Pneumococcal Fusion Protein Vaccines
Malley; Richard ; et al.
U.S. patent application number 16/568646 was filed with the patent office on 2020-03-19 for pneumococcal fusion protein vaccines. This patent application is currently assigned to THE CHILDREN'S MEDICAL CENTER CORPORATION. The applicant listed for this patent is THE CHILDREN'S MEDICAL CENTER CORPORATION. Invention is credited to Yingjie Lu, Richard Malley, Fan ZHANG.
| Application Number | 20200087361 16/568646 |
| Document ID | / |
| Family ID | 69772826 |
| Filed Date | 2020-03-19 |







View All Diagrams
| United States Patent Application | 20200087361 |
| Kind Code | A1 |
| Malley; Richard ; et al. | March 19, 2020 |
PNEUMOCOCCAL FUSION PROTEIN VACCINES
Abstract
Technologies for the prevention and/or treatment of pneumococcal infections.
| Inventors: | Malley; Richard; (Beverly, MA) ; Lu; Yingjie; (West Roxbury, MA) ; ZHANG; Fan; (West Roxbury, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | THE CHILDREN'S MEDICAL CENTER
CORPORATION Boston MA |
||||||||||
| Family ID: | 69772826 | ||||||||||
| Appl. No.: | 16/568646 | ||||||||||
| Filed: | September 12, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 62730199 | Sep 12, 2018 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2319/20 20130101; C07K 14/3156 20130101; C07K 14/47 20130101; C12N 15/62 20130101; C07K 2319/00 20130101; A61K 39/092 20130101 |
| International Class: | C07K 14/47 20060101 C07K014/47; C07K 14/315 20060101 C07K014/315 |
Claims
1. A fusion protein comprising: (i) a biotin-binding moiety; (ii) an SP1500 polypeptide or fragment; and (iii) an SP0785 polypeptide or fragment.
2. A fusion protein comprising: (i) a biotin-binding moiety comprising an amino acid sequence at least 80%, 85%, 90% identical to SEQ ID NO:2 or a biotin binding portion thereof; (ii) a polypeptide comprising an amino acid sequence at least 80%, 85%, 90% identical to SEQ ID NO:7 or an antigenic portion thereof; and (iii) a polypeptide comprising an amino acid sequence at least 80%, 85%, 90% identical to SEQ ID NO:4 or an antigenic portion thereof.
3. A fusion protein comprising: (i) a biotin-binding moiety comprising an amino acid sequence at least 80%, 85%, 90% identical to SEQ ID NO:2 or a biotin binding portion thereof; (ii) a first polypeptide comprising an amino acid sequence at least 80%, 85%, 90% identical to SEQ ID NO:7 or an antigenic portion thereof; (iii) a second polypeptide comprising an amino acid sequence at least 80%, 85%, 90% identical to SEQ ID NO:4 or an antigenic portion thereof, and (iv) a first linker positioned between the biotin-binding moiety and the first polypeptide and/or a second linker positioned between the first polypeptide and the second polypeptide.
4. The fusion protein of claim 3, wherein the first linker comprises the amino acid sequence GGGGSSS (SEQ ID NO: 37).
5. The fusion protein of claim 3 or claim 4, wherein the second linker comprises the amino acid sequence AAA (SEQ ID NO: 38).
6. The fusion protein of any one of claims 1-5, wherein the biotin-binding moiety is C-terminally linked to the first polypeptide.
7. The fusion protein of any one of claims 1-5, wherein the biotin-binding moiety is N-terminally linked to the first polypeptide.
8. The fusion protein of any one of claims 1-7, wherein the first polypeptide is C-terminally linked to the second polypeptide.
9. The fusion protein of any one of claims 1-7, wherein the first polypeptide is N-terminally linked to the second polypeptide.
10. The fusion protein of any one of claims 1-9, wherein the second polypeptide is C-terminally linked to the biotin-binding moiety.
11. The fusion protein of any one of claims 1-9, wherein the second polypeptide is N-terminally linked to the biotin-binding moiety.
12. The fusion protein of any one of claims 1-11, comprising the amino acid sequence of SEQ ID NO:2.
13. The fusion protein of any one of claims 1-12, comprising the amino acid sequence of SEQ ID NO:7.
14. The fusion protein of any one of claims 1-13, comprising the amino acid sequence of SEQ ID NO:4.
15. The fusion protein of any one of claims 1-14, comprising the amino acid sequence at least 80%, 85%, 90% identical to SEQ ID NO:23.
16. The fusion protein of any one of claims 1-15, wherein the fusion protein is CP1.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application claims the benefit of U.S. Provisional Application No. 62/730,199 filed Sep. 12, 2018, the contents of which are hereby incorporated herein in their entirety.
SEQUENCE LISTING
[0002] The instant application contains a Sequence Listing which has been submitted electronically in ASCII format and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Nov. 15, 2019, is named 701039-093500USPT_SL.txt and is 295,804 bytes in size.
BACKGROUND
[0003] Streptococcus pneumoniae remains a leading cause of serious illness, including bacteremia, sepsis, meningitis and pneumonia, among children and adults worldwide. Morbidity and mortality among infants, young children, the elderly and subjects who have certain underlying medical conditions is high.
[0004] S. pneumoniae is a Gram-positive encapsulated coccus that colonizes the nasopharynx in about 5-10% of healthy adults and 20-40% of healthy children. Normal colonization becomes infectious when S. pneumoniae is carried into the Eustachian tubes, nasal sinuses, lungs, bloodstream, meninges, joint spaces, bones and peritoneal cavity. S. pneumoniae infection is the most frequent cause of bacteremia, pneumonia, meningitis, sinusitis and acute otitis media [CDC, 2010].
[0005] Pneumococcal disease can be invasive or noninvasive. The most common form of noninvasive disease, non-bacteremic pneumococcal pneumonia, remains one of the most frequent causes for pneumonia hospitalizations. Invasive pneumococcal disease (IPD) is defined as S. pneumoniae isolated from a normally sterile site (e.g., cerebrospinal fluid, blood, joint fluid, pleural fluid or peritoneal fluid). The highest incidence of IPD is found at the extremes of age--in elderly adults and in young children younger than 2 years of age. In the U.S., prior to advent of the first pneumococcal vaccine, S. pneumoniae caused approximately 17,000 cases of invasive disease each year among children younger than 5 years of age, including 700 cases of meningitis and 200 deaths [CDC, 2000]. The highest morbidity and mortality rates have been reported in developing countries, but the disease burden is also considerable in industrialized countries.
[0006] S. pneumoniae has several virulence factors that enable the organism to evade the immune system. Examples include a polysaccharide capsule that prevents phagocytosis by host immune cells, proteases that inhibit complement-mediated opsonization, and proteins that cause lysis of host cells. In the polysaccharide capsule, the presence of complex polysaccharides forms the basis for dividing pneumococci into different serotypes. To date, close to 100 serotypes of S. pneumoniae have been identified.
[0007] Two vaccines for S. pneumoniae are currently available in the U.S.: Pneumococcal Conjugate Vaccine (PCV13 or Prevnar 13.RTM.) and Pneumococcal Polysaccharide Vaccine (PPSV23 or Pneumovax.RTM.). PCV13 cannot confer protection against most of the known serotypes of S. pneumoniae. While PPSV23 includes polysaccharide components of more serotypes of S. pneumoniae than PCV13, it induces an immune response that is neither long-lasting nor anamnestic upon subsequent challenge. PPSV23 protects adults and the elderly against invasive pneumococcal disease; however, no consistent effect has been observed in the prevention of pneumonia [Gruber et al, 2008].
[0008] Thus, there is a medical need for a vaccine that provides T-cell dependent immunity against a broad range of serotypes of S. pneumoniae.
SUMMARY
[0009] The present disclosure addresses the lack of suitable technologies for the prevention and/or treatment of pneumococcal infection. Among other things, the present disclosure addresses challenges in providing vaccines with sufficient immunogenicity to protect against invasive pneumococcal disease and pneumonia. Technologies described herein can induce a T- and B-cell response and/or provide immunity against a broad range of S. pneumoniae serotypes, including one or more serotypes not included in commercially-available vaccines, e.g., PCV13 or PPSV23.
[0010] In some embodiments, a fusion protein described herein, when administered to a subject, can induce a higher Th17 response by at least 25% or more including, e.g., at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 98%, or more, as compared to that induced by individual antigenic components of the fusion protein. In some embodiments, a fusion protein described herein, when administered to a subject, can induce a higher Th17 response by at least 1.1-fold or more, including, e.g., at least 1.2-fold, at least 1.3-fold, at least 1.4-fold, at least 1.5-fold, at least 2-fold, at least 3-fold, at least 4-fold, at least 5-fold, at least 10-fold, or higher, as compared to that induced by individual antigenic components of the fusion protein.
[0011] In some embodiments, a fusion protein described herein, when administered to a subject, can induce an immune response to one or more representative non-vaccine pneumococcal serotype(s) that is/are not included in a commercially-available vaccine, e.g., PCV13 or PPSV23. In some embodiments, a fusion protein described herein, when administered to a subject, can induce an immune response to one or more non-vaccine pneumococcal serotypes selected from the group consisting of 6B, 16F, 15A, and 35B.
BRIEF DESCRIPTION OF THE DRAWINGS
[0012] The present teachings described herein will be more fully understood from the following description of various illustrative embodiments, when read together with the accompanying drawings. It should be understood that the drawings described below are for illustration purposes only and are not intended to limit the scope of the present teachings in any way.
[0013] FIG. 1 is a schematic of an exemplary CP1 fusion protein. Such an exemplary CP1 fusion protein comprises a biotin-binding protein such as, e.g., a truncated rhizavidin protein (e.g., amino acids 45-179 of a wild-type rhizavidin protein (denoted Rhavi)), a first linker (e.g., a GGGGSSS linker (SEQ ID NO: 37)), a SP1500 polypeptide (e.g., amino acids 27-278 of S. pneumoniae protein SP1500), a second linker (e.g., the amino acid sequence or linker AAA (SEQ ID NO: 38)), and a SP0785 polypeptide (e.g., amino acids 33-399 of S. pneumoniae protein SP0785). In some embodiments, a CP1 fusion protein may further comprise a His tag. For a GGGGSSS linker (SEQ ID NO: 37), the SSS amino acid sequence can be from the Sac I site on a PET21/24b plasmid, with the GGGG amino acid sequence (SEQ ID NO: 61) added to create a flexible linker with minimal steric hindrance. Alternatively, the GGGGSSS linker (SEQ ID NO: 37) can be synthesized. The AAA amino acid sequence (SEQ ID NO: 38) can be from the Not I site on a PET21/24b plasmid, or synthesized.
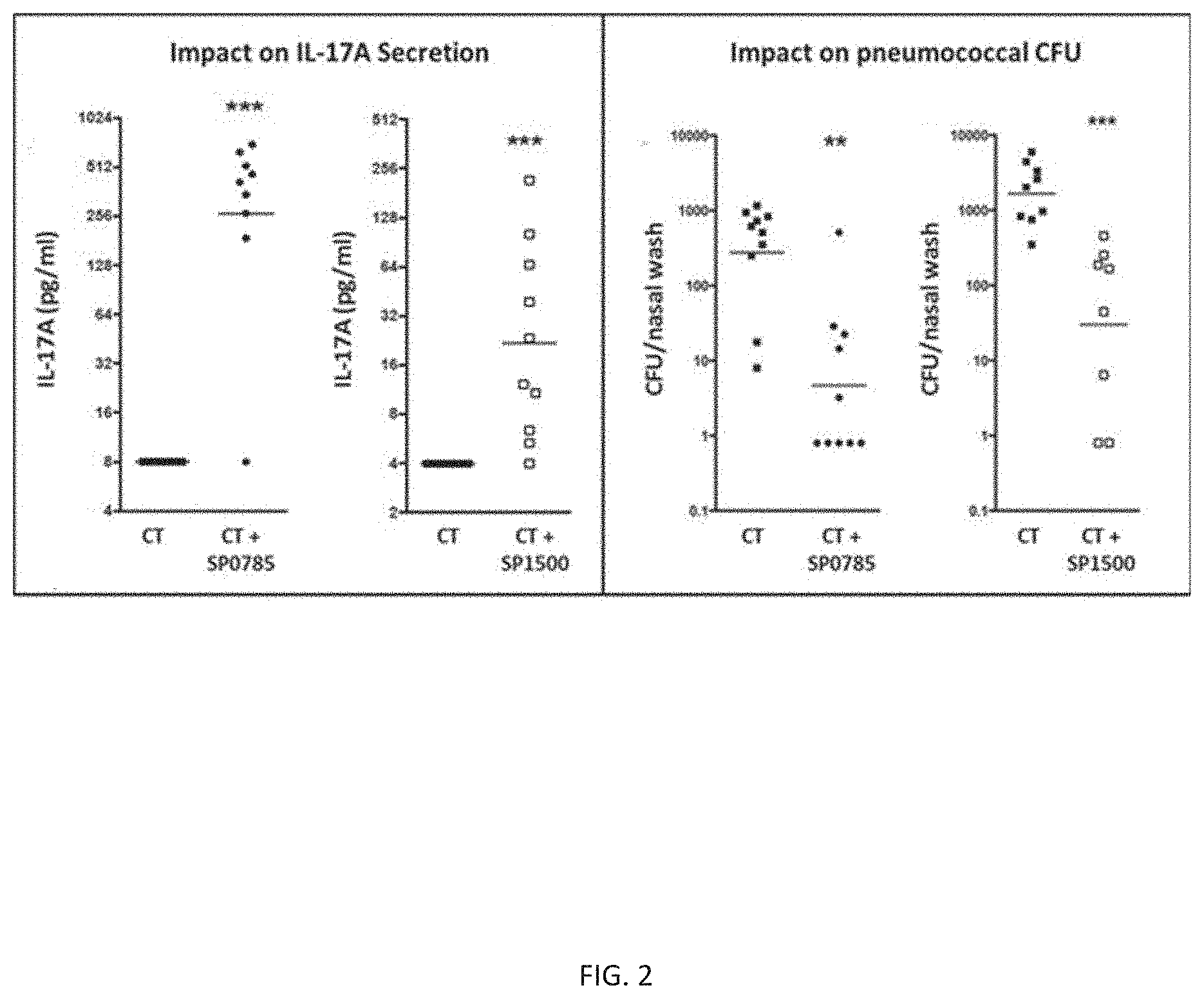
[0014] FIG. 2 illustrates immune responses to SP1500 and SP0785 polypeptide components of an exemplary fusion protein CP1. The left panel shows Th17 response to a SP1500 polypeptide and a SP0785 polypeptide of an exemplary fusion protein CP1. Th17 responses are shown as geometric mean concentration of IL-17A secreted in media after stimulation of peripheral blood samples of mice immunized with cholera toxin alone, or with a SP1500 or SP0785 polypeptide adjuvanted with cholera toxin. Each point on the graph represents secreted IL-17A for one mouse. The right panel shows protection from colonization by S. pneumoniae following immunization with cholera toxin alone, or with a SP1500 or SP0785 polypeptide adjuvanted with cholera toxin and intranasal challenge with S. pneumoniae. Each point on the graph represents S. pneumoniae CFU per nasal wash for one mouse. The horizontal bars in both panels represent the geometric mean of secreted IL-17A (left panel) and geometric mean of CFU per nasal wash for each group (right panel), respectively. The data were statistically analyzed by Mann-Whitney U test. CT: cholera toxin; CFU: colony forming unit; ELISA: enzyme-linked immunosorbent assay; IL-17A: interleukin 17A. **p<0.01; ***p<0.001.
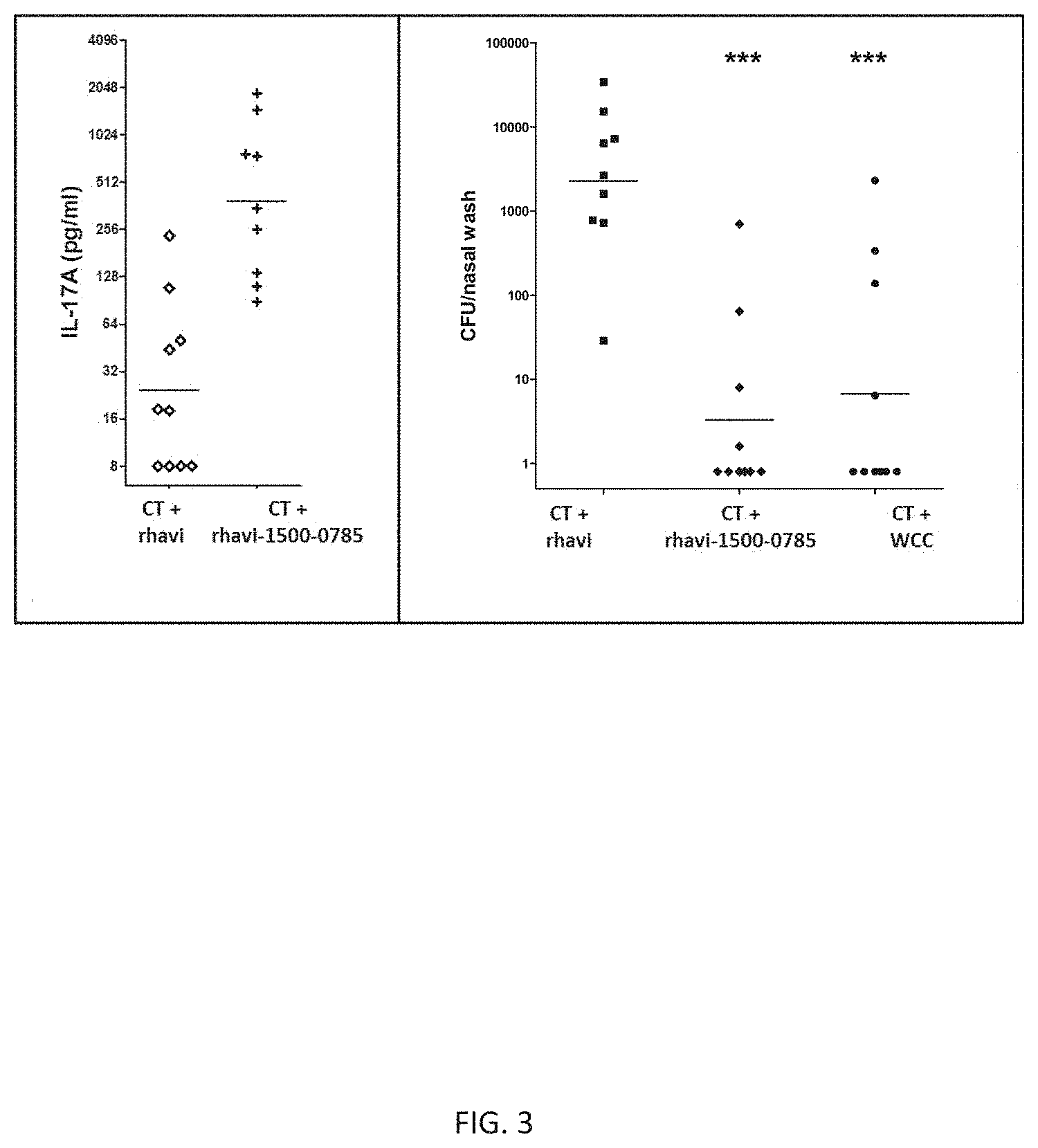
[0015] FIG. 3 illustrates immune responses to an exemplary fusion protein CP1. The left panel shows Th17 response to an exemplary fusion protein CP1. Th17 responses are shown as geometric mean concentration of IL-17 secreted in media after stimulation of peripheral blood samples of mice immunized with a truncated rhizavidin protein (Rhavi) or with CP1, both adjuvanted with cholera toxin. Each point on the graph represents secreted IL-17A for one mouse. The right panel shows protection from colonization by S. pneumoniae following immunization with a truncated rhizavidin protein (Rhavi), CP1, or killed (inactivated) pneumococcal whole cells (WCC), adjuvanted with cholera toxin, and intranasal challenge with S. pneumoniae. Each point on the graph represents S. pneumoniae CFU per nasal wash for one mouse. The horizontal bars in both panels represent the geometric mean of secreted IL-17A (left panel) and geometric mean of CFU per nasal wash for each group (right panel), respectively. The data were statistically analyzed by Mann-Whitney U test. CT: cholera toxin; CFU: colony forming unit; IL-17A: interleukin 17A; rhavi: truncated rhizavidin protein; (amino acids 45-179 of a full-length rhizavidin protein). ***p<0.001.
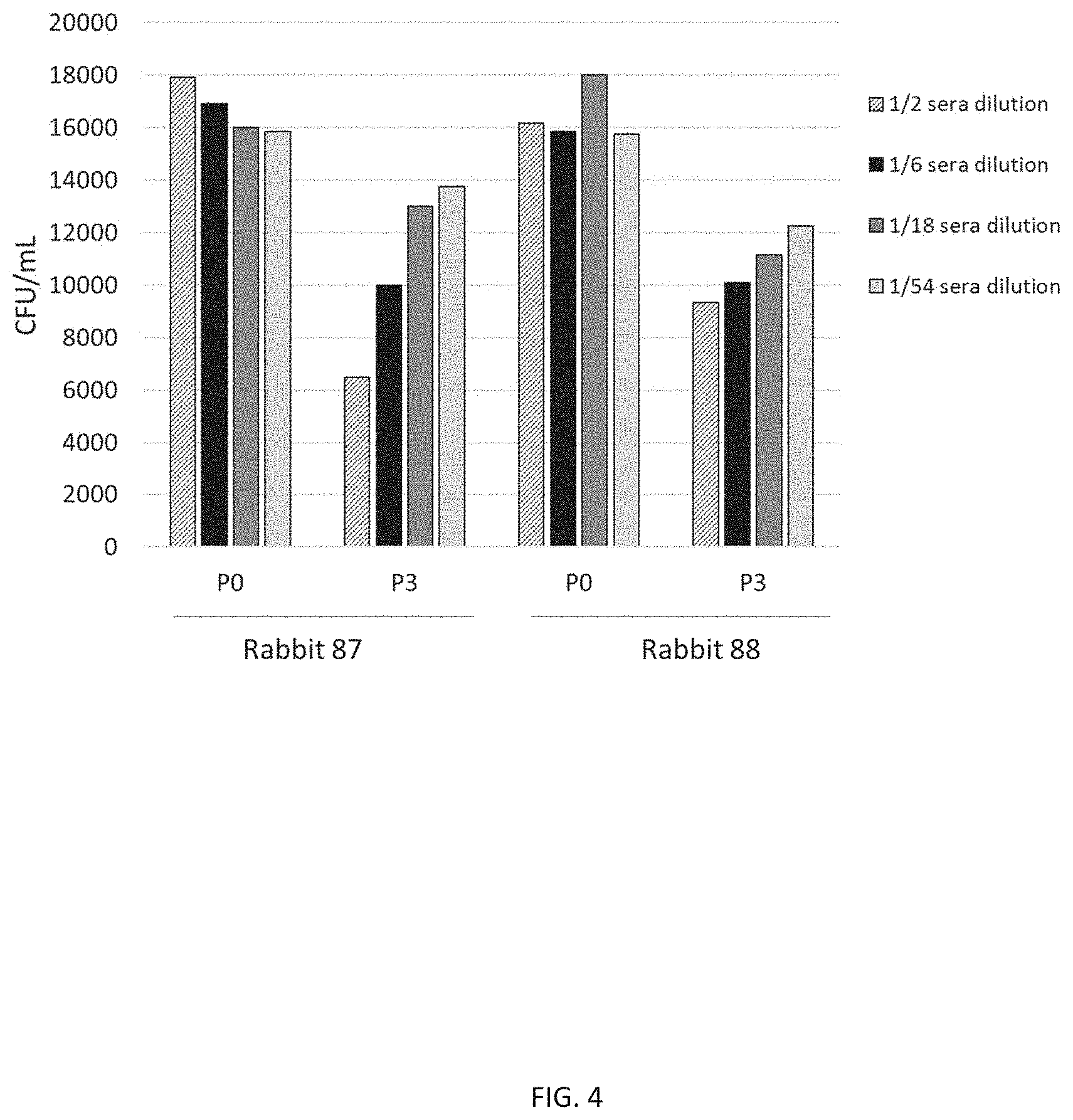
[0016] FIG. 4 illustrates the presence of functional antibodies against a representative S. pneumoniae serotype (e.g., serotype 6B) in CP1 immune sera. S. pneumoniae serotype 6B was incubated in a modified concentrated opsonophagocytic assay (COPA) with heat inactivated pre-immune (P0) and immune (P3) serum, at various dilutions, from each of two rabbits (87 and 88) immunized with CP1 adjuvanted with aluminum phosphate. The colony forming units (CFU) for each dilution and sera combination were enumerated on blood agar plates after overnight incubation. The presence of functional antibodies is shown by killing of S. pneumoniae, i.e., reduction of CFUs following incubation with immune sera. Each vertical bar on the graph represents CFU/ml for each sample of the indicated CP1 serum and dilution, at the indicated timepoint (bottom of graph).
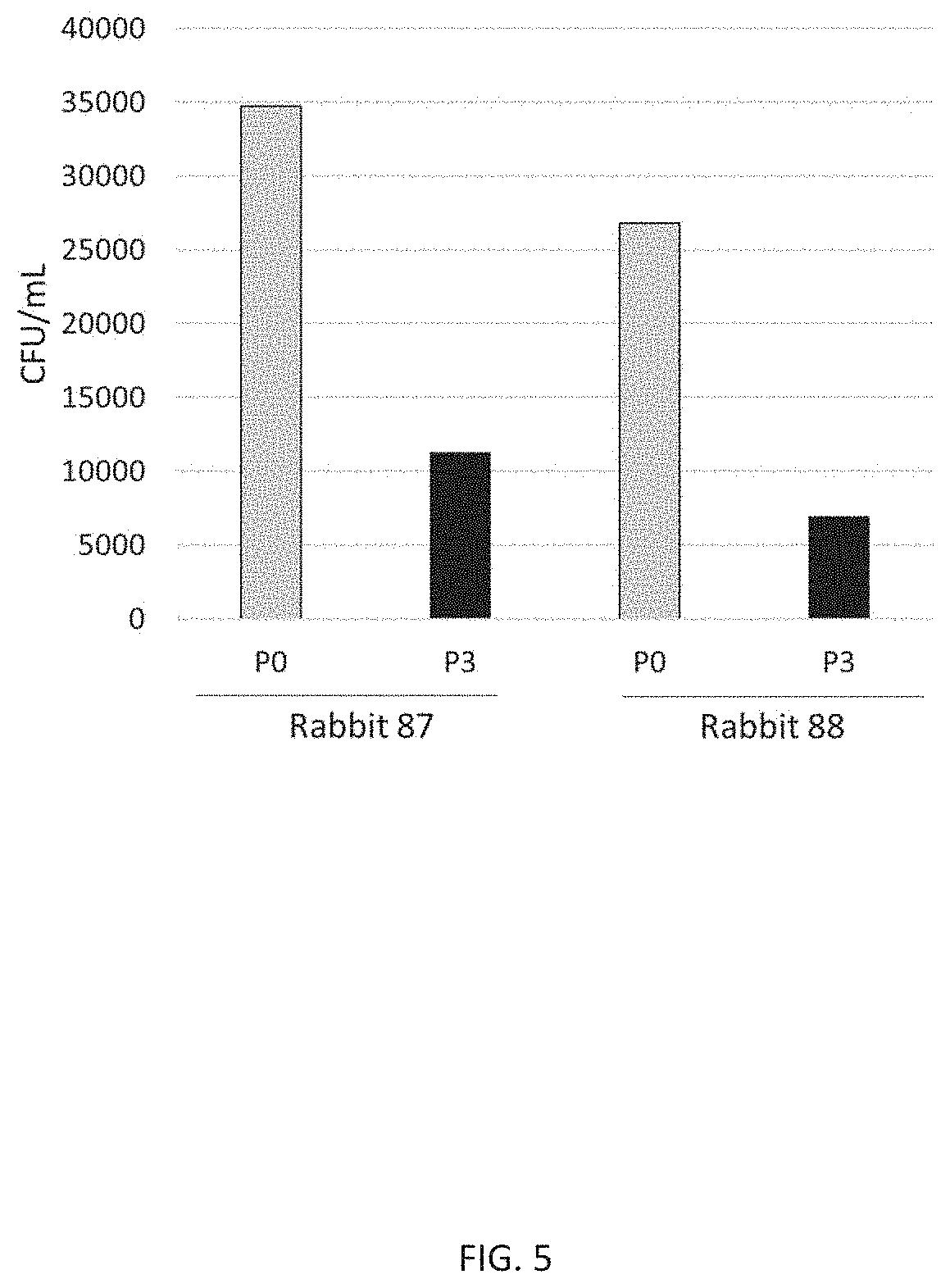
[0017] FIG. 5 illustrates the presence of functional antibodies against a representative S. pneumoniae serotype (e.g., serotype 15A) in CP1 immune sera. S. pneumoniae serotype 15A was incubated in a modified concentrated opsonophagocytic assay (COPA) with heat inactivated pre-immune (P0) and immune (P3) serum, at 1/2 dilution, from each of two rabbits (87 and 88) immunized with CP1 adjuvanted with aluminum phosphate. The colony forming units (CFU) for each serum were enumerated on blood agar plates after overnight incubation. The presence of functional antibodies is shown by killing of S. pneumoniae, i.e., reduction of CFUs following incubation with immune sera. Each vertical bar on the graph represents CFU/ml for each sample of the indicated CP1 serum at the indicated timepoint (bottom of graph).
[0018] FIG. 6 illustrates the presence of functional antibodies against a representative S. pneumoniae serotype (e.g., serotype 35B) in CP1 immune sera. S. pneumoniae serotype 35B was incubated in a modified concentrated opsonophagocytic assay (COPA) with heat inactivated pre-immune (P0) and immune (P3) serum, at 1/2 dilution, from each of two rabbits (87 and 88) immunized with CP1 adjuvanted with aluminum phosphate. The colony forming units (CFU) for each serum were enumerated on blood agar plates after overnight incubation. The presence of functional antibodies is shown by killing of S. pneumoniae, i.e., reduction of CFUs following incubation with immune sera. Each vertical bar on the graph represents CFU/ml for each sample of the indicated CP1 serum at the indicated timepoint (bottom of graph).
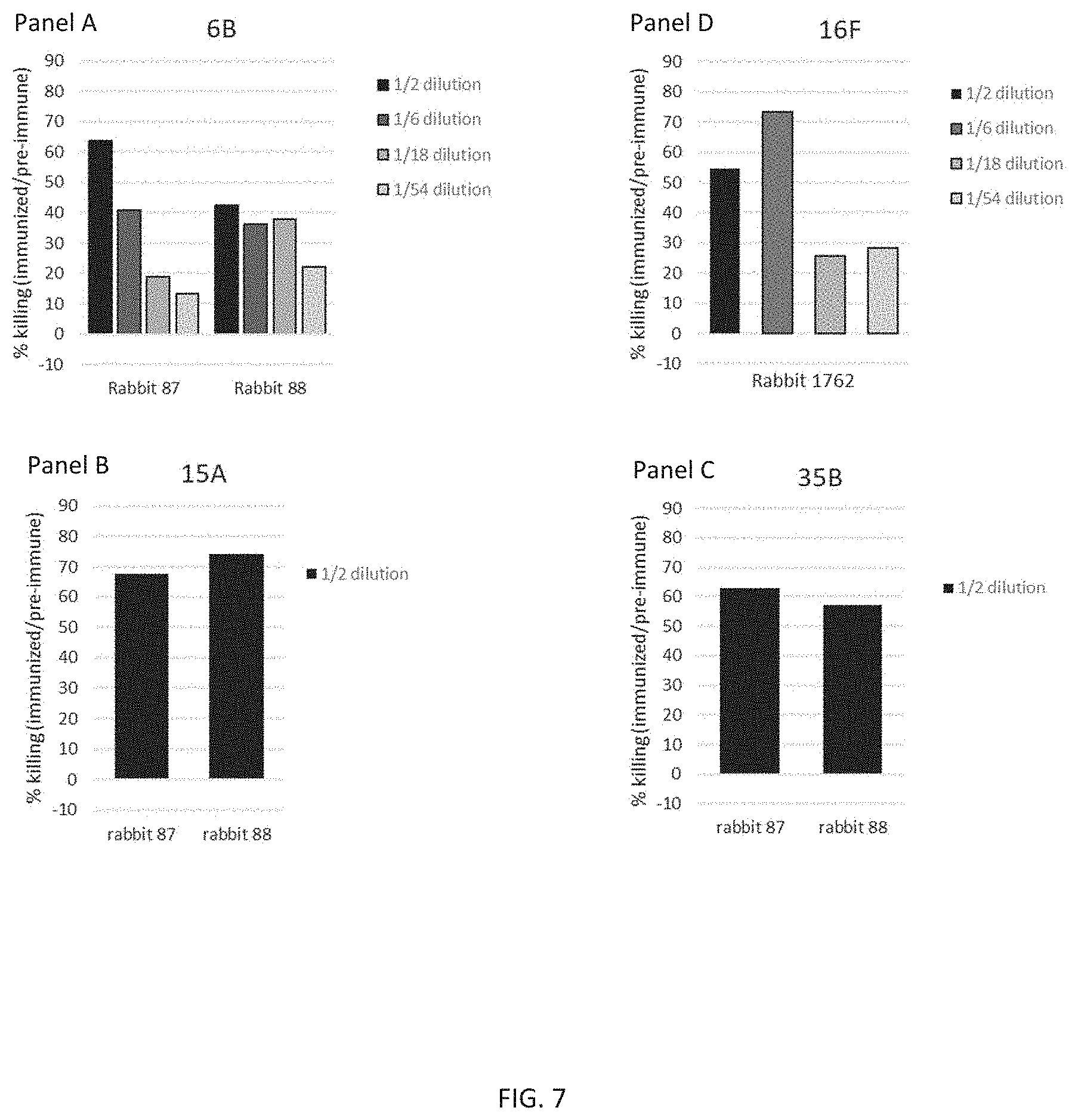
[0019] FIG. 7 illustrates the presence of functional antibodies against representative S. pneumoniae serotypes. S. pneumoniae serotypes 6B (Panel A), 16F (Panel D), 15A (Panel B), and 35B (Panel C) were separately incubated in a modified concentrated opsonophagocytic assay (COPA) with heat-inactivated pre-immune (P0) and immune (P3) sera, at various dilutions, from rabbits (87, 88, and 1762) immunized with CP1 adjuvanted with aluminum phosphate. The presence of functional antibodies is shown by killing of S. pneumoniae. Results are expressed as percent killing activity, i.e., the percent reduction in S. pneumoniae colony forming units (CFU), following incubation with immune (P3) sera, relative to incubation with matched pre-immune (P0) sera. Each vertical bar of Panels A-D represents the percent killing activity observed with the indicated dilution of the indicated CP1 serum (bottom of each graph), against the indicated S. pneumoniae serotype (top of each graph),
[0020] FIG. 8 illustrates immune responses (e.g., Th17 responses) to an exemplary fusion protein CP1 compared to a SP1500 or SP0785 polypeptide. Mice were immunized with CP1, SP1500, or SP0785 polypeptides, adjuvanted with cholera toxin, or were immunized with Rhavi protein adjuvanted with cholera toxin (control). Th17 responses are shown as geometric mean concentration of IL-17A secreted in media after stimulation of peripheral blood samples of immunized mice with purified SP0785 polypeptide (Panel A), purified SP1500 polypeptide (Panel B), or killed (inactivated) pneumococcal whole cells (WCV; Panel C). Each point on the graphs represents secreted IL-17A for one mouse. Horizontal bars represent the geometric mean of secreted IL-17A for each group. CT: cholera toxin.
[0021] FIG. 9 illustrates immune responses (e.g., Th17 responses) to an exemplary fusion protein CP1 compared to a fusion protein SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT. Mice were immunized with CP1 or SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT, adjuvanted with cholera toxin, or were immunized with Rhavi protein adjuvanted with cholera toxin (control). Th17 responses are shown as geometric mean concentration of IL-17A secreted in media after stimulation of peripheral blood samples of immunized mice with purified SP0785 polypeptide (Panel A), or purified SP1500 polypeptide (Panel B). Each point on the graphs represents secreted IL-17A for one mouse. Horizontal bars represent the geometric mean of secreted IL-17A for each group. CT: cholera toxin.
[0022] FIG. 10 illustrates immune responses (e.g., Th17 responses) to an exemplary fusion protein CP1 compared to a mixture (unconjugated) of SP0785, SP1500, and Rhavi polypeptides. Mice were immunized with CP1 or a mixture (unconjugated) of SP0785, SP1500, and Rhavi polypeptides, adjuvanted with cholera toxin, or were immunized with Rhavi protein adjuvanted with cholera toxin (control). Th17 responses are shown as geometric mean concentration of IL-17A secreted in media after stimulation of peripheral blood samples of immunized mice with purified SP0785 polypeptide (Panel A), purified SP1500 polypeptide (Panel B), or killed (inactivated) pneumococcal whole cells (WCV; Panel C). Each point on the graphs represents secreted IL-17A for one mouse. Horizontal bars represent the geometric mean of secreted IL-17A for each group. CT: cholera toxin; mixture:mixture (unconjugated) of SP0785, SP1500, and Rhavi polypeptides.
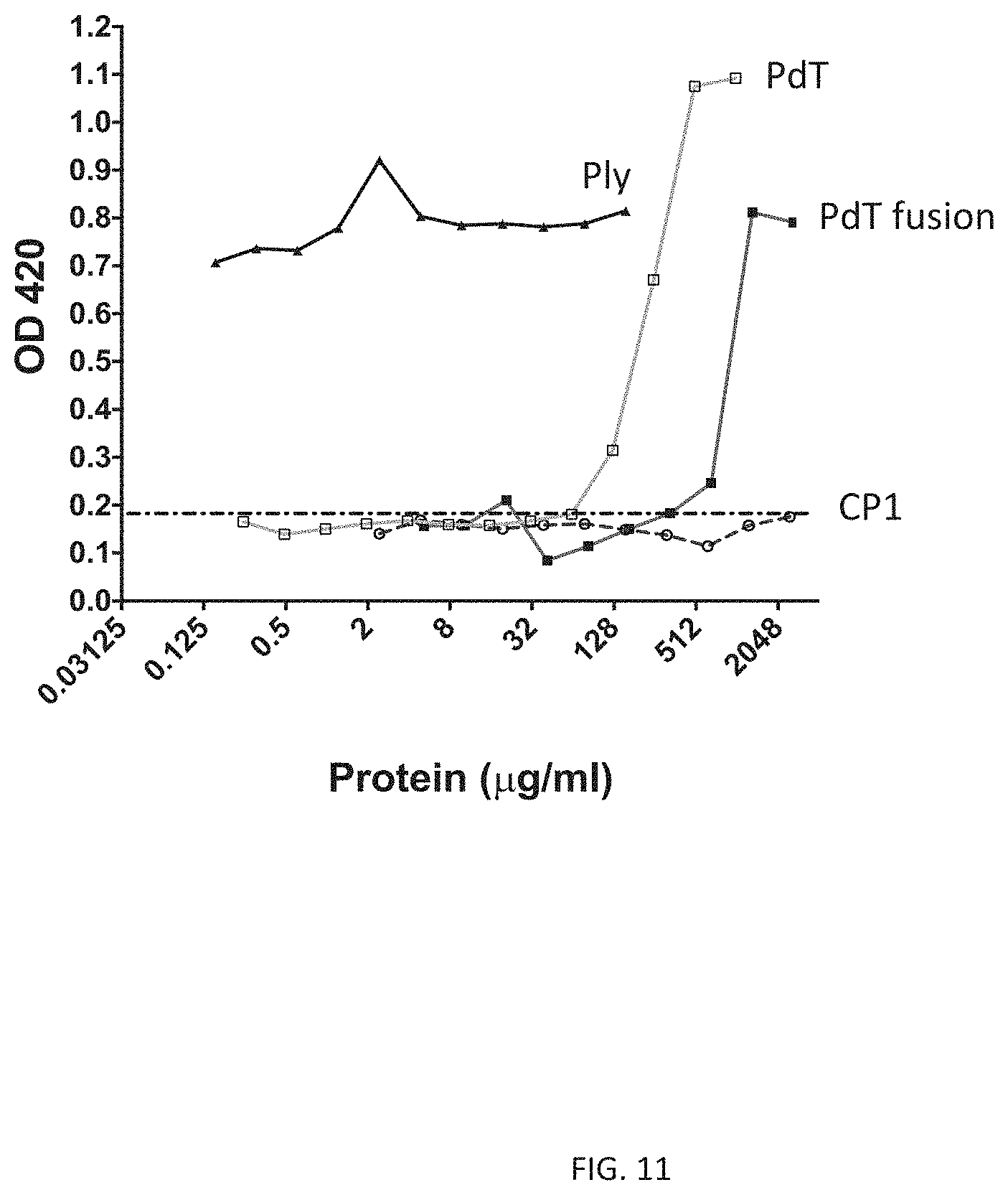
[0023] FIG. 11 illustrates hemolytic activity of an exemplary fusion protein CP1 and fusion protein SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT (PdT fusion) against sheep red blood cells. Sheep red blood cells were incubated with positive control protein pneumolysin (Ply), pneumolysoid PdT, CP1, or fusion protein SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT at the concentrations indicated on the x axis. Hemolytic activity as measured by OD.sub.420 of the supernatants is plotted on the y axis.
CERTAIN DEFINITIONS
[0024] In this application, unless otherwise clear from context, (i) the term "a" may be understood to mean "at least one"; (ii) the term "or" may be understood to mean "and/or"; (iii) the terms "comprising" and "including" may be understood to encompass itemized components or steps whether presented by themselves or together with one or more additional components or steps; and (iv) the terms "about" and "approximately" may be understood to permit standard variation as would be understood by those of ordinary skill in the art; and (v) where ranges are provided, endpoints are included.
[0025] About: The term "about", when used herein in reference to a value, refers to a value that is similar, in context to the referenced value. In general, those skilled in the art, familiar with the context, will appreciate the relevant degree of variance encompassed by "about" in that context. For example, in some embodiments, the term "about" may encompass a range of values that within 25%, 20%, 19%, 18%, 17%, 16%, 15%, 14%, 13%, 12%, 11%, 10%, 9%, 8%, 7%, 6%, 5%, 4%, 3%, 2%, 1%, or less of the referred value.
[0026] Administration: As used herein, the term "administration" typically refers to the administration of a composition to a subject or system to achieve delivery of an agent that is, or is included in, the composition. Those of ordinary skill in the art will be aware of a variety of routes that may, in appropriate circumstances, be utilized for administration to a subject, for example a human. For example, in some embodiments, administration may be ocular, oral, parenteral, topical, etc. In some particular embodiments, administration may be bronchial (e.g., by bronchial instillation), buccal, dermal (which may be or comprise, for example, one or more of topical to the dermis, intradermal, interdermal, transdermal, etc.), enteral, intra-arterial, intradermal, intragastrical, intramedullary, intramuscular, intranasal, intraperitoneal, intrathecal, intravenous, intraventricular, within a specific organ (e.g., intrahepatic), mucosal, nasal, oral, rectal, subcutaneous, sublingual, topical, tracheal (e.g., by intratracheal instillation), vaginal, vitreal, etc. In some embodiments, administration may involve only a single dose. In some embodiments, administration may involve application of a fixed number of doses. In some embodiments, administration may involve dosing that is intermittent (e.g., a plurality of doses separated in time) and/or periodic (e.g., individual doses separated by a common period of time) dosing. In some embodiments, administration may involve continuous dosing (e.g., perfusion) for at least a selected period of time.
[0027] Agent: In general, the term "agent", as used herein, may be used to refer to a compound or entity of any chemical class including, for example, a polypeptide, nucleic acid, saccharide, lipid, small molecule, metal, or combination or complex thereof. In appropriate circumstances, as will be clear from context to those skilled in the art, the term may be utilized to refer to an entity that is or comprises a cell or organism, or a fraction, extract, or component thereof. Alternatively or additionally, as context will make clear, the term may be used to refer to a natural product in that it is found in and/or is obtained from nature. In some instances, again as will be clear from context, the term may be used to refer to one or more entities that is man-made in that it is designed, engineered, and/or produced through action of the hand of man and/or is not found in nature. In some embodiments, an agent may be utilized in isolated or pure form; in some embodiments, an agent may be utilized in crude form. In some embodiments, potential agents may be provided as collections or libraries, for example that may be screened to identify or characterize active agents within them. In some cases, the term "agent" may refer to a compound or entity that is or comprises a polymer; in some cases, the term may refer to a compound or entity that comprises one or more polymeric moieties. In some embodiments, the term "agent" may refer to a compound or entity that is not a polymer and/or is substantially free of any polymer and/or of one or more particular polymeric moieties. In some embodiments, the term may refer to a compound or entity that lacks or is substantially free of any polymeric moiety.
[0028] Amino acid: In its broadest sense, the term "amino acid", as used herein, refers to any compound and/or substance that can be incorporated into a polypeptide chain, e.g., through formation of one or more peptide bonds. In some embodiments, an amino acid has the general structure H.sub.2N--C(H)(R)--COOH. In some embodiments, an amino acid is a naturally-occurring amino acid. In some embodiments, an amino acid is a non-natural amino acid; in some embodiments, an amino acid is a D-amino acid; in some embodiments, an amino acid is an L-amino acid. "Standard amino acid" refers to any of the twenty standard L-amino acids commonly found in naturally occurring peptides. "Non-standard amino acid" refers to any amino acid, other than the standard amino acids, regardless of whether it is prepared synthetically or obtained from a natural source. In some embodiments, an amino acid, including a carboxy- and/or amino-terminal amino acid in a polypeptide, can contain a structural modification as compared with the general structure above. For example, in some embodiments, an amino acid may be modified by methylation, amidation, acetylation, pegylation, glycosylation, phosphorylation, and/or substitution (e.g., of the amino group, the carboxylic acid group, one or more protons, and/or the hydroxyl group) as compared with the general structure. In some embodiments, such modification may, for example, alter the circulating half-life of a polypeptide containing the modified amino acid as compared with one containing an otherwise identical unmodified amino acid. In some embodiments, such modification does not significantly alter a relevant activity of a polypeptide containing the modified amino acid, as compared with one containing an otherwise identical unmodified amino acid. As will be clear from context, in some embodiments, the term "amino acid" may be used to refer to a free amino acid; in some embodiments it may be used to refer to an amino acid residue of a polypeptide.
[0029] Antibody: As used herein, the term "antibody" refers to a polypeptide that includes canonical immunoglobulin sequence elements sufficient to confer specific binding to a particular target antigen. As is known in the art, intact antibodies as produced in nature are approximately 150 kDa tetrameric agents comprised of two identical heavy chain polypeptides (about 50 kDa each) and two identical light chain polypeptides (about 25 kDa each) that associate with each other into what is commonly referred to as a "Y-shaped" structure. Each heavy chain is comprised of at least four domains (each about 110 amino acids long)--an amino-terminal variable (VH) domain (located at the tips of the Y structure), followed by three constant domains: CH1, CH2, and the carboxy-terminal CH3 (located at the base of the Y's stem). A short region, known as the "switch", connects the heavy chain variable and constant regions. The "hinge" connects CH2 and CH3 domains to the rest of the antibody. Two disulfide bonds in this hinge region connect the two heavy chain polypeptides to one another in an intact antibody. Each light chain is comprised of two domains--an amino-terminal variable (VL) domain, followed by a carboxy-terminal constant (CL) domain, separated from one another by another "switch". Intact antibody tetramers are comprised of two heavy chain-light chain dimers in which the heavy and light chains are linked to one another by a single disulfide bond; two other disulfide bonds connect the heavy chain hinge regions to one another, so that the dimers are connected to one another and the tetramer is formed. Naturally-produced antibodies are also glycosylated, typically on the CH2 domain. Each domain in a natural antibody has a structure characterized by an "immunoglobulin fold" formed from two beta sheets (e.g., 3-, 4-, or 5-stranded sheets) packed against each other in a compressed antiparallel beta barrel. Each variable domain contains three hypervariable loops known as "complement determining regions" (CDR1, CDR2, and CDR3) and four somewhat invariant "framework" regions (FR1, FR2, FR3, and FR4). When natural antibodies fold, the FR regions form the beta sheets that provide the structural framework for the domains, and the CDR loop regions from both the heavy and light chains are brought together in three-dimensional space so that they create a single hypervariable antigen binding site located at the tip of the Y structure. The Fc region of naturally-occurring antibodies binds to elements of the complement system, and also to receptors on effector cells, including for example effector cells that mediate cytotoxicity. As is known in the art, affinity and/or other binding attributes of Fc regions for Fc receptors can be modulated through glycosylation or other modification. In some embodiments, antibodies produced and/or utilized in accordance with the present invention include glycosylated Fc domains, including Fc domains with modified or engineered such glycosylation. For purposes of the present invention, in some embodiments, any polypeptide or complex of polypeptides that includes sufficient immunoglobulin domain sequences as found in natural antibodies can be referred to and/or used as an "antibody", whether such polypeptide is naturally produced (e.g., generated by an organism reacting to an antigen), or produced by recombinant engineering, chemical synthesis, or other artificial system or methodology. In some embodiments, an antibody is polyclonal; in some embodiments, an antibody is monoclonal. In some embodiments, an antibody has constant region sequences that are characteristic of mouse, rabbit, primate, or human antibodies. In some embodiments, antibody sequence elements are humanized, primatized, chimeric, etc., as is known in the art. Moreover, the term "antibody" as used herein, can refer in appropriate embodiments (unless otherwise stated or clear from context) to any of the art-known or developed constructs or formats for utilizing antibody structural and functional features in alternative presentation. For example, in some embodiments, an antibody utilized in accordance with the present invention is in a format selected from, but not limited to, intact IgA, IgG, IgE or IgM antibodies; bi- or multi-specific antibodies (e.g., Zybodies.RTM., etc.); antibody fragments such as Fab fragments, Fab' fragments, F(ab')2 fragments, Fd' fragments, Fd fragments, and isolated CDRs or sets thereof; single chain Fvs; polypeptide-Fc fusions; single domain antibodies (e.g., shark single domain antibodies such as IgNAR or fragments thereof); cameloid antibodies; masked antibodies (e.g.,) Probodies.RTM.; Small Modular ImmunoPharmaceuticals ("SMIPs.TM."); single chain or Tandem diabodies (TandAb.RTM.); VHHs; Anticalins.RTM.; Nanobodies.RTM. minibodies; BiTE.RTM. s; ankyrin repeat proteins or DARPINs.RTM., Avimers.RTM., DARTs; TCR-like antibodies; Adnectins.RTM., Affilins.RTM., Trans-bodies.RTM.; Affibodies.RTM.; TrimerX.RTM., MicroProteins; Fynomers.RTM., Centyrins.RTM., and KALBITOR.RTM.s. In some embodiments, an antibody may lack a covalent modification (e.g., attachment of a glycan) that it would have if produced naturally. In some embodiments, an antibody may contain a covalent modification (e.g., attachment of a glycan, a payload [e.g., a detectable moiety, a therapeutic moiety, a catalytic moiety, etc.], or other pendant group [e.g., poly-ethylene glycol, etc.]).
[0030] Antigen: The term "antigen", as used herein, refers to (i) an agent that induces an immune response; and/or (ii) an agent that binds to a T cell receptor (e.g., when presented by an MHC molecule) or to an antibody. In some embodiments, an antigen induces a humoral response (e.g., including production of antigen-specific antibodies); in some embodiments, an antigen induces a cellular response (e.g., involving T cells whose receptors specifically interact with the antigen). In some embodiments, an antigen induces a humoral response and a cellular response. In some embodiments, an antigen binds to an antibody and may or may not induce a particular physiological response in an organism. In general, an antigen may be or include any chemical entity such as, for example, a small molecule, a nucleic acid, a polypeptide, a carbohydrate, a lipid, a polymer (in some embodiments other than a biologic polymer (e.g., other than a nucleic acid or amino acid polymer)), etc. In some embodiments, an antigen is or comprises a polypeptide. In some embodiments, an antigen is or comprises a polysaccharide. Those of ordinary skill in the art will appreciate that, in general, an antigen may be provided in isolated or pure form, or alternatively may be provided in crude form (e.g., together with other materials, for example in an extract such as a cellular extract or other relatively crude preparation of an antigen-containing source). In some embodiments, antigens utilized in accordance with the present invention are provided in a crude form. In some embodiments, an antigen is a recombinant antigen. In some embodiments, an antigen is a polypeptide or a polysaccharide that, upon administration to a subject, induces a specific and/or clinically relevant immune response to such polypeptide or polysaccharide. In some embodiments, an antigen is selected to induce a specific and/or clinically relevant immune response to such polypeptide or polysaccharide.
[0031] Associated with: Two entities are "associated" with one another, as that term is used herein, if the presence, level and/or form of one is correlated with that of the other. In some embodiments, two or more entities are physically "associated" with one another if they interact, directly or indirectly, so that they are and/or remain in physical proximity with one another. In some embodiments, two or more entities that are physically associated with one another are covalently linked to one another. In some embodiments, two or more entities that are physically associated with one another are not covalently linked to one another but are non-covalently associated, for example by means of affinity interactions, electrostatic interactions, hydrogen bonds, van der Waals interaction, hydrophobic interactions, magnetism, and combinations thereof.
[0032] Binding: It will be understood that the term "binding", as used herein, typically refers to a non-covalent association between or among two or more entities. "Direct" binding involves physical contact between entities or moieties; indirect binding involves physical interaction by way of physical contact with one or more intermediate entities. Binding between two or more entities can typically be assessed in any of a variety of contexts--including where interacting entities or moieties are studied in isolation or in the context of more complex systems (e.g., while covalently or otherwise associated with a carrier entity and/or in a biological system or cell).
[0033] Carrier protein: As used herein, the term "carrier protein" refers to a protein or peptide that is coupled, complexed, or otherwise associated with a hapten (e.g., a small peptide or lipid) or less immunogenic antigen (e.g., a polysaccharide) and that induces or improves an immune response to such a coupled, or complexed, or otherwise associated hapten (e.g., a small peptide or lipid) or less immunogenic antigen (e.g., a polysaccharide). In some embodiments, such an immune response is or comprises a response to a hapten or less immunogenic antigen that is coupled, complexed, or otherwise associated with such a carrier protein. In some embodiments, such an immune response is or comprises a response to both a carrier protein and a hapten or less immunogenic antigen that is coupled, complexed, or otherwise associated with such a carrier protein. In some embodiments, no significant immune response to a carrier protein itself occurs. In some embodiments, immune response to a carrier protein may be detected; in some such embodiments, immune response to such a carrier protein is strong. In some embodiments, a carrier protein is coupled, complexed, or otherwise associated with one or more other molecules.
[0034] Colonization: As used herein, the term "colonization" generally refers to the ability of a microbe to grow at a target site or surface. For example, the term "colonization" refers to the ability of a microbe (e.g., a bacterium) to grow at an anatomical site (e.g., a mucosal membrane, gastrointestinal tract, injury site, organ, etc.) of a host.
[0035] Combination therapy: As used herein, the term "combination therapy" refers to those situations in which a subject is exposed to two or more therapeutic regimens (e.g., two or more therapeutic agents). In some embodiments, the two or more regimens may be administered simultaneously; in some embodiments, such regimens may be administered sequentially (e.g., all "doses" of a first regimen are administered prior to administration of any doses of a second regimen); in some embodiments, such agents are administered in overlapping dosing regimens. In some embodiments, "administration" of combination therapy may involve administration of one or more agent(s) or modality(ies) to a subject receiving the other agent(s) or modality(ies) in the combination. For clarity, combination therapy does not require that individual agents be administered together in a single composition (or even necessarily at the same time), although in some embodiments, two or more agents, or active moieties thereof, may be administered together in a combination composition, or even in a combination compound (e.g., as part of a single chemical complex or covalent entity).
