Mating Factor Alpha Pro-Peptide Variants
Noergaard; Per
U.S. patent application number 16/573586 was filed with the patent office on 2020-01-02 for mating factor alpha pro-peptide variants. The applicant listed for this patent is Novo Nordisk A/S. Invention is credited to Per Noergaard.
| Application Number | 20200002388 16/573586 |
| Document ID | / |
| Family ID | 50184799 |
| Filed Date | 2020-01-02 |

| United States Patent Application | 20200002388 |
| Kind Code | A1 |
| Noergaard; Per | January 2, 2020 |
Mating Factor Alpha Pro-Peptide Variants
Abstract
The present invention is related to Mating Factor .alpha. pro-peptide variants useful for the recombinant expression of polypeptides comprising a GLP-1 peptide in yeasts. The invention is also related to DNA sequences, vectors and host cells for use in expressing polypeptides in yeasts.
| Inventors: | Noergaard; Per; (Humlebaek, DK) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 50184799 | ||||||||||
| Appl. No.: | 16/573586 | ||||||||||
| Filed: | September 17, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15121050 | Aug 24, 2016 | |||
| PCT/EP2015/054298 | Mar 2, 2015 | |||
| 16573586 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 14/605 20130101; C12P 21/02 20130101; C07K 2319/00 20130101; C07K 14/395 20130101 |
| International Class: | C07K 14/395 20060101 C07K014/395; C07K 14/605 20060101 C07K014/605; C12P 21/02 20060101 C12P021/02 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Feb 28, 2014 | EP | 14157172.9 |
Claims
1. A Mating Factor .alpha. pro-peptide variant comprising: an amino acid substitution region located at positions 38-42 of Mating Factor .alpha. having the sequence TABLE-US-00017 (SEQ ID NO: 2) MRFPSIFTAVLFAASSALAAPVNTTTEDETAQIPAEAVIGYLDLEGDFDV AVLPFSNSTNNGLLFINTTIASIAAKEEGVSLDKR,
wherein the amino acid substitution region is represented by formula (I): TABLE-US-00018 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein X.sub.38 is selected from F, L or V, X.sub.39 is selected from L, I, V or M, X.sub.40 is selected from A, G or R, X.sub.41 is selected from S, Y, L, I, V or M, X.sub.42 is selected from Y, W, L, I, V or M; or wherein X.sub.38 is selected from I or V, X.sub.39 is selected from L, I, V or M, X.sub.40 is selected from A, G, Y, F, W, R, K, L, I, V or M, X.sub.41 is selected from Y, F or W, and X.sub.42 is selected from L or I; with the proviso that X.sub.38-X.sub.42 is not VIGYL (SEQ ID NO: 3).
2. The Mating Factor .alpha. pro-peptide variant according to claim 1 wherein X.sub.38 is V, X.sub.39 is selected from L, I, V or M, X.sub.40 is selected from G, R or K, X.sub.41 is Y, and X.sub.42 is selected from L or I.
3. The Mating Factor .alpha. pro-peptide variant according to claim 1, wherein X.sub.38 is V, X.sub.39 is selected from L, I, V or M, X.sub.40 is selected from G or R, X.sub.41 is Y, and X.sub.42 is L.
4. The Mating Factor .alpha. pro-peptide variant according to claim 3, wherein X.sub.40 is R.
5. The Mating Factor .alpha. pro-peptide variant according to claim 1, wherein three of the amino acid residues in X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 are identical to the corresponding amino acid residues in VIGYL (SEQ ID NO:3).
6. A GLP-1 precursor which is a fusion polypeptide comprising: a pre-peptide, a Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO: 3) sequence at positions 38-42 to comprise the amino acid sequence of formula (I): TABLE-US-00019 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein X.sub.38 is selected from F, L or V, X.sub.39 is selected from L, I, V or M, X.sub.40 is selected from G or R, X.sub.41 is selected from S, Y, L, I, V or M, X.sub.42 is selected from Y, W, L, I, V or M, with the proviso that X.sub.38-X.sub.42 is not VIGYL (SEQ ID NO:3), or wherein X.sub.38 is selected from I or V, X.sub.39 is selected from L, I, V or M, X.sub.40 is A, G, Y, F, W, R, K, L, I, V or M, X.sub.41 is selected from Y, F or W, and X.sub.42 is selected from L or I; with the proviso that X.sub.38-X.sub.42 is not VIGYL (SEQ ID NO:3), optionally an extension peptide, and a GLP-1 peptide, wherein said Mating Factor .alpha. pro-peptide is amino acid residues 20-85 in the polypeptide: MRFPSIFTAVLFAASSALAAPVNTTTEDETAQIPAEAVIGYLDLEGDFDVAVLPFSNSTNNGLL FINTTIASIAAKEEGVSLDKR (SEQ ID NO:2), and said variant comprises from 1-15 amino acid substitutions, deletions and/or additions relative to said polypeptide.
7. An expression vector comprising a DNA sequence encoding a polypeptide according to claim 1.
8. A host cell comprising the expression vector according to claim 7.
9. A method for recombinant expression of a polypeptide comprising a GLP-1 peptide in yeast comprising the culturing of a yeast strain comprising a DNA sequence encoding a processing and secretion signal upstream of the polypeptide, wherein said processing and secretion signal comprises a Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 of Mating Factor .alpha. pro-peptide to comprise the amino acid sequence of formula (I): TABLE-US-00020 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein X.sub.38 is selected from F, L or V, X.sub.39 is selected from L, I, V or M, X.sub.40 is selected from A, G or R, X.sub.41 is selected from S, Y, L, I, V or M, X.sub.42 is selected from Y, W, L, V or M; with the proviso that X.sub.38-X.sub.42 is not VIGYL (SEQ ID NO:3); or wherein X.sub.38 is selected from I or V, X.sub.39 is selected from L, I, V or M, X.sub.40 is selected from A, G, Y, F, W, R, K, L, I, V or M, X.sub.41 is selected from Y, F or W, and X.sub.42 is selected from L or I; with the proviso that X.sub.38-X.sub.42 is not VIGYL (SEQ ID NO:3); wherein said Mating Factor .alpha. pro-peptide is amino acid residues 20-85 in the polypeptide: TABLE-US-00021 (SEQ ID NO: 2) MRFPSIFTAVLFAASSALAAPVNTTTEDETAQIPAEAVIGYLDLEGDFDV AVLPFSNSTNNGLLFINTTIASIAAKEEGVSLDKR.
10. The method according to claim 9, wherein X.sub.40 is R.
11. The method according to claim 9, wherein said Mating Factor .alpha. pro-peptide variant has less than 10 amino acid residue changes outside of the X.sub.38-X.sub.42 sequence as compared to the Mating Factor .alpha. pro-peptide as set out in SEQ ID NO:2 (amino acid residues 20-85).
12. The method according to claim 9, wherein said yeast carries at least one genetic modification reducing its capacity for O-glycosylation.
13. The method according to claim 9, wherein the PMT1 gene in said yeast is deleted.
14. The method according to claim 9, wherein said polypeptide comprises GLP-1(9-37)[K34R] or GLP-1(9-37)[K34R,G37K].
15. The method according to claim 9, wherein said polypeptide consists of GLP-1(9-37)[K34R] or GLP-1(9-37)[K34R,G37K].
16. The method according to claim 9, wherein said polypeptide has an N-terminal extension.
17. The method according to claim 9, wherein three of the amino acid residues in X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 are identical to the corresponding amino acid residues in VIGYL (SEQ ID NO:3).
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of U.S. application Ser. No. 15/121,050, filed Aug. 24, 2016, which is a 35 U.S.C. .sctn. 371 national stage application of International Patent Application PCT/EP2015/054298 (published as WO 2015/128507), filed Mar. 2, 2015, which claimed priority of European Patent Application 14157172.9, filed Feb. 28, 2014; the contents of all above-named applications are incorporated herein by reference.
SEQUENCE LISTING
[0002] The instant application contains a Sequence Listing which has been submitted in ASCII format via EFS-Web and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Sep. 17, 2019, is named 130089US03_SeqList.txt and is 11 kilobytes in size.
TECHNICAL FIELD
[0003] The present invention relates to the technical fields of protein expression and protein chemistry where a polypeptide is prepared by recombinant expression in yeast.
BACKGROUND
[0004] The techniques of recombinant polypeptide expression allow for the production of large quantities of desirable polypeptides which may be used for e.g. their biological activity. Such polypeptides are often expressed as recombinant fusion polypeptides in microbial host cells. The polypeptide of interest is often attached to a fusion partner polypeptide in order to increase the expression level, facilitate secretion, increase the solubility, promote polypeptide folding, to protect the polypeptide against unintentional proteolysis or to facilitate purification of the polypeptide of interest.
[0005] To ensure secretion of recombinantly expressed polypeptides from yeast, a pre-pro peptide (often called "leader") is normally fused to the N-terminus of the recombinant product. The pre-sequence ensures translocation of the fusion protein into the endoplasmic reticulum (ER), which is the starting point of the secretory pathway. The pro-sequence ensures further transport from the ER to the Golgi apparatus, where an endogenous protease called Kex2p, often are used to cleave off the pro-sequence. The processed recombinant peptide is subsequently secreted to the growth media, from where it can be purified.
