Novel Plant-derived Cis-regulatory Elements For The Development Of Pathogen-responsive Chimeric Promotors
STAHL; Dietmar ; et al.
U.S. patent application number 16/459980 was filed with the patent office on 2020-01-23 for novel plant-derived cis-regulatory elements for the development of pathogen-responsive chimeric promotors. This patent application is currently assigned to KWS SAAT SE. The applicant listed for this patent is KWS SAAT SE. Invention is credited to Reinhard HEHL, Jeannette KOSCHMANN, Julia NIEMEYER, Dietmar STAHL, Fridtjof WELTMEIER.
| Application Number | 20200024613 16/459980 |
| Document ID | / |
| Family ID | 47998116 |
| Filed Date | 2020-01-23 |










View All Diagrams
| United States Patent Application | 20200024613 |
| Kind Code | A1 |
| STAHL; Dietmar ; et al. | January 23, 2020 |
NOVEL PLANT-DERIVED CIS-REGULATORY ELEMENTS FOR THE DEVELOPMENT OF PATHOGEN-RESPONSIVE CHIMERIC PROMOTORS
Abstract
The invention relates to an isolated cis-regulatory element imparting pathogen inducibility or elicitor inducibility, which comprises a nucleic acid molecule, the molecule sequence of which corresponds to one of the core sequence motifs comprising a) vaaagtm, b) aaacca, c) scaaam, d) acrcg, e) sktgkact, f) mrtsack, g) ccaccaa, h) tcgtctcttc (SEQ ID NO: 35), i) wwkgwc or a core sequence motif complementary to a) to i).
| Inventors: | STAHL; Dietmar; (Einbeck, DE) ; WELTMEIER; Fridtjof; (Einbeck, DE) ; HEHL; Reinhard; (Braunscheig, DE) ; KOSCHMANN; Jeannette; (Konigslutter, DE) ; NIEMEYER; Julia; (Aerzen, DE) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | KWS SAAT SE Einbeck DE |
||||||||||
| Family ID: | 47998116 | ||||||||||
| Appl. No.: | 16/459980 | ||||||||||
| Filed: | July 2, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14367521 | Jun 20, 2014 | 10385358 | ||
| PCT/DE2012/001223 | Dec 21, 2012 | |||
| 16459980 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 15/8279 20130101 |
| International Class: | C12N 15/82 20060101 C12N015/82 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Dec 23, 2011 | DE | 102011122267.0 |
Claims
1.-14. (canceled)
15. A plant cell comprising a native promoter which is suitable for effecting an expression of an operatively linked nucleic acid molecule in a plant cell and which comprises a minimal promoter and at least one cis-regulatory element upstream from the minimal promoter, wherein the cis-regulatory element imparts a pathogen inducibility or an elicitor inducibility and comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 138, b) SEQ ID NO: 139, c) SEQ ID NO: 140, d) SEQ ID NO: 141, e) SEQ ID NO: 142, f) SEQ ID NO: 143, g) SEQ ID NO: 144, h) SEQ ID NO: 35, i) SEQ ID NO: 145, and j) a nucleotide sequence complementary to the sequence of a)-i); wherein the distance from the minimal promoter to the first upstream cis-regulatory element is between 0 and 300 base pairs.
16. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 1, b) SEQ ID NO: 2, c) SEQ ID NO: 3, d) SEQ ID NO: 4, e) SEQ ID NO: 5, f) SEQ ID NO: 6, g) SEQ ID NO: 33, h) SEQ ID NO: 34, and i) SEQ ID NO: 41.
17. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 7, b) SEQ ID NO: 8, c) SEQ ID NO: 9, d) SEQ ID NO: 10, e) SEQ ID NO: 11, f) SEQ ID NO: 12, g) SEQ ID NO: 13, h) SEQ ID NO: 14, i) SEQ ID NO: 44, and j) SEQ ID NO: 15.
18. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 16, b) SEQ ID NO: 17, c) SEQ ID NO: 18, and d) SEQ ID NO: 19.
19. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 20, b) SEQ ID NO: 21, c) SEQ ID NO: 22, and d) SEQ ID NO: 23.
20. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 24, b) SEQ ID NO: 25, and c) SEQ ID NO: 26.
21. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 27 and b) SEQ ID NO: 28.
22. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 30, b) SEQ ID NO: 31, and c) SEQ ID NO: 32.
23. The plant cell of claim 15, wherein the cis-regulatory element comprises a nucleotide sequence selected from the group consisting of a) SEQ ID NO: 27, b) SEQ ID NO: 28, c) SEQ ID NO: 42, and d) SEQ ID NO: 43.
24. The plant cell of claim 15, wherein the at least one cis-regulatory element is heterologous to the minimal promoter.
25. The plant cell of claim 15, wherein the at least one cis-regulatory element is localized in a genetic environment that is different from that of the natural promoter of the at least one cis-regulatory element.
26. The plant cell of claim 15, wherein the at least one cis-regulatory element comprises at least 11 nucleotides.
27. The plant cell of claim 15, wherein the plant cell comprises at least two cis-regulatory elements.
28. The plant cell of claim 27, wherein the distance between the two cis-regulatory elements is 0 to 10 base pairs.
29. The plant cell of claim 15, wherein the native promoter is integrated into the genome of the plant cell.
30. A plant cell according to claim 15, wherein the plant cell is a cell of a plant selected from the group consisting of: corn, rice, wheat, rye, barley, oats, sorghum, potatoes, oilseed rape, sunflower, soybean, cotton and Beta vulgaris.
31. A plant comprising the cell of claim 30.
32. Seeds, cells, tissues or a part of a plant comprising the plant cell of claim 30.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of application Ser. No. 14/367,521, filed Jun. 20, 2014, which is a U.S. national phase application under 35 U.S.C. .sctn. 371 of International Patent Application No. PCT/DE2012/001223 filed Dec. 21, 2012, and claims benefit of priority to German Application No. 10 2011 122 267.0 filed on Dec. 23, 2011. The International Application was published on Jun. 27, 2013 as International Publication No. WO 2013/091612 under PCT Article 21(2). The entire contents of these applications are hereby incorporated by reference.
SEQUENCE LISTING
[0002] The instant application contains a Sequence Listing which has been submitted in ASCII format via EFS-Web and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Jul. 2, 2019, is named 245761.000081.sequence.list.txt, and is 31,509 bytes in size.
[0003] The present invention relates to cis-regulatory elements and chimeric promoters, which include pathogen- or elicitor-induced activity in plants, created from said cis regulatory elements. The present invention furthermore relates to a host cell, a transgenic plant cell, a transgenic plant tissue, and a transgenic plant and its seeds. The present invention moreover relates to a method for producing a transgenic plant that in particular is resistant to a pathogen.
[0004] Plant diseases caused by fungi, viruses, nematodes, and bacteria cause considerable crop losses worldwide, negatively affect the quality of crops, and necessitate an expensive use of chemical pesticides because the natural defense mechanisms of plants, with which they can defend themselves against the majority of potential pathogens or delay and limit their spread, are often insufficient. Genetic engineering approaches can be used for generating plants that are resistant to such above-mentioned pathogens. Such plants have increased resistance by generating an expression of proteins (effectors) specifically at the infection location (location of contact between pathogen and plant), which proteins trigger a strong resistance reaction of the plant or of molecules that are themselves toxic to pathogens or inhibit their growth or virulence.
[0005] Effectors that trigger a strong resistance reaction of the plant are, e.g. proteins of auto-activated resistance genes (R-genes) or avirulence genes that lead to the activation of endogenous resistance genes of the plant. The hypersensitive reaction (HR), the controlled cell death of the host tissue at the infection location, the strengthening of the plant cell wall via lignification and callose formation, the formation of phytoalexines, and the production of PR--(pathogenesis-related) proteins are counted among the stronger resistance reactions.
[0006] Since said resistance reactions present a high energy requirement and can lead to the death of plant cells (necrosis), their triggering must be subject to stringent monitoring. The same also applies for the expression of proteins or peptides toxic to pathogens inasmuch as a constitutive expression of these proteins or peptides has a disadvantageous effect on the plant or its agronomic traits such as yield, for example. In a transgenic approach, such control is possible by using promoters having the desired specificity.
[0007] A pathogen-induced expression of transgenes such as effectors can thus be achieved by using various known, natural pathogen-inducible promoters such as of the PR1-promoter, for example (Rushton et al., 1996). The development of the micro-array technology in particular has led to the identification of a multitude of pathogen-inducible promoters (WO 03/00898, WO 02/50293 or JP 2003284566).
[0008] Such natural pathogen-inducible promoters can, however, display very unspecific activities since they can be activated by a large number of different stimuli. Such activities are attributed to a modular construction of the promoter from a plurality of different cis-regulatory elements that integrate diverse-as-possible different signals into a complex expression profile. Natural pathogen-inducible promoters are accordingly also characterized by undesired activities in certain tissues, for example, or by a high degree of background activity.
[0009] Thus, for example, pathogen-inducible promoters of defensin genes from wheat are also active during the seed development and seed germination (Kovalchuk et al., 2010). The already-mentioned PR1 promoter is induced not only by pathogens, but also by senescence (Morris et al. 2000)
[0010] Other research has shown that the natural, rust-inducible Fis1 promoter from flax was not suited after transformation to specifically regulate the expression of auto-active forms of the L6 rust-resistance gene in Linum usitatissimum such that in addition to the rust resistance, no negative agronomic traits such as restricted growth arose (Howles et al., 2005).
[0011] The identification of the cis-regulatory elements responsible for the desired induction and the construction of chimeric promoters from said cis-regulatory elements is a possibility for increasing the desired specificity of a promoter (Venter, 2007). Sequence motifs for other stimuli are, on the other hand, removed.
[0012] The promoters of many pathogen-induced genes have been more closely examined, wherein a plurality of cis-regulatory elements that can impart a pathogen-specific induction were identified (Strittmatter et al., 1996; Eulgem et al., 2000; Kirsch et al., 2000, 2001; Himmelbach et al., 2010). The cis-regulatory elements D-box, S-box or W-box identified by the incremental mutation of natural pathogen-inducible promoters (WO 00/29592) or the left-hand scanning regions LS10 or LS7 (Lebel et al., 1998) are also additional examples. The W-box in particular has been well investigated and its core sequence TTGAC(C/T) can be used to find additional variants of the W-box in natural pathogen-inducible promoters.
[0013] New cis-regulatory elements can furthermore be identified bioinformatically using programs such as MEME (Bailey and Elkan, 1994; Humphry et al., 2010) or BEST (Che et al., 2005). An advantage of this is that a cis-regulatory element is identified not as a short single sequence, but rather as a precisely defined sequence motif via which even more variants of a cis-regulatory element, that is to say many variants of a binding site of transcription factor, are detected. Such bioinformatic approaches are, however, also highly susceptible to yielding false-positive sequences such that these approaches merely lead to a screening of potential sequences or sequence motifs. Proof and verification of the functionality as a cis-regulatory element in general, and specifically as a cis-regulatory element that imparts pathogen-inducibility, remain essential and obligatory. Such experimental analyses are furthermore associated with considerable expense.
[0014] An additional increase in the specificity of a promoter is possible by using combinations of different cis-regulatory elements (Rushton et al., 2002). It is not the combination itself that leads to an increase of the activity (synergism), but rather such a synergism occurs only with specific individual, non-predictable combinations and must in any case be determined empirically. Chimeric promoters having combinations from the cis-regulatory elements D-box and S-box, for example, are known (WO 00/29592). The number of the element repetitions modulates the promoter strength and the background activity.
[0015] A further problem in the development of pathogen-inducible chimeric promoters is their functionality in different plant species. Although generally pathogen inducibility via the known pathogen-inducible chimeric promoters can be determined in nearly all plant varieties analyzed thus far, they still exhibit background activity even when not under attack by a pathogen. Such background activity varies depending on the plant species in which the chimeric promoters are used. The same is true with the induction rate (ratio of the promoter activity in the infected tissue and promoter activity in uninfected tissue) and the absolute activity of the promoters (promoter strength). Thus, for example, when background activity is too strong in non-infected tissue, only a slight pathogen inducibility in the infected tissue is detected.
[0016] According to the current state of knowledge, the used cis-regulatory elements of a promoter are responsible for the described variations in background activity, induction rate, promoter strength, induction kinetics, and the scope of the promoter activation (Rushton et al., 2002, Venter, 2007). Even if the known chimeric promoters are superior to the natural promoters, there is still an optimization need for these chimeric promoters, especially in regard to the cis-regulatory elements and/or to the combinations of cis-regulatory elements. There is still a lack of well-characterized cis-regulatory elements and of suitable combinations of such cis-regulatory elements with which chimeric promoters can be constructed, which ensure a highly specific and controlled, appropriate pathogen-induced expression of transgenes themselves and also in various plant species, wherein the expression should occur only as a result of pathogen attack and almost exclusively at the site of infection (Gurr & Rushton, 2005). The object of the present invention is therefore to provide such novel cis-regulatory elements, and combinations thereof, imparted by pathogen inducibility.
[0017] Some of the terms used in this application will be first be explained below:
[0018] An "elicitor" in the sense of the present invention is an inductor or messenger that induces defenses against plant pathogens such as the synthesis of phytoalexines. Elicitors can be either endogenous or exogenous in origin. An elicitor (exogenous) preferably comes from a pathogen and is recognized by the plant. The PAMPs (pathogen associated molucular pattern) such as flagelin, PEP25, and chitin are also counted among to these elicitors. Elicitors can be used to imitate a pathogen infection or a contact with a pathogen by artificially applying the elicitor in the absence of the pathogen. In the context of the present invention, elicitors should be used in particular to monitor the inducibility of promoters.
[0019] A "single sequence" is a sequence of nucleotides or bases (pairs), wherein each position in the single sequence is determined by only one single pre-defined base (a, c, g or t). A single sequence is isolated from a natural promoter and represents the result of a bioinformatics analysis. A single sequence is composed of a core sequence and flanking sequence regions. The term "single sequence" also means a nucleic acid molecule the nucleotide- or base (pair)-sequences of which correspond to the single sequence.
[0020] A "core sequence" is the sequence of nucleotides or bases (base pairs) in a certain section of a cis-regulatory element, wherein said section is essential for the functionality of the cis-regulatory element. The core sequence represents a part of the single sequence. The term "core sequence" also means a nucleic acid molecule the nucleotide- or base-(pair)-sequences of which correspond to the core sequence.
[0021] A "promoter" means a non-translated DNA sequence, typically upstream from a coded region, which contains the binding site for the RNA polymerase and initiates the transcription of the DNA. A promoter additionally often contains other elements that act as regulators of the gene expression (e.g. cis-regulatory elements).
[0022] A "minimal promoter" is a promoter that has only the basic elements that are used for the transcription initiation (e.g. TATA-box and/or initiator).
[0023] A "chimeric promoter" is a promoter composed of a plurality of elements and that does not occur as such nature. It contains a minimal promoter and includes, upstream from the minimal promoter, at least one cis-regulatory element which serves as a binding site for specific trans-acting factors (e.g. transcription factors). A chimeric promoter is designed according to the desired requirements and is induced or repressed by different factors. The selection of the cis-regulatory element or a combination of cis-regulatory elements is critical for the specificity or the activity level of a promoter. A cis-regulatory element in a chimeric promoter is either heterologous to the minimal promoter used, i.e. the cis-regulatory element is derived from a different organism or from a different species to that of the minimal promoter used (exemplarily represented in FIG. 15A-C), or a cis-regulatory element in a chimeric promoter is homologous to the minimal promoter used, i.e. the cis-regulatory element and the minimal promoter also occur combined in a natural promoter, however the cis-regulatory element is localized itself or as an additional element within the chimeric promoter in a genetic environment that is different from the natural promoter. A chimeric promoter thus also means a (natural) promoter that was altered by multimerization of at least one cis-regulatory element (exemplarily represented in FIG. 15D).
[0024] A "complementary" nucleotide sequence in relation to a double-stranded DNA means that the second DNA strand complementary to the first DNA strand includes the nucleotide bases that correspond to the bases of the first strand in accordance with base-pairing rules and considering the orientation (e.g.: 5'-gcat-3' is complementary to 5'-atgc-3').
[0025] A "pathogen" means an organism that in interactions with a plant leads to illness symptoms in one or more organs of the plant. Animal, fungal, bacterial or viral organisms or oomycetes, for example, are among these pathogens.
[0026] "Pathogen infection" is to be understood as the earliest point in time at which the metabolism of a pathogen is prepared for a penetration of the plant host tissue. Included thereamong in, e.g., fungi or in oomycetes, is the maturation of hyphae or the formation of specific infection structures such as penetration hyphae and appressoria.
[0027] A "pathogen-/elicitor-inducibility" or "pathogen-/elicitor-inducible" in the sense of the present invention means the specific property of a promoter that causes, subsequent to pathogen infection or elicitor application, an at least two-fold stronger transcription of an operatively linked gene. "Pathogen-/elicitor-inducibility" or "pathogen-/elicitor-inducible" in the sense of the present invention furthermore means the property of genes to be transcribed at least two-fold stronger subsequent to pathogen infection or elicitor application.
