Serum Preparation
Masci; Paul ; et al.
U.S. patent application number 16/504033 was filed with the patent office on 2019-12-26 for serum preparation. The applicant listed for this patent is Q-Sera PTY LTD. Invention is credited to John De Jersey, Goce Dimeski, Martin Lavin, Paul Masci, Julie Phillips.
| Application Number | 20190390248 16/504033 |
| Document ID | / |
| Family ID | 45873310 |
| Filed Date | 2019-12-26 |




View All Diagrams
| United States Patent Application | 20190390248 |
| Kind Code | A1 |
| Masci; Paul ; et al. | December 26, 2019 |
SERUM PREPARATION
Abstract
This invention relates to the use of clotting compositions containing prothrombin activators to produce high quality blood serum samples for pathology and other biological assays, and to containers containing such clotting compositions, and related methods of use.
| Inventors: | Masci; Paul; (St. Lucia, AU) ; De Jersey; John; (St. Lucia, AU) ; Lavin; Martin; (St. Lucia, AU) ; Phillips; Julie; (St. Lucia, AU) ; Dimeski; Goce; (Hamilton, AU) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 45873310 | ||||||||||
| Appl. No.: | 16/504033 | ||||||||||
| Filed: | July 5, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 13825047 | Jun 21, 2013 | 10385381 | ||
| PCT/AU2011/001221 | Sep 20, 2011 | |||
| 16504033 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12Q 1/56 20130101; G01N 33/86 20130101 |
| International Class: | C12Q 1/56 20060101 C12Q001/56; G01N 33/86 20060101 G01N033/86 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Sep 20, 2010 | AU | 2010904233 |
Claims
1-9. (canceled)
10. A method of preparing a serum sample for detecting an analyte of interest, the method comprising contacting a blood sample with an isolated or at least partially purified snake venom prothrombin activator, for a time and under conditions sufficient to prepare the serum sample.
11. A method according to claim 10, further comprising separating a blood clot from the serum sample.
12. A method according to claim 11, wherein the blood clot is separated from the serum sample by centrifugation.
13. A method according to claim 11, wherein the blood clot is separated from the serum sample using a physical barrier.
14. A method according to claim 10, wherein the snake venom prothrombin activator is in the form of a dried preparation prior to contacting with the blood sample.
15. A method according to claim 10, wherein the blood sample comprises an anticoagulant.
16. A method according to claim 6, wherein the anticoagulant is heparin, warfarin, citrate or rivaroxaban.
17. A method according to claim 10, wherein the prothrombin activator is a group A prothrombin activator.
18. A method according to claim 8, wherein the prothrombin activator is selected from ecarin and basparin.
19. A method according to claim 10, wherein the prothrombin activator is a group B prothrombin activator.
20. A method according to claim 19, wherein the prothrombin activator is selected from carinactivase-1, carinactivase-2 and multactivase.
21. A method according to claim 10, wherein the prothrombin activator is a group C prothrombin activator.
22. A method according to claim 21, wherein the prothrombin activator is selected from pseutarin C, oscutarin C and omicarin C.
23. A method according to claim 10, wherein the prothrombin activator is a group D prothrombin activator.
24. A method according to claim 23, wherein the prothrombin activator is selected from porpharin D, notecarin D, trocarin D, hopsarin D and notenarin D.
25. A method of detecting an analyte of interest, the method comprising providing a serum sample prepared by a serum preparation method and analysing the serum sample for the presence or amount of the analyte of interest, wherein the serum preparation method comprises contacting a blood sample with an isolated or at least partially purified snake venom prothrombin activator, for a time and under conditions sufficient to prepare the serum sample.
26. A method according to claim 25, wherein the analyte is selected from the group consisting of sodium, potassium, chloride, bicarbonate, glucose, urea, creatine, urate, total protein, albumin, bilirubin, alkaline phosphatase, gamma-glutamyl transferase, alanine aminotransferase, aspartate aminotransferase, lactate dehydrogenase, creatine kinase, calcium, phosphate, magnesium, lipase, cholesterol, triglyceride, high-density lipoprotein cholesterol, iron, transferrin, C reactive protein, cortisol, free thyroxine, thyroid stimulating hormone, ferritin, troponin and free haemoglobin.
27. A method of diagnosing the presence, absence or severity of a disease or condition in a subject, wherein the presence, absence or severity of the disease or condition is associated with the presence, absence or an aberrant amount of an analyte of interest in the subject, the method comprising providing a serum sample prepared by a serum preparation method and detecting the presence, absence or aberrant amount of the analyte in the serum sample to thereby determine the presence, absence or severity of the disease or condition in the subject, wherein the serum preparation method comprises contacting a blood sample with an isolated or at least partially purified snake venom prothrombin activator, for a time and under conditions sufficient to prepare the serum sample.
Description
[0001] This application is a divisional of U.S. patent application Ser. No. 16/504,033, filed Jul. 5, 2019, now U.S. Pat. No. 10,385,381, which is a .sctn. 371 National Entry of International Patent Application PCT/AU2011/001221, filed Sep. 20, 2011, which is incorporated by reference, and which claims priority to Australian Patent Application 20100904233, filed Sep. 20, 2010.
FIELD OF THE INVENTION
[0002] This invention relates generally to using procoagulants to produce high quality blood serum samples for pathology and other biological assays.
BACKGROUND OF THE INVENTION
[0003] Blood collection devices, including tubes, are used to collect blood to produce serum or plasma which is in turn used for biochemical or other pathology assays.
[0004] Serum is produced by allowing the blood sample to clot and then centrifuging the sample to separate the blood clot including cells from the serum. Plastic tubes (in place of glass) are now typically used and require procoagulants (often micronised silica particles) to enhance the clotting process. Serum is usually preferred over plasma for biological testing unless urgent results are required, in which case the clotting time for a serum tube is considered too long. Even with existing procoagulants, in most commercial tubes the minimum required clotting time recommended by manufacturers is 30 minutes for blood samples from normal patients, and much longer (typically 60 minutes or longer) for samples from patients taking anti-clotting therapeutic agents such as warfarin or heparin. For patient samples from emergency situations (emergency departments, intensive care, operating theatres etc.) the time is too long and therefore plasma, which can be produced much faster, is often preferred over serum. An alternative purported to address this issue is a blood collection tube for serum production recently developed by Becton-Dickinson (designated BD Rapid Serum Tube, BDT or BD RST) which contains thrombin designed to increase the rate and extent of blood clotting in blood samples.
[0005] Plasma is formed by collecting blood in tubes containing anticoagulants followed by centrifugation which can be performed immediately after collection to separate the cells and thus obtain plasma for analysis. Lithium heparin is the most commonly used anticoagulant in these tubes. Citrate, sodium fluoride/potassium oxalate and EDTA are other anticoagulants that are used in some tubes to produce plasma for estimation of a small number of other analytes.
[0006] Incomplete Clotting
[0007] The coagulation process in preparing a serum sample consumes fibrinogen and entraps platelets and other cells within a network of fibrin. Upon centrifugation the serum is separated from the clot, either by serum separator in the collection device or by aliquoting the serum into a secondary container, to prevent contact with cells. This separation permits the sample to remain stable for extended periods of time. This stability is particularly important if samples are not analysed immediately, or if re-analysis or additional analyses are required.
[0008] For some serum samples, coagulation is incomplete after the recommended waiting times. This problem of incomplete clotting is especially prevalent in patients on anti-clotting therapy or specimens collected from anticoagulated taps or cannulae. Contamination of the specimen with anticoagulant agents during collection may also occur. Such blood can take much longer than the manufacturer's recommended waiting time to clot, or in fact may never fully clot in a standard serum tube (e.g. blood from cardiac surgery patients who are fully heparinised). If a serum sample is centrifuged before clotting is complete, clotting can continue in the serum, leading to clots, microclots or formation of fibrin strings capable of causing analyser or analyte specific problems. The formation of microclots and fibrinogen strings during sample preparation may also occur in plasma tubes, especially post-storage at low temperatures. Lack of timely inversion of lithium heparin tubes after blood collection can lead to small clot formation around the rubber stopper. Droplets of blood not heparinised in a timely manner will clot, and clots do not disintegrate upon heparinisation.
[0009] Even the smallest clots are capable of producing clinically significant errors. Thus for accuracy, samples must be manually checked by eye or using automated detection systems if available to ensure they are free of fibrin strands or clots. If strands or clots of insoluble material are present, the sample requires sub-sampling into a new container and re-centrifugation prior to test analysis. Samples that exhibit repeated latent clotting may need to be transferred to a lithium heparin tube to stop ongoing clotting. These actions take additional time. Further, fibrin strands or clots are not always detected (e.g. they may even occur post analyser sampling), and consequential sampling errors may lead to patient care decisions being made on inaccurate results.
[0010] Cell Contamination in Plasma Tubes
[0011] Specimens obtained in plasma tubes, lithium heparin plasma specifically, may be contaminated with cells. Lithium heparin gel tubes when centrifuged will always present a small "buffy coat like layer" on top of the gel at the bottom of the plasma. This layer contains fibrin, cells and cell stroma. The rapid gel movement during centrifugation leaves some cells in the plasma. If the plasma specimen is mixed (e.g. during sub-sampling or handling), it will become turbid due to suspension of cell-containing material and fibrin, which decreases the specimen integrity. In addition, platelet aggregates can form which may also contain fibrin and/or white blood cells. These aggregates can be large enough to be visible to the unaided eye and have been termed "white particulate matter" due to their typical white colour, and present similar problems to incomplete clotting discussed above.
[0012] The presence of cells in the sample can affect analyte concentrations. Certain analytes (e.g. glucose) may be decreased by cell activity and others may be increased by leakage or cell lysis (e.g. lactate dehydrogenase, potassium, phosphate).
[0013] Analyte Interference
[0014] Although generally there is no difference in concentration of analytes measured in serum or plasma tubes, there are some exceptions.
[0015] Plasma tubes that use heparin are not suitable for heparin analysis or cell-based assays. Lithium heparin plasma tubes are not suitable for lithium analysis. Plasma may be unreliable for additional testing or re-testing, due to presence of cells and insoluble fibrin formation upon prolonged storage at 2-8.degree. C.
[0016] Further, there have been reports of some serum or plasma tubes producing inaccurate results of analyte levels, due to interaction with the procoagulant or anticoagulant agents within the tubes, or otherwise (Ciuti et al., 1989; Cowley et al., 1985; Davidson et al., 2006; Dimeski et al., 2004; Dimeski et al., 2005; Dimeski et al., 2010; Hartland et al., 1999; Miles et al., 2004; O'Keane et al., 2006; Wannaslip et al., 2006).
[0017] Sample Size
[0018] It is desirable to reduce the sample size needed for testing, especially in critically ill patients, patients receiving blood transfusions, and infants, in order to reduce the volume of blood taken from a patient. It is therefore optimal to be able to run all necessary tests using a sample taken in a single blood collection tube. To achieve this, testing methods have been developed using very small sample volumes (e.g. 2 .mu.L) so that typically one serum or plasma tube is used for at least 21 tests, but can be used for between 50-60 or even 70-80 tests, depending on the volume of sample needed for each test. However, where there is doubt over the accuracy of measuring a particular analyte in a serum or plasma tube, it may be necessary to take both a serum tube and a plasma tube from the patient and doing so defeats the goal of reducing the volume of blood taken from the patient.
[0019] Problems arising from the use of current methodologies for serum and plasma preparation from blood show that improvements are required to achieve timely, reliable analytical results from a wider variety of blood samples generally.
[0020] Snake Venom Prothrombin Activators
[0021] Many snake venoms contain prothrombin activators for the purpose of rapid clotting of the blood of their prey. These prothrombin activators are proteolytic enzymes which convert prothrombin present in blood to thrombin which in turn causes clotting.
[0022] While snake venom prothrombin activators are known procoagulants, they are also known to possess proteolytic trypsin-like activity (Schieck et al., 1972; Parker, H. W. and Grandison A. G. C., 1977; Masci, P. P., 1986; Nicholson et al., 2006; Lavin and Masci, 2009). It has been postulated that there may be an evolutionary reason that prothrombin activators possess both procoagulant and proteolytic properties in that they act to both kill and digest the prey (Masci, P. P., 1986, page 143). For example, ecarin (prothrombin activator purified from Echis carinatus venom) has been shown to have procoagulant activity and as well several other proteolytic activities such as fibrinogenolysis, gelatinolysis, caesionlysis and haemorrhage (Schieck et al., 1972), and a prothrombin activator purified from the venom of Pseudonaja textilis (PtPA) is active against a range of chromogenic peptide substrates designed for different proteolytic enzymes (Masci, P. P., 1986).
[0023] Many analyte tests that may be performed on blood, serum, or plasma samples involve proteins, including tests measuring proteins as analytes (e.g. total protein, albumin); tests measuring enzyme activity of blood proteins (e.g. gamma-glutamyl transpeptidase used in test for gamma-glutamyl transferase, aspartate aminotransferase, lactate dehydrogenase, creatine kinase, lipase); tests using proteins as reagents (e.g. immunoassays); tests using enzymes in the analytical method (e.g. glucose oxidase). Other commonly used tests involving protein include assays for glucose, urea, urate, alanine aminotransferase, creatine kinase, high-density lipoprotein cholesterol, cholesterol, triglycerides, transferrin, C reactive protein, troponin, cortisol, free thyroxine, free triiodothyronine, thyroid stimulating hormone, and ferritin.
[0024] Therefore, despite their procoagulant properties, these snake venom prothrombin activators have never been considered suitable for use in serum tubes for analyte tests, on the basis that their proteolytic activity would degrade analytes being measured (e.g. where the analyte is a protein), or would degrade proteins being used in the reaction to measure analyte levels (e.g. where the analyte test involves use of a protein such as glucose oxidase).
[0025] Thrombin Tubes
[0026] While thrombin-containing tubes have recently become available as `faster` clotting tubes, and thrombin possesses both procoagulant and proteolytic activity, thrombin is known to have high specificity for cutting bonds in fibrinogen, activated protein C (APC) and Factor Va. Therefore, unlike the reported trypsin-like activity of the snake venom prothrombin activators, thrombin would not be expected to interfere with analyte tests.
[0027] In work leading up to the present invention, it was found that thrombin-containing tubes cannot be used with all blood samples. Thrombin is known to be rapidly and completely inhibited by the heparin-antithrombin III complex present in heparinised blood samples. In investigating the BD RST tubes, it was found that these tubes are ineffective in clotting patient samples containing high doses of heparin (Dimeski et al., 2010).
[0028] Development of the Invention
[0029] Surprisingly, the present inventors discovered that when used in blood collection devices, including tubes, prothrombin activators are generally capable of producing high quality serum in an acceptable time from a wide variety of blood samples (including those taken from patients on high concentration of anti-clotting therapy, including heparin), decreasing both the serum sample preparation time and the risk of analysis problems due to incomplete clotting and contamination by cells and cell components.
[0030] Moreover, the inventors also surprisingly discovered that serum samples obtained from blood samples by addition of prothrombin activators give the same results in a wide range of standard biochemistry analytical tests as serum samples produced in existing blood collection tubes.
[0031] These discoveries suggested that prothrombin activators would be suitable for producing serum for the purpose of measuring a wide range of analytes, and have been reduced to practice in blood collection containers for preparing serum samples useful in detecting analytes, related uses and methods, as described hereafter.
SUMMARY OF THE INVENTION
[0032] Accordingly, in one aspect the present invention provides the use of a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator in the preparation of a serum sample that is suitable for detecting an analyte.
[0033] The prothrombin activator (sometimes known as prothrombinase) suitably exhibits trypsin-like activity and activates prothrombin (i.e. converts prothrombin to thrombin).
[0034] The present invention also provides a container for preparing a serum sample that is suitable for detecting an analyte of interest that is present in the sample, wherein the container contains a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator as defined herein.
[0035] In another aspect, the present invention provides the use of a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator as defined herein in the preparation or manufacture of a container for preparing a serum sample suitable for detecting an analyte. In another aspect, the present invention provides a container comprising a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator as defined herein and a blood sample, for preparing a serum sample suitable for detecting an analyte.
[0036] In another aspect the present invention provides a method of preparing a serum sample for detecting an analyte of interest, the method comprising contacting a blood sample with a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator as defined herein for a time and under conditions sufficient to prepare a serum sample. Suitably, the method is carried out in a container as broadly defined above. Suitably, the blood is contacted with the clotting composition for a time and under conditions sufficient to prepare a serum sample and clotted cells. Suitably, the method further comprises separating the serum sample from the clotted cells. In some embodiments the method comprises mixing the clotting composition and blood sample by providing a container containing the blood sample and adding the clotting composition to the container, or providing a container containing the clotting composition and adding or collecting the blood sample into the container.
[0037] The present invention also provides a serum sample produced by contacting a blood sample with a clotting composition as broadly described above for a time and under conditions sufficient to produce the serum sample.
[0038] The present invention further provides methods of detecting an analyte of interest. These methods generally comprise analysing a serum sample prepared by the method of the present invention for the presence or amount of the analyte of interest.
[0039] The present invention also provides methods of diagnosing the presence, absence or severity of a disease or condition in a subject, wherein the presence, absence or severity of the disease or condition is associated with the presence, absence or an aberrant amount of an analyte of interest in the subject. These methods generally comprise providing a serum sample prepared according to the methods broadly described above; and detecting the presence, absence or aberrant amount of the analyte in the serum sample to thereby determine the presence, absence or severity of the disease or condition in the subject.
BRIEF DESCRIPTION OF THE SEQUENCES
[0040] A brief description of the sequences in the sequence listing is provided below.
TABLE-US-00001 SEQUENCE ID NUMBER SEQUENCE SEQ ID NO: 1 Polypeptide sequence for ecarin from Echis carinatus SEQ ID NO: 2 Partial polypeptide sequence for basparin from Bothrops asper venom SEQ ID NO: 3 Partial polypeptide sequence for carinactivase-1 from Echis carinatus venom (prepared as described in Yamada, D., et al., (1996))-62 kDa subunit SEQ ID NO: 4 Partial polypeptide sequence for multactivase from Echis multisquamatus venom (prepared as described in Yamada, D., et al., (1997)) SEQ ID NO: 5 Nucleotide sequence encoding Factor V-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 6 Nucleotide sequence encoding Factor V-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 7 Polypeptide sequence for Factor V-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 8 Polypeptide sequence for Factor V-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 9 Nucleotide sequence encoding Factor V-like component of OsPA (or oscutarin C) from Oxyuranus scutellatus SEQ ID NO: 10 Nucleotide sequence encoding Factor V-like component of OsPA (or oscutarin C) from Oxyuranus scutellatus scutellatus SEQ ID NO: 11 Polypeptide sequence for Factor V-like component of OsPA (or oscutarin C) from Oxyuranus scutellatus scutellatus SEQ ID NO: 12 Polypeptide sequence for Factor V-like component of OsPA (or oscutarin C) from Oxyuranus scutellatus SEQ ID NO: 13 Polypeptide sequence for Factor V-like component of OsPA (or oscutarin C) from Oxyuranus scutellatus SEQ ID NO: 14 Nucleotide sequence encoding Factor V-like component of omicarin C from Oxyuranus microlepidotus SEQ ID NO: 15 Nucleotide sequence encoding factor V from Homo sapiens SEQ ID NO: 16 Polypeptide sequence for factor V from Homo sapiens SEQ ID NO: 17 Nucleotide sequence encoding factor V from Bos taurus SEQ ID NO: 18 Polypeptide sequence for factor V from Bos taurus SEQ ID NO: 19 Nucleotide sequence encoding Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 20 Nucleotide sequence encoding Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 21 Nucleotide sequence encoding Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 22 Nucleotide sequence encoding Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis textilis SEQ ID NO: 23 Nucleotide sequence encoding Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis textilis SEQ ID NO: 24 Nucleotide sequence encoding Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 25 Nucleotide sequence encoding Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 26 Polypeptide sequence for Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 27 Polypeptide sequence for Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 28 Polypeptide sequence for Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis textilis SEQ ID NO: 29 Polypeptide sequence for Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis textilis SEQ ID NO: 30 Polypeptide sequence for Factor X-like component of PtPA (or pseutarin C) from Pseudonaja textilis SEQ ID NO: 31 Nucleotide sequence encoding Factor X-like component of OsPA (or oscutarin C) from Oxyuranus scutellatus SEQ ID NO: 32 Polypeptide sequence for Factor X-like component of OsPA (or oscutarin C) from Oxyuranus scutellatus SEQ ID NO: 33 Nucleotide sequence encoding Factor X-like component of omicarin C from Oxyuranus microlepidotus SEQ ID NO: 34 Polypeptide sequence for Factor X-like component of omicarin C from Oxyuranus microlepidotus SEQ ID NO: 35 Nucleotide sequence encoding Factor X-like component of porpharin D from Pseudechis porphyriacus SEQ ID NO: 36 Polypeptide sequence for Factor X-like component of porpharin D from Pseudechis porphyriacus SEQ ID NO: 37 Nucleotide sequence encoding Factor X-like component of hopsarin D from Hoplocephalus stephensii SEQ ID NO: 38 Polypeptide sequence for Factor X-like component of hopsarin D from Hoplocephalus stephensii SEQ ID NO: 39 Nucleotide sequence encoding Factor X-like component of notecarin D from Notechis scutatus SEQ ID NO: 40 Polypeptide sequence for Factor X-like component of notecarin D from Notechis scutatus SEQ ID NO: 41 Nucleotide sequence encoding Factor X-like component of trocarin D from Tropidechis carinatus SEQ ID NO: 42 Polypeptide sequence for Factor X-like component of trocarin D from Tropidechis carinatus SEQ ID NO: 43 Nucleotide sequence encoding Factor X-like component of prothrombin activator from Demansia vestigiata SEQ ID NO: 44 Polypeptide sequence for Factor X-like component of prothrombin activator from Demansia vestigiata SEQ ID NO: 45 Nucleotide sequence encoding Factor X-like component of prothrombin activator from Demansia vestigiata SEQ ID NO: 46 Polypeptide sequence for Factor X-like component of prothrombin activator from Demansia vestigiata SEQ ID NO: 47 Nucleotide sequence encoding factor X from Homo sapiens SEQ ID NO: 48 Polypeptide sequence for factor X from Homo sapiens SEQ ID NO: 49 Nucleotide sequence encoding factor X from Bos taurus SEQ ID NO: 50 Polypeptide sequence for factor X from Bos taurus SEQ ID NO: 51 Partial polypeptide sequence for carinactivase-1 from Echis carinatus venom (prepared as described in Yamada, D., et al., (1996))-17 kDa subunit SEQ ID NO: 52 Partial polypeptide sequence for carinactivase-1 from Echis carinatus venom (prepared as described in Yamada, D., et al., (1996))-14 kDa subunit
BRIEF DESCRIPTION OF THE DRAWINGS
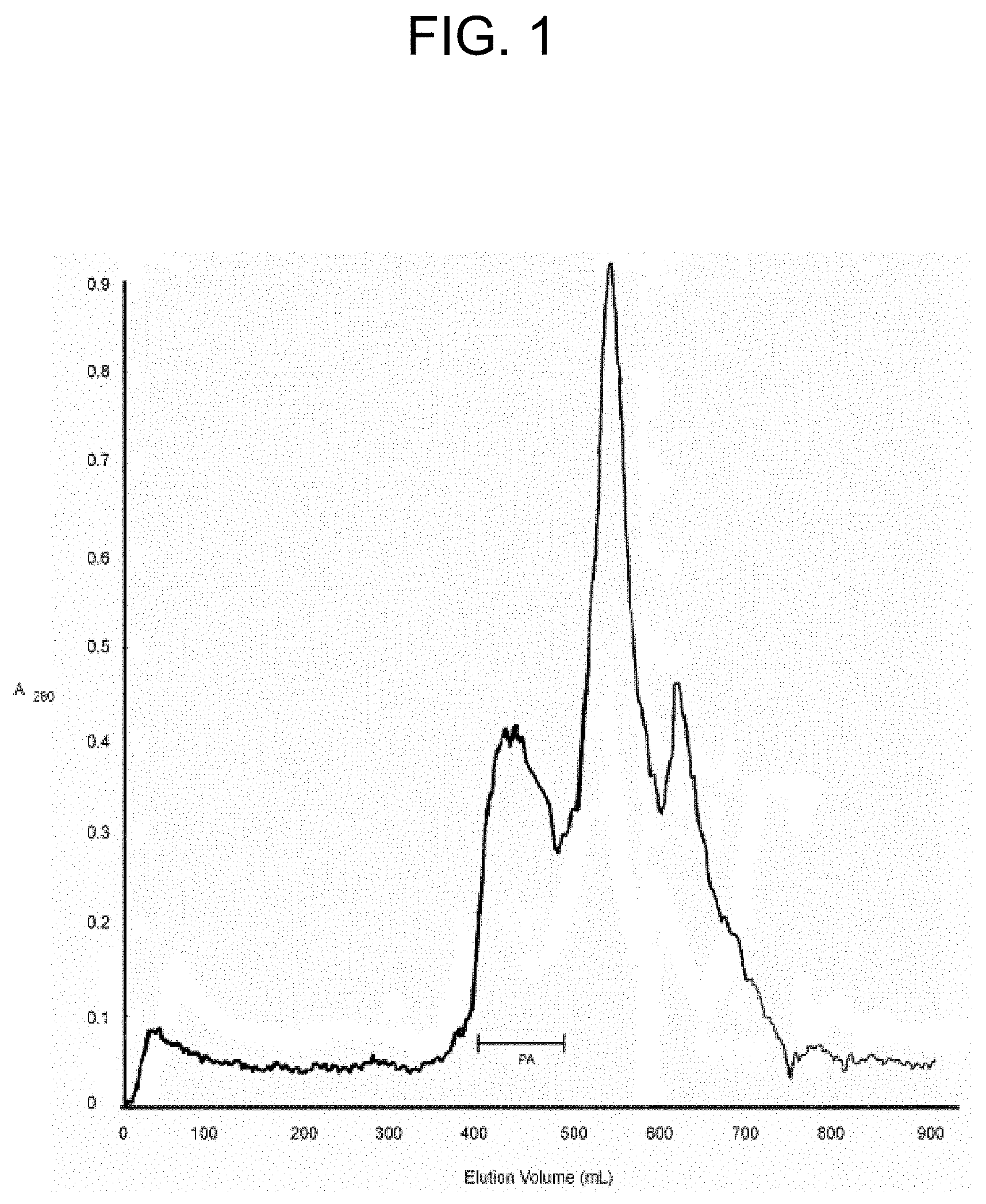
[0041] FIG. 1 shows the elution profile in the isolation of ecarin, carinactivase-1, and carinactivase-2 from E. carinatus venom using gel filtration on a Superdex 200 column as described in Example la. The E. carinatus venom (157 mg, 122 A.sub.280 units) was subjected to gel filtration on Superdex 200 (2.5.times.95 cm) using of 0.05 M Tris-HCl buffer at pH 8.0. The pooled fractions (28.3 A.sub.280 units) of the three prothrombin activators (ecarin, carinactivase-1, and carinactivase-2) are indicated by the bar.
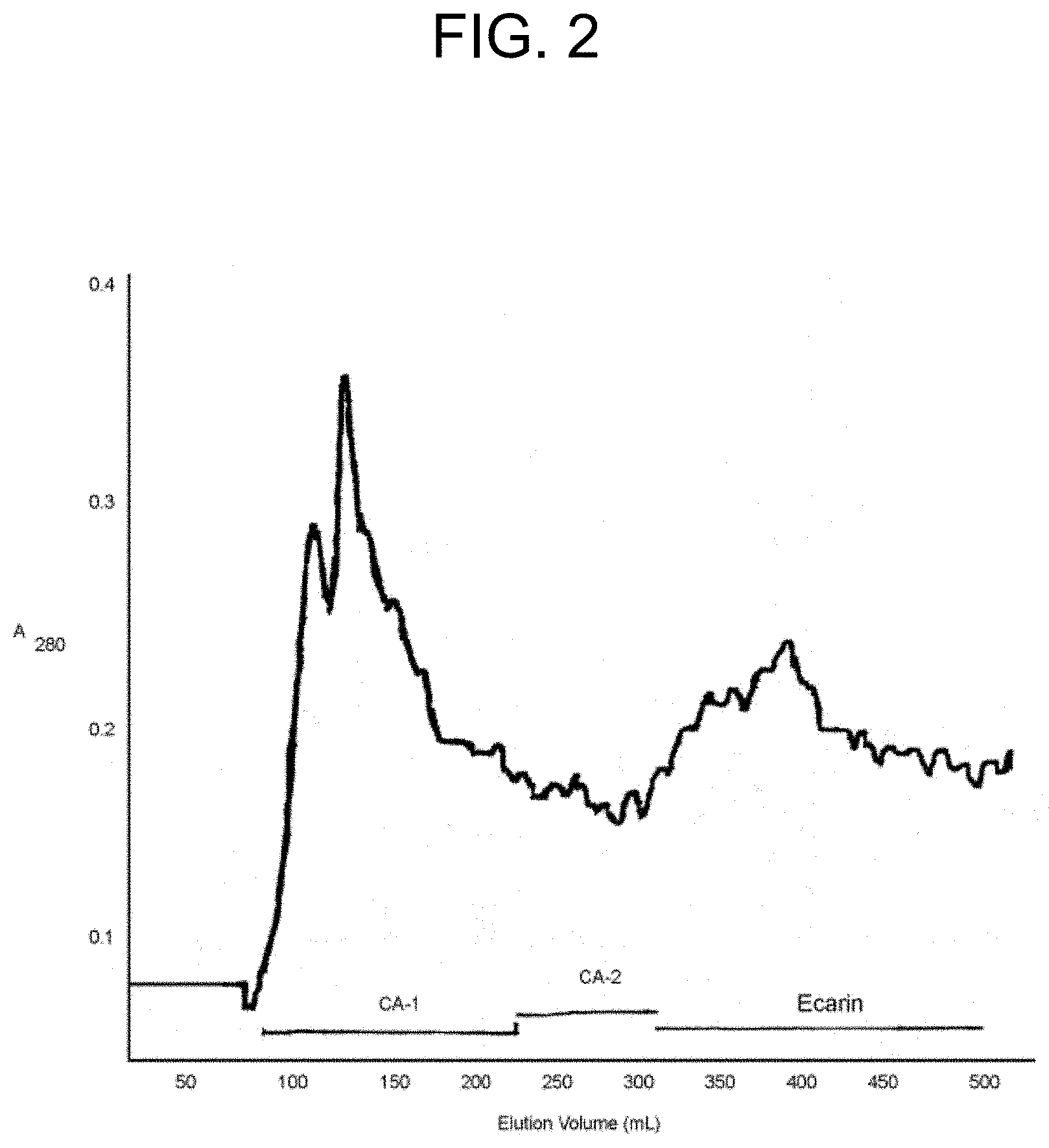
[0042] FIG. 2 shows the elution profile of the Blue Sepharose chromatography of the active (procoagulant) fractions from the Superdex 200 gel filtration chromatography described in Example la (also shown in FIG. 1). The active fractions were subjected to a column of Blue Sepharose and eluted with a linear gradient of NaCl. Three mL fractions were collected. The fractions indicated by the bars were pooled as carinactivase-1, carinactivase-2, and ecarin, respectively.
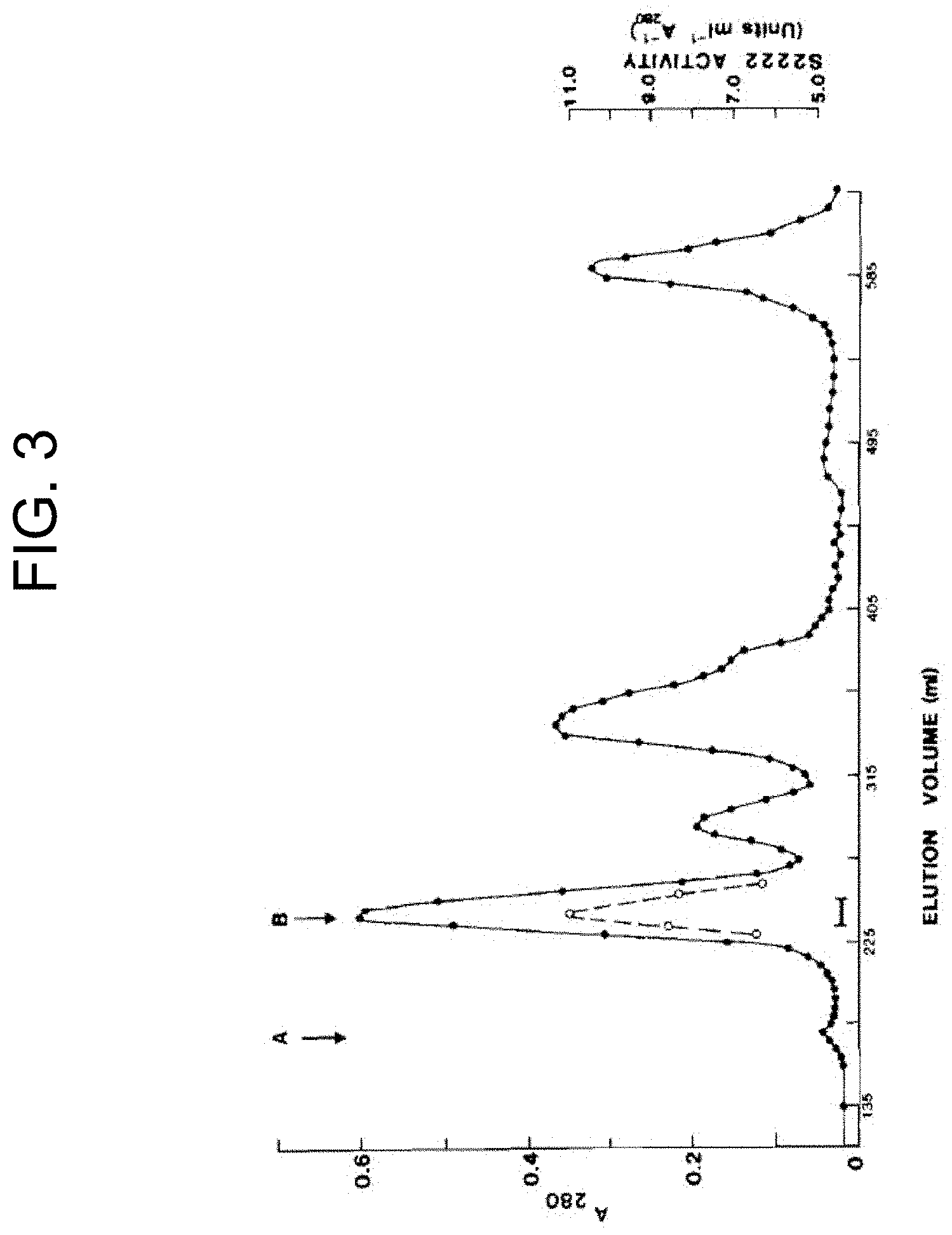
[0043] FIG. 3 shows the elution profile for chromatography of reconstituted P. textilis venom (50 mg in 5 mL) on a column (2.5.times.95 cm) of Sephacryl S-300 in 0.05 Tris-HCl buffer, pH 7.4; 4.degree. C.; flow rate, 17 mL/h; A.sub.280 (.cndot.); S-2222 specific activity (.degree.); `A` and `B` represent the void volume (167 mL) and the elution volume of the peak of S-2222 activity (250 mL) respectively, as described in Example 1b.
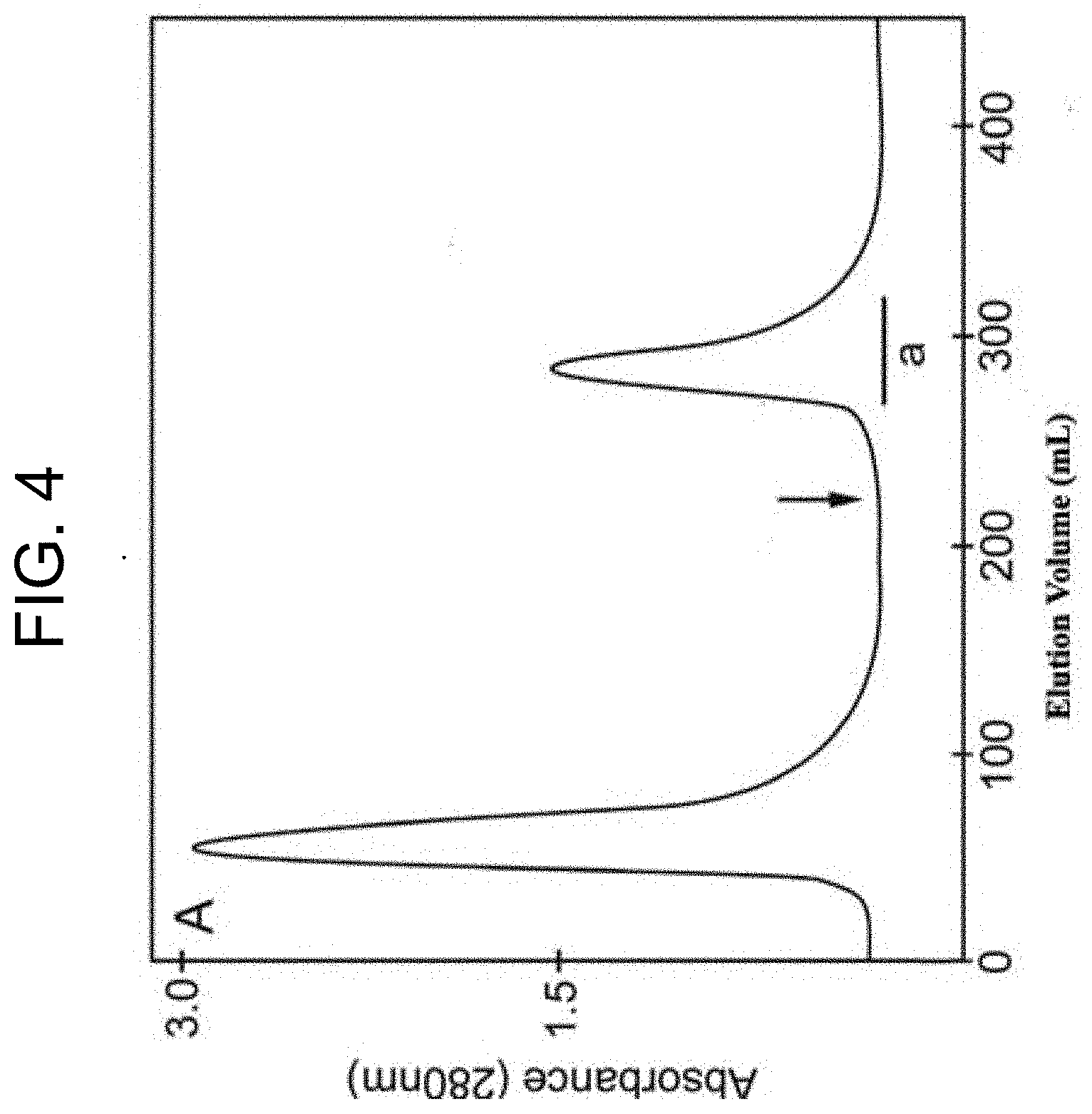
[0044] FIG. 4 shows the elution profile of P. textilis venom (1 g:30 mL) using the Con A-Sepharose affinity chromatography method described in Example 1b. The arrow denotes the position of the application of 0.2 M methyl .alpha.-D-mannopyranoside to elute PtPA (labelled "a").
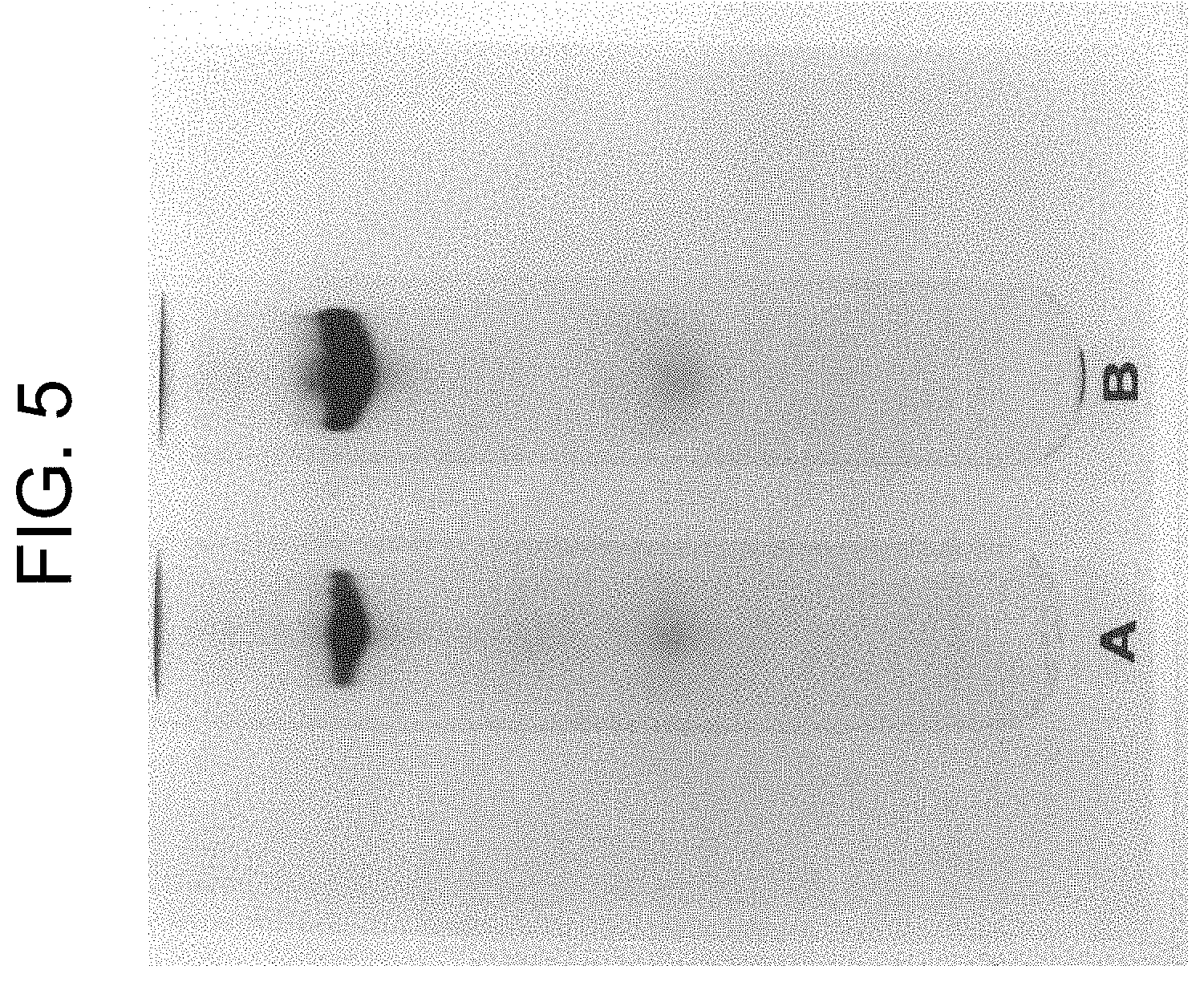
[0045] FIG. 5 shows the results of native PAGE at pH 8.6 of pooled fractions labelled "a" in FIG. 4 where lane A is 25 .mu.g and lane B is 50 .mu.g.
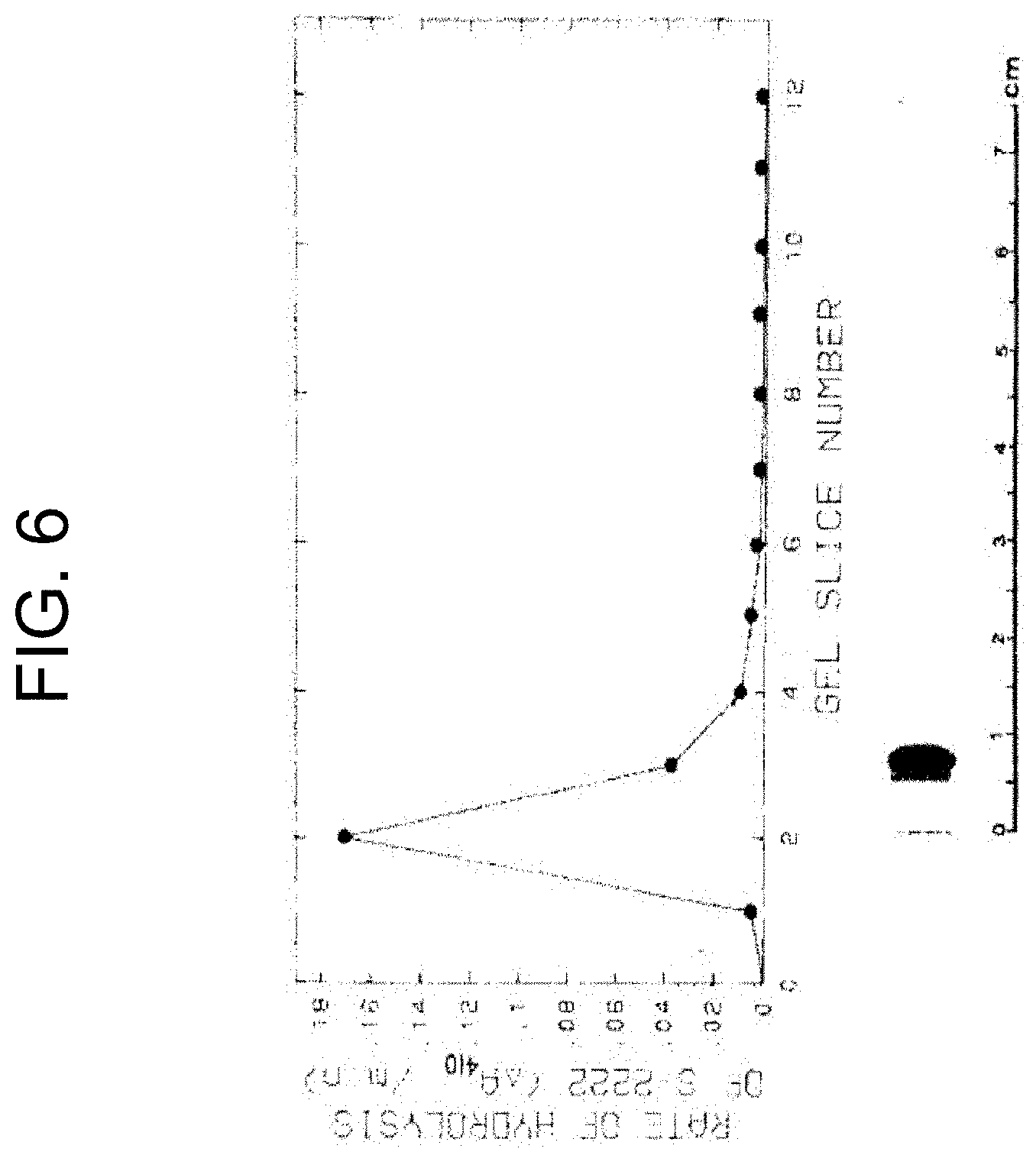
[0046] FIG. 6 shows the results of native PAGE at pH 8.6 of purified PtPA where the gel was stained with Coomassie blue and a duplicate gel was cut into 5 mm slices which were each equilibrated with 1 mL of S-2222 assay mixture to locate the activity, as described in Example 1b. The graph shows a plot of S-2222 activity (Y axis reads: "RATE OF HYDROLYSIS OF S-222 (.DELTA.A.sub.410/min)) against gel slice number (X axis reads: "GEL SLICE NUMBER").
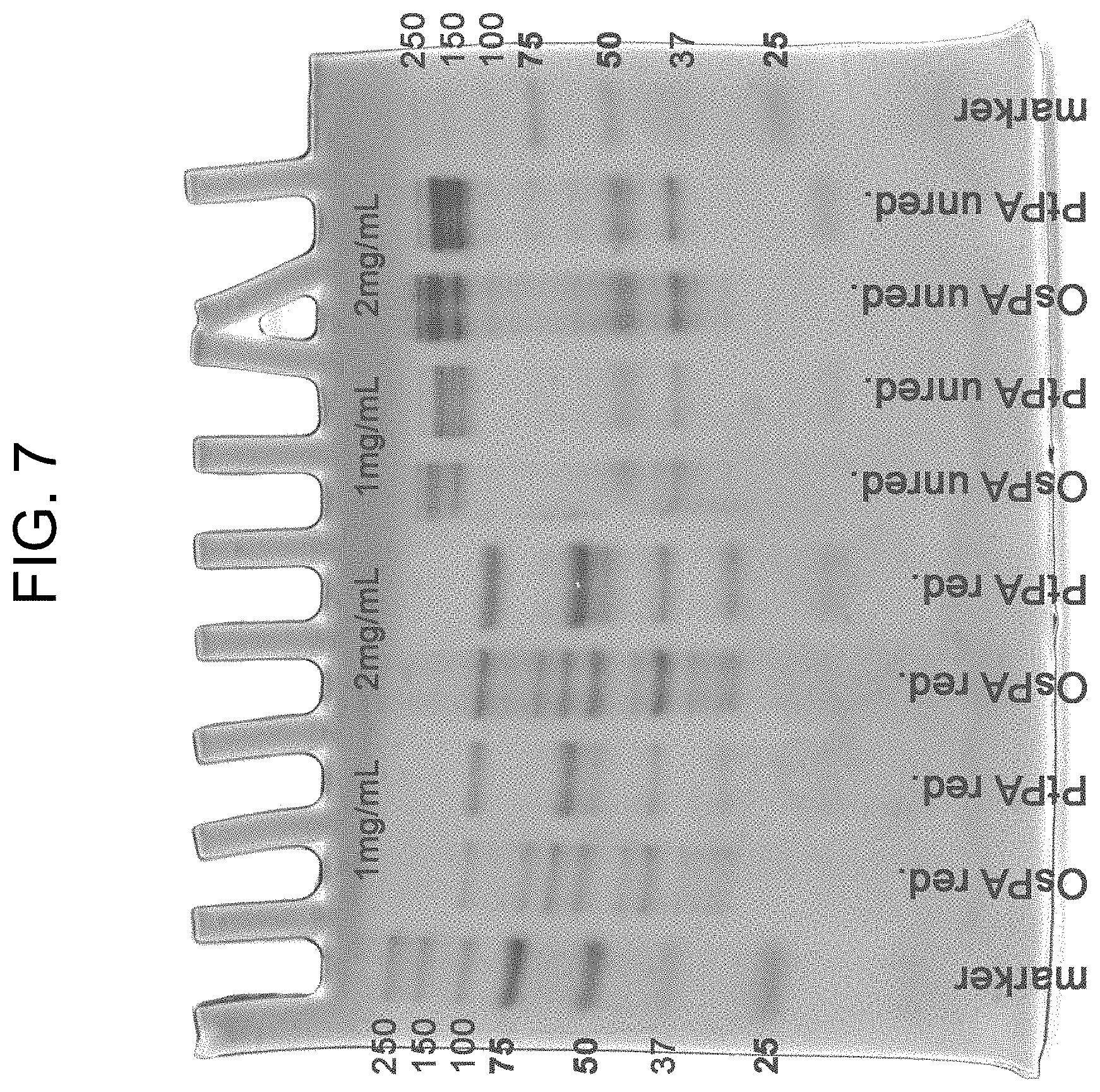
[0047] FIG. 7 shows the SDS-PAGE results of the affinity purified preparations of PtPA and OsPA under reducing and non-reducing conditions as described in Example 1b where the lanes are (from left to right): marker; OsPA red. (1 mg/mL); PtPA red. (1 mg/mL); OsPA red. (2 mg/mL); PtPA red. (2 mg/mL); OsPA unred. (1 mg/mL); PtPA unred. (1 mg/mL); OsPA unred. (2 mg/mL); PtPA unred. (2 mg/mL); marker; where "red." designates the components in the presence of .beta.-mercaptoethanol (i.e. reduced) and "unred." designates the components in the absence of .beta.-mercaptoethanol (i.e. unreduced).
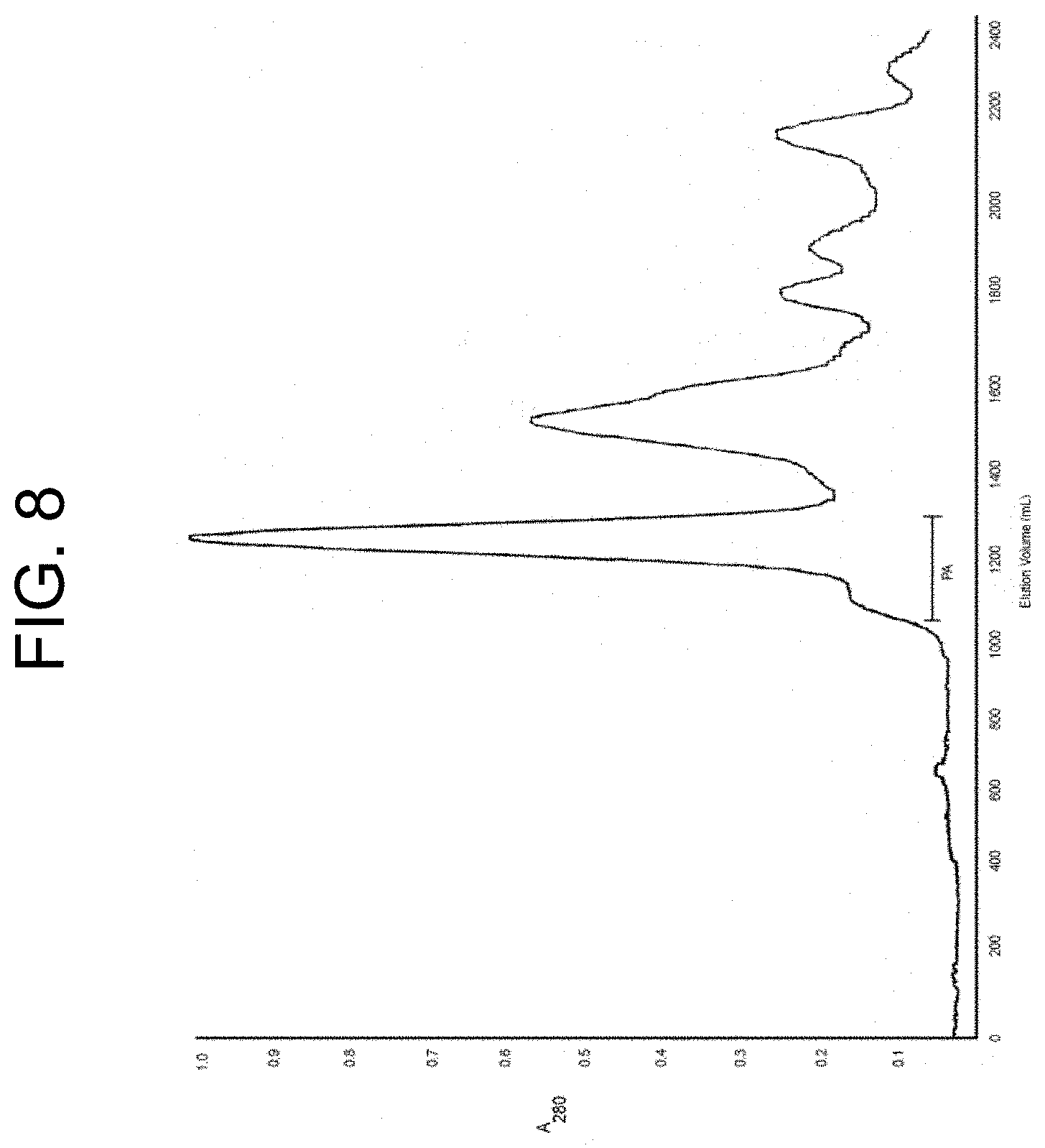
[0048] FIG. 8 shows the elution profile in the isolation of notecarin from N. scutatus venom using Sephacryl S-300 chromatography as described in Example 1c. The pooled fractions of notecarin are indicated by the bar labelled "PA".
[0049] FIG. 9 shows the results of native PAGE at pH 8.9 of the prothrombin activators: carinactivase-1, carinactivase-2, ecarin, PtPA, OsPA, and notecarin, prepared in Examples 1a, 1b, and 1c. In this Figure, the labels represent the following: (I) prothrombin; (II) alpha-thrombin; (III) ecarin; (IV) carinactivase-1; (V) carinactivase-2; (VI) PtPA; (VII) OsPA; (VIII) notecarin (20 .mu.g of each prothrombin activator was loaded).
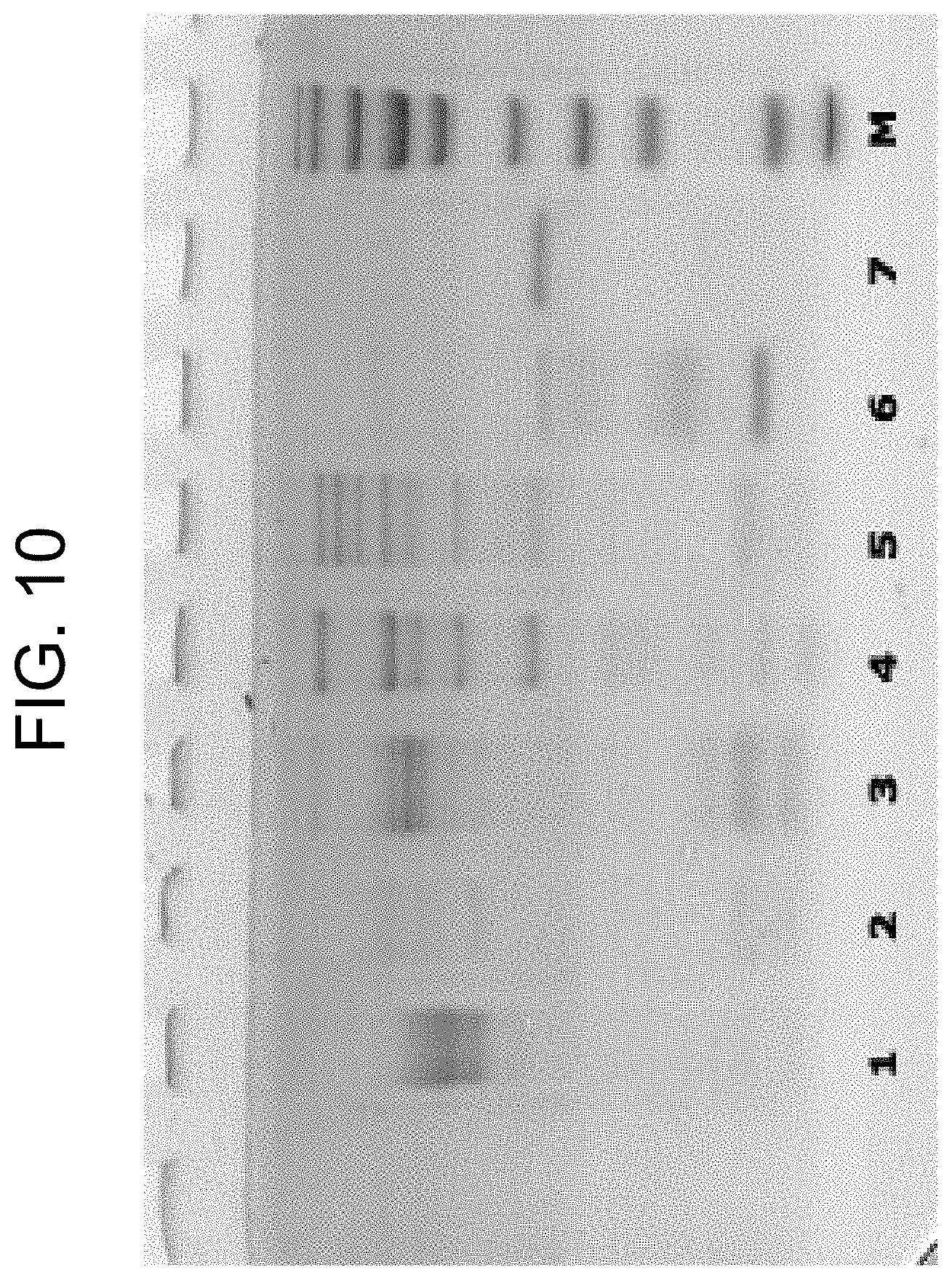
[0050] FIG. 10 shows SDS-PAGE characterisation of the prothrombin activators prepared in Examples 1a, 1b, and 1c, in the presence of .beta.-mercaptoethanol, where the lanes are as follows: (1) carinactivase-1; (2) carinactivase-2; (3) ecarin; (4) PtPA; (5) OsPA; (6) notecarin; (7) thrombin; and (M) molecular weight marker.
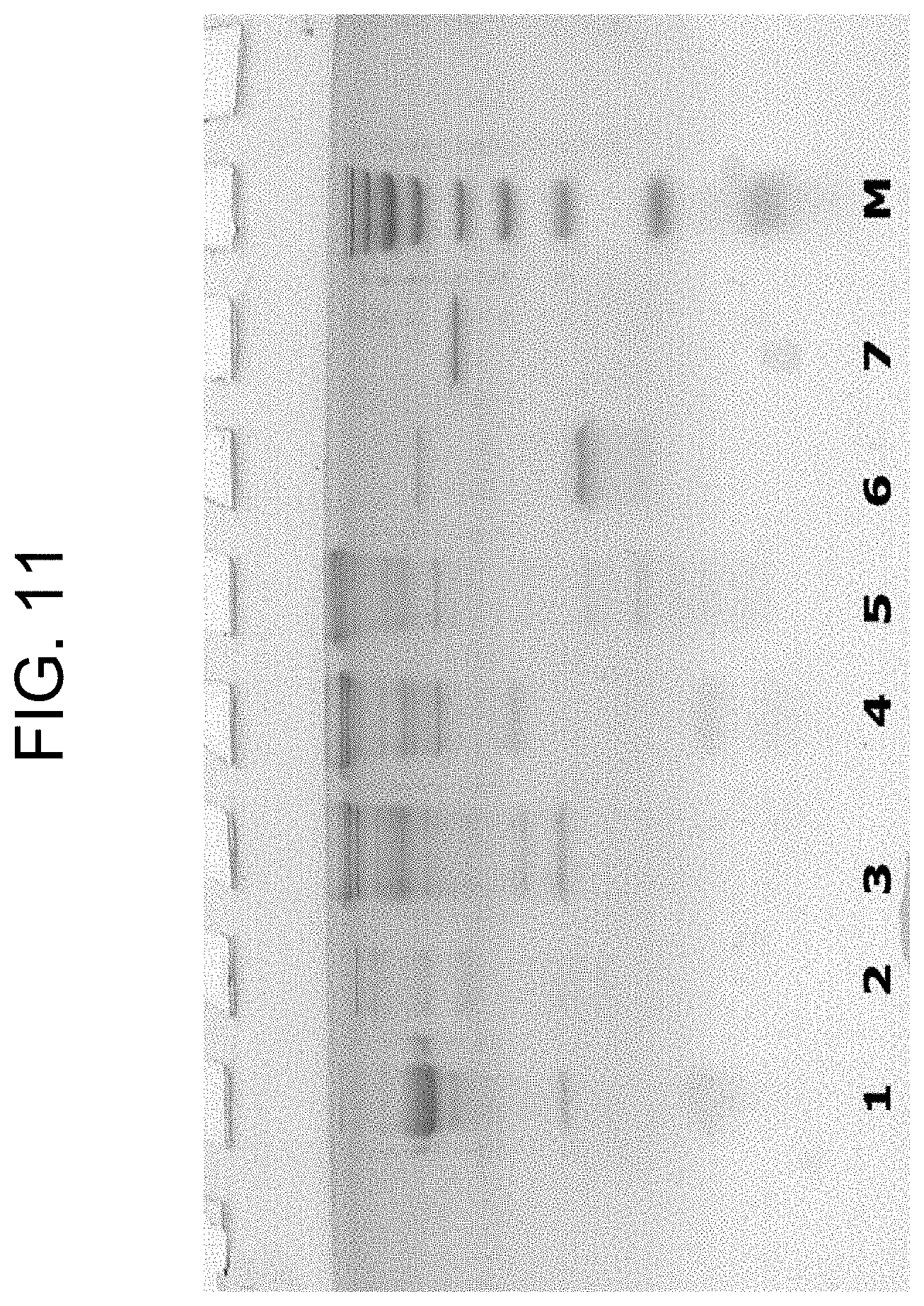
[0051] FIG. 11 shows SDS-PAGE characterisation of the prothrombin activators prepared in Examples 1a, 1b, and 1c, in the absence of .beta.-mercaptoethanol, where the lanes are as follows: (1) carinactivase-1; (2) carinactivase-2; (3) ecarin; (4) PtPA; (5) OsPA; (6) notecarin; (7) thrombin; and (M) molecular weight marker.
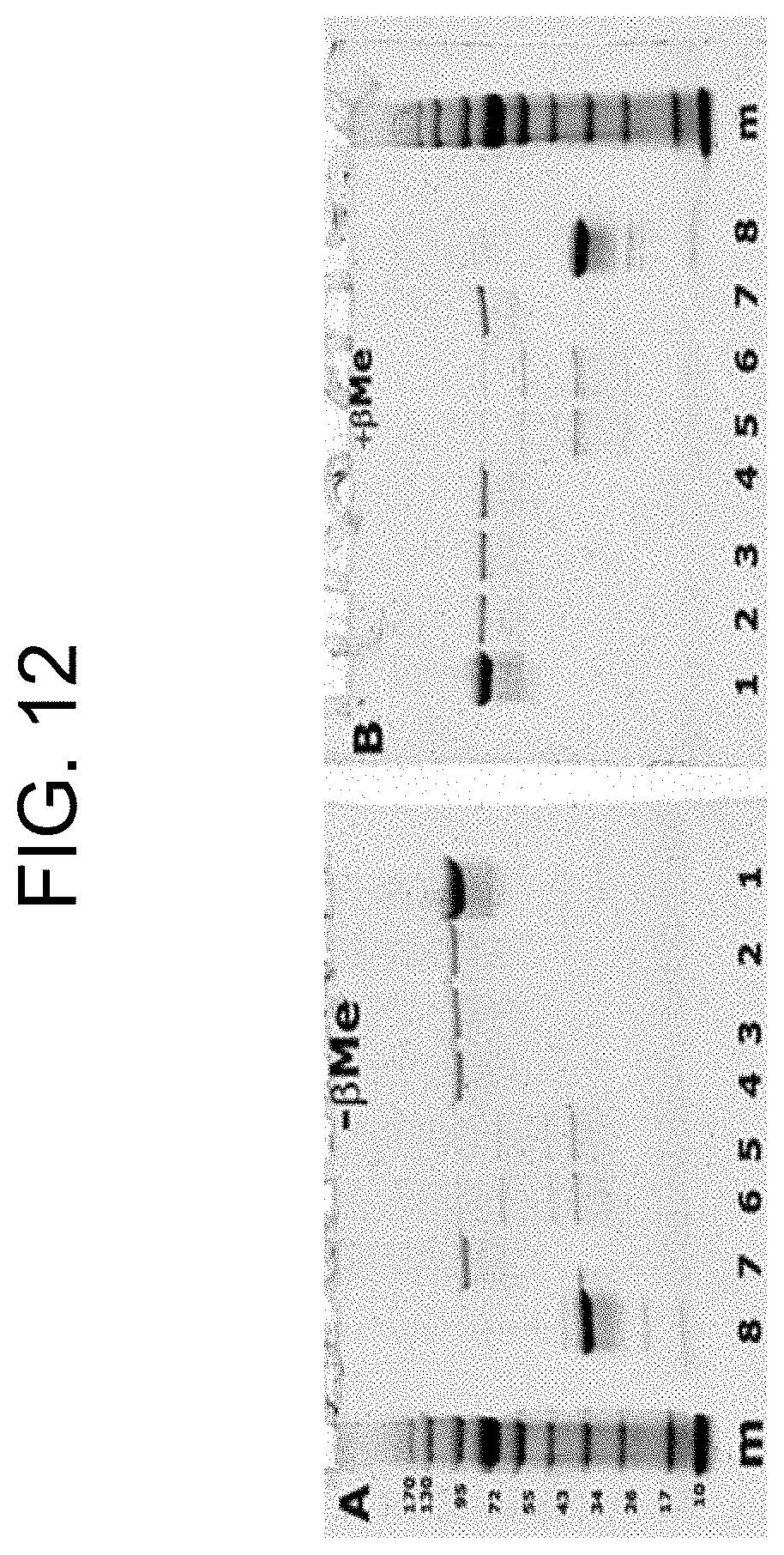
[0052] FIG. 12, panels A and B show the SDS-PAGE without .beta.-mercaptoethanol (A) and with .beta.-mercaptoethanol (B) of samples incubated for 5 minutes at room temperature in the presence of 5 mM Ca.sup.2+, and with the following prothrombin activators: carinactivase-1 (lane 2); carinactivase-2 (lane 3); ecarin (lane 4); PtPA (lane 5); OsPA (lane 6); notecarin (lane 7). Lane 1 contained the sample of human prothrombin alone in buffer (no prothrombin activator), and lane 8 contained the sample of highly purified a-thrombin (no prothrombin activator), and "m" represents the molecular weight marker. This experiment is described in more detail in Example 2a.
[0053] FIG. 13 shows the SDS-PAGE of the time course of the prothrombin (14 .mu.M) to thrombin activation by PtPA (6 nM) and by notecarin (6 nM), as described in Example 2b.
[0054] FIG. 14 is an annotated image of part of the SDS-PAGE results in FIG. 13, where selected bands were eluted and subjected to N-terminal sequencing using mass-spectrometry analysis in order to assign to specific molecular domains.
[0055] FIG. 15 is a graph of the absorbance of the pNA generated from S-2238 by different thrombin concentrations over 155 seconds as described in Example 2d. The calculations on the right hand side of the graph align with each line of the graph, for example the top calculation is the highest line, and so on.
[0056] FIG. 16 is a graph of the standard curve derived from the results in FIG. 15, specifically the slopes of the equations for each reaction up to 180 seconds in FIG. 15 were plotted against the thrombin concentration to provide the linear regression equation shown in FIG. 16.
[0057] FIG. 17 is a graph showing the rate of thrombin generation by 0.6 nM concentrations of the prothrombin activators: PtPA, OsPA, ecarin (Ecr), notecarin (Ntcr), carinactivase-1 (CA-1), and carinactivase-2 (CA-2) over 180 seconds as described in Example 2d.
[0058] FIG. 18 shows the curve fitting analysis of the thrombin catalysed hydrolysis of S-2238 as described in Example 2e. Thrombin concentrations are indicated on progress curves.
[0059] FIG. 19 plots the PtPA activation of prothrombin at the concentrations of PtPA as described in Example 2e.
[0060] FIG. 20 plots the OsPA activation of prothrombin at the concentrations of OsPA at the concentrations of OsPA as described in Example 2e.
[0061] FIG. 21 plots the ecarin activation of prothrombin at the concentrations of ecarin as described in Example 2e.
[0062] FIG. 22 plots the carinactivase-1 activation of prothrombin at the concentrations of carinactivase-1 as described in Example 2e.
[0063] FIG. 23 plots the carinactivase-2 activation of prothrombin at the concentrations of carinactivase-2 as described in Example 2e.
[0064] FIG. 24 plots the notecarin activation of prothrombin at the concentrations of notecarin as described in Example 2e.
[0065] FIG. 25 shows an example of a TEG plot (clot formation part) with the clotting parameters labelled, as discussed in the Examples section.
[0066] FIG. 26 plots the progress curves for hydrolysis of S-2238 by different thrombin concentrations in duplicate from 45-225 nM over a 5 minute incubation period in normal citrated plasma as described in Example 3g.
[0067] FIG. 27 shows the thrombin standard curve derived from the slope of the reactions in FIG. 26 at the 5 minute incubation mark in logarithmic form, where the x-axis shows the slope ln (absorbance units/minute); and the y-axis shows ln (molar concentration of thrombin).
[0068] FIG. 28 shows assays of thrombin remaining after clot removal in sera generated by three different PtPA concentrations (in duplicate), as described in Example 3g.
[0069] FIG. 29 shows assays of thrombin remaining after clot removal in sera generated by three different OsPA concentrations (in duplicate), as described in Example 3g.
[0070] FIG. 30 plots the slopes from FIG. 28 (proportional to thrombin concentrations) against PtPA concentration, as described in Example 3g.
[0071] FIG. 31 plots the slopes from FIG. 29 (proportional to thrombin concentrations) against OsPA concentration, as described in Example 3g.
[0072] FIG. 32 shows progress curves for the hydrolysis of S-2238 by thrombin remaining in serum after clot removal in the presence of 4300 nM (10 IU) heparin, as described in Example 3g. Thrombin was generated by PtPA and OsPA, or was added.
[0073] FIG. 33 shows the reaction curves with samples containing different heparin concentrations using 1.5 nM PtPA as described in Example 3g.
[0074] FIG. 34 shows the reaction curves with samples containing different heparin concentrations using 1.5 nM OsPA as described in Example 3g.
[0075] FIG. 35 shows the changes of absorbances over the 5 minute period in pooled "normal" citrated plasma, with each line representating a BD RST tube filled with a different volume of normal pooled citrate plasma, as described in Example 3h.
[0076] FIG. 36 shows the relationship between the measured slopes from FIG. 35 and the corresponding thrombin concentrations read from the standard curve in FIG. 16, as described in Example 3h.
[0077] FIG. 37 shows the changes of absorbances over the 5 minute period in pooled "normal" citrated plasma, with each line representating a BD RST tube filled with 1 mL or 4 mL normal pooled citrate plasma and a different concentration of heparin, as described in Example 3h.
[0078] FIG. 38 shows TEG traces for comparison of plain tube, commercially available serum tubes, and PtPA-containing tube as described in Example 4c.
[0079] FIG. 39 shows TEG plots of the results in Example 5a.
[0080] FIG. 40, panels A and B show TEG plots of the results for volunteer "W1" in Example 5c.
[0081] FIG. 41 shows TEG plots of the results for volunteer "W2" in Example 5c.
[0082] FIG. 42 shows TEG traces for clotting of recalcified citrated blood from a heparinised participant with PtPA or OsPA as described in Example 5d.
[0083] FIG. 43 shows the TEG traces of the Example 5e results.
[0084] FIG. 44, panels A-D, show the TEG plots of the Example 6a results.
[0085] FIG. 45, panels A-D, show the TEG plots of the Example 6b results.
[0086] FIG. 46 plots the clotting times of prothrombin activator-containing snake venoms from the species: Pseudonaja textilis (Pt), Oxyuranus scutellatus (Os), Oxyuranus microlepidotus (Om), Notechis scutatus (Ns), and Echis carinatus (Ec) at different concentrations as described in Example 7.
[0087] FIG. 47 shows a Greiner plasma tube with gelatinous precipitate as described in Example 8.
[0088] FIG. 48 shows post-centrifugation (latent) clotting in a number of tubes as described in Example 8.
[0089] FIG. 49 shows a comparison of serum samples in different tubes as described in Example 8.
[0090] FIG. 50 shows the range of fibrinogen/fdp/FDP concentrations measured in Greiner serum tubes from 48 randomly selected patients requiring analyte determination as described in Example 9a. The bar shows a mean of 17.5 .mu.g/mL and range is 4.4-32 .mu.g/mL.
[0091] FIG. 51 shows the comparison of the fibrinogen/fdp/FDP concentration measured by ELISA in 36 normal serum samples prepared using Greiner serum (GS) tubes with or without the addition of PtPA as described in Example 9b.
[0092] FIG. 52 shows the fibrinogen/fdp/FDP concentrations measured by ELISA in the sera from 9 normal blood samples collected in four different serum tubes: Greiner serum (GRS), BD serum (BDS), BD RST and PtPA (300 ng/mL) as described in Example 9b.
[0093] FIG. 53 shows the fibrinogen/fdp/FDP concentrations measured by ELISA in the sera from 5 normal blood samples collected in Greiner serum tubes (GRS), Greiner No Additive tubes with 300 ng/mL of PtPA (PTPA), Greiner No Additive tubes containing 125 ng/mL of OsPA (OSPA) and Greiner No Additive tubes containing 0.16 U/mL of purified ecarin (ECARIN) as described in Example 9b. The bars represent mean.+-.standard deviation.
[0094] FIG. 54 shows the fibrinogen/fdp/FDP concentrations measured by ELISA of sera from 3 renal dialysis patients collected in Greiner serum tubes (GRS), BD SST II tubes (BDS), BD RST tubes (BDRST) and Greiner No Additive tubes with added 1.2 .mu.g/4 mL tube PtPA (PTPA) as described in Example 9c.
[0095] FIG. 55 shows the fibrinogen/fdp/FDP concentrations measured by ELISA of plasma and sera from 2 cardiac patients collected in Greiner plasma tubes (GRLH), Greiner serum tubes (GRS), BD RST tubes (BDRST), Greiner Vacuette.TM. No Additive tube with added 300 ng/mL PtPA (PTPA), Greiner Vacuette.TM. No Additive tube with added 125 ng/mL OsPA (OSPA), Greiner Vacuette.TM. No Additive tube with added 0.31 U/mL of ecarin (EC1) and Greiner Vacuette.TM. No Additive tube with added 0.63 U/mL of ecarin (EC2) as described in Example 9c.
[0096] FIG. 56 shows Giemsa-stained Cytospin slides showing cell content above the gel barrier as described in Example 9b where the slides from left to right are: (P)--PtPA serum; (S)--Greiner serum; (LH)--diluted Greiner lithium heparin plasma; and (LH)--undiluted Greiner lithium heparin plasma.
[0097] FIG. 57 shows total protein determination carried out under standard Pathology Queensland procedures on normal serum and plasma samples, (n=26), all serum and plasma samples (n=61) and cardiac patient samples (n=11) as described in Example 12a.
[0098] FIG. 58 shows the time taken for two different PtPA concentrations to clot plasma at selected time points over a period of two weeks (336 hours) when stored at different temperatures, as described in Example 15a.
[0099] FIG. 59 shows the activity of OsPA against the chromogenic substrate S-2765 after irradiation in Greiner plain tubes (P) and in Greiner serum tubes containing silica and surfactant (siP), as described in Example 15b.
[0100] FIG. 60 shows the citrated plasma clotting activity of OsPA in Greiner plain tubes (P) and in Greiner serum tubes containing silica and surfactant, as described in Example 15b.
[0101] FIG. 61 shows the activity (A.sub.405/mm) of OsPA against the chromogenic substrate S-2765 over a period of up to 14 days at 23.degree. C. compared to a fresh dilution of stock, as described in Example 15c.
[0102] FIG. 62 shows an example of a device (point-of-care device in the form of a syringe) designed to produce high quality serum for analysis, as described in Example 17.
DETAILED DESCRIPTION OF THE INVENTION
1. Definitions
[0103] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by those of ordinary skill in the art to which the invention belongs. Although any methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, preferred methods and materials are described. For the purposes of the present invention, the following terms are defined below.
[0104] The articles "a" and "an" are used herein to refer to one or to more than one (i.e. to at least one) of the grammatical object of the article. By way of example, "an element" means one element or more than one element.
[0105] By "about" is meant a quantity, level, value, number, frequency, percentage, dimension, size, amount, weight or length that varies by as much 15, 10, 9, 8, 7, 6, 5, 4, 3, 2 or 1% to a reference quantity, level, value, number, frequency, percentage, dimension, size, amount, weight or length.
[0106] The term "biologically active fragment", as applied to fragments of a reference or full-length polynucleotide or polypeptide sequence, refers to a fragment that has at least about 0.1, 0.5, 1, 2, 5, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, 90, 95, 96, 97, 98, 99% of the activity of a reference sequence. Included within the scope of the present invention are biologically active fragments, including those of at least about 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 40, 50, 60, 70, 80, 90, 100, 120, 140, 160, 180, 200, 250, 300, 400, 500, 600, 700, 800, 900, 1,000, 1,500, 2,000 nucleotides or residues in length, which comprise or encode an activity of a reference polynucleotide or polypeptide. Representative biologically active fragments generally participate in an interaction, e.g. an intramolecular or an inter-molecular interaction. An inter-molecular interaction can be a specific binding interaction or an enzymatic interaction (e.g., the interaction can be transient and a covalent bond is formed or broken). Biologically active fragments of a full-length polypeptide include peptides may comprise amino acid sequences sufficiently similar to or derived from the amino acid sequences of a (putative) full-length polypeptide. Typically, biologically active fragments comprise a domain or motif with at least one activity of a full-length polypeptide. Suitably, the biologically-active fragment has no less than about 1%, 10%, 25% 50% of an activity of the full-length polypeptide from which it is derived.
[0107] By "coding sequence" is meant any nucleic acid sequence that contributes to the code for the polypeptide product of a gene. By contrast, the term "non-coding sequence" refers to any nucleic acid sequence that does not contribute to the code for the polypeptide product of a gene.
[0108] Throughout this specification, unless the context requires otherwise, the words "comprise," "comprises" and "comprising" will be understood to imply the inclusion of a stated step or element or group of steps or elements but not the exclusion of any other step or element or group of steps or elements. Thus, use of the term "comprising" and the like indicates that the listed elements are required or mandatory, but that other elements are optional and may or may not be present. By "consisting of" is meant including, and limited to, whatever follows the phrase "consisting of". Thus, the phrase "consisting of" indicates that the listed elements are required or mandatory, and that no other elements may be present. By "consisting essentially of" is meant including any elements listed after the phrase, and limited to other elements that do not interfere with or contribute to the activity or action specified in the disclosure for the listed elements. Thus, the phrase "consisting essentially of" indicates that the listed elements are required or mandatory, but that other elements are optional and may or may not be present depending upon whether or not they affect the activity or action of the listed elements.
[0109] The terms "complementary" and "complementarity" refer to polynucleotides (i.e., a sequence of nucleotides) related by the base-pairing rules. For example, the sequence "A-G-T," is complementary to the sequence "T-C-A." Complementarity may be "partial," in which only some of the nucleic acids' bases are matched according to the base pairing rules. Or, there may be "complete" or "total" complementarity between the nucleic acids. The degree of complementarity between nucleic acid strands has significant effects on the efficiency and strength of hybridisation between nucleic acid strands.
[0110] By "corresponds to" or "corresponding to" is meant (a) a polynucleotide having a nucleotide sequence that is substantially identical or complementary to all or a portion of a reference polynucleotide sequence or encoding an amino acid sequence identical to an amino acid sequence in a peptide or protein; or (b) a peptide or polypeptide having an amino acid sequence that is substantially identical to a sequence of amino acids in a reference peptide or protein.
[0111] As used herein, the term "detecting an analyte" means determining the presence, absence, amount or concentration of one or more analytes in a sample.
[0112] By "gene" is meant a unit of inheritance that occupies a specific locus on a chromosome and consists of transcriptional and/or translational regulatory sequences and/or a coding region and/or non-translated sequences (i.e., introns, 5' and 3' untranslated sequences).
[0113] "Homology" refers to the percentage number of nucleic or amino acids that are identical or constitute conservative substitutions. Homology may be determined using sequence comparison programs such as GAP (Devereux et al., 1984) which is incorporated herein by reference. In this way sequences of a similar or substantially different length to those cited herein could be compared by insertion of gaps into the alignment, such gaps being determined, for example, by the comparison algorithm used by GAP.
[0114] The term "host cell" includes an individual cell or cell culture which can be or has been a recipient of any recombinant vector(s) or isolated polynucleotide of the invention. Host cells include progeny of a single host cell, and the progeny may not necessarily be completely identical (in morphology or in total DNA complement) to the original parent cell due to natural, accidental, or deliberate mutation and/or change. A host cell includes cells transfected or infected in vivo or in vitro with a recombinant vector or a polynucleotide of the invention. A host cell which comprises a recombinant vector of the invention is a recombinant host cell.
[0115] "Hybridisation" is used herein to denote the pairing of complementary nucleotide sequences to produce a DNA-DNA hybrid or a DNA-RNA hybrid. Complementary base sequences are those sequences that are related by the base-pairing rules. In DNA, A pairs with T and C pairs with G. In RNA U pairs with A and C pairs with G. In this regard, the terms "match" and "mismatch" as used herein refer to the hybridisation potential of paired nucleotides in complementary nucleic acid strands. Matched nucleotides hybridise efficiently, such as the classical A-T and G-C base pair mentioned above. Mismatches are other combinations of nucleotides that do not hybridise efficiently.
[0116] By "isolated" is meant material that is substantially or essentially free from components that normally accompany it in its native state. For example, an "isolated polynucleotide," as used herein, refers to a polynucleotide, which has been purified from the sequences which flank it in a naturally-occurring state, e.g., a DNA fragment which has been removed from the sequences that are normally adjacent to the fragment. Alternatively, an "isolated peptide" or an "isolated polypeptide" and the like, as used herein, refer to in vitro isolation and/or purification of a peptide or polypeptide molecule from its natural cellular environment, and from association with other components of the cell, i.e., it is not associated with in vivo substances.
[0117] By "obtained from" is meant that the polypeptide or complex, for example, is isolated from, or derived from, a particular source.
[0118] The term "oligonucleotide" as used herein refers to a polymer composed of a multiplicity of nucleotide residues (deoxyribonucleotides or ribonucleotides, or related structural variants or synthetic analogues thereof) linked via phosphodiester bonds (or related structural variants or synthetic analogues thereof). Thus, while the term "oligonucleotide" typically refers to a nucleotide polymer in which the nucleotide residues and linkages between them are naturally occurring, it will be understood that the term also includes within its scope various analogues including, but not restricted to, peptide nucleic acids (PNAs), phosphoramidates, phosphorothioates, methyl phosphonates, 2-O-methyl ribonucleic acids, and the like. The exact size of the molecule can vary depending on the particular application. An oligonucleotide is typically rather short in length, generally from about 10 to 30 nucleotide residues, but the term can refer to molecules of any length, although the term "polynucleotide" or "nucleic acid" is typically used for large oligonucleotides.
[0119] The term "operably linked" as used herein means placing a structural gene under the regulatory control of a promoter, which then controls the transcription and optionally translation of the gene. In the construction of heterologous promoter/structural gene combinations, it is generally preferred to position the genetic sequence or promoter at a distance from the gene transcription start site that is approximately the same as the distance between that genetic sequence or promoter and the gene it controls in its natural setting; i.e. the gene from which the genetic sequence or promoter is derived. As is known in the art, some variation in this distance can be accommodated without loss of function. Similarly, the preferred positioning of a regulatory sequence element with respect to a heterologous gene to be placed under its control is defined by the positioning of the element in its natural setting; i.e., the genes from which it is derived.
[0120] The terms "patient", "subject" and "individual" are used interchangeably and refer to patients, subjects and individuals of human or other mammals and includes any one for whom it is desired to detect analyte levels or to diagnose the presence, absence or severity of a disease or condition using the invention. However, it will be understood that "patient" does not imply that symptoms are present. Suitable mammals that fall within the scope of the invention include, but are not restricted to, primates (e.g. humans, chimpanzees), livestock animals (e.g. sheep, cows, horses, donkeys, pigs), laboratory test animals (e.g. rabbits, mice, rats, guinea pigs, hamsters), companion animals (e.g. cats, dogs) and captive wild animals (e.g. foxes, deer, dingoes).
[0121] The term "polynucleotide" or "nucleic acid" as used herein designates mRNA, RNA, cRNA, cDNA or DNA. The term typically refers to polymeric form of nucleotides of at least 10 bases in length, either ribonucleotides or deoxynucleotides or a modified form of either type of nucleotide. The term includes single and double stranded forms of DNA.
[0122] The terms "polynucleotide variant" and "variant" and the like refer to polynucleotides displaying substantial sequence identity with a reference polynucleotide sequence or polynucleotides that hybridise with a reference sequence under stringent conditions that are defined hereinafter. These terms also encompass polynucleotides that are distinguished from a reference polynucleotide by the addition, deletion or substitution of at least one nucleotide. Accordingly, the terms "polynucleotide variant" and "variant" include polynucleotides in which one or more nucleotides have been added or deleted, or replaced with different nucleotides. In this regard, it is well understood in the art that certain alterations inclusive of mutations, additions, deletions and substitutions can be made to a reference polynucleotide whereby the altered polynucleotide retains the biological function or activity of the reference polynucleotide. The terms "polynucleotide variant" and "variant" also include naturally occurring allelic variants.
[0123] "Polypeptide", "peptide" and "protein" are used interchangeably herein to refer to a polymer of amino acid residues and to variants and synthetic analogues of the same. Thus, these terms apply to amino acid polymers in which one or more amino acid residues are synthetic non-naturally occurring amino acids, such as a chemical analogue of a corresponding naturally occurring amino acid, as well as to naturally-occurring amino acid polymers.
[0124] The term "polypeptide variant" refers to polypeptides that are distinguished from a reference polypeptide by the addition, deletion or substitution of at least one amino acid residue. In certain embodiments, a polypeptide variant is distinguished from a reference polypeptide by one or more substitutions, which may be conservative or non-conservative. In certain embodiments, the polypeptide variant comprises conservative substitutions and, in this regard, it is well understood in the art that some amino acids may be changed to others with broadly similar properties without changing the nature of the activity of the polypeptide. Polypeptide variants also encompass polypeptides in which one or more amino acids have been added or deleted, or replaced with different amino acid residues.
[0125] By "primer" is meant an oligonucleotide which, when paired with a strand of DNA, is capable of initiating the synthesis of a primer extension product in the presence of a suitable polymerising agent. The primer is preferably single-stranded for maximum efficiency in amplification but can alternatively be double-stranded. A primer must be sufficiently long to prime the synthesis of extension products in the presence of the polymerization agent. The length of the primer depends on many factors, including application, temperature to be employed, template reaction conditions, other reagents, and source of primers. For example, depending on the complexity of the target sequence, the oligonucleotide primer typically contains 15 to 35 or more nucleotide residues, although it can contain fewer nucleotide residues. Primers can be large polynucleotides, such as from about 200 nucleotide residues to several kilobases or more. Primers can be selected to be "substantially complementary" to the sequence on the template to which it is designed to hybridise and serve as a site for the initiation of synthesis. By "substantially complementary", it is meant that the primer is sufficiently complementary to hybridise with a target polynucleotide. Preferably, the primer contains no mismatches with the template to which it is designed to hybridise but this is not essential. For example, non-complementary nucleotide residues can be attached to the 5' end of the primer, with the remainder of the primer sequence being complementary to the template. Alternatively, non-complementary nucleotide residues or a stretch of non-complementary nucleotide residues can be interspersed into a primer, provided that the primer sequence has sufficient complementarity with the sequence of the template to hybridise therewith and thereby form a template for synthesis of the extension product of the primer.
[0126] "Probe" refers to a molecule that binds to a specific sequence or sub-sequence or other moiety of another molecule. Unless otherwise indicated, the term "probe" typically refers to a polynucleotide probe that binds to another polynucleotide, often called the "target polynucleotide", through complementary base pairing. Probes can bind target polynucleotides lacking complete sequence complementarity with the probe, depending on the stringency of the hybridisation conditions. Probes can be labelled directly or indirectly.
[0127] The term "reference result" includes a result taken from the same subject at a different time, a result from a normal subject or a group of normal subjects, or a reference standard used in an analytical test.
[0128] By "regulatory element" or "regulatory sequence" is meant nucleic acid sequences (e.g., DNA) necessary for expression of an operably linked coding sequence in a particular host cell. The regulatory sequences that are suitable for prokaryotic cells for example, include a promoter, and optionally a cis-acting sequence such as an operator sequence and a ribosome binding site. Control sequences that are suitable for eukaryotic cells include promoters, polyadenylation signals, transcriptional enhancers, translational enhancers, leader or trailing sequences that modulate mRNA stability, as well as targeting sequences that target a product encoded by a transcribed polynucleotide to an intracellular compartment within a cell or to the extracellular environment.
[0129] The term "sequence identity" as used herein refers to the extent that sequences are identical on a nucleotide-by-nucleotide basis or an amino acid-by-amino acid basis over a window of comparison. Thus, a "percentage of sequence identity" is calculated by comparing two optimally aligned sequences over the window of comparison, determining the number of positions at which the identical nucleic acid base (e.g., A, T, C, G, I) or the identical amino acid residue (e.g., Ala, Pro, Ser, Thr, Gly, Val, Leu, Ile, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gln, Cys and Met) occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of sequence identity. For the purposes of the present invention, "sequence identity" may be understood to mean the "match percentage" calculated by the DNASIS computer program (Version 2.5 for Windows; available from Hitachi Software Engineering Co., Ltd., South San Francisco, Calif., USA) using standard defaults as used in the reference material accompanying the software.
[0130] The term "sequence similarity" refers to the percentage number of amino acids that are identical or constitute conservative substitutions as defined in Table 2 infra. Similarity may be determined using sequence comparison programs such as GAP (Devereux et al., 1984). In this way, sequences of a similar or substantially different length to those cited herein might be compared by insertion of gaps into the alignment, such gaps being determined, for example, by the comparison algorithm used by GAP.
[0131] Terms used to describe sequence relationships between two or more polynucleotides or polypeptides include "reference sequence", "comparison window", "sequence identity", "percentage of sequence identity" and "substantial identity". A "reference sequence" is at least 12 but frequently 15 to 18 and often at least 25 monomer units, inclusive of nucleotides and amino acid residues, in length. Because two polynucleotides may each comprise (1) a sequence (i.e., only a portion of the complete polynucleotide sequence) that is similar between the two polynucleotides, and (2) a sequence that is divergent between the two polynucleotides, sequence comparisons between two (or more) polynucleotides are typically performed by comparing sequences of the two polynucleotides over a "comparison window" to identify and compare local regions of sequence similarity. A "comparison window" refers to a conceptual segment of at least 6 contiguous positions, usually about 50 to about 100, more usually about 100 to about 150 in which a sequence is compared to a reference sequence of the same number of contiguous positions after the two sequences are optimally aligned. The comparison window may comprise additions or deletions (i.e., gaps) of about 20% or less as compared to the reference sequence (which does not comprise additions or deletions) for optimal alignment of the two sequences. Optimal alignment of sequences for aligning a comparison window may be conducted by computerized implementations of algorithms (GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package Release 7.0, Genetics Computer Group, 575 Science Drive Madison, Wis., USA) or by inspection and the best alignment (i.e., resulting in the highest percentage homology over the comparison window) generated by any of the various methods selected. Reference also may be made to the BLAST family of programs as for example disclosed by Altschul et al., 1997. A detailed discussion of sequence analysis can be found in Unit 19.3 of Ausubel et al., "Current Protocols in Molecular Biology", John Wiley & Sons Inc, 1994-1998, Chapter 15.
[0132] "Stringency" as used herein, refers to the temperature and ionic strength conditions, and presence or absence of certain organic solvents, during hybridisation and washing procedures. The higher the stringency, the higher will be the degree of complementarity between immobilized target nucleotide sequences and the labelled probe polynucleotide sequences that remain hybridised to the target after washing. The term "high stringency" refers to temperature and ionic conditions under which only nucleotide sequences having a high frequency of complementary bases will hybridise. The stringency required is nucleotide sequence dependent and depends upon the various components present during hybridisation. Generally, stringent conditions are selected to be about 10 to 20.degree. C. lower than the thermal melting point (Tm) for the specific sequence at a defined ionic strength and pH. The Tm is the temperature (under defined ionic strength and pH) at which 50% of a target sequence hybridises to a complementary probe.
[0133] The term "transformation" means alteration of the genotype of an organism, for example a bacterium, yeast, mammal, avian, reptile, fish or plant, by the introduction of a foreign or endogenous nucleic acid.
[0134] By "vector" is meant a polynucleotide molecule, preferably a DNA molecule derived, for example, from a plasmid, bacteriophage, yeast or virus, into which a polynucleotide can be inserted or cloned. A vector preferably contains one or more unique restriction sites and can be capable of autonomous replication in a defined host cell including a target cell or tissue or a progenitor cell or tissue thereof, or be integrable with the genome of the defined host such that the cloned sequence is reproducible. Accordingly, the vector can be an autonomously replicating vector, i.e., a vector that exists as an extra-chromosomal entity, the replication of which is independent of chromosomal replication, e.g., a linear or closed circular plasmid, an extra-chromosomal element, a mini-chromosome, or an artificial chromosome. The vector can contain any means for assuring self-replication. Alternatively, the vector can be one which, when introduced into the host cell, is integrated into the genome and replicated together with the chromosome(s) into which it has been integrated. A vector system can comprise a single vector or plasmid, two or more vectors or plasmids, which together contain the total DNA to be introduced into the genome of the host cell, or a transposon. The choice of the vector will typically depend on the compatibility of the vector with the host cell into which the vector is to be introduced. In the present case, the vector is preferably a viral or viral-derived vector, which is operably functional in animal and preferably mammalian cells. Such vector may be derived from a poxvirus, an adenovirus or yeast. The vector can also include a selection marker such as an antibiotic resistance gene that can be used for selection of suitable transformants. Examples of such resistance genes are known to those of skill in the art and include the nptII gene that confers resistance to the antibiotics kanamycin and G418 (Geneticin.RTM.) and the hph gene which confers resistance to the antibiotic hygromycin B.
[0135] The terms "wild-type" and "naturally occurring" are used interchangeably to refer to a gene or gene product that has the characteristics of that gene or gene product when isolated from a naturally occurring source. A wild type gene or gene product (e.g., a polypeptide) is that which is most frequently observed in a population and is thus arbitrarily designed the "normal" or "wild-type" form of the gene.
2. Prothrombin Activators
[0136] The present invention is based in part on the discovery that prothrombin activators are suitable dotting agents for preparing serum that is used to detect analytes, despite their known proteolytic activity. Prothrombin activators (sometimes known as prothrombinases) exhibit trypsin-like activity and activate prothrombin (i.e. convert prothrombin to thrombin which in turn converts fibrinogen to fibrin and hence cause clot formation).
[0137] In some embodiments, the prothrombin activator is an exogenous prothrombin activator. As used herein, an "exogenous prothrombin activator" means a prothrombin activator obtained from a source other than the blood sample from which the serum sample is to be prepared.
[0138] 2.1 Wild-Type or Naturally-Occurring Prothrombin Activators
[0139] The prothrombin activators used in the present invention may comprise wild-type or naturally-occurring prothrombin activators including those obtained from any suitable organism, including snake, human, bovine and bacterial prothrombin activator. The prothrombin activator may comprise a full-length wild-type or naturally occurring polypeptide.
[0140] In certain embodiments, the prothrombin activator is a snake prothrombin activator. Suitably, the prothrombin activator is a snake venom prothrombin activator. Snake venom prothrombin activators are generally classified in four groups (A, B, C, and D) depending on their structure, function and requirements for co-factors.
[0141] Suitably, the snake venom prothrombin activator is a group A prothrombin activator. Group A prothrombin activators are metalloproteinases consisting of three domains: a metalloproteinase, a disintegrin, and a Cys-rich domain. The metalloproteinase domain contains the consensus sequence HEXXHXXGXXH (SEQ ID NO:53), corresponding to the zinc-chelating active site. These prothrombin activators are found at least in several viper venoms, and include ecarin from Echis carinatus venom and basparin from Bothrops aspen venomSuitably, the snake venom prothrombin activator is a group B prothrombin activator. Group B prothrombin activators are metalloproteinases consisting of two subunits held non-covalently: a metalloproteinase and a C-type lectin-like disulfide-liked dimer. These prothrombin activators are found in several viper venoms, and include carinactivase-1 and carinactivase-2 from Echis carinatus venom and multactivase from Echis multisquamatus venom.
[0142] Suitably, the snake venom prothrombin activator is a group C prothrombin activator. Group C prothrombin activators are serine proteases and resemble the mammalian factor Xa-factor Va complex. Pseutarin C (or PtPA) and oscutarin C (or OsPA) are group C prothrombin activators from the venoms of Pseudonaja textilis and Oxyuranus scutellatus respectively. Omicarin C is the prothrombin activator from Oxyuranus microlepidotus venom.
[0143] Suitably, the snake venom prothrombin activator is a group D prothrombin activator. Group D prothrombin activators are serine proteases and are functionally similar to mammalian factor Xa. Porpharin D (from Pseudechis porphyriacus), notecarin D (from Notechis scutatus scutatus), trocarin D (from Tropidechis carinatus), hopsarin D (from Hoplocephalus stephensi), and notenarin D (from Notechis ater niger) are all group D prothrombin activators.
[0144] A review of snake prothrombin activators is provided in Kini, R. M. (2005), and of those specifically from the venom of Australian Elapids (group C and D prothrombin activators) is in St. Pierre et al. (2005), the contents of each are herein by reference in their entirety. These two reviews use the classification of snake prothrombin activators into groups A-D as described above. This classification supersedes the previous classification system using groups I-III (group I encompasses groups A and B; group II is now group D and group III is now group C), and sometimes additional groups IV (snake venom activators that cleave peptide bonds in prothrombin but do not convert the prothrombin to an enzymatically active product--i.e. thrombin or meizothrombin) and V (bacterial prothrombin activators) as described in earlier review articles, including Rosing, J. et al. (1991) and Rosing, J. et al. (1992), the contents of each being incorporated by reference in their entirety. For an explanation on the change to the classification system, see Kini, R, M., et al. (2001), the contents of which are incorporated by reference in its entirety.
[0145] In specific embodiments, the snake prothrombin activator is obtained from the Family Elapidae, illustrative examples of which include species from the genera Demansia, Hoplocephalus, Notechis, Oxyuranus, Pseudechis, Pseudonaja, Rhinoplocephalus, and Tropidechis including but not limited to Demansia vestigiata, Hoplocephalus stephensii, Notechis ater humphreysi, Notechis ater niger, Notechis ater serventyi, Notechis flinkders, Notechis humphreysi, Notechis niger, Notechis occidentalis, Notechis scutatus, Notechis scutatus scutatus, Notechis serventyi, Oxyuranus microlepidotus, Oxyuranus scutellatus, Pseudechis porphyriacus, Pseudonaja affinis, Pseudonaja inframaculata, Pseudonaja nuchalis, Pseudonaja textilis, Rhinoplocephalus nigrescens, and Tropidechis carinatus.
[0146] In specific embodiments, the snake prothrombin activator is obtained from the Family Viperidae, illustrative examples of which include species from the genera Botrhops, Echis and Trimeresurus, including but not limited to Bothrops alternatus, Bothrops asper, Bothrops atrox, Bothrops atrox asper, Bothrops brasili, Bothrops castelnaudi, Bothrops columbiensis, Bothrops erythromelas, Bothrops fonsecai, Bothrops itapetiningae, Bothrops jararaca, Bothrops neuwiedi, Bothrops venezuelensis, Echis carinatus, Echis coloratus, Echis multisquamatus, and Trimeresurus okinavensis.
[0147] In specific embodiments, the snake prothrombin activator is obtained from the Family Colubridae, illustrative examples of which include species from the genera Dispholidus, Rhabdophis and Thelotornis, including but not limited to Dispholidus typus, Rhabdophis tigrinus tigrinus, Thelotornis kirtlandii, and Thelotornis capensis.
[0148] In some embodiments the snake prothrombin activator is from or is obtained from snake venom. The purification and characterisation of PtPA from P. textilis snake venom is described in Masci (1986) and Masci et al., (1998), and OsPA from O. scutellatus venom is described in Speijer et al., (1986), all of which are incorporated by reference in their entirety. The purification and characterisation of ecarin from Echis carinatus venom is described in Morita, T et al. (1981) and Nishida, S et al. (1995), of carinactivase from Echis carinatus venom is described in Yamada, D et al. (1996), of multactivase from Echis multisquamatus is described in Yamada, D. et al., (1997), and of notecarin from Notechis scutatus is described in Tans, Get al., (1985), each of which are incorporated by reference in their entirety.
[0149] In certain embodiments, the prothrombin activator is a mammalian prothrombin activator. Mammalian prothrombin activators include those derived from human blood and/or tissue and those derived from bovine blood and/or tissue.
[0150] In certain embodiments, the prothrombin activator is a bacterial prothrombin activator. Bacterial prothrombin activators include those from Staphylococcus aureus, Peptococcus indolicus, Bacteroides melaninogenicus, and Pseudomonas aeruginosa (Rosing, J. et al. (1991).
[0151] As will be appreciated by those skilled in the art, the prothrombin activator may comprise, consist essentially of, or consist of one or more polypeptides. In some embodiments, the prothrombin activator comprises, consists essentially of, or consists of a single polypeptide. In other embodiments, the prothrombin activator comprises, consists essentially of, or consists of more than one polypeptide, including but not limited to complexes of polypeptides. Where the prothrombin activator comprises, consists essentially of, or consists of more than one polypeptide, each polypeptide may be from the organisms from the same or different genera, and/or the same or different species.
[0152] In certain embodiments, the prothrombin activator comprises an amino acid sequence selected from those set forth in SEQ ID NOs: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, and 52 or comprises an amino acid sequence encoded by a nucleotide sequence selected from those set forth in SEQ ID NOs: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, and 49.
[0153] 2.2 Chimeric Prothrombin Activators and Fusion Polypeptides
[0154] The present invention also contemplates the use of prothrombin activators comprising a chimeric polypeptide. As used herein, a "chimeric polypeptide" includes a first polypeptide component comprising a polypeptide obtained from a first organism linked to a second polypeptide component obtained from a second organism. In some embodiments, the first organism and the second organism are from different genera. In other embodiments, the first organism and the second organism are different species of the same genus. In certain embodiments, the prothrombin activator comprises a chimeric polypeptide that resembles a factor Xa-factor Va complex, wherein the first polypeptide comprises a factor Xa-like polypeptide and the second polypeptide comprises a factor Va-like polypeptide. In certain specific embodiments, the first polypeptide comprises an amino acid sequence selected from those set forth in SEQ ID NOs: 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, and 50, or comprises an amino acid sequence encoded by a nucleotide sequence selected from those set forth in SEQ ID NOs: 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, and 49, and the second polypeptide comprises an amino acid sequence selected from those set forth in SEQ ID NOs: 7, 8, 11, 12, 13, 16, and 18, or comprises an amino acid sequence encoded by a nucleotide sequence selected from those set forth in SEQ ID NOs: 5, 6, 9, 10, 14, 15, and 17.
[0155] The present invention also contemplates the use of prothrombin activators comprising a fusion polypeptide. As used herein, a "fusion polypeptide" includes a first polypeptide component linked to a second polypeptide component. The first polypeptide component may be obtained from a first organism and the second polypeptide component may be obtained from a second organism. In some embodiments, the first organism and the second organism are from different genera. In other embodiments, the first organism and the second organism are different species of the same genus. The first polypeptide component or the second polypeptide component of the fusion polypeptide can correspond to all or a portion (e.g., a fragment as described herein) of a wild-type or naturally occurring amino acid sequence. The second polypeptide component can be fused to the N-terminus or C-terminus of the first polypeptide component.
[0156] 2.3 Fragments of Wild-Type or Naturally Occurring Polypeptides
[0157] The prothrombin activator may comprise a fragment of a full-length wild-type or naturally occurring polypeptide, wherein the prothrombin activator exhibits prothrombin activating activity.
[0158] Typically, fragments of a full-length polypeptide may participate in an interaction, for example an intramolecular or an intermolecular interaction. Such fragments include peptides comprising the amino acid sequences shown in SEQ ID NOs: 2, 3, 4, 51, and 52 and peptides comprising amino acid sequences sufficiently similar to or derived from the amino acid sequences of a (putative) full-length polypeptide, for example, the amino acid sequences shown in SEQ ID NOs: 1, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, and 50, or the amino acid sequences encoded by a nucleotide sequence selected from those set forth in SEQ ID NOs: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, and 49, which includes less amino acids than a full-length polypeptide, and exhibit one activity of that polypeptide.
[0159] 2.4 Variants of Naturally-Occurring Prothrombin Activators (Polypeptide)
[0160] The present invention also contemplates prothrombin activators comprising polypeptide(s) that is/are variant(s) of the wild-type or naturally-occurring polypeptide(s). Prothrombin activators comprising one or more variant polypeptides encompassed by the present invention are biologically active, that is, they continue to possess prothrombin activating activity.
[0161] Such "variant" prothrombin activators include polypeptides derived from the native polypeptide, wherein the polypeptides are derived from the corresponding native polypeptide(s) by deletion (so-called truncation) or addition of one or more amino acids to the N-terminal and/or C-terminal end of the native polypeptide(s); deletion or addition of one or more amino acids at one or more sites in the native polypeptide(s); or substitution of one or more amino acids at one or more sites in the native polypeptide(s). These variant prothrombin activators may result from, for example, genetic polymorphism or human manipulation.
[0162] Further non-limiting examples of variant polypeptides include precursor polypeptide or polypeptide in zymogen form processed forms of a full-length or precursor polypeptide or polypeptide in zymogen form.
[0163] Variants of a wild-type or naturally-occurring polypeptide will have at least 40%, 50%, 60%, 70%, generally at least 75%, 80%, 85%, usually about 90%, 91%, 92%, 93%, 94%, 95% or more, and typically about 96%, 97%, 98% or more (and all integer percentages in between) sequence similarity or identity with the amino acid sequence for the wild-type or naturally-occurring polypeptide, including but not limited to the sequences in SEQ ID NOs: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, and 52, or the amino acid sequences encoded by the nucleotide sequences in SEQ ID NOs: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, and 49, as determined by sequence alignment programs described elsewhere herein using default parameters. A variant of a wild-type or naturally-occurring polypeptide, which falls within the scope of a variant polypeptide, may differ from that polypeptide generally by as much 200, 100, 50 or 20 amino acid residues or suitably by as few as 1-15 amino acid residues, as few as 1-10, such as 6-10, as few as 5, as few as 4, 3, 2, or even 1 amino acid residue. In some embodiments, a variant polypeptide differs from the corresponding sequences in SEQ ID NOs: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, or 52, or the amino acid sequences encoded by the nucleotide sequences in SEQ ID NOs: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, or 49, by at least 1 but by less than 15, 10 or 5 amino acid residues. In other embodiments, it differs from the corresponding sequences in SEQ ID NOs: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, or 52, or the amino acid sequences encoded by the nucleotide sequences in SEQ ID NOs: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, or 49, by at least one residue but less than 20%, 15%, 10% or 5% of the residues.
[0164] A polypeptide may be altered in various ways including amino acid substitutions, deletions, truncations, and insertions. Methods for such manipulations are generally known in the art. For example, amino acid sequence variants of a polypeptide can be prepared by mutations in the DNA. Methods for mutagenesis and nucleotide sequence alterations are well known in the art. See, for example, Kunkel (1985), Kunkel et al., (1987), U.S. Pat. No. 4,873,192, Watson et al., (1987) and the references cited therein. Guidance as to appropriate amino acid substitutions that do not affect biological activity of the protein of interest may be found in the model of Dayhoff et al. (1978). Methods for screening gene products of combinatorial libraries made by point mutations or truncation, and for screening cDNA libraries for gene products having a selected property are known in the art. Such methods are adaptable for rapid screening of the gene libraries generated by combinatorial mutagenesis of polypeptides. Recursive ensemble mutagenesis (REM), a technique which enhances the frequency of functional mutants in the libraries, can be used in combination with the screening assays to identify polypeptide variants, see for example Arkin et al. (1992) and Delagrave et al. (1993). Conservative substitutions, such as exchanging one amino acid with another having similar properties, may be desirable as discussed in more detail below.
[0165] Variant polypeptides may contain conservative amino acid substitutions at various locations along their sequence, as compared to a parent (e.g., naturally-occurring or reference) amino acid sequence. A "conservative amino acid substitution" is one in which the amino acid residue is replaced with an amino acid residue having a similar side chain. Families of amino acid residues having similar side chains have been defined in the art, which can be generally sub-classified as follows:
[0166] Acidic: The residue has a negative charge due to loss of H ion at physiological pH and the residue is attracted by aqueous solution so as to seek the surface positions in the conformation of a peptide in which it is contained when the peptide is in aqueous medium at physiological pH. Amino acids having an acidic side chain include glutamic acid and aspartic acid.
[0167] Basic: The residue has a positive charge due to association with H ion at physiological pH or within one or two pH units thereof (e.g. histidine) and the residue is attracted by aqueous solution so as to seek the surface positions in the conformation of a peptide in which it is contained when the peptide is in aqueous medium at physiological pH. Amino acids having a basic side chain include arginine, lysine and histidine.
[0168] Charged: The residues are charged at physiological pH and, therefore, include amino acids having acidic or basic side chains (i.e., glutamic acid, aspartic acid, arginine, lysine and histidine).