[0006] The pre-pro sequence from Mating Factor Alpha is often used as leader when secreting recombinant polypeptides. However, many other sequences are able to facilitate the secretory process. The leader sequence has a significant influence, not only on the amounts of secreted peptide, but also on the quality in terms of degradation and post-translational modifications such as O-glycosylation. As degradation and O-glycosylation normally are unwanted events, it is desirable with a leader sequence that reduces these modifications to the lowest possible extent, and at the same time maximizes the yield of secreted polypeptide.
[0007] EP 0121884 A2 describes the recombinant production of human insulin in S. cerevisiae using yeast alpha-factor.
[0008] EP 0324274A1 describes the use of a truncated alpha-factor leader sequence for improved expression and secretion of heterologous proteins in yeast.
[0009] Thim et al. (PNAS 83 (1986) 6766-6770) describe the use of Mating Factor Alpha for secretion and processing of insulin precursors in Saccharomyces cerevisiae.
[0010] WO95/34666 describes synthetic leaders for producing secreted polypeptides in S. cerevisiae.
[0011] Rakestraw et al. (Biotech. Bioeng. 103 (2009) 1192-1201) describe mutant Mating Factor Alpha leader sequences which increase the secretion of single-chain antibody and which increase human IgG1 production levels in S. cerevisiae.
[0012] There is a need for more specific leader sequences which increase the yield of the polypeptide precursor, and which reduce the proportion of the recombinant polypeptide which is O-glycosylated. In particular there is a need for leader sequences for expressing GLP-1 peptides in yeast, which leaders increase the yield of the GLP-1 peptides or precursors thereof and lower the O-glycosylation of the GLP-1 peptides. Such more specific leader sequences may facilitate a higher yield of the recombinant polypeptide as well as a lower the amount of O-glycosylated impurities.
SUMMARY
[0013] It is an object of the present invention to provide yeast cells having increased level of expression of heterologous polypeptides. It is also an object of the present invention to provide yeast cells secreting recombinant polypeptide having a reduced amount of O-glycosylated variants of the recombinant polypeptide. In particular it is an object of the present invention to provide yeast cells having both an increased level of expression of the recombinant polypeptide and a reduced amount of O-glycosylated variants. It is an object of the present invention to provide improved expression system for recombinant expression of GLP-1 peptides in yeast cells.
[0014] According to a first aspect of the invention there is provided a method for recombinant expression of a polypeptide comprising a GLP-1 peptide in yeast comprising the culturing of a yeast strain comprising a DNA sequence encoding a processing and secretion signal upstream of the polypeptide, wherein said processing and secretion signal comprises a Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00001 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein
[0015] X.sub.38 is F, L, I or V;
[0016] X.sub.39 L, I, V or M;
[0017] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0018] X.sub.41 is S, Y, F, W, L, I, V or M;
[0019] X.sub.42 is Y, W, L, I, V, M or S;
[0020] with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4).
[0021] In one embodiment of the method for recombinant expression of a polypeptide in yeast the Mating Factor .alpha. pro-peptide variant has at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein X.sub.38 is V; X.sub.39 L, I, V or M; X.sub.40 is G or R; X.sub.41 is Y, and X.sub.42 is L.
[0022] In another embodiment the polypeptide for recombinant expression is a GLP-1 peptide.
[0023] In another embodiment the polypeptide for recombinant expression comprises GLP-1(7-37)[K34R], GLP-1(9-37)[K34R] or GLP-1(9-37)[K34R,G37K].
[0024] According to a second aspect of the invention there is provided a Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein
[0025] X.sub.38 is F, L, I or V;
[0026] X.sub.39 L, I, V or M;
[0027] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0028] X.sub.41 is S, Y, F, W, L, I, V or M;
[0029] X.sub.42 is Y, W, L, I, V, M or S;
with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4).
[0030] According to a third aspect of the invention there is provided a GLP-1 precursor which is a fusion polypeptide comprising: [0031] A pre-peptide, [0032] A Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00002 [0032] (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
[0033] wherein
[0034] X.sub.38 is F, L, I or V;
[0035] X.sub.39 L, I, V or M;
[0036] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0037] X.sub.41 is S, Y, F, W, L, I, V or M;
[0038] X.sub.42 is Y, W, L, I, V, M or S;
[0039] with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4), [0040] Optionally an extension peptide, and [0041] A GLP-1 polypeptide.
[0042] According to a fourth aspect of the invention there is provided a DNA sequence encoding the Mating Factor .alpha. pro-peptide variant or the GLP-1 precursor.
[0043] According to a fifth aspect of the invention there is provided an expression vector comprising the DNA sequence encoding the Mating Factor .alpha. pro-peptide variant or the GLP-1 precursor.
[0044] According to a sixth aspect of the invention there is provided a host cell comprising the expression vector according to the invention.
[0045] In one embodiment the host cell used for expression has a non-functional pmt1 gene or no pmt1 gene at all. Such a host cell has surprisingly been found to lower the amount of O-glycosylated polypeptide independently of, and in addition to, the lowering obtained from the Mating Factor .alpha. pro-peptide variant of the present invention.
BRIEF DESCRIPTION OF DRAWINGS
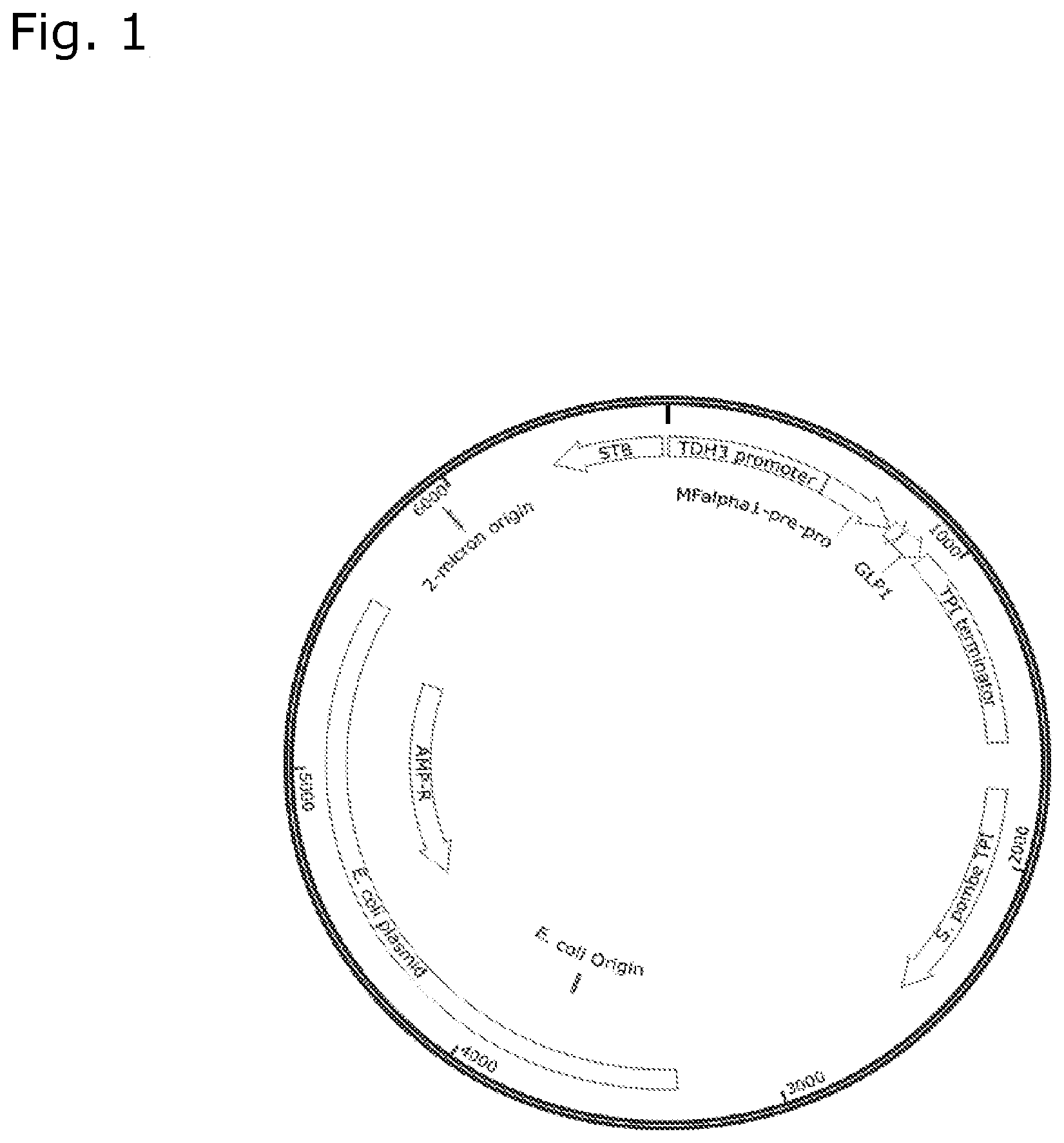
[0046] FIG. 1 shows the minimal expression plasmid as used in Example 1.
[0047] FIG. 2 shows the N-terminally extended GLP-1(7-37)[K34R] precursor including the wildtype Mating Factor .alpha. pre-pro-peptide with indication of the VIGYL (SEQ ID NO:3) subsequence of the pro-peptide.
DESCRIPTION
[0048] According to a first aspect of the invention there is provided a method for recombinant expression of a polypeptide in yeast comprising the culturing of a yeast strain comprising a DNA sequence encoding a processing and secretion signal upstream of the polypeptide, wherein said processing and secretion signal comprises a Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00003 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein
[0049] X.sub.38 is F, L, I or V;
[0050] X.sub.39 L, I, V or M;
[0051] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0052] X.sub.41 is S, Y, F, W, L, I, V or M;
[0053] X.sub.42 is Y, W, L, I, V, M or S;
[0054] with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4).
