Anti-pd-l1 Antibodies
CAMPBELL; Jamie ; et al.
U.S. patent application number 16/311440 was filed with the patent office on 2019-10-31 for anti-pd-l1 antibodies. The applicant listed for this patent is KYMAB LIMITED. Invention is credited to Stephen John ARKINSTALL, Jamie CAMPBELL, Cecilia DEANTONIO, Thomas GALLAGHER, Volker GERMASCHEWSKI, Stephen Douglas GILLIES, Ian KIRBY, Miha KOSMAC, Nikole SANDY, Cassandra VAN KRINKS.
| Application Number | 20190330351 16/311440 |
| Document ID | / |
| Family ID | 57964423 |
| Filed Date | 2019-10-31 |
View All Diagrams
| United States Patent Application | 20190330351 |
| Kind Code | A1 |
| CAMPBELL; Jamie ; et al. | October 31, 2019 |
ANTI-PD-L1 ANTIBODIES
Abstract
The present invention relates to anti-PD-L1 antibodies, bispecific antibodies containing one domain with specificity to PD-L1, and to immunocytokines comprising an anti-PD-L1 antibody fused to a cytokine, such as IL-2. The present invention also provides methods of treatment, uses and pharmaceutical compositions comprising the antibodies, bispecific antibodies and immunocytokines.
| Inventors: | CAMPBELL; Jamie; (Babraham, GB) ; SANDY; Nikole; (Babraham, GB) ; VAN KRINKS; Cassandra; (Babraham, GB) ; ARKINSTALL; Stephen John; (Cambridge, MA) ; GERMASCHEWSKI; Volker; (Babraham, GB) ; KIRBY; Ian; (Babraham, GB) ; KOSMAC; Miha; (Babraham, GB) ; GALLAGHER; Thomas; (Babraham, GB) ; DEANTONIO; Cecilia; (Babraham, GB) ; GILLIES; Stephen Douglas; (Carlisle, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 57964423 | ||||||||||
| Appl. No.: | 16/311440 | ||||||||||
| Filed: | June 20, 2017 | ||||||||||
| PCT Filed: | June 20, 2017 | ||||||||||
| PCT NO: | PCT/GB2017/051796 | ||||||||||
| 371 Date: | December 19, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15354971 | Nov 17, 2016 | 9617338 | ||
| 16311440 | ||||
| 15211504 | Jul 15, 2016 | 9567399 | ||
| 15354971 | ||||
| 62352291 | Jun 20, 2016 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 14/55 20130101; C07K 16/2803 20130101; A61K 2039/505 20130101; C07K 2319/74 20130101; C07K 16/28 20130101; C07K 2317/33 20130101; C07K 2317/524 20130101; C07K 16/2827 20130101; C07K 2319/30 20130101; C07K 2317/567 20130101; C07K 2317/92 20130101; C07K 2319/00 20130101; C07K 2317/56 20130101; C07K 2317/31 20130101; C07K 2317/71 20130101; C07K 2317/565 20130101; C07K 19/00 20130101; C07K 16/468 20130101; C07K 2317/21 20130101; C07K 2319/02 20130101; C07K 2317/76 20130101 |
| International Class: | C07K 16/28 20060101 C07K016/28; C07K 16/46 20060101 C07K016/46; C07K 14/55 20060101 C07K014/55; C07K 19/00 20060101 C07K019/00 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Sep 7, 2016 | GB | 1615224.1 |
| Sep 8, 2016 | GB | 1613683.0 |
| Sep 9, 2016 | GB | 1615335.5 |
| Dec 1, 2016 | GB | 1620414.1 |
| Dec 20, 2016 | GB | 1621782.0 |
| Feb 13, 2017 | GB | 1702338.3 |
| Feb 13, 2017 | GB | 1702339.1 |
| Feb 24, 2017 | GB | 1703071.9 |
Claims
1-61. (canceled)
62. An antibody or antigen binding fragment thereof that specifically binds to hPD-L1, comprising a V.sub.H domain and a V.sub.L domain, wherein the V.sub.H domain comprises an amino acid sequence that is at least 90% identical to SEQ ID NO: 33, and the V.sub.L domain comprises an amino acid sequence that is at least 90% identical to SEQ ID NO: 43.
63. The antibody or antigen binding fragment of claim 62, wherein the V.sub.L domain comprises the amino acid sequence of SEQ ID NO: 43 and the V.sub.H domain comprises an amino acid sequence selected from the group consisting of: a) the V.sub.H domain of heavy chain amino acid sequence SEQ ID NO: 47; b) the V.sub.H domain of heavy chain amino acid sequence SEQ ID NO: 48; c) the V.sub.H domain of heavy chain amino acid sequence SEQ ID NO: 49; d) the V.sub.H domain of heavy chain amino acid sequence SEQ ID NO: 342; and e) SEQ ID NO: 33.
64. The antibody or antigen binding fragment of claim 62, wherein the wherein the V.sub.L domain comprises the amino acid sequence of SEQ ID NO: 43 and the V.sub.H domain comprises the amino acid sequence of SEQ ID NO: 33.
65. The antibody or antigen binding fragment of claim 62, wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO: 33, 47, 48, or 342 and the V.sub.H domain comprises an amino acid sequence selected from the group consisting of: a) the V.sub.L domain of light chain amino acid sequence SEQ ID NO: 50; b) the V.sub.L domain of light chain amino acid sequence SEQ ID NO: 51; and c) the V.sub.L domain of light chain amino acid sequence SEQ ID NO: 298.
66. The antibody or antigen binding fragment of claim 62, wherein the antibody or fragment comprises the CDRH3 sequence of SEQ ID NO: 29 or 32, and/or the CDRH1 sequence of SEQ ID NO: 27 or 30, and/or the CDRH2 sequence of SEQ ID NO: 28 or 31 and/or the CDRL1 sequence of SEQ ID NO: 37 or 40, and/or the CDRL2 sequence of SEQ ID NO: 38 or 41, and/or the CDRL3 sequence of SEQ ID NO: 39 or 42.
67. The antibody or antigen binding fragment of claim 66, wherein the antibody or fragment comprises: a. the CDRH1 amino acid sequence of SEQ ID NO: 27 or SEQ ID NO: 30, the CDRH2 amino acid sequence of SEQ ID NO: 28 or SEQ ID NO: 31, the CDRH3 amino acid sequence of SEQ ID NO: 29 or SEQ ID NO: 32, the CDRL1 amino acid sequence of SEQ ID NO: 37 or SEQ ID NO: 40, the CDRL2 amino acid sequence of SEQ ID NO: 38 or SEQ ID NO: 41, and the CDRL3 amino acid sequence of SEQ ID NO: 39 or SEQ ID NO: 42; b. the CDRH1 amino acid sequence of SEQ ID NO: 27 or SEQ ID NO: 30, the CDRH2 amino acid sequence of SEQ ID NO: 28 or SEQ ID NO: 31, the CDRH3 amino acid sequence of SEQ ID NO: 29 or SEQ ID NO: 32, the CDRL1 amino acid sequence of SEQ ID NO: 37 or SEQ ID NO: 40, the CDRL2 amino acid sequence as defined by the Kabat or IMGT systems from the V.sub.L sequence of SEQ ID NO: 50, and the CDRL3 amino acid sequence of SEQ ID NO: 39 or SEQ ID NO: 42; or c. the CDRH1 amino acid sequence of SEQ ID NO: 27 or SEQ ID NO: 30, the CDRH2 amino acid sequence of SEQ ID NO: 28 or SEQ ID NO: 31, the CDRH3 amino acid sequence of SEQ ID NO: 29 or SEQ ID NO: 32, the CDRL1 amino acid sequence of SEQ ID NO: 37 or SEQ ID NO: 40, the CDRL2 amino acid sequence as defined by the Kabat or IMGT systems from the V.sub.L sequence of SEQ ID No: 298, and the CDRL3 amino acid sequence of SEQ ID NO: 39 or SEQ ID NO: 42.
68. The antibody or antigen binding fragment of claim 62, wherein the antibody or fragment specifically binds to cynomolgus PD-L1 as defined by SEQ ID NO: 2.
69. The antibody or antigen binding fragment of claim 62, wherein the antibody or fragment comprises a constant region.
70. The antibody or antigen binding fragment of claim 69, wherein the constant region is an effector-null human constant region, and wherein the effector-null human constant region is an IgG4 constant region or an IgG1 constant region, optionally wherein the constant region is IgG4-PE (SEQ ID NO: 199) or a disabled IgG1 of SEQ ID NO: 205.
71. A bispecific antibody or fusion protein comprising an antibody or antigen binding fragment of claim 62.
72. The bispecific antibody of claim 71, wherein the bispecific format is selected from the group consisting of DVD-Ig, mAb.sup.2, FIT-Ig, mAb-dAb, dock and lock, SEEDbody, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BiTE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, minibody, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, and charge pairs with common light chain.
73. The bispecific antibody of claim 71, wherein the bispecific antibody specifically binds to hPD-L1 and another target antigen selected from the group consisting of an immune checkpoint inhibitor, an immune modulator, and an immune activator.
74. A method of treating or preventing a hPD-L1 mediated disease or condition in a human, comprising administering to said human a therapeutically effective amount of the antibody or antigen binding fragment of claim 62, wherein the disease or condition is selected from the group consisting of a neurodegenerative disease or condition, neoplastic disease, non-neoplastic disease, chronic viral infection, and cancer, and wherein the hPD-L1 mediated disease or condition is thereby treated or prevented.
75. A pharmaceutical composition comprising the antibody or antigen binding fragment of claim 62 and a pharmaceutically acceptable excipient, diluent or carrier.
76. The pharmaceutical composition of claim 75, further comprising a chemokine receptor antagonist selected from the group consisting of CXCR4, CCR4, and CXCR2.
77. A method of treating or preventing a hPD-L1 mediated disease or condition in a human, comprising administering to said human a therapeutically effective amount of the pharmaceutical composition of claim 75, wherein the hPD-L1-mediated condition or disease selected from the group consisting of a neurodegenerative disease or condition, neoplastic disease, non-neoplastic disease, chronic viral infection, and cancer, and wherein the hPD-L1 mediated disease or condition is thereby treated or prevented.
78. A nucleic acid that encodes a heavy chain or a light chain of the antibody or antigen binding fragment of claim 62.
79. A vector comprising the nucleic acid of claim 78.
80. The vector of claim 79, wherein the vector is a CHO vector or a HEK293 vector.
81. A host cell comprising the vector of claim 80.
Description
FIELD OF THE INVENTION
[0001] Antibodies and methods of using the antibodies are described. In particular, antibodies that specifically bind human PD-L1 antigen and their use in treating various diseases are described.
Introduction
[0002] Immunocytokines (antibody-cytokine fusion proteins) were first reported in the literature in the early 1990s and consisted of whole antibody fusions with cytokines such as lymphotoxin (TNF-.alpha.) or interleukin 2 (IL-2). Subsequent studies in GD2-expressing tumour models in mice indicated that the ch14.18 antibody and ch14.18-IL2 immunocytokine both had anti-tumour activity but that the immunocytokine was far more potent than the antibody, even when combined with free IL-2, (see Sabzevari H et al., Proc. Natl. Acad. Sci. USA, 1994, 91:9626-30; Pancook J D, et al., Cancer Immunol. Immunother., 1996, 42:88-92; Becker J C, et al., Proc. Natl. Acad. Sci. USA, 1996, 93:2702-7). In addition, immune-competent mice treated with the immunocytokine, but not the antibody plus IL-2, developed an adaptive immune response dependent on CD8.sup.+ T-cells that prevented subsequent tumour challenge (Becker J C, et al., J. Exp. Med., 1996, 183:2361-6; Becker J C, et al., Proc. Natl. Acad. Sci. USA, 1996, 93:7826-31). Thus, the targeting of IL-2 to the tumour microenvironment induces an anti-tumour vaccine effect that is not possible with the antibody, either alone or together with the free cytokine. A related humanized immunocytokine, hu 14.18-IL2, achieved clinical proof of concept in relapsed non-bulky neuroblastoma as monotherapy where it induced a significant number of complete responses in patients with no other treatment options (see Shusterman et al., Journal of Clinical Oncology, 2010, 28(33), 4969-4975). A number of publications describe the ability of this molecule to activate several components of the immune system to kill tumour cells (particularly NK cells and CD8.sup.+ T-cells), and develop T-cell memory in order to resist subsequent tumour challenge (Yamane et al. 2009; Expert Opi, Investig. Drugs, 18(7): 991-1000; Neal et al., 2004, Clin. Cancer Res., 1010, 4839-4847).
[0003] As IL-2 based immunocytokines can have significant side effects, recent efforts have focused on the reduction of toxicity whilst maintaining efficacy. One example is Selectikine (EMD 521873), which has a substitution of aspartic acid for threonine at position 20 of IL-2, a key residue in the binding of IL-2R.beta. (Gillies et al., Clinical Cancer Research, 2011, 17(11), 3673-3685). Selectikine, which binds necrotic tissue, has been shown to have good anti-tumour activity, despite its selectivity for the high affinity IL-2R, over the intermediate IL-2R and good tolerability in Phase I studies (Laurent et al., Journal of Translational Medicine, 2013, 11(1), 5. http://doi.org/10.1186/1479-5876-11-5)
[0004] WO2012/178137 (Gillies) and an associated journal article (Gilles, Protein Engineering, Design and Selection, 2013, 26(10), 561-569) describe light chain immunocytokine fusions with tumour targeting antibodies, and modulation of IL-2 activity by the introduction of truncations in the N-terminal part of the cytokine, which decreases signalling through IL-2R.beta..gamma.. IL-2 fusion proteins that specifically target IL-2RP), have been shown to have increased toxicity compared with wild-type (Vasquez-Lombardi et al. Nat Comm, 2017, DOI: 10.1038/ncomms15373), supporting the notion that decreasing IL-2R.beta..gamma. binding may be beneficial in terms of side effects.
[0005] An adaptive immune response involves activation, selection, and clonal proliferation of two major classes of lymphocytes termed T-cells and B-cells. After encountering an antigen, T-cells proliferate and differentiate into antigen-specific effector cells, while B-cells proliferate and differentiate into antibody-secreting cells. T-cell activation is a multi-step process requiring several signalling events between the T-cell and an antigen-presenting cell (APC). For T-cell activation to occur, two types of signals must be delivered to a resting T-cell. The first type is mediated by the antigen-specific. T-cell receptor (TcR), and confers specificity to the immune response. The second signal, a costimulatory type signal, regulates the magnitude of the response and is delivered through accessory receptors on the T-cell.
[0006] A primary costimulatory signal is delivered through the activating CD28 receptor upon engagement of its ligands B7-1 or B7-2. In contrast, engagement of the inhibitory CTLA-4 receptor by the same B7-1 or B7-2 ligands results in attenuation of a T-cell response. Thus, CTLA-4 signals antagonize costimulation mediated by CD28. At high antigen concentrations, CD28 costimulation overrides the CTLA-4 inhibitory effect. Temporal regulation of the CD28 and CTLA-4 expression maintains a balance between activating and Inhibitory signals and ensures the development of an effective immune response, while safeguarding against the development of autoimmunity.
[0007] Programmed death-1 (PD-1) is a 50-55 kDa type I transmembrane receptor that is a member of the CD28 family. PD-1 is involved in the regulation of T-cell activation and is expressed on T-cells, B cells, and myeloid cells. Two ligands for PD-1, PD ligand 1 (PD-L1) and ligand 2 (PD-L2) have been identified and have co-stimulatory features.
[0008] Programmed cell death 1 ligand 1 (PD-L1), also known as cluster of differentiation (CD274) or B7 homolog 1 (B7-H1), is a member of the B7 family that modulates activation or inhibition of the PD-1 receptor. The open reading frame of PD-L1 encodes a putative type 1 transmembrane protein of 290 amino acids, which includes two extracellular Ig domains (a N-terminal V-like domain and a Ig C-like domain), a hydrophobic transmembrane domain and a cytoplasmic tail of 30 amino acids. The 30 amino acid intracellular (cytoplasmic) domain contains no obvious signalling motifs, but does have a potential site for protein kinase C phosphorylation.
[0009] The complete amino acid sequence for PD-L1 can be found in NCBI Reference Sequence: NP_054862.1 (SEQ ID NO: 1), which refers to many journal articles, including, for example, Dong, H., et al. (1999), "PD-L1, a third member of the B7 family, co-stimulates T-cell proliferation and interleukin-10 secretion," Nat. Med. 5 (12), 1365-1369. The PD-L1 gene is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, and zebrafish. The murine form of PD-L1 bears 69% amino acid identity with the human form of PD-L1, and also shares a conserved structure.
[0010] In humans, PD-L1 is expressed on a number of immune cell types including activated and anergic/exhausted T-cells, on naive and activated B-cells, as well as on myeloid dendritic cells (DC), monocytes and mast cells. It is also expressed on non-immune cells including islets of the pancreas, Kupffer cells of the liver, vascular endothelium and selected epithelia, for example airway epithelia and renal tubule epithelia, where its expression is enhanced during inflammatory episodes. PD-L1 expression is also found at increased levels on a number of tumours including, but not limited to breast (including but not limited to triple negative breast cancer and inflammatory breast cancer), ovarian, cervical, colon, colorectal, lung, including non-small cell lung cancer, renal, including renal cell carcinoma, gastric, oesophageal, bladder, hepatocellular cancer, squamous cell carcinoma of the head and neck (SCCHN) and pancreatic cancer, melanoma and uveal melanoma.
[0011] PD-1/PD-L1 signalling is believed to serve a critical non-redundant function within the immune system by negatively regulating T-cell responses. This regulation is involved in T-cell development in the thymus, in regulation of chronic inflammatory responses and in maintenance of both peripheral tolerance and immune privilege. It appears that upregulation of PD-L1 may allow cancers to evade the host immune system and, in many cancers, the expression of PD-L1 is associated with reduced survival and an unfavourable prognosis. Therapeutic monoclonal antibodies that are able to block the PD-1/PD-L1 pathway may enhance anti-tumoural immune responses in patients with cancer. Published clinical data suggest a correlation between clinical responses with tumoural membranous expression of PD-L1 (Brahmer et al., Journal of Clinical Oncology, 2010, Topalian et al, NEJM, 2012) and a stronger correlation between lack of clinical responses and a lack of PD-L1 protein localized to the membrane (Brahmer et al., Journal of Clinical Oncology, 2010, Topalian et al., NEJM, 2012). Thus, PD-L1 expression in tumours or tumour-infiltrating leukocytes (Herbst R S, et al., "Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients", Nature, 2014, Nov. 27, 515(7528):563-7, doi: 10.1038/nature14011) is a candidate molecular marker for use in selecting patients for immunotherapy, for example, immunotherapy using anti-PD-L1 antibodies. Patient enrichment based on surface expression of PD-L1 may significantly enhance the clinical success of treatment with drugs targeting the PD-1/PD-L1 pathway. There is also evidence of an on-going immune response, such as the tumour infiltrating CD8.sup.+ T-cells, or the presence of signature of cytokine activation, such as IFN.gamma..
[0012] Further evidence of PD-L1 expression and correlation to disease will emerge from the numerous ongoing clinical trials. Atezolizumab is the most advanced, and recent data from Phase II trials shows therapeutic effects in metastatic urothelial carcinoma and NSCLC, particularly in patients with PD-L1.sup.+ immune cells in the tumour microenvironment (see Fehrenbacher et al, 2016, The Lancet, http://doi.org/10.1016/S0140-6736(16)00587-0; Rosenberg et al., 2016, The Lancet, http://doi.org/10.101650140-6736(16)00561-4). Recent results from a Phase III trial of 1225 patients with NSCLC showed improved survival in patients taking atezolizumab, compared with chemotherapy, regardless of tumour expression of PD-L1 (Rittmeyer et al, 2017, The Lancet, 389(10066), 255-265).
SUMMARY OF THE INVENTION
Antibodies
[0013] Disclosed herein are antibodies and antigen binding fragments thereof that specifically bind to PD-L1. In one embodiment, the antibody or antigen binding fragment thereof specifically binds to surface expressed PD-L1.
[0014] In a first configuration, there is provided an antibody or a fragment thereof, that specifically binds to hPD-L1 as defined by Seq ID No:1, and competes for binding to said hPD-L1 with the antibody 1D05, wherein the antibody or fragment comprises a V.sub.H domain which comprises a CDRH3 comprising the motif X.sub.1GSGX.sub.2YGX.sub.3X.sub.4FD, wherein X.sub.1, X.sub.2 and X.sub.3 are independently any amino acid, and X.sub.4 is either present or absent, and if present, may be any amino acid.
[0015] In a second configuration, there is provided an antibody or a fragment thereof which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 1D05, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:29 or 32, or the CDRH3 sequence of SEQ ID NO:29 or 32 comprising 6 or fewer amino acid substitutions.
[0016] In a third configuration, there is provided an antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 1D05 specifically binds.
[0017] In a fourth configuration, there is provided an antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 1D05.
[0018] In a fifth configuration, there is provided a bispecific antibody or fusion protein comprising an antibody or fragment thereof as defined in any other configuration, embodiment or concept.
[0019] In a sixth configuration, there is provided an antibody or fragment as defined in any other configuration, embodiment or concept for use in treating or preventing a hPD-L1-mediated disease or condition.
[0020] In a seventh configuration, there is provided the use of an antibody or fragment as defined in any other configuration, embodiment or concept in the manufacture of a medicament for administration to a human for treating or preventing a hPD-L1 mediated disease or condition in the human.
[0021] In an eighth configuration, there is provided a method of treating or preventing a hPD-L1 mediated disease or condition in a human, comprising administering to said human a therapeutically effective amount of an antibody or fragment as defined in any other configuration, embodiment or concept, wherein the hPD-L1 mediated disease or condition is thereby treated or prevented.
[0022] In a ninth configuration, there is provided a pharmaceutical composition comprising an antibody of fragment as defined in any other configuration, embodiment or concept and a pharmaceutically acceptable excipient, diluent or carrier.
[0023] In a tenth configuration, there is provided a kit comprising a pharmaceutical composition comprising an antibody of fragment as defined in any other configuration, embodiment or concept and a pharmaceutically acceptable excipient, diluent or carrier.
[0024] In an eleventh configuration, there is provided a method of modulating PD-1/PD-L1 interaction in a patient, comprising administering an effective amount of an antibody or fragment as defined in any other configuration, embodiment or concept to said patient.
[0025] In a twelfth configuration, there is provided a method of inhibiting PD-L1 activity in a patient, comprising administering an effective amount of an antibody or fragment as defined, in any other configuration, embodiment or concept to said patient.
[0026] In a thirteenth configuration, there is provided a method of treating a proliferative disease in an animal (e.g. a human), comprising administering an effective amount of an antibody or fragment as defined in any other configuration, embodiment or concept to said patient.
[0027] In a fourteenth configuration, there is provided a method of detecting PD-L1 expression in a sample, comprising contacting the sample with an antibody or fragment as defined in any other configuration, embodiment or concept.
[0028] In a fifteenth configuration, there is provided a method comprising contacting a biological sample with an antibody or fragment as defined in any other configuration, embodiment or concept to form a complex with PD-L1 present in the sample and measuring the presence, absence or level of the complex in the biological sample.
[0029] In a sixteenth configuration, there is provided a method of detecting PD-L1 expression in a sample, comprising contacting the sample with an antibody or fragment as defined in any other configuration, embodiment or concept.
[0030] In a seventeenth configuration, there is provided a method comprising contacting a biological sample with an antibody or fragment as defined in any other configuration, embodiment or concept to form a complex with PD-L1 present in the sample and measuring the presence, absence or level of the complex in the biological sample.
[0031] In a eighteenth configuration, there is provided a method for identifying binding partners for PD-L1, the method comprising immunoprecipitating an intact protein complex comprising PD-L1 using an antibody or fragment as defined in any other configuration, embodiment or concept.
[0032] In a nineteenth configuration, there is provided a method of diagnosing a disease in a human subject associated with altered PD-L1 expression comprising the steps of contacting a biological sample from the human subject with an antibody as defined in other configuration, embodiment or concept to form a complex between the antibody and PD-L1 present in the sample; and detecting the amount of the complex.
[0033] In a twentieth configuration, there is provided a nucleic acid that encodes the CDRH3 of an antibody or fragment as defined in any other configuration, embodiment or concept.
[0034] In a twenty-first configuration, there is provided a nucleic acid that encodes a VH domain and/or a VL domain of an antibody or fragment as defined in any other configuration, embodiment or concept.
[0035] In a twenty-second configuration, there is provided a vector comprising the nucleic acid of any other configuration, embodiment or concept; optionally wherein the vector is a CHO or HEK293 vector.
[0036] In a twenty-third configuration, there is provided a host comprising the nucleic acid of any other configuration, embodiment or concept or the vector of any other configuration, embodiment or concept.
Immunocytokines
[0037] In a first configuration, there is provided an immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction:
[0038] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and
[0039] b) A heavy chain constant region; [0040] and wherein the light chain comprises in N- to C-terminal direction:
[0041] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3;
[0042] d) A light chain constant region, (C.sub.L);
[0043] e) Optionally, a linker, (L); and
[0044] f) An IL-2 cytokine; [0045] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to hPD-L1 as defined by Seq ID No:1, and competes for binding to said hPD-L1 with the antibody 1D05; and
[0046] wherein the immunocytokine comprises a V.sub.H domain which comprises a CDRH3 comprising the motif X.sub.1GSGX.sub.2YGX.sub.3X.sub.4FD, wherein X.sub.1, X.sub.2 and X.sub.3 are independently any amino acid, and X.sub.4 is either present or absent, and if present, may be any amino acid.
[0047] In a second configuration, there is provided an immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction:
[0048] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and
[0049] b) A heavy chain constant region;
[0050] and wherein the light chain comprises in N- to C-terminal direction:
[0051] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3;
[0052] d) A light chain constant region, (C.sub.L);
[0053] e) Optionally, a linker, (L); and
[0054] f) An IL-2 cytokine;
[0055] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 1D05, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:29 or 32, or the CDRH3 sequence of SEQ ID NO:29 or 32 comprising 6 or fewer amino acid substitutions.
[0056] In a third configuration, there is provided an immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction:
[0057] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and
[0058] b) A heavy chain constant region; [0059] and wherein the light chain comprises in N- to C-terminal direction:
[0060] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3;
[0061] d) A light chain constant region, (C.sub.L);
[0062] e) Optionally, a linker, (L); and
[0063] f) An IL-2 cytokine; [0064] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to hPD-L1; and [0065] wherein the V.sub.H domain comprises a CDRH3 of from 12 to 20 amino acids and which is derived from the recombination of a human V.sub.H gene segment, a human D gene segment and a human J.sub.H gene segment, wherein the human J.sub.H gene segment is IGHJ5 (e.g. IGHJ5*02).
[0066] In a fourth configuration, there is provided an immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction:
[0067] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and
[0068] b) A heavy chain constant region;
[0069] and wherein the light chain comprises in N- to C-terminal direction:
[0070] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3;
[0071] d) A light chain constant region, (C.sub.L);
[0072] e) Optionally, a linker, (L); and
[0073] f) An IL-2 cytokine;
[0074] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to an epitope that is identical to an epitope to which the antibody 1D05 specifically binds.
[0075] In a fifth configuration, there is provided an immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction:
[0076] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and
[0077] b) A heavy chain constant region;
[0078] and wherein the light chain comprises in N- to C-terminal direction: [0079] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3;
[0080] d) A light chain constant region, (C.sub.L);
[0081] e) Optionally, a linker, (L); and
[0082] f) An IL-2 cytokine;
[0083] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site which competes for binding to hPD-L1 with the antibody 1D05.
[0084] In a sixth configuration, there is provided an immunocytokine as defined in any other configuration, embodiment or aspect for use in treating or preventing a hPD-L1-mediated disease or condition.
[0085] In a seventh configuration, there is provided the use of an immunocytokine as defined in any other configuration, embodiment or aspect in the manufacture of a medicament for administration to a human for treating or preventing a hPD-L1 mediated disease or condition in the human.
[0086] In an eighth configuration, there is provided a method of treating or preventing a hPD-L1 mediated disease or condition in a human, comprising administering to said human a therapeutically effective amount of an immunocytokine as defined in any other configuration, embodiment or aspect, wherein the hPD-L1 mediated disease or condition is thereby treated or prevented.
[0087] In a ninth configuration, there is provided a pharmaceutical composition comprising an immunocytokine as defined in any other configuration, embodiment or aspect, and a pharmaceutically acceptable excipient, diluent or carrier.
[0088] In a tenth configuration, there is provided a kit comprising a pharmaceutical composition comprising an immunocytokine as defined in any other configuration, embodiment or aspect, and a pharmaceutically acceptable excipient, diluent or carrier.
[0089] In an eleventh configuration, there is provided a nucleic acid that encodes a heavy chain and/or a light chain of an immunocytokine as defined in any other configuration, embodiment or aspect.
[0090] In a twelfth configuration, there is provided a vector comprising the nucleic acid that encodes a heavy chain and/or a light chain of an immunocytokine as defined in any other configuration, embodiment or aspect.
[0091] In a thirteenth configuration, there is provided a host comprising the nucleic acid of any other configuration, embodiment or aspect or the vector as defined in any other configuration, embodiment or aspect.
Anti-ICOS Bispecific Antibodies
[0092] In a first configuration, there is provided a multispecific antibody (e.g. bispecific antibody or a dual-binding antibody) which binds (and optionally has specificity for) ICOS (e.g. human ICOS) and another target antigen.
[0093] In a second configuration, there is provided a composition comprising a multispecific, bispecific or dual-binding antibody as described herein and a pharmaceutically acceptable excipient, diluent or carrier.
[0094] In a third configuration, there is provided a multispecific, bispecific or dual-binding antibody as described herein for use in treating or preventing a disease or condition, selected from neurological disease, neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours; such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas).
[0095] In a fourth configuration, there is provided a use of a multispecific, bispecific or dual-binding antibody as described herein in the manufacture of a medicament for administration to a human for treating or preventing a disease or condition in the human selected from neurological disease, neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas).
[0096] In a fifth configuration, there is provided a method of treating or preventing a disease or condition selected from neurological disease, neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas) in a human, comprising administering to said human a therapeutically effective amount of a multispecific, bispecific or dual-binding antibody as described herein, wherein the disease or condition is thereby treated or prevented.
[0097] In a sixth configuration, there is provided a nucleic acid that encodes a heavy chain and/or a light chain of a multispecific antibody as described herein.
[0098] In a seventh configuration, there is provided a vector comprising the nucleic acid that encodes a heavy chain and/or a light chain of a multispecific antibody as described herein.
DESCRIPTION OF THE FIGURES
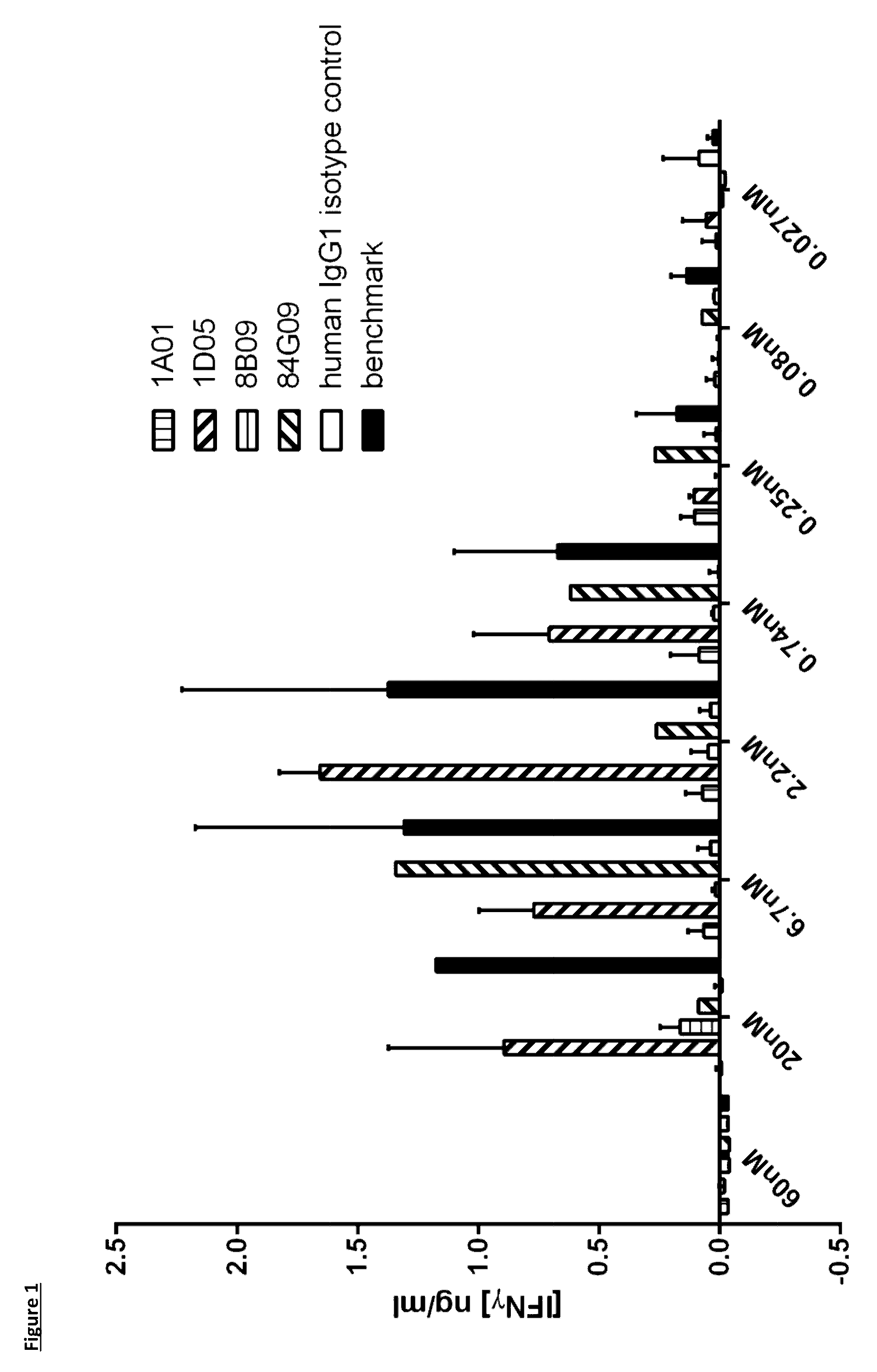
[0099] FIG. 1: Analysis of selected antibodies in a dendritic cell--T-cell mixed lymphocyte reaction. Monocytes were cultured with GM-CSF and IL-4 for seven days, before addition of allogeneic purified CD3.sup.+ T-cells and titrations of antibodies. Supernatants were taken at day 5 for analysis of IFN.gamma. production. Data is shown from one experiment. Note that for 84G09, there is a single point per concentration, as one replicate failed
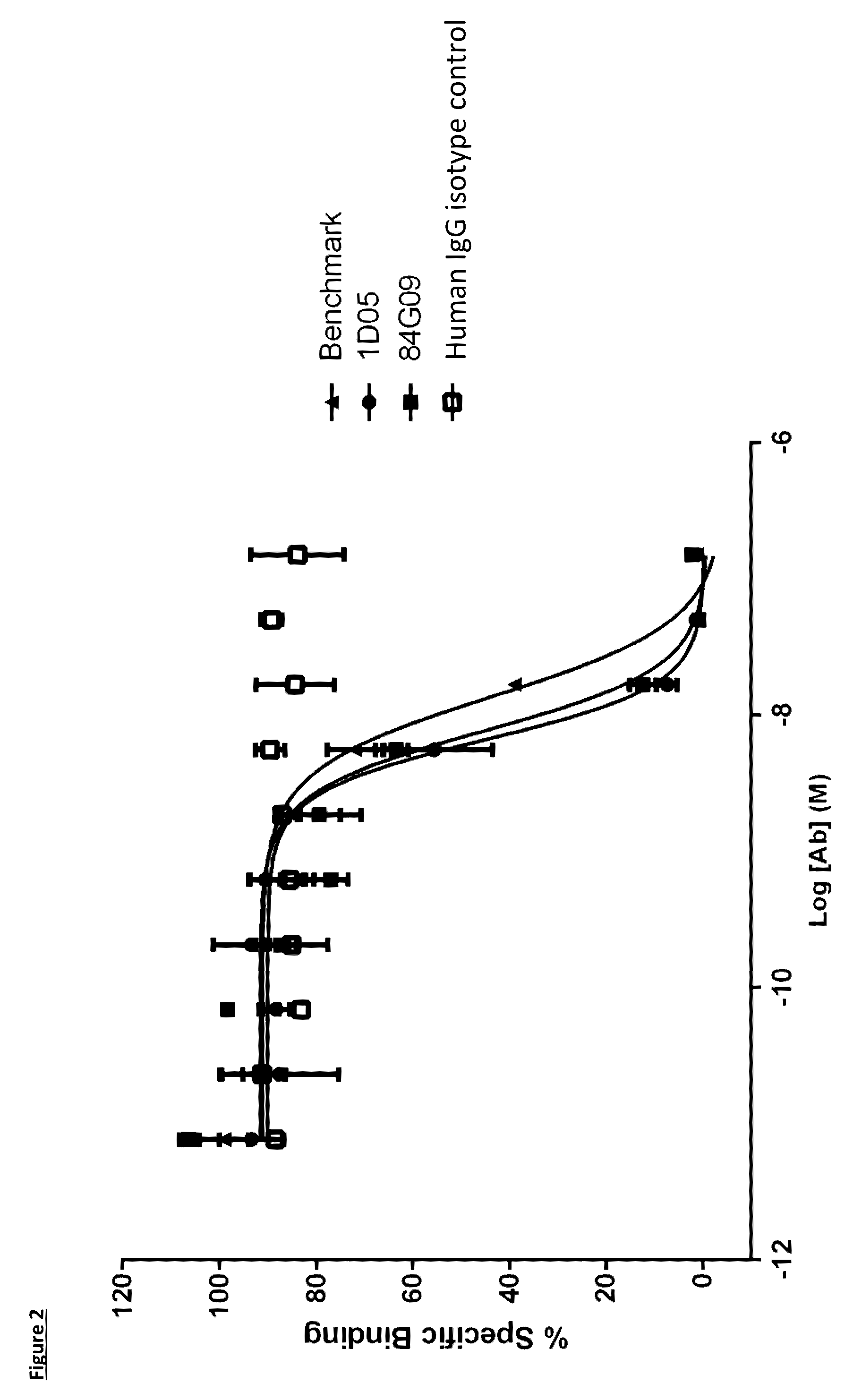
[0100] FIG. 2: PD-L1 direct neutralisation ELISA with PD-1 receptor. Neutralisation profiles of 1D05 and 84G09 compared to a benchmark anti-PD-L1 antibody and isotype control. Data representative of three independent experiments
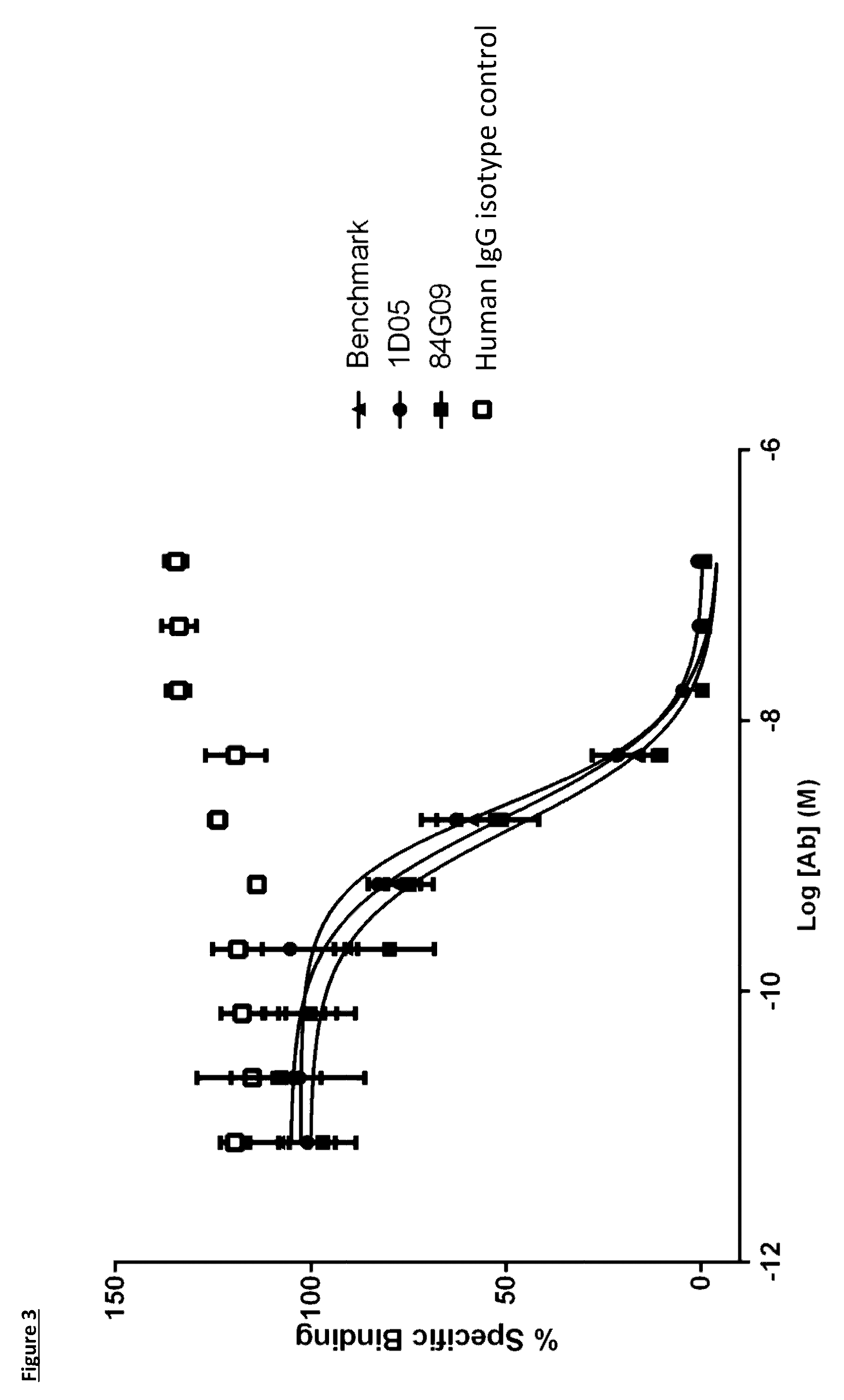
[0101] FIG. 3: Human PD-L1 CHO-S FACS neutralisation with PD-1 receptor. Neutralisation profiles of 1D05 and 84G09 compared to a benchmark anti-PD-L1 antibody and isotype control. Data representative of three independent experiments
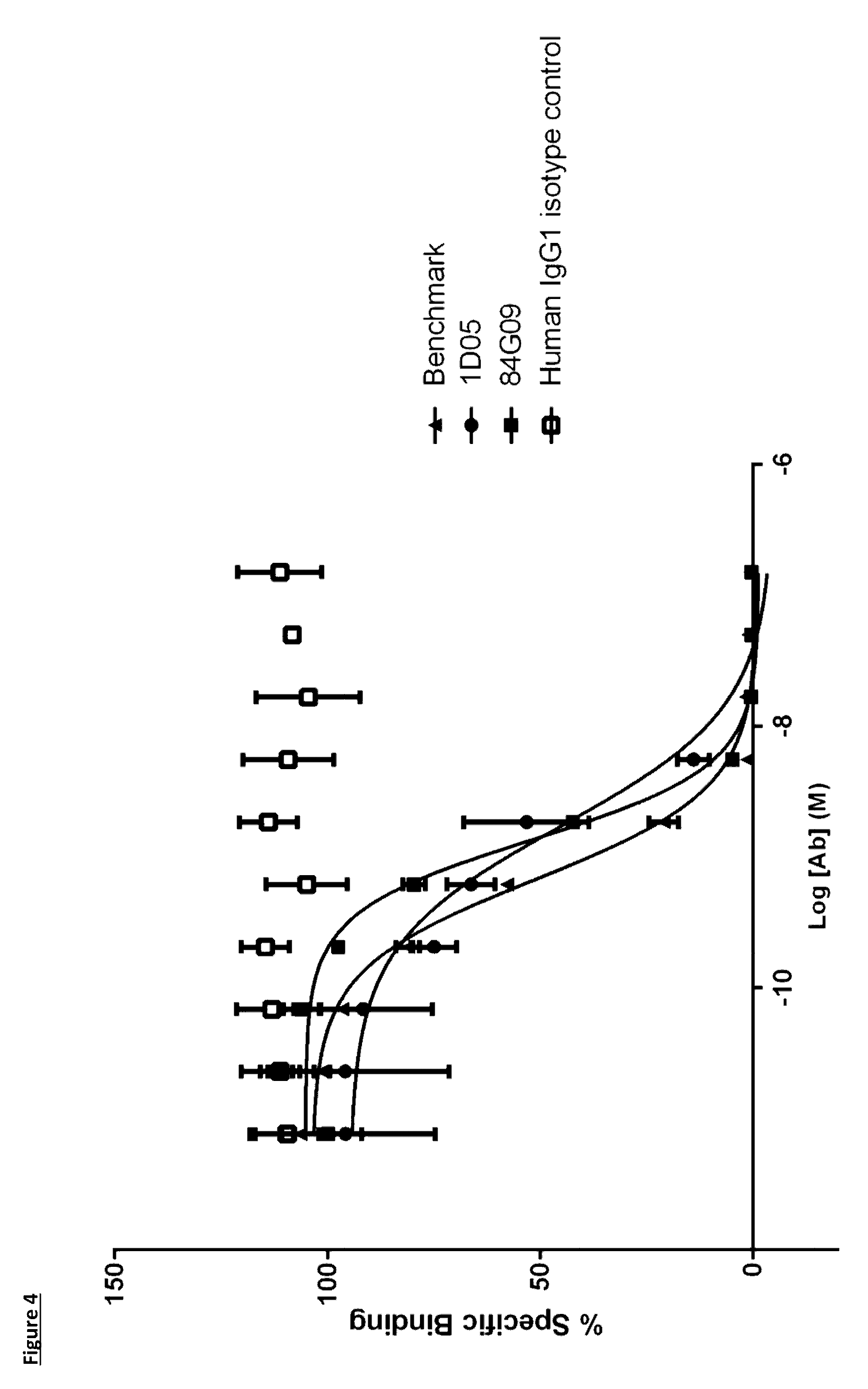
[0102] FIG. 4: Human PD-L1 CHO FACS neutralisation with CD80 receptor. Neutralisation profiles of the 1D05 and 84G09 compared to the benchmark anti-PD-L1 antibody and isotype control. Data representative of three independent experiments
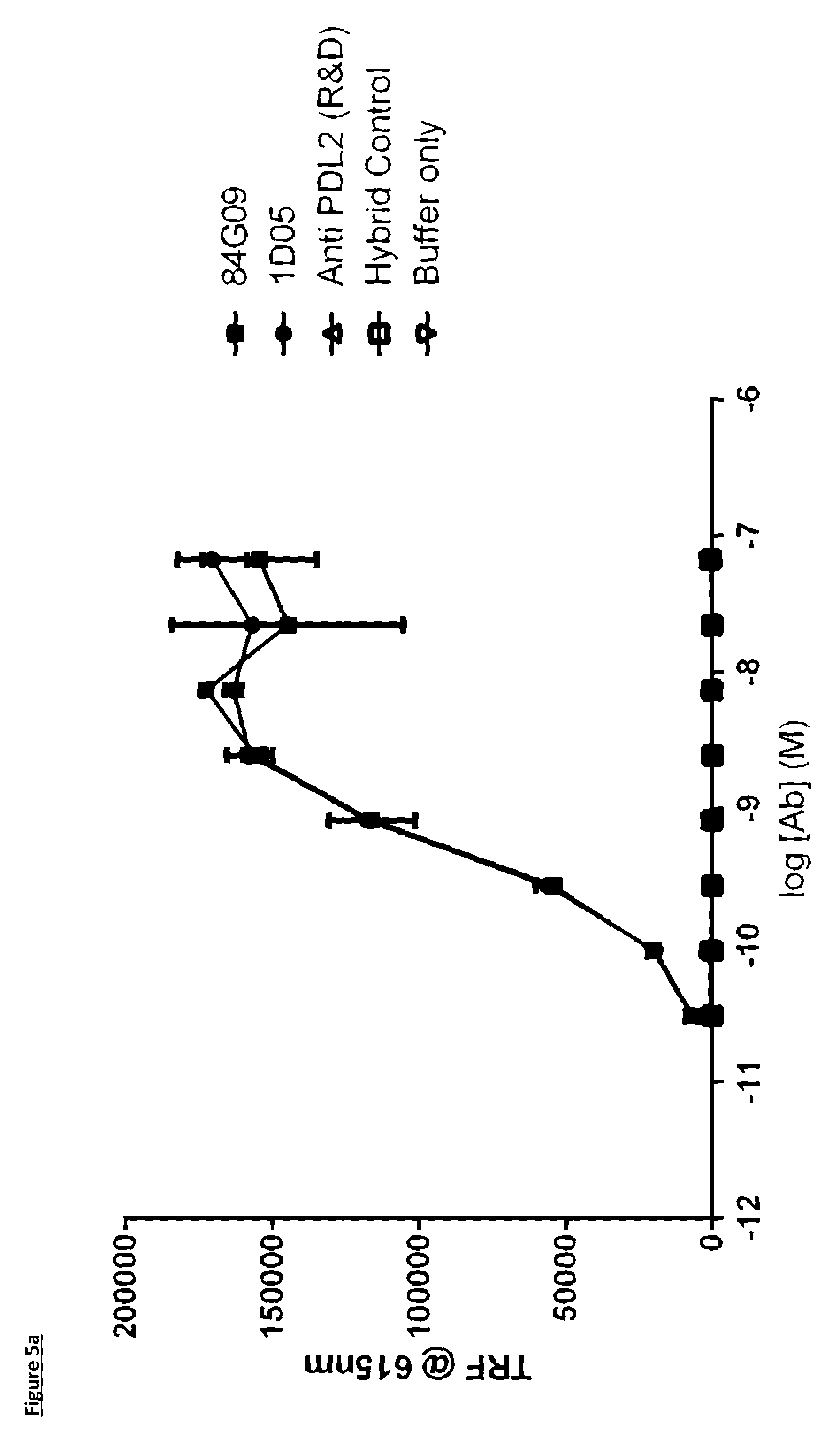
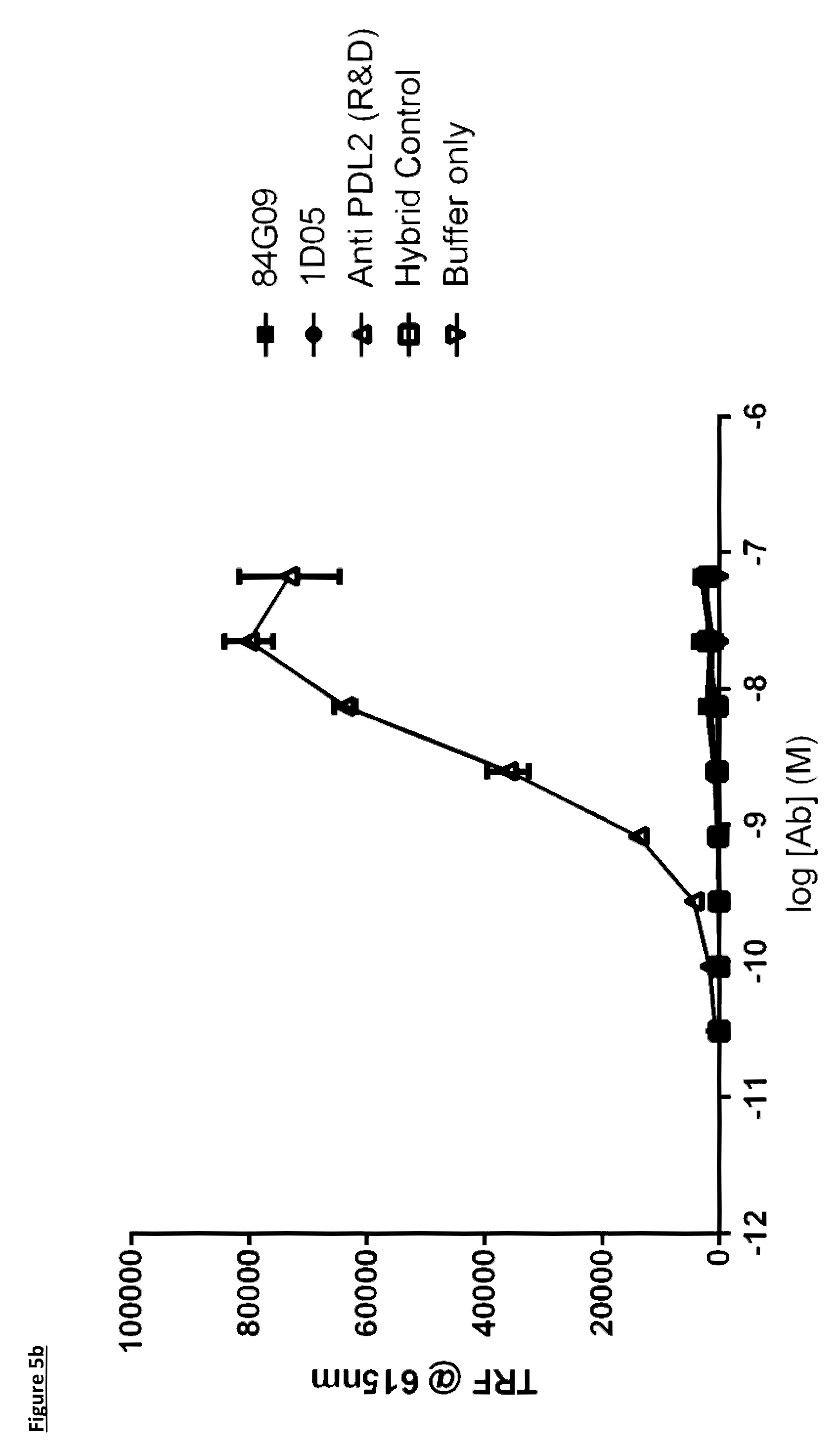
[0103] FIG. 5: Binding of lead antibodies to PD-L1 but not PD-L2. Lead antibodies bind to plate bound PD-L1 (FIG. 5a) but not PD-L2 (FIG. 5b). An anti-PD-L2 antibody was used as a control. Data are expressed as time resolved fluorescence units at 615 nm. Data representative of two independent experiments
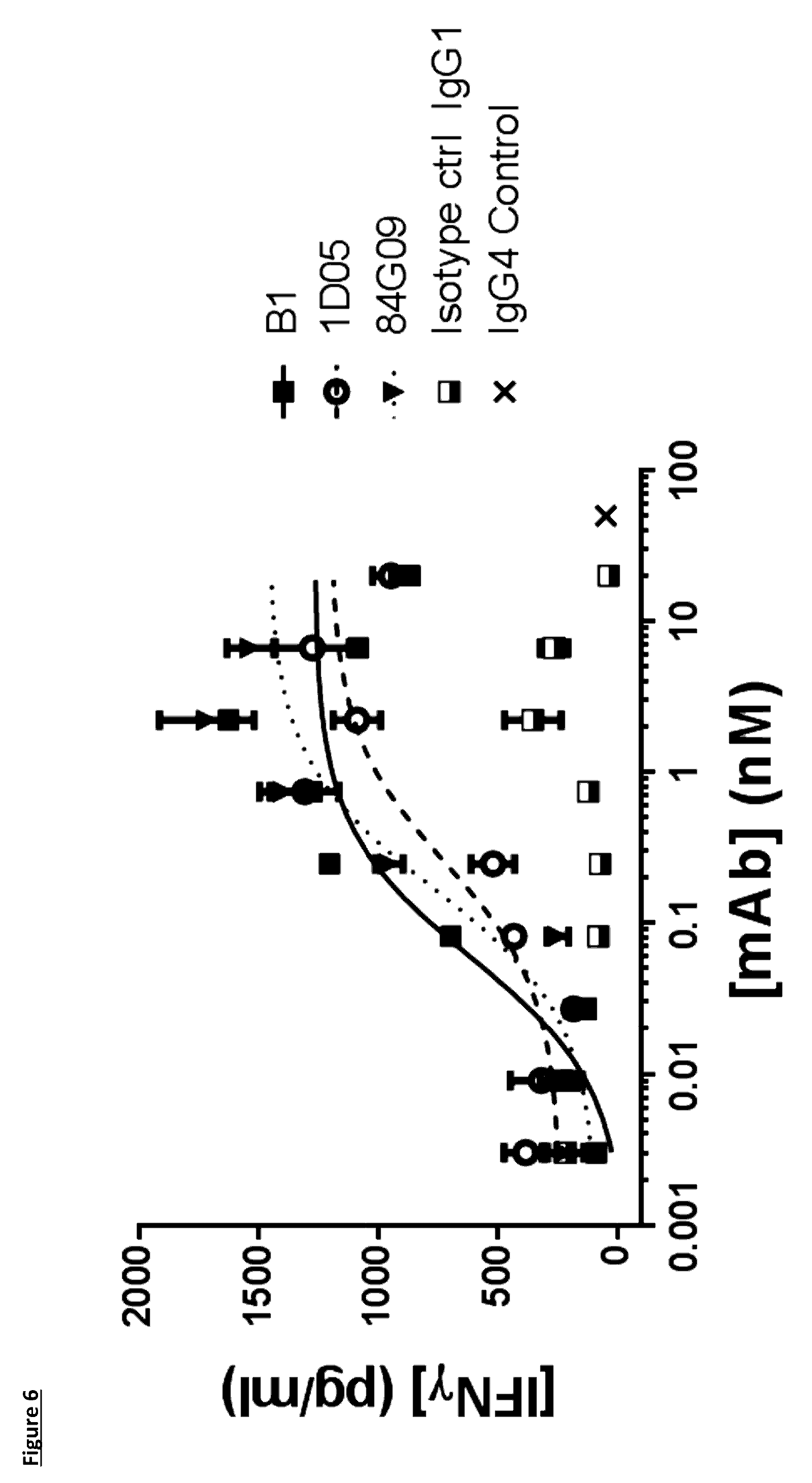
[0104] FIG. 6: Lead antibodies induce IFN.gamma. production in a Dendritic Cell-T-cell mixed lymphocyte reaction. Immature dendritic cells were co-cultured with allogeneic CD4.sup.+ T-cells in the presence of antibodies for 5 days. IFN.gamma. was measured in supernatants by ELISA. Data are representative of three independent experiments. B1 refers to a benchmark antibody
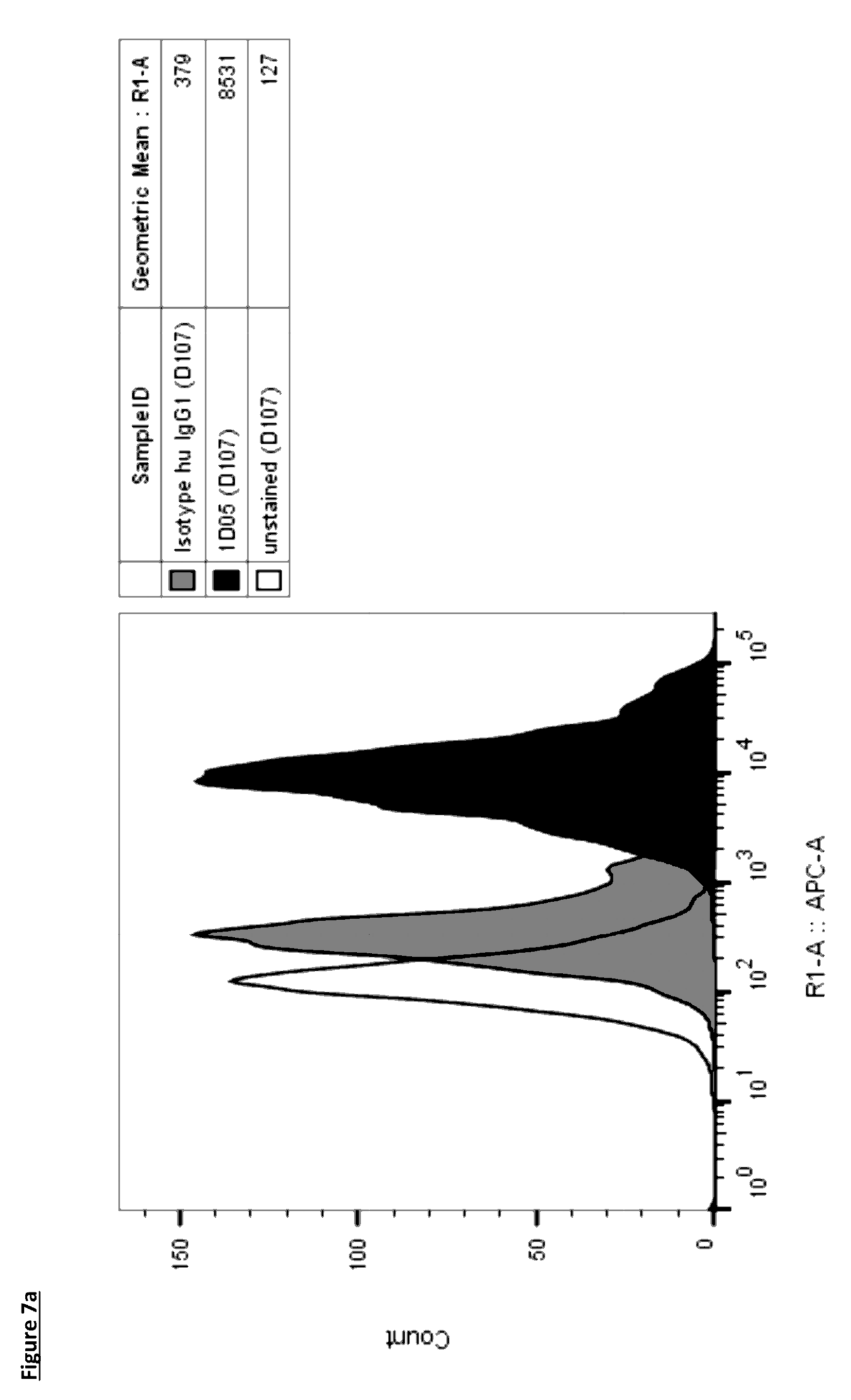
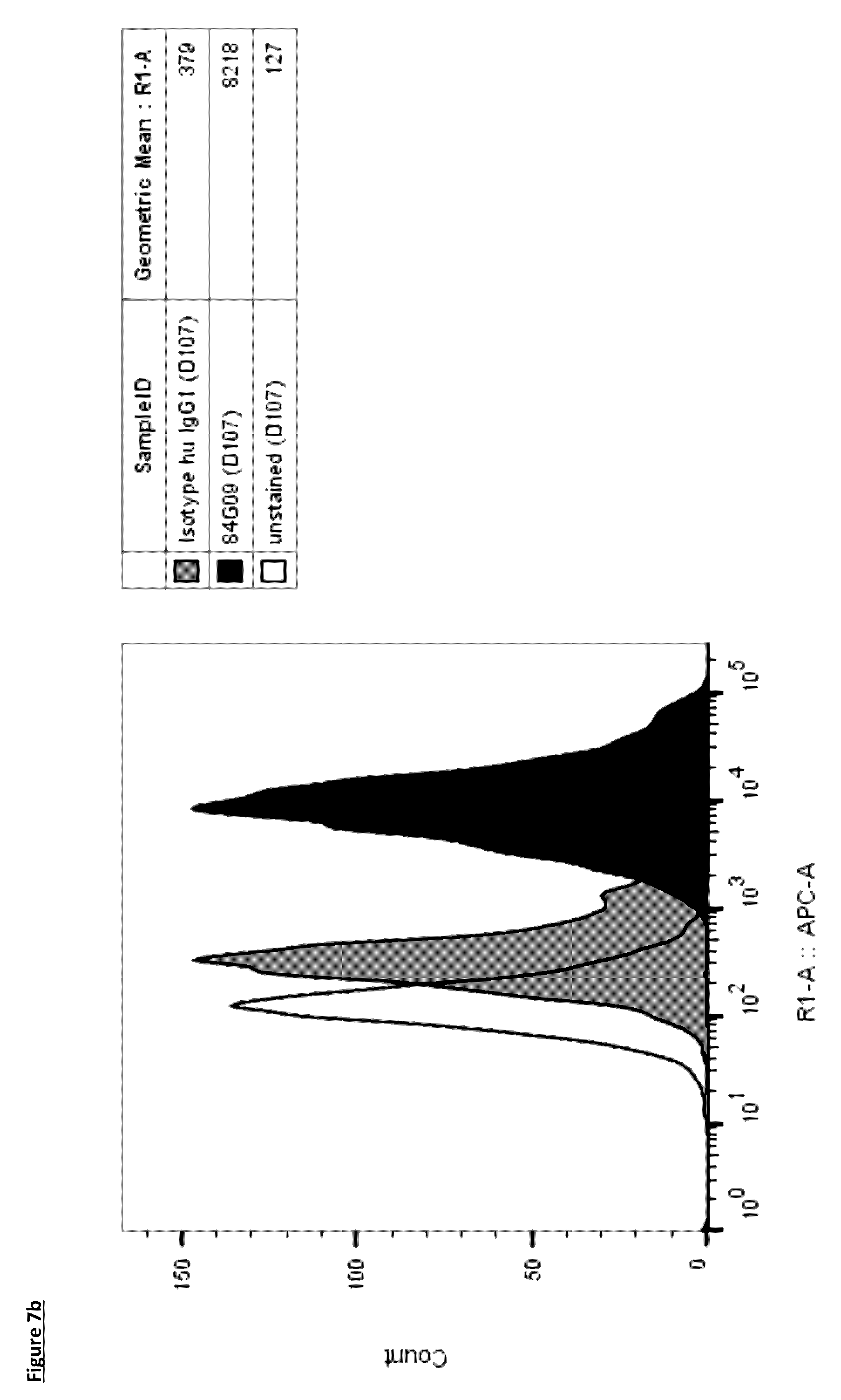
[0105] FIG. 7: Lead antibodies bind to natively expressed PD-L1 on dendritic cells. Dendritic cells were generated from monocyte precursors with GM-CSF and IL-4 and stained with lead antibodies (a) 1D05 and (b) 84G09, and isotype control directly labelled with AlexaFluor647. Data shown is from one blood donor, representative of four donors
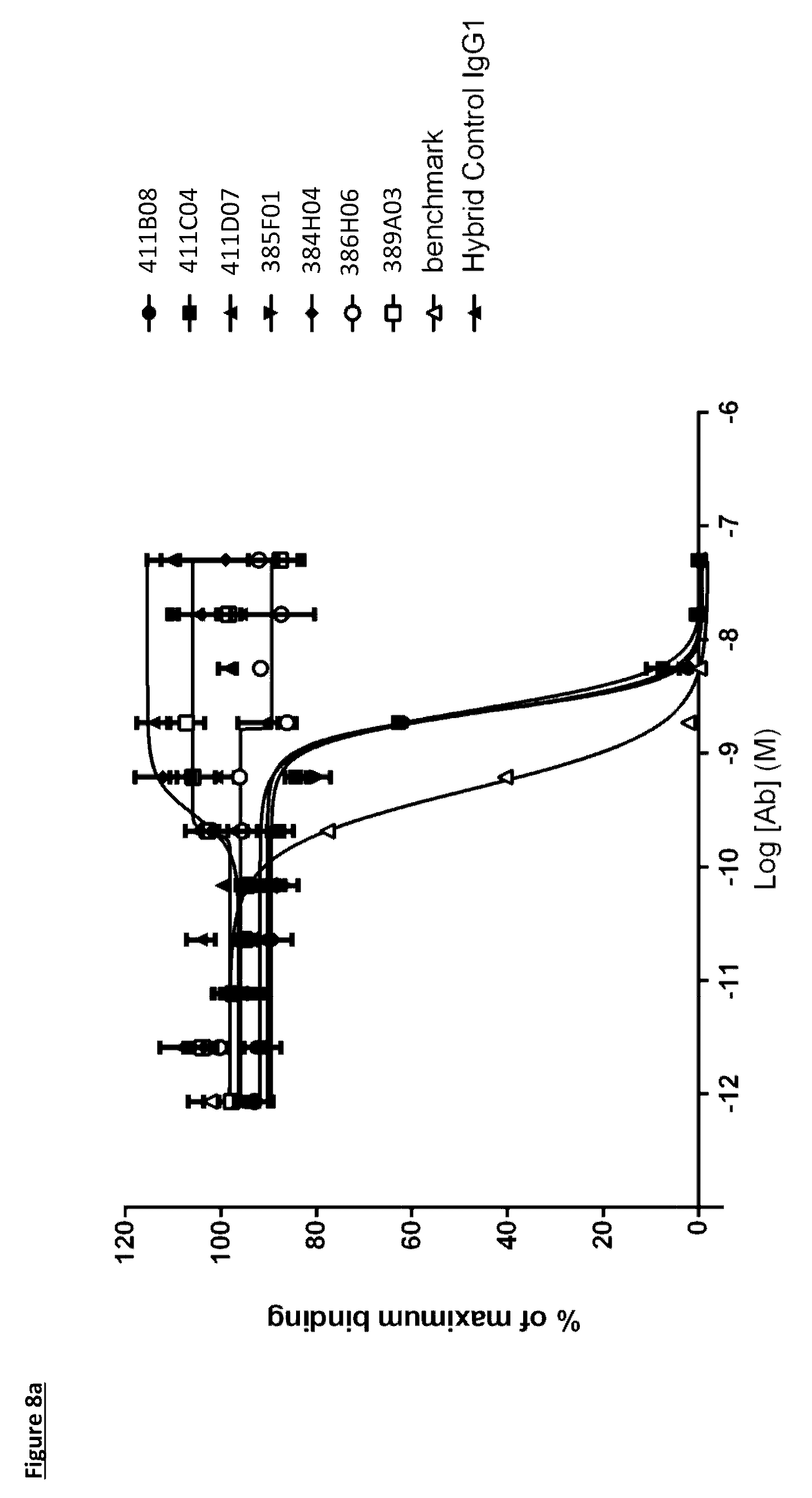
[0106] FIG. 8a: PD-L1 direct neutralisation ELISA with PD-1 receptor. Neutralisation profiles of KM121 hits compared to a benchmark anti-PD-L1 antibody and isotype control. Data representative of three independent experiments
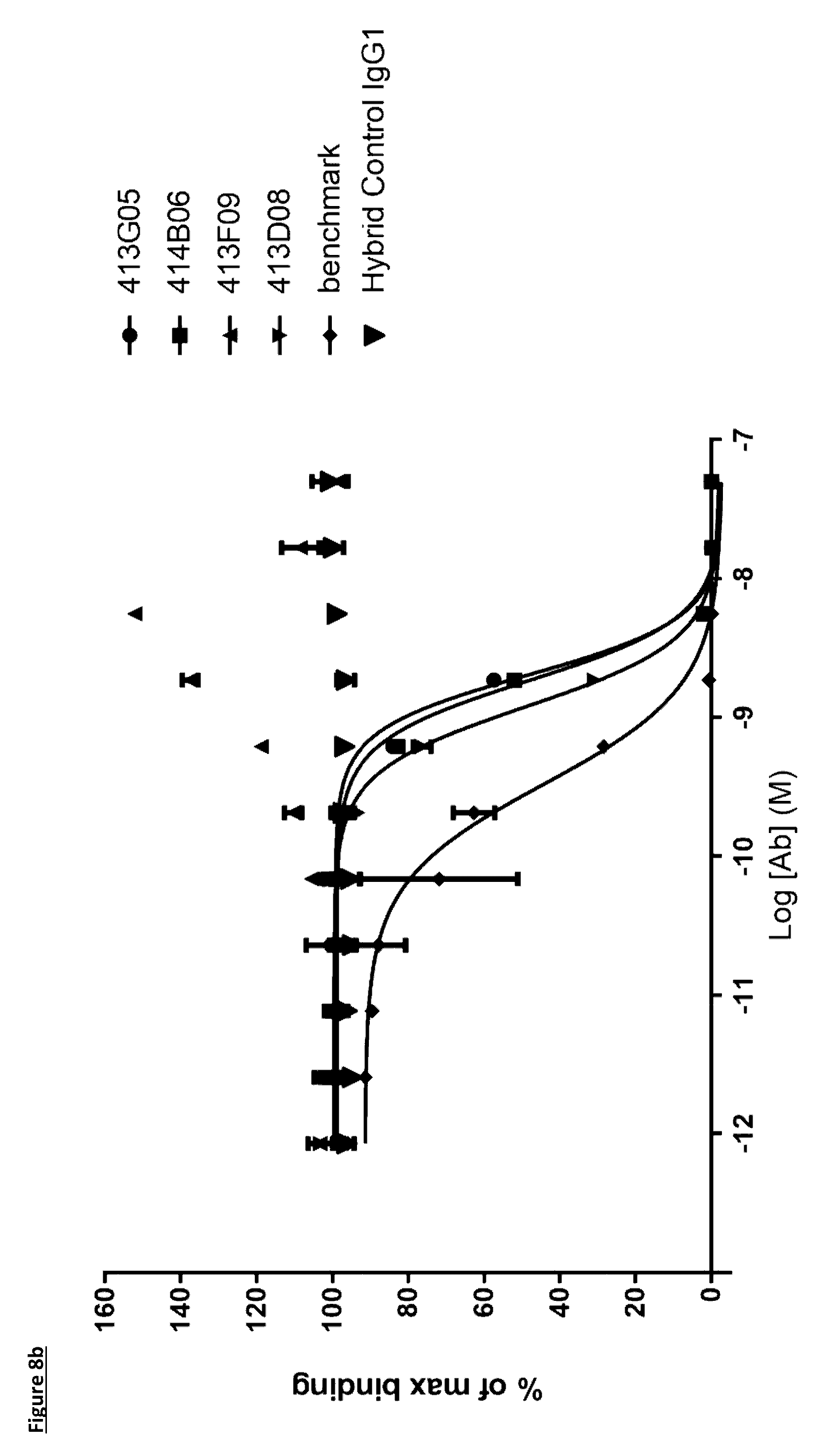
[0107] FIG. 8b: PD-L1 direct neutralisation ELISA with PD-1 receptor. Neutralisation profiles of KM122 lead candidate molecules compared to the benchmark anti-PD-L1 antibody. Data is from a single experiment
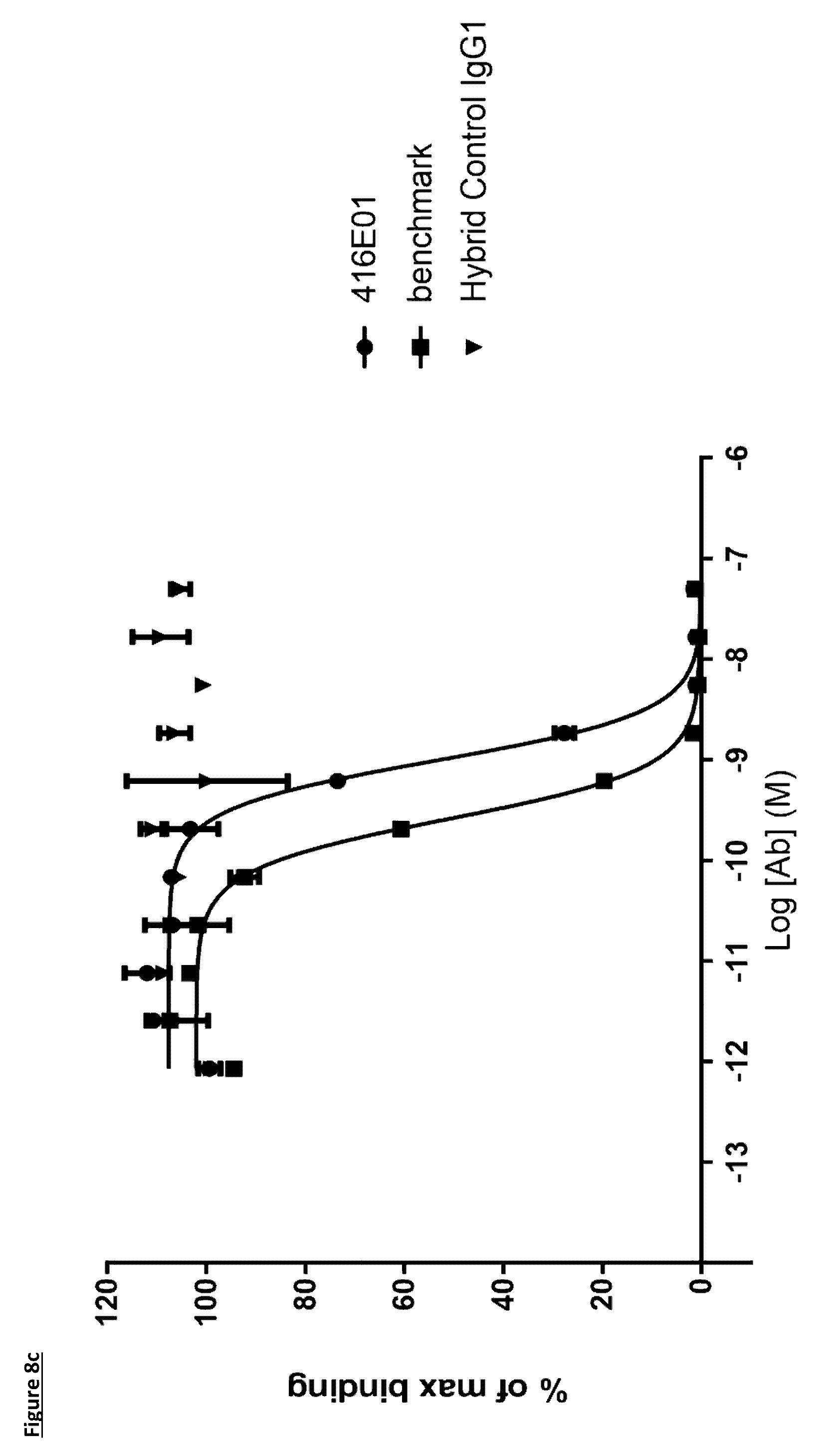
[0108] FIG. 8c: PD-L1 direct neutralisation ELISA with PD-1 receptor. Neutralisation profile of KM122 lead candidate molecule 416E01 compared to the benchmark anti-PD-L1 antibody. Data is from a single experiment
[0109] FIG. 9a: PD-L1 direct neutralisation ELISA with CD80 receptor. Neutralisation profiles of KM121 hits compared to a benchmark anti-PD-L1 antibody and isotype control. Data representative of three independent experiments