[0169] Hydrophobic: The residues are not charged at physiological pH and the residue is repelled by aqueous solution so as to seek the inner positions in the conformation of a peptide in which it is contained when the peptide is in aqueous medium. Amino acids having a hydrophobic side chain include tyrosine, valine, isoleucine, leucine, methionine, phenylalanine and tryptophan.
[0170] Neutral/polar: The residues are not charged at physiological pH, but the residue is not sufficiently repelled by aqueous solutions so that it would seek inner positions in the conformation of a peptide in which it is contained when the peptide is in aqueous medium. Amino acids having a neutral/polar side chain include asparagine, glutamine, cysteine, histidine, serine and threonine.
[0171] This description also characterises certain amino acids as "small" since their side chains are not sufficiently large, even if polar groups are lacking, to confer hydrophobicity. With the exception of proline, "small" amino acids are those with four carbons or less when at least one polar group is on the side chain and three carbons or less when not. Amino acids having a small side chain include glycine, serine, alanine and threonine. The gene-encoded secondary amino acid proline is a special case due to its known effects on the secondary conformation of peptide chains. The structure of proline differs from all the other naturally-occurring amino acids in that its side chain is bonded to the nitrogen of the a-amino group, as well as the a-carbon. Several amino acid similarity matrices (e.g. PAM120 matrix and PAM250 matrix as disclosed for example by Dayhoff et al. (1978) and by Gonnet et al. (1992)), however, include proline in the same group as glycine, serine, alanine and threonine. Accordingly, for the purposes of the present invention, proline is classified as a "small" amino acid.
[0172] The degree of attraction or repulsion required for classification as polar or nonpolar is arbitrary and, therefore, amino acids specifically contemplated by the invention have been classified as one or the other. Most amino acids not specifically named can be classified on the basis of known behaviour.
[0173] Amino acid residues can be further sub-classified as cyclic or non-cyclic, and aromatic or non-aromatic, self-explanatory classifications with respect to the side-chain substituent groups of the residues, and as small or large. The residue is considered small if it contains a total of four carbon atoms or less, inclusive of the carboxyl carbon, provided an additional polar substituent is present; three or less if not. Small residues are, of course, always non-aromatic. Dependent on their structural properties, amino acid residues may fall in two or more classes. For the naturally-occurring protein amino acids, sub-classification according to this scheme is presented in Table 1.
TABLE-US-00002 TABLE 1 AMINO ACID SUB-CLASSIFICATION SUB-CLASSES AMINO ACIDS Acidic Aspartic acid, Glutamic acid Basic Noncyclic: Arginine, Lysine; Cyclic: Histidine Charged Aspartic acid, Glutamic acid, Arginine, Lysine, Histidine Small Glycine, Serine, Alanine, Threonine, Proline Polar/neutral Asparagine, Histidine, Glutamine, Cysteine, Serine, Threonine Polar/large Asparagine, Glutamine Hydrophobic Tyrosine, Valine, Isoleucine, Leucine, Methionine, Phenylalanine, Tryptophan Aromatic Tryptophan, Tyrosine, Phenylalanine Residues that influence Glycine and Proline chain orientation
[0174] Conservative amino acid substitution also includes groupings based on side chains. For example, a group of amino acids having aliphatic side chains is glycine, alanine, valine, leucine, and isoleucine; a group of amino acids having aliphatic-hydroxyl side chains is serine and threonine; a group of amino acids having amide-containing side chains is asparagine and glutamine; a group of amino acids having aromatic side chains is phenylalanine, tyrosine, and tryptophan; a group of amino acids having basic side chains is lysine, arginine, and histidine; and a group of amino acids having sulphur-containing side chains is cysteine and methionine. For example, it is reasonable to expect that replacement of a leucine with an isoleucine or valine, an aspartate with a glutamate, a threonine with a serine, or a similar replacement of an amino acid with a structurally related amino acid will not have a major effect on the properties of the resulting variant polypeptide. Whether an amino acid change results in a functional polypeptide can readily be determined by assaying its activity. Conservative substitutions are shown in Table 2 under the heading of exemplary and preferred substitutions. Amino acid substitutions falling within the scope of the invention, are, in general, accomplished by selecting substitutions that do not differ significantly in their effect on maintaining (a) the structure of the peptide backbone in the area of the substitution, (b) the charge or hydrophobicity of the molecule at the target site, or (c) the bulk of the side chain. After the substitutions are introduced, the variants may be screened for biological activity.
TABLE-US-00003 TABLE 2 EXEMPLARY AND PREFERRED AMINO ACID SUBSTITUTIONS ORIGINAL EXEMPLARY PREFERRED RESIDUE SUBSTITUTIONS SUBSTITUTIONS Ala Val, Leu, Ile Val Arg Lys, Gln, Asn Lys Asn Gln, His, Lys, Arg Gln Asp Glu Glu Cys Ser Ser Gln Asn, His, Lys, Asn Glu Asp, Lys Asp Gly Pro Pro His Asn, Gln, Lys, Arg Arg Ile Leu, Val, Met, Ala, Phe, Norleu Leu Leu Norleu, Ile, Val, Met, Ala, Phe Ile Lys Arg, Gln, Asn Arg Met Leu, Ile, Phe Leu Phe Leu, Val, Ile, Ala Leu Pro Gly Gly Ser Thr Thr Thr Ser Ser Trp Tyr Tyr Tyr Trp, Phe, Thr, Ser Phe Val Ile, Leu, Met, Phe, Ala, Norleu Leu
[0175] Alternatively, similar amino acids for making conservative substitutions can be grouped into three categories based on the identity of the side chains. The first group includes glutamic acid, aspartic acid, arginine, lysine, histidine, which all have charged side chains; the second group includes glycine, serine, threonine, cysteine, tyrosine, glutamine, asparagine; and the third group includes leucine, isoleucine, valine, alanine, proline, phenylalanine, tryptophan, methionine, as described in Zubay, G. (1993).
[0176] Thus, a predicted non-essential amino acid residue in a polypeptide is typically replaced with another amino acid residue from the same side chain family. Alternatively, mutations can be introduced randomly along all or part of a polypeptide gene coding sequence, such as by saturation mutagenesis, and the resultant mutants can be screened for an activity of the parent polypeptide to identify mutants which retain that activity. Following mutagenesis of the coding sequences, the encoded peptide can be expressed recombinantly and the activity of the peptide can be determined. A "non-essential" amino acid residue is a residue that can be altered from the wild-type sequence of a polypeptide without abolishing or substantially altering one or more of its activities. Suitably, the alteration does not substantially alter one of these activities, for example, the activity is at least 20%, 40%, 60%, 70% or 80% of wild-type. An "essential" amino acid residue is a residue that, when altered from the wild-type sequence of a reference polypeptide, results in abolition of an activity of the parent molecule such that less than 20% of the wild-type activity is present.
[0177] Accordingly, the present invention also contemplates variants of the naturally-occurring polypeptide sequences or their biologically-active fragments, wherein the variants are distinguished from the naturally-occurring sequence by the addition, deletion, or substitution of one or more amino acid residues. In general, variants will display at least about 30, 40, 50, 55, 60, 65, 70, 75, 80, 85, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99% similarity to a parent or reference polypeptide sequence as, for example, set forth in SEQ ID NO: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, or 52, or the parent or reference polypeptide sequence as, for example, encoded by the nucleotide sequence set forth in SEQ ID NO: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, or 49. Desirably, variants will have at least 30, 40, 50, 55, 60, 65, 70, 75, 80, 85, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99% sequence identity to a parent polypeptide sequence as, for example, set forth in SEQ ID NO: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, or 52, or the parent polypeptide sequence as, for example, encoded by the nucleotide sequence set forth in SEQ ID NO: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, or 49. Moreover, sequences differing from the native or parent sequences by the addition, deletion, or substitution of 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 30, 40, 50, 60 ,70, 80, 90, 100 or more amino acids but which retain the properties of the parent polypeptide are contemplated. Polypeptides also include polypeptides that are encoded by polynucleotides that hybridise under stringency conditions as defined herein, especially high stringency conditions, to parent-coding polynucleotide sequences, or the non-coding strand thereof, as described below. Illustrative parent polynucleotide sequences are set forth in SEQ ID NO: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, and 49.
[0178] In some embodiments, variant polypeptides differ from a reference sequence by at least one but by less than 50, 40, 30, 20, 15, 10, 8, 6, 5, 4, 3 or 2 amino acid residue(s). In other embodiments, variant polypeptides differ from the corresponding sequences of SEQ ID NO: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, or 52, or the amino acid sequences encoded by the nucleotide sequences of SEQ ID NO: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, or 49, by at least 1% but less than 20%, 15%, 10% or 5% of the residues. (If this comparison requires alignment, the sequences should be aligned for maximum similarity. "Looped" out sequences from deletions or insertions, or mismatches, are considered differences). The differences are, suitably, differences or changes at a non-essential residue or a conservative substitution.
[0179] Variants of a protein can be identified by screening combinatorial libraries of mutants, e.g., truncation mutants, of a protein. Libraries or fragments e.g., N terminal, C terminal, or internal fragments, of a protein coding sequence can be used to generate a variegated population of fragments for screening and subsequent selection of variants of a protein.
[0180] Methods for screening gene products of combinatorial libraries made by point mutation or truncation, and for screening cDNA libraries for gene products having a selected property are known in the art. Such methods are adaptable for rapid screening of the gene libraries generated by combinatorial mutagenesis of proteins.
[0181] Some variants of the snake prothrombin activator ecarin are described in U.S. Pat. No. 6,413,737.
[0182] 2.5 Variants of Naturally-Occurring Prothrombin Activators (Nucleotide)
[0183] The present invention also contemplates prothrombin activators comprising polypeptide(s) that is/are encoded by variant(s) of the wild-type or naturally-occurring polynucleotide(s) encoding the wild-type or naturally-occurring polynucleotide(s).
[0184] Variants of a wild-type or naturally-occurring polynucleotides will have at least 40%, 50%, 60%, 70%, generally at least 75%, 80%, 85%, 86%, 87%, 88%, 89%, usually about 90%, 91%, 92%, 93%, 94%, 95% or more, and typically about 96%, 97%, 98% or more (and all integer percentages in between) sequence similarity or identity with the nucleotide sequence for the wild-type or naturally-occurring polynucleotide, including but not limited to the sequences encoded by the sequences of SEQ ID NOs: 1, 2, 3, 4, 7, 8, 11, 12, 13, 16, 18, 26, 27, 28, 29, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 51, or 52 or the sequences of SEQ ID NOs: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, and 49, or a complement thereof, as determined by sequence alignment programs described elsewhere herein using default parameters.
[0185] Exemplary nucleotide sequences that encode the polypeptides encompass full-length genes as well as portions of the full-length or substantially full-length nucleotide sequences of the genes or their transcripts or DNA copies of these transcripts. Portions of a nucleotide sequence may encode polypeptide portions or segments that retain the biological activity of the native polypeptide. A portion of a nucleotide sequence that encodes a biologically active fragment of a polypeptide may encode at least about 20, 21, 22, 23, 24, 25, 30, 40, 50, 60, 70, 80, 90, 100, 120, 150, 300 or 400 contiguous amino acid residues, or almost up to the total number of amino acids present in a full-length polypeptide.
[0186] Also contemplated are variants of the nucleotide sequences. Nucleic acid variants can be naturally-occurring, such as allelic variants (same locus), homologs (different locus), and orthologs (different organism) or can be non naturally-occurring. Naturally-occurring variants such as these can be identified with the use of well-known molecular biology techniques, as, for example, with polymerase chain reaction (PCR) and hybridisation techniques as known in the art. Non-naturally occurring variants can be made by mutagenesis techniques, including those applied to polynucleotides, cells, or organisms. The variants can contain nucleotide substitutions, deletions, inversions and insertions. Variation can occur in either or both the coding and non-coding regions. The variations can produce both conservative and non-conservative amino acid substitutions (as compared in the encoded product). For nucleotide sequences, conservative variants include those sequences that, because of the degeneracy of the genetic code, encode the amino acid sequence of a reference polypeptide. Variant nucleotide sequences also include synthetically derived nucleotide sequences, such as those generated, for example, by using site-directed mutagenesis but which still encode a polypeptide. Generally, variants of a particular nucleotide sequence will have at least about 30%, 40% 50%, 55%, 60%, 65%, 70%, generally at least about 75%, 80%, 85%, 86%, 87%, 88%, 89%, desirably about 90%, 91%, 92%, 93%, 94%, 95% or more, and more suitably about 96%, 97%, 98% or more sequence identity to that particular nucleotide sequence as determined by sequence alignment programs described elsewhere herein using default parameters.
[0187] Nucleotide sequences can be used to isolate corresponding sequences and alleles from other organisms, particularly other snakes. Methods are readily available in the art for the hybridisation of nucleic acid sequences. Coding sequences from other organisms may be isolated according to well known techniques based on their sequence identity with the coding sequences set forth herein. In these techniques all or part of the known coding sequence is used as a probe which selectively hybridises to other coding sequences present in a population of cloned genomic DNA fragments or cDNA fragments (i.e., genomic or cDNA libraries) from a chosen organism (e.g., a snake). Accordingly, the present invention also contemplates polynucleotides that hybridise to reference nucleotide sequences, or to their complements, under stringency conditions described below. As used herein, the term "hybridises under low stringency, medium stringency, high stringency, or very high stringency conditions" describes conditions for hybridisation and washing.
[0188] Guidance for performing hybridisation reactions can be found in Ausubel et al., supra, sections 6.3.1-6.3.6. Aqueous and non-aqueous methods are described in that reference and either can be used. Reference herein to low stringency conditions include and encompass from at least about 1% v/v to at least about 15% v/v formamide and from at least about 1 M to at least about 2 M salt for hybridisation at 42.degree. C., and at least about 1 M to at least about 2 M salt for washing at 42.degree. C. Low stringency conditions also may include 1% Bovine Serum Albumin (BSA), 1 mM EDTA, 0.5 M NaHPO.sub.4 (pH 7.2), 7% SDS for hybridisation at 65.degree. C., and (i) 2.times.SSC, 0.1% SDS; or (ii) 0.5% BSA, 1 mM EDTA, 40 mM NaHPO.sub.4 (pH 7.2), 5% SDS for washing at room temperature. One embodiment of low stringency conditions includes hybridisation in 6.times.sodium chloride/sodium citrate (SSC) at about 45.degree. C., followed by two washes in 0.2.times.SSC, 0.1% SDS at least at 50.degree. C. (the temperature of the washes can be increased to 55.degree. C. for low stringency conditions). Medium stringency conditions include and encompass from at least about 16% v/v to at least about 30% v/v formamide and from at least about 0.5 M to at least about 0.9 M salt for hybridisation at 42.degree. C., and at least about 0.1 M to at least about 0.2 M salt for washing at 55.degree. C. Medium stringency conditions also may include 1% Bovine Serum Albumin (BSA), 1 mM EDTA, 0.5 M NaHPO.sub.4 (pH 7.2), 7% SDS for hybridisation at 65.degree. C., and (i) 2.times.SSC, 0.1% SDS; or (ii) 0.5% BSA, 1 mM EDTA, 40 mM NaHPO.sub.4 (pH 7.2), 5% SDS for washing at 60-65.degree. C. One embodiment of medium stringency conditions includes hybridising in 6.times.SSC at about 45.degree. C., followed by one or more washes in 0.2.times.SSC, 0.1% SDS at 60.degree. C. High stringency conditions include and encompass from at least about 31% v/v to at least about 50% v/v formamide and from about 0.01 M to about 0.15 M salt for hybridisation at 42.degree. C., and about 0.01 M to about 0.02 M salt for washing at 55.degree. C. High stringency conditions also may include 1% BSA, 1 mM EDTA, 0.5 M NaHPO.sub.4 (pH 7.2), 7% SDS for hybridisation at 65.degree. C., and (i) 0.2.times.SSC, 0.1% SDS; or (ii) 0.5% BSA, 1 mM EDTA, 40 mM NaHPO.sub.4 (pH 7.2), 1% SDS for washing at a temperature in excess of 65.degree. C. One embodiment of high stringency conditions includes hybridising in 6.times.SSC at about 45.degree. C., followed by one or more washes in 0.2.times.SSC, 0.1% SDS at 65.degree. C.
[0189] In certain embodiments, a polypeptide is encoded by a polynucleotide that hybridises to a disclosed nucleotide sequence under low, medium, high, or very high stringency conditions. One embodiment of very high stringency conditions includes hybridising 0.5 M sodium phosphate, 7% SDS at 65.degree. C., followed by one or more washes at 0.2.times.SSC, 1% SDS at 65.degree. C.
[0190] Other stringency conditions are well known in the art and a skilled addressee will recognise that various factors can be manipulated to optimise the specificity of the hybridisation. Optimisation of the stringency of the final washes can serve to ensure a high degree of hybridisation. For detailed examples, see Ausubel et al., supra at pages 2.10.1 to 2.10.16 and Sambrook, J. et al. (2001) at sections 1.101 to 1.104.
[0191] While stringent washes are typically carried out at temperatures from about 42.degree. C. to 68.degree. C., one skilled in the art will appreciate that other temperatures may be suitable for stringent conditions. Maximum hybridisation rate typically occurs at about 20.degree. C. to 25.degree. C. below the Tm for formation of a DNA-DNA hybrid. It is well known in the art that the Tm is the melting temperature, or temperature at which two complementary polynucleotide sequences dissociate. Methods for estimating Tm are well known in the art (see Ausubel et al., supra at page 2.10.8). In general, the Tm of a perfectly matched duplex of DNA may be predicted as an approximation by the formula:
T.sub.m=81.5+16.6 (log.sub.10 M)+0.41(% G+C)-0.63(% formamide)-(600/length)
[0192] wherein: M is the concentration of Na.sup.+, preferably in the range of 0.01 molar to 0.4 molar; % G+C is the sum of guanine and cytosine bases as a percentage of the total number of bases, within the range between 30% and 75% G+C; % formamide is the percent formamide concentration by volume; length is the number of base pairs in the DNA duplex. The T.sub.m of a duplex DNA decreases by approximately 1.degree. C. with every increase of 1% in the number of randomly mismatched base pairs. Washing is generally carried out at T.sub.m-15.degree. C. for high stringency, or T.sub.m-30.degree. C. for moderate stringency.
[0193] In one example of a hybridisation procedure, a membrane (e.g., a nitrocellulose membrane or a nylon membrane) containing immobilized DNA is hybridised overnight at 42.degree. C. in a hybridisation buffer (50% deionized formamide, 5.times.SSC, 5.times.Denhardt's solution (0.1% ficoll, 0.1% polyvinylpyrollidone and 0.1% bovine serum albumin), 0.1% SDS and 200 mg/mL denatured salmon sperm DNA) containing labelled probe. The membrane is then subjected to two sequential medium stringency washes (i.e., 2.times.SSC, 0.1% SDS for 15 min at 45.degree. C., followed by 2.times.SSC, 0.1% SDS for 15 min at 50.degree. C.), followed by two sequential higher stringency washes (i.e., 0.2.times.SSC, 0.1% SDS for 12 min at 55.degree. C. followed by 0.2.times.SSC and 0.1% SDS solution for 12 min at 65-68.degree. C.
[0194] 3. Preparing Prothrombin Activators
[0195] Prothrombin activators may be prepared by any suitable procedure known to those of skill in the art.
[0196] For example, the prothrombin activators may be produced by any convenient method such as by purifying the polypeptide from naturally-occurring reservoirs, including but not limited to snake venom, blood and blood-derived products (e.g. serum). Methods of purification include affinity chromatography, including lectin (e.g. wheat germ agglutinin) affinity chromatography or other separation. The identity and purity of derived prothrombin activator can be determined for example by SDS-polyacrylamide electrophoresis or chromatographically such as by high performance liquid chromatography (HPLC). For example, the purification and characterisation of pseutarin C (also abbreviated to PtPA) from P. textilis snake venom is described in Masci (1986) and Masci et al. (1988), and oscutarin C (OsPA) from O. scutellatus venom is described in Speijer et al. (1986), both of which are incorporated by reference in their entirety. The purification of ecarin from E. carinatus venom is described in Morita, T et al. (1981), the contents of which is also incorporated by reference in its entirety.
[0197] Alternatively, the prothrombin activators may be produced from venom gland cells in culture using methods known in the art, including for example the method described in Yamanouye, N., et al. (2007), which describes the primary culture of secretory cells from the venom gland of Bothrops jararaca for in vitro venom production, the contents of which is incorporated by reference in its entirety.
[0198] Alternatively, the prothrombin activators may be synthesised by chemical synthesis, e.g. using solution synthesis or solid phase synthesis as described, for example, in Chapter 9 of Atherton and Shephard (1989) and in Roberge et al. (1995).
[0199] Alternatively, the prothrombin activators may be prepared by recombinant techniques. For example, the prothrombin activators used in the invention may be prepared by a procedure including the steps of: (a) preparing a construct comprising a polynucleotide sequence that encodes a polypeptide and that is operably linked to a regulatory element; (b) introducing the construct into a host cell; (c) culturing the host cell to express the polypeptide; (d) isolating the polypeptide from the host cell. If the prothrombin activator comprises a complex or two polypeptides, then the prothrombin activator may be prepared by a procedure including the steps of: (a) preparing a construct comprising a polynucleotide sequence that encodes a first polypeptide and that is operably linked to a regulatory element; (b) introducing the construct into a host cell; (c) culturing the host cell to express the first polypeptide; (d) isolating the polypeptide from the host cell; repeating steps (a) to (d) for a second polypeptide; and linking the first polypeptide and the second polypeptide. In illustrative examples, the nucleotide sequence that encodes a polypeptide encodes at least a biologically active portion of the sequences set forth in SEQ ID NO: 5, 6, 9, 10, 14, 15, 17, 19, 20, 21, 22, 23, 24, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, or a variant thereof.
[0200] Recombinant prothrombin activators can be conveniently prepared using standard protocols as described for example in Sambrook, J. et al. (2001), in particular Chapters 16 and 17; Ausubel et al. (1994, supra), in particular Chapters 10 and 16; and Coligan et al., Current Protocols in Protein Science (John Wiley & Sons, Inc. 1995-1997), in particular Chapters 1, 5 and 6. For example, the recombinant production of snake factor V and snake factor X, which can be used to produce group C and group D prothrombin activators, is described in Filippovic, I. et al (2005) and Bos, M. H. A. et al (2009), each of which is incorporated herein in its entirety. An illustrative process for the recombinant production of ecarin and variants of ecarin is provided in Yonemura, H. et al. (2004) and in U.S. Pat. No. 6,413,737, the entire contents of each of which are incorporated herein by reference.
4. Containers
[0201] The present invention contemplates any suitable container for preparing a suitable serum sample. Many suitable containers are well known in the art, including those described in U.S. Pat. Nos. 4,227,620; 4,256,120; 6,416,717; 6,592,613; 6,686,204; 7,488,287; 7,699,828; European patent 0 628 816; and commercially available containers including those used in the Examples of the present specification.
[0202] In some embodiments, the containers used in accordance with the present invention are tubes, including glass or plastic tubes. Suitable plastics include polyvinyl chloride, polypropylene, polyethylene terephthalate, and polystyrene.
[0203] The containers may be evacuated and the end sealed with an appropriate puncturable septum or cap. This allows for a double-ended needle to be used where one end is inserted into a patient's vein and the other end of the needle then punctures the septum or cap covering the end of the tube so that the vacuum in the tube draws the blood sample through the needle into the tube.
[0204] The containers may be of any suitable size. In some embodiments, the containers are designed to hold a blood sample of between 50 .mu.L and 10 mL. Suitably, the containers are designed to hold at least 50 .mu.L, 100 .mu.L, 150 .mu.L, 200 .mu.L, 250 .mu.L, 300 .mu.L, 350 .mu.L, 400 .mu.L, 450 .mu.L, 500 .mu.L, 1 mL, 2 mL, 3 mL, 4 mL, 5 mL, 8 mL, or 10 mL of blood sample.
[0205] In some embodiments, the containers contain a clotting composition comprising, consisting essentially of, or consisting of 0.01 to 100 .mu.g of prothrombin activator. Suitably, the containers contain a clotting composition comprising, consisting essentially of, or consisting of 0.1 to 10 .mu.g of prothrombin activator. In representative embodiments, the containers contain a clotting composition comprising, consisting essentially of, or consisting of 0.1 to 10 .mu.g of prothrombin activator and hold a 4 mL blood sample providing a final concentration of prothrombin activator in the 4 mL blood sample of 25 ng/mL to 2.5 .mu.g/mL.
[0206] In some embodiments, the containers contain a clotting composition comprising, consisting essentially of, or consisting of 0.0005 to 15 .mu.g/mL of prothrombin activator. Suitably, the containers contain a clotting composition comprising, consisting essentially of, or consisting of 0.005 to 10 .mu.g of prothrombin activator.
[0207] In some embodiments, the containers contain a clotting composition comprising, consisting essentially of, or consisting of 0.0005 to 15 U/mL of prothrombin activator. Suitably, the containers contain a clotting composition comprising, consisting essentially of, or consisting of 0.0015 to 10 U/mL of prothrombin activator. In some embodiments, a unit meansurement is a prothrombin activator unit measurement that is defined as follows: 1 unit will activate prothrombin to produce one unit of amidolytic activity at pH 8.4 at 37.degree. C., and 1 amidolytic unit will hydrolyzs 1.0 .mu.mole of N-p-tosyl-Gly-Pro-Arg-p-nitroanilide per min at pH 8.4 at 37.degree. C.
[0208] In some embodiments, the clotting composition may be contained within the container before the blood sample is added to the container. In some embodiments, the clotting composition may be added to the container after the blood sample is added to the container.
[0209] Where the clotting composition is contained within the container before the blood sample is added, it may have been added to the container by any suitable method known in the art. In some embodiments, the clotting composition is dissolved into a suitable solvent and is then added to the container and dried onto the inner surface of the container. The solvent may be a neutral buffer. The clotting composition in solution may be dried onto the inner surface of the container by spray-drying or by freeze-drying or by any other suitable method known in the art. In some other embodiments, the clotting composition is dissolved into a suitable solvent and added to the container without drying so that the container contains an aqueous solution comprising the clotting composition. The solvent may be a neutral buffer.
[0210] In some embodiments, beads are coated with the clotting composition and these beads are added to the container. The beads may be glass beads or synthetic resinous beads, including polystyrene and propylene beads. The beads may have a spherical shape. In some embodiments, the mean diameter of the beads is between 0.1 mm and 1 mm.
[0211] In some embodiments, the container provides for separation of the serum from the clotted cells after clotting has occurred. In some embodiments, the container comprises or contains a gel that provides a barrier between the clotted cells and the serum sample. In some embodiments, the container is a suitable shape and a suitable material to permit centrifugation to separate or assist in maintaining separation of the clotted cells and the serum sample. In some embodiments, the serum sample is removed from the clotted cells, or the clotted cells are removed from the serum sample.
[0212] In some embodiments, the container may comprise one or more further components. The other components may include, for example, one or more co-factors, one or more surfactants, and/or one or more clotting agents in addition to the clotting composition.
5. Further Components
[0213] The clotting composition described herein consists of, consists essentially of, or comprises a prothrombin activator as herein defined.
[0214] As used in the above statement and in similar statements elsewhere in this specification, the term "comprises" (and the like) means the clotting composition includes the prothrombin activator and may also include any one or more further components. Thus, the prothrombin activator is a mandatory component, and any other components are optional and may or may not be present. Other components may include, for example, one or more co-factors, one or more surfactants, and/or one or more additional clotting agents.
[0215] As used in the above statement and in similar statements elsewhere in this specification, the term "consists essentially of" (and the like) means that the clotting composition includes the prothrombin activator, and may also include one or more other components, provided those components do not interfere with or contribute to the activity or action of the prothrombin activator. Thus, the prothrombin activator is a mandatory component, and other components are optional and may or may not be present, depending upon whether or not they affect the activity or action of the prothrombin activator.
[0216] As used in the above statement and in similar statements elsewhere in this specification, the term "consists of" (and the like) means that the clotting composition includes, and is limited to, the prothrombin activator. Thus, the phrase "consists of" indicates that the prothrombin activator is a mandatory component, and that no other components (such as co-factors or clotting agents) may be present.
[0217] Accordingly, in some embodiments, the clotting composition may comprise snake venom, including but not limited to crude snake venom. In some other embodiments, the clotting composition may comprise a preparation of prothrombin activator prepared by partial or full purification of snake venom. Such preparations may be prepared by any suitable method known in the art, including chromatographic and gel filtration methods, including those described herein, and elsewhere. In some other embodiments, the clotting composition may comprise a purified prothrombin activator or isolated prothrombin activator. Purified and isolated prothrombin activators may be prepared by any suitable method known in the art, including those described herein, and elsewhere.
[0218] 5.1 Co-Factors
[0219] The ability of prothrombin activators as herein defined to activate prothrombin to thrombin may be improved with the addition of co-factors, including but not limited to calcium, phospholipid(s) and polypeptides comprising FVa activity.
[0220] 5.2 Surfactants
[0221] Suitable surfactants include any suitable surfactant, including but not limited to sodium dodecyl sulphate (SDS), ammonium lauryl sulphate, sodium laureth sulphate, and sodium myreth sulphate.
[0222] 5.3 Clotting Agents
[0223] Clotting agents or coagulants are classified as either intrinsic clotting agents or extrinsic clotting agents according to the blood cascade stimulated (see for example U.S. Pat. No. 6,686,204).
[0224] Suitable clotting agents include, but not limited to, diatomaceous earth, microparticles or particles of inorganic silicates, microsilica, glass microparticles, ellagic acid, thrombin, heparinase, thromboplastin, batroxobin, hydroyapitite, kaolin, kaolin particles, prothrombin (including microparticulated prothrombin), fibrinogen, and depolymerised collagen.
6. Serum Samples
[0225] As discussed above, the present invention is predicated in part on the discovery that prothrombin activators as defined herein are suitable clotting agents for preparing serum samples that are suitable for detecting analytes. A serum sample that is suitable for detecting analytes is one of suitable quality as discussed herein, and/or one that is prepared within a suitable time as discussed herein.
[0226] 6.1 Serum Quality
[0227] An important factor in the preparation of a serum sample suitable for detecting analytes is the extent to which the clotting process removes fibrinogen from the serum. Serum containing residual fibrinogen or partially degraded fibrinogen, or fibrin as a result of incomplete clotting can lead to analytical accuracy problems because of the formation of precipitates (microclots or strings), latent clotting post-centrifugation and on storage of the serum. Hence complete or substantially complete clotting is pivotal in ensuring highest quality serum and subsequent result accuracy is obtained.
[0228] Accordingly, some embodiments of the present invention provide the use of a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator in the preparation of a serum for detecting an analyte, where the serum comprises <30 .mu.g/mL of fibrinogen or fibrinogen/fibrin related products. In more specific embodiments, the serum comprises <25 .mu.g/mL, <20 .mu.g/mL, <15 .mu.g/mL, <10 .mu.g/mL, <8 .mu.g/mL, or <6 .mu.g/mL of fibrinogen or fibrinogen/fibrin related products.
[0229] In some embodiments, the serum comprises <30%, <20%, <10%, <9%, <8%, <7%, <6%, <5%, <4%, <3%, <2%, <1%, <0.5%, <0.2%, <0.1% of fibrinogen or fibrinogen/fibrin related products present in the original sample from which the serum was produced.
[0230] Levels of fibrinogen and/or fibrinogen/fibrin related products can be detected by any suitable method known in the art, including a sandwich immunoassay using antibodies from MP Biomedicals and standard fibrinogen preparations purchased from NIBSC, Potters Bar, Hertsfordshire, London, UK.
[0231] Another important factor in the preparation of a serum sample suitable for detecting analytes is the activity or number of cells or cellular debris that remain in the serum after clotting. The presence of cells can have two effects during storage and analysis of serum or plasma. Firstly, cells may lyse, releasing cellular contents (e.g. potassium, lactate dehydrogenase) into the serum or plasma. This can lead to significant differences between measurements made immediately after centrifugation and measurements after a period of storage. Secondly, cells continue to be metabolically active and may use up significant amounts of nutrients (e.g. glucose) and release metabolic products (e.g. lactate) on storage. Changes can even be observed in the samples of many tubes when the samples are stored for the usual recommended 30 minute clotting time when the samples are from healthy participants. The degree of cellular contamination is therefore an important quality criterion for serum samples and an important advantage of using serum over plasma.
[0232] Accordingly, in some embodiments, the serum sample comprises less than 50%, 40%, 30%, 20%, 10%, 5%, or 1% of cells in the blood sample from which it has been prepared.
[0233] In some embodiments, the serum sample comprises a change of lactate dehydrogenase activity or phosphate concentration (typically measured in U/L and mmol/L respectively) of <25%, <20%, <15% or <10% over a period of 24 hours, 12 hours, 8 hours, 6 hours, 5 hours, 4 hours, 3 hours, 2 hours, 1 hour or 30 minutes . In some embodiments, the serum sample comprises a change of glucose concentration or potassium concentration (both typically measured in mmol/L) of <5%, <4%, <3%, <2%, <1%, <0.5%, or <0.1% over a period of 24 hours, 12 hours, 8 hours, 6 hours, 5 hours, 4 hours, 3 hours, 2 hours, 1 hour or 30 minutes (for example, from the time of preparing the serum sample). Methods for measuring lactate dehydrogenase activity are well known in the art, see, for example, Dimeski, G., et al. (2004), the contents of which is incorporated by reference in its entirety.
[0234] The haemoglobin concentration of a serum sample can also be used to determine whether the serum sample is suitable for detecting analytes. Accordingly, in some embodiments, the serum sample comprises a haemoglobin concentration of <150 mg/L, <100 mg/L, <90 mg/L, <80 mg/L, <70 mg/L, <60 mg/L, <50 mg/L, <40 mg/L, <30 mg/L, <20 mg/L, or <10 mg/L.
[0235] 6.2 Clotting Time
[0236] As a sample for testing, serum is usually preferred over plasma unless urgent results are required and thus the clotting time for a serum tube is considered too long. Another downside to prolonged clotting time is that it can lead to clinically significant analyte concentration changes due to cellular activity in the blood sample, this problem being most pronounced in leukocytosis.
[0237] Thus in some embodiments, the present invention provides a method of producing a serum sample for detecting an analyte of interest, the method comprising contacting a blood sample with a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator as defined herein, where the serum sample is prepared within 25, 20, 15, 10, 8, 6, 5, 4, 3, 2, 1, or 0.5 minutes from contact with the clotting composition.
7. Blood Samples
[0238] As discussed herein, there is a desire to provide a clotting composition that is suitable for producing a serum sample from all blood samples, or a container comprising a clotting composition that will clot all blood samples, in a suitable time.
[0239] Examples of different types of blood sample for which testing may be desired include blood from healthy individuals, citrated blood, blood with EDTA added, blood from patients on anti-clotting therapy such as heparin, warfarin, citrate, or rivaroxaban, patients taking anti-thrombotic agents including aspirin, thrombocytopenic patients (patients with low platelet counts), and patients with prolonged aPTT.
[0240] In some embodiments, the blood sample is a whole blood sample. In some other embodiments, the blood sample is a serum sample derived from a whole blood sample. Exemplary serum samples in this instance include serum samples where a better quality serum sample is desired, including those where the amount of fibrinogen or fibrinogen/fibrin related products and/or the amount of cells or cellular material in the serum sample and/or the amount of haemoglobin is considered too high for the serum sample to be a sample suitable to detect analytes. For example, the serum sample may exhibit microclots or latent clotting. In some other embodiments, the blood sample is a plasma sample derived from a whole blood sample. For example, the plasma sample may exhibit microclots or latent clotting.
8. Detecting Analytes
[0241] In some embodiments the present invention further provides methods of detecting an analyte, the method comprising analysing a serum sample prepared by the method of the present invention for the presence or amount of the analyte of interest.
[0242] In specific embodiments, the serum sample prepared by the method of the present invention is suitable for more than one analyte test, so that the serum sample can be used to detect more than one analyte. As discussed herein, often a clinician will desire more than one analyte test to be performed on a blood sample from a patient, and it is not uncommon for one serum sample to be used for at least 20 tests, or even more, sometimes between 50-60 or even 70-80 tests. It will be appreciated by those skilled in the art that in specific embodiments the present invention provides for the production of a serum sample where the serum sample is of sufficient volume and quality to enable all desired analyte tests to be performed on the one serum sample. The advantage of this is that both the volume of blood to be taken from the subject and the time taken to perform the analyte tests are reduced.
[0243] Illustrative analyte tests are described below. Methods for performing these analyte tests may be performed in a number of ways and are well known in the art.
[0244] 8.1 Sodium (Na.sup.+)
[0245] This test measures the amount of sodium in a serum or plasma sample. Sodium plays an important role in salt and water balance in the body. Low sodium levels may indicate too much water intake, heart failure, kidney failure, or loss of sodium from the body due to diarrhea or vomitting. High sodium levels may indicate too much salt intake or insufficient water intake.
[0246] 8.2 Potassium (K.sup.+)
[0247] This test measures the amount of potassium in a serum or plasma sample. Levels of potassium that are too high (hyperkalaemia) may be the result of kidney disease, diabetes, ketoacidosis or drugs that decrease the amount of potassium excreted from the body. Levels of potassium that are too low (hypokalaemia) may be caused by dehydration, for example from diarrhoea or vomiting, or excessive sweating. Levels of potassium may also be low as a result of taking drugs that cause the kidneys to lose potassium, for example diuretics.
[0248] Potassium levels are often monitored in those patients that take diuretics or heart medications, those with high blood pressure or kidney disease, critical acidosis and alkalosis conditions, and those receiving kidney dialysis or intravenous therapy on a drip.
[0249] 8.3 Chloride (Cl.sup.-)
[0250] This test measures the amount of chloride in serum or plasma. Chloride is typically measured to assess whether there is an electrolyte imbalance in the patient. Low chloride and normal sodium can be indicative of vomiting or loss of gastric fluid.
[0251] 8.4 Bicarbonate (HCO.sub.3.sup.-)
[0252] This test measures the amount of three forms of carbon dioxide (bicarbonate, carbonic acid, and dissolved carbon dioxide) in serum or plasma. This test is often performed if the patient is having breathing problems. A high level of carbon dioxide may be caused by some diseases including chronic obstructive pulmonary disease, emphysema, pneumonia, Cushing's syndrome or alcoholism, or vomiting. A low level may be caused by some diseases including pneumonia, cirrhosis, hyperventilation, diabetes, dehydration, kidney or heart failure.
[0253] 8.5 Glucose (Gluc)
[0254] This test measures the amount of glucose in serum or plasma. Glucose levels are often tested in those patients exhibiting symptoms of high blood glucose (hyperglycaemia) or hypoglycemia, those who are pregnant, those who have diabetes.
[0255] 8.6 Urea (Urea)
[0256] This test measures the amount of urea in serum or plasma. This test can help evaluate kidney function and monitor the effectiveness of dialysis.
[0257] 8.7 Creatinine (Creat)
[0258] This test measures the amount of creatinine in serum or plasma. This test is pivotal in helping to evaluate kidney function and monitor treatment of kidney disease.
[0259] 8.8 Urate (Urate)
[0260] This test measures the amount of urate (or uric acid) in serum or plasma. High levels of uric acid may be a sign of gout. Uric acid levels are also monitored in patients that are undergoing chemotherapy or radiotherapy to detect tumour lysis syndromes.
[0261] 8.9 Total Protein (TP or T Prot)
[0262] This test measures the total amount of protein in serum or plasma. Although the results of a total protein test will not indicate a specific disease, a high or low protein level often indicates that additional tests are required to determine if there is a problem. Total protein tests are often used to screen for certain liver disorders, kidney disorders, multiple myeloma and hydration status.
[0263] 8.10 Albumin (Alb)
[0264] This test measures the amount of albumin in serum or plasma. Albumin levels are often measured to screen for liver or kidney disease, or to evaluate nutritional status, especially in hospitalised patients.
[0265] 8.11 Total Bilirubin (T Bili)
[0266] This test measures the amount of bilirubin in serum or plasma. Bilirubin levels are measured to screen for and monitor liver disorders, such as jaundice, or liver diseases, such as cirrhosis. Bilirubin levels are also measured in babies to help detect certain rare genetic disorders and to avoid brain damage in those babies with jaundice.
[0267] 8.12 Alkaline Phosphatase (ALP)
[0268] This test measures the amount of alkaline phosphatase in serum or plasma. This test is typically performed to screen for or monitor treatment of a liver or bone disorders.
[0269] 8.13 Gamma-Glutamyl Transferase (GGT)
[0270] This test measures the amount of gamma-glutamyl transferase in serum or plasma. This test is used to screen for liver disease and alcohol abuse. It can also be used to determine if a raised level of ALP is due to liver or bone disease.
[0271] 8.14 Alanine Aminotransferase (ALT)
[0272] This test measures the amount of alanine aminotransferase in serum or plasma. This test is used to screen for liver disease.
[0273] 8.15 Aspartate Aminotransferase (AST)
[0274] This test measures the amount of aspartate aminotransferase in serum or plasma. This test is used to detect liver damage, muscular damage, and other conditions as the enzyme is present in many organs and tissue cells.
[0275] 8.16 Lactate Dehydrogenase (LD)
[0276] This test measures the amount of lactate dehydrogenase in serum or plasma. This test is typically used to identify the cause and location of tissue damage in the body, tissue ischemia, and to monitor its progress.
[0277] 8.17 Creatine Kinase (CK)
[0278] This test measures the amount of creatine kinase in serum or plasma. Creatine kinase is measured in patient's with chest pain or muscle pain or weakness to determine if they have had a heart attack and if other muscles in the body have been damaged.
[0279] 8.18 Total Calcium (TCa)
[0280] This test measures the amount of calcium in serum or plasma. Calcium levels are often measured in patients with kidney, bone or nerve disease, or when symptoms of significantly increased or decreased calcium are present.
[0281] 8.19 Phosphate (Pi or Phos)
[0282] This test measures the amount of phosphate in serum or plasma. Phosphate levels may be measured as a follow-up to a test result of abnormal calcium levels. Phosphate levels may also be measured in patients with kidney disorders, uncontrolled diabetes, or where the patient is taking calcium or phosphate supplements.
[0283] 8.20 Magnesium (Mg.sup.2+)
[0284] This test measures the amount of magnesium in serum or plasma. This test may be performed if the patient has symptoms of too much or too little magnesium, including weakness, irritability, cardiac arrhythmia, nausea or diarrhoea. Magnesium levels may also be measured if abnormal calcium or potassium levels have been detected.
[0285] 8.21 Lipase (Lipase)
[0286] This test measures the amount of lipase in serum or plasma. This test is typically used to diagnose pancreatitis or other pancreatic diseases.
[0287] 8.22 Cholesterol (Chol)
[0288] This test measures the amount of cholesterol in serum or plasma. Cholesterol levels are measured to screen for risk of developing heart disease.
[0289] 8.23 Triglycerides (Trig)
[0290] This test measures the amount of triglycerides in serum or plasma. As for cholesterol levels, this test is typically used to screen for risk of developing heart disease.
[0291] 8.24 High-Density Lipoprotein Cholesterol (HDL-C or HDL)
[0292] This test measures the amount of HDL cholesterol in serum or plasma. This test is typically used to determine the risk of developing heart disease.
[0293] 8.25 Iron (Fe.sup.2+)
[0294] This test measures the amount of iron in serum or plasma. Iron is measured to check if a patient has low or high iron levels. Low iron levels can cause anaemia, and is usually due to long-term or heavy bleeding, pregnancy or rapid growth (in children). High iron levels can be due to a genetic condition or extensive blood transfusions.
[0295] 8.26 Transferrin (Trf)
[0296] This test measures the amount of transferrin in serum or plasma. Transferrin is a plasma protein that transports iron through the blood to the liver, spleen and bone marrow. Thus the blood transferrin level is tested to determine the cause of anaemia, to examine iron metabolism (for example, in iron deficiency anaemia) and to determine the iron-carrying capacity of the blood.
[0297] 8.27 C Reactive Protein (CRP)
[0298] This test measures the amount of C reactive protein in serum or plasma. This test is used to identify the presence of inflammation, to determine its severity, and to monitor response to treatment.
[0299] 8.28 Cortisol (Cortisol)
[0300] This test measures the amount of cortisol in serum or plasma. Cortisol levels are measured to help diagnose Cushing's syndrome or Addison's disease.
[0301] 8.29 Free Thyroxine (fT4)
[0302] This test measures the amount of free thyroxine in serum or plasma. The test is typically used to diagnose hypothyroidism or hyperthyroidism.
[0303] 8.30 Thyroid Stimulating Hormone (TSH)
[0304] This test measures the amount of thyroid stimulating hormone in serum or plasma. The test is typically used to screen for, diagnose and monitor thyroid disorders.
[0305] 8.31 Ferritin
[0306] This test is used to measure ferritin in serum or plasma. Low ferritin levels are indicative of iron deficiency. Elevated levels are indicative of iron overload such as in haematochromatosis.
[0307] 8.32 Troponin (TnI)
[0308] This test measures the amount of troponin in serum or plasma. This test is typically used in a patient with chest pains to determine if the patient has had myocardial damage.
[0309] 8.33 Haemolytic Index
[0310] The haemolytic index test measures the degree of red cell lysis. Haemolysis is the most common interference encountered in a biochemistry laboratory. The test is predominantly used to detect in vitro haemolysis and sample suitability for reporting of certain or all analytes, and in detection of haemolytic anaemias (hereditary spherocytosis, spontaneous haemolysis, RBC enzyme deficiency). Haemolysis or haemolytic index (concentration of free haemoglobin in serum or plasma) is currently estimated by all general chemistry analysers. The value is then used as a guide in determining which analytes and at what haemolysis level may be affected or not reported.
[0311] 8.34 Icteric Index
[0312] The icteric index test returns a value indicating the relative level of bilirubin in a test sample by a purely spectrophotometric method. It is used in determining sample suitability for reporting of certain analytes and cross checking accuracy of bilirubin results in rare cases of interference with the total bilirubin photometric estimation methods. The icteric index has been shown to be of value in detecting cancer paraproteins interference (precipitation and false high total bilirubin) with Roche Total Bilirubin method (Sheppard et al., 2005), where the icteric index has stayed unaffected. Bilirubin can interfere with some creatinine assays at high concentration (e.g. >200 .mu.M/L) as discussed in Dimeski et al., 2008.
[0313] 8.35 Lipemia Index
[0314] The lipemia index has been employed to predict possible interference with assays due to lipaemia.
[0315] 9. Methods of Diagnosis
[0316] The present invention also provides methods of diagnosing the presence, absence or severity of a disease or condition in a subject, wherein the presence, absence or severity of the disease or condition is associated with the presence, absence or an aberrant amount of an analyte of interest in the subject. These methods generally comprise providing a serum sample prepared according to the methods broadly described above; and detecting the presence, absence or aberrant amount of the analyte in the serum sample to thereby determine the presence, absence or severity of the disease or condition in the subject.
[0317] In some embodiments, the methods of the present invention involve comparing the result of the analyte test to a reference result in order to obtain the diagnosis.
[0318] The disease or condition may be any suitable disease or condition that can be diagnosed using a serum sample, including but not limited to, the diseases or conditions outlined above with reference to different analyte tests.
[0319] In some embodiments, the methods may comprise diagnosing the presence or absence of a disease or condition not previously presented by the subject. In other embodiments, the methods may comprise diagnosing the presence, absence or severity of a disease or condition that the subject has previously presented. The methods may comprise reference to a result obtained from the subject at an earlier time. Alternatively, the reference result may be a standard analytical reference.
[0320] In some embodiments, the methods are performed in a testing facility such as a pathology laboratory. In some other embodiments, the methods are "point-of-care" methods. As used herein, a "point-of-care method" means that the method is performed at or near the site of patient care. Point-of-care methods are increasingly popular in hospital and other environments where there is a desire to obtain fast results, and is often accomplished through the use of transportable, portable, and hand-held instruments and test kits.
[0321] The advantages of point of care testing include the ability to obtain rapid analytical results at the bedside in hospitals, especially in emergency situations and the ability to obtain analytical results at home or in doctors' surgeries (e.g., using droplets of capillary blood obtained by skin puncture).
[0322] Devices for point-of-care devices currently available on the market include the immunoassay analyser sold by Siemens called Siemens' DCA Vantage Analyzer, the Retro-STATUS HIV/CD4 350 rapid test device sold by Millenium Biotechnology, Inc. which is used to simultaneously determine a patient's HIV infection status as well as their current immune status, and the Triage PLGF test sold by Alere International which is used for detecting early onset pre-eclampsia.
10. Research Tools
[0323] The present invention also contemplates the use of research tools that employ serum samples produced in accordance with the present invention. These methods generally comprise providing a serum sample prepared according to the methods broadly described above; and employing the serum sample in a research tool study, including but not limited to a genomics, proteomics, metabolomics, systems biology, molecular imaging or assay study.
[0324] Suitable research tools are well known in the art and include those described in Scaros, O. et al., 2005, the entire contents of which are incorporated by reference. Genomics includes pharmacogenomics which studies the correlation between genetics and gene expression patterns with response to therapeutics. Proteomics permits the analysis of the abundance and distribution of proteins in a system. Metabolomics, or biochemical profiling, is the study of metabolites in a system. Systems biology looks at the entire biological system as a functional unit, producing models of behaviour that can potentially predict hwo that system will respond to stimulus. Molecular imaging technologys have the ability to demonstrate both the level of a specific molecular target and the functional state of that target in vivo, and can be used for diagnostic methods.
[0325] In order that the invention may be readily understood and put into practical effect, particular preferred embodiments will now be described by way of the following non-limiting examples.
EXAMPLES
Example 1: Purification and Characterisation of Prothrombin Activators from Snake Venom
Example 1a: Purification and Characterisation of Ecarin, Carinactivase-1 and Carinactivase-2 from Echis Carinatus Venom
[0326] This example describes the purification and characterisation of a group A prothrombin activator termed ecarin and two group B prothrombin activators termed carinactivase-1 (CA-1) and carinactivase-2 (CA-2) from the venom of Echis carinatus.
[0327] Freeze-dried Echis carinatus venom (157 mg; Sigma Chemical Co, USA; cat no. V8250) was dissolved in 8 mL of 0.05 M Tris-HCl pH 8.0 buffer and left standing to allow to dissolve for 30 minutes and then centrifuged (1000 g and 10 minutes) to remove insoluble materials. A volume of 7.7 mL of the reconstituted clarified solution was loaded onto a Superdex 200 gel filtration chromatography column (cat no 17-1043-02) (2.5.times.95 cm) GE Healthcare Bio-Sciences, Sweden) using the same Tris-HCl buffer for the elution. The column flow rate was 20 mL/hr and 4 mL fractions were collected (15 minute fraction). The active fractions were determined by their clotting activity with normal citrated plasma as described in more detail below. The pooled fractions exhibiting prothrombin activator activity contained a total of 100 mL with a total A.sub.280 of 28.3 units. FIG. 1 shows the elution profile in the isolation of ecarin, carinactivase-1, and carinactivase-2 from E. carinatus venom using gel filtration on a Superdex 200 column as described above. The pooled fractions of the three prothrombin activators (ecarin, carinactivase-1, and carinactivase-2) are indicated by the bar in FIG. 1.
[0328] These pooled fractions were immediately applied to a column of Blue Sepharose (2.0.times.12 cm) (Cibacron Blue 3G attached to Sepharose 6 Fast Flow; cat no. 17-0948-01) (GE Healthcare Bio-Sciences, Sweden) pre-equilibrated with the same 0.05 M Tris-HCl buffer pH 8.0. After washing with initial buffer of approximately 2 column volumes, the bound material was eluted with a linear gradient of NaCl in the same buffer (0-1.0 M; 200 mL of each beaker). Carinactivase-1 was recovered in the unbound fractions and carinactivase-2 and ecarin were eluted at 0.2 and 0.5 M NaCl respectively. Prothrombin activator activities were identified by normal citrated plasma clotting assay in the presence of 30 mM calcium and in the absence of added calcium as described in more detail below. Fractions (three mL each) with specific prothrombin activator activities were pooled by their respective specific clotting activity of the clotting times followed by concentrating to 1 mg/mL using a YM 10 membrane in an Amicon stirred cell concentrator Model no. 42. All chromatography was performed at 4.degree. C. using an Gilson peristaltic pump using black to purple chamber set at the required flow rates, an Altex dual wave UV detector set at two attenuation of 280 nm and an LKB 7000 ULTRORAC fraction collector using a time-base fraction collection. FIG. 2 shows the elution profile of this Blue Sepharose chromatography, and the bars indicate the pooled fractions of carinactivase-1, carinactivase-2, and ecarin. The fractions obtained were stored at either 4.degree. C., -20.degree. C. in buffer or -20.degree. C. in 50% glycerol-buffer. These fractions (preparations) containing prothrombin activators were used in all subsequent experiments, except where otherwise indicated.
[0329] The prothrombin activator activity was assayed using the method described by Masci et al., 1988. Essentially, this assay method involved coagulation assays performed using a Hyland-Clotek instrument as described by Austen, D. E., et al. 1975. Fresh-pooled citrated plasma from normal volunteers was used for each group of experiments. The clotting time assays for the preparations were performed based on the criteria defined for group A and group B prothrombin activators in Kini, R. M., 2005. The assays were carried in the presence of calcium (30 mM Ca.sup.2+) to identify the group B prothrombin activators (carinactivase-1 and carinactivase-2) and in the absence of added calcium to identify the group A prothrombin activator (ecarin). The hydrolytic activity against the peptide p-nitroanilide S-2238, measuring the formation of thrombin generation by prothrombin activators, was determined by equilibration of 0.90 mL of 0.02 M Tris-HCl buffer, pH 7.4, with or without 10 mM CaCl.sub.2, and 100 .mu.L of S-2238 (3 mM in water) in the cell compartment of a spectrophotometer (Hitachi U2800) at 25.degree. C. The reaction was initiated by addition of a 25 .mu.L aliquot of prothrombin activator preparation to prothrombin (250 nM), and the release of p-nitroaniline was measured at 405 nm. One unit of activity is equivalent to the hydrolysis of 1 pmol of substrate/minute. Assays using other peptide nitroanilides as substrate were performed in the same way, with a final concentration of 100 .mu.M.
Example 1b: Purification and Characterisation of PtPA from Pseudonaja Textilis Venom and OsPA from Oxyuranus Scutellatus Venom
[0330] This example describes the purification and characterisation of group C prothrombin activators from the venom of Pseudonaja textilis and Oxyuranus scutellatus snakes. These prothrombin activators are abbreviated as PtPA and OsPA respectively.
[0331] Dried lyophilised P. textilis venom was reconstituted in 0.05 M Tris-HCl with a pH of 7.4. FIG. 3 provides an elution profile of the reconstituted P. textilis venom on a Sephracryl S-300 column in 0.05 M Tris-HCl buffer at pH 7.4. `A` refers to void volume and `B` denotes elution position (Ve) for PtPA which is 250 mL on this column and equates to an approximate molecular mass of 250 kDa. The absorbance at 280 nm (A.sub.280) under `B` represents 40% of the total A.sub.280 units applied to the column. A similar gel filtration experiment with O. scutellatus venom showed that approximately 10% of the total A.sub.280 loaded eluted in the 250 kDa peak (corresponding to the prothrombin activator).
[0332] Affinity chromatography using Con A-Sepharose 4B in 0.05 M Tris-HCl, pH 7.4, was used to purify the prothrombin activator from the crude P. textilis and O. scutellatus venom. The eluting buffer was 0.2 M methyl a-D-mannopyranoside with 0.01% sodium azide. The elution profile on Con A-Sepharose for PtPA is shown in FIG. 4. The arrow indicates the application of eluting buffer and "a" indicates the fractions pooled to provide prothrombin activator. FIG. 5 shows the results of native PAGE at pH 8.6. Peak "a" from FIG. 4 showed a major protein band and a trace minor band at loadings of 25 .mu.g (A) and 50 .mu.g (B) per lane. The 250 kDa band was excised from the gel in a parallel experiment and shown to have Factor Xa-like activity against the chromogenic substrate S-2222, confirming its identification as PtPA (FIG. 6). A similar pattern of elution was obtained for OsPA.
[0333] PtPA and OsPA were characterised further by SDS-PAGE in the presence (designated "red") and absence ("unred") of .beta.-mercaptoethanol (FIG. 7). It is evident that several bands are present for both PtPA and OsPA representing component polypeptide chains.
[0334] PtPA- and OsPA-containing preparations prepared in accordance with this method were used in all subsequent experiments, except where otherwise indicated.
Example 1c: Purification and Characterisation of Notecarin from Notechis Scutatus Venom
[0335] The group D prothrombin activator termed notecarin was purified from the venom of Notechis scutatus, using Sephacryl S-300 gel filtration chromatography (5.0.times.9.5 cm) in a similar manner to that described in Example lb above for the group C prothrombin activators. The elution profile is shown in FIG. 8, where "PA" refers to the fraction containing notecarin.
[0336] Notecarin-containing preparations prepared in accordance with this method were used in all subsequent experiments, except where otherwise indicated.
Example 1d: Further Characterisation of Carinactivase-1, Carinactivase-2, Ecarin, PtPA, OsPA, and Notecarin
[0337] The prothrombin activator preparations prepared in Examples 1a, 1b, and 1c were further characterised as follows.
[0338] Native PAGE of prothrombin, alpha-thrombin, and the prothrombin activators at pH 8.9 was performed and the results are shown in FIG. 9, wherein the labels are as follows: (I) prothrombin; (II) alpha-thrombin; (III) ecarin; (IV) carinactivase-1; (V) carinactivase-2; (VI) PtPA; (VII) OsPA; (VIII) notecarin (20 .mu.g of each prothrombin activator was loaded).
[0339] The prothrombin activator preparations were also characterised further by SDS-PAGE in the presence (FIG. 10) and absence (FIG. 11) of .beta.-mercaptoethanol. In each Figure, the lanes are as follows: (1) carinactivase-1; (2) carinactivase-2; (3) ecarin; (4) PtPA; (5) OsPA; (6) notecarin; (7) thrombin; and (M) molecular weight marker. The PtPA-containing sample in lane (4) and the OsPA-containing sample in lane (5) were the same as those used for the results shown in FIG. 7.
[0340] The SDS-PAGE shows that in each case the protein bands in the expected regions are present but also that the preparations are not homogeneous. The protein concentrations of the prothrombin activator preparations were calculated using the relationship that 1 mg/mL solution had an A.sub.280 of 1, and the molar concentrations were calculated by dividing the protein concentrations by the literature values for the molecular weights. The molar concentrations quoted in subsequent experiments described herein that used these preparations are therefore upper estimates because the preparations are not homogenous.
Example 2: Prothrombin to Thrombin Conversion by Prothrombin Activators
Example 2a: Five-Minute Incubation Period
[0341] The aim of this experiment was to determine how much prothrombin was converted to thrombin by six different prothrombin activators from snake venom prepared in Example 1, namely preparations containing ecarin, carinactivase-1, carinactivase-2, PtPA, OsPA, and notecarin, after incubation for five minutes at room temperature.
[0342] The experiment was performed as follows.
[0343] 20 mM HEPES Buffer pH 7.4, 150 mM NaCl, 5 mM CaCl.sub.2, 0.05% surfactant p20 was prepared.
[0344] A solution of human prothrombin in 0.05 M Tris-HCl pH 7.4 was prepared by diluting a stock solution of purified human prothrombin (17.8 mg/mL; # HCP-0200; HTI, USA) 1/50 with distilled water to provide a working solution of 0.36 mg/mL, giving a final concentration in the tube of 72 .mu.g/mL or 0.99 .mu.M. For the gel loading the 0.36 mg/L prothrombin solution was used.
[0345] The stock solutions of the prothrombin activator preparations prepared as described earlier were diluted to provide final concentrations of 6 nM.
[0346] Stock alpha-thrombin (10 mg/mL, # HCT-0020; HTI, USA) was diluted (1/30) with distilled water containing 0.01% Tween 20 to provide a working thrombin solution of 0.33 mg/L which was used to load on the SDS-PAGE gels.
[0347] Samples were then prepared with the volumes shown in Table 3, and incubated for five minutes at room temperature.
TABLE-US-00004 TABLE 3 Volumes in each sample for incubation Sample 1 2 3 4 5 6 7 8 HEPES buffer (.mu.L) 80 60 60 60 60 60 60 80 Prothrombin (.mu.L) 20 20 20 20 20 20 20 0 Prothrombin activator 0 50 50 50 50 50 50 0 .alpha.-thrombin (.mu.L) 0 0 0 0 0 0 0 20 Total volume (.mu.L) 100 100 100 100 100 100 100 100
[0348] The prothrombin activators in the above samples were as follows: (1) no prothrombin activator (human prothrombin alone in buffer); (2) carinactivase-1; (3) carinactivase-2; (4) ecarin; (5) PtPA; (6) OsPA; (7) notecarin; and (8) highly purified .alpha.-thrombin (no prothrombin activator).
[0349] SDS PAGE gels used were nUView Precast Mini Gels (# NB10-420, 4-20%) with running buffer Tris-Glycine (# BG-143). Sample buffer (# BG-145) (NuSEP, Australia) was obtained (i.e., non-reducing sample buffer), and a reducing sample buffer of the sample buffer with 5% .beta.-mercaptoethanol (#44143, BDH, UK) was prepared.
[0350] After the five minute incubation, a 40 .mu.L aliquot from each sample was transferred into equivalent volume of non-reducing sample buffer or reducing sample buffer. The samples were then incubated for 10 minutes at 100.degree. C. in a heating block. An aliquot of 25 .mu.L of each sample and 12 .mu.L of the pre-stained molecular weight marker (#5M0671 Page Ruler Prestained Protein Ladder, Fermentas, Hanover, Md., USA) were loaded on the gels. The gels were run at 100 V using a Mini-Protein II Cell PAGE apparatus (Bio-Rad) until the dye-front reached the bottom of the gel. The gels were stained with Commassie Brilliant Blue G (# B-0770, Sigma) (0.25% w/v Commassie Brilliant Blue G, 45% methanol, 10% acetic acid), and excess stain was removed by destainer (45% methanol, 45% water and 10% acetic acid).
[0351] The results are shown in FIG. 12, where FIG. 12A shows the SDS-PAGE gel using the sample buffer without .beta.-mercaptoethanol (non-reducing sample buffer) and FIG. 12B shows the SDS-PAGE gel using the sample buffer containing 5% .beta.-mercaptoethanol (reducing sample buffer). The sample numbers as used in Table 3 above are the same as the lane numbers in the gels in FIG. 12, and "m" represents the molecular weight marker.
[0352] This experiment showed complete conversion of prothrombin to thrombin in five minutes by PtPA and OsPA, but very little conversion by the other prothrombin activator preparations under these conditions.
Example 2b: Time Course for Conversion by PtPA and by Notecarin
[0353] A time course for conversion of prothrombin (14 .mu.M) to thrombin by PtPA (6 nM) and by notecarin (6 nM) was determined, and the SDS-page gel results are shown in FIG. 13, where each lane is numbered according to the incubation time as shown in Table 4 below, using methodology similar to Example 2a.
TABLE-US-00005 TABLE 4 Incubation time for each lane shown in FIG. 13. Lane Time (min) 1 0 2 0.5 3 1 4 2 5 4 6 6 7 8 8 10 9 15 10 20 11 25 12 30 13 45 14 150
[0354] These results show that PtPA is more efficient than notecarin in converting prothrombin to thrombin, but that notecarin still gave complete cleavage of prothrombin after longer reaction times.
Example 2c: N-Terminal Sequencing of Selected Bands from the Example 2b Results
[0355] Selected bands from the SDS-PAGE results in Example 2b, produced by the action of PtPA and notecarin on prothrombin (i.e., the fragmentation of prothrombin to thrombin), were eluted and subjected to N-terminal sequencing using mass-spectrometry analysis in order to assign to specific molecular domains. These results are shown in FIG. 14. The SDS-PAGE N-terminal sequencing confirmed conversion of prothrombin to alpha-thrombin by PtPA and notecarin, as shown by the major band with N-terminal sequence IVEGSDA which corresponds to the alpha-thrombin heavy chain.
Example 2d: Estimation of Thrombin Generated by Kinetic Analysis
[0356] This experiment was designed to determine the amount of thrombin generated from prothrombin by each of the prothrombin activators: ecarin; carinactivase-1; carinactivase-2; PtPA; OsPA; and notecarin, at 0.6 nM concentration, by way of kinetic analysis.
[0357] The absorbance of the p-nitroaniline (pNA) generated from the chromogenic thrombin substrate S-2238 was continuously monitored at 405 nm, for samples containing different concentrations of thrombin and for samples containing one of the prothrombin activators listed above.
[0358] For the thrombin study, each cuvette contained 50 .mu.L of S-2238 substrate (134 .mu.M), 50 .mu.L of thrombin at different concentrations (range of 0.25-30 nM) and 900 .mu.L of HEPES buffer. FIG. 15 shows progress curves for S-2238 hydrolysis by thrombin used to generate the standard curve shown in FIG. 16.
[0359] For the study of the prothrombin activators, each cuvette contained 50 .mu.L of prothrombin (247 nM), 10 .mu.L (0.6 nM) of the prothrombin activator (ecarin, carinactivase-1, carinactivase-2, PtPA, OsPA, or notecarin), 100 .mu.L (267 .mu.M) S-2238 substrate and 840 .mu.L of HEPES buffer. The reaction slope for each prothrombin activator was determined at 155 seconds and the amount of thrombin generated at 155 seconds read from the standard curve in FIG. 16, as shown in FIG. 17 and Table 5.
TABLE-US-00006 TABLE 5 Amount of thrombin generated by 0.6 nM of the prothrombin activators over 155 seconds Calculated slope Amount of thrombin Prothrombin from standard generated (nM) at 155 activator (0.6 nM) curve in FIG. 16 seconds Ecarin 0.00085 1.74 Carinactivase-1 0.00007 0.07 Carinactivase-2 0.0005 0.99 PtPA 0.0094 20.12 OsPA 0.0077 16.47 Notecarin 0.0001 0.13
[0360] All prothrombin activator preparations produced thrombin as expected, but again, the PtPA and OsPA were the most efficient.
[0361] These results indicate the group C prothrombin activators (PtPA and OsPA) were the most efficient enzymes in hydrolysing prothrombin to thrombin. This was confirmed by the amount of thrombin formed that was able to hydrolyse the S-2238 substrate to form the colour product pNA. The next most efficient prothrombin activator was ecarin which showed it hydrolysed .about.4% of the S-2238, followed by carinactivase-2 (2.5%) then carinactivase-1 and lastly notecarin (<1%) being the least efficient of the prothrombin activators in this purified system. Group D prothrombin activators (e.g., notecarin) require Factor Va, calcium and phospholipid which were absence in the system but all of which are present in blood.
Example 2e: Kinetic Studies on Comparative Rates of Thrombin Generation from Thrombin by Prothrombin Activators
[0362] This experiment was designed to determine the rate of thrombin formation that is generated from pure prothrombin by each of the prothrombin activator preparations.
[0363] Each cuvette contained 50 .mu.L of prothrombin (247 nM), 10 .mu.L of the different prothrombin activators, 100 .mu.L (267 .mu.M) of S-2238 substrate and 840 .mu.L of HEPES buffer.
[0364] The progress curves for S-2238 hydrolysis (thrombin activity) for each prothrombin activator was determined at 3-5 different concentrations ranging from 0.006-6 nM. The data was analysed using DynaFit 4.04.64 Enzyme Kinetic Data Analysis Software (BioKin Ltd, Watertown, Mass., USA) (Kuzmic, P., 1996).
[0365] The thrombin catalysed hydrolysis of S-2238 curve fitting analysis is shown in FIG. 18.
[0366] Kinetic parameters were then calculated as follows:
[0367] Model: B.fwdarw.B+pNA+P (a pseudo first order rate constant for the thrombin hydrolysis of S-2238), where:
[0368] B=thrombin;
[0369] pNA=p-nitroaniline;
[0370] P=inhibition product; and
[0371] S-2238 is in excess and therefore not included in the reaction.
[0372] Differential equations as follows:
d[B]/dt=-kb[B]+kb[B];
d[pNA]/dt=+kb[B]; and
d[P]/dt=+kb[B].
[0373] Table 6 shows the estimated kinetic parameters for thrombin.
TABLE-US-00007 TABLE 6 Estimated kinetic parameters for thrombin Units Parameter Initial Fit CV % s.sup.-1 kb 61.3 (Sonder, et al., 1986) 66.2 0.4
[0374] Estimation of rate constant for prothrombin activators:
[0375] Model:
[0376] A+Z.fwdarw.A+B: ka (second order rate constant for activation of prothrombin);
[0377] B.fwdarw.B+pNA+P: kb; where
[0378] A=prothrombin activator.
[0379] Differential equations as follows:
d[A]/dt=-ka[A][Z]+ka[A][Z];
d[Z]/dt=-ka[A][Z];
d[B]/dt=+ka[A][Z]-kb[B]+kb[B];
d[pNA]/dt=+kb[B]; and
d[P]/dt=+kb[B].
[0380] The kinetic parameters for PtPA were estimated and are shown in Table 7.
TABLE-US-00008 TABLE 7 Estimated kinetic parameters for PtPA Units Parameter Initial Fit CV % s.sup.-1 kb 66.2 s.sup.-1 nM.sup.-1 ka 0.001 6.722e-4 1.8 s.sup.-1 M.sup.-1 ka 6.72 .times. 10.sup.5 1.8
[0381] The rate of thrombin generation:
=ka(s.sup.-1M.sup.-1).times.60(s).times.10.sup.-6
=40.3 .mu.M thrombin/min/M prothrombin/M activator
=40.3 nmol thrombin/mL/min/.mu.mol prothrombin/.mu.mol activator
[0382] FIG. 19 plots the PtPA activation of prothrombin, demonstrating the reaction rates of different PtPA concentrations (0.006-0.12 nM) with prothrombin and then the thrombin generated with the chromogenic substrate S-2238, and the best fit curve for PtPA activation of prothrombin. The dotted lines are the experimental results and the continuous lines are those calculated as best fit. These experimental and calculated progress curves show good agreement.
[0383] The kinetic parameters for OsPA were then estimated and are shown in Table 8.
TABLE-US-00009 TABLE 8 Estimated kinetic parameters for OsPA Units Parameter Initial Fit CV % s.sup.-1 kb 66.2 s.sup.-1 nM.sup.-1 ka 0.001 7.325e-4 3.3 s.sup.-1 M.sup.-1 ka 7.33 .times. 10.sup.5 3.3
[0384] The rate of thrombin generation:
=44.0 nmol thrombin/mL/min/.mu.mol prothrombin/.mu.mol activator
[0385] FIG. 20 plots the OsPA activation of prothrombin, demonstrating the reaction rates of different OsPA concentrations (0.006-0.24 nM) with prothrombin and then the thrombin generated with the chromogenic substrate S-2238, and the best fit curve for OsPA activation of prothrombin. The dotted lines are the experimental results and the continuous lines are those calculated as best fit. These experimental and calculated progress curves show good agreement.
[0386] The kinetic parameters for ecarin were then estimated and are shown in Table 9.
TABLE-US-00010 TABLE 9 Estimated kinetic parameters for ecarin Units Parameter Initial Fit CV % s.sup.-1 kb 66.2 s.sup.-1 nM.sup.-1 ka 0.001 4.607e-005 1.2 s.sup.-1 M.sup.-1 ka 4.61 .times. 10.sup.4 1.2
[0387] The rate of thrombin generation:
=2.76 .mu.M thrombin/min/.mu.M prothrombin/.mu.M activator
=2.76 nmol thrombin/mL/min/.mu.mol prothrombin/.mu.mol activator
[0388] FIG. 21 plots the ecarin activation of prothrombin, demonstrating the reaction rates of different ecarin concentrations (0.24-1.2 nM) with prothrombin and then the thrombin generated with the chromogenic substrate S-2238, and the best fit curve for ecarin activation of prothrombin. The dotted lines are the experimental results and the continuous lines are those calculated as best fit. These experimental and calculated progress curves show good agreement.
[0389] The kinetic parameters for carinactivase-1 were then estimated and are shown in Table 10.
TABLE-US-00011 TABLE 10 Estimated kinetic parameters for carinactivase-1 Units Parameter Initial Fit CV % s.sup.-1 kb 66.2 s.sup.-1 nM.sup.-1 ka 0.001 3.944e-006 0.3 s.sup.-1 M.sup.-1 ka 3.94 .times. 10.sup.3 0.3
[0390] The rate of thrombin generation:
=0.24 .mu.M thrombin/min/.mu.M prothrombin/.mu.M activator
=0.24 nmol thrombin/mL/min/.mu.mol prothrombin/.mu.mol activator
[0391] FIG. 22 plots the carinactivase-1 activation of prothrombin, demonstrating the reaction rates of different carinactivase-1 concentrations (0.24-1.2 nM) with prothrombin and then the thrombin generated with the chromogenic substrate S-2238, and the best fit curve for carinactivase-1 activation of prothrombin. The dotted lines are the experimental results and the continuous lines are those calculated as best fit. These experimental and calculated progress curves show good agreement.
[0392] The kinetic parameters for carinactivase-2 were then estimated and are shown in Table 11.
TABLE-US-00012 TABLE 11 Estimated kinetic parameters for carinactivase-2 Units Parameter Initial Fit CV % s.sup.-1 kb 66.2 s.sup.-1 nM.sup.-1 ka 0.001 3.183e-005 1.8 s.sup.-1 M.sup.-1 ka 3.18 .times. 10.sup.4 1.8
[0393] The rate of thrombin generation:
=1.91 .mu.M thrombin/min/.mu.M prothrombin/.mu.M activator
=1.91 nmol thrombin/mL/min/.mu.mol prothrombin/.mu.mol activator
[0394] FIG. 23 plots the carinactivase-2 activation of prothrombin, demonstrating the reaction rates of different carinactivase-2 concentrations (0.24-1.2 nM) with prothrombin and then the thrombin generated with the chromogenic substrate S-2238, and the best fit curve for carinactivase-2 activation of prothrombin. The dotted lines are the experimental results and the continuous lines are those calculated as best fit. These experimental and calculated progress curves show good agreement.
[0395] The kinetic parameters for notecarin were then estimated and are shown in Table 12.
TABLE-US-00013 TABLE 12 Estimated kinetic parameters for notecarin Units Parameter Initial Fit CV % s.sup.-1 kb 66.2 s.sup.-1 nM.sup.-1 ka 0.001 7.139e-007 0.3 s.sup.-1 M.sup.-1 ka 7.14 .times. 10.sup.2 0.3
[0396] The rate of thrombin generation:
=0.043 .mu.M thrombin/min/.mu.M prothrombin/.mu.M activator;
=0.043 nmol thrombin/mL/min/.mu.mol prothrombin/.mu.mol activator.
[0397] FIG. 24 plots the notecarin activation of prothrombin, demonstrating the reaction rates of different notecarin concentrations (0.24-6.0 nM) with prothrombin and then the thrombin generated with the chromogenic substrate S-2238, and the best fit curve for notecarin activation of prothrombin. The dotted lines are the experimental results and the continuous lines are those calculated as best fit. These experimental and calculated progress curves show good agreement.
[0398] A summary of the estimated activation rate constants and thrombin generation rate for each of the prothrombin activators is provided in Table 13.
TABLE-US-00014 TABLE 13 Summary of the estimated activaton rate constants and the thrombin generation rate for the different prothrombin activators Thrombin generation rate (nmol thrombin/mL/ Prothrombin Activation rate constant min/.mu.mol prothrombin/ activator ka (s.sup.-1 nM.sup.-1) .mu.mol activator) PtPA 6.72 .times. 10.sup.5 40.3 OsPA 7.33 .times. 10.sup.5 44.0 ecarin 4.61 .times. 10.sup.4 2.76 carinactivase-1 3.94 .times. 10.sup.3 0.24 carinactivase-2 3.18 .times. 10.sup.4 1.91 notecarin 7.14 .times. 10.sup.2 0.043
[0399] There was no reaction between the different prothrombin activators and the thrombin substrate S-2238, nor was there any reaction by prothrombin with S-2238. The activation rate data constants confirmed the group C prothrombin activators (PtPA and OsPA) are most potent in hydrolysing the prothrombin to thrombin. In fact they were more than 20 times more effective than ecarin and carinactivase-2, more than 200 times then carinactivase-1 and approximately 1000 times more effective than notecarin. The estimated thrombin generation rates equally show the same efficacy ratio, the group C activators were able to generate the largest amount of thrombin, followed by ecarin, carinactivase-2, carinactivase-1 and notecarin being the least efficient. As noted in Example 2d, notecarin requires cofactors for activity.
[0400] Materials and Methods as Used in Following Examples
[0401] Where indicated in the Examples that follow, the following Materials and Methods were used.
[0402] (A) Containers for Preparing Samples
[0403] A typical container (e.g. tube) for preparation of a serum sample or a plasma sample contains: (1) a gel barrier to separate cells (and the clot in the case of serum) from plasma or serum during centrifugation and to ensure no subsequent re-mixing; (2) a surfactant coating the inner wall to prevent cell and protein adhesion to the tube wall to minimise subsequent cell lysis; and (3) a procoagulant that enhances the dotting process (e.g. silica particles) for preparation of a serum sample or an anticoagulant (e.g. lithium heparin, citrate, or EDTA) for preparation of a plasma sample.
[0404] In the Examples, a number of commercially available tubes were used to prepare serum samples or plasma samples as described below.
[0405] Greiner Vacuette.TM. plasma tube. This is a plasma separator tube supplied by Greiner under reference number 456083. The inner wall of the tube is coated with spray-dried lithium heparin (89 IU) that acts as an anticoagulation agent. The tube also contains a separation gel in the base of the tube. Greiner states that this gel acts to form a stable barrier between the plasma and the blood cells allowing parameters to remain stable for up to 48 hours. The fill volume is 5.0 mL.
[0406] Becton Dickinson (BD) Vacutainer.TM. plasma tube (BD PST II). This is a plasma separator tube supplied by BD under reference number 367375. Each tube contains 77 IU/mL of lithium heparin (0.2-1.0 mg) as an anticoagulant. The tube contains an acrylic gel (0.8-1.9 g) that forms a barrier between the cells and the plasma during centrifugation. BD state that this gel provides enhanced plasma purity demonstrated by a reduction in fibrin and measurable white blood cells, red blood cells and platelets, and enhanced analyte stability as most analytes are stable in the tube for 24 hours at 25.degree. C. The fill volume is 4.0 mL.
[0407] Greiner Vacuette.TM. serum tube. This tube is supplied by Greiner under reference number 456078. The tube contains silica particles as the clot activator. The fill volume is 4.0 mL. The recommended clotting time in this tube is 30 minutes.
[0408] Becton Dickinson (BD) Vacutainer.TM. serum tube (BD SST II). This tube is supplied by BD under reference number 367954. This tube is used to obtain and separate a serum sample. Each tube contains 0.20-2.56 mg amorphous (or fused) silica as the coagulation agent, and also a gel (0.90-3.50 g) that forms a barrier between the clot and the serum after centrifugation. BD recommend a clotting time of 30 minutes for this tube. The fill volume is 4.0 mL.
[0409] Becton Dickinson (BD) Vacutainer.TM. Rapid Serum tube (BD RST). This tube is supplied by BD under reference number 368771. The tube contains a thrombin-based medical clotting agent and gel. BD claims that these tubes will provide a clotted sample within five minutes, (i.e. faster clotting than a standard commercially available serum tube) and will have "minimal fibrin formation". The fill volume is 4.0 mL.
[0410] Greiner Vacuette.TM. No Additive tube. These are plain plastic tubes supplied by Greiner under reference number 45400 to which prothrombin activators at particular concentrations were added as described in the Examples below.
[0411] Sarstedt serum tube. This tube is supplied under catalogue number 5092503 with a light brown top and has a fill volume of 4.7 mL. These tubes contain silica particles and a gel barrier.
[0412] Terumo plain tube. This tube is supplied under the trade mark VENOSAFE under catalogue number VF-076SP and has a fill volume of 6.0 mL. These tubes contain silica particles and a gel barrier.
[0413] Terumo Red Top (RT) tube. This tube is supplied under the trade mark VENOSAFE under catalogue number VF-108SAS and has a fill volume of 8.0 mL. These tubes are internally coated with silica and have a gel barrier.
[0414] Greiner Vacuette.TM. citrate tube. This plastic tube contains 3.2% citrate with a 4.0 mL fill volume. The tube is sold under catalogue number 454327.
[0415] Greiner Vacuette.TM. K2EDTA tube. This plastic tube contains 8 mM EDTA and has a 4.0 mL fill volume. The tube is sold under catalogue number 454023.
[0416] (B) Sample Analyses--Clotek Analyser
[0417] The Clotek analyser (Hyland, USA) uses a sample tube containing a magnetic ball which oscillates vertically when the tube is inserted into the machine. The magnetic steel ball is held in position by a magnetic field while the tube oscillates. The sample, buffer, .+-.calcium and procoagulant are added and the timer is triggered. Thrombin converts fibrinogen into clottable fibrin, and the time required to form the detectable clot is the clotting time. At clot formation the steel ball is removed from the magnetic suspension field and the light path, causing the light beam to strike the photocell and stop the timer (Austen, D. E., et al. 1975 and Starr, H., et al., 1980).
[0418] (C) Sample Analyses--Thromboelastography
[0419] Thromboelastography (TEG) measures clotting of whole blood by determining the rate of blood clot formation and lysis and the elastic properties of the blood clot during its formation and lysis. Where indicated in some Examples below, TEG analysis was performed.
[0420] The TEG analysis was performed using a Thrombelastograph.RTM. 5000 (TEG, Haemoscope Corporation Pty Ltd, Niles, Ill., USA) with TEG Analytical Software version 4.
[0421] The clotting parameters used here for the TEG analysis were as follows:
[0422] R--reaction time (seconds/minutes): time from start of the analysis until the TEG tracing amplitude reaches 2 mm, indicating detectable clot formation;
[0423] .alpha.--angle: measure of the rate of fibrin formation and cross-linking (i.e. clot strengthening);
[0424] MA--maximum amplitude (mm): maximum strength of the clot;
[0425] TMA--time (seconds/minutes) from start of the analysis until MA is reached; and
[0426] K--time (s): time from the start of the clot formation to the time the curve reaches amplitude of 20 mm.
[0427] An example of a TEG plot (clot formation part) with the above clotting parameters labelled is provided in FIG. 25. Only the clotting parameters are presented as the clot lysis parameters were not included.
Example 3: Clotting of Plasma Samples by Venom Prothrombin Activators
Example 3a: Clotting Times of Normal Citrated Plasma by PtPA Versus Bovine Thrombin
[0428] The clotting times of pooled citrated plasma (combined plasma from 6 normal participants) by PtPA (prepared as outlined in Example 1d) and bovine a-thrombin (3276 U/mg; Sigma Chemical Co.), with and without added calcium (10 mM), were measured in duplicate samples.
[0429] The results are shown in Table 14, where the values are the means of duplicate experiments.
TABLE-US-00015 TABLE 14 Clotting times of citrated plasma with PtPA and bovine thrombin, measured in a Hyland-Clotek machine (seconds). Clotting time (seconds) Concentration PtPA + PtPA - thrombin + thrombin - of enzyme (nM) calcium calcium calcium calcium 100 4.2 4.7 4.8 7.9 50 4.2 4.8 6.4 17.9 30 4.3 5.0 13.3 27.2 10 4.5 5.4 33.2 57.2 3 5.0 6.2 78.6 >200.sup.# 1 6.0 6.6 >200.sup.# >200 0.1 8.7 8.8 >200 >200 0.001 (1 pM) 13.5 29.0 >200 >200 0.0001 (0.1 pM) 74.1 98.6 >200 >200 0 >200 >200 >200 >200 #Weak clot observed.
[0430] The PtPA clotted citrated plasma efficiently at concentrations as low as 0.1 pM with or without recalcification. A concentration of 1 nM gave a clotting time of about 6 seconds, close to the minimum clotting time achieved at 100 nM of 4.2 seconds.
[0431] These results also show that PtPA is about 3.times.10.sup.4 times more efficient in the clotting of citrated plasma (with or without re-calcification) on a molar basis than bovine thrombin. For example, in the presence of 10 mM added calcium, 1 pM PtPA gave the same clotting time as 30 nM thrombin and 0.1 pM PtPA had a very similar clotting time to 3 nM thrombin (values in bold in Table 14).
Example 3b: Clotting Times of Normal Citrated Plasma by Prothrombin Activators
[0432] This experiment used the same plasma as in Example 3a.
[0433] Each Clotek tube contained 100 .mu.L of normal "pooled" citrated plasma, 100 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4), 50 .mu.L 0.2 M CaCl.sub.2 or saline, and 50 .mu.L of each procoagulant (either prothrombin activator preparation or thrombin).
[0434] The clotting results from each prothrombin activator preparation and thrombin with the normal "pooled" citrated plasma (with and without re-calcification) are shown in Tables 15 and 16. In these tables (and elsewhere), CA-1 is carinactivase-1 and CA-2 is carinactivase-2.
TABLE-US-00016 TABLE 15 Clotting times of normal "pooled" citrated plasma with different prothrombin activator preparations and thrombin, with calcium. With Ca.sup.2+ (seconds) [Procoagulant] (mM) Ecarin CA-1 CA-2 PtPA OsPA Notecarin Thrombin 100 43.0 108 65.8 3.5 5.7 42.4 4.3 50 56.0 145 85.0 4.6 6.5 52.8 6.7 30 69.0 181 167 4.9 7.9 57.5 10.5 10 105 >300 >300 12.5 12.5 68.5 32.8 3 208 >300 >300 20.1 18.3 99.8 92.3 1 >300 >300 >300 61 30.0 144 >300 0.1 >300 >300 >300 120 108 284 ND 0.01 >300 >300 >300 212 >300 >300 ND 0.001 >300 >300 >300 >300 >300 >300 ND
TABLE-US-00017 TABLE 16 Clotting times of normal "pooled" citrated plasma with different prothrombin activator preparations and thrombin, without calcium Procoagulant Without Ca.sup.2+ (seconds) (mM) Ecarin CA-1 CA-2 PtPA OsPA Notecarin Thrombin 100 94.0 >300 216 3.9 7.3 51.7 5.6 50 129 >300 >300 5.0 8.2 67.6 9.2 30 167 >300 >300 6.1 8.9 76.7 14.6 10 >300 >300 >300 9.0 14.0 115 43.6 3 >300 >300 >300 15.5 21.5 250 145 1 >300 >300 >300 24.6 34.5 >300 >300 0.1 >300 >300 >300 93.0 152 >300 ND 0.01 >300 >300 >300 >300 >300 >300 ND 0.001 >300 >300 >300 >300 >300 >300 ND
[0435] The results show that in the presence of added calcium all the prothrombin activators (representing groups A-D) efficiently clotted normal citrated plasma.
[0436] Citrated plasma is not completely calcium depleted. The total calcium concentration in normal citrated plasma is .about.1.3 mmol/L and the ionised calcium concentration is unmeasurable (<0.25 mmol/L). However, citrate is a chelator of calcium, binding to the calcium in the sample so that free calcium is not readily available to be used for the clot formation. Calcium is part of the prothrombinase complex and plays an important role in the prothrombin to thrombin catalysis. Citrate also binds other metal ions including magnesium and zinc ions. The amount of citrate present in the tube, 0.109 mol/L is more than sufficient to chelate all the metal ions in the tube.
[0437] These results showed that ecarin, the group A prothrombin activator, was more than twice as effective in clotting the plasma in the presence of added calcium, and therefore calcium has a significant enhancement on ecarin's ability to accelerate the clot formation. The group B prothrombin activators, carinactivase-1 and carinactivase-2, were strongly affected by the absence of calcium, with a decrease in activity of .about.4-fold, confirming the calcium enhances clotting activity with these prothrombin activators. The activity of notecarin, the group D prothrombin activator, was stimulated by the presence of added calcium, and the requirement for calcium was most evident with the decrease in the notecarin concentration with the addition of calcium increasing its effectiveness by .about.30%.
[0438] Thrombin itself was affected to a smaller degree by the absence of the added calcium. Thrombin itself generates thrombin formation in the sample via the coagulation cascade of prothrombin which requires calcium. Therefore the amount of thrombin formed from the sample prothrombin decreased with the absence of calcium.
[0439] The calcium dependence of the procoagulants was less when they were in high concentrations due to their potency. The results showed that the group C prothrombin activators, PtPA and OsPA, were the most efficient in clotting the normal "pooled" citrate plasma both in the presence or absence of added calcium. In fact, the absence of calcium made minimal difference on the clotting ability of these two procoagulants. With and without the presence of added calcium the group C prothrombin activators required .about.0.1 nM to achieve clotting in <5 minutes.
Example 3c: Clotting Times of Citrated Plasma Samples and Fibrinogen by PtPA
[0440] The ability of PtPA to clot the following recalcified citrated plasma samples and a sample of purified fibrinogen was determined in the following duplicate samples:
[0441] (i) citrated plasma;
[0442] (ii) citrated plasma adsorbed by Al(OH).sub.3 (adsorption by Al(OH).sub.3 removes prothrombin and other .gamma.-carboxyglutamate (GLA)-containing proteins);
[0443] (iii) citrated plasma from patients on long term warfarin therapy (with International Normalized Ratio (INR) greater than 4.0);
[0444] (iv) citrated plasma from patients on long term warfarin therapy (with INR greater than 4.0) adsorbed by Al(OH).sub.3,
[0445] (v) citrated Factor X-deficient plasma obtained from congenitally deficient patients;
[0446] (vi) citrated Factor V-deficient plasma obtained from congenitally deficient patients; and
[0447] (vii) purified human fibrinogen (2 mg/mL in isotonic saline), adsorbed with Al(OH).sub.3 (to remove traces of prothrombin).
[0448] With respect to (iii) and (iv), warfarin inhibits liver .gamma.-carboxylase, an enzyme which inserts .gamma.-carboxyl groups into 10 N-terminal glutamic acid residues of prothrombin. Warfarin plasma therefore contains an uncarboxylated precursor of prothrombin, descarboxyprothrombin, which is not removed by adsorption with Al(OH).sub.3.
[0449] With respect to (v) and (vi), Factor Xa and Factor Va are essential components of the human prothrombinase complex. The Factor X-deficient and Factor V-deficient plasmas were supplied by Ortho-Diagnostics, USA.
[0450] The results are shown in Table 17.
TABLE-US-00018 TABLE 17 Clotting times of citrated plasma samples (i)-(vi) and fibrinogen sample (vii). PtPA* Clotting time (seconds .+-. 0.5 seconds) for samples (nM) (i) (ii) (iii) (iv) (v) (vi) (vii) 54 12.8 >180 15.6 18.4 16.5 12.8 >180 26 16.4 >180 18.4 26.8 n.d. n.d. n.d. 16 22.5 >180 24.6 28.6 n.d. n.d. n.d. 7.3 34.6 >180 38.8 42.3 n.d. n.d. n.d. 3.8 48.2 >180 55.0 59.8 n.d. n.d. n.d. 0.sup.# 10.0 10.6 7.7 8.6 14.6 10.6 5.4 *In this experiment, a different preparation of PtPA was used, accounting for the lower clotting activity than in Table 14 above. n.d. means that the measurements were not determined. .sup.#Where no PtPA was added, 0.1 Units of bovine thrombin (Parke-Davis, now Pfizer, USA) was added to all samples as a positive control. The rapid clotting observed on addition of thrombin confirmed the presence of fibrinogen in all samples.
[0451] The results in Table 17 show that:
[0452] (1) prothrombin is necessary for clotting by PtPA since Al(OH).sub.3-adsorbed citrated plasma (sample (ii)) was not clotted by PtPA;
[0453] (2) PtPA clots warfarin plasma (samples (iii) and (iv)) indicating that descarboxyprothrombin can be converted to a-thrombin by PtPA;
[0454] (3) the result for warfarin plasma adsorbed by Al(OH).sub.3 (sample (iv)) confirms that descarboxyprothrombin is converted to a-thrombin, since traces of normal prothrombin in warfarin plasma would be removed by Al(OH).sub.3;
[0455] (4) Factor Xa is not required for clotting by PtPA (results for sample (v));
[0456] (5) Factor Va is not required for clotting by PtPA (results for sample (vi)); and
[0457] (6) PtPA is unable to clot a fibrinogen solution and is therefore not functioning as a thrombin-like enzyme.
[0458] In summary, these results shows that clotting induced by PtPA is due to prothrombin activation; that descarboxyprothrombin is converted efficiently to thrombin by PtPA; that Factors Xa and Va are not required for prothrombin activation by PtPA; and PtPA does not convert fibrinogen to fibrin under these conditions.
Example 3d: Clotting of EDTA-Treated Plasma
[0459] A small but significant percentage of samples (serum or lithium heparin plasma samples) received for biochemical analysis is contaminated by EDTA. It is therefore important to determine if the prothrombin activators were effective in clotting EDTA plasma and EDTA whole blood. This example is therefore provided as Example 5b below, so that a comparison of the studies of EDTA plasma and EDTA whole blood could be made.
Example 3e: Clotting of Citrated Plasma from Warfarin Medicated Participants by Prothrombin Activators
[0460] Warfarin is a commonly used anticoagulant, and therefore the purpose of this experiment was to determine if warfarin therapy affects the ability of the prothrombin activators: ecarin; carinactivase-1; carinactivase-2; PtPA; OsPA; and notecarin, to clot samples from patients medicated with warfarin.
[0461] Participant plasma samples with different clotting times (related to warfarin dosage) based on the INR (1.1-7.6) were selected and tested with the different procoagulants (prothrombin activators and thrombin).
[0462] The Clotek tubes contained 100 .mu.L of lithium heparin plasma, 100 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4), 50 .mu.L of 0.2 M CaCl.sub.2, and 50 .mu.L of each procoagulant.
[0463] The results are shown in Tables 18 and 19. As used in these tables and elsewhere, CA-1 is carinactivase-1 and CA-2 is carinactivase-2.
TABLE-US-00019 TABLE 18 Clotting times (seconds) with re-calcified citrated plasma from warfarin medicated patients, by prothrombin activators. Sample INR [Prothrombin activator] 30 nM 1.1 1.2 1.5 2.0 2.9 6.4 7.6 PtPA 5.0 5.7 6.7 6.1 7.0 9.8 10.5 OsPA 9.0 9.1 12.6 16.8 17.2 17.2 19.3 Notecarin 69.4 67.1 68.6 66.6 71.0 91.0 121 Ecarin 72.2 63.4 63.4 69.4 85.6 114 111 CA-1 231 198 232 229 >300 >300 >300 CA-2 115 103 115 109 149.2 242 240
TABLE-US-00020 TABLE 19 Clotting times (seconds) with re-calcified citrated plasma from warfarin medicated patients by PtPA and thrombin. Sample INR [Procoagulant] 30 nM 1.1 1.7 2.3 2.8 5.8 6.4 PtPA 7.4 8.1 8.1 8.3 11.1 13.3 Thrombin 13.4 13.7 14.7 14.0 14.7 16.3
[0464] Warfarin leads to a decrease in the synthesis of normal Factor-X (FX) and prothrombin from their active precursors (descarboxyfactor-X and descarboxyprothrombin). Prothrombin and descarboxyprothrombin are the substrates targeted by the prothrombin activators. The decrease in prothrombin means less thrombin was generated by the prothrombin activators, hence the prolonged clotting times observed. Thrombin itself was not inhibited by warfarin thus the clotting times with thrombin are not affected as has been observed in this experiment. The results in this experiment showed that the carinactivase-1 was the only prothrombin activator unable to clot plasma samples with INR.gtoreq.2.9. The results showed PtPA and OsPA were the most effective procoagulants for plasma from patients taking warfarin.
Example 3f: Clotting of FV-Deficient and FX-Deficient Plasma by Prothrombin Activators
[0465] The prothrombin activator preparations were tested for their ability to clot commercially available FV-deficient plasma (#0008466150) and FX-deficient plasma (#0008466350) (Instrumentation Laboratory, Lexington, Mass., USA) using the Clotek analyser (Hyland USA).
[0466] The Clotek tube contained 100 .mu.L of FV- or FX-deficient plasma, 100 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4), 50 .mu.L, 0.2 M CaCl.sub.2 and 50 .mu.L of each procoagulant. All procoagulants were at a concentration of 30 nM, and an additional sample containing carinactivase-1 at a concentration of 100 nM was also tested.
[0467] The results are shown in Table 20.
TABLE-US-00021 TABLE 20 Clotting times (seconds) of FV- or FX-deficient plasma by the prothrombin activator preparations and thrombin Clotting time (seconds) Normal "pooled" FV-deficient FX-deficient [Procoagulant] 30 nM citrated plasma plasma plasma Ecarin 66.8 74.8 77.5 CA-1 184 >300 >300 CA-1 (100 nM) ND 216 218 CA-2 133 159 159 OsPA 13.3 12.7 12.7 PtPA 9.0 7.4 7.6 Notecarin 47.5 >300 71.9 Thrombin 13.8 44.6 50.2 "ND" means this was not determined.
[0468] Notecarin was not able to clot the FV-deficient plasma in 5 minutes, and its clotting efficiency was also diminished in the absence of FX. Furthermore, the clotting activity of the group A and group B prothrombin activators was also diminished in the absence of FV or FX. The group C prothrombin activators, PtPA and OsPA, were clearly the most efficient in clotting the plasma, and have no requirement for FV or FX from plasma.
[0469] Thrombin clotting time was significantly increased to be more than 3 times longer in the FV- and FX-deficient plasma compared to the clotting time for the normal "pooled" citrated plasma.
Example 3g: Estimation of Amount of Thrombin Generated by PtPA and OsPA in Normal and Heparinised Citrated Plasma
[0470] This experiment was designed to provide an estimate of the amount of thrombin that is generated by PtPA and OsPA using different concentrations and in the presence of heparin.
[0471] Normal citrated plasma was obtained from a healthy participant into a number of tubes (Greiner citrate) which were immediately centrifuged and the citrated plasma pooled.
[0472] The thrombin used was Thrombin-JMI, Topical (Bovine) 5000USP (GenTrac Inc, King Pharmacia, Middleton, Wis., USA). The USP units are equivalent to IU, and one IU is equal to 9 nM. The thrombin substrate used was S-2238 (25 mg stock solution MW 626 Da was dissolved in 15 mL to give 2.67 mM (#820324 Chromogeinix IL Lexington Mass., USA). The heparin used was 5000 IU/5 mL (1000 IU/mL), sodium heparin, Pfizer, NZ. For heparin, 190.9 IU=1 mg (Camenzind, et al. 1997); a MWt of 12000 Da was used to calculate molar concentrations. The buffer used was: 20 mM Hepes buffer; 150 mM NaCl; pH 7.4; 0.05% surfactant p20.
[0473] For the plasma incubation the tubes contained 0.5 mL citrated plasma, 50 .mu.L of 0.5 M CaCl.sub.2, and 5-40 .mu.L thrombin or a prothrombin activator plus 20 .mu.L of heparin to provide concentrations of 860, 4300, and 8600 nM (2, 10 and 20 IU) or saline. The tubes were incubated for 5 minutes at room temperature. Immediately after this 5 minute period, the clot was ringed using two wood applicator sticks, removed and discarded.
[0474] For thrombin activity assays, each spectrophotometric cuvette contained 930 .mu.L of buffer, 50 .mu.L of S-2238 and 20 .mu.L of the clear serum that was remaining after the clot was removed from each tube in the incubation above. The absorbance change was monitored at 405 nm for 5 minutes.
[0475] FIG. 26 shows the activities of remaining thrombin in those tubes to which different concentrations of thrombin had been added. For each curve, the slope at 275 seconds was determined using the time range of 250-300 seconds, and these slopes along with the thrombin concentrations were then used to develop the standard curve in logarithmic format to provide a linear relationship over higher thrombin concentration, as shown in FIGS. 26 and 27.
[0476] The re-calcified plasma (no procoagulant) took .about.37 minutes to clot at room temperature. All the thrombin containing samples exhibited latent clotting. The lowest concentration of thrombin was the slowest to exhibit latent clotting (.about.3 minutes). The time to latent clot decreased as the concentration of thrombin increased. The samples with concentrations >200 nM thrombin exhibited latent clotting immediately after the clot was removed as the serum was being aliquoted. At times, a second clot removal was necessary to ensure sufficient sample was obtained from the tube for the photometric analysis.
[0477] For the samples containing different concentrations of the prothrombin activators PtPA and OsPA, but without heparin, the results were as follows. The spectrophotometric results for the samples containing different concentrations of PtPA are shown in FIG. 28, and for the samples containing different concentrations of OsPA are shown in FIG. 29.
[0478] The thrombin concentrations generated by PtPA was then plotted against the slope at 5 minutes, and the results are shown in FIG. 30, and the thrombin concentrations generated by OsPA was then plotted against the slope at 5 minutes with the results are shown in FIG. 31.
[0479] The concentration of thrombin generated by the different PtPA and OsPA concentrations remaining active in the serum was estimated and is shown in Table 21.
TABLE-US-00022 TABLE 21 Estimation of the active thrombin present in the serum generated by different concentration of PtPA and OsPA determined from the standard curve regression equation, FIG. 27. [Prothrombin activator] Slope at 5 minutes Estimated concentration (nM) per second of active thrombin (nM) PtPA 1.2 0.00098 574 6.1 0.00171 869 12.3 0.00196 965 OsPA 1.2 0.00094 554 6.1 0.00149 784 12.3 0.00195 961
[0480] The results indicated that very significant amounts of thrombin were generated even by 1.2 nM PtPA and OsPA. Very small amounts of PtPA and OsPA were capable of completely clotting the re-calcified normal citrated plasma in <5 minutes. No latent clotting was observed, indicating that the amount of thrombin generated was sufficient to completely catalyse the fibrinogen and prevent latent clotting which would equate to very high quality serum.
[0481] For the samples containing different concentrations of heparin, the results were as follows.
[0482] First, the concentration of thrombin remaining in the serum component from the activity of thrombin, PtPA and OsPA in the presence of 4300 nM (10 IU) heparin was determined and is shown in FIG. 32 (change in A405 for S-2238 hydrolysis/minute on the y-axis is proportional to thrombin concentration). The 135 nM thrombin failed to fully clot the plasma sample, with only a minor 2-3 mm "fluff-like" clot observed. The 6.1 nM concentrations of both prothrombin activators clotted the sample containing 4300 nM (10 IU) of heparin in approximately one minute and no latent clotting was observed even after 24 hours storage at room temperature.
[0483] It was then investigated whether the reaction was of similar nature with different heparin concentrations. Normal pooled citrated plasma was was obtained from the Coagulation Laboratory, Pathology Queensland, Princess Alexandra Hospital as there was insufficient plasma from the normal participant. The results are shown in FIG. 33 (PtPA) and FIG. 34 (OsPA).
[0484] The amount of thrombin generated by 1.5 nM PtPA and OsPA concentrations remaining active in the serum was estimated and is shown in Table 22.
TABLE-US-00023 TABLE 22 Estimation of the active thrombin present in the serum generated by 1.5 nM of PtPA and OsPA determined from the standard curve regression equation, FIG. 27. [Prothrombin Estimated activator] Heparin nM Slope at 5 minutes concentration of (nM) (IU)/mL per second active thrombin (nM) PtPA 1.5 0 0.0007 443 1.5 860 (2) 0.00018 159 1.5 4300 (10) 0.00024 197 1.5 8600 (20) 0.00024 197 OsPA 1.5 0 0.0006 394 1.5 860 (2) 0.0002 172 1.5 4300 (10) 0.00022 185 1.5 8600 (20) 0.00020 172
[0485] The samples containing 860 and 4300 nm (2 and 10 IU) heparin clotted within two minutes and showed no latent clotting. The clot was solid but not as solid as those observed in the samples that did not contain heparin. The 860 nM (2 IU) heparin-containing sample gave a thrombin activity that was about 25% of that of the plasma samples not containing heparin. In contrast, the sample containing 8600 nM (20 IU) heparin produced a "jelly-like" clot (more similar to the clots formed in the thrombin-containing samples) within 3 minutes the sample then exhibited latent clotting within 5 minutes. In FIGS. 33 and 34, the reason for the shape of the progress curves at higher heparin concentrations is unknown.
Example 3h: Estimation of Amount of Thrombin Generated by BD RST Tube using Normal Pooled Citrated Plasma, and the Effect of Heparin
[0486] The purpose of this experiment was to provide an estimate of the amount of thrombin generated and present in the active form in the BD RST tube filled with different volumes of plasma. Additionally the effect of heparin on the clotting ability of the BD RST tubes was also studied.
[0487] Normal pooled citrated plasma (<24 hours old) was obtained from the Coagulation Laboratory, Pathology Queensland, Princess Alexandra Hospital.
[0488] For the standard curve, the BD RST tubes contained 1, 2, 3 and 4 mL of citrated plasma and 50 .mu.L of 0.5 M CaCl.sub.2. The tubes were inverted 8-10 times to mix and incubated for 5 minutes. The clot was ringed using two wood applicator sticks, removed and discarded.
[0489] The spectrophotometric cuvette contained 930 .mu.L of buffer, 50 .mu.L of S-2238 and 20 .mu.L of the clear serum that was remaining after the clot was removed from each BD RST tube above. The absorbance change was monitored at 405 nM for 5 minutes, and the results are shown in FIG. 35. Thrombin concentrations were estimated from the slopes using the standard curve in FIG. 16, and plotted against the slopes from FIG. 35. The original BD RST tube thrombin concentrations and remaining thrombin concentrations in the clot free sera from the four samples were are as follows; 135 versus 4.3; 67.5 versus 1.4; 45 versus 0.5; and 33.8 versus 0.3 nM respectively. It is likely that most of the thrombin was removed during the clot removal as it would be bound to the clot fibrin.
[0490] Unlike the samples with pure thrombin described in Example 3g, each sample in each of the BD RST tubes clotted within one minute and no latent clotting was observed. However, the clot strength was much weaker that those observed with PtPA or OsPA described in Example 3g.
[0491] For the samples containing different volumes of citrated plasma and different heparin concentrations, the absorance change of each sample was monitored at 405 nM for 5 minutes, and the results are shown in FIG. 37. The reason for the shape of the progress curves at high heparin concentrations is unknown.
[0492] There were no samples that produced complete clotting in the presence of any of the three heparin concentrations. The strongest clot was observed with the samples containing 830 nM (2 IU) heparin and one mL of plasma, although the clot was loose by comparison to the solid clots in the absence of heparin. The weakest clots that formed were those in the tube filled with 4 mL of plasma and containing 8600 nM (20 IU) heparin. The clot was "fairy-floss" like and only in part of the sample. All tubes containing heparin showed some latent clotting after 5 minutes. The actual curve shape with the 4300 and 8600 nM (10 and 20 IU) heparin was very similar to the curves observed with PtPA and OsPA.
[0493] The slopes and the estimation of active thrombin in the presence of the different heparin concentrations and the different thrombin concentrations are shown in Table 23.
TABLE-US-00024 TABLE 23 Estimation of the active thrombin in the presence of different concentrations of heparin and different concentrations of thrombin from BD RST tubes, using the regression equation from the standard curve in FIG. 36. [BD RST Estimated Ratio of decrease thrombin] [active BDRST in presence of [Heparin] per mL thrombin] heparin (column (IU) [nM] (nM) Slope (nM) 2/column 4) 2 [830 nM] 135 0.00050 30 4.5 2 [830 nM] 33.8 0.00016 6 5.6 10 [2150 nM] 135 0.00050 30 4.5 10 [2150 nM] 33.8 0.00024 12 3.0 20 [8300 nM] 135 0.00042 25 5.4 20 [8300 nM] 33.8 0.00020 9 3.8
[0494] The results showed that the same nominal thrombin concentrations (in column 2) produced similar active thrombin concentrations in the presence of different heparin concentrations. Thrombin concentrations measured after clotting were lower by several fold than the nominal concentrations.
Example 4: Clotting of Whole Blood Samples from Healthy Participants by Prothrombin Activators
Example 4a: Clotting by Ecarin
[0495] For this experiment, ecarin, purified from Echis carinatus venom, was purchased from Sigma (catalogue number EO 504-1VL; batch number 128K 1536). Citrated blood (pool of three samples with normal coagulation profiles) was obtained from Pathology Queensland, Princess Alexandra Hospital, Queensland, Australia.
[0496] One vial of ecarin (50 units) was reconstituted in 100 .mu.L of H.sub.2O resulting in a stock concentration of 500 units/mL. Dilutions of this stock in distilled water (1:100, i.e. 5 units/mL; 1:1000, i.e. 0.5 units/mL; 1:10000, i.e. 0.05 units/mL) were used in TEG assays. Each assay mixture consisted of 310 .mu.L citrated blood, 20 .mu.L 0.2 M CaCl.sub.2, and 10 .mu.L saline (control) or ecarin dilution. Therefore, the concentration range of ecarin in the assay mixture was 0 to 0.15 units/mL. All experiments were done in duplicate. The results are shown in Table 24.
TABLE-US-00025 TABLE 24 TEG results for clotting of re-calcified citrated blood samples by ecarin. Ecarin (units/mL) R (min) TMA (min) .alpha. (deg) MA (mm) control 6.95 26.45 52.65 65.60 0.0015 7.3 27.60 59.80 65.65 0.015 4.5 17.90 74.15 67.25 0.15 2.1 15.75 69.25 73.80
[0497] Addition of ecarin to recalcified citrated blood resulted in a concentration dependent decrease in both R-time and TMA, and a small increase in MA. At the highest concentration tested, 0.15 units/mL, ecarin reduced the R-time from 6.95 to 2.1 minutes and the TMA from 26.45 to 15.75 minutes. The lowest concentration of ecarin tested, 0.0015 units/mL, had no significant effect on clotting time and clot strength whereas ecarin at a concentration of 0.015 units/mL had an intermediate effect.
[0498] These results suggest that ecarin would also be an effective prothrombin activator to rapidly produce serum for analyte measurements in a clinical laboratory.
Example 4b: Clotting of Normal "pooled" Re-Calcified Citrated Blood by Six Prothrombin Activators
[0499] Purified preparations of ecarin, carinactivase-1, carinactivase-2, PtPA, OsPA, and notecarin prepared as described in Example 1 were used. Clotting of normal pooled citrated blood (as in Example 4a) by the six prothrombin activators, by thrombin, and in BD RST tubes was studied by TEG, as described in Example 4a.
[0500] The results are provided in Table 25.
TABLE-US-00026 TABLE 25 TEG clotting study with normal "pooled" re-calcified citrate blood with the different procoagulants. Procoagulant (nM) R (min) MA (mm) TMA (min) Ecarin 32 1.2 60.5 18.2 9 2.2 61.6 20.1 3.2 3.4 59.0 21.6 0.9 6.5 59.5 29.7 Carinactivase-1 50 3.3 69.1 21.7 32 4.6 68.5 27.1 9 6.2 63.9 31.4 3.2 7.4 64.9 30.3 Carinactivase-2 50 1.8 64.9 19.9 32 2.1 68.2 20.0 9 3.3 67.8 19.7 3.2 4.6 68.0 23.6 OsPA 0.9 0.7 63.7 18.4 0.09 2.9 61.8 20.5 0.009 4.2 63.5 22.5 0.0009 6.2 59.3 24.8 PtPA 0.9 1.5 62.4 22.1 0.09 2.6 63.1 23.6 0.009 3.9 63.9 24.8 0.0009 5.8 53.4 28.1 Notecarin 3.2 1.2 62.1 20.2 0.9 1.8 63.7 18.3 0.09 3.7 63.2 20.3 0.009 7.3 59.0 26.3 Thrombin 32 2.3 65.2 22.8 9 3.5 64.0 23.8 3.2 5.5 62.4 27.6 BD RST 24 0.8 54.8 24.8 6 1.5 59.3 23.7
[0501] These results provided further support to the plasma clotting findings (Example 3) that all prothrombin activators clot blood very efficiently; and that the group C prothrombin activators (PtPA and OsPA) were the most effective in clotting the normal "pooled" re-calcified citrated whole blood. For example, 9 pM PtPA and OsPA achieved R times of about 5 minutes, and the maximum clot strength as confirmed by the MA results. The next most effective prothrombin activator was notecarin, the group D prothrombin activator, which was able to produce an R time of 5 minutes and maximum clot strength with a minimum concentration between 0.09 and 0.009 nM. Although notecarin itself does not contain FVa, the FV present in plasma is converted to active form by the small amount of thrombin formed by the notecarin FXa. The next most effective prothrombin activator was ecarin, the group A prothrombin activator, which was able to achieve the desired R time and maximum clot strength with a minimum concentration between 0.9 and 3.2 nM. The least effective prothrombin activators were the group B prothrombin activators, carinactivase-1 and carinactivase-2 which required between .gtoreq.9 and .gtoreq.3 nM concentrations respectively to achieve R times of <5 minutes and maximum clot strength. The required minimum thrombin concentration to achieve the desired R time and maximum clot strength was .gtoreq.3 nM.