[0036] Derivative: As used herein, the term "derivative", or grammatical equivalents thereof, refers to a structural analogue of a reference substance. That is, a "derivative" is a substance that shows significant structural similarity with the reference substance, for example sharing a core or consensus structure, but also differs in certain discrete ways. Such a substance would be said to be "derived from" said reference substance. In some embodiments, a derivative is a substance that can be generated from the reference substance by chemical manipulation. In some embodiments, a derivative is a substance that can be generated through performance of a synthetic process substantially similar to (e.g., sharing a plurality of steps with) one that generates the reference substance.
[0037] Domain: The term "domain" as used herein refers to a section or portion of an entity. In some embodiments, a "domain" is associated with a particular structural and/or functional feature of the entity so that, when the domain is physically separated from the rest of its parent entity, it substantially or entirely retains the particular structural and/or functional feature. Alternatively or additionally, a domain may be or include a portion of an entity that, when separated from that (parent) entity and linked with a different (recipient) entity, substantially retains and/or imparts on the recipient entity one or more structural and/or functional features that characterized it in the parent entity. In some embodiments, a domain is a section or portion of a molecule (e.g., a small molecule, carbohydrate, lipid, nucleic acid, or polypeptide). In some embodiments, a domain is a section of a polypeptide; in some such embodiments, a domain is characterized by a particular structural element (e.g., a particular amino acid sequence or sequence motif, .alpha.-helix character, .beta.-sheet character, coiled-coil character, random coil character, etc.), and/or by a particular functional feature (e.g., binding activity, enzymatic activity, folding activity, signaling activity, etc.).
[0038] Dosage form or unit dosage form: Those skilled in the art will appreciate that the term "dosage form" may be used to refer to a physically discrete unit of an active agent (e.g., a therapeutic or diagnostic agent) for administration to a subject. Typically, each such unit contains a predetermined quantity of active agent. In some embodiments, such quantity is a unit dosage amount (or a whole fraction thereof) appropriate for administration in accordance with a dosing regimen that has been determined to correlate with a desired or beneficial outcome when administered to a relevant population (i.e., with a therapeutic dosing regimen). Those of ordinary skill in the art appreciate that the total amount of a therapeutic composition or agent administered to a particular subject is determined by one or more attending physicians and may involve administration of multiple dosage forms.
[0039] Dosing regimen: Those skilled in the art will appreciate that the term "dosing regimen" may be used to refer to a set of unit doses (typically more than one) that are administered individually to a subject, typically separated by periods of time. In some embodiments, a given therapeutic agent has a recommended dosing regimen, which may involve one or more doses. In some embodiments, a dosing regimen comprises a plurality of doses each of which is separated in time from other doses. In some embodiments, individual doses are separated from one another by a time period of the same length; in some embodiments, a dosing regimen comprises a plurality of doses and at least two different time periods separating individual doses. In some embodiments, all doses within a dosing regimen are of the same unit dose amount. In some embodiments, different doses within a dosing regimen are of different amounts. In some embodiments, a dosing regimen comprises a first dose in a first dose amount, followed by one or more additional doses in a second dose amount different from the first dose amount. In some embodiments, a dosing regimen comprises a first dose in a first dose amount, followed by one or more additional doses in a second dose amount same as the first dose amount. In some embodiments, a dosing regimen is correlated with a desired or beneficial outcome when administered across a relevant population (i.e., is a therapeutic dosing regimen).
[0040] Fragment: A "fragment" of a material or entity as described herein has a structure that includes a discrete portion of the whole, but lacks one or more moieties found in the whole. In some embodiments, a fragment consists of such a discrete portion. In some embodiments, a fragment includes a discrete portion of the whole which discrete portion shares one or more functional characteristics found in the whole. In some embodiments, a fragment consists of such a discrete portion. In some embodiments, a fragment consists of or comprises a characteristic structural element or moiety found in the whole. In some embodiments, a fragment of a polymer, e.g., a polypeptide or polysaccharide, comprises or consists of at least 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, 90, 95, 100, 110, 120, 130, 140, 150, 160, 170, 180, 190, 200, 210, 220, 230, 240, 250, 275, 300, 325, 350, 375, 400, 425, 450, 475, 500 or more monomeric units (e.g., residues) as found in the whole polymer. In some embodiments, a polymer fragment comprises or consists of at least about 5%, 10%, 15%, 20%, 25%, 30%, 25%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% of the monomeric units (e.g., residues) found in the whole polymer. The whole material or entity may in some embodiments be referred to as the "parent" of the whole.
[0041] Homology: As used herein, the term "homology" refers to the overall relatedness between polymeric molecules, e.g., between nucleic acid molecules (e.g., DNA molecules and/or RNA molecules) and/or between polypeptide molecules. In some embodiments, polymeric molecules are considered to be "homologous" to one another if their sequences are at least 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98% or 99% identical. In some embodiments, polymeric molecules are considered to be "homologous" to one another if their sequences are at least 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98% or 99% similar (e.g., containing residues with related chemical properties at corresponding positions). For example, as is well known by those of ordinary skill in the art, certain amino acids are typically classified as similar to one another as "hydrophobic" or "hydrophilic" amino acids, and/or as having "polar" or "non-polar" side chains. Substitution of one amino acid for another of the same type may often be considered a "homologous" substitution.
[0042] Identity: As used herein, the term "identity" refers to the overall relatedness between polymeric molecules, e.g., between nucleic acid molecules (e.g., DNA molecules and/or RNA molecules) and/or between polypeptide molecules. In some embodiments, polymeric molecules are considered to be "substantially identical" to one another if their sequences are at least 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98% or 99% identical. Calculation of the percent identity of two nucleic acid or polypeptide sequences, for example, can be performed by aligning the two sequences for optimal comparison purposes (e.g., gaps can be introduced in one or both of a first and a second sequence for optimal alignment and non-identical sequences can be disregarded for comparison purposes). In some embodiments, the length of a sequence aligned for comparison purposes is at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.5%, or substantially 100% of the length of a reference sequence. The nucleotides at corresponding positions are then compared. When a position in the first sequence is occupied by the same residue (e.g., nucleotide or amino acid) as the corresponding position in the second sequence, then the molecules are identical at that position. The percent identity between the two sequences is a function of the number of identical positions shared by the sequences, taking into account the number of gaps, and the length of each gap, which needs to be introduced for optimal alignment of the two sequences. The comparison of sequences and determination of percent identity between two sequences can be accomplished using a mathematical algorithm. For example, the percent identity between two nucleotide sequences can be determined using the algorithm of Meyers and Miller, 1989, which has been incorporated into the ALIGN program (version 2.0). In some exemplary embodiments, nucleic acid sequence comparisons made with the ALIGN program use a PAM120 weight residue table, a gap length penalty of 12 and a gap penalty of 4. The percent identity between two nucleotide sequences can, alternatively, be determined using the GAP program in the GCG software package using an NWSgapdna.CMP matrix.
[0043] Improve, increase, inhibit or reduce: As used herein, the terms "improve", "increase", "inhibit`, "reduce", or grammatical equivalents thereof, indicate values that are relative to a baseline or other reference measurement. In some embodiments, an appropriate reference measurement may be or comprise a measurement in a particular system (e.g., in a single subject) under otherwise comparable conditions absent presence of (e.g., prior to and/or after) a particular agent or treatment, or in presence of an appropriate comparable reference agent. In some embodiments, an appropriate reference measurement may be or comprise a measurement in comparable system known or expected to respond in a particular way, in presence of the relevant agent or treatment.
[0044] Immunologically effective amount or immunologically effective dose: As used herein, "immunologically effective amount" or "immunologically effective dose" refers to an amount of an antigenic or immunogenic substance, e.g., an antigen, immunogen, immunogenic complex, immunogenic composition, vaccine, or pharmaceutical composition, which when administered to a subject, either in a single dose or as part of a series of doses, that is sufficient to enhance a subject's own immune response against a subsequent exposure to a pathogen. In some embodiments, the pathogen is S. pneumoniae. In some embodiments, the immune response is against one or more different serotypes of S. pneumoniae. In some embodiments, the immune response is against two or more different serotypes of S. pneumoniae. In some embodiments, the immune response is against nine or more different serotypes of S. pneumoniae. In some embodiments, the immune response is against thirteen or more different serotypes of S. pneumoniae. In some embodiments, the immune response is against fifteen or more different serotypes of S. pneumoniae. In some embodiments, the immune response is against twenty-three or more different serotypes of S. pneumoniae. In some embodiments, the immune response is against twenty-four or more different serotypes of S. pneumoniae. An immunologically effective amount may vary based on the subject to be treated, the species of the subject, the degree of immune response desired to induce, etc. In some embodiments, an immunologically effective amount is sufficient for treatment or protection of a subject having or at risk of having disease. In some embodiments, an immunologically effective amount refers to a non-toxic but sufficient amount that can be an amount to treat, attenuate, or prevent infection and/or disease (e.g., bacterial infection, pneumococcal infection, bacterial colonization, pneumococcal colonization, complications associated with bacterial infection, complications associated with pneumococcal infection, etc.) in any subject. In some embodiments, an immunologically effective amount is sufficient to induce an immunoprotective response upon administration to a subject.
[0045] Immunoprotective response or protective response: As used herein, "immunoprotective response" or "protective response" refers to an immune response that mediates antigen or immunogen-induced immunological memory. In some embodiments, an immunoprotective response is induced by the administration of a substance, e.g., an antigen, immunogen, immunogenic complex, immunogenic composition, vaccine, or pharmaceutical composition to a subject. In some embodiments, immunoprotection involves one or more of active immune surveillance, a more rapid and effective response upon immune activation as compared to a response observed in a naive subject, efficient clearance of the activating agent or pathogen, followed by rapid resolution of inflammation. In some embodiments, an immunoprotective response is an adaptive immune response. In some embodiments, an immunoprotective response is sufficient to protect an immunized subject from productive infection by a particular pathogen or pathogens to which a vaccine is directed (e.g., S. pneumoniae infection).
[0046] Immunization: As used herein, "immunization", or grammatical equivalents thereof, refers to a process of inducing an immune response to an infectious organism or agent in a subject ("active immunization"), or alternatively, providing immune system components against an infectious organism or agent to a subject ("passive immunization"). In some embodiments, immunization involves the administration of one or more antigens, immunogens, immunogenic complexes, vaccines, immune molecules such as antibodies, immune sera, immune cells such as T cells or B cells, or pharmaceutical compositions to a subject. In some embodiments, immunization is performed by administering an immunologically effective amount of a substance, e.g., an antigen, immunogen, immunogenic complex, immunogenic composition, vaccine, immune molecule such as an antibody, immune serum, immune cell such as a T cell or B cell, or pharmaceutical composition to a subject. In some embodiments, immunization results in an immunoprotective response in the subject. In some embodiments, active immunization is performed by administering to a subject an antigenic or immunogenic substance, e.g., an antigen, immunogen, immunogenic complex, vaccine, or pharmaceutical composition. In some embodiments, passive immunization is performed by administering to a subject an immune system component, e.g., an immune molecule such as an antibody, immune serum, or immune cell such as a T cell or B cell.
[0047] Isolated: As used herein, the term "isolated", or grammatical equivalents thereof, refers to a substance and/or entity that has been (1) separated from at least some of the components with which it was associated when initially produced (whether in nature and/or in an experimental setting), and/or (2) designed, produced, prepared, and/or manufactured by the hand of man. Isolated substances and/or entities may be separated from about 10%, about 20%, about 30%, about 40%, about 50%, about 60%, about 70%, about 80%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or more than about 99% of the other components with which they were initially associated. In some embodiments, isolated agents are about 80%, about 85%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or more than about 99% pure. As used herein, a substance is "pure" if it is substantially free of other components. In some embodiments, as will be understood by those skilled in the art, a substance may still be considered "isolated" or even "pure", after having been combined with certain other components such as, for example, one or more carriers or excipients (e.g., buffer, solvent, water, etc.); in such embodiments, percent isolation or purity of the substance is calculated without including such carriers or excipients. To give but one example, in some embodiments, a biological polymer such as a polypeptide or polysaccharide that occurs in nature is considered to be "isolated" when, a) by virtue of its origin or source of derivation is not associated with some or all of the components that accompany it in its native state in nature; b) it is substantially free of other polypeptides or nucleic acids of the same species from the species that produces it in nature; c) is expressed by or is otherwise in association with components from a cell or other expression system that is not of the species that produces it in nature. Thus, for instance, in some embodiments, a polypeptide or polysaccharide that is chemically synthesized or is synthesized in a cellular system different from that which produces it in nature is considered to be an "isolated" polypeptide or polysaccharide. Alternatively or additionally, in some embodiments, a polypeptide or polysaccharide that has been subjected to one or more purification techniques may be considered to be an "isolated" polypeptide or polysaccharide to the extent that it has been separated from other components a) with which it is associated in nature; and/or b) with which it was associated when initially produced.
[0048] Linker: As used herein, the term "linker" is used to refer to an entity that connects two or more elements to form a multi-element agent. For example, those of ordinary skill in the art appreciate that a polypeptide whose structure includes two or more functional or organizational domains often includes a stretch of amino acids between such domains that links them to one another. In some embodiments, a polypeptide comprising a linker element has an overall structure of the general form S1-L-S2, wherein 51 and S2 may be the same or different and represent two domains associated with one another by the linker (L). In some embodiments, a polypeptide linker is at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, 90, 95, 100 or more amino acids in length. In some embodiments, a linker is characterized in that it tends not to adopt a rigid three-dimensional structure, but rather provides flexibility to the polypeptide. A variety of different linker elements that can appropriately be used when engineering polypeptides (e.g., fusion polypeptides) are known in the art (Holliger et al, 1993; Poljak, 1994).
[0049] Pharmaceutical composition: As used herein, the term "pharmaceutical composition" refers to a composition in which an active agent is formulated together with one or more pharmaceutically acceptable carriers. In some embodiments, the active agent is present in unit dose amount appropriate for administration in a therapeutic regimen that shows a statistically significant probability of achieving a predetermined therapeutic effect when administered to a relevant population. In some embodiments, a pharmaceutical composition may be specially formulated for administration in solid or liquid form, including those adapted for the following: oral administration, for example, drenches (aqueous or non-aqueous solutions or suspensions), tablets, e.g., those targeted for buccal, sublingual, and systemic absorption, boluses, powders, granules, pastes for application to the tongue; parenteral administration, for example, by subcutaneous, intramuscular, intravenous or epidural injection as, for example, a sterile solution or suspension, or sustained-release formulation; topical application, for example, as a cream, ointment, or a controlled-release patch or spray applied to the skin, lungs, or oral cavity; intravaginally or intrarectally, for example, as a pessary, cream, or foam; sublingually; ocularly; transdermally; or nasally, pulmonary, and to other mucosal surfaces.
[0050] Pharmaceutically acceptable: As used herein, the term "pharmaceutically acceptable" applied to the carrier, diluent, or excipient used to formulate a composition as disclosed herein means that the carrier, diluent, or excipient must be compatible with the other ingredients of the composition and not deleterious to the recipient thereof.
[0051] Polysaccharide: The term "polysaccharide" as used herein refers to a polymeric carbohydrate molecule composed of long chains of monosaccharide units bound together by glycosidic, phosphodiester, or other linkages, and on hydrolysis give the constituent monosaccharides or oligosaccharides. Polysaccharides range in structure from linear to highly branched. Examples include storage polysaccharides such as starch and glycogen, structural polysaccharides such as cellulose and chitin and microbial polysaccharides, and antigenic polysaccharides found in microorganisms including, but not limited to, capsular polysaccharides (CPS), O polysaccharides (OPS), core O polysaccharides (COPS), and lipopolysaccharides (LPS).
[0052] Polypeptide: The term "polypeptide", as used herein, generally has its art-recognized meaning of a polymer of at least three amino acids, e.g., linked to each other by peptide bonds. Those of ordinary skill in the art will appreciate that the term "polypeptide" is intended to be sufficiently general as to encompass not only polypeptides having a complete sequence recited herein, but also to encompass polypeptides that represent functional fragments (i.e., fragments retaining at least one activity) of such complete polypeptides. Moreover, those of ordinary skill in the art understand that protein sequences generally tolerate some substitution without destroying activity. Thus, any polypeptide that retains activity and shares at least about 30-40% overall sequence identity, often greater than about 50%, 60%, 70%, or 80%, and further usually including at least one region of much higher identity, often greater than 90% or even 95%, 96%, 97%, 98%, or 99% in one or more highly conserved regions, usually encompassing at least 3-4 and often up to 20 or more amino acids, with another polypeptide of the same class, is encompassed within the relevant term "polypeptide" as used herein. Polypeptides may contain L-amino acids, D-amino acids, or both and may contain any of a variety of amino acid modifications or analogs known in the art. Useful modifications include, e.g., terminal acetylation, amidation, methylation, etc. In some embodiments, proteins may comprise natural amino acids, non-natural amino acids, synthetic amino acids, and combinations thereof.
[0053] Prevention: The term "prevent" or "prevention", as used herein in connection with a disease, disorder, and/or medical condition, refers to reducing the risk of developing the disease, disorder and/or condition, and/or a delay of onset, and/or reduction in frequency and/or severity of one or more characteristics or symptoms of a particular disease, disorder or condition. In some embodiments, prevention is assessed on a population basis such that an agent is considered to "prevent" a particular disease, disorder or condition if a statistically significant decrease in the development, frequency, and/or intensity of one or more symptoms of the disease, disorder or condition is observed in a population susceptible to the disease, disorder, or condition. In some embodiments, prevention may be considered complete when onset of a disease, disorder or condition has been delayed for a predefined period of time.
[0054] Protein: As used herein, the term "protein" encompasses a polypeptide. Proteins may include moieties other than amino acids (e.g., may be glycoproteins, proteoglycans, etc.) and/or may be otherwise processed or modified. Those of ordinary skill in the art will appreciate that a "protein" can be a complete polypeptide chain as produced by a cell (with or without a signal sequence), or can be a characteristic portion thereof. Those of ordinary skill will appreciate that a protein can sometimes include more than one polypeptide chain, for example linked by one or more disulfide bonds or associated by other means. Polypeptides may contain 1-amino acids, d-amino acids, or both and may contain any of a variety of amino acid modifications or analogs known in the art. Useful modifications include, e.g., terminal acetylation, amidation, methylation, etc. In some embodiments, proteins may comprise natural amino acids, non-natural amino acids, synthetic amino acids, and combinations thereof. The term "peptide" is generally used to refer to a polypeptide having a length of less than about 100 amino acids, less than about 50 amino acids, less than 20 amino acids, or less than 10 amino acids. In some embodiments, proteins are antibodies, antibody fragments, biologically active portions thereof, and/or characteristic portions thereof.
[0055] Recombinant: As used herein, the term "recombinant" is intended to refer to polypeptides that are designed, engineered, prepared, expressed, created, manufactured, and/or isolated by recombinant means, such as polypeptides expressed using a recombinant expression vector transfected into a host cell; polypeptides isolated from a recombinant, combinatorial human polypeptide library; polypeptides isolated from an animal (e.g., a mouse, rabbit, sheep, fish, etc.) that is transgenic for or otherwise has been manipulated to express a gene or genes, or gene components that encode and/or direct expression of the polypeptide or one or more component(s), portion(s), element(s), or domain(s) thereof; and/or polypeptides prepared, expressed, created or isolated by any other means that involves splicing or ligating selected nucleic acid sequence elements to one another, chemically synthesizing selected sequence elements, and/or otherwise generating a nucleic acid that encodes and/or directs expression of the polypeptide or one or more component(s), portion(s), element(s), or domain(s) thereof. In some embodiments, one or more of such selected sequence elements is found in nature. In some embodiments, one or more of such selected sequence elements is designed in silico. In some embodiments, one or more such selected sequence elements results from mutagenesis (e.g., in vivo or in vitro) of a known sequence element, e.g., from a natural or synthetic source such as, for example, in the germline of a source organism of interest (e.g., of a human, a mouse, etc.).
[0056] Reference: As used herein, the term "reference" describes a standard or control relative to which a comparison is performed. For example, in some embodiments, an agent, animal, subject, population, sample, sequence or value of interest is compared with a reference or control agent, animal, subject, population, sample, sequence or value. In some embodiments, a reference or control is tested and/or determined substantially simultaneously with the testing or determination of interest. In some embodiments, a reference or control is a historical reference or control, optionally embodied in a tangible medium. Typically, as would be understood by those skilled in the art, a reference or control is determined or characterized under comparable conditions or circumstances to those under assessment. Those skilled in the art will appreciate when sufficient similarities are present to justify reliance on and/or comparison to a particular possible reference or control.
[0057] Response: As used herein, a "response" to treatment may refer to any beneficial alteration in a subject's condition that occurs as a result of or correlates with treatment. Such alteration may include stabilization of the condition (e.g., prevention of deterioration that would have taken place in the absence of the treatment), amelioration of symptoms of the condition, and/or improvement in the prospects for cure of the condition, etc. It may refer to a subject's response or to a tumor's response. Subject or tumor response may be measured according to a wide variety of criteria, including clinical criteria and objective criteria. Techniques for assessing response include, but are not limited to, clinical examination, positron emission tomography, chest X-ray CT scan, Mill, ultrasound, endoscopy, laparoscopy, presence or level of biomarkers in a sample obtained from a subject, cytology, and/or histology. The exact response criteria can be selected in any appropriate manner, provided that when comparing groups of subjects and/or tumors, the groups to be compared are assessed based on the same or comparable criteria for determining response rate. One of ordinary skill in the art will be able to select appropriate criteria.
[0058] Risk: As will be understood from context, "risk" of a disease, disorder, and/or condition refers to a likelihood that a particular subject will develop the disease, disorder, and/or condition. In some embodiments, risk is expressed as a percentage. In some embodiments, risk is from 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 30, 40, 50, 60, 70, 80, 90 up to 100%. In some embodiments, risk is expressed as a risk relative to a risk associated with a reference sample or group of reference samples. In some embodiments, a reference sample or group of reference samples have a known risk of a disease, disorder, condition and/or event. In some embodiments a reference sample or group of reference samples are from subjects comparable to a particular subject. In some embodiments, relative risk is 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more.
[0059] Serotype: As used herein, the term "serotype", also referred to as a serovar, refers to a distinct variation within a species of bacteria or virus or among immune cells of different subjects. These microorganisms, viruses, or cells are classified together based on their cell surface antigens, allowing the epidemiologic classification of organisms to the sub-species level. A group of serovars with common antigens may be referred to as a serogroup or sometimes serocomplex.
[0060] Subject: As used herein, the term "subject" refers an organism, typically a mammal (e.g., a human, in some embodiments including prenatal human forms). In some embodiments, a subject is suffering from a relevant disease, disorder or condition. In some embodiments, a subject is susceptible to a disease, disorder, or condition. In some embodiments, a subject displays one or more symptoms or characteristics of a disease, disorder or condition. In some embodiments, a subject does not display any symptom or characteristic of a disease, disorder, or condition. In some embodiments, a subject is someone with one or more features characteristic of susceptibility to or risk of a disease, disorder, or condition. In some embodiments, a subject is a patient. In some embodiments, a subject is a subject to whom diagnosis and/or therapy is and/or has been administered.
[0061] Susceptible to: A subject who is "susceptible to" a disease, disorder, or condition is at risk for developing the disease, disorder, or condition. In some embodiments, a subject who is susceptible to a disease, disorder, or condition does not display any symptoms of the disease, disorder, or condition. In some embodiments, a subject who is susceptible to a disease, disorder, or condition has not been diagnosed with the disease, disorder, and/or condition. In some embodiments, a subject who is susceptible to a disease, disorder, or condition is a subject who has been exposed to conditions associated with development of the disease, disorder, or condition. In some embodiments, a risk of developing a disease, disorder, and/or condition is a population-based risk (e.g., family members of subjects suffering from the disease, disorder, or condition).
[0062] Symptoms are reduced: As used herein, "symptoms are reduced" when one or more symptoms of a particular disease, disorder or condition is reduced in magnitude (e.g., intensity, severity, etc.) and/or frequency, e.g., to a stastistically and/or clinically significant or relevant level. For purposes of clarity, a delay in the onset of a particular symptom is considered one form of reducing the frequency of that symptom.
[0063] Treatment: As used herein, the term "treatment" (also "treat" or "treating") refers to any administration of a therapy that partially or completely alleviates, ameliorates, relieves, inhibits, delays onset of, reduces severity of, and/or reduces incidence of one or more symptoms, features, and/or causes of a particular disease, disorder, and/or condition. In some embodiments, such treatment may be of a subject who does not exhibit signs of the relevant disease, disorder and/or condition and/or of a subject who exhibits only early signs of the disease, disorder, and/or condition. Alternatively or additionally, such treatment may be of a subject who exhibits one or more established signs of the relevant disease, disorder and/or condition. In some embodiments, treatment may be of a subject who has been diagnosed as suffering from the relevant disease, disorder, and/or condition. In some embodiments, treatment may be of a subject known to have one or more susceptibility factors that are statistically correlated with increased risk of development of the relevant disease, disorder, and/or condition.
[0064] Vaccination: As used herein, the term "vaccination" refers to the administration of a composition intended to generate an immune response, for example to a disease-causing agent. For the purposes of the present invention, vaccination can be administered before, during, and/or after exposure to a disease-causing agent, and in some embodiments, before, during, and/or shortly after exposure to the agent. In some embodiments, vaccination includes multiple administrations, appropriately spaced in time, of a vaccinating composition. In some embodiments, vaccination initiates immunization.
DETAILED DESCRIPTION OF CERTAIN EMBODIMENTS
[0065] The present disclosure relates, generally, to novel immunogenic fusion proteins of S. pneumoniae that can be used, e.g., to induce and/or increase an immunoprotective response, or to reduce pneumococcal colonization in subjects at risk of or suffering from pneumococcal infection.
[0066] Two vaccines for S. pneumoniae are currently available in the U.S. PCV13, a 13-valent conjugate vaccine, has been approved for the prevention of invasive pneumococcal disease (IPD) caused by the 13 serotypes contained in the vaccine in children and for the prevention of pneumonia and IPD in adults. In this vaccine, covalent conjugation of saccharides of the 13 pneumococcal serotypes to the CRM197 protein creates saccharide-protein conjugates, which are capable of inducing a T cell-dependent immune response against one or more of the 13 pneumococcal serotypes represented by the saccharides. [PREVNAR 13 prescribing information, 2017]. While infections with S. pneumoniae of multidrug-resistant serotypes contained in PCV13 appeared to decrease after approval of this vaccine, an increase of infections with multidrug-resistant serotypes 35B, 23A, 23B and 15B, which are just a few of the over 84 known serotypes of S. pneumoniae not included in PCV13, was noted. Also, PCV13 was reported to have marginal activity against serotype 3, as its prevalence persists in the population [Richter et al, 2014].
[0067] The second vaccine, PPSV23, is a 23-valent polysaccharide vaccine and is indicated for the prevention of pneumococcal disease in adults greater than 50 years of age, or in persons greater than 2 years of age at increased risk of pneumococcal disease. It is composed of purified capsular polysaccharides from 23 pneumococcal serotypes. While this vaccine has the potential to protect against more serotypes when compared to PCV13, it does not provide protection against the emerging serotypes 35B, 23A and 23B. In addition, PPSV23 elicits a T cell-independent polysaccharide immune response that stimulates mature B-lymphocytes, but not T-lymphocytes. Thus, this vaccine only induces an immune response that is neither long-lasting nor anamnestic upon subsequent challenge. PPSV23 is not effective against colonization. In addition, polysaccharide-type vaccines are not used in infants and children less than 2 years of age, because these children respond poorly to T cell-independent antigens [PNEUMOVAX 23 prescribing information, 2017; CDC, 2010]. Data suggest that PPSV23 may protect adults and the elderly against IPD; however, no consistent effect has been observed in the prevention of pneumonia [Gruber et al, 2008].
[0068] The presently disclosed novel immunogenic proteins represent a substantial advance over the currently available options for immunizing patients against pneumococcal infection. Such immunogenic proteins can be used, e.g., to induce and/or increase an immunoprotective response or to reduce pneumococcal colonization in subjects, such as those at risk of or suffering from pneumococcal infection.
Fusion Proteins
[0069] The present disclosure describes novel immunogenic fusion proteins of S. pneumoniae. In WO2014/124228, the inventors demonstrated that the pneumococcal antigens SP0785 and SP1500 individually elicited a strong IL-17 recall response in re-stimulated human PBMCs and splenocytes of mice that have been exposed to pneumococcus. Immunization of mice with SP0785 plus cholera toxin adjuvant, or with SP1500 plus cholera toxin adjuvant, resulted in significant reduction of pneumococcal colonization (on the order of 100-fold). Immunization with a fusion of SP0785 to the pneumolysoid PdT, or a fusion of SP0785 to the pneumolysoid PdT further conjugated to a polysaccharide of Salmonella typhi, protected 80% of mice from sepsis in a lethal challenge with live S. pneumoniae. Immunization with a fusion of SP0785, SP1500 and the pneumolysoid PdT further conjugated to a polysaccharide of Salmonella typhi, also resulted in significant reduction of pneumococcal colonization (on the order of 10-fold).
[0070] Fusion proteins described and/or utilized herein provide improved immunogenicity and IL-17 response to protein stimulation, as well as further reduction of S. pneumoniae colonization and protection from invasive diseases.
[0071] A fusion protein includes one, two, or more polypeptides that elicit (e.g., primarily elicit) a T cell response, or that elicit both a T cell and a B cell response. In some embodiments, the fusion protein comprises one or more of the polypeptides listed in Table 1. In some embodiments, the fusion protein comprises two of the polypeptides listed in Table 1. In some embodiments, the fusion protein comprises three of the polypeptides listed in Table 1. In some embodiments, the fusion protein comprises one or more of polypeptides encoded by one or more of the genes listed in Table 1. In some embodiments, the fusion protein comprises two of polypeptides encoded by two or more of the genes listed in Table 1. In some embodiments, the fusion protein comprises three polypeptides encoded by three of the genes listed in Table 1.
TABLE-US-00001 TABLE 1 Exemplary Polypeptide Components of Fusion Proteins Protein DNA NCBI SEQ SEQ Accession Locus tag name and description ID No. ID No. No. rhizavidin, full-length 1 9 -- rhizavidin, truncated [aa 45-179] 2 10 n/a (denoted Rhavi) SP0785, full-length (TIGR4 strain) 3 11 ABJ54007.1 SP0785, truncated [aa 33-399] 4 12 n/a SP0785, consensus full-length 5 n/a n/a SP1500, full-length (TIGR4 strain) 6 13 AAK75591.1 SP1500, truncated [aa 27-278] 7 14 n/a SP1500, consensus full-length 8 n/a n/a
[0072] In some embodiments, a fusion protein comprises one or more antigenic polypeptides of S. pneumoniae having an amino acid sequence comprising any of SEQ ID NOs: 3-8, or antigenic fragments thereof. In some embodiments, a fusion protein comprises two antigenic polypeptides having an amino acid sequence comprising any of SEQ ID NOs:3-8, or antigenic fragments thereof. In some embodiments, a fusion protein comprises (i) two antigenic polypeptides having an amino acid sequence comprising any of SEQ ID NOs:3-8, or antigenic fragments thereof, and (i) a biotin-binding moiety comprising SEQ ID NO:1 or 2, or biotin-binding fragments thereof. In some such embodiments, at least one antigenic polypeptide is or comprises an SP0785 polypeptide (e.g., SEQ ID NOs:3-5). In some such embodiments, at least one antigenic polypeptide is or comprises an SP1500 polypeptide (e.g., SEQ ID NOs:6-8).
[0073] In some embodiments, a fusion protein comprises one or more polypeptides homologous to the S. pneumoniae polypeptides listed in Table 1, e.g., an SP0785 polypeptide or an SP1500 polypeptide isolated from different serotypes of S. pneumoniae. Individual serotypes of S. pneumoniae contain numerous mutations relative to each other, and some of these result in different protein sequences between the different serotypes. One of skill in the art may readily substitute an amino acid sequence, or a portion thereof, with the homologous amino acid sequence from a different S. pneumoniae serotype. In some embodiments, antigenic polypeptides have at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.5% identity to the polypeptides listed in Table 1, or antigenic fragments thereof. Serotypic variation may be used to design such variants of the polypeptides listed in Table 1.
[0074] In some embodiments, fusion proteins described herein comprise one or more fragments of polypeptides listed in Table 1, e.g., biotin-binding fragments of rhizavidin, antigenic fragments of a SP0785 polypeptide with or without a signal sequence, or antigenic fragments of a SP1500 polypeptide with or without a signal sequence. In some embodiments, fusion proteins described herein comprise truncation mutants that are close in size to the polypeptides listed in Table 1. For example, they may lack at most one, two, three, four, five, ten, or twenty amino acids from one or both termini (referring to component polypeptides in a fusion protein). In some embodiments, a fragment is a truncated fragment of any of SEQ ID NOs:1-8 lacking 1-5, 1-10, or 1-20 amino acid residues from the N-terminus, C-terminus, or both, of any one of SEQ ID NOs:1-8. In some embodiments, a fragment is a truncated fragment of any of SEQ ID NOs:1-8 lacking 1-10 amino acid residues from the N-terminus, C-terminus, or both, of any one of SEQ ID Nos:1-8. For instance, a fragment may lack 10 amino acid residues at both the N-terminus and C-terminus of any one of SEQ ID NOs:1-8, resulting in a protein lacking 20 amino acid residues. Internal deletions, e.g., of 1-10, 11-20, 21-30, or 31-40 amino acids, are also contemplated.
[0075] In some embodiments, a fusion protein comprises an N-terminal polypeptide and a C-terminal polypeptide. In some embodiments, one or both of the N-terminal polypeptide and the C-terminal polypeptide is an antigenic polypeptide, for example, a polypeptide having an amino acid sequence comprising one or more of SEQ ID NOs:3-8, or an antigenic fragment or variant thereof. In some embodiments, one or both of the N-terminal polypeptide and the C-terminal polypeptide is a biotin-binding moiety, for example a polypeptide having an amino acid sequence comprising SEQ ID NO:1 or 2, or a biotin-binding fragment thereof. In some embodiments, one of the N-terminal polypeptide or the C-terminal polypeptide is a biotin-binding moiety, for example a polypeptide having an amino acid sequence comprising SEQ ID NO:1 or 2, or a biotin-binding fragment thereof, and the other terminal polypeptide is an antigenic polypeptide, for example, a polypeptide having an amino acid sequence comprising one or more of SEQ ID NOs:3-8, or an antigenic fragment or variant thereof.
[0076] In some embodiments, the N-terminal polypeptide and the C-terminal polypeptide are directly bound to each other. In some embodiments, the N-terminal polypeptide and the C-terminal polypeptide are linked via a linker peptide. The length and/or amino acids of a linker, when present, can be adjusted to obtain a more flexible, semi-rigid, or rigid linker. Exemplary flexible peptide linkers are shown as SEQ ID NOs:37-40. A linker can generally be from 1-40, such as 3-10 or 10-30 and specifically 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 amino acids in length. In some embodiments, the fusion protein comprises one linker. In some embodiments, the fusion protein comprises two linkers. In some embodiments, the one or two linkers are selected from SEQ ID NO:37 (GGGGSSS) and SEQ ID NO:38 (AAA). In some embodiments, the fusion protein comprises SEQ ID NO:37 (GGGGSSS) and SEQ ID NO:38 (AAA). In some embodiments, the fusion protein comprises an amino acid sequence AAA (SEQ ID NO: 38) residual from a Not I restriction site. In some embodiments, the fusion protein comprises a linker of SEQ ID NO:37 (GGGGSSS) and an amino acid sequence AAA (SEQ ID NO: 38) residual from a Not I restriction site.
[0077] Exemplary fusion proteins are shown in Table 2.
TABLE-US-00002 TABLE 2 Exemplary Fusion Proteins Protein DNA Locus tag name and description SEQ ID No. SEQ ID No. SP1500-SP0785 17 27 SP0785-SP1500 18 28 Rhavi-SP1500-SP0785 19 29 Rhavi-SP0785-SP1500 20 30 SP1500-SP0785-Rhavi 21 31 SP0785-SP1500-Rhavi 22 32 Rhavi-linker (GGGGSSS)-SP1500- 23 33 linker (AAA)-SP0785 (also denoted CP1) Rhavi-linker (GGGGSSS)-SP0785- 24 34 linker (AAA)-SP1500 SP1500-linker (GGGGSSS)-SP0785- 25 35 linker (AAA)-Rhavi SP0785-linker (GGGGSSS)-SP1500- 26 36 linker (AAA)-Rhavi
[0078] In some embodiments, the present disclosure provides fusion proteins with at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% sequence identity to a fusion protein listed in Table 2. In some embodiments, a fusion protein is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 97%, 98%, 99%, 99.5%, or 100% identity to any one of SEQ ID NOs:17-26. In some embodiments, a fusion protein is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 97%, 98%, 99%, 99.5%, or 100% identity to SEQ ID NO:23. In some embodiments, a fusion protein is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 97%, 98%, 99%, 99.5%, or 100% identity to CP1.
[0079] In some embodiments, a fusion protein described herein comprises an antigenic fragment of a fusion protein shown in Table 2. In some embodiments, a fusion protein is or includes an antigenic fragment of any of SEQ ID NOs:17-26. For example, a fusion protein may lack at most one, two three, four, five, ten, or twenty amino acids from the N-terminus, C-terminus, or both, of any one of SEQ ID NOs:17-26. In some embodiments, the same number of residues is removed from the N-terminus and the C-terminus, while in other embodiments, a different number of residues is removed from the N-terminus compared to the C-terminus. In some embodiments, a fusion protein is or includes an antigenic fragment of SEQ ID NO:23. In some embodiments, a fusion protein is or includes an antigenic fragment of CP1.
[0080] In some embodiments, a fusion protein described herein comprises a biotin-binding moiety. In some embodiments, the fusion protein comprises a biotin-binding moiety, and one or more polypeptide antigens. In some embodiments, the fusion protein comprises a biotin-binding moiety and two or more polypeptide antigens. As used herein, a "biotin-binding moiety" refers to a biotin-binding polypeptide or protein, a biotin-binding fragment thereof, or a biotin-binding domain thereof. In some embodiments, the biotin-binding moiety of the fusion protein comprises rhizavidin or a biotin-binding fragment thereof, as further described in WO 2012/155053, the contents of which are herein incorporated by reference in their entirety.
[0081] In some embodiments, a fusion protein described herein comprises a biotin-binding moiety that is or comprises a polypeptide having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:1 (rhizavidin), or biotin-binding fragment thereof. In some embodiments, the fusion protein comprises a biotin-binding moiety that is or comprises a polypeptide having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:2 (amino acids 45-179 of rhizavidin, denoted Rhavi), or biotin-binding fragment thereof. In some embodiments, the fusion protein comprises a polypeptide comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:4 (amino acids 33-399 of S. pneumoniae SP0785 polypeptide), or an antigenic fragment thereof. In some embodiments, the fusion protein comprises a polypeptide comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:7 (amino acids 27-278 of S. pneumoniae SP1500 polypeptide), or an antigenic fragment thereof.
[0082] In some embodiments, a fusion protein described herein comprises each of: (a) a biotin-binding moiety that is or comprises a polypeptide having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:1 (rhizavidin), or biotin-binding fragment thereof; (b) a polypeptide comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:4 (amino acids 33-399 of S. pneumoniae SP0785 polypeptide), or an antigenic fragment thereof and (c) a polypeptide comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:7 (amino acids 27-278 of S. pneumoniae SP1500 polypeptide) or an antigenic fragment thereof. In some embodiments, the fusion protein further comprises one or more linkers. In some embodiments, the one or more linkers are selected from SEQ ID NO:37 (GGGGSSS) and SEQ ID NO:38 (AAA). In some embodiments, the fusion protein comprises an amino acid sequence AAA (SEQ ID NO: 38) residual from a Not I restriction site. In some embodiments, the fusion protein comprises a linker of SEQ ID NO:37 (GGGGSSS) and an amino acid sequence AAA (SEQ ID NO: 38) residual from a Not I restriction site.
[0083] In some embodiments, a fusion protein described herein comprises each of: (a) a biotin-binding moiety that is or comprises a polypeptide having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:2 (amino acids 45-179 of rhizavidin, denoted Rhavi), or biotin-binding fragment thereof; (b) a polypeptide comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:4 (amino acids 33-399 of S. pneumoniae SP0785 polypeptide) or an antigenic fragment thereof; and (c) a polypeptide comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:5 (amino acids 27-278 of S. pneumoniae SP1500 polypeptide) or an antigenic fragment thereof. In some embodiments, the fusion protein further comprises one or more linkers. In some embodiments, the one or more linkers are selected from SEQ ID NO:37 (GGGGSSS) and SEQ ID NO:38 (AAA). In some embodiments, the fusion protein comprises an amino acid sequence AAA (SEQ ID NO: 38) residual from a Not I restriction site. In some embodiments, the fusion protein comprises a linker of SEQ ID NO:37 (GGGGSSS) and an amino acid sequence AAA (SEQ ID NO: 38) residual from a Not I restriction site. In some embodiments, a fusion protein described herein comprises an amino acid sequence having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence SEQ ID NO:23. In some embodiments, the fusion protein comprises the amino acid sequence SEQ ID NO:23. In some embodiments, the fusion protein consists of the amino acid sequence SEQ ID NO:23 (CP1).
[0084] In some embodiments, a fusion protein described herein includes a variant or fragment of a polypeptide listed in Table 1. In some embodiments, a fusion protein described herein includes a polypeptide encoded by a variant or fragment of a gene listed in Table 1. In some embodiments, a fragment included in a fusion protein described herein is close in size to a full-length polypeptide or a polypeptide listed in Table 1. For example, they may lack at most one, two, three, four, five, ten, twenty, or thirty amino acids from one or both termini. In some embodiments, the fragment is 25-50 amino acids in length, or 50-100, or 100-150, or 150-200, or 200-250, or 250-300, or 300-350 amino acids in length. In some embodiments, the fragments result from processing, or partial processing, of signal sequences by an expression host, e.g. E. coli, an insect cell line (e.g., the baculovirus expression system), or a mammalian (e.g., human or Chinese Hamster Ovary) cell line. The fragments described above or sub-fragments thereof (e.g., fragments of 8-50, 8-30, or 8-20 amino acid residues) preferably have one of the biological activities described below, such as increasing the amount of IL-17 released by at least 1.5 fold or 2 fold or more (e.g., either as an absolute measure or relative to a control protein).
[0085] The DNA and protein sequence of each gene and polypeptide may be identified by searching for the Locus Tag in a publicly available database, e.g., Entrez Gene (on the NCBI NIH web site on the World Wide Web, at www.ncbi.nlm.nih. gov/sites/entrez?db=gene), in the Streptococcus pneumoniae TIGR4 genome, and the indicated sequences are also included within the scope of the present disclosure.
[0086] Certain polypeptides of Table 1, variants thereof, and additional exemplary polypeptides and linkers which constitute components of various embodiments of the fusion proteins are described in greater detail below.
[0087] SP0785 Polypeptides (e.g., SEQ ID NOs:3-5) and Variants Thereof
[0088] SP0785 is a conserved hypothetical S. pneumoniae protein described in WO 2014/124228. In some embodiments, a SP0785 polypeptide is an efflux transporter protein conserved across S. pneumoniae strains. In some embodiments, a SP0785 polypeptide is or comprises a full-length SP0785 polypeptide. For example, in some embodiments, a full-length SP0785 polypeptide has 399 amino acids (38 kDa) and is represented by the amino acid sequence as set forth in SEQ ID NO: 3. Amino acids 1-32 of SEQ ID NO:3 are predicted to be a signal sequence and transmembrane domain of a SP0785 polypeptide (amino acids 1-32 of the full-length protein). In some embodiments, a fusion protein comprises a SP0785 polypeptide of S. pneumoniae. In some embodiments, a fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 350, 367, or 399 consecutive amino acids of a SP0785 polypeptide.
[0089] In some embodiments, a SP0785 polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 350, 367, or 399 consecutive amino acids of the sequence shown in SEQ ID NO:3 [full-length]. In some embodiments, a SP0785 polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 350, 367, or 399 consecutive amino acids of the sequence shown in SEQ ID NO:3 [full-length].
[0090] In some embodiments, a SP0785 polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 350, or 367 consecutive amino acids of the sequence shown in SEQ ID NO:4 [minus signal sequence and transmembrane domain]. In some embodiments, a SP0785 polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 350, or 367 consecutive amino acids of the sequence shown in SEQ ID NO:4 [minus signal sequence and transmembrane domain].
[0091] Sequence variation occurs at the protein level between different S. pneumoniae serotypes, and a consensus sequence illustrating combinations of SP785 sequences from different S. pneumoniae serotypes is provided as SEQ ID NO:5. Accordingly, in some embodiments, the fusion protein includes a polypeptide having an amino acid sequence comprising, or consisting of, SEQ ID NO:5, or an antigenic fragment thereof (e.g., in place of a polypeptide having an amino acid sequence comprising one of SEQ ID NO:3 or 4). In some embodiments, a SP0785 polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 350, or 367 consecutive amino acids of the sequence shown in SEQ ID NO:5 [consensus]. In some embodiments, a SP0785 polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 350, or 367 consecutive amino acids of the sequence shown in SEQ ID NO:5 [consensus].
[0092] An exemplary nucleotide sequence encoding a SP0785 polypeptide is provided herein as SEQ ID NO:11.
[0093] SP1500 Polypeptides (e.g., SEQ ID NOs:6-8) and Variants Thereof
[0094] SP1500 is described in WO 2014/124228. In some embodiments, a SP1500 polypeptide is an Amino Acid ABC Transporter, amino acid-binding polypeptide conserved across S. pneumoniae strains. In some embodiments, a SP1500 polypeptide is or comprises a full-length SP1500 polypeptide. For example, in some embodiments, a full-length SP1500 polypeptide has 278 amino acids (28 kDa) and is represented by the amino acid sequence as set forth in SEQ ID NO: 6. Amino acids 1-26 of SEQ ID NO:6 are predicted to be a signal sequence of a SP1500 polypeptide (amino acids 1-26 of the full-length protein). In some embodiments, a fusion protein comprises a SP1500 polypeptide of S. pneumoniae. In some embodiments, a fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 252, or 278 consecutive amino acids of a SP1500 polypeptide.
[0095] In some embodiments, a SP1500 polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 252, or 278 consecutive amino acids of the sequence shown in SEQ ID NO:6 [full-length]. In some embodiments, a SP1500 polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, 252, or 278 consecutive amino acids of the sequence shown in SEQ ID NO:6 [full-length].
[0096] In some embodiments, a SP1500 polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, or 252 consecutive amino acids of the sequence shown in SEQ ID NO:7 [minus signal sequence]. In some embodiments, a SP1500 polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, or 252 consecutive amino acids of the sequence shown in SEQ ID NO:7 [minus signal sequence].
[0097] Sequence variation occurs at the protein level between different S. pneumoniae serotypes, and a consensus sequence illustrating combinations of SP1500 sequences from different S. pneumoniae serotypes is provided as SEQ ID NO:8. In some embodiments, the fusion protein includes a polypeptide having an amino acid sequence comprising, or consisting of, SEQ ID NO:8, or an antigenic fragment thereof (e.g., in place of a polypeptide having an amino acid sequence comprising one of SEQ ID NO:6 or 7). In some embodiments, a SP1500 polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, or 252 consecutive amino acids of the sequence shown in SEQ ID NO:8 [consensus]. In some embodiments, a SP1500 polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, 200, or 252 consecutive amino acids of the sequence shown in SEQ ID NO:8 [consensus].
[0098] An exemplary nucleotide sequence encoding a SP1500 polypeptide is provided herein as SEQ ID NO:14.
Rhizavidin
[0099] In some embodiments, a fusion protein described herein is a component of non-covalent Multiple Antigen Presenting System (MAPS) immunogenic complexes. In some embodiments, MAPS complexes utilize the high affinity (dissociation constant [K.sub.D].apprxeq.10.sup.-15M) noncovalent binding between biotin, or biotin derivatives, and rhizavidin, a biotin-binding protein that has no significant predicted homology with human proteins.
[0100] Rhizavidin is a naturally occurring dimeric protein in the avidin protein family, was first discovered in Rhizobium etli, a symbiotic bacterium of the common bean. Rhizavidin has only a 22% amino acid identity with chicken avidin, a protein commonly found in eggs, but with high conservation of amino acid residues involved in biotin binding [Helppolainen et al, 2007]. In some embodiments, the nucleotide sequence of rhizavidin is set forth in SEQ ID NO:9. In some embodiments, the amino acid sequence of rhizavidin is set forth in SEQ ID NO:1. Amino acids 1-44 of SEQ ID NO:1 are predicted to be a signal sequence(s) of rhizavidin (amino acids 1-44 of the full-length protein). In some embodiments, a fusion protein comprises rhizavidin. In some embodiments, a fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, or 179 consecutive amino acids of a rhizavidin polypeptide.
[0101] In some embodiments, a rhizavidin polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, or 179 consecutive amino acids of the sequence shown in SEQ ID NO:1 [full-length]. In some embodiments, a rhizavidin polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, 150, or 179 consecutive amino acids of the sequence shown in SEQ ID NO:1 [full-length].
[0102] In some embodiments, a rhizavidin polypeptide of the fusion protein comprises at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, or 135 consecutive amino acids of the sequence shown in SEQ ID NO:2 [minus signal sequence]. In some embodiments, a rhizavidin polypeptide of the fusion protein comprises an amino acid sequence that is at least 60% or more (including, e.g., at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%) identical to at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 45, 50, 60, 65, 70, 75, 80, 85, 90, 95, 100, or 135 consecutive amino acids of the sequence shown in SEQ ID NO:2 [minus signal sequence].
[0103] In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 50-179 of SEQ ID NO:1. In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 55-179 of SEQ ID NO:1. In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 60-179 of SEQ ID NO:1. In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 65-179 of SEQ ID NO:1.
[0104] In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 45-175 of SEQ ID NO:1. In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 45-171 of SEQ ID NO:1. In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 45-167 of SEQ ID NO:1. In some embodiments, a rhizavidin polypeptide of the fusion protein is or comprises amino acids 45-163 of SEQ ID NO:1.
[0105] Linker or Spacer
[0106] In some embodiments, a fusion protein comprises one or more linkers. In some embodiments, a linker is or comprises one or more amino acids. In some embodiments, a fusion protein comprises an antigenic polypeptide joined to a biotin-binding moiety by a linker. In some embodiments, a fusion protein comprises a first antigenic polypeptide, a second antigenic polypeptide, a biotin-binding moiety, and at least one linker. In some embodiments, the first antigenic polypeptide and the second antigenic polypeptide are joined by a linker. In some embodiments, the first antigenic polypeptide or the second antigenic polypeptide are joined to the biotin-binding moiety by a linker. In some embodiments, the first antigenic polypeptide and the second antigenic polypeptide are joined by a first linker; and the first antigenic polypeptide or the second antigenic polypeptide are joined to the biotin-binding moiety by a second linker.
[0107] In some embodiments, a linker interposes a structure between two protein moieties. In some embodiments, the structure is or comprises an .alpha.-helix. In some embodiments the structure is or comprises a .beta.-strand. In some embodiments, the structure is or comprises a coil/bend. In some embodiments, the structure is or comprises a turn. In some embodiments, a linker decreases steric hindrance between two protein moieties joined by the linker. In some embodiments, a linker decreases unfavorable interactions between two protein moieties joined by the linker. In some embodiments, a linker comprises a mixture of glycine and serine residues. In some embodiments, the linker may additionally comprise threonine, proline, and/or alanine residues. In some embodiment a linker is hydrophilic. In some embodiments a linker is hydrophobic. In some embodiments a linker increases the stability of the fusion protein containing the linker.
[0108] In some embodiments, a linker does not interfere with the folding of an antigenic polypeptide to which it is joined. In some embodiments, a linker does not interfere with the antigenicity of an antigenic polypeptide to which it is joined. In some embodiments, a linker does not reduce the antigenicity of an antigenic polypeptide to which it is joined. In some embodiments, a linker does not eliminate the antigenicity of an antigenic polypeptide to which it is joined. In some embodiments the effect of the linker is determined by comparing the polypeptide with the polypeptide joined to the linker.
[0109] In some embodiments, a linker does not interfere with the folding of a biotin-binding moiety to which it is joined. In some embodiments, a linker does not interfere with the biotin-binding ability of a biotin-binding moiety to which it is joined. In some embodiments, a linker does not reduce the biotin-binding ability of a biotin-binding moiety to which it is joined. In some embodiments, a linker does not eliminate the biotin-binding ability of a biotin-binding moiety to which it is joined. In some embodiments the effect of the linker is determined by comparing the biotin-binding moiety with the biotin-binding moiety joined to the linker.
[0110] In some embodiments, a linker is not antigenic. In some embodiments, a linker does not elicit a T cell response. In some embodiments, a linker does not elicit a B cell response. In some embodiments, a linker does not induce a T cell or a B cell response.
[0111] In some embodiments, a linker comprises two or more amino acids. In some embodiments, a linker may be 3-100, 5-100, 10-100, 20-100 30-100, 40-100, 50-100, 60-100, 70-100, 80-100, 90-100, 5-55, 10-50, 10-45, 10-40, 10-35, 10-30, 10-25, 10-20, 10-15, 3-10, 3-9, 3-8, 3-7, 3-6, 3-5, 3-4, or 2-3 amino acids in length. In some embodiments, a linker comprises between 10-100, 10-90, 10-80, 10-70, 10-60, 10-50, 10-40, 10-30, 10-20, 10-15 amino acids. In some embodiments, the linker comprises at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, 90, or 95 amino acids. In some embodiments, a linker is or comprises 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, 90, 95, or 100 amino acids.
[0112] In some embodiments, a linker is a flexible linker. Flexible linkers may be useful for joining domains that require a certain degree of movement or interaction and may include small, non-polar (e.g. Gly) or polar (e.g. Ser or Thr) amino acids. Incorporation of Ser or Thr can also maintain the stability of the linker in aqueous solutions by forming hydrogen bonds with the water molecules, and therefore reduce unfavorable interactions between the linker and the protein moieties. In some embodiments a linker comprises small non-polar (e.g. Gly) or polar (e.g. Ser or Thr) amino acids. In some embodiments, a linker is a Gly-Ser linker.
[0113] In some embodiments, a linker is or comprises an amino acid sequence of GGGGSSS (SEQ ID NO:37). In some embodiments, a linker is or comprises a sequence of (GGGGS).sub.n (SEQ ID NO:39), where n represents the number of repeating GGGGS (SEQ ID NO: 62) units and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments, a polypeptide linker may have an amino acid sequence that is or comprises GGGGSGGGGSGGGGS (SEQ ID NO:41) (i.e., (GGGGS).sub.3 (SEQ ID NO: 41)) or GGGGSGGGGSGGGGSGGGGSGGGGSGGGGS (SEQ ID NO:42) (i.e., (GGGGS).sub.6 (SEQ ID NO: 42)). In some embodiments, a linker comprises one or more of Gly, Ser, Thr, Ala, Lys, and Glu. In some embodiments, a linker is or comprises KESGSVSSEQLAQFRSLD (SEQ ID NO:43). In some embodiments, a linker is or comprises EGKSSGSGSESKST (SEQ ID NO:44). In some embodiments, a linker is or comprises (Gly).sub.n (SEQ ID NO:45) where n represents the number of repeating Gly residues and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments a linker is or comprises GGG. In some embodiments, a linker is or comprises (Gly).sub.6 (SEQ ID NO:40). In some embodiments, a linker is or comprises (Gly).sub.8 (SEQ ID NO:46). In some embodiments, a linker is or comprises GSAGSAAGSGEF (SEQ ID NO:47). In some embodiments, a linker is or comprises an amino acid sequence of AAA (SEQ ID NO:38).
