Control Of Protein To Protein Interactions Of Acid Decarboxylase
Chou; Howard ; et al.
U.S. patent application number 16/464104 was filed with the patent office on 2019-10-31 for control of protein to protein interactions of acid decarboxylase. The applicant listed for this patent is CATHAY R&D CENTER CO., LTD., CIBT America Inc.. Invention is credited to Ling Chen, Howard Chou, Xiucai Liu, Wenqiang Lu.
| Application Number | 20190330613 16/464104 |
| Document ID | / |
| Family ID | 62194569 |
| Filed Date | 2019-10-31 |


| United States Patent Application | 20190330613 |
| Kind Code | A1 |
| Chou; Howard ; et al. | October 31, 2019 |
CONTROL OF PROTEIN TO PROTEIN INTERACTIONS OF ACID DECARBOXYLASE
Abstract
This invention provides acid decarboxylase-prion subunit fusion polypeptides, nucleic acid sequences, expression vectors, and host cells expression such fusion polypeptides to produce various amino acids and derivatives of the amino acids such as polyamines.
| Inventors: | Chou; Howard; (Shanghai, CN) ; Chen; Ling; (Shanghai, CN) ; Lu; Wenqiang; (Shanghai, CN) ; Liu; Xiucai; (Shanghai, CN) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 62194569 | ||||||||||
| Appl. No.: | 16/464104 | ||||||||||
| Filed: | November 24, 2016 | ||||||||||
| PCT Filed: | November 24, 2016 | ||||||||||
| PCT NO: | PCT/CN2016/107083 | ||||||||||
| 371 Date: | May 24, 2019 |
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 14/47 20130101; C07K 2319/00 20130101; C12Y 401/01018 20130101; C12N 9/88 20130101; C12N 15/70 20130101 |
| International Class: | C12N 9/88 20060101 C12N009/88; C07K 14/47 20060101 C07K014/47; C12N 15/70 20060101 C12N015/70 |
Claims
1. A product, which is one of the following products I) through IV): I) a genetically modified host cell comprising a nucleic acid encoding an acid decarboxylase fusion protein comprising an acid decarboxylase polypeptide joined to a prion subunit fused to the carboxyl end of the acid decarboxylase polypeptide, wherein acid decarboxylase fusion protein has increased activity relative to the acid decarboxylase polypeptide not joined to the prion subunit; II) an acid decarboxylase fusion protein comprising an acid decarboxylase polypeptide fused to a prion subunit, wherein the fusion protein has improved acid decarboxylase activity in vitro as measured by the production of polyamines at alkaline pH, relative to a counterpart fusion protein lacking the prion subunit; III) a polynucleotide encoding a fusion protein of II); IV) an expression vector comprising the polynucleotide of III), operably linked to a promoter.
2. A product of claim 1, which is I) the genetically modified host cell, wherein the prion subunit is at least 50 amino acids in length, at least 75 amino acids in length or at least 100 amino acids in length, but 500 amino acids or fewer in length.
3. A product of claim 1, which is I) the genetically modified host cell of claim 1, wherein the prion subunit comprises an amino acid composition having at least 20% glutamine and/or asparagine residues.
4. A product of claim 1, which is I) the genetically modified host cell, where the prion subunit: (i) has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to a Sup35, New1, Ure2, or Rnq1 amino acid sequence; (ii) comprises a Sup35, New1, Ure2, or Rnq1 amino acid sequence; (iii) has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to an amino acid sequence set forth in SEQ ID NO:3 or SEQ ID NO:4; (iv) comprises SEQ ID NO:3 or SEQ ID NO:4; or (v) has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of the prion subunit region of any one of SEQ ID NOS: 7, 8, 11, 13, 14, 15, 17, or 19.
5. (canceled)
6. (canceled)
5. A product of claim 1, which is I) the genetically modified host cell, wherein the prion subunit is joined at the carboxyl terminus to a BST fragment, .lamda.CI fragment, or RecA fragment.
6. A product of claim 1, which is I) the genetically modified host, wherein prion subunit: (a) is joined at the C-terminal end to a BST fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:15, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:15, excluding the linker region; (b) is joined at the C-terminal end to a .DELTA.CI fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:16, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:16, excluding the linker region; (c) is joined at the C-terminal end to a RecA fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:19, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:19, excluding the linker region; or (d) is joined at the carboxyl terminus to an amino acid sequence RRFGEASSAF, ASQWPEETFG, or EGVAETNEDF.
9. (canceled)
7. A product of claim 1, which is I) the genetically modified host cell, wherein the acid decarboxylase is a lysine decarboxylase, ornithine decarboxylase, arginine decarboxylase, or glutamate decarboxylase.
8. A product of claim 1, which is I) the genetically modified host cell of, wherein the acid decarboxylase is a CadA, LdcC, AdiA, SpeA, SpeC, SpeF, GadA, or GadB polypeptide.
12. (canceled)
13. (canceled)
9. A product of claim 8, wherein the lysine decarboxylase is a CadA lysine decarboxylase or a LdcC lysine decarboxylase.
10. A product of claim 8, wherein the host cell is genetically modified to over express one or more lysine biosynthesis polypeptides.
11. A product of claim 1, which is I) the genetically modified host cell, wherein the nucleic acid encoding the acid decarboxylase fusion protein is encoded by an expression vector introduced into the cell, wherein the expression vector comprises the nucleic acid encoding the acid decarboxylase fusion protein operably linked to a promoter, and/or the nucleic acid encoding the acid decarboxylase fusion protein is integrated into the host chromosome.
17. (canceled)
12. A product of claim 1, which is I) the genetically modified host cell, wherein the nucleic acid encoding the acid decarboxylase fusion polypeptide comprises the nucleic acid sequence of any one of SEQ ID NOS:1, 5, 6, 9, 10, 16, 18, or 20.
13. A product of claim 1, which is I) the genetically modified host cell, wherein the host cell is a bacterium.
14. A product of claim 13, wherein the host cell is from the genus Escherichia or Hafnia.
15. A product of claim 14, wherein the host cell is Escherichia coli or Hafnia alvei.
16. A method, which is one of the following methods I) through III): (I) a method for producing an acid decarboxylase fusion protein comprising cultivating a host cell of claim 1 I) under conditions in which the acid decarboxylase fusion protein is expressed; (II) a method of producing an amino acid or an amino acid derivative, the method comprising culturing a host cell of claim 1 I) under conditions in which the acid decarboxylase fusion protein is expressed; (III) a method of improving acid decarboxylase activity in vitro under alkaline pH comprising fusing a prion subunit to the carboxyl terminus of an acid decarboxylase, and subjecting the recombinant protein to alkaline pH.
23. (canceled)
24. (canceled)
25. (canceled)
17. A product of claim 1, which is II) the fusion protein, wherein the prion subunit: (i) is 30 amino acids in length, at least 50 amino acids in length, at least 75 amino acids in length or at least 100 amino acids in length, but 1200 amino acids or fewer in length; (ii) comprises an amino acid composition having at least 20% glutamine and/or asparagine residues; (iii) has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to a Sup35, New1, Ure2, or Rnq1 amino acid sequence; (iv) comprises a Sup35, New1, Ure2, or Rnq1 amino acid sequence; (v) has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to an amino acid sequence set forth in SEQ ID NO: 3 or SEQ ID NO: 4; (vi) comprises SEQ ID NO: 3 or SEQ ID NO: 4; or (vii) has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of the prion subunit region of any one of SEQ ID NOS: 7, 8, 11, 13, 14, 15, 17, or 19.
27. (canceled)
28. (canceled)
29. (canceled)
30. (canceled)
18. A product of claim 1, which is II) the fusion protein wherein the prion subunit: (i) is joined to the C-terminus of the acid decarboxylase; (ii) is joined at the carboxyl terminus to a BST fragment, .DELTA.CI fragment, or RecA fragment; (iii) is joined the carboxyl terminus to an amino acid sequence RRFGEASSAF, ASQWPEETFG, or EGVAETNEDF; (iv) is joined at the C-terminal end to a BST fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO: 15, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO: 15, excluding the linker region; (v) is joined at the C-terminal end to a .DELTA.CI fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO: 16, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO: 16, excluding the linker region; or (vi) is joined at the C-terminal end to a RecA fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO: 19, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO: 19, excluding the linker region.
32. (canceled)
33. (canceled)
34. (canceled)
35. (canceled)
19. A product of claim 1, which is II) the fusion protein, wherein the acid decarboxylase is a CadA, LdcC, AdiA, SpeA, SpeC, SpeF, GadA, or GadB polypeptide.
37. (canceled)
38. (canceled)
39. (canceled)
40. (canceled)
20. A product of claim 1, which is II) the fusion protein, wherein the fusion protein is immobilized to a solid support.
42. (canceled)
43. (canceled)
Description
BACKGROUND OF THE INVENTION
[0001] Most enzymes function optimally within a narrow pH range, because they are amphoteric molecules. The pH of the surrounding environment directly affects the charges on the acidic and basic groups of the amino acids that make up the enzyme. These changes in charge affect the net charge of the enzyme, the pKa of the active site, and the charge distribution across the surface of the enzyme. As a result, changes in pH can affect the activity, solubility, and stability of an enzyme.
[0002] The class of proteins known as acid decarboxylases is a group of enzymes that catalyze the decarboxylation reaction of amino acids, e.g., basic amino acids such as lysine, arginine, ornithine, in order to generate products, e.g., polyamines, as part of the acid stress response in many microorganisms. Escherichia coli has six pyridoxal 5'-phosphate (PLP)-inducible acid decarboxylases: CadA, LdcC, AdiA, SpeA, SpeC, SpeF, GadA, and GadB. All of these enzymes function within a narrow pH range, and the enzyme's activity decreases significantly outside of that pH range (Kanjee et al., Biochemistry 50, 9388-9398, 2011). It has been previously observed that these PLP-dependent decarboxylases dimerize in order to form a complete active site. In some cases, such as CadA, the dimers form decamers that aggregate into higher molecular weight protein complexes required for optimal function. The inhibition of higher molecular weight protein complex formation (e.g., in conditions outside of the optimal pH) leads to a significant decrease in function (Kanjee et al., The EMBO Journal 30, 931-944, 2011).
[0003] Previous studies on the production of polyamines focused on the overexpression of various acid decarboxylases. However, there has not been any study on increasing the stability of the enzyme's activity under various stresses such as alkaline pH. Tolerance to alkaline pH by acid decarboxylases is important, because the polyamines they generate as a product increase the pH of the reaction environment. Therefore, the activity of the acid decarboxylase usually decreases as more polyamines are generated, which can cause the decarboxylation reaction to stop prematurely when the pH of the reaction environment surpasses the pH range tolerated by the acid decarboxylase.
[0004] The typical process to produce polyamines (e.g., cadaverine) uses a process and fermentation medium similar to those used to produce amino acids (e.g., lysine) (Qian et al., Biotechnol. Bioeng. 108, 93-103, 2010). For example, ammonium sulfate is the major nitrogen source due to its ability to provide nitrogen and being slightly acidic (0.1M solution has a pH 5.5). The acidic pH preserves the pH range for an acid decarboxylase, such as lysine decarboxylases, e.g., CadA, to function. However, the use of ammonium sulfate leaves sulfate ions in the medium, which becomes a byproduct that is a salt waste during the fermentation process. The ability to tolerate alkaline pH allows for the use alternative nitrogen sources, and the production of less salt waste during the fermentation process.
[0005] Prions were identified as the infectious agent that causes transmissible spongiform encephalopathy (similar to "mad cow disease", sheep scrapie, human kuru, and Creutzfeldt, Jacob disease). For a review on prions, see Derkatch & Liebman, Prion 1:3, 161-169, 2007. Prions are protein conformations that are infectious. The protein may have other roles in the cell when they are not in the prion conformation. Proteins that form the prion conformation are not homologous, but some are rich in glutamine or asparagine residues. Prion aggregates are highly ordered, and typically form through intermolecular interactions between beta-strands. Therefore, prion conformations are beta-sheet rich, and assemble into structures that resemble amyloid fibers. See, e.g., Derkatch & Liebman, Prion 1:3, 161-169, 2007,
[0006] Prions have also been identified in yeast, and were first observed in two yeast determinants: [PSI.sup.] and [URE3] (Kushnirov & Ter-Avanesyan, Cell 94, 13-16, 1998). It was observed that the formation of prions could be induced by overexpression of the proteins Sup35 or Ure2. It was found that the Sup35 and Ure2 proteins in the yeast cells that have [PSI.sup.+] or [URE3] phenotypes show increase protease resistance and are found in a high-molecular weight aggregated state. In addition to the Sup35 and Ure2 proteins, two other proteins were identified in yeast with the ability to form prion conformations--New1 and Rnq1 (Osherovich et al., PLOS Biology 2, 442-451, 2004) LikeSup35 and Ure2, New1 and Rnq1 also have long series of sequences rich in glutamine and asparagine. Previously, Sup35 has been fused to GST in pGEX-4T-3 (a 25 kD protein) (Ono et al., Biosci. Biotechnol. Biochem. 70, 2813-2823, 2006), and Sup35, New1, and Rnq1 have been fused to GFP (a 27 kD protein) (Garrity et al., PNAS 107, 10596-10601, 2010). These proteins are relatively small, however. An acid decarboxylase monomer is about 81 kD (more than three times larger than GST) and forms higher molecular weight structures, e.g., that are larger than 1620 kD (more than sixty times larger than GST), in order to function. There have been no studies evaluating whether prions can be fused to much larger proteins without affecting function.
BRIEF SUMMARY OF ASPECTS OF THE DISCLOSURE
[0007] This invention is based, in part, on the surprising discovery that fusing a prion protein to an acid decarboxylase increases the stability of the enzyme's activity under various stresses that typically cause the protein complex to transition from a high oligomerization state to a low oligomerization state (e.g., alkaline pH and high temperature).
[0008] In one aspect, the disclosure thus provides a genetically modified host cell comprising a nucleic acid encoding an acid decarboxylase fusion protein comprising an acid decarboxylase polypeptide joined to a prion subunit fused to the carboxyl end of the acid decarboxylase polypeptide, wherein acid decarboxylase fusion polypeptide has increased activity relative to the acid decarboxylase polypeptide not joined to the prion subunit. In some embodiments, the prion subunit is at least 50 amino acids in length, at least 75 amino acids in length or at least 100 amino acids in length, but 500 amino acids or fewer in length. The prion subunit typically has an amino acid composition of 10% or greater glutamine and/or asparagine residues. In some embodiments, the prion subunit comprises an amino acid composition having at least 20% glutamine and/or asparagine residues. In some embodiments, the prion subunit has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to a Sup35, New1, Ure2, or Rnq1 amino acid sequence; or comprises a Sup35, New1, Ure2, or Rnq1 amino acid sequence. In some embodiments, the prion subunit has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to an amino acid sequence set forth in SEQ ID NO:3 or SEQ ID NO:4; or comprises the amino acid sequence of SEQ ID NO:3 or SEQ ID NO:4. In further embodiments, the prion subunit is joined at the carboxyl terminus to a BST fragment, .lamda.CI fragment, or RecA fragment, for example a fragment having the amino acid sequence RRFGEASSAF, ASQWPEETFG, or EGVAETNEDF. In some embodiments, the prion subunit is joined at the C-terminal end to a BST fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:15, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:15, excluding the linker region. In some embodiments, the prion subunit is joined at the C-terminal end to a .lamda.CI fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:16, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:16, excluding the linker region. In some embodiments, the prion subunit is joined at the C-terminal end to a RecA fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:19, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:19, excluding the linker region. In some embodiments, the acid decarboxylase is a lysine decarboxylase, ornithine decarboxylase, glutamate decarboxylase, or arginine decarboxylase. In some embodiments, the acid decarboxylase is a CadA, LdcC, AdiA, SpeA, SpeC, SpeF, GadA, or GadB polypeptide. For example, in some embodiments, the acid decarboxylase is a lysine decarboxylase, such as a CadA lysine decarboxylase polypeptide or a LdcC polypeptide. In some embodiments where the acid decarboxylase is a lysine decarboxylase, the host cell is genetically modified to over express one or more lysine biosynthesis polypeptides. In some embodiments, the nucleic acid encoding the acid decarboxylase fusion protein is encoded by an expression vector introduced into the cell, wherein the expression vector comprises the nucleic acid encoding the acid decarboxylase fusion protein operably linked to a promoter. In alternative embodiments, the nucleic acid encoding the acid decarboxylase fusion protein is integrated into the host chromosome. The host cell may be a bacterium, such as a bacterium from the genus Escherichia or Hafnia. In some embodiments, the host cell is Escherichia coli or Hafnia alvei.
[0009] In a further aspect, the invention provides a method for producing an acid decarboxylase fusion protein comprising cultivating a host cell as described in the preceding paragraph under conditions in which the acid decarboxylase fusion protein is expressed. In another aspect, the invention provides a method of producing an amino acid or an amino acid derivative, the method comprising culturing a host cell as described in the preceding paragraph under conditions in which the acid decarboxylase fusion polypeptide is expressed.
[0010] The invention additionally provides a method of improving acid decarboxylase activity in vitro under alkaline pH and/or high temperature. In some embodiments, the method comprises fusing a prion subunit to the carboxyl terminus of an acid decarboxylase and subjecting the fusion protein to alkaline pH. In some embodiments, the method comprises fusing a prion subunit to the carboxyl terminus of an acid decarboxylase and subjecting the fusion protein to high temperature.
[0011] In an additional aspect, the invention provides an acid decarboxylase fusion protein comprising an acid decarboxylase polypeptide fused to a prion subunit, wherein the fusion protein has improved acid decarboxylase activity in vitro as measured by the production of polyamines at elevated temperature and/or alkaline pH, relative to a counterpart fusion protein lacking the prion subunit. In some embodiments, the prion subunit is 30 amino acids in length, at least 50 amino acids in length, at least 75 amino acids in length or at least 100 amino acids in length, but 1200 amino acids or fewer in length. In some embodiments, the prion subunit comprises an amino acid composition having at least 20% glutamine and/or asparagine residues. In further embodiments, the prion subunit has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to a Sup35, New1, Ure2, or Rnq1 amino acid sequence; or comprises a Sup35, New1, Ure2, or Rnq1 amino acid sequence. In still other embodiments, the prion subunit has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence set forth in SEQ ID NO:3 or SEQ ID NO:4; or comprises the amino acid sequence of SEQ ID NO:3 or SEQ ID NO:4. In other embodiments, the prion subunit has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of the prion subunit region of SEQ ID NO:7, 8, 11, 12, 13, 14, 15, or 19; or comprises the prion subunit region of SEQ ID NO:7, 8, 11, 12, 13, 14, 15, or 19. In some embodiments, the prion subunit is joined to the C-terminus of the acid decarboxylase. In some embodiments, the prion subunit is joined at the carboxyl terminus to a stability fragment, e.g., a BST fragment, .lamda.CI fragment, or RecA fragment, such as a fragments having the amino acid sequence RRFGEASSAF, ASQWPEETFG, or EGVAETNEDF. In some embodiments, the prion subunit is joined at the C-terminal end to a BST fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:15, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:15, excluding the linker region. In some embodiments, the prion subunit is joined at the C-terminal end to a .lamda.CI fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:16, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:16, excluding the linker region. In some embodiments, the prion subunit is joined at the C-terminal end to a RecA fragment and has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to the amino acid sequence of SEQ ID NO:19, excluding the linker region; or comprises the amino acid sequence to SEQ ID NO:19, excluding the linker region. In some embodiments, the acid decarboxylase is a lysine decarboxylase, ornithine decarboxylase, arginine decarboxylase, or glutamate decarboxylase. In still other embodiments, the acid decarboxylase is a CadA, LdcC, AdiA, SpeA, SpeC, SpeF, GadA, or GadB polypeptide. In some embodiments, the acid decarboxylase is a lysine decarboxylase, such as a CadA lysine decarboxylase and the fusion protein has improved lysine decarboxylase activity in vitro as measured by the production of cadaverine at elevated temperature and/or alkaline pH, relative to a counterpart fusion protein lacking the prion subunit. In some embodiments, the fusion protein is immobilized to a solid support.
[0012] In further aspects the invention provides a polynucleotide encoding a fusion protein as described herein and expression vectors that comprise such polynucleotides.
[0013] Other aspects of the invention are further described herein below.
BRIEF DESCRIPTION OF THE DRAWINGS
[0014] FIG. 1. SDS-PAGE results showing the soluble (s) and precipitate (p) fractions of lysed cell cultures from H. avlei transformed with either pCIB128 or pCIB222. The results from two different colonies are shown for each transformant.
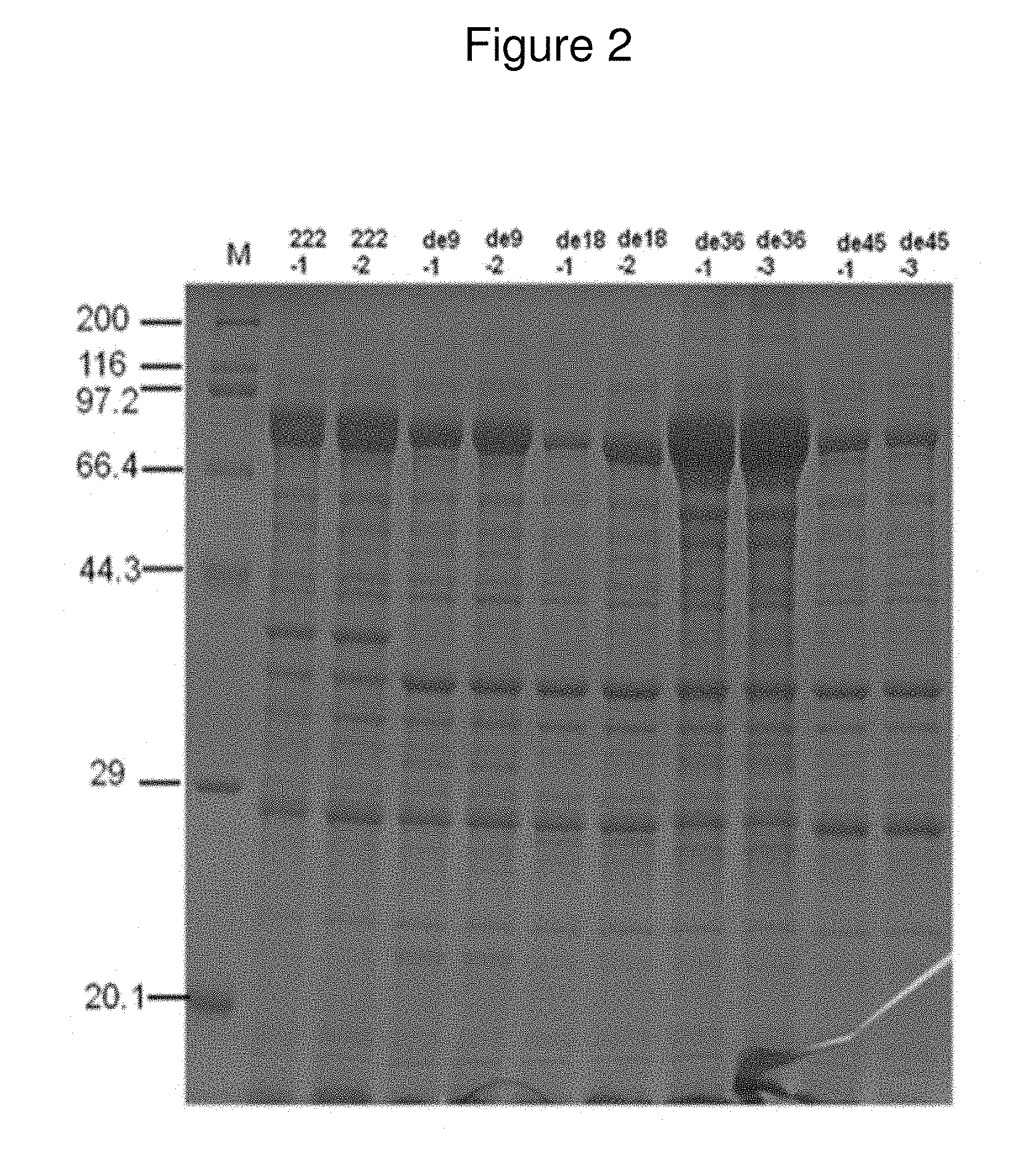
[0015] FIG. 2. SDS-PAGE results showing the total protein of lysed cell cultures from E. coli BL21 transformed with either pCIB222 or one of its truncated variants. The results from two different colonies are shown for each transformant.
DETAILED DESCRIPTION OF ASPECTS OF THE DISCLOSURE
[0016] Before the present invention is described, it is to be understood that this invention is not limited to particular embodiments described, as such may, of course, vary. It is also to be understood that the terminology used herein is for the purpose of describing particular embodiments only, and is not intended to be limiting.
[0017] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Although any methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, the preferred methods and materials are now described. All publications and accession numbers mentioned herein are incorporated herein by reference to disclose and describe the methods and/or materials in connection with which the publications are cited.
Terminology
[0018] A "prion" refers to an infectious agent comprised of protein material that can trigger normal proteins to fold into multiple, structurally distinct conformations (For a review on prions, see Derkatch & Liebman, Prion 1:3, 161-169, 2007). Illustrative examples of yeast prion proteins isolated from S. cerevisiae include Sup35, Ure2, New1, Rnq1, Swi1, Cyc8, Mot3, Spf1 and Mod5. An illustrative example of a filamentous fungal prion protein isolated from Podospora anserine is Het-s. The term "prion variant" as commonly used in the art refers to prion isolates with different properties despite being based on a prion protein with the same sequence. A "prion sequence variant" as used in the present invention refers to a prion amino acid sequence that differs from the amino acid sequence of a native prion amino acid sequence put retains activity of the native prion amino acid sequence when fused to an acid decarboxylase.
[0019] As used herein, the term "prion subunit" refers to a minimal amino acid sequence of a prion protein that is fused to an acid decarboxylase in accordance with the invention and increases the activity of the acid decarboxylase compared to the acid decarboxylase when it is not fused to the prion subunit. The minimal amino acid sequence (and corresponding polynucleotide sequence) comprising the prion subunit can generally be found in the N-terminal region of prion proteins, such as Ure2, Sup35, New1 or Rnq1. In one embodiment, the prion subunit is at least 30 amino acids in length, at least 40 amino acids in length, at least 50 amino acids in length, at least 60 amino acids in length, at least 70 amino acids in length, at least 75 amino acids in length, at least 100 amino acids in length, at least 150 amino acids in length, at least 200 amino acids in length, or at least 300 amino acids in length, or more, but 1200 amino acids or fewer in length. In some embodiments, a prion subunit is 1000 amino acids or fewer in length. In some embodiments, a prion submit is 500 amino acids or fewer in length. In another embodiment, the prion subunit is between 30 and 200 amino acids in length, between 40 and 150 amino acids in length, or between 50 and 120 amino acids in length. A prion subunit when fused to an acid decarboxylase improves activity of the acid decarboxylase in response to stress conditions, such as alkaline pH or elevated temperature compared to the acid decarboxylase that is not fused to the prion subunit.