[0028] The solution of the invention to the object posed is inventively effected by novel cis-regulatory elements imparting pathogen- and/or elicitor-inducibility. Said cis-regulatory elements distinguish themselves in particular within the core sequence significantly from already-known elements and thus do not represent a variation of the known elements. The inventive cis-regulatory elements should accordingly serve as recognition sites and/or binding sites for new transcription factors with the consequence that the imparted pathogen and/or elicitor inducibility has a novel specificity. The cis-regulatory elements according to the invention were identified in promoters of pathogen or PAMP (elicitor)-induced genes from Arabidopsis thaliana via bioinformatics approaches. The isolated single sequences of the cis-regulatory elements could be associated with eight motif groups (motif group 1, 5, 11, 12, 18, 21, 27 and 32) pursuant to various analysis steps. A plurality of isolated single sequences could moreover also be associated with a 21n motif group. Grouped in one motif group are those single sequences that demonstrate a high degree of conservation compared to the identified motifs. Single sequences of a motif group all correspond in a characteristic core sequence motif. The object posed is thus achieved by an isolated cis-regulatory element comprising a nucleic acid molecule, the nucleotide sequence of which corresponds to one of the core sequence motifs from
TABLE-US-00001 a) vaaagtm, b) aaacca, c) scaaam, d) acrcg, e) sktgkact, f) mrtsack, g) ccaccaa, (SEQ ID NO: 35) h) tcgtctcttc or i) wwkgwc.
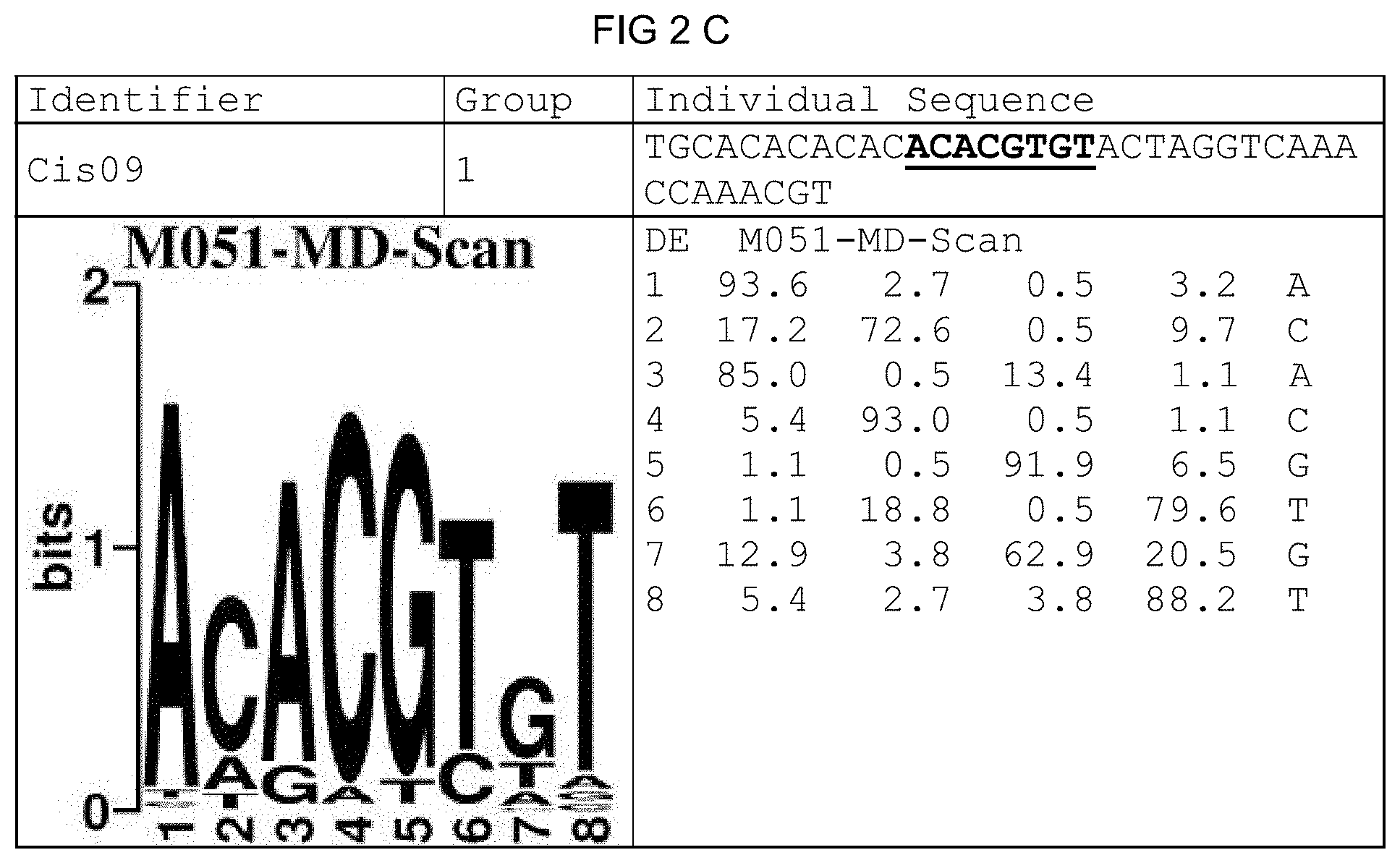
[0029] Less strongly conserved base positions within the characteristic core sequence motifs a) to i) are given as follows: `r` stands for guanine (g) or adenine (a), thus a purine base, `k` stands for guanine (g) or thymine (t)/uracil (u), `s` stands for guanine (g) or cytosine (c), `m` stands for adenine (a) or cytosine (c) and `w` stands for adenine (a) or thymine (t)/uracil (u). A certain core sequence motif reflects at least one partial sequence of the core sequence of one of each of the single sequences of the motif group belonging to the core sequence motif, wherein the partial sequence can constitute at least 30% of the entire core sequence of a single sequence. For those motif groups having core sequence motifs g) and h), the core sequence motif corresponds to the entire core sequence of the single sequences. The invention furthermore also includes an isolated cis-regulatory element comprising a nucleic acid molecule the nucleotide sequence of which corresponds to a core sequence motif complementary to a) to i). A characteristic core sequence motif of a certain motif group can moreover also repeatedly arise in the core sequence of a single sequence, wherein the core sequence motifs can also appear overlapping in the core sequence and/or respectively demonstrate a different orientation. For example, the core sequence of the Cis09 from motif group 1 demonstrates, on the one hand, a partial sequence that corresponds to the core sequence motif acrcg and, on the other hand, a partial sequence overlapping by two bases, which partial sequence corresponds to the complementary core sequence motif of acrcg, namely cgygt. A further example is found in the core sequence of the 21G-2_M1_S2 from motif group 11 where the core sequence has two partial sequences that overlap by one base and each corresponds to the complementary core sequence motif of aaacca, namely tggttt.
[0030] A family motif, in which the characteristic core sequence motif is embedded, could be defined for motif groups 1, 5, 11, 12, 21, 21n, and 27 based on the experimental data regarding functionality. The family motif represents a derived distinguishing feature for a transcription factor or a transcription factor family. The advantage of a family motif is that it subsumes possible variants of a recognition/binding area. A family motif of a motif group advantageously comprises all core sequences of the single sequences grouped in the motif group. To achieve this, the complementary strand of the cis-regulatory element was considered for a part of the single sequences; such complementary sequences are from identified cis-regulatory sequences according to the invention, and are characterized with `(inv)` in the following inasmuch as needed for understanding. A family motif defined in such a manner has a length of at least 15 nucleotides, preferably of at least 13 nucleotides, especially preferred of at least 11 nucleotides. In addition to the corresponding core sequence motif, the family motif has motif flanking sites, wherein when a cis-regulatory element according to the invention is used in a chimeric promoter said motif flanking sites have a substantially quantitative influence on the properties thereof such as background activity and expression strength. The quantitative influence of said motif flanking site reduces with increasing distance from a certain base of the flanking site to the core sequence in a single sequence. Further highly conserved single bases from the flanking sites, which highly conserved single bases go beyond the core sequences of the single sequences and thus expand the family motif, can also be considered in defining the family motif. The family motifs for the motif groups 1, 5, 11, 12, 21, 21n, and 27 are shown in column 2 of Table 1. According to the invention, an isolated cis-regulatory element is also included that comprises a nucleic acid molecule, the nucleotide sequence of which corresponds to [0031] a) a family motif according to SEQ ID NO: 1, including the core sequence motif vaaagtm [0032] b) a family motif according to SEQ ID NO: 2, including the core sequence motif aaacca, [0033] c) a family motif according to SEQ ID NO: 3, including the core sequence motif scaaam, [0034] d) a family motif according to SEQ ID NO: 4, including the core sequence motif acrcg, [0035] e) a family motif according to SEQ ID NO: 5, including the core sequence motif sktgkact, [0036] f) a family motif according to SEQ ID NO: 6, including the core sequence motif mrtsack, [0037] g) a family motif according to SEQ ID NO: 41, including the core sequence motif wwkgwc, or the nucleotide sequence of which corresponds to one of the family motifs complementary to a) to g).
[0038] It is also possible for a family motif to be shorter. In its shortest form, it is defined as a minimal family motif that based on the direction of the single sequences corresponding to the shared core sequence motif groups represents in itself only those base positions that are identified in the core sequences of all single sequences of a motif group. In some cases, the minimal family motif corresponds to the core sequence motif. Column 2, titled Family Motif, of Table 1 shows underlined the minimal family motifs of the motif groups 1, 5, 11, 12, 21, 21n, and 27.
TABLE-US-00002 TABLE 1 Representation of the motif groups 1, 5, 11, 12, 18, 21, 21n, 27, and 32, the core sequence motifs (1) and family motifs (2) underlying the groups, and the single sequences (3) associated with the motif groups; the minimal family motif is underlined, the core sequence motif is in bold typeface within family motif. (`n` stands for an arbitrary base; `h` stands for a, c or t/u; `d` stands for a, g or t/u; `v` stands for a, g or c; `r` stands for g or a, `k` stands for g or t/u, `s` stands for g or c and `m` stands for a or c; `y` stands for t/u or c; `w` stands for a or t/u) 1 3 Core sequence 2 Single sequence motif Family motif identifier Motif group 27 vaaagtm nnhkdnnvaaagtmndhy (SEQ ID NO: 1) 30I-8_M1_S1 (SEQ ID NO: 7) 30I-8_M1_S2 (SEQ ID NO: 8) GG13_M1_S2 (SEQ ID NO: 9) 14S_M1_S1 (SEQ ID NO: 10) 21S_M3_S1 (SEQ ID NO: 11) 30I-8_M1_S3 (SEQ ID NO: 12) Cis02 (SEQ ID NO: 13) Cis05 (SEQ ID NO: 14) sCis05 (SEQ ID NO: 44) Cis13 (SEQ ID NO: 15) Motif group 11 aaacca ynamcnaaaccawwny (SEQ ID NO: 2) GG11_M1_S1 (SEQ ID NO: 16) 22DDD_M1_S1 (SEQ ID NO: 17) 21G-2_M1_S2 (SEQ ID NO: 18) GG6_M1_S1 (SEQ ID NO: 19) Motif group 12 scaaam wnrmscaaamsmw (SEQ ID NO: 3) 18H_M2_S1 (SEQ ID NO: 20) 18H_M2_S3 (SEQ ID NO: 21) 38M_M1_S1 (SEQ ID NO: 22) 26LLL_M1_S2 (SEQ ID NO: 23) Motif group 1 acrcg nnmsacrcgynwm (SEQ ID NO: 4) Cis09 (SEQ ID NO: 24) Cis12 (SEQ ID NO: 25) 12i_M1_S1 (SEQ ID NO: 26) Motif group 21 sktgkact asktgkactwkgwm (SEQ ID NO: 5) GG8_M1_S1 (SEQ ID NO: 27) 27G-8_M1_S1 (SEQ ID NO: 28) Motif group 21n wwkgwc snsnnnwwkgwcnnnsnm (SEQ ID NO: 41) GG8_M1_S1 (SEQ ID NO: 27) 27G-8_M1_S1 (SEQ ID NO: 28) 26WW_M2_S1 (SEQ ID NO: 42) 27B-10_M1_S3 (SEQ ID NO: 43) Motif group 5 mrtsack knwymmrtsackwmn (SEQ ID NO: 6) 20u_M1_S1 (SEQ ID NO: 30) 20u_M1_S2 (SEQ ID NO: 31) 28M-1_M1_S1 (SEQ ID NO: 32) Motif group 18 ccaccaa 12G_M2_S1 (SEQ ID NO: 33) Motif group 32 tcgtctcttc 12r_M1_S1 (SEQ ID NO: 34) (SEQ ID NO: 35)
[0039] The invention furthermore also relates to all identified and isolated single sequences of cis-regulatory elements according to the invention and the core sequences thereof (Tables 1 and 2). The object posed is thus also achieved by an isolated cis-regulatory element, including nucleic acid molecule [0040] a) a nucleic acid molecule having a nucleotide sequence according to SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 29, SEQ ID NO: 30, SEQ ID NO: 31, SEQ ID NO: 32, SEQ ID NO: 33, SEQ ID NO: 34, SEQ ID NO: 42, SEQ ID NO: 43 or SEQ ID NO: 44, [0041] b) a nucleic acid molecule having a nucleotide sequence according to the core sequence from SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 29, SEQ ID NO: 30, SEQ ID NO: 31, SEQ ID NO: 32, SEQ ID NO: 33, SEQ ID NO: 34, SEQ ID NO: 42, SEQ ID NO: 43 or SEQ ID NO: 44, or [0042] c) a nucleic acid molecule having a nucleotide sequence complementary to one of the nucleotide sequences from a) or b).
[0043] A cis-regulatory element including a nucleic acid molecule with a nucleotide sequence according to SEQ ID NO: 24, SEQ ID NO: 25 or SEQ ID NO: 26 is associated with the motif group 1, cis-regulatory element with a nucleotide sequence according to SEQ ID NO: 30, SEQ ID NO: 31 or SEQ ID NO: 32 is associated with the motif group 5, a cis-regulatory element with a nucleotide sequence according to SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 or SEQ ID NO: 19 is associated with the motif group 11, a cis-regulatory element with a nucleotide sequence according to SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22 or SEQ ID NO: 23 is associated with the motif group 12, a cis-regulatory element with a nucleotide sequence according to SEQ ID NO: 33 is associated with the motif group 18, a cis-regulatory element with a nucleotide sequence according to SEQ ID NO: 27 or SEQ ID NO: 28 is associated with the motif group 21, a cis-regulatory element with a nucleotide sequence according to SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 42 or SEQ ID NO: 43 is associated with the motif group 21n, a cis-regulatory element with a nucleotide sequence according to SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 44 or SEQ ID NO: 15 is associated with the motif group 27, and a cis-regulatory element with the nucleotide sequence according to SEQ ID NO: 34 is associated with the motif group 32.
[0044] A cis-regulatory element according to the invention can have a length of fewer than 50 nucleotides, preferably of fewer than 40 nucleotides, and especially preferred of fewer than 30 nucleotides. The core sequence of a cis-regulatory element according to the invention can have a length of fewer than 20 nucleotides, preferably of fewer than 15 nucleotides, and especially preferably of fewer than 10 nucleotides, while the core sequence should not be shorter than 6 nucleotides.
[0045] In addition to having the identified novel core sequence, some single sequences of the cis-regulatory elements according to the invention can also have the core sequence of a known cis-regulatory element imparting pathogen-/elicitor-inducibility, which cis-regulatory element can also possibly influence the specificity of the novel identified core sequence and/or of the identified single sequence. An example is the cis-regulatory element Cis05 (SEQ ID NO: 14) that also has a W-box core sequence between the nucleotide positions 29 and 35 in addition to the identified core sequence between the nucleotide positions 14 and 20. Extensive mutation analyses were conducted pertaining hereto (see FIGS. 5A, 5B and 11).
[0046] A cis-regulatory element according to the invention can be used in a chimeric promoter, wherein the cis-regulatory element imparts a specific pathogen- and/or elicitor inducibility to the chimeric promoter. The present invention thus also includes a chimeric promoter that, when induced by a pathogen infection or a treatment with a pathogenic elicitor, is suitable for effecting an expression of an operatively linked nucleic acid molecule of interest, e.g. of a heterologous DNA sequence, in a plant cell, and which chimeric promoter also includes a minimal promoter and at least one cis-regulatory element according to the invention. A single cis-regulatory element according to the invention in such a chimeric promoter by itself is in the position to impart a significant pathogen- and/or elicitor inducibility. Thus this single cis-regulatory element is itself sufficient for constructing a pathogen-/elicitor-responsive chimeric promoter in combination with a minimal promoter.
[0047] Such a chimeric promoter, containing only one or more inventive cis-regulatory elements as cis-regulatory elements, is preferably only pathogen- and/or elicitor-responsive, i.e. this promoter is not or only minimally inducible by other stimuli such as abiotic stress. After pathogen/elicitor contact, the induction of such a chimeric promoter comprising one or more inventive cis-regulatory elements is at least 2-fold, preferably at least 10-fold, or particularly preferably at least 25-fold greater than the induction without pathogen/elicitor contact (background activity).
[0048] In a preferred embodiment, the induced expression is limited only locally to the infection site, i.e. comparably or to a lesser extent than occurs in the controlled expression of natural PR genes. Especially preferably the transcription activation occurs controlled by a chimeric promoter of the present invention just in the cells that come into contact with the pathogen or the pathogenic elicitor. However, owing to cell-cell interactions, a transcription activation can occur in cells surrounding the infection site(s).
[0049] Chimeric promoters of the present invention, however, are not limited to those that are exclusively pathogen responsive. The induced expression can be further specified by combination with additional regulatory elements, e.g. by combination with a cis-regulatory element that, for example tissue specificity, storage inducibility, cold- or heat inducibility or a specific activity in certain stages of development. Chimeric promoters of the invention can also comprise at least one combination of at least two cis-regulatory elements, wherein said cis-regulatory elements comprise at least one combination of at least one inventive cis-regulatory element. Known cis-regulatory elements imparting pathogen-/elicitor-inducibility, such as W-box, S-box or D-box (see WO 00/29592), can also be used as further cis-regulatory elements in the combination to construct a chimeric promoter.
[0050] The invention moreover also includes a chimeric promoter that comprises one or more monomers and/or one or more multimers of the inventive cis-regulatory elements. Dimers and tetramers are preferred multimeric forms. Monomers separately or individual monomers within a multimer can have different orientations, i.e. they can have a complementary arrangement, for example. Inventive cis-regulatory elements of a multimeric form can be functionally linked to one another, i.e. they show in multimeric form a synergistic or antagonistic effect on, for example, the binding capacity of the transcription factor that distinguishes the characteristic core sequence motif of a certain motif group, among other things. The invention thus likewise includes a chimeric promoter that, when induced by a pathogen infection or treatment with a pathogenic elicitor, is suitable for effecting an expression of an operatively linked nucleic acid molecule of interest in a plant cell and which chimeric promoter furthermore comprises a minimal promoter and at least two inventive cis-regulatory elements, wherein the at least two cis-regulatory elements can be present functionally linked in homo- and/or heteromeric form.
[0051] Subsequent to pathogen/elicitor contact, the induction of a chimeric promoter comprising at least one multimer of the inventive cis-regulatory elements is at least 2-fold, preferably at least 10-fold or particularly preferably at least 25-fold greater than the induction without pathogen/elicitor contact (background activity).
[0052] In a preferred exemplary embodiment of the chimeric promoters of the present invention, the distance from minimal promoter and to the first upstream inventive cis-regulatory element is between 0 and 300 base pairs, preferably between 0 and 70 base pairs and particularly preferably less than 10 base pairs. Additionally or alternatively, the distance between two identical monomers of the inventive cis-regulatory elements in a multimeric form is preferably 0 to 10 base pairs. Preferably two separate multimers in a chimeric promoter of the invention are separated by approximately 0 to 50 base pairs.
[0053] In experimental analyses, specific combinations of inventive cis-regulatory elements with other inventive cis-regulatory elements or with other known regulatory elements or fragments such as S-box, D-box or Gst1 showed an advantageous and surprising effect with regard to promoter properties such as a low background activity or an especially specific or an especially strong inducibility (up to 183-fold 2.times.Cis13-2.times.Cis05). In this case some combinations showed a synergistic effect with regard to a certain promoter property such as, for example, with regard to the induction factor (compare e.g. 2.times.Cis13-2.times.Cis05 in parsley, FIGS. 12 and 13), while although some combinations had an antagonistic effect with regard to the induction factor, for example, in this case an especially low background activity nevertheless developed (e.g. 4.times.Cis05-2.times.D in sugar beet). The invention furthermore includes all combinations and combination possibilities of the inventive cis-regulatory elements with themselves and with known cis-regulatory elements that have an advantageous synergistic or antagonistic effect on the inducibility of the promoter. Advantageous combinations are those that can be selected from the following group: 4.times. sCis05, 4.times. 20u_M1_S1, 4.times. 27G-8_M1_S1, 4.times. 38M_M1_S1, 4.times. 18H_M2_S3, 4.times. 18H_M2_S1, 4.times. GG13_M1_S2, 4.times. 21S_M3_S1, 4.times. 30I-8_M1_S2, 2.times. Cis02-2.times. Cis02, 2.times. Cis02-2.times. Cis05, 2.times. Cis02-2.times. Cis12, 2.times. Cis02-2.times. Cis13, 2.times. Cis02-2.times. D, 2.times. Cis02-2.times.S, 2.times. Cis02-Gst1, 2.times. Cis02-2.times. 30I-8_M1_S2, 2.times. Cis05-2.times. Cis02, 2.times. Cis05-2.times. Cis05, 2.times. Cis05-2.times. Cis12, 2.times. Cis05-2.times. Cis13, 2.times. Cis05-2.times. D, 2.times. Cis05-2.times. S, 2.times. Cis05-Gst1, 2.times. Cis05-2.times. 30I-8_M1_S2, 2.times. Cis12-2.times. Cis02, 2.times. Cis12-2.times. Cis05, 2.times. Cis12-2.times. Cis12, 2.times. Cis12-2.times. Cis13, 2.times. Cis12-2.times. D, 2.times. Cis12-2.times. S, 2.times. Cis12-Gst1, 2.times. Cis12-2.times. 30I-8_M1_S2, 2.times. Cis13-2.times. Cis02, 2.times. Cis13-2.times. Cis05, 2.times. Cis13-2.times. Cis12, 2.times. Cis13-2.times. Cis13, 2.times. Cis13-2.times. D, 2.times. Cis13-2.times. S, 2.times. Cis13 - Gst1, 2.times. Cis13-2.times. 30I-8_M1_S2, 2.times. D-2.times. Cis02, 2.times. D-2.times. Cis05, 2.times. D-2.times. Cis12, 2.times. D-2.times. Cis13, 2.times. D-2.times. 30I-8_M1_S2, 2.times. S-2.times. Cis02, 2.times. S-2.times. Cis05, 2.times. S-2.times. Cis12, 2.times. S-2.times. Cis13, 2.times. S-2.times. 30I-8_M1_S2, Gst1-2.times. Cis02, Gst1-2.times. Cis05, Gst1-2.times. Cis12, Gst1-2.times. Cis13, Gst1-2.times. 30I-8_M1_S2, 2.times. 30I-8_M1_S2-2.times. Cis02, 2.times. 30I-8_M1_S2-2.times. Cis05, 2.times. 30I-8_M1_S2-2.times. Cis12, 2.times. 30I-8_M1_S2-2.times. Cis13, 2.times. 30I-8_M1_S2-2.times. D, 2.times. 30I-8_M1_S2-2.times. S, 2.times. 30I-8_M1_S2-Gst1 and 2.times. 30I-8_M1_S2-2.times. 30I-8_M1_S2.
