Compositions And Methods Comprising Protease Variants
Poulose; Ayrookaran J. ; et al.
U.S. patent application number 16/693876 was filed with the patent office on 2020-03-12 for compositions and methods comprising protease variants. The applicant listed for this patent is DANISCO US INC. Invention is credited to Joshua Roy Basler, Luis G. Cascao-Pereira, David A. Estell, James T. Kellis, JR., Alexander Pisarchik, Ayrookaran J. Poulose, Daniel Esteban Torres Pazmino.
| Application Number | 20200080069 16/693876 |
| Document ID | / |
| Family ID | 44146173 |
| Filed Date | 2020-03-12 |


View All Diagrams
| United States Patent Application | 20200080069 |
| Kind Code | A1 |
| Poulose; Ayrookaran J. ; et al. | March 12, 2020 |
COMPOSITIONS AND METHODS COMPRISING PROTEASE VARIANTS
Abstract
The present invention provides protease variants, compositions comprising protease variants, and methods of using such protease variants and compositions.
| Inventors: | Poulose; Ayrookaran J.; (Belmont, CA) ; Basler; Joshua Roy; (Palo Alto, CA) ; Cascao-Pereira; Luis G.; (Redwood City, CA) ; Kellis, JR.; James T.; (Palo Alto, CA) ; Pisarchik; Alexander; (Palo Alto, CA) ; Torres Pazmino; Daniel Esteban; (Leiden, NL) ; Estell; David A.; (Palo Alto, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 44146173 | ||||||||||
| Appl. No.: | 16/693876 | ||||||||||
| Filed: | November 25, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14843833 | Sep 2, 2015 | |||
| 16693876 | ||||
| 14225292 | Mar 25, 2014 | 9157052 | ||
| 14843833 | ||||
| 12963930 | Dec 9, 2010 | 8728790 | ||
| 14225292 | ||||
| 61392373 | Oct 12, 2010 | |||
| 61285127 | Dec 9, 2009 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 9/52 20130101; C12Y 304/21062 20130101; C11D 3/386 20130101; C12N 9/64 20130101; C12N 9/54 20130101 |
| International Class: | C12N 9/54 20060101 C12N009/54; C11D 3/386 20060101 C11D003/386; C12N 9/64 20060101 C12N009/64; C12N 9/52 20060101 C12N009/52 |
Claims
1. An isolated protease variant of a parent protease, the variant comprising an amino acid sequence comprising three amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2.
2. The variant of claim 1, wherein the variant amino acid sequence comprises four amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q.
3. The variant of claim 1, wherein the variant amino acid sequence comprises five amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q.
4. The variant of claim 1, wherein the variant amino acid sequence comprises six amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q.
5. The variant of any preceding claim, wherein the variant amino acid sequence further comprises amino acid substitution X097A.
6. The variant of any preceding claim, wherein the variant amino acid sequence comprises amino acid substitutions X024G/R+X053G+X078N+X101N+X128A/S+X217L/Q or X097A+X128A/S+X217L/Q.
7. The variant of any preceding claim, wherein the variant amino acid sequence comprises amino acid substitutions S024G/R+S053G+S078N+S101N+G128A/S+Y217L/Q or G097A+G128A/S+Y217L/Q.
8. The variant of claim 7, wherein the variant amino acid sequence comprises amino acid substitutions S024G+S053G+S078N+S101N+G128S+Y217Q or S024G+S053G+S078N+S101N+G128A+Y217Q, and optionally further comprises a substitution selected from the group consisting of N109G, N076D, S033T, N243V, S248A, A088T, and S063G.
9. The variant of claim 7, wherein the variant amino acid sequence further comprises a set of amino acid substitutions selected from the group consisting of: A088T+N109G+A116T+G131H+N243V+L257G, S033T+N076D, S009T+N109G+K141R+N243V, S162G+K256R, N109G+A116T, N109G+L257G, S162G+L257G, N061G+N109G+N243V, N109G+N243V+S248A, S033T+N076D+N109G+N218S+N243V+S248N+K256R, N109G+A116T+N243V+K256R, A088T+N109G+A116T+G131H+N243V, A088T+N109G, N109G+N243V, T158S+L257G, N061S+N109G+N243V, P040A+N109G+N243V+S248N+K256R, S009T+S018T+Y021N+N109G+K141R, A088T+N109G+A116T+T158S+N243V+K256R, A088T+N109G+A116T+T158S+N218S+L257G, N109G+K256R, N109G+N243V+K256R, S063G+K256R, S063G+N109G, S063G, S063G+N076D, S033T+N076D+N218S, and N076D+N218S.
10. The variant of claim 7, wherein the variant amino acid sequence comprises amino acid substitutions S024G+S053G+S078N+S101N+G128A+Y217Q and further comprises a set of amino acid substitutions selected from the group consisting of: A088T+N109G+A116T+G131H+N243V+L257G, S033T+N076D, S009T+N109G+A128S+K141R+N243V, S162G+K256R, N109G+A116T, N109G+L257G, S162G+L257G, N061G+N109G+N243V, N109G+A128S+N243V+S248A, S033T+N076D+N109G+A128S+N218S+N243V+S248N+K256R, N109G+A116T+N243V+K256R, A088T+N109G+A116T+G131H+N243V, A088T+N109G, N109G+N243V, T158S+L257G, N061S+N109G+N243V, P040A+N109G+A128S+N243V+S248N+K256R, S009T+S018T+Y021N+N109G+A128S+K141R, A088T+N109G+A116T+T158S+N243V+K256R, A088T+N109G+A116T+T158S+N218S+L257G, N109G+K256R, N109G+A128S+N243V+K256R, S063G+K256R, S063G+N109G, S063G+A128S, S063G+N076D, S033T+N076D+A128S+N218S, and N076D+N218S.
11. The variant of any preceding claim, wherein the variant amino acid sequence has at least 80% sequence identity to the amino acid sequence of SEQ ID NO:2 or SEQ ID NO:6.
12. The variant of any preceding claim, wherein the variant amino acid sequence has at least 85% sequence identity to the amino acid sequence of SEQ ID NO:2 or SEQ ID NO:6.
13. The variant of any of claims 1-12, wherein the variant has enhanced proteolytic activity and/or cleaning activity compared to the parent protease or enhanced proteolytic activity and/or cleaning activity compared to the proteolytic activity of the BPN' protease having the sequence of SEQ ID NO:2.
14. The variant of any of claims 1-12, wherein the variant has enhanced proteolytic activity compared to the proteolytic activity of the protease having the sequence of SEQ ID NO:4.
15. The variant of any of claims 1-12, wherein the variant has enhanced proteolytic activity compared to the proteolytic activity of the protease having the sequence of SEQ ID NO:6.
16. The variant of any preceding claim, wherein the parent protease is a subtilisin protease.
17. The variant of claim 16, wherein the parent protease has at least 80% sequence identity to the B. amyloliquefaciens subtilisin protease BPN' having the amino acid sequence of SEQ ID NO:2.
18. The variant of claim 16, wherein the parent protease has at least 85% sequence identity to the amino acid sequence of SEQ ID NO:2.
19. The variant of claim 1, wherein the variant comprises the amino acid sequence of SEQ ID NO:6.
20. An isolated protease variant of a parent protease, the protease variant having proteolytic activity and comprising an amino acid sequence which comprises an alteration at one or more amino acid positions corresponding to amino acid positions of SEQ ID NO:2 selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217, wherein the at least one alteration is independently (i) an insertion of one or more amino acid residues upstream or downstream of the amino acid residue which occupies the position, (ii) a deletion of the amino acid residue which occupies the position, or (iii) a substitution of the amino acid residue which occupies the position with a different amino acid residue, wherein each amino acid position is numbered by correspondence with an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin protease BPN' set forth in SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2.
21. The variant of claim 20, wherein the variant comprises an amino acid sequence comprising an alteration at two amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
22. The variant of claim 21, wherein the variant comprises an amino acid sequence comprising an alteration at three amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
23. The variant of claim 22, wherein the variant comprises an amino acid sequence comprising an alteration at four amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
24. The variant of claim 23, wherein the variant comprises an amino acid sequence comprising an alteration at five amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
25. The variant of claim 24, wherein the variant comprises an amino acid sequence comprising an alteration at six amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
26. The variant of claim 25, wherein the variant comprises an amino acid sequence comprising an alteration at each of the amino acid positions corresponding to positions of SEQ ID NO:2 selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
27. The variant of any of claims 20-26, wherein the variant comprises an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at a position selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
28. The variant of claim 27, wherein the variant comprises an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of two positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
29. The variant of claim 28, wherein the variant comprises an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of four positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
30. The variant of claim 29, wherein the variant comprises an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of five positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
31. The variant of claim 30, wherein the variant comprises an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of six positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
32. The variant of claim 31, wherein the variant comprises an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of positions 24, 53, 78, 101, 128, and 217, wherein each position is numbered by correspondence with a position in SEQ ID NO:2.
33. The variant of any of claims 20-32, wherein the variant comprises an amino acid sequence comprising at least one amino acid substitution selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L.
34. The variant of claim 33, wherein the variant comprises an amino acid sequence comprising at least two amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L.
35. The variant of claim 34, wherein the variant comprises an amino acid sequence comprising at least three amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L.
36. The variant of claim 35, wherein the variant comprises an amino acid sequence comprising at least four amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L.
37. The variant of claim 36, wherein the variant comprises an amino acid sequence comprising at least five amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L.
38. The variant of claim 37, wherein the variant comprises an amino acid sequence comprising at least six amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L.
39. The variant of any of claims 20-38, wherein the variant comprises an amino acid sequence comprising a set of substitutions selected from the group consisting of: (a) X128A/S and/or X217L/Q, (b) G128A/S and/or Y217L/Q, and (c) G097A, G128A/S, and Y217L/Q.
40. The variant of any of claims 20-39, wherein the variant comprises an amino acid sequence comprising amino acid substitutions X024G/R+X053G+X078N+X101N+X128A/S+X217Q/L.
41. The variant of any of claims 20-40, wherein the variant comprises an amino acid sequence having at least 80% sequence identity to the amino acid sequence of SEQ ID NO:2.
42. The variant of claim 41, wherein the variant comprises an amino acid sequence having at least 85% sequence identity to SEQ ID NO:2.
43. The variant of any of claims 20-42, wherein the variant amino acid sequence comprises at least one amino acid substitution selected from the group of S024G, S053G, S078N, S101N, G128A/S, and Y217Q.
44. The variant of any of claims 20-43, wherein the variant amino acid sequence comprises amino acid substitutions S024G+S053G+S078N+S101N+G128A+Y217Q.
45. The variant of any of claims 20-44, wherein the variant has enhanced proteolytic activity has enhanced proteolytic activity and/or cleaning activity compared to the parent protease or enhanced proteolytic activity and/or cleaning activity compared to the proteolytic activity of the protease set forth in SEQ ID NO:2.
46. The variant of any of claims 20-44, wherein the variant has enhanced proteolytic activity compared to the proteolytic activity of the protease set forth in SEQ ID NO:4.
47. The variant of any of claims 20-44, wherein the variant has enhanced proteolytic activity compared to the proteolytic activity of the protease set forth in SEQ ID NO:6.
48. The variant of any of claims 20-47, wherein the parent protease is a subtilisin protease.
49. The variant of claim 48, wherein the parent protease has at least 80% sequence identity to the B. amyloliquefaciens subtilisin protease BPN' having the amino acid sequence of SEQ ID NO:2.
50. The variant of claim 49, wherein the parent protease has at least 85% sequence identity to the amino acid sequence of SEQ ID NO:2.
51. An isolated protease variant of a parent protease, wherein (a) the protease variant comprises an amino acid sequence (i) having at least 90% identity to the sequence of SEQ ID NO:2 and (ii) comprising a substitution of glycine at positions 24 and 53, a substitution of asparagine at positions 78 and 101, a substitution of alanine or serine at position 128, and a substitution of glutamine at position 217; (b) the parent protease has at least 90% sequence identity to SEQ ID NO:2; (c) each amino acid position of the variant is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2; and (d) the protease variant has increased proteolytic activity and/or cleaning activity relative to the parent protease.
52. The variant according to any of the preceding claims, wherein the variant is a mature form.
53. An isolated polypeptide having protease activity, said polypeptide comprising an amino acid sequence having at least 85% sequence identity to a polypeptide sequence selected from the group consisting of: a) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+G131H+N243V+L2- 57G; b) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S033T+N076D; c) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S009T+N109G+K141R+N243V; d) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S162G+K256R; e) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+A116T; f) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+L257G; g) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S162G+L257G; h) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N061G+N109G+N243V; i) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+N109G+N243V+S248A; j) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S033T+N076D+N109G+N218S+N243V+S2- 48N+K256R; k) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+A116T+N243V+K256R; l) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+G131H+N243V; m) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G; n) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+N243V; o) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+T158S+L257G; p) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N061S+N109G+N243V; q) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+P040A+N109G+N243V+S248N+K256R; r) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S009T+S018T+Y021N+N109G+K141R- ; s) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+T158S+N243- V+K256R; t) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+T158S+N218S+L2- 57G; u) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+K256R; v) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+N109G+N243V+K256R; w) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+K256R; x) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+N109G; y) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S063G; z) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+N076D; aa) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S033T+N076D+N218S; bb) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N076D+N218S; and cc) BPN-S024G+S053G+S078N+S101N+G128A+Y217Q, wherein each amino acid position of the variant is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2.
54. An isolated nucleic acid comprising a polynucleotide sequence encoding a variant of any of claims 1-52 or a polypeptide of claim 53, or a complementary polynucleotide sequence thereof.
55. An isolated nucleic acid comprising a polynucleotide sequence having at least 80% sequence identity to the polynucleotide sequence set forth in SEQ ID NO:3 or SEQ ID NO:5, or a complementary polynucleotide sequence thereof.
56. An expression vector comprising at least one nucleic acid of claim 54 or 55.
57. The expression vector of claim 56, wherein the at least one nucleic acid is operably linked to a promoter.
58. A recombinant host cell comprising: (a) a nucleic acid of claim 64 or 65, or (b) an expression vector of claim 56 or 57.
59. The recombinant host cell of claim 58, wherein the host cell is a bacterial cell.
60. The recombinant host cell of claim 59, wherein the host cell is a Bacillus cell.
61. The recombinant host cell of claim 70, wherein the host cell is a Bacillus subtilis cell.
62. A cell culture comprising: (a) a nucleic acid of claim 54 or 55, or (b) an expression vector of claim 56 or 57.
63. A method of producing a protease variant, the method comprising cultivating a recombinant host cell of any of claims 58-61 under conditions conducive to produce the variant.
64. The method of claim 63, further comprising recovering the variant from the cell culture.
65. A method of producing a protease variant, the method comprising: (a) introducing the recombinant expression vector of claim 56 or 57 into a population of cells; and (b) culturing the cells in a culture medium under conditions conducive to produce the protease variant encoded by the expression vector.
66. The method of claim 65, further comprising: (c) isolating or recovering the variant from the cells or from the culture medium.
67. A composition comprising a variant of any of claims 1-52 or a polypeptide of claim 53, which composition is not a fabric and home care product.
68. The composition of claim 67, comprising at least one adjunct ingredient or carrier.
69. The composition of claim 67 or 68, wherein the composition comprises an additional enzyme.
70. The composition according to claim 69, wherein the additional enzyme is selected from the group consisting of a hemicellulase, cellulase, amylase, peroxidase, protease, xylanase, lipase, phospholipase, esterase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, -glucanase, arabinosidase, hyaluronidase, chondroitinase, and laccase.
71. The composition of any of claims 67-70, wherein the composition is a detergent composition for cleaning a contact lens.
72. The composition of any of claims 67-70, wherein the composition is a cleaning composition useful in a personal care application.
73. The composition of any of claims 67-72, further comprising at least one builder and/or at least one surfactant.
74. A method for cleaning an item or surface in need of cleaning, the method comprising contacting the item or surface with a variant of any of claims 1-52, a polypeptide of claim 53, or a composition of any of claims 67-73.
75. The method of claim 74, further comprising rinsing the item or surface with water.
76. A method for cleaning a surface or object, the method comprising contacting at least a portion of the item or surface to be cleaned with a variant of any of claims 1-52, a polypeptide of claim 53, or a composition of any of claims 67-73 for a sufficient time and/or under conditions sufficient or effective to clean or wash the item or surface to a desired degree, and optionally comprising rinsing the item or surface with water.
77. A method of treating and/or cleaning a surface or fabric comprising the steps of optionally washing and/or rinsing said surface or fabric, contacting said surface or fabric with a variant of any of claims 1-52, a polypeptide of claim 52, or a composition of any of claims 67-73, then optionally washing and/or rinsing said surface or fabric.
78. The variant of any of claims 1-52, wherein the parent protease is a subtilisin protease, and the variant is a subtilisin protease variant having improved wash performance or improved cleaning performance in a detergent as compared to that of the parent subtilisin protease, or the polypeptide of claim 53, wherein the polypeptide has improved wash performance or improved cleaning performance in a detergent as compared to that of the protease of SEQ ID NO:2, SEQ ID NO:4 or SEQ ID NO:6.
Description
CROSS-REFERENCES TO RELATED APPLICATIONS
[0001] This application is continuation of U.S. patent application Ser. No. 14/843,833, filed Sep. 2, 2015, which is a continuation of U.S. patent application Ser. No. 14/225,292, filed Mar. 25, 2014, which is a divisional of U.S. patent application Ser. No. 12/963,930, filed Dec. 9, 2010, now U.S. Pat. No. 8,728,790, which claims priority to and benefit of U.S. Provisional Patent Application No. 61/285,127, filed on Dec. 9, 2009, and U.S. Provisional Patent Application No. 61/392,373, filed on Oct. 12, 2010, the disclosures of which are each incorporated herein by reference in their entirety for all purposes.
SEQUENCE LISTING
[0002] The sequence listing submitted via EFS, in compliance with 37 C.F.R. .sctn. 1.52(e), is incorporated herein by reference. The sequence listing text file submitted via EFS contains the file "20191125_NB31488USCNT2_SeqLst.txt" created on Nov. 25, 2019 which is 258 KB in size.
FIELD OF THE INVENTION
[0003] The present invention provides protease variants, compositions comprising protease variants, and methods of using such protease variants and compositions thereof.
BACKGROUND OF THE INVENTION
[0004] Although proteases have long been known in the art of industrial enzymes, there remains a need for engineered proteases that are suitable for particular conditions and uses. The present invention fills these and other needs.
SUMMARY OF THE INVENTION
[0005] In a first aspect, the invention provides an isolated protease variant of a parent protease enzyme, the protease variant having proteolytic activity and comprising an amino acid sequence which comprises an alteration at one or more amino acid positions corresponding to amino acid positions of SEQ ID NO:2 selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217, wherein the at least one alteration is independently (i) an insertion of one or more amino acid residues upstream or downstream of the amino acid residue which occupies the position, (ii) a deletion of the amino acid residue which occupies the position, or (iii) a substitution of the amino acid residue which occupies the position with a different amino acid residue, wherein each amino acid position is numbered by correspondence with an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin protease BPN' set forth in SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2.
[0006] In a second aspect, the invention provides an isolated protease variant of a parent protease, the variant comprising an amino acid sequence comprising three amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2.
[0007] In a third aspect, the invention provides an isolated polypeptide having protease activity, said polypeptide comprising an amino acid sequence having at least 85% sequence identity to a polypeptide sequence selected from the group consisting of:
a) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+G131H+N243V+- L257G;
b) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S033T+N076D;
c) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S009T+N109G+K141R+N243V;
d) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S162G+K256R;
e) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+A116T;
f) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+L257G;
g) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S162G+L257G;
h) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N061G+N109G+N243V;
i) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+N109G+N243V+S248A;
j) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S033T+N076D+N109G+N218S+N243V+- S248N+K256R;
k) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+A116T+N243V+K256R;
l) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+G131H+N243V;
m) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G;
n) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+N243V;
o) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+T158S+L257G;
p) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N061S+N109G+N243V;
q) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+P040A+N109G+N243V+S248N+K256R;
r) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S009T+S018T+Y021N+N109G+K141R;
s) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+T158S+N243V+- K256R;
t) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+T158S+N218S+- L257G;
u) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+K256R;
v) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+N109G+N243V+K256R;
w) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+K256R;
x) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+N109G;
y) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S063G;
z) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+N076D;
aa) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S033T+N076D+N218S;
bb) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N076D+N218S; and
[0008] cc) BPN-S024G+S053G+S078N+S101N+G128A+Y217Q, wherein each amino acid position of the variant is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2.
[0009] In a fourth aspect, the invention provides an isolated polypeptide having protease activity selected from the group consisting of: (a) a polypeptide comprising an amino acid sequence having at least 95%, 96%, 97%, 98%, or 99% sequence identity to the polypeptide sequence of SEQ ID NO:6; (b) a polypeptide encoded by a polynucleotide that hybridizes under at least high stringency conditions with (i) the polynucleotide sequence of SEQ ID NO:5 or (ii) a complementary polynucleotide sequence of (i); and (c) a polypeptide encoded by a polynucleotide comprising a polynucleotide sequence having at least 95% sequence identity to the polynucleotide sequence of SEQ ID NO:5.
[0010] In a fifth aspect, the invention provides an isolated protease variant of a parent protease, wherein: (a) the variant comprises an amino acid sequence having no more than 20, 15, or 10 alterations relative to the parent protease, wherein (i) the alterations are independently selected from an insertion, a deletion, or a substitution, and (ii) the alterations include a substitution of glycine at positions 24 and 53, a substitution of asparagine at positions 78 and 101, a substitution of alanine or serine at position 128, and a substitution of glutamine at position 217, (b) the parent protease has at least 90% sequence identity to SEQ ID NO:2, (c) the amino acid sequence of SEQ ID NO:2 is used for determining position numbering; and (d) the variant has increased proteolytic activity relative to the parent protease, wherein each amino acid position is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2.
[0011] In a sixth aspect, the invention provides an isolated protease variant of a parent protease, wherein (a) the variant comprises an amino acid sequence (i) having at least 85% identity to the sequence of SEQ ID NO:2 and (ii) comprising a substitution of glycine at positions 24 and 53, a substitution of asparagine at positions 78 and 101, a substitution of alanine or serine at position 128, and a substitution of glutamine at position 217; (b) the parent protease has at least 85% sequence identity to SEQ ID NO:2; (c) each amino acid position of the variant is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2; and (d) the variant has increased proteolytic activity relative to the parent protease.
[0012] In another aspect, the invention provides an isolated or recombinant nucleic acid comprising a polynucleotide sequence encoding at least one polypeptide variant (e.g., protease variant) of the invention, or a complementary polynucleotide sequence thereof.
[0013] In another aspect, the invention provides an isolated or recombinant nucleic acid comprising a polynucleotide sequence having at least 80% sequence identity to the polynucleotide sequence set forth in SEQ ID NO:3 or SEQ ID NO:5, or a complementary polynucleotide sequence thereof.
[0014] In another aspect, the invention provides an expression vector comprising at least one nucleic acid of the invention. Also provided is a recombinant host cell or cell culture comprising at least one nucleic acid or an expression vector of the invention.
[0015] In another aspect, the invention provides a method of producing at least one polypeptide (e.g., protease variant) of the invention, the method comprising: (a) introducing a recombinant expression vector of the invention which encodes a polypeptide (e.g., protease variant) of the invention into a population of cells; (b) culturing the cells in a culture medium under conditions conducive to produce the polypeptide (e.g., protease variant) encoded by the expression vector; and optionally (c) isolating or recovering the variant from the cells or from the culture medium.
[0016] In another aspect, the invention provides a composition comprising at least one protease variant or polypeptide of the invention, optionally in combination with another enzyme. Such composition may comprise an adjunct ingredient, such as a surfactant and/or builder, or a carrier. Such composition may be a cleaning composition or a detergent composition and may be useful in cleaning methods described elsewhere herein. Such composition may be a fabric and home care product or such composition may not be a fabric and home care product.
[0017] In another aspect, the invention provides a method for cleaning an item, object, or surface in need of cleaning, the method comprising contacting the item, object, or surface with a polypeptide or protease variant of the invention or a composition of the invention, and optionally rinsing the item, object, or surface with water.
[0018] In another aspect, the invention provides a method for cleaning an item or surface (e.g., hard surface), the method comprising contacting at least a portion of the item or surface (e.g., hard surface) to be cleaned with a polypeptide or protease variant of the invention or a composition of the invention for a sufficient time and/or under conditions sufficient or effective to clean or wash the item or surface (e.g., hard surface) to a desired degree, and optionally comprising rinsing the item or surface (e.g., hard surface) with water.
[0019] Other aspects of the invention are described below.
BRIEF DESCRIPTION OF THE DRAWINGS
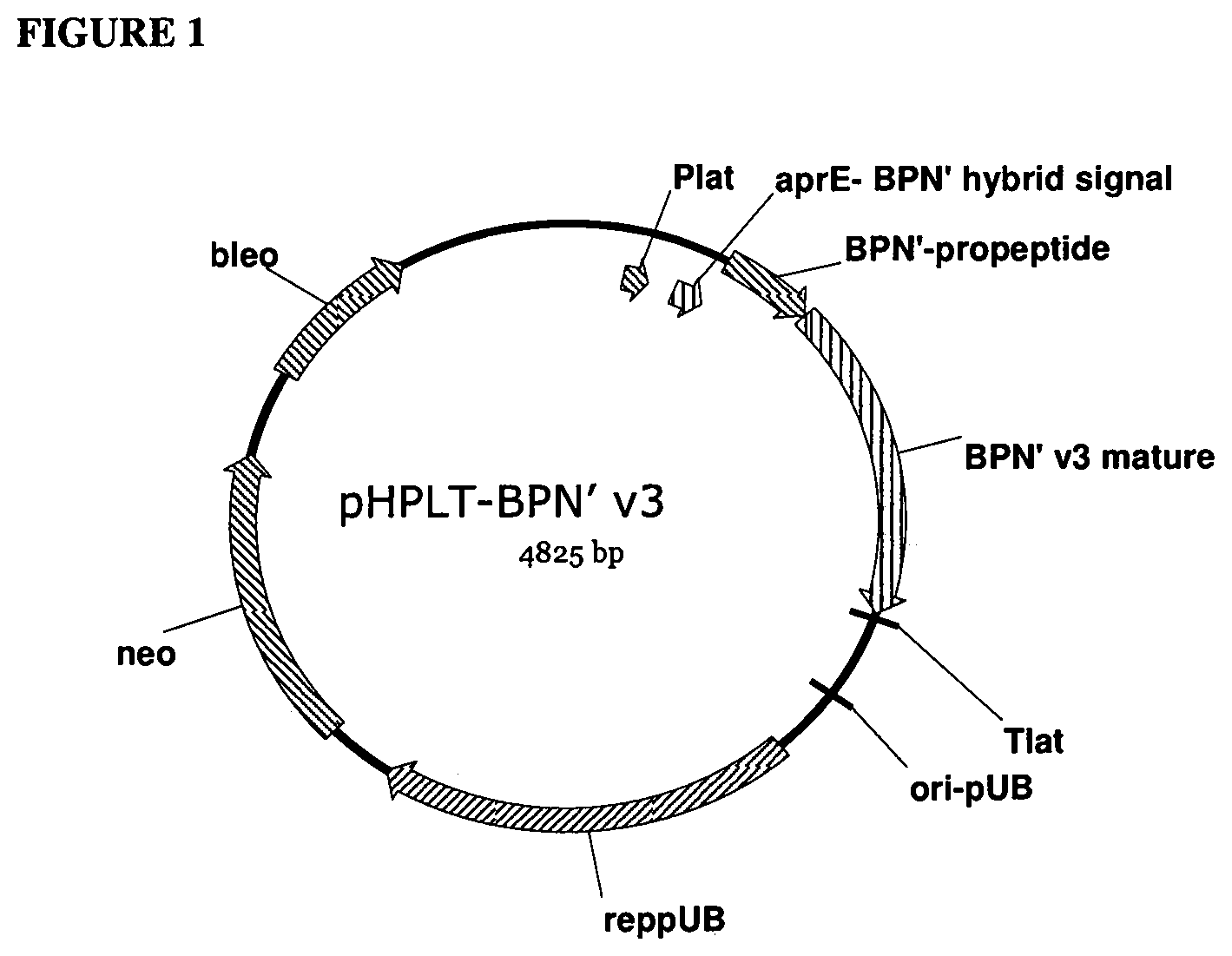
[0020] FIG. 1 provides a plasmid map of pHPLT-BPN'-v3.
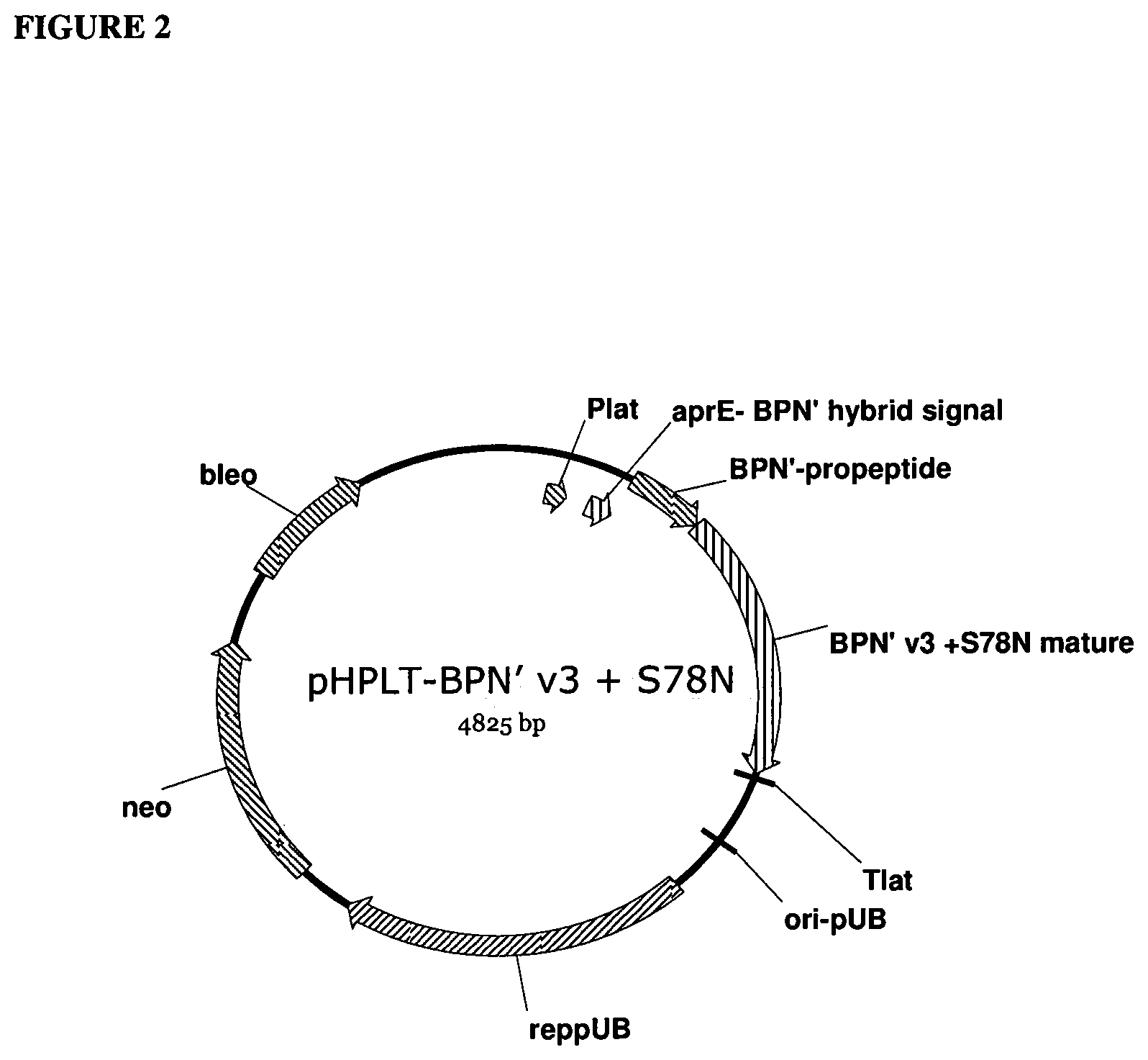
[0021] FIG. 2 provides a plasmid map of pHPLT-BPN'-v3+S78N.
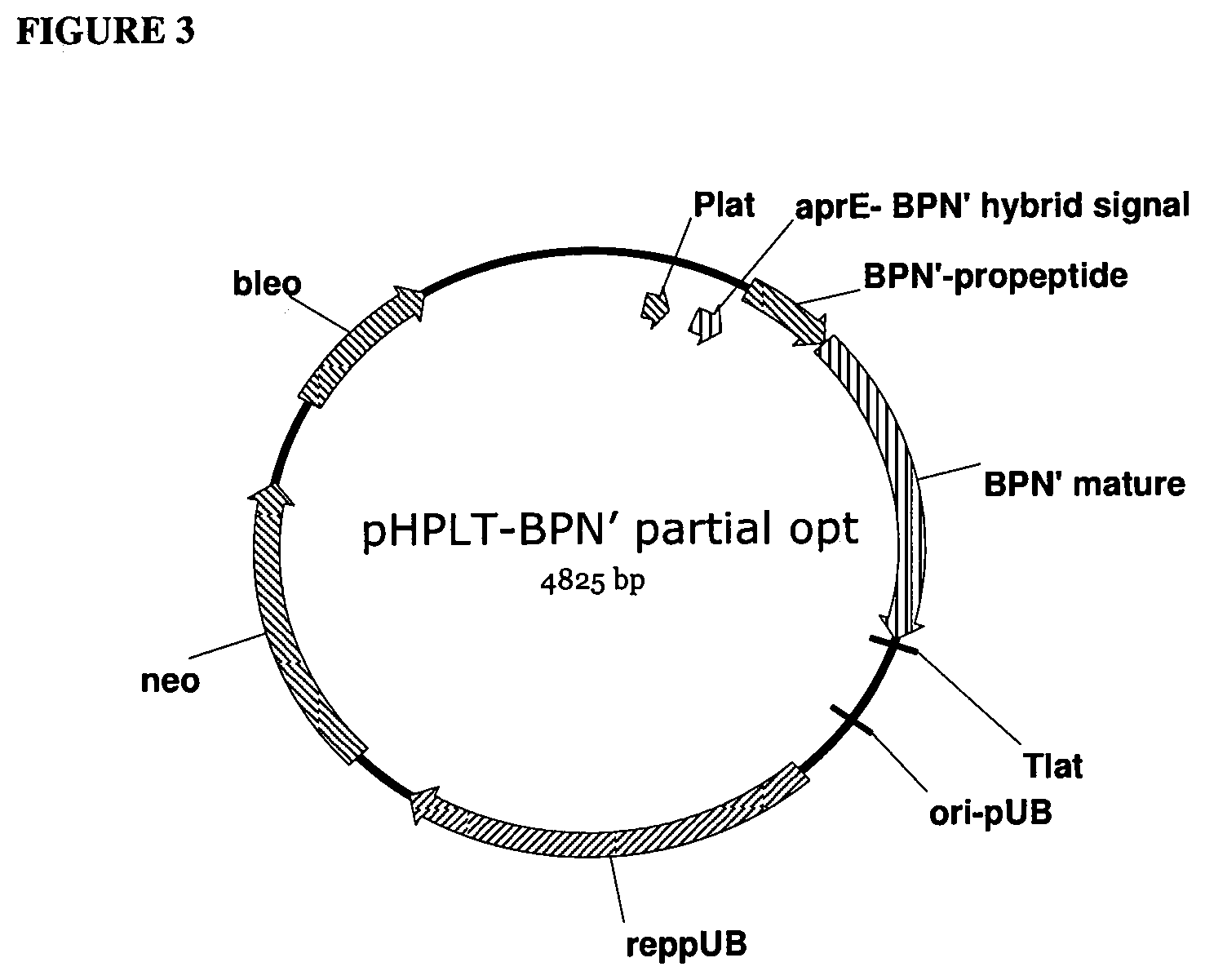
[0022] FIG. 3 provides a plasmid map of pHPLT-BPN' partial opt.
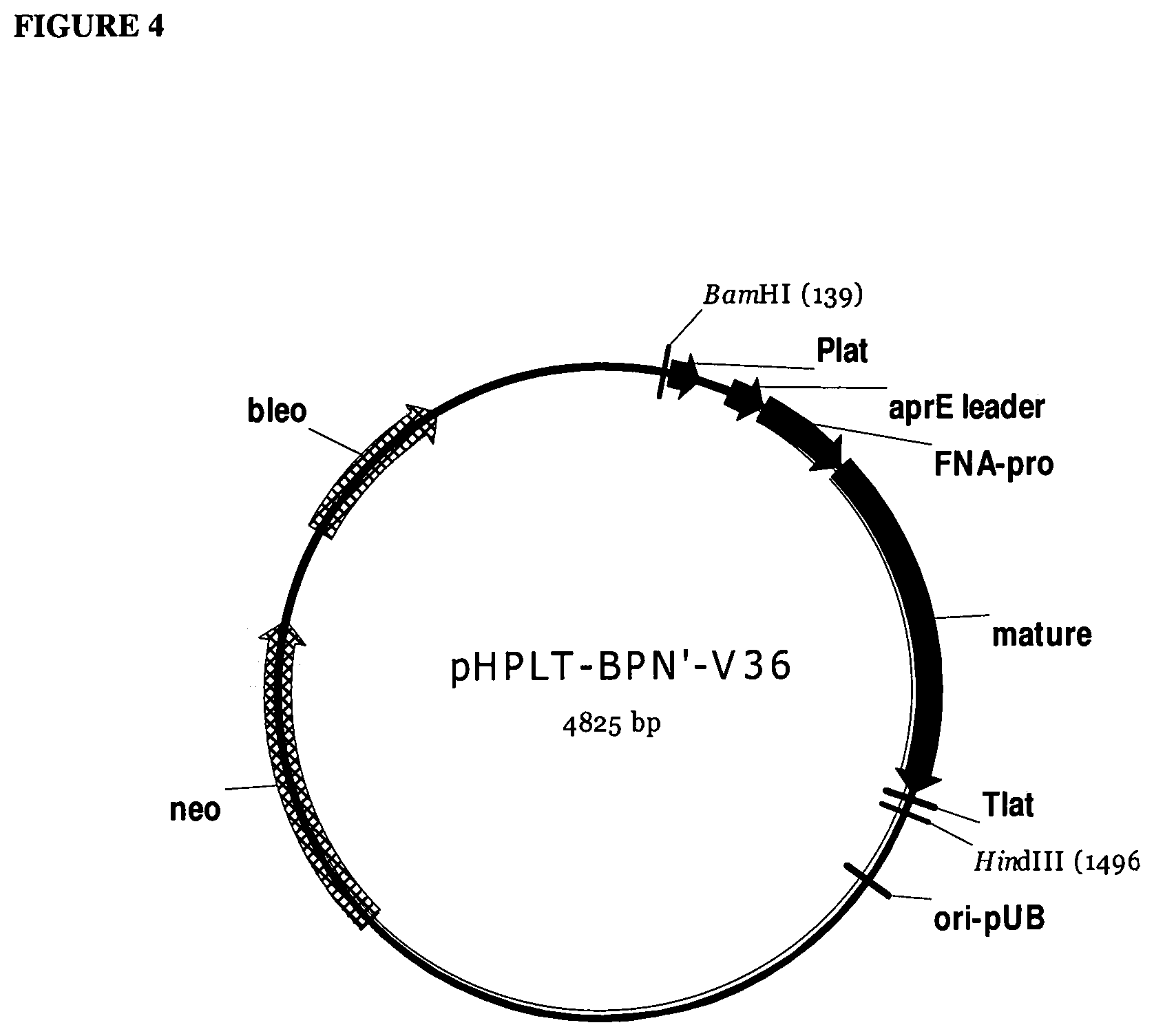
[0023] FIG. 4 provides a plasmid map of pHPLT-BPN'-v36.
[0024] FIG. 5 provides an alignment of the mature reference subtilisin proteases including: BPN' (SEQ ID NO:2) and GG36 (SEQ ID NO:755). Each amino acid position of each protease variant described herein, including each cold water protease variant, is numbered according to the numbering of the corresponding amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin protease BPN' (SEQ ID NO:2), as shown in FIG. 5, as determined by alignment of the protease variant amino acid sequence with the Bacillus amyloliquefaciens subtilisin protease BPN' amino acid sequence. Thus, unless otherwise specified herein, substitution positions are given in relationship to BPN'.
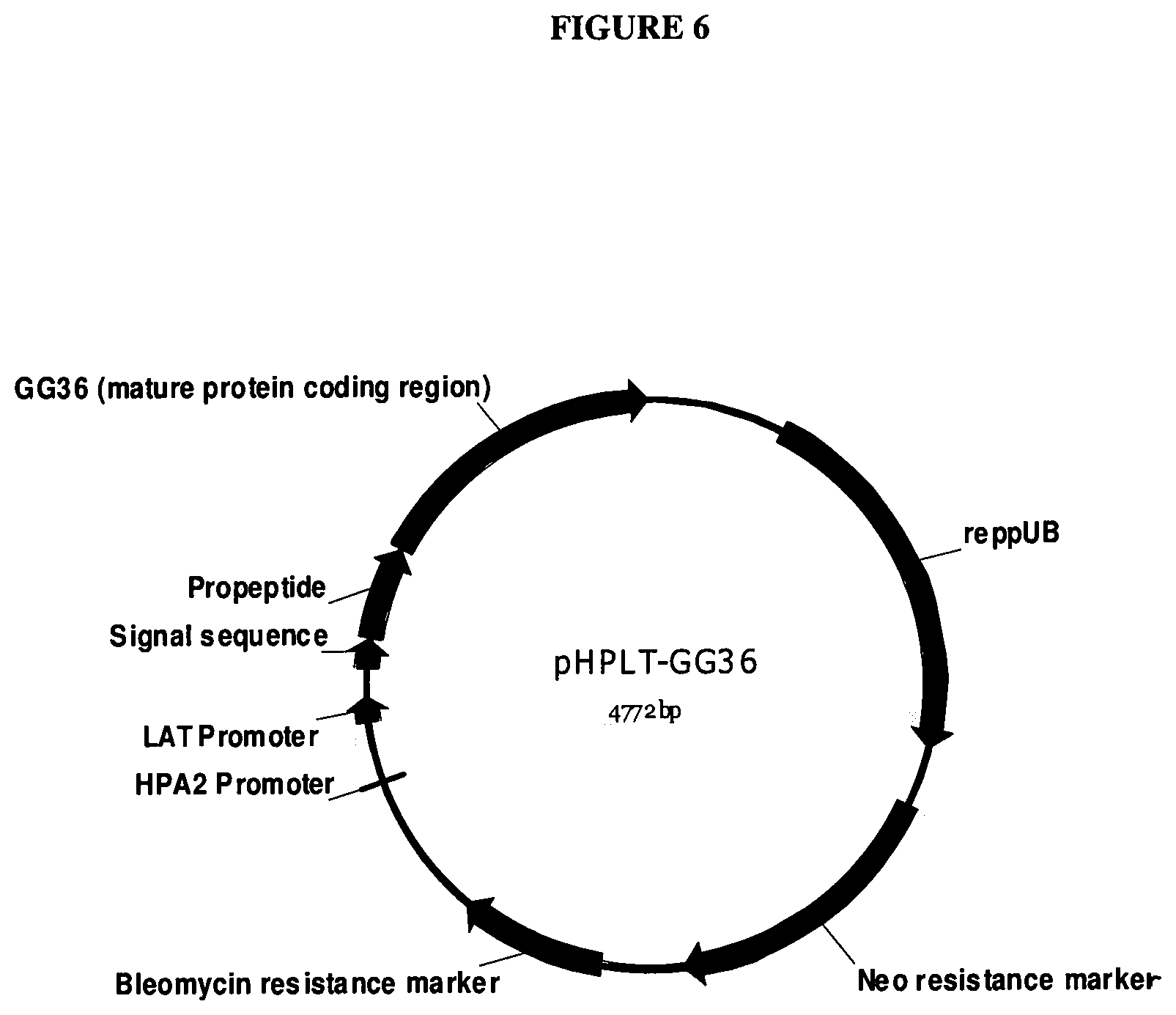
[0025] FIG. 6 provides map of pHPLT-GG36.
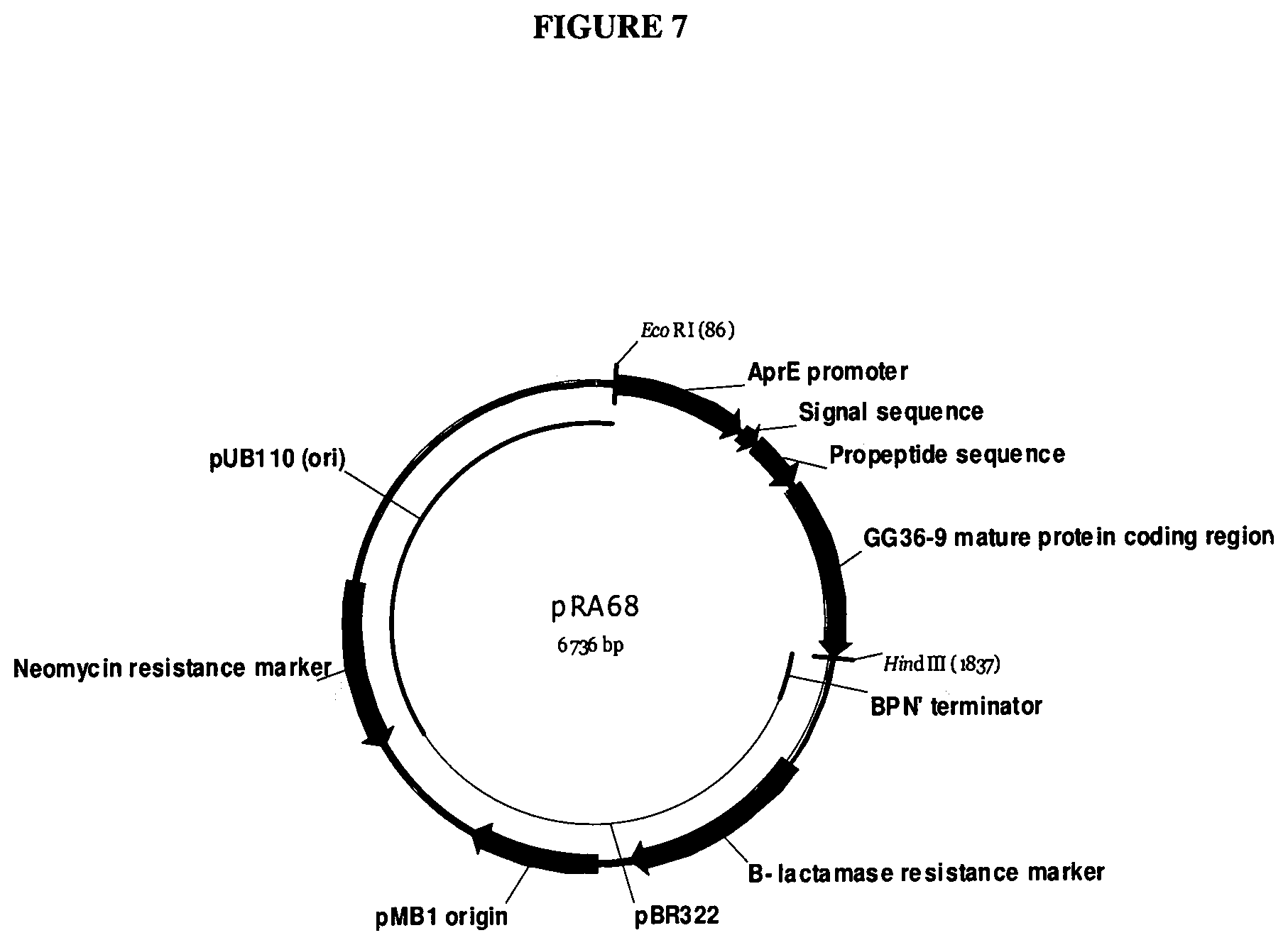
[0026] FIG. 7 provides a map of pRA68.
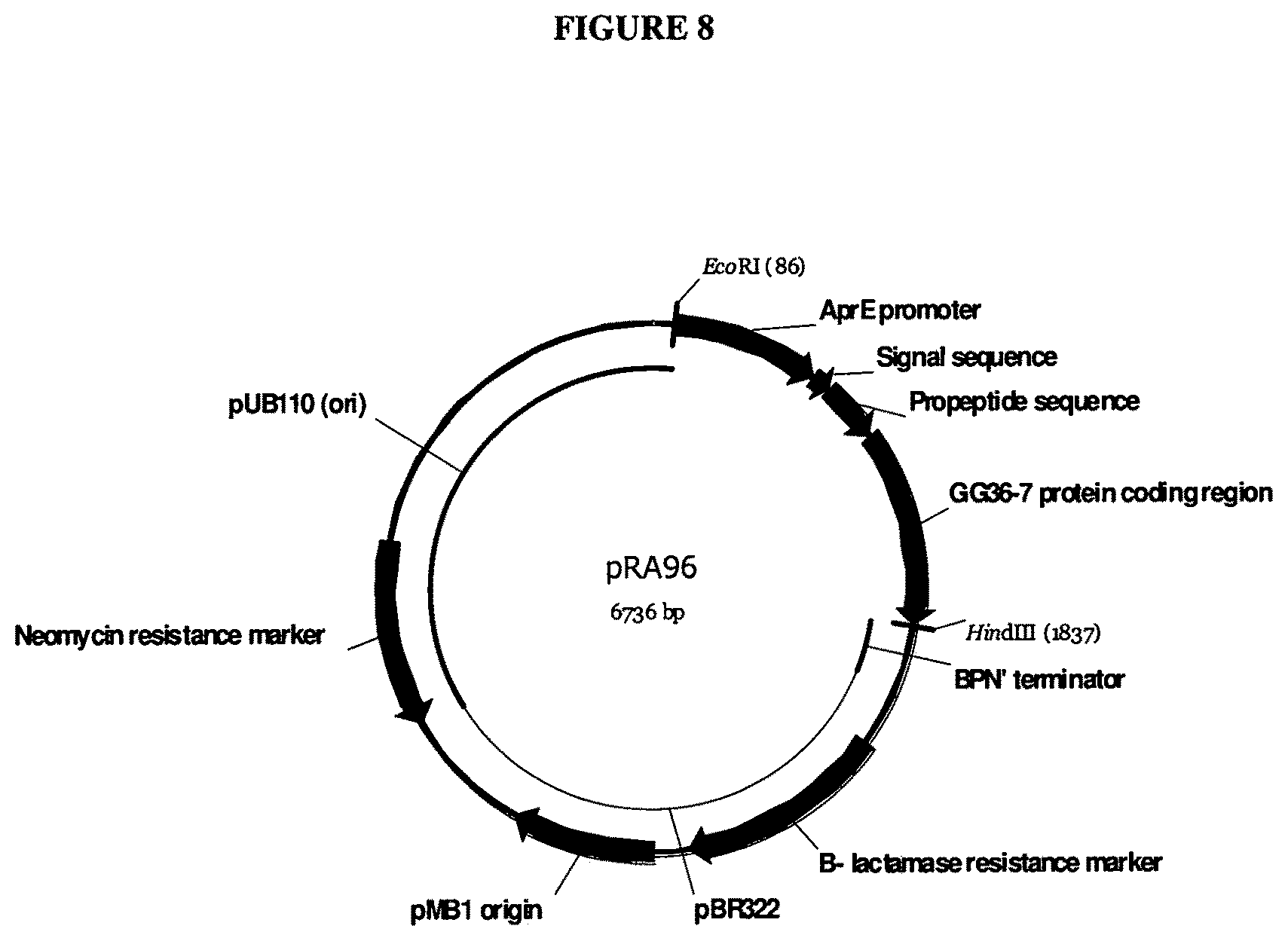
[0027] FIG. 8 provides a map of pRA96.
DESCRIPTION OF THE INVENTION
Definitions
[0028] Unless otherwise indicated, the practice of the present invention involves conventional techniques commonly used in molecular biology, protein engineering, microbiology, and recombinant DNA, which are within the skill of the art. Such techniques are known to those of skill in the art and are described in numerous texts and reference works well known to those of skill in the art. All patents, patent applications, articles and publications mentioned herein, both supra and infra, are hereby expressly incorporated herein by reference.
[0029] Unless defined otherwise herein, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention pertains. Many technical dictionaries are known to those of skill in the art. Although any methods and materials similar or equivalent to those described herein find use in the practice of the present invention, some suitable methods and materials are described herein. Accordingly, the terms defined immediately below are more fully described by reference to the specification as a whole. Numeric ranges are inclusive of the numbers defining the range. Unless otherwise indicated, nucleic acids are written left to right in 5' to 3' orientation; amino acid sequences are written left to right in amino to carboxy orientation, respectively. It is to be understood that this invention is not limited to the particular methodology, protocols, and reagents described, as these may vary, depending upon the context they are used by those of skill in the art.
[0030] The practice of the present invention employs, unless otherwise indicated, conventional techniques of protein purification, molecular biology, microbiology, recombinant DNA techniques and protein sequencing, all of which are within the skill of those in the art.
[0031] Furthermore, the headings provided herein are not limitations of the various aspects of the invention which can be had by reference to the specification as a whole. Accordingly, the terms defined immediately below are more fully defined by reference to the specification as a whole. Nonetheless, in order to facilitate understanding of the invention, a number of terms are defined below.
[0032] As used herein, the terms "protease" and "proteinase" refer to an enzyme protein that has the ability to break down other proteins. A protease has the ability to conduct "proteolysis," which begins protein catabolism by hydrolysis of peptide bonds that link amino acids together in a peptide or polypeptide chain forming the protein. This activity of a protease as a protein-digesting enzyme is referred to as "proteolytic activity." Many well known procedures exist for measuring proteolytic activity (see, e.g., Kalisz, "Microbial Proteinases," In: Fiechter (ed.), Advances in Biochemical Engineering/Biotechnology (1988)). For example, proteolytic activity may be ascertained by comparative assays which analyze the respective protease's ability to hydrolyze a commercial substrate. Exemplary substrates useful in the analysis of protease or proteolytic activity, include, but are not limited to, dimethyl casein (Sigma C-9801), bovine collagen (Sigma C-9879), bovine elastin (Sigma E-1625), and bovine keratin (ICN Biomedical 902111). Colorimetric assays utilizing these substrates are well known in the art (see, e.g., WO 99/34011 and U.S. Pat. No. 6,376,450, both of which are incorporated herein by reference). The pNA assay (see, e.g., Del Mar et al., Anal. Biochem. 99:316-320 [1979]) also finds use in determining the active enzyme concentration for fractions collected during gradient elution. This assay measures the rate at which p-nitroaniline is released as the enzyme hydrolyzes the soluble synthetic substrate, succinyl-alanine-alanine-proline-phenylalanine-p-nitroanilide (suc-AAPF-pNA). The rate of production of yellow color from the hydrolysis reaction is measured at 410 nm on a spectrophotometer and is proportional to the active enzyme concentration. In addition, absorbance measurements at 280 nanometers (nm) can be used to determine the total protein concentration. The active enzyme/total protein ratio gives the enzyme purity.
[0033] As used herein, the term "subtilisin" refers any member of the S8 serine protease family as described in MEROPS--The Peptidase Data base (see Rawlings et al., MEROPS: the peptidase database, Nucl. Acids Res., 34 Database issue, D270-272 [2006]). As described therein, the peptidase family S8 contains the serine endopeptidase subtilisin and its homologues (Rawlings and Barrett, Biochem. J. 290:205-218, [1993]). Family S8, also known as the subtilase family, is the second largest family of serine peptidases. The tertiary structures for several members of family S8 have now been determined. A typical S8 protein structure consists of three layers with a seven-stranded .beta. sheet sandwiched between two layers of helices. Subtilisin (S08.001) is the type structure for clan SB (SB). Despite the different structure, the active sites of subtilisin and chymotrypsin (S01.001) can be superimposed, which suggests the similarity is the result of convergent rather than divergent evolution.
[0034] A "protease variant" (or "variant protease") may refer to a protease that differs in its amino acid sequence from the amino acid sequence of a reference protease or parent protease by at least one amino acid residue. A parent protease or reference protease need not be a wild-type protease, but may itself be a variant of a wild-type protease. It is not intended that the reference or parent protease be limited to any particular amino acid sequence. A protease variant of a reference or parent protease may comprise an amino acid sequence comprising at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the amino acid sequence of the parent protease or reference protease and at least one amino acid substitution, insertion, or deletion relative to the amino acid sequence of the parent protease or reference protease. In one aspect, the invention includes a variant of a serine protease, wherein the variant has at least one mutation relative to the serine protease. In one aspect, the present invention includes a "BPN' variant" (or "BPN' subtilisin variant") comprising an amino acid sequence comprising one or more mutations relative to the mature BPN' sequence of SEQ ID NO:2.
[0035] A parent protease or reference protease can be, but is not limited to, e.g., a known protease (including, but not limited to, e.g., BPN') or a commercially available protease or a variant of the commercially available protease. A parent protease or reference protease may itself be a variant of a known or commercially available protease. A protease variant can be derived from a parent protease that is commercially available or a variant of such commercially available parent protease. Commercially available proteases, include, but are not limited to, e.g., proteases sold under the tradenames SAVINASE.RTM., POLARZYME.RTM., KANNASE.RTM., LIQUANASE.RTM., LIQUANASE ULTRA.RTM., SAVINASE ULTRA.RTM., OVOZYME.RTM., (by Novozymes A/S); MAXACAL.RTM., PROPERASE.RTM., PURAFECT.RTM., FN3.RTM., FN4.RTM. and PURAFECT OXP.RTM., PURAFAST.TM., PURAFECT.RTM. PRIME, PURAMAX.RTM. (by Danisco US Inc., formerly Genencor International, Inc.); and those available from Henkel/Kemira, namely BLAP (amino acid sequence shown in FIG. 29 of U.S. Pat. No. 5,352,604 with the following mutations S99D+S101R+S103A+V104I+G159S, hereinafter referred to as BLAP) and BLAP X (BLAP with S3T+V4I+V205I).
[0036] As used herein, a "cold water protease" is an enzyme that exhibits one or more of the following four criteria: (a) a performance index of at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from 1.1 to about 10, from 1.1 to about 8, or even from 1.1 to about 5 on BMI at pH 8 and 16.degree. C. (60.degree. F.) when compared to PURAFECT.RTM. Prime (SEQ ID NO:2 with the amino acid substitution Y217L), as defined in the "Test Method" set forth herein in Part I Example 1; (b) a performance index of at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from 1.3 to about 10, from 1.3 to about 8, or even from 1.3 to about 5 on BMI at pH 8 and 16.degree. C. (60.degree. F.) when compared to BPN' (SEQ ID NO:2), as defined in the "Test Method" set forth herein in Part I Example 1; (c) a performance index of at least 0.9, at least 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from 0.9 to about 10, from 0.9 to about 8, or even from 0.9 to about 5 on BMI at pH 8 and 16.degree. C. (60.degree. F.) when compared to BPN'-v3 (SEQ ID NO:4), as defined in the "Test Method" set forth herein in Part I Example 1; and/or (d) a performance index of at least 0.9, at least 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from 0.9 to about 10, from 0.9 to about 8, from 0.9 to about 5, from 1.0 to about 10, from 1.0 to about 8, or even from 1.0 to about 5 on BMI at pH 8 and 16.degree. C. (60.degree. F.) when compared to BPN'-v36 (SEQ ID NO:6), as defined in the "Test Method" set forth herein in Part I Example 1.
[0037] Some suitable cold water proteases are derived from subtilisins, particularly those derived from subtilisin BPN' (SEQ ID NO:2). A cold water protease can be a variant of BPN' having the amino acid sequence of SEQ ID NO:2 (e.g., "BPN' variant" or "BPN' subtilisin variant"). Some such cold water proteases comprise one or more of the amino acid substitutions set forth herein.
[0038] As used herein, the genus Bacillus includes all species within the genus Bacillus, as known to those of skill in the art, including but not limited to B. subtilis, B. licheniformis, B. lentus, B. brevis, B. stearothermophilus, B. alkalophilus, B. amyloliquefaciens, B. clausii, B. halodurans, B. megaterium, B. coagulans, B. circulars, B. lautus, and B. thuringiensis. It is recognized that the genus Bacillus continues to undergo taxonomical reorganization. Thus, it is intended that the genus include species that have been reclassified, including but not limited to such organisms as B. stearothermophilus, which is now named "Geobacillus stearothermophilus." The production of resistant endospores in the presence of oxygen is considered the defining feature of the genus Bacillus, although this characteristic also applies to the recently named Alicyclobacillus, Amphibacillus, Aneurinibacillus, Anoxybacillus, Brevibacillus, Filobacillus, Gracilibacillus, Halobacillus, Paenibacillus, Salibacillus, Thermobacillus, Ureibacillus, and Virgibacillus.
[0039] The terms "polynucleotide" and "nucleic acid," which are used interchangeably herein, refer to a polymer of any length of nucleotide monomers covalently bonded in a chain. DNA (deoxyribonucleic acid), a polynucleotide comprising deoxyribonucleotides, and RNA (ribonucleic acid), a polymer of ribonucleotides, are examples of polynucleotides or nucleic acids having distinct biological function. Polynucleotides or nucleic acids include, but are not limited to, a single-, double- or triple-stranded DNA, genomic DNA, cDNA, RNA, DNA-RNA hybrid, or a polymer comprising purine and pyrimidine bases, or other natural, chemically, biochemically modified, non-natural or derivatized nucleotide bases. The following are non-limiting examples of polynucleotides: genes, gene fragments, chromosomal fragments, expressed sequence tag(s) (EST(s)), exons, introns, messenger RNA (mRNA), transfer RNA (tRNA), ribosomal RNA (rRNA), ribozymes, complementary DNA (cDNA), recombinant polynucleotides, branched polynucleotides, plasmids, vectors, isolated DNA of any sequence, isolated RNA of any sequence, nucleic acid probes, and primers. Some polynucleotides comprise modified nucleotides, such as methylated nucleotides and nucleotide analogs, uracyl, other sugars and linking groups such as fluororibose and thioate, and nucleotide branches. A sequence of nucleotides may be interrupted by non-nucleotide components.
[0040] As used herein, the term "vector" refers to a nucleic acid construct or polynucleotide construct used to introduce or transfer nucleic acid(s) or polynucleotide(s) into a target cell or tissue. A vector is typically used to introduce foreign DNA into another cell or tissue. A vector generally comprises a DNA sequence that is a transgene and a larger polynucleotide sequence that serves as the "backbone" of the vector. The vector typically serves to transfers genetic information, such as the inserted transgene, to a target cell or tissue so as to isolate, multiply, or express the insert in the target cell or tissue. Vectors include plasmids, cloning vectors, bacteriophages, viruses (e.g., viral vector), cosmids, expression vectors, shuttle vectors, cassettes, and the like. A vector typically includes an origin of replication, a multicloning site, and a selectable marker. The process of inserting a vector into a target cell is typically referred to as transformation in bacterial and yeast cells and as transfection in mammalian cells. The present invention includes a vector that comprises a DNA sequence encoding a protease variant (e.g., precursor or mature protease variant) that is operably linked to a suitable prosequence (e.g., secretory, signal peptide sequence, etc.) capable of effecting the expression of the DNA sequence in a suitable host.
[0041] As used herein, the term "expression cassette" or "expression vector" refers to a nucleic acid construct or vector generated recombinantly or synthetically for the expression of a nucleic acid of interest (e.g., a foreign nucleic acid or transgene) in a target cell. The nucleic acid of interest typically expresses a protein of interest. An expression vector or expression cassette typically comprises a promoter nucleotide sequence that drives or promotes expression of the foreign nucleic acid. The expression vector or cassette also typically includes any other specified nucleic acid elements that permit transcription of a particular nucleic acid in a target cell. A recombinant expression cassette can be incorporated into a plasmid, chromosome, mitochondrial DNA, plastid DNA, virus, or nucleic acid fragment. Some expression vectors have the ability to incorporate and express heterologous DNA fragments in a host cell. Many prokaryotic and eukaryotic expression vectors are commercially available. Selection of appropriate expression vectors is within the knowledge of those of skill in the art. Selection of appropriate expression vectors for expression of a protein from a nucleic acid sequence incorporated into the expression vector is within the knowledge of those of skill in the art.
[0042] A DNA construct is an artificially constructed segment of nucleic acid that may be introduced into a target cell or tissue. A DNA construct typically comprises a DNA insert comprising a nucleotide sequence encoding a protein of interest that has been subcloned into a vector. The vector may contain bacterial resistance genes for growth in bacteria and a promoter for expression of the protein of interest in an organism. The DNA may be generated in vitro by PCR or any other suitable technique(s) known to those in the art. The DNA construct may comprise a nucleic acid sequence of interest. In one aspect, the sequence is operably linked to additional elements such as control elements (e.g., promoters, etc.). The DNA construct may further comprise a selectable marker and may further comprise an incoming sequence flanked by homology boxes. The construct may comprise other non-homologous sequences, added to the ends (e.g., stuffer sequences or flanks). The ends of the sequence may be closed such that the DNA construct forms a closed circle. The nucleic acid sequence of interest, which is incorporated into the DNA construct, using techniques well known in the art, may be a wild-type, mutant, or modified nucleic acid. The DNA construct may comprise one or more nucleic acid sequences homologous to the host cell chromosome. The DNA construct may comprise one or more non-homologous nucleotide sequences. Once the DNA construct is assembled in vitro, it may be used, e.g., to: 1) insert heterologous sequences into a desired target sequence of a host cell; and/or 2) mutagenize a region of the host cell chromosome (i.e., replace an endogenous sequence with a heterologous sequence); 3) delete target genes; and/or 4) introduce a replicating plasmid into the host. "DNA construct" is used interchangeably herein with "expression cassette."
[0043] As used herein, a "plasmid" refers to an extrachromosomal DNA molecule which is capable of replicating independently from the chromosomal DNA. A plasmid is double stranded (ds) and may be circular and is typically used as a cloning vector.
[0044] As used herein in the context of introducing a nucleic acid sequence into a cell, the term "introduced" refers to any method suitable for transferring the nucleic acid sequence into the cell. Such methods for introduction include but are not limited to protoplast fusion, transfection, transformation, electroporation, conjugation, and transduction (see, e.g., Ferrari et al., "Genetics," in Hardwood et al. (eds.), Bacillus, Plenum Publishing Corp., pp. 57-72 [1989]).
[0045] Transformation refers to the genetic alteration of a cell which results from the uptake, genomic incorporation, and expression of genetic material (e.g., DNA).
[0046] As used herein, a nucleic acid is "operably linked" with another nucleic acid sequence when it is placed into a functional relationship with another nucleic acid sequence. For example, a promoter or enhancer is operably linked to a nucleotide coding sequence if the promoter affects the transcription of the coding sequence. A ribosome binding site may be operably linked to a coding sequence if it is positioned so as to facilitate translation of the coding sequence. Typically, "operably linked" DNA sequences are contiguous. However, enhancers do not have to be contiguous. Linking is accomplished by ligation at convenient restriction sites. If such sites do not exist, synthetic oligonucleotide adaptors or linkers may be used in accordance with conventional practice.
[0047] As used herein, "recombinant" when used with reference to a cell typically indicates that the cell has been modified by the introduction of a heterologous nucleic acid sequence or that the cell is derived from a cell so modified. For example, a recombinant cell may comprise a gene not found in identical form within the native (non-recombinant) form of the cell, or a recombinant cell may comprise a native gene (found in the native form of the cell) but which has been modified and re-introduced into the cell. A recombinant cell may comprise a nucleic acid endogenous to the cell that has been modified without removing the nucleic acid from the cell; such modifications include those obtained by gene replacement, site-specific mutation, and related techniques known to those of ordinary skill in the art. Recombinant DNA (rDNA) is a form of artificial DNA that is created by combining two or more nucleotide sequences that would not normally occur together through the process of gene splicing. Recombinant DNA technology includes techniques for the production of recombinant DNA in vitro and transfer of the recombinant DNA into cells where it may be expressed or propagated, thereby producing a recombinant polypeptide.
[0048] As used herein, the term nucleic acid or gene "amplification" refers to a process by which specific DNA sequences are disproportionately replicated such that the amplified nucleic acid or gene becomes present in a higher copy number than was initially present in the genome. Selection of cells by growth in the presence of a drug (e.g., an inhibitor of an inhibitable enzyme) may result in the amplification of either the endogenous gene encoding the gene product required for growth in the presence of the drug or by amplification of exogenous (i.e., input) sequences encoding this nucleic acid or gene product or both.
[0049] As used herein, the term "primer" refers to an oligonucleotide (a polymer of nucleotide residues), whether occurring naturally as in a purified restriction digest or produced synthetically, which is capable of acting as a point of initiation of synthesis when placed under conditions in which synthesis of a primer extension product which is complementary to a nucleic acid strand is induced (i.e., in the presence of nucleotides and an inducing agent such as DNA polymerase and at a suitable temperature and pH). A primer is preferably single stranded for maximum efficiency in amplification, but may alternatively be double stranded. If double stranded, the primer is first treated to separate its strands before being used to prepare extension products. The primer may comprise an oligodeoxyribonucleotide. The primer must be sufficiently long to prime the synthesis of extension products in the presence of the inducing agent. The exact length of a primer depends on a variety of factors, including temperature, source of primer, and the use of the method.
[0050] As used herein, the term "probe" refers to an oligonucleotide, whether occurring naturally as in a purified restriction digest or produced synthetically, recombinantly or by PCR amplification, which is typically capable of hybridizing to another oligonucleotide of interest. A probe may be single-stranded or double-stranded. Probes are useful in the detection, identification and isolation of particular gene sequences. It is contemplated that any probe used in the present invention will be labeled with any "reporter molecule," so that it is detectable in any detection system, including, but not limited to enzyme (e.g., ELISA, as well as enzyme-based histochemical assays), fluorescent, radioactive, and luminescent systems. It is not intended that the invention be limited to any particular detection system or label.
[0051] As used herein, the term "polymerase chain reaction" (PCR) refers to the methods of U.S. Pat. Nos. 4,683,195 4,683,202, and 4,965,188, hereby incorporated by reference, which include methods for increasing the concentration of a segment of a target sequence in a mixture of genomic DNA without cloning or purification. This process for amplifying the target sequence is well known in the art.
[0052] As used herein, the term "amplification reagents" refers to those reagents (e.g., deoxyribonucleotide triphosphates, buffer, etc.) needed for amplification except for primers, nucleic acid template, and the amplification enzyme. Typically, amplification reagents along with other reaction components are placed and contained in a reaction vessel (test tube, microwell, etc.).
[0053] As used herein, the term "restriction endonuclease" or "restriction enzyme" refers to an enzyme (e.g., bacterial enzyme) that is capable of cutting double-stranded or single-stranded DNA at or near a specific sequence of nucleotides known as a restriction site. The nucleotide sequence comprising the restriction site is recognized and cleaved by a given restriction endonuclease or restriction enzyme and is frequently the site for insertion of DNA fragments. A restriction site can be engineered into an expression vector or DNA construct.
[0054] As is known in the art, a DNA sequence can be transcribed by an RNA polymerase to produce an RNA sequence, but an RNA sequence can be reverse transcribed by reverse transcriptase to produce a DNA sequence.
[0055] "Host strain" or "host cell" refers to a suitable host for an expression vector comprising a DNA sequence of interest. A DNA sequence of interest may express a protein of interest in the host strain or host cell.
[0056] A "protein" or "polypeptide" or "peptide" is a polymeric sequence of amino acid residues. A carboxyl group of one amino acid is linked to the amino group of another. The terms "protein" and "polypeptide" and "peptide" may be used interchangeably herein. A peptide comprises two or more amino acids. Peptides typically contain fewer amino acids than do polypeptides or proteins. The single and 3-letter code for amino acids as defined in conformity with the IUPAC-IUB Joint Commission on Biochemical Nomenclature (JCBN) is used through out this disclosure. The single letter X refers to any of the twenty amino acids. It is also understood that a polypeptide may be coded for by more than one nucleotide sequence due to the degeneracy of the genetic code.
[0057] In describing enzyme variants, the following nomenclature is used typically for ease of reference: Original amino acid(s):position(s):substituted amino acid(s). The accepted IUPAC single letter or triple letter amino acid abbreviation is employed. The single letter "X" refers to any amino acid residue However, when in the context of an amino acid substitution (e.g. "X003C"), it is to be understood that "X" refers to an amino acid residue other than the amino acid residue resulting from the substitution (e.g., X is an amino acid residue other than C). Mutations are typically named by the one letter code for the parent amino acid, followed by a three or two or one digit amino acid position number in an amino acid sequence and then the one letter code for the substituted amino acid. For example, mutating the amino acid glycine (G) at amino acid position 87 in an amino acid sequence by substituting to the amino acid serine (S) for glycine (G) is represented as "G087S" or "G87S". Typically, the substitution of a glycine at position 2 with a threonine is represented as G002T; however, such substitution may also be represented as G02T or G2T. One or two leading zeroes ("0") may be included simply to provide a convenient three number designation for each amino acid position. The amino acid position "001" is the same as "1" and thus "A001C" is the same as "A1C". "X001G" refers to the substitution of glycine (G) at amino acid position 1 in an amino acid sequence, wherein the amino acid that is to be replaced by glycine is any amino acid. Multiple mutations are indicated by inserting a "-" between the mutations or by using a plus (+) sign between the mutations. For example, amino acid substitutions at amino acid residue positions 87 and 90 in an amino acid sequence are represented as either "G087S-A090Y" or "G87S-A90Y" or "G87S+A90Y" or "G087S+A090Y". For deletions, the one letter code "Z" is used. For an insertion relative to the parent sequence, the one letter code "Z" is on the left side of the position number. For a deletion, the one letter code "Z" is on the right side of the position number. For insertions, the position number is the position number before the inserted amino acid(s), plus 0.01 for each amino acid. For example, an insertion of three amino acids alanine (A), serine (S) and tyrosine (Y) between position 87 and 88 is shown as "Z087.01A-Z087.02S-Z087.03Y." Thus, combining all the mutations above plus a deletion at position 100 is: "G087S-Z087.01A-Z087.02S-Z087.03Y-A090Y-A100Z."
[0058] A "prosequence" or "propetide sequence" refers to an amino acid sequence between the signal peptide sequence and mature protease sequence that is necessary for the secretion of the protease. Cleavage of the prosequence or propeptide sequence results in a mature active protease.
[0059] The term "signal sequence" or "signal peptide" refers to a sequence of amino acid residues that may participate in the secretion or direct transport of the mature or precursor form of a protein. The signal sequence is typically located N-terminal to the precursor or mature protein sequence. The signal sequence may also be referred to as a leader sequence. The signal sequence may be endogenous or exogenous. One exemplary exogenous signal sequence comprises the first seven amino acid residues of the signal sequence from Bacillus subtilis subtilisin fused to the remainder of the signal sequence of the subtilisin from Bacillus lentus (ATCC 21536). A signal sequence is normally absent from the mature protein. A signal sequence is typically cleaved from the protein by a signal peptidase after the protein is transported.
[0060] The term "hybrid signal sequence" refers to signal sequences in which part of sequence is obtained from the expression host fused to the signal sequence of the gene to be expressed. Synthetic sequences can be utilized.
[0061] The term "mature" form of a protein, polypeptide, or peptide refers to the functional form of the protein, polypeptide, or peptide without the signal peptide sequence and propeptide sequence.
[0062] The term "precursor" form of a protein or peptide refers to a mature form of the protein having a prosequence operably linked to the amino or carbonyl terminus of the protein. The precursor may also have a "signal" sequence operably linked to the amino terminus of the prosequence. The precursor may also have additional polynucleotides that are involved in post-translational activity (e.g., polynucleotides cleaved therefrom to leave the mature form of a protein, polypeptide, or peptide).
[0063] The term "wild-type" in reference to an amino acid sequence or nucleic acid sequence indicates that the amino acid sequence or nucleic acid sequence is native or naturally occurring sequence. As used herein, the term "naturally-occurring" refers to anything (e.g., proteins, amino acids, or nucleic acid sequences) that is found in nature (e.g., has not been manipulated by means of recombinant or chemical methods). As used herein, the term "non-naturally occurring" refers to anything that is not found in nature (e.g., recombinant or chemically synthesized nucleic acids produced in the laboratory).
[0064] An amino acid residue in a particular amino acid sequence may be numbered by correspondence with an amino acid residue in a position of a reference amino acid sequence. An amino acid residue of an amino acid sequence of interest which is in a position that "corresponds to" or is "corresponding to" or in "correspondence with" the position of an amino acid residue of a reference amino acid sequence indicates that the amino acid residue of the sequence of interest is located at a position that is equivalent or homologous to the position of an amino acid residue in the reference amino acid sequence. One skilled in the art can determine whether a particular residue position in a polypeptide corresponds to a position of a homologous reference sequence. For example, a protease variant may be aligned with that of a reference sequence (e.g., BPN' sequence of SEQ ID NO:2) using known techniques. The positions of the amino acid residues in the reference sequence are used for numbering of the amino acid residues in the sequence of interest. Accordingly, the amino acid residues of the protease variant may be numbered according to the corresponding amino acid residue position numbering of the reference sequence. For example, the amino acid residues in the reference sequence of SEQ ID NO: 2 may be used for determining amino acid residue position numbering of each amino acid residue of a protease variant of interest.
[0065] As used herein, "homology" refers to sequence similarity or identity, with identity being preferred. Homology may be determined using standard techniques known in the art (see, e.g., Smith and Waterman, Adv. Appl. Math. 2:482 [1981]; Needleman and Wunsch, J. Mol. Biol. 48:443 [1970; Pearson and Lipman, Proc. Natl. Acad. Sci. USA 85:2444 [1988]; software programs such as GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package (Genetics Computer Group, Madison, Wis.); and Devereux et al., Nucl. Acid Res. 12:387-395 [1984]). One example of a useful algorithm is PILEUP. PILEUP creates a multiple sequence alignment from a group of related sequences using progressive, pair-wise alignments. It can also plot a tree showing the clustering relationships used to create the alignment. PILEUP uses a simplification of the progressive alignment method of Feng and Doolittle (see Feng and Doolittle, J. Mol. Evol. 35:351-360 [1987]). The method is similar to that described by Higgins and Sharp (see Higgins and Sharp, CABIOS 5:151-153 [1989]). Useful PILEUP parameters including a default gap weight of 3.00, a default gap length weight of 0.10, and weighted end gaps. Another example of a useful algorithm is the BLAST algorithm, described by Altschul et al., (see Altschul et al., J. Mol. Biol. 215:403-410 [1990]; and Karlin and Altschul, Proc. Natl. Acad. Sci. USA 90:5873-5787 [1993]). A particularly useful BLAST program is the WU-BLAST-2 program (see Altschul et al., Meth. Enzymol. 266:460-480 [1996]). WU-BLAST-2 uses several search parameters, most of which are set to the default values. The adjustable parameters are set with the following values: overlap span=1, overlap fraction=0.125, word threshold (T)=11. The HSP S and HSP S2 parameters are dynamic values and are established by the program itself depending upon the composition of the particular sequence and composition of the particular database against which the sequence of interest is being searched. However, the values may be adjusted to increase sensitivity.
[0066] The percent sequence identity (% sequence identity or simply % identity) between a subject polypeptide sequence and a reference polypeptide sequence means that the subject amino acid sequence is identical on an amino acid residue-by-amino acid residue basis by a specified percentage to the reference polypeptide sequence over a comparison length when the sequences are optimally aligned, as determined, for example, by an amino acid sequence comparison algorithm or visual inspection. The percent sequence identity between a subject nucleic acid sequence and a reference nucleic acid sequence similarly means the subject nucleotide sequence is identical on a nucleic acid residue-by-nucleic acid residue basis by a specified percentage to the reference nucleotide sequence over a comparison length when the sequences are optimally aligned.
[0067] The percent sequence identity (percent identity or % sequence identity or % identity) between a reference sequence and a subject sequence of interest may be readily determined by one skilled in the art. The percent identity shared by two polypeptide sequences can be determined, for example, by direct comparison of the amino acid residues in each sequence by aligning the residues of the respective sequences for maximum similarity and determining the number of identical amino acid residues between the sequences by using a sequence comparison algorithm known in the art or by visual inspection. The two optimally aligned polypeptide sequences can be compared over the comparison length and the number of positions in the optimal alignment at which identical amino acid residues occur in both polypeptide sequences can be determined, thereby providing the number of matched positions, and the number of matched positions is then divided by the total number of positions over the comparison length. The resulting number is multiplied by 100 to yield the percent identity of the subject polypeptide sequence to the reference (or query) polypeptide sequence. The percent identity shared by two nucleic acid sequences can be similarly determined by direct comparison of the nucleotide residues in each sequence by aligning the residues of the respective sequences for maximum similarity and determining the number of identical nucleic acid residues between the nucleic acid sequences by using a sequence comparison algorithm or by visual inspection. The percent identity between two or more sequences may also be described as the sequences being a particular percent identical.
[0068] An example of an algorithm that is suitable for determining sequence identity is the BLAST algorithm, (see Altschul, et al., J. Mol. Biol., 215:403-410 [1990]). Software for performing BLAST analyses is publicly available through the National Center for Biotechnology Information. This algorithm involves first identifying high scoring sequence pairs (HSPs) by identifying short words of length W in the query sequence that either match or satisfy some positive-valued threshold score T when aligned with a word of the same length in a database sequence. These initial neighborhood word hits act as starting points to find longer HSPs containing them. The word hits are expanded in both directions along each of the two sequences being compared for as far as the cumulative alignment score can be increased. Extension of the word hits is stopped when: the cumulative alignment score falls off by the quantity X from a maximum achieved value; the cumulative score goes to zero or below; or the end of either sequence is reached. The BLAST algorithm parameters W, T, and X determine the sensitivity and speed of the alignment. The BLAST program uses as defaults a wordlength (W) of 11, the BLOSUM62 scoring matrix (see Henikoff and Henikoff, Proc. Natl. Acad. Sci. USA 89:10915 [1992]) alignments (B) of 50, expectation (E) of 10, M'5, N'-4, and a comparison of both strands.
[0069] The BLAST algorithm then performs a statistical analysis of the similarity between two sequences (see, e.g., Karlin and Altschul, supra). One measure of similarity provided by the BLAST algorithm is the smallest sum probability (P(N)), which provides an indication of the probability by which a match between two nucleotide or amino acid sequences would occur by chance. For example, a nucleic acid is considered similar to a protease-encoding nucleic acid of this invention if the smallest sum probability in a comparison of the test nucleic acid to a protease-encoding nucleic acid is less than about 0.1, more preferably less than about 0.01, and most preferably less than about 0.001. Where the test nucleic acid encodes a protease polypeptide, it is considered similar to a specified protease-encoding nucleic acid if the comparison results in a smallest sum probability of less than about 0.5, and more preferably less than about 0.2.
[0070] "Optimal alignment" or "optimally aligned" refers to the alignment of two (or more) sequences giving the highest percent identity score. For example, optimal alignment of two polypeptide sequences can be achieved by manually aligning the sequences such that the maximum number of identical amino acid residues in each sequence are aligned together or by using software programs or procedures described herein or known in the art. Optimal alignment of two nucleic acid sequences can be achieved by manually aligning the sequences such that the maximum number of identical nucleotide residues in each sequence are aligned together or by using software programs or procedures described herein or known in the art.
[0071] Two sequences (e.g., polypeptide sequences) may be deemed "optimally aligned" when they are aligned using defined parameters, such as a defined amino acid substitution matrix, gap existence penalty (also termed gap open penalty), and gap extension penalty, so as to achieve the highest identity score possible for that pair of sequences. The BLOSUM62 scoring matrix (see Henikoff and Henikoff, supra) is often used as a default scoring substitution matrix in polypeptide sequence alignment algorithms (e.g., BLASTP). The gap existence penalty is imposed for the introduction of a single amino acid gap in one of the aligned sequences, and the gap extension penalty is imposed for each residue position in the gap. Exemplary alignment parameters employed are: BLOSUM62 scoring matrix, gap existence penalty=11, and gap extension penalty=1. The alignment score is defined by the amino acid positions of each sequence at which the alignment begins and ends (e.g., the alignment window), and optionally by the insertion of a gap or multiple gaps into one or both sequences, so as to achieve the highest possible similarity score.
[0072] Optimal alignment between two or more sequences can be determined manually by visual inspection or by using a computer, such as, but not limited to e.g., the BLASTP program for amino acid sequences and the BLASTN program for nucleic acid sequences (see, e.g., Altschul et al., Nucleic Acids Res. 25(17):3389-3402 (1997); see also the National Center for Biotechnology Information (NCBI) website) or CLUSTALW program.
[0073] A polypeptide of interest may be said to be "substantially identical" to a reference polypeptide if the polypeptide of interest comprises an amino acid sequence having at least about 60%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 99.5% sequence identity to the amino acid sequence of the reference polypeptide. The percent identity between two such polypeptides can be determined manually by inspection of the two optimally aligned polypeptide sequences or by using software programs or algorithms (e.g., BLAST, ALIGN, CLUSTAL) using standard parameters. One indication that two polypeptides are substantially identical is that the first polypeptide is immunologically cross-reactive with the second polypeptide. Typically, polypeptides that differ by conservative amino acid substitutions are immunologically cross-reactive. Thus, a polypeptide is substantially identical to a second polypeptide, e.g., where the two peptides differ only by a conservative amino acid substitution or one or more conservative amino acid substitutions.
[0074] A nucleic acid of interest may be said to be "substantially identical" to a reference nucleic acid if the nucleic acid of interest comprises a nucleotide sequence having at least about 60%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 99.5% sequence identity to the nucleotide sequence of the reference nucleic acid. The percent identity between two such nucleic acids can be determined manually by inspection of the two optimally aligned nucleic acid sequences or by using software programs or algorithms (e.g., BLAST, ALIGN, CLUSTAL) using standard parameters. One indication that two nucleic acid sequences are substantially identical is that the two nucleic acid molecules hybridize to each other under stringent conditions (e.g., within a range of medium to high stringency).
[0075] As used herein, "isolated" in reference to a particular component of interest means that component is essentially or substantially free of other components. For example, an "isolated" polypeptide means the polypeptide is essentially or substantially free of other components, including, but not limited to, e.g., other polypeptides and cellular components. An "isolated" nucleic acid means the nucleic acid is essentially or substantially free of other components, including, but not limited to, e.g., other nucleic acids and cellular components. For purposes of this application, "isolated" refers to nucleic acids or polypeptides that are not part of a library (e.g., screening library).
[0076] Purity and homogeneity are typically determined using analytical chemistry techniques, such as polyacrylamide gel electrophoresis or high-performance liquid chromatography. A protein or nucleic acid that is the predominant species present in a preparation is substantially purified. The term "purified" denotes that a nucleic acid or protein gives rise to essentially one band in an electrophoretic gel, chromatographic eluate, and/or a media subjected to density gradient centrifugation. Particularly, "purified" means that when isolated, the isolate contains at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 99.5% or more of nucleic acid or protein by weight of the isolate. Purified polypeptides may be obtained by a number of methods including, but not limited to, e.g., laboratory synthesis, chromatography (e.g., high-performance liquid chromatography) preparative electrophoresis, polyacrylamide gel electrophoresis followed by visualization upon staining, centrifugation, precipitation, affinity purification, etc. (see, generally, R Scopes, Protein Purification, Springer-Verlag, N.Y. (1982), Deutscher, Methods in Enzymol., Vol. 182: Guide to Protein Purification, Academic Press, Inc. N.Y. (1990)). The invention includes an isolated or purified polypeptides (e.g., isolated protease variants or subtilisin variants of the invention) and isolated or purified nucleic acids (e.g., nucleic acids encoding protease variants or subtilisin variants of the invention).
[0077] In a related sense, the invention provides methods of enriching compositions for one or more molecules of the invention, such as one or more polypeptides of the invention (e.g., one or more protease variants of the invention) or one or more nucleic acids of the invention (e.g., one or more nucleic acids encoding one or more protease variants of the invention). A composition is enriched for a molecule when there is a substantial increase in the concentration of the molecule after application of a purification or enrichment technique. A substantially pure polypeptide or nucleic acid will typically comprise at least about 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98, 99%, 99.5% or more by weight (on a molar basis) of all macromolecular species in a particular composition.
[0078] As used herein, the term "combinatorial mutagenesis" refers to methods in which libraries of nucleic acid variants of a reference nucleic acid sequence are generated. In these libraries, the variants contain one or several mutations chosen from a predefined set of mutations. The methods also provide means to introduce random mutations which were not members of the predefined set of mutations. Some such methods include those set forth in U.S. Pat. No. 6,582,914, hereby incorporated by reference. Some such combinatorial mutagenesis methods include and/or encompass methods embodied in commercially available kits (e.g., QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis Kit (Stratagene)).
[0079] As used herein, having "improved properties" used in connection with a protease variant refers to a protease variant having improved properties compared to a reference or parent protease. Protease variants of the invention may exhibit one of more of the following properties: enhanced or improved proteolytic activity, enhanced or improved stability, enhanced or improved ability to clean a surface or item, enhanced or improved cleaning performance, enhanced or improved fabric or laundry cleaning performance or wash performance, enhanced or improved hand wash performance, enhanced or improved hand or manual dishwashing performance, enhanced or improved automatic dishwashing performance, enhanced or improved laundry performance compared to a reference protease or parent protease of interest.
[0080] As used herein, the term "functional assay" refers to an assay that provides an indication of a protein's activity. The term typically refers to assay systems in which a protein is analyzed for its ability to function in its usual capacity. For example, in the case of enzymes, a functional assay involves determining the effectiveness of the enzyme in catalyzing a reaction.
[0081] The term "property" or grammatical equivalents thereof in the context of a molecule may refer to any characteristic or attribute of the molecule that can be selected or detected. For example, in the context of a polypeptide, a property may be enzymatic activity (e.g., proteolytic activity), stability, or other property.
[0082] A "mutant" nucleic acid sequence typically refers to a nucleic acid sequence that has an alteration in at least one codon occurring in a host cell's wild-type sequence such that the expression product of the mutant nucleic acid sequence is a protein with an altered amino acid sequence relative to the wild-type protein. The expression product may have an altered functional capacity (e g, enhanced enzymatic activity).
[0083] As used herein, the term "net charge" is defined as the sum of all charges present in a molecule. "Net charge changes" can be made to a parent protein molecule to provide a protein variant that has a net charge that differs from that of the parent protein molecule (i.e., the variant has a net charge that is not the same as that of the parent molecule). For example, substitution of a neutral amino acid of a protein with a negatively charged amino acid or substitution of a positively charged amino acid of a protein with a neutral amino acid results in net charge of -1 with respect to the unmodified protein. Substitution of a positively charged amino acid of a protein with a negatively charged amino acid results in a net charge of -2 with respect to the unmodified protein. Substitution of a neutral amino acid of a protein with a positively charged amino acid or substitution of a negatively charged amino acid of a protein with a neutral amino acid results in net charge of +1 with respect to the parent. Substitution of a negatively charged amino acid of a protein with a positively charged amino acid results in a net charge of +2 with respect to the unmodified protein. The net charge of a parent protein can also be altered by deletion and/or insertion of one or more charged amino acids.
[0084] The terms "thermally stable" and "thermostable" and "thermostability" in reference to a polypeptide indicates that the polypeptide is resistant to a permanent change in its activity caused solely by heat, such as, e.g., via exposure to higher temperature. For example, a thermally stable enzyme means that the enzyme is resistant to a permanent change in its enzymatic activity caused solely by heat, such as, e.g., via exposure to higher temperature. Typically, a protease that is thermally stable is able to retain at least about 50%, 60%, 70%, 75%, 80%, 85%, 90%, 92%, 95%, 96%, 97%, 98%, or 99% of its proteolytic activity after exposure to increased temperatures over a given time period, e.g., at least about 60 minutes, 120 minutes, 180 minutes, 240 minutes, 300 minutes, etc.
[0085] Cleaning activity of a polypeptide or protease may refer to a cleaning performance achieved by a protease. Cleaning activity may be determined by using various assays for cleaning one or more of various enzyme-sensitive stains on an object, item, or surface (e.g., a stain resulting from food, grass, blood, ink, blood/milk/ink, milk/oil/pigment, egg yolk, milk, oil, and/or egg protein). Cleaning performance of a polypeptide (e.g., a polypeptide of the invention (such as a protease variant) or a reference polypeptide (e.g., reference protease) may be determined by subjecting the stain on the object, item, or surface to standard wash condition(s) and assessing the degree to which the stain is removed by using various chromatographic, spectrophotometric, or other quantitative methodologies. Exemplary cleaning assays and methods are known in the art and include, but are not limited to those described in WO 99/34011 and U.S. Pat. No. 6,605,458, both of which are herein incorporated by reference, as well as those cleaning assays and methods included in the Part I Examples and Part II Examples provided below.
[0086] The term "cleaning effective amount" of a protease variant or reference protease refers to the amount of protease that achieves a desired level of enzymatic activity in a specific cleaning composition. Such effective amounts are readily ascertained by one of ordinary skill in the art and are based on many factors, such as the particular protease used, the cleaning application, the specific composition of the cleaning composition, and whether a liquid or dry (e.g., granular, tablet, bar) composition is required, etc.
[0087] The term "cleaning adjunct material" refers to any liquid, solid, or gaseous material included in cleaning composition other than a protease variant of the invention. The cleaning compositions of the present invention may include one of more cleaning adjunct materials. Each cleaning adjunct material is typically selected depending on the particular type and form of cleaning composition (e.g., liquid, granule, powder, bar, paste, spray, tablet, gel, foam, or other composition). Preferably, each cleaning adjunct material is compatible with the protease enzyme used in the composition.
[0088] The term "enhanced performance" in the context of cleaning activity refers to an increased or greater cleaning activity by an enzyme on certain enzyme sensitive stains such as egg, milk, grass, ink, oil, and/or blood, as determined by usual evaluation after a standard wash cycle and/or multiple wash cycles.
[0089] The term "diminished performance" in the context of cleaning activity refers to a decreased or lesser cleaning activity by an enzyme on certain enzyme sensitive stains such as egg, milk, grass or blood, as determined by usual evaluation after a standard wash cycle.
[0090] As used herein, the term "institutional cleaning composition" refers to products suitable for use in institutions including but not limited to schools, hospitals, factories, stores, corporations, buildings, restaurants, office complexes and buildings, processing and/or manufacturing plants, veterinary hospitals, factory farms, factory ranches, etc.
[0091] As used herein, "fabric and home care product" refers to a product or device generally intended to be used or consumed in the form in which it is sold and that is for treating fabric, hard surface and any other surface in the area of fabric and home care, including: air care including air freshener and scent delivery systems, car care, dishwashing, fabric conditioning (including softening and/or freshening), laundry detergency, laundry and rinse additive and/or care, hard surface cleaning and/or treatment including floor and toilet bowl cleaners, and other cleaning products for consumer and institutional use.
[0092] As used herein, the terms "cleaning composition" and "cleaning formulation" refer to compositions that find use in the removal of undesired compound(s) from an item(s) to be cleaned, such as, but not limited to, e.g., fabric, laundry, dishes, dishware, contact lenses, other solid substrates, hair (including human or animal hair) (shampoos), skin (soaps, cosmetics, and creams), teeth (mouthwashes, toothpastes), non-fabric and home care objects, filters, membranes (e.g., filtration membrane, including, but not limited to, ultrafiltration membranes), hard surfaces and other surfaces, including, but not limited to, e.g., the hard surface of a table (table top or legs), wall, another furniture item or object, floor, ceiling, etc. A cleaning composition or cleaning formulation may be useful in a personal care application and/or in personal care item, including, e.g., but not limited to, shampoo (for cleaning human or animal hair); soap, cream or cosmetics (for skin cleaning and/or skin care); mouthwash (for oral care); toothpaste (for cleaning teeth and/or oral care). The terms encompass any material/compound selected for the particular type of cleaning composition or formulation desired and the form of the product (e.g., liquid, gel, granule, or spray composition), as long as the composition or formulation is compatible with the protease and/or other enzyme(s) used in the composition or formulation. The specific selection of cleaning composition or formulation materials are readily made by considering the surface, object, or item (e.g., fabric) to be cleaned, and the desired form of the composition or formulation for the cleaning conditions during use. In one aspect, a cleaning composition or formulation may be a fabric and home care product (e.g., a cleaning composition for cleaning laundry). In another aspect, a cleaning composition or formulation is not a fabric and home care product (e.g., a cleaning composition for cleaning contact lens, hair, teeth, or skin, or useful in personal care applications and/or personal care items).
[0093] Cleaning compositions and cleaning formulations include any composition that is suited for cleaning, bleaching, disinfecting, and/or sterilizing any object, item, and/or surface. Such compositions and formulations include, but are not limited to, for example, liquid and/or solid compositions, including cleaning or detergent compositions (e.g., liquid, tablet, gel, bar, granule, and/or solid laundry cleaning or detergent compositions and fine fabric detergent compositions; hard surface cleaning compositions and formulations, such as for glass, wood, ceramic and metal counter tops and windows; carpet cleaners; oven cleaners; fabric fresheners; fabric softeners; and textile, laundry booster cleaning or detergent compositions, laundry additive cleaning compositions, and laundry pre-spotter cleaning compositions; dishwashing compositions, including hand or manual dishwash compositions (e.g., "hand" or "manual" dishwashing detergents) and automatic dishwashing compositions (e.g., "automatic dishwashing detergents").
[0094] Cleaning compositions or cleaning formulations, as used herein, include, unless otherwise indicated, granular or powder-form all-purpose or heavy-duty washing agents, especially cleaning detergents; liquid, granular, gel, solid, tablet, or paste-form all-purpose washing agents, especially the so-called heavy-duty liquid (HDL) detergent or heavy-duty powder detergent (HDD) types; liquid fine-fabric detergents; hand or manual dishwashing agents, including those of the high-foaming type; hand or manual dishwashing, automatic dishwashing, or dishware or tableware washing agents, including the various tablet, powder, solid, granular, liquid, gel, and rinse-aid types for household and institutional use; liquid cleaning and disinfecting agents, including antibacterial hand-wash types, cleaning bars, mouthwashes, denture cleaners, car shampoos, carpet shampoos, bathroom cleaners; personal care items, such as, but not limited to, e.g., hair shampoos and/or hair-rinses for humans (and other animals), shower gels and foam baths, skin care items, cosmetics, creams, bath and personal human soaps; metal cleaners; as well as cleaning auxiliaries, such as bleach additives and "stain-stick" or pre-treat types. Some granular compositions are in "compact" form; some liquid compositions are in a "concentrated" form.
[0095] In one aspect, the invention provides a cleaning composition or detergent composition comprising at least one protease variant or polypeptide of the invention, wherein the cleaning composition or detergent composition is useful for cleaning contact lens(es). In another aspect, the invention provides a cleaning composition or detergent composition comprising at least one protease variant or polypeptide of the invention, wherein the cleaning composition or detergent composition is useful in a personal care application. In another aspect, the invention provides a cleaning composition or detergent composition comprising at least one protease variant or polypeptide of the invention, wherein the cleaning composition or detergent composition is useful for cleaning or rinsing hair, including human hair and/or animal hair (e.g., hair shampoo and/or hair-rinse). In another aspect, the invention provides a cleaning composition or detergent composition comprising at least one protease variant or polypeptide of the invention, wherein the cleaning composition or detergent composition is useful for cleaning or treating skin (e.g., human and/or animal skin) (e.g., shower gel, foam bath, skin care cleaner, cosmetic, cream, and/or bath soap). In another aspect, the invention provides a cleaning composition or detergent composition comprising at least one protease variant or polypeptide of the invention, wherein the cleaning composition or detergent composition is useful for cleaning teeth and/or dentures and/or for oral care. Such cleaning compositions or detergent compositions may comprise at least one adjunct ingredient or carrier, at least one additional enzyme, at least one builder, and/or at least one surfactant and may be formulated or in a form appropriate to their use. Such cleaning or detergent compositions may contain phosphate or may be phosphate-free. Additional details regarding compositions of the invention are provided elsewhere herein.
[0096] As used herein, "fabric cleaning compositions" include hand and machine laundry detergent compositions including laundry additive compositions and compositions suitable for use in the soaking and/or pretreatment of stained fabrics (e.g., clothes, linens, and other textile materials).
[0097] As used herein, "non-fabric cleaning compositions" include non-textile (i.e., non-fabric) surface cleaning compositions, including, but not limited to, for example, hand or manual or automatic dishwashing detergent compositions, oral cleaning compositions, denture cleaning compositions, and personal cleansing compositions.
[0098] As used herein, the term "detergent composition" or "detergent formulation" is used in reference to a composition intended for use in a wash medium for the cleaning of soiled or dirty objects, including particular fabric and/or non-fabric objects or items. Such compositions of the present invention are not limited to any particular detergent composition or formulation. Indeed, the detergents of the invention may comprise at least one protease variant of the invention and, in addition, one or more surfactants, transferase(s), hydrolytic enzymes, perhydrolases, oxido reductases, builders (e.g., a builder salt), bleaching agents, bleach activators, bluing agents, fluorescent dyes, caking inhibitors, masking agents, enzyme activators, antioxidants, and/or solubilizers. In some instances, a builder salt is a mixture of a silicate salt and a phosphate salt, preferably with more silicate (e.g., sodium metasilicate) than phosphate (e.g., sodium tripolyphosphate). Some compositions of the invention, such as, but not limited to, cleaning compositions or detergent compositions, do not contain any phosphate (e.g., phosphate salt or phosphate builder).
[0099] As used herein, "dishwashing composition" refers to all forms of compositions for cleaning dishware, including cutlery, including, but not limited to, granular and liquid forms. In some aspects, the dishwashing composition is an "automatic dishwashing" composition that finds use in automatic dishwashing machines. It is not intended that the present invention be limited to any particular type of dishware composition. Indeed, the present invention finds use in cleaning dishware (e.g., dishes, including, but not limited to plates, cups, glasses, bowls, etc.) and cutlery (e.g., utensils, including, but not limited to, spoons, knives, forks, serving utensils, etc.) of any material, including, but not limited to, ceramics, plastics, metals, china, glass, acrylics, etc. The term "dishware" is used herein in reference to both dishes and cutlery.
[0100] As used herein, the term "bleaching" refers to the treatment of a material (e.g., fabric, laundry, pulp, etc.) or surface for a sufficient length of time and/or under appropriate pH and/or temperature conditions to effect a brightening (i.e., whitening) and/or cleaning of the material. Examples of chemicals suitable for bleaching include, but are not limited to, for example, ClO.sub.2, H.sub.2O.sub.2, peracids, NO.sub.2, etc.
[0101] As used herein, "wash performance" of a protease (e.g., a protease variant of the invention) refers to the contribution of a protease variant to washing that provides additional cleaning performance to the detergent as compared to the detergent without the addition of the protease variant to the composition. Wash performance is compared under relevant washing conditions. In some test systems, other relevant factors, such as detergent composition, sud concentration, water hardness, washing mechanics, time, pH, and/or temperature, can be controlled in such a way that condition(s) typical for household application in a certain market segment (e.g., hand or manual dishwashing, automatic dishwashing, dishware cleaning, tableware cleaning, fabric cleaning, etc.) are imitated.
[0102] The term "relevant washing conditions" is used herein to indicate the conditions, particularly washing temperature, time, washing mechanics, sud concentration, type of detergent and water hardness, actually used in households in a hand dishwashing, automatic dishwashing, or laundry detergent market segment.
[0103] The term "improved wash performance" is used to indicate that a better end result is obtained in stain removal under relevant washing conditions, or that less protease variant, on weight basis, is needed to obtain the same end result relative to the corresponding wild-type or starting parent protease.
[0104] As used herein, the term "disinfecting" refers to the removal of contaminants from the surfaces, as well as the inhibition or killing of microbes on the surfaces of items. It is not intended that the present invention be limited to any particular surface, item, or contaminant(s) or microbes to be removed.
[0105] The "compact" form of the cleaning compositions herein is best reflected by density and, in terms of composition, by the amount of inorganic filler salt. Inorganic filler salts are conventional ingredients of detergent compositions in powder form. In conventional detergent compositions, the filler salts are present in substantial amounts, typically about 17 to about 35% by weight of the total composition. In contrast, in compact compositions, the filler salt is present in amounts not exceeding about 15% of the total composition. The filler salt can be present in amounts that do not exceed about 10%, or more preferably, about 5%, by weight of the composition. The inorganic filler salts can be selected from the alkali and alkaline-earth-metal salts of sulfates and chlorides. The filler salt may be sodium sulfate.
[0106] The position of an amino acid residue in a given amino acid sequence is typically numbered herein using the numbering of the position of the corresponding amino acid residue of the B. amyloliquefaciens subtilisin BPN' amino acid sequence shown in SEQ ID NO:2. The B. amyloliquefaciens subtilisin BPN' amino acid sequence of SEQ ID NO:2 thus serves as a reference sequence. A given amino acid sequence, such as a protease variant amino acid sequence described herein, can be aligned with the BPN' sequence (SEQ ID NO:2) using an alignment algorithm as described herein, and an amino acid residue in the given amino acid sequence that aligns (preferably optimally aligns) with an amino acid residue in the BPN' sequence can be conveniently numbered by reference to the corresponding amino acid residue in the subtilisin BPN' sequence.
[0107] Generally, the nomenclature used herein and many of the laboratory procedures in cell culture, molecular genetics, molecular biology, nucleic acid chemistry, and protein chemistry described below are well known and commonly employed by those of ordinary skill in the art. Methods for production and manipulation of recombinant nucleic acid methods, nucleic acid synthesis, cell culture methods, and transgene incorporation (e.g., transfection, electroporation) are known to those skilled in the art and are described in numerous standard texts. Oligonucleotide synthesis and purification steps are typically performed according to specifications. Techniques and procedures are generally performed according to conventional methods well known in the art and various general references that are provided throughout this document. Procedures therein are believed to be well known to those of ordinary skill in the art and are provided for the convenience of the reader.
[0108] A fabric and home care product may comprise a protease (including a protease variant), including one or more protease variants of the invention and a material selected from the group consisting of an encapsulate comprising a perfume, a hueing agent, an amphiphilic cleaning polymer and mixtures thereof, with the balance of any aspects of the aforementioned composition is made up of one or more adjunct materials, is disclosed. In one aspect of the aforementioned fabric and home care product, said fabric and home care product may comprise, based on total fabric and home care product weight, from about 0.005 weight percent (0.0005 wt %) to about 0.1 wt %, from about 0.001 wt % to about 0.05 wt %, or even from about 0.002 wt % to about 0.03 wt % of said protease. In one aspect of the aforementioned fabric and home care product, said fabric and home care product may comprise, based on total fabric and home care product weight, about 0.00003 wt % to about 0.1 wt %, from about 0.00008 wt % to about 0.05 wt %, or even from about 0.0001 wt % to about 0.04 wt % fabric hueing agent. In one aspect of the aforementioned fabric and home care product, said fabric and home care product may comprise, based on total fabric and home care product weight, from about 0.001 wt % to about 5 wt %, from about 0.01 wt % to about 2 wt %, or even from about 0.03 wt % to about 0.5 wt %, perfume capsules. In one aspect of the aforementioned fabric and home care product, said fabric and home care product may comprise, based on total fabric and home care product weight, from about 0.1 wt % to about 5 wt %, from about 0.25 wt % to about 2.5 wt %, or even from about 0.3 wt % to about 1.5 wt % amphiphilic cleaning polymer.
Polypeptides of the Invention
[0109] The present invention provides novel polypeptides, which may be collectively referred to as "polypeptides of the invention". Polypeptides of the invention include isolated, recombinant, substantially pure, or non-naturally occurring protease variants, including, for example, subtilisin protease variant polypeptides, which have enzymatic activity (e.g., proteolytic activity) and/or additional properties discussed in greater detail elsewhere herein (e.g., cleaning activity, stability, etc.). Polypeptides of the invention include isolated, recombinant, substantially pure, or non-naturally occurring cold water proteases having proteolytic activities, cleaning activities, stability, and other properties discussed elsewhere herein. Such polypeptides, including cold water proteases, of the invention may have enhanced performance relative to known proteases (e g, enhanced proteolytic activity, enhanced cleaning performance or activity, enhanced stability, etc). Polypeptides of the invention are useful in cleaning applications and may be incorporated into cleaning compositions that are useful in methods of cleaning an item or a surface (e.g., of surface of an item) in need of cleaning.
[0110] Polypeptides of the invention having proteolytic activity are useful in fabric and home care products, including, e.g., fabric and home care cleaning compositions. Some polypeptides of the invention having proteolytic activity are useful in personal care compositions. Polypeptides of the invention having proteolytic activity are also useful in non-fabric and home care products (i.e., those products that are not fabric and home care products are described herein). A protease variant of the invention may be a subtilisin protease variant. The invention includes Bacillus species protease variants and Bacillus species subtilisin protease variants.
[0111] Polypeptides of the invention are disclosed throughout this specification, including, but not limited to, Part I Examples and Part II Examples provided below. The invention includes an isolated, recombinant, substantially pure, or non-naturally occurring protease variant (e.g., a subtilisin variant) having proteolytic activity, which polypeptide comprises an amino acid sequence having at least about 60%, 70%, 80%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% sequence identity to a specific protease variant amino acid sequence set forth in any of the Part I and Part II Examples below. In one aspect, the invention includes an isolated, recombinant, substantially pure, or non-naturally occurring subtilisin polypeptide having proteolytic activity, wherein said polypeptide is a protease variant of the BPN' sequence of SEQ ID NO:2 and said polypeptide comprises an amino acid sequence having at least about 60%, 70%, 80%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to SEQ ID NO:2 and a set of amino acid substitutions set forth in any of Part I Examples. In one aspect, the invention includes an isolated, recombinant, substantially pure, or non-naturally occurring subtilisin polypeptide having proteolytic activity, wherein said polypeptide is a protease variant of the GG36 sequence of SEQ ID NO:755 and said polypeptide comprises an amino acid sequence having at least about 60%, 70%, 80%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to SEQ ID NO:755 and a set of amino acid substitutions set forth in any of Part II Examples.
[0112] In a first aspect, the invention provides an isolated or non-naturally occurring polypeptide protease variant of a parent protease enzyme, the variant having proteolytic activity and comprising an amino acid sequence which comprises an alteration at one or more amino acid positions corresponding to amino acid positions of SEQ ID NO:2 selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217, wherein the at least one alteration is independently (i) an insertion of one or more amino acid residues upstream or downstream of the amino acid residue which occupies the position, (ii) a deletion of the amino acid residue which occupies the position, or (iii) a substitution of the amino acid residue which occupies the position with a different amino acid residue, wherein each amino acid position is numbered by correspondence with an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin protease BPN' set forth in SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2. A variant of a protease enzyme according to the first aspect of the invention may be termed a protease variant. A variant according to the first aspect of the invention may be a non-naturally occurring protease. A variant according to the first aspect of the invention may be isolated or purified. A variant according to the first aspect of the invention may be a subtilisin variant. A variant according to the first aspect of the invention may have proteolytic activity. A variant according to the first aspect of the invention may have enhanced proteolytic activity compared to the proteolytic activity of the BPN' amino acid sequence set forth in SEQ ID NO:2.
[0113] A variant according to the first aspect of the invention may be a variant of a parent protease that is a subtilisin protease, and the subtilisin protease may be a Bacillus species protease. A parent Bacillus protease according to the first aspect of the invention may be selected from the group consisting of B. amyloliquefaciens subtilisin protease BPN' (SEQ ID NO:2), B. stearothermophilus, B. subtilis, B. licheniformis, B. lentus, B. brevis, B. stearothermophilus, B. alkalophilus, B. amyloliquefaciens, B. clausii, B. halodurans, B. megaterium, B. coagulans, B. circulars, B. lautus, B. species TS-23, B. thuringiensis, BPN'-v3 (SEQ ID N04:), and BPN'-v36 (SEQ ID NO:6). A parent protease according to the first aspect of the invention may comprise an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to the amino acid sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6.
[0114] A variant according to the first aspect of the invention may be a protease variant having a mature form. A variant according to the first aspect of the invention may be a protease variant having a mature form.
[0115] A variant according to the first aspect of the invention may comprise an amino acid sequence comprising an alteration at two amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising an alteration at three amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising an alteration at four amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising an alteration at five amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising an alteration at six amino acid positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising an alteration at each of the amino acid positions corresponding to positions of SEQ ID NO:2 selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at a position selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217.
[0116] A variant according to the first aspect of the invention may comprise an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of two or three positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of four positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of five positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprises an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of six or seven positions selected from the group consisting of positions 24, 53, 78, 97, 101, 128, and 217. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising a substitution of an amino acid residue with a different amino acid residue at each of positions 24, 53, 78, 101, 128, and 217, wherein each position is numbered by correspondence with a position in SEQ ID NO:2.
[0117] A variant according to the first aspect of the invention may comprise an amino acid sequence comprising at least one amino acid substitution selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising at least two amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising at least three amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising at least four amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising at least five amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising at least six amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X097A, X101N, X128A/S, and X217Q/L. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising a set of substitutions selected from the group consisting of: (a) X128A/S and/or X217L/Q, (b) G128A/S and/or Y217L/Q, and (c) G097A, G128A/S, and Y217L/Q. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising amino acid substitutions X024G/R+X053G+X078N+X101N+X128A/S+X217Q/L, wherein the variant has proteolytic activity; and wherein optionally the variant has enhanced proteolytic activity or enhanced cleaning activity compared to the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6 or a performance index in a proteolytic assay (e.g., AAPF assay) or cleaning assay (e.g., BMI, grass, or egg microswatch assay) that is greater than that of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the first aspect of the invention may comprise at least one amino acid substitution selected from the group of S024G, S053G, S078N, S101N, G128A/S, and Y217Q. A variant according to the first aspect of the invention may comprise an amino acid sequence comprising amino acid substitutions S024G+S053G+S078N+S101N+G128A+Y217Q.
[0118] A variant according to the first aspect of the invention may comprise an amino acid sequence having at least 60%, 65%, 70%, 80%, or 85% sequence identity to the amino acid sequence of SEQ ID NO:2. A variant according to the first aspect of the invention may comprise an amino acid sequence having at least 90% sequence identity to SEQ ID NO:2. A variant according to the first aspect of the invention may comprise an amino acid sequence having at least 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:2. A variant according to the first aspect of the invention may comprise an amino acid sequence having at least 95% sequence identity to SEQ ID NO:2.
[0119] A variant according to the first aspect of the invention may have enhanced proteolytic activity compared to the proteolytic activity of the protease set forth in SEQ ID NO:2. A variant according to the first aspect of the invention may have enhanced proteolytic activity compared to the proteolytic activity of the protease set forth in SEQ ID NO:4. A variant according to the first aspect of the invention may have enhanced proteolytic activity compared to the proteolytic activity of the protease set forth in SEQ ID NO:6. A variant according to the first aspect of the invention may have enhanced cleaning activity compared to the cleaning activity of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the first aspect of the invention may a performance index in a proteolytic assay (e.g., AAPF assay) or cleaning assay (e.g., BMI, grass, or egg microswatch assay) that is greater than that of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6.
[0120] In a second aspect, the invention provides an isolated or non-naturally occurring polypeptide protease variant of a parent protease, the variant comprising an amino acid sequence comprising three amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2. A variant according to the first aspect of the invention may have enhanced proteolytic activity or enhanced cleaning activity compared to the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the first aspect of the invention may have a performance index in a proteolytic assay (e.g., AAPF assay) or cleaning assay (e.g., BMI, grass, or egg microswatch assay) that is greater than that of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6.
[0121] A variant according to the second aspect of the invention may comprise an amino acid sequence comprising three amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2. A variant according to the second aspect of the invention may comprise an amino acid sequence comprising four amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q. A variant according to the second aspect of the invention may comprise an amino acid sequence comprising five amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q. A variant according to the second aspect of the invention may comprise an amino acid sequence comprising six amino acid substitutions selected from the group consisting of X024G/R, X053G, X078N, X101N, X128A/S, and X217L/Q. A variant according to the second aspect of the invention may comprise an amino acid sequence further comprising amino acid substitution X097A. A variant according to the second aspect of the invention may comprise an amino acid sequence comprising amino acid substitutions X024G/R+X053G+X078N+X101N+X128A/S+X217L/Q or X097A+X128A/S+X217L/Q.
[0122] A variant according to the second aspect of the invention may comprise an amino acid sequence comprising amino acid substitutions S024G/R+S053G+S078N+S101N+G128A/S+Y217L/Q or G097A+G128A/S+Y217L/Q. A variant according to the second aspect of the invention may comprise an amino acid sequence comprising amino acid substitutions S024G+S053G+S078N+S101N+G128S+Y217Q or S024G+S053G+S078N+S101N+G128A+Y217Q.
[0123] The amino acid sequence of a variant according to the second aspect of the invention may further comprise the substitution N109G. The amino acid sequence of a variant according to the second aspect of the invention may further comprise the substitution N076D. The amino acid sequence of a variant according to the second aspect of the invention may further comprise the substitution S033T. The amino acid sequence of a variant according to the second aspect of the invention may further comprise the substitution N243V. The amino acid sequence of a variant according to the second aspect of the invention may further comprise the substitution S248A. The amino acid sequence of a variant according to the second aspect of the invention may further comprise the substitution A088T. The amino acid sequence of a variant according to the second aspect of the invention may further comprise the substitution S063G. The amino acid sequence of the variant according to the second aspect of the invention may further comprise two or more substitutions selected from the group consisting of N109G, N076D, S033T, N243V, S248A, A088T, and S063G.
[0124] The amino acid sequence of a variant according to the second aspect of the invention may may comprise an amino acid sequence comprising amino acid substitutions S024G+S053G+S078N+S101N+G128S+Y217Q or S024G+S053G+S078N+S101N+G128A+Y217Q and may further comprise a set of amino acid substitutions selected from the group consisting of: A088T+N109G+A116T+G131H+N243V+L257G, S033T+N076D, S009T+N109G+K141R+N243V, S162G+K256R, N109G+A116T, N109G+L257G, S162G+L257G, N061G+N109G+N243V, N109G+N243V+S248A, S033T+N076D+N109G+N218S+N243V+S248N+K256R, N109G+A116T+N243V+K256R, A088T+N109G+A116T+G131H+N243V, A088T+N109G, N109G+N243V, T158S+L257G, N061S+N109G+N243V, P040A+N109G+N243V+S248N+K256R, S009T+S018T+Y021N+N109G+K141R, A088T+N109G+A116T+T158S+N243V+K256R, A088T+N109G+A116T+T158S+N218S+L257G, N109G+K256R, N109G+N243V+K256R, S063G+K256R, S063G+N109G, S063G, S063G+N076D, S033T+N076D+N218S, and N076D+N218S.
[0125] A variant according to the second aspect of the invention may comprise an amino acid sequence comprising amino acid substitutions S024G+S053G+S078N+S101N+G128A+Y217Q and may further comprise a set of amino acid substitutions selected from the group consisting of: A088T+N109G+A116T+G131H+N243V+L257G, S033T+N076D, S009T+N109G+A128S+K141R+N243V, S162G+K256R, N109G+A116T, N109G+L257G, S162G+L257G, N061G+N109G+N243V, N109G+A128S+N243V+S248A, S033T+N076D+N109G+A128S+N218S+N243V+S248N+K256R, N109G+A116T+N243V+K256R, A088T+N109G+A116T+G131H+N243V, A088T+N109G, N109G+N243V, T158S+L257G, N061S+N109G+N243V, P040A+N109G+A128S+N243V+S248N+K256R, S009T+S018T+Y021N+N109G+A128S+K141R, A088T+N109G+A116T+T158S+N243V+K256R, A088T+N109G+A116T+T158S+N218S+L257G, N109G+K256R, N109G+A128S+N243V+K256R, S063G+K256R, S063G+N109G, S063G+A128S, S063G+N076D, S033T+N076D+A128S+N218S, and N076D+N218S, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2. Notably, for example, a variant according to the second aspect of the invention which comprises a sequence comprising substitutions S024G+S053G+S078N+S101N+G128A+Y217Q, and which further comprises the set of substitutions S009T+N109G+A128S+K141R+N243V, has a serine (S) at position 128 because the alanine in position 128 (i.e., G128A substitution) is substituted with S (A128S substitution).
[0126] The amino acid sequence of a variant according to the second aspect of the invention may have at least 60%, 65%, or 70% sequence identity to the amino acid sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. The amino acid sequence of a variant according to the second aspect of the invention may have at least 80% or 85% sequence identity to the amino acid sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. The amino acid sequence of a variant according to the second aspect of the invention may have at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to the amino acid sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6.
[0127] The patent protease according to the second aspect of the invention may be a subtilisin protease. The parent protease according to the second aspect of the invention may comprise an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to the amino acid sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the second aspect of the invention may be a protease variant having a mature form. A variant according to the first aspect of the invention may be a protease variant having a mature form.
[0128] The invention also provides an isolated polypeptide (e.g., protease variant) according to the second aspect of the invention comprising an amino acid sequence having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to the amino acid sequence of SEQ ID NO:6 and comprising a set of amino acid substitutions selected from the group consisting of: [0129] A088T+N109G+A116T+G131H+N243V+L257G, [0130] S033T+N076D, [0131] S009T+N109G+A128S+K141R+N243V, [0132] S162G+K256R, [0133] N109G+A116T, [0134] N109G+L257G, [0135] S162G+L257G, [0136] N061G+N109G+N243V, [0137] N109G+A128S+N243V+S248A, [0138] S033T+N076D+N109G+A128S+N218S+N243V+S248N+K256R, [0139] N109G+A116T+N243V+K256R, [0140] A088T+N109G+A116T+G131H+N243V, [0141] A088T+N109G, [0142] N109G+N243V, [0143] T158S+L257G, [0144] N061S+N109G+N243V, [0145] P040A+N109G+A128S+N243V+S248N+K256R, [0146] S009T+S018T+Y021N+N109G+A128S+K141R, [0147] A088T+N109G+A116T+T158S+N243V+K256R, [0148] A088T+N109G+A116T+T158S+N218S+L257G, [0149] N109G+K256R, [0150] N109G+A128S+N243V+K256R, [0151] S063G+K256R, [0152] S063G+N109G, [0153] S063G+A128S, [0154] S063G+N076D, [0155] S033T+N076D+A128S+N218S, and [0156] N076D+N218S, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2. Such polypeptide may have enhanced proteolytic activity or enhanced cleaning activity compared to the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. Such polypeptide may have a performance index in a proteolytic assay (e.g., AAPF assay) or cleaning assay (e.g., BMI, grass, or egg microswatch assay) that is greater than that of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the second aspect of the invention may have enhanced proteolytic activity compared to the proteolytic activity of the BPN' protease having the sequence of SEQ ID NO:2. A variant according to the second aspect of the invention may have enhanced proteolytic activity compared to the proteolytic activity of the protease having the sequence of SEQ ID NO:4 or SEQ ID NO:6. The parent protease of a variant according to the second aspect of the invention may be a subtilisin protease, and optionally the subtilisin protease may be a Bacillus species. The parent subtilisin protease of a variant according to the second aspect of the invention may be selected from the group consisting of B. amyloliquefaciens subtilisin protease BPN' (SEQ ID NO:2), B. stearothermophilus, B. subtilis, B. licheniformis, B. lentus, B. brevis, B. stearothermophilus, B. alkalophilus, B. amyloliquefaciens, B. clausii, B. halodurans, B. megaterium, B. coagulans, B. circulars, B. lautus, B. species TS-23, B. thuringiensis, BPN'-v3 (SEQ ID N04:), and BPN'-v36 (SEQ ID NO:6). A parent protease according to the second aspect of the invention may comprise an amino acid sequence having at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to the amino acid sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6.
[0157] A variant according to the second aspect of the invention may be a protease variant having a mature form. A parent protease according to the second aspect of the invention may be protease having a mature form.
[0158] A variant according to the second aspect of the invention may comprise an amino acid sequence having at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to SEQ ID NO:2, SEQ ID NO:6, or SEQ ID NO:4.
[0159] In a third aspect, the invention provides an isolated or non-naturally occurring polypeptide having protease activity, said polypeptide comprising an amino acid sequence having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a polypeptide sequence selected from the group consisting of:
a) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+G131H+N243V+- L257G;
b) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S033T+N076D;
c) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S009T+N109G+K141R+N243V;
d) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S162G+K256R;
e) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+A116T;
f) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+L257G;
g) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S162G+L257G;
h) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N061G+N109G+N243V;
i) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+N109G+N243V+S248A;
j) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S033T+N076D+N109G+N218S+N243V+- S248N+K256R;
k) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+A116T+N243V+K256R;
l) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+G131H+N243V;
m) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G;
n) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+N243V;
o) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+T158S+L257G;
p) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N061S+N109G+N243V;
q) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+P040A+N109G+N243V+S248N+K256R;
r) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S009T+S018T+Y021N+N109G+K141R;
s) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+T158S+N243V+- K256R;
t) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+A088T+N109G+A116T+T158S+N218S+- L257G;
u) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N109G+K256R;
v) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+N109G+N243V+K256R;
w) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+K256R;
x) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+N109G;
y) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S063G;
z) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+S063G+N076D;
aa) BPN'-S024G+S053G+S078N+S101N+G128S+Y217Q+S033T+N076D+N218S;
bb) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q+N076D+N218S; and
[0160] cc) BPN'-S024G+S053G+S078N+S101N+G128A+Y217Q, wherein each amino acid position of the variant is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2. A variant according to the third aspect of the invention may have enhanced proteolytic activity or enhanced cleaning activity compared to the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the third aspect of the invention may have a performance index in a proteolytic assay (e.g., AAPF assay) or cleaning assay (e.g., BMI, grass, or egg microswatch assay) that is greater than that of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6.
[0161] A polypeptide according to the third aspect of the invention may comprise or consist of the amino acid sequence set forth in any of a) through cc) above or a fragment of any thereof having proteolytic activity. A variant or parent protease according to the third aspect of the invention may be in a mature form.
[0162] In a fourth aspect, the invention provides an isolated or non-naturally occurring polypeptide having protease activity selected from the group consisting of: (a) a polypeptide comprising an amino acid sequence having at least 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to the polypeptide sequence of SEQ ID NO:6; (b) a polypeptide encoded by a polynucleotide that hybridizes under at least high stringency conditions with (i) the polynucleotide sequence of SEQ ID NO:5 or (ii) a complementary polynucleotide sequence of (i); and (c) a polypeptide encoded by a polynucleotide comprising a nucleotide sequence having at least 90%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to the polynucleotide sequence of SEQ ID NO:5. A polypeptide according to the fourth aspect of the invention may comprise or consist of the amino acid sequence of SEQ ID NO:6, or a fragment thereof having proteolytic activity. The variant according to the fourth aspect of the invention may have increased proteolytic activity and/or cleaning performance or activity compared to that of mature BPN' (SEQ ID NO:2). The variant according to the fourth aspect of the invention may have increased proteolytic activity and/or cleaning performance or activity compared to that of SEQ ID NO:4 or SEQ ID NO:6. A variant or parent protease according to the fourth aspect of the invention may be in a mature form.
[0163] In a fifth aspect, the invention provides an isolated or non-naturally occurring protease variant of a parent protease, wherein: (a) the variant comprises an amino acid sequence having no more than 25, 20, 15, or 10 alterations relative to the parent protease, wherein (i) the alterations are independently selected from an insertion, a deletion, or a substitution, and (ii) the alterations include a substitution of glycine at positions 24 and 53, a substitution of asparagine at positions 78 and 101, a substitution of alanine or serine at position 128, and a substitution of glutamine at position 217, (b) the parent protease has at least 90% sequence identity to SEQ ID NO:2, (c) the amino acid sequence of SEQ ID NO:2 is used for determining position numbering; and (d) the variant has increased proteolytic activity relative to the parent protease, wherein each amino acid position is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2. The amino acid sequence of a variant according to the fifth aspect of the invention may have at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to the amino acid sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6.
[0164] The parent protease according to the fifth aspect of the invention may have at least 80%, 85%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the sequence of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. The parent protease according to the fifth aspect of the invention may be a protease having the amino acid sequence of SEQ ID NO:2. A variant or parent protease according to the fifth aspect of the invention may be in a mature form.
[0165] The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise the substitution N109G. The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise the substitution N076D. The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise the substitution S033T. The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise the substitution N243V. The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise the substitution S248A. The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise the substitution A088T. The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise the substitution S063G.
[0166] The amino acid sequence of a variant according to the fifth aspect of the invention may further comprise a set of amino acid substitutions selected from the group consisting of: A088T+N109G+A116T+G131H+N243V+L257G, S033T+N076D, S009T+N109G+K141R+N243V, S162G+K256R, N109G+A116T, N109G+L257G, S162G+L257G, N061G+N109G+N243V, N109G+N243V+S248A, S033T+N076D+N109G+N218S+N243V+S248N+K256R, N109G+A116T+N243V+K256R, A088T+N109G+A116T+G131H+N243V, A088T+N109G, N109G+N243V, T158S+L257G, N061S+N109G+N243V, P040A+N109G+N243V+S248N+K256R, S009T+S018T+Y021N+N109G+K141R, A088T+N109G+A116T+T158S+N243V+K256R, A088T+N109G+A116T+T158S+N218S+L257G, N109G+K256R, N109G+N243V+K256R, S063G+K256R, S063G+N109G, S063G, S063G+N076D, S033T+N076D+N218S, and N076D+N218S.
[0167] The parent protease according to the fifth aspect of the invention may be a subtilisin protease. The parent protease according to the fifth aspect of the invention may be BPN' set forth in SEQ ID NO:2. The variant according to the fifth aspect of the invention may have increased proteolytic activity and/or cleaning performance or activity compared to that of SEQ ID NO:4 or SEQ ID NO:6.
[0168] In a sixth aspect, the invention provides an isolated or non-naturally occurring protease variant of a parent protease, wherein (a) the variant comprises an amino acid sequence (i) having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, or 97% identity to the sequence of SEQ ID NO:2 and (ii) comprising a substitution of glycine at positions 24 and 53, a substitution of asparagine at positions 78 and 101, a substitution of alanine or serine at position 128, and a substitution of glutamine at position 217; (b) the parent protease has at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to SEQ ID NO:2; (c) each amino acid position of the variant is numbered by correspondence with an amino acid position of the sequence of SEQ ID NO:2; and (d) the variant has increased proteolytic activity relative to the parent protease. The parent protase according to the sixth aspect of the invention may be a subtilisin protease. The parent protease according to the sixth aspect of the invention may be a protease having the amino acid sequence of SEQ ID NO:2. The variant according to the sixth aspect of the invention may have increased proteolytic activity and/or cleaning activity compared to the proteolytic activity and/or cleaning activity, respectively, of the parent protease.
[0169] The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise the substitution N109G. The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise the substitution N076D. The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise the substitution S033T. The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise the substitution N243V. The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise the substitution S248A. The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise the substitution A088T. The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise the substitution S063G.
[0170] The amino acid sequence of a variant according to the sixth aspect of the invention may further comprise a set of amino acid substitutions selected from the group consisting of: A088T+N109G+A116T+G131H+N243V+L257G, S033T+N076D, S009T+N109G+K141R+N243V, S162G+K256R, N109G+A116T, N109G+L257G, S162G+L257G, N061G+N109G+N243V, N109G+N243V+S248A, S033T+N076D+N109G+N218S+N243V+S248N+K256R, N109G+A116T+N243V+K256R, A088T+N109G+A116T+G131H+N243V, A088T+N109G, N109G+N243V, T158S+L257G, N061S+N109G+N243V, P040A+N109G+N243V+S248N+K256R, S009T+S018T+Y021N+N109G+K141R, A088T+N109G+A116T+T158S+N243V+K256R, A088T+N109G+A116T+T158S+N218S+L257G, N109G+K256R, N109G+N243V+K256R, S063G+K256R, S063G+N109G, S063G, S063G+N076D, S033T+N076D+N218S, and N076D+N218S.
[0171] The variant according to the sixth aspect of the invention may have increased proteolytic activity and/or cleaning performance or activity compared to that of SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the sixth aspect of the invention may have a performance index in a proteolytic assay (e.g., AAPF assay) or cleaning assay (e.g., BMI, grass, or egg microswatch assay) that is greater than that of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant or parent protease according to the sixth aspect of the invention may be in a mature form.