[0020] As used in the context of the present disclosure, an "acid decarboxylase" refers to a polypeptide that catalyzes the decarboxylation reaction of basic amino acids (e.g., lysine, arginine, ornithine, glutamate) to generate polyamines. Acid decarboxylases include lysine decarboxylases, e.g., CadA, LdcD; arginine decarboxylases, e.g., AdiA; ornithine decarboxylases, e.g., SpeC, SpeF; and glutamate decarboxylases, e.g., GadA, GadB; that are part of the prokaryotic ornithine decarboxylase subclass of Fold Type I pyridoxal 5'-phosphate (PLP)-dependent decarboxylases. This class of proteins typically contains a N-terminal wing domain, a core domain, and a C-terminal domain. The core domain contains a PLP-binding subdomain. The acid decarboxylase SpeA is also a PLP-dependent decarboxylase, but belongs to a different fold family of the PLP-dependent decarboxylases that contain a TIM barrel domain, .beta.-sandwich, insert, and C-terminal domain (Forouhar, et al., Acta. Cryst. F66, 1562-1566, 2010). Acid decarboxylase monomers may form multimers of various sizes, depending on the acid decarboxylase. For example, the acid decarboyxlases CadA, LdcC, and AdiA form a two-fold symmetric dimer that completes the active site of each monomer. Five dimers associate to form a decamer having a double-ringed structure with five-fold symmetry. The decamerscan associate with other decamers to form higher-order oligomers under favorable pH conditions. Not all acid decarboxylases form decamers to function. For example, the acid decarboxylases GadA and GadB form hexamers, and SpeA forms tetramers. According to Kanjee et al., 2011 the acid decarboxylases CadA, LdcC, AdiA, SpeC, and SpeF share the same structural fold and exist at minimum as homodimers. Crystal structure analysis indicates that GadA and GadB also share the same Type I fold of PLP-dependent enzymes, such as CadA, LdcC, AdiA, SpeC, and SpeF (Capitani, et al., The EMBO Journal 22, 4027-4037, 2003). Similarity between LdcC decamer and CadA decamer is described in Kandia, et al., Sci. Rep. 6, 24601, 2016. AdiA decamer formation is described in Boeker E A & Snell E E, J. Biol. Chem. 243, 1678-1684, 1968 and Andrell, et al., Biochemistry 48, 3915-3927, 2009. A structural description of GadA and GadB is described in Capitani, et al., The EMBO Journal 22, 4027-4037, 2003. The protein data bank IDs for structures of illustrative acid decarboxylases are: 3N75 (CadA), 5FKZ (LdcCd), and 2VYC (AdiA). Other E. coli acid decarboxylases such as SpeC and SpeF form homodimers. SpeA forms homotetramer (PDB ID: 3NZQ), while GadA (PDB ID: 1XEY) and GadB (PDB ID: 1PMM) form homohexamers.
[0021] The term "acid decarboxylase" encompasses biologically active variants, alleles, mutants, and interspecies homologs to the specific polypeptides described herein. A nucleic acid that encodes an acid decarboxylase refers to a gene, pre-mRNA, mRNA, and the like, including nucleic acids encoding variants, alleles, mutants, and interspecies homologs of the particular amino acid sequences described herein.
[0022] An "acid decarboxylase fusion polypeptide" as used herein refers to a polypeptide comprising an acid decarboxylase fused to a prion subunit. An "acid decarboxylase fusion polynucleotide" or "acid decarboxylase fusion gene" refers to a nucleic acid that encodes an acid decarboxylase fusion polypeptide.
[0023] A lysine decarboxylase refers to an enzyme that converts L-lysine into cadaverine. The enzyme is classified as E.C. 4.1.1.18. Lysine decarboxylase polypeptides are well characterized enzymes, the structures of which are well known in the art (see, e.g., Kanjee, et al., EMBO J. 30: 931-944, 2011; and a review by Lemmonier & Lane, Microbiology 144; 751-760, 1998; and references described therein). Illustrative lysine decarboxylase sequences are CadA homologs from Klebsiella sp., WP 012968785.1; Enterobacter aerogenes, YP 004592843.1; Salmonella enterica, WP 020936842.1; Serratia sp., WP 033635725.1; and Raoultella ornithinolytica, YP 007874766.1; and LdcC homologs from Shigella sp., WP 001020968.1; Citrobacter sp., WP 016151770.1; and Salmonella enterica, WP 001021062.1. As used herein, a lysine decarboxylase includes variants of native lysine decarboxylase enzymes that have lysine decarboxylase enzymatic activity. Additional lysine decarboxylase enzyme are described in PCT/CN2014/080873 and PCT/CN2015/072978.
[0024] A "cadA" polypeptide refers to an Escherichia coli cadA polypeptide having the amino acid sequence of SEQ ID NO:2, or a biologically active variant thereof that has acid decarboxylase activity. Biologically active variants include alleles, mutants, and interspecies homologs of the E. coli cadA polypeptide. CadA contains an N-terminal wind domain, a core domain, and a C-terminal domain. Illustrative cadA polypeptides from other species include Salmonella enterica, protein sequence accession number WP 001021062.1. In some embodiments, a "CadA" polypeptide has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, 100, or more, amino acids, or over the length of the cadA polypeptide of SEQ ID NO:2. A "CadA polynucleotide" as used herein refers to a polynucleotide that encodes a CadA polypeptide.
[0025] As used herein, the term "alkaline pH" refers to a solution or surrounding environment having a pH of greater than 7.5. In one embodiment, alkaline pH refers to a solution or surrounding environment having a pH of at least 8.0 or at least 8.5, or higher.
[0026] As used herein, the term "elevated temperature" or "high temperature" refers to a temperature about 35.degree. C. or greater. In some embodiments, a higher temperature is at least 37.degree. C., at least 40.degree. C., at least 42.degree. C., at least 45.degree. C., at least 48.degree. C., at least 50.degree. C., at least 52.degree. C., at least 55.degree. C., or greater. In one embodiment, elevated temperature refers to a temperature of at least 42.degree. C. but less than 60.degree. C.
[0027] The term "enhanced" or "improved" in the context of the production of an amino acid, e.g., lysine, or a lysine derivative, e.g., cadaverine, as used herein refers to an increase in the production of an amino acid or the amino acid derivative produced by a host cell that expresses an acid decarboxylase fusion polypeptide comprising an acid decarboxylase polypeptide fused to a prion subunit, e.g., at the carboxyl end of the acid decarboxylase polypeptide, in comparison to a control counterpart cell, such as a cell of the wildtype strain or a cell of the same strain that expresses the acid decarboxylase protein, but is not fused to the prion subunit. In one embodiment, acid decarboxylase activity of the acid decarboxylase fusion protein, e.g., where the prion subunit is fused to the carboxyl end of the acid decarboxylase, is improved by at least 5%, typically at least 10%, 15% 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or greater compared to the acid decarboxylase activity of a counterpart cell expressing an acid decarboxylase lacking the prion subunit, where activity is assessed by measuring the production of polyamines, such as cadaverine, and lysine produced by the host cell and control cell under identical conditions. In some embodiments, activity of a lysed extract from a host cell culture is measured at an elevated temperature. In some embodiments, activity of a lysed extract from a host cell culture is measure under alkaline conditions. For example, activity of an acid decarboxylase fusion polypeptide of the invention can be assessed by evaluating an aliquot of a culture of host cells transformed with the acid decarboxylase fusion polypeptide compared to a corresponding aliquot from a culture of counterpart host cells of the same strain that expresses the acid decarboxylase without fusion to the prion subunit. By way of illustration, the activity of a lysine decarboxylase fusion polypeptide of the invention compared to the counterpart lysine decarboxylase not fused to the prion subunit can be determined by evaluating the reaction rates of a lysed sample, e.g., from a 100 ml sample, at a pH of 8.0. Reaction rates can be measured using NMR by sampling the amount of lysine converted in the presence of PLP into cadaverine about every 1.5 minutes for a total of 20 minutes, and taking the slope of the linear portion of the yield curve. The samples are diluted so that the reaction rate per volume (U) of lysed sample measured at pH 6.0 and 35.degree. C. is the same. The kinetic constants Vmax and Km for lysine of each lysed samples is measured using the same U at an initial pH of 8. By normalizing for U, the concentration of active enzyme in each sample is the same.
[0028] The terms "numbered with reference to", or "corresponding to," or "determined with reference to" when used in the context of the numbering of a given amino acid or polynucleotide sequence, refers to the numbering of the residues of a specified reference sequence when the given amino acid or polynucleotide sequence is compared to the reference sequence. For example, a segment of a prion subunit polypeptide sequence "corresponds to" a segment in SEQ ID NO:4 when the segment aligns with SEQ ID NO:4 in a maximal alignment.
[0029] The terms "polynucleotide" and "nucleic acid" are used interchangeably and refer to a single or double-stranded polymer of deoxyribonucleotide or ribonucleotide bases read from the 5' to the 3' end. A nucleic acid as used in the present invention will generally contain phosphodiester bonds, although in some cases, nucleic acid analogs may be used that may have alternate backbones, comprising, e.g., phosphoramidate, phosphorothioate, phosphorodithioate, or O-methylphosphoroamidite linkages (see Eckstein, Oligonucleotides and Analogues: A Practical Approach, Oxford University Press); positive backbones; non-ionic backbones, and non-ribose backbones. Nucleic acids or polynucleotides may also include modified nucleotides that permit correct read-through by a polymerase. "Polynucleotide sequence" or "nucleic acid sequence" includes both the sense and antisense strands of a nucleic acid as either individual single strands or in a duplex. As will be appreciated by those in the art, the depiction of a single strand also defines the sequence of the complementary strand; thus the sequences described herein also provide the complement of the sequence. Unless otherwise indicated, a particular nucleic acid sequence also implicitly encompasses variants thereof (e.g., degenerate codon substitutions) and complementary sequences, as well as the sequence explicitly indicated. The nucleic acid may be DNA, both genomic and cDNA, RNA or a hybrid, where the nucleic acid may contain combinations of deoxyribo- and ribo-nucleotides, and combinations of bases, including uracil, adenine, thymine, cytosine, guanine, inosine, xanthine hypoxanthine, isocytosine, isoguanine, etc.
[0030] The term "substantially identical," used in the context of two nucleic acids or polypeptides, refers to a sequence that has at least 40%, 45%, or 50% sequence identity with a reference sequence. Percent identity can be any integer from 50% to 100%. Some embodiments include at least: 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99%, compared to a reference sequence using the programs described herein; preferably BLAST using standard parameters, as described below.
[0031] Two nucleic acid sequences or polypeptide sequences are said to be "identical" if the sequence of nucleotides or amino acid residues, respectively, in the two sequences is the same when aligned for maximum correspondence as described below. The terms "identical" or percent "identity," in the context of two or more nucleic acids or polypeptide sequences, refer to two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues or nucleotides that are the same, when compared and aligned for maximum correspondence over a comparison window, as measured using one of the following sequence comparison algorithms or by manual alignment and visual inspection. When percentage of sequence identity is used in reference to proteins or peptides, it is recognized that residue positions that are not identical often differ by conservative amino acid substitutions, where amino acids residues are substituted for other amino acid residues with similar chemical properties (e.g., charge or hydrophobicity) and therefore do not change the functional properties of the molecule. Where sequences differ in conservative substitutions, the percent sequence identity may be adjusted upwards to correct for the conservative nature of the substitution. Means for making this adjustment are well known to those of skill in the art. Typically this involves scoring a conservative substitution as a partial rather than a full mismatch, thereby increasing the percentage sequence identity. Thus, for example, where an identical amino acid is given a score of 1 and a non-conservative substitution is given a score of zero, a conservative substitution is given a score between zero and 1.
[0032] For sequence comparison, typically one sequence acts as a reference sequence, to which test sequences are compared. When using a sequence comparison algorithm, test and reference sequences are entered into a computer, subsequence coordinates are designated, if necessary, and sequence algorithm program parameters are designated. Default program parameters can be used, or alternative parameters can be designated. The sequence comparison algorithm then calculates the percent sequence identities for the test sequences relative to the reference sequence, based on the program parameters.
[0033] An algorithm that may be used to determine whether an acid decarboxylase fusion polypeptide has sequence identity to a sequence, e.g., SEQ ID NO:2; or any one of SEQ ID NOS:21-28, or another polypeptide reference sequence, is the BLAST algorithm, which is described in Altschul et al., 1990, J. Mol. Biol. 215:403-410, which is incorporated herein by reference. Software for performing BLAST analyses is publicly available through the National Center for Biotechnology Information (on the worldwide web at ncbi.nlm.nih.gov/). For amino acid sequences, the BLASTP program uses as defaults a word size (W) of 3, an expectation (E) of 10, and the BLOSUM62 scoring matrix (see Henikoff & Henikoff, 1989, Proc. Natl. Acad. Sci. USA 89:10915). Other programs that may be used include the Needleman-Wunsch procedure, J. MoI. Biol. 48: 443-453 (1970), using BLOSUM62, a Gap start penalty of 7 and gap extend penalty of 1; and gapped BLAST 2.0 (see Altschul, et al. 1997, Nucleic Acids Res., 25:3389-3402) both
[0034] A "comparison window," as used herein, includes reference to a segment of any one of the number of contiguous positions selected from the group consisting of from 20 to 600, usually about 50 to about 200, more usually about 100 to about 150 in which a sequence may be compared to a reference sequence of the same number of contiguous positions after the two sequences are optimally aligned. Methods of alignment of sequences for comparison are well-known in the art. Optimal alignment of sequences for comparison can be conducted, e.g., by the local homology algorithm of Smith & Waterman, Adv. Appl. Math. 2:482 (1981), by the homology alignment algorithm of Needleman & Wunsch, J. Mol. Biol. 48:443 (1970), by the search for similarity method of Pearson & Lipman, Proc. Nat'l. Acad. Sci. USA 85:2444 (1988), by computerized implementations of these algorithms (GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package, Genetics Computer Group, 575 Science Dr., Madison, Wis.), or by manual alignment and visual inspection.
[0035] Nucleic acid or protein sequences that are substantially identical to a reference sequence include "conservatively modified variants." With respect to particular nucleic acid sequences, conservatively modified variants refers to those nucleic acids which encode identical or essentially identical amino acid sequences, or where the nucleic acid does not encode an amino acid sequence, to essentially identical sequences. Because of the degeneracy of the genetic code, a large number of functionally identical nucleic acids encode any given protein. For instance, the codons GCA, GCC, GCG and GCU all encode the amino acid alanine. Thus, at every position where an alanine is specified by a codon, the codon can be altered to any of the corresponding codons described without altering the encoded polypeptide. Such nucleic acid variations are "silent variations," which are one species of conservatively modified variations. Every nucleic acid sequence herein which encodes a polypeptide also describes every possible silent variation of the nucleic acid. One of skill will recognize that each codon in a nucleic acid (except AUG, which is ordinarily the only codon for methionine) can be modified to yield a functionally identical molecule. Accordingly, each silent variation of a nucleic acid which encodes a polypeptide is implicit in each described sequence.
[0036] The term "polypeptide" as used herein includes reference to polypeptides containing naturally occurring amino acids and amino acid backbones as well as non-naturally occurring amino acids and amino acid analogs.
[0037] As to amino acid sequences, one of skill will recognize that individual substitutions, in a nucleic acid, peptide, polypeptide, or protein sequence which alters a single amino acid or a small percentage of amino acids in the encoded sequence is a "conservatively modified variant" where the alteration results in the substitution of an amino acid with a chemically similar amino acid. Conservative substitution tables providing functionally similar amino acids are well known in the art. Examples of amino acid groups defined in this manner can include: a "charged/polar group" including Glu (Glutamic acid or E), Asp (Aspartic acid or D), Asn (Asparagine or N), Gln (Glutamine or Q), Lys (Lysine or K), Arg (Arginine or R) and His (Histidine or H); an "aromatic or cyclic group" including Pro (Proline or P), Phe (Phenylalanine or F), Tyr (Tyrosine or Y) and Trp (Tryptophan or W); and an "aliphatic group" including Gly (Glycine or G), Ala (Alanine or A), Val (Valine or V), Leu (Leucine or L), Ile (Isoleucine or I), Met (Methionine or M), Ser (Serine or S), Thr (Threonine or T) and Cys (Cysteine or C). Within each group, subgroups can also be identified. For example, the group of charged/polar amino acids can be sub-divided into sub-groups including: the "positively-charged sub-group" comprising Lys, Arg and His; the "negatively-charged sub-group" comprising Glu and Asp; and the "polar sub-group" comprising Asn and Gln. In another example, the aromatic or cyclic group can be sub-divided into sub-groups including: the "nitrogen ring sub-group" comprising Pro, His and Trp; and the "phenyl sub-group" comprising Phe and Tyr. In another further example, the aliphatic group can be sub-divided into sub-groups including: the "large aliphatic non-polar sub-group" comprising Val, Leu and Ile; the "aliphatic slightly-polar sub-group" comprising Met, Ser, Thr and Cys; and the "small-residue sub-group" comprising Gly and Ala. Examples of conservative mutations include amino acid substitutions of amino acids within the sub-groups above, such as, but not limited to: Lys for Arg or vice versa, such that a positive charge can be maintained; Glu for Asp or vice versa, such that a negative charge can be maintained; Ser for Thr or vice versa, such that a free --OH can be maintained; and Gln for Asn or vice versa, such that a free --NH2 can be maintained. The following six groups each contain amino acids that further provide illustrative conservative substitutions for one another. 1) Ala, Ser, Thr; 2) Asp, Glu; 3) Asn, Gln; 4) Arg, Lys; 5) Ile, Leu, Met, Val; and 6) Phe, Try, and Trp (see, e.g., Creighton, Proteins (1984)).
[0038] The term "promoter," as used herein, refers to a polynucleotide sequence capable of driving transcription of a DNA sequence in a cell. Thus, promoters used in the polynucleotide constructs of the invention include cis- and trans-acting transcriptional control elements and regulatory sequences that are involved in regulating or modulating the timing and/or rate of transcription of a gene. For example, a promoter can be a cis-acting transcriptional control element, including an enhancer, a repressor binding sequence and the like. These cis-acting sequences typically interact with proteins or other biomolecules to carry out (turn on/off, regulate, modulate, etc.) gene transcription. Most often the core promoter sequences lie within 1-2 kb of the translation start site, more often within 1 kbp and often within 500 bp or 200 bp or fewer, of the translation start site. By convention, promoter sequences are usually provided as the sequence on the coding strand of the gene it controls. In the context of this application, a promoter is typically referred to by the name of the gene for which it naturally regulates expression. A promoter used in an expression construct of the invention is referred to by the name of the gene. Reference to a promoter by name includes a wild type, native promoter as well as variants of the promoter that retain the ability to induce expression. Reference to a promoter by name is not restricted to a particular species, but also encompasses a promoter from a corresponding gene in other species.
[0039] A "constitutive promoter" in the context of this invention refers to a promoter that is capable of initiating transcription under most conditions in a cell, e.g., in the absence of an inducing molecule. An "inducible promoter" initiates transcription in the presence of an inducer molecule.
[0040] A polynucleotide is "heterologous" to an organism or a second polynucleotide sequence if it originates from a foreign species, or, if from the same species, is modified from its original form. For example, when a polynucleotide encoding a polypeptide sequence is said to be operably linked to a heterologous promoter, it means that the polynucleotide coding sequence encoding the polypeptide is derived from one species whereas the promoter sequence is derived from another, different species; or, if both are derived from the same species, the coding sequence is not naturally associated with the promoter (e.g., is a genetically engineered coding sequence, e.g., from a different gene in the same species, or an allele from a different ecotype or variety). Similarly, a polypeptide is "heterologous" to a host cell if the native wildtype host cell does not produce the polypeptide.
[0041] The term "exogenous" refers generally to a polynucleotide sequence or polypeptide that does not naturally occur in a wild-type cell or organism, but is typically introduced into the cell by molecular biological techniques, i.e., engineering to produce a recombinant microorganism. Examples of "exogenous" polynucleotides include vectors, plasmids, and/or man-made nucleic acid constructs encoding a desired protein or enzyme.
[0042] The term "endogenous" refers to naturally-occurring polynucleotide sequences or polypeptides that may be found in a given wild-type cell or organism. In this regard, it is also noted that even though an organism may comprise an endogenous copy of a given polynucleotide sequence or gene, the introduction of a plasmid or vector encoding that sequence, such as to over-express or otherwise regulate the expression of the encoded protein, represents an "exogenous" copy of that gene or polynucleotide sequence. Any of the pathways, genes, or enzymes described herein may utilize or rely on an "endogenous" sequence, which may be provided as one or more "exogenous" polynucleotide sequences, or both.
[0043] "Recombinant nucleic acid" or "recombinant polynucleotide" as used herein refers to a polymer of nucleic acids wherein at least one of the following is true: (a) the sequence of nucleic acids is foreign to (i.e., not naturally found in) a given host cell; (b) the sequence may be naturally found in a given host cell, but in an unnatural (e.g., greater than expected) amount; or (c) the sequence of nucleic acids comprises two or more subsequences that are not found in the same relationship to each other in nature. For example, regarding instance (c), a recombinant nucleic acid sequence will have two or more sequences from unrelated genes arranged to make a new functional nucleic acid.
[0044] The term "operably linked" refers to a functional relationship between two or more polynucleotide (e.g., DNA) segments. Typically, it refers to the functional relationship of a transcriptional regulatory sequence to a transcribed sequence. For example, a promoter or enhancer sequence is operably linked to a DNA or RNA sequence if it stimulates or modulates the transcription of the DNA or RNA sequence in an appropriate host cell or other expression system. Generally, promoter transcriptional regulatory sequences that are operably linked to a transcribed sequence are physically contiguous to the transcribed sequence, i.e., they are cis-acting. However, some transcriptional regulatory sequences, such as enhancers, need not be physically contiguous or located in close proximity to the coding sequences whose transcription they enhance.
[0045] The term "expression cassette" or "DNA construct" or "expression construct" refers to a nucleic acid construct that, when introduced into a host cell, results in transcription and/or translation of an RNA or polypeptide, respectively. In the case of expression of transgenes, one of skill will recognize that the inserted polynucleotide sequence need not be identical, but may be only substantially identical to a sequence of the gene from which it was derived. As explained herein, these substantially identical variants are specifically covered by reference to a specific nucleic acid sequence. One example of an expression cassette is a polynucleotide construct that comprises a polynucleotide sequence encoding a polypeptide of the invention protein operably linked to a promoter, e.g., its native promoter, where the expression cassette is introduced into a heterologous microorganism. In some embodiments, an expression cassette comprises a polynucleotide sequence encoding a polypeptide of the invention where the polynucleotide that is targeted to a position in the genome of a microorganism such that expression of the polynucleotide sequence is driven by a promoter that is present in the microorganism.
[0046] The term "host cell" as used in the context of this invention refers to a microorganism and includes an individual cell or cell culture that can be or has been a recipient of any recombinant vector(s) or isolated polynucleotide(s) of the invention. Host cells include progeny of a single host cell, and the progeny may not necessarily be completely identical (in morphology or in total DNA complement) to the original parent cell due to natural, accidental, or deliberate mutation and/or change. A host cell includes cells into which a recombinant vector or a polynucleotide of the invention has been introduced, including by transformation, transfection, and the like.
[0047] The term "isolated" refers to a material that is substantially or essentially free from components that normally accompany it in its native state. For example, an "isolated polynucleotide," as used herein, may refer to a polynucleotide that has been isolated from the sequences that flank it in its naturally-occurring or genomic state, e.g., a DNA fragment that has been removed from the sequences that are normally adjacent to the fragment, such as by cloning into a vector. A polynucleotide is considered to be isolated if, for example, it is cloned into a vector that is not a part of the natural environment, or if it is artificially introduced in the genome of a cell in a manner that differs from its naturally-occurring state. Alternatively, an "isolated peptide" or an "isolated polypeptide" and the like, as used herein, refers to a polypeptide molecule that is free of other components of the cell, i.e., it is not associated with in vivo substances.
[0048] The invention employs various routine recombinant nucleic acid techniques. Generally, the nomenclature and the laboratory procedures in recombinant DNA technology described below are commonly employed in the art. Many manuals that provide direction for performing recombinant DNA manipulations are available, e.g., Sambrook & Russell, Molecular Cloning, A Laboratory Manual (3rd Ed, 2001); and Current Protocols in Molecular Biology (Ausubel, et al., John Wiley and Sons, New York, 2009-2016).
SUMMARY OF CERTAIN ASPECTS OF THE DISCLOSURE
[0049] The present disclosure is based, in part, on the discovery that fusing a prion subunit to an acid decarboxylase, e.g., at the carboxyl terminus, to produce an acid decarboxylase fusion protein and expressing the fusion protein in a host cell improves the activity of the acid decarboxylase fusion protein as measured by the production of polyamines under various stress conditions, such as alkaline pH or elevated temperature, as compared to the expression of an acid decarboxylase lacking the prion subunit.
[0050] Additionally, the present disclosure is based, in part, on the discovery that fusion of a prion subunit to an acid decarboxylase, e.g., at the carboxyl terminus, to produce a fusion protein increases the stability of the acid decarboxylase as measured by oligomerization of the fusion protein at elevated temperature or alkaline pH when compared to a counterpart acid decarboxylase lacking the prion subunit.
[0051] Further, the present disclosure is based, in part, on the discovery that fusion of a prion subunit to an acid decarboxylase, e.g., at the carboxyl terminus, to produce a fusion protein increases the solubility of the acid decarboxylase at elevated temperature or alkaline pH. Accordingly, fusion proteins of the present disclosure typically have improved solubility relative to an acid decarboxylase protein lacking the corresponding prion subunit as measured by acid decarboxylase activity at an alkaline pH.
[0052] The ability of an acid decarboxylase fusion protein of the present invention to tolerate alkaline pH also allows the use of alternative nitrogen sources that have higher pH values, such as urea and ammonia (1M solution has a pH 11.6) in fermentation reactions to generate the desired product, e.g., polyamines. These alternative nitrogen sources generate less salt waste byproduct.
Prion Subunit
[0053] Prions are self-propagating and transmissible protein isoforms. A normal cellular protein (PrPc) having an altered confirmation can be infectious, resulting in disease. While there is no protein with homology to PrPc in yeast, several yeast prion proteins have been identified.
[0054] The first prions identified in yeast, [PSI+] and [URE3], were determined to be prion forms of the Ure2 and Sup35 proteins, respectively. Since that time, other yeast prions have been identified in Saccharomyces cerevisiae including [PIN+]/[RNQ+], [SWI+], [OCT+], [MOT+], [ISP+], [BETA], [MOD+] and a fungal prion, [Het-s], identified in Podospora anserina (see, Wickner et al., Microbiology and Molecular Biology Reviews, 2015).
[0055] In some embodiments, a prion subunit can defined as a prion polypeptide or fragment thereof where the percent composition of asparagine (N) and glutamine (Q) is 10% or greater. For example, the percent of Q/N in Sup35 prion subunit is 25% of 154 amino acids, and that of New1 is 27% of 253 amino acids. The 10% is determined with reference to the portion of the fused polypeptide that is considered to have prion activity, i.e., is determined considered in the context of the prion subunit sequence only.
[0056] Prion polypeptide sequences suitable for use in the invention as a prion subunit include amino acid sequences of a prion polypeptide as illustrated in SEQ ID NO:3 or 4, or substantially identical sequence variants thereof. Such a sequence variant typically has at least 50%, or at least 60%, 70%, 75%, 80%, 85%, or 90% identity to one of SEQ ID NOS: 3 or 4, or e.g., a homolog of SEQ ID NO: 3 or 4. In some embodiments, a prion subunit comprises the amino acid sequence of the prion region of SEQ ID NO:7, 8, 11, 12, 13, or 14; or has at least 50%, or has at least 60%, 70%, 75%, 80%, 85%, or 90% identity to the prion region of SEQ ID NO:7, 8, 11, 12, 13, or 14. As used herein, the term "sequence variant" encompasses biologically active polypeptides having one or more substitutions, deletions, or insertions relative to a prion polypeptide reference sequence, such as SEQ ID NO: 3, or 4. Thus, the term "sequence variant" includes biologically active fragments as well as substitution variants.