[0114] In some embodiments, a linker is a rigid linker. Rigid linkers are useful to keep a fixed distance between domains and to maintain their independent functions. Rigid linkers may also be useful when a spatial separation of the domains is critical to preserve the stability or bioactivity of one or more components in the fusion. In some embodiments, a linker is or comprises (EAAAK).sub.n (SEQ ID NO:48) where n represents the number of repeating EAAAK (SEQ ID NO: 63) units and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments, a linker is or comprises A(EAAAK).sub.nA, (SEQ ID NO:49) where n represents the number of repeating EAAAK (SEQ ID NO: 63) units and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments, a linker is or comprises A(EAAAK).sub.nA (SEQ ID NO: 108), where n represents the number of repeating EAAAK (SEQ ID NO: 63) units and is 2, 3, 4, or 5. In some embodiments, a linker is or comprises A(EAAAK).sub.4ALEA(EAAAK).sub.4A (SEQ ID NO:50). In some embodiments, a linker is or comprises [A(EAAAK).sub.nA].sub.m, (SEQ ID NO:51) wherein n is 2, 3, or 4 and m is 1 or 2. In some embodiments, a linker is or comprises AEAAAKEAAAKA (SEQ ID NO:52).
[0115] In some embodiments a linker is or comprises (X-Pro).sub.n (SEQ ID NO:53), with X designating any amino acid, where n represents the number of repeating X-Pro units and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments a linker is or comprises (Ala-Pro).sub.n (SEQ ID NO:54), where n represents the number of repeating Ala-Pro units and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments a linker is or comprises (Ala-Pro).sub.n (SEQ ID NO: 109), where n represents the number of repeating Ala-Pro units and is 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, or 17.
[0116] In some embodiments a linker is or comprises (Lys-Pro).sub.n (SEQ ID NO:55), where n represents the number of repeating Lys-Pro units and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments a linker is or comprises (Gln-Pro).sub.n (SEQ ID NO:56), where n represents the number of repeating Gln-Pro units and is 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30 or more. In some embodiments, a linker is or comprises (Ala-Pro).sub.7 (SEQ ID NO:57).
[0117] In some embodiments a linker is or comprises GAPGGGGGAAAAAGGGGGGAP (GAG linker, SEQ ID NO:58). In some embodiments a linker is or comprises GAPGGGGGAAAAAGGGGGGAPGGGGGAAAAAGGGGGGAP (GAG2 linker, SEQ ID NO:59). In some embodiments a linker is or comprises GAPGGGGGAAAAAGGGGGGAPGGGGGAAAAAGGGGGGAPGGGGGAAAAAGGGGG GAP (GAG3 linker, SEQ ID NO:60).
[0118] Suitable linkers or spacers also include those having an amino acid sequence at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more homologous or identical to the above exemplary linkers.
[0119] Additional linkers suitable for use with some embodiments may be found in U.S. Patent Publication No. 2012/0232021, filed on Mar. 2, 2012, and [Chen, 2013] the disclosures of which is hereby incorporated by reference in their entireties.
Tagged Fusion Proteins
[0120] In some embodiments, a fusion protein described herein may comprise a tag. A tag may be N-terminal or C-terminal. For instance, tags may be added to a polypeptide (via additions or modifications on the encoding DNA sequence) to facilitate purification, detection, solubility, or confer other desirable characteristics on the protein. In some embodiments a tag may be a peptide, oligopeptide, or polypeptide that may be used in affinity purification. In some embodiments, a tag is, comprises, or is derived from one or more of polyhistidine (His), Glutathione S-transferase (GST), tandem affinity purification (TAP), FLAG, myc, human influenza hemagglutinin (HA), maltose binding protein (MBP), vesicular Stomatitis viral glycoprotein (VSV-G), thioredoxin, V5, avidin, streptavidin, biotin carboxyl carrier protein (BCCP), Calmodulin, Nus, S tags, lipoprotein D, and galactosidase. In some embodiments, a His tag is or comprises an amino acid sequence of Hn, wherein n is an integer between 2 and 10 (SEQ ID NO: 64). Exemplary His tags include HHHHHH (SEQ ID NO:15) and MSYYHHHHHH (SEQ ID NO:16). In other embodiments, the fusion protein is free of tags such as protein purification tags, and is purified by a method not relying on affinity for a purification tag. In some embodiments, the fusion protein comprises no more than 1, 2, 3, 4, 5, 10, or 20 additional amino acids on one or both termini of a polypeptide of Table 1 or fusion protein of Table 2.
[0121] In some embodiments, a fusion protein described herein may contain a membrane translocating sequence (MTS), to facilitate introduction of the fusion protein into a mammalian cell and subsequent stimulation of the cell-mediated immune response. Exemplary membrane translocating sequences include the hydrophobic region in the signal sequence of Kaposi fibroblast growth factor, the MTS of a synuclein, the third helix of the Antennapedia homeodomain, SN50, integrin 3 h-region, HIV Tat, pAntp, PR-39, abaecin, apidaecin, Bac5, Bac7, P. berghei CS protein, and those MTSs described in U.S. Pat. Nos. 6,248,558, 6,432,680 and 6,248,558.
Nucleic Acids
[0122] In some embodiments, the present disclosure provides nucleic acids, e.g., DNA, RNA, or analogs thereof, encoding one or more of the polypeptides and/or fusion proteins described herein. An underlying DNA sequence for the polypeptides described herein may be modified in ways that do not affect the sequence of the protein product, and such sequences are included in the invention. In some embodiments, a DNA sequence may be codon-optimized to improve expression in a host such as a bacterial cell line, e.g., E. coli, an insect cell line (e.g., using the baculovirus expression system), or a mammalian (e.g., human or Chinese Hamster Ovary) cell line.
[0123] In some embodiments, the present disclosure provides nucleic acids, e.g., DNA, RNA, or analogs thereof, that are at least 70%, 80%, 85%, 90%, 95%, 97%, 98%, 99%, 99.5%, or 100% identical to a nucleic acid sequence provide in Table 1, Table 2, or a variant or portion thereof. In some embodiments, the nucleic acid is 600-2000, 800-1800, 1000-1600, 1200-1400 nucleotides in length. In some embodiments, the nucleic acid is 600-1600, 800-1800, 1000-2000, 2000-3000, or 3000-4000 nucleotides in length. In some embodiments, a nucleic acid may be used for recombinant production of a polypeptide or fusion protein of Table 1 or Table 2, or antigenic fragments thereof. In some embodiments, a nucleic acid may be used as a vaccine.
[0124] Nucleic acid sequences encoding variants of SP0785 (SEQ ID NOs:3 and 4) are provided as SEQ ID NOs:11 and 12. Nucleic acid sequences encoding variants of SP1500 (SEQ ID NOs:6 and 7) are provided as SEQ ID NOs:13 and 14. Nucleic acid sequences encoding different fusion proteins (SEQ ID NOs:17-26) are provided as SEQ ID NOs:27-36. In all cases, due to degeneracy in the genetic code, other DNA sequences (including multiple codon-optimized sequences) could be contemplated by those of ordinary skill to encode such polypeptides and fusion proteins.
[0125] Nucleic acids encoding polypeptides or fusion proteins of Table 1 or Table 2, or fragments thereof, can be cloned into any of a variety of expression vectors, under the control of a variety of regulatory elements, and fusions can be created with other sequences of interest. Methods of cloning nucleic acids are routine and conventional in the art. For general references describing methods of molecular biology which are mentioned in this application, e.g., isolating, cloning, modifying, labeling, manipulating, sequencing and otherwise treating or analyzing nucleic acids and/or proteins, see, e.g., Sambrook et al, 1989; Ausubel et al, 1995; Davis et al, 1986; Hames et al, 1985; Dracopoli et al, 2018; and Coligan et al, 2018.
Uses of Fusion Proteins
[0126] In some embodiments, a fusion protein described herein does not have, or has minimal, hemolytic activity. For example, in some embodiments, the hemolytic activity of a fusion protein described herein can be established by turbidimetry (OD.sub.420) after incubation of the fusion protein at different dilutions with red blood cells (e.g., sheep erythrocytes), to determine the protein concentration at which 50% of the red blood cells are lysed. In some such embodiments, the hemolytic activity of a fusion protein described herein can be characterized by an OD.sub.420 of less than 0.4 or lower, including, e.g., less than 0.3, less than 0.25, less than 0.2, or lower, for a given protein concentration.
[0127] In some embodiments, polypeptides of S. pneumoniae and fusion proteins described herein, and fragments and variants thereof, are immunogenic. These polypeptides and fusion proteins may be immunogenic in mammals, for example mice, rats, guinea pigs, or humans. An antigenic polypeptide or fusion protein is typically one capable of raising a significant immune response in an assay or in a subject. The immune response may be innate, humoral, cell-mediated, or mucosal (combining elements of innate, humoral and cell-mediated immunity). For instance, an antigenic polypeptide or fusion protein may increase the amount of IL-17 produced by T cells. Alternatively or additionally, an antigenic polypeptide or fusion protein may (i) induce production of antibodies, e.g., neutralizing antibodies, that bind to the polypeptide and/or the whole bacteria, (ii) induce Th17 immunity, (iii) activate the CD4+ T cell response, for example by increasing the number of CD4+ T cells and/or increasing localization of CD4+ T cells to the site of infection or reinfection, (iv) activate the CD8+ T cell response, for example by increasing the number of CD8+ T cells and/or increasing localization of CD8+ T cells to the site of infection or reinfection, (v) activate both the CD4+ and the CD8+ response, (vi) activate CD4-/CD8-immunity, (vii) induce Th1 immunity, (viii) induce anti-microbial peptides, (ix) activate innate immunity, or any combination of the foregoing. In some embodiments, an antigenic polypeptide or fusion protein elicits production of a detectable amount of antibody specific to that antigen.
[0128] In some embodiments, a fusion protein described herein is an antigen or has antigenic properties. In some embodiments, a fusion protein described herein is a carrier protein or has carrier properties. In some embodiments, a fusion protein described herein is both an antigen and a carrier protein. In some embodiments, a fusion protein described herein has both carrier properties and antigenic properties.
[0129] In some embodiments, a fusion protein described herein is an antigen of an immunogenic complex (e.g., a Multiple Antigen Presenting System (MAPS) complex as described in WO 2012/155007, the entire contents of which are incorporated herein by reference for the purposes indicated herein). In some embodiments, a fusion protein described herein is a carrier protein of an immunogenic complex. In some embodiments, a fusion protein described herein is both a carrier protein and an antigen of an immunogenic complex.
[0130] In some embodiments, polypeptides of the fusion proteins described herein have less than 20%, 30%, 40%, 50%, 60% or 70% identity to human auto-antigens and/or gut commensal bacteria (e.g., certain Bacteroides, Clostridium, Fusobacterium, Eubacterium, Ruminococcus, Peptococcus, Peptostreptococcus, Bifidobacterium, Escherichia, and Lactobacillus species). Examples of human autoantigens include insulin, proliferating cell nuclear antigen, cytochrome P450, and myelin basic protein.
[0131] A polypeptide included in a fusion protein described herein may comprise one or more immunogenic portions and one or more non-immunogenic portions. The immunogenic portions may be identified by various methods, including protein microarrays, ELISPOT/ELISA techniques, and/or specific assays on different deletion mutants (e.g., fragments) of the polypeptide in question. Immunogenic portions may also be identified by computer algorithms. Some such algorithms, like EpiMatrix (produced by EpiVax), use a computational matrix approach. Other computational tools for identifying antigenic epitopes include PEPVAC (Promiscuous EPitope-based VACcine, hosted by Dana Farber Cancer Institute on the world wide web at immunax.dfci.harvard.edu/PEPVAC), MHCPred (which uses a partial least squares approach and is hosted by The Jenner Institute on the world wide web at www.jenner. ac.uk/MHCPred), and Immune Epitope Database algorithms on the World Wide Web at tools.immuneepitope.org. An antigenic fragment of a polypeptide described herein comprises at least one immunogenic portion, as measured experimentally or identified by algorithm (for example, the SYFPEITHI algorithm found at www.syfpeithi.de).
[0132] Representative predicted epitopes of SP0785 and SP1500 are presented in Table 3 and Table 4, respectively.
TABLE-US-00003 TABLE 3 Top 20 of 353 total predicted MHC II binding sites (HLA-DRB1*0101) of an exemplary SP0785 polypeptide SEQ Per- ID centile Allele # Start End NO: Sequence Method Used Rank HLA- 1 253 267 68 SPAAGNNTGSKYPYT Consensus 93.69 DRB1*01:01 (CombLib., SMM, NN) HLA- 1 302 316 69 VMDDSKNYVWIVDEQ Consensus 91.95 DRB1*01:01 (comblib., smm, nn) HLA- 1 252 266 70 ASPAAGNNTGSKYPY Consensus 90.60 DRB1*01:01 (comblib., smm, nn) HLA- 1 251 265 71 AASPAAGNNTGSKYP Consensus 90.11 DRB1*01:01 (comblib., smm, nn) HLA- 1 325 339 72 SLGNADAENQEITSG Consensus 89.98 DRB1*01:01 (comblib., smm, nn) HLA- 1 224 238 73 FTSKVYPDKKWTGKL Consensus 89.54 DRB1*01:01 (comblib., smm, nn) HLA- 1 268 282 74 IDVTGEVGDLKQGFS Consensus 89.54 DRB1*01:01 (comblib., smm, nn) HLA- 1 352 366 75 SSLEEGKEVKADEAT Consensus 89.54 DRB1*01:01 (comblib., smm, nn) HLA- 1 353 367 76 SLEEGKEVKADEATN Consensus 89.54 DRB1*01:01 (comblib., smm, nn) HLA- 1 267 281 77 TIDVTGEVGDLKQGF Consensus 89.24 DRB1*01:01 (comblib., smm, nn) HLA- 1 255 269 78 AAGNNTGSKYPYTID Consensus 89.19 DRB1*01:01 (comblib., smm, nn) HLA- 1 110 124 79 PQLPAPVGGEDATVQ Consensus 89.05 DRB1*01:01 (comblib., smm, nn) HLA- 1 111 125 80 QLPAPVGGEDATVQS Consensus 89.05 DRB1*01:01 (comblib., smm, nn) HLA- 1 112 126 81 LPAPVGGEDATVQSP Consensus 89.05 DRB1*01:01 (comblib., smm, nn) HLA- 1 113 127 82 PAPVGGEDATVQSPT Consensus 89.05 DRB1*01:01 (comblib., smm, nn) HLA- 1 114 128 83 APVGGEDATVQSPTP Consensus 89.05 DRB1*01:01 (comblib., smm, nn) HLA- 1 254 268 84 PAAGNNTGSKYPYTI Consensus 88.73 DRB1*01:01 (comblib., smm, nn) HLA- 1 256 270 85 AGNNTGSKYPYTIDV Consensus 88.21 DRB1*01:01 (comblib., smm, nn) HLA- 1 225 239 86 TSKVYPDKKWTGKLS Consensus 87.42 DRB1*01:01 (comblib., smm, nn) HLA- 1 223 237 87 SFTSKVYPDKKWTGK Consensus 86.32 DRB1*01:01 (comblib., smm, nn)
TABLE-US-00004 TABLE 4 Top 20 of 239 total predicted MHC II binding sites (HLA-DRB1*0101) of an exemplary SP1500 polypeptide SEQ Per- ID centile Allele # Start End NO: Sequence Method Used Rank HLA- 1 125 139 88 AQAGSSGYADFEANP Consensus 93.70 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 54 68 89 TVNWQPIDWDLKEAE Consensus 90.32 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 146 160 90 VANKEANQYQTFNEA Consensus 90.02 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 53 67 91 ITVNWQPIDWDLKEA Consensus 89.97 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 52 66 92 GITVNWQPIDWDLKE Consensus 89.54 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 55 69 93 VNWQPIDWDLKEAEL Consensus 89.54 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 228 242 94 KDGKFQEISQKWFGE Consensus 89.54 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 229 243 95 DGKFQEISQKWFGED Consensus 89.54 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 230 244 96 GKFQEISQKWFGEDV Consensus 89.54 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 231 245 97 KFQEISQKWFGEDVA Consensus 89.54 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 239 253 98 WFGEDVATKEVKEGQ Consensus 89.54 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 124 138 99 GAQAGSSGYADFEAN Consensus 89.06 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 227 241 100 YKDGKFQEISQKWFG Consensus 88.56 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 220 234 101 NEAFSSLYKDGKFQE Consensus 87.65 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 51 65 102 YGITVNWQPIDWDLK Consensus 85.94 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 78 92 103 WNGYSATDERREKVA Consensus 85.92 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 208 222 104 ARKEDTNLVKKINEA Consensus 85.34 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 82 96 105 SATDERREKVAFSNS Consensus 84.78 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 79 93 106 NGYSATDERREKVAF Consensus 84.52 DRB1*01:01 (comb.lib./smm/nn) HLA- 1 80 94 107 GYSATDERREKVAFS Consensus 84.52 DRB1*01:01 (comb.lib./smm/nn)
Immunogenic and Vaccine Compositions
[0133] The present disclosure also provides immunogenic compositions (e.g., vaccine compositions) of, or comprising, one or more fusion proteins described herein. In some embodiments, the immunogenic composition comprises one or more fusion proteins with at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% sequence identity to a fusion protein listed in Table 2. In some embodiments, the immunogenic composition comprises a fusion protein that is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 97%, 98%, 99%, 99.5%, or 100% identity to any one of SEQ ID NOs:17-26. In some embodiments, the immunogenic composition comprises a fusion protein that is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 97%, 98%, 99%, 99.5%, or 100% identity to SEQ ID NO:23. In some embodiments, the immunogenic composition comprises a fusion protein that is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 97%, 98%, 99%, 99.5%, or 100% identity to CP1 (e.g., as described in FIG. 1).
[0134] In some embodiments, an immunogenic composition may also comprise portions of fusion proteins described herein, for example internal deletion mutants, truncation mutants, and fragments. In some embodiments, the portions of said fusion proteins are immunogenic. The immunogenicity of a portion of a fusion protein is readily determined using the same assays that are used to determine immunogenicity of the full-length fusion protein. In some embodiments, the portion of the fusion protein has substantially the same immunogenicity as the full-length fusion protein. In some embodiments, the immunogenicity is no less than 10%, 20%, 30%, 40%, or 50% that of the fusion proteins of Table 2.
Multi-Component Immunogenic and Vaccine Compositions
[0135] In some embodiments, an immunogenic composition described herein (e.g., a vaccine composition) includes a fusion protein described herein and additionally one or more, or two or more, known S. pneumoniae antigens. In some instances, the known S. pneumoniae antigens are predominantly antibody targets. In some instances, the known S. pneumoniae antigens are polysaccharides. In some instances, the known S. pneumoniae antigens protect from S. pneumoniae colonization, or from S. pneumoniae-induced sepsis, pneumonia, meningitis, otitis media, sinusitis, or infection of other sites or organs by S. pneumoniae.
[0136] One appropriate art-recognized class of S. pneumoniae antigens is Pneumococcal surface protein A (PspA) and derivatives of PspA. Derivatives of PspA include proline-rich segments with the non-proline block (PR+NPB, also referred to as PRN and further described in Daniels et al, 2010), and related constructs comprising all or a fragment of the proline-rich region of PspA (e.g., regions containing one or more of the sequences PAPAP (SEQ ID NO: 65), PKP, PKEPEQ (SEQ ID NO: 66) and PEKP (SEQ ID NO: 67) and optionally including a non-proline block). In some embodiments, fragments or variants of PspA comprise proline-rich segments with the non-proline block and 10, 20 30, 40 or more additional amino acids of PspA sequence. Peptides containing the NPB are particularly immunogenic, suggesting that the NPB may be an important epitope.
[0137] Another appropriate art-recognized class of S. pneumoniae antigen is the pneumolysoids. Pneumolysoids have homology to the S. pneumoniae protein pneumolysin (PLY or Ply), but have reduced toxicity compared to pneumolysin. Pneumolysoids can be naturally occurring or engineered derivatives of pneumolysin. In some embodiments, a pneumolysoid has at least 70%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% identity to pneumolysin. In some embodiments, the pneumolysoid demonstrates less than one half, 1/10th, 1/20th, 1/50th, 1/100th, 1/200th, 1/500th, or 1/1000th the toxicity of pneumolysin in an assay for one or both of hemolytic activity towards erythrocytes and inhibition of polymorphonuclear leukocytes. Both assays are described in Saunders et al, 1989. Exemplary pneumolysoids include PdT, a triple mutant further described in Berry et al, 1995; Pd-A and Pd-B, further described in Paton et al, 1991; rPd2 and rPd3, further described in Ferreira et al, 2006; Ply8, MPLY, and L460D, further described in, e.g., US 2009/0285846 and L. Mitchell, 2011; or variants thereof. In some embodiments, the pneumolysin has a mutation in the catalytic center, such as at amino acid 428 or 433 or its vicinity.
[0138] Other appropriate S. pneumoniae antigens include choline-binding protein A (CbpA) and derivatives thereof (Ogunniyi et al, 2001); pneumococcal surface adhesin A (PsaA); caseinolytic protease; sortase A (SrtA); pilus 1 RrgA adhesin; PpmA; PrtA; PavA; LytA; Stk-PR; PcsB; RrgB and derivatives thereof. CpbA derivatives include constructs described in WO 2012/134975. Such constructs may comprise one or more copies of the R2 domain, R21 and/or R22 subdomains of CpbA, or active variants and fragments thereof, or any combination thereof. Such constructs may further comprise a pneumolysoid.
[0139] In some embodiments, an immunogenic composition (e.g., a vaccine composition) contains one or more fusion proteins described herein in combination with one or more polypeptides from Table 1, or antigenic fragments or variants thereof, in a mixture. In some embodiments, the mixture contains both full-length polypeptides and fragments resulting from processing, or partial processing, of signal sequences by an expression host, e.g. E. coli, an insect cell line (e.g., the baculovirus expression system), or a mammalian cell line (e.g., human or Chinese Hamster Ovary).
[0140] In some embodiments, an immunogenic composition contains one or more fusion proteins of any of SEQ ID NOs:17-26 in the absence of any other antigens. In some embodiments, an immunogenic composition contains a fusion protein of SEQ ID NO:23 in the absence of any other antigens. In some embodiments, an immunogenic composition contains one or more fusion proteins of any of SEQ ID NOs:17-26 in combination with one or more additional proteins of any of SEQ ID NOs:1-8, in the absence of other antigens. In some embodiments, an immunogenic composition contains a fusion protein of SEQ ID NO:23 in combination with one or more additional proteins of any of SEQ ID NOs:1-8, in the absence of any other antigens.
[0141] In some embodiments, fusion proteins described herein may be conjugated to S. pneumoniae polysaccharides. In some embodiments, fusion proteins described herein may be non-covalently complexed to S. pneumoniae polysaccharides. The S. pneumoniae polysaccharides may be, for example, as described in U.S. Pat. Nos. 5,623,057, 5,371,197, or PCT/US2011/023526. The non-covalent complexes may be, for example, those of the Multiple Antigen Presenting System (MAPS), as described in PCT/US2012/037412, PCT/US2012/037541, and Zhang et al, 2013.
[0142] In some embodiments, a fusion protein described herein is covalently bound to another molecule. This may, for example, increase the half-life, solubility, bioabailability, or immunogenicity of the fusion protein. Molecules that may be covalently bound to the fusion protein include a carbohydrate, biotin, poly(ethylene glycol) (PEG), polysialic acid, N-propionylated polysialic acid, nucleic acids, polysaccharides, and PLGA. There are many different types of PEG, ranging from molecular weights of below 300 g/mol to over 10,000,000 g/mol. PEG chains can be linear, branched, or with comb or star geometries. In some embodiments, the fusion protein is covalently bound to a moeity that stimulates the immune system. An example of such a moeity is a lipid moeity. In some instances, lipid moieties are recognized by a Toll-like receptor (TLR) such as TLR-2 or TLR-4, and activate the innate immune system.
[0143] In some embodiments, a fusion protein and one or more additional components described herein are mixed together using known methods to form a multi-component immunogenic composition. In some embodiments, a fusion protein and one or more additional components described herein are nano-encapsulated using known methods. In some embodiments, a fusion protein and one or more additional components described herein are molded into nano- or micro-particles using known methods. In some embodiments, a fusion protein and one or more additional components described herein are conjugated through a covalent bond using known methods to form a multi-component immunogenic composition. In some embodiments, a fusion protein and one or more additional components described herein are joined non-covalently using known methods to form a multi-component immunogenic composition. Additional methods of combining a fusion protein and one or more additional components are described in, e.g., PCT/US2012/37412 and PCT/US2009/44956.
Nucleic Acid-Based Immunogenic Compositions and Vaccines
[0144] The present disclosure also provides immunogenic compositions (e.g., vaccine compositions) of, or comprising, one or more nucleic acids encoding fusion proteins described herein. In some embodiments, the immunogenic composition comprises one or more nucleic acids encoding fusion proteins with at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% sequence identity to a fusion protein listed in Table 2. In some embodiments, the immunogenic composition comprises a nucleic acid encoding a fusion protein that is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% identity to any one of SEQ ID NOs:17-26. In some embodiments, the immunogenic composition comprises a nucleic acid encoding a fusion protein that is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.5% identity to SEQ ID NO:23. In some embodiments, the immunogenic composition comprises a nucleic acid encoding a fusion protein that is or includes an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% identity to CP1.
[0145] In some embodiments, the immunogenic composition comprises one or more nucleic acids having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% identity to any one of SEQ ID NOs:27-36. In some embodiments, the immunogenic composition comprises a nucleic acid having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% identity to SEQ ID NO:33. In all cases, due to degeneracy in the genetic code, other DNA sequences (including multiple codon-optimized sequences) could encode such fusion proteins. In some embodiments, these nucleic acids are expressed in the immunized individual, resulting in production of the encoded S. pneumoniae fusion proteins, and the S. pneumoniae fusion proteins so produced have an immunostimulatory or immunoprotective effect in the immunized individual.
[0146] Such a nucleic acid-containing immunostimulatory composition may comprise, for example, an origin of replication, and/or a promoter that drives expression of one or more nucleic acids encoding one or more fusion proteins of SEQ ID NOs:27-36. Such a composition may also comprise a bacterial plasmid vector into which is inserted a promoter (sometimes a strong viral promoter), one or more nucleic acids encoding one or more fusion proteins of SEQ ID NOs:17-26, and a polyadenylation/transcriptional termination sequence. In some instances, the nucleic acid is DNA. In some instances, the nucleic acid is RNA.
Uses of Immunogenic and Vaccine Compositions
[0147] In some embodiments, an immunogenic composition or vaccine that includes one or more fusion proteins described herein is characterized in that one or more of the opsonization potential or immune responses to one or more fusion proteins is increased relative to a pre-determined level, as measured by ELISA and/or by a functional antibody assay. In some embodiments, one or more of the opsonization potential or immune response to the one or more fusion proteins is increased by at least 30% or more, including, e.g., at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90% or more, relative to a predetermined level, as measured by ELISA and/or by a functional antibody assay. In some embodiments, one or more of the opsonization potential or immune responses to the one or more fusion proteins is increased at least 1-fold, 2-fold, 3-fold, 4-fold, or 5-fold relative to a pre-determined level, as measured by ELISA and/or by a functional antibody assay. In some embodiments, the pre-determined level is a pre-immune level (e.g., a level observed when a subject is not immunized, or is immunized in the absence of one or more fusion proteins described herein).
[0148] In some embodiments, an immunogenic composition or vaccine that includes one or more fusion proteins described herein, upon administration to a subject, induces an immune response against S. pneumoniae. In some embodiments, the immunogenic composition or vaccine, upon administration to a subject, induces an immune response against one or more serotypes of S. pneumoniae. In some embodiments, the immunogenic composition or vaccine, upon administration to a subject, induces a protective immune response against one or more serotypes of S. pneumoniae. In some embodiments, the immune response is an antibody or B cell response. In some embodiments, the immune response is a T cell response. In some embodiments, the immune response is an innate immune response. In some embodiments, the immune response is a CD4+ T cell response, including Th1, Th2, or Th17 response, or a CD8+ T cell response, or a CD4+ and a CD8+ T cell response, or a CD4-/CD8-T cell response. In some embodiments, the immune response is an antibody or B cell response and a T cell response. In some embodiments, the immune response is an antibody or B cell response, a T cell response, and an innate immune response.
[0149] In some embodiments, an immunogenic composition or vaccine that includes one or more fusion proteins described herein may be used for prophylactic and/or therapeutic treatment of S. pneumoniae. Accordingly, the present disclosure provides a method for immunizing a subject suffering from or susceptible to S. pneumoniae infection, comprising administering an immunologically effective amount of any immunogenic composition or vaccine that includes one or more fusion proteins described herein. The subject receiving the immunization may be a male or a female, and may be an infant, child, adolescent, or adult. In some embodiments, the subject being treated is a human. In other embodiments, the subject is a non-human animal.
[0150] In some embodiments, upon administration to a subject, an immunogenic composition or vaccine comprising a fusion protein described herein treats or prevents infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents Invasive Pneumococcal Disease (IPD) due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents bacteremia due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents sepsis due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents organ damage due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents meningitis due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents pneumonia due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents otitis media due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine treats or prevents sinusitis due to infection by S. pneumoniae.
[0151] In some embodiments, upon administration to a subject, an immunogenic composition or vaccine comprising a fusion protein described herein inhibits or reduces the rate of occurrence of infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of Invasive Pneumococcal Disease (IPD) due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of bacteremia due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of sepsis due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of organ damage due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of meningitis due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of pneumonia due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of otitis media due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits or reduces the rate of occurrence of sinusitis due to infection by S. pneumoniae.
[0152] In some embodiments, upon administration to a subject, an immunogenic composition or vaccine comprising a fusion protein described herein reduces the severity of infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of Invasive Pneumococcal Disease (IPD) due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of bacteremia due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of sepsis due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of organ damage due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of meningitis due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of pneumonia due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of otitis media due to infection by S. pneumoniae. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine reduces the severity of sinusitis due to infection by S. pneumoniae.
[0153] In some embodiments, upon administration to a subject, an immunogenic composition or vaccine comprising a fusion protein described herein inhibits transmission of S. pneumoniae from the subject to another subject. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits colonization by S. pneumoniae in the subject. In some embodiments, upon administration to a subject, the immunogenic composition or vaccine inhibits colonization by S. pneumoniae in the nasopharynx of the subject.
[0154] In some embodiments, an immunogenic composition or vaccine comprising a fusion protein described herein, upon administration to a subject, induces an immune response against S. pneumoniae in the subject at a level greater than a control composition. In some embodiments, the immunogenic composition or vaccine, upon administration to a subject, induces an immune response against one or more serotypes of S. pneumoniae at a level greater than a control composition. In some embodiments, the level greater is about 1%, about 2%, about 3%, about 4%, about 5%, about 6%, about 7%, about 8%, about 9%, about 10%, about 11%, about 12%, about 13%, about 14%, about 15%, about 16%, about 17%, about 18%, about 19%, about 20%, about 21%, about 22%, about 23%, about 24%, about 25%, about 30%, about 35%, about 40%, about 45%, about 50%, about 55%, about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, or about 95% of the control composition.
[0155] In some embodiments, an immunogenic composition or vaccine comprising a fusion protein described herein, upon administration to a subject, induces an immune response that can help protect against the establishment of S. pneumoniae at a level greater than a control composition. In some embodiments, the immunogenic composition or vaccine protects against colonization at a level greater than a control composition. In some embodiments, the immunogenic composition or vaccine inhibits infection by S. pneumoniae in a non-colonized or uninfected subject at a level greater than a control composition. In some embodiments, the immunogenic composition or vaccine reduces the duration of colonization by S. pneumoniae in a subject who is already colonized at a level greater than a control composition. In some embodiments, the level greater is about 1%, about 2%, about 3%, about 4%, about 5%, about 6%, about 7%, about 8%, about 9%, about 10%, about 11%, about 12%, about 13%, about 14%, about 15%, about 16%, about 17%, about 18%, about 19%, about 20%, about 21%, about 22%, about 23%, about 24%, about 25%, about 30%, about 35%, about 40%, about 45%, about 50%, about 55%, about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, or about 95% of the control composition.
Antibody Compositions
[0156] Some embodiments provide for an antibody composition comprising antibodies raised in a mammal immunized with an immunogenic composition or vaccine comprising a fusion protein described herein. In some embodiments, an antibody comprises at least one antibody selected from the group consisting of monoclonal Abs (mAbs) and anti-idiotype antibodies. In some embodiments, an antibody composition comprises an isolated gamma globulin fraction. In some embodiments, an antibody composition comprises polyclonal antibodies. In some embodiments, the antibody composition is administered to a subject.
Vaccine Formulations
[0157] Optimal amounts of components for a particular vaccine comprising a fusion protein described herein can be ascertained by standard studies involving observation of appropriate immune responses in subjects. Following an initial immunization, subjects can receive one or several booster immunizations adequately spaced in time.
[0158] The immunogenic composition or vaccine comprising a fusion protein described herein, and/or preparations thereof, may be formulated in a unit dosage form for ease of administration and uniformity of dosage. The specific therapeutically effective dose level for any particular subject or organism may depend upon a variety of factors including the severity or degree of risk of infection; the activity of the specific vaccine or vaccine composition employed; other characteristics of the specific vaccine or vaccine composition employed; the age, body weight, general health, sex of the subject, diet of the subject, pharmacokinetic condition of the subject, the time of administration (e.g., with regard to other activities of the subject such as eating, sleeping, receiving other medicines including other vaccine doses, etc.), route of administration, rate of excretion of the specific vaccine or vaccine composition employed; vaccines used in combination or coincidental with the vaccine composition employed; and like factors well known in the medical arts.
[0159] An immunogenic composition or vaccine comprising a fusion protein described herein for use in accordance with the present disclosure may be formulated into compositions (e.g., pharmaceutical compositions) according to known techniques. Vaccine preparation is generally described in Vaccine Design (Powell and Newman, 1995). For example, an immunologically amount of a vaccine product can be formulated together with one or more organic or inorganic, liquid or solid, pharmaceutically suitable carrier materials.
[0160] In general, pharmaceutically acceptable carrier(s) include solvents, dispersion media, and the like, which are compatible with pharmaceutical administration. For example, materials that can serve as pharmaceutically acceptable carriers include, but are not limited to sugars such as lactose, glucose, dextrose, and sucrose; starches such as corn starch and potato starch; cellulose and its derivatives such as sodium carboxymethyl cellulose, ethyl cellulose and cellulose acetate; powdered tragacanth; malt; gelatin; talc; excipients such as cocoa butter and suppository waxes; oils such as peanut oil, cottonseed oil, safflower oil, sesame oil, olive oil, corn oil and soybean oil; polyols such as glycerol, propylene glycol, and liquid polyethylene glycol; esters such as ethyl oleate and ethyl laurate; agar; buffering agents such as magnesium hydroxide; alginic acid; pyrogen-free water; isotonic saline; Ringer's solution; ethyl alcohol, and phosphate buffer solutions, as well as other non-toxic compatible lubricants such as sodium lauryl sulfate and magnesium stearate, as well as preservatives, and antioxidants can also be present in the composition, according to the judgment of the formulator (Martin, 1975).
[0161] Vaccines may be formulated by combining one or more fusion proteins described herein with carriers and/or other optional components by any available means including, for example, conventional mixing, granulating, dissolving, lyophilizing, or similar processes.
[0162] Vaccines comprising one or more fusion proteins described herein may be lyophilized up until they are about to be used, at which point they are extemporaneously reconstituted with diluent. In some embodiments, vaccine components or compositions are lyophilized in the presence of one or more other components (e.g., adjuvants), and are extemporaneously reconstituted with saline solution. Alternatively, individual components, or sets of components may be separately lyophilized and/or stored (e.g., in a vaccination kit), the components being reconstituted and either mixed prior to use or administered separately to the subject.
[0163] Lyophilization can produce a more stable composition (for instance by preventing or reducing breakdown of polysaccharide antigens). Lyophilizing of vaccines or vaccine components is well known in the art. Typically, a liquid vaccine or vaccine component is freeze dried, often in the presence of an anti-caking agent (such as, for example, sugars such as sucrose or lactose). In some embodiments, the anti-caking agent is present, for example, at an initial concentration of 10-200 mg/ml. Lyophilization typically occurs over a series of steps, for instance a cycle starting at -69.degree. C., gradually adjusting to -24.degree. C. over 3 h, then retaining this temperature for 18 h, then gradually adjusting to -16.degree. C. over 1 h, then retaining this temperature for 6 h, then gradually adjusting to +34.degree. C. over 3 h, and finally retaining this temperature over 9 h.
[0164] In some embodiments, a vaccine comprising a fusion protein described herein is a liquid. In some embodiments the liquid is a reconstituted lyophylate. In some embodiments a vaccine has a pH of about 5, about 6, about 7, or about 8. In some embodiments a vaccine has a pH between about 5 and about 7.5. In some embodiments a vaccine has a pH between 5 and 7.5. In some embodiments a vaccine has a pH between about 5.3 and about 6.3. In some embodiments a vaccine has a pH between 5.3 and 6.3. In some embodiments a vaccine has a pH of about 5.0, about 5.1, about 5.2, about 5.3, about 5.4, about 5.5, about 5.6, about 5.7, about 5.8, about 5.9, about 6.0, about 6.1, about 6.2, about 6.3, about 6.4, about 6.5, about 6.6, about 6.7, about 6.8, about 6.9, about 7.0, about 7.1, about 7.2, about 7.3, about 7.4, or about 7.5.
[0165] Vaccines or vaccine components for use in accordance with the present disclosure may be incorporated into liposomes, cochleates, biodegradable polymers such as poly-lactide, poly-glycolide and poly-lactide-co-glycolides, or immune-stimulating complexes (ISCOMS).
[0166] In certain situations, it may be desirable to prolong the effect or release of a vaccine for use in accordance with the present invention, for example, by slowing the absorption of one or more vaccine components. Such delay of absorption may be accomplished, for example, by the use of a liquid suspension of crystalline or amorphous material with poor water solubility. The rate of absorption of the product then depends upon its rate of dissolution, which in turn, may depend upon size and form. Alternatively, or additionally, delayed absorption may be accomplished by dissolving or suspending one or more vaccine components in an oil vehicle. Injectable depot forms can also be employed to delay absorption. Such depot forms can be prepared by forming microcapsule matrices of one or more vaccine components a biodegradable polymer network. Depending upon the ratio of polymer to vaccine component, and the nature of the particular polymer(s) employed, the rate of release can be controlled.
[0167] Examples of biodegradable polymers that can be employed in accordance with the present disclosure include, for example, poly(orthoesters) and poly(anhydrides). One particular exemplary polymer is polylactide-polyglycolide.
[0168] Depot injectable formulations may also be prepared by entrapping the product in liposomes or microemulsions, which are compatible with body tissues.
[0169] Polymeric delivery systems can also be employed in non-depot formulations including, for example, oral formulations. For example, biodegradable, biocompatible polymers such as ethylene vinyl acetate, polyanhydrides, polyglycolic acid, collagen, polyorthoesters, and polylactic acid, etc., can be used in oral formulations. Polysaccharide antigens or conjugates may be formulated with such polymers, for example to prepare particles, microparticles, extrudates, solid dispersions, admixtures, or other combinations in order to facilitate preparation of useful formulations (e.g., oral).
[0170] Vaccines comprising one or more fusion proteins described herein for use in accordance with the present disclosure include immunogenic compositions, and may additionally include one or more additional active agents (i.e., agents that exert a biological effect--not inert ingredients). For example, it is common in vaccine preparation to include one or more adjuvants. It will be appreciated that such additional agents may be formulated together with one or more other vaccine components, or may be maintained separately and combined at or near the time of administration. In some embodiments, such additional components may be administered separately from some or all of the other vaccine components, within an appropriate time window for the relevant effect to be achieved.
[0171] Adjuvants
[0172] The vaccine formulations and immunogenic compositions comprising a fusion protein described herein may include an adjuvant. Adjuvants, generally, are agents that enhance the immune response to an antigen. Adjuvants can be broadly separated into two classes, based on their principal mechanisms of action: vaccine delivery systems and immunostimulatory adjuvants (see, e.g., Singh et al, 2003). In most vaccine formulations, the adjuvant provides a signal to the immune system so that it generates a response to the antigen, and the antigen is required for driving the specificity of the response to the pathogen. Vaccine delivery systems are often particulate formulations, e.g., emulsions, microparticles, immune-stimulating complexes (ISCOMs), nanoparticles, which may be, for example, particles and/or matrices, and liposomes. In contrast, immunostimulatory adjuvants are sometimes from or derived from pathogens and can represent pathogen associated molecular patterns (PAMP), e.g., lipopolysaccharides (LPS), monophosphoryl lipid A (MPL), or CpG-containing DNA, which activate cells of the innate immune system.
[0173] Alternatively, adjuvants may be classified as organic and inorganic. Inorganic adjuvants include alum salts such as aluminum phosphate, amorphous aluminum hydroxyphosphate sulfate, and aluminum hydroxide, which are commonly used in human vaccines. Organic adjuvants comprise organic molecules including macromolecules. Non-limiting examples of organic adjuvants include cholera toxin/toxoids, other enterotoxins/toxoids or labile toxins/toxoids of Gram-negative bacteria, interleukins (e.g., IL-1, IL-2, IL-4, IL-5, IL-6, IL-7, IL-12, IL-15, IL-18, etc.), interferons (e.g., gamma interferon), granulocyte macrophage colony stimulating factor (GM-CSF), macrophage colony stimulating factor (M-CSF), and tumor necrosis factor (TNF).
[0174] Adjuvants may also be classified by the response they induce. In some embodiments, the adjuvant induces the generation, proliferation, or activation of Th1 cells or Th2 cells. In other embodiments, the adjuvant induces the generation, proliferation, or activation of B cells. In yet other embodiments, the adjuvant induces the activation of antigen-presenting cells. These categories are not mutually exclusive; in some cases, an adjuvant activates more than one type of cell.
[0175] In some embodiments, the adjuvant induces the generation, proliferation, or activation of Th17 cells. The adjuvant may promote the CD4+ or CD8+ T cells to secrete IL-17. In some embodiments, an adjuvant that induces the generation, proliferation, or activation of Th17 cells is one that produces at least a 2-fold, and in some cases a 10-fold, experimental sample to control ratio in the following assay. In the assay, an experimenter compares the IL-17 levels secreted by two populations of cells: (1) cells from animals immunized with the adjuvant and a polypeptide known to induce Th17 generation, proliferation, or activation, and (2) cells from animals treated with the adjuvant and an irrelevant (control) polypeptide. An adjuvant that induces the generation, proliferation, or activation of Th17 cells may cause the cells of population (1) to produce more than 2-fold, or more than 10-fold more IL-17 than the cells of population (2). IL-17 may be measured, for example, by ELISA or ELISPOT. Certain toxins, such as cholera toxin and labile toxin (produced by enterotoxigenic E. coli, or ETEC), activate a Th17 response. Thus, in some embodiments, the adjuvant is a toxin or toxoid. Cholera toxin was successfully used in a mouse model to induce protective immunity in conjunction with certain polypeptides from Table 1. One form of labile toxin is produced by Intercell. Mutant derivates of labile toxin (toxoids) that are active as adjuvants but significantly less toxic can be used as well. Exemplary detoxified mutant derivatives of labile toxin include mutants lacking ADP-ribosyltransferase activity. Particular detoxified mutant derivatives of labile toxin include LTK7 (Douce et al, 1995) and LTK63 (Williams et al, 2004), LT-G192 (Douce et al, 1999), and LTR72 (Giuliani et al, 1998).
[0176] In some embodiments, the adjuvant comprises a VLP (virus-like particle). One such adjuvant platform, Alphavirus replicons, induces the activation of Th17 cells using alphavirus and is produced by Alphavax. In some embodiments of the Alphavirus replicon system, alphavirus may be engineered to express an antigen of interest, a cytokine of interest (for example, IL-17 or a cytokine that stimulates IL-17 production), or both, and may be produced in a helper cell line. More detailed information may be found in U.S. Pat. Nos. 5,643,576 and 6,783,939. In some embodiments, a vaccine formulation is administered to a subject in combination with a nucleic acid encoding a cytokine.
[0177] Certain classes of adjuvants activate toll-like receptors (TLRs) in order to activate a Th17 response. TLRs are well known proteins that may be found on leukocyte membranes, and recognize foreign antigens (including microbial antigens). Administering a known TLR ligand together with an antigen of interest (for instance, as a fusion protein) can promote the development of an immune response specific to the antigen of interest. One exemplary adjuvant that activates TLRs comprises Monophosphoryl Lipid A (MPL). Traditionally, MPL has been produced as a detoxified lipopolysaccharide (LPS) endotoxin obtained from Gram-negative bacteria, such as S. minnesota. In particular, sequential acid and base hydrolysis of LPS produces an immunoactive lipid A fraction (which is MPL), and lacks the saccharide groups and all but one of the phosphates present in LPS. A number of synthetic TLR agonists (in particular, TLR-4 agonists) are disclosed in Evans et al, 2003. Like MPL adjuvants, these synthetic compounds activate the innate immune system via TLR. Another type of TLR agonist is a synthetic phospholipid dimer, for example E6020 (Ishizaka et al, 2007). Various TLR agonists (including TLR-4 agonists) have been produced and/or sold by, for example, the Infectious Disease Research Institute (IRDI), Corixa, Esai, Avanti Polar Lipids, Inc., and Sigma Aldrich. Another exemplary adjuvant that activates TLRs comprises a mixture of MPL, Trehalose Dicoynomycolate (TDM), and dioctadecyldimethylammonium bromide (DDA). Another TLR-activating adjuvant is R848 (resiquimod).
[0178] In some embodiments, the adjuvant is or comprises a saponin. Typically, the saponin is a triterpene glycoside, such as those isolated from the bark of the Quillaja saponaria tree. A saponin extract from a biological source can be further fractionated (e.g., by chromatography) to isolate the portions of the extract with the best adjuvant activity and with acceptable toxicity. Typical fractions of extract from Quillaja saponaria tree used as adjuvants are known as fractions A and C.
[0179] In some embodiments, combinations of adjuvants are used. Three exemplary combinations of adjuvants are MPL and alum, E6020 and alum, and MPL and an ISCOM.
[0180] Adjuvants may be covalently or non-covalently bound to antigens. In some embodiments, the adjuvant may comprise a protein which induces inflammatory responses through activation of antigen-presenting cells (APCs). In some embodiments, one or more of these proteins can be recombinantly fused with an antigen of choice, such that the resultant fusion molecule promotes dendritic cell maturation, activates dendritic cells to produce cytokines and chemokines, and ultimately, enhances presentation of the antigen to T cells and initiation of T cell responses (e.g., see Wu et al, 2005).
[0181] In some embodiments, an immunogenic composition or vaccine comprising a fusion protein described herein is formulated and/or administered in combination with an adjuvant. In some embodiments, the adjuvant is selected from the group consisting of aluminum phosphate, aluminum hydroxide, and phosphate aluminum hydroxide. In some embodiments, the adjuvant comprises aluminum phosphate. In some embodiments, the adjuvant is aluminum phosphate.
[0182] Typically, the same adjuvant or mixture of adjuvants is present in each dose of a vaccine. Optionally, however, an adjuvant may be administered with the first dose of vaccine and not with subsequent doses (i.e., booster shots). Alternatively, a strong adjuvant may be administered with the first dose of vaccine and a weaker adjuvant or lower dose of the strong adjuvant may be administered with subsequent doses. The adjuvant can be administered before the administration of the antigen, concurrent with the administration of the antigen or after the administration of the antigen to a subject (sometimes within 1, 2, 6, or 12 hours, and sometimes within 1, 2, or 5 days). Certain adjuvants are appropriate for human subjects, non-human animals, or both.
[0183] Vaccines for use in accordance with the present disclosure may include, or be administered concurrently with, antimicrobial therapy. For example, such vaccines may include or be administered with one or more agents that kills or retards growth of a pathogen. Such agents include, for example, penicillin, vancomycin, erythromycin, azithromycin, and clarithromycin, cefotaxime, ceftriaxone, levoflaxin, gatifloxacin.
[0184] Alternatively or additionally, vaccines for use in accordance with the present invention may include, or be administered with, one or more other vaccines or therapies. For example, one or more non-pneumococcal antigens may be included in or administered with the vaccines.
[0185] Additional Components and Excipients
[0186] In addition to the fusion proteins described herein and the adjuvants described above, a vaccine formulation or immunogenic composition may include one or more additional components.
[0187] In some embodiments, the vaccine formulation or immunogenic composition may include one or more stabilizers such as sugars (such as sucrose, glucose, or fructose), phosphate (such as sodium phosphate dibasic, potassium phosphate monobasic, dibasic potassium phosphate, or monosodium phosphate), glutamate (such as monosodium L-glutamate), gelatin (such as processed gelatin, hydrolyzed gelatin, or porcine gelatin), amino acids (such as arginine, asparagine, histidine, L-histidine, alanine, valine, leucine, isoleucine, serine, threonine, lysine, phenylalanine, tyrosine, and the alkyl esters thereof), inosine, or sodium borate.
[0188] In some embodiments, the vaccine formulation or immunogenic composition includes one or more buffers such as a mixture of sodium bicarbonate and ascorbic acid. In some embodiments, the vaccine formulation may be administered in saline, such as phosphate buffered saline (PBS), or distilled water.
[0189] In some embodiments, the vaccine formulation or immunogenic composition includes one or more surfactants, for example, but not limited to, polysorbate 80 (TWEEN 80), polysorbate 20 (TWEEN 20), Polyethylene glycol p-(1,1,3,3-tetramethylbutyl)-phenyl ether (TRITON X-100), and 4-(1,1,3,3-Tetramethylbutyl)phenol polymer with formaldehyde and oxirane (TYLOXAPOL). A surfactant can be ionic or nonionic.
[0190] In some embodiments, the vaccine formulation or immunogenic composition includes one or more salts such as sodium chloride, ammonium chloride, calcium chloride, or potassium chloride.
[0191] In some embodiments, a preservative is included in the vaccine or immunogenic composition. In other embodiments, no preservative is used. A preservative is most often used in multi-dose vaccine vials, and is less often needed in single-dose vaccine vials. In some embodiments, the preservative is 2-phenoxyethanol, methyl and propyl parabens, benzyl alcohol, and/or sorbic acid.
Methods of Administration
[0192] In some embodiments, an immunogenic composition or vaccine comprising a fusion protein described herein is administered to a subject at risk of developing pneumococcal disease, e.g. an infant, a toddler, a juvenile, or an older adult. In some embodiments, the immunogenic composition or vaccine is administered to a subject at elevated risk of developing pneumococcal disease, e.g., immunocompromised subjects, subjects having sickle cell disease or other hemoglobinopathies, congenital or acquired asplenia, splenic dysfunction, chronic renal failure or nephrotic syndrome, diseases associated with treatment with immunosuppressive drugs or radiation therapy, including malignant neoplasm, leukemia, lymphomas, Hodgkin's disease, or solid organ transplantation, congenital or acquired immunodeficiency, HIV infection, cerebrospinal fluid leaks, cochlear implant(s), chronic heart disease, chronic lung disease, diabetes mellitus, alcoholism, chronic liver disease, cigarette smoking, asthma, generalized malignancy, multiple myeloma, or solid organ transplantation. It will be appreciated that a subject can be considered at risk for developing a disease without having been diagnosed with any symptoms of the disease. For example, if the subject is known to have been, or to be intended to be, in situations with relatively high risk of infection, that subject will be considered at risk for developing the disease.
[0193] Any effective route of administration may be utilized such as, for example, oral, nasal, enteral, parenteral, intramuscular or intravenous, subcutaneous, transdermal, intradermal, rectal, vaginal, topical, ocular, pulmonary, or by contact application. In some embodiments, the immunogenic composition or vaccine may be injected (e.g., via intramuscular, intraperitoneal, intradermal and/or subcutaneous routes); or delivered via the mucosa (e.g., to the oral/alimentary, respiratory, and/or genitourinary tracts). Intranasal administration may be particularly useful in some contexts, for example for treatment of pneumonia or otitis media (as nasopharyngeal carriage of pneumococci can be more effectively prevented, thus attenuating infection at its earliest stage). In some embodiments, it may be desirable to administer different doses of the immunogenic composition or vaccine by different routes; in some embodiments, it may be desirable to administer different components of one dose via different routes.
[0194] In some embodiments, pharmaceutical compositions (e.g., immunogenic compositions or vaccines) are administered intradermally. Conventional technique of intradermal injection, the "Mantoux procedure", comprises steps of cleaning the skin, and then stretching with one hand, and with the bevel of a narrow gauge needle (26-31 gauge) facing upwards the needle is inserted at an angle of between 10-15.degree.. Once the bevel of the needle is inserted, the barrel of the needle is lowered and further advanced while providing a slight pressure to elevate it under the skin. The liquid is then injected very slowly thereby forming a bleb or bump on the skin surface, followed by slow withdrawal of the needle.
[0195] Devices that are specifically designed to administer liquid agents into or across the skin have been described, for example the devices described in WO 99/34850 and EP 1092444, also the jet injection devices described for example in WO 01/13977; U.S. Pat. Nos. 5,480,381, 5,599,302, 5,334,144, 5,993,412, 5,649,912, 5,569,189, 5,704,911, 5,383,851, 5,893,397, 5,466,220, 5,339,163, 5,312,335, 5,503,627, 5,064,413, 5,520,639, 4,596,556, 4,790,824, 4,941,880, 4,940,460, WO 97/37705 and WO 97/13537. Other methods of intradermal administration of the immunogenic compositions or vaccines may include conventional syringes and needles, or devices designed for ballistic delivery of solid vaccines (WO 99/27961), or transdermal patches (WO 97/48440; WO 98/28037); or applied to the surface of the skin (transdermal or transcutaneous delivery WO 98/20734; WO 98/28037).
[0196] As described above, pharmaceutical compositions (e.g., immunogenic compositions or vaccines) may be administered as a single dose or as multiple doses. It will be appreciated that an administration is a single "dose" so long as all relevant components are administered to a subject within a window of time; it is not necessary that every component be present in a single composition. For example, administration of two different immunogenic compositions or vaccines, within a period of less than 24 h, is considered a single dose. To give but one example, immunogenic compositions or vaccines having different antigenic components may be administered in separate compositions, but as part of a single dose. As noted above, such separate compositions may be administered via different routes or via the same route. Alternatively or additionally, in embodiments wherein an immunogenic composition or vaccine is combined with additional types of active agents, the immunogenic composition or vaccine may be administered via one route, and a second active agent may be administered by the same route or by a different route.
[0197] Pharmaceutical compositions (e.g., immunogenic compositions or vaccines) are administered in such amounts and for such time as is necessary to achieve a desired result. In some embodiments of the present invention, the immunogenic composition or vaccine comprises an immunologically effective amount of at least immunogenic composition. The exact amount required to achieve an immunologically effective amount may vary, depending on the immunogenic composition, and from subject to subject, depending on the species, age, and general condition of the subject, the stage of the disease, the particular pharmaceutical mixture, its mode of administration, and the like.