[0055] The term "leader sequence" as used herein is intended to mean an amino acid sequence consisting of a pre-peptide (the signal peptide) and a pro-peptide. Non-limiting examples of leader sequences are e.g. the alpha-factor signal leader from S. cerevisiae and the synthetic leader sequences for yeast described in WO95/34666.
[0056] "Pre-peptide" as used herein is intended to mean a signal peptide which is present as an N-terminal sequence on the precursor form of a polypeptide to be expressed. The function of the signal peptide is to facilitate translocation of the polypeptide into the endoplasmic reticulum in the host cell. The signal peptide is normally cleaved off in the course of this process. The signal peptide may be heterologous or homologous to the host cell producing the polypeptide.
[0057] "Pro-peptide" as used herein is intended to mean a peptide sequence whose function is to allow the expressed polypeptide to be directed from the endoplasmic reticulum to the Golgi apparatus and further to a secretory vesicle for secretion into the culture medium (i.e. exportation of the polypeptide across the cell wall or at least through the cellular membrane into the periplasmic space of the yeast cell). Non-limiting examples of a pro-peptide are the yeast .alpha.-factor pro-peptide (vide U.S. Pat. Nos. 4,546,082 and 4,870,008) and the synthetic pro-peptides disclosed in U.S. Pat. Nos. 5,395,922; 5,795,746; 5,162,498 and WO 98/32867. The pro-peptide will preferably contain an endopeptidase processing site at the C-terminal end, such as a Lys-Arg sequence or any functional analog thereof.
[0058] The term "Mating Factor .alpha." (MF.alpha., MFa or MFalpha) as used herein is intended to mean the Saccharomyces cerevisiae prepro-sequence, comprising the Mating Factor .alpha. pre-peptide as amino acid residues 1-19 and Mating Factor .alpha. pro-peptide as amino acid residues 20-85 in the structure:
TABLE-US-00004 (SEQ ID NO: 2) MRFPSIFTAVLFAASSALAAPVNTTTEDETAQIPAEAVIGYLDLEGDFDV AVLPFSNSTNNGLLFINTTIASIAAKEEGVSLDKR.
[0059] The Mating Factor .alpha. comprises the sequence VIGYL (SEQ ID NO:3) as amino acid residues 38-42. A variant of Mating Factor .alpha. has been used for recombinant expression of polypeptides which comprises the sequence VIGYS (SEQ ID NO:4) as amino acid residues 38-42 in the Mating Factor .alpha. sequence.
[0060] In the present context the terms "polypeptide", "protein" and "peptide" may be used interchangeably to designate a polypeptide. It is to be understood that the particular term used has no limitation as to the size of the molecule (unless directly stated in the particular context).
[0061] Amino acid residues are generally designated according to single letter abbreviation according to IUPAC nomenclature, e.g. D meaning aspartic acid (Asp) and G meaning glycine. However, in some instances the corresponding three letter abbreviation is also used.
[0062] "Genetically encoded amino acids" as used herein is intended to mean the group consisting of the following amino acids: G, P, A, V, L, I, M, C, F, Y, W, H, K, R, Q, N, E, D, S, T as well as any biological modification hereof. Non-limiting examples of such biological modifications are e.g. amidation, glycosylation and disulphide bond formation.
[0063] "Analogues" as used herein is intended to mean polypeptides which are derived from the reference polypeptide by means of substitution, deletion and/or addition of one or more amino acid residues from the polypeptide. Non-limiting examples of an analogue of GLP-1(7-37) (SEQ ID NO:5) are GLP-1(7-37)[K34R] (SEQ ID NO: 6) where residue 34 has been substituted by an arginine residue and GLP-1(9-37)[K34R] (SEQ ID NO: 7) where residue 34 has been substituted with an arginine residue and amino acid residues 7-8 have been deleted (using the common numbering of amino acid residues for GLP-1 peptides).
[0064] "Variant" as used herein with reference to a polypeptide is intended to mean a chemical variant of the polypeptide which retains substantially the same main function as the original protein. Hence a variant is typically a modified version of a polypeptide wherein as few modifications are introduced as necessary for the modified polypeptide to have some desirable property while preserving substantially the same main function of the original polypeptide. Non-limiting examples of polypeptide variants are e.g. extended polypeptides, truncated polypeptides, fusion polypeptides and analogues. A non-limiting example of a variant of Mating Factor .alpha. pro-peptide is L42S-Mating Factor .alpha.(20-85) (SEQ ID NO:8). A non-limiting example of a variant of GLP-1(7-37) is GLP-1(7-37)[K34R].
[0065] In one embodiment, a variant of a polypeptide comprises from 1-2 amino acid substitutions, deletions or additions as compared to the unmodified polypeptide. In another embodiment, a variant comprises from 1-5 amino acid substitutions, deletions or additions as compared to the unmodified polypeptide. In another embodiment, a variant comprises from 1-15 amino acid substitution, deletion or additions relative to the corresponding unmodified polypeptide.
[0066] According to a second aspect of the invention there is provided a Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00005 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein
[0067] X.sub.38 is F, L, I or V;
[0068] X.sub.39 L, I, V or M;
[0069] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0070] X.sub.41 is S, Y, F, W, L, I, V or M;
[0071] X.sub.42 is Y, W, L, I, V, M or S;
with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4). In one embodiment the Mating Factor .alpha. pro-peptide variant does not comprise VIGYS (SEQ ID NO:4), VIDYS (SEQ ID NO:45), VATYL (SEQ ID NO:46), VIGYR (SEQ ID NO:47), or AIGYL (SEQ ID NO:48) as X.sub.38-X.sub.42.
[0072] According to a third aspect of the invention there is provided a GLP-1 precursor which is a fusion polypeptide comprising: [0073] A pre-peptide, [0074] A Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00006 [0074] (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
[0075] wherein
[0076] X.sub.38 is F, L, I or V;
[0077] X.sub.39 L, I, V or M;
[0078] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0079] X.sub.41 is S, Y, F, W, L, I, V or M;
[0080] X.sub.42 is Y, W, L, I, V, M or S;
[0081] with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4), [0082] Optionally an extension peptide, and [0083] A GLP-1 peptide.
[0084] The term "GLP-1 peptide", as used herein, is intended to designate GLP-1 (7-37), GLP-1 (7-36) amide as well as analogues thereof, which are capable of being produced by conventional recombinant DNA techniques as well as conventional synthetic methods. Such GLP-1 peptides include but are not limited to native glucagon-like peptide-1, for instance such peptide fragments which comprises GLP-1 (7-37) and functional variants thereof as disclosed in WO 87/06941; such peptide fragments which comprise GLP-1 (7-36) and functional derivatives thereof as disclosed in WO 90/11296; such analogues of the active GLP-1 peptides 7-34, 7-35, 7-36, and 7-37 as disclosed in WO 91/11457; such N-terminal truncated fragments of GLP-1 as disclosed in EP 0699686-A2; and such GLP-1 analogues and derivatives that include an N-terminal imidazole group as disclosed in EP 0708179-A2. Non-limiting examples of a GLP-1 peptide is GLP-1(7-37) and GLP-1(7-37)[K34R].
[0085] The term "GLP-1 precursor" as used herein is intended to mean a polypeptide comprising an extended GLP-1 peptide where the extension serves to facilitate the secretion, expression or recovery of the GLP-1 peptide. Examples of GLP-1 precursors may be found in WO03/010186 and WO09/083549. GLP-1 precursors are intended to include GLP-1 peptides having a small extension, e.g. 2-5 amino acid residues, as well as GLP-1 peptides having longer extensions comprising a pre-peptide and a pro-peptide.
[0086] In one embodiment of the method for recombinant expression of a polypeptide in yeast said amino acid sequence of the general formula (I) has a sequence wherein
[0087] X.sub.38 is F, L or V;
[0088] X.sub.39 L, I, V or M;
[0089] X.sub.40 is G or R;
[0090] X.sub.41 is S, Y, L, I, V or M, and
[0091] X.sub.42 is Y, W, L, V or M.
[0092] In another embodiment said amino acid sequence of the general formula (I) has a sequence wherein
[0093] X.sub.38 is I or V;
[0094] X.sub.39 L, I, V or M;
[0095] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0096] X.sub.41 is Y, F, or W, and
[0097] X.sub.42 is L or I.
[0098] In another embodiment said amino acid sequence of the general formula (I) has a sequence wherein
[0099] X.sub.38 is V;
[0100] X.sub.39 L, I, V or M;
[0101] X.sub.40 is G or R;
[0102] X.sub.41 is Y, and
[0103] X.sub.42 is L.
[0104] In another embodiment said amino acid sequence of the general formula (I) has a sequence wherein X.sub.40 is R. In another embodiment said amino acid sequence of the general formula (I) has a sequence wherein X.sub.40 is R and X.sub.42 is L.
[0105] In yet another embodiment said amino acid sequence of the general formula (I) has a sequence wherein X.sub.38 is V, X.sub.39 is I, X.sub.40 is R, X.sub.41 is Y and X.sub.42 is L.
[0106] In another embodiment the polypeptide being recombinantly expressed is a GLP-1 peptide or a variant thereof, such as GLP-1(7-37)[K34R] or GLP-1(9-37)[K34R].
