material from hydrated starch. Without being bound to a theory, it is believed that the adjacent GG residues at the C-terminus of .alpha.-amylases contribute to starch-binding. While binding tightly to a substrate may be desirable in nature where substrate is limiting, in industrial applications it may be more desirable for the enzyme to release from one starch molecule after hydrolyzing it and find a different starch molecules to hydrolize, rather than remaining associated with the first starch molecule and hydrolysing it processively.
Example 16
Additional C18P-Based Variants with Superior Performance in Secondary Liquifaction
[0673] Site evaluation libraries (SELs) were constructed and screened to determined the effect of mutations at positions E132, Q167, A277, and T400 in CspAmy2-C18P. The residues present at these positions in the best tested variants are shown in Table 6.
TABLE-US-00031 TABLE 6 Combinations of mutations tested Name E132 A277 Q167 T400 C16F E A Q T C25F H F Q T C25B H F E T C25A H F E K
[0674] The relative performance of CspAmy2-C25A, B, and F in a liquefaction assay compared to C18F is shown in FIG. 27. Mutations at positions 132 and 277 increase performance. Additional benefit is observed from mutations at positions 167. In further experiments, it was shown that C25B demonstrated superior liquefaction performance to C16F at pH 5.2 and 5.8, and with or without additional calcium. Variants CspAmy2-C25A, B, and F all outperformed variant CspAmy2-C18P (not shown).
[0675] The amino acid sequence of the mature CspAmy2-C25A amylase polypeptide is shown, below, as SEQ ID NO: 25 [the relevant substitutions are underlined]:
TABLE-US-00032 AATNGTMMQY FEWYVPNDGQ QWNRLRTDAP YLSSVGITAV WTPPAYKGTS QADVGYGPYD LYDLGEFNQK GTVRTKYGTK GELKSAVNTL HSNGIQVYGD VVMNHKAGAD YTENVTAVEV NPSNRYQETS GHYNIQAWTG FNFPGRGTTY SNWKWQWFHF DGTDWDESRS LSRIFKFDGK AWDWEVSSEN GNYDYLMYAD YDYDHPDVVN EMKKWGVWYA NEVGLDGYRL DAVKHIKFQF LKDWVDNARA ATGKEMFTVG EYWQNDLGAL NNYLFKVNYN QSLFDAPLHY NFYAASTGGG YYDMRNILNN TLVASNPTKA VTLVENHDTQ PGQSLESTVQ PWFKPLAYAF ILTRSGGYPS VFYGDMYGTK GTTTREIPAL KSKIEPLLKA RKDYAYGKQR DYIDNPDVIG WTREGDSTKA KSGLATVITD GPGGSKRMYV GTSNAGEIWY DLTGNRTDKI TIGSDGYATF PVNGGSVSVW VQQ
[0676] The amino acid sequence of the mature CspAmy2-C25B amylase polypeptide is shown, below, as SEQ ID NO: 26 [the relevant substitutions are underlined]:
TABLE-US-00033 AATNGTMMQY FEWYVPNDGQ QWNRLRTDAP YLSSVGITAV WTPPAYKGTS QADVGYGPYD LYDLGEFNQK GTVRTKYGTK GELKSAVNTL HSNGIQVYGD VVMNHKAGAD YTENVTAVEV NPSNRYQETS GHYNIQAWTG FNFPGRGTTY SNWKWQWFHF DGTDWDESRS LSRIFKFDGK AWDWEVSSEN GNYDYLMYAD YDYDHPDVVN EMKKWGVWYA NEVGLDGYRL DAVKHIKFQF LKDWVDNARA ATGKEMFTVG EYWQNDLGAL NNYLFKVNYN QSLFDAPLHY NFYAASTGGG YYDMRNILNN TLVASNPTKA VTLVENHDTQ PGQSLESTVQ PWFKPLAYAF ILTRSGGYPS VFYGDMYGTK GTTTREIPAL KSKIEPLLKA RKDYAYGTQR DYIDNPDVIG WTREGDSTKA KSGLATVITD GPGGSKRMYV GTSNAGEIWY DLTGNRTDKI TIGSDGYATF PVNGGSVSVW VQQ
[0677] The amino acid sequence of the mature CspAmy2-C25F amylase polypeptide is shown, below, as SEQ ID NO: 27 [the relevant substitutions are underlined]:
TABLE-US-00034 AATNGTMMQY FEWYVPNDGQ QWNRLRTDAP YLSSVGITAV WTPPAYKGTS QADVGYGPYD LYDLGEFNQK GTVRTKYGTK GELKSAVNTL HSNGIQVYGD VVMNHKAGAD YTENVTAVEV NPSNRYQETS GHYNIQAWTG FNFPGRGTTY SNWKWQWFHF DGTDWDQSRS LSRIFKFDGK AWDWEVSSEN GNYDYLMYAD YDYDHPDVVN EMKKWGVWYA NEVGLDGYRL DAVKHIKFQF LKDWVDNARA ATGKEMFTVG EYWQNDLGAL NNYLFKVNYN QSLFDAPLHY NFYAASTGGG YYDMRNILNN TLVASNPTKA VTLVENHDTQ PGQSLESTVQ PWFKPLAYAF ILTRSGGYPS VFYGDMYGTK GTTTREIPAL KSKIEPLLKA RKDYAYGTQR DYIDNPDVIG WTREGDSTKA KSGLATVITD GPGGSKRMYV GTSNAGEIWY DLTGNRTDKI TIGSDGYATF PVNGGSVSVW VQQ
Example 17
Additional CspAmy2-v5-Based Variants with Superior Performance in Cleaning Applications
[0678] Additional CspAmy2-v5-based variants were made in an effort to further improve performance in cleaning applications. The variants and the mutations are shown in the Table 7. CspAmy2-v171 and CspAmy2-172 were previously described in Example 11. All variants included deletions at positions R178 and G179, indicated by "del (R178, G179)."
TABLE-US-00035 TABLE 7 Additional CspAmy2-v5-based variants Variant Mutations CspAmy2-v5 del (R178, G179) + E187P + I203Y + G476K CspAmy2-v171 del (R178, G179) + T180D + E187P + I203Y + G476K CspAmy2-v172 del (R178, G179) + N126Y + T180D + E187P + I203Y + G476K CspAmy2-v179 del (R178, G179) + N126Y + T180D + E187P + I203Y + Y303D + G476T + G477E CspAmy2-v180 del (R178, G179) + N126Y + T180D + E187P + I203Y + Y303D + N475E + G477Q CspAmy2-v181 del (R178, G179) + N126Y + T180D + E187P + I203Y + Y303R + N475E + G476T + G477R CspAmy2-v186 del (R178, G179) + T38N + N88H + N126Y + T129I + N134M + F153W + L171R + T180D + E187P + I203Y + G476K + G477E CspAmy2-v191 del (R178, G179) + N126Y + E132H + T180D + E187P + I203Y + Y303D + G476T + G477E
[0679] The cleaning performance of the purified variants was analyzed in a microswatch cleaning assay performed essentially as described above. CFT CS-28 swatches were punched to form discs measuring 5.5 mm in diameter. Two discs were placed in each well of 3 each flat-bottom, non-binding 96-well assay plates. The CspAmy2 variants, STAINZYME.RTM., and ACE-QK were each diluted to 0.5 mg/mL in dilution buffer (50 mM MOPS (pH 7.2) and 0.005% Tween), and then further diluted to 18 ppm in a microtiter plate. Several dilutions were made in the microtiter plates, down to 0.27 ppm. 10 .mu.L of each of these samples were added to the three swatch plates, and 170 .mu.L of HEPES buffer (25 mM HEPES, pH 8.0 with 2 mM CaCl.sub.2) and 0.005% Tween-80) was added to each well for a final volume of 180 .mu.L. The final enzyme concentrations ranged from 1 ppm down to 0.015 ppm. The plates were incubated at 25.degree. C. with agitation at 1150 rpm for 15 minutes. Enzyme performance was judged by the amount of color released into the wash liquor. Color release was quantified spectrophotometrically at 488 nm by the transfer of 140 .mu.L of the final wash solution to fresh medium-binding microtiter plates, and triplicate reads were blank-subtracted and averaged.
[0680] The results of the microswatch cleaning assays performed at 0.015 ppm enzyme are shown in FIG. 28. Variants CspAmy2-v179, v186, and v191 all demonstrated superior cleaning performance compared to CspAmy2-v5, and all CspAmy2 combinatorial variants were far superior to STAINZYME.RTM.. The results of the cleaning assays performed at all enzyme concentrations are shown in Table 8. All CspAmy2 demonstrated superior cleaning performance at lower concentrations compared to STAINZYME.RTM..
TABLE-US-00036 TABLE 8 Results of the cleaning assays performed using CspAmy2 variants Enzyme concentration (ppm) Enzyme 0.015 0.032 0.045 0.063 0.090 0.125 0.25 0.5 1.0 CspAmy2-v179 0.25 0.31 0.32 0.35 0.35 0.37 0.37 0.36 0.38 CspAmy2-v186 0.28 0.33 0.34 0.35 0.38 0.38 0.39 0.39 0.39 CspAmy2-v191 0.22 0.30 0.31 0.35 0.37 0.38 0.39 0.39 0.40 CspAmy2-v5 0.19 0.26 0.30 0.34 0.35 0.37 0.38 0.39 0.38 STAINZYME .RTM. 0.07 0.12 0.14 0.19 0.19 0.25 0.30 0.34 0.38 ACE-QK 0.30 0.33 0.36 0.34 0.35 0.36 0.36 0.38 0.38
[0681] Thermal stability assays were performed essentially as described. Stocks of CspAmy2 variants, STAINZYME.RTM., and ACE-QK at 0.5 mg/mL were diluted to 5 ppm, 10 ppm, or 1 ppm, respectively, in dilution buffer (50 mM MOPS (pH 7.2) and 0.005% Tween) to account for their relative specific activities on the soluble substrate. 50 .mu.L of each enzyme were added to each of 12 wells of PCR tubes and sealed. The "unstressed" samples were incubated at room temperature throughout the duration of the experiment. The other samples were incubated in a thermocycler in a gradient from 77.degree. C. to 97.degree. C. for 15 minutes. Samples were transferred to microtiter plates in triplicate, and alpha-amylase activity was measured on all unstressed and stressed samples using the Ceralpha reagent (Megazyme, Inc.). Residual activity was calculated by dividing the activity of each amylase after the thermal stress by the activity of that unstressed amylase.
[0682] The results of the thermostability assay are shown in FIG. 29. Variants CspAmy2-v186 and v191 both demonstrated superior thermal stability compared to CspAmy2-v5. All the CspAmy2 variants demonstrated superior thermal stability compared to STAINZYME.RTM. and ACE-QK.
[0683] The in-detergent storage stability of the CspAmy2 variants was tested in a several commercial detergents, i.e., TIDE.RTM. regular HDL and TIDE.RTM. PODS.TM. (Procter & Gamble) for the USA market, ARIEL.TM. HDL (Procter & Gamble) and OMO Color HDL (Unilever) for the European market, and OMO.TM. (Unilever) and LIBY.TM. HDL (Liby) for the Chinese market. All detergents were heat inactivated at 90.degree. C. for 4 hours to eliminate existing enzyme activities. Enzyme activity in the heat inactivated detergents was measured using the Suc-AAPF-pNA and Ceralpha assays for measuring protease and amylase activity, respectively.
[0684] To prepare the stability samples, 2% w/w protease (PURAFECT.RTM. Prime HA, DuPont Industrial Biosciences) and 0.5% w/w amylase were added to each detergent sample and mixed. Samples were stored in a CO2 incubator (Sanyo) at 37.degree. C. for 28 days. Aliquots were taken from each reaction sample at various time points, diluted in 50 mM MOPS (pH 7.15) buffer with 1% BSA added, and alpha-amylase activity was measured using the Ceralpha substrate (Megazyme, Inc). The activity for each sample was determined using an Arena 20XT Photometric Analyzer (Thermo Scientific) using a calibrated standard. The remaining activity after incubation for 28 days was reported as a percent of the total activity determined at time zero.
[0685] The amount of residual activity of the CspAmy2 variants compared to STAINZYME.RTM. and ACE-QK are shown in FIGS. 30-35. CspAmy2-v179 was particularly stable compared to other tested variants and the controls.
[0686] The amino acid sequence of mature CspAmy2-v179 is shown, below, as SEQ ID NO: 28:
TABLE-US-00037 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFDGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG DYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNTESVSVWVQQ
[0687] The amino acid sequence of mature CspAmy2-v180 is shown, below, as SEQ ID NO: 29:
TABLE-US-00038 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFDGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG DYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVEGQSVSVWVQQ
[0688] The amino acid sequence of mature CspAmy2-v181 is shown, below, as SEQ ID NO: 30:
TABLE-US-00039 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGEYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFDGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG DYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVETRSVSVWVQQ
[0689] The amino acid sequence of mature CspAmy2-v186 is shown, below, as SEQ ID NO: 31:
TABLE-US-00040 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGINAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVHTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQEISGEYMIQAWTGFNFPGRGTTY SNWKWQWFHFDGTDWDQSRSRSRIFKFDGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG YYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNKESVSVWVQQ
[0690] The amino acid sequence of mature CspAmy2-v191 is shown, below, as SEQ ID NO: 32:
TABLE-US-00041 AATNGTMMQYFEWYVPNDGQQWNRLRTDAPYLSSVGITAVWTPPAYKGTS QADVGYGPYDLYDLGEFNQKGTVRTKYGTKGELKSAVNTLHSNGIQVYGD VVMNHKAGADYTENVTAVEVNPSNRYQETSGHYNIQAWTGFNFPGRGTTY SNFKWQWFHFDGTDWDQSRSLSRIFKFDGKAWDWPVSSENGNYDYLMYAD YDYDHPDVVNEMKKWGVWYANEVGLDGYRLDAVKHIKFSFLKDWVDNARA ATGKEMFTVGEYWQNDLGALNNYLAKVNYNQSLFDAPLHYNFYAASTGGG DYDMRNILNNTLVASNPTKAVTLVENHDTQPGQSLESTVQPWFKPLAYAF ILTRSGGYPSVFYGDMYGTKGTTTREIPALKSKIEPLLKARKDYAYGTQR DYIDNPDVIGWTREGDSTKAKSGLATVITDGPGGSKRMYVGTSNAGEIWY DLTGNRTDKITIGSDGYATFPVNTESVSVWVQQ
Example 18
Combinatorial Variants of PcuAmy1
[0691] To determine whether equivalent combinatorial mutations resulted in similar performance gains in a different .alpha.-amylase molecule, equivalent mutations were made in an .alpha.-amylase from Paenibacillus curdlanolyticus (i.e., PcuAmy1). The amino acid sequence of the mature of PcuAmy1 polypeptide is shown, below (SEQ ID NO: 3):
TABLE-US-00042 ADNGTIMQYFEWYLPNDGAHWNRLNNDAQNLKNVGITAVWIPPAYKGGSS ADVGYGVYDTYDLGEFNQKGTVRTKYGTKSELISAVNNLHAKGIAVYGDV VLNHRMNADATELVDAVEVDPNNRNVETTSTYQIQAWTQYDFPGRGNTYS SFKWRWYHFDGVDWDQSRGLNRIYKLRGDGKDWDWEVDSEYGNYDYLMGA DLDFNHPDVVNETKTWGKWFVNTVNLDGVRLDAVKHIKFDFMRDWVNNVR STTGKNLFAVGEYWHYDVNKLNSYITKTNGTMSLFDVPLHFRFYDASNGG GGYDMRNLLNNTLMSSNPMKAVTFVENHDTQPTQALQSTVQSWFKPLAYA TILTREQGYPCVFYGDYYGTSDGKISSYKPIMDKLLNARKVYAYGTQRDY FDHPDIVGWTREGDAAHAGSGLATLITDGPGGSKWMYVGTSKAGQVWTDK TGNRSGTVTIDANGWGNFWVNGGSVSVWAK
[0692] Mutations were made at positions N125, F152, R177, G178, E186, G472, and G473 (using SEQ ID NO: 3 for numbering), corresponding to mutations at positions N126, F153, R178, G179, E187P, G476, and G477, respectively in CspAmy2 (SEQ ID NO: 1). A further mutation was introduced at position N205. Consistent with previous nomenclature "del (R177, G178)" refers to deletions, in this case at positions R177 and G178. In addition to the aforementioned mutations, the PcuAmy1 variants further included mutations at position T333, A335, and Q337E. These mutations, particularly at T333, impart protease resistance to PcuAmy1 but do not affect performance (see Example 19). The variants are shown in Table 9.
