Genetically Encoded Biosensors
Marvin; Jonathan S. ; et al.
U.S. patent application number 16/460112 was filed with the patent office on 2019-10-31 for genetically encoded biosensors. The applicant listed for this patent is The Brigham & Women's Hospital, Inc., Howard Hughes Medical Institute. Invention is credited to Richard T. Lee, Loren Looger, Jonathan S. Marvin, Eric Schreiter.
| Application Number | 20190331678 16/460112 |
| Document ID | / |
| Family ID | 48044437 |
| Filed Date | 2019-10-31 |












View All Diagrams
| United States Patent Application | 20190331678 |
| Kind Code | A1 |
| Marvin; Jonathan S. ; et al. | October 31, 2019 |
GENETICALLY ENCODED BIOSENSORS
Abstract
The present disclosure provides, inter alia, genetically encoded recombinant peptide biosensors comprising analyte-binding framework portions and signaling portions, wherein the signaling portions are present within the framework portions at sites or amino acid positions that undergo a conformational change upon interaction of the framework portion with an analyte.
| Inventors: | Marvin; Jonathan S.; (Arlington, VA) ; Looger; Loren; (Sterling, VA) ; Lee; Richard T.; (Weston, MA) ; Schreiter; Eric; (Ashburn, VA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 48044437 | ||||||||||
| Appl. No.: | 16/460112 | ||||||||||
| Filed: | July 2, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 16112481 | Aug 24, 2018 | 10345297 | ||
| 16460112 | ||||
| 15904574 | Feb 26, 2018 | 10060920 | ||
| 16112481 | ||||
| 15664326 | Jul 31, 2017 | 9939437 | ||
| 15904574 | ||||
| 14350199 | Nov 18, 2014 | 9719992 | ||
| PCT/US2012/059219 | Oct 8, 2012 | |||
| 15664326 | ||||
| 61544867 | Oct 7, 2011 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | G01N 2458/00 20130101; C07K 2319/20 20130101; G01N 33/582 20130101; C07K 14/43595 20130101; C07K 2319/60 20130101; G01N 2400/00 20130101; C07K 14/195 20130101; C07K 14/245 20130101; G01N 33/557 20130101; C07K 2319/24 20130101; G01N 33/6812 20130101; G01N 33/68 20130101 |
| International Class: | G01N 33/557 20060101 G01N033/557; G01N 33/58 20060101 G01N033/58; C07K 14/195 20060101 C07K014/195; C07K 14/245 20060101 C07K014/245; G01N 33/68 20060101 G01N033/68; C07K 14/435 20060101 C07K014/435 |
Claims
1. A recombinant peptide biosensor comprising an analyte-binding framework portion and a signaling portion, wherein the signaling portion is present within the framework portion at a site or amino acid position that undergoes a conformational change upon interaction of the framework portion with a defined, specific, or selected analyte.
Description
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This application is a Divisional of U.S. application Ser. No. 16/112,481, filed Aug. 24, 2018, which is a Divisional of U.S. application Ser. No. 15/904,574, filed Feb. 26, 2018, now U.S. Pat. No. 10,060,920, which is a Divisional of U.S. application Ser. No. 15/664,326, filed Jul. 31, 2017, now U.S. Pat. No. 9,939,437, which is a Divisional of U.S. application Ser. No. 14/350,199, filed Nov. 18, 2014, now U.S. Pat. No. 9,719,992, which is an application under 35 U.S.C. .sctn. 371 of International Application No. PCT/US2012/059219, filed Oct. 8, 2012, which claims priority to U.S.
[0002] Application No. 61/544,867, filed Oct. 7, 2011.
TECHNICAL FIELD
[0003] This disclosure relates to genetically encoded biosensors and methods for the design, production, and use of such biosensors.
BACKGROUND
[0004] Protein-based sensors that transduce microscopic binding events into macroscopically observable signals are available to allow real-time visualization of a variety of biological events and/or molecules (Frommer et al., Chem. Soc. Rev., 38:2833-2841, 2009). Such sensors can be targeted and/or expressed in living cells, tissues, and organisms, and permit imaging with minimally invasive techniques (Okumoto, Curr. Opin. Biotechnol., 21:45-54, 2010). Application of these sensors is limited by the narrow range of analytes that can be detected and/or by their inability to distinguish signal over noise.
SUMMARY
[0005] The present disclosure provides genetically encoded recombinant peptides containing an analyte-binding framework portion linked (e.g., operably linked) to a signaling portion, wherein the signaling portion is allosterically regulated by the framework portion upon interaction of the framework portion with an analyte (e.g., a defined, selected, and/or specific analyte). These constructs can be used as biosensors, e.g., to transduce microscopic binding events into macroscopically observable signals.
[0006] The present disclosure provides, in part, recombinant peptides for use as biosensors (e.g., recombinant peptide biosensors) that include (e.g., comprise, consist essentially of, or consist of), e.g., include at least, an analyte-binding framework portion and a signaling portion. As described in further detail herein, such signaling portions are present within the framework portion at a site or amino acid position that undergoes a conformational change (e.g., a conformational change sufficient to alter a physical and/or functional characteristic of the signaling portion, e.g., a substantial conformational change) upon interaction of the framework portion with a defined, specific, or selected analyte (e.g. such as an analyte to which the framework portion or a region thereof, and/or the biosensor, specifically binds). For example, in some instances, the signaling portion is allosterically regulated by the framework portion such that signaling from the signaling portion is altered (e.g. wherein a first level of signaling is altered or changed to a second level of signaling that can be distinguished using routine methods of detection from the first) upon interaction of the framework portion with the analyte. In some instances, signaling by the signaling portion can detectably increase or decrease upon interaction of the framework portion with the analyte. In some instances, signaling by the signaling portion upon interaction of the biosensor with a defined, specific, or selected analyte (e.g. such as an analyte to which the framework portion or a region thereof, and/or the biosensor, specifically binds) can be proportional or can correlate with to the level of interaction between the framework portion and the analyte such that the level of interaction can be determined from the signaling or alteration thereof.
[0007] In some instances, framework portions of the biosensors disclosed herein have a first structure in the absence of an analyte and a second structure that is detectably distinct from the first structure in the presence of the analyte. In some instances, the conformational change between the first structure and the second structure allosterically regulates the signaling portion.
[0008] In some instances, framework portions of the biosensors disclosed herein can be, or can include (e.g., comprise, consist essentially of, or consist of), periplasmic binding proteins (PBP) or variants of a PBP. In some instances, exemplary PBPs or variants thereof can include, but are not limited to, peptides with at least 90% identity to a peptide selected from the group consisting of SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO:107, SEQ ID NO:108, SEQ ID NO:109, SEQ ID NO: 110, SEQ ID NO:111, SEQ ID NO:113, and SEQ ID NO:114. In some instances, exemplary PBPs or variants thereof can include, but are not limited to, peptides with at least 95% identity to a peptide selected from the group consisting of SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO:107, SEQ ID NO:108, SEQ ID NO:109, SEQ ID NO: 110, SEQ ID NO:111, SEQ ID NO:113, and SEQ ID NO:114. In some instances, exemplary PBPs or variants thereof can include, but are not limited to, peptides selected from the group consisting of SEQ ID NO:105, SEQ ID NO: 106, SEQ ID NO:107, SEQ ID NO:108, SEQ ID NO:109, SEQ ID NO: 110, SEQ ID NO:111, SEQ ID NO:113, and SEQ ID NO:114. In some instances, exemplary PBPs or variants thereof can include, but are not limited to, peptides selected from the group consisting of SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 107, SEQ ID NO:108, SEQ ID NO:109, SEQ ID NO: 110, SEQ ID NO:111, SEQ ID NO:113, and SEQ ID NO: 114 comprising 10 or fewer conservative amino acid substitutions. PBPs or variants thereof disclosed herein can be truncated.
[0009] In some instances, signaling portions of the biosensors disclosed herein can be or can include (e.g., comprise, consist essentially of, or consist of) one or more (e.g., one, two three, four, five, and less than ten) circularly permuted fluorescent proteins (cpFPs). Such cpFPs can be include but are not limited to, for example, green fluorescent proteins, yellow fluorescent proteins, red fluorescent proteins, and/or blue fluorescent proteins.
[0010] In some instances, biosensors disclosed herein, e.g., analyte-binding framework portions of biosensors disclosed herein, can bind (e.g., bind specifically) to glucose. Such sensors can be referred to as glucose binding biosensors or glucose biosensors.
[0011] In some instances, biosensors disclosed herein, e.g., analyte-binding framework portions of biosensors disclosed herein, can bind (e.g., bind specifically) to maltose. Such sensors can be referred to as maltose binding biosensors or maltose biosensors.
[0012] In some instances, biosensors disclosed herein, e.g., analyte-binding framework portions of biosensors disclosed herein, can bind (e.g., bind specifically) to phosphonate. Such sensors can be referred to as phosphonate binding biosensors or phosphonate biosensors.
[0013] In some instances, biosensors disclosed herein, e.g., analyte-binding framework portions of biosensors disclosed herein, can bind (e.g., bind specifically) to glutamate. Such sensors can be referred to as glutamate binding biosensors or glutamte biosensors.
[0014] In some instances, biosensors disclosed herein can include (e.g., comprise, consist essentially of, or consist of): an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and 53, wherein the recombinant peptide biosensor binds specifically to maltose; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and 53 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to maltose; and/or a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and 53.
[0015] In some instances, biosensors disclosed herein can include (e.g., comprise, consist essentially of, or consist of): an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 62 and 63, wherein the recombinant peptide biosensor binds specifically to glutamate; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 62 and 63 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to glutamate; and/or a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 62 and 63.
[0016] In some instances, biosensors disclosed herein can include (e.g., comprise, consist essentially of, or consist of): an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 77 and 78, wherein the recombinant peptide biosensor binds specifically to phosphonate; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 77 and 78 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to phosphonate; and/or a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 77 and 78.
[0017] In some instances, biosensors disclosed herein can include (e.g., comprise, consist essentially of, or consist of): an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 91, 92, 93 and 94, wherein the recombinant peptide biosensor binds specifically to glucose; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 91, 92, 93 and 94 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to glucose; and/or a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 91, 92, 93 and 94.
[0018] In some instances, biosensors disclosed herein can include (e.g., comprise, consist essentially of, or consist of): SEQ ID NO:91; SEQ ID NO:92; SEQ ID NO:93; SEQ ID NO:95.
[0019] In some instances, any recombinant biosensor disclosed herein can be isolated and/or purified. The terms "isolated" or "purified," when applied to a biosensor disclosed herein includes nucleic acid proteins and peptides that are substantially free or free of other cellular material or culture medium when produced by recombinant techniques, or substantially free or free of precursors or other chemicals when chemically synthesized.
[0020] The disclosure also provides, in part, nucleic acids (e.g., isolated and/or purified nucleic acids) encoding any one or more of the recombinant peptide biosensors disclosed herein. For example, nucleic acids can encode: an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and 53, wherein the recombinant peptide biosensor binds specifically to maltose; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and 53 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to maltose; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and 53; an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 62 and 63, wherein the recombinant peptide biosensor binds specifically to glutamate; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 62 and 63 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to glutamate; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 62 and 63; an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 77 and 78, wherein the recombinant peptide biosensor binds specifically to phosphonate; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 77 and 78 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to phosphonate; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 77 and 78; an amino acid sequence with at least 90% identity to a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 91, 92, 93 and 94, wherein the recombinant peptide biosensor binds specifically to glucose; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 91, 92, 93 and 94 comprising 10 or fewer conservative amino acid substitutions, wherein the recombinant peptide biosensor binds specifically to glucose; a recombinant peptide biosensor selected from the group consisting of SEQ ID NO: 91, 92, 93 and 94; and/or SEQ ID NO:91; SEQ ID NO:92; SEQ ID NO:93; SEQ ID NO:95.
[0021] In some instances, the disclosure includes vectors containing one or a plurality of the nucleic acids disclosed herein and cells containing such vectors. In some instances, the disclosure provides cells containing one or a plurality of nucleic acids disclosed herein.
[0022] In some instances, the disclosure includes kits related to the biosensors and nucleic acids disclosed herein Such kits can include or contain, for example, a biosensor, a nucleic acid encoding a biosensor, vectors, and/or cells, provided herein.
[0023] In some instances, the disclosure provides methods related to the biosensors and nucleic acids disclosed herein. Such methods can include methods of making, using, and/or selling the biosensors and nucleic acids disclosed herein. For example, methods can include methods for producing genetically encoded recombinant peptide biosensors. In such instances, methods can include, for example, selecting a framework portion that binds specifically to a target analyte and that undergoes a conformational change upon interacting binding to the target analyte, identifying a site or amino acid position within the selected framework portion where or around which the conformational change occurs, and inserting a signaling portion into the site or amino acid position. In some instances, framework portions include periplasmic binding proteins (PBPs) disclosed herein. Exemplary PBPs include PBPs that bind (e.g., bind specifically) to glucose.
[0024] In some instances, the present disclosure includes methods for detecting glucose, e.g., in a sample containing a level of glucose. Such methods can include, detecting a level of fluorescence emitted by a recombinant peptide biosensor, the peptide biosensor having an amino acid sequence selected from the group consisting of SEQ ID NO: 91, 92, 93 and 94, and correlating the level of fluorescence with the presence of glucose. In some instances, recombinant peptide biosensors used in the methods herein are expressed from nucleic acids. In some instances, methods include contacting the recombinant peptide biosensor with a test sample (e.g., a sample comprising glucose). In some instances, methods can include the level of fluorescence emitted by a biosensor (e.g., a biosensor bound to glucose) with a concentration glucose in the sample. Such correlation can include, for example, comparing the level of fluorescence with a level of fluorescence emitted by the recombinant peptide biosensor in the presence of a sample comprising a known concentration or range of concentrations of glucose. In some instance, the level of fluorescence emitted by the recombinant peptide biosensor in the presence (e.g., bound or bound specifically to) of a sample comprising a known concentration or range of concentrations of glucose is stored on an electronic database.
[0025] One of skill will appreciate that such methods can be adapted for any defined, specific, or selected analyte. For example, in some instances, the disclosure provides methods for detecting a defined, selected, or specific analyte. These methods can include detecting a level of fluorescence emitted by a recombinant peptide biosensor expressed from a nucleic acid and correlating the level of fluorescence with the presence the defined, selected, or specific analyte. In some instances, methods include contacting the recombinant peptide biosensor with a sample comprising the analyte. In some instances, methods include correlating the level of fluorescence with a concentration of the analyte. In some instances, methods include comparing the level of fluorescence with a level of fluorescence emitted by the recombinant peptide biosensor in the presence of a sample comprising a known concentration or range of concentrations of the analyte, wherein the level of fluorescence emitted by the recombinant peptide biosensor in the presence of a sample comprising a known concentration or range of concentrations of the analyte is stored on an electronic database.
[0026] In some instances, the present disclosure provides methods for detecting a defined, selected, or specific analyte, the method comprising detecting a level of fluorescence emitted by a recombinant peptide biosensor of any one of claims 1-36; and correlating the level of fluorescence with the presence of a defined, selected, or specific analyte. In some instances, recombinant peptide biosensors can be expressed from a nucleic acid. In some instances, methods can include contacting the recombinant peptide biosensor with a sample comprising the analyte. In some instances, methods can include correlating the level of fluorescence with a concentration of the analyte and, optionally, comparing the level of fluorescence with a level of fluorescence emitted by the recombinant peptide biosensor in the presence of a sample comprising a known concentration or range of concentrations of the analyte. In some instances, the level of fluorescence emitted by the recombinant peptide biosensor in the presence of a sample comprising a known concentration or range of concentrations of the analyte is stored on an electronic database.
[0027] Methods herein can be performed in vitro.
[0028] In some instances, the present disclosure provides compositions containing any one or a plurality of the peptide biosensors and/or nucleic acids disclosed herein.
[0029] Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Methods and materials are described herein for use in the present invention; other, suitable methods and materials known in the art can also be used. The materials, methods, and examples are illustrative only and not intended to be limiting. All publications, patent applications, patents, sequences, database entries, and other references mentioned herein are incorporated by reference in their entirety. In case of conflict, the present specification, including definitions, will control.
[0030] Other features and advantages of the invention will be apparent from the following detailed description and figures, and from the claims.
DESCRIPTION OF DRAWINGS
[0031] FIG. 1|Cartoon representation showing ligand bound Escherichia Coli malto-dextrin-binding protein (EcMBP) and potential circularly-permuted fluorescent protein (cpFP) insertion sites.
[0032] FIG. 2|Cartoon representation showing ligand bound Pyrococcus furiosus maltotriose binding protein (PfMBP) and potential cpFP insertion sites.
[0033] FIG. 3|Cartoon representation showing ligand bound E. coli glutamate-binding protein (EcYbeJ) and potential cpFP insertion sites.
[0034] FIG. 4|Cartoon representation showing ligand bound E. coli phosphonate-binding protein (EcPhnD) and potential cpFP insertion sites.
[0035] FIG. 5|Cartoon representation showing ligand bound Thermus thermophilus glucose binding protein (TtGBP) and potential cpFP insertion sites.
[0036] FIG. 6A-B|Changes in EcMBP upon maltose binding and locations at which circularly-permuted fluorescent protein (cpFP) was inserted are shown as colored spheres at the C.alpha. positions. Yellow: 165-166, Green: 175-176, Cyan: 311-312, Violet: 317-318(A). (B) shows backbone structural changes. The C.alpha. dihedral is calculated from the four atoms: C.alpha.i+2, C.alpha.i+1, C.alpha.i, C.alpha.i-1. .DELTA.Dihedral is calculated as the difference in dihedrals between the closed (1ANF) and open (1OMP) states of MBP, and corrected to fall within a range of -180.degree. to 180.degree.. The regions near residues 175 and 311 are labeled. There is a crystallographic artifact at the N-terminus resulting in the appearance of significant structural changes.
[0037] FIG. 7A|Amino acid sequence of MBP-165-cpGFP (SEQ ID NO: 1).
[0038] FIG. 7B|Amino acid sequence of MBP-165-cpGFP.PPYF (SEQ ID NO:2).
[0039] FIG. 7C|Amino acid sequence of MBP-165-cpGFP.PCF (SEQ ID NO:3).
[0040] FIG. 8A|Amino acid sequence of MBP-175-cpGFP (SEQ ID NO:4).
[0041] FIG. 8B|Amino acid sequence of MBP-175-cpGFP.L1-HL (SEQ ID NO:5).
[0042] FIG. 9A|Amino acid sequence of MBP-311-cpGFP (SEQ ID NO:6).
[0043] FIG. 9B|Amino acid sequence of MBP-311-cpGFP.L2-NP (SEQ ID NO:7).
[0044] FIG. 10|Amino acid sequence of MBP-317-cpGFP (SEQ ID NO:8).
[0045] FIGS. 11A-11D|Line charts showing EcMBP plot of .DELTA.F/F for clarified lysate screen of cpGFP linker-screens at insertion points 165, 175, 311, and 317. The horizontal dashed line at zero indicates no fluorescence change. Standard deviations in .DELTA.F/F are less than 10% of an average .DELTA.F (repetitions for MBP165-cpGFP.PPYF yields .DELTA.F/F values of 2.51, 2.63, and 2.54).
[0046] FIG. 12|Isothermal titration calorimetry (ITC) of MBP317-cpGFP with maltose.
[0047] FIG. 13|Graph showing EcMBP165-cpGFP.PPYF affinity variant binding maltose-binding curves. Binding curves for affinity variants of MBP165-cpGFP.PPYF. Data is fit to a single-binding site isotherm. Curve-fit affinities are: WT binding pocket, 5 .mu.M (.circle-solid.); W230A, 32 .mu.M (.box-solid.); W62A, 375 .mu.M (.tangle-solidup.); W340A, >1 mM (); I329W, 11 .mu.M (.quadrature.).
[0048] FIGS. 14A-14D|Line graphs showing maltose and sucrose binding curves for wild-type and 5-7 variants of the EcMBP-cpGFP sensors. Maltose (black) and sucrose (red) binding curves for wild-type (filled, solid lines) and 5-7 variants (open, dashed lines) of the MBP-cpGFP sensors. MBP165-cpGFP.PPYF (a); MBP165-cpGFP.PCF (b); MBP175-cpGFP.L1-HL (c); MBP311-cpGFP.L2-NP (d).
[0049] FIGS. 15A-15D|Line graphs showing emission spectra for colored variants of EcMBP sensors. Fluorescence emission spectra of the MBP165-Blue, Cyan, Green, and Yellow wild-type sensors (a) and the 5-7 variants (b) in the absence of ligand (dashed lines, open circles), with 10 mM maltose (solid lines, filled circles), or 10 mM sucrose (solid lines, filed squares). Sensors were excited at 383, 433, 485, and 485 nm, respectively. Titration of maltose and sucrose in the Blue, Cyan, Green, and Yellow MBP165 wild-type sensors (c) and for the 5-7 variants (d). Filled circles are titration of maltose, open circles are titration of sucrose. For the wild-type sensors, Kds for maltose binding are: Blue 3.3 .mu.M, Cyan 13 .mu.M, Green 4.5 .mu.M, Yellow 3.3 .mu.M. No sucrose binding is observed. For the 5-7 variants, Kd of Green is 2.4 mM (sucrose) and 7.1 mM (maltose). Kd of Yellow is 2.5 mM (sucrose) and 4.5 mM (maltose).
[0050] FIG. 16|Plot of .DELTA.F/F for clarified lysate screen of MBP165-cpBFP linker-screen. The horizontal dashed line at zero indicates no fluorescence change.
[0051] FIGS. 17A-17B|Line graphs showing maltose binding. Blue (wt binding pocket) has an affinity of 2.7 .mu.M. Green (W230A) has an affinity of 40 .mu.M. Yellow (W62A) has an affinity of 350 .mu.M. Cyan (W340A) has an affinity of approximately 1.7 mM. Data is plotted at .DELTA.F/F (a) or normalized to Fractional Saturation (b).
[0052] FIGS. 18A-18C|Images bacterial cells expressing (a) EGFP, (b) PPYF, or (c) PPYF.T203V in the absence (top) and presence (bottom) of maltose.
[0053] FIGS. 19A-19B|Line graphs showing EcMBP-cpGFP.PPYF.T203V 2-photon excitation spectra. MBP165-cpAzurite.L2-FE (a), -cpCFP.PCF (a), -cpGFP.PPYF (b), and -cpYFP.PPYF (b) were excited at the wavelengths indicated and emission measured through appropriate wavelength filters. Two graphs are shown to present different y-axis scales. Optimal .DELTA.F/F values for 2-photon excitation of the spectral variants of MBP165 are: -cpAzurite, 1.1 (ex 760 nm); -cpCFP, 2.3 (ex 830-960 nm); -cpGFP, 10.0 (ex 940 nm); -cpYFP, 2.6 (ex 940 nm).
[0054] FIGS. 20A-20C|Images showing EcMBP-cpGFP.PPYF.T203V expressing HEK cells. Images of individual HEK293 cells expressing membrane displayed PPYF.T203V in the absence of maltose (a), in the presence of 1 mM maltose (b), and after washout with maltose-free buffer (c). Scale bars are 10 .mu.m.
[0055] FIGS. 21A-21B|Graphs showing quantification of fluorescence of EcMBP-cpGFP.PPYF.T203V when displayed on the surface of HEK cells. (a) Concentration dependence. (b) Observed fluorescence after a "puff" of HBSS solution containing 1 mM maltose and 2.5 nM Alexa Fluor.RTM. 568 (Invitrogen, Carlsbad, Calif.).
[0056] FIGS. 22A-22D|Cartoon representations and close-up views of inter-domain linkers and selected amino acids of the cpGFP chromophore environment of the structure of MBP175-cpGFP.L1-HL (A and B) and MBP311-cpGFP.L2-NP (C and D) bound to maltose. The MBP domain is colored as in FIG. 1. The cpGFP domain is green and the inter-domain linkers are colored white. The cpGFP chromophore is displayed as sticks and the bound maltose as red and white spheres. Ordered water molecules are represented as red spheres. Selected hydrogen bonds are displayed as dashed black lines. .beta.-strands 10 and 11 of cpGFP are displayed as semi-transparent for clarity. The 2Fo-Fc electron density map calculated with the displayed residues omitted from the model is shown as blue mesh.
[0057] FIGS. 23A-23D|EcMBP-cpGFP: effect of T203V mutation on fluorescence. (a) Emission spectra of 1 .mu.M purified eGFP (filled circles), cpGFP (filled squares), MBP165-cpGFP.PPYF (open circles), and MBP165-cpGFP.PPYF+T203V (open squares) in the absence (dashed lines) or presence (solid lines) of 1 mM maltose. cpGFP is half as bright as eGFP, and the saturated MBP165-cpGFP.PPYF variants are about half as bright as cpGFP. (b) Titration of maltose for MBP165-cpGFP.PPYF (filled squares), and MBP165-cpGFP.PPYF+T203V (filled circles). Affinities for each protein are the same, but with different .DELTA.F/F. (c) Emission spectra of 1 .mu.M purified eGFP (filled circles), cpGFP (filled squares), MBP311-cpGFP.L2-NP (open circles), and MBP311-cpGFP.L2-NP+T203V (open squares) in the absence (dashed lines) or presence (solid lines) of 1 mM maltose. Note that mutation T203V decreases the fluorescence of both the apo-state and the saturated state of MBP311-cpGFP.L2-NP. (d) Titration of maltose for MBP311-cpGFP.L2-NP (filled squares), and MBP311-cpGFP.L2-NP+T203V (filled circles). Affinities for each protein are the same, but with .DELTA.F/F slightly increased for the T203V variant.
[0058] FIG. 24A|Amino acid sequence of PfMBP171-cpGFP (SEQ ID NO:50)
[0059] FIG. 24B|Amino acid sequence of PfMBP171cpGFP.L2-FE (SEQ ID NO:51)
[0060] FIG. 25A|Amino acid sequence of PfMBP316-cpGFP (SEQ ID NO:52)
[0061] FIG. 25B|Amino acid sequence of PfMBP316-cpGFP.L1-NP (SEQ ID NO:53)
[0062] FIG. 26A-26B|Plot of .DELTA.F/F for clarified lysate screen of cpGFP linker-screens at insertion points 171 (A) and 316 (B).
[0063] FIGS. 27A-27D|Plot of Beta-sheet circular dichroism (CD) signal as a function of temperature.
[0064] FIGS. 28A-28B|PfMBP Fluorescence vs. temperature. (A) Plot of fluorescence as a function of temperature in the presence (solid) or absence (dashed) of ligand. (B) Plot of .DELTA.F/F as a function of temperature. Using the data from panel (a), .DELTA.F/F for each protein (Fbound-Fapo/Fapo) was calculated for each temperature.
[0065] FIGS. 28C-28E|Line graphs showing the function of immobilized and soluble proteins.
[0066] FIG. 29A|Amino acid sequence of EcYbeJ253-cpGFP (SEQ ID NO:62).
[0067] FIG. 29B|Amino acid sequence of EcYbeJ253-cpGFP.L1LVL2NP (SEQ ID NO:63).
[0068] FIG. 30|EcYbeJ binding curves. Plot of .DELTA.F/F as a function of [Glutamate], M. The first generation sensor, EcYbeJ253.L1-LV (with the A184V) mutation (grey, solid) has an affinity for glutamate of about 100 .mu.M and a .DELTA.F/F of 1.2. The reversion of that affinity mutation, V184A, in the L1-LV background increases affinity to 1 .mu.M (grey dashed). The second generation sensor, with the L2-NP linker optimization and the A184V mutation, has a .DELTA.F/F of at least 4 and an affinity for glutamate of about 100 .mu.M (black solid).
[0069] FIG. 31|EcYbeJ Hema/cMyc analysis. The effect of N- and C-terminal tags on .DELTA.F/F and glutamate affinity were determined by expressing variously tagged versions of the EcYbeJ253.L1LVL2NP protein in bacteria. The presence of the pRSET leader sequence (black) has no effect on .DELTA.F/F (.about.5) or affinity (.about.120 .mu.M), when compared to the version without a tag (grey). The addition of the cMyc tag to the C-terminus retains .DELTA.F/F and increases affinity slightly, to 60 .mu.M. The addition of the N-terminal hemagglutinin tag, with (green) or without (orange) the cMyc tag, decreases .DELTA.F/F substantially.
[0070] FIGS. 32A-32B|EcYbeJ253-cpGFP.L1LVL2NP.pMinDis expressed in HEK293 cells. (A) Images of the sensor expressing HEK cells in the absence of glutamate (left), with 100 .mu.M glutamate (center), and re-imaged after wash-out of glutamate with buffer (right). (B) By measuring the equilibrium .DELTA.F/F with different concentrations of glutamate in the buffer, an in situ binding affinity (black) can be obtained. The surface displayed sensor has a higher affinity (3 .mu.M) for glutamate than the soluble sensor (grey), which is about 90 .mu.M.
[0071] FIG. 33|EcYbeJ253-cpGFP.L1LVL2NP.pMinDis expressed in neuronal culture, and responds rapidly to added glutamate (green). Red shows signal of 2.5 nM Alexa Fluor.RTM. 568 (Invitrogen, Carlsbad, Calif.), also in pipette.
[0072] FIG. 34A|Amino acid sequence of EcPhnD90-cpGFP (SEQ ID NO:77).
[0073] FIG. 34B|Amino acid sequence of EcPhnD90-cpGFP.L1AD+L297R+L301R (SEQ ID NO: 78).
[0074] FIGS. 35A-35C|EcPhnD90-cpGFP Binding Curves. For both the L1AD and the L1AD+L297R+L301R variants, binding was determined for (A) 2-aminoethylphosphonate (2AEP), (B) methylphosphonate (MP), and (C) ethylphosphonate (EP).
[0075] FIGS. 36A-36C|The crystal structures of the ligand-free (A), open state (with H157A mutation to the binding pocket) and the ligand-bound (B), closed state of EcPhnD clearly shows a large conformational change. Residues in between which cpGFP is inserted in EcPhnD90-cpGFP are marked by red spheres, in the equatorial strand (red). (C) Analysis of the change in C.alpha. dihedral (.DELTA.Dihedral) clearly shows that residues for which there is the greatest .DELTA.Dihedral upon going from the open to the closed state are residues 88 (.DELTA.Dihedral=-75.degree.), 89 (.DELTA.Dihedral=123.degree.), and 90 (.DELTA.Dihedral=52.degree.).
[0076] FIG. 37A|Amino acid sequence of TtGBP326-cpGFP (SEQ ID NO:91).
[0077] FIG. 37B|Amino acid sequence of TtGBP326.L1-PA (SEQ ID NO:92).
[0078] FIG. 37C|Amino acid sequence of TtGBP326.H66A (SEQ ID NO:93).
[0079] FIG. 37D|Amino acid sequence of TtGBP326.H348A (SEQ ID NO:94).
[0080] FIG. 38|TtGBP326-cpGFP Binding Curves. Plot of .DELTA.F/F as a function of [Glucose], mM.
[0081] FIG. 39|An image showing TtGBP326-cpGFP expressed as a transgenic reporter of intracellular glucose in cultured human cells.
[0082] FIGS. 40A-40B|Are line graphs showing that the addition of extracellular glucose increases TtGBP326-cpGFP fluorescence in human cells.
[0083] FIG. 41|Amino acid sequence of Escherichia coli maltodextrin-binding protein (EcMBP) (SEQ ID NO: 105).
[0084] FIG. 42|Amino acid sequence of Pyrococcus furiosus maltose-binding protein (PfMBP) (SEQ ID NO: 106).
[0085] FIG. 43|Amino acid sequence of E. coli glutamate-binding protein (EcYbeJ) (SEQ ID NO:107).
[0086] FIG. 44|Amino acid sequence of E. coli phosphonate-binding protein (EcPhnD) (SEQ ID NO:108).
[0087] FIG. 45|Amino acid sequence of Thermus thermophilus glucose-binding protein (TtGBP) (SEQ ID NO:109).
[0088] FIG. 46|Amino acid sequence of UniProt accession number Q92N37 (SEQ ID NO: 110).
[0089] FIG. 47|Amino acid sequence of UniProt accession number DOVWX8 (SEQ ID NO:111).
[0090] FIG. 48|Amino acid sequence of UniProt accession number Q7CX36 (SEQ ID NO:112).