[0172] In a seventh aspect, the invention provides an isolated or non-naturally occurring protease variant of BPN' subtilisin protease having the amino acid sequence shown in SEQ ID NO:2, said protease variant selected from the group consisting of:
[0173] (a) a protease variant comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% identity to the amino acid sequence of SEQ ID NO:2, said variant having increased proteolytic activity and/or increased cleaning activity compared to the protease of SEQ ID NO:2 and/or SEQ ID NO:4, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2 and wherein said variant comprises at least one set of amino acid substitution(s) relative to SEQ ID NO:2 selected from groups (i) through (x):
[0174] (i) G097A-G128A-P210S-Y217Q-N218A and G097A-G128A-P210S-Y217Q;
[0175] (ii) S063T-S078N-G097A-S101A-G128A-S183T-Y217Q-T244N, N061A-S078N-G097A-G128A-Y217Q-S224A, S053G-S078N-G097A-G128A-P129T-Q185T-Y217Q, S063T-S078N-G097A-S101A-G128A-S183T-Y217Q, S063T-S078N-G097A-S101A-G128A-Y217Q, S063T-S078N-G097A-S101A-G128A-Y217Q-T244I, and S078N-G097A-G128A-P129T-Y217Q;
[0176] (iii) S063T-S078N-G097A-S101A-G128A-S183T-Y217Q, S063T-S078N-G097A-S101A-G128A-S183T-Y217Q-T244N, S063T-S078N-G097A-S101A-G128A-Y217Q, and S063T-S078N-G097A-S101A-G128A-Y217Q-T244I;
[0177] (iv) G097A-I111V-M124V-Y217Q, G097A-I111V-Y167A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128A-V203Y-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128S-V203Y-Y217Q, and V068A-A092G-Y217Q;
[0178] (v) N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128S-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-I111V-G128S-Y217Q;
[0179] (vi) N061P-G097A-G128S-Y217Q, N061P-G097A-S101N-G128A-P210S-Y217Q, N061P-N062Q-G097A-S101N-I111V-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128S-Y217Q, and S024G-N025G-S053G-N061P-S078N-G097A-S101N-I111V-G128S-Y217Q;
[0180] (vii) N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, N025G-G097A-S101N-G128A-Y217Q, N025G-S038G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-G128A-Y217Q, N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, N025G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-S101N-G128A-Y217Q, N061P-S101N-G128A-Y217Q, S024G-N025G-N061P-G097A-G128A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G128A-Y217Q, S024G-N025G-T055P-G097A-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-N061P-G097A-G128A-Y217Q, S024G-S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-T055P-N061P-G097A-G128A-Y217Q, S053G-G097A-S101N-G128A-Y217Q, S053G-N061P-G097A-S101N-G128A-Y217Q-S249N, S053G-N061P-S078N-G097A-G128A-Y217Q, S053G-S078N-G097A-S101N-G128A-Y217Q, S053G-T055P-G097A-S101N-G128A-Y217Q, S053G-T055P-N061P-S101N-G128A-Y217Q, S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, T055P-G097A-S101N-G128A-Y217Q, and T055P-N061P-S078N-G097A-S101N-G128A-Y217Q;
[0181] (viii) S024G-N025G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-T055P-N061P-S078N-S101N-G128A-Y217Q, S053G-G097A-S101N-G128A-Y217Q, and T055P-N061P-S078N-G128A-Y217Q;
[0182] (ix) N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N061P-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-G097A-S101N-G128A-Y217Q, S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-N025G-T055P-G097A-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-G128A-Y217Q; and
[0183] (x) S024G-G097A-S101N-G128A-Y217Q, S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-S101N-G128A-Y217Q, S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-T055P-N061P-G097A-S101N-G128A, S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A, N061P-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-T055P-N061P-S078N-S101N-G128A-Y217Q, N025G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G128A-Y217Q, S024G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, T055P-N061P-S078N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S038G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-G128A-Y217Q, N025G-S053G-N061P-S078N-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S078N-S101N-G128A-Y217Q, T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G128A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-Y217Q, S053G-N061P-G097A-S101N-G128A-Y217Q-S249N, N025G-S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, S024G-T055P-G097A-G128A-Y217Q, T055P-N061P-G097A-A116S-G128A, S024G-N025G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A-Y217Q, S053G-T055P-G097A-S101N-G128A-Y217Q, T055P-G097A-S101N-G128A-Y217Q, S024G-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-G097A-S101N-G128A-Y217Q, G097A-G128S-Y217Q, S024G-S053G-N061P-G097A-G128A-Y217Q, S024G-N025G-N061P-G097A-G128A-Y217Q, S024G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-T055P-N061P-G097A-G128A-Y217Q, S024G-S038G-S053G-S078N-S101N-G128A-Y217Q, S053G-G097A-S101N-G128A-Y217Q, N025G-T055P-G097A-G128A-Y217Q, S024G-T055P-S078N-G097A-S101N-G128A-Y217Q, S053G-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-G097A-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-G128A-Y217Q, N025G-S078N-G097A-S101N-G128A-Y217Q, N025G-G097A-S101N-G128A-Y217Q, S024G-S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-G128A-Y217Q, and S024G-N025G-S053G-N061P-G097A-G128A-S130G-Y217Q; and
[0184] (b) a protease variant comprising an amino acid sequence having at least 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% identity to the amino acid sequence of SEQ ID NO:6, said variant having increased proteolytic activity and/or increased cleaning activity compared to SEQ ID NO:2, SEQ ID NO:4 and/or SEQ ID NO:6, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2 and wherein said variant comprises at least one set of amino acid substitution(s) relative to SEQ ID NO:6 selected from groups (i) through (xv):
[0185] (i) at least one substitution relative to SEQ ID NO:6 selected from the group consisting of A116V, G160S, I111L, I115V, N109S, N117M, P005G, Q059V, T164S, Y262M, A015Q, A015S, A098E, A098N, A098S, A098T, A098V, A098Y, A114S, A114T, A116G, A116L, A116S, A116T, A116W, A133G, A133H, A133T, A133V, A137G, A137I, A137L, A137S, A137T, A138S, A216E, A216F, A216V, D099S, D181E, F261A, F261Q, G024F, G024I, G024Q, G024Y, G097S, G160T, G211L, G211V, H017F, H017W, H039V, H226A, I031V, I111V, I268V, K170R, K265R, L016Q, L016T, L135M, L209T, L209V, L233M, L257T, L257V, L267A, L267V, N025A, N025I, N025Q, N025R, N025T, N025V, N101I, N101Q, N101S, N109A, N109G, N109H, N109L, N109M, N109Q, N109T, N117Q, N184A, N184L, N184T, N184W, N212G, N212L, N212V, N243P, N252G, N252M, P005T, P014S, P040G, P040L, P040Q, P129A, P129S, P172G, P172S, P194Q, P210A, P210S, Q185F, Q185G, Q185I, Q185M, Q185N, Q185S, Q275H, R186K, S009A, S009G, S009H, S009M, S018T, S130T, S132N, S145K, S159T, S161I, S161K, S161N, S161T, S162I, S162M, S162Y, S163G, S182F, S182G, S182V, S182W, S183F, S183L, S183M, S183T, S183V, S183W, S224A, S236T, S249V, T022A, T022G, T022Q, T022V, T208V, T242S, T253N, T253S, T254A, T254S, T255L, T255S, T255V, V004A, V004P, V004W, V084C, V139C, V165M, V203F, Y021K, Y021N, Y021T, Y021V, Y167F, Y171F, Y214F, Y262F, and Y262T;
[0186] (ii) at least one substitution relative to SEQ ID NO:6 selected from the group consisting of A216E, L090I, A098R, A098W, A098Y, A116G, A116R, A116S, A133M, I107L, I115V, M124L, N101I, N109H, N109S, N109T, N117R, P005G, Q185L, S089V, V095A, A015Y, A029G, A098D, A098E, A098G, A098N, A098S, A098T, A098V, A114S, A114T, A116E, A116L, A116T, A116V, A133H, A133L, A133S, A137G, A137I, A137L, A137S, A137V, A138S, A144S, A144V, A176S, A176T, A187T, A216F, A216P, A216Q, A216R, A216S, A216T, A216V, A216Y, D041E, D120A, D120E, D120Q, D120R, D120S, D181S, G020A, G020S, G024A, G097A, G097D, G097S, G131Q, G160S, G166I, G211L, G215N, H039N, H238N, I111L, I111V, I122A, L075I, L075Q, L135M, L209T, L209V, L233V, L235M, L235R, L257A, M119I, N025A, N025G, N025T, N061K, N101F, N101H, N101L, N101Q, N101R, N101S, N101T, N109A, N109G, N109K, N109L, N117E, N117H, N117K, N117S, N212G, N212S, N218F, N218G, N218H, N218L, N218S, N218W, N240Q, N252M, N252R, N252S, P005T, P040A, P040G, P040T, P129D, P129S, P194S, P210R, Q019R, Q019W, Q103L, Q103W, Q185A, Q185G, Q185M, Q185R, Q185T, Q206G, Q206Y, Q217A, Q217E, Q217R, Q217S, Q217T, S003Q, S009H, S018M, S033T, S130A, S130F, S130G, S130T, S130V, S145T, S159A, S161N, S161T, S162V, S162Y, S182L, S182W, S183F, S183L, S183V, S183W, S188K, S188W, S236Q, S236T, S248L, T022H, T022K, T208C, T253H, T255V, V044I, V121I, V139C, V143H, V143Q, V143T, V143W, V143Y, Y006K, Y021A, Y104W, A001F, A001G, A001H, A001K, A001L, A001Q, A001S, A001Y, A013V, A015G, A015K, A015R, A015S, A015T, A015W, A048S, A073N, A073S, A092S, A098K, A098P, A116D, A116W, A128S, A133P, A133T, A133V, A134G, A134S, A137H, A137N, A137T, A144D, A144K, A144L, A144M, A144N, A144R, A179G, A179S, A187V, A216G, A216L, A216W, A223S, A230C, A272K, A272L, A272P, A272S, A272T, A272W, A273G, A273S, A274G, A274M, A274T, D120K, D140E, D181A, D181E, D181G, D181H, D181T, D259E, D259N, D259Q, E054D, E156D, E156T, E251L, E251T, E251V, F058Y, F189W, F261K, F261Q, F261R, G007A, G007S, G020F, G020H, G020N, G020Q, G020T, G020Y, G024F, G024Q, G024R, G024T, G024V, G024W, G024Y, G053T, G097K, G097M, G097R, G097T, G131A, G131H, G131P, G131R, G131T, G131V, G160H, G160T, G166C, G166Q, G166S, G166T, G211A, G211D, G211K, G211M, G211N, G211Q, G211R, G211V, G211W, G215S, G215T, G215W, H017T, H017W, H017Y, H039V, H226A, H226F, H226I, H226L, H226M, H226V, I035V, I079A, 10795, I108V, I205V, I234L, I234V, I268V, K012S, K043P, K136H, K136R, K141A, K141F, K141T, K141W, K170A, K170G, K170R, K213A, K213R, K213S, K237A, K237H, K237L, K237S, K237V, K256A, K256G, K256H, K256M, K256P, K256Q, K256R, L016A, L016Q, L016T, L016V, L042V, L075M, L075T, L082M, L082V, L135F, L196I, L209H, L209Q, L209R, L209S, L209W, L233A, L233M, L233Q, L235I, L235K, L250I, L257S, L257T, L257V, L267A, L267Q, L267T, L267V, M119C, M199V, N025C, N025E, N025F, N025I, N025K, N025L, N025M, N025Q, N025V, N025Y, N061F, N061P, N061S, N061T, N076G, N078S, N101A, N109M, N109Q, N109R, N117A, N117M, N117Q, N118D, N118G, N118H, N118Q, N118R, N118S, N184A, N184C, N184G, N184L, N184R, N184S, N184T, N184V, N184W, N212C, N212F, N212I, N212K, N212L, N212P, N212Q, N212R, N212V, N212W, N212Y, N218A, N218P, N218T, N240A, N240E, N240G, N240H, N240L, N240R, N240S, N240T, N243C, N243Q, N243T, N243V, N252A, N252G, N252K, N252Q, P014G, P014Q, P014R, P014S, P014T, P040F, P040L, P040Q, P040S, P040V, P086C, P086H, P086S, P129A, P129E, P129G, P129K, P129R, P172A, P172K, P172Q, P172S, P194A, P194G, P194H, P194L, P194M, P194Q, P194R, P194V, P194W, P194Y, P210A, P210G, P210L, P210S, P239K, P239R, Q002A, Q002S, Q010A, Q010N, Q010R, Q010T, Q019A, Q019C, Q019D, Q019G, Q019S, Q019T, Q019V, Q059I, Q059V, Q103S, Q185F, Q185H, Q185I, Q185K, Q185N, Q185S, Q185Y, Q206H, Q206L, Q206P, Q206W, Q217F, Q217H, Q217I, Q217K, Q217L, Q217N, Q217V, Q271G, Q271R, Q271T, Q275F, Q275P, Q275R, R186A, R186I, R186K, S003A, S003F, S003G, S003H, S003K, S003R, S003T, S009T, S018N, S018T, S037G, S037T, S037V, S038G, S038Q, S063N, S063Q, S063T, S089M, S089N, S130K, S130L, S130R, S130W, S132N, S145G, S145K, S145M, S145R, S145V, S159C, S159H, S159L, S159Q, S159R, S159T, S159W, S161A, S161C, S161G, S161H, S161I, S161K, S161P, S161Q, S161R, S162F, S162G, S162I, S162L, S162M, S162N, S162P, S162R, S163G, S173A, S173G, S182F, S182G, S182K, S182N, S182Q, S182V, S183A, S183M, S183Q, S183R, S183T, S188A, S188F, S188G, S188P, S188R, S188T, S188V, S190C, S204A, S204G, S204I, S204L, S204Q, S204R, S204V, S207G, S224A, S224T, S236C, S236D, S236E, S236G, S236N, S248A, S248F, S248K, S248M, S248T, S249A, S249R, S249T, S249V, S249W, S249Y, S260G, S260H, S260K, S260N, T022A, T022G, T022Q, T022S, T022V, T022Y, T055A, T055K, T158A, T158S, T208L, T208S, T208V, T220S, T242D, T242N, T242S, T244E, T244G, T244I, I244R, T244V, T244W, T253A, T253G, T253N, T253S, T254S, T254V, T255H, T255I, T255K, T255L, T255Q, T255R, T255S, T255Y, V004A, V004N, V004P, V004W, V008A, V008M, V026I, V045S, V045W, V051I, V081Q, V081T, V084C, V093I, V095C, V143A, V143E, V143F, V143N, V143S, V147I, V147L, V147S, V147T, V148I, V149C, V149I, V165M, V180A, V180C, V180I, V180L, V180T, V192A, V192C, V192I, V192S, V192T, V192Y, V198L, V203A, V203F, V203K, V203L, V203M, V203N, V203Y, W241Y, Y006A, Y006G, Y006H, Y006L, Y006N, Y006P, Y006Q, Y006T, Y021E, Y021K, Y021L, Y021N, Y021Q, Y021R, Y021S, Y021T, Y104F, Y104I, Y104V, Y171F, Y214F, Y214L, Y214W, Y262F, and Y262S;
[0187] (iii) at least one set of amino acid substitutions relative to SEQ ID NO:6 selected from the group consisting of A088T-L257G, A116T-A128S, N061S-N109G-A128S-N243V-S260P, S009T-N109G-A128S-K141R-N243V, S009T-S018T-Y021N-N109G-A128S-K141R, and S162G-K256R;
[0188] (iv) at least one set of amino acid substitution(s) relative to SEQ ID NO:6 selected from the group consisting of A116T, A088T-N243V, G024E-A116T, K043Y, N076D-A116T, N218S-S248N, S033T-N243V, S033T-S063G, S248N-L257G, A001E-S249A, A088T-A116T, A088T-A128S, A088T-G131H, A088T-L257G, A088T-N109G, A088T-S248N, A088T-S249A, A116I-N243V, A116T-T158S, A128S, A128S-K256R, A128S-L257G, A128S-N243V, A128S-S248N, A128S-T158S, G024E-A088T, G024E-A128S, G024E-G131H, G024E-K256R, G024E-L257G, G024E-N218S, G024E-N243V, G024E-S162G, G024E-S249A, G024E-T158S, G131H, G131H-K256R, G131H-S249A, K043Y-A088T, K043Y-A116T, K256R, N076D-K256R, N109G, N109G-A116T, N109G-A128S, N109G-A128S-N243V-K256R, N109G-A128S-N243V-S248A, N109G-G131H, N109G-K256R, N109G-L257G, N109G-N218S, N109G-N243V, N109G-S248N, N218S-L257G, N243V, N243V-K256R, N243V-L257G, N243V-S248N, N243V-S249A, Q103H-A128S, Q103H-G131H, Q103H-K256R, Q103H-L257G, Q103H-N243V, Q103H-S248N, Q103H-S249A, Q103H-T158S, Q206D-N243V, S033T-A128S, S033T-K256R, S033T-N076D, S033T-N218S, S033T-S248N, S033T-T158S, S063G-A128S, S063G-K256R, S063G-N243V, S063G-S162G, S063G-T158S, S162G-K256R, S248N-K256R, S249A, T158S-N243V, and T158S-S249A;
[0189] (v) at least one set of amino acid substitution(s) relative to SEQ ID NO:6 selected from the group consisting of A088T-L257G, G024E-K256R, G024E-L257G, N109G-A116T, N109G-L257G, N243V-K256R, S033T-N109G, S033T-T158S, S063G-L257G, A001E-L257G, A088T-A128S, A088T-G169A, A088T-K256R, A088T-N109G, A088T-N218S, A088T-N243V, A088T-S248N, A088T-T158S, A116T, A116T-A128S, A116T-G131H, A116T-K256R, A116T-L257G, A116T-N218S, A116T-S162G, A116T-T158S, A128S, A128S-G169A, A128S-K256R, A128S-L257G, A128S-N218S, G024E, G024E-A128S, G024E-G131H, G024E-N109G, G024E-N243V, G024E-S033T, G024E-S063G, G024E-S248N, G024E-S249A, G024E-T158S, G131H, G131H-G169A, G131H-K256R, G131H-N218S, G131H-S249A, G169A, G169A-L257G, G169A-N243V, K043Y-A088T, K043Y-N109G, K256R, K256R-L257G, N061G-N109G-N243V, N076D-N109G, N109G, N109G-A128S, N109G-G131H, N109G-K256R, N109G-N218S, N109G-S162G, N109G-S248N, N109G-S249A, N109G-T158S, N218S, N218S-K256R, N218S-L257G, N218S-S248N, N243V, N243V-L257G, N243V-S248N, N243V-S249A, P040A-N109G-A128S-N243V-S248N-K256R, Q103H-K256R, Q103H-L257G, Q103H-N109G, S009T-S018T-Y021N-N109G-A128S-K141R, S033T-A088T, S033T-A116T, S033T-A128S, S033T-G131H, S033T-K043Y, S033T-K256R, S033T-L257G, S033T-N076D, S033T-N218S, S033T-N243V, S033T-Q103H, S033T-S063G, S033T-S162G, S033T-S248N, S033T-S249A, S063G, S063G-A088T, S063G-A116T, S063G-A128S, S063G-G131H, S063G-K256R, S063G-N109G, S063G-N218S, S063G-N243V, S063G-S248N, S063G-S249A, S063G-T158S, S162G-K256R, S162G-N218S, S162G-N243V, S162G-S248N, S162G-S249A, S248N, S249A, S249A-L257G, T158S, T158S-L257G, and T158S-N243V;
[0190] (vi) at least one set of amino acid substitution(s) relative to SEQ ID NO:6 selected from the group consisting of T158S-L257G, K256R, L257G, S033T-N109G, S162G-K256R, S162G-L257G, G024E-K256R, G024E-L257G, G024E-S033T, N109G-A116T, N218S-L257G, S033T-A088T, S033T-A116T, S033T-N243V, S033T-Q103H, S162G-N218S, S162G-N243V, T158S, T158S-N218S, T158S-N243V, A088T, A088T-G169A, A088T-K256R, A088T-L257G, A088T-S162G, A088T-T158S, A116T-K256R, A116T-L257G, A116T-N243V, A128S-L257G, A128S-N218S, A128S-N243V, A128S-S248N, G024E-A116T, G024E-A128S, G024E-G131H, G024E-N243V, G024E-S248N, G024E-S249A, G024E-T158S, G131H-N243V, G131H-T158S, G169A-N218S, G169A-N243V, G169A-S248N, K256R-L257G, N109G-A128S, N109G-G131H, N109G-N218S, N109G-N243V, N109G-S249A, N218S, N218S-K256R, N218S-N243V, N218S-S249A, N243V, N243V-K256R, N243V-L257G, N243V-S248N, Q103H-N109G, Q103H-N218S, S033T-A128S, S033T-L257G, S033T-N218S, S033T-S162G, S033T-S248N, S033T-T158S, S063G-K256R, S063G-L257G, S162G, S162G-G169A, S162G-S248N, S248N, S248N-K256R, S248N-L257G, S249A, T158S-S162G, and T158S-S248N;
[0191] (vii) at least one set of amino acid substitutions relative to SEQ ID NO:6 selected from the group consisting of S033T-N076D-A128S-N218S, A001E-S033T-N109G-N218S, S033T-N218S, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243V, A128S-G169A, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S018T-Y021N-S033T-N109G-A128S-N243V-S248N-K256R, S033T-A128S-G131H-N243P, P040E-N109G-A128S-G131H, S033T-A128S, S033T-N109G-A128S-N243V-S248N-K256R, N109G-G169A, S063G-N109G-A128S-G131H, G169A, N109G-A128S-G131H-N243V-S248N-K256R, S033T-A128S-G131H-N243V, A128S-N218S, A001E-G169A, A088T-G169A, G169A-L257G, N109G-N218S, S033T-N109G-A128S-N243P-S248N-K256R, G169A-K256R, N076D-G169A, A001E-G131H-G169A-N243V, G169A-S249A, S033T-N109G, G169A-S248N, K043Y-G169A, K043Y-N218S, N218S-L257G, N218S-N243V, S063G-G169A, A001E-A128S-G131H-N243V, A001E-S033T-N109G-N243V, A088T-N218S, G024E-N218S, G024E-S033T, G169A-Q206D, N076D-N218S, S033T-L257G, S162G-G169A, A001E-N218S, A116T-N218S, G169A-N243V, N218S, P040A-N109G-A128S-N243V-S248N-K256R, S033T-N076D, A001E-S033T, A128S-G131H, N218S-S248N, S018T-Y021N-N109G-A128S, S033T-K043Y, S033T-N243V, S033T-Q206D, S063G-N218S, S162G-N218S, T158S-G169A, A116T-G169A, G131H-G169A, N061S-N109G-A128S-S260P, N109G-A128S-N243V-K256R, N109G-A128S-N243V-S248A, N109G-A128S-N243V-S248A-K256R, N109G-A128S-N243V-S248N-K256R-L257G, N218S-K256R, S009T-N109G-A128S-K141R, S009T-S018T-Y021N-N109G-A128S-K141R, S033T-A088T, S033T-S063G, S033T-S162G, T158S-N218S, A001E-N076D-N109G-A128S, N109G-A128S-N243V-S248N-K256R, N109G-A128S-S248N-K256R, S009T-N109G-A128S-K141R-N243V, S018T-Y021N-N061S-N109G-A128S-S260P, S033T-A116T, S033T-S248N, S033T-S249A, S033T-T158S, G131H-N218S, N109A-A128S-N243V-K256R, N109G-A128S, N109G-A128S-S162G-N243V-S248N-K256R, N109G-A128S-T158S-N243V-S248N-K256R, N218S-S249A, Q206D-N218S, S018T-Y021N-N109G-A128S-N243V, S018T-Y021N-N109G-A128S-N243V-S248N-K256R, S033T-K256R, A116T-A128S, N061S-N109G-A128S-N243V-S260P, N109G-A128S-N243V-S248N, S009T-N109G-A128S-K141R-N243V-S248N-K256R, G024E-A128S, N061S-N109G-A128S-N243V-S248N-K256R-S260P, N109S-A128S-N243V-K256R, S033T, S033T-G131H, A001E-A128S, A128S, A128S-L257G, A128S-Q206D, N109Q-A128S-N243V-K256R, S009T-A128S-K141R-N243V, S009T-S018T-Y021N-A128S-K141R-N243V, A088T-A128S, A128S-K256R, A128S-N243V, N061P-N109G-N243V, N061S-A128S-N243V-S260P, S018T-Y021N-A128S-N243V, A128S-N243V-S248N-K256R, A128S-S248N, A128S-S249A, N076D-A128S, S063G-A128S, A128S-S162G, A128S-T158S, S018T-Y021N-N061S-A128S-N243V-S260P, S033T-Q103H-A128S-G131H, N061S-N109G-N243V, K043Y-A128S, N061P-N109G-G131H-N243V, N109G-L257G, A001E-G024E-S204E-Q206D, A001E-L257G, A088T-N109G, G024E-N109G, K043Y-N109G, N061G-N109G-N243V, N076D-N109G, N109G, N109G-A116T, N109G-K256R, N109G-N243V-K256R, N109G-N243V-S248A-K256R, N109G-Q206D, S063G-N109G, A001E-A116T, A001E-N109G, A001E-Q206D, A088T-A116T, A088T-N243V, A116T-L257G, G024E-A116T, G024E-L257G, G024E-N243V, G024E-Q206D, N109G-G131H, N109G-N243V, N109G-S162G, N109G-S248N, N109G-S248N-K256R, N109G-S249A, N109G-T158S, N243V-L257G, A001E-A088T, A001E-G024E, A001E-K256R, A001E-N076D, A001E-N243V, A088T, A088T-L257G, A088T-Q206D, A116T, A116T-K256R, A116T-N243V, G024E-A088T, G024E-K043Y, G024E-K256R, G024E-N076D, G024E-S162G, G024E-S248N, K043Y-A088T, K043Y-A116T, K043Y-L257G, K043Y-N243V, K043Y-Q206D, K256R-L257G, N076D-A116T, N076D-L257G, N076D-N243V, N076D-Q206D, N109G-N243V-S248N, N109G-N243V-S248N-K256R, N243V-K256R, Q206D, Q206D-L257G, Q206D-N243V, Q206D-S248N, S063G-K256R, S063G-L257G, T158S-L257G, A001E, A001E-K043Y, A001E-S162G, A001E-S248N, A001E-S249A, A001E-T158S, A088T-K256R, A088T-S162G, A088T-S248N, A088T-S249A, A116T-Q206D, A116T-S248N, A116T-S249A, G024E, G024E-G131H, G024E-S249A, G024E-T158S, G131H, G131H-K256R, G131H-L257G, K043Y-K256R, K043Y-N076D, K256R, L257G, N076D-A088T, N076D-K256R, N076D-S162G, N076D-S248N, N076D-S249A, N109G-N243P-S248A-K256R, N109G-N243P-S248N-K256R, N243V, Q206D-K256R, S033T-P040E-Q103H-N109G, S063G, S063G-A116T, S063G-Q206D, S162G-K256R, S162G-L257G, S162G-N243V, S162G-Q206D, S162G-S248N, S248N, S248N-L257G, S249A, S249A-L257G, T158S, T158S-N243V, and T158S-Q206D;
[0192] (viii) at least one set of amino acid substitution(s) relative to SEQ ID NO:6 selected from the group consisting of A088T-A116T-N243V-K256R-L257G, A088T-A116T-N243V-L257G, A088T-T158S-N218S-K256R, A088T-T158S-N218S-N243V-L257G, A088T-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-A153S-N218S-S248N-L257G, A088T-N109G-A116T-T158S-S248N-K256R-L257G, A088T-N109G-T158S-L257G, A114S-A116T-N218S-N243V-S248N-K256R-L257G, A116T-T158S-K256R, A088T-A116T-G131H-T158S-S248N-L257G, A088T-A116T-T158S, A088T-N109G-A116T-G131H-L257G, A088T-N109G-A116T-T158S-N243V-S248N-L257G, A088T-N109G-N243V-L257G, A088T-N109G-N243V-S248N, A088T-N109G-T158S-N243V-L257G, A088T-N109G-T158S-N243V-S248N-L257G, A116T-T158S-S248N-L257G, Y006H-A116T-G131H-S248N, A088T-A116T-G131H-T158S-N218S-N243V, A088T-A116T-G131H-T158S-N243V, A088T-A116T-G131H-T158S-N243V-K256R-L257G, A088T-A116T-N218S-N243V-K256R-L257G, A088T-A116T-S248N-K256R-L257G, A088T-A116T-T158S-N218S-N243V, A088T-A116T-T158S-N243V-K256R-L257G, A088T-A116T-T158S-N243V-S248N-L257G, A088T-G131H-N243V-S248N-K256R-L257G, A088T-N109G-A116T-T158S-L257G, A088T-N109G-A116T-T158S-N212D-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-T158S-S248N-L257G, A088T-N109G-G131H-V148A-N218S-N243V-K256R-L257G, A088T-N109G-K256R, A088T-N109G-N243V-S248N-L257G, A088T-N109G-T158S-K256R, A088T-N109G-T158S-N243V, A088T-T158S-N243V-K256R-L257G, A116T, A116T-N218S-N243V-L257G-N269S, A116T-T158S-K256R-L257G, N109G-A116T-K256R-L257G, N109G-A116T-N243V, N109G-A116T-T158S-N243V-K256R-L257G, N109G-G131H-L257G, N109G-G131H-S248N-K256R-L257G, N109G-G131H-T158S-K256R-L257G, S003P-A116T-T158S-S248N-K256R, T158S-S248N-K256R, A088T-A116T-G131H-N243V-K256R, A088T-A116T-G131H-S248N-K256R-L257G, A088T-A116T-G131H-V147A-T158S-N218S-N243V-S248N-L257G, A088T-A116T-S248N-L257G, A088T-A116T-T158S-N218S, A088T-A116T-T158S-N218S-K256R-L257G, A088T-A116T-T158S-N218S-L257G, A088T-G131H-N243V-L257G, A088T-G131H-T158S-S248N-L257G, A088T-L257G, A088T-N109G-A116T, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S-S248N-L257G, A088T-N109G-A116T-G131H-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N-K256R-L257G, A088T-N109G-A116T-N218S-N243V-K256R, A088T-N109G-A116T-N218S-N243V-L257G, A088T-N109G-A116T-N243V-S248N-K256R, A088T-N109G-A116T-N243V-S248N-K256R-L257G, A088T-N109G-A116T-T158S-L257G, A088T-N109G-A116T-T158S-N243V-L257G, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-W241R-S248N-K256R, A088T-N109G-K256R-L257G, A088T-N109G-L257G, A088T-N109G-N243V, A088T-N109G-N243V-K256R, A088T-N109G-N243V-K256R-L257G, A088T-N109G-S248N-K256R, A088T-N109G-T158S-N218S-K256R-L257G, A088T-N109G-T158S-N218S-N243V-S248N-K256R, A088T-N109G-T158S-N243V-K256R, A088T-N109G-T158S-N243V-K256R-L257G, A088T-N109G-T158S-N243V-S248N-A274D, A088T-N109G-T158S-S248N-L257G, A088T-T158S-K256R, A088T-T158S-N218S-N243V-K256R-L257G, A088T-T158S-N243V-L257G, A116T-G131H-N218S-N243V-S248N, A116T-G131H-S248N-L257G, A116T-S248N-K256R-L257G, A116T-T158S-N218S-N243V-K256R, A116T-T158S-N218S-S248N-L257G-Q271R, A116T-T158S-N243V-K256R-L257G, A116T-T158S-N243V-S248N-L257G, G131H-S248N, G131H-T158S-I234T-N243V-K256R, G131H-W241L-N243V-S248N-K256R, N109G-A116T-G131H-A137V-T158S-S248N-K256R-L257G, N109G-A116T-G131H-A151S-N218S-K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-K256R, N109G-A116T-G131H-T158S-N218S-S248N, N109G-A116T-G131H-T158S-N243V-S248N, N109G-A116T-S248N, N109G-A116T-T158S-L257G, N109G-A116T-T158S-N218S-W241R-N243V, N109G-A116T-T158S-N243V-S248N-L257G, N109G-A116T-T158S-S248N-K256R-L257G, N109G-A116T-T158S-S248N-L257G, N109G-G131H-N218S-L257G, N109G-G131H-N218S-S248N-K256R-L257G, N109G-G131H-T158S-N218S-S248N-K256R-L257G-A274T, N109G-K256R, N109G-N243V-L257G, N109G-T158S-N218S-K256R-L257G, N109G-T158S-N218S-L257G, N109G-T158S-S248N-K256R, P014L-A015L-L016C-H017T-S018L-Q019K-G020A-Y021T-T022L-G023E, S003F-A088T-N109G-A116T-T158S-N243V-K256R-L257G, V004A-A088T-A116T-T158S-N218S, V004A-N109G-A116T-G131H-S248N-K256R-L257G, V004L-A116T-N218S-N243V-S248N-L257G, Y006H-N109G-N218S-N243V-S248N, A001T-A116T-T158S-N243V-L257G, A088T-A116T, A088T-A116T-G131H-L257G, A088T-A116T-G131H-N218S-L257G, A088T-A116T-G131H-N218S-S248N-K256R-L257G, A088T-A116T-G131H-N218S-S248N-L257G, A088T-A116T-G131H-N243V-K256R-L257G, A088T-A116T-G131H-N243V-L257G, A088T-A116T-G131H-N243V-S248N, A088T-A116T-G131H-T158S-K256R-L257G, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S, A088T-A116T-G131H-T158S-N218S-N243V-K256R-A273T, A088T-A116T-G131H-T158S-N218S-S248N-K256R, A088T-A116T-G131H-T158S-N218S-S248N-L257G, A088T-A116T-G131H-T158S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-S248N, A088T-A116T-G131H-T158S-S248N-L257G, A088T-A116T-K256R, A088T-A116T-K256R-L257G, A088T-A116T-N218S-N243V-L257G, A088T-A116T-N243V-K256R, A088T-A116T-N243V-L257G, A088T-A116T-N243V-S248N-K256R-L257G, A088T-A116T-S248N-K256R, A088T-A116T-T158S-K256R, A088T-A116T-T158S-N218S, A088T-A116T-T158S-N218S-N243V-K256R, A088T-A116T-T158S-N218S-N243V-K256R-N269S, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-N243V-K256R, A088T-A116T-T158S-N243V-L257G, A088T-A116T-T158S-N243V-S248N-K256R, A088T-A116T-T158S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-S248N-K256R, A088T-A116T-V143A-N218S-S248N-K256R, A088T-A116T-V147I-T158S-N218S-N243V-L257G, A088T-G131H-K256R-L257G, A088T-G131H-N218S-N243V-S248N, A088T-G131H-N218S-S248N-L257G, A088T-G131H-S248N-K256R-L257G, A088T-G131H-T158S-L257G, A088T-G131H-T158S-N218S-K256R, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-N243V-L257G, A088T-G131H-T158S-N218S-S248N, A088T-G131H-T158S-N243V, A088T-G131H-T158S-N243V, A088T-G131H-T158S-N243V-S248N, A088T-G131H-T158S-N243V-S248N-K256R, A088T-G131H-T158S-N243V-S248N-L257G, A088T-I107T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-L257G, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V, A088T-N109G-A116T-G131H-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-L257G, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V, A088T-N109G-A116T-G131H-N243V-L257G, A088T-N109G-A116T-G131H-N243V-S248N-L257G, A088T-N109G-A116T-G131H-S248N, A088T-N109G-A116T-G131H-S248N-K256R, A088T-N109G-A116T-G131H-S248N-L257G, A088T-N109G-A116T-G131H-T158S-L257G, A088T-N109G-A116T-G131H-T158S-N218S, A088T-N109G-A116T-G131H-T158S-N218S-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218T-N243V, A088T-N109G-A116T-G131H-T158S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N-L257G, A088T-N109G-A116T-N218S, A088T-N109G-A116T-N218S-L257G, A088T-N109G-A116T-N218S-N243V, A088T-N109G-A116T-N218S-N243V-S248N-L257G, A088T-N109G-A116T-N218S-S248N-K256R, A088T-N109G-A116T-N218T-K256R, A088T-N109G-A116T-N218T-K256R-L257G, A088T-N109G-A116T-N243V, A088T-N109G-A116T-N243V-K256R-L257G, A088T-N109G-A116T-N243V-K256R-L257G-N269D, A088T-N109G-A116T-S248N-K256R, A088T-N109G-A116T-T158S, A088T-N109G-A116T-T158S-N218S-L257G, A088T-N109G-A116T-T158S-N218S-N243V, A088T-N109G-A116T-T158S-N218S-N243V-K256R, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-A116T-T158S-N218S-S248N, A088T-N109G-A116T-T158S-N243V, A088T-N109G-A116T-T158S-N243V-K256R, A088T-N109G-A116T-T158S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-S248N-L257G, A088T-N109G-G131H-L257G, A088T-N109G-G131H-N218S-K256R-L257G, A088T-N109G-G131H-N218S-N243V-K256R, A088T-N109G-G131H-N218S-N243V-L257G, A088T-N109G-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N243V, A088T-N109G-G131H-N243V-L257G, A088T-N109G-G131H-N243V-S248N-K256R, A088T-N109G-G131H-N243V-S248N-L257G, A088T-N109G-G131H-S248N-L257G, A088T-N109G-G131H-T158S-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N243V, A088T-N109G-G131H-T158S-N243V-K256R, A088T-N109G-G131H-T158S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N243V-L257G, A088T-N109G-L257G, A088T-N109G-N218S-K256R, A088T-N109G-N218S-N243V-S248N-L257G, A088T-N109G-N218S-S248N-K256R-L257G, A088T-N109G-N243V-K256R-L257G, A088T-N109G-N243V-S248N-K256R-L257G, A088T-N109G-N243V-S248N-L257G-I268V, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S-N218S-K256R, A088T-N109G-T158S-N218S-N243V-L257G, A088T-N109G-T158S-N218S-N243V-L257G, A088T-N109G-T158S-N243V-K256R-I268V, A088T-N109G-T158S-N243V-S248N-Q275R, A088T-N218S-N243V, A088T-N218S-N243V-S248N-K256R-L257G, A088T-N218S-S248N, A088T-N218S-S248N-L257G, A088T-N243V, A088T-N243V, A088T-N243V-K256R, A088T-N243V-L257G, A088T-S145T-T158S-S248N, A088T-T158S-L257G, A088T-T158S-N218S-S248N-L257G, A088T-T158S-N243V-K256R-L257G-Q271H, A088T-T158S-S248N, A088T-V143A-T158S-K256R, A116T-G131H-K256R, A116T-G131H-N218S, A116T-G131H-N243V, A116T-G131H-N243V-K256R, A116T-G131H-N243V-L257G, A116T-G131H-S248N-K256R, A116T-G131H-T158S-N218S-I234T-N243V-S248N-K256R, A116T-G131H-T158S-N243V-L257G, A116T-G131H-T158S-N243V-S248N-K256R, A116T-G131H-V143F-T158S-N218S, A116T-L257G, A116T-N218S, A116T-N218S-L257G, A116T-N218S-N243V-L257G, A116T-N243V, A116T-N243V-K256R, A116T-N243V-S248N, A116T-N243V-S248N-K256R-L257G, A116T-S248N, A116T-T158S, A116T-T158S-N218S-N243V, A116T-T158S-N218S-S248N, A116T-T158S-N243V, A116T-T158S-N243V-K256R, A116T-T158S-N243V-L257G, A116T-T158S-N243V-S248N, A116T-T158S-S248N-K256R-L257G, A116T-V149I-T158S-N243V-S248N-K256R-Q271H, G131H-N218S-N243V-L257G, G131H-N243V, G131H-N243V-S248N-K256R, G131H-T158S, G131H-T158S-N218S-N243V-K256R, G131H-T158S-N243V-K256R-L257G, G131H-T158S-N243V-S248N-L257G, N109G-A116T-G131H-N218S-K256R-L257G, N109G-A116T-G131H-N218S-L257G, N109G-A116T-G131H-N218S-N243V-K256R-L257G, N109G-A116T-G131H-N218S-S248N-K256R, N109G-A116T-G131H-N243V-K256R, N109G-A116T-G131H-N243V-L257G, N109G-A116T-G131H-N243V-S248N-K256R-L257G, N109G-A116T-G131H-S248N, N109G-A116T-G131H-S248N-I268V, N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, N109G-A116T-G131H-T158S-N218S-S248N-L257G, N109G-A116T-G131H-T158S-S248N, N109G-A116T-G131H-T158S-S248N-K256R, N109G-A116T-N218S, N109G-A116T-N218S-N243V-K256R, N109G-A116T-N218S-N243V-K256R-L257G, N109G-A116T-N218S-S248N-L257G, N109G-A116T-N243V-K256R, N109G-A116T-N243V-S248N-K256R-L257G, N109G-A116T-S248N-L257G, N109G-A116T-T158S-G211V-N243V-S248N-K256R, N109G-A116T-T158S-K256R-L257G, N109G-A116T-T158S-N218S, N109G-A116T-T158S-N218S-N243V-K256R-L257G, N109G-A116T-T158S-N218S-N243V-L257G, N109G-A116T-T158S-N218S-N243V-S248N-L257G, N109G-A116T-T158S-N218S-S248N-K256R-L257G, N109G-A116T-T158S-N243V, N109G-A116T-T158S-Q275R, N109G-G131H-A137V-T158S-N218S-S248N, N109G-G131H-N218S-K237N, N109G-G131H-N218S-N243V-K256R-L257G, N109G-G131H-N218S-S248N-K256R, N109G-G131H-N243V-K256R-L257G, N109G-G131H-S145F-N218S-N243V-K256R-L257G, N109G-G131H-S248N-K256R, N109G-G131H-S248N-L257G, N109G-G131H-T158S-K256R, N109G-G131H-T158S-N218S-N243V-K256R, N109G-G131H-T158S-N243V, N109G-G131H-T158S-N243V-K256R-L257G, N109G-G131H-T158S-N243V-L257G, N109G-G131H-T158S-S248N-L257G, N109G-G131H-T158S-S248N-Q271R, N109G-N218S-L257G, N109G-N218S-N243V, N109G-N243V-K256R-L257G, N109G-N243V-S248N-K256R-L257G, N109G-T158S-I268V, N109G-T158S-K256R, N109G-T158S-N218S-N243V-K256R-L257G, N109G-T158S-N218S-S248N-L257G, N109G-T158S-N243V, N109G-T158S-N243V-K256R-L257G, N109G-T158S-N243V-S248N, N109S-A116T-S248N, N218S, N218S-N243V-S248N-K256R-L257G, N218S-S248N-L257G, N243V, N243V-K256R, N243V-S248N-K256R, N243V-S248N-K256R-L257G, S105P-A116T-T158S-N218S-N243V-S248N-K256R, S248N, T158S-N243V-K256R, and T158S-N243V-L257G;
(ix) at least one set of amino acid substitution(s) relative to SEQ ID NO:6 selected from the group consisting of A088T-N109G-A116T-T158S-N243V-L257G, A116T-N218S-N243V-L257G-N269S, A088T-A116T-K256R, A088T-G131H-K256R, A088T-N109G-A116T-T158S-S248N-K256R-L257G, A088T-N109G-T158S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-K256R-A273T, A088T-A116T-N243V-L257G, A088T-A116T-S248N-K256R-L257G, A088T-A116T-T158S-N243V-L257G, A088T-A116T-T158S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-L257G, A088T-N109G-A116T-G131H-T158S-L257G, A088T-N109G-A116T-T158S-N218S-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N218S-K256R-L257G, A088T-N109G-N218S-S248N-L257G, A088T-T158S-N218S-N243V-K256R-I268V, A088T-T158S-N218S-S248N-L257G, A116T-N218S-K256R-L257G, N109G-A116T, N109G-A116T-G131H-T158S-L257G, N109G-A116T-N243V, N109G-A116T-N243V-K256R, N109G-A116T-T158S-L257G, N109G-K256R, N109G-N243V-K256R-L257G, S003P-N109G-G131H-T158S-K256R, A088T-A116T, A088T-A116T-G131H-N218S-K256R-L257G, A088T-A116T-G131H-N218S-L257G, A088T-A116T-G131H-N218S-N243V-S248N-L257G, A088T-A116T-G131H-N243V-K256R-L257G, A088T-A116T-G131H-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V, A088T-A116T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N, A088T-A116T-G131H-T158S-S248N-K256R-L257G, A088T-A116T-G131H-T158S-S248N-L257G, A088T-A116T-N218S-N243V-L257G, A088T-A116T-N218S-N243V-S248N-K256R-L257G, A088T-A116T-N218S-N243V-S248N-K256R-Q275R, A088T-A116T-T158S-A216S-N218S-N243V-K256R-L257G, A088T-A116T-T158S-K256R, A088T-A116T-T158S-N218S-L257G, A088T-A116T-T158S-N218S-N243V, A088T-A116T-T158S-N218S-N243V-K256R, A088T-A116T-T158S-N218S-N243V-K256R-L257G, A088T-A116T-T158S-N218S-N243V-K256R-N269S, A088T-A116T-T158S-N243V, A088T-A116T-T158S-N243V-K256R, A088T-A116T-V147I-T158S-N218S-N243V-L257G, A088T-G131H-K256R-L257G, A088T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-G131H-S248N-K256R-L257G, A088T-G131H-T158S-N218S-L257G, A088T-G131H-T158S-N218S-N243V-L257G, A088T-I107T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-I107T-N109G-G131H-N218S-S248N-K256R, A088T-N109G-A116T-G131H-A153S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-K256R-L257G, A088T-N109G-A116T-G131H-N218S-K256R-L257G, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V-K256R, A088T-N109G-A116T-G131H-N218S-N243V-L257G, A088T-N109G-A116T-G131H-N243V-L257G, A088T-N109G-A116T-G131H-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218S, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-N218S-K256R-L257G, A088T-N109G-A116T-N218S-L257G, A088T-N109G-A116T-N218S-N243V, A088T-N109G-A116T-N218S-N243V-L257G, A088T-N109G-A116T-N218T-K256R, A088T-N109G-A116T-N218T-K256R-L257G, A088T-N109G-A116T-N243V, A088T-N109G-A116T-N243V-K256R-L257G, A088T-N109G-A116T-N243V-K256R-L257G-N269D, A088T-N109G-A116T-T158S, A088T-N109G-A116T-T158S, A088T-N109G-A116T-T158S-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N, A088T-N109G-A116T-T158S-N243V-S248N-L257G, A088T-N109G-G131H-A138V-T158S-N218S-N243V-S248N-L257G, A088T-N109G-G131H-L257G, A088T-N109G-G131H-N218S, A088T-N109G-G131H-N218S-N243V-L257G, A088T-N109G-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N218S-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-K256R, A088T-N109G-G131H-T158S-N218S-N243V, A088T-N109G-G131H-T158S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N243V-L257G, A088T-N109G-G131H-T158S-N243V-S248N-L257G, A088T-N109G-G131H-V149A-K256R-L257G, A088T-N109G-N218S-N243V-L257G, A088T-N109G-N218S-N243V-S248N-L257G, A088T-N109G-N218S-S248N-K256R-L257G, A088T-N109G-N243V, A088T-N109G-N243V-K256R-L257G, A088T-N109G-N243V-L257G, A088T-N109G-N243V-S248N-L257G, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S-K256R, A088T-N109G-T158S-N218S-N243V-K256R-Q275R, A088T-N109G-T158S-N243V, A088T-N109G-T158S-N243V-K256R-I268V, A088T-N109G-T158S-N243V-L257G, A088T-N109G-T158S-N243V-S248N-L257G, A088T-N218S-N243V-L257G, A088T-N218S-N243V-S248N-K256R-L257G, A088T-N218S-S248N, A088T-T158S-N218S-N243V-K256R-L257G, A088T-V143A-T158S-K256R, A088T-V147I-N218S-N243V-K256R-L257G, A114S-A116T-N218S-N243V-S248N-K256R-L257G, A116T-G131H-N218S-N243V-S248N-K256R-L257G, A116T-G131H-N218S-N243V-S248N-L257G, A116T-G131H-N243V-K256R, A116T-G131H-N243V-L257G, A116T-G131H-N243V-S248N-K256R, A116T-G131H-S248N-L257G, A116T-G131H-T158S-N218S-I234T-N243V-S248N-K256R, A116T-G131H-T158S-N218S-N243V-S248N-L257G, A116T-N218S-L257G, A116T-T158S-N218S-K256R-L257G, A116T-T158S-N218S-S248N, A116T-T158S-N218S-S248N-L257G-Q271R, A116T-T158S-N243V-S248N-L257G, A116T-T158S-S248N-L257G, G131H-N218S-S248N-K256R-L257G, I107T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-G131H-A137V-T158S-S248N-K256R-L257G, N109G-A116T-G131H-A151S-N218S-K256R-L257G, N109G-A116T-G131H-N218S-K256R-L257G, N109G-A116T-G131H-N218S-N243V-L257G, N109G-A116T-G131H-N243V-L257G, N109G-A116T-G131H-S248N-L257G, N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-K256R-L257G, N109G-A116T-N218S-N243V-K256R, N109G-A116T-N218S-N243V-S248N-K256R-L257G, N109G-A116T-N218S-N243V-S248N-L257G, N109G-A116T-N243V-S248N-K256R, N109G-A116T-S248N-L257G, N109G-A116T-T158S-G211V-N243V-S248N-K256R, N109G-A116T-T158S-N218S, N109G-A116T-T158S-N218S-N243V-L257G, N109G-A116T-T158S-N218S-N243V-S248N, N109G-G131H-N218S-K237N, N109G-G131H-N218S-K256R-L257G, N109G-G131H-N218S-L257G, N109G-G131H-N218S-N243V-L257G, N109G-G131H-N243V-K256R-L257G, N109G-G131H-N243V-S248N-L257G, N109G-G131H-S248N-K256R, N109G-G131H-T158S-K256R-L257G, N109G-G131H-T158S-N218S-K256R-L257G, N109G-G131H-T158S-N243V, N109G-N218S, N109G-N218S-N243V-L257G, N109G-T158S-N218S-K256R-L257G, N109G-T158S-N218S-L257G, N109G-T158S-N218S-N243V-K256R, N109G-T158S-N243V-L257G, N243V-K256R-L257G, N243V-L257G, S003F-A088T-N109G-A116T-T158S-N243V-K256R-L257G, T158S-N218S-L233S, T158S-N218S-N243V-S248N-K256R, V004A-N109G-A116T-T158S-N218S-S248N-L257G, Y006H-A116T-G131H-T158S-N218S-N243V-S248N-K256R-A272G, A088T-A098S-N218S-K256R, A088T-A116T-G131H-K256R-L257G-L267M, A088T-A116T-G131H-L257G, A088T-A116T-G131H-N218S-A274T, A088T-A116T-G131H-N218S-K256R, A088T-A116T-G131H-N218S-N243V-K256R, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-N218S-N243V-L257G, A088T-A116T-G131H-N218S-N243V-S248N-K256R, A088T-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-N218S-S248N-L257G, A088T-A116T-G131H-N243V-K256R, A088T-A116T-G131H-N243V-L257G, A088T-A116T-G131H-N243V-S248N, A088T-A116T-G131H-N243V-S248N-A274V, A088T-A116T-G131H-N243V-S248N-K256R-L257G, A088T-A116T-G131H-S248N-K256R-L257G, A088T-A116T-G131H-S248N-L257G, A088T-A116T-G131H-T158S-K256R-L257G, A088T-A116T-G131H-T158S-K256R-L257G, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S, A088T-A116T-G131H-T158S-N218S-K256R, A088T-A116T-G131H-T158S-N218S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-K256R, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N218S-S248N-K256R, A088T-A116T-G131H-T158S-N218S-S248N-L257G, A088T-A116T-G131H-T158S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-N243V-L257G, A088T-A116T-G131H-T158S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-A116T-K256R, A088T-A116T-N218S-K256R, A088T-A116T-N218S-N243V-K256R, A088T-A116T-N218S-N243V-K256R-L257G, A088T-A116T-N218S-N243V-S248N-K256R, A088T-A116T-N218S-N243V-S248N-L257G, A088T-A116T-N243V-K256R, A088T-A116T-N243V-L257G, A088T-A116T-N243V-S248N-K256R, A088T-A116T-N243V-S248N-K256R-L257G, A088T-A116T-S248N, A088T-A116T-S248N-K256R, A088T-A116T-S248N-L257G, A088T-A116T-T158S-N218S, A088T-A116T-T158S-N218S-K256R, A088T-A116T-T158S-N218S-K256R, A088T-A116T-T158S-N218S-K256R-L257G, A088T-A116T-T158S-N218S-N243V-K256R, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N-K256R, A088T-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-N218S-N243V-S248N-L257G, A088T-A116T-T158S-N243V-K256R-L257G, A088T-G131H, A088T-G131H-N218S-K256R-L257G, A088T-G131H-N218S-N243V-K256R, A088T-G131H-N218S-N243V-K256R, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-S248N, A088T-G131H-N218S-S248N-K256R, A088T-G131H-N218S-S248N-K256R-L257G, A088T-G131H-N218S-S248N-L257G, A088T-G131H-N218T-L257G, A088T-G131H-N243V-L257G, A088T-G131H-N243V-S248N-K256R, A088T-G131H-S248N, A088T-G131H-S248N-L257G, A088T-G131H-T158S-N218S-K256R, A088T-G131H-T158S-N218S-K256R-L257G, A088T-G131H-T158S-N218S-N243V-K256R, A088T-G131H-T158S-N218S-N243V-K256R, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-S248N, A088T-G131H-T158S-N218S-S248N-K256R, A088T-G131H-T158S-N218S-S248N-K256R, A088T-G131H-T158S-N218S-S248N-L257G-I268V, A088T-G131H-T158S-N243V, A088T-G131H-T158S-N243V-S248N, A088T-G131H-T158S-N243V-S248N, A088T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-A116T, A088T-N109G-A116T-G131H-D140G-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-K256R, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S-K256R, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V, A088T-N109G-A116T-G131H-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N218S-S248N-L257G, A088T-N109G-A116T-G131H-N218S-S248N-L257G, A088T-N109G-A116T-G131H-N243V-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N, A088T-N109G-A116T-G131H-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N-L257G, A088T-N109G-A116T-G131H-S248N-K256R, A088T-N109G-A116T-G131H-S248N-K256R-L257G, A088T-N109G-A116T-G131H-S248N-L257G, A088T-N109G-A116T-G131H-T158S, A088T-N109G-A116T-G131H-T158S-N218S, A088T-N109G-A116T-G131H-T158S-N218S-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-S248N, A088T-N109G-A116T-G131H-T158S-N218S-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218T-N243V, A088T-N109G-A116T-G131H-T158S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-T158S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N-L257G, A088T-N109G-A116T-K256R, A088T-N109G-A116T-N218S, A088T-N109G-A116T-N218S-N243V, A088T-N109G-A116T-N218S-N243V-K256R, A088T-N109G-A116T-N218S-N243V-K256R-L257G, A088T-N109G-A116T-N218S-N243V-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N243V-S248N, A088T-N109G-A116T-N243V-S248N-K256R, A088T-N109G-A116T-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N243V-S248N-L257G, A088T-N109G-A116T-S248N-K256R, A088T-N109G-A116T-S248N-K256R-L257G, A088T-N109G-A116T-T158S-K256R, A088T-N109G-A116T-T158S-N218S, A088T-N109G-A116T-T158S-N218S-K256R-L257G, A088T-N109G-A116T-T158S-N218S-L257G, A088T-N109G-A116T-T158S-N218S-N243V, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-T158S-N218S-S248N, A088T-N109G-A116T-T158S-N218S-S248N-K256R, A088T-N109G-A116T-T158S-N218S-S248N-L257G, A088T-N109G-A116T-T158S-N243V-K256R, A088T-N109G-A116T-T158S-N243V-S248N, A088T-N109G-A116T-T158S-S248N, A088T-N109G-A116T-T158S-S248N-K256R, A088T-N109G-A116T-T158S-S248N-L257G, A088T-N109G-G131H-N218S-K256R, A088T-N109G-G131H-N218S-K256R, A088T-N109G-G131H-N218S-N243V-K256R, A088T-N109G-G131H-N218S-N243V-K256R, A088T-N109G-G131H-N218S-N243V-K256R-L257G, A088T-N109G-G131H-N218S-N243V-S248N-K256R, A088T-N109G-G131H-N218S-S248N, A088T-N109G-G131H-N218S-S248N-L257G, A088T-N109G-G131H-N243V-K256R-L257G, A088T-N109G-G131H-N243V-K256R-L257G, A088T-N109G-G131H-N243V-L257G, A088T-N109G-G131H-N243V-L257G, A088T-N109G-G131H-N243V-S248N-K256R, A088T-N109G-G131H-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N243V-S248N-L257G, A088T-N109G-G131H-S248N-L257G, A088T-N109G-G131H-T158S-K256R-L257G, A088T-N109G-G131H-T158S-N218S-L257G, A088T-N109G-G131H-T158S-N218S-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-G131H-T158S-N218S-S248N-L257G, A088T-N109G-G131H-T158S-N243V, A088T-N109G-G131H-T158S-N243V-S248N, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-S248N-K256R-L257G, A088T-N109G-G131H-T158S-W241R-S248N-K256R, A088T-N109G-G131H-V148A-N218S-N243V-K256R-L257G, A088T-N109G-G154A-N155P-E156T-G157L-T158M-S159E-G160E-S161L, A088T-N109G-N218S-K256R, A088T-N109G-N218S-N243V-L257G, A088T-N109G-N218S-N243V-S248N, A088T-N109G-N243V-K256R, A088T-N109G-N243V-S248N, A088T-N109G-N243V-S248N-K256R-L257G, A088T-N109G-S248N, A088T-N109G-T158S-N218S,
A088T-N109G-T158S-N218S-K256R, A088T-N109G-T158S-N218S-K256R-L257G, A088T-N109G-T158S-N218S-L257G, A088T-N109G-T158S-N218S-N243V-S248N-K256R, A088T-N109G-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-T158S-N218S-S248N-K256R, A088T-N109G-T158S-N218S-S248N-K256R-L257G, A088T-N109G-T158S-N243V-K256R, A088T-N109G-T158S-N243V-K256R, A088T-N109G-T158S-N243V-S248N-L257G, A088T-N109G-T158S-S248N-K256R-L257G, A088T-N109G-T158S-S248N-L257G, A088T-N109G-T158S-S248N-L257G, A088T-N109G-V147A-N218S-N243V-K256R, A088T-N218S-L257G-I268V, A088T-N218S-N243V, A088T-N218S-N243V-S248N, A088T-N218S-N243V-S248N-N269S, A088T-N218S-S248N-K256R, A088T-N218S-S248N-L257G-Q271R, A088T-N243V, A088T-N243V, A088T-N243V-K256R, A088T-N243V-S248N-K256R, A088T-N243V-S248N-K256R-L257G, A088T-S248N, A088T-T158S-N218S-K256R-L257G, A088T-T158S-N218S-N243V-K256R, A088T-T158S-N218S-N243V-L257G, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N243V-K256R-L257G, A088T-T158S-N243V-K256R-L257G-Q271H, A088T-T158S-N243V-S248N, A088T-T158S-N243V-S248N-L257G, A088T-T158S-S248N, A088T-V147A-K256R, A116T-G131H-K256R, A116T-G131H-N218S, A116T-G131H-N218S-K256R-L257G, A116T-G131H-N218S-L257G, A116T-G131H-N218S-N243V, A116T-G131H-N218S-S248N-K256R, A116T-G131H-N218S-S248N-K256R-L257G, A116T-G131H-N243V-S248N, A116T-G131H-N243V-S248N-L257G, A116T-G131H-S248N-K256R, A116T-G131H-T158S-A231V-N243V-L257G, A116T-G131H-T158S-N218S-K256R, A116T-G131H-T158S-N218S-K256R-L257G, A116T-G131H-T158S-N218S-N243V-K256R-L257G, A116T-G131H-T158S-N218S-N243V-S248N-K256R, A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A116T-G131H-T158S-N218S-S248N, A116T-G131H-T158S-N243V-L257G, A116T-G131H-T158S-N243V-S248N-K256R, A116T-G131H-T158S-S248N-K256R, A116T-G131H-T158S-S248N-L257G, A116T-G131H-V143F-T158S-N218S, A116T-K256R, A116T-N218S, A116T-N218S-K256R, A116T-N218S-N243V-L257G, A116T-N218S-N243V-S248N-K256R, A116T-N218S-N243V-S248N-K256R-L257G, A116T-N218S-N243V-S248N-L257G, A116T-N218S-S248N-L257G, A116T-N243V, A116T-N243V-K256R-L257G, A116T-N243V-S248N, A116T-N243V-S248N-K256R-L257G, A116T-S248N-K256R-L257G, A116T-T158S-K256R-L257G, A116T-T158S-N218S, A116T-T158S-N218S-K256R, A116T-T158S-N218S-N243V, A116T-T158S-N218S-N243V-S248N, A116T-T158S-N218S-S248N-K256R, A116T-T158S-N218S-S248N-K256R-L257G, A116T-T158S-N243V-K256R, A116T-T158S-N243V-L257G, A116T-T158S-N243V-S248N, A116T-T158S-S248N-K256R-L257G, G131H-K141R-T158S-N218S-K256R, G131H-N218S, G131H-N218S-K256R, G131H-N218S-N243V-K256R-L257G, G131H-N218S-N243V-S248N, G131H-N218S-N243V-S248N-L257G, G131H-N243V-S248N-K256R, G131H-N243V-S248N-K256R-L257G, G131H-S248N, G131H-T158S-I234T-N243V-K256R, G131H-T158S-N218S-K256R-L257G, G131H-T158S-N218S-N243V, G131H-T158S-N218S-N243V-S248N-L257G, G131H-T158S-N218S-S248N-K256R-L257G, G131H-T158S-N218S-S248N-L257G, G131H-T158S-N243V-K256R, G131H-T158S-N243V-S248N-L257G, N109G, N109G-A116T-G131H-A144V-T158S-S248N-K256R-L257G, N109G-A116T-G131H-K256R-L257G, N109G-A116T-G131H-N218S-N243V-K256R, N109G-A116T-G131H-N218S-N243V-K256R-L257G, N109G-A116T-G131H-N218S-S248N-K256R, N109G-A116T-G131H-N218S-S248N-L257G, N109G-A116T-G131H-N243V-K256R, N109G-A116T-G131H-N243V-S248N, N109G-A116T-G131H-N243V-S248N-K256R-L257G, N109G-A116T-G131H-S248N, N109G-A116T-G131H-S248N-K256R, N109G-A116T-G131H-T158S-K256R, N109G-A116T-G131H-T158S-K256R-L257G, N109G-A116T-G131H-T158S-N218S-K256R, N109G-A116T-G131H-T158S-N218S-K256R-L257G, N109G-A116T-G131H-T158S-N218S-L257G, N109G-A116T-G131H-T158S-N218S-N243V-K256R, N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, N109G-A116T-G131H-T158S-N218S-S248N, N109G-A116T-G131H-T158S-N243V-L257G, N109G-A116T-G131H-T158S-N243V-S248N, N109G-A116T-G131H-T158S-S248N, N109G-A116T-G131H-T158S-S248N-K256R, N109G-A116T-G131H-T158S-S248N-K256R-L257G, N109G-A116T-G131H-V149A-T158S-N218S-N243V-S248N-L257G, N109G-A116T-N218S, N109G-A116T-N218S-K256R, N109G-A116T-N218S-N243V-K256R-L257G, N109G-A116T-N218S-N243V-S248N-I268V, N109G-A116T-N243V-K256R-L257G, N109G-A116T-N243V-S248N, N109G-A116T-N243V-S248N-L257G, N109G-A116T-T158S, N109G-A116T-T158S-N218S-N243V-K256R-L257G, N109G-A116T-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-T158S-N218S-N243V-S248N-L257G, N109G-A116T-T158S-N218S-S248N-K256R-L257G, N109G-A116T-T158S-N243V-K256R-L257G, N109G-A116T-T158S-N243V-L257G, N109G-A116T-T158S-N243V-S248N-L257G, N109G-A116T-T158S-Q275R, N109G-A116T-T158S-S248N-K256R-L257G, N109G-G131H-L257G, N109G-G131H-N218S-N243V-S248N-K256R-L257G, N109G-G131H-N218S-N243V-S248N-L257G, N109G-G131H-N218S-S248N-K256R-L257G, N109G-G131H-N243V-K256R, N109G-G131H-S145F-N218S-N243V-K256R-L257G, N109G-G131H-S248N-K256R-L257G, N109G-G131H-S248N-L257G, N109G-G131H-T158S-K256R, N109G-G131H-T158S-N218S-L257G, N109G-G131H-T158S-N218S-N243V, N109G-G131H-T158S-N218S-N243V-K256R, N109G-G131H-T158S-N218S-N243V-K256R-L257G, N109G-G131H-T158S-N218S-N243V-S248N-L257G, N109G-G131H-T158S-N218S-S248N-K256R, N109G-G131H-T158S-N218S-S248N-K256R-L257G-A274T, N109G-G131H-T158S-N218S-S248N-L257G, N109G-G131H-T158S-N243V-S248N, N109G-G131H-T158S-N243V-S248N-K256R-L257G, N109G-G131H-T158S-S248N-K256R-L257G, N109G-G131H-T158S-S248N-L257G, N109G-N218S-K256R-L257G, N109G-N218S-L257G, N109G-N218S-N243V-K256R, N109G-N218S-N243V-S248N-S260F, N109G-N218S-S248N, N109G-N243V-K256R, N109G-N243V-L257G, N109G-N243V-S248N, N109G-N243V-S248N-K256R-L257G, N109G-S182F-S204F-S207L-N218S-S236F-S248N-L257G, N109G-S248N-K256R, N109G-T158S-K256R, N109G-T158S-N218S-N243V-K256R-L257G, N109G-T158S-N218S-S248N-L257G, N109G-T158S-N243V, N109G-T158S-N243V-K256R, N109G-T158S-N243V-S248N, N109G-T158S-N243V-S248N-K256R, N109G-T158S-S248N-K256R, N109G-T158S-S248N-L257G, N218S, N218S-N243V-S248N-K256R, N218S-S248N-L257G, N243V-K256R, N243V-S248N-K256R, N243V-S248N-K256R-L257G, N243V-S248N-L257G-Q271R, S003P-A116T-T158S-S248N-K256R, S248N, T158S-N218S, T158S-N218S-A272V, T158S-N218S-L257G, T158S-N218S-N243V-K256R-L257G, T158S-N218S-N243V-L257G, T158S-N218S-S248N-K256R-L257G, T158S-N243V-K256R, T158S-N243V-K256R-L257G, T158S-N243V-S248N-K256R, V004A-N109G-A116T-G131H-S248N-K256R-L257G, and Y006H-A116T-G131H-S248N;
[0194] (x) at least one set of amino acid substitutions relative to SEQ ID NO:6 selected from the group consisting of S018F-S162L, S018P-D120N, P014T-S037T, S009T-K141R, and S161P-S162L;
[0195] (xi) at least one set of amino acid substitutions selected relative to SEQ ID NO:6 from the group consisting of I031V-S038W, P014T-S037T, S018F-S162L, S018P-D120N, and S162L-D181H;
[0196] (xii) at least one set of amino acid substitutions relative to SEQ ID NO:6 selected from the group consisting of Y21H-D259G, S183T-S249R, N61D-Q206R, Y262N-Q275R, K043R-N076S, A133V-D259N, and I079V-Q217H;
[0197] (xiii) at least one set of amino acid substitutions relative to SEQ ID NO:6 selected from the group consisting of N61P-S63G-N109Q-A128S-S224A-N243V, A88T-N109G-A114S-A116T-A128S-N243V, A88T-N109G-A114S-A116T-A128S-S183L-S224A-N243V, N109G-A128S-S183V, N109G-A128S-N243V-K256R, N109M-A128S-S224A, A88T-N109S-A116T-A128S-S224A-N243V, N109Q-A128S-S224A-N243V, A88T-N109M-A116T-A128S-S224A-N243V, N109S-A128S-S224A-N243V, A88T-N109G-A116T-N243V, N101Q-N109Q-A128S-S224A-N243V, N109G-A116T-N243V-K256R, N109G-A128S-P129S-S130T-S224A-N243V, and A88T-N109Q-A116T-A128S-S224A-N243V;
[0198] (xiv) at least one set of amino acid substitutions relative to SEQ ID NO:6 selected from the group consisting of S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V, S33T-N61G-S63G-N109G-A128S-N218S-N243V, S33T-S63G-N109G-A128S-N218S-N243V, N61P-S63G-N109Q-A128S-G131H-G169A-S224A-N243V-S249Q, S33T-N61G-A88T-N109G-A116T-A128S-N218S-N243V, S33T-N109G-A128S-N218S-N243V, S33T-A128S-N218S, A1G-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-S224A-N243V-S249Q, S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-S224A-N243V, A88T-N109G-A114S-A116T-A128S-N243V, A88T-N109G-A114S-A116T-A128S-S183L-S224A-N243V, N109G-A114S-A128S, N109G-A114S-A128S-S183L-S224A, N109G-A114S-A128S-S224A, N109G-A114S-A128S-S224A-N243V, A88T-N109G-A116T-A128S-S224A-N243V, N61G-A88T-N109G-A116T-A128S-S224A-N243V, N109G-A128S-S183V, N109G-A114S-A128S-N243V, N109G-A128S-N243V-S248A, N109G-A128S-S224A-N243V, N109G-A128S-N243V-K256R, N109G-A128S-S224A, N109G-A128S-S183L-S224A, N61G-N109G-A128S-S224A, N109M-A128S-S224A, A88T-N109S-A116T-A128S-S224A-N243V, N109M-A128S-S224A-N243V, S63G-A128S, A88T-N109G-A116T-N243V, N101Q-N109Q-A128S-S224A-N243V, N109G-A116T-N243V-K256R, N109G-A116T, S63G-N109G, A88T-N109G, N109G-K256R, N61G-N109G-N243V, S33T-N109G-A128S-G169A-N218S-N243V, S33T-N109G-A128S-N218S-S224A-N243V, N109G-A128S-P129S-S130T-S224A-N243V, and A88T-N109Q-A116T-A128S-S224A-N243V; and
[0199] (xv) at least one set of amino acid substitutions relative to SEQ ID NO:6 selected from the group consisting of G24S-G53S-N78S-G97A-N101S-A128S, I31L-S33T-S63G-N109G-A128S-G169A-N218S-N243V, A1G-I31L-S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-G169A-S224A-N24- 3V-S249Q, S33T-N61G-S63G-N109G-A128S-G131H-G169A-N218S-N243V, S33T-S63G-N109G-A128S-G169A-N218S-N243V, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V-S249Q, S33T-N61G-S63G-N109G-A128S-N218S-N243V, S33T-S63G-N109G-A128S-N218S-N243V, S33T-T55P-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-G169A-S224A-N243V-S249Q, S33T-N61G-A88T-N109G-A116T-A128S-N218S-N243V, S33T-N109G-A128S-N218S-N243V, S33T-N76D-N109G-A128S-N218S-N243V, S33T-N76D-N109G-A128S-N218S-N243V-S248N-K256R, S33T-N61G-N109G-A128S-N218S-N243V, S33T-N76D-A128S-N218S, S33T-A128S-N218S, A1G-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-S224A-N243V-S249Q, N61P-S63G-N109Q-A128S-G131H-S224A-N243V, S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-S224A-N243V, A88T-N109G-A114S-A116T-A128S-N243V, A88T-N109G-A114S-A116T-A128S-S183L-S224A-N243V, N109G-A114S-A128S, N109G-A114S-A128S-S183L-S224A, N109G-A114S-A128S-S224A, N109G-A114S-A128S-S224A-N243V, A88T-N109G-A116T-A128S-S224A-N243V, N61G-A88T-N109G-A116T-A128S-S224A-N243V, N109G-A128S-S183V, N109G-A114S-A128S-N243V, N109G-A128S-N243V-S248A, N109G-A128S-S224A-N243V, N109G-A128S-N243V-K256R, N109G-A128S-S224A, N109G-A128S-S183L-S224A, N61G-N109G-A128S-S224A, N76D-N109G-A128S-S224A, N109M-A128S-S224A, N109G-A128S-S183L, S33T-N76D, A88T-N109S-A116T-A128S-S224A-N243V, N109Q-A128S-S224A-N243V, N109S-A128S-S224A, A88T-N109M-A116T-A128S-S224A-N243V, N101Q-N109Q-A128S-P129S-S130T-S224A-N243V, S63G-N109Q-A128S-S224A-N243V, N109M-A128S-S224A-N243V, S63G-A128S, N109S-A128S-S224A-N243V, A88T-N109G-A116T-N243V, N61S-N109G-N243V, N101Q-N109Q-A128S-S224A-N243V, N109G-A116T-N243V-K256R, A88T-N109G-A116T-T158S-N243V-K256R, N109G-A116T, S63G-N109G, A88T-N109G, N109G-K256R, N61G-N109G-N243V, S33T-N61P-S63G-N109G-A128S-G131H-G169A-N218S-N243V, S33T-N109G-A128S-G169A-N218S-N243V, S33T-N109G-A128S-N218S-S224A-N243V, N109G-A128S-P129S-S130T-S224A-N243V, and A88T-N109Q-A116T-A128S-S224A-N243V.
[0200] In an eighth aspect, the invention provides an isolated or non-naturally occurring protease variant of a BPN' subtilisin protease having the amino acid sequence of SEQ ID NO:2, said variant having proteolytic activity and comprising at least one set of amino acid substitution(s) selected from the group consisting of: S182E, N109I, N109D-Y217L-S248R, N109D-S188R-Y217L, S87D-Y217L-S248R, S87R-N109D-Y217L-S248R, S87R-N109D-S188D-Y217L-S248R, G128A-Y217Q, I111V-M124V, M124V-Y217Q, N62Q-G97A, S89Y-M124V, V68A, V68A-A92G, V68A-G97A, V68A-I111V, V68A-S89Y, V68A-V227T, V68A-Y217Q, W106F-Y217Q, G97A-G128A-Y217Q, G97A-L126A-Y217Q, G97A-M124V-L126A-Y217Q, G97A-N123G-Y217Q, L96T-G97A-Y217Q, M124V-L126A-Y217Q, N62Q-G128A-Y217Q, N62Q-G97A-Y217Q, G97N-G128A-Y217M, G97G-G128S-Y217E, G97A-G128A-Y217Q, G97M-G128S-Y217E, G97A-G128S-Y217Q, G97D-G128S-Y217Q, G97M-G128G-Y217M, G97G-G128S-Y217Q, G97S-G128S-Y217Q, G97G-G128A-Y217Q, G97S-G128A-Y217E, G97A-G128S-Y217L, G97A-G128A-Y217N, G97Q-G128S-Y217L, G97A-G128A-Y217M, G97A-G128A-Y217S, G97D-G128A-Y217Q, G97M-G128S-Y217Q, G97Q-G128G-Y217D-S87Y, G97S-G128A-Y217N, G97A-G128S-Y217T, G97D-G128S-Y217E, G97D-G128A-Y217L, G97G-G128S-Y217E-S78P-A272T, G97T-G128S-Y217D, G97D-G128A-Y217I, G97Q-G128S-Y217Q, G97G-G128A-Y217D, G97Q-G128A-Y217N, G97S-G128A-Y217M, G97S-G128S-Y217N, G97S-G128S-Y217M, G97E-G128S-Y217M, G97S-G128P-Y217Q, G97T-G128S-Y217Q, G97D-G128S-Y217Q-A73T, G97E-G128S-Y217N, G97G-G128A-Y217I, G97Q-G128A-Y217D, G97Q-G128S-Y217M, G97R-G128T-Y217Q-S162P, G97S-G128S-Y217D, G97T-G128P-Y217I, G97Q-G128G-Y217E, G97C-G128G-Y217N, G97D-G128S-Y217H, G97M-G128S-Y217L, G97M-G128S-Y217N, G97S-G128S-Y217E, G97M-G128S-Y217I, G97A-G128P-Y217A, G97R-G128S-Y217D, G97A-G128A-Y217Q-S145D, G97A-G128A-Y217Q-P239R, G97A-G128A-Y217Q-N61E-P129E-S162K-K213L-N240K, G97A-G128A-Y217Q-N61E, G97A-G128A-Y217Q-P40E-A144K-K213L, G97A-G128A-Y217Q-P129E, G97A-G128A-Y217Q-N61E-P129E-S159K, G97A-G128A-Y217Q-K213L, G97A-G128A-Y217Q-S87D, G97A-G128A-Y217Q-Q206E, G97A-G128A-Y217Q-S24R-P40E-S145D-S159K-K213L, G97A-G128A-Y217Q-K265N, G97A-G128A-Y217Q-S24R, G97A-G128A-Y217Q-P40E, G97A-G128A-Y217Q-Q275E, G97A-G128A-Y217Q-P129E-S145D-N240K, G97A-G128A-Y217Q-A144K, G97A-G128A-Y217Q-S159K, G97A-G128A-Y217Q-S162K, G97A-G128A-Y217Q-N240K, G97A-G128A-Y217Q-S53G, G97A-G128A-Y217Q-S78N, G97A-G128A-Y217Q-S53G-S78N, G97A-G128A-Y217Q-S53G-I111V, G97A-G128A-Y217Q-S53G-N117S, G97A-G128A-Y217Q-S53G-S132N, G97A-G128A-Y217Q-Y104N-S132N, G97A-G128A-Y217Q-S53G-S78N-I111V, G97A-G128A-Y217Q-S53G-S78N-N117S, G97A-G128A-Y217Q-S53G-S78N-S132N, G97A-G128A-Y217Q-S53G-Y104N-S132N, G97A-G128A-Y217Q-S78N-Y104N-S132N, Y217L-V068C-A069G, Y217L-I079F-A098G, Y217L-P086T-S101D-Q103S-V147I, Y217L-A088T-P129S-G146D, Y217L-V093I-G128D-P129R, Y217L-Z096.01D-A098R, Y217L-Z096.01H-A098G, Y217L-G097S-Z097.01S-A098G-A273T, Y217L-A098S-D099G-G100D, Y217L-Z098.01N, Y217L-D099G-Z099.01N, Y217L-D099G-Z099.01S, Y217L-D099V-S101D, Y217L-Z099.01S, Y217L-G100D, Y217L-S101D-Q103H, Y217L-S101G-A151V, Y217L-S101H-G102S, Y217L-S101H-Q103D, Y217L-G102R-Q103C-Y104C-V192I, Y217L-Q103D, Y217L-V121I-I122S-N123C, Y217L-V121L-N123C, Y217L-I122S-N123S, Y217L-M124I, Y217L-M124V, Y217L-L126F-P129Z-S182N, Y217L-L126Y, Y217L-G127S-P129D, Y217L-Z127.01N-G128S-P129S, Y217L-G128H-P129Y, Y217L-G128S-P129D, Y217L-G128S-P129D-S248R, Y217L-G128S-P129G, Y217L-P129G-G131Z, Y217L-P129G-S130H-S132Z, Y217L-P129H-G131Z, Y217L-P129L, Y217L-P129S-S130H-S132Z, Y217L-P129Z, Y217L-P129Z-S130G, Y217L-P129Z-S130G-G131H-S132H, Y217L-P129Z-S130H, Y217L-S130V-G131D-S132I, S87T-A88L-S89G-G97A-G128A-Y217Q, N61P-S63H-G97A-G128A-Y217Q, S87G-A88V-S89A-G97A-G128A-Y217Q, P86S-S87G-A88V-G97A-G128A-Y217Q, Q59S-N61P-G97A-G128A-Y217Q, S24G-N25G-G97A-G128A-Y217Q, N61P-N62S-G97A-G128A-Y217Q, G97A-G128A-P129Q-S130G-G131S-Y217Q, L75S-N76Y-G97A-G128A-Y217Q, G97A-G128A-V203Y-Y217Q, T55P-G97A-G128A-Y217Q, A88V-L901-G97A-G128A-Y217Q, G97A-G128A-G211R-N212S-K213V-Y217Q, G23A-S24G-N25G-G97A-G128A-Y217Q, T22N-S24A-G97A-G128A-Y217Q, S24R-G97A-G128A-Y217Q, G97A-A98S-G128A-Y217Q, G97A-G128A-T158G-S159G-Y217Q, Q59E-N61P-G97A-G128A-Y217Q, G97A-A98E-G128A-Y217Q, G97A-G128A-Y217Q-P86S-S87G-A88V-A116N-N117S-N118G, G97A-G128A-Y217Q-S63T-P86S-S87G-A88V, G97A-G128A-Y217Q-P86S-S87G-A88V-P239R, G97A-G128A-Y217Q-S24G-N25G-N61P-N62S-P194L-A232T, G97A-G128A-Y217Q-P129Q-S130G-G131S-A133V-L267V, G97A-G128A-Y217Q-A134T-L267V, G97A-G128A-Y217Q-S24R-P40E-P129E-S159K-K265R, G97A-G128A-Y217Q-A134T-G211T, G97A-G128A-Y217Q-S24R-P129E, G97A-G128A-Y217Q-I111V-S161P, G97A-G128A-Y217Q-T55P-P129Q, G97A-G128A-Y217Q-I115V-L267V, G97A-G128A-Y217Q-P86S-S87G-A88V-A116S-N117G-N118R, G97A-G128A-Y217Q-V203Y-L267V, G97A-G128A-Y217Q-S24G-N25G-S78N-S101N-V203Y, G97A-G128A-Y217Q-P525-T55P-V203Y, G97A-G128A-Y217Q-Q59S-N61P-A116S-N117G-N118R, G97A-G128A-Y217Q-S24G-N25G-P129Q-S130G-G131S, G97A-G128A-Y217Q-P86S-S87G-A88V-T242R, G97A-G128A-Y217Q-P40E-T55P-N269K, G97A-G128A-Y217Q-G23A-S24G-N25G-A116N-N117S-N118G, G97A-G128A-Y217Q-V8L-N25Y-P129Q-S130G-G131S, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-G211T-L267V, G97A-G128A-Y217Q-S24R-A116N-N117S-N118G, G97A-G128A-Y217Q-S24R-A128S-P129G, G97A-G128A-Y217Q-P129Q-S130G-G131S-N240K, G97A-G128A-Y217Q-N25Y-P129Q-S130G-G131S, G97A-G128A-Y217Q-S87T-A88L-S89G-A134T, G97A-G128A-Y217Q-P129Q-S130G-G131S-L267V, G97A-G128A-Y217Q-S87G-A88V-S89A-A116N-N117S-N118G, G97A-G128A-Y217Q-N61P-P129Q-S130G-G131S, G97A-G128A-Y217Q-N61P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-T55P-P129V-P194S, G97A-G128A-Y217Q-T55P-P129V, G97A-G128A-Y217Q-S24G-N25G-T55P-S78N-S101N, G97A-G128A-Y217Q-T55P-S78N-I115V, G97A-G128A-Y217Q-N25Y-S87G-A88V-S89A, G97A-G128A-Y217Q-A134T-N240K, G97A-G128A-Y217Q-S24R-Q59S-N61P, G97A-G128A-Y217Q-G23A-S24G-N25G-P239R, G97A-G128A-Y217Q-T55P-A116S-N117G-N118R, G97A-G128A-Y217Q-A134T-S161P, G97A-G128A-Y217Q-S24G-N25G-S53G-N61P-S101N-V203Y, G97A-G128A-Y217Q-N25Y-Q59S-N61P, G97A-G128A-Y217Q-N25Y-P129Q-S130G-G131S-A137T, G97A-G128A-Y217Q-G23A-S24G-N25G-N61P-S63H, G97A-G128A-Y217Q-T55P-N61P-S78N-S101N-V203Y, G97A-G128A-Y217Q-P129Q-N240K, G97A-G128A-Y217Q-T55P-A134T, G97A-G128A-Y217Q-N25Y-N61P-S63H, G97A-G128A-Y217Q-S87T-A88L-S89G-P129S, G97A-G128A-Y217Q-T55P-L75H-N76G, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-T55P-I115V, G97A-G128A-Y217Q-T55P-A116N-N117S-N118G, G97A-G128A-Y217Q-S24G-N25G-A116N-N117S-N118G, G97A-G128A-Y217Q-S24R-P129Q-S130G-G131S, G97A-G128A-Y217Q-G23A-S24G-N25G-G211R-N212S-K213V, G97A-G128A-Y217Q-S24G-N25G-T55P-N61P-S78N-S101N-V203Y, G97A-G128A-Y217Q-T55P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-I115V-A273S, G97A-G128A-Y217Q-N25Y-T55P, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N-V203Y- , G97A-G128A-Y217Q-Q59S-N61P-N240K, G97A-G128A-Y217Q-S161P-L267V, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-S24G-N25G-N61P-S101N, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S101N-V203Y, G97A-G128A-Y217Q-N240K, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-T55P-S101N, G97A-G128A-Y217Q-N61P-S63H-A128S-P129Q, G97A-G128A-Y217Q-S89Y-P129Q-S130G-G131S, G97A-G128A-Y217Q-P129Q-S130G-G131S-V203Y, G97A-G128A-Y217Q-I115V-N240K, G97A-G128A-Y217Q-S53G-N61P-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S161P-V203Y, G97A-G128A-Y217Q-S87T-A88L-S89G-N240K, G97A-G128A-Y217Q-S87T-A88L-S89G-P239R, G97A-G128A-Y217Q-T55P-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-I115V-A134T, G97A-G128A-Y217Q-Y6Q-P129Q-S130G-G131S, G97A-G128A-Y217Q-T55P-S78N-S89Y, G97A-G128A-Y217Q-S24G-N25G-T55P-S78N-A88V-S101N, G97A-G128A-Y217Q-N61P-S63H-S78N-I111V-A134T, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S78N-S101N, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S101N-V203Y, G97A-G128A-Y217Q-S53G-N61P-S101N-V203Y, G97A-G128A-Y217Q-S53G-T55P-S78N-S101N-V203Y, G97A-G128A-Y217Q-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-N240K-A273S, G97A-G128A-Y217Q-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-Q59S-N61P-S87T-A88L-S89G, G97A-G128A-Y217Q-N61P-S63H-S78N-S161P, G97A-G128A-Y217Q-N61P-S63H-S78N-I111V, G97A-G128A-Y217Q-T55P-A128S-P129Q, G97A-G128A-Y217Q-N61P-S78N-S101N-V203Y, G97A-G128A-Y217Q-N61E-A144K, G97A-G128A-Y217Q-A134T-P239R, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S101N-V203Y, G97A-G128A-Y217Q-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-T55P-N240K, G97A-G128A-Y217Q-S24G-N25G-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-P129Q-S130G-G131S-P239R, G97A-G128S-Y217Q, G97A-G128A-Y217Q-S53G-N61P-S101N, G97A-G128A-Y217Q-I111V-P129Q-G211T, G97A-G128A-Y217Q-S24G-N25G-S53G-S101N-V203Y, G97A-G128A-Y217Q-Q59S-N61P-S87G-A88V-S89A, G97A-G128A-Y217Q-S24G-N25G-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-P129Q-S130G-G131S-S162K, G97A-G128A-Y217Q-T55P-P129Q-S130G-G131S, G97A-G128A-Y217Q-T55P-V203Y, G97A-G128A-Y217Q-S87G-A88V-S89A-P129Q-S130G-G131S, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G, G97A-G128A-Y217Q-I111V-P129Q-S130G-G131S, G97A-G128A-Y217Q-I111V-A273S, G97A-G128A-Y217Q-N61P-S87T-A88L-S89G, G97A-G128A-Y217Q-T22N-S24A-N61P-S63H, G97A-G128A-Y217Q, G97A-G128A-Y217Q-S53G-S78N-S87T-A88L-S89G-S101N-P129S-V203Y, G97A-G128A-Y217Q-S159K-L267V, G97A-G128A-Y217Q-P40E-S53Y-S78Y-P86S-S87G-A88V, G97A-G128A-Y217Q-S24R-S145D, G97A-G128A-Y217Q-I111V-S159K, G97A-G128A-Y217Q-T55P-P129L, G97A-G128A-Y217Q-Q59S-N61P-V203Y, G97A-G128A-Y217Q-T55P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S101N, G97A-G128A-Y217Q-S53G-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S89Y, G97A-G128A-Y217Q-S24R-P129V, G97A-G128A-Y217Q-S87G-A88V-S89A-A116N-N117S-N118G-P172H, G97A-G128A-Y217Q-I111V-A134T, G97A-G128A-Y217Q-Q59S-N61P-P129Q-S130G-G131S, G97A-G128A-Y217Q-P5S-S87G-A88V-S89A-A116G-N117R, Y217Q-N61P-A97G-G102A-A128G-P129S, G97A-G128A-Y217Q-S24G-N25G-S53G-N61P-S78N, G97A-G128A-Y217Q-S145D-S159K-N240K-Q275E, G97A-G128A-Y217Q-T55P-A128S-P129D, G97A-G128A-Y217Q-G23A-S24G-N25G-A128S-P129D, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S89G-V203Y, G97A-G128A-Y217Q-I111V-P239R, G97A-G128A-Y217Q-S87G-A88V-S89A-S162K, G97A-G128A-Y217Q-S87T-A88L-S89G-I115V, G97A-G128A-Y217Q-S24G-N25G-T55P-S78N, G97A-G128A-Y217Q-T55P-A92G, G97A-G128A-Y217Q-S24G-N25G-S53G-S87T-A88L-S89G-V203Y, G97A-G128A-Y217Q-T22N-S24A-T55P, G97A-G128A-Y217Q-S53G-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S78N-S87T-A88L-S89G, G97A-G128A-Y217Q-P129Q-S130G-G131S-S159K, G97A-Y217Q-N61P-N62Q-G100N-A128G, G97A-G128A-Y217Q-S24R-S78N-S182P-L267V, G97A-G128A-Y217Q-P239R-A273S, G97A-G128A-Y217Q-S53G-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-P129Q-S130G-G131S-T242R, G97A-G128A-Y217Q-S3F-S87T-A88L-S89G-G211T, G97A-G128A-Y217Q-S24G-N25G-L75H-N76G, G97A-G128A-Y217Q-S53G-T55P-N61P-S78N-S87T-A88L-S89G, G97A-G128A-Y217Q-S87T-A88L-S89G-A144K, G97A-G128A-Y217Q-S78N-S87T-A88L-S89G-V203Y, G97A-G128A-Y217Q-Q59S-N61P-A116N-N117S-N118G, G97A-G128A-Y217Q-S87T-A88L-S89G-I111V, G97A-G128A-Y217Q-S24R-S145D-P239R-Q275E, G97A-G128A-Y217Q-S145D-A273S, G97A-G128A-Y217Q-S24G-N25G-K141E-T242R, G97A-G128A-Y217Q-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-A116N-N117S-N118G-P129Q-S130G-G131S, G97A-G128A-Y217Q-S89Y-G211T, G97A-G128A-Y217Q-S87G-A88V-S89A-A116N-N117S-N118G-A144T, G97A-G128A-Y217Q-S24G-N25G-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-P129V, G97A-Y217Q-N61P-A128G-P129S-S130P, G97A-G128A-Y217Q-T55P-N61P-S87T-A88L-S89G-G110C-S130P, G97A-Y217Q-N123G-A128G, G97A-G128A-Y217Q-N61P-N62Q-G100N-G102A-M124I, S78N-G97A-G128A-Y217Q, G97A-S101N-G128A-Y217Q, G97A-G128A-A137V-Y217Q, N61P-G97A-G128A-Y217Q, G97A-G128A-S130P-Y217Q, G97A-Q103N-G128A-Y217Q, S63T-G97A-G128A-Y217Q, G97A-G102A-G128A-Y217Q, G97A-N109D-G128A-Y217Q-S248R, S87R-G97A-G128A-Y217Q, G97A-G128A-S188D-Y217Q, S87D-G97A-G128A-Y217Q-S248R, G97A-G128A-S188D-S248R-Y217Q, G97A-G128A-S248D-Y217Q, S78N-G97A-G128A-Y217Q-L267V, S78N-G97A-G128A-Y217Q-S161P, S78N-G97A-G128A-Y217Q-I115V, S78N-G97A-G128A-Y217Q-A273S, S78N-G97A-G128A-Y217Q-G211T, S78N-G97A-G128A-Y217Q, S78N-G97A-G128A-Y217Q-I111V, S78N-G97A-G128A-Y217Q-V147L, S78N-G97A-G128A-Y217Q-I108V, S78N-G97A-G128A-Y217Q-S89Y, S78N-G97A-G128A-Y217Q-A138T, G97A-G128A-Y217Q-A134T-K213L, G97A-G128A-Y217Q-G23A-S24G-N25G-P129V, G97A-G128A-Y217Q-S24R-P239R, and G97A-G128A-Y217Q-S24R-S87T-A88L-S89G, wherein amino acid positions of the variant are numbered by correspondence with the amino acid sequence of SEQ ID NO:2.
[0201] A variant according to the eighth aspect of the invention may have increased proteolytic activity and/or increased cleaning activity compared to the protease having the sequence of SEQ ID NO:2 and/or SEQ ID NO:4 and may be in a mature form. A variant according to the eighth aspect of the invention may comprise an amino acid sequence having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:4, wherein the variant has proteolytic activity and optionally has enhanced proteolytic activity and/or enhanced cleaning activity compared to the protease having the sequence of SEQ ID NO:2 or SEQ ID NO:4.
[0202] A variant according to the eighth aspect of the invention may comprise three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14 or 15 amino acid substitutions at positions selected from the group of consisting of positions 24, 25, 40, 52, 53, 55, 58, 59, 61, 62, 63, 68, 78, 86, 87, 88, 89, 92, 96, 97, 100, 101, 103, 104, 106, 111, 114, 115, 116, 117, 118, 123, 124, 125, 126, 128, 129, 130, 131, 132, 133, 134, 144, 145, 159, 161, 162, 167, 194, 203, 206, 213, 217, 227, 232, 239, 240, 242, 265, 267, and 275. A variant according to the eighth aspect may comprise a total of three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14 or 15 amino acid substitutions selected from groups (a) and (b): (a) charged amino acid substitutions selected from the group consisting of N61E, A144K, P129E, P239R, P40E, Q103E, Q206E, Q7275E, S145D, S159K, S162K, S24R, S63H, S87D and T242R; and (b) neutral amino acid substitutions selected from the group consisting of A114G, A116N, A133V, A134T, A232T, A88V, A92G, F58G, G100T, G128A, G131S, G97A, I111V, I115V, K213L, K265N, L126A, L267V, L96T, M124V, N117S, N118G, N123G, N240K, N25G, N61P, N62Q, N62R, N62S, P129Q, P129V, P194L, P239V, P52L, P86S, Q59S, S101N, S125A, S130G, S132N, S161P, S24G, S53G, S78N, S87G, S89Y, T55P, V203Y, V227T, V68A, W106F, Y104N, Y167A, and Y217Q. A variant according to the eighth aspect may comprise an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:2.
[0203] In a ninth aspect, the invention provides an isolated or non-naturally occurring protease variant selected from the group of:
[0204] (a) a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 95%, 96%, 97%, or 98% identity to SEQ ID NO:2, and comprising at least one set of amino acid substitution(s) relative to SEQ ID NO:2 selected from groups (i) through (vii):
[0205] (i) Q059V-S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q, S078N-G097A-I108V-G128A-V147Q/-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-Y217Q-N252Q, and S078N-S087E-G097A-M124I-G128A-Y217Q-S224A;
[0206] (ii) Q059V-S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q, S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-Y217Q-N252Q, and S078N-S087E-G097A-M124I-G128A-Y217Q-S224A;
[0207] (iii) G097A-M124V-Y167A-Y217Q, V068A-Y167A-Y217Q, G097A-I111V-M124V-Y167A, I111V-M124V-Y167A-Y217Q, V068A-I111V-Y167A-Y217Q, G097A-I111V-M124V-Y167A-Y217Q, and P052L-V068A-I111V;
[0208] (iv) G097A-N123A-Y217Q, G097A-N123V-Y217Q, N061P-G102A-G128S-Y217Q, N061P-S101N-G102A-G128S-Y217Q, Y217Q, S078N-G097A-I111V-N123Q-Y217Q, and G102A-N123Q-Y217Q;
[0209] (v) G097A-N123A-Y217Q, G097A-N123V-Y217Q, N061P-N062Q-G097A-G100D-Y217Q, N061P-S101N-G102A-G128S-Y217Q, Y217Q, N061P-G102A-G128S-Y217Q, S078N-G097A-I111V-N123Q-Y217Q, and G102A-N123Q-Y217Q;
[0210] (vi) N061P-N062Q-G097A-G100N-Y217Q, N061P-G097A-M124I-Y217Q, G102A-N123Q-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-N123Q-Y217Q, S053G-N061P-G097A-S101N-N123Q-Y217Q, N061P-G097A-S101N-N123Q-Y217Q, N061P-N062Q-G097A-G100Q-P210S-Y217Q, S053G-N061P-G102A-P129S-P210S-Y217Q, G097A-I111V-M124V-P210S-Y217Q, G097A-N123Q-P210S-Y217Q, N061P-G097A-N123Q-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-M124I-Y217Q, S053G-N061P-G097A-S101N-M124I-Y217Q, S053G-N061P-G097A-M124I-Y217Q, N061P-S078N-G097A-I111V-M124I-Y217Q, N061P-G097A-M124V-Y217Q, N061P-N062Q-G100N-G102A-Y217Q, N061P-N062Q-G097A-G100Q-S101N-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, S078N-G097A-I111V-N123Q-Y217Q, S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, N061P-N062Q-G097A-G100N-S101N-Y217Q, N061P-N062Q-S078N-G097A-G100N-I111V-Y217Q, N061P-N062Q-G097A-G100D-Y217Q, N061P-N062Q-G097A-G100Q-Y217Q, N061P-S101N-G102A-G128S-Y217Q, N061P-G102A-G128S-Y217Q, S024G-N025G-S053G-N061P-S101N-G102A-P129S-Y217Q, S053G-N061P-S101N-G102A-P129S-Y217Q, N061P-S078N-G102A-I111V-P129S-Y217Q, G097A-N123V-Y217Q, S053G-N061P-G102A-P129S-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, N061P-N062Q-G097A-S101N-I111V-Y217Q, N061P-N062Q-G097A-I111V-Y217Q, N062Q-S078N-G097A-I111V-Y217Q, N062Q-G097A-I111V-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, N061P-G097A-S101N-I111V-M124V-Y217Q, G097A-N123Q-Y217Q, N061P-G097A-I111V-M124V-Y217Q, S078N-G097A-I111V-M124V-Y217Q, G097A-I111V-M124I-Y217Q, S053G-S078N-G097A-I111V-G128S-Y217Q, S078N-G097A-G128S-Y217Q, S053G-N061P-G097A-G128S-Y217Q, G097A-N123A-Y217Q. and Y217Q; and [0211] (vii) S101N-G128A-Y217Q, S024G-T055P-N061P-G097A-S101N-G128A, S024G-N025G-N061P-S078N-G097A-S101N-G128A, S024G-T055P-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S053G-N061P-G097A-S101N-G128A-Y217Q-S249N, N025G-S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-G097A-S101N-G128A-Y217Q, S024G-S053G-N061P-G097A-G128A-Y217Q, S024G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-T055P-N061P-G097A-G128A-Y217Q, S024G-S038G-S053G-S078N-S101N-G128A-Y217Q, S053G-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-G128A-Y217Q, S024G-N025G-S053G-N061P-G097A-G128A-S130G-Y217Q, and S024G-S053G-S078N-G097A-S101N-G128A-Y217Q; and
[0212] (b) a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% identity to the amino acid sequence of SEQ ID NO:6 and comprising at least one set of amino acid substitution(s) relative to SEQ ID NO:6 selected from groups (i) through (xii):
[0213] (i) A001C, A001D, A001E, A001F, A001H, A001K, A001L, A001M, A001N, A001P, A001Q, A001R, A001S, A001T, A001V, A013C, A013S, A015C, A015D, A015E, A015L, A015R, A048C, A048E, A048S, A073N, A073T, A074G, A074S, A085C, A085G, A085S, A085T, A085V, A088M, A088S, A092S, A098G, A114G, A133E, A133P, A133R, A137E, A137H, A137R, A142C, A144D, A144G, A144H, A144K, A144L, A144N, A144R, A152S, A153G, A153S, A153V, A176C, A179G, A187C, A187F, A187G, A187L, A187N, A187P, A187Q, A187S, A187T, A187V, A187W, A200G, A216C, A216H, A216R, A216W, A223S, A228S, A230G, A230S, A230T, A230V, A231V, A232C, A232V, A272E, A272G, A272K, A272P, A272R, A273D, A273G, A273H, A273L, A273P, A273Q, A273R, A273T, A273V, A274H, A274M, A274R, D036E, D036N, D036Q, D036S, D041C, D060G, D099A, D099H, D099Q, D120E, D181A, D181G, D181H, D181T, D197T, D259A, D259G, D259H, D259N, D259P, D259Q, D259S, D259T, E156A, E156C, E156G, E156H, E156L, E156Q, E156S, E156V, E195G, E251C, E251I, E251L, E251Q, E251T, F058Y, F189A, F189G, F189H, F189L, F189R, F189S, F189T, F189W, F189Y, F261C, F261D, F261E, F261K, F261P, G020C, G020E, G020F, G020H, G020L, G020N, G020Q, G020R, G020T, G020Y, G024A, G024D, G024P, G046D, G053A, G053D, G053E, G053F, G053L, G053M, G053Q, G053R, G053S, G053Y, G097K, G097M, G097R, G131C, G131D, G131R, G146A, G157A, G157N, G157P, G157S, G157T, G160A, G160L, G160R, G160V, G166C, G166I, G166L, G166Q, G166V, G166W, G169A, G178A, G211E, G211K, G215C, G215D, G215E, G215H, G215L, G215S, G215T, G215W, G258A, G258D, G258P, G258S, H017I, H039S, H226L, H238N, H238Y, I011L, I011T, I011V, I031C, I031F, I031L, I079E, I079F, I079K, I079L, I079M, I079Q, I079R, I107L, I111M, I175L, I205A, I205C, I205V, I268L, I268M, K012A, K012C, K012E, K012F, K012G, K012H, K012L, K012N, K012S, K012T, K012W, K027A, K027N, K027S, K027T, K043A, K043C, K043D, K043E, K043F, K043G, K043H, K043I, K043L, K043M, K043N, K043P, K043Q, K043R, K043T, K043V, K043W, K043Y, K136G, K136H, K141A, K141G, K141H, K141L, K141M, K141N, K141Q, K141R, K141T, K141V, K141W, K170A, K170C, K170G, K170Q, K170S, K213A, K213G, K213H, K213I, K213L, K213N, K213Q, K213R, K213S, K213T, K213V, K237A, K237H, K237I, K237L, K237N, K237S, K237T, K237V, K256A, K256C, K256D, K256E, K256G, K256H, K256M, K256P, K256Q, K256S, K256T, K256V, K256W, K265G, K265H, K265N, K265S, K265Y, L016E, L042C, L042F, L042M, L042V, L075G, L075H, L075I, L075T, L082A, L082E, L082F, L082H, L082R, L082S, L082T, L090M, L135F, L196M, L209A, L209C, L209E, L209G, L209H, L209R, L209S, L233E, L233G, L233S, L235M, L235R, L235V, L235W, L257C, L257D, L257E, L257G, L257P, L257R, L257W, L267E, L267F, M050L, M050Y, M119C, M119I, M124L, M222A, M222F, M222L, M222S, M222T, N025C, N025E, N025P, N056D, N056S, N061A, N061C, N061D, N061G, N061I, N061K, N061L, N061Q, N061R, N062A, N062C, N062H, N062L, N062Q, N062R, N062S, N062T, N062V, N062Y, N076A, N076D, N076E, N076L, N076M, N076P, N076Q, N076S, N076T, N076V, N078D, N078E, N078F, N078G, N078H, N078K, N078P, N078Q, N078R, N101D, N101F, N101R, N117G, N117R, N117S, N118A, N118D, N118H, N118Q, N118R, N118S, N118T, N184C, N184E, N184P, N184R, N212C, N212D, N212E, N212R, N212W, N218C, N218D, N218E, N218F, N218G, N218M, N218P, N218R, N218T, N218V, N218W, N240A, N240G, N240Q, N240S, N240W, N243C, N243G, N243S, N252D, N252E, N252V, N269C, N269H, P005A, P005D, P005M, P005Q, P005V, P005W, P014A, P014D, P014F, P014K, P014M, P014R, P014V, P040F, P040R, P040W, P057A, P057W, P129E, P129R, P129V, P172E, P172K, P172R, P194E, P194H, P194R, P194W, P201A, P201G, P201T, P210E, P210L, P239A, P239G, P239H, P239K, P239N, P239R, P239T, Q002D, Q002E, Q002G, Q002I, Q002K, Q002L, Q002P, Q002R, Q002V, Q010D, Q010R, Q010W, Q019C, Q019D, Q019E, Q019H, Q019L, Q019P, Q019R, Q059A, Q059C, Q059D, Q059E, Q059L, Q059R, Q059S, Q059T, Q059W, Q103W, Q185D, Q185E, Q185K, Q185R, Q185W, Q206D, Q206G, Q206H, Q206L, Q206V, Q206W, Q217A, Q217C, Q217E, Q217F, Q217G, Q217H, Q217K, Q217L, Q217R, Q217V, Q245A, Q245D, Q245E, Q245H, Q245M, Q245R, Q271A, Q271C, Q271D, Q271E, Q271F, Q271G, Q271L, Q271P, Q271R, Q271T, Q271W, Q271Y, Q275A, Q275F, Q275G, Q275I, Q275L, Q275P, Q275R, R186A, R186H, R186I, R186L, R186M, R186V, R186W, S003D, S003E, S003F, S003K, S003R, S009C, S009E, S009K, S018C, S018D, S018R, S037A, S037D, S037E, S037G, S037H, S037K, S037L, S037P, S037R, S037Y, S038D, S038M, S038P, S038R, S038Y, S049C, S049T, S063A, S063C, S063D, S063F, S063L, S063M, S063R, S063Y, S087C, S087D, S087K, S087L, S087M, S087N, S087R, S087Y, S089A, S089C, S089D, S089E, S089F, S089G, S089H, S089I, S089K, S089P, S089R, S089V, S089Y, S105T, S125A, S130C, S130D, S130E, S130K, S130R, S130W, S145D, S145G, S145L, S145R, S145T, S159C, S159D, S159L, S159P, S159W, S161C, S161E, S161R, S162C, S162E, S162W, S163A, S173T, S173V, S182C, S182E, S182R, S183C, S183D, S183P, S183R, S188C, S188D, S188E, S188F, S188K, S188L, S188P, S188R, S188W, S190A, S190C, S190G, S190T, S191G, S204E, S204G, S204R, S204Y, S207G, S224G, S224T, S236C, S236D, S236E, S236G, S248C, S248D, S248E, S248H, S248R, S249E, S249L, S249R, S260A, S260C, S260E, S260G, S260K, S260Q, S260R, S260V, S260Y, T022L, T022P, T055C, T055D, T055E, T055I, T055K, T055M, T055R, T055S, T055V, T055W, T055Y, T071A, T158A, T158D, T158E, T158G, T158L, T158P, T158Q, T158R, T158V, T158Y, T164A, T164G, T164K, T164Q, T164R, T208S, T220A, T242D, T242G, T244D, T244E, T244R, T253E, T253R, T253Y, T254G, T255C, T255D, T255E, T255K, T255R, V004D, V004E, V004T, V026A, V028I, V028L, V030I, V044A, V044C, V044P, V044T, V045C, V045D, V045E, V045G, V045I, V045N, V045R, V045T, V051H, V072L, V081A, V081G, V081H, V081R, V081S, V084I, V084M, V095A, V095C, V143A, V143C, V143E, V143F, V143G, V143H, V143Q, V143T, V143W, V147A, V147Q, V147S, V148I, V148L, V149C, V149I, V149L, V165C, V165L, V180A, V180C, V180M, V180S, V192C, V192F, V192G, V192I, V192Q, V192Y, V203A, V203C, V203D, V203E, V203G, V203K, V203M, V203R, V203S, V270C, V270G, V270L, V270P, W241F, W241L, Y006A, Y006C, Y006D, Y006E, Y006M, Y006N, Y006R, Y006S, Y021C, Y091W, Y104T, Y104V, Y104W, Y214H, Y214Q, Y262C, Y262D, Y262E, Y262H, Y262I, Y262R, and Y262V;
[0214] (ii) A001C, A001E, A001P, A015C, A015E, A048C, A048E, A073T, A085C, A085G, A088I, A088M, A114G, A128H, A137R, A142C, A187C, A187F, A187L, A187N, A187P, A187W, A216C, A216H, A230G, A230S, A273D, A273H, A273P, A273Q, A273R, A273T, A274H, D036N, D036Q, D036S, D041C, D041N, D099A, D099H, D099N, D099Q, D099S, D197T, D259H, E156C, E156G, E156H, E156L, E156Q, F058G, F189A, F189G, F189H, F189L, F189R, F189S, F189T, F261C, F261D, F261E, G046D, G053E, G053L, G053M, G053Q, G053R, G131C, G131D, G146A, G157N, G157P, G157T, G160V, G178A, G215C, G215L, G258A, G258P, H039A, H039S, H067T, H238Y, I011L, I031F, I079E, I111M, I175L, I268M, K012A, K012C, K012E, K012F, K012G, K012H, K012L, K012N, K012W, K027N, K027S, K027T, K043C, K043D, K043G, K043L, K043R, K043W, K136E, K136G, K141G, K141L, K141M, K141N, K141R, K170C, K170Q, K213V, K256D, K265G, K265N, K265Q, K265S, K265Y, L042C, L042F, L075G, L075V, L082A, L082E, L082F, L082H, L082R, L082S, L090M, L090T, L126W, L233E, L233G, L233S, L235V, L257D, L257E, L257G, L257P, L257R, L257W, M050L, M222A, M222F, M222L, M222S, M222T, N025R, N056S, N062C, N062H, N062L, N062Q, N062R, N062T, N062Y, N076D, N076P, N078D, N078E, N078F, N078R, N078V, N117G, N118L, N184P, N269C, N269H, N269Q, P005V, P005W, P014K, P057A, P057W, P086F, P129V, P201T, P225G, P225S, P239A, P239G, P239H, P239N, P239T, Q002D, Q002I, Q002K, Q002L, Q002P, Q002R, Q002V, Q010W, Q019L, Q019P, Q059C, Q059D, Q059E, Q059W, Q185W, Q217C, Q245A, Q245H, Q271C, Q271D, Q271E, Q271L, Q271W, Q275A, Q275G, R186M, S009C, S009L, S018C, S018D, S037E, S037H, S037K, S037L, S037P, S038P, S038Y, S049C, S049N, S063C, S063D, S063F, S063L, S063Y, S087C, S087K, S087L, S087N, S087R, S087Y, S089D, S089E, S089F, S089G, S089P, S089W, S105T, S125A, S130C, S145D, S159D, S159P, S163A, S173V, S182P, S183P, S190A, S190G, S191G, S204E, S224G, S248E, S248H, S249E, S260V, S260Y, T022P, T055E, T055M, T055R, T055W, T158D, T158E, T164A, T164G, T164K, T164Q, T164R, T220A, T242G, T242P, T253E, T255C, T255G, V004D, V004T, V044A, V044L, V044P, V044T, V045C, V045G, V045I, V045K, V045L, V045N, V045R, V045T, V045V, V051H, V081A, V081G, V081H, V081R, V084S, V143G, V147A, V148L, V165C, V180S, V203D, V203G, V203S, V270C, V270G, V270P, V270S, W241L, Y104T, Y214H, Y214Q, Y262D, Y262E, Y262G, Y262H, Y262L, Y262N, Y263G, and Y263W;
[0215] (iii) A001E, A001E-A088T, A001E-A116T, A001E-A128S, A001E-A128S-G131H-N243V, A001E-G024E, A001E-G131H, A001E-G131H-G169A-N243V, A001E-G169A, A001E-K043Y, A001E-K256R, A001E-L257G, A001E-N076D, A001E-N076D-N109G-A128S, A001E-N109G, A001E-N218S, A001E-N243V, A001E-Q103H, A001E-Q206D, A001E-S033T, A001E-S033T-N109G-N218S, A001E-S033T-N109G-N243V, A001E-S063G, A001E-S162G, A001E-S248N, A001E-S249A, A001E-T158S, A088T-A128S, A088T-G169A, A088T-N218S, A088T-Q206D, A116T-G169A, A116T-N218S, A116T-Q206D, A128S, A128S-G131H, A128S-G169A, A128S-N218S, A128S-N243V, A128S-Q206D, G024E, G024E-A088T, G024E-A128S, G024E-G131H, G024E-K043Y, G024E-N076D, G024E-N218S, G024E-Q103H, G024E-Q206D, G024E-S033T, G024E-S063G, G024E-S162G, G024E-S248N, G024E-S249A, G131H-G169A, G131H-N218S, G131H-N243V, G131H-Q206D, G169A, G169A-K256R, G169A-L257G, G169A-N218S, G169A-N243V, G169A-Q206D, G169A-S248N, G169A-S249A, K043Y, K043Y-A116T, K043Y-A128S, K043Y-G131H, K043Y-G169A, K043Y-L257G, K043Y-N076D, K043Y-N109G, K043Y-N218S, K043Y-Q103H, K043Y-Q206D, K043Y-S063G, K043Y-S162G, K043Y-S248N, K043Y-S249A, K043Y-T158S, N076D, N076D-A088T, N076D-A116T, N076D-A128S, N076D-G131H, N076D-G169A, N076D-N218S, N076D-N243V, N076D-Q103H, N076D-Q206D, N076D-S162G, N076D-S248N, N076D-S249A, N109G-G169A, N109G-Q206D, N109G-S248N, N218S, N218S-K256R, N218S-L257G, N218S-S248N, N218S-S249A, P040E-N109G-A128S-G131H, Q103H, Q103H-G169A, Q103H-Q206D, Q206D, Q206D-K256R, Q206D-L257G, Q206D-N218S, Q206D-N243V, Q206D-S248N, Q206D-S249A, S018T-Y021N-S033T-N109G-A128S-N243V-S248N-K256R, S033T, S033T-A088T, S033T-A116T, S033T-A128S, S033T-A128S-G131H-N243P, S033T-G131H, S033T-G169A, S033T-K043Y, S033T-K256R, S033T-L257G, S033T-N076D, S033T-N076D-A128S-N218S, S033T-N076D-N109G-A128S-N218S-N243V-S248N-K256R, S033T-N109G, S033T-N109G-A128S-N243V-S248N-K256R, S033T-N218S, S033T-P040E-Q103H-N109G, S033T-Q103H-A128S-G131H, S033T-Q206D, S033T-S063G, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243V, S033T-S162G, S033T-S248N, S033T-S249A, S063G-A116T, S063G-G131H, S063G-G169A, S063G-N109G-A128S-G131H, S063G-N218S, S063G-N243V, S063G-Q206D, S063G-S249A, S162G-G169A, S162G-N218S, S162G-Q206D, S162G-S249A, S248N-K256R, S248N-S249A, S249A-K256R, S249A-L257G, T158S-G169A, T158S-K256R, T158S-Q206D, and T158S-S162G;
[0216] (iv) A001E, A001E-A088T, A001E-A116T, A001E-A128S-G131H-N243V, A001E-G024E, A001E-G131H-G169A-N243V, A001E-G169A, A001E-K043Y, A001E-L257G, A001E-N076D, A001E-N076D-N109G-A128S, A001E-N109G, A001E-Q103H, A001E-Q206D, A001E-S033T-N109G-N218S, A001E-S033T-N109G-N243V, A001E-S248N, A001E-T158S, A088T-G169A, A088T-Q206D, A116T-G169A, A116T-N218S, A116T-Q206D, A128S-G131H, A128S-N243V-S248N-K256R, A128S-Q206D, G024E-K043Y, G024E-N076D, G024E-Q206D, G131H-G169A, G131H-N218S, G131H-N243V-K256R, G131H-Q206D, G131H-S248N, G169A-K256R, G169A-N218S, G169A-N243V, G169A-Q206D, G169A-S249A, K043Y-G169A, K043Y-N076D, K043Y-N218S, K043Y-Q206D, N061P-N109G-G131H-N243V, N061P-N109G-N243V, N061S-N109G-A128S-N243V-S248N-K256R-S260P, N061S-N109G-N243V, N076D, N076D-G131H, N076D-L257G, N076D-N109G, N076D-Q103H, N076D-Q206D, N076D-S249A, N109G-A128S-N243V-S248N, N109G-A128S-N243V-S248N-K256R, N109G-A128S-S248N-K256R, N109G-G169A, N109G-N243P-S248A-K256R, N109G-N243P-S248N-K256R, N109G-N243V-K256R, N109G-N243V-S248A-K256R, N109G-N243V-S248N, N109G-N243V-S248N-K256R, N109G-S248N-K256R, N218S-S249A, N243V-S248N-K256R, P040E-N109G-A128S-G131H, Q103H-Q206D, Q206D, Q206D-K256R, Q206D-L257G, Q206D-N218S, Q206D-S248N, Q206D-S249A, S009T-N109G-A128S-K141R-N243V-S248N-K256R, S009T-S018T-Y021N-A128S-K141R-N243V, S018T-Y021N-A128S-N243V, S018T-Y021N-N061S-A128S-N243V-S260P, S018T-Y021N-N061S-N109G-A128S-S260P, S018T-Y021N-S033T-N109G-A128S-N243V-S248N-K256R, S033T, S033T-A128S-G131H-N243P, S033T-A128S-G131H-N243V, S033T-G169A, S033T-N076D-A128S-N218S, S033T-N076D-N109G-A128S-N218S-N243V-S248N-K256R, S033T-N109G-A128S-N243P-S248N-K256R, S033T-N109G-A128S-N243V-S248N-K256R, S033T-P040E-Q103H-N109G, S033T-Q103H-A128S-G131H, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243V, S063G-G131H, S063G-G169A, S063G-N109G-A128S-G131H, S063G-Q206D, S162G-Q206D, T158S-Q206D, and T158S-S162G;
[0217] (v) A001E-A088T, A001E-A128S, A001E-A128S-G131H-N243V, A001E-G024E, A001E-G024E-S204E-Q206D, A001E-G131H-G169A-N243V, A001E-K043Y, A001E-N076D, A001E-N076D-N109G-A128S, A001E-N218S, A001E-N243V, A001E-Q103H, A001E-Q206D, A001E-S033T, A001E-S033T-N109G-N218S, A001E-S162G, A116T-Q206D, A128S-N243V-S248N-K256R, A128S-Q206D, G024E-Q206D, G131H-Q206D, K043Y-G131H, K043Y-Q103H, K043Y-Q206D, K043Y-S162G, N061S-N109G-A128S-N243V-S260P, N076D-A128S, N076D-Q206D, N076D-S162G, N076D-S248N, N109G-A128S-S248N-K256R, N109G-A128S-T158S-N243V-S248N-K256R, N109G-N243P-S248N-K256R, N109G-Q206D, N243V-S248N-K256R, P040E-N109G-A128S-G131H, Q103H-Q206D, Q206D-K256R, Q206D-N243V, Q206D-S248N, Q206D-S249A, S018T-Y021N-N061S-A128S-N243V-S260P, S018T-Y021N-N061S-N109G-A128S-S260P, S033T-A128S-G131H-N243P, S033T-N076D-A128S-N218S, S033T-N076D-N109G-A128S-N218S-N243V-S248N-K256R, S033T-P040E-Q103H-N109G, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S063G-Q206D, S162G-Q206D, and T158S-Q206D;
[0218] (vi) A001E, A001E-A128S, A001E-G024E, A001E-G131H, A001E-G169A, A001E-K043Y, A001E-K256R, A001E-L257G, A001E-N076D, A001E-N109G, A001E-N218S, A001E-N243V, A001E-Q103H, A001E-Q206D, A001E-S033T, A001E-S063G, A001E-S162G, A001E-S248N, A001E-S249A, A001E-T158S, A088T-A116T, A088T-G131H, A088T-N109G, A088T-N218S, A088T-Q206D, A116T-A128S, A116T-G131H, A116T-Q206D, A116T-S248N, A128S, A128S-G131H, A128S-Q206D, G024E, G024E-N076D, G024E-N109G, G024E-Q206D, G131H, G131H-L257G, G131H-Q206D, G169A-K256R, G169A-Q206D, K043Y, K043Y-A088T, K043Y-A116T, K043Y-A128S, K043Y-G131H, K043Y-G169A, K043Y-K256R, K043Y-L257G, K043Y-N076D, K043Y-N109G, K043Y-N218S, K043Y-N243V, K043Y-Q103H, K043Y-S063G, K043Y-S162G, K043Y-S248N, K043Y-S249A, K043Y-T158S, N076D-A088T, N076D-A116T, N076D-A128S, N076D-G131H, N076D-G169A, N076D-N109G, N076D-N218S, N076D-N243V, N076D-Q103H, N076D-Q206D, N076D-S248N, N076D-T158S, N109G, N109G-G169A, N109G-Q206D, N109G-S162G, N109G-T158S, N218S-S248N, Q103H, Q103H-A128S, Q103H-G131H, Q103H-Q206D, Q103H-S248N, Q103H-S249A, Q103H-T158S, Q206D, Q206D-K256R, Q206D-L257G, Q206D-N218S, Q206D-N243V, Q206D-S248N, Q206D-S249A, S033T, S033T-K256R, S033T-S063G, S033T-S249A, S063G-A088T, S063G-G131H, S063G-G169A, S063G-N076D, S063G-N109G, S063G-N218S, S063G-Q103H, S063G-Q206D, S162G-Q206D, S162G-S249A, S248N-S249A, S249A-K256R, T158S-Q206D, and T158S-S249A;
[0219] (vii) G131H-S162G, G131H-S248N, G131H-T158S, K043Y-G131H, K043Y-S162G, Q103H-A128S, Q103H-N218S, S033T-Q103H, S063G-A088T, S063G-G131H, S063G-S248N, and T158S-S162G;
[0220] (viii) A015S-A088T-N109G-G131H-T158S-N218S-S248N, A088T-A098S-G131H-S248N-K256R-L257G, A088T-A098S-N218S-K256R, A088T-A116T-G131H-G146C, A088T-A116T-G131H-K256R, A088T-A116T-G131H-K256R-L257G-L267M, A088T-A116T-G131H-N218S-N243V-K256R, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-N218S-N243V-S248N, A088T-A116T-G131H-N218S-S248N, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-N243V, A088T-A116T-G131H-S248N, A088T-A116T-G131H-S248N-L257G, A088T-A116T-G131H-S248N-L257G, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S, A088T-A116T-G131H-T158S-N218S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-A116T-G131H-T158S-N218S-S248N, A088T-A116T-G131H-T158S-N218S-S248N-K256R, A088T-A116T-K256R-L257G, A088T-A116T-N218S, A088T-A116T-N218S-I268V, A088T-A116T-N218S-K256R, A088T-A116T-N218S-L257G, A088T-A116T-N218S-N243V-Q271R, A088T-A116T-N218S-N243V-S248N-K256R, A088T-A116T-N218S-N243V-S248N-K256R-Q275R, A088T-A116T-N218S-S248N, A088T-A116T-N218S-S248N-K256R, A088T-A116T-N243V-S248N-K256R, A088T-A116T-T158S, A088T-A116T-T158S-K256R, A088T-A116T-T158S-K256R-L257G, A088T-A116T-T158S-N218S-K256R, A088T-A116T-T158S-N218S-N243V-L257G, A088T-A116T-T158S-N218S-N243V-S248N-E251K-K256R-L257G, A088T-A116T-T158S-N218S-N243V-S248N-K256R, A088T-A116T-T158S-N218S-N243V-S248N-L257G, A088T-A116T-T158S-N218S-S248N, A088T-A116T-T158S-N218S-S248N-K256R, A088T-A116T-T158S-N218S-S248N-K256R-L257G, A088T-A116T-T158S-N218S-S248N-L257G, A088T-A116T-T158S-N243V, A088T-A116T-T158S-N243V-K256R-L257G, A088T-A116T-T158S-S248N-K256R-L257G, A088T-A116T-T158S-S248N-L257G, A088T-A138E-N218S-N243V-K256R, A088T-G131H, A088T-G131H, A088T-G131H-K141E-N218S-N243V-S248N-L257G, A088T-G131H-K256R, A088T-G131H-N218S-K237R-K256R-L257G, A088T-G131H-N218S-K256R, A088T-G131H-N218S-K256R, A088T-G131H-N218S-K256R-L257G, A088T-G131H-N218S-N243V-K256R-L257G, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-N243V-S248N, A088T-G131H-N218S-N243V-S248N-K256R, A088T-G131H-N218S-N243V-S248N-K256R, A088T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-G131H-N218S-N243V-S248N-L257G, A088T-G131H-N218S-S248N, A088T-G131H-N218S-S248N-K256R, A088T-G131H-N218S-S248N-K256R, A088T-G131H-N243V, A088T-G131H-N243V-K256R, A088T-G131H-N243V-K256R-L257G, A088T-G131H-N243V-S248N, A088T-G131H-N243V-S248N, A088T-G131H-N243V-S248N-K256R, A088T-G131H-S248N, A088T-G131H-S248N-K256R, A088T-G131H-T158S-N218S-I234T-S248N-L257G, A088T-G131H-T158S-N218S-K256R-L257G, A088T-G131H-T158S-N218S-L257G, A088T-G131H-T158S-N218S-N243V, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-G131H-T158S-N218S-S248N-K256R, A088T-G131H-T158S-N218S-S248N-K256R-L257G, A088T-G131H-T158S-S248N-K256R, A088T-G131H-T158S-S248N-K256R-L257G, A088T-G131H-T158S-S248N-K256R-L257G, A088T-G131H-V149L-T158S-N243V-S248N-K256R-L257G, A088T-I107T-N109G-G131H-N218S-A223G-S248N-K256R, A088T-I107T-N109G-G131H-N218S-S248N-K256R, A088T-I108T-N109G-G131H-T158S-N218S-S248N-K256R-L257G, A088T-K213N-N243V-S248N-K256R, A088T-K256R-L257G, A088T-L257G, A088T-N109G-A116T-G131H-A232S-N243V-K256R, A088T-N109G-A116T-G131H-D140G-S248N-L257G, A088T-N109G-A116T-G131H-D140G-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-K141E-N218S, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V-K256R, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-N269D, A088T-N109G-A116T-G131H-N218S-N243V-S248N-Q275R, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N, A088T-N109G-A116T-G131H-T158S, A088T-N109G-A116T-G131H-T158S-N218S-L257G-I268V, A088T-N109G-A116T-G131H-T158S-N218S-N243F-S248N, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R-I268V, A088T-N109G-A116T-G131H-T158S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-T158S-S248N, A088T-N109G-A116T-G131H-T158S-S248N, A088T-N109G-A116T-G131H-V149A-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-W241L-S248N-K256R-L257G, A088T-N109G-A116T-K256R, A088T-N109G-A116T-M124I-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-N218S-K256R-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R, A088T-N109G-A116T-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N218S-S248N, A088T-N109G-A116T-T158S-K256R-L257G, A088T-N109G-A116T-T158S-N218S, A088T-N109G-A116T-T158S-N218S-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-T158S-N243V-S248N, A088T-N109G-A116T-T158S-N243V-S248N-K256R, A088T-N109G-A116T-V148A-N218S-N243V, A088T-N109G-D140G-N243V, A088T-N109G-G131H-A138V-T158S-N218S-N243V-S248N-L257G, A088T-N109G-G131H-D140G-T158S-N243V-S248N-K256R, A088T-N109G-G131H-K141E-T158S-N218S-K256R, A088T-N109G-G131H-K256R-L257G, A088T-N109G-G131H-N218S-N243V, A088T-N109G-G131H-N218S-N243V-S248N, A088T-N109G-G131H-N218S-N243V-S248N-L257G, A088T-N109G-G131H-N218S-S248N, A088T-N109G-G131H-N218S-S248N-K256R-L257G, A088T-N109G-G131H-N218S-S248N-K256R-L257G-Q275R, A088T-N109G-G131H-N218S-S248N-K256R-Q271R, A088T-N109G-G131H-N218S-S248N-L257G, A088T-N109G-G131H-N243V-S248N, A088T-N109G-G131H-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S, A088T-N109G-G131H-T158S-K256R, A088T-N109G-G131H-T158S-L233S-N243V-S248N, A088T-N109G-G131H-T158S-N218S-N243V-K256R, A088T-N109G-G131H-T158S-N218S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-L257G, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-S248N-K256R, A088T-N109G-G131H-T158S-S248N-K256R-L257G, A088T-N109G-G131H-V149A-K256R-L257G, A088T-N109G-G131H-V149L-T158S-K256R-L257G, A088T-N109G-N218S-K256R-L257G, A088T-N109G-N218S-N243V-K256R, A088T-N109G-N218S-N243V-L257G, A088T-N109G-N218S-N243V-S248N, A088T-N109G-N218S-N243V-S248N-K256R-L257G, A088T-N109G-N218S-S248N-K256R, A088T-N109G-N218S-S248N-T255K-K256R-L257G, A088T-N109G-N243V-K256R, A088T-N109G-N243V-S248N-K256R, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S, A088T-N109G-T158S-K256R-L257G, A088T-N109G-T158S-N218S, A088T-N109G-T158S-N218S-K256R-L257G-Q271K, A088T-N109G-T158S-N218S-L257G, A088T-N109G-T158S-N218S-N243V-K256R, A088T-N109G-T158S-N218S-N243V-K256R-L257G, A088T-N109G-T158S-N218S-N243V-S248N-K256R, A088T-N109G-T158S-N218S-N243V-S248N-L257G, A088T-N109G-T158S-N218S-S248N, A088T-N109G-T158S-S248N, A088T-N109G-T158S-S248N, A088T-N109G-T158S-S248N-K256R, A088T-N109G-W241R-S248N-K256R, A088T-N218S-N243V-K256R, A088T-N218S-N243V-L257G, A088T-N218S-N243V-S248N, A088T-N218S-N243V-S248N-K256R, A088T-N218S-N243V-S248N-K256R, A088T-N218S-S248N, A088T-N218S-S248N-L257G, A088T-N218S-S248N-L257G-Q271R, A088T-S248N-K256R-L257G, A088T-T158S-K256R, A088T-T158S-N218S-K256R-L257G, A088T-T158S-N218S-L257G, A088T-T158S-N218S-N243V-K256R, A088T-T158S-N218S-N243V-K256R, A088T-T158S-N218S-N243V-L257G, A088T-T158S-N218S-N243V-S248N, A088T-T158S-N218S-N243V-S248N-K256R, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N218S-N243V-S248N-L257G, A088T-T158S-N218S-Q245K-S248N-K256R, A088T-T158S-N218S-S248N, A088T-T158S-N218S-S248N-K256R, A088T-T158S-N218S-S248N-K256R, A088T-T158S-N218S-S248N-L257G, A088T-T158S-N218S-S248N-L257G-Q275K, A088T-T158S-N243V, A088T-T158S-N243V-K256R, A088T-T158S-N243V-S248N, A088T-T158S-S248N-K256R-L257G, A088T-T158S-S248N-L257G, A088T-T158S-V203I-N218S-K256R-L257G, A088T-V147I-N218S-N243V-K256R-L257G, A088T-Y104H-A116T-G131H-N218S-N243V, A116T-D140G-T158S-N218S-N243V-S248N, A116T-G131H-K141E-N218S-N243V-S248N-L257G, A116T-G131H-L257G, A116T-G131H-N218S-N243V, A116T-G131H-N218S-N243V-K256R, A116T-G131H-N218S-N243V-K256R-L257G, A116T-G131H-N218S-N243V-S248N-K256R, A116T-G131H-N218S-N243V-S248N-K256R-L257G, A116T-G131H-N218S-S248N, A116T-G131H-N218S-S248N-K256R, A116T-G131H-N218S-W241R-N243V-S248N-K256R-L257G, A116T-G131H-N243V-S248N-K256R, A116T-G131H-T158S-N218S-L257G, A116T-G131H-T158S-N218S-N243V-S248N, A116T-G131H-T158S-N218S-S248N, A116T-G131H-T158S-N218S-S248N-L257G-N269D, A116T-G131H-T158S-N218S-S248N-Q271R, A116T-G131H-T158S-N243V-K256R, A116T-G131H-T158S-N243V-S248N, A116T-G131H-V139I-N218S-N243V-S248N, A116T-G131H-V150A-T158S-N243V-S248N-K256R-L257G, A116T-G157E-T158S-N243V-S248N-K256R, A116T-K141E-N218S-N243V-S248N-K256R-L257G, A116T-K256R, A116T-N218S-N243V-S248N, A116T-N218T-N243V-S248N, A116T-N243V-K256R-L257G, A116T-S248N-K256R, A116T-T158S-N218S, A116T-T158S-N218S-K256R, A116T-T158S-N218S-K256R-L257G, A116T-T158S-N218S-N243V-L257G, A116T-T158S-N218S-N243V-S248N, A116T-T158S-N218S-S248N-L257G, G024S-G053S-N078S-G097A-N101S, G053S-A088T-N109G-A116T-G131H-T158S-G169S-N218S-S248N-K256R-L257G, G065D-A088T-G131H-N243V-S248N, G131H-K141R-T158S-N218S-K256R, G131H-K256R, G131H-N218S-K256R-L257G, G131H-N218S-L257G, G131H-N218S-N243V-S248N-L257G, G131H-N218S-S248N, G131H-N218S-S248N-K256R, G131H-N243V-S248N, G131H-N243V-S248N-L257G, G131H-T158S-N218S, G131H-T158S-N218S-N240H-N243V-S248N-K256R-L257G, G131H-T158S-N218S-N243V, G131H-T158S-N218S-N243V-S248N-K256R-L257G, G131H-T158S-N218S-N243V-S248N-L257G, G131H-T158S-N218S-S248N-I268V, G131H-T158S-N218S-S248N-K256R-L257G-N269S, G131H-T158S-N243V-L257G, G131H-T158S-N243V-S248N, G131H-T158S-S248N, I107T-G131H-T158S-N243V-S248N-K256R-L257G, I107T-N109G-G131H-N218S-L257G, K256R, K256R-L257G, L090I-N109G-T158S-N243V, L257G, N109G-A116T-G131H-N218S-N243V, N109G-A116T-G131H-N218S-N243V-L257G, N109G-A116T-G131H-N218S-S248N-L257G, N109G-A116T-G131H-N218S-W241R-N243V-K256R, N109G-A116T-G131H-S248N-L257G, N109G-A116T-G131H-T158S-N218S-K256R, N109G-A116T-G131H-T158S-N218S-K256R-L257G-Q271R, N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-S248N, N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, N109G-A116T-G131H-T158S-N243V-K256R-L257G, N109G-A116T-G131H-T158S-N243V-S248N-K256R, N109G-A116T-G131H-T158S-S248N-K256R-L257G, N109G-A116T-I234T-N243V-S248N-K256R-L257G, N109G-A116T-K141E-T158S-N218S-N243V-L257G, N109G-A116T-N218S-N243V-S248N, N109G-A116T-N218S-W241R-N243V-S248N-K256R-L257G, N109G-A116T-N243V-K256R-L257G, N109G-A116T-T158S-N218S-K237R-N243V-S248N, N109G-A116T-T158S-N218S-N243V-S248N, N109G-G131H-K141E-L257G, N109G-G131H-N218S-N243V, N109G-G131H-N218S-N243V-S248N, N109G-G131H-S248N, N109G-G131H-T158S, N109G-G131H-T158S-L257G, N109G-G131H-T158S-N218S-N243V-S248N-K256R, N109G-G131H-T158S-N218S-S248N-K256R, N109G-G131H-T158S-N218S-S248N-L257G, N109G-G131H-T158S-N243V-S248N-K256R-L257G, N109G-G131H-T158S-S248N-K256R, N109G-N218S-K256R-L257G, N109G-N218S-N243V-S248N-K256R, N109G-N218S-S248N-L257G, N109G-S248N, N109G-T158S-N218S, N109G-T158S-N218S-N243V, N109G-T158S-N218S-N243V-L257G, N109G-T158S-N218S-S248N-K256R, N109G-T158S-N243V-L257G, N109G-T158S-N243V-S248N-K256R-L257G, N218S-K256R, N218S-N243V-K256R, N218S-N243V-S248N, N218S-S248N, N218S-S248N-K256R, N218S-S248N-K256R-L257G, N243V-L257G, S003P-N109G-A116T-G131H-N218S-N243V-S248N, S003P-N109G-A116T-G131H-T158S-N218S-K256R, S003P-N109G-G131H-T158S-L257G, S003P-S248N-L257G, S105H-W106G-I107L-I108S-N109A-G110A-I111S-E112N-W113G-A114P, S248N-L257G, T158S, T158S-K256R, T158S-N218S, T158S-N218S-K256R, T158S-N218S-L233S-S248N, T158S-N218S-L257G, T158S-N218S-N243V-K256R, T158S-N218S-N243V-S248N, T158S-N218S-N243V-S248N-L257G, T158S-N218S-S248N-K256R-L257G, T158S-N218S-S248N-L257G, T158S-N243V-S248N-K256R, T158S-N243V-S248N-K256R-L257G, T158S-N243V-S248N-L257G, T158S-S248N, T158S-S248N-K256R-L257G, V004A-A088T-G131H-N218S-N243V-S248N-L257G, V004L-A088T-G131H-T158S-N218S-S248N-L257G, Y006H-N218S-N243V-S248N, and Y104H-N109G-G131H-N243V-S248N;
[0221] (ix) A088T-A098S-G131H-S248N-K256R-L257G, A088T-A116T-G131H-K256R, A088T-A116T-G131H-L257G, A088T-A116T-G131H-N218S-N243V, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-N218S-S248N-L257G, A088T-A116T-G131H-N243V-S248N-L257G, A088T-A116T-G131H-S248N, A088T-A116T-G131H-S248N-K256R-L257G, A088T-A116T-G131H-T158S-K256R, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-N243V-S248N-L257G, A088T-A116T-G131H-V150A-N218S-S248N-L257G, A088T-A116T-K256R-L257G, A088T-A116T-K256R-L257G, A088T-A116T-L257G, A088T-A116T-N218S, A088T-A116T-N218S, A088T-A116T-N218S-N243V-Q271R, A088T-A116T-N218S-N243V-S248N, A088T-A116T-N218S-S248N-K256R, A088T-A116T-T158S, A088T-A116T-T158S, A088T-A116T-T158S-K256R, A088T-A116T-T158S-K256R-L257G, A088T-A116T-T158S-N218S-N243V-L257G, A088T-A116T-T158S-N218S-S248N, A088T-A116T-T158S-N218S-S248N-K256R, A088T-A116T-T158S-N243V-K256R-L257G, A088T-A116T-T158S-N243V-S248N-K256R, A088T-A138E-N218S-N243V-K256R, A088T-G131D-T158S-N243V-S248N, A088T-G131H, A088T-G131H-L257G, A088T-G131H-N218S-K237R-K256R-L257G, A088T-G131H-N218S-K256R, A088T-G131H-N218S-N243V-S248N-K256R, A088T-G131H-N218S-N243V-S248N-L257G, A088T-G131H-N243V-K256R, A088T-G131H-N243V-L257G, A088T-G131H-N243V-S248N, A088T-G131H-N243V-S248N-K256R, A088T-G131H-T158S-N218S-I234T-S248N-L257G, A088T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-N243V-K256R-L257G, A088T-G131H-T158S-S248N, A088T-G131H-T158S-S248N-K256R, A088T-G131H-T158S-S248N-K256R, A088T-G131H-T158S-S248N-K256R-L257G, A088T-G131H-V149L-T158S-N243V-S248N-K256R-L257G, A088T-I108T-N109G-G131H-T158S-N218S-S248N-K256R-L257G, A088T-K213N-N243V-S248N-K256R, A088T-K256R-L257G, A088T-N109G-A116T-G131H-D140G-S248N-L257G, A088T-N109G-A116T-G131H-K141E-N218S, A088T-N109G-A116T-G131H-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-Q275R, A088T-N109G-A116T-G131H-N218S-S248N, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N243V-S248N-K256R, A088T-N109G-A116T-G131H-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-K256R, A088T-N109G-A116T-G131H-T158S-N218S-L257G-I268V, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N, A088T-N109G-A116T-G131H-V149A-T158S-N218S-K256R, A088T-N109G-A116T-G131H-W241L-S248N-K256R-L257G, A088T-N109G-A116T-M124I-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R, A088T-N109G-A116T-N218S-S248N-K256R-L257G, A088T-N109G-A116T-N243V-K256R, A088T-N109G-A116T-N243V-S248N-L257G, A088T-N109G-A116T-S248N-L257G, A088T-N109G-A116T-T158S-K256R-L257G, A088T-N109G-A116T-T158S-N212D-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-A116T-T158S-N218S-S248N, A088T-N109G-A116T-T158S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N243V-S248N, A088T-N109G-A116T-T158S-S248N-L257G-Q271P, A088T-N109G-A116T-V148A-N218S-N243V, A088T-N109G-A137E-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-D140G-N243V, A088T-N109G-G131H-A152S-T158S-N218S-S248N-K256R, A088T-N109G-G131H-D140G-T158S-N243V-S248N-K256R, A088T-N109G-G131H-K256R-L257G, A088T-N109G-G131H-K256R-L257G, A088T-N109G-G131H-N218S, A088T-N109G-G131H-N218S-L257G, A088T-N109G-G131H-N218S-N243V, A088T-N109G-G131H-N218S-N243V-S248N-L257G, A088T-N109G-G131H-N218S-S248N, A088T-N109G-G131H-N243V-S248N, A088T-N109G-G131H-T158S-L233S-N243V-S248N, A088T-N109G-G131H-T158S-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-G131H-T158S-N218S-W241R-S248N-L257G, A088T-N109G-G131H-T158S-N243V-S248N-L257G, A088T-N109G-G131H-T158S-S248N-K256R, A088T-N109G-L257G, A088T-N109G-N218S-K256R-L257G, A088T-N109G-N218S-S248N-K256R, A088T-N109G-N218S-S248N-T255K-K256R-L257G, A088T-N109G-N243V-K256R-L257G, A088T-N109G-N243V-S248N-K256R, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S, A088T-N109G-T158S-N218S-N243V, A088T-N109G-T158S-N218S-N243V-L257G, A088T-N109G-T158S-N218S-N243V-S248N-L257G, A088T-N109G-T158S-N218S-S248N, A088T-N109G-T158S-N218S-S248N, A088T-N109G-T158S-N243V-S248N-K256R, A088T-N109G-T158S-N243V-S248N-Q275R, A088T-N109G-T158S-S248N, A088T-N109G-T158S-S248N-K256R, A088T-N109G-W241R-S248N-K256R, A088T-N218S-N243V-S248N-K256R-L257G, A088T-N218S-S248N-L257G, A088T-N243V-L257G, A088T-S248N, A088T-S248N-K256R-L257G, A088T-S248N-L257G, A088T-S248N-L257G-I268V, A088T-T158S-K256R, A088T-T158S-N218S-K256R, A088T-T158S-N218S-L257G, A088T-T158S-N218S-L257G, A088T-T158S-N218S-N243V-S248N, A088T-T158S-N218S-N243V-S248N-L257G, A088T-T158S-N218S-Q245K-S248N-K256R, A088T-T158S-N218S-S248N-K256R, A088T-T158S-N218S-S248N-L257G, A088T-T158S-N218S-S248N-L257G-Q275K, A088T-T158S-N243V-K256R, A088T-T158S-N243V-K256R-L257G, A088T-T158S-N243V-L257G, A088T-T158S-N243V-S248N-K256R, A088T-T158S-S248N, A088T-T158S-S248N-K256R-L257G, A088T-T158S-S248N-L257G, A088T-T158S-S248N-L257G, A088T-T158S-V203I-N218S-K256R-L257G, A088T-Y104H-A116T-G131H-N218S-N243V, A088T-Y104H-N109G-A116T-A153S-N218S-N243V-S248N-L257G-N269D, A088T-Y104H-N109G-G131H-A137E-T158S-N218S-N243V-S248N-K256R, A098S-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A116T-D140G-T158S-N218S-N243V-S248N, A116T-G131H-K141E-N218S-N243V-S248N-L257G, A116T-G131H-N218S-W241R-N243V-S248N-K256R-L257G, A116T-G131H-N243V, A116T-G131H-T158S-K256R-L257G, A116T-G131H-T158S-N218S-N243V, A116T-G131H-T158S-N218S-S248N-K256R, A116T-G131H-T158S-N243V-S248N, A116T-G131H-V139I-N218S-N243V-S248N, A116T-G157E-T158S-N243V-S248N-K256R, A116T-N218S-S248N, A116T-N218S-S248N-K256R, A116T-N243V-K256R, A116T-S248N-K256R, A116T-T158S, A116T-T158S-L257G-Q271R, A116T-T158S-N218S-N243V-K256R-L257G, G053S-A088T-N109G-A116T-G131H-T158S-G169S-N218S-S248N-K256R-L257G, G065D-A088T-G131H-N243V-S248N, G131H-N218S-L257G, G131H-N218S-S248N, G131H-N243V-K256R, G131H-S248N-K256R, G131H-T158S, G131H-T158S-K256R, G131H-T158S-N243V-L257G, K256R, K256R-L257G, L090I-N109G-T158S-N243V, L257G, N109G-A116T-G131H, N109G-A116T-G131H-N243V, N109G-A116T-G131H-N243V-K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-S248N, N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, N109G-A116T-I234T-N243V-S248N-K256R-L257G, N109G-A116T-N218S-N243V-L257G, N109G-A116T-N218S-W241R-N243V-S248N-K256R-L257G, N109G-A116T-S248N-K256R, N109G-A116T-T158S-K256R-L257G, N109G-A116T-T158S-N218S-K237R-N243V-S248N, N109G-A116T-T158S-N218S-S248N-K256R, N109G-A116T-T158S-N243V, N109G-A116T-T158S-S248N, N109G-A116T-T158S-S248N-K256R, N109G-G131H-A137V-T158S-N218S-S248N, N109G-G131H-K141E-L257G, N109G-G131H-T158S, N109G-G131H-T158S-N243V-K256R-L257G, N109G-G131H-T158S-N243V-S248N-L257G, N109G-G131H-T158S-S248N-Q271R, N109G-N218S-S248N-K256R, N109G-N243V-S248N-L257G, N109G-S248N, N109G-T158S-K256R-L257G, N109G-T158S-N218S-N243V-L257G, N109G-T158S-N243V-K256R-L257G, N109G-T158S-N243V-S248N-K256R-L257G, N218S-N243V-S248N-K256R-L257G, N243V, P014L-A015L-L016C-H017T-S018L-Q019K-G020A-Y021T-T022L-G023E, S003P-N109G-A116T-G131H-T158S-N218S-K256R, S003P-N109G-G131H-T158S-L257G, S003P-S248N-L257G, S105H-W106G-I107L-I108S-N109A-G110A-I111S-E112N-W113G-A114P, T158S-N218S-N243V-K256R, T158S-N218S-N243V-S248N, T158S-N218S-S248N-K256R, T158S-N243V-S248N, T158S-S248N, T158S-S248N-K256R, T158S-S248N-K256R-L257G, V004A-A088T-A116T-T158S-N218S, V004A-A088T-G131H-N218S-N243V-S248N-L257G, V004M-A116T-V148A-T158S-N243V-S248N-K256R, Y006H-N109G-N218S-N243V-S248N, Y104H-A116T-T158S-S248N, Y104H-N109G-G131H-N243V-S248N, and Y104H-N218S-L257G;
[0222] (x) A045S-S236G, A045S-S236Y, A134T-S260G, G024A-S037W, I031V-S038W, 15T-S183T, I115V-N184Y, N025K-P129K, N025K-P129R, N025K-S037P, N061D-S260I, Q010L-S037P, Q010R-S037T, Q019L-S260N, Q019L-S260P, S037P-T254S, S161P-T253A, and S162L-D181H;
[0223] (xi) A045S-S236G, A045S-S236Y, A133V-S260N, A134T-S260G, G024A-S037W, I031V-S038W, I115T-S183T, I115V-N184Y, N025K-P129R, N025K-S037P, N061D-S260I, Q010R-S037T, Q019L-S260N, Q019L-S260P, and S162L-D181H; and
[0224] (xii) A045S-S236G, A045S-S236Y, I115T-S183T, N025K-S037P, N061S-S260P, Q010L-S037P, Q010R-S037T, Q019L-S260N, S037P-T254S, and S161P-S260P; wherein each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2.
[0225] A variant according to the ninth aspect of the invention may have enhanced proteolytic activity and/or enhanced cleaning activity compared to the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6, respectively. A variant according to the ninth aspect of the invention may have a performance index in a proteolytic assay (e.g., AAPF assay) or cleaning assay (e.g., BMI, grass, or egg microswatch assay) that is greater than that of the protease set forth in SEQ ID NO:2, SEQ ID NO:4, or SEQ ID NO:6. A variant according to the ninth aspect of the invention may be a protease variant of BPN' subtilisin protease (SEQ ID NO:2).
[0226] In a twenty-first aspect, the invention provides an isolated or non-naturally occurring cold water protease that is a variant of a parent protease, said cold water protease comprising a total of three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14 or 15 mutations selected from groups (a) and (b) below, wherein at least one mutation is selected from group (a):
(a) 1, 9, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 33, 43, 76, 102, 109, 137, 141, 158, 169, 204, 210, 218, 243, 248, 249, 256, 257, 260, and 269; and (b) 24, 25, 40, 52. 53, 55, 58, 59, 61, 62, 63, 68, 78, 86, 87, 88, 89, 92, 96, 97, 100, 101, 103, 104, 106, 111, 114, 115, 116, 117, 118, 123, 124, 125, 126, 128, 129, 130, 131, 132, 133, 134, 144, 145, 159, 161, 162, 167, 194, 203, 206, 213, 217, 227, 232, 239, 240, 242, 265, 267, and 275, wherein amino acid positions are numbered by correspondence with SEQ ID NO:2. The parent protease according to the twenty-first aspect of the invention may comprise an amino acid sequence having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% identity to the amino acid sequence of SEQ ID NO:2 and the variant according to the twenty-first aspect of the invention, may comprise a total of three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14 or 15 mutations selected from groups (a) and (b) below, wherein at least one mutation is selected from group (a):
(a) A1E/T, S9/T, P14L, A15L, L16C, H17T, S18L/T, Q19K, G20A, Y21N/T, T22L, G23E, S33T, K43Y, N76D, G102A, N109A/S/G, A137V, K141R, T158S, G169A, S204E, P210S, N218S, N243P/V, S248N/A, S249A, K256R, L257G, S260P, and N269D; and
[0227] (b) S24G/R/E, N25G, P40A/E, P52L, S53G, T55P, F58G, Q59S, N61E/P/G/S, N62Q/R/S, S63G/H, V68A, S78N, P86S, S87D/G, A88T/V, S89Y, A92G, L96T, G97A, G100N/Q/T, S101N, Q103E/H, Y104N, W106F, I111V, A114G, I115V, A116N/T, N117S, N118G, N123A/G/Q/V, M124I/V, S125A, L126A, G128A/S, P129E/Q/S/V, S130G, G131S/H, S132N, A133V, A134T, A144K, S145D, S159K, S161P, S162G/K, Y167A, P194L, V203Y, Q206D/E, K213L, Y217Q/L/D, V227T, A232T, P239R/V, N240K, T242R, K265N, L267V, and Q275E. A variant and parent protease according to the second aspect of the invention may be in a mature form.
[0228] A variant according to the twenty-first aspect of the invention may have a total net charge of -1, 0 or +1 relative to the BPN' wild-type.
[0229] In the first, second, fourth, fifth, sixth, seventh, eighth, ninth, or twenty-first aspect of the invention, the variant may be a subtilisin protease variant having improved wash performance or cleaning performance in a detergent as compared to that of the parent subtilisin protease, which may be a subtilisin protease. In the third aspect of the invention, the polypeptide may have improved wash performance or cleaning performance in a detergent as compared to that of the protease of SEQ ID NO:2, SEQ ID NO:4 or SEQ ID NO:6. Exemplary detergents for such aspects are shown in the Part I Examples and Part II Examples.
[0230] A protease variant according to the first, second, third, fourth, fifth, sixth, seventh, eighth, ninth, or twenty-first aspect of the invention may be a cold water protease. In one aspect, a protease variant according to the first, second, third, fourth, fifth, sixth, seventh, eighth, ninth, or twenty-first aspect of the invention may be a cold water protease having: (i) a performance index (PI) greater than 1, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from at least 1 to about 10, from at least 1 to about 8, or from at least 1 to about 5 on BMI at pH 8 and 60.degree. F. when compared to an enzyme having the amino acid sequence of SEQ ID NO:4, as defined in the Test Method set forth in Part I Example 1; or (ii) a performance index of at least 1, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from at least 1 to about 10, from at least 1 to about 8, or from at least 1 to about 5 on BMI at pH 8 and 60.degree. F. when compared to an enzyme having the amino acid sequence of SEQ ID NO:6, as defined in the Test Method set forth in Part I Example 1.
[0231] As noted above, the protease variant polypeptides of the invention have enzymatic activities (e.g., proteolytic activities) and thus are useful in cleaning applications, including but not limited to, methods for cleaning dishware items, tableware items, fabrics, and items having hard surfaces (e.g., the hard surface of a table, table top, wall, furniture item, floor, ceiling, etc.). Exemplary cleaning compositions comprising one or more protease variant polypeptides of the invention are described infra. The enzymatic activity (e.g., protease activity) of a protease variant polypeptide of the invention can be determined readily using procedures well known to those of ordinary skill in the art. The Examples presented infra describe methods for evaluating the enzymatic activity, cleaning performance, and/or washing performance. The performance of protease variants of the invention in removing stains (e.g., a proteinaceous stain), cleaning hard surfaces, or cleaning laundry, dishware or tableware item(s) can be readily determined using procedures well known in the art and/or by using procedures set forth in the Part I Examples and/or Part II Examples.
[0232] A polypeptide of the invention can be subject to various changes, such as one or more amino acid insertions, deletions, and/or substitutions, either conservative or non-conservative, including where such changes do not substantially alter the enzymatic activity of the polypeptide. Similarly, a nucleic acid of the invention can also be subject to various changes, such as one or more substitutions of one or more nucleic acids in one or more codons such that a particular codon encodes the same or a different amino acid, resulting in either a silent variation (e.g., mutation in a nucleotide sequence results in a silent mutation in the amino acid sequence, for example when the encoded amino acid is not altered by the nucleic acid mutation) or non-silent variation, one or more deletions of one or more nucleic acids (or codons) in the sequence, one or more additions or insertions of one or more nucleic acids (or codons) in the sequence, and/or cleavage of or one or more truncations of one or more nucleic acids (or codons) in the sequence. Many such changes in the nucleic acid sequence may not substantially alter the enzymatic activity of the resulting encoded protease variant compared to the protease variant encoded by the original nucleic acid sequence. A nucleic acid of the invention can also be modified to include one or more codons that provide for optimum expression in an expression system (e.g., bacterial expression system), while, if desired, said one or more codons still encode the same amino acid(s).
[0233] The present invention includes a genus of polypeptides comprising protease variant polypeptides having the desired enzymatic activity (e.g., protease activity or cleaning performance activity) which comprise sequences having the amino acid substitutions described herein and also which comprise one or more additional amino acid substitutions, such as conservative and non-conservative substitutions, wherein the polypeptide exhibits, maintains, or approximately maintains the desired enzymatic activity (e.g., protease activity or subtilisin activity, as reflected in the cleaning activity or performance of the protease variant). Amino acid substitutions in accordance with the invention may include, but are not limited to, one or more non-conservative substitutions and/or one or more conservative amino acid substitutions. A conservative amino acid residue substitution typically involves exchanging a member within one functional class of amino acid residues for a residue that belongs to the same functional class (identical amino acid residues are considered functionally homologous or conserved in calculating percent functional homology). A conservative amino acid substitution typically involves the substitution of an amino acid in an amino acid sequence with a functionally similar amino acid. For example, alanine, glycine, serine, and threonine are functionally similar and thus may serve as conservative amino acid substitutions for one another. Aspartic acid and glutamic acid may serve as conservative substitutions for one another. Asparagine and glutamine may serve as conservative substitutions for one another. Arginine, lysine, and histidine may serve as conservative substitutions for one another. Isoleucine, leucine, methionine, and valine may serve as conservative substitutions for one another. Phenylalanine, tyrosine, and tryptophan may serve as conservative substitutions for one another.
[0234] Other conservative amino acid substitution groups can be envisioned. For example, amino acids can be grouped by similar function or chemical structure or composition (e.g., acidic, basic, aliphatic, aromatic, sulfur-containing). For instance, an aliphatic grouping may comprise: Glycine (G), Alanine (A), Valine (V), Leucine (L), Isoleucine (I). Other groups containing amino acids that are considered conservative substitutions for one another include: aromatic: Phenylalanine (F), Tyrosine (Y), Tryptophan (W); sulfur-containing: Methionine (M), Cysteine (C); Basic: Arginine (R), Lysine (K), Histidine (H); Acidic: Aspartic acid (D), Glutamic acid (E); non-polar uncharged residues, Cysteine (C), Methionine (M), and Proline (P); hydrophilic uncharged residues: Serine (S), Threonine (T), Asparagine (N), and Glutamine (Q). Additional groupings of amino acids are well-known to those of skill in the art and described in various standard textbooks. Listing of a polypeptide sequence herein, in conjunction with the above substitution groups, provides an express listing of all conservatively substituted polypeptide sequences.
[0235] More conservative substitutions exist within the amino acid residue classes described above, which also or alternatively can be suitable. Conservation groups for substitutions that are more conservative include: valine-leucine-isoleucine, phenylalanine-tyrosine, lysine-arginine, alanine-valine, and asparagine-glutamine. Thus, for example, the invention includes an isolated or recombinant protease variant polypeptide (e.g., subtilisin variant) having proteolytic activity, said protease variant polypeptide comprising an amino acid sequence having at least about 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.5% sequence identity to the amino acid sequence of SEQ ID NO:2. A conservative substitution of one amino acid for another in a protease variant of the invention is not expected to alter significantly the enzymatic activity or cleaning performance activity of the protease variant. Enzymatic activity or cleaning performance activity of the resultant protease can be readily determined using the standard assays and the assays described herein.
[0236] Conservatively substituted variations of a polypeptide sequence of the invention (e.g., protease variants of the invention) include substitutions of a small percentage, sometimes less than about 25%, 20%, 15%, 14%, 13%, 12%, 11%, 10%, 9%, 8%, 7%, or 6% of the amino acids of the polypeptide sequence, or less than about 5%, 4%, 3%, 2%, or 1%, of the amino acids of the polypeptide sequence, with a conservative amino acid.
[0237] As described elsewhere herein in greater detail and in the Examples provided herein, polypeptides of the invention may have cleaning abilities that may be compared to known proteases, including known subtilisins. Exemplary known subtilisin proteases include, but are not limited to, for example, B. lentus subtilisin GG36, B. amyloliquefaciens subtilisin BPN', B. amyloliquefaciens subtilisin BPN'-Y217L, and B. clausii PB92.
[0238] Numerous polypeptides of the invention, including protease variants, such as, e.g., subtilisin protease variants described and set forth throughout the specification, including, but not limited to, in the Examples. Part I Examples describe protease variants of the invention, including, e.g., but not limited to, BPN' protease variants. Part II Examples describe additional protease variants of the invention, including, e.g., but not limited to, GG36 protease variants.
[0239] The present invention provides protease variants, including serine protease variants, having one or more substitutions as compared to a reference serine protease. In one aspect, the present invention provides cold water proteases. In addition, the present invention provides compositions comprising one or more such protease variants, e.g., serine protease variants. A composition may comprise at least one such protease variant of the invention and an adjunct ingredient, as described elsewhere herein. In one aspect, the present invention provides cleaning compositions comprising at least one of these protease variants, e.g., serine protease variants. A cleaning composition may be a detergent composition. The trend in cleaning is to use lower wash temperatures to save energy. Enzymes have lower activity at lower temperatures resulting in reduced cleaning performance. There is a need in the art for enzymes with enhanced performance at coldwater washing conditions over those currently known in the art. The present invention addresses this need.
[0240] The present invention provides cleaning compositions comprising at least one serine protease variant described herein, such as a subtilisin protease variant. The subtilisin protease variant may be a BPN' protease variant. As discussed in further detail supra, the invention includes a composition comprising a BPN' protease variant and a GG36 protease variant. Some protease variants, including, but not limited to, e.g., subtilisin variants of the invention, are cold water proteases. The cleaning composition may be a laundry detergent. The laundry detergent may be a detergent having a pH between 3 and 11 (e.g., between pH 4, pH 5, pH 6, pH 7, pH 7.5, pH 8, pH 9, pH 10, pH 10.5, etc.) cold water detergent, a low pH detergent (e.g., pH 3-6), neutral pH detergent (e.g., pH 6.5-7.5), alkaline pH detergent (e.g., pH 9-11) or a compact detergent. A detergent may include phosphate or be without phosphate. The cleaning composition may be a dishwashing detergent. In one aspect, the dishwashing detergent is a phosphate-free detergent, while in another aspect, the dishwashing detergent is a phosphate-containing detergent. In one aspect, the cleaning composition of the invention further comprises at least one additional enzyme, which may optionally be selected from the group of a neutral metalloprotease, lipase, cutinase, amylase, carbohydrase, cellulase, pectinase, mannanase, arabinase, galactanase, xylanase, perhydrolase, oxidase, and peroxidase. Also provided are isolated nucleic acids encoding a serine protease variant of the invention, expression vectors comprising one or more such nucleic acids of the invention, and host cells comprising at least one such expression vector of the invention.
[0241] Throughout the specification, for ease of reference, a "set of amino acid substitutions" or "set of substitutions" may refer to a set of multiple amino acid substitutions (i.e., G097A+G128A+Y217Q) or set of a single amino acid substitution (i.e., Y217Q). Thus, a BPN' variant comprising the BPN' sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q and Y217Q indicates the BPN' variant comprises an amino acid sequence comprising BPN'-G097A-G128A-Y217Q or BPN'-Y217Q.
[0242] The amino acid sequence of the mature BPN'-v3 subtilisin protease variant, which is set forth in SEQ ID NO:4, can be represented as BPN'-G097A-G128A-Y217Q, which means the BPN' amino acid sequence of SEQ ID NO:2 with the three substitutions G097A,G128A, and Y217Q. In this format, each dash (-) is equivalent to using a plus sign (+). Thus, BPN'-G097A-G128A-Y217Q can be written alternatively as BPN'+G097A+G128A+Y217Q. The amino acid sequence of the mature BPN'-v36 subtilisin protease variant, which is set forth in SEQ ID NO:6, can be written as BPN'-S24G-S53G-S78N-S101N-G128A-Y217Q or BPN'+S24G+S53G+S78N+S101N+G128A+Y217Q. BPN' variants of the invention may be depicted using these formats.
[0243] In addition, the present invention provides subtilisin variants, wherein each such variant is a mature form having proteolytic activity and comprises the BPN'-v3 amino acid sequence (SEQ ID NO:4) comprising at least one set of amino acid substitutions selected from the group consisting of S87T-A88L-S89G, N61P-S63H, S87G-A88V-S89A, P86S-S87G-A88V, Q59S-N61P, S24G-N25G, N61P-N62S, P129Q-S130G-G131S, L75S-N76Y, V203Y, T55P, A88V-L901, G211R-N212S-K213V, G23A-S24G-N25G, T22N-S24A, S24R, A98S, T158G-S159G, Q59E-N61P, and A98E, and wherein the amino acid positions are numbered by correspondence with the amino acid sequence of B. amyloliquefaciens subtilisin BPN' set forth as SEQ ID NO:2.
[0244] Note that the BPN'-v3 amino acid sequence, which is set forth as SEQ ID NO:4, may be written as BPN'-G97A-G128A-Y217Q. The invention includes a protease variant having proteolytic activity, wherein said variant comprises the amino acid sequence of SEQ ID NO:4 having an amino acid substitution A128S in SEQ ID NO:4. Thus, the invention includes the amino acid sequence BPN'-G97A-G128A-Y217Q. Throughout this specification, polypeptide variants of the invention may be described as a variant of a reference polypeptide comprising one or more particular amino acid substitutions at one or more specified positions in the reference polypeptide sequence, respectively.
[0245] The present invention also provides subtilisin variants, wherein each such variant is a mature form having proteolytic activity and comprises the BPN'-v3 amino acid sequence (SEQ ID NO:4) comprising at least one set of amino acid substitutions selected from the group consisting of P86S-S87G-A88V-A116N-N117S-N118G, S24G-N25G-N61P-S101N, S24G-N25G-S53G-T55P-S87T-A88L-S89G-S101N-V203Y, N61P-S78N-S101N-V203Y, T55P-N61P-S78N-S101N-V203Y, S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N, V203Y-L267V, S24G-N25G-T55P-S101N, A134T-L267V, S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G, S24G-N25G-S53G-N61P-S101N-V203Y, N25Y-Q59S-N61P, I111V-S161P, I115V-L267V, T55P-S78N-S87T-A88L-S89G-S101N-V203Y, N25Y-P129Q-S130G-G131S-A137T, N61P-S63H-A128S-P129Q, S53G-N61P-S101N-V203Y, S24G-N25G-S53G-S78N-S87T-A88L-S89G-S101N, N61P-S78N-S87T-A88L-S89G-S101N, N25Y-N61P-S63H, Q59S-N61P-V203Y, V8L-N25Y-P129Q-S130G-G131S, P86S-S87G-A88V-P239R, S24G-N25G-S53G-T55P-N61P-S101N-V203Y, S24G-N25G-P129Q-S130G-G131S, N240K, G23A-S24G-N25G-G211R-N212S-K213V, N61P-S63H-S78N-I111V-A134T, S63T-P86S-S87G-A88V, G23A-S24G-N25G-A116N-N117S-N118G, S78N-S87T-A88L-S89G-S101N, S24G-N25G-A116N-N117S-N118G, T55P-N240K, T55P-P129V-P194S, N25Y-S87G-A88V-S89A, S24G-N25G-S87T-A88L-S89G-S101N, P129Q-S130G-G131S-V203Y, Q59S-N61P-N240K, S24R-P40E-P129E-S159K-K265R, P525-T55P-V203Y, S24R-P129E, S24G-N25G-S53G-N61P-S78N, S24G-N25G-T55P-S78N-S101N, P86S-S87G-A88V-A116S-N117G-N118R, N61P-S87T-A88L-S89G, S24G-N25G-S53G-T55P-S78N-S87T-A88L-S89G, G23A-S24G-N25G-N61P-S63H, S24R-Q59S-N61P, N61P-P129Q-S130G-G131S, S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N, S24G-N25G-S53G-T55P-S101N-V203Y, N61P-S78N-S87T-A88L-S89G-S101N-V203Y, S24G-N25G-S53G-T55P-S78N-S101N, S24G-N25G-S53G-S101N-V203Y, S24G-N25G-S78N-S101N-V203Y, P129Q-S130G-G131S-A133V-L267V, S87T-A88L-S89G-S101N, G23A-S24G-N25G-P239R, S87G-A88V-S89A-A116N-N117S-N118G, Q59S-N61P-A116S-N117G-N118R, Q59S-N61P-S87T-A88L-S89G, S24G-N25G-S53G-S87T-A88L-S89G-V203Y, A134T-G211T, T55P-A128S-P129Q, T55P-S78N-S87T-A88L-S89G-S101N, P86S-S87G-A88V-T242R, S161P-V203Y, S24G-N25G-T55P-N61P-S78N-S101N-V203Y, G211T-L267V, P40E-T55P-N269K, S24R-A128S-P129G, S24G-N25G-N61P-N62S-P194L-A232T, T55P-A116S-N117G-N118R, S24G-N25G-S53G-S78N-S101N-V203Y, P129Q-S130G-G131S-N240K, S53G-T55P-N61P-S78N-S87T-A88L-S89G, N25Y-P129Q-S130G-G131S, T55P-I115V, N25Y-T55P, G23A-S24G-N25G-A128S-P129D, S53G-S78N-S87T-A88L-S89G-S101N-P129S-V203Y, T55P-A134T, N61P-S63H-S78N-I111V, N61P-A97G-G102A-A128G-P129S, S53G-N61P-S101N, Q59S-N61P-S87G-A88V-S89A, S53G-S87T-A88L-S89G-S101N-V203Y, S87T-A88L-S89G-P129S, S53G-T55P-S78N-S101N-V203Y, T55P-P129Q-S130G-G131S, Q59S-N61P-P129Q-S130G-G131S, A134T-P239R, T55P-V203Y, T55P-S78N-S89Y, T22N-S24A-N61P-S63H, S161P-L267V, T55P-L75H-N76G, A134T-S161P, S87T-A88L-S89G-A134T, T55P-A116N-N117S-N118G, A128S, T55P-S78N-I115V, Y6Q-P129Q-S130G-G131S, S24R-P129Q-S130G-G131S, S24G-N25G-S53G-S78N-S101N, T55P-P129V, N61P-N62Q-G100N-A128G, T55P-P129Q, S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, S87T-A88L-S89G-N240K, A134T-N240K, S87T-A88L-S89G-P239R, P129Q-S130G-G131S-L267V, P129Q-N240K, S78N-S87T-A88L-S89G-V203Y, I111V-A273S, S24G-N25G-T55P-S78N-A88V-S101N, S24G-N25G-T55P-S78N, S24G-N25G-S53G-S78N-S87T-A88L-S101N-V203Y, S24G-N25G-S53G-S78N-S87T-A88L-S89G-V203Y, S24G-N25G-S53G-S78N-S87T-A88L-S89G-S101N-V203Y, S87G-A88V-S89A-P129Q-S130G-G131S, N61P-S63H-S78N-S161P, T55P-N61P-S78N-S87T-A88L-589G-S101N-V203Y, I111V-P129Q-S130G-G131S, T22N-S24A-T55P, I115V-N240K, S87G-A88V-S89A-A116N-N117S-N118G-P172H, S24G-N25G-S78N-S87T-A88L-S89G-S101N, S24G-N25G-I115V-A134T, T55P-A128S-P129D, I111V-S159K, N240K-A273S, S159K-L267V, I111V-P129Q-G211T, I115V-A273S, S89Y, S24R-A116N-N117S-N118G, N61E-A144K, P129Q-S130G-G131S-P239R, S87T-A88L-S89G-I115V, T55P-A92G, S145D-S159K-N240K-Q275E, S89Y-P129Q-S130G-G131S, P129Q-S130G-G131S-S162K, I111V-A134T, P40E-S53Y-S78Y-P86S-S87G-A88V, S24G-N25G-L75H-N76G, N61P-A128G-P129S-S130P, S24R-S145D, S24R-S145D-P239R-Q275E, S24R-S78N-S182P-L267V, S53G-N61P-S87T-A88L-S89G-S101N-V203Y, P5S-S87G-A88V-S89A-A116G-N117R, S53G-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, Q59S-N61P-A116N-N117S-N118G, P239R-A273S, S53G-S78N-S87T-A88L-S89G-S101N-V203Y, S24R-P129V, I111V-P239R, S87T-A88L-S89G-S101N-V203Y, T55P-P129L, S87T-A88L-S89G-I111V, S145D-A273S, P129Q-S130G-G131S-T242R, S3F-S87T-A88L-S89G-G211T, S87G-A88V-S89A-S162K, S89Y-G211T, S87T-A88L-S89G-A144K, P129Q-S130G-G131S-S159K, A116N-N117S-N118G-P129Q-S130G-G131S, S24G-N25G-P129V, S24G-N25G-S78N-S87T-A88L-S89G-S101N-V203Y, N123G-A128G, N61P-N62Q-G100N-G102A-M124I, S24G-N25G-K141E-T242R, S87G-A88V-S89A-A116N-N117S-N118G-A144T, T55P-N61P-S87T-A88L-S89G-G110C-S130P, L75S-N76Y-A116S-N117G-N118R, S145D-S159K-K213L-P239R-N240K, S24R-S87T-A88L-S89G, G23A-S24G-N25G-P129V, A134T-K213L, S89Y-A273S, S24R-P239R, N123G-A128G-P129S, S89Y-P239R, S24G-N25G-A92G, N61P-S63H-I115V-A228V, A97G-A128G-Q217L, L75S-N76Y-P129V, S24R-P129L, S87G-A88V-S89A-P129Q-S182Y-S204Y-P239Q, S24R-A92G, S24R-A116S-N117G-N118R, G23A-S24G-N25G-A116G-N117R, S24G-N25G-P129L, S87T-A88L-S89G-S101G, G23A-S24G-N25G-P129L, S53G-N61P-G102A-V203Y, T55P-V147P, Y6Q-L75S-N76Y, N61P-S63H-V147P, S24R-V147P, S24G-N25G-V68C-A69G, S24G-N25G-N61P-L75S-N76Y-S101N-V203Y, L75 S-N76Y-S78N-S101N-V203Y, L75 S-N76Y-S78N-S87T-A88L-S89G-S101N-S130P, S24G-N25G-S53G-S101N-S130P-V203Y, G47E-M50I-L75S-N76Y-S162K, S53G-T55P-S87T-A88L-S89G-S101N-S130P-V203Y, S24G-N25G-L75S-N76Y-A128T-P129T-S130G-G131Q-S132C-A133G-A134T, P129Q-S130G-G131S-V147P, S53G-T55P-N61P-L75S-N76Y-S87T-A88L-S89G-G102A-S130P-V203Y, and S53G-T55P-N61P-L75 S-N76Y-S101N-S130P-V203Y, and wherein the amino acid positions are numbered by correspondence with the amino acid sequence of B. amyloliquefaciens subtilisin BPN' set forth as SEQ ID NO:2.
[0246] The present invention also provides subtilisin variants, wherein the variant is a mature form having proteolytic activity and comprises the BPN'-v3 amino acid sequence (SEQ ID NO:4) comprising at least one set of amino acid substitutions selected from the group consisting of P40E-S78N-S87D-Y217L, P40E-Y217L, T22V-S78N-Q206E-K213N-Y217L, T22V-S78N-K213N-Y217L, S87D-Y217L, S78N-Y217L, K213N-Y217L, Q206E-Y217L, and T22V-Y217L, and wherein the amino acid positions are numbered by correspondence with the amino acid sequence of B. amyloliquefaciens subtilisin BPN' of SEQ ID NO:2.
[0247] The present invention also provides subtilisin variants, wherein each such variant is a mature form having proteolytic activity and the variant comprises the BPN'-v3 amino acid sequence (SEQ ID NO:4) comprising at least one set of amino acid substitutions selected from the group consisting of S87D-N76D-S78N, P40E-S78N-S87D, P40E-S87D, S78N-P40E, S87D-N76D, P40E, S78N-S87D, S87D, and S78N, and wherein the amino acid positions are numbered by correspondence with the amino acid sequence of B. amyloliquefaciens subtilisin BPN' of SEQ ID NO:2. In one aspect, the substitutions confer improved enzyme stability.
[0248] The present invention also provides subtilisin variants, wherein each such variant is a mature form having proteolytic activity and comprises the BPN'-v3 amino acid sequence (SEQ ID NO:4) comprising at least one set of amino acid substitutions selected from the group consisting of S101N, A137V, N61P, S130P, Q103N, S63T, G102A, N109D-S248R, 587R, S188D, S87D-S248R, S188D-S248R, S248D, N109D-S188D-S248R, N109D, 587R-S248R, N109D-S188R, N76D, S87D-N109D-S188D-S248R, S87R-N109D-S188D-S248R, S87R-S188R-S248R, A187D, N109D-S248D, S87R-N109R-S188R-S248R, F189D, G100N, S87R-N109D-S188D, S87D-N109D-S188D, S87R-S188D-S248D, N62D, and S87D-N109D-S188D-S248D, and wherein the amino acid positions are numbered by correspondence the positions of with the amino acid sequence of B. amyloliquefaciens subtilisin BPN' of SEQ ID NO:2.
[0249] The present invention also provides subtilisin variants, wherein each such variant is a mature form having proteolytic activity and comprises the BPN'-v3 amino acid sequence (SEQ ID NO:4) comprising at least one set of amino acid substitutions selected from S78N-L267V, S78N-S161P, S78N-I115V, S78N-A273S, S78N-G211T, S78N, S78N-I111V, S78N-V147L, S78N-I108V, S78N-S89Y, S78N-A138T, S78N-P172V, S78N-Q59G, P129T-V147Q-S159D-S161P-S183T-Q185T-G211A-S224A, Q059V-I108V-V147Q-G211A-N252Q, S78N-Y167A, S78N-A92G, S78N-P129L, N061A-S087E-M124I-S161P-S224A, S78N-N62Q, S78N-V68A, S063T-S101A-L126V-S183T-T244N, S78N-M124T, P040L-S053G-Q059V-N061A-N062Q-S063T-S087E-G100N, N062Q-G100N-S125A-S159D-N240S, V68A-G102A-G211A-S125A, and S053G-V068A-G102A-P129T-Q185T, and wherein the amino acid positions are numbered by correspondence with the amino acid residue positions of the amino acid sequence of B. amyloliquefaciens subtilisin BPN' set forth as SEQ ID NO:2.
[0250] The present invention also provides subtilisin variants, wherein each such variant is a mature form having proteolytic activity and comprises an amino acid sequence comprising at least one set of amino acid substitutions selected from the group consisting of G97A-G128S-Y217Q-S24G-N25G-N61P-S101N, G97A-G128S-Y217Q-S53G-N61P-S101N-V203Y, G97A-G128S-Y217Q-S24G-N25G-S53G-T55P-N61P-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-S53G-N61P-S101N-V203Y, P52L-V68A-G97A-I111V, I111V-M124V-Y167A-Y217Q, Y104N-G128A-Y217Q, M124V-Y167A-Y217Q, I111V-M124V-Y217Q, P52L-V68A-G97A, G97A-I111V-M124V, V68A-A92G-G97A, G97A-I111V-M124V-Y167A-Y217Q, P52L-V68A-I111V-Y217Q, P52L-V68A-I111V, V68A-A92G-I111V, P52L-V68A-G97A-I111V-Y217Q, V68A-G97A-I111V, G97A-I111V-Y217Q, G97A-I111V-M124V-Y167A, S89Y-I111V-M124V, V68A-S89Y-I111V, V68A-A92G-Y217Q, I111V-Y167A-Y217Q, G97A-I111V-Y167A-Y217Q, G97A-I111V-M124V-Y217Q, V68A-I111V-Y167A-Y217Q, I111V-G128A-Y217Q, G97A-M124V-Y217Q, V68A-Y167A-Y217Q, I111V-M124V-Y167A, N62Q-G97A-I111V, G97A-M124V-Y167A-Y217Q, G97A-L126A-Y217Q, V68A-I111V-Y217Q, S89Y-M124V-Y217Q, and L96T-G97A-Y217Q, and wherein the positions are numbered by correspondence with the amino acid residue positions of the amino acid sequence of B. amyloliquefaciens subtilisin BPN' set forth as SEQ ID NO:2. Some such variants comprise the BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from said group.
[0251] The present invention also provides subtilisin variants, wherein each such variant is a mature form having proteolytic activity and comprises a BPN' amino acid sequence comprising at least one, two, three, four, five, six, seven, eight, nine, ten, eleven or more substitutions selected from S182E, N109I, N117H, K237D, L257Q, P225N, S105H, S236I, L235H, S249E, N76E, S145N, N243D, R247N, E195N, A98K, S182N, S161H, G83H, G131D, T71C, K136Q, P40D, A187H, L250K, S9I, N76M, S132D, Q19F, E112H, S249P, S53D, V68E, D41I, K43H, V4H, A13Y, N62P, L196E, and V44D, and wherein the positions are numbered by correspondence with the amino acid residue positions of amino acid sequence of B. amyloliquefaciens subtilisin BPN' set forth as SEQ ID NO:2.
[0252] The present invention also provides dishwashing compositions comprising the subtilisin variant(s), and fabric cleaning compositions comprising the subtilisin variant(s). In some preferred aspects, the dishwashing and fabric cleaning compositions further comprise at least one additional enzyme. In some preferred aspects, the additional enzyme is selected from: a protease (e.g., a neutral metalloprotease, a wild type serine protease, or a second variant serine protease) a lipase, a cutinase, an amylase, a carbohydrase, a cellulase, a pectinase, a mannanase, an arabinase, a galactanase, a xylanase, a perhydrolase, an oxidase, and a peroxidase. Moreover, the present invention provides dishwashing methods, comprising the steps of: providing i) the dishwashing composition comprising the subtilisin variant, and ii) dishware in need of cleaning; and contacting the dishware with the dishwashing composition under conditions effective to provide cleaning of the dishware. Similarly, the present invention provides fabric cleaning methods, comprising the steps of: providing i) the fabric cleaning composition comprising the subtilisin variant, and ii) laundry in need of cleaning; and contacting the laundry with the fabric cleaning composition under conditions effective to provide cleaning of the laundry. In still further aspects, the present invention provides an isolated nucleic acid encoding the variant, an expression vector comprising the isolated nucleic acid in operable combination with a promoter, and/or host cells comprising the expression vector are provided.
[0253] The present invention also provides cleaning compositions comprising the subtilisin variants provided herein. In one aspect, the cleaning compositions comprise a liquid, gel, tablet, powder and/or granule detergent. In some further aspects, the cleaning compositions are selected from laundry detergents and dish detergents. In some preferred aspects, the cleaning compositions comprise laundry detergents. In some particularly preferred aspects, the cleaning compositions are heavy duty detergents. In some additional aspects, the cleaning compositions comprise dish detergents selected from hand dishwashing and automatic dishwashing detergents. In some further preferred aspects, the cleaning compositions provided herein further comprise at least one bleaching agent. In some additional aspects, the cleaning compositions provided herein are phosphate-free, while in some alternative aspects, the cleaning compositions provided herein are phosphate-containing detergents. In some still further aspects, the cleaning compositions provided herein are cold water detergents. In yet some additional aspects, the cleaning compositions provided herein further comprise at least one additional enzyme. In one aspect, the cleaning compositions comprise at least one additional enzyme selected from the group consisting of a hemicellulase, cellulase, peroxidase, protease, metalloprotease, xylanase, lipase, phospholipase, esterase, perhydrolase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, B-glucanase, arabinosidase, hyaluronidase, chondroitinase, laccase, and amylase; or mixtures of any thereof.
[0254] The present invention also provides methods for cleaning, comprising providing an item to be cleaned and a composition comprising at least one cleaning composition provided herein, and contacting them with the composition, under conditions effective to provide cleaning of the item. In one aspect, the methods further comprise the step of rinsing the item after contacting the item with the cleaning composition. In some preferred aspects, the item to be cleaned comprises dishware. In some alternative aspects, the item to be cleaned comprises fabric.
[0255] In some further aspects, any of the above mutations, deletions, insertions or combinations thereof can be applied to either the enzyme of SEQ ID NO:2 containing the mutation Y217L or the enzyme of SEQ ID NO:2 containing the mutations G97A-G128A-Y217Q.
[0256] In one aspect, the present invention provides a suitable cold water protease derived from a subtilisin, particularly subtilisin BPN' (SEQ ID NO:2) (i.e., a BPN' subtilisin variant). In one aspect, a cold water protease one, two, three, four, five, six, seven, eight, nine, ten, eleven or more of the following amino acid substitutions relative to SEQ ID NO:2: Q002W, P005K, P005L, P005Y, G007T, V008G, V008K, V008P, I011G, I011H, I011S, K012L, A013M, L016W, G023S, V026H, V026W, V026Y, K027P, V028Q, V028S, V028T, A029C, A029D, A029S, A029T, A029V, V030D, V030E, V030G, V030T, I031E, I031G, I031H, I031K, I031N, I031Q, I031S, I031Y, S033F, S033H, D036L, S037P, D041A, D041C, D041M, D041N, D041S, L042S, L042Y, V044H, V044Q, V044T, G046F, G046L, G046M, G046V, G047T, G047W, S049I, S049V, M050D, M050I, M050R, V051A, V051G, V051S, P052C, P052I, P052L, P052M, P052V, P052W, P052Y, E054R, N056G, N056I, N056K, N056M, N056Q, N056R, N056V, N056Y, P057I, P057K, P057L, P057R, P057T, P057V, F058T, Q059P, Q059W, N061P, N062D, N062M, N062Q, S063C, S063Q, S063T, G065Q, V068A, V068G, V068M, V068S, A069C, A069D, A069F, A069H, A069M, A069N, A069P, A069Q, A069R, A069T, T071D, T071E, T071G, T071K, T071M, V072D, V072G, V072K, V072Q, V072S, V072T, A073E, A073I, A073K, A073M, A073Q, A073S, A073V, A074E, A074F, A074H, A074I, A074L, A074M, A074Q, A074R, A074V, A074W, A074Y, L075A, N076A, N077G, N077L, N077P, N077Q, N077R, N077S, N077T, S078W, G080H, V081D, V081F, V081H, V081N, V081Q, V081R, V081W, L082G, L082N, L082W, A085I, A085T, A085V, P086A, P086G, P086M, P086Q, P086R, P086T, P086W, P086Y, S087F, S087I, S087L, S087M, S087Q, S087V, S087W, A088D, A088E, A088K, A088P, A088Q, S089D, S089Q, S089V, L090D, L090E, L090F, L090H, L090P, L090S, L090T, Y091L, Y091Q, Y091T, A092C, A092I, A092M, A092N, A092P, A092V, V093D, V093F, V093L, V093T, K094C, K094R, K094S, V095I, L096F, L096H, L096I, L096M, L096N, L096Q, L096S, L096T, L096V, L096W, L096Y, G097A, G097C, G097D, G097E, G097F, G097L, G097M, G097P, G097Q, G097S, G097V, G097W, G097Y, D099C, D099E, D099I, D099M, D099P, D099V, D099Y, G100D, G100E, G100H, G100I, G100K, G100M, G100Q, G100T, G100V, G100Y, S101A, S101E, S101G, S101N, S101P, S101Q, S101T, S101V, G102A, G102S, Q103E, Q103G, Q103H, Q103K, Q103N, Y104L, Y104M, Y104N, Y104T, Y104V, S105D, S105I, S105R, S105V, W106A, W106C, W106E, W106F, W106G, W106I, W106L, W106M, W106S, W106T, W106V, I107R, I107S, I107T, I108S, I108T, G110S, G110T, I111A, I111C, I111F, I111L, I111M, I111T, I111V, E112I, E112L, E112T, W113H, A114I, A114V, I115A, I115E, I115F, I115H, I115M, I115N, I115P, I115Q, I115R, I115S, I115T, I115V, I115Y, N117K, N117V, N117W, N118I, N118L, N118V, M119F, M119N, D120Y, I122R, I122S, I122T, N123A, N123C, N123G, N123S, M124A, M124H, M124N, M124Q, M124S, M124T, M124V, S125A, L126A, L126Q, L126S, L126T, L126Y, G128E, G128N, G128T, P129V, S130P, S132I, S132N, S132P, S132Q, S132V, A134I, A134L, A134M, L135I, L135T, L135V, L135W, L135Y, K136D, K136F, K136I, K136V, K136Y, A137P, A138D, A138E, A138F, A138H, A138Q, A138Y, A142G, A142I, A142T, A142V, V143W, A144P, G146L, G146T, G146Y, V147D, V147P, V147W, V147Y, V148N, V148Y, V149E, A151T, A153T, A153V, S159K, T164W, V165T, Y167A, Y167D, Y167E, Y167M, Y167P, Y167S, Y167T, P168L, P168T, G169C, K170E, K170N, K170P, K170Q, K170S, K170T, K170Y, Y171C, Y171D, Y171L, Y171N, Y171W, P172E, P172G, P172I, P172L, P172V, P172Y, S173H, S173W, S173Y, V174A, V174I, V174L, V174S, I175A, I175F, I175R, I175T, I175V, A176V, V177W, V180F, V180H, V180Q, S182W, N184A, N184L, N184V, Q185D, Q185E, Q185I, Q185S, Q185T, R186C, R186D, R186E, R186F, R186G, R186I, R186L, R186N, R186P, R186Q, R186S, R186T, R186V, R186W, R186Y, A187D, A187E, A187Q, S190G, S190N, Y192D, Y192E, Y192L, Y192Q, G193H, G193Q, G193T, G193V, P194E, P194I, P194L, P194M, P194N, P194T, P194V, E195C, E195K, E195W, L196I, L196M, L196T, L196V, D197I, D197M, M199F, M199Q, A200C, A200H, A200N, A200T, A200V, A200Y, P201L, P201T, P201V, V203G, I205L, I205T, T208C, T208L, I208M, I208P, I208V, L209C, L209W, P210C, P210D, P210E, P210F, P210G, P210Q, P210R, P210S, P210V, G211A, G211D, G211E, G211P, G211T, G211V, G211W, N212E, N212T, K213Q, K213I, Y214A, Y214D, Y214N, Y214P, Y214S, G215D, G215Q, G215V, A216E, Y217E, Y217L, Y217M, N218P, A223W, S224D, S224N, S224Q, H226E, H226T, V227G, V227L, V227S, A230H, A230N, A231W, A231Y, A232N, I234W, L235N, S236W, H238A, H238G, H238I, P239H, P239S, W241G, W241Q, R247H, R247L, R247W, R247Y, L250E, L250T, N252Q, T253Y, T254D, T254Q, T254R, T255L, T255P, K256G, K256R, G258P, Y263D, Y263K, Y263R, K265P, I268S, 12681, I268W, V270F, A273K, A273P, A273R, A273V, A273W, A274W, S182E, and N109I, wherein positions of the variant sequence are numbered by correspondence to positions of SEQ ID NO:2.