TABLE-US-00043 TABLE 9 Combinatorial variants of PcuAmy1 amylase Variant Mutations PcuAmy1-v1A del (R177, G178) + N125Y + E186P + T333G + A335S + Q337E + G472K PcuAmy1-v6 del (R177, G178) + N125Y + F152W + E186P + T333G + A335S + Q337E + G472K PcuAmy1-v8 del (R177, G178) + N125Y + F152W + E186P + T333G + A335S + Q337E + G472R + G473R PcuAmy1-v16 del (R177, G178) + N125Y + F152W + E186P + N205D + T333G + A335S + Q337E + G472K
[0693] The codon-optimized nucleotide sequence of the PcuAmy1 gene is set forth as SEQ ID NO: 33:
TABLE-US-00044 GCCGACAACGGCACAATCATGCAGTATTTCGAGTGGTACCTGCCGAACGA CGGAGCGCACTGGAACAGACTTAATAACGACGCACAAAACCTGAAAAATG TGGGCATCACGGCAGTGTGGATTCCTCCGGCATACAAGGGCGGCAGCTCA GCAGATGTTGGCTACGGAGTTTACGATACATACGACCTGGGCGAGTTCAA TCAGAAAGGCACGGTCAGAACAAAGTACGGAACGAAGAGCGAACTGATTT CAGCGGTCAACAATCTTCACGCAAAGGGCATTGCGGTTTACGGCGACGTG GTCCTGAACCATAGAATGAATGCGGATGCAACGGAGCTTGTGGATGCGGT TGAGGTGGATCCGAACAACAGAAACGTCGAGACGACAAGCACGTATCAGA TCCAGGCATGGACGCAATACGATTTCCCGGGCAGAGGCAACACGTACAGC AGCTTTAAATGGAGATGGTATCACTTCGACGGCGTCGACTGGGACCAGAG CAGAGGCCTGAACAGAATCTATAAGCTGAGAGGCGATGGCAAGGATTGGG ACTGGGAGGTCGACAGCGAGTACGGCAACTACGATTACCTGATGGGAGCG GACCTGGACTTCAACCACCCGGATGTGGTTAACGAAACAAAGACATGGGG CAAATGGTTTGTGAACACGGTGAACCTGGATGGCGTCAGACTGGACGCGG TTAAGCACATCAAGTTCGACTTCATGAGAGACTGGGTGAACAACGTGAGA AGCACGACGGGCAAGAACCTTTTCGCAGTTGGCGAGTATTGGCACTACGA CGTGAACAAACTGAACAGCTACATCACGAAGACGAATGGCACGATGAGCC TGTTCGACGTGCCGCTGCACTTTAGATTTTATGATGCAAGCAACGGCGGA GGCGGCTACGACATGAGAAACCTGCTGAATAACACGCTGATGAGCAGCAA CCCGATGAAGGCGGTTACATTCGTTGAGAACCATGACACACAACCGACGC AGGCCCTGCAATCAACGGTCCAAAGCTGGTTTAAGCCGCTTGCGTATGCT ACAATCCTGACGAGAGAGCAAGGCTACCCGTGCGTTTTCTACGGCGACTA TTATGGAACAAGCGACGGCAAAATTAGCAGCTACAAGCCGATCATGGATA AGCTTCTTAACGCGAGAAAGGTGTACGCCTACGGCACGCAGAGAGATTAC TTCGATCATCCGGACATCGTTGGCTGGACAAGAGAAGGCGATGCAGCACA TGCTGGCTCAGGACTGGCAACGCTTATCACAGATGGCCCTGGCGGAAGCA AGTGGATGTATGTTGGAACGTCAAAGGCAGGCCAGGTCTGGACGGATAAA ACAGGAAACAGAAGCGGAACGGTGACGATTGATGCCAATGGCTGGGGAAA CTTTTGGGTTAATGGCGGATCAGTTAGCGTTTGGGCAAAATAA
[0694] The amino acid sequence of the mature of PcuAmy1-v1A polypeptide is shown below as SEQ ID NO: 34:
TABLE-US-00045 ADNGTIMQYFEWYLPNDGAHWNRLNNDAQNLKNVGITAVWIPPAYKGGSS ADVGYGVYDTYDLGEFNQKGTVRTKYGTKSELISAVNNLHAKGIAVYGDV VLNHRMNADATELVDAVEVDPNNRYVETTSTYQIQAWTQYDFPGRGNTYS SFKWRWYHFDGVDWDQSRGLNRIYKLDGKDWDWPVDSEYGNYDYLMGADL DFNHPDVVNETKTWGKWFVNTVNLDGVRLDAVKHIKFDFMRDWVNNVRST TGKNLFAVGEYWHYDVNKLNSYITKTNGTMSLFDVPLHFRFYDASNGGGG YDMRNLLNNTLMSSNPMKAVTFVENHDTQPGQSLESTVQSWFKPLAYATI LTREQGYPCVFYGDYYGTSDGKISSYKPIMDKLLNARKVYAYGTQRDYFD HPDIVGWTREGDAAHAGSGLATLITDGPGGSKWMYVGTSKAGQVWTDKTG NRSGTVTIDANGWGNFWVNKGSVSVWAK
[0695] The amino acid sequence of the mature of PcuAmy1-v6 polypeptide is shown below as SEQ ID NO: 35:
TABLE-US-00046 ADNGTIMQYFEWYLPNDGAHWNRLNNDAQNLKNVGITAVWIPPAYKGGSS ADVGYGVYDTYDLGEFNQKGTVRTKYGTKSELISAVNNLHAKGIAVYGDV VLNHRMNADATELVDAVEVDPNNRYVETTSTYQIQAWTQYDFPGRGNTYS SWKWRWYHFDGVDWDQSRGLNRIYKLDGKDWDWPVDSEYGNYDYLMGADL DFNHPDVVNETKTWGKWFVNTVNLDGVRLDAVKHIKFDFMRDWVNNVRST TGKNLFAVGEYWHYDVNKLNSYITKTNGTMSLFDVPLHFRFYDASNGGGG YDMRNLLNNTLMSSNPMKAVTFVENHDTQPGQSLESTVQSWFKPLAYATI LTREQGYPCVFYGDYYGTSDGKISSYKPIMDKLLNARKVYAYGTQRDYFD HPDIVGWTREGDAAHAGSGLATLITDGPGGSKWMYVGTSKAGQVWTDKTG NRSGTVTIDANGWGNFWVNKGSVSVWAK
[0696] The amino acid sequence of the mature of PcuAmy1-v8 polypeptide is shown below as SEQ ID NO: 36:
TABLE-US-00047 ADNGTIMQYFEWYLPNDGAHWNRLNNDAQNLKNVGITAVWIPPAYKGGSS ADVGYGVYDTYDLGEFNQKGTVRTKYGTKSELISAVNNLHAKGIAVYGDV VLNHRMNADATELVDAVEVDPNNRYVETTSTYQIQAWTQYDFPGRGNTYS SWKWRWYHFDGVDWDQSRGLNRIYKLDGKDWDWPVDSEYGNYDYLMGADL DFNHPDVVNETKTWGKWFVNTVNLDGVRLDAVKHIKFDFMRDWVNNVRST TGKNLFAVGEYWHYDVNKLNSYITKTNGTMSLFDVPLHFRFYDASNGGGG YDMRNLLNNTLMSSNPMKAVTFVENHDTQPGQSLESTVQSWFKPLAYATI LTREQGYPCVFYGDYYGTSDGKISSYKPIMDKLLNARKVYAYGTQRDYFD HPDIVGWTREGDAAHAGSGLATLITDGPGGSKWMYVGTSKAGQVWTDKTG NRSGTVTIDANGWGNFWVNRRSVSVWAK
[0697] The amino acid sequence of the mature of PcuAmy1-v16 polypeptide is shown below as SEQ ID NO: 37:
TABLE-US-00048 ADNGTIMQYFEWYLPNDGAHWNRLNNDAQNLKNVGITAVWIPPAYKGGSS ADVGYGVYDTYDLGEFNQKGTVRTKYGTKSELISAVNNLHAKGIAVYGDV VLNHRMNADATELVDAVEVDPNNRYVETTSTYQIQAWTQYDFPGRGNTYS SWKWRWYHFDGVDWDQSRGLNRIYKLDGKDWDWPVDSEYGNYDYLMGADL DFDHPDVVNETKTWGKWFVNTVNLDGVRLDAVKHIKFDFMRDWVNNVRST TGKNLFAVGEYWHYDVNKLNSYITKTNGTMSLFDVPLHFRFYDASNGGGG YDMRNLLNNTLMSSNPMKAVTFVENHDTQPGQSLESTVQSWFKPLAYATI LTREQGYPCVFYGDYYGTSDGKISSYKPIMDKLLNARKVYAYGTQRDYFD HPDIVGWTREGDAAHAGSGLATLITDGPGGSKWMYVGTSKAGQVWTDKTG NRSGTVTIDANGWGNFWVNKGSVSVWAK
[0698] The cleaning performance of PcuAmy1-v1, PcuAmy1-v6, and PcuAmy1-v16, compared to STAINZYME.RTM. and ACE-QK is shown in FIG. 36. The microswatch assay was performed as described in Example 17. PcuAmy1-v6 and PcuAmy1-v16 outperformed PcuAmy1-v1 and STAINZYME.RTM. at low doses (e.g., 0.1 ppm enzyme or less).
[0699] The thermal stability of PcuAmy1-v1, PcuAmy1-v6, and PcuAmy1-v16, compared to STAINZYME.RTM. is shown in FIG. 37. The assays were performed as described in Example 17. PcuAmy1-v16 were more thermostable than the other tested molecules using the same detergents and enzyme doses.
Example 19
Combinatorial Variants in BASE
[0700] As in Example 18, to determine whether equivalent combinatorial mutations resulted in similar performance gains in a different .alpha.-amylase molecule, equivalent mutations were made in an .alpha.-amylase derived from Bacillus sp. TS-23. The amino acid sequence of BASE, a C-terminal-truncated version of the Bacillus sp. TS-23 .alpha.-amylase (see, e.g., US20120045817 and WO2010/115028), is shown below as SEQ ID NO: 5:
TABLE-US-00049 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRNQETSGTYQIQAWTKFDFPGRGN TYSSFKWRWYHFDGTDWDESRKLNRIYKFRSTGKAWDWEVDTENGNYDYL MFADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLT YVRNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTAS KSSGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPL AYAFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQ RDYIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVF YDLTGNRSDTVTINADGWGEFKVNGGSVSIWVAK
[0701] The codon-modified nucleic acid sequence encoding the mature form of BASE (AmyTS23t), is set forth as SEQ ID NO: 38:
TABLE-US-00050 TCTGCAGCT TCAGCAAAC ACCGCGCCG ATTAACGAA ACCATGATG CAGTATTTC GAATGGGAT CTGCCGAAC GATGGCACC CTGTGGACC AAAGTGAAA AACGAAGCG GCGAACCTG AGCAGCCTG GGCATTACC GCGCTGTGG CTGCCGCCG GCATATAAA GGCACCAGC CAGAGCGAT GTGGGCTAT GGCGTGTAT GATCTGTAC GATCTGGGC GAATTTAAC CAGAAAGGC ACCATTCGT ACCAAATAT GGCACCAAA ACCCAGTAT ATTCAGGCG ATCCAGGCG GCGAAAGCG GCGGGTATG CAGGTGTAT GCGGATGTG GTGTTTAAC CATAAAGCG GGTGCGGAT GGCACCGAA TTTGTGGAT GCGGTGGAA GTGGATCCG AGCAACCGT AACCAGGAA ACCAGCGGC ACCTATCAG ATTCAGGCG TGGACCAAA TTTGATTTT CCCGGCCGT GGCAACACC TATAGCAGC TTTAAATGG CGCTGGTAT CATTTTGAT GGCACCGAT TGGGATGAA AGCCGTAAA CTGAACCGC ATCTATAAA TTTCGTAGC ACCGGCAAA GCGTGGGAT TGGGAAGTG GATACCGAA AACGGCAAC TATGATTAC CTGATGTTC GCAGACCTG GATATGGAT CATCCGGAA GTGGTGACC GAACTGAAA AACTGGGGC ACCTGGTAT GTGAACACC ACCAACATT GATGGCTTT CGTCTGGAT GCGGTGAAA CACATCAAA TACAGCTTT TTTCCGGAT TGGCTGACC TATGTGCGT AACCAGACC GGCAAAAAC CTGTTTGCG GTGGGCGAA TTTTGGAGC TATGATGTG AACAAACTG CACAACTAC ATCACCAAA ACCAACGGC AGCATGAGC CTGTTTGAT GCGCCGCTG CATAACAAC TTTTATACC GCGAGCAAA AGCAGCGGC TATTTTGAT ATGCGTTAT CTGCTGAAC AACACCCTG ATGAAAGAT CAGCCGAGC CTGGCCGTG ACCCTGGTG GATAACCAT GATACCCAG CCGGGCCAG AGCCTGCAA AGCTGGGTG GAACCGTGG TTTAAACCG CTGGCCTAC GCGTTTATT CTGACCCGT CAAGAGGGC TATCCGTGC GTTTTTTAT GGCGATTAT TACGGCATC CCGAAATAT AACATTCCG GGCCTGAAA AGCAAAATT GATCCGCTG CTGATTGCG CGTCGTGAT TATGCGTAT GGCACCCAG CGTGATTAT ATTGATCAC CAGGATATT ATTGGCTGG ACCCGTGAA GGCATTGAT ACCAAACCG AACAGCGGC CTGGCCGCG CTGATTACC GATGGCCCG GGTGGCAGC AAATGGATG TATGTGGGC AAAAAACAT GCGGGCAAA GTGTTTTAT GATCTGACC GGCAACCGT AGCGATACC GTGACCATT AACGCGGAT GGCTGGGGT GAGTTTAAA GTGAACGGC GGCAGCGTG AGCATTTGG GTGGCGAAA TAAGTTAAC AGA
[0702] Mutations were made at positions N128, T134, F155, T182, R180, S181, E189, and G475 were made in BASE (using SEQ ID NO: 5 for numbering), which corresponds to mutations at positions N126, E132, F153, R178, G179, E187P, and G476, respectively in CspAmy2 (SEQ ID NO: 1). The variants are shown in Table 10.
TABLE-US-00051 TABLE 10 Combinatorial variants of BASE amylase Variant Mutations BASE-V28 del (R180, G181) + N128Y + E189P + G475R BASE-V29 del (R180, G181) + F155W + E189P + G475R BASE-V30 del (R180, G181) + T134E + T182H + E189P + G475R BASE-V31 del (R180, G181) + N128Y + T134E + T182H + E189P + G475R BASE-V32 del (R180, G181) + N128Y + F155W + E189P + G475R BASE-V33 del (R180, G181) + T134E + F155W + T182H + E189P + G475R BASE-V34 del (R180, G181) + N128Y + T134E + F155W + T182H + E189P + G475R BASE-V35 del (R180, G181) + N128Y + T134H + F155W + T182D + E189P + G475R BASE-V36 del (R180, G181) + N128Y + T134E + F155W + T182G + E189P + G457R ACE-QK del (R180, G181) + S243Q + G475K
[0703] The amino acid sequence of BASE-V28 is shown below as SEQ ID NO: 39:
TABLE-US-00052 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRYQETSGTYQIQAWTKFDFPGRGN TYSSFKWRWYHFDGTDWDESRKLNRIYKFTGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0704] The amino acid sequence of BASE-V29 is shown below as SEQ ID NO: 40:
TABLE-US-00053 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRNQETSGTYQIQAWTKFDFPGRGN TYSSWKWRWYHFDGTDWDESRKLNRIYKFTGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0705] The amino acid sequence of BASE-V30 is shown below as SEQ ID NO: 41:
TABLE-US-00054 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRNQETSGEYQIQAWTKFDFPGRGN TYSSFKWRWYHFDGTDWDESRKLNRIYKFHGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0706] The amino acid sequence of BASE-V31 is shown below as SEQ ID NO: 42:
TABLE-US-00055 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRYQETSGEYQIQAWTKFDFPGRGN TYSSFKWRWYHFDGTDWDESRKLNRIYKFHGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0707] The amino acid sequence of BASE-V32 is shown below as SEQ ID NO: 43:
TABLE-US-00056 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRYQETSGTYQIQAWTKFDFPGRGN TYSSWKWRWYHFDGTDWDESRKLNRIYKFTGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0708] The amino acid sequence of BASE-V33 is shown below as SEQ ID NO: 44:
TABLE-US-00057 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRNQETSGEYQIQAWTKFDFPGRGN TYSSWKWRWYHFDGTDWDESRKLNRIYKFHGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0709] The amino acid sequence of BASE-V34 is shown below as SEQ ID NO: 45:
TABLE-US-00058 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRYQETSGEYQIQAWTKFDFPGRGN TYSSWKWRWYHFDGTDWDESRKLNRIYKFHGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0710] The amino acid sequence of BASE-V35 is shown below as SEQ ID NO: 46:
TABLE-US-00059 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRYQETSGHYQIQAWTKFDFPGRGN TYSSWKWRWYHFDGTDWDESRKLNRIYKFDGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0711] The amino acid sequence of BASE-V36 is shown below as SEQ ID NO: 47:
TABLE-US-00060 NTAPINETMMQYFEWDLPNDGTLWTKVKNEAANLSSLGITALWLPPAYKG TSQSDVGYGVYDLYDLGEFNQKGTIRTKYGTKTQYIQAIQAAKAAGMQVY ADVVFNHKAGADGTEFVDAVEVDPSNRYQETSGEYQIQAWTKFDFPGRGN TYSSWKWRWYHFDGTDWDESRKLNRIYKFGGKAWDWPVDTENGNYDYLMF ADLDMDHPEVVTELKNWGTWYVNTTNIDGFRLDAVKHIKYSFFPDWLTYV RNQTGKNLFAVGEFWSYDVNKLHNYITKTNGSMSLFDAPLHNNFYTASKS SGYFDMRYLLNNTLMKDQPSLAVTLVDNHDTQPGQSLQSWVEPWFKPLAY AFILTRQEGYPCVFYGDYYGIPKYNIPGLKSKIDPLLIARRDYAYGTQRD YIDHQDIIGWTREGIDTKPNSGLAALITDGPGGSKWMYVGKKHAGKVFYD LTGNRSDTVTINADGWGEFKVNRGSVSIWVAK
[0712] The thermal stability of BASE-V28, V29, V30, V31, V32, V33, V34, and V35, compared to ACE-QK (e.g., US20120045817 and WO2010/115028), is shown in FIG. 38. All the BASE variants were more stable than ACE-QK, although BASE-V28 was only marginally more stable.
Example 19
Interactions Between Residues in RG-Deletion Molecules
[0713] A structural interaction between residues 132 and 180 (referring to SEQ ID NO: 1 for numbering) explains the increased stability of some of the variants. Details of the crystal structure of CspAmy2-v1 are shown in FIGS. 39-42. As shown in FIG. 39, the naturally occurring glutamate side chain at position 132 is positioned towards the side chain of the naturally-occurring threonine at position 180. The distance of 5.4 Angstroms, however, is too great for the formation of any stabilizing interaction. As shown in FIG. 40, a T180H variant (e.g., CspAmy2-vC16C) has the histidine imidazole NH group in proximity to the E132 glutamate carboxylate. The distance of 3.2 Angstroms allows the formation of a stabilizing hydrogen bond. At pH of roughly 4.5 to 7.0, a favorable charge interaction (i.e., salt bridge) is also likely between these residues. Referring to, e.g., Examples 6 and 7 and FIGS. 8-10, the observations that CspAmy2-C16E is more stable than CspAmy2-C16C, CspAmy2-C16F is more stable than CspAmy2-C16D, CspAmy2-C16I is more stable than CspAmy2-C16G, and CspAmy2-C16J is more stable than CspAmy2-C16H, which pairs of variants differ only by the presence or absence of the mutation T180H, supports this hypothesis. Similarly, referring to, e.g., Examples 17 and FIG. 29, the observation that CspAmy2-v191 is more stable than CspAmy2-v179, which pair of variants differ only by the presence or absence of the mutation E132H, supports this hypothesis.
[0714] As shown in FIG. 41, an aspartic acid at position 180 may also be capable of hydrogen bonding with the glutamate at position 132, although hydrogen bonding may be overwhelmed by unfavorable like charge interactions. However, the presence of histidine at position 132, in combination with an aspartate at position 180, restores the possibility for a favorable interaction created by a T180D mutation (FIG. 42). The stabilizing effect of the E132H mutation in CspAmy2-v191 and CspAmy2-C25A, B, and F, which all have a T180D mutation, supports this hypothesis (e.g., Example 16 and FIG. 27).
[0715] In BASE, position E132 corresponds to position T134 and position T180 corresponds to position T182. The observations that BASE-V31 is more stable than BASE-V28 and BASE-V33 is more stable than BASE-V29 further supports this hypothesis in the context of a different .alpha.-amylase. In both cases, the mutations T134E and T180H appear to work together to enable the formation of a stabilizing interaction, likely a salt bridge. Similarly, V34 is more stable than V36, because of the stabilizing interaction between the glutamate and histidine in V34, which does not occur between the glutamate and glycine in V36.
[0716] Although the position 132-180 interaction was demonstrated using an "RG" deletion, it can fully be expected to work in the context of an adjacent "DG" or TG'' deletion. It will be appreciated that the conserved amino acid sequence motif X.sub.1G/S.sub.1X.sub.2G.sub.2 (SEQ ID NO: 48), as exemplified by RGTG (SEQ ID NO: 49) in CspAmy2 .alpha.-amylase (SEQ ID NO: 1), is adjacent to the calcium-binding loop in .alpha.-amylases. X1 is typically arginine. G/Si is most often glycine but is serine in the case of BASE. X2 varies but is commonly aspartate or threonine. G2 is highly conserved. Deleting the TG in CspAmy2 .alpha.-amylase, instead of RG, would mean that the remaining residues would be RG rather than TG. Although the arginine would be derived from position 178 as opposed to the threonine, which is derived from position 180, the three-dimensional structure of the resulting variant is indistinguishable from one having an RG deletion plus an R to G substitution, and stabilizing the resulting molecule is simply a matter of selecting a suitable residue at position 132 to form a stabilizing interaction with whatever residue is remaining at the equivalent position in the X.sub.1G/S.sub.1X.sub.2G.sub.2 motif, whether it originally corresponded to position 180 or position 178 in the patent molecule (using SEQ ID NO: 1 for numbering). For convenience, this residue may be referred to as the remaining non-G residue in the aforementioned motif.