[0091] FIG. 49|Amino acid sequence of UniProt accession number POAD96 (SEQ ID NO:113).
[0092] FIG. 50|Amino acid sequence of TtGBP326.L1PA.L2NP.H66A.H348A.L276V (SEQ ID NO: 114).
[0093] FIG. 51|A line graph showing binding of TtGBP326.L1PA.L2NP.H66A.H348A.L276V to glucose.
[0094] FIG. 52|A line graph showing fluorescence increase upon addition of glucose to HEK293 cells expressing TtGBP326.LIPA.L2NP.H66A.H348A.L276V on their extracellular surface.
[0095] FIG. 53|A schematic of Structure I as described herein.
DETAILED DESCRIPTION
[0096] The present disclosure is based, at least in part, on the discovery of structures and methods related to and useful for genetically encoded biosensors. Specifically, the disclosure provides genetically encoded recombinant or chimeric peptides for use as biosensors and methods for the design, production, and use of such biosensors. As described below, these sensors can be employed (e.g., expressed) in biological systems to detect and/or monitor a wide range of target analytes (e.g., a defined, selected, and/or specific analytes) due, in part, to the signal change generated by the sensors upon binding to their respective analyte(s), which signal change allows bound and unbound sensors to be distinguished.
[0097] While the disclosure encompasses generic biosensors and methods related thereto, examples of particular binding sensors, including biosensors for detecting maltose, sucrose, maltotriose, glutamate, phosphonate, and glucose are also disclosed.
Compositions
[0098] Provided herein are genetically encoded biosensors, i.e., nucleic acids encoding peptides, and/or the encoded peptides (e.g., isolated peptides), for use as biosensors. Biosensors herein include genetically encoded recombinant peptides containing an analyte-binding framework portion linked (e.g., operably linked) to at least one independent signaling portion, wherein the independent signaling portion is allosterically modulated or regulated by the framework portion upon interaction of the framework portion with an analyte (e.g., a defined, selected, and/or specific analyte), such that signaling from the signaling portions is altered upon interaction of the framework portion with the analyte.
[0099] In some instances, an independent signaling portion is present at a site within the framework portion that undergoes a conformational change upon interaction of the framework portion with an analyte such that the conformational change allosterically modulates or regulates signaling by the signaling portion. For example, biosensors herein can include structure I (FIG. 53). As described herein, the signaling portion is present at a site within the framework portion that undergoes a conformational change upon interaction of the framework portion with an analyte.
[0100] In some instances, signaling by the signaling portion is detectably altered upon interaction (e.g., binding) of the framework portion with an analyte. For example, signaling by the signaling portion can detectably increase or detectably decrease upon interaction (e.g., binding) of the framework portion with an analyte. In some cases, biosensors have a signal change upon binding (e.g., specific binding) to their respective analyte of at least about, for example, +0.5, and/or an increase or decrease in signal of at least about, for example, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, 250%, 500%, 750%, 1000%, or more than 1000%, e.g., relative to unbound biosensor. In some increases, the level of signal change is linked to background signal. Values represented here can be converted and/or expressed into any conventional units using ordinary skill. For example, units can be expressed as `signal change` (as used above), .DELTA.F/F and/or as signal-to-noise ratio (e.g., .DELTA.F/F multiplied by the square root of the number of photons collected). In some instances, signaling by a biosensor can be intensity based.
[0101] In some instances, biosensors herein are distinguishable from Forster resonance energy transfer, also known as fluorescence resonance energy transfer (FRET)-based sensors, which require donor and acceptor chromophores, e.g., that function in concert, in that they include independently functioning or detectable signaling portions. For example, in some instances, signaling by a first signaling portion of a biosensor herein is independent of signaling by a second signaling portion within the same or a distinct biosensor. As noted above, signaling portions are allosterically regulated by the framework portion to which they are linked upon interaction of the framework portion with an analyte (e.g., a defined, selected, and/or specific analyte).
[0102] Framework Portions
[0103] Framework portions include genetically encoded macromolecules (e.g., proteins or peptides) that undergo conformational alteration (e.g., a structural change) upon interaction (e.g., binding) with, or to, an analyte (e.g., an analyte-binding dependent conformational alteration). For example, genetically encoded framework portions can have a first structure in the absence of an analyte (e.g., in an unbound or open state) and a second structure, that is detectably distinct (e.g., differences in structures before and after a conformational change can be observed using methods known in the art) from the first structure, in the presence of an analyte (e.g., in a bound or closed state), e.g., under physiologic conditions. In some instances, the conformational change that occurs upon interaction with an analyte (e.g., an analyte-binding dependent conformational alteration) is detectably distinct (e.g., can be observed using methods known in the art) from a conformational change that may occur for the same protein or peptide under other physiological conditions (e.g., a change in conformation induced by altered temperature, pH, voltage, ion concentration, phosphorylation).
[0104] Methods for identifying proteins or peptides that exhibit suitable conformational characteristics and/or for observing differences in structure between structures or before and after a conformational change are known in the art and/or are described herein. Such methods can include, for example, one or more of structural analysis, crystallography, NMR, EPR using Spin label techniques, Circular Dichroism (CD), Hydrogen Exchange surface Plasmon resonance, calorimetry, and/or FRET.
[0105] In some instances, framework portions can have a first structure in the absence of an analyte (e.g., in an unbound or open state) and a second structure, that is detectably distinct (e.g., can be observed using methods known in the art) from the first structure, in the presence of an analyte (e.g., in a bound or closed state), e.g., under physiologic conditions, wherein the structural change between the open and closed state can allosterically modulate an independent signaling portion recombinantly (e.g., artificially introduced) present within the framework portion (see, e.g., Structure I in FIG. 53).
[0106] Framework portions can also interact (e.g., bind) with at least one analyte (e.g., at least one defined, specific, and/or selected analyte). In some instances, a framework portion can interact specifically with one analyte (e.g., at least one defined, specific, and/or selected analyte). In such cases, affinity of binding between the framework binding peptide and the analyte can be high or can be controlled (e.g., with millimolar, micromolar, nanomolar, or picomolar affinity). Alternatively, the single framework binding protein can bind two or more analytes (e.g., two or more defined, specific, and/or selected analytes). In such cases, affinity of binding to the two or more analytes can be the same or distinct. For example, the affinity of binding can be greater for one analyte than it is for a second or third, etc., analyte. In some instances, binding between a framework portion and an analyte (e.g., at least one defined, specific, and/or selected analyte) have an affinity of for example, 10 mM to 1 pM.
[0107] As used herein, the term "analyte" can include naturally occurring and/or synthetic sugars, amino acids, proteins (e.g., proteins, peptides, and/or antibodies), hormones, ligands, chemicals (e.g., small molecules), pharmaceuticals, nucleic acids, cells, tissues, and combinations thereof.
[0108] In some instances, biosensors can include one, two, or more framework binding portions that bind (e.g., binds specifically) a single analyte (e.g., a single defined, specific, and/or selected analyte) or distinct analytes (e.g., two or more distinct defined, specific and/or selected analytes). Alternatively or in addition, the framework portion can be chimeric. In such cases, a first part of the framework portion can be a first peptide or can be derived from a first peptide, and a second part of the framework portion can be a second peptide or can be derived from a second peptide, wherein the first a second peptides are combined to result in a single peptide.
[0109] Accordingly, framework portions can include macromolecules that undergo a conformational change upon interaction with an analyte. One non-limiting example of a suitable macromolecule is Calmodulin (CaM). CaM is in an extended shape in the absence of Ca.sup.2+ and in a condensed conformation in the presence of Ca.sup.2+ (Kuboniwa et al., Nat. Struc. Biol., 2:768-776, 1996 and Fallon and Quiocho, Structure, 11:1303-1307, 2003).
[0110] In some instances, a framework binding portion can be a bacterial protein or can be derived from a bacterial protein. Suitable bacterial proteins can include, but are not limited to, for example, periplasmic binding proteins (PBPs).
[0111] PBPs from bacteria are generally useful in the biosensors herein at least because they undergo dramatic conformational changes upon ligand binding (Ouiocho et al. Mol. Microbiol., 20:17-225, 1996). X-ray crystal structures of the apo (open) and bound (closed) forms of various PBPs reveal that these proteins have two (typically, although some have more) domains that undergo a large hinge-twist movement relative to each other in a Venus flytrap manner (Dwyer and Hellinga, Curr. Opin. Struc. Biol., 14:495-504, 2004). This conformational change has been exploited to create a number of FRET-based genetically encoded sensors (see, e.g., Deuschle et al., Pro. Sci, 14:2304-2314, 2005; Deuschle et al., Cytometry, 64:3-9, 2005; Okumoto et al., Proc. Natl. Acad. Sci. USA., 102:8740-8745, 2005; Bogner and Ludewig, J. Fluoresc., 17:350-360, 2007; and Gu et al., FEBS Letters, 580:5885-5893, 2006). In addition, the ligand-binding diversity of the PBP superfamily is large (Dwyer and Hellinga, Curr. Opin. Struc. Biol., 14:495-504, 2004).
[0112] In some instances, framework portions can include, for example, one or more of: arabinose-binding protein(s), glucose/galactose-binding protein(s), histidine-binding protein(s), maltose-binding protein(s), glutamine-binding protein(s), maltotriose-binding protein(s), RBP, ribose-binding protein(s), acetylcholine binding protein(s), choline binding protein(s), lysine binding protein(s), arginine binding protein(s), gamma aminobutyric acid (GABA) binding protein(s), ion-binding protein(s), peptide-binding protein(s), lactate-binding protein(s), histamine-binding protein(s), and/or Leucine/Isoleucine/Valine binding protein(s), including full length proteins, fragments, and/or variants thereof.
[0113] In some instances, exemplary framework portions can include: SEQ ID NO: 105, which is Escherichia coli maltodextrin-binding protein (EcMBP) (UniProt accession number POAEX9); SEQ ID NO: 106, which is Pyrococcus Furiosus maltotriose-binding protein (PfMBP) (UniProt accession number P58300); SEQ ID NO: 107, which is E. coli glutamate-binding protein (EcYbeJ) (UniProt accession number Q1R3F7); SEQ ID NO: 108, which is E. coli phosphonate-binding protein (EcPhnD) (UniProt accession number P37902); and/or SEQ ID NO: 109, which is Thermus thermophilus glucose-binding protein (TtGBP) (UniProt accession number Q72KX2, including full length proteins, fragments, and/or variants thereof.
[0114] In some instances, exemplary framework portions can include SEQ ID NO: 110 (UniProt accession number Q92N37); SEQ ID NO: 111 (UniProt accession number DOVWx8, SEQ ID NO:112 (UniProt accession number Q7CX36), and/or SEQ ID NO: 113 (UniProt accession number POAD96, including full length proteins, fragments, and/or variants thereof.
[0115] In some instances, framework portions, or biosensors, do not include signal peptides, or portions of signal peptides, that would otherwise be present in the peptide from which the framework portion is derived.
[0116] Signaling Portions
[0117] Biosensors herein include one or more genetically encoded signaling portions (e.g., independent signaling portions) within the amino acid sequence of a framework portion at a site(s) within the framework portion that undergo(es) a conformational change upon interaction of the framework portion with an analyte (e.g., a defined, specific, and/or selected analyte).
[0118] Signaling portions (e.g., independent signaling portions) include genetically encoded molecules (e.g., peptides or proteins) that can be allosterically induced to emit a detectable signal (e.g., an analyte-binding dependent signal).
[0119] In some instances, the detectable signal is detectably distinct (e.g., can be distinguished using methods known in the art and/or disclosed herein) from a signal emitted by the molecule prior to allosteric inducement (e.g., signaling portions can emit a detectable signal in two detectably distinct states. For example, first signal can be emitted in unbound state and a second signal can be emitted in bound state). As noted above, in some instances, the detectable signal is proportional to the degree of allosteric inducement. In some instances, if two or more signaling portions are present in a biosensor, then two or more detectably distinct signals can be emitted by the biosensor.
[0120] In some instances, a genetically encoded independent signaling portion is a genetically encoded fluorescent protein (FP), e.g., a macromolecule containing a functional group (e.g., a fluorophore) that absorbs energy of a specific wavelength and re-emits energy at a different (but equally specific) wavelength, including, for example, circularly permuted FP (cpFP).
[0121] As used herein, the term "fluorophore" relates to a functional group in a molecule which will absorb energy of a specific wavelength and re-emit energy at a different (but equally specific) wavelength. In some instances, fluorophore containing molecules include fluorescent proteins. The fluorophore in green fluorescent protein (GFP) includes Ser-Tyr-Gly sequence (i.e., Ser65-dehydroTyr66-Gly67), which is post-translationally modified to a 4-(p-hydroxybenzylidene)-imidazolidin-5. Exemplary genetically encoded fluorescent proteins include, but are not limited to, fluorescent proteins from coelenterate marine organisms, e.g., Aequorea victoria, Trachyphyllia geoffroyi, coral of the Discosoma genus, Rennilla mulleri, Anemonia sulcata, Heteractis crispa, Entacmaea quadricolor, and/or GFP (including the variants S65T and EGFP, Rennilla mulleri GFP), cyan fluorescent protein (CFP), including Cerulean, and mCerulean3 (described by Markwardt et al., PLoS ONE, 6(3) e17896.doi:10.1371/journal.pone.0017896), CGFP (CFP with Thr203Tyr: Has an excitation and emission wavelength that is intermediate between CFP and EGFP), yellow fluorescent protein (YFP, e.g., GFP-Ser65Gly/Ser72Ala/Thr203Tyr; YFP (e.g., GFP-Ser65Gly/Ser72Ala/Thr203Tyr) with Val68Leu/Gln69Lys); Citrine (i.e., YFP-Val68Leu/Gln69Met), Venus (i.e., YFP-Phe46Leu/Phe64Leu/Met153Thr/Val163Ala/Ser175Gly), PA-GFP (i.e., GFP-Val/163Ala/Thr203His), Kaede), red fluorescent protein (RFP, e.g., long wavelength fluorescent protein, e.g., DsRed (DsRed1, DsRed2, DsRed-Express, mRFP1, drFP583, dsFP593, asFP595), eqFP611, and/or other fluorescent proteins known in the art (see, e.g., Zhang et al., Nature Reviews, Molecular and Cellular Biology, 3:906-908, 2002).
[0122] As set forth above, in some instances, fluorophore containing molecules include fluorescent proteins that can be or that are circularly permutated. Circular permutation methods are known in the art (see, e.g., Baird et al., Proc. Natl. Acad. Sci., 96:11241-11246, 1999; Topell and Glockshuber, Methods in Molecular Biology, 183:31-48, 2002).
[0123] In some instances, single-FP sensors have a number of advantages: they preserve spectral bandwidth for multi-analyte imaging; their saturated states may be nearly as bright as the parental FP, and their ligand-free states may be arbitrarily dim, providing large theoretical fluorescence increases. This allows for much greater changes in fluorescence and thus increased signal-to-noise ratios and greater resistance to photobleaching artifacts (Tian et al., Nat. Methods, 6:875-881, 2009).
[0124] In some instances, issues arising from long-term effects such as gene regulation and protein expression and degradation can be identified by simply fusing the intensity-based sensor to a another fluorescent protein of different color, to serve as a reference channel.
[0125] In some instances, biosensors can include circularly permuted YFP (cpYFP) as a cpFP. cpYFP has been used as a reporter element in the creation of sensors for H2O2 (HyPer) (Belousov et al., Nat. Methods, 3:281-286, 2006), cGMP (FlincG) (Nausch et al., Proc. Natl. Acad. Sci. USA., 105:365-370, 2008), ATP:ADP ratio (Perceval) (Berg et al., Nat. Methods., 105:365-370, 2008), and calcium ions (Nakai et al., Nat. Biotechno., 19:137-141, 2001), including full length, fragments, and/or variants thereof.
[0126] Linker Portions
[0127] As shown in Structure I (FIG. 53), biosensors herein can optionally include one or more genetically encoded linkers positioned between or operably linking the framework portion and the signaling portion. Linker portions can include at least one naturally occurring or synthetic amino acid (discussed below) as exemplified by SEQ ID NOs: 9-49, 54-61, 64-76, 79-90, 95-104. In some instances, linker can include one or more of SEQ ID NOs: 9-49, 54-61, 64-76, 79-90, 95-104, and/or portions of SEQ ID NOs: 9-49, 54-61, 64-76, 79-90, 95-104. For example, linkers can include, but are not limited to, one or more of: PxSHNVY (SEQ ID NO:114), xPSHNVY (SEQ ID NO:115), xxSHNVY (SEQ ID NO:116), xxSHNVF (SEQ ID NO:117), PxSHNVF (SEQ ID NO:118), PxSYNVF (SEQ ID NO: 119), xxSYNVF (SEQ ID NO: 120), PxSYNVF (SEQ ID NO:121), xxSYNVF (SEQ ID NO:122), PxSxNVY (SEQ ID NO:123), PxSHxVY (SEQ ID NO: 124), PxSHNxY (SEQ ID NO: 125), PxSHNVx (SEQ ID NO: 126), FNxxY (SEQ ID NO: 127), FNxY (SEQ ID NO: 128), FNY (SEQ ID NO: 129), FxY (SEQ ID NO: 130), xxY (SEQ ID NO:131), WxY (SEQ ID NO: 132), xKY, (SEQ ID NO: 133), FNPxY (SEQ ID NO:134), FNxPY (SEQ ID NO:135), HNS (SEQ ID NO:136), GGS (SEQ ID NO: 137), xxS (SEQ ID NO:138), xxK (SEQ ID NO: 139), GGK (SEQ ID NO:140), PXS (SEQ ID NO:141), xPS (SEQ ID NO:142), Px (SEQ ID NO:143), xP (SEQ ID NO:144), IxxS (SEQ ID NO:145), NxPK (SEQ ID NO:146), NPcK (SEQ ID NO: 147), PPxSH (SEQ ID NO: 148), PPxxSH (SEQ ID NO: 149), PPPxSH (SEQ ID NO: 150), PPxPSH (SEQ ID NO: 151), xxSH (SEQ ID NO: 152), PPxx (SEQ ID NO: 153), FNxKN (SEQ ID NO:154), FNxxKN (SEQ ID NO:155), FNxPKN (SEQ ID NO:156), FNPxKN (SEQ ID NO:157), FNxx (SEQ ID NO:158), N, ADGSSH (SEQ ID NO:159), ADxxSH (SEQ ID NO:160), ADxPSH (SEQ ID NO:161), ADPxSH (SEQ ID NO:162), ADxx (SEQ ID NO: 163), ADxxSH (SEQ ID NO: 164), FNPG (SEQ ID NO: 165), FNxxPG (SEQ ID NO:166), xxPG (SEQ ID NO:167), FNxx (SEQ ID NO:168), FNPx (SEQ ID NO: 169), KYxxSH (SEQ ID NO: 170), KYPxSH (SEQ ID NO: 171), KYxPSH (SEQ ID NO: 172), FxxP (SEQ ID NO: 173), FNxP (SEQ ID NO: 174), and/or FNPx (SEQ ID NO: 175), where "x" indicates any amino acid.
[0128] Exemplary Biosensor Constructs
[0129] As noted above, biosensors herein include genetically encoded biosensors, i.e., nucleic acids encoding biosensors, and/or the encoded biosensors (e.g., isolated biosensors), for use as biosensors. In some instances, nucleic acids encoding biosensors include isolated nucleic acids. In some instances, the portion of a nucleic acid encoding a biosensor can include a single reading frame encoding the biosensor. For example, a biosensor can be encoded by a portion of a nucleic acid that falls within a start codon and a stop codon. In some instances, biosensors are isolated (e.g., biosensors are substantially free of contaminating and/or non-biosensor components).
[0130] In some instances, biosensors can include, for example, one or more framework portions selected from the group consisting of: arabinose-binding protein(s), glucose/galactose-binding protein(s), histidine-binding protein(s), maltose-binding protein(s), maltotriose-binding protein(s), glutamine-binding protein(s), RBP, ribose-binding protein(s), acetylcholine binding protein(s), choline binding protein(s), lysine binding protein(s), arginine binding protein(s), gamma aminobutyric acid (GABA) binding protein(s), ion-binding protein(s), peptide-binding protein(s), lactate-binding protein(s), histamine-binding protein(s), and/or Leucine/Isoleucine/Valine binding protein(s), including full length proteins, fragments, and/or variants thereof, including full length proteins, fragments and/or variants thereof, and at least one independent signaling portion present at a site within the framework portion that undergoes a conformational change upon interaction of the framework portion with an analyte.
[0131] In some instances, biosensors can include, for example, one or more framework portions selected from the group consisting of: SEQ ID NO: 105, which is Escherichia coli maltodextrin-binding protein (EcMBP) (UniProt accession number POAEX9); SEQ ID NO: 106, which is Pyrococcus Furiosus maltose-binding protein (PfMBP) (UniProt accession number P58300); SEQ ID NO: 107, which is E. coli glutamate-binding protein (EcYbeJ) (UniProt accession number Q1R3F7); SEQ ID NO: 108, which is E. coli phosphonate-binding protein (EcPhnD) (UniProt accession number P37902); and/or SEQ ID NO: 109, which is Thermus thermophilus glucose-binding protein (TtGBP) (UniProt accession number Q72KX2), including full length proteins, fragments and/or variants thereof, and at least one independent signaling portion present at a site within the framework portion that undergoes a conformational change upon interaction of the framework portion with an analyte.
[0132] In some instances, biosensors can include, for example, one or more framework portions selected from the group consisting of: SEQ ID NO: 110 (UniProt accession number Q92N37); SEQ ID NO:111 (UniProt accession number DOVWx8, SEQ ID NO: 112 (UniProt accession number Q7CX36), and/or SEQ ID NO: 113 (UniProt accession number POAD96), including full length proteins, fragments and/or variants thereof, and at least one independent signaling portion present at a site within the framework portion that undergoes a conformational change upon interaction of the framework portion with an analyte.
[0133] In some instances, biosensors include any one or more:
[0134] Maltose biosensors SEQ ID NOs: 1-8 (i.e., Escherichia coli maltodextrin-binding protein (EcMBP)) or SEQ ID NOs: 50-53 (Pyrococcus Furiosus maltose-binding protein (PfMBP)), including full length proteins, fragments and/or variants thereof;
[0135] Glutamate biosensors SEQ ID NOs: 62-63 (E. coli glutamate-binding protein (EcYbeJ)), including full length proteins, fragments and/or variants thereof;
[0136] Phosphonate biosensors SEQ ID NOs: 77-78 (E. coli phosphonate-binding protein (EcPhnD)), including full length proteins, fragments and/or variants thereof; and/or
[0137] Glucose biosensors SEQ ID NOs: 91-94 (Thermus thermophilus glucose-binding protein (TtGBP)), including full length proteins, fragments and/or variants thereof.
[0138] In some instances, nucleic acids encoding and/or amino acid sequences of any of the framework portions, signaling portions, linker portions, or biosensors (e.g., SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) (e.g., any amino acid sequence) disclosed herein can be modified to generate fragments (e.g., truncated peptides) and/or variants (e.g., peptides with a defined sequence homology to the peptides disclosed herein). Variants can include framework portions, signaling portions, linker portions, or biosensors with amino acid sequences with homology to the framework portions, signaling portions, linker portions, or biosensors disclosed herein and/or truncated forms of the framework portions, signaling portions, linker portions, or biosensors herein. In some instances, truncated forms of the framework portions, signaling portions, linker portions, or biosensors herein can include 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 50-100, 101-150, fewer amino acids than the framework portions, signaling portions, linker portions, and/or biosensors herein, e.g., wherein the truncated biosensor variants retain at least at portion of the binding and/or signaling properties of same biosensor without truncation (e.g., at least 50%, 60%, 70%, 80%, 90%, or 100% of the binding and/or signaling properties of the same biosensor without truncation). In addition, truncations can be made at the amino-terminus, the carboxy-terminus, and/or within the body of the framework portions, signaling portions, linker portions, and/or biosensors herein.
[0139] While variants are generally observed and discussed at the amino acid level, the actual modifications are typically introduced or performed at the nucleic acid level. For example, variants with 95%, 96%, 97%, 98, or 99% sequence identity to SEQ ID NOs:91, 92, 93, and/or 94 can be generated by modifying the nucleic acids encoding SEQ ID NOs: 91, 92, 93, and/or 94 using techniques (e.g., cloning techniques) known in the art and/or that are disclosed herein.
[0140] As with all peptides, polypeptides, and proteins, including fragments thereof, it is understood that modifications to the amino acid sequence can occur that do not alter the nature or function of the peptides, polypeptides, or proteins. Such modifications include conservative amino acids substitutions and are discussed in greater detail below.
[0141] The peptides, polypeptides, and proteins, including fragments thereof, provided herein are biosensors whose activity can be tested or verified, for example, using the in vitro and/or in vivo assays described herein.
[0142] In some instances, any of the framework portions, signaling portions, or biosensors (e.g., SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) (e.g., any amino acid sequence) described herein can be modified and varied so long as their desired function is maintained. For example, the polypeptides can be modified as long as the resulting variant polypeptides have the same or better characteristics as the polypeptide from which they derived. For example, the variants can have the same or better affinity for their respective analyte.
[0143] In some instances, the interacting face of a modified peptide can be the same (e.g., substantially the same) as an unmodified peptide (methods for identifying the interacting face of a peptide are known in the art (Gong et al., BMC: Bioinformatics, 6:1471-2105 (2007); Andrade and Wei et al., Pure and Appl. Chem., 64(11):1777-1781 (1992); Choi et al., Proteins: Structure, Function, and Bioinformatics, 77(1):14-25 (2009); Park et al., BMC: and Bioinformatics, 10:1471-2105 (2009)), e.g., to maintain binding to an analyte. Alternatively, amino acids within the interacting face can be modified, e.g., to decrease binding to an analyte and/or to change analyte specificity.
[0144] The interacting face of a peptide is the region of the peptide that interacts or associates with other molecules (e.g., other proteins). Generally, amino acids within the interacting face are naturally more highly conserved than those amino acids located outside the interacting face or interface regions of a protein. In some instances, an amino acid within the interacting face region of any of the framework portions or biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) (e.g., any amino acid sequence) disclosed herein can be the same as the amino acid shown in any of the framework portions or biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) (e.g., any amino acid sequence) disclosed herein or can be include conservative amino acid substitutions. In some instances, an amino acid within the interacting face region any of the framework portions or biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) (e.g., any amino acid sequence) disclosed herein can be substituted with an amino acid that increases the interaction between the framework portion or biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) (e.g., any amino acid sequence) and an analyte.
[0145] In some instances, genetically encoded biosensors can include peptides that have at least 80, 85, 90, 95, 96, 97, 98, 99 percent identity to the framework portions, signaling portions, or biosensors (e.g., SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) (e.g., any amino acid sequence) described herein. Those of skill in the art readily understand how to determine the identity of two polypeptides. For example, the identity can be calculated after aligning the two sequences so that the identity is at its highest level.
[0146] Another way of calculating identity can be performed by published algorithms. Optimal alignment of sequences for comparison may be conducted by the local identity algorithm of Smith and Waterman, Adv. Appl. Math, 2:482 (1981), by the identity alignment algorithm of Needleman and Wunsch, J. Mol. Biol. 48:443 (1970), by the search for similarity method of Pearson and Lipman, Proc. Natl. Acad. Sci. USA 85:2444 (1988), by computerized implementations of these algorithms (GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package, Genetics Computer Group, 575 Science Dr., Madison, Wis.), or by inspection.
[0147] The same types of identity can be obtained for nucleic acids by, for example, the algorithms disclosed in Zuker, Science 244:48-52 (1989); Jaeger et al., Proc. Natl. Acad. Sci. USA 86:7706-10 (1989); Jaeger et al., Methods Enzymol. 183:281-306 (1989), which are herein incorporated by reference for at least material related to nucleic acid alignment. It is understood that any of the methods typically can be used and that in certain instances the results of these various methods may differ, but the skilled artisan understands if identity is found with at least one of these methods, the sequences would be said to have the stated identity and to be disclosed herein.
[0148] Amino acid sequence modifications typically fall into one or more of three classes: substitutional, insertional, or deletional modifications. Insertions include amino and/or terminal fusions as well as intra-sequence insertions of single or multiple amino acid residues. Insertions ordinarily will be smaller insertions than those of amino or carboxyl terminal fusions, for example, on the order of one to four residues. Deletions are characterized by the removal of one or more amino acid residues from the protein sequence. Typically, no more than about from 2 to 6 residues are deleted at any one site within the protein molecule. Amino acid substitutions are typically of single residues, but can occur at a number of different locations at once; insertions usually will be on the order of about from 1 to 10 amino acid residues; and deletions will range about from 1 to 30 residues. Deletions or insertions can be made in adjacent pairs, i.e., a deletion of 2 residues or insertion of 2 residues. Substitutions, deletions, insertions or any combination thereof may be combined to arrive at a final construct. The mutations must not place the sequence out of reading frame and preferably will not create complementary regions that could produce secondary mRNA structure. Substitutional modifications are those in which at least one residue has been removed and a different residue inserted in its place. In some instances, substitutions can be conservative amino acid substitutions. In some instances, variants herein can include one or more conservative amino acid substitutions. For example, variants can include 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 20-30, 30-40, or 40-50 conservative amino acid substitutions. Alternatively, variants can include 50 or fewer, 40 or fewer, 30 or fewer, 20 or fewer, 10 or fewer, 9 or fewer, 8 or fewer, 7 or fewer, 6 or fewer, 5 or fewer, 4 or fewer, 3 or fewer, or 2 or fewer conservative amino acid substitutions. Such substitutions generally are made in accordance with the following Table 1 and are referred to as conservative substitutions. Methods for predicting tolerance to protein modification are known in the art (see, e.g., Guo et al., Proc. Natl. Acad. Sci., USA, 101(25):9205-9210 (2004)).
TABLE-US-00001 TABLE 1 Conservative Amino Acid Substitutions Amino Acid Substitutions (others are known in the art) Ala Ser, Gly, Cys Arg Lys, Gln, His Asn Gln, His, Glu, Asp Asp Glu, Asn, Gln Cys Ser, Met, Thr Gln Asn, Lys, Glu, Asp, Arg Glu Asp, Asn, Gln Gly Pro, Ala, Ser His Asn, Gln, Lys Ile Leu, Val, Met, Ala Leu Ile, Val, Met, Ala Lys Arg, Gln, His Met Leu, Ile, Val, Ala, Phe Phe Met, Leu, Tyr, Trp, His Ser Thr, Cys, Ala Thr Ser, Val, Ala Trp Tyr, Phe Tyr Trp, Phe, His Val Ile, Leu, Met, Ala, Thr