[0198] The amount of fusion protein(s) described hereinin each pharmaceutical composition (e.g., immunogenic composition or vaccine) dose is selected to allow the vaccine, when administered as described herein, to induce an appropriate immunoprotective response without significant adverse side effects.
[0199] In some embodiments, a pharmaceutical composition comprising a fusion protein described herein induces a Th1 and/or Th17 cell response upon administration to a subject. In some embodiments, the pharmaceutical composition induces an opsonic/bactericidal response against S. pneumoniae upon administration to a subject. In some embodiments, the pharmaceutical composition comprising a fusion protein disclosed herein reduces rate of transmission and/or colonization of the mucosal surfaces by Streptococcus pneumoniae upon administration to a subject. In some embodiments, the pharmaceutical composition reduces rate of transmission and/or colonization of the nasopharynx or the lungs by S. pneumoniae upon transmission.
[0200] Some embodiments provide for a method of immunizing a subject against S. pneumoniae infection comprising administering to the subject an immunologically effective amount of an immunogenic composition comprising a fusion protein described herein. Some embodiments provide for a method of immunizing a subject against S. pneumoniae infection comprising administering to the subject an immunologically effective amount of a vaccine composition comprising a fusion protein described herein. Some embodiments provide for a method of immunizing a subject against S. pneumoniae infection comprising administering to the subject an immunologically effective amount of a pharmaceutical composition comprising a fusion protein described herein.
[0201] Combination Prophylaxis or Combination Therapy
[0202] In some embodiments, an immunogenic composition or vaccine comprising a fusion protein described herein may be administered in combination with another agent. In some embodiments, the agent is or comprises PCV13. In some embodiments, the agent is or comprises PPSV23. In some embodiments, the agent is or comprises an antibiotic.
[0203] Dosing
[0204] In some embodiments, administration of an immunogenic composition or vaccine comprising a fusion protein described herein may involve the delivery of a single dose. In some embodiments, administration may involve an initial dose followed by one or several additional immunization doses, adequately spaced. An immunization schedule is a program for the administration of one or more specified doses of one or more specified pneumococcal vaccines, by one or more specified routes of administration, at one or more specified ages of a subject.
[0205] The present disclosure provides immunization methods that involve administering at least one dose of a vaccine to an infant subject. In some embodiments, the infant subject is 18 months old or younger. In some embodiments, the infant subject is 12 months old or younger. In some embodiments, the infant subject has previously received one or more doses of a conjugated pneumococcal polysaccharide vaccine; in other embodiments, the infant subject is naive to pneumococcal vaccines. In some embodiments, the infant subject has previously been infected with, or exposed to infection by S. pneumoniae.
[0206] The present disclosure provides immunization methods that involve administering at least one dose of a vaccine to a toddler subject. In some embodiments, the toddler subject is 5 years old or younger. In some embodiments, the toddler subject is 4 years old or younger. In some embodiments, the toddler subject has previously received one or more doses of a conjugated pneumococcal polysaccharide vaccine; in other embodiments, the toddler subject is naive to pneumococcal vaccines. In some embodiments, the toddler subject has previously been infected with, or exposed to infection by S. pneumoniae.
[0207] The present disclosure provides immunization methods that involve administering at least one dose of a vaccine to a juvenile subject. In some embodiments, the juvenile subject is 18 years old or younger. In some embodiments, the juvenile subject is 15 years old or younger. In some embodiments, the juvenile subject has previously received one or more doses of a conjugated pneumococcal polysaccharide vaccine; in other embodiments, the juvenile subject is naive to pneumococcal vaccines. In some embodiments, the juvenile subject has previously been infected with, or exposed to infection by S. pneumoniae.
[0208] The present disclosure provides immunization methods that involve administering at least one dose of a vaccine to an adult subject. In some embodiments, the adult subject is older than about 50 years of age. In some embodiments, the adult subject is older than about 65 years of age. In some embodiments, the adult subject has previously received one or more doses of a conjugated pneumococcal polysaccharide vaccine; in other embodiments, the adult subject is naive to pneumococcal vaccines. In some embodiments, the adult subject has previously been infected with, or exposed to infection by S. pneumoniae.
[0209] Immunization schedules of the present disclosure are provided to induce an immune response (e.g., an immunoprotective response) in a subject sufficient to reduce at least one measure selected from the group consisting of incidence, prevalence, frequency, and/or severity of at least one infection, disease, or disorder, and/or at least one surrogate marker of the infection, disease, or disorder, in a population and/or subpopulation of the subject(s). A supplemental immunization schedule is one which has this effect relative to the standard schedule which it supplements. A supplemental schedule may call for additional administrations and/or supra-immunogenic doses of the immunogenic compositions or vaccines disclosed herein, found in the standard schedule, or for the administration of immunogenic compositions or vaccines not part of the standard schedule. A full immunization schedule of the present invention may comprise both a standard schedule and a supplemental schedule. Exemplary sample immunization schedules are provided for illustrative purposes. Detailed descriptions of methods to assess immunogenic response discussed herein allow one to develop alterations to the sample immunization schedules without undue experimentation.
[0210] In one embodiment of the present disclosure, a first administration of a pneumococcal vaccine usually occurs when a subject is more than about 2 weeks old, more than about 5 weeks old, more than about 1 year old, more than about 2 years old, more than about 15 years old, or more than about 18 years old.
[0211] In one embodiment of the present disclosure, a first administration of a pneumococcal vaccine usually occurs when a subject is more than about 50 years old, more than about 55 years old, more than about 60 years old, more than about 65 years old, or more than about 70 years old.
[0212] In some embodiments of the disclosure, a single administration of vaccine is employed. It is possible that the purposes of the present invention can be served with a single administration, especially when one or more utilized vaccine polypeptides, polysaccharide(s) and/or conjugate(s) or combinations thereof is/are strong, and in such a situation a single dose schedule is sufficient to induce a lasting immune-protective response.
[0213] In some embodiments, it is desirable to administer two or more doses of vaccine, for greater immune-protective efficacy and coverage. Thus, in some embodiments, a number of doses is at least two, at least three or more doses. There is no set maximum number of doses, however it is good clinical practice not to immunize more often than necessary to achieve the desired effect.
[0214] Without being bound by theory, a first dose of vaccine administered according to the disclosure may be considered a "priming" dose. In some embodiments, more than one dose is included in an immunization schedule. In such a scenario, a subsequent dose may be considered a "boosting" dose.
[0215] A priming dose may be administered to a naive subject (a subject who has never previously received a conjugated polysaccharide vaccine). In some embodiments, a priming dose may be administered to a subject who has previously received conjugated polysaccharide vaccine at least five or more years previous to administration of an initial vaccine dose according to the invention. In other embodiments, a priming dose may be administered to a subject who has previously received a conjugated polysaccharide vaccine at least twenty or more years previous to administration of a priming vaccine according to the invention.
[0216] When an immunization schedule calls for two or more separate doses, the interval between doses is considered. The interval between two successive doses may be the same throughout an immunization schedule, or it may change as the subject ages. In immunization schedules of the present invention, once a first vaccine dose has been administered, there is a first interval before administration of a subsequent dose. A first interval is generally at least about 2 weeks, 1 month, 6 weeks, 2 months, 3 months, 6 months, 9 months, 12 months, or longer. Where more than one subsequent dose(s) are administered, second (or higher) intervals may be provided between such subsequent doses. In some embodiments, all intervals between subsequent doses are of the same length; in other embodiments, second intervals may vary in length. In some embodiments, the interval between subsequent doses may be at least about 12 months, at least about 15 months, at least about 18 months, at least about 21 months or at least about 2 years. In some embodiments, the interval between doses may be up to 3 years, up to about 4 years, or up to about 5 years or 10 years or more. In some embodiments, intervals between subsequent doses may decrease as the subject ages.
[0217] It will be appreciated by those skilled in the art that a variety of possible combinations and sub-combinations of the various conditions of timing of the first administration, shortest interval, largest interval and total number of administrations (in absolute terms, or within a stated period) exist, and all of these combinations and sub-combinations should be considered to be within the inventor's contemplation though not explicitly enumerated here.
Assays for Determining Immune Response
[0218] In some embodiments, a method of assessing the immunogenicity of a pharmaceutical composition, immunogenic composition, or vaccine comprising a fusion protein described herein comprises evaluating, measuring, and/or comparing an immune response using one or more in vitro bioassays, including B cell and T cell responses such as antibody levels by ELISA, multiplex ELISA, MSD, Luminex, flow cytometry, Th1/Th17 cell response, cytokine level measurement and functional antibody levels as measured by OPK, serum bactericidal killing (SBA), agglutination, motility, cytotoxicity, or adherence; and in vivo assays in animal models of pneumococcal disease (e.g. pneumonia, bacteremia, meningitis, sepsis, otitis media, nasopharyngeal colonization). Parameters of in vivo assays include bacterial clearance from mucosal surfaces or bloodstream, reduction or prevention of bacteremia, meningitis, sepsis, or otitis media, reduction or prevention of colonization of the nasopharynx, reduction of mortality, and passive and active protection following challenge with the pneumococcal pathogens that are the targets of the immunogenic composition. In some embodiments, the immune response is compared to a control composition.
[0219] In some embodiments, a method of assessing the potency of a pharmaceutical composition, immunogenic composition, or vaccine comprising a fusion protein described herein comprises evaluating, measuring, and/or comparing an immune response using one or more in vitro bioassays, including B cell and T cell responses such as antibody levels by ELISA, multiplex ELISA, MSD, Luminex, flow cytometry, Th1/Th17 cell response, cytokine level measurement and functional antibody levels as measured by OPK, serum bactericidal killing (SBA), internalization, activity neutralization, agglutination, motility, cytotoxicity, or adherence; and in vivo assays in animal models of pneumococcal disease (e.g. pneumonia, bacteremia, meningitis, sepsis, otitis media, nasopharyngeal colonization). Parameters include bacterial clearance or reduction from mucosal surfaces or bloodstream, reduction or prevention of bacteremia, meningitis, sepsis, or otitis media, reduction or prevention of colonization of the nasopharynx, reduction of mortality, and passive and active protection following challenge with the pneumococcal pathogens that are the targets of the immunogenic composition. In some embodiments, the immune response is compared to a control composition.
[0220] Generally speaking, it may be desirable to assess humoral responses, cellular responses, and/or interactions between the two. Where humoral responses are being assessed, antibody titers and/or types (e.g., total IgG, IgG1, IgG2, IgM, IgA, etc.) to specific pathogen antigens (e.g., polypeptides or polysaccharides, either serotype-specific or conserved across two or more serotypes) may be determined, for example before and/or after administration of an initial or a boosting dose of vaccine (and/or as compared with antibody levels in the absence of antigenic stimulation). Cellular responses may be assessed by monitoring reactions such as delayed type hypersensitivity responses, etc. to the antigens. Cellular responses can also be measured directly by evaluating the response of peripheral blood mononuclear cells (PBMCs) monocytes to stimulation with the antigens of interest. Precursor and memory B cell populations may be assessed in enzyme-linked immunospot (ELISpot) assays directed against specific pathogen antigens.
[0221] The RIA method detects specific antibodies through incubation of sera with radio-labeled polysaccharides or polypeptides in suspension (e.g., Schiffiman et al, 1980). The antigen-antibody complexes are then precipitated with ammonium sulfate and the radiolabeled pellets assayed for counts per minute (cpm).
[0222] In the ELISA detection method, specific antibodies from the sera of vaccinated subjects are quantitated by incubation with antigens (e.g., polypeptides or polysaccharides, either serotype-specific or conserved across two or more serotypes) which have been adsorbed to a solid support (e.g., Koskela and Leinonen (1981); Kojima et al, 1990; Concepcion and Frasch, 2001). The bound antibody is detected using enzyme-conjugated secondary detection antibodies. The ELISA also allows isotyping and subclassing of the immune response (i.e., IgM vs. IgG or IgG1 vs. IgG2) by using isotype- or subclass-specific secondary antibodies and can be adapted to evaluate the avidity of the antibodies (Anttila et al, 1998; Romero-Steiner et al, 2005). Multiplex assays (e.g., Luminex) facilitate simultaneous detection of antibodies to multiple antigens. Antigens are conjugated to spectrally distinct microspheres that are mixed and incubated with serum. The antibodies bound to the antigens on the coated microspheres are detected using a secondary antibody (e.g., R-Phycoerythrin-conjugated goat anti-human IgG).
[0223] An approach for assessing functional antibody in serum is the opsonophagocytic assay (OPA) which quantitates only the antibodies that can opsonize the bacteria, leading to ingestion and killing of the bacteria. The standard assay utilizes a human phagocytic effector cell, a source of complement, bacteria, and diluted sera. The assay readout is the serum endpoint titer at which there is >50% killing compared to bacteria incubated with complement and human cells alone (Romero-Steiner et al, 1997). This killing OPA can also be multiplexed by utilizing target strains of pathogen that carry different antibiotic resistance markers (Kim et al, 2003). Another type of multiplex opsonic assay is a nonkilling assay in which the uptake by phagocytic effector cells of fluorescent stained encapsulated pathogen or fluorescent microspheres conjugated with antigens from a target pathogen in the presence of diluted sera plus a complement source is evaluated by FC (Martinez et al, 1999). Opsonic activity of serum antibody plus complement can also be evaluated by measuring the oxidative response of phagocytic human effector cells to ingested pathogen (Munro et al. 1985; Ojo-Amaize et al. 1995).
[0224] Certain in vivo model systems can be used to evaluate the protection afforded by serum antibodies induced by immunogenic compositions or vaccines comprising a fusion protein described herein. In such passive protection systems, mice or rats are challenged with the pathogen plus diluted sera, and the endpoint titer of the sera which provides protection against pneumonia, bacteremia, colonization of organs or tissues, or mortality is determined (Stack et al. 1998; Saeland et al. 2000).
[0225] In some embodiments, efficacy of immunization may be determined by assaying one or more cytokine levels by stimulating T cells from a subject after immunization. The one or more cytokine levels may be compared to the one or more cytokine levels in the same subject before immunization. Increased levels of the one or more cytokine, such as a 1.5 fold, 2-fold, 5-fold, 10-fold, 20-fold, 50-fold or 100-fold or more increase over pre-immunization cytokine levels, would indicate an increased response to the immunogenic composition or vaccine. In some embodiments, the one or more cytokines are selected from GM-CSP; IL-1.alpha.; IL-1.beta.; IL-2; IL-3; IL-4; IL-5; IL-6; IL-7; IL-8; IL-10; IL-12; IL-17A, IL-17F or other members of the IL-17 family; IL-22; IL-23; IFN-.alpha.; IFN-.beta.; IFN-.gamma.; MIP-1.alpha.; MIP-1.beta.; TGF-.beta.; TNF.alpha., or TNF-.beta.. In a non-limiting example, efficacy of immunization may be determined by assaying IL-17 levels (particularly IL-17A) by stimulating T cells from a subject after immunization. The IL-17 levels may be compared to IL-17 levels in the same subject before immunization. Increased IL-17 (e.g., IL-17A) levels, such as a 1.5 fold, 2-fold, 5-fold, 10-fold, 20-fold, 50-fold or 100-fold or more increase, would indicate an increased response to the immunogenic composition or vaccine.
[0226] In some embodiments, one may assay neutrophils in the presence of T cells or antibodies from the patient for pneumococcal killing. Increased pneumococcal killing, such as a 1.5 fold, 2-fold, 5-fold, 10-fold, 20-fold, 50-fold or 100-fold or more increase, would indicate an increased response to the immunogenic composition or vaccine. For example, one may measure Th17 cell activation, where increased Th17 cell activation, such as a 1.5 fold, 2-fold, 5-fold, 10-fold, 20-fold, 50-fold or 100-fold or more increase, correlates with an increased response to the immunogenic composition or vaccine. In another non-limiting example, one may measure Th1 cell activation, where increased Th1 cell activation, such as a 1.5 fold, 2-fold, 5-fold, 10-fold, 20-fold, 50-fold or 100-fold or more increase, correlates with an increased response to the immunogenic composition or vaccine. One may also measure levels of an antibody specific to the immunogenic composition or vaccine, where increased levels of the specific antibody, such as a 1.5 fold, 2-fold, 5-fold, 10-fold, 20-fold, 50-fold or 100-fold or more increase, are correlated with increased efficacy. In some embodiments, two or more of these assays are used. For example, one may measure IL-17 levels and the levels of immunogenic composition- or vaccine-specific antibody. Alternatively, one may follow epidemiological markers such as incidence of, severity of, or duration of pneumococcal infection in vaccinated individuals compared to unvaccinated individuals.
[0227] Immunogenic composition or vaccine efficacy may also be assayed in various model systems such as the mouse challenge model. For instance, BALB/c or C57BL/6 strains of mice may be used. After administering the test immunogenic composition or vaccine to a subject (as a single dose or multiple doses), the experimenter administers a challenge dose of S. pneumoniae. In some cases, a challenge dose administered intranasally is sufficient to cause S. pneumoniae colonization (especially nasal colonization) in an unvaccinated animal, and in some cases a challenge dose administered via aspiration is sufficient to cause sepsis and a high rate of lethality in unvaccinated animals. In some cases, a challenge dose administered via intraperitoneal injection is sufficient to cause sepsis and a high rate of lethality in unvaccinated animals. In some cases, a challenge dose administered via intravenous injection is sufficient to cause sepsis and a high rate of lethality in unvaccinated animals. One can then measure the reduction in colonization or the reduction in lethality in vaccinated animals.
[0228] Certain in vivo model systems can be used to evaluate the protection afforded by serum antibodies induced by vaccines of the present invention. In such passive protection systems, mice or rats are challenged with the pathogen plus diluted sera, and the endpoint titer of the sera which provides protection against bacteremia, colonization of organs or tissues, or mortality is determined (Stack et al. 1998; Saeland et al. 2000).
Exemplification
Example 1: Induction of Th17 Response and Protection from Pneumococcal Nasal Colonization in Mice Mediated by SP0785 and SP1500 Components of Fusion Protein CP1
[0229] Objective:
[0230] This Example compares the ability of individual Streptococcus pneumoniae proteins SP0785 and SP1500 with an exemplary fusion protein CP1 to stimulate a Th17 response and protect mice from nasal colonization with S. pneumoniae after nasal immunization with the adjuvant cholera toxin (CT). An exemplary fusion protein CP1 is a fusion protein comprising a truncated rhizavidin (amino acids [45-179], denoted Rhavi), a SP0785 polypeptide, and a SP1500 polypeptide. In some embodiments, a fusion protein CP1 is or comprises Rhavi-linker (GGGGSSS)-SP1500-linker (AAA)-SP0785.
SUMMARY
[0231] 1. S. pneumoniae proteins SP0785 and SP1500, and fusion protein CP1 can each generate a robust Th17 response in mice as demonstrated by secretion of interleukin 17A (IL-17A) after intranasal immunization with the adjuvant cholera toxin (CT). [0232] 2. S. pneumoniae nasal colonization was significantly reduced by separate intranasal immunizations of mice with SP1500 or SP0785 and CT, or with CP1 and CT.
[0233] Materials and Methods:
[0234] Recombinant Protein Production
[0235] Histidine-tagged recombinant proteins were expressed in Escherichia coli and purified using Ni-nitrilotriacetic acid affinity chromatography. A second purification was executed with size-exclusion chromatography with a Superdex 200 column. Protein concentration was measured using a bicinchoninic acid (BCA) protein assay kit (Bio-Rad).
[0236] Formulation
[0237] Cholera toxin (CT) was used as adjuvant for intranasal immunization to facilitate induction of T-cell responses to proteins alone. Proteins or pneumococcal whole cell vaccine (amount indicated in Table 5) were mixed with 1 .mu.g of CT in saline solution in a final volume of 20 .mu.l per dose prior to administration.
[0238] Intranasal Mouse Immunization Protocol
[0239] For intranasal immunization with CT adjuvanted proteins and pneumococcal whole cell vaccine with chloroform inactivation (WCC), C57BL/6 mice (groups of n=10) received 2 immunizations 1 week apart. Peripheral blood samples were taken 3 weeks after the last immunization for ex vivo IL-17A stimulation in the presence of appropriate antigen(s) as stimulant.
TABLE-US-00005 TABLE 5 Mouse intranasal immunization study groups Dose # of Immunization Schedule Blood Collection Group Antigen (Protein) Adjuvant Mice (days) (days) A -- -- CT 10 0, 7 28 B SP0785 10 .mu.g CT 10 0, 7 28 C -- -- CT 10 0, 7 28 D SP1500 10 .mu.g CT 10 0, 7 28 E CP1 15 .mu.g CT 10 0, 7 28 F Rhavi 15 .mu.g CT 10 0, 7 28 G WCC 100 .mu.g CT 10 0, 7 28 Abbreviations: CP1: fusion protein 1 (Rhavi-linker (GGGGSSS)-SP1500-linker (AAA)-SP0785); Rhavi: truncated rhizavidin, amino acids [45-179]; WCC: pneumococcal whole cell vaccine (chloroform inactivation); CT: cholera toxin Note: all recombinant proteins were His-tagged.
[0240] IL-17A Induction and Measurement from Whole Blood
[0241] Ex vivo stimulation of peripheral blood samples taken 3 weeks after the last intranasal immunization was performed in 96-well round-bottom plates. All stimulants were diluted in stimulation medium (DMEM F-12; 10% FBS, 50 .mu.M 2-mercaptoethanol, 10 .mu.g/ml ciprofloxacin) at a final concentration of 10 .mu.g/ml. In each well, 25 .mu.l of heparinized blood was added to 225 .mu.l of stimulation medium containing indicated stimulants, followed by incubation at 37.degree. C. with 5% CO.sub.2 for 6 days. Supernatants were collected after centrifugation, and IL-17A was analyzed with an ELISA kit (R&D systems).
[0242] Nasopharyngeal S. pneumoniae Infection and Measurement of Colonization
[0243] One to 2 weeks after blood collection, mice were intranasally challenged with 10.sup.7 CFU of type 6B pneumococci (603 strain). Nasopharyngeal wash on euthanized mice was conducted 7 days post infection. The S. pneumoniae CFU per nasal wash were calculated after growth on blood agar plates.
[0244] Statistical Analyses
[0245] Statistical analyses were performed using PRISM (GraphPad Software). All data on IL-17A concentration and nasopharyngeal colonization densities were analyzed using the Mann-Whitney U test. The geometric mean concentrations of IL-17A was calculated for each group, and the geometric mean density of colonization was calculated for each group.
[0246] Results and Discussion:
[0247] IL-17A Response and Reduction in Colonization after Intranasal Immunization with SP1500, SP0785, or CP1.
[0248] As seen in FIG. 2, left panel, intranasal immunization with either SP0785 or SP1500 adjuvanted with CT induced a strong antigen-specific Th17 response compared to immunization with CT alone, as indicated by increased IL-17A production after ex vivo stimulation of peripheral blood with purified SP0785 or SP1500. The increased IL-17A secretion correlated with a corresponding statistically significant reduction in S. pneumoniae CFU recovered from the nasopharyngeal wash 7 days after challenge of immunized mice (FIG. 2, right panel).
[0249] As seen in FIG. 3, left panel, when Rhavi protein was compared to the protein fusion of Rhavi-linker (GGGGSSS)-SP1500-linker (AAA)-SP0785 (CP1) after intranasal immunization with CT, CP1 retained the antigen specific induction of IL-17A. When these mice were challenged with S. pneumoniae and compared to pneumococcal whole cell vaccine intranasal immunization, Rhavi alone had no protective effect as measured by pneumococcal CFU, while CP1 had a protective effect that was comparable to killed (inactivated) pneumococcal whole cells (FIG. 3, right panel).
Example 2: Killing Activity of Anti-Sera Against Fusion Protein CP1
[0250] Materials and Methods:
[0251] Growth of Bacteria
[0252] S. pneumoniae strains 6B, 15A, 16F, and 35B were inoculated into 10 mL cultures of Todd Hewitt Broth (THB) with yeast extract. Cultures were incubated at 37.degree. C. in 5% CO.sub.2 for 4-7 hours until the OD.sub.600 reached 0.5-0.8 (mid-logarithmic phase). The bacteria were harvested by spinning for 7 minutes at 3,000 g at 4.degree. C. and the bacterial pellet was resuspended in 10 mL of THB with 10% glycerol with storage at -80.degree. C. Colony forming unit (CFU) estimations were determined by serially dilutions of frozen stocks on Trypticase soy agar with 5% sheep blood (Becton, Dickenson, and Company) with 37.degree. C. in 5% CO.sub.2 incubation for 18 to 24 hrs.
[0253] Concentrated Opsonophagocytic Assay (COPA)
[0254] Frozen stocks of S. pneumoniae were thawed and resuspended at 2.times.10.sup.5 CFU/ml in assay buffer (Hank's buffered saline with 10% heat inactivated FBS). To a 96 well plate, 10 .mu.l of bacteria suspension was added to each well followed by 20 .mu.l of heat inactivated rabbit serum diluted in assay buffer to be tested in the assay. The bacteria and rabbit serum were incubated at room temperature for 30 min with shaking. To each well, 10 .mu.l of baby rabbit complement (Pel-Freeze Biologicals) was added followed by incubation at room temperature for 30 min with shaking. HL60 cells (ATCC) were washed with assay buffer and resuspend to 1.times.10.sup.7 cells/ml. To each well, 40 .mu.l of HL60 suspension was added (200:1 HL60 to bacteria ratio) followed by incubation with shaking at 37.degree. C. with 5% CO.sub.2 for 1 hour. The plate was transferred to ice and incubated for 20 minutes. Each sample (undiluted, 1/5 and 1/25 dilutions in water) was then plated on 5% blood agar plates. After overnight incubation at 37.degree. C. with 5% CO.sub.2, the CFU were counted for each sample and dilution.
[0255] Rabbit Serum
[0256] New Zealand White rabbits (n=3) were immunized with 100 ug of Rhavi-linker (GGGGSSS)-SP1500-linker (AAA)-SP0785(CP1, His-tagged) and 0.625 mg of elemental aluminum from AlPO.sub.4 per dose for a total of three doses with two weeks between each immunization (rabbits 87, 88, and 1762). Sera were collected prior to immunization (P0) and two weeks after the third immunization (P3) and stored at -80.degree. C.
[0257] Results and Discussion:
[0258] S. pneumoniae Opsonophagocytic Activity of Antibodies Directed Against CP1
[0259] A modified concentrated opsonophagocytic assay (COPA) was established to investigate protein antibody-mediated killing of S. pneumoniae. Sera from two rabbits immunized with CP1 (P3 from rabbits 87 and 88) were assayed in comparison to the pre-immune sera (P0). At all dilutions tested for both rabbits, incubation with the P3 immune serum resulted in a reduction of the CFU (FIG. 4), or in alternative display of the same results, an increase in percent killing activity (FIG. 7, Panel A), compared to incubation with the pre-immune serum (P0) for S. pneumoniae serotype 6B, a type incorporated into the commercially available Prevnar 13 vaccine. The killing activity of the immune serum was dependent on both HL60 cells and active complement (data not shown). The same sera were assayed on two serotypes not incorporated into commercially available vaccines, S. pneumoniae serotype 15A (FIG. 5 and FIG. 7, Panel B: 1/2 dilution) and serotype 35B (FIG. 6 and FIG. 7, Panel C: 1/2 dilution). Serum from a third rabbit immunized with CP1 (P3 from rabbit 1762) was assayed in comparison to the pre-immune serum (P0) against S. pneumoniae serotype 16F, a further serotype not incorporated into commercially available vaccines (FIG. 7, Panel D: 1/2, 1/6, 1/18, 1/54 dilutions). For all rabbits and non-vaccine serotypes tested, incubation with the P3 immune sera resulted in a reduction of the CFU, or alternatively displayed, an increase in percent killing activity, compared to incubation with the pre-immune sera (P0). The killing activity of the immune sera was dependent on both HL60 cells and active complement (data not shown).
Example 3: Comparison of Induction of Th17 Response in Mice Following Immunization with a Mixture (Unconjugated) of SP0785, SP1500, and a Truncated Rhizavidin Protein Rhavi, or with Fusion Proteins CP1 or SP0785-Linker (SSSGG)-SP1500-Linker (SSVDKL)-PdT
[0260] Materials and Methods:
[0261] Recombinant Protein Production
[0262] Histidine-tagged recombinant proteins were expressed in Escherichia coli and purified using Ni-nitrilotriacetic acid affinity chromatography. A second purification was executed with size-exclusion chromatography with a Superdex 200 column. Protein concentration was measured using a bicinchoninic acid (BCA) protein assay kit (Bio-Rad).
[0263] Formulation
[0264] Cholera toxin (CT) was used as adjuvant for intranasal immunization to facilitate induction of T-cell responses to proteins alone. Proteins (amount indicated in Table 6) were mixed with 1 .mu.g of CT in saline solution to a final volume of 20 .mu.l per dose prior to administration.
[0265] Intranasal Mouse Immunization Protocol
[0266] For intranasal immunization with CT adjuvanted proteins, C57BL/6 mice (groups of n=15) received 2 immunizations 1 week apart. Peripheral blood samples were taken 3 weeks after the last immunization for ex vivo IL-17A stimulation in the presence of appropriate antigen(s) as stimulant.
TABLE-US-00006 TABLE 6 Mouse intranasal immunization study groups Blood Dose # of Immunization Collection Group Antigen (Protein) Adjuvant Mice Schedule (days) (days) A -- -- CT 15 0, 7 28 B CP1 10 .mu.g CT 15 0, 7 28 C Rhavi + SP0785 + 10 .mu.g CT 15 0, 7 28 SP1500 mixture D SP0785-linker 17 .mu.g CT 15 0, 7 28 (SSSGG)-SP1500- linker (SSVDKL)-PdT E SP0785 10 .mu.g CT 15 0, 7 28 F SP1500 10 .mu.g CT 15 0, 7 28 Abbreviations: CP1: fusion protein 1 (Rhavi-linker (GGGGSSS)-SP1500-linker (AAA)-SP0785); Rhavi: truncated rhizavidin, amino acids [45-179]; CT: cholera toxin Note: all recombinant proteins were His-tagged.
[0267] IL-17A Induction and Measurement from Whole Blood
[0268] Ex vivo stimulation of peripheral blood samples taken 3 weeks after the last intranasal immunization was performed in 96-well round-bottom plates. All stimulants (purified proteins or killed (inactivated) pneumococcal whole cells) were diluted in stimulation medium (DMEM F-12; 10% FBS, 50 .mu.M 2-mercaptoethanol, 10 .mu.g/ml ciprofloxacin) at a final concentration of 10 .mu.g/ml. In each well, 25 .mu.l of heparinized blood was added to 225 .mu.l of stimulation medium containing indicated stimulants, followed by incubation at 37.degree. C. with 5% CO.sub.2 for 6 days. Supernatants were collected after centrifugation, and IL-17A was analyzed with an ELISA kit (R&D systems).
[0269] Statistical Analyses
[0270] Statistical analyses were performed using PRISM (GraphPad Software). All data on IL-17A concentration were analyzed using the Mann-Whitney U test. The geometric mean concentrations of IL-17A was calculated for each group.
[0271] Results and Discussion:
[0272] IL-17A Responses after Intranasal Immunization with a Mixture of SP 1500, SP0785, and Rhavi Proteins, or with CP1 or SP0785-Linker (SSSGG)-SP1500-Linker (SSVDKL)-PdT.
[0273] As seen in FIG. 8, intranasal immunization with CP1 adjuvanted with CT induced a stronger antigen-specific Th17 response compared to immunization with SP0785 or SP1500 adjuvanted with CT, or with CT alone (control). The Th17 response is indicated by increased IL-17A production after ex vivo stimulation of peripheral blood of immunized mice with purified SP0785 (Panel A), purified SP1500 (Panel B), or killed (inactivated) pneumococcal whole cells (WCV; Panel C).
[0274] As seen in FIG. 9, intranasal immunization with CP1 adjuvanted with CT also induced a stronger antigen-specific Th17 response compared to immunization with fusion protein SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT adjuvanted with CT, or with CT alone (control). The Th17 response is indicated by increased IL-17A production after ex vivo stimulation of peripheral blood of immunized mice with purified SP0785 (Panel A), or purified SP1500 (Panel B).
[0275] As seen in FIG. 10, intranasal immunization with CP1 adjuvanted with CT induced a stronger antigen-specific Th17 response compared to immunization with a combination (mixture) of SP0785, SP1500, and Rhavi adjuvanted with CT, or with CT alone (control). The Th17 response is indicated by increased IL-17A production after ex vivo stimulation of peripheral blood of immunized mice with purified SP0785 (Panel A), purified SP1500 (Panel B), or killed (inactivated) pneumococcal whole cells (WCV; Panel C).
Example 4: Comparison of Hemolytic Activity of CP1 and SP0785-Linker (SSSGG)-SP1500-Linker (SSVDKL)-PdT
[0276] Materials and Methods:
[0277] Recombinant Protein Production
[0278] Histidine-tagged recombinant fusion proteins CP1 and SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT were expressed in Escherichia coli and purified using Ni-nitrilotriacetic acid affinity chromatography. A second purification was executed with size-exclusion chromatography with a Superdex 200 column. Protein concentration was measured using a bicinchoninic acid (BCA) protein assay kit (Bio-Rad).
[0279] Assay for Hemolytic Activity of Fusion Proteins
[0280] Assay was adapted from Benton et al, 1997. Assay buffer contained 10 mM Dithiothreitol, 0.1% Bovine Serum Albumin in PBS pH 7.4, and 2% sheep red blood cells. Sheep red blood cells were prepared as follows: add 200 .mu.l sheep blood+1 ml PBS pH 7.4, mix well, pellet, and wash 3.times. at 8,000 rpm for 30 sec each; finally resuspend blood cells in 10 ml chilled PBS and keep on ice until used. Assay was performed by diluting a pneumolysin standard (Ply), pneumolysoid PdT, and fusion proteins to test (CP1 and SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT) at the indicated concentrations across the plate with 100 .mu.l/well, then adding 50 .mu.l of 2% sheep red blood cells to all wells. Plate was incubated for 30 min at 37.degree. C. After incubation, plate was centrifuged for 5 min at 2,000 rpm at room temperature and 100 .mu.l of supernatant was transferred to an empty 96 well plate to measure absorbance at OD.sub.420. [Benton, K. A., J. C. Paton, and D. E. Briles. 1997. Differences in virulence for mice among Streptococcus pneumoniae strains of capsular types 2, 3, 4, 5, and 6 are not attributable to differences in pneumolysin production. Infect Immun. 65:1237-44.]
[0281] Results and Discussion:
[0282] As seen in FIG. 11, incubation of sheep red blood cells with fusion protein CP1 did not result in hemolysis at any concentration tested. Incubation with fusion protein SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT or pneumolysoid PdT alone, at concentrations above 1 mg/ml and 0.5 mg/ml respectively, resulted in nearly complete hemolysis of sheep red blood cells. These results show that SP0785 and SP1500 moieties of the two fusion proteins do not contribute to hemolytic activity. Hemolytic activity of fusion protein SP0785-linker (SSSGG)-SP1500-linker (SSVDKL)-PdT is attributable instead to the pneumolysoid PdT.
TABLE-US-00007 Sequences SEQ ID NO: 1, rhizavidin protein, full-length [amino acids 1-179]: MIITSLYATFGTIADGRRTSGGKTMIRTNAVAALVFAVATSALAFDASNFKDFSSIASASSSWQ NQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYPLTGRVNGTFIAFSVGWNNSTENCNSATG WTGYAQVNGNNTEIVTSWNLAYEGGSGPAIEQGQDTFQYVPTTENKSLLKD SEQ ID NO: 2, truncated rhizavidin protein, denoted Rhavi [amino acids 45-179]: FDASNFKDFSSIASASSSWQNQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYPLTGRVNGT FIAFSVGWNNSTENCNSATGWTGYAQVNGNNTEIVTSWNLAYEGGSGPAIEQGQDTFQYVPTTE NKSLLKD SEQ ID NO: 3, SP0785 protein, full-length [amino acids 1-399], TIGR4 strain: MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVV AKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSVGDKVSEGQAL VKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGG EDATVQSPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVL STLEGTVVEVNSNVSKSPTGASQVMVHIVSNENLQVKGELSEYNLANLSV GQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTI DVTGEVGDLKQGFSVNIEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQQK AKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVKADEATN SEQ ID NO: 4, SP0785 protein lacking signal sequence [amino acids 33-399]: Note: One T394A mismatch with SP0785 NCBI Sequences ABJ54007.1 and YP816180 FRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSVGDKVSE GQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDATVQSPTP VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQV MVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAA GNNTGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQQKAK KVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVKADEATN SEQ ID NO: 5, consensus SP0785 protein [amino acids 1-399]: (SEQ ID NOS 110-134, respectively, in order of appearance): WP 081570978 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 054387396 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLIVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 097557828 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 000728643 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 061633543 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 050965059 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHFVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 055387306 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 050203943 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 000728633 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPIHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 088799985 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 061764363 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 050259582 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 023396621 1 MKKKNGKAKKWQLYAAIGAASVVILGAGGILLFRQPSQTALKDEPIHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 000728632 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDELTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 061366281 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 061743315 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 057525500 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPIHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 000728639 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 084572368 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 084354434 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPYQTALKDEPTHLVVVKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 050214972 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTAVKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 050208881 1 MKKKNGKAKKWQLYAAIGAASVVILGAGGILLFRQPSQTALKDEPIHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 088793209 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 069123032 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVAKEGSVASSILLSGTVTAKNEQY- VYFDASK 80 WP 000728647 1 MKKKNGKAKKWQLYAAIGAASVVVLGAGGILLFRQPSQTALKDEPTHLVVANEGSVASSVLLSGTVTAKNEQY- VYFDASK 80 WP 081570978 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 054387396 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 097557828 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQNPTP 160 WP 000728643 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 061633543 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPTPQLPAPVGGEDA- TVQSPTP 160 WP 050965059 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 055387306 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 050203943 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 000728633 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 088799985 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 061764363 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASALAPQLPAPVGGEDA- TVQSPTP 160 WP 050259582 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVPSPTP 160 WP 023396621 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 000728632 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 061366281 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 061743315 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVAKADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 057525500 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 000728639 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVAKADRHINELNQARNEAASAQAPQLPAPVGGEDA- TVQSPTP 160 WP 084572368 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARVDCHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 084354434 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 050214972 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVAKADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 050208881 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVAKADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 088793209 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAVYDSASRAVAKADRHINELNQARNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 069123032 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVAKADRHINELNQVRNEAASAPAPQLPAPVGGEDA- TVQSPTP 160 WP 000728647 81 GDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVAKADRHINELNQARNEAASAQAPQLPAPVGGEDA- TVQSPTP 160 WP 081570978 161 VAGNSVASIDAQLGDAFDAFADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIVSNEN- LQVKGEL 240 WP 054387396 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKTVSNEN- LQVKGEL 240 WP 097557829 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN-
1QVKGEL 240 WP 000728643 161 VAGNSVASIDAQLGDARDARAOAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 061633543 161 VAGNSVASIDAQLGDAFDAPADAAAQLSKAOSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIV5NEN- LQVKGEL 240 WP 050965059 161 VAGNSVASIDAQLGDAFDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIVSNEN- LQVKGEL 240 WP 055387306 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 050203943 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- 1QVKGEL 240 WP 000728633 161 VAGNSVASIDAQLGDAFDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIVSNEN- LQVKGEL 240 WP 088799985 161 VAGNSVASIDAQLGDAPDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 061764363 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIVSNEN- LQVKGEL 240 WP 050259582 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVHVHIVSNEN- LQVKGEL 240 WP 023396621 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVHVHIVSNEN- LQVKGEL 240 WP 000728632 161 VAGNSVASIDAQLGDAFDAPADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIVSNEN- LQVKGEL 240 WP 061366281 161 VAGNSVASIDAQLGDAFDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVHVHIVSNEN- LQVKGEL 240 WP 061743315 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 057525500 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVHVHIVSNEN- LQVKGEL 240 WP 000728639 161 VAGNSVASIDAQLGDAFDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIVSNEN- LQVKGEL 240 WP 084572368 161 VAGNSVASIDAQLGDAPDAFADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVKIVSNEN- LQVKGEL 240 WP 084354434 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 050214972 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 050208881 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 088793209 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 069123032 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 000728647 161 VAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNEN- LQVKGEL 240 WP 081570978 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 054387396 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 097557828 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 000728643 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 061633543 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 050965059 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 055387306 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 050203943 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGAAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 000728633 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 088799985 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEGGDLKQGF- SVNIEVK 320 WP 061764363 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 050259582 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 023396621 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 000728632 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 061366281 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPASGNNTGSKYPYTIDVTGEIGDLKQGF- SVNIEVK 320 WP 061743315 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGRLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 057525500 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPASGNNTGSKYPYTIDVTGEIGDLKQGF- SVNIEVK 320 WP 000728639 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 084572368 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 084354434 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 050214972 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGNLKQGF- SVNIEVK 320 WP 050208881 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 088793209 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 069123032 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 000728647 241 SEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGF- SVNIEVK 320 WP 081570978 321 SKTKAILVPVSSLVMDNSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 054387396 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 097557828 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 000728643 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKSKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 061633543 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 050965059 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 055387306 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTFSLEEGKEVK- ADEATN 399 WP 050203943 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 000728633 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 088799985 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 061764363 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 050259582 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 023396621 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 000728632 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 061366281 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 061743315 321
SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 057525500 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 000728639 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 084572368 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 084354434 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 050214972 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 050208881 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 088793209 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- TDEATN 399 WP 069123032 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 WP 000728647 321 SKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK- ADEATN 399 SEQ ID NO: 6, SP1500 protein, full-length [amino acids 1-278], TIGR4 strain: MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGF DIDLATAVFEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQV LVTKKSSGITTAKDMTGKTLGAQAGSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKND RIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGK FQEISQKWFGEDVATKEVKEGQ SEQ ID NO: 7, SP1500 protein lacking signal sequence [amino acids 27-278]: TSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDIDLATAVFEKYGITVNWQPIDWDLK EAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTGKTLGAQAGS SGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYN VFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ SEQ ID NO: 8, consensus SP1500 protein [amino acids 1-278] (SEQ ID NOS 6 and 135-157, respectively, in order of appearance): WP 000759187 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 050213573 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 061814735 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 084843602 1 --------MKKWMHVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 000759185 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 050261378 1 --------MKKWMLVLVSLMTALFLVACGKNTSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 050220771 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 CKA82396 1 ----mkkwMKKWMLVLVSLMTALFLVACGKNTSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 76 WP 088802838 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 049512265 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 CKL85404 1 ----mkkwMKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 76 EHZ28755 1 ------------MLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 68 WP 100128002 1 --------MKKWILVLVSLMTALFLVACGKNSSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 053039665 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 101514844 1 --------MKKWILVLVSLMTALFLVACGKNSSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 050242061 1 --------MKKWMLVLVSLMTALFLVACGKNASEISGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 023941000 1 --------MKKWMFVLVSLMTALFLVACGKNASETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 EHE15463 1 ------------MLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 68 EHE34295 1 ------------MLVLVSLMTALFLVACGKNSSETSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 68 CCM08008 1 --------MKKWMLVLVSLMTALFLVACGKNSSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 KGI27253 1 ----mkkwMKKWMLVLVSLMTALFLVACGKNASETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 76 WP 088777969 1 --------MKKWMLVLVSLMTALFLVACGKNASETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 WP 050223531 1 --------MKKWMLVLVSLMIALFLVACGKNTSETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 72 CKJ33697 1 mkkwmkkwMKKWMLVLVSLMTALFLVACGKNASETSGDNWSKYESNKSITIGFDSTFVPMGFAQKDGSYAGFD- IDLATAV 80 WP 000759187 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTG- KTLGAQA 152 WP 050213573 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTG- KTLGAQA 152 WP 061814735 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTG- KTLGAQA 152 WP 084843602 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMTG- KTLGAQA 152 WP 000759185 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMTG- KTLGAQA 152 WP 050261378 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 WP 050220771 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 CKA82396 77 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 156 WP 088802838 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMTG- KTLGAQA 152 WP 049512265 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 CKL85404 77 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMNG- KTLGAQA 156 EHZ28755 69 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTG- KTLGAQA 148 WP 100128002 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 WP 053039665 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATYERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMTG- KTLGAQA 152 WP 101514844 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSHSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 WP 050242061 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 WP 023941000 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 EHE15463 69 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTG- KTLGAQA 148 EHE34295 69 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTG- KTLGAQA 148 CCM08008 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATYERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMTG- KTLGAQA 152 KGI27253 77 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMTG- KILGAQA 156 WP 088777969 73 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 WP 050223531 73 FEKYGITINWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSHSYMKNEQVLVTKKSSGITTAKDMAG- KTLGAQA 152 CKJ33697 81 FEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKLSGITTAKDMTG- KILGAQA 160 WP 000759187 153 GSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE-
TEAFAVG 232 WP 050213573 153 GSSGYADFEANPELLKNIVANKEVNQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 061814735 153 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 084843602 153 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 000759185 153 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 050261378 153 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 050220771 153 GSSGYADFEANPAILKDIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 CKA82396 157 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 236 WP 088802838 153 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 049512265 153 GSSGYADFEANPAILKDIVANKEANQYQTFNEALIDLKNDRIEGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 CKL85404 157 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 236 EHZ28755 149 GSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 228 WP 100128002 153 GSSGYADFEANPAILKDIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 053039665 153 GSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 101514844 153 GSSGYADFEANPAILKDIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 050242061 153 GSSGYADFEANPAILKDIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 023941000 153 GSSGYADFEANPAILKDIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 EHE15463 149 GSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 228 EHE34295 149 GSSGYADFEANPELLKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 228 CCM08008 153 GSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 KGI27253 157 GSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- IEAFAVG 236 WP 088777969 153 GSSGYADFEANPAILKDIVANKEANQYQIFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 WP 050223531 153 GSSGYADFEANPAILKDIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- TEAFAVG 232 CKJ33697 161 GSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE- IEAFAVG 240 WP 000759187 233 ARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 050213573 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 061814735 233 ARKEDTTLVKKINEDFSSLYKNGKFQEISQKWFGEDVATKEVKEGQ 278 WP 084843602 233 ARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 000759185 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 050261378 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 050220771 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 CKA82396 237 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 282 WP 088802838 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEGVATKEVKEGQ 278 WP 049512265 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 CKL85404 237 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 282 EHZ28755 229 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 274 WP 100128002 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 053039665 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 101514844 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 050242061 233 TRKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 023941000 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 EHE15463 229 SRKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 274 EHE34295 229 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 274 CCM08008 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEG- 277 KGI27253 237 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 282 WP 088777969 233 ARKEDITLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 WP 050223531 233 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 278 CKJ33697 241 ARKEDTTLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ 286 SEQ ID NO: 9, rhizavidin gene encoding full-length rhizavidin protein: TTGATTATTACGAGTTTATATGCAACCTTCGGTACGATCGCCGACGGTCGGCGCACATCAGGAG GAAAAACCATGATCCGTACTAATGCAGTTGCAGCACTTGTGTTTGCTGTAGCAACCTCAGCTCT TGCTTTCGATGCAAGCAACTTCAAGGATTTTTCAAGCATCGCATCGGCTTCCAGCAGTTGGCAG AACCAGTCCGGCTCGACGATGATCATTCAAGTCGACTCGTTCGGAAATGTCTCCGGCCAATATG TAAACAGAGCCCAGGGCACCGGATGCCAGAACTCGCCCTATCCGCTAACAGGAAGGGTAAACGG GACGTTCATCGCATTTTCGGTCGGCTGGAACAATTCGACGGAGAACTGCAATTCCGCAACCGGA TGGACCGGCTACGCACAGGTCAACGGCAACAACACTGAGATAGTCACGAGCTGGAACCTCGCTT ACGAAGGCGGCTCCGGTCCGGCCATTGAGCAAGGACAAGACACTTTCCAGTACGTGCCGACGAC TGAGAACAAAAGCCTCTTGAAGGATTAA SEQ ID NO: 10, rhizavidin gene encoding truncated rhizavidin protein, denoted Rhavi [amino acids 45-179]: TTTGATGCGAGTAACTTTAAAGATTTCAGCTCTATTGCGAGCGCCAGTAGCTCTTGGCAGAATC AGAGTGGCAGCACCATGATTATCCAGGTGGATTCTTTCGGCAACGTTAGTGGCCAGTATGTTAA TCGTGCGCAGGGCACGGGTTGCCAGAACTCTCCGTACCCGCTGACCGGCCGCGTGAATGGCACG TTTATCGCCTTCAGCGTTGGCTGGAACAATTCTACCGAAAACTGTAATAGTGCAACCGGCTGGA CGGGTTATGCGCAGGTGAACGGTAACAATACCGAAATTGTTACGAGCTGGAATCTGGCCTATGA AGGCGGTTCTGGCCCGGCAATCGAACAGGGTCAGGATACCTTTCAGTACGTTCCGACCACGGAA AACAAAAGCCTGCTGAAAGAT SEQ ID NO: 11, SP0785 gene encoding full-length SP0785 protein [amino acids 1-399], TIGR4 strain: ATGAAGAAAAAGAATGGTAAAGCTAAAAAGTGGCAACTGTATGCAGCAAT CGGTGCTGCGAGTGTAGTTGTATTGGGTGCTGGGGGGATTTTACTCTTTA GACAACCTTCTCAGACTGCTCTAAAAGATGAGCCTACTCATCTTGTTGTT GCCAAGGAAGGAAGCGTGGCCTCCTCTGTTTTATTGTCAGGGACAGTAAC AGCAAAAAATGAACAATATGTTTATTTTGATGCTAGTAAGGGTGATTTAG ATGAAATCCTTGTTTCTGTGGGCGATAAGGTCAGCGAAGGGCAGGCTTTA GTCAAGTACAGTAGTTCAGAAGCGCAGGCGGCCTATGATTCAGCTAGTCG AGCAGTAGCTAGGGCAGATCGTCATATCAATGAACTCAATCAAGCACGAA ATGAAGCCGCTTCAGCTCCGGCTCCACAGTTACCAGCGCCAGTAGGAGGA GAAGATGCAACGGTGCAAAGCCCAACTCCAGTGGCTGGAAATTCTGTTGC TTCTATTGACGCTCAATTGGGTGATGCCCGTGATGCGCGTGCAGATGCTG CGGCGCAATTAAGCAAGGCTCAAAGTCAATTGGATGCAACAACTGTTCTC AGTACCCTAGAGGGAACTGTGGTCGAAGTCAATAGCAATGTTTCTAAATC TCCAACAGGGGCGAGTCAAGTTATGGTTCATATTGTCAGCAATGAAAATT TACAAGTCAAGGGAGAATTGTCTGAGTACAATCTAGCCAACCTTTCTGTA GGTCAAGAAGTAAGCTTTACTTCTAAAGTGTATCCTGATAAAAAATGGAC TGGGAAATTAAGCTATATTTCTGACTATCCTAAAAACAATGGTGAAGCAG CTAGTCCAGCAGCCGGGAATAATACAGGTTCTAAATACCCTTATACTATT GATGTGACAGGCGAGGTTGGTGATTTGAAACAAGGTTTTTCTGTCAACAT TGAGGTTAAAAGCAAAACTAAGGCTATTCTTGTTCCTGTTAGCAGTCTAG TAATGGATGATAGTAAAAATTATGTCTGGATTGTGGATGAACAACAAAAG GCTAAAAAAGTTGAGGTTTCATTGGGAAATGCTGACGCAGAAAATCAAGA AATCACTTCTGGTTTAACGAACGGTGCTAAGGTCATCAGTAATCCAACAT CTTCCTTGGAAGAAGGAAAAGAGGTGAAGGCTGATGAAGCAACTAAT SEQ ID NO: 12, SP0785 gene encoding SP0785 protein lacking signal sequence [amino acids 33-399]: TTTAGACAACCTTCTCAGACTGCTCTAAAAGATGAGCCTACTCATCTTGTTGTTGCCAAGGAAGGAAGCGTGGC- CTC CTCTGTTTTATTGTCAGGGACAGTAACAGCAAAAAATGAACAATATGTTTATTTTGATGCTAGTAAGGGTGATT- TAG ATGAAATCCTTGTTTCTGTGGGCGATAAGGTCAGCGAAGGGCAGGCTTTAGTCAAGTACAGTAGTTCAGAAGCG- CAG GCGGCCTATGATTCAGCTAGTCGAGCAGTAGCTAGGGCAGATCGTCATATCAATGAACTCAATCAAGCACGAAA- TGA AGCCGCTTCAGCTCCGGCTCCACAGTTACCAGCGCCAGTAGGAGGAGAAGATGCAACGGTGCAAAGCCCAACTC- CAG TGGCTGGAAATTCTGTTGCTTCTATTGACGCTCAATTGGGTGATGCCCGTGATGCGCGTGCAGATGCTGCGGCG- CAA TTAAGCAAGGCTCAAAGTCAATTGGATGCAACAACTGTTCTCAGTACCCTAGAGGGAACTGTGGTCGAAGTCAA- TAG CAATGTTTCTAAATCTCCAACAGGGGCGAGTCAAGTTATGGTTCATATTGTCAGCAATGAAAATTTACAAGTCA- AGG GAGAATTGTCTGAGTACAATCTAGCCAACCTTTCTGTAGGTCAAGAAGTAAGCTTTACTTCTAAAGTGTATCCT- GAT AAAAAATGGACTGGGAAATTAAGCTATATTTCTGACTATCCTAAAAACAATGGTGAAGCAGCTAGTCCAGCAGC- CGG GAATAATACAGGTTCTAAATACCCTTATACTATTGATGTGACAGGCGAGGTTGGTGATTTGAAACAAGGTTTTT- CTG TCAACATTGAGGTTAAAAGCAAAACTAAGGCTATTCTTGTTCCTGTTAGCAGTCTAGTAATGGATGATAGTAAA- AAT TATGTCTGGATTGTGGATGAACAACAAAAGGCTAAAAAAGTTGAGGTTTCATTGGGAAATGCTGACGCAGAAAA- TCA AGAAATCACTTCTGGTTTAACGAACGGTGCTAAGGTCATCAGTAATCCAACATCTTCCTTGGAAGAAGGAAAAG- AGG TGAAGGCTGATGAAGCAACTAAT SEQ ID NO: 13, SP1500 gene encoding full-length SP1500 protein [amino acids 1-278], TIGR4 strain: ATGAAAAAATGGATGCTTGTATTAGTCAGTCTGATGACTGCTTTGTTCTT AGTAGCTTGTGGGAAAAATTCTAGCGAAACTAGTGGAGATAATTGGTCAA AGTACCAGTCTAACAAGTCTATTACTATTGGATTTGATAGTACTTTTGTT CCAATGGGATTTGCTCAGAAAGATGGTTCTTATGCAGGATTTGATATTGA TTTAGCTACAGCTGTTTTTGAAAAATACGGAATCACGGTAAATTGGCAAC CGATTGATTGGGATTTGAAAGAAGCTGAATTGACAAAAGGAACGATTGAT CTGATTTGGAATGGCTATTCCGCTACAGACGAACGCCGTGAAAAGGTGGC TTTCAGTAACTCATATATGAAGAATGAGCAGGTATTGGTTACGAAGAAAT CATCTGGTATCACGACTGCAAAGGATATGACTGGAAAGACATTAGGAGCT CAAGCTGGTTCATCTGGTTATGCGGACTTTGAAGCAAATCCAGAAATTTT GAAGAATATTGTCGCTAATAAGGAAGCGAATCAATACCAAACCTTTAATG AAGCCTTGATTGATTTGAAAAACGATCGAATTGATGGTCTATTGATTGAC CGTGTCTATGCAAACTATTATTTAGAAGCAGAAGGTGTTTTAAACGATTA TAATGTCTTTACAGTTGGACTAGAAACAGAAGCTTTTGCGGTTGGAGCCC GTAAGGAAGATACAAACTTGGTTAAGAAGATAAATGAAGCTTTTTCTAGT CTTTACAAGGACGGCAAGTTCCAAGAAATCAGCCAAAAATGGTTTGGAGA AGATGTAGCACCAAAGAAGTAAAAGAAGGACAG SEQ ID NO: 14, SP1500 gene encoding SP1500 protein lacking signal sequence [amino acids 27-278]: ACTAGTGGAGATAATTGGTCAAAGTACCAGTCTAACAAGTCTATTACTATTGGATTTGATAGTACTTTTGTTCC- AAT
GGGATTTGCTCAGAAAGATGGTTCTTATGCAGGATTTGATATTGATTTAGCTACAGCTGTTTTTGAAAAATACG- GAA TCACGGTAAATTGGCAACCGATTGATTGGGATTTGAAAGAAGCTGAATTGACAAAAGGAACGATTGATCTGATT- TGG AATGGCTATTCCGCTACAGACGAACGCCGTGAAAAGGTGGCTTTCAGTAACTCATATATGAAGAATGAGCAGGT- ATT GGTTACGAAGAAATCATCTGGTATCACGACTGCAAAGGATATGACTGGAAAGACATTAGGAGCTCAAGCTGGTT- CAT CTGGTTATGCGGACTTTGAAGCAAATCCAGAAATTTTGAAGAATATTGTCGCTAATAAGGAAGCGAATCAATAC- CAA ACCTTTAATGAAGCCTTGATTGATTTGAAAAACGATCGAATTGATGGTCTATTGATTGACCGTGTCTATGCAAA- CTA TTATTTAGAAGCAGAAGGTGTTTTAAACGATTATAATGTCTTTACAGTTGGACTAGAAACAGAAGCTTTTGCGG- TTG GAGCCCGTAAGGAAGATACAAACTTGGTTAAGAAGATAAATGAAGCTTTTTCTAGTCTTTACAAGGACGGCAAG- TTC CAAGAAATCAGCCAAAAATGGTTTGGAGAAGATGTAGCAACCAAAGAAGTAAAAGAAGGACAG SEQ ID NO: 15, His tag 1: HHHHHH SEQ ID NO: 16, His tag 2: MSYYHHHHHH SEQ ID NO: 17, fusion protein SP1500-SP0785: MTSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDIDLATAVFEKYGITVNWQPIDWDL KEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTGKTLGAQAG SSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDY NVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQFRQ PSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSVGDKVSEGQA LVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDATVQSPTPVAG NSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVH IVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNN TGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVE VSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVKADEATN SEQ ID NO: 18, fusion protein SP0785-SP1500: MFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSVGDKVS EGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDATVQSPT PVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQ VMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPA AGNNTGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQQKA KKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVKADEATNTSGDNWSKYQSNKSIT IGFDSTFVPMGFAQKDGSYAGFDIDLATAVFEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGY SATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTGKTLGAQAGSSGYADFEANPEILKNI VANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLETEAFAVGAR KEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ SEQ ID NO: 19, fusion protein Rhavi-SP1500-SP0785: MFDASNFKDFSSIASASSSWQNQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYPLTGRVNG TFIAFSVGWNNSTENCNSATGWTGYAQVNGNNTEIVTSWNLAYEGGSGP AIEQGQDTFQYVPTTENKSLLKDTSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDID LATAVFEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVT KKSSGITTAKDMTGKTLGAQAGSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRID GLLIDRVYANYYLEAEGVLNDYNVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQE ISQKWFGEDVATKEVKEGQFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFD ASKGDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPA PQLPAPVGGEDATVQSPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEG TVVEVNSNVSKSPTGASQVMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKKWTGK LSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVPVSSL VMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLT NGAKVISNPTSSLEEGKEVKADEATN SEQ ID NO: 20, fusion protein Rhavi-SP0785-SP1500: MFDASNFKDFSSIASASSSWQNQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYP LTGRVNGTFIAFSVGWNNSTENCNSATGWTGYAQVNGNNTEIVTSWNLAYEGGSGP AIEQGQDTFQYVPTTENKSLLKDFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQY VYFDASKGDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAA SAPAPQLPAPVGGEDATVQSPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLS TLEGTVVEVNSNVSKSPTGASQVMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKK WTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVP VSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVK ADEATNTSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDIDLATAVFEKYGITVNWQP IDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTGKTL GAQAGSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEG VLNDYNVFTVGLE TEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQ SEQ ID NO: 21, fusion protein SP1500-SP0785-Rhavi: MTSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDIDLATAVFEKYGITVNWQPIDWDL KEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTGKTLGAQAG SSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDY NVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQFRQ PSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSVGDKVSEGQA LVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDATVQSPTPVAG NSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTGASQVMVH IVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAASPAAGNN TGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVE VSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVKADEATNFDASNFKDFSSIASASSSWQ NQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYPLTGRVNGTFIAFSVGWNNSTENCNSATG WTGYAQVNGNNTEIVTSWNLAYEGGSGPAIEQGQDTFQYVPTTENKSLLKD SEQ ID NO: 22, fusion protein SP0785-SP1500-Rhavi: MFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSV GDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAP VGGEDATVQSPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLE GTVVEVNSNVSKSPTGASQVMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKV YPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGFSVN IEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLT NGAKVISNPTSSLEEGKEVKADEATNTSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGF DIDLATAVFEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQV LVTKKSSGITTAKDMTGKTLGAQAGSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKND RIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGK FQEISQKWFGEDVATKEVKEGQFDASNFKDFSSIASASSSWQNQSGSTMIIQVDSFGNVSGQYV NRAQGTGCQNSPYPLTGRVNGTFIAFSVGWNNSTENCNSATGWTGYAQVNGNNTEIVTSWNLAY EGGSGPAIEQGQDTFQYVPTTENKSLLKD SEQ ID NO: 23, fusion protein CP1, Rhavi-linker (GGGGSSS)-SP1500-linker (AAA)-SP0785: MFDASNEKDFSSIASASSSWQNQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYP LTGRVNGTFIAFSVGWNNSTENCNSATGWTGYAQVNGNNTEIVTSWNLAYEGGSGP AIEQGQDTFQYVPTTENKSLLKDGGGGSSSTSGDNWSKYQSNKSITIGFDSTFVPMGF AQKDGSYAGFDIDLATAVFEKYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDE RREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTGKTLGAQAGSSGYADFEANPEILK NIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDYNVFTVGLE TEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQAAAF RQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSV GDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAP VGGEDATVQSPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLE GTVVEVNSNVSKSPTGASQVMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKV YPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGFSVN IEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLT NGAKVISNPTSSLEEGKEVKADEATN SEQ ID NO: 24, fusion protein Rhavi-GGGGSSS-SP0785-AAA-SP1500: MFDASNFKDFSSIASASSSWQNQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYP LTGRVNGTFIAFSVGWNNSTENCNSATGWTGYAQVNGNNTEIVTSWNLAYEGGSGP AIEQGQDTFQYVPTTENKSLLKDGGGGSSSFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTV TAKNEQYVYFDASKGDLDEILVSVGDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELN QARNEAASAPAPQLPAPVGGEDATVQSPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQL DATTVLSTLEGTVVEVNSNVSKSPTGASQVMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTS KVYPDKKWTGKLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGFSVNIEVKSK TKAILVPVSSLVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLINGAKVISNPTSSL EEGKEVKADEATNAAATSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDIDLATAVFE KYGITVNWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGIT TAKDMTGKTLGAQAGSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRV YANYYLEAEGVLNDYNVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFG EDVATKEVKEGQ SEQ ID NO: 25, fusion protein SP1500-GGGGSSS-SP0785-AAA-Rhavi: MTSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDIDLATAVFEKYGITVNWQPIDWDL KEAELTKGTIDLIWNGYSATDERREKVAFSNSYMKNEQVLVTKKSSGITTAKDMTGKTLGAQAG SSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLEAEGVLNDY NVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKEVKEGQGGG GSSSFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSVGD KVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAPVGGEDATVQ SPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLEGTVVEVNSNVSKSPTG ASQVMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKKWTGKLSYISDYPKNNGEAA SPAAGNNTGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVPVSSLVMDDSKNYVWIVDEQ QKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVKADEATNAAAFDASNFKDFS SIASASSSWQNQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYPLTGRVNGTFIAFSVGWNN STENCNSATGWTGYAQVNGNNTEIVTSWNLAYEGGSGPAIEQGQDTFQYVPTTENKSLLKD SEQ ID NO: 26, fusion protein SP0785-linker (GGGGSSS)-SP1500-linker (AAA)-Rhavi: MFRQPSQTALKDEPTHLVVAKEGSVASSVLLSGTVTAKNEQYVYFDASKGDLDEILVSV GDKVSEGQALVKYSSSEAQAAYDSASRAVARADRHINELNQARNEAASAPAPQLPAP VGGEDATVQSPTPVAGNSVASIDAQLGDARDARADAAAQLSKAQSQLDATTVLSTLE GTVVEVNSNVSKSPTGASQVMVHIVSNENLQVKGELSEYNLANLSVGQEVSFTSKVYPDKKWTG KLSYISDYPKNNGEAASPAAGNNTGSKYPYTIDVTGEVGDLKQGFSVNIEVKSKTKAILVPVSS LVMDDSKNYVWIVDEQQKAKKVEVSLGNADAENQEITSGLTNGAKVISNPTSSLEEGKEVKADE ATNGGGGSSSTSGDNWSKYQSNKSITIGFDSTFVPMGFAQKDGSYAGFDIDLATAVFEKYGITV NWQPIDWDLKEAELTKGTIDLIWNGYSATDERREKVAFSNSYMNEQVLVTKKSSGITTAKDMTG KTLGAQAGSSGYADFEANPEILKNIVANKEANQYQTFNEALIDLKNDRIDGLLIDRVYANYYLE AEGVLNDYNVFTVGLETEAFAVGARKEDTNLVKKINEAFSSLYKDGKFQEISQKWFGEDVATKE VKEGQAAAFDASNFKDFSSIASASSSWQNQSGSTMIIQVDSFGNVSGQYVNRAQGTGCQNSPYP LTGRVNGTFIAFSVGWNNSTENCNSATGWTGYAQVNGNNTEIVTSWNLAYEGGSGPAIEQGQDT FQYVPTTENKSLLKD SEQ ID NO: 27, codon-optimized nucleic acid sequence encoding fusion protein SP1500-SP0785 ATGACCAGCGGCGACAATTGGTCCAAATACCAGAGCAACAAGAGCATCACGATCGGCTTCGACA GCACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTACGCGGGTTTCGATATTGACCT GGCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGGCAGCCGATTGATTGGGACCTG AAAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGAATGGTTACTCCGCAACCGATG AGCGTCGCGAAAAAGTTGCCTTCAGCAACAGCTATATGAAGAATGAACAAGTGTTGGTAACCAA GAAATCTAGCGGCATTACGACCGCGAAAGACATGACCGGTAAGACGCTGGGTGCGCAGGCCGGT AGCTCTGGCTATGCGGATTTCGAGGCGAATCCTGAGATTCTGAAAAACATCGTTGCGAATAAAG AGGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCTGAAAAACGATCGCATTGACGG TCTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCCGAGGGCGTTCTGAACGATTAT ATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTGGTGCGCGCAAGGAAGATACCA ACCTGGTTAAAAAGATTAATGAGGCATTTAGCTCACTGTACAAGGACGGCAAGTTCCAAGAAAT TAGCCAGAAGTGGTTCGGTGAAGATGTTGCGACGAAAGAGGTTAAAGAGGGCCAATTTCGCCAA CCGAGCCAGACTGCGTTGAAAGATGAGCCGACCCATCTGGTTGTTGCGAAAGAGGGCAGCGTGG CATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAAACGAACAATACGTGTACTTCGATGC TAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGACAAAGTTAGCGAAGGCCAGGCA CTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACAGCGCAAGCCGCGCAGTGGCGC GTGCCGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGAGGCCGCAAGCGCGCCAGCACC GCAGCTGCCGGCTCCGGTGGGTGGCGAAGATGCGACGGTGCAGAGCCCGACCCCGGTTGCGGGT AATTCGGTCGCCAGCATCGATGCGCAGCTGGGTGACGCGCGTGATGCCCGTGCGGATGCGGCTG CTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGACGGTGCTGAGCACCTTGGAGGGTAC CGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGTGCGAGCCAGGTTATGGTCCAC ATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCGAGTATAACCTGGCGAATCTGA GCGTTGGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGATAAGAAATGGACCGGCAAGTT GAGCTACATCAGCGACTACCCGAAGAACAATGGCGAGGCAGCCTCCCCGGCAGCCGGCAACAAT ACCGGCTCTAAGTATCCGTACACCATCGACGTAACCGGTGAGGTCGGCGACCTGAAACAGGGTT TTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATTTTGGTTCCGGTTAGCTCCCTGGT GATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAACAGAAAGCGAAAAAAGTTGAA GTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGTCTGGTCTGACCAACGGTGCGA AAGTTATTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGAGGTCAAAGCCGACGAAGCTAC GAC SEQ ID NO: 28, codon-optimized nucleic acid sequence encoding fusion protein SP0785-SP1500 ATGTTTCGCCAACCGAGCCAGACTGCGTTGAAAGATGAGCCGACCCATCTGGTTGTTGCGAAAG AGGGCAGCGTGGCATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAAACGAACAATACGT GTACTTCGATGCTAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGACAAAGTTAGC GAAGGCCAGGCACTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACAGCGCAAGCC GCGCAGTGGCGCGTGCCGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGAGGCCGCAAG CGCGCCAGCACCGCAGCTGCCGGCTCCGGTGGGTGGCGAAGATGCGACGGTGCAGAGCCCGACC CCGGTTGCGGGTAATTCGGTCGCCAGCATCGATGCGCAGCTGGGTGACGCGCGTGATGCCCGTG CGGATGCGGCTGCTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGACGGTGCTGAGCAC CTTGGAGGGTACCGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGTGCGAGCCAG GTTATGGTCCACATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCGAGTATAACC TGGCGAATCTGAGCGTTGGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGATAAGAAATG GACCGGCAAGTTGAGCTACATCAGCGACTACCCGAAGAACAATGGCGAGGCAGCCTCCCCGGCA GCCGGCAACAATACCGGCTCTAAGTATCCGTACACCATCGACGTAACCGGTGAGGTCGGCGACC TGAAACAGGGTTTTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATTTTGGTTCCGGT TAGCTCCCTGGTGATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAACAGAAAGCG AAAAAAGTTGAAGTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGTCTGGTCTGA CCAACGGTGCGAAAGTTATTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGAGGTCAAAGC CGACGAAGCTACGAACACCAGCGGCGACAATTGGTCCAAATACCAGAGCAACAAGAGCATCACG ATCGGCTTCGACAGCACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTACGCGGGTT TCGATATTGACCTGGCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGGCAGCCGAT TGATTGGGACCTGAAAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGAATGGTTAC TCCGCAACCGATGAGCGTCGCGAAAAAGTTGCCTTCAGCAACAGCTATATGAAGAATGAACAAG TGTTGGTAACCAAGAAATCTAGCGGCATTACGACCGCGAAAGACATGACCGGTAAGACGCTGGG TGCGCAGGCCGGTAGCTCTGGCTATGCGGATTTCGAGGCGAATCCTGAGATTCTGAAAAACATC GTTGCGAATAAAGAGGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCTGAAAAACG ATCGCATTGACGGTCTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCCGAGGGCGT TCTGAACGATTATAATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTGGTGCGCGC AAGGAAGATACCAACCTGGTTAAAAAGATTAATGAGGCATTTAGCTCACTGTACAAGGACGGCA AGTTCCAAGAAATTAGCCAGAAGTGGTTCGGTGAAGATGTTGCGACGAAAGAGGTTAAAGAGGG CCAA SEQ ID NO: 29, codon-optimized nucleic acid sequence encoding fusion protein Rhavi-SP1500-SP0785: ATGTTCGACGCATCCAACTTTAAAGACTTTAGCAGCATCGCGTCCGCAAGCTCTAGCTGGCAGA ATCAATCTGGTAGCACCATGATTATCCAAGTGGACAGCTTTGGTAACGTCAGCGGTCAATATGT TAATCGTGCACAGGGTACGGGTTGTCAGAATTCTCCGTACCCGCTGACCGGTCGTGTTAACGGC ACGTTCATCGCTTTCAGCGTCGGTTGGAACAATTCTACTGAAAATTGCAACAGCGCGACCGGTT GGACGGGCTATGCACAAGTGAATGGCAATAACACCGAAATCGTCACGTCCTGGAATCTGGCGTA TGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGCCAGGATACCTTCCAATACGTCCCTACGACC GAGAATAAGTCCCTTCTGAAAGACACCAGCGGCGACAATTGGTCCAAATACCAGAGCAACAAGA GCATCACGATCGGCTTCGACAGCACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTA CGCGGGTTTCGATATTGACCTGGCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGG CAGCCGATTGATTGGGACCTGAAAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGA ATGGTTACTCCGCAACCGATGAGCGTCGCGAAAAAGTTGCCTTCAGCAACAGCTATATGAAGAA
TGAACAAGTGTTGGTAACCAAGAAATCTAGCGGCATTACGACCGCGAAAGACATGACCGGTAAG ACGCTGGGTGCGCAGGCCGGTAGCTCTGGCTATGCGGATTTCGAGGCGAATCCTGAGATTCTGA AAAACATCGTTGCGAATAAAGAGGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCT GAAAAACGATCGCATTGACGGTCTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCC GAGGGCGTTCTGAACGATTATAATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTG GTGCGCGCAAGGAAGATACCAACCTGGTTAAAAAGATTAATGAGGCATTTAGCTCACTGTACAA GGACGGCAAGTTCCAAGAAATTAGCCAGAAGTGGTTCGGTGAAGATGTTGCGACGAAAGAGGTT AAAGAGGGCCAATTTCGCCAACCGAGCCAGACTGCGTTGAAAGATGAGCCGACCCATCTGGTTG TTGCGAAAGAGGGCAGCGTGGCATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAAACGA ACAATACGTGTACTTCGATGCTAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGAC AAAGTTAGCGAAGGCCAGGCACTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACA GCGCAAGCCGCGCAGTGGCGCGTGCCGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGA GGCCGCAAGCGCGCCAGCACCGCAGCTGCCGGCTCCGGTGGGTGGCGAAGATGCGACGGTGCAG AGCCCGACCCCGGTTGCGGGTAATTCGGTCGCCAGCATCGATGCGCAGCTGGGTGACGCGCGTG ATGCCCGTGCGGATGCGGCTGCTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGACGGT GCTGAGCACCTTGGAGGGTACCGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGT GCGAGCCAGGTTATGGTCCACATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCG AGTATAACCTGGCGAATCTGAGCGTTGGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGA TAAGAAATGGACCGGCAAGTTGAGCTACATCAGCGACTACCCGAAGAACAATGGCGAGGCAGCC TCCCCGGCAGCCGGCAACAATACCGGCTCTAAGTATCCGTACACCATCGACGTAACCGGTGAGG TCGGCGACCTGAAACAGGGTTTTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATTTT GGTTCCGGTTAGCTCCCTGGTGATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAA CAGAAAGCGAAAAAAGTTGAAGTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGT CTGGTCTGACCAACGGTGCGAAAGTTATTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGA GGTCAAAGCCGACGAAGCTACGAAC SEQ ID NO: 30, codon-optimized nucleic acid sequence encoding fusion protein Rhavi-SP0785-SP1500: ATGTTCGACGCATCCAACTTTAAAGACTTTAGCAGCATCGCGTCCGCAAGCTCTAGCTGGCAGA ATCAATCTGGTAGCACCATGATTATCCAAGTGGACAGCTTTGGTAACGTCAGCGGTCAATATGT TAATCGTGCACAGGGTACGGGTTGTCAGAATTCTCCGTACCCGCTGACCGGTCGTGTTAACGGC ACGTTCATCGCTTTCAGCGTCGGTTGGAACAATTCTACTGAAAATTGCAACAGCGCGACCGGTT GGACGGGCTATGCACAAGTGAATGGCAATAACACCGAAATCGTCACGTCCTGGAATCTGGCGTA TGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGCCAGGATACCTTCCAATACGTCCCTACGACC GAGAATAAGTCCCTTCTGAAAGACTTTCGCCAACCGAGCCAGACTGCGTTGAAAGATGAGCCGA CCCATCTGGTTGTTGCGAAAGAGGGCAGCGTGGCATCGAGCGTGCTGCTGAGCGGTACGGTTAC TGCCAAAAACGAACAATACGTGTACTTCGATGCTAGCAAGGGTGATCTGGATGAAATTCTGGTG AGCGTGGGTGACAAAGTTAGCGAAGGCCAGGCACTGGTGAAGTATTCATCCTCCGAGGCACAGG CAGCGTACGACAGCGCAAGCCGCGCAGTGGCGCGTGCCGACCGTCACATTAACGAATTGAACCA AGCGCGTAACGAGGCCGCAAGCGCGCCAGCACCGCAGCTGCCGGCTCCGGTGGGTGGCGAAGAT GCGACGGTGCAGAGCCCGACCCCGGTTGCGGGTAATTCGGTCGCCAGCATCGATGCGCAGCTGG GTGACGCGCGTGATGCCCGTGCGGATGCGGCTGCTCAACTGAGCAAGGCTCAGAGCCAACTGGA CGCGACGACGGTGCTGAGCACCTTGGAGGGTACCGTTGTCGAAGTCAACAGCAATGTGAGCAAG AGCCCAACGGGTGCGAGCCAGGTTATGGTCCACATTGTGAGCAATGAAAACTTACAGGTCAAGG GTGAGCTGAGCGAGTATAACCTGGCGAATCTGAGCGTTGGTCAAGAGGTCAGCTTTACCAGCAA GGTCTACCCGGATAAGAAATGGACCGGCAAGTTGAGCTACATCAGCGACTACCCGAAGAACAAT GGCGAGGCAGCCTCCCCGGCAGCCGGCAACAATACCGGCTCTAAGTATCCGTACACCATCGACG TAACCGGTGAGGTCGGCGACCTGAAACAGGGTTTTAGCGTGAATATCGAAGTGAAGTCCAAGAC CAAGGCAATTTTGGTTCCGGTTAGCTCCCTGGTGATGGACGATAGCAAGAATTATGTGTGGATT GTCGACGAGCAACAGAAAGCGAAAAAAGTTGAAGTGAGCCTGGGCAATGCTGATGCCGAGAACC AAGAAATCACGTCTGGTCTGACCAACGGTGCGAAAGTTATTAGCAACCCGACCAGCAGCCTGGA AGAGGGTAAAGAGGTCAAAGCCGACGAAGCTACGAACACCAGCGGCGACAATTGGTCCAAATAC CAGAGCAACAAGAGCATCACGATCGGCTTCGACAGCACTTTTGTGCCGATGGGTTTCGCGCAAA AAGACGGTAGCTACGCGGGTTTCGATATTGACCTGGCGACCGCTGTCTTTGAGAAATACGGCAT TACGGTTATTGGCAGCCGATTGATTGGGACCTGAAAGAGGCCGAACTCACCAAAGGCACCATC GACCTGATCTGGAATGGTTACTCCGCAACCGATGAGCGTCGCGAAAAAGTTGCCTTCAGCAACA GCTATATGAAGAATGAACAAGTGTTGGTAACCAAGAAATCTAGCGGCATTACGACCGCGAAAGA CATGACCGGTAAGACGCTGGGTGCGCAGGCCGGTAGCTCTGGCTATGCGGATTTCGAGGCGAAT CCTGAGATTCTGAAAAACATCGTTGCGAATAAAGAGGCGAACCAGTACCAGACCTTTAACGAAG CACTGATCGACCTGAAAAACGATCGCATTGACGGTCTGCTGATCGATCGTGTGTACGCGAACTA TTATCTGGAAGCCGAGGGCGTTCTGAACGATTATAATGTTTTTACCGTGGGTCTGGAGACTGAG GCATTCGCGGTTGGTGCGCGCAAGGAAGATACCAACCTGGTTAAAAAGATTAATGAGGCATTTA GCTCACTGTACAAGGACGGCAAGTTCCAAGAAATTAGCCAGAAGTGGTTCGGTGAAGATGTTGC GACGAAAGAGGTTAAAGAGGGCCAA SEQ ID NO: 31, codon-optimized nucleic acid sequence encoding fusion protein SP1500-SP0785-Rhavi: ATGACCAGCGGCGACAATTGGTCCAAATACCAGAGCAACAAGAGCATCACGATCGGCTTCGACA GCACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTACGCGGGTTTCGATATTGACCT GGCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGGCAGCCGATTGATTGGGACCTG AAAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGAATGGTTACTCCGCAACCGATG AGCGTCGCGAAAAAGTTGCCTTCAGCAACAGCTATATGAAGAATGAACAAGTGTTGGTAACCAA GAAATCTAGCGGCATTACGACCGCGAAAGACATGACCGGTAAGACGCTGGGTGCGCAGGCCGGT AGCTCTGGCTATGCGGATTTCGAGGCGAATCCTGAGATTCTGAAAAACATCGTTGCGAATAAAG AGGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCTGAAAAACGATCGCATTGACGG TCTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCCGAGGGCGTTCTGAACGATTAT AATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTGGTGCGCGCAAGGAAGATACCA ACCTGGTTAAAAAGATTAATGAGGCATTTAGCTCACTGTACAAGGACGGCAAGTTCCAAGAAAT TAGCCAGAAGTGGTTCGGTGAAGATGTTGCGACGAAAGAGGTTAAAGAGGGCCAATTTCGCCAA CCGAGCCAGACTGCGTTGAAAGATGAGCCGACCCATCTGGTTGTTGCGAAAGAGGGCAGCGTGG CATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAAACGAACAATACGTGTACTTCGATGC TAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGACAAAGTTAGCGAAGGCCAGGCA CTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACAGCGCAAGCCGCGCAGTGGCGC GTGCCGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGAGGCCGCAAGCGCGCCAGCACC GCAGCTGCCGGCTCCGGTGGGTGGCGAAGATGCGACGGTGCAGAGCCCGACCCCGGTTGCGGGT AATTCGGTCGCCAGCATCGATGCGCAGCTGGGTGACGCGCGTGATGCCCGTGCGGATGCGGCTG CTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGACGGTGCTGAGCACCTTGGAGGGTAC CGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGTGCGAGCCAGGTTATGGTCCAC ATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCGAGTATAACCTGGCGAATCTGA GCGTTGGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGATAAGAAATGGACCGGCAAGTT GAGCTACATCAGCGACTACCCGAAGAACAATGGCGAGGCAGCCTCCCCGGCAGCCGGCAACAAT ACCGGCTCTAAGTATCCGTACACCATCGACGTAACCGGTGAGGTCGGCGACCTGAAACAGGGTT TTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATTTTGGTTCCGGTTAGCTCCCTGGT GATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAACAGAAAGCGAAAAAAGTTGAA GTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGTCTGGTCTGACCAACGGTGCGA AAGTTATTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGAGGTCAAAGCCGACGAAGCTAC GAACTTCGACGCATCCAACTTTAAAGACTTTAGCAGCATCGCGTCCGCAAGCTCTAGCTGGCAG AATCAATCTGGTAGCACCATGATTATCCAAGTGGACAGCTTTGGTAACGTCAGCGGTCAATATG TTAATCGTGCACAGGGTACGGGTTGTCAGAATTCTCCGTACCCGCTGACCGGTCGTGTTAACGG CACGTTCATCGCTTTCAGCGTCGGTTGGAACAATTCTACTGAAAATTGCAACAGCGCGACCGGT TGGACGGGCTATGCACAAGTGAATGGCAATAACACCGAAATCGTCACGTCCTGGAATCTGGCGT ATGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGCCAGGATACCTTCCAATACGTCCCTACGAC CGAGAATAAGTCCCTTCTGAAAGAC SEQ ID NO: 32, codon-optimized nucleic acid sequence encoding fusion protein SP0785-SP1500-Rhavi: ATGTTTCGCCAACCGAGCCAGACTGCGTTGAAAGATGAGCCGACCCATCTGGTTGTTGCGAAAG AGGGCAGCGTGGCATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAAACGAACAATACGT GTACTTCGATGCTAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGACAAAGTTAGC GAAGGCCAGGCACTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACAGCGCAAGCC GCGCAGTGGCGCGTGCCGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGAGGCCGCAAG CGCGCCAGCACCGCAGCTGCCGGCTCCGGTGGGTGGCGAAGATGCGACGGTGCAGAGCCCGACC CCGGTTGCGGGTAATTCGGTCGCCAGCATCGATGCGCAGCTGGGTGACGCGCGTGATGCCCGTG CGGATGCGGCTGCTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGACGGTGCTGAGCAC CTTGGAGGGTACCGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGTGCGAGCCAG GTTATGGTCCACATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCGAGTATAACC TGGCGAATCTGAGCGTTGGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGATAAGAAATG GACCGGCAAGTTGAGCTACATCAGCGACTACCCGAAGAACAATGGCGAGGCAGCCTCCCCGGCA GCCGGCAACAATACCGGCTCTAAGTATCCGTACACCATCGACGTAACCGGTGAGGTCGGCGACC TGAAACAGGGTTTTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATTTTGGTTCCGGT TAGCTCCCTGGTGATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAACAGAAAGCG AAAAAAGTTGAAGTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGTCTGGTCTGA CCAACGGTGCGAAAGTTATTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGAGGTCAAAGC CGACGAAGCTACGAACACCAGCGGCGACAATTGGTCCAAATACCAGAGCAACAAGAGCATCACG ATCGGCTTCGACAGCACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTACGCGGGTT TCGATATTGACCTGGCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGGCAGCCGAT TGATTGGGACCTGAAAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGAATGGTTAC TCCGCAACCGATGAGCGTCGCGAAAAAGTTGCCTTCAGCAACAGCTATATGAAGAATGAACAAG TGTTGGTAACCAAGAAATCTAGCGGCATTACGACCGCGAAAGACATGACCGGTAAGACGCTGGG TGCGCAGGCCGGTAGCTCTGGCTATGCGGATTTCGAGGCGAATCCTGAGATTCTGAAAAACATC GTTGCGAATAAAGAGGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCTGAAAAACG ATCGCATTGACGGTCTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCCGAGGGCGT TCTGAACGATTATAATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTGGTGCGCGC AAGGAAGATACCAACCTGGTTAAAAAGATTAATGAGGCATTTAGCTCACTGTACAAGGACGGCA AGTTCCAAGAAATTAGCCAGAAGTGGTTCGGTGAAGATGTTGCGACGAAAGAGGTTAAAGAGGG CCAATTCGACGCATCCAACTTTAAAGACTTTAGCAGCATCGCGTCCGCAAGCTCTAGCTGGCAG AATCAATCTGGTAGCACCATGATTATCCAAGTGGACAGCTTTGGTAACGTCAGCGGTCAATATG TTAATCGTGCACAGGGTACGGGTTGTCAGAATTCTCCGTACCCGCTGACCGGTCGTGTTAACGG CACGTTCATCGCTTTCAGCGTCGGTTGGAACAATTCTACTGAAAATTGCAACAGCGCGACCGGT TGGACGGGCTATGCACAAGTGAATGGCAATAACACCGAAATCGTCACGTCCTGGAATCTGGCGT ATGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGCCAGGATACCTTCCAATACGTCCCTACGAC CGAGAATAAGTCCCTTCTGAAAGAC SEQ ID NO: 33, codon-optimized nucleic acid sequence encoding fusion protein CP1 Rhavi-linker (GGGGSSS)-SP1500-linker (AAA)-SP0785: ATGTTCGACGCATCCAACTTTAAAGACTTTAGCAGCATCGCGTCCGCAAGCTCTAGCTGGCAGAATCAATCTGG- TAG CACCATGATTATCCAAGTGGACAGCTTTGGTAACGTCAGCGGTCAATATGTTAATCGTGCACAGGGTACGGGTT- GTC AGAATTCTCCGTACCCGCTGACCGGTCGTGTTAACGGCACGTTCATCGCTTTCAGCGTCGGTTGGAACAATTCT- ACT GAAAATTGCAACAGCGCGACCGGTTGGACGGGCTATGCACAAGTGAATGGCAATAACACCGAAATCGTCACGTC- CTG GAATCTGGCGTATGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGCCAGGATACCTTCCAATACGTCCCTACGA- CCG AGAATAAGTCCCTTCTGAAAGACGGCGGTGGCGGTTCGAGCTCGACCAGCGGCGACAATTGGTCCAAATACCAG- AGC AACAAGAGCATCACGATCGGCTTCGACAGCACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTACGC- GGG TTTCGATATTGACCTGGCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGGCAGCCGATTGATTGGG- ACC TGAAAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGAATGGTTACTCCGCAACCGATGAGCGTCGC- GAA AAAGTTGCCTTCAGCAACAGCTATATGAAGAATGAACAAGTGTTGGTAACCAAGAAATCTAGCGGCATTACGAC- CGC GAAAGACATGACCGGTAAGACGCTGGGTGCGCAGGCCGGTAGCTCTGGCTATGCGGATTTCGAGGCGAATCCTG- AGA TTCTGAAAAACATCGTTGCGAATAAAGAGGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCTGAAA- AAC GATCGCATTGACGGTCTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCCGAGGGCGTTCTGAACGA- TTA TAATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTGGTGCGCGCAAGGAAGATACCAACCTGGTTA- AAA AGATTAATGAGGCATTTAGCTCACTGTACAAGGACGGCAAGTTCCAAGAAATTAGCCAGAAGTGGTTCGGTGAA- GAT GTTGCGACGAAAGAGGTTAAAGAGGGCCAAGCGGCCGCATTTCGCCAACCGAGCCAGACTGCGTTGAAAGATGA- GCC GACCCATCTGGTTGTTGCGAAAGAGGGCAGCGTGGCATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAA- ACG AACAATACGTGTACTTCGATGCTAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGACAAAGTTAGC- GAA GGCCAGGCACTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACAGCGCAAGCCGCGCAGTGGCGCG- TGC CGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGAGGCCGCAAGCGCGCCAGCACCGCAGCTGCCGGCTC- CGG TGGGTGGCGAAGATGCGACGGTGCAGAGCCCGACCCCGGTTGCGGGTAATTCGGTCGCCAGCATCGATGCGCAG- CTG GGTGACGCGCGTGATGCCCGTGCGGATGCGGCTGCTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGAC- GGT GCTGAGCACCTTGGAGGGTACCGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGTGCGAGCCAGG- TTA TGGTCCACATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCGAGTATAACCTGGCGAATCTGAGC- GTT GGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGATAAGAAATGGACCGGCAAGTTGAGCTACATCAGCGA- CTA CCCGAAGAACAATGGCGAGGCAGCCTCCCCGGCAGCCGGCAACAATACCGGCTCTAAGTATCCGTACACCATCG- ACG TAACCGGTGAGGTCGGCGACCTGAAACAGGGTTTTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATT- TTG GTTCCGGTTAGCTCCCTGGTGATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAACAGAAAGCGAA- AAA AGTTGAAGTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGTCTGGTCTGACCAACGGTGCGAAAG- TTA TTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGAGGTCAAAGCCGACGAAGCTACGAAC SEQ ID NO: 34, codon-optimized nucleic acid sequence encoding fusion protein Rhavi-linker (GGGGSSS)-SP0785-linker (AAA)-SP1500: ATGTTCGACGCATCCAACTTTAAAGACTTTAGCAGCATCGCGTCCGCAAGCTCTAGCTGGCAGA ATCAATCTGGTAGCACCATGATTATCCAAGTGGACAGCTTTGGTAACGTCAGCGGTCAATATGT TAATCGTGCACAGGGTACGGGTTGTCAGAATTCTCCGTACCCGCTGACCGGTCGTGTTAACGGC ACGTTCATCGCTTTCAGCGTCGGTTGGAACAATTCTACTGAAAATTGCAACAGCGCGACCGGTT GGACGGGCTATGCACAAGTGAATGGCAATAACACCGAAATCGTCACGTCCTGGAATCTGGCGTA TGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGCCAGGATACCTTCCAATACGTCCCTACGACC GAGAATAAGTCCCTTCTGAAAGACGGCGGTGGCGGTTCGAGCTCGTTTCGCCAACCGAGCCAGA CTGCGTTGAAAGATGAGCCGACCCATCTGGTTGTTGCGAAAGAGGGCAGCGTGGCATCGAGCGT GCTGCTGAGCGGTACGGTTACTGCCAAAAACGAACAATACGTGTACTTCGATGCTAGCAAGGGT GATCTGGATGAAATTCTGGTGAGCGTGGGTGACAAAGTTAGCGAAGGCCAGGCACTGGTGAAGT ATTCATCCTCCGAGGCACAGGCAGCGTACGACAGCGCAAGCCGCGCAGTGGCGCGTGCCGACCG TCACATTAACGAATTGAACCAAGCGCGTAACGAGGCCGCAAGCGCGCCAGCACCGCAGCTGCCG GCTCCGGTGGGTGGCGAAGATGCGACGGTGCAGAGCCCGACCCCGGTTGCGGGTAATTCGGTCG CCAGCATCGATGCGCAGCTGGGTGACGCGCGTGATGCCCGTGCGGATGCGGCTGCTCAACTGAG CAAGGCTCAGAGCCAACTGGACGCGACGACGGTGCTGAGCACCTTGGAGGGTACCGTTGTCGAA GTCAACAGCAATGTGAGCAAGAGCCCAACGGGTGCGAGCCAGGTTATGGTCCACATTGTGAGCA ATGAAAACTTACAGGTCAAGGGTGAGCTGAGCGAGTATAACCTGGCGAATCTGAGCGTTGGTCA AGAGGTCAGCTTTACCAGCAAGGTCTACCCGGATAAGAAATGGACCGGCAAGTTGAGCTACATC AGCGACTACCCGAAGAACAATGGCGAGGCAGCCTCCCCGGCAGCCGGCAACAATACCGGCTCTA AGTATCCGTACACCATCGACGTAACCGGTGAGGTCGGCGACCTGAAACAGGGTTTTAGCGTGAA TATCGAAGTGAAGTCCAAGACCAAGGCAATTTTGGTTCCGGTTAGCTCCCTGGTGATGGACGAT AGCAAGAATTATGTGTGGATTGTCGACGAGCAACAGAAAGCGAAAAAAGTTGAAGTGAGCCTGG GCAATGCTGATGCCGAGAACCAAGAAATCACGTCTGGTCTGACCAACGGTGCGAAAGTTATTAG CAACCCGACCAGCAGCCTGGAAGAGGGTAAAGAGGTCAAAGCCGACGAAGCTACGAACCGGCCG CAACCAGCGGCGACAATTGGTCCAAATACCAGAGCAACAAGAGCATCACGATCGGCTTCGACAG CACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTACGCGGGTTTCGATATTGACCTG GCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGGCAGCCGATTGATTGGGACCTGA AAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGAATGGTTACTCCGCAACCGATGA GCGTCGCGAAAAAGTTGCCTTCAGCAACAGCTATATGAAGAATGAACAAGTGTTGGTAACCAAG AAATCTAGCGGCATTACGACCGCGAAAGACATGACCGGTAAGACGCTGGGTGCGCAGGCCGGTA GCTCTGGCTATGCGGATTTCGAGGCGAATCCTGAGATTCTGAAAAACATCGTTGCGAATAAAGA GGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCTGAAAAACGATCGCATTGACGGT CTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCCGAGGGCGTTCTGAACGATTATA ATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTGGTGCGCGCAAGGAAGATACCAA CCTGGTTAAAAAGATTAATGAGGCATTTAGCTCACTGTACAAGGACGGCAAGTTCCAAGAAATT AGCCAGAAGTGGTTCGGTGAAGATGTTGCGACGAAAGAGGTTAAAGAGGGCCAA SEQ ID NO: 35, codon-optimized nucleic acid sequence encoding fusion protein
SP1500-linker (GGGGSSS)-SP0785-linker (AAA)-Rhavi: ATGACCAGCGGCGACAATTGGTCCAAATACCAGAGCAACAAGAGCATCACGATCGGCTTCGACA GCACTTTTGTGCCGATGGGTTTCGCGCAAAAAGACGGTAGCTACGCGGGTTTCGATATTGACCT GGCGACCGCTGTCTTTGAGAAATACGGCATTACGGTTAATTGGCAGCCGATTGATTGGGACCTG AAAGAGGCCGAACTCACCAAAGGCACCATCGACCTGATCTGGAATGGTTACTCCGCAACCGATG AGCGTCGCGAAAAAGTTGCCTTCAGCAACAGCTATATGAAGAATGAACAAGTGTTGGTAACCAA GAAATCTAGCGGCATTACGACCGCGAAAGACATGACCGGTAAGACGCTGGGTGCGCAGGCCGGT AGCTCTGGCTATGCGGATTTCGAGGCGAATCCTGAGATTCTGAAAAACATCGTTGCGAATAAAG AGGCGAACCAGTACCAGACCTTTAACGAAGCACTGATCGACCTGAAAAACGATCGCATTGACGG TCTGCTGATCGATCGTGTGTACGCGAACTATTATCTGGAAGCCGAGGGCGTTCTGAACGATTAT AATGTTTTTACCGTGGGTCTGGAGACTGAGGCATTCGCGGTTGGTGCGCGCAAGGAAGATACCA ACCTGGTTAAAAAGATTAATGAGGCATTTAGCTCACTGTACAAGGACGGCAAGTTCCAAGAAAT TAGCCAGAAGTGGTTCGGTGAAGATGTTGCGACGAAAGAGGTTAAAGAGGGCCAAGGCGGTGGC GGTTCGAGCTCGTTTCGCCAACCGAGCCAGACTGCGTTGAAAGATGAGCCGACCCATCTGGTTG TTGCGAAAGAGGGCAGCGTGGCATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAAACGA ACAATACGTGTACTTCGATGCTAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGAC AAAGTTAGCGAAGGCCAGGCACTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACA GCGCAAGCCGCGCAGTGGCGCGTGCCGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGA GGCCGCAAGCGCGCCAGCACCGCAGCTGCCGGCTCCGGTGGGTGGCGAAGATGCGACGGTGCAG AGCCCGACCCCGGTTGCGGGTAATTCGGTCGCCAGCATCGATGCGCAGCTGGGTGACGCGCGTG ATGCCCGTGCGGATGCGGCTGCTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGACGGT GCTGAGCACCTTGGAGGGTACCGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGT GCGAGCCAGGTTATGGTCCACATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCG AGTATAACCTGGCGAATCTGAGCGTTGGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGA TAAGAAATGGACCGGCAAGTTGAGCTACATCAGCGACTACCCGAAGAACAATGGCGAGGCAGCC TCCCCGGCAGCCGGCAACAATACCGGCTCTAAGTATCCGTACACCATCGACGTAACCGGTGAGG TCGGCGACCTGAAACAGGGTTTTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATTTT GGTTCCGGTTAGCTCCCTGGTGATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAA CAGAAAGCGAAAAAAGTTGAAGTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGT CTGGTCTGACCAACGGTGCGAAAGTTATTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGA GGTCAAAGCCGACGAAGCTACGAACCGGCCGCATTCGACGCATCCAACTTTAAAGACTTTAGCA GCATCGCGTCCGCAAGCTCTAGCTGGCAGAATCAATCTGGTAGCACCATGATTATCCAAGTGGA CAGCTTTGGTAACGTCAGCGGTCAATATGTTAATCGTGCACAGGGTACGGGTTGTCAGAATTCT CCGTACCCGCTGACCGGTCGTGTTAACGGCACGTTCATCGCTTTCAGCGTCGGTTGGAACAATT CTACTGAAAATTGCAACAGCGCGACCGGTTGGACGGGCTATGCACAAGTGAATGGCAATAACAC CGAAATCGTCACGTCCTGGAATCTGGCGTATGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGC CAGGATACCTTCCAATACGTCCCTACGACCGAGAATAAGTCCCTTCTGAAAGAC SEQ ID NO: 36, codon-optimized nucleic acid sequence encoding fusion protein SP0785-linker (GGGGSSS)-SP1500-linker (AAA)-Rhavi: ATGTTTCGCCAACCGAGCCAGACTGCGTTGAAAGATGAGCCGACCCATCTGGTTGTTGCGAAAG AGGGCAGCGTGGCATCGAGCGTGCTGCTGAGCGGTACGGTTACTGCCAAAAACGAACAATACGT GTACTTCGATGCTAGCAAGGGTGATCTGGATGAAATTCTGGTGAGCGTGGGTGACAAAGTTAGC GAAGGCCAGGCACTGGTGAAGTATTCATCCTCCGAGGCACAGGCAGCGTACGACAGCGCAAGCC GCGCAGTGGCGCGTGCCGACCGTCACATTAACGAATTGAACCAAGCGCGTAACGAGGCCGCAAG CGCGCCAGCACCGCAGCTGCCGGCTCCGGTGGGTGGCGAAGATGCGACGGTGCAGAGCCCGACC CCGGTTGCGGGTAATTCGGTCGCCAGCATCGATGCGCAGCTGGGTGACGCGCGTGATGCCCGTG CGGATGCGGCTGCTCAACTGAGCAAGGCTCAGAGCCAACTGGACGCGACGACGGTGCTGAGCAC CTTGGAGGGTACCGTTGTCGAAGTCAACAGCAATGTGAGCAAGAGCCCAACGGGTGCGAGCCAG GTTATGGTCCACATTGTGAGCAATGAAAACTTACAGGTCAAGGGTGAGCTGAGCGAGTATAACC TGGCGAATCTGAGCGTTGGTCAAGAGGTCAGCTTTACCAGCAAGGTCTACCCGGATAAGAAATG GACCGGCAAGTTGAGCTACATCAGCGACTACCCGAAGAACAATGGCGAGGCAGCCTCCCCGGCA GCCGGCAACAATACCGGCTCTAAGTATCCGTACACCATCGACGTAACCGGTGAGGTCGGCGACC TGAAACAGGGTTTTAGCGTGAATATCGAAGTGAAGTCCAAGACCAAGGCAATTTTGGTTCCGGT TAGCTCCCTGGTGATGGACGATAGCAAGAATTATGTGTGGATTGTCGACGAGCAACAGAAAGCG AAAAAAGTTGAAGTGAGCCTGGGCAATGCTGATGCCGAGAACCAAGAAATCACGTCTGGTCTGA CCAACGGTGCGAAAGTTATTAGCAACCCGACCAGCAGCCTGGAAGAGGGTAAAGAGGTCAAAGC CGACGAAGCTACGAACGGCGGTGGCGGTTCGAGCTCGACCAGCGGCGACAATTGGTCCAAATAC CAGAGCAACAAGAGCATCACGATCGGCTTCGACAGCACTTTTGTGCCGATGGGTTTCGCGCAAA AAGACGGTAGCTACGCGGGTTTCGATATTGACCTGGCGACCGCTGTCTTTGAGAAATACGGCAT TACGGTTAATTGGCAGCCGATTGATTGGGACCTGAAAGAGGCCGAACTCACCAAAGGCACCATC GACCTGATCTGGAATGGTTACTCCGCAACCGATGAGCGTCGCGAAAAAGTTGCCTTCAGCAACA GCTATATGAAGAATGAACAAGTGTTGGTAACCAAGAAATCTAGCGGCATTACGACCGCGAAAGA CATGACCGGTAAGACGCTGGGTGCGCAGGCCGGTAGCTCTGGCTATGCGGATTTCGAGGCGAAT CCTGAGATTCTGAAAAACATCGTTGCGAATAAAGAGGCGAACCAGTACCAGACCTTTAACGAAG CACTGATCGACCTGAAAAACGATCGCATTGACGGTCTGCTGATCGATCGTGTGTACGCGAACTA TTATCTGGAAGCCGAGGGCGTTCTGAACGATTATAATGTTTTTACCGTGGGTCTGGAGACTGAG GCATTCGCGGTTGGTGCGCGCAAGGAAGATACCAACCTGGTTAAAAAGATTAATGAGGCATTTA GCTCACTGTACAAGGACGGCAAGTTCCAAGAAATTAGCCAGAAGTGGTTCGGTGAAGATGTTGC GACGAAAGAGGTTAAAGAGGGCCAACGGCCGCATTCGACGCATCCAACTTTAAAGACTTTAGCA GCATCGCGTCCGCAAGCTCTAGCTGGCAGAATCAATCTGGTAGCACCATGATTATCCAAGTGGA CAGCTTTGGTAACGTCAGCGGTCAATATGTTAATCGTGCACAGGGTACGGGTTGTCAGAATTCT CCGTACCCGCTGACCGGTCGTGTTAACGGCACGTTCATCGCTTTCAGCGTCGGTTGGAACAATT CTACTGAAAATTGCAACAGCGCGACCGGTTGGACGGGCTATGCACAAGTGAATGGCAATAACAC CGAAATCGTCACGTCCTGGAATCTGGCGTATGAGGGTGGCAGCGGTCCGGCTATTGAACAGGGC CAGGATACCTTCCAATACGTCCCTACGACCGAGAATAAGTCCCTTCTGAAAGAC SEQ ID NO: 37, linker sequence [7 amino acids]: GGGGSSS SEQ ID NO: 38, linker sequence [3 amino acids]: AAA SEQ ID NO: 39, linker sequence [5 amino acid repeats]: (GGGGS).sub.n SEQ ID NO: 40, linker sequence [6 amino acids]: GGGGGG SEQ ID NO: 41, linker sequence [15 amino acids]: GGGGSGGGGSGGGGS SEQ ID NO: 42, linker sequence [30 amino acids]: GGGGSGGGGSGGGGSGGGGSGGGGSGGGGS SEQ ID NO: 43, linker sequence [18 amino acids]: KESGSVSSEQLAQFRSLD SEQ ID NO: 44, linker sequence [14 amino acids]: EGKSSGSGSESKST SEQ ID NO: 45, linker sequence: (Gly).sub.n SEQ ID NO: 46, linker sequence [8 amino acids]: GGGGGGGG SEQ ID NO: 47, linker sequence [12 amino acids]: GSAGSAAGSGEF SEQ ID NO: 48, linker sequence [5 amino acid repeats]: (EAAAK).sub.n SEQ ID NO: 49, linker sequence: A(EAAAK).sub.nA SEQ ID NO: 50, linker sequence: A(EAAAK).sub.4ALEA(EAAAK).sub.4A SEQ ID NO: 51, linker sequence: [A(EAAAK).sub.nA].sub.m SEQ ID NO: 52, linker sequence [12 amino acids]: AEAAAKEAAAKA SEQ ID NO: 53, linker sequence [2 amino acid repeats]: (XP).sub.n SEQ ID NO: 54, linker sequence: (AP).sub.n SEQ ID NO: 55, linker sequence: (KP).sub.n SEQ ID NO: 56, linker sequence: (QP).sub.n SEQ ID NO: 57, linker sequence [14 amino acids]: APAPAPAPAPAPAP SEQ ID NO: 58, GAG linker sequence [21 amino acids]: GAPGGGGGAAAAAGGGGGGAP SEQ ID NO: 59, GAG2 linker sequence [39 amino acids]: GAPGGGGGAAAAAGGGGGGAPGGGGGAAAAAGGGGGGAP SEQ ID NO: 60, GAG3 linker sequence [57 amino acids]: GAPGGGGGAAAAAGGGGGGAPGGGGGAAAAAGGGGGGAPGGGGGAAAAAGGGGGGAP