Example 4c: Comparison of Commercially Available Serum Tubes and PtPA-Containing Tubes
[0502] Blood from a healthy participant was drawn sequentially into the following tubes using an inline butterfly needle:
[0503] (1) a 4.0 mL Greiner Vacuette.TM. No Additive tube;
[0504] (3) Greiner Vacuette.TM. serum tube;
[0505] (4) BD Vacutainer.TM. serum tube;
[0506] (5) Sarstedt serum tube;
[0507] (6) Terumo RT tube; and
[0508] (7) Terumo plain tube.
[0509] The tubes were filled with freshly collected blood to the required fill mark and mixed immediately for 30 seconds by gentle inversion 8-10 times. Then a 340 .mu.L sample was transferred into a TEG cup for immediate analysis. This method of blood collection allowed for rapid transfer of a blood sample for TEG analysis immediately after mixing in the commercially available tube so that no anticoagulant or re-calcification in TEG analysis was required. An aliquot, 330 .mu.L of the blood from the plain tube (1) was immediately transferred to a TEG cup and 10 .mu.L (1.41 .mu.g) of PtPA was added (2), and analysis was immediately started.
[0510] All TEG assays were done in duplicate. The TEG parameters (average of the duplicate samples) are shown in Table 26 below.
TABLE-US-00027 TABLE 26 TEG results for blood from a healthy participant in different tubes Sample R (min) TMA (min) .alpha. (deg) MA (mm) (1) Plain tube 14.3 46.5 26.9 49.8 (2) PtPA 0.8 12.2 70.8 54.9 (3) Greiner 3.0 23.5 71.2 68.2 (4) BD serum 4.9 25.3 71.0 68.1 (5) Sarstedt 5.1 23.8 70.8 69.1 (6) Terumo RT 9.6 22.5 63.2 58.1 (7) Terumo 13.4 31.2 38.2 54.6
[0511] The TEG parameters of Table 26 are shown in the TEG plot in FIG. 38 where: trace A represents the plain plastic tube (1); trace B represents the plain tube with added PtPA (2); trace C represents the Greiner serum tube (3); trace D represents the BD serum tube (4); trace E represents the Sarstedt serum tube (5); trace F represents the Terumo RT tube (6); and trace G represents Terumo serum tube (7).
[0512] The blood sample in the PtPA tube (2) clotted most rapidly (R time=0.8 minutes) in the TEG cup. R times for the commercial tubes (3)-(7) varied from 3.0 minutes for the Greiner tube (3) to 13.4 minutes for the Terumo tube (7). These R times can be compared with the R time of 14.3 minutes for the plain tube (1). It is believed that the differences in R times between the commercial tubes reflect the presence of different procoagulants and additives, and/or different amounts of procoagulants or additives in these tubes.
[0513] Clot formation in all tubes was also visually observed. The commercially available tubes were allowed to clot for the standard time of 30 minutes. The PtPA specimen was allowed to stand for 5 minutes. The tubes were visually inspected for clot formation prior to loading in the centrifuge. All tubes were centrifuged at 3000 g for 10 minutes at 20.degree. C. The specimens were stored at room temperature, .about.21.degree. C. for up to 6 hours and inspected on hourly intervals for latent clot formation. All tubes including the PtPA-containing tube formed solid clots, with the PtPA-containing tube forming a solid immobile clot within 2 minutes. No latent clotting was observed in any of the tubes up to 6 hours after collection.
[0514] This example shows that inclusion of a small amount of PtPA led to faster clotting than in any of the commercial tubes tested, demonstrating the potential for rapid production of serum to provide a faster turn-around time for analyte measurements in a clinical laboratory.
Example 4d: Clotting of Normal Citrated Blood by PtPA, OsPA, Ecarin, and Some Commercial Tubes
[0515] The purpose of this experiment was to investigate how three prothrombin activators: PtPA, OsPA, and ecarin compare to three routinely used commercial clotting tubes: Greiner serum, BD SST II and BD RST.
[0516] Blood from a healthy participant was collected into a citrate-containing tube.
[0517] For the prothrombin activators, 30 .mu.L of activator (concentration of PtPA and OsPA 0.56 nM; ecarin 5.3 nM in TEG cup), 20 .mu.L of 0.2 M CaCl.sub.2 and 290 .mu.L of citrated blood was added to the TEG cup.
[0518] For the commercial serum tubes, 1 mL of distilled water was added and allowed to mix on a roller for .about.2 hours to dissolve the content of the tube. The BD RST contains 135 nM bovine thrombin (33.8 nM/mL of blood), whereas the concentration used in this experiment in the TEG cup was 53 nM/mL. For the commercial serum tubes 45 .mu.L of the dissolved content, 20 .mu.L of 0.2 M CaCl.sub.2 and 275 .mu.L of citrated blood was added to the TEG cup.
[0519] The results are shown in Table 27.
TABLE-US-00028 TABLE 27 TEG data with PtPA, OsPA, ecarin, and routinely used commercial serum tubes with native blood from a healthy participant. Tube R time Tube Catalogue # (min) MA TMA Greiner Vacuette Serum 455078 2.8 62.5 20.6 BD SST II 367958 4.3 62.8 23.5 BD RST (18 nM) 368771 0.5 58.8 20.2 PtPA (0.53 nM) 1.4 65.6 19.8 OsPA (0.53 nM) 1.5 64.0 19.7 Ecarin (5.3 nM) 2.6 64.3 19.3 Native blood (plain tube) 6.4 67.6 27.8
[0520] The results demonstrated that the routinely used commercial serum tubes vary in the time at which significant clot formation was detected by the TEG (R time), with the time ranging from 0.5-6.4 minutes for re-calcified citrated blood from a healthy participant. The manufacturers of these tubes recommend a minimum clotting time of 30 minutes prior to centrifugation of the serum tubes. The results also demonstrated that PtPA and OsPA at 0.5 nM gave an R time of <2 minutes with an equivalent clot strength which make them very suitable for evaluation as procoagulant to be used in producing serum rapidly.
Example 5: Clotting of Whole Blood Samples Containing Anticoagulants by Prothrombin Activators
[0521] Many blood samples are taken from patients being treated with anticoagulants such as heparin, warfarin and citrate. In addition, some blood samples are collected into tubes containing anticoagulants such as EDTA and citrate; or are contaminated with anticoagulants during the collection process. The following experiments were performed to study the clotting of blood samples containing anticoagulants by prothrombin activators.
Example 5a: Concentration Dependent Clotting of Citrated or EDTA Treated Blood from a Healthy Participant
[0522] TEG analysis was performed where each TEG assay mixture consisted of 310 .mu.L of citrated or EDTA treated blood, 20 .mu.L of CaCl.sub.2 (0.2 M) and 10 .mu.L of saline or prothrombin activator solution at a range of concentrations (stock solutions--PtPA, 4.8 mg/mL or OsPA, 2.0 mg/mL). The results are shown in FIG. 39 and in Tables 28 and 29 or PtPA and Tables 30 and 31 for OsPA.
TABLE-US-00029 TABLE 28 TEG results for clotting of normal citrated blood with PtPA PtPA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 10.9 40.4 23.8 55.2 0.00141 10.3 39.6 26.2 44.0 0.0141 4.6 22.8 54.0 67.2 0.141 3.8 23.5 67.7 67.0 1.41 1.0 21.2 72.4 64.8 14.1 0.3 20.2 74.7 59.7
TABLE-US-00030 TABLE 29 TEG results for clotting of re-calcified EDTA-treated blood with PtPA PtPA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 9.8 39.2 25.8 54.6 0.00141 9.1 31.5 34.9 60.9 0.0141 6.8 27.0 62.4 64.4 0.141 5.5 25.3 62.0 64.4 1.41 1.3 19.8 72.1 66.3 14.1 0.3 21.2 70.1 60.9
TABLE-US-00031 TABLE 30 TEG results for clotting of normal citrated blood with OsPA OsPA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 9.0 31.3 29.5 55.0 0.000588 9.0 24.0 34.2 49.0 0.00588 5.9 21.3 54.3 62.0 0.0588 2.8 15.3 65.9 54.8 0.588 0.8 17.9 73.6 64.2 5.88 0.3 19.8 72.3 58.2
TABLE-US-00032 TABLE 31 TEG results for clotting of re-calcified EDTA-treated blood with OsPA OsPA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 13.9 40.8 31.2 50.7 0.000588 16.3 45.4 29.8 53.7 0.00588 9.8 30.0 36.2 62.4 0.0588 3.3 18.3 64.3 70.7 0.588 0.8 17.0 73.1 64.0 5.88 0.3 20.8 77.0 61.1
[0523] The R time of re-calcified blood without addition of a prothrombin activator was approximately 10 minutes which decreased to 1 minute or 1.3 minutes in the presence of 1.41 .mu.g/mL PtPA (Tables 28 and 29). The rate of clotting was rapid (angle=54-72.degree.) (between 0.0141 and 1.41) and full strength clots with MA values of 60-70 mm were obtained. The R time of re-calcified blood without addition of a prothrombin activator of approximately 10 minutes was decreased to 0.8 minutes in the presence of 0.59 .mu.g/mL OsPA (Tables 30 and 31).
[0524] It is believed that the very short R-times and large .alpha. observed on addition of the PtPA or OsPA may be explained by generation of a massive burst of thrombin, while downstream reactions (conversion of fibrinogen to fibrin, activation of factor XIII, cross-linking of fibrin monomers) are rate limiting and result in a comparatively smaller improvement of TMA.
[0525] The TEG results from Tables 28, 28, 30 and 31 are shown in FIG. 39, where TEG plot A shows the Table 28 results (citrated blood and PtPA); TEG plot B shows the Table 29 results (EDTA treated blood and PtPA); TEG plot C shows the Table 30 results (citrated blood and OsPA) and TEG plot D shows the Table 31 results (EDTA treated blood and OsPA).
[0526] In FIG. 39, the traces labelled A in each TEG plot represent 14.1 .mu.g/mL for PtPA and 5.88 .mu.g/mL for OsPA ; the traces labelled B represent 1.41 .mu.g/mL for PtPA and 0.588 .mu.g/mL for OsPA; the traces labelled C represent 0.141 .mu.g/mL for PtPA and 0.0588 .mu.g/mL for OsPA; the traces labelled D represent 0.0141 .mu.g/mL for PtPA and 0.00588 .mu.g/mL for OsPA; the traces labelled E represent 0.00141 .mu.g/mL for PtPA and 0.000588 .mu.g/mL for OsPA; and the traces labelled F represent 0 .mu.g/mL PtPA or 0 .mu.g/mL OsPA.
[0527] In summary, PtPA and OsPA both clot re-calcified citrated and EDTA-treated blood in a highly efficient concentration-dependent manner. Thus in practice, blood collected in an EDTA tube could be rapidly clotted to produce serum by adding PtPA or OsPA for biochemical and other laboratory analysis.
Example 5b: Clotting of EDTA Plasma and Blood Obtained from a Healthy Participant
[0528] As mentioned in Example 3d above, a small but significant percentage of samples (serum or lithium heparin plasma samples) received for biochemical analysis is contaminated by EDTA. It is therefore important to determine if prothrombin activators are effective in clotting EDTA plasma and EDTA whole blood. The purpose of this experiment was to do so, and to make a comparison of EDTA plasma and EDTA whole blood.
[0529] Blood from a healthy patient was collected into a Greiner EDTA tube containing 1.8 mg/mL (6.2 mmol) EDTA when filled to the fill mark. At this concentration, the EDTA is able to bind all the metal ions (including Ca.sup.2+, Mg.sup.2+, Fe.sup.2+, Zn.sup.2+, and Cu.sup.2+) present in the tube. To obtain EDTA plasma, a portion of each sample was centrifuged under standard protocols.
[0530] The Clotek tube contained 100 .mu.L of EDTA plasma, 100 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4), 50 .mu.L of 0.2 M CaCl.sub.2 or Tris buffer, and 50 .mu.L of each procoagulant.
[0531] The TEG cup contained 20 .mu.L of 0.2 M CaCl.sub.2 or saline, 60 .mu.L of procoagulant and 260 .mu.L of the EDTA blood.
[0532] For the BD RST experiment the contents of two tubes were dissolved with 1 and 4 mL of distilled water respectively, mixed for 5 minutes and 60 .mu.L of the content used in the TEG cup as the procoagulant.
[0533] The results of the plasma clotting study are shown in Table 32 and in Table 33.
TABLE-US-00033 TABLE 32 Clotting times for EDTA plasma obtained from a healthy participant with different procoagulants (prothrombin activators and thrombin) at different concentrations, with calcium. [Procoagulant] With Ca.sup.2+ (seconds) (nM) Ecarin CA-1 CA-2 PtPA OsPA Notecarin Thrombin 50 68.9 165.4 82.5 6.5 10.0 85.5 12.7 10 127 232 101 13.5 15.8 117 43.0 1 224 >300 151 35.7 42.7 206 >300 0.1 ND ND ND 98.6 123 ND >300 "ND" means this was not determined.
TABLE-US-00034 TABLE 33 Clotting times for EDTA plasma obtained from a healthy participant with different procoagulants (prothrombin activators and thrombin) at different concentrations, without calcium. Without Ca.sup.2+ (seconds) Procoagulant (nM) Ecarin CA-1 CA-2 PtPA OsPA Notecarin Thrombin 50 >300 >300 >300 31.7 56.4 132 17.9 10 >300 48.7 68.2 242 53.2 1 >300 86.4 107 >300 >300 0.1 190 251 >300
[0534] The results of the BD RST plasma clotting study are shown in Table 34.
TABLE-US-00035 TABLE 34 Clotting times for the BD RST tubes, with and without calcium. BDRST Tube (nM) With Ca.sup.2+ (seconds) Without Ca.sup.2+ (seconds) 1 mL (23) 12.3 44.9 4 mL (6) 34.3 101
[0535] EDTA is a metal chelator which chelates the calcium and in turn prevents the normal clotting process from occurring.
[0536] These results show that the calcium-dependent prothrombin activators carinactivase-1 (CA-1) and carinactivase-2 (CA-2) did not clot EDTA plasma even at 50 nM concentration in the pre-requisite time of <5 minutes, although they were able to after re-calcification. The effect on ecarin was significant: without re-calcification the clotting time was more than 5 minutes. The addition of calcium in excess caused only a moderate decrease in clotting time with PtPA and OsPA (for example, PtPA at 0.1 nM, 190 seconds to 98.6 seconds). Although other examples described herein have suggested that notecarin is less effective as a procoagulant than PtPA and OsPA, the results showed it was the prothrombin activator that was the least affected by the absence of calcium. The addition of calcium in excess caused 3-fold increase in the clotting of the plasma with thrombin.
[0537] The results from the whole blood clotting study are shown in Table 35.
TABLE-US-00036 TABLE 35 TEG results showing the effect of different procoagulants on EDTA blood with and without the presence of calcium. Procoagulant (nM) R (min) MA (mm) TMA (min) Ecarin 53 (re-calcified) 2.5 56.8 22.1 53 (not re-calcified) 16.9* ND ND Carinactivase-1 53 (re-calcified) 4.8 57.6 29.5 53 (not re-calcified) 16.9* ND ND Carinactivase-2 53 (re-calcified) 2.6 55.7 25.6 53 (not re-calcified) 34.8* ND ND OsPA 53 (re-calcified) 1.6 41.5 21.6 53 (not re-calcified) 17.7* ND ND PtPA 53 (re-calcified) 0.5 58.2 22.9 53 (not re-calcified) 17.8* ND ND 1 (re-calcified) 1.6 64.0 22.9 PtPA 690 nM (Run 1) (not re-calcified) 50.1* ND ND 690 nM (Run 2) (not re-calcified) 50.3* ND ND Notecarin 53 (re-calcified) 1.9 54.2 25.8 53 (not re-calcified) 17.2* ND ND Thrombin 53 (re-calcified) 2.6 49.9 31.5 53 (not re-calcified) 45.4* ND ND BD RST 23 (re-calcified) 1.6 45.4 24.0 23 (re-calcified) 1.8 57.7 28.3 6 (re-calcified) 30.1* ND ND 6 (re-calcified) 32.5* ND ND Re-calcified EDTA blood alone Run 1 11.2 58.5 40.8 Run 2 10.6 56.7 38.5 *means measurement was stopped, and ND means this was not determined.
[0538] This experiment showed that EDTA whole blood presents a difficult challenge for the procoagulants. Unlike with EDTA plasma, none of the procoagulants were able to clot EDTA blood in the absence of re-calcification. Clotting was not achieved even with very high PtPA concentration (690 nM). However, all procoagulants were able to clot the EDTA blood when it was re-calcified. Rapid and complete clotting was achieved even with a very low PtPA concentration (1nM). EDTA is the anticoagulant of choice for preserving cellular morphology, and by chelating calcium the EDTA also prevents platelet activation and platelet clumping.
[0539] Two additional plasma clotting experiments using 30 nM PtPA were performed. In the first, water was added to the Clotek tube instead of Tris buffer and the sample still clotted in 6.4 seconds. In the second, no buffer was used, only 200 .mu.L EDTA plus 50 .mu.L PtPA and the clotting time was 5.4 seconds. For the TEG study, the following solutions were added to a TEG cup 50 .mu.L Tris buffer, 60 .mu.L PtPA, 230 .mu.L EDTA blood and still no clotting was observed. Additionally, a BD RST tube was filled to the fill mark with EDTA blood and after 60 minutes it showed very partial clotting. It is obvious from these experiments that whenever EDTA may be present, addition of calcium with the procoagulant should be considered in order to achieve clotting.
Example 5c: Clotting of Blood from Warfarin-Treated Participants
[0540] The effect of PtPA and OsPA on coagulation of blood samples from two participants ("W1" and "W2") on warfarin was determined.
[0541] The coagulation parameters of each participant, determined by standard coagulation protocols, are shown in Table 36, where:
[0542] aPTT=activated partial thromboplastin time;
[0543] PT=prothrombin time;
[0544] INR=international normalised ratio; and
[0545] Fib-D=prothrombin time derived fibrinogen.
TABLE-US-00037 TABLE 36 Coagulation parameters for the participants on anticoagulant treatment Participant aPTT (s) PT (s) INR Fib-D (g/L) W1 30.27 21.3 2.08 1.55 W2 36.15 27.4 2.61 5.20 Normal plasma 30-32 10-11 1.0 2.5-4.0
[0546] The results in Table 36 confirm that the blood from participants W1 and W2 was anticoagulated as expected.
[0547] TEG analysis was then performed. Each TEG assay mixture consisted of 310 .mu.L of citrated blood, 20 .mu.L of CaCl.sub.2 (0.2 M) and 10 .mu.L of saline or solution of prothrombin activator (PtPA or OsPA) resulting in final assay concentrations of prothrombin activator of between 0 and 14.1 .mu.g/mL.
[0548] The TEG results are shown in Tables 37, 38 and 39 below. The values therein are derived from the TEG data as shown in FIGS. 40 and 41.
TABLE-US-00038 TABLE 37 TEG results for clotting of citrated blood of W1 with OsPA OsPA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 12.2 35.5 23.6 50.4 0.0588 4.4 22.9 68.1 59.5 0.588 1.6 16.7 73.8 60.8 5.88 0.6 17.8 67.5 53.0
TABLE-US-00039 TABLE 38 TEG results for clotting of citrated blood of W1 with PtPA PtPA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 12.2 35.5 23.6 50.4 0.0141 9.4 31.5 48.4 54.3 1.41 0.8 18.8 69.7 59.6 14.1 0.4 20.6 65.1 53.1
TABLE-US-00040 TABLE 39 TEG results for clotting of citrated blood of W2 with PtPA PtPA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 12.3 37.6 49.1 60.8 0.0141 7.7 26.1 62.9 66.4 1.41 1.3 17.9 77.7 73.3 14.1 0.4 18.4 79.6 69.4
[0549] TEG plots of the results in Tables 37 and 38 are provided in FIG. 40 where TEG plot A shows the results from Table 37 (OsPA) and TEG plot B shows the results from Table 38 (PtPA). In FIG. 40, for the A plot the trace labelled (i) shows the results for 5.88 .mu.g/mL OsPA, the trace labelled (ii) for 0.588 .mu.g/mL OsPA, the trace labelled (iii) for 0.0588 .mu.g/mL OsPA, and the trace labelled (iv) for 0 .mu.g/mL. In FIG. 40, for the B plot the trace labelled (i) shows the results for 14.1 .mu.g/mL PtPA, the trace labelled (ii) for 1.41 .mu.g/mL PtPA, the trace labelled (iii) for 0.0141 .mu.g/mL PtPA, and the trace labelled (iv) for 0 .mu.g/mL.
[0550] In FIG. 41, the trace labelled (i) shows the results for 14.1 .mu.g/mL PtPA, the trace labelled (ii) for 1.41 .mu.g/mL PtPA, the trace labelled (iii) for 0.0141 .mu.g/mL PtPA, and the trace labelled (iv) for 0 .mu.g/mL.
[0551] In summary, both PtPA and OsPA at low concentrations rapidly clotted blood samples from participants on warfarin therapy, with R times, cc values and MA values similar to those in comparable experiments with blood from a healthy participant (see Tables 28-31 above). These results confirm the observation on warfarin plasma in Example 3c.
Example 5d: Clotting of Blood from a High Dose Heparinised Participant with PtPA and OsPA
[0552] Blood was taken from a participant ("H1") who had been given 35,000 IU of heparin 30 minutes prior to the blood sample being taken. The concentration of heparin was calculated to be approximately 7 IU/mL of blood. Blood was collected in a plain syringe and then transferred to a Greiner Vacuette.TM. citrate tube.
[0553] The coagulation parameters of the participant were measured and are shown in Table 40.
TABLE-US-00041 TABLE 40 Coagulation parameters for the heparinised participant. Participant aPTT PT INR TT PTNH REPT Fib-D (s) (s) (s) (s) (s) (g/L) H1 failed 41.2 3.75 failed 17.75 15.28 2.40 All coagulation parameters in Table 40 are known in the art, being: aPTT = activated partial thromboplastin times; PT = prothrombin time; INR = international normalised ratio; TT = thrombin time; PTNH = protamine sulphate time; REPT = repitalise time; Fib-D = prothrombin time derived fibrinogen.
[0554] The results in Table 40 are in the range of expected results for a heavily heparinised sample.
[0555] TEG analysis was then performed. Each TEG assay mixture consisted of 310 .mu.L of citrated blood, 20 .mu.L of CaCl.sub.2 (0.2 M) and 10 .mu.L of saline or solution of prothrombin activator (PA).
[0556] The TEG results are shown in Table 41 below and in FIG. 42.
TABLE-US-00042 TABLE 41 TEG results for clotting of blood from a participant on very high doses of heparin. PA concentration Citrated PA (.mu.g/mL) R (min) TMA (min) .alpha. (deg) MA (mm) 0 n.c. n.c. n.c. n.c. PtPA 14.1 0.4 20.9 72.6 64.3 OsPA 5.9 1.3 12.8 70.6 61.3 In Table 41 above, "n.c." indicates no clotting.
[0557] TEG traces of the results in Tables 41 are provided in FIG. 42 where the trace labelled A shows the results for no prothrombin activator; the trace labelled B shows the results for 14.1 .mu.g/mL PtPA; and the trace labelled C shows the results for 5.9 .mu.g/mL OsPA.
[0558] At the concentration tested both PtPA and OsPA were able to clot citrated blood from the heparinised participant rapidly. The values for the TEG parameters monitored (R-time, TMA, angle, MA) were comparable with the results obtained from blood of a healthy participant, indicating that the activity of the prothrombin activators was not inhibited by the concentration of heparin present in the blood sample.
Example 5e: Clotting of Blood from a Fully Heparinised Participant with a PtPA-Containing Tube and a BD RST Tube
[0559] A 10 mL blood sample was collected from a participant undergoing cardiac surgery who had been given 38,000 IU of heparin 30 minutes before sample collection. The blood was collected in a plain syringe and within 10 minutes delivered to the laboratory for analysis.
[0560] The BD RST tube was filled to the fill mark, inverted/mixed for 30 seconds and 340 .mu.L of this blood was transferred to a TEG cup in channel 1.
[0561] 330 .mu.L of the original heparinised participant blood sample was added to 10 .mu.L of PtPA (final concentration 1.41 .mu.g/mL) in a TEG cup in channel 2.
[0562] The results of the TEG analysis are shown in Table 42 and in FIG. 43.
TABLE-US-00043 TABLE 42 TEG analysis results for heparinised blood in BD RST tube and with PtPA Sample R (min) K (min) Angle (deg) MA (mm) TMA (min) PtPA 0.7 1.3 74.4 43.8 9.8 (outer trace) BD RST 6.2 N/A 7.8 11.5 24.0 tube (inner trace)
[0563] For the BD RST tube, the R time was 6.2 minutes with a maximum amplitude (MA) value of 11.5 mm, indicating that only partial clotting was achieved. This clot strength did not increase further during the 2 hour period of the analysis.
[0564] With PtPA, the R time was 0.7 minutes, the maximum amplitude (MA) was 43.8 mm and the time to maximum amplitude was 9.8 minutes indicating that a strong and stable clot was achieved quickly.
[0565] The results from Table 42 are also shown in FIG. 43 as a TEG plot where trace "A" is the trace of the TEG cup in channel 2 (PtPA) and trace "B" is the trace of the TEG cup in channel 1 (BD RST tube).
[0566] The blood collected in the plain syringe, and the blood in the BD RST tube were also monitored visually for clotting. No clotting was observed in the plain syringe for up to 2 days. Blood in the BD RST tube showed partial clotting at 30 minutes and complete clotting within 24 hours.
[0567] In summary, the heavily heparinised sample clotted rapidly and completely in a tube containing PtPA but only very slowly and incompletely in a BD RST tube.
Example 5f: Clotting of Blood Samples Containing Heparin by Prothrombin Activators
[0568] The ability of venom prothrombin activators to clot blood from a healthy participant collected in a Greiner lithium heparin tube (18 IU heparin per mL of blood) was determined using TEG analysis and the results are shown in Table 43. The citrate sample was collected for the comparison of the MA values.
TABLE-US-00044 TABLE 43 TEG results with blood collected in Greiner lithium heparin tube from a healthy participant. Procoagulant (nM) R (min) MA (mm) TMA (min) Ecarin 56 (black) 9.9 18.8 37.8 32 (green) 13.8 18.6 39.8 Carinactivase-1 100 (black) 35.6* ND ND 50 (green) 34.8* ND ND Carinactivase-2 56 (black) 10.4 23.0 37.8 34 (green) 18.0 18.5 47.5 OsPA (nM) 56 (magenta) 1.9 37.6 24.7 34 (pink) 2.3 36.5 26.1 9 (green) 2.9 29.8 25.2 3 (black) 10.3 6.2 20.0 PtPA 56 (magenta) 2.3 33.5 26.6 34 (brown) 2.3 39.8 27.2 9 (green) 2.8 31.3 25.7 3 (black) 6.8 14.7 25.8 Notecarin 56 (black) 2.5 34.1 30.7 32 (green) 7.9 18.0 28.4 BD RST 25 (black) 35.6* ND ND 6 (green) 35.8* ND ND Re-calcified citrate blood Run 1 6.3 61.2 30.1 Run 2 6.5 61.5 27.1 *means measurement was stopped, and ND means this was not determined.
[0569] The results of this experiment with lithium heparin whole blood showed that some of the procoagulants were capable of clotting the heparinised blood at lower procoagulant concentrations, while other procoagulants were unable to clot the blood even at the highest concentration range tested in this experiment. The results show that the group C procoagulants (PtPA and OsPA) were the most effective in clotting the blood collected in Greiner lithium heparin tubes, requiring .gtoreq.9 nM for OsPA and PtPA to achieve R time within 5 minutes. This was significantly lower than the MAs (.about.61) achieved in the recalcified citrated blood. Although not wishing to be bound by theory, it is believed likely that the ATIII-heparin complex inhibits the thrombin formed, and subsequently prevents proper or stronger clot formation. The minimum concentrations required for the other prothrombin activators are >56 nM for ecarin and carinactivase-2, >100 nM for carinactivase-1, and >32 nM of notecarin. The thrombin concentrations of 6 and 25 nM from the BDRST tubes failed to clot the blood at all.
[0570] Again, although not wishing to be bound by theory, it is believed likely that the group C prothrombin activators (OsPA and PtPA) produced a much larger and more sustained burst of thrombin to overwhelm the ATIII-heparin complex that inhibits both thrombin and FXa and subsequently the clotting process.
Example 5g: Clotting of Normal "Pooled" Citrated Plasma Spiked with Different Concentrations of Heparin
[0571] This experiment was performed to determine the required concentrations of the different prothrombin activators to clot plasma spiked with heparin at different concentrations to mimic levels likely to be found in plasma from heparinised patients.
[0572] Normal "pooled" citrated plasma had the total calcium adjusted to 2.71 mmol/L and then aliquots of heparin were added to give final heparin concentrations in the Clotek Tube of 0.8, 2.0, 10 and 20 IU/mL which is the range that may be encountered in patient blood samples. Additionally 1 mL aliquots of the citrated plasma were dispensed in Greiner lithium heparin tubes, and then mixed on a roller for 15 minutes to give lithium heparin concentration of 90 IU/mL. The Clotek tube contained 100 .mu.L of sodium or lithium heparin plasma, 100 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4) and 50 .mu.L of each procoagulant.
[0573] The clotting results for each prothrombin activator with the heparin spiked plasma samples are shown in Tables 44-49, and for thrombin is shown in Table 50.
TABLE-US-00045 TABLE 44 Clotting by PtPA of normal "pooled" citrated plasma spiked with heparin or dispenses in Greiner lithium heparin tubes PtPA (nM)/Clotting Time (secs) Heparin 120 60 36 30 20 (IU/mL) nM nM nM nM nM 12 nM 0 3.3 4.3 5.2 5.5 6.4 10.1 0.8 3.5 5.3 7.4 7.6 10.4 14.0 2.0 6.2 9.8 12.7 15.7 16.6 26.5 10 25.3 39.1 46.1 60.2 72.3 142 20 38.0 47.2 68.2 80.6 232 >300 90 (Greiner Li 17.4 27.7 53.7 69.5 228 >300 Hep tube)
TABLE-US-00046 TABLE 45 Clotting by OsPA of normal "pooled" citrated plasma spiked with heparin or dispenses in Greiner lithium heparin tubes OsPA (nM)//Clotting Time (secs) Heparin 120 60 36 30 20 12 (IU/mL) nM nM nM nM nM nM 0 6.9 7.2 8.3 8.6 8.9 9.4 0.8 9.1 10.8 12.9 13.7 16.4 21.7 2.0 13.8 15.2 17.6 19.1 22.6 27.9 10 27.3 35.2 49.2 56.5 86.1 270 20 93.9 120 >300 >300 >300 >300 90 (Greiner Li >300 >300 >300 >300 >300 >300 Hep tube)
TABLE-US-00047 TABLE 46 Clotting by notecarin of normal "pooled" citrated plasma spiked with heparin or dispenses in Greiner lithium heparin tubes Notecarin (nM)/Clotting Heparin Time (secs) (IU/mL) 120 nM 60 nM 36 nM 12 nM 0 44.3 55.1 61.6 87.0 0.8 49.8 65.0 78.3 126 2.0 57.0 92.2 129 262 10 >300 >300 >300 >300 20 >300 >300 >300 >300 90 (Greiner Li >300 >300 >300 >300 Hep tube)
TABLE-US-00048 TABLE 47 Clotting by ecarin of normal "pooled" citrated plasma spiked with heparin or dispenses in Greiner lithium heparin tubes Heparin Ecarin (nM)/Clotting Time (secs) (IU/mL) 120 nM 60 nM 36 nM 12 nM 0 28.5 40.0 49.3 92.3 0.8 30.1 41.3 51.8 109 2.0 33.1 50.9 70.7 156.6 10 75.7 142 269 >300 20 140 >300 >300 >300 90 (Greiner Li >300 >300 >300 >300 Hep tube)
TABLE-US-00049 TABLE 48 Clotting by carinactivase-1 of normal "pooled" citrated plasma spiked with heparin or dispenses in Greiner lithium heparin tubes Heparin CA-1(nM)//Clotting Time (secs) (IU/mL) 120 nM 60 nM 36 nM 12 nM 0 170 258 >300 >300 0.8 215 297 >300 >300 2.0 >300 >300 >300 >300 10 >300 >300 >300 >300 20 >300 >300 >300 >300 90 (Greiner Li >300 >300 >300 >300 Hep tube)
TABLE-US-00050 TABLE 49 Clotting by carinactivase-2 of normal "pooled" citrated plasma spiked with heparin or dispenses in Greiner lithium heparin tubes Heparin CA-2 (nM)/Clotting Time (secs) (IU/mL) 120 nM 60 nM 36 nM 12 nM 0 80.6 120 168 267 0.8 87.8 126 181 >300 2.0 130 230 >300 >300 10 >300 >300 >300 >300 20 >300 >300 >300 >300 90 (Greiner Li >300 >300 >300 >300 Hep tube)
TABLE-US-00051 TABLE 50 Clotting by thrombin of normal "pooled" citrated plasma spiked with heparin or dispenses in Greiner lithium heparin tubes Heparin Thrombin (nM)/Clotting Time (secs) (IU/mL) 100 60 30 10 0 5.1 7.0 13.5 24.7 0.8 23.2 >300 >300 >300 2.0 >300 >300 >300 >300 10 >300 >300 >300 >300 20 >300 >300 >300 >300 90 (Greiner Li >300 >300 >300 >300 Hep tube)
[0574] As was shown in the previous experiment (Example 5f) the presence of heparin (and thrombin inhibition due to the ATIII-heparin complex) presents a major challenge for the procoagulants in preventing partial or complete clotting of blood. The results of this experiment showed the relative effectiveness of the procoagulants in clotting of heparinised plasma.
[0575] The results show that PtPA and OsPA were the most effective prothrombin activators in overcoming the anticoagulant effect from the ATIII-heparin complex. PtPA required the lowest concentration (12-20 nM) to clot plasma containing highest heparin concentrations that may be encountered in patient samples. One interesting finding was PtPA was the only prothrombin activator able to effectively clot the heparinised plasma from the Greiner lithium heparin tube in <5 minutes with all but the lowest PtPA concentration of 12 nM even though the heparin concentration was 90 IU/mL compared with the spiked sample containing 20 IU/mL heparin. It was quite possible the PtPA only was aided by the surfactant contained in the Greiner lithium heparin tube. OsPA required >36 nM and ecarin >60 nM to be able to clot the plasma containing 20 IU of heparin, and both were unable to clot the heparinised plasma from the Greiner lithium heparin tube in <5 minutes respectively. Carinactivase-2 and notecarin were unable to clot the plasma containing .gtoreq.10 IU of heparin while the carinactivase-1 was unable to clot the plasma containing >0.8 IU of heparin in the pre-requisite time of <5 minutes even when the concentration was 120 nM. The thrombin study showed thrombin concentration of 100 nM was only able to clot the sample spiked with the lowest heparin concentration, 0.8 IU/mL. Thus, when considering the BD RST tube which contains 135 nM of thrombin it is unlikely to clot samples containing heparin >2 IU/mL in the prerequisite time of 5 minutes.
Example 5h: Clotting of Lithium Heparin Plasma from a Healthy Participant
[0576] Samples collected in Greiner lithium heparin tubes are likely to be the most highly heparinised samples a laboratory will receive, particularly if the tubes are not filled to the recommended fill volume. The purpose of this experiment was to provide a guide on the minimum concentration required of the procoagulants to clot all other samples that may be received from heparin anticoagulated patients and produce high quality serum.
[0577] Greiner lithium heparin tubes were filled to the recommended fill mark, giving a heparin concentration of 18 IU/mL. The Clotek tubes contained 100 .mu.L of lithium heparin plasma, 100 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4) and 50 .mu.L of each procoagulant. The TEG cups contained 60 .mu.L procoagulant and 260 .mu.L of the participant blood collected in Greiner lithium heparin tube.
[0578] For the BD RST tube, deionised water was added to two tubes (BDRST1-1 mL; BDRST2-4 mL), allowed to mix on a roller for 30 minutes to dissolve the thrombin and the content was used as with the other procoagulants.
[0579] The clotting results for each procoagulant with the lithium heparin plasma are shown in Table 51.
TABLE-US-00052 TABLE 51 Clotting study with lithium heparin plasma collected in Greiner Lithium Heparin Tube. [Prothrombin Clotting Time (s) activator] (nM) Ecarin CA-1 CA-2 PtPA OsPA Notecarin Thrombin 120 69.0 >300 100.1 7.5 15.6 47.8 >300 60 122.6 >300 180.3 10.7 18.1 62.7 >300 36 175.2 270.6 13.1 21.0 82.4 >300 12 >300 >300 32.7 37.1 269.3 4 >300 >300 78.5 102.1 >300 1 >300 >300 207.8 >300 >300 0.1 >300 >300 >300 >300 0.01 >300 >300 >300 >300
[0580] The experiment design was to show the effectiveness of the procoagulants in clotting heparinised plasma from blood collection tubes. It was found that the minimum required concentration range to achieve clotting of <5 minutes for ecarin and carinactivase-2 was 12 to 36 nM, >120 nM for carinactivase-1, and 4 to 12 nM for notecarin. For the thrombin, concentration of >120 nM was necessary. The BD RST tube was tested as well with 7.7 and 27 nM of thrombin and both concentrations producing clotting time >5 minutes which suggested that the BDRST may be ineffective in clotting samples from patients on high doses of heparin. The results show the group C prothrombin activators (PtPA and OsPA) were easily the most effective in clotting the lithium heparin plasma obtained from blood of a healthy participant collected in Greiner lithium heparin tubes, requiring between 0.1 to 1 nM of PtPA and between 1 to 4 nM of OsPA respectively to achieve clotting in <5 minutes.
Example 5i: Clotting of Blood and Plasma Obtained from a "Fully Heparinised" Cardiac Surgery Participant
[0581] The highest concentration of heparin given intravenously is in complex surgical procedures such as cardiac surgery. If samples are collected for biochemistry during the maximum heparinisation period the results usually need to be provided in the shortest possible time. If serum is used, this will require the clotting to be completed by the procoagulant by the time these serum samples arrive in the laboratory, say in <10 minutes.
[0582] The heparin that is given to patients is sodium heparin while in blood collection tubes or syringes it is lithium heparin. The maximum amount of heparin infused in such patients is .about.45000 IU. These patients are also haemodiluted (i.e., there is an increase in blood plasma volume to .about.4 L), giving a heparin concentration of .about.10 IU per mL of plasma.
[0583] This experiment was designed to determine whether the selected procoagulant concentrations were able to clot "fully" heparinised cardiac surgery patient blood and produce high quality serum. The participant had received 37000 IU of heparin and the sample was collected .about.30 minutes post heparin infusion, equating to .about.9.3 IU of heparin per mL of plasma. A portion of the blood sample was centrifuged to obtain plasma.
[0584] For plasma clotting, the Clotek tube contained 100 .mu.L of cardiac participant plasma, 100 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4) and 50 .mu.L of each procoagulant.
[0585] For the whole blood clotting, the TEG cup contained 60 .mu.L of procoagulant and 260 .mu.L of the participant blood. Being fully heparinised the blood did not clot on its own.
[0586] For the BD RST-1 experiment, the contents of two BDRST tubes were dissolved with 1 and 4 mL of distilled water respectively, and 60 .mu.L of the solution used in the TEG cup as the procoagulant.
[0587] For the BDRST-2 experiment, the tubes were filled with 1 and 4 mL of blood respectively, the tubes were inverted 10 times (.about.30 secs) and 340 .mu.L of the blood was transferred to the TEG cup.
[0588] The clotting results for each procoagulant with the patient's plasma are shown in Table 52, and the whole blood results are shown in Table 53.
TABLE-US-00053 TABLE 52 Clotting times in seconds of plasma from a "fully heparinised" cardiac surgery participant with the different prothrombin activators, thrombin and BD RST tube contents at different concentrations. Procoagulant cone (nM) PtPA OsPA Ecarin CA-1 CA-2 Notecarin Thrombin BDRST 120 10.0 ND 64.8 175 66.5 ND >300 >300 (27 nM) 60 14.8 18.7 89.7 >300 95.1 68.6 >300 >300 (6 nM) 36 15.9 27.3 >300 >300 124 84 >300 ND 12 36.3 35.5 >300 >300 >300 163 ND ND 6 65.9 ND ND ND ND ND ND ND 4 301.6 100 >300 >300 >300 179 ND ND 3 ND 123 ND ND ND ND ND ND 1 ND >300 ND ND ND >300 ND ND ND means this was not determined.
[0589] The results from this experiment showed the concentrations of each procoagulant required to clot the plasma in the prerequisite time of <5 minutes. The maximum activator concentration giving clotting time of about 5 minutes were: PtPA 4 nM, OsPA 1-3 nM, notecarin 3 nM, carinactivase-2>12 nM, ecarin >36 nM, carinactivase-1>60 nM, and thrombin >120 nM. It was surprising to note that the notecarin efficacy was comparable to the group C activators.
TABLE-US-00054 TABLE 53 Results of the TEG clotting study with blood from a "fully heparinised" cardiac surgery participant with the different procoagulants. Procoagulant (nM) R (min) MA (mm) TMA (min) Ecarin 56 2.3 47.6 19.4 34 3.7 49.0 25.1 11 16.6 35.2* 53.3* 3 28.8* ND ND Carinactivase-1 112 5.3 36.9 23.7 56 9.8 33.1* 37.7* Carinactivase-2 56 2.8 38.1 25.8 34 3.3 36.9 24.3 11 8.9 30.7* 38.5* OsPA (nM) 56 0.2 64.2 18.6 3 2.9 61.4 25.8 2.3 4.2 54.8 28.4 2 11.0 29.4* 37.9* PtPA 56 0.4 62.9 20.1 6 4.4 60.9 27.9 3 9.7 42.8* 48.6* 2 28.7* ND ND Notecarin 56 0.8 51.9 23.3 34 1.1 51.0 19.6 11 2.2 46.6 22.2 3 7.2 39.5 33.6 BD RST-1 25 18.9* ND ND 6 19.3* ND ND BD RST-2 135 (1 mL) 0.4 9.9 13.3 34 (4 mL) 0.3 12.7 8.1 *means measurement was stopped, and ND means this was not determined.
[0590] The concentrations determined to achieve an R time of <5 minutes and maximum clot strength from these results for the different prothrombin activators were: PtPA 6 nM, OsPA 2.3 nM, notecarin 3-11 nM, ecarin 11-34 nM, carinactivase-2 11-34 nM, and carinactivase-1 57-112 nM.
[0591] Surprisingly, the notecarin produced clots with higher MAs than the group A and B prothrombin activators, and without wishing to be bound by theory, it is postulated that additives within the blood of such patients may have aided clotting efficacy.
[0592] The strongest clots formed as indicated by the MA were with the PtPA and OsPA, and the weakest with the carinactivase-1 and carinactivase-2 prothrombin activators.
[0593] The thrombin in the BD RST tubes (BDRST-2 experiment with 135 nM of thrombin) produced incomplete clotting and weak clots as indicated by the MA. Such samples are very likely to form latent clotting in the serum post centrifugation.
Example 5j: Clotting of Normal "Pooled" Citrated Plasma Spiked with Rivaroxaban
[0594] This experiment was designed to examine the effect of one of the new Factor Xa inhibitor anticoagulants, rivaroxaban, on the clotting ability of the procoagulants.
[0595] One rivaroxaban table (10 mg, Mol Wt 435, Xarelto, Bayer Schering Pharm, Germany) tablet was crushed in 5 mL deionised water (2 mg/mL), allowed to mix for 30 minutes, and then centrifuged to remove undissolved particles. The typical therapeutic dose given is 10-40 mg once daily. For a 70 kg person (.about.3000 mL plasma) the total amount is 0.14-0.57 mg/kg, 0.0033-0.013 mg/mL of plasma. The concentrations prepared and tested in the Clotek tube were: 0.0033, 0.0083, 0.017, 0.033, and 0.17 mg/mL.
[0596] The Clotek tubes contained 50 .mu.L of Tris Buffer (150 mM Tris HCl, 150 mM NaCl, pH 7.4), 50 .mu.L 0.2 mM CaCl.sub.2, 100 .mu.L normal "pooled" citrated plasma, 50 .mu.L of Tris buffer or anticoagulant of different concentration, and 50 .mu.L of prothrombin activator (30 nM).
[0597] The clotting times are shown in Table 54.
TABLE-US-00055 TABLE 54 Clotting times (seconds) with normal "pooled" citrated plasma and Rivaroxaban. Pro- coagulant Anticoagulant (30 nM) concentration Rivaroxaban concentration (mg/mL) 0 0.003 0.008 0.017 0.033 0.17 PtPA 4.8 5.8 6.2 7.7 8.8 12.0 OsPA 8.6 8.6 8.8 8.8 9.3 16.9 Ecarin 42.6 50.7 51.6 53.9 58.9 83.9 Notecarin 41.7 42.7 44.9 46.2 47.6 67.3 CA-1 128 133 152 176 194 258 CA-2 60.9 61.1 62.8 66.1 70.6 98.0 Thrombin 7.1 7.0 7.3 7.4 8.0 9.8 BDRST 5.1 5.3 5.2 5.8 5.7 7.7 23 nM
[0598] The results show that rivaroxaban has minimal effect on the procoagulants in the therapeutic concentration range 0.0033-0.013 mg/mL of plasma that may be encountered in patients. However, at higher concentrations, rivaroxaban had a significant inhibitory effect on clotting in the presence of each of the procoagulants. This effect may be due to inhibition of Factor Xa produced from human FX in the plasma by the thrombin generated by the procoagulants.
Example 5k: Clotting of Blood from a Patient Undergoing Citrate Anticoagulation
[0599] In previous experiments, it has been shown that the prothrombin activators are capable of clotting citrated blood collected in Greiner citrate tubes. The purpose of this experiment was to confirm that in vivo citration presented no clotting problems with the prothrombin activators.
[0600] In this example, the patient was on 3.0 mmol of citrate per litre of blood flow. The blood was collected in a 50 mL plain syringe (BD Plastipak #300866) for routine pathology testing from which <3 mL was left to perform the very limited study with PtPA only.
[0601] The coagulation parameters were: PT 13 s (RR 9-13), INR 1.3, APTT 45 s (RR 24-39), fibrinogen (derived) 7.2 g/L (RR 1.7-4.5), confirming anticoagulation.
[0602] The TEG cup contained 20 .mu.L of 0.2 M CaCl.sub.2 or saline, 5 .mu.L of PtPA and 320 .mu.L of the citrated blood.
[0603] The results are shown in Table 55.
TABLE-US-00056 TABLE 55 TEG results for citrated anticoagulated participant. Citrated anticoagulated participant R (min) MA (mm) TMA (min) Re-calcified citrate blood alone 4.8 77.1 18.0 2.8 nM PtPA re-calcified 0.6 74.7 15.8 2.8 nM PtPA non re-calcified 0.8 76.2 18.9
[0604] The results indicated that the citrate concentration used for anticoagulation of patients did not present clotting problems when PtPA was used as the procoagulant in re-calcified and non re-calcified blood. The citrate concentration in blood collected in Greiner Citrate tubes is 3.2% or 109 mM, which is about 36 times the concentration the patient was receiving. Additionally, it was not expected there would be any clotting problems as patients being citrated are continuously monitored for ionised calcium and ionised calcium is replenished as required. The R time in normal "pooled" citrate blood was 6.4 minutes (as shown in Table 43) compared to the 4.8 min in this patient. The re-calcification adds calcium in exceeds to requirement thus theoretically it should not contribute to the difference. It is postulated that the difference is due to higher concentrations of clotting factors including fibrinogen which was 7.2 g/L.
Example 6: Clotting of Whole Blood Samples from Participants with Prolonged Coagulation Profile or Low Platelet Counts
[0605] Some patients present with prolonged coagulation parameters and/or reduced platelet count as a result of drug treatment or genetic factors (e.g. haemophilia). The following experiments were conducted to determine whether PtPA or OsPA would be able to rapidly clot blood samples from such patients.
Example 6a: Participants with Prolonged Clotting Time
[0606] Blood from 4 participants ("A", "B", "C" and "D") with prolonged aPTT indicating a defect in the intrinsic pathway were obtained from Pathology Queensland, Princess Alexandra Hospital, Queensland, Australia. The coagulation parameters of each participant were determined and are shown in Table 56.
TABLE-US-00057 TABLE 56 Coagulation parameters* of each participant Participant aPTT(s) PT(s) INR TT(s) Fib-D (g/L) plat (.times.10.sup.9/L) A 48 16 1.6 19 7.1 453 B 44 34 3.0 16 4.1 97 C >200 n.d. n.d. n.d. n.d. n.d. D 45 16 1.5 15 1.5 96 Reference 25-38 8-14 0.9-1.3 10-15 1.5-4.0 140-400 Interval *aPTT = activated partial thromboplastin times; PT = prothrombin time; INR = international normalised ratio; TT = thrombin time; Fib-D = prothrombin time derived fibrinogen; Plat = platelets, n.d. = not able to be determined. The Reference Intervals were obtained from Pathology Queensland.
[0607] TEG analysis was performed. Each TEG assay mixture consisted of 310 .mu.L citrated blood from A-D participant samples, 20 .mu.L of CaCl.sub.2 (0.2 M) and 10 .mu.L of saline or PtPA (final concentration 1.41 .mu.g/mL). CaCl.sub.2 was added at zero time to initiate clotting.
[0608] TEG traces for the four samples are shown in FIG. 44 and TEG parameters from these traces are shown in Table 57.
TABLE-US-00058 TABLE 57 TEG results for clotting of blood samples from participants with abnormal coagulation parameters R (min) TMA (min) .alpha. (deg) MA (mm) A >35 n.d. n.d. n.d. A + PtPA 1.3 16.6 70.9 67.4 B 23.0 >39 17.0 n.d. B + PtPA 0.9 21.3 70.0 59.0 C >52 n.d. n.d. n.d. C + PtPA 2.1 24.5 67.6 60.4 D 16.1 43.3 26.4 31.7 D + PtPA 1.4 19.2 52.0 40.8 "n.d." indicates that this was not measured
[0609] TEG plots for the four samples are shown in FIG. 44 where TEG plot A represents participant A and in that plot the two traces labelled (i) represent the saline control with added calcium; the two traces labelled (ii) are the samples with PtPA. TEG plot B represents participant B and in that plot the two traces labelled (i) are saline control with added calcium; the two traces labelled (ii) are the samples with PtPA. TEG plot C represents participant C and in that plot the trace labelled (i) is the saline control; and the four traces labelled (ii) are samples with PtPA added. TEG plot D represents participant D and in that plot the two traces labelled (i) are the saline controls; and the two traces labelled (ii) are the samples with PtPA. Triplicate determinations were carried out for patient C due to grossly abnormal clotting parameters. The data in Table 57 represents averages from the TEG data.
[0610] In summary, blood samples from participants A-D show an aPTT time ranging from 44 to >200 seconds indicating a defect in the intrinsic coagulation pathway. Without PtPA the samples did not clot or produced very weak clots. In all cases PtPA (1.41 .mu.g/ml) caused rapid clotting, reducing the clotting time to 1-2 minutes. Thus PtPA is effective in rapidly clotting blood from patients with a deficiency in this coagulation pathway.
Example 6b: Participants with Low Platelet Counts
[0611] In participants with low platelet counts, clotting times are significantly increased compared to blood from the healthy population. This is because platelets provide a phospholipid surface for the formation of the prothrombinase complex and accelerate blood clotting.
[0612] Blood samples from four participants with low platelet counts ("E", "F", "G" and "H") were obtained from Princess Alexandra Hospital, Queensland, Australia. The coagulation parameters of each participant were determined and are shown in Table 58, confirming the low platelet count.
TABLE-US-00059 TABLE 58 Coagulation parameters of each participant Participant aPTT(s) PT(s) INR TT(s) fib (g/L) plat (.times.10.sup.9/L) E 31 10 1.1 n.d. 4.5 10 F 29 14 1.4 n.d. 1.2 29 G 34 12 1.2 n.d. 2.7 18 H 54 11 1.2 21 6.4 46 Reference 25-38 8-14 0.9-1.3 10-15 1.5-4.0 140-400 Interval n.d. indicates that the measurements were not determined.
[0613] TEG analysis was performed. Each TEG assay mixture consisted of 310 .mu.L citrated blood from E-H participant samples, 20 .mu.L of CaCl.sub.2 (0.2 M) and 10 .mu.L of saline or solution of prothrombin activator (either PtPA at a final concentration of 1.41 .mu.g/mL or 14.1 .mu.g/mL, or OsPA at a final concentration of 0.59 .mu.g/mL or 5.9 .mu.g/mL).
[0614] The TEG traces are shown in FIG. 45 and TEG parameters are listed in Tables 59 and 60.
TABLE-US-00060 TABLE 59 TEG results for clotting of blood samples from participants with low platelet counts with PtPA Participant .mu.g/mL R (min) TMA (min) .alpha. (deg) MA (mm) E 0 10 30.3 42.3 34.6 1.41 1.5 22.5 61.6 35.3 14.1 0.4 20.3 73.2 31.1 F 0 9.3 29.8 15.6 28.2 1.41 1.5 24.2 29.0 31.2 14.1 0.6 24.3 38.4 28.0 G 0 12.8 32.8 29.5 30.2 1.41 1.4 23.4 57.1 33.7 14.1 0.4 25.0 66.9 31.3 H 0 9.1 32.0 39.3 48.2 1.41 1.7 24.2 66.3 58.8 14.1 0.3 22.2 73.7 54.7
TABLE-US-00061 TABLE 60 TEG results for clotting of blood samples from participants with low platelet counts with OsPA Participant .mu.g/mL R (min) TMA (min) .alpha. (deg) MA (mm) E 0 10 30.3 42.3 34.6 10 3.8 23.8 41.0 32.7 100 0.9 21.8 63.7 32.9 F 0 9.3 29.8 15.6 28.2 10 3.2 25.3 27.4 28.7 100 1.6 25.6 27.5 29.3 G 0 12.8 32.8 29.5 30.2 10 3.3 24.6 38.8 31.7 100 0.9 24.5 56.5 29.9 H 0 9.1 32.0 39.3 48.2 10 3.4 24.1 59.8 59.7 100 0.8 22.6 66.7 54.9
[0615] The TEG plots are shown in FIG. 45 where plot A shows the results for participant E; plot B shows the results for participant F; plot C shows the results for participant G; and plot D shows the results for participant H. In each plot, the trace labelled (i) represents no PtPA or OsPA; the trace labelled (ii) represents 14.1 .mu.g/mL PtPA; the trace labelled (iii) represents 1.41 .mu.g/mL PtPA; the trace labelled (iv) represents 5.88 .mu.g/mL OsPA; and the blue trace represents 0.588 .mu.g/mL OsPA.
[0616] In summary, both PtPA and OsPA clot blood from participants with low platelet counts rapidly to give high strength clots.
Example 7: Comparison of Clotting Activity of Protrhombin Activator-Containing Venoms
[0617] Five freshly reconstituted venoms from the snakes Pseudonaja textilis (Pt), Oxyuranus scutellatus (Os), Oxyuranus microlepidotus (Om), Notchis scutatus (Ns), and Echis carinatus (Ec) were reconstituted in distilled water with a stock concentration of 50 mg/mL. A working stock dilution of 6 mg/mL was freshly prepared and their clotting activity was measured on the clotting of re-calcified citrated plasma. The venoms were serially diluted in duplicates of each concentration in the assay from 2 mg/mL to 500 pg/mL. The results shown are the average of the duplicate measurements. The assay consisted of: 100 .mu.L of 0.02 M Hepes buffer pH 7.4 with added 10 mM calcium, plus 100 .mu.L of citrated plasma and clotting time (in seconds) was measured from the time of the addition of venom dilution.
[0618] The results are shown in Table 61 and FIG. 46.
TABLE-US-00062 TABLE 61 Clotting times of snake venoms containing prothrombin activators at different concentrations Venom Clotting time (seconds) (mg/mL). Os Pt Om Ns Ec 2 59.5 20.6 14.6 27.4 10.3 1 28.4 9.8 9.8 26.5 9.7 0.5 15 5.2 7.8 29.2 10.9 0.25 9.3 5 6.6 36 14.7 0.125 7.6 4.2 6.1 42.7 17.1 0.063 7.4 3.6 6.8 46.4 19.8 0.031 6.5 3.5 6.9 52.00 24.3 0.015 8.2 3.6 8.0 59.9 28.1 0.0075 8.6 3.7 8.6 67.2 33.4 0.0037 9.2 3.8 9.3 75.7 42.1 0.002 10.6 4.0 10.7 78.8 48.6 0.001 11.2 4.2 11.1 86.00 54.6 0.0005 12.4 4.5 12.6 92.5 61.4 0.00025 12.8 6.2 13.0 105.6 72.3 0.00013 14.5 11.2 14.8 132.4 76.4 0.00006 20.4 12.6 21.1 165.5 82.5 0.00003 21.8 14.3 22.4 192.6 91.3 0.00001 24.6 16.5 25.6 209 106.4 0.0000005 30.7 18.6 31.2 235.4 123.2
[0619] As can be seen in both Table 61 and FIG. 46, all five venoms clotted recalcified citrated plasma very efficiently down to concentration of 500 pg/mL. The Group C prothrombin activator-containing venoms (Pt, Os and Om) were the most active achieving clotting times of less 50 seconds at a concentration of 500 pg/mL.
Example 8: Precipitation and Latent Clotting in Plasma and Serum Samples Prepared Using Commercially Available Tubes or Using PtPA and Effects on Troponin I Estimation
[0620] As discussed in the earlier parts of this specification, a number of problems have been associated with the use of commercial serum and plasma tubes such as latent clotting or no clotting (serum tubes), microclots and fibrinogen strings (serum or plasma tubes), and precipitation of proteins and leakage of cellular material (serum or plasma tubes). To investigate whether a tube containing a prothrombin activator would exhibit some of these problems, blood samples in commercially available tubes were visually inspected and compared against blood samples in PtPA-containing tubes. The results are shown in FIGS. 47-49.
Example 8a: Precipitation and Latent Clotting
[0621] FIG. 47 shows the results of plasma preparation from a blood sample collected in a commercially available Greiner plasma (lithium heparin containing) tube. Four sections of the tube are marked (from top to bottom of the Figure): plasma; gelatinous precipitate; separator gel; cells packed by centrifugation. The gelatinous precipitate is composed of fibrinogen and other plasma proteins and forms when the tube is stored at 2-8.degree. C. This precipitate has the potential to interfere with instrument function and analyte determination. Such precipitates occur in .about.5% of lithium heparin plasma samples.
[0622] FIG. 48 shows the results of serum preparation in three commercially available serum tubes: left tube (GS) is a Greiner serum tube; middle tube (BDS) is a BD serum tube and right tube (BDT) is a BD RST tube. Four sections of the tubes are marked (from top to bottom of the Figure): packed cells; separator gel; latent clotting; serum. Post-centrifugation (latent) clotting is evident in the supernatants or serum component of all three tubes. The tubes are inverted to illustrate the presence of the clots. These latent clots have the potential to interfere with instrumentation and functional assays.
[0623] FIG. 49 is in two halves. The left half shows the results of serum preparation from a single blood sample from the same fully heparinised patient in three commercially available tubes (left to right): a Greiner Vacuette.TM. serum tube (GS), a BD Vacutainer.TM. serum tube/SST II (BDS) and a BD RST tube (BDT), and also in a Greiner Vacuette.TM. No Additive tube with PtPA added. Three sections of the tubes are marked (from top to bottom of the Figure): serum; separator gel; packed cells. All four tubes from FIG. 49 were centrifuged then left standing overnight, after which the serum in each tube was transferred to a clear glass tube and photographed as shown in the right half of FIG. 49 (left to right: GS, BDS, BDT and PtPA tubes). The sera from all three commercially available tubes showed evidence of clotting, by clot formation on the side of the tube and observation of fibrin strands, but there was no evidence of precipitation in the serum prepared in the PtPA-containing tube which had a clarity that considerably exceeded that observed with the other samples.
Example 8b: Troponin I Levels
[0624] It has been observed that at least some false positive results showing elevated troponin I (one of the most specific markers of cardiac events) are due to precipitation/latent clotting in serum and plasma samples.
[0625] Troponin I levels in blood samples from 64 participants were measured using serum and plasma samples prepared in commercially available tubes and PtPA-containing tubes.
[0626] Serum and plasma samples were prepared for each participant in five commercially available tubes: Greiner Vacuette.TM. plasma; BD PST II; Greiner Vacuette.TM. serum; BD SST II; and BD RST, and in Greiner Vacuette.TM. No Additive tubes containing PtPA (1.2 .mu.g/4 mL) and analysed for troponin I (TnI) using the Beckman Coulter AccuTnI assay.
[0627] Four discrepant results were obtained as indicated in Table 62 for participants 18, 20, 38 and 63. Participants 18 and 38 were healthy, and participants 20 and 63 were both patients undergoing cardiac surgery and on heparin. Three false positive results (participants 18, 20 and 38) were obtained. Re-centrifugation of an aliquot from the primary tube followed by re-assay gave negative results, as expected when compared to results from the other tubes from the same blood sample. A further result was exceptionally high (participant 63). Re-centrifugation of an aliquot and re-assay gave a comparable result with the other tubes.
[0628] The fact that centrifugation of samples resolved the discrepancy strongly suggests that the discrepant results were due to latent clotting/precipitate formation causing inaccurate sampling. No discrepant results were obtained with the PtPA serum.
TABLE-US-00063 TABLE 62 Troponin I assay results showing discrepancies. Original TnI result (Repeat TnI result) .mu.g/L Greiner BD Greiner Participant PtPA plasma PST II serum BD SST II BD RST 18 0.003 0.000 0.000 0.000 0.053 0.000 (0.004) 20 0.007 0.015 0.093 0.013 0.013 0.004 (0.015) 38 0.009 0.000 0.006 0.006 1.334 0.004 (0.002) 63 0.675 0.665 0.643 0.652 0.542 1.251 (0.690) (0.704) (0.665) (0.650) (0.579) (0.734)
[0629] In summary, precipitation and latent clotting observed during the preparation of plasma and serum samples in commercially available tubes represent significant problems for analyte determination. No precipitation or evidence of microclot formation was observed when PtPA was used to prepare a serum sample.
Example 9: Fibrinogen, Degraded Fibrinogen and Fibrin Levels in Plasma and Serum Samples
[0630] Soluble fibrinogen, soluble partially degraded fibrinogen (fdp) and unpolymerised fibrin (FDP) are undesirable components in serum and plasma samples for analyte determination as discussed in the earlier parts of this specification. Briefly, fibrinogen/fdp/FDP should be minimal in a high quality serum sample to avoid further conversion into insoluble fibrin (microclots) after standard serum preparation conditions especially on standing. Microclots or fibrin strands can interfere with instrumentation and affect analyte determination. For serum samples prepared by standard methods, the concentration of fibrinogen/fdp/FDP in the sample is believed to depend on the extent to which the patient/individual is anticoagulated, the health status of the patient (e.g. the presence of liver disease), on the type of sample collection container and in rare cases on inadequate mixing of the sample in the tube. In this example, the concentration of fibrinogen/fdp/FDP in serum and plasma samples prepared using different tubes was investigated to assist in establishing conditions for preparation of a "high quality" serum sample.
[0631] Serum and plasma fibrinogen/fdp/FDP concentrations were measured by a sandwich enzyme immunoassay (ELISA) as described below. This assay uses a polyclonal antiserum against fibrinogen which recognises soluble soluble fibrinogen, fdp and FDP.
[0632] The ELISA method requires anti-fibrinogen antiserum (as a purified IgG fraction: AHFAS) and anti-fibrinogen antiserum-horseradish peroxidase conjugate (AHFAS-HRP). Preparations of each were purchased from MP Biochemical, USA. International fibrinogen standard was purchased from NIBSC, London, and diluted from a working stock concentration of 1 mg/mL in 50% glycerol/saline.
[0633] The wash buffer consisted of 0.05 M phosphate buffer, pH 7.4, 0.5 M NaCl, 0.05% Tween 20 and 1% BSA, while the binding buffer used was 0.1 M bicarbonate buffer, pH 9.6.
[0634] Working dilutions of the antibodies were prepared by diluting AHFAS and AHFAS-HRP stocks 1:500 in binding buffer. The HRP substrate solution consisted of 20 mM tetramethylbenzidine, 0.4 mL of 30% H.sub.2O.sub.2 and 50 .mu.L of 0.05 M citrate buffer pH 4.0 in a 50 mL reaction solution.
[0635] Nunc ELISA IMMUNOSORB 96 well plates (Thermo Fisher Scientific, Rochester, N.Y.) were coated with 100 .mu.L of AHFAS working dilution per well by incubating the plates with AHFAS at 4.degree. C. overnight. The plates were then blocked with 100 .mu.L of 2% bovine serum albumin (Sigma Chemical, Co) in wash buffer at 4.degree. C. overnight and subsequently washed three times with wash buffer.
[0636] Blood samples from healthy volunteers (healthy) and patients (cardiac and renal disease) were collected into the following tubes:
[0637] Greiner serum tubes (GS or GRS);
[0638] BD serum tubes (BDS);
[0639] BD RST tubes (BDT or BD RST);
[0640] Greiner No Additive tubes with added 1.2 .mu.g PtPA from a PtPA stock solution 4.8 mg/mL (PtPA);
[0641] Greiner No Additive tubes with added 0.5 .mu.g OsPA from a OsPA stock solution 2.0 mg/mL (OsPA);
[0642] Greiner No Additive tubes with added 0.6 U ecarin;
[0643] Greiner No Additive tubes with added 1.2 U ecarin;
[0644] Greiner No Additive tubes with added 0.63 U/4 mL ecarin (Example 9b);
[0645] Greiner No Additive tubes with added 1.25 U/4 mL ecarin (Example 9c);
[0646] Greiner No Additive tubes with added 2.5 U/4 mL ecarin (Example 9c);
[0647] Greiner Vacuette.TM. plasma tubes (GRLH);
[0648] BD Vacutainer.TM. plasma (PST II) tubes (BDLH);
[0649] Greiner Vacuette.TM. citrate tube (3.2% citrate) for plasma (CIT); and
[0650] Greiner Vacuette.TM. K2EDTA tube (1.5-2.2 mg/mL of EDTA) for plasma (EDTA).
[0651] Plasma and serum samples prepared in each tube above were prepared under standard Pathology Queensland (Australia) analyte sample preparation procedure where blood was collected into each tube, and the tube was left to stand prior to centrifugation at 3,000 g. The BD RST, PtPA, OsPA and ecarin-containing tubes were left to stand for 5 minutes, while all remaining tubes were left to stand for 30 minutes for normal blood and 60 minutes for anticoagulated blood.
[0652] From each tube, 100 .mu.L aliquots of the plasma or serum samples (diluted 1/1,000 dilution and 1/10 dilution, respectively) were plated in triplicate. Serial dilutions of the fibrinogen standard (11 dilutions covering a concentration range of 1,000 ng/mL to 10 ng/mL) were plated out in duplicate. The plates were incubated overnight at 4.degree. C.
[0653] The plates were washed six times with wash buffer to remove unbound components and incubated overnight at 4.degree. C. with 100 .mu.L of AHFAS-HRP working solution per well. The plates were again washed 6 times with wash buffer before adding 100 .mu.L HRP substrate solution per well.
[0654] Development of a blue colour was monitored in a dark environment until A.sub.450 nm reached an absorbance of 1.0 at a fibrinogen concentration of 1 .mu.g/mL. The reaction was then stopped by adding 100 .mu.L per well of 1.0 M sulphuric acid solution, yielding a yellow colour. Plates were then read at A.sub.450 nm in a Thermo Scientific Multiskan Ascent plate reader with Ascent software.
Example 9a: Fibrinogen/fdp/FDP Levels in 48 Serum Samples Collected in Greiner Serum Tubes (GS) from Patients
[0655] Fibrinogen/fdp/FDP concentration was measured using the ELISA method outlined above in serum samples from Greiner serum (GS) tubes from 48 randomly selected patients requiring analyte determination for their clinical conditions.
[0656] FIG. 50 shows the results. The concentration of fibrinogen/fdp/FDP in these samples ranged from 4.4-32 .mu.g/mL, compared with fibrinogen concentration in these plasma samples of 2.0-5.0 mg/mL. These data show that greater than or equal to 99% of fibrinogen is removed by the clotting process in standard commercially available serum tubes. These data also provide a reference interval range of fibrinogen in serum preparation in Greiner serum tubes from blood samples from a hospital population.
Example 9b: Fibrinogen/fdp/FDP Levels in Normal Serum and Plasma Samples
[0657] 36 normal serum samples (as determined by prothrombin time, aPTT and fibrinogen assays) were selected to investigate the effect of PtPA addition (300 ng/mL or 1.2 .mu.g/4 mL tube) on the fibrinogen/fdp/FDP levels. Sera were prepared in Greiner serum tubes (primary tubes) under standard Pathology Queensland procedure prior to aliquoting two equal serum samples of 1 mL from each Greiner serum tube into plain plastic tubes (secondary tubes). PtPA (300 ng/mL or 1.2 .mu.g/4 mL tube) was added to one tube and an equal volume (50 .mu.L) saline to the second to provide matched pairs of secondary tubes.
[0658] The fibrinogen/fdp/FDP concentration in each secondary tube was measured using the ELISA method outlined above and the results are shown in FIG. 51. In both secondary tubes residual fibrinogen was reduced to less than 1% of that present in blood. However, addition of PtPA further reduced the fibrinogen/fdp/FDP levels from a mean of 12.8 .mu.g/mL to 11.8 .mu.g/mL. This reduction was significant (p<0.04) in a paired t-test analysis. Thus the PtPA was capable of further reducing serum fibrinogen/fdp/FDP even in normal individual sera in which residual fibrinogen/fdp/FDP levels were very low. The very small amounts remaining (<1.0%) are likely to be molecules which react with antibody but are not able to polymerise to form insoluble clot.
[0659] 9 normal blood samples were then collected into four different serum tubes: Greiner serum, BD serum, BD RST and PtPA-containing Greiner Vacuette.TM. No Additive tubes. The fibrinogen/fdp/FDP concentration in each secondary tube was measured using the ELISA method outlined above and the results are shown in FIG. 52. The fibrinogen concentration was considerably lower in the sera prepared in PtPA-containing Greiner Vacuette.TM. No Additive tube.
[0660] The concentration of fibrinogen/fdp/FDP in serum produced using OsPA and ecarin was then investigated. Serum samples were prepared from blood of 5 normal participants in Greiner serum tubes (GRS) and in Greiner No Additive tubes containing OsPA (0.50 .mu.g/4 mL tube), ecarin (0.63 units/4 mL tube) and PtPA (1.2 .mu.g/4 mL tube). Fibrinogen/fdp/FDP concentration in the sera of each tube was measured using the ELISA method outlined above. The results are shown in FIG. 53. In all cases residual fibrinogen/fdp/FDP levels were less than 1% that in normal blood or plasma.
Experiment 9c: Fibrinogen/fdp/FDP Levels in Serum Samples Prepared from Heparinised Patients
[0661] Patients undergoing renal dialysis require moderate levels of heparinisation (1,000-10,000 U of heparin during the treatment period) to avoid prothrombotic events during dialysis. 3 patients from this category were chosen to test the ability of PtPA tubes (1.2 .mu.g/4 mL tube) and BD RST tubes to efficiently clot blood from dialysis patients in a 5 minute incubation time, compared to the clotting of blood taken in Greiner serum (GRS) and BD serum (BD SST II) blood collection tubes in a 30 minute clotting time period. The effectiveness of clotting was determined by measuring the fibrinogen/fdp/FDP concentration in the respective sera using the ELISA method outlined above.
[0662] The results are shown in FIG. 54 and reveal that the residual levels of fibrinogen/fdp/FDP in the PtPA-produced sera from the heparinised blood were comparable to those values in the PtPA-produced sera from normal blood (FIGS. 51 and 53). In contrast, residual fibrinogen/fdp/FDP levels in sera produced in the GRS, BDS and BD RST tubes were much higher. At these levels, latent clot formation and microclots are especially likely in the presence of heparin. The values in FIG. 54 for GRS, BDS and BDRST are minimal estimates based on a single dilution of sample.
[0663] Blood samples from 2 cardiac surgery patients (treated with 25,000-41,000 Units of heparin) were collected into the following tubes: Greiner plasma tubes, Greiner serum tubes, BD serum tubes, BD RST tubes, Greiner Vacuette.TM. No Additive tubes with PtPA added (1.2 .mu.g/4mL tube), Greiner Vacuette.TM. No Additive tubes with OsPA added (0.50 .mu.g/4 mL tube), Greiner Vacuette.TM. No Additive tubes with 1.25 units/L of ecarin per 4 mL tube added (EC1), or Greiner Vacuette.TM. No Additive tubes with 2.5 units/L of ecarin per 4 mL tube added (EC2).
[0664] The fibrinogen/fdp/FDP concentration in the sera and plasma of each tube was determined using the ELISA method outlined above and the results are shown in FIG. 55. The fibrinogen/fdp/FDP concentration was greatly reduced in serum obtained using the prothrombin activator-containing tubes compared to the plasma and other serum tubes. Furthermore, the time allowed for clotting was only five minutes with the prothrombin activator-containing tubes compared to 30 minutes in the other tubes or 60 minutes in the tubes for the anticoagulated blood.
[0665] In summary, the tubes containing prothrombin activators were able to produce serum samples with a lower concentration of residual fibrinogen/fdp/FDP than in serum samples produced using commercially available serum or plasma tubes. For normal samples, the effect was relatively small. However, with heparinised samples, the prothrombin activators were able to produce serum with low residual fibrinogen compared to serum prepared in commercial serum tubes. The results suggest that a sufficiently low concentration of residual fibrinogen/fdp/FDP should be achievable by using a prothrombin activator to avoid latent clotting or precipitation in all serum samples. This ability to produce "high quality" serum from patients on high anticoagulant doses (e.g. cardiac care, dialysis) and in "fully heparinised" samples from patients undergoing cardiac surgery is useful.
Example 10: Haemoglobin Concentration in Plasma and Serum Samples
[0666] The haemolytic index is used routinely in chemical pathology as a measure of the haemoglobin present in serum and plasma samples and hence the extent of cell lysis. Lysis of all types of blood cells, including erythrocytes, white cells and platelets, can occur in vivo, during blood collection and serum/plasma preparation and on standing. In vitro cell lysis commonly occurs during collection of samples or transfer of samples using small gauge needles or transfer devices which are normally under pressure. In vivo red cell lysis alone may occur in haemolytic anaemias. During the lysis of cells in vitro, cellular content is released into the serum or plasma, falsely altering the results of some analytes; release of cellular content may even cause dilution of other analytes if the haemolysis is extensive. Haemoglobin in plasma or serum may cause spectral problems during analysis and other cellular analytes may cross react. Serum normally contains slightly higher haemoglobin than lithium heparin plasma and this is considered to be due to the clotting process lysing a small number of cells as the clot expands and contracts. A low haemoglobin concentration is therefore an important criterion for a "high quality" serum sample.
[0667] The following experiment was performed to compare the haemoglobin content of serum produced in the presence and absence of PtPA and in plasma produced using lithium heparin. Blood samples were collected from 2 patients undergoing citrate anticoagulation therapy and 9 healthy participants. Samples were collected in Greiner plasma tubes, Greiner serum tubes, and Greiner Vacuette.TM. No Additive tubes containing PtPA (1.2 .mu.g/4 mL of blood). Tubes were centrifuged 5 minutes after collection for the Greiner plasma tubes and Greiner Vacuette.TM. No Additive tubes with PtPA tubes and 30 minutes for the Greiner Vacuette.TM. No Additive tubes (without PtPA). The samples were analysed for haemoglobin within 30 minutes after centrifugation. Results are shown in Table 63.
TABLE-US-00064 TABLE 63 Haemoglobin concentrations in plasma and serum samples. Haemoglobin concentration (mg/L) Greiner Greiner Greiner Vacuette .TM. No Sample plasma serum Additive tubes with PtPA Number of samples 11 11 11 (2 citrated and 9 healthy participants) Mean 55 67 46 Standard deviation 26 28 25
[0668] The mean haemoglobin concentration in the plasma samples was 55 mg/L, somewhat lower than the 67 mg/L found for the Greiner serum, as expected from the literature. The mean value for the PtPA serum was 46 mg/L, considerably lower than that found for Greiner serum and even lower than that found for plasma. The standard deviations of the mean are large because of the individual variation from person to person. They do not give an estimate of the significance of differences between the three types of tubes for any one person. To check for the significance of these differences, paired two tailed t-tests were performed using the data from the 11 blood samples. The results were as follows:
[0669] Greiner plasma v Greiner serum: p=0.1243
[0670] Greiner serum v Greiner Vacuette.TM. No Additive tubes with PtPA: p=0.0188 (statistically significant P<0.05)
[0671] Greiner plasma v Greiner Vacuette.TM. No Additive tubes with PtPA: p=0.1038
[0672] The conclusions that can be drawn from this Example are that:
[0673] (1) The haemoglobin concentration in the plasma samples prepared in the Greiner plasma tubes was lower than that in serum prepared in the Greiner serum tubes, as expected from the literature;
[0674] (2) The haemoglobin concentration in the sera prepared in the Greiner Vacuette.TM. No Additive tubes with PtPA was significantly lower than that in the Greiner serum. The lower level of haemoglobin in these serum samples may reflect the much faster rate of clotting, limiting the amount of cell lysis which occurs during the clotting process; and
[0675] (3) The mean haemoglobin concentration in the PtPA serum samples was lower than that in the plasma samples and the difference was of borderline significance (p=0.1038).
[0676] Thus, the use of the PtPA tube gave sera of higher quality in terms of haemoglobin content than use of the commercial Greiner serum tube.
Example 11: Presence of Cells and Cellular Content in Plasma and Serum Samples
[0677] Lithium heparin plasma prepared in commercially available tubes contains residual cells in suspension or in the buffy coat layer on top of the gel barrier in contact with the plasma after centrifugation. Serum prepared from healthy participant samples in commercially available tubes contains fewer cells in contact with the serum (compared with plasma). Storage of plasma and serum for at least 7 days at 2-8.degree. C. is a regulatory requirement in Australia under the National Pathology Laboratory Accreditation Advisory Council (NPACC) in case of re-analysis or requests for additional analyses. The presence of cells can have two effects during storage and analysis of serum or plasma. Firstly, cells may lyse, releasing cellular contents (e.g. K.sup.+, lactate dehydrogenase) into the serum or plasma. This can lead to significant differences between measurements made immediately after centrifugation and measurements after a period of storage. Secondly, cells continue to be metabolically active and may use up significant amounts of nutrients (e.g. glucose) and release metabolic products (e.g. lactate) on storage. Changes can even be observed in blood samples of many tubes when the samples are stored for the usual recommended 30 minute clotting time when the samples are from healthy participants.
[0678] The degree of cellular contamination is therefore an important quality criterion for serum samples and an important advantage of using serum over plasma.
[0679] The following experiments were performed to compare serum prepared using PtPA with serum prepared using Greiner serum tubes and lithium heparin plasma prepared using Greiner plasma tubes. The extent of contamination of serum and plasma samples by cells and cell contents can be assessed using several markers of contamination. Three markers of cellular contamination that were used here are: increase in lactate dehydrogenase (LD) activity on storage; decrease in glucose concentration on storage; and direct observation of cells.
Example 11a: Comparison of PtPA Serum with Lithium Heparin Plasma from Healthy Participants Prepared in Commercially Available Tubes
[0680] Blood was collected from 10 healthy participants in Greiner plasma tubes, and Greiner serum tubes containing PtPA, prepared as follows. The insides of the Greiner serum tubes were cleaned to remove the Greiner procoagulants/additives by filling the tubes with sterile de-ionized water, inverted .about.20 times, allowed to stand 10 minutes, and then the inside wall and inner part of tube cap were scrubbed with sterile cotton swab without disturbing the gel barrier. The content was discarded and the tubes were further filled/inverted/rinsed 3 times with de-ionised water. The cleaned tubes were finally placed in a drying oven at 40.degree. C. to completely dry any water droplets before the PtPA was dispensed (1.2 .mu.g/4 mL of blood). Blood was then collected into the tubes. The two tubes (Greiner plasma and PtPA) from each participant were centrifuged and immediately analysed for lactate dehydrogenase (LD) and glucose (zero time) levels. The samples were allowed to stand at room temperature (21.degree. C.) and re-analysed .about.8 hours later on the same analyser. Results are shown in Tables 64 and 65.
TABLE-US-00065 TABLE 64 LD activity (U/L) of samples from 10 participants measured at 0 and 8 hours post centrifugation stored at 21.degree. C. Greiner plasma PtPA % % A 0 hr 8 hr 8-0 hr* 8-0 hr* 0 hr 8 hr 8-0 hr* 8-0 hr* Mean 177 199 22 +12.5 175 173 1.3 -0.6 SD 14 23 16 9.2 19 16 3.4 2.0 *= Difference between 8 hours and 0 hours and percent difference between 8 hours and 0 hours.
TABLE-US-00066 TABLE 65 Glucose concentration (mmol/L) of samples from 10 participants measured at 0 and 8 hours post-centrifugation stored at 21.degree. C. Greiner plasma PtPA % % B 0 hr 8 hr 8-0 hr* 8-0 hr* 0 hr 8 hr 8-0 hr* 8-0 hr* Mean 5.1 4.7 0.5 -9.1 5.2 5.1 0.08 -1.5 SD 0.8 0.9 0.2 5.1 0.8 0.8 0.04 0.9 *= Difference between 8 hours and 0 hours and percent difference between 8 hours and 0 hours.
[0681] In serum samples from the PtPA tubes, the LD and glucose results showed 0.6% and 1.5% change over the 8 hour period compared with the 12.5% and 9.1% change for the Greiner plasma tube samples respectively. In the plasma samples, the presence of cells resulted in glucose consumption, and the LD increased due to leakage from the cells lysed.
[0682] The results confirm that clotting of the blood with PtPA removes cells effectively and this prevents any significant changes in the most affected analytes, glucose and LD, for up to 8 hours. Thus, inclusion of PtPA in a serum tube provided high quality stable serum from the healthy participants' blood samples.
Example 11b: Cellular Contamination of Serum and Plasma Prepared from Anticoagulated Patients
[0683] Blood samples were collected from two participants (P1 and P2) undergoing cardiac surgery who had received in total: P1-30000, and P2-35000 units of heparin just prior to blood collection (within 15 minutes post heparin infusion). The samples were collected in 50 mL plain syringes (BD Plastipak REF #300866) filled with .about.30 mL blood. The syringes were delivered to the laboratory within 15 minutes of collection. The following tubes were filled with blood: Greiner plasma tube, Greiner serum tube, and Greiner serum tube (cleaned as per the procedure above) containing PtPA (1.2 .mu.g/4 mL of blood). The plasma and PtPA containing tubes were centrifuged immediately on arrival at the laboratory. The Greiner serum tube was allowed to stand for 60 minutes before centrifugation. The samples were analysed, then allowed to stand at room temperature (21.degree. C.) and re-analysed 24 hours later on the same analyser. Results are shown in Tables 66 and 67.
TABLE-US-00067 TABLE 66 LD activity (U/L) of samples from 2 cardiac surgery participants measured at 0 and 24 hours post-centrifugation stored at 21.degree. C. Greiner Plasma Greiner Serum PtPA Serum % % % Dfce Dfce Dfce Dfce Dfce Dfce A 0 hr 24 hrs (24-0 hrs) (24-0 hrs) 0 hr 24 hrs (24-0 hrs) (24-0 hrs) 0 hr 24 hrs (24-0 hrs) (24-0 hrs) Mean 87 94 8 +9 87 98 11 +13 84 81 -3 -3 SD 11 1 9 12 8 9 1 1 9 10 1 1
TABLE-US-00068 TABLE 67 Glucose concentration (mmol/L) of samples from 2 cardiac surgery participants measured at 0 and 24 hours post-centrifugation stored at 21.degree. C. Greiner Plasma Greiner Serum PtPA Serum % % % Dfce Dfce Dfce Dfce Dfce Dfce B 0 hr 24 hrs (24-0 hrs) (24-0 hrs) 0 hr 24 hrs (24-0 hrs) (24-0 hrs) 0 hr 24 hrs (24-0 hrs) (24-0 hrs) Mean 5.4 4.6 -0.8 -14.8 5.4 4.7 -0.8 -13.9 5.3 5.3 -0.1 -0.9 SD 0.1 0.1 0.0 0.4 0.1 0.1 0.1 1.0 0.1 0.1 0.1 1.3
[0684] In the PtPA-produced serum samples the LD and glucose results showed a change of -3 and -0.9% at 24 hours compared with the 9%, and -14.8% change in the lithium heparin plasma, and 13% and -13.9% for serum samples respectively. Similar to the healthy participant group of Examples 11a, the presence of cells in plasma resulted in glucose consumption and with cell lysis, the activity of LD was increased. The Greiner serum tubes showed plasma like results, as expected, since the samples never clotted.
[0685] In the Greiner serum tube no blood clotting was observed nor was any latent clotting detected visually or by analysers over the 24 hours post-centrifugation. The samples in the PtPA-containing tubes clotted within 3-5 minutes. The 3 tubes of participant P1 were photographed to indicate the presence of cells on top of the gel barrier. In the PtPA tube, very few cells were present above or within the gel barrier, in contrast to the other two tubes where cells were visible throughout and above the gel barrier.
[0686] From each of the 3 tubes the majority of the plasma or serum content was gently removed without disturbing the layer on top of the gel barrier, leaving about 0.5 mL of the plasma or serum in the tube. The residual plasma or serum in each tube was re-mixed with the buffy coat and transferred into a slide centrifuge, Cytospin (Shandon-Elliott Cytospin, Shandon-Elliott Instruments Limited) to concentrate the cells and produce a Giemsa-stained slide of the cells, cell stroma, etc for microscopic examination. The slides (shown in FIG. 56) clearly indicate presence of cells in abundance in the plasma, slightly fewer in the Greiner serum and minimal cells in the PtPA-produced serum.
[0687] These results confirm clotting of the blood with PtPA was achieved even in so called "fully heparinised" participants, in a very short time, <5 minutes. After centrifugation, the number of residual cells was minimal and insufficient to have an effect on LD activity or glucose concentrations.
Example 11c: Effect of Prolonged Storage of Serum and Plasma Samples from One Cardiac Surgery Participant on Analytes
[0688] The 3 samples from cardiac surgery participant P2 after 24 hours at room temperature (21.degree. C.) were stored in the primary tubes at 4.degree. C. for 13 additional days, (total of 14 days or 336 hours post collection) and then were re-analysed for K.sup.+, glucose, LD and phosphate (Pi). The results are shown in Table 68.
TABLE-US-00069 TABLE 68 Concentration of analytes in serum and plasma samples from a cardiac surgery participant measured at 0, 24 and 336 hours post-centrifugation 24 Dfce % Dfce Dfce % Dfce Analyte 0 hr hrs 336 hrs (24-0) (24-0) (336-0) (336-0) Greiner Plasma K.sup.+ 4.7 4.6 4.9 -01 -2.1 0.2 4.3 (mmol/L) Glucose 5.5 4.1 3.1 -1.4 -25.5 -2.4 -43.6 (mmol/L) LD (U/L) 79 93 208 14 17.7 129 163.0 Pi 0.59 0.6 0.69 0.01 1.7 0.1 16.9 (mmol/L) Greiner Serum K.sup.+ 4.7 4.7 5.2 0 0 0.5 10.6 (mmol/L) Glucose 5.5 4.7 3.8 -0.8 -14.5 -1.7 -30.9 (mmol/L) LD (U/L) 81 91 180 10 12.3 99 122.2 Pi 0.58 0.63 0.74 0.05 7.8 0.16 27.6 (mmol/L) PtPA Serum K.sup.+ 4.6 4.6 4.6 0 0 0 0 (mmol/L) Glucose 5.4 5.3 5.3 0.1 1.9 -0.1 -1.9 (mmol/L) LD (U/L) 77 74 64 3 3.9 -13 -20.3 Pi 0.57 0.58 0.65 -0.01 -1.8 0.08 12.3 (mmol/L)
[0689] Unlike the lithium heparin plasma and "serum" from the Greiner tubes, the PtPA serum sample showed insignificant changes in the H.sup.+ and glucose results, while the LD and Pi results showed minimal changes.
[0690] In this extreme case scenario, serum produced using PtPA provided easily the most stable sample type that will allow laboratories to provide additional analyte testing at later dates and have confidence the results will be accurate. The 20% fall in LD activity in the PtPA sample was probably due to slow denaturation of existing LD on storage.
[0691] Overall Conclusion
[0692] PtPA produced serum offers outstanding stability in analyte concentrations that are most likely to be affected by the incomplete removal of cells from the plasma or serum component above the gel barrier during the clotting and/or centrifugation process in all patients. Our results also show by direct observation the relative absence of cells from the PtPA produced sera compared with Greiner lithium heparin plasma prepared from healthy participants or patients on high doses of heparin.
Example 12: Measurement of Biochemical Analytes in Plasma and Serum Samples Prepared in Commercial Tubes and Serum Samples Prepared Using Venom Prothrombin Activators (Ecarin, PtPA, and OsPA)
[0693] As discussed in earlier parts of this specification, current commercially available blood collection tubes are unable to produce completely clotted serum from all blood samples in a timely manner to meet the quality and turn-around time expectations from biochemistry laboratories for optimal patient care. The results in the above examples demonstrate that prothrombin activators can be employed to produce a quality serum sample rapidly with low levels of fibrinogen/fdp/FDP and without cellular contaminations (as determined by visual inspection, clarity of the serum and analysis) in blood from a wide variety of patients/individuals.
[0694] It was important therefore to determine whether the prothrombin activators might interfere with analysis of analytes commonly used for clinical management of patients. Prothrombin activators are proteolytic enzymes which could in principle cleave serum proteins or proteins involved in analytical methodology, thereby affecting analytical results. Proteins are involved in analytical methods either as the analyte of interest or as reactants (e.g. enzymes, antibodies) used to measure the analytes of interest.
[0695] The purpose of this example was to investigate whether serum prepared using prothrombin activators gives the same analytical results as plasma and serum prepared in current commercially available tubes using current commercial methods.
[0696] It is not uncommon for more than 30 biochemical analytes to be requested and performed from a single serum or plasma tube. As the range of analytes increases and the analytical volumes per analyte decreases with technological improvements, the number of analyses from a single tube will further increase. Hence it is essential tube additives, specifically procoagulants, be inert and not impose any analytical effect on analytes yet provide highest quality sample for the most accurate estimation of analytes.
[0697] Standard analytical test procedures were used in the following experiments for each of the 33 assays which are listed below:
[0698] Test 1: Sodium (Na.sup.+)
[0699] Test 2: Potassium (K.sup.+)
[0700] Test 3: Chloride (Cl.sup.-)
[0701] Test 4: Bicarbonate (HCO.sub.3.sup.-)
[0702] Test 5: Glucose (Gluc)
[0703] Test 6: Urea (Urea)
[0704] Test 7: Creatinine (Creat)
[0705] Test 8: Urate (Urate)
[0706] Test 9: Total Protein (TP or T Prot)
[0707] Test 10: Albumin (Alb)
[0708] Test 11: Total Bilirubin (T Bili)
[0709] Test 12: Alkaline Phosphatase (ALP)
[0710] Test 13: Gamma-Glutamyl Transferase (GGT)
[0711] Test 14: Alanine Aminotransferase (ALT)
[0712] Test 15: Aspartate Aminotransferase (AST)
[0713] Test 16: Lactate Dehydrogenase (LD)
[0714] Test 17: Creatine Kinase (CK)
[0715] Test 18: Total Calcium (TCa)
[0716] Test 19: Phosphate (Pi or Phos)
[0717] Test 20: Magnesium (Mg.sup.2+)
[0718] Test 21: Lipase (Lipase)
[0719] Test 22: Cholesterol (Chol)
[0720] Test 23: Triglycerides*
[0721] Test 24: High-Density Lipoprotein Cholesterol (HDL-C or HDL)
[0722] Test 25: Iron (Fe.sup.2+)
[0723] Test 26: Transferrin (Trf)
[0724] Test 27: C Reactive Protein (CRP)
[0725] Test 28: Cortisol Test (Cortisol)
[0726] Test 29: Free Thyroxine (FT4)
[0727] Test 30: Thyroid Stimulating Hormone (TSH)
[0728] Test 31: Ferritin (Ferritin)
[0729] Test 32: Troponin (TnI)
[0730] Test 33: Haemolytic Index (Haem index)**
[0731] Test 34: Icteric Index*
[0732] Test 35: Lipemia Index*
[0733] *Triglycerides were not measured in these experiments because of the presence of glycerol in the PtPA preparation. Icteric and Lipaemic indices were also not determined in this study.
[0734] ** See also Example 10.
[0735] Test Analysis
[0736] Analysis was performed on Beckman DxC800 general chemistry analysers and a DxI800 immunoassay analyser (Beckman Coulter, Fullerton, Calif., USA). Samples were loaded on the same instruments at the same time and within 1-2 hours post-centrifugation, except where recurrent latent clotting was encountered.
[0737] The analytes tested plus semi-quantitative haemolysis levels are listed in the result tables. The upper limit of imprecision of the between-run coefficient of variations (CVs) from the two and three internal quality control concentrations for the 35 analytes tested on the Beckman DxC800 analysers the DxI800 analyser respectively are shown in Table 69. We also measured the activated partial thromboplastin time (aPTT) on an ACT.sup.TOPS. (Instrumentation Laboratory, Lexington Mass., USA) since it was difficult to ascertain the exact anticoagulant concentration in the participants' blood samples at the time of the specimen collection in the cardiac care unit and dialysis participants. In all cases, results were consistent with listed degree of anticoagulation.
[0738] Data Analysis
[0739] The results of each test were obtained and then data analysis was performed as follows. The mean and standard deviation (SD) were calculated for each test for each type of tube. The difference (actual and %) between each result pair for each test between the different tubes (e.g. PtPA tube result versus Greiner serum result type) was also calculated. The % difference was then compared with the between-run precision values obtained on the analysers (Table 69) to determine if there were analytically significant differences between the different serum tubes and the different serum tubes versus plasma tubes. The participant data was also separated into healthy participants and anticoagulated participants, and the same analysis performed. If a measurement was not obtained for an analyte in any of the three tubes (due to recurrent latent dotting leading to insufficient specimen, insufficient specimen collected, not requested on analyser, or insufficient reagent), the result was not included in the calculation accounting for the variability in the number of specimens analysed per assay.
TABLE-US-00070 TABLE 69 Quality Control (QC) imprecision on the Beckman DxC800 and DxI800 analysers (*DxI800) QC Level 1 QC Level 2 QC Level 3 Analyte Mean SD CV % Mean SD CV % Mean SD CV % Na.sup.+ 132 1.27 1.0 150 1.33 0.9 K.sup.+ 3.9 0.06 1.4 6.0 0.07 1.2 Cl- 85 1.17 1.4 100 1.19 1.2 HCO.sub.3.sup.- 17 0.66 3.8 29 0.85 2.9 Gluc 4.8 0.12 2.6 16.7 0.28 1.8 Urea 5.2 0.17 3.3 15.9 0.41 2.6 Creat 68 3.34 4.9 491 8.93 1.8 Urate 0.23 0.001 1.9 0.49 0.01 1.5 T Prot 41 0.73 1.8 67 1.07 1.6 Alb 26 0.37 1.4 40 0.54 1.3 T Bili 21 1.37 6.4 93 1.96 2.1 ALP 106 2.79 2.6 471 7.93 1.7 GGT 38.7 2.03 5.3 155 2.88 1.9 ALT 25 1.52 6.2 94 2.00 2.1 AST 34 1.35 3.9 201 2.39 1.2 LD 149 3.21 2.2 408 5.65 1.4 CK 139 2.53 1.8 459 5.8 1.3 TCa 2.07 0.03 1.6 2.86 0.04 1.3 Phos 1.00 0.02 1.9 2.86 0.04 1.5 Mg.sup.2+ 0.84 0.02 2.5 1.63 0.04 2.3 Lipase 29 2.39 8.2 57 4.05 7.1 Chol 3.0 0.05 1.7 6.5 0.15 2.3 Trig 1.0 0.04 3.6 2.0 0.05 2.7 HDL 1.1 0.04 3.5 1.9 0.05 2.6 Fe.sup.2+ 10 0.29 2.8 34 1.93 5.7 Trf 1.0 0.04 3.5 2.3 0.08 3.5 CRP 4.6 0.36 7.9 10.0 0.55 5.4 TnI* 0.052 0.01 16.1 0.55 0.04 7.9 10.2 0.69 6.7 Cortisol* 139 8.85 6.4 552 28.48 5.2 906 41.7 4.6 fT4* 7.51 0.62 8.2 23.93 0.94 4.0 50.9 1.92 3.8 TSH* 0.71 0.05 7.1 4.17 0.26 6.3 23.1 1.53 6.6 Ferritin* 19 1.68 8.8 159 9.67 6.1 358 22.42 6.3
Example 12a: Comparison of PtPA Serum with Serum and Lithium Heparin Plasma Prepared in Commercial Tubes
[0740] A total of 61 participants were recruited. All 61 participants were adults, >18 years of age, with a mix of males and females. The participants consisted of two groups: 26 healthy volunteers and 35 anticoagulated patients.
[0741] Of the 35 anticoagulated patients, 1 was an outpatient on low dose warfarin therapy and 34 were inpatients.
[0742] Of the 34 inpatients, 11 were undergoing cardiac surgery, 8 were cardiac care unit patients, and 15 were on dialysis.
[0743] The 11 inpatients undergoing cardiac surgery had received in total 25,000-41,000 units of heparin at the time of the blood collection which was within 30 minutes post heparin infusion. The specimens were collected while participants were on bypass (where blood is pumped by machine not by the heart).
[0744] The 8 cardiac care unit participants were recruited the night before specimen collection and were receiving heparin by IV infusion, 950-1450 units of heparin per hour. Seven remained on IV heparin infusion (blood collection was .gtoreq.12 hours after IV heparin infusion started) at the time of specimens being collected, and the other was due for surgery on the day so had had the infusion stopped approximately three hours prior to the specimen collection (blood collection was >9 hours after IV infusion had started and .about.3 hours after IV heparin infusion was stopped). From the information in the patient records the heparin concentration in the infusate and the infusion rate were unchanged for the participants over the period between when IV heparin infusion started and blood was collected.
[0745] Of the 15 inpatients on dialysis, 12 were on heparin IV infusion, and 1 was on warfarin/heparin (.about.1750-7000 units of heparin, initial bolus plus hourly top up dose), and 2 were on clexane. Blood from these patients was collected at least one hour after dialysis commenced.
[0746] Blood was drawn using a standardised draw order. The blood was drawn by venipuncture from healthy volunteers, via bypass port from cardiac patients, and via bloodline from dialysis patients. The following tubes were used:
[0747] Greiner Vacuette.TM. citrate tube (for coagulation studies);
[0748] Greiner Vacuette.TM. No Additive tube with PtPA added (1.2 .mu.g/4 mL) (PtPA);
[0749] Greiner Vacuette.TM. plasma tube (GLH);
[0750] BD Vacutainer.TM. plasma/PST II tube (BDLH);
[0751] Greiner Vacuette.TM. serum tube (GRS);
[0752] BD serum tube/SST II (BDS); and
[0753] BD RST tube (BD RST).
[0754] The blood for the PtPA-containing tube was collected in a plain syringe and then transferred from the syringe into the Greiner tube containing the PtPA without a needle to minimise cell lysis.
[0755] The Greiner serum and BD SST tubes were allowed to clot for the standard time of 30 minutes for healthy participants and 60 minutes for anticoagulated participants. These tubes were visually inspected for clot formation prior to loading in the centrifuge.
[0756] The BD RST and PtPA tubes were visually inspected for clot formation at 5 minutes for all participants at the point of collection (phlebotomy or clinical unit). The BD RST specimens for the healthy participants and for the anticoagulated participants that formed a solid clot at 5 minutes were centrifuged as soon as delivered to the laboratory (<20 minutes). If clotting was incomplete the specimens were re-checked every 10-15 minutes for clotting and left to clot for 60 minutes maximum. The lithium heparin and PtPA tubes were centrifuged immediately on delivery in the laboratory (<20 minutes from collection).
[0757] All tubes were centrifuged at 3000 g for 10 minutes at 20.degree. C. in swing bucket centrifuges, and then stored at .about.21.degree. C. The tubes were visually inspected for latent clotting immediately after centrifugation and again just prior to loading onto analysers. These tube (primary tubes) were used for analysis except in cases where latent clotting was observed. In tubes where latent clotting was observed, the serum in the primary tubes was transferred to an aliquot tube, re-centrifuged to remove the clots, and the clean serum transferred to another aliquot tube (secondary tube) which was used for analysis. The Greiner No Additive tube does not contain a serum gel barrier, thus the PtPA generated serum was also transferred to a secondary tube for analysis to prevent re-mixing and prolonged contact with the cells.
[0758] Results
[0759] The results are shown in Table 70. Also, an example of results is provided in FIG. 57 for the measurement of total protein (TP). The left graph shows the mean with standard deviation value of all normal patient results (n=26) for each tube type. The centre graph shows the mean value of all patient results (n=61) for each tube type. The right graph shows the mean value of results for cardiac patients (n=11) for each tube type. The units on the y-axis are g/L protein. The error bars are standard deviation.
[0760] Comparison of PtPA sera with commercial tube sera from healthy and anticoagulated participants. No significant differences were observed with any of the analytes. This conclusion is based on comparison of the mean test values in Table 70 and a paired-wise statistical analysis described above.
[0761] Comparison of PtPA sera with commercial tube lithium heparin plasma from healthy and anticoagulated participants. Significant differences were observed in some analytes (K.sup.+, TP, AST and Pi) as expected because of the well established differences between plasma and serum. Sera prepared in the commercially available serum tubes showed similar analytical differences from plasma to those shown with PtPA sera.
TABLE-US-00071 TABLE 70 Analytical data for 32 analytes on serum and plasma samples. Number of Mean .+-. SD samples Analyte Units PtPA GLH BDLH GS BDS BD RST 60 Na.sup.+ mmol/L 136.9 .+-. 2.24 137.4 .+-. 2.44 137.2 .+-. 2.38 137.5 .+-. 2.29 137.5 .+-. 2.34 137.4 .+-. 2.38 61 K.sup.+ mmol/L 4.09 .+-. 0.59 3.91 .+-. 0.63 3.93 .+-. 0.65 4.13 .+-. 0.62 4.15 .+-. 0.60 4.07 .+-. 0.60 61 Cl.sup.- mmol/L 104.1 .+-. 3.8 104.6 .+-. 4.2 104.8 .+-. 4.1 104.6 .+-. 4.0 104.8 .+-. 3.8 104.7 .+-. 3.9 61 HCO.sub.3.sup.- mmol/L 24.8 .+-. 2.8 25.8 .+-. 3.0 25.3 .+-. 2.7 25.7 .+-. 3.1 25.1 .+-. 2.9 25.0 .+-. 3.0 61 Gluc mmol/L 6.2 .+-. 2.09 6.3 .+-. 2.15 6.3 .+-. 2.11 6.2 .+-. 2.07 6.2 .+-. 2.11 6.3 .+-. 2.08 61 Urea mmol/L 5.3 .+-. 2.0 5.3 .+-. 2.0 5.4 .+-. 2.0 5.3 .+-. 2.1 5.4 .+-. 2.1 5.3 .+-. 2.1 61 Creat .mu.mol/L 134.1 .+-. 112.8 135.1 .+-. 115.0 136.3 .+-. 114.7 137.1 .+-. 116.0 136.7 .+-. 116.0 138.1 .+-. 116.6 60 Urate mmol/L 0.27 .+-. 0.11 0.27 .+-. 0.12 0.27 .+-. 0.12 0.28 .+-. 0.11 0.28 .+-. 0.11 0.27 .+-. 0.12 61 TP g/L 62.6 .+-. 13.0 65.0 .+-. 13.2 65.5 .+-. 13.3 63.0 .+-. 12.6 63.0 .+-. 12.5 62.8 .+-. 12.6 61 Alb g/L 36.1 .+-. 8.3 35.9 .+-. 8.1 35.9 .+-. 8.1 36.1 .+-. 8.3 36.1 .+-. 8.3 36.2 .+-. 8.3 58 T Bili .mu.mol/L 13.5 .+-. 3.8 13.6 .+-. 3.7 13.4 .+-. 3.4 13.3 .+-. 3.6 13.4 .+-. 4.0 13.2 .+-. 3.8 61 ALP U/L 74.1 .+-. 26.9 68.9 .+-. 26.5 69.4 .+-. 26.6 71.3 .+-. 26.9 71.7 .+-. 27.5 71.4 .+-. 26.9 61 GGT U/L 27.9 .+-. 17.6 28.3 .+-. 17.0 28.1 .+-. 16.9 27.3 .+-. 17.1 28.1 .+-. 17.1 27.8 .+-. 17.5 61 .sup..sctn.ALT U/L 30.4 .+-. 19.9 29.6 .+-. 18.8 29.6 .+-. 19.1 30.4 .+-. 19.1 29.8 .+-. 19.2 30.7 .+-. 20.1 59 .sup..sctn.AST U/L 34.2 .+-. 37.7 32.5 .+-. 37.5 32.6 .+-. 36.9 32.0 .+-. 34.9 32.6 .+-. 38.7 32.4 .+-. 39.1 58 .sup..sctn.LD U/L 212.9 .+-. 110.0 215.4 .+-. 113.1 215.3 .+-. 110.7 213.8 .+-. 106.0 222.8 .+-. 120.0 212.5 .+-. 119.2 59 .sup..sctn.CK U/L 147.1 .+-. 250.3 146.6 .+-. 250.0 145.3 .+-. 242.8 146.0 .+-. 241.3 149.4 .+-. 259.6 147.7 .+-. 253.3 61 TCa mmol/L 2.19 .+-. 0.26 2.17 .+-. 0.24 2.17 .+-. 0.24 2.20 .+-. 0.26 2.20 .+-. 0.26 2.19 .+-. 0.26 61 Pi mmol/L 1.07 .+-. 0.28 1.02 .+-. 0.28 1.02 .+-. 0.28 1.08 .+-. 0.30 1.08 .+-. 0.30 1.08 .+-. 0.30 57 Mg.sup.2+ mmol/L 0.94 .+-. 0.19 0.95 .+-. 0.20 0.94 0.94 .+-. 0.18 0.94 .+-. 0.20 0.93 .+-. 0.20 0.19 59 Lipase U/L 28.2 .+-. 9.0 28.3 .+-. 9.0 28.3 .+-. 8.9 28.2 .+-. 9.0 28.6 .+-. 9.2 28.2 .+-. 9.0 57 Chol mmol/L 4.2 .+-. 1.6 4.2 .+-. 1.6 4.2 .+-. 1.6 4.3 .+-. 1.6 4.3 .+-. 1.6 4.3 .+-. 1.6 58 HDL-C mmol/L 1.17 .+-. 0.6 1.25 .+-. 0.58 1.26 .+-. 0.60 1.20 .+-. 0.58 1.21 .+-. 0.58 1.21 .+-. 0.59 59 Fe.sup.2+ .mu.mol/L 14.6 .+-. 6.6 14.5 .+-. 6.5 14.6 .+-. 6.7 14.7 .+-. 6.7 14.9 .+-. 6.7 14.6 .+-. 6.6 59 Trf g/L 2.17 .+-. 0.74 2.13 .+-. 0.71 2.14 .+-. 0.72 2.17 .+-. 0.73 2.19 .+-. 0.73 2.18 .+-. 0.77 57 CRP .mu.g/L 14.4 .+-. 49.3 14.5 .+-. 49.5 14.6 .+-. 50.5 14.2 .+-. 46.8 15.0 .+-. 52.3 15.1 .+-. 52.4 59 Cortisol* nmol/L 356 .+-. 247 354 .+-. 241 356 .+-. 254 356 .+-. 249 352 .+-. 248 358 .+-. 250 60 FT4* Pmol/L 13.6 .+-. 4.1 12.7 .+-. 3.7 12.7 .+-. 3.6 12.7 .+-. 3.9 12.7 .+-. 3.9 12.8 .+-. 3.7 61 TSH* .mu.IU/mL 1.81 .+-. 2.12 1.87 .+-. 2.05 1.89 .+-. 2.12 1.83 .+-. 2.10 1.82 .+-. 2.02 1.86 .+-. 2.08 43 Ferritin* .mu.g/L 204 .+-. 191 199 .+-. 191 202 .+-. 192 208 .+-. 203 207 .+-. 198 204 .+-. 187 60 .sup..sctn.TnI* .mu.g/L 0.873 .+-. 5.33 0.994 .+-. 6.06 0.945 .+-. 5.75 0.782 .+-. 4.77 0.869 .+-. 5.25 0.951 .+-. 5.75 61 Haem 0.2 .+-. 0.7 0.2 .+-. 0.7 0.3 .+-. 0.7 0.3 .+-. 0.7 0.2 .+-. 0.7 0.2 .+-. 0.7 Index *Analysis performed on a DxI800 immunoassay analyser. .sup..sctn.signifies non-parametric distribution used. The p value was determined by Wilcoxen Matched-Paris Ranks test.
Example 12b: Comparison of PtPA, OsPA and Ecarin Serum with Serum and Plasma Prepared in Commercial Tubes
[0762] 7 participants were recruited consisting of 5 healthy volunteers with a mix of 4 males and 1 female, and 2 patients undergoing cardiac surgery (all adults >18 years of age). The two cardiac patients had received 34,000 and 43,000 units of heparin within 30 minutes post heparin infusion.
[0763] Blood from the healthy participants was drawn by venipuncture using standardized draw order: Greiner citrate plasma tube; Greiner serum tube; Greiner plasma tube; a plain syringe (Thermo 10 mL # CE0197) for the PtPA, OsPA or ecarin containing Greiner No Additive tubes (the concentration of the prothrombin activators in these 4 mL tubes was PtPA 1.2 .mu.g, OsPA 0.5 .mu.g and ecarin 0.625 units for healthy and 1.25 units for cardiac surgery participants). The blood from the syringe was transferred into the Greiner No Additive tubes containing the prothrombin activators without a needle to minimise cell lysis.
[0764] For the cardiac participants blood was collected in a plain syringe (30 mL) while on bypass. The blood was delivered to the laboratory within 15 minutes and dispensed into the various tubes listed above.
[0765] The Greiner serum tubes were allowed to clot for the standard time of 30 minutes for healthy participants and 60 minutes for the cardiac surgery participants, and then visually inspected prior to loading in the centrifuge for clot formation. The PtPA, OsPA and ecarin specimens were visually inspected for clot formation at 3 and 5 minute mark for all participants at the point of collection (phlebotomy for healthy participants and in the clinical unit for the cardiac surgery participants). The lithium heparin, PtPA, OsPA and ecarin tubes were centrifuged immediately on delivery in the laboratory (<30 minutes from collection).
[0766] All tubes were centrifuged at 3000 g for 10 minutes at 20.degree. C. in a swing bucket centrifuge, and then stored at .about.21.degree. C. The tubes were visually inspected for latent clotting immediately after centrifugation and again just prior to loading onto analysers. The Greiner primary tubes were used for analysis. The Greiner No Additive tube does not contain a serum gel barrier, thus the serum generated using prothrombin activators was transferred to aliquot (secondary) tubes to prevent re-mixing and prolonged contact with the cells.
[0767] Results
[0768] Results are shown in Table 71 and support the following conclusions:
[0769] 1. Comparison of sera produced by the prothrombin activators versus Greiner sera--no significant differences were observed with any of the analytes.
[0770] 2. Comparison of sera produced by prothrombin activators versus Greiner lithium heparin plasma--significant differences were observed in some analytes (K.sup.+, TP, AST and Pi) as expected because of the well established differences between plasma and serum. Sera prepared in commercial serum tubes showed similar analytical differences from plasma to those shown with prothrombin activator sera.