[0149] In some instances, substitutions are not conservative. For example, an amino acid can be replaced with an amino acid that can alter some property or aspect of the peptide. In some instances, non-conservative amino acid substitutions can be made, e.g., to change the structure of a peptide, to change the binding properties of a peptide (e.g., to increase or decrease the affinity of binding of the peptide to an analyte and/or to alter increase or decrease the binding specificity of the peptide).
[0150] Modifications, including the specific amino acid substitutions, are made by known methods. By way of example, modifications are made by site-specific mutagenesis of nucleotides in the DNA encoding the protein, thereby producing DNA encoding the modification, and thereafter expressing the DNA in recombinant cell culture. Techniques for making substitution mutations at predetermined sites in DNA having a known sequence are well known, for example M13 primer mutagenesis and PCR mutagenesis.
[0151] Nucleic Acids
[0152] The disclosure also features nucleic acids encoding the biosensors (e.g., SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) described herein, including variants and/or fragments of the biosensors (e.g., variants and/or fragments of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94). These sequences include all degenerate sequences related to the specific polypeptide sequence, i.e., all nucleic acids having a sequence that encodes one particular polypeptide sequence as well as all nucleic acids, including degenerate nucleic acids, encoding the disclosed variants and derivatives of the polypeptide sequences. Thus, while each particular nucleic acid sequence may not be written out herein, it is understood that each and every sequence is in fact disclosed and described herein through the disclosed polypeptide sequences.
[0153] In some instances, nucleic acids can encode biosensors with 95, 96, 97, 98, or 99 identity to SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94.
[0154] In some instances, nucleic acids can encode SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94 containing 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 20-30, 30-40, or 40-50 conservative amino acid substitutions.
[0155] In some instances, nucleic acids can encode SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94 containing 50 or fewer, 40 or fewer, 30 or fewer, 20 or fewer, 10 or fewer, 9 or fewer, 8 or fewer, 7 or fewer, 6 or fewer, 5 or fewer, 4 or fewer, 3 or fewer, or 2 or fewer conservative amino acid substitutions
[0156] Also provided herein are vectors comprising the biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) described herein, including variants and/or fragments of the biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94). For example:
[0157] Vectors can include nucleic acids that encode biosensors with 95, 96, 97, 98, or 99 identity to SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94.
[0158] Vectors can include nucleic acids that encode SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94 containing 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 20-30, 30-40, or 40-50 conservative amino acid substitutions.
[0159] Vectors can include nucleic acids that encode SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94 containing 50 or fewer, 40 or fewer, 30 or fewer, 20 or fewer, 10 or fewer, 9 or fewer, 8 or fewer, 7 or fewer, 6 or fewer, 5 or fewer, 4 or fewer, 3 or fewer, or 2 or fewer conservative amino acid substitutions Examples of suitable vectors include, but are not limited to, plasmids, artificial chromosomes, such as BACs, YACs, or PACs, and viral vectors. As used herein, vectors are agents that transport the disclosed nucleic acids into a cell without degradation and, optionally, include a promoter yielding expression of the nucleic acid molecule in the cells into which it is delivered.
[0160] Viral vectors can include, for example, Adenovirus, Adeno-associated virus, herpes virus, Vaccinia virus, Polio virus, Sindbis, and other RNA viruses, including these viruses with the HIV backbone. Any viral families which share the properties of these viruses which make them suitable for use as vectors are suitable. Retroviral vectors, in general are described by Coffin et al., Retroviruses, Cold Spring Harbor Laboratory Press (1997), which is incorporated by reference herein for the vectors and methods of making them. The construction of replication-defective adenoviruses has been described (Berkner et al., J. Virology 61:1213-20 (1987); Massie et al., Mol. Cell. Biol. 6:2872-83 (1986); Haj-Ahmad et al., J. Virology 57:267-74 (1986); Davidson et al., J. Virology 61:1226-39 (1987); Zhang et al., BioTechniques 15:868-72 (1993)). Recombinant adenoviruses have been shown to achieve high efficiency after direct, in vivo delivery to airway epithelium, hepatocytes, vascular endothelium, CNS parenchyma, and a number of other tissue sites. Other useful systems include, for example, replicating and host-restricted non-replicating Vaccinia virus vectors.
[0161] Non-viral based vectors can include expression vectors comprising nucleic acid molecules and nucleic acid sequences encoding polypeptides, wherein the nucleic acids are operably linked to an expression control sequence. Suitable vector backbones include, for example, those routinely used in the art such as plasmids, artificial chromosomes, BACs, YACs, or PACs. Numerous vectors and expression systems are commercially available from such corporations as Novagen (Madison, Wis.), Clontech (Pal Alto, Calif.), Stratagene (La Jolla, Calif.), and Invitrogen/Life Technologies (Carlsbad, Calif.). Vectors typically contain one or more regulatory regions. Regulatory regions include, without limitation, promoter sequences, enhancer sequences, response elements, protein recognition sites, inducible elements, protein binding sequences, 5' and 3' untranslated regions (UTRs), transcriptional start sites, termination sequences, polyadenylation sequences, and introns.
[0162] Promoters controlling transcription from vectors in mammalian host cells may be obtained from various sources, for example, the genomes of viruses such as polyoma, Simian Virus 40 (SV40), adenovirus, retroviruses, hepatitis B virus, and most preferably cytomegalovirus (CMV), or from heterologous mammalian promoters, e.g. .beta.-actin promoter or EFlct promoter, or from hybrid or chimeric promoters (e.g., CMV promoter fused to the .beta.-actin promoter). Of course, promoters from the host cell or related species are also useful herein.
[0163] Enhancer generally refers to a sequence of DNA that functions at no fixed distance from the transcription start site and can be either 5' or 3' to the transcription unit. Furthermore, enhancers can be within an intron as well as within the coding sequence itself. They are usually between 10 and 300 base pairs in length, and they function in cis. Enhancers usually function to increase transcription from nearby promoters. Enhancers can also contain response elements that mediate the regulation of transcription. While many enhancer sequences are known from mammalian genes (globin, elastase, albumin, fetoprotein, and insulin), enhancers derived from a eukaryotic cell viruses can be used. Examples of such can include the SV40 enhancer on the late side of the replication origin, the cytomegalovirus early promoter enhancer, the polyoma enhancer on the late side of the replication origin, and adenovirus enhancers.
[0164] The promoter and/or the enhancer can be inducible (e.g. chemically or physically regulated). A chemically regulated promoter and/or enhancer can, for example, be regulated by the presence of alcohol, tetracycline, a steroid, or a metal. A physically regulated promoter and/or enhancer can, for example, be regulated by environmental factors, such as temperature and light. Optionally, the promoter and/or enhancer region can act as a constitutive promoter and/or enhancer to maximize the expression of the region of the transcription unit to be transcribed. In certain vectors, the promoter and/or enhancer region can be active in a cell type specific manner. Optionally, in certain vectors, the promoter and/or enhancer region can be active in all eukaryotic cells, independent of cell type. Promoters of this type can include the CMV promoter, the SV40 promoter, the .beta.-actin promoter, the EF1.alpha. promoter, and the retroviral long terminal repeat (LTR).
[0165] The provided vectors also can include, for example, origins of replication and/or markers. A marker gene can confer a selectable phenotype, e.g., antibiotic resistance, on a cell. The marker product is used to determine if the vector has been delivered to the cell and once delivered is being expressed. Examples of selectable markers for mammalian cells are dihydrofolate reductase (DHFR), thymidine kinase, neomycin, neomycin analog G418, hygromycin, puromycin, and blasticidin. When such selectable markers are successfully transferred into a mammalian host cell, the transformed mammalian host cell can survive if placed under selective pressure. Examples of other markers include, for example, the E. coli lacZ gene, green fluorescent protein (GFP), and luciferase. In addition, an expression vector can include a tag sequence designed to facilitate manipulation or detection (e.g., purification or localization) of the expressed polypeptide. Tag sequences, such as GFP, glutathione S-transferase (GST), polyhistidine, c-myc, hemagglutinin, or FLAG.TM. tag (Kodak; New Haven, Conn.) sequences typically are expressed as a fusion with the encoded polypeptide. Such tags can be inserted anywhere within the polypeptide including at either the carboxyl or amino terminus.
[0166] The disclosure further provides cells comprising the biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) described herein, including variants and/or fragments of the biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94). Cells can include, for example, eukaryotic and/or prokaryotic cells. For example, cells can include, but are not limited to cells of E. coli, Pseudomonas, Bacillus, Streptomyces; fungi cells such as yeasts (Saccharomyces, and methylotrophic yeast such as Pichia, Candida, Hansenula, and Torulopsis); and animal cells, such as CHO, R1.1, B--W and LM cells, African Green Monkey kidney cells (for example, COS 1, COS 7, BSC1, BSC40, and BMT10), insect cells (for example, Sf9), human cells and plant cells. Suitable human cells can include, for example, HeLa cells or human embryonic kidney (HEK) cells. In general, cells that can be used herein are commercially available from, for example, the American Type Culture Collection (ATCC), P.O. Box 1549, Manassas, Va. 20108. See also F. Ausubel et al., Current Protocols in Molecular Biology, John Wiley & Sons, New York, N.Y., (1998).
[0167] Optionally, the biosensors (e.g., SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) described herein, including variants and/or fragments of the biosensors (e.g., SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) can be located in the genome of the cell (e.g., can be stably expressed in the cell) or can be transiently expressed in the cell.
[0168] Methods of making the provided cells are known and the method of transformation and choice of expression vector will depend on the host system selected. Transformation and transfection methods are described, e.g., in F. Ausubel et al., Current Protocols in Molecular Biology, John Wiley & Sons, New York, N.Y., (1998), and, as described above, expression vectors may be chosen from examples known in the art.
[0169] There are a number of compositions and methods which can be used to deliver the nucleic acid molecules and/or polypeptides to cells, either in vitro or in vivo via, for example, expression vectors. These methods and compositions can largely be broken down into two classes: viral based delivery systems and non-viral based deliver systems. Such methods are well known in the art and readily adaptable for use with the compositions and methods described herein.
[0170] By way of example, the provided polypeptides and/or nucleic acid molecules can be delivered via virus like particles. Virus like particles (VLPs) consist of viral protein(s) derived from the structural proteins of a virus. Methods for making and using virus like particles are described in, for example, Garcea and Gissmann, Current Opinion in Biotechnology 15:513-7 (2004). The provided polypeptides can be delivered by subviral dense bodies (DBs). DBs transport proteins into target cells by membrane fusion. Methods for making and using DBs are described in, for example, Pepperl-Klindworth et al., Gene Therapy 10:278-84 (2003). The provided polypeptides can be delivered by tegument aggregates. Methods for making and using tegument aggregates are described in International Publication No. WO 2006/110728.
[0171] Also provided are transgenic animals comprising one or more cells the biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94) described herein, including variants and/or fragments of the biosensors (e.g, SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94). As used herein, the term animal refers to non-human animals, including, mammals, amphibians and birds. Specifically, examples include sheep, feline, bovines, ovines, pigs, horses, rabbits, guinea pigs, mice, hamsters, rats, non-human primates, and the like. As used herein, transgenic animal refers to any animal, in which one or more of the cells of the animal contain a heterologous nucleic acid. The heterologous nucleic acid can be introduced using known transgenic techniques. The nucleic acid is introduced into the cell, directly or indirectly. For example, the nucleic acid can be introduced into a precursor of the cell or by way of deliberate genetic manipulation, such as by microinjection or by infection with a recombinant virus. The nucleic acid may be integrated within a chromosome, or it may be an extrachromosomally replicating DNA.
[0172] Methods for making transgenic animals using a variety of transgenes have been described in Wagner et al. (1981) Proc. Nat. Acad. Sci. USA, 78:5016-5020; Stewart et al. (1982) Science, 217:1046-1048; Constantini et al. (1981) Nature, 294:92-94; Lacy et al. (1983) Cell, 34:343-358; McKnight et al. (1983) Cell, 34:335-341; Brinstar et al. (1983) Nature, 306:332-336; Palmiter et al. (1982) Nature, 300:611-615; Palmiter et al. (1982) Cell, 29:701-710; and Palmiter et al. (1983) Science, 222:809-814. Such methods are also described in U.S. Pat. Nos. 6,175,057; 6,180,849; and 6,133,502.
[0173] By way of example, the transgenic animal can be created by introducing a nucleic acid into, for example, an embryonic stem cell, an unfertilized egg, a fertilized egg, a spermatozoon or a germinal cell containing a primordial germinal cell thereof, preferably in the embryogenic stage in the development of a non-human mammal (more preferably in the single-cell or fertilized cell stage and generally before the 8-cell phase). The nucleic acid can be introduced by known means, including, for example, the calcium phosphate method, the electric pulse method, the lipofection method, the agglutination method, the microinj ection method, the particle gun method, the DEAE-dextran method and other such method. Optionally, the nucleic acid is introduced into a somatic cell, a living organ, a tissue cell or other cell by gene transformation methods. Cells including the nucleic acid may be fused with the above-described germinal cell by a commonly known cell fusion method to create a transgenic animal.
[0174] For embryonic stem (ES) cells, an ES cell line may be employed, or embryonic cells may be obtained freshly from a host, e.g., mouse, rat, guinea pig, and the like. Such cells are grown on an appropriate fibroblast-feeder layer or grown in the presence of appropriate growth factors, such as leukemia inhibiting factor (LIF). When ES cells have been transformed, they may be used to produce transgenic animals. After transformation, the cells are plated onto a feeder layer in an appropriate medium. Cells containing the construct may be detected by employing a selective medium. After sufficient time for colonies to grow, they are picked and analyzed for the occurrence of homologous recombination or integration of the nucleic acid. Those colonies that are positive may then be used for embryo manipulation and blastocyst injection. Blastocysts are obtained from 4 to 6 week old superovulated females. The ES cells are trypsinized, and the modified cells are injected into the blastocoel of the blastocyst. After injection, the blastocysts are returned to each uterine horn of pseudopregnant females. Females are then allowed to go to term and the resulting litters screened for mutant cells having the construct. By providing for a different phenotype of the blastocyst and the ES cells, chimeric progeny can be readily detected. The chimeric animals are screened for the presence of the nucleic acid, and males and females having the modification are mated to produce homozygous progeny transgenic animals.
[0175] Kits comprising one or more containers and the nucleic acid sequences, polypeptides, vectors, cells, provided herein, or combinations thereof, are also provided.
[0176] For example, provided is a kit comprising (i) a nucleic acid sequence encoding a biosensor described herein (e.g, one or more of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94), including variants and/or fragments of the biosensor (e.g, variants or fragments of one or more of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94), (ii) a polypeptide comprising a biosensor described herein (e.g, one or more of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94), including variants and/or fragments of the biosensor (e.g, variants or fragments of one or more of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94), (iii) a vector comprising the nucleic acid of (i), (iv) a cell comprising the nucleic acid or (i) and/or the polypeptide of (ii), (v) a cell comprising the vector of (iii). The kit can comprise any combination of (i)-(v). Optionally, the kit further comprises reagents for using the nucleic acid or peptide biosensors, vectors, and/or cells. For example, if the kit comprises cells, the kit may also comprise cell culture medium. Optionally, the kit further comprises instructions for use. Optionally, the kit further comprises a GPCR, a GPCR-encoding nucleic acid sequence.
Design and Production/Manufacture Methods
[0177] Using the methods described herein, it is possible to design, produce, and/or adapt genetically encoded biosensors to assays for a variety of classes of analytes. The provided materials and methods facilitate the discovery of new compounds targeting a wide array of protein targets, including but not limited to: endogenous targets responsible for disease state progression, targets on pathogens for treating infectious diseases, and endogenous targets to be avoided (thus screening early for potential drug side effects and toxicity).
[0178] Methods herein provide systematic and generic approaches for the design and production of genetically encoded recombinant peptides containing an analyte-binding framework portion linked (e.g., operably linked) to a signaling portion, wherein the signaling portion is allosterically modulated or regulated by the framework portion upon interaction of the framework portion with an analyte. Generally, methods include: (i) selecting one or more target analytes; (ii) selecting a framework portion (e.g., a PBP) that interacts with (e.g., interacts specifically with) or binds to (e.g., binds specifically to) the target analyte and that undergoes a conformational change upon interacting with or binding to the analyte; (iii) identifying sites or amino acid positions within the framework portion (e.g., the PBP) where the conformational change occurs; and (iv) inserting or cloning a signaling portion into the site or amino acid position identified in (iii). Methods can, optionally, further include: (v) modifying or optimizing linker sequences between the framework portion and the signaling portion, for example, by genetic manipulation (e.g., by point mutation); (vi) modifying or optimizing analyte binding; (vii) modifying the signal generated by the biosensor; and/or (viii) cloning the biosensor into a suitable vector.
[0179] In some instances: (iii) includes identification of insertion sites by analysis of the structure (e.g., crystal structure) of the selected framework portion (e.g., the selected PBP) in one or both of its open and closed states to determine amino acid positions at which analyte-binding dependent structural changes occur. In instances where structures for both open and closed states are not available, analysis can be conducted by analogy to a structurally similar framework portion (e.g., PBP); (iv) includes cloning a signaling portion (e.g., a cpFP) at the site identified in (iii) such that the analyte-binding dependent structural change observed in (iii) will result in a conformational change in the signaling portion (e.g., the cpFP) and allosteric modulation of the signaling portion; (v) includes generating a library of mutants of biosensors with distinct linker sequences (e.g., by point mutation), screening the library of mutants to identify mutants with enhanced properties (e.g., improved signal-to-noise ratio), and selecting mutants with enhanced properties (e.g., improved signal-to-noise ratio); (vi) includes increasing or decreasing binding or affinity of the framework portion to the analyte, e.g., by modifying amino acids in the interacting face of the framework portion or regions within the framework portion that are critical for analyte binding; (vii) includes increasing or decreasing signal emission by the signaling portion and/or changing the color of the signal where the signaling portion is a FP (e.g., a cpFP). Methods including (i)-(viii) are exemplified in the Examples section herein.
Methods of Use
[0180] The disclosure further provides methods for using the biosensors disclosed herein (e.g., one or more of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94), including variants and/or fragments of the biosensor (e.g., variants or fragments of one or more of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, 53, 62, 63, 77, 78, 91, 92, 93, and/or 94)) to detect analytes, e.g., in biological systems. Such methods can include, for example:
[0181] Use of a maltose biosensor disclosed herein (e.g., one or more of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and/or 53 including variants and/or fragments of SEQ ID NOs: 1, 2, 3, 4, 5, 6, 7, 8, 50, 51, 52, and/or 53) to detect maltose, e.g., in a biological system;
[0182] Use of a glutamate biosensor disclosed herein (e.g., one or more of SEQ ID NOs: 62 and/or 63 including variants and/or fragments of SEQ ID NOs: 62 and/or 63) to detect glutamate, e.g., in a biological system;
[0183] Use of a phosphonate biosensor disclosed herein (e.g., one or more of SEQ ID NOs: 77 and/or 78 including variants and/or fragments of SEQ ID NOs: 77 and/or 78) to detect phosphonate, e.g., in a biological system; and/or
[0184] Use of a glucose biosensor disclosed herein (e.g., one or more of SEQ ID NOs: 91, 92, 93 and/or 94 including variants and/or fragments of SEQ ID NOs: 91, 92, 93 and/or 94) to detect glucose, e.g., in a biological system.
[0185] Techniques for performing such methods are known in the art and/or are exemplified herein. For example, methods can include introducing one or more biosensors into a biological system (e.g., a cell); expressing the one or more biosensors in the biological system (e.g., the cell); monitoring the signal emitted by the expressed biosensor in the biological system; and correlating the signal emitted by the expressed biosensor in the biological system with a level of the analyte in the biological system.
EXAMPLES
[0186] The invention is further described in the following examples, which do not limit the scope of the invention described in the claims.
Example 1: Maltose Indicators
[0187] Genetically encoded maltose indicators were generated using Escherichia coli maltodextrin-binding protein (EcMBP) as a framework and either circularly permuted 3-lactamase (cpBla) or circularly permuted fluorescent protein (cpFP) as a signal. Data describe below suggest that cpBla and cpFP are not interchangeable.
[0188] Allosteric coupling of ligand binding to fluorescence was hypothesized to require:
[0189] i) that the site in into which cpGFP is inserted have the capacity to transduce the global conformational change the scaffold protein (EcMBP in this example) to the local environment of the chromophore in cpGFP; and
[0190] ii) that the local environment of the chromophore (e.g., linkers) be optimized to maximize the difference in emission between unbound (apo) and the bound (in this example maltose-bound) states.
Example 1A: Identification of cpGFP Insertion Sites in EcMBP
[0191] Potential insertion sites were identified using the crystal structures of the maltose-bound, closed form of EcMBP (Ouiocho et al., Structure, 5:997-1015, 1997) and the ligand-free, open form of EcMBP shown in FIG. 1 (Sharff et al., Biochemistry, 31:10657-10663, 1992) to guide rational design of EcMBP-cpGFP fusions that would result in maltose-dependent GFP fluorescence.
[0192] For (i), the change in dihedral angle (defined by the C.alpha. atoms spanning four residues) was analyzed to identify maltose-dependent structural changes in sequentially adjacent residues (FIG. 6); this analysis showed that the C.alpha. chain is "torqued" around residues 175 (.DELTA.Dihedral=+41.degree.) and 311 (.DELTA.Dihedral=-22.degree.) upon ligand binding. This sequential conformational change was predicted to be coupled to structural changes of an inserted cpGFP, resulting in maltose-dependent fluorescence for the fusion protein.
[0193] Previous studies using randomly digested and reassembled circularly permuted .beta.-lactamase (cpBla) and EcMBP showed maltose-dependent .beta.-lactamase activity in proteins with insertions of cpBla at EcMBP residues 165 and 317 (Guntas et al., Chem. Biol., 11:1483-1487, 2004; Guntas and Ostermeier, J. Mol. Biol., 336:263-273, 2004).
[0194] Since the .DELTA.Dihedral of EcMBP165 is +11.degree. (moderate change) and EcMBP317 is +2.degree. (no real change), four EcMBP-cpGFP templates were constructed by inserting cpGFP into EcMBP at sites 165, 175 (identified herein), 311 (identified herein), and 317 to test our predictive method and the interchangeability of cpBla and cpGFP at sites identified from the EcMBP-cpBla screen. These constructs were named MBP165-cpGFP, MBP175-cpGFP, MBP311-cpGFP, and MBP317-cpGFP (names were modified to encompass variants (e.g., with modified linker sequences). The cpGFP used is cpGFP146 described in Baird et al. (Proc. Natl. Acad. Sci., USA, 96:11241-11246, 1999). PCR assembly was used to construct fusion proteins with GlyGly-linkers between EcMBP and each terminus of cpGFP. The amino acid sequence of each construct is shown in FIGS. 6-9. The sequences of SEQ ID NOs: 1-3 shown in FIGS. 7A-7C (i.e., MBP165-cpGFP) differ in the linker sequence between MBP 1-165 and cpGFP 147-238 (linker 1: see the line ending in amino acid 240)). The sequences of SEQ ID NOs: 4-5 shown in FIGS. 8A-8B (i.e., MBP175-cpGFP) differ in the sequence between MBP 1-175 and cpGFP 147-238 (linker 1: see the line ending in amino acid 240)). The sequences of SEQ ID NOs: 6-7 shown in FIGS. 9A-9B (i.e., MBP311-cpGFP) differ in the sequence between cpGFP 1-146 and MBP 312-370 (linker 2: see the line ending in amino acid 640)). Each construct includes 3 linkers: A linker between the C-terminus of the C-terminal portion of MBP and the N-terminus of cpGFP (i.e., linker 2), a linker between the N-terminus of cpGFP and C-terminus of the N-terminal portion of MBP, and a linker in cpGFP (i.e., linker 3).
Example 1B: Linker Optimization
[0195] Libraries of variants of SEQ ID NOs: 1-8 were generated with randomized linkers by single-stranded uracil template mutagenesis (see Kunkel et al., Methods Enzymol., 204:125-139, 1991) using the primers listed below:
TABLE-US-00002 165 Linker 1 Primers: (SEQ ID NO: 9) PLIAADGxxNVYIM (SEQ ID NO: 10) PLIAADxxNVYIM (SEQ ID NO11) PLIAADGGxxNVYIM (SEQ ID NO: 12) PLIAADGxPNVYIMG (SEQ ID NO: 13) PLIAADGIxNVYIMG (SEQ ID NO: 14) PLIAADPxSHNVYIM (SEQ ID NO: 15) PLIAADxPSHNVYIM (SEQ ID NO: 16) PLIAADxxSHNVYIM (SEQ ID NO: 17) PLIAADxxSHNVFIM (SEQ ID NO: 18) PLIAADPxSHNVFIM (SEQ ID NO: 19) PLIAADPxSYNVFIM (SEQ ID NO: 20) PLIAADxxSYNVFIM (SEQ ID NO: 21) PLIAADPxSYNVFIM (SEQ ID NO: 22) PLIAADxxSYNVFIM (SEQ ID NO: 23) PLIAADPxSxNVYIM (SEQ ID NO: 24) PLIAADPxSHxVYIM (SEQ ID NO: 25) PLIAADPxSHNxYIM (SEQ ID NO: 26) PLIAADPxSHNVxIM 165 Linker 2 Primers: (SEQ ID NO: 27) KLEYNFNxxYAFKYEN (SEQ ID NO: 28) KLEYNFNxYAFKYEN (SEQ ID NO: 29) KLEYNFNYAFKYEN (SEQ ID NO: 30) KLEYNFxYAFKYEN (SEQ ID NO: 31) KLEYNxxYAFKYEN (SEQ ID NO: 32) KLEYNWxYAFKYEN (SEQ ID NO: 33) KLEYNxKYAFKYEN (SEQ ID NO: 34) KLEYNFNPxYAFKYEN (SEQ ID NO: 35) KLEYNFNxPYAFKYEN 175 Linker 1 Primers: (SEQ ID NO: 36) AFKYENxxSHNVYIM 175 Linker 2 Primers: (SEQ ID NO: 37) KLEYNFNxxKYDIKDV 311 Linker 1 Primers: (SEQ ID NO: 38) KSYEELxxSHNVYIM (SEQ ID NO: 39) KSYEELPxSHNVYIM (SEQ ID NO: 40) KSYEELxPSHNVYIM 311 Linker 2 Primers: (SEQ ID NO: 41) KLEYNFNxxAKDPRIA (SEQ ID NO: 42) KLEYNFNPxAKDPRIA (SEQ ID NO: 43) KLEYNFNxPAKDPRIA 317 Linker 1 Primers: (SEQ ID NO: 44) ELAKDPRxSHNVYIM (SEQ ID NO: 45) ELAKDPRxxSHNVYIM (SEQ ID NO: 46) ELAKDPRxxxSHNVYIM 317 Linker 2 Primers: (SEQ ID NO: 47) KLEYNFNxAATMENA (SEQ ID NO: 48) KLEYNFNxxAATMENA (SEQ ID NO: 49) KLEYNFNxxxAATMENA
[0196] Where "x" indicates that a degenerate primer (with DNA sequence "NNS") was used to encode all 20 possible amino acids.
[0197] About 400 variants were screened in semi-high-throughput fashion, measuring fluorescence intensity of clarified cell lysate in the absence and presence of 10 mM maltose.
[0198] Insertion of cpGFP as MBP317, a site previously reported for cpBla, did not show maltose-dependent fluorescence (FIG. 11) even though the framework protein still bound maltose, as determined by isothermal titration calorimetry (FIG. 12). These data demonstrate that identification of insertion sites by a method other than insertion of cpGFP (such as insertion of cpBla) is not sufficient to identify sites that transduce ligand binding to changes in fluorescence intensity
[0199] Insertion of cpGFP at residue 165 of EcMBP (EcMBP165-cpGFP), another position reported in cpBla studies (Guntas and Ostermeier, supra) with -GlyGly-linkers flanking the cpGFP resulted in a protein in which fluorescence increased 20% (.DELTA.F/F=0.2) upon addition of saturating maltose.
[0200] Screening a fully-degenerate, length-two library ("XX") at either the EcMBP-cpGFP linker (linker 1) or the cpGFP-EcMBP linker (linker 2) yielded proteins with maltose-dependent fluorescent increases >300% or decreases >50% (FIG. 11). Many of the variants with increased .DELTA.F/F values had linkers containing proline(s). Subsequent libraries constructed from oligonucleotides encoding XP or PX and randomization of the residues in GFP from residue 146 to 150 were screened, yielding a final variant with: a two-proline EcMBP-cpGFP linker, a two-glycine cpGFP-EcMBP linker, GFP-H148Y, and GFP-Y151F. This variant, called "EcMBP165-cpGFP.PPYF" (abbreviated PPYF (SEQ ID NO:2)) has a .DELTA.F/F=2.5, a Kd for maltose of 3 .mu.M. Screens also identified variant EcMBP311-cpGFP.L2-NP (-AsnPro- at linker 2 (SEQ ID NO:7)), which has a .DELTA.F/F of 1.0 and a Kd for maltose of 2 .mu.M. This variant has an inferior maltose-dependent fluorescence increase than PPYF, but demonstrates generality of the cpFP insertion method.
[0201] EcMBP175-cpGFP was also screened with XX linkers, and a few variants with .DELTA.F/F.apprxeq.1 were identified (FIG. 11). One mutant, with the first linker encoding HL (EcMBP175-cpGFP.L1-HL (SEQ ID NO:5)), has a .DELTA.F/F=0.5 and a Kd for maltose of 1.3 .mu.M.
[0202] These data support that choice of insertion site by structural analysis is preferable to random insertion.
Example 1C: Modifying Ligand Binding and/or Fluorescent Properties of Sensors
[0203] One objective in the development of generic biosensors is for the framework to permit independent optimization of binding and signaling properties. Analysis of whether biosensors herein permit such optimization was tested using the high-SNR sensor PPYF, by: (i) rationally altering maltose-binding affinity; (ii) changing the ligand-binding specificity from maltose to sucrose, and (iii) creating a family of sensors in multiple colors.
[0204] As a first step, the impacts of mutations of three tryptophan side-chains in the maltose-binding pocket (W230, W62, and W340) were tested. These sites have previously been shown to lower the affinity of EcMBP for maltose by one, two, or three orders of magnitude, respectively, when mutated to alanine (Martineau et al., J. Mol. Biol., 214:337-352, 1990). A mutation to the hinge region, I329W, was also made to PPYF, as this has been shown to increase maltose affinity by about 2-fold in both wild-type EcMBP (Marvin and Hellinga, Nat. Struc. Biol., 8:795-798, 2001) and in the EcMBP-cpBla switches (Guntas et al., Chem. Biol., 11:1483-1487, 2004; Kim and Ostermeier, Arch. Biochem. Biophys., 446:44-51, 2006). As shown in FIG. 13, for the PPYF sensor, the three tryptophan-to-alanine binding-pocket mutations behaved as expected, lowering affinity by between one and three orders of magnitude. In contrast, the I329W mutation did not increase affinity as expected, but rather decreased it. .DELTA.F/F also decreased. This data suggests that the mechanism of fluorescence change in this sensor is dependent on subtle interactions between EcMBP and cpGFP that are linked to the I329W mutation.