[0110] FIG. 9b: PD-L1 direct neutralisation ELISA with CD80 receptor. Neutralisation profiles of KM122 lead candidate molecules compared to the benchmark anti-PD-L1 antibody. Data is from a single experiment
[0111] FIG. 9c: PD-L1 direct neutralisation ELISA with CD80 receptor. Neutralisation profiles of KM122 lead candidate molecule 416E01 compared to the benchmark anti-PD-L1 antibody. Data is from a single experiment
[0112] FIG. 10: Bispecific binding as measured by SPR, with PD-L1 as first antigen, and TIGIT as second antigen. A) Bispecific 1, B) Bispecific 2, C) Bispecific 3, D) Bispecific 4. For detailed construction information of each Bispecific construct, see Table 6
[0113] FIG. 11: Bispecific binding as measured by SPR, with TIGIT as first antigen, and PD-L1 as second antigen. A) Bispecific 1, B) Bispecific 2, C) Bispecific 3, D) Bispecific 4. For detailed construction information of each Bispecific construct, see Table 6
[0114] FIG. 12(a): Ability of immunocytokine constructs to induce proliferation in IL-2R.beta..gamma. expressing TF-1 cells, compared with equimolar concentrations of free IL-2. Data shown is from a single experiment, representative of three experiments
[0115] FIG. 12(b): Ability of immunocytokine constructs to induce proliferation in IL-2R.beta..gamma. expressing TF-1 cells, compared with equimolar concentrations of free IL-2. Data shown is from a single experiment, representative of four experiments
[0116] FIG. 13(a): Capacity of 1D05 antibody to neutralise the interaction between PD-1 and PD L1 is unaffected by the fusion of IL-2 to the antibody, as measured in a neutralisation ELISA. Data shown is from a single experiment, representative of three experiments
[0117] FIG. 13(b): Capacity of 1D05 antibody to neutralise the interaction between CD80 and PD-L1 is unaffected by the fusion of IL-2 to the antibody, as measured in a neutralisation ELISA. Data shown is from a single experiment, representative of three experiments
[0118] FIG. 14: Mean group and individual animal growth curves for the NOD/SCID: Xenograft in vivo efficacy study
[0119] FIG. 14(a): shows the group mean (n=8/9) tumour growth curve, for this graph when an animal is removed from the study due to tumour size, the last reading is used for the rest of the study. The shaded area shows the area where the last reading is being used
[0120] FIGS. 14(b) to (e): show the individual animal tumour growth curves for each group. (b) A375 tumours alone; (c) A375 tumours co-injected with CD4.sup.+/8.sup.+ T-cells at a 6:1 ratio. For panels (d) and (e) A375 tumour cell were co-injected with CD4.sup.+/8.sup.+ T-cells at a 6:1 ratio; (d) Isotype Control antibody at 10 mg/kg and (e) anti-PD-L1 antibody 1D05 at 10 mg/kg. Dosing was at 1-hour post tumour/T-cell implantation and on days 3, 6, 8 and 10, shown on the graph by the dotted lines
[0121] FIG. 15: Kaplan-Meier plot for the NOD/SCID:Xenograft in vivo efficacy study showing the number of animals still on study. This plot shows the slight increase in the time on study when CD4.sup.+/CD8.sup.+ T-cells are co-injected with the tumour cells (T-cells/A375) (n=9) when compared to the tumour cells alone A375 group (n=9). Treatment with the isotype control (T-cells/A375--Isotype (n=8)) had no effect on survival when compared to the T-cells co-injected with the tumour cells without antibody. Treatment with 10 mg/kg of the anti-PD-L1 antibody 1D05 (T-cells/A375--anti-PD-L1) (n=8)) significantly increased the time on study when compared to the isotype control group. Dosing was 1-hour post injection of the T-cells/tumour cells and on days 3, 6, 8 and 10, show on the graph by the dotted lines
[0122] FIG. 16: Expansion of lymphocytes in response to dosing with immunocytokines. Fasting blood samples were taken into EDTA treated tubes pre-treatment (0), and 2, 5 and 7 days post-treatment. Cell counts were measured by the Bayer Advia 120. Results are expressed as fold change in lymphocyte count
[0123] FIG. 17: Analysis of standard haematological parameters in response to dosing with immunocytokines. Fasting blood samples were taken into EDTA treated tubes pre-treatment and 7 days post-treatment. Analysis of haemoglobin, haematocrit, red blood cell counts and platelet counts were performed using the Bayer Advia 120. Results are expressed as the percentage change in parameter 7 days post-dosing
[0124] FIG. 18: Cytokine levels in plasma of cynomolgus monkeys dosed with immunocytokine molecules. Plasma samples were obtained pre-treatment (PT) and 3 days after dosing (D3) and analysed by MSD for levels of a) TNF-.alpha.; b) IL-8; c) IL-6; d) IFN.gamma.; e) G-CSF and f) IL-2. Where no bar is included, cytokine levels were below the limit of quantification of the assay. IL4, IL-5 and IL-1.beta. were not detectable in any sample at either timepoint and so are not included in the graphs
[0125] FIG. 19: Levels of soluble CD25 in plasma of cynomolgus monkeys dosed with immunocytokine molecules. Plasma samples were obtained pre-treatment (PT) and 3 days after dosing (D3) and analysed using a commercial ELISA kit. * indicates levels above limit of quantification (20,000 pg/mL)
[0126] FIG. 20: Flow cytometric analysis of PBMC subsets. Whole blood was stained for markers of a) T-cells and b) B-cells, NK cells, neutrophils and monocytes, prior to red blood cell lysis and fixation. Data is expressed as the fold change in cell number 5 days after dosing. Data for 1D05 LC D9-7 ICK is missing due to unusable sample
[0127] FIG. 21: Pharmacokinetic (PK) analysis of immunocytokines. Serum was prepared from blood samples taken at various time points over 96 hours. In panels a) and b), serum was incubated on plates coated with PD-L1 and immunocytokines detected with a biotinylated anti-human Fc detection antibody, and streptavidin-labelled Europium. In panels c) and d), serum was incubated on plates coated with PD-L1 and immunocytokines detected with a biotinylated anti-human IL-2 antibody, and streptavidin-labelled Europium. Results are expressed as ng/mL
[0128] FIG. 22a: Induction of IFN.gamma. production in a monocyte--T-cell co-culture assay by anti-PD-L1 antibodies in human IgG1 format. Each data point represents the mean fold induction from at least three independent experiments, .+-.standard error of the mean
[0129] FIG. 22b: Induction of IFN.gamma. production in a monocyte--T-cell co-culture assay by an anti-PD-L1 antibody in human IgG4(PE) format. Each data point represents the mean fold induction from two independent experiments, .+-.standard deviation
[0130] FIG. 23(a): Induction of IL-2 in a murine T-cell hybridoma assay. Human PD-L1 transfected LK35.2 cells were loaded with ovalbumin peptide and co-cultured overnight with DO-11-10 T-cell hybridoma cells in the presence of anti-PD-L1 antibodies or controls, prior to collection of supernatants and analysis of IL-2 release. Each data point indicates background-corrected mean IL-2 release from three independent experiments.+-.standard deviation
[0131] FIG. 23(b): Induction of IL-2 in a murine T-cell hybridoma assay. Human PD-L1 transfected LK35.2 cells were loaded with ovalbumin peptide and co-cultured overnight with DO-11-10 T-cell hybridoma cells in the presence of ICOS/PD-L1 bi-specific molecules, or individual antibodies, prior to collection of supernatants and analysis of IL-2 release. Each data point indicates background-corrected mean IL-2 release from three independent experiments.+-.standard deviation
[0132] FIG. 24(a): Induction of IFN.gamma., in a DC--T-cell MLR assay. Monocyte derived dendritic cells (DC) were activated with E. coli LPS and co-cultured with allogeneic CD3.sup.+ T-cells at a 1:1 ratio. IFN.gamma., was measured by DELFIA assay after 5 days of co-culture. Data is from a single experiment
[0133] FIG. 24(b): Induction of IL-2 in a DC--T-cell MLR assay. Monocyte derived dendritic cells (DC) were activated with E. coli LPS and co-cultured with allogeneic CD3.sup.+ T-cells at a 1:1 ratio. IL-2 was measured by DELFIA assay after 3 days of co-culture. Data is from a single experiment
[0134] FIG. 25: Titration of FIT-Ig molecules, parental monospecific antibodies, and control antibodies in PD-L1/TIGIT AlphaScreen.RTM. Binding Assay using method one. Antibodies were incubated with PD-L1 and TIGIT proteins for an hour before the addition of AlphaScreen.RTM. acceptor beads for an hour followed by the addition of AlphaScreen.RTM. donor beads for another hour prior to the detection of fluorescence. A) Titration of FIT-Ig molecules; B) Titration of monospecific antibodies. Data shown are representative of one unique experiment
[0135] FIG. 26: Titration of FIT-Ig molecules, parental monospecific antibodies, and control antibodies in PD-L1/TIGIT AlphaScreen.RTM. Binding Assay using method two. AlphaScreen.RTM. donor and acceptor beads were coated for an hour with PD-L1 and TIGIT proteins respectively before the addition of antibodies for an hour followed by the detection of fluorescence. A) Titration of FIT-Ig molecules; B) Titration of monospecific antibodies. Data shown are representative of one unique experiment
[0136] FIG. 27: Titration of FIT-Ig molecules, and control antibody in a PD-L1/TIGIT cell recruitment assay by flow cytometry. CHO human PD-L1 and HEK human TIGIT were stained with CellTrace.TM. Far Red and CellTrace.TM. Violet respectively and co-cultured in presence of antibodies for an hour prior to the detection of fluorescence and identification of double positive population. Data shown are representative of one unique experiment
[0137] FIG. 28: Expansion of lymphocytes in response to dosing with immunocytokines. Fasting blood samples were taken into EDTA treated tubes pre-treatment (0), and 2, 5 and 7, 10, 14 and 23 days post-treatment. Cell counts were measured by the Bayer Advia 120. Results are expressed as fold change in lymphocyte count
[0138] FIG. 29: Levels of soluble CD25 in plasma of cynomolgus monkeys dosed with immunocytokine molecules. Plasma samples were obtained pre-treatment (0) and 3, 7 and 10 days after dosing and analysed using a commercial ELISA kit
[0139] FIG. 30: Analysis of standard haematological parameters in response to dosing with immunocytokines. Fasting blood samples were taken into EDTA treated tubes pre-treatment (0) and 2, 5, 7, 10, 14 and 23 days post-treatment. Analysis of A) haemoglobin, B) haematocrit, C) red blood cell counts and D) platelet counts were performed using the Bayer. Advia 120. Results are expressed as the fold change in parameter at each timepoint
[0140] FIG. 31: Cytokine levels in plasma of cynomolgus monkeys dosed with immunocytokine molecules. Plasma samples were obtained pre-treatment (0) and 1, 3, 7, 10, 14 and 23 days after dosing and analysed by MSD for levels of a) TNF-.alpha.; b) IL-8; c) IL-6; d) IFN.gamma.; e) G-CSF, f) IL-2, g) IL-4 and h) IL-5. Where no bar is included, cytokine levels were below the limit of quantification of the assay. IL-113 was not detectable in any samples and so is not included in the graphs
[0141] FIG. 32: Pharmacokinetic (PK) analysis of immunocytokines. Serum was prepared from blood samples taken at various time points over 96 hours. Serum was incubated on plates coated with PD-L1 and immunocytokines detected with a biotinylated anti-human Fc detection antibody, and streptavidin-labelled Europium. Results are expressed as % peak concentration
[0142] FIG. 33: Expansion of specific T-cell subsets by ICK molecules. Whole blood was incubated with antibodies for staining, before red blood cell lysis, fixation and analysis by flow cytometry. Results are expressed as fold change in absolute (a) CD4.sup.+ T-cell and (b) CD8.sup.+ T-cell numbers at each timepoint.
[0143] FIG. 34: Effector function of lead antibodies in a reporter cell assay. PD-L1 expressing target cells (ES2) were co-cultured overnight with Jurkat cells, engineered to express NFAT-induced luciferase and Fc.gamma.RIIIa, in the presence of PD-L1 antibodies. Each data point indicates mean fold induction of relative light units.+-.standard deviation. Data is from one representative experiment, of three independent experiments
[0144] FIG. 35: Binding of lead antibodies to cell-expressed cynomolgus PD-L1. Antibodies were titrated on CHO cells expressing cynomolgus PD-L1, and bound antibody detected with an anti-human IgG AlexaFluor 647. Data is from a single experiment
[0145] FIG. 36(a): Human PD-L1 CHO-S FACS neutralisation with PD-1 receptor. Neutralisation profiles of lead antibodies compared to a benchmark anti-PD-L1 antibody and isotype control. Data representative of two independent experiments
[0146] FIG. 36(b): Human PD-L1 CHO-S FACS neutralisation with CD80 receptor. Neutralisation profiles of lead antibodies compared to a benchmark anti-PD-L1 antibody and isotype control. Data representative of two independent experiments
[0147] FIG. 37: Induction of IFN.gamma. production in a monocyte--T-cell co-culture assay by anti-PD-L1 antibodies in human IgG1 format. Each data point represents the mean fold induction of IFN.gamma. from at least three independent experiments, .+-.standard error of the mean
DETAILED DESCRIPTION
1. Definitions
[0148] Unless otherwise defined herein, scientific and technical terms shall have the meanings that are commonly understood by those of ordinary skill in the art. Further, unless otherwise required by context, singular terms shall include pluralities and plural terms shall include the singular.
[0149] The singular terms "a," "an," and "the" include plural referents unless context clearly indicates otherwise. Similarly, the word "or" is intended to include "and" unless the context clearly indicates otherwise. Although methods and materials similar or equivalent to those described herein can be used in the practice or testing of this disclosure, suitable methods and materials are described below. The abbreviation, "e.g." is derived from the Latin exempli gratia, and is used herein to indicate a non-limiting example. Thus, the abbreviation "e.g." is synonymous with the term "for example."
[0150] In the specification and claims, the term "about" is used to modify, for example, the quantity of an ingredient in a composition, concentration, volume, process temperature, process time, yield, flow rate, pressure, and like values, and ranges thereof, employed in describing the embodiments of the disclosure. The term "about" refers to variation in the numerical quantity that can occur, for example, through typical measuring and handling procedures used for making compounds, compositions, concentrates or use formulations; through inadvertent error in these procedures; through differences in the manufacture, source, or purity of starting materials or ingredients used to carry out the methods, and like proximate considerations. The term "about" also encompasses amounts that differ due to aging of a formulation with a particular initial concentration or mixture, and amounts that differ due to mixing or processing a formulation with a particular initial concentration or mixture. Where modified by the term "about" the claims appended hereto include equivalents to these quantities.
[0151] As used herein, "administer" or "administration" refers to the act of injecting or otherwise physically delivering a substance as it exists outside the body (e.g., an anti-hPD-L1 antibody provided herein) into a patient, such as by mucosal, intradermal, intravenous, intramuscular delivery and/or any other method of physical delivery described herein or known in the art. When a disease, or a symptom thereof, is being treated, administration of the substance typically occurs after the onset of the disease or symptoms thereof. When a disease, or symptoms thereof, are being prevented, administration of the substance typically occurs before the onset of the disease or symptoms thereof.
[0152] The term "antibody", "immunoglobulin" or "Ig" may be used interchangeably herein and means an immunoglobulin molecule that recognizes and specifically binds to a target, such as a protein, polypeptide, peptide, carbohydrate, polynucleotide, lipid, or combinations of the foregoing through at least one antigen recognition site within the variable region of the immunoglobulin molecule. As used herein, the term "antibody" encompasses intact polyclonal antibodies, intact monoclonal antibodies, antibody fragments (such as Fab, Fab', F(ab').sub.2, and Fv fragments), single chain Fv (scFv) mutants, multispecific antibodies such as bispecific antibodies (including dual binding antibodies), chimeric antibodies, humanized antibodies, human antibodies, fusion proteins comprising an antigen determination portion of an antibody, and any other modified immunoglobulin molecule comprising an antigen recognition site so long as the antibodies exhibit the desired biological activity. The term "antibody" can also refer to a Y-shaped glycoprotein with a molecular weight of approximately 150 kDa that is made up of four polypeptide chains: two light (L) chains and two heavy (H) chains. There are five types of mammalian Ig heavy chain isotypes denoted by the Greek letters alpha (.alpha.), delta (.delta.), epsilon (.epsilon.), gamma (.gamma.), and mu (.mu.). The type of heavy chain defines the class of antibody, i.e., IgA, IgD, IgE, IgG, and IgM, respectively. The .gamma. and .alpha. classes are further divided into subclasses on the basis of differences in the constant domain sequence and function, e.g., IgG1, hIgG2, mIgG2A, mIgG2B, IgG3, IgG4, IgA1 and IgA2. In mammals, there are two types of immunoglobulin light chains, A and K. The "variable region" or "variable domain" of an antibody refers to the amino-terminal domains of the heavy or light chain of the antibody. The variable domains of the heavy chain and light chain may be referred to as "V.sub.H" and "V.sub.L", respectively. These domains are generally the most variable parts of the antibody (relative to other antibodies of the same class) and contain the antigen binding sites.
[0153] The antibodies described herein may be oligoclonal, polyclonal, monoclonal (including full-length monoclonal antibodies), camelised, chimeric, CDR-grafted, multi-specific, bi-specific (including dual-binding antibodies), catalytic, chimeric, humanized, fully human, anti-idiotypic, including antibodies that can be labelled in soluble or bound form as well as fragments, variants or derivatives thereof, either alone or in combination with other amino acid sequences provided by known techniques. An antibody may be from any species. Antibodies described herein can be naked or conjugated to other molecules such as toxins, radioisotopes, etc.
[0154] The term "antigen binding domain," "antigen binding region," "antigen binding fragment," and similar terms refer to that portion of an antibody which comprises the amino acid residues that interact with an antigen and confer on the binding agent its specificity and affinity for the antigen (e.g. the complementarity determining regions (CDRs)). The antigen binding region can be derived from any animal species, such as rodents (e.g. rabbit, rat or hamster) and humans. Preferably, the antigen binding region will be of human origin.
[0155] Antigen binding fragments described herein can include single-chain Fvs (scFv), single-chain antibodies, single domain antibodies, domain antibodies, Fv fragments, Fab fragments, F(ab') fragments, F(ab').sub.2 fragments, antibody fragments that exhibit the desired biological activity, disulfide-stabilised variable region (dsFv), dimeric variable region (diabody), anti-idiotypic (anti-Id) antibodies (including, e.g. anti-Id antibodies to antibodies), intrabodies, linear antibodies, single-chain antibody molecules and multispecific antibodies formed from antibody fragments and epitope-binding fragments of any of the above. In particular, antibodies and antibody fragments described herein can include immunoglobulin molecules and immunologically active fragments of immunoglobulin molecules, i.e., molecules that contain an antigen-binding site. Digestion of antibodies with the enzyme, papain, results in two identical antigen-binding fragments, known also as "Fab" fragments, and a "Fc" fragment, having no antigen-binding activity but having the ability to crystallize. "Fab" when used herein refers to a fragment of an antibody that includes one constant and one variable domain of each of the heavy and light chains. The term "Fc region" herein is used to define a C-terminal region of an immunoglobulin heavy chain, including native-sequence Fc regions and variant Fc regions. The "Fc fragment" refers to the carboxy-terminal portions of both H chains held together by disulfides. The effector functions of antibodies are determined by sequences in the Fc region, the region which is also recognized by Fc receptors (FcR) found on certain types of cells. Digestion of antibodies with the enzyme, pepsin, results in a F(ab').sub.2 fragment in which the two arms of the antibody molecule remain linked and comprise two-antigen binding sites. The F(ab').sub.2 fragment has the ability to crosslink antigen.
[0156] "Fv" when used herein refers to the minimum fragment of an antibody that retains both antigen-recognition and antigen-binding sites. This region consists of a dimer of one heavy and one light chain variable domain in tight, non-covalent or covalent association. It is in this configuration that the three CDRs of each variable domain interact to define an antigen-binding site on the surface of the V.sub.H-V.sub.L dimer. Collectively, the six CDRs confer antigen-binding specificity to the antibody. However, even a single variable domain (or half of an Fv comprising only three CDRs specific for an antigen) has the ability to recognize and bind antigen, although at a lower affinity than the entire binding site.
[0157] The term "monoclonal antibody" as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, i.e. the individual antibodies comprising the population are identical except for possible naturally occurring mutations and/or post-translation modifications (e.g. isomerizations, amidations) that may be present in minor amounts. Monoclonal antibodies are highly specific, and are directed against a single antigentic determinant or epitope. In contrast, polyclonal antibody preparations typically include different antibodies directed against different antigenic determinants (or epitopes). The term "monoclonal antibody" as used herein encompasses both intact and full-length monoclonal antibodies as well as antibody fragments (such as Fab, Fab', F(ab').sub.2, Fv), single chain (scFv) mutants, fusion proteins comprising an antibody portion, and any other modified immunoglobulin molecule comprising an antigen recognition site. Furthermore, "monoclonal antibody" refers to such antibodies made in any number of ways including, but not limited to, hybridoma, phage selection, recombinant expression, and transgenic animals.
[0158] The monoclonal antibodies herein can include "chimeric" antibodies (immunoglobulins) in which a portion of the heavy and/or light chain is identical with or homologous to corresponding sequences in antibodies derived from a particular species or belonging to a particular antibody class or subclass, while the remainder of the chain(s) is(are) identical with or homologous to corresponding sequences in antibodies derived from another species or belonging to another antibody class or subclass, as well as fragments of such antibodies that exhibit the desired biological activity.
[0159] The term "humanized antibody" refers to a subset of chimeric antibodies in which a "hypervariable region" from a non-human immunoglobulin (the donor antibody) replaces residues from a hypervariable region in a human immunoglobulin (recipient antibody). In general, a humanized antibody will include substantially all of at least one, and typically two, variable domains, in which all or substantially all of the hypervariable loops correspond to those of a non-human immunoglobulin sequence, and all or substantially all of the framework regions are those of a human immunoglobulin sequence, although the framework regions may include one or more substitutions that improve antibody performance, such as binding affinity, isomerization, immunogenicity, etc.
[0160] The term "bispecific antibody" means an antibody which comprises specificity for two target molecules, and includes, but is not limited to, formats such as DVD-Ig (see DiGiammarino et al., "Design and generation of DVD-Ig.TM. molecules for dual-specific targeting", Meth. Mo. Biol., 2012, 889, 145-156), mAb.sup.2 (see WO2008/003103, the description of the mAb.sup.2 format is incorporated herein by reference), FIT-Ig (see WO2015/103072, the description of the FIT-Ig scaffold is incorporated herein by reference), mAb-dAb, dock and lock, Fab-arm exchange, SEEDbody, Triomab, LUZ-Y, Fcab, .kappa..lamda.-body, orthogonal Fab, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CH3, Diabody-CH3, Triple body, Miniantibody, minibody, TriBi minibody, scFv-CH3 KIH, scFv-CH-CL-scFv, F(ab')2-scFv, scFv-KIH, Fab-scFv-Fc, tetravalent HCab, ImmTAC, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, DT-IgG, DutaMab, IgG(H)-scFv, scFv-(H)IgG, IgG(L)-scFv, scFv-(L)IgG, IgG(L,H)-Fv, IgG(H)-V, V(H)--IgG, IgG(L)-V, V(L)-IgG, KIH IgG-scFab, 2scFv-IgG, IgG-2scFv, scFv4-Ig and zybody. For a review of bispecific formats, see Spiess, C., et al., Mol. Immunol. (2015). In another embodiment, the bispecific molecule comprises an antibody which is fused to another non-Ig format, for example a T-cell receptor binding domain; an immunoglobulin superfamily domain; an agnathan variable lymphocyte receptor; a fibronectin domain (e.g. an Adnectin.TM.); an antibody constant domain (e.g. a CH.sub.3 domain, e.g., a CH.sub.2 and/or CH.sub.3 of an Fcab.TM.) wherein the constant domain is not a functional CH.sub.1 domain; an scFv; an (scFv).sub.2; an sc-diabody; an scFab; a centyrin and an epitope binding domain derived from a scaffold selected from CTLA-4 (Evibody.TM.); a lipocalin domain; Protein A such as Z-domain of Protein A (e.g. an Affibody.TM. or SpA); an A-domain (e.g. an Avimer.TM. or Maxibodyn.TM.); a heat shock protein (such as and epitope binding domain derived from GroEI and GroES); a transferrin domain (e.g. a trans-body); ankyrin repeat protein (e.g. a DARPin.TM.); peptide aptamer; C-type lectin domain (e.g. Tetranectin.TM.); human .gamma.-crystallin or human ubiquitin (an affilin); a PDZ domain; scorpion toxin; and a kunitz type domain of a human protease inhibitor.
[0161] In one embodiment, the bispecific antibody is a mAb.sup.2. A mAb.sup.2 comprises a V.sub.H and V.sub.L domain from an intact antibody, fused to a modified constant region, which has been engineered to form an antigen-binding site, known as an "Fcab". The technology behind the Fcab/mAb.sup.2 format is described in more detail in WO2008/003103, and the description of the mAb.sup.2 format is incorporated herein by reference.
[0162] In one embodiment, a "bispecific antibody" does not include a FIT-Ig format. In one embodiment, a "bispecific antibody" does not include a mAb.sup.2 format. In one embodiment, a "bispecific antibody" does not include either a FIT-Ig format or a mAb.sup.2 format.
[0163] In another embodiment, the bispecific antibody is a "dual binding antibody". As used herein, the term "dual binding antibody" is a bispecific antibody wherein both antigen-binding domains are formed by a V.sub.H/V.sub.L pair, and includes FIT-Ig (see WO2015/103072, incorporated herein by reference), mAb-dAb, dock and lock, Fab-arm exchange, SEEDbody, Triomab, LUZ-Y, Fcab, .kappa..lamda.-body, orthogonal Fab, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CH3, Diabody-CH3, Triple body, Miniantibody, minibody, scFv-CH.sub.3 KIH, scFv-CH-CL-scFv, F(ab').sub.2-scFv, scFv-KIH, Fab-scFv-Fc, tetravalent HCab, ImmTAC, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, DT-IgG, DutaMab, IgG(H)-scFv, scFv-(H)IgG, IgG(L)-scFv, scFv-(L)IgG, IgG(L,H)-Fv, IgG(H)-V, V(H)--IgG, IgG(L)-V, V(L)-IgG, KIH IgG-scFab, 2scFv-IgG, IgG-2scFv and scFv4-Ig.
[0164] The term "hypervariable region", "CDR region" or "CDR" refers to the regions of an antibody variable domain which are hypervariable in sequence and/or form structurally defined loops. Generally, antigen binding sites of an antibody include six hypervariable regions: three in the V.sub.H (CDRH1, CDRH2, CDRH3), and three in the V.sub.L (CDRL1, CDRL2, CDRL3). These regions of the heavy and light chains of an antibody confer antigen-binding specificity to the antibody. CDRs may be defined according to the Kabat system (see Kabat, E. A. et al., 1991, "Sequences of Proteins of Immunological Interest", 5th edit., NIH Publication no. 91-3242, U.S. Department of Health and Human Services). Other systems may be used to define CDRs, which as the system devised by Chothia et al (see Chothia, C. & Lesk, A. M., 1987, "Canonical structures for the hypervariable regions of immunoglobulins", J. Mol. Biol., 196, 901-917) and the IMGT system (see Lefranc, M. P., 1997, "Unique database numbering system for immunogenetic analysis", Immunol. Today, 18, 50). An antibody typically contains 3 heavy chain CDRs and 3 light chain CDRs. The term CDR or CDRs is used here to indicate one or several of these regions. A person skilled in the art is able to readily compare the different systems of nomenclature and determine whether a particular sequence may be defined as a CDR.
[0165] A "human antibody" is an antibody that possesses an amino-acid sequence corresponding to that of an antibody produced by a human and/or has been made using any of the techniques for making human antibodies and specifically excludes a humanized antibody comprising non-human antigen-binding residues. The term "specifically binds to" refers to measurable and reproducible interactions such as binding between a target and an antibody, which is determinative of the presence of the target in the presence of a heterogeneous population of molecules including biological molecules. For example, an antibody that specifically binds to a target (which can be an epitope) is an antibody that binds this target with greater affinity, avidity, more readily, and/or with greater duration than it binds to other targets. In one embodiment, the extent of binding of an antibody to an unrelated target is less than about 10% of the binding of the antibody to the target as measured, e.g. by a radioimmunoassay (RIA).
[0166] An antibody or a fragment thereof that specifically binds to a hPD-L1 antigen may be cross-reactive with related antigens. Preferably, an antibody or a fragment thereof that specifically binds to a hPD-L1 antigen does not cross-react with other antigens (but may optionally cross-react with PD-L1 of a different species, e.g. rhesus, or murine). An antibody or a fragment thereof that specifically binds to a hPD-L1 antigen can be identified, for example, by immunoassays, BIAcore.TM., or other techniques known to those of skill in the art. An antibody or a fragment thereof binds specifically to a PD-L1 antigen when it binds to a hPD-L1 antigen with higher affinity than to any cross-reactive antigen as determined using experimental techniques, such as radioimmunoassays (RIA) and enzyme-linked immunosorbent assays (ELISAs). Typically, a specific or selective reaction will be at least twice background signal or noise and more typically more than 10 times (such as more than 15 times, more than 20 times, more than 50 times or more than 100 times) background. See, e.g. Paul, ed., 1989, Fundamental Immunology Second Edition, Raven Press, New York at pages 332-336 for a discussion regarding antibody specificity.
[0167] The term "aliphatic amino acid" means that the amino acid R groups are nonpolar and hydrophobic. Hydrophobicity increases with increasing number of C atoms in the hydrocarbon chain. Glycine, Alanine, Valine, Leucine and Isoleucine are aliphatic amino acids.
[0168] The term "aromatic amino acid" means that the amino acid R groups contain an aromatic ring system. Phenylalanine, Tyrosine and Tryptophan are aromatic amino acids.
[0169] The term "hydroxyl-containing amino acid" means that the amino acid R groups contain a hydroxyl group, and are hydrophilic. Serine, Cysteine, Threonine and Methionine are hydroxyl-containing amino acids.
[0170] The term "basic amino acid" means that the amino acid R groups are nitrogen containing and are basic at neutral pH. Histidine, Lysine and Arginine are basic amino acids.
[0171] The term "cyclic amino acid" means that the amino acid R groups have an aliphatic cyclic structure. Proline is the only cyclic aliphatic amino acid.
[0172] The term "acidic amino acid" means that the amino acid R groups are polar and are negatively charged at physiological pH. Aspartate and Glutamate are acidic amino acids.
[0173] The term "amide amino acid" means that the amino acid R groups contain an amide group. Asparagine and Glutamine are amide amino acids.
[0174] As used herein, "authorization number" or "marketing authorization number" refers to a number issued by a regulatory agency upon that agency determining that a particular medical product and/or composition may be marketed and/or offered for sale in the area under the agency's jurisdiction. As used herein "regulatory agency" refers to one of the agencies responsible for evaluating, e.g. the safety and efficacy of a medical product and/or composition and controlling the sales/marketing of such products and/or compositions in a given area. The Food and Drug Administration (FDA) in the US and the European Medicines Agency (EPA) in Europe are but two examples of such regulatory agencies. Other non-limiting examples can include SDA, MPA, MHPRA, IMA, ANMAT, Hong Kong Department of Health-Drug Office, CDSCO, Medsafe, and KFDA.
[0175] As used herein, the term "biomarker" refers to a gene that is differentially expressed in individuals having a disease of interest, for example, a gene that is differentially expressed in individuals having cancer. In one embodiment, PD-L1 is a biomarker whose expression in tumours may be indicative as to whether or not a patient would respond to a particular type of treatment, in particular, whether a patient would response to treatment targeting PD-L1, for example, immunotherapy using anti-PD-L1 antibodies. In one embodiment, PD-L1 is a biomarker whose expression in tumours may be indicative as to whether or not a patient would respond to a particular type of treatment, in particular, whether a patient would response to treatment targeting PD-1, for example, immunotherapy using anti-PD-1 antibodies. In another embodiment, PD-L1 may be free or membrane bound. In another embodiment, PD-L1 may be fixed or unfixed.
[0176] As used herein, a "buffer" refers to a chemical agent that is able to absorb a certain quantity of acid or base without undergoing a strong variation in pH.
[0177] As used herein, the term "carrier" refers to a diluent, adjuvant (e.g., Freund's adjuvant (complete and incomplete)), excipient, or vehicle with which the therapeutic is administered. Such pharmaceutical carriers can be sterile liquids, such as water and oils, including those of petroleum, animal, vegetable or synthetic origin, such as peanut oil, soybean oil, mineral oil, sesame oil and the like. Water is a preferred carrier when the pharmaceutical composition is administered intravenously. Saline solutions and aqueous dextrose and glycerol solutions can also be employed as liquid carriers, particularly for injectable solutions.
[0178] The term "chemotherapeutic agent" or "chemotherapy" refers to a therapeutic agent whose primary purpose is to destroy cancer cells, typically by interfering with the tumour cell's ability to grow or multiply. There are many different types of chemotherapeutic agents, with more than 50 approved chemotherapy drugs available. Chemotherapeutic drugs can be classified based on how they work. Alkylating drugs kill cancer cells by directly attacking DNA, the genetic material of the genes. Cyclophosphamide is an alkylating drug. Antimetabolites interfere/with the production of DNA and keep cells from growing and multiplying. An example of an antimetabolite is 5-fluorouracil (5-FU). Anti-tumour antibiotics are made from natural substances such as fungi in the soil. They interfere with important cell functions, including production of DNA and cell proteins. Doxorubicin and bleomycin belong to this group of chemotherapy drugs. Plant alkaloids prevent cells from dividing normally. Vinblastine and vincristine are plant alkaloids obtained from the periwinkle plant. Steroid hormones slow the growth of some cancers that depend on hormones. For example, tamoxifen is used to treat breast cancers that depend on the hormone estrogen for growth. DNA damage response (DDR) inhibitors, such as PARP inhibitors, block DNA repair mechanisms following single or double stranded breaks.