[0107] In one embodiment the polypeptide being recombinantly expressed has an N-terminal extension, i.e. positioned between the Mating Factor .alpha. variant and the polypeptide to be manufactured. This extension may facilitate the expression or secretion of the polypeptide from the host cell, or it may protect part of the polypeptide from undesirable proteolytical processing in the N-terminal. In another embodiment the N-terminal extension is a polypeptide having from 2-10 amino acid residues or having from about 8 to about 200 amino acid residues. Smaller N-terminal extensions are often used when the extension serves to facilitate the expression of the polypeptide in a host cell, or when the extension serves to protect a polypeptide from being proteolytically processed in the N-terminal. In another embodiment the N-terminal extension is selected from the group consisting of EEK, EEAEK (SEQ ID NO:9), HK, EEAHK (SEQ ID NO:10), E(EA)2HK (SEQ ID NO:11), E(EA)3HK (SEQ ID NO:12), EEGHK (SEQ ID NO:13), EHPK, EEGEPK (SEQ ID NO:14), EEAHELK (SEQ ID NO:15), EEAHEVK (SEQ ID NO:16), EEAHEMK (SEQ ID NO:17), EEAHEFK (SEQ ID NO:18), EEAHEYK (SEQ ID NO:19), EEAHEWKEEGNTTPK (SEQ ID NO:20) and EELDARLEALK (SEQ ID NO:21). In another embodiment the N-terminal extension is selected from the group consisting of QPMYKR (SEQ ID NO:22), GQPMYK (SEQ ID NO:23), PGQPMY (SEQ ID NO:24), KPGQPM (SEQ ID NO:25), LKPGQP (SEQ ID NO:26), QLKPGQ (SEQ ID NO:27), LQLKPG (SEQ ID NO:28), WLQLKP (SEQ ID NO:29), HWLQLK (SEQ ID NO:30), WHWLQL (SEQ ID NO:31), AWHWLQ (SEQ ID NO:32), EAWHWL (SEQ ID NO:33), AEAWHW (SEQ ID NO:34) and EAEAWH (SEQ ID NO:35).
[0108] When the expressed polypeptide includes an N-terminal extension, it is customary to cleave off this N-terminal extension by the use of a protease, a peptidase or by chemical cleavage. Proteases such as trypsin, Acromobacter lyticus protease and Enterokinase may be used. The particular proteolytic enzyme selected for the cleavage is often determined by the polypeptide being manufactured. Hence, the person skilled in the art will often select the proteolytic enzyme based on the polypeptides sequence, specifically the presence of any internal primary or secondary cleavage sites, as well as adapting the N-terminal extension to form a good cleavage site.
[0109] The cleavage efficiency of a protease when used to cleave a polypeptide expressed with an N-terminal extension may be determined by a simple assay as follows: A suitable aqueous solution of the polypeptide is incubated at a pH and temperature which is favourable to the protease and samples are withdrawn from the reaction mixture over time. As soon as the samples are withdrawn the enzyme activity is inactivated. After collecting all the samples covering the time-span of interest, the concentration of the corresponding polypeptide without the N-terminal extension is determined by e.g. HPLC analysis. Depicting the concentration of the cleaved polypeptide as a function of time will indicate the progress of the reaction. Comparing such reaction traces for different N-terminal extension of the polypeptide will allow a ranking of the N-terminal extensions in accordance with the ability of the protease to liberate the polypeptide without N-terminal extension.
[0110] The nucleic acid construct encoding the polypeptide may suitably be of genomic, cDNA or synthetic origin. Amino acid sequence alterations are accomplished by modification of the genetic code by well known techniques.
[0111] The DNA sequence encoding the polypeptide are usually inserted into a recombinant vector which may be any vector, which may conveniently be subjected to recombinant DNA procedures, and the choice of vector will often depend on the host cell into which it is to be introduced. Thus, the vector may be an autonomously replicating vector, i.e. a vector, which exists as an extrachromosomal entity, the replication of which is independent of chromosomal replication, e.g. a plasmid. Alternatively, the vector may be one which, when introduced into a host cell, is integrated into the host cell genome and replicated together with the chromosome(s) into which it has been integrated.
[0112] The vector is preferably an expression vector in which the DNA sequence encoding the polypeptide is operably linked to additional segments required for transcription of the DNA. The term, "operably linked" indicates that the segments are arranged so that they function in concert for their intended purposes, e.g. transcription initiates in a promoter and proceeds through the DNA sequence coding for the polypeptide until it terminates within a terminator.
[0113] Thus, expression vectors for use in expressing the polypeptide will comprise a promoter capable of initiating and directing the transcription of a cloned gene or cDNA. The promoter may be any DNA sequence, which shows transcriptional activity in the host cell of choice and may be derived from genes encoding proteins either homologous or heterologous to the host cell.
[0114] Additionally, expression vectors for use of expression of the polypeptide will also comprise a terminator sequence, a sequence recognized by a host cell to terminate transcription. The terminator sequence is operably linked to the 3' terminus of the nucleic acid sequence encoding the polypeptide. Any terminator which is functional in the host cell of choice may be used in the present invention.
[0115] Expression of the polypeptide is aimed for being directed into the secretory pathway for extracellular expression into the growth medium. Useful signal peptides for use as the pre-peptide in leaders for expression in yeast host cells are obtained e.g. from the genes for Saccharomyces cerevisiae alpha-factor and Saccharomyces cerevisiae invertase. Other examples of useful pre-peptides (signal peptides) are the yeast aspartic protease 3 (Yps1) signal peptide (Egel-Mitani et al. (1990) YEAST 6:127-137 and U.S. Pat. No. 5,726,038), the alpha-factor signal of the MF.alpha.1 gene (Thorner (1981) in The Molecular Biology of the Yeast Saccharomyces cerevisiae, Strathern et al., eds., pp 143-180, Cold Spring Harbor Laboratory, NY) and U.S. Pat. No. 4,870,008), the signal peptide of mouse salivary amylase (O. Hagenbuchle et al., Nature 289, 1981, pp. 643-646), a modified carboxypeptidase signal peptide (L. A. Valls et al., Cell 48, 1987, pp. 887-897) and the yeast BAR1 signal peptide (WO 87/02670).
[0116] The procedures used to ligate the DNA sequences coding for the polypeptide, the promoter, the terminator and secretory signal sequence, respectively, and to insert them into suitable vectors containing the information necessary for replication, are well known to persons skilled in the art (cf., for instance, Sambrook et al., Molecular Cloning: A Laboratory Manual, Cold Spring Harbor, N.Y., 1989).
[0117] Many yeast cells contain an endogenous plasmid, called the 2 micron plasmid that contains various elements that ensure its stable maintenance in the yeast cell (Guerineau et al., 1971, Biochem. Biophys. Res. Comm. 42(3):550-557). The whole or part of this endogenous 2 micron can be used in conjunction with a recombinant gene, as a method for securing stable maintenance of the sequences necessary for the recombinant expression (Beggs J. D., 1978, Transformation of yeast by a replicating hybrid plasmid, Nature, 275:104-109). It has been found by the present inventor that when the endogenous 2 micron plasmid is present in the cell, the expression plasmid for recombinant expression only needs to comprise the replication origin and the STB region. The other factors present on the endogenous 2 micron plasmid can function in trans. The replication origin and the STB region only constitute a small part of the endogenous 2p plasmid.
[0118] In an aspect the present invention provides an expression plasmid containing only the replication origin and the STB region from the 2 micron plasmid. Hence, this minimal expression plasmid does not contain any of the FLP region, the repeat 1 region, the REP1 region, D-protein, repeat 2 region and REP2 region. In an embodiment this plasmid comprises an expression cassette, an E. coli part including an AmpR gene, an S. pombe sequence encoding triose-phosphate-isomerase as described in Russell, P R (1985, Transcription of the triose-phosphate gene of Schizosaccharomyces pombe initiates from a start point different from that in Saccharomyces cerevisiae, Gene, 40:125-130). In another embodiment the minimal plasmid comprises no AmpR or other antibiotic resistance gene. Such an antibiotic resistance gene is useful during the cloning work in e.g. E. coli but it is preferable to eliminate the antibiotic resistance gene in the plasmid used for industrial scale recombinant protein expression. The antibiotic resistance gene can be made non-functional or removed from the host cell by well known procedures, see e.g. WO 00/04172. The minimal expression plasmid is useful for expression of a polyeptide in a yeast.
[0119] The vectors of the present invention preferably contain one or more selectable markers which permit easy selection of transformed cells. A selectable marker is a gene the product of which provides for biocide or viral resistance, resistance to heavy metals, complement auxotrophies, and the like. Examples of bacterial selectable markers are the dal genes from Bacillus subtilis or Bacillus licheniformis, or markers which confer antibiotic resistance such as ampicillin, kanamycin, chloramphenicol or tetracycline resistance. Selectable markers for use in auxotroph yeast cells include ADE2, HIS3, LEU2, LYS2, MET3, TRP1, and URA3. A preferred selectable marker for yeast is the Schizosaccharomyces pombe TPI gene (Russell (1985) Gene 40:125-130).
[0120] In the vector, the polynucleotide sequence is operably connected to a suitable promoter sequence. The promoter may be any nucleic acid sequence which shows transcriptional activity in the host cell of choice including mutant, truncated, and hybrid promoters, and may be obtained from genes encoding extra-cellular or intra-cellular polypeptides either homologous or heterologous to the host cell. Examples of useful promoters in yeast host cells are the Saccharomyces cerevisiae MF.alpha.1, TPI1, ADH2, TDH3 or PGK1 promoters.