[0205] As an alternative test for changing the ligand-binding specificity of the sensor while preserving fluorescence signaling, "5-7" mutations (D14L, K15F, W62Y, E111Y), previously shown to confer EcMBP with an affinity for sucrose (Guntas and Mansell, Proc. Natl. Acad. Sci., 102:11224-11229, 2005), were made to PPYF. As shown in FIG. 14A, the mutations conferred to the sensor about 2 mM affinity for sucrose and.about.3 mM affinity for maltose. To address a discrepancy between expected (micromolar) and observed (millimolar) affinity for disaccharides, the 5-7 mutations were made to sensors with cpGFP inserted at different positions in EcMBP, and with different linker compositions. In the context of EcMBP165-cpGFP.PCF, the 5-7 mutations conferred very low (but observable) binding preference for sucrose over maltose (FIG. 14B). The trend of higher (but still weak) affinity for sucrose (.about.0.6 mM) over maltose (.about.6 mM) continued when the 5-7 mutations are made in the context of EcMBP175-cpGFP.L1-HL (FIG. 14C). In the context of EcMBP311-cpGFP.L2-NP, the 5-7 mutations appeared to eliminate all binding (FIG. 14D). The preference for sucrose over maltose of the 5-7 variants of the sensors is consistent with the binding properties of the 5-7 variants of EcMBP alone and EcMBP-cpBla (Guntas and Mansell, Proc. Natl. Acad. Sci., 102:11224-11229, 2005). The lower affinity for both ligands of the 5-7 variants of the sensors may be the consequence of the inserted cpGFP shifting the open and closed equilibrium.
[0206] These data suggest that ligand binding and fluorescent properties of biosensors can be independently modified.
Example 1D: Modifying Sensor Color
[0207] The color of GFP can be altered by changing the amino acids that either comprise or interact with the chromophore (see Shaner et al., J. Cell. Sci. 120:4247-4260, 2007, for a review).
[0208] Using PPYF as a template, mutations Y66W (to yield a cyan variant, "cpCFP"), L64F+T65G+V68L+T203Y (yellow, "cpYFP"), and Y66H (blue, "cpBFP") mutations were made (see Cubitt et al., Trends Biochem., 20:448-455, 1995, for exemplary methods). As shown in FIG. 15, the variants exhibit fluorescence emission spectra consistent with their respective intended designs.
[0209] The .DELTA.F/F of the color variants in response to maltose is different (in each case inferior) from the .DELTA.F/F of 2.5 observed in PPYF-green. The EcMBP165-cpYFP.PPYF sensor, which has the same covalent chromophore structure as PPYF, has the greatest .DELTA.F/F of the three spectral variants (FIG. 15A). EcMBP165-cpCFP.PPYF has a lower .DELTA.F/F than the green and yellow variants, but by incorporating previously identified mutations, (L1-PC+GFP-Y151F; the resulting protein is called EcMBP165.cpCFP.PCF), a variant with .DELTA.F/F=0.8 was obtained (FIG. 15A).
[0210] The EcMBP165-cpBFP.PPYF variant, while dimly fluorescent, is not a sensor, and a screen of 800 linker variants failed to produce any variant with .DELTA.F/F>0.2 (FIG. 16).
[0211] Since EcMBP165-cpBFP.PPYF was very dim, Azurite mutations T65S+V150I+V224R were included to increase brightness and stability, and make EcMBP165-cpAzurite a good template for linker screening. Using oligonucleotides encoding XX amino acid linkers, a variant was obtained, EcMBP165-cpAzurite.L2-FE, that had .DELTA.F/F=0.8 (FIG. 15).
Example 1E: Modifying Sensor Color and Ligand Specificity/Affinity
[0212] The four sucrose-binding "5-7" mutations described above that conferred weak sucrose affinity in the green sensor (EcMBP165-cpGFP.PPYF) were converted to blue, cyan, and yellow maltose sensors (EcMBP165-cpAzurite.L2-FE, EcMBP165-cpCFP.PCF, and EcMBP165-cpYFP.PPYF). The green and yellow sensors showed increased fluorescence upon addition of 10 mM sucrose, but the cyan and blue proteins did not (FIG. 15A). Like the green variant, the yellow variant had no detectable sucrose affinity with the wild type binding pocket (FIG. 15C) and millimolar affinity for both sugars, with preference for sucrose over maltose (FIG. 15D).
[0213] As seen in FIG. 17, as maltose concentration increased, the blue sensor increased in fluorescence first (Kd .about.2.7 .mu.M), then the green (Kd .about.40 .mu.M), then the yellow (Kd .about.350 .mu.M), and at high maltose concentrations, the cyan variant began to increase its fluorescence (Kd .about.1.7 mM).
Example 1F: Imaging Bacteria
[0214] The ultimate value of genetically encoded fluorescent sensors is in their utility for observing analyte flux in living cells and organisms. In a simple proof-of-principle experiment, Escherichia coli expressing PPYF or PPYF.T203V (see "Second-generation maltose sensors" below) were imaged in the green fluorescence channel in the absence of maltose, and then re-imaged after addition of saturating maltose to the media.
[0215] As shown in FIG. 18, bacteria expressing the sensors clearly became brighter, while control bacteria expressing EGFP appeared unchanged. Increased fluorescence was quantified by measuring the peak (gray-value) pixel intensity of each bacterium. Those expressing PPYF undergo an approximate doubling of fluorescence (bacterium-averaged .DELTA.F/F=1.1.+-.0.4), those expressing PPYF.T203V have slightly increased .DELTA.F/F (.DELTA.F/F=1.29.+-.0.2), while those expressing EGFP have no change in fluorescence (.DELTA.F/F=-0.01.+-.0.05).
Example 1F: 2-Photon Imaging of Mammalian Cells
[0216] Multi-photon microscopy opened new frontiers for in vivo fluorescence imaging, in particular for neuronal activity visualization through the use of genetically encoded calcium indicators (Tian et al., Nat. Methods, 3:281-286, 2009; Denk et al., Science, 248:73-76, 1990; Denk and Svoboda, Neuron, 18:351-357, 1997).
[0217] To demonstrate that the maltose sensors described herein have the potential to be used for 2-photon imaging, fluorescence excitation spectra were collected. As shown in FIG. 19, with a 535 nm bandpass emission filter (50 nm bp), EcMBP165-cpGFP.PPYF showed a 10-fold maltose-dependent increase in fluorescence when excited at 940 nm. All four spectral variants showed a significant maltose-dependent increase in 2-photon fluorescence.
Example 1G: Sub-Cloning Maltose Sensors
[0218] EcMBP165-cpGFP.PPYF.T203V (see "Second-generation maltose sensors" below) were cloned into a modified version of the pDisplay vector (Invitrogen) for extracellular display on the surface of transiently transfected human embryonic kidney (HEK293) cells.
[0219] As shown in FIG. 20, the sensor localized to the plasma membrane and increased in brightness in a concentration-dependent manner when perfused with buffers of varying maltose concentration. The .DELTA.F/F is 5.8-fold, very close to that of the soluble protein produced in E. coli, with the mid-point of the maltose-dependent fluorescence increase being 6.5 .mu.M (FIG. 21A), very similar to the affinity determined on purified protein (5 M). Furthermore, the surface displayed sensor responded rapidly to a pulse of 1 mM maltose (FIG. 21A), indicating that the time course for its action is useful for transient events.
Example 1H: Crystal Structure Analysis of Maltose Sensors
[0220] High-resolution structures of several of the maltose sensors described above were generated. Crystallization trials were performed with EcMBP165-cpGFP.PPYF, EcMBP175-cpGFP.L1-HL, and EcMBP311-cpGFP.L2-NP in the presence and absence of excess maltose, from which both EcMBP175-cpGFP.L1-HL and EcMBP311-cpGFP.L2-NP crystallized in the presence of maltose. X-ray structures were solved to 1.9 and 2.0 .ANG. resolution, respectively, by molecular replacement (FIGS. 22A-22C).
[0221] The structures of the cpGFP and EcMBP domains of the sensors are superimposable with published crystal structures of cpGFP (from GCaMP2; RMSD=0.36 and 0.38 .ANG., respectively, for comparing 221 common C.alpha. atoms) and EcMBP-maltose (RMSD=0.43 and 0.37 .ANG., 370 Ca). The structure of EcMBP is largely unperturbed by insertion of the cpGFP domain; only residues around the 175 and 311 insertion sites showed any significant displacement.
[0222] GFP-H148, which H-bonds the GFP chromophore in the structure of native GFP, also directly H-bonded to the chromophore in the EcMBP175-cpGFP.L1-HL-maltose structure (FIG. 22B), although a different rotamer was observed. In the EcMBP311-cpGFP.L2-NP-maltose structure, GFP-H148 is pulled away from the chromophore and is largely replaced by the Asn from linker 2, which makes H-bond interactions to both strand 8 of the GFP barrel and the chromophore phenolate oxygen (through a water molecule, FIG. 22D). GFP-H148, meanwhile, seemed to stabilize the conformation of linker 2 of EcMBP311-cpGFP.L2-NP by H-bonding the backbone carbonyl of the linker 2 Asn. There is some solvent access to the cpGFP chromophore through the hole in the GFP barrel created by circular permutation, although the inter-domain linkers block much of the opening in both structures. Relatively few contacts are made between the cpGFP and EcMBP domains.
[0223] Based on the structures of two maltose-bound sensors, the sensing mechanism likely involves a shift in the relative position of linker 1 and linker 2 induced by the conformational change in the EcMBP domain associated with maltose binding (FIG. 5). The register shift of interactions between the two linkers could alter the proximity of linker 2 and nearby side-chains to the cpGFP chromophore and change the water structure in the cpGFP opening, leading to a shift in the chromophore protonation equilibrium. This might explain why rigid proline is preferred in either linker, since conformational changes upon ligand binding might be better propagated through the rigid linkers to the cpGFP chromophore environment.
Example 1I: Generation of Second-Generation Maltose Sensors
[0224] In an attempt to increase brightness and .DELTA.F/F of GCaMP, the local environment of the chromophore was altered by randomizing residues within cpGFP, and screening for improved variants (Tian et al., nat. Methods, 6:875-881, 2009).
[0225] As shown in FIG. 23, in the context of EcMBP165-cpGFP.PPYF, the T203V mutation decreases the fluorescence emission of the apo-state by half (FIG. 23A), while saturated fluorescence and affinity are unchanged (FIG. 23B), increasing .DELTA.F/F to 6.5. In the maltose-saturated state, PPYF itself has about a quarter the brightness of EGFP, and half the brightness of cpGFP.
[0226] In the context of EcMBP311-cpGFP.L2-NP, the T203V mutation decreases the brightness of both the apo-state and the saturated-state equally, resulting in no significant change in .DELTA.F/F (FIGS. 23C and D).
[0227] These results indicate that the benefits of the T203V mutation are not universally transferable, and that cpGFP-based fluorescent sensors need to be optimized individually.
Example 2: Maltotriose Indicators
[0228] Genetically encoded maltotriose indicators were created using Pyrococcus furiosus maltotriose binding protein. As described below, only the structure of the ligand-bound state P. furiosus maltotriose binding protein (PfMBP) is available. As shown in FIGS. 1 and 2, PfMBP is homologous to EcMBP (compare FIGS. 1 and 2). Two sensors were made, PfMBP171 and PfMBP316, the insertion points for which were selected based on homology to EcMBP165 and EcMBP311, respectively. Linkers were optimized. PfMBP sensors have a .DELTA.F/F of .about.1.2.
[0229] Pyrococcus furiosus is a thermophilic organism. Proteins from thermophiles have been shown to be more amenable to mutation than those from mesophiles (Bloom et al., Proc. Natl. Acad. Sci., 103:5869-5874, 2006). As an alternative to developing new sensors by inserting cpGFP into PBPs, it should also be possible to generate new sensors by changing the ligand-binding specificity of an existing PBP-based sensor.
[0230] It has previously been shown that the binding sites of PBPs can be reengineered to accommodate novel ligands (Looger et al., Nature, 423:185-190, 2003). However, those re-design efforts used framework proteins from mesophiles and suffered from poor stability. We hypothesized that PfMBP, which is intrinsically more stable than EcMBP, is more tolerant of mutations. To test this hypothesis, we characterized and compared the stability of PfMBP to EcMBP, PfMBP-cpGFP sensors to EcMBP-cpGFP sensors, PfMBP binding site mutants to EcMBP binding site mutants, and PfMBP-cpGFP sensor binding site mutants to EcMBP-cpGFP sensor binding site mutants. Conclusively, the PfMBP variants were more stable than the EcMBP variants. Finally, we demonstrate that the increased thermo-stability of the PfMBP-cpGFP sensors is useful for the measurement of maltotriose at temperatures as high at 60.degree. C., whereas the EcMBP-cpGFP sensors are only useful for the measurement of maltose at temperatures as high as 40.degree. C.
Example 2A: Identification of cpGFP Insertion Sites in PfMBP
[0231] The ligand-bound (closed) structure of PfMBP is available (Evdokimov et al., J. Mol. Biol., 305:891-904, 2001), but the unbound structure is not. Accordingly, insertion sites for the PfMBP-cpGFP sensors were identified by homology to EcMBP.
[0232] Sites were selected based on the structural similarities between PfMBP and EcMBP. Two sites were selected. One of these sites is EcMBP311, which is homologous to PfMBP316. This site is at juncture between the end of the cluster of helices (Helices 8a, 8b, 8c) and the start of the "equatorial" spanning helix (Helix 9). Another site that was made into a sensor in EcMBP was EcMBP165, which is homologous to PfMBP171. cpGFP was inserted into PfMBP at each of these sites. The sequences of the resulting constructs, PfMBP171-cpGFP and PfMBP316-cpGFP, are shown in FIGS. 24 and 25, respectively.
Example 2B: Linker Optimization
[0233] Libraries of variants of SEQ ID NOs: 50-53 were generated with randomized linkers by single-stranded uracil template mutagenesis using the primers listed below:
TABLE-US-00003 175 Linker 1 Primers: (SEQ ID NO: 54) AIAQAFxxSHNVYIMA (SEQ ID NO: 55) AIAQAFPxSHNVYIMA 171 Linker 2 Primers: (SEQ ID NO: 56) KLEYNFNxxYYFDDKTE 316 Linkerl Primers (SEQ ID NO: 57) VLDDPExxHNVYIM (SEQ ID NO: 58) VLDDPEIxxSHNVYIM 316 Linker2 Primers (SEQ ID NO: 59) KLEYNFxxNDPVIY (SEQ ID NO: 60) KLEYNFNxPKNDPVIY (SEQ ID NO: 61) KLEYNFNPxKNDPVIY
[0234] Where "x" indicates that a degenerate primer (with DNA sequence "NNS") was used to encode all 20 possible amino acids.
[0235] Several thousand variants were screened in semi-high-throughput fashion, measuring fluorescence intensity of clarified cell lysate in the absence and presence of 1 mM maltotriose.
[0236] Screening a fully-degenerate, length-two library ("XX") at either the PfMBP171-cpGFP linker (linker 1) or the cpGFP-PfMBP linker (linker 2) yielded proteins with maltotriose-dependent fluorescent increases >100% or decreases >20% (FIG. 26A). A variant from this group with a GlyGly PfMBP-cpGFP linker and a PheGlu cpGFP-PfMBP linker was selected for further characterization. This variant, called "PfMBP171-cpGFP.L2FE" has a .DELTA.F/F=1.2, a Kd for maltotriose of <1 .mu.M.
[0237] Screening a fully-degenerate, length-two library ("XX") at either the PfMBP316-cpGFP linker (linker 1) or the cpGFP-PfMBP linker (linker 2) also yielded proteins with maltotriose-dependent fluorescent increases >100% or decreases >20% (FIG. 26B). A variant from this group with a GlyGly PfMBP-cpGFP linker and a PheGlu cpGFP-PfMBP linker was selected for further characterization. This variant, called "PfMBP316-cpGFP.L1-NP" has a .DELTA.F/F=1.2, a Kd for maltotriose of 40 .mu.M.
[0238] These data support that structurally homologous frameworks can be compared to identify insertion sites for cpGFP.
Example 2C: Characterization of the Thermostability of the PfMBP and PfMBP-cpGFP Compared to EcMBP and EcMBP-cpGFP
[0239] Thermal stability of PfMBP171-cpGFP.L2FE was measured using circular-dichroism (CD) and compared to the original EcMBP and PfMBP binding proteins, along with cpGFP. Following the changes by means of CD allowed determination of whether different transitions happened in alpha, beta, or both kinds of structures.
[0240] Given that cpGFP is a beta barrel, strong transitions in the beta signal alone were associated with changes in this kind of structure. In the same way, transitions in both kinds of signals were associated with the binding protein structure. As shown in FIG. 27A, PfMBP is significantly more thermo-stable than EcMBP. In fact, while EcMBP denatured at about 50.degree. C., PfMBP did not denature at temperatures less than 80.degree. C. Also, the addition of maltose to EcMBP stabilized the protein by about 10.degree. C.
[0241] As shown in FIG. 27B, the stability of the EcMBP component of the EcMBP165-cpGFP.PPYF sensor decreased from 50.degree. C. to 45.degree. C. with insertion of cpGFP, while the intrinsic stability of cpGFP in the sensor remained unchanged. There was little change in the stability of the PfMBP component of the PfMBP171-cpGFP.L2FE sensor with insertion of cpGFP (FIG. 27B). Moreover, PfMBP seemed to exert a small stabilizing effect over the inserted cpGFP, as shown by the change in the steepness and melting point of the curve of the soluble form and the PfMBP171-cpGFP.L2FE sensor. All the associations made between transitions and domain unfolding were supported by CD spectra taken at the beginning and the end of each temperature ramp.
[0242] Analysis of whether the PfMBP scaffold was more tolerant of mutation than the EcMBP scaffold was also performed. Proof-of-principle mutations were made to the ligand-binding sites of EcMBP and PfMBP, and their respective sensors. In EcMBP, Asn12 was mutated to Trp to result in steric clashes with the surrounding residues, and backbone, of the binding pocket. The homologous mutation in PfMBP is Ala13Trp, which would be expected to have the same effect.
[0243] As shown in FIG. 27C, N12W decreased the Tm of EcMBP from 50.degree. C. to 40.degree. C., while the corresponding mutation in PfMBP, A13W, had no noticeable effect. This data confirms that the thermo-philic protein is more tolerant of mutations to the binding site. Furthermore, in the context of the sensors, the N12W mutation to EcMBP165-cpGFP.PPYF completely destabilized the binding protein component of the sensor (FIG. 27D), while the A13W mutation in PfMBP171-cpGFP.L2FE had no effect on stability (FIG. 27D).
Example 2D: Tolerance of PfMBP Sensor to Increased Temperature
[0244] Fluorescence of the protein in the apo and ligand-bound states at was measured at different temperatures.
[0245] As shown in FIG. 28A, fluorescence of the EcMBP165-cpGFP.PPYF sensor in the bound state was higher than it is in the apo-state at lower temperatures, by about 4-fold. However, at around 55.degree. C. (the unfolding transition of the EcMBP component) the fluorescence of the EcMBP165-cpGFP.PPYF sensor dropped precipitously. As a result, EcMBP165-cpGFP.PPYF is unsuitable for detection of maltose at temperatures greater than 50.degree. C. (FIG. 28B). In contrast, PfMBP171-cpGFP.L2FE retained its maltotriose binding capabilities at high temperatures (FIGS. 28A and 28B), and is limited only by the intrinsic fluorescence of the cpGFP component, which decays at about 80.degree. C. (FIG. 28A).
Example 2E: Measurement of Maltodextrins in Hot Liquids
[0246] To demonstrate that the soluble and immobilized sensors function similarly, PfMBP171-cpGFP.L2FE, PfMBP316-cpGFPL1XXX, and EcMBP165-cpGFP.PPYF.T203V were immobilized via their N-terminal poly-histidine tags on to the surface of Ni-NTA coated glass. In a fluorescence plate reader, the immobilized proteins performed similarly to their soluble counterparts (see FIGS. 28C, 28D, and 28F).
[0247] Next, a prototype device was constructed, with a light guide providing the excitation light and returning the fluorescent emitted light back to the photodetector, the bio-sensor protein immobilized to Ni-NTA coated coverslips, and the coverslip attached to the end of the light guide. The "wand" of the detector was dipped into different compositions of solutions, each with varying concentrations of maltose or maltotriose. Experiments were performed at different temperatures. PfMBP-cpGFP sensor performed better at higher temperatures (as high as 60.degree. C.) than the EcMBP-cpGFP sensor.
Example 3: Glutamate Indicators
[0248] Glutamate indicators were created from Escherichia coli glutamate-binding protein (EcYbeJ). As with PfMBP in Example 2, only the structure of the ligand-bound EcYbeJ is available. EcYbeJ is homologous to EcMBP, but to a lesser degree. The best homology match between a site in EcYbeJ and a site in a binding protein for which an intensity-based sensor has already been created is EcYbeJ253 and EcMBP311 (described herein). As shown in FIG. 3, both sites are at the junction of "Rising Helix 8" and the "Equatorial Helix/Coil." The amino acid composition of the cpGFP and EcYbeJ junction was made the same as that of the EcMBP311-cpGFP sensor (Linker 2=NP). The amino acid composition of the EcYbeJ junction and cpGFP was optimized to LV (Linker 1=LV). The variant has a .DELTA.F/F of 5.
Example 3A: Identification of cpGFP Insertion Sites
[0249] The ligand-bound (closed) structure of Shigella flexneri glutamate binding protein is available (Fan et al., Protein Pept. Lett., 13:513-516, 2006). This protein has only 4 amino acid mutations relative to EcYbeJ, and is thus an appropriate model.
[0250] Insertion sites for the EcYbeJ-cpGFP sensors were identified by homology to EcMBP. Based on the topology map (FIG. 3), position 311 in EcMBP was identified as an acceptable insertion site for EcYbeJ. EcMBP311 is equivalent to EcYbeJ253. EcYbeJ253 is at juncture between the end of the cluster of helices (Helices 8a, 8b, 8c) and the start of the "equatorial" spanning helix (Helix 9). In YbeJ, the structure that is homologous to the equatorial helix is the equatorial coil (depicted in red, to match the red coloring of Helix 9).
[0251] Intrinsic affinity of wild-type YbeJ for glutamate (.about.1 .mu.M) was too high to permit high-throughput screening of linker libraries. Endogenous glutamate (from the growth media) saturates the sensor, making measurement of the unbound state technically challenging. A mutation to YbeJ (A184V), in the "hinge" of the protein were made. Mutation of this residue to Trp or Arg have previously been shown to decrease affinity in FRET-based sensors (see Okumoto et al., Proc. Natl. Acad. Sci., 102:8740-8745, 2005). EcYbeJ253 (A184V)-cpGFP has an affinity for glutamate of about 100 .mu.M. All references to EcYbeJ253-cpGFP, unless otherwise noted, refer to the A184V variant. The sequences of the EcYbeJ constructs are shown in FIG. 29.
Example 3B: Linker Optimization
[0252] Libraries of variants of SEQ ID NOs: 62-63 were generated with randomized linkers by single-stranded uracil template mutagenesis using the primers listed below:
TABLE-US-00004 253 Linker 1 Primers: (SEQ ID NO: 64) FKNPIPPxSHNVYIMA (SEQ ID NO: 65) FKNPIPPxxSHNVYIMA (SEQ ID NO: 66) FKNPIPPPxSHNVYIMA (SEQ ID NO: 67) FKNPIPPxPSHNVYIMA (SEQ ID NO: 68) KWFKNPIxxSHNVYIMA (SEQ ID NO: 69) FKNPIPPxxNVYIMAD (SEQ ID NO: 70) KWFKNPIxxNVYIMAD 253 Linker 2 Primers: (SEQ ID NO: 71) KLEYNFNxKNLNMNF (SEQ ID NO: 72) KLEYNFNxxKNLNMNF (SEQ ID NO: 73) KLEYNFNxPKNLNMNF (SEQ ID NO: 74) KLEYNFNPxKNLNMNF (SEQ ID NO: 75) GHKLEYNxxLNMNF (SEQ ID NO: 76) KLEYNFNxxLNMNF
[0253] Where "x" indicates that a degenerate primer (with DNA sequence "NNS") was used to encode all 20 possible amino acids.
[0254] Several thousand variants were screened in semi-high-throughput fashion, measuring fluorescence intensity of clarified cell lysate in the absence and presence of 10 mM glutamate.
[0255] Screening a fully-degenerate, length-two library ("XX") at the EcYbeJ253-cpGFP linker (linker 1) identified a sensor with glutamate-dependent fluorescent increases of .about.100%. This variant has a LeuVal EcYbeJ-cpGFP linker (L1-LV) and was used as the framework for optimization of the cpGFP-EcYbeJ253 linker (linker 2). The results of that screen yielded a protein with glutamate-dependent fluorescent increase of .about.500% and a linker 2 composition of AsnPro. As shown in FIG. 30, this variant, called "EcYbeJ253-cpGFP.L1LVL2NP" has a .DELTA.F/F=5, a Kd for glutamate of 100 .mu.M. Interestingly, the composition of the second linker, AsnPro, is the same as the linker composition of EcMBP311-cpGFP.L2NP.
Example 3C: Detection of Extracellular Glutamate
[0256] EcYbeJ253-cpGFP.L1LVL2NP was cloned into the pDisplay.TM. vector to allow targeting and anchoring of the sensor to the plasma membrane. The resulting construct was transfected into cultured mammalian cells (HEK293) to visualize the addition of glutamate to extracellular media. Constructs were also generated in a bacterial expression vector with the epitope tags individually and in combination.
[0257] As shown in FIG. 31, the hemagglutinin tag interferes with the fluorescence change. EcYbeJ253-cpGFP.L1LVL2NP was re-cloned into a derivative of the pDisplay.TM. vector, lacking the hemagglutinin tag, called pMinDis (for Minimal Display). This new construct, when expressed in HEK293 cells, shows a change in fluorescence intensity under 2-photon excitation that is approximately the same as the soluble protein (see FIG. 32) with higher affinity, of about 1 .mu.M (see FIG. 32).
[0258] To demonstrate that the sensor is functional in neurons, and not just cultured HEK cells, the gene from EcYbeJ253-cpGFP.L1LVL2NP was cloned into an adeno-associated virus vector (AAV) under control of the synapsin promoter. Virus particles were generated and used to infect cultured primary hippocampus neurons from rats 7 days after culturing. 14 days after culturing (and 7 days after infection), the infected neurons were imaged under 2-photon microscopy (FIG. 33).
Example 4: Phosphonate Indicators
[0259] An indicator for phosphonate compounds was created from Escherichia coli phosphonate-binding protein (EcPhnD). In this instance, only the structure of the ligand-bound state was available at the time the sensor was conceived. EcPhnD is homologous to EcMBP to a lesser degree and to EcYbeJ to a greater degree. The best homology match between a site in EcPhnD and a site in a binding protein for which an intensity-based sensor has already been created is EcPhnD90 and EcYbeJ253. There is no "Rising Helix 8" in EcPhnD, but there is an "Equatorial Helix/Coil" (FIG. 4). cpGFP was inserted at the Equatorial Helix/Coil and linkers were optimized to yield a sensor with .DELTA.F/F of 1.2. EcPhnD is a dimmer, so, a pair of mutations (L297R+L301R) were made to convert it to a monomer. The monomer variant has a .DELTA.F/F of 1.6.
Example 4A: Identification of cpGFP Insertion Sites in EcPhnD
[0260] Insertion sites for the EcPhnD-cpGFP sensors were identified using the ligand-bound (closed) structure of EcPhnD by homology to EcMBP. Based on the topology map (FIG. 4), position 311 in EcMBP was identified as an acceptable insertion site in EcPhnD. EcMBP311 corresponds to EcPhnD90. This site is at the point where the rising strand (Strand D) of EcPhnD has a small bend in it that runs equatorial to the rest of the sheets in the protein. Even though it is topologically different from the "equatorial" spanning helix (Helix 9) of EcMBP its equatorial alignment is similar, and with just the closed structure at the time, in an environment that was expected to undergo significant dihedral change upon binding ligand. Sequences of EcPhnD constructs are shown in FIG. 34.
Example 4B: Linker Optimization
[0261] Libraries of variants of SEQ ID NOs: 77-78 were generated with randomized linkers by single-stranded uracil template mutagenesis using the primers listed below:
TABLE-US-00005 90 Linker 1 Primers: (SEQ ID NO: 79) QTVAADGSSHNVYIMA (SEQ ID NO: 80) QTVAADxxSHNVYIMA (SEQ ID NO: 81) QTVAADxPSHNVYIMA (SEQ ID NO: 82) QTVAADPxSHNVYIMA (SEQ ID NO: 83) QTVAADxxNVYIMA (SEQ ID NO: 84) QTVAADxxSHNVYIMA (SEQ ID NO: 85) VFQTVAxxSHNVYIMA 90 Linker 2 Primers: (SEQ ID NO: 86) HKLEYNFNPGYWSVLI (SEQ ID NO: 87) HKLEYNFNxxPGYWSVLI (SEQ ID NO: 88) HKLEYNxxPGYWSVLI (SEQ ID NO: 89) HKLEYNFNxxYWSVLI (SEQ ID NO: 90) HKLEYNFNPxYWSVLI
[0262] Where "x" indicates that a degenerate primer (with DNA sequence "NNS") was used to encode all 20 possible amino acids.
[0263] Several thousand variants were screened in semi-high-throughput fashion, measuring fluorescence intensity of clarified cell lysate in the absence and presence of 100 uM 2AEP.
[0264] Screening a number of fully-degenerate, libraries at the EcPhnD90-cpGFP linker (linker 1) yielded a protein with 2AEP-dependent fluorescent increases of >100%. This variant has a AlaAsp EcPhnD-cpGFP linker (L1-AD) and a .DELTA.F/F of 1.2. The variant came from a linker that also deleted two residues, effectively making the insertion point of cpGFP occur after residue D88, and then skipping to residue P91 at the cpGFP-EcPhnD linker.
[0265] It was observed from the crystal structure that EcPhnD forms a dimer. To disrupt the dimer inter-face and potentially simplify the observable binding behavior of the EcPhnD protein, two mutations, L297R and L301R, were introduced into the dimerization helices. These mutations were expected, by charge repulsion, to disrupt the dimer interface. As shown in FIG. 35, incorporation of L279R and L301R mutations into EcPhnD90-cpGFP.L1AD caused .DELTA.F/F to increases to 1.6 in response to 2AEP.
[0266] Further attempts to crystallize the open, ligand-unbound form of the protein were successful after making a mutation to the binding site, H157A, that substantially decreased affinity for phosphonate compounds. This mutant was crystallized in the absence of ligand, and the open state of the protein solved. The .DELTA.Dihedral analysis (FIG. 36) showed that the region of greatest dihedral change was the group of residues from 88-90, just one amino acid away from the site chosen by homology to the equatorial helix.
[0267] These data further indicate that .DELTA.Dihedral metric is sufficient for identifying sites in PBPs into which cpGFP can be inserted and result in intensity-based fluorescent sensors.
Example 5: Glucose Indicators
[0268] Glucose indicators were created from Thermus thermophilus glucose binding protein (TtGBP). In this instance, only the structure of the ligand-bound state is available. TtGBP is very homologous to EcMBP and PfMBP (compare FIG. 5 with FIGS. 1 and 2). The insertion point (TtGBP326) was chosen by homology to EcMBP311 and PfMBP316. The amino acid composition of the cpGFP and TtGBP junction was made the same as that of the EcMBP311-cpGFP and EcYbeJ253 sensors (Linker 2=NP). Linker 1 was optimized (Linker 1=PA) and the TtGBP326 sensor have a .DELTA.F/F of .about.2.5. To improve its utility for the measuring glucose concentrations in human blood, the affinity was weakened from its native.about.1 .mu.M to 1.5 mM by mutation of two residues in the binding pocket (H66A+H348A).
Example 5A: Identification of cpGFP Insertion Sites in TtGBP
[0269] The ligand-bound (closed) structure of TtGBP is available (Cuneo et al., J. Mol. Biol., 362:259-270, 2006). Accordingly, insertion sites for the TtGBP-cpGFP sensors were identified by homology to EcMBP and PfMBP. Based on the topology map (FIG. 5), it is apparent that TtGBP, PfMBP, and EcMBP are structurally similar in the closed, ligand-bound state. Positions in EcMBP determined by the dihedral analysis (see above) were predicted to be acceptable insertion sites in TtGBP. EcMBP311 is homologous to TtGBP326. This site is at juncture between the end of the cluster of helices (Helices 8a, 8b, 8c) and the start of the "equatorial" spanning helix (Helix 9). The amino acid sequence of the TtGBP construct is shown in FIG. 37.
Example 5B: Linker Optimization
[0270] Libraries of variants of SEQ ID NO:91 were generated with randomized linkers by single-stranded uracil template mutagenesis using the primers listed below:
TABLE-US-00006 326 Linker 1 Primers: (SEQ ID NO: 95) DSDPSKYxxSHNVYIM (SEQ ID NO: 96) DSDPSKYPxSHNVYIM (SEQ ID NO: 97) DSDPSKYxPSHNVYIM (SEQ ID NO: 98) RLDSDPSxxSHNVYIM (SEQ ID NO: 99) DSDPSKYxxNVYIM 326 Linker 2 Primers: (SEQ ID NO: 100) KLEYNFNxxNAYGQSA (SEQ ID NO: 101) KLEYNFxxPNAYGQSA (SEQ ID NO: 102) GHKLEYNxxNAYGQSA (SEQ ID NO: 103) KLEYNFNxPNAYGQSA (SEQ ID NO: 104) KLEYNFNPxNAYGQSA
[0271] Where "x" indicates that a degenerate primer (with DNA sequence "NNS") was used to encode all 20 possible amino acids.
[0272] Several hundred variants were screen in semi-high-throughput fashion, measuring fluorescence intensity of clarified cell lysate in the absence and presence of 10 mM glucose.
[0273] Linker 1 was optimized (Linker 1=PA) and the TtGBP326-cpGFP.L1PAL2NP sensor has a .DELTA.F/F of .about.2.5 (see FIG. 38). Additionally, the TtGBP sensor was tested with and without the N-terminal pRSET tag and no difference was observed. Specifically, both sensors exhibited an affinity for glucose of about 1.5 mM and a .DELTA.F/F of 2.5.
[0274] Data showing that it was possible to construct a glucose sensor by replacing the EcMBP or PfMBP with TtGBP, retaining the composition of linker 2, and optimizing the composition of linker 1, indicates that the methods for generating sensors disclosed herein can be used to generate sensors using any suitable framework.
Example 5C: Detecting Changes in Glucose Concentration In Vivo
[0275] The TtGBP326-cpGFP.L1PAL2NP sensor was cloned into a variant of the pDisplay.TM. vector lacking the N-terminal secretion sequence, the N-terminal hemagglutinin tag, the C-terminal cMyc tag, and the C-terminal PDGFR membrane anchoring domain.