[0257] In another aspect, the invention provides suitable cold water proteases, including subtilisin variants, particularly variant of mature BPN' (SEQ ID NO:2), comprising one, two, three, four, five, six, seven, eight, nine, ten, eleven or more of the following sets of mutations relative to SEQ ID NO:2: N109D-Y217L-S248R, N109D-S188R-Y217L, S87D-Y217L-S248R, S87R-N109D-Y217L-S248R, S87R-N109D-S188D-Y217L-S248R, G128A-Y217Q, I111V-M124V, M124V-Y217Q, N62Q-G97A, S89Y-M124V, V68A, V68A-A92G, V68A-G97A, V68A-I111V, V68A-S89Y, V68A-V227T, V68A-Y217Q, W106F-Y217Q, G97A-G128A-Y217Q, G97A-L126A-Y217Q, G97A-M124V-L126A-Y217Q, G97A-N123G-Y217Q, L96T-G97A-Y217Q, M124V-L126A-Y217Q, N62Q-G128A-Y217Q, N62Q-G97A-Y217Q, G97N-G128A-Y217M, G97G-G128S-Y217E, G97A-G128A-Y217Q, G97M-G128S-Y217E, G97A-G128S-Y217Q, G97D-G128S-Y217Q, G97M-G128G-Y217M, G97G-G128S-Y217Q, G97S-G128S-Y217Q, G97G-G128A-Y217Q, G97S-G128A-Y217E, G97A-G128S-Y217L, G97A-G128A-Y217N, G97Q-G128S-Y217L, G97A-G128A-Y217M, G97A-G128A-Y217S, G97D-G128A-Y217Q, G97M-G128S-Y217Q, G97Q-G128G-Y217D-S87Y, G97S-G128A-Y217N, G97A-G128S-Y217T, G97D-G128S-Y217E, G97D-G128A-Y217L, G97G-G128S-Y217E-S78P-A272T, G97T-G128S-Y217D, G97D-G128A-Y217I, G97Q-G128S-Y217Q, G97G-G128A-Y217D, G97Q-G128A-Y217N, G97S-G128A-Y217M, G97S-G128S-Y217N, G97S-G128S-Y217M, G97E-G128S-Y217M, G97S-G128P-Y217Q, G97T-G128S-Y217Q, G97D-G128S-Y217Q-A73T, G97E-G128S-Y217N, G97G-G128A-Y217I, G97Q-G128A-Y217D, G97Q-G128S-Y217M, G97R-G128T-Y217Q-S162P, G97S-G128S-Y217D, G97T-G128P-Y217I, G97Q-G128G-Y217E, G97C-G128G-Y217N, G97D-G128S-Y217H, G97M-G128S-Y217L, G97M-G128S-Y217N, G97S-G128S-Y217E, G97M-G128S-Y217I, G97A-G128P-Y217A, G97R-G128S-Y217D, G97A-G128A-Y217Q-S145D, G97A-G128A-Y217Q-P239R, G97A-G128A-Y217Q-N61E-P129E-S162K-K213L-N240K, G97A-G128A-Y217Q-N61E, G97A-G128A-Y217Q-P40E-A144K-K213L, G97A-G128A-Y217Q-P129E, G97A-G128A-Y217Q-N61E-P129E-S159K, G97A-G128A-Y217Q-K213L, G97A-G128A-Y217Q-S87D, G97A-G128A-Y217Q-Q206E, G97A-G128A-Y217Q-S24R-P40E-S145D-S159K-K213L, G97A-G128A-Y217Q-K265N, G97A-G128A-Y217Q-S24R, G97A-G128A-Y217Q-P40E, G97A-G128A-Y217Q-Q275E, G97A-G128A-Y217Q-P129E-S145D-N240K, G97A-G128A-Y217Q-A144K, G97A-G128A-Y217Q-S159K, G97A-G128A-Y217Q-S162K, G97A-G128A-Y217Q-N240K, G97A-G128A-Y217Q-S53G, G97A-G128A-Y217Q-S78N, G97A-G128A-Y217Q-S53G-S78N, G97A-G128A-Y217Q-S53G-I111V, G97A-G128A-Y217Q-S53G-N117S, G97A-G128A-Y217Q-S53G-S132N, G97A-G128A-Y217Q-Y104N-S132N, G97A-G128A-Y217Q-S53G-S78N-I111V, G97A-G128A-Y217Q-S53G-S78N-N117S, G97A-G128A-Y217Q-S53G-S78N-S132N, G97A-G128A-Y217Q-S53G-Y104N-S132N, G97A-G128A-Y217Q-S78N-Y104N-S132N, Y217L-V068C-A069G, Y217L-I079F-A098G, Y217L-P086T-S101D-Q103S-V147I, Y217L-A088T-P129S-G146D, Y217L-V093I-G128D-P129R, Y217L-Z096.01D-A098R, Y217L-Z096.01H-A098G, Y217L-G097S-Z097.01S-A098G-A273T, Y217L-A098S-D099G-G100D, Y217L-Z098.01N, Y217L-D099G-Z099.01N, Y217L-D099G-Z099.01S, Y217L-D099V-S101D, Y217L-Z099.01S, Y217L-G100D, Y217L-S101D-Q103H, Y217L-S101G-A151V, Y217L-S101H-G102S, Y217L-S101H-Q103D, Y217L-G102R-Q103C-Y104C-V192I, Y217L-Q103D, Y217L-V121I-I122S-N123C, Y217L-V121L-N123C, Y217L-I122S-N123S, Y217L-M124I, Y217L-M124V, Y217L-L126F-P129Z-S182N, Y217L-L126Y, Y217L-G127S-P129D, Y217L-Z127.01N-G128S-P129S, Y217L-G128H-P129Y, Y217L-G128S-P129D, Y217L-G128S-P129D-S248R, Y217L-G128S-P129G, Y217L-P129G-G131Z, Y217L-P129G-S130H-S132Z, Y217L-P129H-G131Z, Y217L-P129L, Y217L-P129S-S130H-S132Z, Y217L-P129Z, Y217L-P129Z-S130G, Y217L-P129Z-S130G-G131H-S132H, Y217L-P129Z-S130H, Y217L-S130V-G131D-S132I, G97A-G128A-Y217Q-A134T-K213L, G97A-G128A-Y217Q-G23A-S24G-N25G-P129V, G97A-G128A-Y217Q-S24R-P239R, and G97A-G128A-Y217Q-S24R-S87T-A88L-S89G, wherein positions of the variant sequence are numbered by correspondence to positions of SEQ ID NO:2.
[0258] In another aspect, the invention provides suitable cold water proteases, including subtilisin variants, particularly variants of mature BPN' (SEQ ID NO:2) comprise one, two, three, four, five, six, seven, eight, nine, ten, eleven or more of the following sets of mutations relative to SEQ ID NO:2: S87T-A88L-S89G-G97A-G128A-Y217Q, N61P-S63H-G97A-G128A-Y217Q, S87G-A88V-S89A-G97A-G128A-Y217Q, P86S-S87G-A88V-G97A-G128A-Y217Q, Q59S-N61P-G97A-G128A-Y217Q, S24G-N25G-G97A-G128A-Y217Q, N61P-N62S-G97A-G128A-Y217Q, G97A-G128A-P129Q-S130G-G131S-Y217Q, L75S-N76Y-G97A-G128A-Y217Q, G97A-G128A-V203Y-Y217Q, T55P-G97A-G128A-Y217Q, A88V-L901-G97A-G128A-Y217Q, G97A-G128A-G211R-N212S-K213V-Y217Q, G23A-S24G-N25G-G97A-G128A-Y217Q, T22N-S24A-G97A-G128A-Y217Q, S24R-G97A-G128A-Y217Q, G97A-A98S-G128A-Y217Q, G97A-G128A-T158G-S159G-Y217Q, Q59E-N61P-G97A-G128A-Y217Q, G97A-A98E-G128A-Y217Q, G97A-G128A-Y217Q-P86S-S87G-A88V-A116N-N117S-N118G, G97A-G128A-Y217Q-S63T-P86S-S87G-A88V, G97A-G128A-Y217Q-P86S-S87G-A88V-P239R, G97A-G128A-Y217Q-S24G-N25G-N61P-N62S-P194L-A232T, G97A-G128A-Y217Q-P129Q-S130G-G131S-A133V-L267V, G97A-G128A-Y217Q-A134T-L267V, G97A-G128A-Y217Q-S24R-P40E-P129E-S159K-K265R, G97A-G128A-Y217Q-A134T-G211T, G97A-G128A-Y217Q-S24R-P129E, G97A-G128A-Y217Q-I111V-S161P, G97A-G128A-Y217Q-T55P-P129Q, G97A-G128A-Y217Q-I115V-L267V, G97A-G128A-Y217Q-P86S-S87G-A88V-A116S-N117G-N118R, G97A-G128A-Y217Q-V203Y-L267V, G97A-G128A-Y217Q-S24G-N25G-S78N-S101N-V203Y, G97A-G128A-Y217Q-P525-T55P-V203Y, G97A-G128A-Y217Q-Q59S-N61P-A116S-N117G-N118R, G97A-G128A-Y217Q-S24G-N25G-P129Q-S130G-G131S, G97A-G128A-Y217Q-P86S-S87G-A88V-T242R, G97A-G128A-Y217Q-P40E-T55P-N269K, G97A-G128A-Y217Q-G23A-S24G-N25G-A116N-N117S-N118G, G97A-G128A-Y217Q-V8L-N25Y-P129Q-S130G-G131S, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-G211T-L267V, G97A-G128A-Y217Q-S24R-A116N-N117S-N118G, G97A-G128A-Y217Q-S24R-A128S-P129G, G97A-G128A-Y217Q-P129Q-S130G-G131S-N240K, G97A-G128A-Y217Q-N25Y-P129Q-S130G-G131S, G97A-G128A-Y217Q-S87T-A88L-S89G-A134T, G97A-G128A-Y217Q-P129Q-S130G-G131S-L267V, G97A-G128A-Y217Q-S87G-A88V-S89A-A116N-N117S-N118G, G97A-G128A-Y217Q-N61P-P129Q-S130G-G131S, G97A-G128A-Y217Q-N61P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-T55P-P129V-P194S, G97A-G128A-Y217Q-T55P-P129V, G97A-G128A-Y217Q-S24G-N25G-T55P-S78N-S101N, G97A-G128A-Y217Q-T55P-S78N-I115V, G97A-G128A-Y217Q-N25Y-S87G-A88V-S89A, G97A-G128A-Y217Q-A134T-N240K, G97A-G128A-Y217Q-S24R-Q59S-N61P, G97A-G128A-Y217Q-G23A-S24G-N25G-P239R, G97A-G128A-Y217Q-T55P-A116S-N117G-N118R, G97A-G128A-Y217Q-A134T-S161P, G97A-G128A-Y217Q-S24G-N25G-S53G-N61P-S101N-V203Y, G97A-G128A-Y217Q-N25Y-Q59S-N61P, G97A-G128A-Y217Q-N25Y-P129Q-S130G-G131S-A137T, G97A-G128A-Y217Q-G23A-S24G-N25G-N61P-S63H, G97A-G128A-Y217Q-T55P-N61P-S78N-S101N-V203Y, G97A-G128A-Y217Q-P129Q-N240K, G97A-G128A-Y217Q-T55P-A134T, G97A-G128A-Y217Q-N25Y-N61P-S63H, G97A-G128A-Y217Q-S87T-A88L-S89G-P129S, G97A-G128A-Y217Q-T55P-L75H-N76G, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-T55P-I115V, G97A-G128A-Y217Q-T55P-A116N-N117S-N118G, G97A-G128A-Y217Q-S24G-N25G-A116N-N117S-N118G, G97A-G128A-Y217Q-S24R-P129Q-S130G-G131S, G97A-G128A-Y217Q-G23A-S24G-N25G-G211R-N212S-K213V, G97A-G128A-Y217Q-S24G-N25G-T55P-N61P-S78N-S101N-V203Y, G97A-G128A-Y217Q-T55P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-I115V-A273S, G97A-G128A-Y217Q-N25Y-T55P, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N-V203Y- , G97A-G128A-Y217Q-Q59S-N61P-N240K, G97A-G128A-Y217Q-S161P-L267V, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-S24G-N25G-N61P-S101N, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S101N-V203Y, G97A-G128A-Y217Q-N240K, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-T55P-S101N, G97A-G128A-Y217Q-N61P-S63H-A128S-P129Q, G97A-G128A-Y217Q-S89Y-P129Q-S130G-G131S, G97A-G128A-Y217Q-P129Q-S130G-G131-V203Y, G97A-G128A-Y217Q-I115V-N240K, G97A-G128A-Y217Q-S53G-N61P-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S161P-V203Y, G97A-G128A-Y217Q-S87T-A88L-S89G-N240K, G97A-G128A-Y217Q-S87T-A88L-S89G-P239R, G97A-G128A-Y217Q-T55P-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-I115V-A134T, G97A-G128A-Y217Q-Y6Q-P129Q-S130G-G131S, G97A-G128A-Y217Q-T55P-S78N-S89Y, G97A-G128A-Y217Q-S24G-N25G-T55P-S78N-A88V-S101N, G97A-G128A-Y217Q-N61P-S63H-S78N-I111y-A134T, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S78N-S101N, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S101N-V203Y, G97A-G128A-Y217Q-S53G-N61P-S101N-V203Y, G97A-G128A-Y217Q-S53G-T55P-S78N-S101N-V203Y, G97A-G128A-Y217Q-S53G-T55P-N61P-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-N240K-A273S, G97A-G128A-Y217Q-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-Q59S-N61P-S87T-A88L-S89G, G97A-G128A-Y217Q-N61P-S63H-S78N-S161P, G97A-G128A-Y217Q-N61P-S63H-S78N-I111V, G97A-G128A-Y217Q-T55P-A128S-P129Q, G97A-G128A-Y217Q-N61P-S78N-S101N-V203Y, G97A-G128A-Y217Q-N61E-A144K, G97A-G128A-Y217Q-A134T-P239R, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S101N-V203Y, G97A-G128A-Y217Q-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-T55P-N240K, G97A-G128A-Y217Q-S24G-N25G-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-P129Q-S130G-G131S-P239R, G97A-G128S-Y217Q, G97A-G128A-Y217Q-S53G-N61P-S101N, G97A-G128A-Y217Q-I111V-P129Q-G211T, G97A-G128A-Y217Q-S24G-N25G-S53G-S101N-V203Y, G97A-G128A-Y217Q-Q59S-N61P-S87G-A88V-S89A, G97A-G128A-Y217Q-S24G-N25G-S78N-S87T-A88L-S89G-S101N, G97A-G128A-Y217Q-P129Q-S130G-G131S-S162K, G97A-G128A-Y217Q-T55P-P129Q-S130G-G131S, G97A-G128A-Y217Q-T55P-V203Y, G97A-G128A-Y217Q-S87G-A88V-S89A-P129Q-S130G-G131S, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-N61P-S78N-S87T-A88L-S89G, G97A-G128A-Y217Q-I111V-P129Q-S130G-G131S, G97A-G128A-Y217Q-I111V-A273S, G97A-G128A-Y217Q-N61P-S87T-A88L-S89G, G97A-G128A-Y217Q-T22N-S24A-N61P-S63H, G97A-G128A-Y217Q, G97A-G128A-Y217Q-S53G-S78N-S87T-A88L-S89G-S101N-P129S-V203Y, G97A-G128A-Y217Q-S159K-L267V, G97A-G128A-Y217Q-P40E-S53Y-S78Y-P86S-S87G-A88V, G97A-G128A-Y217Q-S24R-S145D, G97A-G128A-Y217Q-I111V-S159K, G97A-G128A-Y217Q-T55P-P129L, G97A-G128A-Y217Q-Q59S-N61P-V203Y, G97A-G128A-Y217Q-T55P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S101N, G97A-G128A-Y217Q-S53G-N61P-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S89Y, G97A-G128A-Y217Q-S24R-P129V, G97A-G128A-Y217Q-S87G-A88V-S89A-A116N-N117S-N118G-P172H, G97A-G128A-Y217Q-I111V-A134T, G97A-G128A-Y217Q-Q59S-N61P-P129Q-S130G-G131S, G97A-G128A-Y217Q-P5S-S87G-A88V-S89A-A116G-N117R, Y217Q-N61P-A97G-G102A-A128G-P129S, G97A-G128A-Y217Q-S24G-N25G-S53G-N61P-S78N, G97A-G128A-Y217Q-S145D-S159K-N240K-Q275E, G97A-G128A-Y217Q-T55P-A128S-P129D, G97A-G128A-Y217Q-G23A-S24G-N25G-A128S-P129D, G97A-G128A-Y217Q-S24G-N25G-S53G-S78N-S87T-A88L-S89G-V203Y, G97A-G128A-Y217Q-I111V-P239R, G97A-G128A-Y217Q-S87G-A88V-S89A-S162K, G97A-G128A-Y217Q-S87T-A88L-S89G-I115V, G97A-G128A-Y217Q-S24G-N25G-T55P-S78N, G97A-G128A-Y217Q-T55P-A92G, G97A-G128A-Y217Q-S24G-N25G-S53G-S87T-A88L-S89G-V203Y, G97A-G128A-Y217Q-T22N-S24A-T55P, G97A-G128A-Y217Q-S53G-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-S53G-T55P-S78N-S87T-A88L-S89G, G97A-G128A-Y217Q-P129Q-S130G-G131S-S159K, G97A-Y217Q-N61P-N62Q-G100N-A128G, G97A-G128A-Y217Q-S24R-S78N-S182P-L267V, G97A-G128A-Y217Q-P239R-A273S, G97A-G128A-Y217Q-S53G-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-P129Q-S130G-G131S-T242R, G97A-G128A-Y217Q-S3F-S87T-A88L-S89G-G211T, G97A-G128A-Y217Q-S24G-N25G-L75H-N76G, G97A-G128A-Y217Q-S53G-T55P-N61P-S78N-S87T-A88L-S89G, G97A-G128A-Y217Q-S87T-A88L-S89G-A144K, G97A-G128A-Y217Q-S78N-S87T-A88L-S89G-V203Y, G97A-G128A-Y217Q-Q59S-N61P-A116N-N117S-N118G, G97A-G128A-Y217Q-S87T-A88L-S89G-I111V, G97A-G128A-Y217Q-S24R-S145D-P239R-Q275E, G97A-G128A-Y217Q-S145D-A273S, G97A-G128A-Y217Q-S24G-N25G-K141E-T242R, G97A-G128A-Y217Q-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-A116N-N117S-N118G-P129Q-S130G-G131S, G97A-G128A-Y217Q-S89Y-G211T, G97A-G128A-Y217Q-S87G-A88V-S89A-A116N-N117S-N118G-A144T, G97A-G128A-Y217Q-S24G-N25G-S78N-S87T-A88L-S89G-S101N-V203Y, G97A-G128A-Y217Q-S24G-N25G-P129V, G97A-Y217Q-N61P-A128G-P129S-S130P, G97A-G128A-Y217Q-T55P-N61P-S87T-A88L-S89G-G110C-S130P, G97A-Y217Q-N123G-A128G, G97A-G128A-Y217Q-N61P-N62Q-G100N-G102A-M124I, S78N-G97A-G128A-Y217Q, G97A-S101N-G128A-Y217Q, G97A-G128A-A137V-Y217Q, N61P-G97A-G128A-Y217Q, G97A-G128A-S130P-Y217Q, G97A-Q103N-G128A-Y217Q, S63T-G97A-G128A-Y217Q, G97A-G102A-G128A-Y217Q, G97A-N109D-G128A-Y217Q-S248R, S87R-G97A-G128A-Y217Q, G97A-G128A-S188D-Y217Q, S87D-G97A-G128A-Y217Q-S248R, G97A-G128A-S188D-S248R-Y217Q, G97A-G128A-S248D-Y217Q, S78N-G97A-G128A-Y217Q-L267V, S78N-G97A-G128A-Y217Q-S161P, S78N-G97A-G128A-Y217Q-I115V, S78N-G97A-G128A-Y217Q-A273S, S78N-G97A-G128A-Y217Q-G211T, S78N-G97A-G128A-Y217Q, S78N-G97A-G128A-Y217Q-I111V, S78N-G97A-G128A-Y217Q-V147L, S78N-G97A-G128A-Y217Q-I108V, S78N-G97A-G128A-Y217Q-S89Y, and S78N-G97A-G128A-Y217Q-A138T, wherein positions of the variant sequence are numbered by correspondence to positions of SEQ ID NO:2.
[0259] In another aspect, the cold water protease is variant of subtilisin BPN' having SEQ ID NO:2, wherein the variant comprises three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen, or more mutations selected from amino acid positions 24, 25, 40, 52, 53, 55, 58, 59, 61, 62, 63, 68, 78, 86, 87, 88, 89, 92, 96, 97, 100, 101, 103, 104, 106, 111, 114, 115, 116, 117, 118, 123, 124, 125, 126, 128, 129, 130, 131, 132, 133, 134, 144, 145, 159, 161, 162, 167, 194, 203, 206, 213, 217, 227, 232, 239, 240, 242, 265, 267, and 275, wherein the positions of the variant sequence are numbered by correspondence to positions in the amino acid sequence of SEQ ID NO:2.
[0260] In another aspect, the cold water protease is variant of subtilisin BPN' comprising SEQ ID NO:2 (i.e., a BPN' subtilisin variant), wherein the variant comprises a total of three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen or more mutations selected from groups (a) and (b): (a) charged mutations selected from the group consisting of N61E, A144K, P129E, P239R, P40E, Q103E, Q206E, Q7275E, S145D, S159K, S162K, S24R, S63H, S87D and T242R; and (b) neutral mutations selected from the group consisting of A114G, A116N, A133V, A134T, A232T, A88V, A92G, F58G, G100T, G128A, G131S, G97A, I111y, I115V, K213L, K265N, L126A, L267V, L96T, M124V, N117S, N118G, N123G, N240K, N25G, N61P, N62Q, N62R, N62S, P129Q, P129V, P194L, P239V, P52L, P86S, Q59S, S101N, S125A, S130G, S132N, S161P, S24G, S53G, S78N, S87G, S89Y, T55P, V203Y, V227T, V68A, W106F, Y104N, Y167A, and Y217Q, wherein the positions of the variant sequence are numbered by correspondence to positions in the amino acid sequence of SEQ ID NO:2.
Nucleic Acids of the Invention
[0261] The invention provides isolated, non-naturally occurring, or recombinant nucleic acids (also referred to herein as "polynucleotides"), which may be collectively referred to as "nucleic acids of the invention" or "polynucleotides of the invention", which encode polypeptides (e.g., protease variants) of the invention. Nucleic acids of the invention, including all described below, are useful in recombinant production (e.g., expression) of polypeptides of the invention, typically through expression of a plasmid expression vector comprising a sequence encoding the polypeptide of interest or fragment thereof. As discussed above, polypeptides include protease variant polypeptides, including subtilisin variant polypeptides having enzymatic activity (e.g., proteolytic activity) which are useful in cleaning applications and cleaning compositions for cleaning an item or a surface (e.g., surface of an item) in need of cleaning.
[0262] The invention includes an isolated, recombinant, substantially pure, or non-naturally occurring nucleic acid comprising a polynucleotide sequence encoding any protein, polypeptide, or protease variant (including any fusion protein, etc.) of the invention described above in the section entitled "Polypeptides of the Invention" and elsewhere herein, including, but not limited, to, the Examples, including Part I Examples and Part II Examples, or a complementary polynucleotide sequence thereof. The invention also provides an isolated, recombinant, substantially pure, or non-naturally-occurring nucleic acid comprising a polynucleotide sequence encoding a combination of two or more of any polypeptides, proteins, or protease variants (including any fusion protein) of the invention described above and elsewhere herein, including, but not limited, to, the Examples, including Part I Examples and Part II Examples.
[0263] In a tenth aspect, the invention provides an isolated, non-naturally occurring, or recombinant nucleic acid comprising a polynucleotide sequence encoding a variant (or polypeptide as in the third aspect) of the first, second, third, fourth, fifth, sixth, seventh, eighth, or ninth aspect of the invention, or a complementary polynucleotide sequence thereof.
[0264] In an eleventh aspect, the invention provides an isolated, non-naturally-occurring, or recombinant nucleic acid comprising a polynucleotide sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to the polynucleotide sequence set forth in SEQ ID NO:3 or SEQ ID NO:5, or a complementary polynucleotide sequence thereof.
[0265] The present invention further provides nucleic acid encoding a protease variant comprising an amino acid sequence which differs from the amino acid sequence of SEQ ID NO:2, and wherein the total net charge of the protease variant is +1, +2, +3, +4, +5, 0, -1, -2, -3, -4, or -5 relative to the total net charge of the B. amyloliquefaciens BPN' subtilisin protease, and wherein amino acid positions of the protease variant are numbered according to the numbering of corresponding amino acid positions in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' shown in SEQ ID NO:2 as determined by alignment of the protease variant amino acid sequence with the Bacillus amyloliquefaciens subtilisin BPN' amino acid sequence.
[0266] The present invention provides nucleic acids encoding a subtilisin variant of B. amyloliquefaciens BPN' subtilisin protease, wherein the BPN' subtilisin protease comprises the amino acid sequence shown in SEQ ID NO:2, and wherein the protease variant comprises an amino acid sequence which differs from the amino acid sequence of SEQ ID NO:2 in no more than two, three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, or 25 mutations at amino acid positions selected from amino acid positions 1-275, wherein the amino acid positions of the subtilisin variant are numbered by correspondence with the amino acid sequence of B. amyloliquefaciens subtilisin BPN' set forth as SEQ ID NO:2, wherein amino acid positions of the protease variant are numbered according to the numbering of corresponding amino acid positions in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' shown in SEQ ID NO:2 as determined by alignment with the protease variant.
[0267] The present invention provides nucleic acids encoding a subtilisin variant comprising an amino acid sequence which differs from the amino acid sequence of SEQ ID NO:2, and wherein the total net charge of the protease variant is +1, +2, +3, +4, +5, 0, -1, -2, -3, -4, or -5 relative to the total net charge of the Bacillus lentus subtilisin BPN' protease, and wherein amino acid positions of the protease variant are numbered according to the numbering of corresponding amino acid positions in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' shown in SEQ ID NO:2 as determined by alignment of the protease variant amino acid sequence with the Bacillus amyloliquefaciens subtilisin BPN' amino acid sequence.
[0268] The above low ionic strength protease variants may form part of a detergent composition that is diluted in water, typically within a laundry washing machine, to form a laundry detergent wash liquor, whose conductivity is from about 0.1 mS/cm to about 3 mS/cm, from about 0.3 mS/cm to about 2.5 mS/cm, or even from about 0.5 mS/cm to about 2 mS/cm. The protease variants may be high ionic strength protease variants. Such high ionic strength protease variants comprise two or more mutations, and have a total net charge of +5, +4, +3, +2, +1 or 0 relative to wild-type BPN' subtilisin protease wild-type shown in SEQ ID NO:2.
[0269] The above high ionic strength protease variants may form part of a detergent composition that is diluted in water, typically within a laundry washing machine, to form a laundry detergent wash liquor, whose conductivity is from about 3 mS/cm to about 30 mS/cm, from about 3.5 mS/cm to about 20 mS/cm, or even from about 4 mS/cm to about 10 mS/cm.
[0270] The charge of the protease variants is expressed relative to BPN' having the amino acid sequence of SEQ ID NO:2. The amino acids that impart a single negative charge are D and E and those that impart a single positive charge are R, H and K. Any amino acid change versus SEQ ID NO:2 that changes a charge is used to calculate the charge of the protease variant. For example, introducing a negative charge mutation from a wild-type neutral position will add a net charge of -1 to the protease variant, whereas introducing a negative charge mutation (D or E) from a wild-type positive amino acid residue (R, H or K) will add a net charge of -2. Summing the charge changes from all the amino acid residues that are different for the protease variant versus BPN' having the amino acid sequence of SEQ ID NO:2 gives the charge change of the protease variant. Without wishing to be bound by theory, it is believed that: the preferred charge range for cold water proteases to be used in low conductivity laundry detergent solutions is -5, -4, -3, -2, -1, 0, particularly -2, -1; the preferred charge range for cold water proteases to be used in high conductivity laundry detergent solutions is +5, +4, +3, +2, +1, 0, particularly +2, +1. By correctly selecting the charge unexpectedly improved levels of cold water cleaning performance can be obtained. "Low conductivity laundry detergent solutions" are defined as having a conductivity of from about 0.1 mS/cm to about 3 mS/cm, from about 0.3 mS/cm to about 2.5 mS/cm, or even from about 0.5 mS/cm to about 2 mS/cm. "High conductivity laundry detergent solutions" are defined as having a conductivity of from about 3 mS/cm to about 30 mS/cm, from about 3.5 mS/cm to about 20 mS/cm, or even from about 4 mS/cm to about 10 mS/cm. It is intended that the above examples be non-limiting. Once mutations are combined to optimize cold water performance, the enzyme charge can also be balanced by mutations in further positions.
[0271] The invention includes cold water proteases and products comprising at least one cold water protease. In one aspect, the cold water protease is a variant of subtilisin BPN' having SEQ ID NO:2, said variant comprising an amino acid sequence having one or more mutations, and having a total net charge of -1, 0 or +1 relative to wild-type BPN' (SEQ ID NO:2). The amino acids that impart negative charge are typically D and E and those that impart positive charge are typically R, H and K. Without wishing to be bound by theory, it is believed that this charge range (-1, 0, +1) is the optimal charge to deliver cold water performance. However, these examples should be viewed as non-limiting. In one aspect, once mutations are combined to optimize cold water performance, the enzyme charge is balanced by mutations in further positions.
[0272] The invention provides an isolated, recombinant, substantially pure, or non-naturally occurring protease variant (e.g., subtilisin variant) having proteolytic activity, said protease variant comprising an amino acid sequence which differs from the amino acid sequence shown in SEQ ID NO:2 by 50, 45, 40, 35, 30, 25, 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, 10, 9, or 8 amino acid residues, wherein amino acid positions are numbered according to the numbering of corresponding amino acid positions in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' shown in SEQ ID NO:2, as determined by alignment of the protease variant amino acid sequence with the Bacillus amyloliquefaciens subtilisin BPN' amino acid sequence, wherein the subtilisin variant includes the amino acid sequence of BPN'-v3 (SEQ ID NO:4) or BPN'-v36 (SEQ ID NO:6).
[0273] In one aspect, the invention provides an isolated, recombinant, substantially pure, or non-naturally occurring protease variant (e.g., subtilisin variant) having proteolytic activity, said protease variant comprising an amino acid sequence which differs from the amino acid sequence shown in SEQ ID NO:2 by no more than 50, 45, 40, 35, 30, 25, 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, 10, 9, 6, 5, 4, 3, 2 amino acid residues, wherein amino acid positions are numbered according to the numbering of corresponding amino acid positions in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' shown in SEQ ID NO:2, as determined by alignment of the protease variant amino acid sequence with the Bacillus amyloliquefaciens subtilisin BPN' amino acid sequence, as set forth herein.
[0274] The invention includes an isolated, recombinant, or non-naturally occurring nucleic acid comprising a polynucleotide sequence encoding: (a) at least one protease variant of the invention, including any protease variant described herein, including, but not limited to, those protease variants set forth in the Part I Examples, (b) at least one cold water protease of the invention, or (c) at least one protease variant of the invention, including any protease variant described herein, including, but not limited to, a Series I GG36 variant set forth in the Part II Examples, or a complementary polynucleotide sequence of any thereof. In another aspect, the invention provides an expression vector comprising at least one nucleic acid of the invention. The expression vector may be operably linked to a promoter.
[0275] In another aspect, the invention provides a nucleic acid comprising a polynucleotide sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to the polynucleotide sequence set forth in SEQ ID NO:3 or SEQ ID NO:5, or a complementary polynucleotide sequence thereof.
[0276] Nucleic acids of the invention can be generated by using any suitable synthesis, manipulation, and/or isolation techniques, or combinations thereof. For example, a polynucleotide of the invention may be produced using standard nucleic acid synthesis techniques, such as solid-phase synthesis techniques that are well-known to those skilled in the art. In such techniques, fragments of up to 50 or more nucleotide bases are typically synthesized, then joined (e.g., by enzymatic or chemical ligation methods, or polymerase mediated recombination methods) to form essentially any desired continuous nucleic acid sequence. The synthesis of the nucleic acids of the invention can be also facilitated (or alternatively accomplished) by any suitable method known in the art, including but not limited to chemical synthesis using the classical phosphoramidite method (see, e.g., Beaucage et al. Tetrahedron Letters 22:1859-69 (1981)); or the method described by Matthes et al., EMBO J. 3:801-805 (1984), as is typically practiced in automated synthetic methods. Nucleic acids of the invention also can be produced by using an automatic DNA synthesizer. Customized nucleic acids can be ordered from a variety of commercial sources (e.g., The Midland Certified Reagent Company, the Great American Gene Company, Operon Technologies Inc., and DNA2.0). Other techniques for synthesizing nucleic acids and related principles are known in the art (see, e.g., Itakura et al., Ann. Rev. Biochem. 53:323 (1984); and Itakura et al., Science 198:1056 (1984)).
[0277] As indicated above, recombinant DNA techniques useful in modification of nucleic acids are well known in the art. For example, techniques such as restriction endonuclease digestion, ligation, reverse transcription and cDNA production, and polymerase chain reaction (e.g., PCR) are known and readily employed by those of skill in the art. Nucleotides of the invention may also be obtained by screening cDNA libraries (e.g., cDNA libraries generated using mutagenesis techniques commonly used in the art, including those described herein) using one or more oligonucleotide probes that can hybridize to or PCR-amplify polynucleotides which encode a protease variant polypeptide(s) of the invention. Procedures for screening and isolating cDNA clones and PCR amplification procedures are well known to those of skill in the art and described in standard references known to those skilled in the art. Some nucleic acids of the invention can be obtained by altering a naturally occurring polynucleotide backbone (e.g., that encodes an enzyme or parent protease) by, for example, a known mutagenesis procedure (e.g., site-directed mutagenesis, site saturation mutagenesis, and in vitro recombination).
[0278] The invention includes nucleic acids that hybridize to a target nucleic acid of the invention (e.g., SEQ ID NO:3 or 5), or a complementary polynucleotide sequence thereof, wherein hybridization is over substantially the entire length of the target nucleic acid. The hybridizing nucleic acid may hybridize to a nucleotide sequence of the invention under at least stringent conditions or under at least high stringency conditions. Moderately stringent, stringent, and highly stringent hybridization conditions for nucleic acid hybridization experiments are known. Examples of factors that can be combined to achieve such levels of stringency are briefly discussed herein.
[0279] Nucleic acids hybridize due to a variety of well-characterized physico-chemical forces, such as hydrogen bonding, solvent exclusion, base stacking and the like. An extensive guide to the hybridization of nucleic acids is found in P. Tijssen (1993) Laboratory Techniques in Biochemistry and Molecular Biology--Hybridization with Nucleic Acid Probes, vol. 24, part I, chapter 2, "Overview of principles of hybridization and the strategy of nucleic acid probe assays," (Elsevier, N.Y.) (hereinafter "Tijssen"). See also Hames and Higgins (1995) Gene Probes 1 and 2. An indication that two nucleic acid sequences are substantially identical is that the two molecules hybridize to each other under at least stringent conditions. Stringent hybridization conditions in the context of nucleic acid hybridization experiments, such as Southern and northern hybridizations, are sequence dependent, and are different under different environmental parameters. High stringency conditions are typically selected such that hybridization occurs at about 5.degree. C. or less than the thermal melting point (Tm) for the specific sequence at a defined ionic strength and pH. The Tm is the temperature (under defined ionic strength and pH) at which 50% of the test sequence hybridizes to a perfectly matched probe. The Tm indicates the temperature at which the nucleic acid duplex is 50% denatured under the given conditions and its represents a direct measure of the stability of the nucleic acid hybrid. Thus, the Tm corresponds to the temperature corresponding to the midpoint in transition from helix to random coil; it depends on length, nucleotide composition, and ionic strength for long stretches of nucleotides. Under stringent conditions, a probe will typically hybridize to its target subsequence, but to no other sequences. Very stringent condition" are selected to be equal to the Tm for a particular probe.
[0280] The Tm of a DNA-DNA duplex can be estimated using equation (1): Tm (.degree. C.)=81.5.degree. C.+16.6 (log.sub.10 M)+0.41 (% G+C)-0.72 (% 0-500/n, where M is the molarity of the monovalent cations (usually Na+), (% G+C) is the percentage of guanosine (G) and cytosine (C) nucleotides, (% 0 is the percentage of formalize and n is the number of nucleotide bases (i.e., length) of the hybrid. The Tm of an RNA-DNA duplex can be estimated using equation (2): Tm (.degree. C.)=79.8.degree. C.+18.5 (log.sub.10 M)+0.58 (% G+C)-11.8(% G+C).sup.2-0.56 (% 0-820/n, where M is the molarity of the monovalent cations (usually Na+), (% G+C) is the percentage of guanosine (G) and cytosine (C) nucleotides, (% f) is the percentage of formamide and n is the number of nucleotide bases (i.e., length) of the hybrid. Equations 1 and 2 above are typically accurate only for hybrid duplexes longer than about 100-200 nucleotides. The Tm of nucleic acid sequences shorter than 50 nucleotides can be calculated as follows: Tm (.degree. C.)=4G+C)+2(A+T), where A (adenine), C, T (thymine), and G are the numbers of the corresponding nucleotides.
[0281] Non-hybridized nucleic acid material is typically removed by a series of washes, the stringency of which can be adjusted depending upon the desired results, in conducting hybridization analysis. Low stringency washing conditions (e.g., using higher salt and lower temperature) increase sensitivity, but can product nonspecific hybridization signals and high background signals. Higher stringency conditions (e.g., using lower salt and higher temperature that is closer to the hybridization temperature) lower the background signal, typically with only the specific signal remaining. For additional guidance, see Hames and Higgins, supra.
[0282] Exemplary stringent conditions for analysis of at least two nucleic acids comprising at least 100 nucleotides include incubation in a solution or on a filter in a Southern or northern blot comprises 50% formalin (or formamide) with 1 milligram (mg) of heparin at 42.degree. C., with the hybridization being carried out overnight. A regular stringency wash can be carried out using a solution comprising 0.2.times.SSC buffer wash at about 65.degree. C. for about 15 minutes (see Sambrook et al. (1989) Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Press, and the third edition thereof (2001) for a description of SSC buffer). The regular stringency wash can be preceded by a low stringency wash to remove background probe signal. A low stringency wash can be carried out in, for example, a solution comprising 2.times.SSC buffer at about 40.degree. C. for about 15 minutes. A highly stringent wash can be carried out using a solution comprising 0.15 M NaCl at about 72.degree. C. for about 15 minutes. Exemplary moderate stringency conditions include overnight incubation at 37.degree. C. in a solution comprising 20% formalin (or formamide), 0.5.times.SSC, 50 mM sodium phosphate (pH 7.6), 5.times.Denhardt's solution, 10% dextran sulfate, and 20 mg/ml denatured sheared salmon sperm DNA, followed by washing the filters in 1.times.SSC at about 37-50.degree. C. High stringency conditions are conditions that (a) use low ionic strength and high temperature for washing, such as 0.015 M sodium chloride/0.0015 M sodium citrate/0.1% sodium dodecyl sulfate (SDS) at 50.degree. C., (b) employ a denaturing agent during hybridization, such as formamide, e.g., 50% (v/v) formamide with 0.1% bovine serum albumin (BSA)/0.1% Ficoll/0.1% polyvinylpyrrolidone (PVP)/50 mM sodium phosphate buffer at pH 6.5 with 750 mM sodium chloride, 75 mM sodium citrate at 42.degree. C., or (c) employ 50% formamide, 5.times.SSC (0.75 M NaCl, 0.075 M sodium citrate), 50 mM sodium phosphate (pH 6.8), 0.1% sodium pyrophosphate, 5.times.Denhardt's solution, sonicated salmon sperm DNA (50m/ml), 0.1% SDS, and 10% dextran sulfate at 42.degree. C. with washes at (1) 42.degree. C. in 0.2.times.SSC, (2) 55.degree. C. in 50% formamide, and (3) 55.degree. C. in 0.1.times.SSC (preferably with EDTA). A signal to noise ratio of 2.times. or 2.5.times.-5.times. that observed for an unrelated probe in the particular hybridization assay indicates detection of a specific hybridization. Detection of at least stringent hybridization between two sequences in the context of the present invention indicates relatively strong structural similarity or homology to a nucleic acid of the invention.
Vectors, Cells, and Methods for Making Protease Variant Polypeptides of the Invention
[0283] A variety of methods are known in the art that are suitable for generating modified polynucleotides of the invention that encode protease variants of the invention (such as cold water proteases of the invention), including, but not limited to, e.g., site-saturation mutagenesis, scanning mutagenesis, insertional mutagenesis, deletion mutagenesis, random mutagenesis, site-directed mutagenesis, and directed-evolution, as well as various other recombinatorial approaches. Methods for making modified polynucleotides and proteins (e.g., protease variants) include DNA shuffling methodologies (see, e.g., Stemmer W P, Proc. Natl. Acad. Sci. USA 91(22):10747-51 (1994)); methods based on non-homologous recombination of genes, e.g., ITCHY (Ostermeier et al., Bioorg. Med. Chem. 7:2139-44 [1999]); SCRATCHY (Lutz et al., Proc. Natl. Acad. Sci. USA 98:11248-53 [2001]); SHIPREC (Sieber et al., Nat. Biotechnol. 19:456-60 [2001]); NRR (Bittker et al., Nat. Biotechnol. 20:1024-9 [2001]; Bittker et al., Proc Natl. Acad. Sci. USA 101:7011-6 [2004]); methods that rely on the use of oligonucleotides to insert random and targeted mutations, deletions and/or insertions (Ness et al., Nat. Biotechnol. 20:1251-5 [2002]; Coco et al., Nat. Biotechnol. 20:1246-50 [2002]; Zha et al., Chembiochem. 4:34-9 [2003]; Glaser et al., J. Immunol. 149:3903-13 [1992]); see also Arkin and Youvan, Biotechnology 10:297-300 (1992); Reidhaar-Olson et al., Methods Enzymol. 208:564-86 (1991).
[0284] In one aspect, a full-length parent polynucleotide is ligated into an appropriate expression plasmid, and the following mutagenesis method is used to facilitate the construction of the modified protease of the present invention, although other methods may be used. The method is based on that described by Pisarchik et al. (Pisarchik et al., Prot. Eng. Des. Select. 20:257-265 [2007]). In one aspect, an added advantage is provided in that the restriction enzyme cuts outside its recognition sequence, which allows digestion of practically any nucleotide sequence and precludes formation of a restriction site scar.
[0285] In one approach, a naturally-occurring gene encoding a full-length protease is obtained and sequenced and scanned for one or more points at which it is desired to make a mutation (e.g., deletion, insertion, substitution) at one or more amino acids. Mutation of the gene in order to change its sequence to conform to the desired sequence is accomplished by primer extension in accord with generally known methods. Fragments to the left and to the right of the desired point(s) of mutation are amplified by PCR and to include the Eam1104I restriction site. The left and right fragments are digested with Eam1104I to generate a plurality of fragments having complementary three base overhangs, which are then pooled and ligated to generate a library of modified sequences containing one or more mutations. This method avoids the occurrence of frame-shift mutations. This method also simplifies the mutagenesis process because all of the oligonucleotides can be synthesized so as to have the same restriction site, and no synthetic linkers are necessary to create the restriction sites as is required by some other methods.
[0286] In one aspect, the invention includes a method of making a cold water protease, the method comprising: (a) providing a library of nucleic acid variants of a DNA substrate molecule that encodes a parent protease; (b) transforming the library of nucleic acid variants into host cells; (c) expressing the library to provide polypeptide expression products; (d) screening the polypeptide expression products to identify a mutant protease having at least one property selected from the group consisting of: (i) having a performance index of from 1.1 to 10 on a blood/milk/ink (BMI) stain at pH 8 and 16.degree. C. compared to PURAFECT.RTM. Prime (SEQ ID NO:2 with the amino acid substitution Y217L) as defined in the "Test Method" set forth in Part I Example 1; (ii) a performance index of from 1.3 to 10 on BMI stain at pH 8 and 16.degree. C. compared to BPN' (SEQ ID NO:2), as defined in the "Test Method" set forth herein in Part I Example 1; (iii) a performance index of from 0.9 to about 10 on BMI stain at pH 8 and 16.degree. C. compared to BPN'-v3 (SEQ ID NO:4), as defined in the "Test Method" set forth herein in Part I Example 1; and (d) a performance index of from 1.0 to about 10 on BMI at pH 8 and 16.degree. C. compared to BPN'-v36 (SEQ ID NO:6), as defined in the "Test Method" set forth herein in Part I Example 1, wherein a cold water protease is identified.
[0287] In another aspect, the invention provides an expression vector comprising at least one nucleic acid of the invention. Such nucleic acid may comprise a polynucleotide sequence encoding: (a) at least one protease variant of the invention, including any protease variant described herein, including, but not limited to, those protease variants set forth in the Part I Examples, (b) at least one cold water protease of the invention, or (c) at least one protease variant of the invention, including any protease variant described herein, including, but not limited to, a Series I GG36 variant set forth in the Part II Examples, or a complementary polynucleotide sequence of any thereof. Such expression vector may be operably linked to a promoter.
[0288] In another aspect, the invention provides a recombinant host cell comprising: (a) a nucleic acid of the invention, (b) an expression vector comprising a nucleic acid of the invention, (c) a protease variant of the invention, including, but not limited to, e.g., a cold water protease of the invention. The recombinant host cell may be a bacterial cell, such as, but not limited to, e.g., a Bacillus cell. An exemplary Bacillus cell is a Bacillus subtilis cell.
[0289] In another aspect, the invention provides a cell culture comprising: (a) a nucleic acid of the invention, (b) an expression vector comprising a nucleic acid of the invention, (c) a protease variant of the invention, including, but not limited to, e.g., a cold water protease of the invention variant.
[0290] In another aspect, the invention provides isolated or recombinant vectors comprising at least one polynucleotide of the invention described herein (e.g., a polynucleotide encoding a protease variant of the invention described herein), isolated or recombinant expression vectors or expression cassettes comprising at least one nucleic acid or polynucleotide of the invention, isolated, substantially pure, or recombinant DNA constructs comprising at least one nucleic acid or polynucleotide of the invention, isolated or recombinant cells comprising at least one polynucleotide of the invention, cell cultures comprising cells comprising at least one polynucleotide of the invention, cell cultures comprising at least one nucleic acid or polynucleotide of the invention, and compositions comprising one or more such vectors, nucleic acids, expression vectors, expression cassettes, DNA constructs, cells, cell cultures, or any combination or mixtures thereof.
[0291] In another aspect, the invention provides recombinant cells comprising at least one vector (e.g., expression vector or DNA construct) of the invention which comprises at least one nucleic acid or polynucleotide of the invention. Some such recombinant cells are transformed or transfected with such at least one vector. Such cells are typically referred to as host cells. Some such cells comprise bacterial cells, including, but are not limited to, e.g., Bacillus sp. cells, such as, e.g., Bacillus subtilis cells. The invention also provides recombinant cells (e.g., recombinant host cells) comprising at least one protease variant of the invention.
[0292] In another aspect, the invention provides a vector comprising a nucleic acid or polynucleotide of the invention. The vector may be an expression vector or expression cassette in which a polynucleotide sequence of the invention which encodes a protease variant of the invention is operably linked to one or additional nucleic acid segments required for efficient gene expression (e.g., a promoter operably linked to the polynucleotide of the invention which encodes a protease variant of the invention). A vector may include a transcription terminator and/or a selection gene, such as an antibiotic resistance gene that enables continuous cultural maintenance of plasmid-infected host cells by growth in antimicrobial-containing media.
[0293] An expression vector may be derived from plasmid or viral DNA, or in alternative aspects, contains elements of both. Exemplary vectors include, but are not limited to pXX, pC194, pJH101, pE194, pHP13 (Harwood and Cutting (eds.)), Molecular Biological Methods for Bacillus, John Wiley & Sons (1990), see, e.g., chapter 3; suitable replicating plasmids for B. subtilis include those listed on p. 92; Perego, M. (1993) Integrational Vectors for Genetic Manipulations in Bacillus subtilis, pp. 615-624; A. L. Sonenshein, J. A. Hoch, and R. Losick (ed.), Bacillus subtilis and other Gram-positive bacteria: biochemistry, physiology and molecular genetics, American Society for Microbiology, Washington, D.C.
[0294] For expression and production of a protein of interest (e.g., protease variant) in a cell, at least one expression vector comprising at least one copy of a polynucleotide encoding the modified protease, and preferably comprising multiple copies, is transformed into the cell under conditions suitable for expression of the protease. In one aspect, a polynucleotide sequence encoding the protease variant (as well as other sequences included in the vector) is integrated into the genome of the host cell, while in another aspect, a plasmid vector comprising a polynucleotide sequence encoding the protease variant remains as autonomous extra-chromosomal element within the cell. The invention provides both extrachromosomal nucleic acid elements as well as incoming nucleotide sequences that are integrated into the host cell genome. The vectors described herein are useful for production of the protease variants of the invention. In one aspect, a polynucleotide construct encoding the protease variant is present on an integrating vector that enables the integration and optionally the amplification of the polynucleotide encoding the protease variant into the bacterial chromosome. Examples of sites for integration include are well known to those skilled in the art. In one aspect, transcription of a polynucleotide encoding a protease variant of the invention is effectuated by a promoter that is the wild-type promoter for the selected precursor protease. In some other aspects, the promoter is heterologous to the precursor protease, but is functional in the host cell. Specifically, examples of suitable promoters for use in bacterial host cells include, but are not limited to, e.g., the amyE, amyQ, amyL, pstS, sacB, pSPAC, pAprE, pVeg, pHpaII promoters, the promoter of the B. stearothermophilus maltogenic amylase gene, the B. amyloliquefaciens (BAN) amylase gene, the B. subtilis alkaline protease gene, the B. clausii alkaline protease gene the B. pumilis xylosidase gene, the B. thuringiensis cryIIIA, and the B. licheniformis alpha-amylase gene. Additional promoters include, but are not limited to the A4 promoter, as well as phage Lambda P.sub.R or P.sub.L promoters, and the E. coli lac, trp or tac promoters.
[0295] Protease variants of the invention can be produced in host cells of any suitable Gram-positive microorganism, including bacteria and fungi. For example, in one aspect, the protease variant is produced in host cells of fungal and/or bacterial origin. In one aspect, the host cells are Bacillus sp., Streptomyces sp., Escherichia sp. or Aspergillus sp. In one aspect, the protease variants are produced by Bacillus sp. host cells. Examples of Bacillus sp. host cells that find use in the production of the protease variants of the invention include, but are not limited to B. licheniformis, B. lentus, B. subtilis, B. amyloliquefaciens, B. lentus, B. brevis, B. stearothermophilus, B. alkalophilus, B. coagulans, B. circulars, B. pumilis, B. thuringiensis, B. clausii, and B. megaterium, as well as other organisms within the genus Bacillus. In one aspect, B. subtilis host cells are used for production of protease variants. U.S. Pat. Nos. 5,264,366 and 4,760,025 (RE 34,606) describe various Bacillus host strains that can be used for producing protease variants of the invention, although other suitable strains can be used.
[0296] Several industrial bacterial strains that can be used to produce protease variants of the invention include non-recombinant (i.e., wild-type) Bacillus sp. strains, as well as variants of naturally-occurring strains and/or recombinant strains. In one aspect, the host strain is a recombinant strain, wherein a polynucleotide encoding a polypeptide of interest has been introduced into the host. In one aspect, the host strain is a B. subtilis host strain and particularly a recombinant Bacillus subtilis host strain. Numerous B. subtilis strains are known, including, but not limited to, e.g., 1A6 (ATCC 39085), 168 (1A01), SB19, W23, Ts85, B637, PB1753 through PB1758, PB3360, JH642, 1A243 (ATCC 39,087), ATCC 21332, ATCC 6051, MI113, DE100 (ATCC 39,094), GX493I, PBT 110, and PEP 211strain (see, e.g., Hoch et al., Genetics 73:215-228 (1973)) (see also U.S. Pat. Nos. 4,450,235 and 4,302,544, and EP 0134048, each of which is incorporated by reference in its entirety). The use of B. subtilis as an expression host cells is well known in the art (see, e.g., Palva et al., Gene 19:81-87 (1982); Fahnestock and Fischer, J. Bacteriol. 165:796-804 (1986); and Wang et al., Gene 69:39-47 (1988)).
[0297] In one aspect, the Bacillus host cell is a Bacillus sp. that includes a mutation or deletion in at least one of the following genes, degU, degS, degR and degQ. The mutation may be in a degU gene, and in some instances the mutation may be degU(Hy)32. See, e.g., Msadek et al., J. Bacteriol. 172:824-834 (1990) and Olmos et al., Mol. Gen. Genet. 253:562-567 (1997)). A typical host strain is a Bacillus subtilis carrying a degU32(Hy) mutation. The Bacillus host may comprise an amino acid mutation (e.g., substitution) or deletion in scoC4 (see, e.g., Caldwell et al., J. Bacteriol. 183:7329-7340 (2001)); spoIIE (see, e.g., Arigoni et al., Mol. Microbiol. 31:1407-1415 (1999)); and/or oppA or other genes of the opp operon (see, e.g., Perego et al., Mol. Microbiol. 5:173-185 (1991)). Indeed, it is contemplated that any mutation in the opp operon that causes the same phenotype as a mutation in the oppA gene will find use in one aspect of the altered Bacillus strain of the invention. Such mutations may occur alone or combinations of mutations may be present. In one aspect, an altered Bacillus host cell strain that can be used to produce a protease variant of the invention is a Bacillus host strain that already includes a mutation in one or more of the above-mentioned genes. In addition, Bacillus sp. host cells that comprise mutation(s) and/or deletions of endogenous protease genes find use. The Bacillus host cell may comprise a deletion of the aprE and the nprE genes. The Bacillus sp. host cell may comprise a deletion of 5 protease genes, or the Bacillus sp. host cell may comprise a deletion of 9 protease genes (see, e.g., U.S. Pat. Appn. Pub. No. 2005/0202535).
[0298] Host cells are transformed with at least one nucleic acid encoding at least one protease variant of the invention using any suitable method known in the art. Whether the nucleic acid is incorporated into a vector or is used without the presence of plasmid DNA, it is typically introduced into a microorganism, in one aspect, preferably an E. coli cell or a competent Bacillus cell. Methods for introducing a nucleic acid (e.g., DNA) into Bacillus cells or E. coli cells utilizing plasmid DNA constructs or vectors and transforming such plasmid DNA constructs or vectors into such cells are well known. In one aspect, the plasmids are subsequently isolated from E. coli cells and transformed into Bacillus cells. However, it is not essential to use intervening microorganisms such as E. coli, and in one aspect, a DNA construct or vector is directly introduced into a Bacillus host.
[0299] Those of skill in the art are well aware of suitable methods for introducing nucleic acid or polynucleotide sequences of the invention into Bacillus cells (see, e.g., Ferrari et al., "Genetics," in Harwood et al. (ed.), Bacillus, Plenum Publishing Corp. (1989), pp. 57-72; Saunders et al., J. Bacteriol. 157:718-726 (1984); Hoch et al., J. Bacteriol. 93:1925-1937 (1967); Mann et al., Current Microbiol. 13:131-135 (1986); and Holubova, Folia Microbiol. 30:97 (1985); Chang et al., Mol. Gen. Genet. 168:11-115 (1979); Vorobjeva et al., FEMS Microbiol. Lett. 7:261-263 (1980); Smith et al., Appl. Env. Microbiol. 51:634 (1986); Fisher et al., Arch. Microbiol. 139:213-217 (1981); and McDonald, J. Gen. Microbiol. 130:203 (1984)). Indeed, such methods as transformation, including protoplast transformation and congression, transduction, and protoplast fusion are well known and suited for use in the present invention. Methods of transformation are used to introduce a DNA construct or vector comprising a nucleic acid encoding a protease variant of the present invention into a host cell. Methods known in the art to transform Bacillus cells include such methods as plasmid marker rescue transformation, which involves the uptake of a donor plasmid by competent cells carrying a partially homologous resident plasmid (Contente et al., Plasmid 2:555-571 (1979); Haima et al., Mol. Gen. Genet. 223:185-191 (1990); Weinrauch et al., J. Bacteriol. 154:1077-1087 (1983); and Weinrauch et al., J. Bacteriol. 169:1205-1211 (1987)). In this method, the incoming donor plasmid recombines with the homologous region of the resident "helper" plasmid in a process that mimics chromosomal transformation.
[0300] In addition to commonly used methods, in one aspect, host cells are directly transformed with a DNA construct or vector comprising a nucleic acid encoding a protease variant of the invention (i.e., an intermediate cell is not used to amplify, or otherwise process, the DNA construct or vector prior to introduction into the host cell). Introduction of the DNA construct or vector of the invention into the host cell includes those physical and chemical methods known in the art to introduce a nucleic acid sequence (e.g., DNA sequence) into a host cell without insertion into a plasmid or vector. Such methods include, but are not limited to calcium chloride precipitation, electroporation, naked DNA, liposomes and the like. In additional aspects, DNA constructs or vector are co-transformed with a plasmid, without being inserted into the plasmid. In further aspects, a selective marker is deleted from the altered Bacillus strain by methods known in the art (see Stahl et al., J. Bacteriol. 158:411-418 (1984); and Palmeros et al., Gene 247:255-264 (2000)).
[0301] In one aspect, the transformed cells of the present invention are cultured in conventional nutrient media. The suitable specific culture conditions, such as temperature, pH and the like are known to those skilled in the art. In addition, some culture conditions may be found in the scientific literature such as Hopwood (2000) Practical Streptomyces Genetics, John Innes Foundation, Norwich U K; Hardwood et al., (1990) Molecular Biological Methods for Bacillus, John Wiley and from the American Type Culture Collection (ATCC). In one aspect, the invention provides a culture (e.g., cell culture) comprising at least one protease variant or at least one nucleic acid of the invention. Also provided is a composition comprising at least one nucleic acid, vector, or DNA construct of the invention.
[0302] Host cells transformed with at least one polynucleotide sequence encoding at least one protease variant of the invention may be cultured in a suitable nutrient medium under conditions permitting the expression of the present protease, after which the resulting protease is recovered from the culture. The medium used to culture the cells comprises any conventional medium suitable for growing the host cells, such as minimal or complex media containing appropriate supplements. Suitable media are available from commercial suppliers or may be prepared according to published recipes (e.g., in catalogues of the American Type Culture Collection). The protease produced by the cells may be recovered from the culture medium by conventional procedures, including, but not limited to, e.g., separating the host cells from the medium by centrifugation or filtration, precipitating the proteinaceous components of the supernatant or filtrate by means of a salt (e.g., ammonium sulfate), chromatographic purification (e.g., ion exchange, gel filtration, affinity, etc.). Any method suitable for recovering or purifying a protease variant of the invention can be used.
[0303] In one aspect, a protease variant produced by a recombinant host cell is secreted into the culture medium. A nucleic acid sequence that encodes a purification facilitating domain may be used to facilitate purification of soluble proteins. A vector or DNA construct comprising a polynucleotide sequence encoding a protease variant may further comprise a nucleic acid sequence encoding a purification facilitating domain to facilitate purification of the protease variant (see, e.g., Kroll, D. J. et al., DNA Cell Biol. 12:441-53 (1993)). Such purification facilitating domains include, but are not limited to, e.g., metal chelating peptides such as histidine-tryptophan modules that allow purification on immobilized metals (Porath J., Protein Expr. Purif 3:263-281 (1992)), protein A domains that allow purification on immobilized immunoglobulin, and the domain utilized in the FLAGS extension/affinity purification system (Immunex Corp., Seattle, Wash.). The inclusion of a cleavable linker sequence such as Factor XA or enterokinase (Invitrogen, San Diego, Calif.) between the purification domain and the heterologous protein also find use to facilitate purification.
[0304] Assays for detecting and measuring the enzymatic activity of an enzyme, such as a protease variant of the invention, are well known. Various assays for detecting and measuring activity of proteases, such as, e.g., protease variants of the invention, are also known to those of ordinary skill in the art. In particular, assays are available for measuring protease activity that are based on the release of acid-soluble peptides from casein or hemoglobin, measured as absorbance at 280 nm or colorimetrically using the Folin method (see, e.g., Bergmeyer et al., "Methods of Enzymatic Analysis" vol. 5, Peptidases, Proteinases and their Inhibitors, Verlag Chemie, Weinheim (1984)). Other exemplary assays involve the solubilization of chromogenic substrates (see, e.g., Ward, "Proteinases," in Fogarty (ed.). Microbial Enzymes and Biotechnology, Applied Science, London, (1983), pp. 251-317). Other exemplary assays include, but are not limited to succinyl-Ala-Ala-Pro-Phe-para nitroanilide assay (SAAPFpNA) and the 2,4,6-trinitrobenzene sulfonate sodium salt assay (TNBS assay). Numerous additional references known to those in the art provide suitable methods (see, e.g., Wells et al., Nucleic Acids Res. 11:7911-7925 (1983); Christianson et al., Anal. Biochem. 223:119-129 (1994); and Hsia et al., Anal Biochem. 242:221-227 (1999)).
[0305] A variety of methods can be used to determine the level of production of a mature protease (e.g., mature protease variant of the invention) in a host cell. Such methods include, but are not limited to, e.g., methods that utilize either polyclonal or monoclonal antibodies specific for the protease. Exemplary methods include, but are not limited, e.g., to enzyme-linked immunosorbent assays (ELISA), radioimmunoassays (RIA), fluorescent immunoassays (FIA), and fluorescent activated cell sorting (FACS). These and other assays are well known in the art (see, e.g., Maddox et al., J. Exp. Med. 158:1211 (1983)).
[0306] In another aspect, the invention provides methods for making or producing a mature protease variant of the invention. A mature protease variant does not include a signal peptide or a propeptide sequence. Some such methods comprising making or producing a protease variant of the invention in a recombinant bacterial host cell, such as, e.g., a Bacillus sp. cell, including, e.g., Bacillus subtilis cell. In one aspect, the invention provides a method of producing a protease variant of the invention, the method comprising cultivating a recombinant host cell comprising a recombinant expression vector comprising a nucleic acid encoding a protease variant of the invention under conditions conducive to the production of the protease variant. Some such methods further comprise recovering the protease variant from the culture.
[0307] In one aspect, the invention provides a method of producing a protease variant of the invention, the method comprising: (a) introducing a recombinant expression vector comprising a nucleic acid encoding a protease variant of the invention into a population of cells (e.g., bacterial cells, such as Bacillus subtilis cells); and (b) culturing the cells in a culture medium under conditions conducive to produce the protease variant encoded by the expression vector. Some such methods further comprise: (c) isolating the protease variant from the cells or from the culture medium.
[0308] In addition to recombinant production, the protease variant polypeptides of the invention may be produced by direct peptide synthesis using solid-phase techniques (see, e.g., Stewart et al. (1969) Solid-Phase Peptide Synthesis, W.H. Freeman Co, San Francisco; Merrifield (1963) J. Am. Chem. Soc 85:2149-2154). Peptide synthesis may be performed using manual or automated techniques. Automated synthesis may be achieved, for example, using Applied Biosystems 431A Peptide Synthesizer (Perkin Elmer, Foster City, Calif.) in accordance with the instructions provided by the manufacturer. For example, subsequences may be chemically synthesized separately and combined using chemical methods to provide protease variants or functional fragments thereof. Alternatively, such variant polypeptide sequences may be ordered from any number of companies that specialize in production of polypeptides. Most commonly, polypeptides of the invention are produced by expressing coding nucleic acids and recovering polypeptides, e.g., as described above and in the Examples.
[0309] In another aspect, the invention provides a method of producing a protease variant, the method comprising cultivating a recombinant host cell of the invention, said host cell comprising an expression vector which comprising at least one nucleic acid of the invention, under conditions conducive to produce the variant. The method may further comprise recovering the variant from the culture.
[0310] In another aspect, the invention includes a method of producing a protease variant, the method comprising: (a) introducing the recombinant expression vector of the invention into a population of cells; and (b) culturing the cells in a culture medium under conditions conducive to produce the subtilisin variant encoded by the expression vector. The method may further comprise (c) isolating the variant from the cells or from the culture medium.
[0311] In another aspect, the invention provides methods for producing a serine protease variant of a Bacillus serine protease, comprising: transforming a host cell with an expression vector comprising a nucleic acid encoding the serine protease variant; and cultivating the transformed host cell under conditions suitable for the production of the serine protease variant. In one aspect, the methods further comprise the step of harvesting the produced serine protease variant. In one aspect, the host cell is a Bacillus cell, and in a subset of these aspects, the Bacillus cell is a B. subtilis cell. Furthermore, the present invention provides methods of cleaning, comprising the step of contacting a surface and/or an article comprising a fabric with a cleaning composition comprising a serine protease variant. In some alternative methods, the present invention provides methods of cleaning, comprising the step of contacting a surface and/or an article comprising dishware with a cleaning composition comprising a serine protease variant.
[0312] In another aspect, the invention provides a method for identifying a wild-type subtilisin protease, said subtilisin protease comprising an amino acid sequence which comprises the following amino acids: a glycine or arginine at amino acid position 24, a glycine at amino acid position 53, an asparagine at amino acid position 78, an alanine at amino acid position 97, an asparagine at amino acid position 101, an alanine or serine at amino acid position 128, and/or a leucine or glutamine at amino acid position 217, wherein said amino acid positions are numbered by correspondence with amino acid positions in the amino acid sequence of BPN' set forth in SEQ ID NO:2, said method comprising: a) providing a sample comprising at least one wild-type subtilisin protease or a library of wild-type subtilisin proteases, or providing a DNA sample comprising one or more DNA sequences encoding wild-type subtilisin proteases; and b) identifying a wild-type subtilisin protease or DNA sequence that encodes a wild-type protease.
[0313] In a twelfth aspect, the invention provides an expression vector comprising at least one nucleic acid of the tenth or eleventh aspect of the invention. An expression vector according to the twelfth aspect of the invention may be operably linked to a promoter. Such expression vector may be in isolated or purified form.
[0314] In a thirteenth aspect, the invention provides a recombinant host cell comprising: (a) a nucleic acid of the tenth or eleventh aspect of the invention or (b) an expression vector of the twelfth aspect of the invention. A recombinant host cell of the thirteenth aspect of the invention may be a bacterial cell, which may optionally be a Bacillus cell. A recombinant host cell of the thirteenth aspect of the invention may be a Bacillus subtilis cell.
[0315] In a fourteenth aspect, the invention provides a cell culture comprising: (a) a nucleic acid of the tenth or eleventh aspect of the invention or (b) an expression vector of the twelfth aspect of the invention.
[0316] In a fifteenth aspect, the invention provides a method of producing a protease variant, the method comprising cultivating a recombinant host cell of the thirteenth aspect of the invention under conditions conducive to produce the variant. A method according to the fifteenth aspect of the invention may further comprise isolating or recovering the variant from the culture.
[0317] In a sixteenth aspect, the invention provides a method of producing a protease variant, the method comprising: (a) introducing the recombinant expression vector of the twelfth aspect of the invention into a population of cells; and (b) culturing the cells in a culture medium under conditions conducive to produce the protease variant encoded by the expression vector. The method according to the sixteenth aspect of the invention may further comprise (c) isolating or recovering the variant from the cells or from the culture medium.
Compositions of the Invention
[0318] The invention includes a composition comprising at least one polypeptide (e.g., at least one protease variant) of the invention. Polypeptides of the invention are described infra and supra, including in the Part I Examples and Part II Examples. Such compositions may comprise at least one excipient, carrier, adjunct ingredient, or other substituent, component, or material.
[0319] For example, in one aspect, the invention includes a composition comprising at least one protease variant and at least one excipient, carrier, adjunct ingredient, or other substituent, component, or material. Such at least one protease variant may be any one of those set forth herein, including, but not limited to, in the Part I Examples. Such at least one protease variant may be a BPN' protease variant. For example, a composition of the invention may be a BPN' protease variant, such as BPN'-v36 (SEQ ID NO:6) or BPN'-v3 (SEQ ID NO:4) or any protease variant set forth herein or in Part I Examples 2-23. Such composition may be a fabric and home care product or fabric and home care composition. Alternatively, such composition may be a fabric and home care product or fabric and home care composition.
[0320] Such composition may further comprise (in addition to a BPN' protease variant) at least one GG36 protease variant, such as at least one Series I GG36 protease variant described in the Part II Examples. Such compositions are useful in a variety of applications as described elsewhere herein.
[0321] In a seventeenth aspect, the invention provides a composition comprising a variant (or polypeptide as in the third aspect) of the first, second, third, fourth, fifth, sixth, seventh, eighth, or ninth aspect of the invention. A composition according to the seventeenth aspect of the invention may comprise an adjunct ingredient or carrier as described in greater detail elsewhere herein. A composition according to the seventeenth aspect of the invention may comprise at least one builder and/or at least one surfactant. A composition according to the seventeenth aspect of the invention may comprise phosphate or may not comprise phosphate.
[0322] A composition according to the seventeenth aspect of the invention may be a cleaning composition or a detergent composition. A composition according to the seventeenth aspect of the invention may be a fabric or home care product. A composition according to the seventeenth aspect of the invention may not be a fabric or home care product. A composition according to the seventeenth aspect of the invention may be a cleaning composition or detergent composition that is a fabric or home care product. A composition according to the seventeenth aspect of the invention may be a cleaning composition or detergent composition that is not a fabric or home care product.
[0323] A composition according to the seventeenth aspect of the invention may comprise may further comprise one, two, three, four, five or more additional enzymes. Such additional enzyme(s) may be selected from the group consisting of additional enzymes selected from the group consisting of hemicellulase, cellulase, amylase, peroxidase, protease, xylanase, lipase, phospholipase, esterase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, B-glucanase, arabinosidase, hyaluronidase, chondroitinase, and laccase.
[0324] The additional enzyme according to the seventeenth aspect of the invention may be a GG36 protease variant having proteolytic activity, such as, e.g., a Series I GG36 protease variant, which Series I GG36 protease variant may comprise an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to the subtilisin Bacillus lentus GG36 of SEQ ID NO:755 and at least one amino acid substitution selected from the group consisting of A1R, Q2S, V4R, V4S, S9A, R10S, P14K, A16S, H17R, N18R, G20R, T22A, T22R, S24R, S24W, G25R, G25V, V26F, L421, N43R, N43A, G46R, P52F, P52E, P52N, T57R, Q59A, N62E, N62Q, V68A, V68C, T71G, I72C, A74C, L75A, L75F, L75R, N76D, 578R, L82R, P86W, E89P, E89T, E89G, E89H, E89I, E89V, E89W, Y91N, K94N, G100S, S101A, S101N, S101G, S101D, S103G, S103N, V104L, V104I, S106V, S106G, A108I, L111V, E112V, G115K, G115R, N117F, G118I, V121F, S128D, S128F, S128L, S128N, P129E, S144R, L148I, A158E. G159E, S160D, S166D, N185E, N185I, R186H, S188E, S188D, D197F, V203E, Y209S, Y209N, Y209F, Y209T, Y209E, Y209H, Y209G, P210R, S212I, S212F, Y214F, A215N, A215D, A215E, L217E, L217N, T224A, A230E, A231I, Q236F, N238R, N238K, P239K, P239G, P239R, P239S, W241R, S242R, S242L, N243R, V244R, N248I, N248V, H249R, L250I, N252R, T253R, L262D, Y263F, S265F, L267V, L267N. N269I, N269R, E271F, E271I, E271H, E271P, E271T, E271V, E271L, and A272F, wherein each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' protease set forth in SEQ ID NO:2. A composition according to the seventeenth aspect of the invention that comprises an additional enzyme (such as a protease, such as, e.g., a Series I G36 protease variant as described herein) may be a fabric and home care product. Alternatively, such composition may not be a fabric and home care product.
[0325] A composition according to the seventeenth aspect of the invention may comprise a Series I GG36 protease variant which comprises an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the subtilisin Bacillus lentus GG36 of SEQ ID NO:755 and at least one amino acid substitution selected from the group consisting of T022R-S024R, S009A-E271L, N018R-W241R, N018R-G115R, N043R-H249R, G020R-H249R, V004R-H249R, G020R-S024R, N018R-H249R, S009A-G020R, G020R-W241R, S009A-S078R, G020R-G115R, N018R-S024R, S024R-S242R, T022R-G115R, N018R-N043R, G020R-N043R, N018R-S242R, S242R-N269R, N018R-V244R, S024R-N269R, G020R-E271L, S024R-E271L, V004R-S009A, G020R-N269R, A001R-S024R, V244R-E271L, S009A-N018R, W241R-E271L, V004R-S024R, S009A-H249R, S009A-T022R, N062E-P129E, N062E-G159E, A016S-L148I, A158E-H249R, A016S-N062E, L111V-S188D, T022A-N062E, N062E-L148I, T022A-P129E, N062E-E271F, N062E-A158E, A016S-G159E, N062E-R186H, S128N-G159E, N062E-S188D, N062E-S128N, L148I-G159E, S103G-A158E, L111V-G159E, A158E-E271F, A016S-S188D, T022A-L111V, S128N-A158E, A016S-A158E, V104L-A158E, S128N-R186H, G159E-Y209E, N062E-S101A, L111V-Y209E, L148I-S188D, S101A-Y209E, T022A-S188D, A016S-T022A, S128N-P129E, A016S-Y209E, A016S-S128N, T022A-E089P, S128N-Y209E, E089P-A158E, N062E-S103G, R186H-E271F, A016S-P129E, E089P-G159E, L111V-H249R, S101A-P129E, L148I-Y209E, T022A-G159E, P129E-H249R, P129E-Y209E, V104L-P129E, S128N-S188D, L111V-A158E, T022A-A158E, N062E-Y209E, N062E-H249R, S101A-R186H, E089P-P129E, P129E-E271, T22A-L111V-G159E, S101A-S103G-V104L-Y209E, S101A-S103G-V104L-G159E, S101A-S103G-V104L-S188D, S101G-S103A-V104I-G159D, T22A-S103G-G159E, T22A-S128N-E271F-Y209E, T22A-Y209E-E271F, T22A-S101A-Y209E, S101A-Y209E-E271F, T22A-L111V-S128N, T22A-S101A-G159E, S101A-S103G-V104L, T22A-S101A-S103G-V104L, S101A-S103G-V104L, S101G-S103A-V104I, S101A-S103G-V104L-S128N, S103A-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N248D, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-S24R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-T253R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159E-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N238R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N76D, and S101G-S103A-V104I-G159E-A232V-Q245R-N248D-E271F, wherein each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' protease set forth in SEQ ID NO:2. Such composition may be a fabric and home care product or fabric and home care composition. In another aspect, such composition may not be a fabric and home care product or fabric and home care composition.
[0326] A composition according to the seventeenth aspect of the invention may comprise an Series I GG36 protease variant which comprises an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to the subtilisin Bacillus lentus GG36 of SEQ ID NO:755 and comprises three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, or even 25 amino acid substitutions selected from the group consisting of: A1R, Q2S, V4R, V4S, S9A, R10S, P14K, A16S, H17R, N18R, G20R, T22A, T22R, S24R, S24W, G25R, G25V, V26F, L421, N43R, N43A, G46R, P52F, P52E, P52N, T57R, Q59A, N62E, N62Q, V68A, V68C, T71G, I72C, A74C, L75A, L75F, L75R, N76D, 578R, L82R, P86W, E89P, E89T, E89G, E89H, E89I, E89V, E89W, Y91N, K94N, G100S, S101A, S101N, S101G, S101D, S103G, S103N, V104L, V104I, S106V, S106G, A108I, L111V, E112V, G115K, G115R, N117F, G118I, V121F, S128D, S128F, S128L, S128N, P129E, S144R, L148I, A158E. G159E, S160D, S166D, N185E, N185I, R186H, S188E, S188D, D197F, V203E, Y209S, Y209N, Y209F, Y209T, Y209E, Y209H, Y209G, P210R, S212I, S212F, Y214F, A215N, A215D, A215E, L217E, L217N, T224A, A230E, A231I, Q236F, N238R, N238K, P239K, P239G, P239R, P239S, W241R, S242R, S242L, N243R, V244R, N248I, N248V, H249R, L250I, N252R, T253R, L262D, Y263F, S265F, L267V, L267N, N269I, N269R, E271F, E271I, E271H, E271P, E271T, E271V, E271L, and A272F; and optionally at least one amino acid substitution selected from the group consisting of: S103A, G159D, Q236H, Q245R, N248D, and N252K, wherein each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' protease set forth in SEQ ID NO:2. Such composition may be a fabric and home care product or fabric and home care composition. In another aspect, such composition may not be a fabric and home care product or fabric and home care composition.
[0327] A composition according to the seventeenth aspect of the invention may comprise a Series I GG36 protease variant which comprises an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to the subtilisin Bacillus lentus GG36 of SEQ ID NO:755 and (a) two or more substitutions selected from the group consisting of A1R, Q2S, V4R, V4S, S9A, R10S, P14K, A16S, T22A, T22R, S24R, G25V, V26F, L421, P52F, P52E, P52N, N62E, N62Q, V68A, V68C, T71G, I72C, A74C, L75A, L75F, 578R, E89P, E89T, E89G, E89H, E89W, Y91N, K94N, G100S, S101A, S101N, S101G, S101D, S103G, S103N, V104L, V104I, A108I, L111V, E112V, G115K, N117F, V121F, S128D, S128F, S128L, S128N, P129E, L148I, A158E. G159E, S160D, S166D, N185E, R186H, S188E, S188D, V203E, Y209S, Y209N, Y209F, Y209T, Y209E, Y209H, Y209G, P210R, S212I, S212F, Y214F, A215N, A215D, A215E, L217E, L217N, T224A, A230E, A231I, Q236F, N238R, N238K, P239K, P239G, P239R, N248V, H249R, L250I, L262D, Y263F, S265F, L267V, L267N. N269I, N269R, E271F, E271I, E271H and A272F, and/or (b) one or more sets of substitutions selected from the group consisting of N062E-P129E, N062E-G159E, A016S-L148I, A158E-H249R, A016S-N062E, L111V-S188D, T022A-N062E, N062E-L148I, T022A-P129E, N062E-E271F, N062E-A158E, A016S-G159E, N062E-R186H, S128N-G159E, N062E-S188D, N062E-S128N, L148I-G159E, S103G-A158E, L111V-G159E, A158E-E271F, A016S-S188D, T022A-L111V, S128N-A158E, A016S-A158E, V104L-A158E, S128N-R186H, G159E-Y209E, N062E-S101A, L111V-Y209E, L148I-S188D, S101A-Y209E, T022A-S188D, A016S-T022A, S128N-P129E, A016S-Y209E, A016S-S128N, T022A-E089P, S128N-Y209E, E089P-A158E, N062E-S103G, R186H-E271F, A016S-P129E, E089P-G159E, L111V-H249R, S101A-P129E, L148I-Y209E, T022A-G159E, P129E-H249R, P129E-Y209E, V104L-P129E, S128N-S188D, L111V-A158E, T022A-A158E, N062E-Y209E, N062E-H249R, S101A-R186H, E089P-P129E, P129E-E271F, T22A-L111V-G159E, S101A-S103G-V104L-Y209E, S101A-S103G-V104L-G159E, S101A-S103G-V104L-S188D, S101G-S103A-V104I-G159D, T22A-S103G-G159E, T22A-S128N-E271F-Y209E, T22A-Y209E-E271F, T22A-S101A-Y209E, S101A-Y209E-E271F, T22A-L111V-S128N, T22A-S101A-G159E, S101A-S103G-V104L, T22A-S101A-S103G-V104L, S101A-S103G-V104L, S101G-S103A-V104I, S101A-S103G-V104L-S128N, S103A-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N248D, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-S24R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-T253R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159E-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N238R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N76D, and S101G-S103A-V104I-G159E-A232V-Q245R-N248D-E271F, wherein each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' protease set forth in SEQ ID NO:2. Such composition may be a fabric and home care product or fabric and home care composition. In another aspect, such composition may not be a fabric and home care product or fabric and home care composition.
[0328] A composition according to the seventeenth aspect of the invention may comprise a Series I GG36 protease variant which comprises an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% sequence identity to the subtilisin Bacillus lentus GG36 of SEQ ID NO:755 and (a) two or more substitutions selected from the group of consisting of V4R, H17R, N18R, G20R, T22R, S24R, S24W, G25R, N43R, N43A, G46R, P52F, P52N, T57R, Q59A, N62Q, T71G, L75R, N76D, S78R, L82R, P86W, E89P, E89W, E89T, E89I, E89H, E89V, V104L, S106V, S106G, G115R, G118I, V121F, S144R, N185I, D197F, Y209N, Y209S, L217E, A231I, P239R, P239S, W241R, S242R, S242L, N243R, V244R, N248I, H249R, N252R, T253R, E271T, E271V, E271L, E271H, E271F, E271P, A1R, S9A, S212F, and N269R; and/or (b) one or more sets of substitutions selected from the group consisting of T022R-S024R, S009A-E271L, N018R-W241R, N018R-G115R, N043R-H249R, G020R-H249R, V004R-H249R, G020R-S024R, N018R-H249R, S009A-G020R, G020R-W241R, S009A-S078R, G020R-G115R, N018R-S024R, S024R-S242R, T022R-G115R, N018R-N043R, G020R-N043R, N018R-S242R, S242R-N269R, N018R-V244R, S024R-N269R, G020R-E271L, S024R-E271L, V004R-S009A, G020R-N269R, A001R-S024R, V244R-E271L, S009A-N018R, W241R-E271L, V004R-S024R, S009A-H249R, S009A-T022R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-S24R, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K, and S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K, wherein each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' protease set forth in SEQ ID NO:2. Such composition may be a fabric and home care product or fabric and home care composition. In another aspect, such composition may not be a fabric and home care product or fabric and home care composition.
[0329] A composition according to the seventeenth aspect of the invention may comprise at least one additional enzyme selected from the group consisting of hemicellulase, cellulase, amylase, peroxidase, protease, xylanase, lipase, phospholipase, esterase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, B-glucanase, arabinosidase, hyaluronidase, chondroitinase, and laccase. A composition according to the seventeenth aspect of the invention may comprise two or more additional enzymes selected from the group consisting of hemicellulase, cellulase, amylase, peroxidase, protease, xylanase, lipase, phospholipase, esterase, cutinase, pectinase, pectate lyase, mannanase, keratinase, reductase, oxidase, phenoloxidase, lipoxygenase, ligninase, pullulanase, tannase, pentosanase, malanase, B-glucanase, arabinosidase, hyaluronidase, chondroitinase, and laccase.
[0330] A composition according to the seventeenth aspect of the invention may comprise phosphate or may not contain phosphate. A composition according to the seventeenth aspect of the invention may comprise at least one builder and/or at least one surfactant.
[0331] As further elsewhere herein, the polypeptides of the invention, including the protease variants of the invention, are useful in a variety of cleaning applications, including laundry cleaning applications, automatic dishwashing applications, hand dishwashing applications, hard surface cleaning applications, personal care applications, and other applications described herein. Thus, for example, in one aspect, the invention provides cleaning compositions comprising at least one polypeptide (e.g., protease variant) of the invention. As noted above, such cleaning compositions include, but are not limited to, e.g., automatic and hand dishwashing detergent compositions, laundry detergent compositions (including, e.g., liquid and powder laundry detergent compositions), fabric cleaning compositions, hard surface cleaning compositions (including, but not limited to, e.g., hard surface of a non-dishware item, non-tableware item, table, table top, furniture item, wall, floor, ceiling, etc.). Such cleaning compositions, which are useful in methods of cleaning an item or a surface in need of cleaning, may comprise, e.g., but not limited to, at least one excipient, carrier, and/or other substituent, component, or material.
[0332] In another aspect, the invention provides a composition comprising any polypeptide of the invention (e.g., any protease variant or subtilisin variant of the invention) described herein, wherein said composition is a fabric and home care composition or a fabric and home care product.
[0333] In another aspect, the invention provides a composition comprising any polypeptide of the invention (e.g., any protease variant or subtilisin variant of the invention) described herein, wherein said composition is not a fabric and home care composition or not a fabric and home care product. A composition of the invention comprising a protease variant of the invention may further comprise at least one adjunct material selected from perfume encapsulate; fabric hueing agent; cold-water soluble brightener; a bleach catalyst that may comprise a material selected from an iminium cation, iminium polyion, iminium zwitterion; modified amine; modified amine oxide; N-sulphonyl imine; N-phosphonyl imine; N-acyl imine; thiadiazole dioxide; perfluoroimine; cyclic sugar ketone; first wash lipase; bacterial cleaning cellulase; Guerbet nonionic surfactant; and mixture of any thereof. Compositions of the invention may further comprise at least one additional non-immunoequivalent protease selected from subtilisins (EC 3.4.21.62); trypsin-like or chymotrypsin-like proteases; metalloproteases; and mixtures thereof.
[0334] Compositions of the invention may further comprise at least one additional non-immunoequivalent protease selected from: subtilisins (EC 3.4.21.62) derived from B. subtilis, B. amyloliquefaciens, B. pumilus and B. gibsonii; trypsin proteases and/or chymotrypsin proteases derived from Cellulomonas; metalloproteases derived from B. amyloliquefaciens; and mixtures thereof.
[0335] Compositions of the invention further comprise at least one additional enzyme selected from first-wash lipases; alpha-amylases; bacterial cleaning cellulases; and mixtures thereof.
[0336] A composition of the invention may further comprise at least one of the following: an encapsulate comprising a perfume comprises a perfume micro capsule; a hueing agent comprising a material selected from basic, acid, hydrophobic, direct and polymeric dyes, and dye-conjugates having a peak absorption wavelength of from 550 nm to 650 nm and mixtures thereof; a detersive surfactant comprising a material selected from anionic detersive surfactants, non-ionic detersive surfactant, cationic detersive surfactants, zwitterionic detersive surfactants and amphoteric detersive surfactants and mixtures thereof; a builder comprising a material selected from zeolites, phosphates and mixtures thereof; a silicate salt comprising a material selected from sodium silicate, potassium silicate and mixtures thereof; a brightener comprising a material selected from cold-water soluble brightener and mixtures thereof; a carboxylate polymer comprising a material selected from maleate/acrylate random copolymer or polyacrylate homopolymer and mixtures thereof; a soil release polymer comprising a material selected from terephthalate co-polymer and mixtures thereof; a cellulosic polymer comprising a material selected from alkyl cellulose, alkyl alkoxyalkyl cellulose, carboxyalkyl cellulose, alkyl carboxyalkyl cellulose and mixtures thereof; a bleach catalyst comprising a material selected from an iminium cation; iminium polyion; iminium zwitterion; modified amine; modified amine oxide; N-sulphonyl imine; N-phosphonyl imine; N-acyl imine; thiadiazole dioxide; perfluoroimine; cyclic sugar ketone and any mixture thereof; a bleach activator comprising a material selected from dodecanoyl oxybenzene sulphonate, decanoyl oxybenzene sulphonate, decanoyl oxybenzoic acid or salt thereof, 3,5,5-trimethyl hexanoyloxybenzene sulphonate, tetraacetyl ethylene diamine (TAED), nonanoyloxybenzene sulphonate (NOBS) and mixtures thereof; a source of hydrogen peroxide comprising a material selected from an inorganic perhydrate salt, including an alkali metal salt, such as sodium salts of perborate (usually mono- or tetra-hydrate), percarbonate, persulphate, perphosphate, persilicate salt, and any mixture thereof; a chelant comprising a material selected from DTPA (diethylene triamine pentaacetic acid), HEDP (hydroxyethane diphosphonic acid), DTPMP (diethylene triamine penta(methylene phosphonic acid)), ethylenediaminedisuccinic acid (EDDS), 1,2-dihydroxybenzene-3,5-disulfonic acid disodium salt hydrate, derivative of said chelant; and any mixture thereof.
[0337] A composition of the invention may comprise a fabric hueing agent selected from the group consisting of a dye; dye-clay conjugate comprising at least one cationic-basic dye and a smectite clay; and any mixture thereof.
[0338] A composition of the invention may comprise at least one fabric hueing agent selected from small molecule dye, polymeric dye, and any mixture thereof; dye-clay conjugate comprising at least one cationic-basic dye and a smectite clay; and any mixture thereof.
[0339] A composition comprising a protease of the invention may be provided in single or multiple-compartment unit doses. The composition may be a multi-compartment unit dose, wherein the protease variant is in a different compartment than any source of hydrogen peroxide and/or chelant and/or additional enzyme. A composition comprising at least one protease variant or polypeptide of the invention may comprise a wash liquor.
[0340] A composition comprising at least one protease variant of the invention may comprise one or more of the following ingredients (based on total composition weight): from about 0.0005 wt % to about 0.1 wt %, from about 0.001 wt % to about 0.05 wt %, or even from about 0.002 wt % to about 0.03 wt % of said protease variant; and one or more of the following: from about 0.00003 wt % to about 0.1 wt % fabric hueing agent; from about 0.001 wt % to about 5 wt %, perfume capsules; from about 0.001 wt % to about 1 wt %, cold-water soluble brighteners; from about 0.00003 wt % to about 0.1 wt % bleach catalysts; from about 0.00003 wt % to about 0.1 wt % first wash lipases; from about 0.00003 wt % to about 0.1 wt % bacterial cleaning cellulases; and/or from about 0.05 wt % to about 20 wt % Guerbet nonionic surfactants.
[0341] A composition may be a granular or powder laundry detergent comprising a cold water protease or comprising a protease variant that is not a cold water protease.
[0342] A composition of the invention may be provided in any suitable form, including a fluid or solid. The composition may be in the form of a unit dose pouch, especially when in the form of a liquid, and the composition may be at least partially, or even completely, enclosed by a water-soluble pouch. In addition, the composition may have any combination of parameters and/or characteristics detailed above.
[0343] Unless otherwise noted, all component or composition levels provided herein are made in reference to the active level of that component or composition, and are exclusive of impurities, for example, residual solvents or by-products, which may be present in commercially available sources. Enzyme components weights are based on total active protein. All percentages and ratios are calculated by weight unless otherwise indicated. All percentages and ratios are calculated based on the total composition unless otherwise indicated. In the exemplified detergent compositions, the enzymes levels are expressed by pure enzyme by weight of the total composition and, unless otherwise specified, the detergent ingredients are expressed by weight of the total compositions.
[0344] As indicated herein, in one aspect, the cleaning compositions of the present invention may further comprise one or more adjunct materials or ingredients including, but not limited to, e.g., one or more surfactants, builders, bleaches, bleach activators, bleach catalysts, other enzymes, enzyme stabilizing systems, chelants, optical brighteners, soil release polymers, dye transfer agents, dispersants, suds suppressors, dyes, perfumes, perfume capsules, colorants, filler salts, hydrotropes, photoactivators, fluorescers, fabric conditioners, fabric softeners, hydrolyzable surfactants, preservatives, anti-oxidants, anti-shrinkage agents, anti-wrinkle agents, germicides, fungicides, color speckles, silvercare, anti-tarnish and/or anti-corrosion agents, alkalinity sources, solubilizing agents, carriers, excipients, processing aids, pigments, rinse aid (e.g., a rinse aid containing at least one surfactant to prevent water droplet formation by making water drain from the surface of the item being cleaned in a thin sheet, rather than forming droplets), solvents, and/or pH control agents (see, e.g., U.S. Pat. Nos. 6,610,642, 6,605,458, 5,705,464, 5,710,115, 5,698,504, 5,695,679, 5,686,014 and 5,646,101, all of which are incorporated herein by reference). Aspects of specific cleaning composition materials are exemplified in detail below. If a cleaning adjunct material(s) is not compatible with a protease variant of the present invention in a desired cleaning composition, then a suitable method of keeping the cleaning adjunct material(s) and the protease variant(s) separated (i.e., not in contact with one anther) until combination of the two components is appropriate is used. Such separation methods include any suitable method known in the art (e.g., gelcaps, encapsulation, tablets, physical separation, etc.).
[0345] The cleaning compositions of the present invention are advantageously employed, for example, in laundry applications, hard surface cleaning applications, hand or manual dishwashing applications, automatic dishwashing applications, eyeglass cleaning applications, as well as cosmetic applications, such as for cleaning dentures, teeth, hair, and skin. Due to the unique advantages of increased effectiveness in lower temperature solutions, the protease variant enzymes of the present invention are suited for laundry applications and dishwashing applications, including hand and automatic dishwashing applications. Furthermore, the protease variant enzymes of the present invention find use in solid, gel, granular, and/or liquid compositions, including solid, gel, granular, and/or liquid detergent compositions and/or formulations.
[0346] The protease variants of the present invention also find use cleaning additive product compositions. In one aspect, a protease variant of the invention is useful in low temperature solution cleaning applications and methods. In one aspect, the invention provides cleaning additive product compositions which include at least one protease variant enzyme of the present invention and which are ideally suited for inclusion in a wash process when additional bleaching effectiveness is desired. Such instances include, but are not limited to, e.g., low temperature solution cleaning applications. In one aspect, the additive product composition is in its simplest form--i.e., one or more protease variants of the invention. In one aspect, the additive product composition is packaged in dosage form for addition to a cleaning process. In one aspect, the additive product composition is packaged in dosage form for addition to a cleaning process where a source of peroxygen is employed and increased bleaching effectiveness is desired. Any suitable single dosage unit form may be used, including but not limited to, e.g., pills, tablets, gelcaps, or other single dosage units, such as pre-measured powders or liquids. Thus, in one aspect, the invention provides a cleaning product composition comprising at least one protease variant of the invention, wherein the product is formulated in suitable form (e.g., as a liquid, powder solid, pill, tablet, gelcap or other suitable form) in a suitable single dosage unit such that a single dose of the protease variant is provided. Such cleaning products are useful in a variety of cleaning methods and applications, including but not limited to, e.g., machine or hand laundry methods and applications, automatic dishwashing or hand dishwashing methods and applications, etc. Such cleaning methods and applications may be conducted at low temperature or low pH conditions. In one aspect, at least one filler and/or at least one carrier material is included to increase the volume of such compositions. Suitable filler or carrier materials include, but are not limited to, e.g., various salts of sulfate, carbonate and silicate as well as talc, clay and the like. Suitable filler or carrier materials for liquid compositions include, but are not limited to, e.g., water or low molecular weight primary and secondary alcohols including polyols and diols. Examples of such alcohols include, but are not limited to, e.g., methanol, ethanol, propanol and isopropanol. In one aspect, the compositions contain from about 5% to about 90% of such filler or carrier materials. Acidic fillers may be included in such compositions to reduce the pH of the resulting solution in the cleaning method or application. Alternatively, in one aspect, the cleaning additive includes one or more adjunct ingredients, as more fully described below.
[0347] The present cleaning compositions and cleaning additives require an effective amount of at least one protease variant of the invention, alone or in combination with other proteases and/or additional enzymes. The required level of enzyme is achieved by the addition of one or more protease variants of the invention. Typically, a cleaning composition comprises at least about 0.0001 weight percent to about 20 weight percent, from about 0.0001 to about 10 weight percent, from about 0.0001 to about 1 weight percent, from about 0.001 to about 1 weight percent, or from about 0.01 to about 0.1 weight percent of at least one protease variant of the invention. In one aspect, a composition of the invention (e.g., cleaning composition of the invention) comprises from about 0.01 milligram (mg) to about 10 mg, about 0.01 to about 5 mg, about 0.01 mg to about 2 mg, about 0.01 to about 1 mg, about 0.5 mg to about 10 mg, about 0.5 to about 5 mg, about 0.5 to about 4 mg, about 0.5 to about 4 mg, about 0.5 to about 3 mg, about 0.5 to about 2 mg, about 0.5 to about 1 mg, about 0.1 to about 10 mg, about 0.1 to about 5 mg, about 0.1 to about 4 mg, about 0.1 to about 3 mg, about 0.1 to about 2 mg, about 0.1 to about 2 mg, about 0.1 to about 1 mg, about 0.1 to about 0.5 mg of at least one active protease variant of the invention per gram of the composition.
[0348] The invention includes a cleaning composition comprising an amount of a protease variant of the invention (said composition optionally comprising one or more adjunct ingredients) such that when the cleaning composition is added to wash water the resultant protease concentration in the resultant wash liquor is 0.01 ppm to 10 ppm, including, e.g., 0.1 ppm to 1 ppm, 0.1 to 5 ppm, 1 to 5 ppm, 1 ppm to 10 ppm, 5 ppm to 10 ppm, 5 ppm to 7 ppm.
[0349] The cleaning compositions of the invention are typically formulated such that, during use in aqueous cleaning operations, the wash water will have a pH of from about 5.0 to about 11.5 or even from about 7.5 to about 10.5. Liquid product compositions or formulations are typically formulated to have a neat pH from about 3.0 to about 9.0 or from about 3.0 to about 5.0. Granular laundry product compositions are typically formulated to have a pH from about 9 to about 11. Hand dishwashing and automatic dishwashing detergent compositions are typically formulated to have a pH from about 8 to about 11.5, including, but not limited to, e.g., pH ranges of about 8 to about 10, from about 9 to about 11.5, and from about 9.5 to about 11.5 depending on the method and specific application. Techniques for controlling pH at recommended usage levels include the use of buffers, alkalis, acids, etc., and are well known to those skilled in the art.
[0350] Suitable low pH cleaning compositions typically have a neat pH of from about 3 to about 5, and are typically free of surfactants that hydrolyze in such a pH environment. Such surfactants include sodium alkyl sulfate surfactants that comprise at least one ethylene oxide moiety or even from about 1 to about 16 moles of ethylene oxide. Such cleaning compositions typically comprise a sufficient amount of a pH modifier, such as sodium hydroxide, monoethanolamine, or hydrochloric acid, to provide such cleaning composition with a neat pH of from about 3 to about 5. Such compositions typically comprise at least one acid stable enzyme. In one aspect, the compositions are liquids, while in other aspects, they are solids. The pH of such liquid compositions is typically measured as a neat pH. The pH of such solid compositions is measured as a 10% solids solution of said composition wherein the solvent is distilled water. In these aspects, all pH measurements are taken at 20.degree. C., unless otherwise indicated.
[0351] In one aspect, when the protease variant(s) of the invention is/are employed in a granular composition or liquid, it is desirable for the protease variant to be in the form of an encapsulated particle to protect the protease variant from other components of the granular composition during storage. In addition, encapsulation is also a means of controlling the availability of the protease variant during the cleaning process. In one aspect, encapsulation enhances the performance of the protease variant(s) and/or additional enzymes. In this regard, the protease variants of the present invention are encapsulated with any suitable encapsulating material known in the art. In one aspect, the encapsulating material typically encapsulates at least part of the catalyst for the protease variant(s) of the invention. Typically, the encapsulating material is water-soluble and/or water-dispersible. In one aspect, the encapsulating material has a glass transition temperature (Tg) of 0.degree. C. or higher. Glass transition temperature is described in more detail in WO 97/11151. The encapsulating material is typically selected from consisting of carbohydrates, natural or synthetic gums, chitin, chitosan, cellulose and cellulose derivatives, silicates, phosphates, borates, polyvinyl alcohol, polyethylene glycol, paraffin waxes, and combinations thereof. When the encapsulating material is a carbohydrate, it is typically selected from monosaccharides, oligosaccharides, polysaccharides, and combinations thereof. In some typical aspects, the encapsulating material is a starch (see, e.g., EP 0 922 499; U.S. Pat. Nos. 4,977,252, 5,354,559, and 5,935,826). In one aspect, the encapsulating material is a microsphere made from plastic such as thermoplastics, acrylonitrile, methacrylonitrile, polyacrylonitrile, polymethacrylonitrile and mixtures thereof; commercially available microspheres that find use include, but are not limited to those supplied by EXPANCEL.RTM. (Stockviksverken, Sweden), and PM 6545, PM 6550, PM 7220, PM 7228, EXTENDOSPHERES.RTM., LUXSIL.RTM., Q-CEL.RTM., and SPHERICEL.RTM. (PQ Corp., Valley Forge, Pa.).
[0352] As described herein, the protease variants of the invention find particular use in the cleaning methods and applications, including, but not limited to, e.g., cleaning, laundry, hand dishwashing, and automatic dishwashing detergent compositions. These applications place enzymes under various environmental stresses. The protease variants of the invention provide advantages over many currently used enzymes in such cleaning applications due to their proteolytic activity and stability under various conditions.
[0353] There are a variety of wash conditions including varying detergent formulations, wash water volumes, wash water temperatures, and lengths of wash time, to which proteases involved in washing are exposed. In addition, detergent formulations used in different geographical areas have different concentrations of their relevant components present in the wash water. For example, European detergents typically have about 4500-5000 parts per million (ppm) of detergent components in the wash water, while Japanese detergents typically have approximately 667 ppm of detergent components in the wash water. In North America, particularly the United States, detergents typically have about 975 ppm of detergent components present in the wash water.
[0354] A low detergent concentration system includes detergents where less than about 800 ppm of the detergent components are present in the wash water. Japanese detergents are typically considered low detergent concentration system as they have approximately 667 ppm of detergent components present in the wash water.
[0355] A medium detergent concentration includes detergents where between about 800 ppm and about 2000 ppm of the detergent components are present in the wash water. North American detergents are generally considered to be medium detergent concentration systems as they have approximately 975 ppm of detergent components present in the wash water. Brazil typically has approximately 1500 ppm of detergent components present in the wash water.
[0356] A high detergent concentration system includes detergents where greater than about 2000 ppm of the detergent components are present in the wash water. European detergents are generally considered to be high detergent concentration systems as they have approximately 4500-5000 ppm of detergent components in the wash water.
[0357] Latin American detergents are generally high suds phosphate builder detergents and the range of detergents used in Latin America can fall in both the medium and high detergent concentrations as they range from 1500 ppm to 6000 ppm of detergent components in the wash water. As mentioned above, Brazil typically has approximately 1500 ppm of detergent components present in the wash water. However, other high suds phosphate builder detergent geographies, not limited to other Latin American countries, may have high detergent concentration systems up to about 6000 ppm of detergent components present in the wash water.
[0358] In light of the foregoing, it is evident that concentrations of detergent compositions in typical wash solutions throughout the world varies from less than about 800 ppm of detergent composition ("low detergent concentration geographies"), e.g., about 667 ppm in Japan, to between about 800 ppm to about 2000 ppm ("medium detergent concentration geographies"), e.g., about 975 ppm in U.S. and about 1500 ppm in Brazil, to greater than about 2000 ppm ("high detergent concentration geographies"), e.g., about 4500 ppm to about 5000 ppm in Europe and about 6000 ppm in high suds phosphate builder geographies.
[0359] The concentrations of the typical wash solutions are determined empirically. For example, in the U.S., a typical washing machine holds a volume of about 64.4 L of wash solution. Accordingly, in order to obtain a concentration of about 975 ppm of detergent within the wash solution about 62.79 g of detergent composition must be added to the 64.4 L of wash solution. This amount is the typical amount measured into the wash water by the consumer using the measuring cup provided with the detergent.
[0360] As a further example, different geographies use different wash temperatures. The temperature of the wash water in Japan is typically less than that used in Europe. For example, the temperature of the wash water in North America and Japan is typically between about 10 and about 30.degree. C. (e.g., about 20.degree. C.), whereas the temperature of wash water in Europe is typically between about 30 and about 60.degree. C. (e.g., about 40.degree. C.). However, in the interest of saving energy, many consumers are switching to using cold water washing. In addition, in some further regions, cold water is typically used for laundry, as well as in dishwashing applications. In one aspect, the "cold water washing" of the present invention utilizes washing at temperatures from about 4.degree. C. to about 10.degree. C., from about 10.degree. C. to about 40.degree. C., or from about 20.degree. C. to about 30.degree. C., from about 15.degree. C. to about 25.degree. C., from about 10.degree. C. to about 20.degree. C., from about 14.degree. C. to about 18.degree. C., as well as all other combinations within the range of about 15.degree. C. to about 35.degree. C., and all ranges within 10.degree. C. to 40.degree. C., and about 16.degree. C.
[0361] As a further example, different geographies typically have different water hardness. Water hardness is usually described in terms of the grains per gallon mixed Ca.sup.2+/Mg.sup.2+. Hardness is a measure of the amount of calcium (Ca.sup.2+) and magnesium (Mg.sup.2+) in the water. Most water in the United States is hard, but the degree of hardness varies. Moderately hard (60-120 ppm) to hard (121-181 ppm) water has 60 to 181 parts per million (parts per million converted to grains per U.S. gallon is ppm # divided by 17.1 equals grains per gallon) of hardness minerals.
TABLE-US-00001 Water Grains per gallon Parts per million Soft less than 1.0 less than 17 Slightly hard 1.0 to 3.5 17 to 60 Moderately hard 3.5 to 7.0 60 to 120 Hard 7.0 to 10.5 120 to 180 Very hard greater than 10.5 greater than 180
[0362] European water hardness is typically greater than about 10.5 (e.g., about 10.5 to about 20.0) grains per gallon mixed Ca.sup.2+/Mg.sup.2+ (e.g., about 15 grains per gallon mixed Ca.sup.2+/Mg.sup.2+). North American water hardness is typically greater than Japanese water hardness, but less than European water hardness. For example, North American water hardness can be between about 3 to about 10 grains, about 3 to about 8 grains or about 6 grains. Japanese water hardness is typically lower than North American water hardness, usually less than about 4, for example about 3 grains per gallon mixed Ca.sup.2+/Mg.sup.2+.
[0363] Accordingly, in one aspect, the invention provides protease variants that show surprising wash performance in at least one set of wash conditions (e.g., water temperature, water hardness, and/or detergent concentration). In one aspect, the protease variants of the invention are comparable in wash performance to other subtilisin proteases. In one aspect, the protease variants of the present invention exhibit enhanced wash performance as compared to subtilisin proteases currently commercially available. Thus, in one aspect of the invention, the protease variants provided herein exhibit enhanced oxidative stability, enhanced thermal stability, enhanced cleaning capabilities under various conditions, and/or enhanced chelator stability. In addition, the protease variants of the invention find use in cleaning compositions that do not include detergents, again either alone or in combination with builders and stabilizers.
[0364] In one aspect, the invention provides a cleaning composition comprising at least one protease variant of the present invention that is present at a level from about 0.00001% to about 10% by weight of the composition with the balance (e.g., about 99.999% to about 90.0%) comprising one or more cleaning adjunct materials by weight of composition. In another aspect, the invention provides a cleaning composition comprising at least one protease variant of the invention that is present at a level of about 0.0001% to about 10%, about 0.001% to about 5%, about 0.001% to about 2%, or about 0.005% to about 0.5% by weight of the composition with the balance of the cleaning composition (e.g., about 99.9999% to about 90.0%, about 99.999% to about 98%, about 99.995% to about 99.5% by weight) comprising one or more cleaning adjunct materials.
[0365] In one aspect, a cleaning composition of the invention comprises, in addition to at least one protease variant of the invention, one or more additional enzymes, which provide cleaning performance and/or fabric care and/or hand or manual dishwashing and/or automatic dishwashing benefits. Examples of suitable enzymes include, but are not limited to, e.g., hemicellulases, cellulases, peroxidases, proteases, xylanases, lipases, phospholipases, esterases, perhydrolases, cutinases, pectinases, pectate lyases, mannanases, keratinases, reductases, oxidases, phenoloxidases, lipoxygenases, ligninases, pullulanases, tannases, pentosanases, malanases, -glucanases, arabinosidases, hyaluronidase, chondroitinase, laccase, and/or amylases, neutral metalloprotease enzymes (abbreviated as "nprE"), or mixtures of any thereof. In one aspect, the cleaning composition comprises, in addition to at least one protease variant of the invention, a combination of additional enzymes (i.e., a "cocktail") comprising conventional applicable enzymes such as, e.g., at least one additional protease, lipase, cutinase, cellulose, and/or amylase.
[0366] In addition to the protease variants provided herein, any other suitable protease may find use and be included in a composition of the invention. In one aspect, the invention provides a composition (e.g., cleaning composition) comprising at least one protease variant of the invention and at least one additional protease. Suitable proteases include those of animal, vegetable, or microbial origin. In one aspect, a microbial protease may be included. A chemically or genetically modified mutant of a protease may be included. In one aspect, the at least one additional protease is a serine protease, preferably an alkaline microbial protease or a trypsin-like protease. Examples of alkaline proteases include subtilisins, especially those derived from Bacillus (e.g., subtilisin, B. lentus subtilisin (i.e., GG36), B. amyloliquefaciens subtilisin (i.e., BPN'), subtilisin Carlsberg, subtilisin 309, subtilisin 147, PB92, and subtilisin 168). Additional examples include those mutant proteases (i.e., protease variants) described in U.S. Pat. Nos. RE 34,606, 5,955,340, 5,700,676, 6,312,936, and 6,482,628, all of which are incorporated herein by reference. Additional proteases include, but are not limited to trypsin (e.g., of porcine or bovine origin), and the Fusarium protease described in WO 89/06270. A composition of the invention comprising at least one protease variant of the invention may also comprise at least one commercially available protease enzyme. Commercially available protease enzymes that find use in compositions of the invention include, but are not limited to, e.g., MAXATASE.RTM., MAXACAL.TM., MAXAPEM.TM., OPTICLEAN.RTM., OPTIMASE.RTM., PROPERASE.RTM., PURAFECT.RTM., PURAFECT.RTM. OXP, PURAMAX.TM., EXCELLASE.TM., and PURAFAST.TM. (Genencor); ALCALASE.RTM., SAVINASE.RTM., PRIMASE.RTM., DURAZYM.TM., POLARZYME.RTM., OVOZYME.RTM., KANNASE.RTM., LIQUANASE.RTM., NEUTRASE.RTM., RELASE.RTM. and ESPERASE.RTM. (Novozymes); KAP Bacillus alkalophilus subtilisin with A230V+S256G+S259N (Kao); and BLAP B. lentus protease, BLAP X, and BLAP S (Henkel Kommanditgesellschaft auf Aktien, Duesseldorf, Germany). Additional proteases that may be included in compositions of the invention include those described in WO95/23221, WO 92/21760, U.S. Pat. Pub. No. 2008/0090747, and U.S. Pat. Nos. 5,801,039, 5,340,735, 5,500,364, 5,855,625, RE 34,606, 5,955,340, 5,700,676, 6,312,936, and 6,482,628, and various other patents. Metalloproteases may be included in compositions of the invention. Such metalloproteases, include, but are not limited to, e.g., the neutral metalloprotease enzyme (nprE) described in WO 07/044993.
[0367] In one aspect, the invention provides a composition (e.g., cleaning composition) comprising at least one protease variant of the invention and at least one lipase. Suitable lipases include, but are not limited to, e.g., those of bacterial or fungal origin. A chemically or genetically modified mutant of a lipase may be included in the composition. Examples of useful lipases include Humicola lanuginosa lipase (see, e.g., EP 258 068, and EP 305 216), Rhizomucor miehei lipase (see, e.g., EP 238 023), Candida lipase, such as C. antarctica lipase (e.g., C. antarctica lipase A or B; see, e.g., EP 214 761), Pseudomonas lipases such as P. alcaligenes lipase and P. pseudoalcaligenes lipase (see, e.g., EP 218 272), P. cepacia lipase (see, e.g., EP 331 376), P. stutzeri lipase (see, e.g., GB 1,372,034), P. fluorescens lipase, Bacillus lipase (e.g., B. subtilis lipase (Dartois et al., Biochem. Biophys. Acta 1131:253-260 (1993)); B. stearothermophilus lipase (see, e.g., JP 64/744992); and B. pumilus lipase (see, e.g., WO 91/16422)).
[0368] Furthermore, a number of cloned lipases find use in compositions (e.g., cleaning compositions) of the present invention, including, but not limited to, e.g., Penicillium camembertii lipase (Yamaguchi et al., Gene 103:61-67 (1991)), Geotricum candidum lipase (Schimada et al., J. Biochem. 106:383-388 (1989)), and various Rhizopus lipases such as R. delemar lipase (Hass et al., Gene 109:117-113 (1991)), a R. niveus lipase (Kugimiya et al., Biosci. Biotech. Biochem. 56:716-719 (1992)) and R. oryzae lipase.
[0369] Other types of lipolytic enzymes, such as cutinases, also find use in one aspect of the present invention, including, but not limited to, e.g., the cutinase derived from Pseudomonas mendocina (see WO 88/09367), and the cutinase derived from Fusarium solani pisi (see WO 90/09446).
[0370] Additional suitable lipases include commercially available lipases such as M1 LIPASE.TM. LUMA FAST.TM., and LIPOMAX.TM. (Genencor); LIPOLASE.RTM. and LIPOLASE.RTM. ULTRA (Novozymes); and LIPASE P.TM. "Amano" (Amano Pharmaceutical Co. Ltd., Japan).
[0371] In one aspect, the invention provides compositions (e.g., cleaning compositions) comprising at least one protease variant of the invention and at least one lipase that is present at a level from about 0.00001% to about 10% of additional lipase by weight of the composition and the balance of one or more cleaning adjunct materials by weight of composition. In one aspect, a cleaning composition of the present invention comprises, in addition to at least one protease variant of the invention, at least one lipase at a level of about 0.0001% to about 10%, about 0.001% to about 5%, about 0.001% to about 2%, or about 0.005% to about 0.5% lipase by weight of the composition.
[0372] Also included is a composition (e.g., cleaning composition) comprising at least one variant of the invention and at least one amylase. Any amylase (e.g., alpha and/or beta) suitable for use in alkaline solutions may be useful to include in such a composition. Suitable amylases include, but are not limited to, e.g., those of bacterial or fungal origin. A chemically or genetically modified mutant of an amylase may be included. Amylases that find use in compositions of the invention, include, but are not limited to, e.g., a-amylases obtained from B. licheniformis (see, e.g., GB 1,296,839). Commercially available amylases that find use in compositions of the invention include, but are not limited to, e.g., DURAMYL.RTM., TERMAMYL.RTM., FUNGAMYL.RTM., STAINZYME.RTM., STAINZYME PLUS.RTM., STAINZYME ULTRA.RTM., and BAN.TM. (Novozymes), as well as POWERASE.TM., RAPIDASE.RTM. and MAXAMYL.RTM. P (Genencor).
[0373] In one aspect, the invention provides a cleaning composition comprising at least one protease variant or at least one amylase, wherein the amylase is present at a level from about 0.00001% to about 10% of additional amylase by weight of the composition and the balance of one or more cleaning adjunct materials by weight of composition. In another aspect, the invention includes a cleaning composition comprising at least one protease variant and at least one amylase that is present at a level of about 0.0001% to about 10%, about 0.001% to about 5%, about 0.001% to about 2%, about 0.005% to about 0.5% amylase by weight of the composition.
[0374] Any suitable cellulase may find used in a cleaning composition of the present invention. In one aspect, the invention provides a cleaning composition comprising at least one protease variant of the invention and one or at least one cellulase. Suitable cellulases include, but are not limited to, e.g., those of bacterial or fungal origin. A chemically or genetically modified mutant of a cellulase may be included in a composition of the invention. Suitable cellulases include, but are not limited to, e.g., Humicola insolens cellulases (see, e.g., U.S. Pat. No. 4,435,307) and cellulases having color care benefits (see, e.g., EP 0 495 257). Additional suitable cellulases are known in the art. Commercially available cellulases that find use and may be included in a composition of the invention include, but are not limited to, e.g., CELLUZYME.RTM., CAREZYME.RTM. (Novozymes), and KAC-500(B).TM. (Kao Corporation), Puradax 7000L, Puradax HA 4000G (Genencor). In one aspect, cellulases are incorporated as portions or fragments of mature wild-type or variant cellulases, wherein a portion of the N-terminus is deleted (see, e.g., U.S. Pat. No. 5,874,276). In one aspect, a cleaning composition of the invention comprises at least one protease variant of the invention and at least one cellulase at a level from about 0.00001% to about 10% of additional cellulase by weight of the composition and the balance of one or more cleaning adjunct materials by weight of composition. In another aspect, the invention provides a cleaning composition comprising at least one protease variant of the invention and at least one cellulase at a level of about 0.0001% to about 10%, about 0.001% to about 5%, about 0.001% to about 2%, about 0.005% to about 0.5% cellulase by weight of the composition.
[0375] Any mannanase suitable for use in detergent compositions also finds use in and thus may be included in a cleaning composition of the invention. In one aspect, the invention provides a cleaning composition comprising at least one protease variant of the invention and least one mannanase. Suitable mannanases include, but are not limited to, e.g., those of bacterial or fungal origin. A chemically or genetically modified mutant of a mannanase may be included in a composition of the invention. Various mannanases are known which are useful and may be included in a composition of invention (see, e.g., mannanases described in U.S. Pat. Nos. 6,566,114, 6,602,842, and 6,440,991, all of which are incorporated herein by reference). In one aspect, the invention provides a cleaning composition comprising at least one protease variant of the invention and at least one mannanase at a level from about 0.00001% to about 10% of additional mannanase by weight of the composition and the balance of one or more cleaning adjunct materials by weight of composition. In some such cleaning compositions, each mannanase is present at a level of about 0.0001% to about 10%, about 0.001% to about 5%, about 0.001% to about 2%, or about 0.005% to about 0.5% mannanase by weight of the composition.
[0376] A peroxidase may be used in combination with hydrogen peroxide or a source thereof (e.g., a percarbonate, perborate or persulfate) in a composition of the invention. An oxidase may be used in combination with oxygen in a composition of the invention. Both such types of enzymes are used for "solution bleaching" (i.e., to prevent transfer of a textile dye from a dyed fabric to another fabric when the fabrics are washed together in a wash liquor), preferably together with an enhancing agent (see, e.g., WO 94/12621 and WO 95/01426). Suitable peroxidases/oxidases that may be included in compositions of the invention include, but are not limited to, e.g., those of plant, bacterial, or fungal origin. A chemically or genetically modified mutant of a peroxidase or oxidase may be included in a composition of the invention. In one aspect, the invention provides a cleaning composition comprising at least one protease variant of the invention and at least one peroxidase and/or at least one oxidase enzyme. Each such peroxidase or oxidase may be present in the composition at a level from about 0.00001% to about 10% of peroxidase or oxidase by weight of the composition and the balance of one or more cleaning adjunct materials by weight of composition. In another aspect, the invention provides a cleaning composition comprising at least one protease variant of the invention and at least one peroxidase and/or at least oxidase enzyme, wherein each such peroxidase or oxidase is present at a level of about 0.0001% to about 10%, about 0.001% to about 5%, about 0.001% to about 2%, or about 0.005% to about 0.5% peroxidase or oxidase enzyme, respectively, by weight of the composition.
[0377] In one aspect, the invention provides a composition (e.g., cleaning composition) comprising at least one protease variant of the invention and one or more additional enzymes find use, including but not limited to, e.g., one or more perhydrolases (see, e.g., WO 05/056782).
[0378] In another aspect, the invention provides a composition (e.g., cleaning composition) comprising at least one protease variant of the invention and one or more mixtures of the above-mentioned enzymes are encompassed, such as, e.g., one or more additional proteases, amylases, lipases, mannanases, and/or cellulases. Indeed, it is contemplated that various mixtures of these enzymes will find use in compositions of the present invention. It is also contemplated that the varying levels of the protease variant(s) and one or more additional enzymes may both independently range to about 10%, the balance of the cleaning composition being one or more cleaning adjunct materials. The specific selection of a cleaning adjunct material is readily made by considering the surface or item (e.g., dishware item, tableware item, or fabric item) or fabric to be cleaned, and the desired form of the composition for the cleaning conditions during use (e.g., through the hand or automatic dishwashing detergent use).
[0379] In one aspect, the invention provides a composition (e.g., cleaning composition) comprising at least one protease variant of the invention (and optionally at least one additional enzyme, if desired) and one or more cleaning adjunct materials. Examples of suitable cleaning adjunct materials include, but are not limited to, e.g., surfactants, builders, bleaches, bleach activators, bleach catalysts, other enzymes, enzyme stabilizing systems, chelants, optical brighteners, soil release polymers, dye transfer agents, dye transfer inhibiting agents, catalytic materials, hydrogen peroxide, sources of hydrogen peroxide, preformed peracids, polymeric dispersing agents, clay soil removal agents, structure elasticizing agents, dispersants, suds suppressors, dyes, perfumes, colorants, filler salts, hydrotropes, perfume, perfume capsule, photoactivators, fluorescers, fabric conditioners, fabric softeners, carriers, hydrotropes, processing aids, solvents, pigments, hydrolyzable surfactants, preservatives, anti-oxidants, anti-shrinkage agents, anti-wrinkle agents, germicides, fungicides, color speckles, silvercare, anti-tarnish and/or anti-corrosion agents, alkalinity sources, solubilizing agents, carriers, processing aids, pigments, and pH control agents (see, e.g., U.S. Pat. Nos. 6,610,642, 6,605,458, 5,705,464, 5,710,115, 5,698,504, 5,695,679, 5,686,014 and 5,646,101, all of which are incorporated herein by reference). Aspects of specific cleaning composition materials are exemplified in detail below. As noted above, if a cleaning adjunct material is not compatible with a protease variant of the present invention included in a desired cleaning composition, then a suitable method of keeping the cleaning adjunct material(s) and the protease(s) separated (i.e., not in contact with each other) until combination of the components is appropriate is used. Such separation methods include any suitable method known in the art (e.g., gelcap, encapsulation, tablets, physical separation, etc.).
[0380] In one aspect, a composition (e.g., cleaning composition) of the invention comprises an effective amount of at least one protease variant of the invention that is useful or effective for cleaning a surface in need of proteinaceous stain removal. Such cleaning compositions include cleaning compositions for such applications as cleaning hard surfaces, laundry, fabrics, dishes, tableware, or dishware (e.g., by hand or manual dishwashing or automatic dishwashing). Indeed, in one aspect, the present invention provides fabric cleaning compositions, while in another aspect, the invention provides non-fabric cleaning compositions. Notably, the invention also provides cleaning compositions comprising at least one protease variant of the invention, wherein such cleaning compositions are suitable for personal care, including oral care (including dentifrices, toothpastes, mouthwashes, etc., as well as denture cleaning compositions), suitable for cleaning skin and hair, and suitable for cleaning eyeglasses. It is intended that the present invention encompass detergent compositions in any form (i.e., liquid, granular, bar, solid, semi-solid, gels, emulsions, tablets, capsules, etc.).
[0381] By way of example, several cleaning compositions wherein the protease variants of the invention find use are described in greater detail below. In one aspect in which the cleaning compositions of the invention are formulated as compositions suitable for use in laundry machine washing method(s), the compositions of the invention preferably contain at least one surfactant and at least one builder compound, as well as one or more cleaning adjunct materials, including, e.g., one or more cleaning adjunct materials selected from the group of organic polymeric compounds, bleaching agents, additional enzymes, suds suppressors, dispersants, lime-soap dispersants, soil suspension and anti-redeposition agents, and corrosion inhibitors. In one aspect, laundry compositions also contain one or more softening agents (i.e., as additional cleaning adjunct materials). Additional exemplary laundry or fabric cleaning compositions and formulations to which one or more protease variants of the invention can be added are presented in the Examples below.
[0382] The compositions of the invention also find use as detergent additive products in solid or liquid form. Such additive products are intended to supplement and/or boost the performance of conventional detergent compositions and can be added at any stage of the cleaning process. In one aspect, the density of the laundry detergent composition ranges from about 400 to about 1200 g/liter, while in other aspects, it ranges from about 500 to about 950 g/liter of composition measured at 20.degree. C.
[0383] In one aspect, the invention provides cleaning compositions, such as those described in U.S. Pat. No. 6,605,458, comprising at least protease variant of the invention. In one aspect, the composition comprising at least one protease variant of the invention is a compact granular fabric cleaning composition, while in other aspects, the composition is a granular fabric cleaning composition useful in the laundering of colored fabrics; in another aspect, the composition is a granular fabric cleaning composition which provides softening through the wash capacity. In another aspect, the composition is a heavy duty liquid fabric cleaning composition. In another aspect, the invention provides a composition comprising at least one protease variant of the invention, wherein the composition is a fabric cleaning composition, such as one described in U.S. Pat. Nos. 6,610,642 and 6,376,450. Also provided are granular laundry detergent compositions of particular utility under European or Japanese washing conditions (see, e.g., U.S. Pat. No. 6,610,642) which comprise at least one protease variant of the invention.
[0384] In one aspect, the invention provides hard surface cleaning compositions comprising at least one protease variant provided herein. Some such compositions comprise hard surface cleaning compositions such as those described in U.S. Pat. Nos. 6,610,642, 6,376,450, and 6,376,450 that include at least one such protease variant.
[0385] The invention includes hand dishwashing or automatic dishwashing detergent compositions comprising at least one protease variant provided herein. Some such compositions comprise hard surface cleaning compositions, such as those described in U.S. Pat. Nos. 6,610,642 and 6,376,450.
[0386] In one aspect, the invention provides cleaning compositions for use in manual or hand dishwashing or automatic dishwashing methods comprising at least one protease variant of the invention and/or at least one surfactant and/or at least one additional cleaning adjunct material selected from the group of organic polymeric compounds, suds enhancing agents, group II metal ions, solvents, hydrotropes, and additional enzymes.
[0387] In one aspect in which the cleaning compositions of the invention are formulated as compositions suitable for use in automatic dishwashing machine method(s), the compositions of the invention typically contain at least one surfactant and/or at least one builder compound and may contain one or more cleaning adjunct materials preferably selected from organic polymeric compounds, bleaching agents, additional enzymes, suds suppressors, dispersants, lime-soap dispersants, soil suspension and anti-redeposition agents and corrosion inhibitors. Additional exemplary dishwashing compositions and formulations to which one or more protease variants of the invention can be added are presented in the Examples below.
[0388] In another aspect, the invention provides oral care compositions comprising at least one protease variant of the present invention that are useful for oral care (e.g., cleaning teeth and dentures); components of oral care compositions that may be useful and included in such compositions include those described in U.S. Pat. No. 6,376,450. Compositions of the invention may further comprise cleaning adjunct materials and compounds described in the U.S. Pat. Nos. 6,376,450, 6,605,458, and 6,610,642, all of which are incorporated by reference herein.
[0389] The cleaning compositions of the invention are formulated into any suitable form and prepared by any process chosen by the formulator, non-limiting examples of which are described in U.S. Pat. Nos. 5,879,584, 5,691,297, 5,574,005, 5,569,645, 5,565,422, 5,516,448, 5,489,392, and 5,486,303, all of which are incorporated herein by reference. When a low pH cleaning composition is desired, the pH of such composition is adjusted via the addition of a material such as monoethanolamine or an acidic material such as hydrogen chloride (HCl).
[0390] While not essential for the purposes of the present invention, the non-limiting list of adjuncts illustrated hereinafter are suitable for use in the instant cleaning compositions. In one aspect, these adjuncts are incorporated, e.g., to assist or enhance cleaning performance for treatment of the substrate to be cleaned or to modify the aesthetics of the cleaning composition as is the case with perfumes, colorants, dyes or the like. It is understood that such adjuncts are in addition to the protease variants of the present invention. The precise nature of these additional components, and levels of incorporation thereof, will depend on the physical form of the composition and the nature of the cleaning operation for which it is to be used. Suitable adjunct materials include, but are not limited to, e.g., surfactants, builders, chelating agents, dye transfer inhibiting agents, deposition aids, dispersants, additional enzymes, and enzyme stabilizers, catalytic materials, bleach activators, bleach boosters, hydrogen peroxide, sources of hydrogen peroxide, preformed peracids, polymeric dispersing agents, clay soil removal/anti-redeposition agents, brighteners, suds suppressors, dyes, perfumes, structure elasticizing agents, fabric softeners, carriers, hydrotropes, processing aids and/or pigments. In addition to the disclosure below, suitable examples of such other adjuncts and levels of use are found in U.S. Pat. Nos. 5,576,282, 6,306,812, and 6,326,348, incorporated by reference. The aforementioned adjunct ingredients may constitute the balance of the cleaning compositions of the present invention.
[0391] In one aspect, cleaning compositions of the invention comprise at least one surfactant and/or a surfactant system, wherein the surfactant is selected from nonionic surfactants, anionic surfactants, cationic surfactants, ampholytic surfactants, zwitterionic surfactants, semi-polar nonionic surfactants and mixtures thereof. In some low pH cleaning composition aspects (e.g., compositions having a neat pH of from about 3 to about 5), the composition typically does not contain alkyl ethoxylated sulfate, as it is believed that such surfactant may be hydrolyzed by such compositions the acidic contents. In one aspect, the surfactant is present at a level of from about 0.1% to about 60%, while in alternative aspects the level is from about 1% to about 50%, while in still further aspects the level is from about 5% to about 40%, by weight of the cleaning composition.
[0392] In one aspect, cleaning compositions of the invention comprise one or more detergent builders or builder systems. In some such compositions incorporating at least one builder, the cleaning compositions comprise at least about 1%, from about 3% to about 60% or even from about 5% to about 40% builder by weight of the cleaning composition. Builders include, but are not limited to, e.g., alkali metal, ammonium and alkanolammonium salts of polyphosphates, alkali metal silicates, alkaline earth and alkali metal carbonates, aluminosilicates, polycarboxylate compounds, ether hydroxypolycarboxylates, copolymers of maleic anhydride with ethylene or vinyl methyl ether, 1, 3, 5-trihydroxy benzene-2, 4, 6-trisulphonic acid, and carboxymethyloxysuccinic acid, the various alkali metal, ammonium and substituted ammonium salts of polyacetic acids such as ethylenediamine tetraacetic acid and nitrilotriacetic acid, as well as polycarboxylates, such as mellitic acid, succinic acid, citric acid, oxydisuccinic acid, polymaleic acid, benzene 1,3,5-tricarboxylic acid, carboxymethyloxysuccinic acid, and soluble salts thereof. It is contemplated that any suitable builder will find use in various compositions of the invention.
[0393] In some such compositions, the builders form water-soluble hardness ion complexes (e.g., sequestering builders), such as citrates and polyphosphates (e.g., sodium tripolyphosphate and sodium tripolyphospate hexahydrate, potassium tripolyphosphate, and mixed sodium and potassium tripolyphosphate, etc.). It is contemplated that any suitable builder will find use in the present invention, including those known in the art (see, e.g., EP 2 100 949).
[0394] Some cleaning compositions of the invention comprise at least one chelating agent in addition to at least one protease variant. Suitable chelating agents include, but are not limited to, e.g., copper, iron, and/or manganese chelating agents and mixtures thereof. In aspects in which at least one chelating agent is used, the cleaning compositions of the present invention comprise from about 0.1% to about 15% or even from about 3% to about 10% chelating agent by weight of the subject cleaning composition.
[0395] Some cleaning compositions provided herein comprise at least one deposition aid in addition to at least one protease variant. Suitable deposition aids include, but are not limited to, e.g., polyethylene glycol, polypropylene glycol, polycarboxylate, soil release polymers such as polytelephthalic acid, clays such as kaolinite, montmorillonite, atapulgite, illite, bentonite, halloysite, and mixtures thereof.
[0396] As indicated herein, in one aspect, anti-redeposition agents find use in one aspect of the present invention. In one aspect, non-ionic surfactants find use. These non-ionic surfactants also find use in preventing the re-deposition of soils. In one aspect, the anti-redeposition agent is a non-ionic surfactant as known in the art (see, e.g., EP 2 100 949).
[0397] In one aspect, the cleaning compositions of the present invention include one or more dye transfer inhibiting agents. Suitable polymeric dye transfer inhibiting agents include, but are not limited to, polyvinylpyrrolidone polymers, polyamine N-oxide polymers, copolymers of N-vinylpyrrolidone and N-vinylimidazole, polyvinyloxazolidones and polyvinylimidazoles or mixtures thereof. In aspects in which at least one dye transfer inhibiting agent is used, the cleaning compositions of the present invention comprise from about 0.0001% to about 10%, from about 0.01% to about 5%, or even from about 0.1% to about 3% by weight of the cleaning composition.
[0398] In one aspect, silicates are included within the compositions of the present invention. In some such aspects, sodium silicates (e.g., sodium disilicate, sodium metasilicate, and crystalline phyllosilicates) find use. In one aspect, silicates are present at a level of from about 1% to about 20%. In one aspect, silicates are present at a level of from about 5% to about 15% by weight of the composition.
[0399] In one aspect, the cleaning compositions of the invention also comprise dispersants. Suitable water-soluble organic materials include, but are not limited to, e.g., the homo- or co-polymeric acids or their salts, in which the polycarboxylic acid comprises at least two carboxyl radicals separated from each other by not more than two carbon atoms.
[0400] In one aspect, the enzymes (e.g., protease variants of the invention or other additional enzymes) used in the cleaning compositions are stabilized any suitable technique. In one aspect, the enzymes employed herein are stabilized by the presence of water-soluble sources of calcium and/or magnesium ions in the finished compositions that provide such ions to the enzymes. In one aspect, the enzyme stabilizers include oligosaccharides, polysaccharides, and inorganic divalent metal salts, including alkaline earth metals, such as calcium salts. It is contemplated that various techniques for enzyme stabilization will find use in the present invention. For example, in one aspect, the enzymes employed herein are stabilized by the presence of water-soluble sources of zinc (II), calcium (II) and/or magnesium (II) ions in the finished compositions that provide such ions to the enzymes, as well as other metal ions (e.g., barium (II), scandium (II), iron (II), manganese (II), aluminum (III), Tin (II), cobalt (II), copper (II), nickel (II), and oxovanadium (IV). Chlorides and sulfates also find use in one aspect of the present invention. Examples of suitable oligosaccharides and polysaccharides (e.g., dextrins) are known in the art (see, e.g., WO 07/145964). In one aspect, reversible protease inhibitors also find use, such as boron-containing compounds (e.g., borate, phenyl boronic acid, 4-formyl phenyl boronic acid, other phenyl boronic acid derivatives, peptide inhibitors, and the like) and/or a tripeptide aldehyde find use in compositions of the invention to further improve stability, as desired.
[0401] In one aspect, one or more bleaches, bleach activators, and/or bleach catalysts are included in the compositions of the invention. In one aspect, the cleaning compositions of the invention comprise inorganic and/or organic bleaching compound(s). Inorganic bleaches include, but are not limited to perhydrate salts (e.g., perborate, percarbonate, perphosphate, persulfate, and persilicate salts). In one aspect, inorganic perhydrate salts are alkali metal salts. In one aspect, inorganic perhydrate salts are included as the crystalline solid, without additional protection, although in some other aspects, the salt is coated. Any suitable salt known in the art finds use in the compositions of the invention (see, e.g., EP 2 100 949).
[0402] In one aspect, bleach activators are used in the compositions of the invention. Bleach activators are typically organic peracid precursors that enhance the bleaching action in the course of cleaning at temperatures of 60.degree. C. and below. Bleach activators suitable for use herein include compounds which, under perhydrolysis conditions, give aliphatic peroxycarboxylic acids having preferably from about 1 to about 10 carbon atoms, in particular from about 2 to about 4 carbon atoms, and/or optionally substituted perbenzoic acid. Additional bleach activators are known in the art and find use in the composition of the invention (see, e.g., EP 2 100 949).
[0403] In one aspect and as further described herein, the cleaning compositions of the invention further comprise at least one bleach catalyst. In one aspect, the manganese triazacyclononane and related complexes find use, as well as cobalt, copper, manganese, and iron complexes. Additional bleach catalysts find use in the present invention (see, e.g., U.S. Pat. Nos. 4,246,612, 5,227,084, and 4,810410; WO 99/06521; and EP 2 100 949).
[0404] In one aspect, the cleaning compositions of the invention comprise one or more catalytic metal complexes. In one aspect, a metal-containing bleach catalyst finds use. In some aspects, the metal bleach catalyst comprises a catalyst system comprising a transition metal cation of defined bleach catalytic activity (e.g., copper, iron, titanium, ruthenium, tungsten, molybdenum, or manganese cations), an auxiliary metal cation having little or no bleach catalytic activity (e.g., zinc or aluminum cations), and a sequestrate having defined stability constants for the catalytic and auxiliary metal cations, particularly ethylenediaminetetraacetic acid, ethylenediaminetetra (methylenephosphonic acid) and water-soluble salts thereof are used (see, e.g., U.S. Pat. No. 4,430,243). In one aspect, the cleaning compositions of the invention are catalyzed by means of a manganese compound. Such compounds and levels of use are well known in the art (see, e.g., U.S. Pat. No. 5,576,282). In additional aspects, cobalt bleach catalysts find use and are included in the cleaning compositions of the invention. Various cobalt bleach catalysts are known in the art (see, e.g., U.S. Pat. Nos. 5,597,936 and 5,595,967) and are readily prepared by known procedures.
[0405] In one aspect, the cleaning compositions of the invention include a transition metal complex of a macropolycyclic rigid ligand (MRL). As a practical matter, and not by way of limitation, in one aspect, the compositions and cleaning processes provided by the invention are adjusted to provide on the order of at least one part per hundred million of the active MRL species in the aqueous washing medium, and in one aspect, provide from about 0.005 ppm to about 25 ppm, from about 0.05 ppm to about 10 ppm, and from about 0.1 ppm to about 5 ppm, of the MRL in the wash liquor.
[0406] In one aspect, transition-metals in the instant transition-metal bleach catalyst include, but are not limited to, e.g., manganese, iron and chromium. Preferred MRLs also include, but are not limited to, e.g., special ultra-rigid ligands that are cross-bridged (e.g., 5,12-diethyl-1,5,8,12-tetraazabicyclo(6.6.2)hexadecane). Suitable transition metal MRLs are readily prepared by known procedures (see, e.g., WO 2000/32601 and U.S. Pat. No. 6,225,464).
[0407] In one aspect, the invention provides an automatic dishwashing detergent composition formulated as a detergent tablet. Such tablet comprises at least one protease variant of the invention and a builder, such as, e.g., a builder salt. Some such tablets have an alkalinity of at least equivalent to 3 grams (g) of sodium hydroxide per 100 grams of the tablet composition and a density of at least 1.4 grams/cubic centimeter. The builder salt can comprise a mixture of a silicate salt and a phosphate salt, preferably with more silicate (e.g., sodium metasilicate) than phosphate (e.g., sodium tripolyphosphate). Some such tablets are free of surfactant materials and are especially adapted for use in automatic dishwashing machines.
[0408] In one aspect, the cleaning compositions of the present invention comprise metal care agents. Metal care agents are useful in preventing and/or reducing the tarnishing, corrosion, and/or oxidation of metals, including aluminum, stainless steel, and non-ferrous metals (e.g., silver and copper). Suitable metal care agents include those described in EP 2 100 949, WO 9426860, and WO 94/26859). In some such cleaning compositions, the metal care agent is a zinc salt. Some such cleaning compositions comprise from about 0.1% to about 5% by weight of one or more metal care agent.
[0409] As indicated above, the cleaning compositions of the present invention are formulated into any suitable form and prepared by any process chosen by the formulator, non-limiting examples of which are described in U.S. Pat. Nos. 5,879,584, 5,691,297, 5,574,005, 5,569,645, 5,516,448, 5,489,392, and 5,486,303, all of which are incorporated herein by reference. In one aspect in which a low pH cleaning composition is desired, the pH of such composition is adjusted via the addition of an acidic material such as hydrogen chloride (HCl).
[0410] The cleaning compositions disclosed herein find use in cleaning a situs (e.g., a surface, dishware, tableware, or fabric). Typically, at least a portion of the situs is contacted with a present cleaning composition of the invention in neat form or diluted in a wash liquor, and then the situs is optionally washed and/or rinsed. For purposes of the present invention, "washing" includes, but is not limited to, e.g., scrubbing and mechanical agitation. In one aspect, the cleaning compositions are employed at concentrations of from about 500 ppm to about 15,000 ppm in solution. When the wash solvent is water, the water temperature typically ranges from about 5.degree. C. to about 90.degree. C. and when the situs comprises a fabric, the water to fabric mass ratio is typically from about 1:1 to about 30:1.
Processes of Making and Using Compositions
[0411] The compositions of the invention described herein and throughout, including, e.g., cleaning compositions, can be formulated into any suitable form and prepared by any suitable process chosen by the formulator (see, e.g., U.S. Pat. Nos. 5,879,584, 5,691,297, 5,574,005, 5,569,645, 5,565,422, 5,516,448, 5,489,392, 5,486,303, 4,515,705, 4,537,706, 4,515,707, 4,550,862, 4,561,998, 4,597,898, 4,968,451, 5,565,145, 5,929,022, 6,294,514 and 6,376,445).
[0412] In one aspect, the cleaning compositions of the invention are provided in unit dose form, including tablets, capsules, sachets, pouches, and multi-compartment pouches. In one aspect, the unit dose format is designed to provide controlled release of the ingredients within a multi-compartment pouch (or other unit dose format). Suitable unit dose and controlled release formats are known in the art (see, e.g., EP 2 100 949, WO 02/102955, U.S. Pat. Nos. 4,765,916 and 4,972,017, and WO 04/111178 for materials suitable for use in unit dose and controlled release formats). In one aspect, the unit dose form is provided by tablets wrapped with a water-soluble film or water-soluble pouches. Various formats for unit doses are provided in EP 2 100 947, and are known in the art.
[0413] Additional aspects of the invention relating to processes of making and using compositions of the invention are described elsewhere herein.
Methods of the Invention
[0414] The invention provides methods for cleaning or washing an item or surface (e.g., hard surface) in need of cleaning, including, but not limited to, e.g., methods for cleaning or washing a dishware item, a tableware item, a fabric item, a laundry item, personal care item, eye glass, etc., or the like, and methods for cleaning or washing a hard or soft surface, such as, e.g., a hard surface of an item.
[0415] In one aspect, the invention provides a method for cleaning an item, object, or surface in need of cleaning, the method comprising contacting the item or surface (or a portion of the item or surface desired to be cleaned) with a protease variant of any of the invention or a composition of the invention for a sufficient time and/or under conditions suitable or effective to clean the item, object, or surface to a desired degree. Some such methods further comprise rinsing the item, object, or surface with water. For some such methods, the cleaning composition is a dishwashing detergent composition and the item or object to be cleaned is a dishware item or tableware item. A dishware item is a dish item generally used in serving or eating food. A dishware item can be, but is not limited to, e.g., a dish, plate, cup, bowl, etc., and the like. Tableware is a broader term that includes, but is not limited, to, e.g., dishes, cutlery, knives, forks, spoons, chopsticks, glassware, pitchers, sauce boats, drinking vessels, etc., and the like; a tableware item includes any of these or similar items for serving or eating food. For some such methods, the cleaning composition is an automatic dishwashing detergent composition or a hand dishwashing detergent composition and the item or object to be cleaned is a dishware or tableware item. For some such methods, the cleaning composition is a laundry detergent composition, such as, e.g., a power laundry detergent composition or a liquid laundry detergent composition, and the item to be cleaned is a fabric item.
[0416] In one aspect, the invention provides methods for cleaning or washing a fabric item optionally in need of cleaning or washing, respectively. Some such methods comprise providing a composition comprising the protease variant (such as, but not limited to, e.g., a fabric or laundry cleaning composition) and a fabric item or laundry item in need of cleaning, and contacting the fabric item or laundry item (or a portion of the item desired to be cleaned) with the composition under conditions sufficient or effective to clean or wash the fabric or laundry item to a desired degree.
[0417] In one aspect, the invention provides a method for cleaning or washing an item or surface (e.g., hard surface) optionally in need of cleaning, the method comprising providing an item or surface to be cleaned or washed and contacting the item or surface (or a portion of the item or surface desired to be cleaned or washed) with at least one protease variant of the invention or a composition of the invention comprising at least one such protease variant for a sufficient time and/or under conditions sufficient or effective to clean or wash the item or surface to a desired degree. Such compositions include, but are not limited to, e.g., a cleaning composition or detergent composition of the invention (including, but not limited to, e.g., hand dishwashing detergent composition, hand dishwashing cleaning composition, laundry detergent or fabric detergent or laundry or fabric cleaning composition, liquid laundry detergent, liquid laundry cleaning composition, powder laundry detergent composition, powder laundry cleaning composition, automatic dishwashing detergent composition, laundry booster cleaning or detergent composition, laundry cleaning additive, and laundry pre-spotter composition, etc.). In some instances, if desired, the method can be repeated one or more times, particularly if additional cleaning or washing is desired. For example, in some instance, the method optionally further comprises allowing the item or surface to remain in contact with the at least one protease variant or composition for a period of time sufficient or effective to clean or wash the item or surface to the desired degree. Some such methods further comprise rinsing the item or surface with water. Some such methods further comprise contacting the item or surface with at least one protease variant of the invention or a composition of the invention again and allowing the item or surface to remain in contact with the at least one protease variant or composition for a period of time sufficient to clean or wash the item or surface to the desired degree. For some such methods, the cleaning composition is a dishwashing detergent composition and the item to be cleaned is a dishware or tableware item. For some such methods, the cleaning composition is an automatic dishwashing detergent composition or a hand dishwashing detergent composition and the item to be cleaned is a dishware or tableware item. For some such methods, the cleaning composition is a laundry detergent composition and the item to be cleaned is a fabric item.
[0418] In one aspect, the invention provides a method of cleaning a tableware or dishware item in an automatic dishwashing machine, the method comprising providing an automatic dishwashing machine, placing an amount of an automatic dishwashing composition comprising at least one protease variant of the invention or a composition of the invention sufficient to clean the tableware or dishware item in the machine (e.g., by placing the composition in an appropriate or provided detergent compartment or dispenser in the machine), putting a dishware or tableware item in the machine, and operating the machine so as to clean the tableware or dishware item (e.g., as per the manufacturer's instructions). Such method can include any automatic dishwashing composition described herein, which comprises, but is not limited to, e.g., any protease variant described herein. The amount of automatic dishwashing composition to be used can be readily determined according to manufacturer's instructions or suggestions and any form of automatic dishwashing composition comprising at least one protease variant of the invention (e.g., liquid, powder, solid, gel, table, etc.), including any described herein, may be employed.
[0419] In one aspect, the invention provides a method for cleaning a surface, item or object optionally in need of cleaning, the method comprises contacting the item or surface (or a portion of the item or surface desired to be cleaned) with at least one protease variant of the invention or a cleaning composition of the invention in neat form or diluted in a wash liquor for a sufficient time and/or under conditions sufficient or effective to clean or wash the item or surface to a desired degree. The surface, item, or object may then be (optionally) washed and/or rinsed if desired. For purposes of the present invention, "washing" includes, but is not limited to, e.g., scrubbing and mechanical agitation. In one aspect, the cleaning compositions are employed at concentrations of from about 500 ppm to about 15,000 ppm in solution (e.g., aqueous solution). When the wash solvent is water, the water temperature typically ranges from about 5.degree. C. to about 90.degree. C. and when the situs comprises a fabric, the water to fabric mass ratio is typically from about 1:1 to about 30:1.
[0420] In one aspect, the invention provides a method of cleaning a laundry or fabric item in an washing machine, the method comprising providing an washing machine, placing an amount of a laundry detergent composition comprising at least one protease variant of the invention sufficient to clean the laundry or fabric item in the machine (e.g., by placing the composition in an appropriate or provided detergent compartment or dispenser in the machine), putting the laundry or fabric item in the machine, and operating the machine so as to clean the tableware or dishware item (e.g., as per the manufacturer's instructions). Such method can include any laundry washing detergent composition described herein, which comprises, but is not limited to, e.g., any protease variant described herein. The amount of laundry detergent composition to be used can be readily determined according to manufacturer's instructions or suggestions and any form of laundry detergent composition comprising at least one protease variant of the invention (e.g., solid, powder, liquid, tablet, gel, etc.), including any described herein, may be employed.
[0421] In an eighteenth aspect, the invention provides a method for cleaning an item, object, or surface in need of cleaning, the method comprising contacting the item, object, or surface with (i) a variant (or polypeptide as in the third aspect) of the first, second, third, fourth, fifth, sixth, seventh, eighth, or ninth aspect of the invention or (ii) a composition of the seventeenth aspect of the invention. A method of the eighteenth aspect of the invention may further comprise rinsing the item, object, or surface with water.
[0422] In a nineteenth aspect, the invention provides a method for cleaning an item or hard surface, the method comprising contacting at least a portion of the item or hard surface to be cleaned with (i) a variant (or polypeptide as in the third aspect) of the first, second, third, fourth, fifth, sixth, seventh, eighth, or ninth aspect of the invention or (ii) a composition of the seventeenth aspect of the invention for a sufficient time and/or under conditions sufficient or effective to clean or wash the item or hard surface to a desired degree, and optionally comprising rinsing the item or hard surface with water.
[0423] In a twentieth aspect, the invention provides a method of treating and/or cleaning a surface or fabric comprising the steps of optionally washing and/or rinsing said surface or fabric, contacting said surface or fabric with (i) a variant (or polypeptide as in the third aspect) of the first, second, third, fourth, fifth, sixth, seventh, eighth, or ninth aspect of the invention or (ii) a composition of the seventeenth aspect of the invention, and then optionally washing and/or rinsing said surface or fabric.
[0424] Included is a use of a protease variant or polypeptide of the invention in a detergent composition, including, but not limited to, in a laundry and/or dishwashing detergent composition.
[0425] Additional exemplary cleaning methods are provided throughout and in the Examples below.
Experimental
[0426] In the experimental disclosure of Part I Examples and Part I Examples which follow, the following abbreviations apply: PI (Performance Index), ppm (parts per million); M (molar); mM (millimolar); .mu.M (micromolar); nM (nanomolar); mol (moles); mmol (millimoles); .mu.mol (micromoles); nmol (nanomoles); gm (grams); mg (milligrams); .mu.g (micrograms); pg (picograms); L (liters); ml and mL (milliliters); .mu.l and .mu.L (microliters); cm (centimeters); mm (millimeters); .mu.m (micrometers); nm (nanometers); U (units); V (volts); MW (molecular weight); sec (seconds); min(s) (minute/minutes); h(s) and hr(s) (hour/hours); .degree. C. (degrees Centigrade); QS (quantity sufficient); ND (not done); rpm (revolutions per minute); GH (degrees German hardness); H.sub.2O (water); dH.sub.2O (deionized water); HCl (hydrochloric acid); aa (amino acid); bp (base pair); kb (kilobase pair); kD (kilodaltons); cDNA (copy or complementary DNA); DNA (deoxyribonucleic acid); ssDNA (single stranded DNA); dsDNA (double stranded DNA); dNTP (deoxyribonucleotide triphosphate); RNA (ribonucleic acid); MgCl.sub.2 (magnesium chloride); NaCl (sodium chloride); w/v (weight to volume); v/v (volume to volume); w/w (weight to weight); g (gravity); OD (optical density); ppm (parts per million); Dulbecco's phosphate buffered solution (DPBS); SOC (2% Bacto-Tryptone, 0.5% Bacto Yeast Extract, 10 mM NaCl, 2.5 mM KCl); Terrific Broth (TB; 12 g/l Bacto-Tryptone, 24 g/l glycerol, 2.31 g/l KH.sub.2PO.sub.4, and 12.54 g/l K.sub.2HPO.sub.4); OD.sub.280 (optical density at 280 nm); OD.sub.600 (optical density at 600 nm); A.sub.405 (absorbance at 405 nm); Vmax (the maximum initial velocity of an enzyme catalyzed reaction); PAGE (polyacrylamide gel electrophoresis); PBS (phosphate buffered saline [150 mM NaCl, 10 mM sodium phosphate buffer, pH 7.2]); PBST (PBS+0.25% TWEEN.RTM.-20); PEG (polyethylene glycol); PCR (polymerase chain reaction); RT-PCR (reverse transcription PCR); SDS (sodium dodecyl sulfate); Tris (tris(hydroxymethyl)aminomethane); HEPES (N-[2-Hydroxyethyl]piperazine-N-[2-ethanesulfonic acid]); HBS (HEPES buffered saline); Tris-HCl (tris[Hydroxymethyl]aminomethane-hydrochloride); Tricine (N-[tris-(hydroxymethyl)-methyl]-glycine); CHES (2-(N-cyclo-hexylamino) ethane-sulfonic acid); TAPS (3-{[tris-(hydroxymethyl)-methyl]-amino}-propanesulfonic acid); CAPS (3-(cyclo-hexylamino)-propane-sulfonic acid; DMSO (dimethyl sulfoxide); DTT (1,4-dithio-DL-threitol); SA (sinapinic acid (s,5-dimethoxy-4-hydroxy cinnamic acid); TCA (trichloroacetic acid); Glut and GSH (reduced glutathione); GSSG (oxidized glutathione); TCEP (Tris[2-carboxyethyl] phosphine); Ci (Curies); mCi (milliCuries); Ci (microCuries); HPLC (high-performance liquid chromatography); RP-HPLC (reverse phase high pressure liquid chromatography); TLC (thin layer chromatography); MALDI-TOF (matrix-assisted laser desorption/ionization--time of flight); Ts (tosyl); Bn (benzyl); Ph (phenyl); Ms (mesyl); Et (ethyl), Me (methyl); Taq (Thermus aquaticus DNA polymerase); Klenow (DNA polymerase I large (Klenow) fragment); EGTA (ethylene glycol-bis( -aminoethyl ether) N, N, N', N'-tetraacetic acid); EDTA (ethylenediaminetetracetic acid); bla (.beta.-lactamase or ampicillin-resistance gene); HDL (heavy duty liquid); HDD (heavy duty powder detergent); HSG (high suds granular detergent); CEE (Central and Eastern Europe); WE (Western Europe); NA, when used in reference to detergents (North America); Japan and JPN, when used in reference to detergents (Japan); MTP (microtiter plate); MJ Research (MJ Research, Reno, Nev.); Baseclear (Baseclear BV, Inc., Leiden, The Netherlands); PerSeptive (PerSeptive Biosystems, Framingham, Mass.); ThermoFinnigan (ThermoFinnigan, San Jose, Calif.); Argo (Argo BioAnalytica, Morris Plains, N.J.); Seitz EKS (SeitzSchenk Filtersystems GmbH, Bad Kreuznach, Germany); Pall (Pall Corp., East Hills, N.Y. and Bad Kreuznach, Germany); Spectrum (Spectrum Laboratories, Dominguez Rancho, Calif.); Molecular Structure (Molecular Structure Corp., Woodlands, Tex.); Accelrys (Accelrys, Inc., San Diego, Calif.); Chemical Computing (Chemical Computing Corp., Montreal, Canada); New Brunswick (New Brunswick Scientific, Co., Edison, N.J.); CFT (Center for Test Materials, Vlaardingen, The Netherlands); P&G and Procter & Gamble (Procter & Gamble, Inc., Cincinnati, Ohio); GE Healthcare (GE Healthcare, Chalfont St. Giles, United Kingdom); DNA2.0 (DNA2.0, Menlo Park, Calif.); OXOID (Oxoid, Basingstoke, Hampshire, UK); Megazyme (Megazyme International Ireland Ltd., Bray Business Park, Bray, Co., Wicklow, Ireland); Finnzymes (Finnzymes Oy, Espoo, Finland); Kelco (CP Kelco, Wilmington, Del.); Corning (Corning Life Sciences, Corning, N.Y.); (NEN (NEN Life Science Products, Boston, Mass.); Pharma AS (Pharma AS, Oslo, Norway); Dynal (Dynal, Oslo, Norway); Bio-Synthesis (Bio-Synthesis, Lewisville, Tex.); ATCC (American Type Culture Collection, Rockville, Md.); Gibco/BRL (Gibco/BRL, Grand Island, N.Y.); Sigma (Sigma Chemical Co., St. Louis, Mo.); Pharmacia (Pharmacia Biotech, Piscataway, N.J.); NCBI (National Center for Biotechnology Information); Applied Biosystems (Applied Biosystems, Foster City, Calif.); BD Biosciences and/or Clontech (BD Biosciences CLONTECH Laboratories, Palo Alto, Calif.); Operon Technologies (Operon Technologies, Inc., Alameda, Calif.); MWG Biotech (MWG Biotech, High Point, N.C.); Oligos Etc (Oligos Etc. Inc, Wilsonville, Oreg.); Bachem (Bachem Bioscience, Inc., King of Prussia, Pa.); Difco (Difco Laboratories, Detroit, Mich.); Mediatech (Mediatech, Herndon, Va.; Santa Cruz (Santa Cruz Biotechnology, Inc., Santa Cruz, Calif.); Oxoid (Oxoid Inc., Ogdensburg, N.Y.); Worthington (Worthington Biochemical Corp., Freehold, N.J.); GIBCO BRL or Gibco BRL (Life Technologies, Inc., Gaithersburg, Md.); Millipore (Millipore, Billerica, Mass.); Bio-Rad (Bio-Rad, Hercules, Calif.); Invitrogen (Invitrogen Corp., San Diego, Calif.); NEB (New England Biolabs, Beverly, Mass.); Sigma (Sigma Chemical Co., St. Louis, Mo.); Pierce (Pierce Biotechnology, Rockford, Ill.); Takara (Takara Bio Inc. Otsu, Japan); Roche (Hoffmann-La Roche, Basel, Switzerland); Gene Oracle (Gene Oracle, Inc., Mountain View, Calif.); EM Science (EM Science, Gibbstown, N.J.); Qiagen (Qiagen, Inc., Valencia, Calif.); Biodesign (Biodesign Intl., Saco, Me.); Aptagen (Aptagen, Inc., Herndon, Va.); Sorvall (Sorvall brand, from Kendro Laboratory Products, Asheville, N.C.); Molecular Devices (Molecular Devices, Corp., Sunnyvale, Calif.); R&D Systems (R&D Systems, Minneapolis, Minn.); Siegfried Handel (Siegfried Handel AG, Zofingen, Switzerland); Stratagene (Stratagene Cloning Systems, La Jolla, Calif.); Marsh (Marsh Biosciences, Rochester, N.Y.); Geneart (Geneart GmbH, Regensburg, Germany); Bio-Tek (Bio-Tek Instruments, Winooski, Vt.); (Biacore (Biacore, Inc., Piscataway, N.J.); PeproTech (PeproTech, Rocky Hill, N.J.); SynPep (SynPep, Dublin, Calif.); New Objective (New Objective brand; Scientific Instrument Services, Inc., Ringoes, N.J.); Waters (Waters, Inc., Milford, Mass.); Matrix Science (Matrix Science, Boston, Mass.); Dionex (Dionex, Corp., Sunnyvale, Calif.); Monsanto (Monsanto Co., St. Louis, Mo.); Wintershall (Wintershall AG, Kassel, Germany); BASF (BASF Co., Florham Park, N.J.); Huntsman (Huntsman Petrochemical Corp., Salt Lake City, Utah); Shell Chemicals (Shell Chemicals, Inc., London, UK); Stepan (Stepan, Northfield, Ill.); Clariant (Clariant, Sulzbach, Germany); Industrial Zeolite (Industrial Zeolite Ltd., Grays, Essex, UK); Jungbunzlauer (Jungbunzlauer, Basel, Switzerland); Solvay (Solvay, Brussels, Belgium); 3V Sigma (3V Sigma, Bergamo, Italy); Innospec (Innospec, Ellesmere Port, UK); Thermphos (Thermphos, Vlissiggen-Ost, The Netherlands); Ciba Specialty (Ciba Specialty Chemicals, Basel, Switzerland); Dow Corning (Dow Corning, Barry, UK); Enichem (Enichem Iberica, Barcelona, Spain); Fluka Chemie AG (Fluka Chemie AG, Buchs, Switzerland); Gist-Brocades (Gist-Brocades, NV, Delft, The Netherlands); Dow Corning (Dow Corning Corp., Midland, Mich.); Mettler-Toledo (Mettler-Toledo Inc, Columbus, Ohio); RB (Reckitt-Benckiser, Slough, UK); and Microsoft (Microsoft, Inc., Redmond, Wash.).