[0057] In one embodiment, prion subunit polypeptide sequences suitable for use in the invention include amino acid sequences encoding Ure2, Sup35, New1, Rnq1, Swi1, Cyc8, Mot3 or Sfp1 or substantially identical variants thereof. Illustrative examples of Ure2 polypeptides include those from S. cerevisiae protein sequence accession number AAM93184; Candida albicans protein sequence accession number AAM91946; S. bayanus protein sequence accession number AAM91939; and Eremothecium gossypii protein sequence accession number AAM91943. Illustrative examples of Sup35 polypeptides include those from S. cerevisiae protein sequence accession number AJV18122; S. boulardii protein sequence accession number KOH51638; and S. bayanus protein sequence accession number AAL15027. Illustrative examples of New1 polypeptides include those from S. cerevisiae protein sequence accession number AHY77957; S. boulardii protein sequence accession number KOH47591; and Sugiyamella lignohabitans protein sequence accession number ANB11767. Illustrative examples of Rnq1 polypeptides include those from S. cerevisiae protein sequence accession number AFU61310; and S. boulardii protein sequence accession number KOH52602. Illustrative examples of Swi1 polypeptides include those from S. cerevisiae protein sequence accession number AJP42124; C. albicans protein sequence accession number AOW28823; and Sugiyamella lignohabitans protein sequence accession number ANB13699. Illustrative examples of Cyc8 polypeptides include those from S. cerevisiae protein sequence accession number CAA85069; and Aspergillus nomius protein sequence accession number KNG81485. Illustrative examples of Mot3 polypeptides include those from S. cerevisiae protein sequence accession number AAC49982; and S. boulardii protein sequence accession number KQC41827. Illustrative examples of Sfp1 polypeptides include those from S. cerevisiae protein sequence accession number AAB82343; S. boulardii protein sequence accession number KOH49283; and S. arboricola protein sequence accession number EJS42621. In one embodiment, polypeptide sequences suitable for use in the invention include amino acid sequences encoding a prion polypeptide that are capable of inducing protein oligomerization in vivo or in vitro.
[0058] In some embodiments, polynucleotide sequences suitable for use in the invention include nucleic acid sequences encoding one or more of the following prion proteins: Ure2, Sup35, New1, Rnq1, Swi1, Cyc8, Mot3 or Sfp1 or homologs thereof. Moreover, suitable polynucleotides for use in the invention include nucleic acid sequences that encode any one of the illustrative prion polypeptides provided herein. In another embodiment, suitable polynucleotides include nucleic acid sequences that encode any one or more of the illustrative prion polypeptides disclosed herein that are capable of inducing protein oligomerization in vivo or in vitro. In one embodiment, polynucleotide sequences suitable for use in the invention include nucleic acid sequences that encode a prion polypeptide as illustrated in SEQ NOs: 5 or 6, or substantially identical variants thereof. Such a variant typically has at least 60%, or at least 70%, 75%, 80%, 85%, or 90% identity to one of SEQ ID NOS: 5 or 6.
[0059] In one embodiment, the invention relates to a genetically modified host cell having a nucleic acid sequence encoding an acid decarboxylase fusion protein, where the acid decarboxylase fusion protein comprises or consists of an acid decarboxylase polypeptide joined to a prion subunit, e.g., at the carboxyl terminus of the acid decarboxylase polypeptide, and where the fusion protein has improved acid decarboxylase activity as measured by the production of polyamines relative to a counterpart host cell that expresses the acid decarboxylase polypeptide not joined to the prion subunit. In one embodiment, the prion subunit is at least 30 amino acids in length, at least 50 amino acids in length, at least 75 amino acids in length or at least 100 amino acids in length, but 1200 amino acids or fewer in length. In some embodiments, a prion subunit comprises an amino acid composition having at least 20% Q or N residues. In another embodiment, the prion subunit has at least 70%, 75%, 80%, 85%, 90%, or 95% identity to a Sup35, New1, Ure2, or Rnq1 amino acid sequence; or comprises a Sup35, New1, Ure2, or Rnq1 amino acid sequence.
Structural Organization of Prions
[0060] Generally, yeast prions are intrinsically disordered in solution and QN-rich. Scrambled PrD's of Sup35 and Ure2 maintaining amino acid composition but not exact sequence, were found to be capable of both generating amyloid in vitro and prions in vivo, and of propagating the prion state, indicating that the amino acid composition plays an important role in prion properties (Ross et al., 2005). Thus, variants of yeast prion sequences of use in the invention as prion subunits include fragments of Sup35 or Ure2 that have less than 50% sequence identity to Sup35 and Ure2, but have a QN composition of 10% or greater.
Sup35
[0061] Yeast protein Sup35 (685 aa) is a subunit of the translation termination factor and terminates translation at stop codons, and residues 254-685 (Sup35C) have been observed to be sufficient to carry out the essential translation termination function. Residues 1-253 (Sup35NM) were observed to regulate general mRNA turnover through interactions with the poly(A) --binding protein and the poly(A)-degrading enzyme. Residues 1-114 (Sup35N) are sufficient to propagate the original [PSI+] variant, while residues 1-61 are sufficient to propagate several variants of this prion (Chang et al., PNAS, 2008). The N-proximal PrD region of Sup35 includes an N-terminal QN-rich region located within the first 40 amino acids, and a region of 5.5 imperfect oligopeptide repeats (ORs) located at positions 41 and 97. The PrD fragment required for aggregation is shorter than the fragment needed for propagation of the prion state and is primarily confined to the QN-rich region (Osherovich et al, 2004).
[0062] Parts of the Sup35M domain (residues 115-253), up to residue 137, were observed as necessary for propagation of some strong and weak [PSI+] variants, and deletions and substitutions within the M domain were observed to alter the character of [PSI+] variant significantly (Liu et al., PNAS, 2002). Solid-state nuclear magnetic resonance (ss-NMR) experiments with Sup35NM filaments showed that Tyrosine (Tyr) residues, all of which are within the N terminal, are in an in-register parallel .beta.-sheet structure (Shewmaker, PNAS, 2006). Additionally, it was observed that there are eight leucine (Leu) residues, i.e., residues 110, 126, 144, 146, 154, 212, 218 and 238. The ss-NMR data suggests that four of these Leu residues are in an in-register parallel structure (Shewmaker et al., Biochemistry, 2009).
Ure2
[0063] Yeast protein Ure2 (354aa) acts to regulate nitrogen catabolism. The part of Ure2 whose overproduction induces the formation of [URE3] was found to be the N-terminal 65 residues and this region proved to be sufficient to propagate [URE3] in the absence of the remainder of the molecule (Masison et al., Science, 1997). The N-terminal prion domain normally functions to stabilize Ure2 against degradation (Shewmaker et al., Genetics, 2007).
Rnq1
[0064] In the case of the yeast protein Rnq1 (405 aa), four QN-rich regions were found within the PrD (Kadnar et al., 2010). While none were essential for prion propagation, two of the four stretches were each found to support prion maintenance if retained alone.
New1
[0065] Yeast protein New1 (1196 aa) consists of a N-terminal prion region (New1N) and a C-terminal region homologous to a translation elongation factor with two ATP-binding motifs.
[0066] Generally, a prion capable of inducing protein aggregation requires a glutamine (Q) and/or asparagine (N) or (NQ)-rich region. For example, the prion protein, New1, contains the sequence "QQQRNWKQGGNYQQYQSYN" and "SNYNNYNNYNNYNNYNNYNNYNKYNGQGYQ". In the prion protein Sup35, the N-terminus contains a NQ-rich region followed by the N domain repeat (NR) region, which contains five complete copies (R1-R5) and one partial copy (R6) of the imperfect oligopeptide repeating sequence "PQGGYQQN".
[0067] In one embodiment, the prion subunit can comprise or consist of a nucleic acid encoding a prion protein selected from the group consisting of Ure2, Sup35, New1, Rnq1, Swi1, Cyc8, Mot3 and Sfp1. In another embodiment, the prion subunit can comprise or consist of a nucleic acid encoding a polypeptide derived from Ure2, Sup35, New1, Rnq1, Swi1, Cyc8, Mot3 or Sfp1 capable of inducing protein oligomerization in vivo or in vitro. In yet another embodiment, the prion subunit can consist of or comprise an amino acid sequence of at least 30 amino acids, at least 40 amino acids, at least 50 amino acids, at least 60 amino acids, at least 70 amino acids, at least 80 amino acids, at least 90 amino acids, at least 100 amino acids, at least 200 amino acids, at least 300 amino acids, but less than 1200 amino acids that when fused to the carboxyl end of an acid decarboxylase improves decarboxylation by the acid decarboxylase under alkaline pH and/or elevated temperature as compared to a counterpart acid decarboxylase lacking the prion subunit under the same conditions. In one embodiment, decarboxylation is measured by the production of polyamines by the acid decarboxylase.
[0068] In one embodiment, the prion subunit can consist of or comprise an amino acid sequence of at least 30 amino acids, at least 40 amino acids, at least 50 amino acids, at least 60 amino acids, at least 70 amino acids, at least 80 amino acids, at least 90 amino acids, at least 100 amino acids, at least 200 amino acids, at least 300 amino acids, but less than 1200 amino acids that when fused to the carboxyl end of a lysine decarboxylase improves decarboxylation by the lysine decarboxylase under alkaline pH and/or elevated temperature as compared to a counterpart lysine decarboxylase lacking the prion subunit under the same conditions. In one embodiment, decarboxylation is measured by the production of cadaverine by the lysine decarboxylase.
[0069] In one embodiment, the prion subunit can consist of or comprise an amino acid sequence of at least 30 amino acids, at least 40 amino acids, at least 50 amino acids, at least 60 amino acids, at least 70 amino acids, at least 80 amino acids, at least 90 amino acids, at least 100 amino acids, at least 200 amino acids, at least 300 amino acids, but less than 1200 amino acids that when fused to the carboxyl end of a arginine decarboxylase improves decarboxylation by the arginine decarboxylase under alkaline pH and/or elevated temperature as compared to a counterpart arginine decarboxylase lacking the prion subunit under the same conditions. In one embodiment, decarboxylation is measured by the production of putrescine by the arginine decarboxylase.
[0070] In one embodiment, the prion subunit can consist of or comprise an amino acid sequence of at least 30 amino acids, at least 40 amino acids, at least 50 amino acids, at least 60 amino acids, at least 70 amino acids, at least 80 amino acids, at least 90 amino acids, at least 100 amino acids, at least 200 amino acids, at least 300 amino acids, but less than 1200 amino acids that when fused to the carboxyl end of an ornithine decarboxylase improves decarboxylation by the ornithine decarboxylase under alkaline pH and/or elevated temperature as compared to a counterpart ornithine decarboxylase lacking the prion subunit under the same conditions. In one embodiment, decarboxylation is measured by the production of spermine by the ornithine decarboxylase.
[0071] In one embodiment, the prion subunit can consist of or comprise an amino acid sequence of at least 30 amino acids, at least 40 amino acids, at least 50 amino acids, at least 60 amino acids, at least 70 amino acids, at least 80 amino acids, at least 90 amino acids, at least 100 amino acids, at least 200 amino acids, at least 300 amino acids, but less than 1200 amino acids that when fused to the carboxyl end of a glutamate decarboxylase improves decarboxylation by the glutamate decarboxylase under alkaline pH and/or elevated temperature as compared to a counterpart glutamate decarboxylase lacking the prion subunit under the same conditions. In one embodiment, decarboxylation is measured by the production of gamma-aminobutyric acid (GABA) by the glutamate decarboxylase.
[0072] A prion subunit has at least 10% Q and N residues, and typically at least 20%, 30%, 40%, 50% or more Q and N residues. In another embodiment, the prion subunit has at least 10% Q residues, and typically at least 20%, 30%, 40%, 50% or more Q residues. In yet another embodiment, the prion subunit has at least 10% N residues, and typically at least 20%, 30%, 40%, 50% or more N residues. In one embodiment, the prion subunit contains a higher percentage of N residues as compared to Q residues. In another embodiment, the prion subunit contains a higher percentage of Q residues as compared to N residues. In another embodiment, the percentage of Q and N residues present in the fusion protein is such that the prion subunit capable of causing protein oligomerization contains at least 10% Q or N residues.
[0073] In one embodiment, the prion subunit of the fusion protein can comprise or consist of a nucleic acid sequence encoding a prion protein selected from the group consisting of Ure2, Sup35, New1, Rnq1, Swi1, Cyc8, Mot3 or Sfp1. In another embodiment, the prion subunit can comprise or consist of a nucleic acid sequence encoding a polypeptide derived from Ure2, Sup35, New1, Rnq1, Swi1, Cyc8, Mot3 or Sfp1 capable of inducing protein oligomerization in vivo or in vitro. In yet another embodiment, the prion subunit can consist of or comprise an amino acid sequence of at least 30 amino acids, at least 40 amino acids, at least 50 amino acids, at least 60 amino acids, at least 70 amino acids, at least 80 amino acids, at least 90 amino acids, at least 100 amino acids, at least 200 amino acids, at least 300 amino acids, but less than 1200 amino acids that when fused to the carboxyl end of an acid decarboxylase increases decarboxylation by the acid decarboxylase under alkaline pH and/or elevated temperature as compared to a counterpart acid decarboxylase lacking the prion subunit under the same conditions. In one embodiment, decarboxylation is measured by the production of polyamines by the acid decarboxylase.
[0074] In another embodiment, the prion subunit represents the minimally required amino acid sequence (and corresponding polynucleotide sequence) necessary to improve acid decarboxylase activity as measured by the production of polyamines by the fusion protein. In one embodiment, the prion subunit can be at least 30 amino acids in length, or at least 40, 50, 60, 70, 80, 90, 100, 200, 300 or more amino acids, but less than 1200 amino acids in length.
[0075] In one embodiment, a prion subunit can be linked to another short amino acid sequence that confers stability. In one embodiment, the short peptide is selected from the group consisting of a BST fragment, a RecA fragment and a .lamda.CI fragment. In another embodiment, the linker polypeptide can comprise the amino acid sequence RRFGEASSAF, ASQWPEETFG, or EGVAETNEDF.
[0076] In one embodiment, the prion subunit represents the minimally required amino acid sequence (and corresponding polynucleotide sequence) necessary to improve acid decarboxylase activity as measured by the production of polyamines by the fusion protein relative to an acid decarboxylase lacking the prion subunit. In one embodiment, the prion subunit can be at least 30 amino acids in length, or at least 40, 50, 60, 70, 80, 90, 100, 200, 300 or more amino acids, but less than 1200 amino acids in length.
[0077] In one embodiment, the prion subunit is capable of inducing protein aggregation (oligomerization) and the prion subunits forma .beta.-sheet structure, such as an in-register parallel (3-sheet structure. In one embodiment, the prion subunit may be capable of inducing protein aggregation of one or more protein monomers fused to the prion subunit and the prion subunits form into a .beta.-helix structure.
[0078] A prion subunit may be fused to an acid decarboxylase at the N-terminus, the C-terminus of an acid decarboxylase, or may be introduced at a surface region of the acid decarboxylase protein. In certain embodiments, the prion subunit is fused at the C-terminus of the acid decarboxylase. A prion subunit is typically joined to the acid decarboxylase by a linker, such as flexible linker comprising amino acids such as Gly, Ser, Ala, and the like.
Acid Decarboxylases
[0079] Various acid decarboxylase activity have been well characterized, both structurally and functionally. These include CadA, LdcC, AdiA, SpeA, SpeC, SpeF, GadA, GadB, and their homologs. The optimal pH for CadA is between 5 and 6, LdcC is between 7 and 8, AdiA is between 4.5 and 5.5, SpeC is between 7.5 and 8.5, and SpeF is between 7 and 8 (Kanjee et al., Biochemistry 50, 9388-9398, 2011). GadA and GadB are activated when the pH of the environment is between 2 and 2.5 (Castanie-Cornet et al., J. Bacteriol. 181, 3525-3535, 1999). However, the decarboxylation of basic amino acids lysine, arginine, ornithine, and glutamateleads to the production of cadaverine, putrescine, spermine, and GABA; and their formation involves the consumption of protons. These are basic molecules that tend to increase the pH of the medium. For example, the pKa's of cadaverine are 9.1 and 10.2, and that of putrescine are 9.7 and 11.2. Therefore, the production of these basic molecules quickly increases the pH of the reaction medium to a pH that is outside of the optimal pH of the acid decarboxylase.
[0080] Acid decarboxylases that are fused to a prion subunit as described herein include lysine decarboxylases, arginine decarboxylases, ornithine decarboxylases, and glutamate decarboxylases, which as detailed above, are part of the prokaryotic ornithine decarboxylase subclass of Fold Type I pyridoxal 5'-phosphate (PLP)-dependent decarboxylases. This class of proteins typically contains a N-terminal wing domain, a core domain, and a C-terminal domain. The core domain contains the PLP-binding subdomain and subdomain 4. The acid decarboxylase SpeA belongs to a different fold family of the PLP-dependent decarboxylases. These share little sequence identity, but the structures are well known. The Protein Data Bank Identification numbers for illustrative acid decarboxylase structures are CadA: 3N75, LdcC: 5FKZ, AdiA: 2VYC, SpeA: 3NZQ, GadA: 1XEY, and GadB: 1PMM.
[0081] For CadA, the N-terminal wind domain (residues 1 to 129 as determined with reference to SEQ ID NO:2) has a flavodoxin-like fold consisting of five-stranded parallel beta-sheets sandwiched between two sets of amphipathic alpha-helices. The core domain (residues 130 to 563 as determined with reference to 563 of SEQ ID NO:2) includes a linker region (amino acid residues 130 to 183 of SEQ ID NO:2) that form a short helical bundle, the PLP-binding subdomain (amino acids 184 to 417 of SEQ ID NO:2) that form a seven-stranded beta-sheet core surrounded by three sets of alpha-helices, and subdomain 4 consists of amino acid residues 418 to 563 that form a four stranded antiparallel beta-sheet core with three alpha-helices facing outward. The C-terminal domain (corresponding to amino acid residues 564 to 715 as determined with referenced to SEQ ID NO:2) forms two sets of beta sheets with an alpha-helical outer surface (Kanjee et al., The EMBO Journal 30, 931-944 2011).
[0082] The CadA protein forms a two-fold symmetric dimer that completes the active site of each monomer. Five dimers associate to form a decamer that consist of a double-ringed structure with five-fold symmetry. The decamer associates with other decamers to form higher-order oligomers. It has been shown that in acidic conditions (pH 5), CadA predominantly exists in the oligomeric state, and less oligomers and decamers are found as the environment becomes more basic. It was estimated that 25% of the enzymes exist as dimers and 75% exist as decamers at pH 6.5, while 95% of the enzymes exist as dimers at pH 8.0 (Kanjee et al., The EMBO Journal 30, 931-944 2011). This decrease in oligomer formation coincides with the decrease in decarboxylase activity observed as the pH of the environment of the enzyme increases above 5.0, suggesting that the decrease in oligomer formation is one of the causes of the decrease in decarboxylase activity.
[0083] Any acid decarboxylase, e.g., lysine decarboxylase, arginine decarboxylase, glutamate decarboxylase, or ornithine decarboxylase, may be fused to a prion protein in accordance with the invention. Suitable acid decarboxylases include CadA, LdcC, AdiA, SpeA, SpeC, SpeF, GadA, GadB, and their homologs. As used herein: a lysine decarboxylase refers to an enzyme that converts L-lysine into cadaverine; the enzyme is classified as E.C. 4.1.1.18; an arginine decarboxylase refers to an enzyme that converts L-arginine to agmatine, which can be further converted to purtrescine through the activity of agmatinase, the enzyme is classified as E.C. 4.1.1.19; an ornithine decarboxylase refers to an enzyme that converts ornithine into putrescine, which can be further converted into spermidine and spermine, the enzyme is classified as E.C. 4.1.1.17; and a glutamate decarboxylase refers to an enzyme that converts glutamate to gamma-aminobutyrate (GABA), the enzyme is classified as 4.1.1.15.
[0084] In some embodiments, the lysine decarboxylase is CadA from E. coli or a CadA homolog from another species, e.g., Klebsiella sp.; Enterobacter aerogenes; Salmonella enterica; Serratia sp.; and Raoultella ornithinolytica. In some embodiments, the lysine decarboxylase is LdcC from E. coli or an LdcC homologs from Shigella sp., Citrobacter sp., and Salmonella enterica. As used herein, a lysine decarboxylase includes variants of native lysine decarboxylase enzymes that have lysine decarboxylase enzymatic activity. Additional lysine decarboxylase enzymes are described in PCT/CN2014/080873 and PCT/CN2015/072978.
[0085] In some embodiments, a lysine decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the length of, the cadA polypeptide of SEQ ID NO:2.
[0086] In some embodiments, a lysine decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity; preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, a homolog of a cadA polypeptide.
[0087] In some embodiments, a lysine decarboxylase polypeptide suitable for in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the length of, the LdcC polypeptide of SEQ ID NO:21.
[0088] In some embodiments, a lysine decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity; preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, a homolog of an LdcC polypeptide. Illustrative homologs and accession numbers for the sequences of the polypeptides are: Shigella multispecies (WP 001020996.1), Escherichia fergusonii (WP 001021009.1), Achromobacter sp ATCC35328 (CUJ86682.1), Enterobacteriaceae multispecies (WP 058668594.1), Citrobacter multispecies (WP 016151770.1), Gammoprobacteria multispecies (WP 046401634.1), and Salmonella bongori (WP 038390535.1).
[0089] In some embodiments, an arginine decarboxylase polypeptide suitable for in the inventionis an Adi or a homolog therefore from another species. In some embodiments, an arginine decarboxylase polypeptide has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, SEQ ID NO:22.
[0090] In some embodiments, an arginine decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, a homolog of an AdiA polypeptide. Illustrative homologs and accession numbers for the sequences of the polypeptides are: Shigella multispecies (WP 000978677.1), Escherichia multispecies (WP 000978709.1), Enterobacteriaceae multispecies (WP 032934133.1), Citrobacter multispecies (WP 008786969.1), Klebsiella oxytoca (SAP84601.1), and Salmonella enterica (WP 048668294.1).
[0091] In some embodiments, an arginine decarboxylase polypeptide suitable for in the inventionis an SpeA or a homolog therefore from another species. In some embodiments, an SpeA polypeptide has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, SEQ ID NO:23.
[0092] In some embodiments, an arginine decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, a homolog of an SpeA polypeptide. Illustrative homologs and accession numbers for the sequences of the polypeptides are: Shigella multispecies (WP 005096955.1), Escherichia multispecies (WP 010350365.1), Gammaproteobacteria multispecies (WP 042998051.1), Salmonella multispecies (WP 001278580.1), Achromobacter sp. ATCC35328 (CUJ95389.1), Citrobacter farmeri (WP 042324083.1), Citrobacter koseri (WP 024130934.1), and Citrobacter amalonaticus (WP 052746994.1).
[0093] In some embodiments, an acid decarboxylase polypeptide suitable for in the inventionis an ornithine decarboxylase, SpeC or a homolog thereof, from another species. In some embodiments, the ornithine decarboxylase has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, SEQ ID NO:24.
[0094] In some embodiments, an ornithinedecarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, a homolog of an SpeC polypeptide. Illustrative homologs and accession numbers for the sequences homologs are: Shigella multispecies (WP 005085661.1), Escherichia multispecies (WP 010352539.1), Citrobacter multispecies (WP 044255681.1), Enterobacteriaceae (WP 047357853.1), Gammaproteobacteria multispecies (WP 044327655.1), Achromobacter sp. ATCC35328 (CUJ95194.1), Klebsiella oxytoca (SBL12331.1), and Klebsiella pneumonia (CDK72259.1).
[0095] In some embodiments, an ornithine decarboxylase polypeptide suitable for in the inventionis SpeF or a homolog thereof, from another species. In some embodiments, the ornithine decarboxylase has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, SEQ ID NO:25.
[0096] In some embodiments, an ornithine decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or at least 500 or more amino acids in length, or over the full-length of, a homolog of an SpeF polypeptide. Illustrative homologs and the accession number of polypeptide sequences are: Shigella multispecies (WP 000040203.1), Enterobacteriaceae multispecies (WP 049009856.1), Escherichia multispecies (WP 001292417.1), Gammaproteobacteria multispecies (WP 046401512.1), Citrobacter koseri (WP 024130539.1), Citrobacter amalonaticus (WP 046274704.1), Citrobacter braakii (WP 047501716.1), and Salmonella enterica (WP 023220629.1).
[0097] In some embodiments, an glutamate decarboxylase polypeptide suitable for in the invention is GadA or a homolog thereof, from another species. In some embodiments, the glutamate decarboxylase has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or more amino acids in length, or over the full-length of, SEQ ID NO:26.
[0098] In some embodiments, an glutamate decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or more amino acids in length, or over the full-length of, a homolog of an GadA polypeptide. Illustrative homologs and the accession numbers of the polypeptide sequences are: Escherichia multispecies (WP 001517297.1), Shigella multispecies (WP 000358931.1), Yersinia multispecies (WP 050085789.1), Achromobacter sp. ATCC35328 (WP 054518524.1), and Rhodococcs gingshengii (KDQ00107.1).
[0099] In some embodiments, an glutamate decarboxylase polypeptide suitable for in the invention is GadB or a homolog thereof, from another species. In some embodiments, the glutamate decarboxylase has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or more amino acids in length, or over the full-length of, SEQ ID NO:27.
[0100] In some embodiments, an glutamate decarboxylase polypeptide suitable for use in the invention has at least 60% amino acid sequence identity, preferably at least 65%, 70%, 75%, 80%, 85%, 90%, preferably 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% or greater amino acid sequence identity, preferably over a region of at least about 50, at least 100, at least 200, at least 300, at least 400, or more amino acids in length, or over the full-length of, a homolog of an GadB polypeptide. Illustrative homologs and the accession numbers of the polypeptide sequences are: Shigella multispecies (WP 000358931.1), Escherichia multispecies (WP 016248697.1), Yersinia multispecies (WP 050085789.1), Achromobacter sp. ATCC35328 (WP 054518524.1), Rhodococcs gingshengii (KDQ00107.1)
Nucleic Acids Encoding Prion Subunits and Acid Decarboxylases
[0101] Isolation or generation of acid decarboxylase polynucleotide sequences can be accomplished by a number of techniques In some embodiments, oligonucleotide probes based on the sequences disclosed here can be used to identify the desired polynucleotide in a cDNA or genomic DNA library from a desired bacteria species. Probes may be used to hybridize with genomic DNA or cDNA sequences to isolate homologous genes from different organisms, e.g., fungal species or plant species.
[0102] Alternatively, the nucleic acids of interest can be amplified from nucleic acid samples using routine amplification techniques. For instance, PCR may be used to amplify the sequences of the genes directly from mRNA, from cDNA, from genomic libraries or cDNA libraries. PCR and other in vitro amplification methods may also be useful, for example, to clone nucleic acid sequences that code for proteins to be expressed, to make nucleic acids to use as probes for detecting the presence of the desired mRNA in samples, for nucleic acid sequencing, or for other purposes.
[0103] Appropriate primers and probes for identifying an acid decarboxylase polynucleotide in bacteria can be generated from comparisons of the sequences provided herein. For a general overview of PCR see PCR Protocols: A Guide to Methods and Applications. (Innis, M, Gelfand, D., Sninsky, J. and White, T., eds.), Academic Press, San Diego (1990). Illustrative primer sequences are shown in the Table of Primers in the Examples section.