[0276] The TtGBP sensor was cloned into a mammalian expression vector (based on the pDisplay.TM. vector described in Example 3 above) with the secretion, epitope, and transmembrane anchoring peptides removed, thus resulting in cytosolic expression of the TtGBP326-cpGFP.L1PAL2NP+H66A+H348A sensor. The construct was transfected into HEK293 cells. As shown in FIG. 39, the TtGBP sensor was expressed in the cytosol.
[0277] As shown in FIG. 40, addition of 10 mM glucose to the media increases fluorescence.
[0278] The TtGBP326-cpGFP.L1PAL2NP+H66A+H348A sensor was further modified by L276V mutation to produce TtGBP326.L1PA.L2NP.H66A.H348A.L276V (see FIG. 50). As shown in FIG. 51, this construct has an affinity for glucose of 6.5 mM.
[0279] Additionally, the TtGBP326.L1P1.L2NP.G66A.H348A.L276V was cloned into the pMinDis derivative of the pDisplay vector and expressed on the extracellular surface of HEK293 cells. After exchanging the HEK293 cell media for PBS, addition of glucose to the PBS led to an increase in fluorescence (see FIG. 52).
[0280] These data indicate, in part, that the pRSET tag is not essential to the function of the sensor and that the TtGBP326-cpGFP.L1PAL2NP sensor is capable of detecting changes in the concentration of glucose inside or on the external surface of human cells.
Other Embodiments
[0281] It is to be understood that while the invention has been described in conjunction with the detailed description thereof, the foregoing description is intended to illustrate and not limit the scope of the invention, which is defined by the scope of the appended claims. Other aspects, advantages, and modifications are within the scope of the following claims.
Sequence CWU 1
1
1761662PRTArtificial Sequencerecombinant peptide biosensor 1Met Arg
Gly Ser His His His His His His Gly Met Ala Ser Met Thr1 5 10 15Gly
Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25
30Arg Trp Gly Ser Lys Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn
35 40 45Gly Asp Lys Gly Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe
Glu 50 55 60Lys Asp Thr Gly Ile Lys Val Thr Val Glu His Pro Asp Lys
Leu Glu65 70 75 80Glu Lys Phe Pro Gln Val Ala Ala Thr Gly Asp Gly
Pro Asp Ile Ile 85 90 95Phe Trp Ala His Asp Arg Phe Gly Gly Tyr Ala
Gln Ser Gly Leu Leu 100 105 110Ala Glu Ile Thr Pro Asp Lys Ala Phe
Gln Asp Lys Leu Tyr Pro Phe 115 120 125Thr Trp Asp Ala Val Arg Tyr
Asn Gly Lys Leu Ile Ala Tyr Pro Ile 130 135 140Ala Val Glu Ala Leu
Ser Leu Ile Tyr Asn Lys Asp Leu Leu Pro Asn145 150 155 160Pro Pro
Lys Thr Trp Glu Glu Ile Pro Ala Leu Asp Lys Glu Leu Lys 165 170
175Ala Lys Gly Lys Ser Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe
180 185 190Thr Trp Pro Leu Ile Ala Ala Asp Gly Gly Ser His Asn Val
Tyr Ile 195 200 205Met Ala Asp Lys Gln Lys Asn Gly Ile Lys Ala Asn
Phe Lys Ile Arg 210 215 220His Asn Ile Glu Asp Gly Gly Val Gln Leu
Ala Tyr His Tyr Gln Gln225 230 235 240Asn Thr Pro Ile Gly Asp Gly
Pro Val Leu Leu Pro Asp Asn His Tyr 245 250 255Leu Ser Thr Gln Ser
Lys Leu Ser Lys Asp Pro Asn Glu Lys Arg Asp 260 265 270His Met Val
Leu Leu Glu Phe Val Thr Ala Ala Gly Ile Thr Leu Gly 275 280 285Met
Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser Met Val Ser Lys 290 295
300Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu Val Glu Leu
Asp305 310 315 320Gly Asp Val Asn Gly His Lys Phe Ser Val Ser Gly
Glu Gly Glu Gly 325 330 335Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys
Phe Ile Cys Thr Thr Gly 340 345 350Lys Leu Pro Val Pro Trp Pro Thr
Leu Val Thr Thr Leu Thr Tyr Gly 355 360 365Val Gln Cys Phe Ser Arg
Tyr Pro Asp His Met Lys Gln His Asp Phe 370 375 380Phe Lys Ser Ala
Met Pro Glu Gly Tyr Ile Gln Glu Arg Thr Ile Phe385 390 395 400Phe
Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu Val Lys Phe Glu 405 410
415Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys Gly Ile Asp Phe Lys
420 425 430Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu Tyr Asn Phe
Asn Gly 435 440 445Gly Tyr Ala Phe Lys Tyr Glu Asn Gly Lys Tyr Asp
Ile Lys Asp Val 450 455 460Gly Val Asp Asn Ala Gly Ala Lys Ala Gly
Leu Thr Phe Leu Val Asp465 470 475 480Leu Ile Lys Asn Lys His Met
Asn Ala Asp Thr Asp Tyr Ser Ile Ala 485 490 495Glu Ala Ala Phe Asn
Lys Gly Glu Thr Ala Met Thr Ile Asn Gly Pro 500 505 510Trp Ala Trp
Ser Asn Ile Asp Thr Ser Lys Val Asn Tyr Gly Val Thr 515 520 525Val
Leu Pro Thr Phe Lys Gly Gln Pro Ser Lys Pro Phe Val Gly Val 530 535
540Leu Ser Ala Gly Ile Asn Ala Ala Ser Pro Asn Lys Glu Leu Ala
Lys545 550 555 560Glu Phe Leu Glu Asn Tyr Leu Leu Thr Asp Glu Gly
Leu Glu Ala Val 565 570 575Asn Lys Asp Lys Pro Leu Gly Ala Val Ala
Leu Lys Ser Tyr Glu Glu 580 585 590Glu Leu Val Lys Asp Pro Arg Ile
Ala Ala Thr Met Glu Asn Ala Gln 595 600 605Lys Gly Glu Ile Met Pro
Asn Ile Pro Gln Met Ser Ala Phe Trp Tyr 610 615 620Ala Val Arg Thr
Ala Val Ile Asn Ala Ala Ser Gly Arg Gln Thr Val625 630 635 640Asp
Glu Asp Leu Lys Asp Ala Gln Thr Arg Ile Thr Lys Gly Ser His 645 650
655His His His His His Gly 6602662PRTArtificial Sequencerecombinant
peptide biosensor 2Met Arg Gly Ser His His His His His His Gly Met
Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp
Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys Ile Glu Glu Gly Lys
Leu Val Ile Trp Ile Asn 35 40 45Gly Asp Lys Gly Tyr Asn Gly Leu Ala
Glu Val Gly Lys Lys Phe Glu 50 55 60Lys Asp Thr Gly Ile Lys Val Thr
Val Glu His Pro Asp Lys Leu Glu65 70 75 80Glu Lys Phe Pro Gln Val
Ala Ala Thr Gly Asp Gly Pro Asp Ile Ile 85 90 95Phe Trp Ala His Asp
Arg Phe Gly Gly Tyr Ala Gln Ser Gly Leu Leu 100 105 110Ala Glu Ile
Thr Pro Asp Lys Ala Phe Gln Asp Lys Leu Tyr Pro Phe 115 120 125Thr
Trp Asp Ala Val Arg Tyr Asn Gly Lys Leu Ile Ala Tyr Pro Ile 130 135
140Ala Val Glu Ala Leu Ser Leu Ile Tyr Asn Lys Asp Leu Leu Pro
Asn145 150 155 160Pro Pro Lys Thr Trp Glu Glu Ile Pro Ala Leu Asp
Lys Glu Leu Lys 165 170 175Ala Lys Gly Lys Ser Ala Leu Met Phe Asn
Leu Gln Glu Pro Tyr Phe 180 185 190Thr Trp Pro Leu Ile Ala Ala Asp
Pro Pro Ser Tyr Asn Val Phe Ile 195 200 205Met Ala Asp Lys Gln Lys
Asn Gly Ile Lys Ala Asn Phe Lys Ile Arg 210 215 220His Asn Ile Glu
Asp Gly Gly Val Gln Leu Ala Tyr His Tyr Gln Gln225 230 235 240Asn
Thr Pro Ile Gly Asp Gly Pro Val Leu Leu Pro Asp Asn His Tyr 245 250
255Leu Ser Thr Gln Ser Lys Leu Ser Lys Asp Pro Asn Glu Lys Arg Asp
260 265 270His Met Val Leu Leu Glu Phe Val Thr Ala Ala Gly Ile Thr
Leu Gly 275 280 285Met Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser
Met Val Ser Lys 290 295 300Gly Glu Glu Leu Phe Thr Gly Val Val Pro
Ile Leu Val Glu Leu Asp305 310 315 320Gly Asp Val Asn Gly His Lys
Phe Ser Val Ser Gly Glu Gly Glu Gly 325 330 335Asp Ala Thr Tyr Gly
Lys Leu Thr Leu Lys Phe Ile Cys Thr Thr Gly 340 345 350Lys Leu Pro
Val Pro Trp Pro Thr Leu Val Thr Thr Leu Thr Tyr Gly 355 360 365Val
Gln Cys Phe Ser Arg Tyr Pro Asp His Met Lys Gln His Asp Phe 370 375
380Phe Lys Ser Ala Met Pro Glu Gly Tyr Ile Gln Glu Arg Thr Ile
Phe385 390 395 400Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu
Val Lys Phe Glu 405 410 415Gly Asp Thr Leu Val Asn Arg Ile Glu Leu
Lys Gly Ile Asp Phe Lys 420 425 430Glu Asp Gly Asn Ile Leu Gly His
Lys Leu Glu Tyr Asn Phe Asn Gly 435 440 445Gly Tyr Ala Phe Lys Tyr
Glu Asn Gly Lys Tyr Asp Ile Lys Asp Val 450 455 460Gly Val Asp Asn
Ala Gly Ala Lys Ala Gly Leu Thr Phe Leu Val Asp465 470 475 480Leu
Ile Lys Asn Lys His Met Asn Ala Asp Thr Asp Tyr Ser Ile Ala 485 490
495Glu Ala Ala Phe Asn Lys Gly Glu Thr Ala Met Thr Ile Asn Gly Pro
500 505 510Trp Ala Trp Ser Asn Ile Asp Thr Ser Lys Val Asn Tyr Gly
Val Thr 515 520 525Val Leu Pro Thr Phe Lys Gly Gln Pro Ser Lys Pro
Phe Val Gly Val 530 535 540Leu Ser Ala Gly Ile Asn Ala Ala Ser Pro
Asn Lys Glu Leu Ala Lys545 550 555 560Glu Phe Leu Glu Asn Tyr Leu
Leu Thr Asp Glu Gly Leu Glu Ala Val 565 570 575Asn Lys Asp Lys Pro
Leu Gly Ala Val Ala Leu Lys Ser Tyr Glu Glu 580 585 590Glu Leu Val
Lys Asp Pro Arg Ile Ala Ala Thr Met Glu Asn Ala Gln 595 600 605Lys
Gly Glu Ile Met Pro Asn Ile Pro Gln Met Ser Ala Phe Trp Tyr 610 615
620Ala Val Arg Thr Ala Val Ile Asn Ala Ala Ser Gly Arg Gln Thr
Val625 630 635 640Asp Glu Asp Leu Lys Asp Ala Gln Thr Arg Ile Thr
Lys Gly Ser His 645 650 655His His His His His Gly
6603659PRTArtificial Sequencerecombinant peptide biosensor 3Met Arg
Gly Ser His His His His His His Gly Met Ala Ser Met Thr1 5 10 15Gly
Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25
30Arg Trp Gly Ser Lys Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn
35 40 45Gly Asp Lys Gly Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe
Glu 50 55 60Lys Asp Thr Gly Ile Lys Val Thr Val Glu His Pro Asp Lys
Leu Glu65 70 75 80Glu Lys Phe Pro Gln Val Ala Ala Thr Gly Asp Gly
Pro Asp Ile Ile 85 90 95Phe Trp Ala His Asp Arg Phe Gly Gly Tyr Ala
Gln Ser Gly Leu Leu 100 105 110Ala Glu Ile Thr Pro Asp Lys Ala Phe
Gln Asp Lys Leu Tyr Pro Phe 115 120 125Thr Trp Asp Ala Val Arg Tyr
Asn Gly Lys Leu Ile Ala Tyr Pro Ile 130 135 140Ala Val Glu Ala Leu
Ser Leu Ile Tyr Asn Lys Asp Leu Leu Pro Asn145 150 155 160Pro Pro
Lys Thr Trp Glu Glu Ile Pro Ala Leu Asp Lys Glu Leu Lys 165 170
175Ala Lys Gly Lys Ser Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe
180 185 190Thr Trp Pro Leu Ile Ala Ala Asp Pro Cys Ser His Asn Val
Phe Ile 195 200 205Met Ala Asp Lys Gln Lys Asn Gly Ile Lys Ala Asn
Phe Lys Ile Arg 210 215 220His Asn Ile Glu Asp Gly Gly Val Gln Leu
Ala Tyr His Tyr Gln Gln225 230 235 240Asn Thr Pro Ile Gly Asp Gly
Pro Val Leu Leu Pro Asp Asn His Tyr 245 250 255Leu Ser Thr Gln Ser
Lys Leu Ser Lys Asp Pro Asn Glu Lys Arg Asp 260 265 270His Met Val
Leu Leu Glu Phe Val Thr Ala Ala Gly Ile Thr Leu Gly 275 280 285Met
Asp Glu Leu Tyr Lys Gly Gly Ser Met Val Ser Lys Gly Glu Glu 290 295
300Leu Phe Thr Gly Val Val Pro Ile Leu Val Glu Leu Asp Gly Asp
Val305 310 315 320Asn Gly His Lys Phe Ser Val Ser Gly Glu Gly Glu
Gly Asp Ala Thr 325 330 335Tyr Gly Lys Leu Thr Leu Lys Phe Ile Cys
Thr Thr Gly Lys Leu Pro 340 345 350Val Pro Trp Pro Thr Leu Val Thr
Thr Leu Thr Tyr Gly Val Gln Cys 355 360 365Phe Ser Arg Tyr Pro Asp
His Met Lys Gln His Asp Phe Phe Lys Ser 370 375 380Ala Met Pro Glu
Gly Tyr Ile Gln Glu Arg Thr Ile Phe Phe Lys Asp385 390 395 400Asp
Gly Asn Tyr Lys Thr Arg Ala Glu Val Lys Phe Glu Gly Asp Thr 405 410
415Leu Val Asn Arg Ile Glu Leu Lys Gly Ile Asp Phe Lys Glu Asp Gly
420 425 430Asn Ile Leu Gly His Lys Leu Glu Tyr Asn Phe Asn Gly Gly
Tyr Ala 435 440 445Phe Lys Tyr Glu Asn Gly Lys Tyr Asp Ile Lys Asp
Val Gly Val Asp 450 455 460Asn Ala Gly Ala Lys Ala Gly Leu Thr Phe
Leu Val Asp Leu Ile Lys465 470 475 480Asn Lys His Met Asn Ala Asp
Thr Asp Tyr Ser Ile Ala Glu Ala Ala 485 490 495Phe Asn Lys Gly Glu
Thr Ala Met Thr Ile Asn Gly Pro Trp Ala Trp 500 505 510Ser Asn Ile
Asp Thr Ser Lys Val Asn Tyr Gly Val Thr Val Leu Pro 515 520 525Thr
Phe Lys Gly Gln Pro Ser Lys Pro Phe Val Gly Val Leu Ser Ala 530 535
540Gly Ile Asn Ala Ala Ser Pro Asn Lys Glu Leu Ala Lys Glu Phe
Leu545 550 555 560Glu Asn Tyr Leu Leu Thr Asp Glu Gly Leu Glu Ala
Val Asn Lys Asp 565 570 575Lys Pro Leu Gly Ala Val Ala Leu Lys Ser
Tyr Glu Glu Glu Leu Val 580 585 590Asp Lys Pro Arg Ile Ala Ala Thr
Met Glu Asn Ala Gln Lys Gly Glu 595 600 605Ile Met Pro Asn Ile Pro
Gln Met Ser Ala Phe Trp Tyr Ala Val Arg 610 615 620Thr Ala Val Ile
Asn Ala Ala Ser Gly Arg Gln Thr Val Asp Glu Asp625 630 635 640Leu
Lys Asp Ala Gln Thr Arg Ile Thr Lys Gly Ser His His His His 645 650
655His His Gly4661PRTArtificial Sequencerecombinant peptide
biosensor 4Met Arg Gly Ser His His His His His His Gly Met Ala Ser
Met Thr1 5 10 15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp
Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys Ile Glu Glu Gly Lys Leu Val
Ile Trp Ile Asn 35 40 45Gly Asp Lys Gly Tyr Asn Gly Leu Ala Glu Val
Gly Lys Lys Phe Glu 50 55 60Lys Asp Thr Gly Ile Lys Val Thr Val Glu
His Pro Asp Lys Leu Glu65 70 75 80Glu Lys Phe Pro Gln Val Ala Ala
Thr Gly Asp Gly Pro Asp Ile Ile 85 90 95Phe Trp Ala His Asp Arg Phe
Gly Gly Tyr Ala Gln Ser Gly Leu Leu 100 105 110Ala Glu Ile Thr Pro
Asp Lys Ala Phe Gln Asp Lys Leu Tyr Pro Phe 115 120 125Thr Trp Asp
Ala Val Arg Tyr Asn Gly Lys Leu Ile Ala Tyr Pro Ile 130 135 140Ala
Val Glu Ala Leu Ser Leu Ile Tyr Asn Lys Asp Leu Leu Pro Asn145 150
155 160Pro Pro Lys Thr Trp Glu Glu Ile Pro Ala Leu Asp Lys Glu Leu
Lys 165 170 175Ala Lys Gly Lys Ser Ala Leu Met Phe Asn Leu Gln Glu
Pro Tyr Phe 180 185 190Thr Trp Pro Leu Ile Ala Ala Asp Gly Gly Tyr
Ala Phe Lys Tyr Glu 195 200 205Asn Gly Gly Ser His Asn Val Tyr Ile
Met Ala Asp Lys Gln Lys Asn 210 215 220Gly Ile Lys Ala Asn Phe Lys
Ile Arg His Asn Ile Glu Asp Gly Gly225 230 235 240Val Gln Leu Ala
Tyr His Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly 245 250 255Pro Val
Leu Leu Pro Asp Asn His Tyr Leu Ser Thr Gln Ser Lys Leu 260 265
270Ser Lys Asp Pro Asn Glu Lys Arg Asp His Met Val Leu Leu Glu Phe
275 280 285Val Thr Ala Ala Gly Ile Thr Leu Gly Met Asp Glu Leu Tyr
Lys Gly 290 295 300Gly Thr Gly Gly Ser Met Val Ser Lys Gly Glu Glu
Leu Phe Thr Gly305 310 315 320Val Val Pro Ile Leu Val Glu Leu Asp
Gly Asp Val Asn Gly His Lys 325 330 335Phe Ser Val Ser Gly Glu Gly
Glu Gly Asp Ala Thr Tyr Gly Lys Leu 340 345 350Thr Leu Lys Phe Ile
Cys Thr Thr Gly Lys Leu Pro Val Pro Trp Pro 355 360 365Thr Leu Val
Thr Thr Leu Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr 370 375 380Pro
Asp His Met Lys Gln His Asp Phe Phe Lys Ser Ala Met Pro Glu385 390
395 400Gly Tyr Ile Gln Glu Arg Thr Ile Phe Phe Lys Asp Asp Gly Asn
Tyr 405 410 415Lys Thr Arg Ala Glu Val Lys Phe Glu Gly Asp Thr Leu
Val Asn Arg 420 425 430Ile Glu Leu Lys Gly Ile Asp Phe Lys Glu Asp
Gly Asn Ile Leu Gly 435 440 445His Lys Leu Glu Tyr Asn Phe Asn Gly
Gly Lys Tyr Asp Ile Lys Asp 450 455
460Val Gly Asp Asn Ala Gly Ala Lys Ala Gly Leu Thr Phe Leu Val
Asp465 470 475 480Leu Ile Lys Asn Lys His Met Asn Ala Asp Thr Asp
Tyr Ser Ile Ala 485 490 495Glu Ala Ala Phe Asn Lys Gly Glu Thr Ala
Met Thr Ile Asn Gly Pro 500 505 510Trp Ala Trp Ser Asn Ile Asp Thr
Ser Lys Val Asn Tyr Gly Val Thr 515 520 525Val Leu Pro Thr Phe Lys
Gly Gln Pro Ser Lys Pro Phe Val Gly Val 530 535 540Leu Ser Ala Gly
Ile Asn Ala Ala Ser Pro Asn Lys Glu Leu Ala Lys545 550 555 560Glu
Phe Leu Glu Asn Tyr Leu Leu Thr Asp Glu Gly Leu Glu Ala Val 565 570
575Asn Lys Asp Lys Pro Leu Gly Ala Val Ala Leu Lys Ser Tyr Glu Glu
580 585 590Glu Leu Val Asp Lys Pro Arg Ile Ala Ala Thr Met Glu Asn
Ala Gln 595 600 605Lys Gly Glu Ile Met Pro Asn Ile Pro Gln Met Ser
Ala Phe Trp Tyr 610 615 620Ala Val Arg Thr Ala Val Ile Asn Ala Ala
Ser Gly Gln Thr Val Asp625 630 635 640Glu Asp Leu Lys Asp Ala Gln
Thr Arg Ile Thr Lys Gly Ser His His 645 650 655His His His His Gly
6605663PRTArtificial Sequencerecombinant peptide biosensor 5Met Arg
Gly Ser His His His His His His Gly Met Ala Ser Met Thr1 5 10 15Gly
Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25
30Arg Trp Gly Ser Lys Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn
35 40 45Gly Asp Lys Gly Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe
Glu 50 55 60Lys Asp Thr Gly Ile Lys Val Thr Val Glu His Pro Asp Lys
Leu Glu65 70 75 80Glu Lys Phe Pro Gln Val Ala Ala Thr Gly Asp Gly
Pro Asp Ile Ile 85 90 95Phe Trp Ala His Asp Arg Phe Gly Gly Tyr Ala
Gln Ser Gly Leu Leu 100 105 110Ala Glu Ile Thr Pro Asp Lys Ala Phe
Gln Asp Lys Leu Tyr Pro Phe 115 120 125Thr Trp Asp Ala Val Arg Tyr
Asn Gly Lys Leu Ile Ala Tyr Pro Ile 130 135 140Ala Val Glu Ala Leu
Ser Leu Ile Tyr Asn Lys Asp Leu Leu Pro Asn145 150 155 160Pro Pro
Lys Thr Trp Glu Glu Ile Pro Ala Leu Asp Lys Glu Leu Lys 165 170
175Ala Lys Gly Lys Ser Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe
180 185 190Thr Trp Pro Leu Ile Ala Ala Asp Gly Gly Tyr Ala Phe Lys
Tyr Glu 195 200 205Asn His Leu Ser His Asn Val Tyr Ile Met Ala Asp
Lys Gln Lys Asn 210 215 220Gly Ile Lys Ala Asn Phe Lys Ile Arg His
Asn Ile Glu Asp Gly Gly225 230 235 240Val Gln Leu Ala Tyr His Tyr
Gln Gln Asn Thr Pro Ile Gly Asp Gly 245 250 255Pro Val Leu Leu Pro
Asp Asn His Tyr Leu Ser Thr Gln Ser Lys Leu 260 265 270Ser Lys Asp
Pro Asn Glu Lys Arg Asp His Met Val Leu Leu Glu Phe 275 280 285Val
Thr Ala Ala Gly Ile Thr Leu Gly Met Asp Glu Leu Tyr Lys Gly 290 295
300Gly Thr Gly Gly Ser Met Val Ser Lys Gly Glu Glu Leu Phe Thr
Gly305 310 315 320Val Val Pro Ile Leu Val Glu Leu Asp Gly Asp Val
Asn Gly His Lys 325 330 335Phe Ser Val Ser Gly Glu Gly Glu Gly Asp
Ala Thr Tyr Gly Lys Leu 340 345 350Thr Leu Lys Phe Ile Cys Thr Thr
Gly Lys Leu Pro Val Pro Trp Pro 355 360 365Thr Leu Val Thr Thr Leu
Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr 370 375 380Pro Asp His Met
Lys Gln His Asp Phe Phe Lys Ser Ala Met Pro Glu385 390 395 400Gly
Tyr Ile Gln Glu Arg Thr Ile Phe Phe Lys Asp Asp Gly Asn Tyr 405 410
415Lys Thr Arg Ala Glu Val Lys Phe Glu Gly Asp Thr Leu Val Asn Arg
420 425 430Ile Glu Leu Lys Gly Ile Asp Phe Lys Glu Asp Gly Asn Ile
Leu Gly 435 440 445His Lys Leu Glu Tyr Asn Phe Asn Gly Gly Lys Tyr
Asp Ile Lys Asp 450 455 460Val Gly Val Asp Asn Ala Gly Ala Lys Ala
Gly Leu Thr Phe Leu Val465 470 475 480Asp Leu Ile Lys Asn Lys His
Met Asn Ala Asp Thr Asp Tyr Ser Ile 485 490 495Ala Glu Ala Ala Phe
Asn Lys Gly Glu Thr Ala Met Thr Ile Asn Gly 500 505 510Pro Trp Ala
Trp Ser Asn Ile Asp Thr Ser Lys Val Asn Tyr Gly Val 515 520 525Thr
Val Leu Pro Thr Phe Lys Gly Gln Pro Ser Lys Pro Phe Val Gly 530 535
540Val Leu Ser Ala Gly Ile Asn Ala Ala Ser Pro Asn Lys Glu Leu
Ala545 550 555 560Lys Glu Phe Leu Glu Asn Tyr Leu Leu Thr Asp Glu
Gly Leu Glu Ala 565 570 575Val Asn Lys Asp Lys Pro Leu Gly Ala Val
Ala Leu Lys Ser Tyr Glu 580 585 590Glu Glu Leu Val Lys Asp Pro Arg
Ile Ala Ala Thr Met Glu Asn Ala 595 600 605Gln Lys Gly Glu Ile Met
Pro Asn Ile Pro Gln Met Ser Ala Phe Trp 610 615 620Tyr Ala Val Arg
Thr Ala Val Ile Asn Ala Ala Ser Gly Arg Gln Thr625 630 635 640Val
Asp Glu Asp Leu Lys Asp Ala Gln Thr Arg Ile Thr Lys Gly Ser 645 650
655His His His His His His Gly 6606655PRTArtificial
Sequencerecombinant peptide biosensor 6Met Arg Gly Ser His His His
His His His Gly Met Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met Gly
Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys
Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn 35 40 45Gly Asp Lys Gly
Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe Glu 50 55 60Lys Asp Thr
Gly Ile Lys Val Thr Val Glu His Pro Asp Lys Leu Glu65 70 75 80Glu
Lys Phe Pro Gln Val Ala Ala Thr Gly Asp Gly Pro Asp Ile Ile 85 90
95Phe Trp Ala His Asp Arg Phe Gly Gly Tyr Ala Gln Ser Gly Leu Leu
100 105 110Ala Glu Ile Thr Pro Asp Lys Ala Phe Gln Asp Lys Leu Tyr
Pro Phe 115 120 125Thr Trp Asp Ala Val Arg Tyr Asn Gly Lys Leu Ile
Ala Tyr Pro Ile 130 135 140Ala Val Glu Ala Leu Ser Leu Ile Tyr Asn
Lys Asp Leu Leu Pro Asn145 150 155 160Pro Pro Lys Thr Trp Glu Glu
Ile Pro Ala Leu Asp Lys Glu Leu Lys 165 170 175Ala Lys Gly Lys Ser
Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe 180 185 190Thr Trp Pro
Leu Ile Ala Ala Asp Gly Gly Tyr Ala Phe Lys Tyr Glu 195 200 205Asn
Gly Lys Tyr Asp Ile Lys Asp Val Gly Val Asp Asn Ala Gly Ala 210 215
220Lys Ala Gly Leu Thr Phe Leu Val Asp Leu Ile Lys Asn Lys His
Met225 230 235 240Asn Ala Asp Thr Asp Tyr Ser Ile Ala Glu Ala Ala
Phe Asn Lys Gly 245 250 255Glu Thr Ala Met Thr Ile Asn Gly Pro Trp
Ala Trp Ser Asn Ile Asp 260 265 270Thr Ser Lys Val Asn Tyr Gly Val
Thr Val Leu Pro Thr Phe Lys Gly 275 280 285Gln Pro Ser Lys Pro Phe
Val Gly Val Leu Ser Ala Gly Ile Asn Ala 290 295 300Ala Ser Pro Asn
Lys Glu Leu Ala Lys Glu Phe Leu Glu Asn Tyr Leu305 310 315 320Leu
Thr Asp Glu Gly Leu Glu Ala Val Asn Lys Asp Lys Pro Leu Gly 325 330
335Ala Val Ala Leu Lys Ser Tyr Glu Glu Glu Leu Gly Gly Ser His Asn
340 345 350Val Tyr Ile Met Ala Asp Lys Gln Arg Asn Gly Ile Lys Ala
Asn Phe 355 360 365Lys Ile Arg His Asn Ile Glu Asp Gly Gly Val Gln
Leu Ala Tyr His 370 375 380Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly
Pro Val Leu Leu Pro Asp385 390 395 400Asn His Tyr Leu Ser Thr Gln
Ser Lys Leu Ser Lys Asp Pro Asn Glu 405 410 415Lys Arg Asp His Met
Val Leu Leu Glu Phe Val Thr Ala Ala Gly Ile 420 425 430Thr Leu Gly
Met Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser Met 435 440 445Val
Ser Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu Val 450 455
460Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser Gly
Glu465 470 475 480Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu
Lys Phe Ile Cys 485 490 495Thr Thr Gly Lys Leu Pro Val Pro Trp Pro
Thr Leu Val Thr Thr Leu 500 505 510Thr Tyr Gly Val Gln Cys Phe Ser
Arg Tyr Pro Asp His Met Lys Gln 515 520 525His Asp Phe Phe Lys Ser
Ala Met Pro Glu Gly Tyr Ile Gln Glu Arg 530 535 540Thr Ile Phe Phe
Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu Val545 550 555 560Lys
Phe Glu Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys Gly Ile 565 570
575Asp Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu Tyr Asn
580 585 590Phe Asn Gly Gly Ala Lys Asp Pro Arg Ile Ala Ala Thr Met
Glu Asn 595 600 605Ala Gln Lys Gly Glu Ile Met Pro Asn Ile Pro Gln
Met Ser Ala Phe 610 615 620Trp Tyr Ala Val Arg Thr Ala Val Ile Asn
Ala Ala Ser Gly Arg Gln625 630 635 640Thr Val Asp Glu Asp Leu Lys
Asp Ala Gln Thr Arg Ile Thr Lys 645 650 6557655PRTArtificial
Sequencerecombinant peptide biosensor 7Met Arg Gly Ser His His His
His His His Gly Met Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met Gly
Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys
Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn 35 40 45Gly Asp Lys Gly
Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe Glu 50 55 60Lys Asp Thr
Gly Ile Lys Val Thr Val Glu His Pro Asp Lys Leu Glu65 70 75 80Glu
Lys Phe Pro Gln Val Ala Ala Thr Gly Asp Gly Pro Asp Ile Ile 85 90
95Phe Trp Ala His Asp Arg Phe Gly Gly Tyr Ala Gln Ser Gly Leu Leu
100 105 110Ala Glu Ile Thr Pro Asp Lys Ala Phe Gln Asp Lys Leu Tyr
Pro Phe 115 120 125Thr Trp Asp Ala Val Arg Tyr Asn Gly Lys Leu Ile
Ala Tyr Pro Ile 130 135 140Ala Val Glu Ala Leu Ser Leu Ile Tyr Asn
Lys Asp Leu Leu Pro Asn145 150 155 160Pro Pro Lys Thr Trp Glu Glu
Ile Pro Ala Leu Asp Lys Glu Leu Lys 165 170 175Ala Lys Gly Lys Ser
Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe 180 185 190Thr Trp Pro
Leu Ile Ala Ala Asp Gly Gly Tyr Ala Phe Lys Tyr Glu 195 200 205Asn
Gly Lys Tyr Asp Ile Lys Asp Val Gly Val Asp Asn Ala Gly Ala 210 215
220Lys Ala Gly Leu Thr Phe Leu Val Asp Leu Ile Lys Asn Lys His
Met225 230 235 240Asn Ala Asp Thr Asp Tyr Ser Ile Ala Glu Ala Ala
Phe Asn Lys Gly 245 250 255Glu Thr Ala Met Thr Ile Asn Gly Pro Trp
Ala Trp Ser Asn Ile Asp 260 265 270Thr Ser Lys Val Asn Tyr Gly Val
Thr Val Leu Pro Thr Phe Lys Gly 275 280 285Gln Pro Ser Lys Pro Phe
Val Gly Val Leu Ser Ala Gly Ile Asn Ala 290 295 300Ala Ser Pro Asn
Lys Glu Leu Ala Lys Glu Phe Leu Glu Asn Tyr Leu305 310 315 320Leu
Thr Asp Glu Gly Leu Glu Ala Val Asn Lys Asp Lys Pro Leu Gly 325 330
335Ala Val Ala Leu Lys Ser Tyr Glu Glu Glu Leu Gly Gly Ser His Asn
340 345 350Val Tyr Ile Met Ala Asp Lys Gln Arg Asn Gly Ile Lys Ala
Asn Phe 355 360 365Lys Ile Arg His Asn Ile Glu Asp Gly Gly Val Gln
Leu Ala Tyr His 370 375 380Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly
Pro Val Leu Leu Pro Asp385 390 395 400Asn His Tyr Leu Ser Thr Gln
Ser Lys Leu Ser Lys Asp Pro Asn Glu 405 410 415Lys Arg Asp His Met
Val Leu Leu Glu Phe Val Thr Ala Ala Gly Ile 420 425 430Thr Leu Gly
Met Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser Met 435 440 445Val
Ser Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu Val 450 455
460Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser Gly
Glu465 470 475 480Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu
Lys Phe Ile Cys 485 490 495Thr Thr Gly Lys Leu Pro Val Pro Trp Pro
Thr Leu Val Thr Thr Leu 500 505 510Thr Tyr Gly Val Gln Cys Phe Ser
Arg Tyr Pro Asp His Met Lys Gln 515 520 525His Asp Phe Phe Lys Ser
Ala Met Pro Glu Gly Tyr Ile Gln Glu Arg 530 535 540Thr Ile Phe Phe
Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu Val545 550 555 560Lys
Phe Glu Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys Gly Ile 565 570
575Asp Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu Tyr Asn
580 585 590Phe Asn Asn Pro Ala Lys Asp Pro Arg Ile Ala Ala Thr Met
Glu Asn 595 600 605Ala Gln Lys Gly Glu Ile Met Pro Asn Ile Pro Gln
Met Ser Ala Phe 610 615 620Trp Tyr Ala Val Arg Thr Ala Val Ile Asn
Ala Ala Ser Gly Arg Gln625 630 635 640Thr Val Asp Glu Asp Leu Lys
Asp Ala Gln Thr Arg Ile Thr Lys 645 650 6558659PRTArtificial
Sequencerecombinant peptide biosensor 8Met Arg Gly Ser His His His
His His His Gly Met Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met Gly
Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys
Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn 35 40 45Gly Asp Lys Gly
Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe Glu 50 55 60Lys Asp Thr
Gly Ile Lys Val Thr Val Glu His Pro Asp Lys Leu Glu65 70 75 80Glu
Lys Phe Pro Gln Val Ala Ala Thr Gly Asp Gly Pro Asp Ile Ile 85 90
95Phe Trp Ala His Asp Arg Phe Gly Gly Tyr Ala Gln Ser Gly Leu Leu
100 105 110Ala Glu Ile Thr Pro Asp Lys Ala Phe Gln Asp Lys Leu Tyr
Pro Phe 115 120 125Thr Trp Asp Ala Val Arg Tyr Asn Gly Lys Leu Ile
Ala Tyr Pro Ile 130 135 140Ala Val Glu Ala Leu Ser Leu Ile Tyr Asn
Lys Asp Leu Leu Pro Asn145 150 155 160Pro Pro Lys Thr Trp Glu Glu
Ile Pro Ala Leu Asp Lys Glu Leu Lys 165 170 175Ala Lys Gly Lys Ser
Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe 180 185 190Thr Trp Pro
Leu Ile Ala Ala Asp Gly Gly Tyr Ala Phe Lys Tyr Glu 195 200 205Asn
Gly Lys Tyr Asp Ile Lys Asp Val Gly Val Asp Asn Ala Gly Ala 210 215
220Lys Ala Gly Leu Thr Phe Leu Val Asp Leu Ile Lys Asn Lys His
Met225 230 235 240Asn Ala Asp Thr Asp Tyr Ser Ile Ala Glu Ala Ala
Phe Asn Lys Gly 245 250 255Glu Thr Ala Met Thr Ile Asn Gly Pro Trp
Ala Trp Ser Asn Ile Asp 260 265
270Thr Ser Lys Val Asn Tyr Gly Val Thr Val Leu Pro Thr Phe Lys Gly
275 280 285Gln Pro Ser Lys Pro Phe Val Gly Val Leu Ser Ala Gly Ile
Asn Ala 290 295 300Ala Ser Pro Asn Lys Glu Leu Ala Lys Glu Phe Leu
Glu Asn Tyr Leu305 310 315 320Leu Thr Asp Glu Gly Leu Glu Ala Val
Asn Lys Asp Lys Pro Leu Gly 325 330 335Ala Val Ala Leu Lys Ser Tyr
Glu Glu Glu Leu Val Lys Asp Pro Arg 340 345 350Ser His Asn Val Tyr
Ile Met Ala Asp Lys Gln Lys Asn Gly Ile Lys 355 360 365Ala Asn Phe
Lys Ile Arg His Asn Ile Glu Asp Gly Gly Val Gln Leu 370 375 380Ala
Tyr His Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly Pro Val Leu385 390
395 400Leu Pro Asp Asn His Tyr Leu Ser Thr Gln Ser Lys Leu Ser Lys
Asp 405 410 415Pro Asn Glu Lys Arg Asp His Met Val Leu Leu Glu Phe
Val Thr Ala 420 425 430Ala Gly Ile Thr Leu Gly Met Asp Glu Leu Tyr
Lys Gly Gly Thr Gly 435 440 445Gly Ser Met Val Ser Lys Gly Glu Glu
Leu Phe Thr Gly Val Val Pro 450 455 460Ile Leu Val Glu Leu Asp Gly
Asp Val Asn Gly His Lys Phe Ser Val465 470 475 480Ser Gly Glu Gly
Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys 485 490 495Phe Ile
Cys Thr Thr Gly Lys Leu Pro Val Pro Trp Pro Thr Leu Val 500 505
510Thr Thr Leu Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr Pro Asp His
515 520 525Met Lys Gln His Asp Phe Phe Lys Ser Ala Met Pro Glu Gly
Tyr Ile 530 535 540Gln Glu Arg Thr Ile Phe Phe Lys Asp Asp Gly Asn
Tyr Lys Thr Arg545 550 555 560Ala Glu Val Lys Phe Glu Gly Asp Thr
Leu Val Asn Arg Ile Glu Leu 565 570 575Lys Gly Ile Asp Phe Lys Glu
Asp Gly Asn Ile Leu Gly His Lys Leu 580 585 590Glu Tyr Asn Phe Asn
Ala Ala Thr Met Glu Asn Ala Gln Lys Gly Glu 595 600 605Ile Met Pro
Asn Ile Pro Gln Met Ser Ala Phe Trp Tyr Ala Val Arg 610 615 620Thr
Ala Val Ile Asn Ala Ala Ser Gly Arg Gln Thr Val Asp Glu Asp625 630
635 640Leu Lys Asp Ala Gln Thr Arg Ile Thr Lys Gly Ser His His His
His 645 650 655His His Gly914PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 9Pro Leu Ile Ala Ala
Asp Gly Xaa Xaa Asn Val Tyr Ile Met1 5 101013PRTArtificial
Sequencesynthetic oligopeptideVARIANT7, 8Xaa = Any Amino Acid 10Pro
Leu Ile Ala Ala Asp Xaa Xaa Asn Val Tyr Ile Met1 5
101115PRTArtificial Sequencesynthetic oligopeptideVARIANT9, 10Xaa =
Any Amino Acid 11Pro Leu Ile Ala Ala Asp Gly Gly Xaa Xaa Asn Val
Tyr Ile Met1 5 10 151215PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 12Pro Leu Ile Ala Ala Asp
Gly Xaa Pro Asn Val Tyr Ile Met Gly1 5 10 151315PRTArtificial
Sequencesynthetic oligopeptideVARIANT9Xaa = Any Amino Acid 13Pro
Leu Ile Ala Ala Asp Gly Ile Xaa Asn Val Tyr Ile Met Gly1 5 10
151415PRTArtificial Sequencesynthetic oligopeptideVARIANT8Xaa = Any
Amino Acid 14Pro Leu Ile Ala Ala Asp Pro Xaa Ser His Asn Val Tyr
Ile Met1 5 10 151515PRTArtificial Sequencesynthetic
oligopeptideVARIANT7Xaa = Any Amino Acid 15Pro Leu Ile Ala Ala Asp
Xaa Pro Ser His Asn Val Tyr Ile Met1 5 10 151615PRTArtificial
Sequencesynthetic oligopeptideVARIANT7, 8Xaa = Any Amino Acid 16Pro
Leu Ile Ala Ala Asp Xaa Xaa Ser His Asn Val Tyr Ile Met1 5 10
151715PRTArtificial Sequencesynthetic oligopeptideVARIANT7, 8Xaa =
Any Amino Acid 17Pro Leu Ile Ala Ala Asp Xaa Xaa Ser His Asn Val
Phe Ile Met1 5 10 151815PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 18Pro Leu Ile Ala Ala Asp
Pro Xaa Ser His Asn Val Phe Ile Met1 5 10 151915PRTArtificial
Sequencesynthetic oligopeptideVARIANT8Xaa = Any Amino Acid 19Pro
Leu Ile Ala Ala Asp Pro Xaa Ser Tyr Asn Val Phe Ile Met1 5 10
152015PRTArtificial Sequencesynthetic oligopeptideVARIANT7, 8Xaa =
Any Amino Acid 20Pro Leu Ile Ala Ala Asp Xaa Xaa Ser Tyr Asn Val
Phe Ile Met1 5 10 152115PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 21Pro Leu Ile Ala Ala Asp
Pro Xaa Ser Tyr Asn Val Phe Ile Met1 5 10 152215PRTArtificial
Sequencesynthetic oligopeptideVARIANT7, 8Xaa = Any Amino Acid 22Pro
Leu Ile Ala Ala Asp Xaa Xaa Ser Tyr Asn Val Phe Ile Met1 5 10
152315PRTArtificial Sequencesynthetic oligopeptideVARIANT8, 10Xaa =
Any Amino Acid 23Pro Leu Ile Ala Ala Asp Pro Xaa Ser Xaa Asn Val
Tyr Ile Met1 5 10 152415PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 11Xaa = Any Amino Acid 24Pro Leu Ile Ala Ala
Asp Pro Xaa Ser His Xaa Val Tyr Ile Met1 5 10 152515PRTArtificial
Sequencesynthetic oligopeptideVARIANT8, 12Xaa = Any Amino Acid
25Pro Leu Ile Ala Ala Asp Pro Xaa Ser His Asn Xaa Tyr Ile Met1 5 10
152615PRTArtificial Sequencesynthetic oligopeptideVARIANT8, 13Xaa =
Any Amino Acid 26Pro Leu Ile Ala Ala Asp Pro Xaa Ser His Asn Val
Xaa Ile Met1 5 10 152716PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 27Lys Leu Glu Tyr Asn
Phe Asn Xaa Xaa Tyr Ala Phe Lys Tyr Glu Asn1 5 10
152815PRTArtificial Sequencesynthetic oligopeptideVARIANT8Xaa = Any
Amino Acid 28Lys Leu Glu Tyr Asn Phe Asn Xaa Tyr Ala Phe Lys Tyr
Glu Asn1 5 10 152914PRTArtificial Sequencesynthetic oligopeptide
29Lys Leu Glu Tyr Asn Phe Asn Tyr Ala Phe Lys Tyr Glu Asn1 5
103014PRTArtificial Sequencesynthetic oligopeptideVARIANT7Xaa = Any
Amino Acid 30Lys Leu Glu Tyr Asn Phe Xaa Tyr Ala Phe Lys Tyr Glu
Asn1 5 103114PRTArtificial Sequencesynthetic oligopeptideVARIANT6,
7Xaa = Any Amino Acid 31Lys Leu Glu Tyr Asn Xaa Xaa Tyr Ala Phe Lys
Tyr Glu Asn1 5 103214PRTArtificial Sequencesynthetic
oligopeptideVARIANT7Xaa = Any Amino Acid 32Lys Leu Glu Tyr Asn Trp
Xaa Tyr Ala Phe Lys Tyr Glu Asn1 5 103314PRTArtificial
Sequencesynthetic oligopeptideVARIANT6Xaa = Any Amino Acid 33Lys
Leu Glu Tyr Asn Xaa Lys Tyr Ala Phe Lys Tyr Glu Asn1 5
103416PRTArtificial Sequencesynthetic oligopeptideVARIANT9Xaa = Any
Amino Acid 34Lys Leu Glu Tyr Asn Phe Asn Pro Xaa Tyr Ala Phe Lys
Tyr Glu Asn1 5 10 153516PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 35Lys Leu Glu Tyr Asn Phe
Asn Xaa Pro Tyr Ala Phe Lys Tyr Glu Asn1 5 10 153615PRTArtificial
Sequencesynthetic oligopeptideVARIANT7, 8Xaa = Any Amino Acid 36Ala
Phe Lys Tyr Glu Asn Xaa Xaa Ser His Asn Val Tyr Ile Met1 5 10
153716PRTArtificial Sequencesynthetic oligopeptideVARIANT8, 9Xaa =
Any Amino Acid 37Lys Leu Glu Tyr Asn Phe Asn Xaa Xaa Lys Tyr Asp
Ile Lys Asp Val1 5 10 153815PRTArtificial Sequencesynthetic
oligopeptideVARIANT7, 8Xaa = Any Amino Acid 38Lys Ser Tyr Glu Glu
Leu Xaa Xaa Ser His Asn Val Tyr Ile Met1 5 10 153915PRTArtificial
Sequencesynthetic oligopeptideVARIANT8Xaa = Any Amino Acid 39Lys
Ser Tyr Glu Glu Leu Pro Xaa Ser His Asn Val Tyr Ile Met1 5 10
154015PRTArtificial Sequencesynthetic oligopeptideVARIANT7Xaa = Any
Amino Acid 40Lys Ser Tyr Glu Glu Leu Xaa Pro Ser His Asn Val Tyr
Ile Met1 5 10 154116PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 41Lys Leu Glu Tyr Asn
Phe Asn Xaa Xaa Ala Lys Asp Pro Arg Ile Ala1 5 10
154216PRTArtificial Sequencesynthetic oligopeptideVARIANT9Xaa = Any
Amino Acid 42Lys Leu Glu Tyr Asn Phe Asn Pro Xaa Ala Lys Asp Pro
Arg Ile Ala1 5 10 154316PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 43Lys Leu Glu Tyr Asn Phe
Asn Xaa Pro Ala Lys Asp Pro Arg Ile Ala1 5 10 154415PRTArtificial
Sequencesynthetic oligopeptideVARIANT8Xaa = Any Amino Acid 44Glu
Leu Ala Lys Asp Pro Arg Xaa Ser His Asn Val Tyr Ile Met1 5 10
154516PRTArtificial Sequencesynthetic oligopeptideVARIANT8, 9Xaa =
Any Amino Acid 45Glu Leu Ala Lys Asp Pro Arg Xaa Xaa Ser His Asn
Val Tyr Ile Met1 5 10 154617PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9, 10Xaa = Any Amino Acid 46Glu Leu Ala Lys
Asp Pro Arg Xaa Xaa Xaa Ser His Asn Val Tyr Ile1 5 10
15Met4715PRTArtificial Sequencesynthetic oligopeptideVARIANT8Xaa =
Any Amino Acid 47Lys Leu Glu Tyr Asn Phe Asn Xaa Ala Ala Thr Met
Glu Asn Ala1 5 10 154816PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 48Lys Leu Glu Tyr Asn
Phe Asn Xaa Xaa Ala Ala Thr Met Glu Asn Ala1 5 10
154917PRTArtificial Sequencesynthetic oligopeptideVARIANT8, 9,
10Xaa = Any Amino Acid 49Lys Leu Glu Tyr Asn Phe Asn Xaa Xaa Xaa
Ala Ala Thr Met Glu Asn1 5 10 15Ala50671PRTArtificial
Sequencerecombinant peptide biosensor 50Met Arg Gly Ser His His His
His His His Gly Met Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met Gly
Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys
Ile Glu Glu Gly Lys Val Val Ile Trp His Ala 35 40 45Met Gln Pro Asn
Glu Leu Glu Val Phe Gln Ser Leu Ala Glu Glu Tyr 50 55 60Met Ala Leu
Cys Pro Glu Val Glu Ile Val Phe Glu Gln Lys Pro Asn65 70 75 80Leu