[0121] The polynucleotide construct of the invention will also typically be operably connected to a suitable terminator. In yeast an example of a suitable terminator is the TPI1 terminator (Alber et al. (1982) J. Mol. Appl. Genet. 1:419-434), but could be any endogenous yeast terminator.
[0122] The procedures used to ligate the polynucleotide sequence of the invention, the promoter and the terminator, respectively, and to insert them into suitable yeast vectors containing the information necessary for yeast replication, are well known to persons skilled in the art. It will be understood that the vector may be constructed either by first preparing a DNA construct containing the entire DNA sequence encoding the polypeptide to be expressed, and subsequently inserting this fragment into a suitable expression vector, or by sequentially inserting DNA fragments containing genetic information for the individual elements (such as the Mating Factor .alpha. variant of the invention, the polypeptide to be expressed optionally including a N-terminal extension) followed by assembly of the elements by ligation, seamless cloning methods or by cloning directly in the yeast cell by homologous recombination.
[0123] The present invention also relates to recombinant host cells, comprising a polynucleotide sequence encoding the Mating Factor .alpha. variant of the invention and the polypeptide to be expressed. A vector comprising such polynucleotide sequence is introduced into the host cell so that the vector is maintained as a chromosomal integrant or as a self-replicating extra-chromosomal vector.
[0124] "Host cell" as used herein is intended to mean a microorganism which is used for the expression of a polypeptide of interest. A host cell encompasses any progeny of a parent cell that is not identical to the parent cell due to mutations that occur during replication.
[0125] A suitable host cell for the present invention is a yeast cell. "Yeast" as used herein includes ascosporogenous yeast (Endomycetales), basidiosporogenous yeast, and yeast belonging to the Fungi Imperfecti (Blastomycetes). The ascosporogenous yeasts are divided into the families Spermophthoraceae and Saccharomycetaceae. The latter is comprised of four subfamilies, Schizosaccharomycoideae (e.g., genus Schizosaccharomyces), Nadsonioideae, Lipomycoideae, and Saccharomycoideae (e.g., genera Pichia, Kluyveromyces and Saccharomyces). The basidiosporogenous yeasts include the genera Leucosporidim, Rhodosporidium, Sporidiobolus, Filobasidium, and Filobasidiella. Yeast belonging to the Fungi Imperfecti are divided into two families, Sporobolomycetaceae (e.g., genera Sorobolomyces and Bullera) and Cryptococcaceae (e.g., genus Candida). Since the classification of yeast may change in the future, for the purposes of this invention, yeast shall be defined as described in Biology and Activities of Yeast (Skinner, F. A., Passmore, S. M., and Davenport, R. R., eds, Soc. App. Bacteriol. Symposium Series No. 9, 1980. The biology of yeast and manipulation of yeast genetics are well known in the art (see, e.g., Biochemistry and Genetics of Yeast, Bacil, M., Horecker, B. J., and Stopani, A. O. M., editors, 2nd edition, 1987; The Yeasts, Rose, A. H., and Harrison, J. S., editors, 2nd edition, 1987; and The Molecular Biology of the Yeast Saccharomyces, Strathern et al., editors, 1981).
[0126] The yeast host cell used in the process of the invention may be any suitable yeast organism which, on cultivation, produces large amounts of the polypeptide to be expressed.
[0127] Examples of suitable yeast organisms are strains selected from a cell of a species of Candida, Kluyveromyces, Saccharomyces, Schizosaccharomyces, Pichia, Hansenula, and Yarrowia. In one embodiment, the yeast host cell is selected from a Saccharomyces carlsbergensis, Saccharomyces cerevisiae, Saccharomyces diastaticus, Saccharomyces douglasii, Saccharomyces kluyveri, Saccharomyces norbensis, Saccharomyces oviformis, Schizosaccharomyces pombe, Sacchoromyces uvarum, Pichia kluyveri, Yarrowia lipolytica, Candida utilis, Candida cacaoi, and Geotrichum fermentans. Other useful yeast host cells are a Kluyveromyces lactis, Kluyveromyces fragilis, Hansenula polymorpha, Pichia pastoris Yarrowia lipolytica, Schizosaccharomyces pombe, Ustilgo maylis, Candida maltose, Pichia guillermondii and Pichia methanoliol (cf. Gleeson et al., J. Gen. Microbiol. 132, 1986, pp. 3459-3465; U.S. Pat. Nos. 4,882,279 and 4,879,231). The transformation of the yeast cells may for instance be effected by protoplast formation followed by transformation in a manner known per se.
[0128] The host cell for expressing the polypeptide is preferably a cell free from any functional antibiotic resistance genes. Although such antibiotic resistance genes are useful during initial cloning steps in e.g. E. coli, the antibiotic resistance genes can be made non-functional or removed from the host cell by well known procedures, see e.g. WO 00/04172.
[0129] "Medium" as used herein is intended to mean a liquid solution for cultivating the host cell, i.e. supporting the growth and product formation of the yeast. A suitable medium for yeast is e.g. YPD or as described in WO2008/037735. The medium contains at least one carbon source, one or several nitrogen sources, essential salts including salts of potassium, sodium, magnesium, phosphate and sulphate, trace metals, water soluble vitamins, and process aids including but not limited to antifoam agents, protease inhibitors, stabilizers, ligands and inducers. Typical carbon sources are e.g. mono- or disaccharides. Typical nitrogen sources are, e.g. ammonia, urea, amino acids, yeast extract, corn steep liquor and fully or partially hydrolysed proteins. Typical trace metals are e.g. Fe, Zn, Mn, Cu, Mo and H.sub.3BO.sub.3. Typical water soluble vitamins are e.g. biotin, pantothenate, niacin, thiamine, p-aminobenzoic acid, choline, pyridoxine, folic acid, riboflavin and ascorbic acid.
[0130] By "fermentation" as used herein is intended to mean an aseptic process used for propagating microorganisms submerged in a liquid medium. The fermentation is preferably carried out in aseptic, stirred tanks with supply lines for addition of compressed, sterile gasses consisting of but not limited to air, oxygen and ammonia. A fermentation tank can contain sensors and devices for monitoring pH, temperature, pressure, agitation rate, dissolved oxygen level, liquid content, foam level, feed addition rates and rates of adding acid and base. Furthermore, the fermentation tank can be equipped with optical devices for monitoring levels of cell density, concentrations of metabolites and products regardless of their physio-chemical form.
[0131] The desired product produced during the fermentation is present as soluble extracellular material or as intracellular material either in the form of soluble material or as insoluble material including aggregated material. It is preferable that it is present as soluble extracellular material. A fermentation process is typically carried out in tanks with a working volume ranging from 100 mL to 200.000 L. A fermentation process can be operated as a batch process, a fed-batch process, a repeated fed-batch process or a continuous process.
[0132] The secreted polypeptides, a significant proportion of which will be present in the medium in correctly processed form, may be recovered from the medium by conventional procedures including separating the yeast cells from the medium by centrifugation, filtration or catching the polypeptide by an ion-exchange matrix or by a reverse phase absorption matrix, precipitating the protein components of the supernatant or filtrate by means of a salt, e.g. ammonium sulphate, followed by purification by a variety of chromatographic procedures, e.g. ion exchange chromatography, affinity chromatography, or the like.
[0133] The novel Mating Factor .alpha. variants of the present invention also facilitate reduced O-glycosylation of the polypeptide during expression in yeast. As such the Mating Factor .alpha. variants of the present invention can be used in an improved method for making polypeptides such as GLP-1 peptides in yeast. Expressing the polypeptide in a yeast cell having reduced capacity for O-glycosylation may maintain the improved yield of said precursor while at the same time reducing even further the fraction of said polypeptide that is O-glycosylated during expression.
[0134] Protein O-mannosyltransferases (PMTs) initiate the assembly of O-mannosyl glycans, an essential protein modification in fungi. PMTs are conserved in fungi and the PMT family is phylogenetically classified into PMT1, PMT2 and PMT4 subfamilies, which differ in protein substrate specificity. The protein O-mannosyltransferases Pmt1p and Pmt2p are catalyzing the O-glycosylation of serine and threonine residues in proteins in the endoplasmic reticulum of yeast by transfer of a mannosyl residue from Dolichyl phosphate-D-mannose (Gentzsch et al., FEBS Lett 1995, 18, pp 128-130). In Saccharomyces cerevisiae as well as in many other yeasts the PMT family is highly redundant, and only the simultaneous deletion of PMT1/PMT2 and PMT4 subfamily members is lethal (Girrbach and Strahl, J. Biol. Chem. 2003, 278, pp 12554-62). U.S. Pat. No. 5,714,377 describe that yeast cells having reduced O-glycosylation capacity from PMT1/PMT2 modification are still viable and show good growth characteristics in industrial fermentation conditions.
NON-LIMITING EMBODIMENTS
[0135] The invention is further described by the following non-limiting embodiments:
1. A method for recombinant expression of a polypeptide in yeast comprising the culturing of a yeast strain comprising a DNA sequence encoding a processing and secretion signal upstream of the polypeptide, wherein said processing and secretion signal comprises a Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00007 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein
[0136] X.sub.38 is F, L, I or V;
[0137] X.sub.39 L, I, V or M;
[0138] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0139] X.sub.41 is S, Y, F, W, L, I, V or M;
[0140] X.sub.42 is Y, W, L, I, V, M or S;
[0141] with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4).
2. The method according to embodiment 1 wherein
[0142] X.sub.38 is F, L or V;
[0143] X.sub.39 L, I, V or M;
[0144] X.sub.40 is G or R;
[0145] X.sub.41 is S, Y, L, I, V or M, and
[0146] X.sub.42 is Y, W, L, V or M.