[0104] Nucleic acid sequences encoding an acid decarboxylase polypeptide for use in the disclosure includes genes and gene products identified and characterized by techniques such as hybridization and/or sequence analysis using illustrative nucleic acid sequences, e.g., a cadA polynucleotide sequence of SEQ ID NO:1. In some embodiments, a host cell is genetically modified by introducing a nucleic acid sequence having at least 60% identity, or at least 70%, 75%, 80%, 85%, or 90% identity, or 100% identity, to an acid decarboxylase polynucleotide, e.g., a cadA polynucleotide of SEQ ID NO:1.
[0105] Nucleic acid sequences encoding an acid decarboxylase fusion protein in accordance with the invention that confers increased production of an amino acid, e.g., lysine, or an amino acid-derived product, e.g., cadaverine, to a host cell, may additionally be codon-optimized for expression in a desired host cell. Methods and databases that can be employed are known in the art. For example, preferred codons may be determined in relation to codon usage in a single gene, a set of genes of common function or origin, highly expressed genes, the codon frequency in the aggregate protein coding regions of the whole organism, codon frequency in the aggregate protein coding regions of related organisms, or combinations thereof. See e.g., Henaut and Danchin in "Escherichia coli and Salmonella," Neidhardt, et al. Eds., ASM Pres, Washington D.C. (1996), pp. 2047-2066; Nucleic Acids Res. 20:2111-2118; Nakamura et al., 2000, Nucl. Acids Res. 28:292).
Preparation of Recombinant Vectors
[0106] Recombinant vectors for expression of an acid decarboxylase fusion protein can be prepared using methods well known in the art. For example, a DNA sequence encoding an acid decarboxylase fusion polypeptide (described in further detail below), can be combined with transcriptional and other regulatory sequences which will direct the transcription of the sequence from the gene in the intended cells, e.g., bacterial cells such as E. coli. In some embodiments, an expression vector that comprises an expression cassette that comprises the gene encoding the acid decarboxylase fusion polypeptide further comprises a promoter operably linked to the nucleic acid sequence encoding the acid decarboxylase fusion polypeptide. In other embodiments, a promoter and/or other regulatory elements that direct transcription of acid decarboxylase fusion polypeptide sequence gene are endogenous to the host cell and an expression cassette comprising the acid decarboxylase fusion gene is introduced, e.g., by homologous recombination, such that the exogenous gene is operably linked to an endogenous promoter and is expression driven by the endogenous promoter.
[0107] As noted above, expression of the polynucleotide encoding an acid decarboxylase fusion protein can be controlled by a number of regulatory sequences including promoters, which may be either constitutive or inducible; and, optionally, repressor sequences, if desired. Examples of suitable promoters, especially in a bacterial host cell, are the promoters obtained from the E. coli lac operon and other promoters derived from genes involved in the metabolism of other sugars, e.g., galactose and maltose. Additional examples include promoters such as the trp promoter, bla promoter bacteriophage lambda PL, and T5. In addition, synthetic promoters, such as the tac promoter (U.S. Pat. No. 4,551,433), can be used. Further examples of promoters include Streptomyces coelicolor agarase gene (dagA), Bacillus subtilis levansucrase gene (sacB), Bacillus licheniformis alpha-amylase gene (amyL), Bacillus stearothermophilus maltogenic amylase gene (amyM), Bacillus amyloliquefaciens alpha-amylase gene (amyQ), Bacillus licheniformis penicillinase gene (penP), Bacillus subtilis xylA and xylB genes. Suitable promoters are also described in Ausubel and Sambrook & Russell, both supra. Additional promoters include promoters described by Jensen & Hammer, Appl. Environ. Microbiol. 64:82, 1998; Shimada, et al., J. Bacteriol. 186:7112, 2004; and Miksch et al., Appl. Microbiol. Biotechnol. 69:312, 2005.
[0108] In some embodiments, a promoter that influences expression of an acid decarboxylase polypeptide may be modified to increase expression. For example, an endogenous acid decarboxylase promoter may be replaced by a promoter that provides for increased expression compared to the native promoter.
[0109] An expression vector may also comprise additional sequences that influence expression of a polynucleotide encoding the acid decarboxylase fusion polypeptide. Such sequences include enhancer sequences, a ribosome binding site, or other sequences such as transcription termination sequences, and the like.
[0110] A vector expressing a polynucleotide encoding an acid decarboxylase fusion polypeptide of the invention may be an autonomously replicating vector, i.e., a vector which exists as an extrachromosomal entity, the replication of which is independent of chromosomal replication, e.g., a plasmid, an extrachromosomal element, a minichromosome, or an artificial chromosome. The vector may contain any means for assuring self-replication. Alternatively, the vector may be one which, when introduced into the host, is integrated into the genome and replicated together with the chromosome(s) into which it has been integrated. Thus, an expression vector may additionally contain an element(s) that permits integration of the vector into the host's genome.
[0111] An expression vector of the invention preferably contains one or more selectable markers which permit easy selection of transformed hosts. For example, an expression vector may comprise a gene that confers antibiotic resistance (e.g., ampicillin, kanamycin, chloramphenicol or tetracycline resistance) to the recombinant host organism, e.g., a bacterial cell such as E. coli.
[0112] Although any suitable expression vector may be used to incorporate the desired sequences, readily available bacterial expression vectors include, without limitation: plasmids such as pSC101, pBR322, pBBR1MCS-3, pUR, pET, pEX, pMR100, pCR4, pBAD24, p15a, pACYC, pUC, e.g., pUC18 or pUC19, or plasmids derived from these plasmids; and bacteriophages, such as Ml 3 phage and .lamda. phage. One of ordinary skill in the art, however, can readily determine through routine experimentation whether any particular expression vector is suited for any given host cell. For example, the expression vector can be introduced into the host cell, which is then monitored for viability and expression of the sequences contained in the vector.
[0113] Expression vectors of the invention may be introduced into the host cell using any number of well-known methods, including calcium chloride-based methods, electroporation, or any other method known in the art.
Host Cells
[0114] The present invention provides for a genetically modified host cell that is engineered to express an acid decarboxylase fusion polypeptide. A genetically modified host strain of the present invention typically comprises at least one additional genetic modification to enhance production of an amino acid or amino acid derivative relative to a control strain that does not have the one additional genetic modification, e.g., a wildtype strain or a cell of the same strain without the one additional genetic modification. An "additional genetic modification to enhance production of an amino acid or amino acid derivative" can be any genetic modification. In some embodiments, the genetic modification is the introduction of a polynucleotide that expresses an enzyme involved in the synthesis of the amino acid or amino acid derivative. In some embodiments, the host cell comprises multiple modifications to increase production, relative to a wildtype host cell, of an amino acid or amino acid derivative.
[0115] In some aspects, genetic modification of a host cell to express an acid decarboxylase fusion polypeptide is performed in conjunction with modifying the host cell to overexpress one or more lysine biosynthesis polypeptides.
[0116] In some embodiments, a host cell may be genetically modified to express one or more polypeptides that affect lysine biosynthesis. Examples of lysine biosynthesis polypeptides include the E. coli genes SucA, Ppc, AspC, LysC, Asd, DapA, DapB, DapD, ArgD, DapE, DapF, LysA, Ddh, PntAB, CyoABE, GadAB, YbjE, GdhA, GltA, SucC, GadC, AcnB, PflB, ThrA, AceA, AceB, GltB, AceE, SdhA, MurE, SpeE, SpeG, PuuA, PuuP, and YgjG, or the corresponding genes from other organisms. Such genes are known in the art (see, e.g., Shah et al., J. Med. Sci. 2:152-157, 2002; Anastassiadia, S. Recent Patents on Biotechnol. 1: 11-24, 2007). See, also, Kind, et al., Appl. Microbiol. Biotechnol. 91: 1287-1296, 2011 for a review of genes involved in cadaverine production. Illustrative genes encoding lysine biosynthesis polypeptides are provided below.
TABLE-US-00001 EC GenBank Protein Gene Number Accession No. .alpha.-ketogultarate sucA 1.2.4.2 YP_489005.1 dehydrogenase (SucA) Phosphoenolpyruvate ppc 4.1.1.31 AAC76938.1 carboxylase (PPC) aspartate transaminase (AspC) aspC 2.6.1.1 AAC74014.1 aspartate kinase (LysC) lysC 2.7.2.4 NP_418448.1 aspartate semialdehyde asd 1.2.1.11 AAC76458.1 dehydrogenase (Asd) dihydrodipicolinate synthase dapA 4.3.3.7 NP_416973.1 (DapA) dihydropicolinate reductase dapB 1.17.1.8 AAC73142.1 (DapB) tetrahydrodipicoinate dapD 2.3.1.117 AAC73277.1 succinylase (DapD) N-succinyldiaminopimelate argD 2.6.1.11 AAC76384.1 aminotransferase (ArgD) N-succinyl-L-diaminopimelate dapE 3.5.1.18 AAC75525.1 deacylase (DapE) diaminopimelate epimerase dapF 5.1.1.7 AAC76812.2 (DapF) diaminopimelate decarboxylase lysA 4.1.1.20 AAC75877.1 (LysA) meso-diaminopimelate ddh NA P04964.1 dehydrogenase (Ddh) pyridine nucleotide pntAB NA AAC74675.1, transhydrogenase (PntAB) AAC74674.1 cytochrome O oxidase cyoABE 1.10.3.10 AAC73535.1, (CyoABE) AAC73534.1, AAC73531.1 glutamate decarboxylase gadAB 4.1.1.15 AAC76542.1, (GadAB) AAC74566.1 L-amino acid efflux transporter ybjE NA AAC73961.2 (YbjE) glutamate dehydrogenase gdhA 1.4.1.4 AAC74831.1 (GdhA) citrate synthase (GltA) gltA 2.3.3.1/ AAC73814.1 2.3.3.16 succinyl-coA synthase (SucC) sucC 6.2.1.5 AAC73822.1 glutamate-GABA antiporter gadC NA AAC74565.1 (GadC) aconitase B (AcnB) acnB 4.2.1.99 AAC73229.1 pyruvate-formate lyase (PflB) pflB NA AAC73989.1 aspartate kinase/homoserine thrA 2.7.2.4 AAC73113.1 dehydrogenase (ThrA) isocitrate lyase (AceA) aceA 4.1.3.1 AAC76985.1 malate synthase (AceB) aceB 2.3.3.9 AAC76984.1 glutmate synthase (GltB) gltB 1.4.1.13 AAC76244.2 pyruvate dehydrogenase (AceE) aceE 1.2.4.1 AAC73225.1 succinate dehydrogenase sdhA 1.3.5.1 AAC73817.1 (SdhA) UDP-N-acetylmuramoyl-L- murE 6.3.2.13 AAC73196.1 alanyl-D-glutamate:meso- diaminopimelate ligase (MurE) putrescine/cadaverine speE 2.5.1.16 AAC73232.1 aminopropyltransferase (SpeE) spermidine acetyltransferase speG NA AAC74656.1 (SpeG) glutamate-putrescine/glutamate- puuA NA AAC74379.2 cadaverine ligase (PuuA) putrescine importer (PuuP) puuP NA AAC74378.2 putrescine/cadaverine ygjG 2.6.1.82 AAC76108.3 aminotransferase (YgjG)
[0117] In some embodiments, a host cell may be genetically modified to attenuate or reduce the expression of one or more polypeptides that affect lysine biosynthesis. Examples of such polypeptides include the E. coli genes Pck, Pgi, DeaD, CitE, MenE, PoxB, AceA, AceB, AceE, RpoC, and ThrA, or the corresponding genes from other organisms. Such genes are known in the art (see, e.g., Shah et al., J. Med. Sci. 2:152-157, 2002; Anastassiadia, S. Recent Patents on Biotechnol. 1: 11-24, 2007). See, also, Kind, et al., Appl. Microbiol. Biotechnol. 91: 1287-1296, 2011 for a review of genes attenuated to increase cadaverine production. Illustrative genes encoding polypeptides whose attenuation increases lysine biosynthesis are provided below.
TABLE-US-00002 EC GenBank Protein Gene Number Accession No. PEP carboxykinase (Pck) pck 4.1.1.49 NP_417862 Glucose-6-phosphate pgi 5.3.1.9 NP_418449 isomerase (Pgi) DEAD-box RNA deaD NP_417631 helicase (DeaD) citrate lyase (CitE) citE 4.1.3.6/ NP_415149 4.1.3.34 o-succinylbenzoate-CoA menE 6.2.1.26 NP_416763 ligase (MenE) pyruvate oxidase (PoxB) poxB 1.2.2.2 NP_415392 isocitrate lyase (AceA) aceA 4.1.3.1 NP_418439 malate synthase A (AceB) aceB 2.3.3.9 NP_418438 pyruvate dehydrogenase (aceE) aceE 1.2.4.1 NP_414656 RNA polymerase b' subunit rpoC 2.7.7.6 NP_418415 (RpoC) aspartokinase I (ThrA) thrA 2.7.2.4/ NP_414543 1.1.1.3
[0118] Nucleic acids encoding a lysine biosynthesis polypeptide may be introduced into the host cell along with acid decarboxylase fusion polynucleotide, e.g., encoded on a single expression vector, or introduced in multiple expression vectors at the same time. Alternatively, the host cell may be genetically modified to overexpress one or more lysine biosynthesis polypeptides before or after the host cells genetically modified express an acid decarboxylase fusion polypeptide.
[0119] A host cell engineered to express an acid decarboxylase fusion polypeptideis typically a bacterial host cell. In typical embodiments, the bacterial host cell is a Gram-negative bacterial host cell. In some embodiments of the invention, the bacterium is an enteric bacterium. In some embodiments of the invention, the bacterium is a species of the genus Corynebacterium, Escherichia, Pseudomonas, Zymomonas, Shewanella, Salmonella, Shigella, Enterobacter, Citrobacter, Cronobacter, Erwinia, Serratia, Proteus, Hafnia, Yersinia, Morganella, Edwardsiella, or Klebsiella taxonomical classes. In some embodiments, the host cells are members of the genus Escherichia, Hafnia, or Corynebacterium. In some embodiments, the host cell is an Escherichia coli, Hafnia alvei, or Corynebacterium glutamicum host cell.
[0120] In some embodiments, the host cell is a gram-positive bacterial host cell, such as a Bacillus sp., e.g., Bacillus subtilis or Bacillus licheniformis; or another Bacillus sp. such as B. alcalophilus, B. aminovorans, B. amyloliquefaciens, B. caldolyticus, B. circulans, B. stearothermophilus, B. thermoglucosidasius, B. thuringiensis or B. vulgatis.
[0121] Host cells modified in accordance with the invention can be screened for increased production of lysine or a lysine derivative, such as cadaverine, as described herein.
[0122] In some embodiments, an acid decarboxylase fusion protein of the present invention may be recovered from a host cell that expresses the fusion protein. In some embodiments, the recovered fusion protein may be immobilized onto a solid substrate or inert material to form an immobilized enzyme. In one embodiment, the immobilized enzyme may have improved thermal stability and/or operational stability than the soluble form of the fusion protein. In one embodiment, the fusion protein comprises a lysine, arginine, ornithine, or glutamate decarboxylase fused at the C-terminal end to a prion subunit.
Methods of Producing Lysine or a Lysine Derivative.
[0123] A host cell genetically modified to express an acid decarboxylase fusion polypeptide can be employed to produce lysine or a derivative of lysine. In some embodiments, the host cell produces cadaverine. To produce lysine or the lysine derivative, a host cell genetically modified to express an acid decarboxylase fusion polypeptide as described herein can be cultured under conditions suitable to allow expression of the polypeptide and expression of genes that encode the enzymes that are used to produce lysine or the lysine derivative. A host cell modified in accordance with the invention to express an acid decarboxylase fusion polypeptide provides a higher yield of lysine or lysine derivatives relative to a non-modified counterpart host cell that expresses the acid decarboxylase that is not fused to a prion subunit.
[0124] Host cells may be cultured using well known techniques (see, e.g., the illustrative conditions provided in the examples section.
[0125] In some embodiments, host cells are cultured using nitrogen sources that are not salts (e.g., ammonium sulfate or ammonium chloride), such as ammonia or urea. Host cells may be cultured at an alkaline pH during cell growth or enzyme production.
[0126] The lysine or lysine derivative then be separated and purified using known techniques. Lysine or lysine derivatives, e.g., cadverine, produced in accordance with the invention may then be used in any known process, e.g., to produce a polyamide.
[0127] In some embodiments, lysine may be converted to caprolactam using chemical catalysts or by using enzymes and chemical catalysts.
[0128] The present invention will be described in greater detail by way of specific examples. The following examples are offered for illustrative purposes, and are not intended to limit the invention in any manner. Those of skill in the art will readily recognize a variety of noncritical parameters, which can be changed or modified to yield essentially the same results.
EXAMPLES
Example 1: Construction of Plasmid Vectors that Encode CadA
[0129] A plasmid vector containing wild-type E. coli cadA (SEQ ID NO: 1), which encodes the lysine decarboxylase CadA (SEQ ID NO: 2), was amplified from the E. coli MG1655 K12 genomic DNA using the PCR primers cadA-F and cadA-R (FIG. 1), digested using the restriction enzymes SacI and XbaI, and ligated into pUC18 to generate the plasmid pCIB60. The 5' sequence upstream of the cadA gene was optimized using the PCR primers cadA-F2 and cadA-R2 to create pCIB71. The chloramphenicol resistance gene cat was amplified using the primers cat-HindIII-F and cat-NdeI-R, and cloned behind cadA in pCIB71 to create pCIB128.
Example 2: Synthesis of Codon Optimized New1 and Sup35 Prion Fragments
[0130] The minimal polypeptide fragment necessary for New1 and Sup35 prion activity was determined based on Osherovich et al., PLOS Biology 2:4, 2004. Therefore, amino acids 2-100 of New1 (SEQ ID NO: 3) and 2-103 of Sup35 (SEQ ID NO: 4) were codon optimized for heterologous expression in E. coli (SEQ ID NO: 5 and 6). In addition, a short polypeptide linker sequence that consists of the amino acids GSGSG was added to the beginning of SEQ ID NO: 3 and 4 (SEQ ID NO: 7 and 8), and their corresponding DNA sequences are presented in SEQ ID NO: 9 and 10. Codon optimization and DNA assembly was performed according to Hoover D M & Lubkowski J, Nucleic Acids Research 30:10, 2002.
Example 3: Construction of a Polynucleotide Encoding a Fragment of New1 Fused 3' of CadA
[0131] The stop codon of cadA in pCIB128 was removed using the primers cadAt-XbaI-R and cat-XbaI-F to create the plasmid pCIB138. The DNA fragment that consists of the prion domain of New1 with the polypeptide linker was amplified using the primers New1-XbaI-F and New1-HindIII-R, digested using the restriction enzymes XbaI and HindIII, and ligated into pCIB138 to make pCIB222.
Example 4: Construction of a Polynucleotide Encoding a Fragment of Sup35 Fused 3' of CadA
[0132] The DNA fragment that consists of the prion domain of Sup35 with the polypeptide linker was amplified using the primers 35-XbaI-F and 35-HindIII-R, digested using the restriction enzymes XbaI and HindIII, and ligated into pCIB138 to make pCIB223.
Example 5: Lysine Decarboxylase Activity of Novel Polypeptide Consisting of Either the New1 or Sup35 Prion Domain Fragment Fused 3' of CadA
[0133] H. alvei was transformed with pCIB128, pCIB222, and pCIB223. Three single colonies from each transformation were grown overnight at 37.degree. C. in 4 mL of LB medium with ampicillin (100 .mu.g/mL). The following day, 0.7 mL of each overnight culture was added to 0.3 mL of lysine-HCl and PLP to a final concentration of 120 g/L and 0.1 mM, respectively. Each mixture was incubated at 37.degree. C. for 2 hours. Cadaverine production from each sample was quantified using NMR, and yield/OD was calculated by dividing the molar amount of cadaverine produced by the molar amount of lysine added. The yield from each sample after 2 hours is presented in Table 1.
TABLE-US-00003 TABLE 1 Production of cadaverine H. alvei strains overproducing a fusion polypeptide consisting of a prion domain fragment and a lysine decarboxylase. Cadaverine Yield Strain Plasmid (%) H. alvei pCIB128 67.4 .+-. 4.7 pCIB222 48.4 .+-. 5.5 pCIB223 50.3 .+-. 4.2
[0134] As shown in Table 1, both polypeptides that consisted of either a New1 or Sup35 prion domain fragment fused 3' of CadA showed significant lysine decarboxylase activity. The lysine decarboxylase activity was lower than the control, where the lysine decarboxylase was expressed by itself without being fused to a prion domain fragment. The decrease in activity could be caused by many factors, some of which are a decrease cell density and total protein, a decrease in lysine decarboxylase activity, a decrease in the amount of functional soluble enzyme, or an increase of the insoluble protein fraction as a result of the fusion polypeptide having difficulty folding.
Example 6: Construction of a Polynucleotide Encoding a Fragment of New1 Fused 5' of CadA
[0135] The cadA gene was amplified using the primers cadA-XbaI-F and cadA-HindIII-R, digested using the restriction enzymes XbaI and HindIII, and ligated into pCIB128 to create the plasmid pCM146 having two copies of the cadA gene. The SacI restriction site was added 5' of the first cadA gene after the promoter using the primers rbs2-SacI-F and rbs2-SacI-R to construct pCIB149. The New1 prion fragment was amplified using the primers New1-SacI-F and New1-XbaI-R, digested using the restriction enzymes SacI and XbaI, and ligated into pCM149 to create pCIB241.
Example 7: Construction of a Polynucleotide Encoding a Fragment of Sup35 Fused 5' of CadA
[0136] The Sup35 prion fragment was amplified using the primers Sup35-SacI-F and Sup35-XbaI-R, digested using the restriction enzymes SacI and XbaI, and ligated into pCM149 to create pCIB242.
Example 8: Lysine Decarboxylase Activity of Novel Polypeptide Consisting of Either the New1 or Sup35 Prion Domain Fragment Fused 5' of CadA
[0137] H. alvei was transformed with pCIB128, pCIB241, and pCIB242. Three single colonies from each transformation were grown overnight at 37.degree. C. in 4 mL of LB medium with ampicillin (100 .mu.g/mL). The following day, 0.7 mL of each overnight culture was added to 0.3 mL of lysine-HCl and PLP to a final concentration of 120 g/L and 0.1 mM, respectively. Each mixture was incubated at 37.degree. C. for 2 hours. Cadaverine production from each sample was quantified using NMR, and yield/OD was calculated by dividing the molar amount of cadaverine produced by the molar amount of lysine added. The yield from each sample after 2 hours is presented in Table 2.
TABLE-US-00004 TABLE 2 Production of cadaverine H. alvei strains overproducing a fusion polypeptide consisting of a prion domain fragment and a lysine decarboxylase. Cadaverine Yield Strain Plasmid (%) H. alvei pCIB128 68.8 .+-. 3.5 pCIB241 5.2 .+-. 1.6 pCIB242 8.3 .+-. 3.2
[0138] As shown in Table 2, both polypeptides that consisted of either a New1 or Sup35 prion domain fragment fused 5' of CadA showed very little lysine decarboxylase activity. The lysine decarboxylase activity was lower than the control, where the lysine decarboxylase was expressed by itself without being fused to a prion domain fragment. The lysine decarboxylase activities of these enzymes were also lower compared to those where the prion domain fragments were fused 3' of CadA. This indicates that the synthetic fusion polypeptide is more functional when the prion domain is fused 3' of the acid decarboxylase after the C-terminal domain, and less functional when the prion domain is fused 5' of the acid decarboxylase in front of the N-terminal wing domain.
Example 9: Cell Density and Enzyme Production by H. Alvei Overexpressing a New1 Prion Domain Fused to CadA
[0139] To determine the cause for the decrease in observed total activity when the New1 prion domain fragment was fused 3' of the lysine decarboxylase, the cell density of an overnight culture was measured at Abs600, and the culture was lysed in order to determine the total protein content using the Bradford assay. The lysed cell culture was analyzed using SDS-PAGE in order to determine how much of the total protein consisted of the synthetic fusion polypeptide. Cell lysis was performed using a combination of freeze thaw and lysozyme. Lysed samples were treated with DNAse in order to remove most DNA. The OD and total protein data is presented in Table 3. The result of the SDS-PAGE is shown in FIG. 1.
TABLE-US-00005 TABLE 3 OD and total protein data of a fusion polypeptide consisting of a prion domain fragment and a lysine decarboxylase. OD Total protein Strain Plasmid (Abs.sub.600) (mg/mL) H. alvei pCIB128 4.1 .+-. 0.5 1.05 pCIB222 3.4 .+-. 0.1 0.65
[0140] As shown in Table 3, both the observed OD and total protein are lower when H. alvei is producing the fusion polypeptide (pCIB222) compared to producing the lysine decarboxylase alone (pCIB128). The OD decreased by 17%, while the total protein decreased by 38%. The SDS-PAGE result indicates that the background protein did not change significantly; instead, the target protein decreased significantly (the dark band at 80 kDa). Furthermore, most of the target protein is found in the soluble fraction, which indicates that protein solubility is not a problem. Therefore, the decrease in total activity is not due to accumulation of inactive protein that is insoluble.
Example 10: Construction of Different Sized Fragments of the New1 Prion Domain Fused to CadA
[0141] It was observed that the fusion of the prion domain fragment to lysine decarboxylase affects the growth of the host cell, and decreases the final cell density (OD measured at Abs600) achieved when compared to the control without the prion domain fragment. As demonstrated above, the decrease in cell density decreases the amount of soluble enzyme and total activity produced. The decrease in OD is not due to the accumulation of poorly folded enzyme, as indicated in the SDS-PAGE result. However, the reduction in cell density could be caused by challenges in folding of the new fusion polypeptide (e.g., slower kinetics).
[0142] Different sized fragments of the New1 prion domain were generated to determine whether truncating a portion of the prion domain would decrease the burden on the cell and improve growth and the final cell density. The N-terminus of the New1 prion domain fragment was truncated by various lengths of amino acids in order to determine whether the portion of the fragment less rich in Q/N residues were important for lysine decarboxylase function of the fusion polypeptide, and whether their removal could improve growth of the host. The first 9, 18, 36, and 45 amino acids of the New1 prion domain fragment in pCIB222 were truncated (SEQ ID NO: 11, 12, 13, and 14) using the PCR primer 222-de-R and the respective PCR primers 222-de-9-F, 222-de18-F, 222-de36-F, and 222-de45-F.
Example 11: Improvements in Cell Density by H. Alvei Overexpressing Variants of the New1 Prion Domain Fused to CadA
[0143] E. coli BL21 was transformed with either pCIB222 or a variant of pCIB222 with the first 9, 18, 27, 36, or 45 amino acids of the New1 prion domain fragment truncated. Three single colonies from each transformation were grown overnight at 37.degree. C. in 4 mL of LB medium with ampicillin (100 .mu.g/mL). The following day, the absorbance of the overnight culture was measured at Abs600. The OD of each sample is presented in Table 4, and the SDS-PAGE analysis of each sample is shown in FIG. 2.