[0427] In the exemplified detergent compositions provided herein, the enzymes levels are expressed by pure enzyme by weight of the total composition and unless otherwise specified, the detergent ingredients are expressed by weight of the total compositions. The abbreviated component identifications therein have the following meanings:
TABLE-US-00002 Abbreviation Ingredient LAS Sodium linear C.sub.11-13 alkyl benzene sulfonate NaC16-17HSAS Sodium C.sub.16-17 highly soluble alkyl sulfate TAS Sodium tallow alkyl sulphate CxyAS Sodium C.sub.1x-C.sub.1y alkyl sulfate CxyEz C.sub.1x-C.sub.1y predominantly linear primary alcohol condensed with an average of z moles of ethylene oxide CxyAEzS C.sub.1x-C.sub.1y sodium alkyl sulfate condensed with an average of z moles of ethylene oxide. Added molecule name in the examples. Nonionic Mixed ethoxylated/propoxylated fatty alcohol e.g. Plurafac LF404 being an alcohol with an average degree of ethoxylation of 3.8 and an average degree of propoxylation of 4.5. QAS R.sub.2.cndot.N.sup.+(CH.sub.3).sub.2(C.sub.2H.sub.4OH) with R.sub.2 = C.sub.12-C.sub.14 Silicate Amorphous Sodium Silicate (SiO.sub.2:Na.sub.2O ratio = 1.6-3.2:1) Metasilicate Sodium metasilicate (SiO.sub.2:Na.sub.2O ratio = 1.0) Zeolite A Hydrated aluminosilicate of formula Na.sub.12(A1O.sub.2SiO.sub.2).sub.12.cndot.27H.sub.2O SKS-6 Crystalline layered silicate of formula .delta.-Na.sub.2Si.sub.2O.sub.5 Sulfate Anhydrous sodium sulphate STPP Sodium Tripolyphosphate MA/AA Random copolymer of 4:1 acrylate/maleate, average molecular weight about 70,000-80,000 AA Sodium polyacrylate polymer of average molecular weight (MW) 4,500 Polycarboxylate Copolymer comprising mixture of carboxylated monomers such as acrylate, maleate and methyacrylate with a MW ranging between 2,000-80,000 such as Sokolan commercially available from BASF, being a copolymer of acrylic acid, MW 4,500 BB1 3-(3,4-Dihydroisoquinolinium)propane sulfonate BB2 1-(3,4-dihydroisoquinolinium)-decane-2-sulfate PB1 Sodium perborate monohydrate PB4 Sodium perborate tetrahydrate of nominal formula NaBO.sub.3.cndot.4H.sub.2O Percarbonate Sodium percarbonate of nominal formula 2Na.sub.2CO.sub.3.cndot.3H.sub.2O.sub.2 TAED Tetraacetyl ethylene diamine NOBS Nonanoyloxybenzene sulfonate in the form of the sodium salt DTPA Diethylene triamine pentaacetic acid HEDP 1,1-hydroxyethane diphosphonic acid DETPMP Diethyltriamine penta (methylene) phosphonate, marketed by Monsanto under the Trade name Dequest 2060 EDDS Ethylenediamine-N,N'-disuccinic acid, (S,S) isomer in the form of its sodium salt Diamine Dimethyl aminopropyl amine; 1,6-hezane diamine; 1,3-propane diamine; 2-methyl-1,5-pentane diamine; 1,3-pentanediamine; 1- methyl-diaminopropane DETBCHD 5, 12- diethyl-1,5,8,12-tetraazabicyclo [6,6,2] hexadecane, dichloride, Mn(II) SALT PAAC Pentaamine acetate cobalt(III) salt Paraffin Paraffin oil sold under the tradename Winog 70 by Wintershall Paraffin Sulfonate A Paraffin oil or wax in which some of the hydrogen atoms have been replaced by sulfonate groups Aldose oxidase Oxidase enzyme sold under the tradename Aldose Oxidase by Novozymes A/S Galactose oxidase Galactose oxidase from Sigma nprE The recombinant form of neutral metalloprotease expressed in Bacillus subtilis (see, e.g., WO 07/044993) PMN Purified neutral metalloprotease from Bacillus amyloliquefaciens Amylase A suitable amylolytic enzyme, such as those sold under the tradenames PURAFECT .RTM. Ox described in WO 94/18314, WO96/05295 sold by Genencor; NATALASE .RTM., TERMAMYL .RTM., FUNGAMYl .RTM. and DURAMYL .TM., all available from Novozymes A/S. Lipase A suitable lipolytic enzyme such as those sold under the tradenames LIPEX .RTM., LIPOLASE .RTM., LIPOLASE .RTM. Ultra by Novozymes A/S and Lipomax .TM. by Gist-Brocades. Cellulase A suitable cellulytic enzyme such as those sold under the tradenames CAREZYME .RTM., CELLUZYME .RTM., and/or ENDOLASE .RTM. by Novozymes A/S. Pectin Lyase A suitable pectin lyase, such as those sold under the tradenames XPECT .RTM., PECTAWAY .RTM. and PECTAWASH .RTM. available from Novozymes A/S. PVP Polyvinylpyrrolidone with an average molecular weight of 60,000 PVNO Polyvinylpyridine-N-Oxide, with an average molecular weight of 50,000 PVPVI Copolymer of vinylimidazole and vinylpyrrolidone, with an average molecular weight of 20,000 Brightener 1 Disodium 4,4'-bis(2-sulphostyryl)biphenyl Silicone antifoam Polydimethylsiloxane foam controller with siloxane-oxyalkylene copolymer as dispersing agent with a ratio of said foam controller to said dispersing agent of 10:1 to 100:1 Suds Suppressor 12% Silicone/silica, 18% stearyl alcohol, 70% starch in granular form SRP 1 Anionically end capped poly esters PEG X Polyethylene glycol of a molecular weight of x PVP K60 .RTM. Vinylpyrrolidone homopolymer (average MW 160,000) Jeffamine .RTM. ED-2001 Capped polyethylene glycol from Huntsman Isachem .RTM. AS A branched alcohol alkyl sulphate from Enichem MME PEG (2000) Monomethyl ether polyethylene glycol (MW 2000) from Fluka Chemie AG DC3225C Silicone suds suppresser, mixture of Silicone oil and Silica from Dow Corning TEPAE Tetreaethylenepentaamine ethoxylate BTA Benzotriazole Betaine (CH.sub.3).sub.3N.sup.+CH.sub.2COO.sup.- Sugar Industry grade D-glucose or food grade sugar CFAA C.sub.12-C.sub.14 alkyl N-methyl glucamide TPKFA C.sub.12-C.sub.14 topped whole cut fatty acids Clay A hydrated aluminumu silicate in a general formula Al.sub.2O.sub.3SiO.sub.2.cndot.xH.sub.2O. Types: Kaolinite, montmorillonite, atapulgite, illite, bentonite, halloysite pH Measured as a 1% solution in distilled water at 20.degree. C.
[0428] For North American (NA) and Western European (WE) heavy duty liquid laundry (HDL) detergents, heat inactivation of the enzymes present in commercially-available detergents is performed by placing pre-weighed liquid detergent (in a glass bottle) in a water bath at 95.degree. C. for 2 hours. The incubation time for heat inactivation of NA and WE auto dish washing (ADW) detergents is 8 hours. Both un-heated and heated detergents are assayed within 5 minutes of dissolving the detergent to accurately determine percentage deactivated. Enzyme activity is tested by the AAPF assay.
[0429] For testing of enzyme activity in heat-inactivated detergents, working solutions of detergents are made from the heat inactivated stocks. Appropriate amounts of water hardness (e.g., 6 gpg or 12 gpg) and buffer are added to the detergent solutions to match the desired conditions. The solutions are mixed by vortexing or inverting the bottles. The following Table provides information regarding some of the commercially-available detergents and test conditions used herein. In some experiments, additional and/or other commercially available detergents find use in the following Examples.
TABLE-US-00003 TABLE 1.1 Laundry and Dish Washing Conditions Region Form Dose Detergent* Buffer gpg pH T (.degree. C.) Laundry (Heavy Duty Liquid and Granular) NA HDL 0.78 g/1 P&G TIDE .RTM. 2X 5 mM HEPES 6 8.0 20 WE HDL 5.0 g/L Henkel PERSIL .TM. 5 mM HEPES 12 8.2 40 WE HDG 8.0 g/L P&G ARIEL .RTM. 2 mM Na.sub.2 CO.sub.3 12 10.5 40 JPN HDG 0.7 g/L P&G TIDE .RTM. 2 mM Na.sub.2 CO.sub.3 6 10.0 20 NA HDG 1.0 g/L P&G TIDE .RTM. 2 mM Na.sub.2 CO.sub.3 6 10.0 20 Automatic Dish Washing WE ADW 3.0 g/L RB CALGONIT .TM. 2 mM Na.sub.2 CO.sub.3 21 10.0 40 NA ADW 3.0 g/L P&G CASCADE .RTM. 2 mM Na.sub.2 CO.sub.3 9 10.0 40
[0430] In some additional aspects, the following solutions find use:
TABLE-US-00004 TABLE 1-2 Working Detergent Solutions Temp Detergent Detergent (C.) g/L pH Buffer gpg TIDE .RTM. 2X Cold 16 0.98 8 5 mM HEPES 6 TIDE .RTM. 2X Cold 32 0.98 8 5 mM HEPES 6 TIDE .RTM. 2X Cold 16 0.98 7 5 mM MOPS 6
[0431] The examples, which are intended to be purely exemplary of the invention and should therefore not be considered to limit the invention in any way, also describe and detail aspects and embodiments of the invention discussed above. The foregoing examples and detailed description are offered by way of illustration and not by way of limitation.
Part I
Table of Detergents
[0432] The compositions of the detergents used in the assays in Part I Examples are shown in Table 1-3. BPN' variant protein samples were added to the detergent compositions as described in Part I Example 1 to assay for the various properties tested.
[0433] The following are liquid laundry detergent compositions suitable for top-loading automatic washing machines (1, 2 & 4) and front loading washing machines (3).
TABLE-US-00005 TABLE 1-3 Composition of Detergents Used in the Assays to Test BPN' Variants Composition (wt % of composition) Ingredient 1 2 3 4 C.sub.12-15 Alkylethoxy(1.8)sulfate 14.7 11.6 16.31 C.sub.11.8 Alkylbenzene sulfonate 4.3 11.6 8.3 7.73 C.sub.16-17 Branched alkyl sulfate 1.7 1.29 3.09 C.sub.12-14 Alkyl -9-ethoxylate 0.9 1.07 1.31 C.sub.12 dimethylamine oxide 0.6 0.64 1.03 Citric acid 3.5 0.65 3 0.66 C.sub.12-18 fatty acid 1.5 2.32 3.6 1.52 Sodium Borate (Borax) 2.5 2.46 1.2 2.53 Sodium C.sub.12-14 alkyl ethoxy 3 sulfate 2.9 C.sub.14-15 alkyl 7-ethoxylate 4.2 C.sub.12-14 Alkyl -7-ethoxylate 1.7 Ca formate 0.09 0.09 0.09 A compound having the following general structure: 1.2 bis((C.sub.2H.sub.5O)(C.sub.2H.sub.4O)n)(CH.sub.3)--N.sup.+--C.sub.xH.sub.- 2x--N.sup.+--(CH.sub.3)- bis((C.sub.2H.sub.5O)(C.sub.2H.sub.4O)n), wherein n = from 20 to 30, and x = from 3 to 8, or sulphated or sulphonated variants thereof Random graft co-polymer.sup.1 1.46 0.5 Ethoxylated Polyethylenimine .sup.2 1.5 1.29 1.44 Diethylene triamine pentaacetic acid 0.34 0.64 0.34 Diethylene triamine penta(methylene phosphonic acid) 0.3 Tinopal AMS-GX 0.06 Tinopal CBS-X 0.2 0.17 0.29 Amphiphilic alkoxylated grease cleaning polymer .sup.3 1.28 1 0.4 1.93 Ethanol 2 1.58 1.6 5.4 Propylene Glycol 3.9 3.59 1.3 4.3 Diethylene glycol 1.05 1.54 1.15 Polyethylene glycol 0.06 0.04 0.1 Monoethanolamine 3.05 2.41 0.4 1.26 NaOH 2.44 1.8 3.01 Sodium Cumene Sulphonate 1 Sodium Formate 0.11 0.09 Water, Aesthetics (Dyes, perfumes) and Minors balance balance balance (Enzymes, solvents, structurants) .sup.1Random graft copolymer is a polyvinyl acetate grafted polyethylene oxide copolymer having a polyethylene oxide backbone and multiple polyvinyl acetate side chains. The molecular weight of the polyethylene oxide backbone is about 6000 and the weight ratio of the polyethylene oxide to polyvinyl acetate is about 40 to 60 and no more than 1 grafting point per 50 ethylene oxide units. .sup.2 Polyethylenimine (MW = 600) with 20 ethoxylate groups per --NH. .sup.3 Amphiphilic alkoxylated grease cleaning polymer is a polyethylenimine (MW = 600) with 24 ethoxylate groups per --NH and 16 propoxylate groups per --NH
PART I EXAMPLES
Example 1
Assays
[0434] Various assays were used as set forth below. Any deviations from the protocols provided below are indicated in the subsequent Examples.
A. TCA Assay for Protein Content Determination in 96-Well Microtiter Plates
[0435] For BPN' and BPN' variants, this assay was started using filtered B. subtilis bacterial culture supernatant from microtiter plates grown 3-4 days at 33-37.degree. C. with shaking at 230-250 rpm and humidified aeration. A fresh 96-well flat bottom microtiter plate (MTP) was used for the assay. First, 100 .mu.L/well of 0.25 N HCl was placed in each well. Then, 25 .mu.L of filtered culture broth was added. The light scattering/absorbance at 405 nm (use 5 sec mixing mode in the plate reader) was then determined in order to provide the "blank" reading. For the test, 100 .mu.L/well of 30% (w/v) trichloroacetic acid (TCA) was placed in the plates and incubated for 10 minutes at room temperature. The light scattering/absorbance at 405 nm (use 5 sec mixing mode in the plate reader) was then determined. The equipment used was a Biomek FX Robot (Beckman Coulter) and a SpectraMAX (type 340; Molecular Devices) MTP Reader; the MTPs were from Costar (type 9017).
[0436] The calculations were performed by subtracting the blank (no TCA) from the test reading with TCA to provide a relative measure of the protein content in the samples. If desired, a standard curve can be created by calibrating the TCA readings with AAPF assays of clones with known conversion factors. However, the TCA results are linear with respect to protein concentration from 250 to 2500 micrograms protein per ml (ppm) and can thus be plotted directly against enzyme performance for the purpose of choosing good-performing variants. The turbidity/light scatter increase in the samples correlates to the total amount of precipitable protein in the culture supernatant.
B. AAPF Protease Assay in 96-well Microtiter Plates
[0437] In order to determine the protease activity of the proteases and variants thereof of the present invention, the hydrolysis of N-succinyl-L-alanyl-L-alanyl-L-prolyl-L-phenyl-p-nitroanilide (suc-AAPF-pNA) was measured. The reagent solutions used were: 100 mM Tris/HCl, pH 8.6, containing 0.005% TWEEN.RTM.-80 (Tris dilution buffer); 100 mM Tris buffer, pH 8.6, containing 10 mM CaCl.sub.2) and 0.005% TWEEN.RTM.-80 (Tris/Ca buffer); and 160 mM suc-AAPF-pNA in DMSO (suc-AAPF-pNA stock solution) (Sigma: S-7388). To prepare a suc-AAPF-pNA working solution, 1 ml suc-AAPF-pNA stock solution was added to 100 ml Tris/Ca buffer and mixed well for at least 10 seconds. The assay was performed by adding 10 .mu.l of diluted protease solution to each well, immediately followed by the addition of 190 .mu.l 1 mg/ml suc-AAPF-pNA working solution. The solutions were mixed for 5 sec., and the absorbance change in kinetic mode (25 readings in 5 minutes) was read at 405 nm in an MTP reader, at 25.degree. C. The protease activity was expressed as AU (activity=.DELTA.ODmin.sup.-1 ml.sup.-1).
C. BMI Microswatch Assay
[0438] Blood, milk and ink (BMI) stained microswatches of 5.5 millimeter circular diameter were obtained from CFT. Before cutting the swatches, the fabric (EMPA 116) was washed with water. One microswatch was placed in each well of a 96-well non-binding microtiter plate (Corning 3641). The detergents used for the assays included Detergent Composition 1, Detergent Composition 2, and Detergent Composition 4. The detergents were diluted in Milli-Q (deionized) water to a working strength concentration of 0.788 g/L. These detergents were buffered with 5 mM HEPES pH 8.2 or pH 7.2, which upon addition to detergent, buffers at pH 8 or pH 7, respectively. Additionally, 6 grains per gallon (gpg) water hardness (3:1 Ca:Mg--CaCl.sub.2: MgCl.sub.2.6H.sub.2O) was added. The detergent solution was pre-equilibrated in an ice-water bath for 16.degree. C. assays (room temperature for 32.degree. C. assays) and pumped into a circulating reservoir (Beckman FX). Then, 190 .mu.l of the desired detergent solution was added to each well of the MTP that contained microswatches. To this mixture, 10 .mu.l of the diluted enzyme master dilution solution was added, providing an approximate enzyme concentration of 0.4-0.5 .mu.g/mL. The master dilution was prepared from the culture supernatants at 8 .mu.g/mL, where the approximate enzyme concentrations of the culture supernatants and BPN'-v3 or BPN'-v36 parent controls were determined using the AAPF protease activity assay, basing the concentration on a purified BPN'-v3 or BPN'-v36 standard of known concentration. The MTP was sealed with tape and placed in the iEMS incubator/shaker (Thermo/Labsystems) pre-set at 16.degree. C. in a refrigerated dairy case or at 32.degree. C. on the benchtop for 20 minutes, with agitation at 1400 rpm. Following incubation under the appropriate conditions, the sealing tape was removed from each plate and 125 .mu.l (150 .mu.l if pipetting by hand for smaller screens) of the solution from each well was transferred into a fresh MTP (Corning 9017). The new MTP containing 125 .mu.l-150 .mu.l of solution/well was read at 600 nm (with 5 sec mixing mode in the plate reader) using a MTP SpectraMax reader (type 340; Molecular Devices). Blank controls containing a microswatch and detergent but no enzyme were also included. The absorbance value obtained was corrected for the blank value (substrate without enzyme), providing a measure of hydrolytic activity. For each sample (variant), the performance index was calculated as described below. This BMI Microswatch Assay, run at 60.degree. F. (16.degree. C.) and pH 8, is referred to herein as the "Test Method."
D. Egg Microswatch Assay
[0439] CS38 aged egg yolk with pigment stained cotton microswatches of 5.5 millimeter circular diameter were obtained from CFT. These swatches were not pre-rinsed in water. One microswatch was placed in each well of a 96-well non-binding microtiter plate (Corning 3641). Detergent Composition 4 was diluted in Milli-Q (deionized) water to a working strength concentration of 0.788 g/L. The detergents were buffered with 5 mM HEPES pH 8.2 which upon addition to detergent, buffers at pH 8. Additionally 6 grains per gallon (gpg) water hardness (3:1 Ca:Mg--CaCl.sub.2: MgCl.sub.2.6H.sub.2O); was added. The detergent solution was pre-equilibrated in an ice-water bath for 16.degree. C. assays (room temperature for 32.degree. C. assays) and pumped into a circulating reservoir (Beckman FX). Then, 185 .mu.l of the desired detergent solution was added to each well of the MTP, containing microswatches. To this mixture, 15 .mu.l of the diluted enzyme master dilution solution was added, providing an approximate enzyme concentration of 0.6 .mu.g/mL in the reaction. The master dilution was prepared from the culture supernatants at 8 .mu.g/mL, where the approximate enzyme concentration of the culture supernatants and BPN'-v3 or BPN'-v36 parent control was determined using the AAPF protease activity assay, basing the concentration on a purified BPN'-v3 or BPN'-v36 standard of known concentration. The MTP was sealed with tape and placed in the iEMS incubator/shaker (Thermo/Labsystems) pre-set at 16.degree. C. in a refrigerated dairy case or at 32.degree. C. on the benchtop for 30 minutes, with agitation at 1400 rpm. Following incubation under the appropriate conditions, the sealing tape was removed from each plate and 125 .mu.l (150 .mu.l if pipetting by hand for smaller screens) of the solution from each well was transferred into a fresh MTP (Corning 9017). The new MTP containing 125 .mu.l-150 .mu.l of solution/well was read at 405 nm (with 5 sec mixing mode in the plate reader) using a MTP SpectraMax reader (type 340; Molecular Devices). Blank controls containing a microswatch and detergent but no enzyme were also included. The absorbance value obtained was corrected for the blank value (substrate without enzyme), providing a measure of hydrolytic activity. For each sample (variant), the performance index was calculated as described below.
E. Grass Microswatch Assay
[0440] Warwick Equest scrubbed grass on woven cotton swatches were obtained from Warwick. These swatches were cut into 5.5 millimeter circular diameter microswatches using a custom made 96-well punch machine that places one microswatch in each well of a 96-well non-binding microtiter plate (Corning 3641). After cutting of the swatches, the fabric was washed in the wells (pre-rinsed) with 100 .mu.L per well of 50% working strength Detergent Composition 4 diluted in water. After 20 minutes of pre-rinsing the 100 .mu.l of 50% detergent rinse was removed carefully pipetting by hand. Detergent Composition 4 was diluted in Milli-Q (deionized) water to a working strength concentration of 0.788 g/L. These detergents were buffered with 5 mM HEPES pH 8.2, which upon addition to detergent, buffers at pH 8. Additionally 6 grains per gallon (gpg) water hardness (3:1 Ca:Mg--CaCl.sub.2: MgCl.sub.2.6H.sub.2O); was added. The detergent solution was pre-equilibrated in an ice-water bath for 16.degree. C. assays (room temperature for 32.degree. C. assays). Then, 180 .mu.l of the desired detergent solution was added to each well of the MTP containing the microswatches, immediately after the pre-rinsing was complete. To this mixture, 20 .mu.l of the diluted enzyme master dilution solution was added making the approximate enzyme in the reaction at 0.8 .mu.g/mL. The master dilution was prepared from the culture supernatants at 8 .mu.g/mL where the approximate enzyme concentration of the culture supernatants and BPN'-v36 parent control was determined using the AAPF protease assay basing the concentration on a purified BPN'-v36 standard of known concentration. The MTP was sealed with tape and placed in the iEMS incubator/shaker (Thermo/Labsystems) pre-set at 16.degree. C. in a refrigerated dairy case or at 32.degree. C. on the benchtop for 30 minutes, with agitation at 1400 rpm. Following incubation under the appropriate conditions, the sealing tape was removed from each plate and 125 .mu.l (150 .mu.l if pipetting by hand for smaller screens) of the solution from each well was transferred into a fresh MTP (Corning 9017). The new MTP containing 125 .mu.l-150 .mu.l of solution/well was read at both 430 nm and 670 nm (with 5 sec mixing mode in the plate reader) using a MTP SpectraMax reader (type 340; Molecular Devices). Blank controls containing a microswatch and detergent but no enzyme were also included. The absorbance value obtained was corrected for the blank value (substrate without enzyme), providing a measure of hydrolytic activity. For each sample (variant), the performance index was calculated as described below.
F. Stability Assay
[0441] The stability of protease variants was determined in the presence of 40% concentrated Detergent Composition 3 diluted in water. The reagents used were Detergent Composition 3 diluted to 50% in Milli-Q water, 10 mM MES 0.01% TWEEN.RTM.-80 pH 5.8 master dilution buffer, AAPF reagents: see protocol AAPF assay. The equipment used was F-bottom MTP (Corning 9017) for dilution of diluted enzyme into detergent as well as for suc-AAPF-pNA plates, Biomek FX (Beckman Coulter), Spectramax Plus 384 MTP Reader (Molecular Devices), iEMS Incubator/Shaker (1 mm amplitude) (Thermo Electron Corporation), sealing tape: Nunc (236366), circulating reservoir (Beckman Fx).
[0442] Detergent Composition 3 was initially diluted to 50% in water. This detergent was kept at room temperature and cycled through the circulating reservoir. The iEMS incubators/shakers (Thermo/Labsystems) were pre-set at 43.degree. C. Culture supernatants were diluted into plates containing master dilution buffer to a concentration of .about.20 ppm (master dilution plate). Then, 40 .mu.l of sample from the master dilution plate was added to plates containing 160 .mu.l 150% Detergent Composition 3 to give a final incubation concentration of 4 ppm. The contents were mixed and kept at room temperature and triplicate AAPF assays were performed immediately on these plates and recorded as unstressed reads. The AAPF assay was modified such that 20 .mu.L of sample from the step above was added to 190 .mu.L of suc-AAPF-pNA working solution. The plates were immediately covered with sealing tape and placed in 43.degree. C. iEMS shakers for 30 min at 650 rpm. Following 30 minutes of incubation, triplicate AAPF assays were performed on these stress plates and recorded as stressed reads. The stability of the samples was determined by calculating the ratio of the residual and initial AAPF activity as follows: Residual Activity (%)=[mODmin.sup.-1 stressed]*100/[mODmin.sup.-1 unstressed]. For each sample (variant), the performance index was calculated as described below.
G. LAS/EDTA Stability Assay
[0443] The stability of protease variants in the presence of a representative anionic surfactant (LAS=linear alkylbenzene sulfonate, specifically, sodium dodecylbenzenesulfonate-DOBS) and di-sodium EDTA was measured after incubation under defined conditions and the residual activity was determined using the AAPF assay. The reagents used were dodecyllbenzene sulfonate, sodium salt (DOBS, Sigma No. D-2525), TWEEN.RTM.-80 (Sigma No. P-8074), di-sodium EDTA (Siegfried Handel No. 164599-02), HEPES (Sigma No. H-7523), unstressed buffer: 50 mM HEPES (11.9 g/1)+0.005% TWEEN.RTM.-80, pH 8.0, Stress buffer: 50 mM HEPES (11.9 g/1), 0.1% (w/v) DOBS (1 g/1), 10 mM EDTA (3.36 g/1), pH 8.0, reference protease and protease variant culture supernatants, containing 200-400 .mu.g/ml protein. The equipment used was V- or U-bottom MTPs as dilution plates (Greiner 651101 and 650161, respectively), F-bottom MTPs (Corning 9017) for unstressed and LAS/EDTA buffer as well as for suc-AAPF-pNA plates, Biomek FX (Beckman Coulter), Spectramax Plus 384 MTP Reader (Molecular Devices), iEMS Incubator/Shaker (1 mm amplitude) (Thermo Electron Corporation), and Nunc sealing tape (236366).
[0444] The iEMS incubator/shaker (Thermo/Labsystems) was set at 29.degree. C. Culture supernatants were diluted into plates containing unstressed buffer to a concentration of -25 ppm (master dilution plate). Then, 20 .mu.l of sample from the master dilution plate was added to plates containing 180 .mu.l unstressed buffer to give a final incubation concentration of 2.5 ppm. The contents were mixed and kept at room temperature and an AAPF assay was performed on this plate. Then, 20 .mu.l of sample from the master dilution plate was also added to plates containing 180 .mu.l stress buffer (50 mM HEPES (11.9 g/1), 0.1% (w/v) DOBS (1 g/1), 10 mM EDTA (3.36 g/1), pH 8.0). The solutions were mixed and immediately placed in 29.degree. C. iEMS shaker for 30 min at 400 rpm. Following 30 minutes of incubation, an AAPF assay was performed on the stress plate. The stability of the samples was determined by calculating the ratio of the residual and initial AAPF activity as follows: Residual Activity (%)=[mODmin-1 stressed]*100/[mODmin-1 unstressed]. For each sample (variant), the performance index was calculated as described below.
Performance Index
[0445] The performance index provides a comparison of the performance of a variant (actual value) and a standard or reference protease enzyme (theoretical value) at the same protein concentration. The theoretical values can be calculated using the parameters of a performance dose response curve (i.e. using a Langmuir equation to generate the performance curve) of the standard/reference protease. A performance index (PI) that is greater than 1 (PI>1) identifies a better variant as compared to the standard or reference protease (which may be, e.g., wild-type protease or another protease variant), while a PI of 1 (PI=1) identifies a variant that performs the same as the standard or reference protease, and a PI that is less than 1 (PI<1) identifies a variant that performs worse than the standard or reference protease. Thus, the PI identifies winners (e.g., variants having enhanced proteolytic activity compared to that of the standard/reference protease) as well as variants that may be less desirable for use under certain circumstances (e.g., variants having proteolytic activity lower than the proteolytic activity of the standard/reference protease).
[0446] It is important to note that protease variants of the invention having performance index values lower than that of a reference or standard protease are nevertheless useful in the applications and methods described herein. For example, protease variants of the invention having performance index values lower than that of a reference or standard protease have proteolytic activity and thus are useful in the compositions of the invention, such as, but not limited to, e.g., cleaning compositions (including, but not limited, to, e.g., detergent cleaning compositions) for cleaning a variety of surfaces and items, including, but not limited to, e.g., laundry, fabrics, and dishware, and in personal care applications and compositions as described elsewhere herein; such protease variants are also useful in fabric and home care products and compositions and in non-fabric and home care products and compositions described elsewhere herein and in methods of the invention, including, but not limited, to, e.g., cleaning methods, methods for personal care, etc., described elsewhere herein.
[0447] Various terms set forth below are used to describe the variant: non-deleterious variants have a PI>0.05; deleterious variants have a PI less than or equal to 0.05; combinable variants are those for which the variant has performance index values greater than or equal to 0.2 for at least one property, and >0.05 for all properties. Combinable variants are those that can be combined to deliver proteins with appropriate performance indices for one or more desired properties. These data find use in engineering any subtilisin/subtilase or protease. Even if the subtilase or protease to be engineered has an amino acid different from that of subtilisin BPN' at one or more particular positions, these data find use in identifying amino acid substitutions that alter the desired properties by identifying the best choices for substitutions, including substitutions of the BPN' wild type amino acid.
Example 2
Construction of BPN' Library and Cleaning Performance of BPN' Variants
[0448] a) Description of the BPN'-v3 Expression Cassette Used for Library Construction
[0449] The BPN'-v3 (BPN' protease containing G097A-G128A-Y217Q substitutions) expression cassette used for combinatorial library construction was generated using the BPN' expression cassette, which comprises the aprE-BPN' hybrid leader sequence (i.e., signal sequence), BPN' pro and BPN' mature sequence from B. amyloliquefaciens. The DNA sequence is shown below as SEQ ID NO:1 and encodes the BPN' precursor protein shown below as SEQ ID NO:168.
TABLE-US-00006 (SEQ ID NO: 1) GTGAGAAGCAAAAAATTGTGGATCAGTTTGCTGTTTGCTTTAGCGTTAAT CTTTACGATGGCGTTCGGCAGCACATCCTCTGCCCAGGCGGCAGGGAAAT CAAACGGGGAAAAGAAATATATTGTCGGGTTTAAACAGACAATGAGCACG ATGAGCGCCGCTAAGAAGAAAGATGTCATTTCTGAAAAAGGCGGGAAAGT GCAAAAGCAATTCAAATATGTAGACGCAGCTTCAGCTACATTAAACGAAA AAGCTGTAAAAGAATTGAAAAAAGACCCGAGCGTCGCTTACGTTGAAGAA GATCACGTAGCACATGCGTACGCGCAGTCCGTGCCTTACGGCGTATCACA AATTAAAGCCCCTGCTCTGCACTCTCAAGGCTACACTGGATCAAATGTTA AAGTAGCGGTTATCGACAGCGGTATCGACTCGAGCCATCCAGATCTTAAA GTCGCTGGAGGGGCTTCTATGGTGCCGTCCGAAACAAACCCGTTTCAAGA TAACAATTCTCATGGCACACACGTCGCAGGAACGGTTGCGGCGTTAAACA ATTCTATTGGCGTGCTTGGTGTAGCCCCGTCTGCTTCGCTCTACGCCGTT AAAGTTCTTGGCGCAGACGGATCAGGCCAATACTCATGGATTATCAACGG CATCGAATGGGCCATCGCGAATAACATGGATGTAATCAACATGAGCCTGG GAGGACCAAGCGGCAGTGCGGCACTTAAAGCAGCAGTTGATAAAGCTGTT GCATCTGGTGTCGTCGTAGTAGCGGCAGCTGGGAATGAGGGAACATCCGG ATCATCGAGTACCGTCGGTTATCCAGGCAAGTACCCTTCAGTGATTGCAG TGGGCGCTGTAGACTCTTCAAATCAACGTGCCTCTTTTTCCTCCGTGGGA CCGGAGCTGGATGTCATGGCCCCTGGCGTTTCTATTCAATCGACGCTTCC AGGGAACAAGTATGGTGCGTATAACGGGACTTCCATGGCCTCGCCGCATG TAGCTGGGGCGGCCGCATTGATTCTTTCTAAGCACCCGAACTGGACAAAC ACTCAAGTCCGCAGCAGTTTAGAAAACACCACTACAAAACTTGGTGATTC TTTCTACTATGGAAAAGGGCTGATCAACGTACAGGCGGCAGCTCAG
[0450] In the nucleotide sequence of SEQ ID NO:1, the DNA sequence encoding the mature protease is shown in bold, the nucleotide sequence encoding leader sequence (aprE-BPN' hybrid leader sequence) is shown in standard (non-underlined) text, and the nucleotide sequence encoding the pro sequence (BPN') is underlined. In the amino acid sequence (aprE-BPN' hybrid leader sequence, BPN' pro sequence, and BPN' mature protein sequence) of the BPN' precursor protein set forth in SEQ ID NO:168, the bolded portion indicates the mature BPN' subtilisin protease.
TABLE-US-00007 (SEQ ID NO: 168) VRSKKLWISLLFALALIFTMAFGSTSSAQAAGKSNGEKKYIVGFKQTMST MSAAKKKDVISEKGGKVQKQFKYVDAASATLNEKAVKELKKDPSVAYVEE DHVAHAYAQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLK VAGGASMVPSETNPFQDNNSHGTHVAGTVAALNNSIGVLGVAPSASLYAV KVLGADGSGQYSWIINGIEWAIANNMDVINMSLGGPSGSAALKAAVDKAV ASGVVVVAAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVG PELDVMAPGVSIQSTLPGNKYGAYNGTSMASPHVAGAAALILSKHPNWTN TQVRSSLENTTTKLGDSFYYGKGLINVQAAAQ
[0451] Thus, the amino acid sequence of the mature BPN' subtilisin protease is:
TABLE-US-00008 (SEQ ID NO: 2) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDNNSHGTHVAGTVAALNNSIGVLGVAPSASLYAVKVLGADG SGQYSWIINGIEWAIANNMDVINMSLGGPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAYNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0452] The nucleotide sequence of the mature BPN'-v3 gene is shown below (the signal sequence and propeptide sequence used in the BPN'-v3 expression cassette is the same as that for BPN' shown in SEQ ID NO:1):
TABLE-US-00009 (SEQ ID NO: 3) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTCAAGATAACAATTCTCATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATTCTATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA TCAGGCCAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGAGCCTGGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGGGAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACACTCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAG
[0453] The protein sequence of the mature BPN'-v3 protease variant is shown below (the signal sequence and propeptide sequence used in the BPN'-v3 expression cassette is the same as that for BPN' shown in SEQ ID NO:168):
TABLE-US-00010 (SEQ ID NO: 4) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDNNSHGTHVAGTVAALNNSIGVLGVAPSASLYAVKVLAADG SGQYSWIINGIEWAIANNMDVINMSLGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0454] b) Construction of Combinatorial Library Using pHPLT-BPN'-v3 Plasmid
[0455] The pHPLT-BPN'-v3 plasmid (see FIG. 1) containing the BPN'-v3 expression cassette described above served as template DNA for cloning to provide variants derived from BPN'-v3. The vector pHPLT (FIG. 4 in U.S. Pat. No. 6,566,112) contains the B. licheniformis LAT promoter ("Plat"); a sequence encoding the LAT signal peptide ("preLAT"). Additional plasmid elements from plasmid pUB110 disclosed in McKenzie et al., Plasmid 15(2): 93-103 (1986): "ori-pUB" is the origin of replication from pUB110; "neo" is the neomycin/kanamycin resistance gene from pUB110; "Terminator" is the transcriptional terminator from B. licheniformis amylase.
[0456] A combinatorial DNA library was synthesized at DNA 2.0 and delivered as individual ligation reactions. In some instances for efficient transformation of B. subtilis, the DNA from the ligation reaction mixtures was amplified by rolling circle amplification (RCA) using the Illustra Templiphi kit (GE Healthcare). The reaction was performed according to the manufacturer's protocol. One microliter of ten-fold diluted amplified DNA was used to transform 50 .mu.L of competent B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). The transformation mixture was shaken at 37.degree. C. for 1 hour. Ten micro-liter aliquots of the transformation mixture were plated on skim milk (1.6%) Luria agar plates supplemented with 10 .mu.g/ml of neomycin (Teknova).
[0457] The transformants that formed halos on the skim milk plates were picked into microtiter plates containing 150 .mu.l Luria broth (LB) medium supplemented with 10 .mu.g/ml neomycin. Plates were grown overnight at 37.degree. C. with 250-300 rpm shaking and 70-80% humidity using Enzyscreen lids for microtiter plates (Enzyscreen). Using a 96 pin replicating tool, (Enzyscreen) the overnight culture plate was used to inoculate a new microtiter plate containing 180 .mu.l of MBD medium (a MOPS based defined medium) with 2.5 .mu.g/ml neomycin. MBD medium was prepared essentially as known in the art (see Neidhardt et al., J. Bacteriol. 119:736-747 [1974]), except that NH.sub.4Cl.sub.2, FeSO.sub.4, and CaCl.sub.2 were omitted from the base medium, 3 mM K.sub.2HPO.sub.4 was used, and the base medium was supplemented with 60 mM urea, and 100 ml of a solution made of 210 g/L glucose, and 350 g/L maltodextrin. 1 g/L of BD Bacto Yeast Extract was added and the pH was adjusted to 7.4 with KOH. The micronutrients were made up as a 100.times. stock solution containing in one liter, 400 mg FeSO.sub.4.7H.sub.2O, 100 mg MnSO.sub.4.H.sub.2O, 100 mg ZnSO.sub.4.7H.sub.2O, 50 mg CuCl.sub.2.2H.sub.2O, 100 mg CoCl.sub.2.6H.sub.2O, 100 mg NaMoO.sub.4.2H.sub.2O, 100 mg Na.sub.2B.sub.4O.sub.7.10H.sub.2O, 10 ml of 1M CaCl.sub.2, and 10 ml of 0.5 M sodium citrate. The MBD medium containing microtiter plates were grown for 64 hours at 37.degree. C., 250-300 rpm, and 70-80% humidity using Enzyscreen lids (Enzyscreen) for protease variant expression. The next day, cultures were filtered through a micro-filter plate (0.22 um; Millipore) and the resulting filtrates containing protease variants were used for biochemical analysis.
[0458] The protease variants were tested for cleaning performance using a BMI microswatch assay in Detergent Composition 1 at 16.degree. C. and pH 8 and BMI microswatch assay in Detergent Composition 2 at 16.degree. C. and pH 8. Protein content was determined using the TCA assay. Assays were performed as described in Example 1 and Performance Indices were calculated relative to BPN'-v3 (with a PI value of 1.0).
[0459] The following BPN' subtilisin protease variant was determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2 to about 5, or from greater than 1.0 to about 5 relative to BPN'-v3 (SEQ ID NO:4) in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising the set of amino acid substitutions G097A-G128A-P210S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variant has a PI value of 1.1 relative to BPN'-v3 in this BMI microswatch cleaning assay, and enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) in this assay, the variant having an amino acid sequence comprising amino acid substitutions G097A-G128A-P210S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Also included is a protease variant having enhanced proteolytic activity compared to SEQ ID NO:2 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising amino acid substitutions G097A-G128A-P210S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value greater than that of BPN' (SEQ ID NO:2) and/or BPN'-v3 in a BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to SEQ ID NO:2, wherein the variant comprises substitutions X097A-X128A-X210S-X217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2, and optionally wherein the variant comprises at least one substitution selected from the group of G097A, G128A, P210S, and Y217Q. Such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising such protease variant and methods for cleaning utilizing such variant as described in greater detail elsewhere herein.
[0460] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q, G097A-G128A-E156S-P210S-Y217Q, G097A-G128A-P210S-Y217Q-N218A, G097A-G128A-P210S-Y217Q-N218S, and G097A-Y104F-G128A-E156S-P210I-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) or a PI value of about 1.0 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value of about 1.0 compared to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2, wherein the variant comprises at least one substitution selected from the group of X097A, X104F, X128A, X156A/S, X210I/S, X217Q, X218A/S, and optionally at least one substitution selected from the group of G097A, Y104F, G128A, E156A/S, P210I/S, Y217Q, and N218A/S, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0461] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-E156A-P210S-Y217Q-N218S and G097A-G128A-Y217Q-N218A, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0462] The following BPN' variants were determined to have a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-P210S-Y217Q-N218A and G097A-G128A-P210S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value of greater than 1, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9 to about 5 compared to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2, wherein the variant comprises at least one substitution selected from the group of X097A, X104F, X128A, X156A/S, X210I/S, X217Q, X218A/S, and optionally at least one substitution selected from the group of G097A, Y104F, G128A, E156A/S, P210I/S, Y217Q, and N218A/S, wherein amino acid positions of the variant are numbered by correspondence with position of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0463] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q (i.e., BPN'-v3), G097A-G128A-E156S-P210S-Y217Q, and G097A-G128A-P210S-Y217Q-N218S, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity and enhanced proteolytic activity compared to BPN' in this assay. The invention includes a protease variant having proteolytic activity, PI value of 1.0 relative to BPN'-v3, and/or enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0464] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-E156A-P210S-Y217Q-N218S, G097A-G128A-Y217Q-N218A, and G097A-Y104F-G128A-E156S-P210I-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 3
Generation of Combinatorial Libraries and Cleaning Performance of Variants of BPN'-v3+S78N
[0465] a) Description of BPN'-v3+S78N Variant and Synthetic Gene Sequences Derived from this Variant
[0466] Gene Oracle synthesized and cloned eight genes into the pHPLT-BPN'-v3+S78N (BPN'-S78N-G97A-G128A-Y217Q) parent plasmid (see FIG. 2). Some of these genes were used as templates (parents) to create combinatorial libraries. The BPN'-v3+S78N variant was generated using standard molecular biology methods known in the art. The nucleotide sequence encoding the BPN'-v3+S78N variant is shown below:
TABLE-US-00011 (SEQ ID NO: 7) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATTGATTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTCAAGATAACAATTCTCATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA TCAGGCCAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGAGCCTGGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGGGAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACACTCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAG
[0467] The amino acid sequence of the BPN'-v3+S78N variant is shown below:
TABLE-US-00012 (SEQ ID NO: 8) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDNNSHGTHVAGTVAALNNNIGVLGVAPSASLYAVKVLAADG SGQYSWIINGIEWAIANNMDVINMSLGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0468] The nucleotide and protein sequences of genes GcM90-96, and GcM100 are shown below. The nucleotide sequence of synthesized gene GcM90 is:
TABLE-US-00013 (SEQ ID NO: 9) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGGGAGAAACAAACCCGTTTCAAGATAACAATTCTCATGGCACACA CGCAGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA TCAGCACAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGAGCCTGGGAGCAACAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATACACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGGGAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACACTCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0469] The amino acid sequence of GcM90 is provided below:
TABLE-US-00014 (SEQ ID NO: 10) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPGETNPFQDNNSHGTHAAGTVAALNNNIGVLGVAPSASLYAVKVLAADG SAQYSWIINGIEWAIANNMDVINMSLGATSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNTRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0470] The nucleotide sequence of synthesized gene GcM91 is provided below:
TABLE-US-00015 (SEQ ID NO: 11) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTGTCGATAACAATTCTCATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA TCAGGCCAATACTCATGGATTGTCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGAGCCTGGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTCAAGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGCAAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACACTCAAGTCCGCAGCAGTTTA GAACAAACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0471] The amino acid sequence of GcM91 is provided below:
TABLE-US-00016 (SEQ ID NO: 12) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFVDNNSHGTHVAGTVAALNNNIGVLGVAPSASLYAVKVLAADG SGQYSWIVNGIEWAIANNMDVINMSLGAPSGSAALKAAVDKAVASGQVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPANKYGAQNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL EQTTTKLGDSFYYGKGLINVQAAAQ
[0472] The nucleotide sequence of synthesized gene GcM92 is provided below:
TABLE-US-00017 (SEQ ID NO: 13) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTCAAGATGCAAATTCTCATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGGAAGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA TCAGGCCAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATCAGCCTGGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGACCTTCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGGGAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCGCACCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACACTCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0473] The amino acid sequence of GcM92 is provided below:
TABLE-US-00018 (SEQ ID NO: 14) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDANSHGTHVAGTVAALNNNIGVLGVAPEASLYAVKVLAADG SGQYSWIINGIEWAIANNMDVINISLGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGPSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMAAPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0474] The nucleotide sequence of synthesized gene of GcM93 is provided below:
TABLE-US-00019 (SEQ ID NO: 15) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTCAAGATAACCAATCTCATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACAAC TCAGGCCAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGGCACTGGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACAGATGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGGGAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGTCATGGACAAACACTCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0475] The amino acid sequence of GcM93 is provided below:
TABLE-US-00020 (SEQ ID NO: 16) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDNQSHGTHVAGTVAALNNNIGVLGVAPSASLYAVKVLAADN SGQYSWIINGIEWAIANNMDVINMALGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTDGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMASPHVAGAAALILSKHPSWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0476] The nucleotide sequence of synthesized gene GcM94 is provided below:
TABLE-US-00021 (SEQ ID NO: 17) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTCAAGATAACAATACACATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA GCAGGCCAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGAGCGTCGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTACAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGGGAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACAACCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0477] The amino acid sequence of GcM94 is provided below:
TABLE-US-00022 (SEQ ID NO: 18) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDNNTHGTHVAGTVAALNNNIGVLGVAPSASLYAVKVLAADG AGQYSWIINGIEWAIANNMDVINMSVGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSTNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMASPHVAGAAALILSKHPNWTNNQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0478] The nucleotide sequence of synthesized gene GcM 95 is provided below:
TABLE-US-00023 (SEQ ID NO: 19) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCTGGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGGGAGAAACAAACCCGTTTGTCGATGCACAAACACATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGGAAGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACAAC GCAGCACAATACTCATGGATTGTCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGAGCCTGGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGGGAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACACTCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0479] The amino acid sequence of GcM 95 is provided below:
TABLE-US-00024 (SEQ ID NO: 20) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHLDLKVAGGASM VPGETNPFVDAQTHGTHVAGTVAALNNNIGVLGVAPEASLYAVKVLAADN AAQYSWIVNGIEWAIANNMDVINMSLGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0480] The nucleotide sequence of synthesized gene GcM96 is provided below:
TABLE-US-00025 (SEQ ID NO: 21) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTCAAGATAACAATTCTCATGGCACACA CGTCGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA TCAGGCCAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGAGCCTGGGAGCAACAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTCAAGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACAGATGGACCTTCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTACAA ATACACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGCAAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCGCACCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGTCATGGACAAACAACCAAGTCCGCAGCAGTTTA GAACAAACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0481] The amino acid sequence of GcM96 is provided below:
TABLE-US-00026 (SEQ ID NO: 22) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDNNSHGTHVAGTVAALNNNIGVLGVAPSASLYAVKVLAADG SGQYSWIINGIEWAIANNMDVINMSLGATSGSAALKAAVDKAVASGQVVV AAAGNEGTDGPSSTVGYPGKYPSVIAVGAVDSTNTRASFSSVGPELDVMA PGVSIQSTLPANKYGAQNGTSMAAPHVAGAAALILSKHPSWTNNQVRSSL EQTTTKLGDSFYYGKGLINVQAAAQ
[0482] The nucleotide sequence of synthesized gene GcM100 is provided below:
TABLE-US-00027 (SEQ ID NO: 23) GCGCAGTCCGTGCCTTACGGCGTATCACAAATTAAAGCCCCTGCTCTGCA CTCTCAAGGCTACACTGGATCAAATGTTAAAGTAGCGGTTATCGACAGCG GTATCGACTCGAGCCATCCAGATCTTAAAGTCGCTGGAGGGGCTTCTATG GTGCCGTCCGAAACAAACCCGTTTCAAGATAACAATTCTCATGGCACACA CGCAGCAGGAACGGTTGCGGCGTTAAACAATAATATTGGCGTGCTTGGTG TAGCCCCGTCTGCTTCGCTCTACGCCGTTAAAGTTCTTGCAGCAGACGGA TCAGCACAATACTCATGGATTATCAACGGCATCGAATGGGCCATCGCGAA TAACATGGATGTAATCAACATGGCACTGGGAGCACCAAGCGGCAGTGCGG CACTTAAAGCAGCAGTTGATAAAGCTGTTGCATCTGGTGTCGTCGTAGTA GCGGCAGCTGGGAATGAGGGAACATCCGGATCATCGAGTACCGTCGGTTA TCCAGGCAAGTACCCTTCAGTGATTGCAGTGGGCGCTGTAGACTCTTCAA ATCAACGTGCCTCTTTTTCCTCCGTGGGACCGGAGCTGGATGTCATGGCC CCTGGCGTTTCTATTCAATCGACGCTTCCAGCAAACAAGTATGGTGCGCA AAACGGGACTTCCATGGCCTCGCCGCATGTAGCTGGGGCGGCCGCATTGA TTCTTTCTAAGCACCCGAACTGGACAAACACTCAAGTCCGCAGCAGTTTA GAAAACACCACTACAAAACTTGGTGATTCTTTCTACTATGGAAAAGGGCT GATCAACGTACAGGCGGCAGCTCAGTAA
[0483] The amino acid sequence of GcM100 is provided below:
TABLE-US-00028 (SEQ ID NO: 24) AQSVPYGVSQIKAPALHSQGYTGSNVKVAVIDSGIDSSHPDLKVAGGASM VPSETNPFQDNNSHGTHAAGTVAALNNNIGVLGVAPSASLYAVKVLAADG SAQYSWIINGIEWAIANNMDVINMALGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPANKYGAQNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0484] b) Construction of Combinatorial Libraries CG1-CG5 and CG8 Using the Synthetic Genes GcM90-94 and GcM100
TABLE-US-00029 TABLE 3-1 List of Possible Substitutions Introduced and Primers Used for the Construction of Combinatorial Libraries CG1-CG5 and CG8 Synthesized Genes (template or Library Substitutions Primer parent) Name Introduced Name Primer Sequence GcM90 CG1 G53S 1 S53 f /5Phos/CTTCTATGGTGCCGTCCGAAACA AACCCGTTTCAAG (SEQ ID NO: 25) A68V 1 V68 f /5Phos/TCATGGCACACACGTCGCAGGAA CGGTTGCGGCG (SEQ ID NO: 26) A102G 1 G102 f /5Phos/AGCAGACGGATCAGGCCAATACT CATGGATTATCAAC (SEQ ID NO: 27) T129P 1 P129 f /5Phos/TGAGCCTGGGAGCACCAAGCGGC AGTGCGGCACTTAAAG (SEQ ID NO: 28) T185Q 1 Q185 f /5Phos/TAGACTCTTCAAATCAACGTGCC TCTTTTTCCTCCGTG (SEQ ID NO: 29) GcM91 CG2 V59Q 2 Q59 f /5Phos/GAAACAAACCCGTTTCAAGATAA CAATTCTCATG (SEQ ID NO: 30) V108I 2 I108 f /5Phos/ATACTCATGGATTATCAACGGCA TCGAATGGGCCATC (SEQ ID NO: 31) Q147V 2 V147 f /5Phos/TGTTGCATCTGGTGTCGTCGTAGT AGCGGCAGCTGG (SEQ ID NO: 32) A211G 2 G211 f /5Phos/ATCGACGCTTCCAGGGAACAAGT ATGGTGCGCAAAAC (SEQ ID NO: 33) Q252N 2 N252 f /5Phos/CAGCAGTTTAGAAAACACCACTA CAAAACTTGGTG (SEQ ID NO: 34) GcM92 CG3 A61N 3 N61 f /5Phos/CAAACCCGTTTCAAGATAACAAT TCTCATGGCACACAC (SEQ ID NO: 35) E87S 3 S87 f /5Phos/TTGGTGTAGCCCCGTCTGCTTCGC TCTACGCCGTTAAAG (SEQ ID NO: 36) I124M 3 M124 f /5Phos/TGGATGTAATCAACATGAGCCTG GGAGCACCAAGCG (SEQ ID NO: 37) P161S 3 S161 f /5Phos/AGGGAACATCCGGATCATCGAGT ACCGTCGGTTATCCAG (SEQ ID NO: 38) A224S 3 S224 f /5Phos/GACTTCCATGGCCTCGCCGCATG TAGCTGGGGCGGC (SEQ ID NO: 39) GcM93 CG4 Q62N 4 N62 f /5Phos/GTTTCAAGATAACAATTCTCATG GCACACACGTCGC (SEQ ID NO: 40) N100G 4 G100 f /5Phos/GTTCTTGCAGCAGACGGATCAGG CCAATACTCATG (SEQ ID NO: 41) A125S 4 S125 f /5Phos/ATGTAATCAACATGAGCCTGGGA GCACCAAGCGGCAG (SEQ ID NO: 42) D159S 4 S159 f /5Phos/GGAATGAGGGAACATCCGGATCA TCGAGTACCGTCGG (SEQ ID NO: 43) S240N 4 N240 f /5Phos/CTTTCTAAGCACCCGAACTGGAC AAACACTCAAGTCCG (SEQ ID NO: 44) GcM94 CG5 T63S 5 S63 f /5Phos/TCAAGATAACAATTCTCATGGCA CACACGTCGCAGG (SEQ ID NO: 45) A101S 5 S101 f /5Phos/TGCAGCAGACGGATCAGGCCAAT ACTCATGGATTATC (SEQ ID NO: 46) V126L 5 L126 f /5Phos/AATCAACATGAGCCTGGGAGCAC CAAGCGGCAGTG (SEQ ID NO: 47) T183S 5 S183 f /5Phos/CGCTGTAGACTCTTCAAATCAAC GTGCCTCTTTTTCC (SEQ ID NO: 48) N244T 5 T244 f /5Phos/GAACTGGACAAACACTCAAGTCC GCAGCAGTTTAG (SEQ ID NO: 49) GcM100 CG8 A68V 1 V68 f /5Phos/TCATGGCACACACGTCGCAGGAA CGGTTGCGGCG (SEQ ID NO: 50) A102G 1 G102 f /5Phos/AGCAGACGGATCAGGCCAATACT CATGGATTATCAAC (SEQ ID NO: 51) A211G 2 G211 f /5Phos/ATCGACGCTTCCAGGGAACAAGT ATGGTGCGCAAAAC (SEQ ID NO: 52) A125S 4 S125 f /5Phos/ATGTAATCAACATGAGCCTGGGA GCACCAAGCGGCAG (SEQ ID NO: 53)
[0485] Each synthesized gene was built into the pHPLT-BPN'-S78N-G97A-G128A-Y217Q parent molecule. Resulting plasmids containing the six synthesized genes GcM90-94, and GcM100 served as templates to make combinatorial libraries at the respective positions (Table 3-1). Two additional genes, GcM95 and GcM96, were also synthesized for analysis, but did not serve as parental DNA for libraries. These genes each have nine mutations on top of the pHPLT-BPN'-S78N-G97A-G128A-Y217Q parent molecule.
[0486] The parent plasmids (template DNA) containing the synthetic genes GcM90-94, and GcM100 were methylated were methylated using two micrograms of DNA and methylase (NEB), according to the NEB protocol. Methylated DNA was then purified using DNA Clean and Concentrator kit (Zymo Research). Combinatorial libraries CG1-5 and CG8 were made using a QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis kit ("QCMS kit"; Stratagene) following the manufacturer's protocol (see Table 3-1 for respective template and primer combinations), with the exception of libraries CG3 and CG4, which used 86.5 ng of each primer in place of the 50 ng suggested in the protocol. All primers used for introducing the desired substitutions in each library are listed in Table 3-1. They were synthesized and provided by Integrated DNA Technologies. After the QCMS reactions were completed for each library, the template DNA was digested by the addition of 0.5-1 .mu.l DpnI (from the QCMS kit) and incubated at 37.degree. C. for 1-4 hours, followed by another addition of 0.5-1 .mu.l DpnI and another incubation at 37.degree. C. for 1-4 hours. For efficient transformation of B. subtilis, DNA from the QCMS reaction mixtures were amplified before transformation and transformants grown as described in Example 2.
[0487] Additional variants of BPN'-v3+S78N were produced by DNA2.0. The following substitutions were introduced individually into the BPN'-v3+S78N parent molecule: Q59G, N62Q, V68A, S89Y, A92G, I108V, I115V, M124T, P129L, A138T, V147L, S161P, Y167A, P172V, G211T, L267V, and A273S.
[0488] All of the combinatorial library variants described above and the variants synthesized at DNA2.0 were tested for cleaning performance using a BMI microswatch assay in Detergent Composition 1 at 16.degree. C. and pH 8 and BMI microswatch assay in Detergent Composition 2 at 16.degree. C. and pH 8. Protein content was determined using the TCA assay. Assays were performed as described in Example 1 and Performance Indices were calculated relative to BPN'-v3 (with a PI value of 1.0).
[0489] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of S063T-S078N-G097A-S101A-G128A-S183T-Y217Q-T244N, N061A-S078N-G097A-G128A-Y217Q-S224A, S053G-S078N-G097A-G128A-P129T-Q185T-Y217Q, S063T-S078N-G097A-S101A-G128A-S183T-Y217Q, S063T-S078N-G097A-S101A-G128A-Y217Q, S063T-S078N-G097A-S101A-G128A-Y217Q-T244I, and S078N-G097A-G128A-P129T-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' and BPN'-v3 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0490] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2, wherein the variant comprises at least one substitution selected from the group of X040E, X053G, X059V, X061A, X062H/Q, X068A, X078N, X087E, X101A, X102A, X108V, X124I, X125A, X126V, X129T, X147Q, X159D, X183T, X185T, X211A, X224A, X244I/N, X252Q, and X274D, and optionally at least one substitution selected from the group of X040E, X053G, X059V, X061A, X062H/Q, X068A, X078N, X087E, X101A, X102A, X108V, M124I, S125A, L126V, P129T, V147Q, S159D, S183T, Q185T, G211A, S224A, T244I/N, N252Q, and A274D, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0491] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q, N061A-S078N-S087E-G097A-G128A-Y217Q-S224A, Q059V-S078N-G097A-G128A-G211A-Y217Q, Q059V-S078N-G097A-G128A-V147Q-Y217Q, Q059V-S078N-G097A-G128A-Y217Q, Q059V-S078N-G097A-I108V-G128A-Y217Q-N252Q, S053G-S078N-G097A-G128A-P129T-Y217Q, S078N-G097A-G128A-G211A-Y217Q, S078N-G097A-G128A-Q185T-Y217Q, S078N-G097A-G128A-V147Q-Y217Q, S078N-G097A-G128A-Y217Q, S078N-G097A-G128A-Y217Q-S224A, and S078N-G097A-G128A-Y217Q-S224A-A274D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0492] The following BPN' variant was determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising the set of amino acid substitutions S078N-G097A-I108V-G128A-V147Q-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising said set of amino acid substitutions above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0493] The following BPN' variants were determined to a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of Q059V-S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q, S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-Y217Q-N252Q, and S078N-S087E-G097A-M124I-G128A-Y217Q-S224A, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0494] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of S063T-S078N-G097A-S101A-G128A-S183T-Y217Q, S063T-S078N-G097A-S101A-G128A-S183T-Y217Q-T244N, S063T-S078N-G097A-S101A-G128A-Y217Q, and S063T-S078N-G097A-S101A-G128A-Y217Q-T244I, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than about 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0495] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q, N061A-S078N-G097A-G128A-Y217Q-S224A, N061A-S078N-S087E-G097A-G128A-Y217Q-S224A, Q059V-S078N-G097A-G128A-G211A-Y217Q, Q059V-S078N-G097A-G128A-V147Q-Y217Q, Q059V-S078N-G097A-G128A-Y217Q, Q059V-S078N-G097A-I108V-G128A-Y217Q-N252Q, S053G-S078N-G097A-G128A-P129T-Q185T-Y217Q, S053G-S078N-G097A-G128A-P129T-Y217Q, S078N-G097A-G128A-G211A-Y217Q, S078N-G097A-G128A-P129T-Y217Q, S078N-G097A-G128A-Q185T-Y217Q, S078N-G097A-G128A-Y217Q, S078N-G097A-G128A-Y217Q-S224A, and S078N-G097A-G128A-Y217Q-S224A-A274D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0496] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of S078N-G097A-G128A-V147Q-Y217Q and S078N-G097A-I108V-G128A-V147Q-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including cleaning compositions, comprising at least one such variant and methods for cleaning an item or surface in need of cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0497] The following BPN' variants were determined to a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of Q059V-S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q, S078N-G097A-I108V-G128A-V147Q-G211A-Y217Q-N252Q, S078N-G097A-I108V-G128A-V147Q-Y217Q-N252Q, and S078N-S087E-G097A-M124I-G128A-Y217Q-S224A, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including cleaning compositions, comprising at least one such variant and methods for cleaning an item or surface in need of cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 4
Generation and Cleaning Performance of BPN' Variants
Generation of BPN' Variants LC1-LC4 Via QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis
[0498] BPN' variants were constructed from different parental plasmids using QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis kits. The parental plasmids (Table 4-1) were methylated using a NEB Dam Methylase Kit in a reaction containing 77.5 .mu.L H.sub.2O+10 .mu.L Buffer 10.times.+0.25 .mu.L SAM+2 .mu.L DAM methylase+10 .mu.L miniprep DNA (.about.150 ng/.mu.L) at 37.degree. C. overnight. The methylated plasmid DNA was purified using a QIAGEN.RTM. PCR purification kit. QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis reactions were set up for each of the DNA templates in a reaction mix containing 2.5 .mu.L Buffer 5.times.+0.5 .mu.L primer 1 (25 .mu.M)+0.5 .mu.L primer 2 (25 .mu.M)+1 .mu.L dNTP's+1 .mu.L enzyme blend+18 .mu.L H.sub.2O+1.5 .mu.L DNA. The PCR program used was: 95.degree. C. for 1 min; (95.degree. C. for 1 min, 53.degree. C. for 1 min, 65.degree. C. for 9:39 min).times.29 cycles; 65.degree. C. for 10 min, 4.degree. C. hold. Primer sequences are shown in Table 4-2. In all reactions, PCR was performed using a MJ Research PTC-200 Peltier thermal cycler. Parental DNA from the PCR samples was removed by addition of 1 .mu.L of DpnI to QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis reactions at 37.degree. C. overnight. To increase transformation frequency, the DpnI-digested reactions were amplified using rolling circle amplification (RCA) using the Illustra TempliPhi kit according to the manufacturer's protocol. B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo) were transformed with 1 .mu.L each of the RCA reaction and the transformed cells were plated onto LA+1.6% skim milk plates containing 10 ppm neomycin and incubated at 37.degree. C. overnight. Colonies from overnight growth were selected to perform colony PCR for sequencing using "puReTaq Ready-To-Go PCR Beads" (Amersham). The PCR and sequencing primers used were pHPLT F1 (/5PHOS/TACATATGAGTTATGCAGTTTG (SEQ ID NO:54)) and pHPLT seq R1 (/5PHOS/TTATCCTTTACCTTGTCTC (SEQ ID NO:55)). Clones with appropriate sequences were frozen. BPN' variant proteins were produced by growing B. subtilis transformants in 96 well microtiter plates at 37.degree. C. for 68 hours in a MOPS based medium containing urea as described in Example 2.
TABLE-US-00030 TABLE 4-1 Parental Plasmids and Primers Used for Generation of BPN' Variants LC1-LC4 Mutations Primers Parental Plasmid Introduced Used BPN'-G097A-G128A-Y217Q-S024G- A128S A128Sf, N025G-N061P-S101N (termed LC1) A128Sr BPN'-G097A-G128A-Y217Q-S053G- A128S A128Sf, N061P-S101N-V203Y (termed LC2) A128Sr BPN'-G097A-G128A-Y217Q-S024G- A128S A128Sf, N025G-S053G-T055P-N061P-S101N- A128Sr V203Y (termed LC3) BPN'-G097A-G128A-Y217Q-S024G- P55T P55Tf, N025G-S053G-T055P-N061P-S101N- P55Tr V203Y (termed LC4)
TABLE-US-00031 TABLE 4-2 Sequences of Primers Used for QUIKCHANGE .RTM. Multi Site-Directed Mutagenesis Reactions to Make BPN' variants LC1-LC4 Primer Name Primer Sequence (5' to 3') A128Sf /5Phos/CAACATGAGCCTGGGATCACCAAGCGGCAGTGCGG (SEQ ID NO: 56) A128Sr /5Phos/CCGCACTGCCGCTTGGTGATCCCAGGCTCATGTTG (SEQ ID NO: 57) P55Tf /5Phos/CTATGGTGCCGGGCGAAACAAACCCGTTTCAAGATCCG (SEQ ID NO: 58) P55Tr /5Phos/CGGATCTTGAAACGGGTTTGTTTCGCCCGGCACCATAG (SEQ ID NO: 59)
Generation of Additional BPN' Variants LC5-LC37
[0499] An additional 33 BPN' variants termed successively LC5 through LC37 were produced by DNA 2.0 using the BPN' nucleic acid as the parent gene contained in the expression plasmid pHPLT-BPN' partial opt (see FIG. 3). LC5 through LC37 BPN' variants are as follows, respectively: BPN-P52L-V68A-G97A-I111V, BPN-I111V-M124V-Y167A-Y217Q, BPN'-Y104N-G128A-Y217Q, BPN'-M124V-Y167A-Y217Q, BPN-I111V-M124V-Y217Q, BPN'-P52L-V68A-G97A, BPN-G97A-I111V-M124V, BPN'-V68A-A92G-G97A, BPN'-G97A-I111V-M124V-Y167A-Y217Q, BPN-P52L-V68A-I111V-Y217Q, BPN-P52L-V68A-I111V, BPN-V68A-A92G-I111V, BPN-P52L-V68A-G97A-I111V-Y217Q, BPN-V68A-G97A-I111V, BPN-G97A-I111V-Y217Q, BPN-G97A-I111V-M124V-Y167A, BPN-S89Y-I111V-M124V, BPN-V68A-S89Y-I111V, BPN'-V68A-A92G-Y217Q, BPN-I111V-Y167A-Y217Q, BPN-G97A-I111V-Y167A-Y217Q, BPN-G97A-I111V-M124V-Y217Q, BPN'-V68A-I111V-Y167A-Y217Q, BPN-I111V-G128A-Y217Q, BPN'-G97A-M124V-Y217Q, BPN'-V68A-Y167A-Y217Q, BPN-I111V-M124V-Y167A, BPN-N62Q-G97A-I111V, BPN'-G97A-M124V-Y167A-Y217Q, BPN'-G97A-L126A-Y217Q, BPN'-V68A-I111V-Y217Q, BPN'-S89Y-M124V-Y217Q, and BPN'-L96T-G97A-Y217Q. Plasmid pHPLT-BPN' partial opt was also created by DNA 2.0.
[0500] Transformants were picked into microtiter plates and grown as described in Example 2. The variants were assayed for cleaning performance using a BMI microswatch assay in Detergent Composition 2 at 16.degree. C. and pH 8. Protein content was determined using the TCA assay. The assays were performed as described in Example 1 and the Performance Indices were calculated relative to BPN'-v3 (with a PI value of 1.0).
[0501] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-I111V-M124V-Y217Q, G097A-I111V-Y167A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128A-V203Y-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128S-V203Y-Y217Q, and V068A-A092G-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2, wherein the variant comprises at least one substitution selected from the group of X024G, X025G, X052L, X053G, X055P, X061P, X062Q, X068A, X089Y, X092G, X096T, X097A, X101N, X104N, X111V, X124V, X126A, X128A/S, X167A, X203Y, and X217Q, and optionally at least one substitution selected from the group of S024G, N025G, P052L, S053G, T055P, N061P, N062Q, V068A, S089Y, A092G, L096T, G097A, S101N, Y104N, I111V, M124V, L126A, G128A/S, Y167A, V203Y, and Y217Q, and wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0502] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q, G097A-G128S-Y217Q, G097A-I111V-Y217Q, I111V-G128A-Y217Q, I111V-M124V-Y167A, I111V-M124V-Y217Q, L096T-G097A-Y217Q, N062Q-G097A-I111V, S053G-N061P-G097A-S101N-G128S-V203Y-Y217Q, S089Y-M124V-Y217Q, and V068A-I111V-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a greater PI value than that of BPN', and a PI value of about 1.0 compared to BPN'-v3 in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0503] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-I111V-M124V, G097A-L126A-Y217Q, G097A-M124V-Y217Q, I111V-Y167A-Y217Q, M124V-Y167A-Y217Q, P052L-V068A-G097A, S089Y-I111V-M124V, V068A-A092G-G097A, V068A-A092G-I111V, V068A-G097A-I111V, V068A-S089Y-I111V, and Y104N-G128A-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0504] The following BPN' variants were determined to a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 2 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-M124V-Y167A-Y217Q, V068A-Y167A-Y217Q, G097A-I111V-M124V-Y167A, I111V-M124V-Y167A-Y217Q, V068A-I111V-Y167A-Y217Q, G097A-I111V-M124V-Y167A-Y217Q, and P052L-V068A-I111V, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Such variants have proteolytic activity. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 5
Cleaning Performance of BPN' Variants
[0505] Variants based on parent BPN' were made by DNA 2.0. The variants were grown as described in Example 2 and tested for cleaning performance on BMI microswatch assay in Detergent Composition 1 at 16.degree. C. and pH 8, BMI microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8, and egg microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8. The protein content was determined using the TCA assay. The assays were performed as described in Example 1 and the Performance Indices were calculated relative to BPN'-v3 (with a PI value of 1.0).
[0506] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128S-Y217Q, and S024G-N025G-S053G-N061P-S078N-G097A-S101N-I111V-G128S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0507] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q, G097A-G128A-P210S-Y217Q, G097A-G128S-P210S-Y217Q, G097A-I111V-M124I-Y217Q, G097A-I111V-M124V-P210S-Y217Q, G097A-N123Q-P210S-Y217Q, G097A-N123Q-Y217Q, N061P-G097A-G128A-P210S-Y217Q, N061P-G097A-G128S-Y217Q, N061P-G097A-I111V-M124V-Y217Q, N061P-G097A-N123Q-Y217Q, N061P-G097A-S101N-I111V-M124V-Y217Q, N061P-G097A-S101N-N123Q-Y217Q, N061P-G102A-P129S-Y217Q, N061P-N062Q-G097A-G100N-S101N-Y217Q, N061P-N062Q-G097A-G100N-Y217Q, N061P-N062Q-G097A-G100Q-P210S-Y217Q, N061P-N062Q-G097A-I111V-Y217Q, N061P-N062Q-G097A-S101N-I111V-Y217Q, N061P-S078N-G097A-I111V-M124I-Y217Q, N061P-S078N-G102A-I111V-P129S-Y217Q, N062Q-G097A-I111V-P210S-Y217Q, N062Q-G097A-I111V-Y217Q, N062Q-S078N-G097A-I111V-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-N123Q-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, S024G-N025G-S053G-N061P-S101N-G102A-P129S-Y217Q, S053G-N061P-G097A-G128S-Y217Q, S053G-N061P-G097A-M124I-Y217Q, S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, S053G-N061P-G102A-P129S-P210S-Y217Q, S053G-N061P-G102A-P129S-Y217Q, S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, S053-N061P-S101N-G102A-P129S-Y217Q, S053G-S078N-G097A-I111V-G128S-Y217Q, S078N-G097A-G128S-Y217Q, and S078N-G097A-I111V-M124V-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0508] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of N061P-G097A-M124I-Y217Q, N061P-G097A-M124V-Y217Q, N061P-N062Q-G097A-G100D-Y217Q, N061P-N062Q-G097A-G100Q-S101N-Y217Q, N061P-N062Q-G097A-G100Q-Y217Q, N061P-N062Q-G100N-G102A-Y217Q, N061P-N062Q-S078N-G097A-G100N-I111V-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-M124I-Y217Q, S053G-N061P-G097A-S101N-M124I-Y217Q, and S053G-N061P-G097A-S101N-N123Q-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0509] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 or 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2, wherein the variant comprises at least one substitution selected from the group of X024G, X025G, X053G, X061P, X062Q, X078N, X097A, X100D/N/Q, X101N, X102A, X111V, X123A/Q/V, X124I/V, X128A/S, X129S, X210S, X217Q, and optionally at least one substitution selected from the group of S024G, N025G, S053G, N061P, N062Q, S078N, G097A, G100D/N/Q, S101N, G102A, I111V, N123A/Q/V, M124I/V, G128A/S, P129S, P210S, and Y217Q, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0510] The following BPN' variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-N123A-Y217Q, G097A-N123V-Y217Q, N061P-G102A-G128S-Y217Q, N061P-S101N-G102A-G128S-Y217Q, Y217Q, S078N-G097A-I111V-N123Q-Y217Q, and G102A-N123Q-Y217Q, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0511] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of N061P-G097A-G128S-Y217Q, N061P-G097A-S101N-G128A-P210S-Y217Q, N061P-N062Q-G097A-S101N-I111V-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128S-Y217Q, and S024G-N025G-S053G-N061P-S078N-G097A-S101N-I111V-G128S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0512] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q, G097A-G128A-P210S-Y217Q, G097A-G128S-P210S-Y217Q, G097A-I111V-M124I-Y217Q, G097A-I111V-M124V-P210S-Y217Q, G097A-N123Q-P210S-Y217Q, G097A-N123Q-Y217Q, N061P-G097A-G128A-P210S-Y217Q, N061P-G097A-I111V-M124V-Y217Q, N061P-G097A-M124V-Y217Q, N061P-G097A-N123Q-Y217Q, N061P-G097A-S101N-I111V-M124V-Y217Q, N061P-G102A-P129S-Y217Q, N061P-N062Q-G097A-G100N-S101N-Y217Q, N061P-N062Q-G097A-G100Q-Y217Q, N061P-N062Q-G097A-I111V-Y217Q, N061P-N062Q-S078N-G097A-G100N-I111V-Y217Q, N061P-S078N-G097A-I111V-M124I-Y217Q, N061P-S078N-G102A-I111V-P129S-Y217Q, N062Q-G097A-I111V-P210S-Y217Q, N062Q-G097A-I111V-Y217Q, N062Q-S078N-G097A-I111V-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-N123Q-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, S024G-N025G-S053G-N061P-S101N-G102A-P129S-Y217Q, S053G-N061P-G097A-G128S-Y217Q, S053G-N061P-G102A-P129S-P210S-Y217Q, S053G-N061P-G102A-P129S-Y217Q, S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, S053G-N061P-S101N-G102A-P129S-Y217Q, S053G-S078N-G097A-I111V-G128S-Y217Q, S078N-G097A-G128S-Y217Q, and S078N-G097A-I111V-M124V-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0513] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of N061P-G097A-M124I-Y217Q, N061P-G097A-S101N-N123Q-Y217Q, N061P-N062Q-G097A-G100N-Y217Q, N061P-N062Q-G097A-G100Q-P210S-Y217Q, N061P-N062Q-G097A-G100Q-S101N-Y217Q, N061P-N062Q-G100N-G102A-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-M124I-Y217Q, S053G-N061P-G097A-M124I-Y217Q, S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, S053G-N061P-G097A-S101N-M124I-Y217Q, S053G-N061P-G097A-S101N-N123Q-Y217Q, and S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0514] The following BPN' variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-N123A-Y217Q, G097A-N123V-Y217Q, N061P-N062Q-G097A-G100D-Y217Q, N061P-S101N-G102A-G128S-Y217Q, Y217Q, N061P-G102A-G128S-Y217Q, S078N-G097A-I111V-N123Q-Y217Q, and G102A-N123Q-Y217Q, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0515] The following BPN' variants were determined to have a PI value greater than 1.0 to about 5 relative to BPN'-v3 in an egg microswatch cleaning assay in Detergent Composition 4 at 16.degree. C. and pH 8: BPN' amino acid sequence (SEQ ID NO:2) comprising the set of amino acid substitutions N061P-G097A-S101N-G128A-P210S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising amino acid substitutions N061P-G097A-S101N-G128A-P210S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0516] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128A-Y217Q, N061P-G102A-P129S-Y217Q, N062Q-G097A-I111V-P210S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128A-P210S-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-P210S-Y217Q, N061P-G097A-G128A-P210S-Y217Q, G097A-G128S-P210S-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-I111V-G128S-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-G128S-Y217Q, N061P-G097A-G128S-Y217Q, and G097A-G128A-P210S-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0517] The following BPN' variants were determined to have a PI value equal to or greater than 0.5 and equal to or less than 0.9 relative to BPN'-v3 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of N061P-G097A-M124I-Y217Q, S053G-N061P-G097A-S101N-N123Q-Y217Q, S053G-N061P-G102A-P129S-P210S-Y217Q, G097A-I111V-M124V-P210S-Y217Q, G097A-N123Q-P210S-Y217Q, S053G-N061P-S101N-G102A-P129S-Y217Q, S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, N061P-N062Q-G097A-S101N-I111V-Y217Q, N061P-N062Q-G097A-I111V-Y217Q, N062Q-G097A-I111V-Y217Q, N061P-G097A-S101N-I111V-M124V-Y217Q, G097A-N123Q-Y217Q, N061P-G097A-I111V-M124V-Y217Q, S078N-G097A-I111V-M124V-Y217Q, S053G-S078N-G097A-I111V-G128S-Y217Q, S078N-G097A-G128S-Y217Q, S053G-N061P-G097A-G128S-Y217Q, N061P-N062Q-G097A-G100N-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-N123Q-Y217Q, N061P-G097A-S101N-N123Q-Y217Q, N061P-N062Q-G097A-G100Q-P210S-Y217Q, N061P-G097A-N123Q-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-M124I-Y217Q, S053G-N061P-G097A-S101N-M124I-Y217Q, S053G-N061P-G097A-M124I-Y217Q, N061P-S078N-G097A-I111V-M124I-Y217Q, N061P-G097A-M124V-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, S024G-N025G-S053G-N061P-S101N-G102A-P129S-Y217Q, N061P-S078N-G102A-I111V-P129S-Y217Q, S053G-N061P-G102A-P129S-Y217Q, S024G-N025G-S053G-N061P-N062Q-G097A-S101N-I111V-Y217Q, N062Q-S078N-G097A-I111V-Y217Q, S024G-N025G-S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, S053G-N061P-G097A-S101N-I111V-M124V-Y217Q, G097A-I111V-M124I-Y217Q, Y217Q, N061P-N062Q-G100N-G102A-Y217Q, S053G-N061P-N062Q-G097A-G100N-S101N-Y217Q, N061P-N062Q-G097A-G100N-S101N-Y217Q, N061P-N062Q-S078N-G097A-G100N-I111V-Y217Q, N061P-N062Q-G097A-G100Q-Y217Q, N061P-S101N-G102A-G128S-Y217Q, G097A-N123V-Y217Q, G097A-N123A-Y217Q, G102A-N123Q-Y217Q, N061P-N062Q-G097A-G100Q-S101N-Y217Q, S078N-G097A-I111V-N123Q-Y217Q, N061P-N062Q-G097A-G100D-Y217Q, and N061P-G102A-G128S-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value of equal to or greater than 0.5 and equal to or less than 0.9 relative to BPN'-v3 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0518] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v3 in this egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2, wherein the variant comprises at least one substitution selected from the group of X024G, X025G, X053G, X061P, X062Q, X078N, X097A, X100D/N/Q, X101N, X102A, X111V, X123A/Q/V, X124I/V, X128A/S, X129S, X210S, and X217Q, and optionally at least one substitution selected from the group of S024G, N025G, S053G, N061P, N062Q, S078N, G097A, G100D/N/Q, S101N, G102A, I111V, N123A/Q/V, M124I/V, G128A/S, P129S, P210S, and Y217Q, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 6
Construction and Cleaning Performance of BPN' Variants
[0519] A BPN' combinatorial library based on BPN' parent was made by DNA2.0 and delivered as a ligation reaction. For efficient transformation of B. subtilis, DNA from the ligation reaction mixtures was amplified before transformation and transformants grown as described in Example 2. These variants were tested for cleaning performance using BMI microswatch assay in Detergent Composition 1 and Detergent Composition 4 at 16.degree. C. and pH 8 as well as egg microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8. Protein content was determined using the TCA assay and protease activity was assayed using the AAPF assay. The assays were performed as described in Example 1 and the Performance Indices were calculated relative to BPN'-v3 (with a PI value of 1.0).
[0520] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, N025G-G097A-S101N-G128A-Y217Q, N025G-S038G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-G128A-Y217Q, N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, N025G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-S101N-G128A-Y217Q, N061P-S101N-G128A-Y217Q, S024G-N025G-N061P-G097A-G128A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G128A-Y217Q, S024G-N025G-T055P-G097A-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-N061P-G097A-G128A-Y217Q, S024G-S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-T055P-N061P-G097A-G128A-Y217Q, S053G-G097A-S101N-G128A-Y217Q, S053G-N061P-G097A-S101N-G128A-Y217Q-S249N, S053G-N061P-S078N-G097A-G128A-Y217Q, S053G-S078N-G097A-S101N-G128A-Y217Q, S053G-T055P-G097A-S101N-G128A-Y217Q, S053G-T055P-N061P-S101N-G128A-Y217Q, S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, T055P-G097A-S101N-G128A-Y217Q, and T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence.
[0521] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128S-Y217Q, G097A-G128A-Y217Q, N025G-S078N-G097A-G128A-Y217Q, N025G-T055P-G097A-G128A-Y217Q, S024G-G097A-S101N-G128A-Y217Q, S024G-I035V-T055P-N061P-S078N-G097A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A, S024G-N025G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-G097A-G128A-S130G-Y217Q, S024G-N025G-S053G-N061P-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G128A-Y217Q, S024G-N025G-S053G-T055P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S101N-G128A-Y217Q, S024G-N025G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-Y217Q, S024G-N061P-G097A-S101N-G128A-Y217Q, S024G-S038G-S053G-S078N-S101N-G128A-Y217Q, S024G-S053G-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-S078N-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-T055P-G097A-G128A-Y217Q, S024G-T055P-N061P-G097A-S101N-G128A, S024G-T055P-N061P-S078N-S101N-G128A-Y217Q, S024G-T055P-S078N-G097A-S101N-G128A-Y217Q, S101N-G128A-Y217Q, T055P-N061P-G097A-A116S-G128A, and T055P-N061P-S078N-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0522] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN' and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v3 in this BMI microswatch cleaning assay in Detergent Composition 1 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2, wherein the variant comprises at least one substitution selected from the group of X024G, X025G, X035V, X038G, X053G, X055P, X061P, X078N, X097A, X101N, X116S, X128A/S, X130G, X216Q, X217Q, and X249N, and optionally at least one substitution selected from the group of S024G, N025G, I035V, S038G, S053G, T055P, N061P, S078N, G097A, S101N, A116S, G128A/S, S130G, Y216Q, Y217Q, and S249N, and wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0523] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of S024G-N025G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-T055P-N061P-S078N-S101N-G128A-Y217Q, S053G-G097A-S101N-G128A-Y217Q, and T055P-N061P-S078N-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence.
[0524] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128S-Y217Q, G097A-G128A-Y217Q, N025G-G097A-S101N-G128A-Y217Q, N025G-S038G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-G128A-Y217Q, N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, N025G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-G128A-Y217Q, N025G-S078N-G097A-S101N-G128A-Y217Q, N025G-T055P-G097A-G128A-Y217Q, N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-S101N-G128A-Y217Q, N061P-S101N-G128A-Y217Q, S024G-G097A-S101N-G128A-Y217Q, S024G-I035V-T055P-N061P-S078N-G097A-Y217Q, S024G-N025G-N061P-G097A-G128A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A, S024G-N025G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-G097A-G128A-S130G-Y217Q, S024G-N025G-S053G-N061P-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G128A-Y217Q, S024G-N025G-S053G-T055P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S101N-G128A-Y217Q, S024G-N025G-T055P-G097A-G128A-Y217Q, S024G-N025G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N061P-G097A-S101N-G128A-Y217Q, S024G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S038G-S053G-S078N-S101N-G128A-Y217Q, S024G-S053G-N061P-G097A-G128A-Y217Q, S024G-S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-S078N-S101N-G128A-Y217Q, S024G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-T055P-G097A-G128A-Y217Q, S024G-T055P-N061P-G097A-G128A-Y217Q, S024G-T055P-N061P-G097A-S101N-G128A, S024G-T055P-S078N-G097A-S101N-G128A-Y217Q, S053G-N061P-G097A-S101N-G128A-Y217Q-S249N, S053G-N061P-S078N-G097A-G128A-Y217Q, S053G-S078N-G097A-S101N-G128A-Y217Q, S053G-T055P-G097A-S101N-G128A-Y217Q, S053G-T055P-N061P-S101N-G128A-Y217Q, S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, S101N-G128A-Y217Q, T055P-G097A-S101N-G128A-Y217Q, and T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0525] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of T055P-N061P-G097A-A116S-G128A and S024G-N025G-T055P-N061P-S078N-G097A-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity and may have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) in this assay. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0526] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N061P-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-G097A-S101N-G128A-Y217Q, S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-N025G-T055P-G097A-G128A-Y217Q, and S024G-N025G-S053G-T055P-G097A-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence.
[0527] The following BPN' variants were determined to have a PI value of about 1.0 relative to BPN'-v3 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of G097A-G128S-Y217Q, G097A-G128A-Y217Q, S024G-G097A-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-S101N-G128A-Y217Q, S053G-T055P-N061P-S101N-G128A-Y217Q, S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, N025G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, T055P-N061P-S078N-G128A-Y217Q, N025G-S038G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-G128A-Y217Q, S024G-N025G-S053G-T055P-S078N-S101N-G128A-Y217Q, T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G128A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-Y217Q, S024G-T055P-G097A-G128A-Y217Q, T055P-N061P-G097A-A116S-G128A, S053G-T055P-G097A-S101N-G128A-Y217Q, T055P-G097A-S101N-G128A-Y217Q, S024G-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-G097A-G128A-Y217Q, S024G-S053G-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S053G-G097A-S101N-G128A-Y217Q, N025G-T055P-G097A-G128A-Y217Q, S024G-T055P-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-S101N-G128A-Y217Q, N025G-G097A-S101N-G128A-Y217Q, S024G-S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-Y217Q, and S024G-I035V-T055P-N061P-S078N-G097A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning comprising utilizing at least one such variant as described in greater detail elsewhere herein.
[0528] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of S101N-G128A-Y217Q, S024G-T055P-N061P-G097A-S101N-G128A, S024G-N025G-N061P-S078N-G097A-S101N-G128A, S024G-T055P-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S053G-N061P-G097A-S101N-G128A-Y217Q-S249N, N025G-S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-G097A-S101N-G128A-Y217Q, S024G-S053G-N061P-G097A-G128A-Y217Q, S024G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-T055P-N061P-G097A-G128A-Y217Q, S024G-S038G-S053G-S078N-S101N-G128A-Y217Q, S053G-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-G128A-Y217Q, and S024G-N025G-S053G-N061P-G097A-G128A-S130G-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0529] The following BPN' variant was determined to have a PI value of about 0.8 relative to BPN'-v3 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising amino acid substitutions S024G-S053G-S078N-G097A-S101N-G128A-Y217Q, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. The invention includes a protease variant having proteolytic activity, a PI value of about 0.8 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising amino acid substitutions S024G-S053G-S078N-G097A-S101N-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0530] The following BPN' variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1 to about 12, from greater than 4 to about 12, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v3 in an AAPF proteolytic assay: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of S024G-G097A-S101N-G128A-Y217Q, S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-S101N-G128A-Y217Q, S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-T055P-N061P-G097A-S101N-G128A, S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A, N061P-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-S101N-G128A-Y217Q, S024G-T055P-N061P-S078N-S101N-G128A-Y217Q, N025G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S101N-G128A-Y217Q, N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-G128A-Y217Q, S024G-S053G-T055P-N061P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, T055P-N061P-S078N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, N025G-S038G-S053G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-N061P-G128A-Y217Q, N025G-S053G-N061P-S078N-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-S078N-S101N-G128A-Y217Q, T055P-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G128A-Y217Q, S024G-N025G-N061P-G097A-S101N-G128A-Y217Q, S053G-N061P-G097A-S101N-G128A-Y217Q-S249N, N025G-S053G-T055P-S078N-G097A-S101N-G128A-Y217Q, S024G-T055P-G097A-G128A-Y217Q, T055P-N061P-G097A-A116S-G128A, S024G-N025G-T055P-G097A-S101N-G128A-Y217Q, S024G-N025G-N061P-S078N-G097A-S101N-G128A-Y217Q, S053G-T055P-G097A-S101N-G128A-Y217Q, T055P-G097A-S101N-G128A-Y217Q, S024G-N061P-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-G097A-S101N-G128A-Y217Q, G097A-G128S-Y217Q, S024G-S053G-N061P-G097A-G128A-Y217Q, S024G-N025G-N061P-G097A-G128A-Y217Q, S024G-N061P-S078N-G097A-S101N-G128A-Y217Q, S024G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-S078N-S101N-G128A-Y217Q, S024G-N025G-S053G-T055P-N061P-S078N-G097A-G128A-Y217Q, S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-T055P-N061P-G097A-G128A-Y217Q, S024G-S038G-S053G-S078N-S101N-G128A-Y217Q, S053G-G097A-S101N-G128A-Y217Q, N025G-T055P-G097A-G128A-Y217Q, S024G-T055P-S078N-G097A-S101N-G128A-Y217Q, S053G-S078N-G097A-S101N-G128A-Y217Q, S024G-N025G-T055P-G097A-G128A-Y217Q, S024G-N025G-S053G-T055P-G097A-G128A-Y217Q, N025G-S078N-G097A-S101N-G128A-Y217Q, N025G-G097A-S101N-G128A-Y217Q, S024G-S053G-N061P-S078N-G097A-G128A-Y217Q, S024G-S053G-S078N-G097A-S101N-G128A-Y217Q, N025G-S078N-G097A-G128A-Y217Q, and S024G-N025G-S053G-N061P-G097A-G128A-S130G-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' protease (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v3 in this AAPF assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence.
[0531] The following BPN' variant was determined to have a PI value of about 1.0 relative to BPN'-v3 in an AAPF proteolytic assay: BPN' amino acid sequence (SEQ ID NO:2) comprising amino acid substitutions G097A-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' protease (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a PI value of about 1.0 relative to BPN'-v3 in this AAPF assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and comprising amino acid substitutions G097A-G128A-Y217Q, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 7
Construction of Site Evaluation Libraries of BPN'-v36 and Cleaning Performance of BPN'-v36 Variants
Construction of the Site Evaluation Libraries of BPN'-v36
[0532] The amino acid sequence of BPN'-v36 is set forth in SEQ ID NO:6 below:
TABLE-US-00032 (SEQ ID NO: 6) AQSVPYGVSQIKAPALHSQGYTGGNVKVAVIDSGIDSSHPDLKVAGGASM VPGETNPFQDNNSHGTHVAGTVAALNNNIGVLGVAPSASLYAVKVLGADG NGQYSWIINGIEWAIANNMDVINMSLGAPSGSAALKAAVDKAVASGVVVV AAAGNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMA PGVSIQSTLPGNKYGAQNGTSMASPHVAGAAALILSKHPNWTNTQVRSSL ENTTTKLGDSFYYGKGLINVQAAAQ
[0533] The nucleic acid sequence encoding the BPN'-v36 protease variant is:
TABLE-US-00033 (SEQ ID NO: 5) Gcgcagtccgtgccttacggcgtatcacaaattaaagcccctgctctgca ctctcaaggctacactggaggcaatgttaaagtagcggttatcgacagcg gtatcgactcgagccatccagatcttaaagtcgctggaggggcttctatg gtgccgggcgaaacaaacccgtttcaagataacaattctcatggcacaca cgtcgcaggaacggttgcggcgttaaacaataatattggcgtgcttggtg tagccccgtctgcttcgctctacgccgttaaagttcttggcgcagacgga aatggccaatactcatggattatcaacggcatcgaatgggccatcgcgaa taacatggatgtaatcaacatgagcctgggagcaccaagcggcagtgcgg cacttaaagcagcagttgataaagctgttgcatctggtgtcgtcgtagta gcggcagctgggaatgagggaacatccggatcatcgagtaccgtcggtta tccaggcaagtacccttcagtgattgcagtgggcgctgtagactcttcaa atcaacgtgcctctttttcctccgtgggaccggagctggatgtcatggcc cctggcgtttctattcaatcgacgcttccagggaacaagtatggtgcgca aaacgggacttccatggcctcgccgcatgtagctggggcggccgcattga ttctttctaagcacccgaactggacaaacactcaagtccgcagcagttta gaaaacaccactacaaaacttggtgattctttctactatggaaaagggct gatcaacgtacaggcggcagctcag
[0534] The amino acid sequence of BPN'-v36 may be represented by reference to the subtilisin BPN' amino acid sequence of SEQ ID NO:2. That is, BPN'-v36 may be represented as the subtilisin BPN' sequence of SEQ ID NO:2 with the six amino acid substitutions S024G-S053G-S078N-S101N-G128A-Y217Q. The BPN'-v36 amino acid sequence may be conveniently designated as BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q or BPN'+S024G+S053G+S078N+S101N+G128A+Y217Q. Throughout this specification, unless otherwise indicated, each amino acid position of an amino acid sequence is numbered according to the numbering of a corresponding amino acid position in the amino acid sequence of Bacillus amyloliquefaciens subtilisin BPN' shown in SEQ ID NO:2 as determined by alignment of the variant amino acid sequence with the Bacillus amyloliquefaciens subtilisin BPN' amino acid sequence.
[0535] Site evaluation libraries (SELs) were created at every single amino acid position in mature BPN'-v36 (i.e., BPN'-S24G-S53G-S78N-S101N-G128A-Y217Q) protein by PCR fusion.
[0536] For each codon to be mutated in the BPN'-v36 protease, a pair of partially overlapping, complementary (mutagenic forward and reverse) primers were designed. Each mutagenic primer contained the NNS (N=A,C,G, or T and S=G or C) mutagenic codon in the center flanked by at least 15 nucleotides on each side. To create a library at a given position, two PCR reactions were carried out using either a common forward gene-flanking primer (P4974, GCCTCACATTTGTGCCACCTA; SEQ ID NO:60) and a mutagenic NNS reverse primer, or the common reverse gene-flanking primer (P4976, CCTCTCGGTTATGAGTTAGTTC; SEQ ID NO:61) and a mutagenic NNS forward primer. These PCR reactions generated two PCR fragments, one encoding the 5' half of the mutant BPN'-v36 gene (5' gene fragment) and the other encoding the 3' half of the mutant BPN'-v36 gene (3' gene fragment).
[0537] Each PCR amplification reaction contained 30 pmol of each primer and 100 ng of the BPN'-v36 parent template DNA (plasmid pHPLT-BPN'-v36, see FIG. 4). Amplifications were carried out using Vent DNA polymerase (NEB). The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 40 sec. Following amplification, the 5' and 3' gene fragments were gel-purified by the QIAGEN.RTM. gel-band purification kit, mixed (50 ng of each fragment), mixed and amplified by PCR once again using the primers P4973 (AAAGGATCCTAATCGGCGCTTTTC; SEQ ID NO:62) and P4950 (CTTGTCTCCAAGCTTAAAATAAAA; SEQ ID NO:63) to generate the full-length gene fragment. The PCR conditions were same as described above, except the extension phase, which was carried out at 72.degree. C. for 2 min. The full-length DNA fragment was gel-purified by the QIAGEN.RTM. gel-band purification kit, digested by the BamHI and HindIII restriction enzymes and ligated with the pHPLT-BPN' partial opt vector that also was digested with the same restriction enzymes. Ligation mixtures were amplified using rolling circle amplification in an Illustra Templiphi kit according to the manufacturer's recommendation (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. For this purpose, 1 .mu.l of the ligation mixture was mixed with 5 .mu.l of the sample buffer, heated to 95.degree. C. for 3 min and cooled on ice. Next, 5 .mu.l of the reaction buffer and 0.2 .mu.l of the enzyme were added to each tube, followed by incubation at 30.degree. C. for 10 hours. Products of the rolling circle amplification were diluted 100 times and used to transform B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C. Subsequently, the colonies with halos were inoculated in 150 .mu.l of LB media containing 10 .mu.g/mL neomycin. The next day, cultures were either frozen with 15% glycerol or grown in MBD medium for biochemical analysis as described in Example 2.
Cleaning Performance of the BPN'-v36 Variants
[0538] Protein variants from BPN'-v36 SEL were tested for cleaning performance using a BMI microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8 and egg microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8. Protein content was determined using the TCA assay. The assays were performed as described in Example 1 and the Performance Indices were calculated relative to BPN'-v36 (i.e., BPN'-S24G-S53G-S78N-S101N-G128A-Y217Q) (with a PI value of 1.0).
[0539] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q (SEQ ID NO:6) (i.e., BPN'-v36) comprising at least one amino acid substitution selected from the group consisting of A116V, G160S, I111L, I115V, N109S, N117M, P005G, Q059V, T164S, Y262M, A015Q, A015S, A098E, A098N, A098S, A098T, A098V, A098Y, A114S, A114T, A116G, A116L, A116S, A116T, A116W, A133G, A133H, A133T, A133V, A137G, A137I, A137L, A137S, A137T, A138S, A216E, A216F, A216V, D099S, D181E, F261A, F261Q, G024F, G024I, G024Q, G024Y, G097S, G160T, G211L, G211V, H017F, H017W, H039V, H226A, I031V, I111V, I268V, K170R, K265R, L016Q, L016T, L135M, L209T, L209V, L233M, L257T, L257V, L267A, L267V, N025A, N025I, N025Q, N025R, N025T, N025V, N101I, N101Q, N101S, N109A, N109G, N109H, N109L, N109M, N109Q, N109T, N117Q, N184A, N184L, N184T, N184W, N212G, N212L, N212V, N243P, N252G, N252M, P005T, P014S, P040G, P040L, P040Q, P129A, P129S, P172G, P172S, P194Q, P210A, P210S, Q185F, Q185G, Q185I, Q185M, Q185N, Q185S, Q275H, R186K, S009A, S009G, S009H, S009M, S018T, S130T, S132N, S145K, S159T, S161I, S161K, S161N, S161T, S162I, S162M, S162Y, S163G, S182F, S182G, S182V, S182W, S183F, S183L, S183M, S183T, S183V, S183W, S224A, S236T, S249V, T022A, T022G, T022Q, T022V, T208V, T242S, T253N, T253S, T254A, T254S, T255L, T255S, T255V, V004A, V004P, V004W, V084C, V139C, V165M, V203F, Y021K, Y021N, Y021T, Y021V, Y167F, Y171F, Y214F, Y262F, and Y262T, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN'-v3 and BPN'-v36, a PI value greater than that of BPN'-v3, and/or a PI value greater than 1 to about 5 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0540] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one amino acid substitution selected from the group consisting of A001G, A001Y, A013G, A013V, A015F, A015G, A015K, A015M, A015P, A015T, A015W, A015Y, A029G, A073S, A088C, A088I, A088L, A088T, A088V, A098D, A098K, A098P, A098R, A098W, A116D, A116E, A116R, A128S, A133L, A133M, A133S, A134G, A134S, A137N, A137V, A144M, A144Q, A144S, A144T, A144V, A151C, A176S, A176T, A179S, A216G, A216L, A216P, A216Q, A216S, A216T, A216Y, A228T, A230C, A231C, A272I, A272L, A272Q, A272S, A272T, A272W, A273S, A274G, A274L, A274Q, A274T, A274V, D041E, D099G, D099N, D120A, D120K, D120Q, D120R, D120S, D140E, D181S, D259E, E054D, E156D, E156T, E251V, F261G, F261H, F261L, F261R, F261S, F261T, F261V, F261W, G007A, G007S, G020A, G020D, G020S, G024N, G024R, G024S, G024T, G024V, G024W, G053H, G053K, G053N, G053T, G097A, G097D, G097T, G131A, G131H, G131P, G131Q, G131T, G131V, G160H, G160P, G166A, G166S, G166T, G211A, G211D, G211M, G211N, G211P, G211Q, G211R, G211W, G215A, G215N, G215V, H017L, H017M, H017T, H017V, H017Y, H039A, H039C, H039N, H226F, H226I, H226M, H226S, H226V, H226Y, I035V, I079A, I079S, I079T, I079V, I079W, I108V, I115L, I122A, I234L, I234V, K012R, K027R, K136R, K141F, K213W, K237R, K256R, K265Q, L016A, L016F, L016I, L016S, L016V, L042I, L075A, L075M, L075Q, L075V, L075Y, L082K, L082M, L082Q, L082V, L090I, L196I, L196V, L209Q, L209W, L233A, L233Q, L233V, L235I, L235K, L250I, L257A, L257H, L257Q, L257S, L257Y, L267Q, L267R, L267S, L267T, M199V, N025F, N025G, N025H, N025K, N025L, N025M, N025S, N025Y, N061F, N061H, N061P, N061S, N061T, N061V, N061W, N076G, N076W, N078S, N078T, N078V, N101A, N101H, N101L, N101T, N109K, N109R, N117A, N117E, N117H, N117K, N118G, N184G, N184H, N184I, N184S, N184V, N212A, N212F, N212I, N212K, N212P, N212Q, N212S, N212Y, N218A, N218H, N218L, N218S, N240E, N240H, N240L, N240R, N240T, N243A, N243Q, N243T, N243V, N252A, N252K, N252L, N252Q, N252R, N252S, N252T, N269Q, N269S, P014G, P014Q, P014T, P040A, P040H, P040S, P040T, P040V, P040Y, P086A, P086C, P086F, P086H, P086S, P129D, P129G, P129K, P129T, P172A, P172Q, P194A, P194G, P194L, P194M, P194S, P194V, P194Y, P210G, P210R, P210V, Q002A, Q002S, Q010A, Q010F, Q010H, Q010I, Q010L, Q010N, Q010S, Q010T, Q019A, Q019G, Q019N, Q019S, Q019T, Q019V, Q019W, Q059I, Q103L, Q103S, Q185A, Q185H, Q185L, Q185T, Q185Y, Q206P, Q206S, Q206Y, Q217I, Q217N, Q217S, Q217T, Q245K, Q275D, Q275S, Q275W, S003A, S003G, S003H, S003M, S003P, S003Q, S003T, S003V, S009I, S009L, S009P, S009T, S009W, S018A, S018G, S018I, S018L, S018M, S018N, S018P, S018V, S018W, S033T, S037Q, S037T, S037V, S038G, S038H, S038K, S038Q, S038T, S063K, S063N, S063Q, S063T, S087A, S087F, S087G, S087Q, S087T, S089L, S089M, S089N, S089Q, S089T, S089W, S130A, S130F, S130G, S130L, S130V, S145A, S145H, S145M, S145V, S159A, S159G, S159H, S159Q, S159R, S161A, S161G, S161H, S161L, S161M, S161P, S161Q, S161W, S162A, S162F, S162G, S162L, S162N, S162P, S162R, S162V, S163P, S173A, S173G, S182A, S182H, S182K, S182L, S182N, S182P, S182Q, S182T, S183A, S183G, S183H, S183Q, S188A, S188G, S188T, S188V, S191A, S204A, S204I, S204L, S204Q, S204V, S224C, S236A, S236N, S236Q, S248A, S248F, S248G, S248I, S248K, S248L, S248M, S248N, S248Q, S248T, S248V, S249A, S249C, S249H, S249Q, S249T, S249W, S249Y, S260H, S260N, S260P, S260T, T022H, T022K, T022N, T022R, T022S, T022Y, T055A, T055G, T055L, T055N, T055P, T055Q, T071S, T158H, T158S, T164N, T208C, T208L, T220S, T242N, T244A, T244G, T244H, T244I, T244Q, T244S, T244V, T244W, T253A, T253G, T253H, T253Q, T254V, T255A, T255G, T255H, T255I, T255Q, T255Y, V004G, V004N, V004R, V008A, V008C, V008M, V026I, V044I, V044L, V045H, V045K, V045L, V045M, V045Q, V045S, V045V, V045W, V045Y, V051I, V081L, V081Q, V081T, V084A, V084S, V084T, V093I, V121I, V143N, V143S, V143Y, V147C, V147I, V147L, V147T, V180I, V180L, V180T, V192A, V192S, V192T, V198I, V198L, V198M, V203H, V203I, V203L, V203N, V203Q, V203T, V203W, V203Y, V270A, V270S, V270T, W241M, W241Y, Y006G, Y006H, Y006I, Y006K, Y006L, Y006P, Y006Q, Y006T, Y006V, Y006W, Y021A, Y021D, Y021E, Y021L, Y021Q, Y021R, Y021S, Y104F, Y104I, Y214L, Y214V, Y214W, Y262A, Y262G, Y262L, Y262N, Y262S, Y262W, Y263G, and Y263W, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Thus, e.g., the invention includes BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising substitution A128S, e.g., BPN'-S024G-S053G-S078N-S101N-G128S-Y217Q. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and/or a PI value of about 1.0 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0541] The following BPN'-v36 variants were determined to have a PI value of about 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one amino acid substitution selected from the group consisting of A001F, A001K, A001L, A001M, A001Q, A001R, A001S, A001T, A001V, A013C, A013S, A015D, A015E, A015L, A015R, A048S, A073N, A073T, A074G, A074S, A085C, A085G, A085S, A085V, A088M, A088S, A092S, A098G, A114G, A133P, A137E, A137H, A144G, A144H, A144K, A144L, A144N, A153S, A153V, A176C, A179G, A187G, A187S, A200G, A216W, A223S, A228S, A230T, A230V, A231V, A232C, A232V, A272E, A272G, A272K, A272P, A273G, A273L, A273V, A274M, A274R, D036E, D099A, D099Q, D120E, D181A, D181G, D259A, D259G, D259Q, D259T, E156A, E156S, E251I, E251L, E251Q, E251T, F058Y, F261C, F261D, F261K, F261P, G020E, G020F, G020H, G020L, G020N, G020Q, G020R, G020T, G020Y, G024A, G024P, G053A, G053D, G053E, G053F, G053L, G053Q, G053S, G053Y, G097K, G097M, G157A, G157S, G160A, G160L, G166C, G166I, G166Q, G169A, G211K, G215H, G215L, G215S, G215T, G215W, G258S, H017I, H039S, H226L, H238N, H238Y, I011L, I011V, I031L, I079F, I079K, I079L, I079M, I079Q, I205A, I205V, I268L, I268M, K012G, K043F, K043H, K043I, K043N, K043Q, K043T, K141A, K141R, K141W, K170A, K213A, K213G, K213H, K213I, K213L, K213N, K213Q, K213R, K213S, K213T, K213V, K237A, K237H, K237I, K237L, K237N, K237S, K256A, K256G, K256H, K256M, K256P, K256Q, K256W, K265H, L016E, L042V, L075G, L075H, L075I, L075T, L082A, L082F, L082H, L082R, L082S, L082T, L090M, L135F, L196M, L209C, L209H, L209S, L233S, L235M, L235R, L235W, L257C, L257G, L267F, M050Y, M119C, M119I, M124L, N025C, N025E, N025P, N061A, N061G, N061I, N061K, N061L, N061Q, N061R, N062S, N062T, N076A, N076P, N076Q, N076S, N076T, N076V, N078G, N078H, N078K, N078P, N078Q, N078R, N101F, N117R, N117S, N118D, N118H, N118Q, N118R, N118S, N118T, N184C, N184E, N184R, N212D, N212R, N212W, N218F, N218G, N218M, N218P, N218T, N218V, N218W, N240A, N240G, N240Q, N240S, N240W, N243C, N243G, N243S, N252V, N269H, P005A, P005D, P005M, P005Q, P014A, P014M, P014R, P014V, P040F, P040R, P040W, P129E, P129R, P172E, P172K, P194H, P194R, P194W, P201A, P201G, P210L, P239K, P239R, Q002D, Q002E, Q002G, Q002I, Q002P, Q002V, Q010D, Q010R, Q019C, Q019D, Q019E, Q019H, Q019L, Q019P, Q019R, Q059A, Q059E, Q059L, Q059S, Q059T, Q103W, Q185D, Q185K, Q185R, Q185W, Q206G, Q206H, Q206L, Q206V, Q206W, Q217E, Q217F, Q217H, Q217L, Q217V, Q245M, Q271A, Q271D, Q271G, Q271L, Q271P, Q271T, Q271Y, Q275F, Q275L, Q275P, Q275R, S003D, S003F, S003K, S003R, S009K, S018D, S018R, S037A, S037G, S037K, S037L, S037P, S038M, S063A, S063F, S063G, S063M, S063R, S063Y, S087C, S087K, S087L, S087M, S087N, S087Y, S089A, S089D, S089F, S089G, S089H, S089I, S089K, S089R, S089V, S089Y, S130D, S130E, S130K, S130W, S145G, S145L, S145R, S145T, S159D, S159L, S159W, S161E, S161R, S162C, S162E, S162W, S163A, S182E, S182R, S183C, S183D, S183P, S183R, S188D, S188P, S204G, S204Y, S207G, S224G, S224T, S236C, S236G, S248D, S248H, S248R, S249E, S249L, S249R, S260A, S260G, S260K, S260Q, S260V, S260Y, T022L, T055D, T055E, T055I, T055K, T055M, T055S, T055V, T055Y, T158A, T158G, T158L, T158Q, T158V, T164K, T164Q, I208S, T244D, T244E, T244R, T253E, T253R, T253Y, T254G, T255D, T255E, T255K, T255R, V026A, V028I, V028L, V030I, V044C, V044P, V045E, V045G, V045N, V072L, V081A, V081G, V081H, V081S, V084I, V084M, V095A, V095C, V143A, V143F, V143H, V143Q, V143T, V143W, V147A, V147Q, V147S, V148I, V148L, V149C, V149I, V149L, V165L, V180A, V180C, V180M, V192C, V192F, V192I, V192Q, V192Y, V203A, V203G, V203K, V203S, V270C, V270L, V270P, W241F, Y006A, Y006M, Y006N, Y006R, Y006S, Y021C, Y091W, Y104V, Y104W, Y262C, Y262D, Y262E, Y262H, Y262I, Y262R, and Y262V, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants may have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and/or a greater PI value than that of BPN' in this assay. The invention includes a protease variant having proteolytic activity and/or a PI value of about 0.9 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0542] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S24G-S53G-S78N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one amino acid substitution selected from the group consisting of A001D, A001H, A001N, A015C, A048C, A048E, A085T, A133R, A137R, A142C, A144D, A144R, A152S, A153G, A187P, A187Q, A187T, A187V, A216R, A230S, A272R, A273H, A273T, A274H, D036N, D036S, D181H, D181T, D259N, D259P, D259S, E156G, E156H, E156L, E156Q, E156V, E251C, F189S, F189T, F189W, F189Y, F261E, G020C, G024D, G053M, G053R, G097R, G131D, G131R, G157N, G160R, G160V, G166L, G166W, G211E, G215D, G258A, G258D, G258P, I011T, I031C, I079E, I079R, I175L, I205C, K012H, K012N, K027A, K027N, K027S, K043A, K043D, K043E, K043G, K043L, K043M, K043P, K043V, K043W, K043Y, K136H, K141H, K141L, K141M, K141N, K141Q, K141T, K141V, K170G, K170S, K237T, K237V, K256D, K256S, K256T, K256V, K265N, K265S, L042F, L042M, L082E, L209A, L209E, L209G, L209R, L233G, L235V, L257D, L257E, L257P, L257R, L257W, L267E, M050L, N056D, N056S, N061C, N061D, N062A, N062H, N062L, N062V, N062Y, N076D, N076L, N076M, N078D, N078F, N101D, N101R, N118A, N212C, N212E, N218C, N218D, N218E, N252D, N252E, P014F, P014K, P057A, P057W, P172R, P194E, P201T, P210E, Q059C, Q059D, Q059R, Q185E, Q206D, Q217A, Q217K, Q217R, Q245A, Q245D, Q245E, Q245H, Q245R, Q271E, Q271F, Q271R, Q271W, Q275G, Q275I, R186I, R186L, R186V, R186W, S003E, S009C, S009E, S018C, S037D, S037E, S037H, S037R, S037Y, S038D, S038P, S038R, S038Y, S063L, S087D, S087R, S089C, S089E, S130C, S130R, S145D, S159C, S159P, S161C, S173T, S182C, S188E, S188F, S188K, S188L, S188R, S188W, S190A, S190G, S190T, S204R, S236D, S236E, S248C, S248E, S260C, S260E, S260R, T022P, T055C, T055W, T071A, T158D, T158E, T158P, T158R, T158Y, T164R, T242D, T242G, T255C, V004E, V004T, V045C, V045D, V045R, V045T, V051H, V081R, V143C, V143E, V143G, V192G, V203C, V203D, V203E, V203M, V203R, V270G, W241L, Y214H, Y214Q, A001E, A133E, A187L, A187N, A216C, A216H, A273Q, D099H, D259H, E156C, E195G, F189H, G131C, G146A, G166V, G215C, G215E, I107L, K012A, K012S, K012T, K043C, K170C, K256C, K256E, K265G, K265Y, L233E, M222F, M222S, N062Q, N076E, N078E, N184P, N218R, P005V, P014D, Q002K, Q002L, Q002R, Q010W, Q271C, R186H, S049C, S063C, S063D, S105T, S188C, S190C, S204E, T055R, T164G, V004D, V044T, V045I, V165C, V180S, Y006C, Y006D, Y006E, Y104T, A001C, A187C, A230G, A273D, A273P, D036Q, F189G, F189L, F189R, G157T, G178A, I031F, I111M, K012F, K012L, K027T, K043R, K136G, K141G, K170Q, M222A, M222L, N062R, N117G, N269C, P005W, P129V, P239A, P239H, P239T, Q059W, Q217G, Q275A, R186A, S191G, T164A, T220A, A001P, A187F, A187W, A273R, D041C, D060G, D197T, F189A, G046D, G157P, K012C, K012E, K012W, L042C, M222T, N062C, P239G, P239N, Q217C, R186M, S049T, S089P, S125A, S173V, and V044A, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0543] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at 16.degree. C. and pH 8: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one amino acid substitution selected from the group consisting of A216E, L090I, A098R, A098W, A098Y, A116G, A116R, A116S, A133M, I107L, I115V, M124L, N101I, N109H, N109S, N109T, N117R, P005G, Q185L, S089V, V095A, A015Y, A029G, A098D, A098E, A098G, A098N, A098S, A098T, A098V, A114S, A114T, A116E, A116L, A116T, A116V, A133H, A133L, A133S, A137G, A137I, A137L, A137S, A137V, A138S, A144S, A144V, A176S, A176T, A187T, A216F, A216P, A216Q, A216R, A216S, A216T, A216V, A216Y, D041E, D120A, D120E, D120Q, D120R, D120S, D181S, G020A, G020S, G024A, G097A, G097D, G097S, G131Q, G160S, G166I, G211L, G215N, H039N, H238N, I111L, I111V, I122A, L075I, L075Q, L135M, L209T, L209V, L233V, L235M, L235R, L257A, M119I, N025A, N025G, N025T, N061K, N101F, N101H, N101L, N101Q, N101R, N101S, N101T, N109A, N109G, N109K, N109L, N117E, N117H, N117K, N117S, N212G, N212S, N218F, N218G, N218H, N218L, N218S, N218W, N240Q, N252M, N252R, N252S, P005T, P040A, P040G, P040T, P129D, P129S, P194S, P210R, Q019R, Q019W, Q103L, Q103W, Q185A, Q185G, Q185M, Q185R, Q185T, Q206G, Q206Y, Q217A, Q217E, Q217R, Q217S, Q217T, S003Q, S009H, S018M, S033T, S130A, S130F, S130G, S130T, S130V, S145T, S159A, S161N, S161T, S162V, S162Y, S182L, S182W, S183F, S183L, S183V, S183W, S188K, S188W, S236Q, S236T, S248L, T022H, T022K, T208C, T253H, T255V, V044I, V121I, V139C, V143H, V143Q, V143T, V143W, V143Y, Y006K, Y021A, Y104W, A001F, A001G, A001H, A001K, A001L, A001Q, A001S, A001Y, A013V, A015G, A015K, A015R, A015S, A015T, A015W, A048S, A073N, A073S, A092S, A098K, A098P, A116D, A116W, A128S, A133P, A133T, A133V, A134G, A134S, A137H, A137N, A137T, A144D, A144K, A144L, A144M, A144N, A144R, A179G, A179S, A187V, A216G, A216L, A216W, A223S, A230C, A272K, A272L, A272P, A272S, A272T, A272W, A273G, A273S, A274G, A274M, A274T, D120K, D140E, D181A, D181E, D181G, D181H, D181T, D259E, D259N, D259Q, E054D, E156D, E156T, E251L, E251T, E251V, F058Y, F189W, F261K, F261Q, F261R, G007A, G007S, G020F, G020H, G020N, G020Q, G020T, G020Y, G024F, G024Q, G024R, G024T, G024V, G024W, G024Y, G053T, G097K, G097M, G097R, G097T, G131A, G131H, G131P, G131R, G131T, G131V, G160H, G160T, G166C, G166Q, G166S, G166T, G211A, G211D, G211K, G211M, G211N, G211Q, G211R, G211V, G211W, G215S, G215T, G215W, H017T, H017W, H017Y, H039V, H226A, H226F, H226I, H226L, H226M, H226V, I035V, I079A, I079S, I108V, I205V, I234L, I234V, I268V, K012S, K043P, K136H, K136R, K141A, K141F, K141T, K141W, K170A, K170G, K170R, K213A, K213R, K213S, K237A, K237H, K237L, K237S, K237V, K256A, K256G, K256H, K256M, K256P, K256Q, K256R, L016A, L016Q, L016T, L016V, L042V, L075M, L075T, L082M, L082V, L135F, L196I, L209H, L209Q, L209R, L209S, L209W, L233A, L233M, L233Q, L235I, L235K, L250I, L257S, L257T, L257V, L267A, L267Q, L267T, L267V, M119C, M199V, N025C, N025E, N025F, N025I, N025K, N025L, N025M, N025Q, N025V, N025Y, N061F, N061P, N061S, N061T, N076G, N078S, N101A, N109M, N109Q, N109R, N117A, N117M, N117Q, N118D, N118G, N118H, N118Q, N118R, N118S, N184A, N184C, N184G, N184L, N184R, N184S, N184T, N184V, N184W, N212C, N212F, N212I, N212K, N212L, N212P, N212Q, N212R, N212V, N212W, N212Y, N218A, N218P, N218T, N240A, N240E, N240G, N240H, N240L, N240R, N240S, N240T, N243C, N243Q, N243T, N243V, N252A, N252G, N252K, N252Q, P014G, P014Q, P014R, P014S, P014T, P040F, P040L, P040Q, P040S, P040V, P086C, P086H, P086S, P129A, P129E, P129G, P129K, P129R, P172A, P172K, P172Q, P172S, P194A, P194G, P194H, P194L, P194M, P194Q, P194R, P194V, P194W, P194Y, P210A, P210G, P210L, P210S, P239K, P239R, Q002A, Q002S, Q010A, Q010N, Q010R, Q010T, Q019A, Q019C, Q019D, Q019G, Q019S, Q019T, Q019V, Q059I, Q059V, Q103S, Q185F, Q185H, Q185I, Q185K, Q185N, Q185S, Q185Y, Q206H, Q206L, Q206P, Q206W, Q217F, Q217H, Q217I, Q217K, Q217L, Q217N, Q217V, Q271G, Q271R, Q271T, Q275F, Q275P, Q275R, R186A, R186I, R186K, S003A, S003F, S003G, S003H, S003K, S003R, S003T, S009T, S018N, S018T, S037G, S037T, S037V, S038G, S038Q, S063N, S063Q, S063T, S089M, S089N, S130K, S130L, S130R, S130W, S132N, S145G, S145K, S145M, S145R, S145V, S159C, S159H, S159L, S159Q, S159R, S159T, S159W, S161A, S161C, S161G, S161H, S161I, S161K, S161P, S161Q, S161R, S162F, S162G, S162I, S162L, S162M, S162N, S162P, S162R, S163G, S173A, S173G, S182F, S182G, S182K, S182N, S182Q, S182V, S183A, S183M, S183Q, S183R, S183T, S188A, S188F, S188G, S188P, S188R, S188T, S188V, S190C, S204A, S204G, S204I, S204L, S204Q, S204R, S204V, S207G, S224A, S224T, S236C, S236D, S236E, S236G, S236N, S248A, S248F, S248K, S248M, S248T, S249A, S249R, S249T, S249V, S249W, S249Y, S260G, S260H, S260K, S260N, T022A, T022G, T022Q, T022S, T022V, T022Y, T055A, T055K, T158A, T158S, T208L, T208S, T208V, T220S, T242D, T242N, T242S, T244E, T244G, T244I, T244R, T244V, T244W, T253A, T253G, T253N, T253S, T254S, T254V, T255H, 12551, T255K, T255L, T255Q, T255R, T255S, T255Y, V004A, V004N, V004P, V004W, V008A, V008M, V026I, V045S, V045W, V051I, V081Q, V081T, V084C, V093I, V095C, V143A, V143E, V143F, V143N, V143S, V147I, V147L, V147S, V147T, V148I, V149C, V149I, V165M, V180A, V180C, V180I, V180L, V180T, V192A, V192C, V192I, V192S, V192T, V192Y, V198L, V203A, V203F, V203K, V203L, V203M, V203N, V203Y, W241Y, Y006A, Y006G, Y006H, Y006L, Y006N, Y006P, Y006Q, Y006T, Y021E, Y021K, Y021L, Y021N, Y021Q, Y021R, Y021S, Y021T, Y104F, Y104I, Y104V, Y171F, Y214F, Y214L, Y214W, Y262F, and Y262S, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN'-v3 and BPN'-v36, a PI value greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0544] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one amino acid substitution selected from the group consisting of A001D, A001M, A001N, A001R, A001T, A001V, A013C, A013G, A013S, A015D, A015F, A015L, A015M, A015P, A015Q, A074G, A074S, A085S, A085T, A085V, A088C, A088L, A088S, A088T, A088V, A133E, A133G, A133R, A137E, A144G, A144H, A144Q, A144T, A151C, A152S, A153G, A153S, A153V, A176C, A187G, A187Q, A187S, A200G, A228S, A228T, A230T, A230V, A231C, A231V, A232C, A232V, A272E, A272G, A272I, A272Q, A272R, A273L, A273V, A274L, A274Q, A274R, A274V, D036E, D259A, D259G, D259P, D259S, D259T, E156A, E156S, E156V, E195G, E251C, E251I, E251Q, F189Y, F261A, F261G, F261H, F261L, F261P, F261S, F261T, F261V, F261W, G020C, G020D, G020E, G020L, G020R, G024D, G024I, G024N, G024P, G024S, G053A, G053D, G053F, G053H, G053K, G053N, G053S, G053Y, G157A, G157S, G160A, G160L, G160P, G160R, G166A, G166L, G166V, G166W, G169A, G211E, G211P, G215A, G215D, G215E, G215H, G215V, G258D, G258S, H017F, H017I, H017L, H017M, H017V, H039C, H226S, H226Y, I011T, I011V, I031C, I031L, I031V, I079F, I079K, I079L, I079M, I079Q, I079R, I079T, I079V, I079W, I115L, I205A, I205C, I268L, K012R, K012T, K027R, K043A, K043E, K043F, K043H, K043I, K043M, K043N, K043Q, K043T, K043V, K043Y, K141H, K141Q, K141V, K170S, K213G, K213H, K213I, K213L, K213N, K213Q, K213T, K213W, K237I, K237N, K237R, K237T, K256C, K256E, K256S, K256T, K256V, K256W, K265H, K265R, L016E, L016F, L016I, L016S, L042I, L042M, L075A, L075H, L075Y, L082K, L082Q, L082T, L196M, L196V, L209A, L209C, L209E, L209G, L235W, L257C, L257H, L257Q, L257Y, L267E, L267F, L267R, L267S, M050Y, N025H, N025P, N025S, N056D, N061A, N061C, N061D, N061G, N061H, N061I, N061L, N061Q, N061R, N061V, N061W, N062A, N062S, N062V, N076A, N076E, N076L, N076M, N076Q, N076S, N076T, N076W, N078G, N078H, N078K, N078P, N078Q, N078T, N101D, N118A, N118T, N184E, N184H, N184I, N212A, N212D, N212E, N218C, N218D, N218E, N218M, N218R, N218V, N240W, N243A, N243G, N243P, N243S, N252D, N252E, N252L, N252T, N252V, N269S, P005A, P005D, P005M, P005Q, P014A, P014D, P014F, P014M, P014V, P040H, P040R, P040W, P040Y, P086A, P129T, P172E, P172G, P172R, P194E, P201A, P201G, P210E, P210V, Q002E, Q002G, Q010D, Q010F, Q010H, Q010I, Q010L, Q010S, Q019E, Q019H, Q019N, Q059A, Q059L, Q059R, Q059S, Q059T, Q185D, Q185E, Q206D, Q206S, Q206V, Q217G, Q245D, Q245E, Q245K, Q245M, Q245R, Q271A, Q271F, Q271P, Q271Y, Q275D, Q275H, Q275I, Q275L, Q275S, Q275W, R186H, R186L, R186V, R186W, S003D, S003E, S003M, S003P, S003V, S009A, S009E, S009G, S009I, S009K, S009M, S009P, S009W, S018A, S018G, S018I, S018L, S018P, S018R, S018V, S018W, S037A, S037D, S037Q, S037R, S037Y, S038D, S038H, S038K, S038M, S038R, S038T, S063A, S063G, S063K, S063M, S063R, S087A, S087D, S087F, S087G, S087Q, S087T, S089A, S089C, S089H, S089I, S089K, S089L, S089Q, S089R, S089T, S089Y, S130D, S130E, S145A, S145H, S145L, S159G, S161E, S161L, S161M, S161W, S162A, S162C, S162E, S162W, S163P, S173T, S182A, S182C, S182E, S182H, S182R, S182T, S183C, S183D, S183G, S183H, S188C, S188D, S188E, S188L, S190T, S191A, S204Y, S224C, S236A, S248C, S248D, S248G, S248I, S248N, S248Q, S248R, S248V, S249C, S249H, S249L, S249Q, S260A, S260C, S260E, S260P, S260Q, S260R, S260T, T022L, T022N, T022R, T055C, T055D, T055G, T055I, T055L, T055N, T055Q, T055S, T055V, T055Y, T071A, T071S, T158G, T158H, T158L, T158P, T158Q, T158R, T158V, T158Y, T164N, T164S, T244A, T244D, T244H, T244Q, T244S, T253Q, T253R, T253Y, T254A, T254G, T255A, T255D, T255E, V004E, V004G, V004R, V008C, V026A, V028L, V030I, V044C, V045D, V045E, V045H, V045M, V045Q, V045Y, V072L, V081L, V081S, V084A, V084I, V084M, V084T, V143C, V147C, V147Q, V149L, V165L, V180M, V192F, V192G, V192Q, V198I, V198M, V203C, V203E, V203H, V203I, V203Q, V203R, V203T, V203W, V270A, V270L, V270T, W241F, W241M, Y006C, Y006D, Y006E, Y006I, Y006M, Y006R, Y006S, Y006V, Y006W, Y021C, Y021D, Y021V, Y091W, Y167F, Y214V, Y262A, Y262C, Y262I, Y262M, Y262R, Y262T, Y262V, and Y262W, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and/or a PI value of about 1.0 relative to BPN'-v36 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0545] The following BPN'-v36 variants were determined to have a PI value of about 0.9 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one amino acid substitution selected from the group consisting of A001E, A015C, A015E, A048C, A048E, A073T, A085C, A085G, A088I, A088M, A114G, A137R, A187L, A187N, A187P, A187W, A216C, A230S, A273D, A273H, A273T, D036N, D099N, D259H, E156C, E156G, E156H, E156Q, F189H, F189R, F189S, F189T, F261C, F261D, F261E, G053E, G053M, G053Q, G131C, G131D, G157N, G160V, G215C, G215L, G258A, H039A, I011L, I079E, I268M, K012A, K012G, K012H, K012N, K027N, K043C, K043D, K043G, K043L, K043W, K136G, K141G, K141L, K141M, K141N, K141R, K170C, K170Q, K213V, K256D, K265G, K265N, K265Q, K265S, L082A, L082F, L082H, L082R, L082S, L090M, L233S, L235V, L257E, L257G, L257R, L257W, M222S, N025R, N062H, N062T, N076D, N076P, N078D, N078E, N078F, N078R, N078V, N269H, N269Q, P014K, P057A, P086F, P201T, Q002D, Q002I, Q002P, Q002V, Q019L, Q019P, Q059C, Q059D, Q059E, Q185W, Q271C, Q271D, Q271E, Q271L, Q271W, Q275G, R186M, S009C, S009L, S018D, S037E, S037H, S037K, S037L, S037P, S038P, S063C, S063D, S063F, S063L, S063Y, S087L, S087N, S087R, S087Y, S089D, S089F, S089G, S089W, S105T, S125A, S130C, S159D, S159P, S163A, S182P, S183P, S190A, S190G, S204E, S224G, S248E, S248H, S249E, S260V, S260Y, T055M, T055R, T055W, T158D, T158E, T164G, T164K, T164Q, T220A, T242G, T253E, T255C, T255G, V004D, V044L, V044P, V045C, V045G, V045L, V045N, V045R, V045V, V081A, V081G, V081H, V084S, V147A, V203D, V203G, V270C, V270P, V270S, W241L, Y104T, Y214Q, Y262D, Y262E, Y262G, Y262H, Y262L, Y262N, Y263G, and Y263W, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), and/or a PI value of about 0.9 relative to BPN'-v36 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0546] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one amino acid substitution selected from the group consisting of A001C, A142C, A187C, A216H, A273Q, A274H, D036Q, D036S, D099S, D197T, E156L, F189A, F189L, G053L, G053R, G157P, G178A, G258P, H039S, H238Y, K012C, K012E, K012L, K012W, K136E, K265Y, L075G, L075V, L082E, L126W, L257D, L257P, M050L, M222A, M222F, M222L, N056S, N062C, N062L, N062Y, N269C, P057W, Q002K, Q002L, Q217C, Q245A, Q245H, S018C, S038Y, S049C, S087C, S087K, S145D, S191G, T022P, T055E, T164A, T164R, V045K, V051H, V081R, V143G, V148L, V180S, V203S, V270G, Y214H, A187F, A273P, F189G, G046D, G146A, G157T, I031F, I175L, K012F, K027T, L042F, L233E, L233G, M222T, N062R, N184P, P005V, P005W, P129V, P239N, P239T, Q010W, Q059W, Q275A, V004T, V165C, A128H, A230G, D041C, H067T, K027S, K043R, L090T, N062Q, N117G, P225G, P225S, P239G, P239H, Q002R, S089E, V044A, V045I, A001P, A273R, D041N, D099A, D099H, D099Q, F058G, I111M, L042C, N118L, P239A, S049N, S089P, S173V, T242P, V044T, and V045T, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value of equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one, two, three, four, five, six or more amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 8
Cleaning Performance of Additional Combinatorial Variants Based on BPN'-v36 Parent
[0547] Additional combinatorial variants based on parent BPN'-v36 (BPN'-S24G-S53G-S78N-S101N-G128A-Y217Q) were made and provided by DNA 2.0. These variants were tested for their cleaning performance using BMI microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8, BMI microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 7, Egg microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8, and Grass microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8. Protein content was determined using TCA assay and protease activity was assayed using AAPF assay. All assays were performed as described in Example 1 and the Performance Indices were calculated relative to BPN'-v36 (with a PI value of 1.0).
[0548] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T-L257G, A116T-A128S, N061S-N109G-A128S-N243V-S260P, S009T-N109G-A128S-K141R-N243V, S009T-S018T-Y021N-N109G-A128S-K141R, and S162G-K256R, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence.
[0549] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T, A088T-A116T, A088T-G131H, A088T-K256R, A088T-N109G, A088T-N243V, A088T-Q103H, A088T-S162G, A088T-S248N, A088T-S249A, A088T-T158S, A116T, A116T-G131H, A116T-K256R, A116T-L257G, A116T-N243V, A116T-S162G, A116T-S248N, A116TS-249A, A116T-T158S, A128S-K256R, A128S-L257G, A128S-N243V-S248N-K256R, A128S-S162G, A128S-S248N, A128S-S249A, A128S-T158S, G024E-A116T, G024E-K256R, G024E-L257G, G024E-N109G, G024E-N243V, G024E-T158S, G131H, G131H-K256R, G131H-L257G, G131H-N243V-K256R, G131H-S162G, G131H-S248N, G131H-S249A, G131H-T158S, K043Y-A088T, K043Y-K256R, K043Y-N243V, K256R, K256R-L257G, L257G, N061G-N109G-N243V, N061P-N109G-G131H-N243V, N061P-N109G-N243V, N061S-A128S-N243V-S260P, N061S-N109G-A128S-N243V-S248N-K256R-S260P, N061S-N109G-A128S-S260P, N061S-N109G-N243V, N076D-K256R, N076D-L257G, N076D-N109G, N076D-T158S, N109A-A128S-N243V-K256R, N109G, N109G-A116T, N109G-A128S, N109G-A128S-G131H-N243V-S248N-K256R, N109G-A128S-N243V-K256R, N109G-A128S-N243V-S248A, N109G-A128S-N243V-S248A-K256R, N109G-A128S-N243V-S248N, N109G-A128S-N243V-S248N-K256R, N109G-A128S-N243V-S248N-K256R-L257G, N109G-A128S-S162G-N243V-S248N-K256R, N109G-A128S-S248N-K256R, N109G-A128S-T158S-N243V-S248N-K256R, N109G-G131H, N109G-K256R, N109G-L257G, N109G-N218S, N109G-N243P-S248A-K256R, N109G-N243P-S248N-K256R, N109G-N243V, N109G-N243V-K256R, N109G-N243V-S248A-K256R, N109G-N243V-S248N, N109G-N243V-S248N-K256R, N109G-S162G, N109G-S248N-K256R, N109G-S249A, N109G-T158S, N109Q-A128S-N243V-K256R, N109S-A128S-N243V-K256R, N218S-N243V, N243V, N243V-K256R, N243V-L257G, N243V-S248N, N243V-S248N-K256R, N243V-S249A, P040A-N109G-A128S-N243V-S248N-K256R, Q103H-A116T, Q103H-A128S, Q103H-G131H, Q103H-K256R, Q103H-L257G, Q103H-N109G, Q103H-N218S, Q103H-N243V, Q103H-S162G, Q103H-S248N, Q103H-S249A, Q103H-T158S, S009T-A128S-K141R-N243V, S009T-N109G-A128S-K141R, S009T-N109G-A128S-K141R-N243V-S248N-K256R, S009T-S018T-Y021N-A128S-K141R-N243V, S018T-Y021N-A128S-N243V, S018T-Y021N-N061S-A128S-N243V-S260P, S018T-Y021N-N061S-N109G-A128S-S260P, S018T-Y021N-N109G-A128S, S018T-Y021N-N109G-A128S-N243V, S018T-Y021N-N109G-A128S-N243V-S248N-K256R, S033T-N109G-A128S-N243P-S248N-K256R, S033T-N243V, S033T-Q103H, S033T-T158S, S063G, S063G-A088T, S063G-A128S, S063G-K256R, S063G-L257G, S063G-N076D, S063G-N109G, S063G-Q103H, S063G-S162G, S063G-S248N, S063G-T158S, S162G, S162G-L257G, S162G-N243V, S162G-S248N, S248N, S248N-L257G, S249A, T158S, T158S-L257G, T158S-N218S, T158S-N243V, T158S-S248N, and T158S-S249A, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and a PI value of about 1.0 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0550] The following BPN'-v36 variants were determined to have a PI value of about 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E-A088T, A001E-A116T, A001E-A128S-G131H-N243V, A001E-G131H-G169A-N243V, A001E-K256R, A001E-N109G, A001E-N243V, A001E-S033T, A001E-S033T-N109G-N218S, A001E-S033T-N109G-N243V, A001E-S162G, A001E-T158S, A088T-A128S, A088T-G169A, A088T-N218S, A088T-Q206D, A116T-G169A, A116T-N218S, A116T-Q206D, A128S, A128S-G131H, A128S-G169A, A128S-N218S, A128S-N243V, A128S-Q206D, G024E, G024E-A088T, G024E-A128S, G024E-G131H, G024E-K043Y, G024E-N218S, G024E-Q103H, G024E-S033T, G024E-S063G, G024E-S162G, G024E-S248N, G024E-S249A, G131H-G169A, G131H-N218S, G131H-N243V, G131H-Q206D, G169A, G169A-K256R, G169A-L257G, G169A-N218S, G169A-N243V, G169A-Q206D, G169A-S248N, G169A-S249A, K043Y, K043Y-A116T, K043Y-A128S, K043Y-G131H, K043Y-G169A, K043Y-L257G, K043Y-N109G, K043Y-N218S, K043Y-Q103H, K043Y-S063G, K043Y-S162G, K043Y-S248N, K043Y-S249A, K043Y-T158S, N076D, N076D-A088T, N076D-A128S, N076D-G131H, N076D-N218S, N076D-N243V, N076D-Q103H, N076D-S162G, N076D-S248N, N076D-S249A, N109G-G169A, N109G-Q206D, N109G-S248N, N218S, N218S-K256R, N218S-L257G, N218S-S248N, N218S-S249A, P040E-N109G-A128S-G131H, Q103H, Q103H-G169A, Q206D, Q206D-K256R, Q206D-L257G, Q206D-N218S, Q206D-N243V, Q206D-S248N, Q206D-S249A, S018T-Y021N-S033T-N109G-A128S-N243V-S248N-K256R, S033T, S033T-A088T, S033T-A116T, S033T-A128S, S033T-A128S-G131H-N243P, S033T-G131H, S033T-K043Y, S033T-K256R, S033T-L257G, S033T-N076D, S033T-N076D-A128S-N218S, S033T-N076D-N109G-A128S-N218S-N243V-S248N-K256R, S033T-N109G, S033T-N109G-A128S-N243V-S248N-K256R, S033T-N218S, S033T-P040E-Q103H-N109G, S033T-Q103H-A128S-G131H, S033T-Q206D, S033T-S063G, S033T-S162G, S033T-S248N, S033T-S249A, S063G-A116T, S063G-G131H, S063G-G169A, S063G-N109G-A128S-G131H, S063G-N218S, S063G-N243V, S063G-Q206D, S063G-S249A, S162G-G169A, S162G-N218S, S162G-Q206D, S162G-S249A, S248N-K256R, S248N-S249A, S249A-K256R, S249A-L257G, T158S-G169A, T158S-K256R, T158S-Q206D, and T158S-S162G, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), and/or a PI value of about 0.9 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0551] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E, A001E-A128S, A001E-G024E, A001E-G131H, A001E-G169A, A001E-L257G, A001E-N218S, A001E-Q103H, A001E-S063G, A001E-S248N, A001E-S249A, G024E-N076D, K043Y-N076D, K043Y-Q206D, N076D-A116T, N076D-G169A, N076D-Q206D, Q103H-Q206D, S033T-G169A, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243V, A001E-K043Y, A001E-N076D, A001E-N076D-N109G-A128S, A001E-Q206D, and G024E-Q206D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity.
The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0552] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A116T, A088T-N243V, G024E-A116T, K043Y, N076D-A116T, N218S-S248N, S033T-N243V, S033T-S063G, S248N-L257G, A001E-S249A, A088T-A116T, A088T-A128S, A088T-G131H, A088T-L257G, A088T-N109G, A088T-S248N, A088T-S249A, A116T-N243V, A116T-T158S, A128S, A128S-K256R, A128S-L257G, A128S-N243V, A128S-S248N, A128S-T158S, G024E-A088T, G024E-A128S, G024E-G131H, G024E-K256R, G024E-L257G, G024E-N218S, G024E-N243V, G024E-S162G, G024E-S249A, G024E-T158S, G131H, G131H-K256R, G131H-S249A, K043Y-A088T, K043Y-A116T, K256R, N076D-K256R, N109G, N109G-A116T, N109G-A128S, N109G-A128S-N243V-K256R, N109G-A128S-N243V-S248A, N109G-G131H, N109G-K256R, N109G-L257G, N109G-N218S, N109G-N243V, N109G-S248N, N218S-L257G, N243V, N243V-K256R, N243V-L257G, N243V-S248N, N243V-S249A, Q103H-A128S, Q103H-G131H, Q103H-K256R, Q103H-L257G, Q103H-N243V, Q103H-S248N, Q103H-S249A, Q103H-T158S, Q206D-N243V, S033T-A128S, S033T-K256R, S033T-N076D, S033T-N218S, S033T-S248N, S033T-T158S, S063G-A128S, S063G-K256R, S063G-N243V, S063G-S162G, S063G-T158S, S162G-K256R, S248N-K256R, S249A, T158S-N243V, and T158S-S249A, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0553] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E-A128S, A001E-G131H, A001E-K256R, A001E-N218S, A001E-N243V, A001E-S033T, A001E-S063G, A001E-S162G, A088T, A088T-K256R, A088T-N218S, A088T-Q103H, A088T-S162G, A088T-T158S, A116T-A128S, A116T-G131H, A116T-K256R, A116T-L257G, A116T-S162G, A116T-S248N, A116T-S249A, A128S-G169A, A128S-N218S, A128S-S162G, A128S-S249A, G024E, G024E-N109G, G024E-Q103H, G024E-S033T, G024E-S063G, G024E-S248N, G131H-L257G, G131H-N243V, G131H-S162G, G131H-T158S, G169A, G169A-L257G, G169A-S248N, K043Y-A128S, K043Y-G131H, K043Y-K256R, K043Y-L257G, K043Y-N109G, K043Y-N243V, K043Y-Q103H, K043Y-S063G, K043Y-S162G, K043Y-S248N, K043Y-S249A, K043Y-T158S, K256R-L257G, L257G, N061G-N109G-N243V, N061S-A128S-N243V-S260P, N061S-N109G-A128S-N243V-S260P, N061S-N109G-A128S-S260P, N076D-A088T, N076D-A128S, N076D-G169A, N076D-N218S, N076D-N243V, N076D-S162G, N076D-S248N, N076D-T158S, N109A-A128S-N243V-K256R, N109G-A128S-G131H-N243V-S248N-K256R, N109G-A128S-N243V-S248A-K256R, N109G-A128S-N243V-S248N-K256R-L257G, N109G-A128S-S162G-N243V-S248N-K256R, N109G-A128S-T158S-N243V-S248N-K256R, N109G-Q206D, N109G-S162G, N109G-S249A, N109G-T158S, N109Q-A128S-N243V-K256R, N109S-A128S-N243V-K256R, N218S, N218S-K256R, N218S-N243V, P040A-N109G-A128S-N243V-S248N-K256R, Q103H, Q103H-A116T, Q103H-G169A, Q103H-N109G, Q103H-N218S, Q103H-S162G, S009T-A128S-K141R-N243V, S009T-N109G-A128S-K141R, S009T-N109G-A128S-K141R-N243V, S009T-S018T-Y021N-N109G-A128S-K141R, S018I-Y021N-N109G-A128S, S018I-Y021N-N109G-A128S-N243V, S018T-Y021N-N109G-A128S-N243V-S248N-K256R, S033T-A088T, S033T-A116T, S033T-G131H, S033T-K043Y, S033T-L257G, S033T-N109G, S033T-Q103H, S033T-Q206D, S033T-S162G, S033T-S249A, S063G, S063G-A088T, S063G-A116T, S063G-L257G, S063G-N076D, S063G-N109G, S063G-N218S, S063G-Q103H, S063G-S248N, S063G-S249A, S162G, S162G-G169A, S162G-L257G, S162G-N218S, S162G-N243V, S162G-S248N, S162G-S249A, S248N, S248N-S249A, S249A-K256R, S249A-L257G, T158S, T158S-G169A, T158S-K256R, T158S-L257G, T158S-N218S, and T158S-S248N, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' protease (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of 1.0 relative to BPN'-v3, and a PI value of about 1.0 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0554] The following BPN'-v36 variants were determined to have a PI value of about 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E, A001E-A088T, A001E-A116T, A001E-G169A, A001E-L257G, A001E-N109G, A001E-S033T-N109G-N243V, A001E-T158S, A088T-G169A, A088T-Q206D, A116T-N218S, A128S-G131H, A128S-N243V-S248N-K256R, A128S-Q206D, G024E-K043Y, G024E-N076D, G024E-Q206D, G131H-G169A, G131H-N218S, G131H-N243V-K256R, G131H-Q206D, G131H-S248N, G169A-K256R, G169A-N218S, G169A-N243V, G169A-Q206D, K043Y-N076D, K043Y-N218S, N061P-N109G-G131H-N243V, N061P-N109G-N243V, N061S-N109G-A128S-N243V-S248N-K256R-S260P, N061S-N109G-N243V, N076D, N076D-G131H, N076D-L257G, N076D-N109G, N076D-Q103H, N076D-S249A, N109G-A128S-N243V-S248N, N109G-A128S-N243V-S248N-K256R, N109G-A128S-S248N-K256R, N109G-N243P-S248A-K256R, N109G-N243P-S248N-K256R, N109G-N243V-K256R, N109G-N243V-S248A-K256R, N109G-N243V-S248N, N109G-N243V-S248N-K256R, N109G-S248N-K256R, N218S-S249A, N243V-S248N-K256R, Q103H-Q206D, Q206D, Q206D-K256R, Q206D-N218S, Q206D-S248N, Q206D-S249A, S009T-N109G-A128S-K141R-N243V-S248N-K256R, S009T-S018T-Y021N-A128S-K141R-N243V, S018T-Y021N-A128S-N243V, S018T-Y021N-N061S-A128S-N243V-S260P, S018T-Y021N-N061S-N109G-A128S-S260P, S018T-Y021N-S033T-N109G-A128S-N243V-S248N-K256R, S033T, S033T-G169A, S033T-N076D-A128S-N218S, S033T-N076D-N109G-A128S-N218S-N243V-S248N-K256R, S033T-N109G-A128S-N243P-S248N-K256R, S033T-N109G-A128S-N243V-S248N-K256R, S063G-G131H, S063G-G169A, S162G-Q206D, and T158S-S162G, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value of about 0.9 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0555] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E-A128S-G131H-N243V, A001E-G024E, A001E-G131H-G169A-N243V, A001E-Q103H, A001E-S033T-N109G-N218S, A001E-S248N, A116T-G169A, A116T-Q206D, G169A-S249A, K043Y-G169A, N109G-G169A, P040E-N109G-A128S-G131H, Q206D-L257G, S033T-A128S-G131H-N243P, S033T-A128S-G131H-N243V, S033T-P040E-Q103H-N109G, S033T-Q103H-A128S-G131H, S063G-N109G-A128S-G131H, S063G-Q206D, T158S-Q206D, A001E-K043Y, A001E-N076D, A001E-Q206D, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243V, A001E-N076D-N109G-A128S, K043Y-Q206D, and N076D-Q206D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0556] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T-L257G, G024E-K256R, G024E-L257G, N109G-A116T, N109G-L257G, N243V-K256R, S033T-N109G, S033T-T158S, S063G-L257G, A001E-L257G, A088T-A128S, A088T-G169A, A088T-K256R, A088T-N109G, A088T-N218S, A088T-N243V, A088T-S248N, A088T-T158S, A116T, A116T-A128S, A116T-G131H, A116T-K256R, A116T-L257G, A116T-N218S, A116T-S162G, A116T-T158S, A128S, A128S-G169A, A128S-K256R, A128S-L257G, A128S-N218S, G024E, G024E-A128S, G024E-G131H, G024E-N109G, G024E-N243V, G024E-S033T, G024E-S063G, G024E-S248N, G024E-S249A, G024E-T158S, G131H, G131H-G169A, G131H-K256R, G131H-N218S, G131H-S249A, G169A, G169A-L257G, G169A-N243V, K043Y-A088T, K043Y-N109G, K256R, K256R-L257G, N061G-N109G-N243V, N076D-N109G, N109G, N109G-A128S, N109G-G131H, N109G-K256R, N109G-N218S, N109G-S162G, N109G-S248N, N109G-S249A, N109G-T158S, N218S, N218S-K256R, N218S-L257G, N218S-S248N, N243V, N243V-L257G, N243V-S248N, N243V-S249A, P040A-N109G-A128S-N243V-S248N-K256R, Q103H-K256R, Q103H-L257G, Q103H-N109G, S009T-S018T-Y021N-N109G-A128S-K141R, S033T-A088T, S033T-A116T, S033T-A128S, S033T-G131H, S033T-K043Y, S033T-K256R, S033T-L257G, S033T-N076D, S033T-N218S, S033T-N243V, S033T-Q103H, S033T-S063G, S033T-S162G, S033T-S248N, S033T-S249A, S063G, S063G-A088T, S063G-A116T, S063G-A128S, S063G-G131H, S063G-K256R, S063G-N109G, S063G-N218S, S063G-N243V, S063G-S248N, S063G-S249A, S063G-T158S, S162G-K256R, S162G-N218S, S162G-N243V, S162G-S248N, S162G-S249A, S248N, S249A, S249A-L257G, T158S, T158S-L257G, and T158S-N243V, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0557] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E, A001E-A116T, A001E-G131H, A001E-G169A, A001E-K256R, A001E-N109G, A001E-S033T-N109G-N243V, A001E-S063G, A001E-S248N, A001E-S249A, A001E-T158S, A088T, A088T-A116T, A088T-G131H, A088T-Q103H, A088T-Q206D, A088T-S162G, A088T-S249A, A116T-G169A, A116T-N243V, A116T-S248N, A116T-S249A, A128S-G131H, A128S-N243V, A128S-S162G, A128S-S248N, A128S-S249A, A128S-T158S, G024E-A088T, G024E-A116T, G024E-K043Y, G024E-N076D, G024E-N218S, G024E-Q103H, G024E-S162G, G131H-L257G, G131H-N243V, G131H-N243V-K256R, G131H-S162G, G131H-S248N, G131H-T158S, G169A-K256R, G169A-N218S, G169A-Q206D, G169A-S248N, G169A-S249A, K043Y, K043Y-A116T, K043Y-A128S, K043Y-G169A, K043Y-K256R, K043Y-L257G, K043Y-N076D, K043Y-N218S, K043Y-N243V, K043Y-S063G, K043Y-S248N, K043Y-S249A, K043Y-T158S, L257G, N061P-N109G-G131H-N243V, N061P-N109G-N243V, N061S-A128S-N243V-S260P, N061S-N109G-A128S-N243V-S248N-K256R-S260P, N061S-N109G-A128S-S260P, N061S-N109G-N243V, N076D, N076D-A088T, N076D-A116T, N076D-G131H, N076D-G169A, N076D-K256R, N076D-L257G, N076D-N218S, N076D-N243V, N076D-Q103H, N076D-S249A, N076D-T158S, N109A-A128S-N243V-K256R, N109G-A128S-G131H-N243V-S248N-K256R, N109G-A128S-N243V-K256R, N109G-A128S-N243V-S248A, N109G-A128S-N243V-S248A-K256R, N109G-A128S-N243V-S248N, N109G-A128S-N243V-S248N-K256R, N109G-A128S-N243V-S248N-K256R-L257G, N109G-A128S-S162G-N243V-S248N-K256R, N109G-G169A, N109G-N243P-S248A-K256R, N109G-N243V-K256R, N109G-N243V-S248A-K256R, N109G-N243V-S248N, N109G-N243V-S248N-K256R, N109G-S248N-K256R, N109Q-A128S-N243V-K256R, N109S-A128S-N243V-K256R, N218S-N243V, N218S-S249A, Q103H, Q103H-A116T, Q103H-A128S, Q103H-G131H, Q103H-G169A, Q103H-N218S, Q103H-N243V, Q103H-S162G, Q103H-S248N, Q103H-S249A, Q103H-T158S, Q206D, Q206D-L257G, Q206D-N218S, S009T-A128S-K141R-N243V, S009T-N109G-A128S-K141R, S009T-N109G-A128S-K141R-N243V, S009T-N109G-A128S-K141R-N243V-S248N-K256R, S009T-S018T-Y021N-A128S-K141R-N243V, S018T-Y021N-A128S-N243V, S018T-Y021N-N109G-A128S, S018T-Y021N-N109G-A128S-N243V, S018T-Y021N-N109G-A128S-N243V-S248N-K256R, S018T-Y021N-S033T-N109G-A128S-N243V-S248N-K256R, S033T, S033T-A128S-G131H-N243V, S033T-G169A, S033T-N109G-A128S-N243P-S248N-K256R, S033T-N109G-A128S-N243V-S248N-K256R, S033T-Q103H-A128S-G131H, S033T-Q206D, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243V, S063G-G169A, S063G-N076D, S063G-N109G-A128S-G131H, S063G-Q103H, S063G-S162G, S162G, S162G-G169A, S162G-L257G, S248N-K256R, S248N-L257G, S248N-S249A, S249A-K256R, T158S-G169A, T158S-K256R, T158S-N218S, T158S-S162G, T158S-S248N, and T158S-S249A, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' protease (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and a PI value of 1.0 relative to BPN'-v36 in this egg microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0558] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E-A088T, A001E-A128S, A001E-A128S-G131H-N243V, A001E-G024E, A001E-G024E-S204E-Q206D, A001E-G131H-G169A-N243V, A001E-K043Y, A001E-N076D, A001E-N076D-N109G-A128S, A001E-N218S, A001E-N243V, A001E-Q103H, A001E-Q206D, A001E-S033T, A001E-S033T-N109G-N218S, A001E-S162G, A116T-Q206D, A128S-N243V-S248N-K256R, A128S-Q206D, G024E-Q206D, G131H-Q206D, K043Y-G131H, K043Y-Q103H, K043Y-Q206D, K043Y-S162G, N061S-N109G-A128S-N243V-S260P, N076D-A128S, N076D-Q206D, N076D-S162G, N076D-S248N, N109G-A128S-S248N-K256R, N109G-A128S-T158S-N243V-S248N-K256R, N109G-N243P-S248N-K256R, N109G-Q206D, N243V-S248N-K256R, P040E-N109G-A128S-G131H, Q103H-Q206D, Q206D-K256R, Q206D-N243V, Q206D-S248N, Q206D-S249A, S018T-Y021N-N061S-A128S-N243V-S260P, S018T-Y021N-N061S-N109G-A128S-S260P, S033T-A128S-G131H-N243P, S033T-N076D-A128S-N218S, S033T-N076D-N109G-A128S-N218S-N243V-S248N-K256R, S033T-P040E-Q103H-N109G, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S063G-Q206D, S162G-Q206D, and T158S-Q206D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0559] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in a grass microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of T158S-L257G, K256R, L257G, S033T-N109G, S162G-K256R, S162G-L257G, G024E-K256R, G024E-L257G, G024E-S033T, N109G-A116T, N218S-L257G, S033T-A088T, S033T-A116T, S033T-N243V, S033T-Q103H, S162G-N218S, S162G-N243V, T158S, T158S-N218S, T158S-N243V, A088T, A088T-G169A, A088T-K256R, A088T-L257G, A088T-S162G, A088T-T158S, A116T-K256R, A116T-L257G, A116T-N243V, A128S-L257G, A128S-N218S, A128S-N243V, A128S-S248N, G024E-A116T, G024E-A128S, G024E-G131H, G024E-N243V, G024E-S248N, G024E-S249A, G024E-T158S, G131H-N243V, G131H-T158S, G169A-N218S, G169A-N243V, G169A-S248N, K256R-L257G, N109G-A128S, N109G-G131H, N109G-N218S, N109G-N243V, N109G-S249A, N218S, N218S-K256R, N218S-N243V, N218S-S249A, N243V, N243V-K256R, N243V-L257G, N243V-S248N, Q103H-N109G, Q103H-N218S, S033T-A128S, S033T-L257G, S033T-N218S, S033T-S162G, S033T-S248N, S033T-T158S, S063G-K256R, S063G-L257G, S162G, S162G-G169A, S162G-S248N, S248N, S248N-K256R, S248N-L257G, S249A, T158S-S162G, and T158S-S248N, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0560] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in a grass microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E-A088T, A001E-A116T, A088T-A128S, A088T-N243V, A088T-Q103H, A088T-S248N, A088T-S249A, A116T, A116T-G169A, A116T-N218S, A116T-S162G, A116T-S249A, A116T-T158S, A128S-G169A, A128S-K256R, A128S-S162G, A128S-S249A, A128S-T158S, G024E-A088T, G024E-K043Y, G024E-N218S, G024E-Q103H, G024E-S063G, G024E-S162G, G131H-G169A, G131H-K256R, G131H-N218S, G131H-S162G, G131H-S248N, G131H-S249A, G169A, G169A-L257G, G169A-S249A, N076D, N076D-K256R, N076D-L257G, N076D-S162G, N076D-S249A, N109G-K256R, N109G-L257G, N109G-S248N, N243V-S249A, Q103H-A116T, Q103H-G169A, Q103H-K256R, Q103H-L257G, Q103H-N243V, Q103H-S162G, S033T-G131H, S033T-G169A, S033T-K043Y, S033T-N076D, S033T-Q206D, S063G, S063G-A116T, S063G-A128S, S063G-N243V, S063G-S162G, S063G-S248N, S063G-S249A, S063G-T158S, S249A-L257G, T158S-G169A, and T158S-K256R, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of 1.0 relative to BPN'-v3, and a PI value of about 1.0 relative to BPN'-v36 in this BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0561] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a grass microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E-G169A, A001E-K256R, A001E-N109G, A001E-N218S, A088T-A116T, A088T-G131H, A088T-N109G, A088T-N218S, A116T-A128S, A116T-G131H, A116T-S248N, A128S, A128S-G131H, G024E, G024E-N109G, G131H, G131H-L257G, G169A-K256R, G169A-Q206D, K043Y, K043Y-A088T, K043Y-L257G, K043Y-N109G, N076D-A088T, N076D-G131H, N076D-G169A, N076D-N243V, N076D-T158S, N109G, N109G-S162G, N109G-T158S, N218S-S248N, Q103H-A128S, Q103H-S248N, Q103H-S249A, Q103H-T158S, Q206D-K256R, Q206D-L257G, Q206D-N218S, Q206D-N243V, Q206D-S248N, S033T, S033T-K256R, S033T-S063G, S033T-S249A, S063G-A088T, S063G-G131H, S063G-G169A, S063G-N109G, S162G-Q206D, S162G-S249A, S248N-S249A, S249A-K256R, T158S-Q206D, T158S-S249A, A001E-L257G, A001E-N243V, A001E-Q103H, A001E-S063G, A001E-S162G, A001E-T158S, G024E-N076D, G131H-Q206D, K043Y-A116T, K043Y-G169A, K043Y-K256R, K043Y-N076D, K043Y-S063G, K043Y-S162G, K043Y-S248N, K043Y-S249A, K043Y-T158S, N076D-A116T, N076D-A128S, N076D-S248N, N109G-G169A, Q103H, Q103H-G131H, Q206D-S249A, S063G-N076D, S063G-N218S, S063G-Q103H, A001E-A128S, A001E-G024E, A001E-G131H, A001E-N076D, A001E-Q206D, A001E-S033T, A001E-S248N, A001E-S249A, A088T-Q206D, A116T-Q206D, A128S-Q206D, G024E-Q206D, K043Y-A128S, K043Y-G131H, K043Y-N218 S, K043Y-Q103H, N076D-N109G, N076D-N218S, N076D-Q103H, N109G-Q206D, Q103H-Q206D, Q206D, A001E, A001E-K043Y, K043Y-N243V, and S063G-Q206D, N076D-Q206D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this grass microswatch cleaning assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0562] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an AAPF proteolytic assay: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of S033T-N076D-A128S-N218S, A001E-S033T-N109G-N218S, S033T-N218S, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243V, A128S-G169A, S033T-S063G-Q103H-N109Q-A128S-G131H-G169A-N243P, S018T-Y021N-S033T-N109G-A128S-N243V-S248N-K256R, S033T-A128S-G131H-N243P, P040E-N109G-A128S-G131H, S033T-A128S, S033T-N109G-A128S-N243V-S248N-K256R, N109G-G169A, S063G-N109G-A128S-G131H, G169A, N109G-A128S-G131H-N243V-S248N-K256R, S033T-A128S-G131H-N243V, A128S-N218S, A001E-G169A, A088T-G169A, G169A-L257G, N109G-N218S, S033T-N109G-A128S-N243P-S248N-K256R, G169A-K256R, N076D-G169A, A001E-G131H-G169A-N243V, G169A-S249A, S033T-N109G, G169A-S248N, K043Y-G169A, K043Y-N218S, N218S-L257G, N218S-N243V, S063G-G169A, A001E-A128S-G131H-N243V, A001E-S033T-N109G-N243V, A088T-N218S, G024E-N218S, G024E-S033T, G169A-Q206D, N076D-N218S, S033T-L257G, S162G-G169A, A001E-N218S, A116T-N218S, G169A-N243V, N218S, P040A-N109G-A128S-N243V-S248N-K256R, S033T-N076D, A001E-S033T, A128S-G131H, N218S-S248N, S018T-Y021N-N109G-A128S, S033T-K043Y, S033T-N243V, S033T-Q206D, S063G-N218S, S162G-N218S, T158S-G169A, A116T-G169A, G131H-G169A, N061S-N109G-A128S-S260P, N109G-A128S-N243V-K256R, N109G-A128S-N243V-S248A, N109G-A128S-N243V-S248A-K256R, N109G-A128S-N243V-S248N-K256R-L257G, N218S-K256R, S009T-N109G-A128S-K141R, S009T-S018T-Y021N-N109G-A128S-K141R, S033T-A088T, S033T-S063G, S033T-S162G, T158S-N218S, A001E-N076D-N109G-A128S, N109G-A128S-N243V-S248N-K256R, N109G-A128S-S248N-K256R, S009T-N109G-A128S-K141R-N243V, S018T-Y021N-N061S-N109G-A128S-S260P, S033T-A116T, S033T-S248N, S033T-S249A, S033T-T158S, G131H-N218S, N109A-A128S-N243V-K256R, N109G-A128S, N109G-A128S-S162G-N243V-S248N-K256R, N109G-A128S-T158S-N243V-S248N-K256R, N218S-S249A, Q206D-N218S, S018T-Y021N-N109G-A128S-N243V, S018T-Y021N-N109G-A128S-N243V-S248N-K256R, S033T-K256R, A116T-A128S, N061S-N109G-A128S-N243V-S260P, N109G-A128S-N243V-S248N, S009T-N109G-A128S-K141R-N243V-S248N-K256R, G024E-A128S, N061S-N109G-A128S-N243V-S248N-K256R-S260P, N109S-A128S-N243V-K256R, S033T, S033T-G131H, A001E-A128S, A128S, A128S-L257G, A128S-Q206D, N109Q-A128S-N243V-K256R, S009T-A128S-K141R-N243V, S009T-S018T-Y021N-A128S-K141R-N243V, A088T-A128S, A128S-K256R, A128S-N243V, N061P-N109G-N243V, N061S-A128S-N243V-S260P, S018T-Y021N-A128S-N243V, A128S-N243V-S248N-K256R, A128S-S248N, A128S-S249A, N076D-A128S, S063G-A128S, A128S-S162G, A128S-T158S, S018T-Y021N-N061S-A128S-N243V-S260P, S033T-Q103H-A128S-G131H, N061S-N109G-N243V, K043Y-A128S, N061P-N109G-G131H-N243V, N109G-L257G, A001E-G024E-S204E-Q206D, A001E-L257G, A088T-N109G, G024E-N109G, K043Y-N109G, N061G-N109G-N243V, N076D-N109G, N109G, N109G-A116T, N109G-K256R, N109G-N243V-K256R, N109G-N243V-S248A-K256R, N109G-Q206D, S063G-N109G, A001E-A116T, A001E-N109G, A001E-Q206D, A088T-A116T, A088T-N243V, A116T-L257G, G024E-A116T, G024E-L257G, G024E-N243V, G024E-Q206D, N109G-G131H, N109G-N243V, N109G-S162G, N109G-S248N, N109G-S248N-K256R, N109G-S249A, N109G-T158S, N243V-L257G, A001E-A088T, A001E-G024E, A001E-K256R, A001E-N076D, A001E-N243V, A088T, A088T-L257G, A088T-Q206D, A116T, A116T-K256R, A116T-N243V, G024E-A088T, G024E-K043Y, G024E-K256R, G024E-N076D, G024E-S162G, G024E-S248N, K043Y-A088T, K043Y-A116T, K043Y-L257G, K043Y-N243V, K043Y-Q206D, K256R-L257G, N076D-A116T, N076D-L257G, N076D-N243V, N076D-Q206D, N109G-N243V-S248N, N109G-N243V-S248N-K256R, N243V-K256R, Q206D, Q206D-L257G, Q206D-N243V, Q206D-S248N, S063G-K256R, S063G-L257G, T158S-L257G, A001E, A001E-K043Y, A001E-S162G, A001E-S248N, A001E-S249A, A001E-T158S, A088T-K256R, A088T-S162G, A088T-S248N, A088T-S249A, A116T-Q206D, A116T-S248N, A116T-S249A, G024E, G024E-G131H, G024E-S249A, G024E-T158S, G131H, G131H-K256R, G131H-L257G, K043Y-K256R, K043Y-N076D, K256R, L257G, N076D-A088T, N076D-K256R, N076D-S162G, N076D-S248N, N076D-S249A, N109G-N243P-S248A-K256R, N109G-N243P-S248N-K256R, N243V, Q206D-K256R, S033T-P040E-Q103H-N109G, S063G, S063G-A116T, S063G-Q206D, S162G-K256R, S162G-L257G, S162G-N243V, S162G-Q206D, S162G-S248N, S248N, S248N-L257G, S249A, S249A-L257G, T158S, T158S-N243V, and T158S-Q206D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0563] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in an AAPF proteolytic assay: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A001E-G131H, A001E-S063G, A088T-G131H, A088T-T158S, A116T-G131H, A116T-S162G, A116T-T158S, G024E-S063G, G131H-N243V, G131H-N243V-K256R, G131H-Q206D, G131H-S249A, K043Y, K043Y-S063G, K043Y-S248N, K043Y-S249A, K043Y-T158S, N076D, N076D-G131H, N076D-T158S, N243V-S248N, N243V-S248N-K256R, N243V-S249A, Q103H-G169A, Q206D-S249A, S063G-N076D, S063G-N243V, S063G-S162G, S063G-S249A, S063G-T158S, S162G, S162G-S249A, S248N-K256R, S248N-S249A, S249A-K256R, T158S-K256R, T158S-S248N, and T158S-S249A, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and a PI value of 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0564] The following BPN'-v36 variants were determined to have a PI value of about 0.9 relative to BPN'-v36 in an AAPF proteolytic assay: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of G131H-S162G, G131H-S248N, G131H-T158S, K043Y-G131H, K043Y-S162G, S063G-A088T, S063G-G131H, S063G-S248N, T158S-S162G, Q103H-N218S, S033T-Q103H, and Q103H-A128S, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value of about 0.9 relative to BPN'-v36 in this proteolytic assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0565] Also provided is a subtilisin protease variant having proteolytic activity, enhanced proteolytic activity compared to BPN', or a PI value greater than that of BPN' (SEQ ID NO:2) in a BMI microswatch cleaning assay, the variant comprising an amino acid sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:6, wherein the variant comprises at least one substitution selected from the group of X001E, X009T, X018T, X021N, X024G, X033T, X040A, X043Y, X061G/P/S, X063G, X076D, X088T, X103H, X109A/G/Q/S, X116T, X128S, X131H, X141R, X158S, X162G, X169A, X204E, X206D, X218S, X243P/V, X248A/N, X249A, X256R, X257G, and X260P, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2, and optionally wherein the variant comprises at least one substitution selected from the group of A001E, S009T, S018T, Y021N, S024G, S033T, P040A, K043Y, N061G/P/S, S063G, N076D, A088T, Q103H, N109A/G/Q/S, A116T, G128S, G131H, K141R, T158S, S162G, G169A, S204E, Q206D, N218S, N243P/V, S248A/N, S249A, K256R, L257G, and S260P, and wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
Example 9
Construction and Cleaning Performance of Variants from a Combinatorial Library Based on BPN'-v36 Parent
[0566] A BPN' combinatorial library based on the BPN'-v36 parent molecule was made by DNA 2.0 and delivered as a ligation reaction. For efficient transformation of B. subtilis, DNA from the ligation reaction mixture was amplified before transformation and transformants grown as described in Example 2. The variants were tested for cleaning performance using BMI microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8 and egg microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8. Protein content was determined using the TCA assay. Assays were performed as in Example 1 and Performance Indices were calculated relative to BPN'-v36 (i.e., BPN'-S24G-S53G-S78N-S101N-G128A-Y217Q) (with a PI value of 1.0).