Glu Asp Ala Leu Lys Ala Ala Ile Pro Thr Gly Gln Gly Pro Asp 85 90
95Leu Phe Ile Trp Ala His Asp Trp Ile Gly Lys Phe Ala Glu Ala Gly
100 105 110Leu Leu Glu Pro Ile Asp Glu Tyr Val Thr Glu Asp Leu Leu
Asn Glu 115 120 125Phe Ala Pro Met Ala Gln Asp Ala Met Gln Tyr Lys
Gly His Tyr Tyr 130 135 140Ala Leu Pro Phe Ala Ala Glu Thr Val Ala
Ile Ile Tyr Ser Lys Glu145 150 155 160Met Val Ser Glu Pro Pro Lys
Thr Phe Asp Glu Met Lys Ala Ile Met 165 170 175Glu Lys Tyr Tyr Asp
Pro Ala Asn Glu Lys Tyr Gly Ile Ala Trp Pro 180 185 190Ile Asn Ala
Tyr Phe Ile Ser Ala Ile Ala Gln Ala Phe Gly Gly Ser 195 200 205His
Asn Val Tyr Ile Met Ala Asp Lys Gln Lys Asn Gly Ile Lys Ala 210 215
220Asn Phe Lys Ile Arg His Asn Ile Glu Asp Gly Gly Val Gln Leu
Ala225 230 235 240Tyr His Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly
Pro Val Leu Leu 245 250 255Pro Asp Asn His Tyr Leu Ser Thr Gln Ser
Lys Leu Ser Lys Asp Pro 260 265 270Asn Glu Lys Arg Asp His Met Val
Leu Leu Glu Phe Val Thr Ala Ala 275 280 285Gly Ile Thr Leu Gly Met
Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly 290 295 300Ser Met Val Ser
Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile305 310 315 320Leu
Val Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser 325 330
335Gly Glu Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys Phe
340 345 350Ile Cys Thr Thr Gly Lys Leu Pro Val Pro Trp Pro Thr Leu
Val Thr 355 360 365Thr Leu Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr
Pro Asp His Met 370 375 380Lys Gln His Asp Phe Phe Lys Ser Ala Met
Pro Glu Gly Tyr Ile Gln385 390 395 400Glu Arg Thr Ile Phe Phe Lys
Asp Asp Gly Asn Tyr Lys Thr Arg Ala 405 410 415Glu Val Lys Phe Glu
Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys 420 425 430Gly Ile Asp
Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu 435 440 445Tyr
Asn Phe Asn Gly Gly Tyr Tyr Phe Asp Asp Lys Thr Glu Gln Pro 450 455
460Gly Leu Asp Lys Pro Glu Thr Ile Glu Gly Phe Lys Phe Phe Phe
Thr465 470 475 480Glu Ile Trp Pro Tyr Met Ala Pro Thr Gly Asp Tyr
Asn Thr Gln Gln 485 490 495Ser Ile Phe Leu Glu Gly Arg Ala Pro Met
Met Val Asn Gly Pro Trp 500 505 510Ser Ile Asn Asp Val Lys Lys Ala
Gly Ile Asn Phe Gly Val Val Pro 515 520 525Leu Pro Pro Ile Ile Lys
Asp Gly Lys Glu Tyr Trp Pro Arg Pro Tyr 530 535 540Gly Gly Val Lys
Leu Ile Tyr Phe Ala Ala Gly Ile Lys Asn Lys Asp545 550 555 560Ala
Ala Trp Lys Phe Ala Lys Trp Leu Thr Thr Ser Glu Glu Ser Ile 565 570
575Lys Thr Leu Ala Leu Glu Leu Gly Tyr Ile Pro Val Leu Thr Lys Val
580 585 590Leu Asp Asp Pro Glu Ile Lys Asn Asp Pro Val Ile Tyr Gly
Phe Gly 595 600 605Gln Ala Val Gln His Ala Tyr Leu Met Pro Lys Ser
Pro Lys Met Ser 610 615 620Ala Val Trp Gly Gly Val Asp Gly Ala Ile
Asn Glu Ile Leu Gln Asp625 630 635 640Pro Gln Asn Ala Asp Ile Glu
Gly Ile Leu Lys Lys Tyr Gln Gln Glu 645 650 655Ile Leu Asn Asn Met
Gln Gly Ser His His His His His His Gly 660 665
67051671PRTArtificial Sequencerecombinant peptide biosensor 51Met
Arg Gly Ser His His His His His His Gly Met Ala Ser Met Thr1 5 10
15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp
20 25 30Arg Trp Gly Ser Lys Ile Glu Glu Gly Lys Val Val Ile Trp His
Ala 35 40 45Met Gln Pro Asn Glu Leu Glu Val Phe Gln Ser Leu Ala Glu
Glu Tyr 50 55 60Met Ala Leu Cys Pro Glu Val Glu Ile Val Phe Glu Gln
Lys Pro Asn65 70 75 80Leu Glu Asp Ala Leu Lys Ala Ala Ile Pro Thr
Gly Gln Gly Pro Asp 85 90 95Leu Phe Ile Trp Ala His Asp Trp Ile Gly
Lys Phe Ala Glu Ala Gly 100 105 110Leu Leu Glu Pro Ile Asp Glu Tyr
Val Thr Glu Asp Leu Leu Asn Glu 115 120 125Phe Ala Pro Met Ala Gln
Asp Ala Met Gln Tyr Lys Gly His Tyr Tyr 130 135 140Ala Leu Pro Phe
Ala Ala Glu Thr Val Ala Ile Ile Tyr Ser Lys Glu145 150 155 160Met
Val Ser Glu Pro Pro Lys Thr Phe Asp Glu Met Lys Ala Ile Met 165 170
175Glu Lys Tyr Tyr Asp Pro Ala Asn Glu Lys Tyr Gly Ile Ala Trp Pro
180 185 190Ile Asn Ala Tyr Phe Ile Ser Ala Ile Ala Gln Ala Phe Gly
Gly Ser 195 200 205His Asn Val Tyr Ile Met Ala Asp Lys Gln Lys Asn
Gly Ile Lys Ala 210 215 220Asn Phe Lys Ile Arg His Asn Ile Glu Asp
Gly Gly Val Gln Leu Ala225 230 235 240Tyr His Tyr Gln Gln Asn Thr
Pro Ile Gly Asp Gly Pro Val Leu Leu 245 250 255Pro Asp Asn His Tyr
Leu Ser Thr Gln Ser Lys Leu Ser Lys Asp Pro 260 265 270Asn Glu Lys
Arg Asp His Met Val Leu Leu Glu Phe Val Thr Ala Ala 275 280 285Gly
Ile Thr Leu Gly Met Asp
Glu Leu Tyr Lys Gly Gly Thr Gly Gly 290 295 300Ser Met Val Ser Lys
Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile305 310 315 320Leu Val
Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser 325 330
335Gly Glu Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys Phe
340 345 350Ile Cys Thr Thr Gly Lys Leu Pro Val Pro Trp Pro Thr Leu
Val Thr 355 360 365Thr Leu Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr
Pro Asp His Met 370 375 380Lys Gln His Asp Phe Phe Lys Ser Ala Met
Pro Glu Gly Tyr Ile Gln385 390 395 400Glu Arg Thr Ile Phe Phe Lys
Asp Asp Gly Asn Tyr Lys Thr Arg Ala 405 410 415Glu Val Lys Phe Glu
Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys 420 425 430Gly Ile Asp
Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu 435 440 445Tyr
Asn Phe Asn Phe Glu Tyr Tyr Phe Asp Asp Lys Thr Glu Gln Pro 450 455
460Gly Leu Asp Lys Pro Glu Thr Ile Glu Gly Phe Lys Phe Phe Phe
Thr465 470 475 480Glu Ile Trp Pro Tyr Met Ala Pro Thr Gly Asp Tyr
Asn Thr Gln Gln 485 490 495Ser Ile Phe Leu Glu Gly Arg Ala Pro Met
Met Val Asn Gly Pro Trp 500 505 510Ser Ile Asn Asp Val Lys Lys Ala
Gly Ile Asn Phe Gly Val Val Pro 515 520 525Leu Pro Pro Ile Ile Lys
Asp Gly Lys Glu Tyr Trp Pro Arg Pro Tyr 530 535 540Gly Gly Val Lys
Leu Ile Tyr Phe Ala Ala Gly Ile Lys Asn Lys Asp545 550 555 560Ala
Ala Trp Lys Phe Ala Lys Trp Leu Thr Thr Ser Glu Glu Ser Ile 565 570
575Lys Thr Leu Ala Leu Glu Leu Gly Tyr Ile Pro Val Leu Thr Lys Val
580 585 590Leu Asp Asp Pro Glu Ile Lys Asn Asp Pro Val Ile Tyr Gly
Phe Gly 595 600 605Gln Ala Val Gln His Ala Tyr Leu Met Pro Lys Ser
Pro Lys Met Ser 610 615 620Ala Val Trp Gly Gly Val Asp Gly Ala Ile
Asn Glu Ile Leu Gln Asp625 630 635 640Pro Gln Asn Ala Asp Ile Glu
Gly Ile Leu Lys Lys Tyr Gln Gln Glu 645 650 655Ile Leu Asn Asn Met
Gln Gly Ser His His His His His His Gly 660 665
67052662PRTArtificial Sequencerecombinant peptide biosensor 52Met
Arg Gly Ser His His His His His His Gly Met Ala Ser Met Thr1 5 10
15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp
20 25 30Arg Trp Gly Ser Lys Ile Glu Glu Gly Lys Val Val Ile Trp His
Ala 35 40 45Met Gln Pro Asn Glu Leu Glu Val Phe Gln Ser Leu Ala Glu
Glu Tyr 50 55 60Met Ala Leu Cys Pro Glu Val Glu Ile Val Phe Glu Gln
Lys Pro Asn65 70 75 80Leu Glu Asp Ala Leu Lys Ala Ala Ile Pro Thr
Gly Gln Gly Pro Asp 85 90 95Leu Phe Ile Trp Ala His Asp Trp Ile Gly
Lys Phe Ala Glu Ala Gly 100 105 110Leu Leu Glu Pro Ile Asp Glu Tyr
Val Thr Glu Asp Leu Leu Asn Glu 115 120 125Phe Ala Pro Met Ala Gln
Asp Ala Met Gln Tyr Lys Gly His Tyr Tyr 130 135 140Ala Leu Pro Phe
Ala Ala Glu Thr Val Ala Ile Ile Tyr Ser Lys Glu145 150 155 160Met
Val Ser Glu Pro Pro Lys Thr Phe Asp Glu Met Lys Ala Ile Met 165 170
175Glu Lys Tyr Tyr Asp Pro Ala Asn Glu Lys Tyr Gly Ile Ala Trp Pro
180 185 190Ile Asn Ala Tyr Phe Ile Ser Ala Ile Ala Gln Ala Phe Gly
Gly Tyr 195 200 205Tyr Phe Asp Asp Lys Thr Glu Gln Pro Gly Leu Asp
Lys Pro Glu Thr 210 215 220Ile Glu Gly Phe Lys Phe Phe Phe Thr Glu
Ile Trp Pro Tyr Met Ala225 230 235 240Pro Thr Gly Asp Tyr Asn Thr
Gln Gln Ser Ile Phe Leu Glu Gly Arg 245 250 255Ala Pro Met Met Val
Asn Gly Pro Trp Ser Ile Asn Asp Val Lys Lys 260 265 270Ala Gly Ile
Asn Phe Gly Val Val Pro Leu Pro Pro Ile Ile Lys Asp 275 280 285Gly
Lys Glu Tyr Trp Pro Arg Pro Tyr Gly Gly Val Lys Leu Ile Tyr 290 295
300Phe Ala Ala Gly Ile Lys Asn Lys Asp Ala Ala Trp Lys Phe Ala
Lys305 310 315 320Trp Leu Thr Thr Ser Glu Glu Ser Ile Lys Thr Leu
Ala Leu Glu Leu 325 330 335Gly Tyr Ile Pro Val Leu Thr Lys Val Leu
Asp Asp Pro Glu Ile Ser 340 345 350His Asn Val Tyr Ile Met Ala Asp
Lys Gln Lys Asn Gly Ile Lys Ala 355 360 365Asn Phe Lys Ile Arg His
Asn Ile Glu Asp Gly Gly Val Gln Leu Ala 370 375 380Tyr His Tyr Gln
Gln Asn Thr Pro Ile Gly Asp Gly Pro Val Leu Leu385 390 395 400Pro
Asp Asn His Tyr Leu Ser Thr Gln Ser Lys Leu Ser Lys Asp Pro 405 410
415Asn Glu Lys Arg Asp His Met Val Leu Leu Glu Phe Val Thr Ala Ala
420 425 430Gly Ile Thr Leu Gly Met Asp Glu Leu Tyr Lys Gly Gly Thr
Gly Gly 435 440 445Ser Met Val Ser Lys Gly Glu Glu Leu Phe Thr Gly
Val Val Pro Ile 450 455 460Leu Val Glu Leu Asp Gly Asp Val Asn Gly
His Lys Phe Ser Val Ser465 470 475 480Gly Glu Gly Glu Gly Asp Ala
Thr Tyr Gly Lys Leu Thr Leu Lys Phe 485 490 495Ile Cys Thr Thr Gly
Lys Leu Pro Val Pro Trp Pro Thr Leu Val Thr 500 505 510Thr Leu Thr
Tyr Gly Val Gln Cys Phe Ser Arg Tyr Pro Asp His Met 515 520 525Lys
Gln His Asp Phe Phe Lys Ser Ala Met Pro Glu Gly Tyr Ile Gln 530 535
540Glu Arg Thr Ile Phe Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg
Ala545 550 555 560Glu Val Lys Phe Glu Gly Asp Thr Leu Val Asn Arg
Ile Glu Leu Lys 565 570 575Gly Ile Asp Phe Lys Glu Asp Gly Asn Ile
Leu Gly His Lys Leu Glu 580 585 590Tyr Asn Phe Asn Lys Asn Asp Pro
Val Ile Tyr Gly Phe Gly Gln Ala 595 600 605Val Gln His Ala Tyr Leu
Met Pro Lys Ser Pro Lys Met Ser Ala Val 610 615 620Trp Gly Gly Val
Asp Gly Ala Ile Asn Glu Ile Leu Gln Asp Pro Gln625 630 635 640Asn
Ala Asp Ile Glu Gly Ile Leu Lys Lys Tyr Gln Gln Glu Ile Leu 645 650
655Asn Asn Met Gln Gly Ser 66053664PRTArtificial
Sequencerecombinant peptide biosensor 53Met Arg Gly Ser His His His
His His His Gly Met Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met Gly
Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys
Ile Glu Glu Gly Lys Val Val Ile Trp His Ala 35 40 45Met Gln Pro Asn
Glu Leu Glu Val Phe Gln Ser Leu Ala Glu Glu Tyr 50 55 60Met Ala Leu
Cys Pro Glu Val Glu Ile Val Phe Glu Gln Lys Pro Asn65 70 75 80Leu
Glu Asp Ala Leu Lys Ala Ala Ile Pro Thr Gly Gln Gly Pro Asp 85 90
95Leu Phe Ile Trp Ala His Asp Trp Ile Gly Lys Phe Ala Glu Ala Gly
100 105 110Leu Leu Glu Pro Ile Asp Glu Tyr Val Thr Glu Asp Leu Leu
Asn Glu 115 120 125Phe Ala Pro Met Ala Gln Asp Ala Met Gln Tyr Lys
Gly His Tyr Tyr 130 135 140Ala Leu Pro Phe Ala Ala Glu Thr Val Ala
Ile Ile Tyr Ser Lys Glu145 150 155 160Met Val Ser Glu Pro Pro Lys
Thr Phe Asp Glu Met Lys Ala Ile Met 165 170 175Glu Lys Tyr Tyr Asp
Pro Ala Asn Glu Lys Tyr Gly Ile Ala Trp Pro 180 185 190Ile Asn Ala
Tyr Phe Ile Ser Ala Ile Ala Gln Ala Phe Gly Gly Tyr 195 200 205Tyr
Phe Asp Asp Lys Thr Glu Gln Pro Gly Leu Asp Lys Pro Glu Thr 210 215
220Ile Glu Gly Phe Lys Phe Phe Phe Thr Glu Ile Trp Pro Tyr Met
Ala225 230 235 240Pro Thr Gly Asp Tyr Asn Thr Gln Gln Ser Ile Phe
Leu Glu Gly Arg 245 250 255Ala Pro Met Met Val Asn Gly Pro Trp Ser
Ile Asn Asp Val Lys Lys 260 265 270Ala Gly Ile Asn Phe Gly Val Val
Pro Leu Pro Pro Ile Ile Lys Asp 275 280 285Gly Lys Glu Tyr Trp Pro
Arg Pro Tyr Gly Gly Val Lys Leu Ile Tyr 290 295 300Phe Ala Ala Gly
Ile Lys Asn Lys Asp Ala Ala Trp Lys Phe Ala Lys305 310 315 320Trp
Leu Thr Thr Ser Glu Glu Ser Ile Lys Thr Leu Ala Leu Glu Leu 325 330
335Gly Tyr Ile Pro Val Leu Thr Lys Val Leu Asp Asp Pro Glu Ile Pro
340 345 350Pro Ser His Asn Val Tyr Ile Met Ala Asp Lys Gln Lys Asn
Gly Ile 355 360 365Lys Ala Asn Phe Lys Ile Arg His Asn Ile Glu Asp
Gly Gly Val Gln 370 375 380Leu Ala Tyr His Tyr Gln Gln Asn Thr Pro
Ile Gly Asp Gly Pro Val385 390 395 400Leu Leu Pro Asp Asn His Tyr
Leu Ser Thr Gln Ser Lys Leu Ser Lys 405 410 415Asp Pro Asn Glu Lys
Arg Asp His Met Val Leu Leu Glu Phe Val Thr 420 425 430Ala Ala Gly
Ile Thr Leu Gly Met Asp Glu Leu Tyr Lys Gly Gly Thr 435 440 445Gly
Gly Ser Met Val Ser Lys Gly Glu Glu Leu Phe Thr Gly Val Val 450 455
460Pro Ile Leu Val Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe
Ser465 470 475 480Val Ser Gly Glu Gly Glu Gly Asp Ala Thr Tyr Gly
Lys Leu Thr Leu 485 490 495Lys Phe Ile Cys Thr Thr Gly Lys Leu Pro
Val Pro Trp Pro Thr Leu 500 505 510Val Thr Thr Leu Thr Tyr Gly Val
Gln Cys Phe Ser Arg Tyr Pro Asp 515 520 525His Met Lys Gln His Asp
Phe Phe Lys Ser Ala Met Pro Glu Gly Tyr 530 535 540Ile Gln Glu Arg
Thr Ile Phe Phe Lys Asp Asp Gly Asn Tyr Lys Thr545 550 555 560Arg
Ala Glu Val Lys Phe Glu Gly Asp Thr Leu Val Asn Arg Ile Glu 565 570
575Leu Lys Gly Ile Asp Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys
580 585 590Leu Glu Tyr Asn Phe Asn Lys Asn Asp Pro Val Ile Tyr Gly
Phe Gly 595 600 605Gln Ala Val Gln His Ala Tyr Leu Met Pro Lys Ser
Pro Lys Met Ser 610 615 620Ala Val Trp Gly Gly Val Asp Gly Ala Ile
Asn Glu Ile Leu Gln Asp625 630 635 640Pro Gln Asn Ala Asp Ile Glu
Gly Ile Leu Lys Lys Tyr Gln Gln Glu 645 650 655Ile Leu Asn Asn Met
Gln Gly Ser 6605416PRTArtificial Sequencesynthetic
oligopeptideVARIANT7, 8Xaa = Any Amino Acid 54Ala Ile Ala Gln Ala
Phe Xaa Xaa Ser His Asn Val Tyr Ile Met Ala1 5 10
155516PRTArtificial Sequencesynthetic oligopeptideVARIANT8Xaa = Any
Amino Acid 55Ala Ile Ala Gln Ala Phe Pro Xaa Ser His Asn Val Tyr
Ile Met Ala1 5 10 155617PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 56Lys Leu Glu Tyr Asn
Phe Asn Xaa Xaa Tyr Tyr Phe Asp Asp Lys Thr1 5 10
15Glu5714PRTArtificial Sequencesynthetic oligopeptideVARIANT7, 8Xaa
= Any Amino Acid 57Val Leu Asp Asp Pro Glu Xaa Xaa His Asn Val Tyr
Ile Met1 5 105816PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 58Val Leu Asp Asp Pro
Glu Ile Xaa Xaa Ser His Asn Val Tyr Ile Met1 5 10
155914PRTArtificial Sequencesynthetic oligopeptideVARIANT7, 8Xaa =
Any Amino Acid 59Lys Leu Glu Tyr Asn Phe Xaa Xaa Asn Asp Pro Val
Ile Tyr1 5 106016PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 60Lys Leu Glu Tyr Asn Phe
Asn Xaa Pro Lys Asn Asp Pro Val Ile Tyr1 5 10 156116PRTArtificial
Sequencesynthetic oligopeptideVARIANT9Xaa = Any Amino Acid 61Lys
Leu Glu Tyr Asn Phe Asn Pro Xaa Lys Asn Asp Pro Val Ile Tyr1 5 10
1562556PRTArtificial Sequencerecombinant peptide biosensor 62Met
Arg Gly Ser His His His His His His Gly Met Ala Ser Met Thr1 5 10
15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp
20 25 30Arg Trp Gly Ser Ala Ala Gly Ser Thr Leu Asp Lys Ile Ala Lys
Asn 35 40 45Gly Val Ile Val Val Gly His Arg Glu Ser Ser Val Pro Phe
Ser Tyr 50 55 60Tyr Asp Asn Gln Gln Lys Val Val Gly Tyr Ser Gln Asp
Tyr Ser Asn65 70 75 80Ala Ile Val Glu Ala Val Lys Lys Lys Leu Asn
Lys Pro Asp Leu Gln 85 90 95Val Lys Leu Ile Pro Ile Thr Ser Gln Asn
Arg Ile Pro Leu Leu Gln 100 105 110Asn Gly Thr Phe Asp Phe Glu Cys
Gly Ser Thr Thr Asn Asn Val Glu 115 120 125Arg Gln Lys Gln Ala Ala
Phe Ser Asp Thr Ile Phe Val Val Gly Thr 130 135 140Arg Leu Leu Thr
Lys Lys Gly Gly Asp Ile Lys Asp Phe Ala Asn Leu145 150 155 160Lys
Asp Lys Ala Val Val Val Thr Ser Gly Thr Thr Ser Glu Val Leu 165 170
175Leu Asn Lys Leu Asn Glu Glu Gln Lys Met Asn Met Arg Ile Ile Ser
180 185 190Ala Lys Asp His Gly Asp Ser Phe Arg Thr Leu Glu Ser Gly
Arg Ala 195 200 205Val Ala Phe Met Met Asp Asp Val Leu Leu Ala Gly
Glu Arg Ala Lys 210 215 220Ala Lys Lys Pro Asp Asn Trp Glu Ile Val
Gly Lys Pro Gln Ser Gln225 230 235 240Glu Ala Tyr Gly Cys Met Leu
Arg Lys Asp Asp Pro Gln Phe Lys Lys 245 250 255Leu Met Asp Asp Thr
Ile Ala Gln Val Gln Thr Ser Gly Glu Ala Glu 260 265 270Lys Trp Phe
Asp Lys Trp Phe Lys Asn Pro Ile Leu Val Ser His Asn 275 280 285Val
Tyr Ile Met Ala Asp Lys Gln Lys Asn Gly Ile Lys Ala Asn Phe 290 295
300Lys Ile Arg His Asn Ile Glu Asp Gly Gly Val Gln Leu Ala Tyr
His305 310 315 320Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly Pro Val
Leu Leu Pro Asp 325 330 335Asn His Tyr Leu Ser Thr Gln Ser Lys Leu
Ser Lys Asp Pro Asn Glu 340 345 350Lys Arg Asp His Met Val Leu Leu
Glu Phe Val Thr Ala Ala Gly Ile 355 360 365Thr Leu Gly Met Asp Glu
Leu Tyr Lys Gly Gly Thr Gly Gly Ser Met 370 375 380Val Ser Lys Gly
Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu Val385 390 395 400Glu
Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser Gly Glu 405 410
415Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys Phe Ile Cys
420 425 430Thr Thr Gly Lys Leu Pro Val Pro Trp Pro Thr Leu Val Thr
Thr Leu 435 440 445Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr Pro Asp
His Met Lys Gln 450 455 460His Asp Phe Phe Lys Ser Ala Met Pro Glu
Gly Tyr Ile Gln Glu Arg465 470 475 480Thr Ile Phe Phe Lys Asp Asp
Gly Asn Tyr Lys Thr Arg Ala Glu Val 485 490 495Lys Phe Glu Gly Asp
Thr Leu Val Asn Arg Ile Glu Leu Lys Gly Ile 500 505 510Asp Phe Lys
Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu
Tyr Asn 515 520 525Phe Asn Asn Pro Leu Asn Met Asn Phe Glu Leu Ser
Asp Glu Met Lys 530 535 540Ala Leu Phe Lys Glu Pro Asn Asp Lys Ala
Leu Lys545 550 55563556PRTArtificial Sequencerecombinant peptide
biosensor 63Met Arg Gly Ser His His His His His His Gly Met Ala Ser
Met Thr1 5 10 15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp
Asp Lys Asp 20 25 30Arg Trp Gly Ser Ala Ala Gly Ser Thr Leu Asp Lys
Ile Ala Lys Asn 35 40 45Gly Val Ile Val Val Gly His Arg Glu Ser Ser
Val Pro Phe Ser Tyr 50 55 60Tyr Asp Asn Gln Gln Lys Val Val Gly Tyr
Ser Gln Asp Tyr Ser Asn65 70 75 80Ala Ile Val Glu Ala Val Lys Lys
Lys Leu Asn Lys Pro Asp Leu Gln 85 90 95Val Lys Leu Ile Pro Ile Thr
Ser Gln Asn Arg Ile Pro Leu Leu Gln 100 105 110Asn Gly Thr Phe Asp
Phe Glu Cys Gly Ser Thr Thr Asn Asn Val Glu 115 120 125Arg Gln Lys
Gln Ala Ala Phe Ser Asp Thr Ile Phe Val Val Gly Thr 130 135 140Arg
Leu Leu Thr Lys Lys Gly Gly Asp Ile Lys Asp Phe Ala Asn Leu145 150
155 160Lys Asp Lys Ala Val Val Val Thr Ser Gly Thr Thr Ser Glu Val
Leu 165 170 175Leu Asn Lys Leu Asn Glu Glu Gln Lys Met Asn Met Arg
Ile Ile Ser 180 185 190Ala Lys Asp His Gly Asp Ser Phe Arg Thr Leu
Glu Ser Gly Arg Ala 195 200 205Val Ala Phe Met Met Asp Asp Val Leu
Leu Ala Gly Glu Arg Ala Lys 210 215 220Ala Lys Lys Pro Asp Asn Trp
Glu Ile Val Gly Lys Pro Gln Ser Gln225 230 235 240Glu Ala Tyr Gly
Cys Met Leu Arg Lys Asp Asp Pro Gln Phe Lys Lys 245 250 255Leu Met
Asp Asp Thr Ile Ala Gln Val Gln Thr Ser Gly Glu Ala Glu 260 265
270Lys Trp Phe Asp Lys Trp Phe Lys Asn Pro Ile Leu Val Ser His Asn
275 280 285Val Tyr Ile Met Ala Asp Lys Gln Lys Asn Gly Ile Lys Ala
Asn Phe 290 295 300Lys Ile Arg His Asn Ile Glu Asp Gly Gly Val Gln
Leu Ala Tyr His305 310 315 320Tyr Gln Gln Asn Thr Pro Ile Gly Asp
Gly Pro Val Leu Leu Pro Asp 325 330 335Asn His Tyr Leu Ser Thr Gln
Ser Lys Leu Ser Lys Asp Pro Asn Glu 340 345 350Lys Arg Asp His Met
Val Leu Leu Glu Phe Val Thr Ala Ala Gly Ile 355 360 365Thr Leu Gly
Met Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser Met 370 375 380Val
Ser Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu Val385 390
395 400Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser Gly
Glu 405 410 415Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys
Phe Ile Cys 420 425 430Thr Thr Gly Lys Leu Pro Val Pro Trp Pro Thr
Leu Val Thr Thr Leu 435 440 445Thr Tyr Gly Val Gln Cys Phe Ser Arg
Tyr Pro Asp His Met Lys Gln 450 455 460His Asp Phe Phe Lys Ser Ala
Met Pro Glu Gly Tyr Ile Gln Glu Arg465 470 475 480Thr Ile Phe Phe
Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu Val 485 490 495Lys Phe
Glu Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys Gly Ile 500 505
510Asp Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu Tyr Asn
515 520 525Phe Asn Asn Pro Leu Asn Met Asn Phe Glu Leu Ser Asp Glu
Met Lys 530 535 540Ala Leu Phe Lys Glu Pro Asn Asp Lys Ala Leu
Lys545 550 5556416PRTArtificial Sequencerecombinant peptide
biosensorVARIANT8Xaa = Any Amino Acid 64Phe Lys Asn Pro Ile Pro Pro
Xaa Ser His Asn Val Tyr Ile Met Ala1 5 10 156517PRTArtificial
Sequencerecombinant peptide biosensorVARIANT8, 9Xaa = Any Amino
Acid 65Phe Lys Asn Pro Ile Pro Pro Xaa Xaa Ser His Asn Val Tyr Ile
Met1 5 10 15Ala6617PRTArtificial Sequencerecombinant peptide
biosensorVARIANT9Xaa = Any Amino Acid 66Phe Lys Asn Pro Ile Pro Pro
Pro Xaa Ser His Asn Val Tyr Ile Met1 5 10 15Ala6717PRTArtificial
Sequencerecombinant peptide biosensorVARIANT8Xaa = Any Amino Acid
67Phe Lys Asn Pro Ile Pro Pro Xaa Pro Ser His Asn Val Tyr Ile Met1
5 10 15Ala6817PRTArtificial Sequencerecombinant peptide
biosensorVARIANT8, 9Xaa = Any Amino Acid 68Lys Trp Phe Lys Asn Pro
Ile Xaa Xaa Ser His Asn Val Tyr Ile Met1 5 10
15Ala6916PRTArtificial Sequencerecombinant peptide
biosensorVARIANT8, 9Xaa = Any Amino Acid 69Phe Lys Asn Pro Ile Pro
Pro Xaa Xaa Asn Val Tyr Ile Met Ala Asp1 5 10 157016PRTArtificial
Sequencerecombinant peptide biosensorVARIANT8, 9Xaa = Any Amino
Acid 70Lys Trp Phe Lys Asn Pro Ile Xaa Xaa Asn Val Tyr Ile Met Ala
Asp1 5 10 157115PRTArtificial Sequencerecombinant peptide
biosensorVARIANT8Xaa = Any Amino Acid 71Lys Leu Glu Tyr Asn Phe Asn
Xaa Lys Asn Leu Asn Met Asn Phe1 5 10 157216PRTArtificial
Sequencerecombinant peptide biosensorVARIANT8, 9Xaa = Any Amino
Acid 72Lys Leu Glu Tyr Asn Phe Asn Xaa Xaa Lys Asn Leu Asn Met Asn
Phe1 5 10 157316PRTArtificial Sequencerecombinant peptide
biosensorVARIANT8Xaa = Any Amino Acid 73Lys Leu Glu Tyr Asn Phe Asn
Xaa Pro Lys Asn Leu Asn Met Asn Phe1 5 10 157416PRTArtificial
Sequencerecombinant peptide biosensorVARIANT9Xaa = Any Amino Acid
74Lys Leu Glu Tyr Asn Phe Asn Pro Xaa Lys Asn Leu Asn Met Asn Phe1
5 10 157514PRTArtificial Sequencerecombinant peptide
biosensorVARIANT8, 9Xaa = Any Amino Acid 75Gly His Lys Leu Glu Tyr
Asn Xaa Xaa Leu Asn Met Asn Phe1 5 107614PRTArtificial
Sequencerecombinant peptide biosensorVARIANT8, 9Xaa = Any Amino
Acid 76Lys Leu Glu Tyr Asn Phe Asn Xaa Xaa Leu Asn Met Asn Phe1 5
1077567PRTArtificial Sequencerecombinant peptide biosensor 77Met
His His His His His His Gly Ser Glu Glu Gln Glu Lys Ala Leu1 5 10
15Asn Phe Gly Ile Ile Ser Thr Glu Ser Gln Gln Asn Leu Lys Pro Gln
20 25 30Trp Thr Pro Phe Leu Gln Asp Met Glu Lys Lys Leu Gly Val Lys
Val 35 40 45Asn Ala Phe Phe Ala Pro Asp Tyr Ala Gly Ile Ile Gln Gly
Met Arg 50 55 60Phe Asn Lys Val Asp Ile Ala Trp Tyr Gly Asn Leu Ser
Ala Met Glu65 70 75 80Ala Val Asp Arg Ala Asn Gly Gln Val Phe Ala
Gln Thr Val Ala Ala 85 90 95Asp Gly Ser Ser His Asn Val Tyr Ile Met
Ala Asp Lys Gln Arg Asn 100 105 110Gly Ile Lys Ala Asn Phe Lys Ile
Arg His Asn Ile Glu Asp Gly Gly 115 120 125Val Gln Leu Ala Tyr His
Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly 130 135 140Pro Val Leu Leu
Pro Asp Asn His Tyr Leu Ser Thr Gln Ser Lys Leu145 150 155 160Ser
Lys Asp Pro Asn Glu Lys Arg Asp His Met Val Leu Leu Glu Phe 165 170
175Val Thr Ala Ala Gly Ile Thr Leu Gly Met Asp Glu Leu Tyr Lys Gly
180 185 190Gly Thr Gly Gly Ser Met Val Ser Lys Gly Glu Glu Leu Phe
Thr Gly 195 200 205Val Val Pro Ile Leu Val Glu Leu Asp Gly Asp Val
Asn Gly His Lys 210 215 220Phe Ser Val Ser Gly Glu Gly Glu Gly Asp
Ala Thr Tyr Gly Lys Leu225 230 235 240Thr Leu Lys Phe Ile Cys Thr
Thr Gly Lys Leu Pro Val Pro Trp Pro 245 250 255Thr Leu Val Thr Thr
Leu Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr 260 265 270Pro Asp His
Met Lys Gln His Asp Phe Phe Lys Ser Ala Met Pro Glu 275 280 285Gly
Tyr Ile Gln Glu Arg Thr Ile Phe Phe Lys Asp Asp Gly Asn Tyr 290 295
300Lys Thr Arg Ala Glu Val Lys Phe Glu Gly Asp Thr Leu Val Asn
Arg305 310 315 320Ile Glu Leu Lys Gly Ile Asp Phe Lys Glu Asp Gly
Asn Ile Leu Gly 325 330 335His Lys Leu Glu Tyr Asn Phe Asn Pro Gly
Tyr Trp Ser Val Leu Ile 340 345 350Val Asn Lys Asp Ser Pro Ile Asn
Asn Leu Asn Asp Leu Leu Ala Lys 355 360 365Arg Lys Asp Leu Thr Phe
Gly Asn Gly Asp Pro Asn Ser Thr Ser Gly 370 375 380Phe Leu Val Pro
Gly Tyr Tyr Val Phe Ala Lys Asn Ile Asn Ile Ser385 390 395 400Ala
Ser Asp Phe Lys Arg Thr Val Asn Ala Gly His Glu Thr Asn Ala 405 410
415Leu Ala Val Ala Asn Lys Gln Val Asp Val Ala Thr Asn Asn Thr Glu
420 425 430Asn Leu Asp Lys Leu Lys Thr Ser Ala Pro Glu Lys Leu Lys
Glu Leu 435 440 445Lys Val Ile Trp Lys Ser Pro Leu Ile Pro Gly Asp
Pro Ile Val Trp 450 455 460Arg Lys Asn Leu Ser Glu Thr Thr Lys Asp
Lys Ile Tyr Asp Phe Phe465 470 475 480Met Asn Tyr Gly Lys Thr Pro
Glu Glu Lys Ala Val Leu Glu Arg Leu 485 490 495Gly Trp Ala Pro Phe
Arg Ala Ser Ser Asp Leu Gln Leu Val Pro Ile 500 505 510Arg Gln Leu
Ala Leu Phe Lys Glu Met Gln Ser Val Lys Asp Asn Lys 515 520 525Gly
Leu Asn Glu Gln Asp Lys Leu Ala Lys Thr Thr Ala Ile Gln Ala 530 535
540Gln Leu Asp Asp Leu Asp Arg Leu Asn Asn Ala Leu Ser Ala Met
Ser545 550 555 560Ser Val Ser Lys Ala Val Gln 56578566PRTArtificial
Sequencerecombinant peptide biosensor 78Met His His His His His His
Gly Ser Glu Glu Gln Glu Lys Ala Leu1 5 10 15Asn Phe Gly Ile Ile Ser
Thr Glu Ser Gln Gln Asn Leu Lys Pro Gln 20 25 30Trp Thr Pro Phe Leu
Gln Asp Met Glu Lys Lys Leu Gly Val Lys Val 35 40 45Asn Ala Phe Phe
Ala Pro Asp Tyr Ala Gly Ile Ile Gln Gly Met Arg 50 55 60Phe Asn Lys
Val Asp Ile Ala Trp Tyr Gly Asn Leu Ser Ala Met Glu65 70 75 80Ala
Val Asp Arg Ala Asn Gly Gln Val Phe Ala Gln Thr Val Ala Ala 85 90
95Asp Ala Asp Asn Val Tyr Ile Met Ala Asp Lys Gln Arg Asn Gly Ile
100 105 110Lys Ala Asn Phe Lys Ile Arg His Asn Ile Glu Asp Gly Gly
Val Gln 115 120 125Leu Ala Tyr His Tyr Gln Gln Asn Thr Pro Ile Gly
Asp Gly Pro Val 130 135 140Leu Leu Pro Asp Asn His Tyr Leu Ser Thr
Gln Ser Lys Leu Ser Lys145 150 155 160Ser Lys Asp Pro Asn Glu Lys
Arg Asp His Met Val Leu Leu Glu Phe 165 170 175Val Thr Ala Ala Gly
Ile Thr Leu Gly Met Asp Glu Leu Tyr Lys Gly 180 185 190Gly Thr Gly
Gly Ser Met Val Ser Lys Gly Glu Glu Leu Phe Thr Gly 195 200 205Val
Val Pro Ile Leu Val Glu Leu Asp Gly Asp Val Asn Gly His Lys 210 215
220Phe Ser Val Ser Gly Glu Gly Glu Gly Asp Ala Thr Tyr Gly Lys
Leu225 230 235 240Thr Leu Lys Phe Ile Cys Thr Thr Gly Lys Leu Pro
Val Pro Trp Pro 245 250 255Thr Leu Val Thr Thr Leu Thr Tyr Gly Val
Gln Cys Phe Ser Arg Tyr 260 265 270Pro Asp His Met Lys Gln His Asp
Phe Phe Lys Ser Ala Met Pro Glu 275 280 285Gly Tyr Ile Gln Glu Arg
Thr Ile Phe Phe Lys Asp Asp Gly Asn Tyr 290 295 300Lys Thr Arg Ala
Glu Val Lys Phe Glu Gly Asp Thr Leu Val Asn Arg305 310 315 320Ile
Glu Leu Lys Gly Ile Asp Phe Lys Glu Asp Gly Asn Ile Leu Gly 325 330
335His Lys Leu Glu Tyr Asn Phe Asn Pro Gly Tyr Trp Ser Val Leu Ile
340 345 350Val Asn Lys Asp Ser Pro Ile Asn Asn Leu Asn Asp Leu Leu
Ala Lys 355 360 365Arg Lys Asp Leu Thr Phe Gly Asn Gly Asp Pro Asn
Ser Thr Ser Gly 370 375 380Phe Leu Val Pro Gly Tyr Tyr Val Phe Ala
Lys Asn Asn Ile Ser Ala385 390 395 400Gly Lys Thr Pro Glu Glu Lys
Ala Val Leu Glu Arg Leu Gly Trp Ala 405 410 415Pro Phe Arg Ala Ser
Ser Asp Leu Gln Leu Val Pro Ile Arg Gln Leu 420 425 430Ala Leu Phe
Lys Glu Met Gln Ser Val Lys Asp Asn Lys Gly Leu Asn 435 440 445Glu
Gln Asp Lys Leu Ala Lys Thr Thr Ala Ile Gln Ala Gln Leu Asp 450 455
460Asp Leu Asp Arg Arg Asn Asn Ala Arg Ser Ala Met Ser Ser Val
Ser465 470 475 480Asn Tyr Gly Lys Thr Pro Glu Glu Lys Ala Val Leu
Glu Arg Leu Gly 485 490 495Trp Ala Pro Phe Arg Ala Ser Ser Asp Leu
Gln Leu Val Pro Ile Arg 500 505 510Gln Leu Ala Leu Phe Lys Glu Met
Gln Ser Val Lys Asp Asn Lys Gly 515 520 525Leu Asn Glu Gln Asp Lys
Leu Ala Lys Thr Thr Ala Ile Gln Ala Gln 530 535 540Leu Asp Asp Leu
Asp Arg Leu Asn Asn Ala Leu Ser Ala Met Ser Ser545 550 555 560Val
Ser Lys Ala Val Gln 5657916PRTArtificial Sequencesynthetic
oligopeptide 79Gln Thr Val Ala Ala Asp Gly Ser Ser His Asn Val Tyr
Ile Met Ala1 5 10 158016PRTArtificial Sequencesynthetic
oligopeptideVARIANT7, 8Xaa = Any Amino Acid 80Gln Thr Val Ala Ala
Asp Xaa Xaa Ser His Asn Val Tyr Ile Met Ala1 5 10
158116PRTArtificial Sequencesynthetic oligopeptideVARIANT7Xaa = Any
Amino Acid 81Gln Thr Val Ala Ala Asp Xaa Pro Ser His Asn Val Tyr
Ile Met Ala1 5 10 158216PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 82Gln Thr Val Ala Ala Asp
Pro Xaa Ser His Asn Val Tyr Ile Met Ala1 5 10 158314PRTArtificial
Sequencesynthetic oligopeptideVARIANT7, 8Xaa = Any Amino Acid 83Gln
Thr Val Ala Ala Asp Xaa Xaa Asn Val Tyr Ile Met Ala1 5
108416PRTArtificial Sequencesynthetic oligopeptideVARIANT7, 8Xaa =
Any Amino Acid 84Gln Thr Val Ala Ala Asp Xaa Xaa Ser His Asn Val
Tyr Ile Met Ala1 5 10 158516PRTArtificial Sequencesynthetic
oligopeptideVARIANT7, 8Xaa = Any Amino Acid 85Val Phe Gln Thr Val
Ala Xaa Xaa Ser His Asn Val Tyr Ile Met Ala1 5 10
158616PRTArtificial Sequencesynthetic oligopeptide 86His Lys Leu
Glu Tyr Asn Phe Asn Pro Gly Tyr Trp Ser Val Leu Ile1 5 10
158718PRTArtificial Sequencesynthetic oligopeptideVARIANT9, 10Xaa =
Any Amino Acid 87His Lys Leu Glu Tyr Asn Phe Asn Xaa Xaa Pro Gly
Tyr Trp Ser Val1 5 10 15Leu Ile8816PRTArtificial Sequencesynthetic
oligopeptideVARIANT7, 8Xaa = Any Amino Acid 88His Lys Leu Glu Tyr
Asn Xaa Xaa Pro Gly Tyr Trp Ser Val Leu Ile1 5 10
158916PRTArtificial Sequencesynthetic oligopeptideVARIANT9, 10Xaa =
Any Amino Acid 89His Lys Leu Glu Tyr Asn Phe Asn Xaa Xaa Tyr Trp
Ser Val Leu Ile1 5 10 159016PRTArtificial Sequencesynthetic
oligopeptideVARIANT10Xaa = Any Amino Acid 90His Lys Leu Glu Tyr Asn
Phe Asn Pro Xaa Tyr Trp Ser Val Leu Ile1 5 10 1591678PRTArtificial
Sequencerecombinant peptide biosensor 91Met Arg Gly Ser His His His
His His His Gly Met Ala Ser Met
Thr1 5 10 15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp
Lys Asp 20 25 30Arg Trp Gly Ser Lys Leu Glu Ile Phe Ser Trp Trp Ala
Gly Asp Glu 35 40 45Gly Pro Ala Leu Glu Ala Leu Ile Arg Leu Tyr Lys
Gln Lys Tyr Pro 50 55 60Gly Val Glu Val Ile Asn Ala Thr Val Thr Gly
Gly Ala Gly Val Asn65 70 75 80Ala Arg Ala Val Leu Lys Thr Arg Met
Leu Gly Gly Asp Pro Pro Asp 85 90 95Thr Phe Gln Val His Ala Gly Met
Glu Leu Ile Gly Thr Trp Val Val 100 105 110Ala Asn Arg Met Glu Asp
Leu Ser Ala Leu Phe Arg Gln Glu Gly Trp 115 120 125Leu Gln Ala Phe
Pro Lys Gly Leu Ile Asp Leu Ile Ser Tyr Lys Gly 130 135 140Gly Ile
Trp Ser Val Pro Val Asn Ile His Arg Ser Asn Val Met Trp145 150 155
160Tyr Leu Pro Ala Lys Leu Lys Glu Trp Gly Val Asn Pro Pro Arg Thr
165 170 175Trp Asp Glu Phe Leu Ala Thr Cys Gln Thr Leu Lys Gln Lys
Gly Leu 180 185 190Glu Ala Pro Leu Ala Leu Gly Glu Asn Trp Thr Gln
Gln His Leu Trp 195 200 205Glu Ser Val Ala Leu Ala Val Leu Gly Pro
Asp Asp Trp Asn Asn Leu 210 215 220Trp Asn Gly Lys Leu Lys Phe Thr
Asp Pro Lys Ala Val Arg Ala Trp225 230 235 240Glu Val Phe Gly Arg
Val Leu Asp Cys Ala Asn Lys Asp Ala Ala Gly 245 250 255Leu Ser Trp
Gln Gln Ala Val Asp Arg Val Val Gln Gly Lys Ala Ala 260 265 270Phe
Asn Val Met Gly Asp Trp Ala Ala Gly Tyr Met Thr Thr Thr Leu 275 280
285Lys Leu Lys Pro Gly Thr Asp Phe Ala Trp Ala Pro Ser Pro Gly Thr
290 295 300Gln Gly Val Phe Met Met Leu Ser Asp Ser Phe Gly Leu Pro
Lys Gly305 310 315 320Ala Lys Asn Arg Gln Asn Ala Ile Asn Trp Leu
Arg Leu Val Gly Ser 325 330 335Lys Glu Gly Gln Asp Thr Phe Asn Pro
Leu Lys Gly Ser Ile Ala Ala 340 345 350Arg Leu Asp Ser Asp Pro Ser
Lys Tyr Gly Gly Ser His Asn Val Tyr 355 360 365Ile Met Ala Asp Lys
Gln Arg Asn Gly Ile Lys Ala Asn Phe Lys Ile 370 375 380Arg His Asn
Ile Glu Asp Gly Gly Val Gln Leu Ala Tyr His Tyr Gln385 390 395
400Gln Asn Thr Pro Ile Gly Asp Gly Pro Val Leu Leu Pro Asp Asn His
405 410 415Tyr Leu Ser Thr Gln Ser Lys Leu Ser Lys Asp Pro Asn Glu
Lys Arg 420 425 430Asp His Met Val Leu Leu Glu Phe Val Thr Ala Ala
Gly Ile Thr Leu 435 440 445Gly Met Asp Glu Leu Tyr Lys Gly Gly Thr
Gly Gly Ser Met Val Ser 450 455 460Lys Gly Glu Glu Leu Phe Thr Gly
Val Val Pro Ile Leu Val Glu Leu465 470 475 480Asp Gly Asp Val Asn
Gly His Lys Phe Ser Val Ser Gly Glu Gly Glu 485 490 495Gly Asp Ala
Thr Tyr Gly Lys Leu Thr Leu Lys Phe Ile Cys Thr Thr 500 505 510Gly
Lys Leu Pro Val Pro Trp Pro Thr Leu Val Thr Thr Leu Thr Tyr 515 520
525Gly Val Gln Cys Phe Ser Arg Tyr Pro Asp His Met Lys Gln His Asp
530 535 540Phe Phe Lys Ser Ala Met Pro Glu Gly Tyr Ile Gln Glu Arg
Thr Ile545 550 555 560Phe Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg
Ala Glu Val Lys Phe 565 570 575Glu Gly Asp Thr Leu Val Asn Arg Ile
Glu Leu Lys Gly Ile Asp Phe 580 585 590Lys Glu Asp Gly Asn Ile Leu
Gly His Lys Leu Glu Tyr Asn Phe Asn 595 600 605Asn Pro Asn Ala Tyr
Gly Gln Ser Ala Met Arg Asp Trp Arg Ser Asn 610 615 620Arg Ile Val
Gly Ser Leu Val His Gly Ala Val Ala Pro Glu Ser Phe625 630 635
640Met Ser Gln Phe Gly Thr Val Met Glu Ile Phe Leu Gln Thr Arg Asn
645 650 655Pro Gln Ala Ala Ala Asn Ala Ala Gln Ala Ile Ala Asp Gln
Val Gly 660 665 670Leu Gly Arg Leu Gly Gln 67592678PRTArtificial
Sequencerecombinant peptide biosensor 92Met Arg Gly Ser His His His
His His His Gly Met Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met Gly
Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys
Leu Glu Ile Phe Ser Trp Trp Ala Gly Asp Glu 35 40 45Gly Pro Ala Leu
Glu Ala Leu Ile Arg Leu Tyr Lys Gln Lys Tyr Pro 50 55 60Gly Val Glu
Val Ile Asn Ala Thr Val Thr Gly Gly Ala Gly Val Asn65 70 75 80Ala
Arg Ala Val Leu Lys Thr Arg Met Leu Gly Gly Asp Pro Pro Asp 85 90
95Thr Phe Gln Val His Ala Gly Met Glu Leu Ile Gly Thr Trp Val Val
100 105 110Ala Asn Arg Met Glu Asp Leu Ser Ala Leu Phe Arg Gln Glu
Gly Trp 115 120 125Leu Gln Ala Phe Pro Lys Gly Leu Ile Asp Leu Ile
Ser Tyr Lys Gly 130 135 140Gly Ile Trp Ser Val Pro Val Asn Ile His
Arg Ser Asn Val Met Trp145 150 155 160Tyr Leu Pro Ala Lys Leu Lys
Glu Trp Gly Val Asn Pro Pro Arg Thr 165 170 175Trp Asp Glu Phe Leu
Ala Thr Cys Gln Thr Leu Lys Gln Lys Gly Leu 180 185 190Glu Ala Pro
Leu Ala Leu Gly Glu Asn Trp Thr Gln Gln His Leu Trp 195 200 205Glu
Ser Val Ala Leu Ala Val Leu Gly Pro Asp Asp Trp Asn Asn Leu 210 215
220Trp Asn Gly Lys Leu Lys Phe Thr Asp Pro Lys Ala Val Arg Ala
Trp225 230 235 240Glu Val Phe Gly Arg Val Leu Asp Cys Ala Asn Lys
Asp Ala Ala Gly 245 250 255Leu Ser Trp Gln Gln Ala Val Asp Arg Val
Val Gln Gly Lys Ala Ala 260 265 270Phe Asn Val Met Gly Asp Trp Ala
Ala Gly Tyr Met Thr Thr Thr Leu 275 280 285Lys Leu Lys Pro Gly Thr
Asp Phe Ala Trp Ala Pro Ser Pro Gly Thr 290 295 300Gln Gly Val Phe
Met Met Leu Ser Asp Ser Phe Gly Leu Pro Lys Gly305 310 315 320Ala
Lys Asn Arg Gln Asn Ala Ile Asn Trp Leu Arg Leu Val Gly Ser 325 330
335Lys Glu Gly Gln Asp Thr Phe Asn Pro Leu Lys Gly Ser Ile Ala Ala
340 345 350Arg Leu Asp Ser Asp Pro Ser Lys Tyr Pro Ala Ser His Asn
Val Tyr 355 360 365Ile Met Ala Asp Lys Gln Arg Asn Gly Ile Lys Ala
Asn Phe Lys Ile 370 375 380Arg His Asn Ile Glu Asp Gly Gly Val Gln
Leu Ala Tyr His Tyr Gln385 390 395 400Gln Asn Thr Pro Ile Gly Asp
Gly Pro Val Leu Leu Pro Asp Asn His 405 410 415Tyr Leu Ser Thr Gln
Ser Lys Leu Ser Lys Asp Pro Asn Glu Lys Arg 420 425 430Asp His Met
Val Leu Leu Glu Phe Val Thr Ala Ala Gly Ile Thr Leu 435 440 445Gly