REFERENCES
[0283] Anttila M, Eskola J, Ahman H, Kayhty H. Avidity of IgG for Streptococcus pneumoniae type 6B and 23F polysaccharides in infants primed with pneumococcal conjugates and boosted with polysaccharide or conjugate vaccines. J Infect Dis. 1998 June; 177(6):1614-21. [0284] Ausubel F M et al. 1995. Current Protocols in Molecular Biology, N.Y., John Wiley & Sons. [0285] Berry, A. M. et al. (1995) Infection and Immunity 63: 1969-74) [0286] Centers for Disease Control and Prevention. Preventing pneumococcal disease among infants and young children. Morbidity and Mortality Weekly Report. 2000; 49:1-55. [0287] Centers for Disease Control and Prevention. Prevention of pneumococcal disease among infants and children--use of 13-valent pneumococcal conjugate vaccine and 23-valent pneumococcal polysaccharide vaccine. Morbidity and Mortality Weekly Report. 2010; 59:1-24. [0288] Coligan J E et al. 2018. Current Protocols in Protein Science, John Wiley & Sons, Inc. [0289] Daniels, C. C. et al. (2010) Infection and Immunity 7S:2163-72 [0290] Davis et al. 1986. Basic Methods in Molecular Biology, Elsevir Sciences Publishing, Inc., New York. [0291] Douce et al. Mutants of Escherichia coli heat-labile toxin lacking ADP-ribosyltransferase activity act as non-toxic, mucosal adjuvants. PNAS Vol. 92, pp. 1644-1648, February 1995. [0292] Douce et al. Genetically detoxified mutants of heat-labile toxin from Escherichia coli are able to act as oral adjuvants" Infect Immun. 1999 September; 67(9):4400-6) [0293] Dracopoli, N et al. 2018. Current Protocols in Human Genetics, John Wiley & Sons, Inc.; [0294] Evans J T et al. Enhancement of antigen-specific immunity via the TLR-4 ligands MPL adjuvant and Ribi.529. Expert Rev Vaccines 2003 April; 2(2):219-29. [0295] Ferreira et al. "DNA vaccines based on genetically detoxified derivatives of pneumolysin fail to protect mice against challenge with Streptococcus pneumoniae" FEMS Immunol Med Microbial (2006) 46: 291-297) [0296] Giuliani M M et al. Mucosal adjuvanticity and immunogenicity of LTR72, a novel mutant of Escherichia coli heat-labile enterotoxin with partial knockout of ADP-ribosyltransferase activity. J Exp Med. 1998 Apr. 6; 187(7):1123-32. [0297] Gruber M F, Pratt D, Haase M. Licensing of pneumococcal conjugate vaccines for children and adults: Regulatory perspective from the European Medicines Agency and the U.S. Food and Drug Administration. In: Siber G R, Klugman K P, Makela P H, eds. Pneumococcal Vaccines: The Impact of Conjugate Vaccine. Washington, D.C.: ASM Press; 2008; 183-96. [0298] Hames et al. 1985. Nucleic Acid Hybridization, IL Press. [0299] Helppolainen S H, Nurminen K P, Maatta J A, Halling K K, Slotte J P, Huhtala T, et al. Rhizavidin from Rhizobium etli: the first natural dimer in the avidin protein family. Biochem J. 2007; 405:397-405. [0300] Holliger P, Prospero T, Winter G. "Diabodies": small bivalent and bispecific antibody fragments. Proc Natl Acad Sci USA. 1993 Jul. 15; 90(14):6444-8. [0301] Ishizaka S T et al. "E6020: a synthetic Toll-like receptor 4 agonist as a vaccine adjuvant." Expert Rev. Vaccines. 2007 October; 6(5):773-84. [0302] Kim K H, Yu J, Nahm M H. Efficiency of a pneumococcal opsonophagocytic killing assay improved by multiplexing and by coloring colonies. Clin Diagn Lab Immunol. 2003 July; 10(4):616-21. [0303] Kojima K, Ishizaka A, Oshika E, Taguchi Y, Tomizawa K, et al. Quantitation of IgG subclass antibodies to pneumococcal capsular polysaccharides by ELISA, using Pneumovax-specific antibodies as a reference. Tohoku J Exp Med. 1990 July; 161(3):209-15. [0304] Koskela M, Leinonen M. Comparison of ELISA and RIA for measurement of pneumococcal antibodies before and after vaccination with 14-valent pneumococcal capsular polysaccharide vaccine. J Clin Pathol. 1981 January; 34(1):93-8. [0305] Martin, E W, Ed. Remington's Pharmaceutical Sciences. 15th ed. Easton, Pa.: Mack Publishing Company, 1975. [0306] Martinez J E, Romero-Steiner S, Pilishvili T, Barnard S, Schinsky J, et al. A flow cytometric opsonophagocytic assay for measurement of functional antibodies elicited after vaccination with the 23-valent pneumococcal polysaccharide vaccine. Clin Diagn Lab Immunol. 1999 July; 6(4):581-6. [0307] Meyers and Miller. CABIOS, 1989, 4:11-17. [0308] Mitchell L. Protective Immune Responses to Streptococcus pneumoniae Pneumolysoids, ASM2011 conference abstract, 2011. [0309] Moffitt K L, Malley R, Lu, Y-J. Identification of Protective Pneumococcal Th17 Antigens from the Soluble Fraction of a Killed Whole Cell Vaccine. 2012 PLoS ONE 7(8):e43445. [0310] Munro C S, Stanley P J, Cole P J. Assessment of biological activity of immunoglobulin preparations by using opsonized micro-organisms to stimulate neutrophil chemiluminescence. Clin Exp Immunol. 1985 July; 61(1):183-8. [0311] Ojo-Amaize E A, Church J A, Barka N E, Agopian M S, Peter J B. A rapid and sensitive chemiluminescence assay for evaluation of functional opsonic activity of Haemophilus influenzae type b-specific antibodies. Clin Diagn Lab Immunol. 1995 May; 2(3):286-90. [0312] Paton J. C. et al. "Purification and immunogenicity of genetically obtained pneumolysin toxoids and their conjugation to Streptococcus pneumoniae type 19F polysaccharide" Infect Immun 1991 July; 59(7):2297-304 [0313] PNEUMOVAX.RTM. 23 (prescribing information). Whitehouse Station, NJ: Merck & Co.; May 2015. [0314] PREVNAR 13.RTM. (prescribing information). New York, N.Y.: Pfizer; August 2017. [0315] Poljak R J. Production and structure of diabodies. Structure. 1994 Dec. 15; 2(12):1121-3. [0316] Powell M F and Newman M J, Eds. 1995. Vaccine Design: The Subunit and Adjuvant Approach. New York, N.Y.: Plenum Press. [0317] Richter S S, Diekema D J, Heilmann K P, Dohrn C L, Riahi F, Doern G V. Changes in pneumococcal serotypes and antimicrobial resistance after introduction of the 13 valent conjugate vaccine in the United States. Antimicrob Agents Chemother. 2014; 58:6484-9. [0318] Romero-Steiner S, Libutti D, Pais L B, Dykes J, Anderson P, et al. Standardization of an opsonophagocytic assay for the measurement of functional antibody activity against Streptococcus pneumoniae using differentiated HL-60 cells. Clin Diagn Lab Immunol. 1997 July; 4(4):415-22. [0319] Romero-Steiner S, Holder P F, Gomez de Leon P, Spear W, Hennessy T W, et al. Avidity determinations for Haemophilus influenzae Type b anti-polyribosylribitol phosphate antibodies. Clin Diagn Lab Immunol. 2005 September; 12(9):1029-35. [0320] Saeland E, Vidarsson G, Jonsdottir I. Pneumococcal pneumonia and bacteremia model in mice for the analysis of protective antibodies. Microb Pathog. 2000 August; 29(2):81-91. [0321] Sambrook J et al. 1989. Molecular Cloning, a Laboratory Manual. Cold Harbor Laboratory Press, Cold Spring Harbor, N.Y. [0322] Saunders F. K. et al. Pneumolysin, the thiol-activated toxin of Streptococcus pneumoniae, does not require a thiol group for in vitro activity. Infect Immun 1989 August; 57(8):2547-52). Singh et al. Curr. HIV Res. 2003 1:309-20. [0323] Stack A M, Malley R, Thompson C M, Kobzik L, Siber G R, et al. Minimum protective serum concentrations of pneumococcal anti-capsular antibodies in infant rats. J Infect Dis. 1998 April; 177(4):986-90. [0324] Williams et al, Innate imprinting by the modified heat-labile toxin of Escherichia coli (LTK63) provides generic protection against lung infectious disease. The Journal of Immunology 2004 173:7435-7443. [0325] Wu W, Huang J, Duan B, Traficante D C, Hong H, et al. Th17-stimulating protein vaccines confer protection against Pseudomonas aeruginosa pneumonia. Am J Respir Crit Care Med. 2012 Sep. 1; 186(5):420-7. [0326] Zhang F, Lu Y J, Malley R. Multiple antigen-presenting system (MAPS) to induce comprehensive B- and T-cell immunity. Proc Natl Acad Sci USA. 2013; 110:13564-9. [0327] WO2012/155007 [0328] WO 2012/155053 [0329] US 2009/0285846
EQUIVALENTS
[0330] Those skilled in the art will recognize, or be able to ascertain using no more than routine experimentation, many equivalents to the specific embodiments of the invention described herein. The scope of the present invention is not intended to be limited to the above Description, but rather is as set forth in the following claims:
Sequence CWU 1
1
1571179PRTUnknownDescription of Unknown Rhizavidin sequence 1Met
Ile Ile Thr Ser Leu Tyr Ala Thr Phe Gly Thr Ile Ala Asp Gly1 5 10
15Arg Arg Thr Ser Gly Gly Lys Thr Met Ile Arg Thr Asn Ala Val Ala
20 25 30Ala Leu Val Phe Ala Val Ala Thr Ser Ala Leu Ala Phe Asp Ala
Ser 35 40 45Asn Phe Lys Asp Phe Ser Ser Ile Ala Ser Ala Ser Ser Ser
Trp Gln 50 55 60Asn Gln Ser Gly Ser Thr Met Ile Ile Gln Val Asp Ser
Phe Gly Asn65 70 75 80Val Ser Gly Gln Tyr Val Asn Arg Ala Gln Gly
Thr Gly Cys Gln Asn 85 90 95Ser Pro Tyr Pro Leu Thr Gly Arg Val Asn
Gly Thr Phe Ile Ala Phe 100 105 110Ser Val Gly Trp Asn Asn Ser Thr
Glu Asn Cys Asn Ser Ala Thr Gly 115 120 125Trp Thr Gly Tyr Ala Gln
Val Asn Gly Asn Asn Thr Glu Ile Val Thr 130 135 140Ser Trp Asn Leu
Ala Tyr Glu Gly Gly Ser Gly Pro Ala Ile Glu Gln145 150 155 160Gly
Gln Asp Thr Phe Gln Tyr Val Pro Thr Thr Glu Asn Lys Ser Leu 165 170
175Leu Lys Asp2135PRTUnknownDescription of Unknown Truncated
Rhizavidin sequence 2Phe Asp Ala Ser Asn Phe Lys Asp Phe Ser Ser
Ile Ala Ser Ala Ser1 5 10 15Ser Ser Trp Gln Asn Gln Ser Gly Ser Thr
Met Ile Ile Gln Val Asp 20 25 30Ser Phe Gly Asn Val Ser Gly Gln Tyr
Val Asn Arg Ala Gln Gly Thr 35 40 45Gly Cys Gln Asn Ser Pro Tyr Pro
Leu Thr Gly Arg Val Asn Gly Thr 50 55 60Phe Ile Ala Phe Ser Val Gly
Trp Asn Asn Ser Thr Glu Asn Cys Asn65 70 75 80Ser Ala Thr Gly Trp
Thr Gly Tyr Ala Gln Val Asn Gly Asn Asn Thr 85 90 95Glu Ile Val Thr
Ser Trp Asn Leu Ala Tyr Glu Gly Gly Ser Gly Pro 100 105 110Ala Ile
Glu Gln Gly Gln Asp Thr Phe Gln Tyr Val Pro Thr Thr Glu 115 120
125Asn Lys Ser Leu Leu Lys Asp 130 1353399PRTStreptococcus
pneumoniae 3Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr
Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly
Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu
Pro Thr His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser
Val Leu Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val
Tyr Phe Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val
Ser Val Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr
Ser Ser Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg
Ala Val Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln
Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro
Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150
155 160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp
Ala 165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys
Ala Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu
Glu Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser
Pro Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn
Glu Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn
Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser
Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265
270Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala
275 280 285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val
Thr Gly 290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn
Ile Glu Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro
Val Ser Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp
Ile Val Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser
Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly
Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser
Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
3954367PRTStreptococcus pneumoniae 4Phe Arg Gln Pro Ser Gln Thr Ala
Leu Lys Asp Glu Pro Thr His Leu1 5 10 15Val Val Ala Lys Glu Gly Ser
Val Ala Ser Ser Val Leu Leu Ser Gly 20 25 30Thr Val Thr Ala Lys Asn
Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys 35 40 45Gly Asp Leu Asp Glu
Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 50 55 60Gly Gln Ala Leu
Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr65 70 75 80Asp Ser
Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn Glu 85 90 95Leu
Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 100 105
110Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro
115 120 125Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly
Asp Ala 130 135 140Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser
Lys Ala Gln Ser145 150 155 160Gln Leu Asp Ala Thr Thr Val Leu Ser
Thr Leu Glu Gly Thr Val Val 165 170 175Glu Val Asn Ser Asn Val Ser
Lys Ser Pro Thr Gly Ala Ser Gln Val 180 185 190Met Val His Ile Val
Ser Asn Glu Asn Leu Gln Val Lys Gly Glu Leu 195 200 205Ser Glu Tyr
Asn Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 210 215 220Thr
Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr225 230
235 240Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala
Ala 245 250 255Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp
Val Thr Gly 260 265 270Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val
Asn Ile Glu Val Lys 275 280 285Ser Lys Thr Lys Ala Ile Leu Val Pro
Val Ser Ser Leu Val Met Asp 290 295 300Asp Ser Lys Asn Tyr Val Trp
Ile Val Asp Glu Gln Gln Lys Ala Lys305 310 315 320Lys Val Glu Val
Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile 325 330 335Thr Ser
Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 340 345
350Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn 355
360 3655399PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptideMOD_RES(37)..(37)Any amino
acidMOD_RES(41)..(41)Any amino acidMOD_RES(45)..(46)Any amino
acidMOD_RES(48)..(49)Any amino acidMOD_RES(51)..(52)Any amino
acidMOD_RES(60)..(60)Any amino acidMOD_RES(111)..(111)Any amino
acidMOD_RES(121)..(122)Any amino acidMOD_RES(124)..(124)Any amino
acidMOD_RES(132)..(132)Any amino acidMOD_RES(140)..(141)Any amino
acidMOD_RES(156)..(157)Any amino acidMOD_RES(269)..(269)Any amino
acidMOD_RES(282)..(282)Any amino acidMOD_RES(288)..(288)Any amino
acidMOD_RES(306)..(306)Any amino acidMOD_RES(308)..(308)Any amino
acidMOD_RES(337)..(337)Any amino acidMOD_RES(351)..(351)Any amino
acidMOD_RES(384)..(384)Any amino acidMOD_RES(394)..(394)Any amino
acid 5Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala
Ala1 5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile
Leu Leu 20 25 30Phe Arg Gln Pro Xaa Gln Thr Ala Xaa Lys Asp Glu Xaa
Xaa His Xaa 35 40 45Xaa Val Xaa Xaa Glu Gly Ser Val Ala Ser Ser Xaa
Leu Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr
Phe Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser
Val Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser
Ser Ser Glu Ala Gln Ala Xaa Tyr 100 105 110Asp Ser Ala Ser Arg Ala
Val Ala Xaa Xaa Asp Xaa His Ile Asn Glu 115 120 125Leu Asn Gln Xaa
Arg Asn Glu Ala Ala Ser Ala Xaa Xaa Pro Gln Leu 130 135 140Pro Ala
Pro Val Gly Gly Glu Asp Ala Thr Val Xaa Xaa Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Xaa Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Xaa Ala Ala Ser Pro Ala Xaa 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Xaa Gly Xaa Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Xaa Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Xaa Lys 340 345 350Lys Val Glu Val Ser Leu Gly
Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr
Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Xaa 370 375 380Ser Leu Glu
Glu Gly Lys Glu Val Lys Xaa Asp Glu Ala Thr Asn385 390
3956278PRTStreptococcus pneumoniae 6Met Lys Lys Trp Met Leu Val Leu
Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys Asn
Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Gln Ser Asn
Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met Gly
Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp Leu
Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn Trp
Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90 95Gly
Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg 100 105
110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val
115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp
Met Thr 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser Ser Gly
Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Glu Ile Leu Lys Asn
Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr Phe Asn
Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp Gly Leu
Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu Ala Glu
Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215 220Glu
Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Asn Leu225 230
235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly
Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly Glu Asp Val
Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
2757252PRTStreptococcus pneumoniae 7Thr Ser Gly Asp Asn Trp Ser Lys
Tyr Gln Ser Asn Lys Ser Ile Thr1 5 10 15Ile Gly Phe Asp Ser Thr Phe
Val Pro Met Gly Phe Ala Gln Lys Asp 20 25 30Gly Ser Tyr Ala Gly Phe
Asp Ile Asp Leu Ala Thr Ala Val Phe Glu 35 40 45Lys Tyr Gly Ile Thr
Val Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys 50 55 60Glu Ala Glu Leu
Thr Lys Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr65 70 75 80Ser Ala
Thr Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr 85 90 95Met
Lys Asn Glu Gln Val Leu Val Thr Lys Lys Ser Ser Gly Ile Thr 100 105
110Thr Ala Lys Asp Met Thr Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
115 120 125Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro Glu Ile Leu Lys
Asn Ile 130 135 140Val Ala Asn Lys Glu Ala Asn Gln Tyr Gln Thr Phe
Asn Glu Ala Leu145 150 155 160Ile Asp Leu Lys Asn Asp Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val 165 170 175Tyr Ala Asn Tyr Tyr Leu Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn 180 185 190Val Phe Thr Val Gly
Leu Glu Thr Glu Ala Phe Ala Val Gly Ala Arg 195 200 205Lys Glu Asp
Thr Asn Leu Val Lys Lys Ile Asn Glu Ala Phe Ser Ser 210 215 220Leu
Tyr Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly225 230
235 240Glu Asp Val Ala Thr Lys Glu Val Lys Glu Gly Gln 245
2508274PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptideMOD_RES(1)..(2)Any amino
acidMOD_RES(9)..(9)Any amino acidMOD_RES(20)..(20)Any amino
acidMOD_RES(23)..(23)Any amino acidMOD_RES(32)..(32)Any amino
acidMOD_RES(76)..(76)Any amino acidMOD_RES(106)..(106)Any amino
acidMOD_RES(116)..(116)Any amino acidMOD_RES(130)..(130)Any amino
acidMOD_RES(140)..(140)Any amino acidMOD_RES(143)..(143)Any amino
acidMOD_RES(161)..(162)Any amino acidMOD_RES(165)..(165)Any amino
acidMOD_RES(172)..(172)Any amino acidMOD_RES(177)..(177)Any amino
acidMOD_RES(191)..(191)Any amino acidMOD_RES(222)..(222)Any amino
acidMOD_RES(229)..(229)Any amino acidMOD_RES(234)..(235)Any amino
acidMOD_RES(243)..(243)Any amino acidMOD_RES(250)..(250)Any amino
acidMOD_RES(264)..(264)Any amino acidMOD_RES(274)..(274)Any amino
acid 8Xaa Xaa Val Leu Val Ser Leu Met Xaa Ala Leu Phe Leu Val Ala
Cys1 5 10 15Gly Lys Asn Xaa Ser Glu Xaa Ser Gly Asp Asn Trp Ser Lys
Tyr Xaa 20 25 30Ser Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr Phe
Val Pro Met 35 40 45Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe
Asp Ile Asp Leu 50 55 60Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr
Xaa Asn Trp Gln Pro65 70 75 80Ile Asp Trp Asp Leu Lys Glu Ala Glu
Leu Thr Lys Gly Thr Ile Asp 85 90 95Leu Ile Trp Asn Gly Tyr Ser Ala
Thr Xaa Glu Arg Arg Glu Lys Val 100 105 110Ala Phe Ser Xaa Ser Tyr
Met Lys Asn Glu Gln Val Leu Val Thr Lys 115 120 125Lys Xaa Ser Gly
Ile Thr Thr Ala Lys Asp Met Xaa Gly Lys Xaa Leu 130 135 140Gly Ala
Gln Ala Gly Ser Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro145 150 155
160Xaa Xaa Leu Lys Xaa Ile Val Ala Asn Lys Glu Xaa Asn Gln Tyr Gln
165 170 175Xaa Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp Arg Ile
Xaa Gly 180
185 190Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu
Gly 195 200 205Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu Glu
Xaa Glu Ala 210 215 220Phe Ala Val Gly Xaa Arg Lys Glu Asp Xaa Xaa
Leu Val Lys Lys Ile225 230 235 240Asn Glu Xaa Phe Ser Ser Leu Tyr
Lys Xaa Gly Lys Phe Gln Glu Ile 245 250 255Ser Gln Lys Trp Phe Gly
Glu Xaa Val Ala Thr Lys Glu Val Lys Glu 260 265 270Gly
Xaa9540DNAUnknownDescription of Unknown Rhizavidin sequence
9ttgattatta cgagtttata tgcaaccttc ggtacgatcg ccgacggtcg gcgcacatca
60ggaggaaaaa ccatgatccg tactaatgca gttgcagcac ttgtgtttgc tgtagcaacc
120tcagctcttg ctttcgatgc aagcaacttc aaggattttt caagcatcgc
atcggcttcc 180agcagttggc agaaccagtc cggctcgacg atgatcattc
aagtcgactc gttcggaaat 240gtctccggcc aatatgtaaa cagagcccag
ggcaccggat gccagaactc gccctatccg 300ctaacaggaa gggtaaacgg
gacgttcatc gcattttcgg tcggctggaa caattcgacg 360gagaactgca
attccgcaac cggatggacc ggctacgcac aggtcaacgg caacaacact
420gagatagtca cgagctggaa cctcgcttac gaaggcggct ccggtccggc
cattgagcaa 480ggacaagaca ctttccagta cgtgccgacg actgagaaca
aaagcctctt gaaggattaa 54010405DNAUnknownDescription of Unknown
Truncated Rhizavidin sequence 10tttgatgcga gtaactttaa agatttcagc
tctattgcga gcgccagtag ctcttggcag 60aatcagagtg gcagcaccat gattatccag
gtggattctt tcggcaacgt tagtggccag 120tatgttaatc gtgcgcaggg
cacgggttgc cagaactctc cgtacccgct gaccggccgc 180gtgaatggca
cgtttatcgc cttcagcgtt ggctggaaca attctaccga aaactgtaat
240agtgcaaccg gctggacggg ttatgcgcag gtgaacggta acaataccga
aattgttacg 300agctggaatc tggcctatga aggcggttct ggcccggcaa
tcgaacaggg tcaggatacc 360tttcagtacg ttccgaccac ggaaaacaaa
agcctgctga aagat 405111197DNAStreptococcus pneumoniae 11atgaagaaaa
agaatggtaa agctaaaaag tggcaactgt atgcagcaat cggtgctgcg 60agtgtagttg
tattgggtgc tggggggatt ttactcttta gacaaccttc tcagactgct
120ctaaaagatg agcctactca tcttgttgtt gccaaggaag gaagcgtggc
ctcctctgtt 180ttattgtcag ggacagtaac agcaaaaaat gaacaatatg
tttattttga tgctagtaag 240ggtgatttag atgaaatcct tgtttctgtg
ggcgataagg tcagcgaagg gcaggcttta 300gtcaagtaca gtagttcaga
agcgcaggcg gcctatgatt cagctagtcg agcagtagct 360agggcagatc
gtcatatcaa tgaactcaat caagcacgaa atgaagccgc ttcagctccg
420gctccacagt taccagcgcc agtaggagga gaagatgcaa cggtgcaaag
cccaactcca 480gtggctggaa attctgttgc ttctattgac gctcaattgg
gtgatgcccg tgatgcgcgt 540gcagatgctg cggcgcaatt aagcaaggct
caaagtcaat tggatgcaac aactgttctc 600agtaccctag agggaactgt
ggtcgaagtc aatagcaatg tttctaaatc tccaacaggg 660gcgagtcaag
ttatggttca tattgtcagc aatgaaaatt tacaagtcaa gggagaattg
720tctgagtaca atctagccaa cctttctgta ggtcaagaag taagctttac
ttctaaagtg 780tatcctgata aaaaatggac tgggaaatta agctatattt
ctgactatcc taaaaacaat 840ggtgaagcag ctagtccagc agccgggaat
aatacaggtt ctaaataccc ttatactatt 900gatgtgacag gcgaggttgg
tgatttgaaa caaggttttt ctgtcaacat tgaggttaaa 960agcaaaacta
aggctattct tgttcctgtt agcagtctag taatggatga tagtaaaaat
1020tatgtctgga ttgtggatga acaacaaaag gctaaaaaag ttgaggtttc
attgggaaat 1080gctgacgcag aaaatcaaga aatcacttct ggtttaacga
acggtgctaa ggtcatcagt 1140aatccaacat cttccttgga agaaggaaaa
gaggtgaagg ctgatgaagc aactaat 1197121101DNAStreptococcus pneumoniae
12tttagacaac cttctcagac tgctctaaaa gatgagccta ctcatcttgt tgttgccaag
60gaaggaagcg tggcctcctc tgttttattg tcagggacag taacagcaaa aaatgaacaa
120tatgtttatt ttgatgctag taagggtgat ttagatgaaa tccttgtttc
tgtgggcgat 180aaggtcagcg aagggcaggc tttagtcaag tacagtagtt
cagaagcgca ggcggcctat 240gattcagcta gtcgagcagt agctagggca
gatcgtcata tcaatgaact caatcaagca 300cgaaatgaag ccgcttcagc
tccggctcca cagttaccag cgccagtagg aggagaagat 360gcaacggtgc
aaagcccaac tccagtggct ggaaattctg ttgcttctat tgacgctcaa
420ttgggtgatg cccgtgatgc gcgtgcagat gctgcggcgc aattaagcaa
ggctcaaagt 480caattggatg caacaactgt tctcagtacc ctagagggaa
ctgtggtcga agtcaatagc 540aatgtttcta aatctccaac aggggcgagt
caagttatgg ttcatattgt cagcaatgaa 600aatttacaag tcaagggaga
attgtctgag tacaatctag ccaacctttc tgtaggtcaa 660gaagtaagct
ttacttctaa agtgtatcct gataaaaaat ggactgggaa attaagctat
720atttctgact atcctaaaaa caatggtgaa gcagctagtc cagcagccgg
gaataataca 780ggttctaaat acccttatac tattgatgtg acaggcgagg
ttggtgattt gaaacaaggt 840ttttctgtca acattgaggt taaaagcaaa
actaaggcta ttcttgttcc tgttagcagt 900ctagtaatgg atgatagtaa
aaattatgtc tggattgtgg atgaacaaca aaaggctaaa 960aaagttgagg
tttcattggg aaatgctgac gcagaaaatc aagaaatcac ttctggttta
1020acgaacggtg ctaaggtcat cagtaatcca acatcttcct tggaagaagg
aaaagaggtg 1080aaggctgatg aagcaactaa t 110113834DNAStreptococcus
pneumoniae 13atgaaaaaat ggatgcttgt attagtcagt ctgatgactg ctttgttctt
agtagcttgt 60gggaaaaatt ctagcgaaac tagtggagat aattggtcaa agtaccagtc
taacaagtct 120attactattg gatttgatag tacttttgtt ccaatgggat
ttgctcagaa agatggttct 180tatgcaggat ttgatattga tttagctaca
gctgtttttg aaaaatacgg aatcacggta 240aattggcaac cgattgattg
ggatttgaaa gaagctgaat tgacaaaagg aacgattgat 300ctgatttgga
atggctattc cgctacagac gaacgccgtg aaaaggtggc tttcagtaac
360tcatatatga agaatgagca ggtattggtt acgaagaaat catctggtat
cacgactgca 420aaggatatga ctggaaagac attaggagct caagctggtt
catctggtta tgcggacttt 480gaagcaaatc cagaaatttt gaagaatatt
gtcgctaata aggaagcgaa tcaataccaa 540acctttaatg aagccttgat
tgatttgaaa aacgatcgaa ttgatggtct attgattgac 600cgtgtctatg
caaactatta tttagaagca gaaggtgttt taaacgatta taatgtcttt
660acagttggac tagaaacaga agcttttgcg gttggagccc gtaaggaaga
tacaaacttg 720gttaagaaga taaatgaagc tttttctagt ctttacaagg
acggcaagtt ccaagaaatc 780agccaaaaat ggtttggaga agatgtagca
accaaagaag taaaagaagg acag 83414756DNAStreptococcus pneumoniae
14actagtggag ataattggtc aaagtaccag tctaacaagt ctattactat tggatttgat
60agtacttttg ttccaatggg atttgctcag aaagatggtt cttatgcagg atttgatatt
120gatttagcta cagctgtttt tgaaaaatac ggaatcacgg taaattggca
accgattgat 180tgggatttga aagaagctga attgacaaaa ggaacgattg
atctgatttg gaatggctat 240tccgctacag acgaacgccg tgaaaaggtg
gctttcagta actcatatat gaagaatgag 300caggtattgg ttacgaagaa
atcatctggt atcacgactg caaaggatat gactggaaag 360acattaggag
ctcaagctgg ttcatctggt tatgcggact ttgaagcaaa tccagaaatt
420ttgaagaata ttgtcgctaa taaggaagcg aatcaatacc aaacctttaa
tgaagccttg 480attgatttga aaaacgatcg aattgatggt ctattgattg
accgtgtcta tgcaaactat 540tatttagaag cagaaggtgt tttaaacgat
tataatgtct ttacagttgg actagaaaca 600gaagcttttg cggttggagc
ccgtaaggaa gatacaaact tggttaagaa gataaatgaa 660gctttttcta
gtctttacaa ggacggcaag ttccaagaaa tcagccaaaa atggtttgga
720gaagatgtag caaccaaaga agtaaaagaa ggacag 756156PRTArtificial
SequenceDescription of Artificial Sequence Synthetic 6xHis tag
15His His His His His His1 51610PRTArtificial SequenceDescription
of Artificial Sequence Synthetic peptide 16Met Ser Tyr Tyr His His
His His His His1 5 1017620PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 17Met Thr Ser Gly Asp Asn
Trp Ser Lys Tyr Gln Ser Asn Lys Ser Ile1 5 10 15Thr Ile Gly Phe Asp
Ser Thr Phe Val Pro Met Gly Phe Ala Gln Lys 20 25 30Asp Gly Ser Tyr
Ala Gly Phe Asp Ile Asp Leu Ala Thr Ala Val Phe 35 40 45Glu Lys Tyr
Gly Ile Thr Val Asn Trp Gln Pro Ile Asp Trp Asp Leu 50 55 60Lys Glu
Ala Glu Leu Thr Lys Gly Thr Ile Asp Leu Ile Trp Asn Gly65 70 75
80Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser
85 90 95Tyr Met Lys Asn Glu Gln Val Leu Val Thr Lys Lys Ser Ser Gly
Ile 100 105 110Thr Thr Ala Lys Asp Met Thr Gly Lys Thr Leu Gly Ala
Gln Ala Gly 115 120 125Ser Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro
Glu Ile Leu Lys Asn 130 135 140Ile Val Ala Asn Lys Glu Ala Asn Gln
Tyr Gln Thr Phe Asn Glu Ala145 150 155 160Leu Ile Asp Leu Lys Asn
Asp Arg Ile Asp Gly Leu Leu Ile Asp Arg 165 170 175Val Tyr Ala Asn
Tyr Tyr Leu Glu Ala Glu Gly Val Leu Asn Asp Tyr 180 185 190Asn Val
Phe Thr Val Gly Leu Glu Thr Glu Ala Phe Ala Val Gly Ala 195 200
205Arg Lys Glu Asp Thr Asn Leu Val Lys Lys Ile Asn Glu Ala Phe Ser
210 215 220Ser Leu Tyr Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys
Trp Phe225 230 235 240Gly Glu Asp Val Ala Thr Lys Glu Val Lys Glu
Gly Gln Phe Arg Gln 245 250 255Pro Ser Gln Thr Ala Leu Lys Asp Glu
Pro Thr His Leu Val Val Ala 260 265 270Lys Glu Gly Ser Val Ala Ser
Ser Val Leu Leu Ser Gly Thr Val Thr 275 280 285Ala Lys Asn Glu Gln
Tyr Val Tyr Phe Asp Ala Ser Lys Gly Asp Leu 290 295 300Asp Glu Ile
Leu Val Ser Val Gly Asp Lys Val Ser Glu Gly Gln Ala305 310 315
320Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr Asp Ser Ala
325 330 335Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn Glu Leu
Asn Gln 340 345 350Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln
Leu Pro Ala Pro 355 360 365Val Gly Gly Glu Asp Ala Thr Val Gln Ser
Pro Thr Pro Val Ala Gly 370 375 380Asn Ser Val Ala Ser Ile Asp Ala
Gln Leu Gly Asp Ala Arg Asp Ala385 390 395 400Arg Ala Asp Ala Ala
Ala Gln Leu Ser Lys Ala Gln Ser Gln Leu Asp 405 410 415Ala Thr Thr
Val Leu Ser Thr Leu Glu Gly Thr Val Val Glu Val Asn 420 425 430Ser
Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val Met Val His 435 440
445Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu Leu Ser Glu Tyr
450 455 460Asn Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe Thr
Ser Lys465 470 475 480Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu
Ser Tyr Ile Ser Asp 485 490 495Tyr Pro Lys Asn Asn Gly Glu Ala Ala
Ser Pro Ala Ala Gly Asn Asn 500 505 510Thr Gly Ser Lys Tyr Pro Tyr
Thr Ile Asp Val Thr Gly Glu Val Gly 515 520 525Asp Leu Lys Gln Gly
Phe Ser Val Asn Ile Glu Val Lys Ser Lys Thr 530 535 540Lys Ala Ile
Leu Val Pro Val Ser Ser Leu Val Met Asp Asp Ser Lys545 550 555
560Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys Lys Val Glu
565 570 575Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile Thr
Ser Gly 580 585 590Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr
Ser Ser Leu Glu 595 600 605Glu Gly Lys Glu Val Lys Ala Asp Glu Ala
Thr Asn 610 615 62018620PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 18Met Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Thr His1 5 10 15Leu Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser 20 25 30Gly Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser 35 40 45Lys Gly Asp
Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser 50 55 60Glu Gly
Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala65 70 75
80Tyr Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn
85 90 95Glu Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro
Gln 100 105 110Leu Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln
Ser Pro Thr 115 120 125Pro Val Ala Gly Asn Ser Val Ala Ser Ile Asp
Ala Gln Leu Gly Asp 130 135 140Ala Arg Asp Ala Arg Ala Asp Ala Ala
Ala Gln Leu Ser Lys Ala Gln145 150 155 160Ser Gln Leu Asp Ala Thr
Thr Val Leu Ser Thr Leu Glu Gly Thr Val 165 170 175Val Glu Val Asn
Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln 180 185 190Val Met
Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu 195 200
205Leu Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser
210 215 220Phe Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys
Leu Ser225 230 235 240Tyr Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu
Ala Ala Ser Pro Ala 245 250 255Ala Gly Asn Asn Thr Gly Ser Lys Tyr
Pro Tyr Thr Ile Asp Val Thr 260 265 270Gly Glu Val Gly Asp Leu Lys
Gln Gly Phe Ser Val Asn Ile Glu Val 275 280 285Lys Ser Lys Thr Lys
Ala Ile Leu Val Pro Val Ser Ser Leu Val Met 290 295 300Asp Asp Ser
Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala305 310 315
320Lys Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu
325 330 335Ile Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn
Pro Thr 340 345 350Ser Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp
Glu Ala Thr Asn 355 360 365Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln
Ser Asn Lys Ser Ile Thr 370 375 380Ile Gly Phe Asp Ser Thr Phe Val
Pro Met Gly Phe Ala Gln Lys Asp385 390 395 400Gly Ser Tyr Ala Gly
Phe Asp Ile Asp Leu Ala Thr Ala Val Phe Glu 405 410 415Lys Tyr Gly
Ile Thr Val Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys 420 425 430Glu
Ala Glu Leu Thr Lys Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr 435 440
445Ser Ala Thr Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr
450 455 460Met Lys Asn Glu Gln Val Leu Val Thr Lys Lys Ser Ser Gly
Ile Thr465 470 475 480Thr Ala Lys Asp Met Thr Gly Lys Thr Leu Gly
Ala Gln Ala Gly Ser 485 490 495Ser Gly Tyr Ala Asp Phe Glu Ala Asn
Pro Glu Ile Leu Lys Asn Ile 500 505 510Val Ala Asn Lys Glu Ala Asn
Gln Tyr Gln Thr Phe Asn Glu Ala Leu 515 520 525Ile Asp Leu Lys Asn
Asp Arg Ile Asp Gly Leu Leu Ile Asp Arg Val 530 535 540Tyr Ala Asn
Tyr Tyr Leu Glu Ala Glu Gly Val Leu Asn Asp Tyr Asn545 550 555
560Val Phe Thr Val Gly Leu Glu Thr Glu Ala Phe Ala Val Gly Ala Arg
565 570 575Lys Glu Asp Thr Asn Leu Val Lys Lys Ile Asn Glu Ala Phe
Ser Ser 580 585 590Leu Tyr Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln
Lys Trp Phe Gly 595 600 605Glu Asp Val Ala Thr Lys Glu Val Lys Glu
Gly Gln 610 615 62019755PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 19Met Phe Asp Ala Ser Asn
Phe Lys Asp Phe Ser Ser Ile Ala Ser Ala1 5 10 15Ser Ser Ser Trp Gln
Asn Gln Ser Gly Ser Thr Met Ile Ile Gln Val 20 25 30Asp Ser Phe Gly
Asn Val Ser Gly Gln Tyr Val Asn Arg Ala Gln Gly 35 40 45Thr Gly Cys
Gln Asn Ser Pro Tyr Pro Leu Thr Gly Arg Val Asn Gly 50 55 60Thr Phe
Ile Ala Phe Ser Val Gly Trp Asn Asn Ser Thr Glu Asn Cys65 70 75
80Asn Ser Ala Thr Gly Trp Thr Gly Tyr Ala Gln Val Asn Gly Asn Asn
85 90 95Thr Glu Ile Val Thr Ser Trp Asn Leu Ala Tyr Glu Gly Gly Ser
Gly 100 105 110Pro Ala Ile Glu Gln Gly Gln Asp Thr Phe Gln Tyr Val
Pro Thr Thr 115 120 125Glu Asn Lys Ser Leu Leu Lys Asp Thr Ser Gly
Asp Asn Trp Ser Lys 130 135 140Tyr Gln Ser Asn Lys Ser Ile Thr Ile
Gly Phe Asp Ser Thr Phe Val145 150 155 160Pro Met Gly Phe Ala Gln
Lys Asp Gly Ser Tyr Ala Gly Phe Asp Ile 165 170 175Asp Leu Ala Thr
Ala Val Phe Glu Lys Tyr Gly Ile Thr Val Asn Trp 180 185 190Gln Pro
Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys Gly Thr 195 200
205Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu
210 215
220Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val Leu
Val225 230 235 240Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp
Met Thr Gly Lys 245 250 255Thr Leu Gly Ala Gln Ala Gly Ser Ser Gly
Tyr Ala Asp Phe Glu Ala 260 265 270Asn Pro Glu Ile Leu Lys Asn Ile
Val Ala Asn Lys Glu Ala Asn Gln 275 280 285Tyr Gln Thr Phe Asn Glu
Ala Leu Ile Asp Leu Lys Asn Asp Arg Ile 290 295 300Asp Gly Leu Leu
Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu Glu Ala305 310 315 320Glu
Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu Glu Thr 325 330
335Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Asn Leu Val Lys
340 345 350Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys
Phe Gln 355 360 365Glu Ile Ser Gln Lys Trp Phe Gly Glu Asp Val Ala
Thr Lys Glu Val 370 375 380Lys Glu Gly Gln Phe Arg Gln Pro Ser Gln
Thr Ala Leu Lys Asp Glu385 390 395 400Pro Thr His Leu Val Val Ala
Lys Glu Gly Ser Val Ala Ser Ser Val 405 410 415Leu Leu Ser Gly Thr
Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe 420 425 430Asp Ala Ser
Lys Gly Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp 435 440 445Lys
Val Ser Glu Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala 450 455
460Gln Ala Ala Tyr Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp
Arg465 470 475 480His Ile Asn Glu Leu Asn Gln Ala Arg Asn Glu Ala
Ala Ser Ala Pro 485 490 495Ala Pro Gln Leu Pro Ala Pro Val Gly Gly
Glu Asp Ala Thr Val Gln 500 505 510Ser Pro Thr Pro Val Ala Gly Asn
Ser Val Ala Ser Ile Asp Ala Gln 515 520 525Leu Gly Asp Ala Arg Asp
Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser 530 535 540Lys Ala Gln Ser
Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu545 550 555 560Gly
Thr Val Val Glu Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly 565 570
575Ala Ser Gln Val Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val
580 585 590Lys Gly Glu Leu Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val
Gly Gln 595 600 605Glu Val Ser Phe Thr Ser Lys Val Tyr Pro Asp Lys
Lys Trp Thr Gly 610 615 620Lys Leu Ser Tyr Ile Ser Asp Tyr Pro Lys
Asn Asn Gly Glu Ala Ala625 630 635 640Ser Pro Ala Ala Gly Asn Asn
Thr Gly Ser Lys Tyr Pro Tyr Thr Ile 645 650 655Asp Val Thr Gly Glu
Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn 660 665 670Ile Glu Val
Lys Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser 675 680 685Leu
Val Met Asp Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln 690 695
700Gln Lys Ala Lys Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala
Glu705 710 715 720Asn Gln Glu Ile Thr Ser Gly Leu Thr Asn Gly Ala
Lys Val Ile Ser 725 730 735Asn Pro Thr Ser Ser Leu Glu Glu Gly Lys
Glu Val Lys Ala Asp Glu 740 745 750Ala Thr Asn
75520755PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 20Met Phe Asp Ala Ser Asn Phe Lys Asp Phe Ser
Ser Ile Ala Ser Ala1 5 10 15Ser Ser Ser Trp Gln Asn Gln Ser Gly Ser
Thr Met Ile Ile Gln Val 20 25 30Asp Ser Phe Gly Asn Val Ser Gly Gln
Tyr Val Asn Arg Ala Gln Gly 35 40 45Thr Gly Cys Gln Asn Ser Pro Tyr
Pro Leu Thr Gly Arg Val Asn Gly 50 55 60Thr Phe Ile Ala Phe Ser Val
Gly Trp Asn Asn Ser Thr Glu Asn Cys65 70 75 80Asn Ser Ala Thr Gly
Trp Thr Gly Tyr Ala Gln Val Asn Gly Asn Asn 85 90 95Thr Glu Ile Val
Thr Ser Trp Asn Leu Ala Tyr Glu Gly Gly Ser Gly 100 105 110Pro Ala
Ile Glu Gln Gly Gln Asp Thr Phe Gln Tyr Val Pro Thr Thr 115 120
125Glu Asn Lys Ser Leu Leu Lys Asp Phe Arg Gln Pro Ser Gln Thr Ala
130 135 140Leu Lys Asp Glu Pro Thr His Leu Val Val Ala Lys Glu Gly
Ser Val145 150 155 160Ala Ser Ser Val Leu Leu Ser Gly Thr Val Thr
Ala Lys Asn Glu Gln 165 170 175Tyr Val Tyr Phe Asp Ala Ser Lys Gly
Asp Leu Asp Glu Ile Leu Val 180 185 190Ser Val Gly Asp Lys Val Ser
Glu Gly Gln Ala Leu Val Lys Tyr Ser 195 200 205Ser Ser Glu Ala Gln
Ala Ala Tyr Asp Ser Ala Ser Arg Ala Val Ala 210 215 220Arg Ala Asp
Arg His Ile Asn Glu Leu Asn Gln Ala Arg Asn Glu Ala225 230 235
240Ala Ser Ala Pro Ala Pro Gln Leu Pro Ala Pro Val Gly Gly Glu Asp
245 250 255Ala Thr Val Gln Ser Pro Thr Pro Val Ala Gly Asn Ser Val
Ala Ser 260 265 270Ile Asp Ala Gln Leu Gly Asp Ala Arg Asp Ala Arg
Ala Asp Ala Ala 275 280 285Ala Gln Leu Ser Lys Ala Gln Ser Gln Leu
Asp Ala Thr Thr Val Leu 290 295 300Ser Thr Leu Glu Gly Thr Val Val
Glu Val Asn Ser Asn Val Ser Lys305 310 315 320Ser Pro Thr Gly Ala
Ser Gln Val Met Val His Ile Val Ser Asn Glu 325 330 335Asn Leu Gln
Val Lys Gly Glu Leu Ser Glu Tyr Asn Leu Ala Asn Leu 340 345 350Ser
Val Gly Gln Glu Val Ser Phe Thr Ser Lys Val Tyr Pro Asp Lys 355 360
365Lys Trp Thr Gly Lys Leu Ser Tyr Ile Ser Asp Tyr Pro Lys Asn Asn
370 375 380Gly Glu Ala Ala Ser Pro Ala Ala Gly Asn Asn Thr Gly Ser
Lys Tyr385 390 395 400Pro Tyr Thr Ile Asp Val Thr Gly Glu Val Gly
Asp Leu Lys Gln Gly 405 410 415Phe Ser Val Asn Ile Glu Val Lys Ser
Lys Thr Lys Ala Ile Leu Val 420 425 430Pro Val Ser Ser Leu Val Met
Asp Asp Ser Lys Asn Tyr Val Trp Ile 435 440 445Val Asp Glu Gln Gln
Lys Ala Lys Lys Val Glu Val Ser Leu Gly Asn 450 455 460Ala Asp Ala
Glu Asn Gln Glu Ile Thr Ser Gly Leu Thr Asn Gly Ala465 470 475
480Lys Val Ile Ser Asn Pro Thr Ser Ser Leu Glu Glu Gly Lys Glu Val
485 490 495Lys Ala Asp Glu Ala Thr Asn Thr Ser Gly Asp Asn Trp Ser
Lys Tyr 500 505 510Gln Ser Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser
Thr Phe Val Pro 515 520 525Met Gly Phe Ala Gln Lys Asp Gly Ser Tyr
Ala Gly Phe Asp Ile Asp 530 535 540Leu Ala Thr Ala Val Phe Glu Lys
Tyr Gly Ile Thr Val Asn Trp Gln545 550 555 560Pro Ile Asp Trp Asp
Leu Lys Glu Ala Glu Leu Thr Lys Gly Thr Ile 565 570 575Asp Leu Ile
Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys 580 585 590Val
Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val Leu Val Thr 595 600
605Lys Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp Met Thr Gly Lys Thr
610 615 620Leu Gly Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp Phe Glu
Ala Asn625 630 635 640Pro Glu Ile Leu Lys Asn Ile Val Ala Asn Lys
Glu Ala Asn Gln Tyr 645 650 655Gln Thr Phe Asn Glu Ala Leu Ile Asp
Leu Lys Asn Asp Arg Ile Asp 660 665 670Gly Leu Leu Ile Asp Arg Val
Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu 675 680 685Gly Val Leu Asn Asp
Tyr Asn Val Phe Thr Val Gly Leu Glu Thr Glu 690 695 700Ala Phe Ala
Val Gly Ala Arg Lys Glu Asp Thr Asn Leu Val Lys Lys705 710 715
720Ile Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys Phe Gln Glu
725 730 735Ile Ser Gln Lys Trp Phe Gly Glu Asp Val Ala Thr Lys Glu
Val Lys 740 745 750Glu Gly Gln 75521755PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
21Met Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln Ser Asn Lys Ser Ile1
5 10 15Thr Ile Gly Phe Asp Ser Thr Phe Val Pro Met Gly Phe Ala Gln
Lys 20 25 30Asp Gly Ser Tyr Ala Gly Phe Asp Ile Asp Leu Ala Thr Ala
Val Phe 35 40 45Glu Lys Tyr Gly Ile Thr Val Asn Trp Gln Pro Ile Asp
Trp Asp Leu 50 55 60Lys Glu Ala Glu Leu Thr Lys Gly Thr Ile Asp Leu
Ile Trp Asn Gly65 70 75 80Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys
Val Ala Phe Ser Asn Ser 85 90 95Tyr Met Lys Asn Glu Gln Val Leu Val
Thr Lys Lys Ser Ser Gly Ile 100 105 110Thr Thr Ala Lys Asp Met Thr
Gly Lys Thr Leu Gly Ala Gln Ala Gly 115 120 125Ser Ser Gly Tyr Ala
Asp Phe Glu Ala Asn Pro Glu Ile Leu Lys Asn 130 135 140Ile Val Ala
Asn Lys Glu Ala Asn Gln Tyr Gln Thr Phe Asn Glu Ala145 150 155
160Leu Ile Asp Leu Lys Asn Asp Arg Ile Asp Gly Leu Leu Ile Asp Arg
165 170 175Val Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu Gly Val Leu Asn
Asp Tyr 180 185 190Asn Val Phe Thr Val Gly Leu Glu Thr Glu Ala Phe
Ala Val Gly Ala 195 200 205Arg Lys Glu Asp Thr Asn Leu Val Lys Lys
Ile Asn Glu Ala Phe Ser 210 215 220Ser Leu Tyr Lys Asp Gly Lys Phe
Gln Glu Ile Ser Gln Lys Trp Phe225 230 235 240Gly Glu Asp Val Ala
Thr Lys Glu Val Lys Glu Gly Gln Phe Arg Gln 245 250 255Pro Ser Gln
Thr Ala Leu Lys Asp Glu Pro Thr His Leu Val Val Ala 260 265 270Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly Thr Val Thr 275 280
285Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys Gly Asp Leu
290 295 300Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu Gly
Gln Ala305 310 315 320Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala
Ala Tyr Asp Ser Ala 325 330 335Ser Arg Ala Val Ala Arg Ala Asp Arg
His Ile Asn Glu Leu Asn Gln 340 345 350Ala Arg Asn Glu Ala Ala Ser
Ala Pro Ala Pro Gln Leu Pro Ala Pro 355 360 365Val Gly Gly Glu Asp
Ala Thr Val Gln Ser Pro Thr Pro Val Ala Gly 370 375 380Asn Ser Val
Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala Arg Asp Ala385 390 395
400Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser Gln Leu Asp
405 410 415Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val Glu
Val Asn 420 425 430Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln
Val Met Val His 435 440 445Ile Val Ser Asn Glu Asn Leu Gln Val Lys
Gly Glu Leu Ser Glu Tyr 450 455 460Asn Leu Ala Asn Leu Ser Val Gly
Gln Glu Val Ser Phe Thr Ser Lys465 470 475 480Val Tyr Pro Asp Lys
Lys Trp Thr Gly Lys Leu Ser Tyr Ile Ser Asp 485 490 495Tyr Pro Lys
Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala Gly Asn Asn 500 505 510Thr
Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly Glu Val Gly 515 520
525Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys Ser Lys Thr
530 535 540Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp Asp
Ser Lys545 550 555 560Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys
Ala Lys Lys Val Glu 565 570 575Val Ser Leu Gly Asn Ala Asp Ala Glu
Asn Gln Glu Ile Thr Ser Gly 580 585 590Leu Thr Asn Gly Ala Lys Val
Ile Ser Asn Pro Thr Ser Ser Leu Glu 595 600 605Glu Gly Lys Glu Val
Lys Ala Asp Glu Ala Thr Asn Phe Asp Ala Ser 610 615 620Asn Phe Lys
Asp Phe Ser Ser Ile Ala Ser Ala Ser Ser Ser Trp Gln625 630 635
640Asn Gln Ser Gly Ser Thr Met Ile Ile Gln Val Asp Ser Phe Gly Asn
645 650 655Val Ser Gly Gln Tyr Val Asn Arg Ala Gln Gly Thr Gly Cys
Gln Asn 660 665 670Ser Pro Tyr Pro Leu Thr Gly Arg Val Asn Gly Thr
Phe Ile Ala Phe 675 680 685Ser Val Gly Trp Asn Asn Ser Thr Glu Asn
Cys Asn Ser Ala Thr Gly 690 695 700Trp Thr Gly Tyr Ala Gln Val Asn
Gly Asn Asn Thr Glu Ile Val Thr705 710 715 720Ser Trp Asn Leu Ala
Tyr Glu Gly Gly Ser Gly Pro Ala Ile Glu Gln 725 730 735Gly Gln Asp
Thr Phe Gln Tyr Val Pro Thr Thr Glu Asn Lys Ser Leu 740 745 750Leu
Lys Asp 75522755PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 22Met Phe Arg Gln Pro Ser Gln Thr
Ala Leu Lys Asp Glu Pro Thr His1 5 10 15Leu Val Val Ala Lys Glu Gly
Ser Val Ala Ser Ser Val Leu Leu Ser 20 25 30Gly Thr Val Thr Ala Lys
Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser 35 40 45Lys Gly Asp Leu Asp
Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser 50 55 60Glu Gly Gln Ala
Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala65 70 75 80Tyr Asp
Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn 85 90 95Glu
Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln 100 105
110Leu Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr
115 120 125Pro Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu
Gly Asp 130 135 140Ala Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu
Ser Lys Ala Gln145 150 155 160Ser Gln Leu Asp Ala Thr Thr Val Leu
Ser Thr Leu Glu Gly Thr Val 165 170 175Val Glu Val Asn Ser Asn Val
Ser Lys Ser Pro Thr Gly Ala Ser Gln 180 185 190Val Met Val His Ile
Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu 195 200 205Leu Ser Glu
Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser 210 215 220Phe
Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser225 230
235 240Tyr Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro
Ala 245 250 255Ala Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile
Asp Val Thr 260 265 270Gly Glu Val Gly Asp Leu Lys Gln Gly Phe Ser
Val Asn Ile Glu Val 275 280 285Lys Ser Lys Thr Lys Ala Ile Leu Val
Pro Val Ser Ser Leu Val Met 290 295 300Asp Asp Ser Lys Asn Tyr Val
Trp Ile Val Asp Glu Gln Gln Lys Ala305 310 315 320Lys Lys Val Glu
Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu 325 330 335Ile Thr
Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr 340 345
350Ser Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn
355 360 365Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln Ser Asn Lys Ser
Ile Thr 370 375 380Ile Gly Phe Asp Ser Thr Phe Val Pro Met Gly Phe
Ala Gln Lys
Asp385 390 395 400Gly Ser Tyr Ala Gly Phe Asp Ile Asp Leu Ala Thr
Ala Val Phe Glu 405 410 415Lys Tyr Gly Ile Thr Val Asn Trp Gln Pro
Ile Asp Trp Asp Leu Lys 420 425 430Glu Ala Glu Leu Thr Lys Gly Thr
Ile Asp Leu Ile Trp Asn Gly Tyr 435 440 445Ser Ala Thr Asp Glu Arg
Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr 450 455 460Met Lys Asn Glu
Gln Val Leu Val Thr Lys Lys Ser Ser Gly Ile Thr465 470 475 480Thr
Ala Lys Asp Met Thr Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser 485 490
495Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro Glu Ile Leu Lys Asn Ile
500 505 510Val Ala Asn Lys Glu Ala Asn Gln Tyr Gln Thr Phe Asn Glu
Ala Leu 515 520 525Ile Asp Leu Lys Asn Asp Arg Ile Asp Gly Leu Leu
Ile Asp Arg Val 530 535 540Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu Gly
Val Leu Asn Asp Tyr Asn545 550 555 560Val Phe Thr Val Gly Leu Glu
Thr Glu Ala Phe Ala Val Gly Ala Arg 565 570 575Lys Glu Asp Thr Asn
Leu Val Lys Lys Ile Asn Glu Ala Phe Ser Ser 580 585 590Leu Tyr Lys
Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly 595 600 605Glu
Asp Val Ala Thr Lys Glu Val Lys Glu Gly Gln Phe Asp Ala Ser 610 615
620Asn Phe Lys Asp Phe Ser Ser Ile Ala Ser Ala Ser Ser Ser Trp
Gln625 630 635 640Asn Gln Ser Gly Ser Thr Met Ile Ile Gln Val Asp
Ser Phe Gly Asn 645 650 655Val Ser Gly Gln Tyr Val Asn Arg Ala Gln
Gly Thr Gly Cys Gln Asn 660 665 670Ser Pro Tyr Pro Leu Thr Gly Arg
Val Asn Gly Thr Phe Ile Ala Phe 675 680 685Ser Val Gly Trp Asn Asn
Ser Thr Glu Asn Cys Asn Ser Ala Thr Gly 690 695 700Trp Thr Gly Tyr
Ala Gln Val Asn Gly Asn Asn Thr Glu Ile Val Thr705 710 715 720Ser
Trp Asn Leu Ala Tyr Glu Gly Gly Ser Gly Pro Ala Ile Glu Gln 725 730
735Gly Gln Asp Thr Phe Gln Tyr Val Pro Thr Thr Glu Asn Lys Ser Leu
740 745 750Leu Lys Asp 75523765PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 23Met Phe Asp Ala Ser Asn
Phe Lys Asp Phe Ser Ser Ile Ala Ser Ala1 5 10 15Ser Ser Ser Trp Gln
Asn Gln Ser Gly Ser Thr Met Ile Ile Gln Val 20 25 30Asp Ser Phe Gly
Asn Val Ser Gly Gln Tyr Val Asn Arg Ala Gln Gly 35 40 45Thr Gly Cys
Gln Asn Ser Pro Tyr Pro Leu Thr Gly Arg Val Asn Gly 50 55 60Thr Phe
Ile Ala Phe Ser Val Gly Trp Asn Asn Ser Thr Glu Asn Cys65 70 75
80Asn Ser Ala Thr Gly Trp Thr Gly Tyr Ala Gln Val Asn Gly Asn Asn
85 90 95Thr Glu Ile Val Thr Ser Trp Asn Leu Ala Tyr Glu Gly Gly Ser
Gly 100 105 110Pro Ala Ile Glu Gln Gly Gln Asp Thr Phe Gln Tyr Val
Pro Thr Thr 115 120 125Glu Asn Lys Ser Leu Leu Lys Asp Gly Gly Gly
Gly Ser Ser Ser Thr 130 135 140Ser Gly Asp Asn Trp Ser Lys Tyr Gln