[0179] Examples of chemotherapeutic agents include Adriamycin, Doxorubicin, 5-Fluorouracil, Cytosine arabinoside (Ara-C), Cyclophosphamide, Thiotepa, Taxotere (docetaxel), Busulfan, Cytoxin, Taxol, Methotrexate, Cisplatin, Melphalan, Vinblastine, Bleomycin, Etoposide, Ifosfamide, Mitomycin C, Mitoxantrone, Vincreistine, Vinorelbine, Carboplatin, Teniposide, Daunomycin, Carminomycin, Aminopterin, Dactinomycin, Mitomycins, Esperamicins (see, U.S. Pat. No. 4,675,187), Melphalan, and other related nitrogen mustards. Suitable toxins and chemotherapeutic agents are described in Remington's Pharmaceutical Sciences, 19th Ed. (Mack Publishing Co. 1995), and in Goodman and Gilman's The Pharmacological Basis of Therapeutics, 7th Ed. (MacMillan Publishing Co. 1985). Another example of chemotherapeutic agents is the class of antibody-conjugated toxins, including, but not limited to pyrrolobenzodiazepiness, maytansanoids, calicheamicin, etc. Other suitable toxins and/or chemotherapeutic agents are known to those of skill in the art.
[0180] As used herein, the term "composition" is intended to encompass a product containing the specified ingredients (e.g. an antibody of the invention) in, optionally, the specified amounts, as well as any product which results, directly or indirectly, from combination of the specified ingredients in, optionally, the specified amounts.
[0181] As used herein the term "comprising" or "comprises" is used with reference to antibodies, fragments, uses, compositions, methods, and respective component(s) thereof, that are essential to the method or composition, yet open to the inclusion of unspecified elements, whether essential or not.
[0182] The term "consisting of" refers to antibodies, fragments, uses, compositions, methods, and respective components thereof as described herein, which are exclusive of any element not recited in that description of the embodiment.
[0183] As used herein the term "consisting essentially of" refers to those elements required for a given embodiment. The term permits the presence of elements that do not materially affect the basic and novel or functional characteristic(s) of that embodiment.
[0184] In the context of a polypeptide, the term "derivative" as used herein refers to a polypeptide that comprises an amino acid sequence of a hPD-L1 polypeptide, a fragment of a hPD-L1 polypeptide, or an antibody that specifically binds to a hPD-L1 polypeptide which has been altered by the introduction of amino acid residue substitutions, deletions or additions. The term "derivative" as used herein also refers to a hPD-L1 polypeptide, a fragment of a hPD-L1 polypeptide, or an antibody that specifically binds to a hPD-L1 polypeptide which has been chemically modified, e.g. by the covalent attachment of any type of molecule to the polypeptide. For example, but not by way of limitation, a hPD-L1 polypeptide, a fragment of a hPD-L1 polypeptide, or a hPD-L1 antibody may be chemically modified, e.g. by glycosylation, acetylation, pegylation, phosphorylation, amidation, derivatization by known protecting/blocking groups, proteolytic cleavage, linkage to a cellular ligand or other protein, etc. The derivatives are modified in a manner that is different from naturally occurring or starting peptide or polypeptides, either in the type or location of the molecules attached. Derivatives further include deletion of one or more chemical groups which are naturally present on the peptide or polypeptide. A derivative of a hPD-L1 polypeptide, a fragment of a hPD-L1 polypeptide, or a hPD-L1 antibody may be chemically modified by chemical modifications using techniques known to those of skill in the art, including, but not limited to specific chemical cleavage, acetylation, formulation, metabolic synthesis of tunicamycin, etc. Further, a derivative of a hPD-L1 polypeptide, a fragment of a hPD-L1 polypeptide, or a hPD-L1 antibody may contain one or more non-classical amino acids. A polypeptide derivative possesses a similar or identical function as a hPD-L1 polypeptide, a fragment of a hPD-L1 polypeptide, or a hPD-L1 antibody described herein.
[0185] The term "effector function" as used herein is meant to refer to one or more of antibody dependant cell mediated cytotoxic activity (ADCC), complement-dependant cytotoxic activity (CDC) mediated responses, Fc-mediated phagocytosis or antibody dependant cellular phagocytosis (ADCP) and antibody recycling via the FcRn receptor.
[0186] An "effective amount" refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired effect, including a therapeutic or prophylactic result. A "therapeutically effective amount" refers to the minimum concentration required to effect a measurable improvement or prevention of a particular disorder. A therapeutically effective amount herein may vary according to factors such as the disease state, age, sex, and weight of the patient, and the ability of the antibody to elicit a desired response in the individual. A therapeutically effective amount is also one in which toxic or detrimental effects of the antibody are outweighed by the therapeutically beneficial effects. A "prophylactically effective amount" refers to an amount effective, at the dosages and for periods of time necessary, to achieve the desired prophylactic result. In some embodiments, the effective amount of an antibody of the invention is from about 0.1 mg/kg (mg of antibody per kg weight of the subject) to about 100 mg/kg. In certain embodiments, an effective amount of an antibody provided therein is about 0.1 mg/kg, about 0.5 mg/kg, about 1 mg/kg, 3 mg/kg, 5 mg/kg, about 10 mg/kg, about 15 mg/kg, about 20 mg/kg, about 25 mg/kg, about 30 mg/kg, about 35 mg/kg, about 40 mg/kg, about 45 mg/kg, about 50 mg/kg, about 60 mg/kg, about 70 mg/kg, about 80 mg/kg about 90 mg/kg or about 100 mg/kg (or a range therein). In some embodiments, "effective amount" as used herein also refers to the amount of an antibody of the invention to achieve a specified result (e.g. inhibition of a hPD-L1 biological activity of a cell).
[0187] The term "epitope" as used herein refers to a localized region on the surface of an antigen, such as hPD-L1 polypeptide or hPD-L1 polypeptide fragment, that is capable of being bound to one or more antigen binding regions of an antibody, and that has antigenic or immunogenic activity in an animal, preferably a. mammal, and most preferably in a human, that is capable of eliciting an immune response. An epitope having immunogenic activity is a portion of a polypeptide that elicits an antibody response in an animal. An epitope having antigenic activity is a portion of a polypeptide to which an antibody specifically binds as determined by any method well known in the art, for example, by the immunoassays described herein. Antigenic epitopes need not necessarily be immunogenic. Epitopes usually consist of chemically active surface groupings of molecules such as amino acids or sugar side chains and have specific three dimensional structural characteristics as well as specific charge characteristics. A region of a polypeptide contributing to an epitope may be contiguous amino acids of the polypeptide or the epitope may come together from two or more non-contiguous regions of the polypeptide. The epitope may or may not be a three-dimensional surface feature of the antigen. In certain embodiments, a hPD-L1 epitope is a three-dimensional surface feature of a hPD-L1 polypeptide (e.g. in a trimeric form of a hPD-L1 polypeptide). In other embodiments, a hPD-L1 epitope is linear feature of a hPD-L1 polypeptide (e.g. in a trimeric form or monomeric form of the hPD-L1 polypeptide). Antibodies provided herein may specifically bind to an epitope of the monomeric (denatured) form of hPD-L1, an epitope of the trimeric (native) form of hPD-L1, or both the monomeric (denatured) form and the trimeric (native) form of hPD-L1. In specific embodiments, the antibodies provided herein specifically bind to an epitope of the trimeric form of hPD-L1 but do not specifically bind the monomeric form of hPD-L1.
[0188] The term "excipients" as used herein refers to inert substances which are commonly used as a diluent, vehicle, preservatives, binders, or stabilizing agent for drugs and includes, but not limited to, proteins (e.g. serum albumin, etc.), amino acids (e.g. aspartic acid, glutamic acid, lysine, arginine, glycine, histidine, etc.), fatty acids and phospholipids (e.g. alkyl sulfonates, caprylate, etc.), surfactants (e.g. SDS, polysorbate, nonionic surfactant, etc.), saccharides (e.g. sucrose, maltose, trehalose, etc.) and polyols (e.g. mannitol, sorbitol, etc.). See, also, Remington's Pharmaceutical Sciences (1990) Mack Publishing Co., Easton, Pa., which is hereby incorporated by reference in its entirety.
[0189] As used herein, the term "fixed" or "fixation" refers to a chemical process by which biological tissues are preserved from decay, to prevent autolysis or putrefaction. In general, fixation involves exposing the tissue to chemical compounds such as alcohols or aldehydes such as formaldehyde to terminate ongoing biochemical reactions. In some instances, fixation may also increase the mechanical strength or stability of the treated tissues. The term "unfixed" refers to a tissue that has not been subjected to a chemical process to prevent tissue decay. As used herein, the term "surface expressed" means that the protein is embedded in or spans a cell membrane or is associated with a protein that is embedded in or spans a cell membrane (i.e. a membrane associated protein). In one embodiment, a surface expressed protein includes one or more transmembrane domains. In another embodiment, the protein is associated with the exterior or interior surface of a cell membrane indirectly via association with another membrane spanning protein (i.e. the surface expressed protein is not spanning the cell membrane itself). In general, surface expressed proteins that are integrated into a cell membrane or expressed endogenously within a cell are more likely to fold in the correct conformation than recombinantly produced free forms of the same protein.
[0190] In the context of a peptide or polypeptide, the term "fragment" as used herein refers to a peptide or polypeptide that comprises less than the full length amino acid sequence. Such a fragment may arise, for example, from a truncation at the amino terminus, a truncation at the carboxy terminus, and/or an internal deletion of a residue(s) from the amino acid sequence. Fragments may, for example, result from alternative RNA splicing or from in vivo protease activity. In certain embodiments, PD-L1 fragments include polypeptides comprising an amino acid sequence of at least 5 contiguous amino acid residues, at least 10 contiguous amino acid residues, at least 15 contiguous amino acid residues, at least 20 contiguous amino acid residues, at least 25 contiguous amino acid residues, at least 40 contiguous amino acid residues, at least 50 contiguous amino acid residues, at least 60 contiguous amino residues, at least 70 contiguous amino acid residues, at least 80 contiguous amino acid residues, at least 90 contiguous amino acid residues, at least contiguous 100 amino acid residues, at least 125 contiguous amino acid residues, at least 150 contiguous amino acid residues, at least 175 contiguous amino acid residues, at least 200 contiguous amino acid residues, or at least 250 contiguous amino acid residues of the amino acid sequence of a hPD-L1 polypeptide or an antibody that specifically binds to a hPD-L1 polypeptide. In a specific embodiment, a fragment of a hPD-L1 polypeptide or an antibody that specifically binds to a hPD-L1 antigen retains at least 1, at least 2, or at least 3 functions of the polypeptide or antibody.
[0191] The term "free" refers to a polypeptide, for example, PD-L1 or fragments and variants thereof, that is combined with a buffer, wherein the polypeptide is not associated with a cell surface or cell membrane. As such, the term "free" can refer to a polypeptide that is capable of surface expression (i.e. includes one or more transmembrane domains or membrane association domains), but that is not, in its present state, expressed on the surface of a cell or bound to a protein that is expressed on the surface of a cell. A free polypeptide can also refer to a free recombinant or native or unbound polypeptide. In the context of phage display, a free antigen can be selected in solution (referred to herein as a "soluble selection") or adsorbed to a surface, for example, adsorbed to the surface of a 96-well plate (referred to herein as "biopanning selection").
[0192] The term "fusion protein" as used herein refers to a polypeptide that comprises an amino acid sequence of an antibody and an amino acid sequence of a heterologous polypeptide or protein (i.e. a polypeptide or protein not normally a part of the antibody (e.g. a non-anti-hPD-L1 antigen antibody)). The term "fusion" when used in relation to hPD-L1 or to an anti-hPD-L1 antibody refers to the joining of a peptide or polypeptide, or fragment, variant and/or derivative thereof, with a heterologous peptide or polypeptide. Preferably, the fusion protein retains the biological activity of the hPD-L1 or anti-hPD-L1 antibody. In certain embodiments, the fusion protein comprises a hPD-L1 antibody VH domain, VL domain, VH CDR (one, two or three VH CDRs), and/or VL CDR (one, two or three VL CDRs), wherein the fusion protein specifically binds to a hPD-L1 epitope.
[0193] The term "heavy chain" when used with reference to an antibody refers to five distinct types, called alpha (.alpha.), delta (.delta.), epsilon (.epsilon.), gamma (.gamma.) and mu (.mu.), based on the amino acid sequence of the heavy chain constant domain. These distinct types of heavy chains are well known and give rise to five classes of antibodies, IgA, IgD, IgE, IgG and IgM, respectively, including four subclasses of IgG, namely IgG1, IgG2, IgG3 and IgG4. Preferably the heavy chain is a human heavy chain. In the human population, multiple heavy chain constant region alleles, of each immunoglobulin or immunoglobulin subclass, exist. The nucleotide and amino acid sequences of these allelic variants are accessible on publicly available databases such as IMGT, ENSEMBL Swiss-Prot and Uniprot. Allelic variants may also be identified in various genome sequencing projects. In one embodiment, the antibodies and antibody fragments disclosed herein comprise a heavy chain encoded by a IgG1 constant region allele, which includes, but is not limited to, human IGHG1*01 (Seq ID Nos:340, 341 & 537), IGHG1*02 (Seq ID Nos:340, 341 &537), IGHG1*03 (Seq ID Nos:523 & 524), IGHG1*04 (Seq ID Nos:525 & 526) and IGHG1*05 (Seq ID Nos:340, 341 & 537). In one embodiment, the antibodies and antibody fragments disclosed herein comprise a protein encoded by a IgG2 constant region allele, which includes, but is not limited to, human IGHG2*01 (Seq ID Nos:527 & 528), IGHG2*02 (Seq ID Nos:529 & 530), IGHG2*03 (Seq ID Nos:527 & 528), IGHG2*04 (Seq ID Nos:531 & 532), IGHG2*05 (Seq ID Nos:527 & 528) and IGHG2*06 (Seq ID Nos:533 & 534). In one embodiment, the antibodies or antibody fragments disclosed herein comprise a protein encoded by a IgG3 constant region allele, which includes but is not limited to human IGHG3*01, IGHG3*02, IGHG3*03, IGHG3*04, IGHG3*05, IGHG3*06, IGHG3*07, IGHG3*08, IGHG3*09, IGHG3*10, IGHG3*11, IGHG3*12, IGHG3*13, IGHG3*14, IGHG3*15, IGHG3*16, IGHG3*17, IGHG3*18 and IGHG3*19. In one embodiment, the antibodies or antibody fragments disclosed herein comprise a protein encoded by a IgG4 constant region allele, which includes but is not limited to human IGHG4*01 (Seq ID Nos:192 & 193), IGHG4*02 (Seq ID Nos:194 & 195), IGHG4*03 (Seq ID Nos:196 & 197) and IGHG4*04 (Seq ID Nos:192 & 193). In another example, the heavy chain is a disabled IgG isotype, e.g. a disabled IgG4. In certain embodiments, the antibodies of the invention comprise a human gamma 4 constant region. In another embodiment, the heavy chain constant region does not bind Fc-.gamma. receptors, and e.g. comprises a Leu235Glu mutation. In another embodiment, the heavy chain constant region comprises a Ser228Pro mutation to increase stability. In another embodiment, the heavy chain constant region is IgG4-PE (SEQ ID No:199. In another embodiment, the antibodies and antibody fragments disclosed herein comprise a heavy chain constant region encoded by a murine IgG1 constant region allele, which includes but is not limited to mouse IGHG1*01 or IGHG1*02. In one embodiment, the antibodies and antibody fragments disclosed herein comprise a heavy chain constant region encoded by a murine IgG2 constant region allele, which includes, but is not limited to, mouse IGHG2A*01, IGHG2A*02, IGHG2B*01, IGHG2B*02, IGHG2C*01, IGHG2C*02 or IGHG2C*03. In one embodiment, the antibodies or antibody fragments disclosed herein comprise a protein encoded by a murine IgG3 constant region allele, which includes but is not limited to mouse IGHG3*01.
[0194] The term "host" as used herein refers to an animal, preferably a mammal, and most preferably a human.
[0195] The term "host cell" as used herein refers to the particular subject cell transfected with a nucleic acid molecule and the progeny or potential progeny of such a cell. Progeny of such a cell may not be identical to the parent cell transfected with the nucleic acid molecule due to mutations or environmental influences that may occur in succeeding generations or integration of the nucleic acid molecule into the host cell genome.
[0196] The term "an IL-2 cytokine" as used herein refers to a cytokine-like molecule which has a similar activity to a wild-type IL-2. It may have activity at the high (.alpha..beta..gamma.) affinity IL-2 receptor and/or the intermediate affinity (.alpha..beta.) IL-2 receptor. The cytokine may be a variant IL-2 cytokine having one or more amino acid deletions, substitutions or additions. Variant cytokines are described in more detail hereinbelow.
[0197] The term "immunomodulatory agent" and variations thereof including, but not limited to, immunomodulatory agents, as used herein refer to an agent that modulates a host's immune system. In certain embodiments, an immunomodulatory agent is an immunosuppressant agent. In certain other embodiments, an immunomodulatory agent is an immunostimulatory agent. In accordance with the invention, an immunomodulatory agent used in the combination therapies of the invention does not include an anti-hPD-L1 antibody or antigen-binding fragment. Immunomodulatory agents include, but are not limited to, small molecules, peptides, polypeptides, proteins, fusion proteins, antibodies, inorganic molecules, mimetic agents, and organic molecules.
[0198] The term "in combination" in the context of the administration of other therapies refers to the use of more than one therapy. The use of the term "in combination" does not restrict the order in which therapies are administered to a subject with a disease. A first therapy can be administered before (e.g. 1 minute, 45 minutes, 30 minutes, 45 minutes, 1 hour, 2 hours, 4 hours, 6 hours, 12 hours, 24 hours, 48 hours, 72 hours, 96 hours, 1 week, 2 weeks, 3 weeks, 4 weeks, 5 weeks, 6 weeks, 8 weeks, or 12 weeks), concurrently, or after (e.g. 1 minute, 45 minutes, 30 minutes, 45 minutes, 1 hour, 2 hours, 4 hours, 6 hours, 12 hours, 24 hours, 48 hours, 72 hours, 96 hours, 1 week, 2 weeks, 3 weeks, 4 weeks, 5 weeks, 6 weeks, 8 weeks, or 12 weeks) the administration of a second therapy to a subject which had, has, or is susceptible to a hPD-L1-mediated disease. Any additional therapy can be administered in any order with the other additional therapies. In certain embodiments, the antibodies of the invention can be administered in combination with one or more therapies (e.g. therapies that are not the antibodies of the invention that are currently administered to prevent, treat, manage, and/or ameliorate a hPD-L1-mediated disease. Non-limiting examples of therapies that can be administered in combination with an antibody of the invention include analgesic agents, anaesthetic agents, antibiotics, or immunomodulatory agents or any other agent listed in the U.S. Pharmacopoeia and/or Physician's Desk Reference.
[0199] The term "immunocytokine", as used herein refers to an antibody format which is fused to a cytokine molecule. The antibody format may be any of those described herein, and the cytokine may be fused directly, or by means of a linker or chemical conjugation to either the N- or C-terminus of the heavy or the light chain of the antibody format.
[0200] As used herein, "injection device" refers to a device that is designed for carrying out injections, an injection including the steps of temporarily fluidically coupling the injection device to a person's tissue, typically the subcutaneous tissue. An injection further includes administering an amount of liquid drug into the tissue and decoupling or removing the injection device from the tissue. In some embodiments, an injection device can be an intravenous device or IV device, which is a type of injection device used when the target tissue is the blood within the circulatory system, e.g. the blood in a vein. A common, but non-limiting example of an injection device is a needle and syringe.
[0201] As used herein, "instructions" refers to a display of written, printed or graphic matter on the immediate container of an article, for example the written material displayed on a vial containing a pharmaceutically active agent, or details on the composition and use of a product of interest included in a kit containing a composition of interest. Instructions set forth the method of the treatment as contemplated to be administered or performed.
[0202] An "isolated" or "purified" antibody or protein is one that has been identified, separated and/or recovered from a component of its production environment (e.g. natural or recombinant). For example, the antibody or protein is substantially free of cellular material or other contaminating proteins from the cell or tissue source from which the antibody is derived, or substantially free of chemical precursors or other chemicals when chemically synthesized. The language "substantially free of cellular material" includes preparations of an antibody in which the antibody is separated from cellular components of the cells from which it is isolated or recombinantly produced. Thus, an antibody that is substantially free of cellular material includes preparations of antibody having less than about 30%, 20%, 10%, or 5% (by dry weight) of heterologous protein (also referred to herein as a "contaminating protein"). When the antibody is recombinantly produced, it is also preferably substantially free of culture medium, i.e. culture medium represents less than about 20%, 10%, or 5% of the volume of the protein preparation. When the antibody is produced by chemical synthesis, it is preferably substantially free of chemical precursors or other chemicals, i.e., it is separated from chemical precursors or other chemicals which are involved in the synthesis of the protein. Accordingly, such preparations of the antibody have less than about 30%, 20%, 10%, 5% (by dry weight) of chemical precursors or compounds other than the antibody of interest. In a preferred embodiment, antibodies of the invention are isolated or purified.
[0203] The terms "Kabat numbering," and like terms are recognized in the art and refer to a system of numbering amino acid residues which are more variable (i.e. hypervariable) than other amino acid residues in the heavy chain variable regions of an antibody, or an antigen binding portion thereof (Kabat et al., (1971) Ann. NY Acad. Sci., 190:382-391 and, Kabat et al. (1991) Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242). For the heavy chain variable region, the hypervariable region typically ranges from amino acid positions 31 to 35 for CDR1, amino acid positions 50 to 65 for CDR2, and amino acid positions 95 to 102 for CDR3.
[0204] "Label" or "labelled" as used herein refers to the addition of a detectable moiety to a polypeptide, for example, a radiolabel, fluorescent label, enzymatic label, chemiluminescent label or a biotinyl group or gold. Radioisotopes or radionuclides may include .sup.3H, .sup.14C, .sup.15N, .sup.35S, .sup.90Y, .sup.99Tc, .sup.115In, .sup.125I, .sup.131I, fluorescent labels may include rhodamine, lanthanide phosphors or FITC and enzymatic labels may include horseradish peroxidase, .beta.-galactosidase, luciferase, alkaline phosphatase. Additional labels include, by way of illustration and not limitation: enzymes, such as glucose-6-phosphate dehydrogenase ("G6PDH"), alpha-D-galactosidase, glucose oxydase, glucose amylase, carbonic anhydrase, acetylcholinesterase, lysozyme, malate dehydrogenase and peroxidase; dyes (e.g. cyanine dyes, e.g. Cy5.TM., Cy5.5.TM.. or Cy7.TM.); additional fluorescent labels or fluorescers include, such as fluorescein and its derivatives, fluorochrome, GFP (GFP for "Green Fluorescent Protein"), other fluorescent proteins (e.g. mCherry, mTomato), dansyl, umbelliferone, phycoerythrin, phycocyanin, allophycocyanin, o-phthaldehyde, and fluorescamine; fluorophores such as lanthanide cryptates and chelates e.g. Europium etc (Perkin Elmer and Cisbio Assays); chemoluminescent labels or chemiluminescers, such as isoluminol, luminol and the dioxetanes; sensitisers; coenzymes; enzyme substrates; particles, such as latex or carbon particles; metal sol; crystallite; liposomes; cells, etc., which may be further labelled with a dye, catalyst or other detectable group; molecules such as biotin, digoxygenin or 5-bromodeoxyuridine; toxin moieties, such as for example a toxin moiety selected from a group of Pseudomonas exotoxin (PE or a cytotoxic fragment or mutant thereof), Diptheria toxin or a cytotoxic fragment or mutant thereof, a botulinum toxin A, B, C, D, E or F, ricin or a cytotoxic fragment thereof e.g. ricin A, abrin or a cytotoxic fragment thereof, saporin or a cytotoxic fragment thereof, pokeweed antiviral toxin or a cytotoxic fragment thereof and bryodin 1 or a cytotoxic fragment thereof.
[0205] The term "light chain" when used in reference to an antibody refers to the immunoglobulin light chains, of which there are two types in mammals, lambda (.lamda.) and kappa (.kappa.). Preferably, the light chain is a human light chain. Preferably the light chain constant region is a human constant region. In the human population, multiple light chain constant region alleles exist. The nucleotide and amino acid sequences of these allelic variants are accessible on publicly available databases such as IMGT, ENSEMBL, Swiss-Prot and Uniprot. In one embodiment, the antibodies or antibody fragments disclosed herein comprise a protein encoded by a human K constant region allele, which includes, but is not limited to, IGKC*01 (Seq ID Nos:206 & 207), IGKC*02 (Seq ID Nos:208 & 209), IGKC*03 (Seq ID Nos:210 & 211), IGKC*04 (Seq ID Nos:212 & 213) and IGKC*05 (Seq ID Nos:214 & 215). In one embodiment, the antibodies or antibody fragments disclosed herein comprise a protein encoded by a human A constant region allele, which includes but is not limited to IGLC1*01 (Seq ID Nos:216 & 217), IGLC1*02 (Seq ID Nos:218, 219 & 220), IGLC2*01 (Seq ID Nos:221, 222 & 538), IGLC2*02 (Seq ID Nos:224 & 225), IGLC2*03 (Seq ID Nos:224 & 225), IGLC3*01 (Seq ID Nos:226 & 227), IGLC3*02 (Seq ID Nos:228 & 229), IGLC3*03 (Seq ID Nos:230 & 231), IGLC3*04 (Seq ID Nos:232 & 233), IGLC6*01 (Seq ID Nos:234 & 235), IGLC7*01 (Seq ID Nos:236 & 237), IGLC7*02 (Seq ID Nos:236 & 237), IGLC7*03 (Seq ID Nos:535 & 536). In another embodiment, the antibodies and antibody fragments disclosed herein comprise a light chain constant region encoded by a mouse .kappa. constant region allele, which includes, but is not limited to, IGKC*01, IGKC*03 or IGKC*03. In another embodiment, the antibodies and antibody fragments disclosed herein comprise a light chain constant region encoded by a mouse A constant region allele, which includes, but is not limited to, IGLC1*01, IGLC2*01 or IGLC3*01.
[0206] "Percent (%) amino acid sequence identity" and "homology" with respect to a peptide, polypeptide or antibody sequence are defined as the percentage of amino acid residues in a candidate sequence that are identical with the amino acid residues in the specific peptide or polypeptide sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN or MEG ALIGN.TM. (DNASTAR) software. In one embodiment, the % homology is about 70%. In one embodiment, the % homology is about 75%. In one embodiment, the % homology is about 80%. In one embodiment, the % homology is about 85%. In one embodiment, the % homology is about 90%. In one embodiment, the % homology is about 92%. In one embodiment, the % homology is about 95%. In one embodiment, the % homology is about 97%. In one embodiment, the % homology is about 98%. In one embodiment, the % homology is about 99%. In one embodiment, the % homology is 100%.
[0207] The term "naturally occurring" or "native" when used in connection with biological materials such as nucleic acid molecules, polypeptides, host cells, and the like, refers to those which are found in nature and not manipulated by a human being.
[0208] As used herein, "packaging" refers to how the components are organized and/or restrained into a unit fit for distribution and/or use. Packaging can include, e.g. boxes, bags, syringes, ampoules, vials, tubes, clamshell packaging, barriers and/or containers to maintain sterility, labelling, etc.
[0209] The term "pharmaceutically acceptable" as used herein means being approved by a regulatory agency of the Federal or a state government, or listed in the U.S. Pharmacopeia, European Pharmacopeia or other generally recognized Pharmacopeia for use in animals, and more particularly in humans.
[0210] As used herein, the term "polynucleotide," "nucleotide," nucleic acid" "nucleic acid molecule" and other similar terms are used interchangeable and include DNA, RNA, mRNA and the like.
[0211] As used herein, the terms "prevent", "preventing", and "prevention" refer to the total or partial inhibition of the development, recurrence, onset or spread of a hPD-L1-mediated disease and/or symptom related thereto, resulting from the administration of a therapy or combination of therapies provided herein (e.g. a combination of prophylactic or therapeutic agents, such as an antibody of the invention).
[0212] The term "soluble" refers to a polypeptide, such as PD-L1 and variants or fragments thereof, that is lacking one or more transmembrane or cytoplasmic domains found in the native or membrane-associated form. In one embodiment, the "soluble" form of PD-L1 lacks both the transmembrane domain and the cytoplasmic domain.
[0213] The term "subject" or "patient" refers to any animal, including, but not limited to, mammals. As used herein, the term "mammal" refers to any vertebrate animal that suckle their young and either give birth to living young (eutharian or placental mammals) or are egg-laying (metatharian or nonplacental mammals). Examples of mammalian species include, but are not limited to, humans and other primates, including non-human primates such as chimpanzees and other apes and monkey species; farm animals such as cattle, sheep, pigs, goats and horses; domestic mammals such as dogs and cats; laboratory animals including rodents such as mice, rats (including cotton rats) and guinea pigs; birds, including domestic, wild and game birds such as chickens, turkeys and other gallinaceous birds, ducks, geese, and the like.
[0214] As used herein "substantially all" refers to refers to at least about 60%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 98%, at least about 99%, or about 100%.
[0215] The term "substantially free of surfactant" as used herein refers to a formulation of an antibody that specifically binds to a hPD-L1 antigen, said formulation containing less than 0.0005%, less than 0.0003%, or less than 0.0001% of surfactants and/or less than 0.0005%, less than 0.0003%, or less than 0.0001% of surfactants.
[0216] The term "substantially free of salt" as used herein refers to a formulation of an antibody that specifically binds to a hPD-L1 antigen, said formulation containing less than 0.0005%, less than 0.0003%, or less than 0.0001% of inorganic salts.
[0217] The term "surfactant" as used herein refers to organic substances having amphipathic structures; namely, they are composed of groups of opposing solubility tendencies, typically an oil-soluble hydrocarbon chain and a water-soluble ionic group. Surfactants can be classified, depending on the charge of the surface-active moiety, into anionic, cationic, and non-ionic surfactants. Surfactants are often used as wetting, emulsifying, solubilizing, and dispersing agents for various pharmaceutical compositions and preparations of biological materials.
[0218] As used herein, the term "tag" refers to any type of moiety that is attached to, e.g. a polypeptide and/or a polynucleotide that encodes a hPD-L1 or hPD-L1 antibody or antigen binding fragment thereof. For example, a polynucleotide that encodes a hPD-L1, hPD-L1 antibody or antigen binding fragment thereof can contain one or more additional tag-encoding nucleotide sequences that encode e.g. a detectable moiety or a moiety that aids in affinity purification. When translated, the tag and the antibody can be in the form of a fusion protein. The term "detectable" or "detection" with reference to a tag refers to any tag that is capable of being visualized or wherein the presence of the tag is otherwise able to be determined and/or measured (e.g. by quantitation). A non-limiting example of a detectable tag is a fluorescent tag.
[0219] As used herein, the term "therapeutic agent" refers to any agent that can be used in the treatment, management or amelioration of a hPD-L1-mediated disease and/or a symptom related thereto. In certain embodiments, the term "therapeutic agent" refers to an antibody of the invention. In certain other embodiments, the term "therapeutic agent" refers to an agent other than an antibody of the invention. Preferably, a therapeutic agent is an agent which is known to be useful for, or has been or is currently being used for the treatment, management or amelioration of a hPD-L1-mediated disease or one or more symptoms related thereto. In specific embodiments, the therapeutic agent is a fully human anti-hPD-L1 antibody, such as a fully human anti-hPD-L1 monoclonal antibody.
[0220] As used herein, the term "therapy" refers to any protocol, method and/or agent that can be used in the prevention, management, treatment and/or amelioration of a hPD-L1-mediated disease (e.g. cancer). In certain embodiments, the terms "therapies" and "therapy" refer to a biological therapy, supportive therapy, and/or other therapies useful in the prevention, management, treatment and/or amelioration of a hPD-L1-mediated disease known to one of skill in the art such as medical personnel.
[0221] The terms "treat", "treatment" and "treating" refer to the reduction or amelioration of the progression, severity, and/or duration of a hPD-L1-mediated disease (e.g. cancer) resulting from the administration of one or more therapies (including, but not limited to, the administration of one or more prophylactic or therapeutic agents, such as an antibody of the invention). In specific embodiments, such terms refer to the reduction or inhibition of the binding of hPD-L1 to PD-1, the reduction or inhibition of the binding of hPD-L1 to CD80, and/or the inhibition or reduction of one or more symptoms associated with a hPD-L1-mediated disease, such as cancer. In specific embodiments, such terms refer to the reduction or inhibition of the binding of hPD-L1 to PD-1 and/or CD80, and/or the inhibition or reduction of one or more symptoms associated with a hPD-L1-mediated disease, such as cancer. In an example, the cell is a human cell. In specific embodiments, a prophylactic agent is a fully human anti-hPD-L1 antibody, such as a fully human anti-hPD-L1 monoclonal antibody.
[0222] The term "variable region" or "variable domain" refers to a portion of the light and heavy chains, typically about the amino-terminal 120 to 130 amino acids in the heavy chain and about 100 to 110 amino acids in the light chain, which differ extensively in sequence among antibodies and are used in the binding and specificity of each particular antibody for its particular antigen. The variability in sequence is concentrated in those regions called complimentarily determining regions (CDRs) while the more highly conserved regions in the variable domain are called framework regions (FR). The CDRs of the PD-L1 and heavy chains are primarily responsible for the interaction of the antibody with antigen. Numbering of amino acid positions used herein is according to the EU Index, as in Kabat et al. (1991) Sequences of proteins of immunological interest. (U.S. Department of Health and Human Services, Washington, D.C.) 5.sup.th ed. ("Kabat et al."). In preferred embodiments, the variable region is a human variable region.
[0223] Definitions of common terms in cell biology and molecular biology can be found in "The Merck Manual of Diagnosis and Therapy", 19th Edition, published by Merck Research Laboratories, 2006 (ISBN 0-911910-19-0); Robert S. Porter et al. (eds.), The Encyclopedia of Molecular Biology, published by Blackwell Science Ltd., 1994 (ISBN 0-632-02182-9); Benjamin Lewin, Genes X, published by Jones & Bartlett Publishing, 2009 (ISBN-10: 0763766321); Kendrew et al. (Eds.), Molecular Biology and Biotechnology: a Comprehensive Desk Reference, published by VCH Publishers, Inc., 1995 (ISBN 1-56081-569-8) and Current Protocols in Protein Sciences 2009, Wiley Intersciences, Coligan et al., eds.
[0224] Unless otherwise stated, the present invention was performed using standard procedures, as described, for example in Sambrook et al., Molecular Cloning: A Laboratory Manual (4 ed.), Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., USA (2012); Davis et al., Basic Methods in Molecular Biology, Elsevier Science Publishing, Inc., New York, USA (1995); or Methods in Enzymology: Guide to Molecular Cloning Techniques Vol. 152, S. L Berger and A. R. Kimmel Eds., Academic Press Inc., San Diego, USA (1987); Current Protocols in Protein Science (CPPS) (John E. Coligan, et al., ed., John Wiley and Sons, Inc.), Current Protocols in Cell Biology (CPCB) (Juan S. Bonifacino et al. ed., John Wiley and Sons, Inc.), and Culture of Animal Cells: A Manual of Basic Technique by R. Ian Freshney, Publisher: Wiley-Liss; 5th edition (2005), Animal Cell Culture Methods (Methods in Cell Biology, Vol. 57, Jennie P. Mather and David Barnes editors, Academic Press, 1st edition, 1998) which are all incorporated by reference herein in their entireties.
[0225] Other terms are defined herein within the description of the various aspects of the invention.
2. PD-L1 Antibodies
[0226] Many tumour cells express surface molecules that are specific to cancer that can serve as diagnostic and/or therapeutic antibody targets. Examples of cell surface proteins expressed by tumour molecules that can be useful as biomarkers include, for example, members of the B7 family of proteins, major histocompatibility complex molecules (MHC), cytokine and growth factor receptors such as the receptor for epidermal growth factor (EGFR). The B7 family is a group of proteins that are members of the immunoglobulin (Ig) superfamily of cell-surface proteins that bind to receptors on lymphocytes to regulate immune responses. The family includes transmembrane or glycosylphosphatidylinositol (GPI)-linked proteins characterized by extracellular Ig-like domains (IgV and IgC domains related to the variable and constant domains of immunoglobulins). All members have short cytoplasmic domains. There are seven known members of the B7 family: B7-1, B7-2, PD-L1 (37-H1), PD-L2, B7-H2, B7-H3, and B7-H4.
[0227] The complete amino acid sequence for PD-L1 can be found in NCBI Reference Sequence: NP_054862.1 (SEQ ID No:1), which refers to many journal articles, including, for example, Dong, H., et al. (1999), "PD-L1, a third member of the B7 family, co-stimulates T-cell proliferation and interleukin-10 secretion," Nat. Med. 5 (12), 1365-1369, the disclosure of which is hereby incorporated by reference herein in its entirety. The amino acid sequence of PD-L1 includes a 30 amino acid long cytoplasmic domain that is unique to PD-L1, which shows little homology to other molecules, including other B7 family members.
[0228] In one embodiment, the antibody is a polyclonal antibody. Methods for generating polyclonal antibodies are known, and include, for example, inoculating a suitable mammal with an antigen to induce the immune system of the animal to produce immunoglobulins (IgGs) that specifically bind the injected antigen. Examples of suitable mammals include, for example, mouse, guinea pig, hamster, rat, rabbit sheep or goat. The polyclonal IgG is then typically purified from the mammal's serum. In one embodiment, the antibody is a polyclonal antibody that binds to a surface expressed protein. In another embodiment, the antibody is a polyclonal antibody that specifically binds to a member of the B7 family of proteins. In a more specific embodiment, the antibody is a polyclonal antibody that specifically binds PD-L1. In another embodiment, the antibody is a polyclonal antibody that specifically binds surface expressed PD-L1. In a more particular embodiment, the polyclonal antibody or antigen binding fragment thereof specifically binds human PD-L1. In another embodiment, the antibody is a polyclonal antibody that specifically binds soluble PD-L1. The term "soluble" also refers to a protein, such as PD-L1 that is lacking one or more transmembrane domain or cytoplasmic domains. In one embodiment, the "soluble" form of PD-L1 lacks both the transmembrane domain and the cytoplasmic domain. In one embodiment, the antibody is a polyclonal antibody that binds "free" PD-L1 (i.e. PD-L1 that is not associated with a cell membrane or surface, either directly or indirectly).
[0229] In another embodiment, the antibody can be a monoclonal antibody. Methods of making monoclonal antibodies are known and include, for example, fusing myeloma cells with the cells from an animal that was immunized with the desired antigen. In other embodiments, the monoclonal antibodies may be generated using recombinant DNA technology. In one embodiment, the antibody is a monoclonal antibody that specifically binds a surface expressed protein. In one embodiment, the antibody is a fully human monoclonal antibody. In another embodiment, the antibody is a monoclonal antibody that specifically binds to a member of the B7 family of proteins. In a more specific embodiment, the antibody is a monoclonal antibody that specifically binds PD-L1. In another embodiment, the antibody is a monoclonal antibody that specifically binds surface expressed PD-L1. In a more particular embodiment, the monoclonal antibody or antigen binding fragment thereof specifically binds human PD-L1. In another embodiment, the antibody is a monoclonal antibody that specifically binds soluble PD-L1. In one embodiment, the antibody is a monoclonal antibody that specifically binds soluble PD-L1 that is lacking one or more transmembrane domain or cytoplasmic domains. In one embodiment, the antibody is a monoclonal antibody that specifically binds soluble PD-L1 that is lacking both the transmembrane domain and the cytoplasmic domain. In one embodiment, the antibody is a monoclonal antibody that binds "free" PD-L1 (i.e. PD-L1 that is not associated with a cell membrane or surface, either directly or indirectly).
[0230] In an example the binding site(s) of the antibody or fragment are selected from a plurality (e.g. library) of binding sites. For example, the plurality of binding sites comprises or consists of a plurality of 4-chain antibodies or fragments thereof, e.g. dAbs, Fabs or scFvs. Suitable methods for producing pluralities of binding sites for screening include phage display (producing a phage display library of antibody binding sites), ribosome display (producing a ribosome display library of antibody binding sites), yeast display (producing a yeast display library of antibody binding sites), or immunisation of a non-human vertebrate (e.g. a rodent, e.g. a mouse or rat, e.g. a Velocimouse.TM., Kymouse.TM., Xenomouse.TM., Aliva Mouse.TM., HuMab Mouse.TM., Omnimouse.TM., Omnirat.TM. or MeMo Mouse.TM.) with hPD-L1 or a hPD-L1 epitope and isolation of a repertoire of antibody-producing cells (e.g. a B-cell, plasma cell or plasmablast repertoire) and/or a repertoire of isolated antibodies, fragments or binding sites.
[0231] PD-L1 binding ability, specificity and affinity (Kd, K.sub.off and/or K.sub.on) can be determined by any routine method in the art, e.g. by surface plasmon resonance (SPR). The term "Kd" or "K.sub.D", as used herein, is intended to refer to the equilibrium dissociation constant of a particular antibody-antigen interaction. Such binding measurements can be made using a variety of binding assays known in the art, e.g. using surface plasmon resonance (SPR), such as by Biacore.TM. or using the ProteOn XPR36.TM. (Bio-Rad.RTM.), using KinExA.RTM. (Sapidyne Instruments, Inc), or using ForteBio Octet (Pall ForteBio Corp.).
[0232] In one embodiment, the surface plasmon resonance (SPR) is carried out at 25.degree. C. In another embodiment, the SPR is carried out at 37.degree. C.
[0233] In one embodiment, the SPR is carried out at physiological pH, such as about pH7 or at pH7.6 (e.g. using Hepes buffered saline at pH 7.6 (also referred to as HBS-EP)).
[0234] In one embodiment, the SPR is carried out at a physiological salt level, e.g. 150 mM NaCl.
[0235] In one embodiment, the SPR is carried out at a detergent level of no greater than 0.05% by volume, e.g. in the presence of P20 (polysorbate 20; e.g. Tween 20.TM.) at 0.05% and EDTA at 3 mM.
[0236] In one example, the SPR is carried out at 25.degree. C. or 37.degree. C. in a buffer at pH 7.6, 150 mM NaCl, 0.05% detergent (e.g. P20) and 3 mM EDTA. The buffer can contain 10 mM Hepes. In one example, the SPR is carried out at 25.degree. C. or 37.degree. C. in HBS-EP. HBS-EP is available from Teknova Inc. (California; catalogue number H8022).
[0237] In an example, the affinity of the antibody or fragment is determined using SPR by: [0238] 1. Coupling anti-mouse (or other relevant human, rat or non-human vertebrate antibody constant region species-matched) IgG (e.g. Biacore.TM. BR-1008-38) to a biosensor chip (e.g. GLM chip) such as by primary amine coupling; [0239] 2. Exposing the anti-mouse IgG (or other matched species antibody) to a test IgG antibody to capture test antibody on the chip; [0240] 3. Passing the test antigen over the chip's capture surface at 1024 nM, 256 nM, 64 nM, 16 nM, 4 nM with a 0 nM (i.e. buffer alone); and [0241] 4. And determining the affinity of binding of test antibody to test antigen using surface plasmon resonance, e.g. under an SPR condition discussed above (e.g. at 25.degree. C. in physiological buffer). SPR can be carried out using any standard SPR apparatus, such as by Biacore.TM. or using the ProteOn XPR36.TM. (Bio-Rad.RTM.).
[0242] Regeneration of the capture surface can be carried out with 10 mM glycine at pH 1.7. This removes the captured antibody and allows the surface to be used for another interaction. The binding data can be fitted to 1:1 model inherent using standard techniques, e.g. using a model inherent to the ProteOn XPR36.TM. analysis software.
[0243] The present inventors have identified a number of antibodies having specificity for hPD-L1, which have a number of potential utilities and benefits over existing antibodies. For example, the antibodies described herein may have one or more of the following properties: [0244] a. Specificity for blocking only one of the ligands of PD-L1 (e.g. blocks CD80/PD-L1 interaction, but not PD-1/PD-L1 interaction) [0245] b. Immunogenicity/lack of side effects [0246] c. Solubility [0247] d. Stability [0248] e. Ease of formulation [0249] f. Frequency of dosing and/or route of administration, for example due to improved half-life over existing anti-PDL1 antibodies [0250] g. Manufacturability (e.g. expression, ease of purification, isoforms)
[0251] 1D05 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:33, comprising the CDRH1 amino acid sequence of Seq ID No:27 (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:34. 1D05 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:43, comprising the CDRL1 amino acid sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), the CDRL2 amino acid sequence of Seq ID No:38 (IMGT) or Seq ID No:41 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:44. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:35 (heavy chain nucleic acid sequence Seq ID No:36). A full length light chain amino acid sequence is Seq ID No:45 (light chain nucleic acid sequence Seq ID No:46).
[0252] 84G09 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:13, comprising the CDRH1 amino acid sequence of Seq ID No:7 (IMGT) or Seq ID No:10 (Kabat), the CDRH2 amino acid sequence of Seq ID No:8 (IMGT) or Seq ID No:11 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:9 (IMGT) or Seq ID No:12 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:14. 84G09 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:23, comprising the CDRL1 amino acid sequence of Seq ID No:17 (IMGT) or Seq ID No:20 (Kabat), the CDRL2 amino acid sequence of Seq ID No:18 (IMGT) or Seq ID No:21 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:19 (IMGT) or Seq ID No:22 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:24. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:15 (heavy chain nucleic acid sequence Seq ID No:16). A full length light chain amino acid sequence is Seq ID No:25 (light chain nucleic acid sequence Seq ID No:26).
[0253] 1D05 HC mutant 1 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:47, comprising the CDRH1 amino acid sequence of Seq ID No:27 (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). 1D05 HC mutant 1 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:43, comprising the CDRL1 amino acid sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), the CDRL2 amino acid sequence of Seq ID No:38 (IMGT) or Seq ID No:41 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:44. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length light chain amino acid sequence is Seq ID No:45 (light chain nucleic acid sequence Seq ID No:46).
[0254] 1D05 HC mutant 2 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:48, comprising the CDRH1 amino acid sequence of Seq ID No:27. (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). 1D05 HC mutant 2 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:43, comprising the CDRL1 amino acid sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), the CDRL2 amino acid sequence of Seq ID No:38 (IMGT) or Seq ID No:41 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:44. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length light chain amino acid sequence is Seq ID No:45 (light chain nucleic acid sequence Seq ID No:46).
[0255] 1D05 HC mutant 3 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:49, comprising the CDRH1 amino acid sequence of Seq ID No:27 (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). 1D05 HC mutant 3 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:43, comprising the CDRL1 amino acid sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), the CDRL2 amino acid sequence of Seq ID No:38 (IMGT) or Seq ID No:41 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:44. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length light chain amino acid sequence is Seq ID No:45 (light chain nucleic acid sequence Seq ID No:46).
[0256] 1D05 HC mutant 4 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:342, comprising the CDRH1 amino acid sequence of Seq ID No:27 (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). 1D05 HC mutant 4 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:43, comprising the CDRL1 amino add sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), the CDRL2 amino acid sequence of Seq ID No:38 (IMGT) or Seq ID No:41 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:44. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length light chain amino acid sequence is Seq ID No:45 (light chain nucleic acid sequence Seq ID No:46).
[0257] 1D05 LC mutant 1 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:33, comprising the CDRH1 amino acid sequence of Seq ID No:27 (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:34. 1D05 LC mutant 1 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:50, comprising the CDRL1 amino acid sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The CDRL2 sequence of 1D05 LC Mutant 1 is as defined by the Kabat or IMGT systems from the V.sub.L sequence of Seq ID No:50. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:35 (heavy chain nucleic acid sequence Seq ID No:36).
[0258] 1D05 LC mutant 2 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:33, comprising the CDRH1 amino acid sequence of Seq ID No:27 (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:34. 1D05 LC mutant 2 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:51, comprising the CDRL1 amino acid sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), the CDRL2 amino acid sequence of Seq ID No:38 (IMGT) or Seq ID No:41 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:35 (heavy chain nucleic acid sequence Seq ID No:36).
[0259] 1D05 LC mutant 3 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:33, comprising the CDRH1 amino acid sequence of Seq ID No:27 (IMGT) or Seq ID No:30 (Kabat), the CDRH2 amino acid sequence of Seq ID No:28 (IMGT) or Seq ID No:31 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:29 (IMGT) or Seq ID No:32 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:34. 1D05 LC mutant 3 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:298, comprising the CDRL1 amino acid sequence of Seq ID No:37 (IMGT) or Seq ID No:40 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:39 (IMGT) or Seq ID No:42 (Kabat). The CDRL2 sequence of 1D05 LC Mutant 3 is as defined by the Kabat or IMGT systems from the V.sub.L sequence of Seq ID No:298. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:44. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:35 (heavy chain nucleic acid sequence Seq ID No:36). A full length light chain amino acid sequence is Seq ID No:45 (light chain nucleic acid sequence Seq ID No:46).
[0260] 411B08 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:58, comprising the CDRH1 amino acid sequence of Seq ID No:52 (IMGT) or Seq ID No:55 (Kabat), the CDRH2 amino acid sequence of Seq ID No:53 (IMGT) or Seq ID No:56 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:54 (IMGT) or Seq ID No:57 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:59. 411B08 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:68, comprising the CDRL1 amino acid sequence of Seq ID No:62 (IMGT) or Seq ID No:65 (Kabat), the CDRL2 amino acid sequence of Seq ID No:63 (IMGT) or Seq ID No:66 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:64 (IMGT) or Seq ID No:67 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:69. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:60 (heavy chain nucleic acid sequence Seq ID No:61). A full length light chain amino acid sequence is Seq ID No:70 (light chain nucleic acid sequence Seq ID No:71).
[0261] 411C04 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:78, comprising the CDRH1 amino acid sequence of Seq ID No:72 (IMGT) or Seq ID No:75 (Kabat), the CDRH2 amino acid sequence of Seq ID No:73 (IMGT) or Seq ID No:76 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:74 (IMGT) or Seq ID No:77 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:79. 411C04 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:88, comprising the CDRL1 amino acid sequence of Seq ID No:82 (IMGT) or Seq ID No:85 (Kabat), the CDRL2 amino acid sequence of Seq ID No:83 (IMGT) or Seq ID No:86 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:84 (IMGT) or Seq ID No:87 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:89. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:80 (heavy chain nucleic acid sequence Seq ID No:81). A full length light chain amino acid sequence is Seq ID No:90 (light chain nucleic acid sequence Seq ID No:91).
[0262] 411D07 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:98, comprising the CDRH1 amino acid sequence of Seq ID No:92 (IMGT) or Seq ID No:95 (Kabat), the CDRH2 amino acid sequence of Seq ID No:93 (IMGT) or Seq ID No:96 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:94 (IMGT) or Seq ID No:97 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:99. 411D07 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:108, comprising the CDRL1 amino acid sequence of Seq ID No:102 (IMGT) or Seq ID No:105 (Kabat), the CDRL2 amino acid sequence of Seq ID No:103 (IMGT) or Seq ID No:106 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:104 (IMGT) or Seq ID No:107 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:109. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:100 (heavy chain nucleic acid sequence Seq ID No:101). A full length light chain amino acid sequence is Seq ID No:110 (light chain nucleic acid sequence Seq ID No:111).
[0263] 385F01 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:118, comprising the CDRH1 amino acid sequence of Seq ID No:112 (IMGT) or Seq ID No:115 (Kabat), the CDRH2 amino acid sequence of Seq ID No:113 (IMGT) or Seq ID No:116 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:114 (IMGT) or Seq ID No:117 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:119. 385F01 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:128, comprising the CDRL1 amino acid sequence of Seq ID No:122 (IMGT) or Seq ID No:125 (Kabat), the CDRL2 amino add sequence of Seq ID No:123 (IMGT) or Seq ID No:126 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:124 (IMGT) or Seq ID No:127 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:129. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:120 (heavy chain nucleic acid sequence Seq ID No:121). A full length light chain amino acid sequence is Seq ID No:130 (light chain nucleic acid sequence Seq ID No:131).
[0264] 386H03 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:158, comprising the CDRH1 amino acid sequence of Seq ID No:152 (IMGT) or Seq ID No:155 (Kabat), the CDRH2 amino acid sequence of Seq ID No:153 (IMGT) or Seq ID No:156 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:154 (IMGT) or Seq ID No:157 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:159. 386H03 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:168, comprising the CDRL1 amino acid sequence of Seq ID No:162 (IMGT) or Seq ID No:165 (Kabat), the CDRL2 amino acid sequence of Seq ID No:163 (IMGT) or Seq ID No:166 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:164 (IMGT) or Seq ID No:167 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:169. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:160 (heavy chain nucleic acid sequence Seq ID No:161). A full length light chain amino acid sequence is Seq ID No:170 (light chain nucleic acid sequence Seq ID No:171).
[0265] 389A03 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:178, comprising the CDRH1 amino acid sequence of Seq ID No:172 (IMGT) or Seq ID No:175 (Kabat), the CDRH2 amino acid sequence of Seq ID No:173 (IMGT) or Seq ID No:176 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:174 (IMGT) or Seq ID No:177 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:179. 389A03 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:188, comprising the CDRL1 amino acid sequence of Seq ID No:182 (IMGT) or Seq ID No:185 (Kabat), the CDRL2 amino acid sequence of Seq ID No:183 (IMGT) or Seq ID No:186 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:184 (IMGT) or Seq ID No:187 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:189. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:180 (heavy chain nucleic acid sequence Seq ID No:181). A full length light chain amino acid sequence is Seq ID No:190 (light chain nucleic acid sequence Seq ID No:191).
[0266] 413D08 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:138, comprising the CDRH1 amino acid sequence of Seq ID No:132 (IMGT) or Seq ID No:135 (Kabat), the CDRH2 amino acid sequence of Seq ID No:133 (IMGT) or Seq ID No:136 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:134 (IMGT) or Seq ID No:137 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:139. 413D08 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:148, comprising the CDRL1 amino acid sequence of Seq ID No:142 (IMGT) or Seq ID No:145 (Kabat), the CDRL2 amino acid sequence of Seq ID No:143 (IMGT) or Seq ID No:146 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:144 (IMGT) or Seq ID No:147 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:149. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No: 140 (heavy chain nucleic acid sequence Seq ID No:141). A full length light chain amino acid sequence is Seq ID No:150 (light chain nucleic acid sequence Seq ID No:151).
[0267] 413G05 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:244, comprising the CDRH1 amino acid sequence of Seq ID No:238 (IMGT) or Seq ID No:241 (Kabat), the CDRH2 amino acid sequence of Seq ID No:239 (IMGT) or Seq ID No:242 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:240 (IMGT) or Seq ID No:243 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:245. 413G05 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:254, comprising the CDRL1 amino acid sequence of Seq ID No:248 (IMGT) or Seq ID No:251 (Kabat), the CDRL2 amino acid sequence of Seq ID No:249 (IMGT) or Seq ID No:252 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:250 (IMGT) or Seq ID No:253 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:255. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:246 (heavy chain nucleic acid sequence Seq ID No:247). A full length light chain amino acid sequence is Seq ID No:256 (light chain nucleic acid sequence Seq ID No:257).
[0268] 413F09 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:264, comprising the CDRH1 amino acid sequence of Seq ID No:258 (IMGT) or Seq ID No:261 (Kabat), the CDRH2 amino acid sequence of Seq ID No:259 (IMGT) or Seq ID No:262 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:260 (IMGT) or Seq ID No:263 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:265. 413F09 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:274, comprising the CDRL1 amino acid sequence of Seq ID No:268 (IMGT) or Seq ID No:271 (Kabat), the CDRL2 amino acid sequence of Seq ID No:269 (IMGT) or Seq ID No:272 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:270 (IMGT) or Seq ID No:273 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:275. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:266 (heavy chain nucleic acid sequence Seq ID No:267). A full length light chain amino acid sequence is Seq ID No:276 (light chain nucleic acid sequence Seq ID No:277).
[0269] 414B06 has a heavy chain variable (V.sub.H) region amino acid sequence of Seq ID No:284, comprising the CDRH1 amino acid sequence of Seq ID No:278 (IMGT) or Seq ID No:281 (Kabat), the CDRH2 amino acid sequence of Seq ID No:279 (IMGT) or Seq ID No:282 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:280 (IMGT) or Seq ID No:283 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:285. 414E306 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:294, comprising the CDRL1 amino acid sequence of Seq ID No:288 (IMGT) or Seq ID No:291(Kabat), the CDRL2 amino acid sequence of Seq ID No:289 (IMGT) or Seq ID No:292 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:290 (IMGT) or Seq ID No:293 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:295. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:286 (heavy chain nucleic acid sequence Seq ID No:287). A full length light chain amino acid sequence is Seq ID No:296 (light chain nucleic acid sequence Seq ID No:297).