Met Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser Met Val Ser 450 455
460Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu Val Glu
Leu465 470 475 480Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser
Gly Glu Gly Glu 485 490 495Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu
Lys Phe Ile Cys Thr Thr 500 505 510Gly Lys Leu Pro Val Pro Trp Pro
Thr Leu Val Thr Thr Leu Thr Tyr 515 520 525Gly Val Gln Cys Phe Ser
Arg Tyr Pro Asp His Met Lys Gln His Asp 530 535 540Phe Phe Lys Ser
Ala Met Pro Glu Gly Tyr Ile Gln Glu Arg Thr Ile545 550 555 560Phe
Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu Val Lys Phe 565 570
575Glu Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys Gly Ile Asp Phe
580 585 590Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu Tyr Asn
Phe Asn 595 600 605Asn Pro Asn Ala Tyr Gly Gln Ser Ala Met Arg Asp
Trp Arg Ser Asn 610 615 620Arg Ile Val Gly Ser Leu Val His Gly Ala
Val Ala Pro Glu Ser Phe625 630 635 640Met Ser Gln Phe Gly Thr Val
Met Glu Ile Phe Leu Gln Thr Arg Asn 645 650 655Pro Gln Ala Ala Ala
Asn Ala Ala Gln Ala Ile Ala Asp Gln Val Gly 660 665 670Leu Gly Arg
Leu Gly Gln 67593678PRTArtificial Sequencerecombinant peptide
biosensor 93Met Arg Gly Ser His His His His His His Gly Met Ala Ser
Met Thr1 5 10 15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp
Asp Lys Asp 20 25 30Arg Trp Gly Ser Lys Leu Glu Ile Phe Ser Trp Trp
Ala Gly Asp Glu 35 40 45Gly Pro Ala Leu Glu Ala Leu Ile Arg Leu Tyr
Lys Gln Lys Tyr Pro 50 55 60Gly Val Glu Val Ile Asn Ala Thr Val Thr
Gly Gly Ala Gly Val Asn65 70 75 80Ala Arg Ala Val Leu Lys Thr Arg
Met Leu Gly Gly Asp Pro Pro Asp 85 90 95Thr Phe Gln Val His Ala Gly
Met Glu Leu Ile Gly Thr Trp Val Val 100 105 110Ala Asn Arg Met Glu
Asp Leu Ser Ala Leu Phe Arg Gln Glu Gly Trp 115 120 125Leu Gln Ala
Phe Pro Lys Gly Leu Ile Asp Leu Ile Ser Tyr Lys Gly 130 135 140Gly
Ile Trp Ser Val Pro Val Asn Ile His Arg Ser Asn Val Met Trp145 150
155 160Tyr Leu Pro Ala Lys Leu Lys Glu Trp Gly Val Asn Pro Pro Arg
Thr 165 170 175Trp Asp Glu Phe Leu Ala Thr Cys Gln Thr Leu Lys Gln
Lys Gly Leu 180 185 190Glu Ala Pro Leu Ala Leu Gly Glu Asn Trp Thr
Gln Gln His Leu Trp 195 200 205Glu Ser Val Ala Leu Ala Val Leu Gly
Pro Asp Asp Trp Asn Asn Leu 210 215 220Trp Asn Gly Lys Leu Lys Phe
Thr Asp Pro Lys Ala Val Arg Ala Trp225 230 235 240Ala Arg Ala Val
Leu Lys Thr Arg Met Leu Gly Gly Asp Pro Pro Asp 245 250 255Thr Phe
Gln Val Ala Ala Gly Met Glu Leu Ile Gly Thr Trp Val Val 260 265
270Ala Asn Arg Met Glu Asp Leu Ser Ala Leu Phe Arg Gln Glu Gly Trp
275 280 285Leu Gln Ala Phe Pro Lys Gly Leu Ile Asp Leu Ile Ser Tyr
Lys Gly 290 295 300Gly Ile Trp Ser Val Pro Val Asn Ile His Arg Ser
Asn Val Met Trp305 310 315 320Ala Lys Asn Arg Gln Asn Ala Ile Asn
Trp Leu Arg Leu Val Gly Ser 325 330 335Lys Glu Gly Gln Asp Thr Phe
Asn Pro Leu Lys Gly Ser Ile Ala Ala 340 345 350Arg Leu Asp Ser Asp
Pro Ser Lys Tyr Gly Gly Ser His Asn Val Tyr 355 360 365Ile Met Ala
Asp Lys Gln Arg Asn Gly Ile Lys Ala Asn Phe Lys Ile 370 375 380Arg
His Asn Ile Glu Asp Gly Gly Val Gln Leu Ala Tyr His Tyr Gln385 390
395 400Gln Asn Thr Pro Ile Gly Asp Gly Pro Val Leu Leu Pro Asp Asn
His 405 410 415Tyr Leu Ser Thr Gln Ser Lys Leu Ser Lys Asp Pro Asn
Glu Lys Arg 420 425 430Asp His Met Val Leu Leu Glu Phe Val Thr Ala
Ala Gly Ile Thr Leu 435 440 445Gly Met Asp Glu Leu Tyr Lys Gly Gly
Thr Gly Gly Ser Met Val Ser 450 455 460Lys Gly Glu Glu Leu Phe Thr
Gly Val Val Pro Ile Leu Val Glu Leu465 470 475 480Asp Gly Asp Val
Asn Gly His Lys Phe Ser Val Ser Gly Glu Gly Glu 485 490 495Gly Asp
Ala Thr Tyr Gly Lys Leu Thr Leu Lys Phe Ile Cys Thr Thr 500 505
510Gly Lys Leu Pro Val Pro Trp Pro Thr Leu Val Thr Thr Leu Thr Tyr
515 520 525Gly Val Gln Cys Phe Ser Arg Tyr Pro Asp His Met Lys Gln
His Asp 530 535 540Phe Phe Lys Ser Ala Met Pro Glu Gly Tyr Ile Gln
Glu Arg Thr Ile545 550 555 560Phe Phe Lys Asp Asp Gly Asn Tyr Lys
Thr Arg Ala Glu Val Lys Phe 565 570 575Glu Gly Asp Thr Leu Val Asn
Arg Ile Glu Leu Lys Gly Ile Asp Phe 580 585 590Lys Glu Asp Gly Asn
Ile Leu Gly His Lys Leu Glu Tyr Asn Phe Asn 595 600 605Asn Pro Asn
Ala Tyr Gly Gln Ser Ala Met Arg Asp Trp Arg Ser Asn 610 615 620Arg
Ile Val Gly Ser Leu Val His Gly Ala Val Ala Pro Glu Ser Phe625 630
635 640Met Ser Gln Phe Gly Thr Val Met Glu Ile Phe Leu Gln Thr Arg
Asn 645 650 655Pro Gln Ala Ala Ala Asn Ala Ala Gln Ala Ile Ala Asp
Gln Val Gly 660 665 670Leu Gly Arg Leu Gly Gln
67594678PRTArtificial Sequencerecombinant peptide biosensor 94Met
Arg Gly Ser His His His His His His Gly Met Ala Ser Met Thr1 5 10
15Gly Gly Gln Gln Met Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp
20 25 30Arg Trp Gly Ser Lys Leu Glu Ile Phe Ser Trp Trp Ala Gly Asp
Glu 35 40 45Gly Pro Ala Leu Glu Ala Leu Ile Arg Leu Tyr Lys Gln Lys
Tyr Pro 50 55 60Gly Val Glu Val Ile Asn Ala Thr Val Thr Gly Gly Ala
Gly Val Asn65 70 75 80Ala Arg Ala Val Leu Lys Thr Arg Met Leu Gly
Gly Asp Pro Pro Asp 85 90 95Thr Phe Gln Val His Ala Gly Met Glu Leu
Ile Gly Thr Trp Val Val 100 105 110Ala Asn Arg Met Glu Asp Leu Ser
Ala Leu Phe Arg Gln Glu Gly Trp 115 120 125Leu Gln Ala Phe Pro Lys
Gly Leu Ile Asp Leu Ile Ser Tyr Lys Gly 130 135 140Gly Ile Trp Ser
Val Pro Val Asn Ile His Arg Ser Asn Val Met Trp145 150 155 160Tyr
Leu Pro Ala Lys Leu Lys Glu Trp Gly Val Asn Pro Pro Arg Thr 165 170
175Trp Asp Glu Phe Leu Ala Thr Cys Gln Thr Leu Lys Gln Lys Gly Leu
180 185 190Glu Ala Pro Leu Ala Leu Gly Glu Asn Trp Thr Gln Gln His
Leu Trp 195 200 205Glu Ser Val Ala Leu Ala Val Leu Gly Pro Asp Asp
Trp Asn Asn Leu 210 215 220Trp Asn Gly Lys Leu Lys Phe Thr Asp Pro
Lys Ala Val Arg Ala Trp225 230 235 240Glu Val Phe Gly Arg Val Leu
Asp Cys Ala Asn Lys Asp Ala Ala Gly 245 250 255Leu Ser Trp Gln Gln
Ala Val Asp Arg Val Val Gln Gly Lys Ala Ala 260 265 270Phe Asn Val
Met Gly Asp Trp Ala Ala Gly Tyr Met Thr Thr Thr Leu 275 280 285Lys
Leu Lys Pro Gly Thr Asp Phe Ala Trp Ala Pro Ser Pro Gly Thr 290 295
300Gln Gly Val Phe Met Met Leu Ser Asp Ser Phe Gly Leu Pro Lys
Gly305 310 315 320Ala Lys Asn Arg Gln Asn Ala Ile Asn Trp Leu Arg
Leu Val Gly Ser 325 330 335Lys Glu Gly Gln Asp Thr Phe Asn Pro Leu
Lys Gly Ser Ile Ala Ala 340 345 350Arg Leu Asp Ser Asp Pro Ser Lys
Tyr Gly Gly Ser His Asn Val Tyr 355 360 365Ile Met Ala Asp Lys Gln
Arg Asn Gly Ile Lys Ala Asn Phe Lys Ile 370 375 380Arg His Asn Ile
Glu Asp Gly Gly Val Gln Leu Ala Tyr His Tyr Gln385 390 395 400Gln
Asn Thr Pro Ile Gly Asp Gly Pro Val Leu Leu Pro Asp Asn His 405 410
415Tyr Leu Ser Thr Gln Ser Lys Leu Ser Lys Asp Pro Asn Glu Lys Arg
420
425 430Asp His Met Val Leu Leu Glu Phe Val Thr Ala Ala Gly Ile Thr
Leu 435 440 445Gly Met Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser
Met Val Ser 450 455 460Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro
Ile Leu Val Glu Leu465 470 475 480Asp Gly Asp Val Asn Gly His Lys
Phe Ser Val Ser Gly Glu Gly Glu 485 490 495Gly Asp Ala Thr Tyr Gly
Lys Leu Thr Leu Lys Phe Ile Cys Thr Thr 500 505 510Gly Lys Leu Pro
Val Pro Trp Pro Thr Leu Val Thr Thr Leu Thr Tyr 515 520 525Gly Val
Gln Cys Phe Ser Arg Tyr Pro Asp His Met Lys Gln His Asp 530 535
540Phe Phe Lys Ser Ala Met Pro Glu Gly Tyr Ile Gln Glu Arg Thr
Ile545 550 555 560Phe Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala
Glu Val Lys Phe 565 570 575Glu Gly Asp Thr Leu Val Asn Arg Ile Glu
Leu Lys Gly Ile Asp Phe 580 585 590Lys Glu Asp Gly Asn Ile Leu Gly
His Lys Leu Glu Tyr Asn Phe Asn 595 600 605Asn Pro Asn Ala Tyr Gly
Gln Ser Ala Met Arg Asp Trp Arg Ser Asn 610 615 620Arg Ile Val Gly
Ser Leu Val Ala Gly Ala Val Ala Pro Glu Ser Phe625 630 635 640Met
Ser Gln Phe Gly Thr Val Met Glu Ile Phe Leu Gln Thr Arg Asn 645 650
655Pro Gln Ala Ala Ala Asn Ala Ala Gln Ala Ile Ala Asp Gln Val Gly
660 665 670Leu Gly Arg Leu Gly Gln 6759516PRTArtificial
Sequencesynthetic oligopeptideVARIANT8, 9Xaa = Any Amino Acid 95Asp
Ser Asp Pro Ser Lys Tyr Xaa Xaa Ser His Asn Val Tyr Ile Met1 5 10
159616PRTArtificial Sequencesynthetic oligopeptideVARIANT9Xaa = Any
Amino Acid 96Asp Ser Asp Pro Ser Lys Tyr Pro Xaa Ser His Asn Val
Tyr Ile Met1 5 10 159716PRTArtificial Sequencesynthetic
oligopeptideVARIANT8Xaa = Any Amino Acid 97Asp Ser Asp Pro Ser Lys
Tyr Xaa Pro Ser His Asn Val Tyr Ile Met1 5 10 159816PRTArtificial
Sequencesynthetic oligopeptideVARIANT8, 9Xaa = Any Amino Acid 98Arg
Leu Asp Ser Asp Pro Ser Xaa Xaa Ser His Asn Val Tyr Ile Met1 5 10
159914PRTArtificial Sequencesynthetic oligopeptideVARIANT8, 9Xaa =
Any Amino Acid 99Asp Ser Asp Pro Ser Lys Tyr Xaa Xaa Asn Val Tyr
Ile Met1 5 1010016PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 100Lys Leu Glu Tyr Asn
Phe Asn Xaa Xaa Asn Ala Tyr Gly Gln Ser Ala1 5 10
1510116PRTArtificial Sequencesynthetic oligopeptideVARIANT7, 8Xaa =
Any Amino Acid 101Lys Leu Glu Tyr Asn Phe Xaa Xaa Pro Asn Ala Tyr
Gly Gln Ser Ala1 5 10 1510216PRTArtificial Sequencesynthetic
oligopeptideVARIANT8, 9Xaa = Any Amino Acid 102Gly His Lys Leu Glu
Tyr Asn Xaa Xaa Asn Ala Tyr Gly Gln Ser Ala1 5 10
1510316PRTArtificial Sequencesynthetic oligopeptideVARIANT8Xaa =
Any Amino Acid 103Lys Leu Glu Tyr Asn Phe Asn Xaa Pro Asn Ala Tyr
Gly Gln Ser Ala1 5 10 1510416PRTArtificial Sequencesynthetic
oligopeptideVARIANT9Xaa = Any Amino Acid 104Lys Leu Glu Tyr Asn Phe
Asn Pro Xaa Asn Ala Tyr Gly Gln Ser Ala1 5 10
15105396PRTEscherichia coli 105Met Lys Ile Lys Thr Gly Ala Arg Ile
Leu Ala Leu Ser Ala Leu Thr1 5 10 15Thr Met Met Phe Ser Ala Ser Ala
Leu Ala Lys Ile Glu Glu Gly Lys 20 25 30Leu Val Ile Trp Ile Asn Gly
Asp Lys Gly Tyr Asn Gly Leu Ala Glu 35 40 45Val Gly Lys Lys Phe Glu
Lys Asp Thr Gly Ile Lys Val Thr Val Glu 50 55 60His Pro Asp Lys Leu
Glu Glu Lys Phe Pro Gln Val Ala Ala Thr Gly65 70 75 80Asp Gly Pro
Asp Ile Ile Phe Trp Ala His Asp Arg Phe Gly Gly Tyr 85 90 95Ala Gln
Ser Gly Leu Leu Ala Glu Ile Thr Pro Asp Lys Ala Phe Gln 100 105
110Asp Lys Leu Tyr Pro Phe Thr Trp Asp Ala Val Arg Tyr Asn Gly Lys
115 120 125Leu Ile Ala Tyr Pro Ile Ala Val Glu Ala Leu Ser Leu Ile
Tyr Asn 130 135 140Lys Asp Leu Leu Pro Asn Pro Pro Lys Thr Trp Glu
Glu Ile Pro Ala145 150 155 160Leu Asp Lys Glu Leu Lys Ala Lys Gly
Lys Ser Ala Leu Met Phe Asn 165 170 175Leu Gln Glu Pro Tyr Phe Thr
Trp Pro Leu Ile Ala Ala Asp Gly Gly 180 185 190Tyr Ala Phe Lys Tyr
Glu Asn Gly Lys Tyr Asp Ile Lys Asp Val Gly 195 200 205Val Asp Asn
Ala Gly Ala Lys Ala Gly Leu Thr Phe Leu Val Asp Leu 210 215 220Ile
Lys Asn Lys His Met Asn Ala Asp Thr Asp Tyr Ser Ile Ala Glu225 230
235 240Ala Ala Phe Asn Lys Gly Glu Thr Ala Met Thr Ile Asn Gly Pro
Trp 245 250 255Ala Trp Ser Asn Ile Asp Thr Ser Lys Val Asn Tyr Gly
Val Thr Val 260 265 270Leu Pro Thr Phe Lys Gly Gln Pro Ser Lys Pro
Phe Val Gly Val Leu 275 280 285Ser Ala Gly Ile Asn Ala Ala Ser Pro
Asn Lys Glu Leu Ala Lys Glu 290 295 300Phe Leu Glu Asn Tyr Leu Leu
Thr Asp Glu Gly Leu Glu Ala Val Asn305 310 315 320Lys Asp Lys Pro
Leu Gly Ala Val Ala Leu Lys Ser Tyr Glu Glu Glu 325 330 335Leu Ala
Lys Asp Pro Arg Ile Ala Ala Thr Met Glu Asn Ala Gln Lys 340 345
350Gly Glu Ile Met Pro Asn Ile Pro Gln Met Ser Ala Phe Trp Tyr Ala
355 360 365Val Arg Thr Ala Val Ile Asn Ala Ala Ser Gly Arg Gln Thr
Val Asp 370 375 380Glu Ala Leu Lys Asp Ala Gln Thr Arg Ile Thr
Lys385 390 395106433PRTPyrococcus furiosus 106Met Arg Arg Ala Thr
Tyr Ala Phe Ala Leu Leu Ala Ile Leu Val Leu1 5 10 15Gly Val Val Ala
Ser Gly Cys Ile Gly Gly Gly Thr Thr Thr Pro Thr 20 25 30Gln Thr Ser
Pro Ala Thr Gln Pro Thr Thr Thr Gln Thr Pro Thr Gln 35 40 45Thr Glu
Thr Gln Ala Val Glu Cys Gly Ser Gly Lys Val Val Ile Trp 50 55 60His
Ala Met Gln Pro Asn Glu Leu Glu Val Phe Gln Ser Leu Ala Glu65 70 75
80Glu Tyr Met Ala Leu Cys Pro Glu Val Glu Ile Val Phe Glu Gln Lys
85 90 95Pro Asn Leu Glu Asp Ala Leu Lys Ala Ala Ile Pro Thr Gly Gln
Gly 100 105 110Pro Asp Leu Phe Ile Trp Ala His Asp Trp Ile Gly Lys
Phe Ala Glu 115 120 125Ala Gly Leu Leu Glu Pro Ile Asp Glu Tyr Val
Thr Glu Asp Leu Leu 130 135 140Asn Glu Phe Ala Pro Met Ala Gln Asp
Ala Met Gln Tyr Lys Gly His145 150 155 160Tyr Tyr Ala Leu Pro Phe
Ala Ala Glu Thr Val Ala Ile Ile Tyr Asn 165 170 175Lys Glu Met Val
Ser Glu Pro Pro Lys Thr Phe Asp Glu Met Lys Ala 180 185 190Ile Met
Glu Lys Tyr Tyr Asp Pro Ala Asn Glu Lys Tyr Gly Ile Ala 195 200
205Trp Pro Ile Asn Ala Tyr Phe Ile Ser Ala Ile Ala Gln Ala Phe Gly
210 215 220Gly Tyr Tyr Phe Asp Asp Lys Thr Glu Gln Pro Gly Leu Asp
Lys Pro225 230 235 240Glu Thr Ile Gly Phe Lys Phe Phe Phe Thr Glu
Ile Trp Pro Tyr Met 245 250 255Ala Pro Thr Gly Asp Tyr Asn Thr Gln
Gln Ser Ile Phe Leu Glu Gly 260 265 270Arg Ala Pro Met Met Val Asn
Gly Pro Trp Ser Ile Asn Asp Val Lys 275 280 285Lys Ala Gly Ile Asn
Phe Gly Val Val Pro Leu Pro Pro Ile Ile Lys 290 295 300Asp Gly Lys
Glu Tyr Trp Pro Arg Pro Tyr Gly Gly Val Lys Leu Ile305 310 315
320Tyr Phe Ala Ala Gly Ile Lys Asn Lys Asp Ala Ala Trp Lys Phe Ala
325 330 335Lys Trp Leu Thr Thr Ser Glu Glu Ser Ile Lys Thr Leu Ala
Leu Glu 340 345 350Leu Gly Tyr Ile Pro Val Leu Thr Lys Val Leu Asp
Asp Pro Glu Ile 355 360 365Lys Asn Asp Pro Val Ile Tyr Gly Phe Gly
Gln Ala Val Gln His Ala 370 375 380Tyr Leu Met Pro Lys Ser Pro Lys
Met Ser Ala Val Trp Gly Gly Val385 390 395 400Asp Gly Ala Ile Asn
Glu Ile Leu Gln Asp Pro Gln Asn Ala Asp Ile 405 410 415Glu Gly Ile
Leu Lys Lys Tyr Gln Gln Glu Ile Leu Asn Asn Met Gln 420 425
430Gly107338PRTEscherichia coli 107Met Asn Ala Lys Ile Ile Ala Ser
Leu Ala Phe Thr Ser Met Phe Ser1 5 10 15Leu Ser Thr Leu Leu Asn Pro
Ala Tyr Ala Glu Glu Gln Glu Lys Ala 20 25 30Leu Asn Phe Gly Ile Ile
Ser Thr Glu Ser Gln Gln Asn Leu Lys Pro 35 40 45Gln Trp Thr Pro Phe
Leu Gln Asp Met Glu Lys Lys Leu Gly Val Lys 50 55 60Val Asn Ala Phe
Phe Ala Pro Asp Tyr Ala Gly Ile Ile Gln Gly Met65 70 75 80Arg Phe
Asn Lys Val Asp Ile Ala Trp Tyr Gly Asn Leu Ser Ala Met 85 90 95Glu
Ala Val Asp Arg Ala Asn Gly Gln Val Phe Ala Gln Thr Val Ala 100 105
110Ala Asp Gly Ser Pro Gly Tyr Trp Ser Val Leu Ile Val Asn Lys Asp
115 120 125Ser Pro Ile Asn Asn Leu Asn Asp Leu Leu Ala Lys Arg Lys
Asp Leu 130 135 140Thr Phe Gly Asn Gly Asp Pro Asn Ser Thr Ser Gly
Phe Leu Val Pro145 150 155 160Gly Tyr Tyr Val Phe Ala Lys Asn Asn
Ile Ser Ala Ser Asp Phe Lys 165 170 175Arg Thr Val Asn Ala Gly His
Glu Thr Asn Ala Leu Ala Val Ala Asn 180 185 190Lys Gln Val Asp Val
Ala Thr Asn Asn Thr Glu Asn Leu Asp Lys Leu 195 200 205Lys Thr Ser
Ala Pro Glu Lys Leu Lys Glu Leu Lys Val Ile Trp Lys 210 215 220Ser
Pro Leu Ile Pro Gly Asp Pro Ile Val Trp Arg Lys Asn Leu Ser225 230
235 240Glu Thr Thr Lys Asp Lys Ile Tyr Asp Phe Phe Met Asn Tyr Gly
Lys 245 250 255Thr Pro Glu Glu Lys Ala Val Leu Glu Arg Leu Gly Trp
Ala Pro Phe 260 265 270Arg Ala Ser Ser Asp Leu Gln Leu Val Pro Ile
Arg Gln Leu Ala Leu 275 280 285Phe Lys Glu Met Gln Ser Val Lys Asp
Asn Lys Gly Leu Asn Glu Gln 290 295 300Asp Lys Leu Ala Lys Thr Thr
Ala Ile Gln Ala Gln Leu Asp Asp Leu305 310 315 320Asp Arg Leu Asn
Asn Ala Leu Ser Ala Met Ser Ser Val Ser Lys Ala 325 330 335Val
Gln108302PRTEscherichia coli 108Met Gln Leu Arg Lys Pro Ala Thr Ala
Ile Leu Ala Leu Ala Leu Ser1 5 10 15Ala Gly Leu Ala Gln Ala Asp Asp
Ala Ala Pro Ala Ala Gly Ser Thr 20 25 30Leu Asp Lys Ile Ala Lys Asn
Gly Val Ile Val Val Gly His Arg Glu 35 40 45Ser Ser Val Pro Phe Ser
Tyr Tyr Asp Asn Gln Gln Lys Val Val Gly 50 55 60Tyr Ser Gln Asp Tyr
Ser Asn Ala Ile Val Glu Ala Val Lys Lys Lys65 70 75 80Leu Asn Lys
Pro Asp Leu Gln Val Lys Leu Ile Pro Ile Thr Ser Gln 85 90 95Asn Arg
Ile Pro Leu Leu Gln Asn Gly Thr Phe Asp Phe Glu Cys Gly 100 105
110Ser Thr Thr Asn Asn Val Glu Arg Gln Lys Gln Ala Ala Phe Ser Asp
115 120 125Thr Ile Phe Val Val Gly Thr Arg Leu Leu Thr Lys Lys Gly
Gly Asp 130 135 140Ile Lys Asp Phe Ala Asn Leu Lys Asp Lys Ala Val
Val Val Thr Ser145 150 155 160Gly Thr Thr Ser Glu Val Leu Leu Asn
Lys Leu Asn Glu Glu Gln Lys 165 170 175Met Asn Met Arg Ile Ile Ser
Ala Lys Asp His Gly Asp Ser Phe Arg 180 185 190Thr Leu Glu Ser Gly
Arg Ala Val Ala Phe Met Met Asp Asp Ala Leu 195 200 205Leu Ala Gly
Glu Arg Ala Lys Ala Lys Lys Pro Asp Asn Trp Glu Ile 210 215 220Val
Gly Lys Pro Gln Ser Gln Glu Ala Tyr Gly Cys Met Leu Arg Lys225 230
235 240Asp Asp Pro Gln Phe Lys Lys Leu Met Asp Asp Thr Ile Ala Gln
Val 245 250 255Gln Thr Ser Gly Glu Ala Glu Lys Trp Phe Asp Lys Trp
Phe Lys Asn 260 265 270Pro Ile Pro Pro Lys Asn Leu Asn Met Asn Phe
Glu Leu Ser Asp Glu 275 280 285Met Lys Ala Leu Phe Lys Glu Pro Asn
Asp Lys Ala Leu Asn 290 295 300109414PRTThermus thermophilus 109Met
Arg Lys Trp Leu Leu Ala Ile Gly Met Val Leu Gly Leu Ser Ala1 5 10
15Leu Ala Gln Gly Gly Lys Leu Glu Ile Phe Ser Trp Trp Ala Gly Asp
20 25 30Glu Gly Pro Ala Leu Glu Ala Leu Ile Arg Leu Tyr Lys Gln Lys
Tyr 35 40 45Pro Gly Val Glu Val Ile Asn Ala Thr Val Thr Gly Gly Ala
Gly Val 50 55 60Asn Ala Arg Ala Val Leu Lys Thr Arg Met Leu Gly Gly
Asp Pro Pro65 70 75 80Asp Thr Phe Gln Val His Ala Gly Met Glu Leu
Ile Gly Thr Trp Val 85 90 95Val Ala Asn Arg Met Glu Asp Leu Ser Ala
Leu Phe Arg Gln Glu Gly 100 105 110Trp Leu Gln Ala Phe Pro Lys Gly
Leu Ile Asp Leu Ile Ser Tyr Lys 115 120 125Gly Gly Ile Trp Ser Val
Pro Val Asn Ile His Arg Ser Asn Val Met 130 135 140Trp Tyr Leu Pro
Ala Lys Leu Lys Glu Trp Gly Val Asn Pro Pro Arg145 150 155 160Thr
Trp Asp Glu Phe Leu Ala Thr Cys Gln Thr Leu Lys Gln Lys Gly 165 170
175Leu Glu Ala Pro Leu Ala Leu Gly Glu Asn Trp Thr Gln Gln His Leu
180 185 190Trp Glu Ser Val Ala Leu Ala Val Leu Gly Pro Asp Asp Trp
Asn Asn 195 200 205Leu Trp Asn Gly Lys Leu Lys Phe Thr Asp Pro Lys
Ala Val Arg Ala 210 215 220Trp Glu Val Phe Gly Arg Val Leu Asp Cys
Ala Asn Lys Asp Ala Ala225 230 235 240Gly Leu Ser Trp Gln Gln Ala
Val Asp Arg Val Val Gln Gly Lys Ala 245 250 255Ala Phe Asn Val Met
Gly Asp Trp Ala Ala Gly Tyr Met Thr Thr Thr 260 265 270Leu Lys Leu
Lys Pro Gly Thr Asp Phe Ala Trp Ala Pro Ser Pro Gly 275 280 285Thr
Gln Gly Val Phe Met Met Leu Ser Asp Ser Phe Gly Leu Pro Lys 290 295
300Gly Ala Lys Asn Arg Gln Asn Ala Ile Asn Trp Leu Arg Leu Val
Gly305 310 315 320Ser Lys Glu Gly Gln Asp Thr Phe Asn Pro Leu Lys
Gly Ser Ile Ala 325 330 335Ala Arg Leu Asp Ser Asp Pro Ser Lys Tyr
Asn Ala Tyr Gly Gln Ser 340 345 350Ala Met Arg Asp Trp Arg Ser Asn
Arg Ile Val Gly Ser Leu Val His 355 360 365Gly Ala Val Ala Pro Glu
Ser Phe Met Ser Gln Phe Gly Thr Val Met 370 375 380Glu Ile Phe Leu
Gln Thr Arg Asn Pro Gln Ala Ala Ala Asn Ala Ala385 390 395 400Gln
Ala Ile Ala Asp Gln Val Gly Leu Gly Arg Leu Gly Gln 405
410110318PRTRhizobium meliloti 110Met Ile Arg Thr Leu Ser Leu Lys
Phe Met Leu Ala Gly Ala Val Cys1 5 10 15Met Ala Thr Leu Thr Ala Gly
Ser Ala Phe Ala Ala Glu Pro Glu
Ser 20 25 30Cys Gly Thr Val Arg Phe Ser Asp Val Gly Trp Thr Asp Ile
Thr Ala 35 40 45Thr Thr Ala Thr Ala Thr Thr Ile Leu Glu Ala Leu Gly
Tyr Glu Thr 50 55 60Asp Val Lys Val Leu Ser Val Pro Val Thr Tyr Thr
Ser Leu Lys Asn65 70 75 80Lys Asp Ile Asp Val Phe Leu Gly Asn Trp
Met Pro Thr Met Glu Ala 85 90 95Asp Ile Ala Pro Tyr Arg Glu Asp Lys
Ser Val Glu Thr Val Arg Glu 100 105 110Asn Leu Ala Gly Ala Lys Tyr
Thr Leu Ala Thr Asn Ala Lys Gly Ala 115 120 125Glu Leu Gly Ile Lys
Asp Phe Lys Asp Ile Ala Ala His Lys Asp Glu 130 135 140Leu Asp Gly
Lys Ile Tyr Gly Ile Glu Pro Gly Asn Asp Gly Asn Arg145 150 155
160Leu Ile Ile Asp Met Val Glu Lys Gly Thr Phe Asp Leu Lys Gly Phe
165 170 175Glu Val Val Glu Ser Ser Glu Gln Gly Met Leu Ala Gln Val
Ala Arg 180 185 190Ala Glu Lys Ser Gly Asp Pro Ile Val Phe Leu Gly
Trp Glu Pro His 195 200 205Pro Met Asn Ala Asn Phe Lys Leu Thr Tyr
Leu Ser Gly Gly Asp Asp 210 215 220Val Phe Gly Pro Asn Tyr Gly Gly
Ala Thr Val His Thr Asn Val Arg225 230 235 240Ala Gly Tyr Thr Thr
Glu Cys Pro Asn Val Gly Lys Leu Leu Gln Asn 245 250 255Leu Ser Phe
Ser Leu Gln Met Glu Asn Glu Ile Met Gly Lys Ile Leu 260 265 270Asn
Asp Gly Glu Asp Pro Glu Lys Ala Ala Ala Ala Trp Leu Lys Asp 275 280
285Asn Pro Gln Ser Ile Glu Pro Trp Leu Ser Gly Val Ala Thr Lys Asp
290 295 300Gly Gly Asp Gly Leu Ala Ala Val Lys Ala Ala Leu Gly
Leu305 310 315111249PRTGeobacillus stearothermophilus 111Met Gly
Gly Gly Arg Ser Thr Glu Thr Ser Ser Ser Ser Gly Gly Asp1 5 10 15Gly
Gly Ala Thr Lys Lys Lys Val Val Val Gly Thr Asp Ala Ala Phe 20 25
30Ala Pro Phe Glu Tyr Met Gln Lys Gly Lys Ile Val Gly Phe Asp Val
35 40 45Asp Leu Leu Asp Ala Val Met Lys Ala Ala Gly Leu Asp Tyr Glu
Leu 50 55 60Lys Asn Ile Gly Trp Asp Pro Leu Phe Ala Ser Leu Gln Ser
Lys Glu65 70 75 80Val Asp Met Gly Ile Ser Gly Ile Thr Ile Thr Asp
Glu Arg Lys Gln 85 90 95Ser Tyr Asp Phe Ser Asp Pro Tyr Phe Glu Ala
Thr Gln Val Ile Leu 100 105 110Val Lys Gln Gly Ser Pro Val Lys Asn
Ala Leu Asp Leu Lys Gly Thr 115 120 125Ile Gly Val Gln Asn Ala Thr
Thr Gly Gln Glu Ala Ala Glu Lys Leu 130 135 140Phe Gly Lys Gly Pro
His Ile Lys Lys Phe Glu Thr Thr Val Val Ala145 150 155 160Ile Met
Glu Leu Leu Asn Gly Gly Val Asp Ala Val Ile Thr Asp Asn 165 170
175Ala Val Ala Asn Glu Tyr Val Lys Asn Asn Pro Asn Lys Lys Leu Gln
180 185 190Val Ile Glu Asp Pro Lys Asn Phe Ala Ser Glu Tyr Tyr Gly
Met Ile 195 200 205Phe Pro Lys Asn Ser Glu Leu Lys Ala Lys Val Asp
Glu Ala Leu Lys 210 215 220Asn Val Ile Asn Ser Gly Lys Tyr Thr Glu
Ile Tyr Lys Lys Trp Phe225 230 235 240Gly Lys Glu Pro Lys Leu Asp
Arg Leu 245112370PRTAgrobacterium tumefaciens 112Met Lys Lys Ser
Leu Leu Ser Ala Val Ala Leu Thr Ala Met Val Ala1 5 10 15Phe Gly Gly
Ser Ala Trp Ala Asp Val Val Ile Ala Val Gly Ala Pro 20 25 30Leu Thr
Gly Pro Asn Ala Ala Phe Gly Ala Gln Ile Gln Lys Gly Ala 35 40 45Glu
Gln Ala Ala Lys Asp Ile Asn Ala Ala Gly Gly Ile Asn Gly Glu 50 55
60Gln Ile Lys Ile Val Leu Gly Asp Asp Val Ser Asp Pro Lys Gln Gly65
70 75 80Ile Ser Val Ala Asn Lys Phe Val Ala Asp Gly Val Lys Phe Val
Val 85 90 95Gly His Phe Asn Ser Gly Val Ser Ile Pro Ala Ser Glu Val
Tyr Ala 100 105 110Glu Asn Gly Ile Leu Glu Ile Thr Pro Ala Ala Thr
Asn Pro Val Phe 115 120 125Thr Glu Arg Gly Leu Trp Asn Thr Phe Arg
Thr Cys Gly Arg Asp Asp 130 135 140Gln Gln Gly Gly Ile Ala Gly Lys
Tyr Leu Ala Asp His Phe Lys Asp145 150 155 160Ala Lys Val Ala Ile
Ile His Asp Lys Thr Pro Tyr Gly Gln Gly Leu 165 170 175Ala Asp Glu
Thr Lys Lys Ala Ala Asn Ala Ala Gly Val Thr Glu Val 180 185 190Met
Tyr Glu Gly Val Asn Val Gly Asp Lys Asp Phe Ser Ala Leu Ile 195 200
205Ser Lys Met Lys Glu Ala Gly Val Ser Ile Ile Tyr Trp Gly Gly Leu
210 215 220His Thr Glu Ala Gly Leu Ile Ile Arg Gln Ala Ala Asp Gln
Gly Leu225 230 235 240Lys Ala Lys Leu Val Ser Gly Asp Gly Ile Val
Ser Asn Glu Leu Ala 245 250 255Ser Ile Ala Gly Asp Ala Val Glu Gly
Thr Leu Asn Thr Phe Gly Pro 260 265 270Asp Pro Thr Leu Arg Pro Glu
Asn Lys Glu Leu Val Glu Lys Phe Lys 275 280 285Ala Ala Gly Phe Asn
Pro Glu Ala Tyr Thr Leu Tyr Ser Tyr Ala Ala 290 295 300Met Gln Ala
Ile Ala Gly Ala Ala Lys Ala Ala Gly Ser Val Glu Pro305 310 315
320Glu Lys Val Ala Glu Ala Leu Lys Lys Gly Ser Phe Pro Thr Ala Leu
325 330 335Gly Glu Ile Ser Phe Asp Glu Lys Gly Asp Pro Lys Leu Pro
Gly Tyr 340 345 350Val Met Tyr Glu Trp Lys Lys Gly Pro Asp Gly Lys
Phe Thr Tyr Ile 355 360 365Gln Gln 370113368PRTEscherichia coli
113Met Asn Ile Lys Gly Lys Ala Leu Leu Ala Gly Cys Ile Ala Leu Ala1
5 10 15Phe Ser Asn Met Ala Leu Ala Glu Asp Ile Lys Val Ala Val Val
Gly 20 25 30Ala Met Ser Gly Pro Val Ala Gln Tyr Gly Asp Gln Glu Phe
Thr Gly 35 40 45Ala Glu Gln Ala Val Ala Asp Ile Asn Ala Lys Gly Gly
Ile Lys Gly 50 55 60Asn Lys Leu Gln Ile Val Lys Tyr Asp Asp Ala Cys
Asp Pro Lys Gln65 70 75 80Ala Val Ala Val Ala Asn Lys Val Val Asn
Asp Gly Ile Lys Tyr Val 85 90 95Ile Gly His Leu Cys Ser Ser Ser Thr
Gln Pro Ala Ser Asp Ile Tyr 100 105 110Glu Asp Glu Gly Ile Leu Met
Ile Thr Pro Ala Ala Thr Ala Pro Glu 115 120 125Leu Thr Ala Arg Gly
Tyr Gln Leu Ile Leu Arg Thr Thr Gly Leu Asp 130 135 140Ser Asp Gln
Gly Pro Thr Ala Ala Lys Tyr Ile Leu Glu Lys Val Lys145 150 155
160Pro Gln Arg Ile Ala Ile Val His Asp Lys Gln Gln Tyr Gly Glu Gly
165 170 175Leu Ala Arg Ala Val Gln Asp Gly Leu Ile Lys Lys Gly Asn
Ala Asn 180 185 190Val Val Phe Phe Asp Gly Ile Thr Ala Gly Glu Lys
Asp Phe Ser Thr 195 200 205Leu Val Ala Arg Leu Lys Lys Glu Asn Ile
Asp Phe Val Tyr Tyr Gly 210 215 220Gly Tyr His Pro Glu Met Gly Gln
Ile Leu Arg Gln Ala Arg Ala Ala225 230 235 240Gly Leu Lys Thr Gln
Phe Met Gly Pro Glu Gly Val Ala Asn Val Ser 245 250 255Leu Ser Asn
Ile Ala Gly Glu Ser Ala Glu Gly Leu Leu Val Thr Lys 260 265 270Pro
Lys Asn Tyr Asp Gln Val Pro Ala Asn Lys Pro Ile Val Asp Ala 275 280
285Ile Lys Ala Lys Lys Gln Asp Pro Ser Gly Ala Phe Val Trp Thr Thr
290 295 300Tyr Ala Ala Leu Gln Ser Leu Gln Ala Gly Leu Asn Gln Ser
Asp Asp305 310 315 320Pro Ala Glu Ile Ala Lys Tyr Leu Lys Ala Asn
Ser Val Asp Thr Val 325 330 335Met Gly Pro Leu Thr Trp Asp Glu Lys
Gly Asp Leu Lys Gly Phe Glu 340 345 350Phe Gly Val Phe Asp Trp His
Ala Asn Gly Thr Ala Thr Asp Ala Lys 355 360 365114677PRTArtificial
Sequencerecombinant peptide biosensor 114Met Arg Gly Ser His His
His His His His Gly Met Ala Ser Met Thr1 5 10 15Gly Gly Gln Gln Met
Gly Arg Asp Leu Tyr Asp Asp Asp Asp Lys Asp 20 25 30Arg Trp Gly Ser
Lys Leu Glu Ile Phe Ser Trp Trp Ala Gly Asp Glu 35 40 45Gly Pro Ala
Leu Glu Ala Leu Ile Arg Leu Tyr Lys Gln Lys Tyr Pro 50 55 60Gly Val
Glu Val Ile Asn Ala Thr Val Thr Gly Gly Ala Gly Val Asn65 70 75
80Ala Arg Ala Val Leu Lys Thr Arg Met Leu Gly Gly Asp Pro Pro Asp
85 90 95Thr Phe Gln Val Ala Ala Gly Met Glu Leu Ile Gly Thr Trp Val
Val 100 105 110Ala Asn Arg Met Glu Asp Leu Ser Ala Leu Phe Arg Gln
Glu Gly Trp 115 120 125Leu Gln Ala Phe Pro Lys Gly Leu Ile Asp Leu
Ile Ser Tyr Lys Gly 130 135 140Gly Ile Trp Ser Val Pro Val Asn Ile
His Arg Ser Asn Val Met Trp145 150 155 160Tyr Leu Pro Ala Lys Leu
Lys Glu Trp Gly Val Asn Pro Pro Arg Thr 165 170 175Trp Glu Phe Leu
Ala Thr Cys Gln Thr Leu Lys Gln Lys Gly Leu Glu 180 185 190Ala Pro
Leu Ala Leu Gly Glu Asn Trp Thr Gln Gln His Leu Trp Glu 195 200
205Ser Val Ala Leu Ala Val Leu Gly Pro Asp Asp Trp Asn Asn Leu Trp
210 215 220Asn Gly Lys Leu Lys Phe Thr Asp Pro Lys Ala Val Arg Ala
Trp Glu225 230 235 240Val Phe Gly Arg Val Leu Asp Cys Ala Asn Lys
Asp Ala Ala Gly Leu 245 250 255Ser Trp Gln Gln Ala Val Asp Arg Val
Val Gln Gly Lys Ala Ala Phe 260 265 270Asn Val Met Gly Asp Trp Ala
Ala Gly Tyr Met Thr Thr Thr Leu Lys 275 280 285Leu Lys Pro Gly Thr
Asp Phe Ala Trp Ala Pro Ser Pro Gly Thr Gln 290 295 300Gly Val Phe
Met Met Val Ser Asp Ser Phe Gly Leu Pro Lys Gly Ala305 310 315
320Lys Asn Arg Gln Asn Ala Ile Asn Trp Leu Arg Leu Val Gly Ser Lys
325 330 335Glu Gly Gln Asp Thr Phe Asn Pro Leu Lys Gly Ser Ile Ala
Ala Arg 340 345 350Leu Asp Ser Asp Pro Ser Lys Tyr Pro Ala Ser His
Asn Val Tyr Ile 355 360 365Met Ala Asp Lys Gln Arg Asn Gly Ile Lys
Ala Asn Phe Lys Ile Arg 370 375 380His Asn Ile Glu Asp Gly Gly Val
Gln Leu Ala Tyr His Tyr Gln Gln385 390 395 400Asn Thr Pro Ile Gly
Asp Gly Pro Val Leu Leu Pro Asp Asn His Tyr 405 410 415Leu Ser Thr
Gln Ser Lys Leu Ser Lys Asp Pro Asn Glu Lys Arg Asp 420 425 430His
Met Val Leu Leu Glu Phe Val Thr Ala Ala Gly Ile Thr Leu Gly 435 440
445Met Asp Glu Leu Tyr Lys Gly Gly Thr Gly Gly Ser Met Val Ser Lys
450 455 460Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu Val Glu
Leu Asp465 470 475 480Gly Asp Val Asn Gly His Lys Phe Ser Val Ser
Gly Glu Gly Glu Gly 485 490 495Asp Ala Thr Tyr Gly Lys Leu Thr Leu
Lys Phe Ile Cys Thr Thr Gly 500 505 510Lys Leu Pro Val Pro Trp Pro
Thr Leu Val Thr Thr Leu Thr Tyr Gly 515 520 525Val Gln Cys Phe Ser
Arg Tyr Pro Asp His Met Lys Gln His Asp Phe 530 535 540Phe Lys Ser
Ala Met Pro Glu Gly Tyr Ile Gln Glu Arg Thr Ile Phe545 550 555
560Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu Val Lys Phe Glu
565 570 575Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys Gly Ile Asp
Phe Lys 580 585 590Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu Tyr
Asn Phe Asn Asn 595 600 605Pro Asn Ala Tyr Gly Gln Ser Ala Met Arg
Asp Trp Arg Ser Asn Arg 610 615 620Ile Val Gly Ser Leu Val Ala Gly
Ala Val Ala Pro Glu Ser Phe Met625 630 635 640Ser Gln Phe Gly Thr
Val Met Glu Ile Phe Leu Gln Thr Arg Asn Pro 645 650 655Gln Ala Ala
Ala Asn Ala Ala Gln Ala Ile Ala Asp Gln Val Gly Leu 660 665 670Gly
Arg Leu Gly Gln 6751157PRTArtificial Sequencesynthetic
oligopeptideVARIANT1Xaa = Any Amino Acid 115Xaa Pro Ser His Asn Val
Tyr1 51167PRTArtificial Sequencesynthetic oligopeptideVARIANT1,
2Xaa = Any Amino Acid 116Xaa Xaa Ser His Asn Val Tyr1
51177PRTArtificial Sequencesynthetic oligopeptideVARIANT1, 2Xaa =
Any Amino Acid 117Xaa Xaa Ser His Asn Val Phe1 51187PRTArtificial
Sequencesynthetic oligopeptideVARIANT2Xaa = Any Amino Acid 118Pro
Xaa Ser His Asn Val Phe1 51197PRTArtificial Sequencesynthetic
oligopeptideVARIANT2Xaa = Any Amino Acid 119Pro Xaa Ser Tyr Asn Val
Phe1 51207PRTArtificial Sequencesynthetic oligopeptideVARIANT1,
2Xaa = Any Amino Acid 120Xaa Xaa Ser Tyr Asn Val Phe1
51217PRTArtificial Sequencesynthetic oligopeptideVARIANT2Xaa = Any
Amino Acid 121Pro Xaa Ser Tyr Asn Val Phe1 51227PRTArtificial
Sequencesynthetic oligopeptideVARIANT1, 2Xaa = Any Amino Acid
122Xaa Xaa Ser Tyr Asn Val Phe1 51237PRTArtificial
Sequencesynthetic oligopeptideVARIANT2, 4Xaa = Any Amino Acid
123Pro Xaa Ser Xaa Asn Val Tyr1 51247PRTArtificial
Sequencesynthetic oligopeptideVARIANT2, 5Xaa = Any Amino Acid
124Pro Xaa Ser His Xaa Val Tyr1 51257PRTArtificial
Sequencesynthetic oligopeptideVARIANT2, 6Xaa = Any Amino Acid
125Pro Xaa Ser His Asn Xaa Tyr1 51267PRTArtificial
Sequencesynthetic oligopeptideVARIANT2, 7Xaa = Any Amino Acid
126Pro Xaa Ser His Asn Val Xaa1 51275PRTArtificial
Sequencesynthetic oligopeptideVARIANT3, 4Xaa = Any Amino Acid
127Phe Asn Xaa Xaa Tyr1 51284PRTArtificial Sequencesynthetic
oligopeptideVARIANT3Xaa = Any Amino Acid 128Phe Asn Xaa
Tyr11293PRTArtificial Sequencesynthetic oligopeptide 129Phe Asn
Tyr11303PRTArtificial Sequencesynthetic oligopeptideVARIANT2Xaa =
Any Amino Acid 130Phe Xaa Tyr11313PRTArtificial Sequencesynthetic
oligopeptideVARIANT1, 2Xaa = Any Amino Acid 131Xaa Xaa
Tyr11323PRTArtificial Sequencesynthetic oligopeptideVARIANT2Xaa =
Any Amino Acid 132Trp Xaa Tyr11333PRTArtificial Sequencesynthetic
oligopeptideVARIANT1Xaa = Any Amino Acid 133Xaa Lys
Tyr11345PRTArtificial Sequencesynthetic oligopeptideVARIANT4Xaa =
Any Amino Acid 134Phe Asn Pro Xaa Tyr1 51355PRTArtificial
Sequencesynthetic oligopeptideVARIANT3Xaa = Any Amino Acid 135Phe
Asn Xaa Pro Tyr1 51363PRTArtificial Sequencesynthetic oligopeptide
136His Asn Ser11373PRTArtificial Sequencesynthetic oligopeptide
137Gly Gly Ser11383PRTArtificial Sequencesynthetic
oligopeptideVARIANT1, 2Xaa = Any Amino Acid 138Xaa Xaa
Ser11393PRTArtificial Sequencesynthetic oligopeptideVARIANT1, 2Xaa
= Any Amino Acid 139Xaa Xaa Lys11403PRTArtificial Sequencesynthetic
oligopeptide 140Gly Gly Lys11413PRTArtificial Sequencesynthetic
oligopeptideVARIANT2Xaa = Any Amino Acid 141Pro Xaa
Ser11423PRTArtificial Sequencesynthetic oligopeptideVARIANT1Xaa =
Any Amino Acid 142Xaa Pro Ser11432PRTArtificial Sequencesynthetic
oligopeptideVARIANT2Xaa = Any Amino Acid 143Pro
Xaa11442PRTArtificial Sequencesynthetic oligopeptideVARIANT1Xaa =
Any Amino Acid 144Xaa Pro11454PRTArtificial Sequencesynthetic
oligopeptideVARIANT2, 3Xaa = Any Amino Acid 145Ile Xaa Xaa
Ser11464PRTArtificial Sequencesynthetic oligopeptideVARIANT2Xaa =
Any Amino Acid 146Asn Xaa Pro Lys11474PRTArtificial
Sequencesynthetic
oligopeptide 147Asn Pro Cys Lys11485PRTArtificial Sequencesynthetic
oligopeptideVARIANT3Xaa = Any Amino Acid 148Pro Pro Xaa Ser His1
51496PRTArtificial Sequencesynthetic oligopeptideVARIANT3, 4Xaa =
Any Amino Acid 149Pro Pro Xaa Xaa Ser His1 51506PRTArtificial
Sequencesynthetic oligopeptideVARIANT4Xaa = Any Amino Acid 150Pro
Pro Pro Xaa Ser His1 51516PRTArtificial Sequencesynthetic
oligopeptideVARIANT3Xaa = Any Amino Acid 151Pro Pro Xaa Pro Ser
His1 51524PRTArtificial Sequencesynthetic oligopeptideVARIANT1,
2Xaa = Any Amino Acid 152Xaa Xaa Ser His11534PRTArtificial
Sequencesynthetic oligopeptideVARIANT3, 4Xaa = Any Amino Acid
153Pro Pro Xaa Xaa11545PRTArtificial Sequencesynthetic
oligopeptideVARIANT3Xaa = Any Amino Acid 154Phe Asn Xaa Lys Asn1
51556PRTArtificial Sequencesynthetic oligopeptideVARIANT3, 4Xaa =
Any Amino Acid 155Phe Asn Xaa Xaa Lys Asn1 51566PRTArtificial
Sequencesynthetic oligopeptideVARIANT3Xaa = Any Amino Acid 156Phe
Asn Xaa Pro Lys Asn1 51576PRTArtificial Sequencesynthetic
oligopeptideVARIANT4Xaa = Any Amino Acid 157Phe Asn Pro Xaa Lys
Asn1 51584PRTArtificial Sequencesynthetic oligopeptideVARIANT3,
4Xaa = Any Amino Acid 158Phe Asn Xaa Xaa11597PRTArtificial
Sequencesynthetic oligopeptide 159Asn Ala Asp Gly Ser Ser His1
51606PRTArtificial Sequencesynthetic oligopeptideVARIANT3, 4Xaa =
Any Amino Acid 160Ala Asp Xaa Xaa Ser His1 51616PRTArtificial
Sequencesynthetic oligopeptideVARIANT3Xaa = Any Amino Acid 161Ala
Asp Xaa Pro Ser His1 51626PRTArtificial Sequencesynthetic
oligopeptideVARIANT4Xaa = Any Amino Acid 162Ala Asp Pro Xaa Ser
His1 51634PRTArtificial Sequencesynthetic oligopeptideVARIANT3,
4Xaa = Any Amino Acid 163Ala Asp Xaa Xaa11646PRTArtificial
Sequencesynthetic oligopeptideVARIANT3, 4Xaa = Any Amino Acid
164Ala Asp Xaa Xaa Ser His1 51654PRTArtificial Sequencesynthetic
oligopeptide 165Phe Asn Pro Gly11666PRTArtificial Sequencesynthetic
oligopeptideVARIANT3, 4Xaa = Any Amino Acid 166Phe Asn Xaa Xaa Pro
Gly1 51674PRTArtificial Sequencesynthetic oligopeptideVARIANT1,
2Xaa = Any Amino Acid 167Xaa Xaa Pro Gly11684PRTArtificial
Sequencesynthetic oligopeptideVARIANT3, 4Xaa = Any Amino Acid
168Phe Asn Xaa Xaa11694PRTArtificial Sequencesynthetic
oligopeptideVARIANT4Xaa = Any Amino Acid 169Phe Asn Pro
Xaa11706PRTArtificial Sequencesynthetic oligopeptideVARIANT3, 4Xaa
= Any Amino Acid 170Lys Tyr Xaa Xaa Ser His1 51716PRTArtificial
Sequencesynthetic oligopeptideVARIANT4Xaa = Any Amino Acid 171Lys
Tyr Pro Xaa Ser His1 51726PRTArtificial Sequencesynthetic
oligopeptideVARIANT3Xaa = Any Amino Acid 172Lys Tyr Xaa Pro Ser
His1 51734PRTArtificial Sequencesynthetic oligopeptideVARIANT2,
3Xaa = Any Amino Acid 173Phe Xaa Xaa Pro11744PRTArtificial
Sequencesynthetic oligopeptideVARIANT3Xaa = Any Amino Acid 174Phe
Asn Xaa Pro11754PRTArtificial Sequencesynthetic
oligopeptideVARIANT4Xaa = Any Amino Acid 175Phe Asn Pro
Xaa11767PRTArtificial Sequencesynthetic oligopeptideVARIANT2Xaa =
Any Amino Acid 176Pro Xaa Ser His Asn Val Tyr1 5
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025