[0054] Particularly advantageous combinations having surprising effect with regard to induction factor and activity are listed in Table 3.
[0055] The present invention additionally also relates to chimeric promoters that comprise at least one of the above-mentioned combinations of cis-regulatory elements.
[0056] In a preferred embodiment of the chimeric promoters of the present invention, the minimal promoter originates, for example, from a CaMV35S promoter, for monocotyledonous plants from, for example, the wheat TaPal promoter (SEQ ID NO: 39), the corn ZmUbiquitin promoter (SEQ ID NO: 40) or the rice OsGns1 promoter (SEQ ID NO: 38), or for dicotyledonous plants from known minimal promoters (WO 07/147395). Minimal promoters from other sources can furthermore also be used to construct a chimeric promoter in the sense of the present invention.
[0057] A chimeric promoter of the present invention regardless fulfills the essential requirements placed on the stringent expression regulation of a transgene in a genetic engineering approach, e.g. for producing a pathogen-/disease-resistant plant. The transgene is a nucleic acid molecule of interest, e.g. a heterologous DNA sequence, operatively linked with the chimeric promoter, which transgene codes, for example, for a resistance gene (R gene), an auto-activated resistance gene, an avirulence gene, a different effector, a protein that is toxic to at least one pathogen, signal transduction components, or a protein which when synthesizing phytoalexins codes a double-stranded RNA for forming siRNAs directed against a pathogen or an antimicrobial peptide. Moreover, numerous experiments on parsley (Petroselinum erispum), Arabidopsis thaliana, wheat (Triticum sp.) and sugar beet (Beta vulgaris) demonstrated that a chimeric promoter of the present invention functions across species and can be used across species (see, for example 4.times.Cis05 in parsley, sugar beet, and wheat).
[0058] The definition of a family motif for each motif group 1, 5, 11, 12, 21, 21n, and 27 creates possibilities for a person skilled in the art in the construction of chimeric promoters of the present invention. The observed activities of the different members of the motif groups 27 and 12 (FIG. 8) demonstrate that the flanking sequence regions can be used for customized fine-tuning of the desired expression level. This also applies to a species-dependent coordination of the expression level. The family motif reproduces the variation possibilities of individual bases in said flanking regions and thus teaches a person skilled in the art the extent to which the flanking regions can be modified. A person skilled in the art moreover obtains information from the family motif about how strongly conserved individual base positions are within the family motif. Here it can be assumed that the modification of a strongly conserved base has a more marked effect on the resulting properties of the chimeric promoter than a weakly conserved base.
[0059] The invention moreover also relates to a recombinant gene comprising a chimeric promoter of the present invention. The recombinant gene is preferably formed such that the chimeric promoter is operatively linked to a nucleic acid molecule, e.g. a heterologous DNA sequence. Such a heterologous DNA sequence codes in particular for a (poly) peptide, a cytotoxic protein (such as Bt toxin, avirulence protein or enzymes such as glucose oxidase that generate reactive oxygen species), an antibody, an antisense RNA, a sense RNA, a transcription factor, a protease, a nuclease, a lipase, an enzyme inhibitor or a measurable marker (such as luciferase, GFP or .beta.-galactosidase). The last-mentioned marker and other markers known from the prior art can be used in test systems to determine the pathogen specificity of a chimeric promoter of the present invention or to identify effectors that promote or inhibit an induction of the chimeric promoter.
[0060] Chimeric promoters of the invention can also be used in RNAi-based processes for "gene silencing", wherein the operatively linked nucleic acid molecule of interest such as an anti-sense RNA, for example, codes a sense-RNA or a double-stranded RNA (dsRNA). The RNA molecule can then represent a short nucleotide sequence (generally at least 10 nucleotides, preferably at least 14 nucleotides and optionally up to 100 or more nucleotides long), which short nucleotide sequence is essentially complementary to a specific mRNA sequence and/or to a DNA sequence of a gene of interest. Standard methods of RNAi technology are described in the prior art.
[0061] In principle, it is possible to modify the operatively linked coding sequence such that the product of the translation is localized in a desired cell compartment such as nucleus, endoplasmic reticulum, mitochondrium, cytoplasm or vacuole or also extracellularly (apoplastic). Suitable methods for such modification are known to a person skilled in the art from the prior art (Gorlich, Science 271 (1996), 1513-1518; Hicks, Plant Physiol. 107 (1995), 1055-1058; Rachubinski, Cell 83 (1995), 525-528; Schatz, Science 271 (1996), 1519-1526; Schnell, Cell 83 (1995), 521-524; Verner, Science 241 (1988), 1307-1313; Vitale, BioEssays 14 (1992), 151-160).
[0062] A recombinant gene of the present invention can be used both alone and as part of a vector. The present invention thus also relates to a vector that comprises the chimeric promoter of this invention or the recombinant gene of the invention. The preferred vector is a plant expression vector that furthermore also preferably comprises a selection marker for plants. Examples of suitable markers have already been specified above. Methods for constructing such vectors are known from the prior art to a person skilled in the art, e.g. described in Sambrook, Molecular Cloning A Laboratory Manual, Cold Spring Harbor Laboratory (1989) N.Y. and Ausubel, Current Protocols in Molecular Biology, Green Publishing Associates and Wiley Interscience, N.V. (1989).
[0063] The present invention additionally relates to a prokaryotic or a eukaryotic host cell that comprises a chimeric promoter, an recombinant gene or an inventive vector, wherein the chimeric promoter itself or as part of the recombinant gene or as part of the vector or respectively as part of the chimeric promoter such as a cis-regulatory element is heterologous to the prokaryotic or eukaryotic host cell, thus for example originates from a cell or from an organism having a different genetic background, or is homologous to the prokaryotic or eukaryotic host cell but is localized in a different genetic environment and thus differs from the naturally occurring chimeric promoter or its part.
[0064] The chimeric promoter, the recombinant gene or the inventive vector can either be integrated in the genome of the prokaryotic or eukaryotic host cell, preferably stably integrated, or can remain in an extrachromosomal form such as a plasmid in the cell.
[0065] The invention moreover provides a method for producing a transgenic plant, which method includes introducing a chimeric promoter, a recombinant gene or a vector according to the present invention into at least one cell of the plant or said method includes introducing a chimeric promoter, a recombinant gene or a vector according to the present invention into at least one plant cell in a cell culture from which the transformed or transgenic plant is subsequently regenerated. The chimeric promoter, the recombinant gene or the vector is preferably integrated, especially preferred stably integrated, into the genome of the plant. The nucleic acid molecule can be connected to additional regulatory sequences like the 3' end to a poly-A tail for the expression of the nucleic acid molecule of interest in plant cells under the control of a chimeric promoter according to the present invention. Methods for introducing genes or genetic material into a plant or into a plant cell as well as methods for regenerating transformed plant cells are known from the prior art, for example Agrobacterium tumefaciens- or Agrobacterium rhizogenes-conveyed transformation of plant cells or tissues with T-DNA, die protoplast fusion, injection, electroporation, vacuum infiltration or biolistic methods. Methods for preparing suitable vectors for introducing genes or genetic material into a plant or into a plant cell are likewise routine for a person skilled in the art (Sambrook, Molecular Cloning A Laboratory Manual, Cold Spring Harbor Laboratory (1989) N.Y. and Ausubel, Current Protocols in Molecular Biology, Green Publishing Associates and Wiley Interscience, N.V. (1989)).
[0066] In an alternative embodiment, a plant cell can be modified such that said plant cell expresses an endogenous gene under the control of an inventive chimeric promoter or under the control a native promoter of the endogenous gene, which native promoter was modified by inventive cis-regulatory elements. Introducing such a chimeric promoter, which naturally does not regulate the expression of a certain gene or of a certain genomic sequence, at the desired location in the plant genome or the introduction of inventive cis-regulatory elements into a native promoter can be effected using known standard methods such as targeted integration (`gene targeting`) via zinc finger-nucleases (Urnov et al., Nature Reviews 2010_Genome editing with engineered zinc finger nucleases; Townsend et al., Nature 2009_High-frequency modification of plant genes using engineered zinc-finger nucleases) or TAL effector nucleases (WO 2010/079430; WO 2011/072246). The modification of a native promoter of an endogenous gene also means the additional introduction of an inventive cis-regulatory element into the native promoter, which already naturally has an inventive cis-regulatory element, and thus a multimerization of present cis-regulatory elements. Such a modified promoter can, compared to the native version, have changed properties with regard to specificity, expression level or background activity, for example.
[0067] Using known methods, modified plants can be regenerated from the modified plant cells.
[0068] The invention thus relates again to transgenic (transformed) plants described below, which plants were transformed with a chimeric promoter, a recombinant gene, a vector according to the present invention, and described plants modified by introducing at least one inventive cis-regulatory element or an inventive chimeric promoter. Transgenic or modified plants can come from any desired plant species. They can be monocotyledonous, dicotyledonous or angiosperm plants, preferably belonging to plant species of agricultural or horticultural interest, for example corn, rice, wheat, rye, barley, oats, sorghum, potatoes, oilseed rape, sunflower, soybean, cotton or sugar beet.
[0069] In a preferred embodiment of the invention, a transgenic or modified plant is resistant or demonstrates an increased resistance to one or more pathogens in comparison to a non-transgenic or non-modified plant of the same species (wild type).
[0070] Included in the present invention are furthermore a plant part, a plant tissue, a plant cell or a seed of the transgenic and of the modified plant of the present invention, wherein said plant part, plant tissue, plant cell or seed likewise has the transgene introduced into the plant or the introduced modification.
BRIEF DESCRIPTION OF THE DRAWINGS
[0071] Embodiments of the present invention are described by way of example with reference to the accompanying figures and sequences:
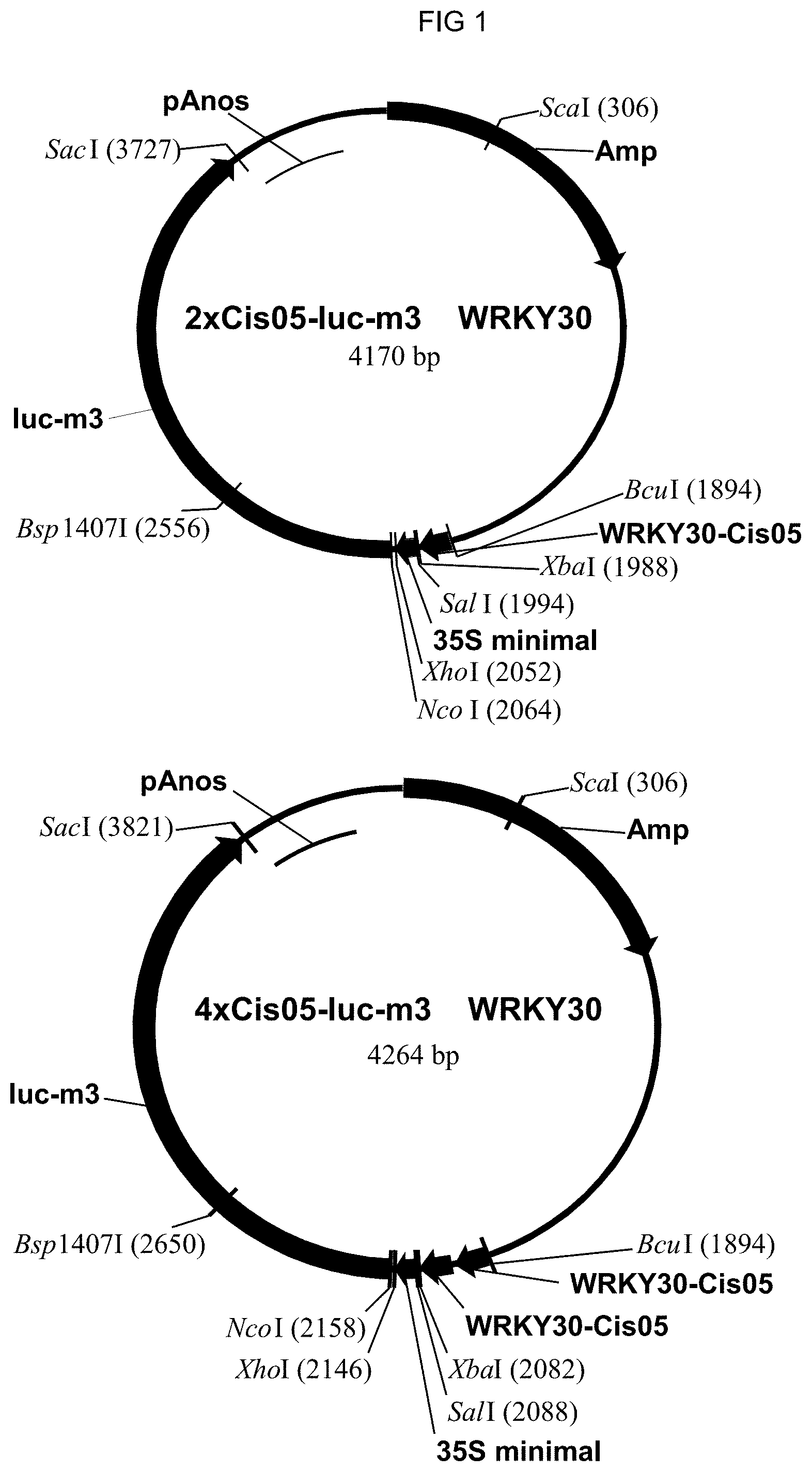
[0072] FIG. 1: The plasmids including 2.times.Cis05 element and the multimerized 4.times.Cis05 element are shown as an example of cloning the single sequences as chimeric promoters. The plasmids are derived from pBT10-GUS (Sprenger and Weisshaar (2000): The Plant Journal 22, 1-8). The structure of the other plasmids is in accordance therewith. Amp: ampicillin resistance; WRKY30-Cis05: doubled single sequence Cis05. 35S-minimal: 35S-minimal promoter; Luc-m3: luciferase reporter gene; pAnos: NOS terminator.
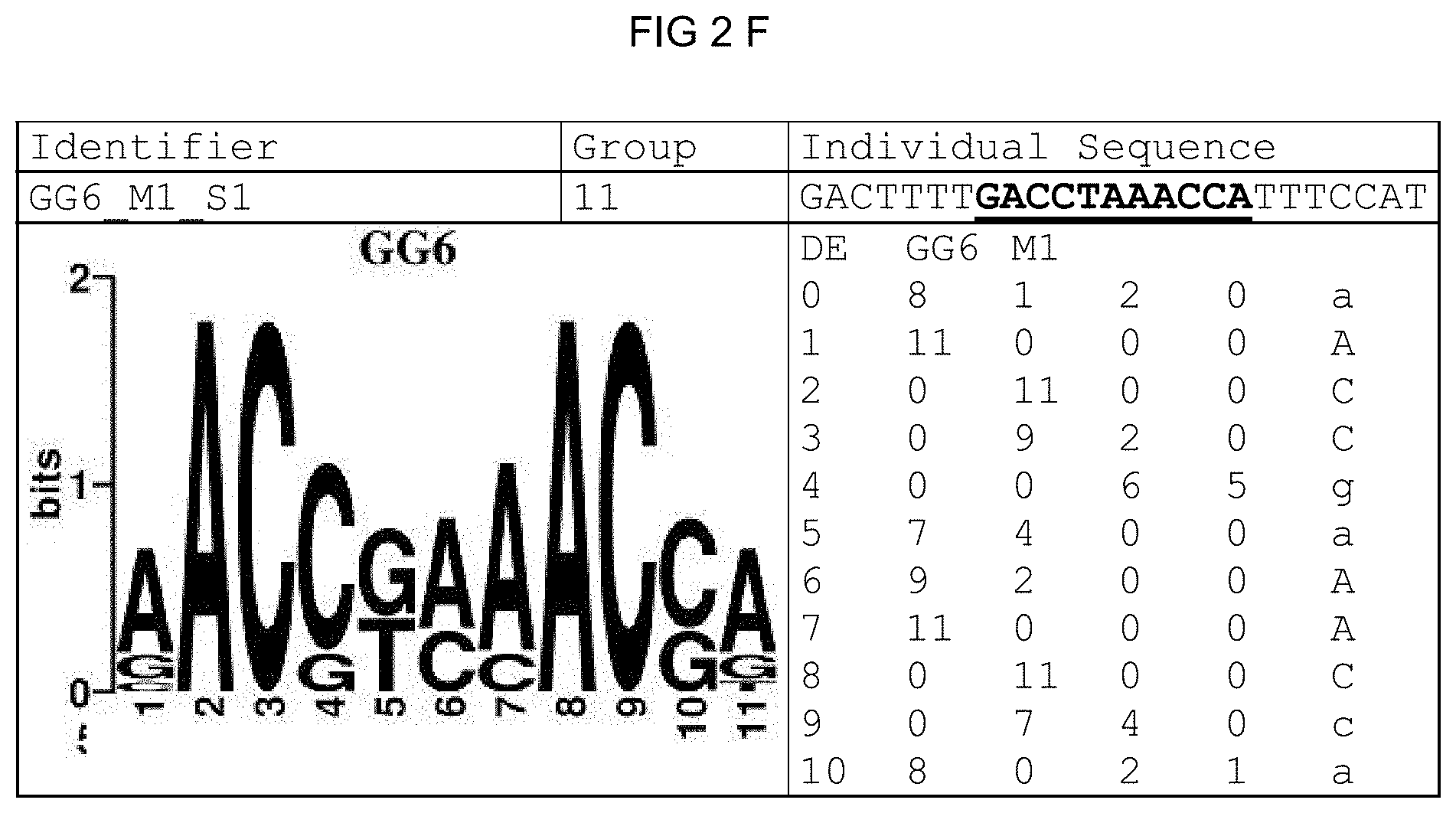
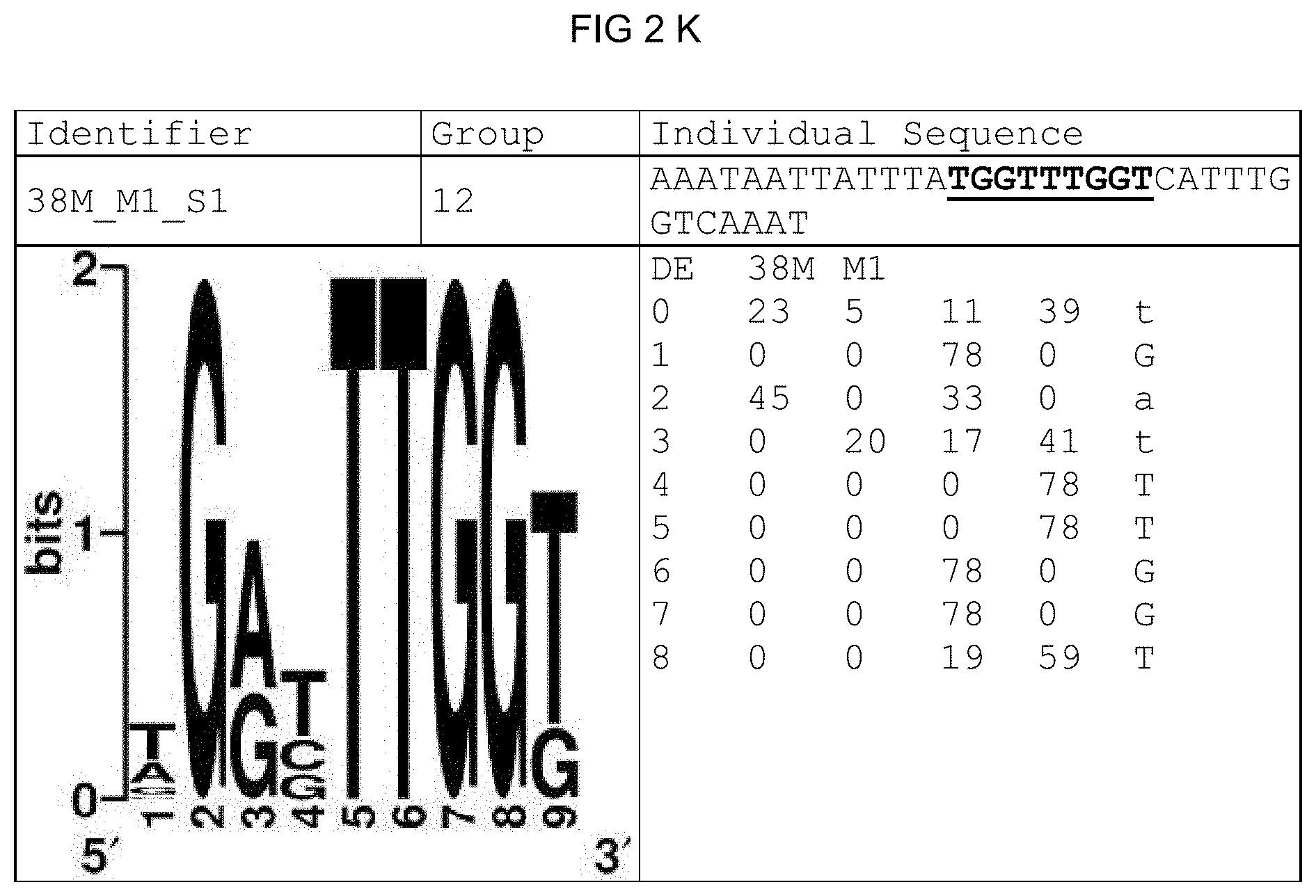
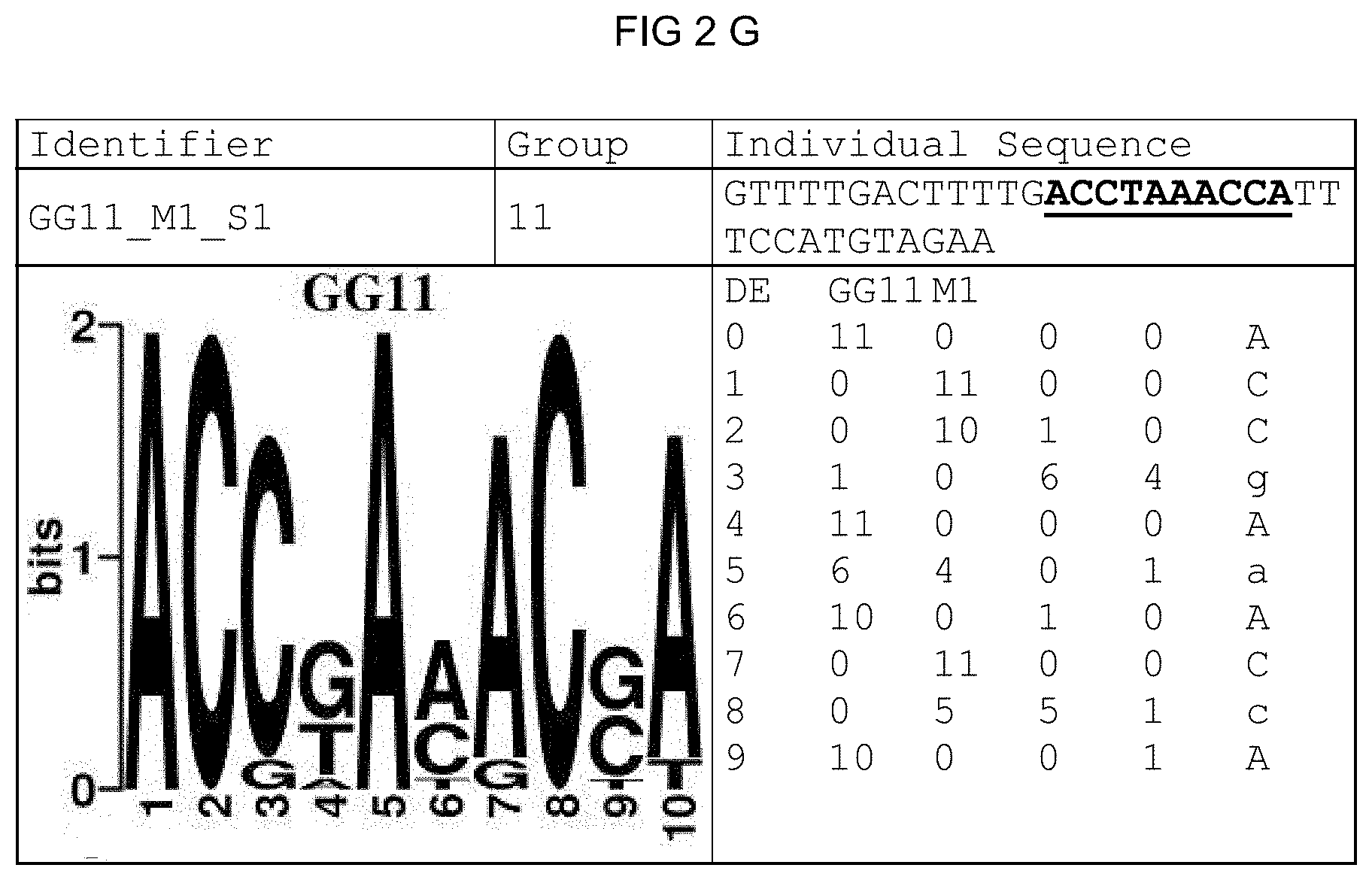
[0073] FIGS. 2A-2X: Species-wide pathogen-inducible single sequences. The upper lines respectively reflect the identifier of the single sequence, the motif group, and the single sequences themselves. The core sequences of the motif that led to the selection of the single sequence is written in boldface type and underlined. Therebeneath are respectively reflected the motif logo of the underlying bioinformatically identified motif and its matrix. If more than one single sequence of a motif was tested, they are summarized. FIG. 2A discloses SEQ ID NO: 26; FIG. 2B discloses SEQ ID NO: 25; FIG. 2C discloses SEQ ID NO: 24; FIG. 2D discloses SEQ ID NOS 30-31; FIG. 2E discloses SEQ ID NO: 32; FIG. 2F discloses SEQ ID NO: 19; FIG. 2G discloses SEQ ID NO: 16; FIG. 2H discloses SEQ ID NO: 17; FIG. 2I discloses SEQ ID NO: 18; FIG. 2J discloses SEQ ID NOS 20-21; FIG. 2K discloses SEQ ID NO: 22; FIG. 2L discloses SEQ ID NO: 23; FIG. 2M discloses SEQ ID NO: 33; FIG. 2N discloses SEQ ID NO: 27; FIG. 2O discloses SEQ ID NO: 28; FIG. 2P discloses SEQ ID NOS 7-8 and 12; FIG. 2Q discloses SEQ ID NO: 9; FIG. 2R discloses SEQ ID NO: 10; FIG. 2S discloses SEQ ID NO: 11; FIG. 2T discloses SEQ ID NOS 13-14; FIG. 2U discloses SEQ ID NO: 15; FIG. 2V discloses SEQ ID NO: 34; FIG. 2W discloses SEQ ID NO: 42; and FIG. 2X discloses SEQ ID NO: 43.
[0074] FIGS. 3A-3G: Summary of all single sequences of a motif group, which sequences tested positive, and representation of the sequence- and family-motifs derived therefrom. The respective motif group is specified in the top line. An alignment of all single sequences that tested positive is shown therebeneath, which alignment includes at least the core sequences and, if present, additional conserved bases. The bases, from which the core sequence motif is derived, are surrounded by a border. FIG. 3A discloses SEQ ID NOS 89-91; FIG. 3B discloses SEQ ID NOS 92-94; FIG. 3C discloses SEQ ID NOS 95 and 95-97; FIG. 3D discloses SEQ ID NOS 98-101; FIG. 3E discloses SEQ ID NOS 102-103; FIG. 3F discloses SEQ ID NOS 104-108, 106 and 109-111; and FIG. 3G discloses SEQ ID NOS 112-115.
[0075] FIG. 4: Phylogenetic tree of the identified motif groups. The phylogenetic tree was generated via a cluster analysis using the STAMP web server. The number of the motifs contained in the motif group is surrounded by parentheses behind the number of the motif group.
[0076] FIG. 5A) Mutagenesis of the Cis05-single sequence. The sequence used contains both the Cis05 motif (bold-face, underlined) and a W-box (bold-face and surrounded by a border). Mutations were introduced in both motifs. As described, parsley assays were conducted on induction with the tetramerized mutated derivatives using the PAMP PEP25. The PAMP-induced activity of the chimeric promoters was measured after 4, 8, and 24 hours (right-hand side). Mutations in the Cis05-motif (Cis05mut1) and in the W-box (Cis05mut2) lead to a significant decline of the induced activity. Only when both motifs are mutated (Cis05mut1+2) is there a complete loss of inducibility.
[0077] FIG. 5B) sCis05 is a shortened variant of Cis05 containing only the Cis05 motif and the W-box no longer. The PEP25 inducibility of sCis05 and mutated derivatives was tested as described in FIG. 5A. Both bars represent two biological replicates (independent transformations). For guidance, the W-box consensus is shown beneath the sCis05 derivatives. FIG. 5A discloses SEQ ID NOS 14 and 116-119, respectively, in order of appearance. FIG. 5B discloses SEQ ID NOS 44 and 120-124, respectively, in order of appearance.
[0078] FIG. 6A: Elicitor-responsive reporter gene expression of the chimeric promoters 4.times.30I-8_M1_S2 and 4.times.18H_M2_S1 with mutations in the single sequences. The mutated bases are underlined. Elicitation was effected by PEP25 in parsley protoplasts. The nucleotide sequences of the mutated derivatives of the single sequences are shown beneath the diagrams as +, with elicitor PEP25; -, without elicitor. 2S2D: Positive control; MS23GUS: negative control (empty vector). FIG. 6A discloses SEQ ID NOS 8, 125-127, 20, and 128-130, respectively, in order of appearance. FIG. 6B discloses SEQ ID NOS 131-137, respectively, in order of appearance.
[0079] FIG. 6B: Elicitor-responsive reporter-gene expression of the chimeric promoters 4.times.20u_M1_S1 and 4.times.27G-8_M1_S1 with mutations in the single sequences. The mutated bases are underlined. Elicitation was effected by PEP25 in parsley protoplasts. The nucleotide sequences of the mutated derivatives of the single sequences are shown beneath the diagrams.
[0080] FIG. 7: Binary vector for the transformation of the luciferase reporter gene under the control of the chimeric promoters in sugar beet. The vector with the chimeric promoter 4.times.Cis05 is shown as an example. nptll: kanamycin resistance; WRKY30-Cis05: double single sequence Cis05. 35S minimal: 35S minimal promoter; luc-m3: luciferase reporter gene; Anos: Nos terminator.
[0081] FIG. 8A: Cercospora beticola induced promoter activity in stably transformed sugar beets. Of the constructs given, the luciferase activity subsequent to C. beticola infection of in-vitro plants was determined for a plurality of independent transformants (4 replicas per transformant and point in time). The median was calculated from the measured values obtained. The top diagram summarizes the results for single sequences from the motif group 12, while the bottom diagram summarizes the results for elements from the motif group 27. Co: Control (mock infection); inf; infection with C. beticola; 1 d -4 d: days post inoculation (d.p.i.). Control: Non-transgenic plants.
[0082] FIG. 8B: Cercospora beticola-induced promoter activity of 4.times.Cis05 and its derivatives in stably transformed sugar beets. For 4.times.Cis05 and its derivatives the luciferase activity after C. beticola infection was determined in a plurality of independent transformants (4 replicas per transformant and point in time). The median was calculated from the measured values obtained. The sequence of the different derivatives is given in FIGS. 5A and 5B. For each of the different derivatives, it is additionally indicated under the diagram whether the derivative contains the Cis05 motif or the W-box. Co: Control (mock infection); inf; infection with C. beticola; 1 d -4 d: days post inoculation (d.p.i.) non transgenic: non-transgenic control.