3. The method according to claim 1 wherein
[0147] X.sub.38 is I or V;
[0148] X.sub.39 L, I, V or M;
[0149] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0150] X.sub.41 is Y, F, or W, and
[0151] X.sub.42 is L or I.
4. The method according to any of embodiments 1-3 wherein
[0152] X.sub.38 is V;
[0153] X.sub.39 L, I, V or M;
[0154] X.sub.40 is G or R;
[0155] X.sub.41 is Y, and
[0156] X.sub.42 is L.
5. The method according to any of embodiments 1-4, wherein four of the amino acid residues in X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 are identical to the corresponding amino acid residue in VIGYL (SEQ ID NO:3) or VIGYS (SEQ ID NO:4). 6. The method according to any of embodiments 1-5, wherein at least three of the amino acid residues in X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 are identical to the corresponding amino acid residue in VIGYL (SEQ ID NO:3) or VIGYS (SEQ ID NO:4). 7. The method according to any of embodiments 1-6, wherein X.sub.41 is Y. 8. The method according to any of embodiments 1-6, wherein X.sub.41 is L. 9. The method according to any of embodiments 1-8, wherein X.sub.42 is L. 10. The method according to any of embodiments 1 and 5-8, wherein X.sub.42 is S. 11. The method according to any of embodiments 1 and 5-8, wherein X.sub.42 is Y, W, L, I, V or M. 12. The method according to any of embodiments 1-11, wherein X.sub.40 is R. 13. The method according to embodiment 12, wherein X.sub.38 is V, X.sub.39 is I, X.sub.41 is Y and X.sub.42 is L. 14. The method according to embodiment 12, wherein X.sub.38 is V, X.sub.39 is I, X.sub.41 is Y and X.sub.42 is S. 15. The method according to any of embodiments 1 and 12, wherein formula (I) is VI-X.sub.40-YL or VI-X.sub.40-YS. 16. The method according to embodiment 15 wherein X.sub.40 is A, Y, F, W, R, K, L, I, V or M. 17. The method according to any of embodiments 1-16, wherein said Mating Factor .alpha. pro-peptide variant has less than 10 amino acid residue changes outside of the X.sub.38-X.sub.42 sequence as compared to the Mating Factor .alpha. pro-peptide as set out in SEQ ID NO:2 (amino acid residues 20-85). 18. The method according to any of embodiments 1-17, wherein said Mating Factor .alpha. pro-peptide variant has less than 5 amino acid residue changes outside of the X.sub.38-X.sub.42 sequence as compared to the Mating Factor .alpha. pro-peptide as set out in SEQ ID NO:2 (amino acid residues 20-85). 19. The method according to any of embodiments 1-18, wherein said Mating Factor .alpha. pro-peptide variant has less than 2 amino acid residue changes outside of the X.sub.38-X.sub.42 sequence as compared to the Mating Factor .alpha. pro-peptide as set out in SEQ ID NO:2 (amino acid residues 20-85). 20. The method according to any of embodiments 1-19, wherein said Mating Factor .alpha. pro-peptide variant is part of a Mating Factor .alpha. prepro-peptide comprising a pre-peptide as the N-terminal part fused to said Mating Factor .alpha. pro-peptide variant as the C-terminal part. 21. The method according to embodiment 20, wherein said pre-peptide is from yeast aspartic protease 3 (YAP3) signal peptide, from the a-factor signal of the MF.alpha.1 gene from S. cerevisiae or a variant thereof. 22. The method according to any of embodiments 1-21, wherein said yeast carries at least one genetic modification reducing its capacity for O-glycosylation. 23. The method according to embodiment 22, wherein said yeast carries at least one genetic modification within the genes for PMT1 or PMT2 reducing its capacity for O-glycosylation. 24. The method according to any of embodiments 22-23, wherein said yeast carries at least one genetic modification reducing its capacity for O-glycosylation by the protein O-mannosyltransferase 1 (PMT1) of the polypeptide GLP-1(7-37)[K34R] when expressed with the alpha leader as compared to the yeast carrying the corresponding unmodified genes. 25. The method according to any of embodiments 22-24, wherein said yeast carries at least one genetic modification reducing its capacity for O-glycosylation by protein O-mannosyltransferase 2 (PMT2) of the polypeptide GLP-1(7-37)[K34R] when expressed with the alpha leader as compared to the yeast carrying the corresponding unmodified genes. 26. The method according to any of embodiments 22-25, wherein said capacity for O-glycosylation is reduced by at least a factor 2. 27. The method according to any of embodiments 22-26, wherein said capacity for O-glycosylation is reduced by at least a factor 4. 28. The method according to any of embodiments 22-27, wherein said at least one genetic modification is located in the coding region of PMT1 or PMT2. 29. The method according to any of embodiments 22-27, wherein said at least one genetic modification is located in the regions responsible for or involved in the expression and/or transcriptional regulation of PMT1 or PMT2. 30. The method according to any of embodiments 22-27, wherein the PMT1 gene in said yeast is deleted. 31. The method according to any of embodiments 22-27, wherein the PMT1 and PMT2 genes in said yeast are both deleted. 32. The method according to any of embodiments 1-31, wherein said polypeptide comprises a GLP-1 peptide. 33. The method according to embodiment 32, wherein said polypeptide comprises GLP-1(9-37)[K34R] or GLP-1(9-37)[K34R]. 34. The method according to embodiment 33, wherein said polypeptide is GLP-1(7-37)[K34R], GLP-1(9-37)[K34R] or GLP-1(9-37)[K34R,G37K] (SEQ ID NO:43). 35. The method according to any of embodiments 32-34, wherein said polypeptide has an N-terminal extension. 36. The method according to embodiment 35, wherein said N-terminal extension is selected from the group consisting of EEK, EEAEK (SEQ ID NO:9), HK, EEAHK (SEQ ID NO:10), E(EA)2HK (SEQ ID NO:11), E(EA)3HK (SEQ ID NO:12), EEGHK (SEQ ID NO:13), EHPK, EEGEPK (SEQ ID NO:14), EEAHELK (SEQ ID NO:15), EEAHEVK (SEQ ID NO:16), EEAHEMK (SEQ ID NO:17), EEAHEFK (SEQ ID NO:18), EEAHEYK (SEQ ID NO:19), EEAHEWKEEGNTTPK (SEQ ID NO:20) and EELDARLEALK (SEQ ID NO:21). 37. The method according to embodiment 35, wherein said N-terminal extension is selected from the group consisting of DV, DVKPGQPLA (SEQ ID NO:36), DVKPGQPEY (SEQ ID NO:37), DVKPGEPLY (SEQ ID NO:38), DVKPGQPLY (SEQ ID NO:39), DVKPGQPLE (SEQ ID NO:40), DVKPGQPMY (SEQ ID NO:41) and DVKPGQPMYDDDDK (SEQ ID NO:42). 38. The method according to embodiment 35, wherein said N-terminal extension is selected from the group consisting of QPMYKR (SEQ ID NO:22), GQPMYK (SEQ ID NO:23), PGQPMY (SEQ ID NO:24), KPGQPM (SEQ ID NO:25), LKPGQP (SEQ ID NO:26), QLKPGQ (SEQ ID NO:27), LQLKPG (SEQ ID NO:28), WLQLKP (SEQ ID NO:29), HWLQLK (SEQ ID NO:30), WHWLQL (SEQ ID NO:31), AWHWLQ (SEQ ID NO:32), EAWHWL (SEQ ID NO:33), AEAWHW (SEQ ID NO:34) and EAEAWH (SEQ ID NO:35). 39. Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00008 (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
wherein
[0157] X.sub.38 is F, L, I or V;
[0158] X.sub.39 L, I, V or M;
[0159] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0160] X.sub.41 is S, Y, F, W, L, I, V or M;
[0161] X.sub.42 is Y, W, L, I, V, M or S;
with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4). 40. The Mating Factor .alpha. pro-peptide variant according to embodiment 39 wherein X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4), VIDYS (SEQ ID NO:45), VATYL (SEQ ID NO:46), VIGYR (SEQ ID NO:47), or AIGYL (SEQ ID NO:48). 41. GLP-1 precursor which is a fusion polypeptide comprising: [0162] A pre-peptide, [0163] A Mating Factor .alpha. pro-peptide variant having at least one substitution in the VIGYL (SEQ ID NO:3) sequence at positions 38-42 to comprise the amino acid sequence of the general formula (I):
TABLE-US-00009 [0163] (SEQ ID NO: 1) X.sub.38-X.sub.39-X.sub.40-X.sub.41-X.sub.42 (I)
[0164] wherein
[0165] X.sub.38 is F, L, I or V;
[0166] X.sub.39 L, I, V or M;
[0167] X.sub.40 is A, G, S, E, Q, Y, F, W, R, K, H, L, I, V or M;
[0168] X.sub.41 is S, Y, F, W, L, I, V or M;
[0169] X.sub.42 is Y, W, L, I, V, M or S;
[0170] with the proviso that X.sub.38-X.sub.42 is not VIGYS (SEQ ID NO:4), [0171] Optionally an extension peptide, and [0172] A GLP-1 peptide. 42. The GLP-1 precursor according to embodiment 41 wherein the peptides comprised by said GLP-1 precursor are fused according to the order in which they are listed, i.e. pre-peptide--Mating Factor .alpha. pro-peptide variant, the optional extension peptide and the GLP-1 peptide. 43. DNA sequence encoding the polypeptide according to any of embodiments 39-42. 44. Expression vector comprising a DNA sequence according to embodiment 43. 45. Expression vector according to embodiment 44, wherein said DNA sequence encoding a polypeptide to be expressed is operatively linked to an upstream promoter and a downstream terminator. 46. Host cell comprising the expression vector according to any of embodiments 44-45. 47. The host cell according to embodiment 46, which is selected from the group consisting of Saccharomyces spp., Pichia spp., Hansenula spp., Arxula spp., Kluyveromyces spp., Yarrowia spp. and Schizosaccharomyces spp.