TABLE-US-00006 TABLE 4 OD of truncated variants of a fusion polypeptide consisting of a prion domain fragment and a lysine decarboxylase. Strain Plasmid Amino acids deleted OD (Abs.sub.600) E. coli pCIB222 none 5.4 .+-. 0.2 pCIB222-de9 FPPKKFKDL 6.5 .+-. 0.4 pCIB222-de18 FPPKKFKDLNSFLDDQPK 6.1 .+-. 0.2 pCIB222-de36 FPPKKFKDLNSFLDDQPK 6.3 .+-. 0.1 DPNLVASPFGGYFKNPAA pCIB222-de45 FPPKKFKDLNSFLDDQPK 6.2 .+-. 0.1 DPNLVASPFGGYFKNPAA DAGSNNASK
[0144] Table 4 shows that the truncated versions of the New1 prion domain fragment improved growth of the E. coli BL21 host compared to the control with the entire prion domain fragment. There was no significant difference in the final OD between the strains producing different versions of the truncated enzymes. Even though no difference could be seen between the truncated versions based on the OD data, the SDS-PAGE analysis shows that significant increases in total protein and the target enzyme are observed for the version with 36 amino acids truncated. The versions of the enzymes with 9, 18, and 45 amino acids truncated show less total protein and target enzyme compared to the control.
Example 12: Modification of the C-Terminal Amino Acids of the Polynucleotide Encoding a Fragment of New1 Fused to CadA
[0145] It is common for synthetic proteins that fold slowly or poorly to be degraded when overexpressed. Therefore, reducing protein degradation in order to increase the time a polypeptide has to fold would increase the amount of functional protein in a cell. Previous work (Trepod C M & Mott J E, J Biotechnol 84, 273-284, 2000) reported that specific sequences of short polypeptides when fused to the C-terminus of a protein can increases protein stability and reduce intracellular protein degradation. These short polypeptides are derived from the C-terminus of proteins such as bovine somatotropin (BST), RpoC, .lamda.CI, Rho, RecA, Bla, and TufA.
[0146] Three different short polypeptides derived from the genes BST (C2), E. coli .lamda.CI (C4), and E. coli recA (C6) were fused to the C-terminus of the New1 prion domain fragment in pCIB222 using the respective PCR primers 222-C2-F, 222-C2-R, 222-C4-F, 222-C4-R, 222-C6-F, and 222-C6-R. The BST fragment consists of the amino acid sequence RRFGEASSAF (SEQ ID NO: 15 and 16), the .lamda.CI fragment consists of the amino acid sequence ASQWPEETFG (SEQ ID NO: 17 and 18), and the RecA fragment consists of the amino acid sequence EGVAETNEDF (SEQ ID NO: 19 and 20).
Example 13: Improvements in Total Enzyme Activity by E. coli Overexpressing C-Terminal Variants of the New1 Prion Domain Fused to CadA
[0147] E. coli BL21 was transformed with either pCIB222 or a variant of pCIB222 where the BST, .lamda.CI, or RecA fragment is fused to the C-terminus of the New1 prion domain fragment. Three single colonies from each transformation were grown overnight at 37.degree. C. in 4 mL of LB medium with ampicillin (100 .mu.g/mL). The following day, 0.6 mL of each overnight culture was added to 0.4 mL of lysine-HCl and PLP to a final concentration of 160 g/L and 0.1 mM, respectively. Each mixture was incubated at 37.degree. C. for 2 hours. Cadaverine production from each sample was quantified using NMR, and yield was calculated by dividing the molar amount of cadaverine produced by the molar amount of lysine added. The yield from each sample after 2 hours is presented in Table 5.
TABLE-US-00007 TABLE 5 Cadaverine production by E. coli producing C-terminal variants of a fusion polypeptide consisting of a prion domain fragment and a lysine decarboxylase. Strain Plasmid Peptide added Cadaverine Yield (%) E. coli pCIB222 none 32.2 .+-. 0.2 pCIB222-C2 RRFGEASSAF 34.8 .+-. 0.1 pCIB222-C4 ASQWPEETFG 41.5 .+-. 0.2 pCIB222-C6 EGVAETNEDF 37.5 .+-. 0.1
[0148] Table 5 shows that modification of the C-terminus of the New1 prion domain fragment with an additional short polypeptide derived from .lamda.CI improved cadaverine yield from lysine. Modification of the prion domain fragment with an additional short polypeptide derived from either BST or RecA showed a less significant change in cadaverine yield. The increase in total activity observed in the strain harboring the .lamda.CI C-terminal fragment suggests an increase in the stability of the triple fusion polypeptide that consists of CadA, the New1 prion domain fragment, and the .lamda.CI polypeptide compared to the double fusion polypeptide that lacks the .lamda.CI polypeptide.
Example 14: Effect of the Addition of Sorbitol on Lysine Decarboxylase Activity by a Polypeptide Consisting of New1 Fused to CadA
[0149] Protein chaperones can be overexpressed in a host in order to improve the amount of functional protein produced in a cell, especially in the case of a synthetic protein that does not fold well or is not native to the host. For example, the common chaperone protein systems used in E. coli are GroEL/GroES, DnaK/DnaJ/GrpE, ClpB, and heat shock proteins/IbpA/IbpB (de Marco et al., BMC Biotechnol 7:32, 2007). Chemical chaperones that can increase the amount of soluble and functional protein produced have also been demonstrated (Prasad, et al., Appl Environ Microbiol 77, 4603-4609, 2011).
[0150] To improve the folding of the synthetic polypeptide that consists of the New1 prion domain fused to a lysine decarboxylase CadA, sorbitol was added during protein production. E. coli BL21 and H. alvei were transformed with pCIB222. Three single colonies from each transformation were grown overnight at 37.degree. C. in 4 mL of LB medium with ampicillin (100 .mu.g/mL), and enough sorbitol to reach a final concentration of either 0, 0.2, or 0.8 M. The following day, 0.6 mL of each overnight culture was added to 0.4 mL of lysine-HCl and PLP to a final concentration of 160 g/L and 0.1 mM, respectively. Each mixture was incubated at 37.degree. C. for 2 hours. Cadaverine production from each sample was quantified using NMR, and yield/OD was calculated by dividing the molar amount of cadaverine produced by the molar amount of lysine added and dividing again by the absorbance of the overnight culture at Abs600. The yield/OD from each sample is presented in Table 6.
TABLE-US-00008 TABLE 6 Production of cadaverine by E. coli and H. alvei strains grown with sorbitol and overproducing the fusion polypeptide. Cadaverine Sorbitol OD Yield/OD Strain Plasmid (mM) (Abs.sub.600) (%) E. coli pCIB222 0 6.2 .+-. 0.3 15.7 .+-. 0.3 0.2 6.0 .+-. 0.2 9.2 .+-. 0.2 0.8 4.0 .+-. 0.1 9.0 .+-. 0.2 H. alvei 0 5.4 .+-. 0.2 7.9 .+-. 0.2 0.2 5 .+-. 0.2 9.3 .+-. 0.2 0.8 4.7 .+-. 0.1 10.0 .+-. 0.3
[0151] As shown in Table 6, the addition of sorbitol affected enzyme activity differently depending on the strain used. In both E. coli and H. alvei, sorbitol negatively affected growth at concentration of 0.8 M compared to 0 M, and the effect on growth is less significant at 0.2 M. The addition of sorbitol increased the yield/OD when H. alvei was the host, whereas the addition of sorbitol decreased the yield/OD when E. coli was the host. Although the addition of sorbitol decreases the OD in H. alvei, sorbitol increased the lysine decarboxylase activity per cell and increased total activity per culture volume.
Example 15: In Vitro Lysine Decarboxylase Activity and Kinetics of a Polypeptide Consisting of New1 Fused to CadA at Different pHs
[0152] According to the literature, the activity of lysine decarboxylase at pH 8 is significantly less than its activity at pH 6 due to a structural change from a high oligomer state to a low oligomer state. The activity of lysine decarboxylase with and without the New1 prion fragment fused to its C-terminus was compared at pH 6 and pH 8, in order to determine whether the prion fragment can increase the tolerance of the polypeptide to alkaline conditions.
[0153] 100 mL samples of H. avlei transformed with either pCIB128 or pCIB222 were lysed with a french press. The lysed samples were centrifuged, and the supernatant was separated from the pellet in order to perform in vitro experiments. The reaction rate of each lysed sample was measured using NMR by sampling the amount of lysine converted in the presence of PLP into cadaverine every 1.6 minutes for a total of 20 minutes, and taking the slope of the linear portion of the yield curve. The samples were diluted so that the reaction rate per volume (U) of lysed sample was the same. The kinetic constants Vmax and Km for lysine of each lysed samples was measured using the same U at an initial pH of either 6 or pH 8. By normalizing for U, the concentration of active enzyme in each sample is the same. The results of the kinetic analysis of the two samples are shown in Table 7.
TABLE-US-00009 TABLE 7 Kinetic analysis of the effect of pH on lysed samples of H. avlei producing lysine decarboxylase with and without the New1 prion domain fragment. pH CIB128 CIB222 6 Vmax (mmol/min) 3.9 4.1 Km (mM) 27 25 8 Vmax (mmol/min) 2.6 3.7 Km (mM) 27 25
[0154] In accordance with the literature, wild-type lysine decarboxylase (CIB128) lost a significant amount of activity at pH 8 compared to pH 6. The reduction in activity at pH 8 compared to pH 6 was 33%. Surprisingly, the lysine decarboxylase fused to the New1 prion domain (CIB222) only lost 10% of its activity when the initial pH was 8 compared to 6. The Km of either wild-type or modified lysine decarboxylase for lysine did not change even though the initial pH changed. Since the amount of active enzyme added is normalized by U, the ability for the modified enzyme to better tolerate alkaline pH is likely due to its ability to maintain a higher oligomer state compared to the wild-type enzyme.
Example 16: In Vitro Lysine Decarboxylase Activity and Kinetics of a Polypeptide Consisting of New1 Fused to CadA at Different Temperatures
[0155] Tolerance to high temperature is another beneficial trait of a biological enzyme in a large-scale production system, especially during the summer months in order to reduce the cost of cooling the reactor. Most enzymes operate within a narrow temperature range, and temperatures higher than that range tend to cause the enzymes to denature or not exist in structural states necessary for function. The lysine decarboxylase fused with a New1 prion domain fragment showed increased tolerance to high pH. It is possible that the increased structural stability provided by the prion domain fragment is useful not only for tolerating high pH, but also high temperature. The activities of lysine decarboxylase with and without the New1 prion domain fragment were determined following incubation for different periods of time at 37.degree. C., 45.degree. C., and 55.degree. C., in order to determine whether the enzymes are able to maintain the structural integrity necessary for function at high temperatures.
[0156] 100 mL samples of H. avlei transformed with either pCIB128 or pCIB222 were lysed with a french press. The lysed samples were centrifuged, and the supernatant was separated from the pellet in order to perform in vitro experiments. The reaction rate of each lysed sample was measured using NMR in the presence of PLP by sampling the amount of lysine converted into cadaverine every 1.6 minutes for a total of 20 minutes, and taking the slope of the linear portion of the yield curve. The samples were diluted so that the reaction rate per volume (U) of lysed sample was the same. Equal amounts of enzyme based on U were incubated at 37.degree. C., 45.degree. C., and 55.degree. C. for 0, 1, 2, 4, and 20 hours. After incubation at the specific temperature and time period, the reaction rate of each enzyme sample was determined at 37.degree. C. The effects of temperature and time on the two different enzymes are shown in Table 8.
TABLE-US-00010 TABLE 8 Relative activity after incubation at different temperatures for different periods of time of lysine decarboxylase with or without the New1 prion domain fragment. Temp Time (h) Strain (.degree. C.) 0 1 2 4 20 CIB128 37 100 100 98 81 70 45 100 95 92 55 37 55 100 8 6 0 0 CIB222 37 100 100 95 93 93 45 100 94 98 89 69 55 100 93 80 71 51
[0157] Table 8 shows the surprising discovery that not only does the New1 prion domain fragment increase the tolerance of lysine decarboxylase for alkaline pH, but it also increases the tolerance of the enzyme for high temperature. The wild-type lysine decarboxylase (CIB128) lost almost all of its activity after incubation at 55.degree. C. for one hour. However, the lysine decarboxylase fused with a fragment of the New1 prion domain (CIB222) was able to maintain 93% of its activity. Furthermore, when no detectable activity by the wild-type enzyme was observed after 4 hours of incubation at 55.degree. C., the fusion polypeptide still maintained 71% of its activity. Therefore, the increased stability imparted on the acid decarboxylase by fusing it with a prion domain fragment is not specific for tolerating a single environmental stress, and can enable the new enzyme to function across a wider range of operating conditions useful for industrial production.
TABLE-US-00011 Table of plasmids used in Examples Host Protein(s) Overexpressed Plasmid Strain CadA pCIB71 CadA, Cat pCIB128 CadA, CadA pCIB146 CadA-New1 pCIB222 CadA-Sup35 pCIB223 New1-CadA pCIB241 Sup35-CadA pCIB242 CadA-New1 (.DELTA.9) pCIB222-de9 CadA-New1 (.DELTA.18) pCIB222-de18 CadA-New1 (.DELTA.36) pCIB222-de36 CadA-New1 (.DELTA.45) pCIB222-de45 CadA-New1-C2 pCIB222-C2 CadA-New1-C4 pCIB222-C4 CadA-New1-C6 pCIB222-C6 E. coli CadA-New1 pCIB222 CIB222-EC E. coli CadA-New1 (.DELTA.9) pCIB222-de9 CIB222-de9 E. coli CadA-New1 (.DELTA.18) pCIB222-de18 CIB222-de18 E. coli CadA-New1 (.DELTA.36) pCIB222-de36 CIB222-de36 E. coli CadA-New1 (.DELTA.45) pCIB222-de45 CIB222-de45 E. coli CadA-New1-C2 pCIB222-C2 CIB222-C2 E. coli CadA-New1-C4 pCIB222-C4 CIB222-C4 E. coli CadA-New1-C6 pCIB222-C6 CIB222-C6 H. avlei CadA, Cat pCIB128 CIB128 H. avlei CadA-New1 pCIB222 CIB222 H. avlei CadA-Sup35 pCIB223 CIB223 H. avlei New1-CadA pCIB241 CIB241 H. avlei Sup35-CadA pCIB242 CIB242
TABLE-US-00012 Table of primer sequences used in Examples. Name Sequence (5'-3') cadA-F ggcgagctcacacaggaaacagaccatgaacgttattgcaatattgaatcac cadA-R ggctctagaccacttcccttgtacgagc cadA-F2 atttcacacaggaaacagctatgaacgttattgcaatattgaat cadA-R2 agctgtttcctgtgtgaaat cat-HindIII-F ggcaagcttgagaaaaaaatcactggatatacc cat-NdeI-R ggccatatgtaagggcaccaataactgcc cadAt-XbaI-R ggctctagatttgctttcttctttcaatacc cat-XbaI-F ggctctagagagaaaaaaatcactggatatacc New1-XbaI-F ggctctagaggttctggctctggttctccg New1-HindIII-R ggcaagcttttactggtagccctgaccgttg Sup35-XbaI-F ggctctagaggtagcggctctggctctga Sup35-HindIII-R ggcaagcttttagccaccctgtgggttaaact cadA-XbaI-F ggctctagaatttcacacaggaaacagct cadA-HindIII-R ggcaagcttcacttcccttgtacgagcta rbs2-SacI-F ggcgagctcatgaacgttattgcaatattgaatc rbs2-SacI-R ggcgagctcctcctgtgtgaaattg New1-SacI-F ggcgagctcatgggttctggctctggttc New1-XbaI-R ggctctagactggtagccctgaccgttg Sup35-SacI-F ggcgagctcatgggtagcggctctggc Sup35-XbaI-R ggctctagagccaccctgtgggttaaact 222-de-R tctagatttgctttcttctttcaatacc 222-de-9-F gaagaaagcaaatctagaaagttcaaagacctgaactctttc ggtg 222-de-18-F gaagaaagcaaatctagagacgaccagccgaaagacccgaac 222-de-36-F gaagaaagcaaatctagaaaaaacccagcggcggacgcggg 222-de-45-F gaagaaagcaaatctagaaacaacgcgtctaagaaatcttc 222-C2-F tttcggcgaagcgagcagcgcgttctaaaagcttaagagacaggatg 222-C2-R cgctgctcgcttcgccgaaacgacgctggtagccctgaccgttgtat 222-C4-F ccagtggccggaagaaaccttcggctaaaagcttaagagacaggatg 222-C4-R aggtttcttccggccactggctcgcctggtagccctgaccgttgtat 222-C6-F cgtggcggaaaccaacgaagatttctaaaagcttaagagacaggatg 222-C6-R cttcgttggtttccgccacgccttcctggtagccctgaccgttgtat
Illustrative Sequences:
TABLE-US-00013 [0158] Escherichia coli cadA nucleic acid sequence SEQ ID NO: 1 ATGAACGTTATTGCAATATTGAATCACATGGGGGTTTATTTTAAAGAAGAACCCATC CGTGAACTTCATCGCGCGCTTGAACGTCTGAACTTCCAGATTGTTTACCCGAACGAC CGTGACGACTTATTAAAACTGATCGAAAACAATGCGCGTCTGTGCGGCGTTATTTTT GACTGGGATAAATATAATCTCGAGCTGTGCGAAGAAATTAGCAAAATGAACGAGAA CCTGCCGTTGTACGCGTTCGCTAATACGTATTCCACTCTCGATGTAAGCCTGAATGA CCTGCGTTTACAGATTAGCTTCTTTGAATATGCGCTGGGTGCTGCTGAAGATATTGCT AATAAGATCAAGCAGACCACTGACGAATATATCAACACTATTCTGCCTCCGCTGACT AAAGCACTGTTTAAATATGTTCGTGAAGGTAAATATACTTTCTGTACTCCTGGTCAC ATGGGCGGTACTGCATTCCAGAAAAGCCCGGTAGGTAGCCTGTTCTATGATTTCTTT GGTCCGAATACCATGAAATCTGATATTTCCATTTCAGTATCTGAACTGGGTTCTCTGC TGGATCACAGTGGTCCACACAAAGAAGCAGAACAGTATATCGCTCGCGTCTTTAAC GCAGACCGCAGCTACATGGTGACCAACGGTACTTCCACTGCGAACAAAATTGTTGGT ATGTACTCTGCTCCAGCAGGCAGCACCATTCTGATTGACCGTAACTGCCACAAATCG CTGACCCACCTGATGATGATGAGCGATGTTACGCCAATCTATTTCCGCCCGACCCGT AACGCTTACGGTATTCTTGGTGGTATCCCACAGAGTGAATTCCAGCACGCTACCATT GCTAAGCGCGTGAAAGAAACACCAAACGCAACCTGGCCGGTACATGCTGTAATTAC CAACTCTACCTATGATGGTCTGCTGTACAACACCGACTTCATCAAGAAAACACTGGA TGTGAAATCCATCCACTTTGACTCCGCGTGGGTGCCTTACACCAACTTCTCACCGATT TACGAAGGTAAATGCGGTATGAGCGGTGGCCGTGTAGAAGGGAAAGTGATTTACGA AACCCAGTCCACTCACAAACTGCTGGCGGCGTTCTCTCAGGCTTCCATGATCCACGT TAAAGGTGACGTAAACGAAGAAACCTTTAACGAAGCCTACATGATGCACACCACCA CTTCTCCGCACTACGGTATCGTGGCGTCCACTGAAACCGCTGCGGCGATGATGAAAG GCAATGCAGGTAAGCGTCTGATCAACGGTTCTATTGAACGTGCGATCAAATTCCGTA AAGAGATCAAACGTCTGAGAACGGAATCTGATGGCTGGTTCTTTGATGTATGGCAGC CGGATCATATCGATACGACTGAATGCTGGCCGCTGCGTTCTGACAGCACCTGGCACG GCTTCAAAAACATCGATAACGAGCACATGTATCTTGACCCGATCAAAGTCACCCTGC TGACTCCGGGGATGGAAAAAGACGGCACCATGAGCGACTTTGGTATTCCGGCCAGC ATCGTGGCGAAATACCTCGACGAACATGGCATCGTTGTTGAGAAAACCGGTCCGTAT AACCTGCTGTTCCTGTTCAGCATCGGTATCGATAAGACCAAAGCACTGAGCCTGCTG CGTGCTCTGACTGACTTTAAACGTGCGTTCGACCTGAACCTGCGTGTGAAAAACATG CTGCCGTCTCTGTATCGTGAAGATCCTGAATTCTATGAAAACATGCGTATTCAGGAA CTGGCTCAGAATATCCACAAACTGATTGTTCACCACAATCTGCCGGATCTGATGTAT CGCGCATTTGAAGTGCTGCCGACGATGGTAATGACTCCGTATGCTGCATTCCAGAAA GAGCTGCACGGTATGACCGAAGAAGTTTACCTCGACGAAATGGTAGGTCGTATTAA CGCCAATATGATCCTTCCGTACCCGCCGGGAGTTCCTCTGGTAATGCCGGGTGAAAT GATCACCGAAGAAAGCCGTCCGGTTCTGGAGTTCCTGCAGATGCTGTGTGAAATCGG CGCTCACTATCCGGGCTTTGAAACCGATATTCACGGTGCATACCGTCAGGCTGATGG CCGCTATACCGTTAAGGTATTGAAAGAAGAAAGCAAAAAATAA CadA polypeptide sequence SEQ ID NO: 2 MNVIAILNHMGVYFKEEPIRELHRALERLNFQIVYPNDRDDLLKLIENNARLCGVIFDWD KYNLELCEEISKMNENLPLYAFANTYSTLDVSLNDLRLQISFFEYALGAAEDIANKIKQT TDEYINTILPPLTKALFKYVREGKYTFCTPGHMGGTAFQKSPVGSLFYDFFGPNTMKSDI SISVSELGSLLDHSGPHKEAEQYIARVFNADRSYMVTNGTSTANKIVGMYSAPAGSTILI DRNCHKSLTHLMMMSDVTPIYFRPTRNAYGILGGIPQSEFQHATIAKRVKETPNATWPV HAVITNSTYDGLLYNTDFIKKTLDVKSIHFDSAWVPYTNFSPIYEGKCGMSGGRVEGKVI YETQSTHKLLAAFSQASMIFIVKGDVNEETFNEAYMMHTTTSPHYGIVASTETAAAMMK GNAGKRLINGSIERAIKFRKEIKRLRTESDGWFFDVWQPDHIDTTECWPLRSDSTWHGFK NIDNEHMYLDPIKVTLLTPGMEKDGTMSDFGIPASIVAKYLDEHGIVVEKTGPYNLLFLF SIGIDKTKALSLLRALTDFKRAFDLNLRVKNMLPSLYREDPEFYENMRIQELAQNIHKLI VHHNLPDLMYRAFEVLPTMVMTPYAAFQKELHGMTEEVYLDEMVGRINANMILPYPP GVPLVMPGEMITEESRPVLEFLQMLCEIGAHYPGFETDIHGAYRQA DGRYTVKVLKEESKK New1 prion subunit polypeptide sequence SEQ ID NO: 3 FPPKKFKDLNSFLDDQPKDPNLVASPFGGYFKNPAADAGSNNASKKSSYQQQRNWKQG GNYQQGGYQSYDSNYNNYNNYNNYNNYNNYNNYNKYNGQGYQ Sup35 prion subunit polypeptide sequence SEQ ID NO: 4 SDSNQGNNQQNYQQYSQNGNQQQGNNRYQGYQAYNAQAQPAGGYYQNYQGYSGYQ QGGYQQYNPQGGYQQDAGYQQQYNPQGGYQQYNPQGGYQQQFNPQGG New1 prion subunit nucleic acid sequence SEQ ID NO: 5 TTTCCGCCGAAAAAGTTCAAAGACCTGAACTCTTTCCTGGACGACCAGCCGAAAGA CCCGAACCTGGTTGCGTCTCCGTTCGGTGGCTACTTCAAAAACCCAGCGGCGGACGC GGGTTCTAACAACGCGTCTAAGAAATCTTCTTACCAGCAGCAGCGTAACTGGAAAC AGGGTGGCAACTATCAGCAAGGTGGTTACCAGTCTTACGACTCTAATTACAACAACT ACAACAACTACAATAACTATAATAACTACAACAACTACAACAATTATAACAAATAC AACGGTCAGGGCTACCAG Sup35 prion subunit nucleic acid sequence SEQ ID NO: 6 TCTGACTCTAACCAAGGTAATAACCAGCAGAACTACCAACAATACTCTCAGAACGG CAACCAGCAGCAGGGCAACAACCGCTATCAAGGCTACCAAGCGTACAACGCGCAGG CACAGCCAGCAGGTGGCTACTACCAGAATTACCAGGGTTACTCTGGTTACCAGCAA GGTGGTTATCAACAGTATAATCCGCAGGGCGGCTATCAGCAGGACGCAGGTTACCA GCAACAATATAACCCTCAGGGCGGCTATCAGCAATACAACCCGCAAGGCGGTTATC AACAACAGTTTAACCCACAGGGTGGC New1 prion subunitand linker polypeptide sequence. The linker sequence is underlined SEQ ID NO: 7 GSGSGFPPKKFKDLNSFLDDQPKDPNLVASPFGGYFKNPAADAGSNNASKKSSYQQQR NWKQGGNYQQGGYQSYDSNYNNYNNYNNYNNYNNYNNYNKYNGQGYQ Sup35 prion subunit and linker polypeptide sequence SEQ ID NO: 8 GSGSGSDSNQGNNQQNYQQYSQNGNQQQGNNRYQGYQAYNAQAQPAGGYYQNYQG YSGYQQGGYQQYNPQGGYQQDAGYQQQYNPQGGYQQYNPQGGYQQQFNPQGG New1 prion subunit and linker nucleic acid sequence. The region encoding the linker is underlined. SEQ ID NO: 9 GGTTCTGGCTCTGGTTTTCCGCCGAAAAAGTTCAAAGACCTGAACTCTTTCCTGGAC GACCAGCCGAAAGACCCGAACCTGGTTGCGTCTCCGTTCGGTGGCTACTTCAAAAAC CCAGCGGCGGACGCGGGTTCTAACAACGCGTCTAAGAAATCTTCTTACCAGCAGCA GCGTAACTGGAAACAGGGTGGCAACTATCAGCAAGGTGGTTACCAGTCTTACGACT CTAATTACAACAACTACAACAACTACAATAACTATAATAACTACAACAACTACAAC AATTATAACAAATACAACGGTCAGGGCTACCAG Sup35 prion subunit and linker nucleic acid sequence. The region encoding the linker is underlined. SEQ ID NO: 10 GGTAGCGGCTCTGGCTCTGACTCTAACCAAGGTAATAACCAGCAGAACTACCAACA ATACTCTCAGAACGGCAACCAGCAGCAGGGCAACAACCGCTATCAAGGCTACCAAG CGTACAACGCGCAGGCACAGCCAGCAGGTGGCTACTACCAGAATTACCAGGGTTAC TCTGGTTACCAGCAAGGTGGTTATCAACAGTATAATCCGCAGGGCGGCTATCAGCAG GACGCAGGTTACCAGCAACAATATAACCCTCAGGGCGGCTATCAGCAATACAACCC GCAAGGCGGTTATCAACAACAGTTTAACCCACAGGGTGGC New1 prion subunit and linker with 9 amino acid truncation polypeptide sequence. The linker is underlined. SEQ ID NO: 11 GSGSGNSFLDDQPKDPNLVASPFGGYFKNPAADAGSNNASKKSSYQQQRNWKQGGNY QQGGYQSYDSNYNNYNNYNNYNNYNNYNNYNKYNGQGYQ New1 prion subunit and linker with 18 amino acid truncation polypeptide sequence. The linker is underlined. SEQ ID NO: 12 GSGSGDPNLVASPFGGYFKNPAADAGSNNASKKSSYQQQRNWKQGGNYQQGGYQSY DSNYNNYNNYNNYNNYNNYNNYNKYNGQGYQ New1 prion subunit and linker with 36 amino acid truncation polypeptide sequence. The linker is underlined. SEQ ID NO: 13 GSGSGDAGSNNASKKSSYQQQRNWKQGGNYQQGGYQSYDSNYNNYNNYNNYNNYN NYNNYNKYNGQGYQ New1 prion subunit and linker with 45 amino acid truncation polypeptide sequence. The linker is underlined. SEQ ID NO: 14 GSGSGKSSYQQQRNWKQGGNYQQGGYQSYDSNYNNYNNYNNYNNYNNYNNYNKYN GQGYQ New1 prion domain fragment and linker with BST C-terminal fragment polypeptide sequence. The linker is underlined. SEQ ID NO: 15 GSGSGFPPKKFKDLNSFLDDQPKDPNLVASPFGGYFKNPAADAGSNNASKKSSYQQQR NWKQGGNYQQGGYQSYDSNYNNYNNYNNYNNYNNYNNYNKYNGQGYQRRFGEASS AF New1 prion subunit and linker with BST C-terminal fragment nucleic acid sequence. The region encoding the linker is underlined SEQ ID NO: 16 GGTTCTGGCTCTGGTTTTCCGCCGAAAAAGTTCAAAGACCTGAACTCTTTCCTGGAC GACCAGCCGAAAGACCCGAACCTGGTTGCGTCTCCGTTCGGTGGCTACTTCAAAAAC CCAGCGGCGGACGCGGGTTCTAACAACGCGTCTAAGAAATCTTCTTACCAGCAGCA GCGTAACTGGAAACAGGGTGGCAACTATCAGCAAGGTGGTTACCAGTCTTACGACT CTAATTACAACAACTACAACAACTACAATAACTATAATAACTACAACAACTACAAC AATTATAACAAATACAACGGTCAGGGCTACCAGCGTCGTTTCGGCGAAGCGAGCAG CGCGTTC New1 prion subunit and linker with AEI C-terminal fragment polypeptide sequence. The linker is underlined. SEQ ID NO: 17 GSGSGFPPKKFKDLNSFLDDQPKDPNLVASPFGGYFKNPAADAGSNNASKKSSYQQQR NWKQGGNYQQGGYQSYDSNYNNYNNYNNYNNYNNYNNYNKYNGQGYQASQWPEE TFG New1 prion domain fragment and linker with AEI C-terminal fragment nucleic acid sequence. The region encoding the linker is underlined. SEQ ID NO: 18 GGTTCTGGCTCTGGTTTTCCGCCGAAAAAGTTCAAAGACCTGAACTCTTTCCTGGAC GACCAGCCGAAAGACCCGAACCTGGTTGCGTCTCCGTTCGGTGGCTACTTCAAAAAC CCAGCGGCGGACGCGGGTTCTAACAACGCGTCTAAGAAATCTTCTTACCAGCAGCA GCGTAACTGGAAACAGGGTGGCAACTATCAGCAAGGTGGTTACCAGTCTTACGACT
CTAATTACAACAACTACAACAACTACAATAACTATAATAACTACAACAACTACAAC AATTATAACAAATACAACGGTCAGGGCTACCAGGCGAGCCAGTGGCCGGAAGAAAC CTTCGGC New1 prion domain fragment and linker with RecA C-terminal fragment polypeptide sequence. The linker is underlined. SEQ ID NO: 19 GSGSGFPPKKFKDLNSFLDDQPKDPNLVASPFGGYFKNPAADAGSNNASKKSSYQQQR NWKQGGNYQQGGYQSYDSNYNNYNNYNNYNNYNNYNNYNKYNGQGYQEGVAETN EDF New1 subunit and linker with RecA C-terminal fragment nucleic acid sequence. The region encoding the linker is underlined. SEQ ID NO: 20 GGTTCTGGCTCTGGTTTTCCGCCGAAAAAGTTCAAAGACCTGAACTCTTTCCTGGAC GACCAGCCGAAAGACCCGAACCTGGTTGCGTCTCCGTTCGGTGGCTACTTCAAAAAC CCAGCGGCGGACGCGGGTTCTAACAACGCGTCTAAGAAATCTTCTTACCAGCAGCA GCGTAACTGGAAACAGGGTGGCAACTATCAGCAAGGTGGTTACCAGTCTTACGACT CTAATTACAACAACTACAACAACTACAATAACTATAATAACTACAACAACTACAAC AATTATAACAAATACAACGGTCAGGGCTACCAGGAAGGCGTGGCGGAAACCAACGA AGATTTC LdcC polypeptide sequence SEQ ID NO: 21 MNIIAIMGPHGVFYKDEPIKELESALVAQGFQIIWPQNSVDLLKFIEHNPRICGVIFDWDE YSLDLCSDINQLNEYLPLYAFINTHSTMDVSVQDMRMALWFFEYALGQAEDIAIRMRQ YTDEYLDNITPPFTKALFTYVKERKYTFCTPGHMGGTAYQKSPVGCLFYDFFGGNTLKA DVSISVTELGSLLDHTGPHLEAEEYIARTFGAEQSYIVTNGTSTSNKIVGMYAAPSGSTLL IDRNCHKSLAHLLMMNDVVPVWLKPTRNALGILGGIPRREFTRDSIEEKVAATTQAQWP VHAVITNSTYDGLLYNTDWIKQTLDVPSIHFDSAWVPYTHFHPIYQGKSGMSGERVAGK VIFETQSTHKMLAALSQASLIHIKGEYDEEAFNEAFMMHTTTSPSYPIVASVETAAAMLR GNPGKRLINRSVERALHFRKEVQRLREESDGWFFDIWQPPQVDEAECWPVAPGEQWHG FNDADADHMFLDPVKVTILTPGMDEQGNMSEEGIPAALVAKFLDERGIVVEKTGPYNLL FLFSIGIDKTKAMGLLRGLTEFKRSYDLNLRIKNMLPDLYAEDPDFYRNMRIQDLAQGIH KLIRKHDLPGLMLRAFDTLPEMIMTPHQAWQRQIKGEVETIALEQLVGRVSANMILPYP PGVPLLMPGEMLTKESRTVLDFLLMLCSVGQHYPGFETDIHGAKQDEDGVYRVRVLKM AG AdiA polypeptide sequence SEQ ID NO: 22 MKVLIVESEFLHQDTWVGNAVERLADALSQQNVTVIKSTSFDDGFAILSSNEAIDCLMFS YQMEHPDEHQNVRQLIGKLHERQQNVPVFLLGDREKALAAMDRDLLELVDEFAWILED TADFIAGRAVAAMTRYRQQLLPPLFSALMKYSDIHEYSWAAPGHQGGVGFTKTPAGRF YHDYYGENLFRTDMGIERTSLGSLLDHTGAFGESEKYAARVFGADRSWSVVVGTSGSN RTIMQACMTDNDVVVVDRNCHKSIEQGLMLTGAKPVYMVPSRNRYGIIGPIYPQEMQP ETLQKKISESPLTKDKAGQKPSYCVVTNCTYDGVCYNAKEAQDLLEKTSDRLHFDEAW YGYARFNPIYADHYAMRGEPGDHNGPTVFATHSTHKLLNALSQASYIHVREGRGAINFS RFNQAYMMHATTSPLYAICASNDVAVSMMDGNSGLSLTQEVIDEAVDFRQAMARLYK EFTADGSWFFKPWNKEVVTDPQTGKTYDFADAPTKLLTTVQDCWVMHPGESWHGFKD IPDNWSMLDPIKVSILAPGMGEDGELEETGVPAALVTAWLGRHGIVPTRTTDFQIMFLFS MGVTRGKWGTLVNTLCSFKRHYDANTPLAQVMPELVEQYPDTYANMGIHDLGDTMF AWLKENNPGARLNEAYSGLPVAEVTPREAYNAIVDNNVELVSIENLPGRIAANSVIPYPP GIPMLLSGENFGDKNSPQVSYLRSLQSWDHHFPGFEHETEGTEIIDGIYHVMCVKA SpeA polypeptide sequence SEQ ID NO: 23 MSDDMSMGLPSSAGEHGVLRSMQEVAMSSQEASKMLRTYNIAWWGNNYYDVNELGH ISVCPDPDVPEARVDLAQLVKTREAQGQRLPALFCFPQILQHRLRSINAAFKRARESYGY NGDYFLVYPIKVNQHRRVIESLIHSGEPLGLEAGSKAELMAVLAHAGMTRSVIVCNGYK DREYIRLALIGEKMGHKVYLVIEKMSEIAIVLDEAERLNVVPRLGVRARLASQGSGKWQ SSGGEKSKFGLAATQVLQLVETLREAGRLDSLQLLHFHLGSQMANIRDIATGVRESARF YVELHKLGVNIQCFDVGGGLGVDYEGTRSQSDCSVNYGLNEYANNIIWAIGDACEENG WLPHPTVITESGRAVTAHHTVLVSNIIGVERNEYTVPTAPAEDAPRALQSMWETWQEMHE PGTRRSLREWLHDSQMDLHDIHIGYSSGIFSLQERAWAEQLYLSMCHEVQKQLDPQNR AHRPIIDELQERMADKMYVNFSLFQSMPDAWGIDQLFPVLPLEGLDQVPERRAVLLDIT CDSDGAIDHYIDGDGIATTMPMPEYDPENPPMLGFFMVGAYQEILGNMHNLFGDTEAV DVFVFPDGSVEVELSDEGDTVADMLQYVQLDPKTLLTQFRDQVKKTDLDAELQQQFLE EFEAGLYGYTYLEDE SpeC polypeptide sequence SEQ ID NO: 24 MKSMNIAASSELVSRLSSHRRVVALGDTDFTDVAAVVITAADSRSGILALLKRTGFHLP VFLYSEHAVELPAGVTAVINGNEQQWLELESAACQYEENLLPPFYDTLTQYVEMGNSTF ACPGHQHGAFFKKHPAGRHFYDFFGENVFRADMCNADVKLGDLLIHEGSAKDAQKFA AKVFHADKTYFVLNGTSAANKVVTNALLTRGDLVLFDRNNHKSNHHGALIQAGATPV YLEASRNPFGFIGGIDAHCFNEEYLRQQIRDVAPEKADLPRPYRLAIIQLGTYDGTVYNA RQVIDTVGHLCDYILFDSAWVGYEQFIPMMADSSPLLLELNENDPGIFVTQSVHKQQA GFSQTSQIHKKDNHIRGQARFCPHKRLNNAFMLHASTSPFYPLFAALDVNAKIHEGESG RRLWAECVEIGIEARKAILARCKLFRPFIPPVVDGKLWQDYPTSVLASDRRFFSFEPGAK WHGFEGYAADQYFVDPCKLLLTTPGIDAETGEYSDFGVPATILAHYLRENGIVPEKCDL NSILFLLTPAESHEKLAQLVAMLAQFEQHIEDDSPLVEVLPSVYNKYPVRYRDYTLRQLC QEMHDLYVSFDVKDLQKAMFRQQSFPSVVMNPQDAHSAYIRGDVELVRIRDAEGRIAA EGALPYPPGVLCVVPGEVWGGAVQRYFLALEEGVNLLPGFSPELQGVYSETDADGVKR LYGYVLK SpeF polypeptide sequence SEQ ID NO: 25 MSKLKIAVSDSCPDCFTTQRECIYINESRNIDVAAIVLSLNDVTCGKLDEIDATGYGIPVFI ATENQERVPAEYLPRISGVFENCESRREFYGRQLETAASHYETQLRPPFFRALVDYVNQ GNSAFDCPGHQGGEFFRRHPAGNQFVEYFGEALFRADLCNADVAMGDLLIHEGAPCIA QQHAAKVFNADKTYFVLNGTSSSNKVVLNALLTPGDLVLFDRNNHKSNHHGALLQAG ATPVYLETARNPYGFIGGIDAHCFEESYLRELIAEVAPQRAKEARPFRLAVIQLGTYDGTI YNARQVVDKIGHLCDYILFDSAWVGYEQFIPMMADCSPLLLDLNENDPGILVTQSVHKQ QAGFSQTSQIHKKDSHIKGQQRYVPHKRMNNAFMMHASTSPFYPLFAALNINAKMHEG VSGRNMWMDCVVNGINARKLILDNCQHIRPFVPELVDGKPWQSYETAQIAVDLRFFQF VPGEHWHSFEGYAENQYFVDPCKLLLTTPGIDARNGEYEAFGVPATILANFLRENGVVP EKCDLNSILFLLTPAEDMAKLQQLVALLVRFEKLLESDAPLAEVLPSIYKQHEERYAGYT LRQLCQEMHDLYARHNVKQLQKEMFRKEHFPRVSMNPQEANYAYLRGEVELVRLPDA GRIAAEGALPYPPGVLCVVPGEIWGGAVLRYFSALEEGINLLPGFAPELQGVYIEEHDG RKQVWCYVIKPRDAQSTLLKGEKL GadA polypeptide sequence SEQ ID NO: 26 MDQKLLTDFRSELLDSRFGAKAISTIAESKRFPLHEMRDDVAFQIINDELYLDGNARQNL ATFCQTWDDENVHKLMDLSINKNWIDKEEYPQSAAIDLRCVNMVADLWHAPAPKNGQ AVGTNTIGSSEACMLGGMAMKWRWRKRMEAAGKPTDKPNLVCGPVQICWHKFARY WDVELREIPMRPGQLFMDPKRMIEACDENTIGVVPTFGVTYTGNYEFPQPLHDALDKFQ ADTGIDIDMHIDAASGGFLAPFVAPDIVWDFRLPRVKSISASGHKFGLAPLGCGWVIWR DEEALPQELVFNVDYLGGQIGTFAINFSRPAGQVIAQYYEFLRLGREGYTKVQNASYQV AAYLADEIAKLGPYEFICTGRPDEGIPAVCFKLKDGEDPGYTLYDLSERLRLRGWQVPA FTLGGEATDIVVMRIMCRRGFEMDFAELLLEDYKASLKYLSDHPKLQGIAQQNSFKHT GadB polypeptide sequence SEQ ID NO: 27 MDKKQVTDLRSELLDSRFGAKSISTIAESKRFPLHEMRDDVAFQIINDELYLDGNARQNL ATFCQTWDDENVHKLMDLSINKNWIDKEEYPQSAAIDLRCVNMVADLWHAPAPKNGQ AVGTNTIGSSEACMLGGMAMKWRWRKRMEAAGKPTDKPNLVCGPVQICWHKFARY WDVELREIPMRPGQLFMDPKRMIEACDENTIGVVPTFGVTYTGNYEFPQPLHDALDKFQ ADTGIDIDMHIDAASGGFLAPFVAPDIVWDFRLPRVKSISASGHKFGLAPLGCGWVIWR DEEALPQELVFNVDYLGGQIGTFAINFSRPAGQVIAQYYEFLRLGREGYTKVQNASYQV AAYLADEIAKLGPYEFICTGRPDEGIPAVCFKLKDGEDPGYTLYDLSERLRLRGWQVPA FTLGGEATDIVVMRIMCRRGFEMDFAELLLEDYKASLKYLSDHPKLQGIAQQNSFKHT
Sequence CWU 1
1
6812148DNAEscherichia coli 1atgaacgtta ttgcaatatt gaatcacatg
ggggtttatt ttaaagaaga acccatccgt 60gaacttcatc gcgcgcttga acgtctgaac
ttccagattg tttacccgaa cgaccgtgac 120gacttattaa aactgatcga
aaacaatgcg cgtctgtgcg gcgttatttt tgactgggat 180aaatataatc
tcgagctgtg cgaagaaatt agcaaaatga acgagaacct gccgttgtac
240gcgttcgcta atacgtattc cactctcgat gtaagcctga atgacctgcg
tttacagatt 300agcttctttg aatatgcgct gggtgctgct gaagatattg
ctaataagat caagcagacc 360actgacgaat atatcaacac tattctgcct
ccgctgacta aagcactgtt taaatatgtt 420cgtgaaggta aatatacttt
ctgtactcct ggtcacatgg gcggtactgc attccagaaa 480agcccggtag
gtagcctgtt ctatgatttc tttggtccga ataccatgaa atctgatatt
540tccatttcag tatctgaact gggttctctg ctggatcaca gtggtccaca
caaagaagca 600gaacagtata tcgctcgcgt ctttaacgca gaccgcagct
acatggtgac caacggtact 660tccactgcga acaaaattgt tggtatgtac
tctgctccag caggcagcac cattctgatt 720gaccgtaact gccacaaatc
gctgacccac ctgatgatga tgagcgatgt tacgccaatc 780tatttccgcc
cgacccgtaa cgcttacggt attcttggtg gtatcccaca gagtgaattc
840cagcacgcta ccattgctaa gcgcgtgaaa gaaacaccaa acgcaacctg
gccggtacat 900gctgtaatta ccaactctac ctatgatggt ctgctgtaca
acaccgactt catcaagaaa 960acactggatg tgaaatccat ccactttgac
tccgcgtggg tgccttacac caacttctca 1020ccgatttacg aaggtaaatg
cggtatgagc ggtggccgtg tagaagggaa agtgatttac 1080gaaacccagt
ccactcacaa actgctggcg gcgttctctc aggcttccat gatccacgtt
1140aaaggtgacg taaacgaaga aacctttaac gaagcctaca tgatgcacac
caccacttct 1200ccgcactacg gtatcgtggc gtccactgaa accgctgcgg
cgatgatgaa aggcaatgca 1260ggtaagcgtc tgatcaacgg ttctattgaa
cgtgcgatca aattccgtaa agagatcaaa 1320cgtctgagaa cggaatctga
tggctggttc tttgatgtat ggcagccgga tcatatcgat 1380acgactgaat
gctggccgct gcgttctgac agcacctggc acggcttcaa aaacatcgat
1440aacgagcaca tgtatcttga cccgatcaaa gtcaccctgc tgactccggg
gatggaaaaa 1500gacggcacca tgagcgactt tggtattccg gccagcatcg
tggcgaaata cctcgacgaa 1560catggcatcg ttgttgagaa aaccggtccg
tataacctgc tgttcctgtt cagcatcggt 1620atcgataaga ccaaagcact
gagcctgctg cgtgctctga ctgactttaa acgtgcgttc 1680gacctgaacc
tgcgtgtgaa aaacatgctg ccgtctctgt atcgtgaaga tcctgaattc
1740tatgaaaaca tgcgtattca ggaactggct cagaatatcc acaaactgat
tgttcaccac 1800aatctgccgg atctgatgta tcgcgcattt gaagtgctgc
cgacgatggt aatgactccg 1860tatgctgcat tccagaaaga gctgcacggt
atgaccgaag aagtttacct cgacgaaatg 1920gtaggtcgta ttaacgccaa
tatgatcctt ccgtacccgc cgggagttcc tctggtaatg 1980ccgggtgaaa
tgatcaccga agaaagccgt ccggttctgg agttcctgca gatgctgtgt
2040gaaatcggcg ctcactatcc gggctttgaa accgatattc acggtgcata
ccgtcaggct 2100gatggccgct ataccgttaa ggtattgaaa gaagaaagca aaaaataa
21482715PRTEscherichia coli 2Met Asn Val Ile Ala Ile Leu Asn His
Met Gly Val Tyr Phe Lys Glu1 5 10 15Glu Pro Ile Arg Glu Leu His Arg
Ala Leu Glu Arg Leu Asn Phe Gln 20 25 30Ile Val Tyr Pro Asn Asp Arg
Asp Asp Leu Leu Lys Leu Ile Glu Asn 35 40 45Asn Ala Arg Leu Cys Gly
Val Ile Phe Asp Trp Asp Lys Tyr Asn Leu 50 55 60Glu Leu Cys Glu Glu
Ile Ser Lys Met Asn Glu Asn Leu Pro Leu Tyr65 70 75 80Ala Phe Ala
Asn Thr Tyr Ser Thr Leu Asp Val Ser Leu Asn Asp Leu 85 90 95Arg Leu
Gln Ile Ser Phe Phe Glu Tyr Ala Leu Gly Ala Ala Glu Asp 100 105
110Ile Ala Asn Lys Ile Lys Gln Thr Thr Asp Glu Tyr Ile Asn Thr Ile
115 120 125Leu Pro Pro Leu Thr Lys Ala Leu Phe Lys Tyr Val Arg Glu
Gly Lys 130 135 140Tyr Thr Phe Cys Thr Pro Gly His Met Gly Gly Thr
Ala Phe Gln Lys145 150 155 160Ser Pro Val Gly Ser Leu Phe Tyr Asp
Phe Phe Gly Pro Asn Thr Met 165 170 175Lys Ser Asp Ile Ser Ile Ser
Val Ser Glu Leu Gly Ser Leu Leu Asp 180 185 190His Ser Gly Pro His
Lys Glu Ala Glu Gln Tyr Ile Ala Arg Val Phe 195 200 205Asn Ala Asp
Arg Ser Tyr Met Val Thr Asn Gly Thr Ser Thr Ala Asn 210 215 220Lys
Ile Val Gly Met Tyr Ser Ala Pro Ala Gly Ser Thr Ile Leu Ile225 230
235 240Asp Arg Asn Cys His Lys Ser Leu Thr His Leu Met Met Met Ser
Asp 245 250 255Val Thr Pro Ile Tyr Phe Arg Pro Thr Arg Asn Ala Tyr
Gly Ile Leu 260 265 270Gly Gly Ile Pro Gln Ser Glu Phe Gln His Ala
Thr Ile Ala Lys Arg 275 280 285Val Lys Glu Thr Pro Asn Ala Thr Trp
Pro Val His Ala Val Ile Thr 290 295 300Asn Ser Thr Tyr Asp Gly Leu
Leu Tyr Asn Thr Asp Phe Ile Lys Lys305 310 315 320Thr Leu Asp Val
Lys Ser Ile His Phe Asp Ser Ala Trp Val Pro Tyr 325 330 335Thr Asn
Phe Ser Pro Ile Tyr Glu Gly Lys Cys Gly Met Ser Gly Gly 340 345
350Arg Val Glu Gly Lys Val Ile Tyr Glu Thr Gln Ser Thr His Lys Leu
355 360 365Leu Ala Ala Phe Ser Gln Ala Ser Met Ile His Val Lys Gly
Asp Val 370 375 380Asn Glu Glu Thr Phe Asn Glu Ala Tyr Met Met His
Thr Thr Thr Ser385 390 395 400Pro His Tyr Gly Ile Val Ala Ser Thr
Glu Thr Ala Ala Ala Met Met 405 410 415Lys Gly Asn Ala Gly Lys Arg
Leu Ile Asn Gly Ser Ile Glu Arg Ala 420 425 430Ile Lys Phe Arg Lys
Glu Ile Lys Arg Leu Arg Thr Glu Ser Asp Gly 435 440 445Trp Phe Phe
Asp Val Trp Gln Pro Asp His Ile Asp Thr Thr Glu Cys 450 455 460Trp
Pro Leu Arg Ser Asp Ser Thr Trp His Gly Phe Lys Asn Ile Asp465 470
475 480Asn Glu His Met Tyr Leu Asp Pro Ile Lys Val Thr Leu Leu Thr
Pro 485 490 495Gly Met Glu Lys Asp Gly Thr Met Ser Asp Phe Gly Ile
Pro Ala Ser 500 505 510Ile Val Ala Lys Tyr Leu Asp Glu His Gly Ile
Val Val Glu Lys Thr 515 520 525Gly Pro Tyr Asn Leu Leu Phe Leu Phe
Ser Ile Gly Ile Asp Lys Thr 530 535 540Lys Ala Leu Ser Leu Leu Arg
Ala Leu Thr Asp Phe Lys Arg Ala Phe545 550 555 560Asp Leu Asn Leu
Arg Val Lys Asn Met Leu Pro Ser Leu Tyr Arg Glu 565 570 575Asp Pro
Glu Phe Tyr Glu Asn Met Arg Ile Gln Glu Leu Ala Gln Asn 580 585
590Ile His Lys Leu Ile Val His His Asn Leu Pro Asp Leu Met Tyr Arg
595 600 605Ala Phe Glu Val Leu Pro Thr Met Val Met Thr Pro Tyr Ala
Ala Phe 610 615 620Gln Lys Glu Leu His Gly Met Thr Glu Glu Val Tyr
Leu Asp Glu Met625 630 635 640Val Gly Arg Ile Asn Ala Asn Met Ile
Leu Pro Tyr Pro Pro Gly Val 645 650 655Pro Leu Val Met Pro Gly Glu
Met Ile Thr Glu Glu Ser Arg Pro Val 660 665 670Leu Glu Phe Leu Gln
Met Leu Cys Glu Ile Gly Ala His Tyr Pro Gly 675 680 685Phe Glu Thr
Asp Ile His Gly Ala Tyr Arg Gln Ala Asp Gly Arg Tyr 690 695 700Thr
Val Lys Val Leu Lys Glu Glu Ser Lys Lys705 710
7153100PRTSaccharomyces cerevisiae 3Phe Pro Pro Lys Lys Phe Lys Asp
Leu Asn Ser Phe Leu Asp Asp Gln1 5 10 15Pro Lys Asp Pro Asn Leu Val
Ala Ser Pro Phe Gly Gly Tyr Phe Lys 20 25 30Asn Pro Ala Ala Asp Ala
Gly Ser Asn Asn Ala Ser Lys Lys Ser Ser 35 40 45Tyr Gln Gln Gln Arg
Asn Trp Lys Gln Gly Gly Asn Tyr Gln Gln Gly 50 55 60Gly Tyr Gln Ser