Ser Asn Lys Ser Ile Thr Ile145 150 155 160Gly Phe Asp Ser Thr Phe
Val Pro Met Gly Phe Ala Gln Lys Asp Gly 165 170 175Ser Tyr Ala Gly
Phe Asp Ile Asp Leu Ala Thr Ala Val Phe Glu Lys 180 185 190Tyr Gly
Ile Thr Val Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu 195 200
205Ala Glu Leu Thr Lys Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser
210 215 220Ala Thr Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser
Tyr Met225 230 235 240Lys Asn Glu Gln Val Leu Val Thr Lys Lys Ser
Ser Gly Ile Thr Thr 245 250 255Ala Lys Asp Met Thr Gly Lys Thr Leu
Gly Ala Gln Ala Gly Ser Ser 260 265 270Gly Tyr Ala Asp Phe Glu Ala
Asn Pro Glu Ile Leu Lys Asn Ile Val 275 280 285Ala Asn Lys Glu Ala
Asn Gln Tyr Gln Thr Phe Asn Glu Ala Leu Ile 290 295 300Asp Leu Lys
Asn Asp Arg Ile Asp Gly Leu Leu Ile Asp Arg Val Tyr305 310 315
320Ala Asn Tyr Tyr Leu Glu Ala Glu Gly Val Leu Asn Asp Tyr Asn Val
325 330 335Phe Thr Val Gly Leu Glu Thr Glu Ala Phe Ala Val Gly Ala
Arg Lys 340 345 350Glu Asp Thr Asn Leu Val Lys Lys Ile Asn Glu Ala
Phe Ser Ser Leu 355 360 365Tyr Lys Asp Gly Lys Phe Gln Glu Ile Ser
Gln Lys Trp Phe Gly Glu 370 375 380Asp Val Ala Thr Lys Glu Val Lys
Glu Gly Gln Ala Ala Ala Phe Arg385 390 395 400Gln Pro Ser Gln Thr
Ala Leu Lys Asp Glu Pro Thr His Leu Val Val 405 410 415Ala Lys Glu
Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly Thr Val 420 425 430Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys Gly Asp 435 440
445Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu Gly Gln
450 455 460Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
Asp Ser465 470 475 480Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His
Ile Asn Glu Leu Asn 485 490 495Gln Ala Arg Asn Glu Ala Ala Ser Ala
Pro Ala Pro Gln Leu Pro Ala 500 505 510Pro Val Gly Gly Glu Asp Ala
Thr Val Gln Ser Pro Thr Pro Val Ala 515 520 525Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala Arg Asp 530 535 540Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser Gln Leu545 550 555
560Asp Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val Glu Val
565 570 575Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val
Met Val 580 585 590His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly
Glu Leu Ser Glu 595 600 605Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe Thr Ser 610 615 620Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr Ile Ser625 630 635 640Asp Tyr Pro Lys Asn
Asn Gly Glu Ala Ala Ser Pro Ala Ala Gly Asn 645 650 655Asn Thr Gly
Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly Glu Val 660 665 670Gly
Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys Ser Lys 675 680
685Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp Asp Ser
690 695 700Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
Lys Val705 710 715 720Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn
Gln Glu Ile Thr Ser 725 730 735Gly Leu Thr Asn Gly Ala Lys Val Ile
Ser Asn Pro Thr Ser Ser Leu 740 745 750Glu Glu Gly Lys Glu Val Lys
Ala Asp Glu Ala Thr Asn 755 760 76524765PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
24Met Phe Asp Ala Ser Asn Phe Lys Asp Phe Ser Ser Ile Ala Ser Ala1
5 10 15Ser Ser Ser Trp Gln Asn Gln Ser Gly Ser Thr Met Ile Ile Gln
Val 20 25 30Asp Ser Phe Gly Asn Val Ser Gly Gln Tyr Val Asn Arg Ala
Gln Gly 35 40 45Thr Gly Cys Gln Asn Ser Pro Tyr Pro Leu Thr Gly Arg
Val Asn Gly 50 55 60Thr Phe Ile Ala Phe Ser Val Gly Trp Asn Asn Ser
Thr Glu Asn Cys65 70 75 80Asn Ser Ala Thr Gly Trp Thr Gly Tyr Ala
Gln Val Asn Gly Asn Asn 85 90 95Thr Glu Ile Val Thr Ser Trp Asn Leu
Ala Tyr Glu Gly Gly Ser Gly 100 105 110Pro Ala Ile Glu Gln Gly Gln
Asp Thr Phe Gln Tyr Val Pro Thr Thr 115 120 125Glu Asn Lys Ser Leu
Leu Lys Asp Gly Gly Gly Gly Ser Ser Ser Phe 130 135 140Arg Gln Pro
Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu Val145 150 155
160Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly Thr
165 170 175Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser
Lys Gly 180 185 190Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys
Val Ser Glu Gly 195 200 205Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu
Ala Gln Ala Ala Tyr Asp 210 215 220Ser Ala Ser Arg Ala Val Ala Arg
Ala Asp Arg His Ile Asn Glu Leu225 230 235 240Asn Gln Ala Arg Asn
Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu Pro 245 250 255Ala Pro Val
Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro Val 260 265 270Ala
Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala Arg 275 280
285Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser Gln
290 295 300Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val
Val Glu305 310 315 320Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly
Ala Ser Gln Val Met 325 330 335Val His Ile Val Ser Asn Glu Asn Leu
Gln Val Lys Gly Glu Leu Ser 340 345 350Glu Tyr Asn Leu Ala Asn Leu
Ser Val Gly Gln Glu Val Ser Phe Thr 355 360 365Ser Lys Val Tyr Pro
Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr Ile 370 375 380Ser Asp Tyr
Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala Gly385 390 395
400Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly Glu
405 410 415Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val
Lys Ser 420 425 430Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu
Val Met Asp Asp 435 440 445Ser Lys Asn Tyr Val Trp Ile Val Asp Glu
Gln Gln Lys Ala Lys Lys 450 455 460Val Glu Val Ser Leu Gly Asn Ala
Asp Ala Glu Asn Gln Glu Ile Thr465 470 475 480Ser Gly Leu Thr Asn
Gly Ala Lys Val Ile Ser Asn Pro Thr Ser Ser 485 490 495Leu Glu Glu
Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn Ala Ala 500 505 510Ala
Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln Ser Asn Lys Ser Ile 515 520
525Thr Ile Gly Phe Asp Ser Thr Phe Val Pro Met Gly Phe Ala Gln Lys
530 535 540Asp Gly Ser Tyr Ala Gly Phe Asp Ile Asp Leu Ala Thr Ala
Val Phe545 550 555 560Glu Lys Tyr Gly Ile Thr Val Asn Trp Gln Pro
Ile Asp Trp Asp Leu 565 570 575Lys Glu Ala Glu Leu Thr Lys Gly Thr
Ile Asp Leu Ile Trp Asn Gly 580 585 590Tyr Ser Ala Thr Asp Glu Arg
Arg Glu Lys Val Ala Phe Ser Asn Ser 595 600 605Tyr Met Lys Asn Glu
Gln Val Leu Val Thr Lys Lys Ser Ser Gly Ile 610 615 620Thr Thr Ala
Lys Asp Met Thr Gly Lys Thr Leu Gly Ala Gln Ala Gly625 630 635
640Ser Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro Glu Ile Leu Lys Asn
645 650 655Ile Val Ala Asn Lys Glu Ala Asn Gln Tyr Gln Thr Phe Asn
Glu Ala 660 665 670Leu Ile Asp Leu Lys Asn Asp Arg Ile Asp Gly Leu
Leu Ile Asp Arg 675 680 685Val Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu
Gly Val Leu Asn Asp Tyr 690 695 700Asn Val Phe Thr Val Gly Leu Glu
Thr Glu Ala Phe Ala Val Gly Ala705 710 715 720Arg Lys Glu Asp Thr
Asn Leu Val Lys Lys Ile Asn Glu Ala Phe Ser 725 730 735Ser Leu Tyr
Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp Phe 740 745 750Gly
Glu Asp Val Ala Thr Lys Glu Val Lys Glu Gly Gln 755 760
76525765PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 25Met Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln
Ser Asn Lys Ser Ile1 5 10 15Thr Ile Gly Phe Asp Ser Thr Phe Val Pro
Met Gly Phe Ala Gln Lys 20 25 30Asp Gly Ser Tyr Ala Gly Phe Asp Ile
Asp Leu Ala Thr Ala Val Phe 35 40 45Glu Lys Tyr Gly Ile Thr Val Asn
Trp Gln Pro Ile Asp Trp Asp Leu 50 55 60Lys Glu Ala Glu Leu Thr Lys
Gly Thr Ile Asp Leu Ile Trp Asn Gly65 70 75 80Tyr Ser Ala Thr Asp
Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser 85 90 95Tyr Met Lys Asn
Glu Gln Val Leu Val Thr Lys Lys Ser Ser Gly Ile 100 105 110Thr Thr
Ala Lys Asp Met Thr Gly Lys Thr Leu Gly Ala Gln Ala Gly 115 120
125Ser Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro Glu Ile Leu Lys Asn
130 135 140Ile Val Ala Asn Lys Glu Ala Asn Gln Tyr Gln Thr Phe Asn
Glu Ala145 150 155 160Leu Ile Asp Leu Lys Asn Asp Arg Ile Asp Gly
Leu Leu Ile Asp Arg 165 170 175Val Tyr Ala Asn Tyr Tyr Leu Glu Ala
Glu Gly Val Leu Asn Asp Tyr 180 185 190Asn Val Phe Thr Val Gly Leu
Glu Thr Glu Ala Phe Ala Val Gly Ala 195 200 205Arg Lys Glu Asp Thr
Asn Leu Val Lys Lys Ile Asn Glu Ala Phe Ser 210 215 220Ser Leu Tyr
Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp Phe225 230 235
240Gly Glu Asp Val Ala Thr Lys Glu Val Lys Glu Gly Gln Gly Gly Gly
245 250 255Gly Ser Ser Ser Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys
Asp Glu 260 265 270Pro Thr His Leu Val Val Ala Lys Glu Gly Ser Val
Ala Ser Ser Val 275 280 285Leu Leu Ser Gly Thr Val Thr Ala Lys Asn
Glu Gln Tyr Val Tyr Phe 290 295 300Asp Ala Ser Lys Gly Asp Leu Asp
Glu Ile Leu Val Ser Val Gly Asp305 310 315 320Lys Val Ser Glu Gly
Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala 325 330 335Gln Ala Ala
Tyr Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg 340 345 350His
Ile Asn Glu Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro 355 360
365Ala Pro Gln Leu Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln
370 375 380Ser Pro Thr Pro Val Ala Gly Asn Ser Val Ala Ser Ile Asp
Ala Gln385 390 395 400Leu Gly Asp Ala Arg Asp Ala Arg Ala Asp Ala
Ala Ala Gln Leu Ser 405 410 415Lys Ala Gln Ser Gln Leu Asp Ala Thr
Thr Val Leu Ser Thr Leu Glu 420 425 430Gly Thr Val Val Glu Val Asn
Ser Asn Val Ser Lys Ser Pro Thr Gly 435 440 445Ala Ser Gln Val Met
Val His Ile Val Ser Asn Glu Asn Leu Gln Val 450 455 460Lys Gly Glu
Leu Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln465 470 475
480Glu Val Ser Phe Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly
485 490 495Lys Leu Ser Tyr Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu
Ala Ala 500 505 510Ser Pro Ala Ala Gly Asn Asn Thr Gly Ser Lys Tyr
Pro Tyr Thr Ile 515 520 525Asp Val Thr Gly Glu Val Gly Asp Leu Lys
Gln Gly Phe Ser Val Asn 530
535 540Ile Glu Val Lys Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser545 550 555 560Leu Val Met Asp Asp Ser Lys Asn Tyr Val Trp Ile
Val Asp Glu Gln 565 570 575Gln Lys Ala Lys Lys Val Glu Val Ser Leu
Gly Asn Ala Asp Ala Glu 580 585 590Asn Gln Glu Ile Thr Ser Gly Leu
Thr Asn Gly Ala Lys Val Ile Ser 595 600 605Asn Pro Thr Ser Ser Leu
Glu Glu Gly Lys Glu Val Lys Ala Asp Glu 610 615 620Ala Thr Asn Ala
Ala Ala Phe Asp Ala Ser Asn Phe Lys Asp Phe Ser625 630 635 640Ser
Ile Ala Ser Ala Ser Ser Ser Trp Gln Asn Gln Ser Gly Ser Thr 645 650
655Met Ile Ile Gln Val Asp Ser Phe Gly Asn Val Ser Gly Gln Tyr Val
660 665 670Asn Arg Ala Gln Gly Thr Gly Cys Gln Asn Ser Pro Tyr Pro
Leu Thr 675 680 685Gly Arg Val Asn Gly Thr Phe Ile Ala Phe Ser Val
Gly Trp Asn Asn 690 695 700Ser Thr Glu Asn Cys Asn Ser Ala Thr Gly
Trp Thr Gly Tyr Ala Gln705 710 715 720Val Asn Gly Asn Asn Thr Glu
Ile Val Thr Ser Trp Asn Leu Ala Tyr 725 730 735Glu Gly Gly Ser Gly
Pro Ala Ile Glu Gln Gly Gln Asp Thr Phe Gln 740 745 750Tyr Val Pro
Thr Thr Glu Asn Lys Ser Leu Leu Lys Asp 755 760
76526764PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 26Met Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys
Asp Glu Pro Thr His1 5 10 15Leu Val Val Ala Lys Glu Gly Ser Val Ala
Ser Ser Val Leu Leu Ser 20 25 30Gly Thr Val Thr Ala Lys Asn Glu Gln
Tyr Val Tyr Phe Asp Ala Ser 35 40 45Lys Gly Asp Leu Asp Glu Ile Leu
Val Ser Val Gly Asp Lys Val Ser 50 55 60Glu Gly Gln Ala Leu Val Lys
Tyr Ser Ser Ser Glu Ala Gln Ala Ala65 70 75 80Tyr Asp Ser Ala Ser
Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn 85 90 95Glu Leu Asn Gln
Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln 100 105 110Leu Pro
Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr 115 120
125Pro Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp
130 135 140Ala Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys
Ala Gln145 150 155 160Ser Gln Leu Asp Ala Thr Thr Val Leu Ser Thr
Leu Glu Gly Thr Val 165 170 175Val Glu Val Asn Ser Asn Val Ser Lys
Ser Pro Thr Gly Ala Ser Gln 180 185 190Val Met Val His Ile Val Ser
Asn Glu Asn Leu Gln Val Lys Gly Glu 195 200 205Leu Ser Glu Tyr Asn
Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser 210 215 220Phe Thr Ser
Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser225 230 235
240Tyr Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala
245 250 255Ala Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp
Val Thr 260 265 270Gly Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val
Asn Ile Glu Val 275 280 285Lys Ser Lys Thr Lys Ala Ile Leu Val Pro
Val Ser Ser Leu Val Met 290 295 300Asp Asp Ser Lys Asn Tyr Val Trp
Ile Val Asp Glu Gln Gln Lys Ala305 310 315 320Lys Lys Val Glu Val
Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu 325 330 335Ile Thr Ser
Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr 340 345 350Ser
Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn 355 360
365Gly Gly Gly Gly Ser Ser Ser Thr Ser Gly Asp Asn Trp Ser Lys Tyr
370 375 380Gln Ser Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr Phe
Val Pro385 390 395 400Met Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala
Gly Phe Asp Ile Asp 405 410 415Leu Ala Thr Ala Val Phe Glu Lys Tyr
Gly Ile Thr Val Asn Trp Gln 420 425 430Pro Ile Asp Trp Asp Leu Lys
Glu Ala Glu Leu Thr Lys Gly Thr Ile 435 440 445Asp Leu Ile Trp Asn
Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys 450 455 460Val Ala Phe
Ser Asn Ser Tyr Met Asn Glu Gln Val Leu Val Thr Lys465 470 475
480Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp Met Thr Gly Lys Thr Leu
485 490 495Gly Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp Phe Glu Ala
Asn Pro 500 505 510Glu Ile Leu Lys Asn Ile Val Ala Asn Lys Glu Ala
Asn Gln Tyr Gln 515 520 525Thr Phe Asn Glu Ala Leu Ile Asp Leu Lys
Asn Asp Arg Ile Asp Gly 530 535 540Leu Leu Ile Asp Arg Val Tyr Ala
Asn Tyr Tyr Leu Glu Ala Glu Gly545 550 555 560Val Leu Asn Asp Tyr
Asn Val Phe Thr Val Gly Leu Glu Thr Glu Ala 565 570 575Phe Ala Val
Gly Ala Arg Lys Glu Asp Thr Asn Leu Val Lys Lys Ile 580 585 590Asn
Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys Phe Gln Glu Ile 595 600
605Ser Gln Lys Trp Phe Gly Glu Asp Val Ala Thr Lys Glu Val Lys Glu
610 615 620Gly Gln Ala Ala Ala Phe Asp Ala Ser Asn Phe Lys Asp Phe
Ser Ser625 630 635 640Ile Ala Ser Ala Ser Ser Ser Trp Gln Asn Gln
Ser Gly Ser Thr Met 645 650 655Ile Ile Gln Val Asp Ser Phe Gly Asn
Val Ser Gly Gln Tyr Val Asn 660 665 670Arg Ala Gln Gly Thr Gly Cys
Gln Asn Ser Pro Tyr Pro Leu Thr Gly 675 680 685Arg Val Asn Gly Thr
Phe Ile Ala Phe Ser Val Gly Trp Asn Asn Ser 690 695 700Thr Glu Asn
Cys Asn Ser Ala Thr Gly Trp Thr Gly Tyr Ala Gln Val705 710 715
720Asn Gly Asn Asn Thr Glu Ile Val Thr Ser Trp Asn Leu Ala Tyr Glu
725 730 735Gly Gly Ser Gly Pro Ala Ile Glu Gln Gly Gln Asp Thr Phe
Gln Tyr 740 745 750Val Pro Thr Thr Glu Asn Lys Ser Leu Leu Lys Asp
755 760271860DNAArtificial SequenceDescription of Artificial
Sequence Synthetic polynucleotide 27atgaccagcg gcgacaattg
gtccaaatac cagagcaaca agagcatcac gatcggcttc 60gacagcactt ttgtgccgat
gggtttcgcg caaaaagacg gtagctacgc gggtttcgat 120attgacctgg
cgaccgctgt ctttgagaaa tacggcatta cggttaattg gcagccgatt
180gattgggacc tgaaagaggc cgaactcacc aaaggcacca tcgacctgat
ctggaatggt 240tactccgcaa ccgatgagcg tcgcgaaaaa gttgccttca
gcaacagcta tatgaagaat 300gaacaagtgt tggtaaccaa gaaatctagc
ggcattacga ccgcgaaaga catgaccggt 360aagacgctgg gtgcgcaggc
cggtagctct ggctatgcgg atttcgaggc gaatcctgag 420attctgaaaa
acatcgttgc gaataaagag gcgaaccagt accagacctt taacgaagca
480ctgatcgacc tgaaaaacga tcgcattgac ggtctgctga tcgatcgtgt
gtacgcgaac 540tattatctgg aagccgaggg cgttctgaac gattataatg
tttttaccgt gggtctggag 600actgaggcat tcgcggttgg tgcgcgcaag
gaagatacca acctggttaa aaagattaat 660gaggcattta gctcactgta
caaggacggc aagttccaag aaattagcca gaagtggttc 720ggtgaagatg
ttgcgacgaa agaggttaaa gagggccaat ttcgccaacc gagccagact
780gcgttgaaag atgagccgac ccatctggtt gttgcgaaag agggcagcgt
ggcatcgagc 840gtgctgctga gcggtacggt tactgccaaa aacgaacaat
acgtgtactt cgatgctagc 900aagggtgatc tggatgaaat tctggtgagc
gtgggtgaca aagttagcga aggccaggca 960ctggtgaagt attcatcctc
cgaggcacag gcagcgtacg acagcgcaag ccgcgcagtg 1020gcgcgtgccg
accgtcacat taacgaattg aaccaagcgc gtaacgaggc cgcaagcgcg
1080ccagcaccgc agctgccggc tccggtgggt ggcgaagatg cgacggtgca
gagcccgacc 1140ccggttgcgg gtaattcggt cgccagcatc gatgcgcagc
tgggtgacgc gcgtgatgcc 1200cgtgcggatg cggctgctca actgagcaag
gctcagagcc aactggacgc gacgacggtg 1260ctgagcacct tggagggtac
cgttgtcgaa gtcaacagca atgtgagcaa gagcccaacg 1320ggtgcgagcc
aggttatggt ccacattgtg agcaatgaaa acttacaggt caagggtgag
1380ctgagcgagt ataacctggc gaatctgagc gttggtcaag aggtcagctt
taccagcaag 1440gtctacccgg ataagaaatg gaccggcaag ttgagctaca
tcagcgacta cccgaagaac 1500aatggcgagg cagcctcccc ggcagccggc
aacaataccg gctctaagta tccgtacacc 1560atcgacgtaa ccggtgaggt
cggcgacctg aaacagggtt ttagcgtgaa tatcgaagtg 1620aagtccaaga
ccaaggcaat tttggttccg gttagctccc tggtgatgga cgatagcaag
1680aattatgtgt ggattgtcga cgagcaacag aaagcgaaaa aagttgaagt
gagcctgggc 1740aatgctgatg ccgagaacca agaaatcacg tctggtctga
ccaacggtgc gaaagttatt 1800agcaacccga ccagcagcct ggaagagggt
aaagaggtca aagccgacga agctacgaac 1860281860DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
28atgtttcgcc aaccgagcca gactgcgttg aaagatgagc cgacccatct ggttgttgcg
60aaagagggca gcgtggcatc gagcgtgctg ctgagcggta cggttactgc caaaaacgaa
120caatacgtgt acttcgatgc tagcaagggt gatctggatg aaattctggt
gagcgtgggt 180gacaaagtta gcgaaggcca ggcactggtg aagtattcat
cctccgaggc acaggcagcg 240tacgacagcg caagccgcgc agtggcgcgt
gccgaccgtc acattaacga attgaaccaa 300gcgcgtaacg aggccgcaag
cgcgccagca ccgcagctgc cggctccggt gggtggcgaa 360gatgcgacgg
tgcagagccc gaccccggtt gcgggtaatt cggtcgccag catcgatgcg
420cagctgggtg acgcgcgtga tgcccgtgcg gatgcggctg ctcaactgag
caaggctcag 480agccaactgg acgcgacgac ggtgctgagc accttggagg
gtaccgttgt cgaagtcaac 540agcaatgtga gcaagagccc aacgggtgcg
agccaggtta tggtccacat tgtgagcaat 600gaaaacttac aggtcaaggg
tgagctgagc gagtataacc tggcgaatct gagcgttggt 660caagaggtca
gctttaccag caaggtctac ccggataaga aatggaccgg caagttgagc
720tacatcagcg actacccgaa gaacaatggc gaggcagcct ccccggcagc
cggcaacaat 780accggctcta agtatccgta caccatcgac gtaaccggtg
aggtcggcga cctgaaacag 840ggttttagcg tgaatatcga agtgaagtcc
aagaccaagg caattttggt tccggttagc 900tccctggtga tggacgatag
caagaattat gtgtggattg tcgacgagca acagaaagcg 960aaaaaagttg
aagtgagcct gggcaatgct gatgccgaga accaagaaat cacgtctggt
1020ctgaccaacg gtgcgaaagt tattagcaac ccgaccagca gcctggaaga
gggtaaagag 1080gtcaaagccg acgaagctac gaacaccagc ggcgacaatt
ggtccaaata ccagagcaac 1140aagagcatca cgatcggctt cgacagcact
tttgtgccga tgggtttcgc gcaaaaagac 1200ggtagctacg cgggtttcga
tattgacctg gcgaccgctg tctttgagaa atacggcatt 1260acggttaatt
ggcagccgat tgattgggac ctgaaagagg ccgaactcac caaaggcacc
1320atcgacctga tctggaatgg ttactccgca accgatgagc gtcgcgaaaa
agttgccttc 1380agcaacagct atatgaagaa tgaacaagtg ttggtaacca
agaaatctag cggcattacg 1440accgcgaaag acatgaccgg taagacgctg
ggtgcgcagg ccggtagctc tggctatgcg 1500gatttcgagg cgaatcctga
gattctgaaa aacatcgttg cgaataaaga ggcgaaccag 1560taccagacct
ttaacgaagc actgatcgac ctgaaaaacg atcgcattga cggtctgctg
1620atcgatcgtg tgtacgcgaa ctattatctg gaagccgagg gcgttctgaa
cgattataat 1680gtttttaccg tgggtctgga gactgaggca ttcgcggttg
gtgcgcgcaa ggaagatacc 1740aacctggtta aaaagattaa tgaggcattt
agctcactgt acaaggacgg caagttccaa 1800gaaattagcc agaagtggtt
cggtgaagat gttgcgacga aagaggttaa agagggccaa 1860292265DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
29atgttcgacg catccaactt taaagacttt agcagcatcg cgtccgcaag ctctagctgg
60cagaatcaat ctggtagcac catgattatc caagtggaca gctttggtaa cgtcagcggt
120caatatgtta atcgtgcaca gggtacgggt tgtcagaatt ctccgtaccc
gctgaccggt 180cgtgttaacg gcacgttcat cgctttcagc gtcggttgga
acaattctac tgaaaattgc 240aacagcgcga ccggttggac gggctatgca
caagtgaatg gcaataacac cgaaatcgtc 300acgtcctgga atctggcgta
tgagggtggc agcggtccgg ctattgaaca gggccaggat 360accttccaat
acgtccctac gaccgagaat aagtcccttc tgaaagacac cagcggcgac
420aattggtcca aataccagag caacaagagc atcacgatcg gcttcgacag
cacttttgtg 480ccgatgggtt tcgcgcaaaa agacggtagc tacgcgggtt
tcgatattga cctggcgacc 540gctgtctttg agaaatacgg cattacggtt
aattggcagc cgattgattg ggacctgaaa 600gaggccgaac tcaccaaagg
caccatcgac ctgatctgga atggttactc cgcaaccgat 660gagcgtcgcg
aaaaagttgc cttcagcaac agctatatga agaatgaaca agtgttggta
720accaagaaat ctagcggcat tacgaccgcg aaagacatga ccggtaagac
gctgggtgcg 780caggccggta gctctggcta tgcggatttc gaggcgaatc
ctgagattct gaaaaacatc 840gttgcgaata aagaggcgaa ccagtaccag
acctttaacg aagcactgat cgacctgaaa 900aacgatcgca ttgacggtct
gctgatcgat cgtgtgtacg cgaactatta tctggaagcc 960gagggcgttc
tgaacgatta taatgttttt accgtgggtc tggagactga ggcattcgcg
1020gttggtgcgc gcaaggaaga taccaacctg gttaaaaaga ttaatgaggc
atttagctca 1080ctgtacaagg acggcaagtt ccaagaaatt agccagaagt
ggttcggtga agatgttgcg 1140acgaaagagg ttaaagaggg ccaatttcgc
caaccgagcc agactgcgtt gaaagatgag 1200ccgacccatc tggttgttgc
gaaagagggc agcgtggcat cgagcgtgct gctgagcggt 1260acggttactg
ccaaaaacga acaatacgtg tacttcgatg ctagcaaggg tgatctggat
1320gaaattctgg tgagcgtggg tgacaaagtt agcgaaggcc aggcactggt
gaagtattca 1380tcctccgagg cacaggcagc gtacgacagc gcaagccgcg
cagtggcgcg tgccgaccgt 1440cacattaacg aattgaacca agcgcgtaac
gaggccgcaa gcgcgccagc accgcagctg 1500ccggctccgg tgggtggcga
agatgcgacg gtgcagagcc cgaccccggt tgcgggtaat 1560tcggtcgcca
gcatcgatgc gcagctgggt gacgcgcgtg atgcccgtgc ggatgcggct
1620gctcaactga gcaaggctca gagccaactg gacgcgacga cggtgctgag
caccttggag 1680ggtaccgttg tcgaagtcaa cagcaatgtg agcaagagcc
caacgggtgc gagccaggtt 1740atggtccaca ttgtgagcaa tgaaaactta
caggtcaagg gtgagctgag cgagtataac 1800ctggcgaatc tgagcgttgg
tcaagaggtc agctttacca gcaaggtcta cccggataag 1860aaatggaccg
gcaagttgag ctacatcagc gactacccga agaacaatgg cgaggcagcc
1920tccccggcag ccggcaacaa taccggctct aagtatccgt acaccatcga
cgtaaccggt 1980gaggtcggcg acctgaaaca gggttttagc gtgaatatcg
aagtgaagtc caagaccaag 2040gcaattttgg ttccggttag ctccctggtg
atggacgata gcaagaatta tgtgtggatt 2100gtcgacgagc aacagaaagc
gaaaaaagtt gaagtgagcc tgggcaatgc tgatgccgag 2160aaccaagaaa
tcacgtctgg tctgaccaac ggtgcgaaag ttattagcaa cccgaccagc
2220agcctggaag agggtaaaga ggtcaaagcc gacgaagcta cgaac
2265302265DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 30atgttcgacg catccaactt taaagacttt
agcagcatcg cgtccgcaag ctctagctgg 60cagaatcaat ctggtagcac catgattatc
caagtggaca gctttggtaa cgtcagcggt 120caatatgtta atcgtgcaca
gggtacgggt tgtcagaatt ctccgtaccc gctgaccggt 180cgtgttaacg
gcacgttcat cgctttcagc gtcggttgga acaattctac tgaaaattgc
240aacagcgcga ccggttggac gggctatgca caagtgaatg gcaataacac
cgaaatcgtc 300acgtcctgga atctggcgta tgagggtggc agcggtccgg
ctattgaaca gggccaggat 360accttccaat acgtccctac gaccgagaat
aagtcccttc tgaaagactt tcgccaaccg 420agccagactg cgttgaaaga
tgagccgacc catctggttg ttgcgaaaga gggcagcgtg 480gcatcgagcg
tgctgctgag cggtacggtt actgccaaaa acgaacaata cgtgtacttc
540gatgctagca agggtgatct ggatgaaatt ctggtgagcg tgggtgacaa
agttagcgaa 600ggccaggcac tggtgaagta ttcatcctcc gaggcacagg
cagcgtacga cagcgcaagc 660cgcgcagtgg cgcgtgccga ccgtcacatt
aacgaattga accaagcgcg taacgaggcc 720gcaagcgcgc cagcaccgca
gctgccggct ccggtgggtg gcgaagatgc gacggtgcag 780agcccgaccc
cggttgcggg taattcggtc gccagcatcg atgcgcagct gggtgacgcg
840cgtgatgccc gtgcggatgc ggctgctcaa ctgagcaagg ctcagagcca
actggacgcg 900acgacggtgc tgagcacctt ggagggtacc gttgtcgaag
tcaacagcaa tgtgagcaag 960agcccaacgg gtgcgagcca ggttatggtc
cacattgtga gcaatgaaaa cttacaggtc 1020aagggtgagc tgagcgagta
taacctggcg aatctgagcg ttggtcaaga ggtcagcttt 1080accagcaagg
tctacccgga taagaaatgg accggcaagt tgagctacat cagcgactac
1140ccgaagaaca atggcgaggc agcctccccg gcagccggca acaataccgg
ctctaagtat 1200ccgtacacca tcgacgtaac cggtgaggtc ggcgacctga
aacagggttt tagcgtgaat 1260atcgaagtga agtccaagac caaggcaatt
ttggttccgg ttagctccct ggtgatggac 1320gatagcaaga attatgtgtg
gattgtcgac gagcaacaga aagcgaaaaa agttgaagtg 1380agcctgggca
atgctgatgc cgagaaccaa gaaatcacgt ctggtctgac caacggtgcg
1440aaagttatta gcaacccgac cagcagcctg gaagagggta aagaggtcaa
agccgacgaa 1500gctacgaaca ccagcggcga caattggtcc aaataccaga
gcaacaagag catcacgatc 1560ggcttcgaca gcacttttgt gccgatgggt
ttcgcgcaaa aagacggtag ctacgcgggt 1620ttcgatattg acctggcgac
cgctgtcttt gagaaatacg gcattacggt taattggcag 1680ccgattgatt
gggacctgaa agaggccgaa ctcaccaaag gcaccatcga cctgatctgg
1740aatggttact ccgcaaccga tgagcgtcgc gaaaaagttg ccttcagcaa
cagctatatg 1800aagaatgaac aagtgttggt aaccaagaaa tctagcggca
ttacgaccgc gaaagacatg 1860accggtaaga cgctgggtgc gcaggccggt
agctctggct atgcggattt cgaggcgaat 1920cctgagattc tgaaaaacat
cgttgcgaat aaagaggcga accagtacca gacctttaac 1980gaagcactga
tcgacctgaa aaacgatcgc attgacggtc tgctgatcga tcgtgtgtac
2040gcgaactatt atctggaagc cgagggcgtt ctgaacgatt ataatgtttt
taccgtgggt 2100ctggagactg aggcattcgc ggttggtgcg cgcaaggaag
ataccaacct ggttaaaaag 2160attaatgagg catttagctc actgtacaag
gacggcaagt tccaagaaat tagccagaag 2220tggttcggtg aagatgttgc
gacgaaagag gttaaagagg gccaa 2265312265DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
31atgaccagcg gcgacaattg gtccaaatac cagagcaaca agagcatcac gatcggcttc
60gacagcactt ttgtgccgat gggtttcgcg caaaaagacg gtagctacgc gggtttcgat
120attgacctgg cgaccgctgt ctttgagaaa tacggcatta cggttaattg
gcagccgatt 180gattgggacc tgaaagaggc cgaactcacc aaaggcacca
tcgacctgat ctggaatggt
240tactccgcaa ccgatgagcg tcgcgaaaaa gttgccttca gcaacagcta
tatgaagaat 300gaacaagtgt tggtaaccaa gaaatctagc ggcattacga
ccgcgaaaga catgaccggt 360aagacgctgg gtgcgcaggc cggtagctct
ggctatgcgg atttcgaggc gaatcctgag 420attctgaaaa acatcgttgc
gaataaagag gcgaaccagt accagacctt taacgaagca 480ctgatcgacc
tgaaaaacga tcgcattgac ggtctgctga tcgatcgtgt gtacgcgaac
540tattatctgg aagccgaggg cgttctgaac gattataatg tttttaccgt
gggtctggag 600actgaggcat tcgcggttgg tgcgcgcaag gaagatacca
acctggttaa aaagattaat 660gaggcattta gctcactgta caaggacggc
aagttccaag aaattagcca gaagtggttc 720ggtgaagatg ttgcgacgaa
agaggttaaa gagggccaat ttcgccaacc gagccagact 780gcgttgaaag
atgagccgac ccatctggtt gttgcgaaag agggcagcgt ggcatcgagc
840gtgctgctga gcggtacggt tactgccaaa aacgaacaat acgtgtactt
cgatgctagc 900aagggtgatc tggatgaaat tctggtgagc gtgggtgaca
aagttagcga aggccaggca 960ctggtgaagt attcatcctc cgaggcacag
gcagcgtacg acagcgcaag ccgcgcagtg 1020gcgcgtgccg accgtcacat
taacgaattg aaccaagcgc gtaacgaggc cgcaagcgcg 1080ccagcaccgc
agctgccggc tccggtgggt ggcgaagatg cgacggtgca gagcccgacc
1140ccggttgcgg gtaattcggt cgccagcatc gatgcgcagc tgggtgacgc
gcgtgatgcc 1200cgtgcggatg cggctgctca actgagcaag gctcagagcc
aactggacgc gacgacggtg 1260ctgagcacct tggagggtac cgttgtcgaa
gtcaacagca atgtgagcaa gagcccaacg 1320ggtgcgagcc aggttatggt
ccacattgtg agcaatgaaa acttacaggt caagggtgag 1380ctgagcgagt
ataacctggc gaatctgagc gttggtcaag aggtcagctt taccagcaag
1440gtctacccgg ataagaaatg gaccggcaag ttgagctaca tcagcgacta
cccgaagaac 1500aatggcgagg cagcctcccc ggcagccggc aacaataccg
gctctaagta tccgtacacc 1560atcgacgtaa ccggtgaggt cggcgacctg
aaacagggtt ttagcgtgaa tatcgaagtg 1620aagtccaaga ccaaggcaat
tttggttccg gttagctccc tggtgatgga cgatagcaag 1680aattatgtgt
ggattgtcga cgagcaacag aaagcgaaaa aagttgaagt gagcctgggc
1740aatgctgatg ccgagaacca agaaatcacg tctggtctga ccaacggtgc
gaaagttatt 1800agcaacccga ccagcagcct ggaagagggt aaagaggtca
aagccgacga agctacgaac 1860ttcgacgcat ccaactttaa agactttagc
agcatcgcgt ccgcaagctc tagctggcag 1920aatcaatctg gtagcaccat
gattatccaa gtggacagct ttggtaacgt cagcggtcaa 1980tatgttaatc
gtgcacaggg tacgggttgt cagaattctc cgtacccgct gaccggtcgt
2040gttaacggca cgttcatcgc tttcagcgtc ggttggaaca attctactga
aaattgcaac 2100agcgcgaccg gttggacggg ctatgcacaa gtgaatggca
ataacaccga aatcgtcacg 2160tcctggaatc tggcgtatga gggtggcagc
ggtccggcta ttgaacaggg ccaggatacc 2220ttccaatacg tccctacgac
cgagaataag tcccttctga aagac 2265322265DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
32atgtttcgcc aaccgagcca gactgcgttg aaagatgagc cgacccatct ggttgttgcg
60aaagagggca gcgtggcatc gagcgtgctg ctgagcggta cggttactgc caaaaacgaa
120caatacgtgt acttcgatgc tagcaagggt gatctggatg aaattctggt
gagcgtgggt 180gacaaagtta gcgaaggcca ggcactggtg aagtattcat
cctccgaggc acaggcagcg 240tacgacagcg caagccgcgc agtggcgcgt
gccgaccgtc acattaacga attgaaccaa 300gcgcgtaacg aggccgcaag
cgcgccagca ccgcagctgc cggctccggt gggtggcgaa 360gatgcgacgg
tgcagagccc gaccccggtt gcgggtaatt cggtcgccag catcgatgcg
420cagctgggtg acgcgcgtga tgcccgtgcg gatgcggctg ctcaactgag
caaggctcag 480agccaactgg acgcgacgac ggtgctgagc accttggagg
gtaccgttgt cgaagtcaac 540agcaatgtga gcaagagccc aacgggtgcg
agccaggtta tggtccacat tgtgagcaat 600gaaaacttac aggtcaaggg
tgagctgagc gagtataacc tggcgaatct gagcgttggt 660caagaggtca
gctttaccag caaggtctac ccggataaga aatggaccgg caagttgagc
720tacatcagcg actacccgaa gaacaatggc gaggcagcct ccccggcagc
cggcaacaat 780accggctcta agtatccgta caccatcgac gtaaccggtg
aggtcggcga cctgaaacag 840ggttttagcg tgaatatcga agtgaagtcc
aagaccaagg caattttggt tccggttagc 900tccctggtga tggacgatag
caagaattat gtgtggattg tcgacgagca acagaaagcg 960aaaaaagttg
aagtgagcct gggcaatgct gatgccgaga accaagaaat cacgtctggt
1020ctgaccaacg gtgcgaaagt tattagcaac ccgaccagca gcctggaaga
gggtaaagag 1080gtcaaagccg acgaagctac gaacaccagc ggcgacaatt
ggtccaaata ccagagcaac 1140aagagcatca cgatcggctt cgacagcact
tttgtgccga tgggtttcgc gcaaaaagac 1200ggtagctacg cgggtttcga
tattgacctg gcgaccgctg tctttgagaa atacggcatt 1260acggttaatt
ggcagccgat tgattgggac ctgaaagagg ccgaactcac caaaggcacc
1320atcgacctga tctggaatgg ttactccgca accgatgagc gtcgcgaaaa
agttgccttc 1380agcaacagct atatgaagaa tgaacaagtg ttggtaacca
agaaatctag cggcattacg 1440accgcgaaag acatgaccgg taagacgctg
ggtgcgcagg ccggtagctc tggctatgcg 1500gatttcgagg cgaatcctga
gattctgaaa aacatcgttg cgaataaaga ggcgaaccag 1560taccagacct
ttaacgaagc actgatcgac ctgaaaaacg atcgcattga cggtctgctg
1620atcgatcgtg tgtacgcgaa ctattatctg gaagccgagg gcgttctgaa
cgattataat 1680gtttttaccg tgggtctgga gactgaggca ttcgcggttg
gtgcgcgcaa ggaagatacc 1740aacctggtta aaaagattaa tgaggcattt
agctcactgt acaaggacgg caagttccaa 1800gaaattagcc agaagtggtt
cggtgaagat gttgcgacga aagaggttaa agagggccaa 1860ttcgacgcat
ccaactttaa agactttagc agcatcgcgt ccgcaagctc tagctggcag
1920aatcaatctg gtagcaccat gattatccaa gtggacagct ttggtaacgt
cagcggtcaa 1980tatgttaatc gtgcacaggg tacgggttgt cagaattctc
cgtacccgct gaccggtcgt 2040gttaacggca cgttcatcgc tttcagcgtc
ggttggaaca attctactga aaattgcaac 2100agcgcgaccg gttggacggg
ctatgcacaa gtgaatggca ataacaccga aatcgtcacg 2160tcctggaatc
tggcgtatga gggtggcagc ggtccggcta ttgaacaggg ccaggatacc
2220ttccaatacg tccctacgac cgagaataag tcccttctga aagac
2265332295DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 33atgttcgacg catccaactt taaagacttt
agcagcatcg cgtccgcaag ctctagctgg 60cagaatcaat ctggtagcac catgattatc
caagtggaca gctttggtaa cgtcagcggt 120caatatgtta atcgtgcaca
gggtacgggt tgtcagaatt ctccgtaccc gctgaccggt 180cgtgttaacg
gcacgttcat cgctttcagc gtcggttgga acaattctac tgaaaattgc
240aacagcgcga ccggttggac gggctatgca caagtgaatg gcaataacac
cgaaatcgtc 300acgtcctgga atctggcgta tgagggtggc agcggtccgg
ctattgaaca gggccaggat 360accttccaat acgtccctac gaccgagaat
aagtcccttc tgaaagacgg cggtggcggt 420tcgagctcga ccagcggcga
caattggtcc aaataccaga gcaacaagag catcacgatc 480ggcttcgaca
gcacttttgt gccgatgggt ttcgcgcaaa aagacggtag ctacgcgggt
540ttcgatattg acctggcgac cgctgtcttt gagaaatacg gcattacggt
taattggcag 600ccgattgatt gggacctgaa agaggccgaa ctcaccaaag
gcaccatcga cctgatctgg 660aatggttact ccgcaaccga tgagcgtcgc
gaaaaagttg ccttcagcaa cagctatatg 720aagaatgaac aagtgttggt
aaccaagaaa tctagcggca ttacgaccgc gaaagacatg 780accggtaaga
cgctgggtgc gcaggccggt agctctggct atgcggattt cgaggcgaat
840cctgagattc tgaaaaacat cgttgcgaat aaagaggcga accagtacca
gacctttaac 900gaagcactga tcgacctgaa aaacgatcgc attgacggtc
tgctgatcga tcgtgtgtac 960gcgaactatt atctggaagc cgagggcgtt
ctgaacgatt ataatgtttt taccgtgggt 1020ctggagactg aggcattcgc
ggttggtgcg cgcaaggaag ataccaacct ggttaaaaag 1080attaatgagg
catttagctc actgtacaag gacggcaagt tccaagaaat tagccagaag
1140tggttcggtg aagatgttgc gacgaaagag gttaaagagg gccaagcggc
cgcatttcgc 1200caaccgagcc agactgcgtt gaaagatgag ccgacccatc
tggttgttgc gaaagagggc 1260agcgtggcat cgagcgtgct gctgagcggt
acggttactg ccaaaaacga acaatacgtg 1320tacttcgatg ctagcaaggg
tgatctggat gaaattctgg tgagcgtggg tgacaaagtt 1380agcgaaggcc
aggcactggt gaagtattca tcctccgagg cacaggcagc gtacgacagc
1440gcaagccgcg cagtggcgcg tgccgaccgt cacattaacg aattgaacca
agcgcgtaac 1500gaggccgcaa gcgcgccagc accgcagctg ccggctccgg
tgggtggcga agatgcgacg 1560gtgcagagcc cgaccccggt tgcgggtaat
tcggtcgcca gcatcgatgc gcagctgggt 1620gacgcgcgtg atgcccgtgc
ggatgcggct gctcaactga gcaaggctca gagccaactg 1680gacgcgacga
cggtgctgag caccttggag ggtaccgttg tcgaagtcaa cagcaatgtg
1740agcaagagcc caacgggtgc gagccaggtt atggtccaca ttgtgagcaa
tgaaaactta 1800caggtcaagg gtgagctgag cgagtataac ctggcgaatc
tgagcgttgg tcaagaggtc 1860agctttacca gcaaggtcta cccggataag
aaatggaccg gcaagttgag ctacatcagc 1920gactacccga agaacaatgg
cgaggcagcc tccccggcag ccggcaacaa taccggctct 1980aagtatccgt
acaccatcga cgtaaccggt gaggtcggcg acctgaaaca gggttttagc
2040gtgaatatcg aagtgaagtc caagaccaag gcaattttgg ttccggttag
ctccctggtg 2100atggacgata gcaagaatta tgtgtggatt gtcgacgagc
aacagaaagc gaaaaaagtt 2160gaagtgagcc tgggcaatgc tgatgccgag
aaccaagaaa tcacgtctgg tctgaccaac 2220ggtgcgaaag ttattagcaa
cccgaccagc agcctggaag agggtaaaga ggtcaaagcc 2280gacgaagcta cgaac
2295342294DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 34atgttcgacg catccaactt taaagacttt
agcagcatcg cgtccgcaag ctctagctgg 60cagaatcaat ctggtagcac catgattatc
caagtggaca gctttggtaa cgtcagcggt 120caatatgtta atcgtgcaca
gggtacgggt tgtcagaatt ctccgtaccc gctgaccggt 180cgtgttaacg
gcacgttcat cgctttcagc gtcggttgga acaattctac tgaaaattgc
240aacagcgcga ccggttggac gggctatgca caagtgaatg gcaataacac
cgaaatcgtc 300acgtcctgga atctggcgta tgagggtggc agcggtccgg
ctattgaaca gggccaggat 360accttccaat acgtccctac gaccgagaat
aagtcccttc tgaaagacgg cggtggcggt 420tcgagctcgt ttcgccaacc
gagccagact gcgttgaaag atgagccgac ccatctggtt 480gttgcgaaag
agggcagcgt ggcatcgagc gtgctgctga gcggtacggt tactgccaaa
540aacgaacaat acgtgtactt cgatgctagc aagggtgatc tggatgaaat
tctggtgagc 600gtgggtgaca aagttagcga aggccaggca ctggtgaagt
attcatcctc cgaggcacag 660gcagcgtacg acagcgcaag ccgcgcagtg
gcgcgtgccg accgtcacat taacgaattg 720aaccaagcgc gtaacgaggc
cgcaagcgcg ccagcaccgc agctgccggc tccggtgggt 780ggcgaagatg
cgacggtgca gagcccgacc ccggttgcgg gtaattcggt cgccagcatc
840gatgcgcagc tgggtgacgc gcgtgatgcc cgtgcggatg cggctgctca
actgagcaag 900gctcagagcc aactggacgc gacgacggtg ctgagcacct
tggagggtac cgttgtcgaa 960gtcaacagca atgtgagcaa gagcccaacg
ggtgcgagcc aggttatggt ccacattgtg 1020agcaatgaaa acttacaggt
caagggtgag ctgagcgagt ataacctggc gaatctgagc 1080gttggtcaag
aggtcagctt taccagcaag gtctacccgg ataagaaatg gaccggcaag
1140ttgagctaca tcagcgacta cccgaagaac aatggcgagg cagcctcccc
ggcagccggc 1200aacaataccg gctctaagta tccgtacacc atcgacgtaa
ccggtgaggt cggcgacctg 1260aaacagggtt ttagcgtgaa tatcgaagtg
aagtccaaga ccaaggcaat tttggttccg 1320gttagctccc tggtgatgga
cgatagcaag aattatgtgt ggattgtcga cgagcaacag 1380aaagcgaaaa
aagttgaagt gagcctgggc aatgctgatg ccgagaacca agaaatcacg
1440tctggtctga ccaacggtgc gaaagttatt agcaacccga ccagcagcct
ggaagagggt 1500aaagaggtca aagccgacga agctacgaac cggccgcaac
cagcggcgac aattggtcca 1560aataccagag caacaagagc atcacgatcg
gcttcgacag cacttttgtg ccgatgggtt 1620tcgcgcaaaa agacggtagc
tacgcgggtt tcgatattga cctggcgacc gctgtctttg 1680agaaatacgg
cattacggtt aattggcagc cgattgattg ggacctgaaa gaggccgaac
1740tcaccaaagg caccatcgac ctgatctgga atggttactc cgcaaccgat
gagcgtcgcg 1800aaaaagttgc cttcagcaac agctatatga agaatgaaca
agtgttggta accaagaaat 1860ctagcggcat tacgaccgcg aaagacatga
ccggtaagac gctgggtgcg caggccggta 1920gctctggcta tgcggatttc
gaggcgaatc ctgagattct gaaaaacatc gttgcgaata 1980aagaggcgaa
ccagtaccag acctttaacg aagcactgat cgacctgaaa aacgatcgca
2040ttgacggtct gctgatcgat cgtgtgtacg cgaactatta tctggaagcc
gagggcgttc 2100tgaacgatta taatgttttt accgtgggtc tggagactga
ggcattcgcg gttggtgcgc 2160gcaaggaaga taccaacctg gttaaaaaga
ttaatgaggc atttagctca ctgtacaagg 2220acggcaagtt ccaagaaatt
agccagaagt ggttcggtga agatgttgcg acgaaagagg 2280ttaaagaggg ccaa
2294352294DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 35atgaccagcg gcgacaattg gtccaaatac
cagagcaaca agagcatcac gatcggcttc 60gacagcactt ttgtgccgat gggtttcgcg
caaaaagacg gtagctacgc gggtttcgat 120attgacctgg cgaccgctgt
ctttgagaaa tacggcatta cggttaattg gcagccgatt 180gattgggacc
tgaaagaggc cgaactcacc aaaggcacca tcgacctgat ctggaatggt
240tactccgcaa ccgatgagcg tcgcgaaaaa gttgccttca gcaacagcta
tatgaagaat 300gaacaagtgt tggtaaccaa gaaatctagc ggcattacga
ccgcgaaaga catgaccggt 360aagacgctgg gtgcgcaggc cggtagctct
ggctatgcgg atttcgaggc gaatcctgag 420attctgaaaa acatcgttgc
gaataaagag gcgaaccagt accagacctt taacgaagca 480ctgatcgacc
tgaaaaacga tcgcattgac ggtctgctga tcgatcgtgt gtacgcgaac
540tattatctgg aagccgaggg cgttctgaac gattataatg tttttaccgt
gggtctggag 600actgaggcat tcgcggttgg tgcgcgcaag gaagatacca
acctggttaa aaagattaat 660gaggcattta gctcactgta caaggacggc
aagttccaag aaattagcca gaagtggttc 720ggtgaagatg ttgcgacgaa
agaggttaaa gagggccaag gcggtggcgg ttcgagctcg 780tttcgccaac
cgagccagac tgcgttgaaa gatgagccga cccatctggt tgttgcgaaa
840gagggcagcg tggcatcgag cgtgctgctg agcggtacgg ttactgccaa
aaacgaacaa 900tacgtgtact tcgatgctag caagggtgat ctggatgaaa
ttctggtgag cgtgggtgac 960aaagttagcg aaggccaggc actggtgaag
tattcatcct ccgaggcaca ggcagcgtac 1020gacagcgcaa gccgcgcagt
ggcgcgtgcc gaccgtcaca ttaacgaatt gaaccaagcg 1080cgtaacgagg
ccgcaagcgc gccagcaccg cagctgccgg ctccggtggg tggcgaagat
1140gcgacggtgc agagcccgac cccggttgcg ggtaattcgg tcgccagcat
cgatgcgcag 1200ctgggtgacg cgcgtgatgc ccgtgcggat gcggctgctc
aactgagcaa ggctcagagc 1260caactggacg cgacgacggt gctgagcacc
ttggagggta ccgttgtcga agtcaacagc 1320aatgtgagca agagcccaac
gggtgcgagc caggttatgg tccacattgt gagcaatgaa 1380aacttacagg
tcaagggtga gctgagcgag tataacctgg cgaatctgag cgttggtcaa
1440gaggtcagct ttaccagcaa ggtctacccg gataagaaat ggaccggcaa
gttgagctac 1500atcagcgact acccgaagaa caatggcgag gcagcctccc
cggcagccgg caacaatacc 1560ggctctaagt atccgtacac catcgacgta
accggtgagg tcggcgacct gaaacagggt 1620tttagcgtga atatcgaagt
gaagtccaag accaaggcaa ttttggttcc ggttagctcc 1680ctggtgatgg
acgatagcaa gaattatgtg tggattgtcg acgagcaaca gaaagcgaaa
1740aaagttgaag tgagcctggg caatgctgat gccgagaacc aagaaatcac
gtctggtctg 1800accaacggtg cgaaagttat tagcaacccg accagcagcc
tggaagaggg taaagaggtc 1860aaagccgacg aagctacgaa ccggccgcat
tcgacgcatc caactttaaa gactttagca 1920gcatcgcgtc cgcaagctct
agctggcaga atcaatctgg tagcaccatg attatccaag 1980tggacagctt
tggtaacgtc agcggtcaat atgttaatcg tgcacagggt acgggttgtc
2040agaattctcc gtacccgctg accggtcgtg ttaacggcac gttcatcgct
ttcagcgtcg 2100gttggaacaa ttctactgaa aattgcaaca gcgcgaccgg
ttggacgggc tatgcacaag 2160tgaatggcaa taacaccgaa atcgtcacgt
cctggaatct ggcgtatgag ggtggcagcg 2220gtccggctat tgaacagggc
caggatacct tccaatacgt ccctacgacc gagaataagt 2280cccttctgaa agac
2294362294DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 36atgtttcgcc aaccgagcca gactgcgttg
aaagatgagc cgacccatct ggttgttgcg 60aaagagggca gcgtggcatc gagcgtgctg
ctgagcggta cggttactgc caaaaacgaa 120caatacgtgt acttcgatgc
tagcaagggt gatctggatg aaattctggt gagcgtgggt 180gacaaagtta
gcgaaggcca ggcactggtg aagtattcat cctccgaggc acaggcagcg
240tacgacagcg caagccgcgc agtggcgcgt gccgaccgtc acattaacga
attgaaccaa 300gcgcgtaacg aggccgcaag cgcgccagca ccgcagctgc
cggctccggt gggtggcgaa 360gatgcgacgg tgcagagccc gaccccggtt
gcgggtaatt cggtcgccag catcgatgcg 420cagctgggtg acgcgcgtga
tgcccgtgcg gatgcggctg ctcaactgag caaggctcag 480agccaactgg
acgcgacgac ggtgctgagc accttggagg gtaccgttgt cgaagtcaac
540agcaatgtga gcaagagccc aacgggtgcg agccaggtta tggtccacat
tgtgagcaat 600gaaaacttac aggtcaaggg tgagctgagc gagtataacc
tggcgaatct gagcgttggt 660caagaggtca gctttaccag caaggtctac
ccggataaga aatggaccgg caagttgagc 720tacatcagcg actacccgaa
gaacaatggc gaggcagcct ccccggcagc cggcaacaat 780accggctcta
agtatccgta caccatcgac gtaaccggtg aggtcggcga cctgaaacag
840ggttttagcg tgaatatcga agtgaagtcc aagaccaagg caattttggt
tccggttagc 900tccctggtga tggacgatag caagaattat gtgtggattg
tcgacgagca acagaaagcg 960aaaaaagttg aagtgagcct gggcaatgct
gatgccgaga accaagaaat cacgtctggt 1020ctgaccaacg gtgcgaaagt
tattagcaac ccgaccagca gcctggaaga gggtaaagag 1080gtcaaagccg
acgaagctac gaacggcggt ggcggttcga gctcgaccag cggcgacaat
1140tggtccaaat accagagcaa caagagcatc acgatcggct tcgacagcac
ttttgtgccg 1200atgggtttcg cgcaaaaaga cggtagctac gcgggtttcg
atattgacct ggcgaccgct 1260gtctttgaga aatacggcat tacggttaat
tggcagccga ttgattggga cctgaaagag 1320gccgaactca ccaaaggcac
catcgacctg atctggaatg gttactccgc aaccgatgag 1380cgtcgcgaaa
aagttgcctt cagcaacagc tatatgaaga atgaacaagt gttggtaacc
1440aagaaatcta gcggcattac gaccgcgaaa gacatgaccg gtaagacgct
gggtgcgcag 1500gccggtagct ctggctatgc ggatttcgag gcgaatcctg
agattctgaa aaacatcgtt 1560gcgaataaag aggcgaacca gtaccagacc
tttaacgaag cactgatcga cctgaaaaac 1620gatcgcattg acggtctgct
gatcgatcgt gtgtacgcga actattatct ggaagccgag 1680ggcgttctga
acgattataa tgtttttacc gtgggtctgg agactgaggc attcgcggtt
1740ggtgcgcgca aggaagatac caacctggtt aaaaagatta atgaggcatt
tagctcactg 1800tacaaggacg gcaagttcca agaaattagc cagaagtggt
tcggtgaaga tgttgcgacg 1860aaagaggtta aagagggcca acggccgcat
tcgacgcatc caactttaaa gactttagca 1920gcatcgcgtc cgcaagctct
agctggcaga atcaatctgg tagcaccatg attatccaag 1980tggacagctt
tggtaacgtc agcggtcaat atgttaatcg tgcacagggt acgggttgtc
2040agaattctcc gtacccgctg accggtcgtg ttaacggcac gttcatcgct
ttcagcgtcg 2100gttggaacaa ttctactgaa aattgcaaca gcgcgaccgg
ttggacgggc tatgcacaag 2160tgaatggcaa taacaccgaa atcgtcacgt
cctggaatct ggcgtatgag ggtggcagcg 2220gtccggctat tgaacagggc
caggatacct tccaatacgt ccctacgacc gagaataagt 2280cccttctgaa agac
2294377PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 37Gly Gly Gly Gly Ser Ser Ser1 5383PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 38Ala
Ala Ala139150PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptideMISC_FEATURE(1)..(150)This sequence
may encompass 1-20, 25 or 30 "Gly Gly Gly Gly Ser" repeating
unitsSee specification as filed for detailed description of
substitutions and preferred embodiments 39Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly1 5 10 15Gly Gly Gly Ser Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly 20 25 30Gly Gly Ser Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly 35 40 45Gly Ser Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 50 55 60Ser Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser65 70 75
80Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly
85 90 95Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly
Gly 100 105 110Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser
Gly Gly Gly 115 120 125Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly
Ser Gly Gly Gly Gly 130 135 140Ser Gly Gly Gly Gly Ser145
150406PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 40Gly Gly Gly Gly Gly Gly1 54115PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 41Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser1 5 10
154230PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 42Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser Gly1 5 10 15Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser 20 25 304318PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 43Lys Glu Ser Gly Ser Val Ser
Ser Glu Gln Leu Ala Gln Phe Arg Ser1 5 10 15Leu
Asp4414PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 44Glu Gly Lys Ser Ser Gly Ser Gly Ser Glu Ser Lys
Ser Thr1 5 104530PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptideMISC_FEATURE(1)..(30)This sequence
may encompass 1-20, 25 or 30 residuesSee specification as filed for
detailed description of substitutions and preferred embodiments
45Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly1
5 10 15Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly 20
25 30468PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 46Gly Gly Gly Gly Gly Gly Gly Gly1
54712PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 47Gly Ser Ala Gly Ser Ala Ala Gly Ser Gly Glu
Phe1 5 1048150PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptideMISC_FEATURE(1)..(150)This sequence
may encompass 1-20, 25 or 30 "Glu Ala Ala Ala Lys" repeating
unitsSee specification as filed for detailed description of
substitutions and preferred embodiments 48Glu Ala Ala Ala Lys Glu
Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu1 5 10 15Ala Ala Ala Lys Glu
Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala 20 25 30Ala Ala Lys Glu
Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala 35 40 45Ala Lys Glu
Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala 50 55 60Lys Glu
Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys65 70 75
80Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu
85 90 95Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu
Ala 100 105 110Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys
Glu Ala Ala 115 120 125Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala
Lys Glu Ala Ala Ala 130 135 140Lys Glu Ala Ala Ala Lys145
15049152PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptideMISC_FEATURE(2)..(151)This region may
encompass 1-20, 25 or 30 "Glu Ala Ala Ala Lys" repeating unitsSee
specification as filed for detailed description of substitutions
and preferred embodiments 49Ala Glu Ala Ala Ala Lys Glu Ala Ala Ala
Lys Glu Ala Ala Ala Lys1 5 10 15Glu Ala Ala Ala Lys Glu Ala Ala Ala
Lys Glu Ala Ala Ala Lys Glu 20 25 30Ala Ala Ala Lys Glu Ala Ala Ala
Lys Glu Ala Ala Ala Lys Glu Ala 35 40 45Ala Ala Lys Glu Ala Ala Ala
Lys Glu Ala Ala Ala Lys Glu Ala Ala 50 55 60Ala Lys Glu Ala Ala Ala
Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala65 70 75 80Lys Glu Ala Ala
Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys 85 90 95Glu Ala Ala
Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu 100 105 110Ala
Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala 115 120
125Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala
130 135 140Ala Lys Glu Ala Ala Ala Lys Ala145 1505046PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
50Ala Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys1
5 10 15Glu Ala Ala Ala Lys Ala Leu Glu Ala Glu Ala Ala Ala Lys Glu
Ala 20 25 30Ala Ala Lys Glu Ala Ala Ala Lys Glu Ala Ala Ala Lys Ala
35 40 455144PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptideMISC_FEATURE(1)..(44)This sequence
may encompass 1-2 "A(EAAAK)nA" repeating units where n=2-4See
specification as filed for detailed description of substitutions
and preferred embodiments 51Ala Glu Ala Ala Ala Lys Glu Ala Ala Ala
Lys Glu Ala Ala Ala Lys1 5 10 15Glu Ala Ala Ala Lys Ala Ala Glu Ala
Ala Ala Lys Glu Ala Ala Ala 20 25 30Lys Glu Ala Ala Ala Lys Glu Ala
Ala Ala Lys Ala 35 405212PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 52Ala Glu Ala Ala Ala Lys Glu
Ala Ala Ala Lys Ala1 5 105360PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptideMOD_RES(1)..(1)Any amino
acidMOD_RES(3)..(3)Any amino acidMOD_RES(5)..(5)Any amino
acidMOD_RES(7)..(7)Any amino acidMOD_RES(9)..(9)Any amino
acidMOD_RES(11)..(11)Any amino acidMOD_RES(13)..(13)Any amino
acidMOD_RES(15)..(15)Any amino acidMOD_RES(17)..(17)Any amino
acidMOD_RES(19)..(19)Any amino acidMOD_RES(21)..(21)Any amino
acidMOD_RES(23)..(23)Any amino acidMOD_RES(25)..(25)Any amino
acidMOD_RES(27)..(27)Any amino acidMOD_RES(29)..(29)Any amino
acidMOD_RES(31)..(31)Any amino acidMOD_RES(33)..(33)Any amino
acidMOD_RES(35)..(35)Any amino acidMOD_RES(37)..(37)Any amino
acidMOD_RES(39)..(39)Any amino acidMOD_RES(41)..(41)Any amino
acidMOD_RES(43)..(43)Any amino acidMOD_RES(45)..(45)Any amino
acidMOD_RES(47)..(47)Any amino acidMOD_RES(49)..(49)Any amino
acidMOD_RES(51)..(51)Any amino acidMOD_RES(53)..(53)Any amino
acidMOD_RES(55)..(55)Any amino acidMOD_RES(57)..(57)Any amino
acidMOD_RES(59)..(59)Any amino acidMISC_FEATURE(1)..(60)This
sequence may encompass 1-20, 25 or 30 "Xaa Pro" repeating unitsSee
specification as filed for detailed description of substitutions
and preferred embodiments 53Xaa Pro Xaa Pro Xaa Pro Xaa Pro Xaa Pro
Xaa Pro Xaa Pro Xaa Pro1 5 10 15Xaa Pro Xaa Pro Xaa Pro Xaa Pro Xaa
Pro Xaa Pro Xaa Pro Xaa Pro 20 25 30Xaa Pro Xaa Pro Xaa Pro Xaa Pro
Xaa Pro Xaa Pro Xaa Pro Xaa Pro 35 40 45Xaa Pro Xaa Pro Xaa Pro Xaa
Pro Xaa Pro Xaa Pro 50 55 605460PRTArtificial SequenceDescription
of Artificial Sequence Synthetic
polypeptideMISC_FEATURE(1)..(60)This sequence may encompass 1-20,
25 or 30 "Ala Pro" repeating unitsSee specification as filed for
detailed description of substitutions and preferred embodiments
54Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro1
5 10 15Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala
Pro 20 25 30Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro
Ala Pro 35 40 45Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro Ala Pro 50
55 605560PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptideMISC_FEATURE(1)..(60)This sequence may
encompass 1-20, 25 or 30 "Lys Pro" repeating unitsSee specification
as filed for detailed description of substitutions and preferred
embodiments 55Lys Pro Lys Pro Lys Pro Lys Pro Lys Pro Lys Pro Lys
Pro Lys Pro1 5 10 15Lys Pro Lys Pro Lys Pro Lys Pro Lys Pro Lys Pro
Lys Pro Lys Pro 20 25 30Lys Pro Lys Pro Lys Pro Lys Pro Lys Pro Lys
Pro Lys Pro Lys Pro 35 40 45Lys Pro Lys Pro Lys Pro Lys Pro Lys Pro
Lys Pro 50 55 605660PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptideMISC_FEATURE(1)..(60)This sequence
may encompass 1-20, 25 or 30 "Gln Pro" repeating unitsSee
specification as filed for detailed description of substitutions
and preferred embodiments 56Gln Pro Gln Pro Gln Pro Gln Pro Gln Pro
Gln Pro Gln Pro Gln Pro1 5 10 15Gln Pro Gln Pro Gln Pro Gln Pro Gln
Pro Gln Pro Gln Pro Gln Pro 20 25 30Gln Pro Gln Pro Gln Pro Gln Pro
Gln Pro Gln Pro Gln Pro Gln Pro 35 40 45Gln Pro Gln Pro Gln Pro Gln
Pro Gln Pro Gln Pro 50 55 605714PRTArtificial SequenceDescription
of Artificial Sequence Synthetic peptide 57Ala Pro Ala Pro Ala Pro
Ala Pro Ala Pro Ala Pro Ala Pro1 5 105821PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 58Gly
Ala Pro Gly Gly Gly Gly Gly Ala Ala Ala Ala Ala Gly Gly Gly1 5 10
15Gly Gly Gly Ala Pro 205939PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 59Gly Ala Pro Gly Gly Gly
Gly Gly Ala Ala Ala Ala Ala Gly Gly Gly1 5 10 15Gly Gly Gly Ala Pro
Gly Gly Gly Gly Gly Ala Ala Ala Ala Ala Gly 20 25 30Gly Gly Gly Gly
Gly Ala Pro 356057PRTArtificial SequenceDescription of Artificial
Sequence Synthetic polypeptide 60Gly Ala Pro Gly Gly Gly Gly Gly
Ala Ala Ala Ala Ala Gly Gly Gly1 5 10 15Gly Gly Gly Ala Pro Gly Gly
Gly Gly Gly Ala Ala Ala Ala Ala Gly 20 25 30Gly Gly Gly Gly Gly Ala