[0270] 416E01 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:349, comprising the CDRH1 amino acid sequence of Seq ID No:343 (IMGT) or Seq ID No:346 (Kabat), the CDRH2 amino acid sequence of Seq ID No:344 (IMGT) or Seq ID No:347 (Kabat), and the CDRH3 amino acid sequence of Seq ID No:345 (IMGT) or Seq ID No:348 (Kabat). The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:350. 416E01 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:359, comprising the CDRL1 amino acid sequence of Seq ID No:353 (IMGT) or Seq ID No:356 (Kabat), the CDRL2 amino acid sequence of Seq ID No:354 (IMGT) or Seq ID No:357 (Kabat), and the CDRL3 amino acid sequence of Seq ID No:355 (IMGT) or Seq ID No:358 (Kabat). The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:360. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:351 (heavy chain nucleic acid sequence Seq ID No:352). A full length light chain amino acid sequence is Seq ID No:361 (light chain nucleic acid sequence Seq ID No:362).
[0271] The antibodies of the invention are described with respect to the following concepts, aspects, sentences, arrangements and embodiments. Unless otherwise stated, all concepts, embodiments, sentences, arrangements and aspects are to be read as being able to be combined with any other concept, aspect, sentence, arrangement or embodiment, unless such combination would not make technical sense or is explicitly stated otherwise.
Concept 1. An antibody or a fragment thereof, which specifically binds to hPD-L1 as defined by Seq ID No:1, and competes for binding to said hPD-L1 with the antibody 1D05, wherein the antibody or fragment comprises a V.sub.H domain which comprises a CDRH3 comprising the motif X.sub.1GSGX.sub.2YGX.sub.3X.sub.4FD, wherein X.sub.1, X.sub.2 and X.sub.3 are independently any amino acid, and X.sub.4 is either present or absent, and if present, may be any amino acid.
[0272] In these concepts, antibodies or fragments may include or may not include bispecific antibodies. In one embodiment, in these concepts, antibodies or fragments includes bispecific antibodies. In one embodiment, a bispecific antibody does not include a FIT-Ig format. In one embodiment, a bispecific antibody does not include a mAb.sup.2 format. In one embodiment, a bispecific antibody does not include either a FIT-Ig format or a mAb.sup.2 format. In one embodiment, the antibody or fragment in these concepts includes a bispecific antibody, but does not include a bispecific antibody having a FIT-Ig format. In one embodiment, the antibody or fragment in these concepts includes a bispecific antibody, but does not include a bispecific antibody having a mAb.sup.2 format. In one embodiment, the antibody or fragment in these concepts includes a bispecific antibody, but does not include a bispecific antibody having a FIT-Ig format or a mAb.sup.2 format. In another embodiment, in these concepts, antibodies or fragments include dual binding antibodies.
[0273] Preferably, an antibody or a fragment thereof that specifically binds to a hPD-L1 antigen does not cross-react with other antigens (but may optionally cross-react with PD-L1 of a different species, e.g., rhesus, cynomolgus, or murine). An antibody or a fragment thereof that specifically binds to a hPD-L1 antigen can be identified, for example, by immunoassays, BIAcore.TM., or other techniques known to those of skill in the art. An antibody or a fragment thereof binds specifically to a hPD-L1 antigen when it binds to a hPD-L1 antigen with higher affinity than to any cross-reactive antigen as determined using experimental techniques, such as radioimmunoassays (RIA) and enzyme-linked immunosorbent assays (ELISAs). Typically, a specific or selective reaction will be at least twice background signal or noise and more typically more than 10 times background. See, e.g. Paul, ed., 1989, Fundamental Immunology Second Edition, Raven Press, New York at pages 332-336 for a discussion regarding antibody specificity.
[0274] In one embodiment, the antibody or fragment is a human antibody. In one embodiment, the antibody or fragment is a human antibody or fragment. In one embodiment, the antibody or fragment is a fully human antibody or fragment. In one embodiment, the antibody or fragment is a fully human monoclonal antibody or fragment.
[0275] There is also provided concept 1a: An antibody or a fragment thereof, that specifically binds to hPD-L1 as defined by Seq ID No:1, and competes for binding to said hPD-L1 with the antibody 411608, wherein the antibody or fragment comprises a V.sub.H domain which comprises a CDRH3 comprising the motif ARX.sub.1RX.sub.2X.sub.3SDX.sub.4X.sub.5D, wherein X.sub.1, X.sub.2, X.sub.3, X.sub.4 and X.sub.5 are independently any amino acid.
[0276] There is also provided concept 1b: An antibody or a fragment thereof, that specifically binds to hPD-L1 as defined by Seq ID No:1, and competes for binding to said hPD-L1 with the antibody 411608, wherein the antibody or fragment comprises a V.sub.H domain which comprises a CDRH3 comprising the motif X.sub.1RDGSGSY, wherein X.sub.1 is any amino acid.
[0277] As provided in the concepts or aspects herein, an anti-PD-L1 antibody or immunocytokine may bind to PD-L1, e.g. human PD-L1 with a K.sub.D of less than 50 nM, less than 40 nM, less than 30 nM as determined by surface plasmon resonance. Another embodiment, anti-PD-L1 antibody or immunocytokine may bind to PD-L1, e.g. human PD-L1 with a K.sub.D of less than 20 nM, less than 15 nM, less than 10 nM as determined by surface plasmon resonance. anti-PD-L1 antibody or immunocytokine may bind to PD-L1, e.g. human PD-L1 with a K.sub.D of less than 8 nM, less than 5 nM, less than 4 nM, less than 3 nM, less than 2 nM or less than 1 nM as determined by surface plasmon resonance. The K.sub.D may be 0.9 nM or less, 0.8 nM or less, 0.7 nM or less, 0.6 nM or less, 0.5 nM or less, 0.4 nM or less, 0.3 nM or less, 0.2 nM or less, or 0.1 nM or less.
[0278] In another embodiment, the K.sub.D is within a range of 0.01 to 1 nM, or a range of 0.05 to 2 nM, or a range of 0.05 to 1 nM. The K.sub.D may be with regard to hPD-L1, cynoPD-L1 and/or mouse PD-L1.
[0279] In another embodiment, the anti-PD-L1 antibodies described herein have a K.sub.ON rate (e.g. as measured by SPR, e.g. at 25.degree. C. or at 37.degree. C.) of approximately 0.5 to 10 .mu.M, for example approximately 1 to 8 .mu.M or approximately 1 to 7 .mu.M. In another embodiment, the K.sub.ON rate is approximately 1 to 5 .mu.M, e.g. approximately 1 .mu.M, approximately 1.5 .mu.M, approximately 2 .mu.M, approximately 2.5 .mu.M or approximately 3 .mu.M. In another embodiment, the K.sub.ON rate is approximately 3.5 .mu.M, approximately 4 .mu.M, approximately 4.5 .mu.M, approximately 5 .mu.M or approximately 5.5 .mu.M.
[0280] In another embodiment, the anti-PD-L1 antibodies described herein have a K.sub.OFF rate (e.g. as measured by SPR, e.g. at 25.degree. C. or at 37.degree. C.) of approximately 0.01 to 100 mM, for example approximately 0.1 to 50 mM or approximately 0.5 to 50 mM. In another embodiment, the K.sub.OFF rate is approximately 0.5 to 10 mM, or approximately 0.5 to 10 mM, e.g. approximately 1 mM, approximately 2 mM, approximately 3 mM, approximately 4 mM or approximately 5 mM. In another embodiment, the K.sub.OFF rate is approximately 0.6 mM, approximately 0.7 mM, approximately 0.8 mM or approximately 0.9 mM.
[0281] In another embodiment, the anti-PD-L1 antibodies (and immunocytokines) described in the concepts and aspects herein provide improved transient expression levels over other anti-PD-L1 antibodies and immunocytokines. Thus, in one embodiment, the anti-PD-L1 antibody (or immunocytokine) is expressed in a HEK293 cell, e.g. a HEK293T cell, at an expression level of approximately 100 .mu.g/mL, or in a range of approximately 100 to 350 .mu.g/mL. In another embodiment, the expression level is above approximately 350 .mu.g/mL.
[0282] In another embodiment, the anti-PD-L1 antibody (or immunocytokine) is expressed in a CHO cell, e.g. an Expi-CHO cell, at an expression level of approximately 100 .mu.g/mL, or in a range of approximately 100 to 350 .mu.g/mL. In another embodiment, the expression level is above approximately 350 .mu.g/mL.
[0283] In another embodiment, the anti-PD-L1 antibody (or immunocytokine) is expressed in a CHO cell, e.g. an Expi-CHO cell or a CHO-E7 EBNA cell, at an expression level of approximately 100 .mu.g/mL, or in a range of approximately 100 to 350 .mu.g/mL. In another embodiment, the expression level is above approximately 350 .mu.g/mL. The antibody described herein as 1D05, formatted as a human IgG1 (Seq ID No:340, at 2L volume in CHO-E7 EBNA cells has an expression level of approximately 115 pg/mL. The antibody described herein as 416E01, formatted as a human IgG1 (Seq ID No:340), at 2L volume in CHO-E7 EBNA cells has an expression level of approximately 160 .mu.g/mL. The antibody described herein as 1414606, formatted as a human IgG1 (Seq ID No:340), at 2L volume in CHO-E7 EBNA cells has an expression level of approximately 783 .mu.g/mL. The antibody described herein as 413G05, formatted as a human IgG1 (Seq ID No:340), at 2L volume in CHO-E7 EBNA cells has an expression level of approximately 383 .mu.g/mL.
[0284] In any of these expression systems, the expression is carried out of a scale of between approximately 0.5 mL and 3 mL, for example between approximately 0.5 mL and 2 mL. In any of these expression systems, the anti-PD-L1 antibody (or immunocytokine) may be expressed from a pTT5 vector. In any of these expression systems, the anti-PD-L1 antibody (or immunocytokine) may be expressed in conjunction with a lipid transfection reagent, and may optionally be expressed in a CHO cell, e.g. an Expi-CHO cell. In any of these expression systems, the anti-PD-L1 antibody (or immunocytokine) may be expressed in conjunction with a PEI transfection reagent, and may optionally be expressed in a CHO cell, e.g. an CHO-E7 EBNA cell. In any of these expression systems, the anti-PD-L1 antibody (or immunocytokine) may be expressed in conjunction with a helper plasmid (e.g. an AKT helper plasmid), and may optionally be expressed in a CHO cell, e.g. an CHO-E7 EBNA cell.
[0285] In any of these expression systems, the expression level is between approximately 100 .mu.g/mL and approximately 1500 .mu.g/mL, for example between approximately 100 .mu.g/mL and approximately 1000 .mu.g/mL, or between approximately 200 .mu.g/mL and approximately 1000 .mu.g/mL, or between approximately 350 .mu.g/mL and approximately 1000 .mu.g/mL. In any of these expression systems, the lower limit of expression may be approximately 100 .mu.g/mL, approximately 200 .mu.g/mL, approximately 300 .mu.g/mL, or approximately 400 .mu.g/mL. In another embodiment, the lower limit of expression may be approximately 500 .mu.g/mL, approximately 600 .mu.g/mL, approximately 700 .mu.g/mL, or approximately 800 .mu.g/mL. In any of these expression systems, the upper limit of expression may be approximately 2000 .mu.g/mL, approximately 1800 .mu.g/mL, approximately 1600 .mu.g/mL, or approximately 1500 .mu.g/mL. In another embodiment, the upper limit of expression may be approximately 1250 .mu.g/mL, approximately 1000 .mu.g/mL, approximately 900 .mu.g/mL, or approximately 800 .mu.g/mL.
[0286] In another embodiment, the expression system is a Lonza expression system, e.g. Lonza X-Ceed.RTM. system. In the Lonza expression system, the expression may be carried out at a scale of approximately 30 mL to 2 L, for example 50 mL to 1 L, or 1 L tp 2 L. In the Lonza expression system, the anti-PD-L1 antibody (or immunocytokine) may be expressed in conjunction with electroporation, and optionally without any helper plasmids. In the Lonza expression system, the anti-PD-L1 antibody (or immunocytokine) may be expressed at a level of approximately 1 g/L, or approximately 900 mg/L, or approximately 800 mg/L, or approximately 700 mg/L. In another embodiment, In the Lonza expression system, the anti-PD-L1 antibody (or immunocytokine) may be expressed at a level of approximately 600 mg/L or approximately 500 mg/L or approximately 400 mg/L. In the Lonza expression system, the anti-PD-L1 antibody (or immunocytokine) may be expressed at a level of between approximately 400 mg/L and approximately 2 g/L, for example between approximately 500 mg/L and approximately 1.5 g/L, or between approximately 500 mg/L and approximately 1 g/L. In another embodiment, the expression level is above 1 g/L. In another embodiment, the anti-PD-L1 antibodies described in the concepts provide improved half-life over other anti-PD-L1 antibodies as further described in Aspect 1 hereinbelow.
Concept 2. The antibody or fragment according to concept 1, wherein X.sub.1 is a hydroxyl-containing amino acid, optionally T.
[0287] In one embodiment, the hydroxyl-containing amino acid is Serine. In one embodiment, the hydroxyl-containing amino acid is Cysteine. In one embodiment, the hydroxyl-containing amino acid is Threonine. In one embodiment, the hydroxyl-containing amino acid is Methionine. In one embodiment, the hydroxyl-containing amino acid is Serine or Cysteine. In one embodiment, the hydroxyl-containing amino acid is Serine or Threonine. In one embodiment, the hydroxyl-containing amino acid is Serine or Methionine. In one embodiment, the hydroxyl-containing amino acid is Cysteine or Threonine. In one embodiment, the hydroxyl-containing amino acid is Cysteine or Methionine. In one embodiment, the hydroxyl-containing amino acid is Threonine or Methionine.
[0288] In one embodiment, the hydroxyl-containing amino acid is selected from serine, cysteine, threonine and methionine.
[0289] Concept 2a. The antibody or fragment according to concept 1a, wherein X.sub.1 is an aliphatic amino acid or an amide amino acid.
[0290] In one embodiment, X.sub.1 is selected from Asparagine (N) and valine (V). In one embodiment, X.sub.1 is valine. In one embodiment, X.sub.1 is asparagine.
[0291] Concept 2b. The antibody or fragment according to concept 1b, wherein X.sub.1 is an aliphatic amino acid.
[0292] In one embodiment, X.sub.1 is selected from alanine (A) or valine (V). In one embodiment, X.sub.1 is valine. In one embodiment, X.sub.1 is alanine.
Concept 3. The antibody or fragment according to concept 1 or concept 2, wherein X.sub.2 is a basic amino acid, optionally K.
[0293] In one embodiment, the hydroxyl-containing amino acid is Histidine. In one embodiment, the hydroxyl-containing amino acid is Lysine. In one embodiment, the hydroxyl-containing amino acid is Arginine. In one embodiment, the hydroxyl-containing amino acid is Histidine or Lysine. In one embodiment, the hydroxyl-containing amino acid is Histidine or Arginine. In one embodiment, the hydroxyl-containing amino acid is Lysine or Arginine.
[0294] In one embodiment, the hydroxyl-containing amino acid is selected from Histidine, Lysine and Arginine.
[0295] Concept 3a. The antibody or fragment according to concept 1a or concept 2a, wherein X.sub.2 is an aliphatic amino acid or an amide amino acid.
[0296] In one embodiment, X.sub.2 is selected from leucine (L), isoleucine (I), Valine (V), Asparagine (N) and glutamine (Q). In one embodiment, X.sub.2 is selected from leucine (L), isoleucine (I) and Valine (V).
[0297] In one embodiment, X.sub.2 is selected from Asparagine (N) and glutamine (Q) In one embodiment, X.sub.2 is selected from leucine (L) and glutamine (Q). In one embodiment, X.sub.2 is leucine (L). In one embodiment, X.sub.2 is glutamine (Q).
Concept 4. The antibody or fragment according to any one of concepts 1 to 3, wherein X.sub.2 is a hydroxyl-containing amino acid, optionally S or T.
[0298] In one embodiment, the hydroxyl-containing amino acid is Serine. In one embodiment, the hydroxyl-containing amino acid is Cysteine. In one embodiment, the hydroxyl-containing amino acid is Threonine. In one embodiment, the hydroxyl-containing amino acid is Methionine. In one embodiment, the hydroxyl-containing amino acid is Serine or Cysteine. In one embodiment, the hydroxyl-containing amino acid is Serine or Threonine. In one embodiment, the hydroxyl-containing amino acid is Serine or Methionine. In one embodiment, the hydroxyl-containing amino acid is Cysteine or Threonine. In one embodiment, the hydroxyl-containing amino acid is Cysteine or Methionine. In one embodiment, the hydroxyl-containing amino acid is Threonine or Methionine.
[0299] In one embodiment, the hydroxyl-containing amino acid is selected from serine, cysteine, threonine and methionine.
[0300] Concept 4a. The antibody or fragment according to any one of concepts 1a, 2a or 3a, wherein X.sub.3 is an aromatic amino acid.
[0301] In one embodiment, X.sub.3 is selected from Phenylalanine (F), Tyrosine (Y) and Tryptophan (W). In one embodiment, X.sub.3 is selected from Tyrosine (Y) and Tryptophan (W). In one embodiment, X.sub.3 is Tyrosine (Y). In one embodiment, X.sub.3 is Tryptophan (W).
Concept 5. The antibody or fragment according to any one of concepts 1 to 4, wherein X.sub.3 is an aromatic amino acid, optionally W.
[0302] In one embodiment, the hydroxyl-containing amino acid is Phenylalanine. In one embodiment, the hydroxyl-containing amino acid is Tyrosine. In one embodiment, the hydroxyl-containing amino acid is Tryptophan. In one embodiment, the hydroxyl-containing amino acid is Phenylalanine or
[0303] Tyrosine. In one embodiment, the hydroxyl-containing amino acid is Phenylalanine or Tryptophan. In one embodiment, the hydroxyl-containing amino acid is Tyrosine or Tryptophan.
[0304] In one embodiment, the hydroxyl-containing amino acid is selected from Phenylalanine, Tyrosine and Tryptophan.
[0305] Concept 5a. The antibody or fragment according to any one of concepts 1a, 2a, 3a or 4a wherein X.sub.4 is an aromatic amino acid.
[0306] In one embodiment, X.sub.4 is selected from Phenylalanine (F), Tyrosine (Y) and Tryptophan (W). In one embodiment, X.sub.4 is selected from Tyrosine (Y) and Phenylalanine (F). In one embodiment, X.sub.4 is Tyrosine (Y). In one embodiment, X.sub.4 is Phenylalanine (F).
Concept 6. The antibody or fragment according to any one of concepts 1 to 5, wherein X.sub.4 is absent.
[0307] Concept 6a. The antibody or fragment according to any one of concepts 1a, 2a, 3a, 4a or 5a wherein X.sub.5 is an aliphatic amino acid or an hydroxyl-containing amino acid.
[0308] In one embodiment, X.sub.5 is selected from leucine (L), isoleucine (I), Valine (V), Serine (S), Cysteine (C) and Threonine (T). In one embodiment, X.sub.5 is selected from leucine (L), isoleucine (I) and Valine (V). In one embodiment, X.sub.5 is selected from Serine (S), Cysteine (C) and Threonine (T). In one embodiment, X.sub.5 is selected from leucine (L) and Serine (S). In one embodiment, X.sub.5 is Serine (S). In one embodiment, X.sub.5 is leucine (L).
Concept 7. The antibody or fragment according to any one of concepts 1 to 5, wherein X.sub.4 is present. Concept 8. The antibody or fragment according to concept 7, wherein X.sub.4 is an aliphatic amino acid, optionally G.
[0309] In one embodiment, the hydroxyl-containing amino acid is selected from Glycine, Alanine, Valine, Leucine and Isoleucine.
[0310] In one embodiment, the hydroxyl-containing amino acid is selected from Glycine and Alanine. In one embodiment, the hydroxyl-containing amino acid is selected from Glycine and Valine. In one embodiment, the hydroxyl-containing amino acid is selected from Glycine and Leucine. In one embodiment, the hydroxyl-containing amino acid is selected from Glycine and Isoleucine. In one embodiment, the hydroxyl-containing amino acid is selected from Alanine and Valine. In one embodiment, the hydroxyl-containing amino acid is selected from Alanine and Leucine. In one embodiment, the hydroxyl-containing amino acid is selected from Alanine and Isoleucine. In one embodiment, the hydroxyl-containing amino acid is selected from Valine and Leucine. In one embodiment, the hydroxyl-containing amino acid is selected from Valine and Isoleucine. In one embodiment, the hydroxyl-containing amino acid is selected from, Leucine and Isoleucine.
[0311] In one embodiment, the hydroxyl-containing amino acid selected from three of each of Glycine, Alanine, Valine, Leucine and Isoleucine. In one embodiment, the hydroxyl-containing amino acid selected from four of each of Glycine, Alanine, Valine, Leucine and Isoleucine.
Concept 9. An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 1D05, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:29 or 32, or the CDRH3 sequence of SEQ ID NO:29 or 32 comprising 6 or fewer amino acid substitutions.
[0312] Concept 9a: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 84G09, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:9 or 12, or the CDRH3 sequence of SEQ ID NO:9 or 12 comprising 6 or fewer amino acid substitutions.
[0313] Concept 9b: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 411B08, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:54 or 57, or the CDRH3 sequence of SEQ ID NO:54 or 57 comprising 6 or fewer amino acid substitutions.
[0314] Concept 9c: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 411C04, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID No:74 or 77, or the CDRH3 sequence of SEQ ID NO:74 or 77 comprising 6 or fewer amino acid substitutions.
[0315] Concept 9d: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 411D07, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:94 or 97, or the CDRH3 sequence of SEQ ID NO:94 or 97 comprising 3 or fewer amino acid substitutions.
[0316] Concept 9e: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 385F01, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:114 or 117, or the CDRH3 sequence of SEQ ID NO:114 or 117 comprising 6 or fewer amino acid substitutions.
[0317] Concept 9f: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 386H03, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:144 or 147, or the CDRH3 sequence of SEQ ID NO:144 or 147 comprising 3 or fewer amino acid substitutions.
[0318] Concept 9g: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 389A03, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:174 or 177, or the CDRH3 sequence of SEQ ID NO:174 or 177 comprising 6 or fewer amino acid substitutions.
[0319] Concept 9h: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 413D08, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:134 or 137, or the CDRH3 sequence of SEQ ID NO:134 or 137 comprising 5 or fewer amino acid substitutions.
[0320] Concept 9i: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 413G05, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:240 or 243, or the CDRH3 sequence of SEQ ID NO:240 or 243 comprising 6 or fewer amino acid substitutions.
[0321] Concept 9j: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 413F09, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:260 or 263, or the CDRH3 sequence of SEQ ID NO:260 or 263 comprising 6 or fewer amino acid substitutions.
[0322] Concept 9k: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 414B06, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:280 or 283, or the CDRH3 sequence of SEQ ID NO:280 or 283 comprising 6 or fewer amino acid substitutions.
[0323] Concept 9l: An antibody or a fragment thereof, optionally according to any one of concepts 1 to 8, which specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 416E01, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID No:345 or 348, or the CDRH3 sequence of SEQ ID No:345 or 348 comprising 6 or fewer amino acid substitutions.
[0324] In all of concepts 9, 9a to I, 17, 17a to I, 18, 18a to I, 19, 19a to I, 22, 22a to I, 23, 23a to I, 24 and 24a to I, in one embodiment, the CDR comprises one amino acid substitution, which may be a conservative amino acid substitution. In all of concepts 9, 9a to I, 17, 17a to I, 18, 18a to I, 19, 19a to I, 22, 22a to I, 23, 23a, 24 and 24a to I, in one embodiment, the CDR comprises two amino acid substitutions, which may be conservative amino acid substitutions. In all of concepts 9, 9a to I, 17, 17a to I, 18, 18a to I, 19, 19a to I, 22, 22a, 22b, 22d, 22f, 22g, 24 and 24a to I, in one embodiment, the CDR comprises three amino acid substitutions, which may be conservative amino acid substitutions. In all of concepts 9, 9a to c, 9e, 9g to I, 17, 17a to c, 17e, 17g to I, 19, 19a, 22, 22d, 22f, 22g, 24 and 24a to I, in one embodiment, the CDR comprises four amino acid substitutions, which may be conservative amino acid substitutions. In all of concepts 9, 9a to c, 9e, 9g to I, 17, 17a to c, 17e, 17g to I, 22d, 22f and 22g, in one embodiment, the CDR comprises five amino acid substitutions, which may be conservative amino acid substitutions. In all of concepts 9, 9a to c, 9e, 9g, 9i to I, 17, 17a to c, 17e, 17g and 17i to I, in one embodiment, the CDR comprises six amino acid substitutions, which may be conservative amino acid substitutions.
[0325] Amino acid substitutions include alterations in which an amino acid is replaced with a different naturally-occurring amino acid residue. Such substitutions may be classified as "conservative", in which case an amino acid residue contained in a polypeptide is replaced with another naturally occurring amino acid of similar character either in relation to polarity, side chain functionality or size. Such conservative substitutions are well known in the art. Substitutions encompassed by the present invention may also be "non-conservative", in which an amino acid residue which is present in a peptide is substituted with an amino acid having different properties, such as naturally-occurring amino acid from a different group (e.g. substituting a charged or hydrophobic amino; acid with alanine), or alternatively, in which a naturally-occurring amino acid is substituted with a non-conventional amino acid.
[0326] In one embodiment, the conservative amino acid substitutions are as described herein. For example, the substitution may be of Y with F, T with S or K, P with A, E with D or Q, N with D or G, R with K, G with N or A, T with S or K, D with N or E, I with L or V, F with Y, S with T or A, R with K, G with N or A, K with R, A with S, K or P. In another embodiment, the conservative amino acid substitutions may be wherein Y is substituted with F, T with A or S, I with L or V, W with Y, M with L, N with D, G with A, T with A or S, D with N, I with L or V, F with Y or L, S with A or T and A with S, G, T or V.
Concept 10. An antibody or fragment which specifically binds to hPD-L1 and comprises a V.sub.H domain comprising a CDRH3 of from 12 to 20 amino acids and which is derived from the recombination of a human V.sub.H gene segment, a human D gene segment and a human J.sub.H gene segment, wherein the human J.sub.H gene segment is IGHJ5 (e.g. IGHJ5*02).
[0327] In one embodiment, the CDRH3 is from 14 to 17 amino acids and the human J.sub.H gene segment is IGHJ5 (e.g. IGHJ5*02).
[0328] There is also provided as concept 10a an antibody or fragment which specifically binds to hPD-L1 and comprises a V.sub.H domain comprising a CDRH3 of from 8 to 16 amino acids and which is derived from the recombination of a human V.sub.H gene segment, a human D gene segment and a human J.sub.H gene segment, wherein the human J.sub.H gene segment is selected from IGHJ4 (e.g. IGHJ4*02), IGHJ5 (e.g. IGHJ5*02) and IGHJ6 (e.g. IGHJ6*02).
[0329] In another embodiment, the human J.sub.H gene segment is IGHJ6 (e.g. IGHJ6*02). In another embodiment, the CDRH3 is of from 10 to 17 amino acids and the human J.sub.H gene segment is IGHJ6 (e.g. IGHJ6*02).
[0330] In another embodiment, the human J.sub.H gene segment is IGHJ4 (e.g. IGHJ4*02). In another embodiment, the CDRH3 is from 7 to 17 amino acids and the human J.sub.H gene segment is IGHJ4 (e.g. IGHJ4*02).
[0331] Optionally, the antibody of concept 10 or 10a has any of the features of concepts 1 to 9, including the binding affinities, Kon and Koff rates, expression levels, half-life etc.
Concept 11. The antibody or fragment according to concept 10 or 10a, wherein the human V.sub.H gene segment is IGHV3 (e.g. IGHV3-9, such as IGHV3-9*01).
[0332] There is also provided as concept 11a an antibody or fragment according to concept 10 or 10a, wherein the human V.sub.H gene segment is selected from IGHV3 (e.g. IGHV3-9, such as IGHV3-9*01 or e.g. IGHV3-7, such as IGHV3-7*01 or e.g. IGHV3-33, such as IGHV3-33*01 or e.g. IGHV3-11, such as IGHV3-11*01 or e.g. IGHV3-23, such as IGHV3-23*04), or IGHV4 (e.g. IGHV4-4, such as IGHV4-4*02 or e.g. IGHV4-39, such as IGHV4-39*01).
[0333] In one embodiment, the human V.sub.H gene segment is IGHV3 (e.g. IGHV3-7, such as IGHV3-7*01). In one embodiment, the human V.sub.H gene segment is IGHV3 (e.g. IGHV3-33, such as IGHV3-33*01). In one embodiment, the human V.sub.H gene segment is IGHV3 (e.g. IGHV3-11, such as IGHV3-11*01). In one embodiment, the human V.sub.H gene segment is IGHV3 (e.g. IGHV3-23, such as IGHV3-23*04).
[0334] In one embodiment, the human V.sub.H gene segment is IGHV4 (e.g. e.g. IGHV4-4, such as IGHV4-4*02). In one embodiment, the human V.sub.H gene segment is IGHV4 (e.g. IGHV4-39, such as IGHV4-39*01).
[0335] There is also provided as concept 11b an antibody or fragment according to concept 10, 10a, 11 or 11a, wherein the human D gene segment is selected from IGHD1 (e.g. IGHD1-20, such as IGHD1-20*01), IGHD3 (e.g. IGHD3-10, such as IGHD3-10*01), IGHD4 (e.g. IGHD4-11, such as IGHD4-11*01), IGHD5 (e.g. IGHD5-7, such as IGHD5-18*01), and IGHD6 (e.g. IGHD6-13, such as IGHD6-13*01).
[0336] In one embodiment, the human D gene segment is IGHD1 (e.g. IGHD1-20, such as IGHD1-20*01). In one embodiment, the human D gene segment is IGHD3 (e.g. IGHD3-10, such as IGHD3-10*01). In one embodiment, the human D gene segment is IGHD4 (e.g. IGHD4-11, such as IGHD4-11*01). In one embodiment, the human D gene segment is IGHD5 (e.g. IGHD5-18, such as IGHD5-19*01). In one embodiment, the human D gene segment is IGHD6 (e.g. IGHD6-13, such as IGHD6-13*01).
[0337] In any of concepts 10, 11 and 11a, the V.sub.H, D.sub.H and J.sub.H gene segments are as described in the combinations for the antibodies in Table 5 hereinbelow. In one embodiment, the antibody heavy chain is derived from a combination of IGHV3 (e.g. IGHV3-7 such as IGHV3-7*01), IGHD4 (e.g. IGHD4-11 such as IGHD4-11*01) and IGHJ4 (e.g. IGHJ4*02). In one embodiment, the antibody heavy chain is derived from a combination of IGHV4 (e.g. IGHV4-4 such as IGHV4-4*02), IGHD3 (e.g. IGHD3-10 such as IGHD3-10*01) and IGHJ4 (e.g. IGHJ4*02). In one embodiment, the antibody heavy chain is derived from a combination of IGHV4 (e.g. IGHV4-39 such as IGHV4-39*01), IGHD6 (e.g. IGHD6-13 such as IGHD6-13*01) and IGHJ1 (e.g. IGHJ1*01). In one embodiment, the antibody heavy chain is derived from a combination of IGHV3 (e.g. IGHV3-33 such as IGHV3-33*01), IGHD5 (e.g. IGHD5-18 such as IGHD5-18*01) and IGHJ6 (e.g. IGHJ6*02). In one embodiment, the antibody heavy chain is derived from a combination of IGHV3 (e.g. IGHV3-11 such as IGHV3-11*01), IGHD1 (e.g. IGHD1-20 such as IGHD1-20*01) and IGHJ6 (e.g. IGHJ6*02). In one embodiment, the antibody heavy chain is derived from a combination of IGHV3 (e.g. IGHV3-23 such as IGHV3-23*04), IGHD5 (e.g. IGHD5-18 such as IGHD5-18*01) and IGHJ4 (e.g. IGHJ4*02). In one embodiment, the antibody heavy chain is derived from a combination of IGHV3 (e.g. IGHV3-7 such as IGHV3-7*01), IGHD5 (e.g. IGHD5-24 such as IGHD5-24*01) and IGHJ4 (e.g. IGHJ4*02). In one embodiment, the antibody heavy chain is derived from a combination of IGHV3 (e.g. IGHV3-23 such as IGHV3-23*04), IGHD6 (e.g. IGHD6-13 such as IGHD6-13*01) and IGHJ4 (e.g. IGHJ4*02).
Concept 12. The antibody or fragment according to concept 10, 10a, 11, 11a or 11b, wherein the antibody or fragment comprises a V.sub.L domain which is derived from the recombination of a human V.sub.K gene segment, and a human J.sub.K gene segment, wherein the human V.sub.K gene segment is IG.kappa.V1D (e.g. IG.kappa.V1D-39, such as IG.kappa.V1D-39*01).
[0338] There is also provided as concept 12a an antibody or fragment according to any of concepts 10, 10a, 11, 11a or 11b, wherein the human V.sub.K gene segment is selected from IG.kappa.V1 (e.g. IG.kappa.V1-17, such as IG.kappa.V1-17*01 or e.g. IG.kappa.V1-9, such as IG.kappa.V1-9*d01 or e.g. IG.kappa.V1D-12, such as IG.kappa.V1D-12*02 or e.g. IG.kappa.V1D-39, such as IG.kappa.V1D-39*01), and IG.kappa.V4 (e.g. IG.kappa.V4-1, such as IG.kappa.V4-1*01).
[0339] In one embodiment, the human V.kappa. gene segment is IG.kappa.V1 (e.g. IG.kappa.V1-17, such as IG.kappa.V1-17*01). In one embodiment, the human V.kappa. gene segment is IG.kappa.V1 (e.g. IG.kappa.V1-9, such as IG.kappa.V1-9*d01). In one embodiment, the human V.kappa. gene segment is IG.kappa.V1 (e.g. IG.kappa.V1D-12, such as IG.kappa.V1D-12*02). In one embodiment, the human V.kappa. gene segment is IG.kappa.V1 (e.g. IG.kappa.V1D-39, such as IG.kappa.V1D-39*01).
[0340] In one embodiment, the human V.kappa. gene segment is IG.kappa.V1 IG.kappa.V4 (e.g. IG.kappa.V4-1, such as IG.kappa.V4-1*01)
[0341] There is also provided as concept 12b an antibody or fragment according to concept 10, 10a, 11 or 11a, wherein the human 3K gene segment is selected from Ian (e.g. IG.kappa.J1*01), IG.kappa.J2 (e.g. IG.kappa.J2*04), IG.kappa.J3 (e.g. IG.kappa.J3*01), IG.kappa.J4 (e.g. IG.kappa.J4*01) or IG.kappa.J5 (e.g. IG.kappa.J5*01).
[0342] In one embodiment, the human J.kappa. gene segment is IG.kappa.J1 (e.g. IG.kappa.J1*01). In one embodiment, the human 3K gene segment is IG.kappa.J2 (e.g. IG.kappa.J2*04). In one embodiment, the human J.kappa. gene segment is IG.kappa.J3 (e.g. IG.kappa.J3*01). In one embodiment, the human J.kappa. gene segment is IG.kappa.J4 (e.g. IG.kappa.J4*01). In one embodiment, the human J.kappa. gene segment is IG.kappa.J5 (e.g. IG.kappa.J5*01).
[0343] In any of concepts 12 and 12a, the V.kappa. and J.kappa. gene segments are as described in the combinations for the antibodies in Table 5 hereinbelow. In one embodiment, the antibody light chain is derived from a combination of IG.kappa.V1D (e.g. IG.kappa.V1D-12 such as IG.kappa.V1D-12*02) and IG.kappa.J3 (e.g. IG.kappa.J3*01). In one embodiment, the antibody light chain is derived from a combination of IG.kappa.V4 (e.g. IG.kappa.V4-1 such as IG.kappa.V14-1*01) and IG.kappa.J2 (e.g. IG.kappa.J2*04). In one embodiment, the antibody light chain is derived from a combination of IG.kappa.V1 (e.g. IG.kappa.V1-17 such as IG.kappa.V1-17*01) and IG.kappa.J1 (e.g. IG.kappa.J1*01). In one embodiment, the antibody light chain is derived from a combination of IG.kappa.V1D (e.g. IG.kappa.V1D-12,such as IG.kappa.V1D-12*02) and IG.kappa.J4 (e.g. IG.kappa.J4*01). In one embodiment, the antibody light chain is derived from a combination of IG.kappa.V1 (e.g. IG.kappa.V1-9 such as IG.kappa.V1-9*d01) and IG.kappa.J5 (e.g. IG.kappa.J5*01). In one embodiment, the antibody light chain is derived from a combination of IG.kappa.V1D (e.g. IG.kappa.V1D-12 such as IG.kappa.V1D-12*02) and IG.kappa.J5 (e.g. IG.kappa.J5*01).
Concept 13. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 1D05 specifically binds.
[0344] Concept 13a. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 84G09 specifically binds.
[0345] Concept 13b. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 411B08 specifically binds.
[0346] Concept 13c. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 411C04 specifically binds.
[0347] Concept 13d. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 411D07 specifically binds.
[0348] Concept 13e. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 385F01 specifically binds.
[0349] Concept 13f. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 386H03 specifically binds.
[0350] Concept 13g. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 389A03 specifically binds.
[0351] Concept 13h. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 413D08 specifically binds.
[0352] Concept 13i. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 413G05 specifically binds.
[0353] Concept 13j. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 413F09 specifically binds.
[0354] Concept 13k. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 414B06 specifically binds.
[0355] Concept 13l. An antibody or fragment thereof which specifically binds to an epitope that is identical to an epitope to which the antibody 416E01 specifically binds.
[0356] The antibodies described in these concepts have the sequences as described hereinabove.
[0357] In one embodiment, there is provided an antibody which specifically binds to an epitope which is substantially similar to an epitope to which any of the antibodies in concept 13, 13 a to 13l bind.
[0358] Contact amino acid residues involved in the interaction of antibody and antigen may be determined by various known methods to those skilled in the art.
[0359] In one embodiment, sequential replacement of the amino acids of the antigen sequence (using standard molecular biology techniques to mutate the DNA of the coding sequence, of the antigen), in this case hPD-L1 with Alanine (a.k.a Alanine scan), or another unrelated amino acid, may provide residues whose mutation would reduce or ablate the ability of the antibody to recognise the antigen in question. Binding may be assessed using standard techniques, such as, but not limited to, SPR, HTRF, ELISA (which are described elsewhere herein). Other substitutions could be made to enhance the disruption of binding such as changing the charge on the side chain of antigen sequence amino acids (e.g. Lysine change to glutamic acid), switching polar and non-polar residues (e.g. Serine change to leucine). The alanine scan or other amino substitution method may be carried out either with recombinant soluble antigen, or where the target is a cell membrane target, directly on cells using transient or stable expression of the mutated versions.
[0360] In one embodiment, protein crystallography may be used to determine contact residues between antibody and antigen (i.e. to determine the epitope to which the antibody binds), crystallography allows the direct visualisation of contact residues involved in the antibody-antigen interaction. As well as standard X-ray crystallography, cryo-electro microscopy has been used to determine contact residues between antibodies and HIV capsid protein (see Lee, Jeong Hyun, et al. "Antibodies to a conformational epitope on gp41 neutralize HIV-1 by destabilizing the Env spike.", Nature communications, 6, (2015)).
[0361] In one embodiment, if the antibody recognises a linear epitope, short peptides based on the antigen sequence can be produced and binding of the antibody to these peptides can be assessed using standard techniques, such as, but not limited to, SPR, HTRF, ELISA (which are described elsewhere herein). Further investigation of the epitope could be provided by performing an Alanine scan on any peptides that show binding. Alternative to linear peptides, conformational scans could be carried out using Pepscan technology (http://www.pepscan.com/) using their chemical linkage of peptides onto scaffolds, which has been used to determine discontinuous epitopes on CD20 targeting antibodies (Niederfellner, Gerhard, et al. "Epitope characterization and crystal structure of GA101 provide insights into the molecular basis for type I/II distinction of CD20 antibodies.", Blood, 118.2, (2011), 358-367.).
[0362] In one embodiment, limited proteolytic digestion and mass spectrophotometry can be used to identify binding epitopes. The antibody-antigen complex is digested by a protease, such as, but not limited to, trypsin. The digested complex peptides are compared to antibody-alone and antigen-alone digestion mass spectrophotometry to determine if a particular epitope is protected by the complexation. Further work involving amino acid substitution, competition binding, may then be employed to narrow down to individual amino acid residues involved in the interaction (see, for example, Suckau, Detlev, et al. "Molecular epitope identification by limited proteolysis of an immobilized antigen-antibody complex and mass spectrometric peptide mapping.", Proceedings of the National Academy of Sciences, 87.24, (1990), 9848-9852).
[0363] Thus, in one embodiment, the contact residues of the epitope are identified with an unrelated amino acid scan (e.g. alanine scan). In another embodiment, an unrelated amino acid scan (e.g. alanine scan) is carried out using a technique selected from SPR, HTRF, ELISA, X-ray crystallography, cryo-electro microscopy and a combination of limited proteolytic digestion and mass spectrometry. In one embodiment, the unrelated amino acid scan (e.g. alanine scan) is carried out using HTRF. In one embodiment, the unrelated amino add scan (e.g. alanine scan) is carried out using ELISA.
[0364] When the alanine scan is carried out with either ELISA or HTRF, an amino add residue is identified as contributing to the epitope if the reduction in signal is at least 25%. In one embodiment, the reduction in signal is at least 30%. In one embodiment, the reduction in signal is at least 35%. In one embodiment, the reduction in signal is at least 40%. In one embodiment, the reduction in signal is at least 45%. In one embodiment, the reduction in signal is at least 50%. In one embodiment, the reduction in signal is at least 55%. In one embodiment, the reduction in signal is at least 60%. In one embodiment, the reduction in signal is at least 70%. In one embodiment, the reduction in signal is at least 75%. In one embodiment, the reduction in signal is at least 80%. In one embodiment, the reduction in signal is at least 85%. In one embodiment, the reduction in signal is at least 90%.
[0365] When the alanine scan is carried out with SPR, an amino acid residue is identified as contributing to the epitope if there is at least a 10-fold reduction in affinity. In one embodiment, the reduction in affinity is at least 15-fold. In one embodiment, the reduction in affinity is at least 20-fold. In one embodiment, the reduction in affinity is at least 30-fold. In one embodiment, the reduction in affinity is at least 40-fold. In one embodiment, the reduction in affinity is at least 50-fold. In one embodiment, the reduction in affinity is at least 100-fold.
[0366] In one embodiment, the contact residues of the epitope are identified by X-ray crystallography. In one embodiment, the contact residues of the epitope are identified by cryo-electro microscopy. In one embodiment, the contact residues of the epitope are identified by a combination of limited proteolytic digestion and mass spectrometry.
Concept 14. The antibody or fragment according to concept 13, wherein the epitope is identified by unrelated amino acid scanning, or by X-ray crystallography. Concept 15. The antibody or fragment according to concept 14, wherein the contact residues of the epitope are defined by a reduction in affinity of at least 10-fold in an unrelated amino acid scan, e.g. an alanine scan as determined by SPR.
[0367] In one embodiment, the reduction in affinity is at least 15-fold. In one embodiment, the reduction in affinity is at least 20-fold. In one embodiment, the reduction in affinity is at least 30-fold. In one embodiment, the reduction in affinity is at least 40-fold. In one embodiment, the reduction in affinity is at least 50-fold. In one embodiment, the reduction in affinity is at least 100-fold.
[0368] SPR may be carried out as described hereinabove.
Concept 16. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 1D05.
[0369] Competition may be determined by surface plasmon resonance (SPR), such techniques being readily apparent to the skilled person. SPR may be carried out using Biacore.TM., Proteon.TM. or another standard SPR technique. Such competition may be due, for example, to the antibodies or fragments binding to identical or overlapping epitopes of hPD-L1. In one embodiment, competition is determined by ELISA, such techniques being readily apparent to the skilled person. In one embodiment, competition is determined by homogenous time resolved fluorescence (HTRF), such techniques being readily apparent to the skilled person. In one embodiment, competition is determined by fluorescence activated cell sorting (FACS), such techniques being readily apparent to the skilled person. In one embodiment, competition is determined by ForteBio Octet.RTM. Bio-Layer Interferometry (BLI) such techniques being readily apparent to the skilled person.
[0370] In one embodiment, the antibody or fragment competes (e.g. in a dose-dependent manner) with hPD-1 (or a fusion protein thereof) for binding to cell surface-expressed hPD-L1. In one embodiment, the antibody or fragment competes (e.g. in a dose-dependent manner) with hPD-1 (or a fusion protein thereof) for binding to soluble hPDL-1.
[0371] In one embodiment, the antibody or fragment partially or completely inhibits binding of PD-1 and/or CD80 to cell surface-expressed PD-L1, such as hPD-L1. In another embodiment, the antibody or fragment partially or completely inhibits binding of hPD-1 and/or CD80 to soluble hPD-L1. In some embodiments, the antibody or fragment partially or completely increases the secretion of IFN.gamma., CD25 and IL-2 from a cell having cell surface-expressed PD-1. In one embodiment, the antibody or fragment partially or completely inhibits binding of CD80 to soluble hPD-L1, but does not show any detectable inhibition of the binding of PD-1 to cell surface-expressed PD-L1. In one embodiment, the antibody or fragment partially or completely inhibits binding of CD80 to soluble hPD-L1, but does not show any detectable inhibition of the binding of PD-1 to soluble PD-L1.
[0372] As used herein, "inhibits", "inhibition", "inhibiting" and the like, as used herein refers to the ability of an antagonist (e.g. an antibody or fragment thereof) to bind to an epitope which either partially or completely prevents the binding of the receptor (e.g. CD80 or PD-1) to the ligand (e.g. PD-L1). If the epitope to which the antagonist binds completely blocks the binding site of the ligand, then ligand binding is completely prevented (which may be a physical blocking--in the case of overlapping epitopes--or steric blocking--where the antagonist is large such that it prevents the ligand binding to its distinct epitope), and the ligand is not removed from circulation. The concentration of circulating ligand may therefore appear to be increased. If the epitope to which the antagonist binds partially blocks the binding site of the ligand, the the ligand may be able to bind, but only weakly (in the case of partial inhibition), or in a different orientation to the natural binding interaction. In this case, some of the ligand may be removed from circulation, but not as much as when the ligand binding site is completely free and available for binding. Inhibition thus refers to the physical interaction of ligand and receptor. Inhibition can be measured by HTRF, which is described in more detail elsewhere herein and in Mathis (1995) Clinical Chemistry 41(9), 1391-1397. Inhibition can also be measured by flow cytometry, where receptor is expressed on cells, or by ELISA, where receptor is adsorbed onto plates.
[0373] Concept 16a. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 84G09.
[0374] Concept 16b. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 411B08.
[0375] Concept 16c. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 411C04.
[0376] Concept 16d. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 411D07.
[0377] Concept 16e. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 385F01.
[0378] Concept 16f. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 386H03.
[0379] Concept 16g. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 389A03.
[0380] Concept 16h. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 413D08.
[0381] Concept 16i. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 413G05.
[0382] Concept 16j. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 413F09.
[0383] Concept 16k. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 414B06.
[0384] Concept 16l. An antibody or fragment thereof which competes for binding to hPD-L1 with the antibody 416E01.
[0385] The antibodies have the sequences as described hereinabove.
Concept 17. The antibody or fragment according to any one of concepts 10 to 16, wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:29 or 32, or the CDRH3 sequence of SEQ ID NO:29 or 32 comprising 6 or fewer amino acid substitutions.
[0386] Concept 17a: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13a, and when dependent on concept 16, it is dependent on concept 16a), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:9 or 12, or the CDRH3 sequence of SEQ ID NO:9 or 12 comprising 6 or fewer amino acid substitutions.
[0387] Concept 17b: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13b, and when dependent on concept 16, it is dependent on concept 16b), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:54 or 57, or the CDRH3 sequence of SEQ ID NO:54 or 57 comprising 6 or fewer amino acid substitutions.
[0388] Concept 17c: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13c, and when dependent on concept 16, it is dependent on concept 16c), wherein the a V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:74 or 77, or the CDRH3 sequence of SEQ ID NO:74 or 77 comprising 6 or fewer amino acid substitutions.
[0389] Concept 17d: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13d, and when dependent on concept 16, it is dependent on concept 16d), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:94 or 97, or the CDRH3 sequence of SEQ ID NO:94 or 97 comprising 3 or fewer amino acid substitutions.
[0390] Concept 17e: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13e, and when dependent on concept 16, it is dependent on concept 16e), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:114 or 117, or the CDRH3 sequence of SEQ ID NO:114 or 117 comprising 6 or fewer amino acid substitutions.
[0391] Concept 17f: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13f, and when dependent on concept 16, it is dependent on concept 16f), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:144 or 147, or the CDRH3 sequence of SEQ ID NO:144 or 147 comprising 3 or fewer amino acid substitutions.
[0392] Concept 17g: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13g, and when dependent on concept 16, it is dependent on concept 16g), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:174 or 177, or the CDRH3 sequence of SEQ ID NO:174 or 177 comprising 6 or fewer amino acid substitutions.
[0393] Concept 17h: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13h, and when dependent on concept 16, it is dependent on concept 16h), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:134 or 137, or the CDRH3 sequence of SEQ ID NO:134 or 137 comprising 5 or fewer amino acid substitutions.
[0394] Concept 17i: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13i, and when dependent on concept 16, it is dependent on concept 16i), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:240 or 243, or the CDRH3 sequence of SEQ ID NO:240 or 243 comprising 6 or fewer amino acid substitutions.
[0395] Concept 17j: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13j, and when dependent on concept 16, it is dependent on concept 16j), wherein the a V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:260 or 263, or the CDRH3 sequence of SEQ ID NO:260 or 263 comprising 6 or fewer amino acid substitutions.
[0396] Concept 17k: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13k, and when dependent on concept 16, it is dependent on concept 16k), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:280 or 283, or the CDRH3 sequence of SEQ ID NO:280 or 283 comprising 6 or fewer amino acid substitutions.
[0397] Concept 17l: An antibody or a fragment thereof according to any one of concepts 10 to 16 (but when dependent on concept 13, it is dependent on concept 13l, and when dependent on concept 16, it is dependent on concept 16l), wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:345 or 348, or the CDRH3 sequence of SEQ ID NO:345 or 348 comprising 6 or fewer amino acid substitutions.
Concept 18. The antibody or fragment according to any preceding concept, wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:27 or 30 or the CDRH1 sequence of SEQ ID NO:27 or 30 comprising 3, 2 or 1 amino acid substitution(s).
[0398] Concept 18a: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9a, when dependent on concept 13, it is dependent on concept 13a, when dependent on concept 16, it is dependent on concept 16a, and when dependent on concept 17, it is dependent on concept 17a), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:7 or 10, or the CDRH1 sequence of SEQ ID NO:7 or 10 comprising 3, 2 or 1 amino acid substitution(s).
[0399] Concept 18b: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9b, when dependent on concept 13, it is dependent on concept 13b, when dependent on concept 16, it is dependent on concept 16b, and when dependent on concept 17, it is dependent on concept 17b), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:52 or 55, or the CDRH1 sequence of SEQ ID NO:52 or 55 comprising 3, 2 or 1 amino acid substitution(s).
[0400] Concept 18c: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9c, when dependent on concept 13, it is dependent on concept 13c, when dependent on concept 16, it is dependent on concept 16c, and when dependent on concept 17, it is dependent on concept 17c), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:72 or 75, or the CDRH1 sequence of SEQ ID NO:72 or 75 comprising 3, 2 or 1 amino acid substitution(s).
[0401] Concept 18d: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9d, when dependent on concept 13, it is dependent on concept 13d, when dependent on concept 16, it is dependent on concept 16d, and when dependent on concept 17, it is dependent on concept 17d), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:92 or 95, or the CDRH1 sequence of SEQ ID NO:92 or 95 comprising 3, 2 or 1 amino acid substitution(s).
[0402] Concept 18e: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9e, when dependent on concept 13, it is dependent on concept 13e, when dependent on concept 16, it is dependent on concept 16e, and when dependent on concept 17, it is dependent on concept 17e), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:112 or 115, or the CDRH1 sequence of SEQ ID NO:112 or 115 comprising 3, 2 or 1 amino acid substitution(s).
[0403] Concept 18f: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9f, when dependent on concept 13, it is dependent on concept 13f, when dependent on concept 16, it is dependent on concept 16f, and when dependent on concept 17, it is dependent on concept 17f), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:142 or 145, or the CDRH1 sequence of SEQ ID NO:142 or 145 comprising 3, 2 or 1 amino acid substitution(s).
[0404] Concept 18g: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9g, when dependent on concept 13, it is dependent on concept 13g, when dependent on concept 16, it is dependent on concept 16g, and when dependent on concept 17, it is dependent on concept 17g), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:172 or 175, or the CDRH1 sequence of SEQ ID NO:172 or 175 comprising 3, 2 or 1 amino acid substitution(s).
[0405] Concept 18h: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9h, when dependent on concept 13, it is dependent on concept 13h, when dependent on concept 16, it is dependent on concept 16h, and when dependent on concept 17, it is dependent on concept 17h), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:132 or 135, or the CDRH1 sequence of SEQ ID NO:132 or 135 comprising 3, 2 or 1 amino acid substitution(s).
[0406] Concept 18i: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9i, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16i, and when dependent on concept 17, it is dependent on concept 17i), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:238 or 241, or the CDRH1 sequence of SEQ ID NO:238 or 241 comprising 3, 2 or 1 amino acid substitution(s).
[0407] Concept 18j: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9j, when dependent on concept 13, it is dependent on concept 13j, when dependent on concept 16, it is dependent on concept 16j, and when dependent on concept 17, it is dependent on concept 17j), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:258 or 261, or the CDRH1 sequence of SEQ ID NO:258 or 261 comprising 3, 2 or 1 amino acid substitution(s).
[0408] Concept 18k: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9k, when dependent on concept 13, it is dependent on concept 13k, when dependent on concept 16, it is dependent on concept 16k, and when dependent on concept 17, it is dependent on concept 17k), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO: 278 or 281, or the CDRH1 sequence of SEQ ID NO: 278 or 281 comprising 3, 2 or 1 amino acid substitution(s).
[0409] Concept 18l: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9l, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16l, and when dependent on concept 17, it is dependent on concept 17l), wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO: 343 or 346, or the CDRH1 sequence of SEQ ID NO: 343 or 346 comprising 3, 2 or 1 amino acid substitution(s).
Concept 19. The antibody or fragment according to any preceding concept, wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:28 or 31, or the CDRH2 sequence of SEQ ID NO:28 or 31 comprising 4 or fewer amino acid substitutions.
[0410] Concept 19a: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9a, when dependent on concept 13, it is dependent on concept 13a, when dependent on concept 16, it is dependent on concept 16a, when dependent on concept 17, it is dependent on concept 17a, and when dependent on concept 18, it is dependent on concept 18a), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:8 or 11, or the CDRH2 sequence of SEQ ID NO:8 or 11 comprising 4 or fewer amino acid substitutions.
[0411] Concept 19b: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9b, when dependent on concept 13, it is dependent on concept 13b, when dependent on concept 16, it is dependent on concept 16b, when dependent on concept 17, it is dependent on concept 17b, and when dependent on concept 18, it is dependent on concept 18b), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:53 or 56, or the CDRH2 sequence of SEQ ID NO:53 or 56 comprising 3, 2 or 1 amino acid substitution(s).
[0412] Concept 19c: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9c, when dependent on concept 13, it is dependent on concept 13c, when dependent on concept 16, it is dependent on concept 16c, when dependent on concept 17, it is dependent on concept 17c, and when dependent on concept 18, it is dependent on concept 18c), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:73 or 76, or the CDRH2 sequence of SEQ ID NO:73 or 76 comprising 3, 2 or 1 amino acid substitution(s).