Tyr Asp Ser Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn65 70 75 80Asn Tyr
Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Lys Tyr Asn Gly 85 90 95Gln
Gly Tyr Gln 1004102PRTSaccharomyces cerevisiae 4Ser Asp Ser Asn Gln
Gly Asn Asn Gln Gln Asn Tyr Gln Gln Tyr Ser1 5 10 15Gln Asn Gly Asn
Gln Gln Gln Gly Asn Asn Arg Tyr Gln Gly Tyr Gln 20 25 30Ala Tyr Asn
Ala Gln Ala Gln Pro Ala Gly Gly Tyr Tyr Gln Asn Tyr 35 40 45Gln Gly
Tyr Ser Gly Tyr Gln Gln Gly Gly Tyr Gln Gln Tyr Asn Pro 50 55 60Gln
Gly Gly Tyr Gln Gln Asp Ala Gly Tyr Gln Gln Gln Tyr Asn Pro65 70 75
80Gln Gly Gly Tyr Gln Gln Tyr Asn Pro Gln Gly Gly Tyr Gln Gln Gln
85 90 95Phe Asn Pro Gln Gly Gly 1005300DNAArtificial SequenceNew1
prion subunit nucleic acid sequence.Codon optimized. 5tttccgccga
aaaagttcaa agacctgaac tctttcctgg acgaccagcc gaaagacccg 60aacctggttg
cgtctccgtt cggtggctac ttcaaaaacc cagcggcgga cgcgggttct
120aacaacgcgt ctaagaaatc ttcttaccag cagcagcgta actggaaaca
gggtggcaac 180tatcagcaag gtggttacca gtcttacgac tctaattaca
acaactacaa caactacaat 240aactataata actacaacaa ctacaacaat
tataacaaat acaacggtca gggctaccag 3006306DNAArtificial SequenceSup35
prion subunit nucleic acid sequence.Codon optimized 6tctgactcta
accaaggtaa taaccagcag aactaccaac aatactctca gaacggcaac 60cagcagcagg
gcaacaaccg ctatcaaggc taccaagcgt acaacgcgca ggcacagcca
120gcaggtggct actaccagaa ttaccagggt tactctggtt accagcaagg
tggttatcaa 180cagtataatc cgcagggcgg ctatcagcag gacgcaggtt
accagcaaca atataaccct 240cagggcggct atcagcaata caacccgcaa
ggcggttatc aacaacagtt taacccacag 300ggtggc 3067105PRTArtificial
SequenceNew1 prion subunit and linker polypeptide sequence. The
linker sequence is GSGSG(in the front of the sequence). 7Gly Ser
Gly Ser Gly Phe Pro Pro Lys Lys Phe Lys Asp Leu Asn Ser1 5 10 15Phe
Leu Asp Asp Gln Pro Lys Asp Pro Asn Leu Val Ala Ser Pro Phe 20 25
30Gly Gly Tyr Phe Lys Asn Pro Ala Ala Asp Ala Gly Ser Asn Asn Ala
35 40 45Ser Lys Lys Ser Ser Tyr Gln Gln Gln Arg Asn Trp Lys Gln Gly
Gly 50 55 60Asn Tyr Gln Gln Gly Gly Tyr Gln Ser Tyr Asp Ser Asn Tyr
Asn Asn65 70 75 80Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn
Tyr Asn Asn Tyr 85 90 95Asn Lys Tyr Asn Gly Gln Gly Tyr Gln 100
1058107PRTArtificial SequenceSup35 prion subunit and linker
polypeptide sequence.The linker polypeptide sequence is GSGSG(in
the front of the sequence) 8Gly Ser Gly Ser Gly Ser Asp Ser Asn Gln
Gly Asn Asn Gln Gln Asn1 5 10 15Tyr Gln Gln Tyr Ser Gln Asn Gly Asn
Gln Gln Gln Gly Asn Asn Arg 20 25 30Tyr Gln Gly Tyr Gln Ala Tyr Asn
Ala Gln Ala Gln Pro Ala Gly Gly 35 40 45Tyr Tyr Gln Asn Tyr Gln Gly
Tyr Ser Gly Tyr Gln Gln Gly Gly Tyr 50 55 60Gln Gln Tyr Asn Pro Gln
Gly Gly Tyr Gln Gln Asp Ala Gly Tyr Gln65 70 75 80Gln Gln Tyr Asn
Pro Gln Gly Gly Tyr Gln Gln Tyr Asn Pro Gln Gly 85 90 95Gly Tyr Gln
Gln Gln Phe Asn Pro Gln Gly Gly 100 1059315DNAArtificial
SequenceNew1 prion subunit and linker nucleic acid sequence.The
region encoding the linker is GGTTCTGGCTCTGGT(in the front of
sequence). 9ggttctggct ctggttttcc gccgaaaaag ttcaaagacc tgaactcttt
cctggacgac 60cagccgaaag acccgaacct ggttgcgtct ccgttcggtg gctacttcaa
aaacccagcg 120gcggacgcgg gttctaacaa cgcgtctaag aaatcttctt
accagcagca gcgtaactgg 180aaacagggtg gcaactatca gcaaggtggt
taccagtctt acgactctaa ttacaacaac 240tacaacaact acaataacta
taataactac aacaactaca acaattataa caaatacaac 300ggtcagggct accag
31510321DNAArtificial SequenceSup35 prion subunit and linker
nucleic acid sequence.The region encoding the linker is
GGTAGCGGCTCTGGC(in the front of the sequence). 10ggtagcggct
ctggctctga ctctaaccaa ggtaataacc agcagaacta ccaacaatac 60tctcagaacg
gcaaccagca gcagggcaac aaccgctatc aaggctacca agcgtacaac
120gcgcaggcac agccagcagg tggctactac cagaattacc agggttactc
tggttaccag 180caaggtggtt atcaacagta taatccgcag ggcggctatc
agcaggacgc aggttaccag 240caacaatata accctcaggg cggctatcag
caatacaacc cgcaaggcgg ttatcaacaa 300cagtttaacc cacagggtgg c
3211196PRTArtificial SequenceNew1 prion subunit and linker with 9
amino acid truncation polypeptide sequence.The linker is GSGSG(in
the front of the sequence). 11Gly Ser Gly Ser Gly Asn Ser Phe Leu
Asp Asp Gln Pro Lys Asp Pro1 5 10 15Asn Leu Val Ala Ser Pro Phe Gly
Gly Tyr Phe Lys Asn Pro Ala Ala 20 25 30Asp Ala Gly Ser Asn Asn Ala
Ser Lys Lys Ser Ser Tyr Gln Gln Gln 35 40 45Arg Asn Trp Lys Gln Gly
Gly Asn Tyr Gln Gln Gly Gly Tyr Gln Ser 50 55 60Tyr Asp Ser Asn Tyr
Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn65 70 75 80Tyr Asn Asn
Tyr Asn Asn Tyr Asn Lys Tyr Asn Gly Gln Gly Tyr Gln 85 90
951287PRTArtificial SequenceNew1 prion subunit and linker with 18
amino acid truncation polypeptide sequence.The linker is GSGSG(in
the front of the sequence). 12Gly Ser Gly Ser Gly Asp Pro Asn Leu
Val Ala Ser Pro Phe Gly Gly1 5 10 15Tyr Phe Lys Asn Pro Ala Ala Asp
Ala Gly Ser Asn Asn Ala Ser Lys 20 25 30Lys Ser Ser Tyr Gln Gln Gln
Arg Asn Trp Lys Gln Gly Gly Asn Tyr 35 40 45Gln Gln Gly Gly Tyr Gln
Ser Tyr Asp Ser Asn Tyr Asn Asn Tyr Asn 50 55 60Asn Tyr Asn Asn Tyr
Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Lys65 70 75 80Tyr Asn Gly
Gln Gly Tyr Gln 851369PRTArtificial SequenceNew1 prion subunit and
linker with 36 amino acid truncation polypeptide sequence.The
linker is GSGSG(in the front of the sequence). 13Gly Ser Gly Ser
Gly Asp Ala Gly Ser Asn Asn Ala Ser Lys Lys Ser1 5 10 15Ser Tyr Gln
Gln Gln Arg Asn Trp Lys Gln Gly Gly Asn Tyr Gln Gln 20 25 30Gly Gly
Tyr Gln Ser Tyr Asp Ser Asn Tyr Asn Asn Tyr Asn Asn Tyr 35 40 45Asn
Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Lys Tyr Asn 50 55
60Gly Gln Gly Tyr Gln651460PRTArtificial SequenceNew1 prion subunit
and linker with 45 amino acid truncation polypeptide sequence.The
linker is GSGSG(in the front of the sequence). 14Gly Ser Gly Ser
Gly Lys Ser Ser Tyr Gln Gln Gln Arg Asn Trp Lys1 5 10 15Gln Gly Gly
Asn Tyr Gln Gln Gly Gly Tyr Gln Ser Tyr Asp Ser Asn 20 25 30Tyr Asn
Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr 35 40 45Asn
Asn Tyr Asn Lys Tyr Asn Gly Gln Gly Tyr Gln 50 55
6015115PRTArtificial SequenceNew1 prion domain fragment and linker
with BST C-terminal fragment polypeptide sequence.The linker is
GSGSG(in the front of the sequence). 15Gly Ser Gly Ser Gly Phe Pro
Pro Lys Lys Phe Lys Asp Leu Asn Ser1 5 10 15Phe Leu Asp Asp Gln Pro
Lys Asp Pro Asn Leu Val Ala Ser Pro Phe 20 25 30Gly Gly Tyr Phe Lys
Asn Pro Ala Ala Asp Ala Gly Ser Asn Asn Ala 35 40 45Ser Lys Lys Ser
Ser Tyr Gln Gln Gln Arg Asn Trp Lys Gln Gly Gly 50 55 60Asn Tyr Gln
Gln Gly Gly Tyr Gln Ser Tyr Asp Ser Asn Tyr Asn Asn65 70 75 80Tyr
Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr 85 90
95Asn Lys Tyr Asn Gly Gln Gly Tyr Gln Arg Arg Phe Gly Glu Ala Ser
100 105 110Ser Ala Phe 11516345DNAArtificial SequenceNew1 prion
subunit and linker with BST C-terminal fragment nucleic acid
sequence.The region encoding the linker is GGTTCTGGCTCTGGT(in the
front of the sequence). 16ggttctggct ctggttttcc gccgaaaaag
ttcaaagacc tgaactcttt cctggacgac 60cagccgaaag acccgaacct ggttgcgtct
ccgttcggtg gctacttcaa aaacccagcg 120gcggacgcgg gttctaacaa
cgcgtctaag aaatcttctt accagcagca gcgtaactgg 180aaacagggtg
gcaactatca gcaaggtggt taccagtctt acgactctaa ttacaacaac
240tacaacaact acaataacta taataactac aacaactaca acaattataa
caaatacaac 300ggtcagggct accagcgtcg tttcggcgaa gcgagcagcg cgttc
34517115PRTArtificial SequenceNew1 prion subunit and linker with
??CI C-terminal fragment polypeptide sequence.The linker is
GSGSG(in the front of the sequence). 17Gly Ser Gly Ser Gly Phe Pro
Pro Lys Lys Phe Lys Asp Leu Asn Ser1 5 10 15Phe Leu Asp Asp Gln Pro
Lys Asp Pro Asn Leu Val Ala Ser Pro Phe 20 25 30Gly Gly Tyr Phe Lys
Asn Pro Ala Ala Asp Ala Gly Ser Asn Asn Ala 35 40 45Ser Lys Lys Ser
Ser Tyr Gln Gln Gln Arg Asn Trp Lys Gln Gly Gly 50 55 60Asn Tyr Gln
Gln Gly Gly Tyr Gln Ser Tyr Asp Ser Asn Tyr Asn Asn65 70 75 80Tyr
Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr 85 90
95Asn Lys Tyr Asn Gly Gln Gly Tyr Gln Ala Ser Gln Trp Pro Glu Glu
100 105 110Thr Phe Gly 11518345DNAArtificial SequenceNew1 prion
domain fragment and linker with ??CI C-terminal fragment nucleic
acid sequence.The region encoding the linker is GGTTCTGGCTCTGGT(in
the front of the sequence). 18ggttctggct ctggttttcc gccgaaaaag
ttcaaagacc tgaactcttt cctggacgac 60cagccgaaag acccgaacct ggttgcgtct
ccgttcggtg gctacttcaa aaacccagcg 120gcggacgcgg gttctaacaa
cgcgtctaag aaatcttctt accagcagca gcgtaactgg 180aaacagggtg
gcaactatca gcaaggtggt taccagtctt acgactctaa ttacaacaac
240tacaacaact acaataacta taataactac aacaactaca acaattataa
caaatacaac 300ggtcagggct accaggcgag ccagtggccg gaagaaacct tcggc
34519115PRTArtificial SequenceNew1 prion domain fragment and linker
with RecA C-terminal fragment polypeptide sequence.The linker is
GSGSG(in the front of the sequence). 19Gly Ser Gly Ser Gly Phe Pro
Pro Lys Lys Phe Lys Asp Leu Asn Ser1 5 10 15Phe Leu Asp Asp Gln Pro
Lys Asp Pro Asn Leu Val Ala Ser Pro Phe 20 25 30Gly Gly Tyr Phe Lys
Asn Pro Ala Ala Asp Ala Gly Ser Asn Asn Ala 35 40 45Ser Lys Lys Ser
Ser Tyr Gln Gln Gln Arg Asn Trp Lys Gln Gly Gly 50 55 60Asn Tyr Gln
Gln Gly Gly Tyr Gln Ser Tyr Asp Ser Asn Tyr Asn Asn65 70 75 80Tyr
Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr 85 90
95Asn Lys Tyr Asn Gly Gln Gly Tyr Gln Glu Gly Val Ala Glu Thr Asn
100 105 110Glu Asp Phe 11520345DNAArtificial SequenceNew1 subunit
and linker with RecA C-terminal fragment nucleic acid sequence.The
region encoding the linker is GGTTCTGGCTCTGGT(in the front of the
sequence). 20ggttctggct ctggttttcc gccgaaaaag ttcaaagacc tgaactcttt
cctggacgac 60cagccgaaag acccgaacct ggttgcgtct ccgttcggtg gctacttcaa
aaacccagcg 120gcggacgcgg gttctaacaa cgcgtctaag aaatcttctt
accagcagca gcgtaactgg 180aaacagggtg gcaactatca gcaaggtggt
taccagtctt acgactctaa ttacaacaac 240tacaacaact acaataacta
taataactac aacaactaca acaattataa caaatacaac 300ggtcagggct
accaggaagg cgtggcggaa accaacgaag atttc 34521713PRTEscherichia coli
21Met Asn Ile Ile Ala Ile Met Gly Pro His Gly Val Phe Tyr Lys Asp1
5 10 15Glu Pro Ile Lys Glu Leu Glu Ser Ala Leu Val Ala Gln Gly Phe
Gln 20 25 30Ile Ile Trp Pro Gln Asn Ser Val Asp Leu Leu Lys Phe Ile
Glu His 35 40 45Asn Pro Arg Ile Cys Gly Val Ile Phe Asp Trp Asp Glu
Tyr Ser Leu 50 55 60Asp Leu Cys Ser Asp Ile Asn Gln Leu Asn Glu Tyr
Leu Pro Leu Tyr65 70 75 80Ala Phe Ile Asn Thr His Ser Thr Met Asp
Val Ser Val Gln Asp Met 85 90 95Arg Met Ala Leu Trp Phe Phe Glu Tyr
Ala Leu Gly Gln Ala Glu Asp 100 105 110Ile Ala Ile Arg Met Arg Gln
Tyr Thr Asp Glu Tyr Leu Asp Asn Ile 115 120 125Thr Pro Pro Phe Thr
Lys Ala Leu Phe Thr Tyr Val Lys Glu Arg Lys 130 135 140Tyr Thr Phe
Cys Thr Pro Gly His Met Gly Gly Thr Ala Tyr Gln Lys145 150 155
160Ser Pro Val Gly Cys Leu Phe Tyr Asp Phe Phe Gly Gly Asn Thr Leu
165 170 175Lys Ala Asp Val Ser Ile Ser Val Thr Glu Leu Gly Ser Leu
Leu Asp 180 185 190His Thr Gly Pro His Leu Glu Ala Glu Glu Tyr Ile
Ala Arg Thr Phe 195 200 205Gly Ala Glu Gln Ser Tyr Ile Val Thr Asn
Gly Thr Ser Thr Ser Asn 210 215 220Lys Ile Val Gly Met Tyr Ala Ala
Pro Ser Gly Ser Thr Leu Leu Ile225 230 235 240Asp Arg Asn Cys His
Lys Ser Leu Ala His Leu Leu Met Met Asn Asp 245 250 255Val Val Pro
Val Trp Leu Lys Pro Thr Arg Asn Ala Leu Gly Ile Leu 260 265 270Gly
Gly Ile Pro Arg Arg Glu Phe Thr Arg Asp Ser Ile Glu Glu Lys 275 280
285Val Ala Ala Thr Thr Gln Ala Gln Trp Pro Val His Ala Val Ile Thr
290 295 300Asn Ser Thr Tyr Asp Gly Leu Leu Tyr Asn Thr Asp Trp Ile
Lys Gln305 310 315 320Thr Leu Asp Val Pro Ser Ile His Phe Asp Ser
Ala Trp Val Pro Tyr 325 330 335Thr His Phe His Pro Ile Tyr Gln Gly
Lys Ser Gly Met Ser Gly Glu 340 345 350Arg Val Ala Gly Lys Val Ile
Phe Glu Thr Gln Ser Thr His Lys Met 355 360 365Leu Ala Ala Leu Ser
Gln Ala Ser Leu Ile His Ile Lys Gly Glu Tyr 370 375 380Asp Glu Glu
Ala Phe Asn Glu Ala Phe Met Met His Thr Thr Thr Ser385 390 395
400Pro Ser Tyr Pro Ile Val Ala Ser Val Glu Thr Ala Ala Ala Met Leu
405 410 415Arg Gly Asn Pro Gly Lys Arg Leu Ile Asn Arg Ser Val Glu
Arg Ala 420 425 430Leu His Phe Arg Lys Glu Val Gln Arg Leu Arg Glu
Glu Ser Asp Gly 435 440 445Trp Phe Phe Asp Ile Trp Gln Pro Pro Gln
Val Asp Glu Ala Glu Cys 450 455 460Trp Pro Val Ala Pro Gly Glu Gln
Trp His Gly Phe Asn Asp Ala Asp465 470 475 480Ala Asp His Met Phe
Leu Asp Pro Val Lys Val Thr Ile Leu Thr Pro 485 490 495Gly Met Asp
Glu Gln Gly Asn Met Ser Glu Glu Gly Ile Pro Ala Ala 500 505 510Leu
Val Ala Lys Phe Leu Asp Glu Arg Gly Ile Val Val Glu Lys Thr 515 520
525Gly Pro Tyr Asn Leu Leu Phe Leu Phe Ser Ile Gly Ile Asp Lys Thr
530 535 540Lys Ala Met Gly Leu Leu Arg Gly Leu Thr Glu Phe Lys Arg
Ser Tyr545 550 555 560Asp Leu Asn Leu Arg Ile Lys Asn Met Leu Pro
Asp Leu Tyr Ala Glu 565 570 575Asp Pro Asp Phe Tyr Arg Asn Met Arg
Ile Gln Asp Leu Ala Gln Gly 580 585 590Ile His Lys Leu Ile Arg Lys
His Asp Leu Pro Gly Leu Met Leu Arg 595 600 605Ala Phe Asp Thr Leu
Pro Glu Met Ile Met Thr Pro His Gln Ala Trp 610 615 620Gln Arg Gln
Ile Lys Gly Glu Val Glu Thr Ile Ala Leu Glu Gln Leu625 630 635
640Val Gly Arg Val Ser Ala Asn Met Ile Leu Pro Tyr Pro Pro Gly Val
645 650 655Pro Leu Leu Met Pro Gly Glu Met Leu Thr Lys Glu Ser Arg
Thr Val 660 665 670Leu Asp Phe Leu Leu Met Leu Cys Ser Val Gly Gln
His Tyr Pro Gly 675 680 685Phe Glu Thr Asp Ile His Gly Ala Lys Gln
Asp Glu Asp Gly Val Tyr 690 695 700Arg Val Arg Val Leu Lys Met Ala
Gly705 71022755PRTEscherichia coli 22Met Lys Val Leu Ile Val Glu
Ser Glu Phe Leu His Gln Asp Thr Trp1 5 10 15Val Gly Asn Ala Val Glu
Arg Leu Ala Asp Ala Leu Ser Gln Gln Asn 20 25 30Val Thr Val Ile Lys
Ser Thr Ser Phe Asp Asp Gly Phe Ala Ile Leu 35 40 45Ser Ser Asn Glu
Ala Ile Asp Cys Leu Met Phe Ser Tyr Gln Met Glu 50 55 60His Pro Asp
Glu His Gln Asn Val Arg Gln Leu Ile Gly Lys Leu His65 70 75 80Glu
Arg Gln Gln Asn Val Pro Val Phe Leu Leu Gly Asp Arg Glu Lys 85 90
95Ala Leu Ala Ala Met Asp Arg Asp Leu Leu Glu Leu Val Asp Glu Phe
100 105 110Ala Trp Ile Leu Glu Asp Thr Ala Asp Phe Ile Ala Gly Arg
Ala Val 115 120 125Ala Ala Met Thr Arg Tyr Arg Gln Gln Leu Leu Pro
Pro Leu Phe Ser 130 135 140Ala Leu Met Lys Tyr Ser Asp Ile His Glu
Tyr Ser Trp Ala Ala Pro145 150 155 160Gly His Gln Gly Gly Val Gly
Phe Thr Lys Thr Pro Ala Gly Arg Phe 165 170 175Tyr His Asp Tyr Tyr
Gly Glu Asn Leu Phe Arg Thr Asp Met Gly Ile 180 185 190Glu Arg Thr
Ser Leu Gly Ser Leu Leu Asp His Thr Gly Ala Phe Gly 195 200 205Glu
Ser Glu Lys Tyr Ala Ala Arg Val Phe Gly Ala Asp Arg Ser Trp 210 215
220Ser Val Val Val Gly Thr Ser Gly Ser Asn Arg Thr Ile Met Gln
Ala225 230 235 240Cys Met Thr Asp Asn Asp Val Val Val Val Asp Arg
Asn Cys His Lys 245 250 255Ser Ile Glu Gln Gly Leu Met Leu Thr Gly
Ala Lys Pro Val Tyr Met 260 265 270Val Pro Ser Arg Asn Arg Tyr Gly
Ile Ile Gly Pro Ile Tyr Pro Gln 275 280 285Glu Met Gln Pro Glu Thr
Leu Gln Lys Lys Ile Ser Glu Ser Pro Leu 290 295 300Thr Lys Asp Lys
Ala Gly Gln Lys Pro Ser Tyr Cys Val Val Thr Asn305 310 315 320Cys
Thr Tyr Asp Gly Val Cys Tyr Asn Ala Lys Glu Ala Gln Asp Leu 325 330
335Leu Glu Lys Thr Ser Asp Arg Leu His Phe Asp Glu Ala Trp Tyr Gly
340 345 350Tyr Ala Arg Phe Asn Pro Ile Tyr Ala Asp His Tyr Ala Met
Arg Gly 355 360 365Glu Pro Gly Asp His Asn Gly Pro Thr Val Phe Ala
Thr His Ser Thr 370 375 380His Lys Leu Leu Asn Ala Leu Ser Gln Ala
Ser Tyr Ile His Val Arg385 390 395 400Glu Gly Arg Gly Ala Ile Asn
Phe Ser Arg Phe Asn Gln Ala Tyr Met 405 410 415Met His Ala Thr Thr
Ser Pro Leu Tyr Ala Ile Cys Ala Ser Asn Asp 420 425 430Val Ala Val
Ser Met Met Asp Gly Asn Ser Gly Leu Ser Leu Thr Gln 435 440 445Glu
Val Ile Asp Glu Ala Val Asp Phe Arg Gln Ala Met Ala Arg Leu 450 455
460Tyr Lys Glu Phe Thr Ala Asp Gly Ser Trp Phe Phe Lys Pro Trp
Asn465 470 475 480Lys Glu Val Val Thr Asp Pro Gln Thr Gly Lys Thr
Tyr Asp Phe Ala 485 490 495Asp Ala Pro Thr Lys Leu Leu Thr Thr Val
Gln Asp Cys Trp Val Met 500 505 510His Pro Gly Glu Ser Trp His Gly
Phe Lys Asp Ile Pro Asp Asn Trp 515 520 525Ser Met Leu Asp Pro Ile
Lys Val Ser Ile Leu Ala Pro Gly Met Gly 530 535 540Glu Asp Gly Glu
Leu Glu Glu Thr Gly Val Pro Ala Ala Leu Val Thr545 550 555 560Ala
Trp Leu Gly Arg His Gly Ile Val Pro Thr Arg Thr Thr Asp Phe 565 570
575Gln Ile Met Phe Leu Phe Ser Met Gly Val Thr Arg Gly Lys Trp Gly
580 585 590Thr Leu Val Asn Thr Leu Cys Ser Phe Lys Arg His Tyr Asp
Ala Asn 595 600 605Thr Pro Leu Ala Gln Val Met Pro Glu Leu Val Glu
Gln Tyr Pro Asp 610 615 620Thr Tyr Ala Asn Met Gly Ile His Asp Leu
Gly Asp Thr Met Phe Ala625 630 635 640Trp Leu Lys Glu Asn Asn Pro
Gly Ala Arg Leu Asn Glu Ala Tyr Ser 645 650 655Gly Leu Pro Val Ala
Glu Val Thr Pro Arg Glu Ala Tyr Asn Ala Ile 660 665 670Val Asp Asn
Asn Val Glu Leu Val Ser Ile Glu Asn Leu Pro Gly Arg 675 680 685Ile
Ala Ala Asn Ser Val Ile Pro Tyr Pro Pro Gly Ile Pro Met Leu 690 695
700Leu Ser Gly Glu Asn Phe Gly Asp Lys Asn Ser Pro Gln Val Ser
Tyr705 710 715 720Leu Arg Ser Leu Gln Ser Trp Asp His His Phe Pro
Gly Phe Glu His 725 730 735Glu Thr Glu Gly Thr Glu Ile Ile Asp Gly
Ile Tyr His Val Met Cys 740 745 750Val Lys Ala
75523658PRTEscherichia coli 23Met Ser Asp Asp Met Ser Met Gly Leu
Pro Ser Ser Ala Gly Glu His1 5 10 15Gly Val Leu Arg Ser Met Gln Glu
Val Ala Met Ser Ser Gln Glu Ala 20 25 30Ser Lys Met Leu Arg Thr Tyr
Asn Ile Ala Trp Trp Gly Asn Asn Tyr 35 40 45Tyr Asp Val Asn Glu Leu
Gly His Ile Ser Val Cys Pro Asp Pro Asp 50 55 60Val Pro Glu Ala Arg
Val Asp Leu Ala Gln Leu Val Lys Thr Arg Glu65 70 75 80Ala Gln Gly
Gln Arg Leu Pro Ala Leu Phe Cys Phe Pro Gln Ile Leu 85 90 95Gln His
Arg Leu Arg Ser Ile Asn Ala Ala Phe Lys Arg Ala Arg Glu 100 105
110Ser Tyr Gly Tyr Asn Gly Asp Tyr Phe Leu Val Tyr Pro Ile Lys Val
115 120 125Asn Gln His Arg Arg Val Ile Glu Ser Leu Ile His Ser Gly
Glu Pro 130 135 140Leu Gly Leu Glu Ala Gly Ser Lys Ala Glu Leu Met
Ala Val Leu Ala145 150 155 160His Ala Gly Met Thr Arg Ser Val Ile