[0083] FIG. 9: Cercospora beticola-induced promoter activity of the chimeric promoter 4.times.GG6_M1_S1 in stably transformed sugar beets. The luciferase activity after C. beticola infection of in-vitro plants was determined for 10 independent transformants with the construct 4.times.GG6_M1_S1-luc (4 replicas per transformant and time period). Co: Control (mock infection); inf; infection with C. beticola; 2 d, 3 d, 4 d and 7 d: days post inoculation (d.p.i.); 3DC4156: Non-transgenic control plants.
[0084] FIG. 10: Plasmid card of the plasmids used for the transient tests in wheat. The plasmid with the chimeric 4.times.Cis05 promoter is shown as an example. Ruc: Renila-luciferase reporter gene. AMP: Ampicillin resistance. WRKY30-Cis05: Double single sequence Cis05.
[0085] FIG. 11: Test of the induction by Fusarium of the chimeric promoter 4.times.Cis05 and its mutated derivatives with mutations in the Cis05 motif (Cis05mut1), the W-box (Cis05mut2) or in both motifs (Cis05mut1+2). The corresponding constructs were transiently transformed in wheat and the luciferase activity was measured 20 hours post incubation with Fusarium. 4.times.Cis05-dam/dcm identifies an experiment in which the plasmid DNA of a non-methylated E. Coli strain was used to exclude an induction by dam/dcm-methylated DNA (likewise a potential PAMP). If the core sequence of Cis05 mutated, the inducibility is entirely lost. In contrast, the mutation in the W-box has no effect. The sequences of Cis05 and its mutated derivatives are reproduced on the right-hand side. Mutated bases are highlighted in red. FIG. 11 discloses SEQ ID NOS 116-117, respectively, in order of appearance.
[0086] FIG. 12: Induced and non-induced activity of the chimeric combinatorial promoters after PEP25 induction in parsley. The tests were conducted in three biological replicates; the blue line reproduces the induction factor. In the bottom row beneath the diagram, the elements in 5' position are given, while in the top row the elements in 3' position are given. 30I8b is a different identifier for the single sequence 30I-8_M1_S2.
[0087] FIG. 13: Synergistic and antagonistic interactions of single sequences in the chimeric combinatorial promoters after PEP25 induction in parsley. The actual measured induction factor is reproduced in violet, while the expected induction factor based on the induction factors of the single elements is reproduced in blue. The ratios of both values are shown by the yellow line. If the dots of the yellow line are above the value 1, then the single elements show a synergistic interaction.
[0088] FIG. 14: Transgenic sugar beets with a 4.times.Cis05-RFP construct were infected with Cercospora beticola. The infection leads to activation of the chimeric 4.times.Cis05 promoter, which leads to the formation of the red fluorescing RFP protein. The protein is seen as red fluorescent under the microscope. As can be seen, the induction and thus the fluorescence are limited to the region around the penetration site or the infection location.
[0089] FIGS. 15A-15D: Exemplary schematic representations of a chimeric promoter in the sense of the invention. The chimeric promoter is operatively linked with a nucleic acid molecule of interest and includes (FIG. 15A) a heterologous cis-regulatory element next to a minimal promoter, (FIG. 15B) a dimer/multimer of a heterologous cis-regulatory element next to a minimal promoter or (FIG. 15C) two dimers/multimers, each having different heterologous cis-regulatory elements, next to a minimal promoter. Furthermore, (FIG. 15D) shows as an exemplary chimeric promoter a natural promoter including an endogenous minimal promoter and an endogenous cis-regulatory element, wherein said promoter was modified through integration of an additional homologous cis-regulatory element.
[0090] FIG. 16: Transgenic Arabidopsis plants with a tetramer of the cis-regulatory element Cis05 in a chimeric promoter that controls the expression of the GUS reporter gene to investigate the pathogen-induced, wound-induced, and tissue-specific activity of the element. Ten independent transformants were investigated for the promoter. One respectively representative line is shown. The image on the left-hand side (5 d.p.i. with H. arabidopsidis) shows the activity of the promoter subsequent to infection with the compatible pathogen Hyaloperonospora arabidopsidis, while the image on the right-hand side (Mock control) is the corresponding control. The promoter shows a clear induction through Hyaloperonospora arabidopsidis (dark coloration of the plant tissue).
[0091] On the right-hand image, a leaf is additionally cut into. A blue coloration at this cut surface would indicate a wound inducibility of the promoter with the tetramer of the cis-regulatory element Cis05.
Bioinformatic Identification of the Inventive Cis-Regulatory Elements:
[0092] Publicly available microarray-expression data are the basis for the bioinformatic identification of the novel cis-regulatory elements. Said expression data are deposited in databases such as TAIR, NASCArrays, Geo or ArrayExpress or can be obtained directly from corresponding publications of microarray experiments (e.g. Rhee et al., 2003; Craigon et al., 2004; Barret and Edgar, 2006; Brazma et al., 2006, Zipfel et al., 2004, 2006; Bulow et al., 2007; Wan et al., 2008). For the bioinformatic identification of novel cis-regulatory elements, initially the publicly available microarray-expression data were used to define groups of genes of the Arabidopsis thaliana plant, the expression of which is induced by pathogens such as P. syringae or B. cinerea or PAMPs like flg22 or chitin. Then the sequences of the promoters of these gene groups were extracted from the genome sequence of Arabidopsis thaliana (TAIR; http://www.arabidopsis.org). Using different known algorithms (MEME, Bioprospector, Alignace, BEST and the like the promoter sequences were examined for concentrated motifs.
Database Queries
[0093] A software tool was written for the database queries for identifying co-regulated genes, which tool permits identification of genes that are upregulated or induced together by up to six different stimuli. More than 700 database queries for identifying co-regulated genes were performed. This query process delivered more than 400 groups of genes induced together that are suitable for an identification of shared cis-regulatory motifs with the BEST software package (Che et al., 2005). By raising the necessary induction factor, the number of shared regulated genes in 77 groups was reduced to 120 genes. The total number of gene groups of co-regulated genes (2-120) was 510 thereafter.
[0094] Of these 510 gene groups, 500-bp or 1000-bp long promoter sequences upstream from the transcription start [SITE--what "TSS" stands for)] (TSS) of the co-regulated genes were extracted of which the TSS was known. The promoter sequences of the co-regulated gene groups were examined in accordance with conserved sequence motifs by the BEST, Cismodule, MD-Scan, BioProspector or MEME programs. Motif lengths of 5-10 nt, 10-15 nt and 15-20 nt were selected. Approximately 500.times.3 (motif lengths)=1500 queries were performed. In most cases, by lengthening the motif lengths, no further motifs were identified that did not already occur in the shorter motif lengths. In the BEST analyses, a motif was always classified when it was found by at least two of the four total BEST programs.
[0095] Given certain different stimuli, the same co-regulated genes, and therefore also the same motifs, were obtained. A summary of the identical motifs of redundant gene groups (GG) to novel motifs (gene groups table) followed, so that only unique motifs were compared in the comparative, systematizing analysis of all motifs among themselves.
[0096] A catalog was created that contains all identified motifs plus evaluation dataset and the sequence logos of the individual motifs. The sequence logos (Crooks et al., 2004), created under http://weblogo.berkeley.edu/, reflect the conservation of the nucleotide at the individual positions of the motif. The matrix was created from the sequences forming the motif, the sequences and the related genes thereof likewise being reflected.
[0097] The STAMP program (Mahony and Benos, 2007) that can be invoked from the Internet address http://www.benoslab.pitt.edu/stamp was used to compare the identified motifs to already known cis-regulatory elements (PLACE, Agris, Athamap). With the STAMP web server, similar motifs can furthermore be grouped into a motif group. The program issues a phylogenetic tree in which the similarities of the motif groups are shown as an overview (FIG. 4).
Proof of Pathogen Inducibility of the Identified Cis-Regulatory Elements
[0098] Since bioinformatoric approaches are prone to identifying false-positive sequences, the pathogen inducibility must be experimentally confirmed. For said experimental confirmation, the bioinformatically-identified sequences were cloned as a tetramer in front of a luciferase reporter gene using standard DNA cloning procedures and tested for their inducibility in a transient expression system in parsley by the PAMP PEP25.
[0099] Elements were selected for the experimental test that are novel and show no similarity to known cis-regulatory elements of the pathogen-conveyed induction. The single sequences given in Table 2 were synthesized in vitro and cloned into the plasmid MS23 with SpeI and XbaI or with SpeI and SalI linkers. MS23 carries either a .beta.-glucuronidase (GUS)-reporter gene or a luciferase reporter gene with a 35S minimal promoter. All elements were tetramerized and sequenced for testing. The plasmids including 2.times.Cis05 element and the multimerized 4.times.Cis05 element are shown in FIG. 1 as an example of cloning the single sequences as chimeric promoters.
TABLE-US-00003 TABLE 2 Examined Single Sequences. The table lists the novel identifiers and sequences of the examined potentially cis-regulatory elements. The bioinformatically identified core sequences are emphasized (bold-face type and underlined). The result of the PEP25/parsley test is reproduced in the last column (-: no induction; +: inducible). Single- sequence Motif SEQ identifier group AGI Single sequence Inducibility ID NO: 12i_M1_S1 1 At5g04340 TCTCATCTCTCGACACGCAACTTCC + 26 Cis09 1 At1g27730 TGCACACACACACACGTGTACTAGGTCAAACCAAACGT + 24 Cis12 1 At2g33580 CAAAAAGTCAACACATACGACGCGTTTCCATTGACTAAATA + 25 12c_M1_S1 4 At1g21100 TCTACTAGAGGCCCATTAGGACCGGCAT - 45 20u_M1_S1 5 At1g13990 TGTTGAGTCGTTTACGTCACGTCGAGAATTTTCTC + 30 20u_M1_S2 5 At4g05020 TGTCATTATTAATACGTGACGAAACTGTAGCTCTG + 31 28M-1_M1_S1 5 At1g09080 TTACGTGTCAAGAAGTGATTGGAGAGGACACTCTAC + 32 28M-1_M1_S2 5 At4g17500 AAGACAAGTTGAGAGAGACGAGACCAATCACAACA - 46 28M-8_M1_S1 6 At3g21520 ATCCAACATCTCGGACCGGATCAATGATTTATCAT - 47 24F_M1_S1 7 At2g40750 TCATCAATGTGACATAAGCAAAGCT - 48 3C_M1_S1 11 At3g51440 TTTGATACGGTTACGGTTAATTAACG - 49 GG6_M1_S1 11 At2g40140 GACTTTTGACCTAAACCATTTCCAT + 19 GG11_M1_S1 11 At2g40140 GTTTTGACTTTTGACCTAAACCATTTCCATGTAGAA + 16 GG6_M1_S2 11 At5g59820 AAGATTCTCATCCAACCGAAACGACTCTTTCGTTTT - 50 GG11_M1_S2 11 At5g59820 AGATTCTCATCCAACCGAAACGACTCTTTCGTTTTT - 51 GG11_M1_S3 11 At1g27730 TCTTCTTCATTTTACCAACACCACTTGCACACACAC - 52 22DDD_M1_S1 11 At3g14990 CCGTCTTAGTTTACCGAAACCAAAGTGGCTTTTTCT + 17 21G-2_M1_S1 11 At1g01560 AGTTGAATTAGTTCGGTTCGGTTCGGTTGATATTG - 53 21G-2_M1_S2 11 At3g55470 CGTAATAATGGTTTGGTTTGGTTTGATCAAGTCTT + 18 37E_M1_S1 12 At2g39200 CGATAAACTTGCGAAACCCTAAAA - 54 18H_M2_S1 12 At1g70170 CAACACAAAACGCAAACGCAGACCTC + 20 18H_M2_S2 12 At2g35980 TATTGGAAGTTTGGGGCAACATCAC - 55 16MM_M1_S1 12 At5g05340 TTCAACCCTATAAACCAAAACAAATAACAGAATGC - 56 38M_M1_S1 12 At4g23810 AAATAATTATTTATGGTTTGGTCATTTGGTCAAAT + 22 3I_M3_S1 12 At1g26390 CGCCTCAATCATGAAAACGAATCCTCTGTAGTAGTG - 57 18H_M2_S3 12 At1g70170 AATTGACAAAAGACACGCAAACGATTCCAACGACC + 21 26LLL_M1_S1 12 At1g67920 TTACCGACACGTAACCAAAACTCACCGAACACCGT - 58 38M_M1_S2 12 At2g29460 GTTTCGAACGGGAACCAAACCATAATATGCGATGC - 59 38M_M1_S3 12 At3g54960 ACACTATTGGTCTTGGTTTGGTTTATATGCACGAC - 60 23LLL_M1_S1 12 At2g30770 GAAAACGATGGGTTCCAAAACTGTCGCTAATAAACT - 61 37D_M1_S1 12 At1g32940 TCTCCACTCGTTGTGATTTGGTCTGCAAGAAAACTA - 62 23LLL_M1_S2 12 At1g01480 ACGTTTTGAAATATTGTTTTGGATGGAGATTTTTTC - 63 34G-4_M1_S1 12 At3g28740 ATTTTTCATTTCGCCCAAAACAATTATCCTAACGTT - 64 26LLL_M1_S2 12 At4g01700 AGTCAAAACGTAGACCAAAACAAAAACATGTAACT + 23 27H-8_M1_S1 12 At4g18430 TTTTATAACACTACCAAAACCAATAAGCCCTTTCGT - 65 26KKK_M1_S1 12 At2g43510 TCATCAAACCAATCGGTTTGGTCCTAAAGATAATT - 66 19Q_M1_S1 13 At4g35180 GTCAATATACACAGCCACCGAACAAATTACTCTAT - 67 21S_M1_S1 14 At2g14610 AAGCGATGTTTACGAACCCCAAAATC - 68 28H-9_M1_S1 14 At1g19020 AGATTTGTTCGAGAACCTTGAGAAA - 69 28H-9_M1_S2 14 At4g37370 TTGCTACTTCGAGAACATTGGTCAA - 70 20a_M1_S1 15 At5g04340 TTAGAAGTGGCTCGAGTGTTCTACTT - 71 20a_M1_S2 15 At5g20230 AAGAAAGACAATCGAGCCTAGAAATT - 72 12G_M2_S1 18 At1g61800 CCATACAATATAAACCACCAAACCATAACCACAAA + 33 30_M1_S1 18 At4g39950 AATAATGTTCAACGTTGGTGGTGGTACTCAAGATGG - 73 41J_M1_S1 19 At3g09940 TCAAATACAGGCAACCAAGACTCGAGATCCTCATCG - 74 37C_M1_S1 20 At3g51440 AGAAAAATATTGGGCCTACTGGGAA - 75 3M_M3_S1 20 At3g02800 AGATTCCTGAAGTGAGGTCCACCCTAAAATCCATTT - 76 GG8_M1_S1 21/21n At1g27730 CACACACGTGTACTAGGTCAAACCA + 27 27G-8_M1_S1 21/21n At2g38860 AGGACTTTTCACCAGTTGGACTTTGAAGCCACCAA + 28 27G-8_M1_S2 21 At1g72060 GGTTTAGTCAAAGTAAACAAGACTTTGACTGTTCA - 77 27B-10_M1_S1 21 At1g22890 TGAACTTAATCACTGTCATTGTTTTCGTAACAATTT - 78 26WW_M2_S1 21n At4g02380 CTCAAAGGCCAGAATTGACGCAGCCGTTT + 42 27B-10_M1_S3 21n At1g21120 CCTTGGCCCAGTCCTTGGTCGTCGTATC + 43 GG4_M2_S1 22 At3g14990 GAAAAATGTGTGTGTTTGTGTTAATT - 79 GG4_M2_S2 22 At5g59820 ATAGTTCCCAAACGGACACGAACACA - 80 30A-8_M1_S1 24 At3g49620 GCAAACTAACGCCGGCGGCCGTCTTG - 81 30I-8_M1_S1 27 At5g12930 ACAACAGACGACTTTTCATAATTCA + 7 30I-8_M1_S2 27 At1g26390 CTATATGACAAAAGTCAAACATAAA + 8 GG13_M1_S2 27 At3g26830 TGTTCACTTTGAAAAGTATTCTTTGAG + 9 30H-8_M1_S1 27 At1g35230 TAGCTGTTGAAATTTCCAAGAAAAT - 82 14S_M1_S1 27 At1g76960 CGATCAGACTTTTCTACGCAAGAGAA + 10 21S_M3_S1 27 At1g76960 TAATTTCTCTTGCGTAGAAAAGTCTGATCGGGAAG + 11 30I-8_M1_S3 27 At5g24110 TCGTTCTTCAGTCAAAAAGTCAAACTATCTCTCTC + 12 Cis02 27 At5g64905 GAGCGTGAATTGACTTTGACCAAAACCAAA + 13 Cis05 27 At5g24110 GGTCAGCATGTTGGACTTTCCAAATTCATTGACCAAAG + 14 Cis13 27 At1g26380 AAAATAAACAGCTACTTGACGAAAAGTCAAACCAAATTC + 15 22AAA_M1_S1 31 At2g40000 TTTTTCTCGTCCCCATCCTCTATCC - 83 12r_M1_S1 32 At1g73480 CAATCTACTCGTCTCTTCTCTTACAT + 34 GG3_M1_S1 32 At5g44420 TAGGTTCCTGCCCTCTCCGTTCCTCC - 84 GG3_M1_S2 32 At4g39980 TCGAAACCAACCCTCTCCCTTATAAA - 85 20EE_M1_S1 32 At4g39980 GAAACCAACCCTCTCCCTTATAAATA - 86 24P_M1_S1 32 At1g30135 TGTTTTGTTTCCACCGTCTCTCCCGTGTCCTCTCTC - 87
[0100] The tetramerized single sequences in transiently transformed parsley cells from a cell culture were tested in the following for their inducibility by the PAMP PEP25. In addition to a general validation, it should be ensured that the elements in different plant species can be induced and by different elicitors.
[0101] For the test in parsley, protoplasts were isolated from a five-day-old parsley cell culture. 35 ml cell culture were removed by centrifugation and resuspended in 90 ml of a sterile-filtered solution with 0.5% cellulase, 0.2% macerozyme R-10, and 0.24 M CaCl.sub.2 and incubated with stirring overnight at 26.degree. C. Thereafter, the released protoplasts were pelletized though centrifugation, washed with 40 ml 0.24 M CaCl.sub.2 and subsequently resuspended in 50 ml P5 medium (1 bag prepared media Gamborgs B-5, 1 mg 2.4 D, 96.9 g saccharose, pH 5.7 with 1M KOH, sterile filtered). After centrifugation, the protoplasts float on the surface of the P5 medium and can be removed. The purification with P5 medium was repeated 2.times..
[0102] For the transformation, 5 .mu.g of the promoter construct to be tested, 2.5 .mu.g of a constitutive renilla luciferase-expressing normalizing vector, and 200 .mu.l PEG in 10 ml were put in screw-cap test tubes. 200 .mu.l protoplasts were added, after which the mixture was carefully stirred and incubated for 20 min in the dark at room temperature. The reaction was subsequently stopped by the addition of 5 ml 0.275M CaNO.sub.3-Lsg. The transformed protoplasts were removed by centrifugation, held in 6 ml P5 medium and divided into 2 aliquots. One aliquot was elicited with Pep25 (final concentration: 300 ng/ml; sequence: VTAGAEVWNQPVRGFKVYEQTEMT) (SEQ ID NO: 88)) and the other served as control. After overnight incubation, the protoplasts were retrieved through centrifugation. The luciferase activity was determined with the Dual Luciferase Kit (Promega, Mannheim, Germany) in a Sirius Luminometer (Berthold Detection System GmbH, Pforzheim, Germany). The parsley cells were pelletized through centrifugation and lysed for 20 minutes at 4.degree. C. in 150 .mu.l PLB buffer (Passive Lysis Buffer; Promega, Mannheim, Germany). The cell residues were removed by centrifugation for 20 minutes at 13,000 rpm and 4.degree. C. in a bench centrifuge.
[0103] Various procedures were applied with the lysate depending on whether the luciferase reporter gene or the GUS reporter gene were used. If the luciferase reporter gene was used, a 5 .mu.l sample of the supernatant with the released luciferase is mixed with 50 .mu.l LARII buffer (Promega, Mannheim, Germany) in 5 ml test tubes (Sarstedt, item no. 55.476). The buffer contains the substrate of the luciferase so that the activity of the enzyme and thus of the promoter can be measured with the Luminometer. The measurement occurs with 2 seconds pre-measurement time and 10 seconds luciferase-measurement time. Thereafter, 50 .mu.l Stop & Glo buffer (Promega, Mannheim, Germany) is added and carefully mixed by drawing up. This buffer stops the luciferase activity and renders measurable the constitutive renilla-luciferase activity of the normalizing vector. This measured value is used for normalizing the different transformation efficiencies. The measurement occurs with 2 seconds premeasurement time and 10 seconds luciferase-measurement time.
[0104] If the .beta.-glucuronidase (GUS) reporter gene from E. coli was used (Jefferson et al., 1987), the proof is effected by an enzyme reaction in which the Substrate MUG is hydrolized into 4-MU. Then 4-MU is proven and quantified by its fluorescence. Using these methods, the activity with and without the PAMP PEP25 was measured for all single sequences examined.
[0105] Table 2 lists all tested single sequences. It additionally lists whether they were inducible by the PAMP PEP25 in parsley. The cis-regulatory elements (single sequences) identified as pathogen-inducible and the motifs thereof are summarized in FIG. 2.
Mutation Analyses of the Identified Core Sequences
[0106] Mutation analyses were conducted for selected single sequences to prove that the respective identified core sequence is responsible for the PAMP- and pathogen-inducibility of the single sequences. Mutated derivatives of the single sequences were created therefore. The mutated single sequences were synthesized as oligonucleotides. The cloning of the plasmids was conducted corresponding to the constructs with chimeric promoters without mutations. Thereafter, the constructs were tested, as previously described above, for their PEP25 inducibility in parsley. The results of the mutation analyses are reproduced in FIG. 5A, FIG. 5B, FIG. 6A, and FIG. 6B. It could be shown for all five elements examined that the identified core sequence is responsible for the inducibility.