[0717] Therefore, in general, if position 132 is negatively charged (i.e., D or E), then the remaining non-G residue should be positively charged (i.e., H, R, or K). If position 132 is positively charged (i.e., H, R, or K), then the remaining non-G residue should be negatively charged (i.e., D or E).
Example 19
Proteolytic Cleavage of PcuAmy1 and Variants
[0718] Incubation of wild-type PcuAmy1 amylase, or the PcuAmy-v1 variant, with subtilisin proteases leads to cleavage of the proteins, as observed when the reaction products are subjected to SDS/PAGE electrophoresis. FIG. 43 is an image of an SDS/PAGE gel showing the cleavage of 20 .mu.g of PcuAmy1-v1 in the presence of increasing amounts of GG36 protease, (from 0 to 40 .mu.g as indicated above the gel). The letters on the right side of the gel indicate (A) intact full-length PcuAmy1-v1, (B) a first cleavage product of PcuAmy1-v1, (C) GG36 protease, (D) a contaminant in the GG36 protein preparation, and (E) a second cleavage product of PcuAmy1-v1. The main degradation products observed after incubation of PcuAmy-v1 amylase with a subtilisin protease have a molecular weight of about 38 and 16 kDa (B and E, respectively). The amount of proteolytic degradation is dependent on the concentration of protease used. This makes PcuAmy1 amylase suboptimal for inclusion in enzyme detergent formulations that contain commonly-used subtilisin proteases.
[0719] A sample of PcuAmy-v1 protein was incubated with GG36 protease (Bacillus lentis subtilisin) and the reaction products were analyzed by mass spectroscopy. The results were consistent with hydrolysis occurring between residues Q334 and L336 (not shown).
[0720] To determine whether protease-stable variants of PcuAmy could be engineered, PcuAmy1-v3 and further variants 3A to 3L were constructed and tested. PcuAmy1-v3 is a variant of PcuAmy1 with the mutations E186P, G472K and lacking R177 and G178 (using SEQ ID NO: 3 for numbering). The substitution E186P and the deletions at R177 and G178 increase the detergent stability of PcuAmy1. The substitution G472K improves cleaning performance. None of these mutations has any effect on protease sensitivity (data not shown). Therefore, including these mutations in variants made to explore the effect of other mutations on protease stability does not interfere with the results.
[0721] The mature form of PcuAmy1-v3 is shown, below, as (SEQ ID NO: 50):
TABLE-US-00061 ADNGTIMQYFEWYLPNDGAHWNRLNNDAQNLKNVGITAVWIPPAYKGGSS ADVGYGVYDTYDLGEFNQKGTVRTKYGTKSELISAVNNLHAKGIAVYGDV VLNHRMNADATELVDAVEVDPNNRNVETTSTYQIQAWTQYDFPGRGNTYS SFKWRWYHFDGVDWDQSRGLNRIYKLDGKDWDWPVDSEYGNYDYLMGADL DFNHPDVVNETKTWGKWFVNTVNLDGVRLDAVKHIKFDFMRDWVNNVRST TGKNLFAVGEYWHYDVNKLNSYITKTNGTMSLFDVPLHFRFYDASNGGGG YDMRNLLNNTLMSSNPMKAVTFVENHDTQPTQALQSTVQSWFKPLAYATI LTREQGYPCVFYGDYYGTSDGKISSYKPIMDKLLNARKVYAYGTQRDYFD HPDIVGWTREGDAAHAGSGLATLITDGPGGSKWMYVGTSKAGQVWTDKTG NRSGTVTIDANGWGNFWVNKGSVSVWAK
[0722] To identify the reason for PcuAmy1 protease sensitivity, the amino acid sequence of PcuAmy1 was compared to that of other CAZy Family GH-13 amylases which show protease-resistance, such as PURASTAR.RTM. ST (B. licheniformis amylase or AmyL), SPEZYME.RTM. XTRA (Geobacillus stearothermophilus amylase or AmyS), ACE-QK (WO2010/115021), and STAINZYME.RTM. (Novozymes). Based on observed differences in sequence, PcuAmy1 variants PcuAmy1-v3A to PcuAmy1-v3L (i.e., 3A-3L) were designed and tested for protease resistance. The mutations present in each variant are listed in Table 11. They were introduced into PcuAmy1-v3 (SEQ ID NO: 49), using standard methods, many of which are described above.
TABLE-US-00062 TABLE 11 List of mutations introduced in PcuAmy1-v3 resulting in variants 3A to 3L Position wt 3A 3B 3C 3D 3E 3F 3G 3H 3I 3J 3K 3L 319 M T 333 T G G G G G G 335 A S S S S S S 337 Q E E E E E 339 T W 341 Q E 342 S P P P P T 351 T F F W
[0723] The 12 PcuAmy1 variants were expressed in B. subtilis as described, above. 40 filtered supernatant from each PcuAmy1 variant culture broth was incubated with 100 .mu.g GG36 protease for 6 hours at room temperature, and subsequently analyzed for remaining amylase activity using the Megazyme Ceralpha substrate assay (Megazyme International Ireland, Co. Wicklow, Ireland). Residual activity after protease incubation was compared to the amylase activity of each sample after incubation with buffer alone. The results are shown in FIG. 44. A subset of the samples was also analyzed by SDS-PAGE (FIG. 45), with the protein standard, SEEBLUE.RTM. Plus2 (Invitrogen). The commercially available amylases were included for comparison.
[0724] PcuAmy1 variants 3A, 3B, 3C, 3D, and 3L maintained >70% of their enzymatic activity after incubation with GG36 protease. PcuAmy1 variants 3J and 3K maintained >65% of their enzymatic activity after incubation with GG36 protease. PcuAmy1 variants 3E, 3F, 3G, 3H, and 3I did not show an appreciable increase of stability compared to the wild-type enzyme. Samples of the 3B, 3C, 3D and 3L incubations were analyzed by SDS/PAGE and showed significant reduction in degradation products when incubated with GG36 protease, confirming that the increase in residual amylase activity was due to decreased proteolytic cleavage.
[0725] These results of the small-scale experiment indicate that the introduction of a mutation at position T333 significantly reduces the proteolytic cleavage of PcuAmy1, and that additional mutations at A335, Q337, and 5342 further reduce proteolytic cleavage. The T351W mutation but not the T351F mutation, also appears to reduce the proteolytic cleavage of PcuAmy1.
[0726] To better characterize the relative contributions of these mutations to protease resistance, the following additional variants were made:
[0727] PcuAmy1-v10: del (R177, G178)+E186P+T333G+Q337E+G472K
[0728] PcuAmy1-v11: del (R177, G178)+E186P+T333G+A335S+G472K
[0729] PcuAmy1-v12: del (R177, G178)+E186P+A335S+Q337E+G472K
[0730] PcuAmy1-v13: del (R177, G178)+E186P+T333G+A335S+Q337E+T351W+G472K
[0731] Variants PcuAmy1-v10, PcuAmy1-v11, and PcuAmy1-v12 include pair-wise combination of mutations at positions T333, A335, and Q337. PcuAmy1-v3-v13 includes mutations at all the aforementioned positions and includes the additional mutation T351W. These variants were compared in a large scale detergent stability assay to the following previously-described variants:
[0732] PcuAmy1-v3B: del (R177, G178)+E186P+T333G+A335S+Q337E+G472K
[0733] PcuAmy1-v3L: del (R177, G178)+E186P+T333G+A335S+Q337E+S342T+G472K
[0734] The commercial detergents Total Color (MIFA Ag Frenkendorf, Switzerland) and Omo (Unilever, London, UK) were heat inactivated at 90.degree. C. for 4 hours to eliminate existing enzyme activities. Enzyme activity in the heat inactivated detergents was measured using the Suc-AAPF-pNA and Ceralpha assays for measuring protease and amylase activity, respectively. To prepare the stability samples, 2% w/w protease (PURAFECT.RTM. Prime 4000L, Danisco US Inc.) and 0.5% w/w amylase were added to each detergent sample and mixed. Samples were stored in a CO2 incubator (Sanyo) at 37.degree. C. for 14 days. Aliquots were taken from each reaction sample at various time points, diluted in 50 mM MOPS, pH 7.15 buffer with 1% BSA added, and alpha-amylase activity was measured using the Ceralpha substrate (Megazyme, Inc). The activity for each sample was determined using a Arena 20XT Photometric Analyzer (Thermo Scientific) using a calibrated standard. The remaining activity at each time point was reported as a percent (%) of the total activity determined at time zero.
[0735] The results of the detergent stability assays performed in MIFA Total (MIFA Ag Frenkendorf, Switzerland) and Unilever OMO (Unilever, London, UK) are shown in FIGS. 46-50. FIGS. 46 and 48 show the residual activity of PcuAmy1 variants over time in MIFA Total and Unilever Omo, respectively, supplemented with FNA protease. FIGS. 47 and 49 summarize the data for the 3 and 14 day time points. PcuAmy1 variants that include the T333 mutation, i.e., 3B, 3L, v10, v13, and to a lesser degree, v11 were the most stable. Variants that did not include the T333 mutation, i.e., V1 and v12, were the least stable. As evidenced by v10 and v11, the presence of a mutation at Q337E further improves stability.
[0736] The cleaning performance of purified PcuAmy1-v3B and PcuAmy1-v3L was analyzed in a microswatch cleaning assay. CFT CS-28 rice starch on cotton swatches (Center for Testmaterials, BV, Vlaardingen, Netherlands) containing an indicator dye bound to the starch were punched to form discs measuring 5.5 mm in diameter. Two discs were placed in each well of three flat-bottom non-binding 96-well assay plates.
[0737] Both enzymes and two commercial amylase products: PURASTAR.RTM. ST (alpha-amylase from Bacillus licheniformis; DuPont Industrial Biosciences, Palo Alto, Calif., USA), and STAINZYME.RTM. (Novozymes, Copenhagen, Denmark) were diluted to 0.5 mg/mL in dilution buffer (50 mM MOPS, pH 7.2, 0.005% Tween), and then further diluted to 2 ppm in a microtiter plate. 200 .mu.L of these samples were transferred into the first row of each of three swatch plates. 100 .mu.L of HEPES buffer (25 mM HEPES, pH 8.0 with 2 mM CaCl.sub.2) and 0.005% Tween-80) was then added to each well of the next five rows of the swatch plates, and serial dilutions were made to result in final enzyme concentrations of 2, 1, 0.5, 0.25, and 0.125 ppm as well as a row of blank (buffer only) wells with 200 .mu.L in every well. Plates were incubated at 25.degree. C. with agitation at 1150 rpm for 15 minutes. The wash liquor was transferred to fresh microtiter plates and enzyme performance was judged by the amount of color released into the wash liquor. Color release was quantified spectrophotometrically at 488 nm, and triplicate reads were blank-subtracted and averaged.
[0738] The results are shown in FIG. 50. Both PcuAmy1-v3B and PcuAmy1-v3L demonstrated excellent cleaning performance.
[0739] Commercial detergent Persil Universal Gel Gold (Henkel, Dusseldorf, Germany) was heat inactivated at 90.degree. C. for 4 hours to eliminate existing enzyme activities. Following inactivation, enzyme activity in the heat inactivated detergents was measured using the Suc-AAPF-pNA substrate-based and Ceralpha assay (Megazyme, Wicklow, Ireland) to ensure that any protease and amylase activities, respectively, had been abolished. A 10% solution of detergent was then made in water.
[0740] 100 .mu.L of each of the enzyme stocks (at 0.5 mg/mL) were added to 400 .mu.L of the 10% detergent solutions. The enzymes tested were PURASTAR.RTM., STAINZYME.RTM., PcuAmy1-v3B and PcuAmy-v3L. 50 .mu.L of enzyme stock solution was added to PCR tubes and incubated at either 60, 70, 80, or 90.degree. C. for 15 minutes. Prior to incubation, 10 .mu.L was removed and incubated at room temperature throughout the duration of the experiment to serve as the "unstressed" samples. Following incubation, an additional 1:10 dilution of each sample was made in dilution buffer. Samples were then transferred to microtiter plates in triplicate, and alpha-amylase activity was measured on all unstressed and stressed samples using the Ceralpha assay. Residual activity was calculated by dividing the activity of each amylase after the thermal stress by the activity of the unstressed amylase.
[0741] The results are shown in FIGS. 51 and 52. PcuAmy1-v3B and PcuAmy-v3L demonstrated similar thermostability compared to STAINZYME.RTM., and significantly better stability than PURASTAR.RTM..
[0742] Although the foregoing compositions and methods have been described in some detail by way of illustration and examples for purposes of clarity of understanding, it will be apparent to those skilled in the art that certain changes and modifications may be made. Therefore, the description should not be construed as limiting the scope of the invention, which is delineated by the appended claims.
[0743] All publications, patents, and patent applications cited herein are hereby incorporated by reference in their entireties for all purposes and to the same extent as if each individual publication, patent, or patent application were specifically and individually indicated to be so incorporated by reference.
Sequence CWU 1
1
501485PRTArtificial SequenceSynthetic amino acid sequence of the
mature form of the CspAmy2 alpha-amylase polypeptide 1Ala Ala Thr
Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp
Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser
Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40
45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu
50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr
Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn
Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys Ala Gly
Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val Asn Pro
Ser Asn Arg Asn Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn Ile Gln
Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr Thr Tyr
Ser Asn Phe Lys Trp Gln Trp Phe His Phe145 150 155 160Asp Gly Thr
Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170 175Phe
Arg Gly Thr Gly Lys Ala Trp Asp Trp Glu Val Ser Ser Glu Asn 180 185
190Gly Asn Tyr Asp Tyr Leu Met Tyr Ala Asp Ile Asp Tyr Asp His Pro
195 200 205Asp Val Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu 210 215 220Val Gly Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe225 230 235 240Ser Phe Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys 245 250 255Glu Met Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu 260 265 270Asn Asn Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala 275 280 285Pro Leu His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr 290 295 300Asp
Met Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr305 310
315 320Lys Ala Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln
Ser 325 330 335Leu Glu Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala
Tyr Ala Phe 340 345 350Ile Leu Thr Arg Ser Gly Gly Tyr Pro Ser Val
Phe Tyr Gly Asp Met 355 360 365Tyr Gly Thr Lys Gly Thr Thr Thr Arg
Glu Ile Pro Ala Leu Lys Ser 370 375 380Lys Ile Glu Pro Leu Leu Lys
Ala Arg Lys Asp Tyr Ala Tyr Gly Thr385 390 395 400Gln Arg Asp Tyr
Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu 405 410 415Gly Asp
Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp 420 425
430Gly Pro Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
435 440 445Glu Ile Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile 450 455 460Gly Ser Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly
Gly Ser Val Ser465 470 475 480Val Trp Val Gln Gln
4852483PRTArtificial SequenceSynthetic amino acid sequence of the
mature CspAmy2-v1 a-amylase polypeptide 2Ala Ala Thr Asn Gly Thr
Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln
Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly
Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln
Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu
Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75
80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln
85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr
Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg
Asn Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr
Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe
Lys Trp Gln Trp Phe His Phe145 150 155 160Asp Gly Thr Asp Trp Asp
Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170 175Phe Thr Gly Lys
Ala Trp Asp Trp Glu Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp
Tyr Leu Met Tyr Ala Asp Ile Asp Tyr Asp His Pro Asp Val 195 200
205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly
210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe
Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala
Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn
Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr
Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr
Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile
Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315
320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu
325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe
Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly
Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro
Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys
Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn
Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys
Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly
Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440
445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser
450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly Gly Ser Val Ser
Val Trp465 470 475 480Val Gln Gln3480PRTArtificial
SequenceSynthetic amino acid sequence of the mature form of the
PcuAmy1 alpha-amylase polypeptide 3Ala Asp Asn Gly Thr Ile Met Gln
Tyr Phe Glu Trp Tyr Leu Pro Asn1 5 10 15Asp Gly Ala His Trp Asn Arg
Leu Asn Asn Asp Ala Gln Asn Leu Lys 20 25 30Asn Val Gly Ile Thr Ala
Val Trp Ile Pro Pro Ala Tyr Lys Gly Gly 35 40 45Ser Ser Ala Asp Val
Gly Tyr Gly Val Tyr Asp Thr Tyr Asp Leu Gly 50 55 60Glu Phe Asn Gln
Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys Ser65 70 75 80Glu Leu
Ile Ser Ala Val Asn Asn Leu His Ala Lys Gly Ile Ala Val 85 90 95Tyr
Gly Asp Val Val Leu Asn His Arg Met Asn Ala Asp Ala Thr Glu 100 105
110Leu Val Asp Ala Val Glu Val Asp Pro Asn Asn Arg Asn Val Glu Thr
115 120 125Thr Ser Thr Tyr Gln Ile Gln Ala Trp Thr Gln Tyr Asp Phe
Pro Gly 130 135 140Arg Gly Asn Thr Tyr Ser Ser Phe Lys Trp Arg Trp