Pro Gly Gly Gly Gly Gly Ala Ala Ala Ala 35 40 45Ala Gly Gly Gly Gly
Gly Gly Ala Pro 50 55614PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptide 61Gly Gly Gly
Gly1625PRTArtificial SequenceDescription of Artificial Sequence
Synthetic peptide 62Gly Gly Gly Gly Ser1 5635PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 63Glu
Ala Ala Ala Lys1 56410PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptideMISC_FEATURE(1)..(10)This
sequence may encompass 2-10 residues 64His His His His His His His
His His His1 5 10655PRTStreptococcus pneumoniae 65Pro Ala Pro Ala
Pro1 5666PRTStreptococcus pneumoniae 66Pro Lys Glu Pro Glu Gln1
5674PRTStreptococcus pneumoniae 67Pro Glu Lys
Pro16815PRTStreptococcus pneumoniae 68Ser Pro Ala Ala Gly Asn Asn
Thr Gly Ser Lys Tyr Pro Tyr Thr1 5 10 156915PRTStreptococcus
pneumoniae 69Val Met Asp Asp Ser Lys Asn Tyr Val Trp Ile Val Asp
Glu Gln1 5 10 157015PRTStreptococcus pneumoniae 70Ala Ser Pro Ala
Ala Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr1 5 10
157115PRTStreptococcus pneumoniae 71Ala Ala Ser Pro Ala Ala Gly Asn
Asn Thr Gly Ser Lys Tyr Pro1 5 10 157215PRTStreptococcus pneumoniae
72Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile Thr Ser Gly1 5 10
157315PRTStreptococcus pneumoniae 73Phe Thr Ser Lys Val Tyr Pro Asp
Lys Lys Trp Thr Gly Lys Leu1 5 10 157415PRTStreptococcus pneumoniae
74Ile Asp Val Thr Gly Glu Val Gly Asp Leu Lys Gln Gly Phe Ser1 5 10
157515PRTStreptococcus pneumoniae 75Ser Ser Leu Glu Glu Gly Lys Glu
Val Lys Ala Asp Glu Ala Thr1 5 10 157615PRTStreptococcus pneumoniae
76Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn1 5 10
157715PRTStreptococcus pneumoniae 77Thr Ile Asp Val Thr Gly Glu Val
Gly Asp Leu Lys Gln Gly Phe1 5 10 157815PRTStreptococcus pneumoniae
78Ala Ala Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp1 5 10
157915PRTStreptococcus pneumoniae 79Pro Gln Leu Pro Ala Pro Val Gly
Gly Glu Asp Ala Thr Val Gln1 5 10 158015PRTStreptococcus pneumoniae
80Gln Leu Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser1 5 10
158115PRTStreptococcus pneumoniae 81Leu Pro Ala Pro Val Gly Gly Glu
Asp Ala Thr Val Gln Ser Pro1 5 10 158215PRTStreptococcus pneumoniae
82Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr1 5 10
158315PRTStreptococcus pneumoniae 83Ala Pro Val Gly Gly Glu Asp Ala
Thr Val Gln Ser Pro Thr Pro1 5 10 158415PRTStreptococcus pneumoniae
84Pro Ala Ala Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile1 5 10
158515PRTStreptococcus pneumoniae 85Ala Gly Asn Asn Thr Gly Ser Lys
Tyr Pro Tyr Thr Ile Asp Val1 5 10 158615PRTStreptococcus pneumoniae
86Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser1 5 10
158715PRTStreptococcus pneumoniae 87Ser Phe Thr Ser Lys Val Tyr Pro
Asp Lys Lys Trp Thr Gly Lys1 5 10 158815PRTStreptococcus pneumoniae
88Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro1 5 10
158915PRTStreptococcus pneumoniae 89Thr Val Asn Trp Gln Pro Ile Asp
Trp Asp Leu Lys Glu Ala Glu1 5 10 159015PRTStreptococcus pneumoniae
90Val Ala Asn Lys Glu Ala Asn Gln Tyr Gln Thr Phe Asn Glu Ala1 5 10
159115PRTStreptococcus pneumoniae 91Ile Thr Val Asn Trp Gln Pro Ile
Asp Trp Asp Leu Lys Glu Ala1 5 10 159215PRTStreptococcus pneumoniae
92Gly Ile Thr Val Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu1 5 10
159315PRTStreptococcus pneumoniae 93Val Asn Trp Gln Pro Ile Asp Trp
Asp Leu Lys Glu Ala Glu Leu1 5 10 159415PRTStreptococcus pneumoniae
94Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly Glu1 5 10
159515PRTStreptococcus pneumoniae 95Asp Gly Lys Phe Gln Glu Ile Ser
Gln Lys Trp Phe Gly Glu Asp1 5 10 159615PRTStreptococcus pneumoniae
96Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly Glu Asp Val1 5 10
159715PRTStreptococcus pneumoniae 97Lys Phe Gln Glu Ile Ser Gln Lys
Trp Phe Gly Glu Asp Val Ala1 5 10 159815PRTStreptococcus pneumoniae
98Trp Phe Gly Glu Asp Val Ala Thr Lys Glu Val Lys Glu Gly Gln1 5 10
159915PRTStreptococcus pneumoniae 99Gly Ala Gln Ala Gly Ser Ser Gly
Tyr Ala Asp Phe Glu Ala Asn1 5 10 1510015PRTStreptococcus
pneumoniae 100Tyr Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp
Phe Gly1 5 10 1510115PRTStreptococcus pneumoniae 101Asn Glu Ala Phe
Ser Ser Leu Tyr Lys Asp Gly Lys Phe Gln Glu1 5 10
1510215PRTStreptococcus pneumoniae 102Tyr Gly Ile Thr Val Asn Trp
Gln Pro Ile Asp Trp Asp Leu Lys1 5 10 1510315PRTStreptococcus
pneumoniae 103Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys
Val Ala1 5 10 1510415PRTStreptococcus pneumoniae 104Ala Arg Lys Glu
Asp Thr Asn Leu Val Lys Lys Ile Asn Glu Ala1 5 10
1510515PRTStreptococcus pneumoniae 105Ser Ala Thr
Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser1 5 10
1510615PRTStreptococcus pneumoniae 106Asn Gly Tyr Ser Ala Thr Asp
Glu Arg Arg Glu Lys Val Ala Phe1 5 10 1510715PRTStreptococcus
pneumoniae 107Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys Val Ala
Phe Ser1 5 10 1510827PRTArtificial SequenceDescription of
Artificial Sequence Synthetic peptideMISC_FEATURE(2)..(26)This
region may encompass 2-5 "Glu Ala Ala Ala Lys" repeating unitsSee
specification as filed for detailed description of substitutions
and preferred embodiments 108Ala Glu Ala Ala Ala Lys Glu Ala Ala
Ala Lys Glu Ala Ala Ala Lys1 5 10 15Glu Ala Ala Ala Lys Glu Ala Ala
Ala Lys Ala 20 2510934PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptideMISC_FEATURE(1)..(34)This
sequence may encompass 5-17 "Ala Pro" repeating unitsSee
specification as filed for detailed description of substitutions
and preferred embodiments 109Ala Pro Ala Pro Ala Pro Ala Pro Ala
Pro Ala Pro Ala Pro Ala Pro1 5 10 15Ala Pro Ala Pro Ala Pro Ala Pro
Ala Pro Ala Pro Ala Pro Ala Pro 20 25 30Ala
Pro110399PRTStreptococcus pneumoniae 110Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asn Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395111399PRTStreptococcus pneumoniae
111Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr
His Leu 35 40 45Ile Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly
Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr
Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu
Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
395112399PRTStreptococcus pneumoniae 112Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Asn Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395113399PRTStreptococcus pneumoniae
113Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr
His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Ser Lys 340 345 350Lys Val Glu Val Ser Leu Gly
Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr
Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu
Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
395114399PRTStreptococcus pneumoniae 114Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Thr Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395115399PRTStreptococcus pneumoniae
115Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr
His Phe 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245
250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser
Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser
Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr
Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe
Ser Val Asn Ile Glu Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile
Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn
Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val
Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360
365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser
370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr
Asn385 390 395116399PRTStreptococcus pneumoniae 116Met Lys Lys Lys
Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala
Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg
Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val
Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55
60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65
70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser
Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala
Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg
His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser
Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp
Ala Thr Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser
Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala
Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln
Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200
205Glu Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val
210 215 220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly
Glu Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly
Gln Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys
Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn
Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly
Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly
Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315
320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp
325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys
Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu
Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val
Ile Ser Asn Pro Thr Phe 370 375 380Ser Leu Glu Glu Gly Lys Glu Val
Lys Ala Asp Glu Ala Thr Asn385 390 395117399PRTStreptococcus
pneumoniae 117Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu
Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly
Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp
Glu Pro Thr His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser
Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr
Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu
Val Ser Val Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys
Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser
Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn
Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135
140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr
Pro145 150 155 160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln
Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln
Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu
Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val
Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile
Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser
Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250
255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr
260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn Gly Ala Ala Ala Ser Pro
Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile
Asp Val Thr Gly 290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser
Val Asn Ile Glu Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu
Val Pro Val Ser Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr
Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu
Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr
Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375
380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385
390 395118399PRTStreptococcus pneumoniae 118Met Lys Lys Lys Asn Gly
Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser
Val Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro
Ser Gln Thr Ala Leu Lys Asp Glu Pro Ile His Leu 35 40 45Val Val Ala
Lys Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val
Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75
80Gly Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu
85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala
Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His
Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala
Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala
Thr Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val
Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg
Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu
Asp Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200
205Glu Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val
210 215 220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly
Glu Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly
Gln Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys
Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn
Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly
Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly
Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315
320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp
325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys
Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu
Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val
Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val
Lys Ala Asp Glu Ala Thr Asn385 390 395119399PRTStreptococcus
pneumoniae 119Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu
Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly
Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp
Glu Pro Thr His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser
Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr
Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu
Val Ser Val Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys
Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser
Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn
Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135
140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr
Pro145 150 155 160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln
Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln
Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu
Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val
Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile
Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser
Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250
255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr
260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro
Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile
Asp Val Thr Gly 290 295 300Glu Gly Gly Asp Leu Lys Gln Gly Phe Ser
Val Asn Ile Glu Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu
Val Pro Val Ser Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr
Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu
Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr
Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375
380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385
390 395120399PRTStreptococcus pneumoniae 120Met Lys Lys Lys Asn Gly
Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser
Val Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro
Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Ala
Lys Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val
Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75
80Gly Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu
85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala
Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His
Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala
Leu Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala
Thr Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val
Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg
Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu
Asp Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200
205Glu Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val
210 215 220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly
Glu Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly
Gln Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys
Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn
Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly
Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly
Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315
320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp
325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys
Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu
Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val
Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val
Lys Ala Asp Glu Ala Thr Asn385 390 395121399PRTStreptococcus
pneumoniae 121Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu
Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly
Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp
Glu Pro Thr His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser
Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr
Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu
Val Ser Val Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys
Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser
Arg Ala Val Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn
Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135
140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr Val Pro Ser Pro Thr
Pro145 150 155 160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln
Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln
Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu
Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val
Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile
Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser
Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250
255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr
260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro
Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile
Asp Val Thr Gly 290 295
300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val
Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser
Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp
Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly Asn
Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn
Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu
Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
395122399PRTStreptococcus pneumoniae 122Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Ile Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Ile His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395123399PRTStreptococcus pneumoniae
123Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Leu Thr
His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Arg Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly
Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr
Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu
Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
395124399PRTStreptococcus pneumoniae 124Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ser 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Ile Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395125399PRTStreptococcus pneumoniae
125Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr
His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Lys Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Arg Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly
Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr
Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu
Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
395126399PRTStreptococcus pneumoniae 126Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Ile His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ser 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Ile Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395127399PRTStreptococcus pneumoniae
127Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr
His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Lys Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Gln Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Ala Lys 340 345
350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln Glu Ile
355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser Asn Pro
Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala Asp Glu
Ala Thr Asn385 390 395128399PRTStreptococcus pneumoniae 128Met Lys
Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile
Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25
30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu
35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser
Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala
Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp
Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu
Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg
Val Asp Cys His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu
Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly
Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155 160Val Ala
Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170
175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser
180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr
Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly
Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu Asn Leu
Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn
Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr
Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp
Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly
Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295
300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val
Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser
Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp
Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly Asn
Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn
Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu
Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
395129399PRTStreptococcus pneumoniae 129Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Tyr
Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Val Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Arg Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395130399PRTStreptococcus pneumoniae
130Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Val Lys Asp Glu Pro Thr
His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Ala Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Lys Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Val Gly Asn Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly
Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr
Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu
Glu Gly Lys Glu Val Lys Ala Asp Glu Ala Thr Asn385 390
395131399PRTStreptococcus pneumoniae 131Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Ile Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Ile His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Lys Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395132399PRTStreptococcus pneumoniae
132Met Lys Lys Lys Asn Gly Lys Ala Lys Lys Trp Gln Leu Tyr Ala Ala1
5 10 15Ile Gly Ala Ala Ser Val Val Val Leu Gly Ala Gly Gly Ile Leu
Leu 20 25 30Phe Arg Gln Pro Ser Gln Thr Ala Leu Lys Asp Glu Pro Thr
His Leu 35 40 45Val Val Ala Lys Glu Gly Ser Val Ala Ser Ser Val Leu
Leu Ser Gly 50 55 60Thr Val Thr Ala Lys Asn Glu Gln Tyr Val Tyr Phe
Asp Ala Ser Lys65 70 75 80Gly Asp Leu Asp Glu Ile Leu Val Ser Val
Gly Asp Lys Val Ser Glu 85 90 95Gly Gln Ala Leu Val Lys Tyr Ser Ser
Ser Glu Ala Gln Ala Val Tyr 100 105 110Asp Ser Ala Ser Arg Ala Val
Ala Lys Ala Asp Arg His Ile Asn Glu 115 120 125Leu Asn Gln Ala Arg
Asn Glu Ala Ala Ser Ala Pro Ala Pro Gln Leu 130 135 140Pro Ala Pro
Val Gly Gly Glu Asp Ala Thr Val Gln Ser Pro Thr Pro145 150 155
160Val Ala Gly Asn Ser Val Ala Ser Ile Asp Ala Gln Leu Gly Asp Ala
165 170 175Arg Asp Ala Arg Ala Asp Ala Ala Ala Gln Leu Ser Lys Ala
Gln Ser 180 185 190Gln Leu Asp Ala Thr Thr Val Leu Ser Thr Leu Glu
Gly Thr Val Val 195 200 205Glu Val Asn Ser Asn Val Ser Lys Ser Pro
Thr Gly Ala Ser Gln Val 210 215 220Met Val His Ile Val Ser Asn Glu
Asn Leu Gln Val Lys Gly Glu Leu225 230 235 240Ser Glu Tyr Asn Leu
Ala Asn Leu Ser Val Gly Gln Glu Val Ser Phe 245 250 255Thr Ser Lys
Val Tyr Pro Asp Lys Lys Trp Thr Gly Lys Leu Ser Tyr 260 265 270Ile
Ser Asp Tyr Pro Lys Asn Asn Gly Glu Ala Ala Ser Pro Ala Ala 275 280
285Gly Asn Asn Thr Gly Ser Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly
290 295 300Glu Val Gly Asp Leu Lys Gln Gly Phe Ser Val Asn Ile Glu
Val Lys305 310 315 320Ser Lys Thr Lys Ala Ile Leu Val Pro Val Ser
Ser Leu Val Met Asp 325 330 335Asp Ser Lys Asn Tyr Val Trp Ile Val
Asp Glu Gln Gln Lys Ala Lys 340 345 350Lys Val Glu Val Ser Leu Gly
Asn Ala Asp Ala Glu Asn Gln Glu Ile 355 360 365Thr Ser Gly Leu Thr
Asn Gly Ala Lys Val Ile Ser Asn Pro Thr Ser 370 375 380Ser Leu Glu
Glu Gly Lys Glu Val Lys Thr Asp Glu Ala Thr Asn385 390
395133399PRTStreptococcus pneumoniae 133Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Ala Lys
Glu Gly Ser Val Ala Ser Ser Ile Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Lys Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Val Arg Asn Glu Ala Ala Ser Ala Pro
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390
395134399PRTStreptococcus pneumoniae 134Met Lys Lys Lys Asn Gly Lys
Ala Lys Lys Trp Gln Leu Tyr Ala Ala1 5 10 15Ile Gly Ala Ala Ser Val
Val Val Leu Gly Ala Gly Gly Ile Leu Leu 20 25 30Phe Arg Gln Pro Ser
Gln Thr Ala Leu Lys Asp Glu Pro Thr His Leu 35 40 45Val Val Ala Asn
Glu Gly Ser Val Ala Ser Ser Val Leu Leu Ser Gly 50 55 60Thr Val Thr
Ala Lys Asn Glu Gln Tyr Val Tyr Phe Asp Ala Ser Lys65 70 75 80Gly
Asp Leu Asp Glu Ile Leu Val Ser Val Gly Asp Lys Val Ser Glu 85 90
95Gly Gln Ala Leu Val Lys Tyr Ser Ser Ser Glu Ala Gln Ala Ala Tyr
100 105 110Asp Ser Ala Ser Arg Ala Val Ala Lys Ala Asp Arg His Ile
Asn Glu 115 120 125Leu Asn Gln Ala Arg Asn Glu Ala Ala Ser Ala Gln
Ala Pro Gln Leu 130 135 140Pro Ala Pro Val Gly Gly Glu Asp Ala Thr
Val Gln Ser Pro Thr Pro145 150 155 160Val Ala Gly Asn Ser Val Ala
Ser Ile Asp Ala Gln Leu Gly Asp Ala 165 170 175Arg Asp Ala Arg Ala
Asp Ala Ala Ala Gln Leu Ser Lys Ala Gln Ser 180 185 190Gln Leu Asp
Ala Thr Thr Val Leu Ser Thr Leu Glu Gly Thr Val Val 195 200 205Glu
Val Asn Ser Asn Val Ser Lys Ser Pro Thr Gly Ala Ser Gln Val 210 215
220Met Val His Ile Val Ser Asn Glu Asn Leu Gln Val Lys Gly Glu
Leu225 230 235 240Ser Glu Tyr Asn Leu Ala Asn Leu Ser Val Gly Gln
Glu Val Ser Phe 245 250 255Thr Ser Lys Val Tyr Pro Asp Lys Lys Trp
Thr Gly Lys Leu Ser Tyr 260 265 270Ile Ser Asp Tyr Pro Lys Asn Asn
Gly Glu Ala Ala Ser Pro Ala Ala 275 280 285Gly Asn Asn Thr Gly Ser
Lys Tyr Pro Tyr Thr Ile Asp Val Thr Gly 290 295 300Glu Val Gly Asp
Leu Lys Gln Gly Phe Ser Val Asn Ile Glu Val Lys305 310 315 320Ser
Lys Thr Lys Ala Ile Leu Val Pro Val Ser Ser Leu Val Met Asp 325 330
335Asp Ser Lys Asn Tyr Val Trp Ile Val Asp Glu Gln Gln Lys Ala Lys
340 345 350Lys Val Glu Val Ser Leu Gly Asn Ala Asp Ala Glu Asn Gln
Glu Ile 355 360 365Thr Ser Gly Leu Thr Asn Gly Ala Lys Val Ile Ser
Asn Pro Thr Ser 370 375 380Ser Leu Glu Glu Gly Lys Glu Val Lys Ala
Asp Glu Ala Thr Asn385 390 395135278PRTStreptococcus pneumoniae
135Met Lys Lys Trp Met Leu Val Leu Val Ser Leu Met Thr Ala Leu Phe1
5 10 15Leu Val Ala Cys Gly Lys Asn Ser Ser Glu Thr Ser Gly Asp Asn
Trp 20 25 30Ser Lys Tyr Gln Ser Asn Lys Ser Ile Thr Ile Gly Phe Asp
Ser Thr 35 40 45Phe Val Pro Met Gly Phe Ala Gln Lys Asp Gly Ser Tyr
Ala Gly Phe 50 55 60Asp Ile Asp Leu Ala Thr Ala Val Phe Glu Lys Tyr
Gly Ile Thr Val65 70 75 80Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys
Glu Ala Glu Leu Thr Lys 85 90 95Gly Thr Ile Asp Leu Ile Trp Asn Gly
Tyr Ser Ala Thr Asp Glu Arg 100 105 110Arg Glu Lys Val Ala Phe Ser
Asn Ser Tyr Met Lys Asn Glu Gln Val 115 120 125Leu Val Thr Lys Lys
Ser Ser Gly Ile Thr Thr Ala Lys Asp Met Thr 130 135 140Gly Lys Thr
Leu Gly Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp Phe145 150 155
160Glu Ala Asn Pro Glu Leu Leu Lys Asn Ile Val Ala Asn Lys Glu Val
165 170 175Asn Gln Tyr Gln Thr Phe Asn Glu Ala Leu Ile Asp Leu Lys
Asn Asp 180 185 190Arg Ile Asp Gly Leu Leu Ile Asp Arg Val Tyr Ala
Asn Tyr Tyr Leu 195 200 205Glu Ala Glu Gly Val Leu Asn Asp Tyr Asn
Val Phe Thr Val Gly Leu 210 215 220Glu Thr Glu Ala Phe Ala Val Gly
Ala Arg Lys Glu Asp Thr Thr Leu225 230 235 240Val Lys Lys Ile Asn
Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys 245 250 255Phe Gln Glu
Ile Ser Gln Lys Trp Phe Gly Glu Asp Val Ala Thr Lys 260 265 270Glu
Val Lys Glu Gly Gln 275136278PRTStreptococcus pneumoniae 136Met Lys
Lys Trp Met Leu Val Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu
Val Ala Cys Gly Lys Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25
30Ser Lys Tyr Gln Ser Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr
35 40 45Phe Val Pro Met Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly
Phe 50 55 60Asp Ile Asp Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile
Thr Val65 70 75 80Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala
Glu Leu Thr Lys 85 90 95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser
Ala Thr Asp Glu Arg 100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser
Tyr Met Lys Asn Glu Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser
Gly Ile Thr Thr Ala Lys Asp Met Thr 130 135 140Gly Lys Thr Leu Gly
Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala
Asn Pro Glu Leu Leu Lys Asn Ile Val Ala Asn Lys Glu Ala 165 170
175Asn Gln Tyr Gln Thr Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp
180 185 190Arg Ile Asp Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr
Tyr Leu 195 200 205Glu Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe
Thr Val Gly Leu 210 215 220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg
Lys Glu Asp Thr Thr Leu225 230 235 240Val Lys Lys Ile Asn Glu Asp
Phe Ser Ser Leu Tyr Lys Asn Gly Lys 245 250 255Phe Gln Glu Ile Ser
Gln Lys Trp Phe Gly Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys
Glu Gly Gln 275137278PRTStreptococcus pneumoniae 137Met Lys Lys Trp
Met His Val Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala
Cys Gly Lys Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys
Tyr Gln Ser Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe
Val Pro Met Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55
60Asp Ile Asp Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65
70 75 80Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr
Lys 85 90 95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp
Glu Arg 100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys
Asn Glu Gln Val 115 120 125Leu Val Thr Lys Lys Leu Ser Gly Ile Thr
Thr Ala Lys Asp Met Thr 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala
Gly Ser Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Glu
Leu Leu Lys Asn Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr
Gln Thr Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg
Ile Asp Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200
205Glu Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu
210 215 220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr
Asn Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu
Tyr Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe
Gly Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275138278PRTStreptococcus pneumoniae 138Met Lys Lys Trp Met Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Gln Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Leu Ser Gly Ile Thr Thr Ala
Lys Asp Met Thr 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Glu Leu Leu
Lys Asn Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275139278PRTStreptococcus pneumoniae 139Met Lys Lys Trp Met Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Thr Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala
Lys Asp Met Ala 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Glu Leu Leu
Lys Asn Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275140278PRTStreptococcus pneumoniae 140Met Lys Lys Trp Met Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala
Lys Asp Met Ala 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu
Lys Asp Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275141282PRTStreptococcus pneumoniae 141Met Lys Lys Trp Met Lys Lys
Trp Met Leu Val Leu Val Ser Leu Met1 5 10 15Thr Ala Leu Phe Leu Val
Ala Cys Gly Lys Asn Thr Ser Glu Thr Ser 20 25 30Gly Asp Asn Trp Ser
Lys Tyr Glu Ser Asn Lys Ser Ile Thr Ile Gly 35 40 45Phe Asp Ser Thr
Phe Val Pro Met Gly Phe Ala Gln Lys Asp Gly Ser 50 55 60Tyr Ala Gly
Phe Asp Ile Asp Leu Ala Thr Ala Val Phe Glu Lys Tyr65 70 75 80Gly
Ile Thr Val Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala 85 90
95Glu Leu Thr Lys Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala
100 105 110Thr Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr
Met Lys 115 120 125Asn Glu Gln Val Leu Val Thr Lys Lys Ser Ser Gly
Ile Thr Thr Ala 130 135 140Lys Asp Met Ala Gly Lys Thr Leu Gly Ala
Gln Ala Gly Ser Ser Gly145 150 155 160Tyr Ala Asp Phe Glu Ala Asn
Pro Glu Leu Leu Lys Asn Ile Val Ala 165 170 175Asn Lys Glu Ala Asn
Gln Tyr Gln Thr Phe Asn Glu Ala Leu Ile Asp 180 185 190Leu Lys Asn
Asp Arg Ile Asp Gly Leu Leu Ile Asp Arg Val Tyr Ala 195 200 205Asn
Tyr Tyr Leu Glu Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe 210 215
220Thr Val Gly Leu Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys
Glu225 230 235 240Asp Thr Thr Leu Val Lys Lys Ile Asn Glu Ala Phe
Ser Ser Leu Tyr 245 250 255Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln
Lys Trp Phe Gly Glu Asp 260 265 270Val Ala Thr Lys Glu Val Lys Glu
Gly Gln 275 280142278PRTStreptococcus pneumoniae 142Met Lys Lys Trp
Met Leu Val Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala
Cys Gly Lys Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys
Tyr Gln Ser Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe
Val Pro Met Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55
60Asp Ile Asp Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65
70 75 80Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr
Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Leu Ser Gly Ile Thr Thr Ala
Lys Asp Met Thr 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Glu Leu Leu
Lys Asn Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Gly Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275143278PRTStreptococcus pneumoniae 143Met Lys Lys Trp Met Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala
Lys Asp Met Ala 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu
Lys Asp Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Glu
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275144282PRTStreptococcus pneumoniae 144Met Lys Lys Trp Met Lys Lys
Trp Met Leu Val Leu Val Ser Leu Met1 5 10 15Thr Ala Leu Phe Leu Val
Ala Cys Gly Lys Asn Ser Ser Glu Thr Ser 20 25 30Gly Asp Asn Trp Ser
Lys Tyr Gln Ser Asn Lys Ser Ile Thr Ile Gly 35 40 45Phe Asp Ser Thr
Phe Val Pro Met Gly Phe Ala Gln Lys Asp Gly Ser 50 55 60Tyr Ala Gly
Phe Asp Ile Asp Leu Ala Thr Ala Val Phe Glu Lys Tyr65 70 75 80Gly
Ile Thr Val Asn Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala 85 90
95Glu Leu Thr Lys Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala
100 105 110Thr Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr
Met Lys 115 120 125Asn Glu Gln Val Leu Val Thr Lys Lys Leu Ser Gly
Ile Thr Thr Ala 130 135 140Lys Asp Met Asn Gly Lys Thr Leu Gly Ala
Gln Ala Gly Ser Ser Gly145 150 155 160Tyr Ala Asp Phe Glu Ala Asn
Pro Glu Leu Leu Lys Asn Ile Val Ala 165 170 175Asn Lys Glu Ala Asn
Gln Tyr Gln Thr Phe Asn Glu Ala Leu Ile Asp 180 185 190Leu Lys Asn
Asp Arg Ile Asp Gly Leu Leu Ile Asp Arg Val Tyr Ala 195 200 205Asn
Tyr Tyr Leu Glu Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe 210 215
220Thr Val Gly Leu Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys
Glu225 230 235 240Asp Thr Thr Leu Val Lys Lys Ile Asn Glu Ala Phe
Ser Ser Leu Tyr 245 250 255Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln
Lys Trp Phe Gly Glu Asp 260 265 270Val Ala Thr Lys Glu Val Lys Glu
Gly Gln 275 280145274PRTStreptococcus pneumoniae 145Met Leu Val Leu
Val Ser Leu Met Thr Ala Leu Phe Leu Val Ala Cys1 5 10 15Gly Lys Asn
Ser Ser Glu Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln 20 25 30Ser Asn
Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr Phe Val Pro Met 35 40 45Gly
Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe Asp Ile Asp Leu 50 55
60Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val Asn Trp Gln Pro65
70 75 80Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys Gly Thr Ile
Asp 85 90 95Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu
Lys Val 100 105 110Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val
Leu Val Thr Lys 115 120 125Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp
Met Thr Gly Lys Thr Leu 130 135 140Gly Ala Gln Ala Gly Ser Ser Gly
Tyr Ala Asp Phe Glu Ala Asn Pro145 150 155 160Glu Ile Leu Lys Asn
Ile Val Ala Asn Lys Glu Ala Asn Gln Tyr Gln 165 170 175Thr Phe Asn
Glu Ala Leu Ile Asp Leu Lys Asn Asp Arg Ile Asp Gly 180 185 190Leu
Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu Gly 195 200
205Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu Glu Thr Glu Ala
210 215 220Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr Leu Val Lys
Lys Ile225 230 235 240Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly
Lys Phe Gln Glu Ile 245 250 255Ser Gln Lys Trp Phe Gly Glu Asp Val
Ala Thr Lys Glu Val Lys Glu 260 265 270Gly
Gln146278PRTStreptococcus pneumoniae 146Met Lys Lys Trp Ile Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala
Lys Asp Met Ala 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu
Lys Asp Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275147278PRTStreptococcus pneumoniae 147Met Lys Lys Trp Met Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Tyr Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Leu Ser Gly Ile Thr Thr Ala
Lys Asp Met Thr 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Glu Ile Leu
Lys Asn Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275148278PRTStreptococcus pneumoniae 148Met Lys Lys Trp Ile Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ser Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser His Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala
Lys Asp Met Ala 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu
Lys Asp Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275149278PRTStreptococcus pneumoniae 149Met Lys Lys Trp Met Leu Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ala Ser Glu Ile Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Gln Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala
Lys Asp Met Ala 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu
Lys Asp Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Thr Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275150278PRTStreptococcus pneumoniae 150Met Lys Lys Trp Met Phe Val
Leu Val Ser Leu Met Thr Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys
Asn Ala Ser Glu Thr Ser Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser
Asn Lys Ser Ile Thr Ile Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met
Gly Phe Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp
Leu Ala Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn
Trp Gln Pro Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90
95Gly Thr Ile Asp Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg
100 105 110Arg Glu Lys Val Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu
Gln Val 115 120 125Leu Val Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala
Lys Asp Met Ala 130 135 140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser
Ser Gly Tyr Ala Asp Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu
Lys Asp Ile Val Ala Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr
Phe Asn Glu Ala Leu Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp
Gly Leu Leu Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu
Ala Glu Gly Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215
220Glu Thr Glu Ala Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr
Leu225 230 235 240Val Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr
Lys Asp Gly Lys 245 250 255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly
Glu Asp Val Ala Thr Lys 260 265 270Glu Val Lys Glu Gly Gln
275151274PRTStreptococcus pneumoniae 151Met Leu Val Leu Val Ser Leu
Met Thr Ala Leu Phe Leu Val Ala Cys1 5 10 15Gly Lys Asn Ser Ser Glu
Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln 20 25 30Ser Asn Lys
Ser Ile Thr Ile Gly Phe Asp Ser Thr Phe Val Pro Met 35 40 45Gly Phe
Ala Gln Lys Asp Gly Ser Tyr Ala Gly Phe Asp Ile Asp Leu 50 55 60Ala
Thr Ala Val Phe Glu Lys Tyr Gly Ile Thr Val Asn Trp Gln Pro65 70 75
80Ile Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys Gly Thr Ile Asp
85 90 95Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys
Val 100 105 110Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val Leu
Val Thr Lys 115 120 125Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp Met
Thr Gly Lys Thr Leu 130 135 140Gly Ala Gln Ala Gly Ser Ser Gly Tyr
Ala Asp Phe Glu Ala Asn Pro145 150 155 160Glu Ile Leu Lys Asn Ile
Val Ala Asn Lys Glu Ala Asn Gln Tyr Gln 165 170 175Thr Phe Asn Glu
Ala Leu Ile Asp Leu Lys Asn Asp Arg Ile Asp Gly 180 185 190Leu Leu
Ile Asp Arg Val Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu Gly 195 200
205Val Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu Glu Thr Glu Ala
210 215 220Phe Ala Val Gly Ser Arg Lys Glu Asp Thr Thr Leu Val Lys
Lys Ile225 230 235 240Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly
Lys Phe Gln Glu Ile 245 250 255Ser Gln Lys Trp Phe Gly Glu Asp Val
Ala Thr Lys Glu Val Lys Glu 260 265 270Gly
Gln152274PRTStreptococcus pneumoniae 152Met Leu Val Leu Val Ser Leu
Met Thr Ala Leu Phe Leu Val Ala Cys1 5 10 15Gly Lys Asn Ser Ser Glu
Thr Ser Gly Asp Asn Trp Ser Lys Tyr Gln 20 25 30Ser Asn Lys Ser Ile
Thr Ile Gly Phe Asp Ser Thr Phe Val Pro Met 35 40 45Gly Phe Ala Gln
Lys Asp Gly Ser Tyr Ala Gly Phe Asp Ile Asp Leu 50 55 60Ala Thr Ala
Val Phe Glu Lys Tyr Gly Ile Thr Val Asn Trp Gln Pro65 70 75 80Ile
Asp Trp Asp Leu Lys Glu Ala Glu Leu Thr Lys Gly Thr Ile Asp 85 90
95Leu Ile Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg Arg Glu Lys Val
100 105 110Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val Leu Val
Thr Lys 115 120 125Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp Met Thr
Gly Lys Thr Leu 130 135 140Gly Ala Gln Ala Gly Ser Ser Gly Tyr Ala
Asp Phe Glu Ala Asn Pro145 150 155 160Glu Leu Leu Lys Asn Ile Val
Ala Asn Lys Glu Ala Asn Gln Tyr Gln 165 170 175Thr Phe Asn Glu Ala
Leu Ile Asp Leu Lys Asn Asp Arg Ile Asp Gly 180 185 190Leu Leu Ile
Asp Arg Val Tyr Ala Asn Tyr Tyr Leu Glu Ala Glu Gly 195 200 205Val
Leu Asn Asp Tyr Asn Val Phe Thr Val Gly Leu Glu Thr Glu Ala 210 215
220Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr Leu Val Lys Lys
Ile225 230 235 240Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys
Phe Gln Glu Ile 245 250 255Ser Gln Lys Trp Phe Gly Glu Asp Val Ala
Thr Lys Glu Val Lys Glu 260 265 270Gly Gln153277PRTStreptococcus
pneumoniae 153Met Lys Lys Trp Met Leu Val Leu Val Ser Leu Met Thr
Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys Asn Ser Ser Glu Thr Ser
Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser Asn Lys Ser Ile Thr Ile
Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met Gly Phe Ala Gln Lys Asp
Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp Leu Ala Thr Ala Val Phe
Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn Trp Gln Pro Ile Asp Trp
Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90 95Gly Thr Ile Asp Leu Ile
Trp Asn Gly Tyr Ser Ala Thr Tyr Glu Arg 100 105 110Arg Glu Lys Val
Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val 115 120 125Leu Val
Thr Lys Lys Leu Ser Gly Ile Thr Thr Ala Lys Asp Met Thr 130 135
140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp
Phe145 150 155 160Glu Ala Asn Pro Glu Ile Leu Lys Asn Ile Val Ala
Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr Phe Asn Glu Ala Leu
Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp Gly Leu Leu Ile Asp
Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu Ala Glu Gly Val Leu
Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215 220Glu Thr Glu Ala
Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr Leu225 230 235 240Val
Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys 245 250
255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly Glu Asp Val Ala Thr Lys
260 265 270Glu Val Lys Glu Gly 275154282PRTStreptococcus pneumoniae
154Met Lys Lys Trp Met Lys Lys Trp Met Leu Val Leu Val Ser Leu Met1
5 10 15Thr Ala Leu Phe Leu Val Ala Cys Gly Lys Asn Ala Ser Glu Thr
Ser 20 25 30Gly Asp Asn Trp Ser Lys Tyr Glu Ser Asn Lys Ser Ile Thr
Ile Gly 35 40 45Phe Asp Ser Thr Phe Val Pro Met Gly Phe Ala Gln Lys
Asp Gly Ser 50 55 60Tyr Ala Gly Phe Asp Ile Asp Leu Ala Thr Ala Val
Phe Glu Lys Tyr65 70 75 80Gly Ile Thr Val Asn Trp Gln Pro Ile Asp
Trp Asp Leu Lys Glu Ala 85 90 95Glu Leu Thr Lys Gly Thr Ile Asp Leu
Ile Trp Asn Gly Tyr Ser Ala 100 105 110Thr Asp Glu Arg Arg Glu Lys
Val Ala Phe Ser Asn Ser Tyr Met Lys 115 120 125Asn Glu Gln Val Leu
Val Thr Lys Lys Leu Ser Gly Ile Thr Thr Ala 130 135 140Lys Asp Met
Thr Gly Lys Ile Leu Gly Ala Gln Ala Gly Ser Ser Gly145 150 155
160Tyr Ala Asp Phe Glu Ala Asn Pro Glu Ile Leu Lys Asn Ile Val Ala
165 170 175Asn Lys Glu Ala Asn Gln Tyr Gln Thr Phe Asn Glu Ala Leu
Ile Asp 180 185 190Leu Lys Asn Asp Arg Ile Asp Gly Leu Leu Ile Asp
Arg Val Tyr Ala 195 200 205Asn Tyr Tyr Leu Glu Ala Glu Gly Val Leu
Asn Asp Tyr Asn Val Phe 210 215 220Thr Val Gly Leu Glu Ile Glu Ala
Phe Ala Val Gly Ala Arg Lys Glu225 230 235 240Asp Thr Thr Leu Val
Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr 245 250 255Lys Asp Gly
Lys Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly Glu Asp 260 265 270Val
Ala Thr Lys Glu Val Lys Glu Gly Gln 275 280155278PRTStreptococcus
pneumoniae 155Met Lys Lys Trp Met Leu Val Leu Val Ser Leu Met Thr
Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys Asn Ala Ser Glu Thr Ser
Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser Asn Lys Ser Ile Thr Ile
Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met Gly Phe Ala Gln Lys Asp
Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp Leu Ala Thr Ala Val Phe
Glu Lys Tyr Gly Ile Thr Val65 70 75 80Asn Trp Gln Pro Ile Asp Trp
Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90 95Gly Thr Ile Asp Leu Ile
Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg 100 105 110Arg Glu Lys Val
Ala Phe Ser Asn Ser Tyr Met Lys Asn Glu Gln Val 115 120 125Leu Val
Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp Met Ala 130 135
140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp
Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu Lys Asp Ile Val Ala
Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Ile Phe Asn Glu Ala Leu
Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp Gly Leu Leu Ile Asp
Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu Ala Glu Gly Val Leu
Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215 220Glu Thr Glu Ala
Phe Ala Val Gly Ala Arg Lys Glu Asp Ile Thr Leu225 230 235 240Val
Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys 245 250
255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly Glu Asp Val Ala Thr Lys
260 265 270Glu Val Lys Glu Gly Gln 275156278PRTStreptococcus
pneumoniae 156Met Lys Lys Trp Met Leu Val Leu Val Ser Leu Met Ile
Ala Leu Phe1 5 10 15Leu Val Ala Cys Gly Lys Asn Thr Ser Glu Thr Ser
Gly Asp Asn Trp 20 25 30Ser Lys Tyr Glu Ser Asn Lys Ser Ile Thr Ile
Gly Phe Asp Ser Thr 35 40 45Phe Val Pro Met Gly Phe Ala Gln Lys Asp
Gly Ser Tyr Ala Gly Phe 50 55 60Asp Ile Asp Leu Ala Thr Ala Val Phe
Glu Lys Tyr Gly Ile Thr Ile65 70 75 80Asn Trp Gln Pro Ile Asp Trp
Asp Leu Lys Glu Ala Glu Leu Thr Lys 85 90 95Gly Thr Ile Asp Leu Ile
Trp Asn Gly Tyr Ser Ala Thr Asp Glu Arg 100 105 110Arg Glu Lys Val
Ala Phe Ser His Ser Tyr Met Lys Asn Glu Gln Val 115 120 125Leu Val
Thr Lys Lys Ser Ser Gly Ile Thr Thr Ala Lys Asp Met Ala 130 135
140Gly Lys Thr Leu Gly Ala Gln Ala Gly Ser Ser Gly Tyr Ala Asp
Phe145 150 155 160Glu Ala Asn Pro Ala Ile Leu Lys Asp Ile Val Ala
Asn Lys Glu Ala 165 170 175Asn Gln Tyr Gln Thr Phe Asn Glu Ala Leu
Ile Asp Leu Lys Asn Asp 180 185 190Arg Ile Asp Gly Leu Leu Ile Asp
Arg Val Tyr Ala Asn Tyr Tyr Leu 195 200 205Glu Ala Glu Gly Val Leu
Asn Asp Tyr Asn Val Phe Thr Val Gly Leu 210 215 220Glu Thr Glu Ala
Phe Ala Val Gly Ala Arg Lys Glu Asp Thr Thr Leu225 230 235 240Val
Lys Lys Ile Asn Glu Ala Phe Ser Ser Leu Tyr Lys Asp Gly Lys 245 250
255Phe Gln Glu Ile Ser Gln Lys Trp Phe Gly Glu Asp Val Ala Thr Lys
260 265 270Glu Val Lys Glu Gly Gln 275157286PRTStreptococcus
pneumoniae 157Met Lys Lys Trp Met Lys Lys Trp Met Lys Lys Trp Met
Leu Val Leu1 5 10 15Val Ser Leu Met Thr Ala Leu Phe Leu Val Ala Cys
Gly Lys Asn Ala 20 25 30Ser Glu Thr Ser Gly Asp Asn Trp Ser Lys Tyr
Glu Ser Asn Lys Ser 35 40 45Ile Thr Ile Gly Phe Asp Ser Thr Phe Val
Pro Met Gly Phe Ala Gln 50 55 60Lys Asp Gly Ser Tyr Ala Gly Phe Asp
Ile Asp Leu Ala Thr Ala Val65 70 75 80Phe Glu Lys Tyr Gly Ile Thr
Val Asn Trp Gln Pro Ile Asp Trp Asp 85 90 95Leu Lys Glu Ala Glu Leu
Thr Lys Gly Thr Ile Asp Leu Ile Trp Asn 100 105 110Gly Tyr Ser Ala
Thr Asp Glu Arg Arg Glu Lys Val Ala Phe Ser Asn 115 120 125Ser Tyr
Met Lys Asn Glu Gln Val Leu Val Thr Lys Lys Leu Ser Gly 130 135
140Ile Thr Thr Ala Lys Asp Met Thr Gly Lys Ile Leu Gly Ala Gln
Ala145 150 155 160Gly Ser Ser Gly Tyr Ala Asp Phe Glu Ala Asn Pro
Glu Ile Leu Lys 165 170 175Asn Ile Val Ala Asn Lys Glu Ala Asn Gln
Tyr Gln Thr Phe Asn Glu 180 185 190Ala Leu Ile Asp Leu Lys Asn Asp
Arg Ile Asp Gly Leu Leu Ile Asp 195 200 205Arg Val Tyr Ala Asn Tyr
Tyr Leu Glu Ala Glu Gly Val Leu Asn Asp 210 215 220Tyr Asn Val Phe
Thr Val Gly Leu Glu Ile Glu Ala Phe Ala Val Gly225 230 235 240Ala
Arg Lys Glu Asp Thr Thr Leu Val Lys Lys Ile Asn Glu Ala Phe 245 250
255Ser Ser Leu Tyr Lys Asp Gly Lys Phe Gln Glu Ile Ser Gln Lys Trp
260 265 270Phe Gly Glu Asp Val Ala Thr Lys Glu Val Lys Glu Gly Gln
275 280 285
References
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.