[0567] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T-A116T-N243V-K256R-L257G, A088T-A116T-N243V-L257G, A088T-T158S-N218S-K256R, A088T-T158S-N218S-N243V-L257G, A088T-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-A153S-N218S-S248N-L257G, A088T-N109G-A116T-T158S-S248N-K256R-L257G, A088T-N109G-T158S-L257G, A114S-A116T-N218S-N243V-S248N-K256R-L257G, A116T-T158S-K256R, A088T-A116T-G131H-T158S-S248N-L257G, A088T-A116T-T158S, A088T-N109G-A116T-G131H-L257G, A088T-N109G-A116T-T158S-N243V-S248N-L257G, A088T-N109G-N243V-L257G, A088T-N109G-N243V-S248N, A088T-N109G-T158S-N243V-L257G, A088T-N109G-T158S-N243V-S248N-L257G, A116T-T158S-S248N-L257G, Y006H-A116T-G131H-S248N, A088T-A116T-G131H-T158S-N218S-N243V, A088T-A116T-G131H-T158S-N243V, A088T-A116T-G131H-T158S-N243V-K256R-L257G, A088T-A116T-N218S-N243V-K256R-L257G, A088T-A116T-S248N-K256R-L257G, A088T-A116T-T158S-N218S-N243V, A088T-A116T-T158S-N243V-K256R-L257G, A088T-A116T-T158S-N243V-S248N-L257G, A088T-G131H-N243V-S248N-K256R-L257G, A088T-N109G-A116T-T158S-L257G, A088T-N109G-A116T-T158S-N212D-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-T158S-S248N-L257G, A088T-N109G-G131H-V148A-N218S-N243V-K256R-L257G, A088T-N109G-K256R, A088T-N109G-N243V-S248N-L257G, A088T-N109G-T158S-K256R, A088T-N109G-T158S-N243V, A088T-T158S-N243V-K256R-L257G, A116T, A116T-N218S-N243V-L257G-N269S, A116T-T158S-K256R-L257G, N109G-A116T-K256R-L257G, N109G-A116T-N243V, N109G-A116T-T158S-N243V-K256R-L257G, N109G-G131H-L257G, N109G-G131H-S248N-K256R-L257G, N109G-G131H-T158S-K256R-L257G, S003P-A116T-T158S-S248N-K256R, T158S-S248N-K256R, A088T-A116T-G131H-N243V-K256R, A088T-A116T-G131H-S248N-K256R-L257G, A088T-A116T-G131H-V147A-T158S-N218S-N243V-S248N-L257G, A088T-A116T-S248N-L257G, A088T-A116T-T158S-N218S, A088T-A116T-T158S-N218S-K256R-L257G, A088T-A116T-T158S-N218S-L257G, A088T-G131H-N243V-L257G, A088T-G131H-T158S-S248N-L257G, A088T-L257G, A088T-N109G-A116T, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S-S248N-L257G, A088T-N109G-A116T-G131H-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N-K256R-L257G, A088T-N109G-A116T-N218S-N243V-K256R, A088T-N109G-A116T-N218S-N243V-L257G, A088T-N109G-A116T-N243V-S248N-K256R, A088T-N109G-A116T-N243V-S248N-K256R-L257G, A088T-N109G-A116T-T158S-L257G, A088T-N109G-A116T-T158S-N243V-L257G, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-W241R-S248N-K256R, A088T-N109G-K256R-L257G, A088T-N109G-L257G, A088T-N109G-N243V, A088T-N109G-N243V-K256R, A088T-N109G-N243V-K256R-L257G, A088T-N109G-S248N-K256R, A088T-N109G-T158S-N218S-K256R-L257G, A088T-N109G-T158S-N218S-N243V-S248N-K256R, A088T-N109G-T158S-N243V-K256R, A088T-N109G-T158S-N243V-K256R-L257G, A088T-N109G-T158S-N243V-S248N-A274D, A088T-N109G-T158S-S248N-L257G, A088T-T158S-K256R, A088T-T158S-N218S-N243V-K256R-L257G, A088T-T158S-N243V-L257G, A116T-G131H-N218S-N243V-S248N, A116T-G131H-S248N-L257G, A116T-S248N-K256R-L257G, A116T-T158S-N218S-N243V-K256R, A116T-T158S-N218S-S248N-L257G-Q271R, A116T-T158S-N243V-K256R-L257G, A116T-T158S-N243V-S248N-L257G, G131H-S248N, G131H-T158S-I234T-N243V-K256R, G131H-W241L-N243V-S248N-K256R, N109G-A116T-G131H-A137V-T158S-S248N-K256R-L257G, N109G-A116T-G131H-A151S-N218S-K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-K256R, N109G-A116T-G131H-T158S-N218S-S248N, N109G-A116T-G131H-T158S-N243V-S248N, N109G-A116T-S248N, N109G-A116T-T158S-L257G, N109G-A116T-T158S-N218S-W241R-N243V, N109G-A116T-T158S-N243V-S248N-L257G, N109G-A116T-T158S-S248N-K256R-L257G, N109G-A116T-T158S-S248N-L257G, N109G-G131H-N218S-L257G, N109G-G131H-N218S-S248N-K256R-L257G, N109G-G131H-T158S-N218S-S248N-K256R-L257G-A274T, N109G-K256R, N109G-N243V-L257G, N109G-T158S-N218S-K256R-L257G, N109G-T158S-N218S-L257G, N109G-T158S-S248N-K256R, P014L-A015L-L016C-H017T-S018L-Q019K-G020A-Y021T-T022L-G023E, S003F-A088T-N109G-A116T-T158S-N243V-K256R-L257G, V004A-A088T-A116T-T158S-N218S, V004A-N109G-A116T-G131H-S248N-K256R-L257G, V004L-A116T-N218S-N243V-S248N-L257G, Y006H-N109G-N218S-N243V-S248N, A001T-A116T-T158S-N243V-L257G, A088T-A116T, A088T-A116T-G131H-L257G, A088T-A116T-G131H-N218S-L257G, A088T-A116T-G131H-N218S-S248N-K256R-L257G, A088T-A116T-G131H-N218S-S248N-L257G, A088T-A116T-G131H-N243V-K256R-L257G, A088T-A116T-G131H-N243V-L257G, A088T-A116T-G131H-N243V-S248N, A088T-A116T-G131H-T158S-K256R-L257G, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S, A088T-A116T-G131H-T158S-N218S-N243V-K256R-A273T, A088T-A116T-G131H-T158S-N218S-S248N-K256R, A088T-A116T-G131H-T158S-N218S-S248N-L257G, A088T-A116T-G131H-T158S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-S248N, A088T-A116T-G131H-T158S-S248N-L257G, A088T-A116T-K256R, A088T-A116T-K256R-L257G, A088T-A116T-N218S-N243V-L257G, A088T-A116T-N243V-K256R, A088T-A116T-N243V-L257G, A088T-A116T-N243V-S248N-K256R-L257G, A088T-A116T-S248N-K256R, A088T-A116T-T158S-K256R, A088T-A116T-T158S-N218S, A088T-A116T-T158S-N218S-N243V-K256R, A088T-A116T-T158S-N218S-N243V-K256R-N269S, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-N243V-K256R, A088T-A116T-T158S-N243V-L257G, A088T-A116T-T158S-N243V-S248N-K256R, A088T-A116T-T158S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-S248N-K256R, A088T-A116T-V143A-N218S-S248N-K256R, A088T-A116T-V147I-T158S-N218S-N243V-L257G, A088T-G131H-K256R-L257G, A088T-G131H-N218S-N243V-S248N, A088T-G131H-N218S-S248N-L257G, A088T-G131H-S248N-K256R-L257G, A088T-G131H-T158S-L257G, A088T-G131H-T158S-N218S-K256R, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-N243V-L257G, A088T-G131H-T158S-N218S-S248N, A088T-G131H-T158S-N243V, A088T-G131H-T158S-N243V, A088T-G131H-T158S-N243V-S248N, A088T-G131H-T158S-N243V-S248N-K256R, A088T-G131H-T158S-N243V-S248N-L257G, A088T-I107T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-L257G, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V, A088T-N109G-A116T-G131H-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-L257G, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V, A088T-N109G-A116T-G131H-N243V-L257G, A088T-N109G-A116T-G131H-N243V-S248N-L257G, A088T-N109G-A116T-G131H-S248N, A088T-N109G-A116T-G131H-S248N-K256R, A088T-N109G-A116T-G131H-S248N-L257G, A088T-N109G-A116T-G131H-T158S-L257G, A088T-N109G-A116T-G131H-T158S-N218S, A088T-N109G-A116T-G131H-T158S-N218S-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218T-N243V, A088T-N109G-A116T-G131H-T158S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N-L257G, A088T-N109G-A116T-N218S, A088T-N109G-A116T-N218S-L257G, A088T-N109G-A116T-N218S-N243V, A088T-N109G-A116T-N218S-N243V-S248N-L257G, A088T-N109G-A116T-N218S-S248N-K256R, A088T-N109G-A116T-N218T-K256R, A088T-N109G-A116T-N218T-K256R-L257G, A088T-N109G-A116T-N243V, A088T-N109G-A116T-N243V-K256R-L257G, A088T-N109G-A116T-N243V-K256R-L257G-N269D, A088T-N109G-A116T-S248N-K256R, A088T-N109G-A116T-T158S, A088T-N109G-A116T-T158S-N218S-L257G, A088T-N109G-A116T-T158S-N218S-N243V, A088T-N109G-A116T-T158S-N218S-N243V-K256R, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-A116T-T158S-N218S-S248N, A088T-N109G-A116T-T158S-N243V, A088T-N109G-A116T-T158S-N243V-K256R, A088T-N109G-A116T-T158S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-S248N-L257G, A088T-N109G-G131H-L257G, A088T-N109G-G131H-N218S-K256R-L257G, A088T-N109G-G131H-N218S-N243V-K256R, A088T-N109G-G131H-N218S-N243V-L257G, A088T-N109G-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N243V, A088T-N109G-G131H-N243V-L257G, A088T-N109G-G131H-N243V-S248N-K256R, A088T-N109G-G131H-N243V-S248N-L257G, A088T-N109G-G131H-S248N-L257G, A088T-N109G-G131H-T158S-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N243V, A088T-N109G-G131H-T158S-N243V-K256R, A088T-N109G-G131H-T158S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N243V-L257G, A088T-N109G-L257G, A088T-N109G-N218S-K256R, A088T-N109G-N218S-N243V-S248N-L257G, A088T-N109G-N218S-S248N-K256R-L257G, A088T-N109G-N243V-K256R-L257G, A088T-N109G-N243V-S248N-K256R-L257G, A088T-N109G-N243V-S248N-L257G-I268V, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S-N218S-K256R, A088T-N109G-T158S-N218S-N243V-L257G, A088T-N109G-T158S-N218S-N243V-L257G, A088T-N109G-T158S-N243V-K256R-I268V, A088T-N109G-T158S-N243V-S248N-Q275R, A088T-N218S-N243V, A088T-N218S-N243V-S248N-K256R-L257G, A088T-N218S-S248N, A088T-N218S-S248N-L257G, A088T-N243V, A088T-N243V, A088T-N243V-K256R, A088T-N243V-L257G, A088T-S145T-T158S-S248N, A088T-T158S-L257G, A088T-T158S-N218S-S248N-L257G, A088T-T158S-N243V-K256R-L257G-Q271H, A088T-T158S-S248N, A088T-V143A-T158S-K256R, A116T-G131H-K256R, A116T-G131H-N218S, A116T-G131H-N243V, A116T-G131H-N243V-K256R, A116T-G131H-N243V-L257G, A116T-G131H-S248N-K256R, A116T-G131H-T158S-N218S-I234T-N243V-S248N-K256R, A116T-G131H-T158S-N243V-L257G, A116T-G131H-T158S-N243V-S248N-K256R, A116T-G131H-V143F-T158S-N218S, A116T-L257G, A116T-N218S, A116T-N218S-L257G, A116T-N218S-N243V-L257G, A116T-N243V, A116T-N243V-K256R, A116T-N243V-S248N, A116T-N243V-S248N-K256R-L257G, A116T-S248N, A116T-T158S, A116T-T158S-N218S-N243V, A116T-T158S-N218S-S248N, A116T-T158S-N243V, A116T-T158S-N243V-K256R, A116T-T158S-N243V-L257G, A116T-T158S-N243V-S248N, A116T-T158S-S248N-K256R-L257G, A116T-V149I-T158S-N243V-S248N-K256R-Q271H, G131H-N218S-N243V-L257G, G131H-N243V, G131H-N243V-S248N-K256R, G131H-T158S, G131H-T158S-N218S-N243V-K256R, G131H-T158S-N243V-K256R-L257G, G131H-T158S-N243V-S248N-L257G, N109G-A116T-G131H-N218S-K256R-L257G, N109G-A116T-G131H-N218S-L257G, N109G-A116T-G131H-N218S-N243V-K256R-L257G, N109G-A116T-G131H-N218S-S248N-K256R, N109G-A116T-G131H-N243V-K256R, N109G-A116T-G131H-N243V-L257G, N109G-A116T-G131H-N243V-S248N-K256R-L257G, N109G-A116T-G131H-S248N, N109G-A116T-G131H-S248N-I268V, N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, N109G-A116T-G131H-T158S-N218S-S248N-L257G, N109G-A116T-G131H-T158S-S248N, N109G-A116T-G131H-T158S-S248N-K256R, N109G-A116T-N218S, N109G-A116T-N218S-N243V-K256R, N109G-A116T-N218S-N243V-K256R-L257G, N109G-A116T-N218S-S248N-L257G, N109G-A116T-N243V-K256R, N109G-A116T-N243V-S248N-K256R-L257G, N109G-A116T-S248N-L257G, N109G-A116T-T158S-G211V-N243V-S248N-K256R, N109G-A116T-T158S-K256R-L257G, N109G-A116T-T158S-N218S, N109G-A116T-T158S-N218S-N243V-K256R-L257G, N109G-A116T-T158S-N218S-N243V-L257G, N109G-A116T-T158S-N218S-N243V-S248N-L257G, N109G-A116T-T158S-N218S-S248N-K256R-L257G, N109G-A116T-T158S-N243V, N109G-A116T-T158S-Q275R, N109G-G131H-A137V-T158S-N218S-S248N, N109G-G131H-N218S-K237N, N109G-G131H-N218S-N243V-K256R-L257G, N109G-G131H-N218S-S248N-K256R, N109G-G131H-N243V-K256R-L257G, N109G-G131H-S145F-N218S-N243V-K256R-L257G, N109G-G131H-S248N-K256R, N109G-G131H-S248N-L257G, N109G-G131H-T158S-K256R, N109G-G131H-T158S-N218S-N243V-K256R, N109G-G131H-T158S-N243V, N109G-G131H-T158S-N243V-K256R-L257G, N109G-G131H-T158S-N243V-L257G, N109G-G131H-T158S-S248N-L257G, N109G-G131H-T158S-S248N-Q271R, N109G-N218S-L257G, N109G-N218S-N243V, N109G-N243V-K256R-L257G, N109G-N243V-S248N-K256R-L257G, N109G-T158S-I268V, N109G-T158S-K256R, N109G-T158S-N218S-N243V-K256R-L257G, N109G-T158S-N218S-S248N-L257G, N109G-T158S-N243V, N109G-T158S-N243V-K256R-L257G, N109G-T158S-N243V-S248N, N109S-A116T-S248N, N218S, N218S-N243V-S248N-K256R-L257G, N218S-S248N-L257G, N243V, N243V-K256R, N243V-S248N-K256R, N243V-S248N-K256R-L257G, S105P-A116T-T158S-N218S-N243V-S248N-K256R, S248N, T158S-N243V-K256R, and T158S-N243V-L257G, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0568] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T, A088T-A116T-G131H-L257G, A088T-A116T-G131H-N218S-A274T, A088T-A116T-G131H-N218S-K256R, A088T-A116T-G131H-N218S-K256R-L257G, A088T-A116T-G131H-N218S-N243V, A088T-A116T-G131H-N218S-N243V-L257G, A088T-A116T-G131H-N218S-N243V-S248N-K256R, A088T-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-N218S-N243V-S248N-L257G, A088T-A116T-G131H-N218S-S248N, A088T-A116T-G131H-N218S-S248N-L257G, A088T-A116T-G131H-N243V, A088T-A116T-G131H-N243V-K256R, A088T-A116T-G131H-N243V-S248N, A088T-A116T-G131H-N243V-S248N-A274V, A088T-A116T-G131H-N243V-S248N-K256R-L257G, A088T-A116T-G131H-N243V-S248N-K256R-L257G, A088T-A116T-G131H-N243V-S248N-L257G, A088T-A116T-G131H-S248N-K256R, A088T-A116T-G131H-S248N-K256R-L257G, A088T-A116T-G131H-T158S-K256R, A088T-A116T-G131H-T158S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-K256R, A088T-A116T-G131H-T158S-N218S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-K256R, A088T-A116T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N, A088T-A116T-G131H-T158S-N218S-N243V-S248N, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N243V-K256R, A088T-A116T-G131H-T158S-N243V-K256R, A088T-A116T-G131H-T158S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-N243V-L257G, A088T-A116T-G131H-T158S-N243V-S248N, A088T-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-N243V-S248N-L257G, A088T-A116T-G131H-T158S-S248N-K256R, A088T-A116T-G131H-T158S-S248N-K256R-L257G, A088T-A116T-K256R, A088T-A116T-L257G, A088T-A116T-N218S, A088T-A116T-N218S-N243V-K256R, A088T-A116T-N218S-N243V-N269D, A088T-A116T-N218S-N243V-S248N, A088T-A116T-N218S-N243V-S248N-K256R-L257G, A088T-A116T-N218S-N243V-S248N-L257G, A088T-A116T-N218S-S248N, A088T-A116T-N218S-S248N-L257G, A088T-A116T-N243V-K256R-L257G, A088T-A116T-N243V-S248N-K256R, A088T-A116T-S248N, A088T-A116T-S248N-K256R, A088T-A116T-T158S-A216S-N218S-N243V-K256R-L257G, A088T-A116T-T158S-N218S-K256R, A088T-A116T-T158S-N218S-N243V-K256R, A088T-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-N218S-S248N, A088T-A116T-T158S-N243V-K256R, A088T-A116T-T158S-N243V-S248N, A088T-A116T-T158S-N243V-S248N-K256R, A088T-A116T-T158S-N243V-S248N-L257G, A088T-A116T-T158S-S248N, A088T-G131D-T158S-N243V-S248N, A088T-G131H-A138V-N218S-L257G, A088T-G131H-K256R, A088T-G131H-N218S-N243V-K256R, A088T-G131H-N218S-N243V-K256R, A088T-G131H-N218S-S248N, A088T-G131H-N218S-S248N-K256R-L257G, A088T-G131H-N218T-L257G, A088T-G131H-N243V-L257G, A088T-G131H-N243V-S248N-K256R, A088T-G131H-S248N, A088T-G131H-S248N-L257G, A088T-G131H-T158S-K256R, A088T-G131H-T158S-K256R-L257G, A088T-G131H-T158S-N218S, A088T-G131H-T158S-N218S, A088T-G131H-T158S-N218S-K256R-L257G, A088T-G131H-T158S-N218S-N243V-K256R, A088T-G131H-T158S-N218S-N243V-K256R, A088T-G131H-T158S-N218S-N243V-S248N, A088T-G131H-T158S-N218S-S248N, A088T-G131H-T158S-N218S-S248N-K256R, A088T-G131H-T158S-N218S-S248N-L257G-I268V, A088T-G131H-T158S-N243V-K256R, A088T-G131H-T158S-N243V-K256R-L257G, A088T-G131H-T158S-N243V-S248N, A088T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-S248N, A088T-G131H-T158S-S248N-K256R, A088T-N109G-A116T-G131H-K256R, A088T-N109G-A116T-G131H-K256R-L257G, A088T-N109G-A116T-G131H-K256R-L257G, A088T-N109G-A116T-G131H-N218S-K256R, A088T-N109G-A116T-G131H-N218S-K256R, A088T-N109G-A116T-G131H-N218S-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-L257G, A088T-N109G-A116T-G131H-N218S-S248N, A088T-N109G-A116T-G131H-N218S-S248N-L257G, A088T-N109G-A116T-G131H-N243V-K256R, A088T-N109G-A116T-G131H-N243V-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N243V-S248N-K256R, A088T-N109G-A116T-G131H-S248N-K256R-L257G, A088T-N109G-A116T-G131H-S248N-K256R-L257G, A088T-N109G-A116T-G131H-S248N-L257G, A088T-N109G-A116T-G131H-T158S-K256R, A088T-N109G-A116T-G131H-T158S-L257G, A088T-N109G-A116T-G131H-T158S-N218S, A088T-N109G-A116T-G131H-T158S-N218S-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218S-S248N, A088T-N109G-A116T-G131H-T158S-N218T-K256R, A088T-N109G-A116T-G131H-T158S-N243V, A088T-N109G-A116T-G131H-T158S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-V149A-T158S-N218S-K256R, A088T-N109G-A116T-K256R, A088T-N109G-A116T-N218S-K256R, A088T-N109G-A116T-N218S-N243V, A088T-N109G-A116T-N218S-N243V-K256R-L257G, A088T-N109G-A116T-N218S-N243V-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R, A088T-N109G-A116T-N218S-S248N, A088T-N109G-A116T-N218S-S248N-K256R-L257G, A088T-N109G-A116T-N243V-K256R, A088T-N109G-A116T-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N243V-S248N-L257G, A088T-N109G-A116T-N243V-S248N-L257G, A088T-N109G-A116T-S248N, A088T-N109G-A116T-S248N-K256R-L257G, A088T-N109G-A116T-S248N-L257G, A088T-N109G-A116T-T158S, A088T-N109G-A116T-T158S-K256R, A088T-N109G-A116T-T158S-N218S, A088T-N109G-A116T-T158S-N218S-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N, A088T-N109G-A116T-T158S-N218S-N243V-S248N, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-T158S-N218S-S248N, A088T-N109G-A116T-T158S-N218S-S248N-K256R, A088T-N109G-A116T-T158S-N218S-S248N-L257G, A088T-N109G-A116T-T158S-N218S-S248N-L257G, A088T-N109G-A116T-T158S-N243V-K256R, A088T-N109G-A116T-T158S-N243V-S248N, A088T-N109G-A116T-T158S-S248N, A088T-N109G-A116T-T158S-S248N-K256R, A088T-N109G-A137E-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-A152S-T158S-N218S-S248N-K256R, A088T-N109G-G131H-K256R-L257G, A088T-N109G-G131H-N218S, A088T-N109G-G131H-N218S, A088T-N109G-G131H-N218S-K256R, A088T-N109G-G131H-N218S-K256R, A088T-N109G-G131H-N218S-L257G, A088T-N109G-G131H-N218S-N243V, A088T-N109G-G131H-N218S-N243V-K256R, A088T-N109G-G131H-N218S-N243V-K256R-L257G, A088T-N109G-G131H-N218S-N243V-S248N-K256R, A088T-N109G-G131H-N218S-S248N, A088T-N109G-G131H-N218S-S248N-K256R, A088T-N109G-G131H-N218S-S248N-K256R, A088T-N109G-G131H-N218S-S248N-K256R-L257G, A088T-N109G-G131H-N243V-K256R, A088T-N109G-G131H-N243V-K256R-L257G, A088T-N109G-G131H-N243V-K256R-L257G, A088T-N109G-G131H-N243V-L257G, A088T-N109G-G131H-N243V-S248N-K256R, A088T-N109G-G131H-S248N-K256R, A088T-N109G-G131H-S248N-L257G, A088T-N109G-G131H-T158S, A088T-N109G-G131H-T158S-K256R, A088T-N109G-G131H-T158S-K256R-L257G, A088T-N109G-G131H-T158S-N218S-K256R, A088T-N109G-G131H-T158S-N218S-K256R, A088T-N109G-G131H-T158S-N218S-L257G, A088T-N109G-G131H-T158S-N218S-L257G, A088T-N109G-G131H-T158S-N218S-N243V, A088T-N109G-G131H-T158S-N218S-N243V-K256R, A088T-N109G-G131H-T158S-N218S-N243V-S248N, A088T-N109G-G131H-T158S-N218S-S248N-L257G, A088T-N109G-G131H-T158S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N243V-S248N, A088T-N109G-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N243V-S248N-L257G, A088T-N109G-G131H-T158S-N243V-S248N-L257G, A088T-N109G-G131H-T158S-S248N, A088T-N109G-G131H-T158S-S248N-L257G, A088T-N109G-G131H-V149A-K256R-L257G, A088T-N109G-G154A-N155P-E156T-G157L-T158M-S159E-G160E-S161L, A088T-N109G-K256R-L257G, A088T-N109G-N218S-K256R, A088T-N109G-N218S-N243V-L257G, A088T-N109G-N218S-N243V-S248N-K256R, A088T-N109G-N218S-N243V-S248N-K256R, A088T-N109G-N218S-S248N, A088T-N109G-N218S-S248N-L257G, A088T-N109G-N218S-S248N-L257G, A088T-N109G-N243V-S248N-K256R, A088T-N109G-S248N, A088T-N109G-S248N, A088T-N109G-T158S-N218S, A088T-N109G-T158S-N218S-K256R-Q271H, A088T-N109G-T158S-N218S-N243V, A088T-N109G-T158S-N218S-N243V-K256R-Q275R, A088T-N109G-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-T158S-N218S-S248N, A088T-N109G-T158S-N218S-S248N-K256R, A088T-N109G-T158S-N218S-S248N-N269D, A088T-N109G-T158S-N243V-K256R, A088T-N109G-T158S-N243V-S248N-K256R, A088T-N109G-T158S-N243V-S248N-K256R-L257G-N269D, A088T-N109G-T158S-N243V-S248N-L257G, A088T-N109G-T158S-S248N-K256R-L257G, A088T-N109G-T158S-S248N-L257G, A088T-N109G-V147A-N218S-N243V-K256R, A088T-N218S, A088T-N218S-K256R, A088T-N218S-L257G-I268V, A088T-N218S-N243V, A088T-N218S-N243V-K256R, A088T-N218S-N243V-K256R-L257G, A088T-N218S-N243V-L257G, A088T-N218S-N243V-S248N-K256R-L257G, A088T-N218S-N243V-S248N-L257G, A088T-N218S-N243V-S248N-N269S, A088T-N218S-S248N-K256R, A088T-N243V-S248N, A088T-N243V-S248N-K256R, A088T-N243V-S248N-K256R, A088T-N243V-S248N-K256R-L257G, A088T-N243V-S248N-L257G, A088T-S248N, A088T-S248N, A088T-S248N-K256R-L257G, A088T-S248N-L257G, A088T-S248N-L257G-I268V, A088T-T158S, A088T-T158S, A088T-T158S-N218S, A088T-T158S-N218S-K256R, A088T-T158S-N218S-L257G, A088T-T158S-N218S-N243V-K256R-I268V, A088T-T158S-N218S-N243V-K256R-L257G, A088T-T158S-N218S-N243V-S248N-L257G, A088T-T158S-N218S-S248N, A088T-T158S-N243V-K256R, A088T-T158S-N243V-K256R-L257G, A088T-T158S-N243V-S248N-K256R, A088T-T158S-N243V-S248N-L257G, A088T-T158S-N243V-S248N-L257G, A088T-T158S-S248N, A088T-T158S-S248N-L257G, A088T-V147A-K256R, A098S-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A116T-G131H-N218S-K256R, A116T-G131H-N218S-K256R-L257G, A116T-G131H-N218S-L257G, A116T-G131H-N218S-N243V-S248N-L257G, A116T-G131H-N218S-S248N-K256R-L257G, A116T-G131H-N243V-S248N, A116T-G131H-N243V-S248N-L257G, A116T-G131H-T158S-A231V-N243V-L257G, A116T-G131H-T158S-K256R, A116T-G131H-T158S-K256R-L257G, A116T-G131H-T158S-N218S-K256R, A116T-G131H-T158S-N218S-K256R-L257G, A116T-G131H-T158S-N218S-N243V, A116T-G131H-T158S-N218S-N243V-K256R, A116T-G131H-T158S-N218S-N243V-K256R-L257G, A116T-G131H-T158S-N218S-N243V-L257G, A116T-G131H-T158S-N218S-N243V-S248N-K256R, A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A116T-G131H-T158S-N218S-N243V-S248N-L257G, A116T-G131H-T158S-N218S-S248N-K256R, A116T-G131H-T158S-N218T-L257G, A116T-G131H-T158S-S248N-K256R, A116T-G131H-T158S-S248N-L257G, A116T-N218S-K256R, A116T-N218S-K256R-L257G, A116T-N218S-N243V-S248N-K256R, A116T-N218S-N243V-S248N-K256R-L257G, A116T-N218S-N243V-S248N-L257G, A116T-N218S-S248N, A116T-N218S-S248N-K256R, A116T-N218S-S248N-L257G, A116T-N243V-S248N-L257G, A116T-S248N-L257G, A116T-T158S-L257G-Q271R, A116T-T158S-N218S-L257G, A116T-T158S-N218S-N243V-K256R-L257G, A116T-T158S-N218S-S248N-K256R, A116T-T158S-N218S-S248N-K256R-L257G, A116T-T158S-N243V-S248N-K256R, A116T-T158S-N243V-S248N-K256R-L257G, G024S-G053S-N078S-G097A-N101S-A128S, G131H, G131H-N218S, G131H-N218S-K256R, G131H-N218S-N243V-K256R, G131H-N218S-N243V-K256R-L257G, G131H-N218S-N243V-S248N, G131H-N218S-N243V-S248N-K256R, G131H-N218S-S248N-K256R-L257G, G131H-N218S-S248N-L257G, G131H-N243V-K256R, G131H-N243V-K256R-L257G, G131H-N243V-S248N-K256R-L257G, G131H-S248N-K256R, G131H-T158S-K256R, G131H-T158S-K256R-L257G, G131H-T158S-N218S-K256R, G131H-T158S-N218S-K256R-L257G, G131H-T158S-N218S-N243V-K256R-L257G, G131H-T158S-N218S-N243V-S248N, G131H-T158S-N218S-S248N-K256R-L257G, G131H-T158S-N218S-S248N-L257G, G131H-T158S-N243V-K256R, G131H-T158S-N243V-S248N-K256R, G131H-T158S-S248N-L257G, G131H-V147I-N218S-S248N-K256R, I107T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, I107T-N109G-G131H-N218S-N243V-K256R-L257G, N109G, N109G-A116T-G131H, N109G-A116T-G131H-A144V-T158S-S248N-K256R-L257G, N109G-A116T-G131H-K256R-L257G, N109G-A116T-G131H-L257G, N109G-A116T-G131H-N218S-N243V-K256R, N109G-A116T-G131H-N218S-N243V-S248N-K256R, N109G-A116T-G131H-N243V, N109G-A116T-G131H-N243V-K256R-L257G, N109G-A116T-G131H-N243V-S248N, N109G-A116T-G131H-N243V-S248N-K256R, N109G-A116T-G131H-S248N-K256R, N109G-A116T-G131H-T158S-K256R, N109G-A116T-G131H-T158S-K256R-L257G, N109G-A116T-G131H-T158S-L257G, N109G-A116T-G131H-T158S-N218S, N109G-A116T-G131H-T158S-N218S-K256R-L257G, N109G-A116T-G131H-T158S-N218S-L257G, N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-G131H-T158S-N218S-S248N-K256R, N109G-A116T-G131H-T158S-N218S-S248N-K256R-L257G, N109G-A116T-G131H-T158S-N243V-L257G, N109G-A116T-G131H-T158S-S248N-L257G, N109G-A116T-G131H-V147A-T158S-N218S-K256R-L257G, N109G-A116T-G131H-V149A-T158S-N218S-N243V-S248N-L257G, N109G-A116T-K256R, N109G-A116T-N218S-K256R, N109G-A116T-N218S-N243V, N109G-A116T-N218S-N243V-L257G, N109G-A116T-N218S-N243V-S248N-I268V, N109G-A116T-N218S-N243V-S248N-K256R-L257G, N109G-A116T-N218S-N243V-S248N-L257G, N109G-A116T-N243V-S248N, N109G-A116T-N243V-S248N-K256R, N109G-A116T-N243V-S248N-L257G, N109G-A116T-S248N-K256R, N109G-A116T-T158S, N109G-A116T-T158S-N218S-N243V-S248N-K256R, N109G-A116T-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-T158S-N218S-S248N-K256R, N109G-A116T-T158S-N243V-L257G, N109G-A116T-T158S-N243V-S248N, N109G-A116T-T158S-S248N, N109G-A116T-T158S-S248N-K256R, N109G-G131H, N109G-G131H-K256R, N109G-G131H-N218S-K256R, N109G-G131H-N218S-K256R-L257G, N109G-G131H-N218S-N243V-L257G, N109G-G131H-N218S-N243V-S248N-K256R, N109G-G131H-N218S-N243V-S248N-K256R-L257G, N109G-G131H-N218S-N243V-S248N-L257G, N109G-G131H-N218S-S248N-L257G, N109G-G131H-N243V, N109G-G131H-N243V-K256R, N109G-G131H-N243V-S248N, N109G-G131H-N243V-S248N-K256R-L257G, N109G-G131H-N243V-S248N-L257G, N109G-G131H-T158S-N218S-K256R-L257G, N109G-G131H-T158S-N218S-L257G, N109G-G131H-T158S-N218S-N243V, N109G-G131H-T158S-N218S-N243V-K256R-L257G, N109G-G131H-T158S-N218S-N243V-S248N, N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, N109G-G131H-T158S-N218S-N243V-S248N-L257G, N109G-G131H-T158S-N218S-S248N-K256R-L257G, N109G-G131H-T158S-N243V-K256R-I268V, N109G-G131H-T158S-N243V-S248N, N109G-G131H-T158S-N243V-S248N-K256R, N109G-G131H-T158S-N243V-S248N-L257G, N109G-G131H-T158S-S248N, N109G-G131H-T158S-S248N-K256R-L257G, N109G-K141E-N218S-S248N-L257G, N109G-N218S, N109G-N218S-N243V-K256R, N109G-N218S-N243V-L257G, N109G-N218S-N243V-S248N-S260F,
N109G-N218S-S248N, N109G-N218S-S248N-K256R, N109G-N243V-K256R, N109G-N243V-S248N, N109G-N243V-S248N-K256R, N109G-N243V-S248N-L257G, N109G-N243V-S248N-L257G-Q275R, N109G-S182F-S204F-S207L-N218S-S236F-S248N-L257G, N109G-S248N-K256R, N109G-T158S-K256R-L257G, N109G-T158S-L257G, N109G-T158S-N218S-N243V-K256R, N109G-T158S-N218S-N243V-S248N, N109G-T158S-N218S-N243V-S248N-L257G, N109G-T158S-N243V-K256R, N109G-T158S-N243V-S248N-K256R, N109G-T158S-N243V-S248N-L257G, N109G-T158S-S248N-L257G, N218S-N243V-L257G, N218S-N243V-S248N-K256R, N243V-K256R-L257G, N243V-S248N-L257G-Q271R, P057Q-A088T-N109G-A116T-G131H-T158S-N218S-S248N, S003P-A116T-N218S-K256R, S003P-N109G-G131H-N218S-N243V-S248N-K256R-L257G, S248N-K256R-L257G, T158S-K256R-L257G, T158S-N218S-A272V, T158S-N218S-K256R-L257G, T158S-N218S-L233S, T158S-N218S-N243V, T158S-N218S-N243V-K256R-L257G, T158S-N218S-N243V-L257G, T158S-N218S-N243V-S248N-K256R, T158S-N218S-S248N-K256R, T158S-N243V, T158S-N243V-K256R-L257G, T158S-N243V-S248N, T158S-N243V-S248N-K256R-N269D, and V004A-N109G-A116T-T158S-N218S-S248N-L257G, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of 1.0 relative to BPN'-v3, and a PI value of about 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0569] The following BPN'-v36 variants were determined to have a PI value of about 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T-A098S-N218S-K256R, A088T-A116T-G131H-K256R, A088T-A116T-G131H-K256R-L257G-L267M, A088T-A116T-G131H-N218S-N243V-K256R, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-N218S-N243V-S248N, A088T-A116T-G131H-N218S-S248N, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-N243V, A088T-A116T-G131H-S248N, A088T-A116T-G131H-S248N-L257G, A088T-A116T-G131H-S248N-L257G, A088T-A116T-G131H-T158S-N218S, A088T-A116T-G131H-T158S-N218S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-A116T-G131H-T158S-N218S-S248N, A088T-A116T-G131H-T158S-N218S-S248N-K256R, A088T-A116T-K256R-L257G, A088T-A116T-N218S-I268V, A088T-A116T-N218S-K256R, A088T-A116T-N218S-N243V-Q271R, A088T-A116T-N218S-N243V-S248N-K256R, A088T-A116T-N218S-N243V-S248N-K256R-Q275R, A088T-A116T-N218S-S248N, A088T-A116T-N218S-S248N-K256R, A088T-A116T-N243V-S248N-K256R, A088T-A116T-T158S, A088T-A116T-T158S-N218S-K256R, A088T-A116T-T158S-N218S-N243V-L257G, A088T-A116T-T158S-N218S-N243V-S248N-L257G, A088T-A116T-T158S-N218S-S248N, A088T-A116T-T158S-N218S-S248N-K256R, A088T-A116T-T158S-N218S-S248N-K256R-L257G, A088T-A116T-T158S-N218S-S248N-L257G, A088T-A116T-T158S-S248N-L257G, A088T-G131H, A088T-G131H, A088T-G131H-N218S-K237R-K256R-L257G, A088T-G131H-N218S-K256R, A088T-G131H-N218S-K256R-L257G, A088T-G131H-N218S-N243V-K256R-L257G, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-N243V-S248N, A088T-G131H-N218S-N243V-S248N-K256R, A088T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-G131H-N218S-S248N, A088T-G131H-N243V, A088T-G131H-N243V-K256R, A088T-G131H-N243V-K256R-L257G, A088T-G131H-N243V-S248N, A088T-G131H-N243V-S248N, A088T-G131H-N243V-S248N-K256R, A088T-G131H-S248N, A088T-G131H-S248N-K256R, A088T-G131H-T158S-N218S-K256R-L257G, A088T-G131H-T158S-N218S-L257G, A088T-G131H-T158S-N218S-N243V, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-G131H-T158S-N218S-S248N-K256R, A088T-G131H-T158S-N218S-S248N-K256R-L257G, A088T-G131H-T158S-S248N-K256R-L257G, A088T-L257G, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-N269D, A088T-N109G-A116T-G131H-N218S-N243V-S248N-Q275R, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N, A088T-N109G-A116T-G131H-T158S, A088T-N109G-A116T-G131H-T158S-N218S-L257G-I268V, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-S248N, A088T-N109G-A116T-G131H-T158S-S248N, A088T-N109G-A116T-G131H-W241L-S248N-K256R-L257G, A088T-N109G-A116T-K256R, A088T-N109G-A116T-N218S-K256R-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R, A088T-N109G-A116T-N218S-S248N, A088T-N109G-A116T-T158S-N218S, A088T-N109G-A116T-T158S-N218S-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-T158S-N243V-S248N, A088T-N109G-A116T-T158S-N243V-S248N-K256R, A088T-N109G-G131H-A138V-T158S-N218S-N243V-S248N-L257G, A088T-N109G-G131H-K256R-L257G, A088T-N109G-G131H-N218S-N243V, A088T-N109G-G131H-N218S-N243V-S248N-L257G, A088T-N109G-G131H-N218S-S248N-K256R-L257G, A088T-N109G-G131H-N218S-S248N-K256R-L257G-Q275R, A088T-N109G-G131H-N243V-S248N, A088T-N109G-G131H-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S, A088T-N109G-G131H-T158S-L233S-N243V-S248N, A088T-N109G-G131H-T158S-N218S-N243V-K256R, A088T-N109G-G131H-T158S-N218S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-L257G, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-S248N-K256R, A088T-N109G-G131H-T158S-S248N-K256R-L257G, A088T-N109G-G131H-V149A-K256R-L257G, A088T-N109G-N218S-K256R-L257G, A088T-N109G-N218S-N243V-K256R, A088T-N109G-N218S-N243V-L257G, A088T-N109G-N218S-N243V-S248N, A088T-N109G-N218S-N243V-S248N-K256R-L257G, A088T-N109G-N218S-S248N-K256R, A088T-N109G-N243V-K256R, A088T-N109G-N243V-S248N-K256R, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S, A088T-N109G-T158S-K256R-L257G, A088T-N109G-T158S-N218S-N243V-K256R, A088T-N109G-T158S-N218S-N243V-K256R-L257G, A088T-N109G-T158S-N218S-N243V-S248N-K256R, A088T-N109G-T158S-N218S-N243V-S248N-L257G, A088T-N109G-T158S-N218S-S248N, A088T-N109G-T158S-S248N, A088T-N109G-T158S-S248N, A088T-N218S-N243V-K256R, A088T-N218S-N243V-L257G, A088T-N218S-N243V-S248N, A088T-N218S-N243V-S248N-K256R, A088T-N218S-N243V-S248N-K256R, A088T-N218S-S248N, A088T-N218S-S248N-L257G, A088T-S248N-K256R-L257G, A088T-T158S-K256R, A088T-T158S-N218S-N243V-K256R, A088T-T158S-N218S-N243V-L257G, A088T-T158S-N218S-N243V-S248N, A088T-T158S-N218S-N243V-S248N-K256R, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N218S-N243V-S248N-L257G, A088T-T158S-N218S-S248N, A088T-T158S-N218S-S248N-K256R, A088T-T158S-N218S-S248N-L257G, A088T-T158S-N218S-S248N-L257G-Q275K, A088T-T158S-N243V, A088T-T158S-N243V-K256R, A088T-T158S-N243V-S248N, A088T-T158S-S248N-K256R-L257G, A088T-T158S-S248N-L257G, A088T-V147I-N218S-N243V-K256R-L257G, A116T-G131H-L257G, A116T-G131H-N218S-N243V, A116T-G131H-N218S-N243V-K256R-L257G, A116T-G131H-N218S-N243V-S248N-K256R-L257G, A116T-G131H-N218S-S248N, A116T-G131H-N218S-S248N-K256R, A116T-G131H-N243V-S248N-K256R, A116T-G131H-T158S-N218S-N243V-S248N, A116T-G131H-T158S-N218S-S248N, A116T-G131H-T158S-N243V-K256R, A116T-G131H-V139I-N218S-N243V-S248N, A116T-K141E-N218S-N243V-S248N-K256R-L257G, A116T-K256R, A116T-N218S-N243V-S248N, A116T-N218T-N243V-S248N, A116T-N243V-K256R-L257G, A116T-S248N-K256R, A116T-T158S-N218S-K256R, A116T-T158S-N218S-K256R-L257G, A116T-T158S-N218S-N243V-L257G, A116T-T158S-N218S-N243V-S248N, A116T-T158S-N218S-S248N-L257G, G024S-G053S-N078S-G097A-N101S, G053S-A088T-N109G-A116T-G131H-T158S-G169S-N218S-S248N-K256R-L257G, G131H-K141R-T158S-N218S-K256R, G131H-K256R, G131H-N218S-K256R-L257G, G131H-N218S-N243V-S248N-L257G, G131H-N218S-S248N-K256R, G131H-N243V-S248N, G131H-N243V-S248N-L257G, G131H-T158S-N218S, G131H-T158S-N218S-N240H-N243V-S248N-K256R-L257G, G131H-T158S-N218S-N243V, G131H-T158S-N218S-S248N-K256R-L257G-N269S, G131H-T158S-N243V-L257G, G131H-T158S-N243V-S248N, G131H-T158S-S248N, K256R, K256R-L257G, N109G-A116T-G131H-N218S-N243V, N109G-A116T-G131H-N218S-N243V-L257G, N109G-A116T-G131H-N218S-S248N-L257G, N109G-A116T-G131H-N218S-W241R-N243V-K256R, N109G-A116T-G131H-S248N-L257G, N109G-A116T-G131H-T158S-N218S-K256R, N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-S248N, N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, N109G-A116T-G131H-T158S-N243V-K256R-L257G, N109G-A116T-G131H-T158S-N243V-S248N-K256R, N109G-A116T-G131H-T158S-S248N-K256R-L257G, N109G-A116T-I234T-N243V-S248N-K256R-L257G, N109G-A116T-N218S-N243V-S248N, N109G-A116T-N243V-K256R-L257G, N109G-A116T-T158S-N218S-K237R-N243V-S248N, N109G-G131H-N218S-N243V-S248N, N109G-G131H-S248N, N109G-G131H-T158S, N109G-G131H-T158S-L257G, N109G-G131H-T158S-N218S-N243V-S248N-K256R, N109G-G131H-T158S-N218S-S248N-K256R, N109G-G131H-T158S-N218S-S248N-L257G, N109G-G131H-T158S-N243V-S248N-K256R-L257G, N109G-G131H-T158S-S248N-K256R, N109G-N218S-K256R-L257G, N109G-N218S-N243V-S248N-K256R, N109G-N218S-S248N-L257G, N109G-S248N, N109G-T158S-N218S, N109G-T158S-N218S-N243V, N109G-T158S-N243V-L257G, N218S-K256R, N218S-N243V-K256R, N218S-N243V-S248N, N218S-S248N, N218S-S248N-K256R, N243V-L257G, S003P-N109G-A116T-G131H-N218S-N243V-S248N, S003P-N109G-A116T-G131H-T158S-N218S-K256R, S105H-W106G-I107L-I108S-N109A-G110A-I111S-E112N-W113G-A114P, S248N-L257G, T158S, T158S-K256R, T158S-N218S, T158S-N218S-K256R, T158S-N218S-L233S-S248N, T158S-N218S-L257G, T158S-N218S-N243V-K256R, T158S-N218S-N243V-S248N, T158S-N218S-N243V-S248N-L257G, T158S-N218S-S248N-K256R-L257G, T158S-N218S-S248N-L257G, T158S-N243V-S248N-L257G, T158S-S248N, and V004L-A088T-G131H-T158S-N218S-S248N-L257G, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value of about 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0570] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A015S-A088T-N109G-G131H-T158S-N218S-S248N, A088T-A098S-G131H-S248N-K256R-L257G, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-A116T-N218S-L257G, A088T-A116T-T158S-K256R, A088T-A116T-T158S-S248N-K256R-L257G, A088T-G131H-K141E-N218S-N243V-S248N-L257G, A088T-G131H-K256R, A088T-G131H-N218S-K256R, A088T-G131H-N218S-N243V-S248N-K256R, A088T-G131H-N218S-N243V-S248N-L257G, A088T-G131H-N218S-S248N-K256R, A088T-G131H-N218S-S248N-K256R, A088T-G131H-T158S-S248N-K256R, A088T-G131H-T158S-S248N-K256R-L257G, A088T-I107T-N109G-G131H-N218S-S248N-K256R, A088T-N109G-A116T-G131H-D140G-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-N218S-N243V-K256R, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R-I268V, A088T-N109G-A116T-G131H-V149A-N218S-S248N-K256R-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-T158S-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-D140G-N243V, A088T-N109G-G131H-D140G-T158S-N243V-S248N-K256R, A088T-N109G-G131H-K141E-T158S-N218S-K256R, A088T-N109G-G131H-N218S-S248N, A088T-N109G-G131H-N218S-S248N-K256R-Q271R, A088T-N109G-G131H-N218S-S248N-L257G, A088T-N109G-G131H-T158S-K256R, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-G131H-V149L-T158S-K256R-L257G, A088T-N109G-T158S-N218S, A088T-N109G-T158S-N218S-K256R-L257G-Q271K, A088T-N109G-T158S-N218S-L257G, A088T-N109G-T158S-S248N-K256R, A088T-N218S-S248N-L257G-Q271R, A088T-T158S-N218S-K256R-L257G, A088T-T158S-N218S-N243V-K256R, A088T-Y104H-A116T-G131H-N218S-N243V, A116T-G131H-K141E-N218S-N243V-S248N-L257G, A116T-G131H-N218S-N243V-S248N-K256R, A116T-G131H-T158S-N218S-S248N-L257G-N269D, A116T-G131H-T158S-N218S-S248N-Q271R, A116T-G131H-T158S-N243V-S248N, A116T-G157E-T158S-N243V-S248N-K256R, A116T-T158S-N218S, G131H-N218S-L257G, G131H-N218S-S248N, G131H-T158S-N218S-N243V-S248N-K256R-L257G, G131H-T158S-N218S-N243V-S248N-L257G, G131H-T158S-N218S-S248N-I268V, I107T-N109G-G131H-N218S-L257G, L090I-N109G-T158S-N243V, L257G, N109G-A116T-G131H-T158S-N218S-K256R-L257G-Q271R, N109G-A116T-N218S-W241R-N243V-S248N-K256R-L257G, N109G-G131H-K141E-L257G, N109G-G131H-N218S-N243V, N109G-T158S-N218S-N243V-L257G, N109G-T158S-N218S-S248N-K256R, N109G-T158S-N243V-S248N-K256R-L257G, N218S-S248N-K256R-L257G, S003P-N109G-G131H-T158S-L257G, S003P-S248N-L257G, T158S-S248N-K256R-L257G, V004A-A088T-G131H-N218S-N243V-S248N-L257G, Y006H-N218S-N243V-S248N, Y104H-N109G-G131H-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N-K256R, A088T-A116T-T158S-N243V, A088T-G131H-T158S-N218S-I234T-S248N-L257G, A088T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-G131H-V149L-T158S-N243V-S248N-K256R-L257G, A088T-I107T-N109G-G131H-N218S-A223G-S248N-K256R, A088T-K213N-N243V-S248N-K256R, A088T-K256R-L257G, A088T-N109G-A116T-G131H-A232S-N243V-K256R, A088T-N109G-A116T-G131H-D140G-S248N-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-L257G, A088T-N109G-A116T-M124I-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-V148A-N218S-N243V, A088T-N109G-G131H-N218S-N243V-S248N, A088T-N109G-N218S-S248N-T255K-K256R-L257G, A088T-T158S-N218S-L257G, A088T-T158S-N218S-Q245K-S248N-K256R, A088T-T158S-N218S-S248N-K256R, A116T-G131H-N218S-N243V-K256R, A116T-G131H-N218S-W241R-N243V-S248N-K256R-L257G, A116T-G131H-T158S-N218S-L257G, A116T-G131H-V150A-T158S-N243V-S248N-K256R-L257G, I107T-G131H-T158S-N243V-S248N-K256R-L257G, N109G-A116T-K141E-T158S-N218S-N243V-L257G, N109G-A116T-T158S-N218S-N243V-S248N, T158S-N243V-S248N-K256R, T158S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-G146C, A088T-A116T-N218S, A088T-A116T-T158S-N243V-K256R-L257G, A088T-A138E-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243F-S248N, A088T-T158S-V203I-N218S-K256R-L257G, A116T-D140G-T158S-N218S-N243V-S248N, A088T-A116T-T158S-K256R-L257G, A088T-A116T-T158S-N218S-N243V-S248N-E251K-K256R-L257G, A088T-I108T-N109G-G131H-T158S-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-K141E-N218S, A088T-N109G-W241R-S248N-K256R, and G065D-A088T-G131H-N243V-S248N, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0571] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T-N109G-A116T-T158S-N243V-L257G, A116T-N218S-N243V-L257G-N269S, A088T-A116T-K256R, A088T-G131H-K256R, A088T-N109G-A116T-T158S-S248N-K256R-L257G, A088T-N109G-T158S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-K256R-A273T, A088T-A116T-N243V-L257G, A088T-A116T-S248N-K256R-L257G, A088T-A116T-T158S-N243V-L257G, A088T-A116T-T158S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-L257G, A088T-N109G-A116T-G131H-T158S-L257G, A088T-N109G-A116T-T158S-N218S-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N218S-K256R-L257G, A088T-N109G-N218S-S248N-L257G, A088T-T158S-N218S-N243V-K256R-I268V, A088T-T158S-N218S-S248N-L257G, A116T-N218S-K256R-L257G, N109G-A116T, N109G-A116T-G131H-T158S-L257G, N109G-A116T-N243V, N109G-A116T-N243V-K256R, N109G-A116T-T158S-L257G, N109G-K256R, N109G-N243V-K256R-L257G, S003P-N109G-G131H-T158S-K256R, A088T-A116T, A088T-A116T-G131H-N218S-K256R-L257G, A088T-A116T-G131H-N218S-L257G, A088T-A116T-G131H-N218S-N243V-S248N-L257G, A088T-A116T-G131H-N243V-K256R-L257G, A088T-A116T-G131H-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V, A088T-A116T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N, A088T-A116T-G131H-T158S-S248N-K256R-L257G, A088T-A116T-G131H-T158S-S248N-L257G, A088T-A116T-N218S-N243V-L257G, A088T-A116T-N218S-N243V-S248N-K256R-L257G, A088T-A116T-N218S-N243V-S248N-K256R-Q275R, A088T-A116T-T158S-A216S-N218S-N243V-K256R-L257G, A088T-A116T-T158S-K256R, A088T-A116T-T158S-N218S-L257G, A088T-A116T-T158S-N218S-N243V, A088T-A116T-T158S-N218S-N243V-K256R, A088T-A116T-T158S-N218S-N243V-K256R-L257G, A088T-A116T-T158S-N218S-N243V-K256R-N269S, A088T-A116T-T158S-N243V, A088T-A116T-T158S-N243V-K256R, A088T-A116T-V147I-T158S-N218S-N243V-L257G, A088T-G131H-K256R-L257G, A088T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-G131H-S248N-K256R-L257G, A088T-G131H-T158S-N218S-L257G, A088T-G131H-T158S-N218S-N243V-L257G, A088T-I107T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-I107T-N109G-G131H-N218S-S248N-K256R, A088T-N109G-A116T-G131H-A153S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-K256R-L257G, A088T-N109G-A116T-G131H-N218S-K256R-L257G, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V-K256R, A088T-N109G-A116T-G131H-N218S-N243V-L257G, A088T-N109G-A116T-G131H-N243V-L257G, A088T-N109G-A116T-G131H-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218S, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-N218S-K256R-L257G, A088T-N109G-A116T-N218S-L257G, A088T-N109G-A116T-N218S-N243V, A088T-N109G-A116T-N218S-N243V-L257G, A088T-N109G-A116T-N218T-K256R, A088T-N109G-A116T-N218T-K256R-L257G, A088T-N109G-A116T-N243V, A088T-N109G-A116T-N243V-K256R-L257G, A088T-N109G-A116T-N243V-K256R-L257G-N269D, A088T-N109G-A116T-T158S, A088T-N109G-A116T-T158S, A088T-N109G-A116T-T158S-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N, A088T-N109G-A116T-T158S-N243V-S248N-L257G, A088T-N109G-G131H-A138V-T158S-N218S-N243V-S248N-L257G, A088T-N109G-G131H-L257G, A088T-N109G-G131H-N218S, A088T-N109G-G131H-N218S-N243V-L257G, A088T-N109G-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N218S-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-K256R, A088T-N109G-G131H-T158S-N218S-N243V, A088T-N109G-G131H-T158S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N243V-L257G, A088T-N109G-G131H-T158S-N243V-S248N-L257G, A088T-N109G-G131H-V149A-K256R-L257G, A088T-N109G-N218S-N243V-L257G, A088T-N109G-N218S-N243V-S248N-L257G, A088T-N109G-N218S-S248N-K256R-L257G, A088T-N109G-N243V, A088T-N109G-N243V-K256R-L257G, A088T-N109G-N243V-L257G, A088T-N109G-N243V-S248N-L257G, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S-K256R, A088T-N109G-T158S-N218S-N243V-K256R-Q275R, A088T-N109G-T158S-N243V, A088T-N109G-T158S-N243V-K256R-I268V, A088T-N109G-T158S-N243V-L257G, A088T-N109G-T158S-N243V-S248N-L257G, A088T-N218S-N243V-L257G, A088T-N218S-N243V-S248N-K256R-L257G, A088T-N218S-S248N, A088T-T158S-N218S-N243V-K256R-L257G, A088T-V143A-T158S-K256R, A088T-V147I-N218S-N243V-K256R-L257G, A114S-A116T-N218S-N243V-S248N-K256R-L257G, A116T-G131H-N218S-N243V-S248N-K256R-L257G, A116T-G131H-N218S-N243V-S248N-L257G, A116T-G131H-N243V-K256R, A116T-G131H-N243V-L257G, A116T-G131H-N243V-S248N-K256R, A116T-G131H-S248N-L257G, A116T-G131H-T158S-N218S-I234T-N243V-S248N-K256R, A116T-G131H-T158S-N218S-N243V-S248N-L257G, A116T-N218S-L257G, A116T-T158S-N218S-K256R-L257G, A116T-T158S-N218S-S248N, A116T-T158S-N218S-S248N-L257G-Q271R, A116T-T158S-N243V-S248N-L257G, A116T-T158S-S248N-L257G, G131H-N218S-S248N-K256R-L257G, I107T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-G131H-A137V-T158S-S248N-K256R-L257G, N109G-A116T-G131H-A151S-N218S-K256R-L257G, N109G-A116T-G131H-N218S-K256R-L257G, N109G-A116T-G131H-N218S-N243V-L257G, N109G-A116T-G131H-N243V-L257G, N109G-A116T-G131H-S248N-L257G, N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-K256R-L257G, N109G-A116T-N218S-N243V-K256R, N109G-A116T-N218S-N243V-S248N-K256R-L257G, N109G-A116T-N218S-N243V-S248N-L257G, N109G-A116T-N243V-S248N-K256R, N109G-A116T-S248N-L257G, N109G-A116T-T158S-G211V-N243V-S248N-K256R, N109G-A116T-T158S-N218S, N109G-A116T-T158S-N218S-N243V-L257G, N109G-A116T-T158S-N218S-N243V-S248N, N109G-G131H-N218S-K237N, N109G-G131H-N218S-K256R-L257G, N109G-G131H-N218S-L257G, N109G-G131H-N218S-N243V-L257G, N109G-G131H-N243V-K256R-L257G, N109G-G131H-N243V-S248N-L257G, N109G-G131H-S248N-K256R, N109G-G131H-T158S-K256R-L257G, N109G-G131H-T158S-N218S-K256R-L257G, N109G-G131H-T158S-N243V, N109G-N218S, N109G-N218S-N243V-L257G, N109G-T158S-N218S-K256R-L257G, N109G-T158S-N218S-L257G, N109G-T158S-N218S-N243V-K256R, N109G-T158S-N243V-L257G, N243V-K256R-L257G, N243V-L257G, S003F-A088T-N109G-A116T-T158S-N243V-K256R-L257G, T158S-N218S-L233S, T158S-N218S-N243V-S248N-K256R, V004A-N109G-A116T-T158S-N218S-S248N-L257G, Y006H-A116T-G131H-T158S-N218S-N243V-S248N-K256R-A272G, A088T-A098S-N218S-K256R, A088T-A116T-G131H-K256R-L257G-L267M, A088T-A116T-G131H-L257G, A088T-A116T-G131H-N218S-A274T, A088T-A116T-G131H-N218S-K256R, A088T-A116T-G131H-N218S-N243V-K256R, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-N218S-N243V-K256R-L257G, A088T-A116T-G131H-N218S-N243V-L257G, A088T-A116T-G131H-N218S-N243V-S248N-K256R, A088T-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-N218S-S248N-L257G, A088T-A116T-G131H-N243V-K256R, A088T-A116T-G131H-N243V-L257G, A088T-A116T-G131H-N243V-S248N, A088T-A116T-G131H-N243V-S248N-A274V, A088T-A116T-G131H-N243V-S248N-K256R-L257G, A088T-A116T-G131H-S248N-K256R-L257G, A088T-A116T-G131H-S248N-L257G, A088T-A116T-G131H-T158S-K256R-L257G, A088T-A116T-G131H-T158S-K256R-L257G, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S, A088T-A116T-G131H-T158S-N218S-K256R, A088T-A116T-G131H-T158S-N218S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-K256R-L257G, A088T-A116T-G131H-T158S-N218S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-K256R, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N218S-S248N-K256R, A088T-A116T-G131H-T158S-N218S-S248N-L257G, A088T-A116T-G131H-T158S-N243V-K256R-L257G, A088T-A116T-G131H-T158S-N243V-L257G, A088T-A116T-G131H-T158S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-A116T-K256R, A088T-A116T-N218S-K256R, A088T-A116T-N218S-N243V-K256R, A088T-A116T-N218S-N243V-K256R-L257G, A088T-A116T-N218S-N243V-S248N-K256R, A088T-A116T-N218S-N243V-S248N-L257G, A088T-A116T-N243V-K256R, A088T-A116T-N243V-L257G, A088T-A116T-N243V-S248N-K256R, A088T-A116T-N243V-S248N-K256R-L257G, A088T-A116T-S248N, A088T-A116T-S248N-K256R, A088T-A116T-S248N-L257G, A088T-A116T-T158S-N218S, A088T-A116T-T158S-N218S-K256R, A088T-A116T-T158S-N218S-K256R, A088T-A116T-T158S-N218S-K256R-L257G, A088T-A116T-T158S-N218S-N243V-K256R, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N, A088T-A116T-T158S-N218S-N243V-S248N-K256R, A088T-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-N218S-N243V-S248N-L257G, A088T-A116T-T158S-N243V-K256R-L257G, A088T-G131H, A088T-G131H-N218S-K256R-L257G, A088T-G131H-N218S-N243V-K256R, A088T-G131H-N218S-N243V-K256R, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-N243V-L257G, A088T-G131H-N218S-S248N, A088T-G131H-N218S-S248N-K256R, A088T-G131H-N218S-S248N-K256R-L257G, A088T-G131H-N218S-S248N-L257G, A088T-G131H-N218T-L257G, A088T-G131H-N243V-L257G, A088T-G131H-N243V-S248N-K256R, A088T-G131H-S248N, A088T-G131H-S248N-L257G, A088T-G131H-T158S-N218S-K256R, A088T-G131H-T158S-N218S-K256R-L257G, A088T-G131H-T158S-N218S-N243V-K256R, A088T-G131H-T158S-N218S-N243V-K256R, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-G131H-T158S-N218S-S248N, A088T-G131H-T158S-N218S-S248N-K256R, A088T-G131H-T158S-N218S-S248N-K256R, A088T-G131H-T158S-N218S-S248N-L257G-I268V, A088T-G131H-T158S-N243V, A088T-G131H-T158S-N243V-S248N, A088T-G131H-T158S-N243V-S248N, A088T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-A116T, A088T-N109G-A116T-G131H-D140G-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-K256R, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S, A088T-N109G-A116T-G131H-N218S-K256R, A088T-N109G-A116T-G131H-N218S-L257G, A088T-N109G-A116T-G131H-N218S-N243V, A088T-N109G-A116T-G131H-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-N218S-N243V-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N218S-S248N-L257G, A088T-N109G-A116T-G131H-N218S-S248N-L257G, A088T-N109G-A116T-G131H-N243V-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N, A088T-N109G-A116T-G131H-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N-L257G, A088T-N109G-A116T-G131H-S248N-K256R, A088T-N109G-A116T-G131H-S248N-K256R-L257G, A088T-N109G-A116T-G131H-S248N-L257G, A088T-N109G-A116T-G131H-T158S, A088T-N109G-A116T-G131H-T158S-N218S, A088T-N109G-A116T-G131H-T158S-N218S-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-S248N, A088T-N109G-A116T-G131H-T158S-N218S-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-G131H-T158S-N218T-N243V, A088T-N109G-A116T-G131H-T158S-N243V-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-T158S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N-L257G, A088T-N109G-A116T-K256R, A088T-N109G-A116T-N218S, A088T-N109G-A116T-N218S-N243V, A088T-N109G-A116T-N218S-N243V-K256R, A088T-N109G-A116T-N218S-N243V-K256R-L257G, A088T-N109G-A116T-N218S-N243V-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N243V-S248N, A088T-N109G-A116T-N243V-S248N-K256R, A088T-N109G-A116T-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N243V-S248N-K256R-L257G, A088T-N109G-A116T-N243V-S248N-L257G, A088T-N109G-A116T-S248N-K256R, A088T-N109G-A116T-S248N-K256R-L257G, A088T-N109G-A116T-T158S-K256R, A088T-N109G-A116T-T158S-N218S, A088T-N109G-A116T-T158S-N218S-K256R-L257G, A088T-N109G-A116T-T158S-N218S-L257G, A088T-N109G-A116T-T158S-N218S-N243V, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-A116T-T158S-N218S-N243V-S248N, A088T-N109G-A116T-T158S-N218S-N243V-S248N-K256R, A088T-N109G-A116T-T158S-N218S-N243V-S248N-L257G, A088T-N109G-A116T-T158S-N218S-S248N, A088T-N109G-A116T-T158S-N218S-S248N-K256R, A088T-N109G-A116T-T158S-N218S-S248N-L257G, A088T-N109G-A116T-T158S-N243V-K256R, A088T-N109G-A116T-T158S-N243V-S248N, A088T-N109G-A116T-T158S-S248N, A088T-N109G-A116T-T158S-S248N-K256R, A088T-N109G-A116T-T158S-S248N-L257G, A088T-N109G-G131H-N218S-K256R, A088T-N109G-G131H-N218S-K256R, A088T-N109G-G131H-N218S-N243V-K256R, A088T-N109G-G131H-N218S-N243V-K256R, A088T-N109G-G131H-N218S-N243V-K256R-L257G, A088T-N109G-G131H-N218S-N243V-S248N-K256R, A088T-N109G-G131H-N218S-S248N, A088T-N109G-G131H-N218S-S248N-L257G, A088T-N109G-G131H-N243V-K256R-L257G, A088T-N109G-G131H-N243V-K256R-L257G, A088T-N109G-G131H-N243V-L257G, A088T-N109G-G131H-N243V-L257G, A088T-N109G-G131H-N243V-S248N-K256R, A088T-N109G-G131H-N243V-S248N-K256R-L257G, A088T-N109G-G131H-N243V-S248N-L257G, A088T-N109G-G131H-S248N-L257G, A088T-N109G-G131H-T158S-K256R-L257G, A088T-N109G-G131H-T158S-N218S-L257G, A088T-N109G-G131H-T158S-N218S-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-G131H-T158S-N218S-S248N-L257G, A088T-N109G-G131H-T158S-N243V, A088T-N109G-G131H-T158S-N243V-S248N, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N243V-S248N-K256R, A088T-N109G-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-S248N-K256R-L257G,
A088T-N109G-G131H-T158S-W241R-S248N-K256R, A088T-N109G-G131H-V148A-N218S-N243V-K256R-L257G, A088T-N109G-G154A-N155P-E156T-G157L-T158M-S159E-G160E-S161L, A088T-N109G-N218S-K256R, A088T-N109G-N218S-N243V-L257G, A088T-N109G-N218S-N243V-S248N, A088T-N109G-N243V-K256R, A088T-N109G-N243V-S248N, A088T-N109G-N243V-S248N-K256R-L257G, A088T-N109G-S248N, A088T-N109G-T158S-N218S, A088T-N109G-T158S-N218S-K256R, A088T-N109G-T158S-N218S-K256R-L257G, A088T-N109G-T158S-N218S-L257G, A088T-N109G-T158S-N218S-N243V-S248N-K256R, A088T-N109G-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-T158S-N218S-S248N-K256R, A088T-N109G-T158S-N218S-S248N-K256R-L257G, A088T-N109G-T158S-N243V-K256R, A088T-N109G-T158S-N243V-K256R, A088T-N109G-T158S-N243V-S248N-L257G, A088T-N109G-T158S-S248N-K256R-L257G, A088T-N109G-T158S-S248N-L257G, A088T-N109G-T158S-S248N-L257G, A088T-N109G-V147A-N218S-N243V-K256R, A088T-N218S-L257G-I268V, A088T-N218S-N243V, A088T-N218S-N243V-S248N, A088T-N218S-N243V-S248N-N269S, A088T-N218S-S248N-K256R, A088T-N218S-S248N-L257G-Q271R, A088T-N243V, A088T-N243V, A088T-N243V-K256R, A088T-N243V-S248N-K256R, A088T-N243V-S248N-K256R-L257G, A088T-S248N, A088T-T158S-N218S-K256R-L257G, A088T-T158S-N218S-N243V-K256R, A088T-T158S-N218S-N243V-L257G, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-T158S-N243V-K256R-L257G, A088T-T158S-N243V-K256R-L257G-Q271H, A088T-T158S-N243V-S248N, A088T-T158S-N243V-S248N-L257G, A088T-T158S-S248N, A088T-V147A-K256R, A116T-G131H-K256R, A116T-G131H-N218S, A116T-G131H-N218S-K256R-L257G, A116T-G131H-N218S-L257G, A116T-G131H-N218S-N243V, A116T-G131H-N218S-S248N-K256R, A116T-G131H-N218S-S248N-K256R-L257G, A116T-G131H-N243V-S248N, A116T-G131H-N243V-S248N-L257G, A116T-G131H-S248N-K256R, A116T-G131H-T158S-A231V-N243V-L257G, A116T-G131H-T158S-N218S-K256R, A116T-G131H-T158S-N218S-K256R-L257G, A116T-G131H-T158S-N218S-N243V-K256R-L257G, A116T-G131H-T158S-N218S-N243V-S248N-K256R, A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A116T-G131H-T158S-N218S-S248N, A116T-G131H-T158S-N243V-L257G, A116T-G131H-T158S-N243V-S248N-K256R, A116T-G131H-T158S-S248N-K256R, A116T-G131H-T158S-S248N-L257G, A116T-G131H-V143F-T158S-N218S, A116T-K256R, A116T-N218S, A116T-N218S-K256R, A116T-N218S-N243V-L257G, A116T-N218S-N243V-S248N-K256R, A116T-N218S-N243V-S248N-K256R-L257G, A116T-N218S-N243V-S248N-L257G, A116T-N218S-S248N-L257G, A116T-N243V, A116T-N243V-K256R-L257G, A116T-N243V-S248N, A116T-N243V-S248N-K256R-L257G, A116T-S248N-K256R-L257G, A116T-T158S-K256R-L257G, A116T-T158S-N218S, A116T-T158S-N218S-K256R, A116T-T158S-N218S-N243V, A116T-T158S-N218S-N243V-S248N, A116T-T158S-N218S-S248N-K256R, A116T-T158S-N218S-S248N-K256R-L257G, A116T-T158S-N243V-K256R, A116T-T158S-N243V-L257G, A116T-T158S-N243V-S248N, A116T-T158S-S248N-K256R-L257G, G131H-K141R-T158S-N218S-K256R, G131H-N218S, G131H-N218S-K256R, G131H-N218S-N243V-K256R-L257G, G131H-N218S-N243V-S248N, G131H-N218S-N243V-S248N-L257G, G131H-N243V-S248N-K256R, G131H-N243V-S248N-K256R-L257G, G131H-S248N, G131H-T158S-I234T-N243V-K256R, G131H-T158S-N218S-K256R-L257G, G131H-T158S-N218S-N243V, G131H-T158S-N218S-N243V-S248N-L257G, G131H-T158S-N218S-S248N-K256R-L257G, G131H-T158S-N218S-S248N-L257G, G131H-T158S-N243V-K256R, G131H-T158S-N243V-S248N-L257G, N109G, N109G-A116T-G131H-A144V-T158S-S248N-K256R-L257G, N109G-A116T-G131H-K256R-L257G, N109G-A116T-G131H-N218S-N243V-K256R, N109G-A116T-G131H-N218S-N243V-K256R-L257G, N109G-A116T-G131H-N218S-S248N-K256R, N109G-A116T-G131H-N218S-S248N-L257G, N109G-A116T-G131H-N243V-K256R, N109G-A116T-G131H-N243V-S248N, N109G-A116T-G131H-N243V-S248N-K256R-L257G, N109G-A116T-G131H-S248N, N109G-A116T-G131H-S248N-K256R, N109G-A116T-G131H-T158S-K256R, N109G-A116T-G131H-T158S-K256R-L257G, N109G-A116T-G131H-T158S-N218S-K256R, N109G-A116T-G131H-T158S-N218S-K256R-L257G, N109G-A116T-G131H-T158S-N218S-L257G, N109G-A116T-G131H-T158S-N218S-N243V-K256R, N109G-A116T-G131H-T158S-N218S-N243V-S248N-L257G, N109G-A116T-G131H-T158S-N218S-S248N, N109G-A116T-G131H-T158S-N243V-L257G, N109G-A116T-G131H-T158S-N243V-S248N, N109G-A116T-G131H-T158S-S248N, N109G-A116T-G131H-T158S-S248N-K256R, N109G-A116T-G131H-T158S-S248N-K256R-L257G, N109G-A116T-G131H-V149A-T158S-N218S-N243V-S248N-L257G, N109G-A116T-N218S, N109G-A116T-N218S-K256R, N109G-A116T-N218S-N243V-K256R-L257G, N109G-A116T-N218S-N243V-S248N-I268V, N109G-A116T-N243V-K256R-L257G, N109G-A116T-N243V-S248N, N109G-A116T-N243V-S248N-L257G, N109G-A116T-T158S, N109G-A116T-T158S-N218S-N243V-K256R-L257G, N109G-A116T-T158S-N218S-N243V-S248N-K256R-L257G, N109G-A116T-T158S-N218S-N243V-S248N-L257G, N109G-A116T-T158S-N218S-S248N-K256R-L257G, N109G-A116T-T158S-N243V-K256R-L257G, N109G-A116T-T158S-N243V-L257G, N109G-A116T-T158S-N243V-S248N-L257G, N109G-A116T-T158S-Q275R, N109G-A116T-T158S-S248N-K256R-L257G, N109G-G131H-L257G, N109G-G131H-N218S-N243V-S248N-K256R-L257G, N109G-G131H-N218S-N243V-S248N-L257G, N109G-G131H-N218S-S248N-K256R-L257G, N109G-G131H-N243V-K256R, N109G-G131H-S145F-N218S-N243V-K256R-L257G, N109G-G131H-S248N-K256R-L257G, N109G-G131H-S248N-L257G, N109G-G131H-T158S-K256R, N109G-G131H-T158S-N218S-L257G, N109G-G131H-T158S-N218S-N243V, N109G-G131H-T158S-N218S-N243V-K256R, N109G-G131H-T158S-N218S-N243V-K256R-L257G, N109G-G131H-T158S-N218S-N243V-S248N-L257G, N109G-G131H-T158S-N218S-S248N-K256R, N109G-G131H-T158S-N218S-S248N-K256R-L257G-A274T, N109G-G131H-T158S-N218S-S248N-L257G, N109G-G131H-T158S-N243V-S248N, N109G-G131H-T158S-N243V-S248N-K256R-L257G, N109G-G131H-T158S-S248N-K256R-L257G, N109G-G131H-T158S-S248N-L257G, N109G-N218S-K256R-L257G, N109G-N218S-L257G, N109G-N218S-N243V-K256R, N109G-N218S-N243V-S248N-S260F, N109G-N218S-S248N, N109G-N243V-K256R, N109G-N243V-L257G, N109G-N243V-S248N, N109G-N243V-S248N-K256R-L257G, N109G-S182F-S204F-S207L-N218S-S236F-S248N-L257G, N109G-S248N-K256R, N109G-T158S-K256R, N109G-T158S-N218S-N243V-K256R-L257G, N109G-T158S-N218S-S248N-L257G, N109G-T158S-N243V, N109G-T158S-N243V-K256R, N109G-T158S-N243V-S248N, N109G-T158S-N243V-S248N-K256R, N109G-T158S-S248N-K256R, N109G-T158S-S248N-L257G, N218S, N218S-N243V-S248N-K256R, N218S-S248N-L257G, N243V-K256R, N243V-S248N-K256R, N243V-S248N-K256R-L257G, N243V-S248N-L257G-Q271R, S003P-A116T-T158S-S248N-K256R, S248N, T158S-N218S, T158S-N218S-A272V, T158S-N218S-L257G, T158S-N218S-N243V-K256R-L257G, T158S-N218S-N243V-L257G, T158S-N218S-S248N-K256R-L257G, T158S-N243V-K256R, T158S-N243V-K256R-L257G, T158S-N243V-S248N-K256R, V004A-N109G-A116T-G131H-S248N-K256R-L257G, and Y006H-A116T-G131H-S248N, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0572] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A015S-A088T-N109G-G131H-T158S-N218S-S248N, A088T, A088T-A116T-G131H-N218S-N243V-S248N, A088T-A116T-G131H-N218S-S248N, A088T-A116T-G131H-N218S-S248N, A088T-A116T-G131H-N218S-S248N-K256R-L257G, A088T-A116T-G131H-N243V, A088T-A116T-G131H-N243V-K256R, A088T-A116T-G131H-N243V-S248N, A088T-A116T-G131H-S248N-K256R, A088T-A116T-G131H-T158S-N218S-N243V-S248N, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-A116T-G131H-T158S-N218S-S248N, A088T-A116T-G131H-T158S-N218S-S248N-K256R, A088T-A116T-G131H-T158S-N243V, A088T-A116T-G131H-T158S-N243V-K256R, A088T-A116T-G131H-T158S-N243V-S248N, A088T-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-A116T-G131H-T158S-S248N, A088T-A116T-G131H-T158S-S248N-K256R, A088T-A116T-G131H-T158S-S248N-L257G, A088T-A116T-G131H-V147A-T158S-N218S-N243V-S248N-L257G, A088T-A116T-N218S-I268V, A088T-A116T-N218S-L257G, A088T-A116T-N218S-N243V-N269D, A088T-A116T-N218S-S248N, A088T-A116T-N218S-S248N, A088T-A116T-N218S-S248N-L257G, A088T-A116T-N243V-K256R-L257G, A088T-A116T-N243V-S248N-K256R, A088T-A116T-S248N-K256R, A088T-A116T-T158S-N218S, A088T-A116T-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-N218S-S248N, A088T-A116T-T158S-N218S-S248N-K256R-L257G, A088T-A116T-T158S-N243V-S248N, A088T-A116T-T158S-N243V-S248N-K256R, A088T-A116T-T158S-N243V-S248N-K256R-L257G, A088T-A116T-T158S-N243V-S248N-L257G, A088T-A116T-T158S-S248N, A088T-A116T-T158S-S248N-K256R-L257G, A088T-A116T-T158S-S248N-L257G, A088T-A116T-V143A-N218S-S248N-K256R, A088T-G131H-A138V-N218S-L257G, A088T-G131H-K141E-N218S-N243V-S248N-L257G, A088T-G131H-K256R, A088T-G131H-N218S-K256R, A088T-G131H-N218S-N243V-K256R-L257G, A088T-G131H-N218S-N243V-S248N, A088T-G131H-N218S-N243V-S248N, A088T-G131H-N218S-N243V-S248N-K256R, A088T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-G131H-N218S-S248N, A088T-G131H-N218S-S248N-K256R, A088T-G131H-N243V-K256R-L257G, A088T-G131H-N243V-S248N, A088T-G131H-N243V-S248N-K256R-L257G, A088T-G131H-S248N, A088T-G131H-S248N-K256R, A088T-G131H-T158S-K256R, A088T-G131H-T158S-L257G, A088T-G131H-T158S-N218S, A088T-G131H-T158S-N218S-N243V-S248N, A088T-G131H-T158S-N218S-N243V-S248N-K256R, A088T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-N218S-N243V-S248N-L257G, A088T-G131H-T158S-N218S-S248N, A088T-G131H-T158S-N218S-S248N-K256R-L257G, A088T-G131H-T158S-N243V, A088T-G131H-T158S-N243V-K256R, A088T-G131H-T158S-N243V-S248N-K256R, A088T-G131H-T158S-N243V-S248N-L257G, A088T-G131H-T158S-S248N-K256R-L257G, A088T-G131H-T158S-S248N-L257G, A088T-I107T-N109G-G131H-N218S-A223G-S248N-K256R, A088T-L257G, A088T-L257G, A088T-N109G-A116T-G131H-A232S-N243V-K256R, A088T-N109G-A116T-G131H-K256R-L257G, A088T-N109G-A116T-G131H-L257G, A088T-N109G-A116T-G131H-N218S-K256R, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R, A088T-N109G-A116T-G131H-N218S-N243V-S248N-N269D, A088T-N109G-A116T-G131H-N243V-K256R, A088T-N109G-A116T-G131H-S248N, A088T-N109G-A116T-G131H-T158S-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243F-S248N, A088T-N109G-A116T-G131H-T158S-N218S-N243V, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218T-K256R, A088T-N109G-A116T-G131H-T158S-N243V, A088T-N109G-A116T-G131H-T158S-N243V-K256R, A088T-N109G-A116T-G131H-T158S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R-I268V, A088T-N109G-A116T-G131H-T158S-N243V-S248N-L257G, A088T-N109G-A116T-G131H-T158S-S248N, A088T-N109G-A116T-G131H-T158S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-V149A-N218S-S248N-K256R-L257G, A088T-N109G-A116T-K256R, A088T-N109G-A116T-N218S-K256R, A088T-N109G-A116T-N218S-N243V-S248N-K256R, A088T-N109G-A116T-N218S-N243V-S248N-L257G, A088T-N109G-A116T-N218S-S248N, A088T-N109G-A116T-N218S-S248N-K256R, A088T-N109G-A116T-S248N, A088T-N109G-A116T-T158S-L257G, A088T-N109G-A116T-T158S-N218S, A088T-N109G-A116T-T158S-N243V, A088T-N109G-A116T-T158S-N243V-K256R, A088T-N109G-A116T-T158S-N243V-S248N-K256R, A088T-N109G-G131H-N218S-N243V-S248N, A088T-N109G-G131H-N218S-S248N-K256R, A088T-N109G-G131H-N218S-S248N-K256R-L257G, A088T-N109G-G131H-N218S-S248N-K256R-L257G-Q275R, A088T-N109G-G131H-N218S-S248N-K256R-Q271R, A088T-N109G-G131H-N243V, A088T-N109G-G131H-N243V-K256R, A088T-N109G-G131H-N243V-S248N-K256R, A088T-N109G-G131H-S248N-K256R, A088T-N109G-G131H-S248N-L257G, A088T-N109G-G131H-T158S, A088T-N109G-G131H-T158S, A088T-N109G-G131H-T158S-K256R, A088T-N109G-G131H-T158S-K256R, A088T-N109G-G131H-T158S-N218S-N243V-K256R, A088T-N109G-G131H-T158S-N218S-N243V-K256R, A088T-N109G-G131H-T158S-N218S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-L257G, A088T-N109G-G131H-T158S-N243V-K256R, A088T-N109G-G131H-T158S-N243V-K256R-L257G, A088T-N109G-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-S248N, A088T-N109G-G131H-T158S-S248N-L257G, A088T-N109G-G131H-V149A-K256R-L257G, A088T-N109G-G131H-V149L-T158S-K256R-L257G, A088T-N109G-K256R, A088T-N109G-K256R-L257G, A088T-N109G-K256R-L257G, A088T-N109G-L257G, A088T-N109G-N218S-K256R, A088T-N109G-N218S-N243V-K256R, A088T-N109G-N218S-N243V-S248N-K256R, A088T-N109G-N218S-N243V-S248N-K256R, A088T-N109G-N218S-S248N, A088T-N109G-N218S-S248N-L257G, A088T-N109G-N243V-K256R, A088T-N109G-N243V-S248N-L257G-I268V, A088T-N109G-S248N, A088T-N109G-S248N-K256R, A088T-N109G-T158S-K256R-L257G, A088T-N109G-T158S-N218S, A088T-N109G-T158S-N218S-K256R-Q271H, A088T-N109G-T158S-N218S-N243V-K256R, A088T-N109G-T158S-N218S-N243V-K256R-L257G, A088T-N109G-T158S-N218S-N243V-L257G, A088T-N109G-T158S-N218S-N243V-S248N-K256R, A088T-N109G-T158S-N218S-S248N-N269D, A088T-N109G-T158S-N243V-K256R-L257G, A088T-N109G-T158S-N243V-S248N-A274D, A088T-N109G-T158S-N243V-S248N-K256R-L257G-N269D, A088T-N218S-K256R, A088T-N218S-N243V, A088T-N218S-N243V-K256R, A088T-N218S-N243V-K256R-L257G, A088T-N218S-N243V-S248N-K256R, A088T-N218S-N243V-S248N-L257G, A088T-N218S-S248N-L257G, A088T-N243V-S248N, A088T-N243V-S248N-K256R, A088T-N243V-S248N-L257G, A088T-S145T-T158S-S248N, A088T-S248N-K256R-L257G, A088T-S248N-L257G, A088T-T158S, A088T-T158S, A088T-T158S-K256R, A088T-T158S-L257G, A088T-T158S-N218S, A088T-T158S-N218S-K256R, A088T-T158S-N218S-N243V-K256R, A088T-T158S-N218S-N243V-K256R-L257G, A088T-T158S-N218S-N243V-L257G, A088T-T158S-N218S-S248N, A088T-T158S-N218S-S248N, A088T-T158S-N243V, A088T-T158S-N243V-K256R, A088T-T158S-N243V-S248N-L257G, A116T, A116T-G131H-L257G, A116T-G131H-N218S-K256R, A116T-G131H-N218S-N243V-K256R, A116T-G131H-N218S-N243V-K256R-L257G, A116T-G131H-N218S-N243V-S248N, A116T-G131H-N218S-N243V-S248N-K256R, A116T-G131H-N218S-S248N, A116T-G131H-T158S-K256R, A116T-G131H-T158S-N218S-L257G, A116T-G131H-T158S-N218S-N243V-K256R, A116T-G131H-T158S-N218S-N243V-L257G, A116T-G131H-T158S-N218S-N243V-S248N, A116T-G131H-T158S-N218S-S248N-L257G-N269D, A116T-G131H-T158S-N218S-S248N-Q271R, A116T-G131H-T158S-N218T-L257G, A116T-G131H-T158S-N243V-K256R, A116T-G131H-V150A-T158S-N243V-S248N-K256R-L257G, A116T-K141E-N218S-N243V-S248N-K256R-L257G, A116T-L257G, A116T-N218S-N243V-S248N, A116T-N218T-N243V-S248N, A116T-N243V-S248N-L257G, A116T-S248N, A116T-S248N-L257G, A116T-T158S-K256R, A116T-T158S-N218S-L257G, A116T-T158S-N218S-N243V-K256R, A116T-T158S-N218S-N243V-L257G, A116T-T158S-N243V, A116T-T158S-N243V-K256R-L257G, A116T-T158S-N243V-S248N-K256R, A116T-T158S-N243V-S248N-K256R-L257G, A116T-V149I-T158S-N243V-S248N-K256R-Q271H, G024S-G053S-N078S-G097A-N101S, G024S-G053S-N078S-G097A-N101S-A128S, G131H-K256R, G131H-N218S-K256R-L257G, G131H-N218S-N243V-L257G, G131H-N218S-N243V-S248N-K256R, G131H-N218S-S248N-K256R, G131H-N218S-S248N-L257G, G131H-N243V, G131H-N243V-S248N, G131H-N243V-S248N-L257G, G131H-T158S-K256R-L257G, G131H-T158S-N218S, G131H-T158S-N218S-K256R, G131H-T158S-N218S-N240H-N243V-S248N-K256R-L257G, G131H-T158S-N218S-N243V-K256R, G131H-T158S-N218S-N243V-K256R-L257G, G131H-T158S-N218S-N243V-S248N, G131H-T158S-N218S-N243V-S248N-K256R-L257G, G131H-T158S-N218S-S248N-I268V, G131H-T158S-N218S-S248N-K256R-L257G-N269S, G131H-T158S-N243V-K256R-L257G, G131H-T158S-N243V-S248N, G131H-T158S-N243V-S248N-K256R, G131H-T158S-S248N, G131H-T158S-S248N-L257G, G131H-W241L-N243V-S248N-K256R, I107T-G131H-T158S-N243V-S248N-K256R-L257G, I107T-N109G-G131H-N218S-L257G, N109G-A116T-G131H-L257G, N109G-A116T-G131H-N218S-L257G, N109G-A116T-G131H-N218S-N243V, N109G-A116T-G131H-N218S-N243V-S248N-K256R, N109G-A116T-G131H-N218S-W241R-N243V-K256R, N109G-A116T-G131H-N243V-S248N-K256R, N109G-A116T-G131H-S248N-I268V, N109G-A116T-G131H-T158S-N218S, N109G-A116T-G131H-T158S-N218S-S248N-K256R, N109G-A116T-G131H-T158S-N218S-S248N-K256R-L257G, N109G-A116T-G131H-T158S-N218S-S248N-L257G, N109G-A116T-G131H-T158S-N243V-K256R-L257G, N109G-A116T-G131H-T158S-S248N-L257G, N109G-A116T-K141E-T158S-N218S-N243V-L257G, N109G-A116T-K256R, N109G-A116T-N218S-N243V, N109G-A116T-N218S-N243V-S248N, N109G-A116T-N218S-S248N-L257G, N109G-A116T-N243V-S248N-K256R-L257G, N109G-A116T-S248N, N109G-A116T-T158S-N218S-N243V-S248N-K256R, N109G-A116T-T158S-N218S-W241R-N243V, N109G-A116T-T158S-N243V-S248N, N109G-A116T-T158S-S248N-L257G, N109G-G131H-K256R, N109G-G131H-N218S-K256R, N109G-G131H-N218S-N243V, N109G-G131H-N218S-N243V-K256R-L257G, N109G-G131H-N218S-N243V-S248N-K256R, N109G-G131H-N218S-S248N-K256R, N109G-G131H-N243V, N109G-G131H-N243V-S248N, N109G-G131H-N243V-S248N-K256R-L257G, N109G-G131H-S248N, N109G-G131H-T158S-L257G, N109G-G131H-T158S-N218S-N243V-S248N, N109G-G131H-T158S-N218S-N243V-S248N-K256R, N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, N109G-G131H-T158S-N243V-L257G, N109G-G131H-T158S-N243V-S248N-K256R, N109G-G131H-T158S-S248N-K256R, N109G-K141E-N218S-S248N-L257G, N109G-N218S-N243V, N109G-N218S-N243V-S248N-K256R, N109G-N218S-S248N-K256R-L257G, N109G-N218S-S248N-L257G, N109G-N243V-S248N-L257G-Q275R, N109G-T158S-I268V, N109G-T158S-N218S, N109G-T158S-N218S-N243V, N109G-T158S-N218S-N243V-S248N, N109G-T158S-N218S-S248N-K256R, N109S-A116T-S248N, N218S-K256R, N218S-N243V-K256R, N218S-N243V-L257G, N218S-N243V-S248N, N218S-S248N, N218S-S248N-K256R, N218 S-S248N-K256R-L257G, S003P-N109G-G131H-N218S-N243V-S248N-K256R-L257G, S248N-K256R-L257G, T158S-K256R, T158S-K256R-L257G, T158S-N218S-K256R-L257G, T158S-N218S-L233S-S248N, T158S-N243V, T158S-N243V-S248N-K256R-N269D, T158S-N243V-S248N-L257G, V004L-A116T-N218S-N243V-S248N-L257G, and Y006H-N218S-N243V-S248N, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and a PI value of 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0573] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A088T-A098S-G131H-S248N-K256R-L257G, A088T-A116T-G131H-K256R, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-N243V-S248N-L257G, A088T-A116T-G131H-S248N, A088T-A116T-G131H-S248N-K256R-L257G, A088T-A116T-G131H-T158S-K256R, A088T-A116T-G131H-T158S-L257G, A088T-A116T-G131H-T158S-N218S-L257G, A088T-A116T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-A116T-K256R-L257G, A088T-A116T-K256R-L257G, A088T-A116T-N218S-N243V-Q271R, A088T-A116T-N218S-S248N-K256R, A088T-A116T-T158S, A088T-A116T-T158S, A088T-A116T-T158S-K256R, A088T-A116T-T158S-N218S-N243V-L257G, A088T-A116T-T158S-N218S-S248N-K256R, A088T-G131H, A088T-G131H-L257G, A088T-G131H-N218S-N243V-S248N-L257G, A088T-G131H-N243V-K256R, A088T-G131H-N243V-L257G, A088T-G131H-N243V-S248N, A088T-G131H-N243V-S248N-K256R, A088T-G131H-T158S-N218S-I234T-S248N-L257G, A088T-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-G131H-T158S-N243V-K256R-L257G, A088T-G131H-T158S-S248N, A088T-G131H-T158S-S248N-K256R, A088T-G131H-T158S-S248N-K256R-L257G, A088T-G131H-V149L-T158S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-K141E-N218S, A088T-N109G-A116T-G131H-N218S-N243V-S248N-Q275R, A088T-N109G-A116T-G131H-N218S-S248N, A088T-N109G-A116T-G131H-N243V-S248N-K256R, A088T-N109G-A116T-G131H-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-K256R, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-G131H-T158S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-S248N, A088T-N109G-A116T-G131H-V149A-T158S-N218S-K256R, A088T-N109G-A116T-N218S-S248N-K256R-L257G, A088T-N109G-A116T-N243V-K256R, A088T-N109G-A116T-N243V-S248N-L257G, A088T-N109G-A116T-S248N-L257G, A088T-N109G-A116T-T158S-N212D-N243V-K256R-L257G, A088T-N109G-A137E-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-D140G-N243V, A088T-N109G-G131H-A152S-T158S-N218S-S248N-K256R, A088T-N109G-G131H-D140G-T158S-N243V-S248N-K256R, A088T-N109G-G131H-K256R-L257G, A088T-N109G-G131H-N218S, A088T-N109G-G131H-N218S-N243V, A088T-N109G-G131H-T158S-L257G, A088T-N109G-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A088T-N109G-G131H-T158S-N218S-W241R-S248N-L257G, A088T-N109G-G131H-T158S-S248N-K256R, A088T-N109G-N218S-S248N-K256R, A088T-N109G-N243V-S248N-K256R, A088T-N109G-T158S, A088T-N109G-T158S-N243V-S248N-K256R, A088T-N109G-T158S-N243V-S248N-Q275R, A088T-N109G-T158S-S248N, A088T-N218S-N243V-S248N-K256R-L257G, A088T-N243V-L257G, A088T-S248N, A088T-T158S-K256R, A088T-T158S-N218S-K256R, A088T-T158S-N218S-L257G, A088T-T158S-N218S-L257G, A088T-T158S-N218S-N243V-S248N, A088T-T158S-N218S-N243V-S248N-L257G, A088T-T158S-N218S-Q245K-S248N-K256R, A088T-T158S-N218S-S248N-L257G-Q275K, A088T-T158S-N243V-K256R, A088T-T158S-N243V-K256R-L257G, A088T-T158S-N243V-S248N-K256R, A088T-T158S-S248N, A088T-T158S-S248N-K256R-L257G, A088T-T158S-S248N-L257G, A088T-T158S-S248N-L257G, A098S-G131H-T158S-N218S-N243V-S248N-K256R-L257G, A116T-G131H-K141E-N218S-N243V-S248N-L257G, A116T-G131H-N218S-W241R-N243V-S248N-K256R-L257G, A116T-G131H-N243V, A116T-G131H-T158S-N218S-S248N-K256R, A116T-G131H-V139I-N218S-N243V-S248N, A116T-N218S-S248N-K256R, A116T-T158S-L257G-Q271R, A116T-T158S-N218S-N243V-K256R-L257G, G053S-A088T-N109G-A116T-G131H-T158S-G169S-N218S-S248N-K256R-L257G, G131H-N218S-L257G, G131H-T158S, G131H-T158S-K256R, K256R, L090I-N109G-T158S-N243V, L257G, N109G-A116T-G131H, N109G-A116T-G131H-N243V, N109G-A116T-G131H-T158S-N218S-N243V-S248N-K256R, N109G-A116T-S248N-K256R, N109G-A116T-T158S-N218S-K237R-N243V-S248N, N109G-A116T-T158S-S248N, N109G-G131H-T158S, N109G-G131H-T158S-N243V-K256R-L257G, N109G-G131H-T158S-S248N-Q271R, N109G-N218S-S248N-K256R, N109G-N243V-S248N-L257G, N109G-S248N, N109G-T158S-N218S-N243V-L257G, N109G-T158S-N243V-K256R-L257G, N109G-T158S-N243V-S248N-K256R-L257G, N218S-N243V-S248N-K256R-L257G, S003P-N109G-A116T-G131H-T158S-N218S-K256R, S003P-N109G-G131H-T158S-L257G, S105H-W106G-I107L-I108S-N109A-G110A-I111S-E112N-W113G-A114P, T158S-N218S-S248N-K256R, T158S-N243V-S248N, T158S-S248N, T158S-S248N-K256R-L257G, V004A-A088T-A116T-T158S-N218S, V004A-A088T-G131H-N218S-N243V-S248N-L257G, Y006H-N109G-N218S-N243V-S248N, Y104H-A116T-T158S-S248N, A088T-A116T-G131H-N218S-S248N-L257G, A088T-A116T-G131H-T158S-N218S-N243V-L257G, A088T-A116T-G131H-T158S-N243V-K256R-L257G, A088T-A116T-N218S, A088T-G131D-T158S-N243V-S248N, A088T-G131H-N218S-K237R-K256R-L257G, A088T-G131H-N218S-K256R, A088T-K213N-N243V-S248N-K256R, A088T-K256R-L257G, A088T-N109G-A116T-G131H-D140G-S248N-L257G, A088T-N109G-A116T-G131H-L257G, A088T-N109G-A116T-G131H-N218S-N243V-S248N-K256R-L257G, A088T-N109G-A116T-G131H-T158S-N218S-N243V-S248N, A088T-N109G-A116T-M124I-G131H-T158S-N218S-S248N-L257G, A088T-N109G-A116T-T158S-N218S-N243V-K256R, A088T-N109G-A116T-T158S-N218S-N243V-L257G, A088T-N109G-A116T-T158S-N218S-S248N, A088T-N109G-G131H-K256R-L257G, A088T-N109G-G131H-N218S-N243V-S248N-L257G, A088T-N109G-G131H-N243V-S248N, A088T-N109G-G131H-T158S-L233S-N243V-S248N, A088T-N109G-G131H-T158S-N218S-S248N-K256R, A088T-N109G-L257G, A088T-N109G-N218S-K256R-L257G, A088T-N109G-N218S-S248N-T255K-K256R-L257G, A088T-N109G-S248N-K256R-L257G, A088T-N109G-T158S-N218S-N243V, A088T-N109G-T158S-N218S-N243V-L257G, A088T-N109G-T158S-N218S-N243V-S248N-L257G, A088T-N109G-T158S-N218S-S248N, A088T-S248N-K256R-L257G, A088T-S248N-L257G-I268V, A088T-T158S-N243V-L257G, A088T-T158S-V203I-N218S-K256R-L257G, A088T-Y104H-A116T-G131H-N218S-N243V, A116T-D140G-T158S-N218S-N243V-S248N, A116T-G131H-T158S-K256R-L257G, A116T-G131H-T158S-N243V-S248N, A116T-G157E-T158S-N243V-S248N-K256R, A116T-S248N-K256R, G131H-N218S-S248N, G131H-N243V-K256R, G131H-T158S-N243V-L257G, K256R-L257G, N109G-A116T-G131H-T158S-N218S-N243V-S248N, N109G-A116T-N218S-N243V-L257G, N109G-A116T-N218S-W241R-N243V-S248N-K256R-L257G, N109G-A116T-T158S-N243V, N109G-A116T-T158S-S248N-K256R, N109G-N243V, N109G-T158S-K256R-L257G, N243V, P014L-A015L-L016C-H017T-S018L-Q019K-G020A-Y021T-T022L-G023E, S003P-S248N-L257G, T158S-N218S-N243V-S248N, T158S-S248N-K256R, Y104H-N109G-G131H-N243V-S248N, A088T-A116T-G131H-L257G, A088T-A116T-G131H-N218S-N243V, A088T-A116T-G131H-N218S-S248N-K256R, A088T-A116T-G131H-T158S-N243V-S248N-L257G, A088T-A116T-L257G, A088T-A116T-N218S, A088T-A116T-N218S-N243V-S248N, A088T-A116T-T158S-N218S-S248N, A088T-A116T-T158S-N243V-S248N-K256R, A088T-A138E-N218S-N243V-K256R, A088T-G131H-N218S-N243V-S248N-K256R, A088T-G131H-T158S-S248N-K256R, A088T-N109G-A116T-G131H-N218S-S248N-K256R-L257G, A088T-N109G-A116T-G131H-N243V-S248N-K256R, A088T-N109G-A116T-G131H-T158S-N218S-L257G-I268V, A088T-N109G-A116T-G131H-W241L-S248N-K256R-L257G, A088T-N109G-A116T-N218S-N243V-S248N-K256R, A088T-N109G-A116T-T158S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-N243V-S248N, A088T-N109G-G131H-N218S-L257G, A088T-N109G-G131H-N218S-S248N, A088T-N109G-G131H-T158S-N243V-S248N-L257G, A088T-N109G-N243V-K256R-L257G, A088T-N109G-T158S-S248N-K256R, A088T-N109G-W241R-S248N-K256R, A088T-N218S-S248N-L257G, A088T-T158S-N218S-S248N-K256R, A088T-T158S-N218S-S248N-L257G, A116T-G131H-T158S-N218S-N243V, A116T-N218S-S248N, A116T-N243V-K256R, A116T-T158S, G131H-S248N-K256R, N109G-A116T-G131H-N243V-K256R-L257G, N109G-A116T-I234T-N243V-S248N-K256R-L257G, N109G-A116T-T158S-K256R-L257G, N109G-A116T-T158S-N218S-S248N-K256R, N109G-G131H, N109G-G131H-A137V-T158S-N218S-S248N, N109G-G131H-K141E-L257G, N109G-G131H-T158S-N243V-S248N-L257G, T158S-N218S-N243V-K256R, A088T-A116T-T158S-N243V-K256R-L257G, A088T-N109G-A116T-T158S-K256R-L257G, A088T-N109G-T158S-N218S-S248N, A088T-S248N-L257G, A088T-Y104H-N109G-G131H-A137E-T158S-N218S-N243V-S248N-K256R, G065D-A088T-G131H-N243V-S248N, Y104H-N218S-L257G, A088T-A116T-G131H-V150A-N218S-S248N-L257G, A088T-A116T-T158S-K256R-L257G, A088T-I108T-N109G-G131H-T158S-N218S-S248N-K256R-L257G, A088T-N109G-A116T-T158S-S248N-L257G-Q271P, A088T-N109G-A116T-V148A-N218S-N243V, A088T-Y104H-N109G-A116T-A153S-N218S-N243V-S248N-L257G-N269D, and V004M-A116T-V148A-T158S-N243V-S248N-K256R, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0574] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN'-v36 and/or BPN-'v3 and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v36 in a BMI microswatch or egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:6, wherein the variant comprises at least one substitution selected from the group of X001R/T, X002R, X003F/P, X004A/L/M/P, X005S, X006H, X014L, X015L/S, X016C, X017T, X018L, X019K, X020A, X021T, X022L, X023E, X024S, X034S, X053S, X057Q, X065D, X078S, X086L, X088T, X090I, X097A, X098S, X101S, X104H, X105H/P, X106G, X107L/T, X108S/T, X109A/G/S, X110A, X111S, X112N, X113G, X114P/S, X116T, X124I, X128S, X131D/H, X137EN, X138EN, X139I, X140G, X141E/R, X143A/F, X144V, X145F/P/T, X146C, X147A/I, X148A, X149A/I/L, X150A, X151S, X152S, X153S, X154A, X155P, X156T, X157E/L, X158M/S, X159E, X160E, X161L, X169S, X182F, X203I, X204F, X207L, X211V, X212D, X213N, X216S, X218S/T, X223G, X231V, X232S, X233S, X234T, X235P, X236F/P, X237N/R, X240H, X241L/R, X243F/V, X245K, X248N, X251K, X255K, X256R, X257G, X260F, X267M, X268V, X269D/S, X271H/K/P/R, X272G/V, X273T, X274D/L/T/V, and X275K/R/S, and optionally at least one substitution selected from the group of A001R/T, Q002R, S003F/P, V004A/L/M/P, P005S, Y006H, P014L, A015L/S, L016C, H017T, S018L, Q019K, G020A, Y021T, T022L, G023E, G024S, G034S, G053S, P057Q, G065D, N078S, P086L, A088T, L090I, G097A, A098S, N101S, Y104H, S105H/P, W106G, I107L/T, I108S/T, N109A/G/S, G110A, I111S, E112N, W113G, A114P/S, A116T, M124I, A128S, G131D/H, A137E/V, A138E/V, V139I, D140G, K141E/R, V143A/F, A144V, S145F/P/T, G146C, V147A/I, V148A, V149A/I/L, V150A, A151S, A152S, A153S, G154A, N155P, E156T, G157E/L, T158M/S, S159E, G160E, S161L, G169S, S182F, V203I, S204F, S207L, G211V, N212D, K213N, A216S, N218S/T, A223G, A231V, A232S, L233S, I234T, L235P, S236F/P, K237N/R, N240H, W241L/R, N243F/V, Q245K, S248N, E251K, T255K, K256R, L257G, S260F, L267M, I268V, N269D/S, Q271H/K/P/R, A272G/V, A273T, A274D/L/T/V, Q275K/R/S, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) BPN'-v3, and BPN'-v36 and a PI value greater than that of BPN', BPN'-v3, and BPN'-v36 in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 10
Construction and Cleaning Performance of Additional Variants of BPN'-v36
[0575] The DNA from the site evaluation libraries of the BPN'-v36 (described in Example 7) was further mutagenized by error-prone PCR. These libraries were amplified with primers P4973 and P4950 (described in Example 7) using Taq DNA polymerase (Promega). Each PCR amplification reaction contained 30 pmol of each primer, 100 ng of the template DNA (SELs of the BPN'-v36) and various amount of MnCl.sub.2. The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 2 min. The DNA fragment was gel-purified by the QIAGEN.RTM. gel-band purification kit, digested by the BamHI and HindIII restriction enzymes and ligated with the pHPLT-BPN' partial opt vector that also was digested with the same restriction enzymes. Ligation mixtures were amplified using rolling circle amplification in an Illustra Templiphi kit according to the manufacturer's recommendation (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. For this purpose, 1 .mu.l of the ligation mixture was mixed with 5 .mu.l of the sample buffer, heated to 95.degree. C. for 3 min and cooled on ice. Next, 5 .mu.l of the reaction buffer and 0.2 .mu.l of the enzyme were added to each tube, followed by incubation at 30.degree. C. for 10 hours. Products of the rolling circle amplification were diluted 100 times and used to transform B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C.
[0576] About 500,000 clones were pre-screened on skim milk plates. Very few of them formed halos (i.e., indicative of the presence of functional protease). Colonies with halos were picked, inoculated in 150 .mu.l of LB media containing 10 .mu.g/mL neomycin and sequenced (Quintara). Sequences of these clones were analyzed by looking for combination of mutations, which occurred in this pool multiple times and might provide performance benefits. In order to assess the performance of these mutation combinations, double mutants were created in the BPN'-v36 background by PCR fusion as described below. For this purpose, two or three partially overlapping fragments were amplified by mutagenic primers. Primer combinations used to generate the respective variants are shown in Table 10-1 and primer sequences are shown in Table 10-2.
TABLE-US-00034 TABLE 10-1 Primers Pairs Used to Amplify Fragments Variant Mutation 1 Mutation 2 Fragment 1 Fragment 2 Fragment 3 1 A45S S236Y P4974, P6645 P6644, P6647 P6646, P4976 2 A45S S236G P4974, P6645 P6644, P6649 P6648, P4976 3 I115T S183T P4974, P6651 P6650, P6655 P6654, P4976 4 I115V N184Y P4974, P6653 P6652, P6657 P6656, P4976 5 I31T S37P P4974, P6659 P6658, P4976 6 I31T I35L P4974, P6661 P6660, P4976 7 I31V S38W P4974, P6663 P6662, P4976 8 N25K P129R P4974, P6665 P6664, P6667 P6666, P4976 9 N25K P129K P4974, P6665 P6664, P6669 P6668, P4976 10 P14T S37T P4974, P6671 P6670, P6673 P6672, P4976 11 P5L Q217K P4974, P6679 P6678, P6681 P6680, P4976 12 P5L Q217G P4974, P6679 P6678, P6683 P6682, P4976 13 Q10L S37P P4974, P6685 P6684, P6675 P6674, P4976 14 Q10R S37T P4974, P6687 P6686, P6673 P6672, P4976 15 S37P T254S P4974, P6675 P6674, P6689 P6688, P4976 16 N25K S37P P4974, P6665 P6664, P6675 P6674, P4976 17 G24A S37W P4974, P6691 P6690, P6677 P6676, P4976 18 N25K P129R P4974, P6665 P6664, P6667 P6666, P4976 20 S161P S162L P4974, P6695 P6694, P4976 21 S161P T253A P4974, P6693 P6692, P6701 P6700, P4976 22 S161P S260P P4974, P6693 P6692, P6703 P6702, P4976 23 S162L D181H P4974, P6697 P6696, P6711 P6710, P4976 24 S162L D181G P4974, P6697 P6696, P6713 P6712, P4976 25 S18F S162L P4974, P6715 P6714, P6697 P6696, P4976 26 S18T S162P P4974, P6717 P6716, P6699 P6698, P4976 27 S18P D120N P4974, P6719 P6718, P6727 P6726, P4976 28 S18Y K213R P4974, P6721 P6720, P6729 P6728, P4976 29 S18L Y21S P4974, P6731 P6730, P4976 30 S18T Y21N P4974, P6733 P6732, P4976 31 S9T K141F P4974, P6635 P6734, P6737 P6736, P4976 32 S9T K141R P4974, P6635 P6734, P6739 P6738, P4976 33 Q19L S260N P4974, P6725 P6724, P6705 P6704, P4976 34 Q19L S260P P4974, P6725 P6724, P6703 P6702, P4976 35 N61S S260P P4974, P6741 P6740, P6703 P6702, P4976 36 N61D S260I P4974, P6743 P6742, P6707 P6706, P4976 37 T253A S260P P4974, P6701 P6700, P6703 P6702, P4976 38 A134T S260G P4974, P6745 P6744, P6709 P6708, P4976 39 A133V S260N P4974, P6648 P6746, P6705 P6704, P4976
TABLE-US-00035 TABLE 10-2 Primer Sequences Used for Generation of Double Mutants of BPN'-v36 Primer Name Primer Sequence SEQ ID NO: P6644 CAGATCTTAAAGTCTCTGGAGGGGCTTCTATGGTGC SEQ ID NO: 64 P6645 CATAGAAGCCCCTCCAGAGACTTTAAGATCTGGATGGCTC SEQ ID NO: 65 P6646 GCATTGATTCTTTACAAGCACCCGAACTGGACAAAC SEQ ID NO: 66 P6647 CAGTTCGGGTGCTTGTAAAGAATCAATGCGGCCGCCCCA SEQ ID NO: 67 P6648 GCATTGATTCTTGGTAAGCACCCGAACTGGACAAAC SEQ ID NO: 68 P6649 CCAGTTCGGGTGCTTACCAAGAATCAATGCGGCCGCCCCA SEQ ID NO: 69 P6650 CATCGAATGGGCCACAGCGAATAACATGGATGTAATCAAC SEQ ID NO: 70 P6651 CATCCATGTTATTCGCTGTGGCCCATTCGATGCCGTTGAT SEQ ID NO: 71 P6652 CATCGAATGGGCCGTAGCGAATAACATGGATGTAATCAAC SEQ ID NO: 72 P6653 CATCCATGTTATTCGCTACGGCCCATTCGATGCCGTTGAT SEQ ID NO: 73 P6654 CTGTAGACTCTACAAATCAACGTGCCTCTTTTTCCT SEQ ID NO: 74 P6655 AAAGAGGCACGTTGATTTGTAGAGTCTACAGCGCCCACTG SEQ ID NO: 75 P6656 CTGTAGACTCTTCATACCAACGTGCCTCTTTTTCCTCC SEQ ID NO: 76 P6657 GAAAAAGAGGCACGTTGGTATGAAGAGTCTACAGCGCCCA SEQ ID NO: 77 P6658 TAGCGGTTACAGACAGCGGTATCGACCCAAGCCATCCAGATC SEQ ID NO: 78 TTAAAGTCG P6659 ATGGCTTGGGTCGATACCGCTGTCTGTAACCGCTACTTTAACA SEQ ID NO: 79 TTGCCTC P6660 TAAAGTAGCGGTTACAGACAGCGGTTTAGACTCGAGCCATCC SEQ ID NO: 80 AGATCTT P6661 ATGGCTCGAGTCTAAACCGCTGTCTGTAACCGCTACTTTAACA SEQ ID NO: 81 TTGCCTC P6662 GGTTGTAGACAGCGGTATCGACTCGTGGCATCCAGATCTTAA SEQ ID NO: 82 AGTCGCTG P6663 ATGCCACGAGTCGATACCGCTGTCTACAACCGCTACTTTAACA SEQ ID NO: 83 TTGCCTC P6664 CTACACTGGAGGCAAAGTTAAAGTAGCGGTTATCGACA SEQ ID NO: 84 P6665 ATAACCGCTACTTTAACTTTGCCTCCAGTGTAGCCTTGAG SEQ ID NO: 85 P6666 GAGCCTGGGAGCACGTAGCGGCAGTGCGGCACTTAAA SEQ ID NO: 86 P6667 GTGCCGCACTGCCGCTACGTGCTCCCAGGCTCATGTTGAT SEQ ID NO: 87 P6668 TGAGCCTGGGAGCAAAGAGCGGCAGTGCGGCACTTAAA SEQ ID NO: 88 P6669 GTGCCGCACTGCCGCTCTTTGCTCCCAGGCTCATGTTGAT SEQ ID NO: 89 P6670 ATCACAAATTAAAGCCACAGCTCTGCACTCTCAAGGCTAC SEQ ID NO: 90 P6671 AGAGTGCAGAGCTGTGGCTTTAATTTGTGATACGCCGTAA SEQ ID NO: 91 P6672 GACAGCGGTATCGACACAAGCCATCCAGATCTTAAAGTCG SEQ ID NO: 92 P6673 TAAGATCTGGATGGCTTGTGTCGATACCGCTGTCGATAAC SEQ ID NO: 93 P6674 GACAGCGGTATCGACCCAAGCCATCCAGATCTTAAAGTCG SEQ ID NO: 94 P6675 TAAGATCTGGATGGCTTGGGTCGATACCGCTGTCGATAAC SEQ ID NO: 95 P6676 GACAGCGGTATCGACTGGAGCCATCCAGATCTTAAAGTCG SEQ ID NO: 96 P6677 TAAGATCTGGATGGCTCCAGTCGATACCGCTGTCGATAAC SEQ ID NO: 97 P6678 ACGCGCAGTCCGTGTTATACGGCGTATCACAAATTAAAGC SEQ ID NO: 98 P6679 ATTTGTGATACGCCGTATAACACGGACTGCGCGTACGCAT SEQ ID NO: 99 P6680 ACAAGTATGGTGCGAAAAACGGGACTTCCATGGCCTC SEQ ID NO: 100 P6681 CATGGAAGTCCCGTTTTTCGCACCATACTTGTTCCCTG SEQ ID NO: 101 P6682 ACAAGTATGGTGCGGGAAACGGGACTTCCATGGCCTC SEQ ID NO: 102 P6683 CCATGGAAGTCCCGTTTCCCGCACCATACTTGTTCCCTG SEQ ID NO: 103 P6684 CTTACGGCGTATCATTAATTAAAGCCCCTGCTCTGCAC SEQ ID NO: 104 P6685 GAGCAGGGGCTTTAATTAATGATACGCCGTAAGGCACGGA SEQ ID NO: 105 P6686 CTTACGGCGTATCACGTATTAAAGCCCCTGCTCTGCAC SEQ ID NO: 106 P6687 GAGCAGGGGCTTTAATACGTGATACGCCGTAAGGCACGGA SEQ ID NO: 107 P6688 TTTAGAAAACACCTCTACAAAACTTGGTGATTCTTTCTAC SEQ ID NO: 108 P6689 TCACCAAGTTTTGTAGAGGTGTTTTCTAAACTGCTGCGGA SEQ ID NO: 109 P6690 AGGCTACACTGGAGCAAATGTTAAAGTAGCGGTTATCGAC SEQ ID NO: 110 P6691 GCTACTTTAACATTTGCTCCAGTGTAGCCTTGAGAGTG SEQ ID NO: 111 P6692 GAGGGAACATCCGGACCATCGAGTACCGTCGGTTATCCA SEQ ID NO: 112 P6693 ACCGACGGTACTCGATGGTCCGGATGTTCCCTCATTCCCA SEQ ID NO: 113 P6694 AGGGAACATCCGGACCATTAAGTACCGTCGGTTATCCAGG SEQ ID NO: 114 P6695 ACCGACGGTACTTAATGGTCCGGATGTTCCCTCATTCCCA SEQ ID NO: 115 P6696 GAACATCCGGATCATTAAGTACCGTCGGTTATCCAGGCA SEQ ID NO: 116 P6697 ATAACCGACGGTACTTAATGATCCGGATGTTCCCTCATTC SEQ ID NO: 117 P6698 GAACATCCGGATCACCAAGTACCGTCGGTTATCCAGGCA SEQ ID NO: 118 P6699 ATAACCGACGGTACTTGGTGATCCGGATGTTCCCTCATTC SEQ ID NO: 119 P6700 GTTTAGAAAACGCAACTACAAAACTTGGTGATTCTTTC SEQ ID NO: 120 P6701 CACCAAGTTTTGTAGTTGCGTTTTCTAAACTGCTGCGGAC SEQ ID NO: 121 P6702 CAAAACTTGGTGATCCATTCTACTATGGAAAAGGGCTGAT SEQ ID NO: 122 P6703 TTTCCATAGTAGAATGGATCACCAAGTTTTGTAGTGGTGT SEQ ID NO: 123 P6704 CAAAACTTGGTGATAACTTCTACTATGGAAAAGGGCTGAT SEQ ID NO: 124 P6705 TTTCCATAGTAGAAGTTATCACCAAGTTTTGTAGTGGTGT SEQ ID NO: 125 P6706 CAAAACTTGGTGATATCTTCTACTATGGAAAAGGGCTGAT SEQ ID NO: 126 P6707 TTTCCATAGTAGAAGATATCACCAAGTTTTGTAGTGGTGT SEQ ID NO: 127 P6708 CAAAACTTGGTGATGGATTCTACTATGGAAAAGGGCTGAT SEQ ID NO: 128 P6709 TTTCCATAGTAGAATCCATCACCAAGTTTTGTAGTGGTGT SEQ ID NO: 129 P6710 GTGGGCGCTGTACACTCTTCAAATCAACGTGCCTCTT SEQ ID NO: 130 P6711 CACGTTGATTTGAAGAGTGTACAGCGCCCACTGCAATCAC SEQ ID NO: 131 P6712 GTGGGCGCTGTAGGATCTTCAAATCAACGTGCCTCTT SEQ ID NO: 132 P6713 CACGTTGATTTGAAGATCCTACAGCGCCCACTGCAATCAC SEQ ID NO: 133 P6714 CCTGCTCTGCACTTCCAAGGCTACACTGGAGGCAATG SEQ ID NO: 134 P6715 CTCCAGTGTAGCCTTGGAAGTGCAGAGCAGGGGCTTTAAT SEQ ID NO: 135 P6716 CCTGCTCTGCACACACAAGGCTACACTGGAGGCAATG SEQ ID NO: 136 P6717 CTCCAGTGTAGCCTTGTGTGTGCAGAGCAGGGGCTTTAAT SEQ ID NO: 137 P6718 CCTGCTCTGCACCCACAAGGCTACACTGGAGGCAATG SEQ ID NO: 138 P6719 CTCCAGTGTAGCCTTGTGGGTGCAGAGCAGGGGCTTTAAT SEQ ID NO: 139 P6720 CCTGCTCTGCACTACCAAGGCTACACTGGAGGCAATG SEQ ID NO: 140 P6721 CTCCAGTGTAGCCTTGGTAGTGCAGAGCAGGGGCTTTAAT SEQ ID NO: 141 P6722 CCTGCTCTGCACTTACAAGGCTACACTGGAGGCAATG SEQ ID NO: 142 P6723 CTCCAGTGTAGCCTTGTAAGTGCAGAGCAGGGGCTTTAAT SEQ ID NO: 143 P6724 TGCTCTGCACTCTTTAGGCTACACTGGAGGCAATGTTA SEQ ID NO: 144 P6725 TTGCCTCCAGTGTAGCCTAAAGAGTGCAGAGCAGGGGCTT SEQ ID NO: 145 P6726 ATCGCGAATAACATGAACGTAATCAACATGAGCCTGGGA SEQ ID NO: 146 P6727 CTCATGTTGATTACGTTCATGTTATTCGCGATGGCCCAT SEQ ID NO: 147 P6728 CTTCCAGGGAACCGTTATGGTGCGCAAAACGGGACTT SEQ ID NO: 148 P6729 GTTTTGCGCACCATAACGGTTCCCTGGAAGCGTCGATTG SEQ ID NO: 149 P6730 CTGCACTTACAAGGCTCTACTGGAGGCAATGTTAAAGTAG SEQ ID NO: 150 P6731 TAACATTGCCTCCAGTAGAGCCTTGTAAGTGCAGAGCAGGGG SEQ ID NO: 151 CTTTAAT P6732 GCTCTGCACTTACAAGGCAACACTGGAGGCAATGTTAAAGTAG SEQ ID NO: 152 P6733 AACATTGCCTCCAGTGTTGCCTTGTAAGTGCAGAGCAGGGGC SEQ ID NO: 153 TTTAAT P6734 CTTACGGCGTAACACAAATTAAAGCCCCTGCTCTG SEQ ID NO: 154 P6735 AGGGGCTTTAATTTGTGTTACGCCGTAAGGCACGGACT SEQ ID NO: 155 P6736 TTAAAGCAGCAGTTGATTTCGCTGTTGCATCTGGTGTCGT SEQ ID NO: 156 P6737 AGATGCAACAGCGAAATCAACTGCTGCTTTAAGTGCCGCA SEQ ID NO: 157 P6738 TTAAAGCAGCAGTTGATCGTGCTGTTGCATCTGGTGTCGT SEQ ID NO: 158 P6739 AGATGCAACAGCACGATCAACTGCTGCTTTAAGTGCCGCA SEQ ID NO: 159 P6740 AAACCCGTTTCAAGATTCTAATTCTCATGGCACACACGTC SEQ ID NO: 160 P6741 TGTGCCATGAGAATTAGAATCTTGAAACGGGTTTGTTTCG SEQ ID NO: 161 P6742 AAACCCGTTTCAAGATGATAATTCTCATGGCACACACGTC SEQ ID NO: 162 P6743 TGTGCCATGAGAATTATCATCTTGAAACGGGTTTGTTTCG SEQ ID NO: 163 P6744 AAGCGGCAGTGCGACACTTAAAGCAGCAGTTGATAAAGC SEQ ID NO: 164 P6745 TCAACTGCTGCTTTAAGTGTCGCACTGCCGCTTGGTGCTC SEQ ID NO: 165 P6746 CAAGCGGCAGTGTTGCACTTAAAGCAGCAGTTGATAA SEQ ID NO: 166 P6747 ACTGCTGCTTTAAGTGCAACACTGCCGCTTGGTGCTCCCA SEQ ID NO: 167
[0577] Each PCR amplification reaction contained 30 pmol of each primer and 100 ng of the BPN'-v36 parent template DNA (plasmid pHPLT-BPN'-v36) (see FIG. 4). Amplifications were carried out using Vent DNA polymerase (NEB). The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 40 sec. Following amplification, the 5' and 3' gene fragments were gel-purified by the QIAGEN.RTM. gel-band purification kit, mixed (50 ng of each fragment), mixed and amplified by PCR once again using the primers P4973 and P4950 to generate the full-length gene fragment. The PCR conditions were same as described above, except the extension phase, which was carried out at 72.degree. C. for 2 min. The full-length DNA fragment was gel-purified by the QIAGEN.RTM. gel-band purification kit, digested by the BamHI and HindIII restriction enzymes and ligated with the pHPLT-BPN' partial opt vector that also was digested with the same restriction enzymes. Ligation mixtures were amplified using rolling circle amplification in an Illustra Templiphi kit according to the manufacturer's recommendation (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. For this purpose, 1 .mu.l of the ligation mixture was mixed with 5 .mu.l of the sample buffer, heated to 95.degree. C. for 3 min and cooled on ice. Next, 5 .mu.l of the reaction buffer and 0.2 .mu.l of the enzyme were added to each tube, followed by incubation at 30.degree. C. for 10 hours. Products of the rolling circle amplification were diluted 100 times and used to transform B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C. Subsequently, the colonies with halos were inoculated in 150 .mu.l of LB media containing 10 .mu.g/mL neomycin. The next day, the cultures were either frozen with 15% glycerol or grown in MBD medium for biochemical analysis as described in Example 2.
[0578] The variants were tested for cleaning performance using BMI microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8, BMI microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 7, and Egg microswatch assay in Detergent Composition 4 at 16.degree. C. and pH 8. Protein content was determined using TCA assay. All assays were performed as described in Example 1 and Performance Indices were calculated relative to BPN'-v36 (i.e., BPN'-S24G-S53G-S78N-S101N-G128A-Y217Q) (with a PI value of 1.0).
[0579] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A133V-S260N, N061S-S260P, P014T-S037T, S009T-K141F, S009T-K141R, S018F-S162L, S018L-Y021S, S018P-D120N, S018T-S162P, S018T-Y021N, S018Y-K213R, S161P-S162L, S161P-S260P, and T253A-S260P, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and a PI value of about 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0580] The following BPN'-v36 variants were determined to have a PI value of about 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A134T-S260G, I115V-N184Y, N025K-S037P, Q010L-S037P, Q019L-S260N, Q019L-S260P, S037P-T254S, and S161P-T253A, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value of about 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0581] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A045S-S236G, G024A-S037W, I031V-S038W, N061D-S260I, Q010R-S037T, I115T-S183T, N025K-P129K, N025K-P129R, A045S-S236Y, and S162L-D181H, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0582] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of S018F-S162L, S018P-D120N, P014T-S037T, S009T-K141R, and S161P-S162L, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0583] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of N061S-S260P, Q010L-S037P, S009T-K141F, S018L-Y021S, S018T-S162P, S018T-Y021N, S018Y-K213R, S037P-T254S, S161P-S260P, S161P-T253A, and T253A-S260P, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and a PI value of about 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0584] The following BPN' variants were determined to have a PI value of about 0.9 relative to BPN'-v3 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN' amino acid sequence (SEQ ID NO:2) comprising at least one set of amino acid substitutions selected from the group consisting of A133V-S260N, A134T-S260G, 15T-S183T, 15V-N184Y, N061D-S260I, Q019L-S260N, and Q019L-S260P, wherein positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, a PI value of about 0.9 relative to BPN'-v3, and/or an enhanced proteolytic activity compared to BPN' in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0585] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 7 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A045S-S236G, G024A-S037W, Q010R-S037T, A045S-S236Y, I031V-S038W, N025K-S037P, S162L-D181H, and N025K-P129R, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0586] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of I031V-S038W, P014T-S037T, S018F-S162L, S018P-D120N, and S162L-D181H, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0587] The following BPN'-v36 variants were determined to have a PI value of about 1.0 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A133V-S260N, A134T-S260G, G024A-S037W, I115V-N184Y, N025K-P129K, N025K-P129R, N061D-S260I, Q019L-S260P, S009T-K141F, S009T-K141R, S018L-Y021S, S018T-S162P, S018T-Y021N, S018Y-K213R, S161P-S162L, S161P-T253A, and T253A-S260P, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of about 1.0 relative to BPN'-v3, and/or a PI value of about 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0588] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.8 and equal to or less than 0.9 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of A045S-S236G, A045S-S236Y, I115T-S183T, N025K-S037P, N061S-S260P, Q010L-S037P, Q010R-S037T, Q019L-S260N, S037P-T254S, and S161P-S260P, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.8 and equal to or less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
[0589] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN'-v36 and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v36 in a BMI or egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:6, wherein the variant comprises at least one substitution comprising at least one substitution selected from the group of X009T, X010L/R, X014T, X018F/L/P/T/Y, X019L, X021N/S, X024A, X025K, X031V, X037P/T/W, X038W, X045S, X061D/S, X115TN, X120N, X129K/R, X133V, X134T, X141F/R, X161P, X162L/P, X181H, X183T, X184Y, X213R, X236G/Y, X253A, X254S, and X260G/I/N/P, and optionally at least one substitution selected from the group of S009T, Q010L/R, P014T, S018F/L/P/T/Y, Q019L, Y021N/S, G024A, N025K, I031V, S037P/T/W, S038W, A045S, N061D/S, 15TN, D120N, P129K/R, A133V, A134T, K141F/R, S161P, S162L/P, D181H, S183T, N184Y, K213R, S236G/Y, T253A, T254S, and S260G/I/N/P, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) BPN'-v3, and BPN'-v36 and a PI value greater than that of BPN', BPN'-v3, and BPN'-v36 in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such variant as described in greater detail elsewhere herein.
Example 11
1. Generation of Combinatorial Libraries FS1-FS3
[0590] The pHPLT-BPN'-v3 plasmid containing the BPN' expression cassette served as template DNA (parent plasmid) for cloning. Three separate combinatorial libraries (FS1, FS2, and FS3) were synthesized by DNA2.0, and were delivered as individual ligation reactions. A list of libraries and possible substitutions are shown in Table 11-1. The libraries were designed to allow the incorporation of either the wild type residues or the substitutions at each site described in Table 11-1.
[0591] For efficient transformation of B. subtilis, the DNA from the ligation reaction was amplified by rolling circle amplification (RCA) using the Illustra Templiphi kit (GE Healthcare). Reactions were performed according to the manufacturer's protocol. One microliter often-fold diluted amplified DNA was used to transform 50 .mu.L of competent B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). The transformation mixture was shaken at 37.degree. C. for 1 hour. Ten microliter aliquots of the transformation mixture were plated on skim milk (1.6%) Luria Agar plates supplemented with 10 .mu.g/ml of neomycin (Teknova).
[0592] Transformants were picked into microtiter plates containing 125-150 .mu.l Luria broth medium supplemented with 10 .mu.g/ml neomycin. Plates were grown overnight at 37.degree. C. with 250-300 rpm shaking and 70-80% humidity using Enzyscreen lids for microtiter plates (Enzyscreen). Between 7 and 10 microliters from the overnight culture plate were used to inoculate a new microtiter plate containing 190 .mu.l of MBD medium (a MOPS based defined medium) with 10 .mu.g/ml neomycin. MBD medium was prepared essentially as known in the art (see Neidhardt et al., J. Bacteriol. 119:736-747 (1974)), except that NH.sub.4Cl.sub.2, FeSO.sub.4, and CaCl.sub.2 were omitted from the base medium, 3 mM K.sub.2HPO.sub.4 was used, and the base medium was supplemented with 60 mM urea, and 100 ml of a solution made of 210 g/L glucose, and 350 g/L maltodextrin. The micronutrients were made up as a 100.times. stock solution containing in one liter, 400 mg FeSO.sub.4.7H.sub.2O, 100 mg MnSO.sub.4.H.sub.2O, 100 mg ZnSO.sub.4.7H.sub.2O, 50 mg CuCl.sub.2.2H.sub.2O, 100 mg CoCl.sub.2.6H.sub.2O, 100 mg NaMoO.sub.4.2H.sub.2O, 100 mg Na.sub.2B.sub.4O.sub.7.10H.sub.2O, 10 ml of 1 M CaCl.sub.2, and 10 ml of 0.5 M sodium citrate. The MBD medium containing microtiter plates were grown for 60-70 hours at 37.degree. C., 250-300 rpm, and 70-80% humidity using Enzyscreen lids (Enzyscreen) for determining protein expression. The next day, cultures were filtered through a micro-filter plate (0.22 .mu.m; Millipore) and the resulting filtrate was used for biochemical analysis.
TABLE-US-00036 TABLE 11-1 Possible Substitutions for Combinatorial Libraries FS1-FS3 FS1 FS2 FS3 1 G102A A97G N61E 2 S130P A128G P129E 3 T55P N123G K213L 4 V203Y G102A S145D 5 N61P L126V Q275E 6 S101N G100N P40E 7 S53G N62Q S159K 8 S78N M124I S24R 9 S87T-A88L-S89G N61P A144K 10 S24G-N25G S130P N240K 11 L75S-N76Y P129S P239R
2. Generation of Variants to Improve BPN' Stability
[0593] To improve BPN' stability, variants were constructed using either parent molecules pHPLT-BPN' G97A-G128A-Y217Q-S87D or pHPLT-BPN' G97A-G128A-Y217Q-P40E, both synthesized by Gene Oracle, or parent molecules pHPLT-BPN' G97A-G128A-Y217Q-S78N and pHPLT-partial opt FNA (B. amyloliquefaciens subtilisin BPN'-Y217L) synthesized by GeneArt.
[0594] The information listed in Tables 11-2 and 11-3 summarizes the parent molecule used, the mutations added, and the primers used to construct variants provided herein.
TABLE-US-00037 TABLE 11-2 Primer Sequences Used for the Generation of BPN' Stability Mutants Primer Primer Sequence 5' to 3' SEQ ID NO: p31 /5PHOS/CGGCGTTAAACAATAACATTGGCGTGCTTGGTGTAG 169 p32 /5PHOS/ACCAAGCACGCCAATGTTATTGTTTAACGCCGCAACC 170 p25 /5PHOS/GTATCGACTCGAGCCATGAAGATCTTAAAGTCGCTGG 171 AG p26 /5PHOS/CAGCGACTTTAAGATCTTCATGGCTCGAGTCGATACCG 172 p33 /5PHOS/TTGGTGTAGCCCCGGATGCTTCGCTCTACGCCGTTAA 173 AG p34 /5PHOS/CGTAGAGCGAAGCATCCGGGGCTACACCAAGCACG 174 p29 /5PHOS/GAACGGTTGCGGCGTTAGATAATTCTATTGGCGTGCT 175 TG p30 /5PHOS/AGCACGCCAATAGAATTATCTAACGCCGCAACCGTTC 176 p27 /5PHOS/AACGGTTGCGGCGTTAGATAATAACATTGGCGTGCTT 177 GGTGTAG p28 /5PHOS/ACACCAAGCACGCCAATGTTATTATCTAACGCCGCAA 178 CCGTTCCTG p7 /5PHOS/CGGCGTTAAACAATAATATTGGCGTGCTTGG 179 p8 /5PHOS/CCAAGCACGCCAATATTATTGTTTAACGCCG 180 p21 /5PHOS/GGTGTAGCCCCGGATGCTTCGCTCTACG 181 p22 /5PHOS/CGTAGAGCGAAGCATCCGGGGCTACACC 182 p11 /5PHOS/GTATCGACTCGAGCCATGAAGATCTTAAAG 183 p12 /5PHOS/CTTTAAGATCTTCATGGCTCGAGTCGATAC 184 p13 /5PHOS/CGCTTCCAGGGAACAACTATGGTGCGTA 185 p14 /5PHOS/TACGCACCATAGTTGTTCCCTGGAAGCG 186 p15 /5PHOS/CACTCTCAAGGCTACGTTGGATCAAATGTTA 187 p16 /5PHOS/TAACATTTGATCCAACGTAGCCTTGAGAGTG 188 p17 /5PHOS/TGGCGTTTCTATTGAATCGACGCTTCCAG 189 p18 /5PHOS/CTGGAAGCGTCGATTCAATAGAAACGCCA 190
TABLE-US-00038 TABLE 11-3 Templates Used, Mutations to be Incorporated, and Primers Used in the Generation of Variants for Improved Stability Mutation to Primers Parent Molecules (Templates) Incorporate Used 1 pHPLT-BPN' G97A-G128A-Y217Q-S87D S78N p31, p32 2 pHPLT-BPN' G97A-G128A-Y217Q-S78N P40E p25, p26 3 pHPLT-BPN' G97A-G128A-Y217Q-P40E S87D p33, p34 4 pHPLT-BPN' G97A-G128A-Y217Q-S78N P40E, S87D p25, p33 5 pHPLT-BPN' G97A-G128A-Y217Q-S87D N76D p29, p30 6 pHPLT-BPN' G97A-G128A-Y217Q-S87D N76D, S78N p27, p28 7 pHPLT-partial opt FNA S78N p7, p8 8 pHPLT-partial opt FNA S87D p21, p22 9 pHPLT-partial opt FNA P40E p11, p12 10 pHPLT-partial opt FNA K213N p13, p14 11 pHPLT-partial opt FNA T22V p15, p16 12 pHPLT-partial opt FNA Q206E p17, p18 13 pHPLT-partial opt FNA P40E, S87D, p11, p21, S78N p7
[0595] Bacillus subtilis strains expressing plasmids were streaked onto 1.6% skim milk plates containing 10 ppm neomycin and grown overnight at 37.degree. C. Single colonies from the plates were grown overnight at 37.degree. C. with shaking at 250 rpm in 5 mL Luria broth containing 10 ppm neomycin. Plasmids were isolated using the QIAGEN.RTM. Miniprep kit protocol adding 1 microliter of Ready Lyse lysozyme (Epicentre) for 15 minutes at 37.degree. C. in buffer P1 to aid in cell lysis. The plasmids were sequenced to ensure correct DNA sequences before proceeding. The plasmids were methylated using NEB's Dam Methylase Kit in a reaction containing 77.5 .mu.L water+10 .mu.L Buffer 10.times.+0.25 .mu.L SAM+2 .mu.L DAM methylase+10 .mu.L miniprep DNA (-150 ng/.mu.L) at 37.degree. C. overnight. The methylated plasmid DNA was purified using a DNA Clean and Concentrator Kit (Zymo) or with a QIAGEN.RTM. PCR purification kit. Multi-Site QUIKCHANGE.RTM. mutagenesis reactions were set up for each of the DNA templates in a reaction mix containing 2.5 .mu.L Buffer 5.times.+0.75 .mu.L Quik Solution+0.5 .mu.L primer 1 (25 .mu.M)+0.5 .mu.L primer 2 (25 .mu.M)+1.5 .mu.L dNTP's+1 .mu.L enzyme blend+16.25 .mu.L H.sub.2O+2 .mu.L DNA. The PCR program used was: 95.degree. C. for 1 min; (95.degree. C. for 1 min, 53.degree. C. for 1 min, 65.degree. C. for 10 min).times.29 cycles; 65.degree. C. for 10 min, 4.degree. C. hold. In all reactions, PCR was performed using a MJ Research PTC-200 Peltier thermal cycler. The parental DNA from the PCR samples was removed by addition of 1 .mu.L of DpnI to QUIKCHANGE.RTM. mutagenesis kit reactions at 37.degree. C. overnight. To increase the transformation frequency, the DpnI digested reactions were amplified using rolling circle amplification (RCA) using the Illustra TempliPhi kit according to the manufacturer's protocol. B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo) were transformed with 1 .mu.L each of the RCA reaction and the transformed cells were plated onto LA+1.6% skim milk plates containing 10 ppm neomycin and incubated at 37.degree. C. overnight. Colonies from overnight growth were selected to perform colony PCR for sequencing using "puReTaq Ready-To-Go PCR Beads" (Amersham). The PCR and sequencing primers used were pHPLT F1 (SEQ ID NO:54) and pHPLT seq R1 (SEQ ID NO:55). Clones with appropriate sequences were frozen. The BPN' variant proteins were produced by growing the B. subtilis transformants in 96 well microtiter plates at 37.degree. C. for 68 hours in a MOPS based medium containing urea.
3. Generation of BPN' Variants Derived from Five Different Parent Plasmids
[0596] BPN' variants were constructed using a total of five different templates: BPN'-v3 (G97A-G128A-Y217Q), BPN'-v4 (G97A-N123G-Y217Q), BPN' variant 8, (S87D-G97A-N109D-G128A-S188D-S248R-Y217Q), BPN' variant 16 (S87D-G97A-N109D-G128A-S188D-Y217Q), and BPN' variant 21 (S87R-G97A-N109R-G128A-S188R-Y217Q-S248R) as shown in Table 11-4. The generation of BPN'-v4 and BPN'-v3 are described in PCT App. No. PCT/US09/46066 (WO 09/149144), filed on Jun. 3, 2009, hereby incorporated herein by reference for such description. BPN' variants 8, 16, 21 were synthesized by Gene Oracle and served as parent plasmids to build additional variants. All variants were generated using QUIKCHANGE.RTM. mutagenesis kits, except two (variants 5 and 33), which were generated using fusion PCR as described below. Primers (listed in Table 11-5) for the generation of variants were synthesized at Integrated DNA Technologies. The mutations introduced (shown in bold) and the primers and template used are shown in Table 11-4.
TABLE-US-00039 TABLE 11-4 Mutations Introduced (bold) & Parent Plasmids Used to Generate BPN' Variants Variant Variants Constructed Parent Plasmid Primers Used 1 G97A-N123C-Y217Q G97A-N123G-Y217Q N123C f, N123C r 2 N76D-G97A-G128A-Y217Q G97A-G128A-Y217Q N76D f, N76D r 3 G97A-N109D-G128A-Y217Q G97A-G128A-Y217Q N109D f1, N109D r 4 G97A-G128A-S188D-Y217Q G97A-G128A-Y217Q S188D f, S188D r 5 G97A-G128A-S248D-Y217Q G97A-G128A-Y217Q pHPLT F1, S248D forfus, pHPLT R1, S248D revfus 6 G97A-G128A-S188D-S248R- G97A-G128A-Y217Q S188D f, S248R f1 Y217Q 7 G97A-N109D-G128A-S188D- S87D-G97A-N109D- D87S f, D87S r S248R-Y217Q G128A-S188D-S248R- Y217Q 8 S87D-G97A-N109D-G128A- Synthesized by Gene Synthesized by Gene S188D-S248R-Y217Q Oracle Oracle 9 S87R-G97A-G128A-S188D- S87D-G97A-N109D- S87R f, S248D f1, S248D-Y217Q G128A-S188D-Y217Q D109N f 10 S87R-G97A-N109D-G128A- S87D-G97A-N109D- R248Sfor, R248Srev S188D-Y217Q G128A-S188D-S248R- Y217Q 11 G97A-N109D-G128A-S188R- G97A-G128A-Y217Q N109D f2, S188R f Y217Q 12 S87R-G97A-N109D-G128A- S87D-G97A-N109D- S87R f, S87R r S188D-Y217Q-S248R G128A-S188D-S248R- Y217Q 13 G97A-N109D-G128A- G97A-G128A-Y217Q N109D f2, S248R f2 Y217Q-S248R 14 S87D-G97A-G128A-Y217Q- G97A-G128A-Y217Q S87D f, S248R f1 S248R 15 S87D-G97A-N109D-G128A- S87D-G97A-N109D- S248D f1, S248D r S188D-Y217Q-S248D G128A-S188D-Y217Q 16 S87D-G97A-N109D-G128A- Synthesized by Gene Synthesized by Gene S188D-Y217Q Oracle Oracle 17 G97A-N109D-G128A- G97A-G128A-Y217Q N109D f2, S248D f2 Y217Q-S248D 18 S87R-G97A-G128A-Y217Q G97A-G128A-Y217Q S87R f, S87R r 19 S87R-G97A-G128A-Y217Q- G97A-G128A-Y217Q S87R f, S248R f1 S248R 20 S87R-G97A-G128A-S188R- S87R-G97A-N109R- D109N f, D109N r Y217Q-S248R G128A-S188R-Y217Q- S248R 21 S87R-G97A-N109R-G128A- Synthesized by Gene Synthesized by Gene S188R-Y217Q-S248R Oracle Oracle 22 G97A-G102A-G128A-Y217Q G97A-G128A-Y217Q G102A f, G102A r 23 G97A-G128A-S130P-Y217Q G97A-G128A-Y217Q S130P f, S130P r 24 G97A-S101N-G128A-Y217Q G97A-G128A-Y217Q S101N f, S101N r 25 G97A-G100N-G128A-Y217Q G97A-G128A-Y217Q G100N f, G100N r 26 N61P-G97A-G128A-Y217Q G97A-G128A-Y217Q N61P f, N61P r 27 G97A-G128A-A187D-Y217Q G97A-G128A-Y217Q A187D f, A187D r 28 G97A-G128A-F189D-Y217Q G97A-G128A-Y217Q f189D f, f189D r 29 G97A-G128A-A137V-Y217Q G97A-G128A-Y217Q A137V f, A137V r 30 S63T-G97A-G128A-Y217Q G97A-G128A-Y217Q S63T f, S63T r 31 G97A-Q103N-G128A-Y217Q G97A-G128A-Y217Q Q103N f, Q103N r 32 N62D-G97A-G128A-Y217Q G97A-G128A-Y217Q N62D f, N62D r 33 G97A-G100E-G128A-Y217Q G97A-G128A-Y217Q G100E Fsfor, pHPLT F1, G100E Fsrev, pHPLT R1
Generation of BPN' Variants via QUIKCHANGE.RTM. Mutagenesis
[0597] Bacillus subtilis strains containing plasmids expressing BPN'-v3, BPN'-v4, BPN' variant 8, BPN' variant 16, and BPN' variant 21 were streaked onto 1.6% skim milk plates containing 10 ppm neomycin and grown overnight at 37.degree. C. Single colonies from the plates were grown overnight at 37.degree. C. with shaking at 250 rpm in 5 mL Luria broth containing 10 ppm neomycin. Plasmids expressing BPN'-v3, BPN'-v4, BPN' variant 8, BPN' variant 16, and BPN' variant 21 were isolated using QIAGEN.RTM. Miniprep kit protocol except following cell lysis, 1 microliter of Ready Lyse lysozyme was added and incubated for 15 minutes at 37.degree. C. The plasmids were methylated using NEB's Dam Methylase Kit in a reaction containing 77.75 .mu.L H.sub.2O+10 .mu.L Buffer 10.times.+0.25 .mu.L SAM+2 .mu.L DAM methylase+104 miniprep DNA at 37.degree. C. overnight. The methylated DNA was purified using the QIAGEN.RTM. PCR purification kit. Variants 14, 18, and 19 listed in Table 11-4 were generated using QUIKCHANGE LIGHTNING.TM. Multi Site-Directed Mutagenesis kits (Stratagene) in a reaction mix containing 2.5 .mu.L Buffer 5.times.+0.75 .mu.L Quik Solution+0.5 .mu.L primer 1 (25 .mu.M)+0.5 .mu.L primer 2 (25 .mu.M)+1.5 .mu.L, dNTP's+1 .mu.L enzyme blend+16.25 .mu.L H.sub.2O+2 .mu.L DNA. The PCR program used was as follows: 95.degree. C. for 2 min; (95.degree. C. for 20 sec, 55.degree. C. for 30 sec, 64.degree. C. for 2 min 30 sec).times.29 cycles; 64.degree. C. for 5 min, 4.degree. C. hold.
[0598] The remaining variants were created using QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis kits in a reaction mix containing 2.5 .mu.L Buffer 5.times.+0.75 .mu.L Quik Solution+0.5 .mu.L primer 1 (25 .mu.M)+0.5 .mu.L primer 2 (25 .mu.M)+1.5 .mu.L dNTP's+1 .mu.L enzyme blend+16.25 .mu.L H.sub.2O+2 .mu.L DNA. The PCR program used was as follows: 95.degree. C. for 1 min; (95.degree. C. for 1 min, 53.degree. C. for 1 min, 65.degree. C. for 10 min).times.29 cycles; 65.degree. C. for 10 min, 4.degree. C. hold. In all reactions, PCR was performed using a MJ Research PTC-200 Peltier thermal cycler. The primers used for the Quik-Change reactions are provided in Table 11-5.
TABLE-US-00040 TABLE 11-5 Primers Used for Quik-Change Reactions Primer SEQ ID Name Primer Sequence (5' to 3') NO: N76D f /5PHOS/GAACGGTTGCGGCGTTAGATAATTCTATTGGCGTGCTTG 191 N76D r /5PHOS/AGCACGCCAATAGAATTATCTAACGCCGCAACCGTTC 192 S87D f /5PHOS/TTGGTGTAGCCCCGGATGCTTCGCTCTACGCCGTTAAAG 193 S87D r /5PHOS/CGTAGAGCGAAGCATCCGGGGCTACACCAAGCACG 194 G102A f /5PHOS/GCAGCAGACGGATCAGCACAATACTCATGGATTAT 195 G102A r /5PHOS/ATAATCCATGAGTATTGTGCTGATCCGTCTGCTGC 196 S130P f /5PHOS/CAACATGAGCCTGGGAGCACCACCGGGCAGTGCGGCACT 197 TAAAGC S130P r /5PHOS/GCTTTAAGTGCCGCACTGCCCGGTGGTGCTCCCAGGCTCA 198 TGTTG S101N f /5PHOS/GTTCTTGCAGCAGACGGAAATGGCCAATACTCATGGATT 199 S101N r /5PHOS/AATCCATGAGTATTGGCCATTTCCGTCTGCTGCAAGAAC 200 G100N f /5PHOS/TTAAAGTTCTTGCAGCAGACAATTCAGGCCAATACTCATG 201 GA G100N r /5PHOS/TCCATGAGTATTGGCCTGAATTGTCTGCTGCAAGAACTTT 202 AA N61P f /5PHOS/CCGTTTCAAGATCCGAATTCTCATGGCACACACGTC 203 N61P r /5PHOS/TGCCATGAGAATTCGGATCTTGAAACGGGTTTGTTTCG 204 A187D f /5PHOS/TTCAAATCAACGTGATTCTTTTTCCTCCGTGGGACCGGAG 205 A187D r /5PHOS/ACGGAGGAAAAAGAATCACGTTGATTTGAAGAGTCTACAG 206 F189D f /5PHOS/CAAATCAACGTGCCTCTGATTCCTCCGTGGGACCGGAG 207 F189D r /5PHOS/CTCCGGTCCCACGGAGGAATCAGAGGCACGTTGATTTG 208 A137V f /5PHOS/AGCGGCAGTGCGGCACTTAAAGTTGCAGTTGATAAAGCT 209 GTTGC A137V r /5PHOS/GCAACAGCTTTATCAACTGCAACTTTAAGTGCCGCACTGC 210 CGCT S63T f /5PHOS/TCAAGATAACAATACACATGGCACACACGTCGCAGGAAC 211 S63T r /5PHOS/ACGTGTGTGCCATGTGTATTGTTATCTTGAAACGGGTTTG 212 Q103N f /5PHOS/AGACGGATCAGGCAATTACTCATGGATTATCAACGGCATC 213 Q103N r /5PHOS/TAATCCATGAGTAATTGCCTGATCCGTCTGCTGCAAG 214 N62D f /5PHOS/ACCCGTTTCAAGATAACGATTCTCATGGCACACACGTC 215 N62D r /5PHOS/GACGTGTGTGCCATGAGAATCGTTATCTTGAAACGGGT 216 N109D /5PHOS/CAATACTCATGGATTATCGATGGCATCGAATGGGCCA 217 f1 N109D r /5PHOS/TGGCCCATTCGATGCCATCGATAATCCATGAGTATTG 218 S188D f /5PHOS/CTCTTCAAATCAACGTGCCGATTTTTCCTCCGTGGGACC 219 S188D r /5PHOS/GGTCCCACGGAGGAAAAATCGGCACGTTGATTTGAAGAG 220 S248R f1 /5PHOS/CAAACACTCAAGTCCGCAGAAGTTTAGAAAACACCAC 221 S87R f /5PHOS/GTGCTTGGTGTAGCCCCGAGAGCTTCGCTCTACGCCGT 222 S87R r /5PHOS/ACGGCGTAGAGCGAAGCTCTCGGGGCTACACCAAGCAC 223 S248D f1 /5PHOS/CAAACACTCAAGTCCGCGATAGTTTAGAAAACACCAC 224 S248D r /5PHOS/GTGGTGTTTTCTAAACTATCGCGGACTTGAGTGTTTG 225 D87S f /5PHOS/GTGCTTGGTGTAGCCCCGTCTGCTTCGCTCTACGCCGT 226 D87S r /5PHOS/ACGGCGTAGAGCGAAGCAGACGGGGCTACACCAAGCAC 227 D109N f /5PHOS/CAATACTCATGGATTATCAACGGCATCGAATGGGCCA 228 D109N r /5PHOS/TGGCCCATTCGATGCCGTTGATAATCCATGAGTATTG 229 N109D /5PHOS/ACTCATGGATTATCGATGGCATCGAATGGGCCATCGC 230 f2 S248R f2 /5PHOS/CACTCAAGTCCGCAGAAGTTTAGAAAACACCACTAC 231 S248D f2 /5PHOS/CACTCAAGTCCGCGATAGTTTAGAAAACACCACTAC 232 S188R f /5PHOS/CAAATCAACGTGCCAGATTTTCCTCCGTGGGACCGGAG 233 QC AAAGGGGAGGAAAATCGTGAAACA 234 FUSION_For1 QC GTTCTAAATCGTGTTTTTCTTG 235 FUSION_Rev1 S248D GAACTGGACAAACACTCAAGTCCGCGATAGTTTAGAAAACACCA 236 forfus CTAC S248D GACTTGAGTGTTTGTCCAGTTCGGGTGCTTAGAAAG 237 revfus R248Sfor /5PHOS/CAAACACTCAAGTCCGCAGCAGTTTAGAAAACACCAC 238 R248S /5PHOS/GTGGTGTTTTCTAAACTGCTGCGGACTTGAGTGTTTG 239 rev G100E_Fsfor ACGCCGTTAAAGTTCTTGCAGCAGACGAATCAGGCCAATACTCAT 240 GGAT G100E_Fsrev TGCTGCAAGAACTTTAACGGCGTAGAGCGAAGCAGA 241 N123C f /5PHOS/ACATGGATGTAATCTGCATGAGCCTGGGAGGACCAAG 242 N123C r /5PHOS/TCCTCCCAGGCTCATGCAGATTACATCCATGTTATTCG 243 pHpLT /5PHOS/GTTATGAGTTAGTTCAAATTCG 244 R1
[0599] The parental DNA from the PCR samples was removed by addition of 14 of DpnI to Quik-Change reactions at 37.degree. C. overnight. One micro-liter of the DpnI-digested reactions were amplified using rolling circle amplification (RCA) using the Illustra TempliPhi kit according to the manufacturer's protocol. B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo) were transformed with 1 .mu.L each of the RCA reaction and the transformed cells were plated onto LA+1.6% skim milk plates containing 10 ppm neomycin and incubated at 37.degree. C. overnight. Colonies from overnight growth were selected to perform colony PCR using "puReTaq Ready-To-Go PCR Beads" (Amersham). The PCR primers used were pHPLT F1 (SEQ ID NO:54) and pHPLT seq R1 (SEQ ID NO:55). Clones with appropriate sequences were frozen. BPN' variants were expressed by growing the B. subtilis transformants in 96 well microtiter plates at 37.degree. C. for 68 hours in a MOPS based medium containing urea.