[0413] Concept 19d: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9d, when dependent on concept 13, it is dependent on concept 13d, when dependent on concept 16, it is dependent on concept 16d, when dependent on concept 17, it is dependent on concept 17d, and when dependent on concept 18, it is dependent on concept 18d), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:93 or 96, or the CDRH2 sequence of SEQ ID NO:93 or 96 comprising 3, 2 or 1 amino acid substitution(s).
[0414] Concept 19e: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9e, when dependent on concept 13, it is dependent on concept 13e, when dependent on concept 16, it is dependent on concept 16e, when dependent on concept 17, it is dependent on concept 17e, and when dependent on concept 18, it is dependent on concept 18e), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:113 or 116, or the CDRH2 sequence of SEQ ID NO:113 or 116 comprising 3, 2 or 1 amino acid substitution(s).
[0415] Concept 19f: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9f, when dependent on concept 13, it is dependent on concept 13f, when dependent on concept 16, it is dependent on concept 16f, when dependent on concept 17, it is dependent on concept 17f, and when dependent on concept 18, it is dependent on concept 18f), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:143 or 146, or the CDRH2 sequence of SEQ ID NO:143 or 146 comprising 3, 2 or 1 amino acid substitution(s).
[0416] Concept 19g: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9g, when dependent on concept 13, it is dependent on concept 13g, when dependent on concept 16, it is dependent on concept 16g, when dependent on concept 17, it is dependent on concept 17g, and when dependent on concept 18, it is dependent on concept 18g), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:173 or 176, or the CDRH2 sequence of SEQ ID NO:173 or 176 comprising 3, 2 or 1 amino acid substitution(s).
[0417] Concept 19h: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9h, when dependent on concept 13, it is dependent on concept 13h, when dependent on concept 16, it is dependent on concept 16h, when dependent on concept 17, it is dependent on concept 17h, and when dependent on concept 18, it is dependent on concept 18h), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:133 or 136, or the CDRH2 sequence of SEQ ID NO:133 or 136 comprising 3, 2 or 1 amino acid substitution(s).
[0418] Concept 19i: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9i, when dependent on concept 13, it is dependent on concept 13i, when dependent on concept 16, it is dependent on concept 16i, when dependent on concept 17, it is dependent on concept 17i, and when dependent on concept 18, it is dependent on concept 18i), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:239 or 242, or the CDRH2 sequence of SEQ ID NO:239 or 242 comprising 3, 2 or 1 amino acid substitution(s).
[0419] Concept 19j: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9j, when dependent on concept 13, it is dependent on concept 13j, when dependent on concept 16, it is dependent on concept 16j, when dependent on concept 17, it is dependent on concept 17j, and when dependent on concept 18, it is dependent on concept 18j), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:259 or 262, or the CDRH2 sequence of SEQ ID NO:259 or 262 comprising 3, 2 or 1 amino acid substitution(s).
[0420] Concept 19k: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9k, when dependent on concept 13, it is dependent on concept 13k, when dependent on concept 16, it is dependent on concept 16k, when dependent on concept 17, it is dependent on concept 17k, and when dependent on concept 18, it is dependent on concept 18k), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:279 or 282, or the CDRH2 sequence of SEQ ID NO:279 or 282 comprising 3, 2 or 1 amino acid substitution(s).
[0421] Concept 19l: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9l, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16l, when dependent on concept 17, it is dependent on concept 17l, and when dependent on concept 18, it is dependent on concept 18l), wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:344 or 347, or the CDRH2 sequence of SEQ ID NO:344 or 347 comprising 3, 2 or 1 amino acid substitution(s).
Concept 20. The antibody or fragment according to any preceding concept, wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:33, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:33.
[0422] Concept 20a: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9a, when dependent on concept 13, it is dependent on concept 13a, when dependent on concept 16, it is dependent on concept 16a, when dependent on concept 17, it is dependent on concept 17a, when dependent on concept 18, it is dependent on concept 18a, and when dependent on concept 19, it is dependent on concept 19a), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:13, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:13.
[0423] Concept 20b: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9b, when dependent on concept 13, it is dependent on concept 13b, when dependent on concept 16, it is dependent on concept 16b, when dependent on concept 17, it is dependent on concept 17b, when dependent on concept 18, it is dependent on concept 18b, and when dependent on concept 19, it is dependent on concept 19b), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:58, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:58.
[0424] Concept 20c: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9c, when dependent on concept 13, it is dependent on concept 13c, when dependent on concept 16, it is dependent on concept 16c, when dependent on concept 17, it is dependent on concept 17c, when dependent on concept 18, it is dependent on concept 18c, and when dependent on concept 19, it is dependent on concept 19c), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:78, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:78.
[0425] Concept 20d: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9d, when dependent on concept 13, it is dependent on concept 13d, when dependent on concept 16, it is dependent on concept 16d, when dependent on concept 17, it is dependent on concept 17d, when dependent on concept 18, it is dependent on concept 18d, and when dependent on concept 19, it is dependent on concept 19d), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:98, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:98.
[0426] Concept 20e: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9e, when dependent on concept 13, it is dependent on concept 13e, when dependent on concept 16, it is dependent on concept 16e, when dependent on concept 17, it is dependent on concept 17e, when dependent on concept 18, it is dependent on concept 18e, and when dependent on concept 19, it is dependent on concept 19e), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:118, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:118.
[0427] Concept 20f: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9f, when dependent on concept 13, it is dependent on concept 13f, when dependent on concept 16, it is dependent on concept 16f, when dependent on concept 17, it is dependent on concept 17f, when dependent on concept 18, it is dependent on concept 18f, and when dependent on concept 19, it is dependent on concept 19f), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:158, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:158.
[0428] Concept 20g: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9g, when dependent on concept 13, it is dependent on concept 13g, when dependent on concept 16, it is dependent on concept 16g, when dependent on concept 17, it is dependent on concept 17g, when dependent on concept 18, it is dependent on concept 18g, and when dependent on concept 19, it is dependent on concept 19g), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:178, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:178.
[0429] Concept 20h: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9h, when dependent on concept 13, it is dependent on concept 13h, when dependent on concept 16, it is dependent on concept 16h, when dependent on concept 17, it is dependent on concept 17h, when dependent on concept 18, it is dependent on concept 18h, and when dependent on concept 19, it is dependent on concept 19h), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:138, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:138.
[0430] Concept 20i: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9i, when dependent on concept 13, it is dependent on concept 13i, when dependent on concept 16, it is dependent on concept 16i, when dependent on concept 17, it is dependent on concept 17i, when dependent on concept 18, it is dependent on concept 18i, and when dependent on concept 19, it is dependent on concept 19i), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:244, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:244.
[0431] Concept 20j: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9j, when dependent on concept 13, it is dependent on concept 13j, when dependent on concept 16, it is dependent on concept 16j, when dependent on concept 17, it is dependent on concept 17j, when dependent on concept 18, it is dependent on concept 18j, and when dependent on concept 19, it is dependent on concept 19j), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:264, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:264.
[0432] Concept 20k: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9k, when dependent on concept 13, it is dependent on concept 13k, when dependent on concept 16, it is dependent on concept 16k, when dependent on concept 17, it is dependent on concept 17k, when dependent on concept 18, it is dependent on concept 18k, and when dependent on concept 19, it is dependent on concept 19k), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:284, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:284.
[0433] Concept 20l: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9l, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16l, when dependent on concept 17, it is dependent on concept 17l, when dependent on concept 18, it is dependent on concept 18l, and when dependent on concept 19, it is dependent on concept 19l), wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:349, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:349.
[0434] In one embodiment, the amino acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99.5% identical to the specified Seq ID No.
Concept 21. The antibody or fragment according to any preceding concept comprising first and second copies of said V.sub.H domain. Concept 22. The antibody or fragment according to any preceding concept, comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:37 or 40, or the CRDL1 sequence of SEQ ID NO:37 or 40 comprising 3 or fewer amino acid substitutions.
[0435] Concept 22a: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9a, when dependent on concept 13, it is dependent on concept 13a, when dependent on concept 16, it is dependent on concept 16a, when dependent on concept 17, it is dependent on concept 17a, when dependent on concept 18, it is dependent on concept 18a, when dependent on concept 19, it is dependent on concept 19a, and when dependent on concept 20, it is dependent on concept 20a), comprising a V.sub.L domain, which comprises the CDRL1 sequence of SEQ ID NO:17 or 20, or the CDRL1 sequence of SEQ ID NO:17 or 20 comprising 3 or fewer amino acid substitutions.
[0436] Concept 22b: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9b, when dependent on concept 13, it is dependent on concept 13b, when dependent on concept 16, it is dependent on concept 16b, when dependent on concept 17, it is dependent on concept 17b, when dependent on concept 18, it is dependent on concept 18b, when dependent on concept 19, it is dependent on concept 19b, and when dependent on concept 20, it is dependent on concept 20b), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:62 or 65, or the CDRL1 sequence of SEQ ID NO:62 or 65 comprising 3 or fewer amino acid substitutions.
[0437] Concept 22c: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9c, when dependent on concept 13, it is dependent on concept 13c, when dependent on concept 16, it is dependent on concept 16c, when dependent on concept 17, it is dependent on concept 17c, when dependent on concept 18, it is dependent on concept 18c, when dependent on concept 19, it is dependent on concept 19c, and when dependent on concept 20, it is dependent on concept 20c), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:82 or 85, or the CDRL1 sequence of SEQ ID NO:82 or 85 comprising 2 or 1 amino acid substitution(s).
[0438] Concept 22d: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9d, when dependent on concept 13, it is dependent on concept 13d, when dependent on concept 16, it is dependent on concept 16d, when dependent on concept 17, it is dependent on concept 17d, when dependent on concept 18, it is dependent on concept 18d, when dependent on concept 19, it is dependent on concept 19d, and when dependent on concept 20, it is dependent on concept 20d), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:102 or 105, or the CDRL1 sequence of SEQ ID NO:102 or 105 comprising 5 or fewer amino acid substitutions.
[0439] Concept 22e: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9e, when dependent on concept 13, it is dependent on concept 13e, when dependent on concept 16, it is dependent on concept 16e, when dependent on concept 17, it is dependent on concept 17e, when dependent on concept 18, it is dependent on concept 18e, when dependent on concept 19, it is dependent on concept 19e, and when dependent on concept 20, it is dependent on concept 20e), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:122 or 125, or the CDRL1 sequence of SEQ ID NO:122 or 125 comprising 2 or 1 amino acid substitution(s).
[0440] Concept 22f: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9f, when dependent on concept 13, it is dependent on concept 13f, when dependent on concept 16, it is dependent on concept 16f, when dependent on concept 17, it is dependent on concept 17f, when dependent on concept 18, it is dependent on concept 18f, when dependent on concept 19, it is dependent on concept 19f, and when dependent on concept 20, it is dependent on concept 20f), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:162 or 165, or the CDRL1 sequence of SEQ ID NO:162 or 165 comprising 5 or fewer amino acid substitutions.
[0441] Concept 22g: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9g, when dependent on concept 13, it is dependent on concept 13g, when dependent on concept 16, it is dependent on concept 16g, when dependent on concept 17, it is dependent on concept 17g, when dependent on concept 18, it is dependent on concept 18g, when dependent on concept 19, it is dependent on concept 19g, and when dependent on concept 20, it is dependent on concept 20g), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:182 or 185, or the CDRL1 sequence of SEQ ID NO:182 or 185 comprising 5 or fewer amino acid substitutions.
[0442] Concept 22h: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9h, when dependent on concept 13, it is dependent on concept 13h, when dependent on concept 16, it is dependent on concept 16h, when dependent on concept 17, it is dependent on concept 17h, when dependent on concept 18, it is dependent on concept 18h, when dependent on concept 19, it is dependent on concept 19h, and when dependent on concept 20, it is dependent on concept 20h), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:142 or 145, or the CDRL1 sequence of SEQ ID NO:142 or 145 comprising 2 or 1 amino acid substitution(s).
[0443] Concept 22i: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9i, when dependent on concept 13, it is dependent on concept 13i, when dependent on concept 16, it is dependent on concept 16i, when dependent on concept 17, it is dependent on concept 17i, when dependent on concept 18, it is dependent on concept 18i, when dependent on concept 19, it is dependent on concept 19i, and when dependent on concept 20, it is dependent on concept 20i), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:248 or 251, or the CDRL1 sequence of SEQ ID NO:248 or 251 comprising 2 or 1 amino acid substitution(s).
[0444] Concept 22j: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9j, when dependent on concept 13, it is dependent on concept 13j, when dependent on concept 16, it is dependent on concept 16j, when dependent on concept 17, it is dependent on concept 17j, when dependent on concept 18, it is dependent on concept 18j, when dependent on concept 19, it is dependent on concept 19j, and when dependent on concept 20, it is dependent on concept 20j), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:268 or 271, or the CDRL1 sequence of SEQ ID NO:268 or 271 comprising 2 or 1 amino acid substitution(s).
[0445] Concept 22k: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9k, when dependent on concept 13, it is dependent on concept 13k, when dependent on concept 16, it is dependent on concept 16k, when dependent on concept 17, it is dependent on concept 17k, when dependent on concept 18, it is dependent on concept 18k, when dependent on concept 19, it is dependent on concept 19k, and when dependent on concept 20, it is dependent on concept 20k), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:288 or 291, or the CDRL1 sequence of SEQ ID NO:288 or 291 comprising 2 or 1 amino acid substitution(s).
[0446] Concept 22l: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9l, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16l, when dependent on concept 17, it is dependent on concept 17l, when dependent on concept 18, it is dependent on concept 18l, when dependent on concept 19, it is dependent on concept 19l, and when dependent on concept 20, it is dependent on concept 20l), comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:353 or 356, or the CDRL1 sequence of SEQ ID NO:353 or 356 comprising 2 or 1 amino acid substitution(s).
Concept 23. The antibody or fragment according to any preceding concept, comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:38 or 41, or the CRDL2 sequence of SEQ ID NO:38 or 41 comprising 2 or 1 amino acid substitution(s), for example a CDRL2 sequence of Seq ID No:50.
[0447] Concept 23a: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9a, when dependent on concept 13, it is dependent on concept 13a, when dependent on concept 16, it is dependent on concept 16a, when dependent on concept 17, it is dependent on concept 17a, when dependent on concept 18, it is dependent on concept 18a, when dependent on concept 19, it is dependent on concept 19a, when dependent on concept 20, it is dependent on concept 20a, and when dependent on concept 22, it is dependent on concept 22a), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:18 or 21, or the CDRL2 sequence of SEQ ID NO:18 or 21 comprising 2 or 1 amino acid substitution(s).
[0448] Concept 23b: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9b, when dependent on concept 13, it is dependent on concept 13b, when dependent on concept 16, it is dependent on concept 16b, when dependent on concept 17, it is dependent on concept 17b, when dependent on concept 18, it is dependent on concept 18b, when dependent on concept 19, it is dependent on concept 19b, when dependent on concept 20, it is dependent on concept 20b, and when dependent on concept 22, it is dependent on concept 22b), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:63 or 66, or the CDRL2 sequence of SEQ ID NO:63 or 66 comprising one amino acid substitution.
[0449] Concept 23c: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9c, when dependent on concept 13, it is dependent on concept 13c, when dependent on concept 16, it is dependent on concept 16c, when dependent on concept 17, it is dependent on concept 17c, when dependent on concept 18, it is dependent on concept 18c, when dependent on concept 19, it is dependent on concept 19c, when dependent on concept 20, it is dependent on concept 20c, and when dependent on concept 22, it is dependent on concept 22c), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:83 or 86, or the CDRL2 sequence of SEQ ID NO:83 or 86 comprising one amino acid substitution.
[0450] Concept 23d: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9d, when dependent on concept 13, it is dependent on concept 13d, when dependent on concept 16, it is dependent on concept 16d, when dependent on concept 17, it is dependent on concept 17d, when dependent on concept 18, it is dependent on concept 18d, when dependent on concept 19, it is dependent on concept 19d, when dependent on concept 20, it is dependent on concept 20d, and when dependent on concept 22, it is dependent on concept 22d), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:103 or 106, or the CDRL2 sequence of SEQ ID NO:103 or 106 comprising one amino acid substitution.
[0451] Concept 23e: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9e, when dependent on concept 13, it is dependent on concept 13e, when dependent on concept 16, it is dependent on concept 16e, when dependent on concept 17, it is dependent on concept 17e, when dependent on concept 18, it is dependent on concept 18e, when dependent on concept 19, it is dependent on concept 19e, when dependent on concept 20, it is dependent on concept 20e, and when dependent on concept 22, it is dependent on concept 22e), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:123 or 126, or the CDRL2 sequence of SEQ ID NO:123 or 126 comprising one amino acid substitution.
[0452] Concept 23f: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9f, when dependent on concept 13, it is dependent on concept 13f, when dependent on concept 16, it is dependent on concept 16f, when dependent on concept 17, it is dependent on concept 17f, when dependent on concept 18, it is dependent on concept 18f, when dependent on concept 19, it is dependent on concept 19f, when dependent on concept 20, it is dependent on concept 20f, and when dependent on concept 22, it is dependent on concept 220, comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:153 or 156, or the CDRL2 sequence of SEQ ID NO:153 or 156 comprising one amino acid substitution.
[0453] Concept 23g: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9g, when dependent on concept 13, it is dependent on concept 13g, when dependent on concept 16, it is dependent on concept 16g, when dependent on concept 17, it is dependent on concept 17g, when dependent on concept 18, it is dependent on concept 18g, when dependent on concept 19, it is dependent on concept 19g, when dependent on concept 20, it is dependent on concept 20g, and when dependent on concept 22, it is dependent on concept 22g), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:183 or 186, or the CDRL2 sequence of SEQ ID NO:183 or 186 comprising one amino acid substitution.
[0454] Concept 23h: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9h, when dependent on concept 13, it is dependent on concept 13h, when dependent on concept 16, it is dependent on concept 16h, when dependent on concept 17, it is dependent on concept 17h, when dependent on concept 18, it is dependent on concept 18h, when dependent on concept 19, it is dependent on concept 19h, when dependent on concept 20, it is dependent on concept 20h, and when dependent on concept 22, it is dependent on concept 22h), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:143 or 146, or the CDRL2 sequence of SEQ ID NO:143 or 146 comprising one amino acid substitution.
[0455] Concept 23i: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9i, when dependent on concept 13, it is dependent on concept 13i, when dependent on concept 16, it is dependent on concept 16i, when dependent on concept 17, it is dependent on concept 17i, when dependent on concept 18, it is dependent on concept 18i, when dependent on concept 19, it is dependent on concept 19i, when dependent on concept 20, it is dependent on concept 20i, and when dependent on concept 22, it is dependent on concept 22i), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:249 or 252, or the CDRL2 sequence of SEQ ID NO:249 or 252 comprising one amino acid substitution.
[0456] Concept 23j: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9j, when dependent on concept 13, it is dependent on concept 13j, when dependent on concept 16, it is dependent on concept 16j, when dependent on concept 17, it is dependent on concept 17j, when dependent on concept 18, it is dependent on concept 18j, when dependent on concept 19, it is dependent on concept 19j, when dependent on concept 20, it is dependent on concept 20j, and when dependent on concept 22, it is dependent on concept 22j), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:269 or 272, or the CDRL2 sequence of SEQ ID NO:269 or 272 comprising one amino acid substitution.
[0457] Concept 23k: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9k, when dependent on concept 13, it is dependent on concept 13k, when dependent on concept 16, it is dependent on concept 16k, when dependent on concept 17, it is dependent on concept 17k, when dependent on concept 18, it is dependent on concept 18k, when dependent on concept 19, it is dependent on concept 19k, when dependent on concept 20, it is dependent on concept 20k, and when dependent on concept 22, it is dependent on concept 22k), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:289 or 292, or the CDRL2 sequence of SEQ ID NO:289 or 292 comprising one amino acid substitution.
[0458] Concept 23l: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9l, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16l, when dependent on concept 17, it is dependent on concept 17l, when dependent on concept 18, it is dependent on concept 18l, when dependent on concept 19, it is dependent on concept 19l, when dependent on concept 20, it is dependent on concept 20l, and when dependent on concept 22, it is dependent on concept 22l), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of SEQ ID NO:354 or 357, or the CDRL2 sequence of SEQ ID NO:354 or 357 comprising one amino acid substitution.
Concept 24. The antibody or fragment according to any preceding concept, comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:39 or 42, or the CRDL3 sequence of SEQ ID NO:39 or 42 comprising 4 or fewer amino acid substitutions.
[0459] Concept 24a: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9a, when dependent on concept 13, it is dependent on concept 13a, when dependent on concept 16, it is dependent on concept 16a, when dependent on concept 17, it is dependent on concept 17a, when dependent on concept 18, it is dependent on concept 18a, when dependent on concept 19, it is dependent on concept 19a, when dependent on concept 20, it is dependent on concept 20a, when dependent on concept 22, it is dependent on concept 22a, and when dependent on concept 23, it is dependent on concept 23a), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:19 or 22, or the CDRL3 sequence of SEQ ID NO: 19 or 22 comprising 4 or fewer amino acid substitutions.
[0460] Concept 24b: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9b, when dependent on concept 13, it is dependent on concept 13b, when dependent on concept 16, it is dependent on concept 16b, when dependent on concept 17, it is dependent on concept 17b, when dependent on concept 18, it is dependent on concept 18b, when dependent on concept 19, it is dependent on concept 19b, when dependent on concept 20, it is dependent on concept 20b, when dependent on concept 22, it is dependent on concept 22b, and when dependent on concept 23, it is dependent on concept 23b), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:64 or 67, or the CDRL3 sequence of SEQ ID NO:64 or 67 comprising 4 or fewer amino acid substitutions.
[0461] Concept 24c: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9c, when dependent on concept 13, it is dependent on concept 13c, when dependent on concept 16, it is dependent on concept 16c, when dependent on concept 17, it is dependent on concept 17c, when dependent on concept 18, it is dependent on concept 18c, when dependent on concept 19, it is dependent on concept 19c, when dependent on concept 20, it is dependent on concept 20c, when dependent on concept 22, it is dependent on concept 22c, and when dependent on concept 23, it is dependent on concept 23c), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:84 or 87, or the CDRL3 sequence of SEQ ID NO:84 or 87 comprising 4 or fewer amino acid substitutions.
[0462] Concept 24d: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9d, when dependent on concept 13, it is dependent on concept 13d, when dependent on concept 16, it is dependent on concept 16d, when dependent on concept 17, it is dependent on concept 17d, when dependent on concept 18, it is dependent on concept 18d, when dependent on concept 19, it is dependent on concept 19d, when dependent on concept 20, it is dependent on concept 20d, when dependent on concept 22, it is dependent on concept 22d, and when dependent on concept 23, it is dependent on concept 23d), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:104 or 107, or the CDRL3 sequence of SEQ ID NO:104 or 107 comprising 4 or fewer amino acid substitutions.
[0463] Concept 24e: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9e, when dependent on concept 13, it is dependent on concept 13e, when dependent on concept 16, it is dependent on concept 16e, when dependent on concept 17, it is dependent on concept 17e, when dependent on concept 18, it is dependent on concept 18e, when dependent on concept 19, it is dependent on concept 19e, when dependent on concept 20, it is dependent on concept 20e, when dependent on concept 22, it is dependent on concept 22e, and when dependent on concept 23, it is dependent on concept 23e), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:124 or 127, or the CDRL3, sequence of SEQ ID NO:124 or 127 comprising 4 or fewer amino acid substitutions.
[0464] Concept 24f: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9f, when dependent on concept 13, it is dependent on concept 13f, when dependent on concept 16, it is dependent on concept 16f, when dependent on concept 17, it is dependent on concept 17f, when dependent on concept 18, it is dependent on concept 18f, when dependent on concept 19, it is dependent on concept 19f, when dependent on concept 20, it is dependent on concept 20f, when dependent on concept 22, it is dependent on concept 22f, and when dependent on concept 23, it is dependent on concept 23f), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:164 or 167, or the CDRL3 sequence of SEQ ID NO:164 or 167 comprising 4 or fewer amino acid substitutions.
[0465] Concept 24g: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9g, when dependent on concept 13, it is dependent on concept 13g, when dependent on concept 16, it is dependent on concept 16g, when dependent on concept 17, it is dependent on concept 17g, when dependent on concept 18, it is dependent on concept 18g, when dependent on concept 19, it is dependent on concept 19g, when dependent on concept 20, it is dependent on concept 20g, when dependent on concept 22, it is dependent on concept 22g, and when dependent on concept 23, it is dependent on concept 23g), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:184 or 187, or the CDRL3 sequence of SEQ ID NO:184 or 187 comprising 4 or fewer amino acid substitutions.
[0466] Concept 24h: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9h, when dependent on concept 13, it is dependent on concept 13h, when dependent on concept 16, it is dependent on concept 16h, when dependent on concept 17, it is dependent on concept 17h, when dependent on concept 18, it is dependent on concept 18h, when dependent on concept 19, it is dependent on concept 19h, when dependent on concept 20, it is dependent on concept 20h, when dependent on concept 22, it is dependent on concept 22h, and when dependent on concept 23, it is dependent on concept 23h), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:144 or 147, or the CDRL3 sequence of SEQ ID NO:144 or 147 comprising 4 or fewer amino acid substitutions.
[0467] Concept 24i: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9i, when dependent on concept 13, it is dependent on concept 13i, when dependent on concept 16, it is dependent on concept 16i, when dependent on concept 17, it is dependent on concept 17i, when dependent on concept 18, it is dependent on concept 18i, when dependent on concept 19, it is dependent on concept 19i, when dependent on concept 20, it is dependent on concept 20i, when dependent on concept 22, it is dependent on concept 22i, and when dependent on concept 23, it is dependent on concept 23i), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:250 or 253, or the CDRL3 sequence of SEQ ID NO:250 or 253 comprising 4 or fewer amino acid substitutions.
[0468] Concept 24j: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9j, when dependent on concept 13, it is dependent on concept 13j, when dependent on concept 16, it is dependent on concept 16j, when dependent on concept 17, it is dependent on concept 17j, when dependent on concept 18, it is dependent on concept 18j, when dependent on concept 19, it is dependent on concept 19j, when dependent on concept 20, it is dependent on concept 20j, when dependent on concept 22, it is dependent on concept 22j, and when dependent on concept 23, it is dependent on concept 23j), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:270 or 273, or the CDRL3 sequence of SEQ ID NO:270 or 273 comprising 4 or fewer amino acid substitutions.
[0469] Concept 24k: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9k, when dependent on concept 13, it is dependent on concept 13k, when dependent on concept 16, it is dependent on concept 16k, when dependent on concept 17, it is dependent on concept 17k, when dependent on concept 18, it is dependent on concept 18k, when dependent on concept 19, it is dependent on concept 19k, when dependent on concept 20, it is dependent on concept 20k, when dependent on concept 22, it is dependent on concept 22k, and when dependent on concept 23, it is dependent on concept 23k), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:290 or 293, or the CDRL3 sequence of SEQ ID NO:290 or 293 comprising 4 or fewer amino acid substitutions.
[0470] Concept 24l: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9l, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16l, when dependent on concept 17, it is dependent on concept 17l, when dependent on concept 18, it is dependent on concept 18l, when dependent on concept 19, it is dependent on concept 19l, when dependent on concept 20, it is dependent on concept 20l, when dependent on concept 22, it is dependent on concept 22l, and when dependent on concept 23, it is dependent on concept 23l), comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of SEQ ID NO:355 or 358, or the CDRL3 sequence of SEQ ID NO:355 or 358 comprising 4 or fewer amino acid substitutions.
Concept 25. The antibody or fragment according to any preceding concept, comprising a or said V.sub.L domain, which V.sub.L domain comprises an amino acid sequence of SEQ ID NO:43, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:43 (for example the V.sub.L domain sequence in the light chain sequence of Seq ID No:50, 51 or 298).
[0471] Concept 25a: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9a, when dependent on concept 13, it is dependent on concept 13a, when dependent on concept 16, it is dependent on concept 16a, when dependent on concept 17, it is dependent on concept 17a, when dependent on concept 18, it is dependent on concept 18a, when dependent on concept 19, it is dependent on concept 19a, when dependent on concept 20, it is dependent on concept 20a, when dependent on concept 22, it is dependent on concept 22a, when dependent on concept 23, it is dependent on concept 23a, and when dependent on concept 24, it is dependent on concept 24a), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:23, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:23.
[0472] Concept 25b: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9b, when dependent on concept 13, it is dependent on concept 13b, when dependent on concept 16, it is dependent on concept 16b, when dependent on concept 17, it is dependent on concept 17b, when dependent on concept 18, it is dependent on concept 18b, when dependent on concept 19, it is dependent on concept 19b, when dependent on concept 20, it is dependent on concept 20b, when dependent on concept 22, it is dependent on concept 22a, when dependent on concept 23, it is dependent on concept 23b, and when dependent on concept 24, it is dependent on concept 24b), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:68, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:68.
[0473] Concept 25c: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9c, when dependent on concept 13, it is dependent on concept 13c, when dependent on concept 16, it is dependent on concept 16c, when dependent on concept 17, it is dependent on concept 17c, when dependent on concept 18, it is dependent on concept 18c, when dependent on concept 19, it is dependent on concept 19c, when dependent on concept 20, it is dependent on concept 20c, when dependent on concept 22, it is dependent on concept 22c, when dependent on concept 23, it is dependent on concept 23c, and when dependent on concept 24, it is dependent on concept 24c), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:88, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:88.
[0474] Concept 25d: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9d, when dependent on concept 13, it is dependent on concept 13d, when dependent on concept 16, it is dependent on concept 16d, when dependent on concept 17, it is dependent on concept 17d, when dependent on concept 18, it is dependent on concept 18d, when dependent on concept 19, it is dependent on concept 19d, when dependent on concept 20, it is dependent on concept 20d, when dependent on concept 22, it is dependent on concept 22d, when dependent on concept 23, it is dependent on concept 23d, and when dependent on concept 24, it is dependent on concept 24d), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:108, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:108.
[0475] Concept 25e: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9e, when dependent on concept 13, it is dependent on concept 13e, when dependent on concept 16, it is dependent on concept 16e, when dependent on concept 17, it is dependent on concept 17e, when dependent on concept 18, it is dependent on concept 18e, when dependent on concept 19, it is dependent on concept 19e, when dependent on concept 20, it is dependent on concept 20e, when dependent on concept 22, it is dependent on concept 22e, when dependent on concept 23, it is dependent on concept 23e, and when dependent on concept 24, it is dependent on concept 24e), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:128, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:128.
[0476] Concept 25f: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9f, when dependent on concept 13, it is dependent on concept 13f, when dependent on concept 16, it is dependent on concept 16f, when dependent on concept 17, it is dependent on concept 17f, when dependent on concept 18, it is dependent on concept 18f, when dependent on concept 19, it is dependent on concept 19f, when dependent on concept 20, it is dependent on concept 20f, when dependent on concept 22, it is dependent on concept 22f, when dependent on concept 23, it is dependent on concept 23f, and when dependent on concept 24, it is dependent on concept 240, wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:168, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:168.
[0477] Concept 25g: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9g, when dependent on concept 13, it is dependent on concept 13g, when dependent on concept 16, it is dependent on concept 16g, when dependent on concept 17, it is dependent on concept 17g, when dependent on concept 18, it is dependent on concept 18g, when dependent on concept 19, it is dependent on concept 19g, when dependent on concept 20, it is dependent on concept 20g, when dependent on concept 22, it is dependent on concept 22g, when dependent on concept 23, it is dependent on concept 23g, and when dependent on concept 24, it is dependent on concept 24g), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:188, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:188.
[0478] Concept 25h: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9h, when dependent on concept 13, it is dependent on concept 13h, when dependent on concept 16, it is dependent on concept 16h, when dependent on concept 17, it is dependent on concept 17h, when dependent on concept 18, it is dependent on concept 18h, when dependent on concept 19, it is dependent on concept 19h, when dependent on concept 20, it is dependent on concept 20h, when dependent on concept 22, it is dependent on concept 22h, when dependent on concept 23, it is dependent on concept 23h, and when dependent on concept 24, it is dependent on concept 24h), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:148, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:148.
[0479] Concept 25i: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9i, when dependent on concept 13, it is dependent on concept 13i, when dependent on concept 16, it is dependent on concept 16i, when dependent on concept 17, it is dependent on concept 17i, when dependent on concept 18, it is dependent on concept 18i, when dependent on concept 19, it is dependent on concept 19i, when dependent on concept 20, it is dependent on concept 20i, when dependent on concept 22, it is dependent on concept 22i, when dependent on concept 23, it is dependent on concept 23i, and when dependent on concept 24, it is dependent on concept 24i), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:254, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:254.
[0480] Concept 25j: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9j, when dependent on concept 13, it is dependent on concept 13j, when dependent on concept 16, it is dependent on concept 16j, when dependent on concept 17, it is dependent on concept 17j, when dependent on concept 18, it is dependent on concept 18j, when dependent on concept 19, it is dependent on concept 19j, when dependent on concept 20, it is dependent on concept 20j, when dependent on concept 22, it is dependent on concept 22j, when dependent on concept 23, it is dependent on concept 23j, and when dependent on concept 24, it is dependent on concept 24j), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:274, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:274.
[0481] Concept 25k: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9k, when dependent on concept 13, it is dependent on concept 13k, when dependent on concept 16, it is dependent on concept 16k, when dependent on concept 17, it is dependent on concept 17k, when dependent on concept 18, it is dependent on concept 18k, when dependent on concept 19, it is dependent on concept 19k, when dependent on concept 20, it is dependent on concept 20k, when dependent on concept 22, it is dependent on concept 22k, when dependent on concept 23, it is dependent on concept 23k, and when dependent on concept 24, it is dependent on concept 24k), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:294, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:294.
[0482] Concept 25l: An antibody or a fragment thereof according to any preceding concept (but when dependent on concept 9, it is dependent on concept 9l, when dependent on concept 13, it is dependent on concept 13l, when dependent on concept 16, it is dependent on concept 16l, when dependent on concept 17, it is dependent on concept 17l, when dependent on concept 18, it is dependent on concept. 18l, when dependent on concept 19, it is dependent on concept 19l, when dependent on concept 20, it is dependent on concept 20l, when dependent on concept 22, it is dependent on concept 22l, when dependent on concept 23, it is dependent on concept 23l, and when dependent on concept 24, it is dependent on concept 24l), wherein the V.sub.L domain comprises an amino acid sequence of SEQ ID NO:359, or a light chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:359.
[0483] In one embodiment, the amino acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99.5% identical to the specified Seq ID No.
Concept 26. The antibody or fragment according to any one of concepts 12 to 21, comprising first and second copies of a or said V.sub.L domain. Concept 27. The antibody or fragment according to any preceding concept which specifically binds to cynomolgus PD-L1 as defined by Seq ID No:2.
[0484] In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of less than 1 nM (e.g. from 1 nM to 0.01 pM or from 1 nM to 0.1 pM, or from 1 nM to 1 pM). In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of less than 10 nM (e.g. from 10 nM to 0.01 pM or from 10 nM to 0.1 pM, or from 10 nM to 1 pM). In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of less than 0.1 nM (e.g. from 0.1 nM to 0.01 pM or from 0.1 nM to 0.1 pM, or from 0.1 nM to 1 pM). In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of less than 0.01 nM (e.g. from 0.011 nM to 0.01 pM or from 0.01 nM to 0.1 pM).
[0485] In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of within 2-fold of the affinity to hPD-L1. In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of within 4-fold of the affinity to hPD-L1. In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of within 5-fold of the affinity to hPD-L1. In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of within 6-fold of the affinity to hPD-L1. In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of within 8-fold of the affinity to hPD-L1. In one embodiment, the antibody or fragment binds to cynomolgus PD-L1 with an affinity of within 10-fold of the affinity to hPD-L1.
[0486] In one embodiment, the antibody or fragment does not detectably bind to cynomolgus PD-L1. In one embodiment, the antibody or fragment does not detectably bind to murine PD-L1.
[0487] In one embodiment, the antibody or fragment binds to murine PD-L1 with an affinity of less than 1 nM (e.g. from 1 nM to 0.01 pM or from 1 nM to 0.1 pM, or from 1 nM to 1 pM). In one embodiment, the antibody or fragment binds to murine PD-L1 with an affinity of less than 10 nM (e.g. from 10 nM to 0.01 pM or from 10 nM to 0.1 pM, or from 10 nM to 1 pM). In one embodiment, the antibody or fragment binds to murine PD-L1 with an affinity of less than 0.1 nM (e.g. from 0.1 nM to 0.01 pM or from 0.1 nM to 0.1 pM, or from 0.1 nM to 1 pM). In one embodiment, the antibody or fragment binds to murine PD-L1 with an affinity of less than 0.01 nM (e.g. from 0.011 nM to 0.01 pM or from 0.01 nM to 0.1 pM).
Concept 28. The antibody or fragment according to any preceding concept, wherein the antibody or fragment comprises a kappa light chain.
[0488] Kappa light chain constant region amino acid and nucleotide sequences can be found in Seq ID Nos:206 to 215.
[0489] In one embodiment, the light chain may be a lambda light chain. Lambda light chain constant region amino acid and nucleotide sequences can be found in Seq ID Nos:216 to 237 and Seq ID No:535, Seq ID No:536 and Seq ID No:538.
Concept 29. The antibody or fragment according to any one of concepts 9 to 28, wherein the amino acid substitutions are conservative amino acid substitutions, optionally wherein the conservative substitutions are from one of six groups (each group containing amino acids that are conservative substitutions for one another) selected from: [0490] 1) Alanine (A), Serine (S), Threonine (T); [0491] 2) Aspartic acid (D), Glutamic acid (E); [0492] 3) Asparagine (N), Glutamine (Q); [0493] 4) Arginine (R), Lysine (K); [0494] 5) Isoleucine (I), Leucine (L), Methionine (M), Valine (V); and [0495] 6) Phenylalanine (F), Tyrosine (Y), Tryptophan (W).
[0496] Conservative substitutions may be as described above in concept 9.
Concept 30. The antibody or fragment according to any preceding concept, wherein the antibody or fragment comprises a constant region, such as a human constant region, for example an effector-null human constant region, e.g. an IgG4 constant region or an IgG1 constant region, optionally wherein the constant region is IgG4-PE (Seq ID No:199), or a disabled IgG1 as defined in Seq ID No:205.
[0497] In other embodiments, the antibody or fragment is any of the isotypes or constant regions as defined hereinabove. In one embodiment, the constant region is wild-type human IgG1 (Seq ID No:340). For example, the constant region is an effector-enabled IgG1 constant region, optionally having ADCC and/or CDC activity. In one embodiment, the constant region is engineered for enhanced ADCC and/or CDC and/or ADCP. In another embodiment, the constant region is engineered for enhanced effector function.
[0498] The IgG4 constant region may be any of the IgG4 constant region amino acid sequences, or encoded by any of the nucleic acid sequences of Seq ID Nos:192 to 203. A heavy chain constant region may be an IgG4 comprising both the Leu235Glu mutation and the Ser228Pro mutation. This "IgG4-PE" heavy chain constant region (Seq ID Nos:198, encoded by Seq ID Nos:199, 200 and 201) is effector null.
[0499] An alternative effector null human constant region is a disabled IgG1 being an IgG1*01 allele comprising the L235A and/or G237A mutations (e.g. LAGA, Seq ID No:204, encoded by Seq ID No:205). In one embodiment, the antibodies or antibody fragments disclosed herein comprise an IgG1 heavy chain constant region, wherein the sequence contains alanine at position 235 and/or 237 (EU index numbering).
[0500] The antibody-dependent cell phagocytosis (ADCP) mechanism is discussed in Gul et al., "Antibody-Dependent Phagocytosis of Tumor Cells by Macrophages: A Potent Effector Mechanism of Monoclonal Antibody Therapy of Cancer", Cancer Res., 75(23), Dec. 1, 2015.
[0501] The potency of Fc-mediated effects may be enhanced by engineering the Fc domain by various established techniques. Such methods increase the affinity for certain Fc-receptors, thus creating potential diverse profiles of activation enhancement. This can be achieved by modification of one or several amino acid residues (e.g. as described in Lazar et al., 2006, Proc. Natl. Acad. Sci. U.S.A., March 14; 103(11):4005-10; the modifications disclosed therein are incorporated herein by reference). Human IgG1 constant regions containing specific mutations or altered glycosylation on residue Asn297 (e.g. N297Q, EU index numbering) have been shown to enhance binding to Fc receptors. In one embodiment, such mutations are one or more of the residues selected from 239, 332 and 330 for human IgG1 constant regions (or the equivalent positions in other IgG isotypes). In one embodiment, the antibody or fragment comprises a human IgG1 constant region having one or more mutations independently selected from N297Q, S239D, I332E and A330L (EU index numbering).
[0502] In another embodiment, the increase in affinity for Fc-receptors is achieved by altering the natural glycosylation profile of the Fc domain by, for example, generating under fucosylated or defucosylated variants (as described in Natsume et al., 2009, Drug Des. Devel. Ther., 3:7-16 or by Zhou Q., Biotechnol. Bioeng., 2008, Feb. 15, 99(3):652-65, the modifications described therein are incorporated herein by reference). Non-fucosylated antibodies harbour a tri-mannosyl core structure of complex-type N-glycans of Fc without fucose residue. These glycoengineered antibodies that lack core fucose residue from the Fc N-glycans may exhibit stronger ADCC than fucosylated equivalents due to enhancement of Fc.gamma.RIIIa binding capacity. For example, to increase ADCC, residues in the hinge region can be altered to increase binding to Fc-.gamma.RIII (see, for example, Shields et al., 2001, J. Biol. Chem., March 2; 276(9):6591-604; the modifications described therein are incorporated herein by reference). Thus, in one embodiment, the antibody or fragment comprises a human IgG heavy chain constant region that is a variant of a wild-type human IgG heavy chain constant region, wherein the variant human IgG heavy chain constant region binds to human Fc.gamma. receptors selected from the group consisting of Fc.gamma.RIIB and Fc.gamma.RIIA with higher affinity than the wild type human IgG heavy chain constant region binds to the human Fc.gamma. receptors. In one embodiment, the antibody or fragment comprises a human IgG heavy chain constant region that is a variant of a wild type human IgG heavy chain constant region, wherein the variant human IgG heavy chain constant region binds to human Fc.gamma.RIIB with higher affinity than the wild type human IgG heavy chain constant region binds to human Fc.gamma.RIIB. In one embodiment, the variant human IgG heavy chain constant region is a variant human IgG1, a variant human IgG2, or a variant human IgG4 heavy chain constant region. In one embodiment, the variant human IgG heavy chain constant region comprises one or more amino acid mutations selected from G236D, P238D, S239D, S267E, L328F, and L328E (EU index numbering system). In another embodiment, the variant human IgG heavy chain constant region comprises a set of amino acid mutations selected from the group consisting of: S267E and L328F; P238D and L328E; P238D and one or more substitutions selected from the group consisting of E233D, G237D, H268D, P271G, and A330R; P238D, E233D, G237D, H268D, P271G, and A330R; G236D and S267E; S239D and S267E; V262E, S267E, and L328F; and V264E, S267E, and L328F (EU index numbering system). In another embodiment, the variant human IgG heavy chain constant region further comprises one or more amino acid mutations that reduce the affinity of the IgG for human Fc.gamma.RIIIA, human Fc.gamma.RIIA, or human Fc.gamma.RI. In one embodiments, the Fc.gamma.RIIB is expressed on a cell selected from the group consisting of macrophages, monocytes, B-cells, dendritic cells, endothelial cells, and activated T-cells. In one embodiment, the variant human IgG heavy chain constant region comprises one or more of the following amino acid mutations G236A, S239D, F243L, T256A, K290A, R292P, S298A, Y300L, V305I, A330L, I332E, E333A, K334A, A339T, and P396L (EU index numbering system). In one embodiment, the variant human IgG heavy chain constant region comprises a set of amino acid mutations selected from the group consisting of: S239D; T256A; K290A; S298A; I332E; E333A; K334A; A339T; S239D and I332E; S239D, A330L, and I332E; S298A, E333A, and K334A; G236A, S239D, and I332E; and F243L, R292P, Y300L, V305I, and P396L (EU index numbering system). In one embodiment, the variant human IgG heavy chain constant region comprises a S239D, A330L, or I332E amino acid mutations (EU index numbering system). In one embodiment, the variant human IgG heavy chain constant region comprises an S239D and I332E amino acid mutations (EU index numbering system). In one embodiment, the variant human IgG heavy chain constant region is a variant human IgG1 heavy chain constant region comprising the S239D and I332E amino acid mutations (EU index numbering system). In one embodiment, the antibody or fragment comprises an afucosylated Fc region. In another embodiment, the antibody or fragment thereof is defucosylated. In another embodiment, the antibody or fragment is under fucosylated.
[0503] In another embodiment, the antibodies and fragments disclosed herein may comprise a triple mutation (M252Y/S254T/T256E) which enhances binding to FcRn. See DaII et al., Immunol 2002; 169:5171-5180 for a discussion of mutations affection FcRn binding in table 2, the mutations described therein are incorporated herein by reference.
[0504] Equally, the enhancement of CDC may be achieved by amino acid changes that increase affinity for C1q, the first component of the classic complement activation cascade (see Idusogie et al., J. Immunol., 2001, 166:2571-2575; the modifications described are incorporated herein by reference). Another approach is to create a chimeric Fc domain created from human IgG1 and human IgG3 segments that exploit the higher affinity if IgG3 for C1q (Natsume et al., 2008, Cancer Res., 68: 3863-3872; the modifications are incorporated herein by reference). In another embodiment, the antibody or antibody fragments disclosed herein may comprise mutated amino acids at residues 329, 331 and/or 322 to alter the C1q binding and/or reduced or abolished CDC activity. In another embodiment, the antibodies or antibody fragments disclosed herein may contain Fc regions with modifications at residues 231 and 239, whereby the amino acids are replaced to alter the ability of the antibody to fix complement. In one embodiment, the antibody or fragment has a constant region comprising one or more mutations selected from E345K, E430G, R344D and D356R, in particular a double mutation comprising R344D and D356R (EU index numbering system).
[0505] An antibody may have a heavy chain constant region that binds one or more types of Fc receptor but does not induce cellular effector functions, i.e. which does not mediate ADCC, CDC or ADCP activity. Such a constant region may be unable to bind the particular Fc receptor(s) responsible for triggering ADCC, CDC or ADCP activity. An antibody may have a heavy chain constant region that does not bind Fc.gamma. receptors. Thus, in one embodiment, the constant region may comprise a Leu235Glu mutation (EU index numbering system).
[0506] In another embodiment, the antibodies and fragments disclosed herein are modified to increase or decrease serum half-life. In one embodiment, one or more of the following mutations: T252L, T254S or T256F are introduced to increase biological half-life of the antibody. Biological half-life can also be increased by altering the heavy chain constant region CH.sub.1 domain or CL region to contain a salvage receptor binding epitope taken from two loops of a CH.sub.2 domain of an Fc region of an IgG, as described in U.S. Pat. Nos. 5,869,046 and 6,121,022, the modifications described therein are incorporated herein by reference. In another embodiment, the Fc hinge region of an antibody or antigen-binding fragment of the invention is mutated to decrease the biological half-life of the antibody or fragment. One or more amino acid mutations are introduced into the CH.sub.2-CH.sub.3 domain interface region of the Fc-hinge fragment such that the antibody or fragment has impaired Staphylococcyl protein A (SpA) binding relative to native Fc-hinge domain SpA binding. Other methods of increasing serum half-life are known to those skilled in the art. Thus, in one embodiment, the antibody or fragment is PEGylated. In another embodiment, the antibody or fragment is fused to an albumin-bidnig domain, e.g. an albumin binding single domain antibody (dAb). In another embodiment, the antibody or fragment is PASylated (i.e. genetic fusion of polypeptide sequences composed of PAS (XL-Protein GmbH) which forms uncharged random coil structures with large hydrodynamic volume). In another embodiment, the antibody or fragment is XTENylated.RTM./rPEGylated (i.e. genetic fusion of non-exact repeat peptide sequence (Amunix, Versartis) to the therapeutic peptide). In another embodiment, the antibody or fragment is ELPylated (i.e. genetic fusion to ELP repeat sequence (PhaseBio)). These various half-life extending fusions are described in more detail in Strohl, BioDrugs (2015) 29:215-239, which fusions, e.g. in Tables 2 and 6, are incorporated herein by reference.
[0507] The antibody may have a modified constant region which increases stability. Thus, in one embodiment, the heavy chain constant region comprises a Ser228Pro mutation. In another embodiment, the antibodies and fragments disclosed herein comprise a heavy chain hinge region that has been modified to alter the number of cysteine residues. This modification can be used to facilitate assembly of the light and heavy chains or to increase or decrease the stability of the antibody.
Concept 31. The antibody or fragment according to concept 30, wherein the constant region is a murine constant region.
[0508] In other embodiments, the constant region may be of any non-human mammalian origin, e.g. rat, mouse, hamster, guinea pig, dog, cat, horse, chicken, llama, dromedary, etc. In one embodiment, the constant region is a rat constant region. In another embodiment, the constant region is a llama constant region. The murine constant region may be any of the isotypes or alleles described hereinabove.
Concept 32. The antibody or fragment according to concept 30 or concept 31, wherein the constant region has CDC and/or ADCC activity. Concept 33. The antibody according to any preceding concept wherein the: [0509] a) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0510] b) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:33, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:43; [0511] c) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0512] d) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0513] e) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0514] f) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:342 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0515] g) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0516] h) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0517] i) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0518] j) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0519] k) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:342 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0520] l) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0521] m) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0522] n) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0523] o) V.sub.H domain comprise an amino acid sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0524] p) V.sub.H domain comprise an amino acid sequence of the V.sub.H domain of SEQ ID No:342 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0525] q) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0526] r) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0527] s) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0528] t) V.sub.H domain comprise an amino acid sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0529] u) V.sub.H domain comprise an amino acid sequence of the V.sub.H domain of SEQ ID No:342 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0530] v) V.sub.H domain comprises an amino acid sequence of SEQ ID No:58 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:68; [0531] w) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:58, and the V.sub.L domain comprise an amino acid sequence that is at least 85% identical to SEQ ID No:68; [0532] x) V.sub.H domain comprises an amino acid sequence of SEQ ID No:78 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:88; [0533] y) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:78, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:88; [0534] z) V.sub.H domain comprises an amino acid sequence of SEQ ID No:98 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:108; [0535] aa) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:98, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:108; [0536] bb) V.sub.H domain comprises an amino acid sequence of SEQ ID No:118 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:128; [0537] cc) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:118, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:128; [0538] dd) V.sub.H domain comprises an amino acid sequence of SEQ ID No:158 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:168; [0539] ee) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:158, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:168; [0540] ff) V.sub.H domain comprises an amino acid sequence of SEQ ID No:178 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:188; [0541] gg) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:178, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:188; [0542] hh) V.sub.H domain comprises an amino acid sequence of SEQ ID No:138 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:148; [0543] ii) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:138 and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:148; [0544] jj) V.sub.H domain comprises an amino acid sequence of SEQ ID No:244 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:254; [0545] kk) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:244, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:254; [0546] ll) V.sub.H domain comprises an amino acid sequence of SEQ ID No:264 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:274; [0547] mm) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:264, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:274; [0548] nn) V.sub.H domain comprises an amino acid sequence of SEQ ID No:284 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:294; and [0549] oo) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:284, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:294; [0550] pp) V.sub.H domain comprises an amino acid sequence of SEQ ID No:349 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:359; and [0551] qq) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:349, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:359.
[0552] In one embodiment, the amino acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99.5% identical to the specified Seq ID No.
Concept 34. The antibody according to any preceding concept wherein the antibody comprises a heavy chain and a light chain, and [0553] a) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:35 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:45; [0554] b) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:35 and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:45; [0555] c) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:47 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:45; [0556] d) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:48 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:45; [0557] e) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:49 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:45; [0558] f) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:342 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:45; [0559] g) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:35 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:50; [0560] h) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:47 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:50; [0561] i) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:48 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:50; [0562] j) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:49 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:50; [0563] k) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:342 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:50; [0564] l) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:35 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:51; [0565] m) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:47 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:51; [0566] n) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:48 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:51; [0567] o) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:49 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:51; [0568] p) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:342 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:51; [0569] q) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:35 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:298; [0570] r) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:47 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:298; [0571] s) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:48 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:298; [0572] t) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:49 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:298; [0573] u) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:342 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:298; [0574] v) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:60 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:70; [0575] w) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:60, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:70; [0576] x) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:80 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:90; [0577] y) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:80, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:90; [0578] z) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:100 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:110; [0579] aa) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:100, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:110; [0580] bb) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:120 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:130; [0581] cc) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:120, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:130; [0582] dd) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:160 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:170; [0583] ee) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:160, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:170; [0584] ff) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:180 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:190; [0585] gg) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:180, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:190 [0586] hh) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:140 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:150; [0587] ii) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:140, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:150; [0588] jj) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:246 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:256; [0589] kk) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:246, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:256; [0590] ll) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:266 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:276; [0591] mm) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:266, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:276; [0592] nn) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:286 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:296; and [0593] oo) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:286, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:296; [0594] pp) the heavy chain amino acid sequence comprises an amino acid sequence of SEQ ID No:351 and the light chain amino acid sequence comprises an amino acid sequence of SEQ ID No:361; and [0595] qq) the heavy chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:351, and the light chain amino acid sequence comprises an amino acid sequence that is at least 85% identical to SEQ ID No:361.
[0596] In one embodiment, the amino acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99.5% identical to the specified Seq ID No.
Concept 35. The antibody or fragment according to any preceding concept which competes for binding to hPD-L1 with the antibody 1D05, optionally wherein the competition for binding to hPD-L1 is conducted using SPR.
[0597] SPR may be carried out as described hereinabove, or as described in concept 16.
Concept 36. The antibody or fragment according to any preceding concept wherein the antibody or fragment is capable of inhibiting PD-L1-mediated suppression of T-cells, optionally wherein the suppression of T-cells is measured by an increase in one or more of IFN.gamma., IL-2, CD25 or proliferation of T-cells in an assay that provides co-stimulation by either direct CD3/CD28 stimulation, superantigen stimulation or provides co-stimulation by co-incubation with cells capable of inducing a T-cell response.
[0598] The measurements may be carried out with any suitable technique. For example, the measurements may be taken with ELISA, HTRF, BRDU incorporation (proliferation), electrochemiluminescence (ECL) or flow cytometry (e.g. FACS). These techniques are well-known to those skilled in the art and are described elsewhere herein. In one embodiment, the assay is flow cytometry. In one embodiment, the assay is ELISA. In one embodiment, the assay is HTRF.
[0599] In one embodiment, the suppression of T-cells is measured by an increase in IFN.gamma.. In one embodiment, the suppression of T-cells is measured by an increase in IL-2. In one embodiment, the suppression of T-cells is measured by an increase in CD25. In one embodiment, the suppression of T-cells is measured by an increase in IFN.gamma. and IL-2. In one embodiment, the suppression of T-cells is measured by an increase in IFN.gamma. and CD25. In one embodiment, the suppression of T-cells is measured by an increase in CD25 and IL-2. In one embodiment, the suppression of T-cells is measured by an increase in IFN.gamma., IL-2 and CD25.
[0600] In one embodiment, the co-stimulation is provided by direct CD3/CD28 stimulation.
[0601] In one embodiment, the co-stimulation is provided by a superantigen, such as staphylococcal enterotoxin B (SEB).
[0602] In one embodiment, the assay provides co-stimulation by co-incubation with cells capable of inducing a T-cell response. Such cells may be antigen-presenting cells (APCs), for example monocytes, B-cells or dendritic cells. In one embodiment, the assay provides co-stimulation by co-incubation with APCs. In one embodiment, the assay provides co-stimulation by co-incubation with monocytes. In one embodiment, the assay provides co-stimulation by co-incubation with B-cells. In one embodiment, the assay provides co-stimulation by co-incubation with dendritic cells.
Concept 37. A bispecific antibody or fusion protein comprising an antibody or fragment thereof as defined in any preceding concept.
[0603] Concept 37a. A dual binding antibody or fusion protein comprising an antibody or fragment thereof as defined in any preceding concept.
[0604] A dual binding antibody has the meaning as set out above.
Concept 38. The bispecific antibody according to concept 37, wherein the bispecific format is selected from DVD-Ig, mAb.sup.2, FIT-Ig, mAb-dAb, dock and lock, SEEDbody, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BiTE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, minibody, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, in particular mAb.sup.2, knob-in-holes, knob-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs and FIT-Ig, e.g. mAb.sup.2 and FIT-Ig.