[0107] This is especially significant for single sequences Cis05 and 30I-8 M1 S2 of the group 27. The core sequence of 30I-8_M1_S2 partially overlaps with a sequence (GTCAA) complementary to the W-box (TTGAC; Rushton et al., 1996). An overlapping W-box sequence was also found in the single sequences 30I-8_M1_S3, Cis02, and Cis13.
[0108] However, it could be shown through the mutation analyses, in particular through the variants 30I-8_M1_S2_mut2, that bases outside the W-box are vital for the PAMP inducibility (FIG. 6A). Other members of the group 27 show no overlapping W-box sequence. Correspondingly, the mandatory core sequence TGAC for the W-box is not part of the sequence- or family motif of the group 27 (FIG. 3F). Thus the motifs or the single sequences of the group 27 do not involve variants of the W-box.
[0109] A W-box sequence is found outside the core sequence in the single sequence Cis05. The mutation analyses could show here that this W-box does indeed convey PAMP inducibility. However, the mutation leads only the Cis05 core sequence or only the W-box to a reduction in the inducibility and only mutation of both elements leads to a complete loss of the inducibility. Thus a single sequence with two functional, PAMP- and pathogen-inducible cis-regulatory elements, the known W-box, and the novel Cis05-Motiv (FIG. 2T) are concerned. The combination of both elements shows a considerably higher activity than the single elements alone.
[0110] To isolate the sequence region of the element Cis05 motif (FIG. 2T) that conveys the inducibility, said element was tetramerized separately from the W-box (sCis05) and four additional mutated derivatives of the Cis05 motif were created. The derivatives were tested for their inducibility through the PAMP PEP25 in parsley as described above (FIG. 5B). It could be shown that in addition to the critical importance of the core sequence for the inducibility, both bases in 5'-direction prior to the core sequence are also essential for the inducibility of Cis05, which further confirms that it is not a W-box variant.
[0111] The core sequences of the elements 20u_M1_S1 and 27G-8_M1_S1 could also be confirmed through additional mutation analyses. The tests were conducted as already described above. The results of these mutation analyses are reproduced in FIG. 6B (on top: 20u_M1_S1; on the bottom: 27G-8_M1_S1).
Derivation of Family Motifs
[0112] Because the cis-regulatory elements identified in Arabidopsis promoters were tested in parsley, a biological functionality, which spans all plant species, of the identified cis-regulatory elements was ensured. The experiments showed, as expected, that not all of the bioinformatically identified sequences are functional. Roughly one third of the tested DNA sequences were inducible by PEP25 in parsley. The other sequences were false-positive sequences from the bioinformatic analysis or did not exhibit the desired species-spanning biological functionality. These important results could be used to identify the family motifs of the motif groups 1, 5, 11, 12, 21, 21n and 27 as well as the corresponding strongly conserved, characteristic core sequences (FIG. 3) on the basis of the functionally effective single sequences. To this end, all functionally effective single sequences belonging to a motif group were summarized and a consensus and a motif were derived therefrom. First the region was that is part of the core sequence in all underlying single sequences was defined as a characteristic core sequence motif of the family motif.
[0113] The core sequence of motif group 27 is also in the sequence LS10 (Lebel et al., 1998). In said latter sequence, a mutation including ten bases was generated in the native PR-1 promoter in that sequence region, which mutation led to a sharp decline of a SA- or INA-inducibility of the native promoter. The results shown there, however, do not enable the derivation of a motif or of a core sequence. Furthermore, the usability of LS10 for chimeric promotes is not shown. Moreover, the family motif of the motif group 27 differentiates the motif group from the LS10 sequence. The family motif excludes a C on the position 5, while in LS10 a C is present at said position. Furthermore, it excludes a G from position 17, while in LS10 a G is present as said position. Finally it necessitates a T or C at position 18, while an A is present in LS10 at this position.
Proof of the Inducibility by the Pathogenic Fungus Cercospora beticola in Stably Transformed Sugar Beets
[0114] The novel cis-regulatory elements should distinguish themselves in that they can be induced in different plant species by different PAMPs and pathogens. To demonstrate the inducibility in an agronomically important plant by an agronomically important pathogen, the single sequences that tested positive in parsley were stably transformed in sugar beets. To this end, the chimeric promoters were recloned with the tetramerized single sequences including the luc gene at the ascI- and sacI-interfaces in the binary vector 1.times.W1-luc-kan, a plasmid based on the binary vector pGPTV. In FIG. 7, the vector 4.times.Cis05-luc-kan is shown as an example. Corresponding vectors were created for all tested elements. The plasmid DNA of the binary vectors was isolated from E. Coli and transformed into the Agrobacterium tumefaciens strain GV3101 by a Gene Pulser.RTM. II. Electroporation System at a setting of 25 mF and 2.5 kV. The selection of recombinant A. tumefaciens-{2)clones occurred by using the antibiotic kanamycin (50 mg/l). The transformation of the sugar beets was effected according to Lindsey et al. (1991) with the use of the antibiotic kanamycin.
[0115] The transgenesis of the plants was tested by PCR. The use of the primers GTGGAGAGGCTATTCGGTA (SEQ ID NO: 36) and CCACCATGATATTCGGCAAG (SEQ ID NO: 37) led to the amplification of a 553-bp-sized DNA fragment from the nptlI-gene. The PCR was conducted using 10 ng genomic DNA, a primer concentration of 0.2 .mu.M at an annealing temperature of 55.degree. C. in a multicycler PTC-200 (MJ Research, Watertown, USA).
[0116] Ten to twenty independent transgenic lines were clonally increased in in vitro culture and infected with Cercospora beticola (Ahlburg isolate). After one, two, three, and four days, four plants/line were harvested and their luciferase activity was measured with the Promega Luciferase Assay System, 100 assays, cat. no. E1500 (LAR). The samples were housed in four volumes CCLR buffer (Cell Culture Lysis Reagent, 5.times.) and processed using a Heidolph (RZR 2020). The plant residue was removed by centrifugation for ten min at 4.degree. C. and 14000 rpm and the supernatant was used for the luciferase measurement. A 10-.mu.l sample was pipetted into a 5 ml test tube (Sarstedt, item no. 55.476) for measurement and 100 .mu.l Luciferase Assay Reagent (LAR; Promega, Mannheim, Germany) was added. After careful mixing, the luciferase activity was determined in a Sirius Luminometer (Berthold Detection System GmbH, Pforzheim, Germany).
[0117] Novel elements from groups 12 and 27 (see FIGS. 3D and F for group classification) were tested, among other things. The median was calculated of the measured luciferase activities of the approximately 10 to 20 independent transgenic lines with a chimeric promoter. The tests of the different promoters are summarized according to groups from which the elements originated. The results are shown in FIG. 8A. The direct comparison shows that the single sequences of a motif group have significantly differing activities despite the high homology of the motifs they contain. The nucleotides of the family motifs that border the core sequence can thus heavily influence the promoter strength and the background activity. The different sequences of a motif group are thus not identical in their function, but rather permit the development of chimeric promoters with quantitative differences in the regulation of the gene expression.
[0118] To test a further motif group, the element GG6_M1 (single sequence GG6_M1_S1) was stably transformed in sugar beet as specified above and tested for inducibility by Cercospora. Ten to twenty independent transgenic lines were clonally increased in vitro culture and infected with Cercospora beticola (Ahlburg isolate). After two, three, four, and seven days, four plants/line were harvested and their luciferase activity was measured with the Promega Luciferase Assay System, 100 assays, cat. no. E1500 (LAR). The results are shown in FIG. 9 and demonstrate a clear inducibility of the single sequence GG6_M1_S1 in sugar beet by Cercospora beticola. Furthermore, a histochemical analysis showed that the induction largely occurred in vascular tissue.
[0119] To investigate the influence of the W-box sequence in the single sequence Cis05 in stably transformed plants, the mutated derivatives Cis05mut1 and Cis05mut2 and the shortened single sequence sCis05 from the parsley tests were stably transformed in sugar beet. Construct creation and transformation were conducted as described above. Eighteen independent transgenic lines were clonally propagated in vitro and infected with Cercospora beticola (Ahlburg isolate). After one, two, three, and four days, four plants/line were harvested and their luciferase activity was measured as described above (FIG. 8B).
[0120] The mutation analysis in stably transformed plants shows that this W-box alone, i. e. with mutated Cis05 motif, conveys only a weak pathogen inducibility by Cercospora. The Cis05 motif alone (mutated W-box or shortened element without W-box) is, on the other hand, significantly inducible. The complete Cis05 single sequence with W-box is likewise inducible, but demonstrates an increased background activity relative to the derivatives without W-box. Thus the Cis05 motif alone in stably transformed sugar beet is equivalent or even superior to the combination with the W-box.
[0121] Another important feature of the inventive chimeric promoters is that the pathogen-induced activity is limited to the region of the infection site. This is shown in FIG. 14 using the example of the inventive promoter 4.times.Cis05. Said promoter was merged with the red fluorescing reporter gene and the construct thereby obtained stably transformed in sugar beet. The activity can be observed as red fluorescence under the laser scanning microscope. In FIG. 14 shows the local induction of the 4.times.Cis05 promoter around the penetration site of a Cercospora hyphae.
Enlarged Species-Wide Pathogen Inducibility
[0122] As a further example for broad applicability of the cis-regulatory elements in different plant species, the inducibility was demonstrated for Cis05 in transient experiments in the monocotyledonous plant wheat through the fungus Fusarium culmorum. Since the 35S minimal promoter in wheat conveys insufficient activity, the Cis05 elements had to be recloned. To that end, they were excised from the plasmids used for the parsley tests with the enzymes Eco31I and XbaI and cloned in the vector pubiTATARucII opened with Eco31I and BcuI (FIG. 10). Corresponding constructs were generated for Cis05 as well as for the mutated Cis05 single sequences Cis05mut1 and Cis05mut2 in which either the Cis05 motif or the W-box is mutated. These constructs were biolistically transformed in the primary leaves of the wheat variety "taifun", which leaves were infected with Fusarium graminearum, as well as in non-infected control leaves.
[0123] For the infection, Fusarium graminearum mycel with a specimen slide covered in fungal plate was scraped off, briefly ground in 200 ml water with an Ultraturrax, and suspended. Thereafter, 200 .mu.l 2% triton was added and the wheat primary leaves were stirred in the suspension for one minute. Subsequent thereto, the infected leaf pieces and non-infected control leaf pieces were placed on H.sub.2O agar plates and transiently transformed with a Bio-Rad particle gun according to manufacturer's instruction using 1100 psi rupture disks. A constitutive luciferase-expressing vector was used as a normalizing vector.
[0124] The wheat leaves were then incubated over night at 25.degree. C. To determine the luciferase activities, each of the leaves was ground up in a mortar in 1 ml PLB buffer with sea sand. After 20 minutes of centrifugation at 4.degree. C., the a 5 .mu.l sample of the supernatant with the released luciferase is mixed in 5 ml test tubes (Sarstedt, item no. 55.476) with 50 .mu.l LARII buffer (Promega, Mannheim, Germany). The buffer contains the substrate of the luciferase so that the activity of the normalizing vectors can be measured. This measured value is used for normalizing the different transformation efficiencies. The measurement is effected with two seconds pre-measurement time and ten seconds luciferase measurement time. Subsequent thereto, 50 .mu.l Stop & Glo buffer (Promega, Mannheim, Germany) is added and carefully mixed by drawing up. This buffer stops the luciferase activity and renders measurable the renilla-luciferase activity that corresponds to the activity of the Cis05 promoters. The measurement likewise occurs with two seconds pre-measurement time and ten seconds luciferase-measurement time.
[0125] The induced or non-induced activities of the 4.times.Cis05 promoter and of its mutated derivatives were measured in five biological repetitions (FIG. 11). Thereafter, the complete experiment was repeated once again. Both repetitions demonstrate that in wheat, the pathogen-induced activity stems from the Cis05 motif, while the W-box motif does not convey any activity induced by Fusarium graminearum.
[0126] Analysis of the pathogen- or wound-induced and tissue-specific activity of the elements Cis02, Cis05, Cis09, Cis12 or Cis13 in Arabidopsis.
[0127] The novel cis-regulatory elements should thereby distinguish themselves in that they can be induced in different plant species by different PAMPs and pathogens. Six promoters with tetramerized single sequences were stably transformed in Arabidopsis as an additional control for the activity of the chimeric promoters. The tetramerized elements Cis02, Cis05, Cis09, Cis12 or Cis13 were additionally cloned with 35S minimal promoter prior to the GUS reporter gene in the transformation vector pBIN-GUS. The final construct was transformed in agrobacteria. A floral-dip transformation (Clough and Bent, 1998) of Arabidopsis plants was conducted in the following to stably integrate the promoter-GUS constructs in the plant genome. The selection of transgenic plants was made using the kanamycin antibiotic.
[0128] Ten independent transformants were investigated for each element.
[0129] The activity of the GUS reporter gene and of the promoters can be made visible with the use of a GUS dye (Jefferson et al., 1987). The activation of the respective chimeric promoters leads to the expression of an enzyme (GUS) that creates a blue dye. The coloration thus shows the activity of the respective chimeric promoters.
[0130] To test the pathogen inducibility of the promoters, the transgenic Arabidopsis plants were infected with the compatible pathogen Hyaloperonospora arabidopsidis. Five days after infection, the activity of the promoters was proven by a GUS dye. Individual leaves were additionally wounded by cutting with scissors to test the wound inducibility of the promoters (FIG. 16).
[0131] The element Cis02 demonstrated favorable pathogen inducibility and only a minimal wound inducibility. The element Cis05 demonstrated a generally strong activity and is likewise strongly induced by H. Arabidopsidis. The element Cis09 demonstrated a favorable induction of the promoter subsequent to infection and hardly any undesired activity after wounding. Like element Cis09, the element Cis12 demonstrated in all examined lines hardly any undesired activity after wounding, while an obvious induction through the pathogen H. arabidopsidis was observed. The elements Cis12 and Cis09 share a common family motif. The element Cis13 is also heavily induced through infection with H. arabidopsidis. An obvious wound inducibility is not, however, observed. FIG. 16 shows the exemplary GUS dyeing of transgenic A. thaliana plants that express the GUS reporter gene under the control of a chimeric promoter with 4.times. Cis05.
Combinatorials of the Cis-Regulatory Elements
[0132] Chimeric combinatorial promoters that are composed of different cis-regulatory elements can demonstrate a greater specificity and/or activity than the single elements present therein through a synergistic interaction or through integration of different signaling pathways (a promoter would be conceivable that would have to perceive and integrate signals "EF-Tu" AND "flg22" in order to be activated) (Rushton et al., 2002). To identify optimal combinations of the novel cis-regulatory elements for chimeric combinatorial promoters, chimeric combinatorial promoters, including all possible 2.times.2 combinations of the herein-described cis-regulatory elements Cis02, Cis05, Cis12, Cis13 and 30I-8_M1_S2 and the already published elements D-box (Rushton et al., 2002), S-box (Kirsch et al., 2000) and Gst1-box (Strittmatter et al., 1996), were generated using standard DNA cloning procedures. The approach taken followed that for the tetramerization of the single sequences described, only not the two identical dimers (e.g. 2.times.30I-8_M1_S2 and 2.times.30I-8_M1_S2), but rather two different dimers (e.g. 2.times.Cis05 and 2.times.30I-8_M1_S2) are cloned together. The thus-resulting chimeric combinatorial promoters were tested in the transient expression systems in parsley for their inducibility through the PAMP PEP25. The results (absolute non-induced and induced activity of the chimeric combinatorial promoters and the induction factor) are reproduced in FIG. 12. Chimeric combinatorial promoters with especially stronger and more specific inducibility could be identified, the induction factor of and activity of which are greater than in chimeric promoters that are constructed from only repetitions of the individual cis-regulatory elements (FIG. 13). The chimeric combinatorial promoters having the greatest induction factors and the strongest activities are summarized in Table 3.
TABLE-US-00004 TABLE 3 Combinations of pathogen-inducible single sequences having the greatest induction factors and induced activities. Standard Coefficient Average deviation of variation Induction Combinations with the greatest induction factor 2xCis05-2xCis05 - 3.23 2.06 0.64 149.58 2xCis05-2xCis05 + 482.53 398.05 0.82 2xCis05-2xS - 2.00 1.51 0.76 115.63 2xCis05-2xS + 231.82 48.39 0.21 2xCis05-2x30I8b - 2.47 1.42 0.58 123.16 2xCis05-2x30I8b + 304.21 45.38 0.15 2xCis13-2xCis02 - 2.50 0.50 0.20 124.53 2xCis13-2xCis02 + 311.48 37.76 0.12 2xCis13-2xCis05 - 2.33 0.41 0.18 182.59 2xCis13-2xCis05 + 425.23 258.29 0.61 2xCis13-2xS - 4.55 0.65 0.14 139.85 2xCis13-2xS + 636.85 364.80 0.57 2xS-2xCis02 - 5.28 3.36 0.64 106.32 2xS-2xCis02 + 561.76 440.81 0.78 2xS-2xS - 2.76 0.49 0.18 138.80 2xS-2xS + 382.69 92.20 0.24 Gst1-2xS - 2.41 0.87 0.36 258.83 Gst1-2xS + 624.18 274.19 0.44 2x30I8b-2xCis05 - 0.90 0.30 0.34 135.12 2x30I8b-2xCis05 + 121.20 88.90 0.73 Combinations with the greatest induction factor (induced) 2xD-2xCis05 - 84.57 49.56 0.59 17.88 2xD-2xCis05 + 1511.90 580.33 0.38 2xD-2xS - 23.96 17.66 0.74 57.72 2xD-2xS + 1382.94 1180.84 0.85 2xD-Gst1 - 19.25 20.18 1.05 62.10 2xD-Gst1 + 1195.21 486.92 0.41 2xS-2xCis12 - 36.84 19.34 0.53 29.05 2xS-2xCis12 + 1070.08 1043.46 0.98 2xS-2xD - 22.13 23.38 1.06 65.56 2xS-2xD + 1450.50 1085.69 0.75