Generation of BPN' Variants Via Fusion PCR
[0600] Variants 5 and 33 were generated using Fusion PCR, with fragments amplified from template pHPLT-BPN'-v3. The PCR primers used to generate these variants are included in Table 11-5. For Variant 5, a 5' fragment of the BPN' gene was amplified using forward primer pHPLT F1(SEQ ID NO:54), and reverse primer S248D revfus. The 3' fragment of the BPN' gene was amplified using the forward primer S248D forfus containing the mutation of interest and the reverse primer pHPLT R1. The two products contained 20 bp of overlapping sequence, and were fused by combining 1 .mu.L of each fragment and fusion primers QC FUSION_For1 and QC FUSION_Rev1 in a final PCR reaction. All PCR reactions were performed using standard conditions of the Herculase II PCR Kit (Stratagene). The PCR mix contained 1 .mu.L DNA polymerase, 1 .mu.L plasmid DNA (or fragment DNA for fusion), 0.5 .mu.L dNTP's, 1.25 .mu.L 25 .mu.M forward primer, 1.25 .mu.L 25 .mu.M reverse primer, 10 .mu.L Buffer 5.times., 35 .mu.L H.sub.2O and the PCR program used was as follows: 95.degree. C. for 2 min, (95.degree. C. for 30 sec, 55.degree. C. for 30 sec, 72.degree. C. for "X" sec) for 29 cycles, 72.degree. C. for 1 min, 4.degree. C. hold (the "X" is 15 seconds per 1 kB of DNA to amplify).
[0601] For Variant 33, a 5' fragment of the BPN' gene was amplified using the template pHPLT-BPN'-v3 and primers pHPLT F1 (SEQ ID NO:54), and G100E_Fsrev. The 3' fragment that contained the variant was amplified using primers G100E_Fsfor and pHPLT R1. The two products contained 20 bp of overlapping sequence, and were fused by combining 1 .mu.l of each fragment and fusion primers QC FUSION_For1 and QC FUSION_Rev1 in a final PCR reaction. The PCR conditions were the same as listed above.
[0602] The two fusion products were purified using a QIAGEN.RTM. PCR purification column with conditions provided by the manufacturer, and digested overnight using Bgl I and HindIII enzymes. The plasmid pHPLT-BPN' partial opt was digested using the same enzymes and the vector band was gel extracted and purified over a QIAGEN.RTM. gel purification column using the manufacturer's recommendations. The restriction enzyme mix contained: 10 .mu.L purified DNA, 5 .mu.L Roche Buffer B, 0.54 HindIII, 0.54 Bgl I, 34 .mu.L H.sub.2O and the reactions were carried out at 37.degree. C. for 8 hours followed by 65.degree. C. for 20 min. The digest was purified using a QIAGEN.RTM. PCR purification column and ligated to the cut vector backbone overnight at 16.degree. C. using the Mighty Mix Ligase kit (Tekara). Following incubation, 1 .mu.L of the ligation mix was amplified using the Illustra TempliPhi kit.
[0603] For the amplification reaction, 14, of the ligation reaction mix was mixed with 5 .mu.L of sample buffer from the TempliPhi kit and heated for 3 minutes at 95.degree. C. to denature the DNA. The reaction was placed on ice to cool for 2 minutes and then spun down briefly. Five microliters of reaction buffer and 0.2 .mu.L of phi29 polymerase from the TempliPhi kit were added, and the reactions were incubated at 30.degree. C. in an MJ Research PCR machine for 4 hours. The phi29 enzyme was heat inactivated in the reactions by incubation at 65.degree. C. for 10 min in the PCR machine. Bacillus subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo) were transformed using 14 of the reaction mix and the transformants were grown overnight at 37.degree. C. on 1.6% skim milk plates containing 10 ppm neomycin. Transformants were selected to perform colony PCR and sequencing using "puReTaq Ready-To-Go PCR Beads" (Amersham) and primers pHPLT F1 (SEQ ID NO:54) and pHPLT seqR1 (SEQ ID NO:55).
4. Generation of BPN' Variants from Libraries RCL4-RCL7
RCL4 Library
[0604] "RCL4" refers to a group of site saturation libraries created by PCR fusion that simultaneously randomize three contiguous codons in the BPN'-v3-encoding (BPN''-G97A-G128A-Y217Q) gene. The amino acid positions corresponding to the three mutated codons in each library are provided in Table 11-6. Two partially overlapping, complementary mutagenic primers, each containing three degenerate codons were used to introduce mutations within each library as shown in Table 11-6 below. Only the first two nucleotides of each degenerate codon (NNX, N=A, C, T, or G and X is unchanged nucleotide) of interest were mutated in each primer (Table 11-6).
[0605] To create each library, two PCR reactions were carried out using either the common 3' gene-flanking primer (P4976, CCTCTCGGTTATGAGTTAGTTC; (SEQ ID NO:61)) and mutagenic primer, or the common 5'gene-flanking primer (P4974, GCCTCACATTTGTGCCACCTA; (SEQ ID NO:60)) and mutagenic primer as shown for each library in Table 11-6. These PCR reactions generated two PCR fragments, one encoding the 5' half of the mutant BPN'-v3 gene (5' gene fragment) and the other encoding the 3' half of the mutant BPN'-v3 gene (3' gene fragment). Each PCR amplification reaction contained 30 pmol of each primer and 100 ng of the BPN'-v3 parent template DNA (plasmid pHPLT-BPN'-v3) (see FIG. 1). Amplifications were carried out using Vent DNA polymerase (NEB). The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 40 sec. Following amplification, the 5' and 3' gene fragments were gel-purified by the QIAGEN.RTM. gel-band purification kit, mixed (50 ng of each fragment) and amplified by PCR once again using the primers P4973 (AAAGGATCCTAATCGGCGCTTTTC; SEQ ID NO:62) and P4950 (CTTGTCTCCAAGCTTAAAATAAAA; SEQ ID NO:63) to generate the full-length gene fragment. The PCR conditions were same as described above, except the extension phase, which was carried out at 72.degree. C. for 2 min. The full-length DNA fragment was gel-purified by the QIAGEN.RTM. gel-band purification kit, digested by the BamHI and HindIII restriction enzymes and ligated with the pHPLT-BPN' partial opt vector that also was digested with the same restriction enzymes. Ligation mixtures were amplified using rolling circle amplification in an Illustra Templiphi kit according to the manufacturer's recommendation (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. For this purpose, 1 .mu.l of the ligation mixture was mixed with 5 .mu.l of the sample buffer, heated to 95.degree. C. for 3 min and cooled on ice. Next, 5 .mu.l of the reaction buffer and 0.2 .mu.l of the enzyme were added to each tube, followed by incubation at 30.degree. C. for 10 hours. Products of the rolling circle amplification were diluted 100 times and used to transform B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C. Subsequently, the colonies with halos were inoculated in 120 .mu.l of LB media containing 10 .mu.g/mL neomycin.
TABLE-US-00041 TABLE 11-6 List of Primers Used to Create RCL4 Libraries Mutated Common residues flanking Mutagenic SEQ in BPN'- Gene primer primer Mutagenic primer ID Library # v3 fragment names names sequence NO: 1 5-7 3' P4976 P5119 TACGCGCAGTCCGT 245 GNNTNNCNNCGTAT CACAAATTAAAGCC CCTG 5' P4974 P5120 TTTAATTTGTGATA 246 CGNNGNNANNCAC GGACTGCGCGTACG CAT 2 11-13 3' P4976 P5121 TACGGCGTATCACA 247 ANNTNNANNCCCTG CTCTGCACTCTCAAG 5' P4974 P5122 AGAGTGCAGAGCA 248 GGGNNTNNANNTTG TGATACGCCGTAAG GCAC 3 20-22 3' P4976 P5123 GCTCTGCACTCTCA 249 ANNCNNCNNTGGAT CAAATGTTAAAGTA GCGGT 5' P4974 P5124 TTTAACATTTGATC 250 CANNGNNGNNTTGA GAGTGCAGAGCAG GGGCTT 4 21-23 3' P4976 P5125 CTGCACTCTCAAGG 251 CNNCNNTNNATCAA ATGTTAAAGTAGCG GTTATC 5' P4974 P5126 TACTTTAACATTTG 252 ATNNANNGNNGCCT TGAGAGTGCAGAGC AG 5 22-24 3' P4976 P5127 CACTCTCAAGGCTA 253 CNNTNNANNAAATG TTAAAGTAGCGGTT ATCGA 5' P4974 P5128 CGCTACTTTAACAT 254 TTNNTNNANNGTAG CCTTGAGAGTGCAG AG 6 23-25 3' P4976 P5129 TCTCAAGGCTACAC 255 TNNANNANNTGTTA AAGTAGCGGTTATC GACA 5' P4974 P5130 AACCGCTACTTTAA 256 CANNTNNTNNAGTG TAGCCTTGAGAGTG CAG 7 24-26 3' P4976 P5131 CAAGGCTACACTGG 257 ANNANNTNNTAAA GTAGCGGTTATCGA CAGC 5' P4974 P5132 GATAACCGCTACTT 258 TANNANNTNNTCCA GTGTAGCCTTGAGA GTG 8 25-27 3' P4976 P5133 GGCTACACTGGATC 259 ANNTNNTNNAGTAG CGGTTATCGACAGC GGT 5' P4974 P5134 GTCGATAACCGCTA 260 CTNNANNANNTGAT CCAGTGTAGCCTTG AGA 9 26-28 3' P4976 P5135 TACACTGGATCAAA 261 TNNTNNANNAGCGG TTATCGACAGCGGT AT 5' P4974 P5136 GCTGTCGATAACCG 262 CTNNTNNANNATTT GATCCAGTGTAGCC TTGA 10 27-29 3' P4976 P5137 ACTGGATCAAATGT 263 TNNANNANNGGTTA TCGACAGCGGTATC GAC 5' P4974 P5138 ACCGCTGTCGATAA 264 CCNNTNNTNNAACA TTTGATCCAGTGTA GCCT 11 28-30 3' P4976 P5139 GGATCAAATGTTAA 265 ANNANNGNNTATCG ACAGCGGTATCGAC TC 5' P4974 P5140 GATACCGCTGTCGA 266 TANNCNNTNNTTTA ACATTTGATCCAGT GTAGC 12 29-31 3' P4976 P5141 TCAAATGTTAAAGT 267 ANNGNNTNNCGAC AGCGGTATCGACTC GAGCCAT 5' P4974 P5142 GTCGATACCGCTGT 268 CGNNANNCNNTACT TTAACATTTGATCC AGTGTA 13 30-32 3' P4976 P5143 AATGTTAAAGTAGC 269 GNNTNNCNNCAGCG GTATCGACTCGAGC CAT 5' P4974 P5144 CGAGTCGATACCGC 270 TGNNGNNANNCGCT ACTTTAACATTTGA TCCAG 14 31-33 3' P4976 P5145 GTTAAAGTAGCGGT 271 TNNCNNCNNCGGTA TCGACTCGAGCCAT CCA 5' P4974 P5146 GCTCGAGTCGATAC 272 CGNNGNNGNNAAC CGCTACTTTAACAT TTGATC 15 32-34 3' P4976 P5147 AAAGTAGCGGTTAT 273 CNNCNNCNNTATCG ACTCGAGCCATCCA GAT 5' P4974 P5148 ATGGCTCGAGTCGA 274 TANNGNNGNNGAT AACCGCTACTTTAA CATTTG 16 33-35 3' P4976 P5149 GTAGCGGTTATCGA 275 CNNCNNTNNCGACT CGAGCCATCCAGAT CT 5' P4974 P5150 TGGATGGCTCGAGT 276 CGNNANNGNNGTC GATAACCGCTACTT TAACA 17 34-36 3' P4976 P5151 GCGGTTATCGACAG 277 CNNTNNCNNCTCGA GCCATCCAGATCTT AAAG 5' P4974 P5152 ATCTGGATGGCTCG 278 AGNNGNNANNGCT GTCGATAACCGCTA CTTT 18 35-37 3' P4976 P5153 GTTATCGACAGCGG 279 TNNCNNCNNGAGCC ATCCAGATCTTAAA GTC 5' P4974 P5154 AAGATCTGGATGGC 280 TCNNGNNGNNACCG CTGTCGATAACCGC TA 19 36-38 3' P4976 P5155 ATCGACAGCGGTAT 281 CNNCNNGNNCCATC CAGATCTTAAAGTC GCTG 5' P4974 P5156 TTTAAGATCTGGAT 282 GGNNCNNGNNGAT ACCGCTGTCGATAA CCGCTA 20 37-39 3' P4976 P5157 GACAGCGGTATCGA 283 CNNGNNCNNTCCAG ATCTTAAAGTCGCT GGA 5' P4974 P5158 GACTTTAAGATCTG 284 GANNGNNCNNGTC GATACCGCTGTCGA TAAC 21 38-40 3' P4976 P5159 AGCGGTATCGACTC 285 GNNCNNTNNAGATC TTAAAGTCGCTGGA GG 5' P4974 P5160 AGCGACTTTAAGAT 286 CTNNANNGNNCGA GTCGATACCGCTGT CGA 22 41-43 3' P4976 P5161 GACTCGAGCCATCC 287 ANNTNNTNNAGTCG CTGGAGGGGCTTCT AT 5' P4974 P5162 AGCCCCTCCAGCGA 288 CTNNANNANNTGGA TGGCTCGAGTCGAT AC 23 43-45 3' P4976 P5163 AGCCATCCAGATCT 289 TNNANNCNNTGGAG GGGCTTCTATGGTG CCGT 5' P4974 P5164 CATAGAAGCCCCTC 290 CNAGCGACTTTNAA GATCTGGATGGCTC GAGTC 24 44-46 3' P4976 P5165 CATCCAGATCTTAA 291 ANNCNNTNNAGGG GCTTCTATGGTGCC GT 5' P4974 P5166 CACCATAGAAGCCC 292 CTNNANNGNNTTTA AGATCTGGATGGCT CGAG 25 51-53 3' P4976 P5167 GGAGGGGCTTCTAT 293 GNNGNNGNNCGAA ACAAACCCGTTTCA AGATAA 5' P4974 P5168 AAACGGGTTTGTTT 294 CGNNCNNCNNCATA GAAGCCCCTCCAGC GA 26 52-54 3' P4976 P5169 GGGGCTTCTATGGT 295 GNNGNNCNNAACA AACCCGTTTCAAGA TAACAA 5' P4974 P5170 TTGAAACGGGTTTG 296 TTNNGNNCNNCACC ATAGAAGCCCCTCC AG 27 53-55 3' P4976 P5171 GCTTCTATGGTGCC 297 GNNCNNANNAAAC CCGTTTCAAGATAA CAATTC 5' P4974 P5172 ATCTTGAAACGGGT 298 TTNNTNNGNNCGGC ACCATAGAAGCCCC TC
28 54-56 3' P4976 P5173 TCTATGGTGCCGTC 299 CNNANNANNCCCGT TTCAAGATAACAAT TCTCA 5' P4974 P5174 GTTATCTTGAAACG 300 GGNNTNNTNNGGAC GGCACCATAGAAGC CCCT 29 55-57 3' P4976 P5175 ATGGTGCCGTCCGA 301 ANNANNCNNGTTTC AAGATAACAATTCT CATGG 5' P4974 P5176 ATTGTTATCTTGAA 302 ACNNGNNTNNTTCG GACGGCACCATAGA AG 30 56-58 3' P4976 P5177 GTGCCGTCCGAAAC 303 ANNCNNGNNTCAA GATAACAATTCTCA TGGCAC 5' P4974 P5178 AGAATTGTTATCTT 304 GANNCNNGNNTGTT TCGGACGGCACCAT AGA 31 57-59 3' P4976 P5179 CCGTCCGAAACAAA 305 CNNGNNTNNAGATA ACAATTCTCATGGC ACAC 5' P4974 P5180 ATGAGAATTGTTAT 306 CTNNANNCNNGTTT GTTTCGGACGGCAC CA 32 58-60 3' P4976 P5181 TCCGAAACAAACCC 307 GNNTNNANNTAACA ATTCTCATGGCACA CAC 5' P4974 P5182 GCCATGAGAATTGT 308 TANNTNNANNCGGG TTTGTTTCGGACGG CA 33 59-61 3' P4976 P5183 GAAACAAACCCGTT 309 TNNANNTNNCAATT CTCATGGCACACAC GTC 5' P4974 P5184 TGTGCCATGAGAAT 310 TGNNANNTNNAAAC GGGTTTGTTTCGGA CG 34 60-62 3' P4976 P5185 ACAAACCCGTTTCA 311 ANNTNNCNNTTCTC ATGGCACACACGTCG 5' P4974 P5186 GTGTGTGCCATGAG 312 AANNGNNANNTTG AAACGGGTTTGTTT CGGAC 35 61-63 3' P4976 P5187 AACCCGTTTCAAGA 313 TNNCNNTNNTCATG GCACACACGTCGCAG 5' P4974 P5188 GACGTGTGTGCCAT 314 GANNANNGNNATCT TGAAACGGGTTTGT TTCG 36 62-64 3' P4976 P5189 CCGTTTCAAGATAA 315 CNNTNNTNNTGGCA CACACGTCGCAGGAA 5' P4974 P5190 TGCGACGTGTGTGC 316 CANNANNANNGTTA TCTTGAAACGGGTT TGTTT 37 67-69 3' P4976 P5191 AATTCTCATGGCAC 317 ANNCNNCNNAGGA ACGGTTGCGGCGTT AAA 5' P4974 P5192 CGCCGCAACCGTTC 318 CTNNGNNGNNTGTG CCATGAGAATTGTT ATCTT 38 68-70 3' P4976 P5193 TCTCATGGCACACA 319 CNNCNNANNAACG GTTGCGGCGTTAAA CAAT 5' P4974 P5194 TAACGCCGCAACCG 320 TTNNTNNGNNGTGT GTGCCATGAGAATT GTTA 39 71-73 3' P4976 P5195 ACACACGTCGCAGG 321 ANNGNNTNNGGCGT TAAACAATTCTATT GGCGT 5' P4974 P5196 AGAATTGTTTAACG 322 CCNNANNCNNTCCT GCGACGTGTGTGCC AT 40 74-76 3' P4976 P5197 GCAGGAACGGTTGC 323 GNNGNNANNCAATT CTATTGGCGTGCTT GGTG 5' P4974 P5198 CACGCCAATAGAAT 324 TGNNTNNCNNCGCA ACCGTTCCTGCGAC GT 41 77-79 3' P4976 P5199 ACGGTTGCGGCGTT 325 ANNCNNTNNTATTG GCGTGCTTGGTGTA GC 5' P4974 P5200 ACCAAGCACGCCAA 326 TANNANNGNNTAAC GCCGCAACCGTTCC TG 42 81-83 3' P4976 P5201 AACAATTCTATTGG 327 CNNGNNTNNTGTAG CCCCGTCTGCTTCG CT 5' P4974 P5202 AGCAGACGGGGCTA 328 CANNANNCNNGCC AATAGAATTGTTTA ACGCCGCAA 43 83-85 3' P4976 P5203 TCTATTGGCGTGCT 329 TNNTNNANNCCCGT CTGCTTCGCTCTACG 5' P4974 P5204 GAGCGAAGCAGAC 330 GGGNNTNNANNAA GCACGCCAATAGAA TTGTTTA 44 84-86 3' P4976 P5205 ATTGGCGTGCTTGG 331 TNNANNCNNGTCTG CTTCGCTCTACGCC GT 5' P4974 P5206 GTAGAGCGAAGCA 332 GACNNGNNTNNACC AAGCACGCCAATAG AATTG 45 85-87 3' P4976 P5207 TGGCGTGCTTGGTG 333 TANNCNNGNNTGCT TCGCTCTACGCCGT TAA 5' P4974 P5208 GGCGTAGAGCGAA 334 GCANNCNNGNNTAC ACCAAGCACGCCAA TAGA 46 86-88 3' P4976 P5209 GTGCTTGGTGTAGC 335 CNNGNNTNNTTCGC TCTACGCCGTTAAA GTT 5' P4974 P5210 AACGGCGTAGAGCG 336 AANNANNCNNGGC TACACCAAGCACGC CAA 47 87-89 3' P4976 P5211 CTTGGTGTAGCCCC 337 GNNTNNTNNGCTCT ACGCCGTTAAAGTT CTT 5' P4974 P5212 TTTAACGGCGTAGA 338 GCNNANNANNCGG GGCTACACCAAGCA CGCCAAT 48 88-90 3' P4976 P5213 GGTGTAGCCCCGTC 339 TNNTNNGNNCTACG CCGTTAAAGTTCTT GCAG 5' P4974 P5214 AACTTTAACGGCGT 340 AGNNCNNANNAGA CGGGGCTACACCAA GCA 49 89-91 3' P4976 P5215 GTAGCCCCGTCTGC 341 TNNGNNCNNCGCCG TTAAAGTTCTTGCA GCA 5' P4974 P5216 AAGAACTTTAACGG 342 CGNNGNNCNNAGC AGACGGGGCTACAC CAA 50 90-92 3' P4976 P5217 GCCCCGTCTGCTTC 343 GNNCNNCNNCGTTA AAGTTCTTGCAGCA GAC 5' P4974 P5218 TGCAAGAACTTTAA 344 CGNNGNNGNNCGA AGCAGACGGGGCTA CAC 51 91-93 3' P4976 P5219 CCGTCTGCTTCGCT 345 CNNCNNCNNTAAAG TTCTTGCAGCAGAC GGA 5' P4974 P5220 TGCTGCAAGAACTT 346 TANNGNNGNNGAG CGAAGCAGACGGG GCT 52 92-94 3' P4976 P5221 TCTGCTTCGCTCTAC 347 NNCNNTNNAGTTCT TGCAGCAGACGGATC 5' P4974 P5222 GTCTGCTGCAAGAA 348 CTNNANNGNNGTAG AGCGAAGCAGACG GGGCTA 53 93-95 3' P4976 P5223 GCTTCGCTCTACGC 349 CNNTNNANNTCTTG CAGCAGACGGATCAG 5' P4974 P5224 TCCGTCTGCTGCAA 350 GANNTNNANNGGC GTAGAGCGAAGCA GACG 54 94-96 3' P4976 P5225 TCGCTCTACGCCGT 351 TNNANNTNNTGCAG CAGACGGATCAGGC CA 5' P4974 P5226 TGATCCGTCTGCTG 352 CANNANNTNNAAC GGCGTAGAGCGAA GCAG 55 95-97 3' P4976 P5227 CTCTACGCCGTTAA 353 ANNTNNTNNAGCAG ACGGATCAGGCCAA TA 5' P4974 P5228 GCCTGATCCGTCTG 354 CTNNANNANNTTTA ACGGCGTAGAGCGA AG 56 96-98 3' P4976 P5229 TACGCCGTTAAAGT 355 TNNTNNANNAGACG GATCAGGCCAATAC TC 5' P4974 P5230 TTGGCCTGATCCGT 356
CTNNTNNANNAACT TTAACGGCGTAGAG CGA 57 97-99 3' P4976 P5231 GCCGTTAAAGTTCT 357 TNNANNANNCGGAT CAGGCCAATACTCA TG 5' P4974 P5232 GTATTGGCCTGATC 358 CGNNTNNTNNAAGA ACTTTAACGGCGTA GAG 58 98-100 3' P4976 P5233 GTTAAAGTTCTTGC 359 ANNANNCNNATCA GGCCAATACTCATG GATTA 5' P4974 P5234 TGAGTATTGGCCTG 360 ATNNGNNTNNTGCA AGAACTTTAACGGC GTAG 59 99-101 3' P4976 P5235 AAAGTTCTTGCAGC 361 ANNCNNANNAGGC CAATACTCATGGAT TATC 5' P4974 P5236 CCATGAGTATTGGC 362 CTNNTNNGNNTGCT GCAAGAACTTTAAC GGCGTA 60 100-102 3' P4976 P5237 GTTCTTGCAGCAGA 363 CNNANNANNCCAAT ACTCATGGATTATC AAC 5' P4974 P5238 AATCCATGAGTATT 364 GGNNTNNTNNGTCT GCTGCAAGAACTTT AACG 61 101-103 3' P4976 P5239 CTTGCAGCAGACGG 365 ANNANNCNNATACT CATGGATTATCAAC GGCA 5' P4974 P5240 GATAATCCATGAGT 366 ATNNGNNTNNTCCG TCTGCTGCAAGAAC TTT 62 102-104 3' P4976 P5241 GCAGCAGACGGATC 367 ANNCNNANNCTCAT GGATTATCAACGGC ATC 5' P4974 P5242 TTGATAATCCATGA 368 GNNTNNGNNTGATC CGTCTGCTGCAAGA AC 63 103-105 3' P4976 P5243 GCAGACGGATCAGG 369 CNNANNCNNATGG ATTATCAACGGCAT CGAAT 5' P4974 P5244 GCCGTTGATAATCC 370 ATNNGNNTNNGCCT GATCCGTCTGCTGC AA 64 104-106 3' P4976 P5245 GACGGATCAGGCCA 371 ANNCNNANNGATTA TCAACGGCATCGAA TGG 5' P4974 P5246 GATGCCGTTGATAA 372 TCNNTNNGNNTTGG CCTGATCCGTCTGC TG 65 105-107 3' P4976 P5247 GGATCAGGCCAATA 373 CNNANNGNNTATCA ACGGCATCGAATGG GCCAT 5' P4974 P5248 TTCGATGCCGTTGA 374 TANNCNNTNNGTAT TGGCCTGATCCGTC TG 66 106-108 3' P4976 P5249 TCAGGCCAATACTC 375 ANNGNNTNNCAAC GGCATCGAATGGGC CAT 5' P4974 P5250 CCATTCGATGCCGT 376 TGNNANNCNNTGAG TATTGGCCTGATCC GTC 67 107-109 3' P4976 P5251 GGCCAATACTCATG 377 GNNTNNCNNCGGCA TCGAATGGGCCATC GCGAAT 5' P4974 P5252 GGCCCATTCGATGC 378 CGNNGNNANNCCAT GAGTATTGGCCTGA TCC 68 108-110 3' P4976 P5253 CAATACTCATGGAT 379 TNNCNNCNNCATCG AATGGGCCATCGCG AA 5' P4974 P5254 GATGGCCCATTCGA 380 TGNNGNNGNNAATC CATGAGTATTGGCC TGAT 69 109-111 3' P4976 P5255 TACTCATGGATTAT 381 CNNCNNCNNCGAAT GGGCCATCGCGAAT AA 5' P4974 P5256 CGCGATGGCCCATT 382 CGNNGNNGNNGAT AATCCATGAGTATT GGCCT 70 110-112 3' P4976 P5257 TCATGGATTATCAA 383 CNNCNNCNNATGGG CCATCGCGAATAAC ATG 5' P4974 P5258 ATTCGCGATGGCCC 384 ATNNGNNGNNGTTG ATAATCCATGAGTA TTGG 71 111-113 3' P4976 P5259 TGGATTATCAACGG 385 CNNCNNANNGGCC ATCGCGAATAACAT GGA 5' P4974 P5260 GTTATTCGCGATGG 386 CCNNTNNGNNGCCG TTGATAATCCATGA GTAT 72 112-114 3' P4976 P5261 ATTATCAACGGCAT 387 CNNANNGNNCATCG CGAATAACATGGAT GTAA 5' P4974 P5262 CATGTTATTCGCGA 388 TGNNCNNTNNGATG CCGTTGATAATCCA TGAG 73 113-115 3' P4976 P5263 ATCAACGGCATCGA 389 ANNGNNCNNCGCG AATAACATGGATGT AATCAA 5' P4974 P5264 ATCCATGTTATTCG 390 CGNNGNNCNNTTCG ATGCCGTTGATAAT CCAT 74 114-116 3' P4976 P5265 AACGGCATCGAATG 391 GNNCNNCNNGAAT AACATGGATGTAAT CAACAT 5' P4974 P5266 TACATCCATGTTAT 392 TCNNGNNGNNCCAT TCGATGCCGTTGAT AATC 75 115-117 3' P4976 P5267 GGCATCGAATGGGC 393 CNNCNNGNNTAACA TGGATGTAATCAAC ATGAG 5' P4974 P5268 GATTACATCCATGT 394 TANNCNNGNNGGCC CATTCGATGCCGTT GA 76 116-118 3' P4976 P5269 ATCGAATGGGCCAT 395 CNNGNNTNNCATGG ATGTAATCAACATG AGCCT 5' P4974 P5270 GTTGATTACATCCA 396 TGNNANNCNNGATG GCCCATTCGATGCC GT 77 117-119 3' P4976 P5271 GAATGGGCCATCGC 397 GNNTNNCNNGGATG TAATCAACATGAGC CTG 5' P4974 P5272 CATGTTGATTACAT 398 CCNNGNNANNCGC GATGGCCCATTCGA TGC 78 118-120 3' P4976 P5273 TGGGCCATCGCGAA 399 TNNCNNGNNTGTAA TCAACATGAGCCTG GGA 5' P4974 P5274 GCTCATGTTGATTA 400 CANNCNNGNNATTC GCGATGGCCCATTC GA 79 119-121 3' P4976 P5275 GCCATCGCGAATAA 401 CNNGNNTNNAATCA ACATGAGCCTGGGA GCA 5' P4974 P5276 CAGGCTCATGTTGA 402 TTNNANNCNNGTTA TTCGCGATGGCCCA TTC 80 120-122 3' P4976 P5277 ATCGCGAATAACAT 403 GNNTNNANNCAAC ATGAGCCTGGGAGC AC 5' P4974 P5278 TCCCAGGCTCATGT 404 TGNNTNNANNCATG TTATTCGCGATGGC CCAT 81 121-123 3' P4976 P5279 GCGAATAACATGGA 405 TNNANNCNNCATGA GCCTGGGAGCACCA AG 5' P4974 P5280 TGCTCCCAGGCTCA 406 TGNNGNNTNNATCC ATGTTATTCGCGAT GGCCCATT 82 122-124 3' P4976 P5281 AATAACATGGATGT 407 ANNCNNCNNGAGC CTGGGAGCACCAAG CGGCA 5' P4974 P5282 TGGTGCTCCCAGGC 408 TCNNGNNGNNTACA TCCATGTTATTCGC GATG 83 123-125 3' P4976 P5283 AACATGGATGTAAT 409 CNNCNNGNNCCTGG GAGCACCAAGCGGCA 5' P4974 P5284 GCTTGGTGCTCCCA 410 GGNNCNNGNNGATT ACATCCATGTTATT CGCGA 84 124-126 3' P4976 P5285 ATGGATGTAATCAA 411 CNNGNNCNNGGGA GCACCAAGCGGCAG TG 5' P4974 P5286 GCCGCTTGGTGCTC 412
CCNNGNNCNNGTTG ATTACATCCATGTT ATTCG 85 125-127 3' P4976 P5287 GATGTAATCAACAT 413 GNNCNNGNNAGCA CCAAGCGGCAGTGC GGCA 5' P4974 P5288 ACTGCCGCTTGGTG 414 CTNNCNNGNNCATG TTGATTACATCCAT GTTATT 86 126-128 3' P4976 P5289 GTAATCAACATGAG 415 CNNGNNANNACCA AGCGGCAGTGCGGC ACT 5' P4974 P5290 CGCACTGCCGCTTG 416 GTNNTNNCNNGCTC ATGTTGATTACATC CATG 87 129-131 3' P4976 P5291 ATGAGCCTGGGAGC 417 ANNANNCNNCAGT GCGGCACTTAAAGC AGCA 5' P4974 P5292 TTTAAGTGCCGCAC 418 TGNNGNNTNNTGCT CCCAGGCTCATGTT GAT 88 132-134 3' P4976 P5293 GGAGCACCAAGCG 419 GCNNTNNGNNACTT AAAGCAGCAGTTGA TAAAG 5' P4974 P5294 AACTGCTGCTTTAA 420 GTNNCNNANNGCCG CTTGGTGCTCCCAG GCT 89 133-135 3' P4976 P5295 GCACCAAGCGGCAG 421 TNNGNNANNTAAA GCAGCAGTTGATAA AGCTG 5' P4974 P5296 ATCAACTGCTGCTT 422 TANNTNNCNNACTG CCGCTTGGTGCTCC CA 90 134-136 3' P4976 P5297 CCAAGCGGCAGTGC 423 GNNANNTNNAGCA GCAGTTGATAAAGC TGTTG 5' P4974 P5298 TTTATCAACTGCTG 424 CTNNANNTNNCGCA CTGCCGCTTGGTGC TC 91 135-137 3' P4976 P5299 AGCGGCAGTGCGGC 425 ANNTNNANNAGCA GTTGATAAAGCTGT TGCAT 5' P4974 P5300 AGCTTTATCAACTG 426 CTNNTNNANNTGCC GCACTGCCGCTTGG TG 92 136-138 3' P4976 P5301 GGCAGTGCGGCACT 427 TNNANNANNAGTTG ATAAAGCTGTTGCA TCTG 5' P4974 P5302 AACAGCTTTATCAA 428 CTNNTNNTNNAAGT GCCGCACTGCCGCT TG 93 137-139 3' P4976 P5303 AGTGCGGCACTTAA 429 ANNANNANNTGAT AAAGCTGTTGCATC TGGTG 5' P4974 P5304 TGCAACAGCTTTAT 430 CANNTNNTNNTTTA AGTGCCGCACTGCC GCTT 94 138-140 3' P4976 P5305 GCGGCACTTAAAGC 431 ANNANNTNNTAAA GCTGTTGCATCTGG TGTC 5' P4974 P5306 AGATGCAACAGCTT 432 TANNANNTNNTGCT TTAAGTGCCGCACT GC 95 139-141 3' P4976 P5307 GCACTTAAAGCAGC 433 ANNTNNTNNAGCTG TTGCATCTGGTGTC GT 5' P4974 P5308 ACCAGATGCAACAG 434 CTNNANNANNTGCT GCTTTAAGTGCCGC AC 96 140-142 3' P4976 P5309 CTTAAAGCAGCAGT 435 TNNTNNANNTGTTG CATCTGGTGTCGTC GT 5' P4974 P5310 GACACCAGATGCAA 436 CANNTNNANNAACT GCTGCTTTAAGTGC CGCA 97 141-143 3' P4976 P5311 AAAGCAGCAGTTGA 437 TNNANNTNNTGCAT CTGGTGTCGTCGTA GT 5' P4974 P5312 GACGACACCAGATG 438 CANNANNTNNATCA ACTGCTGCTTTAAG TGC 98 142-144 3' P4976 P5313 GCAGCAGTTGATAA 439 ANNTNNTNNATCTG GTGTCGTCGTAGTA GC 5' P4974 P5314 TACGACGACACCAG 440 ATNNANNANNTTTA TCAACTGCTGCTTT AAGTG 99 143-145 3' P4976 P5315 GCAGTTGATAAAGC 441 TNNTNNANNTGGTG TCGTCGTAGTAGCG GCA 5' P4974 P5316 TACTACGACGACAC 442 CANNTNNANNAGCT TTATCAACTGCTGC TTTAA 100 144-146 3' P4976 P5317 GTTGATAAAGCTGT 443 TNNANNTNNTGTCG TCGTAGTAGCGGCA GCT 5' P4974 P5318 CGCTACTACGACGA 444 CANNANNTNNAAC AGCTTTATCAACTG CTGCT 101 145-147 3' P4976 P5319 GATAAAGCTGTTGC 445 ANNTNNTNNCGTCG TAGTAGCGGCAGCTG 5' P4974 P5320 TGCCGCTACTACGA 446 CGNNANNANNTGC AACAGCTTTATCAA CTGCT 102 146-148 3' P4976 P5321 AAAGCTGTTGCATC 447 TNNTNNCNNCGTAG TAGCGGCAGCTGGG AA 5' P4974 P5322 AGCTGCCGCTACTA 448 CGNNGNNANNAGA TGCAACAGCTTTAT CAACTG 103 147-149 3' P4976 P5323 GCTGTTGCATCTGG 449 TNNCNNCNNAGTAG CGGCAGCTGGGAAT GA 5' P4974 P5324 CCCAGCTGCCGCTA 450 CTNNGNNGNNACCA GATGCAACAGCTTT ATCA 104 148-150 3' P4976 P5325 GTTGCATCTGGTGT 451 CNNCNNANNAGCG GCAGCTGGGAATGA GGGAA 5' P4974 P5326 ATTCCCAGCTGCCG 452 CTNNTNNGNNGACA CCAGATGCAACAGC TTT 105 149-151 3' P4976 P5327 GCATCTGGTGTCGT 453 CNNANNANNGGCA GCTGGGAATGAGGG AAC 5' P4974 P5328 CTCATTCCCAGCTG 454 CCNNTNNTNNGACG ACACCAGATGCAAC AG 106 150-152 3' P4976 P5329 TCTGGTGTCGTCGT 455 ANNANNGNNAGCT GGGAATGAGGGAA CATC 5' P4974 P5330 TCCCTCATTCCCAG 456 CTNNCNNTNNTACG ACGACACCAGATGC AAC 107 151-153 3' P4976 P5331 GGTGTCGTCGTAGT 457 ANNGNNANNTGGG AATGAGGGAACATC CGGAT 5' P4974 P5332 TGTTCCCTCATTCCC 458 ANNTNNCNNTACTA CGACGACACCAGAT GCA 108 158-160 3' P4976 P5333 GCTGGGAATGAGGG 459 ANNANNCNNATCAT CGAGTACCGTCGGT TAT 5' P4974 P5334 GACGGTACTCGATG 460 ATNNGNNTNNTCCC TCATTCCCAGCTGC CGCTA 109 159-161 3' P4976 P5335 GGGAATGAGGGAA 461 CANNCNNANNATCG AGTACCGTCGGTTA TCCA 5' P4974 P5336 ACCGACGGTACTCG 462 ATNNTNNGNNTGTT CCCTCATTCCCAGC TG 110 160-162 3' P4976 P5337 AATGAGGGAACATC 463 CNNANNANNGAGT ACCGTCGGTTATCC AGG 5' P4974 P5338 ATAACCGACGGTAC 464 TCNNTNNTNNGGAT GTTCCCTCATTCCC AG 111 163-165 3' P4976 P5339 ACATCCGGATCATC 465 GNNTNNCNNCGGTT ATCCAGGCAAGTAC CCTT 5' P4974 P5340 CTTGCCTGGATAAC 466 CGNNGNNANNCGA TGATCCGGATGTTC CCT 112 164-166 3' P4976 P5341 TCCGGATCATCGAG 467 TNNCNNCNNTTATC CAGGCAAGTACCCT TCA 5' P4974 P5342 GTACTTGCCTGGAT 468
AANNGNNGNNACT CGATGATCCGGATG TTCC 113 167-169 3' P4976 P5343 TCGAGTACCGTCGG 469 TNNTNNANNCAAGT ACCCTTCAGTGATT GCA 5' P4974 P5344 CACTGAAGGGTACT 470 TGNNTNNANNACCG ACGGTACTCGATGA TC 114 168-170 3' P4976 P5345 AGTACCGTCGGTTA 471 TNNANNCNNGTACC CTTCAGTGATTGCA GTG 5' P4974 P5346 AATCACTGAAGGGT 472 ACNNGNNTNNATAA CCGACGGTACTCGA TGA 115 169-171 3' P4976 P5347 ACCGTCGGTTATCC 473 ANNCNNGNNCCCTT CAGTGATTGCAGTGG 5' P4974 P5348 TGCAATCACTGAAG 474 GGNNCNNGNNTGG ATAACCGACGGTAC TCGA 116 170-172 3' P4976 P5349 GTCGGTTATCCAGG 475 CNNGNNCNNTTCAG TGATTGCAGTGGGC GCT 5' P4974 P5350 CACTGCAATCACTG 476 AANNGNNCNNGCCT GGATAACCGACGGT AC 117 171-173 3' P4976 P5351 GGTTATCCAGGCAA 477 GNNCNNTNNAGTGA TTGCAGTGGGCGCT GTA 5' P4974 P5352 GCCCACTGCAATCA 478 CTNNANNGNNCTTG CCTGGATAACCGAC GGTA 118 172-174 3' P4976 P5353 TATCCAGGCAAGTA 479 CNNTNNANNGATTG CAGTGGGCGCTGTA GA 5' P4974 P5354 AGCGCCCACTGCAA 480 TCNNTNNANNGTAC TTGCCTGGATAACC GAC 119 182-184 3' P4976 P5355 GTGGGCGCTGTAGA 481 CNNTNNANNTCAAC GTGCCTCTTTTTCCTC 5' P4974 P5356 AAAAGAGGCACGTT 482 GANNTNNANNGTCT ACAGCGCCCACTGC AA 120 183-185 3' P4976 P5357 GGCGCTGTAGACTC 483 TNNANNTNNACGTG CCTCTTTTTCCTCCGT 5' P4974 P5358 GGAAAAAGAGGCA 484 CGTNNANNTNNAGA GTCTACAGCGCCCA CTG 121 184-186 3' P4976 P5359 GCTGTAGACTCTTC 485 ANNTNNANNTGCCT CTTTTTCCTCCGTGG GA 5' P4974 P5360 GGAGGAAAAAGAG 486 GCANNTNNANNTGA AGAGTCTACAGCGC CCA 122 185-187 3' P4976 P5361 GTAGACTCTTCAAA 487 TNNANNTNNCTCTT TTTCCTCCGTGGGAC 5' P4974 P5362 CACGGAGGAAAAA 488 GAGNNANNTNNATT TGAAGAGTCTACAG CGCCCA 123 186-188 3' P4976 P5363 GACTCTTCAAATCA 489 ANNTNNCNNTTTTT CCTCCGTGGGACCG GA 5' P4974 P5364 TCCCACGGAGGAAA 490 AANNGNNANNTTG ATTTGAAGAGTCTA CAGCGCCCA 124 192-194 3' P4976 P5365 GCCTCTTTTTCCTCC 491 NNGNNANNGGAGC TGGATGTCATGGCC CCT 5' P4974 P5366 CATGACATCCAGCT 492 CCNNTNNCNNGGAG GAAAAAGAGGCAC GTTG 125 194-196 3' P4976 P5367 TTTTCCTCCGTGGG 493 ANNGNNGNNGGAT GTCATGGCCCCTGG CGTT 5' P4974 P5368 AGGGGCCATGACAT 494 CCNNCNNCNNTCCC ACGGAGGAAAAAG AGG 126 195-197 3' P4976 P5369 TCCTCCGTGGGACC 495 GNNGNNGNNTGTCA TGGCCCCTGGCGTT TCTATT 5' P4974 P5370 GCCAGGGGCCATGA 496 CANNCNNCNNCGGT CCCACGGAGGAAA AAG 127 196-198 3' P4976 P5371 TCCGTGGGACCGGA 497 GNNGNNTNNCATGG CCCCTGGCGTTTCT ATT 5' P4974 P5372 AACGCCAGGGGCCA 498 TGNNANNCNNCTCC GGTCCCACGGAGGA AAAA 128 197-199 3' P4976 P5373 GTGGGACCGGAGCT 499 GNNTNNCNNGGCCC CTGGCGTTTCTATTC AA 5' P4974 P5374 AGAAACGCCAGGG 500 GCCNNGNNANNCA GCTCCGGTCCCACG GAGGAAA 129 198-200 3' P4976 P5375 GGACCGGAGCTGGA 501 TNNCNNGNNCCCTG GCGTTTCTATTCAA TCGA 5' P4974 P5376 AATAGAAACGCCAG 502 GGNNCNNGNNATCC AGCTCCGGTCCCAC GGA 130 203-205 3' P4976 P5377 GTCATGGCCCCTGG 503 CNNTNNTNNTCAAT CGACGCTTCCAGGG AA 5' P4974 P5378 TGGAAGCGTCGATT 504 GANNANNANNGCC AGGGGCCATGACAT CCA 131 210-212 3' P4976 P5379 ATTCAATCGACGCT 505 TNNANNGNNCAAGT ATGGTGCGCAAAAC GGGA 5' P4974 P5380 TTGCGCACCATACT 506 TGNNCNNTNNAAGC GTCGATTGAATAGA AACG 132 211-213 3' P4976 P5381 CAATCGACGCTTCC 507 ANNGNNCNNGTATG GTGCGCAAAACGGG ACT 5' P4974 P5382 GTTTTGCGCACCAT 508 ACNNGNNCNNTGG AAGCGTCGATTGAA TAGAA 133 216-218 3' P4976 P5383 GGGAACAAGTATGG 509 TNNGNNANNCGGG ACTTCCATGGCCTC GCCGCAT 5' P4974 P5384 GGCCATGGAAGTCC 510 CGNNTNNCNNACCA TACTTGTTCCCTGG AAG 134 217-219 3' P4976 P5385 AACAAGTATGGTGC 511 GNNANNCNNGACTT CCATGGCCTCGCCG CAT 5' P4974 P5386 CGAGGCCATGGAAG 512 TCNNGNNTNNCGCA CCATACTTGTTCCCTG 135 218-220 3' P4976 P5387 AAGTATGGTGCGCA 513 ANNCNNGNNTTCCA TGGCCTCGCCGCATG 5' P4974 P5388 CGGCGAGGCCATGG 514 AANNCNNGNNTTGC GCACCATACTTGTT CCC 136 219-221 3' P4976 P5389 TATGGTGCGCAAAA 515 CNNGNNTNNCATGG CCTCGCCGCATGTAG 5' P4974 P5390 ATGCGGCGAGGCCA 516 TGNNANNCNNGTTT TGCGCACCATACTT GTTC 137 230-232 3' P4976 P5391 CCGCATGTAGCTGG 517 GNNGNNCNNATTGA TTCTTTCTAAGCAC CCGAA 5' P4974 P5392 CTTAGAAAGAATCA 518 ATNNGNNCNNCCCA GCTACATGCGGCGA GGCCAT 138 231-233 3' P4976 P5393 CATGTAGCTGGGGC 519 GNNCNNANNGATTC TTTCTAAGCACCCG AACT 5' P4974 P5394 GTGCTTAGAAAGAA 520 TCNNTNNGNNCGCC CCAGCTACATGCGG CGAGGCCAT 139 232-234 3' P4976 P5395 GTAGCTGGGGCGGC 521 CNNANNGNNTCTTT CTAAGCACCCGAAC TG 5' P4974 P5396 CGGGTGCTTAGAAA 522 GANNCNNTNNGGCC GCCCCAGCTACATGC 140 238-240 3' P4976 P5397 TTGATTCTTTCTAAG 523 NNCNNGNNCTGGAC AAACACTCAAGTCC GCA 5' P4974 P5398 TTGAGTGTTTGTCC 524 AGNNCNNGNNCTTA GAAAGAATCAATGC GGC 141 240-242 3' P4976 P5399 CTTTCTAAGCACCC 525 GNNCNNGNNAAAC ACTCAAGTCCGCAG
CAGT 5' P4974 P5400 GCGGACTTGAGTGT 526 TTNNCNNGNNCGGG TGCTTAGAAAGAAT CAAT 142 246-248 3' P4976 P5401 TGGACAAACACTCA 527 ANNCNNCNNCAGTT TAGAAAACACCACT ACAAAA 5' P4974 P5402 GGTGTTTTCTAAAC 528 TGNNGNNGNNTTGA GTGTTTGTCCAGTT CGGGT 143 255-257 3' P4976 P5403 TTAGAAAACACCAC 529 TNNANNANNTGGTG ATTCTTTCTACTATG GAAA 5' P4974 P5404 GTAGAAAGAATCAC 530 CANNTNNTNNAGTG GTGTTTTCTAAACT GCTG 144 258-260 3' P4976 P5405 ACCACTACAAAACT 531 TNNTNNTNNTTTCT ACTATGGAAAAGGG CTGA 5' P4974 P5406 TTTTCCATAGTAGA 532 AANNANNANNAAG TTTTGTAGTGGTGTT TTCTAA 145 265-267 3' P4976 P5407 GATTCTTTCTACTAT 533 NNANNANNGCTGAT CAACGTACAGGCGG CA 5' P4974 P5408 CTGTACGTTGATCA 534 GCNNTNNTNNATAG TAGAAAGAATCACC AAGTTT
RCL5 Variants
[0606] "RCL5" refers to a set of combinatorial variants created by PCR fusion using several BPN' mutants as parent (template) molecules. The mutations introduced in each parent plasmid are shown in Table 11-7 and the mutagenic primers used to create the mutants are indicated in Table 11-8.
TABLE-US-00042 TABLE 11-7 List of Parent Plasmids and Mutations Introduced in the RCL5 Variants Combinatorial Mutations Variant # Parent Plasmids Introduced 1 G97A-G128A-Y217Q-T22N-S24A N61P-N62S 2 G97A-G128A-Y217Q-T22N-S24A T55P 3 G97A-G128A-Y217Q-T22N-S24A N61P-S63H 4 G97A-G128A-Y217Q-T22N-S24A Q59S-N61P 5 G97A-G128A-Y217Q-T22N-S24A L75S-N76Y 6 G97A-G128A-Y217Q-T22N-S24A P86S-S87G-A88V 7 G97A-G128A-Y217Q-T22N-S24A S87G-A88V-S89A 8 G97A-G128A-Y217Q-T22N-S24A S87T-A88L-S89G 9 G97A-G128A-Y217Q-T22N-S24A P129Q-S130G-G131S 10 G97A-G128A-Y217Q-T22N-S24A V203Y 11 G97A-G128A-Y217Q-T22N-S24A G211R-N212S-K213V 12 G97A-G128A-Y217Q-S24G-N25G N61P-N62S 13 G97A-G128A-Y217Q-S24G-N25G T55P 14 G97A-G128A-Y217Q-S24G-N25G N61P-S63H 15 G97A-G128A-Y217Q-S24G-N25G Q59S-N61P 16 G97A-G128A-Y217Q-S24G-N25G L75S-N76Y 17 G97A-G128A-Y217Q-S24G-N25G P86S-S87G-A88V 18 G97A-G128A-Y217Q-S24G-N25G S87G-A88V-S89A 19 G97A-G128A-Y217Q-S24G-N25G S87T-A88L-S89G 20 G97A-G128A-Y217Q-S24G-N25G P129Q-S130G-G131S 21 G97A-G128A-Y217Q-S24G-N25G V203Y 22 G97A-G128A-Y217Q-S24G-N25G G211R-N212S-K213V 23 G97A-G128A-Y217Q-S24R N61P-N62S 24 G97A-G128A-Y217Q-S24R T55P 25 G97A-G128A-Y217Q-S24R N61P-S63H 26 G97A-G128A-Y217Q-S24R Q59S-N61P 27 G97A-G128A-Y217Q-S24R L75S-N76Y 28 G97A-G128A-Y217Q-S24R P86S-S87G-A88V 29 G97A-G128A-Y217Q-S24R S87G-A88V-S89A 30 G97A-G128A-Y217Q-S24R S87T-A88L-S89G 31 G97A-G128A-Y217Q-S24R P129Q-S130G-G131S 32 G97A-G128A-Y217Q-S24R V203Y 33 G97A-G128A-Y217Q-S24R G211R-N212S-K213V 34 G97A-G128A-Y217Q-G23A-S24G-N25G N61P-N62S 35 G97A-G128A-Y217Q-G23A-S24G-N25G T55P 36 G97A-G128A-Y217Q-G23A-S24G-N25G N61P-S63H 37 G97A-G128A-Y217Q-G23A-S24G-N25G Q59S-N61P 38 G97A-G128A-Y217Q-G23A-S24G-N25G L75S-N76Y 39 G97A-G128A-Y217Q-G23A-S24G-N25G P86S-S87G-A88V 40 G97A-G128A-Y217Q-G23A-S24G-N25G S87G-A88V-S89A 41 G97A-G128A-Y217Q-G23A-S24G-N25G S87T-A88L-S89G 42 G97A-G128A-Y217Q-G23A-S24G-N25G P129Q-S130G-G131S 43 G97A-G128A-Y217Q-G23A-S24G-N25G V203Y 44 G97A-G128A-Y217Q-G23A-S24G-N25G G211R-N212S-K213V 45 G97A-G128A-Y217Q-N61P-N62S L75S-N76Y 46 G97A-G128A-Y217Q-N61P-N62S P86S-S87G-A88V 47 G97A-G128A-Y217Q-N61P-N62S S87G-A88V-S89A 48 G97A-G128A-Y217Q-N61P-N62S S87T-A88L-S89G 49 G97A-G128A-Y217Q-N61P-N62S P129Q-S130G-G131S 50 G97A-G128A-Y217Q-N61P-N62S V203Y 51 G97A-G128A-Y217Q-N61P-N62S G211R-N212S-K213V 52 G97A-G128A-Y217Q-T55P L75S-N76Y 53 G97A-G128A-Y217Q-T55P P86S-S87G-A88V 54 G97A-G128A-Y217Q-T55P S87G-A88V-S89A 55 G97A-G128A-Y217Q-T55P S87T-A88L-S89G 56 G97A-G128A-Y217Q-T55P P129Q-S130G-G131S 57 G97A-G128A-Y217Q-T55P V203Y 58 G97A-G128A-Y217Q-T55P G211R-N212S-K213V 59 G97A-G128A-Y217Q-N61P-S63H L75S-N76Y 60 G97A-G128A-Y217Q-N61P-S63H P86S-S87G-A88V 61 G97A-G128A-Y217Q-N61P-S63H S87G-A88V-S89A 62 G97A-G128A-Y217Q-N61P-S63H S87T-A88L-S89G 63 G97A-G128A-Y217Q-N61P-S63H P129Q-S130G-G131S 64 G97A-G128A-Y217Q-N61P-S63H V203Y 65 G97A-G128A-Y217Q-N61P-S63H G211R-N212S-K213V 66 G97A-G128A-Y217Q-Q59S-N61P L75S-N76Y 67 G97A-G128A-Y217Q-Q59S-N61P P86S-S87G-A88V 68 G97A-G128A-Y217Q-Q59S-N61P S87G-A88V-S89A 69 G97A-G128A-Y217Q-Q59S-N61P S87T-A88L-S89G 70 G97A-G128A-Y217Q-Q59S-N61P P129Q-S130G-G131S 71 G97A-G128A-Y217Q-Q59S-N61P V203Y 72 G97A-G128A-Y217Q-Q59S-N61P G211R-N212S-K213V 73 G97A-G128A-Y217Q-L75S-N76Y P86S-S87G-A88V 74 G97A-G128A-Y217Q-L75S-N76Y S87G-A88V-S89A 75 G97A-G128A-Y217Q-L75S-N76Y S87T-A88L-S89G 76 G97A-G128A-Y217Q-L75S-N76Y P129Q-S130G-G131S 77 G97A-G128A-Y217Q-L75S-N76Y V203Y 78 G97A-G128A-Y217Q-L75S-N76Y G211R-N212S-K213V 79 G97A-G128A-Y217Q-P86S-S87G-A88V P129Q-S130G-G131S 80 G97A-G128A-Y217Q-P86S-S87G-A88V V203Y 81 G97A-G128A-Y217Q-P86S-S87G-A88V G211R-N212S-K213V 82 G97A-G128A-Y217Q-S87G-A88V-S89A P129Q-S130G-G131S 83 G97A-G128A-Y217Q-S87G-A88V-S89A V203Y 84 G97A-G128A-Y217Q-S87G-A88V-S89A G211R-N212S-K213V 85 G97A-G128A-Y217Q-S87T-A88L-S89G P129Q-S130G-G131S 86 G97A-G128A-Y217Q-S87T-A88L-S89G V203Y 87 G97A-G128A-Y217Q-S87T-A88L-S89G G211R-N212S-K213V 88 G97A-G128A-Y217Q-P129Q-S130G-G131S V203Y 89 G97A-G128A-Y217Q-P129Q-S130G-G131S G211R-N212S-K213V 90 G97A-G128A-Y217Q-V203Y G211R-N212S-K213V
TABLE-US-00043 TABLE 11-8 Primers Used to Create RCL5 Combinatorial Variants Common 3' & 5' Gene Flanking SEQ Mutations Gene Primer Primer ID Introduced Fragments Names Name Primer Sequence NO: T22N- 3' P4976 P5432 TCAAGGCTACAATGGAGCAAATG 535 S24A TTAAAGTAGCGGTTATCGA 5' P4974 P5433 TTTAACATTTGCTCCATTGTAGCC 536 TTGAGAGTGCAGAG S24G- 3' P4976 P5434 CTACACTGGAGGAGGTGTTAAAG 537 N25G TAGCGGTTATCGACA 5' P4974 P5435 CTACTTTAACACCTCCTCCAGTGT 538 AGCCTTGAGAGTG S24R 3' P4976 P5436 AGGCTACACTGGAAGAAATGTTA 539 AAGTAGCGGTTATCGAC 5' P4974 P5437 CTTTAACATTTCTTCCAGTGTAGC 540 CTTGAGAGTG G23A- 3' P4976 P5438 AAGGCTACACTGCAGGAGGTGTT 541 S24G- AAAGTAGCGGTTATCGACA N25G 5' P4974 P5439 CTACTTTAACACCTCCTGCAGTGT 542 AGCCTTGAGAGTGCAG N61P- 3' P4976 P5440 CGTTTCAAGATCCCTCTTCTCATG 543 N62S GCACACACGTCGC 5' P4974 P5441 TGTGCCATGAGAAGAGGGATCTT 544 GAAACGGGTTTGTTTCG T55P 3' P4976 P5442 TGCCGTCCGAACCAAACCCGTTT 545 CAAGATAACAATTCT 5' P4974 P5443 TCTTGAAACGGGTTTGGTTCGGA 546 CGGCACCATAGAAG N61P- 3' P4976 P5444 CCGTTTCAAGATCCCAATCATCA 547 S63H TGGCACACACGTCGCAG 5' P4974 P5445 TGTGTGCCATGATGATTGGGATC 548 TTGAAACGGGTTTGTTTCG Q59S- 3' P4976 P5446 ACAAACCCGTTTTCAGATCCCAA 549 N61P TTCTCATGGCACACACGTCGCA 5' P4974 P5447 CCATGAGAATTGGGATCTGAAAA 550 CGGGTTTGTTTCGGACGGCA L75S- 3' P4976 P5448 GGTTGCGGCGTCATACAATTCTA 551 N76Y TTGGCGTGCTTGGTG 5' P4974 P5449 GCCAATAGAATTGTATGACGCCG 552 CAACCGTTCCTGCGA P86S- 3' P4976 P5450 TGGTGTAGCCTCGGGTGTTTCGCT 553 S87G- CTACGCCGTTAAAGTT A88V 5' P4974 P5451 CGTAGAGCGAAACACCCGAGGCT 554 ACACCAAGCACGCCAA S87G- 3' P4976 P5452 GTGTAGCCCCGGGTGTTGCACTC 555 A88V- TACGCCGTTAAAGTTCTTG S89A 5' P4974 P5453 ACGGCGTAGAGTGCAACACCCGG 556 GGCTACACCAAGCACGCCAA S87T- 3' P4976 P5454 TGGTGTAGCCCCGACTCTTGGAC 557 A88L- TCTACGCCGTTAAAGTTCTTG S89G 5' P4974 P5455 ACGGCGTAGAGTCCAAGAGTCGG 558 GGCTACACCAAGCACGCCAA P129Q- 3' P4976 P5456 GCCTGGGAGCACAAGGCTCTAGT 559 S130G- GCGGCACTTAAAGCAGCA G131S 5' P4974 P5457 AGTGCCGCACTAGAGCCTTGTGC 560 TCCCAGGCTCATGTTGAT V203Y 3' P4976 P5458 ATGGCCCCTGGCTATTCTATTCAA 561 TCGACGCTTCCAG 5' P4974 P5459 TCGATTGAATAGAATAGCCAGGG 562 GCCATGACATCCA G211R- 3' P4976 P5460 TCGACGCTTCCAAGGTCCGTGTA 563 N212S- TGGTGCGCAAAACGGGACT K213V 5' P4974 P5461 TTGCGCACCATACACGGACCTTG 564 GAAGCGTCGATTGAATAGAAA
[0607] To create each mutant, two PCR reactions were carried out using either the common 3' gene-flanking primer (P4976, CCTCTCGGTTATGAGTTAGTTC; SEQ ID NO:61) and the mutagenic primer, or the common 5'gene-flanking primer (P4974, GCCTCACATTTGTGCCACCTA; SEQ ID NO:60) and mutagenic primer as shown for each library in Table 11-8. These PCR reactions generated two PCR fragments, one encoding the 5' half of the mutant BPN' gene (5' gene fragment) and the other encoding the 3' half of the mutant BPN' gene (3' gene fragment). Each PCR amplification reaction contained 30 pmol of each primer and 100 ng of the parent molecules listed in Table 11-7. Amplifications were carried out using Vent DNA polymerase (NEB). The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 40 sec. Following amplification, the 5' and 3' gene fragments were gel-purified by the QIAGEN.RTM. gel-band purification kit, mixed (50 ng of each fragment) and amplified by PCR once again using the primers P4973 (AAAGGATCCTAATCGGCGCTTTTC; SEQ ID NO:62) and P4950 (CTTGTCTCCAAGCTTAAAATAAAA; SEQ ID NO:63) to generate the full-length gene fragment. The PCR conditions were same as described above, except the extension phase, which was carried out at 72.degree. C. for 2 min. The full-length DNA fragment was gel-purified by the QIAGEN.RTM. gel-band purification kit, digested by the BamHI and HindIII restriction enzymes and ligated with the pHPLT-BPN' partial opt that also was digested with the same restriction enzymes. Ligation mixtures were amplified using rolling circle amplification by Illustra Templiphi kit according to the manufacturer's instructions (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. For this purpose, 1 .mu.l of the ligation mixture was mixed with 5 .mu.l of the sample buffer, heated to 95.degree. C. for 3 min and cooled on ice. Next, 5 .mu.l of the reaction buffer and 0.2 .mu.m of the enzyme were added to each tube, followed by incubation at 30.degree. C. for 10 hours. Products of the rolling circle amplification were diluted 100 times and used to transform B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C. Subsequently, the colonies with halos were inoculated in 120 .mu.m of LB media containing 10 .mu.g/mL neomycin.
RCL 6 Combinatorial Libraries
[0608] "RCL6" refers to a group of combinatorial libraries created by PCR fusion using several BPN' mutants as parent (template) molecules. A mixture of BPN' mutants were used as templates (parent molecules) in the construction of each of these libraries. The five different mixes of parent molecules used to create these libraries are provided in Table 11-9, and the mutations introduced in each library are listed in Table 11-10.
[0609] To create each mutant, two PCR reactions were carried out using either the common 3' gene-flanking primer (P4976, CCTCTCGGTTATGAGTTAGTTC; SEQ ID NO:61) and the mutagenic primer, or the common 5'gene-flanking primer (P4974, GCCTCACATTTGTGCCACCTA; SEQ ID NO:60) and mutagenic primer as shown for each library in Table 11-10. These PCR reactions generated two PCR fragments, one encoding the 5' half of the mutant BPN' gene (5' gene fragment) and the other encoding the 3' half of the mutant BPN' gene (3' gene fragment). Each PCR amplification reaction contained 30 pmol of each primer and 100 ng of the parent molecules listed in Table 11-9. Amplifications were carried out using Vent DNA polymerase (NEB). The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 40 sec. Following amplification, the 5' and 3' gene fragments were gel-purified by the QIAGEN.RTM. gel-band purification kit, mixed (50 ng of each fragment) and amplified by PCR once again using the primers P4973 (AAAGGATCCTAATCGGCGCTTTTC; SEQ ID NO:62) and P4950 (CTTGTCTCCAAGCTTAAAATAAAA; SEQ ID NO:63) to generate the full-length gene fragment. The PCR conditions were same as described above, except the extension phase, which was carried out at 72.degree. C. for 2 min. The full-length DNA fragment was gel-purified by the QIAGEN.RTM. gel-band purification kit, digested using BamHI and HindIII restriction enzymes and ligated with the pHPLT-BPN' partial opt vector that also was digested with the same restriction enzymes. Ligation mixtures were amplified using rolling circle amplification by Illustra Templiphi kit according to the manufacturer's instructions (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. For this purpose, 1 .mu.l of the ligation mixture was mixed with 5 .mu.l of the sample buffer, heated to 95.degree. C. for 3 min and cooled on ice. Next, 5 .mu.l of the reaction buffer and 0.2 .mu.m of the enzyme were added to each tube, followed by incubation at 30.degree. C. for 10 hours. Products of the rolling circle amplification were diluted 100 times and used to transform B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C. Subsequently, the colonies with halos were inoculated in 120 .mu.m of LB media containing 10 .mu.g/mL neomycin.
TABLE-US-00044 TABLE 11-9 Parent Molecules of BPN' Used to Create RCL6 Libraries Mixes of Parent Molecules Mix 1 Mix 2 Mix 3 Mix 4 Mix 5 G97A-G128A- G97A-G128A- G97A-G128A- G97A-G128A- G97A-G128A- Y217Q-S24G- Y217Q-T55P Y217Q-L75S- Y217Q-P86S- Y217Q-P129Q- N25G N76Y S87G-A88V S130G-G131S G97A-G128A- G97A-G128A- G97A-G128A- Y217Q-S24R Y217Q-N61P- Y217Q-S87G- S63H A88V-S89A G97A-G128A- G97A-G128A- G97A-G128A- Y217Q-G23A- Y217Q-Q59S- Y217Q-S87T- S24G-N25G N61P A88L-S89G
TABLE-US-00045 TABLE 11-10 Mutations Introduced in the RCL6 Libraries Common 5' & 3' Gene Flanking Mutagenic Mutations Gene Primer Primer SEQ ID Introduced Fragment Names Name Mutagenic Primer Sequence NO: V68C, 3' P4976 P5462 CATGGCACACACTGCGGA 565 A69G GGAACGGTTGCGGCGTTA AAC 5' P4974 P5463 GCAACCGTTCCTCCGCAG 566 TGTGTGCCATGAGAATTG TTA V72I, A73G, 3' P4976 P5464 GTCGCAGGAACGATTGGT 567 delA74, L75S TCAAACAATTCTATTGGC GTGCTTG 5' P4974 P5465 CAATAGAATTGTTTGAAC 568 CAATCGTTCCTGCGACGT GTGTGCCAT L75H, 3' P4976 P5466 AACGGTTGCGGCGCATGG 569 N76G AAATTCTATTGGCGTGCTT GGTG 5' P4974 P5467 CAATAGAATTTCCATGCG 570 CCGCAACCGTTCCTGCGA CGTGT L75R, N76G, 3' P4976 P5468 AACGGTTGCGGCGAGAGG 571 N77S AGGTTCTATTGGCGTGCTT GGTGTA 5' P4974 P5469 CACGCCAATAGAACCTCC 572 TCTCGCCGCAACCGTTCCT GCGACGTGT L75G, 3' P4976 P5470 AACGGTTGCGGCGGGAGG 573 N76G, CGGTTCTATTGGCGTGCTT N77G GGTGTA 5' P4974 P5471 CACGCCAATAGAACCGCC 574 TCCCGCCGCAACCGTTCCT GCGACGTGT A92G 3' P4976 P5472 TGCTTCGCTCTACGGCGTT 575 AAAGTTCTTGCAGCAGAC 5' P4974 P5473 CAAGAACTTTAACGCCGT 576 AGAGCGAAGCAGACGGG GCTA delV93, 3' P4976 P5474 TTCGCTCTACGCCTCATGT 577 K94S, TCTGCAGCAGACGGATCA V95C, L96S GGCCAA 5' P4974 P5475 ATCCGTCTGCTGCAGAAC 578 ATGAGGCGTAGAGCGAAG CAGACG V121I_I122S_N123C 3' P4976 P5476 AATAACATGGATATATCT 579 TGCATGAGCCTGGGAGCA CCAAG 5' P4974 P5477 CAGGCTCATGCAAGATAT 580 ATCCATGTTATTCGCGATG GCCCATT V121L_N123C 3' P4976 P5478 CGAATAACATGGATCTTA 581 TCTGCATGAGCCTGGGAG CACCAAG 5' P4974 P5479 CCAGGCTCATGCAGATAA 582 GATCCATGTTATTCGCGAT GGCCCATT I122C_N123S_M124L 3' P4976 P5480 TAACATGGATGTATGCTC 583 ATTGAGCCTGGGAGCACC AAGCGGCA 5' P4974 P5481 TGCTCCCAGGCTCAATGA 584 GCATACATCCATGTTATTC GCGATG N123C 3' P4976 P5482 ACATGGATGTAATCTGCA 585 TGAGCCTGGGAGCACCAAG 5' P4974 P5483 TCCCAGGCTCATGCAGAT 586 TACATCCATGTTATTCGCG AT M124I 3' P4976 P5484 GGATGTAATCAACATCAG 587 CCTGGGAGCACCAAGCGG CA 5' P4974 P5485 TGCTCCCAGGCTGATGTT 588 GATTACATCCATGTTATTCG M124V 3' P4976 P5486 GGATGTAATCAACGTAAG 589 CCTGGGAGCACCAAGCGG CA 5' P4974 P5487 TGCTCCCAGGCTTACGTTG 590 ATTACATCCATGTTATTCG M124V- 3' P4976 P5488 GGATGTAATCAACGTAAG 591 L126A CGCGGGAGCACCAAGCGG CAGTG 5' P4974 P5489 TTGGTGCTCCCGCGCTTAC 592 GTTGATTACATCCATGTTA TTCG L126F, 3' P4976 P5490 AATCAACATGAGCTTCGG 593 delP129 AGCAAGCGGCAGTGCGGC ACTTAA 5' P4974 P5491 CACTGCCGCTTGCTCCGA 594 AGCTCATGTTGATTACATC CATGT G127Y 3' P4976 P5492 AACATGAGCCTGTACGCA 595 CCAAGCGGCAGTGCGGCA CTTA 5' P4974 P5493 CACTGCCGCTTGGTGCGT 596 ACAGGCTCATGTTGATTA CATCC G127S_P129D 3' P4976 P5494 CAACATGAGCCTGTCAGC 597 AGATAGCGGCAGTGCGGC ACTTAAA 5' P4974 P5495 GCACTGCCGCTATCTGCT 598 GACAGGCTCATGTTGATT ACATCC G127N, P129R 3' P4976 P5496 CAACATGAGCCTGAACGC 599 ACGTAGCGGCAGTGCGGC ACTTAAA 5' P4974 P5497 GCACTGCCGCTACGTGCG 600 TTCAGGCTCATGTTGATTA CATCC G128N, 3' P4976 P5498 ATGAGCCTGGGAAATTCA 601 insS, TCTAGCGGCAGTGCGGCA P129S CTTAAA 5' P4974 P5499 GCACTGCCGCTAGATGAA 602 TTTCCCAGGCTCATGTTGA TTAC G128S_P129V 3' P4976 P5500 CATGAGCCTGGGATCAGT 603 TAGCGGCAGTGCGGCACT TAAA 5' P4974 P5501 GCACTGCCGCTAACTGAT 604 CCCAGGCTCATGTTGATT AC G128S, 3' P4976 P5502 CATGAGCCTGGGATCAGA 605 P129D TAGCGGCAGTGCGGCACT TAAA 5' P4974 P5503 GCACTGCCGCTATCTGAT 606 CCCAGGCTCATGTTGATT AC G128S, 3' P4976 P5504 CATGAGCCTGGGATCAGG 607 P129G TAGCGGCAGTGCGGCACT TAAA 5' P4974 P5505 GCACTGCCGCTACCTGAT 608 CCCAGGCTCATGTTGATT AC H128H, 3' P4976 P5506 CATGAGCCTGGGACACTA 609 P129Y TAGCGGCAGTGCGGCACT TAAA 5' P4974 P5507 GCACTGCCGCTATAGTGT 610 CCCAGGCTCATGTTGATT AC P129D 3' P4976 P5508 GAGCCTGGGAGCAGACAG 611 CGGCAGTGCGGCACTTAA 5' P4974 P5509 TGCCGCACTGCCGCTGTCT 612 GCTCCCAGGCTCATGTTG AT P129E 3' P4976 P5510 GAGCCTGGGAGCAGAAAG 613 CGGCAGTGCGGCACTTAA 5' P4974 P5511 TGCCGCACTGCCGCTTTCT 614 GCTCCCAGGCTCATGTTG AT P129V 3' P4976 P5512 GAGCCTGGGAGCAGTAAG 615 CGGCAGTGCGGCACTTAA 5' P4974 P5513 TGCCGCACTGCCGCTTACT 616 GCTCCCAGGCTCATGTTG AT P129G, 3' P4976 P5514 GAGCCTGGGAGCAGGAGG 617 delS130 CAGTGCGGCACTTAAAGC 5' P4974 P5515 AGTGCCGCACTGCCTCCT 618 GCTCCCAGGCTCATGTTG AT P129H, 3' P4976 P5516 AGCCTGGGAGCACACGGC 619 delS130, AATGCGGCACTTAAAGCA S132N GCAGTT 5' P4974 P5517 TTTAAGTGCCGCATTGCC 620 GTGTGCTCCCAGGCTCAT GTTGAT A134T 3' P4976 P5518 AAGCGGCAGTGCGACACT 621 TAAAGCAGCAGTTGATAA AG 5' P4974 P5519 AACTGCTGCTTTAAGTGTC 622 GCACTGCCGCTTGGTGCTC G97R, 3' P4976 P5520 GTTAAAGTTCTTCGTGGTT 623 insG, A98C GTGACGGATCAGGCCAAT ACTC 5' P4974 P5521 CTGATCCGTCACAACCAC 624 GAAGAACTTTAACGGCGT AGAGC A98G, 3' P4976 P5522 TTAAAGTTCTTGCAGGAG 625 D99G GCGGATCAGGCCAATACT CATG 5' P4974 P5523 TATTGGCCTGATCCGCCTC 626 CTGCAAGAACTTTAACGG CGTAG A98G, insR 3' P4976 P5524 TTAAAGTTCTTGCAGGAC 627 GTGACGGATCAGGCCAAT ACTCA 5' P4974 P5525 CTGATCCGTCACGTCCTGC 628 AAGAACTTTAACGGCGTAG A98D, 3' P4976 P5526 TTAAAGTTCTTGCAGACG 629 D99G GCGGATCAGGCCAATACT CATG 5' P4974 P5527 TATTGGCCTGATCCGCCGT 630 CTGCAAGAACTTTAACGG CGTAG A98H, 3' P4976 P5528 TAAAGTTCTTGCACATGG 631 D99G, AGATTCAGGCCAATACTC G100D ATGGATTAT 5' P4974 P5529 AGTATTGGCCTGAATCTC 632 CATGTGCAAGAACTTTAA CGGCGTAG D99R, insN 3' P4976 P5530 AAGTTCTTGCAGCACGTA 633 ACGGATCAGGCCAATACT CATG 5' P4974 P5531 TATTGGCCTGATCCGTTAC 634 GTGCTGCAAGAACTTTAA CGGCGTA D99V, 3' P4976 P5532 AGTTCTTGCAGCAGTAGG 635 S101D AGATGGCCAATACTCATG GATTATCAA 5' P4974 P5533 TGAGTATTGGCCATCTCCT 636 ACTGCTGCAAGAACTTTA ACGGCGTA
D99C, insS 3' P4976 P5534 TTAAAGTTCTTGCAGCAT 637 GTAGCGGATCAGGCCAAT ACTCATG 5' P4974 P5535 TATTGGCCTGATCCGCTAC 638 ATGCTGCAAGAACTTTAA CGGCGTA G100S 3' P4976 P5536 AAGTTCTTGCAGCAGACT 639 CTTCAGGCCAATACTCAT GGATTAT 5' P4974 P5537 ATGAGTATTGGCCTGAAG 640 AGTCTGCTGCAAGAACTT TAACG G100S, 3' P4976 P5538 TTCTTGCAGCAGACTCTGT 641 S101V AGGCCAATACTCATGGAT TATCA 5' P4974 P5539 CATGAGTATTGGCCTACA 642 GAGTCTGCTGCAAGAACT TTAACG G100D 3' P4976 P5540 AAGTTCTTGCAGCAGACG 643 ATTCAGGCCAATACTCAT GGATTAT 5' P4974 P5541 ATGAGTATTGGCCTGAAT 644 CGTCTGCTGCAAGAACTT TAACG G100N 3' P4976 P5542 AAGTTCTTGCAGCAGACA 645 ATTCAGGCCAATACTCAT GGATTAT 5' P4974 P5543 ATGAGTATTGGCCTGAAT 646 TGTCTGCTGCAAGAACTTT AACG S100N, 3' P4976 P5544 TTCTTGCAGCAGACAATC 647 S101L TAGGCCAATACTCATGGA TTATCA 5' P4974 P5545 CATGAGTATTGGCCTAGA 648 TTGTCTGCTGCAAGAACTT TAACG S101G 3' P4976 P5546 TTCTTGCAGCAGACGGAG 649 GAGGCCAATACTCATGGA TTATCAA 5' P4974 P5547 ATGAGTATTGGCCTCCTCC 650 GTCTGCTGCAAGAACTTTA S101D 3' P4976 P5548 TTCTTGCAGCAGACGGAG 651 ATGGCCAATACTCATGGA TTATCAA 5' P4974 P5549 ATGAGTATTGGCCATCTC 652 CGTCTGCTGCAAGAACTT TA S101V, 3' P4976 P5550 TGCAGCAGACGGAGTAGG 653 Q103N CAACTACTCATGGATTAT CAACGGCAT 5' P4974 P5551 ATAATCCATGAGTAGTTG 654 CCTACTCCGTCTGCTGCAA GAACTTTA S101E 3' P4976 P5552 TTCTTGCAGCAGACGGAC 655 GTGGCCAATACTCATGGA TTATCAA 5' P4974 P5553 ATGAGTATTGGCCACGTC 656 CGTCTGCTGCAAGAACTT TA A116S, N117G, 3' P4976 P5554 AATGGGCCATCTCTGGTA 657 N118R GAATGGATGTAATCAACA TGAGCCT 5' P4974 P5555 GATTACATCCATTCTACCA 658 GAGATGGCCCATTCGATG CCGTT A116G, N117R 3' P4976 P5556 AATGGGCCATCGGACGTA 659 ACATGGATGTAATCAACA TGAG 5' P4974 P5557 GATTACATCCATGTTACGT 660 CCGATGGCCCATTCGATG CCGTT A116N, N117S, 3' P4976 P5558 AATGGGCCATCAATTCTG 661 N118G GAATGGATGTAATCAACA TGAGCCT 5' P4974 P5559 GATTACATCCATTCCAGA 662 ATTGATGGCCCATTCGAT GCCGTT M222Q 3' P4976 P5560 AAAACGGGACTTCCCAGG 663 CCTCGCCGCATGTAGCTG 5' P4974 P5561 TACATGCGGCGAGGCCTG 664 GGAAGTCCCGTTTTGCGC AC S24R 3' P4976 P5562 AAGGCTACACTGGAAGAA 665 ATGTTAAAGTAGCGGTTA TCGA 5' P4974 P5563 CTACTTTAACATTTCTTCC 666 AGTGTAGCCTTGAGAGTG N25Y 3' P4976 P5564 CTACACTGGATCATATGTT 667 AAAGTAGCGGTTATCGACA 5' P4974 P5565 TAACCGCTACTTTAACAT 668 ATGATCCAGTGTAGCCTT GAGA P52D 3' P4976 P5566 CTTCTATGGTGGATTCCGA 669 AACAAACCCGTTTCAAG 5' P4974 P5567 GGTTTGTTTCGGAATCCAC 670 CATAGAAGCCCCTCCAG S63T 3' P4976 P5568 TTTCAAGATAACAATACA 671 CATGGCACACACGTCGCA GGA 5' P4974 P5569 TGTGTGCCATGTGTATTGT 672 TATCTTGAAACGGGTTTGT N61E 3' P4976 P5570 GTTTCAAGATGAAAATTC 673 TCATGGCACACACGTC 5' P4974 P5571 TGTGCCATGAGAATTTTC 674 ATCTTGAAACGGGTTTGTT TCG N61P 3' P4976 P5572 AACCCGTTTCAAGATCCA 675 AATTCTCATGGCACACAC GTC 5' P4974 P5573 TGCCATGAGAATTTGGAT 676 CTTGAAACGGGTTTGTTTCG N62Q 3' P4976 P5574 GTTTCAAGATAACCAATC 677 TCATGGCACACACGTCGC AGGAA 5' P4974 P5575 TGTGTGCCATGAGATTGG 678 TTATCTTGAAACGGGTTTG TTT N62D 3' P4976 P5576 GTTTCAAGATAACGATTC 679 TCATGGCACACACGTCGC AGGAA 5' P4974 P5577 TGTGTGCCATGAGAATCG 680 TTATCTTGAAACGGGTTTG TTT S63Q 3' P4976 P5578 TCAAGATAACAATCAACA 681 TGGCACACACGTCGCAGG 5' P4974 P5579 ACGTGTGTGCCATGTTGA 682 TTGTTATCTTGAAACGGGT TTG V68A 3' P4976 P5580 TCATGGCACACACGCAGC 683 AGGAACGGTTGCGGCGTT AA 5' P4974 P5581 CAACCGTTCCTGCTGCGT 684 GTGTGCCATGAGAATTGT TA S87D 3' P4976 P5582 TGTAGCCCCGGATGCTTC 685 GCTCTACGCCGTTAA 5' P4974 P5583 CGTAGAGCGAAGCATCCG 686 GGGCTACACCAAGCACG L96T 3' P4976 P5584 CGTTAAAGTTACAGCAGC 687 AGACGGATCAGGCCAATA 5' P4974 P5585 TGATCCGTCTGCTGCTGTA 688 ACTTTAACGGCGTAGAGC GAA L126A 3' P4976 P5586 TAATCAACATGAGCGCGG 689 GAGCACCAAGCGGCAGTG 5' P4974 P5587 TTGGTGCTCCCGCGCTCAT 690 GTTGATTACATCCATG L126T 3' P4976 P5588 TAATCAACATGAGCACGG 691 GAGCACCAAGCGGCAGTG 5' P4974 P5589 TTGGTGCTCCCGTGCTCAT 692 GTTGATTACATCCATG S125A 3' P4976 P5590 ATGTAATCAACATGGCAC 693 TGGGAGCACCAAGCGGCA GT 5' P4974 P5591 TTGGTGCTCCCAGTGCCAT 694 GTTGATTACATCCATGTTA TT S130P 3' P4976 P5592 TGGGAGCACCACCAGGCA 695 GTGCGGCACTTAAAGC 5' P4974 P5593 GTGCCGCACTGCCTGGTG 696 GTGCTCCCAGGCTCATGT P129L 3' P4976 P5594 TGAGCCTGGGAGCACTTA 697 GCGGCAGTGCGGCACTTAA 5' P4974 P5595 TGCCGCACTGCCGCTAAG 698 TGCTCCCAGGCTCATGTTG AT P129E 3' P4976 P5596 TGAGCCTGGGAGCAGAAA 699 GCGGCAGTGCGGCACTTAA 5' P4974 P5597 TGCCGCACTGCCGCTTTCT 700 GCTCCCAGGCTCATGTTG AT P129S 3' P4976 P5598 TGAGCCTGGGAGCATCTA 701 GCGGCAGTGCGGCACTTAA 5' P4974 P5599 TGCCGCACTGCCGCTAGA 702 TGCTCCCAGGCTCATGTTG AT P40E 3' P4976 P5600 GACTCGAGCCATGAAGAT 703 CTTAAAGTCGCTGGAGG 5' P4974 P5601 GACTTTAAGATCTTCATG 704 GCTCGAGTCGATACCGCT Y6Q 3' P4976 P5602 GCAGTCCGTGCCTCAAGG 705 CGTATCACAAATTAAAGC CCCT 5' P4974 P5603 ATTTGTGATACGCCTTGA 706 GGCACGGACTGCGCGTAC GCAT G102A 3' P4976 P5604 CAGACGGATCAGCACAAT 707 ACTCATGGATTATCAACG GCAT 5' P4974 P5605 TAATCCATGAGTATTGTG 708 CTGATCCGTCTGCTGCAA GAAC S101N 3' P4976 P5606 GCAGCAGACGGAAACGGC 709 CAATACTCATGGATTATC AA 5' P4974 P5607 CATGAGTATTGGCCGTTTC 710 CGTCTGCTGCAAGAACTT TA G100E 3' P4976 P5608 TTCTTGCAGCAGACGAAT 711 CAGGCCAATACTCATGGA TTAT 5' P4974 P5609 TGAGTATTGGCCTGATTC 712 GTCTGCTGCAAGAACTTT AACG I115V 3' P4976 P5610 ATCGAATGGGCCGTAGCG 713 AATAACATGGATGTAATC AA 5' P4974 P5611 CATCCATGTTATTCGCTAC 714 GGCCCATTCGATGCCGTT GAT A144K 3' P4976 P5612 GTTGATAAAGCTGTTAAA 715
TCTGGTGTCGTCGTAGTA GC 5' P4974 P5613 GACGACACCAGATTTAAC 716 AGCTTTATCAACTGCTGCTT S145D 3' P4976 P5614 GATAAAGCTGTTGCAGAT 717 GGTGTCGTCGTAGTAGCG GCA 5' P4974 P5615 TACTACGACGACACCATC 718 TGCAACAGCTTTATCAAC TGCT S159K 3' P4976 P5616 AATGAGGGAACAAAAGG 719 ATCATCGAGTACCGTCGG TTA 5' P4974 P5617 ACGGTACTCGATGATCCT 720 TTTGTTCCCTCATTCCCAG CTG S162K 3' P4976 P5618 AACATCCGGATCAAAAAG 721 TACCGTCGGTTATCCAGG CAA 5' P4974 P5619 ATAACCGACGGTACTTTTT 722 GATCCGGATGTTCCCTCATT V147P 3' P4976 P5620 TGTTGCATCTGGTCCAGTC 723 GTAGTAGCGGCAGCTGGG AAT 5' P4974 P5621 TGCCGCTACTACGACTGG 724 ACCAGATGCAACAGCTTT ATCA S161P 3' P4976 P5622 AGGGAACATCCGGACCAT 725 CGAGTACCGTCGGTTATC CA 5' P4974 P5623 ACCGACGGTACTCGATGG 726 TCCGGATGTTCCCTCATTC CCA A187D 3' P4976 P5624 CTTCAAATCAACGTGACT 727 CTTTTTCCTCCGTGGGACC GGA 5' P4974 P5625 ACGGAGGAAAAAGAGTC 728 ACGTTGATTTGAAGAGTC TACAG F189D 3' P4976 P5626 TCAACGTGCCTCTGATTCC 729 TCCGTGGGACCGGAGCTG GAT 5' P4974 P5627 TCCCACGGAGGAATCAGA 730 GGCACGTTGATTTGAAGAG L267V 3' P4976 P5628 TACTATGGAAAAGGGGTA 731 ATCAACGTACAGGCGGCA GC 5' P4974 P5629 CTGTACGTTGATTACCCCT 732 TTTCCATAGTAGAAAGAAT Q206E 3' P4976 P5630 TGGCGTTTCTATTGAATCG 733 ACGCTTCCAGGGAACAA 5' P4974 P5631 CTGGAAGCGTCGATTCAA 734 TAGAAACGCCAGGGGCCAT K213T 3' P4976 P5632 CTTCCAGGGAACACATAT 735 GGTGCGCAAAACGGGACT 5' P4974 P5633 GTTTTGCGCACCATATGTG 736 TTCCCTGGAAGCGTCGATT K213L 3' P4976 P5634 CTTCCAGGGAACCTTTAT 737 GGTGCGCAAAACGGGACT 5' P4974 P5635 GTTTTGCGCACCATAAAG 738 GTTCCCTGGAAGCGTCGA TT K265N 3' P4976 P5636 TTTCTACTATGGAAACGG 739 GCTGATCAACGTACAGGC GGCA 5' P4974 P5637 ACGTTGATCAGCCCGTTTC 740 CATAGTAGAAAGAATCAC CAA N240K 3' P4976 P5638 TTTCTAAGCACCCGAAAT 741 GGACAAACACTCAAGTCC GCA 5' P4974 P5639 GAGTGTTTGTCCATTTCGG 742 GTGCTTAGAAAGAATCAAT P239R 3' P4976 P5640 TTCTTTCTAAGCACCGTAA 743 CTGGACAAACACTCAAGT CC 5' P4974 P5641 TGTTTGTCCAGTTACGGTG 744 CTTAGAAAGAATCAATGCG T242R 3' P4976 P5642 CACCCGAACTGGCGTAAC 745 ACTCAAGTCCGCAGCAGT 5' P4974 P5643 TGCGGACTTGAGTGTTAC 746 GCCAGTTCGGGTGCTTAG AAAG S89Y 3' P4976 P5644 CGTCTGCTTACCTCTACGC 747 CGTTAAAGTTCTTG 5' P4974 P5645 ACTTTAACGGCGTAGAGG 748 TAAGCAGACGGGGCTACA CCAA P129Q 3' P4976 P5646 AGCCTGGGAGCACAAAGC 749 GGCAGTGCGGCACTTAAA 5' P4974 P5647 CACTGCCGCTTTGTGCTCC 750 CAGGCTCATGTTGAT G211T 3' P4976 P5648 TTCAATCGACGCTTCCAA 751 CGAACAAGTATGGTGCGC AAAAC 5' P4974 P5649 CACCATACTTGTTCGTTGG 752 AAGCGTCGATTGAATAGA AA I111V 3' P4976 P5650 TGGATTATCAACGGCGTA 753 GAATGGGCCATCGCGAAT AAC 5' P4974 P5651 CGATGGCCCATTCTACGC 754 CGTTGATAATCCATGAGT ATT
RCL 7 Combinatorial Variants
[0610] "RCL7" refers to a set of combinatorial variants created by PCR fusion using several BPN' mutants as parent (template) plasmid. The mutations introduced in each parent plasmid are listed in Table 11-11, and the mutagenic primers used to create the mutants are described in Table 11-10.
[0611] To create each mutant, two PCR reactions were carried out using either the common 3' gene-flanking primer (P4976, CCTCTCGGTTATGAGTTAGTTC; SEQ ID NO:61) and the mutagenic primer, or the common 5'gene-flanking primer (P4974, GCCTCACATTTGTGCCACCTA; SEQ ID NO:60) and mutagenic primer as shown for each library in Table 11-10. These PCR reactions generated two PCR fragments, one encoding the 5' half of the mutant BPN' gene (5' gene fragment) and the other encoding the 3' half of the mutant BPN' gene (3' gene fragment). Each PCR amplification reaction contained 30 pmol of each primer and 100 ng of the parent molecules listed in Table 11-11. Amplifications were carried out using Vent DNA polymerase (NEB). The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 40 sec. Following amplification, the 5' and 3' gene fragments were gel-purified using a QIAGEN.RTM. gel-band purification kit, mixed (50 ng of each fragment) and amplified by PCR once again using the primers P4973 (AAAGGATCCTAATCGGCGCTTTTC; SEQ ID NO:62) and P4950 (CTTGTCTCCAAGCTTAAAATAAAA; SEQ ID NO:63) to generate the full-length gene fragment. The PCR conditions were same as described above, except the extension phase, which was carried out at 72.degree. C. for 2 min. The full-length DNA fragment was gel-purified using a QIAGEN.RTM. gel-band purification kit, digested using BamHI and HindIII restriction enzymes, and ligated with the pHPLT-BPN' partial opt that also was digested with the same restriction enzymes. Ligation mixtures were amplified using rolling circle amplification by Illustra Templiphi kit according to the manufacturer's instructions (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. For this purpose, 1 .mu.m of the ligation mixture was mixed with 5 .mu.l of the sample buffer, heated to 95.degree. C. for 3 min and cooled on ice. Next, 5 .mu.l of the reaction buffer and 0.2 .mu.m of the enzyme were added to each tube, followed by incubation at 30.degree. C. for 10 hours. Products of the rolling circle amplification were diluted 100 times and used to transform B. subtilis cells (AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C. Subsequently, the colonies with halos were inoculated in 120 .mu.m of LB media containing 10 .mu.g/mL neomycin.
TABLE-US-00046 TABLE 11-11 List of Parent Plasmids and Introduced Mutations in the RCL7 variants Combinatorial Mutation(s) Variant # Parent Plasmid Introduced 1 G97A-G128A-Y217Q-T55P S24R 2 G97A-G128A-Y217Q-N61E S24R 3 G97A-G128A-Y217Q-N61P-S63H S24R 4 G97A-G128A-Y217Q-L75S-N76Y S24R 5 G97A-G128A-Y217Q-S87T-A88L-S89G S24R 6 G97A-G128A-Y217Q-S89Y S24R 7 G97A-G128A-Y217Q-I111V S24R 8 G97A-G128A-Y217Q-I115V S24R 9 G97A-G128A-Y217Q-P129Q-S130G-G131S S24R 10 G97A-G128A-Y217Q-P129Q S24R 11 G97A-G128A-Y217Q-A134T S24R 12 G97A-G128A-Y217Q-A144K S24R 13 G97A-G128A-Y217Q-S145D S24R 14 G97A-G128A-Y217Q-S159K S24R 15 G97A-G128A-Y217Q-S162K S24R 16 G97A-G128A-Y217Q-S161P-S78N S24R 17 G97A-G128A-Y217Q-V203Y S24R 18 G97A-G128A-Y217Q-G211T-S78N S24R 19 G97A-G128A-Y217Q-K213T S24R 20 G97A-G128A-Y217Q-P239R S24R 21 G97A-G128A-Y217Q-N240K S24R 22 G97A-G128A-Y217Q-L267V-S78N S24R 23 G97A-G128A-Y217Q-A273S-S78N S24R 24 G97A-G128A-Y217Q-L75S-N76Y T55P 25 G97A-G128A-Y217Q-S87T-A88L-S89G T55P 26 G97A-G128A-Y217Q-S89Y T55P 27 G97A-G128A-Y217Q-I111V T55P 28 G97A-G128A-Y217Q-I115V T55P 29 G97A-G128A-Y217Q-P129Q-S130G-G131S T55P 30 G97A-G128A-Y217Q-P129Q T55P 31 G97A-G128A-Y217Q-A134T T55P 32 G97A-G128A-Y217Q-A144K T55P 33 G97A-G128A-Y217Q-S145D T55P 34 G97A-G128A-Y217Q-S159K T55P 35 G97A-G128A-Y217Q-S162K T55P 36 G97A-G128A-Y217Q-S161P T55P 37 G97A-G128A-Y217Q-V203Y T55P 38 G97A-G128A-Y217Q-G211T T55P 39 G97A-G128A-Y217Q-K213T T55P 40 G97A-G128A-Y217Q-P239R T55P 41 G97A-G128A-Y217Q-N240K T55P 42 G97A-G128A-Y217Q-L267V T55P 43 G97A-G128A-Y217Q-A273S T55P 44 G97A-G128A-Y217Q-L75S-N76Y N61E 45 G97A-G128A-Y217Q-S87T-A88L-S89G N61E 46 G97A-G128A-Y217Q-S89Y N61E 47 G97A-G128A-Y217Q-I111V N61E 48 G97A-G128A-Y217Q-I115V N61E 49 G97A-G128A-Y217Q-P129Q-S130G-G131S N61E 50 G97A-G128A-Y217Q-P129Q N61E 51 G97A-G128A-Y217Q-A134T N61E 52 G97A-G128A-Y217Q-A144K N61E 53 G97A-G128A-Y217Q-S145D N61E 54 G97A-G128A-Y217Q-S159K N61E 55 G97A-G128A-Y217Q-S162K N61E 56 G97A-G128A-Y217Q-S161P N61E 57 G97A-G128A-Y217Q-V203Y N61E 58 G97A-G128A-Y217Q-G211T N61E 59 G97A-G128A-Y217Q-K213T N61E 60 G97A-G128A-Y217Q-P239R N61E 61 G97A-G128A-Y217Q-N240K N61E 62 G97A-G128A-Y217Q-L267V N61E 63 G97A-G128A-Y217Q-A273S N61E 64 G97A-G128A-Y217Q-L75S-N76Y N61P-S63H 65 G97A-G128A-Y217Q-S87T-A88L-S89G N61P-S63H 66 G97A-G128A-Y217Q-S89Y N61P-S63H 67 G97A-G128A-Y217Q-I111V N61P-S63H 68 G97A-G128A-Y217Q-I115V N61P-S63H 69 G97A-G128A-Y217Q-P129Q-S130G-G131S N61P-S63H 70 G97A-G128A-Y217Q-P129Q N61P-S63H 71 G97A-G128A-Y217Q-A134T N61P-S63H 72 G97A-G128A-Y217Q-A144K N61P-S63H 73 G97A-G128A-Y217Q-S145D N61P-S63H 74 G97A-G128A-Y217Q-S159K N61P-S63H 75 G97A-G128A-Y217Q-S162K N61P-S63H 76 G97A-G128A-Y217Q-S161P N61P-S63H 77 G97A-G128A-Y217Q-V203Y N61P-S63H 78 G97A-G128A-Y217Q-G211T N61P-S63H 79 G97A-G128A-Y217Q-K213T N61P-S63H 80 G97A-G128A-Y217Q-P239R N61P-S63H 81 G97A-G128A-Y217Q-N240K N61P-S63H 82 G97A-G128A-Y217Q-L267V N61P-S63H 83 G97A-G128A-Y217Q-A273S N61P-S63H 84 G97A-G128A-Y217Q-S87T-A88L-S89G L75S-N76Y 85 G97A-G128A-Y217Q-S89Y L75S-N76Y 86 G97A-G128A-Y217Q-I111V L75S-N76Y 87 G97A-G128A-Y217Q-I115V L75S-N76Y 88 G97A-G128A-Y217Q-P129Q-S130G-G131S L75S-N76Y 89 G97A-G128A-Y217Q-P129Q L75S-N76Y 90 G97A-G128A-Y217Q-A134T L75S-N76Y 91 G97A-G128A-Y217Q-A144K L75S-N76Y 92 G97A-G128A-Y217Q-S145D L75S-N76Y 93 G97A-G128A-Y217Q-S159K L75S-N76Y 94 G97A-G128A-Y217Q-S162K L75S-N76Y 95 G97A-G128A-Y217Q-S161P L75S-N76Y 96 G97A-G128A-Y217Q-V203Y L75S-N76Y 97 G97A-G128A-Y217Q-G211T L75S-N76Y 98 G97A-G128A-Y217Q-K213T L75S-N76Y 99 G97A-G128A-Y217Q-P239R L75S-N76Y 100 G97A-G128A-Y217Q-N240K L75S-N76Y 101 G97A-G128A-Y217Q-L267V L75S-N76Y 102 G97A-G128A-Y217Q-A273S L75S-N76Y 103 G97A-G128A-Y217Q-I111V S87T-A88L-S89G 104 G97A-G128A-Y217Q-I115V S87T-A88L-S89G 105 G97A-G128A-Y217Q-P129Q-S130G-G131S S87T-A88L-S89G 106 G97A-G128A-Y217Q-P129Q S87T-A88L-S89G 107 G97A-G128A-Y217Q-A134T S87T-A88L-S89G 108 G97A-G128A-Y217Q-A144K S87T-A88L-S89G 109 G97A-G128A-Y217Q-S145D S87T-A88L-S89G 110 G97A-G128A-Y217Q-S159K S87T-A88L-S89G 111 G97A-G128A-Y217Q-S162K S87T-A88L-S89G 112 G97A-G128A-Y217Q-S161P S87T-A88L-S89G 113 G97A-G128A-Y217Q-V203Y S87T-A88L-S89G 114 G97A-G128A-Y217Q-G211T S87T-A88L-S89G 115 G97A-G128A-Y217Q-K213T S87T-A88L-S89G 116 G97A-G128A-Y217Q-P239R S87T-A88L-S89G 117 G97A-G128A-Y217Q-N240K S87T-A88L-S89G 118 G97A-G128A-Y217Q-L267V S87T-A88L-S89G 119 G97A-G128A-Y217Q-A273S S87T-A88L-S89G 120 G97A-G128A-Y217Q-I111V S89Y 121 G97A-G128A-Y217Q-I115V S89Y 122 G97A-G128A-Y217Q-P129Q-S130G-G131S S89Y 123 G97A-G128A-Y217Q-P129Q S89Y 124 G97A-G128A-Y217Q-A134T S89Y 125 G97A-G128A-Y217Q-A144K S89Y 126 G97A-G128A-Y217Q-S145D S89Y 127 G97A-G128A-Y217Q-S159K S89Y 128 G97A-G128A-Y217Q-S162K S89Y 129 G97A-G128A-Y217Q-S161P S89Y 130 G97A-G128A-Y217Q-V203Y S89Y 131 G97A-G128A-Y217Q-G211T S89Y 132 G97A-G128A-Y217Q-K213T S89Y 133 G97A-G128A-Y217Q-P239R S89Y 134 G97A-G128A-Y217Q-N240K S89Y 135 G97A-G128A-Y217Q-L267V S89Y 136 G97A-G128A-Y217Q-A273S S89Y 137 G97A-G128A-Y217Q-P129Q-S130G-G131S I111V 138 G97A-G128A-Y217Q-P129Q I111V 139 G97A-G128A-Y217Q-A134T I111V 140 G97A-G128A-Y217Q-A144K I111V 141 G97A-G128A-Y217Q-S145D I111V 142 G97A-G128A-Y217Q-S159K I111V 143 G97A-G128A-Y217Q-S162K I111V 144 G97A-G128A-Y217Q-S161P I111V 145 G97A-G128A-Y217Q-V203Y I111V 146 G97A-G128A-Y217Q-G211T I111V 147 G97A-G128A-Y217Q-K213T I111V 148 G97A-G128A-Y217Q-P239R I111V 149 G97A-G128A-Y217Q-N240K I111V 150 G97A-G128A-Y217Q-L267V I111V 151 G97A-G128A-Y217Q-A273S I111V 152 G97A-G128A-Y217Q-P129Q-S130G-G131S I115V 153 G97A-G128A-Y217Q-P129Q I115V 154 G97A-G128A-Y217Q-A134T I115V 155 G97A-G128A-Y217Q-A144K I115V 156 G97A-G128A-Y217Q-S145D I115V 157 G97A-G128A-Y217Q-S159K I115V 158 G97A-G128A-Y217Q-S162K I115V 159 G97A-G128A-Y217Q-S161P I115V 160 G97A-G128A-Y217Q-V203Y I115V 161 G97A-G128A-Y217Q-G211T I115V 162 G97A-G128A-Y217Q-K213T I115V 163 G97A-G128A-Y217Q-P239R I115V 164 G97A-G128A-Y217Q-N240K I115V 165 G97A-G128A-Y217Q-L267V I115V 166 G97A-G128A-Y217Q-A273S I115V 167 G97A-G128A-Y217Q-A144K P129Q-S130G-G131S 168 G97A-G128A-Y217Q-S145D P129Q-S130G-G131S 169 G97A-G128A-Y217Q-S159K P129Q-S130G-G131S 170 G97A-G128A-Y217Q-S162K P129Q-S130G-G131S 171 G97A-G128A-Y217Q-S161P P129Q-S130G-G131S 172 G97A-G128A-Y217Q-V203Y P129Q-S130G-G131S 173 G97A-G128A-Y217Q-G211T P129Q-S130G-G131S 174 G97A-G128A-Y217Q-K213T P129Q-S130G-G131S 175 G97A-G128A-Y217Q-P239R P129Q-S130G-G131S 176 G97A-G128A-Y217Q-N240K P129Q-S130G-G131S 177 G97A-G128A-Y217Q-L267V P129Q-S130G-G131S 178 G97A-G128A-Y217Q-A273S P129Q-S130G-G131S 179 G97A-G128A-Y217Q-A144K P129Q 180 G97A-G128A-Y217Q-S145D P129Q 181 G97A-G128A-Y217Q-S159K P129Q 182 G97A-G128A-Y217Q-S162K P129Q 183 G97A-G128A-Y217Q-S161P P129Q 184 G97A-G128A-Y217Q-V203Y P129Q 185 G97A-G128A-Y217Q-G211T P129Q 186 G97A-G128A-Y217Q-K213T P129Q 187 G97A-G128A-Y217Q-P239R P129Q 188 G97A-G128A-Y217Q-N240K P129Q 189 G97A-G128A-Y217Q-L267V P129Q 190 G97A-G128A-Y217Q-A273S P129Q 191 G97A-G128A-Y217Q-A144K A134T 192 G97A-G128A-Y217Q-S145D A134T 193 G97A-G128A-Y217Q-S159K A134T 194 G97A-G128A-Y217Q-S162K A134T 195 G97A-G128A-Y217Q-S161P A134T 196 G97A-G128A-Y217Q-V203Y A134T 197 G97A-G128A-Y217Q-G211T A134T 198 G97A-G128A-Y217Q-K213T A134T 199 G97A-G128A-Y217Q-P239R A134T 200 G97A-G128A-Y217Q-N240K A134T 201 G97A-G128A-Y217Q-L267V A134T 202 G97A-G128A-Y217Q-A273S A134T 203 G97A-G128A-Y217Q-S159K A144K 204 G97A-G128A-Y217Q-S162K A144K 205 G97A-G128A-Y217Q-S161P A144K 206 G97A-G128A-Y217Q-V203Y A144K 207 G97A-G128A-Y217Q-G211T A144K 208 G97A-G128A-Y217Q-K213T A144K 209 G97A-G128A-Y217Q-P239R A144K 210 G97A-G128A-Y217Q-N240K A144K 211 G97A-G128A-Y217Q-L267V A144K 212 G97A-G128A-Y217Q-A273S A144K 213 G97A-G128A-Y217Q-S159K S145D 214 G97A-G128A-Y217Q-S162K S145D 215 G97A-G128A-Y217Q-S161P S145D 216 G97A-G128A-Y217Q-V203Y S145D 217 G97A-G128A-Y217Q-G211T S145D 218 G97A-G128A-Y217Q-K213T S145D 219 G97A-G128A-Y217Q-P239R S145D 220 G97A-G128A-Y217Q-N240K S145D 221 G97A-G128A-Y217Q-L267V S145D 222 G97A-G128A-Y217Q-A273S S145D 223 G97A-G128A-Y217Q-V203Y S159K 224 G97A-G128A-Y217Q-G211T S159K 225 G97A-G128A-Y217Q-K213T S159K 226 G97A-G128A-Y217Q-P239R S159K 227 G97A-G128A-Y217Q-N240K S159K 228 G97A-G128A-Y217Q-L267V S159K 229 G97A-G128A-Y217Q-A273S S159K 230 G97A-G128A-Y217Q-V203Y S162K 231 G97A-G128A-Y217Q-G211T S162K 232 G97A-G128A-Y217Q-K213T S162K 233 G97A-G128A-Y217Q-P239R S162K 234 G97A-G128A-Y217Q-N240K S162K 235 G97A-G128A-Y217Q-L267V S162K 236 G97A-G128A-Y217Q-A273S S162K 237 G97A-G128A-Y217Q-V203Y S161P 238 G97A-G128A-Y217Q-G211T S161P 239 G97A-G128A-Y217Q-K213T S161P 240 G97A-G128A-Y217Q-P239R S161P 241 G97A-G128A-Y217Q-N240K S161P 242 G97A-G128A-Y217Q-L267V S161P 243 G97A-G128A-Y217Q-A273S S161P 244 G97A-G128A-Y217Q-G211T V203Y
245 G97A-G128A-Y217Q-K213T V203Y 246 G97A-G128A-Y217Q-P239R V203Y 247 G97A-G128A-Y217Q-N240K V203Y 248 G97A-G128A-Y217Q-L267V V203Y 249 G97A-G128A-Y217Q-A273S V203Y 250 G97A-G128A-Y217Q-P239R G211T 251 G97A-G128A-Y217Q-N240K G211T 252 G97A-G128A-Y217Q-L267V G211T 253 G97A-G128A-Y217Q-A273S G211T 254 G97A-G128A-Y217Q-P239R K213T 255 G97A-G128A-Y217Q-N240K K213T 256 G97A-G128A-Y217Q-L267V K213T 257 G97A-G128A-Y217Q-A273S K213T 258 G97A-G128A-Y217Q-L267V P239R 259 G97A-G128A-Y217Q-A273S P239R 260 G97A-G128A-Y217Q-L267V N240K 261 G97A-G128A-Y217Q-A273S N240K
Example 12
Table of Detergents
[0612] The compositions of the detergents used in the assays for Part I Example 12 are shown in Table 12-1. BPN' variant protein samples were added to the detergent compositions as described in Part I Example 1 to assay for the various properties listed.
TABLE-US-00047 TABLE 12-1 Composition of Detergents Used in the Assays to Test BPN' Variants Composition (wt % of Composition) Ingredient 1 2 3 C.sub.12-15 Alkylethoxy(1.8)sulphate 14.7 11.6 C.sub.11.8 Alkylbenzene sulfonate 4.3 11.6 8.3 C.sub.16-17 Branched alkyl sulphate 1.7 1.29 C.sub.12-14 Alkyl -9-ethoxylate 0.9 1.07 C.sub.12 dimethylamine oxide 0.6 0.64 Citric acid 3.5 0.65 3 C.sub.12-14 fatty acid 1.5 2.32 3.6 Sodium Borate (Borax) 2.5 2.46 1.2 Sodium C.sub.12-14 alkyl ethoxy 3 sulfate 2.9 C.sub.14-15 alkyl 7-ethoxylate 4.2 C.sub.12-14 Alkyl -7-ethoxylate 1.7 Ca formate 0.09 0.09 A compound having the following general structure: 1.2 bis((C.sub.2H.sub.5O)(C.sub.2H.sub.4O)n)(CH.sub.3)--N.sup.+--C.sub.xH.sub.- 2x--N.sup.+--(CH.sub.3)- bis((C.sub.2H.sub.5O)(C.sub.2H.sub.4O)n), wherein n = from 20 to 30, and x = from 3 to 8, or sulphated or sulphonated variants thereof Random graft co-polymer.sup.1 1.46 0.5 Ethoxylated Polyethylenimine .sup.2 1.5 1.29 Diethylene triamine pentaacetic acid 0.34 0.64 Diethylene triamine penta(methylene phosphonic acid) 0.3 Tinopal AMS-GX 0.06 Tinopal CBS-X 0.2 0.17 Amphiphilic alkoxylated grease cleaning polymer .sup.3 1.28 1 0.4 Ethanol 2 1.58 1.6 Propylene Glycol 3.9 3.59 1.3 Diethylene glycol 1.05 1.54 Polyethylene glycol 0.06 0.04 Monoethanolamine 3.05 2.41 0.4 NaOH 2.44 1.8 Sodium Cumene Sulphonate 1 Sodium Formate 0.11 Water, Aesthetics (Dyes, perfumes) and Minors balance balance balance (Enzymes, solvents, structurants) .sup.1"Random graft copolymer" is a polyvinyl acetate grafted polyethylene oxide copolymer having a polyethylene oxide backbone and multiple polyvinyl acetate side chains. The molecular weight of the polyethylene oxide backbone is about 6000 and the weight ratio of the polyethylene oxide to polyvinyl acetate is about 40 to 60 and no more than 1 grafting point per 50 ethylene oxide units. .sup.2 Polyethylenimine (MW = 600) with 20 ethoxylate groups per --NH. .sup.3 Amphiphilic alkoxylated grease cleaning polymer is a polyethylenimine (MW = 600) with 24 ethoxylate groups per --NH and 16 propoxy late groups per --NH.
Stain Removal Performance of BPN' Combinatorial Variants
[0613] Experiments to evaluate the stain removal performance of BPN' combinatorial variants generated as described in Example 11 were performed using BMI stained microswatches. The assay was performed as described in Example 1 (BMI microswatch assay). Table 12-2 provides Performance Index (PI) values of variants generated from RCL4 library using BMI microswatch assay in Detergent Composition 1 at pH 8 and 16.degree. C. and Detergent Composition 1 at pH 8 and 32.degree. C. and BMI microswatch assay in heat deactivated commercial TIDE.RTM. 2.times. Cold (Procter & Gamble) detergent at 16.degree. C. and pH 8. Heat inactivation of commercial detergent formulas serves to destroy the endogenous enzymatic activity of any protein components while retaining the properties of nonenzymatic components. Heat inactivation of the detergents was performed by placing pre-weighed amounts of liquid detergent (in a glass bottle) in a water bath at 95.degree. C. for 2 hours. The TIDE.RTM. 2.times. Cold detergent was purchased from local supermarket stores. Both unheated and heated detergents were assayed within 5 minutes of dissolving the detergent, in order to accurately determine percentage deactivated. Enzyme activity was tested by AAPF assay. Working solutions were made from the heat inactivated stock. Appropriate amounts of water hardness and buffer were added to the detergent solutions to match the desired conditions (Table 12-2). The solutions were mixed by vortexing or inverting the bottles.
TABLE-US-00048 TABLE 12-2 Working Detergent Solutions Temp Detergent Hardness Detergent (.degree. C.) g/L pH Buffer Gpg TIDE .RTM. 2X Cold 16, 32 0.98 8 5 mM HEPES 6
[0614] The sequences of the variants listed in Table 12-3 are relative to BPN'-v3: G97A-G128A-Y217Q. The PI values are calculated relative to BPN'-v3. All mutants in this list have a PI cutoff equal or greater than 0.5 for at least one property tested. "Det. Comp." means Detergent Composition.
TABLE-US-00049 TABLE 12-3 Performance Index Values of Variants Generated from RCL4 Library Sequence TIDE .RTM. Det. Comp. Relative to BPN'- Detergent pH 1, pH 8, Sequence Relative to v3: G97A- 8, 16.degree. C., BMI Det. Comp. 1, pH 32.degree. C., BMI BPN' G128A-Y217Q PI 8, 16.degree. C., BMI PI PI S87T-A88L-S89G- S87T-A88L- 1.00 1.12 0.99 G97A-G128A-Y217Q S89G N61P-S63H-G97A- N61P-S63H 1.03 1.12 1.01 G128A-Y217Q S87G-A88V-S89A- S87G-A88V- 1.02 1.11 0.99 G97A-G128A-Y217Q S89A P86S-S87G-A88V- P86S-S87G- 1.00 1.10 1.00 G97A-G128A-Y217Q A88V Q59S-N61P-G97A- Q59S-N61P 1.01 1.09 1.00 G128A-Y217Q S24G-N25G-G97A- S24G-N25G 0.99 1.09 1.02 G128A-Y217Q N61P-N62S-G97A- N61P-N62S 0.99 1.06 0.98 G128A-Y217Q G97A-G128A-P129Q- P129Q-S130G- 0.96 1.06 0.99 S130G-G131S-Y217Q G131S L75S-N76Y-G97A- L75S-N76Y 0.99 1.06 1.00 G128A-Y217Q G97A-G128A-V203Y- V203Y 0.99 1.05 1.01 Y217Q T55P-G97A-G128A- T55P 0.98 1.04 0.98 Y217Q A88V-L90I-G97A- A88V-L90I 0.99 1.04 1.00 G128A-Y217Q G97A-G128A-G211R- G211R-N212S- 0.97 1.04 0.98 N212S-K213V-Y217Q K213V G23A-S24G-N25G- G23A-S24G- 0.98 1.04 0.98 G97A-G128A-Y217Q N25G T22N-S24A-G97A- T22N-S24A 0.98 1.03 0.97 G128A-Y217Q S24R-G97A-G128A- S24R 0.95 1.02 0.99 Y217Q G97A-A98S-G128A- A98S 0.95 1.02 0.99 Y217Q BPN'-v3: G97A- BPN'-v3 1.00 1.00 1.00 G128A-Y217Q G97A-G128A-T158G- T158G-S159G 0.95 0.99 0.97 S159G-Y217Q Q59E-N61P-G97A- Q59E-N61P 0.90 0.94 0.90 G128A-Y217Q G97A-A98E-G128A- A98E 0.92 0.91 0.90 Y217Q
[0615] The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from those in Table 12-3, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. The proteolytic activity of such protease variant may be greater than that of the BPN' or BPN'-v3 protease. Each such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0616] Table 12-4 provides Performance Index (PI) values of variants generated from RCL 5-7 and FS1-3 using BMI microswatch assay in Detergent Composition 1 at 16.degree. C. and pH 8, BMI microswatch assay in Detergent Composition 2 at 16.degree. C. and pH 8, and stability measured in Detergent Composition 3. PI values for specific activity by AAPF hydrolysis (Specific AAPF PI) were also determined. All assays were performed as described in Example 1. The sequences of the variants listed are relative to BPN'-v3: G97A-G128A-Y217Q. PI values were calculated relative to BPN'-v3. All mutants in this list have a PI cutoff equal or greater than 0.5 for at least one property tested. PI values less than 0.01 were modified to display 0.01 in bold italics.
TABLE-US-00050 TABLE 12-4 Performance Index Values of Variants Generated From RCL 5-7 and FS1-3 Det. Det. Sequence Comp. 1, Comp. 2, Relative to pH 8, pH 8, Det. Source BPN'-v3: 16.degree. C., 16.degree. C., Specific Comp. 3, of Sequence Relative G97A-G128A- BMI BMI AAPF Stability Variants to BPN' Y217Q PI PI PI PI RCL6 G97A-G128A- P86S-S87G- 1.21 1.14 1.17 0.03 Y217Q-P86S- A88V-A116N- S87G-A88V- N117S-N118G A116N-N117S- N118G FS1 G97A-G128A- S24G-N25G- 1.05 1.12 1.96 1.00 Y217Q-S24G- N61P-S101N N25G-N61P- S101N FS1 G97A-G128A- S24G-N25G- 1.04 1.10 1.71 0.03 Y217Q-S24G- S53G-T55P- N25G-S53G- S87T-A88L- T55P-S87T-A88L- S89G-S101N- S89G-S101N- V203Y V203Y FS1 G97A-G128A- N61P-S78N- 1.03 1.10 1.73 0.33 Y217Q-N61P- S101N-V203Y S78N-S101N- V203Y FS1 G97A-G128A- T55P-N61P- 1.06 1.10 1.97 0.39 Y217Q-T55P- S78N-S101N- N61P-S78N- V203Y S101N-V203Y FS1 G97A-G128A- S53G-T55P- 1.03 1.10 2.22 0.33 Y217Q-S53G- N61P-S78N- T55P-N61P-S78N- S87T-A88L- S87T-A88L-S89G- S89G-S101N S101N RCL7 G97A-G128A- V203Y-L267V 1.10 1.09 1.12 0.09 Y217Q-V203Y- L267V FS1 G97A-G128A- S24G-N25G- 1.04 1.09 1.82 1.13 Y217Q-S24G- T55P-S101N N25G-T55P- S101N RCL7 G97A-G128A- A134T-L267V 1.13 1.08 0.93 0.66 Y217Q-A134T- L267V FS1 G97A-G128A- S24G-N25G- 1.01 1.08 1.10 0.29 Y217Q-S24G- S53G-T55P- N25G-S53G- N61P-S78N- T55P-N61P-S78N- S87T-A88L- S87T-A88L-S89G S89G FS1 G97A-G128A- S24G-N25G- 1.07 1.08 2.11 0.09 Y217Q-S24G- S53G-N61P- N25G-S53G- S101N-V203Y N61P-S101N- V203Y RCL6 G97A-G128A- N25Y-Q59S- 1.07 1.08 0.85 0.66 Y217Q-N25Y- N61P Q59S-N61P RCL7 G97A-G128A- I111V-S161P 1.10 1.08 0.66 0.98 Y217Q-I111V- S161P RCL7 G97A-G128A- I115V-L267V 1.10 1.08 1.07 0.63 Y217Q-I115V- L267V FS1 G97A-G128A- T55P-S78N- 0.99 1.08 1.39 0.16 Y217Q-T55P- S87T-A88L- S78N-S87T-A88L- S89G-S101N- S89G-S101N- V203Y V203Y RCL6 G97A-G128A- N25Y-P129Q- 1.07 1.08 0.88 0.67 Y217Q-N25Y- S130G-G131S- P129Q-S130G- A137T G131S-A137T RCL7 G97A-G128A- N61P-S63H- 1.04 1.08 1.59 0.78 Y217Q-N61P- A128S-P129Q S63H-A128S- P129Q FS1 G97A-G128A- S53G-N61P- 1.03 1.08 1.91 0.09 Y217Q-S53G- S101N-V203Y N61P-S101N- V203Y FS1 G97A-G128A- S24G-N25G- 1.08 1.07 1.44 0.33 Y217Q-S24G- S53G-S78N- N25G-S53G- S87T-A88L- S78N-S87T-A88L- S89G-S101N S89G-S101N FS1 G97A-G128A- N61P-S78N- 1.07 1.07 1.80 0.31 Y217Q-N61P- S87T-A88L- S78N-S87T-A88L- S89G-S101N S89G-S101N RCL6 G97A-G128A- N25Y-N61P- 1.06 1.07 0.74 0.60 Y217Q-N25Y- S63H N61P-S63H RCL5 G97A-G128A- Q59S-N61P- 0.99 1.07 0.69 0.09 Y217Q-Q59S- V203Y N61P-V203Y RCL6 G97A-G128A- V8L-N25Y- 1.08 1.07 1.22 0.03 Y217Q-V8L- P129Q-S130G- N25Y-P129Q- G131S S130G-G131S RCL6 G97A-G128A- P86S-S87G- 1.16 1.07 1.26 0.03 Y217Q-P86S- A88V-P239R S87G-A88V- P239R FS1 G97A-G128A- S24G-N25G- 1.02 1.07 2.07 0.11 Y217Q-S24G- S53G-T55P- N25G-S53G- N61P-S101N- T55P-N61P- V203Y S101N-V203Y RCL5 G97A-G128A- S24G-N25G- 1.09 1.06 1.25 1.00 Y217Q-S24G- P129Q-S130G- N25G-P129Q- G131S S130G-G131S FS3 G97A-G128A- N240K 1.04 1.06 0.90 0.94 Y217Q-N240K RCL5 G97A-G128A- G23A-S24G- 1.06 1.06 1.05 0.06 Y217Q-G23A- N25G-G211R- S24G-N25G- N212S-K213V G211R-N212S- K213V RCL7 G97A-G128A- N61P-S63H- 1.03 1.06 0.43 1.14 Y217Q-N61P- S78N-I111V- S63H-S78N- A134T I111V-A134T RCL6 G97A-G128A- S63T-P86S- 1.17 1.06 1.46 0.02 Y217Q-S63T- S87G-A88V P86S-S87G-A88V RCL6 G97A-G128A- G23A-S24G- 1.08 1.06 0.91 0.21 Y217Q-G23A- N25G-A116N- S24G-N25G- N117S-N118G A116N-N117S- N118G FS1 G97A-G128A- S78N-S87T- 1.03 1.06 1.40 0.33 Y217Q-S78N- A88L-S89G- S87T-A88L-S89G- S101N S101N RCL6 G97A-G128A- S24G-N25G- 1.06 1.06 0.74 1.05 Y217Q-S24G- A116N- N25G-A116N- N117S-N118G N117S-N118G RCL7 G97A-G128A- T55P-N240K 1.02 1.06 0.91 0.93 Y217Q-T55P- N240K RCL6 G97A-G128A- T55P-P129V- 1.07 1.06 0.85 0.86 Y217Q-T55P- P194S P129V-P194S RCL6 G97A-G128A- N25Y-S87G- 1.07 1.05 1.14 0.08 Y217Q-N25Y- A88V-S89A S87G-A88V-S89A FS1 G97A-G128A- S24G-N25G- 1.02 1.05 1.19 0.06 Y217Q-S24G- S87T-A88L- N25G-S87T- S89G-S101N A88L-S89G- S101N RCL7 G97A-G128A- P129Q-S130G- 1.04 1.05 0.94 0.05 Y217Q-P129Q- G131S-V203Y S130G-G131S- V203Y RCL6 G97A-G128A- Q59S-N61P- 1.05 1.05 0.80 0.84 Y217Q-Q59S- N240K N61P-N240K FS3 G97A-G128A- S24R-P40E- 1.13 1.05 1.08 1.31 Y217Q-S24R- P129E-S159K- P40E-P129E- K265R S159K-K265R RCL7 G97A-G128A- P52S-T55P- 1.09 1.05 0.53 0.15 Y217Q-P52S- V203Y T55P-V203Y RCL6 G97A-G128A- S24R-P129E 1.11 1.05 0.88 0.59 Y217Q-S24R- P129E FS1 G97A-G128A- S24G-N25G- 0.98 1.05 1.17 1.16 Y217Q-S24G- S53G-N61P- N25G-S53G- S78N N61P-S78N FS1 G97A-G128A- S24G-N25G- 1.07 1.05 1.71 1.29 Y217Q-S24G- T55P-S78N- N25G-T55P- S101N S78N-S101N RCL6 G97A-G128A- P86S-S87G- 1.10 1.05 0.76 0.05 Y217Q-P86S- A88V-A116S- S87G-A88V- N117G-N118R A116S-N117G- N118R RCL6 G97A-G128A- N61P-S87T- 1.00 1.05 1.12 0.05 Y217Q-N61P- A88L-S89G S87T-A88L-S89G FS1 G97A-G128A- S24G-N25G- 0.96 1.05 0.96 0.29 Y217Q-S24G- S53G-T55P- N25G-S53G- S78N-S87T- T55P-S78N-S87T- A88L-S89G A88L-S89G RCL5 G97A-G128A- G23A-S24G- 1.06 1.05 0.89 0.13 Y217Q-G23A- N25G-N61P- S24G-N25G- S63H N61P-S63H RCL6 G97A-G128A- S24R-Q59S- 1.07 1.05 0.94 0.49 Y217Q-S24R- N61P Q59S-N61P RCL6 G97A-G128A- N61P-P129Q- 1.07 1.05 1.13 0.78 Y217Q-N61P- S130G-G131S P129Q-S130G- G131S FS1 G97A-G128A- S24G-N25G- 1.05 1.04 1.98 0.32 Y217Q-S24G- S53G-T55P- N25G-S53G- N61P-S78N- T55P-N61P-S78N- S87T-A88L- S87T-A88L-S89G- S89G-S101N S101N FS1 G97A-G128A- S24G-N25G- 1.05 1.04 1.57 0.15 Y217Q-S24G- S53G-T55P- N25G-S53G- S101N-V203Y T55P-S101N- V203Y FS1 G97A-G128A- N61P-S78N- 1.02 1.04 1.60 0.12 Y217Q-N61P- S87T-A88L- S78N-S87T-A88L- S89G-S101N- S89G-S101N- V203Y V203Y FS1 G97A-G128A- S24G-N25G- 1.03 1.04 2.01 1.27 Y217Q-S24G- S53G-T55P- N25G-S53G- S78N-S101N T55P-S78N- S101N FS1 G97A-G128A- S24G-N25G- 1.01 1.04 1.77 0.07 Y217Q-S24G- S53G-S101N- N25G-S53G- V203Y S101N-V203Y FS1 G97A-G128A- S24G-N25G- 1.09 1.04 1.54 0.28 Y217Q-S24G- S78N-S101N- N25G-S78N- V203Y S101N-V203Y RCL7 G97A-G128A- P129Q-S130G- 1.14 1.04 1.37 0.56 Y217Q-P129Q- G131S- S130G-G131S- A133V-L267V A133V-L267V FS1 G97A-G128A- S87T-A88L- 1.05 1.04 1.44 0.07 Y217Q-S87T- S89G-S101N A88L-S89G- S101N RCL6 G97A-G128A- G23A-S24G- 1.07 1.04 0.92 0.16 Y217Q-G23A- N25G-P239R S24G-N25G- P239R RCL6 G97A-G128A- S87G-A88V- 1.07 1.04 0.92 0.37 Y217Q-S87G- S89A-A116N- A88V-S89A- N117S-N118G
A116N-N117S- N118G RCL6 G97A-G128A- Q59S-N61P- 1.09 1.04 0.68 0.91 Y217Q-Q59S- A116S- N61P-A116S- N117G-N118R N117G-N118R RCL5 G97A-G128A- Q59S-N61P- 1.03 1.04 0.81 0.08 Y217Q-Q59S- S87T-A88L- N61P-S87T-A88L- S89G S89G FS1 G97A-G128A- S24G-N25G- 0.96 1.04 0.73 0.04 Y217Q-S24G- S53G-S87T- N25G-S53G- A88L-S89G- S87T-A88L-S89G- V203Y V203Y RCL7 G97A-G128A- A134T-G211T 1.11 1.04 0.71 0.35 Y217Q-A134T- G211T RCL7 G97A-G128A- T55P-A128S- 1.03 1.04 2.71 1.02 Y217Q-T55P- P129Q A128S-P129Q FS1 G97A-G128A- T55P-S78N- 1.05 1.04 1.56 0.33 Y217Q-T55P- S87T-A88L- S78N-S87T-A88L- S89G-S101N S89G-S101N RCL6 G97A-G128A- P86S-S87G- 1.09 1.04 0.83 0.04 Y217Q-P86S- A88V-T242R S87G-A88V- T242R RCL7 G97A-G128A- S161P-V203Y 1.04 1.04 0.81 0.09 Y217Q-S161P- V203Y FS1 G97A-G128A- S24G-N25G- 1.05 1.04 2.01 0.44 Y217Q-S24G- T55P-N61P- N25G-T55P- S78N-S101N- N61P-S78N- V203Y S101N-V203Y RCL7 G97A-G128A- G211T-L267V 1.08 1.03 1.16 0.30 Y217Q-G211T- L267V RCL6 G97A-G128A- P40E-T55P- 1.08 1.03 0.71 0.05 Y217Q-P40E- N269K T55P-N269K RCL6 G97A-G128A- S24R-A128S- 1.08 1.03 1.63 0.64 Y217Q-S24R- P129G A128S-P129G RCL6 G97A-G128A- S24G-N25G- 1.15 1.03 3.06 0.35 Y217Q-S24G- N61P-N62S- N25G-N61P- P194L-A232T N62S-P194L- A232T RCL6 G97A-G128A- T55P-A116S- 1.07 1.03 0.83 1.06 Y217Q-T55P- N117G-N118R A116S-N117G- N118R FS1 G97A-G128A- S24G-N25G- 1.03 1.03 1.76 0.32 Y217Q-S24G- S53G-S78N- N25G-S53G- S101N-V203Y S78N-S101N- V203Y RCL7 G97A-G128A- P129Q-S130G- 1.08 1.03 0.90 0.83 Y217Q-P129Q- G131S-N240K S130G-G131S- N240K FS1 G97A-G128A- S53G-T55P- 0.93 1.03 1.00 0.33 Y217Q-S53G- N61P-S78N- T55P-N61P-S78N- S87T-A88L- S87T-A88L-S89G S89G RCL6 G97A-G128A- N25Y-P129Q- 1.08 1.03 0.93 0.69 Y217Q-N25Y- S130G-G131S P129Q-S130G- G131S RCL6 G97A-G128A- T55P-I115V 1.06 1.03 1.07 1.07 Y217Q-T55P- I115V RCL6 G97A-G128A- N25Y-T55P 1.05 1.03 1.00 0.80 Y217Q-N25Y- T55P RCL6 G97A-G128A- G23A-S24G- 0.97 1.03 1.75 0.15 Y217Q-G23A- N25G-A128S- S24G-N25G- P129D A128S-P129D FS1 G97A-G128A- S53G-S78N- 1.00 1.03 1.68 0.13 Y217Q-S53G- S87T-A88L- S78N-S87T-A88L- S89G-S101N- S89G-S101N- P129S-V203Y P129S-V203Y RCL7 G97A-G128A- T55P-A134T 1.06 1.03 0.94 1.05 Y217Q-T55P- A134T RCL7 G97A-G128A- N61P-S63H- 1.03 1.02 0.49 1.18 Y217Q-N61P- S78N-I111V S63H-S78N-I111V FS2 Y217Q-N61P- N61P-A97G- 0.98 1.02 NA NA A97G-G102A- G102A- A128G-P1295 A128G-P129S FS1 G97A-G128A- S53G-N61P- 1.02 1.02 2.08 0.88 Y217Q-S53G- S101N N61P-S101N RCL5 G97A-G128A- Q59S-N61P- 1.01 1.02 0.90 0.20 Y217Q-Q59S- S87G-A88V- N61P-S87G- S89A A88V-S89A FS1 G97A-G128A- S53G-S87T- 0.96 1.02 1.11 0.04 Y217Q-S53G- A88L-S89G- S87T-A88L-S89G- S101N-V203Y S101N-V203Y RCL6 G97A-G128A- S87T-A88L- 1.06 1.02 1.21 0.10 Y217Q-S87T- S89G-P129S A88L-S89G- P129S FS1 G97A-G128A- S53G-T55P- 1.03 1.02 1.70 0.48 Y217Q-S53G- S78N-S101N- T55P-S78N- V203Y S101N-V203Y RCL5 G97A-G128A- T55P-P129Q- 1.01 1.02 1.18 0.91 Y217Q-T55P- S130G-G131S P129Q-S130G- G131S RCL5 G97A-G128A- Q59S-N61P- 0.98 1.02 0.83 0.80 Y217Q-Q59S- P129Q-S130G- N61P-P129Q- G131S S130G-G131S RCL7 G97A-G128A- A134T-P239R 1.03 1.02 0.53 0.98 Y217Q-A134T- P239R RCL5 G97A-G128A- T55P-V203Y 1.01 1.01 1.02 0.23 Y217Q-T55P- V203Y RCL7 G97A-G128A- T55P-S78N- 1.03 1.01 1.05 1.25 Y217Q-T55P- S89Y S78N-S89Y RCL5 G97A-G128A- T22N-S24A- 1.00 1.01 0.69 0.46 Y217Q-T22N- N61P-S63H S24A-N61P-S63H RCL7 G97A-G128A- S161P-L267V 1.05 1.01 1.00 0.66 Y217Q-S161P- L267V RCL6 G97A-G128A- T55P-L75H- 1.06 1.01 0.69 0.59 Y217Q-T55P- N76G L75H-N76G RCL7 G97A-G128A- A134T-S161P 1.07 1.01 0.73 1.00 Y217Q-A134T- S161P RCL7 G97A-G128A- S87T-A88L- 1.08 1.01 0.66 0.13 Y217Q-S87T- S89G-A134T A88L-S89G- A134T RCL6 G97A-G128A- T55P-A116N- 1.06 1.01 1.06 1.11 Y217Q-T55P- N117S-N118G A116N-N117S- N118G v12 G97A-G128S- A128S 1.02 1.01 1.65 1.00 Y217Q RCL7 G97A-G128A- T55P-S78N- 1.07 1.00 1.00 1.32 Y217Q-T55P- I115V S78N-I115V RCL6 G97A-G128A- Y6Q-P129Q- 1.03 1.00 0.98 0.23 Y217Q-Y6Q- S130G-G131S P129Q-S130G- G131S RCL7 G97A-G128A- S24R-P129Q- 1.06 1.00 1.11 0.61 Y217Q-S24R- S130G-G131S P129Q-S130G- G131S FS1 G97A-G128A- S24G-N25G- 0.99 1.00 1.61 1.21 Y217Q-S24G- S53G-S78N- N25G-S53G- S101N S78N-S101N RCL6 G97A-G128A- T55P-P129V 1.07 1.00 0.70 1.00 Y217Q-T55P- P129V v3 G97A-G128A- BPN'-v3 1.00 1.00 1.00 1.00 Y217Q FS2 G97A-Y217Q- N61P-N62Q- 0.95 1.00 NA NA N61P-N62Q- G100N-A128G G100N-A128G RCL6 G97A-G128A- T55P-P129Q 1.10 1.00 1.77 1.15 Y217Q-T55P- P129Q FS1 G97A-G128A- S24G-N25G- 1.05 1.00 1.84 0.12 Y217Q-S24G- S53G-T55P- N25G-S53G- N61P-S78N- T55P-N61P-S78N- S87T-A88L- S87T-A88L-S89G- S89G-S101N- S101N-V203Y V203Y RCL6 G97A-G128A- S87T-A88L- 1.04 1.00 0.83 0.10 Y217Q-S87T- S89G-N240K A88L-S89G- N240K RCL7 G97A-G128A- A134T-N240K 1.07 0.99 0.61 0.96 Y217Q-A134T- N240K RCL7 G97A-G128A- S87T-A88L- 1.04 0.99 0.68 0.14 Y217Q-S87T- S89G-P239R A88L-S89G- P239R RCL7 G97A-G128A- P129Q-S130G- 1.08 0.99 1.35 0.61 Y217Q-P129Q- G131S-L267V S130G-G131S- L267V RCL7 G97A-G128A- P129Q-N240K 1.06 0.99 1.39 1.00 Y217Q-P129Q- N240K FS1 G97A-G128A- S78N-S87T- 0.93 0.99 0.74 0.15 Y217Q-S78N- A88L-S89G- S87T-A88L-S89G- V203Y V203Y RCL7 G97A-G128A- I111V-A273S 1.00 0.99 0.48 0.27 Y217Q-I111V- A273S FS1 G97A-G128A- S24G-N25G- 1.03 0.99 2.17 0.71 Y217Q-S24G- T55P-S78N- N25G-T55P- A88V-S101N S78N-A88V- S101N FS1 G97A-G128A- S24G-N25G- 0.96 0.98 1.15 1.20 Y217Q-S24G- T55P-S78N N25G-T55P-S78N FS1 G97A-G128A- S24G-N25G- 1.02 0.98 1.27 0.10 Y217Q-S24G- S53G-S78N- N25G-S53G- S87T-A88L- S78N-S87T-A88L- S101N-V203Y S101N-V203Y FS1 G97A-G128A- S24G-N25G- 0.97 0.98 0.75 0.14 Y217Q-S24G- S53G-S78N- N25G-S53G- S87T-A88L- S78N-S87T-A88L- S89G-V203Y S89G-V203Y FS1 G97A-G128A- S24G-N25G- 1.06 0.98 1.21 0.14 Y217Q-S24G- S53G-S78N- N25G-S53G- S87T-A88L- S78N-S87T-A88L- S89G-S101N- S89G-S101N- V203Y V203Y RCL5 G97A-G128A- S87G-A88V- 1.01 0.98 1.18 0.29 Y217Q-S87G- S89A-P129Q- A88V-S89A- S130G-G131S P129Q-S130G- G131S RCL7 G97A-G128A- N61P-S63H- 1.03 0.98 0.73 1.16 Y217Q-N61P- S78N-S161P S63H-S78N- S161P FS1 G97A-G128A- T55P-N61P- 1.04 0.98 1.75 0.14 Y217Q-T55P- S78N-S87T- N61P-S78N-S87T- A88L-S89G- A88L-S89G- S101N-V203Y S101N-V203Y RCL7 G97A-G128A- I111V-P129Q- 1.00 0.98 0.52 0.97 Y217Q-I111V- S130G-G131S
P129Q-S130G- G131S RCL5 G97A-G128A- T22N-S24A- 0.96 0.98 0.94 0.64 Y217Q-T22N- T55P S24A-T55P RCL7 G97A-G128A- I115V-N240K 1.04 0.98 0.66 0.90 Y217Q-I115V- N240K RCL6 G97A-G128A- S87G-A88V- 0.98 0.98 0.66 0.40 Y217Q-S87G- S89A-A116N- A88V-S89A- N117S- A116N-N117S- N118G-P172H N118G-P172H FS1 G97A-G128A- S24G-N25G- 1.01 0.98 1.28 0.31 Y217Q-S24G- S78N-S87T- N25G-S78N- A88L-S89G- S87T-A88L-S89G- S101N S101N RCL6 G97A-G128A- S24G-N25G- 1.03 0.98 0.59 1.13 Y217Q-S24G- I115V-A134T N25G-I115V- A134T RCL6 G97A-G128A- T55P-A128S 0.97 0.97 1.39 1.00 Y217Q-T55P- P129D A128S-P129D RCL7 G97A-G128A- I111V-S159K 0.99 0.97 0.64 1.12 Y217Q-I111V- S159K RCL7 G97A-G128A- N240K-A273S 1.03 0.96 0.64 0.19 Y217Q-N240K- A273S RCL7 G97A-G128A- S159K-L267V 1.00 0.96 1.08 0.79 Y217Q-S159K- L267V RCL7 G97A-G128A- I111V-P129Q- 1.01 0.96 0.83 0.42 Y217Q-I111V- G211T P129Q-G211T RCL7 G97A-G128A- I115V-A273S 1.05 0.95 0.74 0.20 Y217Q-I115V- A273S RCL6 G97A-G128A- S89Y 0.99 0.95 0.70 0.88 Y217Q-S89Y RCL6 G97A-G128A- S24R-A116N- 1.08 0.95 0.91 0.60 Y217Q-S24R- N117S-N118G A116N-N117S- N118G RCL7 G97A-G128A- N61E-A144K 1.03 0.95 0.97 1.10 Y217Q-N61E- A144K RCL6 G97A-G128A- P129Q-S130G- 1.02 0.95 0.85 0.99 Y217Q-P129Q- G131S-P239R S130G-G131S- P239R RCL7 G97A-G128A- S87T-A88L- 0.97 0.95 0.59 0.13 Y217Q-S87T- S89G-I115V A88L-S89G- I115V RCL6 G97A-G128A- T55P-A92G 0.96 0.94 0.56 0.91 Y217Q-T55P- A92G FS3 G97A-G128A- S145D-S159K- 0.98 0.94 0.76 1.08 Y217Q-S145D- N240K-Q275E S159K-N240K- Q275E RCL7 G97A-G128A- S89Y-P129Q- 1.04 0.94 0.71 0.70 Y217Q-S89Y- S130G-G131S P129Q-S130G- G131S RCL7 G97A-G128A- P129Q-S130G- 1.01 0.94 1.00 0.93 Y217Q-P129Q- G131S-S162K S130G-G131S- S162K RCL7 G97A-G128A- I111V-A134T 0.98 0.94 0.37 1.06 Y217Q-I111V- A134T RCL6 G97A-G128A- P40E-S53Y- 0.99 0.94 0.97 0.35 Y217Q-P40E- S78Y-P86S- S53Y-S78Y-P86S- S87G-A88V S87G-A88V RCL6 G97A-G128A- S24G-N25G- 0.93 0.93 0.58 0.52 Y217Q-S24G- L75H-N76G N25G-L75H- N76G FS2 G97A-Y217Q- N61P-A128G- 0.85 0.93 0.65 0.92 N61P-A128G- P129S-S130P P129S-S130P RCL6 G97A-G128A- S24R-S145D 0.99 0.93 0.89 0.59 Y217Q-S24R- S145D FS3 G97A-G128A- S24R-S145D- 0.92 0.92 0.63 0.68 Y217Q-S24R- P239R-Q275E S145D-P239R- Q275E RCL7 G97A-G128A- S24R-S78N- 0.95 0.92 1.16 0.55 Y217Q-S24R- S182P-L267V S78N-S182P- L267V FS1 G97A-G128A- S53G-N61P- 1.04 0.92 1.67 0.02 Y217Q-S53G- S87T-A88L- N61P-S87T-A88L- S89G-S101N- S89G-S101N- V203Y V203Y RCL6 G97A-G128A- P5S-S87G- 0.98 0.92 0.64 0.07 Y217Q-P5S- A88V-S89A- S87G-A88V- A116G-N117R S89A-A116G- N117R FS1 G97A-G128A- S53G-N61P- 0.99 0.92 1.65 0.11 Y217Q-S53G- S78N-S87T- N61P-S78N-S87T- A88L-S89G- A88L-S89G- S101N-V203Y S101N-V203Y RCL6 G97A-G128A- Q59S-N61P- 0.93 0.92 0.42 1.04 Y217Q-Q59S- A116N- N61P-A116N- N117S-N118G N117S-N118G RCL7 G97A-G128A- P239R-A273S 0.94 0.91 0.52 0.23 Y217Q-P239R- A273S FS1 G97A-G128A- S53G-S78N- 0.94 0.91 1.16 0.13 Y217Q-S53G- S87T-A88L- S78N-S87T-A88L- S89G-S101N- S89G-S101N- V203Y V203Y RCL6 G97A-G128A- S24R-P129V 0.98 0.91 0.52 0.58 Y217Q-S24R- P129V RCL7 G97A-G128A- I111V-P239R 0.97 0.91 0.38 1.08 Y217Q-I111V- P239R FS1 G97A-G128A- S87T-A88L- 0.90 0.91 0.79 0.05 Y217Q-S87T- S89G-S101N- A88L-S89G- V203Y S101N-V203Y RCL6 G97A-G128A- T55P-P129L 0.99 0.91 0.62 0.98 Y217Q-T55P- P129L RCL7 G97A-G128A- S87T-A88L- 0.92 0.90 0.42 0.13 Y217Q-S87T- S89G-I111V A88L-S89G- I111V RCL7 G97A-G128A- S145D-A273S 0.91 0.90 0.66 0.17 Y217Q-S145D- A273S RCL6 G97A-G128A- P129Q-S130G- 0.94 0.90 0.51 0.89 Y217Q-P129Q- G131S-T242R S130G-G131S- T242R RCL7 G97A-G128A- S3F-S87T- 0.93 0.89 0.55 0.07 Y217Q-S3F-S87T- A88L-S89G- A88L-S89G- G211T G211T RCL6 G97A-G128A- S87G-A88V- 0.97 0.89 1.21 0.37 Y217Q-S87G- S89A-S162K A88V-S89A- S162K RCL7 G97A-G128A- S89Y-G211T 0.89 0.88 0.53 0.41 Y217Q-S89Y- G211T RCL7 G97A-G128A- S87T-A88L- 0.93 0.88 0.74 0.12 Y217Q-S87T- S89G-A144K A88L-S89G- A144K RCL6 G97A-G128A- P129Q-S130G- 0.95 0.88 1.21 0.96 Y217Q-P129Q- G131S-S159K S130G-G131S- S159K RCL6 G97A-G128A- A116N- 0.90 0.88 0.54 0.95 Y217Q-A116N- N117S- N117S-N118G- N118G- P129Q-S130G- P129Q-S130G- G1315 G1315 RCL6 G97A-G128A- S24G-N25G- 0.86 0.87 0.44 1.02 Y217Q-S24G- P129V N25G-P129V FS1 G97A-G128A- S24G-N25G- 0.87 0.86 0.83 0.12 Y217Q-S24G- S78N-S87T- N25G-S78N- A88L-S89G- S87T-A88L-S89G- S101N-V203Y S101N-V203Y FS2 G97A-Y217Q- N123G-A128G 0.83 0.86 1.00 0.14 N123G-A128G FS2 G97A-G128A- N61P-N62Q- 0.82 0.86 5.52 0.40 Y217Q-N61P- G100N- N62Q-G100N- G102A-M124I G102A-M124I RCL6 G97A-G128A- S24G-N25G- 0.90 0.85 0.65 1.10 Y217Q-S24G- K141E-T242R N25G-K141E- T242R RCL6 G97A-G128A- S87G-A88V- 0.89 0.85 0.44 0.32 Y217Q-S87G- S89A-A116N- A88V-S89A- N117S- A116N-N117S- N118G-A144T N118G-A144T FS1 G97A-G128A- T55P-N61P- 0.83 0.85 0.92 0.30 Y217Q-T55P- S87T-A88L- N61P-S87T-A88L- S89G-G110C- S89G-G110C- S130P S130P RCL6 G97A-G128A- L75S-N76Y- 0.84 0.83 0.41 0.13 Y217Q-L75S- A116S- N76Y-A116S- N117G-N118R N117G-N118R Y217Q FS3 G97A-G128A- S145D-S159K- 0.74 0.81 0.38 0.65 Y217Q-S145D- K213L-P239R- S159K-K213L- N240K P239R-N240K RCL6 G97A-G128A- S24R-S87T- 0.86 0.80 0.53 0.08 Y217Q-S24R- A88L-S89G S87T-A88L-S89G RCL6 G97A-G128A- G23A-S24G- 0.88 0.79 0.53 0.20 Y217Q-G23A- N25G-P129V S24G-N25G- P129V RCL7 G97A-G128A- A134T-K213L 0.87 0.79 0.51 0.81 Y217Q-A134T- K213L RCL7 G97A-G128A- S89Y-A273S 0.81 0.79 0.48 0.19 Y217Q-S89Y- A2735 RCL7 G97A-G128A- S24R-P239R 0.90 0.78 0.71 0.69 Y217Q-S24R- P239R FS2 G97A-Y217Q- N123G- 0.76 0.78 0.71 0.12 N123G-A128G- A128G-P129S P129S RCL7 G97A-G128A- S89Y-P239R 0.77 0.76 0.38 0.90 Y217Q-S89Y- P239R RCL6 G97A-G128A- S24G-N25G- 0.71 0.76 0.25 0.59 Y217Q-S24G- A92G N25G-A92G RCL6 G97A-G128A- N61P-S63H- 0.73 0.74 0.24 0.42 Y217Q-N61P- I115V-A228V S63H-I115V- A228V FNA Y217L A97G-A128G- 0.73 0.74 1.81 1.21 Q217L RCL6 G97A-G128A- L75S-N76Y- 0.73 0.73 0.34 0.09 Y217Q-L75S- P129V N76Y-P129V RCL6 G97A-G128A- S24R-P129L 0.80 0.73 0.35 0.58 Y217Q-S24R- P129L RCL6 G97A-G128A- S87G-A88V- 0.69 0.73 0.90 0.36 Y217Q-S87G- S89A-P129Q- A88V-S89A- S182Y-S204Y- P129Q-S182Y- P239Q S204Y-P239Q
RCL6 G97A-G128A- S24R-A92G 0.75 0.73 0.28 0.44 Y217Q-S24R- A92G RCL6 G97A-G128A- S24R-A116S- 0.81 0.72 0.48 0.58 Y217Q-S24R- N117G-N118R A116S-N117G- N118R RCL6 G97A-G128A- G23A-S24G- 0.85 0.66 0.63 0.20 Y217Q-G23A- N25G-A116G- S24G-N25G- N117R A116G-N117R RCL6 G97A-G128A- S24G-N25G- 0.72 0.66 0.39 1.03 Y217Q-S24G- P129L N25G-P129L RCL6 G97A-G128A- S87T-A88L- 0.78 0.66 0.65 0.11 Y217Q-S87T- S89G-S101G A88L-S89G- S101G RCL6 G97A-G128A- G23A-S24G- 0.52 0.59 0.37 0.20 Y217Q-G23A- N25G-P129L S24G-N25G- P129L FS1 G97A-G128A- S53G-N61P- 0.55 0.58 0.29 0.16 Y217Q-S53G- G102A-V203Y N61P-G102A- V203Y RCL6 G97A-G128A- T55P-V147P 0.56 0.57 0.22 1.09 Y217Q-T55P- V147P RCL6 G97A-G128A- Y6Q-L75S- 0.55 0.52 0.24 0.17 Y217Q-Y6Q- N76Y L75S-N76Y RCL6 G97A-G128A- N61P-S63H- 0.36 0.37 0.14 0.95 Y217Q-N61P- V147P S63H-V147P RCL6 G97A-G128A- S24R-V147P 0.16 0.22 0.08 0.85 Y217Q-S24R- V147P RCL6 G97A-G128A- S24G-N25G- 0.21 0.18 0.06 0.71 Y217Q-S24G- V68C-A69G N25G-V68C- A69G FS1 G97A-G128A- S24G-N25G- 0.19 0.18 0.08 0.38 Y217Q-S24G- N61P-L75S- N25G-N61P- N76Y-S101N- L75S-N76Y- V203Y S101N-V203Y FS1 G97A-G128A- L75S-N76Y- 0.13 0.12 0.05 0.72 Y217Q-L75S- S78N-S101N- N76Y-S78N- V203Y S101N-V203Y FS1 G97A-G128A- L75S-N76Y- 0.04 0.04 0.07 0.73 Y217Q-L75S- S78N-S87T- N76Y-S78N- A88L-S89G- S87T-A88L-S89G- S101N-S130P S101N-S130P FS1 G97A-G128A- S24G-N25G- 0.06 0.04 0.06 0.33 Y217Q-S24G- S53G-S101N- N25G-S53G- S130P-V203Y S101N-S130P- V203Y RCL6 G97A-G128A- G47E-M50I- 0.01 0.03 0.02 1.21 Y217Q-G47E- L75S-N76Y- M50I-L75S- S162K N76Y-S162K FS1 G97A-G128A- S53G-T55P- 0.01 0.02 0.05 0.45 Y217Q-S53G- S87T-A88L- T55P-S87T-A88L- S89G-S101N- S89G-S101N- S130P-V203Y S130P-V203Y FS1 G97A-G128A- S24G-N25G- 0.01 0.01 0.02 1.21 Y217Q-S24G- L75S-N76Y- N25G-L75S- A128T-P129T- N76Y-A128T- S130G- P129T-S130G- G131Q-S132C- G131Q-S132C- A133G-A134T A133G-A134T RCL6 G97A-G128A- P129Q-S130G- 0.01 0.01 0.03 1.11 Y217Q-P129Q- G131S-V147P S130G-G131S- V147P FS1 G97A-G128A- S53G-T55P- 0.01 0.01 0.02 1.05 Y217Q-S53G- N61P-L75S- T55P-N61P-L75S- N76Y-S87T- N76Y-S87T- A88L-S89G- A88L-S89G- G102A-S130P- G102A-S130P- V203Y V203Y FS1 G97A-G128A- S53G-T55P- 0.03 0.01 0.02 1.03 Y217Q-S53G- N61P-L75S- T55P-N61P-L75S- N76Y-S101N- N76Y-S101N- S130P-V203Y S130P-V203Y
[0617] The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from those listed in Table 12-4, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Each such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0618] Table 12-5 provides the Performance Index (PI) values of BPN' variants (generated as described in "Generation of Variants to Improve BPN' Stability"; see Table 11-3) for stain removal in BMI microswatch assay in Detergent Composition 1 at 16.degree. C. and pH 8 (Det. Comp. 1, pH 8, 16.degree. C., BMI PI) and for stability in LAS/EDTA (LAS/EDTA Stability PI). Assays were performed as described in Example 1 (BMI microswatch assay, LAS/EDTA stability assay). The sequences of the variants are shown relative to both BPN' and FNA. That is, each variant sequence is the BPN' or FNA sequence with the specified variant amino acid substitutions. PI values are shown relative to FNA parent, which is BPN'-Y217L.
TABLE-US-00051 TABLE 12-5 Performance Index of Stability-Improved BPN' Variants Det. Comp. 1 Sequence Relative to Sequence pH 8, 16.degree. C., LAS/EDTA FNA: BPN' Y217L Relative to BPN' BMI PI Stability PI P40E-S78N-S87D P40E-S78N- 0.71 11.65 S87D-Y217L P40E P40E-Y217L 0.96 8.33 T22V-S78N-Q206E- T22V-S78N- 0.75 5.95 K213N Q206E-K213N- Y217L T22V-S78N-K213N T22V-S78N- 0.86 5.71 K213N-Y217L S87D S87D-Y217L 0.90 4.04 S78N S78N-Y217L 0.87 3.86 K213N K213N-Y217L 0.91 1.84 Q206E Q206E-Y217L 0.86 1.76 T22V T22V-Y217L 0.97 1.46 FNA Y217L 1.00 1.00
[0619] The invention includes a protease variant having proteolytic activity and/or improved stability relative to FNA, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from those listed in Table 12-5, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Such protease variant may have a proteolytic activity greater than that of BPN' or BPN'-v3. Each such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0620] Table 12-6 provides the Performance Index (PI) values of BPN' variants generated from Library Parent: BPN'-v3: G97A-G128A-Y217Q (as described in "Generation of Variants to Improve BPN' Stability"; see Table 11-3) for stain removal in a BMI microswatch assay in Detergent Composition 1 at 16.degree. C. and pH 8 and for stability in LAS/EDTA. Assays were performed as described in Example 1 (BMI microswatch assay and LAS/EDTA stability assay). The Performance Index was calculated relative to BPN'-v3: G97A-G128A-Y217Q. All mutants in this list have a PI cutoff equal or greater than 0.5 for at least one property tested.
TABLE-US-00052 TABLE 12-6 Performance Index of Stability-Improved BPN' Variants Sequence Relative to BPN'-v3: BPN' Det. Comp. 1 G97A-G128A- pH 8, 16.degree. C., LAS/EDTA Y217Q Sequence Relative to BPN' BMI PI Stability PI S87D-N76D-S78N S87D-N76D-S78N-G97A-G128A-Y217Q 0.62 2.27 P40E-S78N-S87D P40E-S78N-S87D-G97A-G128A-Y217Q 0.21 2.18 P40E-S87D P40E-S87D-G97A-G128A-Y217Q 0.18 2.14 S78N-P40E S78N-P40E-G97A-G128A-Y217Q 0.80 2.03 S87D-N76D S87D-N76D-G97A-G128A-Y217Q 0.55 1.89 P40E P40E-G97A-G128A-Y217Q 0.84 1.79 S78N-S87D S78N-S87D-G97A-G128A-Y217Q 0.79 1.71 S87D S87D-G97A-G128A-Y217Q 0.78 1.20 S78N S78N-G97A-G128A-Y217Q 0.93 1.14 BPN'-v3 G97A-G128A-Y217Q 1.00 1.00
[0621] The invention includes a protease variant having proteolytic activity and having improved stability relative to BPN'-v3, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from those listed in Table 12-6, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Each such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0622] Table 12-7 provides the Performance Index (PI) values of BPN' variants (generated as described in "Generation of BPN' Variants from Five Different Plasmids"; see Table 11-4) for stain removal in a BMI microswatch assay in Detergent Composition lat 16.degree. C. and pH 8. Assays were performed as described in Example 1 (BMI microswatch assay). The Performance Index of each variant was calculated relative to BPN'-v3: G97A-G128A-Y217Q. All mutants in this list have a PI cutoff equal or greater than 0.5 for at least one property tested.
TABLE-US-00053 TABLE 12-7 Performance Index of BPN' Variants Det. Comp. 1, Sequence Relative to BPN'- pH 8, 16.degree. C., v3 G97A-G128A-Y217Q Sequence Relative to BPN' BMI PI S101N G97A-S101N-G128A-Y217Q 1.12 A137V G97A-G128A-A137V-Y217Q 1.12 N61P N61P-G97A-G128A-Y217Q 1.11 S130P G97A-G128A-S130P-Y217Q 1.09 Q103N G97A-Q103N-G128A-Y217Q 1.07 S63T S63T-G97A-G128A-Y217Q 1.03 G102A G97A-G102A-G128A-Y217Q 1.02 BPN'-v3 BPN'-v3 (G97A-G128A-Y217Q) 1.00 N109D-S248R G97A-N109D-G128A-Y217Q-S248R 0.96 S87R S87R-G97A-G128A-Y217Q 0.95 S188D G97A-G128A-S188D-Y217Q 0.95 S87D-S248R S87D-G97A-G128A-Y217Q-S248R 0.94 S188D-S248R G97A-G128A-S188D-S248R-Y217Q 0.93 S248D G97A-G128A-S248D-Y217Q 0.86 N109D-S188D-S248R G97A-N109D-G128A-S188D-S248R- 0.83 Y217Q N109D G97A-N109D-G128A-Y217Q 0.81 S87R-S248R S87R-G97A-G128A-Y217Q-S248R 0.79 N109D-S188R G97A-N109D-G128A-S188R-Y217Q 0.77 N76D N76D-G97A-G128A-Y217Q 0.75 S87D-N109D-S188D- S87D-G97A-N109D-G128A-S188D- 0.58 S248R S248R-Y217Q S87R-N109D-S188D- S87R-G97A-N109D-G128A-S188D- 0.55 S248R Y217Q-S248R S87R-S188R-S248R S87R-G97A-G128A-S188R-Y217Q-S248R 0.52 A187D G97A-G128A-A187D-Y217Q 0.48 N109D-S248D G97A-N109D-G128A-Y217Q-S248D 0.47 S87R-N109R-S188R-S248R S87R-G97A-N109R-G128A-S188R- 0.39 Y217Q-S248R F189D G97A-G128A-F189D-Y217Q 0.31 G100N G97A-G100N-G128A-Y217Q 0.28 S87R-N109D-S188D S87R-G97A-N109D-G128A-S188D- 0.24 Y217Q S87D-N109D-S188D S87D-G97A-N109D-G128A-S188D- 0.12 Y217Q S87R-S188D-S248D S87R-G97A-G128A-S188D-S248D-Y217Q 0.09 N62D N62D-G97A-G128A-Y217Q 0.09 S87D-N109D-S188D- S87D-G97A-N109D-G128A-S188D- 0.08 S248D Y217Q-S248D
[0623] The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from those listed in Table 12-7, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Each such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0624] Table 12-8 provides the Performance index (PI) values of BPN' variants (generated as described in "Generation of Combinatorial Libraries and Variants of BPN'-v3+578N" as described in Example 3) for stain removal using a BMI microswatch assay in Detergent Composition 1 at 16.degree. C. and pH 8. Assays were performed as described in Example 1 (BMI microswatch assay). The Performance Index of each variant was calculated relative to BPN'-S78N-G97A-G128A-Y217Q. PI values less than 0.01 were modified and are indicated as "0.01" in bold italics.