Val Cys Asn Gly Tyr Lys Asp 165 170 175Arg Glu Tyr Ile Arg Leu Ala
Leu Ile Gly Glu Lys Met Gly His Lys 180 185 190Val Tyr Leu Val Ile
Glu Lys Met Ser Glu Ile Ala Ile Val Leu Asp 195 200 205Glu Ala Glu
Arg Leu Asn Val Val Pro Arg Leu Gly Val Arg Ala Arg 210 215 220Leu
Ala Ser Gln Gly Ser Gly Lys Trp Gln Ser Ser Gly Gly Glu Lys225 230
235 240Ser Lys Phe Gly Leu Ala Ala Thr Gln Val Leu Gln Leu Val Glu
Thr 245 250 255Leu Arg Glu Ala Gly Arg Leu Asp Ser Leu Gln Leu Leu
His Phe His 260 265 270Leu Gly Ser Gln Met Ala Asn Ile Arg Asp Ile
Ala Thr Gly Val Arg 275 280 285Glu Ser Ala Arg Phe Tyr Val Glu Leu
His Lys Leu Gly Val Asn Ile 290 295 300Gln Cys Phe Asp Val Gly Gly
Gly Leu Gly Val Asp Tyr Glu Gly Thr305 310 315 320Arg Ser Gln Ser
Asp Cys Ser Val Asn Tyr Gly Leu Asn Glu Tyr Ala 325 330 335Asn Asn
Ile Ile Trp Ala Ile Gly Asp Ala Cys Glu Glu Asn Gly Leu 340 345
350Pro His Pro Thr Val Ile Thr Glu Ser Gly Arg Ala Val Thr Ala His
355 360 365His Thr Val Leu Val Ser Asn Ile Ile Gly Val Glu Arg Asn
Glu Tyr 370 375 380Thr Val Pro Thr Ala Pro Ala Glu Asp Ala Pro Arg
Ala Leu Gln Ser385 390 395 400Met Trp Glu Thr Trp Gln Glu Met His
Glu Pro Gly Thr Arg Arg Ser 405 410 415Leu Arg Glu Trp Leu His Asp
Ser Gln Met Asp Leu His Asp Ile His 420 425 430Ile Gly Tyr Ser Ser
Gly Ile Phe Ser Leu Gln Glu Arg Ala Trp Ala 435 440 445Glu Gln Leu
Tyr Leu Ser Met Cys His Glu Val Gln Lys Gln Leu Asp 450 455 460Pro
Gln Asn Arg Ala His Arg Pro Ile Ile Asp Glu Leu Gln Glu Arg465 470
475 480Met Ala Asp Lys Met Tyr Val Asn Phe Ser Leu Phe Gln Ser Met
Pro 485 490 495Asp Ala Trp Gly Ile Asp Gln Leu Phe Pro Val Leu Pro
Leu Glu Gly 500 505 510Leu Asp Gln Val Pro Glu Arg Arg Ala Val Leu
Leu Asp Ile Thr Cys 515 520 525Asp Ser Asp Gly Ala Ile Asp His Tyr
Ile Asp Gly Asp Gly Ile Ala 530 535 540Thr Thr Met Pro Met
Pro Glu Tyr Asp Pro Glu Asn Pro Pro Met Leu545 550 555 560Gly Phe
Phe Met Val Gly Ala Tyr Gln Glu Ile Leu Gly Asn Met His 565 570
575Asn Leu Phe Gly Asp Thr Glu Ala Val Asp Val Phe Val Phe Pro Asp
580 585 590Gly Ser Val Glu Val Glu Leu Ser Asp Glu Gly Asp Thr Val
Ala Asp 595 600 605Met Leu Gln Tyr Val Gln Leu Asp Pro Lys Thr Leu
Leu Thr Gln Phe 610 615 620Arg Asp Gln Val Lys Lys Thr Asp Leu Asp
Ala Glu Leu Gln Gln Gln625 630 635 640Phe Leu Glu Glu Phe Glu Ala
Gly Leu Tyr Gly Tyr Thr Tyr Leu Glu 645 650 655Asp
Glu24711PRTEscherichia coli 24Met Lys Ser Met Asn Ile Ala Ala Ser
Ser Glu Leu Val Ser Arg Leu1 5 10 15Ser Ser His Arg Arg Val Val Ala
Leu Gly Asp Thr Asp Phe Thr Asp 20 25 30Val Ala Ala Val Val Ile Thr
Ala Ala Asp Ser Arg Ser Gly Ile Leu 35 40 45Ala Leu Leu Lys Arg Thr
Gly Phe His Leu Pro Val Phe Leu Tyr Ser 50 55 60Glu His Ala Val Glu
Leu Pro Ala Gly Val Thr Ala Val Ile Asn Gly65 70 75 80Asn Glu Gln
Gln Trp Leu Glu Leu Glu Ser Ala Ala Cys Gln Tyr Glu 85 90 95Glu Asn
Leu Leu Pro Pro Phe Tyr Asp Thr Leu Thr Gln Tyr Val Glu 100 105
110Met Gly Asn Ser Thr Phe Ala Cys Pro Gly His Gln His Gly Ala Phe
115 120 125Phe Lys Lys His Pro Ala Gly Arg His Phe Tyr Asp Phe Phe
Gly Glu 130 135 140Asn Val Phe Arg Ala Asp Met Cys Asn Ala Asp Val
Lys Leu Gly Asp145 150 155 160Leu Leu Ile His Glu Gly Ser Ala Lys
Asp Ala Gln Lys Phe Ala Ala 165 170 175Lys Val Phe His Ala Asp Lys
Thr Tyr Phe Val Leu Asn Gly Thr Ser 180 185 190Ala Ala Asn Lys Val
Val Thr Asn Ala Leu Leu Thr Arg Gly Asp Leu 195 200 205Val Leu Phe
Asp Arg Asn Asn His Lys Ser Asn His His Gly Ala Leu 210 215 220Ile
Gln Ala Gly Ala Thr Pro Val Tyr Leu Glu Ala Ser Arg Asn Pro225 230
235 240Phe Gly Phe Ile Gly Gly Ile Asp Ala His Cys Phe Asn Glu Glu
Tyr 245 250 255Leu Arg Gln Gln Ile Arg Asp Val Ala Pro Glu Lys Ala
Asp Leu Pro 260 265 270Arg Pro Tyr Arg Leu Ala Ile Ile Gln Leu Gly
Thr Tyr Asp Gly Thr 275 280 285Val Tyr Asn Ala Arg Gln Val Ile Asp
Thr Val Gly His Leu Cys Asp 290 295 300Tyr Ile Leu Phe Asp Ser Ala
Trp Val Gly Tyr Glu Gln Phe Ile Pro305 310 315 320Met Met Ala Asp
Ser Ser Pro Leu Leu Leu Glu Leu Asn Glu Asn Asp 325 330 335Pro Gly
Ile Phe Val Thr Gln Ser Val His Lys Gln Gln Ala Gly Phe 340 345
350Ser Gln Thr Ser Gln Ile His Lys Lys Asp Asn His Ile Arg Gly Gln
355 360 365Ala Arg Phe Cys Pro His Lys Arg Leu Asn Asn Ala Phe Met
Leu His 370 375 380Ala Ser Thr Ser Pro Phe Tyr Pro Leu Phe Ala Ala
Leu Asp Val Asn385 390 395 400Ala Lys Ile His Glu Gly Glu Ser Gly
Arg Arg Leu Trp Ala Glu Cys 405 410 415Val Glu Ile Gly Ile Glu Ala
Arg Lys Ala Ile Leu Ala Arg Cys Lys 420 425 430Leu Phe Arg Pro Phe
Ile Pro Pro Val Val Asp Gly Lys Leu Trp Gln 435 440 445Asp Tyr Pro
Thr Ser Val Leu Ala Ser Asp Arg Arg Phe Phe Ser Phe 450 455 460Glu
Pro Gly Ala Lys Trp His Gly Phe Glu Gly Tyr Ala Ala Asp Gln465 470
475 480Tyr Phe Val Asp Pro Cys Lys Leu Leu Leu Thr Thr Pro Gly Ile
Asp 485 490 495Ala Glu Thr Gly Glu Tyr Ser Asp Phe Gly Val Pro Ala
Thr Ile Leu 500 505 510Ala His Tyr Leu Arg Glu Asn Gly Ile Val Pro
Glu Lys Cys Asp Leu 515 520 525Asn Ser Ile Leu Phe Leu Leu Thr Pro
Ala Glu Ser His Glu Lys Leu 530 535 540Ala Gln Leu Val Ala Met Leu
Ala Gln Phe Glu Gln His Ile Glu Asp545 550 555 560Asp Ser Pro Leu
Val Glu Val Leu Pro Ser Val Tyr Asn Lys Tyr Pro 565 570 575Val Arg
Tyr Arg Asp Tyr Thr Leu Arg Gln Leu Cys Gln Glu Met His 580 585
590Asp Leu Tyr Val Ser Phe Asp Val Lys Asp Leu Gln Lys Ala Met Phe
595 600 605Arg Gln Gln Ser Phe Pro Ser Val Val Met Asn Pro Gln Asp
Ala His 610 615 620Ser Ala Tyr Ile Arg Gly Asp Val Glu Leu Val Arg
Ile Arg Asp Ala625 630 635 640Glu Gly Arg Ile Ala Ala Glu Gly Ala
Leu Pro Tyr Pro Pro Gly Val 645 650 655Leu Cys Val Val Pro Gly Glu
Val Trp Gly Gly Ala Val Gln Arg Tyr 660 665 670Phe Leu Ala Leu Glu
Glu Gly Val Asn Leu Leu Pro Gly Phe Ser Pro 675 680 685Glu Leu Gln
Gly Val Tyr Ser Glu Thr Asp Ala Asp Gly Val Lys Arg 690 695 700Leu
Tyr Gly Tyr Val Leu Lys705 71025732PRTEscherichia coli 25Met Ser
Lys Leu Lys Ile Ala Val Ser Asp Ser Cys Pro Asp Cys Phe1 5 10 15Thr
Thr Gln Arg Glu Cys Ile Tyr Ile Asn Glu Ser Arg Asn Ile Asp 20 25
30Val Ala Ala Ile Val Leu Ser Leu Asn Asp Val Thr Cys Gly Lys Leu
35 40 45Asp Glu Ile Asp Ala Thr Gly Tyr Gly Ile Pro Val Phe Ile Ala
Thr 50 55 60Glu Asn Gln Glu Arg Val Pro Ala Glu Tyr Leu Pro Arg Ile
Ser Gly65 70 75 80Val Phe Glu Asn Cys Glu Ser Arg Arg Glu Phe Tyr
Gly Arg Gln Leu 85 90 95Glu Thr Ala Ala Ser His Tyr Glu Thr Gln Leu
Arg Pro Pro Phe Phe 100 105 110Arg Ala Leu Val Asp Tyr Val Asn Gln
Gly Asn Ser Ala Phe Asp Cys 115 120 125Pro Gly His Gln Gly Gly Glu
Phe Phe Arg Arg His Pro Ala Gly Asn 130 135 140Gln Phe Val Glu Tyr
Phe Gly Glu Ala Leu Phe Arg Ala Asp Leu Cys145 150 155 160Asn Ala
Asp Val Ala Met Gly Asp Leu Leu Ile His Glu Gly Ala Pro 165 170
175Cys Ile Ala Gln Gln His Ala Ala Lys Val Phe Asn Ala Asp Lys Thr
180 185 190Tyr Phe Val Leu Asn Gly Thr Ser Ser Ser Asn Lys Val Val
Leu Asn 195 200 205Ala Leu Leu Thr Pro Gly Asp Leu Val Leu Phe Asp
Arg Asn Asn His 210 215 220Lys Ser Asn His His Gly Ala Leu Leu Gln
Ala Gly Ala Thr Pro Val225 230 235 240Tyr Leu Glu Thr Ala Arg Asn
Pro Tyr Gly Phe Ile Gly Gly Ile Asp 245 250 255Ala His Cys Phe Glu
Glu Ser Tyr Leu Arg Glu Leu Ile Ala Glu Val 260 265 270Ala Pro Gln
Arg Ala Lys Glu Ala Arg Pro Phe Arg Leu Ala Val Ile 275 280 285Gln
Leu Gly Thr Tyr Asp Gly Thr Ile Tyr Asn Ala Arg Gln Val Val 290 295
300Asp Lys Ile Gly His Leu Cys Asp Tyr Ile Leu Phe Asp Ser Ala
Trp305 310 315 320Val Gly Tyr Glu Gln Phe Ile Pro Met Met Ala Asp
Cys Ser Pro Leu 325 330 335Leu Leu Asp Leu Asn Glu Asn Asp Pro Gly
Ile Leu Val Thr Gln Ser 340 345 350Val His Lys Gln Gln Ala Gly Phe
Ser Gln Thr Ser Gln Ile His Lys 355 360 365Lys Asp Ser His Ile Lys
Gly Gln Gln Arg Tyr Val Pro His Lys Arg 370 375 380Met Asn Asn Ala
Phe Met Met His Ala Ser Thr Ser Pro Phe Tyr Pro385 390 395 400Leu
Phe Ala Ala Leu Asn Ile Asn Ala Lys Met His Glu Gly Val Ser 405 410
415Gly Arg Asn Met Trp Met Asp Cys Val Val Asn Gly Ile Asn Ala Arg
420 425 430Lys Leu Ile Leu Asp Asn Cys Gln His Ile Arg Pro Phe Val
Pro Glu 435 440 445Leu Val Asp Gly Lys Pro Trp Gln Ser Tyr Glu Thr
Ala Gln Ile Ala 450 455 460Val Asp Leu Arg Phe Phe Gln Phe Val Pro
Gly Glu His Trp His Ser465 470 475 480Phe Glu Gly Tyr Ala Glu Asn
Gln Tyr Phe Val Asp Pro Cys Lys Leu 485 490 495Leu Leu Thr Thr Pro
Gly Ile Asp Ala Arg Asn Gly Glu Tyr Glu Ala 500 505 510Phe Gly Val
Pro Ala Thr Ile Leu Ala Asn Phe Leu Arg Glu Asn Gly 515 520 525Val
Val Pro Glu Lys Cys Asp Leu Asn Ser Ile Leu Phe Leu Leu Thr 530 535
540Pro Ala Glu Asp Met Ala Lys Leu Gln Gln Leu Val Ala Leu Leu
Val545 550 555 560Arg Phe Glu Lys Leu Leu Glu Ser Asp Ala Pro Leu
Ala Glu Val Leu 565 570 575Pro Ser Ile Tyr Lys Gln His Glu Glu Arg
Tyr Ala Gly Tyr Thr Leu 580 585 590Arg Gln Leu Cys Gln Glu Met His
Asp Leu Tyr Ala Arg His Asn Val 595 600 605Lys Gln Leu Gln Lys Glu
Met Phe Arg Lys Glu His Phe Pro Arg Val 610 615 620Ser Met Asn Pro
Gln Glu Ala Asn Tyr Ala Tyr Leu Arg Gly Glu Val625 630 635 640Glu
Leu Val Arg Leu Pro Asp Ala Glu Gly Arg Ile Ala Ala Glu Gly 645 650
655Ala Leu Pro Tyr Pro Pro Gly Val Leu Cys Val Val Pro Gly Glu Ile
660 665 670Trp Gly Gly Ala Val Leu Arg Tyr Phe Ser Ala Leu Glu Glu
Gly Ile 675 680 685Asn Leu Leu Pro Gly Phe Ala Pro Glu Leu Gln Gly
Val Tyr Ile Glu 690 695 700Glu His Asp Gly Arg Lys Gln Val Trp Cys
Tyr Val Ile Lys Pro Arg705 710 715 720Asp Ala Gln Ser Thr Leu Leu
Lys Gly Glu Lys Leu 725 73026466PRTEscherichia coli 26Met Asp Gln
Lys Leu Leu Thr Asp Phe Arg Ser Glu Leu Leu Asp Ser1 5 10 15Arg Phe
Gly Ala Lys Ala Ile Ser Thr Ile Ala Glu Ser Lys Arg Phe 20 25 30Pro
Leu His Glu Met Arg Asp Asp Val Ala Phe Gln Ile Ile Asn Asp 35 40
45Glu Leu Tyr Leu Asp Gly Asn Ala Arg Gln Asn Leu Ala Thr Phe Cys
50 55 60Gln Thr Trp Asp Asp Glu Asn Val His Lys Leu Met Asp Leu Ser
Ile65 70 75 80Asn Lys Asn Trp Ile Asp Lys Glu Glu Tyr Pro Gln Ser
Ala Ala Ile 85 90 95Asp Leu Arg Cys Val Asn Met Val Ala Asp Leu Trp
His Ala Pro Ala 100 105 110Pro Lys Asn Gly Gln Ala Val Gly Thr Asn
Thr Ile Gly Ser Ser Glu 115 120 125Ala Cys Met Leu Gly Gly Met Ala
Met Lys Trp Arg Trp Arg Lys Arg 130 135 140Met Glu Ala Ala Gly Lys
Pro Thr Asp Lys Pro Asn Leu Val Cys Gly145 150 155 160Pro Val Gln
Ile Cys Trp His Lys Phe Ala Arg Tyr Trp Asp Val Glu 165 170 175Leu
Arg Glu Ile Pro Met Arg Pro Gly Gln Leu Phe Met Asp Pro Lys 180 185
190Arg Met Ile Glu Ala Cys Asp Glu Asn Thr Ile Gly Val Val Pro Thr
195 200 205Phe Gly Val Thr Tyr Thr Gly Asn Tyr Glu Phe Pro Gln Pro
Leu His 210 215 220Asp Ala Leu Asp Lys Phe Gln Ala Asp Thr Gly Ile
Asp Ile Asp Met225 230 235 240His Ile Asp Ala Ala Ser Gly Gly Phe
Leu Ala Pro Phe Val Ala Pro 245 250 255Asp Ile Val Trp Asp Phe Arg
Leu Pro Arg Val Lys Ser Ile Ser Ala 260 265 270Ser Gly His Lys Phe
Gly Leu Ala Pro Leu Gly Cys Gly Trp Val Ile 275 280 285Trp Arg Asp
Glu Glu Ala Leu Pro Gln Glu Leu Val Phe Asn Val Asp 290 295 300Tyr
Leu Gly Gly Gln Ile Gly Thr Phe Ala Ile Asn Phe Ser Arg Pro305 310
315 320Ala Gly Gln Val Ile Ala Gln Tyr Tyr Glu Phe Leu Arg Leu Gly
Arg 325 330 335Glu Gly Tyr Thr Lys Val Gln Asn Ala Ser Tyr Gln Val
Ala Ala Tyr 340 345 350Leu Ala Asp Glu Ile Ala Lys Leu Gly Pro Tyr
Glu Phe Ile Cys Thr 355 360 365Gly Arg Pro Asp Glu Gly Ile Pro Ala
Val Cys Phe Lys Leu Lys Asp 370 375 380Gly Glu Asp Pro Gly Tyr Thr
Leu Tyr Asp Leu Ser Glu Arg Leu Arg385 390 395 400Leu Arg Gly Trp
Gln Val Pro Ala Phe Thr Leu Gly Gly Glu Ala Thr 405 410 415Asp Ile
Val Val Met Arg Ile Met Cys Arg Arg Gly Phe Glu Met Asp 420 425
430Phe Ala Glu Leu Leu Leu Glu Asp Tyr Lys Ala Ser Leu Lys Tyr Leu
435 440 445Ser Asp His Pro Lys Leu Gln Gly Ile Ala Gln Gln Asn Ser
Phe Lys 450 455 460His Thr46527466PRTEscherichia coli 27Met Asp Lys
Lys Gln Val Thr Asp Leu Arg Ser Glu Leu Leu Asp Ser1 5 10 15Arg Phe
Gly Ala Lys Ser Ile Ser Thr Ile Ala Glu Ser Lys Arg Phe 20 25 30Pro
Leu His Glu Met Arg Asp Asp Val Ala Phe Gln Ile Ile Asn Asp 35 40
45Glu Leu Tyr Leu Asp Gly Asn Ala Arg Gln Asn Leu Ala Thr Phe Cys
50 55 60Gln Thr Trp Asp Asp Glu Asn Val His Lys Leu Met Asp Leu Ser
Ile65 70 75 80Asn Lys Asn Trp Ile Asp Lys Glu Glu Tyr Pro Gln Ser
Ala Ala Ile 85 90 95Asp Leu Arg Cys Val Asn Met Val Ala Asp Leu Trp
His Ala Pro Ala 100 105 110Pro Lys Asn Gly Gln Ala Val Gly Thr Asn
Thr Ile Gly Ser Ser Glu 115 120 125Ala Cys Met Leu Gly Gly Met Ala
Met Lys Trp Arg Trp Arg Lys Arg 130 135 140Met Glu Ala Ala Gly Lys
Pro Thr Asp Lys Pro Asn Leu Val Cys Gly145 150 155 160Pro Val Gln
Ile Cys Trp His Lys Phe Ala Arg Tyr Trp Asp Val Glu 165 170 175Leu
Arg Glu Ile Pro Met Arg Pro Gly Gln Leu Phe Met Asp Pro Lys 180 185
190Arg Met Ile Glu Ala Cys Asp Glu Asn Thr Ile Gly Val Val Pro Thr
195 200 205Phe Gly Val Thr Tyr Thr Gly Asn Tyr Glu Phe Pro Gln Pro
Leu His 210 215 220Asp Ala Leu Asp Lys Phe Gln Ala Asp Thr Gly Ile
Asp Ile Asp Met225 230 235 240His Ile Asp Ala Ala Ser Gly Gly Phe
Leu Ala Pro Phe Val Ala Pro 245 250 255Asp Ile Val Trp Asp Phe Arg
Leu Pro Arg Val Lys Ser Ile Ser Ala 260 265 270Ser Gly His Lys Phe
Gly Leu Ala Pro Leu Gly Cys Gly Trp Val Ile 275 280 285Trp Arg Asp
Glu Glu Ala Leu Pro Gln Glu Leu Val Phe Asn Val Asp 290 295 300Tyr
Leu Gly Gly Gln Ile Gly Thr Phe Ala Ile Asn Phe Ser Arg Pro305 310
315 320Ala Gly Gln Val Ile Ala Gln Tyr Tyr Glu Phe Leu Arg Leu Gly
Arg 325 330 335Glu Gly Tyr Thr Lys Val Gln Asn Ala Ser Tyr Gln Val
Ala Ala Tyr 340 345 350Leu Ala Asp Glu Ile Ala Lys Leu Gly Pro Tyr
Glu Phe Ile Cys Thr 355 360 365Gly Arg Pro Asp Glu Gly Ile Pro Ala
Val Cys Phe Lys Leu Lys Asp 370 375 380Gly Glu Asp Pro Gly Tyr Thr
Leu Tyr Asp Leu Ser Glu Arg Leu Arg385 390 395 400Leu Arg Gly Trp
Gln Val Pro Ala Phe Thr Leu Gly Gly Glu Ala Thr 405 410 415Asp Ile
Val Val Met Arg Ile Met Cys Arg Arg Gly Phe Glu Met Asp 420 425
430Phe Ala Glu Leu Leu Leu Glu Asp Tyr Lys Ala Ser Leu Lys Tyr Leu
435
440 445Ser Asp His Pro Lys Leu Gln Gly Ile Ala Gln Gln Asn Ser Phe
Lys 450 455 460His Thr4652819PRTSaccharomyces cerevisiae 28Gln Gln
Gln Arg Asn Trp Lys Gln Gly Gly Asn Tyr Gln Gln Tyr Gln1 5 10 15Ser
Tyr Asn2930PRTSaccharomyces cerevisiae 29Ser Asn Tyr Asn Asn Tyr
Asn Asn Tyr Asn Asn Tyr Asn Asn Tyr Asn1 5 10 15Asn Tyr Asn Asn Tyr
Asn Lys Tyr Asn Gly Gln Gly Tyr Gln 20 25 30308PRTSaccharomyces
cerevisiae 30Pro Gln Gly Gly Tyr Gln Gln Asn1 5319PRTSaccharomyces
cerevisiae 31Phe Pro Pro Lys Lys Phe Lys Asp Leu1
53218PRTSaccharomyces cerevisiae 32Phe Pro Pro Lys Lys Phe Lys Asp
Leu Asn Ser Phe Leu Asp Asp Gln1 5 10 15Pro Lys3336PRTSaccharomyces
cerevisiae 33Phe Pro Pro Lys Lys Phe Lys Asp Leu Asn Ser Phe Leu
Asp Asp Gln1 5 10 15Pro Lys Asp Pro Asn Leu Val Ala Ser Pro Phe Gly
Gly Tyr Phe Lys 20 25 30Asn Pro Ala Ala 353445PRTSaccharomyces
cerevisiae 34Phe Pro Pro Lys Lys Phe Lys Asp Leu Asn Ser Phe Leu
Asp Asp Gln1 5 10 15Pro Lys Asp Pro Asn Leu Val Ala Ser Pro Phe Gly
Gly Tyr Phe Lys 20 25 30Asn Pro Ala Ala Asp Ala Gly Ser Asn Asn Ala
Ser Lys 35 40 453510PRTBovine adenovirus type 1 35Arg Arg Phe Gly
Glu Ala Ser Ser Ala Phe1 5 103610PRTEscherichia coli 36Ala Ser Gln
Trp Pro Glu Glu Thr Phe Gly1 5 103710PRTEscherichia coli 37Glu Gly
Val Ala Glu Thr Asn Glu Asp Phe1 5 103852DNAArtificial
SequencePrimer cadA-F 38ggcgagctca cacaggaaac agaccatgaa cgttattgca
atattgaatc ac 523928DNAArtificial SequencePrimer cadA-R
39ggctctagac cacttccctt gtacgagc 284044DNAArtificial SequencePrimer
cadA-F2 40atttcacaca ggaaacagct atgaacgtta ttgcaatatt gaat
444120DNAArtificial SequencePrimer cadA-R2 41agctgtttcc tgtgtgaaat
204233DNAArtificial SequencePrimer cat-HindIII-F 42ggcaagcttg
agaaaaaaat cactggatat acc 334329DNAArtificial SequencePrimer
cat-NdeI-R 43ggccatatgt aagggcacca ataactgcc 294431DNAArtificial
SequencePrimer cadAt-XbaI-R 44ggctctagat ttgctttctt ctttcaatac c
314533DNAArtificial SequencePrimer cat-XbaI-F 45ggctctagag
agaaaaaaat cactggatat acc 334630DNAArtificial SequencePrimer
New1-XbaI-F 46ggctctagag gttctggctc tggttctccg 304731DNAArtificial
SequencePrimer New1-HindIII-R 47ggcaagcttt tactggtagc cctgaccgtt g
314829DNAArtificial SequencePrimer Sup35-XbaI-F 48ggctctagag
gtagcggctc tggctctga 294932DNAArtificial SequencePrimer
Sup35-HindIII-R 49ggcaagcttt tagccaccct gtgggttaaa ct
325029DNAArtificial SequencePrimer cadA-XbaI-F 50ggctctagaa
tttcacacag gaaacagct 295129DNAArtificial SequencePrimer
cadA-HindIII-R 51ggcaagcttc acttcccttg tacgagcta
295234DNAArtificial SequencePrimer rbs2-SacI-F 52ggcgagctca
tgaacgttat tgcaatattg aatc 345325DNAArtificial SequencePrimer
rbs2-SacI-R 53ggcgagctcc tcctgtgtga aattg 255429DNAArtificial
SequencePrimer New1-SacI-F 54ggcgagctca tgggttctgg ctctggttc
295528DNAArtificial SequencePrimer New1-XbaI-R 55ggctctagac
tggtagccct gaccgttg 285627DNAArtificial SequencePrimer Sup35-SacI-F
56ggcgagctca tgggtagcgg ctctggc 275729DNAArtificial SequencePrimer
Sup35-XbaI-R 57ggctctagag ccaccctgtg ggttaaact 295828DNAArtificial
SequencePrimer 222-de-R 58tctagatttg ctttcttctt tcaatacc
285946DNAArtificial SequencePrimer 222-de-9-F 59gaagaaagca
aatctagaaa gttcaaagac ctgaactctt tcggtg 466042DNAArtificial
SequencePrimer 222-de-18-F 60gaagaaagca aatctagaga cgaccagccg
aaagacccga ac 426141DNAArtificial SequencePrimer 222-de-36-F
61gaagaaagca aatctagaaa aaacccagcg gcggacgcgg g 416241DNAArtificial
SequencePrimer 222-de-45-F 62gaagaaagca aatctagaaa caacgcgtct
aagaaatctt c 416347DNAArtificial SequencePrimer 222-C2-F
63tttcggcgaa gcgagcagcg cgttctaaaa gcttaagaga caggatg
476447DNAArtificial SequencePrimer 222-C2-R 64cgctgctcgc ttcgccgaaa
cgacgctggt agccctgacc gttgtat 476547DNAArtificial SequencePrimer
222-C4-F 65ccagtggccg gaagaaacct tcggctaaaa gcttaagaga caggatg
476647DNAArtificial SequencePrimer 222-C4-R 66aggtttcttc cggccactgg
ctcgcctggt agccctgacc gttgtat 476747DNAArtificial SequencePrimer
222-C6-F 67cgtggcggaa accaacgaag atttctaaaa gcttaagaga caggatg
476847DNAArtificial SequencePrimer 222-C6-R 68cttcgttggt ttccgccacg
ccttcctggt agccctgacc gttgtat 47
D00000

D00001

D00002

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.