[0605] In one embodiment, the bispecific format is selected from DVD-Ig, mAb.sup.2, FIT-Ig, mAb-dAb, dock and lock, Fab-arm exchange, SEEDbody, Triomab, LUZ-Y, Fcab, .kappa..lamda.-body, orthogonal Fab, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, Triple body, Miniantibody, minibody, TriBi minibody, scFv-CH.sub.3 KIH, scFv-CH-CL-scFv, F(ab').sub.2-scFv, scFv-KIH, Fab-scFv-Fc, tetravalent HCab, ImmTAC, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, DT-IgG, DutaMab, IgG(H)-scFv, scFv-(H)IgG, IgG(L)-scFv, scFv-(L)IgG, IgG(L,H)-Fv, IgG(H)-V, V(H)--IgG, IgG(L)-V, V(L)-IgG, KIH IgG-scFab, 2scFv-IgG, IgG-2scFv, scFv4-Ig and zybody.
[0606] In one embodiment, the bispecific format is selected from DVD-Ig, FIT-Ig, mAb-dAb, dock and lock, Fab-arm exchange, SEEDbody, Triomab, LUZ-Y, Fcab, .kappa..lamda.-body, orthogonal Fab, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, Triple body, Miniantibody, minibody, TriBi minibody, scFv-CH.sub.3 KIH, scFv-CH-CL-scFv, F(ab')z-scFv, scFv-KIH, Fab-scFv-Fc, tetravalent HCab, ImmTAC, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, DT-IgG, DutaMab, IgG(H)-scFv, scFv-(H)IgG, IgG(L)-scFv, scFv-(L)IgG, IgG(L,H)-Fv, IgG(H)-V, V(H)-IgG, IgG(L)-V, V(L)-IgG, KIH IgG-scFab, 2scFv-IgG, IgG-2scFv, scFv4-Ig and zybody, for example DVD-Ig, FIT-Ig, mAb-dAb, dock and lock, SEEDbody, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, minibody, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, in particular knob-in-holes, knob-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs and FIT-Ig, e.g. FIT-Ig.
[0607] In one embodiment, the bispecific format is selected from DVD-Ig, mAb.sup.2, mAb-dAb, dock and lock, Fab-arm exchange, SEEDbody, Triomab, LUZ-Y, Fcab, .kappa..lamda.-body, orthogonal Fab, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, Triple body, Miniantibody, minibody, TriBi minibody, scFv-CH.sub.3 KIH, scFv-CH-CL-scFv, F(ab').sub.2-scFv, scFv-KIH, Fab-scFv-Fc, tetravalent HCab, ImmTAC, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, DT-IgG, DutaMab, IgG(H)-scFv, scFv-(H)IgG, IgG(L)-scFv, scFv-(L)IgG, IgG(L,H)-Fv, IgG(H)-V, V(H)--IgG, IgG(L)-V, V(L)-IgG, KIH IgG-scFab, 2scFv-IgG, IgG-2scFv, scFv4-Ig and zybody, for example DVD-Ig, mAb.sup.2, mAb-dAb, dock and lock, SEEDbody, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, minibody, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, in particular mAb.sup.2, knob-in-holes, knobs-in-holes with common light chain and charge pairs, and knob-in-holes with common light chain, e.g. mAb.sup.2.
[0608] In one embodiment, the bispecific format is selected from DVD-Ig, mAb-dAb, dock and lock, Fab-arm exchange, SEEDbody, Triomab, LUZ-Y, Fcab, .kappa..lamda.-body, orthogonal Fab, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BITE, diabody, DART, TandAb, scDiabody, scDiabody-CHs, Diabody-CH.sub.3, Triple body, Miniantibody, minibody, TriBi minibody, scFv-CH.sub.3 KIH, scFv-CH-CL-scFv, F(ab').sub.2-scFv, scFv-KIH, Fab-scFv-Fc, tetravalent HCab, ImmTAC, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, DT-IgG, DutaMab, IgG(H)-scFv, scFv-(H)IgG, IgG(L)-scFv, scFv-(L)IgG, IgG(L,H)-Fv, IgG(H)-V, V(H)-IgG, IgG(L)-V, V(L)-IgG, KIH IgG-scFab, 2scFv-IgG, IgG-2scFv, scFv4-Ig and zybody, for example DVD-Ig, mAb-dAb, dock and lock, SEEDbody, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BiTE, diabody, DART, TandAb, scDiabody, scDiabody-CH.sub.3, Diabody-CH.sub.3, minibody, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, in particular knob-in-holes, knobs-in-holes with common light chain and charge pairs, and knob-in-holes with common light chain.
Concept 39. The bispecific antibody according to concept 37 or concept 38, wherein the bispecific antibody specifically binds to hPD-L1 and another target antigen selected from immune checkpoint inhibitors (such as PD-1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, TIM-3 and LAG-3), immune modulators (such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10, CXCL11 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R), immune activators (such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD27, CD3, ICOS (e.g. agonistic anti-ICOS antibodies), for example. ICOS, CD137, GITR and OX40).
[0609] Concept 39a. A bispecific antibody which binds to hPD-L1 with a V.sub.H, a V.sub.L, or a paired V.sub.H and V.sub.L comprising one or more of the CDRs (e.g. CDRH3 and CDRL3) or variable region sequences of any of the antibodies described in Aspect 1a hereinbelow, and another target antigen selected from immune checkpoint inhibitors (such as PD-1, CRA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, TIM-3 and LAG-3), immune modulators (such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10, CXCL11 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R), immune activators (such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD27, CD3, ICOS (e.g. agonistic anti-ICOS antibodies), for example. ICOS, CD137, GITR and OX40).
[0610] Concept 39b. The bispecific antibody according to concept 37 or concept 38, wherein the bispecific antibody specifically binds to hPD-L1 and another target antigen selected from immune checkpoint inhibitors (such as PD-1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, TIM-3 and LAG-3), immune modulators (such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R), immune activators (such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD3, ICOS (e.g. agonistic anti-ICOS antibodies), for example. ICOS, CD137, GITR and OX40).
[0611] In one embodiment, the another target antigen is an immune checkpoint inhibitor, such as PD-1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, CTLA-4, TIM-3 and LAG-3. In one embodiment, the another target antigen is an immune modulator, such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10, CXCL11 and CD155, or such as such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10 and CD155 e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R. In one embodiment, the another target antigen is an immune activator, such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD27, CD3 and ICOS (e.g. agonistic anti-ICOS antibodies), or CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD3 and ICOS (e.g. agonistic anti-ICOS antibodies), for example ICOS, CD137, GITR and OX40). In one embodiment, the another target antigen is CTLA-4. In one embodiment, the another target antigen is TIGIT. In one embodiment, the another target antigen is TIM-3. In one embodiment, the another target antigen is LAG-3. In one embodiment, the another target antigen is GITR. In one embodiment, the another target antigen is VISTA. In one embodiment, the another target antigen is CD137. In one embodiment, the another target antigen is SIRP.alpha.. In one embodiment, the another target antigen is CXCL10. In one embodiment, the another target antigen is CD155. In one embodiment, the another target antigen is CD40.
[0612] In another embodiment, the bispecific antibody binds another target antigen which is PD-1 and the binding to PD-1 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0613] In another embodiment, the bispecific antibody binds another target antigen which is CTLA4 and the binding to CTLA4 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0614] In another embodiment, the bispecific antibody binds another target antigen which is TIGIT and the binding to TIGIT is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0615] In another embodiment, the bispecific antibody binds another target antigen which is TIM-3 and the binding to TIM-3 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0616] In another embodiment, the bispecific antibody binds another target antigen which is LAG3 and the binding to LAG3 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0617] In another embodiment, the bispecific antibody binds another target antigen which is VISTA and the binding to VISTA is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0618] In another embodiment, the bispecific antibody binds another target antigen which is BTLA and the binding to BTLA is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0619] In another embodiment, the bispecific antibody binds another target antigen which is hHVEM and the binding to hHVEM is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0620] In another embodiment, the bispecific antibody binds another target antigen which is CSF1R and the binding to CSF1R is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0621] In another embodiment, the bispecific antibody binds another target antigen which is CCR4 and the binding to CCR4 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0622] In another embodiment, the bispecific antibody binds another target antigen which is CD39 and the binding to CD39 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0623] In another embodiment, the bispecific antibody binds another target antigen which is CD40 and the binding to CD40 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0624] In another embodiment, the bispecific antibody binds another target antigen which is CD73 and the binding to CD73 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0625] In another embodiment, the bispecific antibody binds another target antigen which is CD96 and the binding to CD96 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0626] In another embodiment, the bispecific antibody binds another target antigen which is CXCR2 and the binding to CXCR2 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0627] In another embodiment, the bispecific antibody binds another target antigen which is CXCR4 and the binding to CXCR4 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0628] In another embodiment, the bispecific antibody binds another target antigen which is CD200 and the binding to CD200 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0629] In another embodiment, the bispecific antibody binds another target antigen which is GARP and the binding to GARP is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0630] In another embodiment, the bispecific antibody binds another target antigen which is SIRP.alpha. and the binding to SIRP.alpha. is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0631] In another embodiment, the bispecific antibody binds another target antigen which is CXCL9 and the binding to CXCL9 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0632] In another embodiment, the bispecific antibody binds another target antigen which is CXCL10 and the binding to CXCL10 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0633] In another embodiment, the bispecific antibody binds another target antigen which is CXCL11 and the binding to CXCL11 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0634] In another embodiment, the bispecific antibody binds another target antigen which is CD155 and the binding to CD155 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0635] In another embodiment, the bispecific antibody binds another target antigen which is CD137 and the binding to CD137 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0636] In another embodiment, the bispecific antibody binds another target antigen which is GITR and the binding to GITR is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0637] In another embodiment, the bispecific antibody binds another target antigen which is OX40 and the binding to OX40 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0638] In another embodiment, the bispecific antibody binds another target antigen which is CD40 and the binding to CD40 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0639] In another embodiment, the bispecific antibody binds another target antigen which is CXCR3 and the binding to CXCR3 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0640] In another embodiment, the bispecific antibody binds another target antigen which is CD27 and the binding to CD27 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0641] In another embodiment, the bispecific antibody binds another target antigen which is CD3 and the binding to CD3 is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in Aspect 1A hereinbelow.
[0642] In another embodiment, the bispecific antibody binds another target antigen which is ICOS and the binding to ICOS is provided by an antigen-binding domain (for example, a V.sub.H, a V.sub.L or a paired V.sub.H and V.sub.L) having any of the sequences, including CDR sequences (for example CDRH3 and/or CDRL3) or variable region sequences as described in arrangement 5 and arrangement 5a hereinbelow, and any of the anti-ICOS antibodies described in sentences 1 to 102 and sentences 1a to 21a.
[0643] In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds GITR (optionally wherein the GITR Fab has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds GITR (optionally wherein the GITR antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the FIT-Ig is effector-enabled (e.g. as described in any of concepts 30 to 32). In another embodiment, the FIT-Ig is effector-disabled (e.g. is an IgG4 format, or as described in any of concepts 30 to 31).
[0644] In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds ICOS (e.g. binds with agonistic activity and optionally wherein the ICOS Fab has a sequence--including CDRs and variable regions--as defined in arrangement 5, or in arrangement 5a, or in sentences 1 to 102, or in sentences 1a to 21a hereinbelow). In one embodiment, the ICOS Fab has a sequence of any of the ICOS antibodies described herein in sentences 1 to 102 or in sentences 1a to 21a) In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds ICOS (e.g. binds with agonistic activity or optionally wherein the ICOS antibody has a sequence--including CDRs and variable regions--as defined in arrangement 5, or in arrangement 5a, or in sentences 1 to 102, or in sentences 1a to 21a hereinbelow) and a Fab which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1A hereinbelow). In one embodiment, the FIT-Ig is effector-enabled (e.g. as described in any of concepts 30 to 32). In another embodiment, the FIT-Ig is effector-disabled (e.g. is an IgG4 format, or as described in any of concepts 30 or 31).
[0645] In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds TIM-3 (optionally wherein the TIM-3 Fab has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds TIM-3 (optionally wherein the TIM-3 antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the FIT-Ig is effector-enabled (e.g. as described in any of concepts 30 to 32). In another embodiment, the FIT-Ig is effector-disabled (e.g. is an IgG4 format, or as described in any of concepts 30 or 31).
[0646] In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one, of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds CD137 (optionally wherein the CD137 Fab has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds CD137 (optionally wherein the CD137 antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the FIT-Ig is effector-enabled (e.g. as described in any of concepts 30 to 32). In another embodiment, the FIT-Ig is effector-disabled (e.g. is an IgG4 format, or as described in any of concepts 30 or 31).
[0647] In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.3, CH.sub.2 and CH.sub.3) which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds CD3 (optionally wherein the CD3 Fab has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds CD3 (optionally wherein the CD3 antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the FIT-Ig is effector-enabled (e.g. as described in any of concepts 30 to 32). In another embodiment, the FIT-Ig is effector-disabled (e.g. is an IgG4 format, or as described in any of concepts 30 or 31).
[0648] Any of the targets listed above (and the Fabs and/or full antibodies described in more detail in Aspect 1A) may be applied to the FIT-Ig structure.
Concept 40. The bispecific antibody according to concept 39, wherein the another target antigen is TIGIT or LAG3.
[0649] In any of concepts 37 to 40, if the antibody or fragment thereof has the heavy and light variable region sequences of 84G09, then the bispecific antibody shall be interpreted as not including a mAb.sup.2 format wherein the Fcab has binding affinity to LAG3.
[0650] In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds TIGIT (optionally wherein the TIGIT Fab has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds TIGIT (optionally wherein the TIGIT antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the FIT-Ig is effector-enabled (e.g. as described in any of concepts 30 to 32). In another embodiment, the FIT-Ig is effector-disabled (e.g. is an IgG4 format, or as described in any of concepts 30 or 31).
[0651] In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds LAG3 (optionally wherein the LAG3 Fab has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the bispecific antibody has a FIT-Ig format which comprises a full antibody (e.g. an antibody comprising a light chain comprising a V.sub.L and C.sub.L and a heavy chain comprising V.sub.H, CH.sub.1, CH.sub.2 and CH.sub.3) which binds LAG3 (optionally wherein the LAG3 antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow) and a Fab which binds hPD-L1 (optionally wherein the antibody has a structure as defined in any one of concepts 1 to 40, or wherein the antibody has a sequence--including CDRs and variable regions--as defined in Aspect 1a hereinbelow). In one embodiment, the FIT-Ig is effector-enabled (e.g. as described in any of concepts 30 to 32). In another embodiment, the FIT-Ig is effector-disabled (e.g. is an IgG4 format, or as described in any of concepts 30 or 31).
Concept 41. An antibody or fragment as defined in any preceding concept for use in treating or preventing a hPD-L1-mediated disease or condition, e.g. selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma). (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas). Concept 42. Use of an antibody or fragment as defined in any one of concepts 1 to 40 in the manufacture of a medicament for administration to a human for treating or preventing a hPD-L1 mediated disease or condition in the human, e.g. selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas). Concept 43. A method of treating or preventing a hPD-L1 mediated disease or condition, e.g. selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas) in a human, comprising administering to said human a therapeutically effective amount of an antibody or fragment as defined in any one of concepts 1 to 40, wherein the hPD-L1 mediated disease or condition is thereby treated or prevented.
[0652] In any of concepts 41 to 43, the hPD-L1 mediated disease may be any of those as described herein. In one embodiment, in any of concepts 41 to 43, the hPD-L1 mediated disease is a virally induced cancer, such as cervical cancer and nasopharyngeal cancer, for example cervical cancers caused by HPV infection. In one embodiment, in any of concepts 41 to 43, the hPD-L1 mediated disease is a chronic viral infection. In one embodiment, in any of concepts 41 to 43, the hPD-L1 mediated disease is a neoplastic disease. In one embodiment, in any of concepts 41 to 43, the hPD-L1 mediated disease is a non-neoplastic disease. In one embodiment, in any of concepts 41 to 43, the hPD-L1 mediated disease is a malignant tumour. In one embodiment, in any of concepts 41 to 43, the hPD-L1 mediated disease is a cancer which is known to be responsive to PD-L1 therapy, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma. In one embodiment, in any of concepts 41 to 43, the hPD-L1 mediated disease is a cancer which is a soft tissue sarcoma.
Concept 44. The antibody or fragment according to concept 41, the use according to concept 42 or the method according to concept 43, wherein the hPD-L1-mediated disease or condition is cancer.
[0653] Concept 44a. The antibody or fragment according to concept 41, the use according to concept 42 or the method according to concept 43, wherein the hPD-L1-mediated disease or condition is a neurodegenerative disease, disorder or condition, optionally wherein the neurodegenerative disease, disorder or condition is selected from Alzheimer's disease, amyotrophic lateral sclerosis, Parkinson's disease, Huntington's disease, primary progressive multiple sclerosis, secondary progressive multiple sclerosis, corticobasal degeneration, Rett syndrome, a retinal degeneration disorder selected from age-related macular degeneration and retinitis pigmentosa; anterior ischemic optic neuropathy, glaucoma, uveitis, depression, trauma-associated stress or post-traumatic stress disorder, frontotemporal dementia, Lewy body dementias, mild cognitive impairments, posterior cortical atrophy, primary progressive aphasia and progressive supranuclear palsy or aged-related dementia, in particular Alzheimer's disease, amyotrophic lateral sclerosis, Parkinson's disease and Huntington's disease, and e.g. Alzheimer's disease.
[0654] In concept 44a, the therapeutically effective amount of an antibody or fragment may comprise an antigen-binding site that specifically binds PD-L1, e.g. hPD-L1.
[0655] In one embodiment, the antigen-binding site specifically binds PD-L1, e.g. hPD-L1. In one embodiment, the PD-L1 antigen-binding site comprises the CDRH1, CDRH2, CDR3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L. region from any one of the anti-PD-L1 antibodies selected from atezolizumab (Roche), avelumab (Merck), BMS-936559/MDX-1105 (BMS), durvalumab/Medi4736 (Medimmune), KN-035, CA-170, FAZ-053 M7824, ABBV-368, LY-3300054, GNS-1480, YW243.55.S70, REGN3504 and any of the PD-L1 antibodies disclosed in WO2017/034916, WO2017/020291, WO2017/020858, WO2017/020801, WO2016/111645, WO2016/197367, WO2016/061142, WO2016/149201, WO2016/000619, WO2016/160792, WO2016/022630, WO2016/007235, WO2015/179654, WO2015/173267, WO2015/181342, WO2015/109124, WO2015/112805, WO2015/061668, WO2014/159562, WO2014/165082, WO2014/100079, WO2014/055897, WO2013/181634, WO2013/173223, WO2013/079174, WO2012/145493, WO2011/066389, WO2010/077634, WO2010/036959, WO2010/089411 or WO2007/005874, which antibodies and sequences are incorporated herein by reference.
[0656] In another embodiment of concept 44a, the PD-L1 antigen-binding site comprises the CDRH1, CDRH2, CDR3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L. region from any one of the anti-PD-L1 antibodies selected from an anti-PD-L1 antibody disclosed herein, particularly the anti-PD-L1 antibody clones disclosed in concepts 16a through 16l, and more particularly anti-PD-L1 antibody clone 84G09.
[0657] In another embodiment of concept 44a, the PD-L1 antigen-binding site comprises the CDRH1, CDRH2, CDR3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from anti-PD-L1 antibody clone 84G09 and the hPD-L1-mediated disease or condition is Alzheimer's disease.
Concept 45. The antibody or fragment, the use or the method according to concept 44, wherein the cancer is selected from melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or is selected from virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas. Concept 46. The antibody or fragment, use or the method according to any one of concepts 41 to 45, further comprising administering to the human a further therapy, for example a further therapeutic agent, optionally wherein the further therapeutic agent is independently selected from the group consisting of: [0658] a. other immune checkpoint inhibitors (such as anti-TIM-3 antibodies, anti-CTLA-4 antibodies, anti-TIGIT antibodies and anti-LAG-3 antibodies); [0659] b. immune stimulators (such as anti-OX40 antibodies, anti-GITR antibodies, anti-CD137 antibodies, anti-ICOS antibodies and anti-CD40 antibodies); [0660] c. chemokine receptor antagonists (such as CXCR4, CCR4 and CXCR2); [0661] d. targeted kinase inhibitors (such as CSF-1R or VEGFR inhibitors); [0662] e. angiogenesis inhibitors (such as anti-VEGF-A or Delta-like Ligand-4); [0663] f. immune stimulating peptides or chemokines (such as CXCL9 or CXCL10); [0664] g. cytokines (such as IL-15 and IL-21); [0665] h. bispecific T-cell engagers (BiTEs) having at least one specificity against CD3 (e.g. CD3/CD19 BiTE); [0666] i. other bi-specific molecules (for example IL-15-containing molecules targeted towards tumour associated antigens, for example Epidermal growth factor receptors such as EGFR, Her-2, New York Esophageal Cancer-1 (NY-ESO-1), GD2, EpCAM or Melanoma Associated Antigen-3 (MAGE-A3)); [0667] j. oncolytic viruses (such as HSV virus (optionally which secretes GMCSF), Newcastle disease virus and Vaccinia virus); [0668] k. vaccination with tumour associated antigens (such as New York Esophageal Cancer-1 [NY-ESO-1], Melanoma Associated Antigen-3 [MAGE-3]); [0669] l. cell-based therapies (such as chimeric Antigen Receptor-T-cells (CAR-T) for example expressing anti-CD19, anti-EpCam or anti-mesothelin); [0670] m. bi-specific NK cell engagers having a specificity against an activating MK receptor such as NKG2D or CD16a; and [0671] n. adoptive transfer of tumour specific T-cells or LAK cells, or optionally wherein the further therapy is chemotherapy, radiotherapy and surgical removal of tumours.
[0672] Radiotherapy may be single dose or in fractionated doses, either delivered to affected tissues directly or to the whole body.
[0673] Chemotherapeutic agents may any as described hereinabove, in particular, agents that induce immunogenic cell death, for example platinum therapies, such as oxaliplatin. In one embodiment, the chemotherapy is a standard of care cytotoxic chemotherapy for the cancer being treated.
[0674] In this aspect, the bispecific molecules include "bispecific antibodies" and antibody fusion proteins, including those formats and molecules described in concepts 37 to 40.
[0675] The antibodies may be any of the sequences or antibodies described in arrangement 5, 5a or detailed in Aspect 1a.
[0676] The further therapeutic agents of this concept may be delivered by any method, which methods are well-known to those skilled in the art. For example, the further therapeutic agents may be delivered orally, systemically or locally (to the tumour environment). In one embodiment, the further therapeutic agent is delivered orally. In one embodiment, the further therapeutic agent is delivered systemically (e.g. intravenously). In one embodiment, the further therapeutic agent is delivered locally to the tumour environment.
[0677] Compositions and routes of administration are described in more detail hereinbelow.
Concept 47. The antibody or fragment, use or the method according to concept 46, wherein the further therapeutic agent is administered sequentially or simultaneously with the anti-hPD-L1 antibody or fragment. Concept 48. A pharmaceutical composition comprising an antibody of fragment as defined in any one of concepts 1 to 40 and a pharmaceutically acceptable excipient, diluent or carrier and optionally further comprising a further therapeutic agent independently selected from the group consisting of: [0678] a) other immune checkpoint inhibitors (such as anti-TIM-3 antibodies, anti-CTLA-4 antibodies, anti-TIGIT antibodies and anti-LAG-3 antibodies); [0679] b) immune stimulators (such as anti-OX40 antibodies, anti-GITR antibodies, anti-CD137 antibodies, anti-ICOS antibodies and anti-CD40 antibodies); [0680] c) chemokine receptor antagonists (such as CXCR4, CCR4 and CXCR2); [0681] d) targeted kinase inhibitors (such as CSF-1R or VEGFR inhibitors); [0682] e) angiogenesis inhibitors (such as anti-VEGF-A or Delta-like Ligand-4); [0683] f) immune stimulating peptides or chemokines (such as CXCL9 or CXCL10); [0684] g) cytokines (such as IL-15 and IL-21); [0685] h) bispecific T-cell engagers (BiTEs) having at least one specificity against CD3 (e.g. CD3/CD19 BiTE); [0686] i) other bi-specific molecules (for example IL-15-containing molecules targeted towards tumour associated antigens, for example Epidermal growth factor receptors such as EGFR, Her-2, New York Esophageal Cancer-1 (NY-ESO-1), GD2, EpCAM or Melanoma Associated Antigen-3 (MAGE-A3)); [0687] j) oncolytic viruses (such as HSV virus (optionally which secretes GMCSF), Newcastle disease virus and Vaccinia virus); [0688] k) vaccination with tumour associated antigens (such as New York Esophageal Cancer-1 [NY-ESO-1], Melanoma Associated Antigen-3 [MAGE-3]); [0689] l) cell-based therapies (such as chimeric Antigen Receptor-T-cells (CAR-T) for example expressing anti-CD19, anti-EpCam or anti-mesothelin); [0690] m) bi-specific NK cell engagers having a specificity against an activating MK receptor such as NKG2D or CD16a; and [0691] n) adoptive transfer of tumour specific T-cells or LAK cells.
[0692] Pharmaceutical formulations are well-known to those skilled in the art. In one embodiment, the antibody or fragment is administered intravenously. In one embodiment, the antibody or fragment is administered subcutaneously.
[0693] In an example, an antibody or fragment as disclosed herein is contained in a medical container, e.g. a vial, syringe, IV container or an injection device (such as an intraocular or intravitreal injection device). In an example, the antibody or fragment is in vitro, for example, in a sterile container.
[0694] In one embodiment, the composition is formulated in accordance with routine procedures as a pharmaceutical composition adapted for intravenous administration to human beings. Typically, compositions for intravenous administration are solutions in sterile isotonic aqueous buffer. Where necessary, the composition may also include a solubilizing agent and a local anesthetic such as lignocamne to ease pain at the site of the injection. Such compositions, however, may be administered by a route other than intravenous.
[0695] Generally, the ingredients of compositions are supplied either separately or mixed together in unit dosage form, for example, as a dry lyophilized powder or water free concentrate in a hermetically sealed container such as an ampoule or sachette indicating the quantity of active agent. Where the composition is to be administered by infusion, it can be dispensed with an infusion bottle containing sterile pharmaceutical grade water or saline. Where the composition is administered by injection, an ampoule of sterile water for injection or saline can be provided so that the ingredients may be mixed prior to administration.
[0696] In this aspect, the bispecific molecules include "bispecific antibodies" and antibody fusion proteins, including those formats and molecules described in concepts 37 to 40.
[0697] The further therapeutic agents of this concept may be delivered by any method, which methods are well-known to those skilled in the art. For example, the further therapeutic agents may be delivered orally, systemically or locally (to the tumour environment). In one embodiment, the further therapeutic agent is delivered orally. In one embodiment, the further therapeutic agent is delivered systemically (e.g. intravenously). In one embodiment, the further therapeutic agent is delivered locally to the tumour environment.
[0698] The antibodies may have any of the sequences or may be any of the antibodies described in arrangement 5, 5a or detailed in aspect 1a.
Concept 49. A pharmaceutical composition according to concept 48, or a kit comprising a pharmaceutical composition as defined in concept 48, wherein the composition is for treating and/or preventing a hPD-L1-mediated condition or disease, e.g. selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease, diffuse large B-cell lymphoma. Concept 50. A pharmaceutical composition according to concept 48 or concept 49 in combination with, or kit according to concept 49 comprising, a label or instructions for use to treat and/or prevent said disease or condition in a human; optionally wherein the label or instructions comprise a marketing authorisation number (e.g. an FDA or EMA authorisation number); optionally wherein the kit comprises an IV or injection device that comprises the antibody or fragment. Concept 51. A method of modulating PD-1/PD-L1 interaction in a patient, comprising administering an effective amount of an antibody or fragment as defined in any one of concepts 1 to 40 to said patient.
[0699] In another embodiment, there is provided a method of modulating CD80/PD-L1 interaction in a patient, comprising administering an effective amount of an antibody or fragment as defined in any one of concepts 1 to 40 to said patient. In another embodiment, the antibody or fragment modulates CD80/PD-L1 interaction, but does not modulate PD-1/PD-L1 interaction. In another embodiment, the antibody or fragment blocks CD80/PD-L1 interaction, but does not block PD-1/PD-L1 interaction. In another embodiment, the antibody or fragment inhibits CD80/PD-L1 interaction, but does not inhibit PD-1/PD-L1 interaction.
Concept 52. A method of inhibiting PD-L1 activity in a patient, comprising administering an effective amount of an antibody or fragment as defined in any one of concepts 1 to 40 to said patient.
[0700] In one embodiment, the antibody or fragment blocks or inhibits PD-1 binding to PD-L1. In one embodiment, the antibody or fragment blocks or inhibits CD80 binding to PD-L1.
Concept 53. A method of treating a proliferative disease in an animal (e.g. a human), comprising administering an effective amount of an antibody or fragment as defined in any one of concepts 1 to 40 to said patient.
[0701] Proliferative diseases may be any as described elsewhere herein.
Concept 54. A method of detecting PD-L1 expression in a sample, comprising contacting the sample with an antibody or fragment as defined in any one of concepts 1 to 40. Concept 55. A method comprising contacting a biological sample with an antibody or fragment as defined in any one of concepts 1 to 40 to form a complex with PD-L1 present in the sample and measuring the presence, absence or level of the complex in the biological sample. Concept 56. The method according to concept 55, wherein the presence, absence and/or level of PD-L1 expression is detected prior to treatment and a high level of surface expressed PD-L1 is indicative of successful treatment. Concept 57. The method according to concept 55, wherein the presence, absence and/or level of PD-L1 expression is detected during treatment as an early response biomarker. Concept 58. The method according to concept 55 or concept 57, wherein the presence, absence and/or level of PD-L1 expression is detected during or after treatment to help determine one or more of: whether treatment has been successful, whether treatment should continue, and/or whether treatment should be modified. Concept 59. The method according to any one of concepts 55 to 58, wherein therapy comprises treatment with an anti-PD-L1 antibody, optionally as defined in any one of concepts 1 to 40. Concept 60. A method for monitoring therapy efficacy, the method comprising detecting expression of surface expressed PD-L1 in a patient prior to therapy, and during or after therapy, wherein an antibody or fragment as defined in any one of concepts 1 to 40 is used to detect expression of surface expressed PD-L1. Concept 61. The method according to concept 60, wherein surface expressed PD-L1 expression is detected in vivo. Concept 62. The method according to concept 60, wherein surface expressed PD-L1 expression is detected in a tissue sample in vitro. Concept 63. A method for identifying binding partners for PD-L1, the method comprising immunoprecipitating an intact protein complex comprising PD-L1 using an antibody or fragment as defined in any one of concepts 1 to 40. Concept 64. A method of diagnosing a disease in a human subject associated with altered PD-L1 expression comprising the steps of contacting a biological sample from the human subject with an antibody as defined in concepts 1 to 40 to form a complex between the antibody and PD-L1 present in the sample; and detecting the amount of the complex. Concept 65. A nucleic acid that encodes the CDRH3 of an antibody or fragment as defined in any one of concepts 1 to 40.
[0702] Concept 65a. There is also provided a nucleic acid that encodes the CDRH2 of an antibody or fragment as defined in any one of concepts 1 to 40.
[0703] Concept 65b. There is also provided a nucleic acid that encodes the CDRH1 of an antibody or fragment as defined in any one of concepts 1 to 40.
[0704] Concept 65c. There is also provided a nucleic acid that encodes the CDRL1 of an antibody or fragment as defined in any one of concepts 1 to 40.
[0705] Concept 65d. There is also provided a nucleic acid that encodes the CDRL2 of an antibody or fragment as defined in any one of concepts 1 to 40.
[0706] Concept 65e. There is also provided a nucleic acid that encodes the CDRL3 of an antibody or fragment as defined in any one of concepts 1 to 40.
[0707] In one embodiment, the nucleic acid is an isolated and purified nucleic acid.
Concept 66. A nucleic acid that encodes a V.sub.H domain and/or a V.sub.L domain of an antibody or fragment as defined in any one of concepts 1 to 40.
[0708] The V.sub.H and V.sub.L domain nucleic acid sequences of the invention are provided in the sequence listing. In one embodiment, the nucleic acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 99.5% identical to the specified Seq ID No.
Concept 67. The nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:36 and/or SEQ ID NO:46.
[0709] Concept 67a. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:16 and/or SEQ ID NO:26.
[0710] Concept 67b. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:61 and/or SEQ ID NO:71.
[0711] Concept 67c. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:81 and/or SEQ ID NO:91.
[0712] Concept 67d. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:101 and/or SEQ ID NO:111.
[0713] Concept 67e. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:121 and/or SEQ ID NO:131.
[0714] Concept 67f. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:161 and/or SEQ ID NO:171.
[0715] Concept 67g. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:181 and/or SEQ ID NO:191.
[0716] Concept 67h. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:141 and/or SEQ ID NO:151.
[0717] Concept 67i. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:247 and/or SEQ ID NO:257.
[0718] Concept 67j. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:267 and/or SEQ ID NO:277.
[0719] Concept 67k. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:287 and/or SEQ ID NO:297.
[0720] Concept 67l. A nucleic acid according to concept 66 comprising a nucleotide sequence that is at least 80% identical to the sequence of SEQ ID NO:352 and/or SEQ ID NO:362.
[0721] In one embodiment, the nucleic acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the nucleic acid sequence is at least 99.5% identical to the specified Seq ID No.
Concept 68. A nucleic acid that encodes a heavy chain or a light chain of an antibody as defined in any one of concepts 1 to 40. Concept 69. A vector comprising the nucleic acid of any one of concepts 65 to 68; optionally wherein the vector is a CHO or HEK293 vector. Concept 70. A host comprising the nucleic acid of any one of concepts 65 to 68 or the vector of concept 69.
3. Immunocytokines
[0722] The inventors have described immunocytokines which comprise an antibody which binds to an immune checkpoint inhibitor, such as PD-L1 fused to either the N-terminus or C-terminus of the heavy chain or the light chain (for example, the C-terminus of the heavy or light chain, and in particular the light chain). The immunocytokines comprise a cytokine molecule, which may be IL-2 or a variant thereof (including variant having a 1 to 10 amino acid deletion at the N-terminus). The antibodies as described hereinabove may be used in any immunocytokine described herein.
[0723] Without being bound by theory, immunocytokines of the invention may provide one or more of the following advantageous properties: [0724] synergistic activity (by virtue of the therapeutic activity of antibody Fab portion in combination with the cytokine) [0725] improved tumour targeting [0726] ability to retain effector functions such as CDC, ADCC and/or ADCP [0727] reduced off-target effects [0728] reduced toxicity (e.g. compared to free cytokine or cytokine when fused to the heavy chain of an immunocytokine) [0729] reduced immunogenicity [0730] lower dose/frequency of dosing, in particular due to improved half life of light chain cytokine fusions as compared to heavy chain fusion equivalents [0731] Specificity for blocking only one of the ligands of PD-L1 (e.g. blocks CD80/PD-L1 interaction, but not PD-1/PD-L1 interaction) [0732] Solubility [0733] Stability [0734] Ease of formulation [0735] Frequency of dosing and/or route of administration [0736] Manufacturability (e.g. expression, ease of purification, isoforms)
[0737] 1D05 ICK comprises a heavy chain amino acid sequence of Seq ID No:299, and a light chain amino acid sequence of Seq ID No:300. The light chain comprises a V.sub.L domain comprising the CDRs and V.sub.L sequence of antibody 1D05 described hereinabove, fused at the heavy chain to full length, wild-type, human IL-2 cytokine. It does not contain a linker peptide. The heavy chain comprises a V.sub.H domain comprising the CDRs and V.sub.H sequence of antibody 1D05 described hereinabove, fused to a disabled IgG constant region (Seq ID No:205).
[0738] 1D05 D5-9 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D5-9 (Seq ID No:303), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0739] 1D05 D1-9 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-9 (Seq ID No:304), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0740] 1D05 D5-7 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D5-7 (Seq ID No:305), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0741] 1D05 D1 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1 (Seq ID No:306), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0742] 1D05 D1-2 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-2 (Seq ID No:307), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0743] 1D05 D1-3 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-3 (Seq ID No:308), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0744] 1D05 D1-4 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-4 (Seq ID No:309), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0745] 1D05 D1-5 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-5 (Seq ID No:310), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0746] 1D05 D1-6 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-6 (Seq ID No:311), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0747] 1D05 D1-7 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-7 (Seq ID No:312), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0748] 1D05 D1-8 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-8 (Seq ID No:313), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0749] 1D05 D9 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9 (Seq ID No:314), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0750] 1D05 D9-8 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-8 (Seq ID No:315), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0751] 1D05 D9-7 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-7 (Seq ID No:316), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0752] 1D05 D9-6 ICK comprises a heavy chain comprising a V.sub.L region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-6 (Seq ID No:317), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0753] 1D05 D9-4 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-4 (Seq ID No:318), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0754] 1D05 D9-3 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-3 (Seq ID No:319), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0755] 1D05 D9-2 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-2 (Seq ID No:320), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0756] 1D05 D2-6 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D2-6 (Seq ID No:321), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0757] 1D05 D3-7 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D3-7 (Seq ID No:322), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0758] 1D05 D4-8 ICK comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:33 (comprising the CDRs of 1D05 as described hereinabove) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D4-8 (Seq ID No:323), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0759] In any of the ICK constructs above, the IL-2 binding portion may be a variant IL-2, in particular an IL-2 having an R38A mutation (as described in amino acids 21-133 of the variant IL-2 described as SEQ ID NO:517) or an R38Q mutation (as described in amino acids 21-133 of the variant IL-2 described as SEQ ID NO:518).
[0760] In any of the ICK constructs above, the V.sub.H region of the 1D05 antibody may be exchanged for the V.sub.H region of mutated 1D05--Heavy Chain mutant 1 (Seq ID No:47), mutated 1D05--Heavy Chain mutant 2 (Seq ID No:48), mutated 1D05--Heavy Chain mutant 3 (Seq ID No:49) or mutated 1D05--Heavy Chain mutant 4 (Seq ID No:342). A preferred mutated heavy chain V.sub.H region of 1D05 is mutated 1D05--Heavy Chain mutant 4 (Seq ID No:342).
[0761] Thus, certain ICK constructs comprise:
[0762] Mutated 1D05--Heavy Chain mutant 4 D5-9 ICK, which comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:342 (comprising the CDRs of mutated 1D05--Heavy Chain mutant 4 as described herein) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D5-9 (Seq ID No:303), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0763] Mutated 1D05--Heavy Chain mutant 4 D1-9 ICK, which comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:342 (comprising the CDRs of mutated 1D05--Heavy Chain mutant 4 as described herein) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-9 (Seq ID No:304), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0764] Mutated 1D05--Heavy Chain mutant 4 D1-8 ICK, which comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:342 (comprising the CDRs of mutated 1D05--Heavy Chain mutant 4 as described herein) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D1-8 (Seq ID No:313), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0765] Mutated 1D05--Heavy Chain mutant 4 D9-7 ICK, which comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:342 (comprising the CDRs of mutated 1D05--Heavy Chain mutant 4 as described herein) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-7 (Seq ID No:316), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0766] Mutated 1D05--Heavy Chain mutant 4 D9-2 ICK, which comprises a heavy chain comprising a V.sub.H region amino acid sequence of Seq ID No:342 (comprising the CDRs of mutated 1D05--Heavy Chain mutant 4 as described herein) fused to a disabled IgG1 constant region with an amino acid sequence of Seq ID No:205. The light chain comprises a V.sub.L amino acid sequence of Seq ID No:43 (comprising the CDRs of 1D05 as described hereinabove) directly fused at the C-terminus to IL-2 D9-2 (Seq ID No:320), which is directly fused to amino acids 21 to 133 of hIL-2 (Seq ID No:324).
[0767] In any of the ICK constructs above, the V.sub.L region of the 1D05 antibody may be exchanged for the V.sub.L region of mutated 1D05--Light Chain mutant 1 (Seq ID No:50), mutated 1D05--Light Chain mutant 2 (Seq ID No:51) or mutated 1D05--Light Chain mutant 3 (Seq ID No:298).
[0768] In any of the ICK constructs above, both the V.sub.H and V.sub.L region of the 1D05 antibody may be exchanged for both the V.sub.H and V.sub.L regions of any of the other antibodies described herein, i.e. 84G09, 411608, 411C04, 411D07, 385F01, 413D08, 386H03, 389A03, 413G05, 413F09 and 414606.
[0769] In any of the ICK constructs above, the heavy chain constant region of Seq ID No:205 may be exchanged for any of the heavy chain constant regions of Seq ID Nos:193, 195, 197, 199, 203, 205, 340, 524, 526, 528, 530, 532 or 534.
[0770] Immunocytokines may be described in the following sentences or aspects. Unless otherwise apparent, the features of any of the concepts described hereinabove apply mutatis matandis to any of the aspects hereinbelow.
Aspect 1. An immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction: [0771] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and [0772] b) A heavy chain constant region; [0773] and wherein the light chain comprises in N- to C-terminal direction: [0774] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3; [0775] d) A light chain constant region, (C.sub.L); [0776] e) Optionally, a linker, (L); and [0777] f) An IL-2 cytokine; [0778] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to hPD-L1 as defined by Seq ID No:1, and competes for binding to said hPD-L1 with the antibody 1D05; and [0779] wherein the immunocytokine comprises a V.sub.H domain which comprises a CDRH3 comprising the motif X.sub.1GSGX.sub.2YGX.sub.3X.sub.4FD, wherein X.sub.1, X.sub.2 and X.sub.3 are independently any amino acid, and X.sub.4 is either present or absent, and if present, may be any amino acid.
[0780] In the aspects described herein, CDR sequences may be determined according to any method known to those skilled in the art, such as using the Kabat method, the IMGT method or the Chothia method, each of which are described in more detail herein. In one embodiment, the CDR regions are human CDR regions.
[0781] In addition to the CDR regions, the V.sub.H and/or V.sub.L domains may further comprise framework regions, such as FW1, FW2 and FW3. The V.sub.H and/or V.sub.L domains may be of any origin described herein, and may be for example, fully human, humanised, murine or camelid. In one embodiment, the V.sub.H and/or V.sub.L domains are human V.sub.H and/or V.sub.L domains. CDRs may be of a non-human origin (e.g. mouse origin) and be grafted onto human framework regions. In another embodiment, the CDRs are synthetic.
[0782] In another embodiment, V.sub.H regions may be selected from the group consisting of an antibody variable domain (e.g. a V.sub.L or a V.sub.H, an antibody single variable domain (domain antibody or dAb), a camelid V.sub.HH antibody single variable domain, a shark immunoglobulin single variable domain (NARV), a Nanobody.TM. or a camelised V.sub.H single variable domain); a T-cell receptor binding domain; an immunoglobulin superfamily domain; an agnathan variable lymphocyte receptor; a fibronectin domain (e.g. an Adnectin.TM.); an antibody constant domain (e.g. a CH.sub.3 domain, e.g. a CH.sub.2 and/or CH.sub.3 of an Fcab.TM.) wherein the constant domain is not a functional CH.sub.1 domain; an scFv; an (scFv)2; an sc-diabody; an scFab; a centyrin and an epitope binding domain derived from a scaffold selected from CTLA-4 (Evibody.TM.); a lipocalin domain; Protein A such as Z-domain of Protein A (e.g. an Affibody.TM. or SpA); an A-domain (e.g. an Avimer.TM. or Maxibody.TM.); a heat shock protein (such as and epitope binding domain derived from GroEI and GroES); a transferrin domain (e.g. a trans-body); ankyrin repeat protein (e.g. a DARPin.TM.); peptide aptamer; C-type lectin domain (e.g. Tetranectin.TM.); human .gamma.-crystallin or human ubiquitin (an affilin); a PDZ domain; scorpion toxin; and a kunitz type domain of a human protease inhibitor.
[0783] The constant region comprises at least two heavy chain constant region domains selected from CH.sub.1, CH.sub.2, CH.sub.3 and CH.sub.4. In one embodiment, the constant region comprises (or consists of) a CHI domain and a CH.sub.2 domain. In one embodiment, the constant region comprises (or consists of) a CHI domain, a hinge region and a CH.sub.2 domain. In one embodiment, the constant region comprises (or consists of) a CH.sub.1 domain and a CH.sub.3 domain, and optionally a hinge region. In one embodiment, the constant region comprises (or consists of) a CH.sub.1 domain and a CH.sub.4 domain, and optionally a hinge region. In one embodiment, the constant region comprises (or consists of) a CH.sub.1 domain, a CH.sub.2 domain and a CH.sub.3 domain, and optionally a hinge region. In one embodiment, the constant region comprises (or consists of) a CH.sub.1 domain, a CH.sub.2 domain and a CH.sub.4 domain, and optionally a hinge region. In one embodiment, the constant region comprises (or consists of) a CH.sub.1 domain, a CH.sub.3 domain and a CH.sub.4 domain, and optionally a hinge region. In one embodiment, the constant region comprises (or consists of) a full constant region.
[0784] The constant region may be of any isotype described herein, e.g. IgA, IgD, IgE, IgG, and IgM. In one embodiment, the constant region is of any origin described herein, and may be for example, human, murine or camelid. In one embodiment, the constant region is a (full) human constant region. In one embodiment, the constant region is a human IgG constant region. In one embodiment, the constant region is a (full) human IgG1 constant region. In one embodiment, the constant region is an effector null (full) human IgG1 constant region. In one embodiment, the constant region has CDC and/or ADCC and/or ADCP activity. In one embodiment, the constant region is engineered to enhance the CDC and/or ADCC and/or ADCP activity. The constant region may be any of the constant regions described in concepts 30 to 32 hereinabove.
[0785] The light chain constant region may be a kappa or lambda light chain constant region. The light chain constant region may be as described in concept 28 hereinabove.
[0786] An IL-2 cytokine is a cytokine molecule which confers IL-2 activity on one or both of the intermediate affinity IL-2 Receptor (.alpha..beta.) and the high affinity IL-2 receptor (.alpha..beta..gamma.). An IL-2 cytokine includes variant IL-2 cytokines. An IL-2 cytokine may be of human origin or of non-human origin, for example of a non-human mammal, including, but not limited to, primates (e.g. monkeys such a rhesus macaque or cynomolgus), rodents (such as mice, rats and guinea pigs) farm animals, (such as cattle, sheep, pigs, goats, horses, chickens, turkeys, ducks and geese), and domestic mammals (such as dogs and cats). In one embodiment, an IL-2 cytokine is a human IL-2 cytokine.
[0787] As used herein, a "variant IL-2 cytokine" is a cytokine having up to 10 amino acids deleted from the N terminal sequence, in combination with up to 5 amino acid substitutions, deletions or additions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 10 (e.g. 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 5 (e.g. 1, 2, 3, 4 or 5) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 10 (e.g. 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acid deletions from the N-terminal sequence (e.g. within the first 15 amino acids of the wild-type IL-2 sequence in question), in combination with up to 5 (e.g. 1, 2, 3, 4 or 5) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 10 (e.g. 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acid deletions from the N-terminal sequence (e.g. within the first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 5 (e.g. 1, 2, 3, 4 or 5) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 10 (e.g. 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 10 (e.g. 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 10 (e.g. 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 10 (e.g. 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with 1 amino acid substitution elsewhere in the IL-2 cytokine.
[0788] In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 9 (e.g. 1, 2, 3, 4, 5, 6, 7, 8 or 9) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 9 (e.g. 1, 2, 3, 4, 5, 6, 7, 8 or 9) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 9 (e.g. 1, 2, 3, 4, 5, 6, 7, 8 or 9) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 9 (e.g. 1, 2, 3, 4, 5, 6, 7, 8 or 9) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino adds of the wild-type IL-2 sequence in question), in combination with one amino acid substitution elsewhere in the IL-2 cytokine.
[0789] In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 8 (e.g. 1, 2, 3, 4, 5, 6, 7 or 8) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 8 (e.g. 1, 2, 3, 4, 5, 6, 7 or 8) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 8 (e.g. 1, 2, 3, 4, 5, 6, 7 or 8) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 8 (e.g. 1, 2, 3, 4, 5, 6, 7 or 8) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with one amino acid substitution elsewhere in the IL-2 cytokine.
[0790] In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 7 (e.g. 1, 2, 3, 4, 5, 6 or 7) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 7 (e.g. 1, 2, 3, 4, 5, 6 or 7) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 7 (e.g. 1, 2, 3, 4, 5, 6 or 7) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 7 (e.g. 1, 2, 3, 4, 5, 6 or 7) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with one amino acid substitution elsewhere in the IL-2 cytokine.
[0791] In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 6 (e.g. 1, 2, 3, 4, 5 or 6) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 6 (e.g. 1, 2, 3, 4, 5 or 6) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 6 (e.g. 1, 2, 3, 4, 5 or 6) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine.
[0792] In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 5 (e.g. 1, 2, 3, 4 or 5) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 5 (e.g. 1, 2, 3, 4 or 5) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 5 (e.g. 1, 2, 3, 4 or 5) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 6 (e.g. 1, 2, 3, 4, 5 or 6) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with one amino acid substitution elsewhere in the IL-2 cytokine.
[0793] In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 4 (e.g. 1, 2, 3 or 4) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 4 (e.g. 1, 2, 3 or 4) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 4 (e.g. 1, 2, 3 or 4) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 4 (e.g. 1, 2, 3 or 4) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with one amino acid substitution elsewhere in the IL-2 cytokine.
[0794] In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 3 (e.g. 1, 2 or 3) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 3 (e.g. 1, 2 or 3) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 3 (e.g. 1, 2 or 3) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) up to 3 (e.g. 1, 2 or 3) amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with one amino acid substitution elsewhere in the IL-2 cytokine.
[0795] In one embodiment, the variant IL-2 cytokine comprises (or consists of) 1 or 2 amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 4 (e.g. 1, 2, 3 or 4) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) 1 or 2 amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 3 (e.g. 1, 2 or 3) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) 1 or 2 amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with up to 2 (e.g. 1 or 2) amino acid substitutions elsewhere in the IL-2 cytokine. In one embodiment, the variant IL-2 cytokine comprises (or consists of) 1 or 2 amino acid deletions from the N-terminal sequence (e.g. within the first 20, or first 15, or first 10 amino acids of the wild-type IL-2 sequence in question), in combination with one amino acid substitution elsewhere in the IL-2 cytokine.
[0796] Substitutions elsewhere in the IL-2 cytokine are defined further in aspect 44 hereinbelow. Particular IL-2 cytokines and variant IL-2 cytokines are further defined in aspects 40 to 45 hereinbelow.
[0797] The amino acid sequence of the .alpha.-chain of human IL-2 is provided in Seq ID No:327. The amino acid sequence of the .beta.-chain of human IL-2 is provided in Seq ID No:328. The amino acid sequence of the .gamma.-chain of human IL-2 is provided in Seq ID No:239.
[0798] In any of the aspects or concepts herein, an immunocytokine or anti-PDL1 antibody or fragment may have a half-life of at least 4 hours, 5 hours, 6 hours, 7 hours or 8 hours. In another embodiment, the half-life of any of the immunocytokines or anti-PD-L1 antibodies or fragments provided herein is at least 9 hours, or at least 10 hours, or at least 11 hours, or at least 12 hours. In another embodiment, the half-life of any of the immunocytokines or anti-PD-L1 antibodies or fragments provided herein is at least 13 hours, or at least 14 hours, or at least 15 hours, or at least 16 hours. In another embodiment, the half-life of any of the immunocytokines or anti-PD-L1 antibodies or fragments provided herein is at least 17 hours, or at least 18 hours, or at least 19 hours, or at least 20 hours. In another embodiment, the half-life of any of the immunocytokines or anti-PD-L1 antibodies or fragments provided herein is at least 21 hours, or at least 22 hours, or at least 23 hours, or at least 24 hours. In another embodiment, the half-life of any of the immunocytokines or anti-PD-L1 antibodies or fragments provided herein is at least 25 hours, or at least 26 hours, or at least 27 hours, or at least 30 hours. In another embodiment, the half-life of any of the immunocytokines or anti-PD-L1 antibodies or fragments provided herein is at least 32 hours, or at least 34 hours, or at least 36 hours, or at least 40 hours. In one embodiment, the half-life is determined in a mouse model (for example a human PD-L1 knock-in mouse, e.g. as described in Example 22 hereinbelow, or in an immunocompromised mouse xenografted with human T-cells). In another embodiment, the half life is determined in a single dose study in cynomolgus monkeys (e.g. as described in Example 18 or Example 23 hereinbelow). In another embodiment, the half life is determined in an extended single dose study in cynomolgus monkeys (e.g. as described in Example 19 or Example 26 hereinbelow).
Aspect 1a. An immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction: [0799] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and [0800] b) A heavy chain constant region; and wherein the light chain comprises in N- to C-terminal direction: [0801] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3; [0802] d) A light chain constant region, (C.sub.L); [0803] e) Optionally, a linker, (L); and [0804] f) An IL-2 cytokine;
[0805] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to an antigen selected from: an immune checkpoint inhibitor (such as PD-1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, TIM-3 and LAG-3), an immune modulator (such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10, CXCL11 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R), and an immune activator (such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic activity against CXCR3), CD27, CD3 and ICOS (e.g. agonistic activity against ICOS), for example, ICOS, CD137, GITR and OX40).
[0806] In another embodiment, the antigen-binding site that specifically binds to an antigen selected from: an immune checkpoint inhibitor (such as PD-1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, TIM-3 and LAG-3), an immune modulator (such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R), and an immune activator (such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic activity against CXCR3), CD3 and ICOS (e.g. agonistic activity against ICOS), for example, ICOS, CD137, GITR and OX40).
[0807] Any of the embodiments of aspect 1 apply mutatis mutandis aspect 1a. Any of the features or embodiments of aspects 2 to 54 apply mutatis mutandis to aspect 1a. Any of the features of the antibodies or other embodiments or features of concepts 1 to 70 apply mutatis mutandis to aspect 1a.
[0808] In one embodiment, the antigen-binding site specifically binds PD-L1, e.g. hPD-L1. In one embodiment, the PD-L1 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from any one of the anti-PD-L1 antibodies selected from atezolizumab/MPDL3280A (Roche), avelumab/MSB0010718C (Merck), BMS-936559/MDX-1105 (BMS), durvalumab/Medi4736 (Medimmune), KN-035, CA-170, FAZ-053 M7824, ABBV-368, LY-3300054, GNS-1480, YW243.55.S70, REGN3504 and any of the PD-L1 antibodies disclosed in WO2017/034916, WO2017/020291, WO2017/020858, WO2017/020801, WO2016/111645, WO2016/050721, WO2016/197367, WO2016/061142, WO2016/149201, WO2016/000619, WO2016/160792, WO2016/022630, WO2016/007235, WO2015/179654, WO2015/173267, WO2015/181342, WO2015/109124, WO2015/195163, WO2015/112805, WO2015/061668, WO2014/159562, WO2014/165082, WO2014/100079, WO2014/055897, WO2013/181634, WO2013/173223, WO2013/079174, WO2012/145493, WO2011/066389, WO2010/077634, WO2010/036959, WO2010/089411 or WO2007/005874, which antibodies and sequences are incorporated herein by reference.
[0809] In one embodiment, the antigen-binding site specifically binds ICOS, e.g. hICOS. In one embodiment, the antigen-binding site specifically binds ICOS, e.g. hICOS and is an agonist to ICOS, e.g. hICOS. In one embodiment, the antigen-binding site specifically binds ICOS, e.g. hICOS and is an antagonist to ICOS, e.g. hICOS. In one embodiment, the ICOS antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from any one of the anti-ICOS antibodies described in arrangement 5 and arrangement 5a hereinbelow, and any of the anti-ICOS antibodies described in sentences 1 to 102 and sentences 1a to 21a.
[0810] In any of the following embodiments, a particular antigen-binding site specifically binds to a human target. In one embodiment, the antigen-binding site specifically binds an immune checkpoint inhibitor. In one embodiment, the antigen-binding site specifically binds an immune checkpoint inhibitor selected from PD-1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA. In one embodiment, the antigen-binding site specifically binds an immune checkpoint inhibitor selected from TIGIT, CTLA-4, TIM-3 and LAG-3.