[0173] All references, including publications, patent applications, and patents, cited herein are hereby incorporated by reference in their entirety and to the same extent as if each reference were individually and specifically indicated to be incorporated by reference and were set forth in its entirety herein (to the maximum extent permitted by law).All headings and sub-headings are used herein for convenience only and should not be construed as limiting the invention in any way. The use of any and all examples, or exemplary language (e.g., "such as") provided herein, is intended merely to better illuminate the invention and does not pose a limitation on the scope of the invention unless otherwise claimed. No language in the specification should be construed as indicating any non-claimed element as essential to the practice of the invention. The citation and incorporation of patent documents herein is done for convenience only and does not reflect any view of the validity, patentability, and/or enforceability of such patent documents. This invention includes all modifications and equivalents of the subject matter recited in the claims appended hereto as permitted by applicable law.
EXAMPLES
Examples 1-45
[0174] We constructed a plasmid, that contains the TDH3 promoter, the gene encoding the MFalpha pre-pro-peptide, the gene encoding an N-terminal extended GLP-1 (DVKPGQPMYDDDDK-GLP-1(7-37)[K34R]) (SEQ ID NO:44), a minimal 2 micron region for maintenance in yeast, and a selectable marker, namely the TPI gene from S. pombe (see FIG. 1).
[0175] In this plasmid background we introduced various single mutations in the region encoding position 38-42 of the MFalpha-pro-peptide region, the X.sub.38-X.sub.42=VIGYL (SEQ ID NO:3) sequence (see FIG. 2). The experiment using the wt VIGYL (SEQ ID NO:3) sequence is Example 1, c.f. tables 1-5. Other reference experiments are Examples 5-6, 10-11, 28-29, 37-38 and 44-45.
[0176] These plasmids were introduced into a Saccharomyces cerevisiae strain lacking the TPI1 gene, allowing for selection of transformants harbouring the plasmid on media containing glucose as sole carbon source. Transformants were thereafter cultivated for 3 days in shake flasks in 5 ml relevant media and the cultivation supernatants were analyzed by LCMS for concentration of secreted GLP-1 peptide, and degree of O-glycosylation of the peptide. In tables 1-5 are shown the yield and O-glycosylation degree of the wild type and of the single amino acid residue mutants of the X.sub.38-X.sub.42 sequence (a couple of reference examples on each position being included).
TABLE-US-00010 TABLE 1 Data from expression of N-terminal extended GLP-1 under control of Mating Factor .alpha. propeptide mutants having a single amino acid mutation in position 38 (X.sub.38). Concentrations of N-terminal extended GLP-1 and O-glyco impurity are normalised against the data from same expression but under control of the Mating Factor .alpha. propeptide having the wildtype Val as X.sub.38. Position 38 % conc. % O-glyco Example Mutation relative to wt relative to wt 1 Val 100 100 2 Ile 92 91 3 Phe 107 103 4 Leu 106 104 5 Asp 33 277 6 Pro 47 354
TABLE-US-00011 TABLE 2 Data from expression of N-terminal extended GLP-1 under control of Mating Factor .alpha. propeptide mutants having a single amino acid mutation in position 39 (X.sub.39). Concentrations of N-terminal extended GLP-1 and O-glyco impurity are normalised against the data from same expression but under control of the Mating Factor .alpha. propeptide having the wildtype Ile as X.sub.39. Position 39 % conc. % O-glyco Example Mutation relative to wt relative to wt 1 Ile 100 100 7 Leu 113 82 8 Val 108 76 9 Met 126 88 10 Glu 40 337 11 His 25 313
TABLE-US-00012 TABLE 3 Data from expression of N-terminal extended GLP-1 under control of Mating Factor .alpha. propeptide mutants having a single amino acid mutation in position 40 (X.sub.40). Concentrations of N-terminal extended GLP-1 and O-glyco impurity are normalised against the data from same expression but under control of the Mating Factor .alpha. propeptide having the wildtype Gly as X.sub.40. Position 40 % conc. % O-glyco Example Mutation relative to wt relative to wt 1 Gly 100 100 12 Ala 84 48 13 Ser 79 89 14 Thr 80 67 15 Asp 51 90 16 Glu 58 92 17 Gln 87 98 18 Tyr 60 49 19 Phe 50 23 20 Trp 53 18 21 Arg 106 14 22 Lys 93 36 23 His 74 83 24 Leu 42 13 25 Ile 18 1 26 Val 26 1 27 Met 60 36 28 Asn 81 112 29 Pro 46 253
TABLE-US-00013 TABLE 4 Data from expression of N-terminal extended GLP-1 under control of Mating Factor .alpha. propeptide mutants having a single amino acid mutation in position 41 (X.sub.41). Concentrations of N-terminal extended GLP-1 and O-glyco impurity are normalised against the data from same expression but under control of the Mating Factor .alpha. propeptide having the wildtype Tyr as X.sub.41. Position 41 % conc. % O-glyco Example Mutation relative to wt relative to wt 1 Tyr 100 100 30 Phe 69 74 31 Trp 79 73 32 Ser 107 182 33 Leu 129 104 34 Ile 111 171 35 Val 134 136 36 Met 131 151 37 Gly 50 366 38 Asp 27 590
TABLE-US-00014 TABLE 5 Data from expression of N-terminal extended GLP-1 under control of Mating Factor .alpha. propeptide mutants having a single amino acid mutation in position 42 (X.sub.42). Concentrations of N-terminal extended GLP-1 and O-glyco impurity are normalised against the data from same expression but under control of the Mating Factor .alpha. propeptide having the wildtype Leu as X.sub.42. Position 42 % conc. % O-glyco Example Mutation relative to wt relative to wt 1 Leu 100 100 39 Ile 98 64 40 Tyr 123 156 41 Trp 116 163 42 Val 103 111 43 Met 109 142 44 Asp 19 525 45 Pro 47 472
Example 46
[0177] A plasmid similar to those in Examples 1-45, containing the TDH3 promoter, the gene encoding the MFalpha pre-pro-peptide, the gene encoding an N-terminal extended GLP-1 (DVKPGQPMYDDDDK-GLP-1(7-37)[K34R]) (SEQ ID NO:44), a minimal 2 micron region for maintenance in yeast, and a selectable marker (FIG. 1) was transformed into two different strain backgrounds--one containing the PMT1 gene (PMT+), the other deleted for the PMT1 gene (PMT-). The PMT1 gene is encoding a protein mannosyltransferase involved in O-glycosylation.
[0178] The MFalpha pro-peptide was either wildtype (40G) or mutated (G40R). The strains were cultivated under identical conditions in continuous cultures, with a dilution rate=0.1, and pH 5.8. Samples were analyzed by HPLC to determine the concentration of GLP-1 precursor in the spent medium and by LCMS to determine the proportion of 0-glycosylated GLP-1 precursor.
[0179] The results demonstrate that the effects indeed are additive.
[0180] The G40R mutation reduced O-glycosylation with more than 80%. Deletion of PMT1 reduces O-glycosylation by further more than 80%, in this case giving levels of O-glycosylation close to the detection limit. Furthermore, the G40R mutation gives a surprising increase in yield of the N-terminally extended GLP-1 as compared to the 40G wildtype version, both in PMT+ strain as well as in pmt1 deleted strain (see Table 6).
[0181] These results demonstrate that the combination of a beneficial mutation in the MFalpha propeptide, e.g. G40R, and a PMT1 deleted host strain results in an additive reduction in O-glycosylation.
TABLE-US-00015 TABLE 6 Normalised maximum concentration (yield) of the peptide DVKPGQPMYDDDDK-GLP-1(7-37)[K34R] (SEQ ID NO. 44) in continuous cultures when using 40R mutated MFalpha pro-peptide as compared to the corresponding wildtype 40G. PMT+ strain PMT- strain 40R 40R (relative to 40G (relative to 40G 40G in same strain) 40G in same strain) 100% 133% 100% 169%
Example 47
[0182] To examine the effect of one of the mutations during expression of other GLP-1 peptides, we constructed plasmids, that contained the TDH3 promoter, the gene encoding the MFalpha pre-pro-peptide in the form of either the wildtype (40G) or a 40R mutation, the gene encoding N-terminal extended GLP-1 (DVKPGQPMYDDDDK-GLP-1(9-37)[K34R] (SEQ ID NO:7) or DVKPGQPMYDDDDK-GLP-1(9-37)[K34R,G37K]) (SEQ ID NO:43), a minimal 2 micron region for maintenance in yeast, and a selectable marker, namely the TPI gene from S. pombe (see FIG. 1).
[0183] These plasmids were introduced into a Saccharomyces cerevisiae strain lacking the TPI1 gene, allowing for selection of transformants harbouring the plasmid on media containing glucose as sole carbon source. Transformants were thereafter cultivated for 3 days in shake flasks in 5 ml relevant media and the cultivation supernatants were analyzed by LCMS for concentration of secreted GLP-1 peptide, and degree of O-glycosylation of the peptide. Table 7 shows the results including the yield of GLP-1 peptide and O-glycosylation degree of GLP-1 peptide when expressed of the wild type (wt) and of the 40R mutation.