Tyr His Phe Asp145 150 155 160Gly Val Asp Trp Asp Gln Ser Arg Gly
Leu Asn Arg Ile Tyr Lys Leu 165 170 175Arg Gly Asp Gly Lys Asp Trp
Asp Trp Glu Val Asp Ser Glu Tyr Gly 180 185 190Asn Tyr Asp Tyr Leu
Met Gly Ala Asp Leu Asp Phe Asn His Pro Asp 195 200 205Val Val Asn
Glu Thr Lys Thr Trp Gly Lys Trp Phe Val Asn Thr Val 210 215 220Asn
Leu Asp Gly Val Arg Leu Asp Ala Val Lys His Ile Lys Phe Asp225 230
235 240Phe Met Arg Asp Trp Val Asn Asn Val Arg Ser Thr Thr Gly Lys
Asn 245 250 255Leu Phe Ala Val Gly Glu Tyr Trp His Tyr Asp Val Asn
Lys Leu Asn 260 265 270Ser Tyr Ile Thr Lys Thr Asn Gly Thr Met Ser
Leu Phe Asp Val Pro 275 280 285Leu His Phe Arg Phe Tyr Asp Ala Ser
Asn Gly Gly Gly Gly Tyr Asp 290 295 300Met Arg Asn Leu Leu Asn Asn
Thr Leu Met Ser Ser Asn Pro Met Lys305 310 315 320Ala Val Thr Phe
Val Glu Asn His Asp Thr Gln Pro Thr Gln Ala Leu 325 330 335Gln Ser
Thr Val Gln Ser Trp Phe Lys Pro Leu Ala Tyr Ala Thr Ile 340 345
350Leu Thr Arg Glu Gln Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr Tyr
355 360 365Gly Thr Ser Asp Gly Lys Ile Ser Ser Tyr Lys Pro Ile Met
Asp Lys 370 375 380Leu Leu Asn Ala Arg Lys Val Tyr Ala Tyr Gly Thr
Gln Arg Asp Tyr385 390 395 400Phe Asp His Pro Asp Ile Val Gly Trp
Thr Arg Glu Gly Asp Ala Ala 405 410 415His Ala Gly Ser Gly Leu Ala
Thr Leu Ile Thr Asp Gly Pro Gly Gly 420 425 430Ser Lys Trp Met Tyr
Val Gly Thr Ser Lys Ala Gly Gln Val Trp Thr 435 440 445Asp Lys Thr
Gly Asn Arg Ser Gly Thr Val Thr Ile Asp Ala Asn Gly 450 455 460Trp
Gly Asn Phe Trp Val Asn Gly Gly Ser Val Ser Val Trp Ala Lys465 470
475 4804478PRTArtificial SequenceSynthetic amino acid sequence of a
variant form of PcuAmy1 alpha-amylase having a deletion of both
R177 and R178 4Ala Asp Asn Gly Thr Ile Met Gln Tyr Phe Glu Trp Tyr
Leu Pro Asn1 5 10 15Asp Gly Ala His Trp Asn Arg Leu Asn Asn Asp Ala
Gln Asn Leu Lys 20 25 30Asn Val Gly Ile Thr Ala Val Trp Ile Pro Pro
Ala Tyr Lys Gly Gly 35 40 45Ser Ser Ala Asp Val Gly Tyr Gly Val Tyr
Asp Thr Tyr Asp Leu Gly 50 55 60Glu Phe Asn Gln Lys Gly Thr Val Arg
Thr Lys Tyr Gly Thr Lys Ser65 70 75 80Glu Leu Ile Ser Ala Val Asn
Asn Leu His Ala Lys Gly Ile Ala Val 85 90 95Tyr Gly Asp Val Val Leu
Asn His Arg Met Asn Ala Asp Ala Thr Glu 100 105 110Leu Val Asp Ala
Val Glu Val Asp Pro Asn Asn Arg Asn Val Glu Thr 115 120 125Thr Ser
Thr Tyr Gln Ile Gln Ala Trp Thr Gln Tyr Asp Phe Pro Gly 130 135
140Arg Gly Asn Thr Tyr Ser Ser Phe Lys Trp Arg Trp Tyr His Phe
Asp145 150 155 160Gly Val Asp Trp Asp Gln Ser Arg Gly Leu Asn Arg
Ile Tyr Lys Leu 165 170 175Asp Gly Lys Asp Trp Asp Trp Glu Val Asp
Ser Glu Tyr Gly Asn Tyr 180 185 190Asp Tyr Leu Met Gly Ala Asp Leu
Asp Phe Asn His Pro Asp Val Val 195 200 205Asn Glu Thr Lys Thr Trp
Gly Lys Trp Phe Val Asn Thr Val Asn Leu 210 215 220Asp Gly Val Arg
Leu Asp Ala Val Lys His Ile Lys Phe Asp Phe Met225 230 235 240Arg
Asp Trp Val Asn Asn Val Arg Ser Thr Thr Gly Lys Asn Leu Phe 245 250
255Ala Val Gly Glu Tyr Trp His Tyr Asp Val Asn Lys Leu Asn Ser Tyr
260 265 270Ile Thr Lys Thr Asn Gly Thr Met Ser Leu Phe Asp Val Pro
Leu His 275 280 285Phe Arg Phe Tyr Asp Ala Ser Asn Gly Gly Gly Gly
Tyr Asp Met Arg 290 295 300Asn Leu Leu Asn Asn Thr Leu Met Ser Ser
Asn Pro Met Lys Ala Val305 310 315 320Thr Phe Val Glu Asn His Asp
Thr Gln Pro Thr Gln Ala Leu Gln Ser 325 330 335Thr Val Gln Ser Trp
Phe Lys Pro Leu Ala Tyr Ala Thr Ile Leu Thr 340 345 350Arg Glu Gln
Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr Tyr Gly Thr 355 360 365Ser
Asp Gly Lys Ile Ser Ser Tyr Lys Pro Ile Met Asp Lys Leu Leu 370 375
380Asn Ala Arg Lys Val Tyr Ala Tyr Gly Thr Gln Arg Asp Tyr Phe
Asp385 390 395 400His Pro Asp Ile Val Gly Trp Thr Arg Glu Gly Asp
Ala Ala His Ala 405 410 415Gly Ser Gly Leu Ala Thr Leu Ile Thr Asp
Gly Pro Gly Gly Ser Lys 420 425 430Trp Met Tyr Val Gly Thr Ser Lys
Ala Gly Gln Val Trp Thr Asp Lys 435 440 445Thr Gly Asn Arg Ser Gly
Thr Val Thr Ile Asp Ala Asn Gly Trp Gly 450 455 460Asn Phe Trp Val
Asn Gly Gly Ser Val Ser Val Trp Ala Lys465 470 4755484PRTArtificial
SequenceSynthetic amino acid sequence of a C-terminal- truncated
version of the Bacillus sp. TS-23 alpha-amylase 5Asn Thr Ala Pro
Ile Asn Glu Thr Met Met Gln Tyr Phe Glu Trp Asp1 5 10 15Leu Pro Asn
Asp Gly Thr Leu Trp Thr Lys Val Lys Asn Glu Ala Ala 20 25 30Asn Leu
Ser Ser Leu Gly Ile Thr Ala Leu Trp Leu Pro Pro Ala Tyr 35 40 45Lys
Gly Thr Ser Gln Ser Asp Val Gly Tyr Gly Val Tyr Asp Leu Tyr 50 55
60Asp Leu Gly Glu Phe Asn Gln Lys Gly Thr Ile Arg Thr Lys Tyr Gly65
70 75 80Thr Lys Thr Gln Tyr Ile Gln Ala Ile Gln Ala Ala Lys Ala Ala
Gly 85 90 95Met Gln Val Tyr Ala Asp Val Val Phe Asn His Lys Ala Gly
Ala Asp 100 105 110Gly Thr Glu Phe Val Asp Ala Val Glu Val Asp Pro
Ser Asn Arg Asn 115 120 125Gln Glu Thr Ser Gly Thr Tyr Gln Ile Gln
Ala Trp Thr Lys Phe Asp 130 135 140Phe Pro Gly Arg Gly Asn Thr Tyr
Ser Ser Phe Lys Trp Arg Trp Tyr145 150 155 160His Phe Asp Gly Thr
Asp Trp Asp Glu Ser Arg Lys Leu Asn Arg Ile 165 170 175Tyr Lys Phe
Arg Ser Thr Gly Lys Ala Trp Asp Trp Glu Val Asp Thr 180 185 190Glu
Asn Gly Asn Tyr Asp Tyr Leu Met Phe Ala Asp Leu Asp Met Asp 195 200
205His Pro Glu Val Val Thr Glu Leu Lys Asn Trp Gly Thr Trp Tyr Val
210 215 220Asn Thr Thr Asn Ile Asp Gly Phe Arg Leu Asp Ala Val Lys
His Ile225 230 235 240Lys Tyr Ser Phe Phe Pro Asp Trp Leu Thr Tyr
Val Arg Asn Gln Thr 245 250 255Gly Lys Asn Leu Phe Ala Val Gly Glu
Phe Trp Ser Tyr Asp Val Asn 260 265 270Lys Leu His Asn Tyr Ile Thr
Lys Thr Asn Gly Ser Met Ser Leu Phe 275 280 285Asp Ala Pro Leu His
Asn Asn Phe Tyr Thr Ala Ser Lys Ser Ser Gly 290 295 300Tyr Phe Asp
Met Arg Tyr Leu Leu Asn Asn Thr Leu Met Lys Asp Gln305 310 315
320Pro Ser Leu Ala Val Thr Leu Val Asp Asn His Asp Thr Gln Pro Gly
325 330 335Gln Ser Leu Gln Ser Trp Val Glu Pro Trp Phe Lys Pro Leu
Ala Tyr 340 345 350Ala Phe Ile Leu Thr Arg Gln Glu Gly Tyr Pro Cys
Val Phe Tyr Gly 355 360 365Asp Tyr Tyr Gly Ile Pro Lys Tyr Asn Ile
Pro Gly Leu Lys Ser Lys 370 375 380Ile Asp Pro Leu Leu Ile Ala Arg
Arg Asp Tyr Ala Tyr Gly Thr Gln385 390 395 400Arg Asp Tyr Ile Asp
His Gln Asp Ile Ile Gly Trp Thr Arg Glu Gly 405 410 415Ile Asp Thr
Lys Pro Asn Ser Gly Leu Ala Ala Leu Ile Thr Asp Gly 420 425 430Pro
Gly Gly Ser Lys Trp Met Tyr Val Gly Lys Lys His Ala Gly Lys 435 440
445Val Phe Tyr Asp Leu Thr Gly Asn Arg Ser Asp Thr Val Thr Ile Asn
450 455
460Ala Asp Gly Trp Gly Glu Phe Lys Val Asn Gly Gly Ser Val Ser
Ile465 470 475 480Trp Val Ala Lys6482PRTArtificial
SequenceSynthetic amino acid sequence of a variant form of BASE
alpha-amylase having a deletion of both R180 and S181 6Asn Thr Ala
Pro Ile Asn Glu Thr Met Met Gln Tyr Phe Glu Trp Asp1 5 10 15Leu Pro
Asn Asp Gly Thr Leu Trp Thr Lys Val Lys Asn Glu Ala Ala 20 25 30Asn
Leu Ser Ser Leu Gly Ile Thr Ala Leu Trp Leu Pro Pro Ala Tyr 35 40
45Lys Gly Thr Ser Gln Ser Asp Val Gly Tyr Gly Val Tyr Asp Leu Tyr
50 55 60Asp Leu Gly Glu Phe Asn Gln Lys Gly Thr Ile Arg Thr Lys Tyr
Gly65 70 75 80Thr Lys Thr Gln Tyr Ile Gln Ala Ile Gln Ala Ala Lys
Ala Ala Gly 85 90 95Met Gln Val Tyr Ala Asp Val Val Phe Asn His Lys
Ala Gly Ala Asp 100 105 110Gly Thr Glu Phe Val Asp Ala Val Glu Val
Asp Pro Ser Asn Arg Asn 115 120 125Gln Glu Thr Ser Gly Thr Tyr Gln
Ile Gln Ala Trp Thr Lys Phe Asp 130 135 140Phe Pro Gly Arg Gly Asn
Thr Tyr Ser Ser Phe Lys Trp Arg Trp Tyr145 150 155 160His Phe Asp
Gly Thr Asp Trp Asp Glu Ser Arg Lys Leu Asn Arg Ile 165 170 175Tyr
Lys Phe Thr Gly Lys Ala Trp Asp Trp Glu Val Asp Thr Glu Asn 180 185
190Gly Asn Tyr Asp Tyr Leu Met Phe Ala Asp Leu Asp Met Asp His Pro
195 200 205Glu Val Val Thr Glu Leu Lys Asn Trp Gly Thr Trp Tyr Val
Asn Thr 210 215 220Thr Asn Ile Asp Gly Phe Arg Leu Asp Ala Val Lys
His Ile Lys Tyr225 230 235 240Ser Phe Phe Pro Asp Trp Leu Thr Tyr
Val Arg Asn Gln Thr Gly Lys 245 250 255Asn Leu Phe Ala Val Gly Glu
Phe Trp Ser Tyr Asp Val Asn Lys Leu 260 265 270His Asn Tyr Ile Thr
Lys Thr Asn Gly Ser Met Ser Leu Phe Asp Ala 275 280 285Pro Leu His
Asn Asn Phe Tyr Thr Ala Ser Lys Ser Ser Gly Tyr Phe 290 295 300Asp
Met Arg Tyr Leu Leu Asn Asn Thr Leu Met Lys Asp Gln Pro Ser305 310
315 320Leu Ala Val Thr Leu Val Asp Asn His Asp Thr Gln Pro Gly Gln
Ser 325 330 335Leu Gln Ser Trp Val Glu Pro Trp Phe Lys Pro Leu Ala
Tyr Ala Phe 340 345 350Ile Leu Thr Arg Gln Glu Gly Tyr Pro Cys Val
Phe Tyr Gly Asp Tyr 355 360 365Tyr Gly Ile Pro Lys Tyr Asn Ile Pro
Gly Leu Lys Ser Lys Ile Asp 370 375 380Pro Leu Leu Ile Ala Arg Arg
Asp Tyr Ala Tyr Gly Thr Gln Arg Asp385 390 395 400Tyr Ile Asp His
Gln Asp Ile Ile Gly Trp Thr Arg Glu Gly Ile Asp 405 410 415Thr Lys
Pro Asn Ser Gly Leu Ala Ala Leu Ile Thr Asp Gly Pro Gly 420 425
430Gly Ser Lys Trp Met Tyr Val Gly Lys Lys His Ala Gly Lys Val Phe
435 440 445Tyr Asp Leu Thr Gly Asn Arg Ser Asp Thr Val Thr Ile Asn
Ala Asp 450 455 460Gly Trp Gly Glu Phe Lys Val Asn Gly Gly Ser Val
Ser Ile Trp Val465 470 475 480Ala Lys71536DNAArtificial
SequenceSynthetic DNA fragment encoding CspAmy2-v1 7atgaaacaac
aaaaacggct ttacgcccga ttgctgacgc tgttatttgc gctcatcttc 60ttgctgcctc
attctgcagc tagcgcagca gcgacaaacg gaacaatgat gcagtatttc
120gagtggtatg tacctaacga cggccagcaa tggaacagac tgagaacaga
tgccccttac 180ttgtcatctg ttggtattac agcagtatgg acaccgccgg
cttataaggg cacgtctcaa 240gcagatgtgg ggtacggccc gtacgatctg
tatgatttag gcgagtttaa tcaaaaaggt 300acagtcagaa cgaagtatgg
cacaaaagga gaacttaaat ctgctgttaa cacgctgcat 360tcaaatggaa
tccaagtgta tggtgatgtc gtgatgaatc ataaagcagg tgctgattat
420acagaaaacg taacggcggt ggaggtgaat ccgtctaata gaaatcagga
aacgagcggc 480gaatataata ttcaggcatg gacaggcttc aactttccgg
gcagaggaac aacgtattct 540aacttcaaat ggcagtggtt ccattttgat
ggaacggatt gggaccagag cagaagcctc 600tctagaatct tcaaattcac
gggaaaggcg tgggactggg aggtttcttc agaaaacgga 660aattatgact
atctgatgta cgcggacatt gattatgacc atccggatgt cgtgaatgaa
720atgaaaaagt ggggcgtctg gtatgccaac gaagttgggt tagatggata
cagacttgac 780gcggtcaaac atattaaatt tagctttctc aaagactggg
tggataacgc aagagcagcg 840acgggaaaag aaatgtttac ggttggcgaa
tattggcaaa atgatttagg ggccctgaat 900aactacctgg caaaggtaaa
ttacaaccaa tctctttttg atgcgccgtt gcattacaac 960ttttacgctg
cctcaacagg gggtggatat tacgatatga gaaatattct taataacacg
1020ttagtcgcaa gcaatccgac aaaggctgtt acgttagttg agaatcatga
cacacagcct 1080ggacaatcac tggaatcaac agtccaaccg tggtttaaac
cgttagccta cgcgtttatt 1140ctcacgagaa gcggaggcta tccttctgta
ttttatggag atatgtacgg tacaaaagga 1200acgacaacaa gagagatccc
tgctcttaaa tctaaaatcg aacctttgct taaggctaga 1260aaagactatg
cttatggaac acagagagac tatattgata acccggatgt cattggctgg
1320acgagagaag gggactcaac gaaagccaag agcggtctgg ccacagtgat
tacagatggg 1380ccgggcggtt caaaaagaat gtatgttggc acgagcaatg
cgggtgaaat ctggtatgat 1440ttgacaggga atagaacaga taaaatcacg
attggaagcg atggctatgc aacatttcct 1500gtcaatgggg gctcagtttc
agtatgggtg cagcaa 15368483PRTArtificial SequenceSynthetic amino
acid sequence of the mature form of CspAmy2-v5 8Ala Ala Thr Asn Gly
Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln
Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val
Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser
Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly
Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75
80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln
85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr
Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg
Asn Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr
Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe
Lys Trp Gln Trp Phe His Phe145 150 155 160Asp Gly Thr Asp Trp Asp
Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170 175Phe Thr Gly Lys
Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp
Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200
205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly
210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe
Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala
Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn
Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr
Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr
Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile
Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315
320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu
325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe
Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly
Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro
Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys
Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn
Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys
Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly
Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440
445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser
450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Lys Gly Ser Val Ser
Val Trp465 470 475 480Val Gln Gln9483PRTArtificial
SequenceSynthetic amino acid sequence of the mature form of
CspAmy2-v6 9Ala Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr
Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala
Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro
Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr
Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg
Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn
Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met
Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala
Val Glu Val Asn Pro Ser Asn Arg Asn Gln Glu 115 120 125Thr Ser Gly
Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly
Arg Gly Thr Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe His Phe145 150
155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe
Lys 165 170 175Phe Thr Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu
Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr
Asp His Pro Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val
Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp
Ala Val Lys His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp
Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr
Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265
270Tyr Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu
275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr
Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn
Pro Thr Lys Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr
Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe
Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly
Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly
Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu
Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390
395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly
Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr
Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser
Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Asn Ser
Thr Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro
Val Asn Lys Gly Ser Val Ser Val Trp465 470 475 480Val Gln
Gln10483PRTArtificial SequenceSynthetic amino acid sequence of
mature CspAmy2-16A 10Ala Ala Thr Asn Gly Thr Met Met Gln Tyr Phe
Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg
Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp
Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr
Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly
Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser
Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp
Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn
Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Tyr Gln Glu 115 120
125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro
130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe
His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu
Ser Arg Ile Phe Lys 165 170 175Phe Thr Gly Lys Ala Trp Asp Trp Pro
Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala
Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200 205Val Asn Glu Met Lys
Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly
Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe Ser Phe225 230 235
240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met
245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu
Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu Phe
Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly
Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr Leu
Val Ala Ser Asn Pro Thr Lys Ala305 310 315 320Val Thr Leu Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr Val
Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr
Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360
365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile
370 375 380Glu Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr
Gln Arg385 390 395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp
Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala
Thr Val Ile Thr Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr
Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr
Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr
Ala Thr Phe Pro Val Asn Gly Gly Ser Val Ser Val Trp465 470 475
480Val Gln Gln11483PRTArtificial SequenceSynthetic amino acid
sequence of mature CspAmy2-16B 11Ala Ala Thr Asn Gly Thr Met Met
Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn
Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr
Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp
Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn
Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu
Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val
Tyr Gly Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105
110Glu Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Tyr Gln Glu
115 120 125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn
Phe Pro 130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe Lys Trp Gln