[0811] In one embodiment, the antigen-binding site specifically binds PD-1, e.g. human PD-1. In one embodiment, the PD-1 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from pembrolizumab (Keytruda.RTM./MK-3475), nivolumab (Opdivo.RTM./BMS-936558/MDX-1106), MEDI-0680/AMP514, PDR001, Lambrolizumab, BMS-936558, REGN2810, BGB-A317, BGB-108, PDR-001, SHR-1210, JS-001, JNJ-63723283, AGEN-2034, PF-06801591, genolimzumab, MGA-012, IBI-308, BCD-100, TSR-042 ANA011, AUNP-12, KD033, MCLA-134, mDX400, muDX400, STI-A1110, AB011, 244C8, 388D4, XCE853, or pidilizumab/CT-011, or from any one of the anti-PD-1 antibodies described in WO2015/112800 & US2015/0203579 (including the antibodies in Tables 1 to 3), U.S. Pat. Nos. 9,394,365, 5,897,862 and 7,488,802, WO2017/087599 (including antibody SSI-361 and SHB-617), WO2017/079112, WO2017/071625 (including deposit C2015132, hybridoma LT004, and antibodies 6F5/6 F5 (Re), 6F5H1 L1 and 6F5 H2L2), WO2017/058859 (including PD1AB-1 to PD1AB-6), WO2017/058115 (including 67D9, c67D9, and hu67D9), WO2017/055547 (including 12819.15384, 12748.15381, 12748.16124, 12865.15377, 12892.15378, 12796.15376, 12777.15382, 12760.15375 and 13112.15380), WO2017/040790 (including AGEN2033w, AGEN2034w, AGEN2046w, AGEN2047w, AGEN2001w and AGEN2002w), WO2017/025051 & WO2017/024515 (including 1.7.3 hAb, 1.49.9 hAb, 1.103.11 hAb, 1.103.11-v2 hAb, 1.139.15 hAb and 1.153.7 hAb), WO2017/025016 & WO2017/024465 (including antibody A to antibody I), WO2017/020858 & WO2017/020291 (including 1.4.1, 1.14.4, 1.20.15 and 1.46.11), WO2017/019896 & WO2015/112900 & US2015/0210769 (including BAP049-hum01 to BAP049-hum16 and BAP049-Clone-A to BAP049-Clone-E), WO2017/019846 (including PD-1 mAb 1 to PD-1 mAb 15), WO2017/016497 (including MHC723, MHC724, MHC725, MHC728, MHC729, m136-M13, m136-M19, m245-M3, m245-M5 and m136-M14), WO2016/201051 (including antibody EH12.2H7, antibody hPD-1 mAb2, antibody hPD-1 mAb7, antibody hPD-1 mAb9, antibody hPD-1 mAb15, or an anti-PD-1 antibody selected from Table 1), WO2016/197497 (including DFPD1-1 to DFPD1-13), WO2016/197367 (including 2.74.15 and 2.74.15.hAb4 to 2.74.15.hAb8), WO2016/196173 (including the antibodies in Table 5, and FIGS. 1-5), WO2016/127179 (including R3A1, R3A2, R4B3, and R3D6), WO2016/077397 (including the antibodies described in Table 1 of Example 9), WO2016/106159 (including the murine antibodies in Table 3 of Example 2 and the humanised antibodies in Tables 7, 8 and 9 of Example 3), WO2016/092419 (including C1, C2, C3, EH12.1, mAb7-G4, mAb15-G4, mAb-AAA, mAb15-AAA), WO2016/068801 (including clone A3 and its variants and the other antibodies described in FIGS. 1 to 4), WO2016/014688 (including 10D1, 4C10, 7D3, 13F1, 15H5, 14A6, 22A5, 6E1, 5A8, 7A4, and 7A4D and the humanised antibodies of Examples 9/10), WO2016/015685 (including 10F8, BA08-1, BA-08-2 and 15H6), WO2015/091911 & WO2015/091910 (including the anti-canine PD-1 antibodies in Examples 2, 3 and 4), WO2015/091914 (including the anti-canine PD-1 antibodies in Table 3), WO2015/085847 (including mAb005, H005-1 to H005-4), WO2015/058573 (including cAB7), WO2015/036394 (including LOPD180), WO2015/035606 (including the antibodies in Table 1 of Example 2, in Tables 14, 15 and 16 of Example 7 and in tables 20, 21 and 22 of Example 11), WO2014/194302 (including GA2, RG1B3, RG1H10, RG2A7, RG2H10, SH-A4, RG4A6, GA1, GB1, GB6, GH1, A2, C7, H7, SH-A4, SH-A9, RG1H11, and RG6B), WO2014/179664 (including 9A2, 10611, 6E9, APE1922, APE1923, APE1924, APE1950, APE1963 and APE2058), WO2014/206107 (including clone 1, 10, 11, 55, 64, 38, 39, 41 and 48), WO2012/135408 (including h409A11, h409A16, and h409A17), WO2012/145493 (including antibodies 1E3, 1E8, 1H3 and h1H3 Var 1 to h1H3 Var 14), WO2011/110621 (including antibody 949 and the modified versions disclosed in FIGS. 1 to 11), WO2011/110604 (including antibody 948 and the modified versions disclosed in FIGS. 3 to 11), WO2010/089411 (including CNCM deposit number 1-4122, 1-4080 or 1-4081), WO2010/036959 (including the antibodies in Table 1 of Example 1), WO2010/029435 & WO2010/029434 (including clones 2, 10 and 19), WO2008/156712 (including hPD-1.08A, hPD-1.09A, h409A11, h409A16 and h409A17 and the antibodies described in Example 2, Table H, Example 4 and table IV), WO2006/121168 (including clones 17D8, 4H1, 5C4, 4A11, 7D3, 5F4, and 2D3), WO2004/004771 or WO2004/056875 (including PD1-17, PD1-28, PD1-33, PD1-35, PD1-F2 and the Abs described in Table 1); the sequences and features of the anti-PD-1 antibodies are incorporated herein by reference.
[0812] In one embodiment, the antigen-binding site specifically binds CTLA-4, e.g. hCTLA-4. In one embodiment, the CTLA-4 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from ipilimumab (MDX-010, CAS No. 477202-00-9), tremelimumab (ticilimumab/CP-675,206), antibody clone 2F1, clone 1F4 (Abnova Corporation), clone 9H10 (EMD Millipore), clone BNU3 (GeneTex), clone 1 E2, clone AS32 (Lifespan Biosciences) clone A3.4H2.H12 (Acris Antibodies), clone 060 (Sino Biological), clone BU5G3 (Creative Diagnostics), clone MIH8 (MBL International), clone A3.6B10.G1, or clone L3D10 (BioLegend) or from any one of the anti-CTLA-4 antibodies described in WO2017/087588 (ISVs disclosed in FIG. 2), WO2017/084078 (clones C2, C4, C10, C11, C12 and C13, and FIGS. 4-7), WO2016/196237 (including AGEN1884w, AGEN2041w, the sequences in FIGS. 19A, 19B and Tables 1-6), WO2016/130986 & WO2016/130898 (including E8, F7 and the Abs described in Table 4), WO2016/015675 (including hybridoma LT001 and antibodies 8D2, 8D2H1L1, 8D2H2L2, 8D2H3L3, 8D2H2L15 and 8D2H2L17), WO2012/120125 (including 3B10, 8H5, and the Abs identified in Examples 1, 2, 3 and 5), WO2010/097597 (including JMW-3B3 and the variants and fragments disclosed), WO2009/100140 (including 10D1, 1H5, 3A4, 6C10 and the antibodies described in FIGS. 1 to 6), WO2007/008463 & WO2006/101692 & WO2006/101691 & WO2006/048749 & WO2005/09238, (including 4.1.1, 4.8.1, 4.10.2, 4.13.1, 4.14.3, 6.1.1, 11.2.1, 11.6.1, 11.7.1, 12.3.1.1, 12.9.1.1, and 10D1), WO2006/096491 (including ATCC Deposit No. 11.2.1 11.2.1.4 PTA-5169 and 4.1.1 4.1.1.1 PTA-5166), WO2006/066568 (including TGN2122.C, TGN2422.C, 4.8H10H5 and 4.3F6B5 and the antibodies described in tables 3 to 14), WO2006/029219 (including L3D10, L1B11, K4G4, KM10, and YL2), WO2004/029069 (including ATCC deposit number PTA-4537), WO01/54732 (including antibodies 25, 26, 27, 29, 33, 34, 35, 36 and 38), WO01/14424 (including 3A4, 9A5, 2E2, 2E7, 4B6, 4E10, 5C4, 5G1, 11E8, and 11G1 and the antibodies identified in Examples 3 and 4 and table 3) and WO00/37504 (including 3.1.1, 4.1.1, 4.8.1, 4.10.2, 4.13.1, 4.14.3, 6.1.1, 11.2.1, 11.6.1, 11.7.1, 12.3.1.1, and 12.9.1.1); the sequences and features of the anti-CTLA-4 antibodies are incorporated herein by reference.
[0813] In one embodiment, the antigen-binding site specifically binds TIGIT, e.g. human TIGIT. In one embodiment, the TIGIT antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from RG-6058 (MTIG-7192A) or from any one of the anti-TIGIT antibodies described in WO2017/053748 (including 1A4, 1D3, 4A3, 10A7, 4.1D3.Q1E, h10A7.K4G3, 4.1D3 and the other antibodies described in Examples 1 and 2), WO2017/037707 (including VSIG9#1 and 258-csl#4), WO2017/030823 (including 14D7, 26610 and humanized versions in Example 3), WO2016/191643 (including 313R11, 313R12, 313R14, 313R19, 313R20, ATCC PTA-122180 and ATCC PTA-122181), WO2016/106302 (including 14B2, 13E6, 6F9, 11G11, 10C9, 16F6, 11C9, 27A9, 10D7, 20G6, 24E8, 24G1, 27F1, 15A6, 4E4, 13D1, 9B11, 10B8, 22G2, 19H2, 8C8, 17G4, 25E7, 26D8 and 16A8), WO2016/028656 (including 14A6, 28H5 or 3106 and humanized versions from Example 6), and WO2009/126688 (US2013/0251720, including 10A7 and 1F4); the sequences and features of the anti-TIGIT antibodies are incorporated herein by reference.
[0814] In one embodiment, the antigen-binding site specifically binds TIM-3, e.g. human TIM-3. In one embodiment, the TIM-3 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from F38-2E2 (BioLegend), clone 2E2 (Merck Millipore), clone 6B6E2, clone 024 (Sino Biological) clone 344801 (R&D Systems), clone E-18, clone H-191 (Santa Cruz Biotechnology), or clone 13A224 (United States Biological), TSR-022 (Tesaro) or from any one of the anti-TIM-3 antibodies described in WO2017/079115 (including anti-TIM3 antibodies listed in tables 30-38), WO2017/055404 (including PD1TIM3-0389, PD1TIM3-0168, PD1TIM3-0166, TIM3-0038, TIM3-0018, TIM3-0028, TIM3-0438--Table C), WO2017/031242 (Table 10), WO2016/179194 (including antibodies in FIG. 1b, including mAb F38-2E2 and 2E2), WO2016/171722 (including 344823 and antibodies from the hybridomas 7D11, 10G12, 11G8, 8B.2C12 and 25F.1D6), WO2016/161270 (including APE5137 and APE5121), WO2016/111947 (including mAb5, mAb13, mAb15, mAb17, mAb21, mAb22, mAb26, mAb27, mAb48, mAb58 and mAb91), WO2016/071448 (including TIM3-0016, TIM3-0018, TIM3-0021, TIM3-0022, TIM3-0026, TIM3-0028, TIM3-0030, TIM3-0033, TIM3-0038, TIM3-0433, TIM3-0434, TIM3-0438 and TIM3-0443), WO2016/068802 (including 1B9, 1H9, 1H10, 2C7, 2F4, 2G6, 1D9, 1F4 and 2C8--FIGS. 1, 2 & 3), WO2016/068803 (including A3, B10, G6, G7, G9, A11 and A11_gl--FIGS. 1, 2 & 3), WO2015/117002 (including ABTIM3, ABTIM3-hum02, ABTIM3-hum05, ABTIM3-hum06, ABTIM3-hum09, ABTIM3-hum10, ABTIM3-hum12, ABTIM-hum01, ABTIM-hum04, ABTIM3-hum07, ABTIM3-hum08, ABTIM3-hum04, ABTIM3-hum21, ABTIM3-hum03, ABTIM3-hum11 and antibodies listed in Table 9), WO2015/048312 (including 5D12), WO2014/022332 (including 2C12), WO2013/006490 (including antibodies in Table 1), WO2011/155607 (including 512, 644, 4545, 4177, 8213, 344823 and 34823), WO2003/063792 (including antibody 8B.2C12 and 25F.1D6), WO2017/019897 (including antibody molecules disclosed in Tables 1-4, including ABTIM3, ABTIM3-hum20, ABTIM3-hum22 and ABTIM3-hum23), WO2016/079050 & WO2016/079050 (including 11m3_0022, Tim3_0016, Tim3_0018, Tim3_00122, Tim3_0022, Tim3_0021, Tim3_0028, Tim3_0026, Tim3_0033, Tim3_0038, Tim3_0030, 1.7. E10, F38-2EL and 27-12E12); the sequences and features of the anti-TIM-3 antibodies are incorporated herein by reference.
[0815] In one embodiment, the antigen-binding site specifically binds LAG-3, e.g. human LAG-3. In one embodiment, the LAG-3 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from antibody clone 17B4 (Enzo Life Sciences), or clone 333210 (R&D Systems), or clone 14L676 (United States Biological), or C9B7W (PharMingen), or 11E, or IM0321, or mAb C9B7W (BioXcell) or from any one of the anti-LAG-3 antibodies described in WO95/30750, WO2004/078928, WO2008/132601 (including IMP731 Lag-3 Ab, IMP321, A9H12 Lag-3 mAb and 31G11), WO2010/019570 (including 25F7, 26H10, 25E3, 8B7, 11F2 and 17E5), WO2014/140180 (including H5L7, H5L7BW, IMP731 and antibodies in Tables 3 & Table 7), WO2014/179664 (including APE03109), WO2014/008218 (including Lag3.1, Lag3.5, Lag3.6, Lag3.7 and Lag3.8), WO2015/042246, WO2015/116539 (including BMS-986016), WO2015/138920 (including BAP050-hum01 to BAP050-hum20, huBAP050(Ser), BAP050-hum01-Ser to BAP050-hum20-Ser, BAP050-Clone-F, BAP050-Clone-G, BAP050-Clone-H, BAP050-Clone-I, BAP050-Clone-J, BAP050 and BAP050-chi), WO2015/198312, WO2016/028672 (including Ab1, Ab2, Ab3, Ab4, Ab5, Ab6, Ab7, Ab8 and Ab9), WO2016/126858, WO2016/200782 (including LAG-3 mAb1, to LAG-3 mAb6), WO2017/015560 (including L32D10, L3E3, L3C5, L35D4, L35G6, L33H11, L32A9, L32A4, L3A1 and the antibodies listed in Table 3), WO2017/062888 (including mAb1, H4H15477P, H4H15483P, H4H15484P, H4H15491, H4H17823P, H4H17826P2, H4H17828P2, H4sH15460P, H4sH15462P, H4sH15463P, H4sH15464P, H4sH15466P, H4sH15467P, H4sH15470P, H4sH15475P, H4sH15479P, H4sH15480P, H4sH15482P, H4sH15488P, H4sH15496P2, H4sH15498P2, H4sH15505P2, H4sH15518P2, H4sH15523P2, H4sH15530P2, H4sH15555P2, H4sH15558P2, H4sH15567P2 and H4H17819P), WO2017/019894, WO2017/037203 (including 8E2, 13E2, 34F4, 1764 and IMP761), WO2017/087589 (including 11609) or WO2017/087901; the sequences and features of the anti-LAG-3 antibodies are incorporated herein by reference.
[0816] In one embodiment, the antigen-binding site specifically binds VISTA, e.g. human VISTA. In one embodiment, the VISTA antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from any one of the anti-VISTA antibodies described in WO2016/207717 & WO2015/097536 (including VSTB50, VSTB53, VSTB60, VSTB95, VSTB112, VSTB116, VSTB174, VSTB175, VSTB149, VSTB140 and the antibodies in Table 1A and Examples 7 and 8) and WO2014/190356 (including clone 2D3 and 18C3); the sequences and features of the anti-VISTA antibodies are incorporated herein by reference.
[0817] In one embodiment, the antigen-binding site specifically binds an immune modulator. In one embodiment, the antigen-binding site specifically binds an immune modulator selected from BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10, CXCL11 and CD155, or from BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10 and CD155. In one embodiment, the antigen-binding site specifically binds an immune modulator selected from GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R.
[0818] In one embodiment, the antigen-binding site specifically binds GARP, e.g. human GARP. In one embodiment, the GARP antigen-binding site comprises the CDRH1, CDRH2, CDR3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from G14D9, Plato-1, 272, G6, 50 G10 or 7611 or from any of one of the anti-GARP antibodies described in WO2007/113301 & WO2015/015003 (including MHGARP8, LHG-10, LHG-10-D, LHG-10.3-D, LHG-10.4-D, LHG-10.5-D, LHG-10.6-D, LHG-10.3, LHG-10.4, LHG-10.5, LHG-10.6, 27E10, MHGARP1, MHGARP2, MHGARP3, MHGARP4, MHGARP5, MHGARP6, MHGARP7 and MHGARP9), WO2017/051888 (including 110F, 105F, c151D, c198D, h198D, h151D, h151D-H1L1 and h198D-H3L4); the sequences and features of the anti-GARP antibodies are incorporated herein by reference.
[0819] In one embodiment, the antigen-binding site specifically binds SIRP.alpha., e.g. human SIRP.alpha.. In one embodiment, the SIRP.alpha. antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from ED9 (ThermoFisher), or 602411 (Novus Biologicals), or from any one of the anti-SIRP.alpha. antibodies described in WO97/48723, WO00/24869 (including 10C4), WO00/66159 (including ED9 and ED17), WO01/40307, WO02/092784 (including SE5A5, SE7C2 and SE12C3), WO2004/108923 (including SE12C3 and 2F34), WO2009/046541 (including P84), WO2011/076781, WO2012/172521, WO2012/040207 (including SE5A5 and mouse P84), WO2013/056352 (including 29-AM4-5, Ab AM4-5, AM5-1, AM5-3, AM5-5, AM5-6, SIRPalpha-AM3-35, AM4-1, SIRP29-AM3-35, SIRP29-AM4-5, SIRP29-AM4-1, 29-AM2-2, 29-AM4-4, 29-AM4-1, 29-AM4-5, 29-AM3-35 and SIRP29-AM3-63), WO2016/063233, WO2016/205042 (including P362) or WO2015/138600 (including KWAR23); the sequences and features of the anti-SIRP.alpha. antibodies are incorporated herein by reference.
[0820] In one embodiment, the antigen-binding site specifically binds CXCR4, e.g. human CXCR4. In one embodiment, the CXCR4 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region of ulocuplumab/BMS-936564, clone 44717.111 or PF-06747143 or from any one of the anti-CXCR4 antibodies described in WO97/49424 (including MAB12G5), WO99/50461, WO01/42308, WO03/066830 & WO2003/066830 (including Ab124 and Ab125), WO2004/059285 (including ALX40-4C), WO2006/089141 (including mAbs 2N, 6R, 18, 19, 20, 33 and 48), WO2007/005605, WO2008/142303 (including MAB170, MAB171, MAB173 and MAB172), WO2008/060367 & WO2013/071068 & WO2015/015401 (including BMS-936564/MDX-1338), WO2009/140124 (including antibody I, II, III, IV and V), WO2009/117706 (including 701, 708, 716, 717, 718 and 4G10), WO2011/161266 (including 4CXCR100, 4CXCR103, 4CXCR104, 4CXCR101, 4CXCR238D2 and 4CXCR238D4), WO2011/098762 (including C-9P21 (Table 1), B-1M22 (Table 2), C1124 (Table 3), D-1K21 (Table 4) and 9N10 (Table 5)), WO2012/175576, WO2013/013025 (including 2A4, 6C7, 4C1, 7C8, 5C9 and 5E1), WO2013/017566 (including Mab 427aB1 and 515H7), WO2013/017562 (including 1-3859 Mab and 515H7), WO2015/069874 (including antibodies corresponding to Seq ID numbers 25 and 29), WO2015/015401 (including 12A11, 6B6, 3G10, m3G10.hIgG1, m3G10.hIgG4, h3G10.A57.hIgG1, h3G10.A57.A58A.hIgG1, h3G10.1.91.A58A.hIgG1, h3G10.1.91.A58B.hIgG1 and h3G10.2.37.2.72.hIgG1), WO2016/156570 (including 281F12, 281A6 and 281D4), WO2016/109872 (including antibodies listed in tables 1, 2, 9 & 12, M3-114-6H, AM4-272-6H, AM3-523-6H, AM4-272, AM3-114, AM3-523, AM4-746 and AM4-1121), WO2017/071625, WO2012/175576, WO2010/125162 & WO2012/055980 & WO2011/121040 & WO2010/037831 (including c414H5 (414H5), c515H7 (515H7) and 301aE5), WO2009/138519 (including ALX40-4C, 238D2, 238D4, 237B5 antibodies and sequences listed in table 1, table 1.1, table A-I, table B-1.1 & B-5), WO2011/042398 (including 238D2 and 238D4), WO2011/083140 (including those disclosed in Tables C-2, C-3, C-4 & C-5, FIG. 2 and ALX-0651, 15H3, 10E12, 10G10, 238B6, 10E9, 281E10, 10A10, 14A2 and 15A1) or WO2011/083141; the sequences and features of the anti-CXCR4 antibodies are incorporated herein by reference.
[0821] In one embodiment, the antigen-binding site specifically binds BTLA, e.g. hBTLA. In one embodiment, the BTLA antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from antibody clone 167, clone 2G8, clone 4C5 (Abnova Corporation), clone 4B8 (antibodies-online), clone MIH26 (Thermo Scientific Pierce Antibodies), clone UMAB61 (OriGene Technologies), clone 330104 (R&D Systems), clone 1B4 (Lifespan Biosciences), clone 440205, clone 5E7 (Creative Diagnostics) or from any one of the anti-BTLA antibodies described in WO2016/176583 (including clone 6F4), WO2011/014438 (including 8D5, 8A3, 20H4, 21H6, 15C5, 19A7 and 4C7), WO2010/106051 (including CNCM deposit number 1-4123) and WO2008/076560 (including 164, E4H9, 3C2, 3C2a, 6A5, 11E2, E8D9, 10H6 and 4C9 as detailed in Example 2); the sequences and features of the anti-BTLA antibodies are incorporated herein by reference.
[0822] In one embodiment, the antigen-binding site specifically binds hVEM, e.g. human hVEM. In one embodiment, the HVEM antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from any one of the anti-HVEM antibodies described in WO2008/083169 (including LBH1); the sequences and features of the anti-BTLA antibodies are incorporated herein by reference.
[0823] In one embodiment, the antigen-binding site specifically binds CSF1R. In one embodiment, the CSF1R antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from any one of the anti-CSF1R antibodies described in WO2009/026303 (including 1.2, 1.109, 2.360 and 1.2.SM and the antibodies in FIGS. 1 and 2), WO2009/112245 (including CXIIG6), WO2011/070024 (including Mab 2F11, 2E10, 2H7 and 1G10, and their derivatives), WO2011/107553 (including 7H5.2G10/DSM ACC2922), WO2011/123381 (including antibody 1 and antibody 2), WO2011/131407 (including 7G5.3B6/DSM ACC2921), WO2011/140249 (including 0301, 0302, and 0311 their derivatives and the antibodies in tables 2, 3 and 5), WO2013/169264 & WO2014/036357 & WO2016/106180 & WO2016/168149 (including huAb1 to huAb16), WO2012/110360 & WO2013/057281 (including CXIIG6, H19K12, H27K5 and H27K15 and the humanised antibodies of tables 1 and 2), WO2013/087699 (including 9D11.2E8 and 10H2.2F12), WO2014/072441 (including H27K15), WO2014/173814 & WO2013/132044 (including Mab 2F11, Mab 2E10, Mab 2H7, Mab 1G10 and sc2-4A5 and the antibodies in Table 3 and 3b), WO2015/028455 & WO2015/028454 (including Ab535, Ab969, and derivatives, e.g. Ab969.g2), WO2015/036511 & WO2016/207312 (including 2F11, 2E10 and the derivatives described in embodiment 33) and WO2017/049038 (including ALM-423 and the antibodies listed in Table 2); the sequences and features of the anti-CSF1R antibodies are incorporated herein by reference.
[0824] In one embodiment, the antigen-binding site specifically binds CD39, e.g. human CD39. In one embodiment, the CD39 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from BY40, BY12, BA54g (Biolegend), BU61 (Santa Cruz Biotech), A1 (Ebiosciences), AC2 (Immunotech), 22A9 (Abcam), 24DMS1 or any one of the anti-CD39 antibodies described in WO96/32471, WO00/04041, WO01/10205 (including CD39L4), WO2009/09547 (including CNCM-I-3889/BY40), WO2014/169255, WO2012/085132 (including antibodies VY12, BY40 and BA54g), WO2016/073845 (including R29-5-13A, R29-5-71A, R29-5-165C and R29-9-8B), WO2017/089334 (including 1-391, 1-392 and antibodies produced from hybridomas 1-3889 and CNCM 1-41171) and WO2009/095478; the sequences and features of the anti-CD39 antibodies are incorporated herein by reference.
[0825] In one embodiment, the antigen-binding site specifically binds CD40, e.g. human CD40. In one embodiment, the CD40 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from BMS3h-56-269, CP-870,893, dacetuzumab, SEA-CD40, ADC-1013, R07009789 and Chi Lob 7/4, or from any one of the anti-CD40 antibodies described in WO2017/059243, WO2017/059196, WO2017/040932, WO2017/040566, WO2017/004016, WO2017/004006, WO2016/196314, WO2016/028810, WO2016/023960, WO2016/023875, WO2015/134988, WO2015/091853, WO2014/070934, WO2014/065403, WO2014/065402, WO2014/04298, WO2013/164789, WO2013/034904, WO2012/149356, WO2012/145673, WO2012/125569, WO2012/111762, WO2012/075111, WO2012/065950, WO2012/041635, WO2011/123489, WO2010/123012, WO2010/104761, WO2010/121231, WO2009/062125, WO2010/104747, WO2010/104748, WO2010/104749, WO2010/024676, WO2009/094391, WO2009/062054, WO2008/091954, WO2007/130493, WO2007/129895, WO2007/124299, WO2007/053767, WO2007/053661, WO2006/128103, WO2006/073443, WO2005/063981, WO2005/063289 (US2012/0263732), WO2005/044855, WO2005/044306, WO2005/044294, WO2005/044307, WO2005/044304, WO2005/044854, WO2005/044305, WO03/040170 (U.S. Pat. Nos. 7,563,442B, 7,618,633B, 7,338,660B, 7,288,251B, 7,626,012B, 8,388,971B, 2013/0024956), WO03/029296, WO02/088186, WO01/83755, WO02/28905, WO02/28480, WO02/28481, WO02/28904, WO01/37870, WO01/16180, WO00/75348 WO99/61057, WO99/42075, WO97/31025, WO95/17202 and WO95/09653; the sequences and features of the anti-CD40 antibodies are incorporated herein by reference.
[0826] In one embodiment, the antigen-binding site specifically binds CD73, e.g. human CD73. In one embodiment, the CD73 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from 1E9 (Santa Cruz Biotechnology), AD2, 7G2, 4G4 or from any one of the anti-CD73 antibodies described in WO2017/064043 (including 7H10, 12F9, 15D7, 4B11, 11D9 and 9D2), WO2016/081748 (including 4C3, 7A11, 6E11, 5F8, 4C3, 11F11, 11A6, CD73.4-1, CD73.4-2, CD73.3, 11F11-1, 11F11-2, 11F11, 4C3-1, 4C3-2, 4C3-3, 4D4, 10D2-1, 10D2-2, 11A6, 24H2, 5F8-1, 5F8-2 and 5F8-3), WO2016/131950 (including 11E1, 8C7, 3C12 and 6E1), WO2016/075176 (including MEDI9447, clone 10.3 and clone 2C5) & WO2016/075099 (including CD730004, CD730008, CD7300011, CD730021, CD730042, CD730046, CD730047, CD730068 and CD730069), WO2016/055609 (including 11E1, 6E1, 3C12 and 8C7); the sequences and features of the anti-CD73 antibodies are incorporated herein by reference.
[0827] In one embodiment, the antigen-binding site specifically binds CD96, e.g. human CD96. In one embodiment, the CD96 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region of 6A6, or NK92.39 (E bioscience), 1C8, 3H8, MAA6359 or from any one of the anti-CD96 antibodies described in WO2008/073316, WO2009/007124, WO2013/184912, WO2014/089169, WO2014/149310 (including antibody 3.3), WO2015/024060 or WO2015/024042, WO2015/024060 (including mAb 3.3); the sequences and features of the anti-CD96 antibodies are incorporated herein by reference.
[0828] In one embodiment, the antigen-binding site specifically binds CXCR2, e.g. human CXCR2. In one embodiment, the CXCR2 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from any one of the anti-CXCR2 antibodies described in WO2015/169811 (including HY29 and HY29GL), WO2014/170317 (including CX2-Mab#1 to #19), WO2012/062713, WO2013/168108 (including 163D2-127D1, 163E3-127D1, 163E3-54B12, 163D2-54B12, 2B2-163E3, 2B2-163D2, 97A9-2B2, 97A9-54B12, 127D1-163D2, 127D1-163E3, 2B2-97A9, 54B12-163D2, 54B12-163E3, 163D2-2B2, 163E3-2B2, 127D1-97A9, 54B12-97A9, 97A9-127D1 and derivatives thereof), WO2009/117706 (including 48311.211, 5E8/CXCR2, clone 19 and derivatives thereof), WO2009/120186 (including RII115, 48311 and derivatives thereof) and WO2002/26249; the sequences and features of the anti-CXCR2 antibodies are incorporated herein by reference.
[0829] In one embodiment, the antigen-binding site specifically binds CD200, e.g. human CD200. In one embodiment, the CD200 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from DX-109, samalizumab/ALXN-6000, 111-200.7 or from any one of the anti-CD200 antibodies described in WO99/24565 (including M3B5 and the antibodies in Examples 4 and 5), WO02/11762 (including 366 and the antibodies in the Examples), WO2004/060295 (US2004/0213783), WO2004/078938 (including scFv-9), WO2006/020266 (U.S. Pat. No. 8,840,885B2, including CG1R3A10, cG2aR3A10, cG2aR3B7, dGIR3A5, dGIR3B5, and dGIR3B10 and the antibodies described in FIGS. 9A-9C, FIGS. 21A and 216), WO2007/084321 (U.S. Pat. No. 8,709,415B2, including ALXN5200, hB7VH3VL2, C2aB7G1, C2aB7G2/G4, V3V2-G1 and V3V2-G2/G4), WO2009/014745 (including OX90mG2a (FIG. 10), OX90NE and OX90NE-AG), and WO2011/100538 & US2013/0189258 (including Antibody 1 and Antibody 2); the sequences and features of the anti-CD200 antibodies are incorporated herein by reference.
[0830] In one embodiment, the antigen-binding site specifically binds CCR4, e.g. human CCR4. In one embodiment, the CCR4 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from mogamulizumab, KM3060 (see Niwa et al., 2004, Cancer Research 64, 2127-2133), and KW-0761 (see Ishida et al., Annals of Oncology 2008, vol 19, supplement 4, 513) or from any one of the anti-CCR4 antibodies described in WO2016/178779 & WO2016/057488 (including mAb2-3, 1-44, 1-49, 2-1 and 2-2), WO2015/179236 (including KW-0761), WO2013/166500 (including mAb1567, c1567, h1567, mAb 1-4 and 2-3 and the antibodies in Examples 6 and 13), WO2012/076883 (including antibodies 208, 306, 308, 406, 501, 503, 601, 603 and 803--Tables 1-9), WO2010/142952 (including 17G, 9E, 11F, 9E10, 9E103 and 9E1D--see Tables 1-16), WO2009/086514 (including mAb1567 and the humanised mAbs in Example 14), WO2005/035582 (including the DG44/CCR4 antibody and the Ms705/CCR4 antibody (FERM BP-8467)), WO2005/053741 & WO01/64754 (U.S. Pat. Nos. 6,989,145B, 7,666,418B, 8,197,814B, 8,632,996B, including KM2160 (FERM BP-10090), KM2760 (FERM deposit BP-7054)), WO2003/018635 (including KM2160, KM8759 (FERM BP-8129) and KM8760 (FERM BP-8130), WO00/42074 (U.S. Pat. Nos. 6,488,930B, 7,138,117B, including 2B10, 10E4, 1G1 and the antibodies deposited as ATCC accession number HB-12624 and HB-12625) and WO00/41724 (U.S. Pat. Nos. 6,881,406B, 6,245,332B, including 1G1 and the antibody deposited under ATCC accession number HB-12624); the sequences and features of the anti-CCR4 antibodies are incorporated herein by reference.
[0831] In one embodiment, the antigen-binding site specifically binds CXCL9, e.g. human CXCL9. In one embodiment, the CXCL9 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from mAb 392-100 or AF392 (R&D Systems).
[0832] In one embodiment, the antigen-binding site specifically binds CXCL10, e.g. human CXCL10. In one embodiment, the CXCL10 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region of mAb266 (R & D systems) or from any one of the anti-CXCL10 antibodies described in WO017/8708 (including CR.G (IP-10) (IgG1) (PharMingen) and IP-10 (IgG)(A.Luster), WO02/15932, WO03/006045, WO2004/082714, WO2004/045525, WO2004/045526, WO2004/101511 (including antibodies in table 1 and AIP12, HuAIP12, MuAIP12, AIP13, HuAIP13, MuAIP13, AIP6, AIP8, AIP14, AIP18, AIP21, AIP22, AIP5 and AIP17) WO2005/060457 (including AIP5, AIP6, AIP8, AIP10, AIP12, AIP13, AIP14, AIP17, AIP18, AIP21, AIP22, AIP32 and AIP36), WO2005/011605, WO2005/023201, WO2005/058815 (including 1D4, 1E1, 2G1, 3C4, 6A5, 6A8, 6610, 7C10, 8F6, 10A12 and 10A12S13C4), WO2005/084708, WO2006/039819, WO2006/118085, WO2008/047486, WO2008/044824 (including antibodies #124, #31, #28, #43 and #137), WO2008/106200, WO2009/023566, WO2012/149320 (including MSX-1100 and 6A5), WO2014/003742 (including the antibody of Example 14), WO2013/170735, WO2014/189306, WO2015/063187; the sequences and features of the anti-CXCL10 antibodies are incorporated herein by reference.
[0833] In one embodiment, the antigen-binding site specifically binds CD155, e.g. human CD155. In one embodiment, the CD155 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from clone SKII.4 (BioLegend).
[0834] In one embodiment, the antigen-binding site specifically binds an immune activator. In one embodiment, the antigen-binding site specifically binds an immune activator selected from CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic activity against CXCR3), CD3 and ICOS (e.g. agonistic activity against ICOS). In one embodiment, the antigen-binding site specifically binds an immune activator selected from ICOS, CD137, GITR and OX40.
[0835] In one embodiment, the antigen-binding site specifically binds CD137, e.g. hCD137. In one embodiment, the CD137 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from urelumab, BMS-663513, PF-05082566 (Pfizer), 1D8 and 3E1, 4B4 (BioLegend 309809), H4-1BB-M127 (BD Pharmingen 552532), BBK.2 (Thermo Fisher M S621PABX), 145501 (Leinco Technologies B591), the antibody produced by cell line deposited as ATCC No. HB-11248 (U.S. Pat. No. 6,974,863) or XmAb-5592, or from any one of the anti-CD137 antibodies described in WO2017/04945, WO2016/134358, WO2015/179236, WO2012/177788, WO2012/145183, WO2012/032433, WO2009/135019, WO2005/035584, U.S. Pat. No. 6,974,863, WO2004/055513 and WO2004/010947; the sequences and features of the anti-CD137 antibodies are incorporated herein by reference.
[0836] In one embodiment, the antigen-binding site specifically binds GITR, e.g. hGITR. In one embodiment, the GITR antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from MK4166, TRX518, TRX385, MAB689 (R & D Systems), YGITR765 (Novus Biologicals) or 1D8 (Novus Biologicals), or from any one of the anti-GITR antibodies described in WO2015/187835 (including 28F3, 3C3-1, 3C3-2, 2G6, 8A6, 9G7-1, 9G7-2, 14E3, 19H8-1, 19H8-2, 19D3, 18E10, and 6G10), WO2015/184099 (including 1042-7, 32-15, 1039-45, 1333-21, 231-1039-45, 231-32-15, Hum231#1, Hum231#2, m6C8, pab1964, to pab1973, pab1975 to pab1977, pab1979 to pab1981, pab1983, pab2159, pab2160, pab2161 and the antibodies in tables 1 and 2), WO2015/031667 (including antibodies Abl to Ab59 in table 1), WO2015/026684 (including an antibody with a CDR sequence of Seq ID 1-66), WO2013/039954 (including, 2155, 1718, 1649, 1362, 954, 827, 698, 706 and antibodies listed in Tables 1 & 3), WO2011/051726 (including antibodies containing CDRs a-f listed on page 17), WO2011/028683 (including antibodies 36E5, 61F6, 61G6, 3D6, 6H6, 1D8, 17F10, 35D8, 49A1, 9E5, 31H6 and antibodies from hybridomas PTA-9889, PTA-9890, PTA-9891, PTA-9892, PTA-9893, PTA-10286, PTA-10287, PTA-10288, PTA-10289, PTA-10290, and PTA-10291), WO2009/009116 (including antibody 2F8), WO2007/133822 (including antibodies listed in Table 1), WO2006/105021 (including 6C8, 2F8, HuN6C8-Agly, HuQ6C8-Gly, and HuQ6C8-Agly), WO2006/050172 & WO2004/084942 (including DTA-1), WO03/006058 (including anti-GITR/TNFRSF18# AF524), WO2016/054638 (including mAb #1-81, #3-167, #5-139, #7-192, #10-116, #11-126, #12-46, #13-169, #14-182, #15-68 and #17-60), WO2016/196792 (including 6G10, 28F3, 19D3, 18E10, 3C3, 2G6, 8A6, 9G7, 14E3 and 19H8), WO2017/087678 (including 28F3, 19D3, 18E10, 30-1, 3C3-2, 2G6, 8A6, 9G7-1, 9G7-2, 14E3, 19H8-1, 19H8-2 and 6G10); the sequences and features of the anti-GITR antibodies are incorporated herein by reference.
[0837] In one embodiment, the antigen-binding site specifically binds OX40, e.g. hOX40. In one embodiment, the OX40 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from GSK3174998, L106 BD (Pharmingen Product #340420), ACT35 (Santa Cruz Biotechnology, Catalog #20073), MOXR0916, MEDI-6469, MEDI-0562, 9612 (Weinberg, A. D., et al., Immunother 29, 575-585 (2006)), the humanised anti-OX40 Ab described in Morris et al., Mol Immunol. May 2007; 44(12):3112-3121, or from any one of the anti-OX40 antibodies described in WO2017/077085 (including SAPS, SAP28.2, SAP15.3, SAP29-50, SAP25-29 and SAP29-23 and humanised versions described in Examples 4 and 5), WO2017/063162 (including O3, O19, O21 and the affinity matured version in Example 5--Table 2, including 21#H28H33, 21#H65, 21#H96, 21#VHnew-L80, 21#H96-L80), WO2017/050729 (including SP197), WO2017/021912 & WO2017/021910 (including ANTIBODY 106-222, OX86, and the antibodies described in FIGS. 6 and 7), WO2016/200836 & WO2016/200835 (including MOXR0916/1A7.gr1 IgG1), WO2016/196228 (including 3F4, 1466-1, 1466-2, 23H3, 18E9, 8V11, 20B3, 20C1, 6E1-1, 6E1-2, 14A2, 14A2-1, 14A2-2, L106, OX40.1, OX40.5, OX40.8, OX40.6, and OX40.16 and OX40.21--FIGS. 1 to 10), WO2016/179517 (including 11D4, pab1949, pab1949-1, pab2044, pab2193-1, Tables 1 to 4), WO2016/057667 (including 9612 and OX40mAb24), WO2015/153513 (including 3C8, 1D2, 1A7 and their variants described in the sequence listing, including A1A7.gr1 and 3C8.gr.5, the antibodies described in FIG. 1), WO2014/148895 (including ACT35, 12H3, 12H3 (FIG. 25)--and humanised versions VL1H1, VL1VH2, VL1VH3, VL2H1, VL2VH2 and VL2VH3 (FIGS. 43 & 44) and 20E5 (FIG. 24)), WO2013/068563 (including A26 [FIG. 2]), WO2013/038191 (including ACT35, 12H3 and 12H3), WO2013/028231 (including 119-122, 119-43-1, 106-222 and the antibodies in Table 1), WO2013/008171 (including 2F8, 1D4 and their derivatives, including VH6/VL9, and the antibodies in FIGS. 4 and 5 and tables 6 and 7), WO2012/027328 (including 119-122, 119-43-1, Hu106 and Hu106-222), WO2010/096418 (including A26), WO2008/106116 (including the antibodies in Tables 1 and 2, and A10 (inc A10A-F), B66--FIG. 14--B2, B24, B36, B37, and B39) and WO2007/062245 (including 112V8 (ATCC No. PTA-7219), 112Y55 (ATCC No. PTA-7220), 112Y131 (ATCC No. PTA-7218), 112F32 (ATCC No. PTA-7217) and 112Z5 (ATCC No. PTA-7216); the sequences and features of the anti-OX40 antibodies are incorporated herein by reference.
[0838] In one embodiment, the antigen-binding site specifically binds CXCR3, e.g. CXCR3. In one embodiment, the CXCR3 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from GSK3174998 or from any one of the anti-CXCR3 antibodies described in WO2016/200836, WO2016/200835, WO2016/196228, WO2016/179517, WO2016/057667, WO2015/153513, WO2014/148895, WO2013/068563, WO2013/038191, WO2013/028231, WO2013/008171, WO2012/027328, WO2010/096418, WO2011/073180, WO2008/106116 and WO2007/062245; the sequences and features of the anti-CXCR3 antibodies are incorporated herein by reference.
[0839] In one embodiment, the antigen-binding site specifically binds CD27, e.g. hCD27. In one embodiment, the CD27 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from any one of the anti-CD27 antibodies described in WO2016/145085 (including 1F5), WO2015/016718 (including hCD27.15 and 1F5), WO2014/140374 (including 2F2, 5F24, 5F32, 10F13, 10F31, 11F26, 1052 to 015, F2A4B2 and their derivatives, including hz5F24VH+V5Q, hz5F24VL+K45Q), WO2013/138586 (including C2177, C2186, C2191, and C2192 and the derivatives in Examples 8 to 12, and tables 7 to 42), WO2012/004367 (including hCD27.15/ATCC number PTA-11008), WO2011/130434 (including 1G5, 1H8, 3H12, 3H8, 2G9, 1F5, 3A10, 2C2, ms 1A4, ms 9F4 and ms M-T271), WO2011/081164 & WO2010/001908 (including KM4027, KM4028, KM4026, KM4030, KM4032 and derivatives thereof), WO2008/051424 (including LG3A10 and AT124-1); the sequences and features of the anti-CD27 antibodies are incorporated herein by reference.
[0840] In one embodiment, the antigen-binding site specifically binds CD3, e.g. hCD3. In one embodiment, the CD3 antigen-binding site comprises the CDRH1, CDRH2, CDRH3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region from OKT3 antibody, otelixizumab, teplizumab or visilizumab, or from any one of the anti-CD3 antibodies described in WO2017/010874, WO2017/009442, WO2016/204966, WO2016/180721, WO2016/179003, WO2016/116626, WO2016/014974, WO2015/104346, WO2015/095392, WO2015/001085, WO2014/047231, WO2013/188693, WO2013/186613, WO2013/158856, WO2012/173819, WO2012/162067, WO2005/118635, WO2004/108158, WO2004/052397, WO2004/024771, WO01/51644, WO00/05268, WO97/44362, WO93/19196, WO92/06193 and WO91/09968; the sequences and features of the anti-CD3 antibodies are incorporated herein by reference.
Aspect 1b. An immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction: [0841] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and [0842] b) A heavy chain constant region; [0843] c) Optionally, a linker, (L); and [0844] d) An IL-2 cytokine;
[0845] and wherein the light chain comprises in N- to C-terminal direction: [0846] e) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3; and [0847] f) A light chain constant region, (C.sub.L); [0848] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to an antigen selected from: an immune checkpoint inhibitor (such as PD-1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, TIM-3 and LAG-3), an immune modulator (such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R), and an immune activator (such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD27, CD3 and ICOS (e.g. agonistic anti-ICOS antibodies), for example ICOS, CD137, GITR and OX40).
[0849] In another embodiment, the antigen-binding site that specifically binds to an antigen selected from: an immune checkpoint inhibitor (such as PD-1, CRA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. TIGIT, TIM-3 and LAG-3), an immune modulator (such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R), and an immune activator (such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD3 and ICOS (e.g. agonistic anti-ICOS antibodies), for example ICOS, CD137, GITR and OX40).
[0850] Any of the embodiments of aspect 1 and/or aspect 1a apply mutatis mutandis to aspect 1b. Any of the features or embodiments of aspects 2 to 54 apply mutatis mutandis to aspect 1b. Any of the features of the antibodies or other embodiments or features of concepts 1 to 70 apply mutatis mutandis to aspect 1b.
[0851] In one embodiment, the antigen binding site specifically binds any of the antigens as set out in aspect 1a.
[0852] In one embodiment, the antigen-binding site specifically bind to hPD-L1 as defined by Seq ID No:1, and competes for binding to said hPD-L1 with the antibody 1D05; and wherein the immunocytokine comprises a V.sub.H domain which comprises a CDRH3 comprising the motif X.sub.1GSGX.sub.2YGX.sub.3X.sub.4FD, wherein X.sub.1, X.sub.2 and X.sub.3 are independently any amino acid, and X.sub.4 is either present or absent, and if present, may be any amino acid.
[0853] In either of aspect 1 or 1a, the wording of part f) may be substituted to read: "f) a cytokine, e.g. selected from IL-7, IL-15, IL-21, IL-12, GM-CSF, TNF.alpha., TGF.beta., CXCL9, CXCL10 and interferon-a". In 1b, the wording of part d) may be substituted for "d) a cytokine, e.g. selected from IL-7, IL-15, IL-21, IL-12, GM-CSF, TNF.alpha., TGF.beta., CXCL9, CXCL10 and interferon-.alpha.". Thus, the immunocytokines as disclosed herein may contain cytokines other than a cytokine having IL-2 cytokine activity. In one embodiment, the cytokine is IL-7 (Seq ID No:330). In one embodiment, the cytokine is IL-15 (Seq ID No:331). In one embodiment, the cytokine is IL-21 (Seq ID No:332). In one embodiment, the cytokine is IL-12, comprising the .alpha.-chain (Seq ID No:336) and the .beta.-chain (Seq ID No:337). In one embodiment, the cytokine is GM-CSF (Seq ID No:333). In one embodiment, the cytokine is TNF.alpha. (Seq ID No:335). In one embodiment, the cytokine is TGF.beta.. In one embodiment, the cytokine is CXCL9 (Seq ID No:338). In one embodiment, the cytokine is CXCL10 (Seq ID No:339). In one embodiment, the cytokine is interferon-.alpha. (Seq ID No:334).
[0854] In another embodiment, the cytokine is an immune-stimulating cytokine. In another embodiment, the cytokine is a T-cell stimulating cytokine.
Aspect 2. An immunocytokine according to aspect 1, wherein X.sub.1 is a hydroxyl-containing amino acid, optionally T. Aspect 3. An immunocytokine according to aspect 1 or aspect 2, wherein X.sub.2 is a basic amino acid, optionally K. Aspect 4. An immunocytokine according to any one of aspects 1 to 3, wherein X.sub.2 is a hydroxyl-containing amino acid, optionally S or T. Aspect 5. The immunocytokine according to any one of claims 1 to 4, wherein X.sub.3 is an aromatic amino acid, optionally W. Aspect 6. An immunocytokine according to any one of aspects 1 to 5, wherein X.sub.4 is absent. Aspect 7. An immunocytokine according to any one of aspects 1 to 5, wherein X.sub.4 is present. Aspect 8. An immunocytokine according to aspect 7, wherein X.sub.4 is an aliphatic amino acid, optionally G.
[0855] The features of aspects 2 to 7 may be as defined in any of concepts 2 to 7 hereinabove.
Aspect 9. An immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction: [0856] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and [0857] b) A heavy chain constant region; [0858] and wherein the light chain comprises in N- to C-terminal direction: [0859] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3; [0860] d) A light chain constant region, (C.sub.L); [0861] e) Optionally, a linker, (L); and [0862] f) An IL-2 cytokine; [0863] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to hPD-L1, and competes for binding to said hPD-L1 with the antibody 1D05, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:29 or 32, or the CDRH3 sequence of SEQ ID NO:29 or 32 comprising 6 or fewer amino acid substitutions, optionally, wherein the immunocytokine is according to any one of aspects 2 to 8.
[0864] In this aspect, any of the features of CDRH3 described in concepts 9, and 9a to I, and any of the embodiments of concept 9 apply mutatis mutandis.
Aspect 10. An immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction: [0865] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and [0866] b) A heavy chain constant region; [0867] and wherein the light chain comprises in N- to C-terminal direction: [0868] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3; [0869] d) A light chain constant region, (C.sub.L); [0870] e) Optionally, a linker, (L); and [0871] f) An IL-2 cytokine; [0872] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to hPD-L1; and [0873] wherein the V.sub.H domain comprises a CDRH3 of from 12 to 20 amino acids and which is derived from the recombination of a human V.sub.H gene segment, a human D gene segment and a human J.sub.H gene segment, wherein the human J.sub.H gene segment is IGHJ5 (e.g. IGHJ5*02).
[0874] In this aspect, any of the features of CDRH3 described in concepts 10 and 10a apply mutatis mutandis.
Aspect 11. An immunocytokine according to aspect 10, wherein the human V.sub.H gene segment is IGHV3 (e.g. IGHV3-9, such as IGHV3-9*01).
[0875] In this aspect, any of the features of the gene segments described in concept 11, 11a or 11b apply mutatis mutandis.
[0876] Aspect 12. An immunocytokine according to aspect 10 or aspect 11, wherein the antibody or fragment comprises a V.sub.L domain which is derived from the recombination of a human V.sub.K gene segment, and a human J.sub.K gene segment, wherein the human V.sub.L gene segment is IG.kappa.V1D (e.g. IG.kappa.V1D-39, such as IG.kappa.V1D-39*01).
[0877] In this aspect, any of the features of the gene segments described in concept 12, 12a or 12b apply mutatis mutandis.
Aspect 13. An immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction: [0878] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and [0879] b) A heavy chain constant region; [0880] and wherein the light chain comprises in N- to C-terminal direction: [0881] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3; [0882] d) A light chain constant region, (C.sub.L); [0883] e) optionally, a linker, (L); and [0884] f) An IL-2 cytokine; [0885] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site that specifically binds to an epitope that is identical to an epitope to which the antibody 1D05 specifically binds.
[0886] In this aspect, any of the features of the epitopes, assays and other embodiments described in any of concepts 13 and 13a to 13l apply mutatis mutandis.
Aspect 14. An immunocytokine according to aspect 13, wherein the epitope is identified by an unrelated amino acid scan, or by X-ray crystallography. Aspect 15. An immunocytokine according to aspect 14, wherein the contact residues of the epitope are defined by a reduction in affinity of at least 10-fold in an unrelated amino acid scan, e.g. an alanine scan as determined by SPR.
[0887] In this aspect, any of the features of concept 15 apply mutatis mutandis Aspect 16. An immunocytokine comprising an immunoglobulin heavy chain and an immunoglobulin light chain, wherein the heavy chain comprises in N- to C-terminal direction: [0888] a) A V.sub.H domain comprising CDRH1, CDRH2 and CDRH3; and [0889] b) A heavy chain constant region; [0890] and wherein the light chain comprises in N- to C-terminal direction: [0891] c) A V.sub.L domain comprising CDRL1, CDRL2 and CDRL3; [0892] d) A light chain constant region, (C.sub.L); [0893] e) Optionally, a linker, (L); and [0894] f) An IL-2 cytokine; [0895] wherein the V.sub.H domain and V.sub.L domain are comprised by an antigen-binding site which competes for binding to hPD-L1 with the antibody 1D05.
[0896] In this aspect, any of the features of the antibodies of concepts 16a to 16l or any of the competitive-assays and other embodiments described in concept 16, or the features of concept 35 apply mutatis mutandis.
[0897] Aspect 17. An immunocytokine according to any one of aspects 10 to 16, wherein the V.sub.H domain comprises the CDRH3 sequence of SEQ ID NO:29 or 32, or the CDRH3 sequence of SEQ ID NO:29 or 32 comprising 6 or fewer amino acid substitutions.
[0898] In this aspect, any of the features of the antibodies of concepts 17a to 17l apply mutatis mutandis.
Aspect 18. An immunocytokine according to any preceding aspect, wherein the V.sub.H domain comprises the CDRH1 sequence of SEQ ID NO:27 or 30 or the CDRH1 sequence of SEQ ID NO:27 or 30 comprising 3, 2 or 1 amino acid substitution(s).
[0899] In this aspect, any of the features of the antibodies of concepts 18a to 18l apply mutatis mutandis.
Aspect 19. An immunocytokine according to any preceding aspect, wherein the V.sub.H domain comprises the CDRH2 sequence of SEQ ID NO:28 or 31, or the CDRH2 sequence of SEQ ID NO:28 or 31 comprising 4 or fewer amino acid substitutions.
[0900] In this aspect, any of the features of the antibodies of concepts 19a to 19l apply mutatis mutandis.
Aspect 20. An immunocytokine according to any preceding aspect, wherein the V.sub.H domain comprises an amino acid sequence of SEQ ID NO:33, or a heavy chain variable domain amino acid sequence that is at least 80% (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:33 (for example the V.sub.H domain sequence in any of the heavy chain sequences of Seq ID Nos:47 to 49).
[0901] In this aspect, any of the features of the antibodies of concepts 20a to 201 or any of the embodiments of concept 20 apply mutatis mutandis.
Aspect 21. An immunocytokine according to any preceding aspect comprising first and second copies of said heavy chain. Aspect 22. An immunocytokine according to any preceding aspect, comprising a V.sub.L domain which comprises the CDRL1 sequence of SEQ ID NO:37 or 40, or the CRDL1 sequence of SEQ ID NO:37 or 40 comprising 3 or fewer amino acid substitutions.
[0902] In this aspect, any of the features of the antibodies of concepts 22a to 22l apply mutatis mutandis.
Aspect 23. An immunocytokine according to any preceding aspect, comprising a V.sub.L domain which comprises the CDRL2 sequence of SEQ ID NO:38 or 41, or the CRDL2 sequence of SEQ ID NO:38 or 41 comprising 2 or 1 amino acid substitution(s), for example a CDRL2 sequence of Seq ID No:50.
[0903] In this aspect, any of the features of the antibodies of concepts 23a to 23l apply mutatis mutandis.
Aspect 24. An immunocytokine according to any preceding aspect, comprising a V.sub.L domain which comprises the CDRL3 sequence of SEQ ID NO:39 or 42, or the CRDL3 sequence of SEQ ID NO:39 or 42 comprising 4 or fewer amino acid substitutions.
[0904] In this aspect, any of the features of the antibodies of concepts 24a to 241 apply mutatis mutandis.
Aspect 25. An immunocytokine according to any preceding aspect, comprising a V.sub.L domain which comprises an amino acid sequence of SEQ ID NO:43, or a light chain variable domain amino acid sequence that is at least 80%, (e.g. at least 85%, or at least 90%) identical to SEQ ID NO:43 (for example the V.sub.L domain sequence in the light chain sequence of Seq ID No:50 or 51).
[0905] In this aspect, any of the features of the antibodies of concepts 25a to 251 or any of the embodiments of concept 25 apply mutatis mutandis.
Aspect 26. An immunocytokine according to any preceding aspect comprising first and second copies of said light chain. Aspect 27. An immunocytokine according to any preceding aspect which specifically binds to cynomolgus PD-L1 as defined by Seq ID No:2.
[0906] In this aspect, any of embodiments of concept 27 apply mutatis mutandis.