TABLE-US-00016 TABLE 7 Data from expression of specific N-terminal extended GLP-1 peptide under control of Mating Factor .alpha. propeptide mutants with or without a single amino acid substitution to Arg in position 40 (X.sub.40). Concentrations of N-terminal extended GLP-1 peptide (yield) and O-glycosylated impurity are normalised against the data from expression of the same GLP-1 peptide but under control of the Mating Factor .alpha. propeptide having the wildtype Gly as X.sub.40. X.sub.40 of MFalpha Yield O-glycosylation pre-pro- (% relative (% relative GLP-1 peptide peptide to wt) to wt) GLP-1(9-37)[K34R] Gly (wt) 100 100 GLP-1(9-37)[K34R] Arg 127 <1 (BDL) GLP-1(9-37)[K34R, G37K] Gly (wt) 100 BDL GLP-1(9-37)[K34R, G37K] Arg 143 BDL BDL: below detection limit
[0184] While certain features of the invention have been illustrated and described herein, many modifications, substitutions, changes, and equivalents will now occur to those of ordinary skill in the art. It is, therefore, to be understood that the appended claims are intended to cover all such modifications and changes as fall within the true spirit of the invention.
Sequence CWU 1
1
4815PRTArtificial SequenceMFalpha propeptide variant
subsequenceMISC_FEATURE(1)..(1)Amino acid residue which is F, L, I
or V.MISC_FEATURE(2)..(2)Amino acid residue which is L, I, V or
M.MISC_FEATURE(3)..(3)Amino acid residue which is A, G, S, E, Q, Y,
F, W, R, K, H, L, I, V or M.MISC_FEATURE(4)..(4)Amino acid residue
which is S, Y, F, W, L, I, V or M.MISC_FEATURE(5)..(5)Amino acid
residue which is Y, W, L, I, V, M or S. 1Xaa Xaa Xaa Xaa Xaa1
5285PRTSaccharomyces cerevisiae 2Met Arg Phe Pro Ser Ile Phe Thr
Ala Val Leu Phe Ala Ala Ser Ser1 5 10 15Ala Leu Ala Ala Pro Val Asn
Thr Thr Thr Glu Asp Glu Thr Ala Gln 20 25 30Ile Pro Ala Glu Ala Val
Ile Gly Tyr Leu Asp Leu Glu Gly Asp Phe 35 40 45Asp Val Ala Val Leu
Pro Phe Ser Asn Ser Thr Asn Asn Gly Leu Leu 50 55 60Phe Ile Asn Thr
Thr Ile Ala Ser Ile Ala Ala Lys Glu Glu Gly Val65 70 75 80Ser Leu
Asp Lys Arg 8535PRTArtificial SequenceSynthetic 3Val Ile Gly Tyr
Leu1 545PRTArtificial SequenceSynthetic 4Val Ile Gly Tyr Ser1
5531PRTHomo sapiens 5His Ala Glu Gly Thr Phe Thr Ser Asp Val Ser
Ser Tyr Leu Glu Gly1 5 10 15Gln Ala Ala Lys Glu Phe Ile Ala Trp Leu
Val Lys Gly Arg Gly 20 25 30631PRTArtificial sequenceAnalogue of
GLP-1(7-37) 6His Ala Glu Gly Thr Phe Thr Ser Asp Val Ser Ser Tyr
Leu Glu Gly1 5 10 15Gln Ala Ala Lys Glu Phe Ile Ala Trp Leu Val Arg
Gly Arg Gly 20 25 30729PRTArtificial sequenceAnalogue of
GLP-1(9-37) 7Glu Gly Thr Phe Thr Ser Asp Val Ser Ser Tyr Leu Glu
Gly Gln Ala1 5 10 15Ala Lys Glu Phe Ile Ala Trp Leu Val Arg Gly Arg
Gly 20 25866PRTArtificial sequenceMFalpha propeptide variant
L42S-MFa(20-85) 8Ala Pro Val Asn Thr Thr Thr Glu Asp Glu Thr Ala
Gln Ile Pro Ala1 5 10 15Glu Ala Val Ile Gly Tyr Ser Asp Leu Glu Gly
Asp Phe Asp Val Ala 20 25 30Val Leu Pro Phe Ser Asn Ser Thr Asn Asn
Gly Leu Leu Phe Ile Asn 35 40 45Thr Thr Ile Ala Ser Ile Ala Ala Lys
Glu Glu Gly Val Ser Leu Asp 50 55 60Lys Arg6595PRTArtificial
SequenceSynthetic 9Glu Glu Ala Glu Lys1 5105PRTArtificial
SequenceSynthetic 10Glu Glu Ala His Lys1 5117PRTArtificial
SequenceSynthetic 11Glu Glu Ala Glu Ala His Lys1 5129PRTArtificial
SequenceSynthetic 12Glu Glu Ala Glu Ala Glu Ala His Lys1
5135PRTArtificial SequenceSynthetic 13Glu Glu Gly His Lys1
5146PRTArtificial SequenceSynthetic 14Glu Glu Gly Glu Pro Lys1
5157PRTArtificial SequenceSynthetic 15Glu Glu Ala His Glu Leu Lys1
5167PRTArtificial SequenceSynthetic 16Glu Glu Ala His Glu Val Lys1
5177PRTArtificial SequenceSynthetic 17Glu Glu Ala His Glu Met Lys1
5187PRTArtificial SequenceSynthetic 18Glu Glu Ala His Glu Phe Lys1
5197PRTArtificial SequenceSynthetic 19Glu Glu Ala His Glu Tyr Lys1
52015PRTArtificial SequenceSynthetic 20Glu Glu Ala His Glu Trp Lys
Glu Glu Gly Asn Thr Thr Pro Lys1 5 10 152111PRTArtificial
SequenceSynthetic 21Glu Glu Leu Asp Ala Arg Leu Glu Ala Leu Lys1 5
10226PRTArtificial SequenceSynthetic 22Gln Pro Met Tyr Lys Arg1
5236PRTArtificial SequenceSynthetic 23Gly Gln Pro Met Tyr Lys1
5246PRTArtificial SequenceSynthetic 24Pro Gly Gln Pro Met Tyr1
5256PRTArtificial SequenceSynthetic 25Lys Pro Gly Gln Pro Met1
5266PRTArtificial SequenceSynthetic 26Leu Lys Pro Gly Gln Pro1
5276PRTArtificial SequenceSynthetic 27Gln Leu Lys Pro Gly Gln1
5286PRTArtificial SequenceSynthetic 28Leu Gln Leu Lys Pro Gly1
5296PRTArtificial SequenceSynthetic 29Trp Leu Gln Leu Lys Pro1
5306PRTArtificial SequenceSynthetic 30His Trp Leu Gln Leu Lys1
5316PRTArtificial SequenceSynthetic 31Trp His Trp Leu Gln Leu1
5326PRTArtificial SequenceSynthetic 32Ala Trp His Trp Leu Gln1
5336PRTArtificial SequenceSynthetic 33Glu Ala Trp His Trp Leu1
5346PRTArtificial SequenceSynthetic 34Ala Glu Ala Trp His Trp1
5356PRTArtificial SequenceSynthetic 35Glu Ala Glu Ala Trp His1
5369PRTArtificial SequenceSynthetic 36Asp Val Lys Pro Gly Gln Pro
Leu Ala1 5379PRTArtificial SequenceSynthetic 37Asp Val Lys Pro Gly
Gln Pro Glu Tyr1 5389PRTArtificial SequenceSynthetic 38Asp Val Lys
Pro Gly Glu Pro Leu Tyr1 5399PRTArtificial SequenceSynthetic 39Asp
Val Lys Pro Gly Gln Pro Leu Tyr1 5409PRTArtificial
SequenceSynthetic 40Asp Val Lys Pro Gly Gln Pro Leu Glu1
5419PRTArtificial SequenceSynthetic 41Asp Val Lys Pro Gly Gln Pro
Met Tyr1 54214PRTArtificial SequenceSynthetic 42Asp Val Lys Pro Gly
Gln Pro Met Tyr Asp Asp Asp Asp Lys1 5 104329PRTArtificial
sequenceAnalogue of GLP-1(9-37) 43Glu Gly Thr Phe Thr Ser Asp Val
Ser Ser Tyr Leu Glu Gly Gln Ala1 5 10 15Ala Lys Glu Phe Ile Ala Trp
Leu Val Arg Gly Arg Lys 20 254445PRTArtificial sequenceExtended
analogue of GLP-1(7-37) 44Asp Val Lys Pro Gly Gln Pro Met Tyr Asp
Asp Asp Asp Lys His Ala1 5 10 15Glu Gly Thr Phe Thr Ser Asp Val Ser
Ser Tyr Leu Glu Gly Gln Ala 20 25 30Ala Lys Glu Phe Ile Ala Trp Leu
Val Arg Gly Arg Gly 35 40 45455PRTArtificial SequenceSynthetic
45Val Ile Asp Tyr Ser1 5465PRTArtificial SequenceSynthetic 46Val
Ala Thr Tyr Leu1 5475PRTArtificial SequenceSynthetic 47Val Ile Gly
Tyr Arg1 5485PRTArtificial SequenceSynthetic 48Ala Ile Gly Tyr Leu1
5
D00001

D00002

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.