TABLE-US-00072 TABLE 71 Data for each analyte from the three different snake procoagulants and the Greiner tubes. No of Mean .+-. SD Samples Analyte Units GLH GS PtPA OsPA Ecarin 7 Na.sup.+ mmol/L 138.1 .+-. 1.22 138.1 .+-. 1.46 137.3 .+-. 1.25 136.7 .+-. 2.36 137.1 .+-. 1.57 7 K.sup.+ mmol/L 4.40 .+-. 0.82 4.61 .+-. 0.71 4.53 .+-. 00.77 4.50 .+-. 0.76 4.49 .+-. 0.79 7 Cl.sup.- mmol/L 104.0 .+-. 2.8 104.0 .+-. 2.6 103.0 .+-. 2.6 102.9 .+-. 2.34 103.4 .+-. 2.4 7 HCO.sub.3.sup.- mmol/L 25.1 .+-. 3.2 25.3 .+-. 3.5 23.9 .+-. 2.4 24.1 .+-. 2.7 23.3 .+-. 2.2 7 Gluc mmol/L 5.76 .+-. 0.88 5.69 .+-. 0.94 5.63 .+-. 0.0.81 5.60 .+-. 0.80 5.64 .+-. 0.0.90 7 Urea mmol/L 4.63 .+-. 0.82 4.66 .+-. 0.80 4.64 .+-. 0.85 4.54 .+-. 0.83 4.54 .+-. 0.88 7 Creat .mu.mol/L 75.1 .+-. 10.9 76.7 .+-. 12.35 75.9 .+-. 12.10 76.29 .+-. 12.54 75.0 .+-. 12.7 7 Urate mmol/L 0.33 .+-. 0.08 0.33 .+-. 0.08 0.33 .+-. 0.08 0.32 .+-. 0.07 0.32 .+-. 0.08 7 TP g/L 68.5 .+-. 11.0 62.7 .+-. 11.6 63.9 .+-. 11.8 62.4 .+-. 14.1 63.4 .+-. 13.4 7 Alb g/L 38.3 .+-. 5.5 38.3 .+-. 5.5 38.6 .+-. 5.8 38.4 .+-. 6.0 38.6 .+-. 5.8 7 T Bili .mu.mol/L 15.6 .+-. 4.6 16.6 .+-. 4.7 16.4 .+-. 4.5 16.1 .+-. 4.6 15.7 .+-. 5.2 7 ALP U/L 65.7 .+-. 18.6 66.0 .+-. 18.4 66.1 .+-. 20.7 69.3 .+-. 16.7 67.9 .+-. 18.7 7 GGT U/L 27.4 .+-. 15.2 25.1 .+-. 15.0 25.4 .+-. 14.7 25.7 .+-. 15.8 24.9 .+-. 15.8 7 .sup..sctn.ALT U/L 30.4 .+-. 11.4 31.0 .+-. 11.8 32.0 .+-. 11.4 31.7 .+-. 12.2 32.1 .+-. 11.8 7 .sup..sctn.AST1 U/L 22.6 .+-. 7.8 25.0 .+-. 8.3 24.1 .+-. 7.2 24.9 .+-. 6.5 24.9 .+-. 6.7 7 .sup..sctn.LD U/L 194.4 .+-. 23.1 193.6 .+-. 41.6 195.6 .+-. 22.4 189.0 .+-. 18.9 192.7 .+-. 21.2 7 .sup..sctn.CK U/L 98.9 .+-. 34.9 97.9 .+-. 37.5 98.6 .+-. 36.8 97.1 .+-. 34.0 98.0 .+-. 36.1 7 TCa mmol/L 2.20 .+-. 0.236 2.23 .+-. 0.245 2.22 .+-. 0.255 2.20 .+-. 0260 2.21 .+-. 0.25 7 Pi mmol/L 1.13 .+-. 0.13 1.18 .+-. 0.14 1.19 .+-. 0.13 1.16 .+-. 0.14 1.17 .+-. 0.13 7 Mg.sup.2+ mmol/L 1.04 .+-. 0.23 1.06 .+-. 0.22 1.03 .+-. 0.20 1.03 .+-. 0.19 1.04 .+-. 0.21 7 Chol mmol/L 4.3 .+-. 1.7 4.3 .+-. 1.7 4.3 .+-. 1.7 4.3 .+-. 1.7 4.3 .+-. 1.7 7 HDL-C mmol/L 1.07 .+-. 0.27 1.04 .+-. 0.23 1.01 .+-. 0.27 1.03 .+-. 0.28 1.01 .+-. 0.06 7 Fe.sup.2+ .mu.mol/L 16.1 .+-. 5.8 16.3 .+-. 5.9 16.4 .+-. 5.8 16.4 .+-. 5.7 16.1 .+-. 5.8 7 Trf g/L 2.18 .+-. 0.51 2.18 .+-. 0.51 2.21 .+-. 0.54 2.20 .+-. 0.53 2.21 .+-. 0.55 7 CRP .mu.g/L 1.7 .+-. 1.5 1.9 .+-. 1.6 1.7 .+-. 1.5 1.9 .+-. 1.6 1.7 .+-. 1.5 7 Cortisol* nmol/L 382.1 .+-. 251.4 379.7 .+-. 247.2 373.6 .+-. 256.9 366.3 .+-. 241.2 359.3 .+-. 237.7 7 FT4* Pmol/L 13.53 .+-. 2.63 13.56 .+-. 3.26 13.46 .+-. 1.2.3 13.76 .+-. 3.17 14.23 .+-. 3.13 7 FT3* Pmol/L 4.83 .+-. 0.35 4.66 .+-. 0.43 4.91 .+-. 0.51 4.74 .+-. 0.35 4.97 .+-. 0.48 7 TSH* .mu.IU/mL 1.86 .+-. 0.48 1.88 .+-. 0.56 1.72 .+-. 0.30 1.75 .+-. 0.30 1.73 .+-. 0.32 7 Ferritin* .mu.g/L 236.6 .+-. 297.2 234.9 .+-. 304.8 226.4 .+-. 280.8 207.3 .+-. 236.5 225.3 .+-. 280.1 7 .sup..sctn.TnI* .mu.g/L 0.079 .+-. 0.136 0.071 .+-. 0.137 0.091 .+-. 0.165 0.092 .+-. 0.170 0.093 .+-. 0.174 7 Haem 0.4 .+-. 0.5 0.6 .+-. 0.8 0.6 .+-. 0.5 0.4 .+-. 0.5 0.4 .+-. 0.5 Index .sup..sctn.signifies non-parametric distribution used. The p value was determined by Wilcoxen Matched-Paris Ranks test. *Analysis performed on a DxI800 immunoassay analyser.
[0771] Summary of Example 12
[0772] The blood of all the healthy and anticoagulated participants ranging from the lowest dose to the highest "fully" heparinised participant clotted within 5 minutes in the PtPA, OsPA and ecarin-containing tubes to give firm immobile clots. No latent clotting was visually observed or detected by the analysers in any of the sera produced by the prothrombin activators. Despite the prothrombin activators being proteolytic enzymes, they did not produce any analytically or clinically significant effect on any of the analytes measured irrespective of whether the analytes were proteins or whether proteins were used as reactive compounds in the analytical methods. There were no significant differences observed in any of the analytes between the sera prepared in the commercial tube and the sera prepared using prothrombin activators. The analytical and clinical differences observed in analytes between lithium heparin plasma prepared in a commercial tube and sera produced using prothrombin activators were in line with published data.
Example 13: Further Measurement of Biochemical Analytes in Plasma and Serum Samples Prepared in Commercial Tubes and Serum Samples Prepared Using Venom Prothrombin Activators (Notecarin and Carinactivase-2)
[0773] This Example follows from Example 12 and uses the same methodology described therein.
Example 13a: Analyte Measurement
[0774] Blood from 5 healthy participants was collected into Greiner No Additive tubes (Cat No 454001) to which 25 .mu.L of notecarin or carinactivase-2 had been added to give concentrations of 12 nmol/mL and 45 nmol/mL respectively. Greiner Vacuette serum tubes (Cat No 456078; GS) were used as controls. The GS tubes were allowed to clot for 30 minutes as per manufacturer's recommendation prior to centrifugation. The tubes containing the prothrombin activators were observed for clotting immediately upon addition of the blood.
[0775] Notecarin and carinactivase-2 tubes clotted within 2 minutes and the tubes were brought to the laboratory and centrifuged within 7 to 15 minutes of collection. The clots formed in the tubes containing the activators were solid, and no latent clot formation was observed or detected.
[0776] The samples were then analysed for 31 analytes, and the results are shown in Table 72. For 27 analytes, the values obtained were equal within experimental error, with differences between paired results less than the least significant change (LSC %). The results where the differences were greater than the stated LSC (highlighted) can be explained as follows:
[0777] (1) For LD, the slightly lower activities in the two prothrombin activator samples may reflect decreased cellular contamination of these compared to Greiner serum;
[0778] (2) The high triglyceride (Trig) level in the Notecarin sample is due to interference by glycerol in which the Notecarin was stored;
[0779] (3) Notecarin serum produced higher AST levels, however the differences were clinically insignificant;
[0780] (4) Troponin results for the five patients were too low for accurate differentiation.
TABLE-US-00073 TABLE 72 Analytical results for serum samples generated by notecarin and carinactivase-2. CA2-GS Notec-GS Mean Mean dffce dffce No of Mean .+-. SD between between LSC CAL Samples Analyte Units GS Notecarin CA-2 pairs (%) pairs (%) (%) (%) 5 Na.sup.+ mmol/L 138.6 .+-. 1.5 138 .+-. 1.6 138 .+-. 1.5 -0.4 -0.4 2.8 3 5 K.sup.+ mmol/L 3.94 .+-. 0.17 3.94 .+-. 0.18 3.83 .+-. 0.18 0 -2.9 3.3 5 5 Cl.sup.- mmol/L 104.2 .+-. 2.4 104.2 .+-. 2.7 106.5 .+-. 2.6 0 2.2 3.3 5 5 HCO.sub.3.sup.- mmol/L 26.8 .+-. 2.6 25.6 .+-. 2.5 25.6 .+-. 2.2 -4.5 -4.5 4.8 10 5 Gluc mmol/L 4.86 .+-. 0.86 4.76 .+-. 0.82 4.74 .+-. 0.65 -2.1 -2.4 4.4 10 5 Urea mmol/L 5.58 .+-. 2.49 5.56 .+-. 2.44 5.55 .+-. 2.42 -0.4 -0.5 5.0 10 5 Creat .mu.mol/L 69.2 .+-. 25.0 69.2 .+-. 22.9 67.2 .+-. 22.9 -2.9 -2.9 5.5 10 5 Urate mmol/L 0.242 .+-. 0.026 0.240 .+-. 0.026 0.240 .+-. 0.023 -0.8 -1.2 3.8 10 5 TP g/L 66.0 .+-. 2.0 65.0 .+-. 2.2 63.6 .+-. 2.4 -1.5 -3.3 3.5 5 5 Alb g/L 40.8 .+-. 1.9 40.2 .+-. 1.9 39.5 .+-. 1.4 -1.5 -3.1 3.2 5 5 T Bili .mu.mol/L 11.4 .+-. 8.8 11.6 .+-. 9.0 10.6 .+-. 6.9 1.8 -7.0 7.0 10 5 ALP U/L 60.0 .+-. 4.1 60.4 .+-. 4.2 58.4 .+-. 4.3 0.7 -2.6 4.8 10 5 GGT U/L 14.6 .+-. 4.9 14.4 .+-. 6.5 14.8 .+-. 5.0 -1.4 1.2 7.3 10 5 ALT U/L 22.6 .+-. 8.0 23.2 .+-. 7.6 23.9 .+-. 7.6 2.7 5.8 6.8 10 5 AST U/L 17.2 .+-. 3.8 19.8 .+-. 4.4 17.3 .+-. 3.5 .sup. 15.1.sup..sctn. 0.4 6.2 10 5 LD U/L 192.2 .+-. 17.0 184.8 .+-. 18.3 179.9 .+-. 17.6 .sup. -3.9.sup..sctn. -6.3.sup..sctn. 3.7 15 5 CK U/L 81.4 .+-. 15.9 81.0 .+-. 15.4 78.8 .+-. 14.1 -0.5 -3.2 3.7 15 5 TCa mmol/L 2.330 .+-. 0.07 2.290 .+-. 0.07 2.267 .+-. 0.07 -1.7 -2.7 3.5 5 5 Pi mmol/L 1.314 .+-. 0.118 1.282 .+-. 0.117 1.271 .+-. 0.099 -2.4 -3.3 3.8 10 5 Mg.sup.2+ mmol/L 0.804 .+-. 0.046 0.802 .+-. 0.051 0.792 .+-. 0.054 -0.3 -1.4 4.4 10 5 Chol mmol/L 4.84 .+-. 0.35 4.78 .+-. 0.39 4.70 .+-. 0.34 -1.2 -2.9 3.6 10 5 Trig mmol/L 1.52 .+-. 0.55 5.61 .+-. 0.96 1.56 .+-. 0.65 269#.sup..sctn. 2.6 5.3 10 5 HDL-C mmol/L 1.420 .+-. 0.399 1.424 .+-. 0.293 1.388 .+-. 0.277 0.3 -2.6 5.2 10 5 Fe.sup.2+ .mu.mol/L 17.2 .+-. 3.8 16.8 .+-. 4.1 16.6 .+-. 3.6 -2.3 -3.3 3.7 10 5 Trf g/L 2.596 .+-. 0.187 2.566 .+-. 0.188 2.508 .+-. 0.188 -1.16 -3.4 4.8 10 5 Cortisol* nmol/L 236.0 .+-. 27.2 244.4 .+-. 21.5 231.9 .+-. 297.7 3.6 -1.7 6.4 15 5 FT4* Pmol/L 10.54 .+-. 1.45 11.38 .+-. 2.05 11.08 .+-. 1.05 -2.7 5.1 6.5 30 5 TSH* .mu.IU/mL 1.142 .+-. 0.783 1.142 .+-. 0.783 1.080 .+-. 0.783 1.1 -4.5 7.3 15 5 Ferritin* .mu.g/L 43.6 .+-. 20 43.6 .+-. 17.5 14.2 .+-. 17.7 0.0 1.4 7.2 15 5 TnI* .mu.g/L 0.009 .+-. 0.003 0.010 .+-. 0.005 0.011 .+-. 0.002 11.6 25.6 14 30 5 Haem 0 .+-. 0 0 .+-. 0 0 .+-. 0 0 0 Index Analysis performed on Beckman DxC800 and *DxI800 immunoassay analysers. .sup..sctn.= Those results which differ by more than the LSC and/or CAL.
Example 13b: Analyte Stability
[0781] Greiner Serum tubes were washed (x5 with distilled water) to remove the surfactant and procoagulant but retain the gel barrier then oven dried. Notecarin and carinactivase-2 (25 .mu.L aliquots) were added to the tubes to give concentrations of 12 nmol/mL and 45 nmol/mL respectively. Blood (5 mL) was collected from a healthy volunteer into one tube containing notecarin and one containing carinactivase-2. The tubes were centrifuged after 5 minutes. K.sup.+, glucose, LD and phosphate levels were determined at 3 time intervals after serum preparation. The samples were stored at 23.degree. C. for 5 hours then from 5 to 26.5 hours at 4.degree. C. The level of haemoglobin was also measured.
[0782] The results are shown in Table 73.
TABLE-US-00074 TABLE 73 Stability of analytes in serum samples generated by Notecarin and Carinactivase-2 Analyte Zero time 5 hrs 26.5 hrs Serum Hb mg/L Notecarin 32 K.sup.+ 4.0 4.1 4.0 Glucose 5.4 5.3 5.1 LD 176 175 173 Phosphate 1.05 1.13 1.05 Carinactivase 2 46 K.sup.+ 3.9 3.9 3.9 Glucose 5.3 5.0 4.9 LD 176 177 170 Phosphate 1.03 1.04 1.06
[0783] A haemoglobin level of <50 mg/L is considered clinically insignificant. These results therefore indicate that the serum samples produced by the prothrombin activators were effectively free of haemoglobin (therefore there is very little haemolysis during the dotting process), and free of cellular contamination. If cells or cell debris had been present, the concentration of K.sup.+, LD and phosphate would be expected to increase and the concentration of glucose to decrease on storage. Samples of the sera were taken and analysed for cells using the Cytospin method. Very few cells were detected.
[0784] In summary, serum samples prepared by using a group B prothrombin activator (carinactivase-2) and a group D prothrombin activator (notecarin) were of high quality based on completeness of clotting, analytical results and absence of cellular contaminants.
Example 14: Measurement of Biochemical Analytes in Plasma and Serum Samples Prepared in Commercial Tubes and Serum Samples Prepared Using Prothrombin Activator-Containing Venoms
[0785] This Example uses the same analyte analysis methodology described in Examples 12 and 13.
[0786] The discoveries made in the preceding examples surprisingly demonstrated that snake venom prothrombin activators would be suitable for use as procoagulants in clinical environments. This then raised the question whether the crude venoms which contain these prothrombin activators could be used in blood clotting devices without prior purification of the prothrombin activators.
[0787] The following experiments were therefore designed and conducted to show whether the venoms could be used to rapidly produce high quality serum from human blood samples. The amount of venom used was approximately four times the amount of purified prothrombin activators used in the preceding experiments, reflecting the fact that the venoms contain other proteins than the prothrombin activators.
Example 14a: Use of Crude P. Textilis and O. Scutellatus Venoms as Procoagulants in Clotting Tubes
[0788] Blood was collected from 2 healthy volunteers into 10 Greiner No Additive tubes (#454001, Griener Bio-One, Kremsmuster, Austraia) (4 mL capacity) containing 2 and 4 .mu.g (in duplicate for the 4 .mu.g O. scutellatus venom) of either P. textilis or O. scutellatus venom. Blood samples were also collected into Greiner standard serum tubes (#456071,Greiner Bio-One, Kremsmuster, Austraia) for comparison.
[0789] Clotting in the venom-containing tubes appeared complete by visual observation in 2 minutes. Samples in the venom-containing tubes were centrifuged after 15 minutes and those in the Greiner serum tubes after 30 minutes. Samples were centrifuged at 3000 g, 10 min and 20.degree. C. (Hereaus 1 S-R centrifuge, Germany). The sera from the Greiner No Additive tubes were immediately transferred into Beckman plain plastic tubes (#448778, Beckman coulter, Brea, Calif., USA) for observation and analysis. Latent clotting was observed in about half (6 out of the 10) of the venom-containing tubes but not in either of the Greiner serum tubes. This indicates that in some tubes, clotting was incomplete at the concentrations of venom used. Samples in the plain plastic tubes were re-centrifuged to remove cells, cell stroma and latent clots and the clear sera obtained analysed within 2 hours for 31 analytes. Sample analysis was performed within two hour post centrifugation for 31 analytes on the Beckman DxC800 general chemistry analyser and DxI800 immuno-analyser (Beckman Coulter, Brea, Calif., USA). There were no clinically significant differences at the concentration levels tested between results obtained for the 12 sera (4 P. textilis sera and 6 0. scutellatus sera and the 2 Greiner sera).
[0790] The results are shown in Table 74.
TABLE-US-00075 TABLE 74 Analytical results for serum samples generated by P. textilis sera and O. scutellatus venom. Mean .+-. SD Mean difference between pairs (%) No of Brown Brown Taipan Taipan Taipan Br2- Samples Analyte Units GS 2.0 .mu.g 4.0 .mu.g 2.0 .mu.g 4.0 .mu.g 4.0 .mu.g GS BR4-GS T2-GS T4-GS T4R-GS LSC (%) CAL (%) 2 Na.sup.+ mmol/L 136.5 .+-. 0.71 137.0 .+-. 1.41 136.5 .+-. 0.71 137.0 .+-. 1.41 136 .+-. 0.00 136.0 .+-. 0.00 -0.4 0.0 0.4 -0.4 -0.4 2.8 3 2 K.sup.+ mmol/L 4.05 .+-. 0.21 4.05 .+-. 0.07 4.00 .+-. 0.00 4.20 .+-. 0.28 4.00 .+-. 0.00 4.25 .+-. 0.35 0.0 -1.2 3.7 -1.2 4.9 3.3 5 2 Cl.sup.- mmol/L 103.5 .+-. 0.71 103.5 .+-. 0.71 103.5 .+-. 0.71 103.5 .+-. 0.71 104.0 .+-. 0.00 103.5 .+-. 0.71 0.0 0.0 0.0 0.5 0.0 3.3 5 2 HCO.sub.3.sup.- mmol/L 26.0 .+-. 0.0 24.5 .+-. 0.7 24.0 .+-. 0.0 24.5 .+-. 0.7 23.5 .+-. 0.7 24.0 .+-. 0.0 -5.8 -7.7 -5.8 -9.6 -7.7 4.8 10 2 Gluc mmol/L 5.45 .+-. 0.35 5.20 .+-. 0.42 5.30 .+-. 0.14 5.35 .+-. 0.50 5.2 .+-. 0.28 5.4 .+-. 0.42 -4.6 -2.8 -1.8 -4.6 -0.9 4.4 10 2 Urea mmol/L 7.45 .+-. 2.76 7.30 .+-. 2.83 7.25 .+-. 2.90 7.30 .+-. 2.97 7.3 .+-. 2.97 7.35 .+-. 2.90 -2.0 -2.7 -2.0 -1.3 -2.0 5.0 10 2 Creat .mu.mol/L 106.0 .+-. 2.8 101.5 .+-. 12.0 104.0 .+-. 7.1 103.5 .+-. 6.4 107.0 .+-. 0.00 105.0 .+-. 8.5 -4.2 -1.9 -2.4 0.9 -0.9 5.5 10 2 Urate mmol/L 0.31 .+-. 0.13 0.30 .+-. 0.14 0.30 .+-. 0.14 0.30 .+-. 0.14 0.30 .+-. 0.14 0.30 .+-. 0.14 -3.2 -3.2 -3.2 -3.2 -3.2 3.8 10 2 TP g/L 65.0 .+-. 0.0 64.5 .+-. 0.71 65.5 .+-. 0.71 66.0 .+-. 0.0 63.5 .+-. 0.71 65.5 .+-. 0.71 -0.8 0.8 1.5 -2.3 0.8 3.5 5 2 Alb g/L 39.5 .+-. 0.7 40.5 .+-. 0.7 40.5 .+-. 0.7 40.5 .+-. 0.71 40.0 .+-. 0.0 40.0 .+-. 0.0 2.5 2.5 2.5 1.3 1.3 3.2 5 2 T Bili .mu.mol/L 13.5 .+-. 5.0 11.5 .+-. 5.0 11.0 .+-. 4.2 11.0 .+-. 5.7 12.5 .+-. 5.0 12.5 .+-. 5.0 -14.8 -18.5 -18.5 -7.4 -7.4 7.0 20 2 ALP U/L 66.5 .+-. 17.7 65.5 .+-. 20.5 69.0 .+-. 21.2 67.5 .+-. 21.9 66.0 .+-. 21.2 68.0 .+-. 19.8 -1.5 3.8 1.5 -0.8 2.3 4.8 10 2 GGT U/L 14.5 .+-. 6.4 15.0 .+-. 2.8 15.0 .+-. 7.1 16.5 .+-. 5.0 15.5 .+-. 9.2 14.0 .+-. 7.1 3.4 3.4 13.8.sup..sctn. 6.9 -3.4 7.3 10 2 ALT U/L 32.0 .+-. 2.8 30.5 .+-. 6.4 30.5 .+-. 5.0 31.5 .+-. 3.5 31.5 .+-. 5.0 30.5 .+-. 2.1 -4.7 -4.7 -1.6 -1.6 -4.7 6.8 10 2 AST U/L 23.5 .+-. 0.7 26.5 .+-. 0.7 30.0 .+-. 0.0 29.5 .+-. 2.1 29.5 .+-. 2.1 29.0 .+-. 0.0 12.8 27.7.sup..sctn. 25.5.sup..sctn. 25.5.sup..sctn. 23.4.sup..sctn. 6.2 15 2 LD U/L 189.5 .+-. 10.6 189.5 .+-. 20.5 193.5 .+-. 6.4 197.5 .+-. 20.5 201.5 .+-. 14.8 201.5 .+-. 13.4 0.0 2.1 4.2 6.3 6.3 3.7 15 2 CK U/L 152.0 .+-. 83.4 154.5 .+-. 84.0 145.5 .+-. 90.0 145.5 .+-. 89.8 147.5 .+-. 92.6 145.5 .+-. 94.0 1.6 -4.3 -4.3 -3.0 -4.3 3.7 15 2 TCa mmol/L 2.17 .+-. 0.06 2.18 .+-. 0.09 2.17 .+-. 0.09 2.18 .+-. 0.06 2.15 .+-. 0.08 2.17 .+-. 0.09 0.5 0.0 0.2 -1.2 -0.2 3.5 5 2 Pi mmol/L 1.22 .+-. 0.04 1.22 .+-. 0.04 1.21 .+-. 0.07 1.21 .+-. 0.06 1.21 .+-. 0.04 1.20 .+-. 0.04 0.0 -0.8 -1.2 -1.2 -2.0 3.8 10 2 Mg.sup.2+ mmol/L 0.915 .+-. 0.049 0.935 .+-. 0.05 0.935 .+-. 0.049 0.930 .+-. 0.057 0.950 .+-. 0.057 0.945 .+-. 0.049 2.2 2.2 1.6 3.8 3.3 4.4 10 2 Chol mmol/L 4.60 .+-. 0.00 4.70 .+-. 0.00 4.70 .+-. 0.14 4.70 .+-. 0.00 4.65 .+-. 0.07 4.70 .+-. 0.00 2.2 2.9 2.2 1.1 2.2 3.6 10 2 Trig mmol/L 2.40 .+-. 0.28 2.50 .+-. 0.57 2.55 .+-. 0.50 2.50 .+-. 0.57 2.50 .+-. 0.42 2.50 .+-. 0.42 4.2 6.2 4.2 4.2 4.2 5.3 10 2 HDL-C mmol/L 1.50 .+-. 0.42 1.35 .+-. 0.07 1.50 .+-. 0.14 1.45 .+-. 0.07 1.50 .+-. 0.28 1.50 .+-. 0.14 -10.0 0.0 -3.3 0.0 0.0 5.2 10 2 Fe.sup.2+ .mu.mol/L 13.0 .+-. 5.7 13.0 .+-. 5.7 13.5 .+-. 5.0 13.0 .+-. 5.7 13.0 .+-. 5.7 13.5 .+-. 5.0 0.0 3.8 0.0 0.0 3.8 3.7 10 2 Trf g/L 2.59 .+-. 0.46 2.61 .+-. 0.44 2.58 .+-. 0.35 2.62 .+-. 0.43 2.56 .+-. 0.44 2.55 .+-. 0.36 1.0 -0.2 1.2 -1.0 -1.5 4.8 10 2 Cortisol* nmol/L 236.0 .+-. 27.2 .+-. .+-. .+-. .+-. .+-. 3.6 -1.7 6.4 15 2 FT4* Pmol/L 11.55 .+-. 1.34 12.80 .+-. 0.28 12.90 .+-. 2.55 12.70 .+-. 2.12 13.25 .+-. 4.03 12.75 .+-. 1.34 10.8 11.7 10.0 14.7 10.4 6.5 30 2 TSH* .mu.IU/mL 2.185 .+-. 1.011 2.135 .+-. 0.841 2.430 .+-. 1.202 2.160 .+-. 0.834 2.070 .+-. 0.891 2.085 .+-. 0.884 -2.3 11.2 -1.1 -5.3 -4.6 7.3 15 2 Ferritin* .mu.g/L 50.0 .+-. 0.0 56.0 .+-. 0.0 56.5 .+-. 2.1 51.5 .+-. 9.2 55 .+-. 9.9 51.0 .+-. 7.1 12.0 13.0 3.0 10.0 2.0 7.2 15 2 TnI* .mu.g/L 0.010 .+-. 0.005 0.012 .+-. 0.008 0.010 .+-. 0.002 0.012 .+-. 0.007 0.010 .+-. 0.003 0.007 .+-. 0.002 26.3 0.0 26.3 5.3 -31.6 14 30 2 Haem 0 .+-. 0 0 .+-. 0 0 .+-. 0 0 .+-. 0 0.5 .+-. 0.7 0.5 .+-. 0.7 Index .sup..sctn.= results differed by more than the CAL. Analysis was performed on Beckman DxC800 and *DxI800 analysers Br2 - Brown snake venom 2.0 .mu.g; Br4 - Brown snake venom 4.0 .mu.g; T2 - Taipan snake venom 2.0 .mu.g; T4 and T4R - Taipan snake venom 4.0 .mu.g.
[0791] In summary, we have shown that crude P. textilis and O. scutellatus venoms are potential procoagulants for producing serum in blood clotting tubes. The optimal concentrations need to be determined to avoid latent clotting and maintain analytical integrity.
Example 14b: Use of Crude E. Carinatus Venom as Procoagulant in Clotting Tubes
[0792] The following experiments were conducted to show whether E. carinatus venom could be used to rapidly produce high quality serum from human blood samples.
[0793] Blood was collected from 2 healthy volunteers into 10 Greiner No Additive tubes (#454001, Griener Bio-One, Kremsmuster, Austraia) (4 mL capacity) containing 4 .mu.g of E. carinatus venom. At the same time blood was collected into Greiner standard serum tubes (#456071, Greiner Bio-One, Kremsmuster, Austraia) for comparison.
[0794] Clotting in the venom-containing tubes appeared complete by visual observation in 3 minutes. Samples in the venom-containing tubes were centrifuged after 10 minutes and those in the Greiner serum tubes after 30 minutes. Samples were centrifuged at 3000 g, 10 min and 20.degree. C. (Hereaus 1 S-R centrifuge, Germany). The sera from the Greiner No Additive tubes were immediately transferred into Beckman plain plastic tubes (#448778, Beckman coulter, Brea, Calif., USA) for observation and analysis. No latent clotting was observed in the venom-containing tubes or in the Greiner serum tubes. Samples in the plain plastic tubes were re-centrifuged to remove any cells during the transfer of serum and the clear sera obtained analysed within 2 hours for 31 analytes. Sample analysis was performed within two hour post centrifugation for 31 analytes on the Beckman DxC800 general chemistry analyser and DxI800 immuno-analyser (Beckman Coulter, Brea, Calif., USA). The results are shown in Table 75. There were clinically significant differences with AST and HDL and no other analyte at the concentration level tested between results obtained for the E. carinatus and the Greiner sera.
TABLE-US-00076 TABLE 75 Analytical results for serum samples generated by Echis carinatus snake venom. Mean difference between No of Mean .+-. SD pairs (%) LSC CAL Samples Analyte Units GS E. carinatus 4.0 .mu.g EC4-GS (%) (%) 2 Na.sup.+ mmol/L 138.5 .+-. 0.71 137.5 .+-. 0.71 -0.7 2.8 3 2 K.sup.+ mmol/L 4.0 .+-. 0.14 4.0 .+-. 0.14 0.0 3.3 5 2 Cl.sup.- mmol/L 103 .+-. 0.0 103.5 .+-. 0.7 -0.5 3.3 5 2 HCO.sub.3.sup.- mmol/L 27.4 .+-. 0.92 26.1 .+-. 0.14 -4.6 4.8 10 2 Gluc mmol/L 5.45 .+-. 0.07 5.35 .+-. 0.07 -1.8 4.4 10 2 Urea mmol/L 7.25 .+-. 0.64 7.30 .+-. 0.71 0.7 5.0 10 2 Creat .mu.mol/L 88.5 .+-. 12.1 86.0 .+-. 17.0 -2.8 5.5 10 2 Urate mmol/L 0.28 .+-. 0.09 0.28 .+-. 0.09 0 3.8 10 2 TP g/L 63.5 .+-. 0.7 63.5 .+-. 0.7 0 3.5 5 2 Alb g/L 39.5 .+-. 0.7 40.5 .+-. 0.7 2.5 3.2 5 2 T Bili .mu.mol/L 13.5 .+-. 6.4 12.0 .+-. 5.7 -11.1 7.0 20 2 ALP U/L 71.5 .+-. 23.3 70.0 .+-. 19.8 -2.1 4.8 10 2 GGT U/L 14.0 .+-. 4.2 14.5 .+-. 7.8 3.6 7.3 10 2 ALT U/L 34.0 .+-. 0.0 35.5 .+-. 2.1 4.4 6.8 10 2 AST U/L 20.5 .+-. 2.1 27.0 .+-. 00 31.7.sup..sctn. 6.2 15 2 LD U/L 185 .+-. 5.7 193 .+-. 11.3 4.3 3.7 15 2 CK U/L 135 .+-. 50.9 133.5 .+-. 48.8 -1.1 3.7 15 2 TCa mmol/L 2.20 .+-. 0.014 2.17 .+-. 0.021 -1.6 3.5 5 2 Pi mmol/L 1.215 .+-. 0.247 1.17 .+-. 0.226 -3.7 3.8 10 2 Mg.sup.2+ mmol/L 0.91 .+-. 0.0 0.93 .+-. 0.04 1.6 4.4 10 2 Chol mmol/L 4.75 .+-. 0.64 4.75 .+-. 0.64 0.0 3.6 10 2 Trig mmol/L 1.45 .+-. 0.50 1.50 .+-. 0.14 3.4 5.3 10 2 HDL-C mmol/L 1.25 .+-. 0.21 1.80 .+-. 0.57 44.0.sup..sctn. 5.2 10 2 Fe.sup.2+ .mu.mol/L 13.5 .+-. 5.0 13.5 .+-. 5.0 0 3.7 10 2 Trf g/L 2.62 .+-. 0.49 2.60 .+-. 0.44 1.3 4.8 10 2 Cortisol* nmol/L 232 .+-. 10.6 238 .+-. 7.8 2.6 6.4 15 2 FT4* Pmol/L 12.2 .+-. 0.6 12.6 .+-. 0.7 3.7 6.5 30 2 TSH* .mu.IU/mL 2.16 .+-. 0.76 2.10 .+-. 0.67 -3.0 7.3 15 2 Ferritin* .mu.g/L 50 .+-. 2.8 47 .+-. 2.8 -6.0 7.2 15 2 TnI* .mu.g/L 0.024 .+-. 0.000 0.020 .+-. 0.006 -18.8 14 30 2 Haem 0 .+-. 0 0 .+-. 0 Index .sup..sctn.= results were different by more than the CAL. Analysis was performed on Beckman DxC800 and *DxI800 analysers EC4 - Echis carinatus venom 4.0 .mu.g.
[0795] As these are preliminary results, further investigation is needed to establish why there were clinically significant differences for the AST and HDL results.
[0796] In summary, we have shown that use of crude E. carinatus venom as procoagulant in blood clotting tubes may be feasible. The optimal concentration and further studies on analyte interference need to be accurately determined to avoid latent clotting and maintain analytical integrity.
Example 15: Stability Studies
[0797] For a venom prothrombin activator to be useful as a component of blood collection devices, it must meet three types of stability requirement:
[0798] (1) Stability during bulk storage: It needs to be stable for the period after purification before use in manufacture of the tubes. For certainty of supply, this storage period may be for several months. Storage could be as a concentrated aqueous solution or as lyophilized solid. Storage could be under refrigeration if required.
[0799] (2) Stability during manufacture: It needs to be stable during the manufacture of the blood collection device. This process is likely to involve addition of an aliquot of a stock solution of the prothrombin activator to tubes which already contain other components such as the separating gel, the surfactant and possible a particulate procoagulant. This would be followed by drying to give a surface layer of prothrombin activator, sealing the tubes under vacuum and sterilization by radiation.
[0800] (3) Stability during storage of the device prior to its use: It would be necessary for the activator to retain close to full activity for a period of at least 12 months at room temperature, say 23 C, preferably for a longer period at a higher temperature.
[0801] Experiments performed to date suggest that conditions will be found to meet these requirements for at least some of the venom prothrombin activators.
Example 15a: Stability During Bulk Storage
[0802] The stability of PtPA when maintained at a range of temperatures as a dilute solution in phosphate buffered saline at pH 7 in plain plastic Eppendorf tubes was determined. Results of plasma clotting assays are shown in FIG. 58. The results show that there was no significant loss of clotting activity after two weeks (336 hours) at 4 or 25.degree. C. Similar results were obtained when the activity of PtPA was measured against the Factor Xa chromogenic substrate S-2222. At 37.degree. C. PtPA lost a significant amount of its activity after 168 hours.
[0803] In related experiments, it was demonstrated that pH 7.4 was the optimal temperature for storage and that the addition of calcium ions to the storage buffer stabilized the plasma clotting activity.
[0804] These experiments indicate that it should be possible to find conditions for the bulk storage of PtPA and the closely related OsPA so that they retain all or almost all of their activity on storage for extended periods.
Example 15b: Stability During Manufacture
[0805] An experiment was performed to determine the effect of gamma radiation on OsPA added to blood collection tubes as a dilute solution.
[0806] Stock OsPA (1.089 mg/mL; 4.36 mM) was diluted 1 in 25 with 0.02 M Hepes buffer pH 7.4. 50 .mu.L was added to each of 5 plain blood collection tubes with no other additives were gamma irradiated (16 megaBeq) and to 5 similar tubes which were not irradiated. Similarly, 50 .mu.L of diluted OsPA was added to each of 10 Greiner silica-containing blood collection tubes (Greiner serum tubes) and 5 of these tubes were irradiated.
[0807] After gamma irradiation, all tubes were made up to 2.0 mL with 0.02 M Hepes buffer pH 7.4. Aliquots were then taken and tested for hydrolytic activity against S-2765 and clotting activity using normal citrated plasma with added 10 mM calcium.
[0808] The results are shown in FIGS. 59 and 60, and show that irradiation had no effect on either clotting activity against recalcified citrated plasma or hydrolytic activity against the chromogenic substrate S-2765.
[0809] The addition of OsPA (1.5 nM) to plain blood collection tube showed a significant loss of activity as compared to the freshly diluted OsPA. This can be explained by hydrophobic binding of the OsPA to the plain plastic tube. Adding OsPA to silica-containing blood collection tubes showed recovery of greater than 95% the hydrolytic and clotting activities. Again, no loss activity was observed with gamma irradiation treatment.
[0810] In conclusion, gamma irradiation of OsPA in a blood collection tube did not affect either hydrolytic or clotting activity. Adding OsPA to a clotting tube containing silica, gel barrier, and surfactant is the most effective combination determined to date to clot normal blood in less than 1 minute.
Example 15c: Stability During Storage of the Device
[0811] Aliquots (50 .mu.L) of a diluted solution of OsPA in 0.02 M Hepes buffer, pH 7.4 (1.5 nM) were placed in Greiner standard serum tubes. Each tube was rotated to coat the surface with the OsPA and the tubes were placed in a vacuum dessicator and purged by a stream of dry nitrogen before applying a low vacuum for 3 days at 23.degree. C. to dry. The dessicator was opened, flushed with nitrogen and the tubes re-sealed. The tubes were then maintained at room temperature (23.degree. C.). After specified times, three tubes were opened, the contents redissolved in 2 mL water and assayed against the Factor Xa specific substrate S-2222. The results are shown in FIG. 61, and show that the activity (expressed as increase in A405/min) of OsPA towards S-2222 was effectively unchanged by storage at 23.degree. C. as a dry surface layer in the Greiner serum tubes for 14 days. All assays were done in triplicate. In FIG. 61, the right hand bar is the activity of a freshly diluted sample of OsPA stock solution (4.05 uM) and its activity was the same as that found in the tubes, showing that the process used to prepare the tubes did not cause any significant loss of OsPA activity.
Example 16: Preparation of Blood Collection Tube
[0812] All commercial serum and plasma tubes are now plastic and contain a number of components which have been developed by the suppliers to enhance the quality of the samples prepared in them. As discussed above, some container (e.g. tube) for preparation of a serum sample or a plasma sample contains:
[0813] (1) a gel barrier which aims to separate cells (and clot in the case of serum) from the supernatant (serum or plasma) after centrifugation and to limit re-mixing. The gel is added as a warm liquid which solidifies on cooling to room temperature;
[0814] (2) a surfactant sprayed on the inner surface of the plastic tube above the gel barrier to decrease attachment of cells, debris and clot material and cell lysis; and
[0815] (3) a procoagulant/anticoagulant, usually particulate, sprayed as a suspension onto the inner surface of the tube on top of the surfactant.
[0816] The present invention envisages the a serum tube (e.g., a plastic tube) comprising components (1), (2), and (3) as described above, where the tube is prepared by adding components (1) and (2) to the tube first, and subsequently adding component (3) to the tube. Component (3) comprises a procoagulant comprising a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator as herein defined, for example, component (3) may comprise existing particulate procoagulants used in some commercial serum tubes. The amount of prothrombin activator within each tube can be determined using the experimental results described in detail above, and routine techniques known in the art. The tubes are then evacuated and sterile capped. The tubes are then stored at room temperature.
[0817] Also envisaged a serum tube (e.g., a plastic tube) comprising component (3) and optionally comprising components (1) and/or (2). The amount of prothrombin activator within each tube can be determined using the experimental results described in detail above, and routine techniques known in the art. The tubes are evacuated and sterile capped and can be stored at room temperature.
Example 17: Point of Care Embodiment
[0818] As discussed above, "point-of-care" testing means that the testing is performed at or near the site of patient care. Point of care testing is becoming increasingly popular in hospital and other environments where there is a need for fast results. It is accomplished using a variety of devices some of which are relatively inexpensive, small, and portable.
[0819] In current practice, many point-of-care devices work as follows: a droplet of blood is placed on a membrane which retains blood cells but allows plasma to diffuse through the membrane into a microfluidic device with multiple channels in which a number of analyses are performed. As discussed elsewhere, serum is preferred to plasma as the sample for biochemical or other pathology assays. In this embodiment, the present invention contemplates the use of clotting compositions comprising, consisting essentially of, or consisting of a prothrombin activator to produce serum in devices designed for point of care testing.
[0820] In one such device, suitable for hospital and doctors' surgery use, a blood sample obtained by venipuncture is collected into a syringe (or similar) containing a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator, where composition is achieved in .about.30 seconds. The device is designed to allow either filtration or centrifugation to rapidly separate the serum from the clot. The serum so obtained is used in existing microfluidic devices for biochemical or other pathology assays, with the range of analyses increased as the sample being analysed is serum rather than plasma that is used in existing point-of-care devices. Furthermore, a sample of the serum may be retained to allow further analysis in the pathology laboratory.
[0821] The filtration device in the syringe described above may comprise a filtration mechanism that allows the serum to be pushed through after 5 minutes for analysis. Suitably, the syringe may be divided in compartments as illustrated in FIG. 62, with the syringe labelled as follows:
[0822] A--entry port that will have blood collection needle adapter that would unscrew to minimising prodruding component;
[0823] B--port for dispensing serum or entry point for point of care sample needles;
[0824] C--valves to allow for one way entry of blood and prevent blood being pushed into serum compartment;
[0825] D--plunger filter that would allow for serum to be pushed out into the serum compartment (front part of the syringe);
[0826] E--opening for blood to enter blood compartment.
[0827] F--compartment ion which clotting occurs.
[0828] In FIG. 62, G is coated with the clotting composition on the inside to set the clotting process in place. It is then pushed to force the serum across the filter (D).
[0829] In another such device, the point-of-care device involves a capillary system, where part of the device is coated with a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator and an absorbant that can draw the serum into the reaction chamber for analysis.
[0830] In another such device, a clotting composition comprising, consisting essentially of, or consisting of a prothrombin activator could be coupled to the membrane of a microfluidic device so as to achieve rapid clotting of a blood droplet placed on the membrane. The serum so formed diffuses into the device and is analysed in place of the plasma currently analysed in such devices.
SUMMARY OF EXAMPLES
[0831] Obtaining a suitable sample for analysis in a timely manner is pivotal in clinical chemistry service provision. Serum is recognised as being the cleaner sample but because of time required for completion of clotting (30 minutes to ensure complete clotting in most currently available commercial tubes plus analysis time in the standard protocols) and an ever-increasing number of patients on anti-coagulant therapy, plasma is commonly used, especially in hospital environments. However, latent clotting (clotting after centrifugation) can occur in plasma samples leading to analytical errors and delays in delivery of critical results. Some of these problems are illustrated in the above examples.
[0832] The results in the above examples also show that use of prothrombin activators can produce high quality serum very quickly from blood samples from a wide variety of patients, including patients being treated with anticoagulants. They provide suitable alternatives to lithium heparin plasma and to conventional serum tubes by: [0833] having the clotting process completed in .about.<5 minutes (time taken to deliver samples to the laboratory) allowing rapid analysis after blood sample collection and hence rapid turn around times comparable to the times for plasma tubes and better than those for most current serum tubes; [0834] at the same time providing high quality serum samples for blood from both healthy and all anticoagulated individuals that may be encountered in a clinical environment with minimal analyte interference from cellular debris and/or latent microclotting; and [0835] producing a single serum sample that is suitable for a very wide range of assays typically used in clinical environment, the serum sample being superior to plasma for analyte analysis, and the single sample reducing the burden on scarce resources (blood in critically ill patients, staff time and consumables by needing to collect fewer blood tubes).
[0836] Throughout the specification the aim has been to describe the preferred embodiments of the invention without limiting the invention to any one embodiment or specific collection of features. Those of skill in the art will therefore appreciate that, in light of the instant disclosure, various modifications and changes can be made in the particular embodiments exemplified without departing from the scope of the present invention. All such modifications and changes are intended to be included within the scope of the appended claims.
BIBLIOGRAPHY
[0837] Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., Miller, W., Lipman, D. J., "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs." Nucleic Acids Res. 1997 Sep. 1; 25(17): 3389-402. [0838] Arkin, A. P., Youvan, D. C., "An algorithm for protein engineering: simulations of recursive ensemble mutagenesis" 1992 Proc. Natl. Acad. Sci. USA 89: 7811-7815. [0839] Atherton, E., Shephard, R. C., Solid Phase Peptide Synthesis--A Practical Approach 1989, IRL Press, Oxford England. [0840] Austen, D. E., Rhymes, I. L., "Laboratory manual of blood coagulation" (1975) Blackwell Scientific Publications, London, p 37. [0841] Bos, M. H. A., Boltz, M., St Pierre, L., Masci, P. P., de Jersey, J., Lavin, M. F., Camire, R. M., "Venom factor V from the common brown snake escapes hemostatic regulation through procoagulant adaptations" Blood 16 Jul. 2009; 114(3): 686-692. [0842] Camenzind, E., Bakker, W. H., Reijs, A., van Geijlswijk, I. M., Boersma, E., Kutryk, M. J., Krenning, E. P., Roelandt, J. R., Serruys, P. W., "Site-specific intracoronary heparin delivery in humans after balloon angioplasty. A radioisotopic assessment of regional pharmacokinetics." Circulation 1997; 96: 154-65. [0843] Ciuti, R., Rinaldi, G., "Serum and plasma compared for use in 19 common chemical tests performed in the Hitachi 737 analyzer." Clin Chem. 1989; 35: 1562-3. [0844] Cowley, D. M., Nagle, B. A., Chalmers, A. H., Sinton, T. J., "Effects of platelets on collection of specimens for assay of ammonia in plasma." Clin Chem 1985; 31: 332. [0845] Davidson, A. S., Darn, S. M., Sodi, R., "Can lithium heparin plasma be used for protein electrophoresis and paraprotein identification?" Ann Clin Biochem 2006; 43: 31-34. [0846] Dayhoff, M. O., Schwartz, R. M., Orcutt, B. C., et al., "A model of evolutionary change in proteins. Matrices for determining distance relationships" 1978 In Atlas of protein sequence and structure (Dayhoff, M. O. ed.), vol5, pp 345-358, National Biomedical Research Foundation, Washington DC. [0847] Delagrave, S., Goldman, E. R., Youvan, D. C., "Recursive ensemble mutagenesis" April 1993 Protein Eng. 6(3): 327-31. [0848] Devereux, J., Haeberli, P., Smithies, O., "A comprehensive set of sequence analysis programs for the VAX." Nucleic Acids Res. 1984 Jan. 11; 12(1 Pt 1): 387-95. [0849] Dimeski, G., Masci, P. P., Trabi, M., Lavin, M. F., de Jersey, J., "Evaluation of the Becton-Dickinson rapid serum tube: does it provide a suitable alternative to lithium heparin plasma tubes?" Clin Chem Lab Med 2010; 48(5): 2111-2120. [0850] Dimeski, G., Badrick, T., Flatman, R., Ormiston, B., "Roche IFCC Methods for Lactate Dehydrogenase Tested for Duplicate Errors with Greiner and Becton-Dickinson Lithium-Heparin and Greiner Serum Samples" Clin Chem 2004; 50(12): 2391-2392. [0851] Dimeski, G., Carter, A., "Rare IgM Interference with Roche/Hitachi Modular Glucose and .gamma.-Glutamyltransferase Methods in Heparin Samples" Clin Chem 2005; 51(11): 2202-2204. [0852] Dimeski, G., Clague, A. E., Hickman, P. E., "Correcting and reporting of potassium results in haemolysed samples" Ann. Clin. Biochem. 2005; 42: 119-123. [0853] Dimeski, G., McWhinney, B., Jones, B., Mason, R., Carter, A., "Extent of bilirubin interference in Beckman-Coulter creatinine methods." Ann Clin Biochem 2008; 45:91-92. [0854] Filippovich, I., Sorokina, N., St Pierre, L., Filght, S., de Jersey, J., Perry, N., Masci, P. P., Lavin, M. F. "Cloning and functional expression of venom prothrombin activator protease from Pseudonaja textilis with whole blood procoagulant activity" British Journal of Haematology 2005; 131: 237-246. [0855] Gonnet, G. H., Cohen, M. A., Benner, S. A., "Exhaustive matching of the entire protein sequence database" Jun. 5 1992 Science 256(5062): 1443-5. [0856] Hartland, A. J., Neary, R. H., "Serum potassium is unreliable as an estimate of in vivo plasma potassium" Clin Chem 1999; 45:1091-1092. [0857] Kini, R. M., Morita, T., Rosing, J., "Classification and Nomenclature of Prothrombin Activators Isolated from Snake Venoms" Thromb. Haemost. 2001 85:710-711. [0858] Kini, R. M., "The intriguing world of prothrombin activators from snake venom" Toxicon (2005) 45: 1133-1145. [0859] Kunkel, T. A., "Rapid and Efficient site-specific mutagenesis without phenotypic selection" 1985 Proc. Natl. Acad. Sci. USA, 82: 488-492. [0860] Kunkel, T. A., Roberts, J. D., Zakour, R. A., "Rapid and Efficient site-specific mutagenesis without phenotypic selection" 1987 Methods in Enzymol. 154: 367-382. [0861] Kuzmic, P., "Program DYNAFIT for the Analysis of Enzyme Kinetic Data: Application to HIV Proteinase" 1996 Anal. Biochem. 237: 260-273. [0862] Lavin, M. F., Masci, P. P., "Prothrombinase complexes with different physiological roles" Thromb Haemost 2009; 102: 421-423. [0863] Masci, P. P., "The Effects of Australian Snake Venoms on Coagulation and Fibrinolysis" Thesis for Masters of Science in the subject of Biochemistry, July 1986, University of Queensland, St Lucia, Brisbane, Australia. [0864] Masci, P. P., Whitaker, A. N., de Jersey, J., "Purification and characterization of a prothrombin activator from the venom of the Australian brown snake, Pseudonaj a textilis textilis" Biochem. Int. 1988; 17(5):825-835. [0865] Miles, R. R., Roberts, R. F., Putnam, A. R., Roberts, W. L., "Comparison of serum and heparinized plasma samples for measurement of chemistry analytes." Clin Chem 2004; 50: 1704-5. [0866] National Pathology Laboratory Accreditation Advisory Council--Requirements for the Retention of Laboratory Records and Diagnostic Material (Fifth Edition 2009), [0867] Nishida, S., Fujita, T., Kohno, N., Atoda, H., Morita, T., Takeya, H., Kido, I., Paine, M. J. I., Kawabata, S-i., Iwanaga, S. "cDNA Cloning and Deduced Amino Acid Sequence of Prothrombin Activator (Ecarin) from Kenyan Echis carinatus venom." Biochemistry 1995; 34: 1771-1778. [0868] Morita, T., Iwanaga, S., "Prothrombin activator from Echis carinatus venom" Meth Enzymol 1981; 80-pt. C: 303-311. [0869] Nicholson, J., Mirtschin, P., Madaras, F., Venning, M., Kokkinn, M., "Digestive properties of the venom of the Australian Coastal Taipan, Oxyuranus scultellatus (Peters, 1867)" Toxicon 2006: 48: 422-428. [0870] O'Keane M. P., Cunningham, S. K., "Evaluation of three different specimen types (serum, plasma lithium heparin and serum gel separator) for analysis of certain analytes: clinical significance of differences in results and efficiency in use." Clin Chem Lab Med. 2006; 44: 662-8. [0871] Parker, H. W., Grandison, A. G. C., "Snakes: A natural history" University of Queensland Press, St. Lucia, Queensland 1977: pp 5, 59 and 87. [0872] Roberge, J. Y., Beebe, X., Danishefsky, S. J., "A strategy for a convergent synthesis of N-linked glycopeptides on a solid support" Science 1995 269(5221); 202-204. [0873] Rosing, J., Tans, G., "Inventory of Exogenous Prothrombin Activators" Thrombosis and Haemostasis (Stuttgart) 1991 65(2): 627-630. [0874] Rosing, J., Tans, G., "Structural and Functional Properties of Snake Venom Prothrombin Activators" Toxicon 30(12): 1515-1527, 1992. [0875] Sambrook, J., Russell, D. W., Molecular cloning. A laboratory manual (third edition) 2001; Cold Spring Harbor Laboratory Press. [0876] Schieck, A., Kornalik, F., Habermann, E. "The prothrombin-activating principle from Echis carinatus venom. I. Preparation and biochemical properties" Naunyn-Schmiedeberg's Arch Pharmacol. 1972; 272: 402-416. [0877] Sheppard, C. A., Allen, R. C., Austin, G. E., Young, A. N., Ribeiro, M. A., Fantz, C. R., "Paraprotein interference in automated chemistry analyzers." Clin Chem 2005 June; 51(6): 1077-8. [0878] Smith, G. F., Craft, T. J., "Heparin reacts stoichiometrically with thrombin during thrombin inhibition in human plasma." Biochem. Biophys. Res. Commun. 1976; 71: 738-45. [0879] Sonder, S. A., Fenton, J. W., "Thrombin specificity with tripeptide chromogenic substrates: comparison of human and bovine thrombins with and without fibrinogen clotting activities" Clin. Chem. 1986;32:934-7. [0880] Speijer, H., Goversriemslag, J. W. P., Zwaal, R. F. A., Rosing, J., "Prothrombin activation by an activator from the venom of Oxyuranus scutellatus (Taipan snake)" J. Biol. Chem. 1986; 261(28): 3258-3267. [0881] Starr, H., Rhodes, P., Lam-Po-Tang, P. R., Archer, G. T., "Prothrombin times: an evaluation of four thromboplastins and four machines." Pathology 1980; 12: 567-574. [0882] St Pierre, L., Masci, P., Filippovich, I., Sorokina, N., Marsh, N., Miller, D. J., Lavin, M. F., "Comparative Analysis of Prothrombin Activators from the Venom of Australian Elapids" Mol. Biol. Evol. 2005; 22(9): 1853-1864. [0883] Tans, G., Govers-Riemslag, J. W., van Rijn, J. L., Rosing, J. J., "Purification and properties of a prothrombin activator from the venom of Notechis scutatus scutatus" Biol. Chem. 1985 Aug. 5; 260(16): 9366-72. [0884] Thorelli, E., Kaufman, R. J., Dahlback, B., "Cleavage requirements of factor V in tissue-factor induced thrombin generation." Thromb. Haemost. 1998 July; 80(1) 92-98. [0885] Wannaslip, N., Sribhen, K., Pussara, N., Hwanpuch, T., Wangchaijaroekit, S., Opartkiattikul, N., "Heparin is unsuitable anticoagulant for the detection of plasma ammonia." Clin Chimica Acta 2006; 371: 196-7. [0886] Watson, J. D., Hopkins, N. H., Roberts, J. W., Steitz, J. A., Weiner, A. M. Molecular Biology of the Gene. fourth edition, 1987 The Benjamin/Cummings Publishing Co., Inc., Menlo Park, Calif. [0887] Yamada, D., Sekiya, F., Morita, T., "Isolation and Characterization of Carinactivase, a Novel Prothrombin Activator in Echis carinatus Venom with a Unique Catalytic Mechanism" J. Biol. Chem. 1996; 271(9): 5200-5207. [0888] Yamada, D., Morita, T., "Purification and Characterization of a Ca.sup.2+-Dependent Prothrombin Activator, Multactivase, from the Venom of Echis multisquamatus" J. Biochem. 1997; 122: 991-997. [0889] Yamanouye, N., Kerchove, C. M., Moura-da-Silva, A. M., Carneiro, S. M., Markus, R. P., "Long-term primary culture of secretory cells of Bothrops jararaca gland for venom production in vitro" Nature Protocols 2007; 1: 2763-2766. [0890] Yonemura, H., Imamura, T., Soejima, K., Nakahara, Y., Morikawa, W., Ushio, Y., Kamachi, Y., Nakatake, H., Sugawara, K., Nakagaki, T., Nozaki, C., "Preparation of Recombinant .alpha.-Thrombin: High-Level Expression of Recombinant Human Prethrombin-2 and Its Activation by Recombinant Ecarin" J. Biochem. 2004; 135: 577-582. [0891] Zubay, G., Biochemistry, third edition Wm.C. Brown Publishers, Oxford (1993).
Sequence CWU 1
1
531609PRTEchis carinatus 1Met Ile Gln Ile Leu Leu Val Ile Ile Cys
Leu Ala Val Phe Pro Tyr1 5 10 15Gln Gly Cys Ser Ile Ile Leu Gly Ser
Gly Asn Val Asn Asp Tyr Glu 20 25 30Val Val Tyr Pro Gln Lys Val Thr
Ala Leu Pro Lys Gly Ala Val Gln 35 40 45Gln Pro Glu Gln Lys Tyr Glu
Asp Ala Met Gln Tyr Glu Phe Glu Val 50 55 60Lys Gly Glu Pro Val Val
His Leu Glu Lys Asn Lys Glu Leu Phe Ser65 70 75 80Glu Asp Tyr Ser
Glu Thr His Tyr Ser Ser Asp Asp Arg Glu Ile Thr 85 90 95Thr Asn Pro
Ser Val Glu Asp His Cys Tyr Tyr His Gly Arg Ile Gln 100 105 110Asn
Asp Ala Glu Ser Thr Ala Ser Ile Ser Ala Cys Asn Gly Leu Lys 115 120
125Gly His Phe Lys Leu Arg Gly Glu Thr Tyr Phe Ile Glu Pro Leu Lys
130 135 140Ile Pro Asp Ser Glu His Ala Val Tyr Lys Tyr Glu Asn Ile
Glu Asn145 150 155 160Glu Asp Glu Ala Pro Lys Met Cys Gly Val Thr
Gln Asp Asn Trp Glu 165 170 175Ser Asp Glu Pro Ile Lys Lys Thr Leu
Gly Leu Ile Val Pro Pro His 180 185 190Glu Arg Lys Phe Glu Lys Lys
Phe Ile Glu Leu Val Val Val Val Asp 195 200 205His Ser Met Val Thr
Lys Tyr Asn Asn Asp Ser Thr Ala Ile Arg Thr 210 215 220Trp Ile Tyr
Glu Leu Asn Thr Val Asn Glu Ile Tyr Leu Pro Phe Asn225 230 235
240Ile Arg Val Ala Leu Val Gly Leu Glu Phe Trp Cys Asn Gly Asp Leu
245 250 255Ile Asn Val Thr Ser Thr Ala Asp Asp Thr Leu His Ser Phe
Gly Glu 260 265 270Trp Arg Ala Ser Asp Leu Leu Asn Arg Lys Arg His
Asp His Ala Gln 275 280 285Leu Leu Thr Asn Val Thr Leu Asp His Ser
Thr Leu Gly Ile Thr Phe 290 295 300Val Tyr Gly Cys Lys Ser Asp Arg
Ser Val Glu Leu Ile Leu Asp Tyr305 310 315 320Ser Asn Ile Thr Phe
Asn Met Ala Tyr Ile Ile Ala His Glu Met Gly 325 330 335His Ser Leu
Gly Met Leu His Asp Thr Lys Phe Cys Thr Cys Gly Ala 340 345 350Lys
Pro Cys Ile Met Phe Gly Lys Glu Ser Ile Pro Pro Pro Lys Glu 355 360
365Phe Ser Ser Cys Ser Tyr Asp Gln Tyr Asn Lys Tyr Leu Leu Lys Tyr
370 375 380Asn Pro Cys Ile Leu Asp Pro Pro Leu Arg Lys Asp Ile Ala
Ser Pro385 390 395 400Ala Val Cys Gly Asn Glu Ile Trp Glu Glu Gly
Glu Glu Cys Asp Cys 405 410 415Gly Ser Pro Ala Asp Cys Arg Asn Pro
Cys Cys Asp Ala Ala Thr Cys 420 425 430Lys Leu Lys Pro Gly Ala Glu
Cys Gly Asn Gly Glu Cys Cys Asp Lys 435 440 445Cys Lys Ile Arg Lys
Ala Gly Thr Glu Cys Arg Pro Ala Arg Asp Asp 450 455 460Cys Val Ala
Glu His Cys Thr Gly Gln Ser Ala Glu Cys Pro Arg Asn465 470 475
480Glu Phe Gln Arg Asn Gly Gln Pro Cys Leu Asn Asn Ser Gly Tyr Cys
485 490 495Tyr Asn Gly Asp Cys Pro Ile Met Leu Asn Gln Cys Ile Ala
Leu Phe 500 505 510Ser Pro Ser Ala Thr Val Ala Gln Asp Ser Cys Phe
Gln Arg Asn Leu 515 520 525Gln Gly Ser Tyr Tyr Gly Tyr Cys Thr Lys
Glu Ile Gly Tyr Tyr Gly 530 535 540Arg Phe Pro Cys Ala Pro Gln Asp
Val Lys Cys Gly Arg Leu Tyr Cys545 550 555 560Leu Asp Asn Ser Phe
Lys Lys Asn Met Arg Cys Lys Asn Asp Tyr Ser 565 570 575Tyr Ala Asp
Glu Asn Lys Gly Ile Val Glu Pro Gly Thr Lys Cys Glu 580 585 590Asp
Gly Lys Val Cys Ile Asn Arg Lys Cys Val Asp Val Asn Thr Ala 595 600
605Tyr263PRTBothrops asper 2Ser His Asp Asn Ala Gln Leu Leu Thr Ala
Ile Lys Ala Tyr Ile Ala1 5 10 15Thr Met Cys Asp Pro Lys Met Ala Val
Ile Met Ala His Glu Ile Gly 20 25 30His Gly Gly Tyr Tyr Gly Tyr Cys
Arg Lys Ile Pro Cys Ala Pro Glu 35 40 45Asp Val Lys Asp Asp Asp Ile
Gly Met Val Leu Pro Gly Thr Lys 50 55 60332PRTEchis
carinatusmisc_feature(23)..(23)Xaa can be any naturally occurring
amino acidmisc_feature(25)..(25)Xaa can be any naturally occurring
amino acid 3Ser Arg Lys Gln Lys Phe Asp Lys Lys Phe Ile Lys Leu Val
Ile Val1 5 10 15Val Asp His Ser Met Val Xaa Lys Xaa Asn Asn Asp Leu
Ile Ala Ile 20 25 30429PRTEchis multisquamatus 4Asp Cys Leu Pro Gly
Trp Ser Val Tyr Glu Gly Arg Cys Tyr Lys Val1 5 10 15Phe Asn Gln Lys