Trp Phe His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg
Ser Leu Ser Arg Ile Phe Lys 165 170 175Phe Thr Gly Lys Ala Trp Asp
Trp Glu Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr
Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200
205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly
210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe
Gln Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala
Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn
Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr
Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr
Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile
Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315
320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu
325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe
Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly
Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro
Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys
Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn
Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys
Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly
Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440
445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser
450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly Gly Ser Val Ser
Val Trp465 470 475 480Val Gln Gln12483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-16C 12Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr Tyr Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170
175Phe Thr Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly
Gly Ser Val Ser Val Trp465 470 475 480Val Gln Gln13483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-16D 13Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr Tyr Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170
175Phe Thr Gly Lys Ala Trp Asp Trp Glu Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Gln Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly
Gly Ser Val Ser Val Trp465 470 475 480Val Gln Gln14483PRTArtificial
SequenceSynthetic amino acid sequence of the mature CspAmy2-16E
14Ala Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1
5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr
Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr
Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu
Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys
Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu
His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His
Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu
Val Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr
Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly
Thr Thr Tyr Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155
160Asp Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys
165 170 175Phe His Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn
Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp
His Pro Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp
Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala
Val Lys His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val
Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val
Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr
Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280
285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met
290 295 300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr
Lys Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro
Gly Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro
Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro
Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr
Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu
Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395
400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp
405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp
Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn
Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp
Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val
Asn Gly Gly Ser Val Ser Val Trp465 470 475 480Val Gln
Gln15483PRTArtificial SequenceSynthetic amino acid sequence of
mature CspAmy2-16F 15Ala Ala Thr Asn Gly Thr Met Met Gln Tyr Phe
Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg
Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp
Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr
Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly
Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser
Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp
Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn
Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Tyr Gln Glu 115 120
125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro
130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Trp Lys Trp Gln Trp Phe
His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu
Ser Arg Ile Phe Lys 165 170 175Phe His Gly Lys Ala Trp Asp Trp Glu
Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala
Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200 205Val Asn Glu Met Lys
Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly
Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe Gln Phe225 230 235
240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met
245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu
Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu Phe
Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly
Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr Leu
Val Ala Ser Asn Pro Thr Lys Ala305 310 315 320Val Thr Leu Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr Val
Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr
Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360
365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile
370 375 380Glu Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr
Gln Arg385 390 395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp
Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala
Thr Val Ile Thr Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr
Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr
Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr
Ala Thr Phe Pro Val Asn Gly Gly Ser Val Ser Val Trp465 470 475
480Val Gln Gln16483PRTArtificial SequenceSynthetic amino acid
sequence of mature CspAmy2-16G 16Ala Ala Thr Asn Gly Thr Met Met
Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn
Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr
Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp
Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn
Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu
Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val
Tyr Gly Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105
110Glu Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Tyr Gln Glu
115 120 125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn
Phe Pro 130 135 140Gly Arg Gly Thr Thr His Ser Asn Trp Lys Trp Gln
Trp Phe His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg
Ser Asn Ser Arg Ile Phe Lys 165 170 175Phe Thr Gly Lys Ala Trp Asp
Trp Pro Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met
Tyr Ala Asp Tyr Asp Tyr Asp His Pro Asp Val 195
200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala Asn Glu Val
Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys
Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala
Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln
Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn
Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe
Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn
Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315
320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu
325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe
Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly
Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro
Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys
Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn
Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys
Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly
Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440
445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser
450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly Gly Ser Val Ser
Val Trp465 470 475 480Val Gln Gln17483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-16H 17Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr His Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Asn Ser Arg Ile Phe Lys 165 170
175Phe Thr Gly Lys Ala Trp Asp Trp Glu Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Gln Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly
Gly Ser Val Ser Val Trp465 470 475 480Val Gln Gln18483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-16I 18Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr His Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Asn Ser Arg Ile Phe Lys 165 170
175Phe His Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly
Gly Ser Val Ser Val Trp465 470 475 480Val Gln Gln19483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-16J 19Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr His Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Asn Ser Arg Ile Phe Lys 165 170
175Phe His Gly Lys Ala Trp Asp Trp Glu Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Gln Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly
Gly Ser Val Ser Val Trp465 470 475 480Val Gln Gln20483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-v1-E187P
20Ala Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1
5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr
Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr
Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu
Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys
Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu
His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His
Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu
Val Asn Pro Ser Asn Arg Asn Gln Glu 115 120 125Thr Ser Gly Glu Tyr
Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly
Thr Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe His Phe145 150 155
160Asp Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys
165 170 175Phe Thr Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn
Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Ile Asp Tyr Asp
His Pro Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp
Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala
Val Lys His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val
Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val
Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr
Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280
285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met
290 295 300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr
Lys Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro
Gly Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro
Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro
Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr
Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu
Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395
400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp
405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp
Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn
Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp
Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val
Asn Gly Gly Ser Val Ser Val Trp465 470 475 480Val Gln
Gln21483PRTArtificial SequenceSynthetic amino acid sequence of
mature CspAmy2-v1-S241Q 21Ala Ala Thr Asn Gly Thr Met Met Gln Tyr
Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu
Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val
Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly
Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys
Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys
Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly
Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu
Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Asn Gln Glu 115 120
125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro
130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe
His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu
Ser Arg Ile Phe Lys 165 170 175Phe Thr Gly Lys Ala Trp Asp Trp Glu
Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala
Asp Ile Asp Tyr Asp His Pro Asp Val 195 200 205Val Asn Glu Met Lys
Lys Trp
Gly Val Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg
Leu Asp Ala Val Lys His Ile Lys Phe Gln Phe225 230 235 240Leu Lys
Asp Trp Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250
255Phe Thr Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn
260 265 270Tyr Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala
Pro Leu 275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly
Tyr Tyr Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala
Ser Asn Pro Thr Lys Ala305 310 315 320Val Thr Leu Val Glu Asn His
Asp Thr Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro
Trp Phe Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser
Gly Gly Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr
Lys Gly Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375
380Glu Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln
Arg385 390 395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr
Arg Glu Gly Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr
Val Ile Thr Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr Val
Gly Thr Ser Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly
Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala
Thr Phe Pro Val Asn Gly Gly Ser Val Ser Val Trp465 470 475 480Val
Gln Gln22483PRTArtificial SequenceSynthetic amino acid sequence of
mature CspAmy2-v171 22Ala Ala Thr Asn Gly Thr Met Met Gln Tyr Phe
Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg
Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp
Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr
Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly
Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser
Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp
Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn
Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Asn Gln Glu 115 120
125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro
130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe
His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu
Ser Arg Ile Phe Lys 165 170 175Phe Asp Gly Lys Ala Trp Asp Trp Pro
Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala
Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200 205Val Asn Glu Met Lys
Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly
Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe Ser Phe225 230 235
240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met
245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu
Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu Phe
Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly
Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr Leu
Val Ala Ser Asn Pro Thr Lys Ala305 310 315 320Val Thr Leu Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr Val
Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr
Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360
365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile
370 375 380Glu Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr
Gln Arg385 390 395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp
Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala
Thr Val Ile Thr Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr
Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr
Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr
Ala Thr Phe Pro Val Asn Lys Gly Ser Val Ser Val Trp465 470 475
480Val Gln Gln23483PRTArtificial SequenceSynthetic amino acid
sequence of mature CspAmy2-v172 23Ala Ala Thr Asn Gly Thr Met Met
Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn
Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr
Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp
Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn
Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu
Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val
Tyr Gly Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105
110Glu Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Tyr Gln Glu
115 120 125Thr Ser Gly Glu Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn
Phe Pro 130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe Lys Trp Gln
Trp Phe His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Gln Ser Arg
Ser Leu Ser Arg Ile Phe Lys 165 170 175Phe Asp Gly Lys Ala Trp Asp
Trp Pro Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met
Tyr Ala Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200 205Val Asn Glu
Met Lys Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu
Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe Ser Phe225 230
235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu
Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala
Leu Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr Asn Gln Ser Leu
Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr
Gly Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr
Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315 320Val Thr Leu Val
Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr
Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345