TABLE-US-00054 TABLE 12-8 Performance Index Values of BPN' Variants Sequence Relative to Det. Comp. 1 BPN'-v3: G97A- pH 8, 16.degree. C., Variant Sequence Relative to BPN' G128A-Y217Q BMI PI v3/S78N/L267V S78N-G97A-G128A- S78N-L267V 1.12 Y217Q-L267V v3/S78N/S161P S78N-G97A-G128A- S78N-S161P 1.05 Y217Q-S161P v3/S78N/I115V S78N-G97A-G128A- S78N-I115V 1.04 Y217Q-I115V v3/S78N/A273S S78N-G97A-G128A- S78N-A273S 1.03 Y217Q-A273S v3/S78N/G211T S78N-G97A-G128A- S78N-G211T 1.00 Y217Q-G211T V3 + S78N S78N-G97A-G128A- S78N 1.00 Y217Q v3/S78N/I111V S78N-G97A-G128A- S78N-I111V 0.98 Y217Q-I111V v3/S78N/V147L S78N-G97A-G128A- S78N-V147L 0.97 Y217Q-V147L v3/S78N/I108V S78N-G97A-G128A- S78N-I108V 0.97 Y217Q-I108V v3/S78N/S89Y S78N-G97A-G128A- S78N-S89Y 0.94 Y217Q-S89Y v3/S78N/A138T S78N-G97A-G128A- S78N-A138T 0.92 Y217Q-A138T v3/S78N/P172V S78N-G97A-G128A- S78N-P172V 0.74 Y217Q-P172V v3/S78N/Q59G S78N-G97A-G128A- S78N-Q59G 0.64 Y217Q-Q59G GcM96 G97A-G128A-Y217Q- P129T-V147Q- 0.57 P129T-V147Q-S159D- S159D-S161P- S161P-S183T-Q185T- S183T-Q185T- G211A-S224A G211A-S224A GcM91 G97A-G128A-Y217Q- Q059V-I108V- 0.55 Q059V-I108V-V147Q- V147Q-G211A- G211A-N252Q N252Q v3/S78N/Y167A S78N-G97A-G128A- S78N-Y167A 0.53 Y217Q-Y167A v3/S78N/A92G S78N-G97A-G128A- S78N-A92G 0.49 Y217Q-A92G v3/S78N/P129L S78N-G97A-G128A- S78N-P129L 0.48 Y217Q-P129L GcM92 G97A-G128A-Y217Q- N061A-S087E- 0.36 N061A-S087E-M124I- M124I-S161P-S224A S161P-S224A v3/S78N/N62Q S78N-G97A-G128A- S78N-N62Q 0.27 Y217Q-N62Q v3/S78N/V68A S78N-G97A-G128A- S78N-V68A 0.24 Y217Q-V68A GcM94 G97A-G128A-Y217Q- S063T-S101A- 0.12 S063T-S101A-L126V- L126V-S183T- S183T-T244N T244N v3/S78N/M124T S78N-G97A-G128A- S78N-M124T 0.05 Y217Q-M124T GcM95 G97A-G128A-Y217Q- P040L-S053G- P040L-S053G-Q059V- Q059V-N061A- N061A-N062Q-S063T- N062Q-S063T- S087E-G100N S087E-G100N GcM93 G97A-G128A-Y217Q- N062Q-G100N- N062Q-G100N-S125A- S125A-S159D- S159D-N240S N240S GcM100 G97A-G128A-Y217Q- V68A-G102A- V68A-G102A-G211A- G211A-S125A S125A GcM90 G97A-G128A-Y217Q- S053G-V068A- S053G-V068A-G102A- G102A-P129T- P129T-Q185T Q185T
[0625] The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one set of amino acid substitutions selected from those listed in Table 12-8, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Each such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0626] Table 12-9 provides Performance index (PI) values of BPN' single variants (constructed using PCR fusion as described in PCT App. No. PCT/US09/46156, filed Jun. 3, 2009, which is incorporated by reference herein for such description) for stain removal in a BMI microswatch assay in Detergent Composition 2 at 16.degree. C. and pH 8 and for stability measured in Detergent Composition 3. PI values for specific activity by AAPF hydrolysis (PI specific AAPF) were determined. All assays were performed as described in Example 1. Performance index values were calculated relative to BPN wild type. PI values less than 0.01 are indicated as "0.01" in bold italics. "Det. Comp." means Detergent Composition.
TABLE-US-00055 TABLE 12-9 Performance Index Values for BPN' Single Variants Det. Comp. 2 BPN' pH 8, 16.degree. C., PI Specific Det. Comp. 3 Variant BMI PI AAPF Stability PI S182E 1.34 1.05 0.50 N109I 1.28 1.22 0.20 N117H 1.15 0.25 0.20 K237D 1.15 0.60 0.70 L257Q 1.14 0.94 0.80 P225N 1.13 1.02 0.70 S105H 1.11 1.07 S236I 1.10 0.58 0.90 L235H 1.10 0.65 0.70 S249E 1.07 0.72 0.30 N76E 1.07 0.76 0.20 S145N 1.06 1.16 1.10 N243D 1.05 1.03 0.90 R247N 1.04 1.04 0.50 E195N 1.04 1.05 0.40 A98K 1.03 0.75 0.70 S182N 0.99 1.14 0.90 S161H 0.97 1.07 0.90 G83H 0.95 0.72 0.60 G131D 0.95 1.11 1.30 T71C 0.93 1.00 1.30 K136Q 0.93 1.02 0.80 P40D 0.93 1.20 1.20 A187H 0.91 0.95 0.60 L250K 0.90 1.02 0.40 S9I 0.87 0.20 N76M 0.85 0.60 0.60 S132D 0.85 0.88 0.70 Q19F 0.83 0.47 0.30 E112H 0.83 1.02 S249P 0.80 1.02 0.50 S53D 0.78 0.29 0.10 V68E 0.78 0.59 1.70 D41I 0.72 1.15 0.90 K43H 0.70 0.28 0.10 V4H 0.66 0.37 0.60 A13Y 0.64 0.48 N62P 0.61 0.60 1.30 L196E 0.56 0.70 0.70 V44D 0.51 0.18
[0627] The invention includes a protease variant having proteolytic activity, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 and at least one substitution selected from those listed in Table 12-9, wherein positions of the variant are numbered by correspondence with positions of the SEQ ID NO:2 sequence. Each such protease variant may be an isolated, recombinant, substantially pure, or non-naturally occurring protease variant. Also included are compositions, including cleaning compositions, comprising at least one such protease variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
Example 13
Liquid Laundry Detergent Compositions
[0628] In this Example, various formulations for liquid laundry detergent compositions are provided. The following liquid laundry detergent compositions of the present invention are prepared as shown below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00056 TABLE 13-1 Liquid Laundry Detergent Composition Formulations Compound I II III IV V LAS 24.0 32.0 6.0 3.0 6.0 NaC.sub.16-C.sub.17 HSAS -- -- -- 5.0 -- C.sub.12-C.sub.15 AE.sub.1.8S -- -- 8.0 7.0 5.0 C.sub.8-C.sub.10 propyl dimethyl 2.0 2.0 2.0 2.0 1.0 amine C.sub.12-C.sub.14 alkyl dimethyl -- -- -- -- 2.0 amine oxide C.sub.12-C.sub.15 AS -- -- 17.0 -- 8.0 CFAA -- 5.0 4.0 4.0 3.0 C.sub.12-C.sub.14 Fatty alcohol 12.0 6.0 1.0 1.0 1.0 ethoxylate C.sub.12-C.sub.18 Fatty acid 3.0 -- 4.0 2.0 3.0 Citric acid (anhydrous) 4.5 5.0 3.0 2.0 1.0 DETPMP -- -- 1.0 1.0 0.5 Monoethanolamine 5.0 5.0 5.0 5.0 2.0 Sodium hydroxide -- -- 2.5 1.0 1.5 1N HCl aqueous solution #1 #1 -- -- -- Propanediol 12.7 14.5 13.1 10. 8.0 Ethanol 1.8 2.4 4.7 5.4 1.0 DTPA 0.5 0.4 0.3 0.4 0.5 Pectin Lyase -- -- -- 0.005 -- Amylase 0.001 0.002 -- -- Cellulase -- -- 0.0002 0.0001 Lipase 0.1 -- 0.1 -- 0.1 NprE (optional) 0.05 0.3 -- 0.5 0.2 PMN -- -- 0.08 -- -- Protease A (optional) -- -- -- -- 0.1 Aldose Oxidase -- -- 0.3 -- 0.003 ZnCl2 0.1 0.05 0.05 0.05 0.02 Ca formate 0.05 0.07 0.05 0.06 0.07 DETBCHD -- -- 0.02 0.01 -- SRP1 0.5 0.5 -- 0.3 0.3 Boric acid -- -- -- -- 2.4 Sodium xylene sulfonate -- -- 3.0 -- -- Sodium cumene -- -- -- 0.3 0.5 sulfonate DC 3225C 1.0 1.0 1.0 1.0 1.0 2-butyl-octanol 0.03 0.04 0.04 0.03 0.03 Brightener 1 0.12 0.10 0.18 0.08 0.10 Balance to 100% perfume/dye and/or water #1: Add 1N HCl aqueous solution to adjust the neat pH of the formula in the range from about 3 to about 5.
[0629] The pH of Formulations (I)-(II) in Table 13-1 is about 5 to about 7 and of Formulations (III)-(V) in Table 13-1 is about 7.5 to about 8.5.
Example 14
Hand Dish Liquid Detergent Compositions
[0630] In this Example, various hand dish liquid detergent formulations are provided. The following hand dish liquid detergent compositions of the present invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00057 TABLE 14-1 Hand Dish Liquid Detergent Compositions Formulations Compound I II III IV V VI C.sub.12-C.sub.15AE.sub.1.8S 30.0 28.0 25.0 -- 15.0 10.0 LAS -- -- -- 5.0 15.0 12.0 Paraffin Sulfonate -- -- -- 20.0 -- -- C.sub.10-C.sub.18 Alkyl Dimethyl Amine 5.0 3.0 7.0 -- -- -- Oxide Betaine 3.0 -- 1.0 3.0 1.0 -- C.sub.12 poly-OH fatty acid amide -- -- -- 3.0 -- 1.0 C.sub.14 poly-OH fatty acid amide -- 1.5 -- -- -- -- C.sub.11E.sub.9 2.0 -- 4.0 -- -- 20.0 DTPA -- -- -- -- 0.2 -- Tri-sodium Citrate dehydrate 0.25 -- -- 0.7 -- -- Diamine 1.0 5.0 7.0 1.0 5.0 7.0 MgCl.sub.2 0.25 -- -- 1.0 -- -- nprE (optional) 0.02 0.01 -- 0.01 -- 0.05 PMN -- -- 0.03 -- 0.02 -- Protease A (optional) -- 0.01 -- -- -- -- Amylase 0.001 -- -- 0.002 -- 0.001 Aldose Oxidase 0.03 -- 0.02 -- 0.05 -- Sodium Cumene Sulphonate -- -- -- 2.0 1.5 3.0 PAAC 0.01 0.01 0.02 -- -- -- DETBCHD -- -- -- 0.01 0.02 0.01 Balance to 100% perfume/dye and/or water
[0631] The pH of Formulations (I)-(VI) in Table 14-1 is about 8 to about 11.
Example 15
Liquid Automatic Dishwashing Detergent Compositions
[0632] In this Example, various liquid automatic dishwashing detergent formulations are provided. The following hand dish liquid detergent compositions of the present invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00058 TABLE 15-1 Liquid Automatic Dishwashing Detergent Compositions Formulations Compound I II III IV V STPP 16 16 18 16 16 Potassium Sulfate -- 10 8 -- 10 1,2 propanediol 6.0 0.5 2.0 6.0 0.5 Boric Acid -- -- -- 4.0 3.0 CaCl.sub.2 dihydrate 0.04 0.04 0.04 0.04 0.04 Nonionic 0.5 0.5 0.5 0.5 0.5 nprE (optional) 0.1 0.03 -- 0.03 -- PMN -- -- 0.05 -- 0.06 Protease B (optional) -- -- -- 0.01 -- Amylase 0.02 -- 0.02 0.02 -- Aldose Oxidase -- 0.15 0.02 -- 0.01 Galactose Oxidase -- -- 0.01 -- 0.01 PAAC 0.01 -- -- 0.01 -- DETBCHD -- 0.01 -- -- 0.01 Balance to 100% perfume/dye and/or water
Example 16
Granular and/or Tablet Laundry Compositions
[0633] This Example provides various formulations for granular and/or tablet laundry detergents. The following laundry compositions of present invention, which may be in the form of granules or tablet, are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00059 TABLE 16-1 Granular and/or Tablet Laundry Compositions Formulations Compound I II III IV V Base Product C.sub.14-C.sub.15AS or TAS 8.0 5.0 3.0 3.0 3.0 LAS 8.0 -- 8.0 -- 7.0 C.sub.12-C.sub.15AE.sub.3S 0.5 2.0 1.0 -- -- C.sub.12-C.sub.15E.sub.5 or E.sub.3 2.0 -- 5.0 2.0 2.0 QAS -- -- -- 1.0 1.0 Zeolite A 20.0 18.0 11.0 -- 10.0 SKS-6 (dry add) -- -- 9.0 -- -- MA/AA 2.0 2.0 2.0 -- -- AA -- -- -- -- 4.0 3Na Citrate.cndot.2H.sub.2O -- 2.0 -- -- -- Citric Acid (Anhydrous) 2.0 -- 1.5 2.0 -- DTPA 0.2 0.2 -- -- -- EDDS -- -- 0.5 0.1 -- HEDP -- -- 0.2 0.1 -- PB1 3.0 4.8 -- -- 4.0 Percarbonate -- -- 3.8 5.2 -- NOBS 1.9 -- -- -- -- NACA OBS -- -- 2.0 -- -- TAED 0.5 2.0 2.0 5.0 1.00 BB1 0.06 -- 0.34 -- 0.14 BB2 -- 0.14 -- 0.20 -- Anhydrous Na Carbonate 15.0 18.0 -- 15.0 15.0 Sulfate 5.0 12.0 5.0 17.0 3.0 Silicate -- 1.0 -- -- 8.0 nprE (optional) 0.03 -- 0.1 0.06 -- PMN -- 0.05 -- -- 0.1 Protease B (optional) -- 0.01 -- -- -- Protease C (optional) -- -- -- 0.01 -- Lipase -- 0.008 -- -- -- Amylase 0.001 -- -- -- 0.001 Cellulase -- 0.0014 -- -- -- Pectin Lyase 0.001 0.001 0.001 0.001 0.001 Aldose Oxidase 0.03 -- 0.05 -- -- PAAC -- 0.01 -- -- 0.05 Balance to 100% Moisture and/or Minors* *Perfume, dye, brightener/SRP1/Na carboxymethylcellulose/photobleach/MgSO.sub.4/PVPVI/suds suppressor/high molecular PEG/clay.
Example 17
Liquid Laundry Detergents
[0634] This Example provides various formulations for liquid laundry detergents. The following liquid laundry detergent formulations of the present invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00060 TABLE 17-1 Liquid Laundry Detergents Formulations Compound I II III IV V VI LAS 11.5 11.5 9.0 -- 4.0 -- C.sub.12-C.sub.15AE.sub.2.85S -- -- 3.0 18.0 -- 16.0 C.sub.14-C.sub.15E.sub.2.5S 11.5 11.5 3.0 -- 16.0 -- C.sub.12-C.sub.13E.sub.9 -- -- 3.0 2.0 2.0 1.0 C.sub.12-C.sub.13E.sub.7 3.2 3.2 -- -- -- -- CFAA -- -- -- 5.0 -- 3.0 TPKFA 2.0 2.0 -- 2.0 0.5 2.0 Citric Acid 3.2 3.2 0.5 1.2 2.0 1.2 (Anhydrous) Ca formate 0.1 0.1 0.06 0.1 -- -- Na formate 0.5 0.5 0.06 0.1 0.05 0.05 ZnCl2 0.1 0.05 0.06 0.03 0.05 0.05 Na Culmene 4.0 4.0 1.0 3.0 1.2 -- Sulfonate Borate 0.6 0.6 1.5 -- -- -- Na Hydroxide 6.0 6.0 2.0 3.5 4.0 3.0 Ethanol 2.0 2.0 1.0 4.0 4.0 3.0 1,2 3.0 3.0 2.0 8.0 8.0 5.0 Propanediol Mono- 3.0 3.0 1.5 1.0 2.5 1.0 ethanolamine TEPAE 2.0 2.0 -- 1.0 1.0 1.0 nprE 0.03 0.05 -- 0.03 -- 0.02 (optional) PMN -- -- 0.01 -- 0.08 -- Protease A -- -- 0.01 -- -- -- (optional) Lipase -- -- -- 0.002 -- -- Amylase -- -- -- -- 0.002 -- Cellulase -- -- -- -- -- 0.0001 Pectin Lyase 0.005 0.005 -- -- -- Aldose 0.05 -- -- 0.05 -- 0.02 Oxidase Galactose -- 0.04 oxidase PAAC 0.03 0.03 0.02 -- -- -- DETBCHD -- -- -- 0.02 0.01 -- SRP 1 0.2 0.2 -- 0.1 -- -- DTPA -- -- -- 0.3 -- -- PVNO -- -- -- 0.3 -- 0.2 Brightener 1 0.2 0.2 0.07 0.1 -- -- Silicone 0.04 0.04 0.02 0.1 0.1 0.1 antifoam Balance to 100% perfume/dye and/or water
Example 18
High Density Dishwashing Detergents
[0635] This Example provides various formulations for high density dishwashing detergents. The following compact high density dishwashing detergents of the present invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00061 TABLE 18-1 High Density Dishwashing Detergents Formulations Compound I II III IV V VI STPP -- 45.0 45.0 -- -- 40.0 3Na Citrate.cndot.2H.sub.2O 17.0 -- -- 50.0 40.2 -- Na Carbonate 17.5 14.0 20.0 -- 8.0 33.6 Bicarbonate -- -- -- 26.0 -- -- Silicate 15.0 15.0 8.0 -- 25.0 3.6 Metasilicate 2.5 4.5 4.5 -- -- -- PB1 -- -- 4.5 -- -- -- PB4 -- -- -- 5.0 -- -- Percarbonate -- -- -- -- -- 4.8 BB1 -- 0.1 0.1 -- 0.5 -- BB2 0.2 0.05 -- 0.1 -- 0.6 Nonionic 2.0 1.5 1.5 3.0 1.9 5.9 HEDP 1.0 -- -- -- -- -- DETPMP 0.6 -- -- -- -- -- PAAC 0.03 0.05 0.02 -- -- -- Paraffin 0.5 0.4 0.4 0.6 -- -- nprE (optional) 0.072 0.053 -- 0.026 -- 0.01 PMN -- -- 0.053 -- 0.059 -- Protease B -- -- -- -- -- 0.01 (optional) Amylase 0.012 -- 0.012 -- 0.021 0.006 Lipase -- 0.001 -- 0.005 -- -- Pectin Lyase 0.001 0.001 0.001 -- -- -- Aldose Oxidase 0.05 0.05 0.03 0.01 0.02 0.01 BTA 0.3 0.2 0.2 0.3 0.3 0.3 Polycarboxylate 6.0 -- -- -- 4.0 0.9 Perfume 0.2 0.1 0.1 0.2 0.2 0.2 Balance to 100% Moisture and/or Minors* *Brightener/dye/SRP1/Na carboxymethylcellulose/photobleach/MgSO.sub.4/PVPVI/suds suppressor/high molecular PEG/clay.
[0636] The pH of Formulations (I) through (VI) in Table 18-1 is from about 9.6 to about 11.3.
Example 19
Tablet Detergent Compositions
[0637] This Example provides various tablet detergent formulations. The following tablet detergent compositions of the present invention are prepared by compression of a granular dishwashing detergent composition at a pressure of 13KN/cm.sup.2 using a standard 12 head rotary press. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00062 TABLE 19-1 Tablet Detergent Compositions Formulations Compound I II III IV V VI VII VIII STPP -- 48.8 44.7 38.2 -- 42.4 46.1 46.0 3Na Citrate.cndot.2H.sub.2O 20.0 -- -- -- 35.9 -- -- -- Na Carbonate 20.0 5.0 14.0 15.4 8.0 23.0 20.0 -- Silicate 15.0 14.8 15.0 12.6 23.4 2.9 4.3 4.2 Lipase 0.001 -- 0.01 -- 0.02 -- -- -- Protease B 0.01 -- -- -- -- -- -- -- (optional) Protease C -- -- -- -- -- 0.01 -- -- (optional) nprE (optional) 0.01 0.08 -- 0.04 -- 0.023 -- 0.05 PMN -- -- 0.05 -- 0.052 -- 0.023 -- Amylase 0.012 0.012 0.012 -- 0.015 -- 0.017 0.002 Pectin Lyase 0.005 -- -- 0.002 -- -- -- -- Aldose Oxidase -- 0.03 -- 0.02 0.02 -- 0.03 -- PB1 -- -- 3.8 -- 7.8 -- -- 4.5 Percarbonate 6.0 -- -- 6.0 -- 5.0 -- -- BB1 0.2 -- 0.5 -- 0.3 0.2 -- -- BB2 -- 0.2 -- 0.5 -- -- 0.1 0.2 Nonionic 1.5 2.0 2.0 2.2 1.0 4.2 4.0 6.5 PAAC 0.01 0.01 0.02 -- -- -- -- -- DETBCHD -- -- -- 0.02 0.02 -- -- -- TAED -- -- -- -- -- 2.1 -- 1.6 HEDP 1.0 -- -- 0.9 -- 0.4 0.2 -- DETPMP 0.7 -- -- -- -- -- -- -- Paraffin 0.4 0.5 0.5 0.5 -- -- 0.5 -- BTA 0.2 0.3 0.3 0.3 0.3 0.3 0.3 -- Polycarboxylate 4.0 -- -- -- 4.9 0.6 0.8 -- PEG 400-30,000 -- -- -- -- -- 2.0 -- 2.0 Glycerol -- -- -- -- -- 0.4 -- 0.5 Perfume -- -- -- 0.05 0.2 0.2 0.2 0.2 Balance to 100% Moisture and/or Minors* *Brightener/SRP1/Na carboxymethylcellulose/photobleach/MgSO.sub.4/PVPVI/suds suppressor/high molecular PEG/clay.
[0638] The pH of Formulations (I) through (VII) in Table 19-1 is from about 10 to about 11.5 and the pH of Formulation (VIII) in Table 19-1 is from 8-10. The tablet weight of Formulations (I) through (VIII) in Table 19-1 is from about 20 grams to about 30 grams.
Example 20
Liquid Hard Surface Cleaning Detergents
[0639] This Example provides various formulations for liquid hard surface cleaning detergents. The following liquid hard surface cleaning detergent compositions of the present invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00063 TABLE 20-1 Liquid Hard Surface Cleaning Detergents Formulations Compound I II III IV V VI VII C.sub.9-C.sub.11E.sub.5 2.4 1.9 2.5 2.5 2.5 2.4 2.5 C.sub.12-C.sub.14E.sub.5 3.6 2.9 2.5 2.5 2.5 3.6 2.5 C.sub.7-C.sub.9E.sub.6 -- -- -- -- 8.0 -- -- C.sub.12-C.sub.14E.sub.21 1.0 0.8 4.0 2.0 2.0 1.0 2.0 LAS -- -- -- 0.8 0.8 -- 0.8 Sodium culmene sulfonate 1.5 2.6 -- 1.5 1.5 1.5 1.5 Isachem .RTM. AS 0.6 0.6 -- -- -- 0.6 -- Na.sub.2CO.sub.3 0.6 0.13 0.6 0.1 0.2 0.6 0.2 3Na Citrate.cndot.2H.sub.2O 0.5 0.56 0.5 0.6 0.75 0.5 0.75 NaOH 0.3 0.33 0.3 0.3 0.5 0.3 0.5 Fatty Acid 0.6 0.13 0.6 0.1 0.4 0.6 0.4 2-butyl octanol 0.3 0.3 -- 0.3 0.3 0.3 0.3 PEG DME-2000 .RTM. 0.4 -- 0.3 0.35 0.5 -- -- PVP 0.3 0.4 0.6 0.3 0.5 -- -- MME PEG (2000) .RTM. -- -- -- -- -- 0.5 0.5 Jeffamine .RTM. ED-2001 -- 0.4 -- -- 0.5 -- -- PAAC -- -- -- 0.03 0.03 0.03 -- DETBCHD 0.03 0.05 0.05 -- -- -- -- nprE (optional) 0.07 -- 0.08 0.03 -- 0.01 0.04 PMN -- 0.05 -- -- 0.06 -- -- Protease B (optional) -- -- -- -- -- 0.01 -- Amylase 0.12 0.01 0.01 -- 0.02 -- 0.01 Lipase -- 0.001 -- 0.005 -- 0.005 -- Pectin Lyase 0.001 -- 0.001 -- -- -- 0.002 ZnCl2 0.02 0.01 0.03 0.05 0.1 0.05 0.02 Calcium Formate 0.03 0.03 0.01 -- -- -- -- PB1 -- 4.6 -- 3.8 -- -- -- Aldose Oxidase 0.05 -- 0.03 -- 0.02 0.02 0.05 Balance to 100% perfume/dye and/or water
The pH of Formulations (I) through (VII) in Table 20-1 is from about 7.4 to about 9.5.
Example 21
Cleaning Performance of BPN'-v36 Polypeptide Variants
[0640] BPN'-v36 polypeptide variants comprising two amino acid substitutions were constructed by standard PCR fusion using the BPN'-v36 variant as a backbone or parent sequence. For this purpose, two or three partially overlapping fragments were amplified by mutagenic primers prepared such that the primer encoded a desired substitution. PCR amplification reactions were carried out as described in Example 7 of Part I supra. The following BPN'-v36 double mutant variants (i.e., BPN'-v36 with the following two amino acid substitution) were constructed: Q019R-N025D, A001Y-Q275R, V004A-S249N, V004E-S260P, V004A-T55A, Y006F-S249C, Y006D-T55A, V008L-Q275R, Q010R-Q275K, L016Q-Q217H, H017R-T158A, S183D-Q206R, P210S-N212D, S018Y-V203A, S018K-V203I, Y021H-D259G, Y021H-D259R, K027R-N269D, K027R-N269T, S037P-S260F, S037T-S260P, D041E-N077D, D041G-N077E, G166V-S183T, N252S-L257H, V044A-Q206H, V044A-Q206K, V044A-Q206R, N076T-N212D, N076P-N212S, N077D-N252D, N077D-N252T, K141I-S248N, T158I-D259N, T158A-D259P, S161E-Q185H, K237M-H238R, G160A-D259G, G160R-D259V, G215R-D259R, G215D-D259V, N061D-Q206R, N061L-Q206H, S009L-N218S, S161E-S260T, Q019A-N109S, T022S-G166V, Y021H-N252H, P129S-K136R, T022S-T242S, N025K-H238R, N025D-Q185R, S037G-Q275H, K043R-N076S, K043N-Q217R, K043N-S163T, T055A-V147A, N061K-N252K, N062Y-G097D, Y021H-V084E, Y021H-S037E, N062Y-T244A, K027E-Y091F, A074S-P129Q, S249R-Q275R, I079V-Q217H, A098T-T158A, K027R-D120H, Q019R-Q185R, G131S-K265N, A133V-D259N, A144H-T244A, I035V-K043N, G160R-T244A, S161P-T253A, S163T-Q245L, K170R-D259G, S183T-S249R, N184Y-Y262N, V198L-D259G, A200T-H226L, Q206R-S260P, G211V-T244A, Q217R-T244A, L751-N76D, S260P-Q275L, S260P-Q275R, Y262N-Q275R, V004A-Y006F, H017L-Q019A, N025D-V026A, N118G-V121A, V072F-L075I, S183T-R186K, V203A-Q217R, and S249R-Y262H. The cleaning performance of each of these variants was tested in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C. and egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C. as described in Example 1 of Part I. Results are provided below.
[0641] The following BPN' protease variants were determined to have a PI value equal to or greater than 0.9 and equal or less than 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of S183T-S249R, N61D-Q206R, Y262N-Q275R, K43R-N765, K170R-D259G, Y6F-S249C, Q19A-N109S, H17L-Q19A, Q19R-Q185R, S18Y-V203A, S161E-S260T, S18K-V203I, V4A-T55A, N252S-L257H, S249R-Y262H, N61L-Q206H, N184Y-Y262N, Q19R-N25D, A74S-P129Q, K27R-D120H, Y21H-N252H, K27R-N269D, A98T-T158A, 179V-Q217H, S9L-N218S, V4A-Y6F, S161P-T253A, V203A-Q217R, T22S-T242S, N76P-N212S, S37T-S260P, T55A-V147A, G160R-T244A, N25D-Q185R, G211V-T244A, A144H-T244A, Y21H-N252H, A1Y-Q275R, V198L-D259G, K141I-S248N, S183T-R186K, S161E-Q185H, P129S-K136R, K43N-S163T, S37G-Q275H, N62Y-T244A, and S260P-Q275R, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to BPN' (SEQ ID NO:2) and a greater PI value than that of BPN' in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), a PI value of equal to or greater than 0.9 and equal to or less than 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0642] Also provided is a subtilisin protease variant having enhanced proteolytic activity and a PI value of greater than 1.0 to about 5 compared to BPN' in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:6, wherein the variant comprises at least one substitution comprising at least one substitution selected from the group of X1Y, X4A, X6F, X9L, X17L, X18K/Y, X19A/R, X21H, X22S, X25D, X27R, X37G/T, X43N/R, X55A, X61D/L, X62Y, X74S, X76P/S, X79V, X98T, X109S, X120H, X129Q/S, X136R, X141I, X144H, X147A, X158A, X160R, X161E/P, X163T, X170R, X183T, X184Y, X185H/R, X186K, X198L, X203A/I, X206H/R, X211V, X212S, X217H/R, X218S, X242S, X244A, X248N, X249C/R, X252H/S, X253A, X257H, X259G, X260P/T, X262H/N, X269D, and X275H/R, and optionally at least one substitution selected from the group of A1Y, V4A, Y6F, S9L, H17L, S18K/Y, Q19A/R, Y21H, T22S, N25D, K27R, S37G/T, K43N/R, T55A, N61D/L, N62Y, A74S, N76P/S, I79V, A98T, N109S, D120H, P129Q/S, K136R, K141I, A144H, V147A, T158A, G160R, S161E/P, S163T, K170R, S183T, N184Y, Q185H/R, R186K, V198L, V203A/I, Q206H/R, G211V, N212S, Q217H/R, N218S, T242S, T244A, S248N, S249C/R, N252H/S, T253A, L257H, D259G, S260P/T, Y262H/N, N269D, and Q275H/R, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN' (SEQ ID NO:2) and a PI value greater than that of BPN' in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0643] The following BPN' subtilisin protease variants were determined to have a PI value equal or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of Y21H-D259G, A133V-D259N, 179V-Q217H, 518K-V203I, T158A-D259P, N61K-N252K, K43N-Q217R, T158A-D259P, Q206R-S260P, A133V-D259N, V198L-D259G, N61K-N252K, S161E-S260T, G160A-D259G, K43N-Q217R, A1Y-Q275R, A200T-H226L, Q217R-T244A, S260P-Q275R, Q206R-S260P, T158I-D259N, Q217R-T244A, L751-N76D, S161E-Q185H, Y21H-S37E, S249R-Q275R, T158I-D259N, Y21H-S37E, N76T-N212D, S260P-Q275L, G131S-K265N, V4A-S249N, N25D-Q185R, K43R-N765, S183D-Q206R, Q10R-Q275K, K43N-S163T, Q10R-Q275K, N25D-V26A, G131S-K265N, S260P-Q275L, K141I-S248N, L16Q-Q217H, S249R-Q275R, K27R-N269T, P210S-N212D, L751-N76D, S183D-Q206R, N118G-V121A, G215D-D259V, N76T-N212D, K27R-N269T, N62Y-G97D, V4E-S260P, G215D-D259V, K27E-Y91F, Y6D-T55A, N77D-N252T, V4E-S260P, Y6D-T55A, N25K-H238R, V44A-Q206H, L16Q-Q217H, V44A-Q206R, V44A-Q206H, and S37P-S260F, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, having enhanced proteolytic activity greater than that of BPN', and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0644] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of Y21H-D259G, S183T-S249R, N61D-Q206R, Y262N-Q275R, K043R-N076S, A133V-D259N, and I079V-Q217H, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0645] Also provided is a subtilisin protease variant having enhanced proteolytic activity and/or a PI value of greater than 1.0 to about 5 compared to BPN' in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:6, wherein the variant comprises at least one substitution comprising at least one substitution selected from the group of X021H, X043R, X061D, X076S, X079V, X133V, X183T, X206R, X217H, X249R, X259G/N, X262N, and X275R, and optionally at least one substitution selected from the group of Y021H, K043R, N061D, N076S, I079V, A133V, S183T, Q206R, Q217H, S249R, D259G/N, Y262N, and Q275R, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN' (SEQ ID NO:2) and a PI value greater than that of BPN' in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0646] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than or equal to 1.0 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of K170R-D259G, S18K-V203I, Y6F-S249C, Q19A-N109S, H17L-Q19A, Q19R-Q185R, S18Y-V203A, N61D-Q206R, S161E-S260T, S18K-V203I, V4A-T55A, N252S-L257H, S249R-Y262H, N61L-Q206H, N184Y-Y262N, Q19R-N25D, S249R-Y262H, A74S-P129Q, T158A-D259P, H17L-Q19A, K27R-D120H, V4A-T55A, N61K-N252K, Y21H-N252H, K27R-N269D, K43N-Q217R, T158A-D259P, Q206R-S260P, K27R-N269D, A98T-T158A, 179V-Q217H, S9L-N218S, V4A-Y6F, S161P-T253A, V203A-Q217R, T22S-T242S, N76P-N212S, A133V-D259N, S37T-S260P, T55A-V147A, V198L-D259G, Q19R-Q185R, V4A-Y6F, Q19A-N109S, Y262N-Q275R, G160R-T244A, Q19R-N25D, N25D-Q185R, N61K-N252K, S161E-S260T, A98T-T158A, N61L-Q206H, G211V-T244A, S9L-N218S, A144H-T244A, A144H-T244A, 518Y-V203A, Y21H-N252H, A74S-P129Q, G160A-D259G, K43N-Q217R, A1Y-Q275R, A1Y-Q275R, A200T-H226L, Q217R-T244A, S260P-Q275R, Q206R-S260P, K141I-S248N, S183T-R186K, T158I-D259N, S37T-S260P, K27R-D120H, T22S-T242S, Q217R-T244A, S161E-Q185H, P129S-K136R, G211V-T244A, N76P-N212S, L751-N76D, S161E-Q185H, Y21H-S37E, S249R-Q275R, G160A-D259G, K43N-S163T, T158I-D259N, Y21H-S37E, S37G-Q275H, S161P-T253A, N76T-N212D, S260P-Q275L, Y6F-S249C, N184Y-Y262N, G131S-K265N, V4A-S249N, N25D-Q185R, N252S-L257H, K43R-N765, S183D-Q206R, G160R-T244A, Q10R-Q275K, S37G-Q275H, K43N-S163T, Q10R-Q275K, N25D-V26A, P129S-K136R, G131S-K265N, S260P-Q275L, K141I-S248N, T22S-G166V, N62Y-T244A, L16Q-Q217H, S249R-Q275R, S260P-Q275R, K27R-N269T, P210S-N212D, L751-N76D, S183D-Q206R, N118G-V121A, G215D-D259V, N76T-N212D, V4A-S249N, K27R-N269T, G166V-S183T, N62Y-G97D, V4E-S260P, G215D-D259V, K27E-Y91F, Y21H-D259R, Y6D-T55A, N77D-N252T, V4E-S260P, Y6D-T55A, N77D-N252T, Y21H-D259R, N25K-H238R, N77D-N252D, V44A-Q206H, L16Q-Q217H, V72F-L751, S37P-S260F, V72F-L751, N77D-N252D, V44A-Q206R, S163T-Q245L, V44A-Q206H, V44A-Q206R, S37P-S260F, G215R-D259R, V44A-Q206K, and V44A-Q206K, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and equal to or less than 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
Example 22
Cleaning Performance of Additional BPN'-v36 Polypeptide Variants
[0647] The following BPN'-v36 variants were synthesized at DNA2.0 (Menlo Park, Calif.) using the pHPLT-BPN'-v36 plasmid containing the BPN' expression cassette served as template DNA (parent plasmid) for cloning: N109G-A128S-S224A, N109G-A128S-S224A-N243V, A88T-N109G-A116T-A128S-S224A-N243V, N61G-N109G-A128S-S224A, N61G-N109G-A128S-S224A-N243V, N61G-A88T-N109G-A116T-A128S-S224A-N243V, N109Q-A128S-S224A, N109Q-A128S-S224A-N243V, A88T-N109Q-A116T-A128S-S224A-N243V, N109S-A128S-S224A, N109S-A128S-S224A-N243V, A88T-N109S-A116T-A128S-S224A-N243V, N109M-A128S-S224A, N109M-A128S-S224A-N243V, A88T-N109M-A116T-A128S-S224A-N243V, N109G-A114S-A128S, N109G-A114S-A128S-N243V, A88T-N109G-A114S-A116T-A128S-N243V, N109G-A114S-A128S-S224A, N109G-A114S-A128S-S224A-N243V, N109G-A128S-S183V, N109G-A128S-S183L, N109G-A128S-S183L-S224A, N109G-A114S-A128S-S183L-S224A, A88T-N109G-A114S-A116T-A128S-S183L-S224A-N243V, N76D-N109G-A128S-S224A, N101Q-N109Q-A128S-S224A-N243V, N101Q-N109Q-A128S-P129S-S130T-S224A-N243V, N109G-A128S-P129S-S130T-S224A-N243V, S33T-A128S-N218S, S33T-N109G-A128S-N218S-N243V, S33T-N61G-N109G-A128S-N218S-N243V, S33T-N109G-A128S-G169A-N218S-N243V, S33T-S63G-N109G-A128S-N218S-N243V, S33T-N76D-N109G-A128S-N218S-N243V, S33T-S63G-N109G-A128S-G169A-N218S-N243V, I31L-S33T-S63G-N109G-A128S-G169A-N218S-N243V, S33T-N61G-S63G-N109G-A128S-N218S-N243V, S33T-N61G-A88T-N109G-A116T-A128S-N218S-N243V, S33T-N61G-S63G-N109G-A128S-G131H-G169A-N218S-N243V, S33T-N61P-S63G-N109G-A128S-G131H-G169A-N218S-N243V, S33T-N109G-A128S-N218S-S224A-N243V, S63G-N109Q-A128S-S224A-N243V, S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-S224A-N243V, A1G-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, I31L-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-S224A-N243V-S249Q, S33T-T55P-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V-S249Q, A1G-I31L-S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V-S24- 9Q, N61P-S63G-N109Q-A128S-G131H-G169A-S224A-N243V-S249Q, and A1G-I31L-S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-G169A-S224A-N24- 3V-S249Q.
[0648] The variants were grown for protein expression as described in Example 11 of Part I. These variants were tested for their performance in the BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., and the AAPF assay as described in Example 1 of Part I. Results are provided below.
[0649] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of N61P-S63G-N109Q-A128S-S224A-N243V, A88T-N109G-A114S-A116T-A128S-N243V, A88T-N109G-A114S-A116T-A128S-S183L-S224A-N243V, N109G-A128S-S183V, N109G-A128S-N243V-K256R, N109M-A128S-S224A, A88T-N109S-A116T-A128S-S224A-N243V, N109Q-A128S-S224A-N243V, A88T-N109M-A116T-A128S-S224A-N243V, N109S-A128S-S224A-N243V, A88T-N109G-A116T-N243V, N101Q-N109Q-A128S-S224A-N243V, N109G-A116T-N243V-K256R, N109G-A128S-P129S-S130T-S224A-N243V, and A88T-N109Q-A116T-A128S-S224A-N243V, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0650] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN'-v36 and/or a PI value of equal to or greater than 1.0 to about 5 compared to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NON:6, wherein the variant comprises at least one substitution comprising at least one substitution selected from the group of X61G/P/S, X63G, X88T, X101Q, X109G/M/Q/S, X114S, X116T, X128S, X129S, X130T, X158S, X183L/V, X224A, X243V, X248A, and X256R, and optionally at least one substitution selected from the group of N61G/P/S, S63G, A88T, N101Q, N109G/M/Q/S, A114S, A116T, A128S, P129S, S130T, T158S, S183L/V, S224A, N243V, S248A, and K256R, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN' (SEQ ID NO:2) BPN'-v3, and BPN'-v36 and a PI value greater than that of BPN', BPN'-v3, and BPN'-v36 in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0651] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.9 and equal to or less than 1.0 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of G24S-G53S-N78S-G97A-N101S-A128S, G24S-G53S-N78S-G97A-N101S, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V, S33T-N61G-S63G-N109G-A128S-N218S-N243V, S33T-S63G-N109G-A128S-N218S-N243V, S33T-T55P-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-G169A-S224A-N243V-S249Q, S33T-N61G-A88T-N109G-A116T-A128S-N218S-N243V, S33T-N109G-A128S-N218S-N243V, S33T-N76D-N109G-A128S-N218S-N243V, S33T-N76D-N109G-A128S-N218S-N243V-S248N-K256R, S33T-N61G-N109G-A128S-N218S-N243V, S33T-A128S-N218S, A1G-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-S224A-N243V-S249Q, N61P-S63G-N109Q-A128S-G131H-S224A-N243V, S63G-N109Q-A128S-G131H-S224A-N243V, N109G-A114S-A128S, N109G-A114S-A128S-S183L-S224A, N109G-A114S-A128S-S224A, N109G-A114S-A128S-S224A-N243V, A88T-N109G-A116T-A128S-S224A-N243V, N61G-A88T-N109G-A116T-A128S-S224A-N243V, N109G-A114S-A128S-N243V, N109G-A128S-S224A-N243V, N109G-A128S-S224A, N109G-A128S-S183L-S224A, N61G-N109G-A128S-S224A, N109G-A128S-S183L, S33T-N76D, N109S-A128S-S224A, N101Q-N109Q-A128S-P129S-S130T-S224A-N243V, S63G-N109Q-A128S-S224A-N243V, N109M-A128S-S224A-N243V, S63G-N109G, N109G-K256R, S63G-N76D, S33T-N109G-A128S-G169A-N218S-N243V, and S33T-N109G-A128S-N218S-S224A-N243V, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity, enhanced proteolytic activity compared to BPN', and/or a PI value equal to or greater than 0.9 and less or equal to 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0652] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in a BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of I31L-S33T-S63G-N109G-A128S-G169A-N218S-N243V, A1G-I31L-S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-G169A-S224A-N24- 3V-S249Q, S33T-N61G-S63G-N109G-A128S-G131H-G169A-N218S-N243V, S33T-S63G-N109G-A128S-G169A-N218S-N243V, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V-S249Q, S33T-N76D-A128S-N218S, N76D-N109G-A128S-S224A, and S33T-N61P-S63G-N109G-A128S-G131H-G169A-N218S-N243V, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0653] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V, S33T-N61G-S63G-N109G-A128S-N218S-N243V, S33T-S63G-N109G-A128S-N218S-N243V, N61P-S63G-N109Q-A128S-G131H-G169A-S224A-N243V-S249Q, S33T-N61G-A88T-N109G-A116T-A128S-N218S-N243V, S33T-N109G-A128S-N218S-N243V, S33T-A128S-N218S, A1G-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-S224A-N243V-S249Q, S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-S224A-N243V, A88T-N109G-A114S-A116T-A128S-N243V, A88T-N109G-A114S-A116T-A128S-S183L-S224A-N243V, N109G-A114S-A128S, N109G-A114S-A128S-S183L-S224A, N109G-A114S-A128S-S224A, N109G-A114S-A128S-S224A-N243V, A88T-N109G-A116T-A128S-S224A-N243V, N61G-A88T-N109G-A116T-A128S-S224A-N243V, N109G-A128S-S183V, N109G-A114S-A128S-N243V, N109G-A128S-N243V-S248A, N109G-A128S-S224A-N243V, N109G-A128S-N243V-K256R, N109G-A128S-S224A, N109G-A128S-S183L-S224A, N61G-N109G-A128S-S224A, N109M-A128S-S224A, A88T-N109S-A116T-A128S-S224A-N243V, N109M-A128S-S224A-N243V, S63G-A128S, A88T-N109G-A116T-N243V, N101Q-N109Q-A128S-S224A-N243V, N109G-A116T-N243V-K256R, N109G-A116T, S63G-N109G, A88T-N109G, N109G-K256R, N61G-N109G-N243V, S33T-N109G-A128S-G169A-N218S-N243V, S33T-N109G-A128S-N218S-S224A-N243V, N109G-A128S-P129S-S130T-S224A-N243V, and A88T-N109Q-A116T-A128S-S224A-N243V, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0654] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN'-v36 and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v36 in an egg microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C., the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:6, wherein the variant comprises at least one substitution comprising at least one substitution selected from the group of X1G, X33T, X55P, X61G/P/S, X63G, X76D, X88T, X101Q, X109G/M/Q/S, X114S, X116T, X128S, X129S, X130T, X131H, X158S, X169A, X183L/V, X218S, X224A, X243V, X248A/N, X249Q, X256R, and optionally at least one substitution selected from the group of A1G, S33T, T55P, N61G/P/S, S63G, N76D, A88T, N101Q, N109G/M/Q/S, A114S, A116T, A128S, P129S, S130T, G131H, T158S, G169A, S183L/V, N218S, S224A, N243V, S248A/N, S249Q, K256R, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN', BPN'-v3, and BPN'-v36 and a PI value greater than that of BPN', BPN'-v3, and BPN'-v36 in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0655] The following BPN'-v36 variant was determined to have a PI value equal to or greater than 0.5 and equal to or less than 1.0 relative to BPN'-v36 in an egg BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising amino acid substitutions selected from the group consisting of substitutions S33T-N76D-A128S-N218S, N76D-N109G-A128S-S224A and S063G-N76D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and equal to or less than 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising amino acid substitutions selected from the group consisting of S33T-N76D-A128S-N218S, N76D-N109G-A128S-S224A, and S063G-N076D, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0656] The following BPN'-v36 variants were determined to have a PI value greater than 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from greater than 1.0 to about 10, from greater than 1.0 to about 8, or from greater than 1.0 to about 5 relative to BPN'-v36 in an AAPF proteolytic assay: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of G24S-G53S-N78S-G97A-N101S-A128S, I31L-S33T-S63G-N109G-A128S-G169A-N218S-N243V, A1G-I31L-S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-G169A-S224A-N24- 3V-S249Q, S33T-N61G-S63G-N109G-A128S-G131H-G169A-N218S-N243V, S33T-S63G-N109G-A128S-G169A-N218S-N243V, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V, S33T-T55P-N61P-S63G-A88T-N109G-A116T-A128S-G131H-S224A-N243V-S249Q, S33T-N61G-S63G-N109G-A128S-N218S-N243V, S33T-S63G-N109G-A128S-N218S-N243V, S33T-T55P-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-G169A-S224A-N243V-S249Q, S33T-N61G-A88T-N109G-A116T-A128S-N218S-N243V, S33T-N109G-A128S-N218S-N243V, S33T-N76D-N109G-A128S-N218S-N243V, S33T-N76D-N109G-A128S-N218S-N243V-S248N-K256R, S33T-N61G-N109G-A128S-N218S-N243V, S33T-N76D-A128S-N218S, S33T-A128S-N218S, A1G-N61P-S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-G131H-S224A-N243V-S249Q, N61P-S63G-N109Q-A128S-G131H-S224A-N243V, S63G-N109Q-A128S-G131H-S224A-N243V, N61P-S63G-N109Q-A128S-S224A-N243V, A88T-N109G-A114S-A116T-A128S-N243V, A88T-N109G-A114S-A116T-A128S-S183L-S224A-N243V, N109G-A114S-A128S, N109G-A114S-A128S-S183L-S224A, N109G-A114S-A128S-S224A, N109G-A114S-A128S-S224A-N243V, A88T-N109G-A116T-A128S-S224A-N243V, N61G-A88T-N109G-A116T-A128S-S224A-N243V, N109G-A128S-S183V, N109G-A114S-A128S-N243V, N109G-A128S-N243V-S248A, N109G-A128S-S224A-N243V, N109G-A128S-N243V-K256R, N109G-A128S-S224A, N109G-A128S-S183L-S224A, N61G-N109G-A128S-S224A, N76D-N109G-A128S-S224A, N109M-A128S-S224A, N109G-A128S-S183L, S33T-N76D, A88T-N109S-A116T-A128S-S224A-N243V, N109Q-A128S-S224A-N243V, N109S-A128S-S224A, A88T-N109M-A116T-A128S-S224A-N243V, N101Q-N109Q-A128S-P129S-S130T-S224A-N243V, S63G-N109Q-A128S-S224A-N243V, N109M-A128S-S224A-N243V, S63G-A128S, N109S-A128S-S224A-N243V, A88T-N109G-A116T-N243V, N61S-N109G-N243V, N101Q-N109Q-A128S-S224A-N243V, N109G-A116T-N243V-K256R, A88T-N109G-A116T-T158S-N243V-K256R, N109G-A116T, S63G-N109G, A88T-N109G, N109G-K256R, N61G-N109G-N243V, S33T-N61P-S63G-N109G-A128S-G131H-G169A-N218S-N243V, S33T-N109G-A128S-G169A-N218S-N243V, S33T-N109G-A128S-N218S-S224A-N243V, N109G-A128S-P129S-S130T-S224A-N243V, and A88T-N109Q-A116T-A128S-S224A-N243V, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN', BPN'-v3, and BPN'-v36, and a greater PI value than that of BPN', BPN'-v3 and BPN'-v36 in this assay. The invention includes a protease variant having enhanced proteolytic activity compared to BPN' (SEQ ID NO:2), enhanced proteolytic activity compared to BPN', BPN'-v3, and BPN'-v36, a PI value of greater than 1.0 to about 5 relative to BPN'-v3, and/or a PI value of greater than 1.0 to about 5 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of amino acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0657] Also provided is a subtilisin protease variant having enhanced proteolytic activity compared to BPN'-v36 and/or a PI value of greater than 1.0 to about 5 compared to BPN'-v36 in An AAPF proteolytic assay, the variant comprising an amino acid sequence having at least 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to SEQ ID NO:2 or SEQ ID NO:6, wherein the variant comprises at least one substitution comprising at least one substitution selected from the group of X1G, X31L, X33T, X55P, X61G/P/S, X63G, X76D, X88T, X101Q, X109G/M/Q/S, X114S, X116T, X128S, X129S, X130T, X131H, X158S, X169A, X183L/V, X218S, X224A, X243V, X248A/N, X249Q, and X256R, and optionally at least one substitution selected from the group of A1G, I31L, S33T, T55P, N61G/P/S, S63G, N76D, A88T, N101Q, N109G/M/Q/S, A114S, A116T, A128S, P129S, S130T, G131H, T158S, G169A, S183L/V, N218S, S224A, N243V, S248A/N, S249Q, and K256R, wherein amino acid positions of the variant are numbered by correspondence with positions of the sequence of SEQ ID NO:2. Such variants have enhanced proteolytic activity compared to the BPN' (SEQ ID NO:2) BPN'-v3, and BPN'-v36 and a PI value greater than that of BPN', BPN'-v3, and BPN'-v36 in this assay. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
[0658] The following BPN'-v36 variant was determined to have a PI value equal to or greater than 0.5 and equal to or less than 1.0 relative to BPN'-v36 in an AAPF proteolytic assay: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising amino acid substitutions G24S-G53S-N78S-G97A-N101S or S063G-N76D, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and equal to or less than 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising substitutions G24S-G53S-N78S-G97A-N101S or S063G-N76D, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
Example 23
Cleaning Performance of Polypeptide BPN'-v36 Polypeptides Variants from Combinatorial Libraries Based on BPN'-v36 Polypeptide
[0659] Two separate combinatorial libraries (AJ1 and AJ2) were synthesized by DNA2.0 (Menlo Park, Calif.) and were delivered as individual ligation reactions. The pHPLT-BPN'-v36 plasmid (FIG. 4) containing the BPN' expression cassette served as template DNA (parent plasmid) for library construction. A list of the possible amino acid positions and substitutions for each library is shown in Table 23-1. The ligation reactions for each library were used to transform B. subtilis, and the library variants were grown up for protein expression as described in Example 11. The variants were tested for performance in the BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C. as described in Example 1 of Part I.
TABLE-US-00064 TABLE 23-1 Possible Substitutions for Combinatorial Libraries AJ1 and AJ2 AJ1 AJ2 Position Possible Substitutions Position Possible Substitutions S33 G, S E54 E, Q D60 D, G D99 D, N N62 N, L, S D120 D, N S63 S, R, L, N, G D140 D, N S125 S, A E156 E, Q Q217 Q, R, E, L, G D197 D, N M222 M, L, S K12 K, T K27 K, S K43 K, T K141 K, Y K213 K, Q K237 K, A K256 K, Q
[0660] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less or equal to 1.0 relative to BPN'-v36 in the BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of G024S-G053S-N078S-G097A-N101S (i.e., BPN'-v3), G024S-G053S-N078S-G097A-N101S-A128S (i.e., BPN'-v12), N062L, N062L-S063G, N062S, N062S-S063G-Q217L, N062S-S063L-Q217L, N062S-S063N, N062S-S063R, Q217E, S063G, S063G-Q217L, S063G-Q217L-M222S, S063L-Q217L, S063N, S063N-Q217L, D099N-K141Y-K213Q, D099N-K141Y-K256Q, K043T, K043T-K141Y-E156Q, N062L-Q217E, N062L-Q217L, N062L-S063G-Q217E, N062L-S063L, N062L-S063N-Q217L, N062S-Q217L, N062S-S063G, N062S-S063L, N062S-S063N-Q217L, N062S-S063R-Q217E, Q217L, S063G-Q217E, S063N-Q217E, S063R, S063R-Q217E, S063R-Q217L,), D099N-K141Y-K213Q, D099N-K141Y-K256Q, K043T, K043T-K141Y-E156Q, N062L-Q217E, N062L-Q217L, N062L-S063G-Q217E, N062L-S063L, N062S-Q217L, N062S-S063G, N062S-S063L, N062S-S063N-Q217L, Q217L, S063G-Q217E, S063N-Q217E, S063R, S063R-Q217E, and S063R-Q217L, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and equal to or less than 1.0 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, or 98% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein. Note that a protease variant which is BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising the amino acid substitutions G024S-G053S-N078S-G097A-N101S is BPN'-v3 (SEQ ID NO:4), because the G at position 24, G at position 53, N at position 78, and N at position 101 of SEQ ID NO:6 have been substituted with S at each of positions 24, 53, 78, and 101 of SEQ ID NO:6. In addition, G at position 97 of SEQ ID NO:6 has been substituted with A. Thus, the resultant sequence is SEQ ID NO:4.
[0661] The following BPN'-v36 variants were determined to have a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in the BMI microswatch cleaning assay in Detergent Composition 4 at pH 8 and 16.degree. C.: BPN'-S024G-S053G-S078N-S101N-G128A-Y217Q amino acid sequence (SEQ ID NO:6) comprising at least one set of amino acid substitutions selected from the group consisting of D099N, K141Y-E156Q, N062L-S063L-Q217L, N062L-S063N, N062L-S063N-Q217E, N062L-S063R, N062L-S063R-Q217L, N062S-Q217E, N062S-S063G-Q217E, N062S-S063G-Q217R, N062S-S063N-Q217R, S063G-S125A, D060G-Q217L, D120N-K141Y-K213Q, K043T-D099N-D120N-K141Y, K043T-D099N-K141Y-K256Q, K043T-K237A, N062L-S063G-Q217R, N062L-S063G-S125A, N062L-S063L-Q217E, N062L-S063N-S125A-Q217L, N062S-Q217R, N062S-S063L-Q217E, N062S-S063R-Q217L, S063G-M222S, S063G-Q217R, D120N-E156Q-K256Q, K141Y-D197N, N062L-Q217R, N062L-S063G-Q217L-M222S, N062L-S063L-Q217R, N062L-S063N-Q217R, N062S-Q217G, N062S-S063G-Q217G, N062S-S063G-Q217L-M222L, N062S-S063G-S125A-Q217L, N062S-S063N-Q217E, Q217G, S033G-N062S-S063G, S063G-Q217G, S063G-Q217L-M222L, S063G-S125A-Q217R, S063L-Q217R, S063N-M222S, S063N-Q217R, S063N-S125A-Q217L, S063R-Q217R, S063R-S125A-Q217L, D099N-E156Q-K256Q, E156Q, K012T-D099N-K213Q, K012T-K256Q, K043T-D099N-K141Y-K213Q, K043T-E156Q, K141Y-K213Q, N062L-Q217G, N062L-Q217L-M222L, N062L-Q217L-M222S, N062L-S063G-M222S, N062L-S063G-Q217L-M222L, N062L-S063G-Q217R-M222S, N062L-S063N-Q217L-M222S, N062L-S063N-S125A, N062L-S063R-S125A, N062L-S125A, N062S-S063G-M222S, N062S-S063G-Q217G-M222S, N062S-S063G-S125A, N062S-S063N-Q217L-M222L, N062S-S063N-S125A-Q217L, N062S-S063R-Q217G, N062S-S063R-Q217L-M222S, Q217G-M222S, Q217L-M222S, Q217R, S033G-S063G-Q217R, S063G-Q217E-M222S, S063G-S125A-Q217G, S063L-Q217E, S063N-Q217G, S063N-Q217G-M222S, S063N-Q217L-M222S, S063R-Q217L-M222S, and S063R-S125A, wherein amino acid positions of the variant are numbered by correspondence with the sequence of SEQ ID NO:2. Such variants have proteolytic activity. The invention includes a protease variant having proteolytic activity and/or a PI value equal to or greater than 0.5 and less than 0.9 relative to BPN'-v36 in this assay, the variant comprising an amino acid sequence having at least 60%, 70%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity to the sequence of SEQ ID NO:2 or SEQ ID NO:6 and comprising at least one set of acid substitutions selected from said group above, wherein amino acid positions of the variant are numbered by correspondence with amino acid positions of the SEQ ID NO:2 sequence. Also included are compositions, including, but not limited to, e.g., cleaning compositions, comprising at least one such variant and methods for cleaning utilizing at least one such protease variant as described in greater detail elsewhere herein.
Part II
Series I GG36 Cold Water Protease Variants
[0662] The amino acid sequence of wild-type mature Bacillus lentus GG36 protease is:
TABLE-US-00065 (SEQ ID NO: 755) AQSVPWGISRVQAPAAHNRGLTGSGVKVAVLDTGISTHPDLNIRGGASFV PGEPSTQDGNGHGTHVAGTIAALNNSIGVLGVAPSAELYAVKVLGASGSG SVSSIAQGLEWAGNNGMHVANLSLGSPSPSATLEQAVNSATSRGVLVVAA SGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGLDIVAPGVNVQ STYPGSTYASLNGTSMATPHVAGAAALVKQKNPSWSNVQIRNHLKNTATS LGSTNLYGSGLVNAEAATR
[0663] As indicated herein, suitable Series I GG36 cold water protease variants include enzyme variants derived from a parent protease, said parent protease's sequence being at least 60%, 70%, 75%, 80%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 100% identical to the amino acid sequence of SEQ ID NO:755, said protease variant having one or more of the following characteristics: [0664] a) Test Method 2 performance index of at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from 1.1 to about 10, from 1.1 to about 8, or even from 1.1 to about 5; [0665] b) a Test Method 3 performance index of at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from 1.1 to about 10, from 1.1 to about 8, or even from 1.1 to about 5; [0666] c) a Test Method 4 performance index of at least 1.0, at least 1.1, at least 1.2, at least 1.3, at least 1.4, at least 1.5, at least 1.6, at least 1.7, at least 1.8, at least 1.9, at least 2, from 1.0 to about 10, from 1.0 to about 8, or even from 1.0 to about 5.
[0667] Test Method 2, Test Method 3, and Test Method 4 are explicitly described in the "Test Methods" section of Part II Example 1. All mutations referenced herein utilize the BPN' numbering scheme as shown in FIG. 5. In some aspects, the variants referenced herein refer to variants having amino acid sequences compared to the amino acid sequence of SEQ ID NO:755, using the BPN' numbering scheme.
[0668] Suitable Series I GG36 cold water proteases can be variants of subtilisins or derived from subtilisins, particularly those derived from subtilisin Bacillus lentus GG36 of SEQ ID NO:755 and in some aspects, comprise one or more of the following mutations: A1R, Q2S, V4R, V4S, S9A, R10S, P14K, A16S, H17R, N18R, G20R, T22A, T22R, S24R, S24W, G25R, G25V, V26F, L421, N43R, N43A, G46R, P52F, P52E, P52N, T57R, Q59A, N62E, N62Q, V68A, V68C, T71G, I72C, A74C. L75A, L75F, L75R, N76D, S78R, L82R, P86W, E89P, E89T, E89G, E89H, E89I, E89V, E89W, Y91N, K94N, G100S, S101A, S101N, S101G, S101D, S103G, S103N, V104L, V104I, S106V, S106G, A108I, L111V, E112V, G115K, G115R, N117F, G118I, V121F, S128D, S128F, S128L, S128N, P129E, S144R, L148I, A158E. G159E, S160D, S166D, N185E, N185I, R186H, S188E, S188D, D197F, V203E, Y209S, Y209N, Y209F, Y209T, Y209E, Y209H, Y209G, P210R, S212I, S212F, Y214F, A215N, A215D, A215E, L217E, L217N, T224A, A230E, A231I, Q236F, N238R, N238K, P239K, P239G, P239R, P239S, W241R, S242R, S242L, N243R, V244R, N248I, N248V, H249R, L250I, N252R, T253R, L262D, Y263F, S265F, L267V, L267N. N269I, N269R, E271F, E271I, E271H, E271P, E271T, E271V, E271L and/or A272F, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 (BPN') as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2.
[0669] In one aspect, suitable Series I GG36 cold water protease variants include subtilisins, particularly those derived from Bacillus lentus GG36 of SEQ ID NO:755, comprising one or more of the following sets of substitutions: T022R-S024R, S009A-E271L, N018R-W241R, N018R-G115R, N043R-H249R, G020R-H249R, V004R-H249R, G020R-S024R, N018R-H249R, S009A-G020R, G020R-W241R, S009A-S078R, G020R-G115R, N018R-S024R, S024R-S242R, T022R-G115R, N018R-N043R, G020R-N043R, N018R-S242R, S242R-N269R, N018R-V244R, S024R-N269R, G020R-E271L, S024R-E271L, V004R-S009A, G020R-N269R, A001R-S024R, V244R-E271L, S009A-N018R, W241R-E271L, V004R-S024R, S009A-H249R, S009A-T022R, N062E-P129E, N062E-G159E, A016S-L148I, A158E-H249R, A016S-N062E, L111V-S188D, T022A-N062E, N062E-L148I, T022A-P129E, N062E-E271F, N062E-A158E, A016S-G159E, N062E-R186H, S128N-G159E, N062E-S188D, N062E-S128N, L148I-G159E, S103G-A158E, L111V-G159E, A158E-E271F, A016S-S188D, T022A-L111V, S128N-A158E, A016S-A158E, V104L-A158E, S128N-R186H, G159E-Y209E, N062E-S101A, L111V-Y209E, L148I-S188D, S101A-Y209E, T022A-S188D, A016S-T022A, S128N-P129E, A016S-Y209E, A016S-S128N, T022A-E089P, S128N-Y209E, E089P-A158E, N062E-S103G, R186H-E271F, A016S-P129E, E089P-G159E, L111V-H249R, S101A-P129E, L148I-Y209E, T022A-G159E, P129E-H249R, P129E-Y209E, V104L-P129E, S128N-S188D, L111V-A158E, T022A-A158E, N062E-Y209E, N062E-H249R, S101A-R186H, E089P-P129E, P129E-E271, T22A-L111V-G159E, S101A-S103G-V104L-Y209E, S101A-S103G-V104L-G159E, S101A-S103G-V104L-S188D, S101G-S103A-V104I-G159D, T22A-S103G-G159E, T22A-S128N-E271F-Y209E, T22A-Y209E-E271F, T22A-S101A-Y209E, S101A-Y209E-E271F, T22A-L111V-S128N, T22A-S101A-G159E, S101A-S103G-V104L, T22A-S101A-S103G-V104L, S101A-S103G-V104L, S101G-S103A-V104I, S101A-S103G-V104L-S128N, S103A-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N248D, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-S24R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-T253R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159E-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N238R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N76D, and/or S101G-S103A-V104I-G159E-A232V-Q245R-N248D-E271F, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 (BPN') as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2.
[0670] In another aspect, suitable Series I GG36 cold water protease variants include variants of subtilisins, particularly those derived from Bacillus lentus GG36 of SEQ ID NO:755, wherein the variants comprise three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, or even 25 mutations within the group of positions comprising positions 1, 2, 4, 9, 10, 14, 16, 17, 18, 20, 22, 24, 25, 26, 42, 43, 46, 52, 57, 59, 62, 68, 71, 72, 74, 75, 76, 78, 82, 86, 89, 91, 94, 100, 101, 103, 104, 106, 108, 111, 112, 115, 117, 118, 121, 128, 129, 144, 148, 158, 159, 160, 166, 185, 186, 188, 197, 203, 209, 210, 212, 214, 215, 217, 224, 230, 231, 236, 238, 239, 241, 242, 243, 244, 248, 249, 250, 252, 253, 262, 263, 265, 267, 269, 271 and 272.
[0671] In another aspect, suitable Series I GG36 cold water protease variants include variants of subtilisins, particularly those derived from Bacillus lentus GG36 of SEQ ID NO:755, wherein the variants comprise a total of three, four, five, six, seven, eight, nine, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, or even 25 mutations selected from: A1R, Q2S, V4R, V4S, S9A, R10S, P14K, A165, H17R, N18R, G20R, T22A, T22R, S24R, S24W, G25R, G25V, V26F, L421, N43R, N43A, G46R, P52F, P52E, P52N, T57R, Q59A, N62E, N62Q, V68A, V68C, T71G, I72C, A74C. L75A, L75F, L75R, N76D, S78R, L82R, P86W, E89P, E89T, E89G, E89H, E89I, E89V, E89W, Y91N, K94N, G100S, S101A, S101N, S101G, S101D, S103G, S103N, V104L, V104I, S106V, S106G, A108I, Lilly, E112V, G115K, G115R, N117F, G118I, V121F, S128D, S128F, S128L, S128N, P129E, S144R, L148I, A158E. G159E, S160D, S166D, N185E, N185I, R186H, S188E, S188D, D197F, V203E, Y209S, Y209N, Y209F, Y209T, Y209E, Y209H, Y209G, P210R, S212I, S212F, Y214F, A215N, A215D, A215E, L217E, L217N, T224A, A230E, A231I, Q236F, N238R, N238K, P239K, P239G, P239R, P239S, W241R, S242R, S242L, N243R, V244R, N248I, N248V, H249R, L250I, N252R, T253R, L262D, Y263F, S265F, L267V, L267N, N269I, N269R, E271F, E271I, E271H, E271P, E271T, E271V, E271L and A272F; and optionally one or more of the following mutations: S103A, G159D, Q236H, Q245R, N248D and N252K, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 (BPN') as determined by alignment of the amino acid sequence of the variant with SEQ ID NO:2.
[0672] In some aspects, the Series I GG36 cold water protease variant comprises one or more mutations, and having a total net charge of -5, -4, -3, -2, -1 or 0 relative to B. lentus subtilisin GG36 wild-type (SEQ ID NO:755).
[0673] In another aspect, the Series I GG36 cold water protease variants are low ionic strength Series I GG36 cold water protease variants. Such low ionic strength Series I GG36 cold water protease variants comprising one or more mutations, and having a total net charge of -5, -4, -3, -2, -1 or 0 relative to B. lentus subtilisin GG36 protease wild-type (SEQ ID NO:755). These mutations can be selected from: (a) two or more of the following mutations: A1R, Q2S, V4R, V4S, S9A, R10S, P14K, A16S, T22A, T22R, S24R, G25V, V26F, L42I, P52F, P52E, P52N, N62E, N62Q, V68A, V68C, T71G, I72C, A74C. L75A, L75F, 578R, E89P, E89T, E89G, E89H, E89W, Y91N, K94N, G100S, S101A, S101N, S101G, S101D, S103G, S103N, V104L, V104I, A108I, L111V, E112V, G115K, N117F, V121F, S128D, S128F, S128L, S128N, P129E, L148I, A158E. G159E, S160D, S166D, N185E, R186H, S188E, S188D, V203E, Y209S, Y209N, Y209F, Y209T, Y209E, Y209H, Y209G, P210R, S212I, S212F, Y214F, A215N, A215D, A215E, L217E, L217N, T224A, A230E, A231I, Q236F, N238R, N238K, P239K, P239G, P239R, N248V, H249R, L250I, L262D, Y263F, S265F, L267V, L267N. N269I, N269R, E271F, E271I, E271H and A272F; and/or (b) one or more of the following sets of mutations: N062E-P129E, N062E-G159E, A016S-L148I, A158E-H249R, A016S-N062E, L111V-S188D, T022A-N062E, N062E-L148I, T022A-P129E, N062E-E271F, N062E-A158E, A016S-G159E, N062E-R186H, S128N-G159E, N062E-S188D, N062E-S128N, L148I-G159E, S103G-A158E, L111V-G159E, A158E-E271F, A016S-S188D, T022A-L111V, S128N-A158E, A016S-A158E, V104L-A158E, S128N-R186H, G159E-Y209E, N062E-S101A, L111V-Y209E, L148I-S188D, S101A-Y209E, T022A-S188D, A016S-T022A, S128N-P129E, A016S-Y209E, A016S-S128N, T022A-E089P, S128N-Y209E, E089P-A158E, N062E-S103G, R186H-E271F, A016S-P129E, E089P-G159E, L111V-H249R, S101A-P129E, L148I-Y209E, T022A-G159E, P129E-H249R, P129E-Y209E, V104L-P129E, S128N-S188D, L111V-A158E, T022A-A158E, N062E-Y209E, N062E-H249R, S101A-R186H, E089P-P129E, P129E-E271F, T22A-L111V-G159E, S101A-S103G-V104L-Y209E, S101A-S103G-V104L-G159E, S101A-S103G-V104L-S188D, S101G-S103A-V104I-G159D, T22A-S103G-G159E, T22A-S128N-E271F-Y209E, T22A-Y209E-E271F, T22A-S101A-Y209E, S101A-Y209E-E271F, T22A-L111V-S128N, T22A-S101A-G159E, S101A-S103G-V104L, T22A-S101A-S103G-V104L, S101A-S103G-V104L, S101G-S103A-V104I, S101A-S103G-V104L-S128N, S103A-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-G159D-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-Q236H-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q245R-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-N248D-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N248D, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-S24R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-T253R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-H249R, T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159E-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N238R, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N76D and S101G-S103A-V104I-G159E-A232V-Q245R-N248D-E271F, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 (BPN').
[0674] In another aspect, the above low ionic strength Series I GG36 cold water protease variants form part of a detergent composition that is diluted in water, typically within a laundry washing machine, to form a laundry detergent wash liquor, whose conductivity is from about 0.1 mS/cm to about 3 mS/cm, from about 0.3 mS/cm to about 2.5 mS/cm, or even from about 0.5 mS/cm to about 2 mS/cm.
[0675] In another aspect, the Series I GG36 cold water protease variants are high ionic strength Series I GG36 cold water protease variants. Such high ionic strength Series I GG36 cold water protease variants comprise two or more mutations, and have a total net charge of +5, +4, +3, +2, +1 or 0 relative to B. lentus subtilisin GG36 protease wild-type (SEQ ID NO:755). These mutations can be selected from: (a) two or more of the following mutations V4R, H17R, N18R, G20R, T22R, S24R, S24W, G25R, N43R, N43A, G46R, P52F, P52N, T57R, Q59A, N62Q, T71G, L75R, N76D, S78R, L82R, P86W, E89P, E89W, E89T, E89I, E89H, E89V, V104L, S106V, S106G, G115R, G118I, V121F, S144R, N185I, D197F, Y209N, Y209S, L217E, A231I, P239R, P239S, W241R, S242R, S242L, N243R, V244R, N248I, H249R, N252R, T253R, E271T, E271V, E271L, E271H, E271F, E271P, A1R, S9A, S212F and N269R; and/or (b) one or more of the following sets of mutations T022R-S024R, S009A-E271L, N018R-W241R, N018R-G115R, N043R-H249R, G020R-H249R, V004R-H249R, G020R-S024R, N018R-H249R, S009A-G020R, G020R-W241R, S009A-S078R, G020R-G115R, N018R-S024R, S024R-S242R, T022R-G115R, N018R-N043R, G020R-N043R, N018R-S242R, S242R-N269R, N018R-V244R, S024R-N269R, G020R-E271L, S024R-E271L, V004R-S009A, G020R-N269R, A001R-S024R, V244R-E271L, S009A-N018R, W241R-E271L, V004R-S024R, S009A-H249R, S009A-T022R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-E271F, S101G-S103A-V104I-A158E-A232V-Q245R-N248D-H249R, S101G-S103A-V104I-G159D-A232V-Q245R-N248D-S24R, S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K, S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K, wherein the variant has proteolytic activity and each amino acid position of the variant is numbered by correspondence to an amino acid position in the amino acid sequence of SEQ ID NO:2 (BPN').
[0676] In another aspect, the above high ionic strength Series I GG36 cold water protease variants form part of a detergent composition that is diluted in water, typically within a laundry washing machine, to form a laundry detergent wash liquor, whose conductivity is from about 3 mS/cm to about 30 mS/cm, from about 3.5 mS/cm to about 20 mS/cm, or even from about 4 mS/cm to about 10 mS/cm.
[0677] The charge of the Series I GG36 cold water protease variants is expressed relative to B. lentus subtilisin GG36 protease wild-type having the amino acid sequence of SEQ ID NO:755. The amino acids that impart a single negative charge are D and E and those that impart a single positive charge are R, H and K. Any amino acid change versus SEQ ID NO:755 that changes a charge is used to calculate the charge of the Series I GG36 cold water protease variant. For example, introducing a negative charge mutation from a wild-type neutral position will add a net charge of -1 to the Series I GG36 cold water protease variant, whereas introducing a negative charge mutation (D or E) from a wild-type positive amino acid residue (R, H or K) will add a net charge of -2. Summing the charge changes from all the amino acid residues that are different for the Series I GG36 cold water protease variant versus B. lentus subtilisin GG36 protease wild-type having the amino acid sequence of SEQ ID NO:755 gives the charge change of the Series I GG36 cold water protease variant. Without wishing to be bound by theory, it is believed that: (a) the preferred charge range for Series I GG36 cold water protease variants to be used in low conductivity laundry detergent solutions is -5, -4, -3, -2, -1, 0, particularly -2, -1; and (b) the preferred charge range for Series I GG36 cold water protease variants to be used in high conductivity laundry detergent solutions is +5, +4, +3, +2, +1, 0, particularly +2, +1. By correctly selecting the charge unexpectedly improved levels of cold water cleaning performance can be obtained. "Low conductivity laundry detergent solutions" are defined as having a conductivity of from about 0.1 mS/cm to about 3 mS/cm, from about 0.3 mS/cm to about 2.5 mS/cm, or even from about 0.5 mS/cm to about 2 mS/cm. "High conductivity laundry detergent solutions" are defined as having a conductivity of from about 3 mS/cm to about 30 mS/cm, from about 3.5 mS/cm to about 20 mS/cm, or even from about 4 mS/cm to about 10 mS/cm. It is intended that the above examples be non-limiting. Once mutations are combined to optimize cold water performance, the enzyme charge can also be balanced by mutations in further positions.
Methods for Making GG36 Cold Water Protease Variants
[0678] A variety of methods are known in the art that are suitable for generating modified polynucleotides of the invention that encode GG36 cold water protease variants, including, but not limited to, for example, site-saturation mutagenesis, scanning mutagenesis, insertional mutagenesis, deletion mutagenesis, random mutagenesis, site-directed mutagenesis, and directed-evolution, as well as various other recombinatorial approaches. Non-limiting exemplary methods of making the Series I GG36 cold water protease variants are provided in the section above entitled "Vectors, Cells, and Methods for Making Protease Variant Polypeptides of the Invention."
[0679] For testing of enzyme activity in heat-inactivated detergents, working solutions of detergents are made from the heat inactivated stocks. Appropriate amounts of water hardness (e.g., 6 gpg or 12 gpg) and buffer are added to the detergent solutions to match the desired conditions. The solutions are mixed by vortexing or inverting the bottles. The following Table provides information regarding some of the commercially-available detergents and test conditions used herein. In some experiments, additional and/or other commercially available detergents find use in the following Examples.
TABLE-US-00066 TABLE A Laundry and Dish Washing Conditions Region Form Dose Detergent* Buffer Gpg pH T (.degree. C.) Laundry (Heavy Duty Liquid and Granular) NA HDL 0.78 g/l P&G TIDE .RTM. 2X 5 mM HEPES 6 8.0 20 WE HDL 5.0 g/L Henkel PERSIL .TM. 5 mM HEPES 12 8.2 40 WE HDG 8.0 g/L P&G ARIEL .RTM. 2 mM Na.sub.2 CO.sub.3 12 10.5 40 JPN HDG 0.7 g/L P&G TIDE .RTM. 2 mM Na.sub.2 CO.sub.3 6 10.0 20 NA HDG 1.0 g/L P&G TIDE .RTM. 2 mM Na.sub.2 CO.sub.3 6 10.0 20 Automatic Dish Washing WE ADW 3.0 g/L RB CALGONIT .TM. 2 mM Na.sub.2 CO.sub.3 21 10.0 40 NA ADW 3.0 g/L P&G CASCADE .RTM. 2 mM Na.sub.2 CO.sub.3 9 10.0 40
In some additional Examples, the following solutions find use:
TABLE-US-00067 TABLE B Working Detergent Solutions Temp Detergent Detergent (C) g/L pH Buffer Gpg TIDE .RTM. 2X Cold 16 0.98 8 5 mM 6 HEPES TIDE .RTM. 2X Cold 32 0.98 8 5 mM 6 HEPES TIDE .RTM. 2X Cold 16 0.98 7 5 mM 6 MOPS
[0680] Table C provides granular laundry detergent compositions produced in accordance with the invention suitable for laundering fabrics.
TABLE-US-00068 TABLE C Granular Laundry Detergent Compositions and Their Components Detergent Compositions Component 1 2 3 4 5 6 Linear alkylbenzenesulfonate 15 12 20 10 12 13 with aliphatic carbon chain length C.sub.11-C.sub.12 Other surfactants 1.6 1.2 1.9 3.2 0.5 1.2 Phosphate builder(s) 2 3 4 Zeolite 1 1 4 1 Silicate 4 5 2 3 3 5 Sodium Carbonate 2 5 5 4 0 3 Polyacrylate (MW 4500) 1 0.6 1 1 1.5 1 Carboxymethyl cellulose 1 -- 0.3 -- 1.1 -- (Finnfix BDA ex CPKelco) Celluclean .RTM. (15.6 mg/g) 0.23 0.17 0.5 0.2 0.2 0.6 Cold Water Protease variant* 0.23 0.17 0.05 0.2 0.03 0.1 Stainzyme Plus .RTM. (14 mg/g) 0.23 0.17 0.5 0.2 0.2 0.6 Mannaway 4.0T (4 mg/g) 0.1 0.1 0.1 Lipex 100T (18.6 mg/g) 0.2 0.1 0.3 Fluorescent Brightener(s) 0.16 0.06 0.16 0.18 0.16 0.16 Diethylenetriamine 0.6 0.6 0.25 0.6 0.6 pentaacetic acid or Ethylene diamine tetraacetic acid MgSO.sub.4 1 1 1 0.5 1 1 Bleach(es) and Bleach 6.88 6.12 2.09 1.17 4.66 activator(s) Ethoxylated thiophene Hueing 0.002 0.001 0.003 0.003 -- -- Dye.sup.5 Direct Violet 9 ex Ciba 0.0006 0.0004 0.0006 Specialty Chemicals Sulfate/Citric Acid/Sodium Balance to 100% Bicarbonate/ Moisture/perfume .sup.1Random graft copolymer is a polyvinyl acetate grafted polyethylene oxide copolymer having a polyethylene oxide backbone and multiple polyvinyl acetate side chains. The molecular weight of the polyethylene oxide backbone is about 6000 and the weight ratio of the polyethylene oxide to polyvinyl acetate is about 40 to 60 and no more than 1 grafting point per 50 ethylene oxide units. .sup.2Polyethylenimine (MW = 600) with 20 ethoxylate groups per --NH. .sup.3Amphiphilic alkoxylated grease cleaning polymer is a polyethylenimine (MW = 600) with 24 ethoxylate groups per --NH and 16 propoxylate groups per --NH .sup.4Reversible protease inhibitor of structure: ##STR00001## .sup.5Ethoxylated thiophene Hueing Dye is as described in U.S. Pat. No. 7,208,459 B2.
[0681] In Table C, all enzyme levels expressed as % enzyme raw material, except for cold water protease variant (of this invention) which is expressed as % of active protein added to the product. Table D provides granular laundry detergent compositions suitable for top-loading automatic washing machines (detergent compositions 7-9) and front loading washing machines (detergent compositions 10-11). The GG36 protease variant tested and/or cold water protease variant of the present invention is added separately to these formulations.
TABLE-US-00069 TABLE D Granular Laundry Detergent Compositions and Their Components Detergent Composition Component 7 8 9 10 11 Surfactants C.sub.16-17 Branched alkyl sulfate 3.55 15.8 C.sub.12-14 alkyl sulphate 1.5 Sodium linear alkylbenzenesulfonate 9.6 10.6 7.5 9 with aliphatic chain length C.sub.11-C.sub.12 Sodium C.sub.14/15 alcohol ethoxy-3- 1.15 2.88 sulfate Sodium C.sub.14/15 alkyl sulphate 2.37 C.sub.14/15 alcohol ethoxylate with average 7 1.17 1 moles of ethoxylation mono-C.sub.8-10 alkyl mono-hydroxyethyl di- 0.45 methyl quaternary ammonium chloride Di methyl hydroxyl ethyl lauryl 0.18 ammonium chloride Zeolite A 13.9 4.7 0.01 2.9 1.8 Sodium Silicate 1.6.ratio 4 0.2 4 4 Sodium Silicate 2.35.ratio 8 Citric Acid 2.5 1.4 Sodium tripolyphosphate 5 Sodium Carbonate 24.1 30 16.9 24.4 21 Nonanoyloxybenzenesuplhonate 5.78 2.81 0.96 Oxaziridinium-based bleach booster 0.03 0.017 Tetrasodium S,S,- 0.2 ethylenediaminedisuccinate Diethylenetriamine penta (methylene 0.61 0.33 phosphonic acid), heptasodium salt Hydroxyethane dimethylene phosphonic 0.29 0.45 acid Ethylene diamine tetraacetate 0.27 MgSO4 0.47 0.5994 0.782 Sodium Percarbonate 7 4.4 15.9 19.1 Tetra Acetyl Ethylene Diamine 3.3 4.6 Sodium Perborate Monohydrate 1.2 Carboxymethyl cellulose 0.1 0.17 1.69 0.23 (e.g., Finnfix BDA ex CPKelco) Sodium Acrylic acid/maleic acid co- 0.0236 3.8 2 2.5 polymer (70/30) Sodium polyacrylate (Sokalan PA30 CL) 4 0.84 Terephthalate polymer 0.23 Polyethylene glycol/vinyl acetate 0.89 0.89 0.91 random graft co polymer Photobleach-zinc phthalocyanine 0.005 0.001 0.002 tetrasulfonate C.I. Fluorescent Brightener 260 0.11 0.15 0.04 0.23 0.15 C.I. Fluorescent Brightener 351 0.1 (Tinopal .RTM. CBS) Suds suppressor granule 0.25 0.07 0.04 Hydrophobically modified carboxy 0.019 0.028 methyl cellulose (Finnifix .RTM. SH-1) Bentonite 8.35 Miscellaneous (Dyes, perfumes, process Balance Balance Balance Balance Balance aids, moisture and sodium sulphate)
[0682] In Table D, surfactant ingredients can be obtained from any suitable supplier, including but not limited to BASF (e.g., LUTENSOL.RTM.), Shell Chemicals, Stepan, Huntsman, and Clariant (e.g., PRAEPAGEN.RTM.). Zeolite can be obtained from sources such as Industrial Zeolite. Citric acid and sodium citrate can be obtained from sources such as Jungbunzlauer. Sodium percarbonate, sodium carbonate, sodium bicarbonate and sodium sesquicarbonate can be obtained from sources such as Solvay. Acrylate/maleate copolymers can be obtained from sources such as BASF. Carboxymethylcellulose and hydrophobically modified carboxymethyl cellulose can be obtained from sources such as CPKelco. C.I. Fluorescent Brightener 260 can be obtained from 3V Sigma (e.g., OPTIBLANC.RTM., OPTIBLANC.RTM. 2M/G, OPTIBLANC.RTM. 2MG/LT Extra, or OPTIBLANC.RTM. Ecobright. Tetrasodium S,S-ethylenediamine disuccinate can be obtained from sources such as Innospec. Terephthalate co-polymer can be obtained from Clariant (e.g., REPELOTEX SF 2). In addition, 1-Hydroxyethane-1,1-diphosphonic acid can be obtained from Thermphos. Oxaziridinium-based bleach booster has the following structure, where R1=2-butyloctyl, and was produced according to US 2006/0089284A1.
##STR00002##
[0683] The enzymes NATALASE.RTM., TERMAMYL.RTM., STAINZYME PLUS.RTM., CELLUCLEAN.RTM. and MANNAWAY.RTM. can be obtained from Novozymes. Zinc phthalocyanine tetrasulfonate can be obtained from Ciba Specialty Chemicals (e.g., TINOLUX.RTM. BMC). Suds suppressor granule can be obtained from Dow Corning. In these detergent compositions, random graft copolymer is a polyvinyl acetate grafted polyethylene oxide copolymer having a polyethylene oxide backbone and multiple polyvinyl acetate side chains. The molecular weight of the polyethylene oxide backbone is about 6000 and the weight ratio of the polyethylene oxide to polyvinyl acetate is about 40 to 60 and no more than 1 grafting point per 50 ethylene oxide units.
PART II EXAMPLES
Example 1
Assays and Test Methods
[0684] This Example describes the various Test Methods and assays used in the development of the variants described in these Examples. Any deviations from the protocols provided are indicated in the pertinent Examples. The assays were performed using a Biomek FX Robot (Beckman Coulter) or a multichannel pipettor (e.g., Rainin PipetLite, Mettler-Toledo) and a SpectraMAX MTP Reader (type 340; Molecular Devices).
A. Test Methods
Test Method 1
[0685] A protocol to define whether a dye or pigment material is a fabric hueing agent for the purpose of the invention is provided below:
[0686] 1) Fill two tergotometer pots with 800 ml of Newcastle upon Tyne, UK, City Water (.about.12 grains per US gallon total hardness, supplied by Northumbrian Water, Pity Me, Durham, Co. Durham, UK).
[0687] 2) Insert pots into tergotometer, with water temperature controlled at 30.degree. C. and agitation set at 40 rpm for the duration of the experiment.
[0688] 3) Add 4.8 g of IEC-B detergent (IEC 60456 Washing Machine Reference Base Detergent Type B), supplied by wfk, Bruggen-Bracht, Germany, to each pot.
[0689] 4) After two minutes, add 2.0 mg active colorant to the first pot.
[0690] 5) After one minute, add 50 g of flat cotton vest (supplied by Warwick Equest, Consett, County Durham, UK), cut into 5 cm.times.5 cm swatches, to each pot.
[0691] 6) After 10 minutes, drain the pots and re-fill with cold Water (16.degree. C.) having a water hardness of 14.4 English Clark Degrees Hardness with a 3:1 Calcium to Magnesium molar ratio. 7) After 2 minutes rinsing, remove fabrics.
[0692] 8) Repeat steps 3-7 for a further three cycles using the same treatments.
[0693] 9) Collect and line dry the fabrics indoors for 12 hours.
[0694] 10) Analyze the swatches using a Hunter Miniscan spectrometer fitted with D65 illuminant and UVA cutting filter, to obtain Hunter a (red-green axis) and Hunter b (yellow-blue axis) values.
[0695] 11) Average the Hunter a and Hunter b values for each set of fabrics. If the fabrics treated with colorant under assessment show an average difference in hue of greater than 0.2 units on either the a axis or b axis, it is deemed to be a fabric hueing agent for the purpose of the invention.
Test Method 2
[0696] For Test Method 2, the BMI microswatch assay provided below is run using the granular detergent composition 10 (see Table D above). The laundry detergent is dissolved in water that has a hardness of 12 gpg and adjusted to a temperature of 16.degree. C., and the protease variant enzyme of interest is added. Performance of the protease variant enzymes is then determined as per the BMI microswatch assay described. The performance index is determined by comparing the performance of the protease variant enzyme with that of the B. lentus GG36 subtilisin enzyme having the amino acid sequence of SEQ ID NO:755, with in all cases the enzyme dosage being 1.6 ppm. Protease variant enzymes having a performance index of 1.1 or greater are viewed to be cold water protease variants.
Test Method 3
[0697] For Test Method 3, the BMI microswatch assay provided below is run using the granular laundry detergent composition 7 (see Table D above). The laundry detergent is dissolved in water that has a hardness of 6 gpg and adjusted to a temperature of 16.degree. C., the GG36 protease variant enzyme of interest is added. Performance of the GG36 protease variant enzymes is then determined as per the BMI microswatch assay described. The performance index is determined by comparing the performance of the GG36 protease variant enzyme with that of the B. lentus GG36 subtilisin enzyme having the amino acid sequence of SEQ ID NO:755, with in all cases the enzyme dosage being 4 ppm. GG36 protease variant enzymes having a performance index of 1.1 or greater are viewed to be cold water protease variants.
Test Method 4
[0698] For Test Method 4, the BMI microswatch assay provided below is run using the granular laundry detergent composition 7 (see Table D above). The laundry detergent is dissolved in water that has a hardness of 6 gpg and adjusted to a temperature of 16.degree. C., and the GG36 protease variant enzyme of interest is added. Performance of the GG36 protease variant enzymes is then determined as per the BMI microswatch assay described. The performance index is determined by comparing the performance of the GG36 protease variant enzyme with that of a reference enzyme GG36-A158E, said GG36-A158E reference enzyme consisting of the B. lentus subtilisin GG36 protease amino acid sequence of SEQ ID NO:755 with a single substitution of glutamic acid for alanine at position 158 (i.e., the A158E mutation), with in all cases the enzyme dosage being 4 ppm. GG36 protease variant enzymes having a performance index of 1.0 or greater are viewed to be cold water protease variants.
Test Method 5
[0699] Electrical conductivity of an aqueous solution is assayed according to the standard method ASTM D1125 and reported in units of milliSiemens/cm, abbreviated to mS/cm herein.
B. Assays
TCA Assay for Protein Content Determination in 96-Well Microtiter Plates
[0700] For GG36 and GG36 variants, this assay was started using filtered B. subtilis culture supernatants from microtiter plates grown 2-3 days at 37.degree. C. with shaking at 250 rpm and humidified aeration. A fresh 96-well flat bottom microtiter plate (MTP; Costar 9017 medium binding clear polystyrene plate) was used for the assay. First, 100 .mu.L/well of 0.25 N HCl was placed in each well. Then, 20-25 .mu.L of filtered culture supernatant were added and the solution was mixed on a table top mixer (e.g., Lab line Instruments, Titer plate shaker, model 4825) for 5-10 seconds. The light scattering/absorbance at 405 nm was then determined, in order to provide the "blank" reading. For the "test" reading, 100 .mu.L/well of 30% (w/v) trichloroacetic acid (TCA) was added to each well containing the mixture of HCl and culture supernatant, and the plate was incubated for 10 minutes at room temperature. After briefly mixing the solution on a table top mixer for no more than 2-3 sec, the light scattering/absorbance at 405 nm was determined. The turbidity/light scatter increase in the samples correlates to the total amount of precipitable protein in the culture supernatant. The calculations were performed by subtracting the "blank" reading (obtained after addition of HCl only, no TCA) from the "test" reading (obtained after addition of TCA, as described above) to provide a relative measure of the protein content in the samples. If desired, a standard curve can be created by calibrating the TCA readings with AAPF assays of clones with known conversion factors. However, the TCA results are linear with respect to protein concentration from 50 to 500 parts per million (ppm) of protein (where 1 ppm corresponds to 1 mg/L) and can thus be plotted directly against enzyme performance for the purpose of choosing variants with desired performance.
AAPF Protease Assay in 96-well Microtiter Plates
[0701] In order to determine the protease activity of the serine protease variants, the hydrolysis of N-succinyl-L-alanyl-L-alanyl-L-prolyl-L-phenyl-p-nitroanilide (suc-AAPF-pNA) was measured. The reagent solutions used were: 100 mM Tris/HCl, pH 8.6, containing 0.005% TWEEN.RTM.-80 (Tris dilution buffer); 100 mM Tris buffer, pH 8.6, containing 1 mM CaCl.sub.2 and 0.005% TWEEN.RTM.-80 (Tris/Ca buffer); and 160 mM suc-AAPF-pNA in DMSO (suc-AAPF-pNA stock solution) (Sigma: S-7388). To prepare a suc-AAPF-pNA working solution, 1 ml suc-AAPF-pNA stock solution was added to 100 ml Tris/Ca buffer and mixed well for at least 10 seconds. The assay was performed by adding 10 .mu.l of diluted protease solution to each well of a 96-well MTP, immediately followed by the addition of 190 .mu.l of 1 mg/ml suc-AAPF-pNA working solution. The solutions were mixed for 5 sec, and the absorbance change in kinetic mode (25 readings in 5 minutes) was read at 405 nm in an MTP reader, at 25.degree. C. The protease activity was expressed as AU (activity=.DELTA.ODmin.sup.-1 ml.sup.-1).
Eglin C Inhibition Assay
[0702] As described herein, serine protease concentration and specific activity was determined by titration with an inhibitor called eglin c. Eglin c from the leech Hirudo medicinalis is a tight-binding protein inhibitor of subtilisins and ASP protease (Heinz et al., Biochemistry, 31: 8755-66 [1992]), and can therefore be used to measure protease enzyme concentration, which in turn permits specific activity to be calculated. The gene for eglin c was synthesized and expressed in E. coli by standard methods. Its properties and inhibitory potency were the same as eglin c purchased from Sigma.
(i) Concentration Determination of an Eglin C Stock Solution
[0703] A sample of Bacillus lentus subtilisin of known specific activity was diluted in 100 mM Tris buffer, pH 8.6, containing 1 mM CaCl.sub.2) and 0.005% TWEEN.RTM.-80 (Tris/Ca buffer), to a concentration appropriate for AAPF protease assay described above. Several dilutions of the eglin c stock solution were also made in the Tris/Ca buffer. An aliquot of each diluted eglin c solution was mixed with an equal volume of the diluted Bacillus lentus subtilisin solution. An aliquot of the Tris/Ca buffer only, without eglin c, was also mixed with an equal volume of the diluted Bacillus lentus subtilisin solution, in order to measure uninhibited subtilisin activity in the absence of eglin c. The mixed solutions were incubated at room temperature for 15-30 minutes and the protease activity of each sample was then measured by AAPF assay described above. Using the known specific activity of Bacillus lentus subtilisin, the concentration of active protease in each sample was determined. The concentration of eglin c in each sample was then calculated based on the decrease of the observed protease activity as compared to the uninhibited subtilisin sample that was mixed with Tris/Ca buffer only (without eglin c). Thus, using the known dilutions and volumes of the eglin c solutions, the concentration of eglin c in the stock solution was determined.
(ii) Concentration and Specific Activity Determination of Subtilisin Variants
[0704] Samples of subtilisin variants were diluted in 100 mM Tris buffer, pH 8.6, containing 1 mM CaCl.sub.2) and 0.005% TWEEN.RTM.-80 (Tris/Ca buffer). Several dilutions of the eglin c stock solution of known concentration were also made in the Tris/Ca buffer. An aliquot of each diluted eglin c solution was mixed with an equal volume of a subtilisin variant solution. The mixed solutions were incubated at room temperature for 15-30 minutes and the protease activity of each sample was then measured by AAPF assay. Using the observed decrease of the protease activity upon addition of each eglin c sample and the known concentration of the eglin c, the concentration of the eglin c necessary for the complete inhibition of each subtilisin enzyme variant was calculated. This concentration is equivalent to the enzyme concentration in the sample. An aliquot of the Tris/Ca buffer only, without eglin c, was also mixed with each subtilisin variant sample and the protease activity in the absence of eglin c was measured by AAPF assay. The specific activity of the subtilisin variants was then calculated using the enzyme concentrations as determined above.
BMI Microswatch Assay
[0705] Blood milk and ink (BMI) stained microswatches (EMPA116) of 5.5 millimeter circular diameter were obtained from CFT. In one method, the EMPA116 BMI fabric is pre-rinsed in water prior to cutting them into a 96 well microtiter plate (Corning 3641), one microswatch per well. In the second method the EMPA 116 cloth is cut directly into a 96 well microtiter plate (Corning 3641) where the swatches are then rinsed with two water washes. The rinses are carried out by adding 200 .mu.l of Milli Q water to each well/swatch and mixing them on a table top mixer (Lab line instruments, Titer plate shaker, model 4825) for 15 minutes at a setting of 7. The wash liquor is removed and 200 .mu.l of water is added again to the swatch for another 15 minute rinse. The wash water is removed and the swatches are then air dried in the microtiter plate.
[0706] Detergent compositions 7-11 were diluted in Milli-Q (deionized) water to final working concentrations described in Table 1-1. These detergents were buffered with 2 mM sodium carbonate, pH 10.3. Additionally, a water hardness composition (3:1 Ca:Mg.-CaCl.sub.2:MgCl2.6H.sub.2O) was added to each detergent solution to the final concentration described in Table 1-1. The detergent solutions were mixed at room temperature for 0.5 to 2 hours, centrifuged in 50 mL polypropylene conical tubes at 3000.times.g for 5-10 minutes and were kept at room temperature for the 32.degree. C. assays or pre-equilibrated in an ice-water bath for the 16.degree. C. assays. Then, 190 .mu.l of the desired detergent solution was added to each well of the MTP containing BMI microswatches. To this mixture, 5-15 .mu.l of the diluted enzyme master dilution solution were added, making the approximate concentration of enzyme in the reaction 0.25-2 .mu.g/ml. The enzyme master dilution solution was prepared from the filtered culture supernatants (see TCA assay described above) at .about.2.5-20 .mu.g/mL. The MTP was sealed with tape and placed in the iEMS incubator/shaker (Thermo/Labsystems) pre-set at 16.degree. C. in a refrigerator for 30 minutes or at 32.degree. C. on the benchtop for 30 minutes, with agitation at 1400 rpm. Following incubation under the appropriate conditions, 120-125 .mu.l of the solution from each well was transferred into a fresh MTP (Corning 9017). The new MTP containing 125 .mu.l of solution/well was read at 600 nm (with 5 sec mixing mode in the plate reader) using the MTP SpectraMax reader. Blank controls containing a microswatch and detergent without any enzyme were also included. The absorbance value obtained was corrected for the blank value (substrate without enzyme), providing a measure of hydrolytic activity. For each sample (variant) the performance index was calculated. The performance index compares the performance of the variant (measured value) and the standard enzyme (theoretical value) at the same protein concentration. In addition, the theoretical values can be calculated, using the parameters of a performance dose response curve of the standard protease. A performance index (PI) that is greater than 1 (PI>1) identifies a better variant as compared to the standard (e.g., wild-type), while a PI of 1 (PI=1) identifies a variant that performs the same as the standard, and a PI that is less than 1 (PI<l) identifies a variant that performs worse than the standard. Thus, the PI identifies winners as well as variants that are less desirable for use under certain circumstances.
TABLE-US-00070 TABLE 1-1 Final Detergent, Water Hardness, and Buffer Concentrations Used for BMI Microswatch Assays Final Detergent Final Sodium Detergent Concentration Final Water Carbonate Buffer Composition (g/L) Hardness* (gpg) Concentration (mM) 7 0.808 6 2 8 1 3 2 9 2.3 12 2 10 5.9 12 2 11 8.3 12 2
LAS/EDTA Stability Assay
[0707] The stability of protease variants in the presence of a representative anionic surfactant (LAS=linear alkylbene sulfonate, sodium dodecylbenzenesulfonate-DOBS) and di-sodium EDTA is measured after incubation under defined conditions and the residual activity is determined using the AAPF assay described above. The reagents used were dodecyllbenzene sulfonate, sodium salt (DOBS; Sigma No. D-2525), TWEEN.RTM.-80 (Sigma No. P-8074), di-sodium EDTA (Siegfried Handel No. 164599-02), HEPES (Sigma No. H-7523), unstressed buffer: 50 mM HEPES (11.9 g/1)+0.005% TWEEN.RTM.-80, pH 8.0, Stress buffer: 50 mM HEPES (11.9 g/1), 0.1% (w/v) DOBS (1 g/1), 10 mM EDTA (3.36 g/1), pH 8.0, reference protease and protease variant culture supernatants, containing 200-400 .mu.g/ml protein. The equipment used is V- or U-bottom MTP as dilution plates (Greiner 651101 and 650161 respectively), F-bottom MTP (Corning 9017) for unstress and LAS/EDTA buffer as well as for suc-AAPF-pNA plates, Biomek FX (Beckman Coulter), Spectramax Plus 384 MTP Reader (Molecular Devices), and iEMS Incubator/Shaker (Thermo/Labsystems).
[0708] The iEMS incubator/shaker (Thermo/Labsystems) is set at 29.degree. C. Culture supernatants were diluted into plates containing unstress buffer to a concentration of .about.25 ppm (master dilution plate). For the assay, 20 .mu.l of sample from the master dilution plate is added to plates containing 180 .mu.l unstress buffer to give a final incubation concentration of 2.5 ppm. The contents were mixed and kept at room temperature and the AAPF assay is performed on this plate. In addition, 20 .mu.l of sample from the master dilution plate is also added to plates containing 180 .mu.l stress buffer (50 mM HEPES (11.9 g/1), 0.1% (w/v) DOBS (1 g/1), 10 mM EDTA (3.36 g/1), pH 8.0). The solutions were mixed and immediately placed in 29.degree. C. iEMS shaker for 30 min at 400 rpm. Following 30 minutes of incubation, the AAPF assay is performed on the stress plate. The stability of the samples is determined by calculating the ratio of the residual and initial AAPF activity as follows: Residual Activity (%)=[mODmin-1 stressed]*100/[mODmin-1 unstressed].
[0709] The final detergent, water hardness and buffer concentrations are determined based on the assay system to be used (e.g., North American, Japanese, Western European, or Central European conditions). In some aspects, the stain removal performance of the protease variants is determined in commercially available detergents. Heat inactivation of commercial detergent formulas serves to destroy the enzymatic activity of any protein components while retaining the properties of non-enzymatic components. Thus, this method is suitable for preparing commercially purchased detergents for use in testing the enzyme variants of the present invention.
Baked Egg Microtiter Assay
[0710] For this assay, 96-well baked egg yolk substrate plates are prepared from chicken egg yolks. Chicken egg yolks are separated from the whites, released from the membrane sac, and diluted 20% (vol/weight) with Milli-Q water. The diluted yolk is stirred for 15 min at room temperature using a magnetic stirrer. Five .mu.L are carefully pipetted into the center of each well of a 96-well V-bottom plate (Costar #3894) using an 8-channel pipette. The plates are baked at 90.degree. C. for 1 hour and cooled at room temperature. The baked egg yolk substrate plates are stored at room temperature and used within one week of preparation. Automatic dish detergents are prepared as described herein and pre-heated to 50.degree. C. A190 .mu.L aliquot of detergent is added to each well of the 96-well plate using an 8-channel pipette. Ten .mu.L of diluted enzyme is added to each well using a 96-channel pipetting device. The plate is carefully sealed with an adhesive foil sealer and incubated at 50.degree. C. with shaking for 30 min. 120 .mu.L of the reaction mixture is transferred to a new 96-well flat-bottom plate, and the absorbance/light scattering is determined at 405 nm. The absorbance/light scattering at 405 nm is proportional to egg yolk removal.
Egg Yolk Microswatch Assay ("CS-38 Microswatch Assay"; or "EGG" or "Dish")
[0711] Automatic dish detergents are prepared as described herein. The equipment used included a New Brunswick Innova 4230 shaker/incubator and a SpectraMAX (type 340) MTP reader. The MTPs are obtained from Costar (type 9017). Aged egg yolk with pigment swatches (CS-38) are obtained from Center for Test Materials (Vlaardingen, Netherlands). Before cutting 0.25-inch circular microswatches, the fabric is washed with water. One microswatch is placed in each well of a 96-well microtiter plate. The test detergent is equilibrated at 50.degree. C. 190 .mu.l of detergent solution is added to each well of the MTP, containing microswatches. To this mixture, 10 .mu.l of the diluted enzyme solution is added. The MTP is sealed with adhesive foil and placed in the incubator for 30 minutes, with agitation. Following incubation, 100 .mu.l of the solution from each well is transferred into a fresh MTP. This MTP is read at 405 nm using a SpectraMax MTP reader. Blank controls, as well as controls containing microswatches and detergent but no enzyme are also included.
[0712] In some aspects, pre-washed microswatches find use. This type of microswatch is pre-washed in deionised water for 20 minutes at ambient temperature. After the pre-washing step, the swatches are put on top of paper towels to dry. The air-dried swatches are then punched using a 1/4'' circular die on an expulsion press. Finally two microswatches are put into each well of a 96-well MTP vertically to expose the whole surface area (i.e. not flat on the bottom of the well).
[0713] Samples of protease variants to be tested are obtained from filtered culture broth of cultures grown in MTP plates. The equipment used is a Biomek FX Robot (Beckman Coulter), a SpectraMAX MTP Reader (type 340; Molecular Devices), an iEMS incubator/shaker (Thermo/Labsystems); F-bottom MTPs (Costar type 9017 used for reading reaction plates after incubation); and V-bottom MTPs (Greiner 651101 used for pre-dilution of supernatant). In this assay, the proteases hydrolyze the substrate and liberate pigment and insoluble particles from the substrate. Thus the rate of turbidity is a measure of enzyme activity.
[0714] The stain removal performance of reference serine proteases and variants therefrom on microswatches is determined on a MTP scale in commercially available detergent (Calgonit 5 in1). CS-38 microswatches (egg-yolk with pigment, aged by heating), obtained from CFT Vlaardingen are used as substrate. Two swatches are used per well. ADW tablets from Calgonit 5in1 are used to prepare the detergent solution. To inactivate the protease activity present in the tablets, a 21 g tablet is dissolved in Milli-Q water heated in a water bath to a temperature of 60.degree. C. The solution is cooled to room temperature and the volume of water adjusted to 700 mL. The solution is further diluted with water to achieve a final concentration of 3 g/1. Water hardness is adjusted to 21.degree. GH by adding 1.46 ml of the Ca/Mg-mixture (Ca/Mg mixture [(3:1), 1.92 M CaCl.sub.2)=282.3 g/L CaCl.sub.2.2H.sub.20; 0.64 M MgCl.sub.2=130.1 g/L MgCl.sub.2.6H.sub.2O), 15000 gpg]. The enzyme samples are prediluted in 10 mM NaCl, 0.1 mM CaCl.sub.2), 0.005% TWEEN.RTM.-80 solution and tested at appropriate concentrations.
[0715] The incubator is set at the desired temperature of 50.degree. C. 72 .mu.l of dilution buffer is added to the empty V-bottom plate (i.e., a "dilution plate") followed by 8 .mu.l supernatant. 9 .mu.l from the dilution plate is added to plates containing the microswatches incubated in 171 .mu.l detergent solution. 9 .mu.l from the dilution plate is added to plates containing the microswatches to give a total dilution of supernatant of 200.times.. The microswatch plate (with detergent and enzyme) is covered with tape and placed in the incubator/shaker for 30 minutes at 1400 rpm. Following incubation, 75 .mu.l of the reaction mixture is transferred to an empty F-bottom plate and the absorbance is read in a MTP Reader at 405 nm after de-bubbling with a hair dryer. Blank controls, containing one or two microswatches and detergent without the addition of reference protease containing samples are also included in the test.
[0716] The absorbance value obtained is corrected for the blank value (substrate without enzyme), providing a measure of hydrolytic activity. For each sample (variant) the performance index is calculated. The performance index compares the performance of the variant (actual value) and the standard enzyme (theoretical value) at the same protein concentration. In addition, the theoretical values can be calculated, using the parameters of the Langmuir equation of the standard enzyme.
Egg Yolk Stains on Stainless Steel
[0717] The stainless steel sheets (10.times.15 cm; brushed on one side) used in these experiments are thoroughly washed at 95.degree. C. in a laboratory dishwasher with a high-alkalinity commercial detergent (e.g., ECOLAB.RTM. detergent; Henkel) to provide sheets that are clean and grease-free. These sheets are deburred prior to their first use. The sheets are dried for 30 minutes at 80.degree. C. in a thermal cabinet before being soiled with egg yolk. The surfaces to be brushed are not touched prior to soiling. Also, no water stains or fluff on the surfaces are permitted. The cooled sheets are weighed before soiling.
The egg yolks are prepared by separating the yolks of approximately 10-11 eggs (200 g of egg yolk) from the whites. The yolks are stirred with a fork in a glass beaker to homogenize the yolk suspension. The yolks are then strained (approx. 0.5 mm mesh) to remove coarse particles and any egg shell fragments. A flat brush (2.5'') is used to apply 1.0.+-.0.1 g egg yolk suspension as uniformly as possible over an area of 140 cm.sup.2 on the brushed sides of each of the stainless steel sheets, leaving an approx. 1 cm wide unsoiled rim (adhesive tape is used if needed). The soiled sheets are dried horizontally (to prevent formation of droplets on the edges of the sheets), at room temperature for 4 hours (max. 24 h).
[0718] For denaturation, the sheets are immersed for 30 seconds in boiling, demineralized water (using a holding device if necessary). Then, the sheets are dried again for 30 min at 80.degree. C. After drying and cooling, the sheets are weighed. After weighing, the sheets are left for at least 24 hours (20.degree. C., 40-60% relatively humidity) before submitting them to the wash test. In order to meet the testing requirements, only sheets with 500.+-.100 mg/140 cm.sup.2 (egg yolk after denaturation), are used in the testing After the wash tests are conducted, the sheets are dried for 30 min at 80.degree. C., in the thermal cabinet, and weighed again after cooling. The percent cleaning performance is determined by dividing the (mg of egg yolk released by washing.times.100) by the (mg of denatured egg yolk applied).
Minced Meat on Porcelain Plates
[0719] For these experiments, dessert plates (Arzberg, white, glazed porcelain) conforming to EN 50242, form 1495, No. 0219, diameter 19 cm are used. A total of 225 g lean pork and beef (half and half) is finely chopped and cooled, after removing visible fat. The mixture is twice run through a mincer. Temperatures above 35.degree. C. are avoided. Then, 225 g of the minced meat is mixed with 75 g of egg (white and yolk mixed together). The preparation is then frozen up to three months at -18.degree. C., prior to use. If pork is not available, beef is used.
[0720] The minced meat and egg mixture (300 g) is brought up to room temperature and mixed with 80 ml synthetic water. The mixture is then homogenized using a kitchen hand blender for 2 min. Then, a fork is used to spread 3 g of the minced meat/egg/water mixture on each white porcelain plate, leaving an approx. 2 cm wide unsoiled margin around the rim. The amount applied is 11.8.+-.0.5 mg/cm.sup.2. The plates are dried for 2 hours at 120.degree. C. in a preheated thermal cabinet. As soon as the plates are cooled, they are ready for use. The plates are stacked with paper towels between each of the plates.
[0721] After washing, the plates are sprayed with ninhydrin solution (1% ethanol) for better identification of the minced meat residues. To promote the color reaction, the plates are heated for 10 min at 80.degree. C. in the thermal cabinet. Evaluation of the washing performance is done by visually inspecting the color reactions of the minced meat residues with reference to the IKW photographic catalogue (IKW).
Egg/Milk Stains on Stainless Steel
[0722] The stainless steel sheets (10.times.15 cm; brushed on one side) used in these experiments are thoroughly washed at 95.degree. C. in a laboratory dishwasher with a high-alkalinity commercial detergent to remove grease and clean the sheets. The sheets are polished dry with a cellulose cloth. The surfaces to be brushed are not touched prior to soiling. Also, no water stains or fluff on the surfaces are permitted. Before soiling, the sheets are placed in a thermal cabinet at 80.degree. C., for 30 min. The cooled sheets are weighed before soiling.
[0723] The egg yolks and whites of whole raw eggs (3-4 eggs; 160 g/egg) are placed in a bowl and beaten with an egg whisk. Then, 50 ml semi-skimmed UHT (1.5% fat, ultra-high temperature, homogenized) milk are added to the mixture. The milk and egg are mixed without generating froth. A flat brush is used to uniformly distribute 1.0.+-.0.1 g of the egg/milk mixture on the brushed side of the stainless steel sheets, using a balance to check the distribution. A margin of approximately 1.0 cm is left around the short sides of the sheets. The soiled sheets are dried horizontally (to prevent formation of droplets on the edges of the sheets), at room temperature for 4 hours (max. 24 h).
[0724] The sheets are then immersed for 30 seconds in boiling, demineralized water (using a holding device if necessary). Then, the sheets are dried again for 30 min at 80.degree. C. After drying and cooling, the sheets are weighed. After weighing, the sheets are left for at least 24 hours (20.degree. C., 40-60% relatively humidity), before submitting them to the wash test. In order to meet the testing requirements, only sheets with 190.+-.10 mg egg yolk are used.
[0725] After the wash tests are conducted, the sheets are dried for 30 min at 80.degree. C., in the thermal cabinet, and weighed again after cooling. The percentage cleaning performance is determined by dividing the (mg of egg/milk released by washing.times.100) by the (mg of egg/milk applied).
Preparation of the Spaghetti Mix Stain on Porcelain Plates
[0726] Pasta sauce (390 g) is mixed with 150 g of boiled spaghetti pasta, 25 g of minced meat (improved IKW composition-a combination of 225 gram fat free minced meat and 75 gram egg yolk) and 50 g of Grozette Formaggio cheese. A spoon is used to spread 3 g of this mixture on each white porcelain plate (Arzberg, 19 cm diameter, white, glazed porcelain, conforming to EN 50242, form 1495, No. 0219) leaving an approximately 2 cm wide unsoiled margin around the rim. The plates are dried by baking them for 2 hours at 120.degree. C. in an oven. As soon as the plates are cooled, they are ready for use. The plates are stacked with paper towels between each of the plates for storage. After washing, the plates are sprayed with iodine solution (0.05N) for better identification of the carbohydrate residues. Evaluation of the washing performance is done by visually inspecting the color reactions of the carbohydrate residues with reference to the IKW photographic catalogue (IKW) and rated on a scale of 0-10 (10 being clean).
Performance Index
[0727] The performance index compares the performance of the variant (measured value) and the standard enzyme (theoretical value) at the same protein concentration. In addition, the theoretical values can be calculated, using the parameters of a performance dose response curve of the standard protease.
[0728] Various terms set forth below are used to describe the variant: non-deleterious variants have a PI>0.05; deleterious variants have a PI=0.05; combinable variants are those for which the variant has performance index values greater than or equal to 0.2 for at least one property, and >0.05 for all properties. Combinable variants are those that can be combined to deliver proteins with appropriate performance indices for one or more desired properties. These data find use in engineering any subtilisin/subtilase. Even if the subtilase to be engineered has an amino acid different from that of subtilisin GG36 at particular positions, these data find use in finding substitutions that will alter the desired properties by identifying the best choices for substitutions, including substitutions to the GG36 wild type amino acid.
Example 2
Generation of GG36 Single Mutants Using Site Evaluation Libraries (SELs)
[0729] The construction of GG36 SELs described in this example was performed by GENEART using their proprietary methods and technology platform for gene optimization, gene synthesis, library generation and analysis (WO 2004/059556A3, European Patent Nos. 0 200 362 and 0 201 184; and U.S. Pat. Nos. 4,683,195, 4,683,202 and 6,472,184). The GG36 SELs were produced at positions pre-selected by the inventors using the pHPLT-GG36 B. subtilis expression plasmid (see FIG. 6). This B. subtilis expression plasmid contains the GG36 expression cassette shown below, the B. licheniformis LAT promoter (Plat), and additional elements from pUB110 (McKenzie et al., Plasmid, 15:93-103, 1986) including a replicase gene (reppUB), a neomycin/kanamycin resistance gene (neo) and a bleomycin resistance marker (bleo) (FIG. 4 in U.S. Pat. No. 6,566,112). The pHPLT-GG36 plasmid map is provided at FIG. 6. The GG36 expression cassette sequence is provided below.
[0730] The DNA sequence of GG36 (the signal sequence is shown in lower case letters, propeptide in lower case, underlined text, and GG36 mature sequence in uppercase letters) is provided below:
TABLE-US-00071 (SEQ ID NO: 756) Gtgagaagcaaaaaattgtggatcgtcgcgtcgaccgcactactcatttc tgttgctttcagttcatcgatcgcatcggctgctgaagaagcaaaagaaa aatatttaattggctttaatgagcaggaagctgtcagtgagtttgtagaa caagtagaggcaaatgacgaggtcgccattctctctgaggaagaggaagt cgaaattgaattgcttcatgaatttgaaacgattcctgttttatccgttg agttaagcccagaagatgtggacgcgcttgagctcgatccagcgatttct tatattgaagaggatgcagaagtaacgacaatgGCGCAATCAGTGCCATG GGGAATTAGCCGTGTGCAAGCCCCAGCTGCCCATAACCGTGGATTGACAG GTTCTGGTGTAAAAGTTGCTGTCCTCGATACAGGTATTTCCACTCATCCA GACTTAAATATTCGTGGTGGCGCTAGCTTTGTACCAGGGGAACCATCCAC TCAAGATGGGAATGGGCATGGCACGCATGTGGCCGGGACGATTGCTGCTT TAAACAATTCGATTGGCGTTCTTGGCGTAGCGCCGAGCGCGGAACTATAC GCTGTTAAAGTATTAGGGGCGAGCGGTTCAGGTTCGGTCAGCTCGATTGC CCAAGGATTGGAATGGGCAGGGAACAATGGCATGCACGTTGCTAATTTGA GTTTAGGAAGCCCTTCGCCAAGTGCCACACTTGAGCAAGCTGTTAATAGC GCGACTTCTAGAGGCGTTCTTGTTGTAGCGGCATCTGGAAATTCAGGTGC AGGCTCAATCAGCTATCCGGCCCGTTATGCGAACGCAATGGCAGTCGGAG CTACTGACCAAAACAACAACCGCGCCAGCTTTTCACAGTATGGCGCAGGG CTTGACATTGTCGCACCAGGTGTAAACGTGCAGAGCACATACCCAGGTTC AACGTATGCCAGCTTAAACGGTACATCGATGGCTACTCCTCATGTTGCAG GTGCAGCAGCCCTTGTTAAACAAAAGAACCCATCTTGGTCCAATGTACAA ATCCGCAATCATCTAAAGAATACGGCAACGAGCTTAGGAAGCACGAACTT GTATGGAAGCGGACTTGTCAATGCAGAAGCTGCAACTCGTTAA
[0731] The protein sequence of GG36 (the signal sequence is shown in lower case letters, propeptide in lower case, underlined text, and GG36 mature protease sequence in uppercase letters) is provided below:
TABLE-US-00072 (SEQ ID NO: 757) vrskklwivastallisvafsssiasaaeeakekyligfneqeavsefve qveandevailseeeeveiellhefetipvlsvelspedvdaleldpais yieedaevttmAQSVPWGISRVQAPAAHNRGLTGSGVKVAVLDTGISTHP DLNIRGGASFVPGEPSTQDGNGHGTHVAGTIAALNNSIGVLGVAPSAELY AVKVLGASGSGSVSSIAQGLEWAGNNGMHVANLSLGSPSPSATLEQAVNS ATSRGVLVVAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAG LDIVAPGVNVQSTYPGSTYASLNGTSMATPHVAGAAALVKQKNPSWSNVQ IRNHLKNTATSLGSTNLYGSGLVNAEAATR.
[0732] The method of mutagenesis was based on the codon-specific mutation approach in which all possible amino acid substitutions are simultaneously created at a specific codon of interest using forward and reverse mutagenesis primers that contain a degenerate codon, NNS ((A,C,T or G), (A,C,T or G), (C or G)) at the site of interest. To construct each of the GG36 SELs, three PCR reactions were performed: two mutagenesis reactions (primary PCR1 and PCR2) to introduce the mutated codon of interest in the mature GG36 DNA sequence using the NNS forward and reverse mutagenesis primers (25-45 nucleotides long), and a third reaction to fuse the two mutagenesis PCR products together to construct the pHPLT-GG36 expression vector having the desired mutated codons in the mature GG36 sequence.
[0733] The primer sequences used in this Example are provided below:
TABLE-US-00073 TABLE 2-1 Primers Sequence Primer Name CGCGCTTGAGCTCGATCCAGCGATTTC (SEQ ID NO: 758) SacI-Fw GTCTCCAAGCTTTAACGAGTTGCAG (SEQ ID NO: 759) HindIII-Rv GCAATTCAGATCTTCCTTCAGGTTATGACC (SEQ ID NO: 760) pHPLT-BglII-Fw GCATCGAAGATCTGATTGCTTAACTGCTTC (SEQ ID NO: 761) pHPLT-BglII-Rv
[0734] The Phusion High-Fidelity DNA Polymerase (Finnzymes catalog no. F-530L) was used for all PCRs, and the reactions were executed according to manufacturer's protocols that were supplied with the polymerase. In particular, for primary PCR 1, 1 .mu.L (10 .mu.M) of each of the pHPLT-BglII-Fw primer and a NNS reverse mutagenesis primer were used, and for primary PCR 2, 1 .mu.L (10 .mu.M) of the pHPLT-BglII-Rv primer and a NNS forward mutagenesis primer were used. Each reaction also included 1 .mu.L of the pHPLT-GG36 plasmid template DNA (0.1-1 ng/.mu.L). An MJ Research PTC-200 Peltier thermal cycler was used for the PCRs. The reactions yielded two fragments of approximately 2 to 3 kb having approximately 30 nucleotide overlap surrounding the GG36 codon of interest. The fragments obtained were fused in a third PCR similar to the ones described above using 1 .mu.L of primary PCR 1 reaction mix, 1 .mu.L, of primary PCR 2 reaction mix and 1 .mu.L (10 .mu.M) of each of the forward and reverse SacI-Fw and HindIII-Rv primers. The amplified linear 859 bp fragment encoding the GG36 variant gene was purified (using QIAGEN.RTM. Qiaquick PCR purification kit) and digested with the SacI and HindIIII restriction enzymes to create cohesive ends on both sides of the fusion fragment. About 50 ng of plasmid pHPLT-GG36 was also purified after digestion with SacI and HindIIII, resulting in a 3.9 kb vector backbone fragment. The digested vector fragment was ligated with 50 ng of the digested 859 bp fragment encoding the variant enzyme using the T4 DNA ligase (Invitrogen) following the manufacturer's protocol for cloning of cohesive ends. Subsequently, the ligation mixture was used to transform B. subtilis cells (.DELTA.aprE, .DELTA.nprE, oppA, .DELTA.spoIIE, degUHy32, .DELTA.amyE::[xylR,pxylA-comK]) as described (WO 2002/014490).
[0735] To express the variant proteins for further biochemical analyses, the B. subtilis strains carrying the GG36 variant plasmids were inoculated into microtiter plates containing 150 .mu.l Luria broth medium supplemented with 10 .mu.g/ml neomycin. Plates were grown overnight at 37.degree. C. with 300 rpm shaking and 80% humidity using Enzyscreen lids for microtiter plates (Enzyscreen). Ten microliters from the overnight culture plate were used to inoculate a new microtiter plate containing 190 .mu.l of MBD medium (a MOPS based defined medium) with 10 ug/ml neomycin. MBD medium was prepared essentially as known in the art (see Neidhardt et al., J. Bacteriol. 119:736-747 [1974]), except that NH.sub.4Cl.sub.2, FeSO.sub.4, and CaCl.sub.2 were omitted from the base medium, 3 mM K.sub.2HPO.sub.4 was used, and the base medium was supplemented with 60 mM urea, and 100 ml of a solution made of 210 g/L glucose, and 350 g/L maltodextrin. The micronutrients were made up as a 100.times. stock solution containing in one liter, 400 mg FeSO.sub.4.7H.sub.2O, 100 mg MnSO.sub.4.H.sub.2O, 100 mg ZnSO.sub.4.7H.sub.2O, 50 mg CuCl.sub.2.2H.sub.2O, 100 mg CoCl.sub.2.6H.sub.2O, 100 mg NaMoO.sub.4.2H.sub.2O, 100 mg Na.sub.2B.sub.4O.sub.7.10H.sub.2O, 10 ml of 1M CaCl.sub.2, and 10 ml of 0.5 M sodium citrate. The MBD medium containing microtiter plates were grown for 68 hours at 37.degree. C., 300 rpm, and 80% humidity using Enzyscreen lids (Enzyscreen) for determining protein expression. The next day, cultures were filtered through a micro-filter plate (0.22 .mu.m; Millipore) and the resulting filtrate was used for biochemical analysis. The TCA and BMI microswatch assays for the detergent compositions 7-11 were carried out as described in Example 1. Performance indices were also calculated as described under the BMI assay description in Example 1, and they are shown in Table 2-2 relative to GG36. In the following Tables, the detergent compositions ("Det.") correspond to those shown in Table D, above. Also, as indicated, the amino acid position is listed according to BPN' numbering.
TABLE-US-00074 TABLE 2-2 Single Variants of GG36 with Performance Indices of at Least 0.2 Relative to GG36 in Either TCA or BMI Microswatch Cleaning at 16.degree. C. in Detergents 7-11. GG36 Amino Acid Position (BPN' WT Mutant Numbering) Residue Residue 1 A R 2 Q A 2 Q R 2 Q S 2 Q M 2 Q W 3 S R 4 V R 4 V S 4 V C 8 I A 9 S W 9 S F 9 S A 10 R A 10 R M 10 R S 10 R H 12 Q F 12 Q R 14 P F 14 P K 14 P Q 15 A R 15 A F 16 A S 17 H R 17 H F 17 H M 18 N R 18 N K 20 G R 20 G K 20 G F 22 T R 22 T Q 22 T L 22 T V 22 T W 22 T Y 22 T A 23 G A 23 G S 23 G F 24 S R 24 S W 24 S H 24 S L 24 S Q 24 S F 25 G R 25 G F 25 G V 26 V F 27 K R 27 K L 27 K V 27 K F 28 V A 28 V E 28 V N 29 A T 30 V E 31 L F 33 T S 33 T G 33 T D 34 G P 35 I M 36 S T 36 S F 36 S R 38 T R 38 T F 38 T L 40 P H 40 P W 40 P R 40 P N 40 P T 40 P L 42 L I 43 N R 43 N A 43 N S 43 N W 43 N F 43 N I 43 N D 43 N M 45 R T 46 G R 48 A R 50 F C 51 V W 51 V F 51 V H 52 P F 52 P N 52 P E 55 P Y 57 T R 59 Q A 59 Q F 59 Q R 60 D P 60 D A 60 D Q 62 N Q 62 N E 63 G S 63 G A 63 G M 63 G V 63 G T 63 G H 63 G Q 63 G I 63 G D 63 G E 63 G P 64 H F 64 H T 68 V A 68 V C 69 A N 69 A T 69 A W 69 A P 71 T G 72 I C 74 A C 75 L R 75 L A 75 L E 75 L F 78 S R 78 S I 78 S N 79 I Q 79 I W 81 V R 82 L R 82 L T 82 L M 82 L F 82 L V 85 A M 86 P W 86 P I 86 P L 89 E P 89 E W 89 E T 89 E I 89 E H 89 E V 89 E F 89 E L 89 E W 89 E G 91 Y F 91 Y N 92 A F 94 K N 99 S F 99 S T 99 S M 99 S G 99 S P 100 G I 100 G S 100 G N 100 G Q 101 S N 101 S G 101 S T 101 S A 101 S D 101 S F 101 S D 101 S E 101 S P 102 G A 102 G N 102 G T 102 G E 102 G H 103 S N 103 S G 103 S D 104 V L 104 V I 104 V E 104 V D 105 S T 105 S Q 105 S E 106 S V 106 S G 106 S T 106 S A 106 S E 106 S D 106 S F 107 I F 107 I M 108 A I 108 A G 109 Q M 111 L V 111 L I 112 E V 112 E L 112 E Q 114 A G 115 G R 115 G K 116 N L 116 N A 116 N K 117 N F 118 G I 118 G R 119 M C 120 H A 120 H F 120 H R 121 V E 121 V F 123 N G 123 N E 124 L S 128 S N 128 S M 128 S H 128 S Q 128 S I
128 S F 128 S L 128 S D 129 P E 132 S A 132 S E 138 A G 144 S R 147 V L 148 L I 158 A E 159 G C 159 G E 160 S D 166 S E 166 S D 167 Y W 175 M V 177 V C 181 D A 182 Q R 183 N D 183 N R 183 N I 183 N F 183 N M 185 N I 185 N E 185 N V 186 R H 186 R K 188 S R 188 S E 188 S D 192 Y W 192 Y H 194 A V 194 A F 194 A E 197 D F 198 I L 198 I F 203 V E 203 V C 208 T S 209 Y N 209 Y S 209 Y F 209 Y T 209 Y H 209 Y L 209 Y G 209 Y E 210 P V 210 P R 210 P L 211 G R 211 G Q 212 S I 212 S F 212 S M 213 T A 214 Y F 215 A F 215 A N 215 A H 215 A E 215 A D 216 S F 216 S A 217 L E 217 L N 217 L D 218 N P 218 N E 218 N D 224 T A 224 T G 227 V I 230 A E 231 A I 231 A C 233 L C 234 V F 235 K F 236 Q N 236 Q F 238 N R 238 N K 238 N L 239 P R 239 P S 239 P R 239 P H 239 P N 239 P K 239 P T 239 P F 239 P G 240 S R 241 W R 242 S R 242 S L 243 N R 243 N F 244 V R 246 I S 248 N I 248 N V 248 N R 249 H R 249 H T 250 L I 251 K R 251 K S 252 N R 252 N F 252 N H 252 N I 253 T R 253 T F 253 T I 254 A C 256 S N 258 G R 260 T V 260 T I 262 L H 262 L D 263 Y F 265 S F 267 L N 267 L M 267 L V 269 N R 269 N I 270 A C 271 E T 271 E V 271 E L 271 E H 271 E F 271 E P 271 E A 271 E M 271 E I 272 A F 272 A R 273 A I 273 A F 274 T G
TABLE-US-00075 TABLE 2-3 GG36 Single Variants with Performance Indices of at Least 1.5 Relative to GG36 BMI Microswatch Cleaning at 32.degree. C. in Detergent 7. GG36 Variant N62E A158E G159E
TABLE-US-00076 TABLE 2-4 GG36 Single Variants with Performance Indices of at least 1.2 Relative to GG36 BMI Microswatch Cleaning at 32.degree. C. in Detergent 10. GG36 Variant A1R S78R V244R N269R E271L
Example 3
Construction and Cleaning Performance of the NHJ1 and WCE1 Sets of GG36 Variants
[0736] The NHJ1 and WCE1 set of GG36 variants described herein were constructed at DNA 2.0, Inc., using the pHPLT-GG36 B. subtilis expression plasmid described above (FIG. 6). The variants were expressed in B. subtilis cells (genotype: AaprE, AnprE, amyE::xylRPxylAcomK-phleo) as described in Example 2, and were further characterized using the TCA assay for protein content determination, LAS/EDTA stability assay, and BMI microswatch cleaning assay as described in Example 1. These results are shown in Tables 3-1 and 3-2. In the following Tables, the detergent compositions ("Det.") correspond to those shown in Table D, above. Also, as indicated, the amino acid position is listed according to BPN' numbering.
TABLE-US-00077 TABLE 3-1 NHJ1 Variants with Performance Indices of at least 0.25 Relative to GG36 in Any One of TCA, LAS/EDTA Stability, or BMI Microswatch Cleaning at 16.degree. C. in Detergents 7, 8 or 9. GG36 Variant (BPN' Numbering) N062E-A158E S103G-A158E S128N-A158E A016S-A158E V104L-A158E E089P-A158E L111V-A158E T022A-A158E S101A-A158E L148I-A158E P129E-A158E T022A-E089P A016S-E089P N062E-E089P N062E-E271F A158E-E271F R186H-E271F P129E-E271F L111V-E271F Y209E-E271F A016S-E271F S188D-E271F T022A-E271F G159E-E271F V104L-E271F S101A-E271F E089P-E271F S128N-E271F S103G-E271F L148I-E271F H249R-E271F N062E-G159E A016S-G159E S128N-G159E L148I-G159E L111V-G159E E089P-G159E T022A-G159E P129E-G159E S103G-G159E V104L-G159E A158E-G159E S101A-G159E A158E-H249R L111V-H249R P129E-H249R N062E-H249R A016S-H249R R186H-H249R L148I-H249R G159E-H249R S101A-H249R S188D-H249R V104L-H249R Y209E-H249R T022A-H249R S128N-H249R S103G-H249R E089P-H249R T022A-L111V S101A-L111V A016S-L111V V104L-L111V N062E-L111V S103G-L111V E089P-L111V A016S-L148I N062E-L148I T022A-L148I P129E-L148I V104L-L148I S103G-L148I S128N-L148I S101A-L148I E089P-L148I L111V-L148I A016S-N062E T022A-N062E N062E-P129E T022A-P129E S128N-P129E A016S-P129E S101A-P129E V104L-P129E E089P-P129E S103G-P129E L111V-P129E N062E-R186H S128N-R186H S101A-R186H T022A-R186H A016S-R186H A158E-R186H E089P-R186H P129E-R186H G159E-R186H S103G-R186H V104L-R186H L111V-R186H L148I-R186H N062E-S101A T022A-S101A A016S-S101A E089P-S101A N062E-S103G T022A-S103G A016S-S103G S101A-S103G E089P-S103G N062E-S128N A016S-S128N T022A-S128N S101A-S128N V104L-S128N E089P-S128N S103G-S128N L111V-S128N L111V-S188D N062E-S188D A016S-S188D L148I-S188D T022A-S188D S128N-S188D S101A-S188D V104L-S188D E089P-S188D P129E-S188D G159E-S188D R186H-S188D S103G-S188D A158E-S188D A016S-T022A A016S-V104L T022A-V104L S101A-V104L N062E-V104L S103G-V104L E089P-V104L G159E-Y209E L111V-Y209E S101A-Y209E A016S-Y209E S128N-Y209E L148I-Y209E P129E-Y209E N062E-Y209E T022A-Y209E S103G-Y209E A158E-Y209E S188D-Y209E V104L-Y209E E089P-Y209E R186H-Y209E GG36
TABLE-US-00078 TABLE 3-2 WCE1 Variants with Performance Indices of at least 0.2 Relative to GG36 in Any One of TCA, LAS/EDTA Stability, or BMI Microswatch Cleaning at 16.degree. C. in Detergents 10 or 11. GG36 Variant (BPN' Numbering) N018R-W241R G020R-W241R S024R-W241R S009A-W241R G020R-W241R V004R-W241R N043R-W241R S078R-W241R T022R-W241R G115R-W241R A001R-W241R S212F-W241R L082R-W241R N018R-V244R S024R-V244R S078R-V244R G020R-V244R S212F-V244R S009A-V244R L082R-V244R A001R-V244R N043R-V244R T022R-V244R V004R-V244R G115R-V244R W241R-V244R S242R-V244R A001R-V004R S009A-T022R N018R-T022R G020R-T022R V004R-T022R A001R-T022R S024R-S242R N018R-S242R V004R-S242R G020R-S242R S212F-S242R L082R-S242R S078R-S242R A001R-S242R S009A-S242R T022R-S242R G115R-S242R N043R-S242R W241R-S242R N018R-S212F T022R-S212F V004R-S212F S024R-S212F A001R-S212F G115R-S212F G020R-S212F S009A-S212F N043R-S212F S078R-S212F L082R-S212F S009A-S078R G020R-S078R S024R-S078R T022R-S078R N018R-S078R V004R-S078R A001R-S078R N043R-S078R T022R-S024R G020R-S024R N018R-S024R A001R-S024R V004R-S024R S009A-S024R V004R-S009A A001R-S009A S242R-N269R S024R-N269R G020R-N269R T022R-N269R H249R-N269R S212F-N269R N043R-N269R V244R-N269R A001R-N269R N018R-N269R S078R-N269R S009A-N269R G115R-N269R W241R-N269R V004R-N269R L082R-N269R N018R-N043R G020R-N043R V004R-N043R T022R-N043R S009A-N043R A001R-N043R S024R-N043R S009A-N018R V004R-N018R A001R-N018R S024R-L082R S009A-L082R N018R-L082R A001R-L082R S078R-L082R G020R-L082R T022R-L082R V004R-L082R N043R-L082R N043R-H249R G020R-H249R V004R-H249R N018R-H249R S009A-H249R S212F-H249R T022R-H249R S024R-H249R G115R-H249R A001R-H249R L082R-H249R S242R-H249R W241R-H249R V244R-H249R S078R-H249R N018R-G115R G020R-G115R T022R-G115R S078R-G115R S009A-G115R V004R-G115R A001R-G115R L082R-G115R N043R-G115R S024R-G115R S009A-G020R N018R-G020R V004R-G020R A001R-G020R S009A-E271L G020R-E271L S024R-E271L V244R-E271L W241R-E271L N043R-E271L T022R-E271L H249R-E271L S212F-E271L G115R-E271L S242R-E271L S078R-E271L V004R-E271L N269R-E271L A001R-E271L N018R-E271L L082R-E271L GG36
Example 4
Construction and Cleaning Performance of NHJ4 Set of GG36 Variants
[0737] The NHJ4 set of GG36 variants described in Table 4-4 below were constructed using the pHPLT-GG36 B. subtilis expression plasmid (FIG. 6) using PCR fusion or the QUIKCHANGE.RTM. Multi Site-directed mutagenesis kit ("QCMS kit"; Stratagene) as described below.
[0738] a) Construction of NHJ4 Variants by QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis
[0739] Variants created using the QUIKCHANGE.RTM. Multi Site-Directed Mutagenesis are shown in Table 4-1. The parent plasmid pHPLT-GG36 (template DNA) was methylated using two micrograms of DNA and Dam methylase (NEB), according to the manufacturer's instructions. Site-directed mutants were made by a QuikChange.RTM. Multi Site-Directed Mutagenesis Kit ("QCMS kit"; Stratagene) following the manufacturer's protocol. The following mutations were introduced in the parent plasmid S101A-S103G-V104L, G159E, T22A, Y209E, E271F, S101A, S103G, L111V, S128N, N62E, and S188D, For efficient transformation of B. subtilis, DNA from the QCMS reaction mixtures was amplified by rolling circle amplification (RCA) using the Illustra Templiphi kit (GE Healthcare) and the reaction was performed according to the manufacturer's protocol. One microliter of ten-fold diluted amplified DNA was used to transform 50 .mu.L of competent B. subtilis cells (genotype: .DELTA.aprE, .DELTA.nprE, amyE::xylRPxylAcomK-phleo). The transformation mixture was shaken at 37.degree. C. for 1 hour. Ten microliter aliquots of the transformation mixture were plated on skim milk (1.6%) Luria agar plates supplemented with 10 .mu.g/ml of neomycin (Teknova). Subsequently, the colonies with halos were inoculated in 120 .mu.m of LB media containing 10 .mu.g/mL neomycin for plasmid DNA extraction (QIAprep Spin Miniprep kit, Qiagen). The extracted plasmids were sequenced to confirm the presence of the desired mutations.
[0740] b) Construction of NHJ4 Variants by Extension PCR
[0741] Ten combinatorial mutants of GG36 were created by extension PCR. The list of mutations introduced in the GG36 gene contained in the pHPLT plasmid were T22A, N62E, S103G, S103G-L111V, S101G-S103A-V104I, S101A-S103G-V104L, S101A, S128N, G159D, G159E, Y209E, and L111V. To create each mutant, PCR fragments containing the desired mutations were amplified using mutagenic primers as well as forward and reverse primers to amplify the entire GG36 variant. Each PCR amplification reaction contained 30 pmol of each mutagenic primer and 100 ng of the DNA template, pHPLT-GG36 plasmid. Amplifications were carried out using Vent DNA polymerase (NEB). The PCR reaction (20 .mu.L) was initially heated at 95.degree. C. for 2.5 min followed by 30 cycles of denaturation at 94.degree. C. for 15 sec., annealing at 55.degree. C. for 15 sec. and extension at 72.degree. C. for 1 min. Following amplification, 2 to 4 PCR fragments for each variant were gel-purified, using a QIAGEN.RTM. gel-band purification kit and mixed (50 ng of each fragment). These mixtures served as DNA templates for the extension PCR to generate the full-length gene fragments. The PCR conditions were same as described above, except the extension phase, which was carried out at 72.degree. C. for 2 min. The full-length DNA fragment was gel-purified using a QIAGEN.RTM. gel-band purification kit, digested with the BamHI and HindIII restriction enzymes and ligated with the pHPLT-GG36, which was digested with the same restriction enzymes. One microliter of the ligation mixtures was amplified using rolling circle amplification by Illustra Templiphi kit according to the manufacturer's instructions (GE Healthcare) to generate multimeric DNA for transformation into Bacillus subtilis. Products of the rolling circle amplification were diluted 100-times and used to transform B. subtilis cells (genotype: AaprE, AnprE, amyE::xylRPxylAcomK-phleo). An aliquot of the transformation mix was plated on LB plates containing 1.6% skim milk and 10 .mu.g/mL neomycin and incubated overnight at 37.degree. C. Subsequently, the colonies with halos were inoculated in 120 .mu.l of Luria broth medium containing 10 .mu.g/mL neomycin for plasmid DNA extraction (QIAprep Spin Miniprep kit, Qiagen). The extracted plasmids were sequenced to confirm the presence of the desired mutations. Variants created by the extension PCR are shown in Table 4-1.
[0742] To express the NHJ4 set of variant proteins for further biochemical analyses, the B. subtilis strains carrying the variant plasmids were inoculated into microtiter plates containing 150 .mu.l Luria broth medium supplemented with 10 .mu.g/ml neomycin. The cultures were grown up for protein expression as described in Example 2, and they were filtered through a micro-filter plate (0.22 .mu.m; Millipore) also as described in Example 2. The resulting filtrate was used for biochemical analysis. The eglin c inhibition assay for protein content determination and BMI microswatch assays tested in various detergents were carried out as described in Example 1. Performance indices are also calculated as described under the BMI assay description in Example 1. Table 4-1 provides information regarding these multiple mutation variants and the results obtained for them. The PI values are relative to GG36. In the following Tables, the detergent compositions ("Det.") correspond to those shown in Table D, above. Also, as indicated, the amino acid position is listed according to BPN' numbering.
TABLE-US-00079 TABLE 4-1 NHJ4 Multiple Mutation Variants with BMI Cleaning Performance Indices of at Least 0.2 Relative to GG36 in Detergents 7, 8 or 9 at 16.degree. C. Mutations, (BPN' Numbering) Variant Name Created by GG36 NHJ4-1 Extension PCR S101G S103A V104I NHJ4-10 Extension PCR T22A S101A Y209E NHJ4-11 Extension PCR S103G L111V G159E NHJ4-12 Extension PCR T22A S103G G159E NHJ4-13 QCMS T22A L111V G159E NHJ4-14 QCMS T22A S128N E271F Y209E NHJ4-15 QCMS T22A S103G L111V NHJ4-16 QCMS N62E L111V S128N NHJ4-17 QCMS T22A L111V S128N NHJ4-18 Extension PCR T22A N62E L111V NHJ4-19 QCMS S101A S103G V104L S188D NHJ4-2 Extension PCR S101G S103A V104I G159D NHJ4-20 Extension PCR S101A S103G V104L S128N NHJ4-24 QCMS T22A S101A G159E NHJ4-3 Extension PCR S101A S103G V104L NHJ4-4 Extension PCR S101A S103G V104L G159E NHJ4-5 Extension PCR T22A S101A S103G V104L NHJ4-6 QCMS S101A S103G V104L Y209E NHJ4-7 QCMS T22A Y209E E271F NHJ4-8 QCMS T22A S101A E271F NHJ4-9 QCMS S101A Y209E E271F
Example 5
Construction and Cleaning Performance of NHJ3 Set of GG36 Variants
[0743] The NHJ3 set of variants described herein are based on a variant of GG36 (referred to as GG36-9) containing the following mutations: S101G, S103A, V104I, G159D, A232V, Q236H, Q245R, N248D, and N252K (BPN' numbering). These variants were created using the QUIKCHANGE.RTM. Lightning Site-Directed Mutagenesis Kit (QCLDS kit; Stratagene), with the pRA68 plasmid (see FIG. 7) as the DNA template. Plasmid pRA68 was derived from the pBN3 vector (see Babe et al., Biotech. Appl. Biochem. 27:117-124 [1998]).
[0744] The DNA sequence of GG36-9 variant (the signal sequence is shown in lower case letters, propeptide in lower case, underlined text, and GG36-9 mature sequence in uppercase letters) is provided below:
TABLE-US-00080 (SEQ ID NO: 762) Gtgagaagcaaaaaattgtggatcgtcgcgtcgaccgcactactcatttc tgttgcttttagttcatcgatcgcatcggctgctgaagaagcaaaagaaa aatatttaattggctttaatgagcaggaagctgtcagtgagtttgtagaa caagtagaggcaaatgacgaggtcgccattctctctgaggaagaggaagt cgaaattgaattgcttcatgaatttgaaacgattcctgttttatccgttg agttaagcccagaagatgtggacgcgcttgaactcgatccagcgatttct tatattgaagaggatgcagaagtaacgacaatgGCGCAATCAGTGCCATG GGGAATTAGCCGTGTGCAAGCCCCGGCTGCCCATAACCGTGGATTGACAG GTTCTGGTGTAAAAGTTGCTGTCCTCGATACAGGTATTTCCACTCATCCA GACTTAAATATTCGTGGTGGCGCTAGCTTTGTACCAGGGGAACCATCCAC TCAAGATGGGAATGGGCATGGCACGCATGTGGCCGGGACGATTGCTGCTC TAAACAATTCGATTGGCGTACTTGGCGTAGCGCCGAGCGCGGAACTATAC GCTGTTAAAGTATTAGGGGCGAGCGGTGGGGGCGCCATCAGCTCGATTGC CCAAGGATTGGAATGGGCAGGGAACAATGGCATGCACGTTGCTAATTTGA GTTTAGGAAGCCCTTCGCCAAGTGCCACACTTGAGCAAGCTGTTAATAGC GCGACTTCTAGGGGCGTTCTTGTTGTAGCGGCATCTGGAAATTCGGGTGC AGACTCAATCAGCTATCCGGCCCGTTATGCGAACGCAATGGCAGTCGGAG CTACTGACCAAAACAACAACCGCGCCAGCTTTTCACAGTATGGCGCAGGG CTTGACATCGTCGCACCAGGTGTAAACGTGCAGAGCACATACCCAGGTTC AACGTATGCCAGCTTAAACGGTACATCGATGGCTACTCCTCATGTTGCAG GTGCAGCAGTCCTTGTTAAACATAAGAACCCATCTTGGTCCAATGTACGA ATCCGCGATCATCTAAAGAAAACGGCAACGAGCTTAGGAAGCACGAACTT GTATGGAAGCGGACTTGTCAATGCCGAAGCTGCAACTCGTTAA
[0745] The protein sequence of the GG36-9 variant (the signal sequence is shown in lower case letters, propeptide in lower case, underlined text, and GG36-9 mature protease sequence in uppercase letters) is provided below:
TABLE-US-00081 (SEQ ID NO: 763) vrskklwivastallisvafsssiasaaeeakekyligfneqeavsefve qveandevailseeeeveiellhefetipvlsvelspedvdaleldpais yieedaevttmAQSVPWGISRVQAPAAHNRGLTGSGVKVAVLDTGISTHP DLNIRGGASFVPGEPSTQDGNGHGTHVAGTIAALNNSIGVLGVAPSAELY AVKVLGASGGGAISSIAQGLEWAGNNGMHVANLSLGSPSPSATLEQAVNS ATSRGVLVVAASGNSGADSISYPARYANAMAVGATDQNNNRASFSQYGAG LDIVAPGVNVQSTYPGSTYASLNGTSMATPHVAGAAVLVKHKNPSWSNVR IRDHLKKTATSLGSTNLYGSGLVNAEAATR
[0746] To create the NHJ3 variants using the QCLSD kit, mutagenic primers were designed for each of the variants according to the manufacturer's protocol. The mutagenesis reaction for each variant consisted of 0.5 .mu.l of 10.times. Buffer, 0.5 .mu.L of pRA68 plasmid DNA (168 ng/.mu.L), 0.50 forward mutagenic primer (25 .mu.M), 0.5 .mu.l reverse mutagenesis primer (25 .mu.M), 1 .mu.l dNTPs (supplied in the QCLSD kit), 1.5 .mu.l Quik solution (supplied in the QCLMS kit), 1 .mu.l Enzyme blend (supplied in the QCLSD kit), and 40 .mu.l of distilled, deionized water to make up a 50 .mu.L reaction volume as per the manufacturer's instructions. The cycling program was 1 cycle at 95.degree. C. for 2 minutes, 18 cycles of 95.degree. C. for 20 seconds, 60.degree. C. for 10 seconds and 68.degree. C. for 3 minutes, 22 seconds, and a final cycle of 68.degree. C. for 5 minutes. Next, 1 .mu.L of DpnI restriction enzyme supplied in the kit was used to digest the plasmid DNA in the reaction, and then 2 .mu.L of the reaction was used to transform TOP 10 E. coli competent cells (Invitrogen). The E. coli transformants were selected on Luria broth medium plates containing 50 .mu.g/mL(ppm) carbenicillin after overnight growth at 37.degree. C. Plasmid DNA was extracted from 4-8 E. coli colonies grown in LA medium containing 50 .mu.g/mL(ppm) carbenicillin using the QIAprep spin miniprep kit (Qiagen). The plasmids were sequenced to confirm the presence of the desired mutations. The variant plasmids were then transformed into B. subtilis cells as described in Example 2. The B. subtilis variant strains were grown up as described in Example 2 for further biochemical analysis, such as protein content determination using the eglin c inhibition assay (Example 1) and a BMI microswatch cleaning assay (Example 1). The results are provided below in Tables 5-1 and 5-2. The PIs are relative to GG36. In the following Tables, the detergent compositions ("Det.") correspond to those shown in Table D, above. Also, as indicated, the amino acid position is listed according to BPN' numbering.
TABLE-US-00082 TABLE 5-1 NHJ3 Multiple Mutation Variants with BMI Cleaning Performance Indices of at Least 1.1 Relative to GG36 in Detergents 7, 8, or 9 at 16.degree. C. Variant Sequence Relative to GG36 (Using BPN' Numbering) Variant GG36 NHJ3-1 S103A-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K NHJ3-2 S101G-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K NHJ3-3 S101G-S103A-G159D-A232V-Q236H-Q245R-N248D-N252K NHJ3-4 S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K NHJ3-5 S101G-S103A-V104L-G159D-Q236H-Q245R-N248D-N252K NHJ3-6 S101G-S103A-V104L-G159D-A232V-Q245R-N248D-N252K NHJ3-7 S101G-S103A-V104L-G159D-A232V-Q236H-N248D-N252K NHJ3-8 S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K NHJ3-9 S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N248D
TABLE-US-00083 TABLE 5-2 NHJ3 Multiple Mutation Variants with BMI Cleaning Performance Indices of at Least 0.3 Relative to GG36 in Detergents 10 or 11 at 16.degree. C. Variant Sequence Relative to GG36 (Using BPN' Numbering) Variant GG36 NHJ3-1 S103A-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K NHJ3-2 S101G-V104I-G159D-A232V-Q236H-Q245R-N248D-N252K NHJ3-3 S101G-S103A-G159D-A232V-Q236H-Q245R-N248D-N252K NHJ3-4 S101G-S103A-V104L-A232V-Q236H-Q245R-N248D-N252K NHJ3-5 S101G-S103A-V104L-G159D-Q236H-Q245R-N248D-N252K NHJ3-6 S101G-S103A-V104L-G159D-A232V-Q245R-N248D-N252K NHJ3-7 S101G-S103A-V104L-G159D-A232V-Q236H-N248D-N252K NHJ3-8 S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N252K NHJ3-9 S101G-S103A-V104L-G159D-A232V-Q236H-Q245R-N248D
Example 6
Construction and Cleaning Performance of NHJ5 Set of GG36 Variants
[0747] The NHJ5 set of variants described herein are based on a variant of GG36 (referred to as GG36-7) containing the following mutations: S101G, S103A, V104I, G159D, A232V, Q245R, N248D, and (BPN' numbering). These variants were created using the QUIKCHANGE.RTM. Lightning Multi Site-Directed Mutagenesis Kit ("QCLMS kit") with the pRA96 plasmid as the DNA template (see FIG. 8). The mutations incorporated included: H243R (H249R), E265F (E271F), D157E (D159E), A156E (A158E), A156E-D157G (A158E-D159E), T22A, N60E (N62E), N232R (N238R), D242R (D248R), T247R (T253R), S24R, N74D (N76D) {GG36 Numbering and BPN' Numbering Shown in Parentheses.
[0748] The variants were generated using the methods described in Example 5. The B. subtilis variant strains were grown up as described in Example 2 for further biochemical analysis, such as protein content determination using the eglin c inhibition assay (Example 1) and the BMI microswatch cleaning assay (Example 1). The results are provided below in Table 6-1. The PI values are relative to GG36. In the following Tables, the detergent compositions ("Det.") correspond to those shown in Table D, above. Also, as indicated, the amino acid position is listed according to BPN' numbering.
[0749] The DNA sequence of GG36-7 variant (the signal sequence is shown in lower case letters, propeptide in lower case, underlined text, and GG36-7 mature protease sequence in uppercase letters) is provided below:
TABLE-US-00084 (SEQ ID NO: 764) gtgagaagcaaaaaattgtggatcgtcgcgtcgaccgcactactcatttc tgttgcttttagttcatcgatcgcatcggctgctgaagaagcaaaagaaa aatatttaattggctttaatgagcaggaagctgtcagtgagtttgtagaa caagtagaggcaaatgacgaggtcgccattctctctgaggaagaggaagt cgaaattgaattgcttcatgaatttgaaacgattcctgttttatccgttg agttaagcccagaagatgtggacgcgcttgaactcgatccagcgatttct tatattgaagaggatgcagaagtaacgacaatgGCGCAATCAGTGCCATG GGGAATTAGCCGTGTGCAAGCCCCGGCTGCCCATAACCGTGGATTGACAG GTTCTGGTGTAAAAGTTGCTGTCCTCGATACAGGTATTTCCACTCATCCA GACTTAAATATTCGTGGTGGCGCTAGCTTTGTACCAGGGGAACCATCCAC TCAAGATGGGAATGGGCATGGCACGCATGTGGCCGGGACGATTGCTGCTC TAAACAATTCGATTGGCGTACTTGGCGTAGCGCCGAGCGCGGAACTATAC GCTGTTAAAGTATTAGGGGCGAGCGGTGGGGGCGCCATCAGCTCGATTGC CCAAGGATTGGAATGGGCAGGGAACAATGGCATGCACGTTGCTAATTTGA GTTTAGGAAGCCCTTCGCCAAGTGCCACACTTGAGCAAGCTGTTAATAGC GCGACTTCTAGGGGCGTTCTTGTTGTAGCGGCATCTGGAAATTCGGGTGC AGACTCAATCAGCTATCCGGCCCGTTATGCGAACGCAATGGCAGTCGGAG CTACTGACCAAAACAACAACCGCGCCAGCTTTTCACAGTATGGCGCAGGG CTTGACATCGTCGCACCAGGTGTAAACGTGCAGAGCACATACCCAGGTTC AACGTATGCCAGCTTAAACGGTACATCGATGGCTACTCCTCATGTTGCAG GTGCAGCAGTCCTTGTTAAACAAAAGAACCCATCTTGGTCCAATGTACGA ATCCGCGATCATCTAAAGAATACGGCAACGAGCTTAGGAAGCACGAACTT GTATGGAAGCGGACTTGTCAATGCCGAAGCTGCAACTCGT
[0750] The protein sequence of GG36-7 variant (signal sequence is shown in lower case letters, propeptide in lower case, underlined text, and GG36-7 mature protease sequence in uppercase letters) is provided below:
TABLE-US-00085 (SEQ ID NO: 765) vrskklwivastallisvafsssiasaaeeakekyligfneqeavsefve qveandevailseeeeveiellhefetipvlsvelspedvdaleldpais yieedaevttmAQSVPWGISRVQAPAAHNRGLTGSGVKVAVLDTGISTHP DLNIRGGASFVPGEPSTQDGNGHGTHVAGTIAALNNSIGVLGVAPSAELY AVKVLGASGGGAISSIAQGLEWAGNNGMHVANLSLGSPSPSATLEQAVNS ATSRGVLVVAASGNSGADSISYPARYANAMAVGATDQNNNRASFSQYGAG LDIVAPGVNVQSTYPGSTYASLNGTSMATPHVAGAAVLVKQKNPSWSNVR IRDHLKNTATSLGSTNLYGSGLVNAEAATR
TABLE-US-00086 TABLE 6-1 NHJ5 Multiple Mutation Variants with BMI Cleaning Performance Indices of at Least 0.6 Relative to GG36 in Detergents 7-11 at 16.degree. C. Variant Sequence Relative to GG36 (BPN' Numbering) Variant GG36 GG36-7 S101G-S103A-V104I-G159D-A232V-Q245R-N248D NHJ5-1 S101G-S103A-V104I-G159D-A232V-Q245R-N248D-E271F NHJ5-2 S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N238R NHJ5-3 S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N248R NHJ5-4 S101G-S103A-V104I-G159D-A232V-Q245R-N248D-T253R NHJ5-5 S101G-S103A-V104I-G159D-A232V-Q245R-N248D-S24R NHJ5-6 S101G-S103A-V104I-G159D-A232V-Q245R-N248D-N76D NHJ5-7 S101G-S103A-V104I-G159E-A232V-Q245R-N248D-H249R NHJ5-8 S101G-S103A-V104I-G159E-A232V-Q245R-N248D-E271F NHJ5-9 S101G-S103A-V104I-A158E-A232V-Q245R-N248D-H249R NHJ5-10 S101G-S103A-V104I-A158E-A232V-Q245R-N248D-E271F NHJ5-11 T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D- H249R NHJ5-12 T22A-S101G-S103A-V104I-G159D-A232V-Q245R-N248D- E271F NHJ5-13 N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D- H249R NHJ5-14 N62E-S101G-S103A-V104I-G159D-A232V-Q245R-N248D- E271F
Example 7
Construction of NHJ2 Combinatorial Library
[0751] This Example describes the construction of a GG36 combinatorial library involving one or more of the following mutations: A16S, T22A, S101A, S103G, V104L, L111V, S128N, and L148I (BPN' numbering). The pHPLT-GG36 B. subtilis expression plasmid was provided to DNA 2.0 Inc. for the generation of NHJ2 combinatorial library. A ligation reaction of the constructed NHJ2 library was provided by DNA 2.0, Inc. for transformation in the B. subtilis strain (genotype: AaprE, AnprE, amyE::xylRPxylAcomK-phleo). The variants generated containing one or several of the mutations described herein are tested for cold water cleaning applications using methods and detergent compositions described herein.