Thr Trp Lys Ala Ala Glu Lys Phe Cys 20 2554737DNAPseudonaja
textilis 5gagccacaga atacatttat gtggggaagt ggcaagttgc tgcaggcaga
actgactttt 60gtatatcttt cagcacatta gcctcaatgg gaagatacag tgtgagccct
gtccccaaat 120gtcttctact gatgttcctg ggttggtcag ggctgaagta
ttacgaagtg aatgcagctc 180agctcaggga gtaccatata gctgctcagc
tggaagactg ggattacaac ccccaacctg 240aggagctatc cagattatca
gagtcagatc ttacgtttaa aaaaattgtc tatagagaat 300atgaactaga
tttcaaacaa gagaagccaa gagatgagct ctcagggctc ctagggccaa
360cactacgtgg agaagtggga gacatcctca taatttattt caagaatttt
gctactcagc 420ctgtgagcat tcacccgcag agtgccgtgt acaacaaatg
gtcagaaggt tcttcatatt 480ctgatggaac atcagatgtg gaaagactgg
atgatgctgt gcctccaggc cagtcgttca 540agtatgtgtg gaatatcact
gcagaaattg ggccaaagaa agctgatcct ccctgtctca 600cttatgcgta
ctactcacat gtaaacatgg tgcgagactt taattctggt ctcattggtg
660ctttgctgat atgtaaagaa ggaagcctga atgcaaatgg ttcacaaaaa
ttcttcaaca 720gagaatatgt gctgatgttt tctgtgtttg atgaaagcaa
gaactggtac agaaagccct 780cactacagta cacaattaat gggtttgcca
atggaacatt gcctgatgtt caggcttgtg 840cttatgatca tattagctgg
catttgatag gaatgagttc cagtcctgag atcttctctg 900ttcacttcaa
tggacaaacc ttggaacaaa accattacaa agtgtcaacc atcaaccttg
960tcggaggtgc ctcagtaaca gccaacatgt cagtgagcag gacaggaaaa
tggctaatat 1020cttctctggt tgcaaagcat ctacaagctg ggatgtatgg
ttatctaaat atcaaagact 1080gtggaaatcc agatacttta acaagaaagt
tatcctttag agaactgagg aggattatga 1140actgggaata tttcattgct
gcagaagaaa tcacctggga ttatgctcca gaaattccta 1200gcagtgttga
cagaagatac aaagctcagt atctggataa tttttcaaat tttattggca
1260agaaatacaa aaaggcagtt ttcaggcaat ataaagacag caatttcact
aaaccgacct 1320atgccatttg gcccaaagaa cgtggaattc tgggccccgt
tatcagagct aaagtcagag 1380acacaataag tattgtattc aaaaatctgg
ccagtcgacc ttacagcatt tatgtgcatg 1440gagtttccgt ttcaaaagat
gcagaaggag ctatttatcc ttcagatccc aaagagaata 1500taactcatgg
caaagcagtt gaaccaggac aggtctacac atataaatgg actgtgctgg
1560atacagatga acctacagta aaggattctg agtgcattac taaattatat
catagtgctg 1620tggacatgac aagagatatt gcttcaggac ttattgggcc
acttctggtt tgtaaacaca 1680aggcactcag cgtcaagggc gtacagaata
aagctgatgt ggaacagcat gcagtcttcg 1740cagtgtttga tgaaaacaag
agctggtact tggaagacaa tatcaagaaa tactgcagca 1800atccttccac
tgttaagaaa gatgacccta aattttacaa gtccaatgtt atgtacacac
1860tcaatggcta tgcatcagat agaacagagg ttttggggtt tcatcagtct
gaagttgttg 1920aatggcacct caccagcgta ggtacagtgg atgagattgt
tccagtacat ctttctggtc 1980acaccttctt atccaaggga aaacatcaag
atattttaaa tcttttcccc atgagtggtg 2040aatcggctac tgtaacaatg
gacaatctag gaacctggct tctgtcatca tggggctcct 2100gtgagatgag
caatggcatg agattgagat ttttggatgc caattatgat gatgaagatg
2160agggaaatga agaagaggaa gaagatgatg gcgatatttt tgccgacatt
ttcattcctc 2220cagaagtagt aaaaaagaaa gaaaaggacc ccgtaaattt
tgtatcagac ccagaatcgg 2280ataagatagc aaaagaatta ggattattag
atgacgagga taatcaagaa gagtcacaca 2340atgtacagac agaggatgat
gaagaacagc taatgatagc tacaatgctt gggtttcgat 2400catttaaggg
gtcagttgct gaagaagaat tgaatctcac agctctagct ttagaagaag
2460atgcccatgc ttctgatcct cgaattgaca gtaatagtgc acgtaatcct
gatgacatag 2520ctggacgcta cctgcgtact atcaaccgtg gaaataaaag
gaggtactac attgcagcag 2580aagaagtttt gtgggactac tcaccgatcg
gaaaaagtca agtgagaagt cgcgcagcca 2640agaccacatt caaaaaagct
attttccgaa gttatcttga tgatactttc cagacaccta 2700gcactggagg
agaatatgaa aagcatcttg gtatactggg tcctatcatt agggctgagg
2760tggatgatgt aatcgaagtt cagttcagaa atttggcctc cagaccatac
tcacttcatg 2820ctcatggcct tctctatgag aaatcttctg aaggcagaag
ctatgatgac aagtctcctg 2880aattgttcaa aaaggatgat gctatcatgc
caaacggcac atacacatat gtctggcaag 2940tccctccacg gtcaggacca
acagacaata cagaaaaatg taaatcatgg gcctattact 3000ctggtgtaaa
tccggaaaaa gatattcact ctggcttaat tggacctatt ttgatctgcc
3060agaaaggcat gattgacaag tacaacagga caatagacat aagggaattt
gtcttgtttt 3120ttatggtctt tgatgaggag aaaagctggt actttccaaa
atctgacaaa agcactcgtg 3180cagagaaact tataggagtc caatctcgcc
acacatttcc tgcaattaat gggatccctt 3240atcagctgca aggcttgacg
atgtacaaag atgagaatgt ccactggcat ttgctgaaca 3300tgggtgggcc
caaagatatc catgttgtta attttcatgg tcagacattc actgaagagg
3360gaagggaaga taatcaactt ggagtccttc ctcttcttcc tggtacattc
gcctccatca 3420aaatgaaacc atccaaaatt ggcacatggc ttttagaaac
agaagttggt gaaaatcagg 3480aaagaggaat gcaggctctc tttactgtca
ttgacaaaga ttgtaaatta ccaatgggac 3540tggcaagtgg gataatacaa
gactcacaga tcagtgcttc aggtcatgtt ggatattggg 3600agcctaagct
agcaagactg aataatactg gaaaatataa tgcttggagc atcataaaga
3660aggaacatga acatccgtgg atccagatag acctacaaag acaagttgtc
atcacaggca 3720ttcagaccca aggagccatg caactactga aacatttgta
tactgtggaa tattttttta 3780cctacagcaa agatgggcaa aactggatta
cttttaaagg aagacattcc gaaacacaaa 3840tgcattttga gggtaattca
gatggcacca cagtaaaaga aaaccacatt gatcctccta 3900ttattgccag
atatattagg ctgcatccaa ccaagttcca caacagacct actttccgca
3960ttgaactgtt aggttgtgaa gttgaaggct gctcagtgcc attgggaatg
gaaagtgggg 4020ctatcaagaa ttcagagatt acagcctctt cttataagaa
gacttggtgg agttcatggg 4080aaccatccct tgcacgactc aatctgaaag
gacgaacaaa tgcttggcaa ccaaaggtaa 4140acaacaaaga tcaatggcta
caaattgacc tgcaacatct tacaaaaata acaagcataa 4200taactcaagg
agccacatca atgactacat caatgtatgt gaaaacattc tccatccatt
4260atactgatga caattcaaca tggaagcctt atttggatgt tcgcacttcc
atggaaaagg 4320ttttcacagg aaatattaac agtgatggtc atgtcaaaca
ttttttcaaa ccccctatat 4380tgtccaggtt cattcgtatc atccctaaaa
catggaatca atatattgca ctccggatag 4440aattgtttgg ttgtgaagtt
ttttaaggct tggacagaag actatcaaat caagcaactt 4500caatgtttca
agttttctta ttactaactc tgctttttaa aaggaaacaa aaacaaaagc
4560ataataaaac tgtcttagca taaaaaagct atccttctca attttcagct
atagctttca 4620aatagctttg aaaatatcaa tcaaaatatc ataactgaag
tgactttaca atgattaatt 4680ctagtgccac tttaatcatg actgtaatcc
taatacataa accttatttt ttttgcc 473764383DNAPseudonaja textilis
6atgggaagat acagtgtgag ccctgtcccc aaatgtcttc tactgatgtt cctgggttgg
60tcagggctga agtattacca agtgaatgca gctcagctca gggagtacca tatagctgct
120cagctggaag actgggatta caacccccaa cctgaggagc tatccagatt
atcagagtca 180gatcttacgt ttaaaaaaat tgtctataga gaatatgaac
tagatttcaa acaagaggag 240ccaagagatg cgctctcagg gctcctaggg
ccaacactac gtggagaagt gggagacagc 300ctcataattt atttcaagaa
ttttgctact cagcctgtga gcattcaccc gcagagtgcc 360gtgtacaaca
aatggtcaga aggttcttca tattctgatg gaacatcaga tgtggaaaga
420ctggatgatg ctgtgcctcc aggccagtcg ttcaagtatg tgtggaatat
cactgcagaa 480attgggccaa agaaagctga tcctccctgt ctcacttatg
cgtactactc acatgtaaac 540atggtgcgag actttaattc tggtctcatt
ggtgctttgc tgatatgtaa agaaggaagc 600ctgaatgcaa atggttcaca
aaaattcttc aacagagaat atgtgctgat gttttctgtg 660tttgatgaaa
gcaagaactg gtacagaaag ccctcactac agtacacaat taatgggttt
720gccaatggaa cattgcctga tgttcaggct tgtgcttatg atcatattag
ctggcatttg 780ataggaatga gttccagtcc tgagatcttc tctgttcact
tcaatggaca aaccttggaa 840caaaaccatt acaaagtgtc aaccatcaac
cttgtcggag gtgcctcagt aacagccgac 900atgtcagtga gcaggacagg
aaaatggcta atatcttctc tggttgcaaa gcatctacaa 960gctgggatgt
atggttatct aaatatcaaa gactgtggaa atccagatac tttaacaaga
1020aagttatcct ttagagaact gatgaagatt aagaactggg aatatttcat
tgctgcagaa 1080gaaatcacct gggattatgc tccagaaatt cctagcagtg
ttgacagaag atacaaagct 1140cagtatctgg ataatttttc aaattttatt
ggcaagaaat acaaaaaggc agttttcagg 1200caatatgaag acggcaattt
cactaaaccg acctatgcca tttggcccaa agaacgtgga 1260attctgggcc
ccgttatcaa agctaaagtc agagacacag taacaattgt attcaaaaat
1320ctggccagtc gaccttacag catttatgtg catggagttt ccgtttcaaa
agatgcagaa 1380ggagctattt atccttcaga tcccaaagag aatataactc
atggcaaagc agttgaacca 1440ggacaggtct acacatataa atggactgtg
ctggatacag atgaacctac agtaaaggat 1500tctgagtgca ttactaaatt
atatcatagt gctgtggaca tgacaagaga tattgcttca 1560ggacttattg
ggccacttct ggtttgtaaa cacaaggcac tcagcgtcaa gggggtacag
1620aataaagctg atgtggaaca gcatgcagtc ttcgcagtgt ttgatgaaaa
caagagctgg 1680tacttggaag acaatatcaa gaaatactgc agcaatcctt
ccgctgttaa gaaagatgac 1740cctaaatttt acaagtccaa tgttatgtac
acactcaatg gctatgcatc agatagaaca 1800gaggttttga ggtttcatca
gtctgaagtt gttcaatggc acctcaccag cgtaggtaca 1860gtggatgaga
ttgttccagt acatctttct ggtcacacct tcttatccaa gggaaaacat
1920caagatattt taaatctttt ccccatgagt ggtgaatctg ctactgtaac
aatggacaat 1980ctaggaacct ggcttctgtc atcatggggc tcctgtgaga
tgagcaatgg catgagattg 2040agatttttgg atgccaatta tgatgatgaa
gatgagggaa atgaagaaga ggaagaagat 2100gatggtgata tttttgccga
cattttcatt ccttcagaag tagtaaaaaa gaaagaagag 2160gttcccgtaa
attttgtacc agacccagaa tcggatgcgc tagcaaaaga attaggatta
2220atagatgacg agggtaatcc aataatacag ccacgcaggg aacagacaga
ggatgatgaa 2280gaacagctaa tgaaagcttc aatgcttggg cttcgatcat
ttaaggggtc agttgctgaa 2340gaagaattga aacacacagc tctagcttta
gaagaagatg cccatgcttc tgatcctcga 2400attgacagta atagtgcacg
taatcctgac gacatagctg gacgctacct gcgtactatc 2460aaccgtggaa
ataaaaggag gtactacatt gcagcagaag aagttttgtg ggactactca
2520ccgatcggaa aaagtcaagt gagaagtcgc gcagccaaga ccacattcaa
aaaagctatt 2580ttccgaagtt atcttgatga tactttccag acacctagca
ctggaggaga atatgaaaag 2640catcttggta tactgggtcc tatcattagg
gctgaggtgg atgatgtaat cgaaattcag 2700ttcaaaaatt tggcctctag
accatactca cttcatgctc atggccttct ctatgagaaa 2760tcttctgaag
gcagaagcta tgacgacaag tctcctgaat tgttcaaaaa ggatgatgct
2820atcatgccaa atggcacata cacatatgtc tggcaagtcc ctccacggtc
aggaccaaca 2880gacaatacag aaaaatgtaa atcatgggcc tattactctg
gtgtaaatcc ggaaaaagat 2940attcactctg gcttaattgg acctattttg
atctgccaga aaggcatgat tgacaagtac 3000aacaggacaa tagacataag
ggaatttgtc ttgtttttta tggtctttga tgaggagaaa 3060agctggtact
ttccaaaatc tgacaaaagc acttgtgaag agaaacttat aggagtccaa
3120tctctccaca catttcctgc aattaatggg atcccttatc agctgcaagg
cttgacgatg 3180tacaaagatg agaatgtcca ctggcatttg ctgaacatgg
gtgggcccaa agatatccat 3240gttgttaatt ttcatggtca gacattcact
gaagagggaa gggaagataa tcaacttgga 3300gtccttcctc ttcttcctgg
tacattcgcc tccatcaaaa tgaaaccatc caaaattggc 3360acatggcttt
tagaaacaga agttggtgaa aatcaggaaa gaggaatgca ggctctcttt
3420actgtcattg acaaagattg taaattacca atgggactgg caagtgggat
aatacaagac 3480tcacagatca gtgcttcagg tcatgttgga tattgggagc
ctaagctagc aagactgaat 3540aatactggaa aatataatgc ttggagcatc
ataaagaagg aacatgaaca tccgtggatc 3600cagatagacc tacaaagaca
agttgtcatc acaggcattc agacccaagg aaccgtgcaa 3660ctactgcaac
attcgtatac tgtggaatat tttgttacct acagcgaaga tgggcaaaac
3720tggattactt ttaaaggaag acattccgaa acacaaatgc attttgaggg
taattcagat 3780ggcaccacag taaaagaaaa ccacattgat cctcctatta
ttgccagata tattagactg 3840catccaacca agttctacaa cagacctact
ttccgcattg aactgttagg ttgtgaagtt 3900gaaggttgct cagtgccatt
gggaatggaa agtggggcta tcaagaattc agagattaca 3960gcctcttctt
ataagaagac ttggtggagt tcatgggaac catcccttgc acgactcaat
4020ctggaaggag gaacaaatgc ttggcaacca gaggtaaaca acaaagatca
atggttacaa 4080attgacctgc aacatcttac aaaaataaca agcataataa
ctcaaggagc cacatcaatg 4140actacatcaa tgtatgtgaa aacattctcc
atccattata ctgatgacaa ttcaacatgg 4200aagccttatt tggatgttcg
cacttccatg gaaaaggttt tcacaggaaa tattaacagt 4260gatggtcatg
tcaaacattt tttcaaaccc cctatattgt ccaggttcat tcgtatcatc
4320cctaaaacat ggaatcaata tattgcactc cggatagaat tgtttggttg
tgaagttttt 4380taa 438371459PRTPseudonaja textilis 7Met Gly Arg Tyr
Ser Val Ser Pro Val Pro Lys Cys Leu Leu Leu Met1 5 10 15Phe Leu Gly
Trp Ser Gly Leu Lys Tyr Tyr Glu Val Asn Ala Ala Gln 20 25 30Leu Arg
Glu Tyr His Ile Ala Ala Gln Leu Glu Asp Trp Asp Tyr Asn 35 40 45Pro
Gln Pro Glu Glu Leu Ser Arg Leu Ser Glu Ser Asp Leu Thr Phe 50 55
60Lys Lys Ile Val Tyr Arg Glu Tyr Glu Leu Asp Phe Lys Gln Glu Lys65
70 75 80Pro Arg Asp Glu Leu Ser Gly Leu Leu Gly Pro Thr Leu Arg Gly
Glu 85 90 95Val Gly Asp Ile Leu Ile Ile Tyr Phe Lys Asn Phe Ala Thr
Gln Pro 100 105 110Val Ser Ile His Pro Gln Ser Ala Val Tyr Asn Lys
Trp Ser Glu Gly 115 120 125Ser Ser Tyr Ser Asp Gly Thr Ser Asp Val
Glu Arg Leu Asp Asp Ala 130 135 140Val Pro Pro Gly Gln Ser Phe Lys
Tyr Val Trp Asn Ile Thr Ala Glu145 150 155 160Ile Gly Pro Lys Lys
Ala Asp Pro Pro Cys Leu Thr Tyr Ala Tyr Tyr 165 170 175Ser His Val
Asn Met Val Arg Asp Phe Asn Ser Gly Leu Ile Gly Ala 180
185 190Leu Leu Ile Cys Lys Glu Gly Ser Leu Asn Ala Asn Gly Ser Gln
Lys 195 200 205Phe Phe Asn Arg Glu Tyr Val Leu Met Phe Ser Val Phe
Asp Glu Ser 210 215 220Lys Asn Trp Tyr Arg Lys Pro Ser Leu Gln Tyr
Thr Ile Asn Gly Phe225 230 235 240Ala Asn Gly Thr Leu Pro Asp Val
Gln Ala Cys Ala Tyr Asp His Ile 245 250 255Ser Trp His Leu Ile Gly
Met Ser Ser Ser Pro Glu Ile Phe Ser Val 260 265 270His Phe Asn Gly
Gln Thr Leu Glu Gln Asn His Tyr Lys Val Ser Thr 275 280 285Ile Asn
Leu Val Gly Gly Ala Ser Val Thr Ala Asn Met Ser Val Ser 290 295
300Arg Thr Gly Lys Trp Leu Ile Ser Ser Leu Val Ala Lys His Leu
Gln305 310 315 320Ala Gly Met Tyr Gly Tyr Leu Asn Ile Lys Asp Cys
Gly Asn Pro Asp 325 330 335Thr Leu Thr Arg Lys Leu Ser Phe Arg Glu
Leu Arg Arg Ile Met Asn 340 345 350Trp Glu Tyr Phe Ile Ala Ala Glu
Glu Ile Thr Trp Asp Tyr Ala Pro 355 360 365Glu Ile Pro Ser Ser Val
Asp Arg Arg Tyr Lys Ala Gln Tyr Leu Asp 370 375 380Asn Phe Ser Asn
Phe Ile Gly Lys Lys Tyr Lys Lys Ala Val Phe Arg385 390 395 400Gln
Tyr Lys Asp Ser Asn Phe Thr Lys Pro Thr Tyr Ala Ile Trp Pro 405 410
415Lys Glu Arg Gly Ile Leu Gly Pro Val Ile Arg Ala Lys Val Arg Asp
420 425 430Thr Ile Ser Ile Val Phe Lys Asn Leu Ala Ser Arg Pro Tyr
Ser Ile 435 440 445Tyr Val His Gly Val Ser Val Ser Lys Asp Ala Glu
Gly Ala Ile Tyr 450 455 460Pro Ser Asp Pro Lys Glu Asn Ile Thr His
Gly Lys Ala Val Glu Pro465 470 475 480Gly Gln Val Tyr Thr Tyr Lys
Trp Thr Val Leu Asp Thr Asp Glu Pro 485 490 495Thr Val Lys Asp Ser
Glu Cys Ile Thr Lys Leu Tyr His Ser Ala Val 500 505 510Asp Met Thr
Arg Asp Ile Ala Ser Gly Leu Ile Gly Pro Leu Leu Val 515 520 525Cys
Lys His Lys Ala Leu Ser Val Lys Gly Val Gln Asn Lys Ala Asp 530 535
540Val Glu Gln His Ala Val Phe Ala Val Phe Asp Glu Asn Lys Ser
Trp545 550 555 560Tyr Leu Glu Asp Asn Ile Lys Lys Tyr Cys Ser Asn
Pro Ser Thr Val 565 570 575Lys Lys Asp Asp Pro Lys Phe Tyr Lys Ser
Asn Val Met Tyr Thr Leu 580 585 590Asn Gly Tyr Ala Ser Asp Arg Thr
Glu Val Leu Gly Phe His Gln Ser 595 600 605Glu Val Val Glu Trp His
Leu Thr Ser Val Gly Thr Val Asp Glu Ile 610 615 620Val Pro Val His
Leu Ser Gly His Thr Phe Leu Ser Lys Gly Lys His625 630 635 640Gln
Asp Ile Leu Asn Leu Phe Pro Met Ser Gly Glu Ser Ala Thr Val 645 650
655Thr Met Asp Asn Leu Gly Thr Trp Leu Leu Ser Ser Trp Gly Ser Cys
660 665 670Glu Met Ser Asn Gly Met Arg Leu Arg Phe Leu Asp Ala Asn
Tyr Asp 675 680 685Asp Glu Asp Glu Gly Asn Glu Glu Glu Glu Glu Asp
Asp Gly Asp Ile 690 695 700Phe Ala Asp Ile Phe Ile Pro Pro Glu Val
Val Lys Lys Lys Glu Lys705 710 715 720Asp Pro Val Asn Phe Val Ser
Asp Pro Glu Ser Asp Lys Ile Ala Lys 725 730 735Glu Leu Gly Leu Leu
Asp Asp Glu Asp Asn Gln Glu Glu Ser His Asn 740 745 750Val Gln Thr
Glu Asp Asp Glu Glu Gln Leu Met Ile Ala Thr Met Leu 755 760 765Gly
Phe Arg Ser Phe Lys Gly Ser Val Ala Glu Glu Glu Leu Asn Leu 770 775
780Thr Ala Leu Ala Leu Glu Glu Asp Ala His Ala Ser Asp Pro Arg
Ile785 790 795 800Asp Ser Asn Ser Ala Arg Asn Pro Asp Asp Ile Ala
Gly Arg Tyr Leu 805 810 815Arg Thr Ile Asn Arg Gly Asn Lys Arg Arg
Tyr Tyr Ile Ala Ala Glu 820 825 830Glu Val Leu Trp Asp Tyr Ser Pro
Ile Gly Lys Ser Gln Val Arg Ser 835 840 845Arg Ala Ala Lys Thr Thr
Phe Lys Lys Ala Ile Phe Arg Ser Tyr Leu 850 855 860Asp Asp Thr Phe
Gln Thr Pro Ser Thr Gly Gly Glu Tyr Glu Lys His865 870 875 880Leu
Gly Ile Leu Gly Pro Ile Ile Arg Ala Glu Val Asp Asp Val Ile 885 890
895Glu Val Gln Phe Arg Asn Leu Ala Ser Arg Pro Tyr Ser Leu His Ala
900 905 910His Gly Leu Leu Tyr Glu Lys Ser Ser Glu Gly Arg Ser Tyr
Asp Asp 915 920 925Lys Ser Pro Glu Leu Phe Lys Lys Asp Asp Ala Ile
Met Pro Asn Gly 930 935 940Thr Tyr Thr Tyr Val Trp Gln Val Pro Pro
Arg Ser Gly Pro Thr Asp945 950 955 960Asn Thr Glu Lys Cys Lys Ser
Trp Ala Tyr Tyr Ser Gly Val Asn Pro 965 970 975Glu Lys Asp Ile His
Ser Gly Leu Ile Gly Pro Ile Leu Ile Cys Gln 980 985 990Lys Gly Met
Ile Asp Lys Tyr Asn Arg Thr Ile Asp Ile Arg Glu Phe 995 1000
1005Val Leu Phe Phe Met Val Phe Asp Glu Glu Lys Ser Trp Tyr Phe
1010 1015 1020Pro Lys Ser Asp Lys Ser Thr Arg Ala Glu Lys Leu Ile
Gly Val 1025 1030 1035Gln Ser Arg His Thr Phe Pro Ala Ile Asn Gly
Ile Pro Tyr Gln 1040 1045 1050Leu Gln Gly Leu Thr Met Tyr Lys Asp
Glu Asn Val His Trp His 1055 1060 1065Leu Leu Asn Met Gly Gly Pro
Lys Asp Ile His Val Val Asn Phe 1070 1075 1080His Gly Gln Thr Phe
Thr Glu Glu Gly Arg Glu Asp Asn Gln Leu 1085 1090 1095Gly Val Leu
Pro Leu Leu Pro Gly Thr Phe Ala Ser Ile Lys Met 1100 1105 1110Lys
Pro Ser Lys Ile Gly Thr Trp Leu Leu Glu Thr Glu Val Gly 1115 1120
1125Glu Asn Gln Glu Arg Gly Met Gln Ala Leu Phe Thr Val Ile Asp
1130 1135 1140Lys Asp Cys Lys Leu Pro Met Gly Leu Ala Ser Gly Ile
Ile Gln 1145 1150 1155Asp Ser Gln Ile Ser Ala Ser Gly His Val Gly
Tyr Trp Glu Pro 1160 1165 1170Lys Leu Ala Arg Leu Asn Asn Thr Gly
Lys Tyr Asn Ala Trp Ser 1175 1180 1185Ile Ile Lys Lys Glu His Glu
His Pro Trp Ile Gln Ile Asp Leu 1190 1195 1200Gln Arg Gln Val Val
Ile Thr Gly Ile Gln Thr Gln Gly Ala Met 1205 1210 1215Gln Leu Leu
Lys His Leu Tyr Thr Val Glu Tyr Phe Phe Thr Tyr 1220 1225 1230Ser
Lys Asp Gly Gln Asn Trp Ile Thr Phe Lys Gly Arg His Ser 1235 1240
1245Glu Thr Gln Met His Phe Glu Gly Asn Ser Asp Gly Thr Thr Val
1250 1255 1260Lys Glu Asn His Ile Asp Pro Pro Ile Ile Ala Arg Tyr
Ile Arg 1265 1270 1275Leu His Pro Thr Lys Phe His Asn Arg Pro Thr
Phe Arg Ile Glu 1280 1285 1290Leu Leu Gly Cys Glu Val Glu Gly Cys
Ser Val Pro Leu Gly Met 1295 1300 1305Glu Ser Gly Ala Ile Lys Asn
Ser Glu Ile Thr Ala Ser Ser Tyr 1310 1315 1320Lys Lys Thr Trp Trp
Ser Ser Trp Glu Pro Ser Leu Ala Arg Leu 1325 1330 1335Asn Leu Lys
Gly Arg Thr Asn Ala Trp Gln Pro Lys Val Asn Asn 1340 1345 1350Lys
Asp Gln Trp Leu Gln Ile Asp Leu Gln His Leu Thr Lys Ile 1355 1360
1365Thr Ser Ile Ile Thr Gln Gly Ala Thr Ser Met Thr Thr Ser Met
1370 1375 1380Tyr Val Lys Thr Phe Ser Ile His Tyr Thr Asp Asp Asn
Ser Thr 1385 1390 1395Trp Lys Pro Tyr Leu Asp Val Arg Thr Ser Met
Glu Lys Val Phe 1400 1405 1410Thr Gly Asn Ile Asn Ser Asp Gly His
Val Lys His Phe Phe Lys 1415 1420 1425Pro Pro Ile Leu Ser Arg Phe
Ile Arg Ile Ile Pro Lys Thr Trp 1430 1435 1440Asn Gln Tyr Ile Ala
Leu Arg Ile Glu Leu Phe Gly Cys Glu Val 1445 1450
1455Phe81460PRTPseudonaja textilis 8Met Gly Arg Tyr Ser Val Ser Pro
Val Pro Lys Cys Leu Leu Leu Met1 5 10 15Phe Leu Gly Trp Ser Gly Leu
Lys Tyr Tyr Gln Val Asn Ala Ala Gln 20 25 30Leu Arg Glu Tyr His Ile
Ala Ala Gln Leu Glu Asp Trp Asp Tyr Asn 35 40 45Pro Gln Pro Glu Glu
Leu Ser Arg Leu Ser Glu Ser Asp Leu Thr Phe 50 55 60Lys Lys Ile Val
Tyr Arg Glu Tyr Glu Leu Asp Phe Lys Gln Glu Glu65 70 75 80Pro Arg
Asp Ala Leu Ser Gly Leu Leu Gly Pro Thr Leu Arg Gly Glu 85 90 95Val
Gly Asp Ser Leu Ile Ile Tyr Phe Lys Asn Phe Ala Thr Gln Pro 100 105
110Val Ser Ile His Pro Gln Ser Ala Val Tyr Asn Lys Trp Ser Glu Gly
115 120 125Ser Ser Tyr Ser Asp Gly Thr Ser Asp Val Glu Arg Leu Asp
Asp Ala 130 135 140Val Pro Pro Gly Gln Ser Phe Lys Tyr Val Trp Asn
Ile Thr Ala Glu145 150 155 160Ile Gly Pro Lys Lys Ala Asp Pro Pro
Cys Leu Thr Tyr Ala Tyr Tyr 165 170 175Ser His Val Asn Met Val Arg
Asp Phe Asn Ser Gly Leu Ile Gly Ala 180 185 190Leu Leu Ile Cys Lys
Glu Gly Ser Leu Asn Ala Asn Gly Ser Gln Lys 195 200 205Phe Phe Asn
Arg Glu Tyr Val Leu Met Phe Ser Val Phe Asp Glu Ser 210 215 220Lys
Asn Trp Tyr Arg Lys Pro Ser Leu Gln Tyr Thr Ile Asn Gly Phe225 230
235 240Ala Asn Gly Thr Leu Pro Asp Val Gln Ala Cys Ala Tyr Asp His
Ile 245 250 255Ser Trp His Leu Ile Gly Met Ser Ser Ser Pro Glu Ile
Phe Ser Val 260 265 270His Phe Asn Gly Gln Thr Leu Glu Gln Asn His
Tyr Lys Val Ser Thr 275 280 285Ile Asn Leu Val Gly Gly Ala Ser Val
Thr Ala Asp Met Ser Val Ser 290 295 300Arg Thr Gly Lys Trp Leu Ile
Ser Ser Leu Val Ala Lys His Leu Gln305 310 315 320Ala Gly Met Tyr
Gly Tyr Leu Asn Ile Lys Asp Cys Gly Asn Pro Asp 325 330 335Thr Leu
Thr Arg Lys Leu Ser Phe Arg Glu Leu Met Lys Ile Lys Asn 340 345
350Trp Glu Tyr Phe Ile Ala Ala Glu Glu Ile Thr Trp Asp Tyr Ala Pro
355 360 365Glu Ile Pro Ser Ser Val Asp Arg Arg Tyr Lys Ala Gln Tyr
Leu Asp 370 375 380Asn Phe Ser Asn Phe Ile Gly Lys Lys Tyr Lys Lys
Ala Val Phe Arg385 390 395 400Gln Tyr Glu Asp Gly Asn Phe Thr Lys
Pro Thr Tyr Ala Ile Trp Pro 405 410 415Lys Glu Arg Gly Ile Leu Gly
Pro Val Ile Lys Ala Lys Val Arg Asp 420 425 430Thr Val Thr Ile Val
Phe Lys Asn Leu Ala Ser Arg Pro Tyr Ser Ile 435 440 445Tyr Val His
Gly Val Ser Val Ser Lys Asp Ala Glu Gly Ala Ile Tyr 450 455 460Pro
Ser Asp Pro Lys Glu Asn Ile Thr His Gly Lys Ala Val Glu Pro465 470
475 480Gly Gln Val Tyr Thr Tyr Lys Trp Thr Val Leu Asp Thr Asp Glu
Pro 485 490 495Thr Val Lys Asp Ser Glu Cys Ile Thr Lys Leu Tyr His
Ser Ala Val 500 505 510Asp Met Thr Arg Asp Ile Ala Ser Gly Leu Ile
Gly Pro Leu Leu Val 515 520 525Cys Lys His Lys Ala Leu Ser Val Lys
Gly Val Gln Asn Lys Ala Asp 530 535 540Val Glu Gln His Ala Val Phe
Ala Val Phe Asp Glu Asn Lys Ser Trp545 550 555 560Tyr Leu Glu Asp
Asn Ile Lys Lys Tyr Cys Ser Asn Pro Ser Ala Val 565 570 575Lys Lys
Asp Asp Pro Lys Phe Tyr Lys Ser Asn Val Met Tyr Thr Leu 580 585
590Asn Gly Tyr Ala Ser Asp Arg Thr Glu Val Leu Arg Phe His Gln Ser
595 600 605Glu Val Val Gln Trp His Leu Thr Ser Val Gly Thr Val Asp
Glu Ile 610 615 620Val Pro Val His Leu Ser Gly His Thr Phe Leu Ser
Lys Gly Lys His625 630 635 640Gln Asp Ile Leu Asn Leu Phe Pro Met
Ser Gly Glu Ser Ala Thr Val 645 650 655Thr Met Asp Asn Leu Gly Thr
Trp Leu Leu Ser Ser Trp Gly Ser Cys 660 665 670Glu Met Ser Asn Gly
Met Arg Leu Arg Phe Leu Asp Ala Asn Tyr Asp 675 680 685Asp Glu Asp
Glu Gly Asn Glu Glu Glu Glu Glu Asp Asp Gly Asp Ile 690 695 700Phe
Ala Asp Ile Phe Ile Pro Ser Glu Val Val Lys Lys Lys Glu Glu705 710
715 720Val Pro Val Asn Phe Val Pro Asp Pro Glu Ser Asp Ala Leu Ala
Lys 725 730 735Glu Leu Gly Leu Ile Asp Asp Glu Gly Asn Pro Ile Ile
Gln Pro Arg 740 745 750Arg Glu Gln Thr Glu Asp Asp Glu Glu Gln Leu
Met Lys Ala Ser Met 755 760 765Leu Gly Leu Arg Ser Phe Lys Gly Ser
Val Ala Glu Glu Glu Leu Lys 770 775 780His Thr Ala Leu Ala Leu Glu
Glu Asp Ala His Ala Ser Asp Pro Arg785 790 795 800Ile Asp Ser Asn
Ser Ala Arg Asn Pro Asp Asp Ile Ala Gly Arg Tyr 805 810 815Leu Arg
Thr Ile Asn Arg Gly Asn Lys Arg Arg Tyr Tyr Ile Ala Ala 820 825
830Glu Glu Val Leu Trp Asp Tyr Ser Pro Ile Gly Lys Ser Gln Val Arg
835 840 845Ser Arg Ala Ala Lys Thr Thr Phe Lys Lys Ala Ile Phe Arg
Ser Tyr 850 855 860Leu Asp Asp Thr Phe Gln Thr Pro Ser Thr Gly Gly
Glu Tyr Glu Lys865 870 875 880His Leu Gly Ile Leu Gly Pro Ile Ile
Arg Ala Glu Val Asp Asp Val 885 890 895Ile Glu Ile Gln Phe Lys Asn
Leu Ala Ser Arg Pro Tyr Ser Leu His 900 905 910Ala His Gly Leu Leu
Tyr Glu Lys Ser Ser Glu Gly Arg Ser Tyr Asp 915 920 925Asp Lys Ser
Pro Glu Leu Phe Lys Lys Asp Asp Ala Ile Met Pro Asn 930 935 940Gly
Thr Tyr Thr Tyr Val Trp Gln Val Pro Pro Arg Ser Gly Pro Thr945 950
955 960Asp Asn Thr Glu Lys Cys Lys Ser Trp Ala Tyr Tyr Ser Gly Val
Asn 965 970 975Pro Glu Lys Asp Ile His Ser Gly Leu Ile Gly Pro Ile
Leu Ile Cys 980 985 990Gln Lys Gly Met Ile Asp Lys Tyr Asn Arg Thr
Ile Asp Ile Arg Glu 995 1000 1005Phe Val Leu Phe Phe Met Val Phe
Asp Glu Glu Lys Ser Trp Tyr 1010 1015 1020Phe Pro Lys Ser Asp Lys
Ser Thr Cys Glu Glu Lys Leu Ile Gly 1025 1030 1035Val Gln Ser Leu
His Thr Phe Pro Ala Ile Asn Gly Ile Pro Tyr 1040 1045 1050Gln Leu
Gln Gly Leu Thr Met Tyr Lys Asp Glu Asn Val His Trp 1055 1060
1065His Leu Leu Asn Met Gly Gly Pro Lys Asp Ile His Val Val Asn
1070 1075 1080Phe His Gly Gln Thr Phe Thr Glu Glu Gly Arg Glu Asp
Asn Gln 1085 1090 1095Leu Gly Val Leu Pro Leu Leu Pro Gly Thr Phe
Ala Ser Ile Lys 1100 1105 1110Met Lys Pro Ser Lys Ile Gly Thr Trp
Leu Leu Glu Thr Glu Val 1115 1120 1125Gly Glu Asn Gln Glu Arg Gly
Met Gln Ala Leu Phe Thr Val Ile 1130 1135 1140Asp Lys Asp Cys Lys
Leu Pro Met Gly Leu Ala Ser Gly Ile Ile 1145 1150 1155Gln Asp Ser
Gln Ile Ser Ala Ser Gly His Val Gly Tyr Trp Glu 1160 1165 1170Pro
Lys Leu Ala Arg Leu Asn Asn Thr Gly Lys Tyr Asn Ala Trp 1175 1180
1185Ser Ile Ile Lys Lys Glu
His Glu His Pro Trp Ile Gln Ile Asp 1190 1195 1200Leu Gln Arg Gln
Val Val Ile Thr Gly Ile Gln Thr Gln Gly Thr 1205 1210 1215Val Gln
Leu Leu Gln His Ser Tyr Thr Val Glu Tyr Phe Val Thr 1220 1225
1230Tyr Ser Glu Asp Gly Gln Asn Trp Ile Thr Phe Lys Gly Arg His
1235 1240 1245Ser Glu Thr Gln Met His Phe Glu Gly Asn Ser Asp Gly
Thr Thr 1250 1255 1260Val Lys Glu Asn His Ile Asp Pro Pro Ile Ile
Ala Arg Tyr Ile 1265 1270 1275Arg Leu His Pro Thr Lys Phe Tyr Asn
Arg Pro Thr Phe Arg Ile 1280 1285 1290Glu Leu Leu Gly Cys Glu Val
Glu Gly Cys Ser Val Pro Leu Gly 1295 1300 1305Met Glu Ser Gly Ala
Ile Lys Asn Ser Glu Ile Thr Ala Ser Ser 1310 1315 1320Tyr Lys Lys
Thr Trp Trp Ser Ser Trp Glu Pro Ser Leu Ala Arg 1325 1330 1335Leu
Asn Leu Glu Gly Gly Thr Asn Ala Trp Gln Pro Glu Val Asn 1340 1345
1350Asn Lys Asp Gln Trp Leu Gln Ile Asp Leu Gln His Leu Thr Lys
1355 1360 1365Ile Thr Ser Ile Ile Thr Gln Gly Ala Thr Ser Met Thr
Thr Ser 1370 1375 1380Met Tyr Val Lys Thr Phe Ser Ile His Tyr Thr
Asp Asp Asn Ser 1385 1390 1395Thr Trp Lys Pro Tyr Leu Asp Val Arg
Thr Ser Met Glu Lys Val 1400 1405 1410Phe Thr Gly Asn Ile Asn Ser
Asp Gly His Val Lys His Phe Phe 1415 1420 1425Lys Pro Pro Ile Leu
Ser Arg Phe Ile Arg Ile Ile Pro Lys Thr 1430 1435 1440Trp Asn Gln
Tyr Ile Ala Leu Arg Ile Glu Leu Phe Gly Cys Glu 1445 1450 1455Val
Phe 146094380DNAOxyuranus scutellatus 9atgggaagat acagtgtgag
ccctgtcccc aaatgtcttc tactgatgtt cctgggttgg 60tcagggctga agtattacca
agtgaatgca gctcagctca gggagtaccg tttagctgct 120cagctggaag
actgggatta caacccccaa cctgaggagc tatccagatt atcagagtca
180gatcttacgt ttaaaaaaat tgtctataga gaatatgaac tagatttcaa
acaagagaag 240ccaagagatg agctctcagg gctcctaggg ccaacactac
gtggagaagt gggagacagc 300ctcataattt atttcaagaa ttttgctact
cagcctgtga gcattcaccc gcagagtgcc 360gtgtacaaca aatggtcaga
aggttcttca tattctgatg gaacatcaga tgtggaaaga 420ctggatgatg
ctgtgcctcc aggccagtcg ttcaagtatg tgtggaatat cactgcagaa
480attgggccaa agaaagctga tcctccctgt ctcacttatg cgtactactc
acatgtaaac 540atggtgcgag actttaattc tggtctcatt ggtgctttgc
tgatatgtaa agaaggaagc 600ctgaatgcag atggtgcaca aaaattcttc
aacagagaat atgtgctgat gttttctgtg 660tttgatgaaa gcaagaactg
gtacagaaag ccctcattac agtacacaat taatgggttt 720gccaatggaa
cattgcctga tgttcaggct tgtgcttatg atcatattag ctggcatttg
780ataggaatga gttccagtcc tgagatcttc tctgttcact tcaatggaca
aaccttggaa 840caaaaccatt acaaagtgtc aaccatcaac cttgtcggag
gtgcctcagt aacagccaac 900atgtcagtga gcaggacagg aaaatggcta
atatcttctc tggttgcaaa gcatctacaa 960gctgggatgt atggttatct
aaatatcaaa gactgtggaa atccagatac tttaacaaga 1020aagttatcct
ttagagaacg gaggaggatt atgaaatggg aatatttcat tgctgcagaa
1080gaaatcacct gggattatgc tccagaaatt cctagcagtg ttgacagaag
atacaaagct 1140cagtatctgg ataatttttc aaattttatt ggcaagaaat
acaaaaaggc agttttcagg 1200caatatgaag acagcaattt cactaaaccg
acctatgcca tttggcccaa agaacgtgga 1260attctgggcc ccgttatcaa
agctaaagtc agagacacag taacaattgt attcaaaaat 1320ctggccagtc
gaccttacag catttatgtg catggagttt ccgtttcaaa agatgcagaa
1380ggagctgttt atccttcaga tcccaaagag aatataactc atggcaaagc
agttgaacca 1440ggacaggtct acacatataa atggactgtg ctggatacag
atgaacctac agtaaaggat 1500tctgagtgca ttactaaatt atatcatagt
gctgtggaca tgacaagaga tattgcttca 1560ggacttattg ggccacctct
ggtttgtaaa cgcaaggcac tcagcatcag gggggtacag 1620aataaagctg
atgtggaaca gcatgcagtc ttcgcagtgt ttgatgaaaa caagagctgg
1680tacttggaag acaatatcaa gaaatactgc agcaatcctt ccagtgttaa
gaaagatgac 1740cctaaatttt acaagtccaa tgttatgtac acactcaatg
gctatgcatc agatagaaca 1800gaggtttggg ggtttcatca gtctgaagtt
gttgaatggc acctcaccag cgtaggtaca 1860gtggatgaga ttgttccagt
acatctttct ggtcacacct tcttatccaa gggaaaacat 1920caagatattt
taaatctttt tcccatgagt ggtgaatcgg ctactgtaac aatggacaat
1980ctaggaacct ggcttctgtc atcatggggc tcctgtgaga tgagcaatgg
catgagattg 2040agatttttgg atgccaatta tgatgatgaa gatgagggaa
atgaagaaga ggaagaagat 2100gatggtgata tttttgccga cattttcaat
cctccagaag tagtaataaa gaaagaagag 2160gttcccgtaa attttgtacc
agacccagaa tcggatgcgc tagcaaaaga attaggatta 2220tttgatgacg
aggataatcc aaaacagtca cgcagtgaac agacagagga tgatgaagaa
2280cagctaatga tagcttcaat gcttgggctt cgatcattta aggggtcagt
tgctgaagaa 2340gaattgaaac acacagctct agctttagaa gaagatgccc
atgcttctga tcctcgaatt 2400gacagtaata gtgcacataa ttctgacgac
atagctggac gctacctgcg tactatcaac 2460cgcagaaata aaaggaggta
ctacattgca gcagaagaag ttttgtggga ctactcaccg 2520atcggaaaaa
gtcaagtgag aagtctccca gccaagacca cattcaaaaa agctattttc
2580cgaagttatc ttgatgatac tttccagaca cctagcactg gaggagaata
tgaaaagcat 2640cttggtatac tgggtcctat cattagggct gaggtggatg
atgtaatcga agttcagttc 2700agaaatttgg cctctagacc atactcactt
catgctcatg gccttctcta tgagaaatct 2760tctgaaggca gaagctatga
cgacaactct cctgaattgt tcaaaaaaga tgatgctatc 2820atgccaaacg
gcacatacac atatgtctgg caagtccctc cacggtcagg accaacagac
2880aatacagaaa aatgtaaatc atgggcctat tactctggtg taaatccgga
aaaagatatt 2940cactctggct taattggacc tattttgatc tgccagaaag
gcatgattga caagtacaac 3000aggacaatag acataaggga atttgtcttg
ttttttatgg tctttgatga ggagaaaagc 3060tggtactttc caaaatctga
caaaagcact tgtgaagaga aacttatagg agtccaatct 3120cgccacacat
ttcctgcaat taatgggatc ccttatcagc tgcaaggctt gatgatgtac
3180aaagatgaga atgtccactg gcatttgctg aacatgggtg ggcccaaaga
tgtccatgtt 3240gttaattttc atggtcagac attcactgaa gagggaaggg
aagataatca acttggagtc 3300cttcctcttc ttcctggtac attcgcctcc
atcaaaatga aaccatccaa aattggcaca 3360tggcttttag aaacagaagt
tggtgaaaat caggaaagag gaatgcaggc tctctttact 3420gtcattgaca
aagattgtaa attaccaatg ggactggcaa gtgggataat acaagactca
3480cagatcagtg cttcaggtca tgttggatat tgggagccta agctagcaag
actgaataat 3540actggaatgt ttaatgcttg gagcatcata aagaaggaac
atgaacatcc gtggatccag 3600atagacctac aaagacaagt tgtcatcaca
ggcattcaga cccagggaac cgtgcaacta 3660ctgaaacatt cgtatactgt
ggaatatttt gttacctaca gcaaagatgg gcaaaactgg 3720attactttta
aaggaagaca ttccaaaaca caaatgcatt ttgagggtaa ttcagatggc
3780accacagtaa aagaaaacca cattgatcct cctattattg ccagatatat
taggctgcat 3840ccaaccaagt tctacaacac acctactttc cgcattgaac
tgttaggttg tgaagttgaa 3900ggttgctcag tgccattggg aatggaaagt
ggggctatca aggattcaga gattacagcc 3960tcttcttata aaaagacttg
gtggagttca tgggaaccat tccttgcacg actcaatctg 4020aaaggacgaa
caaatgcttg gcaaccaaag gtaaacaaca aagatcaatg gctacaaatt
4080gacctgcaac atcttacaaa aataacaagc ataataactc aaggagccac
atcaatgact 4140acatcaatgt atgtgaaaac attctccatc cattatactg
atgacaattc aacatggaag 4200ccttatttgg atgttcgcac ttccatggaa
aaggttttca caggaaatat taacagtgat 4260ggtcatgtca aacatttttt
caacccccct atattgtcca ggttcattcg tatcatccct 4320aaaacatgga
atcaatatat tgcactccgg atagaattgt ttggttgtga agttttttaa
4380104675DNAOxyuranus scutellatus scutellatus 10atgggaagat
acagtgtgag ccctgtcccc aaatgtcttc tactgatgtt cctgggttgg 60tcagggctga
agtattacca agtgaatgca gctcagctca gggagtaccg tatagctgct
120cagctggaag actgggatta caacccccaa cctgaggagc tatccagatt
atcagagtca 180gatcttacgt ttaaaaaaat tgtctataga gaatatgaac
tagatttcaa acaagagaag 240ccaagagatg agctctcagg gctcctaggg
ccaacactac gtggagaagt gggagacagc 300ctcataattt atttcaagaa
ttttgctact cagcctgtga gcattcaccc gcagagtgcc 360gtgtacaaca
aatggtcaga aggttcttca tattctgatg gaacatcaga tgtggaaaga
420ctggatgatg ctgtgcctcc aggccagtcg ttcaagtatg tgtggaatat
cactgcagaa 480attgggccaa agaaagctga tcctccctgt ctcacttatg
cgtactactc acatgtaaac 540atggtgcgag actttaattc tggtctcatt
ggtgctttgc tgatatgtaa agaaggaagc 600ctgaatgcag atggtgcaca
aaaattcttc aacagagaat atgtgctgat gttttctgtg 660tttgatgaaa
gcaagaactg gtacagaaag ccctcattac agtacacaat taatgggttt
720gccaatggaa cattgcctga tgttcaggct tgtgcttatg atcatattag
ctggcatttg 780ataggaatga gttccagtcc tgagatcttc tctgttcact
tcaatggaca aaccttggaa 840caaaaccatt acaaagtgtc aaccatcaac
cttgtcggag gtgcctcagt aacagccaac 900atgtcagtga gcaggacagg
aaaatggcta atatcttctc tggttgcaaa gcatctacaa 960gctgggatgt
atggttatct taatatcaaa gactgtggaa atccagatac tttaacaaga
1020aagttatcct ttagagaatg gaggaggatt atgaaatggg aatatttcat
tgctgcagaa 1080gaaatcacct gggattatgc tccagaaatt cctagcagtg
ttgacagaag atacaaagct 1140cagtatctgg atttttcaaa ttttattggc
aagaaataca aaaaggcagt tttcaggcaa 1200tatgaagaca gcaatttcac
taaaccgacc tatgccattt ggcccaaaga acgtggaatt 1260ctgggccccg
ttatcaaagc taaagtcaga gacacagtaa caattgtatt caaaaatctg
1320gccagtcgac cttacagcat ttatgtgcat ggagtttccg tttcaaaaga
tgcagaagga 1380gctgtttatc cttcagatcc caaagagaat ataactcatg
gcaaagcagt tgaaccagga 1440caggtctaca catataaatg gactgtgctg
gatacagatg aacctacagt aaaggattct 1500gagtgcatta ctaaattata
tcatagtgct gtggacatga caagagatat tgcttcagga 1560cttattgggc
cacttctggt ttgtaaacgc aaggcactca gcatcagggg ggtacagaat
1620aaagctgatg tggaacagca tgcagtcttc gcagtgtttg atgaaaacaa
gagctggtac 1680ttggaagaca atatcaagaa atactgcagc aatccttcca
gtgttaagaa agatgaccct 1740aaattttaca agtccaatgt tatgtacaca
ctcaatggct atgcatcaga tagaacagag 1800gtttgggggt ttcatcagtc
tgaagttgtt gaatggcacc tcaccagcgt aggtacagtg 1860gatgagattg
ttccagtaca tctttctggt cacaccttct tatccaaggg aaaacatcaa
1920gatattttaa atctttttcc catgagtggt gaatccgcta ctgtaacaat
ggacaatcta 1980ggaacctggc ttctgtcatc atggggctcc tgtgagatga
gcaatggcat gagattgaga 2040tttttggatg ccaattatga tgatgaagat
gagggaaatg aagaagagga agaagatgat 2100ggtgatattt ttgccgacat
tttcaatcct ccagaagtag taataaagaa agaagaggtt 2160cccgtaaatt
ttgtaccaga cccagaatcg gatgcgctag caaaagaatt aggattattt
2220gatgacgagg ataatccaaa acagtcacgc agtgaacaga cagaggatga
tgaagaacag 2280ctaatgatag cttcaatgct tgggcttcga tcatttaagg
ggtcagttgc tgaagaagaa 2340ttgaaacaca cagctctagc tttagaagaa
gatgcccatg cttctgatcc tcgaattgac 2400agtaatagtg cacataattc
tgacgacata gctggacgct acctgcgtac tatctaccgc 2460agaaataaaa
ggaggtacta cattgcagca gaagaagttt tgtgggacta ctcaccgatc
2520ggaaaaagtc aagtgagaag tctcccagcc aagaccacat tcaaaaaagc
tattttccga 2580agttatcttg atgatacttt ccagacacct agcactggag
gagaatatga aaagcatctt 2640ggtatactgg gtcctatcat tagggctgag
gtggatgatg taatcgaagt tcagttcaga 2700aatttggcct ctagaccata
ctcacttcat gctcatggcc ttctctatga gaaatcttct 2760gaaggcagaa
gctatgacga caactctcct gaattgttca aaaaggatga tgctatcatg
2820ccaaacggca catacacata tgtctggcaa gtccctccac ggtcaggacc
aacagacaat 2880acagaaaaat gtaaatcatg ggcctattac tctggtgtaa
atccggaaaa agatattcac 2940tctgggctta ttggacctat tttgatctgc
cagaaaggca tgattgacaa gtacaacagg 3000acaatagaca taagggaatt
tgtcttgttt tttatggtct ttgatgagga gaaaagctgg 3060tactttccaa
aatctgacaa aagcacttgt gaagagaaac ttataggagt ccaatctcgc
3120cacacatttc ctgcaattaa tgggatccct tatcagctgc aaggcttgat
gatgtacaaa 3180gatgagaatg tccactggca tttgctgaac atgggtgggc
ccaaagatgt ccatgttgtt 3240aattttcatg gtcagacatt cactgaagag
ggaagggaag ataatcaact tggagtcctt 3300cctcttcttc ctggtacatt
cgcctccatc aaaatgaaac catccaaaat tggcacatgg 3360cttttagaaa
cagaagttgg tgaaaatcag gaaagaggaa tgcaggctct ctttactgtc
3420attgacaaag attgtaaatt accaatggga ctggcaagtg ggataataca
agactcacag 3480atcagtgctt caggtcatgt tggatattgg gagcctaagc
tagcaagact gaataatact 3540ggaatgttta atgcttggag catcataaag
aaggaacatg aacatccgtg gatccagatc 3600gacctacaaa gacaagttgt
catcacaggc attcagaccc agggaaccgt gcacctactg 3660aaacattcgt
atactgtgga atattttgtt acctacagca aagatgggca aaactggatt
3720acttttaaag gaagacattc caaaacacaa atgcattttg agggtaattc
agatggcacc 3780acagtaaaag aaaaccacat tgatcctcct attattgcca
gatatattag gctgcatcca 3840accaagttct acaacacacc tactttccgc
attgaactgt taggttgtga agttgaaggt 3900tgctcagtgc cattgggaat
ggaaagtggg gctatcaagg attcagagat tacagcctct 3960tcttataaaa
agacttggtg gagttcatgg gaaccattcc ttgcacgact caatctgaaa
4020ggacgaacaa atgcttggca accaaaggta aacaacaaag atcaatggct
acaaattgac 4080ctgcaacatc ttacaaaaat aacaagcata ataactcaag
gagccacatc aatgactaca 4140tcaatgtatg tgaaaacatt ctccatccat
tatactgatg acaattcaac atggaagcct 4200tatttggatg ttcgcacttc
catggaaaag gttttcacag gaaatattaa cagtgatggt 4260catgtcaaac
attttttcaa cccccctata ttgtccaggt tcattcgtat catccctaaa
4320acatggaatc aatatattgc actccggata gaattgtttg gttgtgaagt
tttttaaggc 4380ttggacagaa gactgtcaaa tcaagcaact tcaatgtttc
aagttttctt attactaact 4440ctgcttttta aaaggaaaca aaaacaaaag
cataataaaa ctgtcttagc ataaaaaaaa 4500ctatccttct caattttcag
ccatagcttt caaatagctt tgaaaatatc aatcaaaata 4560tcataactga
agtgacgttt acaatgatta attcgtagtg ccacgtttaa tcatgaatgt
4620aatcctaata caataaaatc gttattgttt ttgccccaaa aaaaaaaaaa aaaaa
4675111458PRTOxyuranus scutellatus scutellatus 11Met Gly Arg Tyr
Ser Val Ser Pro Val Pro Lys Cys Leu Leu Leu Met1 5 10 15Phe Leu Gly
Trp Ser Gly Leu Lys Tyr Tyr Gln Val Asn Ala Ala Gln 20 25 30Leu Arg
Glu Tyr Arg Ile Ala Ala Gln Leu Glu Asp Trp Asp Tyr Asn 35 40 45Pro
Gln Pro Glu Glu Leu Ser Arg Leu Ser Glu Ser Asp Leu Thr Phe 50 55
60Lys Lys Ile Val Tyr Arg Glu Tyr Glu Leu Asp Phe Lys Gln Glu Lys65
70 75 80Pro Arg Asp Glu Leu Ser Gly Leu Leu Gly Pro Thr Leu Arg Gly
Glu 85 90 95Val Gly Asp Ser Leu Ile Ile Tyr Phe Lys Asn Phe Ala Thr
Gln Pro 100 105 110Val Ser Ile His Pro Gln Ser Ala Val Tyr Asn Lys
Trp Ser Glu Gly 115 120 125Ser Ser Tyr Ser Asp Gly Thr Ser Asp Val
Glu Arg Leu Asp Asp Ala 130 135 140Val Pro Pro Gly Gln Ser Phe Lys
Tyr Val Trp Asn Ile Thr Ala Glu145 150 155 160Ile Gly Pro Lys Lys
Ala Asp Pro Pro Cys Leu Thr Tyr Ala Tyr Tyr 165 170 175Ser His Val
Asn Met Val Arg Asp Phe Asn Ser Gly Leu Ile Gly Ala 180 185 190Leu
Leu Ile Cys Lys Glu Gly Ser Leu Asn Ala Asp Gly Ala Gln Lys 195 200
205Phe Phe Asn Arg Glu Tyr Val Leu Met Phe Ser Val Phe Asp Glu Ser
210 215 220Lys Asn Trp Tyr Arg Lys Pro Ser Leu Gln Tyr Thr Ile Asn
Gly Phe225 230 235 240Ala Asn Gly Thr Leu Pro Asp Val Gln Ala Cys
Ala Tyr Asp His Ile 245 250 255Ser Trp His Leu Ile Gly Met Ser Ser
Ser Pro Glu Ile Phe Ser Val 260 265 270His Phe Asn Gly Gln Thr Leu
Glu Gln Asn His Tyr Lys Val Ser Thr 275 280 285Ile Asn Leu Val Gly
Gly Ala Ser Val Thr Ala Asn Met Ser Val Ser 290 295 300Arg Thr Gly
Lys Trp Leu Ile Ser Ser Leu Val Ala Lys His Leu Gln305 310 315
320Ala Gly Met Tyr Gly Tyr Leu Asn Ile Lys Asp Cys Gly Asn Pro Asp
325 330 335Thr Leu Thr Arg Lys Leu Ser Phe Arg Glu Trp Arg Arg Ile
Met Lys 340 345 350Trp Glu Tyr Phe Ile Ala Ala Glu Glu Ile Thr Trp
Asp Tyr Ala Pro 355 360 365Glu Ile Pro Ser Ser Val Asp Arg Arg Tyr
Lys Ala Gln Tyr Leu Asp 370 375 380Phe Ser Asn Phe Ile Gly Lys Lys
Tyr Lys Lys Ala Val Phe Arg Gln385 390 395 400Tyr Glu Asp Ser Asn
Phe Thr Lys Pro Thr Tyr Ala Ile Trp Pro Lys 405 410 415Glu Arg Gly
Ile Leu Gly Pro Val Ile Lys Ala Lys Val Arg Asp Thr 420 425 430Val
Thr Ile Val Phe Lys Asn Leu Ala Ser Arg Pro Tyr Ser Ile Tyr 435 440
445Val His Gly Val Ser Val Ser Lys Asp Ala Glu Gly Ala Val Tyr Pro
450 455 460Ser Asp Pro Lys Glu Asn Ile Thr His Gly Lys Ala Val Glu
Pro Gly465 470 475 480Gln Val Tyr Thr Tyr Lys Trp Thr Val Leu Asp
Thr Asp Glu Pro Thr 485 490 495Val Lys Asp Ser Glu Cys Ile Thr Lys
Leu Tyr His Ser Ala Val Asp 500 505 510Met Thr Arg Asp Ile Ala Ser
Gly Leu Ile Gly Pro Leu Leu Val Cys 515 520 525Lys Arg Lys Ala Leu
Ser Ile Arg Gly Val Gln Asn Lys Ala Asp Val 530 535 540Glu Gln His
Ala Val Phe Ala Val Phe Asp Glu Asn Lys Ser Trp Tyr545 550 555
560Leu Glu Asp Asn Ile Lys Lys Tyr Cys Ser Asn Pro Ser Ser Val Lys
565 570 575Lys Asp Asp Pro Lys Phe Tyr Lys Ser Asn Val Met Tyr Thr
Leu Asn 580 585 590Gly Tyr Ala Ser Asp Arg Thr Glu Val Trp Gly Phe
His Gln Ser Glu 595 600 605Val Val Glu Trp His Leu Thr Ser Val Gly
Thr Val Asp Glu Ile Val 610 615 620Pro Val His Leu Ser Gly His Thr
Phe Leu Ser Lys Gly Lys His Gln625 630 635 640Asp Ile Leu Asn Leu
Phe Pro Met Ser Gly Glu Ser Ala Thr Val Thr 645 650 655Met Asp Asn
Leu Gly Thr Trp Leu Leu Ser Ser Trp Gly Ser Cys Glu 660 665 670Met
Ser Asn Gly Met Arg Leu Arg Phe Leu Asp Ala Asn Tyr Asp Asp 675 680
685Glu Asp Glu Gly Asn Glu Glu Glu Glu
Glu Asp Asp Gly Asp Ile Phe 690 695 700Ala Asp Ile Phe Asn Pro Pro
Glu Val Val Ile Lys Lys Glu Glu Val705 710 715 720Pro Val Asn Phe
Val Pro Asp Pro Glu Ser Asp Ala Leu Ala Lys Glu 725 730 735Leu Gly
Leu Phe Asp Asp Glu Asp Asn Pro Lys Gln Ser Arg Ser Glu 740 745
750Gln Thr Glu Asp Asp Glu Glu Gln Leu Met Ile Ala Ser Met Leu Gly
755 760 765Leu Arg Ser Phe Lys Gly Ser Val Ala Glu Glu Glu Leu Lys
His Thr 770 775 780Ala Leu Ala Leu Glu Glu Asp Ala His Ala Ser Asp
Pro Arg Ile Asp785 790 795 800Ser Asn Ser Ala His Asn Ser Asp Asp
Ile Ala Gly Arg Tyr Leu Arg 805 810 815Thr Ile Tyr Arg Arg Asn Lys
Arg Arg Tyr Tyr Ile Ala Ala Glu Glu 820 825 830Val Leu Trp Asp Tyr
Ser Pro Ile Gly Lys Ser Gln Val Arg Ser Leu 835 840 845Pro Ala Lys
Thr Thr Phe Lys Lys Ala Ile Phe Arg Ser Tyr Leu Asp 850 855 860Asp
Thr Phe Gln Thr Pro Ser Thr Gly Gly Glu Tyr Glu Lys His Leu865 870
875 880Gly Ile Leu Gly Pro Ile Ile Arg Ala Glu Val Asp Asp Val Ile
Glu 885 890 895Val Gln Phe Arg Asn Leu Ala Ser Arg Pro Tyr Ser Leu
His Ala His 900 905 910Gly Leu Leu Tyr Glu Lys Ser Ser Glu Gly Arg
Ser Tyr Asp Asp Asn 915 920 925Ser Pro Glu Leu Phe Lys Lys Asp Asp
Ala Ile Met Pro Asn Gly Thr 930 935 940Tyr Thr Tyr Val Trp Gln Val
Pro Pro Arg Ser Gly Pro Thr Asp Asn945 950 955 960Thr Glu Lys Cys
Lys Ser Trp Ala Tyr Tyr Ser Gly Val Asn Pro Glu 965 970 975Lys Asp
Ile His Ser Gly Leu Ile Gly Pro Ile Leu Ile Cys Gln Lys 980 985
990Gly Met Ile Asp Lys Tyr Asn Arg Thr Ile Asp Ile Arg Glu Phe Val
995 1000 1005Leu Phe Phe Met Val Phe Asp Glu Glu Lys Ser Trp Tyr
Phe Pro 1010 1015 1020Lys Ser Asp Lys Ser Thr Cys Glu Glu Lys Leu
Ile Gly Val Gln 1025 1030 1035Ser Arg His Thr Phe Pro Ala Ile Asn
Gly Ile Pro Tyr Gln Leu 1040 1045 1050Gln Gly Leu Met Met Tyr Lys
Asp Glu Asn Val His Trp His Leu 1055 1060 1065Leu Asn Met Gly Gly
Pro Lys Asp Val His Val Val Asn Phe His 1070 1075 1080Gly Gln Thr
Phe Thr Glu Glu Gly Arg Glu Asp Asn Gln Leu Gly 1085 1090 1095Val
Leu Pro Leu Leu Pro Gly Thr Phe Ala Ser Ile Lys Met Lys 1100 1105
1110Pro Ser Lys Ile Gly Thr Trp Leu Leu Glu Thr Glu Val Gly Glu
1115 1120 1125Asn Gln Glu Arg Gly Met Gln Ala Leu Phe Thr Val Ile
Asp Lys 1130 1135 1140Asp Cys Lys Leu Pro Met Gly Leu Ala Ser Gly
Ile Ile Gln Asp 1145 1150 1155Ser Gln Ile Ser Ala Ser Gly His Val
Gly Tyr Trp Glu Pro Lys 1160 1165 1170Leu Ala Arg Leu Asn Asn Thr
Gly Met Phe Asn Ala Trp Ser Ile 1175 1180 1185Ile Lys Lys Glu His
Glu His Pro Trp Ile Gln Ile Asp Leu Gln 1190 1195 1200Arg Gln Val
Val Ile Thr Gly Ile Gln Thr Gln Gly Thr Val His 1205 1210 1215Leu
Leu Lys His Ser Tyr Thr Val Glu Tyr Phe Val Thr Tyr Ser 1220 1225
1230Lys Asp Gly Gln Asn Trp Ile Thr Phe Lys Gly Arg His Ser Lys
1235 1240 1245Thr Gln Met His Phe Glu Gly Asn Ser Asp Gly Thr Thr
Val Lys 1250 1255 1260Glu Asn His Ile Asp Pro Pro Ile Ile Ala Arg
Tyr Ile Arg Leu 1265 1270 1275His Pro Thr Lys Phe Tyr Asn Thr Pro
Thr Phe Arg Ile Glu Leu 1280 1285 1290Leu Gly Cys Glu Val Glu Gly
Cys Ser Val Pro Leu Gly Met Glu 1295 1300 1305Ser Gly Ala Ile Lys
Asp Ser Glu Ile Thr Ala Ser Ser Tyr Lys 1310 1315 1320Lys Thr Trp
Trp Ser Ser Trp Glu Pro Phe Leu Ala Arg Leu Asn 1325 1330 1335Leu
Lys Gly Arg Thr Asn Ala Trp Gln Pro Lys Val Asn Asn Lys 1340 1345
1350Asp Gln Trp Leu Gln Ile Asp Leu Gln His Leu Thr Lys Ile Thr
1355 1360 1365Ser Ile Ile Thr Gln Gly Ala Thr Ser Met Thr Thr Ser
Met Tyr 1370 1375 1380Val Lys Thr Phe Ser Ile His Tyr Thr Asp Asp
Asn Ser Thr Trp 1385 1390 1395Lys Pro Tyr Leu Asp Val Arg Thr Ser
Met Glu Lys Val Phe Thr 1400 1405 1410Gly Asn Ile Asn Ser Asp Gly
His Val Lys His Phe Phe Asn Pro 1415 1420 1425Pro Ile Leu Ser Arg
Phe Ile Arg Ile Ile Pro Lys Thr Trp Asn 1430 1435 1440Gln Tyr Ile
Ala Leu Arg Ile Glu Leu Phe Gly Cys Glu Val Phe 1445 1450
1455121459PRTOxyuranus scutellatus 12Met Gly Arg Tyr Ser Val Ser
Pro Val Pro Lys Cys Leu Leu Leu Met1 5 10 15Phe Leu Gly Trp Ser Gly
Leu Lys Tyr Tyr Gln Val Asn Ala Ala Gln 20 25 30Leu Arg Glu Tyr Arg
Leu Ala Ala Gln Leu Glu Asp Trp Asp Tyr Asn 35 40 45Pro Gln Pro Glu
Glu Leu Ser Arg Leu Ser Glu Ser Asp Leu Thr Phe 50 55 60Lys Lys Ile
Val Tyr Arg Glu Tyr Glu Leu Asp Phe Lys Gln Glu Lys65 70 75 80Pro
Arg Asp Glu Leu Ser Gly Leu Leu Gly Pro Thr Leu Arg Gly Glu 85 90
95Val Gly Asp Ser Leu Ile Ile Tyr Phe Lys Asn Phe Ala Thr Gln Pro
100 105 110Val Ser Ile His Pro Gln Ser Ala Val Tyr Asn Lys Trp Ser
Glu Gly 115 120 125Ser Ser Tyr Ser Asp Gly Thr Ser Asp Val Glu Arg
Leu Asp Asp Ala 130 135 140Val Pro Pro Gly Gln Ser Phe Lys Tyr Val
Trp Asn Ile Thr Ala Glu145 150 155 160Ile Gly Pro Lys Lys Ala Asp
Pro Pro Cys Leu Thr Tyr Ala Tyr Tyr 165 170 175Ser His Val Asn Met
Val Arg Asp Phe Asn Ser Gly Leu Ile Gly Ala 180 185 190Leu Leu Ile
Cys Lys Glu Gly Ser Leu Asn Ala Asp Gly Ala Gln Lys 195 200 205Phe
Phe Asn Arg Glu Tyr Val Leu Met Phe Ser Val Phe Asp Glu Ser 210 215
220Lys Asn Trp Tyr Arg Lys Pro Ser Leu Gln Tyr Thr Ile Asn Gly
Phe225 230 235 240Ala Asn Gly Thr Leu Pro Asp Val Gln Ala Cys Ala
Tyr Asp His Ile 245 250 255Ser Trp His Leu Ile Gly Met Ser Ser Ser
Pro Glu Ile Phe Ser Val 260 265 270His Phe Asn Gly Gln Thr Leu Glu
Gln Asn His Tyr Lys Val Ser Thr 275 280 285Ile Asn Leu Val Gly Gly
Ala Ser Val Thr Ala Asn Met Ser Val Ser 290 295 300Arg Thr Gly Lys
Trp Leu Ile Ser Ser Leu Val Ala Lys His Leu Gln305 310 315 320Ala
Gly Met Tyr Gly Tyr Leu Asn Ile Lys Asp Cys Gly Asn Pro Asp 325 330
335Thr Leu Thr Arg Lys Leu Ser Phe Arg Glu Arg Arg Arg Ile Met Lys
340 345 350Trp Glu Tyr Phe Ile Ala Ala Glu Glu Ile Thr Trp Asp Tyr
Ala Pro 355 360 365Glu Ile Pro Ser Ser Val Asp Arg Arg Tyr Lys Ala
Gln Tyr Leu Asp 370 375 380Asn Phe Ser Asn Phe Ile Gly Lys Lys Tyr
Lys Lys Ala Val Phe Arg385 390 395 400Gln Tyr Glu Asp Ser Asn Phe
Thr Lys Pro Thr Tyr Ala Ile Trp Pro 405 410 415Lys Glu Arg Gly Ile
Leu Gly Pro Val Ile Lys Ala Lys Val Arg Asp 420 425 430Thr Val Thr
Ile Val Phe Lys Asn Leu Ala Ser Arg Pro Tyr Ser Ile 435 440 445Tyr
Val His Gly Val Ser Val Ser Lys Asp Ala Glu Gly Ala Val Tyr 450 455
460Pro Ser Asp Pro Lys Glu Asn Ile Thr His Gly Lys Ala Val Glu
Pro465 470 475 480Gly Gln Val Tyr Thr Tyr Lys Trp Thr Val Leu Asp
Thr Asp Glu Pro 485 490 495Thr Val Lys Asp Ser Glu Cys Ile Thr Lys
Leu Tyr His Ser Ala Val 500 505 510Asp Met Thr Arg Asp Ile Ala Ser
Gly Leu Ile Gly Pro Pro Leu Val 515 520 525Cys Lys Arg Lys Ala Leu
Ser Ile Arg Gly Val Gln Asn Lys Ala Asp 530 535 540Val Glu Gln His
Ala Val Phe Ala Val Phe Asp Glu Asn Lys Ser Trp545 550 555 560Tyr
Leu Glu Asp Asn Ile Lys Lys Tyr Cys Ser Asn Pro Ser Ser Val 565 570
575Lys Lys Asp Asp Pro Lys Phe Tyr Lys Ser Asn Val Met Tyr Thr Leu
580 585 590Asn Gly Tyr Ala Ser Asp Arg Thr Glu Val Trp Gly Phe His
Gln Ser 595 600 605Glu Val Val Glu Trp His Leu Thr Ser Val Gly Thr
Val Asp Glu Ile 610 615 620Val Pro Val His Leu Ser Gly His Thr Phe
Leu Ser Lys Gly Lys His625 630 635 640Gln Asp Ile Leu Asn Leu Phe
Pro Met Ser Gly Glu Ser Ala Thr Val 645 650 655Thr Met Asp Asn Leu
Gly Thr Trp Leu Leu Ser Ser Trp Gly Ser Cys 660 665 670Glu Met Ser
Asn Gly Met Arg Leu Arg Phe Leu Asp Ala Asn Tyr Asp 675 680 685Asp
Glu Asp Glu Gly Asn Glu Glu Glu Glu Glu Asp Asp Gly Asp Ile 690 695
700Phe Ala Asp Ile Phe Asn Pro Pro Glu Val Val Ile Lys Lys Glu
Glu705 710 715 720Val Pro Val Asn Phe Val Pro Asp Pro Glu Ser Asp
Ala Leu Ala Lys 725 730 735Glu Leu Gly Leu Phe Asp Asp Glu Asp Asn
Pro Lys Gln Ser Arg Ser 740 745 750Glu Gln Thr Glu Asp Asp Glu Glu
Gln Leu Met Ile Ala Ser Met Leu 755 760 765Gly Leu Arg Ser Phe Lys
Gly Ser Val Ala Glu Glu Glu Leu Lys His 770 775 780Thr Ala Leu Ala
Leu Glu Glu Asp Ala His Ala Ser Asp Pro Arg Ile785 790 795 800Asp
Ser Asn Ser Ala His Asn Ser Asp Asp Ile Ala Gly Arg Tyr Leu 805 810
815Arg Thr Ile Asn Arg Arg Asn Lys Arg Arg Tyr Tyr Ile Ala Ala Glu
820 825 830Glu Val Leu Trp Asp Tyr Ser Pro Ile Gly Lys Ser Gln Val
Arg Ser 835 840 845Leu Pro Ala Lys Thr Thr Phe Lys Lys Ala Ile Phe
Arg Ser Tyr Leu 850 855 860Asp Asp Thr Phe Gln Thr Pro Ser Thr Gly
Gly Glu Tyr Glu Lys His865 870 875 880Leu Gly Ile Leu Gly Pro Ile
Ile Arg Ala Glu Val Asp Asp Val Ile 885 890 895Glu Val Gln Phe Arg
Asn Leu Ala Ser Arg Pro Tyr Ser Leu His Ala 900 905 910His Gly Leu
Leu Tyr Glu Lys Ser Ser Glu Gly Arg Ser Tyr Asp Asp 915 920 925Asn
Ser Pro Glu Leu Phe Lys Lys Asp Asp Ala Ile Met Pro Asn Gly 930 935
940Thr Tyr Thr Tyr Val Trp Gln Val Pro Pro Arg Ser Gly Pro Thr
Asp945 950 955 960Asn Thr Glu Lys Cys Lys Ser Trp Ala Tyr Tyr Ser
Gly Val Asn Pro 965 970 975Glu Lys Asp Ile His Ser Gly Leu Ile Gly
Pro Ile Leu Ile Cys Gln 980 985 990Lys Gly Met Ile Asp Lys Tyr Asn
Arg Thr Ile Asp Ile Arg Glu Phe 995 1000 1005Val Leu Phe Phe Met
Val Phe Asp Glu Glu Lys Ser Trp Tyr Phe 1010 1015 1020Pro Lys Ser
Asp Lys Ser Thr Cys Glu Glu Lys Leu Ile Gly Val 1025 1030 1035Gln
Ser Arg His Thr Phe Pro Ala Ile Asn Gly Ile Pro Tyr Gln 1040 1045
1050Leu Gln Gly Leu Met Met Tyr Lys Asp Glu Asn Val His Trp His
1055 1060 1065Leu Leu Asn Met Gly Gly Pro Lys Asp Val His Val Val
Asn Phe 1070 1075 1080His Gly Gln Thr Phe Thr Glu Glu Gly Arg Glu
Asp Asn Gln Leu 1085 1090 1095Gly Val Leu Pro Leu Leu Pro Gly Thr
Phe Ala Ser Ile Lys Met 1100 1105 1110Lys Pro Ser Lys Ile Gly Thr
Trp Leu Leu Glu Thr Glu Val Gly 1115 1120 1125Glu Asn Gln Glu Arg
Gly Met Gln Ala Leu Phe Thr Val Ile Asp 1130 1135 1140Lys Asp Cys
Lys Leu Pro Met Gly Leu Ala Ser Gly Ile Ile Gln 1145 1150 1155Asp
Ser Gln Ile Ser Ala Ser Gly His Val Gly Tyr Trp Glu Pro 1160 1165
1170Lys Leu Ala Arg Leu Asn Asn Thr Gly Met Phe Asn Ala Trp Ser
1175 1180 1185Ile Ile Lys Lys Glu His Glu His Pro Trp Ile Gln Ile
Asp Leu 1190 1195 1200Gln Arg Gln Val Val Ile Thr Gly Ile Gln Thr
Gln Gly Thr Val 1205 1210 1215Gln Leu Leu Lys His Ser Tyr Thr Val
Glu Tyr Phe Val Thr Tyr 1220 1225 1230Ser Lys Asp Gly Gln Asn Trp
Ile Thr Phe Lys Gly Arg His Ser 1235 1240 1245Lys Thr Gln Met His
Phe Glu Gly Asn Ser Asp Gly Thr Thr Val 1250 1255 1260Lys Glu Asn
His Ile Asp Pro Pro Ile Ile Ala Arg Tyr Ile Arg 1265 1270 1275Leu
His Pro Thr Lys Phe Tyr Asn Thr Pro Thr Phe Arg Ile Glu 1280 1285
1290Leu Leu Gly Cys Glu Val Glu Gly Cys Ser Val Pro Leu Gly Met
1295 1300 1305Glu Ser Gly Ala Ile Lys Asp Ser Glu Ile Thr Ala Ser
Ser Tyr 1310 1315 1320Lys Lys Thr Trp Trp Ser Ser Trp Glu Pro Phe
Leu Ala Arg Leu 1325 1330 1335Asn Leu Lys Gly Arg Thr Asn Ala Trp
Gln Pro Lys Val Asn Asn 1340 1345 1350Lys Asp Gln Trp Leu Gln Ile
Asp Leu Gln His Leu Thr Lys Ile 1355 1360 1365Thr Ser Ile Ile Thr
Gln Gly Ala Thr Ser Met Thr Thr Ser Met 1370 1375 1380Tyr Val Lys
Thr Phe Ser Ile His Tyr Thr Asp Asp Asn Ser Thr 1385 1390 1395Trp
Lys Pro Tyr Leu Asp Val Arg Thr Ser Met Glu Lys Val Phe 1400 1405
1410Thr Gly Asn Ile Asn Ser Asp Gly His Val Lys His Phe Phe Asn
1415 1420 1425Pro Pro Ile Leu Ser Arg Phe Ile Arg Ile Ile Pro Lys
Thr Trp 1430 1435 1440Asn Gln Tyr Ile Ala Leu Arg Ile Glu Leu Phe
Gly Cys Glu Val 1445 1450 1455Phe131460PRTOxyuranus scutellatus
13Met Gly Arg Tyr Ser Val Ser Pro Val Pro Lys Cys Leu Leu Leu Met1
5 10 15Phe Leu Gly Trp Ser Gly Leu Lys Tyr Tyr Gln Val Asn Ala Ala
Gln 20 25 30Leu Arg Glu Tyr Arg Ile Ala Ala Gln Leu Glu Asp Trp Asp
Tyr Asn 35 40 45Pro Gln Pro Glu Glu Leu Ser Arg Leu Ser Glu Ser Glu
Leu Thr Phe 50 55 60Lys Lys Ile Val Tyr Arg Glu Tyr Glu Leu Asp Phe
Lys Gln Glu Lys65 70 75 80Pro Arg Asp Glu Leu Ser Gly Leu Leu Gly
Pro Thr Leu Arg Gly Glu 85 90 95Val Gly Asp Ile Leu Ile Ile Tyr Phe
Lys Asn Phe Ala Thr Gln Pro 100 105 110Val Ser Ile His Pro Gln Ser
Ala Val Tyr Asn Lys Trp Ser Glu Gly 115 120 125Ser Ser Tyr Ser Asp
Gly Thr Ser Asp Val Glu Arg Leu Asp Asp Ala 130 135 140Val Pro Pro
Gly Gln Ser Phe Lys Tyr Val Trp Asn Ile Thr Ala Glu145 150 155
160Ile Gly Pro Lys Lys Ala Asp Pro Pro Cys Leu Thr Tyr Ala Tyr Tyr
165 170 175Ser His Val Asn Met Val Arg Asp Phe Asn Ser Gly Leu Ile
Gly Ala 180 185 190Leu Leu Ile Cys Lys Glu Gly Ser Leu Asn Ala Asn
Gly Ala Gln Lys 195 200 205Phe Phe Asn Arg Glu Tyr Val Leu Met Phe
Ser Val Phe Asp Glu Ser 210 215 220Lys Asn Trp Tyr Arg Lys Pro Ser
Leu Gln Tyr
Thr Ile Asn Gly Phe225 230 235 240Ala Asn Gly Thr Leu Pro Asp Val
Gln Ala Cys Ala Tyr Asp His Ile 245 250 255Ser Trp His Leu Ile Gly
Met Ser Ser Ser Pro Glu Ile Phe Ser Val 260 265 270His Phe Asn Gly
Gln Thr Leu Glu Gln Asn His Tyr Lys Val Ser Thr 275 280 285Ile Asn
Leu Val Gly Gly Ala Ser Val Thr Ala Asn Met Ser Val Ser 290 295
300Arg Thr Gly Lys Trp Leu Ile Ser Ser Leu Val Ala Lys His Leu
Gln305 310 315 320Ala Gly Met Tyr Gly Tyr Leu Asn Ile Lys Asp Cys
Gly His Pro Asn 325 330 335Thr Leu Thr Arg Lys Leu Ser Phe Arg Glu
Leu Arg Arg Ile Met Asn 340 345 350Trp Glu Tyr Phe Ile Ala Ala Glu