350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly
355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser
Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr
Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile
Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly
Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg
Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp
Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser 450 455 460Asp
Gly Tyr Ala Thr Phe Pro Val Asn Lys Gly Ser Val Ser Val Trp465 470
475 480Val Gln Gln24483PRTArtificial SequenceSynthetic amino acid
sequence of the mature CspAmy2-C18P amylase polypeptide 24Ala Ala
Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn
Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25
30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly
35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp
Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly
Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His Ser
Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys Ala
Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val Asn
Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn Ile
Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr Thr
Tyr Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp Gly
Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170
175Phe Asp Gly Lys Ala Trp Asp Trp Glu Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Gln Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly
Gly Ser Val Ser Val Trp465 470 475 480Val Gln Gln25483PRTArtificial
SequenceSynthetic amino acid sequence of the mature CspAmy2-C25A
amylase polypeptide 25Ala Ala Thr Asn Gly Thr Met Met Gln Tyr Phe
Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg
Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp
Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr
Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly
Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser
Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp
Val Val Met Asn His Lys Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn
Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg Tyr Gln Glu 115 120
125Thr Ser Gly His Tyr Asn Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro
130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Trp Lys Trp Gln Trp Phe
His Phe145 150 155 160Asp Gly Thr Asp Trp Asp Glu Ser Arg Ser Leu
Ser Arg Ile Phe Lys 165 170 175Phe Asp Gly Lys Ala Trp Asp Trp Glu
Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp Tyr Leu Met Tyr Ala
Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200 205Val Asn Glu Met Lys
Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly
Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe Gln Phe225 230 235
240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met
245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu
Asn Asn 260 265 270Tyr Leu Phe Lys Val Asn Tyr Asn Gln Ser Leu Phe
Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly
Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr Leu
Val Ala Ser Asn Pro Thr Lys Ala305 310 315 320Val Thr Leu Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr Val
Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr
Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360
365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile
370 375 380Glu Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Lys
Gln Arg385 390 395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp
Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala
Thr Val Ile Thr Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr
Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr
Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr
Ala Thr Phe Pro Val Asn Gly Gly Ser Val Ser Val Trp465 470 475
480Val Gln Gln26483PRTArtificial SequenceSynthetic amino acid
sequence of the mature CspAmy2-C25B amylase polypeptide 26Ala Ala
Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn
Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25
30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly
35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp
Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly
Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His Ser
Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys Ala
Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val Asn
Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly His Tyr Asn Ile
Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr Thr
Tyr Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp Gly
Thr Asp Trp Asp Glu Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170
175Phe Asp Gly Lys Ala Trp Asp Trp Glu Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val
Trp Tyr Ala Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp
Ala Val Lys His Ile Lys Phe Gln Phe225 230 235 240Leu Lys Asp Trp
Val Asp Asn Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr
Val Gly Glu Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265
270Tyr Leu Phe Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu
275 280 285His Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Tyr Tyr
Asp Met 290 295 300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn
Pro Thr Lys Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr
Gln Pro Gly Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe
Lys Pro Leu Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly
Tyr Pro Ser Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly
Thr Thr Thr Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu
Pro Leu Leu Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390
395 400Asp Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly
Asp 405 410 415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr
Asp Gly Pro 420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser
Asn Ala Gly Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr
Asp Lys Ile Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro
Val Asn Gly Gly Ser Val Ser Val Trp465 470 475 480Val Gln
Gln27483PRTArtificial SequenceSynthetic amino acid sequence of the
mature CspAmy2-C25F amylase polypeptide 27Ala Ala Thr Asn Gly Thr
Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln Gln
Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val Gly
Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser Gln
Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly Glu
Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75
80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln
85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr
Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg
Tyr Gln Glu 115 120 125Thr Ser Gly His Tyr Asn Ile Gln Ala Trp Thr
Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Trp
Lys Trp Gln Trp Phe His Phe145 150 155 160Asp Gly Thr Asp Trp Asp
Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170 175Phe Asp Gly Lys
Ala Trp Asp Trp Glu Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp
Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200
205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly
210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe
Gln Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala
Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn
Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Phe Lys Val Asn Tyr
Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr
Ala Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile
Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315
320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu
325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe
Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly
Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro
Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys
Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn
Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys
Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly
Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440
445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser
450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Gly Gly Ser Val Ser
Val Trp465 470 475 480Val Gln Gln28483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-v179 28Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170
175Phe Asp Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Asp Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Thr
Glu Ser Val Ser Val Trp465 470 475 480Val Gln Gln29483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-v180 29Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170
175Phe Asp Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Asp Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Glu Gly
Gln Ser Val Ser Val Trp465 470 475 480Val Gln Gln30483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-v181 30Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Thr Ser Gly Glu Tyr Asn
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr Tyr Ser Asn Phe Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170
175Phe Asp Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys
His Ile Lys Phe Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn
Ala Arg Ala Ala Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu
Tyr Trp Gln Asn Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala
Lys Val Asn Tyr Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His
Tyr Asn Phe Tyr Ala Ala Ser Thr Gly Gly Gly Asp Tyr Asp Met 290 295
300Arg Asn Ile Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys
Ala305 310 315 320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly
Gln Ser Leu Glu 325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu
Ala Tyr Ala Phe Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser
Val Phe Tyr Gly Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr
Arg Glu Ile Pro Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu
Lys Ala Arg Lys Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp
Tyr Ile Asp Asn Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410
415Ser Thr Lys Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro
420 425 430Gly Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly
Glu Ile 435 440 445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile
Thr Ile Gly Ser 450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Glu Thr
Arg Ser Val Ser Val Trp465 470 475 480Val Gln Gln31483PRTArtificial
SequenceSynthetic amino acid sequence of mature CspAmy2-v186 31Ala
Ala Thr Asn Gly Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10
15Asn Asp Gly Gln Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu
20 25 30Ser Ser Val Gly Ile Asn Ala Val Trp Thr Pro Pro Ala Tyr Lys
Gly 35 40 45Thr Ser Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr
Asp Leu 50 55 60Gly Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr
Gly Thr Lys65 70 75 80Gly Glu Leu Lys Ser Ala Val His Thr Leu His
Ser Asn Gly Ile Gln 85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys
Ala Gly Ala Asp Tyr Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val
Asn Pro Ser Asn Arg Tyr Gln Glu 115 120 125Ile Ser Gly Glu Tyr Met
Ile Gln Ala Trp Thr Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr
Thr Tyr Ser Asn Trp Lys Trp Gln Trp Phe His Phe145 150 155 160Asp
Gly Thr Asp Trp Asp Gln Ser Arg Ser Arg Ser Arg Ile Phe Lys 165 170
175Phe Asp Gly Lys Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn
180 185 190Tyr Asp Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro
Asp Val 195 200 205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala
Asn Glu Val Gly 210
215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe Ser
Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala Thr
Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn Asp
Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr Asn
Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr Ala
Ala Ser Thr Gly Gly Gly Tyr Tyr Asp Met 290 295 300Arg Asn Ile Leu
Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315 320Val
Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu 325 330
335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe Ile Leu
340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly Asp Met
Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro Ala Leu
Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys Asp Tyr
Ala Tyr Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn Pro Asp
Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys Ala Lys
Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly Gly Ser
Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440 445Trp
Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser 450 455
460Asp Gly Tyr Ala Thr Phe Pro Val Asn Lys Glu Ser Val Ser Val
Trp465 470 475 480Val Gln Gln32483PRTArtificial SequenceSynthetic
amino acid sequence of mature CspAmy2-v191 32Ala Ala Thr Asn Gly
Thr Met Met Gln Tyr Phe Glu Trp Tyr Val Pro1 5 10 15Asn Asp Gly Gln
Gln Trp Asn Arg Leu Arg Thr Asp Ala Pro Tyr Leu 20 25 30Ser Ser Val
Gly Ile Thr Ala Val Trp Thr Pro Pro Ala Tyr Lys Gly 35 40 45Thr Ser
Gln Ala Asp Val Gly Tyr Gly Pro Tyr Asp Leu Tyr Asp Leu 50 55 60Gly
Glu Phe Asn Gln Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys65 70 75
80Gly Glu Leu Lys Ser Ala Val Asn Thr Leu His Ser Asn Gly Ile Gln
85 90 95Val Tyr Gly Asp Val Val Met Asn His Lys Ala Gly Ala Asp Tyr
Thr 100 105 110Glu Asn Val Thr Ala Val Glu Val Asn Pro Ser Asn Arg
Tyr Gln Glu 115 120 125Thr Ser Gly His Tyr Asn Ile Gln Ala Trp Thr
Gly Phe Asn Phe Pro 130 135 140Gly Arg Gly Thr Thr Tyr Ser Asn Phe
Lys Trp Gln Trp Phe His Phe145 150 155 160Asp Gly Thr Asp Trp Asp
Gln Ser Arg Ser Leu Ser Arg Ile Phe Lys 165 170 175Phe Asp Gly Lys
Ala Trp Asp Trp Pro Val Ser Ser Glu Asn Gly Asn 180 185 190Tyr Asp
Tyr Leu Met Tyr Ala Asp Tyr Asp Tyr Asp His Pro Asp Val 195 200
205Val Asn Glu Met Lys Lys Trp Gly Val Trp Tyr Ala Asn Glu Val Gly
210 215 220Leu Asp Gly Tyr Arg Leu Asp Ala Val Lys His Ile Lys Phe
Ser Phe225 230 235 240Leu Lys Asp Trp Val Asp Asn Ala Arg Ala Ala
Thr Gly Lys Glu Met 245 250 255Phe Thr Val Gly Glu Tyr Trp Gln Asn
Asp Leu Gly Ala Leu Asn Asn 260 265 270Tyr Leu Ala Lys Val Asn Tyr
Asn Gln Ser Leu Phe Asp Ala Pro Leu 275 280 285His Tyr Asn Phe Tyr
Ala Ala Ser Thr Gly Gly Gly Asp Tyr Asp Met 290 295 300Arg Asn Ile
Leu Asn Asn Thr Leu Val Ala Ser Asn Pro Thr Lys Ala305 310 315
320Val Thr Leu Val Glu Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu
325 330 335Ser Thr Val Gln Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe
Ile Leu 340 345 350Thr Arg Ser Gly Gly Tyr Pro Ser Val Phe Tyr Gly
Asp Met Tyr Gly 355 360 365Thr Lys Gly Thr Thr Thr Arg Glu Ile Pro
Ala Leu Lys Ser Lys Ile 370 375 380Glu Pro Leu Leu Lys Ala Arg Lys
Asp Tyr Ala Tyr Gly Thr Gln Arg385 390 395 400Asp Tyr Ile Asp Asn
Pro Asp Val Ile Gly Trp Thr Arg Glu Gly Asp 405 410 415Ser Thr Lys
Ala Lys Ser Gly Leu Ala Thr Val Ile Thr Asp Gly Pro 420 425 430Gly
Gly Ser Lys Arg Met Tyr Val Gly Thr Ser Asn Ala Gly Glu Ile 435 440
445Trp Tyr Asp Leu Thr Gly Asn Arg Thr Asp Lys Ile Thr Ile Gly Ser
450 455 460Asp Gly Tyr Ala Thr Phe Pro Val Asn Thr Glu Ser Val Ser
Val Trp465 470 475 480Val Gln Gln331443DNAArtificial
SequenceSynthetic codon-optimized nucleotide sequence of the
PcuAmy1 gene 33gccgacaacg gcacaatcat gcagtatttc gagtggtacc
tgccgaacga cggagcgcac 60tggaacagac ttaataacga cgcacaaaac ctgaaaaatg
tgggcatcac ggcagtgtgg 120attcctccgg catacaaggg cggcagctca
gcagatgttg gctacggagt ttacgataca 180tacgacctgg gcgagttcaa
tcagaaaggc acggtcagaa caaagtacgg aacgaagagc 240gaactgattt
cagcggtcaa caatcttcac gcaaagggca ttgcggttta cggcgacgtg
300gtcctgaacc atagaatgaa tgcggatgca acggagcttg tggatgcggt
tgaggtggat 360ccgaacaaca gaaacgtcga gacgacaagc acgtatcaga
tccaggcatg gacgcaatac 420gatttcccgg gcagaggcaa cacgtacagc
agctttaaat ggagatggta tcacttcgac 480ggcgtcgact gggaccagag
cagaggcctg aacagaatct ataagctgag aggcgatggc 540aaggattggg
actgggaggt cgacagcgag tacggcaact acgattacct gatgggagcg
600gacctggact tcaaccaccc ggatgtggtt aacgaaacaa agacatgggg
caaatggttt 660gtgaacacgg tgaacctgga tggcgtcaga ctggacgcgg
ttaagcacat caagttcgac 720ttcatgagag actgggtgaa caacgtgaga
agcacgacgg gcaagaacct tttcgcagtt 780ggcgagtatt ggcactacga
cgtgaacaaa ctgaacagct acatcacgaa gacgaatggc 840acgatgagcc
tgttcgacgt gccgctgcac tttagatttt atgatgcaag caacggcgga
900ggcggctacg acatgagaaa cctgctgaat aacacgctga tgagcagcaa
cccgatgaag 960gcggttacat tcgttgagaa ccatgacaca caaccgacgc
aggccctgca atcaacggtc 1020caaagctggt ttaagccgct tgcgtatgct
acaatcctga cgagagagca aggctacccg 1080tgcgttttct acggcgacta
ttatggaaca agcgacggca aaattagcag ctacaagccg 1140atcatggata
agcttcttaa cgcgagaaag gtgtacgcct acggcacgca gagagattac
1200ttcgatcatc cggacatcgt tggctggaca agagaaggcg atgcagcaca
tgctggctca 1260ggactggcaa cgcttatcac agatggccct ggcggaagca
agtggatgta tgttggaacg 1320tcaaaggcag gccaggtctg gacggataaa
acaggaaaca gaagcggaac ggtgacgatt 1380gatgccaatg gctggggaaa
cttttgggtt aatggcggat cagttagcgt ttgggcaaaa 1440taa
144334478PRTArtificial SequenceSynthetic amino acid sequence of the
mature of PcuAmy1-v1A polypeptide 34Ala Asp Asn Gly Thr Ile Met Gln
Tyr Phe Glu Trp Tyr Leu Pro Asn1 5 10 15Asp Gly Ala His Trp Asn Arg
Leu Asn Asn Asp Ala Gln Asn Leu Lys 20 25 30Asn Val Gly Ile Thr Ala
Val Trp Ile Pro Pro Ala Tyr Lys Gly Gly 35 40 45Ser Ser Ala Asp Val
Gly Tyr Gly Val Tyr Asp Thr Tyr Asp Leu Gly 50 55 60Glu Phe Asn Gln
Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys Ser65 70 75 80Glu Leu
Ile Ser Ala Val Asn Asn Leu His Ala Lys Gly Ile Ala Val 85 90 95Tyr
Gly Asp Val Val Leu Asn His Arg Met Asn Ala Asp Ala Thr Glu 100 105
110Leu Val Asp Ala Val Glu Val Asp Pro Asn Asn Arg Tyr Val Glu Thr