REFERENCES
[0133] Bailey, T. L., C. Elkan (1994). "Fitting a mixture model by expectation maximization to discover motifs in biopolymers." Proc Int Conf Intell Syst Mol Biol. 2: 28-36. [0134] Bulow, L., M. Schindler, et al. (2007). "PathoPlant: a platform for microarray expression data to analyze co-regulated genes involved in plant defense responses." Nucleic Acids Res 35(Database issue): D841-845. [0135] Che, D., S. Jensen, et al. (2005). "BEST: binding-site estimation suite of tools." Bioinformatics 21(12): 2909-2911. [0136] Clough, S. J. and Bent, A. F. (1998). " Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana." Plant J. 16(6):735-43. [0137] Eulgem, T., P. J. Rushton, et al. (2000). "The WRKY superfamily of plant transcription factors." Trends Plant Sci 5(5): 199-206. [0138] Gorlich, D. and I. W. Mattaj (1996). "Nucleocytoplasmic transport." Science 271(5255): 1513-1518. [0139] Gurr, S. J. and P. J. Rushton (2005). "Engineering plants with increased disease resistance: how are we going to express it?" Trends Biotechnol 23(6): 283-290. [0140] Hahlbrock, K., D. Scheel, et al. (1995). "Oligopeptide elicitor-mediated defense gene activation in cultured parsley cells." Proc Natl Acad Sci USA 92(10): 4150-4157. [0141] Hicks, G. R., H. M. Smith, et al. (1995). "Three classes of nuclear import signals bind to plant nuclei." Plant Physiol 107(4): 1055-1058. [0142] Himmelbach, A., L. Liu et al. (2010). "Promoters of the Barley Germin-Like GER4 Gene Cluster Enable Strong Transgene Expression in Response to Pathogen Attack" The Plant Cell 22(3): 937-952. [0143] Howles, P., G. Lawrence, et al. (2005). "Autoactive Alleles of the Flax L6 Rust Resistance Gene Induce Non-Race-Specific Rust Resistance Associated with the Hypersensitive Response." Molecular Plant-Microbe Interactions 18(6): 570-582. [0144] Humphry, M., P. Bednarek, et al. (2010). "A regulon conserved in monocot and dicot plants defines a functional module in antifungal plant immunity." Proc Natl Acad Sci USA 107(50): 21896-21901. [0145] Jefferson, R. A., Kavanagh, T. A & Bevan, M. W. (1987). "GUS fusions, 8-glucuronidase as a sensitive and versatile gene fusion marker in higher plants." EMBO J 6(13): 3901-3907. [0146] Kirsch, C., M. Takamiya-Wik, et al. (2000) "A novel regulatory element involved in rapid activation of parsley ELI7 gene family members by fungal elicitor or pathogen infection." Mol Plant Pathol. 1(4): 243-51. [0147] Kirsch, C., E. Logemann, et al. (2001) "A highly specific pathogen-responsive promoter element from the immediate-early activated CMPG1 gene in Petroselinum crispum." Plant J. 26(2): 217-27. [0148] Kovalchuk, N., M. Li, et al. (2010). "Defensin promoters as potential tools for engineering disease resistance in cereal grains." Plant Biotechnol J 8(1): 47-64. [0149] Lebel, E., P. Heifetz, et al. (1998). "Functional analysis of regulatory sequences controlling PR-1 gene expression in Arabidopsis." Plant J 16(2): 223-233. [0150] Liu, X., D. L. Brutlag, et al. (2001). "BioProspector: discovering conserved DNA motifs in upstream regulatory regions of co-expressed genes." Pac Symp Biocomput: 127-138. [0151] Morris, K., S. A. MacKerness, et al. (2000). "Salicylic acid has a role in regulating gene expression during leaf senescence." Plant J 23(5): 677-685. [0152] Rushton, P. J., A. Reinstadler, et al. (2002). "Synthetic plant promoters containing defined regulatory elements provide novel insights into pathogen- and wound-induced signaling." Plant Cell 14(4): 749-762. [0153] Rushton, P. J., J. T. Torres, et al. (1996). "Interaction of elicitor-induced DNA-binding proteins with elicitor response elements in the promoters of parsley PR1 genes." EMBO J 15(20): 5690-5700. [0154] Schatz, G. and B. Dobberstein (1996). "Common principles of protein translocation across membranes." Science 271(5255): 1519-1526. [0155] Schumacher, S. B., O. Van den Hauwe, et al. (2003). "Development of a dual luciferase reporter screening assay for the detection of synthetic glucocorticoids in animal tissues." Analyst 128(12): 1406-1412. [0156] Strittmatter, G., G. Gheysen, et al. (1996). "Infections with various types of organisms stimulate transcription from a short promoter fragment of the potato gst1 gene." Mol Plant Microbe Interact 9(1): 68-73. [0157] Venter, M. (2007). "Synthetic promoters: genetic control through cis engineering." Trends Plant Sci 12(3): 118-124. [0158] Verner, K. and G. Schatz (1988). "Protein translocation across membranes." Science 241(4871): 1307-1313. [0159] Wan, J., X. C. Zhang, et al. (2008). "A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis." Plant Cell 20(2): 471-481. [0160] Zipfel, C., G. Kunze, et al. (2006). "Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation." Cell 125(4): 749-760. [0161] Zipfel, C., S. Robatzek, et al. (2004). "Bacterial disease resistance in Arabidopsis through flagellin perception." Nature 428(6984): 764-767. [0162] WO 00/29592 (Max-Planck-Gesellschaft zur Forderung der Wissenschaften e.V.). Chimeric promoters capable of mediating gene expression in plants upon pathogen infection and uses thereof. [0163] WO 02/50293 (Max-Planck-Gesellschaft zur Forderung der Wissenschaften e.V.). Pflanzen mit verbesserter Widerstandskraft. [0164] WO 03/00898 (Syngenta Participations AG). Plant genes involved in defense against pathogens. [0165] WO 07/147395 (KWS SAAT AG). Pathogen induzierbarer synthetischer Promotor [0166] WO 10/079430 (U. Bonas et al.). Modular DNA-binding domains and methods of use. [0167] WO 11/072246 (Regents of the University of Minnesota; Iowa State University Research Foundation Inc.). TAL effector-mediated DNA modification.
Sequence CWU 1
1
145118DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotidemodified_base(1)..(2)a, c, t or
gmisc_feature(1)..(2)n is a, c, g, or tmodified_base(6)..(7)a, c, t
or gmisc_feature(6)..(7)n is a, c, g, or tmisc_feature(9)..(14)Core
sequence motifmodified_base(15)..(15)a, c, t or
gmisc_feature(15)..(15)n is a, c, g, or t 1nnhkdnnvaa agtmndhy
18216DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotidemodified_base(2)..(2)a, c, t or
gmisc_feature(2)..(2)n is a, c, g, or tmodified_base(6)..(6)a, c, t
or gmisc_feature(6)..(6)n is a, c, g, or tmisc_feature(7)..(12)Core
sequence motifmodified_base(15)..(15)a, c, t or
gmisc_feature(15)..(15)n is a, c, g, or t 2ynamcnaaac cawwny
16313DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotidemodified_base(2)..(2)a, c, t or
gmisc_feature(2)..(2)n is a, c, g, or tmisc_feature(5)..(10)Core
sequence motif 3wnrmscaaam smw 13413DNAArtificial
SequenceDescription of Artificial Sequence Synthetic
oligonucleotidemodified_base(1)..(2)a, c, t or
gmisc_feature(1)..(2)n is a, c, g, or tmisc_feature(5)..(9)Core
sequence motifmodified_base(11)..(11)a, c, t or
gmisc_feature(11)..(11)n is a, c, g, or t 4nnmsacrcgy nwm
13514DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotidemisc_feature(2)..(9)Core sequence motif
5asktgkactw kgwm 14615DNAArtificial SequenceDescription of
Artificial Sequence Synthetic
oligonucleotidemodified_base(2)..(2)a, c, t or
gmisc_feature(2)..(2)n is a, c, g, or tmisc_feature(6)..(12)Core
sequence motifmodified_base(15)..(15)a, c, t or
gmisc_feature(15)..(15)n is a, c, g, or t 6knwymmrtsa ckwmn
15725DNAArabidopsis thalianamisc_feature(10)..(16)Core sequence
7acaacagacg acttttcata attca 25825DNAArabidopsis
thalianamisc_feature(10)..(16)Core sequence 8ctatatgaca aaagtcaaac
ataaa 25927DNAArabidopsis thalianamisc_feature(7)..(21)Core
sequence 9tgttcacttt gaaaagtatt ctttgag 271026DNAArabidopsis
thalianamisc_feature(7)..(20)Core sequence 10cgatcagact tttctacgca
agagaa 261135DNAArabidopsis thalianamisc_feature(12)..(24)Core
sequence 11taatttctct tgcgtagaaa agtctgatcg ggaag
351235DNAArabidopsis thalianamisc_feature(15)..(21)Core sequence
12tcgttcttca gtcaaaaagt caaactatct ctctc 351330DNAArabidopsis
thalianamisc_feature(12)..(18)Core sequence 13gagcgtgaat tgactttgac
caaaaccaaa 301438DNAArabidopsis thalianamisc_feature(14)..(20)Core
sequence 14ggtcagcatg ttggactttc caaattcatt gaccaaag
381539DNAArabidopsis thalianamisc_feature(20)..(29)Core sequence
15aaaataaaca gctacttgac gaaaagtcaa accaaattc 391636DNAArabidopsis
thalianamisc_feature(14)..(23)Core sequence 16gttttgactt ttgacctaaa
ccatttccat gtagaa 361736DNAArabidopsis
thalianamisc_feature(13)..(24)Core sequence 17ccgtcttagt ttaccgaaac
caaagtggct ttttct 361835DNAArabidopsis
thalianamisc_feature(11)..(25)Core sequence 18cgtaataatg gtttggtttg
gtttgatcaa gtctt 351925DNAArabidopsis
thalianamisc_feature(8)..(18)Core sequence 19gacttttgac ctaaaccatt
tccat 252026DNAArabidopsis thalianamisc_feature(9)..(17)Core
sequence 20caacacaaaa cgcaaacgca gacctc 262135DNAArabidopsis
thalianamisc_feature(14)..(22)Core sequence 21aattgacaaa agacacgcaa
acgattccaa cgacc 352235DNAArabidopsis
thalianamisc_feature(14)..(22)Core sequence 22aaataattat ttatggtttg
gtcatttggt caaat 352335DNAArabidopsis
thalianamisc_feature(15)..(21)Core sequence 23agtcaaaacg tagaccaaaa
caaaaacatg taact 352438DNAArabidopsis
thalianamisc_feature(12)..(19)Core sequence 24tgcacacaca cacacgtgta
ctaggtcaaa ccaaacgt 382541DNAArabidopsis
thalianamisc_feature(20)..(25)Core sequence 25caaaaagtca acacatacga
cgcgtttcca ttgactaaat a 412625DNAArabidopsis
thalianamisc_feature(9)..(17)Core sequence 26tctcatctct cgacacgcaa
cttcc 252725DNAArabidopsis thalianamisc_feature(7)..(19)Core
sequence 27cacacacgtg tactaggtca aacca 252835DNAArabidopsis
thalianamisc_feature(14)..(22)Core sequence 28aggacttttc accagttgga
ctttgaagcc accaa 352934DNAArabidopsis
thalianamisc_feature(11)..(19)Core sequence 29aagtctaaat ctttgacccc
aaaaaagaga gcaa 343035DNAArabidopsis
thalianamisc_feature(15)..(21)Core sequence 30tgttgagtcg tttacgtcac
gtcgagaatt ttctc 353135DNAArabidopsis
thalianamisc_feature(15)..(21)Core sequence 31tgtcattatt aatacgtgac
gaaactgtag ctctg 353236DNAArabidopsis
thalianamisc_feature(11)..(25)Core sequence 32ttacgtgtca agaagtgatt
ggagaggaca ctctac 363335DNAArabidopsis
thalianamisc_feature(15)..(21)Core sequence 33ccatacaata taaaccacca
aaccataacc acaaa 353426DNAArabidopsis
thalianamisc_feature(9)..(18)Core sequence 34caatctactc gtctcttctc
ttacat 263510DNAArabidopsis thaliana 35tcgtctcttc
103619DNAArtificial SequenceDescription of Artificial Sequence
Synthetic primer 36gtggagaggc tattcggta 193720DNAArtificial
SequenceDescription of Artificial Sequence Synthetic primer
37ccaccatgat attcggcaag 2038182DNAOryza sativa 38tcctctctac
tcctcagcat ctatataagg gccttgcagg cagtggctct cacaccaacg 60aaacaaggag
agactcagag aggaggcgtg ttggttagct cattggcagc agcacacaca
120aaccacatct cctatatata gctcattttt agcttttgga attgagagag
gttttgagag 180aa 18239149DNATriticum aestivum 39atctgcaggc
gctgcctatt taatcccttc ccctccctcc attcccctcc aagaagagcc 60acagcttcat
ctgcagctac agctcctctt cgtcttcgac acacaagtat tttttcagga
120caaagatcaa tccagataca catacacct 149401133DNAZea
maysIntron(129)..(1133) 40cttcctcgcc cgccgtaata aatagacacc
ccctccacac cctctttccc caacctcgtg 60ttgttcggag cgcacacaca cacaaccaga
tctcccccaa atccacccgt cggcacctcc 120gcttcaaggt acgccgctcg
tcctcccccc ccctctctac cttctctaga tcggcgttcc 180ggtccatggt
tagggcccgg tagttctact tctgttcatg tttgtgttag atccgtgttt
240gtgttagatc cgtgctgcta gcgttcgtac acggatgcga cctgtacgtc
agacacgttc 300tgattgctaa cttgccagtg tttctctttg gggaatcctg
ggatggctct agccgttccg 360cagacgggat cgatttcatg attttttttg
tttcgttgca tagggtttgg tttgcccttt 420tcctttattt caatatatgc
cgtgcacttg tttgtcgggt catcttttca tgcttttttt 480tgtcttggtt
gtgatgatgt ggtctggttg ggcggtcgtt ctagatcgga gtagaattct
540gtttcaaact acctggtgga tttattaatt ttggatctgt atgtgtgtgc
catacatatt 600catagttacg aattgaagat gatggatgga aatatcgatc
taggataggt atacatgttg 660atgcgggttt tactgatgca tatacagaga
tgcttttgtt cgcttggttg tgatgatgtg 720gtgtggttgg gcggtcgttc
attcgttcta gatcggagta gaatactgtt tcaaactacc 780tggtgtattt
attaattttg gaactgtatg tgtgtgtcat acatcttcat agttacgagt
840ttaagatgga tggaaatatc gatctaggat aggtatacat gttgatgtgg
gttttactga 900tgcatataca tgatggcata tgcagcatct attcatatgc
tctaaccttg agtacctatc 960tattataata aacaagtatg ttttataatt
attttgatct tgatatactt ggatgatggc 1020atatgcagca gctatatgtg
gattttttta gccctgcctt catacgctat ttatttgctt 1080ggtactgttt
cttttgtcga tgctcaccct gttgtttggt gttacttctg cag
11334118DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotidemodified_base(2)..(2)a, c, t or
gmisc_feature(2)..(2)n is a, c, g, or tmodified_base(4)..(6)a, c, t
or gmisc_feature(4)..(6)n is a, c, g, or tmisc_feature(7)..(12)Core
sequence motifmodified_base(13)..(15)a, c, t or
gmisc_feature(13)..(15)n is a, c, g, or tmodified_base(17)..(17)a,
c, t or gmisc_feature(17)..(17)n is a, c, g, or t 41snsnnnwwkg
wcnnnsnm 184229DNAArabidopsis thalianamisc_feature(5)..(25)Core
sequence motif 42ctcaaaggcc agaattgacg cagccgttt
294328DNAArabidopsis thalianamisc_feature(7)..(22)Core sequence
motif 43ccttggccca gtccttggtc gtcgtatc 284411DNAArabidopsis
thalianamisc_feature(5)..(11)Core sequence motif 44gttggacttt c
114528DNAArabidopsis thaliana 45tctactagag gcccattagg accggcat
284635DNAArabidopsis thaliana 46aagacaagtt gagagagacg agaccaatca
caaca 354735DNAArabidopsis thaliana 47atccaacatc tcggaccgga
tcaatgattt atcat 354825DNAArabidopsis thaliana 48tcatcaatgt
gacataagca aagct 254926DNAArabidopsis thaliana 49tttgatacgg
ttacggttaa ttaacg 265036DNAArabidopsis thaliana 50aagattctca
tccaaccgaa acgactcttt cgtttt 365136DNAArabidopsis thaliana
51agattctcat ccaaccgaaa cgactctttc gttttt 365236DNAArabidopsis
thaliana 52tcttcttcat tttaccaaca ccacttgcac acacac
365335DNAArabidopsis thaliana 53agttgaatta gttcggttcg gttcggttga
tattg 355424DNAArabidopsis thaliana 54cgataaactt gcgaaaccct aaaa
245525DNAArabidopsis thaliana 55tattggaagt ttggggcaac atcac
255635DNAArabidopsis thaliana 56ttcaacccta taaaccaaaa caaataacag
aatgc 355736DNAArabidopsis thaliana 57cgcctcaatc atgaaaacga
atcctctgta gtagtg 365835DNAArabidopsis thaliana 58ttaccgacac
gtaaccaaaa ctcaccgaac accgt 355935DNAArabidopsis thaliana
59gtttcgaacg ggaaccaaac cataatatgc gatgc 356035DNAArabidopsis
thaliana 60acactattgg tcttggtttg gtttatatgc acgac
356136DNAArabidopsis thaliana 61gaaaacgatg ggttccaaaa ctgtcgctaa
taaact 366236DNAArabidopsis thaliana 62tctccactcg ttgtgatttg
gtctgcaaga aaacta 366336DNAArabidopsis thaliana 63acgttttgaa
atattgtttt ggatggagat tttttc 366436DNAArabidopsis thaliana
64atttttcatt tcgcccaaaa caattatcct aacgtt 366536DNAArabidopsis
thaliana 65ttttataaca ctaccaaaac caataagccc tttcgt
366635DNAArabidopsis thaliana 66tcatcaaacc aatcggtttg gtcctaaaga
taatt 356735DNAArabidopsis thaliana 67gtcaatatac acagccaccg
aacaaattac tctat 356826DNAArabidopsis thaliana 68aagcgatgtt
tacgaacccc aaaatc 266925DNAArabidopsis thaliana 69agatttgttc
gagaaccttg agaaa 257025DNAArabidopsis thaliana 70ttgctacttc
gagaacattg gtcaa 257126DNAArabidopsis thaliana 71ttagaagtgg
ctcgagtgtt ctactt 267226DNAArabidopsis thaliana 72aagaaagaca
atcgagccta gaaatt 267336DNAArabidopsis thaliana 73aataatgttc
aacgttggtg gtggtactca agatgg 367436DNAArabidopsis thaliana
74tcaaatacag gcaaccaaga ctcgagatcc tcatcg 367525DNAArabidopsis
thaliana 75agaaaaatat tgggcctact gggaa 257636DNAArabidopsis
thaliana 76agattcctga agtgaggtcc accctaaaat ccattt
367735DNAArabidopsis thaliana 77ggtttagtca aagtaaacaa gactttgact
gttca 357836DNAArabidopsis thaliana 78tgaacttaat cactgtcatt
gttttcgtaa caattt 367926DNAArabidopsis thaliana 79gaaaaatgtg
tgtgtttgtg ttaatt 268026DNAArabidopsis thaliana 80atagttccca
aacggacacg aacaca 268126DNAArabidopsis thaliana 81gcaaactaac
gccggcggcc gtcttg 268225DNAArabidopsis thaliana 82tagctgttga
aatttccaag aaaat 258325DNAArabidopsis thaliana 83tttttctcgt
ccccatcctc tatcc 258426DNAArabidopsis thaliana 84taggttcctg
ccctctccgt tcctcc 268526DNAArabidopsis thaliana 85tcgaaaccaa
ccctctccct tataaa 268626DNAArabidopsis thaliana 86gaaaccaacc
ctctccctta taaata 268736DNAArabidopsis thaliana 87tgttttgttt
ccaccgtctc tcccgtgtcc tctctc 368824PRTArtificial
SequenceDescription of Artificial Sequence Synthetic peptide 88Val
Thr Ala Gly Ala Glu Val Trp Asn Gln Pro Val Arg Gly Phe Lys1 5 10
15Val Tyr Glu Gln Thr Glu Met Thr 208913DNAArabidopsis thaliana
89ctcgacacgc aac 139013DNAArabidopsis thaliana 90acacacacgt gta
139113DNAArabidopsis thaliana 91tacgacgcgt ttc 139215DNAArabidopsis
thaliana 92gtttacgtca cgtcg 159315DNAArabidopsis thaliana
93taatacgtga cgaaa 159415DNAArabidopsis thaliana 94tctccaatca cttct
159516DNAArabidopsis thaliana 95tgacctaaac catttc
169616DNAArabidopsis thaliana 96ttaccgaaac caaagt
169716DNAArabidopsis thaliana 97caaaccaaac caaacc
169813DNAArabidopsis thaliana 98aaacgcaaac gca 139913DNAArabidopsis
thaliana 99acacgcaaac gat 1310013DNAArabidopsis thaliana
100tagaccaaaa caa 1310113DNAArabidopsis thaliana 101atgaccaaac cat
1310214DNAArabidopsis thaliana 102acgtgtacta ggtc
1410314DNAArabidopsis thaliana 103agttggactt tgaa
1410418DNAArabidopsis thaliana 104aattatgaaa agtcgtct
1810518DNAArabidopsis thaliana 105atatgacaaa agtcaaac
1810618DNAArabidopsis thaliana 106tgcgtagaaa agtctgat
1810718DNAArabidopsis thaliana 107cactttgaaa agtattct
1810818DNAArabidopsis thaliana 108tcagtcaaaa agtcaaac
1810918DNAArabidopsis thaliana 109cttgacgaaa agtcaaac
1811018DNAArabidopsis thaliana 110ttttggtcaa agtcaatt
1811118DNAArabidopsis thaliana 111gaatttggaa agtccaac
1811221DNAArabidopsis thaliana 112cacgtgtact aggtcaaacc a
2111321DNAArabidopsis thaliana 113tttcaccagt tggactttga a
2111421DNAArabidopsis thaliana 114aaggccagaa ttgacgcagc c
2111521DNAArabidopsis thaliana 115gcccagtcct tggtcgtcgt a
2111644DNAArabidopsis thaliana 116aactggtcag catgttggac tttccaaatt
cattgaccaa agac 4411744DNAArabidopsis thaliana 117aactggtcag
catgttggtc gttccaaatt cattgaccaa agac 4411844DNAArabidopsis
thaliana 118aactggtcag catgttggac tttccaaatt catttcccaa agac
4411944DNAArabidopsis thaliana 119aactggtcag catgttggtc gttccaaatt
catttcccaa agac 4412011DNAArabidopsis thaliana 120gttggtcgtt c
1112111DNAArabidopsis thaliana 121gttcgacttt c
1112211DNAArabidopsis thaliana 122gtcggacttt c
1112311DNAArabidopsis thaliana
123gttggactgc c 1112411DNAArabidopsis thaliana 124gttggacttt a
1112525DNAArabidopsis thaliana 125ctatatgaca aaagagaaac ataaa
2512625DNAArabidopsis thaliana 126ctatatgaca ttagtcaaac ataaa
2512725DNAArabidopsis thaliana 127ctatatctca aaagtcaaac ataaa
2512826DNAArabidopsis thaliana 128caacacaaaa cgctttcgca gacctc
2612926DNAArabidopsis thaliana 129caacacaaat gccaaacgca gacctc
2613026DNAArabidopsis thaliana 130caacacaaaa cgcaaaccgt gacctc
2613129DNAArabidopsis thaliana 131tgagtcgttt acgtcacgtc gagaatttt
2913229DNAArabidopsis thaliana 132tgagtcgttt acgctgtgtc gagaatttt
2913329DNAArabidopsis thaliana 133tgagtcgttg ccgtcacgtc gagaatttt
2913429DNAArabidopsis thaliana 134tgagtcgttt acgtcacggt gagaatttt
2913529DNAArabidopsis thaliana 135acttttcacc agttggactt tgaagccac
2913629DNAArabidopsis thaliana 136acttttcacc agtcaggtct tgaagccac
2913729DNAArabidopsis thaliana 137acttttcacc ggttggactt taaagccac
291387DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotide 138vaaagtm 71396DNAArtificial
SequenceDescription of Artificial Sequence Synthetic
oligonucleotide 139aaacca 61406DNAArtificial SequenceDescription of
Artificial Sequence Synthetic oligonucleotide 140scaaam
61415DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotide 141acrcg 51428DNAArtificial
SequenceDescription of Artificial Sequence Synthetic
oligonucleotide 142sktgkact 81437DNAArtificial SequenceDescription
of Artificial Sequence Synthetic oligonucleotide 143mrtsack
71447DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotide 144ccaccaa 71456DNAArtificial
SequenceDescription of Artificial Sequence Synthetic
oligonucleotide 145wwkgwc 6
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
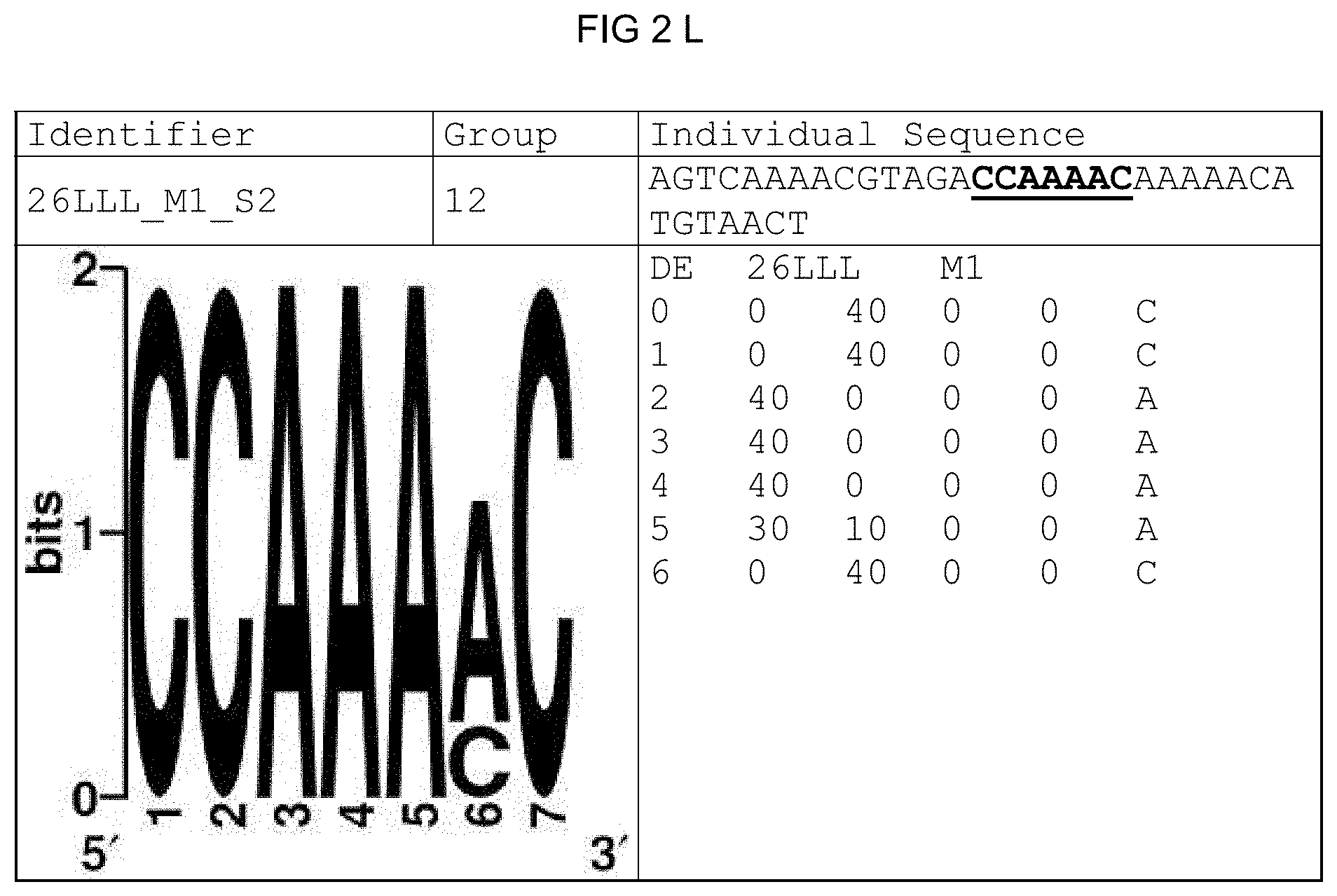