D00026

D00027

D00028

D00029
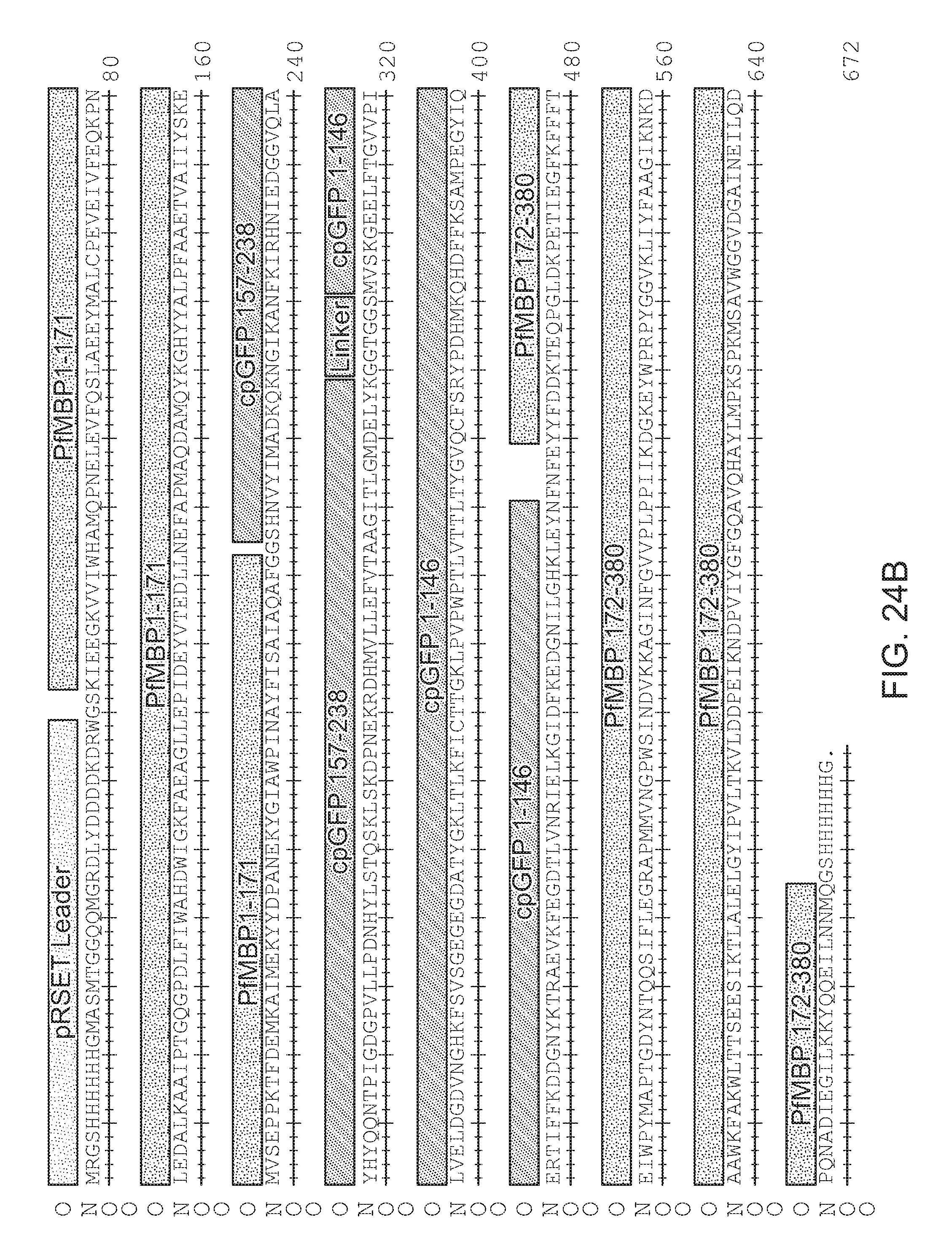
D00030

D00031

D00032

D00033

D00034

D00035

D00036

D00037

D00038

D00039

D00040

D00041

D00042

D00043

D00044

D00045

D00046

D00047

D00048

D00049

D00050

D00051

D00052

D00053

D00054

D00055

D00056

D00057

D00058

D00059

D00060

D00061

D00062
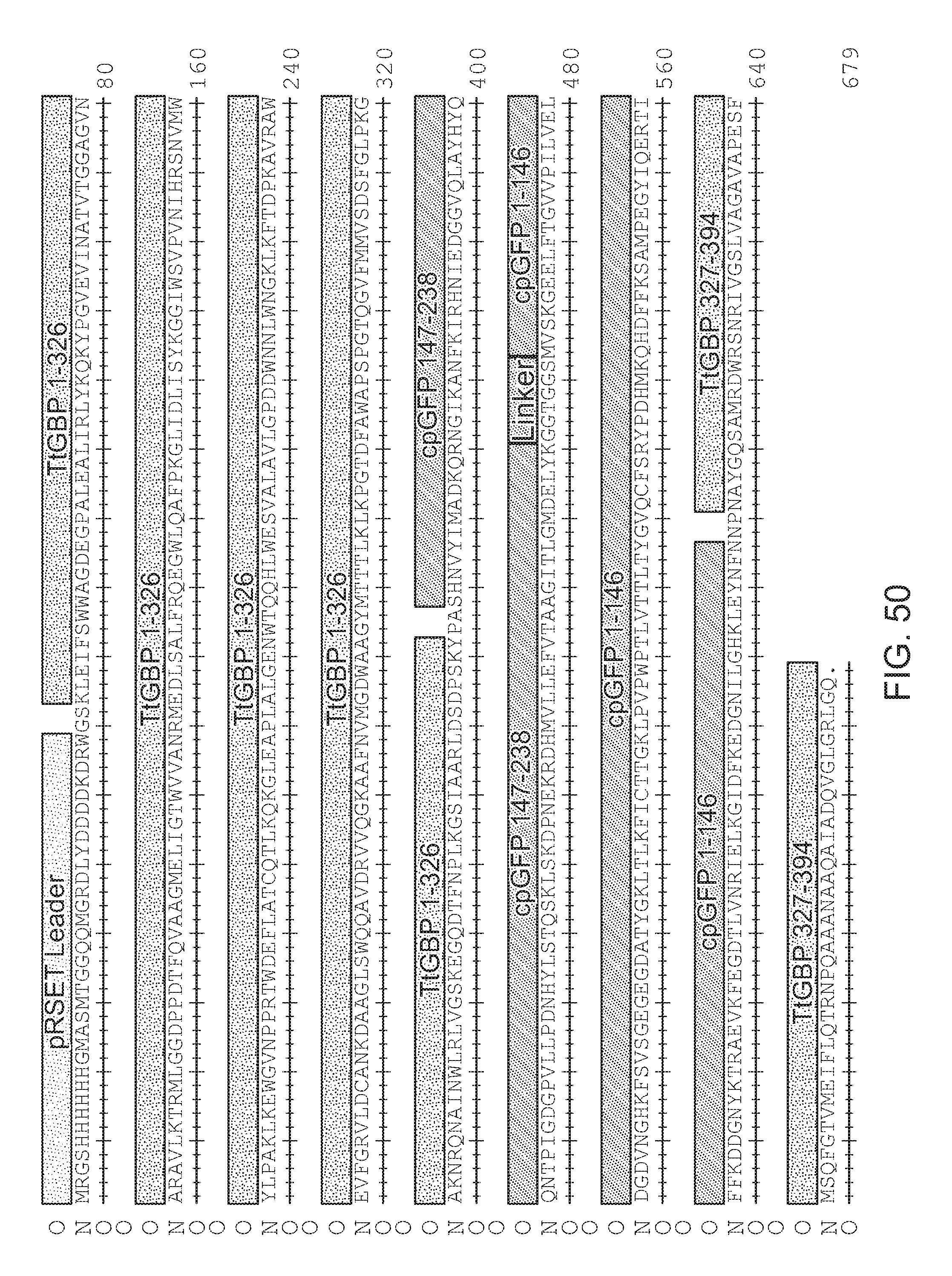
D00063

D00064

D00065

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.