115 120 125Thr Ser Thr Tyr Gln Ile Gln Ala Trp Thr Gln Tyr Asp Phe
Pro Gly 130 135 140Arg Gly Asn Thr Tyr Ser Ser Phe Lys Trp Arg Trp
Tyr His Phe Asp145 150 155 160Gly Val Asp Trp Asp Gln Ser Arg Gly
Leu Asn Arg Ile Tyr Lys Leu 165 170 175Asp Gly Lys Asp Trp Asp Trp
Pro Val Asp Ser Glu Tyr Gly Asn Tyr 180 185 190Asp Tyr Leu Met Gly
Ala Asp Leu Asp Phe Asn His Pro Asp Val Val 195 200 205Asn Glu Thr
Lys Thr Trp Gly Lys Trp Phe Val Asn Thr Val Asn Leu 210 215 220Asp
Gly Val Arg Leu Asp Ala Val Lys His Ile Lys Phe Asp Phe Met225 230
235 240Arg Asp Trp Val Asn Asn Val Arg Ser Thr Thr Gly Lys Asn Leu
Phe 245 250 255Ala Val Gly Glu Tyr Trp His Tyr Asp Val Asn Lys Leu
Asn Ser Tyr 260 265 270Ile Thr Lys Thr Asn Gly Thr Met Ser Leu Phe
Asp Val Pro Leu His 275 280 285Phe Arg Phe Tyr Asp Ala Ser Asn Gly
Gly Gly Gly Tyr Asp Met Arg 290 295 300Asn Leu Leu Asn Asn Thr Leu
Met Ser Ser Asn Pro Met Lys Ala Val305 310 315 320Thr Phe Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu Ser 325 330 335Thr Val
Gln Ser Trp Phe Lys Pro Leu Ala Tyr Ala Thr Ile Leu Thr 340 345
350Arg Glu Gln Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr Tyr Gly Thr
355 360 365Ser Asp Gly Lys Ile Ser Ser Tyr Lys Pro Ile Met Asp Lys
Leu Leu 370 375 380Asn Ala Arg Lys Val Tyr Ala Tyr Gly Thr Gln Arg
Asp Tyr Phe Asp385 390 395 400His Pro Asp Ile Val Gly Trp Thr Arg
Glu Gly Asp Ala Ala His Ala 405 410 415Gly Ser Gly Leu Ala Thr Leu
Ile Thr Asp Gly Pro Gly Gly Ser Lys 420 425 430Trp Met Tyr Val Gly
Thr Ser Lys Ala Gly Gln Val Trp Thr Asp Lys 435 440 445Thr Gly Asn
Arg Ser Gly Thr Val Thr Ile Asp Ala Asn Gly Trp Gly 450 455 460Asn
Phe Trp Val Asn Lys Gly Ser Val Ser Val Trp Ala Lys465 470
47535478PRTArtificial SequenceSynthetic amino acid sequence of the
mature of PcuAmy1-v6 polypeptide 35Ala Asp Asn Gly Thr Ile Met Gln
Tyr Phe Glu Trp Tyr Leu Pro Asn1 5 10 15Asp Gly Ala His Trp Asn Arg
Leu Asn Asn Asp Ala Gln Asn Leu Lys 20 25 30Asn Val Gly Ile Thr Ala
Val Trp Ile Pro Pro Ala Tyr Lys Gly Gly 35 40 45Ser Ser Ala Asp Val
Gly Tyr Gly Val Tyr Asp Thr Tyr Asp Leu Gly 50 55 60Glu Phe Asn Gln
Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys Ser65 70 75 80Glu Leu
Ile Ser Ala Val Asn Asn Leu His Ala Lys Gly Ile Ala Val 85 90 95Tyr
Gly Asp Val Val Leu Asn His Arg Met Asn Ala Asp Ala Thr Glu 100 105
110Leu Val Asp Ala Val Glu Val Asp Pro Asn Asn Arg Tyr Val Glu Thr
115 120 125Thr Ser Thr Tyr Gln Ile Gln Ala Trp Thr Gln Tyr Asp Phe
Pro Gly 130 135 140Arg Gly Asn Thr Tyr Ser Ser Trp Lys Trp Arg Trp
Tyr His Phe Asp145 150 155 160Gly Val Asp Trp Asp Gln Ser Arg Gly
Leu Asn Arg Ile Tyr Lys Leu 165 170 175Asp Gly Lys Asp Trp Asp Trp
Pro Val Asp Ser Glu Tyr Gly Asn Tyr 180 185 190Asp Tyr Leu Met Gly
Ala Asp Leu Asp Phe Asn His Pro Asp Val Val 195 200 205Asn Glu Thr
Lys Thr Trp Gly Lys Trp Phe Val Asn Thr Val Asn Leu 210 215 220Asp
Gly Val Arg Leu Asp Ala Val Lys His Ile Lys Phe Asp Phe Met225 230
235 240Arg Asp Trp Val Asn Asn Val Arg Ser Thr Thr Gly Lys Asn Leu
Phe 245 250 255Ala Val Gly Glu Tyr Trp His Tyr Asp Val Asn Lys Leu
Asn Ser Tyr 260 265 270Ile Thr Lys Thr Asn Gly Thr Met Ser Leu Phe
Asp Val Pro Leu His 275 280 285Phe Arg Phe Tyr Asp Ala Ser Asn Gly
Gly Gly Gly Tyr Asp Met Arg 290 295 300Asn Leu Leu Asn Asn Thr Leu
Met Ser Ser Asn Pro Met Lys Ala Val305 310 315 320Thr Phe Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu Ser 325 330 335Thr Val
Gln Ser Trp Phe Lys Pro Leu Ala Tyr Ala Thr Ile Leu Thr 340 345
350Arg Glu Gln Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr Tyr Gly Thr
355 360 365Ser Asp Gly Lys Ile Ser Ser Tyr Lys Pro Ile Met Asp Lys
Leu Leu 370 375 380Asn Ala Arg Lys Val Tyr Ala Tyr Gly Thr Gln Arg
Asp Tyr Phe Asp385 390 395 400His Pro Asp Ile Val Gly Trp Thr Arg
Glu Gly Asp Ala Ala His Ala 405 410 415Gly Ser Gly Leu Ala Thr Leu
Ile Thr Asp Gly Pro Gly Gly Ser Lys 420 425 430Trp Met Tyr Val Gly
Thr Ser Lys Ala Gly Gln Val Trp Thr Asp Lys 435 440 445Thr Gly Asn
Arg Ser Gly Thr Val Thr Ile Asp Ala Asn Gly Trp Gly 450 455 460Asn
Phe Trp Val Asn Lys Gly Ser Val Ser Val Trp Ala Lys465 470
47536478PRTArtificial SequenceSynthetic amino acid sequence of the
mature of PcuAmy1-v8 polypeptide 36Ala Asp Asn Gly Thr Ile Met Gln
Tyr Phe Glu Trp Tyr Leu Pro Asn1 5 10 15Asp Gly Ala His Trp Asn Arg
Leu Asn Asn Asp Ala Gln Asn Leu Lys 20 25 30Asn Val Gly Ile Thr Ala
Val Trp Ile Pro Pro Ala Tyr Lys Gly Gly 35 40 45Ser Ser Ala Asp Val
Gly Tyr Gly Val Tyr Asp Thr Tyr Asp Leu Gly 50 55 60Glu Phe Asn Gln
Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys Ser65 70 75 80Glu Leu
Ile Ser Ala Val Asn Asn Leu His Ala Lys Gly Ile Ala Val 85 90 95Tyr
Gly Asp Val Val Leu Asn His Arg Met Asn Ala Asp Ala Thr Glu 100 105
110Leu Val Asp Ala Val Glu Val Asp Pro Asn Asn Arg Tyr Val Glu Thr
115 120 125Thr Ser Thr Tyr Gln Ile Gln Ala Trp Thr Gln Tyr Asp Phe
Pro Gly 130 135 140Arg Gly Asn Thr Tyr Ser Ser Trp Lys Trp Arg Trp
Tyr His Phe Asp145 150 155 160Gly Val Asp Trp Asp Gln Ser Arg Gly
Leu Asn Arg Ile Tyr Lys Leu 165 170 175Asp Gly Lys Asp Trp Asp Trp
Pro Val Asp Ser Glu Tyr Gly Asn Tyr 180 185 190Asp Tyr Leu Met Gly
Ala Asp Leu Asp Phe Asn His Pro Asp Val Val 195 200 205Asn Glu Thr
Lys Thr Trp Gly Lys Trp Phe Val Asn Thr Val Asn Leu 210 215 220Asp
Gly Val Arg Leu Asp Ala Val Lys His Ile Lys Phe Asp Phe Met225 230
235 240Arg Asp Trp Val Asn Asn Val Arg Ser Thr Thr Gly Lys Asn Leu
Phe 245 250 255Ala Val Gly Glu Tyr Trp His Tyr Asp Val Asn Lys Leu
Asn Ser Tyr 260 265 270Ile Thr Lys Thr Asn Gly Thr Met Ser Leu Phe
Asp Val Pro Leu His 275 280 285Phe Arg Phe Tyr Asp Ala Ser Asn Gly
Gly Gly Gly Tyr Asp Met Arg 290 295 300Asn Leu Leu Asn Asn Thr Leu
Met Ser Ser Asn Pro Met Lys Ala Val305 310 315 320Thr Phe Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu Ser 325 330 335Thr Val
Gln Ser Trp Phe Lys Pro Leu Ala Tyr Ala Thr Ile Leu Thr 340 345
350Arg Glu Gln Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr Tyr Gly Thr
355 360 365Ser Asp Gly Lys Ile Ser Ser Tyr Lys Pro Ile Met Asp Lys
Leu Leu 370 375 380Asn Ala Arg Lys Val Tyr Ala Tyr Gly Thr Gln Arg
Asp Tyr Phe Asp385 390 395 400His Pro Asp Ile Val Gly Trp Thr Arg
Glu Gly Asp Ala Ala His Ala 405 410 415Gly Ser Gly Leu Ala Thr Leu
Ile Thr Asp Gly Pro Gly Gly Ser Lys 420 425 430Trp Met Tyr Val Gly
Thr Ser Lys Ala Gly Gln Val Trp Thr Asp Lys 435 440 445Thr Gly Asn
Arg Ser Gly Thr Val Thr Ile Asp Ala Asn Gly Trp Gly 450 455 460Asn
Phe Trp
Val Asn Arg Arg Ser Val Ser Val Trp Ala Lys465 470
47537478PRTArtificial SequenceSynthetic amino acid sequence of the
mature of PcuAmy1-v16 polypeptide 37Ala Asp Asn Gly Thr Ile Met Gln
Tyr Phe Glu Trp Tyr Leu Pro Asn1 5 10 15Asp Gly Ala His Trp Asn Arg
Leu Asn Asn Asp Ala Gln Asn Leu Lys 20 25 30Asn Val Gly Ile Thr Ala
Val Trp Ile Pro Pro Ala Tyr Lys Gly Gly 35 40 45Ser Ser Ala Asp Val
Gly Tyr Gly Val Tyr Asp Thr Tyr Asp Leu Gly 50 55 60Glu Phe Asn Gln
Lys Gly Thr Val Arg Thr Lys Tyr Gly Thr Lys Ser65 70 75 80Glu Leu
Ile Ser Ala Val Asn Asn Leu His Ala Lys Gly Ile Ala Val 85 90 95Tyr
Gly Asp Val Val Leu Asn His Arg Met Asn Ala Asp Ala Thr Glu 100 105
110Leu Val Asp Ala Val Glu Val Asp Pro Asn Asn Arg Tyr Val Glu Thr
115 120 125Thr Ser Thr Tyr Gln Ile Gln Ala Trp Thr Gln Tyr Asp Phe
Pro Gly 130 135 140Arg Gly Asn Thr Tyr Ser Ser Trp Lys Trp Arg Trp
Tyr His Phe Asp145 150 155 160Gly Val Asp Trp Asp Gln Ser Arg Gly
Leu Asn Arg Ile Tyr Lys Leu 165 170 175Asp Gly Lys Asp Trp Asp Trp
Pro Val Asp Ser Glu Tyr Gly Asn Tyr 180 185 190Asp Tyr Leu Met Gly
Ala Asp Leu Asp Phe Asp His Pro Asp Val Val 195 200 205Asn Glu Thr
Lys Thr Trp Gly Lys Trp Phe Val Asn Thr Val Asn Leu 210 215 220Asp
Gly Val Arg Leu Asp Ala Val Lys His Ile Lys Phe Asp Phe Met225 230
235 240Arg Asp Trp Val Asn Asn Val Arg Ser Thr Thr Gly Lys Asn Leu
Phe 245 250 255Ala Val Gly Glu Tyr Trp His Tyr Asp Val Asn Lys Leu
Asn Ser Tyr 260 265 270Ile Thr Lys Thr Asn Gly Thr Met Ser Leu Phe
Asp Val Pro Leu His 275 280 285Phe Arg Phe Tyr Asp Ala Ser Asn Gly
Gly Gly Gly Tyr Asp Met Arg 290 295 300Asn Leu Leu Asn Asn Thr Leu
Met Ser Ser Asn Pro Met Lys Ala Val305 310 315 320Thr Phe Val Glu
Asn His Asp Thr Gln Pro Gly Gln Ser Leu Glu Ser 325 330 335Thr Val
Gln Ser Trp Phe Lys Pro Leu Ala Tyr Ala Thr Ile Leu Thr 340 345
350Arg Glu Gln Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr Tyr Gly Thr
355 360 365Ser Asp Gly Lys Ile Ser Ser Tyr Lys Pro Ile Met Asp Lys
Leu Leu 370 375 380Asn Ala Arg Lys Val Tyr Ala Tyr Gly Thr Gln Arg
Asp Tyr Phe Asp385 390 395 400His Pro Asp Ile Val Gly Trp Thr Arg
Glu Gly Asp Ala Ala His Ala 405 410 415Gly Ser Gly Leu Ala Thr Leu
Ile Thr Asp Gly Pro Gly Gly Ser Lys 420 425 430Trp Met Tyr Val Gly
Thr Ser Lys Ala Gly Gln Val Trp Thr Asp Lys 435 440 445Thr Gly Asn
Arg Ser Gly Thr Val Thr Ile Asp Ala Asn Gly Trp Gly 450 455 460Asn
Phe Trp Val Asn Lys Gly Ser Val Ser Val Trp Ala Lys465 470
475381479DNAArtificial SequenceSynthetic codon-modified nucleic
acid sequence encoding the mature form of BASE 38tctgcagctt
cagcaaacac cgcgccgatt aacgaaacca tgatgcagta tttcgaatgg 60gatctgccga
acgatggcac cctgtggacc aaagtgaaaa acgaagcggc gaacctgagc
120agcctgggca ttaccgcgct gtggctgccg ccggcatata aaggcaccag
ccagagcgat 180gtgggctatg gcgtgtatga tctgtacgat ctgggcgaat
ttaaccagaa aggcaccatt 240cgtaccaaat atggcaccaa aacccagtat
attcaggcga tccaggcggc gaaagcggcg 300ggtatgcagg tgtatgcgga
tgtggtgttt aaccataaag cgggtgcgga tggcaccgaa 360tttgtggatg
cggtggaagt ggatccgagc aaccgtaacc aggaaaccag cggcacctat
420cagattcagg cgtggaccaa atttgatttt cccggccgtg gcaacaccta
tagcagcttt 480aaatggcgct ggtatcattt tgatggcacc gattgggatg
aaagccgtaa actgaaccgc 540atctataaat ttcgtagcac cggcaaagcg
tgggattggg aagtggatac cgaaaacggc 600aactatgatt acctgatgtt
cgcagacctg gatatggatc atccggaagt ggtgaccgaa 660ctgaaaaact
ggggcacctg gtatgtgaac accaccaaca ttgatggctt tcgtctggat
720gcggtgaaac acatcaaata cagctttttt ccggattggc tgacctatgt
gcgtaaccag 780accggcaaaa acctgtttgc ggtgggcgaa ttttggagct
atgatgtgaa caaactgcac 840aactacatca ccaaaaccaa cggcagcatg
agcctgtttg atgcgccgct gcataacaac 900ttttataccg cgagcaaaag
cagcggctat tttgatatgc gttatctgct gaacaacacc 960ctgatgaaag
atcagccgag cctggccgtg accctggtgg ataaccatga tacccagccg
1020ggccagagcc tgcaaagctg ggtggaaccg tggtttaaac cgctggccta
cgcgtttatt 1080ctgacccgtc aagagggcta tccgtgcgtt ttttatggcg
attattacgg catcccgaaa 1140tataacattc cgggcctgaa aagcaaaatt
gatccgctgc tgattgcgcg tcgtgattat 1200gcgtatggca cccagcgtga
ttatattgat caccaggata ttattggctg gacccgtgaa 1260ggcattgata
ccaaaccgaa cagcggcctg gccgcgctga ttaccgatgg cccgggtggc
1320agcaaatgga tgtatgtggg caaaaaacat gcgggcaaag tgttttatga
tctgaccggc 1380aaccgtagcg ataccgtgac cattaacgcg gatggctggg
gtgagtttaa agtgaacggc 1440ggcagcgtga gcatttgggt ggcgaaataa
gttaacaga 147939482PRTArtificial SequenceSynthetic amino acid
sequence of BASE-V28 39Asn Thr Ala Pro Ile Asn Glu Thr Met Met Gln
Tyr Phe Glu Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu Trp Thr Lys
Val Lys Asn Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly Ile Thr Ala
Leu Trp Leu Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln Ser Asp Val
Gly Tyr Gly Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu Phe Asn Gln
Lys Gly Thr Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys Thr Gln Tyr
Ile Gln Ala Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met Gln Val Tyr
Ala Asp Val Val Phe Asn His Lys Ala Gly Ala Asp 100 105 110Gly Thr
Glu Phe Val Asp Ala Val Glu Val Asp Pro Ser Asn Arg Tyr 115 120
125Gln Glu Thr Ser Gly Thr Tyr Gln Ile Gln Ala Trp Thr Lys Phe Asp
130 135 140Phe Pro Gly Arg Gly Asn Thr Tyr Ser Ser Phe Lys Trp Arg
Trp Tyr145 150 155 160His Phe Asp Gly Thr Asp Trp Asp Glu Ser Arg
Lys Leu Asn Arg Ile 165 170 175Tyr Lys Phe Thr Gly Lys Ala Trp Asp
Trp Pro Val Asp Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr Leu Met
Phe Ala Asp Leu Asp Met Asp His Pro 195 200 205Glu Val Val Thr Glu
Leu Lys Asn Trp Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr Asn Ile
Asp Gly Phe Arg Leu Asp Ala Val Lys His Ile Lys Tyr225 230 235
240Ser Phe Phe Pro Asp Trp Leu Thr Tyr Val Arg Asn Gln Thr Gly Lys
245 250 255Asn Leu Phe Ala Val Gly Glu Phe Trp Ser Tyr Asp Val Asn
Lys Leu 260 265 270His Asn Tyr Ile Thr Lys Thr Asn Gly Ser Met Ser
Leu Phe Asp Ala 275 280 285Pro Leu His Asn Asn Phe Tyr Thr Ala Ser
Lys Ser Ser Gly Tyr Phe 290 295 300Asp Met Arg Tyr Leu Leu Asn Asn
Thr Leu Met Lys Asp Gln Pro Ser305 310 315 320Leu Ala Val Thr Leu
Val Asp Asn His Asp Thr Gln Pro Gly Gln Ser 325 330 335Leu Gln Ser
Trp Val Glu Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe 340 345 350Ile
Leu Thr Arg Gln Glu Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr 355 360
365Tyr Gly Ile Pro Lys Tyr Asn Ile Pro Gly Leu Lys Ser Lys Ile Asp
370 375 380Pro Leu Leu Ile Ala Arg Arg Asp Tyr Ala Tyr Gly Thr Gln
Arg Asp385 390 395 400Tyr Ile Asp His Gln Asp Ile Ile Gly Trp Thr
Arg Glu Gly Ile Asp 405 410 415Thr Lys Pro Asn Ser Gly Leu Ala Ala
Leu Ile Thr Asp Gly Pro Gly 420 425 430Gly Ser Lys Trp Met Tyr Val
Gly Lys Lys His Ala Gly Lys Val Phe 435 440 445Tyr Asp Leu Thr Gly
Asn Arg Ser Asp Thr Val Thr Ile Asn Ala Asp 450 455 460Gly Trp Gly
Glu Phe Lys Val Asn Arg Gly Ser Val Ser Ile Trp Val465 470 475
480Ala Lys40482PRTArtificial SequenceSynthetic amino acid sequence
of BASE-V29 40Asn Thr Ala Pro Ile Asn Glu Thr Met Met Gln Tyr Phe
Glu Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu Trp Thr Lys Val Lys
Asn Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly Ile Thr Ala Leu Trp
Leu Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln Ser Asp Val Gly Tyr
Gly Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu Phe Asn Gln Lys Gly
Thr Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys Thr Gln Tyr Ile Gln
Ala Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met Gln Val Tyr Ala Asp
Val Val Phe Asn His Lys Ala Gly Ala Asp 100 105 110Gly Thr Glu Phe
Val Asp Ala Val Glu Val Asp Pro Ser Asn Arg Asn 115 120 125Gln Glu
Thr Ser Gly Thr Tyr Gln Ile Gln Ala Trp Thr Lys Phe Asp 130 135
140Phe Pro Gly Arg Gly Asn Thr Tyr Ser Ser Trp Lys Trp Arg Trp
Tyr145 150 155 160His Phe Asp Gly Thr Asp Trp Asp Glu Ser Arg Lys
Leu Asn Arg Ile 165 170 175Tyr Lys Phe Thr Gly Lys Ala Trp Asp Trp
Pro Val Asp Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr Leu Met Phe
Ala Asp Leu Asp Met Asp His Pro 195 200 205Glu Val Val Thr Glu Leu
Lys Asn Trp Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr Asn Ile Asp
Gly Phe Arg Leu Asp Ala Val Lys His Ile Lys Tyr225 230 235 240Ser
Phe Phe Pro Asp Trp Leu Thr Tyr Val Arg Asn Gln Thr Gly Lys 245 250
255Asn Leu Phe Ala Val Gly Glu Phe Trp Ser Tyr Asp Val Asn Lys Leu
260 265 270His Asn Tyr Ile Thr Lys Thr Asn Gly Ser Met Ser Leu Phe
Asp Ala 275 280 285Pro Leu His Asn Asn Phe Tyr Thr Ala Ser Lys Ser
Ser Gly Tyr Phe 290 295 300Asp Met Arg Tyr Leu Leu Asn Asn Thr Leu
Met Lys Asp Gln Pro Ser305 310 315 320Leu Ala Val Thr Leu Val Asp
Asn His Asp Thr Gln Pro Gly Gln Ser 325 330 335Leu Gln Ser Trp Val
Glu Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe 340 345 350Ile Leu Thr
Arg Gln Glu Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr 355 360 365Tyr
Gly Ile Pro Lys Tyr Asn Ile Pro Gly Leu Lys Ser Lys Ile Asp 370 375
380Pro Leu Leu Ile Ala Arg Arg Asp Tyr Ala Tyr Gly Thr Gln Arg
Asp385 390 395 400Tyr Ile Asp His Gln Asp Ile Ile Gly Trp Thr Arg
Glu Gly Ile Asp 405 410 415Thr Lys Pro Asn Ser Gly Leu Ala Ala Leu
Ile Thr Asp Gly Pro Gly 420 425 430Gly Ser Lys Trp Met Tyr Val Gly
Lys Lys His Ala Gly Lys Val Phe 435 440 445Tyr Asp Leu Thr Gly Asn
Arg Ser Asp Thr Val Thr Ile Asn Ala Asp 450 455 460Gly Trp Gly Glu
Phe Lys Val Asn Arg Gly Ser Val Ser Ile Trp Val465 470 475 480Ala
Lys41482PRTArtificial SequenceSynthetic amino acid sequence of
BASE-V30 41Asn Thr Ala Pro Ile Asn Glu Thr Met Met Gln Tyr Phe Glu
Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu Trp Thr Lys Val Lys Asn
Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly Ile Thr Ala Leu Trp Leu
Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln Ser Asp Val Gly Tyr Gly
Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu Phe Asn Gln Lys Gly Thr
Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys Thr Gln Tyr Ile Gln Ala
Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met Gln Val Tyr Ala Asp Val
Val Phe Asn His Lys Ala Gly Ala Asp 100 105 110Gly Thr Glu Phe Val
Asp Ala Val Glu Val Asp Pro Ser Asn Arg Asn 115 120 125Gln Glu Thr
Ser Gly Glu Tyr Gln Ile Gln Ala Trp Thr Lys Phe Asp 130 135 140Phe
Pro Gly Arg Gly Asn Thr Tyr Ser Ser Phe Lys Trp Arg Trp Tyr145 150
155 160His Phe Asp Gly Thr Asp Trp Asp Glu Ser Arg Lys Leu Asn Arg
Ile 165 170 175Tyr Lys Phe His Gly Lys Ala Trp Asp Trp Pro Val Asp
Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr Leu Met Phe Ala Asp Leu
Asp Met Asp His Pro 195 200 205Glu Val Val Thr Glu Leu Lys Asn Trp
Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr Asn Ile Asp Gly Phe Arg
Leu Asp Ala Val Lys His Ile Lys Tyr225 230 235 240Ser Phe Phe Pro
Asp Trp Leu Thr Tyr Val Arg Asn Gln Thr Gly Lys 245 250 255Asn Leu
Phe Ala Val Gly Glu Phe Trp Ser Tyr Asp Val Asn Lys Leu 260 265
270His Asn Tyr Ile Thr Lys Thr Asn Gly Ser Met Ser Leu Phe Asp Ala
275 280 285Pro Leu His Asn Asn Phe Tyr Thr Ala Ser Lys Ser Ser Gly
Tyr Phe 290 295 300Asp Met Arg Tyr Leu Leu Asn Asn Thr Leu Met Lys
Asp Gln Pro Ser305 310 315 320Leu Ala Val Thr Leu Val Asp Asn His
Asp Thr Gln Pro Gly Gln Ser 325 330 335Leu Gln Ser Trp Val Glu Pro
Trp Phe Lys Pro Leu Ala Tyr Ala Phe 340 345 350Ile Leu Thr Arg Gln
Glu Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr 355 360 365Tyr Gly Ile
Pro Lys Tyr Asn Ile Pro Gly Leu Lys Ser Lys Ile Asp 370 375 380Pro
Leu Leu Ile Ala Arg Arg Asp Tyr Ala Tyr Gly Thr Gln Arg Asp385 390
395 400Tyr Ile Asp His Gln Asp Ile Ile Gly Trp Thr Arg Glu Gly Ile