D00014

D00015

D00016

D00017

D00018
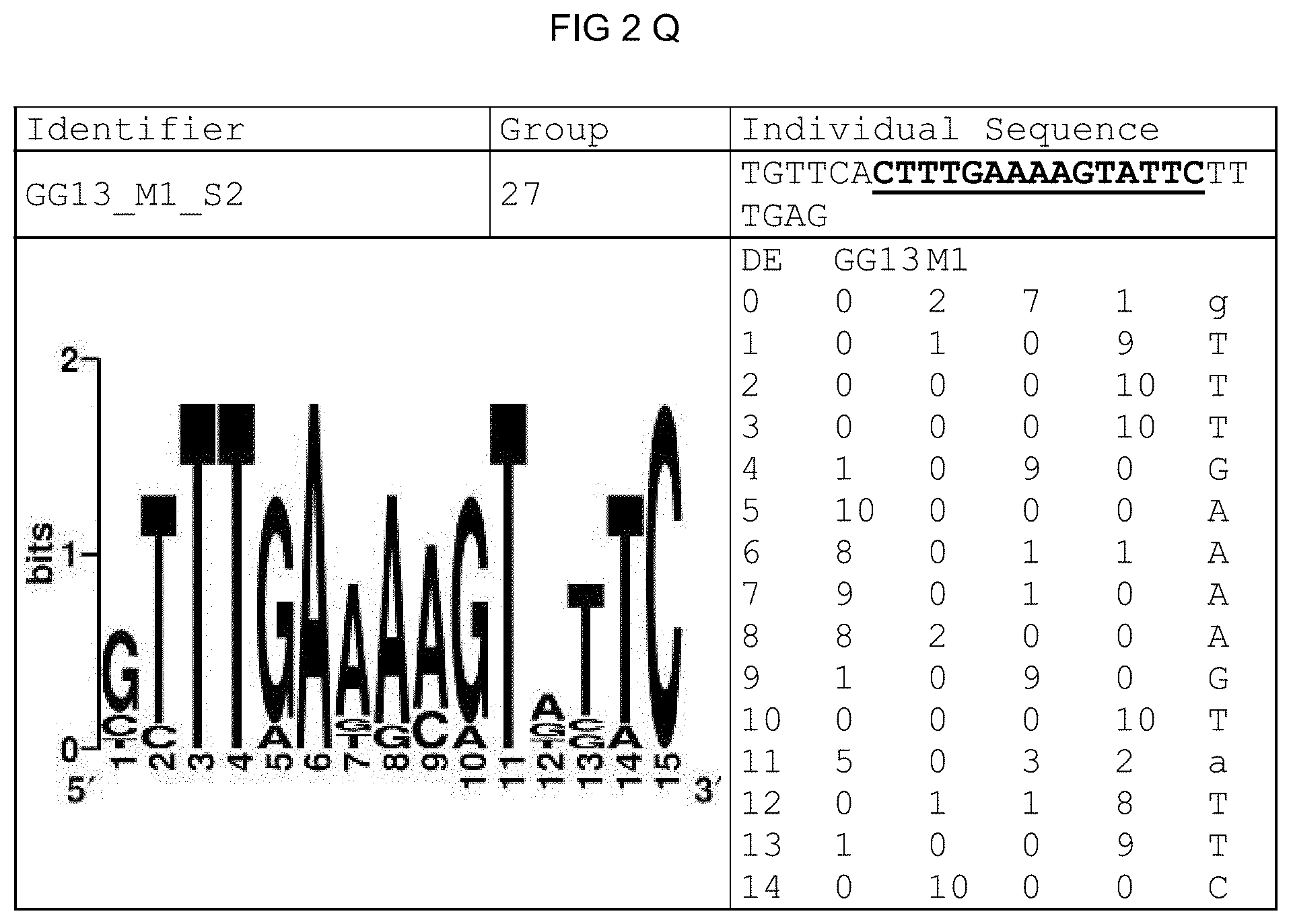
D00019

D00020

D00021

D00022
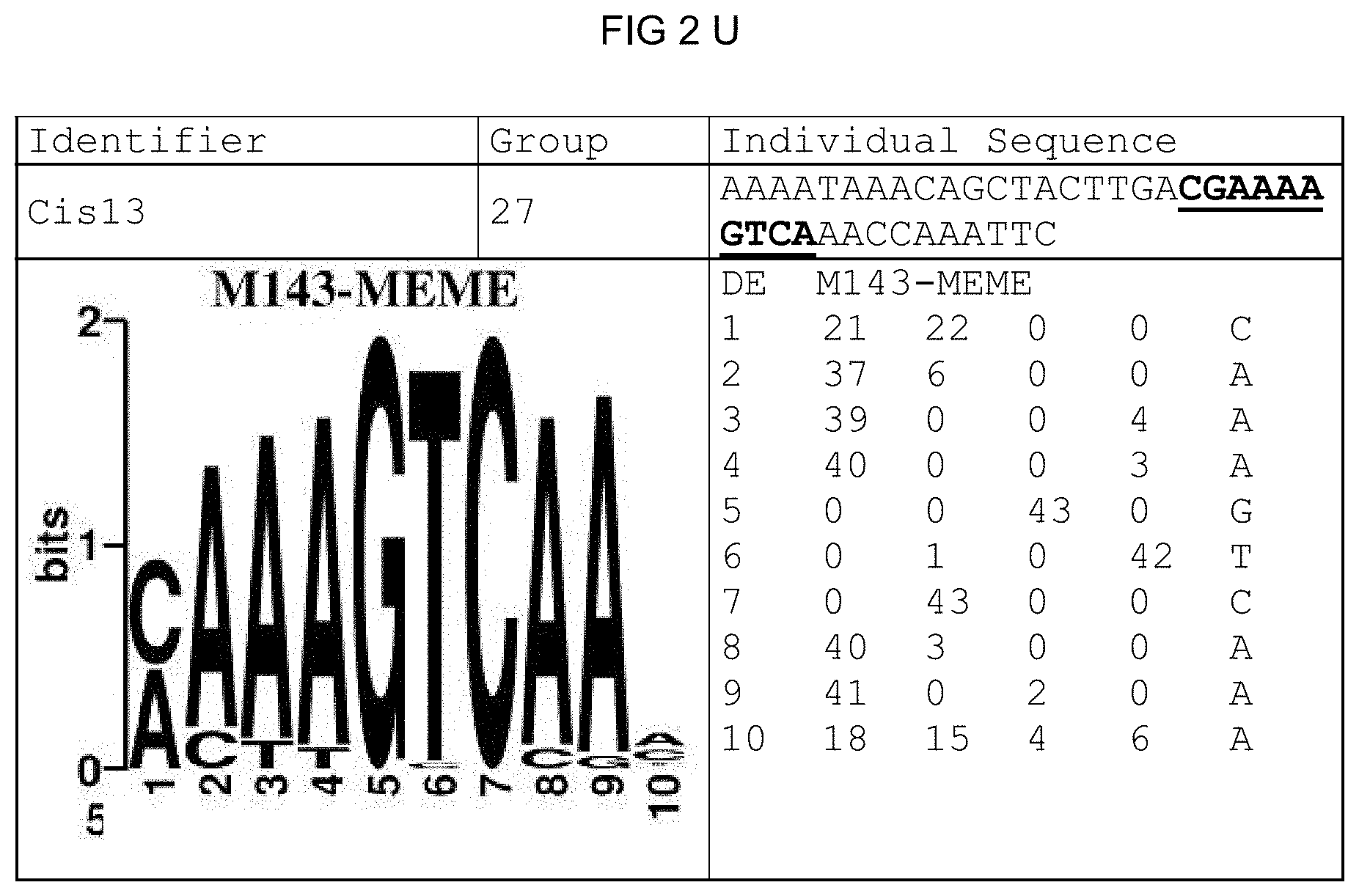
D00023

D00024

D00025

D00026

D00027
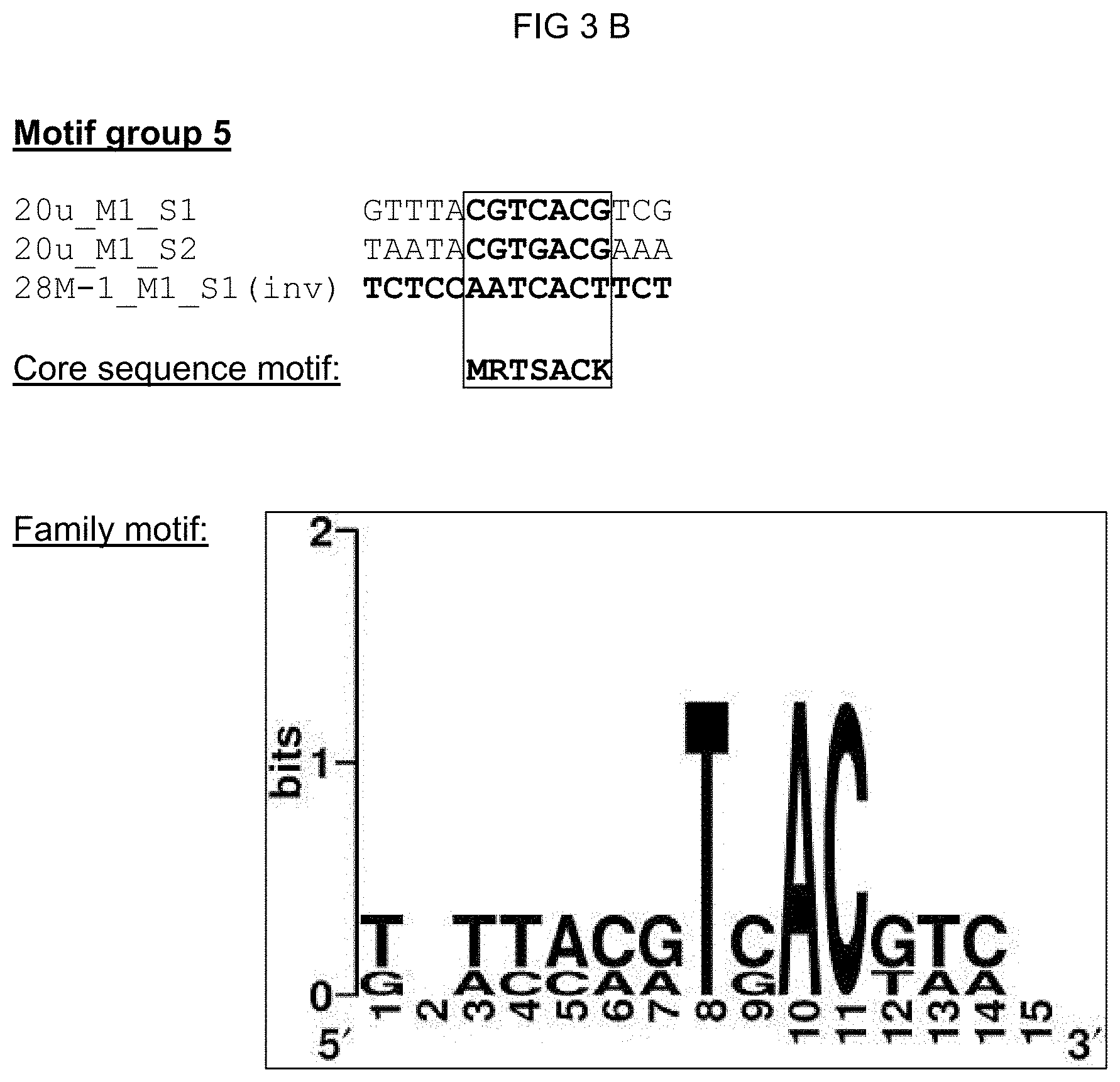
D00028
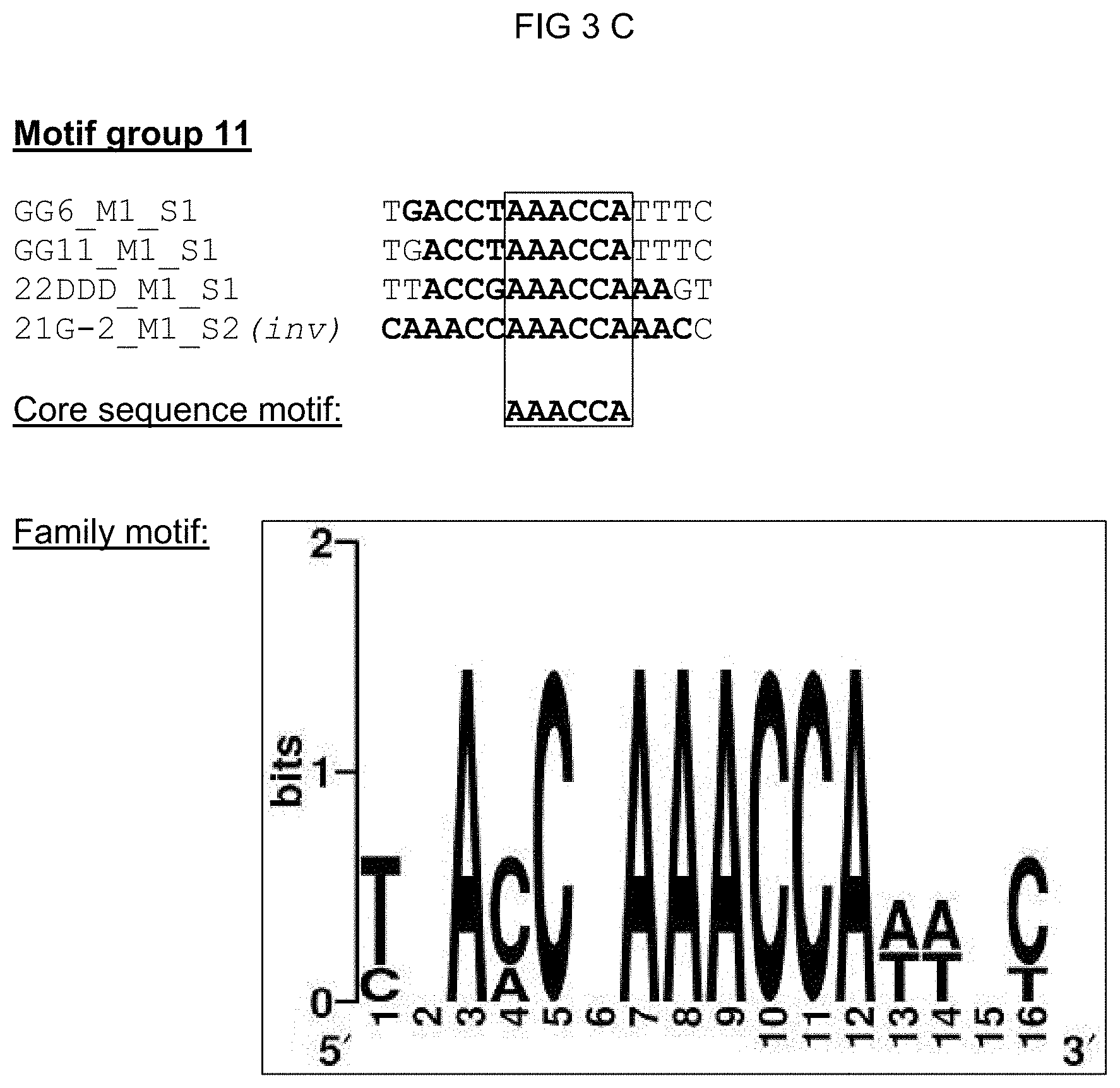

D00029
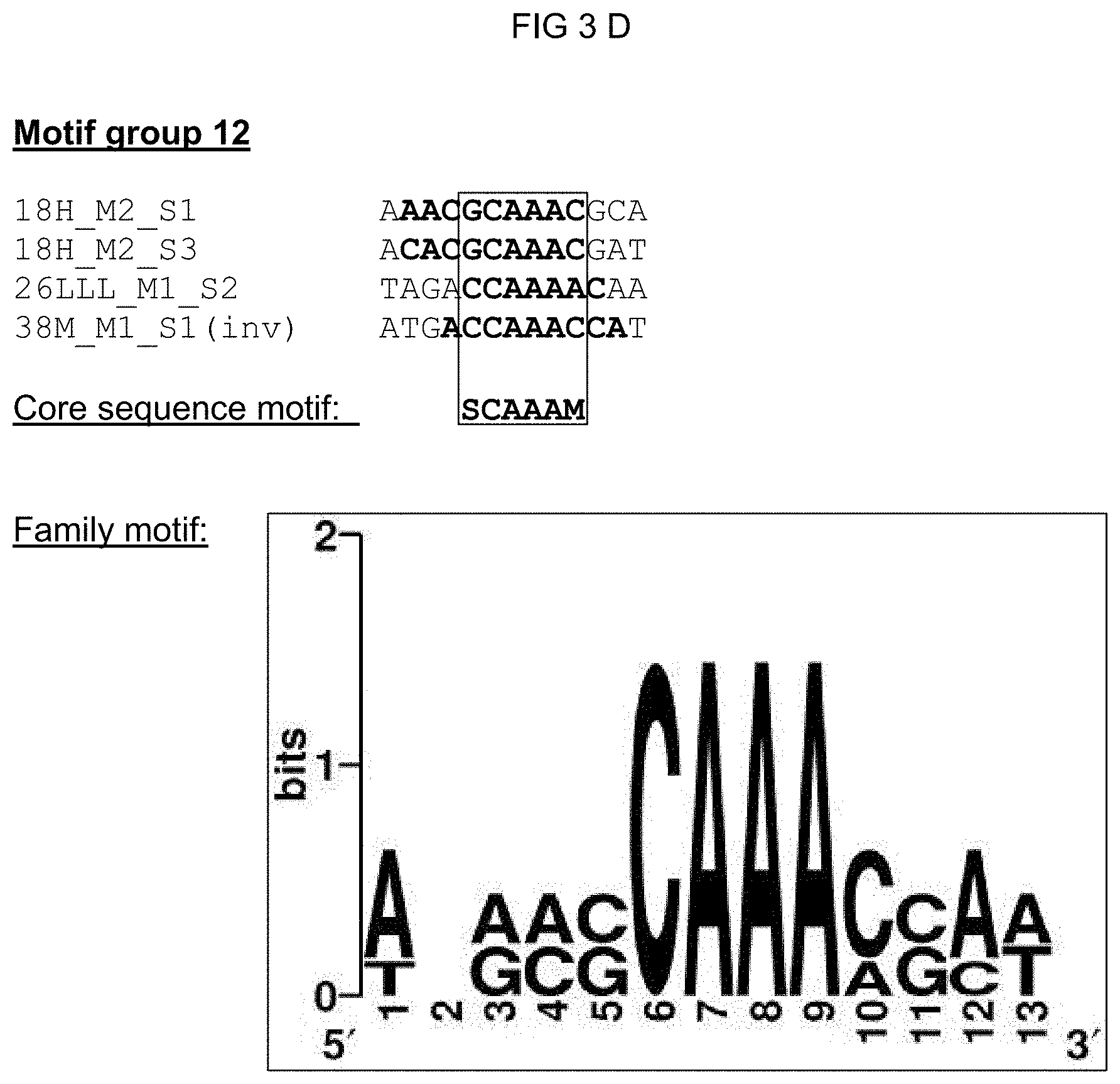
D00030
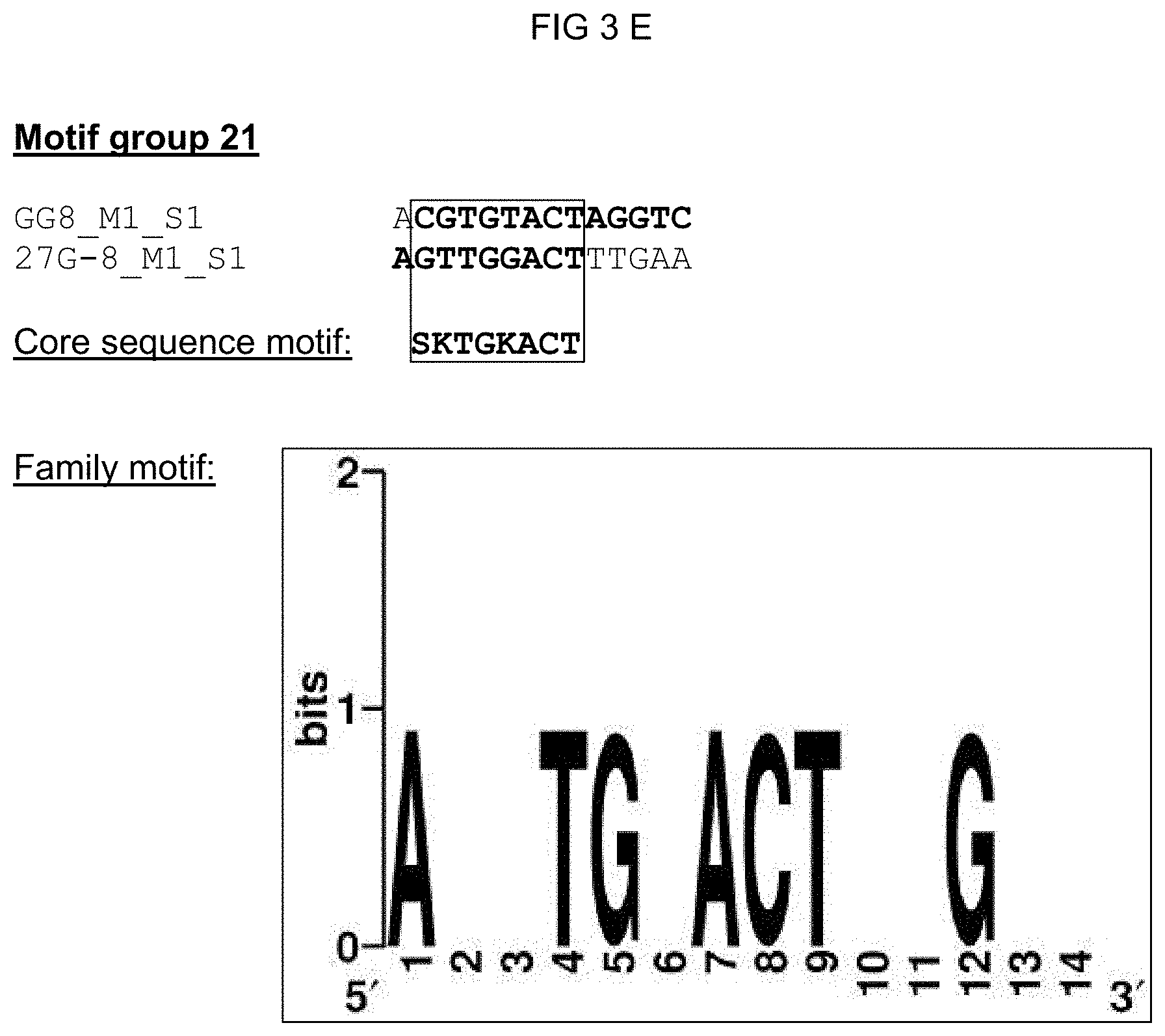
D00031