Glu Ile Thr Trp Asp Tyr Ala Pro 355 360 365Glu Ile Pro Ser Ser Val
Asp Arg Arg Tyr Lys Ala Gln Tyr Leu Asp 370 375 380Asn Phe Ser Asn
Phe Ile Gly Lys Lys Tyr Lys Lys Ala Val Phe Arg385 390 395 400Gln
Tyr Glu Asp Gly Asn Phe Thr Lys Pro Thr Tyr Ala Ile Trp Pro 405 410
415Lys Glu Arg Gly Ile Leu Gly Pro Val Ile Lys Ala Lys Val Arg Asp
420 425 430Thr Val Thr Ile Val Phe Lys Asn Leu Ala Ser Arg Pro Tyr
Ser Ile 435 440 445Tyr Val His Gly Val Ser Val Ser Lys Asp Ala Glu
Gly Ala Ile Tyr 450 455 460Pro Ser Asp Pro Lys Glu Asn Ile Thr His
Gly Lys Ala Val Glu Pro465 470 475 480Gly Gln Val Tyr Thr Tyr Lys
Trp Thr Val Leu Asp Thr Asp Glu Pro 485 490 495Thr Val Lys Asp Ser
Glu Cys Ile Thr Lys Leu Tyr His Ser Ala Val 500 505 510Asp Met Thr
Arg Asp Ile Ala Ser Gly Leu Ile Gly Pro Leu Leu Val 515 520 525Cys
Lys Leu Lys Ala Leu Ser Val Lys Gly Val Gln Asn Lys Ala Asp 530 535
540Val Glu Gln His Ala Val Phe Ala Val Phe Asp Glu Asn Lys Ser
Trp545 550 555 560Tyr Leu Glu Asp Asn Ile Lys Lys Tyr Cys Ser Asn
Pro Ser Ser Val 565 570 575Lys Lys Asp Asp Pro Lys Phe Tyr Lys Ser
Asn Val Met Tyr Thr Leu 580 585 590Asn Gly Tyr Ala Ser Asp Arg Thr
Glu Val Leu Gly Phe His Gln Ser 595 600 605Glu Val Val Gln Trp His
Leu Thr Ser Val Gly Thr Val Asp Glu Ile 610 615 620Val Pro Val His
Leu Ser Gly His Thr Phe Leu Ser Lys Gly Lys His625 630 635 640Gln
Asp Ile Leu Asn Leu Phe Pro Met Ser Gly Glu Ser Ala Thr Val 645 650
655Thr Met Asp Asn Leu Gly Thr Trp Leu Leu Ser Ser Trp Gly Ser Cys
660 665 670Glu Met Ser Asn Gly Met Arg Leu Arg Phe Leu Asp Ala Asn
Tyr Asp 675 680 685Asp Glu Asp Glu Gly Asn Glu Glu Glu Glu Glu Asp
Asp Gly Asp Ile 690 695 700Phe Ala Asp Ile Phe Ser Pro Pro Glu Val
Val Lys Lys Lys Glu Glu705 710 715 720Val Pro Val Asn Phe Val Pro
Asp Pro Glu Ser Asp Ala Leu Ala Lys 725 730 735Glu Leu Gly Leu Leu
Asp Asp Glu Asp Asn Pro Glu Gln Ser Arg Ser 740 745 750Glu Gln Thr
Glu Asp Asp Glu Glu Gln Leu Met Ile Ala Ser Val Leu 755 760 765Gly
Leu Arg Ser Phe Lys Gly Ser Val Ala Glu Glu Glu Leu Lys His 770 775
780Thr Ala Leu Ala Leu Glu Glu Asp Ala His Ala Ser Asp Pro Arg
Ile785 790 795 800Asp Ser Asn Ser Ala Arg Asn Ser Asp Asp Ile Ala
Gly Arg Tyr Leu 805 810 815Arg Thr Ile Asn Arg Arg Asn Lys Arg Arg
Tyr Tyr Ile Ala Ala Glu 820 825 830Glu Val Leu Trp Asp Tyr Ser Pro
Ile Gly Lys Ser Gln Val Arg Ser 835 840 845Leu Pro Ala Lys Thr Thr
Phe Lys Lys Ala Ile Phe Arg Ser Tyr Leu 850 855 860Asp Asp Thr Phe
Gln Thr Pro Ser Thr Gly Gly Glu Tyr Glu Lys His865 870 875 880Leu
Gly Ile Leu Gly Pro Ile Ile Arg Ala Glu Val Asp Asp Val Ile 885 890
895Glu Val Gln Phe Arg Asn Leu Ala Ser Arg Pro Tyr Ser Leu His Ala
900 905 910His Gly Leu Leu Tyr Glu Lys Ser Ser Glu Gly Arg Ser Tyr
Asp Asp 915 920 925Asn Ser Pro Glu Leu Phe Lys Lys Asp Asp Ala Ile
Met Pro Asn Gly 930 935 940Thr Tyr Thr Tyr Val Trp Gln Val Pro Pro
Arg Ser Gly Pro Thr Asp945 950 955 960Asn Thr Glu Lys Cys Lys Ser
Trp Ala Tyr Tyr Ser Gly Val Asn Pro 965 970 975Glu Lys Asp Ile His
Ser Gly Leu Ile Gly Pro Ile Leu Ile Cys Gln 980 985 990Lys Gly Met
Ile Asp Lys Tyr Asn Arg Thr Ile Asp Ile Arg Glu Phe 995 1000
1005Val Leu Phe Phe Met Val Phe Asp Glu Glu Lys Ser Trp Tyr Phe
1010 1015 1020Pro Lys Ser Asp Lys Ser Thr Cys Glu Glu Lys Leu Ile
Gly Val 1025 1030 1035Gln Ser Ser His His Thr Phe Pro Ala Ile Asn
Gly Ile Pro Tyr 1040 1045 1050Gln Leu Gln Gly Leu Met Met Tyr Lys
Asp Glu Asn Val His Trp 1055 1060 1065His Leu Leu Asn Met Gly Gly
Pro Lys Asp Ile His Val Val Asn 1070 1075 1080Phe His Gly Gln Thr
Phe Thr Glu Glu Gly Arg Glu Asp Asn Gln 1085 1090 1095Leu Gly Val
Leu Pro Leu Leu Pro Gly Thr Phe Ala Ser Ile Lys 1100 1105 1110Met
Lys Pro Ser Lys Ile Gly Thr Trp Leu Leu Glu Thr Glu Val 1115 1120
1125Gly Glu Asn Gln Glu Arg Gly Met Gln Ala Leu Phe Thr Val Ile
1130 1135 1140Asp Lys Asp Cys Lys Leu Pro Met Gly Leu Ala Ser Gly
Ile Ile 1145 1150 1155Gln Asp Ser Gln Ile Ser Ala Ser Gly His Val
Glu Tyr Trp Glu 1160 1165 1170Pro Lys Leu Ala Arg Leu Asn Asn Thr
Gly Met Phe Asn Ala Trp 1175 1180 1185Ser Ile Ile Lys Lys Glu His
Glu His Pro Trp Ile Gln Ile Asp 1190 1195 1200Leu Gln Arg Gln Val
Val Ile Thr Gly Ile Gln Thr Gln Gly Thr 1205 1210 1215Val Gln Leu
Leu Lys His Ser Tyr Thr Val Glu Tyr Phe Val Thr 1220 1225 1230Tyr
Ser Lys Asp Gly Gln Asn Trp Ile Thr Phe Lys Gly Arg His 1235 1240
1245Ser Glu Thr Gln Met His Phe Glu Gly Asn Ser Asp Gly Thr Thr
1250 1255 1260Val Lys Glu Asn His Ile Asp Pro Pro Ile Ile Ala Arg
Tyr Ile 1265 1270 1275Arg Leu His Pro Thr Lys Phe Tyr Asn Thr Pro
Thr Phe Arg Ile 1280 1285 1290Glu Leu Leu Gly Cys Glu Val Glu Gly
Cys Ser Val Pro Leu Gly 1295 1300 1305Met Glu Ser Gly Ala Ile Lys
Asn Ser Glu Ile Thr Ala Ser Ser 1310 1315 1320Tyr Lys Lys Thr Trp
Trp Ser Ser Trp Glu Pro Phe Leu Ala Arg 1325 1330 1335Leu Asn Leu
Glu Gly Gly Thr Asn Ala Trp Gln Pro Glu Val Asn 1340 1345 1350Asn
Lys Asp Gln Trp Leu Gln Ile Asp Leu Gln His Leu Thr Lys 1355 1360
1365Ile Thr Ser Ile Ile Thr Gln Gly Ala Thr Ser Met Thr Thr Ala
1370 1375 1380Met Tyr Val Lys Thr Phe Ser Ile His Tyr Thr Asp Asp
Asn Ser 1385 1390 1395Thr Trp Lys Pro Tyr Leu Asp Val Arg Thr Ser
Met Glu Lys Val 1400 1405 1410Phe Thr Gly Asn Ile Asn Ser Asp Gly
His Val Lys His Phe Phe 1415 1420 1425Lys Pro Pro Ile Leu Ser Arg
Phe Ile Arg Ile Ile Pro Lys Thr 1430 1435 1440Trp Asn Gln Tyr Ile
Ala Leu Arg Ile Glu Leu Phe Gly Cys Glu 1445 1450 1455Val Phe
1460144383DNAOxyuranus microlepidotus 14atgggaagat acagtgtgag
ccctgtcccc aaatgtcttc tactgatgtt cctgggttgg 60tcagggctga agtattacca
agtgaatgca gctcagctca gggagtaccg tatagctgct 120cagctggaag
actgggatta caacccccaa cctgaggagc tatccagatt atcagagtca
180gagcttacgt ttaaaaaaat tgtctataga gaatatgaac tagatttcaa
acaagagaag 240ccaagagatg agctctcagg gctcctaggg ccaacactac
gtggagaagt gggagacatc 300ctcataattt atttcaagaa ttttgctact
cagcctgtga gcattcaccc gcagagtgcc 360gtgtacaaca aatggtcaga
aggttcttca tattctgatg gaacatcaga tgtggaaaga 420ctggatgatg
ctgtgcctcc aggccagtcg ttcaagtatg tgtggaatat cactgcagaa
480attgggccaa agaaagctga tcctccctgt ctcacttatg cgtactactc
acatgtaaac 540atggtgcgag actttaattc tggtctcatt ggtgctttgc
tgatatgtaa agaaggaagc 600ctgaatgcaa atggtgcaca aaaattcttc
aacagagaat atgtgctgat gttttctgtg 660tttgatgaaa gcaagaactg
gtacagaaag ccctcactac agtacacaat taatgggttt 720gccaatggaa
cattgcctga tgttcaggct tgtgcttatg atcatattag ctggcatttg
780ataggaatga gttccagtcc tgagatcttc tctgttcact tcaatggaca
aaccttggaa 840caaaaccatt acaaagtgtc aaccatcaac cttgtcggag
gtgcctcagt aacagccaac 900atgtcagtga gcaggacagg aaaatggcta
atatcttctc tggttgcaaa gcatctacaa 960gctgggatgt atggttatct
aaatatcaaa gactgtggac atccaaatac tttaacaaga 1020aagttatcct
ttagagaact gaggaggatt atgaactggg aatatttcat tgctgcagaa
1080gaaatcacct gggattatgc tccagaaatt cctagcagtg ttgacagaag
atacaaagct 1140cagtatctgg ataatttttc aaattttatt ggcaagaaat
acaaaaaggc agttttcagg 1200caatatgaag acggcaattt cactaaaccg
acctatgcca tttggcccaa agaacgtgga 1260attctgggcc ccgttatcaa
agctaaagtc agagacacag taacaattgt attcaaaaat 1320ctggccagtc
gaccttacag catttatgtg catggagttt ccgtttcaaa agatgcagaa
1380ggagctattt atccttcaga tcccaaagag aatataactc atggcaaagc
agttgaacca 1440ggacaggtct acacatataa atggactgtg ctggatacag
atgaacctac agtaaaggat 1500tctgagtgta ttactaaatt atatcatagt
gctgtggaca tgacaagaga tattgcttca 1560ggacttattg ggccacttct
ggtttgtaaa ctcaaggcac tcagcgtcaa gggggtacag 1620aataaagctg
atgtggaaca gcatgcagtc ttcgcagtgt ttgatgaaaa taagagctgg
1680tacttggaag acaatatcaa gaaatactgc agcaatcctt ccagtgttaa
gaaagatgac 1740cctaaatttt acaagtccaa tgttatgtac acactcaatg
gctatgcatc agatagaaca 1800gaggttttgg ggtttcatca gtctgaagtt
gttcaatggc acctcaccag cgtaggtaca 1860gtggatgaga ttgttccagt
acatctttct ggtcacacct tcttatccaa gggaaaacat 1920caagatattt
taaatctttt tcccatgagt ggtgaatcgg ctactgtaac aatggacaat
1980ctaggaacct ggcttctgtc atcatggggc tcctgtgaga tgagcaatgg
catgagattg 2040agatttttgg atgccaatta tgatgatgaa gatgagggaa
atgaagaaga ggaagaagat 2100gatggtgata tttttgccga cattttcagt
cctccagaag tagtaaaaaa gaaagaagag 2160gttcccgtaa attttgtacc
agacccagaa tcggatgcgc tagcaaaaga attaggatta 2220ttagatgacg
aggataatcc agaacagtca cgcagtgaac agacagagga tgatgaagaa
2280cagctaatga tagcttcagt gcttgggctt cgatcattta aggggtcagt
tgctgaagaa 2340gaattgaaac acacagctct agctttagaa gaagatgccc
atgcttctga tcctcgaatt 2400gacagtaata gtgcacgtaa ttctgacgac
atagctggac gctacctgcg tactatcaac 2460cgcagaaata aaaggaggta
ctacattgca gcagaagaag ttttgtggga ctactcaccg 2520atcggaaaaa
gtcaagtgag aagtctccca gccaagacca cattcaaaaa agctattttc
2580cgaagttatc ttgatgatac tttccagaca cctagcactg gaggagaata
tgaaaagcat 2640cttggtatac tgggtcctat cattagggct gaggtggatg
atgtaatcga agttcagttc 2700agaaatttgg cctctagacc atactcactt
catgctcatg gccttctcta tgagaaatct 2760tctgaaggca gaagctatga
cgacaactct cctgaattgt tcaaaaagga tgatgctatc 2820atgccaaacg
gcacatacac atatgtctgg caagtccctc cacggtcagg accaacagac
2880aatacagaaa aatgtaaatc atgggcctat tactctggtg taaatccgga
aaaagatatt 2940cactctggct taattggacc tattttgatc tgccagaaag
gcatgattga caagtacaac 3000aggacaatag acataaggga atttgtcttg
ttttttatgg tctttgatga ggagaaaagc 3060tggtactttc ccaaatctga
caaaagcact tgtgaagaga aacttatagg agtccaatct 3120tctcaccaca
catttcctgc aattaatggg atcccttatc agctgcaagg cttgatgatg
3180tacaaagatg agaatgtcca ctggcatttg ctgaacatgg gtgggcccaa
agatatccat 3240gttgttaatt ttcatggtca gacattcact gaagagggaa
gggaagataa tcaacttgga 3300gtccttcctc ttcttcctgg tacattcgcc
tccatcaaaa tgaaaccatc caaaattggc 3360acatggcttt tagaaacaga
agttggtgaa aatcaggaaa gaggaatgca ggctctcttt 3420actgtcattg
acaaagattg taaattacca atgggactgg caagtgggat aatacaagac
3480tcacagatca gtgcttcagg tcatgttgaa tattgggagc ctaagctagc
aagactgaat 3540aatactggaa tgtttaatgc ttggagcatc ataaagaagg
aacatgaaca tccgtggatc 3600cagatagacc tacaaagaca agttgtcatc
acaggcattc agacccaggg aaccgtgcaa 3660ctactgaaac attcgtatac
tgtggaatat tttgttacct acagcaaaga tgggcaaaac 3720tggattactt
ttaaaggaag acattccgaa acacaaatgc attttgaggg taattcagat
3780ggcaccacag taaaagaaaa ccacattgat cctcctatta ttgccagata
tattaggctg 3840catccaacca agttctacaa cacacctact ttccgcattg
aactgttagg ttgtgaagtt 3900gaaggttgct cagtgccatt gggaatggaa
agtggggcta tcaagaattc agagattaca 3960gcctcttctt ataagaagac
ttggtggagt tcatgggaac cattccttgc acgactcaat 4020ctggaaggag
gaacaaatgc ttggcaacca gaggtaaaca acaaagatca atggctacaa
4080attgacctgc aacatcttac aaaaataaca agcataataa ctcaaggagc
cacatcaatg 4140actacagcaa tgtatgtgaa aacattctcc atccattata
ctgatgacaa ttcaacatgg 4200aagccttatt tggatgttcg cacttccatg
gaaaaggttt tcacaggaaa tattaacagt 4260gatggtcatg tcaaacattt
tttcaaaccc cctatattgt ccaggttcat tcgtatcatc 4320cctaaaacat
ggaatcaata tattgcactc cggatagaat tgtttggttg tgaagttttt 4380taa
4383159179DNAHomo sapiens 15gcaagaactg caggggagga ggacgctgcc
acccacagcc tctagagctc attgcagctg 60ggacagcccg gagtgtggtt agcagctcgg
caagcgctgc ccaggtcctg gggtggtggc 120agccagcggg agcaggaaag
gaagcatgtt cccaggctgc ccacgcctct gggtcctggt 180ggtcttgggc
accagctggg taggctgggg gagccaaggg acagaagcgg cacagctaag
240gcagttctac gtggctgctc agggcatcag ttggagctac cgacctgagc
ccacaaactc 300aagtttgaat ctttctgtaa cttcctttaa gaaaattgtc
tacagagagt atgaaccata 360ttttaagaaa gaaaaaccac aatctaccat
ttcaggactt cttgggccta ctttatatgc 420tgaagtcgga gacatcataa
aagttcactt taaaaataag gcagataagc ccttgagcat 480ccatcctcaa
ggaattaggt acagtaaatt atcagaaggt gcttcttacc ttgaccacac
540attccctgcg gagaagatgg acgacgctgt ggctccaggc cgagaataca
cctatgaatg 600gagtatcagt gaggacagtg gacccaccca tgatgaccct
ccatgcctca cacacatcta 660ttactcccat gaaaatctga tcgaggattt
caactcgggg ctgattgggc ccctgcttat 720ctgtaaaaaa gggaccctaa
ctgagggtgg gacacagaag acgtttgaca agcaaatcgt 780gctactattt
gctgtgtttg atgaaagcaa gagctggagc cagtcatcat ccctaatgta
840cacagtcaat ggatatgtga atgggacaat gccagatata acagtttgtg
cccatgacca 900catcagctgg catctgctgg gaatgagctc ggggccagaa
ttattctcca ttcatttcaa 960cggccaggtc ctggagcaga accatcataa
ggtctcagcc atcacccttg tcagtgctac 1020atccactacc gcaaatatga
ctgtgggccc agagggaaag tggatcatat cttctctcac 1080cccaaaacat
ttgcaagctg ggatgcaggc ttacattgac attaaaaact gcccaaagaa
1140aaccaggaat cttaagaaaa taactcgtga gcagaggcgg cacatgaaga
ggtgggaata 1200cttcattgct gcagaggaag tcatttggga ctatgcacct
gtaataccag cgaatatgga 1260caaaaaatac aggtctcagc atttggataa
tttctcaaac caaattggaa aacattataa 1320gaaagttatg tacacacagt
acgaagatga gtccttcacc aaacatacag tgaatcccaa 1380tatgaaagaa
gatgggattt tgggtcctat tatcagagcc caggtcagag acacactcaa
1440aatcgtgttc aaaaatatgg ccagccgccc ctatagcatt taccctcatg
gagtgacctt 1500ctcgccttat gaagatgaag tcaactcttc tttcacctca
ggcaggaaca acaccatgat 1560cagagcagtt caaccagggg aaacctatac
ttataagtgg aacatcttag agtttgatga 1620acccacagaa aatgatgccc
agtgcttaac aagaccatac tacagtgacg tggacatcat 1680gagagacatc
gcctctgggc taataggact acttctaatc tgtaagagca gatccctgga
1740caggcgagga atacagaggg cagcagacat cgaacagcag gctgtgtttg
ctgtgtttga 1800tgagaacaaa agctggtacc ttgaggacaa catcaacaag
ttttgtgaaa atcctgatga 1860ggtgaaacgt gatgacccca agttttatga
atcaaacatc atgagcacta tcaatggcta 1920tgtgcctgag agcataacta
ctcttggatt ctgctttgat gacactgtcc agtggcactt 1980ctgtagtgtg
gggacccaga atgaaatttt gaccatccac ttcactgggc actcattcat
2040ctatggaaag aggcatgagg acaccttgac cctcttcccc atgcgtggag
aatctgtgac 2100ggtcacaatg gataatgttg gaacttggat gttaacttcc
atgaattcta gtccaagaag 2160caaaaagctg aggctgaaat tcagggatgt
taaatgtatc ccagatgatg atgaagactc 2220atatgagatt tttgaacctc
cagaatctac agtcatggct acacggaaaa tgcatgatcg 2280tttagaacct
gaagatgaag agagtgatgc tgactatgat taccagaaca gactggctgc
2340agcattagga atcaggtcat tccgaaactc atcattgaat caggaagaag
aagagttcaa 2400tcttactgcc ctagctctgg agaatggcac tgaattcgtt
tcttcaaaca cagatataat 2460tgttggttca aattattctt ccccaagtaa
tattagtaag ttcactgtca ataaccttgc 2520agaacctcag aaagcccctt
ctcaccaaca agccaccaca gctggttccc cactgagaca 2580cctcattggc
aagaactcag ttctcaattc ttccacagca gagcattcca gcccatattc
2640tgaagaccct atagaggatc ctctacagcc agatgtcaca gggatacgtc
tactttcact 2700tggtgctgga gaattcaaaa gtcaagaaca tgctaagcat
aagggaccca aggtagaaag 2760agatcaagca gcaaagcaca ggttctcctg
gatgaaatta ctagcacata aagttgggag 2820acacctaagc caagacactg
gttctccttc cggaatgagg ccctgggagg accttcctag 2880ccaagacact
ggttctcctt ccagaatgag gccctggaag gaccctccta gtgatctgtt
2940actcttaaaa caaagtaact catctaagat tttggttggg agatggcatt
tggcttctga 3000gaaaggtagc tatgaaataa tccaagatac
tgatgaagac acagctgtta acaattggct 3060gatcagcccc cagaatgcct
cacgtgcttg gggagaaagc acccctcttg ccaacaagcc 3120tggaaagcag
agtggccacc caaagtttcc tagagttaga cataaatctc tacaagtaag
3180acaggatgga ggaaagagta gactgaagaa aagccagttt ctcattaaga
cacgaaaaaa 3240gaaaaaagag aagcacacac accatgctcc tttatctccg
aggacctttc accctctaag 3300aagtgaagcc tacaacacat tttcagaaag
aagacttaag cattcgttgg tgcttcataa 3360atccaatgaa acatctcttc
ccacagacct caatcagaca ttgccctcta tggattttgg 3420ctggatagcc
tcacttcctg accataatca gaattcctca aatgacactg gtcaggcaag
3480ctgtcctcca ggtctttatc agacagtgcc cccagaggaa cactatcaaa
cattccccat 3540tcaagaccct gatcaaatgc actctacttc agaccccagt
cacagatcct cttctccaga 3600gctcagtgaa atgcttgagt atgaccgaag
tcacaagtcc ttccccacag atataagtca 3660aatgtcccct tcctcagaac
atgaagtctg gcagacagtc atctctccag acctcagcca 3720ggtgaccctc
tctccagaac tcagccagac aaacctctct ccagacctca gccacacgac
3780tctctctcca gaactcattc agagaaacct ttccccagcc ctcggtcaga
tgcccatttc 3840tccagacctc agccatacaa ccctttctcc agacctcagc
catacaaccc tttctttaga 3900cctcagccag acaaacctct ctccagaact
cagtcagaca aacctttctc cagccctcgg 3960tcagatgccc ctttctccag
acctcagcca tacaaccctt tctctagact tcagccagac 4020aaacctctct
ccagaactca gccatatgac tctctctcca gaactcagtc agacaaacct
4080ttccccagcc ctcggtcaga tgcccatttc tccagacctc agccatacaa
ccctttctct 4140agacttcagc cagacaaacc tctctccaga actcagtcaa
acaaaccttt ccccagccct 4200cggtcagatg cccctttctc cagaccccag
ccatacaacc ctttctctag acctcagcca 4260gacaaacctc tctccagaac
tcagtcagac aaacctttcc ccagacctca gtgagatgcc 4320cctctttgca
gatctcagtc aaattcccct taccccagac ctcgaccaga tgacactttc
4380tccagacctt ggtgagacag atctttcccc aaactttggt cagatgtccc
tttccccaga 4440cctcagccag gtgactctct ctccagacat cagtgacacc
acccttctcc cggatctcag 4500ccagatatca cctcctccag accttgatca
gatattctac ccttctgaat ctagtcagtc 4560attgcttctt caagaattta
atgagtcttt tccttatcca gaccttggtc agatgccatc 4620tccttcatct
cctactctca atgatacttt tctatcaaag gaatttaatc cactggttat
4680agtgggcctc agtaaagatg gtacagatta cattgagatc attccaaagg
aagaggtcca 4740gagcagtgaa gatgactatg ctgaaattga ttatgtgccc
tatgatgacc cctacaaaac 4800tgatgttagg acaaacatca actcctccag
agatcctgac aacattgcag catggtacct 4860ccgcagcaac aatggaaaca
gaagaaatta ttacattgct gctgaagaaa tatcctggga 4920ttattcagaa
tttgtacaaa gggaaacaga tattgaagac tctgatgata ttccagaaga
4980taccacatat aagaaagtag tttttcgaaa gtacctcgac agcactttta
ccaaacgtga 5040tcctcgaggg gagtatgaag agcatctcgg aattcttggt
cctattatca gagctgaagt 5100ggatgatgtt atccaagttc gttttaaaaa
tttagcatcc agaccgtatt ctctacatgc 5160ccatggactt tcctatgaaa
aatcatcaga gggaaagact tatgaagatg actctcctga 5220atggtttaag
gaagataatg ctgttcagcc aaatagcagt tatacctacg tatggcatgc
5280cactgagcga tcagggccag aaagtcctgg ctctgcctgt cgggcttggg
cctactactc 5340agctgtgaac ccagaaaaag atattcactc aggcttgata
ggtcccctcc taatctgcca 5400aaaaggaata ctacataagg acagcaacat
gcctatggac atgagagaat ttgtcttact 5460atttatgacc tttgatgaaa
agaagagctg gtactatgaa aagaagtccc gaagttcttg 5520gagactcaca
tcctcagaaa tgaaaaaatc ccatgagttt cacgccatta atgggatgat
5580ctacagcttg cctggcctga aaatgtatga gcaagagtgg gtgaggttac
acctgctgaa 5640cataggcggc tcccaagaca ttcacgtggt tcactttcac
ggccagacct tgctggaaaa 5700tggcaataaa cagcaccagt taggggtctg
gccccttctg cctggttcat ttaaaactct 5760tgaaatgaag gcatcaaaac
ctggctggtg gctcctaaac acagaggttg gagaaaacca 5820gagagcaggg
atgcaaacgc catttcttat catggacaga gactgtagga tgccaatggg
5880actaagcact ggtatcatat ctgattcaca gatcaaggct tcagagtttc
tgggttactg 5940ggagcccaga ttagcaagat taaacaatgg tggatcttat
aatgcttgga gtgtagaaaa 6000acttgcagca gaatttgcct ctaaaccttg
gatccaggtg gacatgcaaa aggaagtcat 6060aatcacaggg atccagaccc
aaggtgccaa acactacctg aagtcctgct ataccacaga 6120gttctatgta
gcttacagtt ccaaccagat caactggcag atcttcaaag ggaacagcac
6180aaggaatgtg atgtatttta atggcaattc agatgcctct acaataaaag
agaatcagtt 6240tgacccacct attgtggcta gatatattag gatctctcca
actcgagcct ataacagacc 6300tacccttcga ttggaactgc aaggttgtga
ggtaaatgga tgttccacac ccctgggtat 6360ggaaaatgga aagatagaaa
acaagcaaat cacagcttct tcgtttaaga aatcttggtg 6420gggagattac
tgggaaccct tccgtgcccg tctgaatgcc cagggacgtg tgaatgcctg
6480gcaagccaag gcaaacaaca ataagcagtg gctagaaatt gatctactca
agatcaagaa 6540gataacggca attataacac agggctgcaa gtctctgtcc
tctgaaatgt atgtaaagag 6600ctataccatc cactacagtg agcagggagt
ggaatggaaa ccatacaggc tgaaatcctc 6660catggtggac aagatttttg
aaggaaatac taataccaaa ggacatgtga agaacttttt 6720caacccccca
atcatttcca ggtttatccg tgtcattcct aaaacatgga atcaaagtat
6780tgcacttcgc ctggaactct ttggctgtga tatttactag aattgaacat
tcaaaaaccc 6840ctggaagaga ctctttaaga cctcaaacca tttagaatgg
gcaatgtatt ttacgctgtg 6900ttaaatgtta acagttttcc actatttctc
tttcttttct attagtgaat aaaattttat 6960acaagaagct tttataatgt
aactccttgc taccagtaag taagataatg gctattactt 7020ctgcattaat
ttgaatacag gtaggaaaat atcaagaacc aacaagaaaa gggcttatct
7080ttcttaatga ttgaaaatgc tatgaagtaa tatttatgta gttaaaatgc
ttcattataa 7140ctcttttaaa tcctttacac actagtaaaa cagatattac
tttaaataat aattgataga 7200cctggataac tttcacaaac acatgatttt
ttaatggttt ttcttgagtg aagagaaaaa 7260caatattatc aaatgaaata
agtacttaaa atatcctgtc tttcccatat aacaatgatt 7320tttctgactt
tccatgagta aaaaaacagc caagcatctt tccagtagcc ccattgaaat
7380tgtgaatccg tcctggtctc cctaaggact gcacacattg atattcaagg
ttggtggtca 7440ttagatatgg aacagaactg aaataaccat ggtagaactg
aatgtgtaat gttggcttta 7500ttctagctgg tactacatgg cacacagttt
caaaacataa tttcacctac tggaaagctc 7560agacctgtaa aacagagcat
gggaactgct ggtctaaatg cagttgttcc tgctcaaaga 7620gacctctggc
caaactggca agcagttaaa gttttctttc agggccttcc tctctatggc
7680ctcaacttcc tcctctctct tcttccagca acttcccctt tcatcattcc
tttccctggg 7740gacttggcat tcagtgatcc tgtagatatt gcacaactgg
ggaaccttta gacatcctta 7800aaatcacatg agatagacag tcatttgggg
tgtctgaaat aaaccacccc aaaacttagt 7860gttaaaagag caaccaaaaa
aaatttatgt gagattatgg atttgttact tagcttgatt 7920taatcatcct
gtaacgtgta catatatcaa aatgttatgt ataccataaa tatataaaat
7980tttatcaacg aaattcataa caatctctca gaccacagag aaatcaaatt
agaactgagg 8040actaagaaac tcactcgaaa ccacacaact acatggaaac
tgaacaacct gctcctgaat 8100gactactggg taaataatga aattaaggca
gaaataaata agttccttaa aaccaatgag 8160aacaaagaga caacatacca
gaatctctag gagacagggc tttgcttttg ctgcattcta 8220ttcgttgtga
acacaaatta caggccagtc tcgattcagt gtagaaggga actgcataag
8280gaccacatac caggaggcat aattcactgg gagcatcttt agaaactacc
agagttacct 8340gttgcccata ccagtggggt aagccctatg aatgtatatg
agagtttcaa acatccacaa 8400aacattggct ttctaatatt cgtattccca
ctattccttt cttttcatga ttcatgtcat 8460tgtcccatca acatttctaa
gatttccatt ccgttaagag caaaagagaa tgttggaagg 8520tgggggaaaa
catttctttg ttttctacag ggccagcttc ttggatgtgt gtgatctgtt
8580cagttgcaaa gggtcacatg ctcagaagga ccgcatgcta aatttaatgc
tttgcagtta 8640ccctcttgaa atcctttatt ttttaagaag gaattcgaca
tttccatttt tcaatgagcc 8700ccacaaatta cgcagctagt cctgggcttc
tctactctga aattgggcag gatctctctt 8760gatctagaat ttactaaggc
ataatagggg caagaaaatc ttatgaaata atggggggta 8820gggaagagat
gggaatggag catgagatcc agcttcgtta ttctctactt gagaaaaata
8880aggccccaaa gattaaacaa cttgcccaag gatattgctt gttagtgtca
gaactgaaac 8940cagaaaccaa atgatcatat ccctagactt ttagtctgct
ttctcttcca taaaatgaaa 9000cttataatgt ttctaatcca ttgctcagac
aggtagacat gaatattaat tgataatgac 9060tattaattga tctggaaaat
acttgtttgg ggatcaataa tatgtttggg ctattatcta 9120atgctgtgta
gaaatattaa aacccctgtt attttgaaat aaaaaagata cccactttt
9179162224PRTHomo sapiens 16Met Phe Pro Gly Cys Pro Arg Leu Trp Val
Leu Val Val Leu Gly Thr1 5 10 15Ser Trp Val Gly Trp Gly Ser Gln Gly
Thr Glu Ala Ala Gln Leu Arg 20 25 30Gln Phe Tyr Val Ala Ala Gln Gly
Ile Ser Trp Ser Tyr Arg Pro Glu 35 40 45Pro Thr Asn Ser Ser Leu Asn
Leu Ser Val Thr Ser Phe Lys Lys Ile 50 55 60Val Tyr Arg Glu Tyr Glu
Pro Tyr Phe Lys Lys Glu Lys Pro Gln Ser65 70 75 80Thr Ile Ser Gly
Leu Leu Gly Pro Thr Leu Tyr Ala Glu Val Gly Asp 85 90 95Ile Ile Lys
Val His Phe Lys Asn Lys Ala Asp Lys Pro Leu Ser Ile 100 105 110His
Pro Gln Gly Ile Arg Tyr Ser Lys Leu Ser Glu Gly Ala Ser Tyr 115 120
125Leu Asp His Thr Phe Pro Ala Glu Lys Met Asp Asp Ala Val Ala Pro
130 135 140Gly Arg Glu Tyr Thr Tyr Glu Trp Ser Ile Ser Glu Asp Ser
Gly Pro145 150 155 160Thr His Asp Asp Pro Pro Cys Leu Thr His Ile
Tyr Tyr Ser His Glu 165 170 175Asn Leu Ile Glu Asp Phe Asn Ser Gly
Leu Ile Gly Pro Leu Leu Ile 180 185 190Cys Lys Lys Gly Thr Leu Thr
Glu Gly Gly Thr Gln Lys Thr Phe Asp 195 200 205Lys Gln Ile Val Leu
Leu Phe Ala Val Phe Asp Glu Ser Lys Ser Trp 210 215 220Ser Gln Ser
Ser Ser Leu Met Tyr Thr Val Asn Gly Tyr Val Asn Gly225 230 235
240Thr Met Pro Asp Ile Thr Val Cys Ala His Asp His Ile Ser Trp His
245 250 255Leu Leu Gly Met Ser Ser Gly Pro Glu Leu Phe Ser Ile His
Phe Asn 260 265 270Gly Gln Val Leu Glu Gln Asn His His Lys Val Ser
Ala Ile Thr Leu 275 280 285Val Ser Ala Thr Ser Thr Thr Ala Asn Met
Thr Val Gly Pro Glu Gly 290 295 300Lys Trp Ile Ile Ser Ser Leu Thr
Pro Lys His Leu Gln Ala Gly Met305 310 315 320Gln Ala Tyr Ile Asp
Ile Lys Asn Cys Pro Lys Lys Thr Arg Asn Leu 325 330 335Lys Lys Ile
Thr Arg Glu Gln Arg Arg His Met Lys Arg Trp Glu Tyr 340 345 350Phe
Ile Ala Ala Glu Glu Val Ile Trp Asp Tyr Ala Pro Val Ile Pro 355 360
365Ala Asn Met Asp Lys Lys Tyr Arg Ser Gln His Leu Asp Asn Phe Ser
370 375 380Asn Gln Ile Gly Lys His Tyr Lys Lys Val Met Tyr Thr Gln
Tyr Glu385 390 395 400Asp Glu Ser Phe Thr Lys His Thr Val Asn Pro
Asn Met Lys Glu Asp 405 410 415Gly Ile Leu Gly Pro Ile Ile Arg Ala
Gln Val Arg Asp Thr Leu Lys 420 425 430Ile Val Phe Lys Asn Met Ala
Ser Arg Pro Tyr Ser Ile Tyr Pro His 435 440 445Gly Val Thr Phe Ser
Pro Tyr Glu Asp Glu Val Asn Ser Ser Phe Thr 450 455 460Ser Gly Arg
Asn Asn Thr Met Ile Arg Ala Val Gln Pro Gly Glu Thr465 470 475
480Tyr Thr Tyr Lys Trp Asn Ile Leu Glu Phe Asp Glu Pro Thr Glu Asn
485 490 495Asp Ala Gln Cys Leu Thr Arg Pro Tyr Tyr Ser Asp Val Asp
Ile Met 500 505 510Arg Asp Ile Ala Ser Gly Leu Ile Gly Leu Leu Leu
Ile Cys Lys Ser 515 520 525Arg Ser Leu Asp Arg Arg Gly Ile Gln Arg
Ala Ala Asp Ile Glu Gln 530 535 540Gln Ala Val Phe Ala Val Phe Asp
Glu Asn Lys Ser Trp Tyr Leu Glu545 550 555 560Asp Asn Ile Asn Lys
Phe Cys Glu Asn Pro Asp Glu Val Lys Arg Asp 565 570 575Asp Pro Lys
Phe Tyr Glu Ser Asn Ile Met Ser Thr Ile Asn Gly Tyr 580 585 590Val
Pro Glu Ser Ile Thr Thr Leu Gly Phe Cys Phe Asp Asp Thr Val 595 600
605Gln Trp His Phe Cys Ser Val Gly Thr Gln Asn Glu Ile Leu Thr Ile
610 615 620His Phe Thr Gly His Ser Phe Ile Tyr Gly Lys Arg His Glu
Asp Thr625 630 635 640Leu Thr Leu Phe Pro Met Arg Gly Glu Ser Val
Thr Val Thr Met Asp 645 650 655Asn Val Gly Thr Trp Met Leu Thr Ser
Met Asn Ser Ser Pro Arg Ser 660 665 670Lys Lys Leu Arg Leu Lys Phe
Arg Asp Val Lys Cys Ile Pro Asp Asp 675 680 685Asp Glu Asp Ser Tyr
Glu Ile Phe Glu Pro Pro Glu Ser Thr Val Met 690 695 700Ala Thr Arg
Lys Met His Asp Arg Leu Glu Pro Glu Asp Glu Glu Ser705 710 715
720Asp Ala Asp Tyr Asp Tyr Gln Asn Arg Leu Ala Ala Ala Leu Gly Ile
725 730 735Arg Ser Phe Arg Asn Ser Ser Leu Asn Gln Glu Glu Glu Glu
Phe Asn 740 745 750Leu Thr Ala Leu Ala Leu Glu Asn Gly Thr Glu Phe
Val Ser Ser Asn 755 760 765Thr Asp Ile Ile Val Gly Ser Asn Tyr Ser
Ser Pro Ser Asn Ile Ser 770 775 780Lys Phe Thr Val Asn Asn Leu Ala
Glu Pro Gln Lys Ala Pro Ser His785 790 795 800Gln Gln Ala Thr Thr
Ala Gly Ser Pro Leu Arg His Leu Ile Gly Lys 805 810 815Asn Ser Val
Leu Asn Ser Ser Thr Ala Glu His Ser Ser Pro Tyr Ser 820 825 830Glu
Asp Pro Ile Glu Asp Pro Leu Gln Pro Asp Val Thr Gly Ile Arg 835 840
845Leu Leu Ser Leu Gly Ala Gly Glu Phe Lys Ser Gln Glu His Ala Lys
850 855 860His Lys Gly Pro Lys Val Glu Arg Asp Gln Ala Ala Lys His
Arg Phe865 870 875 880Ser Trp Met Lys Leu Leu Ala His Lys Val Gly
Arg His Leu Ser Gln 885 890 895Asp Thr Gly Ser Pro Ser Gly Met Arg
Pro Trp Glu Asp Leu Pro Ser 900 905 910Gln Asp Thr Gly Ser Pro Ser
Arg Met Arg Pro Trp Lys Asp Pro Pro 915 920 925Ser Asp Leu Leu Leu
Leu Lys Gln Ser Asn Ser Ser Lys Ile Leu Val 930 935 940Gly Arg Trp
His Leu Ala Ser Glu Lys Gly Ser Tyr Glu Ile Ile Gln945 950 955
960Asp Thr Asp Glu Asp Thr Ala Val Asn Asn Trp Leu Ile Ser Pro Gln
965 970 975Asn Ala Ser Arg Ala Trp Gly Glu Ser Thr Pro Leu Ala Asn
Lys Pro 980 985 990Gly Lys Gln Ser Gly His Pro Lys Phe Pro Arg Val
Arg His Lys Ser 995 1000 1005Leu Gln Val Arg Gln Asp Gly Gly Lys
Ser Arg Leu Lys Lys Ser 1010 1015 1020Gln Phe Leu Ile Lys Thr Arg
Lys Lys Lys Lys Glu Lys His Thr 1025 1030 1035His His Ala Pro Leu
Ser Pro Arg Thr Phe His Pro Leu Arg Ser 1040 1045 1050Glu Ala Tyr
Asn Thr Phe Ser Glu Arg Arg Leu Lys His Ser Leu 1055 1060 1065Val
Leu His Lys Ser Asn Glu Thr Ser Leu Pro Thr Asp Leu Asn 1070 1075
1080Gln Thr Leu Pro Ser Met Asp Phe Gly Trp Ile Ala Ser Leu Pro
1085 1090 1095Asp His Asn Gln Asn Ser Ser Asn Asp Thr Gly Gln Ala
Ser Cys 1100 1105 1110Pro Pro Gly Leu Tyr Gln Thr Val Pro Pro Glu
Glu His Tyr Gln 1115 1120 1125Thr Phe Pro Ile Gln Asp Pro Asp Gln
Met His Ser Thr Ser Asp 1130 1135 1140Pro Ser His Arg Ser Ser Ser
Pro Glu Leu Ser Glu Met Leu Glu 1145 1150 1155Tyr Asp Arg Ser His
Lys Ser Phe Pro Thr Asp Ile Ser Gln Met 1160 1165 1170Ser Pro Ser
Ser Glu His Glu Val Trp Gln Thr Val Ile Ser Pro 1175 1180 1185Asp
Leu Ser Gln Val Thr Leu Ser Pro Glu Leu Ser Gln Thr Asn 1190 1195
1200Leu Ser Pro Asp Leu Ser His Thr Thr Leu Ser Pro Glu Leu Ile
1205 1210 1215Gln Arg Asn Leu Ser Pro Ala Leu Gly Gln Met Pro Ile
Ser Pro 1220 1225 1230Asp Leu Ser His Thr Thr Leu Ser Pro Asp Leu
Ser His Thr Thr 1235 1240 1245Leu Ser Leu Asp Leu Ser Gln Thr Asn
Leu Ser Pro Glu Leu Ser 1250 1255 1260Gln Thr Asn Leu Ser Pro Ala
Leu Gly Gln Met Pro Leu Ser Pro 1265 1270 1275Asp Leu Ser His Thr
Thr Leu Ser Leu Asp Phe Ser Gln Thr Asn 1280 1285 1290Leu Ser Pro
Glu Leu Ser His Met Thr Leu Ser Pro Glu Leu Ser 1295 1300 1305Gln
Thr Asn Leu Ser Pro Ala Leu Gly Gln Met Pro Ile Ser Pro 1310 1315
1320Asp Leu Ser His Thr Thr Leu Ser Leu Asp Phe Ser Gln Thr Asn
1325 1330 1335Leu Ser Pro Glu Leu Ser Gln Thr Asn Leu Ser Pro Ala
Leu Gly 1340 1345 1350Gln Met Pro Leu Ser Pro Asp Pro Ser His Thr
Thr Leu Ser Leu 1355 1360 1365Asp Leu Ser Gln Thr Asn Leu Ser Pro
Glu Leu Ser Gln Thr Asn 1370 1375 1380Leu Ser Pro Asp Leu Ser Glu
Met Pro Leu Phe Ala Asp Leu Ser 1385 1390 1395Gln Ile Pro Leu Thr
Pro Asp Leu Asp Gln Met Thr Leu Ser Pro 1400 1405 1410Asp Leu Gly
Glu Thr Asp Leu Ser Pro Asn Phe Gly Gln Met Ser 1415 1420 1425Leu
Ser Pro Asp Leu Ser Gln Val Thr Leu Ser Pro Asp Ile Ser 1430 1435
1440Asp Thr Thr Leu Leu Pro Asp Leu Ser Gln Ile Ser Pro Pro Pro
1445
1450 1455Asp Leu Asp Gln Ile Phe Tyr Pro Ser Glu Ser Ser Gln Ser
Leu 1460 1465 1470Leu Leu Gln Glu Phe Asn Glu Ser Phe Pro Tyr Pro
Asp Leu Gly 1475 1480 1485Gln Met Pro Ser Pro Ser Ser Pro Thr Leu
Asn Asp Thr Phe Leu 1490 1495 1500Ser Lys Glu Phe Asn Pro Leu Val
Ile Val Gly Leu Ser Lys Asp 1505 1510 1515Gly Thr Asp Tyr Ile Glu
Ile Ile Pro Lys Glu Glu Val Gln Ser 1520 1525 1530Ser Glu Asp Asp
Tyr Ala Glu Ile Asp Tyr Val Pro Tyr Asp Asp 1535 1540 1545Pro Tyr
Lys Thr Asp Val Arg Thr Asn Ile Asn Ser Ser Arg Asp 1550 1555
1560Pro Asp Asn Ile Ala Ala Trp Tyr Leu Arg Ser Asn Asn Gly Asn
1565 1570 1575Arg Arg Asn Tyr Tyr Ile Ala Ala Glu Glu Ile Ser Trp
Asp Tyr 1580 1585 1590Ser Glu Phe Val Gln Arg Glu Thr Asp Ile Glu
Asp Ser Asp Asp 1595 1600 1605Ile Pro Glu Asp Thr Thr Tyr Lys Lys
Val Val Phe Arg Lys Tyr 1610 1615 1620Leu Asp Ser Thr Phe Thr Lys
Arg Asp Pro Arg Gly Glu Tyr Glu 1625 1630 1635Glu His Leu Gly Ile
Leu Gly Pro Ile Ile Arg Ala Glu Val Asp 1640 1645 1650Asp Val Ile
Gln Val Arg Phe Lys Asn Leu Ala Ser Arg Pro Tyr 1655 1660 1665Ser
Leu His Ala His Gly Leu Ser Tyr Glu Lys Ser Ser Glu Gly 1670 1675
1680Lys Thr Tyr Glu Asp Asp Ser Pro Glu Trp Phe Lys Glu Asp Asn
1685 1690 1695Ala Val Gln Pro Asn Ser Ser Tyr Thr Tyr Val Trp His
Ala Thr 1700 1705 1710Glu Arg Ser Gly Pro Glu Ser Pro Gly Ser Ala
Cys Arg Ala Trp 1715 1720 1725Ala Tyr Tyr Ser Ala Val Asn Pro Glu
Lys Asp Ile His Ser Gly 1730 1735 1740Leu Ile Gly Pro Leu Leu Ile
Cys Gln Lys Gly Ile Leu His Lys 1745 1750 1755Asp Ser Asn Met Pro
Val Asp Met Arg Glu Phe Val Leu Leu Phe 1760 1765 1770Met Thr Phe
Asp Glu Lys Lys Ser Trp Tyr Tyr Glu Lys Lys Ser 1775 1780 1785Arg
Ser Ser Trp Arg Leu Thr Ser Ser Glu Met Lys Lys Ser His 1790 1795
1800Glu Phe His Ala Ile Asn Gly Met Ile Tyr Ser Leu Pro Gly Leu
1805 1810 1815Lys Met Tyr Glu Gln Glu Trp Val Arg Leu His Leu Leu
Asn Ile 1820 1825 1830Gly Gly Ser Gln Asp Ile His Val Val His Phe
His Gly Gln Thr 1835 1840 1845Leu Leu Glu Asn Gly Asn Lys Gln His
Gln Leu Gly Val Trp Pro 1850 1855 1860Leu Leu Pro Gly Ser Phe Lys
Thr Leu Glu Met Lys Ala Ser Lys 1865 1870 1875Pro Gly Trp Trp Leu
Leu Asn Thr Glu Val Gly Glu Asn Gln Arg 1880 1885 1890Ala Gly Met
Gln Thr Pro Phe Leu Ile Met Asp Arg Asp Cys Arg 1895 1900 1905Met
Pro Met Gly Leu Ser Thr Gly Ile Ile Ser Asp Ser Gln Ile 1910 1915
1920Lys Ala Ser Glu Phe Leu Gly Tyr Trp Glu Pro Arg Leu Ala Arg
1925 1930 1935Leu Asn Asn Gly Gly Ser Tyr Asn Ala Trp Ser Val Glu
Lys Leu 1940 1945 1950Ala Ala Glu Phe Ala Ser Lys Pro Trp Ile Gln
Val Asp Met Gln 1955 1960 1965Lys Glu Val Ile Ile Thr Gly Ile Gln
Thr Gln Gly Ala Lys His 1970 1975 1980Tyr Leu Lys Ser Cys Tyr Thr
Thr Glu Phe Tyr Val Ala Tyr Ser 1985 1990 1995Ser Asn Gln Ile Asn
Trp Gln Ile Phe Lys Gly Asn Ser Thr Arg 2000 2005 2010Asn Val Met
Tyr Phe Asn Gly Asn Ser Asp Ala Ser Thr Ile Lys 2015 2020 2025Glu
Asn Gln Phe Asp Pro Pro Ile Val Ala Arg Tyr Ile Arg Ile 2030 2035
2040Ser Pro Thr Arg Ala Tyr Asn Arg Pro Thr Leu Arg Leu Glu Leu
2045 2050 2055Gln Gly Cys Glu Val Asn Gly Cys Ser Thr Pro Leu Gly
Met Glu 2060 2065 2070Asn Gly Lys Ile Glu Asn Lys Gln Ile Thr Ala
Ser Ser Phe Lys 2075 2080 2085Lys Ser Trp Trp Gly Asp Tyr Trp Glu
Pro Phe Arg Ala Arg Leu 2090 2095 2100Asn Ala Gln Gly Arg Val Asn
Ala Trp Gln Ala Lys Ala Asn Asn 2105 2110 2115Asn Lys Gln Trp Leu
Glu Ile Asp Leu Leu Lys Ile Lys Lys Ile 2120 2125 2130Thr Ala Ile
Ile Thr Gln Gly Cys Lys Ser Leu Ser Ser Glu Met 2135 2140 2145Tyr
Val Lys Ser Tyr Thr Ile His Tyr Ser Glu Gln Gly Val Glu 2150 2155
2160Trp Lys Pro Tyr Arg Leu Lys Ser Ser Met Val Asp Lys Ile Phe
2165 2170 2175Glu Gly Asn Thr Asn Thr Lys Gly His Val Lys Asn Phe
Phe Asn 2180 2185 2190Pro Pro Ile Ile Ser Arg Phe Ile Arg Val Ile
Pro Lys Thr Trp 2195 2200 2205Asn Gln Ser Ile Ala Leu Arg Leu Glu
Leu Phe Gly Cys Asp Ile 2210 2215 2220Tyr176910DNABos Taurus
17agcctctggg agctcactgc agcccggcct gcggacagcc tcgcagaggc agccctaacc
60cacccccggg gtggtggcgg caggcaagag aaggaaagga accatgttcc tcgcttgccc
120tggcttctgg gtcctcgtgg tcctaggcag cagctgggca ggctggggga
acctaggggc 180tgaagcagca aagctaaggc agttctacgt agctgctcag
agcatcagat ggaactaccg 240ccccgagtcc acacacctca gttcgaaacc
ttttgaaacc tcctttaaga aaattgtcta 300cagggagtat gaagcatatt
ttcagaaaga aaaaccacaa tccagaactt caggacttct 360tgggcctact
ttgtatgctg aagttggaga catcatgaaa gttcacttta agaataaagc
420acacaagccc ttaagcatcc atgctcaagg aattaagtac agtaaattct
cagaaggtgc 480gtcttactct gaccacacac tccccatgga gaagatggat
gatgctgtag ctccgggcca 540agaatatacc tatgagtgga ttatcagtga
gcacagtggg cccacccacg atgaccctcc 600atgcctcaca cacatctatt
actcctatgt aaatctggtg gaggacttca actctggact 660gattggacct
ctgcttattt gtaagaaagg caccctaacc gaggatggaa ctcagaaaat
720gtttgagaag caacatgtac tgatgtttgc tgtgtttgat gaaagtaaaa
gctggaacca 780gacatcatcc ttaatgtaca cagtcaatgg ctatgtgaat
gggacgatgc cagatataac 840agtctgtgcc catgaccaca tcagttggca
tctgattgga atgagctctg ggccagaact 900gttctccatc catttcaatg
gtcaggtcct ggagcagaac catcataaga tctcagccat 960cactctcgtc
agcgccacgt ccacaaccgc aaacatgacc gtgagccccg agggaaggtg
1020gaccatagct tctctcatcc ccagacattt tcaagctggg atgcaggctt
acatagacat 1080taaaaactgt gcaaagaaaa ccagaaatcc taagaaacta
actcgagacc agaggcggca 1140cattaagaga tgggaatact tcattgctgc
agaggaagtc atttgggact atgcacctat 1200aataccagca aacatggaca
aaaaatacag atctctgcat ttggataatt tctcaaaccg 1260aattggaaaa
cattataaga aggttgtcta caaacagtac caagatgact ccttcaccaa
1320acgcctggag gatcccagta gtgaaggaga tgggatcttg ggccctatta
tcagagccca 1380ggtcagagac acactgaaaa tcgtgttcaa aaatatggcc
agccgctcct acagcattta 1440ccctcacggt gtgacattct ctccttatga
caatgaagta aactcttcct caacctcagg 1500cagcaacacc atgatcagag
cagttcgacc aggggaaacc tacacttata agtggaacat 1560cctagaatct
gatgaaccca cagaaaatga tgctcagtgc ttaacaagac catactacag
1620taatgtggac atcacaaggg accttgcttc tggactgata gggcttcttc
taatttgtaa 1680gagcagatcc ttggatagac gaggcataca gagggcagca
gacatcgagc agcaggctgt 1740gtttgccgtg tttgacgaga acaagagctg
gtacattgag gacaacatct acaagttttg 1800tgaaaatcct gagaaagtga
aacgtgatga ccccaagttt tatgagtcaa acatcatgag 1860taatttcact
cttccagcta ttaacggcta tgtgcctgag agtataccca tactagggtt
1920ctgctttgat gacactgtcc agtggcactt ctgcagtgtg ggaacccaga
atgacatttt 1980gaccattcac ttcactgggc actcattcat ctatggaaag
aggcacgagg acaccttgac 2040ccttttcccc atgcaggggg aatccgtgac
tgtcacaatg gataatgttg gaacttggat 2100gttaaccacc atgaattcca
atccaagaag caaaaaacta cggctgaggt tcagggatgc 2160taagtgtatc
cggaatgatg atgatgactc ctatgagatt atatatgaac cttcaggatc
2220tacagccatg actacaaaga aaattcatga ttcttcagaa atcgaagatg
aaaatgatgc 2280tgactctgat taccaggacg aactggcttt aatactaggt
cttaggtcat tcagaaattc 2340atcactgaat caggagaaag atgagctcaa
tcttaccgcc ctagctctgg agaaagactc 2400tgaattcatt cctccgagtg
ccaacagatc tcttgattca aattcttctt cccgaagtca 2460tgttagcagg
cttattgcca aaaactttgc agaatctctg aaaactcttc tgcacctgga
2520agcccctgca gctggttccc ccctggaaca cgctggctta gataagaact
cagctctcaa 2580ccctcccatg gcagagcatt ccagccctta ttctgaagac
cctagagaag atcatccact 2640ctcagatgtc acaggggtaa gcctacttcc
atttggcaca ggattcaaaa atcgaaaacc 2700tgccaaacat caaagattcc
aggtaggaag aggccaagca gcaaagcata agttctccca 2760gacgcgattc
ccagcacata aaaccaggac acgtttaagc caagacaact cttcttcttc
2820cagaatgggg ccctgggagg acattcccag tgatctgtta ctcttacaac
aaaaggatcc 2880atataagatt ctgaatggag aatggcattt ggtttctgag
aaaggcagtt atgaaataat 2940ccaagatgct aatgaaaaca agactgttaa
taagttgcca aacagccccc agaatgactc 3000aaggacttgg ggagaaaaca
tccctttcaa aaacagtcat ggaaagcaga gtggccaccc 3060aacatttttg
gtaactagac gtaaacctct acaagacaga caggatagaa gaaatagtag
3120attgaaggaa ggccttccgt taattaggac acgaagaaag aaaaaggaag
agaagcctgc 3180ataccatgtt cctctatctc caaggagttt tcatcctctg
agaggagagg tcaatgcctc 3240attttcagac agaagacata atcattcatt
gttactccat gcgtccaatg aaacatctct 3300ttccatagac ctcaatcaga
cattcccctc tatgaatctt agccttgcag cctcacttcc 3360tgaccatgac
cagacctcac caaatgacac caccagtcag actagctccc ctccagatct
3420ttatccgaca gtgagcccag aggaacacta tcaaatattc cctattcaag
actctgatcc 3480aacacattct actacagccc ccagtaacag atctcctgat
ccaacacatt ctactacagc 3540ccccagtaac agatctcctc ccacacagcc
cagccagata cccaactatg acctaagaaa 3600cagggccatc cctactgatg
tgagtcaaat tttcccttcc ttggaactcg aagtctggca 3660gacagctacc
tctctagacc tcagtcaacc atccatctcc ccagaccttg gccagatggc
3720actttcccca gaccccggcc aggagtctct ctctccagac cttggccaga
cgtccctctc 3780tccagacctc agccaggagt ctctctcccc agaccttggc
cagacagccc tttccccaga 3840ccccagccag gagtctctct ccccagacct
tggccagaca gccctttccc cagaccccag 3900ccaggagtct ctctccccag
accttggcca gacagccctt tccccagacc ccggccagga 3960gtctctctct
ccagaccttg gccagacgtc cctctctcca gacctcagcc aggagtctct
4020ctccccagac cttggccaga cagccctttc cccagacccc agccaggagt
ctctctcccc 4080agaccttggc cagacagccc tttccccaga ccccagccag
gagtctctct ctccagacct 4140tggccagacg tccctctctc cagaccttgg
ccaggagtct ctctccccag accttggcca 4200gacagccctt tccccagacc
ccagccagga gtctctctct ccagaccttg gccagacgtc 4260cctctctcca
gaccttggcc aggagtctct ctccccagac cttggccaga cagccctttc
4320cccagacctc agccaggagt ctctctctcc agatcttggc cagacacccc
tctctccaga 4380cctcagcctg gagtctcttt ctccagacct cagccagctt
gatctcaagc agacatcacc 4440tcctctagat cttaatcaga catcccacac
ttctgaatca agtcagtcat tgcctcttcc 4500agaatttggt cagactttcc
ctaatgcaga tattggtcag atgccatctc ctccaccaga 4560ctctacacta
aataacactt ttataccaga agaatttaat ccgctggttg tagtaggcct
4620cagtagagat gatggagatt atattgaaat tattccaagg cagaaggaag
agagcagtga 4680agaagactat ggtgaatttg agtttgtagc ctataatgac
ccttaccaaa ctgatcttag 4740gacagatatc aactcctcca gaaatcctga
caacattgca gcatggtacc tccgcagcaa 4800cactggaaac agaaaatatt
attacattgc agctgaagaa atatcctggg attattcaaa 4860atttgtgcaa
agtgatgacg ttgactatgt tccagaggac accgtataca agaaagtagt
4920tttccgaaag taccttgata gcacttttac caaacttgat cctcaggggg
agtatgaaga 4980gcatcttggc atacttggtc cagtcattag agctgaagtg
gatgatgtta tccaagttcg 5040ttttaaaaat ttagcatcca gaccatattc
tcttcatgcc catgggcttt cctatgaaaa 5100atcatcagaa ggaaagactt
atgaagatga ctctcctgaa tggtttaagg aggacaatgc 5160tattcagccc
aataaaactt acacctatgt atggcacgcc actacgcgat ccgggccaga
5220aaaccctgga tctgcctgtc gggcttgggc ctactactca gcagtgaacc
cagaaaaaga 5280catccattca ggcttgatag ggcctcttct gatctgccga
aaagggacac ttgataagga 5340gaccaacatg cctgtggaca tgagagaatt
tgtcctgctt tttatggtct ttgatgaaaa 5400gaagagctgg tattatgaca
agaagcccac aaggtcttgg agacgtgcat cctcagaagt 5460aaaaaactcc
catgagtttc atgccatcaa tgggatgatc tacaacttgc ctggcttgag
5520aatgtacgag caagagtggg tgaggttgca cctgctgaac ttaggcggct
cccgagacat 5580tcacgtggtt cactttcatg gccagacctt gctagaaaac
ggcactcaac agcaccagtt 5640aggggtctgg ccccttctgc ctggttcatt
taaaactctt gaaatgaagg catcaaaacc 5700tggctggtgg ctcctagaca
cggaagttgg agaaattcag agagcaggga tgcagacacc 5760atttctcatt
gtagacagag aatgtaagat gccaatggga ctaagcactg gcctgatagc
5820tgactcacag atccaggctt ctgagttttg gggttattgg gaacccaaat
tagcaaggtt 5880aaacaatggt ggatcataca atgcttggat tgcagaaaaa
ctttcaacgg aatttaaccc 5940tgaaccttgg atccaggtag acatgcaaaa
ggaagtcctg ctcacgggga tccagaccca 6000gggcgccaaa cactacctga
agccctacta caccaccgag ttctgtgtgg cttacagctt 6060ggatcggaaa
aactggcgta tcttcaaagg gaacagcaca aggaatgtga tgtattttgg
6120tggcaattca gatgcttcta caataaaaga gaatcagatt gacccacctg
ttgtggctag 6180atacattagg atctctccaa ctggatccta taacaaacct
gcccttcgat tggagctgca 6240aggttgtgag gttaatggat gctccacacc
gctgggtatg gaaagtggaa agatagaaaa 6300caagcaaatc accgcttcct
cgtttaaaaa gtcttggtgg ggaaattact gggaaccctt 6360ccttgcacgt
cttaatgccc agggccgtgt aaatgcctgg caagctaagg caaacaacaa
6420caatcagtgg ttacaaattg atctgctcaa aatcaagaag ataactgcga
ttgtaacaca 6480aggatgcaag tctctgtcct ctgaaatgta tgtgaagagc
tacaccatcc actacagtga 6540ccagggaacg gactggaaac cttacaggga
gaaatcctca atggtggaca agattttcga 6600aggaaataat aatgtcagag
gacatgtgaa gaactttttc aacccaccaa tcatctccag 6660gtttatacgc
atcattccta aaacatggaa tcagagtatt gcacttcgct tggaactctt
6720tggctgtgat atgtactaga attgaatatt ttaaaagata ggagggactc
aaagatatca 6780aaccacttag agtgggcaat gcattttgta gctattttaa
gtataaaaaa atttccatta 6840tttctctttt ttctattaga gaataaaatt
ttatatgcaa aacctttatg atataactcc 6900tgataaccac 6910182211PRTBos
Taurus 18Met Phe Leu Ala Cys Pro Gly Phe Trp Val Leu Val Val Leu
Gly Ser1 5 10 15Ser Trp Ala Gly Trp Gly Asn Leu Gly Ala Glu Ala Ala
Lys Leu Arg 20 25 30Gln Phe Tyr Val Ala Ala Gln Ser Ile Arg Trp Asn
Tyr Arg Pro Glu 35 40 45Ser Thr His Leu Ser Ser Lys Pro Phe Glu Thr
Ser Phe Lys Lys Ile 50 55 60Val Tyr Arg Glu Tyr Glu Ala Tyr Phe Gln
Lys Glu Lys Pro Gln Ser65 70 75 80Arg Thr Ser Gly Leu Leu Gly Pro
Thr Leu Tyr Ala Glu Val Gly Asp 85 90 95Ile Met Lys Val His Phe Lys
Asn Lys Ala His Lys Pro Leu Ser Ile 100 105 110His Ala Gln Gly Ile
Lys Tyr Ser Lys Phe Ser Glu Gly Ala Ser Tyr 115 120 125Ser Asp His
Thr Leu Pro Met Glu Lys Met Asp Asp Ala Val Ala Pro 130 135 140Gly
Gln Glu Tyr Thr Tyr Glu Trp Ile Ile Ser Glu His Ser Gly Pro145 150
155 160Thr His Asp Asp Pro Pro Cys Leu Thr His Ile Tyr Tyr Ser Tyr
Val 165 170 175Asn Leu Val Glu Asp Phe Asn Ser Gly Leu Ile Gly Pro
Leu Leu Ile 180 185 190Cys Lys Lys Gly Thr Leu Thr Glu Asp Gly Thr
Gln Lys Met Phe Glu 195 200 205Lys Gln His Val Leu Met Phe Ala Val
Phe Asp Glu Ser Lys Ser Trp 210 215 220Asn Gln Thr Ser Ser Leu Met
Tyr Thr Val Asn Gly Tyr Val Asn Gly225 230 235 240Thr Met Pro Asp
Ile Thr Val Cys Ala His Asp His Ile Ser Trp His 245 250 255Leu Ile
Gly Met Ser Ser Gly Pro Glu Leu Phe Ser Ile His Phe Asn 260 265
270Gly Gln Val Leu Glu Gln Asn His His Lys Ile Ser Ala Ile Thr Leu
275 280 285Val Ser Ala Thr Ser Thr Thr Ala Asn Met Thr Val Ser Pro
Glu Gly 290 295 300Arg Trp Thr Ile Ala Ser Leu Ile Pro Arg His Phe
Gln Ala Gly Met305 310 315 320Gln Ala Tyr Ile Asp Ile Lys Asn Cys
Ala Lys Lys Thr Arg Asn Pro 325 330 335Lys Lys Leu Thr Arg Asp Gln
Arg Arg His Ile Lys Arg Trp Glu Tyr 340 345 350Phe Ile Ala Ala Glu
Glu Val Ile Trp Asp Tyr Ala Pro Ile Ile Pro 355 360 365Ala Asn Met
Asp Lys Lys Tyr Arg Ser Leu His Leu Asp Asn Phe Ser 370 375 380Asn
Arg Ile Gly Lys His Tyr Lys Lys Val Val Tyr Lys Gln Tyr Gln385 390
395 400Asp Asp Ser Phe Thr Lys Arg Leu Glu Asp Pro Ser Ser Glu Gly
Asp 405 410 415Gly Ile Leu Gly Pro Ile Ile Arg Ala Gln Val Arg Asp
Thr Leu Lys 420 425 430Ile Val Phe Lys Asn Met Ala Ser Arg Ser Tyr
Ser Ile Tyr Pro His 435 440 445Gly Val Thr Phe Ser Pro Tyr Asp Asn
Glu Val Asn Ser Ser Ser Thr 450 455 460Ser Gly Ser Asn Thr Met Ile
Arg Ala Val Arg Pro Gly Glu Thr Tyr465 470 475 480Thr Tyr Lys Trp
Asn Ile Leu Glu Ser Asp Glu Pro Thr Glu Asn Asp 485 490 495Ala Gln
Cys Leu Thr Arg Pro Tyr Tyr Ser Asn Val Asp Ile Thr Arg 500 505
510Asp Leu Ala Ser Gly Leu Ile Gly Leu Leu Leu Ile Cys Lys Ser Arg
515 520 525Ser Leu Asp Arg Arg Gly Ile Gln Arg Ala Ala Asp Ile Glu
Gln Gln 530 535
540Ala Val Phe Ala Val Phe Asp Glu Asn Lys Ser Trp Tyr Ile Glu
Asp545 550 555 560Asn Ile Tyr Lys Phe Cys Glu Asn Pro Glu Lys Val
Lys Arg Asp Asp 565 570 575Pro Lys Phe Tyr Glu Ser Asn Ile Met Ser
Asn Phe Thr Leu Pro Ala 580 585 590Ile Asn Gly Tyr Val Pro Glu Ser
Ile Pro Ile Leu Gly Phe Cys Phe 595 600 605Asp Asp Thr Val Gln Trp
His Phe Cys Ser Val Gly Thr Gln Asn Asp 610 615 620Ile Leu Thr Ile
His Phe Thr Gly His Ser Phe Ile Tyr Gly Lys Arg625 630 635 640His
Glu Asp Thr Leu Thr Leu Phe Pro Met Gln Gly Glu Ser Val Thr 645 650
655Val Thr Met Asp Asn Val Gly Thr Trp Met Leu Thr Thr Met Asn Ser
660 665 670Asn Pro Arg Ser Lys Lys Leu Arg Leu Arg Phe Arg Asp Ala
Lys Cys 675 680 685Ile Arg Asn Asp Asp Asp Asp Ser Tyr Glu Ile Ile
Tyr Glu Pro Ser 690 695 700Gly Ser Thr Ala Met Thr Thr Lys Lys Ile
His Asp Ser Ser Glu Ile705 710 715 720Glu Asp Glu Asn Asp Ala Asp
Ser Asp Tyr Gln Asp Glu Leu Ala Leu 725 730 735Ile Leu Gly Leu Arg
Ser Phe Arg Asn Ser Ser Leu Asn Gln Glu Lys 740 745 750Asp Glu Leu
Asn Leu Thr Ala Leu Ala Leu Glu Lys Asp Ser Glu Phe 755 760 765Ile
Pro Pro Ser Ala Asn Arg Ser Leu Asp Ser Asn Ser Ser Ser Arg 770 775
780Ser His Val Ser Arg Leu Ile Ala Lys Asn Phe Ala Glu Ser Leu
Lys785 790 795 800Thr Leu Leu His Leu Glu Ala Pro Ala Ala Gly Ser
Pro Leu Glu His 805 810 815Ala Gly Leu Asp Lys Asn Ser Ala Leu Asn
Pro Pro Met Ala Glu His 820 825 830Ser Ser Pro Tyr Ser Glu Asp Pro
Arg Glu Asp His Pro Leu Ser Asp 835 840 845Val Thr Gly Val Ser Leu
Leu Pro Phe Gly Thr Gly Phe Lys Asn Arg 850 855 860Lys Pro Ala Lys
His Gln Arg Phe Gln Val Gly Arg Gly Gln Ala Ala865 870 875 880Lys
His Lys Phe Ser Gln Thr Arg Phe Pro Ala His Lys Thr Arg Thr 885 890
895Arg Leu Ser Gln Asp Asn Ser Ser Ser Ser Arg Met Gly Pro Trp Glu
900 905 910Asp Ile Pro Ser Asp Leu Leu Leu Leu Gln Gln Lys Asp Pro
Tyr Lys 915 920 925Ile Leu Asn Gly Glu Trp His Leu Val Ser Glu Lys
Gly Ser Tyr Glu 930 935 940Ile Ile Gln Asp Ala Asn Glu Asn Lys Thr
Val Asn Lys Leu Pro Asn945 950 955 960Ser Pro Gln Asn Asp Ser Arg
Thr Trp Gly Glu Asn Ile Pro Phe Lys 965 970 975Asn Ser His Gly Lys
Gln Ser Gly His Pro Thr Phe Leu Val Thr Arg 980 985 990Arg Lys Pro
Leu Gln Asp Arg Gln Asp Arg Arg Asn Ser Arg Leu Lys 995 1000
1005Glu Gly Leu Pro Leu Ile Arg Thr Arg Arg Lys Lys Lys Glu Glu
1010 1015 1020Lys Pro Ala Tyr His Val Pro Leu Ser Pro Arg Ser Phe
His Pro 1025 1030 1035Leu Arg Gly Glu Val Asn Ala Ser Phe Ser Asp
Arg Arg His Asn 1040 1045 1050His Ser Leu Leu Leu His Ala Ser Asn
Glu Thr Ser Leu Ser Ile 1055 1060 1065Asp Leu Asn Gln Thr Phe Pro
Ser Met Asn Leu Ser Leu Ala Ala 1070 1075 1080Ser Leu Pro Asp His
Asp Gln Thr Ser Pro Asn Asp Thr Thr Ser 1085 1090 1095Gln Thr Ser
Ser Pro Pro Asp Leu Tyr Pro Thr Val Ser Pro Glu 1100 1105 1110Glu
His Tyr Gln Ile Phe Pro Ile Gln Asp Ser Asp Pro Thr His 1115 1120
1125Ser Thr Thr Ala Pro Ser Asn Arg Ser Pro Asp Pro Thr His Ser
1130 1135 1140Thr Thr Ala Pro Ser Asn Arg Ser Pro Pro Thr Gln Pro
Ser Gln 1145 1150 1155Ile Pro Asn Tyr Asp Leu Arg Asn Arg Ala Ile
Pro Thr Asp Val 1160 1165 1170Ser Gln Ile Phe Pro Ser Leu Glu Leu
Glu Val Trp Gln Thr Ala 1175 1180 1185Thr Ser Leu Asp Leu Ser Gln
Pro Ser Ile Ser Pro Asp Leu Gly 1190 1195 1200Gln Met Ala Leu Ser
Pro Asp Pro Gly Gln Glu Ser Leu Ser Pro 1205 1210 1215Asp Leu Gly
Gln Thr Ser Leu Ser Pro Asp Leu Ser Gln Glu Ser 1220 1225 1230Leu
Ser Pro Asp Leu Gly Gln Thr Ala Leu Ser Pro Asp Pro Ser 1235 1240
1245Gln Glu Ser Leu Ser Pro Asp Leu Gly Gln Thr Ala Leu Ser Pro
1250 1255 1260Asp Pro Ser Gln Glu Ser Leu Ser Pro Asp Leu Gly Gln
Thr Ala 1265 1270 1275Leu Ser Pro Asp Pro Gly Gln Glu Ser Leu Ser
Pro Asp Leu Gly 1280 1285 1290Gln Thr Ser Leu Ser Pro Asp Leu Ser
Gln Glu Ser Leu Ser Pro 1295 1300 1305Asp Leu Gly Gln Thr Ala Leu
Ser Pro Asp Pro Ser Gln Glu Ser 1310 1315 1320Leu Ser Pro Asp Leu
Gly Gln Thr Ala Leu Ser Pro Asp Pro Ser 1325 1330 1335Gln Glu Ser
Leu Ser Pro Asp Leu Gly Gln Thr Ser Leu Ser Pro 1340 1345 1350Asp
Leu Gly Gln Glu Ser Leu Ser Pro Asp Leu Gly Gln Thr Ala 1355 1360
1365Leu Ser Pro Asp Pro Ser Gln Glu Ser Leu Ser Pro Asp Leu Gly
1370 1375 1380Gln Thr Ser Leu Ser Pro Asp Leu Gly Gln Glu Ser Leu
Ser Pro 1385 1390 1395Asp Leu Gly Gln Thr Ala Leu Ser Pro Asp Leu
Ser Gln Glu Ser 1400 1405 1410Leu Ser Pro Asp Leu Gly Gln Thr Pro
Leu Ser Pro Asp Leu Ser 1415 1420 1425Leu Glu Ser Leu Ser Pro Asp
Leu Ser Gln Leu Asp Leu Lys Gln 1430 1435 1440Thr Ser Pro Pro Leu
Asp Leu Asn Gln Thr Ser His Thr Ser Glu 1445 1450 1455Ser Ser Gln
Ser Leu Pro Leu Pro Glu Phe Gly Gln Thr Phe Pro 1460 1465 1470Asn
Ala Asp Ile Gly Gln Met Pro Ser Pro Pro Pro Asp Ser Thr 1475 1480
1485Leu Asn Asn Thr Phe Ile Pro Glu Glu Phe Asn Pro Leu Val Val
1490 1495 1500Val Gly Leu Ser Arg Asp Asp Gly Asp Tyr Ile Glu Ile
Ile Pro 1505 1510 1515Arg Gln Lys Glu Glu Ser Ser Glu Glu Asp Tyr
Gly Glu Phe Glu 1520 1525 1530Phe Val Ala Tyr Asn Asp Pro Tyr Gln
Thr Asp Leu Arg Thr Asp 1535 1540 1545Ile Asn Ser Ser Arg Asn Pro
Asp Asn Ile Ala Ala Trp Tyr Leu 1550 1555 1560Arg Ser Asn Thr Gly
Asn Arg Lys Tyr Tyr Tyr Ile Ala Ala Glu 1565 1570 1575Glu Ile Ser
Trp Asp Tyr Ser Lys Phe Val Gln Ser Asp Asp Val 1580 1585 1590Asp
Tyr Val Pro Glu Asp Thr Val Tyr Lys Lys Val Val Phe Arg 1595 1600
1605Lys Tyr Leu Asp Ser Thr Phe Thr Lys Leu Asp Pro Gln Gly Glu
1610 1615 1620Tyr Glu Glu His Leu Gly Ile Leu Gly Pro Val Ile Arg
Ala Glu 1625 1630 1635Val Asp Asp Val Ile Gln Val Arg Phe Lys Asn
Leu Ala Ser Arg 1640 1645 1650Pro Tyr Ser Leu His Ala His Gly Leu
Ser Tyr Glu Lys Ser Ser 1655 1660 1665Glu Gly Lys Thr Tyr Glu Asp
Asp Ser Pro Glu Trp Phe Lys Glu 1670 1675 1680Asp Asn Ala Ile Gln
Pro Asn Lys Thr Tyr Thr Tyr Val Trp His 1685 1690 1695Ala Thr Thr
Arg Ser Gly Pro Glu Asn Pro Gly Ser Ala Cys Arg 1700 1705 1710Ala
Trp Ala Tyr Tyr Ser Ala Val Asn Pro Glu Lys Asp Ile His 1715 1720
1725Ser Gly Leu Ile Gly Pro Leu Leu Ile Cys Arg Lys Gly Thr Leu
1730 1735 1740Asp Lys Glu Thr Asn Met Pro Val Asp Met Arg Glu Phe
Val Leu 1745 1750 1755Leu Phe Met Val Phe Asp Glu Lys Lys Ser Trp
Tyr Tyr Asp Lys 1760 1765 1770Lys Pro Thr Arg Ser Trp Arg Arg Ala
Ser Ser Glu Val Lys Asn 1775 1780 1785Ser His Glu Phe His Ala Ile
Asn Gly Met Ile Tyr Asn Leu Pro 1790 1795 1800Gly Leu Arg Met Tyr
Glu Gln Glu Trp Val Arg Leu His Leu Leu 1805 1810 1815Asn Leu Gly
Gly Ser Arg Asp Ile His Val Val His Phe His Gly 1820 1825 1830Gln
Thr Leu Leu Glu Asn Gly Thr Gln Gln His Gln Leu Gly Val 1835 1840
1845Trp Pro Leu Leu Pro Gly Ser Phe Lys Thr Leu Glu Met Lys Ala
1850 1855 1860Ser Lys Pro Gly Trp Trp Leu Leu Asp Thr Glu Val Gly
Glu Ile 1865 1870 1875Gln Arg Ala Gly Met Gln Thr Pro Phe Leu Ile
Val Asp Arg Glu 1880 1885 1890Cys Lys Met Pro Met Gly Leu Ser Thr
Gly Leu Ile Ala Asp Ser 1895 1900 1905Gln Ile Gln Ala Ser Glu Phe
Trp Gly Tyr Trp Glu Pro Lys Leu 1910 1915 1920Ala Arg Leu Asn Asn
Gly Gly Ser Tyr Asn Ala Trp Ile Ala Glu 1925 1930 1935Lys Leu Ser
Thr Glu Phe Asn Pro Glu Pro Trp Ile Gln Val Asp 1940 1945 1950Met
Gln Lys Glu Val Leu Leu Thr Gly Ile Gln Thr Gln Gly Ala 1955 1960
1965Lys His Tyr Leu Lys Pro Tyr Tyr Thr Thr Glu Phe Cys Val Ala
1970 1975 1980Tyr Ser Leu Asp Arg Lys Asn Trp Arg Ile Phe Lys Gly
Asn Ser 1985 1990 1995Thr Arg Asn Val Met Tyr Phe Gly Gly Asn Ser
Asp Ala Ser Thr 2000 2005 2010Ile Lys Glu Asn Gln Ile Asp Pro Pro
Val Val Ala Arg Tyr Ile 2015 2020 2025Arg Ile Ser Pro Thr Gly Ser
Tyr Asn Lys Pro Ala Leu Arg Leu 2030 2035 2040Glu Leu Gln Gly Cys
Glu Val Asn Gly Cys Ser Thr Pro Leu Gly 2045 2050 2055Met Glu Ser
Gly Lys Ile Glu Asn Lys Gln Ile Thr Ala Ser Ser 2060 2065 2070Phe
Lys Lys Ser Trp Trp Gly Asn Tyr Trp Glu Pro Phe Leu Ala 2075 2080
2085Arg Leu Asn Ala Gln Gly Arg Val Asn Ala Trp Gln Ala Lys Ala
2090 2095 2100Asn Asn Asn Asn Gln Trp Leu Gln Ile Asp Leu Leu Lys
Ile Lys 2105 2110 2115Lys Ile Thr Ala Ile Val Thr Gln Gly Cys Lys
Ser Leu Ser Ser 2120 2125 2130Glu Met Tyr Val Lys Ser Tyr Thr Ile
His Tyr Ser Asp Gln Gly 2135 2140 2145Thr Asp Trp Lys Pro Tyr Arg
Glu Lys Ser Ser Met Val Asp Lys 2150 2155 2160Ile Phe Glu Gly Asn
Asn Asn Val Arg Gly His Val Lys Asn Phe 2165 2170 2175Phe Asn Pro
Pro Ile Ile Ser Arg Phe Ile Arg Ile Ile Pro Lys 2180 2185 2190Thr
Trp Asn Gln Ser Ile Ala Leu Arg Leu Glu Leu Phe Gly Cys 2195 2200
2205Asp Met Tyr 2210191031DNAPseudonaja textilis 19cgacggcccg
ggctggtctg ccaggaagat ttcatgggac tccttatttg cggagacttt 60gcccctgtgg
atccagagag tagaaactct cactcatctc ctcatagggt gtcctttcta
120tgatgctcca agatgtgttt taatcacacc aatcatttcc aaataaggct
gcagagataa 180atgttatgtt tattatattt caaaggtaac ctcacactca
catatcagat ggccaaatat 240cttcgagttc tacaactacc caaaactttt