Asp 405 410 415Thr Lys Pro Asn Ser Gly Leu Ala Ala Leu Ile Thr Asp
Gly Pro Gly 420 425 430Gly Ser Lys Trp Met Tyr Val Gly Lys Lys His
Ala Gly Lys Val Phe 435 440 445Tyr Asp Leu Thr Gly Asn Arg Ser Asp
Thr Val Thr Ile Asn Ala Asp 450 455 460Gly Trp Gly Glu Phe Lys Val
Asn Arg Gly Ser Val Ser Ile Trp Val465 470 475 480Ala
Lys42482PRTArtificial SequenceSynthetic amino acid sequence of
BASE-V31 42Asn Thr Ala Pro Ile Asn Glu Thr Met Met Gln Tyr Phe Glu
Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu Trp Thr Lys Val Lys Asn
Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly Ile Thr Ala Leu Trp Leu
Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln Ser Asp Val Gly Tyr Gly
Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu Phe Asn Gln Lys Gly Thr
Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys Thr Gln Tyr Ile Gln Ala
Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met Gln Val Tyr Ala Asp Val
Val Phe Asn His Lys Ala Gly Ala Asp 100 105 110Gly Thr Glu Phe Val
Asp Ala Val Glu Val Asp Pro Ser Asn Arg Tyr 115 120 125Gln Glu Thr
Ser Gly Glu Tyr Gln Ile Gln Ala Trp Thr Lys Phe Asp 130 135 140Phe
Pro Gly Arg Gly Asn Thr Tyr Ser Ser Phe Lys Trp Arg Trp Tyr145 150
155 160His Phe Asp Gly Thr Asp Trp Asp Glu Ser Arg Lys Leu Asn Arg
Ile 165 170 175Tyr Lys Phe His Gly Lys Ala Trp Asp Trp Pro Val Asp
Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr Leu Met Phe Ala Asp Leu
Asp Met Asp His Pro 195 200 205Glu Val Val Thr Glu Leu Lys Asn Trp
Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr Asn Ile Asp Gly Phe Arg
Leu Asp Ala
Val Lys His Ile Lys Tyr225 230 235 240Ser Phe Phe Pro Asp Trp Leu
Thr Tyr Val Arg Asn Gln Thr Gly Lys 245 250 255Asn Leu Phe Ala Val
Gly Glu Phe Trp Ser Tyr Asp Val Asn Lys Leu 260 265 270His Asn Tyr
Ile Thr Lys Thr Asn Gly Ser Met Ser Leu Phe Asp Ala 275 280 285Pro
Leu His Asn Asn Phe Tyr Thr Ala Ser Lys Ser Ser Gly Tyr Phe 290 295
300Asp Met Arg Tyr Leu Leu Asn Asn Thr Leu Met Lys Asp Gln Pro
Ser305 310 315 320Leu Ala Val Thr Leu Val Asp Asn His Asp Thr Gln
Pro Gly Gln Ser 325 330 335Leu Gln Ser Trp Val Glu Pro Trp Phe Lys
Pro Leu Ala Tyr Ala Phe 340 345 350Ile Leu Thr Arg Gln Glu Gly Tyr
Pro Cys Val Phe Tyr Gly Asp Tyr 355 360 365Tyr Gly Ile Pro Lys Tyr
Asn Ile Pro Gly Leu Lys Ser Lys Ile Asp 370 375 380Pro Leu Leu Ile
Ala Arg Arg Asp Tyr Ala Tyr Gly Thr Gln Arg Asp385 390 395 400Tyr
Ile Asp His Gln Asp Ile Ile Gly Trp Thr Arg Glu Gly Ile Asp 405 410
415Thr Lys Pro Asn Ser Gly Leu Ala Ala Leu Ile Thr Asp Gly Pro Gly
420 425 430Gly Ser Lys Trp Met Tyr Val Gly Lys Lys His Ala Gly Lys
Val Phe 435 440 445Tyr Asp Leu Thr Gly Asn Arg Ser Asp Thr Val Thr
Ile Asn Ala Asp 450 455 460Gly Trp Gly Glu Phe Lys Val Asn Arg Gly
Ser Val Ser Ile Trp Val465 470 475 480Ala Lys43482PRTArtificial
SequenceSynthetic amino acid sequence of BASE-V32 43Asn Thr Ala Pro
Ile Asn Glu Thr Met Met Gln Tyr Phe Glu Trp Asp1 5 10 15Leu Pro Asn
Asp Gly Thr Leu Trp Thr Lys Val Lys Asn Glu Ala Ala 20 25 30Asn Leu
Ser Ser Leu Gly Ile Thr Ala Leu Trp Leu Pro Pro Ala Tyr 35 40 45Lys
Gly Thr Ser Gln Ser Asp Val Gly Tyr Gly Val Tyr Asp Leu Tyr 50 55
60Asp Leu Gly Glu Phe Asn Gln Lys Gly Thr Ile Arg Thr Lys Tyr Gly65
70 75 80Thr Lys Thr Gln Tyr Ile Gln Ala Ile Gln Ala Ala Lys Ala Ala
Gly 85 90 95Met Gln Val Tyr Ala Asp Val Val Phe Asn His Lys Ala Gly
Ala Asp 100 105 110Gly Thr Glu Phe Val Asp Ala Val Glu Val Asp Pro
Ser Asn Arg Tyr 115 120 125Gln Glu Thr Ser Gly Thr Tyr Gln Ile Gln
Ala Trp Thr Lys Phe Asp 130 135 140Phe Pro Gly Arg Gly Asn Thr Tyr
Ser Ser Trp Lys Trp Arg Trp Tyr145 150 155 160His Phe Asp Gly Thr
Asp Trp Asp Glu Ser Arg Lys Leu Asn Arg Ile 165 170 175Tyr Lys Phe
Thr Gly Lys Ala Trp Asp Trp Pro Val Asp Thr Glu Asn 180 185 190Gly
Asn Tyr Asp Tyr Leu Met Phe Ala Asp Leu Asp Met Asp His Pro 195 200
205Glu Val Val Thr Glu Leu Lys Asn Trp Gly Thr Trp Tyr Val Asn Thr
210 215 220Thr Asn Ile Asp Gly Phe Arg Leu Asp Ala Val Lys His Ile
Lys Tyr225 230 235 240Ser Phe Phe Pro Asp Trp Leu Thr Tyr Val Arg
Asn Gln Thr Gly Lys 245 250 255Asn Leu Phe Ala Val Gly Glu Phe Trp
Ser Tyr Asp Val Asn Lys Leu 260 265 270His Asn Tyr Ile Thr Lys Thr
Asn Gly Ser Met Ser Leu Phe Asp Ala 275 280 285Pro Leu His Asn Asn
Phe Tyr Thr Ala Ser Lys Ser Ser Gly Tyr Phe 290 295 300Asp Met Arg
Tyr Leu Leu Asn Asn Thr Leu Met Lys Asp Gln Pro Ser305 310 315
320Leu Ala Val Thr Leu Val Asp Asn His Asp Thr Gln Pro Gly Gln Ser
325 330 335Leu Gln Ser Trp Val Glu Pro Trp Phe Lys Pro Leu Ala Tyr
Ala Phe 340 345 350Ile Leu Thr Arg Gln Glu Gly Tyr Pro Cys Val Phe
Tyr Gly Asp Tyr 355 360 365Tyr Gly Ile Pro Lys Tyr Asn Ile Pro Gly
Leu Lys Ser Lys Ile Asp 370 375 380Pro Leu Leu Ile Ala Arg Arg Asp
Tyr Ala Tyr Gly Thr Gln Arg Asp385 390 395 400Tyr Ile Asp His Gln
Asp Ile Ile Gly Trp Thr Arg Glu Gly Ile Asp 405 410 415Thr Lys Pro
Asn Ser Gly Leu Ala Ala Leu Ile Thr Asp Gly Pro Gly 420 425 430Gly
Ser Lys Trp Met Tyr Val Gly Lys Lys His Ala Gly Lys Val Phe 435 440
445Tyr Asp Leu Thr Gly Asn Arg Ser Asp Thr Val Thr Ile Asn Ala Asp
450 455 460Gly Trp Gly Glu Phe Lys Val Asn Arg Gly Ser Val Ser Ile
Trp Val465 470 475 480Ala Lys44482PRTArtificial SequenceSynthetic
amino acid sequence of BASE-V33 44Asn Thr Ala Pro Ile Asn Glu Thr
Met Met Gln Tyr Phe Glu Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu
Trp Thr Lys Val Lys Asn Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly
Ile Thr Ala Leu Trp Leu Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln
Ser Asp Val Gly Tyr Gly Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu
Phe Asn Gln Lys Gly Thr Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys
Thr Gln Tyr Ile Gln Ala Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met
Gln Val Tyr Ala Asp Val Val Phe Asn His Lys Ala Gly Ala Asp 100 105
110Gly Thr Glu Phe Val Asp Ala Val Glu Val Asp Pro Ser Asn Arg Asn
115 120 125Gln Glu Thr Ser Gly Glu Tyr Gln Ile Gln Ala Trp Thr Lys
Phe Asp 130 135 140Phe Pro Gly Arg Gly Asn Thr Tyr Ser Ser Trp Lys
Trp Arg Trp Tyr145 150 155 160His Phe Asp Gly Thr Asp Trp Asp Glu
Ser Arg Lys Leu Asn Arg Ile 165 170 175Tyr Lys Phe His Gly Lys Ala
Trp Asp Trp Pro Val Asp Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr
Leu Met Phe Ala Asp Leu Asp Met Asp His Pro 195 200 205Glu Val Val
Thr Glu Leu Lys Asn Trp Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr
Asn Ile Asp Gly Phe Arg Leu Asp Ala Val Lys His Ile Lys Tyr225 230
235 240Ser Phe Phe Pro Asp Trp Leu Thr Tyr Val Arg Asn Gln Thr Gly
Lys 245 250 255Asn Leu Phe Ala Val Gly Glu Phe Trp Ser Tyr Asp Val
Asn Lys Leu 260 265 270His Asn Tyr Ile Thr Lys Thr Asn Gly Ser Met
Ser Leu Phe Asp Ala 275 280 285Pro Leu His Asn Asn Phe Tyr Thr Ala
Ser Lys Ser Ser Gly Tyr Phe 290 295 300Asp Met Arg Tyr Leu Leu Asn
Asn Thr Leu Met Lys Asp Gln Pro Ser305 310 315 320Leu Ala Val Thr
Leu Val Asp Asn His Asp Thr Gln Pro Gly Gln Ser 325 330 335Leu Gln
Ser Trp Val Glu Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe 340 345
350Ile Leu Thr Arg Gln Glu Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr
355 360 365Tyr Gly Ile Pro Lys Tyr Asn Ile Pro Gly Leu Lys Ser Lys
Ile Asp 370 375 380Pro Leu Leu Ile Ala Arg Arg Asp Tyr Ala Tyr Gly
Thr Gln Arg Asp385 390 395 400Tyr Ile Asp His Gln Asp Ile Ile Gly
Trp Thr Arg Glu Gly Ile Asp 405 410 415Thr Lys Pro Asn Ser Gly Leu
Ala Ala Leu Ile Thr Asp Gly Pro Gly 420 425 430Gly Ser Lys Trp Met
Tyr Val Gly Lys Lys His Ala Gly Lys Val Phe 435 440 445Tyr Asp Leu
Thr Gly Asn Arg Ser Asp Thr Val Thr Ile Asn Ala Asp 450 455 460Gly
Trp Gly Glu Phe Lys Val Asn Arg Gly Ser Val Ser Ile Trp Val465 470
475 480Ala Lys45482PRTArtificial SequenceSynthetic amino acid
sequence of BASE-V34 45Asn Thr Ala Pro Ile Asn Glu Thr Met Met Gln
Tyr Phe Glu Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu Trp Thr Lys
Val Lys Asn Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly Ile Thr Ala
Leu Trp Leu Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln Ser Asp Val
Gly Tyr Gly Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu Phe Asn Gln
Lys Gly Thr Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys Thr Gln Tyr
Ile Gln Ala Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met Gln Val Tyr
Ala Asp Val Val Phe Asn His Lys Ala Gly Ala Asp 100 105 110Gly Thr
Glu Phe Val Asp Ala Val Glu Val Asp Pro Ser Asn Arg Tyr 115 120
125Gln Glu Thr Ser Gly Glu Tyr Gln Ile Gln Ala Trp Thr Lys Phe Asp
130 135 140Phe Pro Gly Arg Gly Asn Thr Tyr Ser Ser Trp Lys Trp Arg
Trp Tyr145 150 155 160His Phe Asp Gly Thr Asp Trp Asp Glu Ser Arg
Lys Leu Asn Arg Ile 165 170 175Tyr Lys Phe His Gly Lys Ala Trp Asp
Trp Pro Val Asp Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr Leu Met
Phe Ala Asp Leu Asp Met Asp His Pro 195 200 205Glu Val Val Thr Glu
Leu Lys Asn Trp Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr Asn Ile
Asp Gly Phe Arg Leu Asp Ala Val Lys His Ile Lys Tyr225 230 235
240Ser Phe Phe Pro Asp Trp Leu Thr Tyr Val Arg Asn Gln Thr Gly Lys
245 250 255Asn Leu Phe Ala Val Gly Glu Phe Trp Ser Tyr Asp Val Asn
Lys Leu 260 265 270His Asn Tyr Ile Thr Lys Thr Asn Gly Ser Met Ser
Leu Phe Asp Ala 275 280 285Pro Leu His Asn Asn Phe Tyr Thr Ala Ser
Lys Ser Ser Gly Tyr Phe 290 295 300Asp Met Arg Tyr Leu Leu Asn Asn
Thr Leu Met Lys Asp Gln Pro Ser305 310 315 320Leu Ala Val Thr Leu
Val Asp Asn His Asp Thr Gln Pro Gly Gln Ser 325 330 335Leu Gln Ser
Trp Val Glu Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe 340 345 350Ile
Leu Thr Arg Gln Glu Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr 355 360
365Tyr Gly Ile Pro Lys Tyr Asn Ile Pro Gly Leu Lys Ser Lys Ile Asp
370 375 380Pro Leu Leu Ile Ala Arg Arg Asp Tyr Ala Tyr Gly Thr Gln
Arg Asp385 390 395 400Tyr Ile Asp His Gln Asp Ile Ile Gly Trp Thr
Arg Glu Gly Ile Asp 405 410 415Thr Lys Pro Asn Ser Gly Leu Ala Ala
Leu Ile Thr Asp Gly Pro Gly 420 425 430Gly Ser Lys Trp Met Tyr Val
Gly Lys Lys His Ala Gly Lys Val Phe 435 440 445Tyr Asp Leu Thr Gly
Asn Arg Ser Asp Thr Val Thr Ile Asn Ala Asp 450 455 460Gly Trp Gly
Glu Phe Lys Val Asn Arg Gly Ser Val Ser Ile Trp Val465 470 475
480Ala Lys46482PRTArtificial SequenceSynthetic amino acid sequence
of BASE-V35 46Asn Thr Ala Pro Ile Asn Glu Thr Met Met Gln Tyr Phe
Glu Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu Trp Thr Lys Val Lys
Asn Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly Ile Thr Ala Leu Trp
Leu Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln Ser Asp Val Gly Tyr
Gly Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu Phe Asn Gln Lys Gly
Thr Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys Thr Gln Tyr Ile Gln
Ala Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met Gln Val Tyr Ala Asp
Val Val Phe Asn His Lys Ala Gly Ala Asp 100 105 110Gly Thr Glu Phe
Val Asp Ala Val Glu Val Asp Pro Ser Asn Arg Tyr 115 120 125Gln Glu
Thr Ser Gly His Tyr Gln Ile Gln Ala Trp Thr Lys Phe Asp 130 135
140Phe Pro Gly Arg Gly Asn Thr Tyr Ser Ser Trp Lys Trp Arg Trp
Tyr145 150 155 160His Phe Asp Gly Thr Asp Trp Asp Glu Ser Arg Lys
Leu Asn Arg Ile 165 170 175Tyr Lys Phe Asp Gly Lys Ala Trp Asp Trp
Pro Val Asp Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr Leu Met Phe
Ala Asp Leu Asp Met Asp His Pro 195 200 205Glu Val Val Thr Glu Leu
Lys Asn Trp Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr Asn Ile Asp
Gly Phe Arg Leu Asp Ala Val Lys His Ile Lys Tyr225 230 235 240Ser
Phe Phe Pro Asp Trp Leu Thr Tyr Val Arg Asn Gln Thr Gly Lys 245 250
255Asn Leu Phe Ala Val Gly Glu Phe Trp Ser Tyr Asp Val Asn Lys Leu
260 265 270His Asn Tyr Ile Thr Lys Thr Asn Gly Ser Met Ser Leu Phe
Asp Ala 275 280 285Pro Leu His Asn Asn Phe Tyr Thr Ala Ser Lys Ser
Ser Gly Tyr Phe 290 295 300Asp Met Arg Tyr Leu Leu Asn Asn Thr Leu
Met Lys Asp Gln Pro Ser305 310 315 320Leu Ala Val Thr Leu Val Asp
Asn His Asp Thr Gln Pro Gly Gln Ser 325 330 335Leu Gln Ser Trp Val
Glu Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe 340 345 350Ile Leu Thr
Arg Gln Glu Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr 355 360 365Tyr
Gly Ile Pro Lys Tyr Asn Ile Pro Gly Leu Lys Ser Lys Ile Asp 370 375
380Pro Leu Leu Ile Ala Arg Arg Asp Tyr Ala Tyr Gly Thr Gln Arg
Asp385 390 395 400Tyr Ile Asp His Gln Asp Ile Ile Gly Trp Thr Arg
Glu Gly Ile Asp 405 410 415Thr Lys Pro Asn Ser Gly Leu Ala Ala Leu
Ile Thr Asp Gly Pro Gly 420 425 430Gly Ser Lys Trp Met Tyr Val Gly
Lys Lys His Ala Gly Lys Val Phe 435 440 445Tyr Asp Leu Thr Gly Asn
Arg Ser Asp Thr Val Thr Ile Asn Ala Asp 450 455 460Gly Trp Gly Glu
Phe Lys Val Asn Arg Gly Ser Val Ser Ile Trp Val465 470 475 480Ala
Lys47482PRTArtificial SequenceSynthetic amino acid sequence of
BASE-V36 47Asn Thr Ala Pro Ile Asn Glu Thr Met Met Gln Tyr Phe Glu
Trp Asp1 5 10 15Leu Pro Asn Asp Gly Thr Leu Trp Thr Lys Val Lys Asn
Glu Ala Ala 20 25 30Asn Leu Ser Ser Leu Gly Ile Thr Ala Leu Trp Leu
Pro Pro Ala Tyr 35 40 45Lys Gly Thr Ser Gln Ser Asp Val Gly Tyr Gly
Val Tyr Asp Leu Tyr 50 55 60Asp Leu Gly Glu Phe Asn Gln Lys Gly Thr
Ile Arg Thr Lys Tyr Gly65 70 75 80Thr Lys Thr Gln Tyr Ile Gln Ala
Ile Gln Ala Ala Lys Ala Ala Gly 85 90 95Met Gln Val Tyr Ala Asp Val
Val Phe Asn His Lys Ala Gly Ala Asp 100 105 110Gly Thr Glu Phe Val
Asp Ala Val Glu Val Asp Pro Ser Asn Arg Tyr 115 120 125Gln Glu Thr
Ser Gly Glu Tyr Gln Ile Gln Ala Trp Thr Lys Phe Asp 130 135 140Phe
Pro Gly Arg Gly Asn Thr Tyr Ser Ser Trp Lys Trp Arg Trp Tyr145 150
155 160His Phe Asp Gly Thr Asp Trp Asp Glu Ser Arg Lys Leu Asn Arg
Ile 165 170 175Tyr Lys Phe Gly Gly Lys Ala Trp Asp Trp Pro Val Asp
Thr Glu Asn 180 185 190Gly Asn Tyr Asp Tyr Leu Met Phe Ala Asp Leu
Asp Met Asp His Pro 195 200 205Glu Val Val Thr Glu Leu Lys Asn Trp
Gly Thr Trp Tyr Val Asn Thr 210 215 220Thr Asn Ile Asp Gly Phe Arg
Leu Asp Ala Val Lys His Ile Lys Tyr225 230 235 240Ser Phe Phe Pro
Asp Trp Leu Thr Tyr Val Arg Asn Gln Thr Gly Lys
245 250 255Asn Leu Phe Ala Val Gly Glu Phe Trp Ser Tyr Asp Val Asn
Lys Leu 260 265 270His Asn Tyr Ile Thr Lys Thr Asn Gly Ser Met Ser
Leu Phe Asp Ala 275 280 285Pro Leu His Asn Asn Phe Tyr Thr Ala Ser
Lys Ser Ser Gly Tyr Phe 290 295 300Asp Met Arg Tyr Leu Leu Asn Asn
Thr Leu Met Lys Asp Gln Pro Ser305 310 315 320Leu Ala Val Thr Leu
Val Asp Asn His Asp Thr Gln Pro Gly Gln Ser 325 330 335Leu Gln Ser
Trp Val Glu Pro Trp Phe Lys Pro Leu Ala Tyr Ala Phe 340 345 350Ile
Leu Thr Arg Gln Glu Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr 355 360
365Tyr Gly Ile Pro Lys Tyr Asn Ile Pro Gly Leu Lys Ser Lys Ile Asp
370 375 380Pro Leu Leu Ile Ala Arg Arg Asp Tyr Ala Tyr Gly Thr Gln
Arg Asp385 390 395 400Tyr Ile Asp His Gln Asp Ile Ile Gly Trp Thr
Arg Glu Gly Ile Asp 405 410 415Thr Lys Pro Asn Ser Gly Leu Ala Ala
Leu Ile Thr Asp Gly Pro Gly 420 425 430Gly Ser Lys Trp Met Tyr Val
Gly Lys Lys His Ala Gly Lys Val Phe 435 440 445Tyr Asp Leu Thr Gly
Asn Arg Ser Asp Thr Val Thr Ile Asn Ala Asp 450 455 460Gly Trp Gly
Glu Phe Lys Val Asn Arg Gly Ser Val Ser Ile Trp Val465 470 475
480Ala Lys484PRTArtificial SequenceSynthetic conserved amino acid
sequence motifmisc_feature(1)..(1)Xaa can be any naturally
occurring amino acidmisc_feature(2)..(2)Xaa is Gly or
Sermisc_feature(3)..(3)Xaa can be any naturally occurring amino
acid 48Xaa Xaa Xaa Gly1494PRTArtificial SequenceSynthetic peptide
49Arg Gly Thr Gly150478PRTArtificial SequenceSynthetic mature form
of PcuAmy1-v3 50Ala Asp Asn Gly Thr Ile Met Gln Tyr Phe Glu Trp Tyr
Leu Pro Asn1 5 10 15Asp Gly Ala His Trp Asn Arg Leu Asn Asn Asp Ala
Gln Asn Leu Lys 20 25 30Asn Val Gly Ile Thr Ala Val Trp Ile Pro Pro
Ala Tyr Lys Gly Gly 35 40 45Ser Ser Ala Asp Val Gly Tyr Gly Val Tyr
Asp Thr Tyr Asp Leu Gly 50 55 60Glu Phe Asn Gln Lys Gly Thr Val Arg
Thr Lys Tyr Gly Thr Lys Ser65 70 75 80Glu Leu Ile Ser Ala Val Asn
Asn Leu His Ala Lys Gly Ile Ala Val 85 90 95Tyr Gly Asp Val Val Leu
Asn His Arg Met Asn Ala Asp Ala Thr Glu 100 105 110Leu Val Asp Ala
Val Glu Val Asp Pro Asn Asn Arg Asn Val Glu Thr 115 120 125Thr Ser
Thr Tyr Gln Ile Gln Ala Trp Thr Gln Tyr Asp Phe Pro Gly 130 135
140Arg Gly Asn Thr Tyr Ser Ser Phe Lys Trp Arg Trp Tyr His Phe
Asp145 150 155 160Gly Val Asp Trp Asp Gln Ser Arg Gly Leu Asn Arg
Ile Tyr Lys Leu 165 170 175Asp Gly Lys Asp Trp Asp Trp Pro Val Asp
Ser Glu Tyr Gly Asn Tyr 180 185 190Asp Tyr Leu Met Gly Ala Asp Leu
Asp Phe Asn His Pro Asp Val Val 195 200 205Asn Glu Thr Lys Thr Trp
Gly Lys Trp Phe Val Asn Thr Val Asn Leu 210 215 220Asp Gly Val Arg
Leu Asp Ala Val Lys His Ile Lys Phe Asp Phe Met225 230 235 240Arg
Asp Trp Val Asn Asn Val Arg Ser Thr Thr Gly Lys Asn Leu Phe 245 250
255Ala Val Gly Glu Tyr Trp His Tyr Asp Val Asn Lys Leu Asn Ser Tyr
260 265 270Ile Thr Lys Thr Asn Gly Thr Met Ser Leu Phe Asp Val Pro
Leu His 275 280 285Phe Arg Phe Tyr Asp Ala Ser Asn Gly Gly Gly Gly
Tyr Asp Met Arg 290 295 300Asn Leu Leu Asn Asn Thr Leu Met Ser Ser
Asn Pro Met Lys Ala Val305 310 315 320Thr Phe Val Glu Asn His Asp
Thr Gln Pro Thr Gln Ala Leu Gln Ser 325 330 335Thr Val Gln Ser Trp
Phe Lys Pro Leu Ala Tyr Ala Thr Ile Leu Thr 340 345 350Arg Glu Gln
Gly Tyr Pro Cys Val Phe Tyr Gly Asp Tyr Tyr Gly Thr 355 360 365Ser
Asp Gly Lys Ile Ser Ser Tyr Lys Pro Ile Met Asp Lys Leu Leu 370 375
380Asn Ala Arg Lys Val Tyr Ala Tyr Gly Thr Gln Arg Asp Tyr Phe
Asp385 390 395 400His Pro Asp Ile Val Gly Trp Thr Arg Glu Gly Asp
Ala Ala His Ala 405 410 415Gly Ser Gly Leu Ala Thr Leu Ile Thr Asp
Gly Pro Gly Gly Ser Lys 420 425 430Trp Met Tyr Val Gly Thr Ser Lys
Ala Gly Gln Val Trp Thr Asp Lys 435 440 445Thr Gly Asn Arg Ser Gly
Thr Val Thr Ile Asp Ala Asn Gly Trp Gly 450 455 460Asn Phe Trp Val
Asn Lys Gly Ser Val Ser Val Trp Ala Lys465 470 475
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020
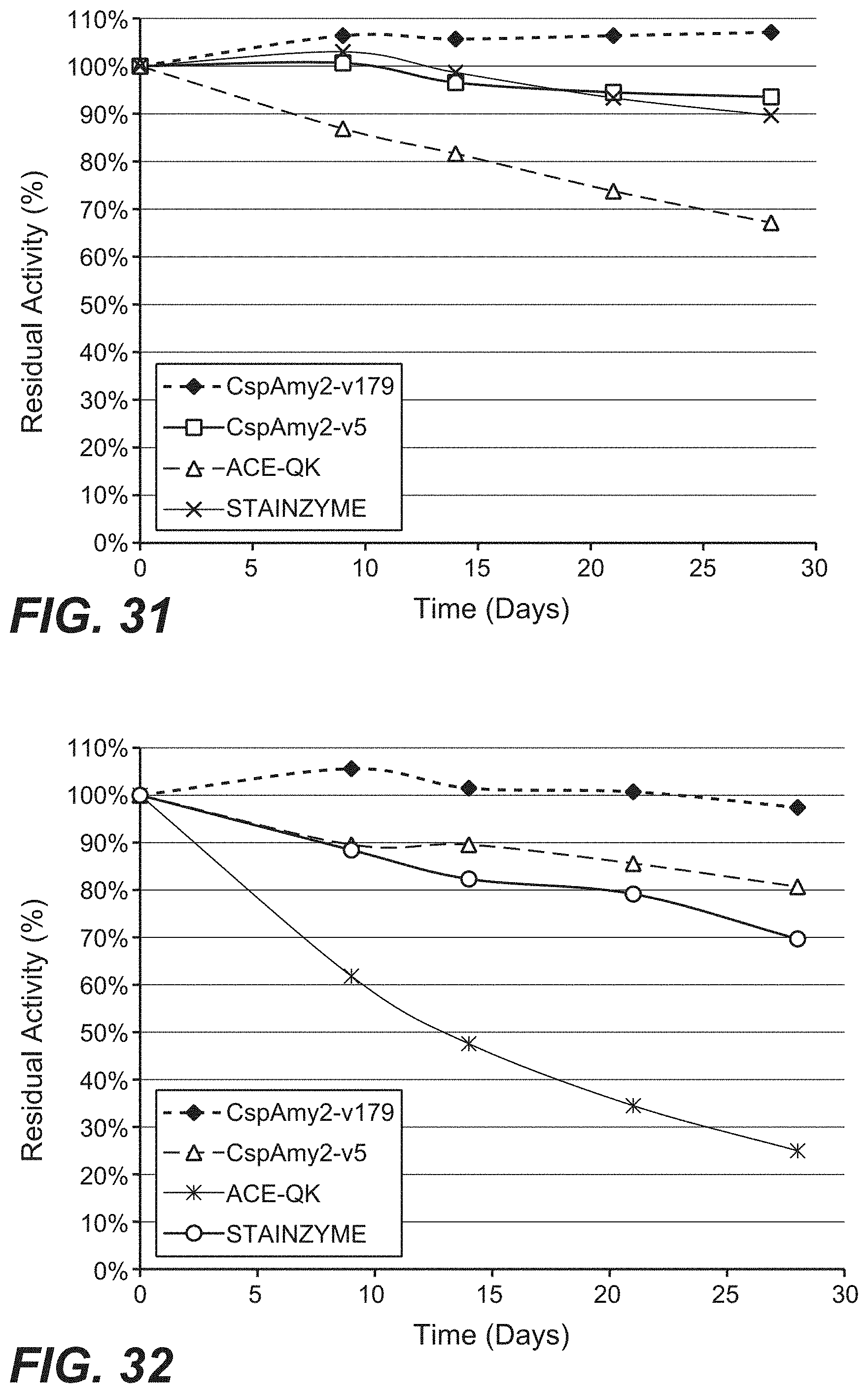
D00021

D00022

D00023

D00024

D00025

D00026

D00027

D00028

D00029

D00030

D00031

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.