Example 8
Construction of Additional Libraries and GG36 Variants
[0752] Additional libraries and variants are constructed using the following set of mutations: A1R, A230E, E271L, G115R, G20R, H249R, K235F, K27V/F/L, L75E, L82R, N18R, N269R, N43D, N43R, N76D, R45T, S212F, S242R, S24R, S78R, S9A, T22R, V121E, V244R, V28E, V30E, V4R, W241R (BPN' numbering). The variants generated containing one or more of these mutations are tested for cold water cleaning applications using methods and detergent compositions described herein.
[0753] Additional sets of GG36 variants are constructed and tested for cold water cleaning applications using methods and detergent compositions described herein include: G20R-N43R-H249R, G20R-T22R-N43R, G20R-N43R-S242R, G20R-N43R-E271L, G20R-N43R-V244R, G20R-S24R-N43R-S242R, S9A-T22R-S78R-S212F-W241R, S9A-G20R-N43R-S212F, S9A-N43R-S212F, G20R-N43R-S212F, G20R-T22R-N43R-S212F, S24R-S78R-S212F, S9A-N43R-S78R, S9A-N43R-S78R-S242R, S9A-G20R-N43R-S78R, G20R-S24R-N43R-S78R-S242R, T22R-S24R-S78R-S212F, S9A-G20R-N43R-S78R-S242R, G20R-N43R-S78R-H249R, G20R-N43R-S78R, S9A-S78R-S212F, S9A-T22R-N43R-S78R, S9A-G20R-S24R-N43R, S9A-T22R-S78R-S212F, V4R-S9A-T22R-S78R-S212F, G20R-S24R-N43R, A1R-S9A-N43R, G20R-S24R-N43R-G115R, S9A-S24R-N43R, G20R-T22R-S24R-N43R, A1R-S24R-N43R, S9A-G20R-S24R-N43R-S242R, S9A-G20R-T22R-S78R-S212F, S9A-S24R-N43R-V244R, S9A-S24R-N43R-S242R, V4R-S9A-T22R-S24R-S212F, and T22R-S24R-N43R (BPN' numbering).
Example 9
Construction and Cleaning Performance of the GG36 Library WCE2
[0754] The WCE2 combinatorial library was generated by DNA 2.0, Inc. using the pHPLT-GG36 B. subtilis expression plasmid. A ligation reaction of the constructed WCE2 library was provided by DNA 2.0, Inc. for transformation in the B. subtilis strain (genotype: AaprE, AnprE, amyE::xylRPxylAcomK-phleo). The set of mutations used to generate the WCE2 library are A230E, G20R, H249R, N18R, N43R/D, N76D, R45T, S242R, and S24R (BPN' numbering). The variants generated containing one or more of these mutations are tested for cold water cleaning applications using methods and detergent compositions described herein.
Example 10
Construction and Cleaning Performance of the WCE3 Set of GG36 Variants
[0755] This Example describes the WCE3 set of mutants based on the GG36 variants, GG36-7 (Example 5) and GG36-9 (Example 4). These variants are: S101G-S103A-V104I-A232V-Q245R-N248D, S101G-S103A-V104I-G159D-A232V-Q245R, S101G-S103A-V104I-G159R-A232V-Q245R-N248D, S101G-S103A-V104I-G159D-A232V-Q245R-N248R, S101G-S103A-V104I-A232V-Q245R, S101G-S103A-V104I-A232V-Q245R-N248R, S101G-S103A-V104I-G159R-A232V-Q245R-N248R, and S101G, S103A, V104I, A232V, Q236H, Q245R, and N252K. They were created using the QuikChange.RTM. Lightning Multi Site-Directed Mutagenesis Kit (QCLMS kit; Stratagene) with the pRA96 plasmid as the DNA template described in Example 5. The variants generated will be tested for cold water cleaning applications using methods and detergent compositions described in this application.
Example 11
Construction of Additional Libraries and Variants of GG36
[0756] This Example describes the construction of GG36 variants and libraries using one or more of the following mutations: A16S, T22A, S24R, N62E, N76D, E89P, S101A/G, S103G/A, V104L/I, L111V, S128N, P129E, A232V, L148I, A158E, G159D/E, S166D, R186H, S188D, Y209E, Q236H, N238R, Q245R, N248D/R, H249R, N252K/R, T253R, E271F (BPN' numbering) using a B. subtilis expression plasmid (e.g., pHPLT-GG36; FIG. 6). The variants generated containing one or more of these mutations are tested for cold water cleaning applications using methods and detergent compositions described herein.
Example 12
Construction of Additional Libraries and Variants of GG36
[0757] This Example describes the construction of GG36 variants and libraries in B. subtilis using one or more of the following mutations (BPN' numbering): A1R, Q2S, Q2M, Q2A, Q2R, Q2W, S3R, V4R, V4S, V4C, I8A, S9A, S9F, S9W, R10S, R10A, R10H, R10M, Q12F, Q12R, P14K, P14F, P14Q, A15R, A15F, A16S, H17R, H17M, H17F, N18R, N18K, G20F, G20K, G20R, T22A, T22R, T22Y, T22V, T22Q, T22L, T22W, G23A, G23S, G23F, S24R, S24F, S24W, S24Q, S24H, S24L, G25V, G25F, G25R, V26F, K27L, K27F, K27R, K27V, V28A, V28N, V28E, A29T, V30E, L31F, T33S, T33G, T33D, G34P, I35M, S36T, S36F, S36R, T38L, T38F, T38R, P40N, P40L, P40T, P40W, P40H, P40R, L421, N43A, N43F, N43I, N43S, N43R, N43M, N43W, N43D, R45T, G46R, A48R, F50C, V51W, V51F, V51H, P52F, P52E, P52N, P55Y, T57R, Q59A, Q59F, Q59R, D60P, D60Q, D60A, N62E, N62Q, G63V, G63M, G63T, G631, G63A, G63S, G63H, G63Q, G63D, G63E, G63P, H64F, H64T, V68A, V68C, A69N, A69T, A69P, A69W, T71G, I72C, A74C, L75A, L75F, L75E, L75R, N76D, S78R, S78N, S781, 179W, I79Q, V81R, L82F, L82T, L82V, L82R, L82M, A85M, P86W, P86L, P861, E89P, E89T, E89G, E89H, E89L, E89V, E89W, E89F, E89I, Y91N, Y91F, A92F, K94N, S99F, S99T, S99P, S99G, S99M, G100S, G100N, G100Q, G100I, S101A, S101N, S101G, S101T, S101D, S101E, S101P, S101F, G102A, G102T, G102N, G102H, G102E, S103G, S103N, S103D, S103A, V104L, V104I, V104E, V104D, S105T, S105E, S105Q, S106G, S106T, S106E, S106D, S106A, S106V, S106F, I107M, I107F, A108I, A108G, Q109M, L111V, L111I, E112V, E112L, E112Q, A114G, G115K, G115R, N116K, N116A, N116L, N117F, G118R, G118I, M119C, H120A, H120F, H120R, V121F, V121E, N123G, N123E, L124S, S128D, S128F, S128L, S128N, S128H, S128M, S128I, S128Q, P129E, S132A, S132E, A138G, S144R, V147L, L148I, A158E, G159D, G159E, G159C, S160D, S166D, S166E, Y167W, M175V, V177C, D181A, Q182R, N183I, N183D, N183M, N183F, N183R, N185E, N185V, N185I, R186H, R186K, S188E, S188D, S188R, Y192H, Y192W, A194E, A194V, A194F, D197F, I198L, I198F, V203E, V203C, T208S, Y209S, Y209N, Y209F, Y209T, Y209E, Y209H, Y209G, Y209L, P210R, P210V, P210L, G211Q, G211R, S212I, S212M, S212F, T213A, Y214F, A215N, A215D, A215E, A215H, A215F, S216F, S216A, L217E, L217N, L217D, N218D, N218P, N218E, T224A, T224G, V227I, A230E, A231I, A231C, A232V, L233C, V234F, K235F, Q236F, Q236N, Q236H, N238R, N238K, N238L, P239K, P239G, P239R, P239H, P239T, P239N, P239S, P239F, S240R, W241R, S242L, S242R, N243F, N243R, V244R, Q245R, I246S, N248D, N248V, N248I, N248R, H249R, H249T, L250I, K251R, K251S, N252I, N252F, N252R, N252K, N252H, T253I, T253R, T253F, A254C, S256N, G258R, T260V, T260I, L262D, L262H, Y263F, S265F, L267V, L267N, L267M, N269I, N269R, A270C, E271I, E271V, E271H, E271M, E271L, E271P, E271A, E271F, E271T, A272F, A272F, A272R, A273F, A273I, and T274G. The variants generated containing one or more of these mutations are tested for cold water cleaning applications using methods and detergent compositions described herein.
Example 13
Automatic Dishwash Performance Tests
[0758] In this Example, methods used to determine the wash performance of protease variants using some commercially available dish detergents are described. These protease variants are tested under various conditions. These detergents are commercially available from WFK and are referred to by the designations provided below. The protocols for each of the stain types (minced meat, egg yolk, and egg yolk with milk) are provided below. Before the individual soil types can be applied to the test dishes, the dishes must be thoroughly washed. This is particularly necessary, as residues of certain persistent stains may still be present on the dishes from previous tests. New dishes were also subjected to three thorough washes before being used for the first time in a test.
[0759] The washing tests are typically performed in an automatic dishwasher (e.g., Miele: G690SC), equipped with soiled dishes and stainless steel sheets, as described above. A defined amount of the detergent is used, as indicated in the tables of results below. In some experiments, the temperatures tested are 45.degree. C., 55.degree. C. and 65.degree. C. In some experiments, the water hardness is 9.degree. or 21.degree. GH (German hardness) (374 ppm Ca).
[0760] As indicated above, after washing, the plates soiled with minced meat or pasta/sauce/meat/cheese are visually assessed using a photo rating scale of from 0 to 10, wherein "0" designated a completely dirty plate and "10" designated a clean plate. These values correspond to the stain or soil removal (SR) capability of the enzyme-containing detergent.
[0761] The washed stainless steel plates soiled with egg yolk and/or egg yolk milk (are analyzed gravimetrically to determine the amount of residual stain after washing.
[0762] Some exemplary detergents are provided below.
TABLE-US-00087 Phosphate-Free Detergent, IEC-60436 WFK Type B (pH = 10.4 in 3 g/l) Component Wt % Sodium citrate dehydrate, 30.0 Maleic acid/acrylic acid copolymer sodium Salt 12.0 (SOKALAN .RTM. CP5; BASF) Sodium perborate monohydrate, 5.0 TAED, 2.0 Sodium disilicate: Protil A (Cognis), 25.0 Linear fatty alcohol ethoxylate, 2.0 Sodium carbonate anhydrous, add to 100
TABLE-US-00088 Phosphate-Containing Detergent:, IEC-60436 WFK Type C (pH = 10.5 in 3 g/l)) Component Wt % Sodium tripolyphosphate 23.0 Sodium citrate dehydrate 22.3 Maleic acid/Acrylic Acid Copolymer Sodium 4.0 Salt Sodium perborate monohydrate 6.0 TAED 2.0 Sodium disilicate: Protil A (Cognis) 5.0 Linear Fatty Alcohol Ethoxylate 2.0 Sodium Carbonate anhydrous, add to 100,
Example 14
Granular and/or Tablet Laundry Compositions
[0763] This Example provides various formulations for granular and/or tablet laundry detergents. The following laundry compositions of present invention, which may be in the form of granules or tablet, are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00089 TABLE 14-1 Granular and/or Tablet Laundry Compositions Formulations Compound I II III IV V Base Product C.sub.14-C.sub.15AS or TAS 8.0 5.0 3.0 3.0 3.0 LAS 8.0 -- 8.0 -- 7.0 C.sub.12-C.sub.15AE.sub.3S 0.5 2.0 1.0 -- -- C.sub.12-C.sub.15E.sub.5 or E.sub.3 2.0 -- 5.0 2.0 2.0 QAS -- -- -- 1.0 1.0 Zeolite A 20.0 18.0 11.0 -- 10.0 SKS-6 (dry add) -- -- 9.0 -- -- MA/AA 2.0 2.0 2.0 -- -- AA -- -- -- -- 4.0 3Na Citrate 2H.sub.2O -- 2.0 -- -- -- Citric Acid (Anhydrous) 2.0 -- 1.5 2.0 -- DTPA 0.2 0.2 -- -- -- EDDS -- -- 0.5 0.1 -- HEDP -- -- 0.2 0.1 -- PB1 3.0 4.8 -- -- 4.0 Percarbonate -- -- 3.8 5.2 -- NOBS 1.9 -- -- -- -- NACA OBS -- -- 2.0 -- -- TAED 0.5 2.0 2.0 5.0 1.00 BB1 0.06 -- 0.34 -- 0.14 BB2 -- 0.14 -- 0.20 -- Anhydrous Na Carbonate 15.0 18.0 -- 15.0 15.0 Sulfate 5.0 12.0 5.0 17.0 3.0 Silicate -- 1.0 -- -- 8.0 nprE (optional) 0.03 -- 0.1 0.06 -- PMN -- 0.05 -- -- 0.1 Protease B (optional) -- 0.01 -- -- -- Protease C (optional) -- -- -- 0.01 -- Lipase -- 0.008 -- -- -- Amylase 0.001 -- -- -- 0.001 Cellulase -- 0.0014 -- -- -- Pectin Lyase 0.001 0.001 0.001 0.001 0.001 Aldose Oxidase 0.03 -- 0.05 -- -- PAAC -- 0.01 -- -- 0.05 Balance to 100% Moisture and/or Minors* *Perfume, dye, brightener/SRP1/Na carboxymethylcellulose/photobleach/MgSO.sub.4/PVPVI/suds suppressor/high molecular PEG/clay.
Example 15
Tablet Detergent Compositions
[0764] This Example provides various tablet detergent formulations. The following tablet detergent compositions of the present invention are prepared by compression of a granular dishwashing detergent composition at a pressure of 13KN/cm.sup.2 using a standard 12 head rotary press. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00090 TABLE 15-1 Tablet Detergent Compositions Formulations Compound I II III IV V VI VII VIII STPP -- 48.8 44.7 38.2 -- 42.4 46.1 46.0 3Na Citrate 2H.sub.2O 20.0 -- -- -- 35.9 -- -- -- Na Carbonate 20.0 5.0 14.0 15.4 8.0 23.0 20.0 -- Silicate 15.0 14.8 15.0 12.6 23.4 2.9 4.3 4.2 Lipase 0.001 -- 0.01 -- 0.02 -- -- -- Protease B 0.01 -- -- -- -- -- -- -- (optional) Protease C -- -- -- -- -- 0.01 -- -- (optional) nprE (optional) 0.01 0.08 -- 0.04 -- 0.023 -- 0.05 PMN -- -- 0.05 -- 0.052 -- 0.023 -- Amylase 0.012 0.012 0.012 -- 0.015 -- 0.017 0.002 Pectin Lyase 0.005 -- -- 0.002 -- -- -- -- Aldose Oxidase -- 0.03 -- 0.02 0.02 -- 0.03 -- PB1 -- -- 3.8 -- 7.8 -- -- 4.5 Percarbonate 6.0 -- -- 6.0 -- 5.0 -- -- BB1 0.2 -- 0.5 -- 0.3 0.2 -- -- BB2 -- 0.2 -- 0.5 -- -- 0.1 0.2 Nonionic 1.5 2.0 2.0 2.2 1.0 4.2 4.0 6.5 PAAC 0.01 0.01 0.02 -- -- -- -- -- DETBCHD -- -- -- 0.02 0.02 -- -- -- TAED -- -- -- -- -- 2.1 -- 1.6 HEDP 1.0 -- -- 0.9 -- 0.4 0.2 -- DETPMP 0.7 -- -- -- -- -- -- -- Paraffin 0.4 0.5 0.5 0.5 -- -- 0.5 -- BTA 0.2 0.3 0.3 0.3 0.3 0.3 0.3 -- Polycarboxylate 4.0 -- -- -- 4.9 0.6 0.8 -- PEG 400-30,000 -- -- -- -- -- 2.0 -- 2.0 Glycerol -- -- -- -- -- 0.4 -- 0.5 Perfume -- -- -- 0.05 0.2 0.2 0.2 0.2
[0765] The pH of Examples 14(I) through 14(VII) is from about 10 to about 11.5; pH of 14(VIII) is from 8-10. The tablet weight of Examples 14(I) through 14(VIII) is from about 20 grams to about 30 grams.
Example 16
Liquid Laundry Detergent Compositions
[0766] In this Example, various formulations for liquid laundry detergent compositions are provided. The following liquid laundry detergent compositions of the present invention are prepared as shown below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00091 TABLE 16-1 Liquid Laundry Detergent Compositions Formulations Compound I II III IV V LAS 24.0 32.0 6.0 3.0 6.0 NaC.sub.16-C.sub.17 HSAS -- -- -- 5.0 -- C.sub.12-C.sub.15 AE.sub.1.8S -- -- 8.0 7.0 5.0 C.sub.8-C.sub.10 propyl dimethyl 2.0 2.0 2.0 2.0 1.0 amine C.sub.12-C.sub.14 alkyl dimethyl -- -- -- -- 2.0 amine oxide C.sub.12-C.sub.15 AS -- -- 17.0 -- 8.0 CFAA -- 5.0 4.0 4.0 3.0 C.sub.12-C.sub.14 Fatty alcohol 12.0 6.0 1.0 1.0 1.0 ethoxylate C.sub.12-C.sub.18 Fatty acid 3.0 -- 4.0 2.0 3.0 Citric acid (anhydrous) 4.5 5.0 3.0 2.0 1.0 DETPMP -- -- 1.0 1.0 0.5 Monoethanolamine 5.0 5.0 5.0 5.0 2.0 Sodium hydroxide -- -- 2.5 1.0 1.5 1N HCl aqueous solution #1 #1 -- -- -- Propanediol 12.7 14.5 13.1 10. 8.0 Ethanol 1.8 2.4 4.7 5.4 1.0 DTPA 0.5 0.4 0.3 0.4 0.5 Pectin Lyase -- -- -- 0.005 -- Amylase 0.001 0.002 -- -- Cellulase -- -- 0.0002 0.0001 Lipase 0.1 -- 0.1 -- 0.1 NprE (optional) 0.05 0.3 -- 0.5 0.2 PMN -- -- 0.08 -- -- Protease A (optional) -- -- -- -- 0.1 Aldose Oxidase -- -- 0.3 -- 0.003 ZnCl2 0.1 0.05 0.05 0.05 0.02 Ca formate 0.05 0.07 0.05 0.06 0.07 DETBCHD -- -- 0.02 0.01 -- SRP1 0.5 0.5 -- 0.3 0.3 Boric acid -- -- -- -- 2.4 Sodium xylene sulfonate -- -- 3.0 -- -- Sodium cumene -- -- -- 0.3 0.5 sulfonate DC 3225C 1.0 1.0 1.0 1.0 1.0 2-butyl-octanol 0.03 0.04 0.04 0.03 0.03 Brightener 1 0.12 0.10 0.18 0.08 0.10 Balance to 100% perfume/dye and/or water #1: Add 1N HCl aqueous solution to adjust the neat pH of the formula in the range from about 3 to about 5.
[0767] The pH of Examples above 15(I)-(II) is about 5 to about 7, and of 15(III)-(V) is about 7.5 to about 8.5.
TABLE-US-00092 TABLE 16-2 Liquid Laundry Detergents Formulations Compound I II III IV V VI LAS 11.5 11.5 9.0 -- 4.0 -- C.sub.12-C.sub.15AE.sub.2.85S -- -- 3.0 18.0 -- 16.0 C.sub.14-C.sub.15E.sub.2.5S 11.5 11.5 3.0 -- 16.0 -- C.sub.12-C.sub.13E.sub.9 -- -- 3.0 2.0 2.0 1.0 C.sub.12-C.sub.13E.sub.7 3.2 3.2 -- -- -- -- CFAA -- -- -- 5.0 -- 3.0 TPKFA 2.0 2.0 -- 2.0 0.5 2.0 Citric Acid 3.2 3.2 0.5 1.2 2.0 1.2 (Anhydrous) Ca formate 0.1 0.1 0.06 0.1 -- -- Na formate 0.5 0.5 0.06 0.1 0.05 0.05 ZnCl2 0.1 0.05 0.06 0.03 0.05 0.05 Na Culmene 4.0 4.0 1.0 3.0 1.2 -- Sulfonate Borate 0.6 0.6 1.5 -- -- -- Na Hydroxide 6.0 6.0 2.0 3.5 4.0 3.0 Ethanol 2.0 2.0 1.0 4.0 4.0 3.0 1,2 3.0 3.0 2.0 8.0 8.0 5.0 Propanediol Mono- 3.0 3.0 1.5 1.0 2.5 1.0 ethanolamine TEPAE 2.0 2.0 -- 1.0 1.0 1.0 nprE 0.03 0.05 -- 0.03 -- 0.02 (optional) PMN -- -- 0.01 -- 0.08 -- Protease A -- -- 0.01 -- -- -- (optional) Lipase -- -- -- 0.002 -- -- Amylase -- -- -- -- 0.002 -- Cellulase -- -- -- -- -- 0.0001 Pectin Lyase 0.005 0.005 -- -- -- Aldose 0.05 -- -- 0.05 -- 0.02 Oxidase Galactose -- 0.04 oxidase PAAC 0.03 0.03 0.02 -- -- -- DETBCHD -- -- -- 0.02 0.01 -- SRP 1 0.2 0.2 -- 0.1 -- -- DTPA -- -- -- 0.3 -- -- PVNO -- -- -- 0.3 -- 0.2 Brightener 1 0.2 0.2 0.07 0.1 -- -- Silicone 0.04 0.04 0.02 0.1 0.1 0.1 antifoam Balance to 100% perfume/dye and/or water
Example 17
Hand Dish Liquid Detergent Compositions
[0768] In this Example, various hand dish liquid detergent formulations are provided. The following hand dish liquid detergent compositions of the invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some alternative aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00093 TABLE 17-1 Hand Dish Liquid Detergent Compositions Formulations Compound I II III IV V VI C.sub.12-C.sub.15 AE.sub.1.8S 30.0 28.0 25.0 -- 15.0 10.0 LAS -- -- -- 5.0 15.0 12.0 Paraffin Sulfonate -- -- -- 20.0 -- -- C.sub.10-C.sub.18 Alkyl Dimethyl Amine 5.0 3.0 7.0 -- -- -- Oxide Betaine 3.0 -- 1.0 3.0 1.0 -- C.sub.12 poly-OH fatty acid amide -- -- -- 3.0 -- 1.0 C.sub.14 poly-OH fatty acid amide -- 1.5 -- -- -- -- C.sub.11E.sub.9 2.0 -- 4.0 -- -- 20.0 DTPA -- -- -- -- 0.2 -- Tri-sodium Citrate dehydrate 0.25 -- -- 0.7 -- -- Diamine 1.0 5.0 7.0 1.0 5.0 7.0 MgCl.sub.2 0.25 -- -- 1.0 -- -- nprE (optional) 0.02 0.01 -- 0.01 -- 0.05 PMN -- -- 0.03 -- 0.02 -- Protease A (optional) -- 0.01 -- -- -- -- Amylase 0.001 -- -- 0.002 -- 0.001 Aldose Oxidase 0.03 -- 0.02 -- 0.05 -- Sodium Cumene Sulphonate -- -- -- 2.0 1.5 3.0 PAAC 0.01 0.01 0.02 -- -- -- DETBCHD -- -- -- 0.01 0.02 0.01 Balance to 100% perfume/dye and/or water
[0769] The pH of Examples 16(I)-(VI) is about 8 to about 11.
Example 18
Liquid Automatic Dishwashing Detergent Compositions
[0770] In this Example, various liquid automatic dishwashing detergent formulations are provided. The following hand dish liquid detergent compositions of the invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00094 TABLE 18-1 Liquid Automatic Dishwashing Detergent Compositions Formulations Compound I II III IV V STPP 16 16 18 16 16 Potassium Sulfate -- 10 8 -- 10 1,2 propanediol 6.0 0.5 2.0 6.0 0.5 Boric Acid -- -- -- 4.0 3.0 CaCl.sub.2 dihydrate 0.04 0.04 0.04 0.04 0.04 Nonionic 0.5 0.5 0.5 0.5 0.5 nprE (optional) 0.1 0.03 -- 0.03 -- PMN -- -- 0.05 -- 0.06 Protease B (optional) -- -- -- 0.01 -- Amylase 0.02 -- 0.02 0.02 -- Aldose Oxidase -- 0.15 0.02 -- 0.01 Galactose Oxidase -- -- 0.01 -- 0.01 PAAC 0.01 -- -- 0.01 -- DETBCHD -- 0.01 -- -- 0.01 Balance to 100% perfume/dye and/or water
Example 19
High Density Dishwashing Detergents
[0771] This Example provides various formulations for high density dishwashing detergents. The following compact high density dishwashing detergents of the invention are provided below. In each formulation, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00095 TABLE 19-1 High Density Dishwashing Detergents Formulations Compound I II III IV V VI STPP -- 45.0 45.0 -- -- 40.0 3Na Citrate 2H.sub.2O 17.0 -- -- 50.0 40.2 -- Na Carbonate 17.5 14.0 20.0 -- 8.0 33.6 Bicarbonate -- -- -- 26.0 -- -- Silicate 15.0 15.0 8.0 -- 25.0 3.6 Metasilicate 2.5 4.5 4.5 -- -- -- PB1 -- -- 4.5 -- -- -- PB4 -- -- -- 5.0 -- -- Percarbonate -- -- -- -- -- 4.8 BB1 -- 0.1 0.1 -- 0.5 -- BB2 0.2 0.05 -- 0.1 -- 0.6 Nonionic 2.0 1.5 1.5 3.0 1.9 5.9 HEDP 1.0 -- -- -- -- -- DETPMP 0.6 -- -- -- -- -- PAAC 0.03 0.05 0.02 -- -- -- Paraffin 0.5 0.4 0.4 0.6 -- -- nprE (optional) 0.072 0.053 -- 0.026 -- 0.01 PMN -- -- 0.053 -- 0.059 -- Protease B -- -- -- -- -- 0.01 (optional) Amylase 0.012 -- 0.012 -- 0.021 0.006 Lipase -- 0.001 -- 0.005 -- -- Pectin Lyase 0.001 0.001 0.001 -- -- -- Aldose Oxidase 0.05 0.05 0.03 0.01 0.02 0.01 BTA 0.3 0.2 0.2 0.3 0.3 0.3 Polycarboxylate 6.0 -- -- -- 4.0 0.9 Perfume 0.2 0.1 0.1 0.2 0.2 0.2 Balance to 100% Moisture and/or Minors* *Brightener/dye/SRP 1/Na carboxymethylcellulose/photobleach/MgSO.sub.4/PVPVI/suds suppressor/high molecular PEG/clay.
The pH of Examples 18(I) through (VI) is from about 9.6 to about 11.3.
Example 20
Liquid Hard Surface Cleaning Detergents
[0772] This Example provides various formulations for liquid hard surface cleaning detergents. The following liquid hard surface cleaning detergent compositions of the invention are provided below. In each of these formulations, at least one protease variant provided herein is included at a concentration of from about 0.0001 to about 10 weight percent. In some aspects, other concentrations will find use, as determined by the formulator, based on their needs.
TABLE-US-00096 TABLE 20-1 Liquid Hard Surface Cleaning Detergents Formulations Compound I II III IV V VI VII C.sub.9-C.sub.11E.sub.5 2.4 1.9 2.5 2.5 2.5 2.4 2.5 C.sub.12-C.sub.14E.sub.5 3.6 2.9 2.5 2.5 2.5 3.6 2.5 C.sub.7-C.sub.9E.sub.6 -- -- -- -- 8.0 -- -- C.sub.12-C.sub.14E.sub.21 1.0 0.8 4.0 2.0 2.0 1.0 2.0 LAS -- -- -- 0.8 0.8 -- 0.8 Sodium culmene sulfonate 1.5 2.6 -- 1.5 1.5 1.5 1.5 Isachem .RTM. AS 0.6 0.6 -- -- -- 0.6 -- Na.sub.2CO.sub.3 0.6 0.13 0.6 0.1 0.2 0.6 0.2 3Na Citrate.cndot.2H.sub.2O 0.5 0.56 0.5 0.6 0.75 0.5 0.75 NaOH 0.3 0.33 0.3 0.3 0.5 0.3 0.5 Fatty Acid 0.6 0.13 0.6 0.1 0.4 0.6 0.4 2-butyl octanol 0.3 0.3 -- 0.3 0.3 0.3 0.3 PEG DME-2000 .RTM. 0.4 -- 0.3 0.35 0.5 -- -- PVP 0.3 0.4 0.6 0.3 0.5 -- -- MME PEG (2000) .RTM. -- -- -- -- -- 0.5 0.5 Jeffamine .RTM. ED-2001 -- 0.4 -- -- 0.5 -- -- PAAC -- -- -- 0.03 0.03 0.03 -- DETBCHD 0.03 0.05 0.05 -- -- -- -- nprE (optional) 0.07 -- 0.08 0.03 -- 0.01 0.04 PMN -- 0.05 -- -- 0.06 -- -- Protease B (optional) -- -- -- -- -- 0.01 -- Amylase 0.12 0.01 0.01 -- 0.02 -- 0.01 Lipase -- 0.001 -- 0.005 -- 0.005 -- Pectin Lyase 0.001 -- 0.001 -- -- -- 0.002 ZnCl2 0.02 0.01 0.03 0.05 0.1 0.05 0.02 Calcium Formate 0.03 0.03 0.01 -- -- -- -- PB1 -- 4.6 -- 3.8 -- -- -- Aldose Oxidase 0.05 -- 0.03 -- 0.02 0.02 0.05 Balance to 100% perfume/dye and/or water
[0773] The pH of Examples 19(I) through (VII) is from about 7.4 to about 9.5.
[0774] All publications, patent applications, and patents cited in this specification are herein incorporated by reference as if each individual publication, patent application, or patent were specifically and individually indicated to be incorporated by reference. In particular, all publications cited herein are expressly incorporated herein by reference for the purpose of describing and disclosing compositions and methodologies which might be used in connection with the invention. Although the foregoing invention has been described in some detail by way of illustration and example for purposes of clarity of understanding, it will be readily apparent to those of ordinary skill in the art in light of the teachings of this invention that certain changes and modifications may be made thereto without departing from the spirit or scope of the appended claims.
Sequence CWU 1
1
76611146DNABacillus amyloliquefaciens 1gtgagaagca aaaaattgtg
gatcagtttg ctgtttgctt tagcgttaat ctttacgatg 60gcgttcggca gcacatcctc
tgcccaggcg gcagggaaat caaacgggga aaagaaatat 120attgtcgggt
ttaaacagac aatgagcacg atgagcgccg ctaagaagaa agatgtcatt
180tctgaaaaag gcgggaaagt gcaaaagcaa ttcaaatatg tagacgcagc
ttcagctaca 240ttaaacgaaa aagctgtaaa agaattgaaa aaagacccga
gcgtcgctta cgttgaagaa 300gatcacgtag cacatgcgta cgcgcagtcc
gtgccttacg gcgtatcaca aattaaagcc 360cctgctctgc actctcaagg
ctacactgga tcaaatgtta aagtagcggt tatcgacagc 420ggtatcgact
cgagccatcc agatcttaaa gtcgctggag gggcttctat ggtgccgtcc
480gaaacaaacc cgtttcaaga taacaattct catggcacac acgtcgcagg
aacggttgcg 540gcgttaaaca attctattgg cgtgcttggt gtagccccgt
ctgcttcgct ctacgccgtt 600aaagttcttg gcgcagacgg atcaggccaa
tactcatgga ttatcaacgg catcgaatgg 660gccatcgcga ataacatgga
tgtaatcaac atgagcctgg gaggaccaag cggcagtgcg 720gcacttaaag
cagcagttga taaagctgtt gcatctggtg tcgtcgtagt agcggcagct
780gggaatgagg gaacatccgg atcatcgagt accgtcggtt atccaggcaa
gtacccttca 840gtgattgcag tgggcgctgt agactcttca aatcaacgtg
cctctttttc ctccgtggga 900ccggagctgg atgtcatggc ccctggcgtt
tctattcaat cgacgcttcc agggaacaag 960tatggtgcgt ataacgggac
ttccatggcc tcgccgcatg tagctggggc ggccgcattg 1020attctttcta
agcacccgaa ctggacaaac actcaagtcc gcagcagttt agaaaacacc
1080actacaaaac ttggtgattc tttctactat ggaaaagggc tgatcaacgt
acaggcggca 1140gctcag 11462275PRTBacillus amyloliquefaciens 2Ala
Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1 5 10
15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile Asp
20 25 30Ser Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly Gly
Ala 35 40 45Ser Met Val Pro Ser Glu Thr Asn Pro Phe Gln Asp Asn Asn
Ser His 50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn Asn
Ser Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu Tyr
Ala Val Lys Val Leu 85 90 95Gly Ala Asp Gly Ser Gly Gln Tyr Ser Trp
Ile Ile Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met Asp
Val Ile Asn Met Ser Leu Gly Gly 115 120 125Pro Ser Gly Ser Ala Ala
Leu Lys Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val Val
Val Val Ala Ala Ala Gly Asn Glu Gly Thr Ser Gly145 150 155 160Ser
Ser Ser Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala 165 170
175Val Gly Ala Val Asp Ser Ser Asn Gln Arg Ala Ser Phe Ser Ser Val
180 185 190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser Ile Gln
Ser Thr 195 200 205Leu Pro Gly Asn Lys Tyr Gly Ala Tyr Asn Gly Thr
Ser Met Ala Ser 210 215 220Pro His Val Ala Gly Ala Ala Ala Leu Ile
Leu Ser Lys His Pro Asn225 230 235 240Trp Thr Asn Thr Gln Val Arg
Ser Ser Leu Glu Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp Ser Phe
Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala Ala Gln
2753825DNABacillus amyloliquefaciens 3gcgcagtccg tgccttacgg
cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat caaatgttaa
agtagcggtt atcgacagcg gtatcgactc gagccatcca 120gatcttaaag
tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc gtttcaagat
180aacaattctc atggcacaca cgtcgcagga acggttgcgg cgttaaacaa
ttctattggc 240gtgcttggtg tagccccgtc tgcttcgctc tacgccgtta
aagttcttgc agcagacgga 300tcaggccaat actcatggat tatcaacggc
atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca tgagcctggg
agcaccaagc ggcagtgcgg cacttaaagc agcagttgat 420aaagctgttg
catctggtgt cgtcgtagta gcggcagctg ggaatgaggg aacatccgga
480tcatcgagta ccgtcggtta tccaggcaag tacccttcag tgattgcagt
gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc tccgtgggac
cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc gacgcttcca
gggaacaagt atggtgcgca aaacgggact 660tccatggcct cgccgcatgt
agctggggcg gccgcattga ttctttctaa gcacccgaac 720tggacaaaca
ctcaagtccg cagcagttta gaaaacacca ctacaaaact tggtgattct
780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcag
8254275PRTBacillus amyloliquefaciens 4Ala Gln Ser Val Pro Tyr Gly
Val Ser Gln Ile Lys Ala Pro Ala Leu1 5 10 15His Ser Gln Gly Tyr Thr
Gly Ser Asn Val Lys Val Ala Val Ile Asp 20 25 30Ser Gly Ile Asp Ser
Ser His Pro Asp Leu Lys Val Ala Gly Gly Ala 35 40 45Ser Met Val Pro
Ser Glu Thr Asn Pro Phe Gln Asp Asn Asn Ser His 50 55 60Gly Thr His
Val Ala Gly Thr Val Ala Ala Leu Asn Asn Ser Ile Gly65 70 75 80Val
Leu Gly Val Ala Pro Ser Ala Ser Leu Tyr Ala Val Lys Val Leu 85 90
95Ala Ala Asp Gly Ser Gly Gln Tyr Ser Trp Ile Ile Asn Gly Ile Glu
100 105 110Trp Ala Ile Ala Asn Asn Met Asp Val Ile Asn Met Ser Leu
Gly Ala 115 120 125Pro Ser Gly Ser Ala Ala Leu Lys Ala Ala Val Asp
Lys Ala Val Ala 130 135 140Ser Gly Val Val Val Val Ala Ala Ala Gly
Asn Glu Gly Thr Ser Gly145 150 155 160Ser Ser Ser Thr Val Gly Tyr
Pro Gly Lys Tyr Pro Ser Val Ile Ala 165 170 175Val Gly Ala Val Asp
Ser Ser Asn Gln Arg Ala Ser Phe Ser Ser Val 180 185 190Gly Pro Glu
Leu Asp Val Met Ala Pro Gly Val Ser Ile Gln Ser Thr 195 200 205Leu
Pro Gly Asn Lys Tyr Gly Ala Gln Asn Gly Thr Ser Met Ala Ser 210 215
220Pro His Val Ala Gly Ala Ala Ala Leu Ile Leu Ser Lys His Pro
Asn225 230 235 240Trp Thr Asn Thr Gln Val Arg Ser Ser Leu Glu Asn
Thr Thr Thr Lys 245 250 255Leu Gly Asp Ser Phe Tyr Tyr Gly Lys Gly
Leu Ile Asn Val Gln Ala 260 265 270Ala Ala Gln
2755825DNAArtificialsynthesized BPN'-v36 variant 5gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggag
gcaatgttaa agtagcggtt atcgacagcg gtatcgactc gagccatcca
120gatcttaaag tcgctggagg ggcttctatg gtgccgggcg aaacaaaccc
gtttcaagat 180aacaattctc atggcacaca cgtcgcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc
tacgccgtta aagttcttgg cgcagacgga 300aatggccaat actcatggat
tatcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tgagcctggg agcaccaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtgt cgtcgtagta gcggcagctg ggaatgaggg
aacatccgga 480tcatcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gggaacaagt atggtgcgca aaacgggact 660tccatggcct
cgccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgaac
720tggacaaaca ctcaagtccg cagcagttta gaaaacacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcag
8256275PRTArtificialsynthesized BPN'-v36 variant 6Ala Gln Ser Val
Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1 5 10 15His Ser Gln
Gly Tyr Thr Gly Gly Asn Val Lys Val Ala Val Ile Asp 20 25 30Ser Gly
Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly Gly Ala 35 40 45Ser
Met Val Pro Gly Glu Thr Asn Pro Phe Gln Asp Asn Asn Ser His 50 55
60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn Asn Asn Ile Gly65
70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu Tyr Ala Val Lys Val
Leu 85 90 95Gly Ala Asp Gly Asn Gly Gln Tyr Ser Trp Ile Ile Asn Gly
Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met Asp Val Ile Asn Met
Ser Leu Gly Ala 115 120 125Pro Ser Gly Ser Ala Ala Leu Lys Ala Ala
Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val Val Val Val Ala Ala
Ala Gly Asn Glu Gly Thr Ser Gly145 150 155 160Ser Ser Ser Thr Val
Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala 165 170 175Val Gly Ala
Val Asp Ser Ser Asn Gln Arg Ala Ser Phe Ser Ser Val 180 185 190Gly
Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser Ile Gln Ser Thr 195 200
205Leu Pro Gly Asn Lys Tyr Gly Ala Gln Asn Gly Thr Ser Met Ala Ser
210 215 220Pro His Val Ala Gly Ala Ala Ala Leu Ile Leu Ser Lys His
Pro Asn225 230 235 240Trp Thr Asn Thr Gln Val Arg Ser Ser Leu Glu
Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp Ser Phe Tyr Tyr Gly Lys
Gly Leu Ile Asn Val Gln Ala 260 265 270Ala Ala Gln
2757825DNAArtificialsynthesized BPN'-v3+S78N variant 7gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat
caaatgttaa agtagcggtt atcgacagcg gtattgattc gagccatcca
120gatcttaaag tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc
gtttcaagat 180aacaattctc atggcacaca cgtcgcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc
tacgccgtta aagttcttgc agcagacgga 300tcaggccaat actcatggat
tatcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tgagcctggg agcaccaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtgt cgtcgtagta gcggcagctg ggaatgaggg
aacatccgga 480tcatcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gggaacaagt atggtgcgca aaacgggact 660tccatggcct
cgccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgaac
720tggacaaaca ctcaagtccg cagcagttta gaaaacacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcag
8258275PRTArtificialsynthesized BPN'-v3+S78N variant 8Ala Gln Ser
Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1 5 10 15His Ser
Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile Asp 20 25 30Ser
Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly Gly Ala 35 40
45Ser Met Val Pro Ser Glu Thr Asn Pro Phe Gln Asp Asn Asn Ser His
50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn Asn Asn Ile
Gly65 70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu Tyr Ala Val
Lys Val Leu 85 90 95Ala Ala Asp Gly Ser Gly Gln Tyr Ser Trp Ile Ile
Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met Asp Val Ile
Asn Met Ser Leu Gly Ala 115 120 125Pro Ser Gly Ser Ala Ala Leu Lys
Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val Val Val Val
Ala Ala Ala Gly Asn Glu Gly Thr Ser Gly145 150 155 160Ser Ser Ser
Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala 165 170 175Val
Gly Ala Val Asp Ser Ser Asn Gln Arg Ala Ser Phe Ser Ser Val 180 185
190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser Ile Gln Ser Thr
195 200 205Leu Pro Gly Asn Lys Tyr Gly Ala Gln Asn Gly Thr Ser Met
Ala Ser 210 215 220Pro His Val Ala Gly Ala Ala Ala Leu Ile Leu Ser
Lys His Pro Asn225 230 235 240Trp Thr Asn Thr Gln Val Arg Ser Ser
Leu Glu Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp Ser Phe Tyr Tyr
Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala Ala Gln
2759828DNAArtificialsynthesized GcM90 gene 9gcgcagtccg tgccttacgg
cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat caaatgttaa
agtagcggtt atcgacagcg gtatcgactc gagccatcca 120gatcttaaag
tcgctggagg ggcttctatg gtgccgggag aaacaaaccc gtttcaagat
180aacaattctc atggcacaca cgcagcagga acggttgcgg cgttaaacaa
taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc tacgccgtta
aagttcttgc agcagacgga 300tcagcacaat actcatggat tatcaacggc
atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca tgagcctggg
agcaacaagc ggcagtgcgg cacttaaagc agcagttgat 420aaagctgttg
catctggtgt cgtcgtagta gcggcagctg ggaatgaggg aacatccgga
480tcatcgagta ccgtcggtta tccaggcaag tacccttcag tgattgcagt
gggcgctgta 540gactcttcaa atacacgtgc ctctttttcc tccgtgggac
cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc gacgcttcca
gggaacaagt atggtgcgca aaacgggact 660tccatggcct cgccgcatgt
agctggggcg gccgcattga ttctttctaa gcacccgaac 720tggacaaaca
ctcaagtccg cagcagttta gaaaacacca ctacaaaact tggtgattct
780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82810275PRTArtificialamino acid encoded by synthesized GcM90 gene
10Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile
Asp 20 25 30Ser Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly
Gly Ala 35 40 45Ser Met Val Pro Gly Glu Thr Asn Pro Phe Gln Asp Asn
Asn Ser His 50 55 60Gly Thr His Ala Ala Gly Thr Val Ala Ala Leu Asn
Asn Asn Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu
Tyr Ala Val Lys Val Leu 85 90 95Ala Ala Asp Gly Ser Ala Gln Tyr Ser
Trp Ile Ile Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met
Asp Val Ile Asn Met Ser Leu Gly Ala 115 120 125Thr Ser Gly Ser Ala
Ala Leu Lys Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val
Val Val Val Ala Ala Ala Gly Asn Glu Gly Thr Ser Gly145 150 155
160Ser Ser Ser Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala
165 170 175Val Gly Ala Val Asp Ser Ser Asn Thr Arg Ala Ser Phe Ser
Ser Val 180 185 190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser
Ile Gln Ser Thr 195 200 205Leu Pro Gly Asn Lys Tyr Gly Ala Gln Asn
Gly Thr Ser Met Ala Ser 210 215 220Pro His Val Ala Gly Ala Ala Ala
Leu Ile Leu Ser Lys His Pro Asn225 230 235 240Trp Thr Asn Thr Gln
Val Arg Ser Ser Leu Glu Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp
Ser Phe Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala
Ala Gln 27511828DNAArtificialsynthesized GcM91 gene 11gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat
caaatgttaa agtagcggtt atcgacagcg gtatcgactc gagccatcca
120gatcttaaag tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc
gtttgtcgat 180aacaattctc atggcacaca cgtcgcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc
tacgccgtta aagttcttgc agcagacgga 300tcaggccaat actcatggat
tgtcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tgagcctggg agcaccaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtca agtcgtagta gcggcagctg ggaatgaggg
aacatccgga 480tcatcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gcaaacaagt atggtgcgca aaacgggact 660tccatggcct
cgccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgaac
720tggacaaaca ctcaagtccg cagcagttta gaacaaacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82812275PRTArtificialamino acid encoded by synthesized GcM91 gene
12Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile
Asp 20 25 30Ser Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly
Gly Ala 35 40 45Ser Met Val Pro Ser Glu Thr Asn Pro Phe Val Asp Asn
Asn Ser His 50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn
Asn Asn Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu
Tyr Ala Val Lys Val Leu 85 90 95Ala Ala Asp Gly Ser Gly Gln Tyr Ser
Trp Ile Val Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met
Asp Val Ile Asn Met
Ser Leu Gly Ala 115 120 125Pro Ser Gly Ser Ala Ala Leu Lys Ala Ala
Val Asp Lys Ala Val Ala 130 135 140Ser Gly Gln Val Val Val Ala Ala
Ala Gly Asn Glu Gly Thr Ser Gly145 150 155 160Ser Ser Ser Thr Val
Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala 165 170 175Val Gly Ala
Val Asp Ser Ser Asn Gln Arg Ala Ser Phe Ser Ser Val 180 185 190Gly
Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser Ile Gln Ser Thr 195 200
205Leu Pro Ala Asn Lys Tyr Gly Ala Gln Asn Gly Thr Ser Met Ala Ser
210 215 220Pro His Val Ala Gly Ala Ala Ala Leu Ile Leu Ser Lys His
Pro Asn225 230 235 240Trp Thr Asn Thr Gln Val Arg Ser Ser Leu Glu
Gln Thr Thr Thr Lys 245 250 255Leu Gly Asp Ser Phe Tyr Tyr Gly Lys
Gly Leu Ile Asn Val Gln Ala 260 265 270Ala Ala Gln
27513828DNAArtificialsynthesized GcM92 gene 13gcgcagtccg tgccttacgg
cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat caaatgttaa
agtagcggtt atcgacagcg gtatcgactc gagccatcca 120gatcttaaag
tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc gtttcaagat
180gcaaattctc atggcacaca cgtcgcagga acggttgcgg cgttaaacaa
taatattggc 240gtgcttggtg tagccccgga agcttcgctc tacgccgtta
aagttcttgc agcagacgga 300tcaggccaat actcatggat tatcaacggc
atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca tcagcctggg
agcaccaagc ggcagtgcgg cacttaaagc agcagttgat 420aaagctgttg
catctggtgt cgtcgtagta gcggcagctg ggaatgaggg aacatccgga
480ccttcgagta ccgtcggtta tccaggcaag tacccttcag tgattgcagt
gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc tccgtgggac
cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc gacgcttcca
gggaacaagt atggtgcgca aaacgggact 660tccatggccg caccgcatgt
agctggggcg gccgcattga ttctttctaa gcacccgaac 720tggacaaaca
ctcaagtccg cagcagttta gaaaacacca ctacaaaact tggtgattct
780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82814275PRTArtificialamino acid encoded by synthesized GcM92 gene
14Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile
Asp 20 25 30Ser Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly
Gly Ala 35 40 45Ser Met Val Pro Ser Glu Thr Asn Pro Phe Gln Asp Ala
Asn Ser His 50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn
Asn Asn Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Glu Ala Ser Leu
Tyr Ala Val Lys Val Leu 85 90 95Ala Ala Asp Gly Ser Gly Gln Tyr Ser
Trp Ile Ile Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met
Asp Val Ile Asn Ile Ser Leu Gly Ala 115 120 125Pro Ser Gly Ser Ala
Ala Leu Lys Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val
Val Val Val Ala Ala Ala Gly Asn Glu Gly Thr Ser Gly145 150 155
160Pro Ser Ser Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala
165 170 175Val Gly Ala Val Asp Ser Ser Asn Gln Arg Ala Ser Phe Ser
Ser Val 180 185 190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser
Ile Gln Ser Thr 195 200 205Leu Pro Gly Asn Lys Tyr Gly Ala Gln Asn
Gly Thr Ser Met Ala Ala 210 215 220Pro His Val Ala Gly Ala Ala Ala
Leu Ile Leu Ser Lys His Pro Asn225 230 235 240Trp Thr Asn Thr Gln
Val Arg Ser Ser Leu Glu Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp
Ser Phe Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala
Ala Gln 27515828DNAArtificialsynthesized GcM93 gene 15gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat
caaatgttaa agtagcggtt atcgacagcg gtatcgactc gagccatcca
120gatcttaaag tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc
gtttcaagat 180aaccaatctc atggcacaca cgtcgcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc
tacgccgtta aagttcttgc agcagacaac 300tcaggccaat actcatggat
tatcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tggcactggg agcaccaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtgt cgtcgtagta gcggcagctg ggaatgaggg
aacagatgga 480tcatcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gggaacaagt atggtgcgca aaacgggact 660tccatggcct
cgccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgtca
720tggacaaaca ctcaagtccg cagcagttta gaaaacacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82816275PRTArtificialamino acid encoded by synthesized GcM93 gene
16Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile
Asp 20 25 30Ser Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly
Gly Ala 35 40 45Ser Met Val Pro Ser Glu Thr Asn Pro Phe Gln Asp Asn
Gln Ser His 50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn
Asn Asn Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu
Tyr Ala Val Lys Val Leu 85 90 95Ala Ala Asp Asn Ser Gly Gln Tyr Ser
Trp Ile Ile Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met
Asp Val Ile Asn Met Ala Leu Gly Ala 115 120 125Pro Ser Gly Ser Ala
Ala Leu Lys Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val
Val Val Val Ala Ala Ala Gly Asn Glu Gly Thr Asp Gly145 150 155
160Ser Ser Ser Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala
165 170 175Val Gly Ala Val Asp Ser Ser Asn Gln Arg Ala Ser Phe Ser
Ser Val 180 185 190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser
Ile Gln Ser Thr 195 200 205Leu Pro Gly Asn Lys Tyr Gly Ala Gln Asn
Gly Thr Ser Met Ala Ser 210 215 220Pro His Val Ala Gly Ala Ala Ala
Leu Ile Leu Ser Lys His Pro Ser225 230 235 240Trp Thr Asn Thr Gln
Val Arg Ser Ser Leu Glu Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp
Ser Phe Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala
Ala Gln 27517828DNAArtificialsynthesized GcM94 gene 17gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat
caaatgttaa agtagcggtt atcgacagcg gtatcgactc gagccatcca
120gatcttaaag tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc
gtttcaagat 180aacaatacac atggcacaca cgtcgcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc
tacgccgtta aagttcttgc agcagacgga 300gcaggccaat actcatggat
tatcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tgagcgtcgg agcaccaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtgt cgtcgtagta gcggcagctg ggaatgaggg
aacatccgga 480tcatcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactctacaa atcaacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gggaacaagt atggtgcgca aaacgggact 660tccatggcct
cgccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgaac
720tggacaaaca accaagtccg cagcagttta gaaaacacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82818275PRTArtificialamino acid encoded by synthesized GcM94 gene
18Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile
Asp 20 25 30Ser Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly
Gly Ala 35 40 45Ser Met Val Pro Ser Glu Thr Asn Pro Phe Gln Asp Asn
Asn Thr His 50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn
Asn Asn Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu
Tyr Ala Val Lys Val Leu 85 90 95Ala Ala Asp Gly Ala Gly Gln Tyr Ser
Trp Ile Ile Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met
Asp Val Ile Asn Met Ser Val Gly Ala 115 120 125Pro Ser Gly Ser Ala
Ala Leu Lys Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val
Val Val Val Ala Ala Ala Gly Asn Glu Gly Thr Ser Gly145 150 155
160Ser Ser Ser Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala
165 170 175Val Gly Ala Val Asp Ser Thr Asn Gln Arg Ala Ser Phe Ser
Ser Val 180 185 190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser
Ile Gln Ser Thr 195 200 205Leu Pro Gly Asn Lys Tyr Gly Ala Gln Asn
Gly Thr Ser Met Ala Ser 210 215 220Pro His Val Ala Gly Ala Ala Ala
Leu Ile Leu Ser Lys His Pro Asn225 230 235 240Trp Thr Asn Asn Gln
Val Arg Ser Ser Leu Glu Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp
Ser Phe Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala
Ala Gln 27519828DNAArtificialsynthesized GcM95 gene 19gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat
caaatgttaa agtagcggtt atcgacagcg gtatcgactc gagccatctg
120gatcttaaag tcgctggagg ggcttctatg gtgccgggag aaacaaaccc
gtttgtcgat 180gcacaaacac atggcacaca cgtcgcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgga agcttcgctc
tacgccgtta aagttcttgc agcagacaac 300gcagcacaat actcatggat
tgtcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tgagcctggg agcaccaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtgt cgtcgtagta gcggcagctg ggaatgaggg
aacatccgga 480tcatcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gggaacaagt atggtgcgca aaacgggact 660tccatggcct
cgccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgaac
720tggacaaaca ctcaagtccg cagcagttta gaaaacacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82820275PRTArtificialamino acid encoded by synthesized GcM95 gene
20Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile
Asp 20 25 30Ser Gly Ile Asp Ser Ser His Leu Asp Leu Lys Val Ala Gly
Gly Ala 35 40 45Ser Met Val Pro Gly Glu Thr Asn Pro Phe Val Asp Ala
Gln Thr His 50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn
Asn Asn Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Glu Ala Ser Leu
Tyr Ala Val Lys Val Leu 85 90 95Ala Ala Asp Asn Ala Ala Gln Tyr Ser
Trp Ile Val Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met
Asp Val Ile Asn Met Ser Leu Gly Ala 115 120 125Pro Ser Gly Ser Ala
Ala Leu Lys Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Val
Val Val Val Ala Ala Ala Gly Asn Glu Gly Thr Ser Gly145 150 155
160Ser Ser Ser Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala
165 170 175Val Gly Ala Val Asp Ser Ser Asn Gln Arg Ala Ser Phe Ser
Ser Val 180 185 190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser
Ile Gln Ser Thr 195 200 205Leu Pro Gly Asn Lys Tyr Gly Ala Gln Asn
Gly Thr Ser Met Ala Ser 210 215 220Pro His Val Ala Gly Ala Ala Ala
Leu Ile Leu Ser Lys His Pro Asn225 230 235 240Trp Thr Asn Thr Gln
Val Arg Ser Ser Leu Glu Asn Thr Thr Thr Lys 245 250 255Leu Gly Asp
Ser Phe Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala
Ala Gln 27521828DNAArtificialsynthesized GcM96 gene 21gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat
caaatgttaa agtagcggtt atcgacagcg gtatcgactc gagccatcca
120gatcttaaag tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc
gtttcaagat 180aacaattctc atggcacaca cgtcgcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc
tacgccgtta aagttcttgc agcagacgga 300tcaggccaat actcatggat
tatcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tgagcctggg agcaacaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtca agtcgtagta gcggcagctg ggaatgaggg
aacagatgga 480ccttcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactctacaa atacacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gcaaacaagt atggtgcgca aaacgggact 660tccatggccg
caccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgtca
720tggacaaaca accaagtccg cagcagttta gaacaaacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82822275PRTArtificialamino acid encoded by synthesized GcM96 gene
22Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly Ser Asn Val Lys Val Ala Val Ile
Asp 20 25 30Ser Gly Ile Asp Ser Ser His Pro Asp Leu Lys Val Ala Gly
Gly Ala 35 40 45Ser Met Val Pro Ser Glu Thr Asn Pro Phe Gln Asp Asn
Asn Ser His 50 55 60Gly Thr His Val Ala Gly Thr Val Ala Ala Leu Asn
Asn Asn Ile Gly65 70 75 80Val Leu Gly Val Ala Pro Ser Ala Ser Leu
Tyr Ala Val Lys Val Leu 85 90 95Ala Ala Asp Gly Ser Gly Gln Tyr Ser
Trp Ile Ile Asn Gly Ile Glu 100 105 110Trp Ala Ile Ala Asn Asn Met
Asp Val Ile Asn Met Ser Leu Gly Ala 115 120 125Thr Ser Gly Ser Ala
Ala Leu Lys Ala Ala Val Asp Lys Ala Val Ala 130 135 140Ser Gly Gln
Val Val Val Ala Ala Ala Gly Asn Glu Gly Thr Asp Gly145 150 155
160Pro Ser Ser Thr Val Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala
165 170 175Val Gly Ala Val Asp Ser Thr Asn Thr Arg Ala Ser Phe Ser
Ser Val 180 185 190Gly Pro Glu Leu Asp Val Met Ala Pro Gly Val Ser
Ile Gln Ser Thr 195 200 205Leu Pro Ala Asn Lys Tyr Gly Ala Gln Asn
Gly Thr Ser Met Ala Ala 210 215 220Pro His Val Ala Gly Ala Ala Ala
Leu Ile Leu Ser Lys His Pro Ser225 230 235 240Trp Thr Asn Asn Gln
Val Arg Ser Ser Leu Glu Gln Thr Thr Thr Lys 245 250 255Leu Gly Asp
Ser Phe Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala 260 265 270Ala
Ala Gln 27523828DNAArtificialsynthesized GcM100 gene 23gcgcagtccg
tgccttacgg cgtatcacaa attaaagccc ctgctctgca ctctcaaggc 60tacactggat
caaatgttaa agtagcggtt atcgacagcg gtatcgactc gagccatcca
120gatcttaaag tcgctggagg ggcttctatg gtgccgtccg aaacaaaccc
gtttcaagat 180aacaattctc atggcacaca cgcagcagga acggttgcgg
cgttaaacaa taatattggc 240gtgcttggtg tagccccgtc tgcttcgctc
tacgccgtta aagttcttgc agcagacgga 300tcagcacaat actcatggat
tatcaacggc atcgaatggg ccatcgcgaa taacatggat 360gtaatcaaca
tggcactggg agcaccaagc ggcagtgcgg cacttaaagc agcagttgat
420aaagctgttg catctggtgt cgtcgtagta gcggcagctg ggaatgaggg
aacatccgga 480tcatcgagta ccgtcggtta tccaggcaag tacccttcag
tgattgcagt gggcgctgta 540gactcttcaa atcaacgtgc ctctttttcc
tccgtgggac cggagctgga tgtcatggcc 600cctggcgttt ctattcaatc
gacgcttcca gcaaacaagt atggtgcgca aaacgggact 660tccatggcct
cgccgcatgt agctggggcg gccgcattga ttctttctaa gcacccgaac
720tggacaaaca ctcaagtccg cagcagttta gaaaacacca ctacaaaact
tggtgattct 780ttctactatg gaaaagggct gatcaacgta caggcggcag ctcagtaa
82824275PRTArtificialamino acid encoded by synthesized GcM100 gene
24Ala Gln Ser Val Pro Tyr Gly Val Ser Gln Ile Lys Ala Pro Ala Leu1
5 10 15His Ser Gln Gly Tyr Thr Gly
Ser Asn Val Lys Val Ala Val Ile Asp 20 25 30Ser Gly Ile Asp Ser Ser
His Pro Asp Leu Lys Val Ala Gly Gly Ala 35 40 45Ser Met Val Pro Ser
Glu Thr Asn Pro Phe Gln Asp Asn Asn Ser His 50 55 60Gly Thr His Ala
Ala Gly Thr Val Ala Ala Leu Asn Asn Asn Ile Gly65 70 75 80Val Leu
Gly Val Ala Pro Ser Ala Ser Leu Tyr Ala Val Lys Val Leu 85 90 95Ala
Ala Asp Gly Ser Ala Gln Tyr Ser Trp Ile Ile Asn Gly Ile Glu 100 105
110Trp Ala Ile Ala Asn Asn Met Asp Val Ile Asn Met Ala Leu Gly Ala
115 120 125Pro Ser Gly Ser Ala Ala Leu Lys Ala Ala Val Asp Lys Ala
Val Ala 130 135 140Ser Gly Val Val Val Val Ala Ala Ala Gly Asn Glu
Gly Thr Ser Gly145 150 155 160Ser Ser Ser Thr Val Gly Tyr Pro Gly
Lys Tyr Pro Ser Val Ile Ala 165 170 175Val Gly Ala Val Asp Ser Ser
Asn Gln Arg Ala Ser Phe Ser Ser Val 180 185 190Gly Pro Glu Leu Asp
Val Met Ala Pro Gly Val Ser Ile Gln Ser Thr 195 200 205Leu Pro Ala
Asn Lys Tyr Gly Ala Gln Asn Gly Thr Ser Met Ala Ser 210 215 220Pro
His Val Ala Gly Ala Ala Ala Leu Ile Leu Ser Lys His Pro Asn225 230
235 240Trp Thr Asn Thr Gln Val Arg Ser Ser Leu Glu Asn Thr Thr Thr
Lys 245 250 255Leu Gly Asp Ser Phe Tyr Tyr Gly Lys Gly Leu Ile Asn
Val Gln Ala 260 265 270Ala Ala Gln 2752536DNAArtificialsynthesized
primer 25cttctatggt gccgtccgaa acaaacccgt ttcaag
362634DNAArtificialsynthesized primer 26tcatggcaca cacgtcgcag
gaacggttgc ggcg 342737DNAArtificialsynthesized primer 27agcagacgga
tcaggccaat actcatggat tatcaac 372839DNAArtificialsynthesized primer
28tgagcctggg agcaccaagc ggcagtgcgg cacttaaag
392938DNAArtificialsynthesized primer 29tagactcttc aaatcaacgt
gcctcttttt cctccgtg 383034DNAArtificialsynthesized primer
30gaaacaaacc cgtttcaaga taacaattct catg
343137DNAArtificialsynthesized primer 31atactcatgg attatcaacg
gcatcgaatg ggccatc 373236DNAArtificialsynthesized primer
32tgttgcatct ggtgtcgtcg tagtagcggc agctgg
363337DNAArtificialsynthesized primer 33atcgacgctt ccagggaaca
agtatggtgc gcaaaac 373435DNAArtificialsynthesized primer
34cagcagttta gaaaacacca ctacaaaact tggtg
353538DNAArtificialsynthesized primer 35caaacccgtt tcaagataac
aattctcatg gcacacac 383639DNAArtificialsynthesized primer
36ttggtgtagc cccgtctgct tcgctctacg ccgttaaag
393736DNAArtificialsynthesized primer 37tggatgtaat caacatgagc
ctgggagcac caagcg 363839DNAArtificialsynthesized primer
38agggaacatc cggatcatcg agtaccgtcg gttatccag
393936DNAArtificialsynthesized primer 39gacttccatg gcctcgccgc
atgtagctgg ggcggc 364036DNAArtificialsynthesized primer
40gtttcaagat aacaattctc atggcacaca cgtcgc
364135DNAArtificialsynthesized primer 41gttcttgcag cagacggatc
aggccaatac tcatg 354237DNAArtificialsynthesized primer 42atgtaatcaa
catgagcctg ggagcaccaa gcggcag 374337DNAArtificialsynthesized primer
43ggaatgaggg aacatccgga tcatcgagta ccgtcgg
374438DNAArtificialsynthesized primer 44ctttctaagc acccgaactg
gacaaacact caagtccg 384536DNAArtificialsynthesized primer
45tcaagataac aattctcatg gcacacacgt cgcagg
364637DNAArtificialsynthesized primer 46tgcagcagac ggatcaggcc
aatactcatg gattatc 374735DNAArtificialsynthesized primer
47aatcaacatg agcctgggag caccaagcgg cagtg
354837DNAArtificialsynthesized primer 48cgctgtagac tcttcaaatc
aacgtgcctc tttttcc 374935DNAArtificialsynthesized primer
49gaactggaca aacactcaag tccgcagcag tttag
355034DNAArtificialsynthesized primer 50tcatggcaca cacgtcgcag
gaacggttgc ggcg 345137DNAArtificialsynthesized primer 51agcagacgga
tcaggccaat actcatggat tatcaac 375237DNAArtificialsynthesized primer
52atcgacgctt ccagggaaca agtatggtgc gcaaaac
375337DNAArtificialsynthesized primer 53atgtaatcaa catgagcctg
ggagcaccaa gcggcag 375422DNAArtificialsynthesized primer
54tacatatgag ttatgcagtt tg 225519DNAArtificialsynthesized primer
55ttatccttta ccttgtctc 195635DNAArtificialsynthesized primer
56caacatgagc ctgggatcac caagcggcag tgcgg
355735DNAArtificialsynthesized primer 57ccgcactgcc gcttggtgat
cccaggctca tgttg 355838DNAArtificialsynthesized primer 58ctatggtgcc
gggcgaaaca aacccgtttc aagatccg 385938DNAArtificialsynthesized
primer 59cggatcttga aacgggtttg tttcgcccgg caccatag
386021DNAArtificialsynthesized primer 60gcctcacatt tgtgccacct a
216122DNAArtificialsynthesized primer 61cctctcggtt atgagttagt tc
226224DNAArtificialsynthesized primer 62aaaggatcct aatcggcgct tttc
246324DNAArtificialsynthesized primer 63cttgtctcca agcttaaaat aaaa
246436DNAArtificialsynthesized primer 64cagatcttaa agtctctgga
ggggcttcta tggtgc 366540DNAArtificialsynthesized primer
65catagaagcc cctccagaga ctttaagatc tggatggctc
406636DNAArtificialsynthesized primer 66gcattgattc tttacaagca
cccgaactgg acaaac 366739DNAArtificialsynthesized primer
67cagttcgggt gcttgtaaag aatcaatgcg gccgcccca
396836DNAArtificialsynthesized primer 68gcattgattc ttggtaagca
cccgaactgg acaaac 366940DNAArtificialsynthesized primer
69ccagttcggg tgcttaccaa gaatcaatgc ggccgcccca
407040DNAArtificialsynthesized primer 70catcgaatgg gccacagcga
ataacatgga tgtaatcaac 407140DNAArtificialsynthesized primer
71catccatgtt attcgctgtg gcccattcga tgccgttgat
407240DNAArtificialsynthesized primer 72catcgaatgg gccgtagcga
ataacatgga tgtaatcaac 407340DNAArtificialsynthesized primer
73catccatgtt attcgctacg gcccattcga tgccgttgat
407436DNAArtificialsynthesized primer 74ctgtagactc tacaaatcaa
cgtgcctctt tttcct 367540DNAArtificialsynthesized primer
75aaagaggcac gttgatttgt agagtctaca gcgcccactg
407638DNAArtificialsynthesized primer 76ctgtagactc ttcataccaa
cgtgcctctt tttcctcc 387740DNAArtificialsynthesized primer
77gaaaaagagg cacgttggta tgaagagtct acagcgccca
407851DNAArtificialsynthesized primer 78tagcggttac agacagcggt
atcgacccaa gccatccaga tcttaaagtc g 517950DNAArtificialsynthesized
primer 79atggcttggg tcgataccgc tgtctgtaac cgctacttta acattgcctc
508049DNAArtificialsynthesized primer 80taaagtagcg gttacagaca
gcggtttaga ctcgagccat ccagatctt 498150DNAArtificialsynthesized
primer 81atggctcgag tctaaaccgc tgtctgtaac cgctacttta acattgcctc
508250DNAArtificialsynthesized primer 82ggttgtagac agcggtatcg
actcgtggca tccagatctt aaagtcgctg 508350DNAArtificialsynthesized
primer 83atgccacgag tcgataccgc tgtctacaac cgctacttta acattgcctc
508438DNAArtificialsynthesized primer 84ctacactgga ggcaaagtta
aagtagcggt tatcgaca 388540DNAArtificialsynthesized primer
85ataaccgcta ctttaacttt gcctccagtg tagccttgag
408637DNAArtificialsynthesized primer 86gagcctggga gcacgtagcg
gcagtgcggc acttaaa 378740DNAArtificialsynthesized primer
87gtgccgcact gccgctacgt gctcccaggc tcatgttgat
408838DNAArtificialsynthesized primer 88tgagcctggg agcaaagagc
ggcagtgcgg cacttaaa 388940DNAArtificialsynthesized primer
89gtgccgcact gccgctcttt gctcccaggc tcatgttgat
409040DNAArtificialsynthesized primer 90atcacaaatt aaagccacag
ctctgcactc tcaaggctac 409140DNAArtificialsynthesized primer
91agagtgcaga gctgtggctt taatttgtga tacgccgtaa
409240DNAArtificialsynthesized primer 92gacagcggta tcgacacaag
ccatccagat cttaaagtcg 409340DNAArtificialsynthesized primer
93taagatctgg atggcttgtg tcgataccgc tgtcgataac
409440DNAArtificialsynthesized primer 94gacagcggta tcgacccaag
ccatccagat cttaaagtcg 409540DNAArtificialsynthesized primer
95taagatctgg atggcttggg tcgataccgc tgtcgataac
409640DNAArtificialsynthesized primer 96gacagcggta tcgactggag
ccatccagat cttaaagtcg 409740DNAArtificialsynthesized primer
97taagatctgg atggctccag tcgataccgc tgtcgataac
409840DNAArtificialsynthesized primer 98acgcgcagtc cgtgttatac
ggcgtatcac aaattaaagc 409940DNAArtificialsynthesized primer
99atttgtgata cgccgtataa cacggactgc gcgtacgcat
4010037DNAArtificialsynthesized primer 100acaagtatgg tgcgaaaaac
gggacttcca tggcctc 3710138DNAArtificialsynthesized primer
101catggaagtc ccgtttttcg caccatactt gttccctg
3810237DNAArtificialsynthesized primer 102acaagtatgg tgcgggaaac
gggacttcca tggcctc 3710339DNAArtificialsynthesized primer
103ccatggaagt cccgtttccc gcaccatact tgttccctg
3910438DNAArtificialsynthesized primer 104cttacggcgt atcattaatt
aaagcccctg ctctgcac 3810540DNAArtificialsynthesized primer
105gagcaggggc tttaattaat gatacgccgt aaggcacgga
4010638DNAArtificialsynthesized primer 106cttacggcgt atcacgtatt
aaagcccctg ctctgcac 3810740DNAArtificialsynthesized primer
107gagcaggggc tttaatacgt gatacgccgt aaggcacgga
4010840DNAArtificialsynthesized primer 108tttagaaaac acctctacaa
aacttggtga ttctttctac 4010940DNAArtificialsynthesized primer
109tcaccaagtt ttgtagaggt gttttctaaa ctgctgcgga
4011040DNAArtificialsynthesized primer 110aggctacact ggagcaaatg
ttaaagtagc ggttatcgac 4011138DNAArtificialsynthesized primer
111gctactttaa catttgctcc agtgtagcct tgagagtg
3811239DNAArtificialsynthesized primer 112gagggaacat ccggaccatc
gagtaccgtc ggttatcca 3911340DNAArtificialsynthesized primer
113accgacggta ctcgatggtc cggatgttcc ctcattccca
4011440DNAArtificialsynthesized primer 114agggaacatc cggaccatta
agtaccgtcg gttatccagg 4011540DNAArtificialsynthesized primer
115accgacggta cttaatggtc cggatgttcc ctcattccca
4011639DNAArtificialsynthesized primer 116gaacatccgg atcattaagt
accgtcggtt atccaggca 3911740DNAArtificialsynthesized primer
117ataaccgacg gtacttaatg atccggatgt tccctcattc
4011839DNAArtificialsynthesized primer 118gaacatccgg atcaccaagt
accgtcggtt atccaggca 3911940DNAArtificialsynthesized primer
119ataaccgacg gtacttggtg atccggatgt tccctcattc
4012038DNAArtificialsynthesized primer 120gtttagaaaa cgcaactaca
aaacttggtg attctttc 3812140DNAArtificialsynthesized primer
121caccaagttt tgtagttgcg ttttctaaac tgctgcggac
4012240DNAArtificialsynthesized primer 122caaaacttgg tgatccattc
tactatggaa aagggctgat 4012340DNAArtificialsynthesized primer
123tttccatagt agaatggatc accaagtttt gtagtggtgt
4012440DNAArtificialsynthesized primer 124caaaacttgg tgataacttc
tactatggaa aagggctgat 4012540DNAArtificialsynthesized primer
125tttccatagt agaagttatc accaagtttt gtagtggtgt
4012640DNAArtificialsynthesized primer 126caaaacttgg tgatatcttc
tactatggaa aagggctgat 4012740DNAArtificialsynthesized primer
127tttccatagt agaagatatc accaagtttt gtagtggtgt
4012840DNAArtificialsynthesized primer 128caaaacttgg tgatggattc
tactatggaa aagggctgat 4012940DNAArtificialsynthesized primer
129tttccatagt agaatccatc accaagtttt gtagtggtgt
4013037DNAArtificialsynthesized primer 130gtgggcgctg tacactcttc
aaatcaacgt gcctctt 3713140DNAArtificialsynthesized primer
131cacgttgatt tgaagagtgt acagcgccca ctgcaatcac
4013237DNAArtificialsynthesized primer 132gtgggcgctg taggatcttc
aaatcaacgt gcctctt 3713340DNAArtificialsynthesized primer
133cacgttgatt tgaagatcct acagcgccca ctgcaatcac
4013437DNAArtificialsynthesized primer 134cctgctctgc acttccaagg
ctacactgga ggcaatg 3713540DNAArtificialsynthesized primer
135ctccagtgta gccttggaag tgcagagcag gggctttaat
4013637DNAArtificialsynthesized primer 136cctgctctgc acacacaagg
ctacactgga ggcaatg 3713740DNAArtificialsynthesized primer
137ctccagtgta gccttgtgtg tgcagagcag gggctttaat
4013837DNAArtificialsynthesized primer 138cctgctctgc acccacaagg
ctacactgga ggcaatg 3713940DNAArtificialsynthesized primer
139ctccagtgta gccttgtggg tgcagagcag gggctttaat
4014037DNAArtificialsynthesized primer 140cctgctctgc actaccaagg
ctacactgga ggcaatg 3714140DNAArtificialsynthesized primer
141ctccagtgta gccttggtag tgcagagcag gggctttaat
4014237DNAArtificialsynthesized primer 142cctgctctgc acttacaagg
ctacactgga ggcaatg 3714340DNAArtificialsynthesized primer
143ctccagtgta gccttgtaag tgcagagcag gggctttaat
4014438DNAArtificialsynthesized primer 144tgctctgcac tctttaggct
acactggagg caatgtta 3814540DNAArtificialsynthesized primer
145ttgcctccag tgtagcctaa agagtgcaga gcaggggctt
4014639DNAArtificialsynthesized primer 146atcgcgaata acatgaacgt
aatcaacatg agcctggga 3914739DNAArtificialsynthesized primer
147ctcatgttga ttacgttcat gttattcgcg atggcccat
3914837DNAArtificialsynthesized primer 148cttccaggga accgttatgg
tgcgcaaaac gggactt 3714939DNAArtificialsynthesized primer
149gttttgcgca ccataacggt tccctggaag cgtcgattg
3915040DNAArtificialsynthesized primer 150ctgcacttac aaggctctac
tggaggcaat gttaaagtag 4015149DNAArtificialsynthesized primer
151taacattgcc tccagtagag ccttgtaagt gcagagcagg ggctttaat
4915243DNAArtificialsynthesized primer 152gctctgcact tacaaggcaa
cactggaggc aatgttaaag tag 4315348DNAArtificialsynthesized primer
153aacattgcct ccagtgttgc cttgtaagtg cagagcaggg gctttaat
4815435DNAArtificialsynthesized primer 154cttacggcgt aacacaaatt
aaagcccctg ctctg 3515538DNAArtificialsynthesized primer
155aggggcttta atttgtgtta cgccgtaagg cacggact
3815640DNAArtificialsynthesized primer 156ttaaagcagc agttgatttc
gctgttgcat ctggtgtcgt 4015740DNAArtificialsynthesized primer
157agatgcaaca gcgaaatcaa ctgctgcttt aagtgccgca
4015840DNAArtificialsynthesized primer 158ttaaagcagc agttgatcgt
gctgttgcat ctggtgtcgt 4015940DNAArtificialsynthesized primer
159agatgcaaca gcacgatcaa ctgctgcttt aagtgccgca
4016040DNAArtificialsynthesized primer 160aaacccgttt caagattcta
attctcatgg cacacacgtc 4016140DNAArtificialsynthesized primer
161tgtgccatga gaattagaat cttgaaacgg gtttgtttcg
4016240DNAArtificialsynthesized primer 162aaacccgttt caagatgata
attctcatgg cacacacgtc 4016340DNAArtificialsynthesized primer
163tgtgccatga gaattatcat cttgaaacgg gtttgtttcg
4016439DNAArtificialsynthesized primer 164aagcggcagt gcgacactta
aagcagcagt tgataaagc 3916540DNAArtificialsynthesized primer
165tcaactgctg ctttaagtgt cgcactgccg cttggtgctc
4016637DNAArtificialsynthesized primer 166caagcggcag tgttgcactt
aaagcagcag ttgataa 3716740DNAArtificialsynthesized primer
167actgctgctt taagtgcaac actgccgctt ggtgctccca 40168382PRTBacillus
amyloliquefaciens 168Val Arg Ser Lys Lys Leu Trp Ile Ser Leu Leu
Phe Ala Leu Ala Leu1 5 10 15Ile Phe Thr Met Ala Phe Gly Ser Thr Ser
Ser Ala Gln Ala Ala Gly 20 25 30Lys Ser Asn Gly Glu Lys Lys Tyr Ile
Val Gly Phe Lys Gln Thr Met 35 40 45Ser Thr Met Ser Ala Ala Lys Lys
Lys Asp Val Ile Ser Glu Lys Gly 50 55 60Gly Lys Val Gln Lys Gln Phe
Lys Tyr Val Asp Ala Ala Ser Ala Thr65 70 75 80Leu Asn Glu Lys Ala
Val Lys Glu Leu Lys Lys Asp Pro Ser Val Ala 85 90 95Tyr Val Glu Glu
Asp His Val Ala His Ala Tyr Ala Gln Ser Val Pro 100 105 110Tyr Gly
Val Ser Gln Ile Lys Ala Pro Ala Leu His Ser Gln Gly Tyr 115 120
125Thr Gly Ser Asn Val Lys Val Ala Val Ile Asp Ser Gly Ile Asp Ser
130 135 140Ser His Pro Asp Leu Lys Val Ala Gly Gly Ala Ser Met Val
Pro Ser145 150 155 160Glu Thr Asn Pro Phe Gln Asp Asn Asn Ser His
Gly Thr His Val Ala 165 170 175Gly Thr Val Ala Ala Leu Asn Asn Ser
Ile Gly Val Leu Gly Val Ala 180 185 190Pro Ser Ala Ser Leu Tyr Ala
Val Lys Val Leu Gly Ala Asp Gly Ser 195 200 205Gly Gln Tyr Ser Trp
Ile Ile Asn Gly Ile Glu Trp Ala Ile Ala Asn 210 215 220Asn Met Asp
Val Ile Asn Met Ser Leu Gly Gly Pro Ser Gly Ser Ala225 230 235
240Ala Leu Lys Ala Ala Val Asp Lys Ala Val Ala Ser Gly Val Val Val
245 250 255Val Ala Ala Ala Gly Asn Glu Gly Thr Ser Gly Ser Ser Ser
Thr Val 260 265 270Gly Tyr Pro Gly Lys Tyr Pro Ser Val Ile Ala Val
Gly Ala Val Asp 275 280 285Ser Ser Asn Gln Arg Ala Ser Phe Ser Ser
Val Gly Pro Glu Leu Asp 290 295 300Val Met Ala Pro Gly Val Ser Ile
Gln Ser Thr Leu Pro Gly Asn Lys305 310 315 320Tyr Gly Ala Tyr Asn
Gly Thr Ser Met Ala Ser Pro His Val Ala Gly 325 330 335Ala Ala Ala
Leu Ile Leu Ser Lys His Pro Asn Trp Thr Asn Thr Gln 340 345 350Val
Arg Ser Ser Leu Glu Asn Thr Thr Thr Lys Leu Gly Asp Ser Phe 355 360
365Tyr Tyr Gly Lys Gly Leu Ile Asn Val Gln Ala Ala Ala Gln 370 375
38016936DNAArtificialsynthesized primer 169cggcgttaaa caataacatt
ggcgtgcttg gtgtag 3617037DNAArtificialsynthesized primer
170accaagcacg ccaatgttat tgtttaacgc cgcaacc
3717139DNAArtificialsynthesized primer 171gtatcgactc gagccatgaa
gatcttaaag tcgctggag 3917238DNAArtificialsynthesized primer
172cagcgacttt aagatcttca tggctcgagt cgataccg
3817339DNAArtificialsynthesized primer 173ttggtgtagc cccggatgct
tcgctctacg ccgttaaag 3917435DNAArtificialsynthesized primer
174cgtagagcga agcatccggg gctacaccaa gcacg
3517539DNAArtificialsynthesized primer 175gaacggttgc ggcgttagat
aattctattg gcgtgcttg 3917637DNAArtificialsynthesized primer
176agcacgccaa tagaattatc taacgccgca accgttc
3717744DNAArtificialsynthesized primer 177aacggttgcg gcgttagata
ataacattgg cgtgcttggt gtag 4417846DNAArtificialsynthesized primer
178acaccaagca cgccaatgtt attatctaac gccgcaaccg ttcctg
4617931DNAArtificialsynthesized primer 179cggcgttaaa caataatatt
ggcgtgcttg g 3118031DNAArtificialsynthesized primer 180ccaagcacgc
caatattatt gtttaacgcc g 3118128DNAArtificialsynthesized primer
181ggtgtagccc cggatgcttc gctctacg 2818228DNAArtificialsynthesized
primer 182cgtagagcga agcatccggg gctacacc
2818330DNAArtificialsynthesized primer 183gtatcgactc gagccatgaa
gatcttaaag 3018430DNAArtificialsynthesized primer 184ctttaagatc
ttcatggctc gagtcgatac 3018528DNAArtificialsynthesized primer
185cgcttccagg gaacaactat ggtgcgta 2818628DNAArtificialsynthesized
primer 186tacgcaccat agttgttccc tggaagcg
2818731DNAArtificialsynthesized primer 187cactctcaag gctacgttgg
atcaaatgtt a 3118831DNAArtificialsynthesized primer 188taacatttga
tccaacgtag ccttgagagt g 3118929DNAArtificialsynthesized primer
189tggcgtttct attgaatcga cgcttccag 2919029DNAArtificialsynthesized
primer 190ctggaagcgt cgattcaata gaaacgcca
2919139DNAArtificialsynthesized primer 191gaacggttgc ggcgttagat
aattctattg gcgtgcttg 3919237DNAArtificialsynthesized primer
192agcacgccaa tagaattatc taacgccgca accgttc
3719339DNAArtificialsynthesized primer 193ttggtgtagc cccggatgct
tcgctctacg ccgttaaag 3919435DNAArtificialsynthesized primer
194cgtagagcga agcatccggg gctacaccaa gcacg
3519535DNAArtificialsynthesized primer 195gcagcagacg gatcagcaca
atactcatgg attat 3519635DNAArtificialsynthesized primer
196ataatccatg agtattgtgc tgatccgtct gctgc
3519745DNAArtificialsynthesized primer 197caacatgagc ctgggagcac
caccgggcag tgcggcactt aaagc 4519845DNAArtificialsynthesized primer
198gctttaagtg ccgcactgcc cggtggtgct cccaggctca tgttg
4519939DNAArtificialsynthesized primer 199gttcttgcag cagacggaaa
tggccaatac tcatggatt 3920039DNAArtificialsynthesized primer
200aatccatgag tattggccat ttccgtctgc tgcaagaac
3920142DNAArtificialsynthesized primer 201ttaaagttct tgcagcagac
aattcaggcc aatactcatg ga 4220242DNAArtificialsynthesized primer
202tccatgagta ttggcctgaa ttgtctgctg caagaacttt aa
4220336DNAArtificialsynthesized primer 203ccgtttcaag atccgaattc
tcatggcaca cacgtc 3620438DNAArtificialsynthesized primer
204tgccatgaga attcggatct tgaaacgggt ttgtttcg
3820540DNAArtificialsynthesized primer 205ttcaaatcaa cgtgattctt
tttcctccgt gggaccggag 4020640DNAArtificialsynthesized primer
206acggaggaaa aagaatcacg ttgatttgaa gagtctacag
4020738DNAArtificialsynthesized primer 207caaatcaacg tgcctctgat
tcctccgtgg gaccggag 3820838DNAArtificialsynthesized primer
208ctccggtccc acggaggaat cagaggcacg ttgatttg
3820944DNAArtificialsynthesized primer 209agcggcagtg cggcacttaa
agttgcagtt gataaagctg ttgc 4421044DNAArtificialsynthesized primer
210gcaacagctt tatcaactgc aactttaagt gccgcactgc cgct
4421139DNAArtificialsynthesized primer 211tcaagataac aatacacatg
gcacacacgt cgcaggaac 3921240DNAArtificialsynthesized primer
212acgtgtgtgc catgtgtatt gttatcttga aacgggtttg
4021340DNAArtificialsynthesized primer 213agacggatca ggcaattact
catggattat caacggcatc 4021437DNAArtificialsynthesized primer
214taatccatga gtaattgcct gatccgtctg ctgcaag
3721538DNAArtificialsynthesized primer 215acccgtttca agataacgat
tctcatggca cacacgtc 3821638DNAArtificialsynthesized primer
216gacgtgtgtg ccatgagaat cgttatcttg aaacgggt
3821737DNAArtificialsynthesized primer 217caatactcat ggattatcga
tggcatcgaa tgggcca 3721837DNAArtificialsynthesized primer
218tggcccattc gatgccatcg ataatccatg agtattg
3721939DNAArtificialsynthesized primer 219ctcttcaaat caacgtgccg
atttttcctc cgtgggacc 3922039DNAArtificialsynthesized primer
220ggtcccacgg aggaaaaatc ggcacgttga tttgaagag
3922137DNAArtificialsynthesized primer 221caaacactca agtccgcaga
agtttagaaa acaccac 3722238DNAArtificialsynthesized primer
222gtgcttggtg tagccccgag agcttcgctc tacgccgt
3822338DNAArtificialsynthesized primer 223acggcgtaga gcgaagctct
cggggctaca ccaagcac 3822437DNAArtificialsynthesized primer
224caaacactca agtccgcgat agtttagaaa acaccac
3722537DNAArtificialsynthesized primer 225gtggtgtttt ctaaactatc
gcggacttga gtgtttg 3722638DNAArtificialsynthesized primer
226gtgcttggtg tagccccgtc tgcttcgctc tacgccgt
3822738DNAArtificialsynthesized primer 227acggcgtaga gcgaagcaga
cggggctaca ccaagcac 3822837DNAArtificialsynthesized primer
228caatactcat ggattatcaa cggcatcgaa tgggcca
3722937DNAArtificialsynthesized primer 229tggcccattc gatgccgttg
ataatccatg agtattg 3723037DNAArtificialsynthesized primer
230actcatggat tatcgatggc atcgaatggg ccatcgc
3723136DNAArtificialsynthesized primer 231cactcaagtc cgcagaagtt
tagaaaacac cactac 3623236DNAArtificialsynthesized primer
232cactcaagtc cgcgatagtt tagaaaacac cactac
3623338DNAArtificialsynthesized primer 233caaatcaacg tgccagattt
tcctccgtgg gaccggag 3823424DNAArtificialsynthesized primer
234aaaggggagg aaaatcgtga aaca 2423522DNAArtificialsynthesized
primer 235gttctaaatc gtgtttttct tg 2223648DNAArtificialsynthesized
primer 236gaactggaca aacactcaag tccgcgatag tttagaaaac accactac
4823736DNAArtificialsynthesized primer 237gacttgagtg tttgtccagt
tcgggtgctt agaaag 3623837DNAArtificialsynthesized primer
238caaacactca agtccgcagc agtttagaaa acaccac
3723937DNAArtificialsynthesized primer 239gtggtgtttt ctaaactgct
gcggacttga gtgtttg 3724049DNAArtificialsynthesized primer
240acgccgttaa agttcttgca gcagacgaat caggccaata ctcatggat
4924136DNAArtificialsynthesized primer 241tgctgcaaga actttaacgg
cgtagagcga agcaga 3624237DNAArtificialsynthesized primer
242acatggatgt aatctgcatg agcctgggag gaccaag
3724338DNAArtificialsynthesized primer 243tcctcccagg ctcatgcaga
ttacatccat gttattcg 3824422DNAArtificialsynthesized primer
244gttatgagtt agttcaaatt cg 2224546DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 245tacgcgcagt ccgtgnntnn cnncgtatca caaattaaag cccctg
4624644DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 246tttaatttgt gatacgnngn
nanncacgga ctgcgcgtac gcat 4424743DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 247tacggcgtat cacaanntnn annccctgct ctgcactctc aag
4324845DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 248agagtgcaga gcagggnntn
nannttgtga tacgccgtaa ggcac 4524947DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 249gctctgcact ctcaanncnn cnntggatca aatgttaaag
tagcggt 4725047DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 250tttaacattt gatccanngn ngnnttgaga gtgcagagca
ggggctt 4725148DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 251ctgcactctc aaggcnncnn tnnatcaaat gttaaagtag
cggttatc 4825244DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 252tactttaaca tttgatnnan ngnngccttg agagtgcaga gcag
4425347DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 253cactctcaag gctacnntnn
annaaatgtt aaagtagcgg ttatcga 4725444DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 254cgctacttta acatttnntn nanngtagcc ttgagagtgc agag
4425546DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 255tctcaaggct acactnnann
anntgttaaa gtagcggtta tcgaca 4625645DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 256aaccgctact ttaacanntn ntnnagtgta gccttgagag tgcag
4525745DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 257caaggctaca ctggannann
tnntaaagta gcggttatcg acagc 4525845DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 258gataaccgct actttannan ntnntccagt gtagccttga gagtg
4525945DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 259ggctacactg gatcanntnn
tnnagtagcg gttatcgaca gcggt 4526045DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 260gtcgataacc gctactnnan nanntgatcc agtgtagcct tgaga
4526144DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 261tacactggat caaatnntnn
annagcggtt atcgacagcg gtat 4426246DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 262gctgtcgata accgctnntn nannatttga tccagtgtag ccttga
4626345DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 263actggatcaa atgttnnann
annggttatc gacagcggta tcgac 4526446DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 264accgctgtcg ataaccnntn ntnnaacatt tgatccagtg tagcct
4626544DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 265ggatcaaatg ttaaannann
gnntatcgac agcggtatcg actc 4426647DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 266gataccgctg tcgatanncn ntnntttaac atttgatcca
gtgtagc 4726748DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 267tcaaatgtta aagtanngnn tnncgacagc ggtatcgact
cgagccat 4826848DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 268gtcgataccg ctgtcgnnan ncnntacttt aacatttgat
ccagtgta 4826945DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 269aatgttaaag tagcgnntnn cnncagcggt atcgactcga gccat
4527047DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 270cgagtcgata ccgctgnngn
nanncgctac tttaacattt gatccag 4727145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 271gttaaagtag cggttnncnn cnncggtatc gactcgagcc atcca
4527247DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 272gctcgagtcg ataccgnngn
ngnnaaccgc tactttaaca tttgatc 4727345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 273aaagtagcgg ttatcnncnn cnntatcgac tcgagccatc cagat
4527447DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 274atggctcgag tcgatanngn
ngnngataac cgctacttta acatttg 4727544DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 275gtagcggtta tcgacnncnn tnncgactcg agccatccag atct
4427646DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 276tggatggctc gagtcgnnan
ngnngtcgat aaccgctact ttaaca 4627746DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 277gcggttatcg acagcnntnn cnnctcgagc catccagatc ttaaag
4627845DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 278atctggatgg ctcgagnngn
nanngctgtc gataaccgct acttt 4527945DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 279gttatcgaca gcggtnncnn cnngagccat ccagatctta aagtc
4528044DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 280aagatctgga tggctcnngn
ngnnaccgct gtcgataacc gcta 4428146DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 281atcgacagcg gtatcnncnn gnnccatcca gatcttaaag tcgctg
4628247DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 282tttaagatct ggatggnncn
ngnngatacc gctgtcgata accgcta 4728345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 283gacagcggta tcgacnngnn cnntccagat cttaaagtcg ctgga
4528445DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 284gactttaaga tctgganngn
ncnngtcgat accgctgtcg ataac 4528544DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 285agcggtatcg actcgnncnn tnnagatctt aaagtcgctg gagg
4428644DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 286agcgacttta agatctnnan
ngnncgagtc gataccgctg tcga 4428744DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 287gactcgagcc atccanntnn tnnagtcgct ggaggggctt ctat
4428844DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 288agcccctcca gcgactnnan
nanntggatg gctcgagtcg atac 4428946DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 289agccatccag atcttnnann cnntggaggg gcttctatgg tgccgt
4629047DNAArtificialsynthesized primermisc_feature(16)..(16)n is a,
c, g, or tmisc_feature(26)..(26)n is a, c, g, or t 290catagaagcc
cctccnagcg actttnaaga tctggatggc tcgagtc
4729143DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 291catccagatc ttaaanncnn
tnnaggggct tctatggtgc cgt 4329246DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 292caccatagaa gcccctnnan ngnntttaag atctggatgg ctcgag
4629347DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 293ggaggggctt ctatgnngnn
gnncgaaaca aacccgtttc aagataa 4729444DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 294aaacgggttt gtttcgnncn ncnncataga agcccctcca gcga
4429547DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 295ggggcttcta tggtgnngnn
cnnaacaaac ccgtttcaag ataacaa 4729644DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 296ttgaaacggg tttgttnngn ncnncaccat agaagcccct ccag
4429747DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 297gcttctatgg tgccgnncnn
annaaacccg tttcaagata acaattc 4729844DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 298atcttgaaac gggtttnntn ngnncggcac catagaagcc cctc
4429947DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 299tctatggtgc cgtccnnann
anncccgttt caagataaca attctca 4730046DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 300gttatcttga aacgggnntn ntnnggacgg caccatagaa gcccct
4630147DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 301atggtgccgt ccgaannann
cnngtttcaa gataacaatt ctcatgg 4730244DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 302attgttatct tgaaacnngn ntnnttcgga cggcaccata gaag
4430347DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 303gtgccgtccg aaacanncnn
gnntcaagat aacaattctc atggcac 4730445DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 304agaattgtta tcttganncn ngnntgtttc ggacggcacc ataga
4530546DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 305ccgtccgaaa caaacnngnn
tnnagataac aattctcatg gcacac 4630644DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 306atgagaattg ttatctnnan ncnngtttgt ttcggacggc acca
4430745DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 307tccgaaacaa acccgnntnn
anntaacaat tctcatggca cacac 4530844DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 308gccatgagaa ttgttanntn nanncgggtt tgtttcggac ggca
4430945DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 309gaaacaaacc cgtttnnann
tnncaattct catggcacac acgtc 4531044DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 310tgtgccatga gaattgnnan ntnnaaacgg gtttgtttcg gacg
4431143DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 311acaaacccgt ttcaanntnn
cnnttctcat ggcacacacg tcg 4331246DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 312gtgtgtgcca tgagaanngn nannttgaaa cgggtttgtt tcggac
4631343DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 313aacccgtttc aagatnncnn
tnntcatggc acacacgtcg cag 4331446DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 314gacgtgtgtg ccatgannan ngnnatcttg aaacgggttt gtttcg
4631543DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 315ccgtttcaag ataacnntnn
tnntggcaca cacgtcgcag gaa 4331647DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 316tgcgacgtgt gtgccannan nanngttatc ttgaaacggg
tttgttt 4731744DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 317aattctcatg gcacanncnn cnnaggaacg gttgcggcgt taaa
4431847DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 318cgccgcaacc gttcctnngn
ngnntgtgcc atgagaattg ttatctt 4731945DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 319tctcatggca cacacnncnn annaacggtt gcggcgttaa acaat
4532046DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 320taacgccgca accgttnntn
ngnngtgtgt gccatgagaa ttgtta 4632147DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 321acacacgtcg cagganngnn tnnggcgtta aacaattcta
ttggcgt 4732244DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 322agaattgttt aacgccnnan ncnntcctgc gacgtgtgtg ccat
4432346DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 323gcaggaacgg ttgcgnngnn
anncaattct attggcgtgc ttggtg 4632444DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 324cacgccaata gaattgnntn ncnncgcaac cgttcctgcg acgt
4432544DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g,
or tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n
is a, c, g, or t 325acggttgcgg cgttanncnn tnntattggc gtgcttggtg
tagc 4432644DNAArtificialsynthesized primermisc_feature(17)..(18)n
is a, c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 326accaagcacg ccaatannan
ngnntaacgc cgcaaccgtt cctg 4432744DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 327aacaattcta ttggcnngnn tnntgtagcc ccgtctgctt cgct
4432850DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 328agcagacggg gctacannan
ncnngccaat agaattgttt aacgccgcaa 5032943DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 329tctattggcg tgcttnntnn anncccgtct gcttcgctct acg
4333047DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 330gagcgaagca gacgggnntn
nannaagcac gccaatagaa ttgttta 4733144DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 331attggcgtgc ttggtnnann cnngtctgct tcgctctacg ccgt
4433246DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 332gtagagcgaa gcagacnngn
ntnnaccaag cacgccaata gaattg 4633345DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 333tggcgtgctt ggtgtanncn ngnntgcttc gctctacgcc gttaa
4533445DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 334ggcgtagagc gaagcanncn
ngnntacacc aagcacgcca ataga 4533545DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 335gtgcttggtg tagccnngnn tnnttcgctc tacgccgtta aagtt
4533644DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 336aacggcgtag agcgaannan
ncnnggctac accaagcacg ccaa 4433745DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 337cttggtgtag ccccgnntnn tnngctctac gccgttaaag ttctt
4533848DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 338tttaacggcg tagagcnnan
nanncggggc tacaccaagc acgccaat 4833946DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 339ggtgtagccc cgtctnntnn gnnctacgcc gttaaagttc ttgcag
4634044DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 340aactttaacg gcgtagnncn
nannagacgg ggctacacca agca 4434145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 341gtagccccgt ctgctnngnn cnncgccgtt aaagttcttg cagca
4534244DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 342aagaacttta acggcgnngn
ncnnagcaga cggggctaca ccaa 4434345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 343gccccgtctg cttcgnncnn cnncgttaaa gttcttgcag cagac
4534444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 344tgcaagaact ttaacgnngn
ngnncgaagc agacggggct acac 4434545DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 345ccgtctgctt cgctcnncnn cnntaaagtt cttgcagcag acgga
4534643DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 346tgctgcaaga actttanngn
ngnngagcga agcagacggg gct 4334744DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 347tctgcttcgc tctacnncnn tnnagttctt gcagcagacg gatc
4434847DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 348gtctgctgca agaactnnan
ngnngtagag cgaagcagac ggggcta 4734943DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 349gcttcgctct acgccnntnn anntcttgca gcagacggat cag
4335044DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 350tccgtctgct gcaaganntn
nannggcgta gagcgaagca gacg 4435144DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 351tcgctctacg ccgttnnann tnntgcagca gacggatcag gcca
4435244DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 352tgatccgtct gctgcannan
ntnnaacggc gtagagcgaa gcag 4435344DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 353ctctacgccg ttaaanntnn tnnagcagac ggatcaggcc aata
4435444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 354gcctgatccg tctgctnnan
nanntttaac ggcgtagagc gaag 4435544DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 355tacgccgtta aagttnntnn annagacgga tcaggccaat actc
4435645DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 356ttggcctgat ccgtctnntn
nannaacttt aacggcgtag agcga 4535744DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 357gccgttaaag ttcttnnann anncggatca ggccaatact catg
4435845DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 358gtattggcct gatccgnntn
ntnnaagaac tttaacggcg tagag 4535946DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 359gttaaagttc ttgcannann cnnatcaggc caatactcat ggatta
4636046DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 360tgagtattgg cctgatnngn
ntnntgcaag aactttaacg gcgtag 4636145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 361aaagttcttg cagcanncnn annaggccaa tactcatgga ttatc
4536248DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 362ccatgagtat tggcctnntn
ngnntgctgc aagaacttta acggcgta 4836345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 363gttcttgcag cagacnnann annccaatac tcatggatta tcaac
4536446DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 364aatccatgag tattggnntn
ntnngtctgc tgcaagaact ttaacg 4636546DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 365cttgcagcag acggannann cnnatactca tggattatca acggca
4636645DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 366gataatccat gagtatnngn
ntnntccgtc tgctgcaaga acttt 4536745DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 367gcagcagacg gatcanncnn annctcatgg attatcaacg gcatc
4536844DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 368ttgataatcc atgagnntnn
gnntgatccg tctgctgcaa gaac 4436946DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 369gcagacggat caggcnnann cnnatggatt atcaacggca tcgaat
4637044DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 370gccgttgata atccatnngn
ntnngcctga tccgtctgct gcaa 4437145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 371gacggatcag gccaanncnn anngattatc aacggcatcg aatgg
4537244DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 372gatgccgttg ataatcnntn
ngnnttggcc tgatccgtct gctg 4437347DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 373ggatcaggcc aatacnnann gnntatcaac ggcatcgaat
gggccat 4737444DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 374ttcgatgccg ttgatanncn ntnngtattg gcctgatccg tctg
4437544DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 375tcaggccaat actcanngnn
tnncaacggc atcgaatggg ccat 4437645DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 376ccattcgatg ccgttgnnan ncnntgagta ttggcctgat ccgtc
4537748DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 377ggccaatact catggnntnn
cnncggcatc gaatgggcca tcgcgaat 4837845DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 378ggcccattcg atgccgnngn nannccatga gtattggcct gatcc
4537944DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 379caatactcat ggattnncnn
cnncatcgaa tgggccatcg cgaa 4438046DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 380gatggcccat tcgatgnngn ngnnaatcca tgagtattgg cctgat
4638144DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 381tactcatgga ttatcnncnn
cnncgaatgg gccatcgcga ataa 4438246DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 382cgcgatggcc cattcgnngn ngnngataat ccatgagtat tggcct
4638345DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 383tcatggatta tcaacnncnn
cnnatgggcc atcgcgaata acatg 4538446DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 384attcgcgatg gcccatnngn ngnngttgat aatccatgag tattgg
4638544DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 385tggattatca acggcnncnn
annggccatc gcgaataaca tgga 4438646DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 386gttattcgcg atggccnntn ngnngccgtt gataatccat gagtat
4638746DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 387attatcaacg gcatcnnann
gnncatcgcg aataacatgg atgtaa 4638846DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 388catgttattc gcgatgnncn ntnngatgcc gttgataatc catgag
4638947DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 389atcaacggca tcgaanngnn
cnncgcgaat aacatggatg taatcaa 4739046DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 390atccatgtta ttcgcgnngn ncnnttcgat gccgttgata atccat
4639147DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 391aacggcatcg aatggnncnn
cnngaataac atggatgtaa tcaacat 4739246DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 392tacatccatg ttattcnngn ngnnccattc gatgccgttg ataatc
4639347DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 393ggcatcgaat gggccnncnn
gnntaacatg gatgtaatca acatgag 4739444DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 394gattacatcc atgttanncn ngnnggccca ttcgatgccg ttga
4439547DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 395atcgaatggg ccatcnngnn
tnncatggat gtaatcaaca tgagcct 4739644DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 396gttgattaca tccatgnnan ncnngatggc ccattcgatg
ccgt
4439745DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 397gaatgggcca tcgcgnntnn
cnnggatgta atcaacatga gcctg 4539844DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 398catgttgatt acatccnngn nanncgcgat ggcccattcg atgc
4439945DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 399tgggccatcg cgaatnncnn
gnntgtaatc aacatgagcc tggga 4540044DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 400gctcatgttg attacanncn ngnnattcgc gatggcccat tcga
4440145DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 401gccatcgcga ataacnngnn
tnnaatcaac atgagcctgg gagca 4540245DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 402caggctcatg ttgattnnan ncnngttatt cgcgatggcc cattc
4540343DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 403atcgcgaata acatgnntnn
anncaacatg agcctgggag cac 4340446DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 404tcccaggctc atgttgnntn nanncatgtt attcgcgatg gcccat
4640544DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 405gcgaataaca tggatnnann
cnncatgagc ctgggagcac caag 4440650DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 406tgctcccagg ctcatgnngn ntnnatccat gttattcgcg
atggcccatt 5040746DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 407aataacatgg atgtanncnn cnngagcctg ggagcaccaa gcggca
4640846DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 408tggtgctccc aggctcnngn
ngnntacatc catgttattc gcgatg 4640943DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 409aacatggatg taatcnncnn gnncctggga gcaccaagcg gca
4341047DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 410gcttggtgct cccaggnncn
ngnngattac atccatgtta ttcgcga 4741143DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 411atggatgtaa tcaacnngnn cnngggagca ccaagcggca gtg
4341247DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 412gccgcttggt gctcccnngn
ncnngttgat tacatccatg ttattcg 4741345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 413gatgtaatca acatgnncnn gnnagcacca agcggcagtg cggca
4541448DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 414actgccgctt ggtgctnncn
ngnncatgtt gattacatcc atgttatt 4841544DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 415gtaatcaaca tgagcnngnn annaccaagc ggcagtgcgg cact
4441646DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 416cgcactgccg cttggtnntn
ncnngctcat gttgattaca tccatg 4641745DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 417atgagcctgg gagcannann cnncagtgcg gcacttaaag cagca
4541845DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 418tttaagtgcc gcactgnngn
ntnntgctcc caggctcatg ttgat 4541946DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 419ggagcaccaa gcggcnntnn gnnacttaaa gcagcagttg ataaag
4642045DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 420aactgctgct ttaagtnncn
nanngccgct tggtgctccc aggct 4542146DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 421gcaccaagcg gcagtnngnn anntaaagca gcagttgata aagctg
4642244DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 422atcaactgct gctttanntn
ncnnactgcc gcttggtgct ccca 4442346DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 423ccaagcggca gtgcgnnann tnnagcagca gttgataaag ctgttg
4642444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 424tttatcaact gctgctnnan
ntnncgcact gccgcttggt gctc 4442546DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 425agcggcagtg cggcanntnn annagcagtt gataaagctg ttgcat
4642644DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 426agctttatca actgctnntn
nanntgccgc actgccgctt ggtg 4442746DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 427ggcagtgcgg cacttnnann annagttgat aaagctgttg catctg
4642844DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 428aacagcttta tcaactnntn
ntnnaagtgc cgcactgccg cttg 4442946DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 429agtgcggcac ttaaannann anntgataaa gctgttgcat ctggtg
4643046DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 430tgcaacagct ttatcanntn
ntnntttaag tgccgcactg ccgctt 4643145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 431gcggcactta aagcannann tnntaaagct gttgcatctg gtgtc
4543244DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 432agatgcaaca gctttannan
ntnntgcttt aagtgccgca ctgc 4443344DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 433gcacttaaag cagcanntnn tnnagctgtt gcatctggtg tcgt
4443444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 434accagatgca acagctnnan
nanntgctgc tttaagtgcc gcac 4443544DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 435cttaaagcag cagttnntnn anntgttgca tctggtgtcg tcgt
4443646DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 436gacaccagat gcaacanntn
nannaactgc tgctttaagt gccgca 4643744DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 437aaagcagcag ttgatnnann tnntgcatct ggtgtcgtcg tagt
4443845DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 438gacgacacca gatgcannan
ntnnatcaac tgctgcttta agtgc 4543944DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 439gcagcagttg ataaanntnn tnnatctggt gtcgtcgtag tagc
4444047DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 440tacgacgaca ccagatnnan
nanntttatc aactgctgct ttaagtg 4744145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 441gcagttgata aagctnntnn anntggtgtc gtcgtagtag cggca
4544247DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 442tactacgacg acaccanntn
nannagcttt atcaactgct gctttaa 4744345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 443gttgataaag ctgttnnann tnntgtcgtc gtagtagcgg cagct
4544446DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 444cgctactacg acgacannan
ntnnaacagc tttatcaact gctgct 4644543DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 445gataaagctg ttgcanntnn tnncgtcgta gtagcggcag ctg
4344646DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 446tgccgctact acgacgnnan
nanntgcaac agctttatca actgct 4644744DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 447aaagctgttg catctnntnn cnncgtagta gcggcagctg ggaa
4444847DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 448agctgccgct actacgnngn
nannagatgc aacagcttta tcaactg 4744944DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 449gctgttgcat ctggtnncnn cnnagtagcg gcagctggga atga
4445046DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 450cccagctgcc gctactnngn
ngnnaccaga tgcaacagct ttatca 4645146DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 451gttgcatctg gtgtcnncnn annagcggca gctgggaatg agggaa
4645245DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 452attcccagct gccgctnntn
ngnngacacc agatgcaaca gcttt 4545344DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 453gcatctggtg tcgtcnnann annggcagct gggaatgagg gaac
4445444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 454ctcattccca gctgccnntn
ntnngacgac accagatgca acag 4445544DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 455tctggtgtcg tcgtannann gnnagctggg aatgagggaa catc
4445645DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 456tccctcattc ccagctnncn
ntnntacgac gacaccagat gcaac 4545746DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 457ggtgtcgtcg tagtanngnn anntgggaat gagggaacat ccggat
4645846DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 458tgttccctca ttcccanntn
ncnntactac gacgacacca gatgca 4645945DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 459gctgggaatg agggannann cnnatcatcg agtaccgtcg gttat
4546047DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 460gacggtactc gatgatnngn
ntnntccctc attcccagct gccgcta 4746145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 461gggaatgagg gaacanncnn annatcgagt accgtcggtt atcca
4546244DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 462accgacggta ctcgatnntn
ngnntgttcc ctcattccca gctg 4446344DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 463aatgagggaa catccnnann anngagtacc gtcggttatc cagg
4446444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 464ataaccgacg gtactcnntn
ntnnggatgt tccctcattc ccag 4446546DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 465acatccggat catcgnntnn cnncggttat ccaggcaagt accctt
4646644DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 466cttgcctgga taaccgnngn
nanncgatga tccggatgtt ccct 4446745DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 467tccggatcat cgagtnncnn cnnttatcca ggcaagtacc cttca
4546845DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 468gtacttgcct
ggataanngn
ngnnactcga tgatccggat gttcc 4546945DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 469tcgagtaccg tcggtnntnn anncaagtac ccttcagtga ttgca
4547044DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 470cactgaaggg tacttgnntn
nannaccgac ggtactcgat gatc 4447145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 471agtaccgtcg gttatnnann cnngtaccct tcagtgattg cagtg
4547245DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 472aatcactgaa gggtacnngn
ntnnataacc gacggtactc gatga 4547343DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 473accgtcggtt atccanncnn gnncccttca gtgattgcag tgg
4347445DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 474tgcaatcact gaagggnncn
ngnntggata accgacggta ctcga 4547545DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 475gtcggttatc caggcnngnn cnnttcagtg attgcagtgg gcgct
4547644DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 476cactgcaatc actgaanngn
ncnngcctgg ataaccgacg gtac 4447745DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 477ggttatccag gcaagnncnn tnnagtgatt gcagtgggcg ctgta
4547846DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 478gcccactgca atcactnnan
ngnncttgcc tggataaccg acggta 4647944DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 479tatccaggca agtacnntnn anngattgca gtgggcgctg taga
4448045DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 480agcgcccact gcaatcnntn
nanngtactt gcctggataa ccgac 4548144DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 481gtgggcgctg tagacnntnn anntcaacgt gcctcttttt cctc
4448244DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 482aaaagaggca cgttganntn
nanngtctac agcgcccact gcaa 4448344DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 483ggcgctgtag actctnnann tnnacgtgcc tctttttcct ccgt
4448444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 484ggaaaaagag gcacgtnnan
ntnnagagtc tacagcgccc actg 4448545DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 485gctgtagact cttcanntnn anntgcctct ttttcctccg tggga
4548644DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 486ggaggaaaaa gaggcanntn
nanntgaaga gtctacagcg ccca 4448743DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 487gtagactctt caaatnnann tnnctctttt tcctccgtgg gac
4348847DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 488cacggaggaa aaagagnnan
ntnnatttga agagtctaca gcgccca 4748944DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 489gactcttcaa atcaanntnn cnntttttcc tccgtgggac cgga
4449050DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 490tcccacggag gaaaaanngn
nannttgatt tgaagagtct acagcgccca 5049145DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 491gcctcttttt cctccnngnn annggagctg gatgtcatgg cccct
4549245DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 492catgacatcc agctccnntn
ncnnggagga aaaagaggca cgttg 4549345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 493ttttcctccg tggganngnn gnnggatgtc atggcccctg gcgtt
4549444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 494aggggccatg acatccnncn
ncnntcccac ggaggaaaaa gagg 4449548DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 495tcctccgtgg gaccgnngnn gnntgtcatg gcccctggcg
tttctatt 4849644DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 496gccaggggcc atgacanncn ncnncggtcc cacggaggaa aaag
4449745DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 497tccgtgggac cggagnngnn
tnncatggcc cctggcgttt ctatt 4549846DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 498aacgccaggg gccatgnnan ncnnctccgg tcccacggag gaaaaa
4649945DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 499gtgggaccgg agctgnntnn
cnnggcccct ggcgtttcta ttcaa 4550047DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 500agaaacgcca ggggccnngn nanncagctc cggtcccacg
gaggaaa 4750146DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 501ggaccggagc tggatnncnn gnnccctggc gtttctattc aatcga
4650245DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 502aatagaaacg ccagggnncn
ngnnatccag ctccggtccc acgga 4550344DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 503gtcatggccc ctggcnntnn tnntcaatcg acgcttccag ggaa
4450444DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 504tggaagcgtc gattgannan
nanngccagg ggccatgaca tcca 4450546DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 505attcaatcga cgcttnnann gnncaagtat ggtgcgcaaa acggga
4650646DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 506ttgcgcacca tacttgnncn
ntnnaagcgt cgattgaata gaaacg 4650745DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 507caatcgacgc ttccanngnn cnngtatggt gcgcaaaacg ggact
4550846DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 508gttttgcgca ccatacnngn
ncnntggaag cgtcgattga atagaa 4650948DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 509gggaacaagt atggtnngnn anncgggact tccatggcct
cgccgcat 4851045DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 510ggccatggaa gtcccgnntn ncnnaccata cttgttccct ggaag
4551145DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 511aacaagtatg gtgcgnnann
cnngacttcc atggcctcgc cgcat 4551244DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 512cgaggccatg gaagtcnngn ntnncgcacc atacttgttc cctg
4451343DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 513aagtatggtg cgcaanncnn
gnnttccatg gcctcgccgc atg 4351445DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 514cggcgaggcc atggaanncn ngnnttgcgc accatacttg ttccc
4551543DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 515tatggtgcgc aaaacnngnn
tnncatggcc tcgccgcatg tag 4351646DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 516atgcggcgag gccatgnnan ncnngttttg cgcaccatac ttgttc
4651747DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 517ccgcatgtag ctgggnngnn
cnnattgatt ctttctaagc acccgaa 4751848DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 518cttagaaaga atcaatnngn ncnncccagc tacatgcggc
gaggccat 4851946DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 519catgtagctg gggcgnncnn anngattctt tctaagcacc cgaact
4652051DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 520gtgcttagaa agaatcnntn
ngnncgcccc agctacatgc ggcgaggcca t 5152144DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 521gtagctgggg cggccnnann gnntctttct aagcacccga actg
4452243DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 522cgggtgctta gaaaganncn
ntnnggccgc cccagctaca tgc 4352346DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 523ttgattcttt ctaagnncnn gnnctggaca aacactcaag tccgca
4652445DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 524ttgagtgttt gtccagnncn
ngnncttaga aagaatcaat gcggc 4552545DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 525ctttctaagc acccgnncnn gnnaaacact caagtccgca gcagt
4552646DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 526gcggacttga gtgtttnncn
ngnncgggtg cttagaaaga atcaat 4652748DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 527tggacaaaca ctcaanncnn cnncagttta gaaaacacca
ctacaaaa 4852847DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 528ggtgttttct aaactgnngn ngnnttgagt gtttgtccag
ttcgggt 4752947DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 529ttagaaaaca ccactnnann anntggtgat tctttctact
atggaaa 4753046DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 530gtagaaagaa tcaccanntn ntnnagtggt gttttctaaa ctgctg
4653146DNAArtificialsynthesized primermisc_feature(16)..(17)n is a,
c, g, or tmisc_feature(19)..(20)n is a, c, g, or
tmisc_feature(22)..(23)n is a, c, g, or t 531accactacaa aacttnntnn
tnntttctac tatggaaaag ggctga 4653248DNAArtificialsynthesized
primermisc_feature(17)..(18)n is a, c, g, or
tmisc_feature(20)..(21)n is a, c, g, or tmisc_feature(23)..(24)n is
a, c, g, or t 532ttttccatag tagaaannan nannaagttt tgtagtggtg
ttttctaa 4853345DNAArtificialsynthesized
primermisc_feature(16)..(17)n is a, c, g, or
tmisc_feature(19)..(20)n is a, c, g, or tmisc_feature(22)..(23)n is
a, c, g, or t 533gattctttct actatnnann anngctgatc aacgtacagg cggca
4553448DNAArtificialsynthesized primermisc_feature(17)..(18)n is a,
c, g, or tmisc_feature(20)..(21)n is a, c, g, or
tmisc_feature(23)..(24)n is a, c, g, or t 534ctgtacgttg atcagcnntn
ntnnatagta gaaagaatca ccaagttt 4853542DNAArtificialsynthesized
primer 535tcaaggctac aatggagcaa atgttaaagt agcggttatc ga
4253638DNAArtificialsynthesized primer 536tttaacattt gctccattgt
agccttgaga gtgcagag 3853738DNAArtificialsynthesized primer
537ctacactgga ggaggtgtta aagtagcggt tatcgaca
3853837DNAArtificialsynthesized primer 538ctactttaac acctcctcca
gtgtagcctt gagagtg 3753940DNAArtificialsynthesized primer
539aggctacact ggaagaaatg ttaaagtagc ggttatcgac
4054034DNAArtificialsynthesized primer 540ctttaacatt tcttccagtg
tagccttgag agtg 3454142DNAArtificialsynthesized primer
541aaggctacac tgcaggaggt gttaaagtag cggttatcga ca
4254240DNAArtificialsynthesized primer 542ctactttaac acctcctgca
gtgtagcctt gagagtgcag 4054337DNAArtificialsynthesized primer
543cgtttcaaga tccctcttct catggcacac acgtcgc
3754440DNAArtificialsynthesized primer 544tgtgccatga gaagagggat
cttgaaacgg gtttgtttcg 4054538DNAArtificialsynthesized primer
545tgccgtccga accaaacccg tttcaagata acaattct
3854637DNAArtificialsynthesized primer 546tcttgaaacg ggtttggttc
ggacggcacc atagaag 3754740DNAArtificialsynthesized primer
547ccgtttcaag atcccaatca tcatggcaca cacgtcgcag
4054842DNAArtificialsynthesized primer 548tgtgtgccat gatgattggg
atcttgaaac gggtttgttt cg 4254945DNAArtificialsynthesized primer
549acaaacccgt tttcagatcc caattctcat ggcacacacg tcgca
4555043DNAArtificialsynthesized primer 550ccatgagaat tgggatctga
aaacgggttt gtttcggacg gca 4355138DNAArtificialsynthesized primer
551ggttgcggcg tcatacaatt ctattggcgt gcttggtg
3855238DNAArtificialsynthesized primer 552gccaatagaa ttgtatgacg
ccgcaaccgt tcctgcga 3855340DNAArtificialsynthesized primer
553tggtgtagcc tcgggtgttt cgctctacgc cgttaaagtt
4055439DNAArtificialsynthesized primer 554cgtagagcga aacacccgag
gctacaccaa gcacgccaa 3955542DNAArtificialsynthesized primer
555gtgtagcccc gggtgttgca ctctacgccg ttaaagttct tg
4255643DNAArtificialsynthesized primer 556acggcgtaga gtgcaacacc
cggggctaca ccaagcacgc caa 4355744DNAArtificialsynthesized primer
557tggtgtagcc ccgactcttg gactctacgc cgttaaagtt cttg
4455843DNAArtificialsynthesized primer 558acggcgtaga gtccaagagt
cggggctaca ccaagcacgc caa 4355941DNAArtificialsynthesized primer
559gcctgggagc acaaggctct agtgcggcac ttaaagcagc a
4156041DNAArtificialsynthesized primer 560agtgccgcac tagagccttg
tgctcccagg ctcatgttga t 4156137DNAArtificialsynthesized primer
561atggcccctg gctattctat tcaatcgacg cttccag
3756236DNAArtificialsynthesized primer 562tcgattgaat agaatagcca
ggggccatga catcca 3656342DNAArtificialsynthesized primer
563tcgacgcttc caaggtccgt gtatggtgcg caaaacggga ct
4256444DNAArtificialsynthesized primer 564ttgcgcacca tacacggacc
ttggaagcgt cgattgaata gaaa 4456539DNAArtificialsynthesized primer
565catggcacac actgcggagg aacggttgcg gcgttaaac
3956639DNAArtificialsynthesized primer 566gcaaccgttc ctccgcagtg
tgtgccatga gaattgtta 3956743DNAArtificialsynthesized primer
567gtcgcaggaa cgattggttc aaacaattct attggcgtgc ttg
4356845DNAArtificialsynthesized primer 568caatagaatt gtttgaacca
atcgttcctg cgacgtgtgt gccat 4556941DNAArtificialsynthesized primer
569aacggttgcg gcgcatggaa attctattgg cgtgcttggt g
4157041DNAArtificialsynthesized primer 570caatagaatt tccatgcgcc
gcaaccgttc ctgcgacgtg t 4157143DNAArtificialsynthesized primer
571aacggttgcg gcgagaggag gttctattgg cgtgcttggt gta
4357246DNAArtificialsynthesized primer 572cacgccaata gaacctcctc
tcgccgcaac cgttcctgcg acgtgt 4657343DNAArtificialsynthesized primer
573aacggttgcg gcgggaggcg gttctattgg cgtgcttggt gta
4357446DNAArtificialsynthesized primer 574cacgccaata gaaccgcctc
ccgccgcaac cgttcctgcg acgtgt 4657537DNAArtificialsynthesized primer
575tgcttcgctc tacggcgtta aagttcttgc agcagac
3757639DNAArtificialsynthesized primer 576caagaacttt aacgccgtag
agcgaagcag acggggcta 3957743DNAArtificialsynthesized primer
577ttcgctctac gcctcatgtt ctgcagcaga cggatcaggc caa
4357842DNAArtificialsynthesized primer 578atccgtctgc tgcagaacat
gaggcgtaga gcgaagcaga cg 4257941DNAArtificialsynthesized primer
579aataacatgg atatatcttg catgagcctg ggagcaccaa g
4158044DNAArtificialsynthesized primer 580caggctcatg caagatatat
ccatgttatt cgcgatggcc catt 4458143DNAArtificialsynthesized primer
581cgaataacat ggatcttatc tgcatgagcc tgggagcacc aag
4358245DNAArtificialsynthesized primer 582ccaggctcat gcagataaga
tccatgttat tcgcgatggc ccatt 4558344DNAArtificialsynthesized primer
583taacatggat gtatgctcat tgagcctggg agcaccaagc ggca
4458443DNAArtificialsynthesized primer 584tgctcccagg ctcaatgagc
atacatccat gttattcgcg atg 4358537DNAArtificialsynthesized primer
585acatggatgt aatctgcatg agcctgggag caccaag
3758639DNAArtificialsynthesized primer 586tcccaggctc atgcagatta
catccatgtt attcgcgat 3958738DNAArtificialsynthesized primer
587ggatgtaatc aacatcagcc tgggagcacc aagcggca
3858838DNAArtificialsynthesized primer 588tgctcccagg ctgatgttga
ttacatccat gttattcg 3858938DNAArtificialsynthesized primer
589ggatgtaatc aacgtaagcc tgggagcacc aagcggca
3859038DNAArtificialsynthesized primer 590tgctcccagg cttacgttga
ttacatccat gttattcg 3859141DNAArtificialsynthesized primer
591ggatgtaatc aacgtaagcg cgggagcacc aagcggcagt g
4159242DNAArtificialsynthesized primer 592ttggtgctcc cgcgcttacg
ttgattacat ccatgttatt cg 4259342DNAArtificialsynthesized primer
593aatcaacatg agcttcggag caagcggcag tgcggcactt aa
4259442DNAArtificialsynthesized primer 594cactgccgct tgctccgaag
ctcatgttga ttacatccat gt 4259540DNAArtificialsynthesized primer
595aacatgagcc tgtacgcacc aagcggcagt gcggcactta
4059641DNAArtificialsynthesized primer 596cactgccgct tggtgcgtac
aggctcatgt tgattacatc c 4159743DNAArtificialsynthesized primer
597caacatgagc ctgtcagcag atagcggcag tgcggcactt aaa
4359842DNAArtificialsynthesized primer 598gcactgccgc tatctgctga
caggctcatg ttgattacat cc 4259943DNAArtificialsynthesized primer
599caacatgagc ctgaacgcac gtagcggcag tgcggcactt aaa
4360042DNAArtificialsynthesized primer 600gcactgccgc tacgtgcgtt
caggctcatg ttgattacat cc 4260142DNAArtificialsynthesized primer
601atgagcctgg gaaattcatc tagcggcagt gcggcactta aa
4260241DNAArtificialsynthesized primer 602gcactgccgc tagatgaatt
tcccaggctc atgttgatta c 4160340DNAArtificialsynthesized primer
603catgagcctg ggatcagtta gcggcagtgc ggcacttaaa
4060438DNAArtificialsynthesized primer 604gcactgccgc taactgatcc
caggctcatg ttgattac 3860540DNAArtificialsynthesized primer
605catgagcctg ggatcagata gcggcagtgc ggcacttaaa
4060638DNAArtificialsynthesized primer 606gcactgccgc tatctgatcc
caggctcatg ttgattac 3860740DNAArtificialsynthesized primer
607catgagcctg ggatcaggta gcggcagtgc ggcacttaaa
4060838DNAArtificialsynthesized primer 608gcactgccgc tacctgatcc
caggctcatg ttgattac 3860940DNAArtificialsynthesized primer
609catgagcctg ggacactata gcggcagtgc ggcacttaaa
4061038DNAArtificialsynthesized primer 610gcactgccgc tatagtgtcc
caggctcatg ttgattac 3861136DNAArtificialsynthesized primer
611gagcctggga gcagacagcg gcagtgcggc acttaa
3661239DNAArtificialsynthesized primer 612tgccgcactg ccgctgtctg
ctcccaggct catgttgat 3961336DNAArtificialsynthesized primer
613gagcctggga gcagaaagcg gcagtgcggc acttaa
3661439DNAArtificialsynthesized primer 614tgccgcactg ccgctttctg
ctcccaggct catgttgat 3961536DNAArtificialsynthesized primer
615gagcctggga gcagtaagcg gcagtgcggc acttaa
3661639DNAArtificialsynthesized primer 616tgccgcactg ccgcttactg
ctcccaggct catgttgat 3961736DNAArtificialsynthesized primer
617gagcctggga gcaggaggca gtgcggcact taaagc
3661838DNAArtificialsynthesized primer 618agtgccgcac tgcctcctgc
tcccaggctc atgttgat 3861942DNAArtificialsynthesized primer
619agcctgggag cacacggcaa tgcggcactt aaagcagcag tt
4262042DNAArtificialsynthesized primer 620tttaagtgcc gcattgccgt
gtgctcccag gctcatgttg at 4262138DNAArtificialsynthesized primer
621aagcggcagt gcgacactta aagcagcagt tgataaag
3862238DNAArtificialsynthesized primer 622aactgctgct ttaagtgtcg
cactgccgct tggtgctc 3862341DNAArtificialsynthesized primer
623gttaaagttc ttcgtggttg tgacggatca ggccaatact c
4162441DNAArtificialsynthesized primer 624ctgatccgtc acaaccacga
agaactttaa cggcgtagag c 4162540DNAArtificialsynthesized primer
625ttaaagttct tgcaggaggc ggatcaggcc aatactcatg
4062642DNAArtificialsynthesized primer 626tattggcctg atccgcctcc
tgcaagaact ttaacggcgt ag 4262741DNAArtificialsynthesized primer
627ttaaagttct tgcaggacgt gacggatcag gccaatactc a
4162838DNAArtificialsynthesized primer 628ctgatccgtc acgtcctgca
agaactttaa cggcgtag 3862940DNAArtificialsynthesized primer
629ttaaagttct tgcagacggc ggatcaggcc aatactcatg
4063042DNAArtificialsynthesized primer 630tattggcctg atccgccgtc
tgcaagaact ttaacggcgt ag 4263145DNAArtificialsynthesized primer
631taaagttctt gcacatggag attcaggcca atactcatgg attat
4563244DNAArtificialsynthesized primer 632agtattggcc tgaatctcca
tgtgcaagaa ctttaacggc gtag 4463340DNAArtificialsynthesized primer
633aagttcttgc agcacgtaac ggatcaggcc aatactcatg
4063444DNAArtificialsynthesized primer 634tattggcctg atccgttacg
tgctgcaaga actttaacgg cgta 4463545DNAArtificialsynthesized primer
635agttcttgca gcagtaggag atggccaata ctcatggatt atcaa
4563645DNAArtificialsynthesized primer 636tgagtattgg ccatctccta
ctgctgcaag aactttaacg gcgta 4563743DNAArtificialsynthesized primer
637ttaaagttct tgcagcatgt agcggatcag gccaatactc atg
4363844DNAArtificialsynthesized primer 638tattggcctg atccgctaca
tgctgcaaga actttaacgg cgta 4463943DNAArtificialsynthesized primer
639aagttcttgc agcagactct tcaggccaat actcatggat tat
4364041DNAArtificialsynthesized primer 640atgagtattg gcctgaagag
tctgctgcaa gaactttaac g 4164142DNAArtificialsynthesized primer
641ttcttgcagc agactctgta ggccaatact catggattat ca
4264242DNAArtificialsynthesized primer 642catgagtatt ggcctacaga
gtctgctgca agaactttaa cg 4264343DNAArtificialsynthesized primer
643aagttcttgc agcagacgat tcaggccaat actcatggat tat
4364441DNAArtificialsynthesized primer 644atgagtattg gcctgaatcg
tctgctgcaa gaactttaac g 4164543DNAArtificialsynthesized primer
645aagttcttgc agcagacaat tcaggccaat actcatggat tat
4364641DNAArtificialsynthesized primer 646atgagtattg gcctgaattg
tctgctgcaa gaactttaac g 4164742DNAArtificialsynthesized primer
647ttcttgcagc agacaatcta ggccaatact catggattat ca
4264842DNAArtificialsynthesized primer 648catgagtatt ggcctagatt
gtctgctgca agaactttaa cg 4264943DNAArtificialsynthesized primer
649ttcttgcagc agacggagga ggccaatact catggattat caa
4365038DNAArtificialsynthesized primer 650atgagtattg gcctcctccg
tctgctgcaa gaacttta 3865143DNAArtificialsynthesized primer
651ttcttgcagc agacggagat ggccaatact catggattat caa
4365238DNAArtificialsynthesized primer 652atgagtattg gccatctccg
tctgctgcaa gaacttta 3865345DNAArtificialsynthesized primer
653tgcagcagac ggagtaggca actactcatg gattatcaac ggcat
4565445DNAArtificialsynthesized primer 654ataatccatg agtagttgcc
tactccgtct gctgcaagaa cttta 4565543DNAArtificialsynthesized primer
655ttcttgcagc agacggacgt ggccaatact catggattat caa
4365638DNAArtificialsynthesized primer 656atgagtattg gccacgtccg
tctgctgcaa gaacttta 3865743DNAArtificialsynthesized primer
657aatgggccat ctctggtaga atggatgtaa tcaacatgag cct
4365842DNAArtificialsynthesized primer 658gattacatcc attctaccag
agatggccca ttcgatgccg tt 4265940DNAArtificialsynthesized primer
659aatgggccat cggacgtaac atggatgtaa tcaacatgag
4066042DNAArtificialsynthesized primer 660gattacatcc atgttacgtc
cgatggccca ttcgatgccg tt 4266143DNAArtificialsynthesized primer
661aatgggccat caattctgga atggatgtaa tcaacatgag cct
4366242DNAArtificialsynthesized primer 662gattacatcc attccagaat
tgatggccca ttcgatgccg tt 4266336DNAArtificialsynthesized primer
663aaaacgggac ttcccaggcc tcgccgcatg tagctg
3666438DNAArtificialsynthesized primer 664tacatgcggc gaggcctggg
aagtcccgtt ttgcgcac 3866540DNAArtificialsynthesized primer
665aaggctacac tggaagaaat gttaaagtag cggttatcga
4066637DNAArtificialsynthesized primer 666ctactttaac atttcttcca
gtgtagcctt gagagtg 3766738DNAArtificialsynthesized primer
667ctacactgga tcatatgtta aagtagcggt tatcgaca
3866840DNAArtificialsynthesized primer 668taaccgctac tttaacatat
gatccagtgt agccttgaga 4066936DNAArtificialsynthesized primer
669cttctatggt ggattccgaa acaaacccgt ttcaag
3667036DNAArtificialsynthesized primer 670ggtttgtttc ggaatccacc
atagaagccc ctccag 3667139DNAArtificialsynthesized primer
671tttcaagata acaatacaca tggcacacac gtcgcagga
3967238DNAArtificialsynthesized primer 672tgtgtgccat gtgtattgtt
atcttgaaac gggtttgt 3867334DNAArtificialsynthesized primer
673gtttcaagat gaaaattctc atggcacaca cgtc
3467440DNAArtificialsynthesized primer 674tgtgccatga gaattttcat
cttgaaacgg gtttgtttcg 4067539DNAArtificialsynthesized primer
675aacccgtttc aagatccaaa ttctcatggc acacacgtc
3967638DNAArtificialsynthesized primer 676tgccatgaga atttggatct
tgaaacgggt ttgtttcg 3867741DNAArtificialsynthesized primer
677gtttcaagat aaccaatctc atggcacaca cgtcgcagga a
4167840DNAArtificialsynthesized primer 678tgtgtgccat gagattggtt
atcttgaaac gggtttgttt 4067941DNAArtificialsynthesized primer
679gtttcaagat aacgattctc atggcacaca cgtcgcagga a
4168040DNAArtificialsynthesized primer 680tgtgtgccat gagaatcgtt
atcttgaaac gggtttgttt 4068136DNAArtificialsynthesized primer
681tcaagataac aatcaacatg gcacacacgt cgcagg
3668240DNAArtificialsynthesized primer 682acgtgtgtgc catgttgatt
gttatcttga aacgggtttg 4068338DNAArtificialsynthesized primer
683tcatggcaca cacgcagcag gaacggttgc ggcgttaa
3868438DNAArtificialsynthesized primer 684caaccgttcc tgctgcgtgt
gtgccatgag aattgtta 3868533DNAArtificialsynthesized primer
685tgtagccccg gatgcttcgc tctacgccgt taa
3368635DNAArtificialsynthesized primer 686cgtagagcga agcatccggg
gctacaccaa gcacg 3568736DNAArtificialsynthesized primer
687cgttaaagtt acagcagcag acggatcagg ccaata
3668840DNAArtificialsynthesized primer 688tgatccgtct gctgctgtaa
ctttaacggc gtagagcgaa 4068936DNAArtificialsynthesized primer
689taatcaacat gagcgcggga gcaccaagcg gcagtg
3669035DNAArtificialsynthesized primer 690ttggtgctcc cgcgctcatg
ttgattacat ccatg 3569136DNAArtificialsynthesized primer
691taatcaacat gagcacggga gcaccaagcg gcagtg
3669235DNAArtificialsynthesized primer 692ttggtgctcc cgtgctcatg
ttgattacat ccatg 3569338DNAArtificialsynthesized primer
693atgtaatcaa catggcactg ggagcaccaa gcggcagt
3869440DNAArtificialsynthesized primer 694ttggtgctcc cagtgccatg
ttgattacat ccatgttatt 4069534DNAArtificialsynthesized primer
695tgggagcacc accaggcagt gcggcactta aagc
3469636DNAArtificialsynthesized primer 696gtgccgcact gcctggtggt
gctcccaggc tcatgt 3669737DNAArtificialsynthesized primer
697tgagcctggg agcacttagc ggcagtgcgg cacttaa
3769839DNAArtificialsynthesized primer 698tgccgcactg ccgctaagtg
ctcccaggct catgttgat 3969937DNAArtificialsynthesized primer
699tgagcctggg agcagaaagc ggcagtgcgg cacttaa
3770039DNAArtificialsynthesized primer 700tgccgcactg ccgctttctg
ctcccaggct catgttgat 3970137DNAArtificialsynthesized primer
701tgagcctggg agcatctagc ggcagtgcgg cacttaa
3770239DNAArtificialsynthesized primer 702tgccgcactg ccgctagatg
ctcccaggct catgttgat 3970335DNAArtificialsynthesized primer
703gactcgagcc atgaagatct taaagtcgct ggagg
3570436DNAArtificialsynthesized primer 704gactttaaga tcttcatggc
tcgagtcgat accgct 3670540DNAArtificialsynthesized primer
705gcagtccgtg cctcaaggcg tatcacaaat taaagcccct
4070640DNAArtificialsynthesized primer 706atttgtgata cgccttgagg
cacggactgc gcgtacgcat 4070740DNAArtificialsynthesized primer
707cagacggatc agcacaatac tcatggatta tcaacggcat
4070840DNAArtificialsynthesized primer 708taatccatga gtattgtgct
gatccgtctg ctgcaagaac 4070938DNAArtificialsynthesized primer
709gcagcagacg gaaacggcca atactcatgg attatcaa
3871039DNAArtificialsynthesized primer 710catgagtatt ggccgtttcc
gtctgctgca agaacttta 3971140DNAArtificialsynthesized primer
711ttcttgcagc agacgaatca ggccaatact catggattat
4071240DNAArtificialsynthesized primer 712tgagtattgg cctgattcgt
ctgctgcaag aactttaacg 4071338DNAArtificialsynthesized primer
713atcgaatggg ccgtagcgaa taacatggat gtaatcaa
3871440DNAArtificialsynthesized primer 714catccatgtt attcgctacg
gcccattcga tgccgttgat 4071538DNAArtificialsynthesized primer
715gttgataaag ctgttaaatc tggtgtcgtc gtagtagc
3871638DNAArtificialsynthesized primer 716gacgacacca gatttaacag
ctttatcaac tgctgctt 3871739DNAArtificialsynthesized primer
717gataaagctg ttgcagatgg tgtcgtcgta gtagcggca
3971840DNAArtificialsynthesized primer 718tactacgacg acaccatctg
caacagcttt atcaactgct 4071938DNAArtificialsynthesized primer
719aatgagggaa caaaaggatc atcgagtacc gtcggtta
3872040DNAArtificialsynthesized primer 720acggtactcg atgatccttt
tgttccctca ttcccagctg 4072139DNAArtificialsynthesized primer
721aacatccgga tcaaaaagta ccgtcggtta tccaggcaa
3972239DNAArtificialsynthesized primer 722ataaccgacg gtactttttg
atccggatgt tccctcatt 3972340DNAArtificialsynthesized primer
723tgttgcatct ggtccagtcg tagtagcggc agctgggaat
4072440DNAArtificialsynthesized primer 724tgccgctact acgactggac
cagatgcaac agctttatca 4072538DNAArtificialsynthesized primer
725agggaacatc cggaccatcg agtaccgtcg gttatcca
3872640DNAArtificialsynthesized primer 726accgacggta ctcgatggtc
cggatgttcc ctcattccca 4072740DNAArtificialsynthesized primer
727cttcaaatca acgtgactct ttttcctccg tgggaccgga
4072840DNAArtificialsynthesized primer 728acggaggaaa aagagtcacg
ttgatttgaa gagtctacag 4072940DNAArtificialsynthesized primer
729tcaacgtgcc tctgattcct ccgtgggacc ggagctggat
4073037DNAArtificialsynthesized primer 730tcccacggag gaatcagagg
cacgttgatt tgaagag 3773138DNAArtificialsynthesized primer
731tactatggaa aaggggtaat caacgtacag gcggcagc
3873238DNAArtificialsynthesized primer 732ctgtacgttg attacccctt
ttccatagta gaaagaat 3873336DNAArtificialsynthesized primer
733tggcgtttct attgaatcga cgcttccagg gaacaa
3673437DNAArtificialsynthesized primer 734ctggaagcgt cgattcaata
gaaacgccag gggccat 3773536DNAArtificialsynthesized primer
735cttccaggga acacatatgg tgcgcaaaac gggact
3673638DNAArtificialsynthesized primer 736gttttgcgca ccatatgtgt
tccctggaag cgtcgatt 3873736DNAArtificialsynthesized primer
737cttccaggga acctttatgg tgcgcaaaac gggact
3673838DNAArtificialsynthesized primer 738gttttgcgca ccataaaggt
tccctggaag cgtcgatt 3873940DNAArtificialsynthesized primer
739tttctactat ggaaacgggc tgatcaacgt acaggcggca
4074040DNAArtificialsynthesized primer 740acgttgatca gcccgtttcc
atagtagaaa gaatcaccaa 4074139DNAArtificialsynthesized primer
741tttctaagca cccgaaatgg acaaacactc aagtccgca
3974238DNAArtificialsynthesized primer 742gagtgtttgt ccatttcggg
tgcttagaaa gaatcaat 3874339DNAArtificialsynthesized primer
743ttctttctaa gcaccgtaac tggacaaaca ctcaagtcc
3974438DNAArtificialsynthesized primer 744tgtttgtcca gttacggtgc
ttagaaagaa tcaatgcg 3874536DNAArtificialsynthesized primer
745cacccgaact ggcgtaacac tcaagtccgc agcagt
3674640DNAArtificialsynthesized primer 746tgcggacttg agtgttacgc
cagttcgggt gcttagaaag 4074733DNAArtificialsynthesized primer
747cgtctgctta cctctacgcc gttaaagttc ttg
3374840DNAArtificialsynthesized primer 748actttaacgg cgtagaggta
agcagacggg gctacaccaa 4074936DNAArtificialsynthesized primer
749agcctgggag cacaaagcgg cagtgcggca cttaaa
3675034DNAArtificialsynthesized primer 750cactgccgct ttgtgctccc
aggctcatgt tgat 3475141DNAArtificialsynthesized primer
751ttcaatcgac gcttccaacg aacaagtatg gtgcgcaaaa c
4175239DNAArtificialsynthesized primer 752caccatactt gttcgttgga
agcgtcgatt gaatagaaa 3975339DNAArtificialsynthesized primer
753tggattatca acggcgtaga atgggccatc gcgaataac
3975439DNAArtificialsynthesized primer 754cgatggccca ttctacgccg
ttgataatcc atgagtatt 39755269PRTBacillus lentus 755Ala Gln Ser Val
Pro Trp Gly Ile Ser Arg Val Gln Ala Pro Ala Ala1 5 10 15His Asn Arg
Gly Leu Thr Gly Ser Gly Val Lys Val Ala Val Leu Asp 20 25 30Thr Gly
Ile Ser Thr His Pro Asp Leu Asn Ile Arg Gly Gly Ala Ser 35 40 45Phe
Val Pro Gly Glu Pro Ser Thr Gln Asp Gly Asn Gly His Gly Thr 50 55
60His Val Ala Gly Thr Ile Ala Ala Leu Asn Asn Ser Ile Gly Val Leu65
70 75 80Gly Val Ala Pro Ser Ala Glu Leu Tyr Ala Val Lys Val Leu Gly
Ala 85 90 95Ser Gly Ser Gly Ser Val Ser Ser Ile Ala Gln Gly Leu Glu
Trp Ala 100 105 110Gly Asn Asn Gly Met His Val Ala Asn Leu Ser Leu
Gly Ser Pro Ser 115 120 125Pro Ser Ala Thr Leu Glu Gln Ala Val Asn
Ser Ala Thr Ser Arg Gly 130 135 140Val Leu Val Val Ala Ala Ser Gly
Asn Ser Gly Ala Gly Ser Ile Ser145 150 155 160Tyr Pro Ala Arg Tyr
Ala Asn Ala Met Ala Val Gly Ala Thr Asp Gln 165 170 175Asn Asn Asn
Arg Ala Ser Phe Ser Gln Tyr Gly Ala Gly Leu Asp Ile 180 185 190Val
Ala Pro Gly Val Asn Val Gln Ser Thr Tyr Pro Gly Ser Thr Tyr 195 200
205Ala Ser Leu Asn Gly Thr Ser Met Ala Thr Pro His Val Ala Gly Ala
210 215 220Ala Ala Leu Val Lys Gln Lys Asn Pro Ser Trp Ser Asn Val
Gln Ile225 230 235 240Arg Asn His Leu Lys Asn Thr Ala Thr Ser Leu
Gly Ser Thr Asn Leu 245 250 255Tyr Gly Ser Gly Leu Val Asn Ala Glu
Ala Ala Thr Arg 260 2657561143DNABacillus lentus 756gtgagaagca
aaaaattgtg gatcgtcgcg tcgaccgcac tactcatttc tgttgctttc 60agttcatcga
tcgcatcggc tgctgaagaa gcaaaagaaa aatatttaat tggctttaat
120gagcaggaag ctgtcagtga gtttgtagaa caagtagagg caaatgacga
ggtcgccatt 180ctctctgagg aagaggaagt cgaaattgaa ttgcttcatg
aatttgaaac gattcctgtt 240ttatccgttg agttaagccc agaagatgtg
gacgcgcttg agctcgatcc agcgatttct 300tatattgaag aggatgcaga
agtaacgaca atggcgcaat cagtgccatg gggaattagc 360cgtgtgcaag
ccccagctgc ccataaccgt ggattgacag gttctggtgt aaaagttgct
420gtcctcgata caggtatttc cactcatcca gacttaaata ttcgtggtgg
cgctagcttt 480gtaccagggg aaccatccac tcaagatggg aatgggcatg
gcacgcatgt ggccgggacg 540attgctgctt taaacaattc gattggcgtt
cttggcgtag cgccgagcgc ggaactatac 600gctgttaaag tattaggggc
gagcggttca ggttcggtca gctcgattgc ccaaggattg 660gaatgggcag
ggaacaatgg catgcacgtt gctaatttga gtttaggaag cccttcgcca
720agtgccacac ttgagcaagc tgttaatagc gcgacttcta gaggcgttct
tgttgtagcg 780gcatctggaa attcaggtgc aggctcaatc agctatccgg
cccgttatgc gaacgcaatg 840gcagtcggag ctactgacca aaacaacaac
cgcgccagct tttcacagta tggcgcaggg 900cttgacattg tcgcaccagg
tgtaaacgtg cagagcacat acccaggttc aacgtatgcc 960agcttaaacg
gtacatcgat ggctactcct catgttgcag gtgcagcagc ccttgttaaa
1020caaaagaacc catcttggtc caatgtacaa atccgcaatc atctaaagaa
tacggcaacg 1080agcttaggaa gcacgaactt gtatggaagc ggacttgtca
atgcagaagc tgcaactcgt 1140taa 1143757380PRTBacillus lentus 757Val
Arg Ser Lys Lys Leu Trp Ile Val Ala Ser Thr Ala Leu Leu Ile1 5 10
15Ser Val Ala Phe Ser Ser Ser Ile Ala Ser Ala Ala Glu Glu Ala Lys
20 25 30Glu Lys Tyr Leu Ile Gly Phe Asn Glu Gln Glu Ala Val Ser Glu
Phe 35 40 45Val Glu Gln Val Glu Ala Asn Asp Glu Val Ala Ile Leu Ser
Glu Glu 50 55 60Glu Glu Val Glu Ile Glu Leu Leu His Glu Phe Glu Thr
Ile Pro Val65 70 75 80Leu Ser Val Glu Leu Ser Pro Glu Asp Val Asp
Ala Leu Glu Leu Asp 85 90 95Pro Ala Ile Ser Tyr Ile Glu Glu Asp Ala
Glu Val Thr Thr Met Ala 100 105 110Gln Ser Val Pro Trp Gly Ile Ser
Arg Val Gln Ala Pro Ala Ala His 115 120 125Asn Arg Gly Leu Thr Gly
Ser Gly Val Lys Val Ala Val Leu Asp Thr 130 135 140Gly Ile Ser Thr
His Pro Asp Leu Asn Ile Arg Gly Gly Ala Ser Phe145 150 155 160Val
Pro Gly Glu Pro Ser Thr Gln Asp Gly Asn Gly His Gly Thr His 165 170
175Val Ala Gly Thr Ile Ala Ala Leu Asn Asn Ser Ile Gly Val Leu Gly
180 185 190Val Ala Pro Ser Ala Glu Leu Tyr Ala Val Lys Val Leu Gly
Ala Ser 195 200 205Gly Ser Gly Ser Val Ser Ser Ile Ala Gln Gly Leu
Glu Trp Ala Gly 210 215 220Asn Asn Gly Met His Val Ala Asn Leu Ser
Leu Gly Ser Pro Ser Pro225 230 235 240Ser Ala Thr Leu Glu Gln Ala
Val Asn Ser Ala Thr Ser Arg Gly Val 245 250 255Leu Val Val Ala Ala
Ser Gly Asn Ser Gly Ala Gly Ser Ile Ser Tyr 260 265 270Pro Ala Arg
Tyr Ala Asn Ala Met Ala Val Gly Ala Thr Asp Gln Asn 275 280 285Asn
Asn Arg Ala Ser Phe Ser Gln Tyr Gly Ala Gly Leu Asp Ile Val 290 295
300Ala Pro Gly Val Asn Val Gln Ser Thr Tyr Pro Gly Ser Thr Tyr
Ala305 310 315 320Ser Leu Asn Gly Thr Ser Met Ala Thr Pro His
Val
Ala Gly Ala Ala 325 330 335Ala Leu Val Lys Gln Lys Asn Pro Ser Trp
Ser Asn Val Gln Ile Arg 340 345 350Asn His Leu Lys Asn Thr Ala Thr
Ser Leu Gly Ser Thr Asn Leu Tyr 355 360 365Gly Ser Gly Leu Val Asn
Ala Glu Ala Ala Thr Arg 370 375 38075827DNAArtificialsynthesized
primer 758cgcgcttgag ctcgatccag cgatttc
2775925DNAArtificialsynthesized primer 759gtctccaagc tttaacgagt
tgcag 2576030DNAArtificialsynthesized primer 760gcaattcaga
tcttccttca ggttatgacc 3076130DNAArtificialsynthesized primer
761gcatcgaaga tctgattgct taactgcttc
307621143DNAArtificialsynthesized Bacillus lentus variant
762gtgagaagca aaaaattgtg gatcgtcgcg tcgaccgcac tactcatttc
tgttgctttt 60agttcatcga tcgcatcggc tgctgaagaa gcaaaagaaa aatatttaat
tggctttaat 120gagcaggaag ctgtcagtga gtttgtagaa caagtagagg
caaatgacga ggtcgccatt 180ctctctgagg aagaggaagt cgaaattgaa
ttgcttcatg aatttgaaac gattcctgtt 240ttatccgttg agttaagccc
agaagatgtg gacgcgcttg aactcgatcc agcgatttct 300tatattgaag
aggatgcaga agtaacgaca atggcgcaat cagtgccatg gggaattagc
360cgtgtgcaag ccccggctgc ccataaccgt ggattgacag gttctggtgt
aaaagttgct 420gtcctcgata caggtatttc cactcatcca gacttaaata
ttcgtggtgg cgctagcttt 480gtaccagggg aaccatccac tcaagatggg
aatgggcatg gcacgcatgt ggccgggacg 540attgctgctc taaacaattc
gattggcgta cttggcgtag cgccgagcgc ggaactatac 600gctgttaaag
tattaggggc gagcggtggg ggcgccatca gctcgattgc ccaaggattg
660gaatgggcag ggaacaatgg catgcacgtt gctaatttga gtttaggaag
cccttcgcca 720agtgccacac ttgagcaagc tgttaatagc gcgacttcta
ggggcgttct tgttgtagcg 780gcatctggaa attcgggtgc agactcaatc
agctatccgg cccgttatgc gaacgcaatg 840gcagtcggag ctactgacca
aaacaacaac cgcgccagct tttcacagta tggcgcaggg 900cttgacatcg
tcgcaccagg tgtaaacgtg cagagcacat acccaggttc aacgtatgcc
960agcttaaacg gtacatcgat ggctactcct catgttgcag gtgcagcagt
ccttgttaaa 1020cataagaacc catcttggtc caatgtacga atccgcgatc
atctaaagaa aacggcaacg 1080agcttaggaa gcacgaactt gtatggaagc
ggacttgtca atgccgaagc tgcaactcgt 1140taa
1143763380PRTArtificialsynthesized Bacillus lentus variant 763Val
Arg Ser Lys Lys Leu Trp Ile Val Ala Ser Thr Ala Leu Leu Ile1 5 10
15Ser Val Ala Phe Ser Ser Ser Ile Ala Ser Ala Ala Glu Glu Ala Lys
20 25 30Glu Lys Tyr Leu Ile Gly Phe Asn Glu Gln Glu Ala Val Ser Glu
Phe 35 40 45Val Glu Gln Val Glu Ala Asn Asp Glu Val Ala Ile Leu Ser
Glu Glu 50 55 60Glu Glu Val Glu Ile Glu Leu Leu His Glu Phe Glu Thr
Ile Pro Val65 70 75 80Leu Ser Val Glu Leu Ser Pro Glu Asp Val Asp
Ala Leu Glu Leu Asp 85 90 95Pro Ala Ile Ser Tyr Ile Glu Glu Asp Ala
Glu Val Thr Thr Met Ala 100 105 110Gln Ser Val Pro Trp Gly Ile Ser
Arg Val Gln Ala Pro Ala Ala His 115 120 125Asn Arg Gly Leu Thr Gly
Ser Gly Val Lys Val Ala Val Leu Asp Thr 130 135 140Gly Ile Ser Thr
His Pro Asp Leu Asn Ile Arg Gly Gly Ala Ser Phe145 150 155 160Val
Pro Gly Glu Pro Ser Thr Gln Asp Gly Asn Gly His Gly Thr His 165 170
175Val Ala Gly Thr Ile Ala Ala Leu Asn Asn Ser Ile Gly Val Leu Gly
180 185 190Val Ala Pro Ser Ala Glu Leu Tyr Ala Val Lys Val Leu Gly
Ala Ser 195 200 205Gly Gly Gly Ala Ile Ser Ser Ile Ala Gln Gly Leu
Glu Trp Ala Gly 210 215 220Asn Asn Gly Met His Val Ala Asn Leu Ser
Leu Gly Ser Pro Ser Pro225 230 235 240Ser Ala Thr Leu Glu Gln Ala
Val Asn Ser Ala Thr Ser Arg Gly Val 245 250 255Leu Val Val Ala Ala
Ser Gly Asn Ser Gly Ala Asp Ser Ile Ser Tyr 260 265 270Pro Ala Arg
Tyr Ala Asn Ala Met Ala Val Gly Ala Thr Asp Gln Asn 275 280 285Asn
Asn Arg Ala Ser Phe Ser Gln Tyr Gly Ala Gly Leu Asp Ile Val 290 295
300Ala Pro Gly Val Asn Val Gln Ser Thr Tyr Pro Gly Ser Thr Tyr
Ala305 310 315 320Ser Leu Asn Gly Thr Ser Met Ala Thr Pro His Val
Ala Gly Ala Ala 325 330 335Val Leu Val Lys His Lys Asn Pro Ser Trp
Ser Asn Val Arg Ile Arg 340 345 350Asp His Leu Lys Lys Thr Ala Thr
Ser Leu Gly Ser Thr Asn Leu Tyr 355 360 365Gly Ser Gly Leu Val Asn
Ala Glu Ala Ala Thr Arg 370 375 3807641140DNAArtificialsynthesized
Bacillus lentus variant 764gtgagaagca aaaaattgtg gatcgtcgcg
tcgaccgcac tactcatttc tgttgctttt 60agttcatcga tcgcatcggc tgctgaagaa
gcaaaagaaa aatatttaat tggctttaat 120gagcaggaag ctgtcagtga
gtttgtagaa caagtagagg caaatgacga ggtcgccatt 180ctctctgagg
aagaggaagt cgaaattgaa ttgcttcatg aatttgaaac gattcctgtt
240ttatccgttg agttaagccc agaagatgtg gacgcgcttg aactcgatcc
agcgatttct 300tatattgaag aggatgcaga agtaacgaca atggcgcaat
cagtgccatg gggaattagc 360cgtgtgcaag ccccggctgc ccataaccgt
ggattgacag gttctggtgt aaaagttgct 420gtcctcgata caggtatttc
cactcatcca gacttaaata ttcgtggtgg cgctagcttt 480gtaccagggg
aaccatccac tcaagatggg aatgggcatg gcacgcatgt ggccgggacg
540attgctgctc taaacaattc gattggcgta cttggcgtag cgccgagcgc
ggaactatac 600gctgttaaag tattaggggc gagcggtggg ggcgccatca
gctcgattgc ccaaggattg 660gaatgggcag ggaacaatgg catgcacgtt
gctaatttga gtttaggaag cccttcgcca 720agtgccacac ttgagcaagc
tgttaatagc gcgacttcta ggggcgttct tgttgtagcg 780gcatctggaa
attcgggtgc agactcaatc agctatccgg cccgttatgc gaacgcaatg
840gcagtcggag ctactgacca aaacaacaac cgcgccagct tttcacagta
tggcgcaggg 900cttgacatcg tcgcaccagg tgtaaacgtg cagagcacat
acccaggttc aacgtatgcc 960agcttaaacg gtacatcgat ggctactcct
catgttgcag gtgcagcagt ccttgttaaa 1020caaaagaacc catcttggtc
caatgtacga atccgcgatc atctaaagaa tacggcaacg 1080agcttaggaa
gcacgaactt gtatggaagc ggacttgtca atgccgaagc tgcaactcgt
1140765380PRTArtificialsynthesized Bacillus lentus variant 765Val
Arg Ser Lys Lys Leu Trp Ile Val Ala Ser Thr Ala Leu Leu Ile1 5 10
15Ser Val Ala Phe Ser Ser Ser Ile Ala Ser Ala Ala Glu Glu Ala Lys
20 25 30Glu Lys Tyr Leu Ile Gly Phe Asn Glu Gln Glu Ala Val Ser Glu
Phe 35 40 45Val Glu Gln Val Glu Ala Asn Asp Glu Val Ala Ile Leu Ser
Glu Glu 50 55 60Glu Glu Val Glu Ile Glu Leu Leu His Glu Phe Glu Thr
Ile Pro Val65 70 75 80Leu Ser Val Glu Leu Ser Pro Glu Asp Val Asp
Ala Leu Glu Leu Asp 85 90 95Pro Ala Ile Ser Tyr Ile Glu Glu Asp Ala
Glu Val Thr Thr Met Ala 100 105 110Gln Ser Val Pro Trp Gly Ile Ser
Arg Val Gln Ala Pro Ala Ala His 115 120 125Asn Arg Gly Leu Thr Gly
Ser Gly Val Lys Val Ala Val Leu Asp Thr 130 135 140Gly Ile Ser Thr
His Pro Asp Leu Asn Ile Arg Gly Gly Ala Ser Phe145 150 155 160Val
Pro Gly Glu Pro Ser Thr Gln Asp Gly Asn Gly His Gly Thr His 165 170
175Val Ala Gly Thr Ile Ala Ala Leu Asn Asn Ser Ile Gly Val Leu Gly
180 185 190Val Ala Pro Ser Ala Glu Leu Tyr Ala Val Lys Val Leu Gly
Ala Ser 195 200 205Gly Gly Gly Ala Ile Ser Ser Ile Ala Gln Gly Leu
Glu Trp Ala Gly 210 215 220Asn Asn Gly Met His Val Ala Asn Leu Ser
Leu Gly Ser Pro Ser Pro225 230 235 240Ser Ala Thr Leu Glu Gln Ala
Val Asn Ser Ala Thr Ser Arg Gly Val 245 250 255Leu Val Val Ala Ala
Ser Gly Asn Ser Gly Ala Asp Ser Ile Ser Tyr 260 265 270Pro Ala Arg
Tyr Ala Asn Ala Met Ala Val Gly Ala Thr Asp Gln Asn 275 280 285Asn
Asn Arg Ala Ser Phe Ser Gln Tyr Gly Ala Gly Leu Asp Ile Val 290 295
300Ala Pro Gly Val Asn Val Gln Ser Thr Tyr Pro Gly Ser Thr Tyr
Ala305 310 315 320Ser Leu Asn Gly Thr Ser Met Ala Thr Pro His Val
Ala Gly Ala Ala 325 330 335Val Leu Val Lys Gln Lys Asn Pro Ser Trp
Ser Asn Val Arg Ile Arg 340 345 350Asp His Leu Lys Asn Thr Ala Thr
Ser Leu Gly Ser Thr Asn Leu Tyr 355 360 365Gly Ser Gly Leu Val Asn
Ala Glu Ala Ala Thr Arg 370 375 3807664PRTArtificialsynthetic
reagent 766Ala Ala Pro Phe1


D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

P00001
P00002
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.