aattaaatat attgttttat tgccccgttc 300tttcctattc tcaataacaa
atgttgtgtt aaactatttc tctcttcact gtactgtaaa 360agtactgtac
tatactttta ttgtgaaggc tgatcaaaga ttgtaatcaa gatattttag
420ttgaaaaaaa tatgttttac tgtgctctac tttctgctgc tttttgagat
gtgtgagaga 480gagatgcaaa atgcaattct ctctcatgtt caaatattgc
agaaatatac ttacgatagg 540tagtagcatt agcatttgct aggtcttcct
taggaacaag ttgctctgga tgtaggatgt 600ttctttaagg tttctttatg
aaaaactcag agaggaggca gtgaagctct tcccctaagt 660acaatctgtt
ttcaacttct gggtgagctt cctttcaagg tcactatctg tgcttagcag
720tgaggggcag ctctcctttg aggtatccat cccacacccc atactattaa
tcttgtactg 780actcaaatga ccttacttgg taaagacccg cattttgaat
tagtcagcac aatgatctga 840agcatccata gtcaaacaca aacaggcttt
ggaggacatg ataagggctg gagcagaaca 900acaggaagct tgattgcttg
aaccttgttc atagccagcc ctgaaagaga acaaactgtt 960cttttccatc
gataggcacc atggcccctc agctactcct ctgtctgatc ctcacttttc
1020tagggagtct c 1031201010DNAPseudonaja textilis 20cgacggcccg
ggctggtctg ccaggaagat ttcatgggac tccttatttg cggagacttt 60gcccctgtgg
atccagagag tagaaactct cactcatctc ctcatagggt gtcctttcta
120tgatgctcca agatgtgttt taatcacacc aatcatttcc aaataaggct
gcagagataa 180atgttatgtt tattatattt caaaggtaac ctcacactca
catatcagat ggccaaatat 240cttcgagttc tacaactacc caaaactttt
aattaaatat attgttttat tgccccgttc 300tttcctattc tcaataacaa
atgttgtgtt aaactatttc tctcttcact gtactgtaaa 360agtactgtac
tatactttta ttgtgaaggc tgatcaaaga ttgtaatcaa gatattttag
420ttgaaaaaaa tatgttttac tgtgctctac tttctgctgc tttttgagat
gtgtgagaga 480gagatgcaaa atgcaattct ctctcatgtt caaatattgc
agaaatatac ttacgatagg 540tagtagcatt agcatttgct aggtcttcct
taggaacaag ttgctctgga tgtaggatgt 600ttctttatga aaactcagag
aggaggcagt gaagctcttc ccctaagtac aatctgtttt 660caacttctgg
gtgagcttcc tttcaaggtc actatctgtg cttagcagtg aggggcagct
720ctcctttgag gtatccatcc cacaccccat actattaatc ttgtactgac
tcaaatgacc 780ttacttggta aagacccgca ttttgaatta gtcagcacaa
tgatctgaag catccatagt 840caaacacaaa caggctttgg aggacatgat
aagggctgga gcagaacaac aggaagcttg 900attgcttgaa ccttgttcat
agccagccct gaaagagaac aaactgttct tttccatcga 960taggcaccat
ggcccctcag ctactcctct gtctgatcct cacttttcta 1010211299DNAPseudonaja
textilis 21cgacggcccg ggctggtctg ccaggaagat ttcatgggac tccttatttg
cggagacttt 60gcccctgtgg atccagagag tagaaactct cactcatctc ctcatagggt
gtcctttcta 120tgatgctcca agatgtgttt taatcacacc aatcatttcc
aaataaggct gcagagataa 180atgttatgtt tattatattt caaaggtaac
ctcacactca catatcagat ggccaaatat 240cttcgagttc tacaactacc
caaaactttt aattaaatat attgttttat tgccccgttc 300tttcctattc
tcaataacaa atgttgtgtt aaactatttc tctcttcact gtactgtaaa
360agtactgtac tatactttta ttgtgaaggc tgatcaaaga ttgtaatcaa
gatattttag 420ttgaaaaaaa tatgttttac tgtgctctac tttctgctgc
tttttgggat gtgtgagaga 480gagatgcaaa atgcaattct ctctcatgtt
caaatattgc agaaatatac ttacgatagg 540tagtagcatt agcatttgct
aggtcttcct taggaaccag gtgctctgga tgtagggtgt 600ttctttaagg
tttctttatg aagaactcag agaggaggca gggaagctct tccccctaag
660taatctgttt tcaacttctg ggtgagcttc ctttcaaggt cactatctgt
gcttagcagt 720gaggggcagc tctcctttga ggtatccatc ccacgcccca
tactattaat cttgtactga 780ctcaaatgac cttacttggt aaagacccgc
attttgaatt agtcagcaca atgatctgaa 840gcatccatag ccaaactcaa
acaggctttg gaggacatga taagggctgg agcagaacaa 900caggaagctt
gattgcttga accttgttca tagccagccc tgtagtgtac ttgtttgcat
960actcataata ctgcattcct attggacaga tactatcgct taacgattgg
tagataacaa 1020cagttctaat tggacgccta agcagtggga gttttaaata
aatgccattg gttgcgagcc 1080gcgagcagcc gctataaaag ggactgccgc
ggctcgactt tagttgaagt tactgacagt 1140taataaagag ctgaattcaa
ctccggtctc gagtctgctt tgttctggcg atagaacaag 1200aacaagaact
gaaagagaac aaaccgttct tttccatcga taggcaccat ggctcctcaa
1260ctactcctct gtctgatcct cacttttcta tggagtctc
1299222231DNAPseudonaja textilis textilis 22acgcggggga agttactgac
agttaataaa gagctgaatt aactccggtc tcgagtctgc 60tttgttctgg cgatagaaca
agaacaagaa ctgaaagaga acaaactgtt cttttccatc 120gataggcacc
atggctcctc aactactcct ctgtctgatc ctcacttttc tgtggagtct
180cccagaggct gaaagtaatg tattcttaaa aagcaaagtg gcaaatagat
ttttgcaaag 240aacaaaacga gctaattcac tggttgagga atttaaatct
ggaaacattg aaagggaatg 300cattgaggag agatgttcaa aagaagaagc
cagggaggta tttgaagatg acgagaaaac 360tgagaccttc tggaatgttt
atgtagatgg ggatcagtgt tcatcaaacc cctgtcatta 420tcgcgggata
tgcaaagatg gcattggtag ctatacctgt acctgcttgt ctggctatga
480agggaaaaac tgtgaacgtg tcttatataa gtcctgcaga gtggacaatg
gtaactgttg 540gcacttctgc aaatctgttc aaaacgatat tcaatgttca
tgcgctgaag gttacctttt 600gggagaggat gggcactctt gtgttgctgg
aggtaacttt tcatgtggta gaaatatcaa 660aacaaggaac aagagggaag
caagtctgcc tgactttgtg cagtcccata atgcaacttt 720gctgaaaaaa
tctgataatc caagccctga tatcagaatt gttaacggaa tggactgcaa
780actgggtgaa tgtccgtggc aggcagctct ggtagatgac aagaaaggtg
tgttttgtgg 840aggaacaatt ttgagtccca tctatgtgct tactgcagcc
cactgcatta atgagaccga 900gacgatttca gttgttgtag gagaaataga
cagatcaaga gcagaaaccg gacctcttct 960ttctgtggat aaagtatatg
tgcataaaaa atttgttcct cccaaaaaaa gccaggaatt 1020ctatgaaaag
tttgatcttg tcagctatga ctatgatata gccatcatcc aaatgaagac
1080ccctatccag ttctctgaaa atgtggttcc tgcctgcctt cccacagctg
attttgccaa 1140ccaagtcctc atgaaacaag attttggcat cgttagtgga
tttgggggta ttttcgaaag 1200aggaccgaac tctaaaacac ttaaagtcct
taaggttcct
tatgtggaca ggcacacctg 1260catgctttcc agcaattttc caattactcc
aactatgttc tgtgctggct atgatactct 1320gcctcaagat gcatgccaag
gagacagcgg ggggccccac atcactgcat acagagatac 1380ccactttatt
actgggattg tcagctgggg ggaaggatgt gcacggaaag gcagatatgg
1440tatttacaca aaattgtcca aattcatccc ttggataaaa agaataatgc
gtcaaaagct 1500acccagtaca gagtcaagca ctggtcggct ctaaaaatca
tccagtgaca tatttcatgc 1560agctataatg cattgggtta gaacattcat
gatatccact ttggttcaga actcttcaga 1620tgtagggcca tttttaaata
taacattcaa gtcatgtagc tttcctattt atcgagacct 1680tttttcttct
ggtattaatc ccttctggaa catagaatga gtaggcgatt tcatttcagc
1740tcttgtctct cgtgtcctat cttttatgac cttttctaaa gatttataaa
ggtttataat 1800ttataatcct tcaaatagaa gctcagcagg aatatttggt
ccctttgtaa tgcaacctcc 1860agttcccttg agaccatcag ttgggttaat
caaggtagtg cccaattcag ctgaattgtt 1920gtccaattta atttacctca
aaccaagcct tcagtactgt tgccttctac ttctatggag 1980ggggagttag
ggacgtcata aaaccttgct ctccgaatcc aacacttcat gtcaaaaatt
2040tcttgaagaa agtgtacaga attctgtatt tcccaaatgg ttattccact
cgcgtgctca 2100cattttgggt tattttgtgt gatcaaaatt tccagtgaca
ggatctgatt gagatgatca 2160ctaactgggt tataggaccc gaataaaagt
gatatattct aaaaaaaaaa aaaaaaaaaa 2220aaaaaaaaaa a
2231232219DNAPseudonaja textilis textilis 23acgcggggga agttactgac
agttaataaa gagctgaatt aactccggtc tcgagtctgc 60tttgttctgg cgatagaaca
agaacaagaa ctgaaagaga acaaactgtt cttttccatc 120gataggcacc
atggctcctc aactactcct ctgtctgatc ctcacttttc tgtggagtct
180cccagaggct gaaagtaatg tattcttaaa aagcaaagtg gcaaatagat
ttttgcaaag 240aacagaacga gctaattcac tggttgagga atttaaatct
ggaaacattg aaagggaatg 300cattgaggag agatgttcaa aagaagaagc
cagggaggta tttgaagatg acgagaaaac 360tgagaccttc tggaatgttt
atgtagatgg ggatcagtgt tcatcaaacc cctgtcatta 420tcgcgggata
tgcaaagatg gcattggtag ctatacctgt acctgcttgt ctggctatga
480agggaaaaac tgtgaacgtg tcttatataa gtcctgcaga gtggacaatg
gtaactgttg 540gcacttctgc aaacatgttc aaaatgatat tcagtgttca
tgtgctgaag gttacctttt 600gggagaggat gggcactctt gtgttgctgg
aggtaacttt tcatgtggta gaaatatcaa 660aacaaggaac aagagggaag
caaatctgcc tgactttgtg cagtcccaga atgcaacttt 720gctgaaaaaa
tctgataatc caagccctga tatcagaatt gttaatggaa tggattgcaa
780actgggtgaa tgtccgtggc aggcagctct ggtagatgaa aaggaaggtg
tgttttgtgg 840aggaacaatt ttgagtccca tctatgtgct tactgcagcc
cactgcatta atgagaccga 900gacgatttca gttgttgtag gggaaataga
caaatcaaga atagaaaccg gacctcttct 960ttctgtggat aaaatatatg
tgcataaaaa atttgttcct cctcaaaaag cctataagtt 1020tgatcttgcc
gcctatgact atgacatagc catcatccaa atgaagaccc ctatccagtt
1080ctctgaaaat gtggttcctg cctgccttcc cacagctgat tttgccaacc
aagtcctcat 1140gaaacaagat tttggcatcg ttagtggatt tgggcgtatt
gtcgaaaaag gaccaaaatc 1200taaaacactt aaagtcctta aggttcctta
tgtggacagg cacacctgca tggtttccag 1260cgaaactcca attactccaa
atatgttctg tgctggctat gatactctgc ctcgagatgc 1320atgccaggga
gacagtgggg ggccccacac cactgtatac agagataccc actttattac
1380tgggattgtc agctcggggg aaggatgtgc aaggaatggc aaatatggta
attacacaaa 1440actgtccaaa ttcatccctt ggataaaaag aataatgcgt
caaaagctac ccagtacaga 1500gtcaagcact ggtcggctct aaaaatcatc
cagtgacata tttcatgcag ctataatgca 1560ttgggttaga acattcatga
tatccacttt ggttcagaac tcttcagatg tagggccatt 1620tttaaatata
acattcaagt catgtagctt tcctatttat cgagaccttt tttcttctgg
1680tattaatccc ttctggaaca tagaatgagt aggcgatttc atttcagctc
ttgtctctcg 1740tgtcctatct tttatgacct tttctaaaga tttataaagg
tttataattt ataatccttc 1800aaatagaagc tcagcaggaa tatttggtcc
ctttgtaatg caacctccag ttcccttgag 1860accatcagtt gggttaatca
aggtagtgcc caattcagct gaattgttgt ccaatttaat 1920ttacctcaaa
ccaagccttc agtactgttg ccttctactt ctatggaggg ggagttaggg
1980acgtcataaa accttgctct ccgaatccaa cacttcatgt caaaaatttc
ttgaagaaag 2040tgtacagaat tctgtatttc ccaaatggtt attccactcg
cgtgctcaca ttttgggtta 2100ttttgtgtga tcaaaatttc cagtgacagg
atctgattga gatgatcact aactgggtta 2160taggacccga ataaaagtga
tatattctaa aaaaaaaaaa aaaaaaaaaa aaaaaaaaa 2219241392DNAPseudonaja
textilis 24atggctcctc agctactcct ctgtctgatc ctcacttttc tgtggagtct
cccagaggct 60gaaagtaatg tattcttaaa aagcaaagtg gcaaatagat ttttgcaaag
aacaaaacga 120gctaattcac tggttgagga atttaaatct ggaaacattg
aaagggaatg cattgaggag 180agatgttcaa aagaagaagc cagggaggca
tttgaagatg acgagaaaac tgagaccttc 240tggaatgttt atgtagatgg
ggatcagtgt tcatcaaacc cctgtcatta tcgcgggata 300tgcaaagatg
gcattggtag ctatacctgt acctgcttgt ctggctatga agggaaaaac
360tgtgaacgtg tcttatataa gtcctgcaga gtggacaatg gtaactgttg
gcacttctgc 420aaacatgttc aaaatgatat tcagtgttca tgtgctgaag
gttacctttt gggagaggat 480gggcactctt gtgttgctgg aggtaacttt
tcatgtggta gaaatatcaa aacaaggaac 540aagagggaag caaatctgcc
tgactttgtg cagtcccaga atgcaacttt gctgaaaaaa 600tctgataatc
caagccctga tatcagaatt gttaatggaa tggattgcaa actgggtgaa
660tgtccgtggc aggcagctct ggtagatgaa aaggaaggtg tgttttgtgg
aggaacaatt 720ttgagtccca tctatgtgct tactgcagcc cactgcatta
atgagaccga gacgatttca 780gttgttgtag gggaaataga caaatcaaga
atagaaaccg gacctcttct ttctgtggat 840aaaatatatg tgcataaaaa
atttgttcct cctcaaaaag cctataagtt tgatcttgcc 900gcctatgact
atgacatagc catcatccaa atgaagaccc ctatccagtt ctctgaaaat
960gtggttcctg cctgccttcc cacagctgat tttgccaacc aagtcctcat
gaaacaagat 1020tttggcatcg ttagtggatt tgggcgtatt ttcgaaaaag
gaccaaaatc taaaacactt 1080aaagtcctta aggttcctta tgtggacagg
cacacctgca tggtttccag cgaaactcca 1140attactccaa atatgttctg
tgctggctat gatactctgc ctcgagatgc atgccaggga 1200gacagtgggg
ggccccacac cactgtatac agagataccc actttattac tgggattgtc
1260agctcggggg aaggatgtgc aaggaatggc aaatatggta tttacacaaa
actgtccaaa 1320ttcatccctt ggataaaaag aataatgcgt caaaagctac
ccagtacaga gtcaagcacc 1380ggtcggctct aa 1392251452DNAPseudonaja
textilis 25atggctcctc agctactcct ctgtctgatc ctcacttttc tgtggagtct
cccagaggct 60gaaagtaatg tattcttaaa aagcaaagtg gcaaatagat ttttgcaaag
aacaaaacga 120gccaattcac tgtttgagga atttaaatct ggaaacattg
aaagggaatg cattgaggag 180agatgttcaa aagaagaagc cagggaggca
tttgaagatg acgagaaaac tgagaccttc 240tggaatgttt atgtagatgg
ggatcagtgt tcatcaaacc cctgtcatta tggcgggaca 300tgcaaagatg
gcattggtag ctatacctgt acctgcttgt ctggctatga agggaaaaac
360tgtgaatatg tcttatataa gtcctgcaga gtggacaatg gtgactgttg
gcacttctgc 420aaacctgttc aaaacggaat tcagtgttca tgtgctgaaa
gttacctttt gggagaggat 480gggcactctt gtgttgctgg aggtgacttt
tcatgtggta gaaatataaa aacaaggaac 540aagcgggaag caaatctgcc
tgactttcaa acagattttt ctgatgacta cgatgagatt 600gatgaaaata
attttgttga aactcctaca aatttctctg gcttagttct cactgtgcag
660tcccagaatg caactttgct gaaaaaatct gataatccaa gccctgatat
cagagttgtt 720aatggaacag actgcaaact aggtgaatgt ccatggcagg
cacttctgct aaatgatgaa 780ggagatgggt tttgtggagg aacaattttg
agtcccatct atgtgcttac tgcagcccac 840tgcattaacc agaccaagta
cattacagtt gttgtagggg aaatagacat atcaagcaaa 900aaaaccggac
gtcttcattc tgtggataaa atatatgtgc atcaaaaatt tgttcctgcc
960acgtatgact atgacatagc catcatccaa ctgaagaccc ctatccagtt
ctctgaaaat 1020gtggttcctg cctgccttcc cactgctgat tttgccaacc
aagtcctcat gaaacaaaat 1080tttggcatcg ttagtggatt tgggcgtact
cgagaaagag gaaagacctc taacacactt 1140aaagttgtta cgcttcctta
tgtggacagg cacacctgca tgctttccag caattttcca 1200attactcaaa
atatgttctg tgctggctat gatactctgc ctcaagatgc atgccaggga
1260gacagcggag ggccccacat cactgcatac agagataccc actttattac
tgggattgtc 1320agctgggggg aaggatgtgc acagacaggc aaatatggtg
tttacacaaa agtgtccaaa 1380ttcatccttt ggataaaaag aataatacgt
caaaagcaac ccagtacaga gtcaagcacc 1440ggtcggctct aa
145226483PRTPseudonaja textilis 26Met Ala Pro Gln Leu Leu Leu Cys
Leu Ile Leu Thr Phe Leu Trp Ser1 5 10 15Leu Pro Glu Ala Glu Ser Asn
Val Phe Leu Lys Ser Lys Val Ala Asn 20 25 30Arg Phe Leu Gln Arg Thr
Lys Arg Ala Asn Ser Leu Phe Glu Glu Phe 35 40 45Lys Ser Gly Asn Ile
Glu Arg Glu Cys Ile Glu Glu Arg Cys Ser Lys 50 55 60Glu Glu Ala Arg
Glu Ala Phe Glu Asp Asp Glu Lys Thr Glu Thr Phe65 70 75 80Trp Asn
Val Tyr Val Asp Gly Asp Gln Cys Ser Ser Asn Pro Cys His 85 90 95Tyr
Gly Gly Thr Cys Lys Asp Gly Ile Gly Ser Tyr Thr Cys Thr Cys 100 105
110Leu Ser Gly Tyr Glu Gly Lys Asn Cys Glu Tyr Val Leu Tyr Lys Ser
115 120 125Cys Arg Val Asp Asn Gly Asp Cys Trp His Phe Cys Lys Pro
Val Gln 130 135 140Asn Gly Ile Gln Cys Ser Cys Ala Glu Ser Tyr Leu
Leu Gly Glu Asp145 150 155 160Gly His Ser Cys Val Ala Gly Gly Asp
Phe Ser Cys Gly Arg Asn Ile 165 170 175Lys Thr Arg Asn Lys Arg Glu
Ala Asn Leu Pro Asp Phe Gln Thr Asp 180 185 190Phe Ser Asp Asp Tyr
Asp Glu Ile Asp Glu Asn Asn Phe Val Glu Thr 195 200 205Pro Thr Asn
Phe Ser Gly Leu Val Leu Thr Val Gln Ser Gln Asn Ala 210 215 220Thr
Leu Leu Lys Lys Ser Asp Asn Pro Ser Pro Asp Ile Arg Val Val225 230
235 240Asn Gly Thr Asp Cys Lys Leu Gly Glu Cys Pro Trp Gln Ala Leu
Leu 245 250 255Leu Asn Asp Glu Gly Asp Gly Phe Cys Gly Gly Thr Ile
Leu Ser Pro 260 265 270Ile Tyr Val Leu Thr Ala Ala His Cys Ile Asn
Gln Thr Lys Tyr Ile 275 280 285Thr Val Val Val Gly Glu Ile Asp Ile
Ser Ser Lys Lys Thr Gly Arg 290 295 300Leu His Ser Val Asp Lys Ile
Tyr Val His Gln Lys Phe Val Pro Ala305 310 315 320Thr Tyr Asp Tyr
Asp Ile Ala Ile Ile Gln Leu Lys Thr Pro Ile Gln 325 330 335Phe Ser
Glu Asn Val Val Pro Ala Cys Leu Pro Thr Ala Asp Phe Ala 340 345
350Asn Gln Val Leu Met Lys Gln Asn Phe Gly Ile Val Ser Gly Phe Gly
355 360 365Arg Thr Arg Glu Arg Gly Lys Thr Ser Asn Thr Leu Lys Val
Val Thr 370 375 380Leu Pro Tyr Val Asp Arg His Thr Cys Met Leu Ser
Ser Asn Phe Pro385 390 395 400Ile Thr Gln Asn Met Phe Cys Ala Gly
Tyr Asp Thr Leu Pro Gln Asp 405 410 415Ala Cys Gln Gly Asp Ser Gly
Gly Pro His Ile Thr Ala Tyr Arg Asp 420 425 430Thr His Phe Ile Thr
Gly Ile Val Ser Trp Gly Glu Gly Cys Ala Gln 435 440 445Thr Gly Lys
Tyr Gly Val Tyr Thr Lys Val Ser Lys Phe Ile Leu Trp 450 455 460Ile
Lys Arg Ile Ile Arg Gln Lys Gln Pro Ser Thr Glu Ser Ser Thr465 470
475 480Gly Arg Leu27463PRTPseudonaja textilis 27Met Ala Pro Gln Leu
Leu Leu Cys Leu Ile Leu Thr Phe Leu Trp Ser1 5 10 15Leu Pro Glu Ala
Glu Ser Asn Val Phe Leu Lys Ser Lys Val Ala Asn 20 25 30Arg Phe Leu
Gln Arg Thr Lys Arg Ala Asn Ser Leu Val Glu Glu Phe 35 40 45Lys Ser
Gly Asn Ile Glu Arg Glu Cys Ile Glu Glu Arg Cys Ser Lys 50 55 60Glu
Glu Ala Arg Glu Ala Phe Glu Asp Asp Glu Lys Thr Glu Thr Phe65 70 75
80Trp Asn Val Tyr Val Asp Gly Asp Gln Cys Ser Ser Asn Pro Cys His
85 90 95Tyr Arg Gly Ile Cys Lys Asp Gly Ile Gly Ser Tyr Thr Cys Thr
Cys 100 105 110Leu Ser Gly Tyr Glu Gly Lys Asn Cys Glu Arg Val Leu
Tyr Lys Ser 115 120 125Cys Arg Val Asp Asn Gly Asn Cys Trp His Phe
Cys Lys His Val Gln 130 135 140Asn Asp Ile Gln Cys Ser Cys Ala Glu
Gly Tyr Leu Leu Gly Glu Asp145 150 155 160Gly His Ser Cys Val Ala
Gly Gly Asn Phe Ser Cys Gly Arg Asn Ile 165 170 175Lys Thr Arg Asn
Lys Arg Glu Ala Asn Leu Pro Asp Phe Val Gln Ser 180 185 190Gln Asn
Ala Thr Leu Leu Lys Lys Ser Asp Asn Pro Ser Pro Asp Ile 195 200
205Arg Ile Val Asn Gly Met Asp Cys Lys Leu Gly Glu Cys Pro Trp Gln
210 215 220Ala Ala Leu Val Asp Glu Lys Glu Gly Val Phe Cys Gly Gly
Thr Ile225 230 235 240Leu Ser Pro Ile Tyr Val Leu Thr Ala Ala His
Cys Ile Asn Glu Thr 245 250 255Glu Thr Ile Ser Val Val Val Gly Glu
Ile Asp Lys Ser Arg Ile Glu 260 265 270Thr Gly Pro Leu Leu Ser Val
Asp Lys Ile Tyr Val His Lys Lys Phe 275 280 285Val Pro Pro Gln Lys
Ala Tyr Lys Phe Asp Leu Ala Ala Tyr Asp Tyr 290 295 300Asp Ile Ala
Ile Ile Gln Met Lys Thr Pro Ile Gln Phe Ser Glu Asn305 310 315
320Val Val Pro Ala Cys Leu Pro Thr Ala Asp Phe Ala Asn Gln Val Leu
325 330 335Met Lys Gln Asp Phe Gly Ile Val Ser Gly Phe Gly Arg Ile
Phe Glu 340 345 350Lys Gly Pro Lys Ser Lys Thr Leu Lys Val Leu Lys
Val Pro Tyr Val 355 360 365Asp Arg His Thr Cys Met Val Ser Ser Glu
Thr Pro Ile Thr Pro Asn 370 375 380Met Phe Cys Ala Gly Tyr Asp Thr
Leu Pro Arg Asp Ala Cys Gln Gly385 390 395 400Asp Ser Gly Gly Pro
His Thr Thr Val Tyr Arg Asp Thr His Phe Ile 405 410 415Thr Gly Ile
Val Ser Ser Gly Glu Gly Cys Ala Arg Asn Gly Lys Tyr 420 425 430Gly
Ile Tyr Thr Lys Leu Ser Lys Phe Ile Pro Trp Ile Lys Arg Ile 435 440
445Met Arg Gln Lys Leu Pro Ser Thr Glu Ser Ser Thr Gly Arg Leu 450
455 46028467PRTPseudonaja textilis textilis 28Met Ala Pro Gln Leu
Leu Leu Cys Leu Ile Leu Thr Phe Leu Trp Ser1 5 10 15Leu Pro Glu Ala
Glu Ser Asn Val Phe Leu Lys Ser Lys Val Ala Asn 20 25 30Arg Phe Leu
Gln Arg Thr Lys Arg Ala Asn Ser Leu Val Glu Glu Phe 35 40 45Lys Ser
Gly Asn Ile Glu Arg Glu Cys Ile Glu Glu Arg Cys Ser Lys 50 55 60Glu
Glu Ala Arg Glu Val Phe Glu Asp Asp Glu Lys Thr Glu Thr Phe65 70 75
80Trp Asn Val Tyr Val Asp Gly Asp Gln Cys Ser Ser Asn Pro Cys His
85 90 95Tyr Arg Gly Ile Cys Lys Asp Gly Ile Gly Ser Tyr Thr Cys Thr
Cys 100 105 110Leu Ser Gly Tyr Glu Gly Lys Asn Cys Glu Arg Val Leu
Tyr Lys Ser 115 120 125Cys Arg Val Asp Asn Gly Asn Cys Trp His Phe
Cys Lys Ser Val Gln 130 135 140Asn Asp Ile Gln Cys Ser Cys Ala Glu
Gly Tyr Leu Leu Gly Glu Asp145 150 155 160Gly His Ser Cys Val Ala
Gly Gly Asn Phe Ser Cys Gly Arg Asn Ile 165 170 175Lys Thr Arg Asn
Lys Arg Glu Ala Ser Leu Pro Asp Phe Val Gln Ser 180 185 190His Asn
Ala Thr Leu Leu Lys Lys Ser Asp Asn Pro Ser Pro Asp Ile 195 200
205Arg Ile Val Asn Gly Met Asp Cys Lys Leu Gly Glu Cys Pro Trp Gln
210 215 220Ala Ala Leu Val Asp Asp Lys Lys Gly Val Phe Cys Gly Gly
Thr Ile225 230 235 240Leu Ser Pro Ile Tyr Val Leu Thr Ala Ala His
Cys Ile Asn Glu Thr 245 250 255Glu Thr Ile Ser Val Val Val Gly Glu
Ile Asp Arg Ser Arg Ala Glu 260 265 270Thr Gly Pro Leu Leu Ser Val
Asp Lys Val Tyr Val His Lys Lys Phe 275 280 285Val Pro Pro Lys Lys
Ser Gln Glu Phe Tyr Glu Lys Phe Asp Leu Val 290 295 300Ser Tyr Asp
Tyr Asp Ile Ala Ile Ile Gln Met Lys Thr Pro Ile Gln305 310 315
320Phe Ser Glu Asn Val Val Pro Ala Cys Leu Pro Thr Ala Asp Phe Ala
325 330 335Asn Gln Val Leu Met Lys Gln Asp Phe Gly Ile Val Ser Gly
Phe Gly 340 345 350Gly Ile Phe Glu Arg Gly Pro Asn Ser Lys Thr Leu
Lys Val Leu Lys 355 360 365Val Pro Tyr Val Asp Arg His Thr Cys Met
Leu Ser Ser Asn Phe Pro 370 375 380Ile Thr Pro Thr Met Phe Cys Ala
Gly Tyr Asp Thr Leu Pro Gln Asp385 390 395 400Ala Cys Gln Gly Asp
Ser Gly Gly Pro His Ile Thr Ala Tyr Arg Asp 405 410 415Thr His Phe
Ile Thr Gly Ile Val Ser Trp Gly Glu Gly Cys Ala Arg 420 425 430Lys
Gly Arg Tyr Gly Ile Tyr Thr Lys Leu Ser Lys Phe Ile Pro Trp 435 440
445Ile Lys Arg Ile Met Arg Gln Lys Leu Pro Ser Thr Glu Ser Ser Thr
450 455 460Gly Arg Leu46529463PRTPseudonaja textilis textilis 29Met
Ala Pro Gln Leu Leu Leu Cys Leu Ile Leu Thr Phe Leu Trp Ser1 5
10
15Leu Pro Glu Ala Glu Ser Asn Val Phe Leu Lys Ser Lys Val Ala Asn
20 25 30Arg Phe Leu Gln Arg Thr Glu Arg Ala Asn Ser Leu Val Glu Glu
Phe 35 40 45Lys Ser Gly Asn Ile Glu Arg Glu Cys Ile Glu Glu Arg Cys
Ser Lys 50 55 60Glu Glu Ala Arg Glu Val Phe Glu Asp Asp Glu Lys Thr
Glu Thr Phe65 70 75 80Trp Asn Val Tyr Val Asp Gly Asp Gln Cys Ser
Ser Asn Pro Cys His 85 90 95Tyr Arg Gly Ile Cys Lys Asp Gly Ile Gly
Ser Tyr Thr Cys Thr Cys 100 105 110Leu Ser Gly Tyr Glu Gly Lys Asn
Cys Glu Arg Val Leu Tyr Lys Ser 115 120 125Cys Arg Val Asp Asn Gly
Asn Cys Trp His Phe Cys Lys His Val Gln 130 135 140Asn Asp Ile Gln
Cys Ser Cys Ala Glu Gly Tyr Leu Leu Gly Glu Asp145 150 155 160Gly
His Ser Cys Val Ala Gly Gly Asn Phe Ser Cys Gly Arg Asn Ile 165 170
175Lys Thr Arg Asn Lys Arg Glu Ala Asn Leu Pro Asp Phe Val Gln Ser
180 185 190Gln Asn Ala Thr Leu Leu Lys Lys Ser Asp Asn Pro Ser Pro
Asp Ile 195 200 205Arg Ile Val Asn Gly Met Asp Cys Lys Leu Gly Glu
Cys Pro Trp Gln 210 215 220Ala Ala Leu Val Asp Glu Lys Glu Gly Val
Phe Cys Gly Gly Thr Ile225 230 235 240Leu Ser Pro Ile Tyr Val Leu
Thr Ala Ala His Cys Ile Asn Glu Thr 245 250 255Glu Thr Ile Ser Val
Val Val Gly Glu Ile Asp Lys Ser Arg Ile Glu 260 265 270Thr Gly Pro
Leu Leu Ser Val Asp Lys Ile Tyr Val His Lys Lys Phe 275 280 285Val
Pro Pro Gln Lys Ala Tyr Lys Phe Asp Leu Ala Ala Tyr Asp Tyr 290 295
300Asp Ile Ala Ile Ile Gln Met Lys Thr Pro Ile Gln Phe Ser Glu
Asn305 310 315 320Val Val Pro Ala Cys Leu Pro Thr Ala Asp Phe Ala
Asn Gln Val Leu 325 330 335Met Lys Gln Asp Phe Gly Ile Val Ser Gly
Phe Gly Arg Ile Val Glu 340 345 350Lys Gly Pro Lys Ser Lys Thr Leu
Lys Val Leu Lys Val Pro Tyr Val 355 360 365Asp Arg His Thr Cys Met
Val Ser Ser Glu Thr Pro Ile Thr Pro Asn 370 375 380Met Phe Cys Ala
Gly Tyr Asp Thr Leu Pro Arg Asp Ala Cys Gln Gly385 390 395 400Asp
Ser Gly Gly Pro His Thr Thr Val Tyr Arg Asp Thr His Phe Ile 405 410
415Thr Gly Ile Val Ser Ser Gly Glu Gly Cys Ala Arg Asn Gly Lys Tyr
420 425 430Gly Asn Tyr Thr Lys Leu Ser Lys Phe Ile Pro Trp Ile Lys
Arg Ile 435 440 445Met Arg Gln Lys Leu Pro Ser Thr Glu Ser Ser Thr
Gly Arg Leu 450 455 46030449PRTPseudonaja textilis 30Met Ala Pro
Gln Leu Leu Leu Cys Leu Ile Leu Thr Phe Leu Trp Ser1 5 10 15Leu Pro
Glu Ala Glu Ser Asn Val Phe Leu Lys Ser Lys Val Ala Asn 20 25 30Arg
Phe Leu Gln Arg Thr Lys Arg Ala Asn Ser Leu Val Glu Glu Phe 35 40
45Lys Ser Gly Asn Ile Glu Arg Glu Cys Ile Glu Glu Arg Cys Ser Lys
50 55 60Glu Glu Ala Arg Glu Ala Phe Glu Asp Asp Glu Lys Thr Glu Thr
Phe65 70 75 80Trp Asn Val Tyr Val Asp Gly Asp Gln Cys Ser Ser Asn
Pro Cys His 85 90 95Tyr Arg Gly Ile Cys Lys Asp Gly Ile Gly Ser Tyr
Thr Cys Thr Cys 100 105 110Leu Ser Gly Tyr Glu Gly Lys Asn Cys Glu
Arg Val Leu Tyr Lys Ser 115 120 125Cys Arg Val Asp Asn Gly Asn Cys
Trp His Phe Cys Lys Ser Val Gln 130 135 140Asn Asp Ile Gln Cys Ser
Cys Ala Glu Gly Tyr Leu Leu Gly Glu Asp145 150 155 160Gly His Ser
Cys Val Ala Gly Gly Asn Phe Ser Cys Gly Arg Asn Ile 165 170 175Lys
Thr Arg Asn Lys Arg Glu Ala Ser Leu Pro Asp Phe Val Gln Ser 180 185
190Gln Asn Ala Pro Leu Leu Lys Ile Ser Asp Asn Pro Ser Pro Asp Ile
195 200 205Arg Ile Val Asn Gly Met Asp Cys Lys Leu Gly Glu Cys Pro
Trp Gln 210 215 220Ala Ala Leu Val Asp Asp Lys Lys Gly Val Phe Cys
Gly Gly Thr Ile225 230 235 240Leu Ser Pro Ile Tyr Val Leu Thr Ala
Ala His Cys Ile Asn Glu Thr 245 250 255Glu Thr Ile Ser Val Val Val
Gly Glu Ile Asp Arg Ser Arg Ala Glu 260 265 270Thr Gly Pro Leu Leu
Ser Val Asp Lys Val Tyr Val His Lys Lys Phe 275 280 285Val Pro Pro
Lys Lys Ser Gln Glu Phe Tyr Glu Lys Phe Asp Leu Val 290 295 300Ser
Tyr Asp Tyr Asp Ile Ala Ile Ile Gln Met Lys Thr Pro Ile Gln305 310
315 320Phe Ser Glu Asn Val Val Pro Ala Cys Leu Pro Thr Ala Asp Phe
Ala 325 330 335Asn Gln Val Leu Met Lys Gln Asp Phe Gly Ile Val Ser
Gly Phe Gly 340 345 350Gly Ile Phe Glu Arg Gly Pro Asn Ser Lys Thr
Leu Lys Val Leu Lys 355 360 365Val Pro Tyr Val Asp Arg His Thr Cys
Met Leu Ser Ser Asn Phe Pro 370 375 380Ile Thr Pro Thr Met Phe Cys
Ala Gly Tyr Asp Thr Leu Pro Gln Asp385 390 395 400Ala Cys Gln Gly
Asp Ser Gly Gly Pro His Ile Thr Ala Tyr Arg Asp 405 410 415Thr His
Phe Ile Thr Gly Ile Val Ser Trp Gly Glu Gly Cys Ala Arg 420 425
430Lys Gly Arg Tyr Gly Ile Tyr Thr Lys Leu Ser Lys Phe Ile Pro Trp
435 440 445Ile311404DNAOxyuranus scutellatus 31atggctcctc
aactactcct ctgtctgatc ctcacttttc tgtggagtct cccagaggcc 60gaaagtaatg
tattcttaaa aagcaaagtg gcaaatagat ttttgcaaag aacaaaacga
120gctaattcac tgtatgagga atttagatct ggaaacattg aaagggaatg
cattgaggag 180agatgttcaa aagaagaagc cagggaggta tttgaagatg
acgagaaaac tgagaccttc 240tggaatgttt atgtagatgg ggatcagtgt
tcatcaaacc cctgtcatta tcgcgggaca 300tgcaaagatg gcattggtag
ctatacctgt acctgcttgt ctggctatga agggaaaaac 360tgtgaacgtg
tcttatataa gtcctgcaga gtggacaatg gtaactgttg gcacttctgc
420aaacctgttc aaaacgatat tcagtgttca tgtgctgaag gttacctttt
gggagaggat 480gggcactctt gtgttgctgg aggtaacttt tcatgtggta
gaaatatcaa aacaaggaac 540aagagggaag caagtctgcc tgactttgtg
cagtcccaga atgcaatttt gctgaaaaaa 600tctgataatc caagccctga
tatcagaatt gttaatggaa tggactgcaa actgggtgaa 660tgtccgtggc
aggcagttct ggtagatgaa aaggaagatg cgttttgtgg aggaacaatt
720ttgagtccca tctatgtgct tactgcagcc cactgcatta accagaccaa
gatgatttca 780gttgttgtag gggaaataaa catatcaaga aaaaaccccg
gacgtcttct ttctgtggat 840aaaatatatg tgcatcaaaa atttgttcct
cccaaaaaag gctatgaatt ctatgaaaag 900tttgatcttg tcagctatga
ctatgatata gccatcctcc aaatgaagac ccctatccag 960ttctctgaaa
atgtggttcc tgcctgcctt cccacagctg attttgccaa ccaagtcctc
1020atgaaacaag attttggcat cgttagtgga tttgggcgta ttttcgaaaa
aggacctcaa 1080tctaaaacac ttaaagtcct taaggttcct tatgtggaca
ggcacacctg catgctttcc 1140agcgaatctc caattactcc aactatgttc
tgtgctggct atgatactct gcctcgagat 1200gcatgccagg gagacagtgg
ggggccccac atcactgcat acagagatac ccactttatt 1260actgggattg
tcagctgggg ggaaggatgt gcacagacag gcaaatatgg tgtttacaca
1320aaagtgtcca aattcatcct ttggataaaa agaataatgc gtcaaaagct
acccagtaca 1380gagtcaagca ctggtcggct ctaa 140432467PRTOxyuranus
scutellatus 32Met Ala Pro Gln Leu Leu Leu Cys Leu Ile Leu Thr Phe
Leu Trp Ser1 5 10 15Leu Pro Glu Ala Glu Ser Asn Val Phe Leu Lys Ser
Lys Val Ala Asn 20 25 30Arg Phe Leu Gln Arg Thr Lys Arg Ala Asn Ser
Leu Tyr Glu Glu Phe 35 40 45Arg Ser Gly Asn Ile Glu Arg Glu Cys Ile
Glu Glu Arg Cys Ser Lys 50 55 60Glu Glu Ala Arg Glu Val Phe Glu Asp
Asp Glu Lys Thr Glu Thr Phe65 70 75 80Trp Asn Val Tyr Val Asp Gly
Asp Gln Cys Ser Ser Asn Pro Cys His 85 90 95Tyr Arg Gly Thr Cys Lys
Asp Gly Ile Gly Ser Tyr Thr Cys Thr Cys 100 105 110Leu Ser Gly Tyr
Glu Gly Lys Asn Cys Glu Arg Val Leu Tyr Lys Ser 115 120 125Cys Arg
Val Asp Asn Gly Asn Cys Trp His Phe Cys Lys Pro Val Gln 130 135
140Asn Asp Ile Gln Cys Ser Cys Ala Glu Gly Tyr Leu Leu Gly Glu
Asp145 150 155 160Gly His Ser Cys Val Ala Gly Gly Asn Phe Ser Cys
Gly Arg Asn Ile 165 170 175Lys Thr Arg Asn Lys Arg Glu Ala Ser Leu
Pro Asp Phe Val Gln Ser 180 185 190Gln Asn Ala Ile Leu Leu Lys Lys
Ser Asp Asn Pro Ser Pro Asp Ile 195 200 205Arg Ile Val Asn Gly Met
Asp Cys Lys Leu Gly Glu Cys Pro Trp Gln 210 215 220Ala Val Leu Val
Asp Glu Lys Glu Asp Ala Phe Cys Gly Gly Thr Ile225 230 235 240Leu
Ser Pro Ile Tyr Val Leu Thr Ala Ala His Cys Ile Asn Gln Thr 245 250
255Lys Met Ile Ser Val Val Val Gly Glu Ile Asn Ile Ser Arg Lys Asn
260 265 270Pro Gly Arg Leu Leu Ser Val Asp Lys Ile Tyr Val His Gln
Lys Phe 275 280 285Val Pro Pro Lys Lys Gly Tyr Glu Phe Tyr Glu Lys
Phe Asp Leu Val 290 295 300Ser Tyr Asp Tyr Asp Ile Ala Ile Leu Gln
Met Lys Thr Pro Ile Gln305 310 315 320Phe Ser Glu Asn Val Val Pro
Ala Cys Leu Pro Thr Ala Asp Phe Ala 325 330 335Asn Gln Val Leu Met
Lys Gln Asp Phe Gly Ile Val Ser Gly Phe Gly 340 345 350Arg Ile Phe
Glu Lys Gly Pro Gln Ser Lys Thr Leu Lys Val Leu Lys 355 360 365Val
Pro Tyr Val Asp Arg His Thr Cys Met Leu Ser Ser Glu Ser Pro 370 375
380Ile Thr Pro Thr Met Phe Cys Ala Gly Tyr Asp Thr Leu Pro Arg
Asp385 390 395 400Ala Cys Gln Gly Asp Ser Gly Gly Pro His Ile Thr
Ala Tyr Arg Asp 405 410 415Thr His Phe Ile Thr Gly Ile Val Ser Trp
Gly Glu Gly Cys Ala Gln 420 425 430Thr Gly Lys Tyr Gly Val Tyr Thr
Lys Val Ser Lys Phe Ile Leu Trp 435 440 445Ile Lys Arg Ile Met Arg
Gln Lys Leu Pro Ser Thr Glu Ser Ser Thr 450 455 460Gly Arg
Leu465331404DNAOxyuranus microlepidotus 33atggctcctc aactactcct
ctgtctgatc ctcacttttc tgtggagtct cccagaggct 60gaaagtaatg tattcttaaa
aagcaaagtg gcaaatagat ttttgcaaag aacaaaacga 120gctaattcac
tgtttgagga atttagatct ggaaacattg aaagggaatg cattgaggag
180agatgttcaa aagaagaagc cagggaggta tttgaagatg acgagaaaac
tgagaccttc 240tggaatgttt atgtagatgg ggatcagtgt tcatcaaacc
cctgtcatta tcgcgggaca 300tgcaaagatg gcattggtag ctatacctgt
acctgcttgt ttggctatga agggaaaaac 360tgtgaacgtg tcttatataa
gtcctgcaga gtggacaatg gtaactgttg gcacttctgc 420aaacctgttc
aaaacgatat tcagtgttca tgtgctgaag gttacctttt gggagaggat
480gggcactctt gtgttgctgg aggtaacttt tcatgtggta gaaatatcaa
aacaaggaac 540aagagggaag caagtctgcc tgactttgtg cagtcccaga
atgcaacttt gctgaaaaaa 600tctgataatc caagccctga tatcagaatt
gttaatggaa tggactgcaa actgggtgaa 660tgtccgtggc aggcagttct
ggtagatgaa aaggaaggtg tgttttgtgg aggaacaatt 720ttgagtccca
tctatgtgct tactgcagcc cactgcatta accagaccga gaagatttca
780gttgttgtag gggaaataga caaatcaaga gtagaaaccg gacatcttct
ttctgtggat 840aaaatatatg tgcataaaaa atttgttcct cccaaaaaag
gctataaatt ctatgaaaag 900tttgatcttg tcagctatga ctatgatata
gccatcatcc aaatgaagac ccctatccag 960ttctctgaaa atgtggttcc
tgcctgcctt cccacagctg attttgccaa ccaagtcctc 1020atgaaacaag
attttggcat cattagtgga tttgggcgta ttttcgaaaa aggaccgaaa
1080tctaacacac ttaaagtcct taaggttcct tatgtggaca ggcacacctg
catggtttcc 1140agcgaatctc caattactcc aactatgttc tgtgctggct
atgatactct gcctcgagat 1200gcatgccagg gagacagtgg ggggccccac
atcactgcat acagagatac ccactttatt 1260actgggattg tcagctgggg
ggaaggatgt gctaagaaag gcaaatatgg tatttacaca 1320aaagtgtcca
aattcatcct ttggataaaa agaataatgc gtcaaaagct acccagtaca
1380gagtcaagca ctggtcggct ctaa 140434467PRTOxyuranus microlepidotus
34Met Ala Pro Gln Leu Leu Leu Cys Leu Ile Leu Thr Phe Leu Trp Ser1
5 10 15Leu Pro Glu Ala Glu Ser Asn Val Phe Leu Lys Ser Lys Val Ala
Asn 20 25 30Arg Phe Leu Gln Arg Thr Lys Arg Ala Asn Ser Leu Phe Glu
Glu Phe 35 40 45Arg Ser Gly Asn Ile Glu Arg Glu Cys Ile Glu Glu Arg
Cys Ser Lys 50 55 60Glu Glu Ala Arg Glu Val Phe Glu Asp Asp Glu Lys
Thr Glu Thr Phe65 70 75 80Trp Asn Val Tyr Val Asp Gly Asp Gln Cys
Ser Ser Asn Pro Cys His 85 90 95Tyr Arg Gly Thr Cys Lys Asp Gly Ile
Gly Ser Tyr Thr Cys Thr Cys 100 105 110Leu Phe Gly Tyr Glu Gly Lys
Asn Cys Glu Arg Val Leu Tyr Lys Ser 115 120 125Cys Arg Val Asp Asn
Gly Asn Cys Trp His Phe Cys Lys Pro Val Gln 130 135 140Asn Asp Ile
Gln Cys Ser Cys Ala Glu Gly Tyr Leu Leu Gly Glu Asp145 150 155
160Gly His Ser Cys Val Ala Gly Gly Asn Phe Ser Cys Gly Arg Asn Ile
165 170 175Lys Thr Arg Asn Lys Arg Glu Ala Ser Leu Pro Asp Phe Val
Gln Ser 180 185 190Gln Asn Ala Thr Leu Leu Lys Lys Ser Asp Asn Pro
Ser Pro Asp Ile 195 200 205Arg Ile Val Asn Gly Met Asp Cys Lys Leu
Gly Glu Cys Pro Trp Gln 210 215 220Ala Val Leu Val Asp Glu Lys Glu
Gly Val Phe Cys Gly Gly Thr Ile225 230 235 240Leu Ser Pro Ile Tyr
Val Leu Thr Ala Ala His Cys Ile Asn Gln Thr 245 250 255Glu Lys Ile
Ser Val Val Val Gly Glu Ile Asp Lys Ser Arg Val Glu 260 265 270Thr
Gly His Leu Leu Ser Val Asp Lys Ile Tyr Val His Lys Lys Phe 275 280
285Val Pro Pro Lys Lys Gly Tyr Lys Phe Tyr Glu Lys Phe Asp Leu Val
290 295 300Ser Tyr Asp Tyr Asp Ile Ala Ile Ile Gln Met Lys Thr Pro
Ile Gln305 310 315 320Phe Ser Glu Asn Val Val Pro Ala Cys Leu Pro
Thr Ala Asp Phe Ala 325 330 335Asn Gln Val Leu Met Lys Gln Asp Phe
Gly Ile Ile Ser Gly Phe Gly 340 345 350Arg Ile Phe Glu Lys Gly Pro
Lys Ser Asn Thr Leu Lys Val Leu Lys 355 360 365Val Pro Tyr Val Asp
Arg His Thr Cys Met Val Ser Ser Glu Ser Pro 370 375 380Ile Thr Pro
Thr Met Phe Cys Ala Gly Tyr Asp Thr Leu Pro Arg Asp385 390 395
400Ala Cys Gln Gly Asp Ser Gly Gly Pro His Ile Thr Ala Tyr Arg Asp
405 410 415Thr His Phe Ile Thr Gly Ile Val Ser Trp Gly Glu Gly Cys
Ala Lys 420 425 430Lys Gly Lys Tyr Gly Ile Tyr Thr Lys Val Ser Lys
Phe Ile Leu Trp 435 440 445Ile Lys Arg Ile Met Arg Gln Lys Leu Pro
Ser Thr Glu Ser Ser Thr 450 455 460Gly Arg
Leu465351365DNAPseudechis porphyriacus 35atggctcctc aactactcct
ctgtctgatc ctcacttttc tctggagtct cccggaggct 60gaaagtaatg tattcttaaa
aagcaaagag gcaaatagat ttttgcaaag aacaaaacga 120tctaattcac
tgtttgagga atttagacct ggaaacattg aaagggaatg cattgaggag
180aaatgttcaa aagaagaagc cagggagata tttaaagata acgagaaaac
tgaggccttt 240tggaatgttt atgtagatgg ggatcagtgt tcatcaaacc
cctgtcatta tggtgggaca 300tgcaaagatg gcattggtag ctatacctgt
acctgcttgc ctaactatga agggaaaaac 360tgtgaacatc tcttatttaa
gtcctgcaga tttttcaatg gtaactgttg gcacttctgc 420aaacctgttc
aaaacgacac tcagtgttca tgtgctgaaa gttaccgttt gggagatgat
480gggcactctt gtgttgctga aggtgacttt tcatgtggta gaaatataaa
agcaaggaac 540aagagggaag caagtctgcc tgactttgtg cagtcccaga
atgcaacttt gctgaaaaaa 600tctgataatc caagccctga tatcagaatt
attaatggaa tggactgcaa actgggtgaa 660tgtccatggc aggcagttct
gctagataaa gaaggagatg tgttttgtgg aggaacaatt 720ttgagtccca
tctatgtgct tactgcagcc cactgcatta cccagtccaa gcacatttca
780gttgttgtag gggaaataga tatatcaaga aaagaaacca gacatcttct
ttctgtagat 840aaagcatatg tgcatacaaa atttgttctt gccacctatg
actatgatat agccatcatc 900caattgaaga
cccctatcca gttctctgaa aatgtggttc ctgcctgtct tcccactgct
960gattttgcca accaagtcct catgaaacaa gattttggca tcattagtgg
atttgggcat 1020actcgatctg gaggacagac ctctaacaca cttaaagtcg
ttacgattcc ttatgtggac 1080aggcacacct gcatgctttc cagcgatttt
cgaattactc caaatatgtt ctgtgctggt 1140tatgatactc tgcctcgaga
tgcatgccag ggagacagtg gggggcccca catcactgca 1200tacagagata
cccactttat tactgggatt atcagctggg gggaaggatg tgcaaagaaa
1260ggcaaatatg gtgtttacac aaaagtgtcc aacttcatcc cttggataaa
agcagtaatg 1320cgtaaacatc aacccagtac agagtcaagc actggtcggc tctaa
136536450PRTPseudechis porphyriacus 36Met Ala Pro Gln Leu Leu Leu
Cys Leu Ile Leu Thr Phe Leu Trp Ser1 5 10 15Leu Pro Glu Ala Glu Ser
Asn Val Phe Leu Lys Ser Lys Glu Ala Asn 20 25 30Arg Phe Leu Gln Arg
Thr Lys Arg Ser Asn Ser Leu Phe Glu Glu Phe 35 40 45Arg Pro Gly Asn
Ile Glu Arg Glu Cys Ile Glu Glu Lys Cys Ser Lys 50 55 60Glu Glu Ala
Arg Glu Ile Phe Lys Asp Asn Glu Lys Thr Glu Ala Phe65 70 75 80Trp
Asn Val Tyr Val Asp Gly Asp Gln Cys Ser Ser Asn Pro Cys His 85 90
95Tyr Gly Gly Thr Cys Lys Asp Gly Ile Gly Ser Tyr Thr Cys Thr Cys
100 105 110Leu Pro Asn Tyr Glu Gly Lys Asn Cys Glu His Leu Leu Phe
Lys Ser 115 120 125Cys Arg Phe Phe Asn Gly Asn Cys Trp His Phe Cys
Lys Pro Val Gln 130 135 140Asn Asp Thr Gln Cys Ser Cys Ala Glu Ser
Tyr Arg Leu Gly Asp Asp145 150 155 160Gly His Ser Cys Val Ala Glu
Gly Asp Phe Ser Cys Gly Arg Asn Ile 165 170 175Lys Ala Arg Asn Lys
Arg Glu Ala Ser Leu Pro Asp Phe Val Gln Ser 180 185 190Gln Asn Ala
Thr Leu Leu Lys Lys Ser Asp Asn Pro Ser Pro Asp Ile 195 200 205Arg
Ile Ile Asn Gly Met Asp Cys Lys Leu Gly Glu Cys Pro Trp Gln 210 215
220Ala Val Leu Leu Asp Lys Glu Gly Asp Val Phe Cys Gly Gly Thr
Ile225 230 235 240Leu Ser Pro Ile Tyr Val Leu Thr Ala Ala His Cys
Ile Thr Gln Ser 245 250 255Lys His Ile Ser Val Val Val Gly Glu Ile
Asp Ile Ser Arg Lys Glu 260 265 270Thr Arg His Leu Leu Ser Val Asp
Lys Ala Tyr Val His Thr Lys Phe 275 280 285Val Leu Ala Thr Tyr Asp
Tyr Asp Ile Ala Ile Ile Gln Leu Lys Thr 290 295 300Pro Ile Gln Phe
Ser Glu Asn Val Val Pro Ala Cys Leu Pro Thr Ala305 310 315 320Asp
Phe Ala Asn Gln Val Leu Met Lys Gln Asp Phe Gly Ile Ile Ser 325 330
335Gly Phe Gly His Thr Arg Ser Gly Gly Gln Thr Ser Asn Thr Leu Lys
340 345 350Val Val Thr Ile Pro Tyr Val Asp Arg His Thr Cys Met Leu
Ser Ser 355 360 365Asp Phe Arg Ile Thr Pro Asn Met Phe Cys Ala Gly
Tyr Asp Thr Leu 370 375 380Pro Arg Asp Ala Cys Gln Gly Asp Ser Gly
Gly Pro His Ile Thr Ala385 390 395 400Tyr Arg Asp Thr His Phe Ile
Thr Gly Ile Ile Ser Trp Gly Glu Gly 405 410 415Cys Ala Lys Lys Gly
Lys Tyr Gly Val Tyr Thr Lys Val Ser Asn Phe 420 425 430Ile Pro Trp
Ile Lys Ala Val Met Arg Lys His Gln Pro Ser Thr Glu 435 440 445Ser
Ser 450371368DNAHoplocephalus stephensii 37atggctcctc aactactcct
ctgtctgatc ctcacttttc tgtggagtgt cccagaggct 60gaaagtaatg tattcttaaa
aagcaaagtg gcaaatagat ttttgcaaag aacaaaacga 120tctaattcac
tgtttgagga aattagacct ggaaacattg aaagggaatg cattgaggag
180aaatgttcaa aagaagaagc cagggaggta tttgaagata acgagaaaac
tgagaccttc 240tggaatgttt atgtagatgg ggatcagtgt tcatcaaacc
cctgtcatta tcacgggaca 300tgcaaagatg gcattggtag ctatacctgt
acctgcttgc ctaactatga agggaaaaac 360tgtgaaaaag tcttatttaa
gtcctgcaga gcgttcaatg gtaactgttg gcacttctgc 420aaacgtgttc
aaagtgaaac tcagtgttca tgtgctgaaa gttaccgttt gggagttgat
480gggcactctt gtgttgctga aggtgacttt tcatgtggta gaaatataaa
agcaaggaac 540aagagggaag caagtctgcc tgactttgtg cagtcccaga
aggcaacttt gctgaaaaaa 600tctgataatc caagccctga tatcagaatt
gttaatggaa tggactccaa actgggtgaa 660tgtccatggc aggcagttct
gataaatgaa aaaggagaag tgttttgtgg aggaacaatt 720ttgagtccca
tccatgtgct tactgcagcc cactgcatta accagaccaa gagcgtttca
780gttattgtag gggaaataga catatcaaga aaagaaacca gacgtcttct
ttctgtggat 840aaaatatatg tgcatacaaa atttgttcct cccaactatt
actatgggca tcaaaacttt 900gatcgtgtcg cctatgacta tgatatagcc
atcatccgaa tgaagacccc tatccagttc 960tctgaaaatg tggttcctgc
ctgccttccc actgctgatt ttgccaacga agtcctcatg 1020aaacaagatt
ctggcatcgt tagtggattt gggcgtattc gatttaaaga accgacctct
1080aacacactta aagtcattac ggttccttat gtggacaggc acacctgcat
gctttccagt 1140gattttcgaa ttactcaaaa tatgttctgt gctggctatg
atactctgcc tcaagatgca 1200tgcgagggag acagtggggg gccccacatc
actgcatacg gagataccca ctttattact 1260gggattgtca gctgggggga
aggatgtgca cggaaaggca aatatggtgt ttacacaaaa 1320gtgtccagat
tcatcccttg gataaaaaaa ataatgagtc taaagtaa 136838455PRTHoplocephalus
stephensii 38Met Ala Pro Gln Leu Leu Leu Cys Leu Ile Leu Thr Phe
Leu Trp Ser1 5 10 15Val Pro Glu Ala Glu Ser Asn Val Phe Leu Lys Ser
Lys Val Ala Asn 20 25 30Arg Phe Leu Gln Arg Thr Lys Arg Ser Asn Ser
Leu Phe Glu Glu Ile 35 40 45Arg Pro Gly Asn Ile Glu Arg Glu Cys Ile
Glu Glu Lys Cys Ser Lys 50 55 60Glu Glu Ala Arg Glu Val Phe Glu Asp
Asn Glu Lys Thr Glu Thr Phe65 70 75 80Trp Asn Val Tyr Val Asp Gly
Asp Gln Cys Ser Ser Asn Pro Cys His 85 90 95Tyr His Gly Thr Cys Lys
Asp Gly Ile Gly Ser Tyr Thr Cys Thr Cys 100 105 110Leu Pro Asn Tyr
Glu Gly Lys Asn Cys Glu Lys Val Leu Phe Lys Ser 115 120 125Cys Arg
Ala Phe Asn Gly Asn Cys Trp His Phe Cys Lys Arg Val Gln 130 135
140Ser Glu Thr Gln Cys Ser Cys Ala Glu Ser Tyr Arg Leu Gly Val
Asp145 150 155 160Gly His Ser Cys Val Ala Glu Gly Asp Phe Ser Cys
Gly Arg Asn Ile 165 170 175Lys Ala Arg Asn Lys Arg Glu Ala Ser Leu
Pro Asp Phe Val Gln Ser 180 185 190Gln Lys Ala Thr Leu Leu Lys Lys
Ser Asp Asn Pro Ser Pro Asp Ile 195 200 205Arg Ile Val Asn Gly Met
Asp Ser Lys Leu Gly Glu Cys Pro Trp Gln 210 215 220Ala Val Leu Ile
Asn Glu Lys Gly Glu Val Phe Cys Gly Gly Thr Ile225 230 235 240Leu
Ser Pro Ile His Val Leu Thr Ala Ala His Cys Ile Asn Gln Thr 245 250
255Lys Ser Val Ser Val Ile Val Gly Glu Ile Asp Ile Ser Arg Lys Glu
260 265 270Thr Arg Arg Leu Leu Ser Val Asp Lys Ile Tyr Val His Thr
Lys Phe 275 280 285Val Pro Pro Asn Tyr Tyr Tyr Gly His Gln Asn Phe
Asp Arg Val Ala 290 295 300Tyr Asp Tyr Asp Ile Ala Ile Ile Arg Met
Lys Thr Pro Ile Gln Phe305 310 315 320Ser Glu Asn Val Val Pro Ala
Cys Leu Pro Thr Ala Asp Phe Ala Asn 325 330 335Glu Val Leu Met Lys
Gln Asp Ser Gly Ile Val Ser Gly Phe Gly Arg 340 345 350Ile Arg Phe
Lys Glu Pro Thr Ser Asn Thr Leu Lys Val Ile Thr Val 355 360 365Pro
Tyr Val Asp Arg His Thr Cys Met Leu Ser Ser Asp Phe Arg Ile 370 375
380Thr Gln Asn Met Phe Cys Ala Gly Tyr Asp Thr Leu Pro Gln Asp
Ala385 390 395 400Cys Glu Gly Asp Ser Gly Gly Pro His Ile Thr Ala
Tyr Gly Asp Thr 405 410 415His Phe Ile Thr Gly Ile Val Ser Trp Gly
Glu Gly Cys Ala Arg Lys 420 425 430Gly Lys Tyr Gly Val Tyr Thr Lys
Val Ser Arg Phe Ile Pro Trp Ile 435 440 445Lys Lys Ile Met Ser Leu
Lys 450 455391362DNANotechis scutatus 39atggctcctc aactactcct
ctgtctgatc ctcacttttc tgtggagtct cccagaggct 60gaaagtaatg tattcttaaa
aagcaaagtg gcaaatagat ttttgcaaag aacaaaacga 120tctaattcac
tgtttgagga aattagacct ggaaacattg aaagggaatg cattgaggag
180aaatgttcaa aagaagaagc cagggaggta tttgaagata acgagaaaac
tgagaccttc 240tggaatgttt atgtagatgg ggatcagtgt tcatcaaacc
cctgtcatta tcgcgggaca 300tgcaaagatg gcattggtag ctatacctgt
acctgcttgc ctaactatga agggaaaaac 360tgtgaaaaag tcttatttaa
gtcctgcaga gcattcaatg gtaactgttg gcacttctgc 420aaacgtgttc
aaagtgaaac tcagtgttca tgtgctgaaa gttacctttt gggagttgat
480gggcactctt gtgttgctga aggtgacttt tcatgtggta gaaatataaa
agcaaggaac 540aagagggaag caagtctgcc tgactttgtg cagtcccaga
aggcaactgt gctgaaaaaa 600tctgataatc caagccctga tatcagaatt
gttaatggaa tggactgcaa actgggtgaa 660tgtccatggc aggcagttct
gataaatgaa aaaggagaag tgttttgtgg aggaacaatt 720ttgagcccca
tccatgtgct tactgcagcc cactgcatta accagaccaa gagcgtttca
780gttattgtag gggaaataga catatcaaga aaagaaacca gacgtcttct
ttctgtggat 840aaaatatatg tgcataaaaa atttgttcct cccaactctt
actatcaaaa cattgatcgt 900ttcgcctatg actatgatat agccatcatc
cgaatgaaga cccctatcca gttctctgaa 960aatgtggttc ctgcctgcct
tcccactgct gattttgcca aggaagtcct catgaaacaa 1020gattctggca
tcgttagtgg atttgggcgt actcaatcta taggatatac ctctaacata
1080cttaaagtca ttacggttcc ttatgtggac aggcacacct gcatgctttc
cagtaatttt 1140cgaattactc aaaatatgtt ctgtgctggc tatgatactc
tgcctcaaga tgcatgccag 1200ggagacagtg gggggcccca catcactgca
tacggagata cccactttgt tactgggatt 1260atcagctggg gggaaggatg
tgcacggaaa ggcaaatatg gtgtttacac aaaagtgtcc 1320aatttcatcc
cttggataaa aaaaataatg agtctaaagt aa 136240453PRTNotechis scutatus
40Met Ala Pro Gln Leu Leu Leu Cys Leu Ile Leu Thr Phe Leu Trp Ser1
5 10 15Leu Pro Glu Ala Glu Ser Asn Val Phe Leu Lys Ser Lys Val Ala
Asn 20 25 30Arg Phe Leu Gln Arg Thr Lys Arg Ser Asn Ser Leu Phe Glu
Glu Ile 35 40 45Arg Pro Gly Asn Ile Glu Arg Glu Cys Ile Glu Glu Lys
Cys Ser Lys 50 55 60Glu Glu Ala Arg Glu Val Phe Glu Asp Asn Glu Lys
Thr Glu Thr Phe65 70 75 80Trp Asn Val Tyr Val Asp Gly Asp Gln Cys
Ser Ser Asn Pro Cys His 85 90 95Tyr Arg Gly Thr Cys Lys Asp Gly Ile
Gly Ser Tyr Thr Cys Thr Cys 100 105 110Leu Pro Asn Tyr Glu Gly Lys
Asn Cys Glu Lys Val Leu Phe Lys Ser 115 120 125Cys Arg Ala Phe Asn
Gly Asn Cys Trp His Phe Cys Lys Arg Val Gln 130 135 140Ser Glu Thr
Gln Cys Ser Cys Ala Glu Ser Tyr Leu Leu Gly Val Asp145 150 155
160Gly His Ser Cys Val Ala Glu Gly Asp Phe Ser Cys Gly Arg Asn Ile
165 170 175Lys Ala Arg Asn Lys Arg Glu Ala Ser Leu Pro Asp Phe Val
Gln Ser 180 185 190Gln Lys Ala Thr Val Leu Lys Lys Ser Asp Asn Pro
Ser Pro Asp Ile 195 200 205Arg Ile Val Asn Gly Met Asp Cys Lys Leu
Gly Glu Cys Pro Trp Gln 210 215 220Ala Val Leu Ile Asn Glu Lys Gly
Glu Val Phe Cys Gly Gly Thr Ile225 230 235 240Leu Ser Pro Ile His
Val Leu Thr Ala Ala His Cys Ile Asn Gln Thr 245 250 255Lys Ser Val
Ser Val Ile Val Gly Glu Ile Asp Ile Ser Arg Lys Glu 260 265 270Thr
Arg Arg Leu Leu Ser Val Asp Lys Ile Tyr Val His Lys Lys Phe 275 280
285Val Pro Pro Asn Ser Tyr Tyr Gln Asn Ile Asp Arg Phe Ala Tyr Asp
290 295 300Tyr Asp Ile Ala Ile Ile Arg Met Lys Thr Pro Ile Gln Phe
Ser Glu305 310 315 320Asn Val Val Pro Ala Cys Leu Pro Thr Ala Asp
Phe Ala Lys Glu Val 325 330 335Leu Met Lys Gln Asp Ser Gly Ile Val
Ser Gly Phe Gly Arg Thr Gln 340 345 350Ser Ile Gly Tyr Thr Ser Asn
Ile Leu Lys Val Ile Thr Val Pro Tyr 355 360 365Val Asp Arg His Thr
Cys Met Leu Ser Ser Asn Phe Arg Ile Thr Gln 370 375 380Asn Met Phe
Cys Ala Gly Tyr Asp Thr Leu Pro Gln Asp Ala Cys Gln385 390 395
400Gly Asp Ser Gly Gly Pro His Ile Thr Ala Tyr Gly Asp Thr His Phe
405 410 415Val Thr Gly Ile Ile Ser Trp Gly Glu Gly Cys Ala Arg Lys
Gly Lys 420 425 430Tyr Gly Val Tyr Thr Lys Val Ser Asn Phe Ile Pro
Trp Ile Lys Lys 435 440 445Ile Met Ser Leu Lys
450411368DNATropidechis carinatus 41atggctcctc aactactcct
ctgtctgatc ctcacttttc tgtggagtct cccagaggct 60gaaagtaatg tattcttaaa
aagcaaagtg gcaaatagat ttttgcaaag aacaaaacga 120tctaattcac
tgtttgagga aattagacct ggaaacattg aaagggaatg cattgaggag
180aaatgttcaa aagaagaagc cagggaggta tttgaagata acgagaaaac
tgagaccttc 240tggaatgttt atgtagatgg ggatcagtgt tcatcaaacc
cctgtcatta tcgcgggaca 300tgcaaagatg gcattggtag ctatacctgt
acctgcttgc ctaactatga agggaaaaac 360tgtgaaaaag tcttatatca
gtcctgcaga gtggacaatg gtaactgttg gcacttctgc 420aaacgtgttc
aaagtgaaac tcagtgttca tgtgctgaaa gttaccgttt gggagttgat
480gggcactctt gtgttgctga aggtgacttt tcatgtggta gaaatataaa
agcaaggaac 540aagagggaag caagtctgcc tgactttgtg cagtcccaaa
aggcaacttt gctgaaaaaa 600tctgataatc caagccctga tatcagaatt
gttaatggaa tggactgcaa actgggtgaa 660tgtccatggc aggcagttct
gataaatgaa aaaggagaag tgttttgtgg aggaacaatt 720ttgagtccca
tccatgtgct tactgcagcc cactgcatta accagaccaa gagcgtttca
780gttattgtag gggaaataga catatcaaga aaagaaacca gacgtcttct
ttctgtggat 840aaaatatatg tgcatacaaa atttgttcct cccaactatt
actatgtgca tcaaaacttt 900gatcgtgtcg cctatgacta tgatatagcc
atcatccgaa tgaagacccc tatccagttc 960tctgaaaatg tggttcctgc
ctgccttccc actgctgatt ttgccaacga agtcctcatg 1020aaacaagatt
ctggcatcgt tagtggattt gggcgtattc aatttaaaca accgacctct
1080aacacactta aagtcattac ggttccttat gtggacaggc acacctgcat
gctttccagt 1140gattttcgaa ttactcaaaa tatgttctgt gctggctatg
atactctgcc tcaagatgca 1200tgccagggag acagtggggg gccccacatc
actgcataca gagataccca ctttattact 1260gggattatca gctgggggga
aggatgtgca cggaaaggca aatatggtgt ttacacaaaa 1320gtgtccaaat
tcatcccttg gataaaaaaa ataatgagtc taaagtaa 136842455PRTTropidechis
carinatus 42Met Ala Pro Gln Leu Leu Leu Cys Leu Ile Leu Thr Phe Leu
Trp Ser1 5 10 15Leu Pro Glu Ala Glu Ser Asn Val Phe Leu Lys Ser Lys
Val Ala Asn 20 25 30Arg Phe Leu Gln Arg Thr Lys Arg Ser Asn Ser Leu
Phe Glu Glu Ile 35 40 45Arg Pro Gly Asn Ile Glu Arg Glu Cys Ile Glu
Glu Lys Cys Ser Lys 50 55 60Glu Glu Ala Arg Glu Val Phe Glu Asp Asn
Glu Lys Thr Glu Thr Phe65 70 75 80Trp Asn Val Tyr Val Asp Gly Asp
Gln Cys Ser Ser Asn Pro Cys His 85 90 95Tyr Arg Gly Thr Cys Lys Asp
Gly Ile Gly Ser Tyr Thr Cys Thr Cys 100 105 110Leu Pro Asn Tyr Glu
Gly Lys Asn Cys Glu Lys Val Leu Tyr Gln Ser 115 120 125Cys Arg Val
Asp Asn Gly Asn Cys Trp His Phe Cys Lys Arg Val Gln 130 135 140Ser
Glu Thr Gln Cys Ser Cys Ala Glu Ser Tyr Arg Leu Gly Val Asp145 150
155 160Gly His Ser Cys Val Ala Glu Gly Asp Phe Ser Cys Gly Arg Asn
Ile 165 170 175Lys Ala Arg Asn Lys Arg Glu Ala Ser Leu Pro Asp Phe
Val Gln Ser 180 185 190Gln Lys Ala Thr Leu Leu Lys Lys Ser Asp Asn
Pro Ser Pro Asp Ile 195 200 205Arg Ile Val Asn Gly Met Asp Cys Lys
Leu Gly Glu Cys Pro Trp Gln 210 215 220Ala Val Leu Ile Asn Glu Lys
Gly Glu Val Phe Cys Gly Gly Thr Ile225 230 235 240Leu Ser Pro Ile
His Val Leu Thr Ala Ala His Cys Ile Asn Gln Thr 245 250 255Lys Ser
Val Ser Val Ile Val Gly Glu Ile Asp Ile Ser Arg Lys Glu 260 265
270Thr Arg Arg Leu Leu Ser Val Asp Lys Ile Tyr Val His Thr Lys Phe
275 280 285Val Pro Pro Asn Tyr Tyr Tyr Val His Gln Asn Phe Asp Arg
Val Ala 290 295 300Tyr Asp Tyr Asp Ile Ala Ile Ile Arg Met Lys Thr
Pro Ile Gln Phe305 310 315 320Ser Glu Asn Val Val Pro Ala
Cys Leu Pro Thr Ala Asp Phe Ala Asn 325 330 335Glu Val Leu Met Lys
Gln Asp Ser Gly Ile Val Ser Gly Phe Gly Arg 340 345 350Ile Gln Phe
Lys Gln Pro Thr Ser Asn Thr Leu Lys Val Ile Thr Val 355 360 365Pro
Tyr Val Asp Arg His Thr Cys Met Leu Ser Ser Asp Phe Arg Ile 370 375
380Thr Gln Asn Met Phe Cys Ala Gly Tyr Asp Thr Leu Pro Gln Asp
Ala385 390 395 400Cys Gln Gly Asp Ser Gly Gly Pro His Ile Thr Ala
Tyr Arg Asp Thr 405 410 415His Phe Ile Thr Gly Ile Ile Ser Trp Gly
Glu Gly Cys Ala Arg Lys 420 425 430Gly Lys Tyr Gly Val Tyr Thr Lys
Val Ser Lys Phe Ile Pro Trp Ile 435 440 445Lys Lys Ile Met Ser Leu
Lys 450 45543671DNADemansia vestigiata 43atggctcctc aactactcct
ctgtctgatc cagacttttc tgtggagtct cccagaggct 60gaaagtaatg tattcttaaa
aagcaatgtg gcaaatagat ttttgcaaag aacaaaacga 120gctaattcag
ggtttgagga aatttaccct gcaaactttg aaagggaatg cgttgaggag
180agatgttcaa aagaagaagc cagggaggta tttgaagatg acgagaaaac
tgaggccttc 240tggactgttt atgtagatgg ggatcagtgt ttatcaaacc
cctgtcatta tggcgggaca 300tgcaaagatg gcattggtag ctatacctgt
acctgcttgg ctggctatga agggaaaaac 360tgtgaacatg acttacttaa
gtcctgcaga gtggacaatg gtaactgttg gcacttctgc 420aaacctgttc
aaaacgacac tcagtgttca tgtgctgaag gttaccgttt gggagataat
480gggttctctt gtattgctga aggtgagttt tcatgtggca gaaatataaa
atcaaggaac 540aagagggaag caagtctgcc tgactttcaa acagattttt
ctgatgacta tgatgcgatt 600gatgaaaata atttgattga aactgtgcag
tcccagagtg caactttgct gaaaaaatct 660gataatccaa a
67144473PRTDemansia vestigiata 44Met Ala Pro Gln Leu Leu Leu Cys
Leu Ile Gln Thr Phe Leu Trp Ser1 5 10 15Leu Pro Glu Ala Glu Ser Asn
Val Phe Leu Lys Ser Asn Val Ala Asn 20 25 30Arg Phe Leu Gln Arg Thr
Lys Arg Ala Asn Ser Gly Phe Glu Glu Ile 35 40 45Tyr Pro Ala Asn Phe
Glu Arg Glu Cys Val Glu Glu Arg Cys Ser Lys 50 55 60Glu Glu Ala Arg
Glu Val Phe Glu Asp Asp Glu Lys Thr Glu Ala Phe65 70 75 80Trp Thr
Val Tyr Val Asp Gly Asp Gln Cys Leu Ser Asn Pro Cys His 85 90 95Tyr
Gly Gly Thr Cys Lys Asp Gly Ile Gly Ser Tyr Thr Cys Thr Cys 100 105
110Leu Ala Gly Tyr Glu Gly Lys Asn Cys Glu His Asp Leu Leu Lys Ser
115 120 125Cys Arg Val Asp Asn Gly Asn Cys Trp His Phe Cys Lys Pro
Val Gln 130 135 140Asn Asp Thr Gln Cys Ser Cys Ala Glu Gly Tyr Arg
Leu Gly Asp Asn145 150 155 160Gly Phe Ser Cys Ile Ala Glu Gly Glu
Phe Ser Cys Gly Arg Asn Ile 165 170 175Lys Ser Arg Asn Lys Arg Glu
Ala Ser Leu Pro Asp Phe Gln Thr Asp 180 185 190Phe Ser Asp Asp Tyr
Asp Ala Ile Asp Glu Asn Asn Leu Ile Glu Thr 195 200 205Val Gln Ser
Gln Ser Ala Thr Leu Leu Lys Lys Ser Asp Asn Pro Asn 210 215 220Pro
Asp Ile Arg Ile Val Asn Gly Leu Asp Cys Lys Leu Gly Glu Cys225 230
235 240Pro Trp Gln Ala Val Leu Ile Asp Glu Lys Gly Thr Ala Phe Gly
Gly 245 250 255Gly Thr Ile Leu Ser Pro Tyr Phe Val Leu Thr Ala Ala
His Cys Ile 260 265 270Asn Lys Thr Lys Ser Ile Ala Val Val Val Gly
Gln Val Asp Ile Ser 275 280 285Arg Lys Glu Thr Arg Arg Leu Leu Ser
Val Asp Lys Val Tyr Thr His 290 295 300Pro Lys Tyr Val His Val Thr
Asn Asp Tyr Asp Ile Ala Ile Ile Gln305 310 315 320Leu Lys Thr Pro
Ile Gln Phe Ser Glu Asn Val Val Pro Ala Cys Leu 325 330 335Pro Thr
Ala Asp Phe Ala Asn His Val Leu Met Lys Gln Asp Phe Gly 340 345
350Ile Val Ser Gly Phe Gly Arg Ile Glu Glu Lys Gly Pro Thr Ser Asn
355 360 365Ile Leu Lys Val Val Met Val Pro Tyr Val Asp Arg His Thr
Cys Ile 370 375 380Leu Ser Thr Lys Ile Pro Ile Thr Arg Asn Met Phe
Cys Ala Gly Tyr385 390 395 400Gly Asn Gln Pro Glu Asp Ala Cys Glu
Gly Asp Ser Gly Gly Pro His 405 410 415Ile Thr Ala Tyr Lys Asp Thr
His Phe Leu Thr Gly Ile Val Ser Trp 420 425 430Gly Glu Gly Cys Gly
Arg Asp Gly Lys Tyr Gly Ile Tyr Thr Lys Val 435 440 445Ser Asn Phe
Leu Pro Trp Ile Lys Thr Ile Met Arg Arg Lys Gln Pro 450 455 460Ser
Thr Glu Ser Ser Thr Gly Arg Leu465 470451416DNADemansia vestigiata
45atggctcctc aactactcct ctgtctgatc ctcacttttc tgtggagtct cccagaggct
60gaaagtaatg tattcttaaa aagcaatgtg gcaaatagat ttttgcaaag aacaaaacga
120gctaattcaa tatttgaaga aattagacct ggaaacattg aaagggaatg
cgttgaggaa 180aaatgttcaa aagaagaagc cagggaggta tttcaagata
atgagaaaac tgaggccttc 240tggactgttt atgtagatgg ggatcagtgt
ttatcaaacc cctgtcatta tcgtgggaca 300tgcaaagatg gcattggtag
ctatacctgt acctgcttgc ctggctatga agggaaaaac 360tgtgaacatg
tcgtagttaa gtcctgcaga ctgttcaatg gtaactgttg gcacttctgc
420aaaactgttc aaaacgacac tcagtgttca tgtgctgaag gttaccgttt
gggagttgat 480gggttctcct gtattgctga aggtgacttt tcatgtggca
gaattataaa atcaaggaac 540aagagggaag caagtctgcc tgactttcat
ttttctgatg actatgatgc gattgatgaa 600aataatttgg ttgaaactgt
gcagtcccag agtgcaactt tgctgaaaaa atctgataat 660ccaagccctg
atatcagaat tgttagtgga ttggactgca aactgggtga atgtccatgg
720caggcagttc tgatagatga acatggaaaa gcgtttggtg gaggaacaat
tttgagtccc 780tactttgtgc ttactgcagc ccactgcctt aaccagacca
aaagcattgc agttgttgta 840gggcaagtag acatatcaag aaaagaaacc
agacatcttc tccatgtgga taaagcatat 900atgcattcaa aatatgttcg
tgccacctat gaccatgata tagccatcct cagactgagg 960acccctatcc
agttctctga aaatgtggtt cctgcctgcc ttcccactgc tgattttgcc
1020gacgaagtcc tcatgaaaca agattttggc atcgttagtg gatttgggcg
tttgcatgaa 1080agaggatcga cctctgacat acttaaagtc attagggttc
cttatgtgga caggtacacc 1140tgcatgcttt ccagcaacta tcgaattact
ccaagtatgt tctgtgctgg ctatggtaat 1200cagcctcaag atgcatgcca
gggagacagt ggggggcccc acatcactgc atacggagat 1260acccacttta
ttactgggat tatcagctgg ggggaaggtt gtggaaggaa aggcaaatat
1320ggtatttaca caaaagtgtc caatttcatc ccttggataa aaacaataat
gcgtcgaaat 1380caacccagta cagagtcaag cactggtcgg ctctaa
141646471PRTDemansia vestigiata 46Met Ala Pro Gln Leu Leu Leu Cys
Leu Ile Leu Thr Phe Leu Trp Ser1 5 10 15Leu Pro Glu Ala Glu Ser Asn
Val Phe Leu Lys Ser Asn Val Ala Asn 20 25 30Arg Phe Leu Gln Arg Thr
Lys Arg Ala Asn Ser Ile Phe Glu Glu Ile 35 40 45Arg Pro Gly Asn Ile