D00032

D00033
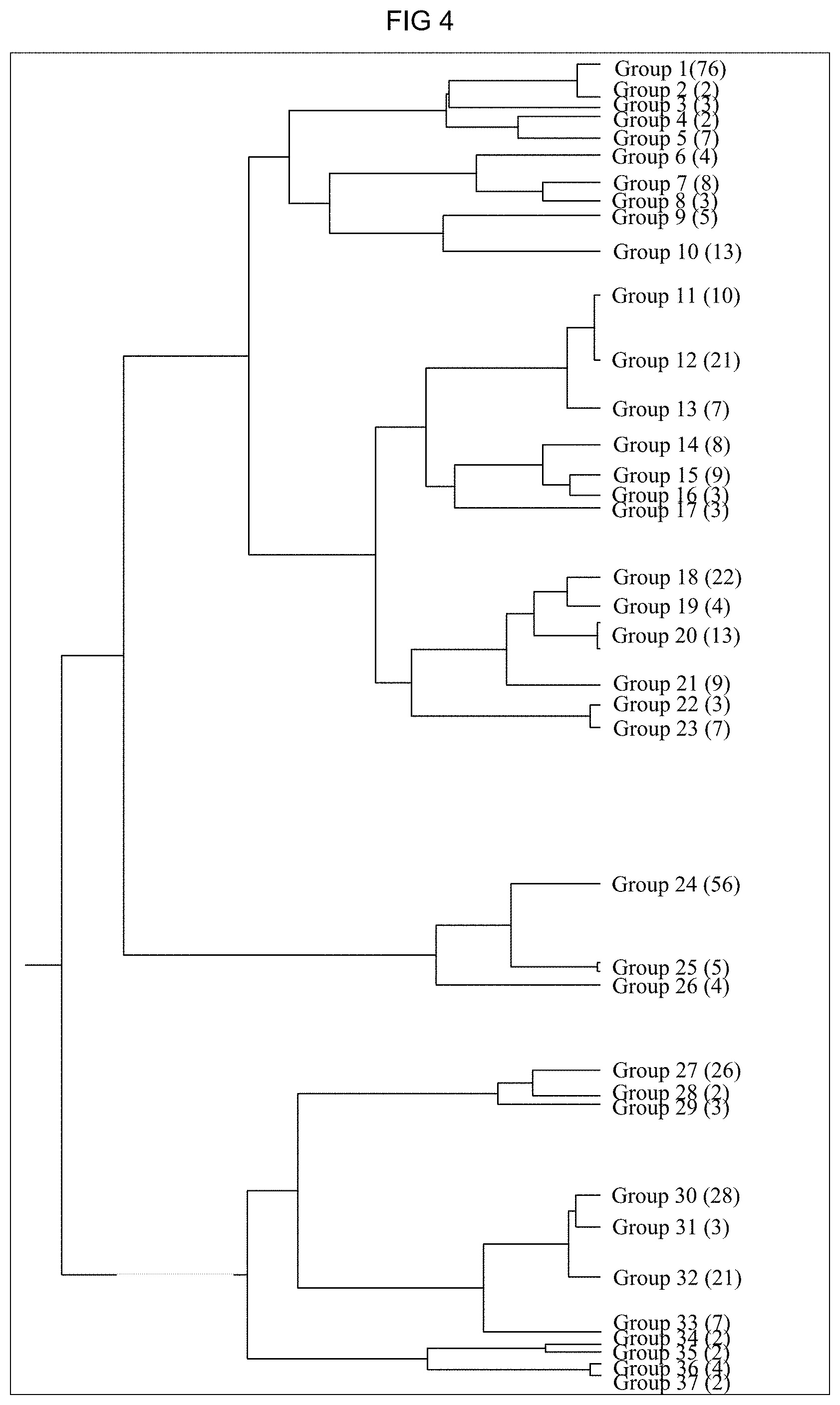
D00034

D00035
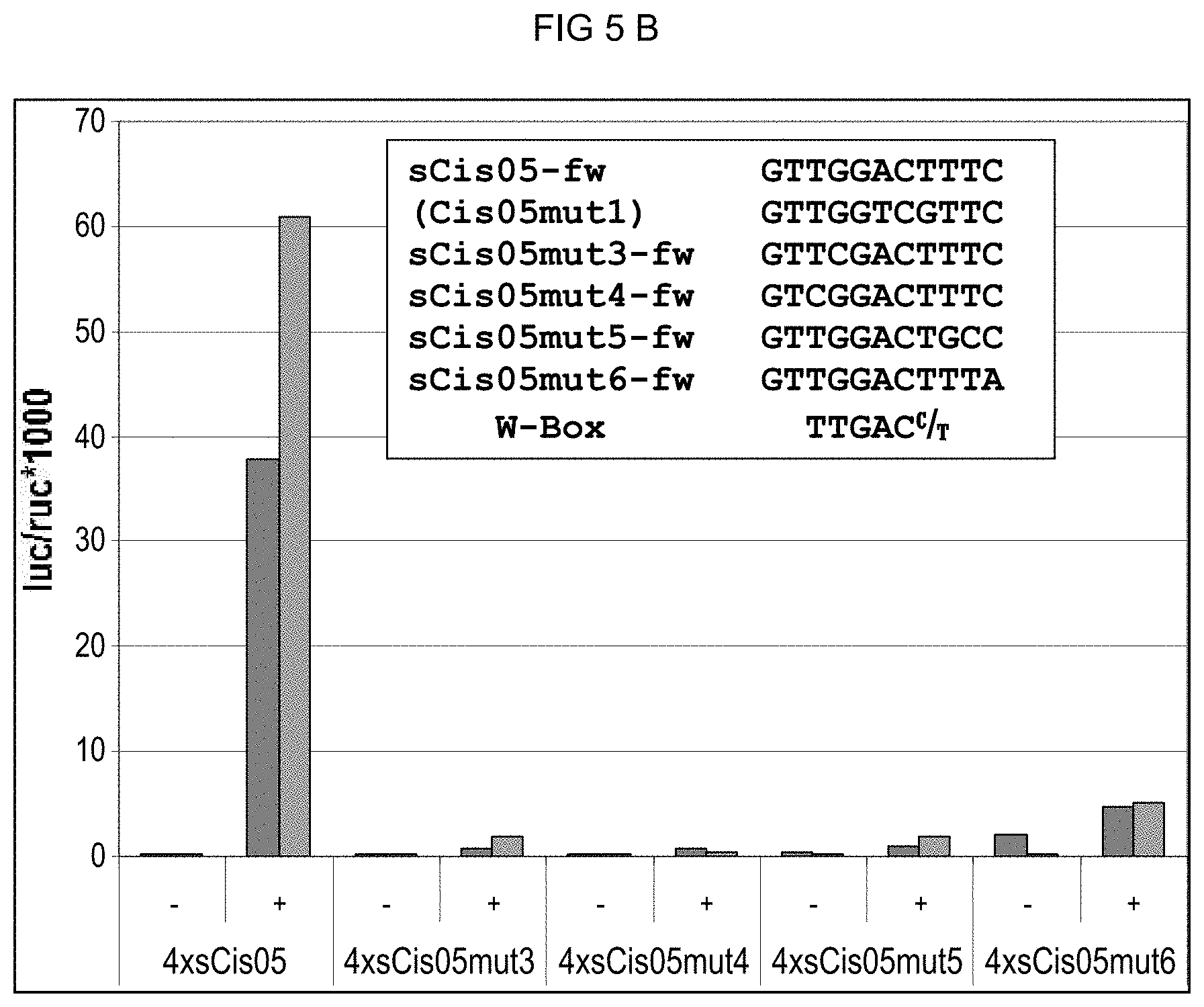
D00036

D00037

D00038
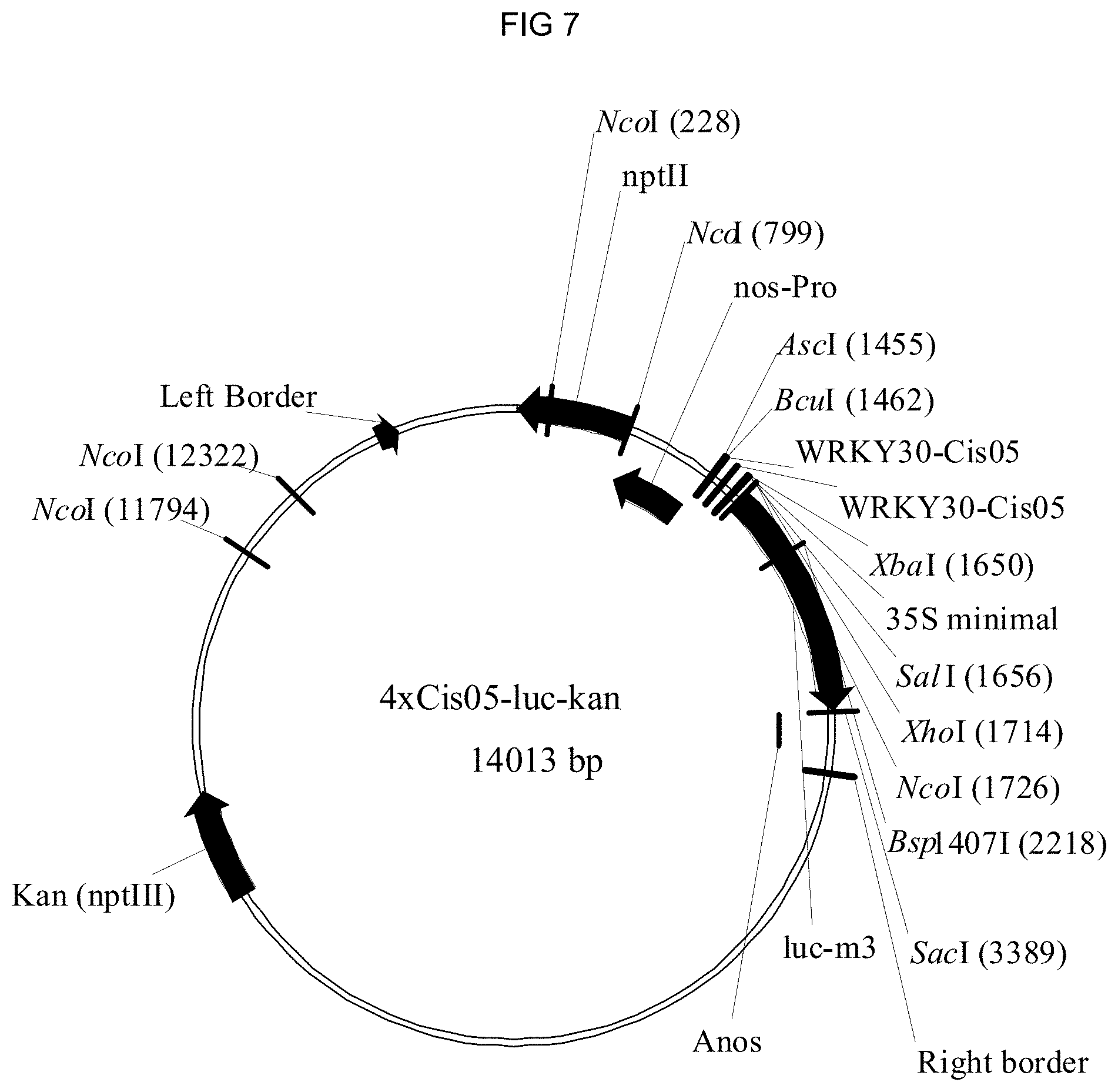
D00039

D00040
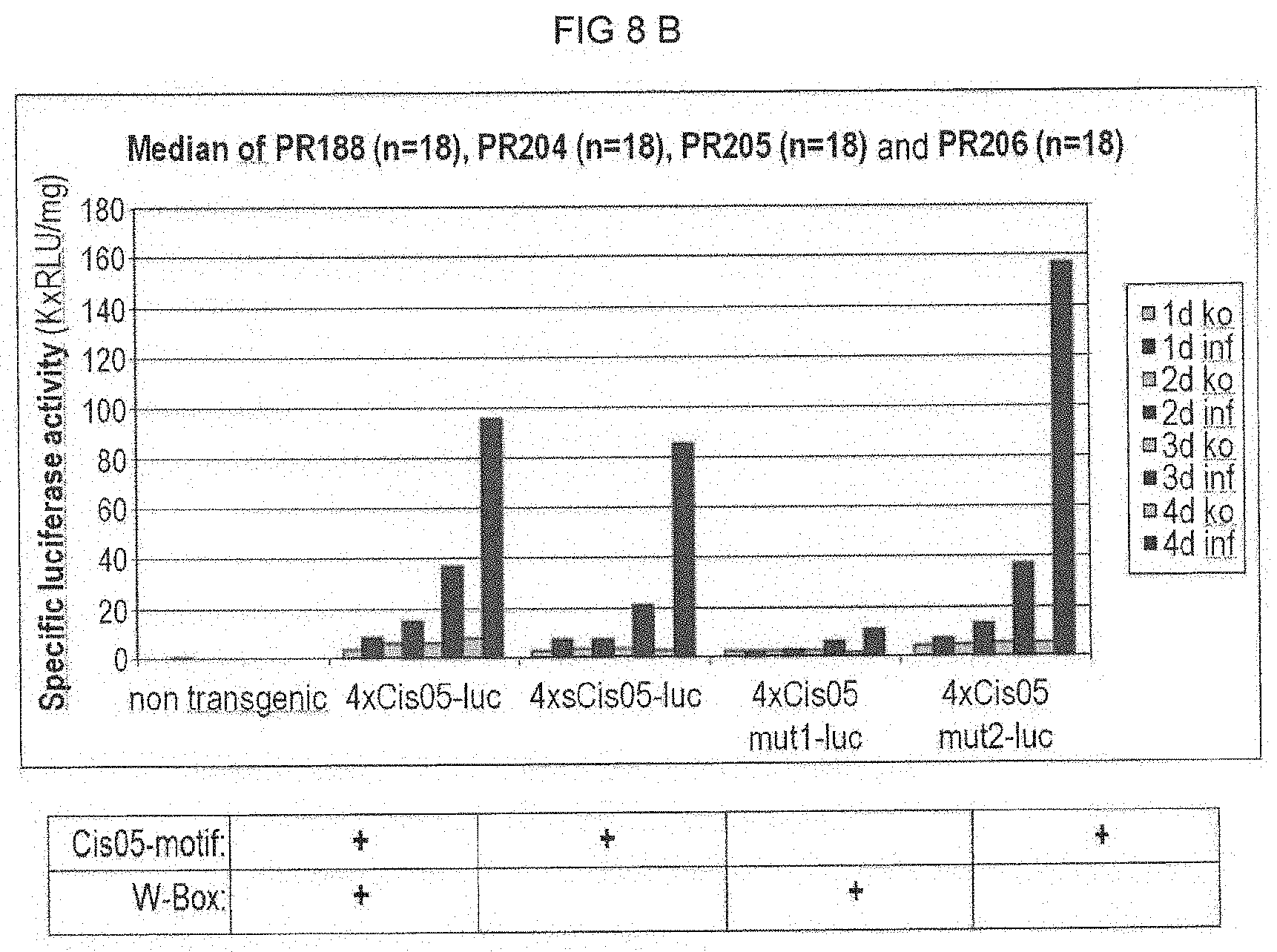
D00041

D00042

D00043
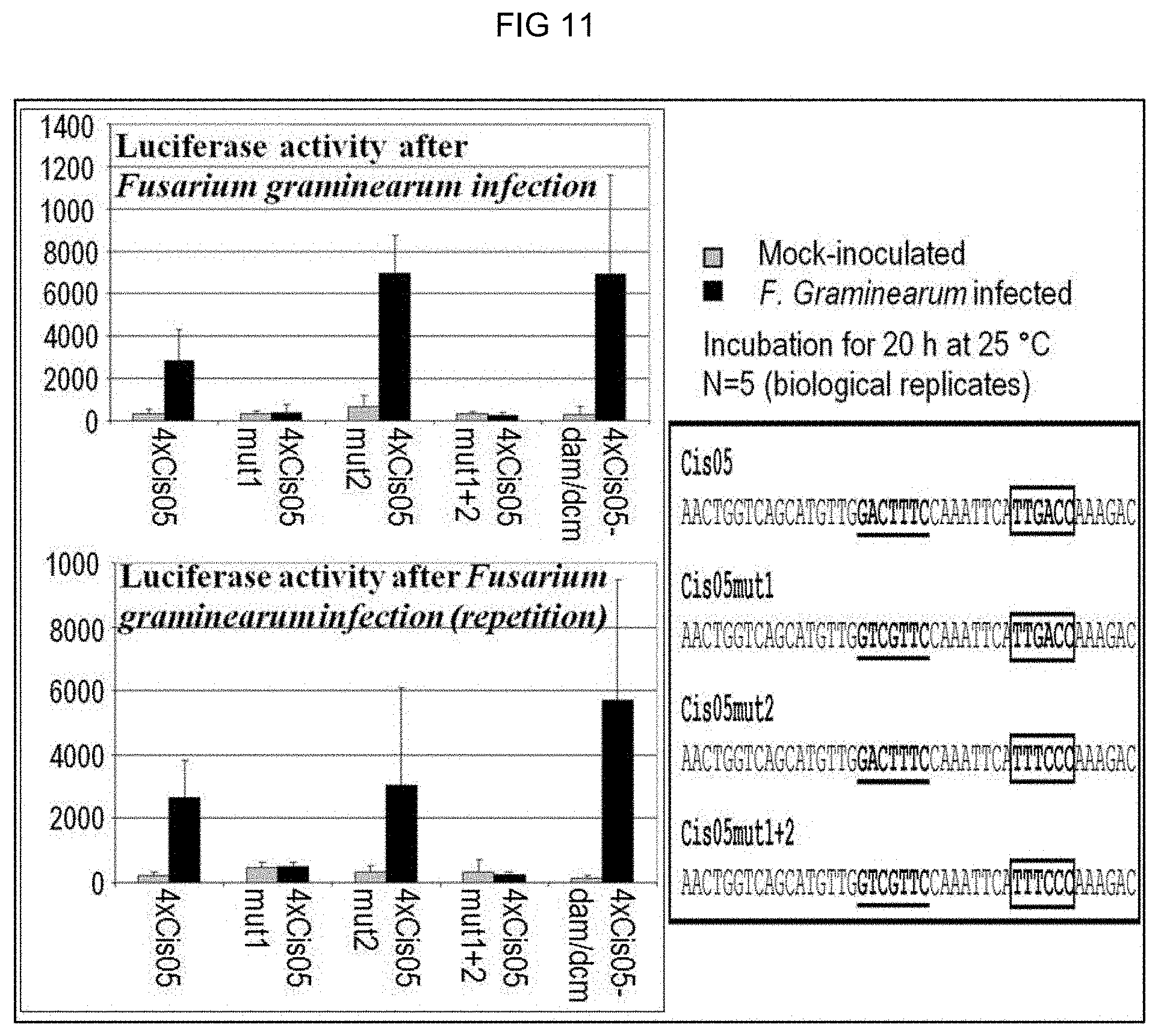
D00044

D00045
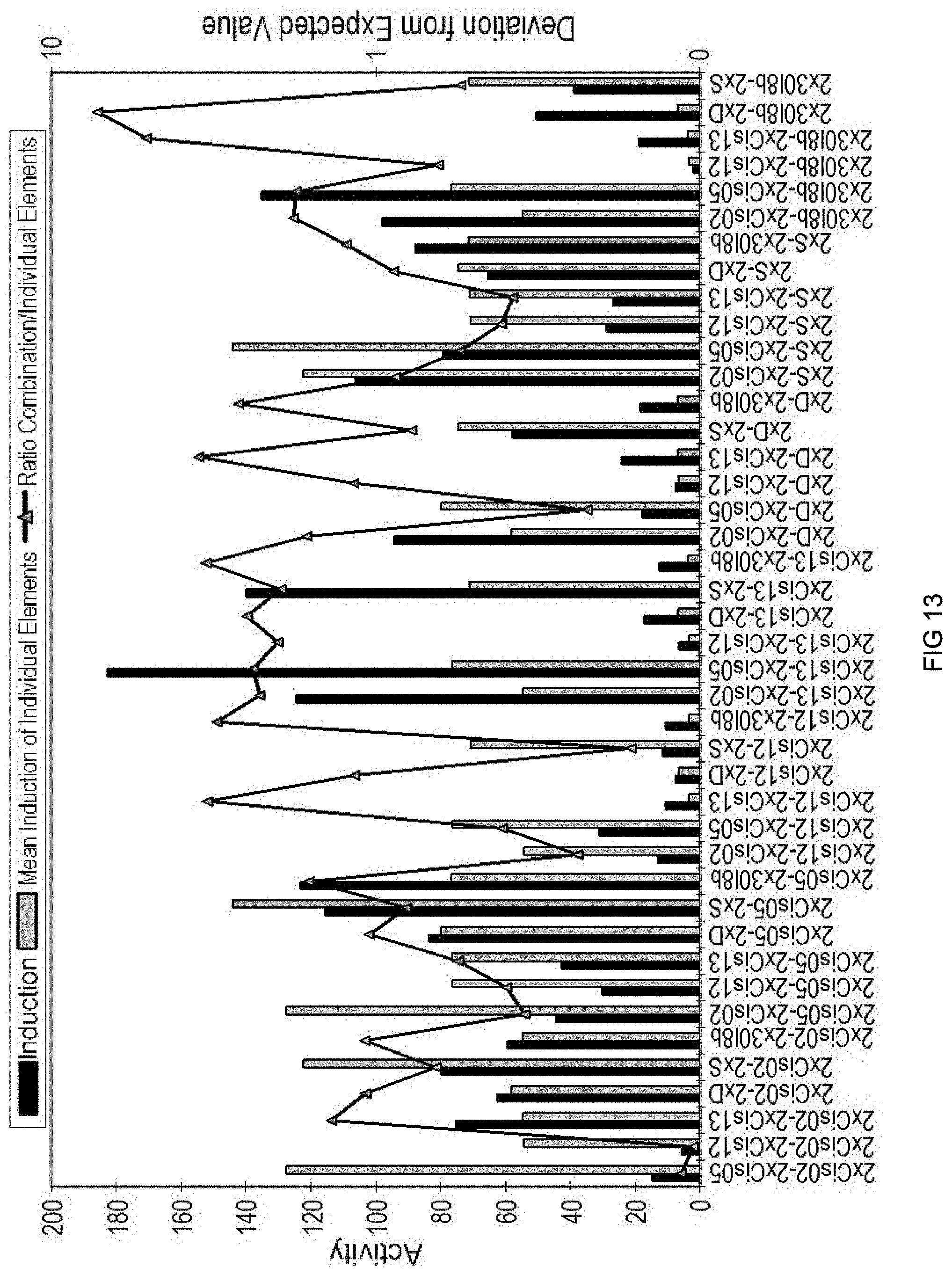
D00046
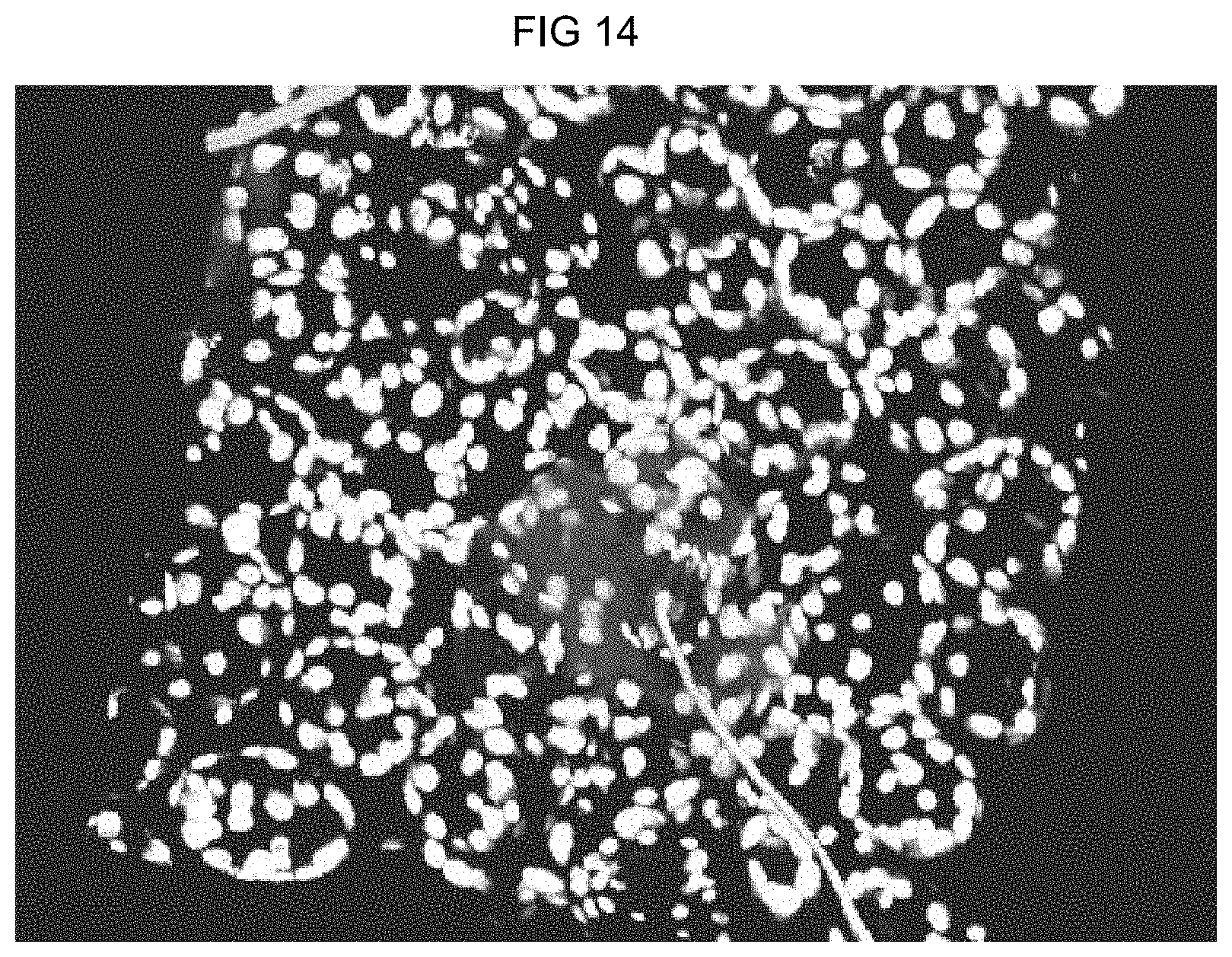

D00047
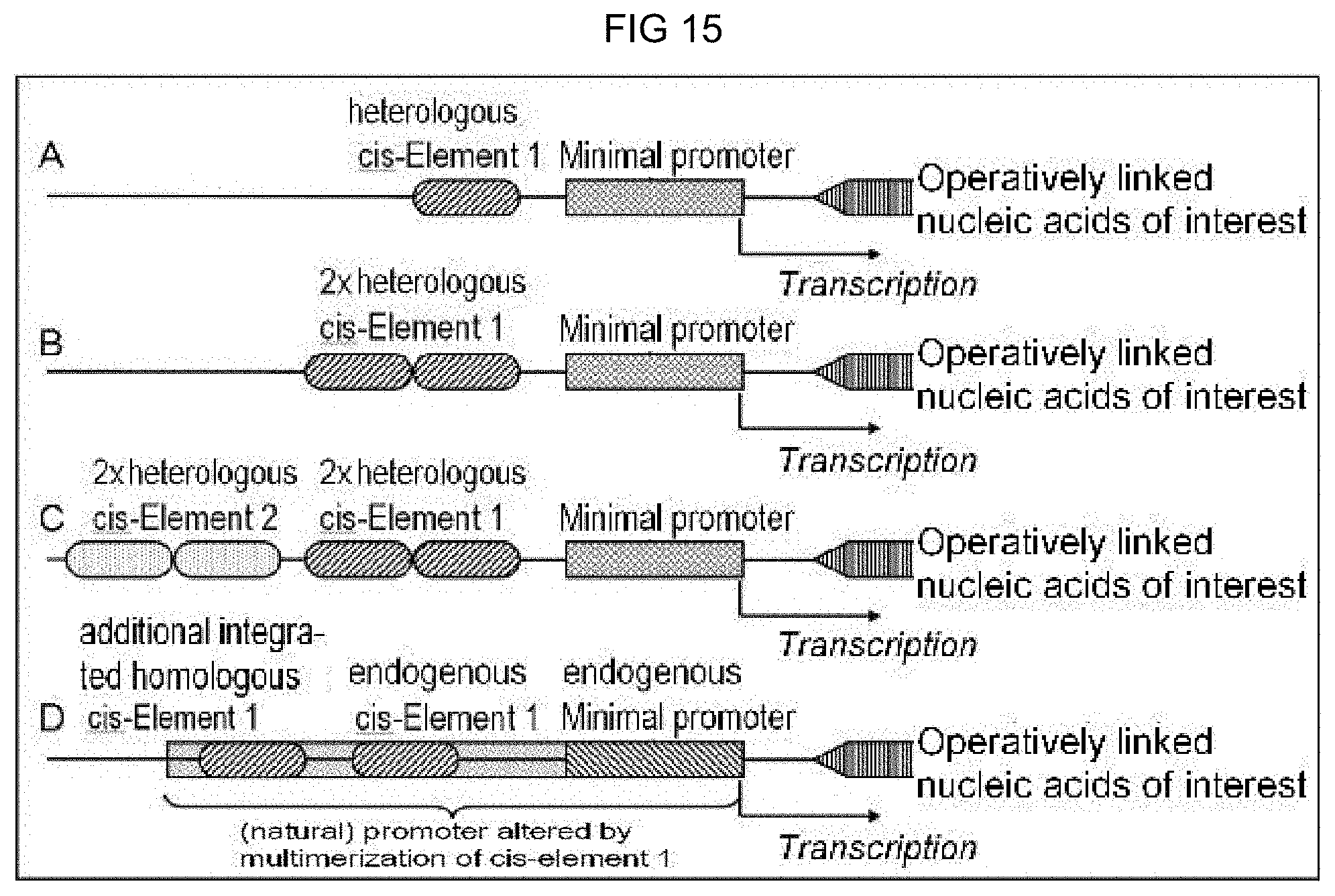
D00048

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.