Glu Arg Glu Cys Val Glu Glu Lys Cys Ser Lys 50 55 60Glu Glu Ala Arg
Glu Val Phe Gln Asp Asn Glu Lys Thr Glu Ala Phe65 70 75 80Trp Thr
Val Tyr Val Asp Gly Asp Gln Cys Leu Ser Asn Pro Cys His 85 90 95Tyr
Arg Gly Thr Cys Lys Asp Gly Ile Gly Ser Tyr Thr Cys Thr Cys 100 105
110Leu Pro Gly Tyr Glu Gly Lys Asn Cys Glu His Val Val Val Lys Ser
115 120 125Cys Arg Leu Phe Asn Gly Asn Cys Trp His Phe Cys Lys Thr
Val Gln 130 135 140Asn Asp Thr Gln Cys Ser Cys Ala Glu Gly Tyr Arg
Leu Gly Val Asp145 150 155 160Gly Phe Ser Cys Ile Ala Glu Gly Asp
Phe Ser Cys Gly Arg Ile Ile 165 170 175Lys Ser Arg Asn Lys Arg Glu
Ala Ser Leu Pro Asp Phe His Phe Ser 180 185 190Asp Asp Tyr Asp Ala
Ile Asp Glu Asn Asn Leu Val Glu Thr Val Gln 195 200 205Ser Gln Ser
Ala Thr Leu Leu Lys Lys Ser Asp Asn Pro Ser Pro Asp 210 215 220Ile
Arg Ile Val Ser Gly Leu Asp Cys Lys Leu Gly Glu Cys Pro Trp225 230
235 240Gln Ala Val Leu Ile Asp Glu His Gly Lys Ala Phe Gly Gly Gly
Thr 245 250 255Ile Leu Ser Pro Tyr Phe Val Leu Thr Ala Ala His Cys
Leu Asn Gln 260 265 270Thr Lys Ser Ile Ala Val Val Val Gly Gln Val
Asp Ile Ser Arg Lys 275 280 285Glu Thr Arg His Leu Leu His Val Asp
Lys Ala Tyr Met His Ser Lys 290 295 300Tyr Val Arg Ala Thr Tyr Asp
His Asp Ile Ala Ile Leu Arg Leu Arg305 310 315 320Thr Pro Ile Gln
Phe Ser Glu Asn Val Val Pro Ala Cys Leu Pro Thr 325 330 335Ala Asp
Phe Ala Asp Glu Val Leu Met Lys Gln Asp Phe Gly Ile Val 340 345
350Ser Gly Phe Gly Arg Leu His Glu Arg Gly Ser Thr Ser Asp Ile Leu
355 360 365Lys Val Ile Arg Val Pro Tyr Val Asp Arg Tyr Thr Cys Met
Leu Ser 370 375 380Ser Asn Tyr Arg Ile Thr Pro Ser Met Phe Cys Ala
Gly Tyr Gly Asn385 390 395 400Gln Pro Gln Asp Ala Cys Gln Gly Asp
Ser Gly Gly Pro His Ile Thr 405 410 415Ala Tyr Gly Asp Thr His Phe
Ile Thr Gly Ile Ile Ser Trp Gly Glu 420 425 430Gly Cys Gly Arg Lys
Gly Lys Tyr Gly Ile Tyr Thr Lys Val Ser Asn 435 440 445Phe Ile Pro
Trp Ile Lys Thr Ile Met Arg Arg Asn Gln Pro Ser Thr 450 455 460Glu
Ser Ser Thr Gly Arg Leu465 4704733731DNAHomo sapiens 47ctgtctgacc
gcagctctca agtgtctcag gggctgtggc tctgggcttc gtgctgtcac 60ttccacagac
agacagacat ccccaaaagg ggagcaacca tgctgggcac gactgctgtg
120gccaccgtgc tctcagccac tttcccatgc ccaaataaaa cgataaaaga
ctgggggctt 180ctgcccatcc tgcctcactt gaccaagagc ccagaagagg
atgcgacacc cagggcctca 240tgggaccacc ggctggcagg ggttctgctc
actgggttta tgggtgagac gagcactccc 300aggagggcca ctgggccggg
aagaactgtg gagaatcggg gcacgccctg tcctcccagc 360tgccagggca
cagcatccct tccccacctc aacacccaga ccccagattc accccagttc
420acttgtcccc acacgagcca caggctgcca cctggggcag gctggcccca
ccttggggtt 480agatgcaggt ccccttgccc cagaaggaga ctgcagcccc
tgcagaccta gaaatggcca 540cagcccatcc ccatgcacca gggggtgagg
tggcaggtgg tggaaagggc ctgagggggg 600cttcttcctt ccaggcgagc
acgacctcag cgagcacgac ggggatgagc agagccggcg 660ggtggcgcag
gtcatcatcc ccagcacgta cgtcccgggc accaccaacc acgacatcgc
720gctgctccgc ctgcaccagc ccgtggtcct cactgaccat gtggtgcccc
tctgcctgcc 780cgaacggacg ttctctgaga ggacgctggc cttcgtgcgc
ttctcattgg tcagcggctg 840gggccagctg ctggaccgtg gcgccacggc
cctggagctc atggtcctca acgtgccccg 900gctgatgacc caggactgcc
tgcagcagtc acggaaggtg ggagactccc caaatatcac 960ggagtacatg
ttctgtgccg gctactcgga tggcagcaag gactcctgca agggggacag
1020tggaggccca catgccaccc actaccgggg cacgtggtac ctgacgggca
tcgtcagctg 1080gggccagggc tgcgcaaccg tgggccactt tggggtgtac
accagggtct cccagtacat 1140cgagtggctg caaaagctca tgcgctcaga
gccacgccca ggagtcctcc tgcgagcccc 1200atttccctag cccagcagcc
ctggcctgtg gagagaaagc caaggctgcg tcgaactgtc 1260ctggcaccaa
atcccatata ttcttctgca gttaatgggg tagaggaggg catgggaggg
1320agggagaggt ggggagggag acagagacag aaacagagag agacagagac
agagagagac 1380tgagggagag actctgagga catggagaga gactcaaaga
gactccaaga ttcaaagaga 1440ctaatagaga cacagagatg gaatagaaaa
gatgagaggc agaggcagac aggcgctgga 1500cagaggggca ggggagtgcc
aaggttgtcc tggaggcaga cagcccagct gagcctcctt 1560acctcccttc
agccaagccc acctgcacgt gatctgctgg cctcaggctg ctgctctgcc
1620ttcattgctg gagacagtag aggcatgaac acacatggat gcacacacac
acacgccaat 1680gcacacacac agagatatgc acacacacgg atgcacacac
agatggtcac acagagatac 1740gcaaacacac cgatgcacac gcacatagag
atatgcacac acagatgcac acacagatat 1800acacatggat gcacgcacat
gccaatgcac gcacacatca gtgcacacgg atgcacagag 1860atatgcacac
accgatgtgc gcacacacag atatgcacac acatggatga gcacacacac
1920accaatgcgc acacacaccg atgtacacac acagatgcac acacagatgc
acacacaccg 1980atgctgactc catgtgtgct gtcctctgaa ggcggttgtt
tagctctcac ttttctggtt 2040cttatccatt atcatcttca cttcagacaa
ttcagaagca tcaccatgca tggtggcgaa 2100tgcccccaaa ctctccccca
aatgtatttc tcccttcgct gggtgccggg ctgcacagac 2160tattccccac
ctgcttccca gcttcacaat aaacggctgc gtctcctccg cacacctgtg
2220gtgcctgcca cccactgggt tgcccatgat tcatttttgg agcccccggt
gctcatcctc 2280tgagatgctc ttttctttca caattttcaa catcactgaa
atgaaccctc acatggaagc 2340tattttttaa aaacaaaagc tgtttgatag
atgtttgagg ctgtagctcc caggatcctg 2400tggaattgga tgttctctcc
ctgccacagc ccttgtcaat gatatttcac agagaccctg 2460ggagcacctg
ctcaagagtc agggacacac gcatcactaa atgcaagttc ccaggccctg
2520gctgcagtgg gaggacctgg caagctgcac tcttgctgag tccccagggt
ggtggaagaa 2580gaatgagaaa cacatgaaca gagaaatggg gaggtgacaa
acagtgcccc cactcagact 2640ccggcaagca cggctcagag agtggactcg
atgccatccc tgcagggccg tcctgggcac 2700cactggcact cacagcagca
aggtgggcac cattggcact cacagcagca aggcaggcac 2760cagcaaccca
cctcgggggc actcaggcat catctacttc agagcagaca gggtctatga
2820actacagccg tgggctgctt ccaaggcacc ctgctcttgt aaataaagtt
ttatgggaac 2880acacccatat tagtgtccat ggagtggccg tggcagagac
gtccagccgg acagaccagc 2940tgacccgcca agcccagcat ggttagtgtc
aggacctctg ctgaagatgc ttgctgaccc 3000tggccagacc ccggttccta
atgcccccta aacgggacgg gagccagtgg cgggccctga 3060tccaggtcag
agctggctct gctttctctt ttgtccgagt gaccatgcct cagtttcctc
3120atgtgtaaaa caggagccca ccgtgatgct tatggtggga tgagatcagc
atggatggaa 3180caaggccctg gaagggccca tgccatggtc atcgacagca
aagccactct gcagacagat 3240gcttcagtga attggtagaa aattctgcaa
ccagaatgcc cggggctcct gagggcctaa 3300gcccagccca gggttctgga
agccactctg acttcttggg agtggaagtt ggcaggactc 3360ttcctgggaa
gaagcggagg gtggggatga gaggacagtt caggagccca cccagaccca
3420caggaggaaa ctaggggagt catgcggggt cctggtggag cgccagcctc
ccttcctgcc 3480aatgggaaat gcaggcgccc acctcatggt gctgccggag
gagggggccc gggactcccc 3540agaggcttcg ctgaagggcc tgggcgcccc
caaaggctac atgtttcata tgggacgtgc 3600cacctgccac ggctcagctc
cagctttctg tgagtggcga gatagaatac ggggaggcca 3660ctggccatgg
gcctgggaca gggtgggatg aggcggcagg cttgggccac caaagccagc
3720atcgccaccc agcattgatg acaaagactg cgtgtctgcc atgagcatcc
tgctgttggt 3780gcacacaccg cattggtctc tccatacaaa catgcctaga
ggcgatgtca gagggtggag 3840accaggagag gcaggagtca gacatctggt
gccaccagga aggcccttct cagaggacca 3900ggctgtgcgt ggtgcccgcc
gtgggaggcc agcctggcgt tggcatccag catcatcagt 3960ttgtgcagtc
gggtggggct cagtgagtgc ctcctgtgtg ccaggcacaa tgacgcacaa
4020tgtgtgcaca ccaggctcat gtgcaggtgg ctgcgagaca gggcgaccca
tcaaggcaga 4080tgcaccatga ggcagtggcc agtgctgtgg gtgttagggg
cattgctccc cggccactac 4140ggcatagcag gcagtgatcg ccacactggc
caagctttag accatttatt ccagagaccc 4200cagaggcaaa aagcccggct
gcacctccca gtgactccca cagccattga gcagagacac 4260tcaggacctt
gtgatgggag gtttctgcac tggagaacga gcccagaagc cctctcagcc
4320tcggaacagt gtggccagtg gtgggcaggt caggaggggc ttcagacaca
gcctgtccct 4380ccagatggtc acgggaaggt cactccccac agaagtacgt
tttggggcca tgcgggcaca 4440gaaggtttgg gggtgggtgg ggcaggtgcc
agcctggcct gtgggaggcc atggtgcaga 4500tgccaagccc cccccgtgac
atgagaccac ctgataccac ccagagagtg gctgtgagcg 4560gaagggcccg
cccagaaaca agcagggcct tggggcagaa gtcctgggct cagatcccac
4620gctcactgcc agcggcctcg gctcaggctt ctgcgctctc taaacttagt
tttctcttct 4680ggaaaaatga tggggaaaat gatatttgta tgtgaggact
gagagttaaa tgtaaacatc 4740tggaaactac aaaatgagca cgaaatgatg
tttttattct tagaacagaa agtccccaca 4800cccgcggccc tggtgactga
tgaggatgag gttctgcggg gcctctctgg ccgcccagct 4860ctgcctgggg
aaggtggggc cagagtggat gtgttcccag cgtggtcact cccctgcctc
4920gccagcaggt ctcggctcca atcaggaggc ctaagccaag tgataagcag
ccagacaaca 4980gccatcccag ctggggcgtg gactttgctc cagcagcctg
tcccagtgag gacagggaca 5040cagtactcgg ccacaccatg gggcgcccac
tgcacctcgt cctgctcagt gcctccctgg 5100ctggcctcct gctgctcggg
gaaagtcgta agtgcccctc gcccttcaga cccaaaagca 5160gcgccaggga
gcagggaggg gcggcagttg gggaaaccct ctcatctctg cagcctggac
5220ggtgggtgcc ttgagtgctg ccagaggctg ggctcggatg gctgggcttg
gcctttccag 5280ccaacggcat cctcaaggcc agctgtggct ccctggggct
gagagtcaga cgggcggatc 5340agaggtcaca gagacaaaaa cacaaggaca
gagtcagaga gagaaaggga gagggaagga 5400gaaacggaga cacagtgaga
tgggaggcca agaggcagag acagaggtag aaagacggag 5460acagagagag
agggaggggt tggggcaggc agagacagga cagttagcca tctgccacca
5520cagggaggca caggacgagg ggcacagcag aggagctccc agggaggagg
aggctgagcc 5580gagccagtgc caccactctc ggactggctc cgtcggggaa
ggagctgcct aatgcacagc 5640tggacaggtg ggggcagcag ggctgtccag
gacccccggg tctgtccaaa agcagaggcc 5700cagacaggac agaagccagg
caagcctggg gacagcggag gaagaggagg cccctctggt 5760ggggacacga
gagacaggga ccctagactt gtttgcatcc tggacaaagt ggacaggcag
5820gggcaccaag gggacccagg cctgggaagg gaatgtgtga gggagagacg
gagcaggggg 5880agaccctcgt ggggtggaaa ggggagaccc ctgggaaggc
tgagtggatc ctcagtgcat 5940cactgaccta aacggcccct ccgcctggtg
acttggagct ccagtcacat cacacggggg 6000tcttctccat cccaccctca
accccctgcc ctccccagcc tctgtcccct gagccacatc 6060ttcctgtctc
ccacgcggaa cgggactccc gtcttcatgg ggtactgtgt ggctccaact
6120cgcccagcct tcttcctccc cctcaggcca cactgccccc tgcaggagcc
cactgtgatg 6180cttgtggtgg gatgagatca gcgtgggtgg aacaaggccc
tggaaggacc
catgccatca 6240tcatcgacag caaagctact ctgcaaacag acagatgctt
ccgcgaattg gtagaaaatt 6300ctgcagccgg aatgctctag gatcctgagg
ccctaagccc ggcccggggc tcccgaggcc 6360ctaagcccgg cccgggactc
tggaagctgc tctggcttat tgggaatgga agttggcagg 6420actcctcttc
ctgggaagaa gcagagggtg gggatgagaa gacaggccag gagcccaccc
6480agacccacag gaggaaacta ggggagttat gcagagtcct ggtggagcgc
cagcttccct 6540tcctgccaat gggaaatgca ggcgcccacc tcgcggacct
ctgggtccac agggtattgg 6600cacccttagc tgtgtgatgc gggcctggct
cataaccatg gcctgtggtg tccccggggg 6660ccggcctgga ccctgggtgg
acatggccag ccccggagag ccagggccag gccatctctc 6720tcccctactc
tgcctcagag gcctcggcag ctgcactgtg gggtgggtgg ggctgaacac
6780aggtcccaga aggtcccact caggaccctg ctgtgcacac ttttgatttt
aataaaatca 6840gaatgcgcac agcatctgca gtctagcctt taaacgagca
cagctgtcct ggcagtcacg 6900gaagttcttc tggggcggtg ggacctcagc
attcctttgc tggtactgct acaagaaagg 6960acaatggacc aagtagctta
aagcaacaga aacatttctc ccacagctct ggaggctgga 7020agttcaaaat
caaggcgtca gcggggctgg ttcctccaga ggctgggaga gagtctgctc
7080taggcctgtt tcctggcctc ggggtgccag cggcctcaca gcccatggct
tgtgggagca 7140ttgctcccat cgctgcctcc atcatcgcat ggggttctca
ccatggctgt ttttctgtct 7200tcttttctct cagaaggaca ccagtcattg
gatttagggc tcactctact cccataaaat 7260gtcttcctaa ctaaatacat
cacaaaagtt ctatttccaa ttaaggtcag gttctgaggt 7320tctgggaaag
acatgaattt ggagggacat tattcagccc tgtcctgcca cctgtgagtg
7380ttttctgcaa tccaactttt tattttaata aaatcagaat acgcagagca
cctgcagtgc 7440cagcctttaa actactgctg ttgtactggc aatcattaaa
gctacgtggc ttcagtttca 7500atctttacat tcaacaagtt taaacccatt
cttcatgagt ttggacctta ctgactgaaa 7560attttgcttg ctggtaaaac
ttgctcaaat gcagttgctg actgtggaat tcactgatgt 7620tgccaaaaca
acaaacacaa ctgtgtgctc gaggattgca atgctcccaa cagcttctga
7680agaaacaaac cacacgacaa atgtctacca atctggatgc tccatatcag
agttctagag 7740tgttccatta atttcttgag acaagtgcct aaaaaccttg
ttttaatttc gttttgccaa 7800aatcccattt tactcacatc agaaagtgtg
gccacgtggc ccagacccgg cctgctcagt 7860ctgactgaag cgttgatgcg
actcagccat aacagatagc agaagcgccc agattcagtc 7920cagagggctg
agccaggcac gccatctttc cttcattccc tcaacacata ttggttaagt
7980tccggccgtg ctaggcacgg gcatacagct gtgaccaaac acgtcaagtc
tttttccagc 8040ggagggaagg ataagcctgt cagcatgtaa tgtcagagag
gggttgggtg ctagaagaca 8100aatagcacag tgtaagggga tgaaagagac
acgagtgggg gcaacgtcag agggcgtggt 8160cagggagggt gtctcggagg
gcgctgggcc tgagccactt gggtatctgt ggaaagaatg 8220ttccagacag
ggaggtgact ggtgcaaaag tcctggggtg tgagtgttgg gctcagccag
8280ggtcagcgca gagcccagtg tggcagagag aggtgaatga gggcagagtt
gaaggtggtg 8340aggcctggga ggcctggcca tgggaagacc ctgggctctg
ttctaagacc actagaggca 8400gattctgggt agtccttgac ttccttgcat
caccctttcc accccgtgcc tgccacctgt 8460accctcttcc tcacacagtc
cagctcaacc ttataggccg tgtcctaccc ccatgagctg 8520gggagagctg
agcaggcttc agggagatgg gaaaaggcgg gaactggaca ggggctgcaa
8580aggaaaggtg acttcttact ggtcaatcag cctggggatg ctcgggggtg
gatgccaagg 8640ggaacagagc tgtggccgct atcacagaac agcgagttcc
tctaagaggt cagaggaggc 8700gcaagggatc gaccagagac agtgagggcg
tcaggctcca gttgagtggg gaccaatcct 8760tgtggcaagt ctgtgaacca
tcactgtggc tctagggtag cagagaaaaa agcaggcata 8820tgtccatctg
gccacaagga aggagaccaa ggggaagaga gaaggtacca agagaggtgt
8880tcacatggag gtgcgtcaga acaccgaggg caaggcagaa cgcggtcttc
agaccccaac 8940tggagcccag gaggcccgcg agtcccagtt tgggaaacac
taagccaggc ttggataact 9000tgtctgaggc tgtggtcatc ccaacatgag
agccagaggc cccaagggag atgggcattc 9060cccacccctc agcttcctca
gtgccttttg tggagtggag gtgacatgag gctgcaggtt 9120gcagggagcc
acgtgtgggc tgcatttcag agcaagtgtg tgggagtgga gcagacacgc
9180agagtaatgg ggcagggtca agaaataata aatctaagtc tagcgttggt
gggataatgt 9240cggtgtcact aaaagaaatg gggaatttgg aaggtaaagt
aactggaggt aggaatgaac 9300ccaagattta ctaagcacct tctgtatacc
aaactcaaca ctaggagttt atacaatctt 9360taaaacagct ctaagagtag
atattataat tcccttctgc caataataaa taataagtga 9420ctaaggtgtt
tttgaagctc aagattgctt gattctgcag tcttgtttta aattttggac
9480atggagagtt tggagttctg agatgaggga aggcatctgt catagtgaga
gagctgaaaa 9540catggagctc aggacaaaag tcagtgctag agagatcagt
tggaagagtt atctttattg 9600tagtgagaca ttgaattttt cagtggaaaa
aaagcacaga aaaagactac agagggccaa 9660gaaaagaaac tgaggctgtc
aaaatgaaaa gaggaggagg agtgggtgag agacaaagga 9720gacacaggca
gagagatgtg gacagcacag cccacagaca cacactctac aagggacaca
9780gactctcgga tccagggcat tttaaccagc agatcaagac tcatttgcca
gtcatgaaat 9840caactcagta ttttttttta aaagagtaga atcaaataga
aaactttgta ctaagtactg 9900tattttagga aatactaata aatactattt
cttgaaagtt ggcatgtatg tgtgtcctgg 9960cacttcataa attgtactat
tatagtttgt aattgaaaca gcatcaccaa tccccacata 10020acaaagagca
gagaccttaa agacgagtgg ggccaggctg aaggggcaca gcttgagcca
10080aagcacagaa cagtgggctg aaagcaccct agggagggag aattcaagga
agggggcttg 10140ggtggccagg ctgagccagc gtgagcatgt gaggaggctg
gtgagctgat tgggtgggat 10200gtgtctactt ctccagagaa gggtgcatgg
gtctagagca ggtgcctcat gctgtacctc 10260atagagaact ggggggaggg
gggaaatggg catttccctc cacagccccc aaagtgcctg 10320aggaaagtgt
tgataaagaa gccaaactct gtaaaatatt tgaagagatt tattctgagc
10380caaatgtgag gaccacgacc catgacacag ccttggaagg tcctgagaac
atgtgtccaa 10440ggtagttggg tcacagcttg attttatgta ttttaggggg
acagaagtta catacagaca 10500ccaatcaata agcataagtt gtacactggt
tcgttccaga aaaggggaag gtgggggctt 10560ccaggtcata ggtggcttca
aagatattct gattggcaat cagttgaaag agttattatc 10620taaagacctg
gaataaatgg aaaggagtat ctgggttaag ataagaggtt gtggagacca
10680aggttcttgt tatgtagatg aagactcata ggtggccacc cttagaggga
atagatggca 10740actgtttcct cttcagacct ttaaaaggtg ctacacacat
ggccaggcgc cttggctcat 10800cctgtaatcc cagcactttg ggaggctgag
gcaggtggat cacttaaggt caggagttca 10860agaccagcct ggccaacatg
gtgaaactcc atctctacta aaaatacaaa aattagccgg 10920gtgtggtggt
gcttgcctgt agtttcagct actcgggagg ctgagacagt agaatctcct
10980cctgaggcag gacaatggct tgaacctggg aggcagatgt tgcagtgagc
tgagattgtg 11040ccattgcact ccagcctgga tgacagagca aaacaccatc
ttaaagagaa aaaaaaaaaa 11100aaaaaaaaaa aaaggtgcta gactctcagc
tcagaaaaag acctggaatg gtaagggggt 11160tctctacaga atgtggattt
ccctgagata gctttgcagg gccatttcaa aatatgtcaa 11220acaaatacaa
tttggagtaa aatcatttat tttagggcct gctatatgtc atgtgatcct
11280atactagaga agtcaggttg gaaactggta tcttattgct acaaagactc
tgtttggtca 11340gcctcaaggt ctcttaacgt gaatgctggt cagctgtgcc
cgaattccaa aggaagaaat 11400aatgaggcgt gtgggacctg cttcccctca
tggcctcaac tagtctttca ggttcctatg 11460gaattccctt ggcagagagg
acgggtccac tcagtgagtt gggggcttag aattttattt 11520ttggtttaca
agagagtgat ccgccttttg tgatctggat atggcaaggg acatggcagt
11580cagggagcat aggtgagggg gagcctgggt gagggtgacc agagctttta
accctgtcct 11640ccctgccttc cagtgttcat ccgcagggag caggccaaca
acatcctggc gagggtcacg 11700agggccaatt cctttcttga agagatgaag
aaaggacacc tcgaaagaga gtgcatggaa 11760gagacctgct catacgaaga
ggcccgcgag gtctttgagg acagcgacaa gacggtaagg 11820gctggggata
gcctggctgt tggtaaggag ctcaggccac agcgccctcg ctggccccgc
11880tgctccgtcc atccaggggg gcggcctgga ggaaggggca gcgtgcgcga
aggctttcag 11940gggcggggcc cagcaaatcg aggcctcggc ggagtcctgc
ccacagggac atcagtgccg 12000cccccgcgct gactccttcc cggcgaggac
tcagcgggga gggatgcgcc caagtccctt 12060gagggtcaca gggcttctgc
cagagttaag ttctatttaa aaataaaatg ttaacctaaa 12120aaccaatagt
catggtctcg gccagcgcct cgccgagttg cagtgagctg agatcgtgcc
12180ctcccacgcc cgcagcccgc gtcctgcctt ggcctccgta gtcgctgaga
gccacagcct 12240agagcgccag cgcgcaggcg cacaactgac gccaggccac
gaacccagta ctgctcctgc 12300acagcagaag cactagcact gaggccgggc
ggcgaacccg gcactgcgcc tgcgcagcaa 12360aaggacacgc actgaggcca
ggccgcgaac ccagcacggt gcctgcgcag caggaagacc 12420ggcatccaca
ccggacgacg aacccagcat cgcgcctgcg cagtaggagg agagcaatgc
12480caccaggccg cgattgcgca gccgcagcag ccccgcgcgg aagacgctac
cctcctctcc 12540cccgaagagg cggggcttcg aacgaacctg gaaatggccg
aggggtctcg acttcctcac 12600cccaggcatc aggaaaggtg cctgcaggac
agggctctga agtggaagtg ggcggtggtg 12660atgccgaact gacaagaatg
agctgaagag aagaaagtag ccgagaaggg ggccaagccg 12720gagctcagtg
agcaacagct aggccagccg cggctggtgc catcagccac accactaact
12780gtggacccaa atggaaactt tgaggcccct tacaggtccc ccagcgcagg
ccaccgccga 12840aggttactgg ggaagacccc tgcagcttcc atgtggacat
tactcgcttg cttcatcccg 12900gagtgcagtc acctgcagcc tgggaccacc
tgactgatgt caccctggag atccatgcca 12960aaagagctgg ggaaagttca
ccttctaatg acccctgcgg accagggtga agttacaagt 13020tatggccagg
tcaaggaatt ataagtcaat aaataaactg cctcagggag acgtactcag
13080agtccaggaa aatctaggaa acaccaagaa gggcgagcgc agcatcagcc
cctgcaaaat 13140tgcactgctg cctcctggct cacatgttgc cccatcctta
ctttagcctc aaaggcaagg 13200agacccaata ttatttgagt acttggattt
gatcctgagt gactttctga gaaagcagtg 13260tatcatctgc cctaagatca
ccacatatat gacaaggttt ttggatgagt tgagatttct 13320agacattgaa
accctcatga tgaacgtcat cccgggtagc aggggctaag cctttcatca
13380cctgctacaa caagctggac attaacttgt atttgagaat tgctccagga
ctctaccttc 13440agatgctggt tagtagcatt gactggattt atgaaattgg
atgccagtta aggaataaga 13500gaatgtattt gatttgccac cctatatggt
ctgtgcagac tgtcatgatc tcattgaaat 13560ccaagaaaag attatgtcag
ggatgaagca catcacaggt ggttacaagg tcacccacca 13620tccagatggc
tcagaaagcc aaacctatgg gttgacttct cctcaccctt ctggagaatc
13680agtgtggtag aagagcttga aaaagtgctg ggtgtgacgc tgccagaaac
taacctcttt 13740ggaactgaag aaactcaaaa aattattggt gatatctatg
taacaaaagc ttttgaaagt 13800cttctacctc agaacatatt caggcgcctt
gataaacttg tcaaggagtt cctgaaagtg 13860acttacatca gtcccacatt
tatctgttat cacctgcaga taatgagctc tttgaccaaa 13920tgatctctct
aagagggtct cactgagcac tttgagctat ttgtcatgaa gatagctgaa
13980tgtttccatg cagcaacagc agcagctgtt gacaaacaag acaaggccag
caaagatgat 14040gaggccacat tcatagatga aagcttccat gcaaccctgg
aatatgggct tccgcccaca 14100gctggttgga gcgtgacaat caaatgtgtt
accatgtttc tcacagactc ccacaacatc 14160aaggaaatat ttctgactcc
tggcatgaaa cttgaagaaa gagaatgtag cagccactaa 14220tacaatggaa
agcgcaacat tgacacgtct atctagaaaa ttttaattgt ctaagttgtg
14280tgactcagat atctttgcat ttctgcaaaa gatcaaggtc tactctaatt
cttaattaaa 14340ttaagaattc ctttttatta cttgttagca aataaatggc
ttgtctctaa cagaaaaaat 14400ttagaatttt cggaaatatt ttcaaatact
tcttatatat acatatattt ttttccactg 14460gtagaatttt tctttagtaa
aagtaaataa tgctgatcca agtttatgtt tcactcagca 14520tcgtttctca
aacactcttc tttacttata tatagctacc ctatagctaa gctatatttt
14580attgtatgat gcatttactc ttttcagagt ttggccatat aagttatttc
taaatattgc 14640tattaggaaa acacatatgc atgcatttct tctagattat
catctaagag tggcttctcc 14700agagagagac gactgaatta aaggttatca
acaagttcca attccagata agatgaagaa 14760atcacattcc acactgcctc
tcccactgag tgtagctcca aaacatggat agaatgcatg 14820tagcagctat
ttgacgaccc taaaaagtaa atcgcagtgt attgcagaat aagactacaa
14880ttagatgtat gatatgatac aactggctgt gagtttatca ttttttcctc
cagtcttcca 14940gacatcactt gacctgaatc taatggacat ttataggatt
ctcaacaata gcaaagtaca 15000ctttccttcc acatatggaa aattcctcaa
ggtagactat atcctgtgtc ttaaagcata 15060cctcaataaa aagattgaac
tcacataaag tatgttttct gaccataatg gaattaaagt 15120aaaaattact
aacagaaaaa taactggaaa cttccctaag tactcggaaa ttaagtcaca
15180catgtataaa taatctgtga gtcaaagaga aaattttaag gggagtaaga
aagtattttg 15240agctgaacaa aaatgaatat gtaacataaa atctgtggga
tgcagctaaa aaagcagtgt 15300ttcaagggaa atttatagca ttaaatgctc
acatgggaaa agaaagacgg tctcaaattg 15360tttatgtaag cttccacttt
aataaactag aaaaaaagaa aaaaataaac caaaaggaaa 15420ttgaaaaagc
agaaatcaaa gaaatttaaa acaaaaataa tagacaaaat taataagctg
15480atgaaactca aacaagactg acaggaataa aaacaaacaa acaaaaacaa
gaaaaaggac 15540ctatgttgga aatggaagag aggggacatc actacagaaa
ctgtagatgt taaatgtata 15600ataagaaaat actttgaaca actctgcata
tataaatttg catgagattt gaacttggat 15660gaaatgagcc tattcttcaa
taccacaagc caccaaaaca tacacaaggt gaaagagata 15720cctgccaatt
caattcttaa tttaaaacct tctgaaaaag taatgttcag gtacagatgg
15780tttcactggt agaattttac caaacatttc aaaaagaaca ccaattctat
acaactcttc 15840cagaacatag aagagggaac acttcttagt ttgtcttagg
ccagcattac cctgatgtca 15900aaaccagaca aatactgaaa acaaaaacca
ccctacgtaa caatatctct catgaatcta 15960gacataaaaa tcctcaacaa
aatattagca aacggtgcag caatatattt ttaaaagagt 16020aataatacac
catgaccaag tgagtttttc tggggcacac atgactggct caatatttaa
16080aaataattat gtaatccacc atataaacaa aagagaacat ccacataatc
atgtcaattg 16140atgcaacaaa caaatctggc aaaatttaac atccatttat
gattttataa aaaacctatc 16200agcagaatat gaataggagg gaattttatg
aacataataa agttcatcta caaagagtct 16260acagttgata ttatacttaa
aggtgaaaac tgaaggtttt ctccctgaga ctggaacaac 16320acaagaatgt
ccattcccaa cactcctaat tcaacattat actggaagtc ctagctctaa
16380ggaaggcctt cagtaagtca agaaaaagaa ataaagttat cactatttga
agatgacatg 16440atcatgcata tagaaaatcc taaagaatgt gaaggggaaa
aaagcttgtt ttagtccctt 16500ctcacgctgc tgtgaagaac tacccgagac
tgggtaattt ataaaggaaa aaaggtttaa 16560ttgactcagt tctacatgtc
taaggagacc tcagtaaact tacaatcatg gcagaaaagg 16620aagcaaacgt
gctcttcttc acatggctgt aggagggaga agaatgagag ccgagcaaaa
16680ggggaatcct cttaaaaaaa atcagatctc atgagaacat actcccacga
gaacagcatg 16740gaggaaccac cctcacgatt cagttacctc ccacttggtc
cctctcacta cacatgggga 16800ttatgggaac tacaattcaa gatgagattt
gggtggggac agagccaaac catatcaatg 16860ctcctaaaat ttgcaaatga
gtgtaacaag gtcacagaat acaaggtcag cacatgtgtt 16920aatcacattt
ttatgtaata gcaatgcaca gttatttgta agccaaaaat ttttaaatgc
16980catttacaat tgcttcaaag aaaattatat acttatatgt aaagctaata
aaacatatac 17040aggatcttta tcccaaaatc tacaaaattc caatgaaagt
atttaaacag acctaaataa 17100atagagacac atacagtgtt catggattga
aagactcaac atattaagat atcaattttc 17160ggccgggcgc ggtggctcat
gcctgtaatc ccagcacttt gggagaccga ggtgggtgga 17220tcacctaagg
tcgggagttc gagaccagcc tggccaacat ggtgaaaccc cgtctctact
17280aaaaaaatac aaaaattagc tgggcgtggt ggtgtgcgcc tgtaatccca
gctactcggg 17340aggctgaggc aggagaatca cttgaacctg ggaggtgaag
gttgcagtga gccaagatca 17400agccattgca ctccagcctg ggcaacaaga
gcgaaactct gtctccaaaa aaaaaaaaac 17460aaaagaaaag aaagaattgt
ctttttcaac aaattatatt agtctcagtc tgtttgtgct 17520tctataacaa
aatagatcag actgggtaaa ttataaacag aataaggtta ttgctcacag
17580ttatggaggc tgggaagtcc tccaagatca agaaaccagc agatatgggg
cctgatgagg 17640gcctggtctc tgcttccaag acggtgcctc atggctgtat
cctcacctga cagaaggcag 17700aagcacagaa gggacaaaca ctgtgtgaag
cctcttttat aaggacatta atcctattca 17760caagggcaga gccttcatgg
cctaatcacc tcctaaagat ctcaccctta atactattac 17820attgtcgatt
aaattttaac atatgtatgg ggggcatgtt gagaccatag cagtgttgga
17880acaattatat atttatatgc aaaaaaatga acctgaccta aacttcacaa
ttatacaaaa 17940attaacacaa tatagataat agatccaaac ataaaataca
aaactataaa acttttagga 18000gaaaatacaa caaaatttat gacatggagc
taggcaaaaa ttcttagaca ttgacaccaa 18060aagaatgatt aataaaagaa
aaaagtcata aattggactt tatcaaaatt aaaacctttt 18120gcacttcaga
aataaacact gttaagagga tgaaaataca agctacaaac taagagaaaa
18180tatttgcaaa tcacatatcc aacaaaggaa tcatattcgg aatatataaa
gaaatcttaa 18240cagatcagaa gaagaaaata aacactcagt taaacaaaag
accttaacag ccaactcgcc 18300aaagaggata tatggataga aaataaacat
gtgagaagat actcaacatt attagctctt 18360acagaaatgc agataaaaac
cacaataaga acgactatat actcatagag taaaaaacac 18420tgacacagaa
cagcgctggt taagacacgg agaaagcaga actttgatac actgctcgtg
18480ggaatgcaaa atggcacggc cactttgaaa aggaatttga cagtttctta
taaagttata 18540taaggttacc acaggactcg gcaatcccat ttctgggcat
ttaccctaga gaaatgaaaa 18600cttatttcca cataaaatcc tgtacataaa
tgtctatagc aactctagtc tttttttttt 18660tttttaattt ttattttttg
agacagagtc ttcccgttgc ccaggctggg gtgcaatggc 18720acaatctcgg
ctcactacaa cctcctcctc tcaggttcaa gtgattctcc tgcctcagcc
18780tcccaagtag ctgggattac aggtgtgtgc caccatggca ggctaattct
tgtacttttt 18840tttttttttt tttttttttt tttttttttt gagacggagt
ctcgctctgt cgcccaggct 18900ggagtgcagt ggcgggatct cggctcactg
caagctccgc ctcccgggtt cacgccattc 18960tcctgcctca gcctcccaag
tagctgggac tacaggcgcc cgccactacg cccggctaat 19020tttttgtatt
tttagtagag acggggtttc accgttttag ctgggatggt ctcgatctcc
19080tgacctcgtg atccgcccgc ctcggcctcc caaagtgctg ggattacagg
cgtgagccac 19140cgcgcccggc caattcttgt acttttagta gagatggggt
ttcaccatgt tggccaggct 19200ggtctcgaac tcctgacctc aagtgatcca
cccgcctcag cctcccaaag tgctgggatt 19260ataggcgtga ggcaccacgc
ctggccagca actctattct taattgccaa aagctggaag 19320taagataaat
gtcttttgct gggtgtaccc atacaatata acagttgtca gcaacaacaa
19380gtaagaaagt attgatacaa cttgcatgaa tttcaaaggc tttgtattga
atgaaaagct 19440agtttcacaa ggtcctataa taaacacttt catttacatg
acatgctcaa taggttgtta 19500tcattgtgat aaagaataga ttagtggtgg
gacagggttt gccaagggtt ggcaggtggg 19560ggagatggca tggggtattt
gggggggaag gatggacctg tactgcatcc tgatgatgct 19620ggtggctaca
gaagtctctc catgtgttga aattcataga agtgtacacc aaaacattgg
19680ttttgctgtg tgagaatttt aaaagtaaaa aagtaagaag atagtgtatt
tgcttcctag 19740ggcaaccata acaaagcact acaaacgcct taaaataaca
gggatttatt gtcgcagctt 19800tggagacaag tctgaaatta gggtgtcagc
agtgttggtt ccttctggag gctctgggag 19860agtctctgtc ccaggctctc
tgctcgcttc caggagcacc cggcaatcat gggcatcctt 19920gggctgcgga
cgcgtcgctc ctgtctctgc tttcatcttc gcatggcctt ctctctctgt
19980gcctctgtgt gactttttct gtctcttata aggactttct cctttattta
gggcccacac 20040tgacccagca tgatctcttc tacagccttg gcttagttaa
catctgcaaa gacccgattt 20100ccaattaagg ttctattctg aggctgcagg
tggacctgaa ttagcaggga gggcactatt 20160caaccaactg tagagagtta
aaaaacaata agcctgtgga cattttttag cgtaatctag 20220gctcttgatg
acctgtttta aactaatcag caatgaatat ttttcagcta acgtaatgac
20280tattgacaag cacgtgaccc ttgtctgaat gttaactcag gcatagcaac
taaaaaccat 20340ccattgacca gctcgggagt agcaaacaga gcaagccatt
cttggtgcaa cctgtttcta 20400ggtaattaac ttgaaaatat tttcaatatt
caacaaagat ggttcattta agatgactga 20460agccacatct tcacagatgc
agaagatctg aatagctttc ctctttagat tgaatagttc 20520tagaacaatt
cattcctaaa agtgacttcc attggggaaa atatcctatt cagcttgagt
20580cacttaatta tggttgttat tggtataaaa tgtctctgtt ttccctaata
tatttttaaa 20640tttctttttt ccttttagaa tgaattctgg aataaataca
aaggtcagta ttttttctgt 20700tttaaccttc agtgagaggg gttcatcagg
atatttgaat tttgaaaata gttcctgaat 20760ttcctttctg cttttgttct
cattttactc atttaagact ttttccctca gggtgtttcc 20820ataatagtta
ttgtaaaaga gtttttagag taattttata ctaatcctag ttttgttatt
20880gagttagaga tatatattta aatcagttca ttctcatttg aggataccaa
attccatgat 20940aacttttctt aaataaaagt gtattcggta aaagcaaaaa
acagagtctg aaagattaga 21000ttcccgacta aggtaaccac cttgatttaa
tgcttaatag catctgaagt ggcctcagtc 21060atgactacct ggtaacagta
ttcacatttc tcaaaatgac aactgggcct atctctaaat 21120gagattgtgt
aaatcctcca agaaatggga agccccgtgt tagtgtttgc cttctccttt
21180tgccccagga tgatttggaa agaggaaccc taacctcctc tcccgtcaag
gcccagccca 21240gaaatgagca tcaggctctc acctttcctc catccttcca
gttggtccct
gtggtcacct 21300ctgactgtaa acacactgca aaacaccggc aaaaatcaaa
aagctgggcc ggtgatccac 21360ctagataaag gcatcacgta cacatggcca
caaaaggggg tggatcaaat aaagtccaaa 21420gagggggagt tgtttacaga
gaaaccggaa gactcttcca gttatctgaa cggcagggcc 21480aaggttagca
cagcaaaact gtttccatga tgccggaaac agcttgcaga ctccagtttc
21540gaaatcctct ctttgcagat ggcgaccagt gtgagaccag tccttgccag
aaccagggca 21600aatgtaaaga cggcctcggg gaatacacct gcacctgttt
agaaggattc gaaggcaaaa 21660actgtgaatt atgtaggttc ctctgcttgg
tataccttca gatcagatgc ccctgaagag 21720tggcaggtgg gcgggggaag
aagtgaaaac gcctaatgaa acaatcttaa gtcatttctg 21780atttacaaag
tctgggctct attataccta ttatactgtg ccactatagc aatagaaaaa
21840aaagccccaa tatgtccccc aaacgattcg gtttgggggc atgatgagag
agacacagtc 21900acttctctgc tcctccgaga gagactgtag aacattgatg
aagcgtgtga tccattcatg 21960tgtaaacagg agtggactct ctgttttcct
tggggccaag tgcattgccc tgttattcct 22020gctccttgtg accctgtgca
gtgattctaa atcacctctt atttatgtgt atggatgcag 22080gtgtcaatat
ttgtgaatat ttgtgattgg ccaattataa aaatttgata catttaatta
22140gttctacgtg gaaaaatcac taagtgcttt ctctaatgtg gtgattaagt
tttaaataaa 22200aagttaggct actgttagat caatttccct aaggaaaaag
atttgcattt cttttaaagt 22260acttaattga tcatcttttt tttttttttt
tttgagatgg agtctcgctc tgtggcccag 22320gctagagtgc agtggcacga
tctcagctca ccgcaagctc cgcctgccag gttcacgcca 22380ttctcctgcc
tcagcctccc aagtagctgg gactataggc cccggccacc agtcccggct
22440aatttttttt ttttttaatt ttttagtgga gacggggttt caccgtgtta
gccaggatgg 22500tctcgatctc ctgacctggt gatccgcccg cttcgacctc
ccaaagtgct gggattacag 22560gcgtgaggca cagcgccggc ctaattgatc
atctttagac tgtgttctta gattggatta 22620cttttgagtt ttccctgatg
agaatatcaa ttacgcatca ttccattcca agtccgcagt 22680cgcctccctg
gaacaccatt tggtaactta tgaggcataa ccctgttcag gctcccaggg
22740ctattatgca cattttctaa aatttcaggc atgttgatct ttgcactgtg
attacttttt 22800catcaaaagc cacacagagg gatgtggagt gaccgtaatg
tgagtgctgc tggggcaggg 22860ggtaccggcc atcccggagg tgtgaggggc
aggtacctgg agcctggctt ctggctacac 22920cgggcactgc accatgagct
ccccgtgacc cgtgaggttg cccttcaagg caagtgtacc 22980tgtcgcctgg
ctctggccct ttgctcaacc caatggccgc tttgtggctg acaggcaagt
23040ggatgtagct ggcacccttg ggccagccca gcctccattt ctccagctgt
ccccagagcc 23100aacgtgcctc tcctttgcag tcacacggaa gctctgcagc
ctggacaacg gggactgtga 23160ccagttctgc cacgaggaac agaactctgt
ggtgtgctcc tgcgcccgcg ggtacaccct 23220ggctgacaac ggcaaggcct
gcattcccac aggtaggagg cacgttgggc cacagccacc 23280cgctgccgct
gggccgggcc agggaggaca agcccgtgcc agggggtggg gacacaggca
23340tgttctgggc gggcctggca ggtaacagtg acaccaagag gacaggactg
agccctgggc 23400tccgggccca ggtggttcaa acatgaagac catgaggttt
ggaaacagac ccattatttc 23460tgtaagccag atctgctgtt taacctcagc
ttccccatct gacaaatggg accaacacta 23520ttgcctgact gcttgggtga
tccctggagc actttgcatg atgcctggcc caccgcaggc 23580cctcagtctg
cattgggact gtggggggat ccagtgcaag ggctcaaagc accagggcag
23640gcaaagggca gagctggccc gaggaactgg agctaaggtg cggggctggg
ataggagtca 23700ggggacgctc aggctctgag ctccttttac caggaccagt
gttcattgaa cgtagttttt 23760cttttccttg atgaatgtgg acaacaggcg
gccagagggc agtgagcaca ggacaggcag 23820gggactgggc agggtgggga
cgagcctccc tgtcctgacc ccgtgggcat tgcctacgct 23880gggcttgcct
ggctgccggc acttccacac ggccagcaca catgaggccc tcgaaggcgg
23940ggcctaggcg tcacagctgc accttgcaca gcaaccccac tcccactcat
agctggcccg 24000acccgcagcg ttggcctcac ccgggggcat attcgaaggg
cagagttcca ggcccgcctt 24060ttcaagagcc tggtgaccca gctcaccttc
cggcttcagg tgcggctcag cccccagacc 24120gtgttctgcc cccggctacc
atgactgtcc cctccagaca caggttactc ccgagtgttc 24180tgtcactctt
cctttcatat ccttcttacc gaaaacaatt tacttccaaa gatgagtgat
24240cacgaaaaga ccgggttcca tatgcatcct tcaagcgctg cttcaattat
gtgcctgaaa 24300catctcagca agtgaaagac actgtggctg accttgctac
tggcaatgac attcaagctt 24360aagctggtta aaaaatattt taactgaagt
catttcttga catacacacg aatatttttt 24420aattctagaa acaatcacaa
atccatttaa aaccaagtgt gggccgggtg cagtagctca 24480tgcctgtaat
cccagcattt tgggaggcca aggcgggcgg atcatgaggt caggagatcg
24540agaccatcct ggccaacaca gtgaaacccc gtctctagta aaaatacaaa
aaaaaaaaaa 24600aattagctgg gcatggtggt gcacgcctgt agtcccagct
gctcaggagg ctgaggcaag 24660agaatggcgt gaacctggaa ggcggagctt
gcactgagca ctgagccgag attgcgccac 24720tgcgctccag cctgggcaac
agaatgagac tctgtctcaa aaaaaaaaaa aaaaaaaaaa 24780tcaaaaggca
aatgtgatgt gtgaaaataa aattacataa tctactttgt agtgcaaaaa
24840gttcaggctg ggcaaggtgg ctcacgcctg taatcccagc actttgggaa
gccaaggtgg 24900gtggatcacc tgaggtcagg agttcaagac cagcctggcc
aacatggtga aaccctgtct 24960ctataaaaaa aaaaatacaa aaacttagct
gggcgtggtg gcgcacgcct gtaatcccag 25020ctactcggga ggctgaggca
ggagaatcgc ctgaacccag gaggtggagg ttgcaatgag 25080ccaagatcat
gccattgcac tccagcctgg gagacaagag agaaactcca tctcaaaaaa
25140aaaaaaaagt tcagttccaa ataatggatg aactcagaac ttggaagggt
ggtgactgca 25200cacatggaca gagctgaggc acggcggggt ggaggcccct
gcggctggca gattcaccgg 25260agcctcctca gactgcgcag gagcacagca
agtaaacagc taagctgtgc ccatctgacc 25320ccagacacgt gtggccacag
agaagcccct tgccatccat tcccccctcc tctcctctcc 25380tgctccccca
cacccctgcc ttcctccaac atgtttcagc cattctcttg gccttggtgc
25440cctaattggc cgttatacaa aaggaagctt cctaacatct cggcgtggcc
tctctgggag 25500ctgtgctatt ccagacgctc tcctgtgcct ccagttgttt
gcgtgcgcca ttccttctgc 25560ctgaaaactt ttttttcttc aatgtttcat
taggaaaagt tttccaacac acagcacact 25620ggaaagaatt ttgcagggag
tcgcacacgc ccagcacttg ggttctcctg ttggcatcct 25680ccggccagat
gcattcatcc catttcgccc ggcccgtttg tctctgtcca tccgtcaagc
25740tttcttgact tcttggtgca ttttcaggca aaccgcagac gccaacactc
ccctcgctgc 25800ctgggttgct gcctggcgtc cattgttcac aggcggtcac
ctgaggggag gccaacgctc 25860ggacagctgc gctcacctgc agatccgacc
cctgccgacg acgtggggcc tcgccctgca 25920agcccgctgc ccctccgggt
gcccctgcgc tctgcctccc ggctctctga ctcttctccc 25980tcagggtgag
ctgtgcaggc tatggggagc ctctctctgt gctgaaggcc ccggccgtcc
26040tctttctttc agggccctac ccctgtggga aacagaccct ggaacgcagg
aagaggtcag 26100tggcccaggc caccagcagc agcggggagg cccctgacag
catcacatgg aagccatatg 26160atgcagccga cctggacccc accgagaacc
ccttcgacct gcttgacttc aaccagacgc 26220agcctgagag gggcgacaac
aacctcacca ggatcgtggg aggccaggaa tgcaaggacg 26280gggagtgtcc
ctggcaggta acagtaggat gtcccctcgg gcctgctgga gagaccacct
26340gtcccgctgt gcacctcggg gaggccagcc tgacacttgg aatagcaatc
cgggaaggaa 26400ctgttccgaa ctaggacaga ggggctccgc cacccaagcc
tgcctgcctg tcccctccct 26460ccgggcagcc aaggaggctg tgagctccac
agggaagtgg ccggggctga gggagaggct 26520gggcccaggc aacgcccccc
tcagcccctt cccactgggc atttccatgg ctgcccgtgg 26580catgcccagg
acgatgctgt cctgtgaaac agaagagagg gagaaggcgc agccacacgc
26640tcaagtgtcc tcaaacctcc cctacaccag gagacaaggc taaagccagg
gagccaccca 26700cactgcaggg gcatcagcgg gcaggaggac ggtgccgggt
gggcaaggcc tccatctgct 26760cttctgtttg acgggaggca gaaagagttg
gtgtcctcgc ttcatttcta attttggaat 26820ttttttaccc aaacacctaa
atcctatgga ggtagatagt accttagaga aaaacacatc 26880tacttatttt
caaaggtaaa aaagaaaatc actctttgag gcttttttgt taagagacag
26940taccttgctc tgttgcccag gctggagtgc agtgtcgcga tctcggctca
ctgcaacctc 27000cacctcctgg gttcaagcga ttctcatgcc tcagactccc
aagtagctgg aattacgggc 27060gcccgctact tacgcctggc taattttttt
ttttttttga gacggagtct cactctgtcg 27120cccaggctgg agtgcagtgg
cgcgatctcg gctcactgca acctccacct cccaggttca 27180tgtcattctc
ctgcttcagc ctcccgagta gctgggacta caggcgcctg ccaccacgcc
27240cagctaattt tttttgtatt tttagtagag acggggtttc accgtgttag
ccaggatggt 27300ctccatctcc tgacctggtg atctgcccac ctcagcctcc
caaagtgctg ggattacagg 27360cgtgagacaa tgtgcccggc catgcctggc
taattttttt atttttaata gagacaggaa 27420tttcaccatg ttggccaggc
tggtctcaaa ctccaggcct catgtgatcc accctcctca 27480gccacccaaa
gtgctggagt tacaggtgtg agccactata ccaggtccta atctttgatt
27540gttgatttgg actaatgctg ccagattaaa caaataaaag cacaatactt
tcaattaaat 27600ttcaatttca cataaactag aaatacatta aacaaaagca
caatactttc aattaaattt 27660caatttcaca taaactagaa atactttcag
tgtaagtatg ttccaagtat cgcatgaagc 27720atacatatgc gaaaaattat
ttgctgttta tctgattcaa gtcaaactag gtgtattagt 27780cagttttcac
actgctgaca catacatacc cgagactggg taatttataa agaaaaagag
27840gttgaatgaa ctcacagttc cacgtggctg gggaggcctc accgtcacgg
tggaaggcgc 27900aaggcacgtc ttacatggcg gcagcaagac agagaatgag
agaacaagca aaaggggttt 27960ccccttagaa aaccatcagc tcttgtgaga
cttattcact tccaccagaa cagcatgggg 28020aaaccgccct cacgattcag
ttacctccca ccaggtccct cccacaacac acgggaaata 28080tgggagctac
agtttgagat gagatttggg tggggacaca gccaaacctt gttgctgggc
28140atcctgtatt ttctctggca atcctcactt ggacttgaat tttcagcgcc
caaaaccaga 28200atgtcctctc ctacaagcaa gaatctcaga gctgccagcg
cccccatgaa ttcccccagg 28260tcttccccca ccccagaccg tgtggcgggt
gagcctctgt ctaactataa agagccaagc 28320gagagaggga tgcactgagg
tggctctgca atgcatgttt gttgagggcc ttctgtgtgt 28380caggcactga
gccgggtgct gtgtaggtgg gatatgaaac catgaagcct ctctgtgacc
28440aatacacaga aatctcaacc tagttaggga gctgagaccg aaatcctccc
agtcccaggc 28500actgtgtggt tggggcaaga acctcgatgc aggagacccc
accgaggatg agcaggaaaa 28560gcctcttgtg gggctgagga gctggacttg
gagctgcagg cggggttttg gaggggttcc 28620tgggctgggg gaccagggtg
gggcgccctg gagggctcac tggaggggcc ctcgcccagc 28680ctgttgaggt
ttgcgattct tgtttcctgg ttcgagtctt ggcaagtggg cctcatctgc
28740atctttagga agaatggttg gtgttcgtgt cttagaaagc ctgactttcc
ctcatgtaag 28800ctggatgatg agttgacaaa ttatgcaaaa aagaggcaaa
aacatgaccc cttttctagc 28860catgaatgtt ttaagaaatg ttttaagact
cggtattgtc agtagtttca ttggtctgta 28920catgtgccca gccactatca
caggacggga aactccccag agaaaagaaa accaaaatat 28980gcccgggctc
caaacttgca agtccagctc cctagggaca gcatgtggca cccctgtcag
29040tgcttgctcc cctgggaccg tgttccaagt cctggcaggt aggagaccct
tcacaggagc 29100tgccacaggg acccccagga agtcacctgg gatggaggtg
tccgtgcacc atgggggaca 29160ggctcacact gctgaaccgt cgggacacca
ggcaggcaca ccggttgagg cagatgatgt 29220ttctgcacag actggcgtct
cctggtccca ggtagaaatc ctgccacaga gacgggaaag 29280gctgctccca
cagggagcat cttttccaaa gcatggacag atgtgtcgtg tgcatgagac
29340tttagagagc tctgtgatgg agttggtaga aagaagagat gactccctat
atcagtgagt 29400gtgtggcaca ggcagagaaa agagacagac aaggaactgt
ccttgggtgg atggcaggag 29460accgaagagg acagcttggc atggggaggg
ccgggcagtg ccacctgaag agctggcttc 29520tcagtcaggc aacacctgtc
cacctggcca gccacactga gcctgtcacg tctgtcacag 29580gccctgctca
tcaatgagga aaacgagggt ttctgtggtg gaaccattct gagcgagttc
29640tacatcctaa cggcagccca ctgtctctac caagccaaga gattcaaggt
gagggtaggt 29700aagtgaccaa cagcccccag ggccgtggtg aggggcaccg
tcactgtctg cttttcagaa 29760accactaaag ctgatggaat ttgttgggaa
cactggttga aatcctgaaa tcctatttgt 29820aggggttagg ggcatttcac
agaggaagaa gatgaggaag cagaggaagg ggaagagtgg 29880ggaggaggac
ggggagggga ggcgaaccag cccagccctt ctcccactgg gtgtccaggt
29940ctcgggtctc cgagtctctg ggtcccgggt ctctgggtct gcatgtccag
ctaatgttct 30000gtgtctcagt gtcttttatt gggagccttc cagacctccc
tttctcttta acatactctg 30060aacaccaagc acctctgtct cttctatttt
tatttgtggg attgtttcat taacatctgt 30120ctttgtccac tagaccctag
agctgctcag tacaaaccca acacaagcta caaatgcaag 30180caatatatgt
aaactgatat ttttctaaca ttaaatgttc tattatacat tttaaaatat
30240aaaaaaacag gctgggtgtg gtggctcatg cctgtaatcc cagcactttg
ggaggctgag 30300gcgggcagat cacctgaggt caggagtttg agatcagcct
ggccagcatg gcgaaacccc 30360atctctacta aaaaatacaa aaattagccg
ggcatggtgg cacgggcctg taatcccagc 30420tacttgggaa gctgaggcag
gagaaccact tgaacccggg aggtggaggt tgcagtgagc 30480tgagattgtg
ccattgcact ccagcccggg caacagagca aaactctgtc tcaaaaaaaa
30540aaaaaaaata tatatatata tatatgtata tatatatgtg tatatatata
tacatatata 30600tacacacaca cacacaattt ccataatata tcttatttaa
ctcaacatat tgaaaatatt 30660actttttcca tgtgtaatca tgttaaaggt
gtaataacac attccgcaca ttttctttca 30720tgctaagtct ctattttacg
ttcatggcac aactatttta cactctcagc cagcggccac 30780accgcacaac
ctgggtctgg gatgccaaaa gccttcggtc ctgggacgcc tcgttggtgc
30840ccacgactgg cacagacgat gcacccgcca aaggacacag gagtggcggc
cgtctaaaga 30900accaaacgtg tgagacagga ccagtggttc cctgggcagc
aaggctgaca ggcactttta 30960tttgctgctt tgcacttccc tctatttttc
aaattttcaa aagtgatcac gtgccatttt 31020taatttaaaa aaatatatat
aacttcctta aaaagcaacg gatgtgcgag agcatgtccc 31080tggctgagct
gagcacagtc ccactcgtct gtcccagggg accggaacac ggagcaggag
31140gagggcggtg aggcggtgca cgaggtggag gtggtcatca agcacaaccg
gttcacaaag 31200gagacctatg acttcgacat cgccgtgctc cggctcaaga
cccccatcac cttccgcatg 31260aacgtggcgc ctgcctgcct ccccgagcgt
gactgggccg agtccacgct gatgacgcag 31320aagacgggga ttgtgagcgg
cttcgggcgc acccacgaga agggccggca gtccaccagg 31380ctcaagatgc
tggaggtgcc ctacgtggac cgcaacagct gcaagctgtc cagcagcttc
31440atcatcaccc agaacatgtt ctgtgccggc tacgacacca agcaggagga
tgcctgccag 31500ggggacagcg ggggcccgca cgtcacccgc ttcaaggaca
cctacttcgt gacaggcatc 31560gtcagctggg gagagggctg tgcccgtaag
gggaagtacg ggatctacac caaggtcacc 31620gccttcctca agtggatcga
caggtccatg aaaaccaggg gcttgcccaa ggccaagagc 31680catgccccgg
aggtcataac gtcctctcca ttaaagtgag atcccactca aggcctggtt
31740tgtctctcga ttgccgcctt gccctggctt ctcccgccct gttgaggtgg
gaaggtgaag 31800tgtctgtctg gaacaccagc ttccgccctt cccagctagg
ctggggattc ctccagggaa 31860tattctagtc tgtgggggca ggatggaggc
tccagggatg atactgtgcc atgactgcca 31920tgggcattcc tttccccaga
taccttcctg catctgggtc acgcccagag gcagatggga 31980gcctgtgcag
gccccgtggc gtcgggaggg gcccacacgt tggcgcagcc tccccaagac
32040cccccacttg gcctggtctc tcttgttcct cttgggaatt ggacacctcc
ccggtgactg 32100cctatgaccc gcagactccc tgggagggaa acgtccagaa
agcttctcat tggggcggac 32160attttacatt aacttaaaca accaggtgct
cttcaactgc acggtgccag gccccacccc 32220agctcaggct tgtgtggtgg
gggccacagg catcccccgg gcaggtgacc tgctcaccag 32280gcagcgacct
gacctggcac agttggcccc caccgtggcc acccttagaa ccccctgtgg
32340gctttagcat gcctgcatcc aggccacagc ctggccactg aaatcagtct
ctggagtgaa 32400gctggccagg agcttctgga agcttctgga gctcctcagg
tgctgagtgg tggtggcgtg 32460gcaggcgggg cttcgggggg ctcctcctct
cctagggtcc agatgtttag tccttgccct 32520gctgcaatcc ggcactgtcc
ctaggcctca agttaactgg ccatgaaaat caaatgaact 32580ttcggtaaac
agaaaagatt ccggacaagg cctgccgtgt gtctcccaaa cgtctcctgc
32640agtttgcgtc ttgtgtaatg tccctaagca aagttcaaca gttctagtac
aaaaactccc 32700caaaaaagtc atgagctggg caaaaccgtt cgtaaacaga
tgttgcgaag tcagggaaaa 32760tcaaagtgga caggtgttcg acctcccaga
aacggtctga ggaggggccg gtctcccagg 32820gtgggcggga gggcattcct
ggcctgcccg ctctgaggcc ttctccgtgg agctggctgt 32880cgggctcctc
gccggccctt cctggagaaa aggcttctgc ctcggagcta gcctgctgtt
32940gggctgcgtt tcctaggcag ccacgtggtc cccagggccc cagaggtaaa
ccctggactt 33000ggattcccgt ttctggaaat caaaggttga gtggggtcca
gagagaactc tgggaaaata 33060attacaattg aaacccccca tcgccatcac
tgtctgcacc ctggttcctg ccgcactggg 33120tgtctggtgc ccgtgcccgt
ctcaggatag aaaggaaact ggaggctgca gagagaagga 33180cctgatgggt
cgtagctcag catctgccga agccccatct agaaataggt tctcgtcctg
33240ggaggtgtgg gagggagcct cgggagggag acagcaggag gagaggcccc
agtcctggac 33300acgcgctggg ggttgaagtc tcggctctgc aggctcctgt
gctgcgtggc agggattttc 33360tctctgccta aatatcgtct tcataagtaa
aggcaagtgg gctaaaccta tgtcatctcc 33420gtgttaactc agaatagtct
aggcctgggc caggggacac tttgtgatct gagaccccca 33480gaattccctg
agggaggccc agctctgttt cgggaaataa ctgaagcggc tgtttgtgcg
33540aggtgagacc ctgaggaccg agagcagcag gaggtcatgg tggggagcaa
aaacgggaaa 33600agtgattccg cctgagactg agggagagag aacccaggtg
agaccctgag gactgtgagc 33660agcaggaggt cacggtgggg agtaaagatg
ggaaaagtga ttccgcctga gagtgaggga 33720gagagaacag a
3373148467PRTHomo sapiens 48Leu Leu Gly Glu Ser Leu Phe Ile Arg Arg
Glu Gln Ala Asn Asn Ile1 5 10 15Leu Ala Arg Val Thr Arg Ala Asn Ser
Phe Leu Glu Glu Met Lys Lys 20 25 30Gly His Leu Glu Arg Glu Cys Met
Glu Glu Thr Cys Ser Tyr Glu Glu 35 40 45Ala Arg Glu Val Phe Glu Asp
Ser Asp Lys Thr Asn Glu Phe Trp Asn 50 55 60Lys Tyr Lys Asp Gly Asp
Gln Cys Glu Thr Ser Pro Cys Gln Asn Gln65 70 75 80Gly Lys Cys Lys
Asp Gly Leu Gly Glu Tyr Thr Cys Thr Cys Leu Glu 85 90 95Gly Phe Glu
Gly Lys Asn Cys Glu Leu Phe Thr Arg Lys Leu Cys Ser 100 105 110Leu
Asp Asn Gly Asp Cys Asp Gln Phe Cys His Glu Glu Gln Asn Ser 115 120
125Val Val Cys Ser Cys Ala Arg Gly Tyr Thr Leu Ala Asp Asn Gly Lys
130 135 140Ala Cys Ile Pro Thr Gly Pro Tyr Pro Cys Gly Lys Gln Thr
Leu Glu145 150 155 160Arg Arg Lys Arg Ser Val Ala Gln Ala Thr Ser
Ser Ser Gly Glu Ala 165 170 175Pro Asp Ser Ile Thr Trp Lys Pro Tyr
Asp Ala Ala Asp Leu Asp Pro 180 185 190Thr Glu Asn Pro Phe Asp Leu
Leu Asp Phe Asn Gln Thr Gln Pro Glu 195 200 205Arg Gly Asp Asn Asn
Leu Thr Arg Ile Val Gly Gly Gln Glu Cys Lys 210 215 220Asp Gly Glu
Cys Pro Trp Gln Ala Leu Leu Ile Asn Glu Glu Asn Glu225 230 235
240Gly Phe Cys Gly Gly Thr Ile Leu Ser Glu Phe Tyr Ile Leu Thr Ala
245 250 255Ala His Cys Leu Tyr Gln Ala Lys Arg Phe Glu Gly Asp Arg
Asn Thr 260 265 270Glu Gln Glu Glu Gly Gly Glu Ala Val His Glu Val
Glu Val Val Ile 275 280 285Lys His Asn Arg Phe Thr Lys Glu Thr Tyr
Asp Phe Asp Ile Ala Val 290 295 300Leu Arg Leu Lys Thr Pro Ile Thr
Phe Arg Met Asn Val Ala Pro Ala305 310 315 320Cys Leu Pro Glu Arg
Asp Trp Ala Glu Ser Thr Leu Met Thr Gln Lys 325 330 335Thr Gly Ile
Val Ser Gly Phe Gly Arg Thr His Glu Lys Gly Arg Gln 340 345 350Ser
Thr Arg Leu Lys Met Leu Glu Val Pro Tyr Val Asp Arg Asn Ser 355 360
365Cys Lys Leu Ser Ser Ser Phe Ile Ile Thr Gln Asn Met Phe Cys Ala
370 375 380Gly Tyr Asp Thr Lys Gln Glu Asp Ala Cys Gln Gly Asp Ser
Gly Gly385 390 395 400Pro His Val Thr Arg Phe Lys Asp Thr Tyr Phe
Val Thr Gly Ile Val 405 410 415Ser Trp Gly Glu Gly
Cys Ala Arg Lys Gly Lys Tyr Gly Ile Tyr Thr 420 425 430Lys Val Thr
Ala Phe Leu Lys Trp Ile Asp Arg Ser Met Lys Thr Arg 435 440 445Gly
Leu Pro Lys Ala Lys Ser His Ala Pro Glu Val Ile Thr Ser Ser 450 455
460Pro Leu Lys465491529DNABos Taurus 49ctcagcaccg ccctgggcgg
cctcctgcgg ccggcgggga gcgtgttcct gccccgggac 60caggcccacc gtgtcctgca
gagagcccgc agggccaact cattcttgga ggaggtgaag 120cagggaaacc
tggagcgaga gtgcctggag gaggcctgct cactagagga ggcccgcgag
180gtcttcgagg acgcagagca gacggatgaa ttctggagta aatacaaaga
tggagaccag 240tgtgaaggcc acccgtgcct gaatcagggc cactgtaaag
acggcatcgg agactacacc 300tgcacctgtg cggaagggtt tgaaggcaaa
aactgcgagt tctccacgcg tgagatctgc 360agcctggaca atgggggctg
cgaccagttc tgcagggagg agcgcagcga ggtgcggtgc 420tcctgcgcgc
acggctacgt gctgggcgac gacagcaagt cctgcgtgtc cacagagcgc
480ttcccctgtg ggaagttcac gcagggacgc agccggcggt gggccatcca
caccagcgag 540gacgcgcttg acgccagcga gctggagcac tacgaccctg
cagacctgag ccccacagag 600agctccttgg acctgctggg cctcaacagg
accgagccca gcgccgggga ggacggcagc 660caggtggtcc ggatagtggg
cggcagggac tgcgcggagg gcgagtgccc atggcaggct 720ctgctggtca
acgaagagaa cgagggattc tgcgggggca ccatcctgaa cgagttctac
780gtcctcacgg ctgcccactg cctgcaccag gccaagaggt tcacggtgag
ggtcggcgac 840cggaacacag agcaggagga gggcaacgag atggcacacg
aggtggagat gactgtgaag 900cacagccgct ttgtcaagga gacctacgac
ttcgacatcg cggtgctgag gctcaagacg 960cccatccggt tccgccggaa
cgtggcgccc gcctgcctgc ccgagaagga ctgggcggag 1020gccacgctga
tgacccagaa gacgggcatc gtcagcggct tcgggcgcac gcacgagaag
1080ggccgcctgt cgtccacgct caagatgctg gaggtgccct acgtggaccg
cagcacctgt 1140aagctgtcca gcagcttcac catcacgccc aacatgttct
gcgccggcta cgacacccag 1200cccgaggacg cctgccaggg cgacagtggc
ggcccccacg tcacccgctt caaggacacc 1260tacttcgtca caggcatcgt
cagctgggga gaagggtgcg cgcgcaaggg caagttcggc 1320gtctacacca
aggtctccaa cttcctcaag tggatcgaca agatcatgaa ggccagggca
1380ggggccgcgg gcagccgcgg ccacagtgaa gcccctgcca cctggacggt
cccgcccccc 1440cttccgctct gagcgggctc cctccctgcc tgattagagc
tgtgtcctct ccttaaaaaa 1500aaaaaaaaaa aaaaaaaaaa aaaaaaaaa
152950492PRTBos Taurus 50Met Ala Gly Leu Leu His Leu Val Leu Leu
Ser Thr Ala Leu Gly Gly1 5 10 15Leu Leu Arg Pro Ala Gly Ser Val Phe
Leu Pro Arg Asp Gln Ala His 20 25 30Arg Val Leu Gln Arg Ala Arg Arg
Ala Asn Ser Phe Leu Glu Glu Val 35 40 45Lys Gln Gly Asn Leu Glu Arg
Glu Cys Leu Glu Glu Ala Cys Ser Leu 50 55 60Glu Glu Ala Arg Glu Val
Phe Glu Asp Ala Glu Gln Thr Asp Glu Phe65 70 75 80Trp Ser Lys Tyr
Lys Asp Gly Asp Gln Cys Glu Gly His Pro Cys Leu 85 90 95Asn Gln Gly
His Cys Lys Asp Gly Ile Gly Asp Tyr Thr Cys Thr Cys 100 105 110Ala
Glu Gly Phe Glu Gly Lys Asn Cys Glu Phe Ser Thr Arg Glu Ile 115 120
125Cys Ser Leu Asp Asn Gly Gly Cys Asp Gln Phe Cys Arg Glu Glu Arg
130 135 140Ser Glu Val Arg Cys Ser Cys Ala His Gly Tyr Val Leu Gly
Asp Asp145 150 155 160Ser Lys Ser Cys Val Ser Thr Glu Arg Phe Pro
Cys Gly Lys Phe Thr 165 170 175Gln Gly Arg Ser Arg Arg Trp Ala Ile
His Thr Ser Glu Asp Ala Leu 180 185 190Asp Ala Ser Glu Leu Glu His
Tyr Asp Pro Ala Asp Leu Ser Pro Thr 195 200 205Glu Ser Ser Leu Asp
Leu Leu Gly Leu Asn Arg Thr Glu Pro Ser Ala 210 215 220Gly Glu Asp
Gly Ser Gln Val Val Arg Ile Val Gly Gly Arg Asp Cys225 230 235
240Ala Glu Gly Glu Cys Pro Trp Gln Ala Leu Leu Val Asn Glu Glu Asn
245 250 255Glu Gly Phe Cys Gly Gly Thr Ile Leu Asn Glu Phe Tyr Val
Leu Thr 260 265 270Ala Ala His Cys Leu His Gln Ala Lys Arg Phe Thr
Val Arg Val Gly 275 280 285Asp Arg Asn Thr Glu Gln Glu Glu Gly Asn
Glu Met Ala His Glu Val 290 295 300Glu Met Thr Val Lys His Ser Arg
Phe Val Lys Glu Thr Tyr Asp Phe305 310 315 320Asp Ile Ala Val Leu
Arg Leu Lys Thr Pro Ile Arg Phe Arg Arg Asn 325 330 335Val Ala Pro
Ala Cys Leu Pro Glu Lys Asp Trp Ala Glu Ala Thr Leu 340 345 350Met
Thr Gln Lys Thr Gly Ile Val Ser Gly Phe Gly Arg Thr His Glu 355 360
365Lys Gly Arg Leu Ser Ser Thr Leu Lys Met Leu Glu Val Pro Tyr Val
370 375 380Asp Arg Ser Thr Cys Lys Leu Ser Ser Ser Phe Thr Ile Thr
Pro Asn385 390 395 400Met Phe Cys Ala Gly Tyr Asp Thr Gln Pro Glu
Asp Ala Cys Gln Gly 405 410 415Asp Ser Gly Gly Pro His Val Thr Arg
Phe Lys Asp Thr Tyr Phe Val 420 425 430Thr Gly Ile Val Ser Trp Gly
Glu Gly Cys Ala Arg Lys Gly Lys Phe 435 440 445Gly Val Tyr Thr Lys
Val Ser Asn Phe Leu Lys Trp Ile Asp Lys Ile 450 455 460Met Lys Ala
Arg Ala Gly Ala Ala Gly Ser Arg Gly His Ser Glu Ala465 470 475
480Pro Ala Thr Trp Thr Val Pro Pro Pro Leu Pro Leu 485
4905130PRTEchis carinatus 51Asp Cys Leu Pro Gly Trp Ser Ser His Glu
Gly His Cys Tyr Lys Val1 5 10 15Phe Asn Gln Glu Met Tyr Trp Ala Asp
Ala Glu Lys Phe Cys 20 25 305230PRTEchis carinatus 52Asp Cys Leu
Pro Asp Trp Phe His Tyr Glu Gly His Cys Tyr Arg Val1 5 10 15Phe Asp
Glu Pro Lys Lys Trp Ala Asp Ala Glu Lys Phe Cys 20 25
305311PRTArtificial sequenceSyntheticmisc_feature(3)..(4)Xaa can be
any naturally occurring amino acidmisc_feature(6)..(7)Xaa can be
any naturally occurring amino acidmisc_feature(9)..(10)Xaa can be
any naturally occurring amino acid 53His Glu Xaa Xaa His Xaa Xaa
Gly Xaa Xaa His1 5 10
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
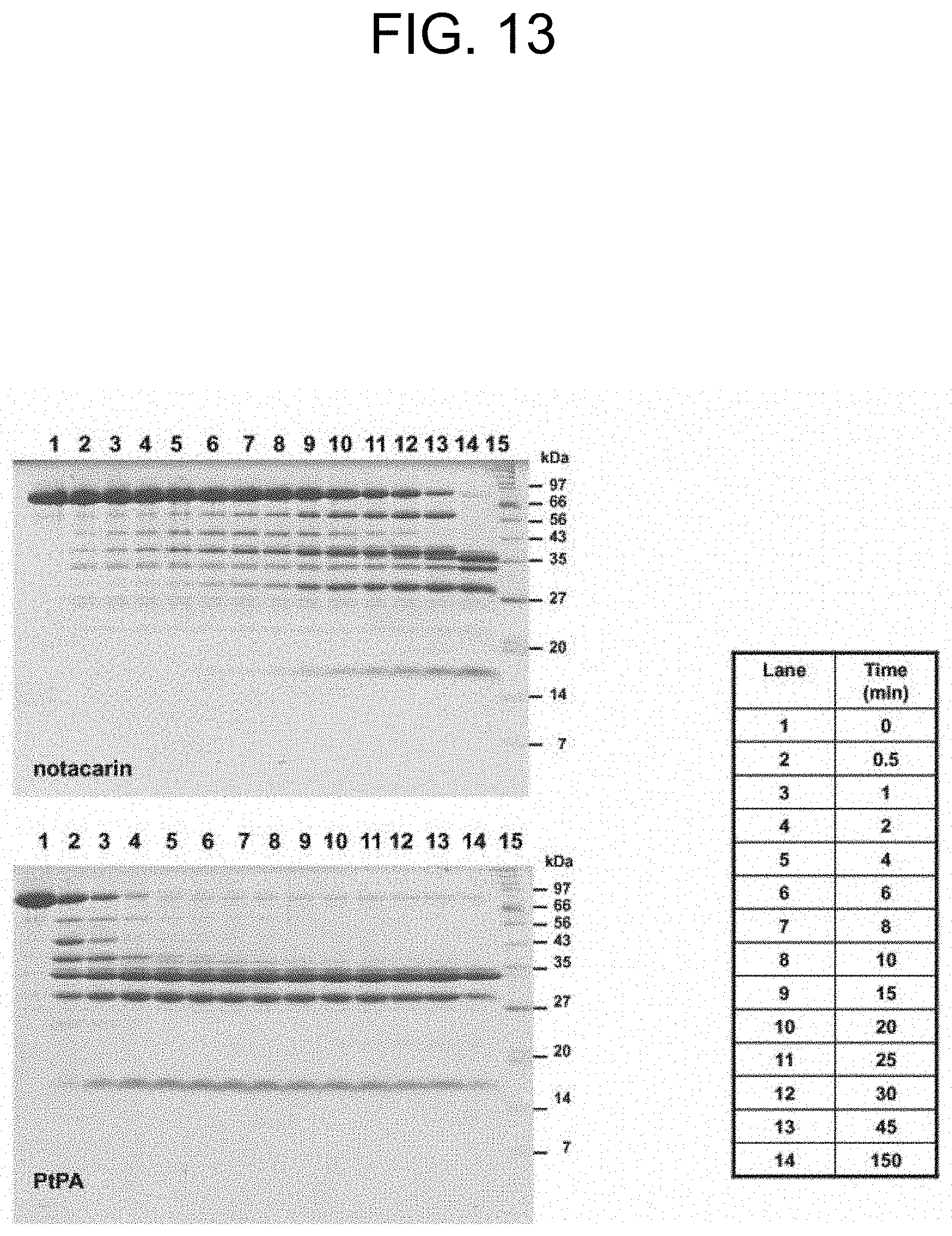
D00014
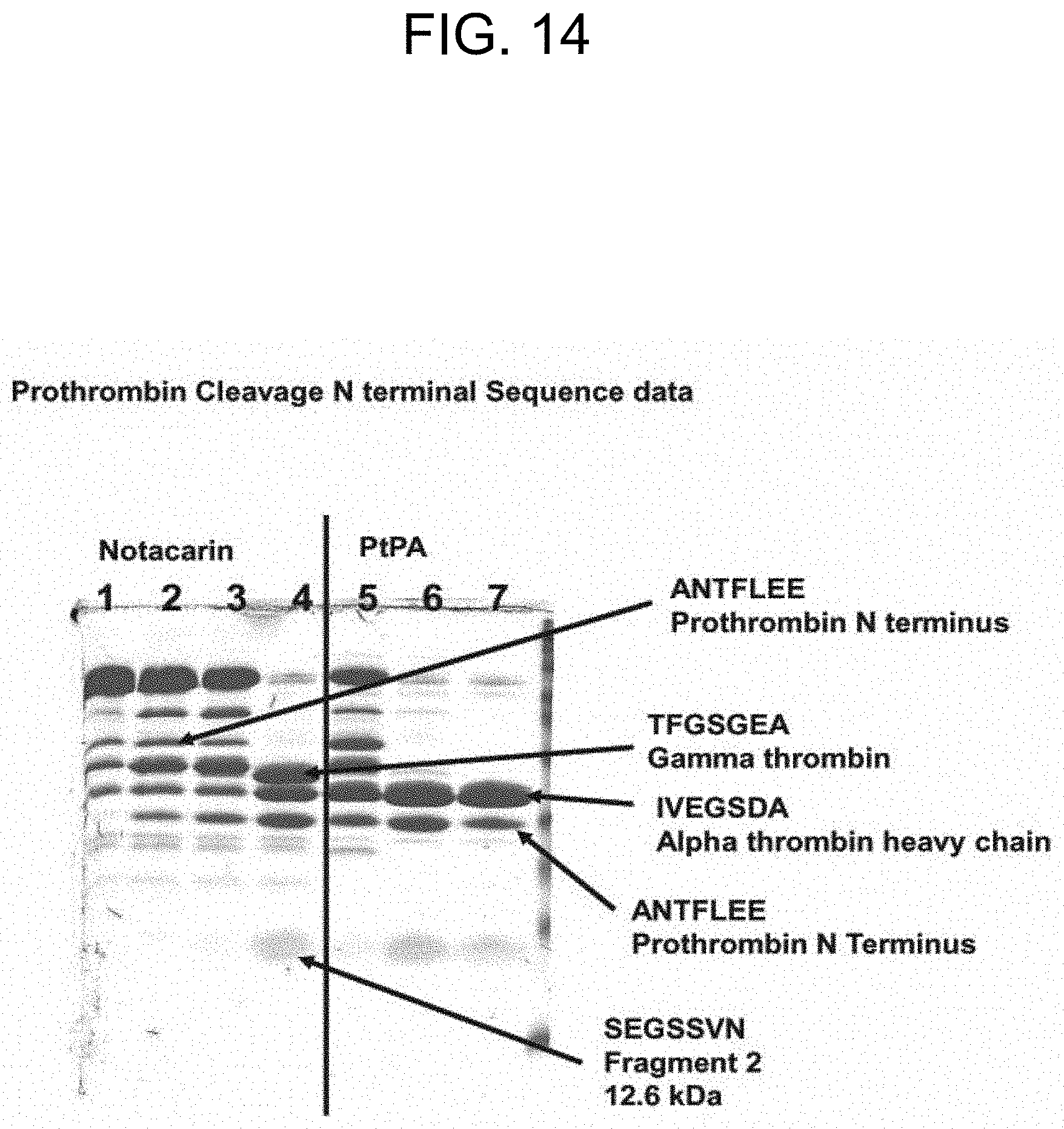
D00015

D00016
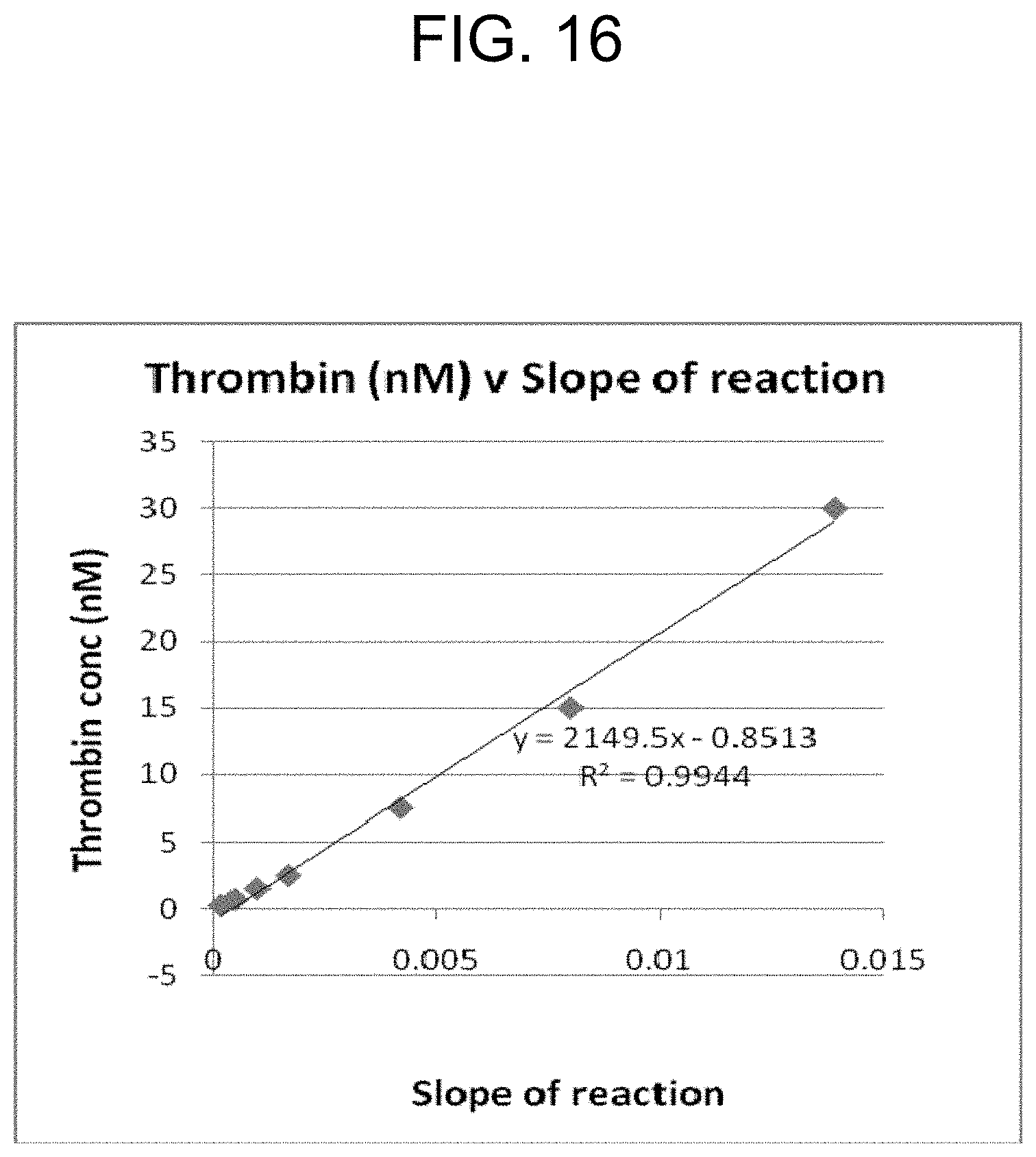
D00017
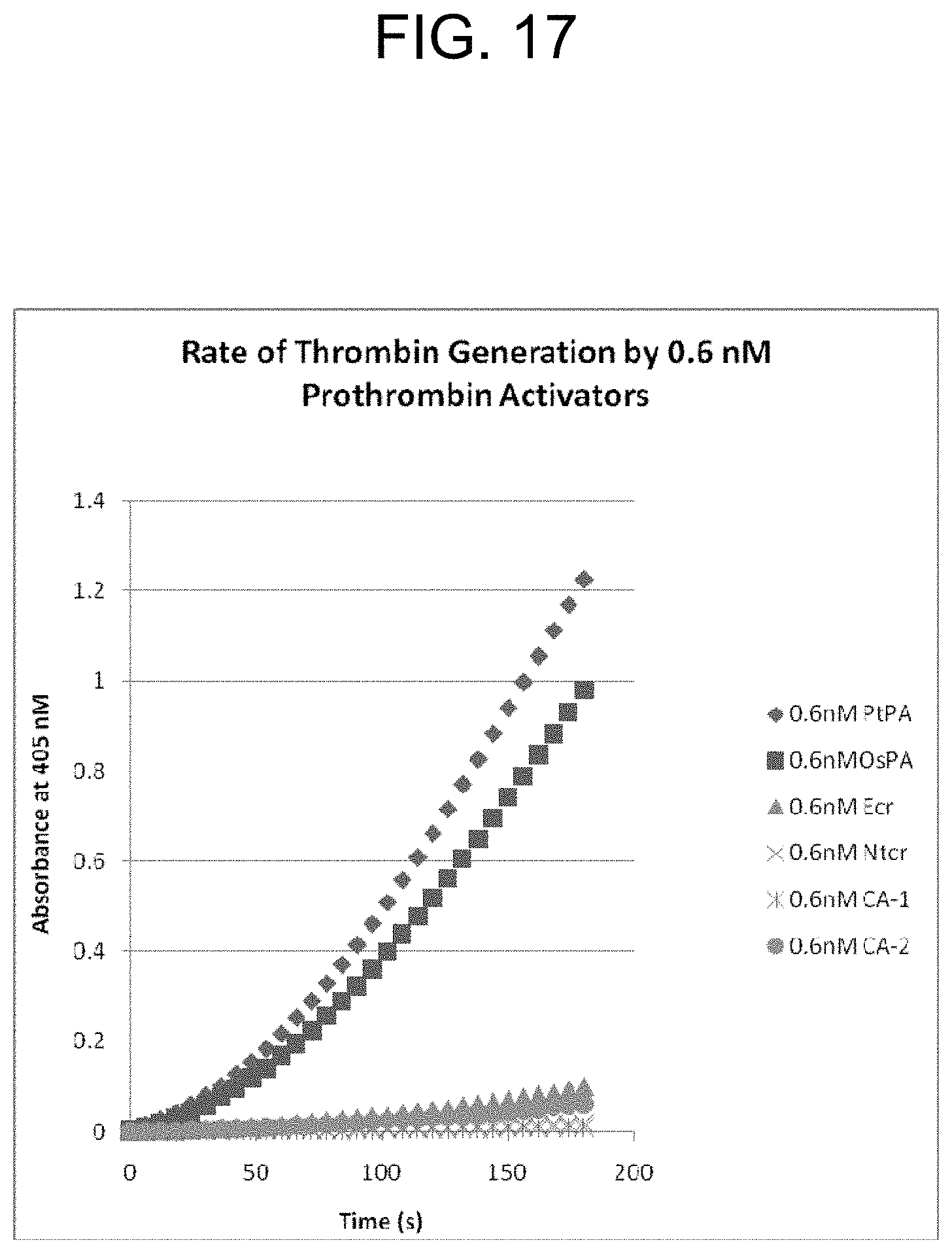
D00018
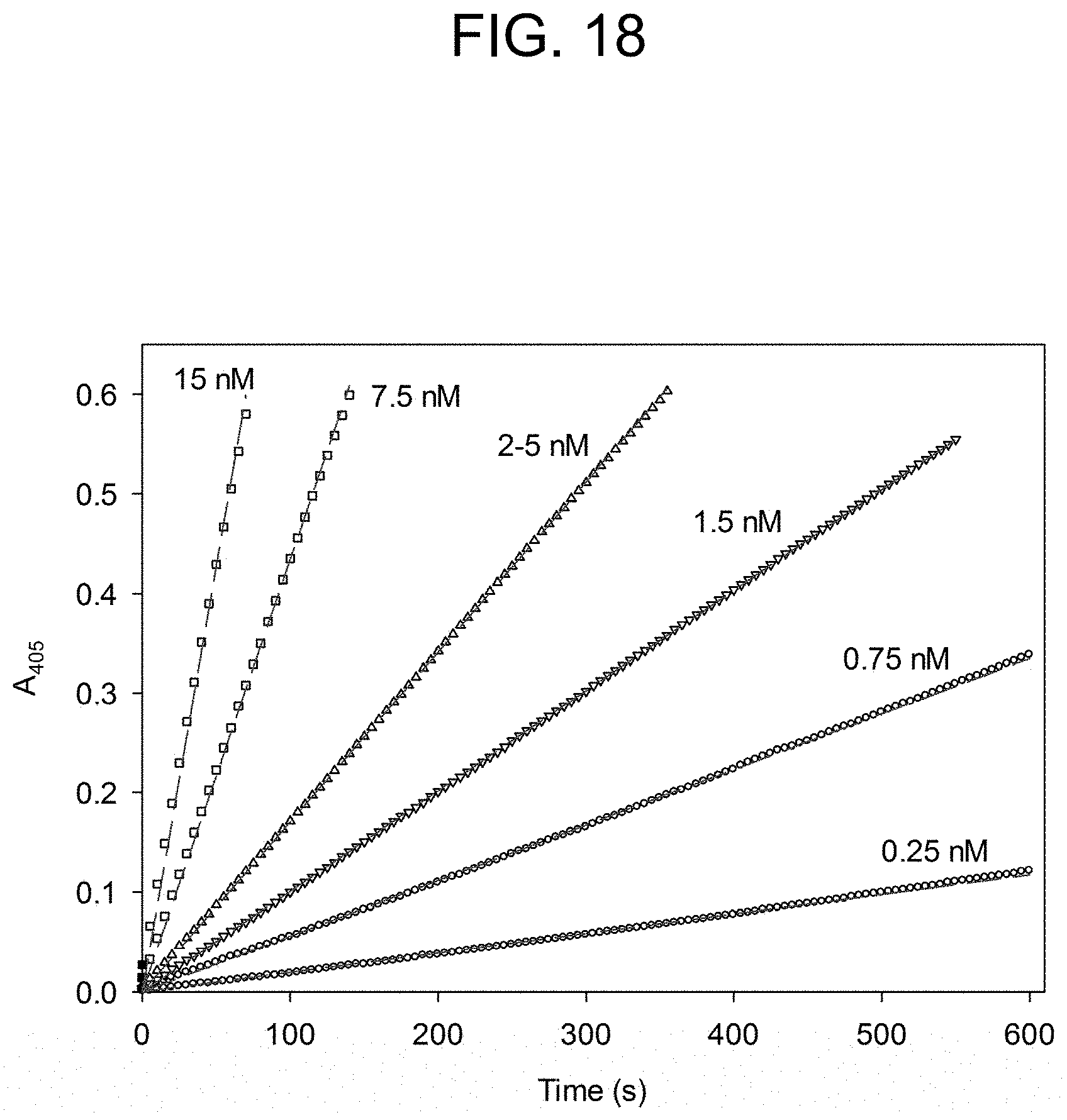
D00019
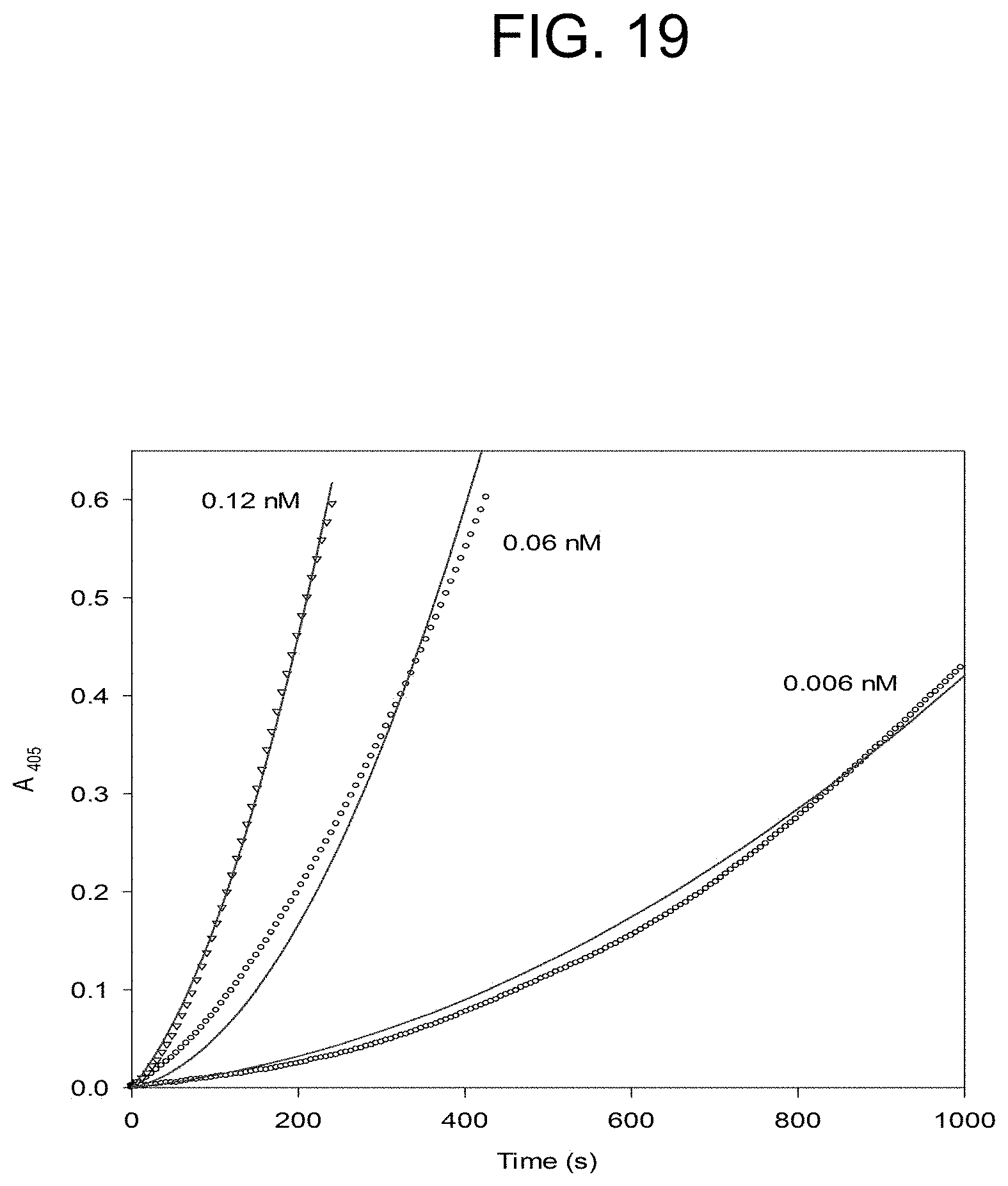
D00020
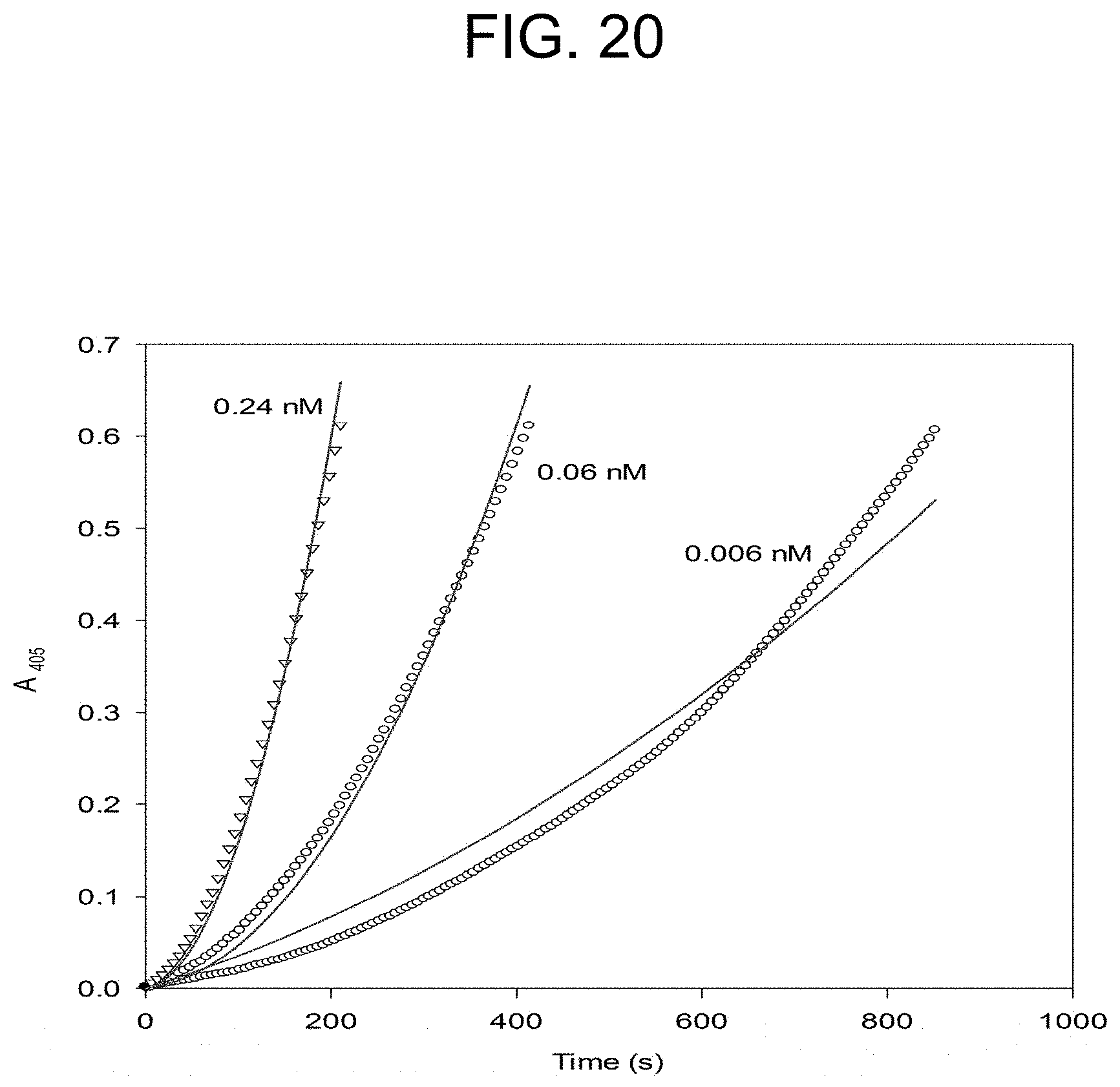
D00021
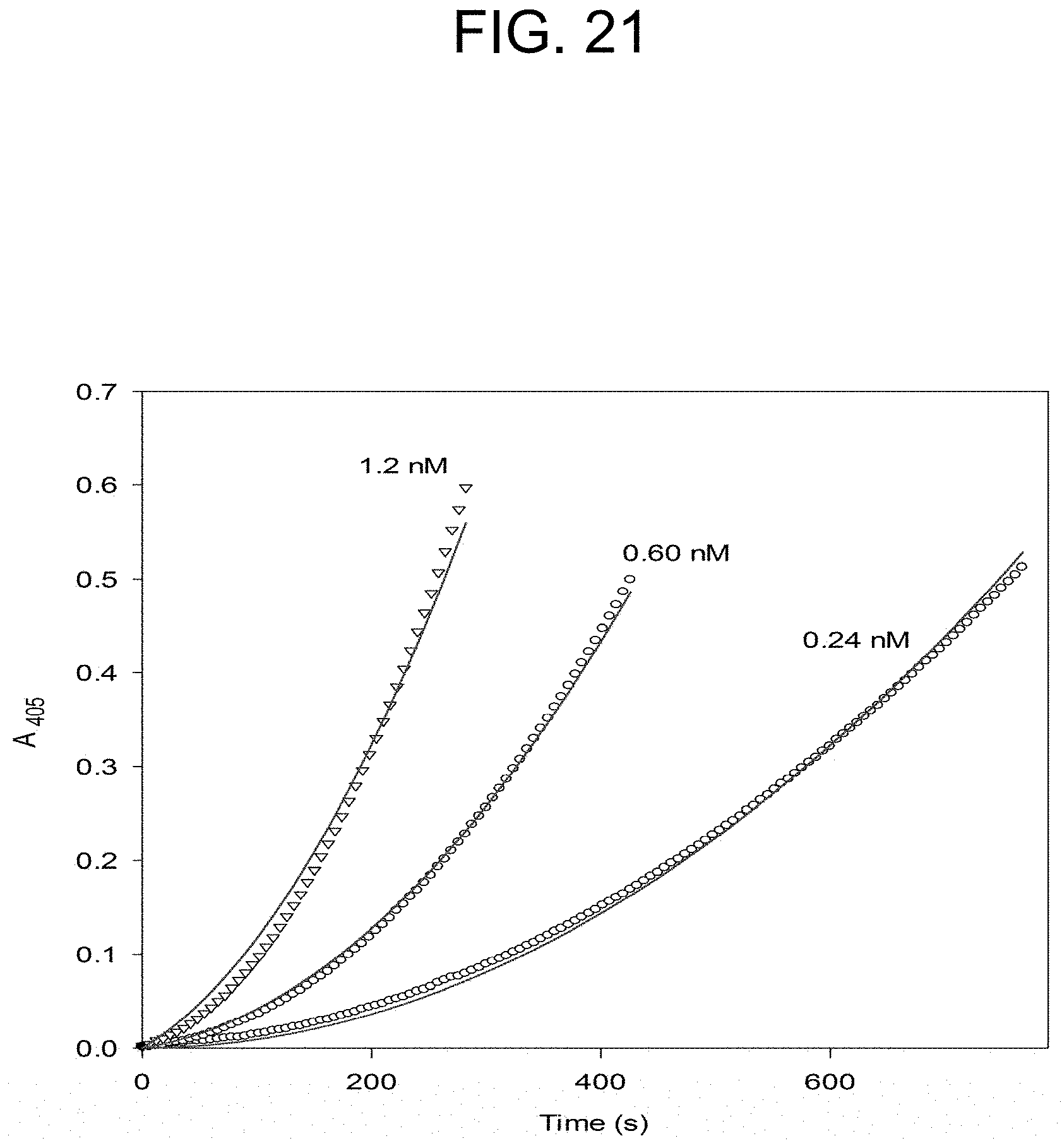
D00022
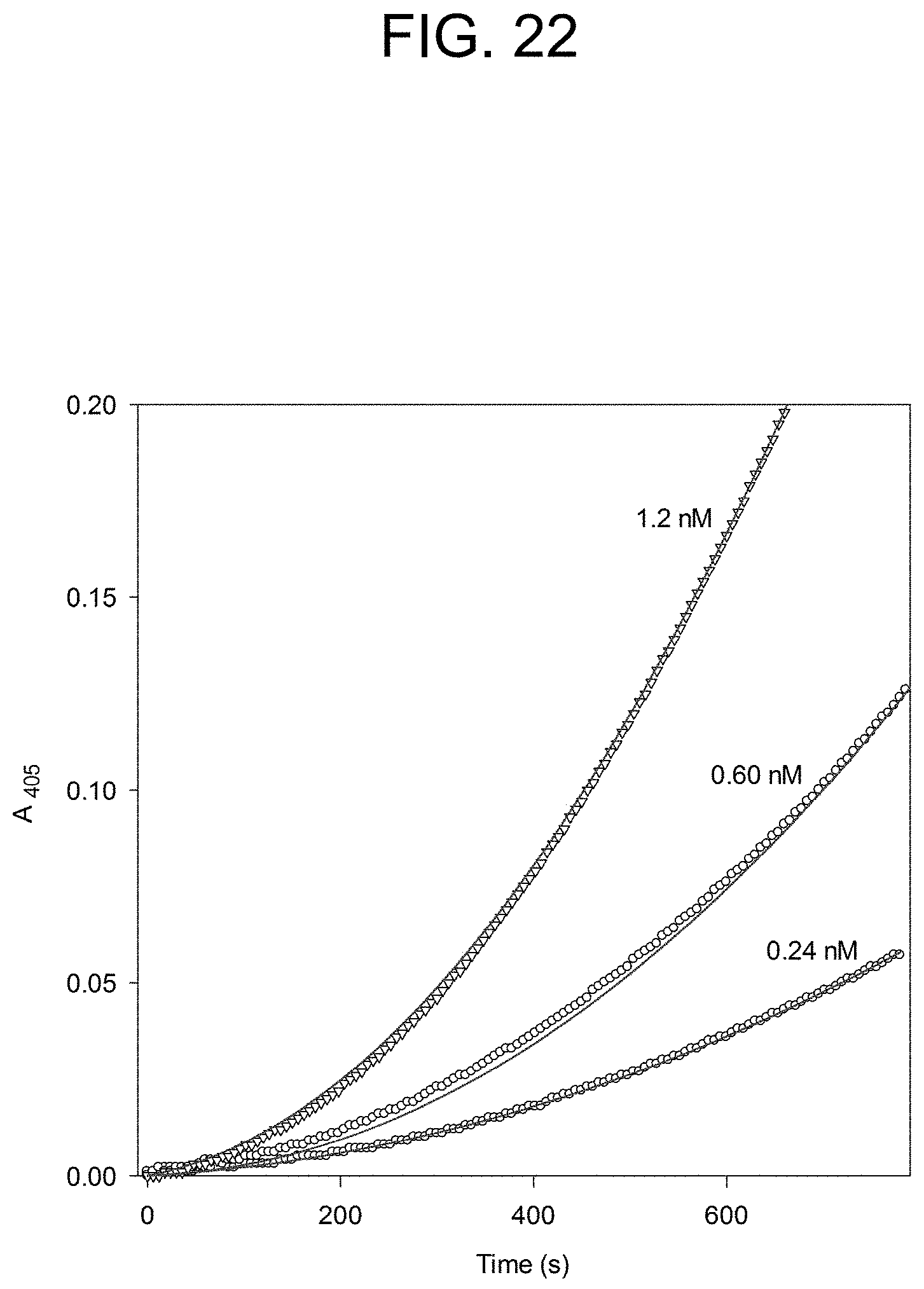
D00023
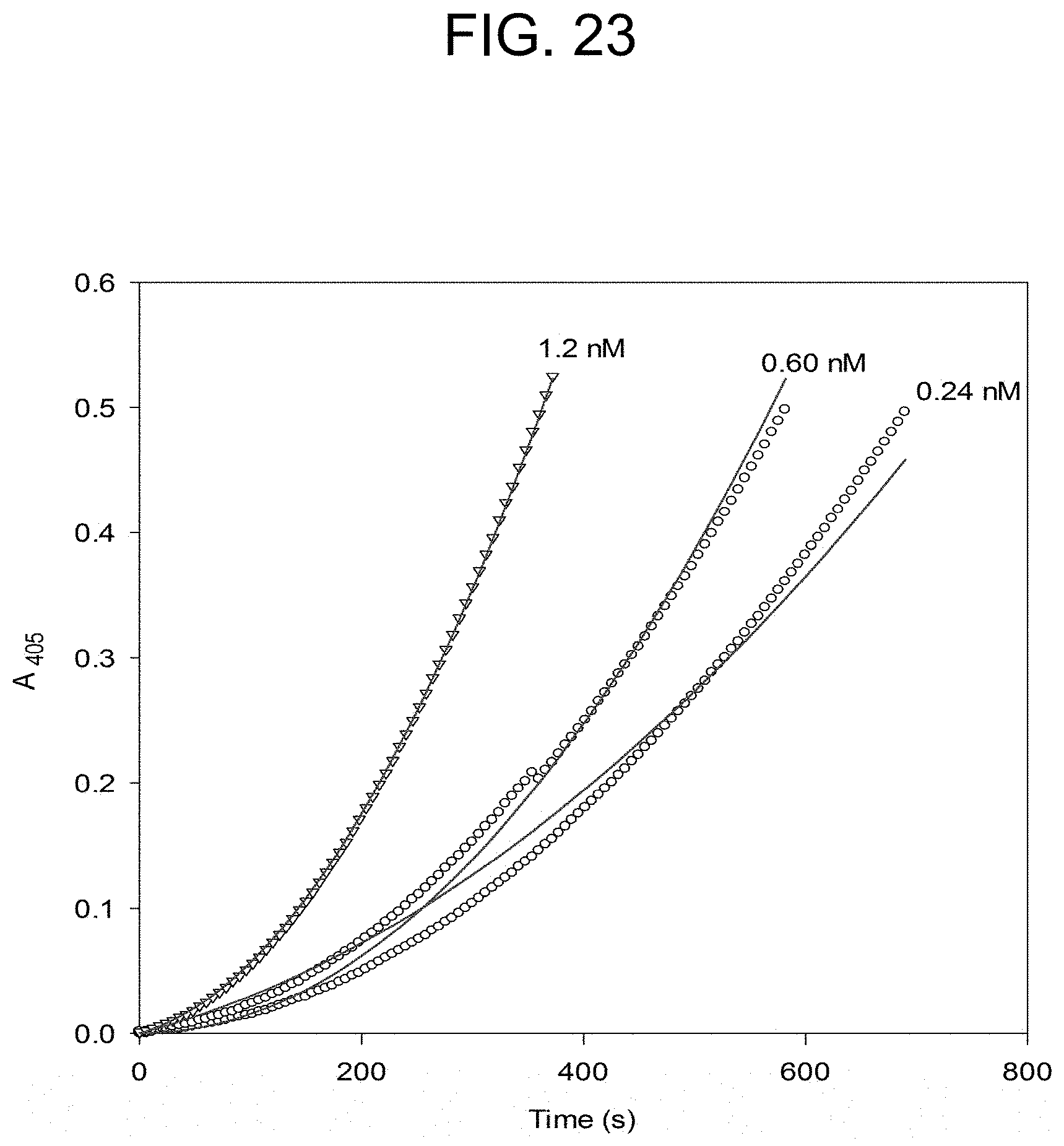
D00024
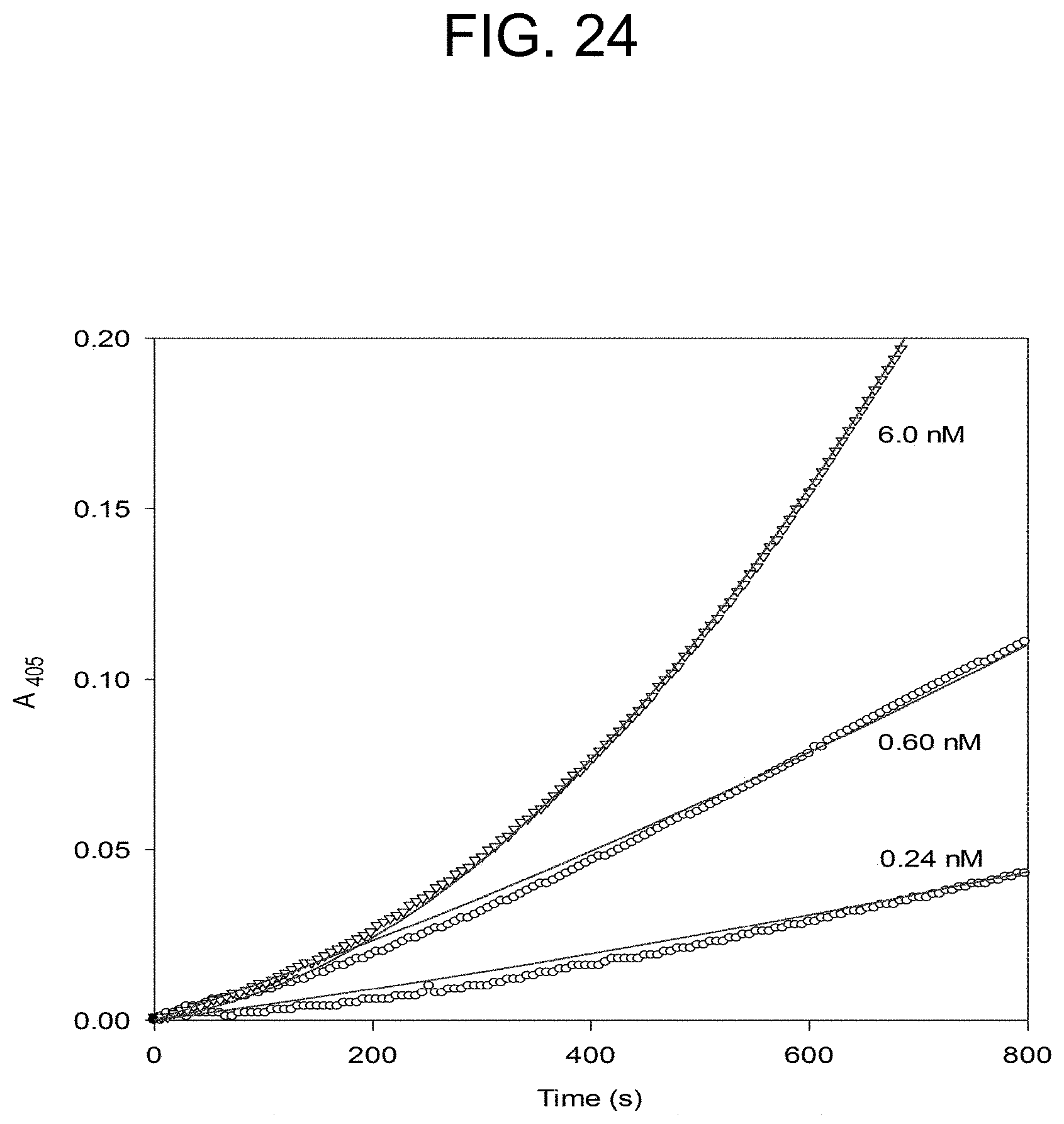
D00025
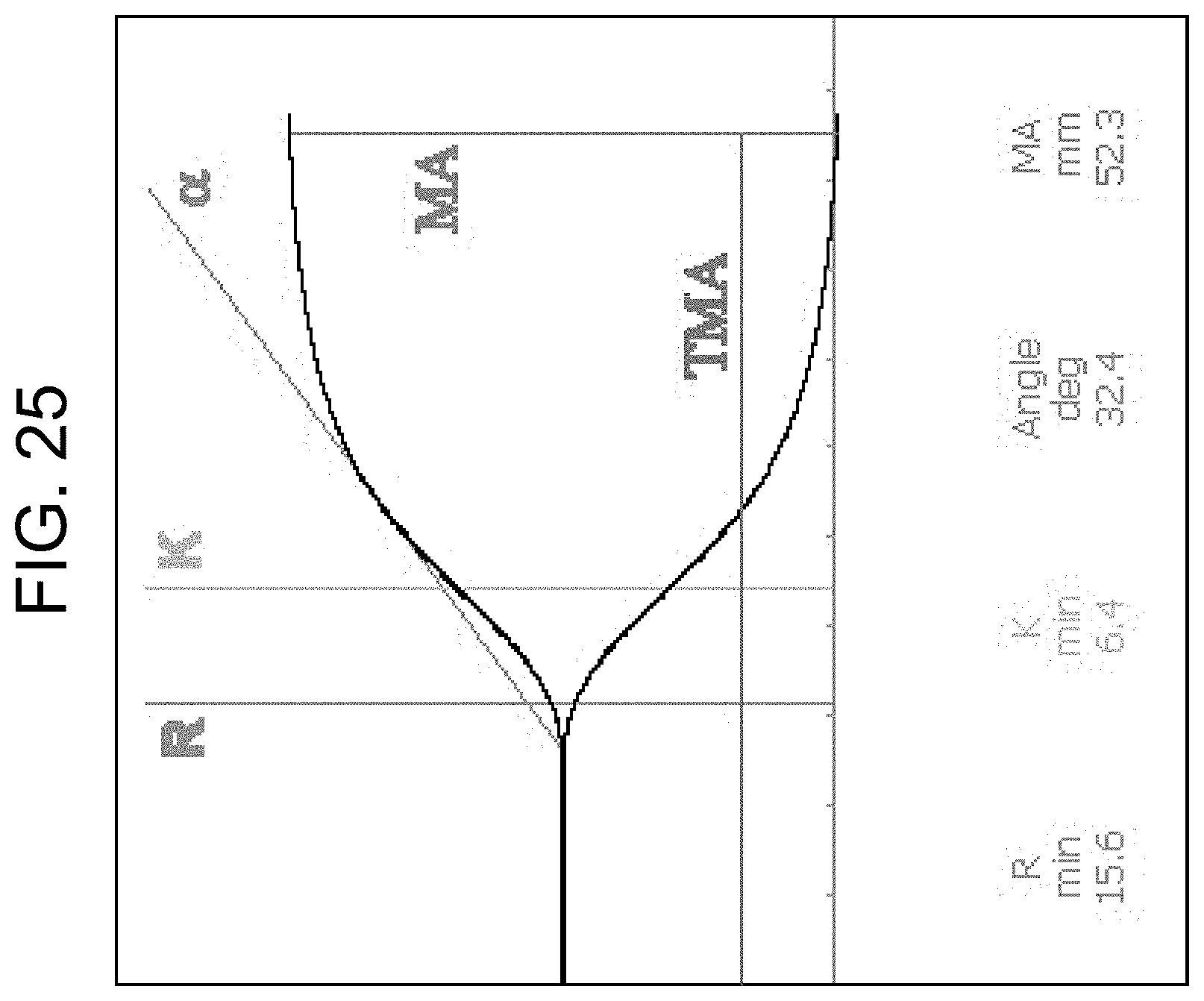
D00026
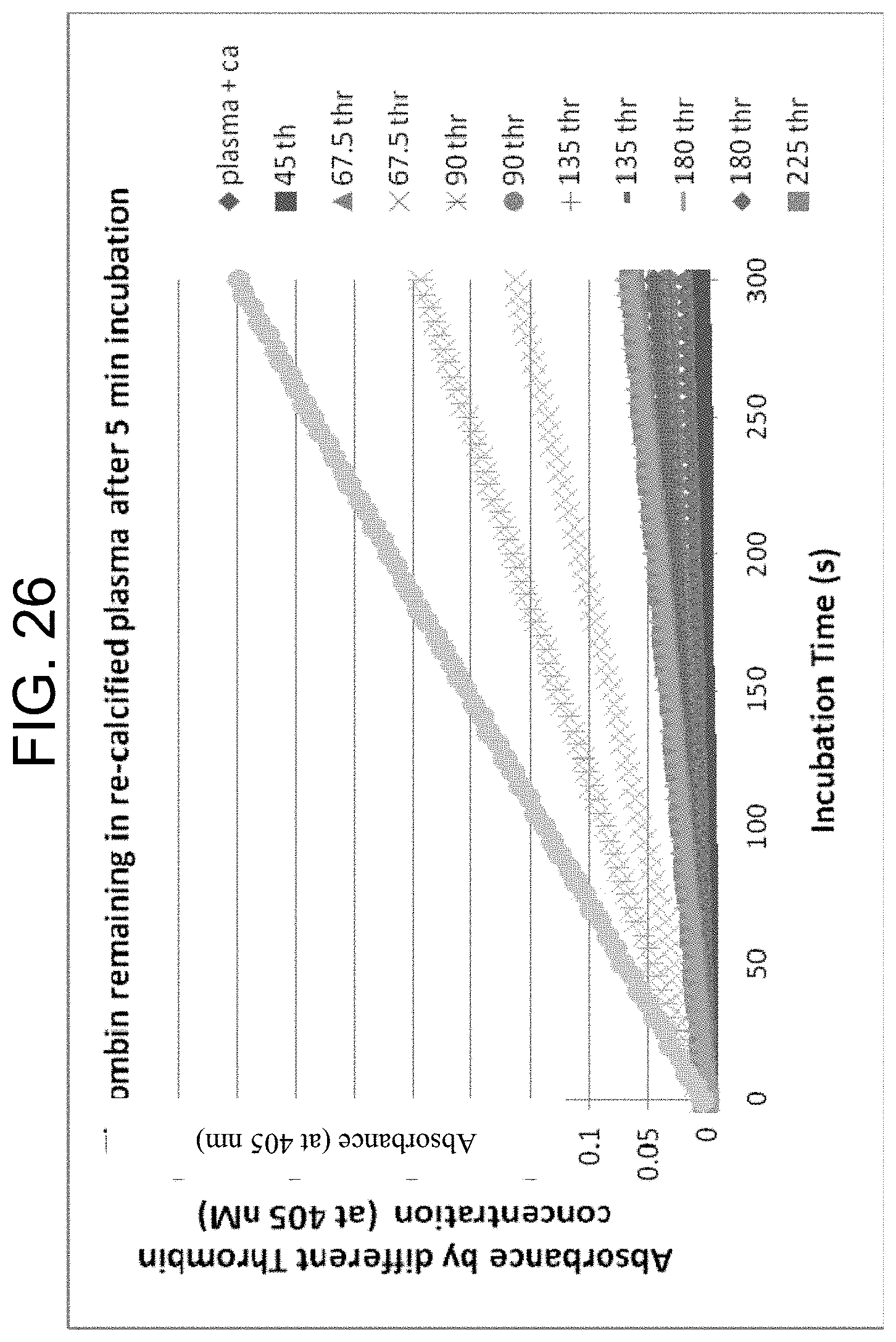
D00027

D00028
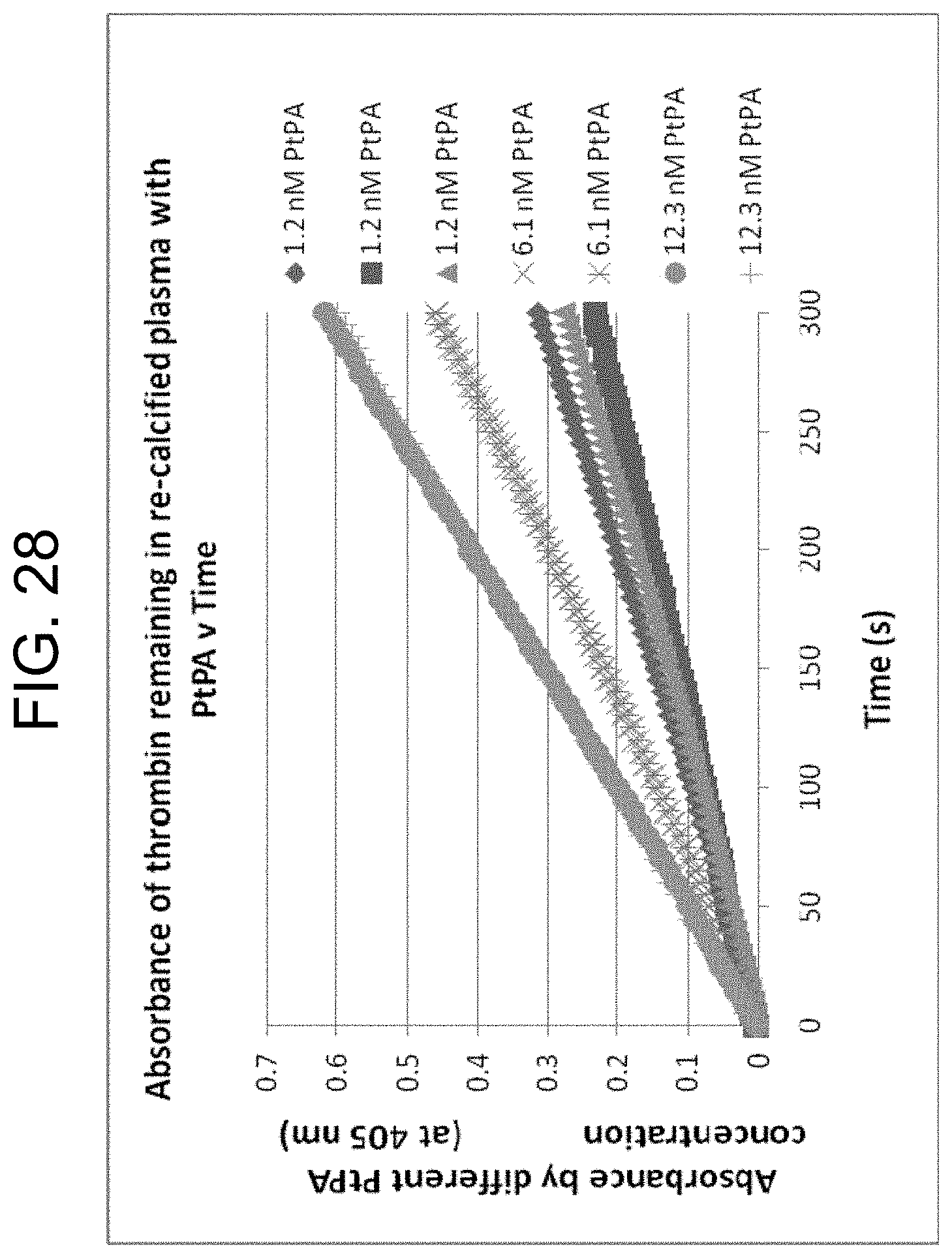
D00029
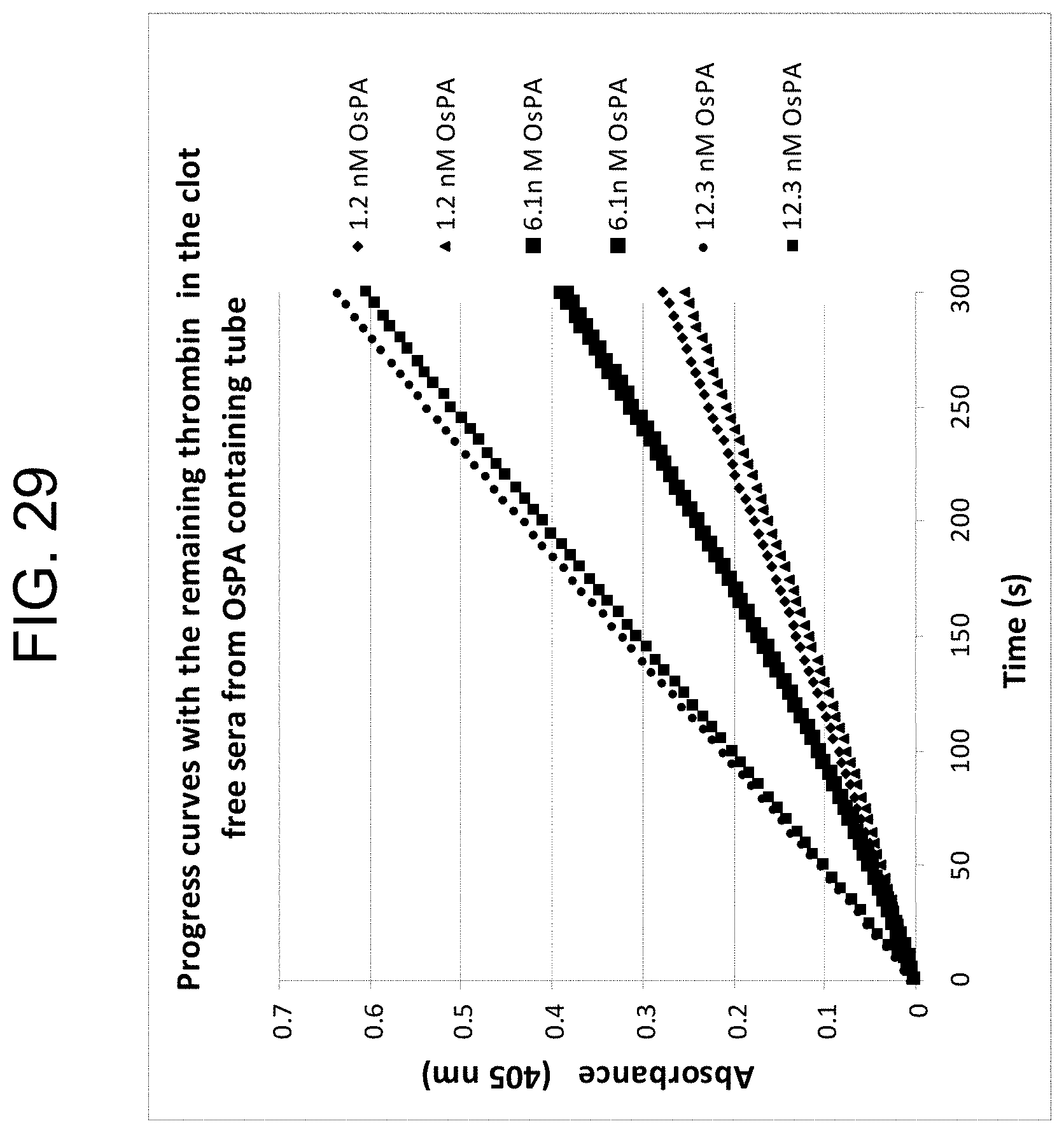
D00030
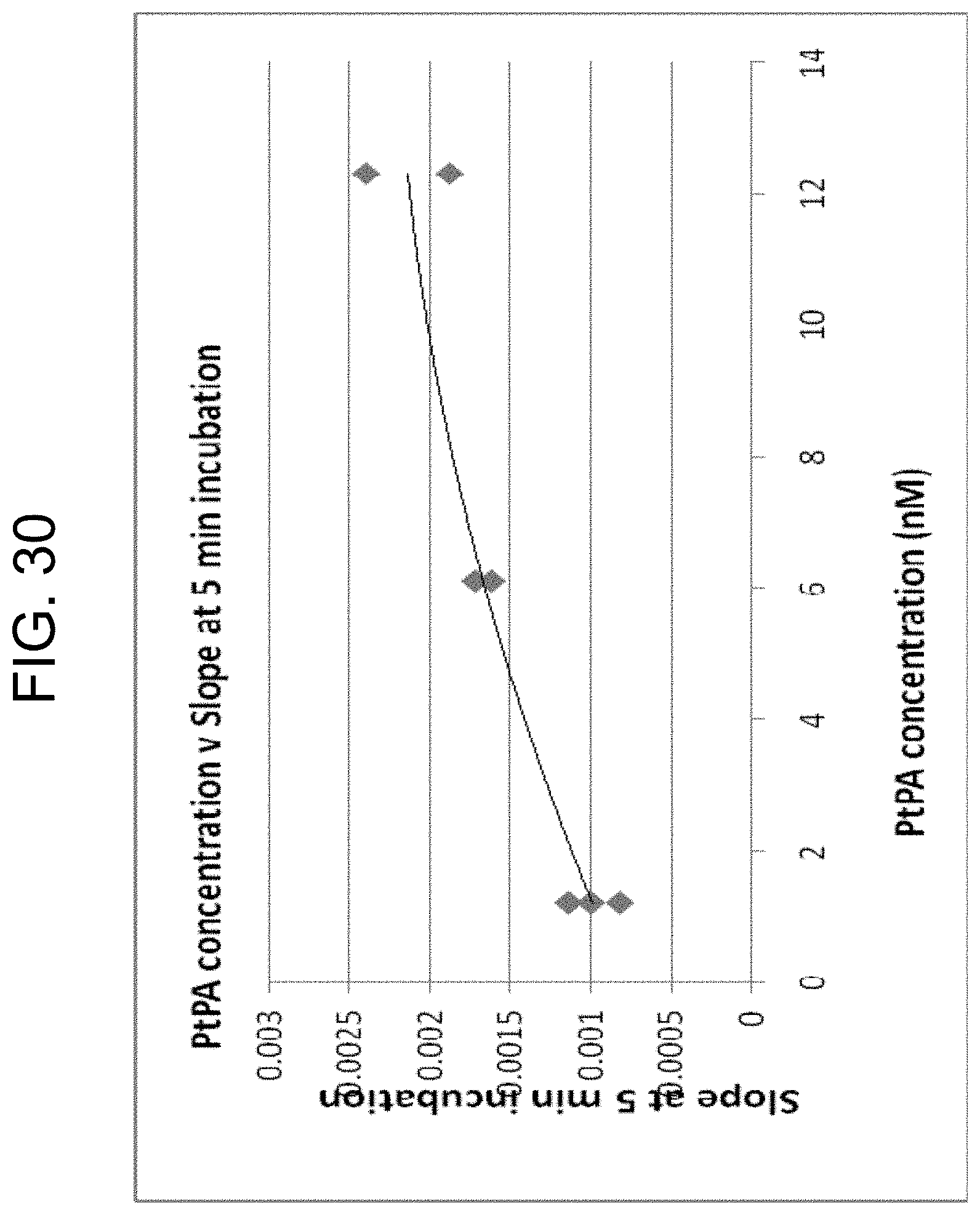
D00031
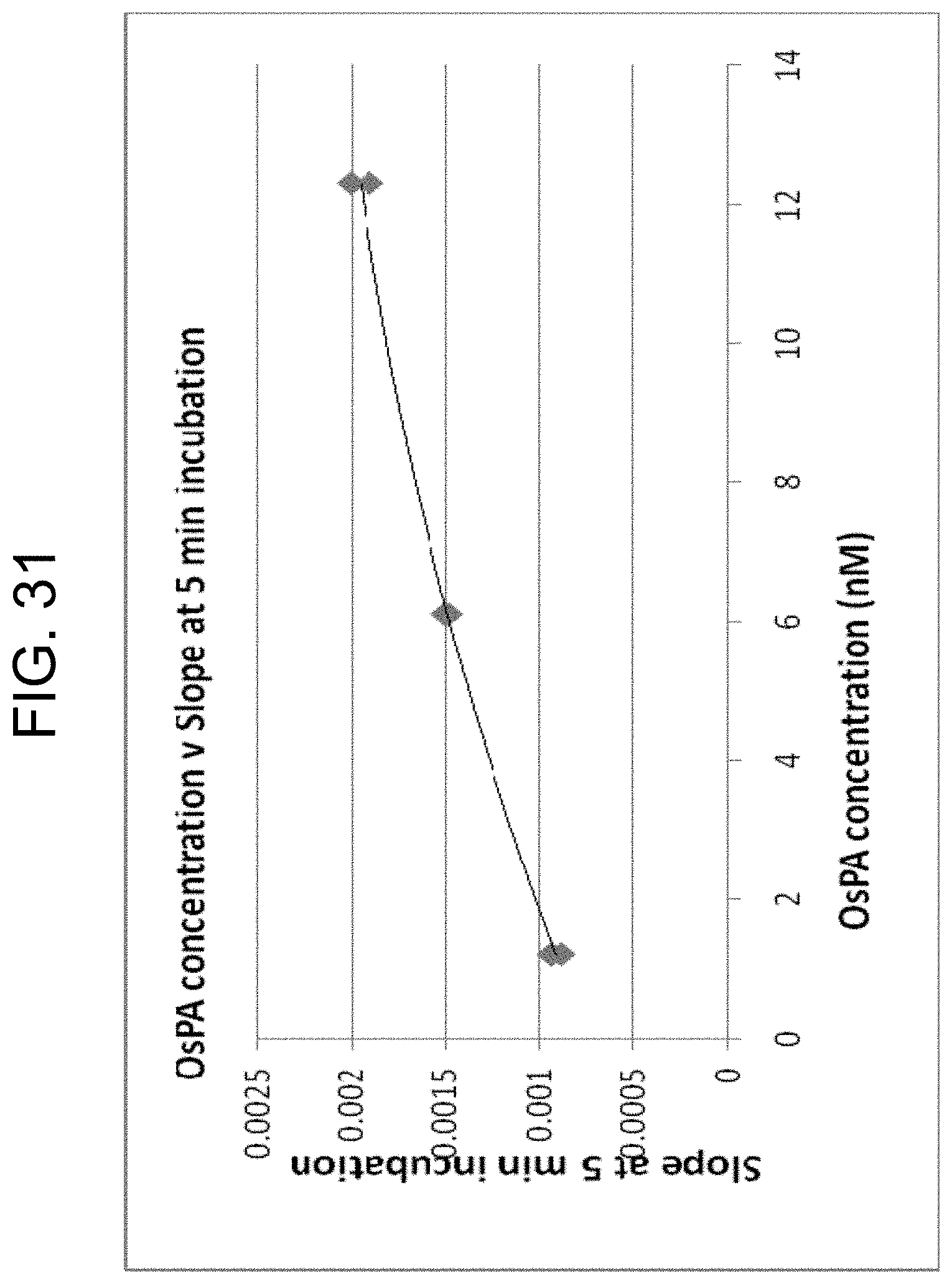
D00032
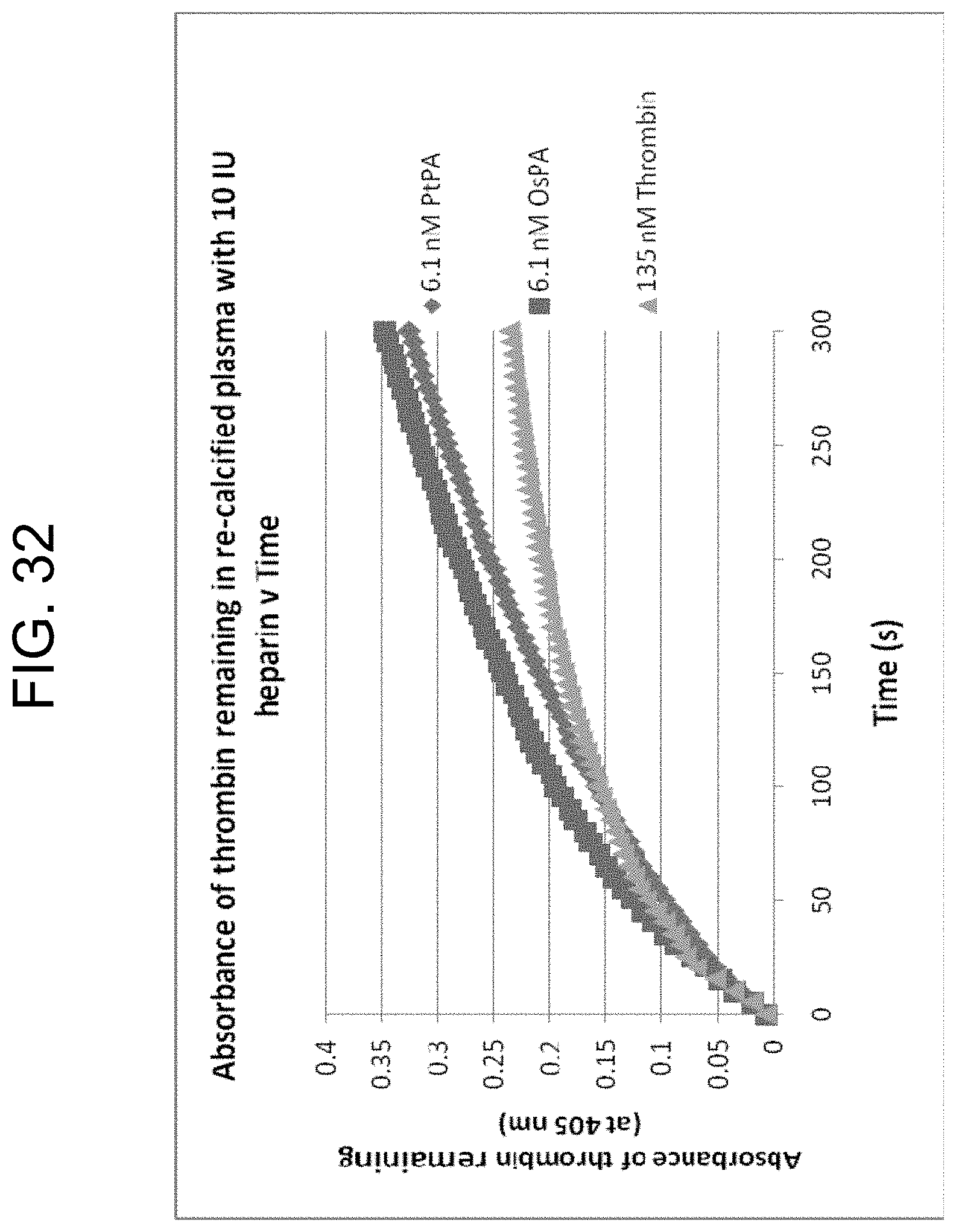
D00033
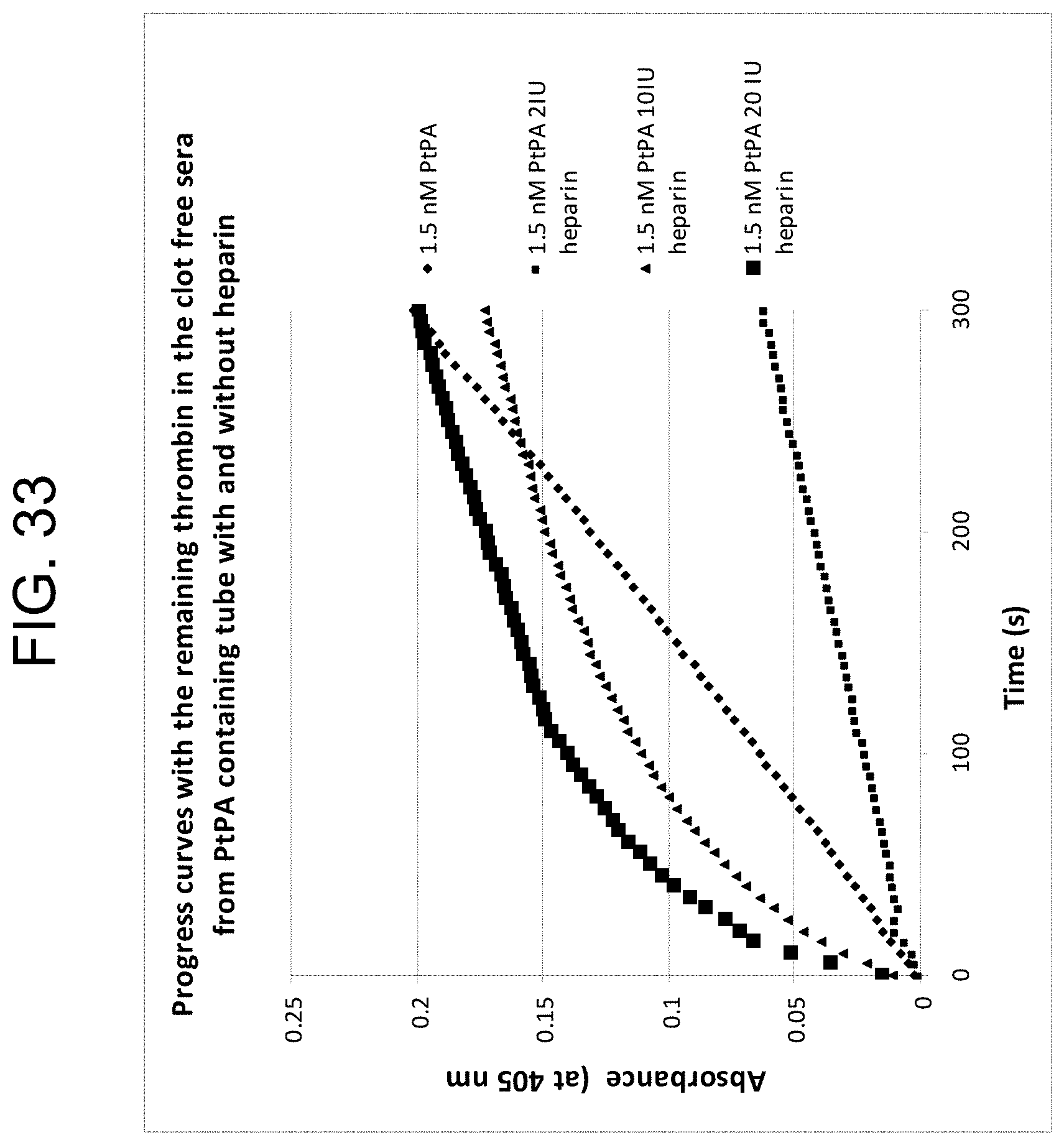
D00034
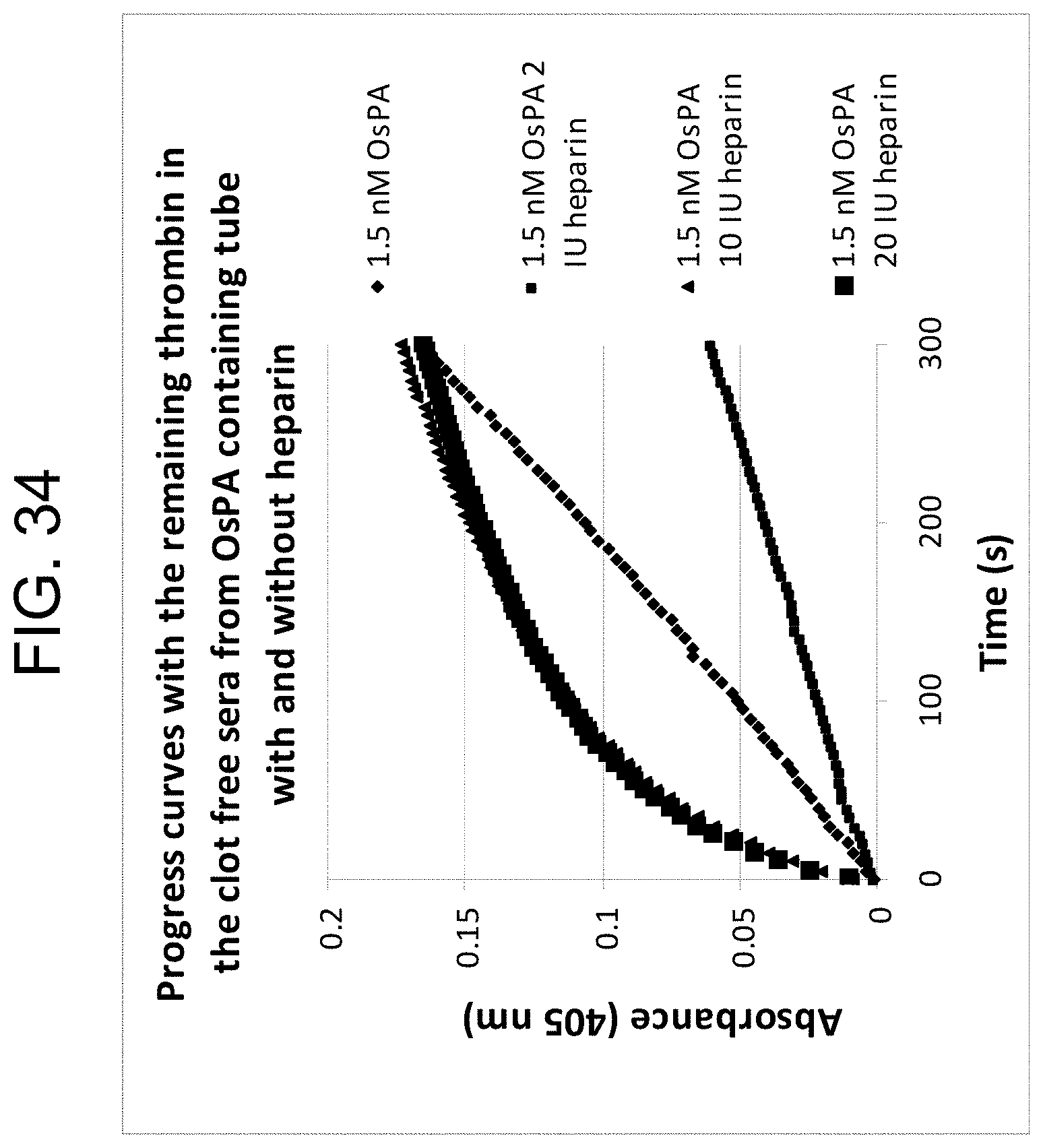
D00035
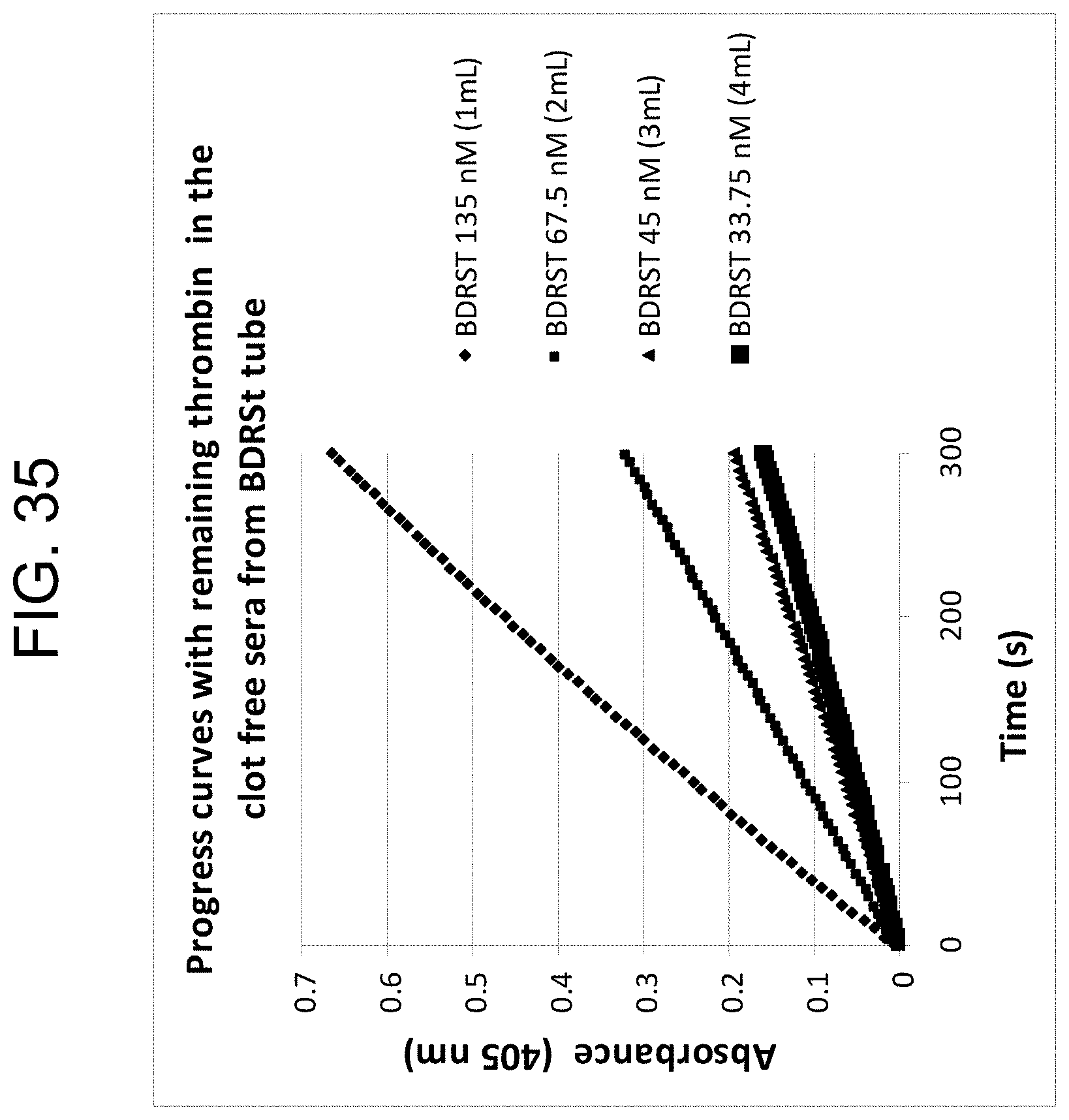
D00036
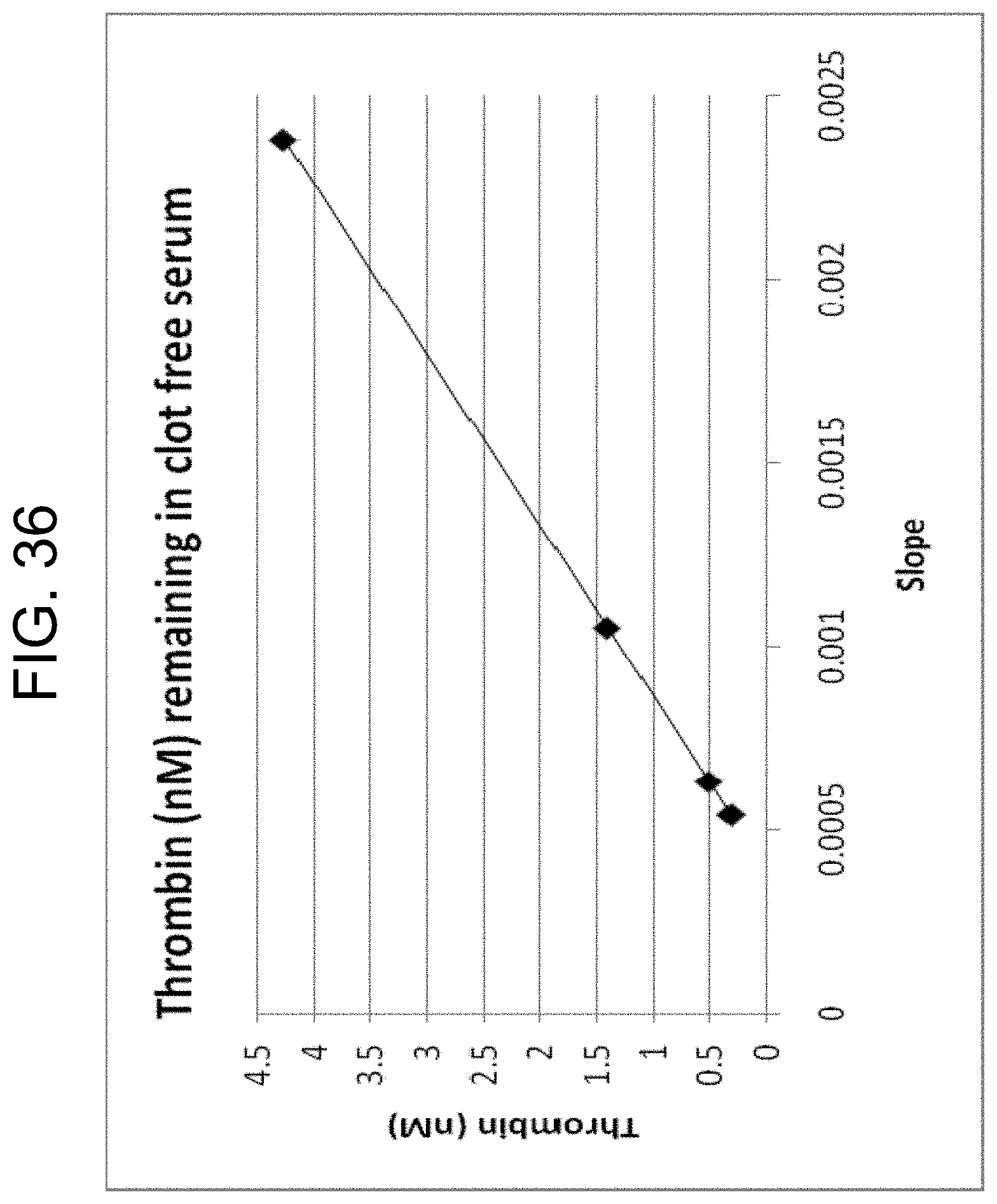
D00037
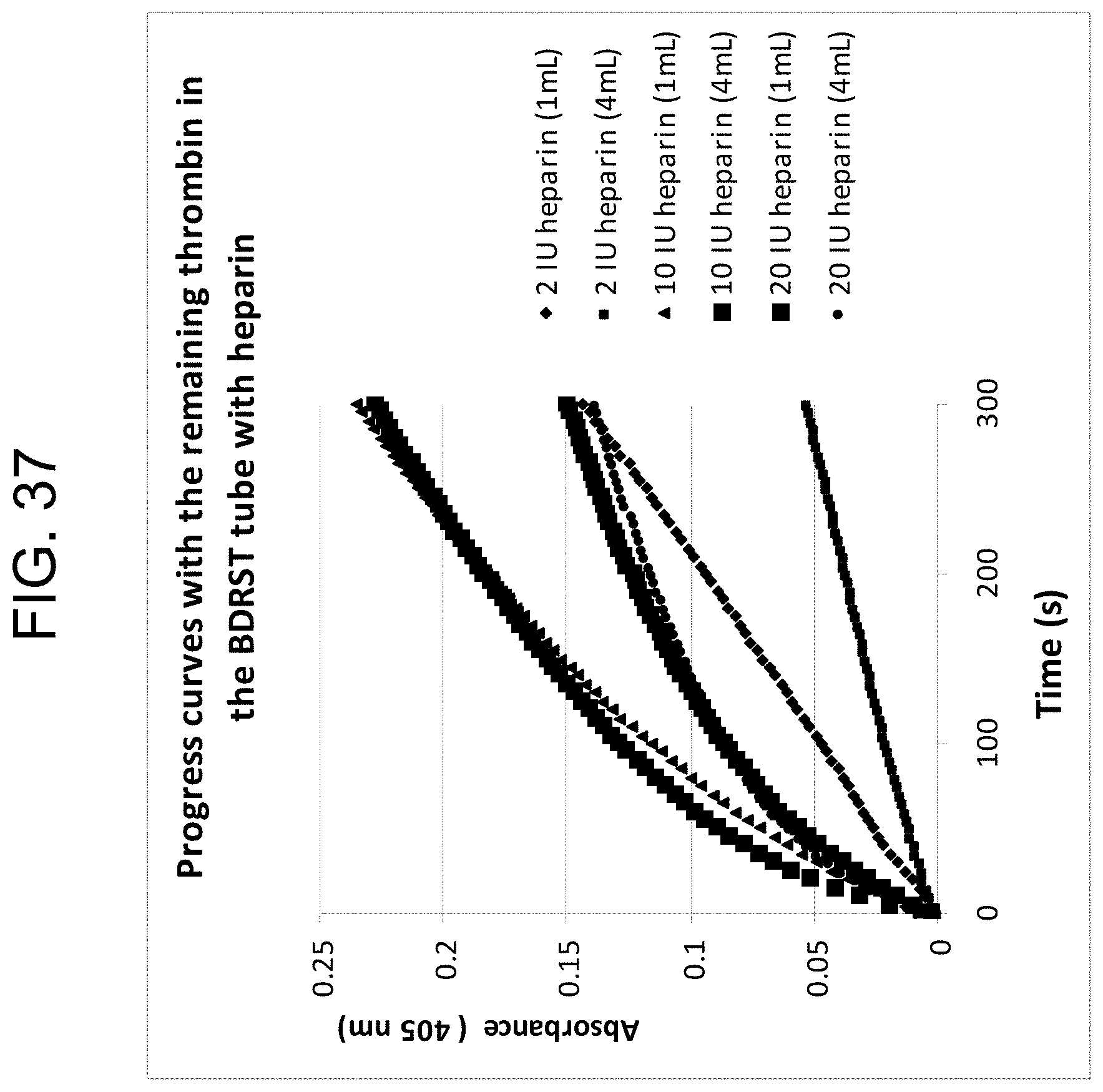
D00038
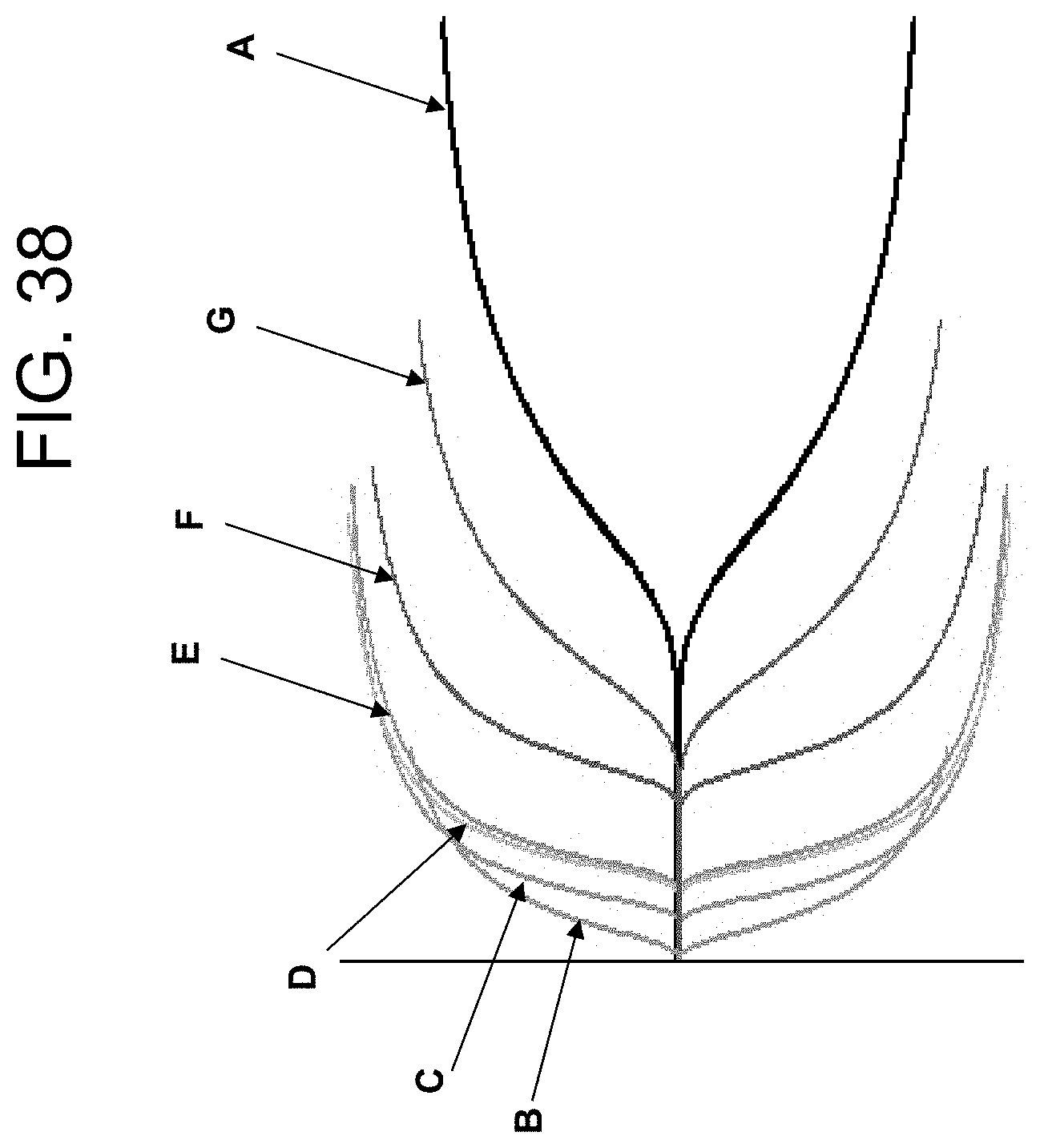
D00039
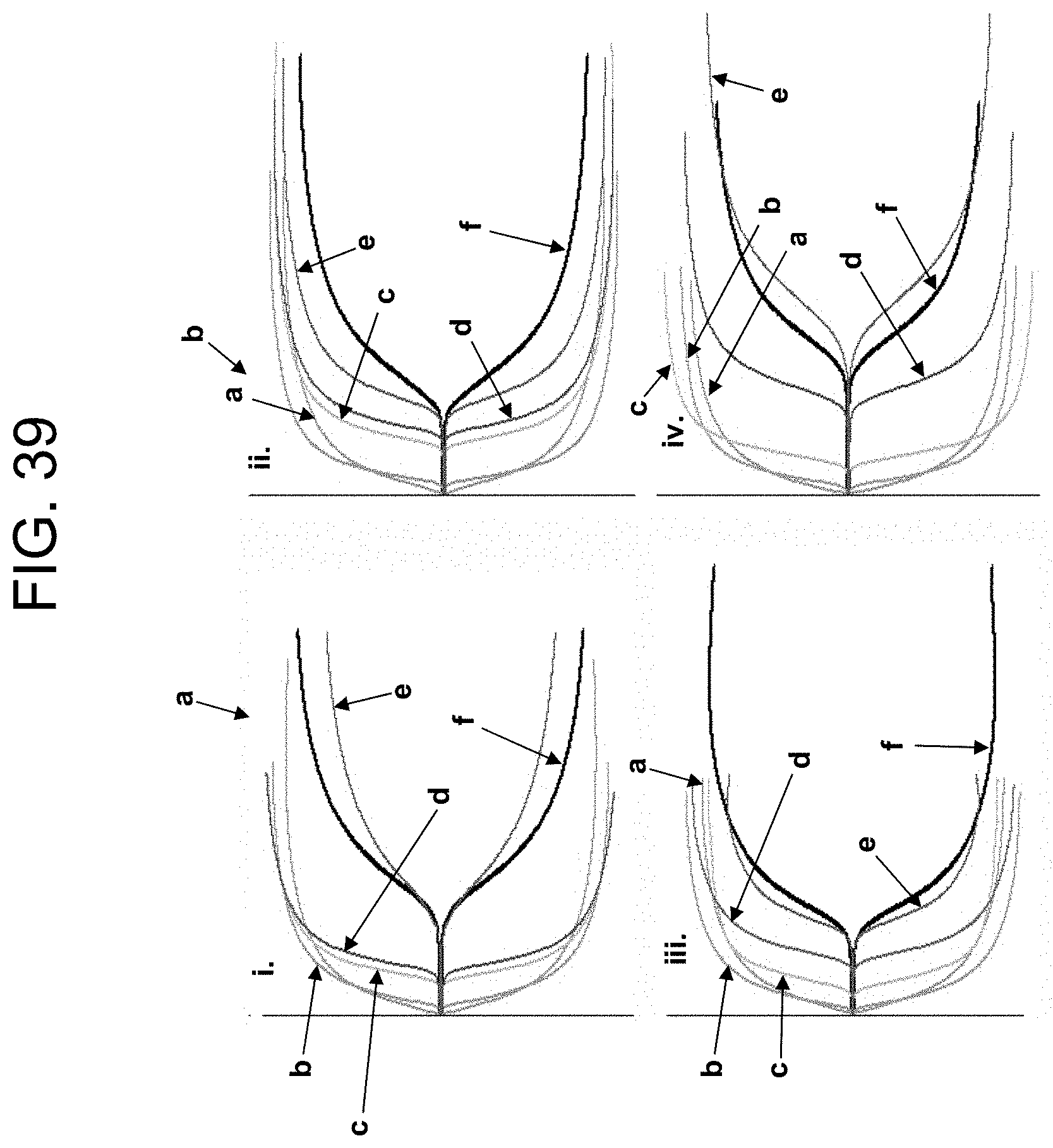
D00040
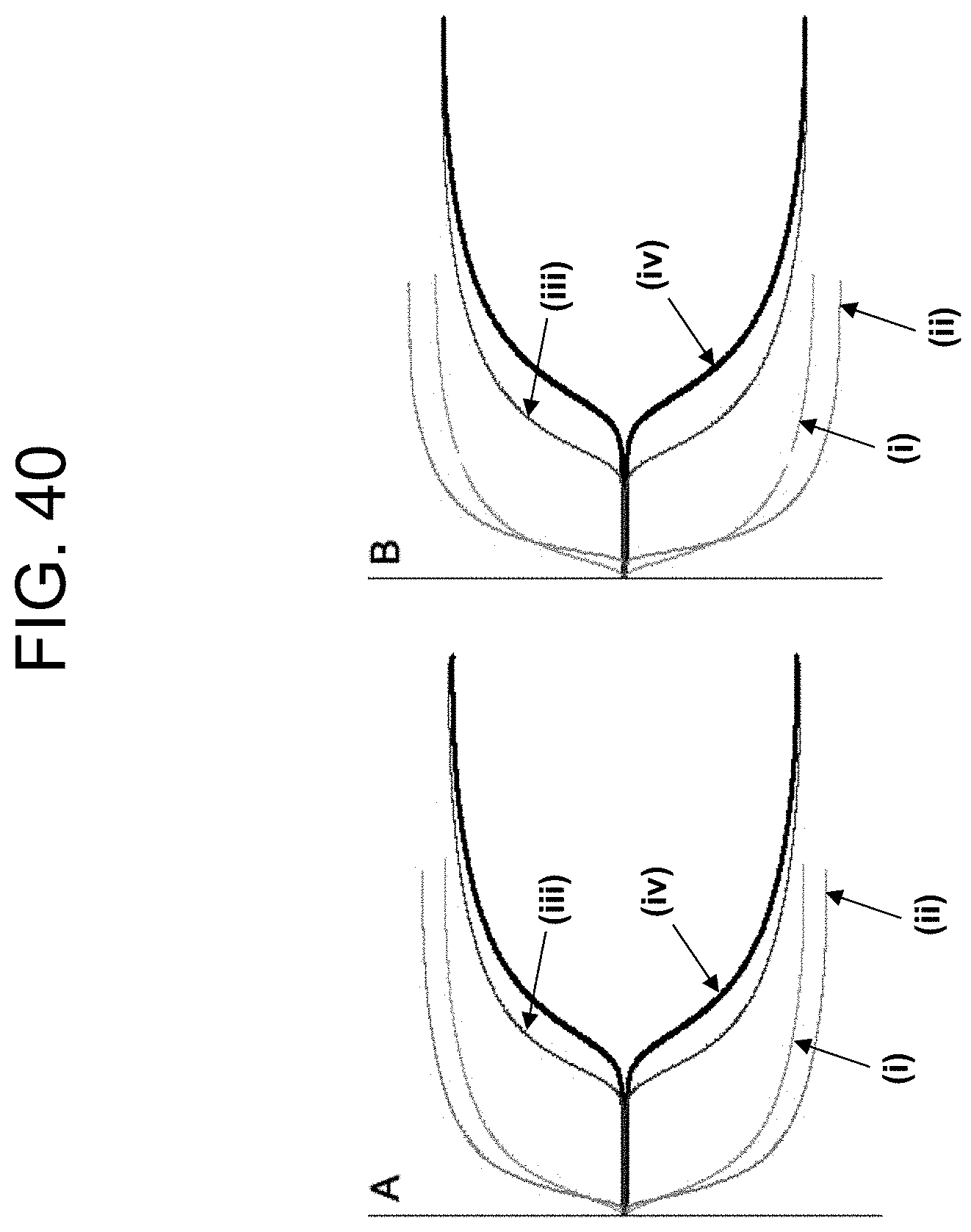
D00041
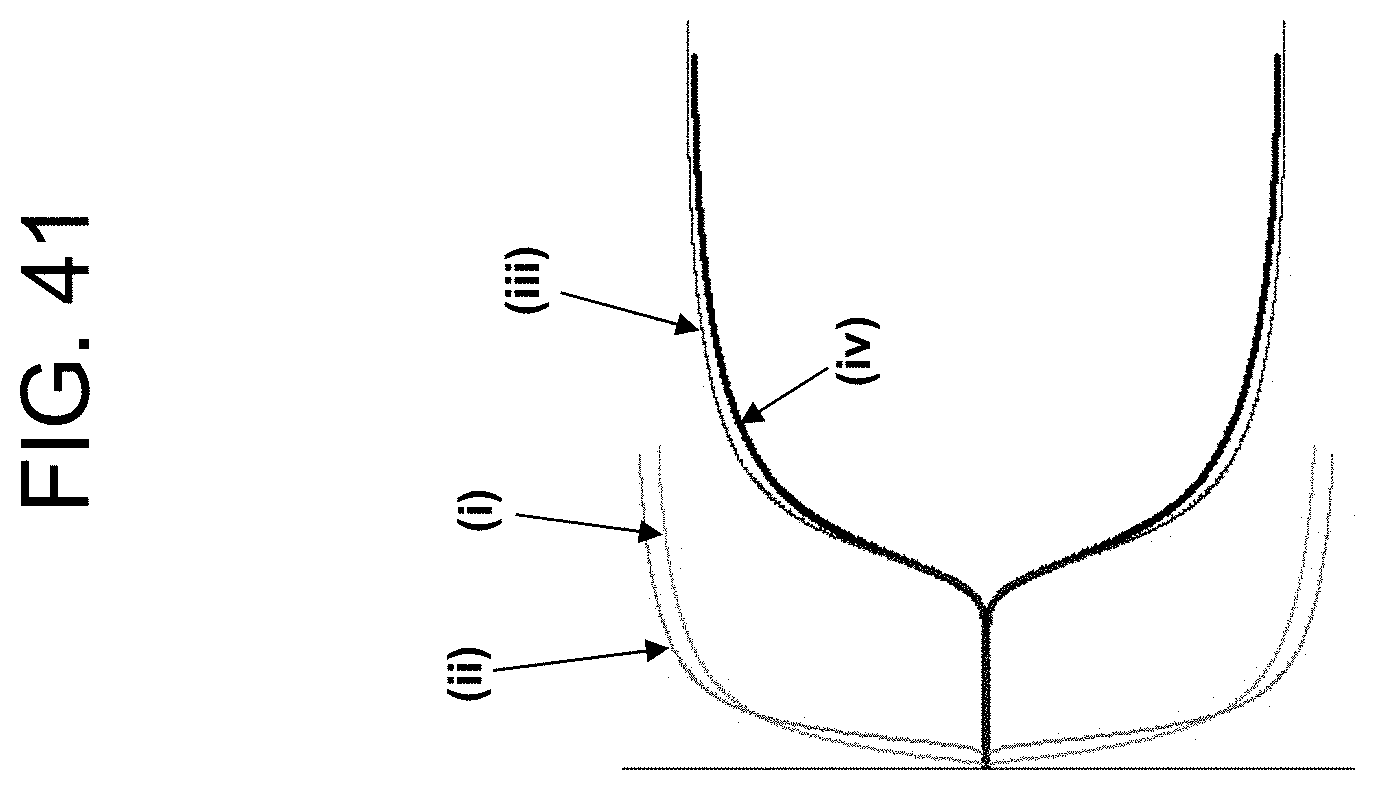
D00042
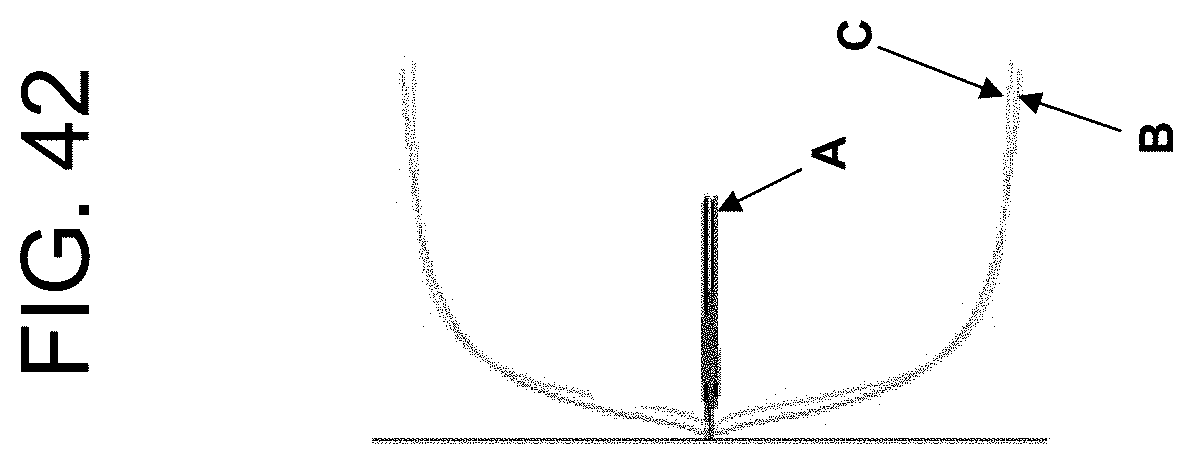
D00043
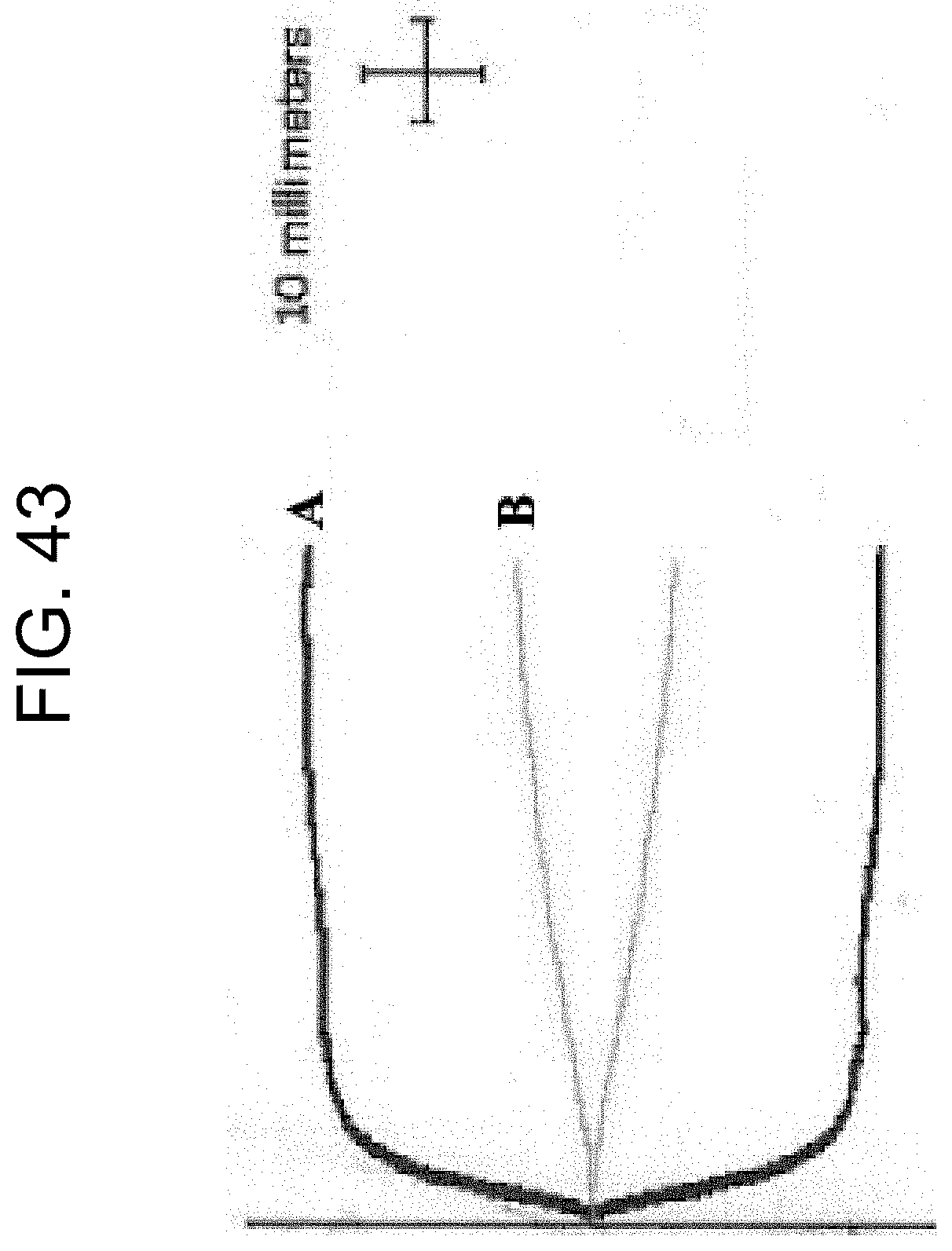
D00044
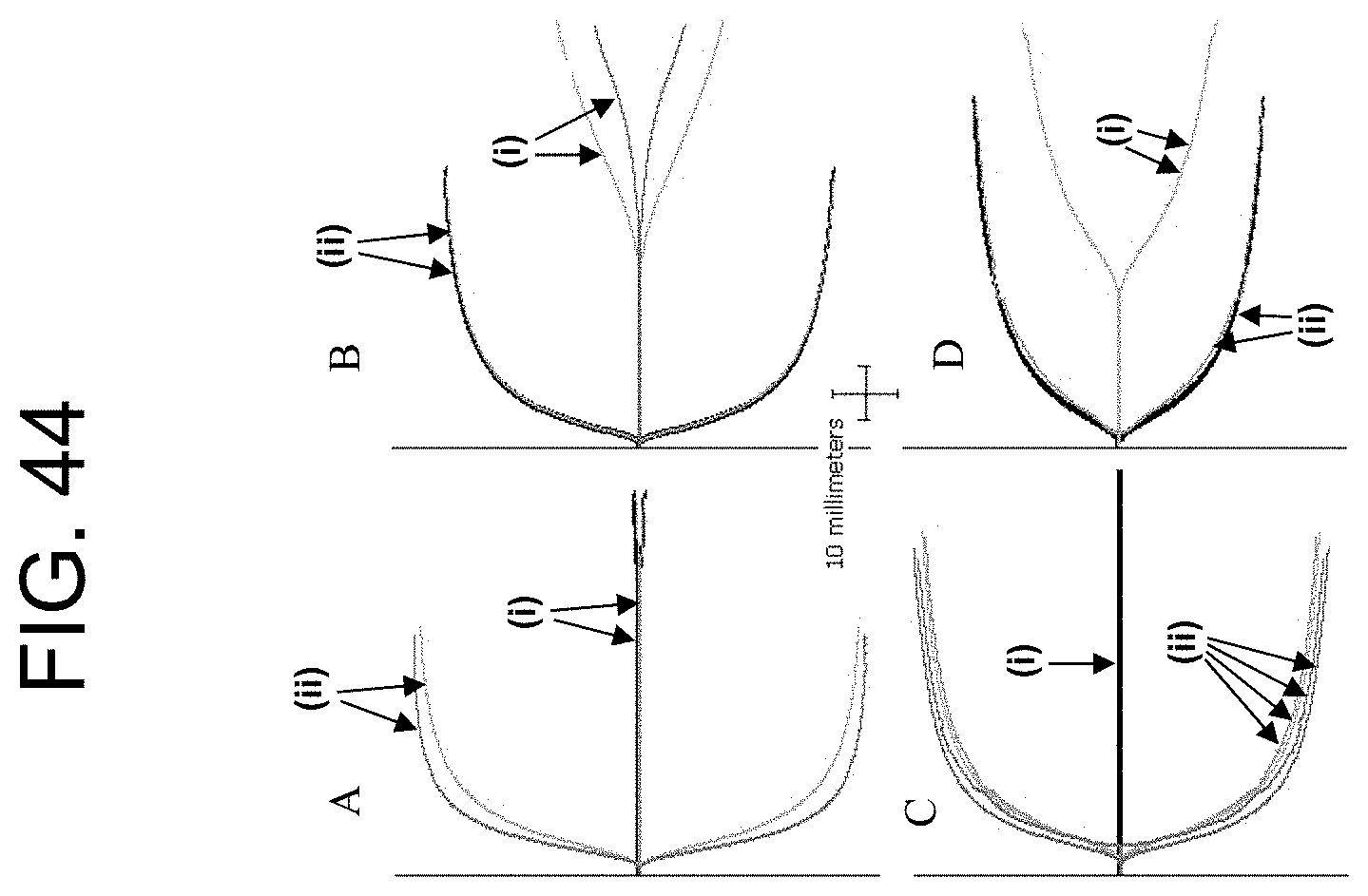
D00045
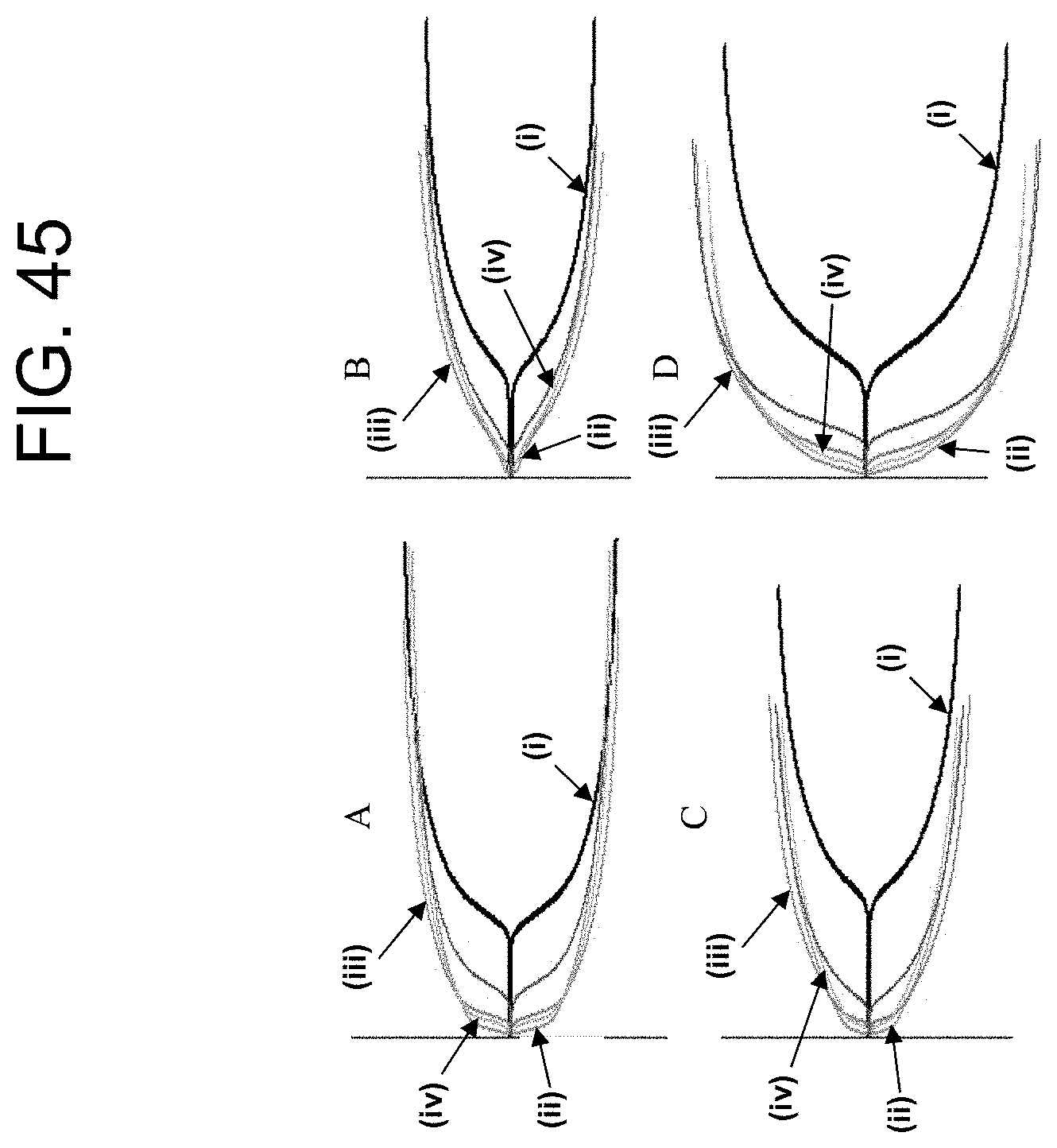
D00046
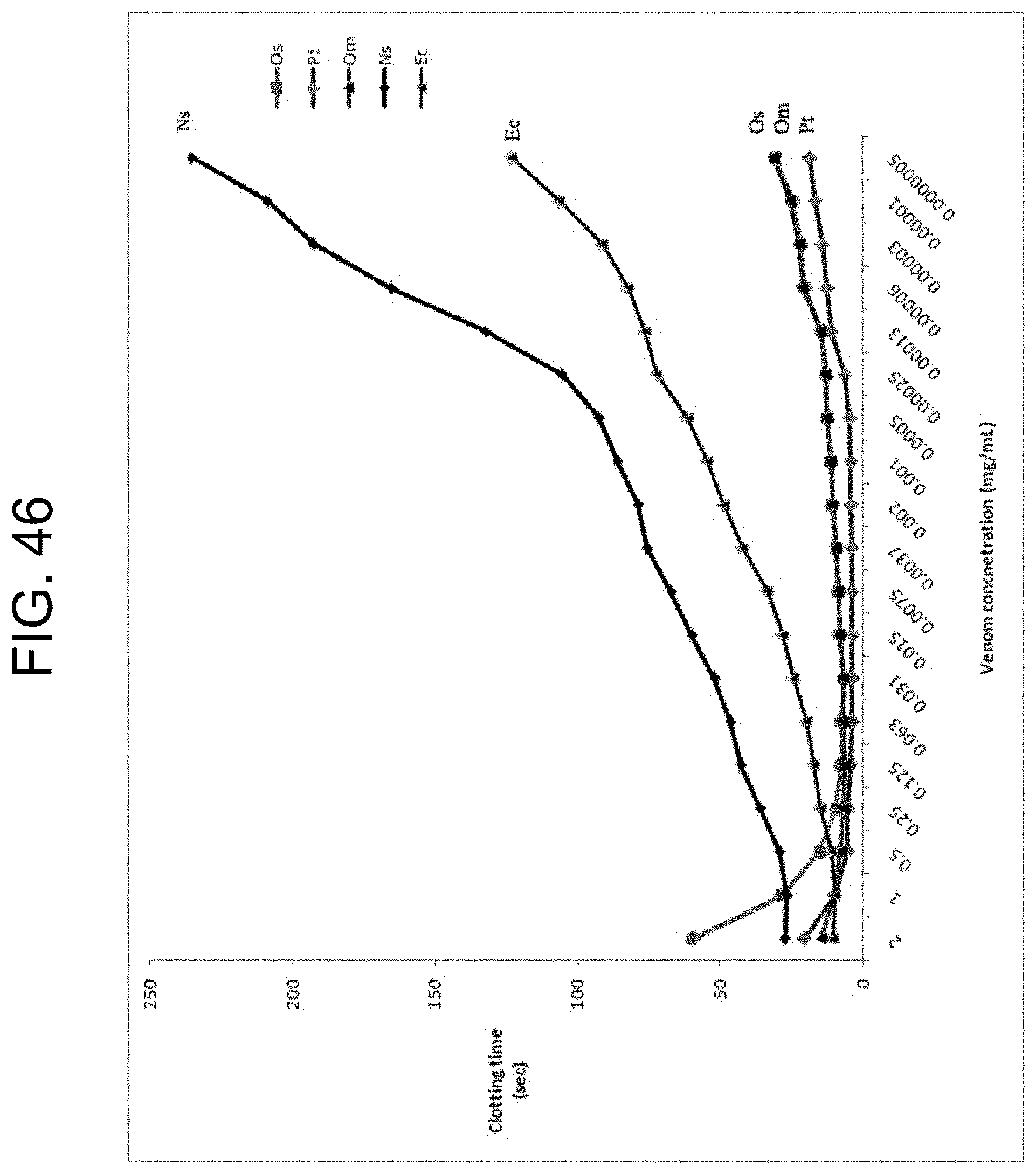
D00047
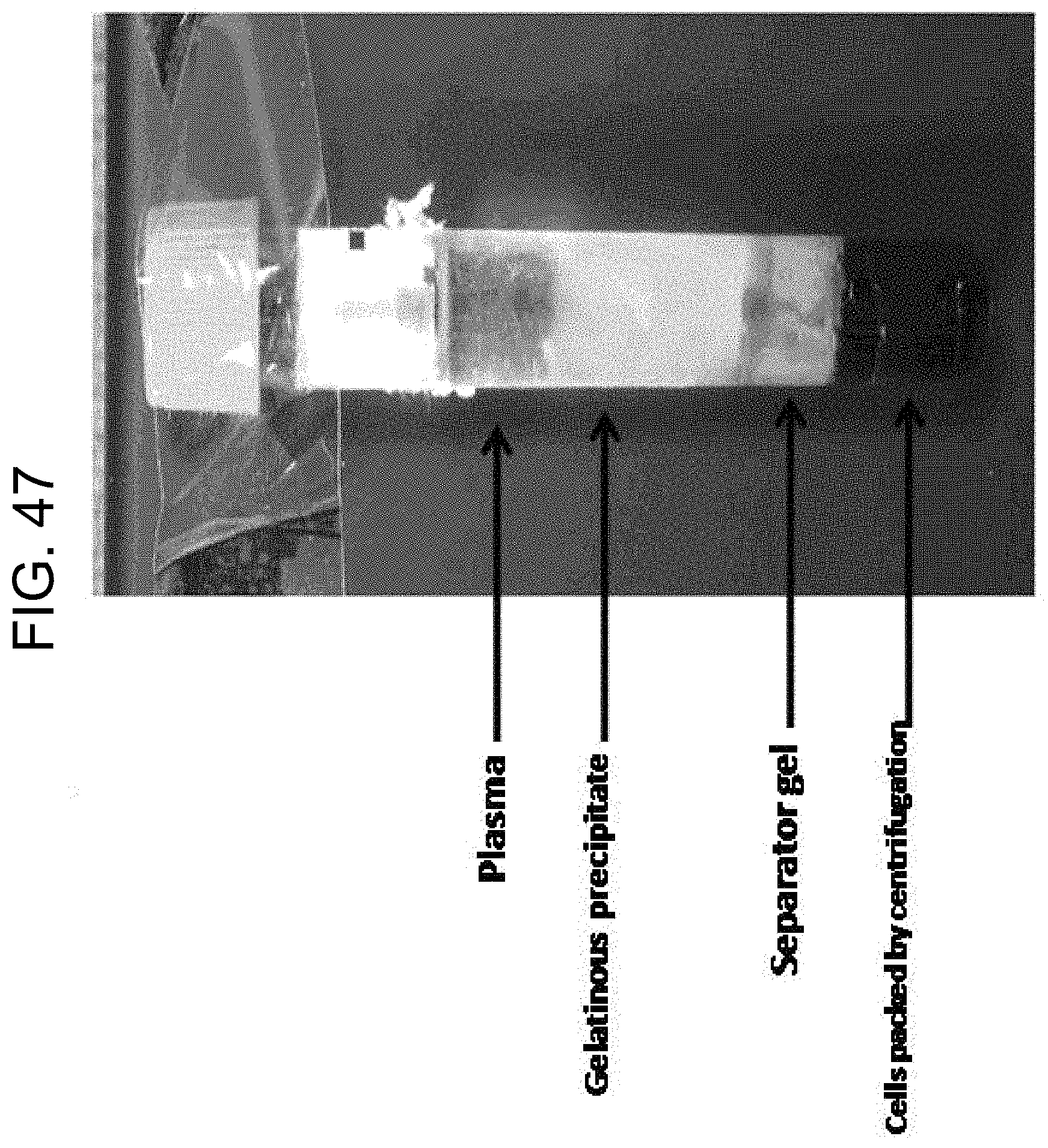

D00048
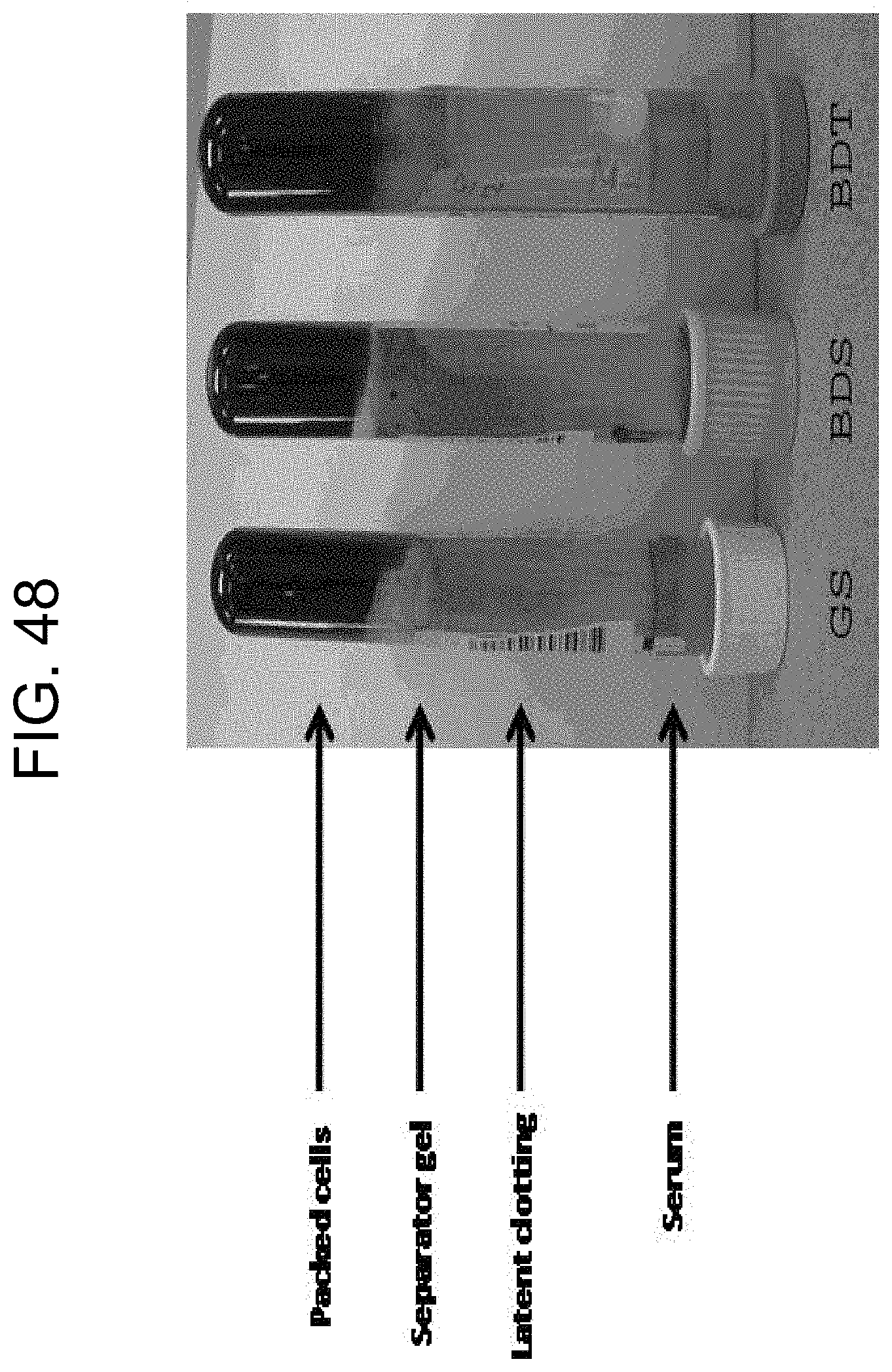
D00049
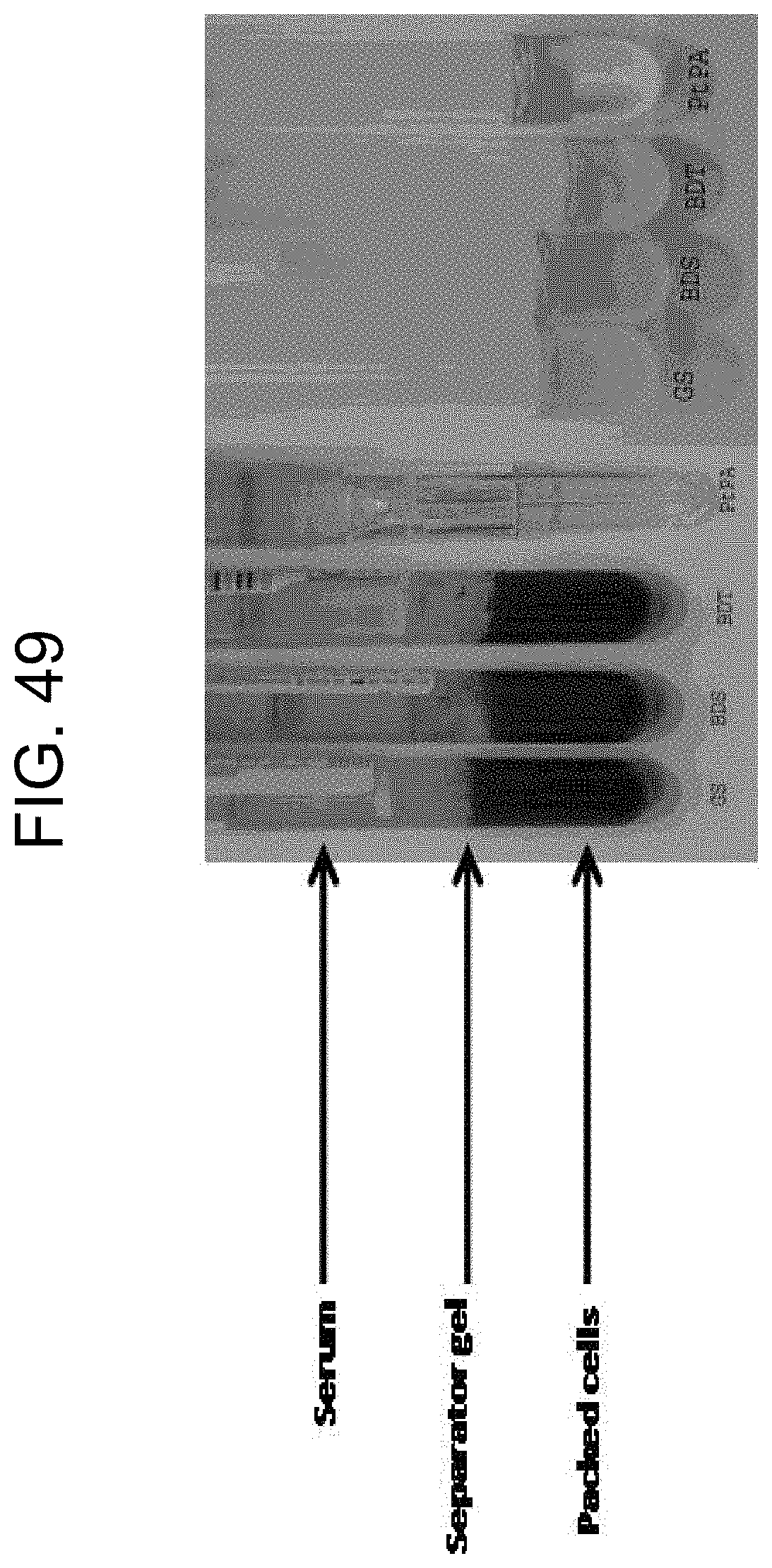
D00050
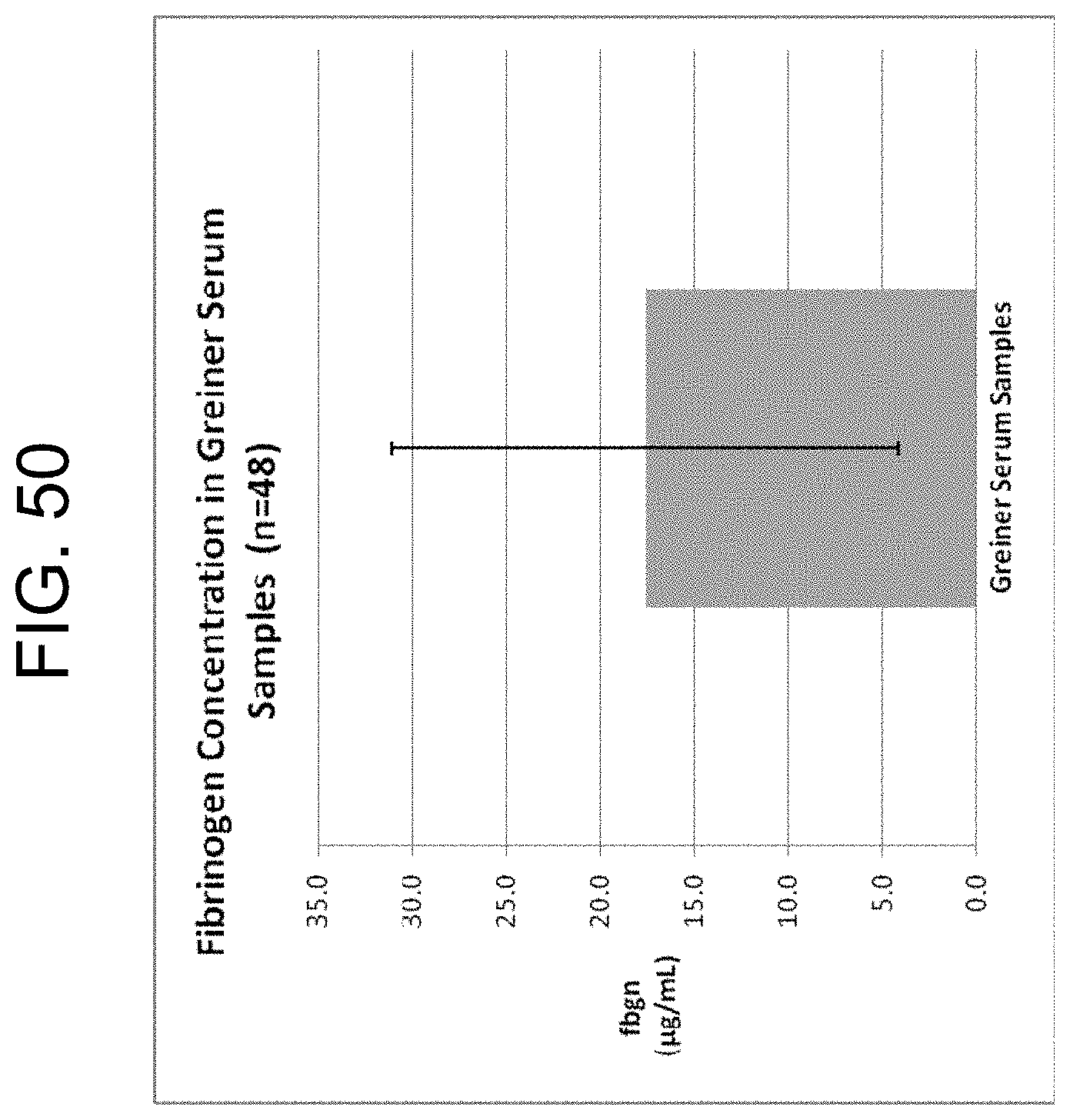
D00051
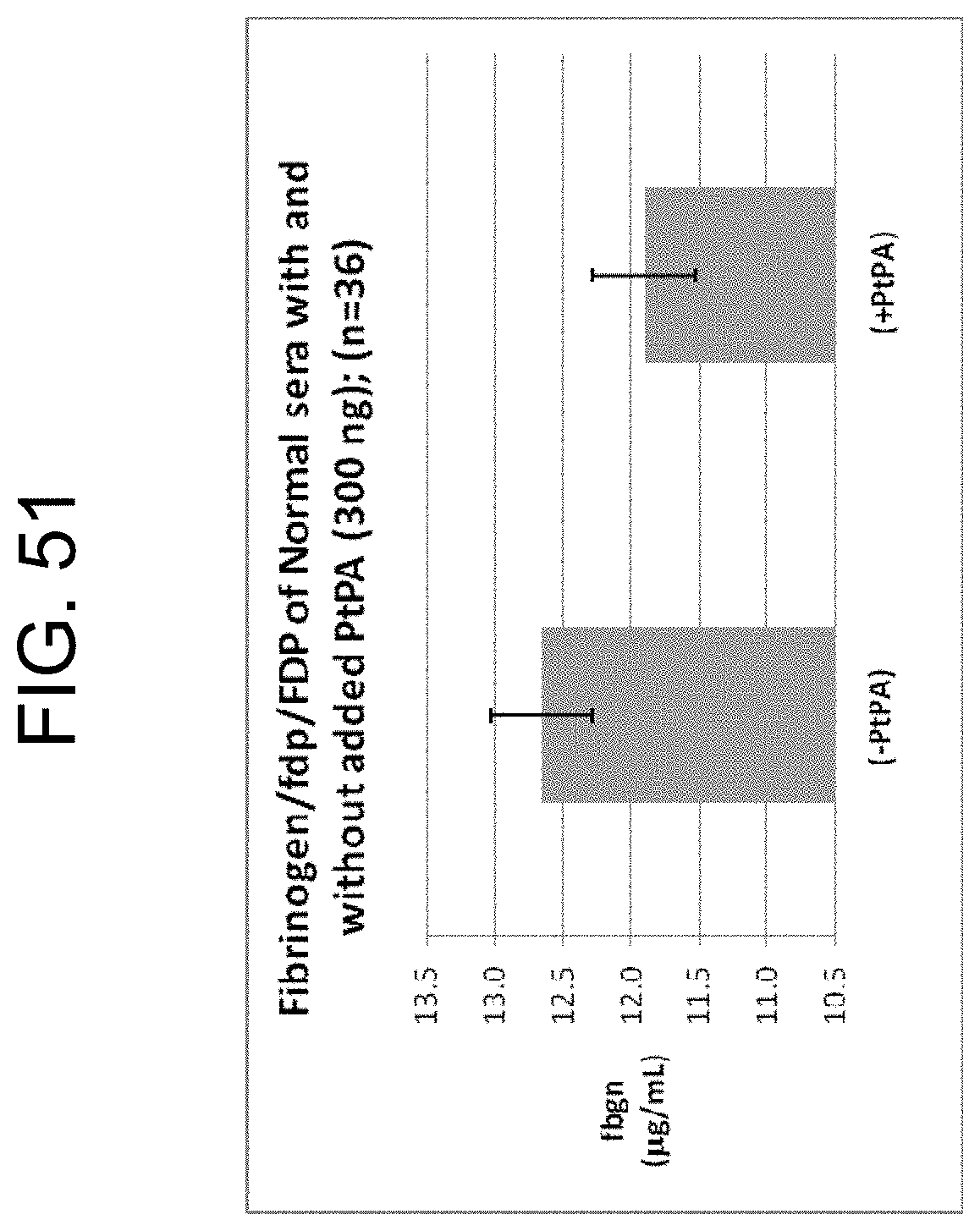
D00052
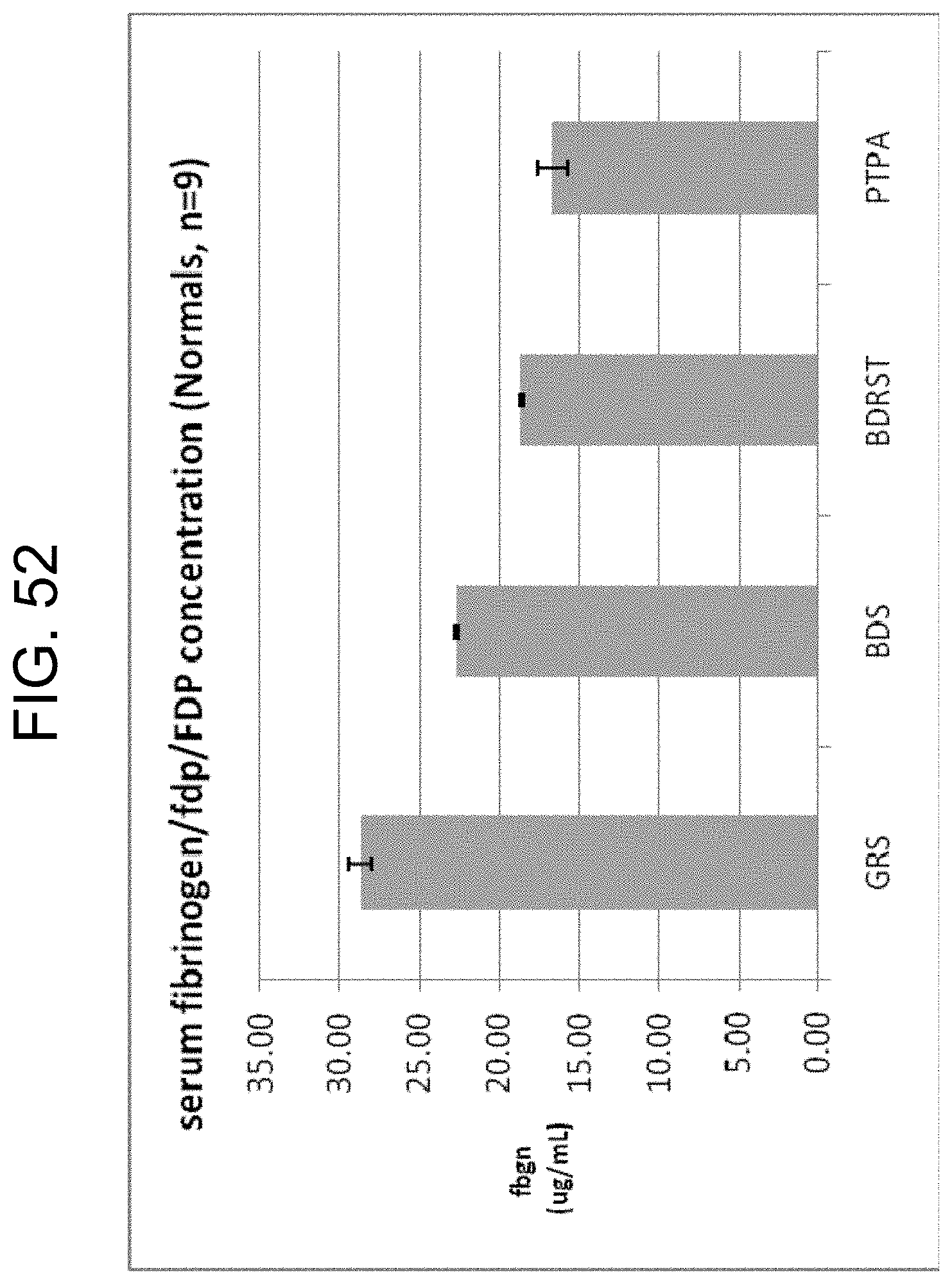
D00053
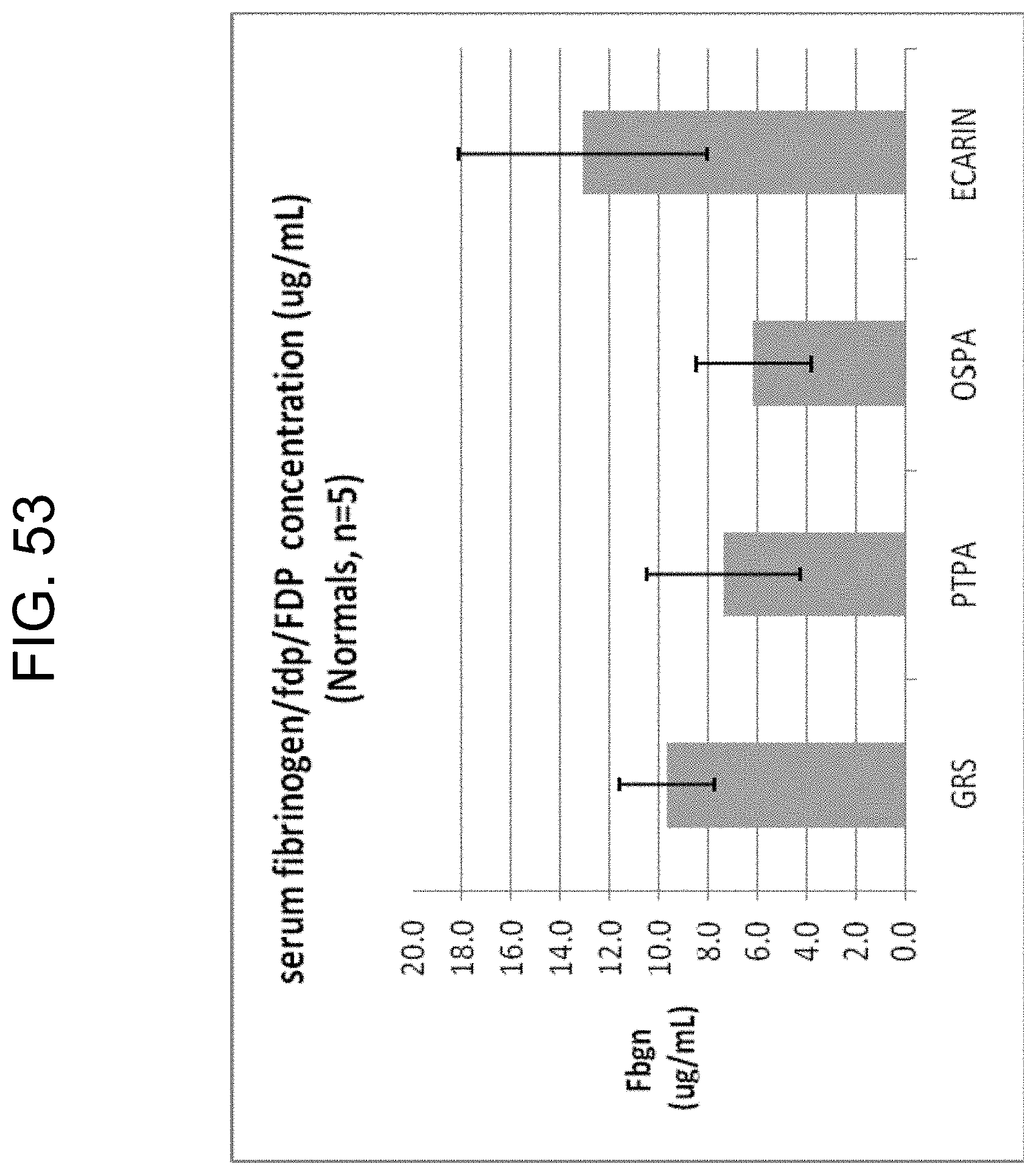
D00054
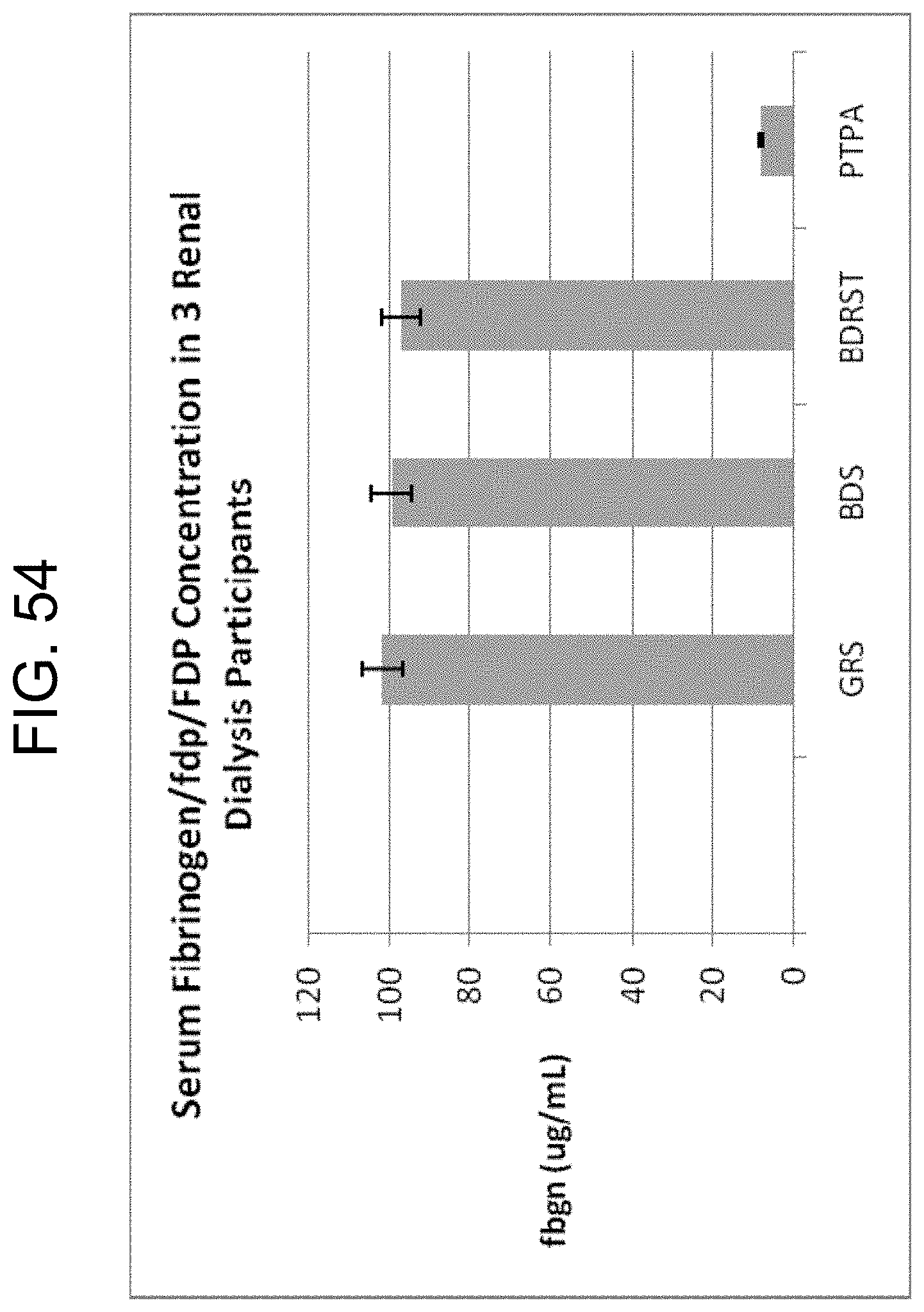

D00055
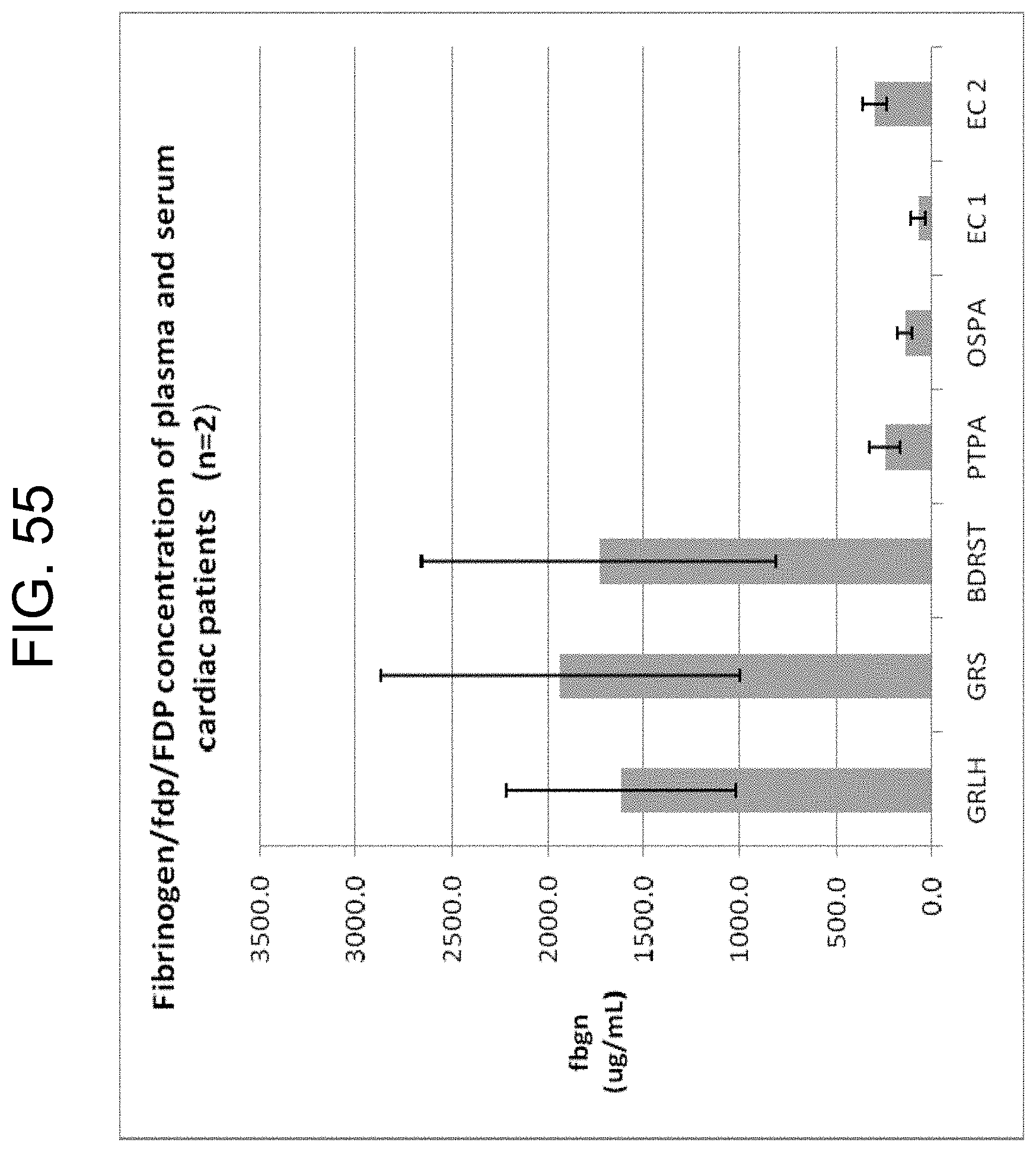
D00056

D00057
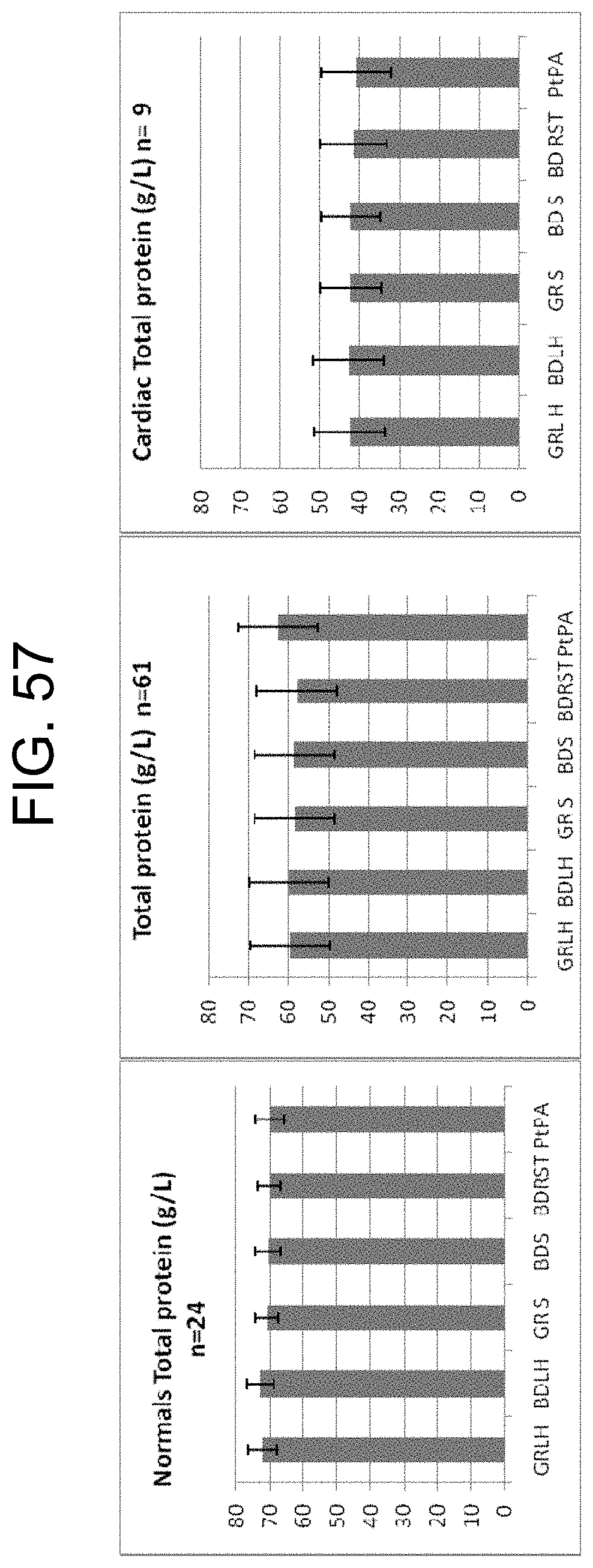
D00058
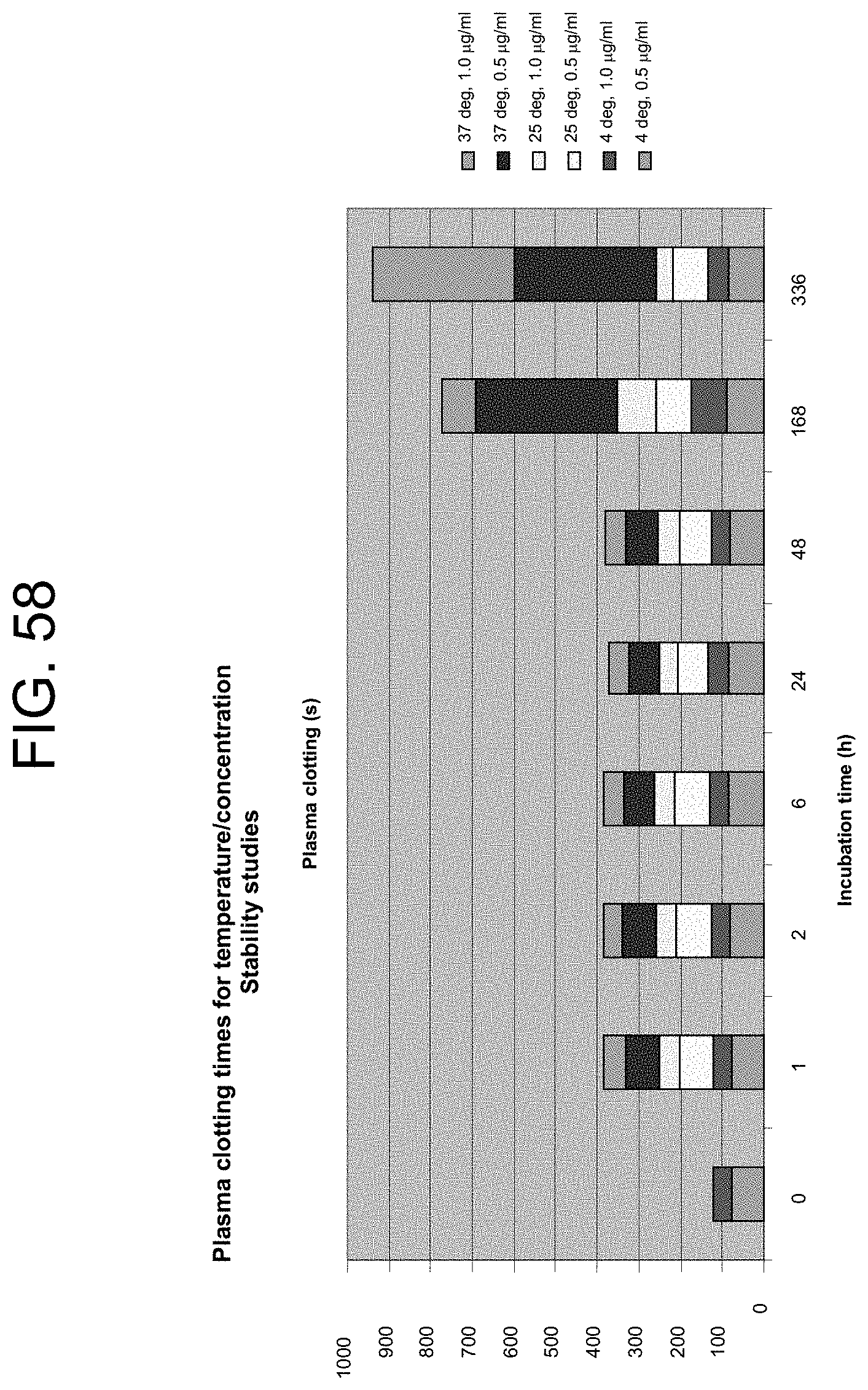
D00059
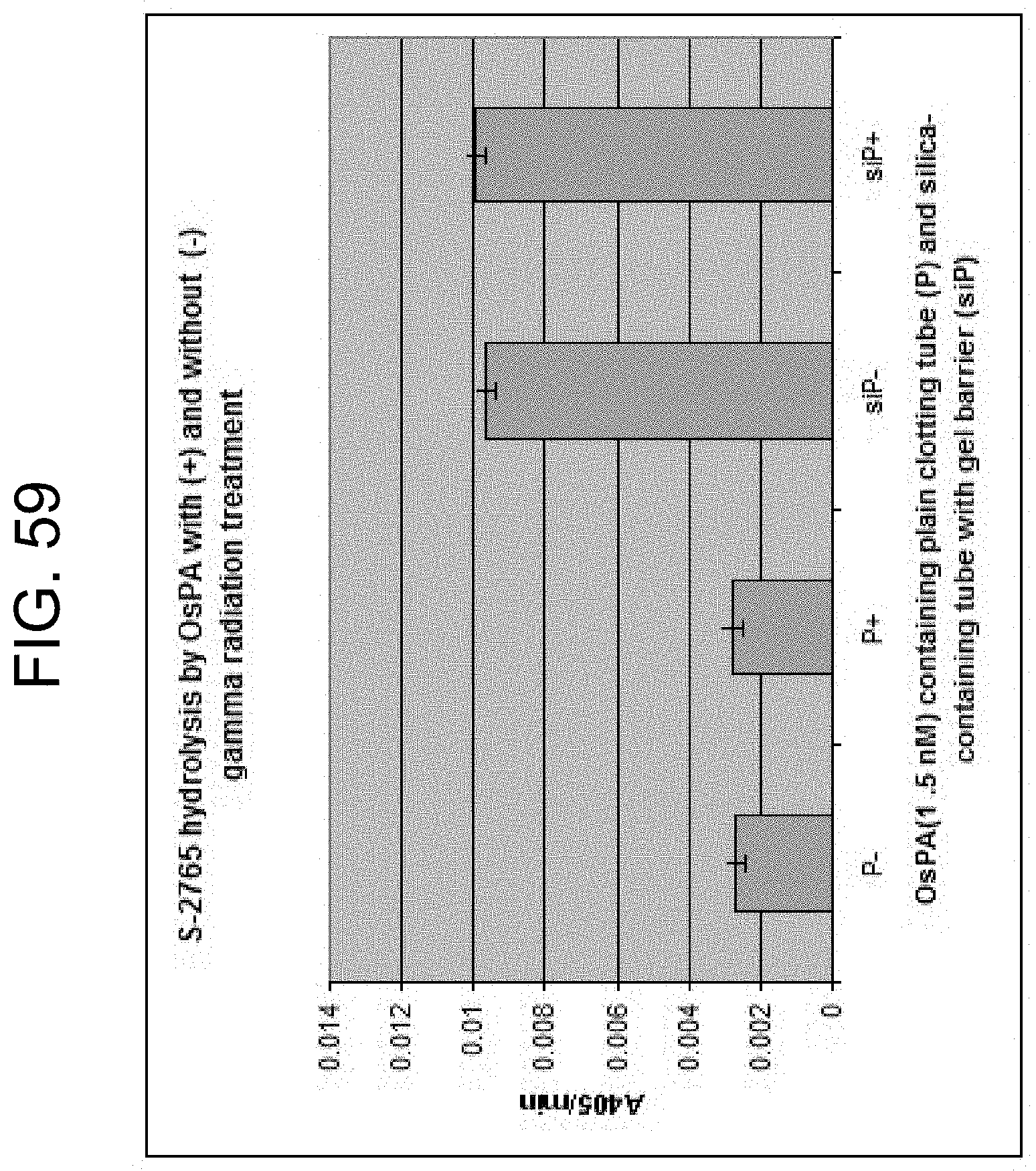
D00060
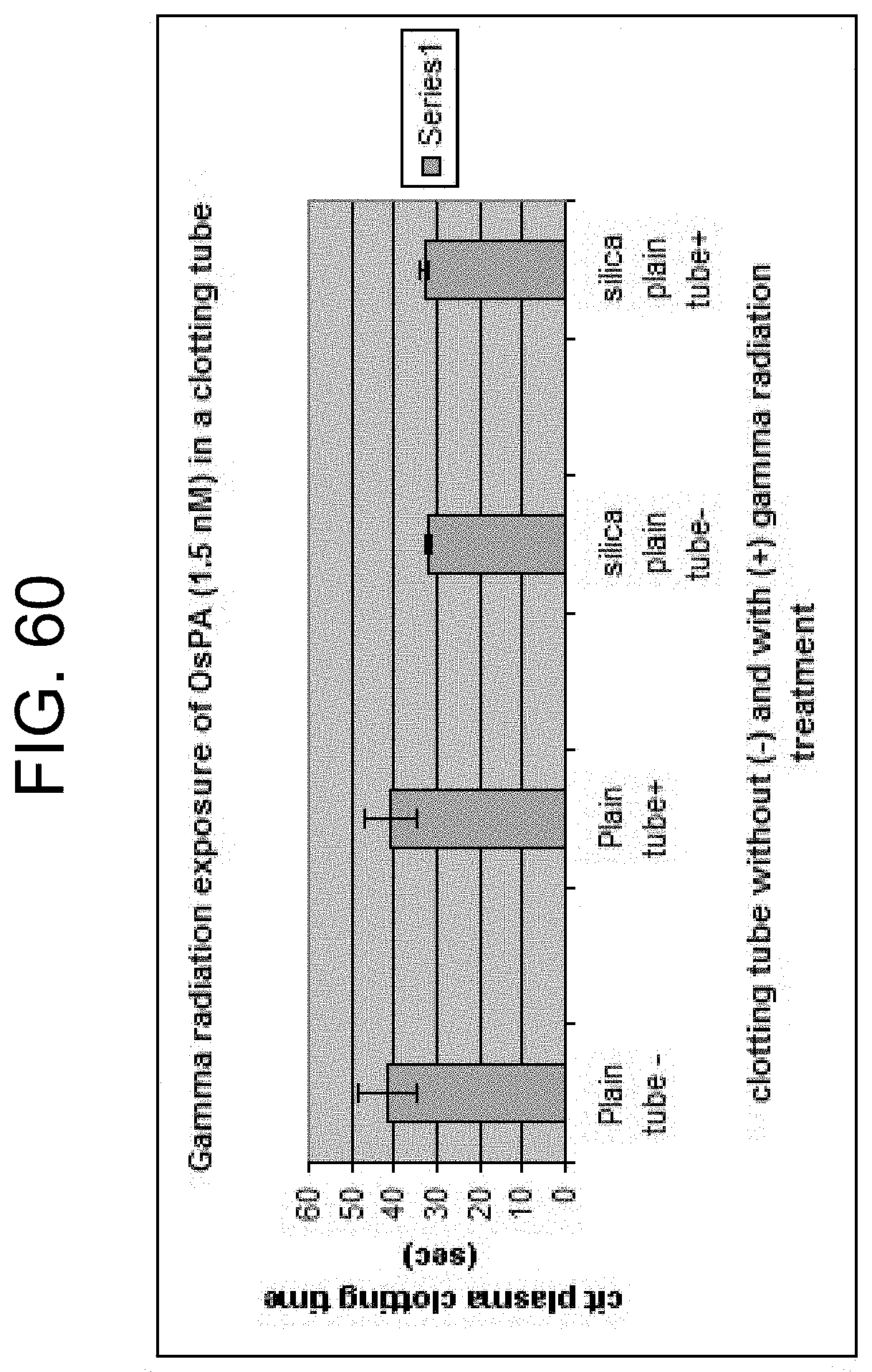
D00061
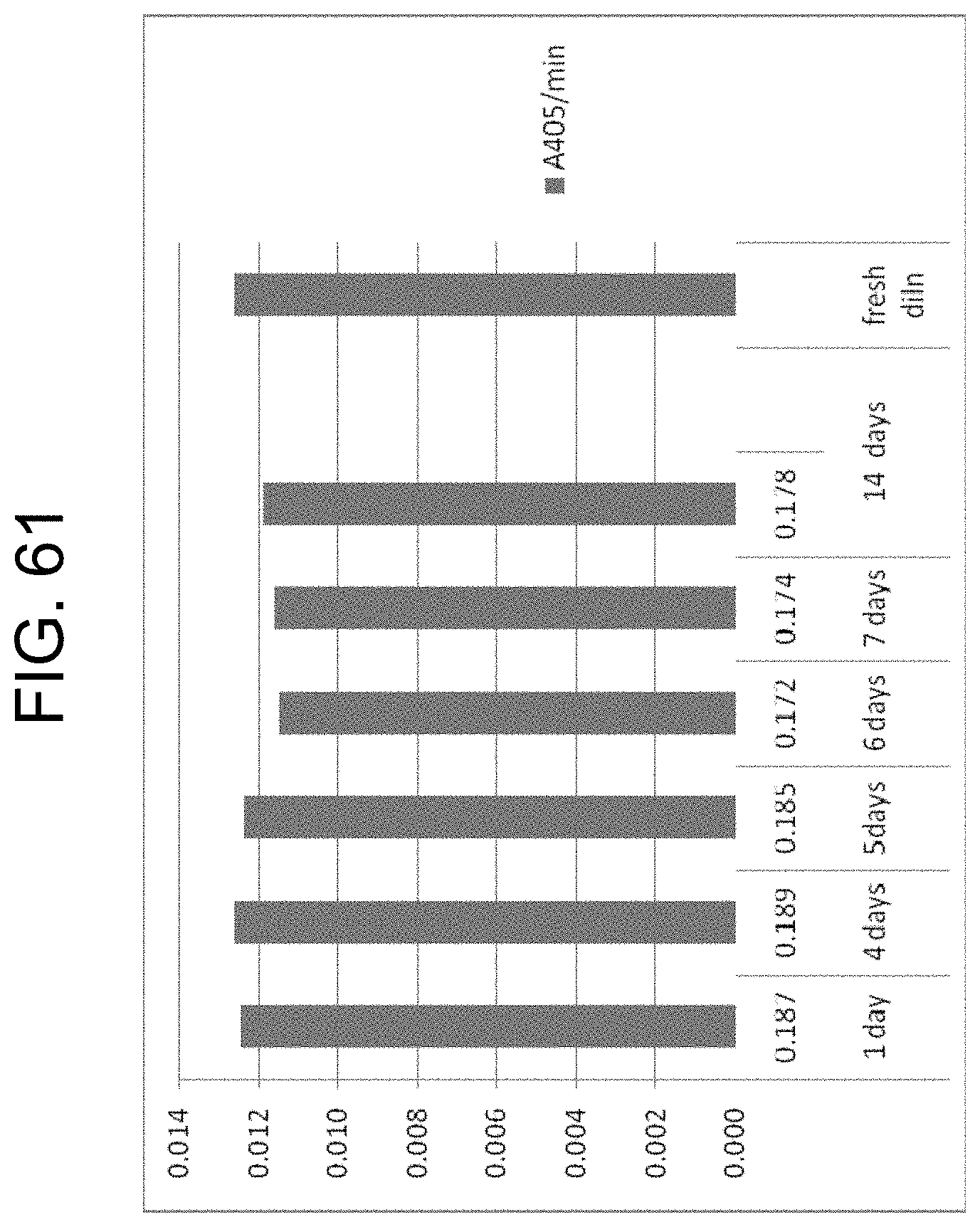
D00062
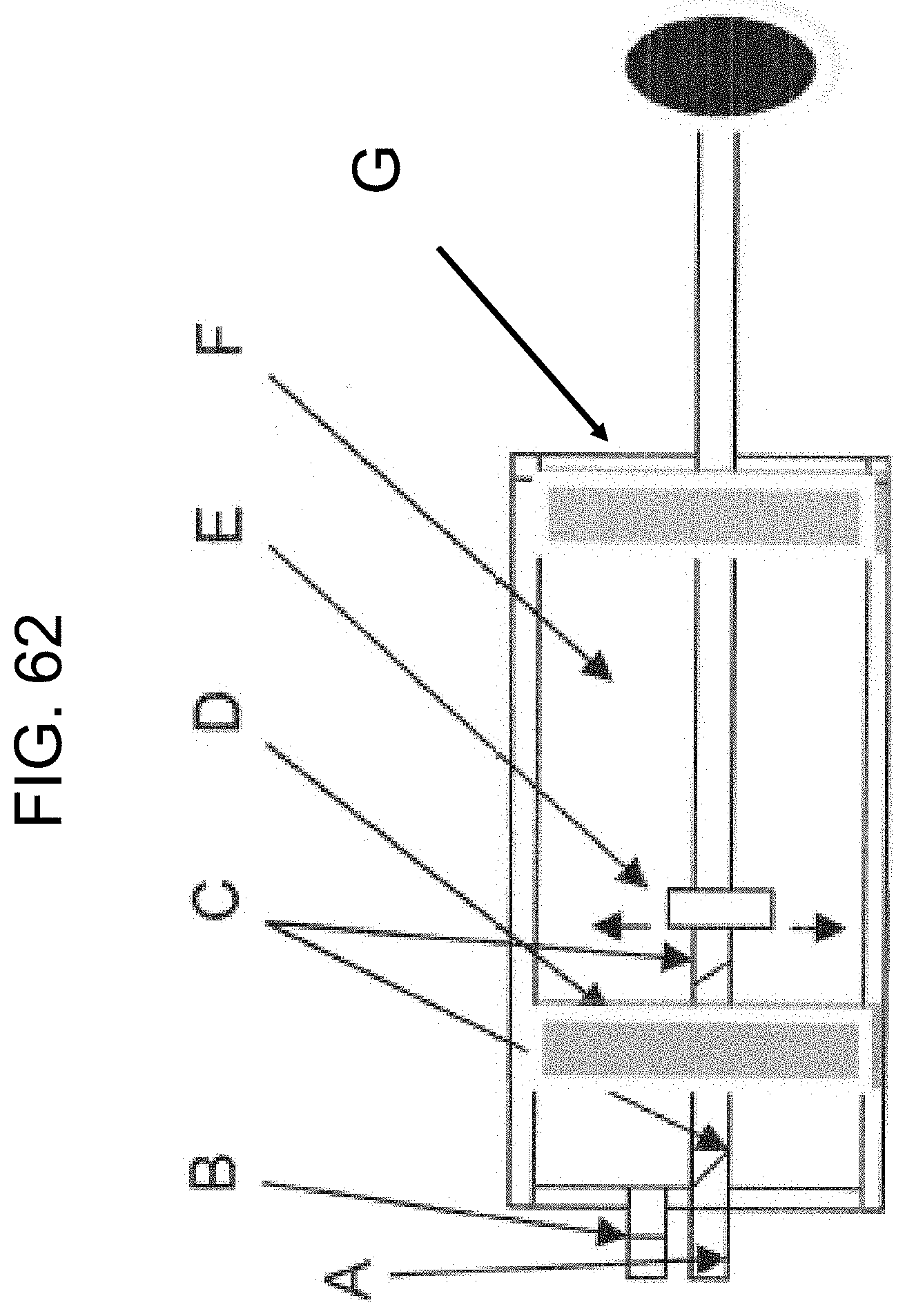
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.