Aspect 28. An immunocytokine according to any preceding aspect, wherein the antibody or fragment comprises a kappa light chain.
[0907] In this aspect, any of the embodiments of concept 28 apply mutatis mutandis.
Aspect 29. An immunocytokine according to any one of aspects 9 to 28, wherein the amino acid substitutions are conservative amino acid substitutions, optionally wherein the conservative substitutions are from one of six groups (each group containing amino acids that are conservative substitutions for one another) selected from: [0908] 1) Alanine (A), Serine (S), Threonine (T); [0909] 2) Aspartic acid (D), Glutamic acid (E); [0910] 3) Asparagine (N), Glutamine (Q); [0911] 4) Arginine (R), Lysine (K); [0912] 5) Isoleucine (I), Leucine (L), Methionine (M), Valine (V); and [0913] 6) Phenylalanine (F), Tyrosine (Y), Tryptophan (W).
[0914] In this aspect, any of the embodiments of concept 9 apply mutatis mutandis.
Aspect 30. An immunocytokine according to any preceding aspect, wherein the antibody or fragment comprises a constant region, e.g. an IgG1 constant region, optionally wherein the constant region is a disabled IgG1 as defined in Seq ID No:205.
[0915] In this aspect, any of the features or the embodiments of concepts 30, 31 or 32 apply mutatis mutandis.
Aspect 31. An immunocytokine according to any preceding aspect wherein the: [0916] A) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0917] B) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:33, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:43; [0918] C) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0919] D) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0920] E) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:43; [0921] F) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0922] G) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0923] H) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0924] I) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:50; [0925] J) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0926] K) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0927] L) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0928] M) V.sub.H domain comprise an amino acid sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:51; [0929] N) V.sub.H domain comprises an amino acid sequence of SEQ ID No:33 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0930] O) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:47 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0931] P) V.sub.H domain comprises an amino acid sequence of the V.sub.H domain of SEQ ID No:48 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0932] Q) V.sub.H domain comprise an amino add sequence of the V.sub.H domain of SEQ ID No:49 and the V.sub.L domain comprises an amino acid sequence of the V.sub.L domain of SEQ ID No:298; [0933] R) V.sub.H domain comprises an amino acid sequence of SEQ ID No:58 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:68; [0934] S) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:58, and the V.sub.L domain comprise an amino acid sequence that is at least 85% identical to SEQ ID No:68; [0935] T) V.sub.H domain comprises an amino acid sequence of SEQ ID No:78 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:88; [0936] U) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:78, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:88; [0937] V) V.sub.H domain comprises an amino acid sequence of SEQ ID No:98 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:108; [0938] W) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:98, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:108; [0939] X) V.sub.H domain comprises an amino acid sequence of SEQ ID No:118 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:128; [0940] Y) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:118, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:128; [0941] Z) V.sub.H domain comprises an amino acid sequence of SEQ ID No:158 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:168; [0942] AA) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:158, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:168; [0943] BB) V.sub.H domain comprises an amino acid sequence of SEQ ID No:178 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:188; [0944] CC)V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:178, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:188; [0945] DD) V.sub.H domain comprises an amino acid sequence of SEQ ID No:138 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:148; [0946] EE) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:13, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:148; [0947] FF) V.sub.H domain comprises an amino acid sequence of SEQ ID No:244 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:254; [0948] GG) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:244, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:254; [0949] HH) V.sub.H domain comprises an amino acid sequence of SEQ ID No:264 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:274; [0950] II) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:264, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:274; [0951] JJ) V.sub.H domain comprises an amino acid sequence of SEQ ID No:284 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:294; [0952] KK) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:284, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:294; [0953] LL) V.sub.H domain comprises an amino acid sequence of SEQ ID No:13 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:23; and [0954] MM) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:13, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:23; [0955] NN) V.sub.H domain comprises an amino acid sequence of SEQ ID No:349 and the V.sub.L domain comprises an amino acid sequence of SEQ ID No:359; and [0956] OO) V.sub.H domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:349, and the V.sub.L domain comprises an amino acid sequence that is at least 85% identical to SEQ ID No:359.
[0957] In one embodiment, the amino acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99.5% identical to the specified Seq ID No.
Aspect 32. An immunocytokine according to any preceding aspect wherein the: [0958] A) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:299 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:45; [0959] B) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:299, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:45; [0960] C) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:47 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:45; [0961] D) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:48 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:45; [0962] E) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:49 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:45; [0963] F) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:342 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:45; [0964] G) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:238 and V.sub.L. and C.sub.L comprise an amino acid sequence of SEQ ID No:50; [0965] H) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:47 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:50; [0966] I) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:48 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:50; [0967] J) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:49 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:50; [0968] K) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:342 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:50; [0969] L) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:299 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:51; [0970] M) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:47 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:51; [0971] N) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:48 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:51; [0972] O) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:49 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:51; [0973] P) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:342 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:51; [0974] Q) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:299 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:298; [0975] R) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:47 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:298; [0976] S) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:48 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:298; [0977] T) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:49 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:298; [0978] U) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:342 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:298; [0979] V) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:60 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:70; [0980] W) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:60, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:70; [0981] X) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:80 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:90; [0982] Y) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:80, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:90; [0983] Z) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:100 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:110; [0984] AA) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:100, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:110; [0985] BB) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:120 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:130; [0986] CC) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:120, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:130; [0987] DD) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:160 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:170; [0988] EE) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:160, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:170; [0989] FF) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:180 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:190; [0990] GG) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:180, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:190; [0991] HH) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:140 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:150; [0992] II) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:140, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:150; [0993] 33) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:246 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:256; [0994] KK) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:246, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:256; [0995] LL) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:266 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:276; [0996] MM) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:266, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:276; [0997] NN) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:286 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:296; and [0998] OO) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:286, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:296; [0999] PP) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:15 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:25; and [1000] QQ) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:15, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:25; [1001] RR) V.sub.H and the constant region comprise an amino acid sequence of SEQ ID No:351 and V.sub.L and C.sub.L comprise an amino acid sequence of SEQ ID No:361; and [1002] SS) V.sub.H and the constant region comprise an amino acid sequence that is at least 85% identical to SEQ ID No:351, and the V.sub.L and C.sub.L comprise an amino acid sequence that is at least 85% identical to SEQ ID No:361.
[1003] In one embodiment, the amino acid sequence is at least 70% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 75% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 95% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 96% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 97% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 98% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99% identical to the specified Seq ID No. In one embodiment, the amino acid sequence is at least 99.5% identical to the specified Seq ID No.
Aspect 33. An immunocytokine according to any preceding aspect wherein the antigen-binding site specifically binds PD-L1, whilst the IL-2 cytokine binds the high affinity (.alpha..beta..gamma.) IL-2 receptor (IL-2R).
[1004] In one embodiment, the antigen binding site binds PD-L1 simultaneously to the IL-2 cytokine binding the .alpha..beta..gamma. IL-2R. In one embodiment, the antigen binding site binds PD-L1 sequentially to the IL-2 cytokine binding the .alpha..beta..gamma. IL-2R. In one embodiment, the IL-2 cytokine additionally binds the intermediate (.beta..gamma.) IL-2R.
Aspect 34. An immunocytokine according to any preceding aspect wherein the immunocytokine is capable of inhibiting PD-L1-mediated suppression of T-cells.
[1005] In one embodiment, the immunocytokine inhibits PD-L1-mediated suppression of T-cells. In one embodiment, the immunocytokine inhibits PD-L1-mediated suppression of T-cells in an in vitro assay. In another embodiment, the antigen binding site has any of the features or embodiments of concept 51 or 52.
[1006] In another embodiment, the antigen binding site blocks or inhibits PD-1 binding to PD-L1. In one embodiment, the antigen binding site blocks or inhibits CD80 binding to PD-L1.
Aspect 35. An immunocytokine according to any preceding aspect wherein the immunocytokine is capable of increasing IL-2R-mediated T-cell activation.
[1007] In one embodiment, the immunocytokine increases IL-2R-mediated T-cell activation. In one embodiment, the immunocytokine increases IL-2R-mediated T-cell activation in an in vitro assay.
Aspect 36. An immunocytokine according to aspect 34 or aspect 35, wherein the suppression of T-cells or the increase in IL-2R-mediated T-cell activation is measured by an increase in one or more of IFN.gamma., IL-2, CD25 or proliferation of T-cells in an assay that provides co-stimulation by either direct CD3/CD28 stimulation, superantigen stimulation or provides co-stimulation by co-incubation with cells capable of inducing a T-cell response.
[1008] The measurements may be carried out with any suitable technique. For example, the measurements may be taken with ELISA, HTRF, BRDU incorporation (proliferation), electrochemiluminescence (ECL) or flow cytometry (e.g. FACS). These techniques are well-known to those skilled in the art and are described elsewhere herein. In one embodiment, the assay is flow cytometry. In one embodiment, the assay is ELISA. In one embodiment, the assay is HTRF.
[1009] In this aspect, when aspect 36 is dependent on aspect 34, any of the features or embodiments of concept 36 apply mutatis mutandis.
[1010] When Aspect 36 is dependent on Aspect 35, in one embodiment, the increase in IL-2R-mediated T-cell activation is measured by an increase in one or both of IFN.gamma. and CD25.
[1011] When Aspect 36 is dependent on Aspect 35, in one embodiment, the co-stimulation is provided by direct CD3/CD28 stimulation.
[1012] When Aspect 36 is dependent on Aspect 35, in one embodiment, the co-stimulation is provided by a superantigen, such as staphylococcal enterotoxin B (SEB).
[1013] When Aspect 36 is dependent on Aspect 35, in one embodiment, the assay provides co-stimulation by co-incubation with cells capable of inducing a T-cell response. Such cells may be antigen-presenting cells (APCs), for example monocytes, B-cells or dendritic cells. In one embodiment, the assay provides co-stimulation by co-incubation with APCs. In one embodiment, the assay provides co-stimulation by co-incubation with monocytes. In one embodiment, the assay provides co-stimulation by co-incubation with B-cells. In one embodiment, the assay provides co-stimulation by co-incubation with dendritic cells.
Aspect 37. An immunocytokine according to any preceding aspect which does not comprise a linker (L), or an immunocytokine according to any preceding claim wherein the C.sub.L of d) is directly fused to the cytokine of f).
[1014] In one embodiment, the C.sub.L of the light chain or the heavy chain is directly fused to the cytokine.
[1015] In one embodiment of aspect 1b, the Q. of b) is directly fused to the cytokine of d).
Aspect 38. An immunocytokine according to any one of aspects 1 to 37, wherein the linker is a peptide linker of 1 to 20 amino acids in length.
[1016] In one embodiment, the linker is peptide linker of 1 to 15 amino acids in length. In one embodiment, the linker is peptide linker of 1 to 10 amino acids in length. In one embodiment, the linker is peptide linker of 1 to 5 amino acids in length.
[1017] In one embodiment, the linker may be a chemical linker. In the case of recombinant fusion proteins, the linkers are encoded by nucleic acid sequences located in frame, in between the coding regions for the different immunocytokine portions. In the case of synthetic proteins, the linker peptides are introduced during synthesis.
[1018] Linkers are well-known to those skilled in the art. For example, see described in Denardo et al., 1998, Clin. Cancer Res., 4(10):2483-90; Peterson et al., 1999, Bioconjug. Chem. 10(4):553-7; and Zimmerman et al., 1999, Nucl. Med. Biol., 26(8):943-50, the modifications described therein are incorporated herein by reference.
Aspect 39. An immunocytokine according to aspect 38, wherein the linker peptide is selected from poly-G or (G4S)x, wherein X is 1, 2, 3 or 4.
[1019] In other embodiments, the linker may be selected from STG, GSTG, RS, TVAAPS, GGGGS, GSTVAAPS, TVAAPSGS or GSTVAAPSGS. In another embodiment, the linker is Gln-Arg-Val-Asp (derived from N-terminus of canine kappa constant region). In another embodiment, the linker is GGNGT or YGNGT.
Aspect 40. An immunocytokine according to any preceding aspect wherein the IL-2 cytokine is human IL-2 (hIL-2) or a variant thereof.
[1020] IL-2 variants are as described in aspect 1.
[1021] There is also provided a variant cytokine, which may be any of the non-IL-2 cytokines described herein (including the non-IL-2 cytokines described in aspect 1, e.g. selected from IL-7, IL-15, IL-21, IL-12, GM-CSF, TNF.alpha., CXCL9, CXCL10 and interferon-.alpha.). The definition of a variant IL-2 cytokine applies mutatis mutandis to the other cytokines (including immune stimulating cytokines and T-cell stimulating cytokines) described herein, e.g. comprising any of the N-terminal deletions described for IL-2 in aspect 1.
Aspect 41. An immunocytokine according to aspect 40, wherein the hIL-2 comprises or consists of the amino acid sequence of Seq ID No:301. Aspect 42. An immunocytokine according to aspect 40, wherein the hIL-2 comprises a variant of IL-2 which comprises a modification at the N-terminus, optionally a deletion of from 1 to 10 amino acids.
[1022] As used in this aspect, a modification at the N-terminus of any of the cytokines described herein (including the non-IL-2 cytokines described in aspect 1, e.g. selected from IL-7, IL-15, IL-21, IL-12, GM-CSF, TNF.alpha., CXCL9, CXCL10 and interferon-.alpha.) refers to one or more (such as 1 to 10, e.g. 1 to 5) amino acid substitutions, deletions or additions.
[1023] In one embodiment, the modification is one or more (such as 1 to 10, e.g. 1 to 5) amino acid substitutions at the N-terminus of the cytokine. Substitutions may be conservative substitutions, for example, as defined in concept 9, concept 29 or aspect 29. In one embodiment, the modification is a deletion. In another embodiment, the modification is an N-terminal deletion, for example, any of the deletions described in concept 9 and aspect 1. In one embodiment, the modification (such as a deletion of 1 to 10 amino acids) is within the final 50 amino acids of the N-terminus of the cytokine, e.g. the IL-2 cytokine. In one embodiment, the modification (such as a deletion of 1 to 10 amino acids) within the final 30 amino acids of the N-terminus of the cytokine, e.g. the IL-2 cytokine. In one embodiment, the modification (such as a deletion of 1 to 10 amino acids) within the final 25 amino acids of the N-terminus of the cytokine, e.g. the IL-2 cytokine. In one embodiment, the modification (such as a deletion of 1 to 10 amino acids) within the final 20 amino acids of the N-terminus of the cytokine, e.g. the IL-2 cytokine. In one embodiment, the modification (such as a deletion of 1 to 10 amino acids) within the final 15 amino acids of the N-terminus of the cytokine, e.g. the IL-2 cytokine. In one embodiment, the modification (such as a deletion of 1 to 10 amino acids) within the final 10 amino acids of the N-terminus of the cytokine, e.g. the IL-2 cytokine.
[1024] In one embodiment, the modification is a deletion of 1 to 9 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 to 9 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion of 1 to 8 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 to 8 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion of 1 to 7 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 to 7 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion of 1 to 6 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 to 6 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion of 1 to 5 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 to 5 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion of 1 to 4 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 to 4 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion or 1 to 3 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 to 3 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion of 1 or 2 amino acids from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final 1 or 2 amino acids of the N-terminus of the cytokine. In one embodiment, the modification is a deletion of 1 amino acid from within the final 10 amino acids of the N-terminus of the cytokine, such as a deletion of the final amino acid of the N-terminus of the cytokine. In a particular embodiment, the cytokine is an IL-2 cytokine, such as a human IL-2 cytokine.
[1025] In one embodiment, the deletion is of the 9.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 8.sup.th and 9.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 7.sup.th, 8.sup.th and 9.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 6th to 9.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 4.sup.th to 9.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 3.sup.rd to 9.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 2.sup.nd to 9.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 2.sup.nd to 6.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 3.sup.rd to 7.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. In one embodiment, the deletion is of the 4.sup.th to 8.sup.th amino acid from the N-terminus of the cytokine, e.g. the IL-2 cytokine, for example the human IL-2 cytokine. Any of the deletions described in Aspect 1 hereinabove may be applied mutatis mutandis to the non-IL-2 cytokines of this aspect.
[1026] Aspect 42a. A variant hIL-2 comprising an N-terminal modification of any of the aspects or features of aspect 42. In one embodiment of aspect 42a, the variant hIL-2 is a purified variant hIL-2. In another embodiment of aspect 42a, the variant hIL-2 is an isolated and purified variant hIL-2. Aspect 42b. A variant cytokine selected from IL-7, IL-15, IL-21, IL-12, GM-CSF, INF.alpha., CXCL9, CXCL10 and interferon-a comprising an N-terminal modification of any of the aspects or features of aspect 42. In one embodiment of aspect 42a, the variant cytokine is a purified variant cytokine. In another embodiment of aspect 42a, the variant cytokine is an isolated and purified variant cytokine.
Aspect 43. An immunocytokine according to aspect 40 or aspect 42, wherein the hIL-2 comprises a variant IL-2 comprising an N-terminal sequence selected from Seq ID No:303 to 323.
[1027] Aspect 43a. A variant hIL-2 comprising an N-terminal sequence selected from Seq ID No:303 to 323.
[1028] In one embodiment of aspect 43a, the variant hIL-2 is a purified variant hIL-2. In another embodiment of aspect 43a, the variant hIL-2 is an isolated and purified variant hIL-2. In one embodiment, the variant hIL-2 comprises (or consists) of an N-term terminal sequence selected from Seq ID No:303 to 323 directly fused to an IL-2 sequence selected from Seq ID No:324, 517 and 518.
Aspect 44. An immunocytokine according to any one of aspects 40, 42 or 43 wherein the hIL-2 variant comprises one or more (such as 1 to 5, e.g. one or two) mutations independently selected from the following: [1029] 1) D20 (such as D20T); [1030] 2) R38 (such as R38W, R38A or R38Q); [1031] 3) F42 (such as F42A or F42K); [1032] 4) Y45 (such as Y45A); [1033] 5) E62 (such as E62A); [1034] 6) N88 (such as N88R); [1035] 7) C125 (such as C125S); [1036] 8) Q126 (such as Q126W); or [1037] 9) R38 and F42 (such as R38W and F42K or R38A and F42A); [1038] wherein the residue numbering is defined with reference to the human wild-type IL-2 sequence, Seq ID No:301. Aspect 44a. A variant hIL-2 according to any one of aspects 42a or 43a wherein the hIL-2 variant comprises one or more (such as 1 to 5, e.g. one or two) mutations independently selected from the following: [1039] 1) D20 (such as D20T); [1040] 2) R38 (such as R38W, R38A or R38Q); [1041] 3) F42 (such as F42A or F42K); [1042] 4) Y45 (such as Y45A); [1043] 5) E62 (such as E62A); [1044] 6) N88 (such as N88R); [1045] 7) C125 (such as C1255); [1046] 8) Q126 (such as Q126W); or [1047] 9) R38 and F42 (such as R38W and F42K or R38A and F42A); [1048] wherein the residue numbering is defined with reference to the human wild-type IL-2 sequence, Seq ID No:301.
[1049] In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q, e.g. R38A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an F42 (such as F42A or F42K, e.g. F42A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) or a Y45 (such as Y45A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an E62 (such as E62A) mutation.
[1050] In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q, e.g. R38A) mutation and an F42 (such as F42A or F42K, e.g. F42A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q e.g. R38A) and a Y45 (such as Y45A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q, e.g. R38A) mutation and an E62 (such as E62A). In one embodiment, the variant hIL-2 comprises (or consists) of a Y45 (such as Y45A) mutation and an E62 (such as E62A). In one embodiment, the variant hIL-2 comprises (or consists) of an F42 (such as F42A or F42K, e.g. F42A) mutation and an E62 (such as E62A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an F42 (such as F42A or F42K, e.g. F42A) mutation and a Y45 (such as Y45A) mutation.
[1051] In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q, e.g. R38A) mutation, an F42 (such as F42A or F42K, e.g. F42A) mutation and a Y45 (such as Y45A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q, e.g. R38A) mutation, an F42 (such as F42A or F42K, e.g. F42A) mutation and an E62 (such as E62A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q; e.g. R38A) mutation, a Y45 (such as Y45A) mutation and an E62 (such as E62A) mutation.
[1052] In one embodiment, the variant hIL-2 comprises (or consists) of an R38 (such as R38W, R38A or R38Q e.g. R38A) mutation, an F42 (such as F42A or F42K, e.g. F42A) mutation, a Y45 (such as Y45A) mutation and an E62 (such as E62A) mutation. In one embodiment, the variant hIL-2 comprises (or consists) of an R38A, F42A, Y45A and an E62A mutation.
[1053] Other hIL-2 mutations are known to those skilled in the art. In one embodiment, the hIL-2 mutations are those described in WO2012/062228 (see claims 2 to 7, incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO1999/60128 (see claims 6, 7, 8, 10, 11 and 12 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO1993/20849 (see claims 4 and 5 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2003/015697 (see claims 7 and 10 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2005/007121 (see claims 9 to 14 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2005/086798 (see claims 5 to 10 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2005/086751 (see claims 5 to 9 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2009/061853 (see claim 5 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2012/088446 (see claims 3 to 8 and 11 to 13 incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2012/107417 (see claims 2, 4, 6 and 9, incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2012/119093 (see claims 1 to 7, incorporated herein by reference). In one embodiment, the hIL-2 mutations are those described in WO2015/164815 (see claims 3 to 19, incorporated herein by reference).
[1054] In these aspects, where the residue numbering is defined with reference to the human wild-type IL-2 sequence, if, for example, there is a single amino acid deletion from the N-terminus of the cytokine, and the claim described an N88 amino acid mutation, then, for the variant IL-2 having the single amino acid deletion, the N will in fact be at position 87. If the cytokine has 3 amino acids deleted from the N-terminus, and the mutation is an F42A mutation, then the position to be mutated, will in fact be F39 in the variant sequence.
[1055] Aspect 45. An immunocytokine according to aspect 40, wherein the hIL-2 comprises a variant IL-2 consists of an N-terminal sequence selected from Seq ID No:242 to 262 fused to the amino acid sequence of Seq ID No:324.
[1056] In one embodiment, the variant hIL-2 comprises (or consists) of an N-terminal sequence selected from Seq ID No:303 to 323 fused to the amino acid sequence selected from Seq ID No:324, 517 and 518.
[1057] In one embodiment, the immunocytokine is 1D05 D1-9 ICK. In one embodiment, the immunocytokine is 1D05 D1-9. In one embodiment, the immunocytokine is 1D05 D9-2 ICK. In one embodiment, the immunocytokine is 1D05 D9-7 ICK.
[1058] Aspect 45a. A variant hIL-2 comprising an N-terminal sequence selected from Seq ID No:303 to 323 fused to the amino acids sequence of Seq ID No:324.
[1059] In one embodiment of aspect 45a, the variant hIL-2 is a purified variant hIL-2. In another embodiment of aspect 44a, the variant hIL-2 is an isolated and purified variant hIL-2.
Aspect 46. An immunocytokine according to any preceding aspect, wherein the IL-2 cytokine binds to the high affinity (.alpha..beta..gamma.) IL-2 receptor with a potency less than free IL-2, for example with an EC.sub.50 of greater than 20 pM, greater than 50 pM or greater than 100 pM, e.g. when measured in a cell-based proliferative assay.
[1060] Free IL-2 has a potency of approximately 10 pM against the .alpha..beta..gamma. (high affinity) receptor in a cell-based proliferative assay. As used herein, EC.sub.50 refers to the effective concentration to give 50% of maximal activation of the IL2R. The higher the EC.sub.50, the less potent the substance is, thus a substance having an EC.sub.50 of 1 pM is more potent than a substance with an EC.sub.50 of 1 nM. The sequences of the .alpha.-chain, .beta.-chain and .gamma.-chain are provided in Seq ID Nos:327, 328 and 329 respectively.
[1061] In one embodiment, the IL-2 cytokine has an EC.sub.50 in the range of 5 pM to 20 pM. In one embodiment, the EC.sub.50 is in the range of 5 pM to 1 nM. In one embodiment, the EC.sub.50 is in the range of 5 pM to 750 pM, 5 pM to 500 pM, 5 pM to 250 pM or 5 pm to 100 pM, e.g. 5 pM to 50 pM.
[1062] In one embodiment, the EC.sub.50 is in the range of 10 pM to 1 nM. In one embodiment, the EC.sub.50 is in the range of 10 pM to 750 pM, 10 pM to 500 pM, 10 pM to 250 pM or 10 pm to 100 pM, e.g. 10 pM to 50 pM, or 10 pM to 30 pM.
[1063] In one embodiment, the EC.sub.50 is in the range of 20 pM to 1 nM. In one embodiment, the EC.sub.50 is in the range of 20 pM to 750 pM, 20 pM to 500 pM, 20 pM to 250 pM or 20 pm to 100 pM, e.g. 20 pM to 50 pM.
[1064] In another embodiment, the IL-2 cytokine has an EC.sub.50 in the range of 50 pM to 1 nM. In one embodiment, the EC.sub.50 is in the range of 50 pM to 750 pM, 50 pM to 500 pM, 50 pM to 250 pM or 50 pm to 100 pM, e.g. 50 pM to 75 pM. In another embodiment, the IL-2 cytokine has an EC.sub.50 in the range of 100 pM to 1 nM. In one embodiment, the EC.sub.50 is in the range of 100 pM to 800 pM, 100 pM to 700 pM, 100 pM to 600 pM or 100 pm to 500 pM, e.g. 100 pM to 400 pM. In another embodiment, the IL-2 cytokine has an EC.sub.50 in the range of 100 pm to 300 pM. In another embodiment, the IL-2 cytokine has an EC.sub.50 in the range of 100 pm to 200 pM.
[1065] In another embodiment, the EC.sub.50 is greater than 5 pM. In another embodiment, the EC.sub.50 is greater than 10 pM. In another embodiment, the EC.sub.50 is greater than 20 pM. In another embodiment, the EC.sub.50 is greater than 30 pM, greater than 40 pM, greater than 50 pM, greater than 60 pM or greater than 70 pM. In another embodiment, the EC.sub.50 is greater than 100 pM, greater than 125 pM, greater than 150 pM, greater than 175 pM or greater than 200 pM. In another embodiment, the EC.sub.50 is greater than 250 pM, greater than 300 pM, greater than 350 pM, greater than 400 pM. In another embodiment, the EC.sub.50 is greater than 500 pM, greater than 600 pM, greater than 700 pM or greater than 800 pM.
[1066] In one embodiment, the EC.sub.50 is less than 5 nM. In one embodiment, the EC.sub.50 is less than 1 nM. In one embodiment, the EC.sub.50 is less than 800 pM. In one embodiment, the EC.sub.50 is less than 700 pM. In one embodiment, the EC.sub.50 is less than 600 pM. In one embodiment, the EC.sub.50 is less than 500 pM. In one embodiment, the EC.sub.50 is less than 400 pM. In one embodiment, the EC.sub.50 is less than 300 pM. In one embodiment, the EC.sub.50 is less than 200 pM. In one embodiment, the EC.sub.50 is less than 100 pM. In one embodiment, the EC.sub.50 is less than 50 pM.
[1067] The potency of the immunocytokine against the .alpha..beta..gamma. IL-2R may be measured in a cell-based proliferative assay, which are well-known to those skilled in the art and are detailed more in the Examples hereinbelow (see Example 13 and FIG. 12).
Aspect 47. An immunocytokine according to any preceding aspect, wherein the IL-2 binds to the intermediate affinity (.beta..gamma.) IL-2 receptor with a potency less than free IL-2, for example with an EC.sub.50 of greater than 1 nM, greater than 5 nM or greater than 10 nM, e.g. when measured in a cell-based proliferative assay.
[1068] Free IL-2 has a potency of approximately 100 pM against the .beta..gamma. (intermediate affinity) receptor in a cell-based proliferative assay. As used herein, EC.sub.50 refers to the effective concentration to give 50% of maximal activation of the IL-2R. The higher the EC.sub.50, the less potent the substance is, thus a substance having an EC.sub.50 of 1 pM is more potent than a substance with an EC.sub.50 of 1 nM. The sequences of the .alpha.-chain, .beta.-chain and .gamma.-chain are provided in Seq ID Nos:327, 328 and 329 respectively.
[1069] In one embodiment, the EC.sub.50 is in the range of 1 to 100 nM. In one embodiment, the EC.sub.50 is in the range of 10 nM to 100 nM. In one embodiment, the EC.sub.50 is in the range of 20 nM to 100 nM. In another embodiment, the IL-2 cytokine has an EC.sub.50 in the range of 30 nM to 100 nM, 40 nM to 100 nM, 50 nM to 100 nM. In one embodiment, the EC.sub.50 is in the range of 50 nM to 100 nM, 60 nM to 100 nM, 70 nM to 100 nM.
[1070] In one embodiment, the EC.sub.50 is in the range of 1 to 50 nM. In one embodiment, the EC.sub.50 is in the range of 10 nM to 50 nM. In one embodiment, the EC.sub.50 is in the range of 20 nM to 50 nM. In another embodiment; the IL-2 cytokine has an EC.sub.50 in the range of 30 nM to 50 nM or 40 nM to 50 nM.
[1071] In one embodiment, the EC.sub.50 is in the range of 1 to 10 nM. In one embodiment, the EC.sub.50 is in the range of 1 to 20 nM. In one embodiment, the EC.sub.50 is in the range of 1 to 30 nM. In one embodiment, the EC.sub.50 is in the range of 1 nM to 9 nM. In one embodiment, the EC.sub.50 is in the range of 1 nM to 8 nM. In another embodiment, the IL-2 cytokine has an EC.sub.50 in the range of 1 nM to 7 nM, 1 nM to 6 nM or 1 nM to 5 nM.
[1072] In another embodiment, the EC.sub.50 is greater than 0.5 nM, greater than 0.6 nM, greater than 0.7 nM, greater than 0.8 nM or greater than 0.9 nM. In another embodiment, the EC.sub.50 is greater than 1 nM, greater than 1.25 nM, greater than 1.5 nM, greater than 1.75 nM or greater than 2 nM. In another embodiment, the EC.sub.50 is greater than 2.5 nM, greater than 3 nM, greater than 3.5 nM, greater than 4 nM. In another embodiment, the EC.sub.50 is greater than 5 nM, greater than 6 nM, greater than 7 nM or greater than 8 nM. In a particular embodiment, the EC.sub.50 is greater than 1 nM.
[1073] In one embodiment, the EC.sub.50 is less than 10 nM. In one embodiment, the EC.sub.50 is less than 20 nM. In one embodiment, the EC.sub.50 is less than 30 nM. In one embodiment, the EC.sub.50 is less than 40 nM. In one embodiment, the EC.sub.50 is less than 50 nM.
[1074] In one embodiment, the EC.sub.50 is less than 100 nM. In one embodiment, the EC.sub.50 is less than 200 nM. In one embodiment, the EC.sub.50 is less than 300 nM.
[1075] In another embodiment, the EC.sub.50 is less than 75 nM or less than 50 nM.
[1076] In one embodiment, the IL-2 shows no detectable potency against the .beta..gamma. IL-2R in a cell-based proliferative assay.
[1077] The potency of the immunocytokine against the .beta..gamma. IL-2R may be measured in a cell-based proliferative assay, which are well-known to those skilled in the art and are detailed more in the Examples hereinbelow (see Example 13 and FIG. 12).
Aspect 48. An immunocytokine according to any preceding aspect, wherein the IL-2 preferentially binds to the high affinity (.alpha..beta..gamma.) IL-2 receptor over the intermediate affinity (.beta..gamma.) IL-2 receptor. Aspect 49. An immunocytokine according to aspect 48, wherein the ratio of IL-2 potency against the high affinity (.alpha..beta..gamma.) IL-2 receptor:intermediate affinity (.beta..gamma.) IL-2 receptor is at least 2:1.
[1078] In one embodiment, the ratio is at least 3:1. In one embodiment, the ratio is at least 4:1. In one embodiment, the ratio is at least 5:1. In one embodiment, the ratio is at least 7.5:1. In one embodiment, the ratio is at least 10:1. In one embodiment, the ratio is at least 12.5:1. In one embodiment, the ratio is at least 15:1. In one embodiment, the ratio is at least 17.5:1. In one embodiment, the ratio is at least 20:1.
[1079] In another embodiment, the ratio is at least 50:1. In another embodiment, the ratio is at least 75:1. In another embodiment, the ratio is at least 100:1. In another embodiment, the ratio is at least 250:1. In another embodiment, the ratio is at least 500:1. In another embodiment, the ratio is at least 750:1. In another embodiment, the ratio is at least 1000:1.
[1080] In another embodiment, the ratio is at least 1250:1. In another embodiment, the ratio is at least 1500:1. In another embodiment, the ratio is at least 1750:1. In another embodiment, the ratio is at least 2000:1.
Aspect 50. An immunocytokine according to any preceding aspect, wherein the antigen binding site binds to hPD-L1 with an affinity of less than 500 pM (e.g. less than 300 pM or less than 200 pM), optionally wherein the immunocytokine provides a ratio of the potency of the IL-2 cytokine against the high affinity (.alpha..beta..gamma.) receptor: affinity of the anti-PD-L1 antigen binding site against hPD-L1 of at least 2:1.
[1081] In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of less than 200 pM. In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of less than 100 pM, or less than 50 pM.
[1082] In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of between 50 pM and 500 pM, or between 75 pM and 500 pM, or between 100 pM and 500 pM or between 200 pM and 500 pM.
[1083] In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of between 50 pM and 400 pM, or between 50 pM and 300 pM, or between 50 pM and 200 pM or between 50 pM and 100 pM.
[1084] In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of between 100 pM and 500 pM, or between 100 pM and 400 pM, or between 100 pM and 300 pM.
[1085] In one embodiment, the ratio of the potency of the IL-2 cytokine against the high affinity (.alpha..beta..gamma.) receptor: affinity of the anti-PD-L1 antigen binding site against hPD-L1 is at least 3:1. In one embodiment, the ratio of the potency of the IL-2 cytokine against the high affinity (.alpha..beta..gamma.) receptor: affinity of the anti-PD-L1 antigen binding site against hPD-L1 is at least 4:1. In one embodiment, the ratio of the potency of the IL-2 cytokine against the high affinity (.alpha..beta..gamma.) receptor: affinity of the anti-PD-L1 antigen binding site against hPD-L1 is at least 5:1. In one embodiment, the ratio of the potency of the IL-2 cytokine against the high affinity (.alpha..beta..gamma.) receptor: affinity of the anti-PD-L1 antigen binding site against hPD-L1 is at least 7:1. In one embodiment, the ratio of the potency of the IL-2 cytokine against the high affinity (.alpha..beta..gamma.) receptor: affinity of the anti-PD-L1 antigen binding site against hPD-L1 is at least 10:1.
[1086] Any of the half-life, K.sub.ON rates, K.sub.OFF rates, or binding characteristics of the anti-PD-L1 antibodies in concepts 1 to 40 apply mutatis mutandis to the immunocytokines disclosed herein.
Aspect 50a. An immunocytokine according to any preceding aspect, wherein the antigen binding site binds to mPD-L1 (Seq ID No:325) with an affinity of less than 500 nM (e.g. less than 100 nM, less than 10 nM or less than 1 nm).
[1087] In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of between 1 nM and 500 nM, or between 1 nM and 250 nM, or between 1 nM and 100 nM, or between 1 nM and 50 nM.
[1088] In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of between 10 nM and 500 nM, or between 10 nM and 250 nM, or between 10 nM and 100 nM, or between 1 nM and 50 nM, in particular between 10 nM and 100 nM.
[1089] In one embodiment, the antigen binding site binds to hPD-L1 with an affinity of between 100 nM and 500 nM, or between 100 nM and 400 nM, or between 100 nM and 300 nM, or between 100 nM and 200 nM.
[1090] The affinity of the antigen-binding site to hPD-L1 or mPD-L1 may be measured by any technique well-known to those skilled in the art. In one embodiment, the affinity is measured using SPR, the details of which are provided hereinabove.
Aspect 51. An immunocytokine as defined in any preceding aspect for use in treating or preventing a hPD-L1-mediated disease or condition, e.g. selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease, diffuse large B-cell lymphoma (for example melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas). Aspect 52. Use of an immunocytokine as defined in any one of aspects 1 to 50 in the manufacture of a medicament for administration to a human for treating or preventing a hPD-L1 mediated disease or condition in the human, e.g. wherein the hPD-L1 mediated disease or condition is selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease, diffuse large B-cell lymphoma (for example melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas). Aspect 53. A method of treating or preventing a hPD-L1 mediated disease or condition, e.g. selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease, diffuse large B-cell lymphoma (for example melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas) in a human, comprising administering to said human a therapeutically effective amount of an immunocytokine as defined in any one of aspects 1 to 50, wherein the hPD-L1 mediated disease or condition is thereby treated or prevented.
[1091] In any of aspects 51 to 53, the hPD-L1 mediated disease may be any of those as described herein. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a virally induced cancer, such as cervical cancer and nasopharyngeal cancer, for example cervical cancers caused by HPV infection. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a chronic viral infection. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a neoplastic disease. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a non-neoplastic disease. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a malignant tumour. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a cancer which is known to be responsive to PD-L1 therapy, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a cancer which is a soft tissue sarcoma. In one embodiment, in any of aspects 51 to 53, the hPD-L1 mediated disease is a neurodegenerative disease, disorder or condition, optionally wherein the neurodegenerative disease, disorder or condition is selected from Alzheimer's disease, amyotrophic lateral sclerosis, Parkinson's disease, Huntington's disease, primary progressive multiple sclerosis, secondary progressive multiple sclerosis, corticobasal degeneration, Rett syndrome, a retinal degeneration disorder selected from age-related macular degeneration and retinitis pigmentosa; anterior ischemic optic neuropathy, glaucoma, uveitis, depression, trauma-associated stress or post-traumatic stress disorder, frontotemporal dementia, Lewy body dementias, mild cognitive impairments, posterior cortical atrophy, primary progressive aphasia and progressive supranuclear palsy or aged-related dementia, in particular Alzheimer's disease, amyotrophic lateral sclerosis, Parkinson's disease and Huntington's disease, and e.g. Alzheimer's disease.
[1092] Aspect 54. The immunocytokine according to aspect 51, the use according to aspect 52 or the method according to aspect 53, wherein the hPD-L1-mediated disease or condition is cancer.
Aspect 55. The immunocytokine, the use or the method according to aspect 54, wherein the cancer is selected from melanoma, Merkel cell cancer, non-small cell lung cancer, bladder cancer, Non-Hodgkin's lymphomas, colorectal cancer with microsatellite instability (MSI) or a cancer selected from breast cancer, ovarian cancer, colorectal cancer (without MSI or microsatellite instability), in particular melanoma and renal cell cancer.
[1093] In one embodiment, the cancer is a cancer which is known to be responsive to both IL-2 therapy and PD-L1 therapy, such as melanoma and renal cell cancer.
[1094] In one embodiment, the cancer is colorectal cancer with microsatellite instability (MSI). In one embodiment, the cancer is breast cancer. In one embodiment, the cancer is ovarian cancer.
Aspect 56. The immunocytokine, use or the method according to any one of aspects 51 to 55, further comprising administering to the human a further therapy, for example a further therapeutic agent, optionally wherein the further therapeutic agent is independently selected from the group consisting of: [1095] A) other immune checkpoint inhibitors (such as anti-TIM-3 antibodies, anti-CTLA-4 antibodies, anti-TIGIT antibodies and anti-LAG-3 antibodies); [1096] B) immune stimulators (such as anti-OX40 antibodies, anti-GITR antibodies, anti-CD137 antibodies, anti-ICOS antibodies and anti-CD40 antibodies); [1097] C) chemokine receptor antagonists (such as CXCR4, CCR4 and CXCR2); [1098] D) targeted kinase inhibitors (such as CSF-1R or VEGFR inhibitors); [1099] E) angiogenesis inhibitors (such as anti-VEGF-A or Delta-like Ligand-4); [1100] F) immune stimulating peptides or chemokines (such as CXCL9 or CXCL10); [1101] G) cytokines (such as IL-15 and IL-21); [1102] H) bispecific T-cell engagers (Bins) having at least one specificity against CD3 (e.g. CD3/CD19 BiTE); [1103] I) other bi-specific molecules (for example IL-15-containing molecules targeted towards tumour associated antigens, for example Epidermal growth factor receptors such as EGFR, Her-2, New York Esophageal Cancer-1 (NY-ESO-1), GD2, EpCAM or Melanoma Associated Antigen-3 (MAGE-A3)); [1104] J) oncolytic viruses (such as HSV virus (optionally which secretes GMCSF), Newcastle disease virus and Vaccinia virus); [1105] K) vaccination with tumour associated antigens (such as New York Esophageal Cancer-1 [NY-ESO-1], Melanoma Associated Antigen-3 [MAGE-3]); [1106] L) cell-based therapies (such as chimeric Antigen Receptor-T-cells (CAR-T) for example expressing anti-CD19, anti-EpCam or anti-mesothelin); [1107] M) bi-specific NK cell engagers having a specificity against an activating MK receptor such as NKG2D or CD16a; and [1108] N) adoptive transfer of tumour specific T-cells or LAK cells, or optionally wherein the further therapy is chemotherapy, radiotherapy and surgical removal of tumours.
[1109] Radiotherapy may be single dose or in fractionated doses, either delivered to affected tissues directly or to the whole body.
[1110] In this aspect, any of the features and embodiments of concept 46 apply mutatis mutandis
[1111] In this aspect, the bispecific molecules include "bispecific antibodies" and antibody fusion proteins, including those formats and molecules described in concepts 37 to 40.
[1112] The antibodies may be any of the sequences or antibodies described in arrangement 5, 5a or detailed in Aspect 1a.
[1113] Aspect 57. A pharmaceutical composition comprising an immunocytokine as defined in any one of aspects 1 to 50 and a pharmaceutically acceptable excipient, diluent or carrier and optionally further comprising a further therapeutic agent independently selected from the group consisting of: [1114] A) other immune checkpoint inhibitors (such as anti-TIM-3 antibodies, anti-CTLA-4 antibodies, anti-TIGIT antibodies and anti-LAG-3 antibodies); [1115] B) immune stimulators (such as anti-OX40 antibodies, anti-G1TR antibodies, anti-CD137 antibodies, anti-ICOS antibodies and anti-CD40 antibodies); [1116] C) chemokine receptor antagonists (such as CXCR4, CCR4 and CXCR2); [1117] D) targeted kinase inhibitors (such as CSF-1R or VEGFR inhibitors); [1118] E) angiogenesis inhibitors (such as anti-VEGF-A or Delta-like Ligand-4); [1119] F) immune stimulating peptides or chemokines (such as CXCL9 or CXCL10); [1120] G) cytokines (such as IL-15 and IL-21); [1121] H) bispecific T-cell engagers (BiTEs) having at least one specificity against CD3 (e.g. CD3/CD19 BITE); [1122] I) other bi-specific molecules (for example IL-15-containing molecules targeted towards tumour associated antigens, for example Epidermal growth factor receptors such as EGFR, Her-2, New York Esophageal Cancer-1 (NY-ESO-1), GD2, EpCAM or Melanoma Associated Antigen-3 (MAGE-A3)); [1123] J) oncolytic viruses (such as HSV virus (optionally which secretes GMCSF), Newcastle disease virus and Vaccinia virus); [1124] K) vaccination with tumour associated antigens (such as New York Esophageal Cancer-1 [NY-ESO-1], Melanoma Associated Antigen-3 [MAGE-3]); [1125] L) cell-based therapies (such as chimeric Antigen Receptor-T-cells (CAR-T) for example expressing anti-CD19, anti-EpCam or anti-mesothelin); [1126] M) bi-specific NK cell engagers having a specificity against an activating MK receptor such as NKG2D or CD16a; and [1127] N) adoptive transfer of tumour specific T-cells or LAK cells.
[1128] In one embodiment, the further therapeutic agent is administered sequentially or simultaneously with the immunocytokine.
[1129] In this aspect, any of the features and embodiments of concept 48 apply mutatis mutandis
[1130] In this aspect, the bispecific molecules include "bispecific antibodies" and antibody fusion proteins, including those formats and molecules described in concepts 37 to 40.
[1131] The antibodies may be any of the sequences or antibodies described in arrangement 5, 5a or detailed in Aspect 1a.
Aspect 58. A pharmaceutical composition according to aspect 57, or a kit comprising a pharmaceutical composition as defined in aspect 57, wherein the composition is for treating and/or preventing a hPD-L1 mediated disease or condition, e.g. selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease, diffuse large B-cell lymphoma (for example melanoma, breast cancer, ovarian cancer, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, colorectal cancer (without MSI or microsatellite instability), head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas). Aspect 59. A pharmaceutical composition according to aspect 57 or aspect 58 in combination with, or kit according to aspect 58 comprising a label or instructions for use to treat and/or prevent said disease or condition in a human; optionally wherein the label or instructions comprise a marketing authorisation number (e.g., an FDA or EMA authorisation number); optionally wherein the kit comprises an IV or injection device that comprises the immunocytokine. Aspect 60. A method of treating a proliferative disease in an animal (e.g. a human), comprising administering an effective amount of an immunocytokine as defined in any one of aspects 1 to 50 to said patient.
[1132] Proliferative diseases may be any as described elsewhere herein.
Aspect 61. A nucleic acid that encodes a heavy chain and/or a light chain of an immunocytokine as defined in any one of aspects 1 to 50.
[1133] In one embodiment, the nucleic acid encodes a light chain of an immunocytokine as defined in any one of aspects 1 to 50.
Aspect 62. A vector comprising the nucleic acid as defined in aspect 61; optionally wherein the vector is a CHO or HEK293 vector: Aspect 63. A host comprising the nucleic acid as defined in aspect 61 or the vector as defined in aspect 62.
4. ICOS Antibodies
[1134] ICOS antibodies are provided herein. The ICOS antibodies may be any of those described in GB patent application 1620414.1 (filed 1 Dec. 2016), the sequences of the anti-ICOS antibodies disclosed therein are incorporated herein by reference.
[1135] STIM001 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:366, comprising the CDRH1 amino acid sequence of Seq ID No:363, the CDRH2 amino acid sequence of Seq ID No:364, and the CDRH3 amino acid sequence of Seq ID No:365. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:367. STIM001 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:373, comprising the CDRL1 amino acid sequence of Seq ID No:370, the CDRL2 amino acid sequence of Seq ID No:371, and the CDRL3 amino acid sequence of Seq ID No:372. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:374. The V.sub.H, domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:368 (heavy chain nucleic acid sequence Seq ID No:369). A full length light chain amino acid sequence is Seq ID No:375 (light chain nucleic acid sequence Seq ID No:376).
[1136] STIM002 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:380, comprising the CDRH1 amino acid sequence of Seq ID No:377, the CDRH2 amino acid sequence of Seq ID No:378, and the CDRH3 amino acid sequence of Seq ID No:379. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:381. STIM002 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:387, comprising the CDRL1 amino acid sequence of Seq ID No:384, the CDRL2 amino acid sequence of Seq ID No:385, and the CDRL3 amino acid sequence of Seq ID No:386. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:388 or Seq ID No:519. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:382 (heavy chain nucleic acid sequence Seq ID No:383). A full length light chain amino acid sequence is Seq ID No:389 (light chain nucleic acid sequence Seq ID No:390 or or Seq ID No:520).
[1137] STIM002-B has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:394, comprising the CDRH1 amino acid sequence of Seq ID No:391, the CDRH2 amino acid sequence of Seq ID No:392, and the CDRH3 amino acid sequence of Seq ID No:393. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:395. STIM002-13 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:401, comprising the CDRL1 amino acid sequence of Seq ID No:398, the CDRL2 amino acid sequence of Seq ID No:399, and the CDRL3 amino acid sequence of Seq ID No:400. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:402. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:396 (heavy chain nucleic acid sequence Seq ID No:397). A full length light chain amino acid sequence is Seq ID No:403 (light chain nucleic acid sequence Seq ID No:404).
[1138] STIM003 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:408, comprising the CDRH1 amino acid sequence of Seq ID No:405, the CDRH2 amino acid sequence of Seq ID No:406, and the CDRH3 amino acid sequence of Seq ID No:407. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:409 or Seq ID No:521. STIM003 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:415, comprising the CDRL1 amino acid sequence of Seq ID No:412, the CDRL2 amino acid sequence of Seq ID No:413, and the CDRL3 amino acid sequence of Seq ID No:414. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:4416. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:410 (heavy chain nucleic acid sequence Seq ID No:411 or Seq ID No:522). A full length light chain amino acid sequence is Seq ID No:417 (light chain nucleic acid sequence Seq ID No:418).
[1139] STIM004 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:422, comprising the CDRH1 amino acid sequence of Seq ID No:419, the CDRH2 amino acid sequence of Seq ID No:420, and the CDRH3 amino acid sequence of Seq ID No:421. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:423. STIM004 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:429, comprising the CDRL1 amino acid sequence of Seq ID No:426, the CDRL2 amino acid sequence of Seq ID No:427, and the CDRL3 amino acid sequence of Seq ID No:428. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:430 or Seq ID No:431. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:424 (heavy chain nucleic acid sequence Seq ID No:425). A full length light chain amino acid sequence is Seq ID No:432 (light chain nucleic acid sequence Seq ID No:433 or Seq ID no: 434).
[1140] STIM005 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:438, comprising the CDRH1 amino acid sequence of Seq ID No:435, the CDRH2 amino acid sequence of Seq ID No:436, and the CDRH3 amino acid sequence of Seq ID No:437. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:439. STIM005 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:445, comprising the CDRL1 amino acid sequence of Seq ID No:442, the CDRL2 amino acid sequence of Seq ID No:443, and the CDRL3 amino acid sequence of Seq ID No:444. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:446. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:440 (heavy chain nucleic acid sequence Seq ID No:441). A full length light chain amino acid sequence is Seq ID No:447 (light chain nucleic acid sequence Seq ID No:448).
[1141] STIM006 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:452, comprising the CDRH1 amino acid sequence of Seq ID No:449, the CDRH2 amino acid sequence of Seq ID No:450, and the CDRH3 amino acid sequence of Seq ID No:451. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:453. STIM006 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:459, comprising the CDRL1 amino acid sequence of Seq ID No:456, the CDRL2 amino acid sequence of Seq ID No:457, and the CDRL3 amino acid sequence of Seq ID No:458. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:460. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:454 (heavy chain nucleic acid sequence Seq ID No:455). A full length light chain amino acid sequence is Seq ID No:461 (light chain nucleic acid sequence Seq ID No:462).
[1142] STIM007 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:466, comprising the CDRH1 amino acid sequence of Seq ID No:463, the CDRH2 amino acid sequence of Seq ID No:464, and the CDRH3 amino acid sequence of Seq ID No:465. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:467. 511M007 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:473, comprising the CDRL1 amino acid sequence of Seq ID No:470, the CDRL2 amino acid sequence of Seq ID No:471, and the CDRL3 amino acid sequence of Seq ID No:472. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:474. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:468 (heavy chain nucleic acid sequence Seq ID No:469). A full length light chain amino acid sequence is Seq ID No:475 (light chain nucleic acid sequence Seq ID No:476).
[1143] STIM008 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:480, comprising the CDRH1 amino acid sequence of Seq ID No:477, the CDRH2 amino add sequence of Seq ID No:478, and the CDRH3 amino add sequence of Seq ID No:479. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:481. STIM008 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:487, comprising the CDRL1 amino acid sequence of Seq ID No:484, the CDRL2 amino acid sequence of Seq ID No:485, and the CDRL3 amino acid sequence of Seq ID No:486. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:488. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:482 (heavy chain nucleic acid sequence Seq ID No:483). A full length light chain amino acid sequence is Seq ID No:489 (light chain nucleic acid sequence Seq ID No:490).
[1144] STIM009 has a heavy chain variable region (V.sub.H) amino acid sequence of Seq ID No:494, comprising the CDRH1 amino acid sequence of Seq ID No:491, the CDRH2 amino acid sequence of Seq ID No:492, and the CDRH3 amino acid sequence of Seq ID No:493. The heavy chain nucleic acid sequence of the V.sub.H domain is Seq ID No:495. STIM009 has a light chain variable region (V.sub.L) amino acid sequence of Seq ID No:501, comprising the CDRL1 amino acid sequence of Seq ID No:498, the CDRL2 amino acid sequence of Seq ID No:499, and the CDRL3 amino acid sequence of Seq ID No:500. The light chain nucleic acid sequence of the V.sub.L domain is Seq ID No:502. The V.sub.H domain may be combined with any of the heavy chain constant region sequences described herein, e.g. Seq ID No:193, Seq ID No:195, Seq ID No:197, Seq ID No:199, Seq ID No:201, Seq ID No:203, Seq ID No:205, Seq ID No:340, Seq ID No:524, Seq ID No:526, Seq ID No:528, Seq ID No:530, Seq ID No:532 or Seq ID No:534. The V.sub.L domain may be combined with any of the light chain constant region sequences described herein, e.g. Seq ID Nos:207, 209, 211, 213, 215, 217, 219, 221, 223, 225, 227, 229, 231, 233, 235, 237, 536 and 538. A full length heavy chain amino acid sequence is Seq ID No:496 (heavy chain nucleic acid sequence Seq ID No:497). A full length light chain amino acid sequence is Seq ID No:503 (light chain nucleic acid sequence Seq ID No:504).
[1145] Antibodies STIM001-009 are described in more detail in GB patent application 1620414.1 (filed 1 Dec. 2016), the contents of which are incorporated herein by reference. ICOS antibodies may also be described as in the following numbered sentences below:
Sentence 1. An isolated antibody that binds the extracellular domain of human and/or mouse ICOS, comprising: [1146] an antibody V.sub.H domain comprising complementarity determining regions (CDRs) HCDR1, HCDR2 and HCDR3, and [1147] an antibody V.sub.L domain comprising complementarity determining regions LCDR1, LCDR2 and LCDR3, wherein [1148] HCDR1 is the HCDR1 of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, or comprises that HCDR1 with 1, 2, 3, 4 or 5 amino acid alterations, [1149] HCDR2 is the HCDR2 of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, or comprises that HCDR2 with 1, 2, 3, 4 or 5 amino acid alterations, and/or [1150] HCDR3 is the HCDR3 of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009 or comprises that HCDR3 with 1, 2, 3, 4 or 5 amino acid alterations. Sentence 2. An antibody according to sentence 1, wherein the antibody heavy chain CDRs are those of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009 or comprise the STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009 heavy chain CDRs with 1, 2, 3, 4 or 5 amino acid alterations. Sentence 3. An antibody according to sentence 2, wherein the antibody V.sub.H domain has the heavy chain CDRs of STIM003. Sentence 4. An isolated antibody that binds the extracellular domain of human and/or mouse ICOS, comprising: [1151] an antibody V.sub.H domain comprising complementarity determining regions HCDR1, HCDR2 and HCDR3, and [1152] an antibody V.sub.L domain comprising complementarity determining regions LCDR1, LCDR2 and LCDR3, [1153] wherein LCDR1 is the LCDR1 of STIM001, STIM002, STIM002-13, STIM003, STIM004 STIM005, STIM006, STIM007, STIM008 or STIM009, or comprises that LCDR1 with 1, 2, 3, 4 or 5 amino acid alterations, [1154] LCDR2 is the LCDR2 of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, or comprises that LCDR2 with 1, 2, 3, 4 or 5 amino acid alterations, and/or [1155] LCDR3 is the LCDR3 of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009 or comprises that LCDR3 with 1, 2, 3, 4 or 5 amino acid alterations. Sentence 5. An antibody according to any preceding sentence, wherein the antibody light chain CDRs are those of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, or comprise the STIM001, STIM002, STIM002-13, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009 light chain CDRs with 1, 2, 3, 4 or 5 amino acid alterations. Sentence 6. An antibody according to sentence 5, wherein the antibody V.sub.L domain has the light chain CDRs of STIM003. Sentence 7. An antibody according to any of the preceding sentences, comprising V.sub.H and/or V.sub.L domain framework regions of human germline gene segment sequences. Sentence 8. An antibody according to any of the preceding sentences, comprising a V.sub.H domain which [1156] (i) is derived from recombination of a human heavy chain V gene segment, a human heavy chain D gene segment and a human heavy chain J gene segment, wherein [1157] the V segment is IGHV1-18 (e.g., V1-18*01), IGVH3-20 (e.g. V3-20*d01), IGVH3-11 (e.g, V3-11*01) or IGVH2-5 (e.g., V2-5*10); [1158] the D gene segment is IGHD6-19 (e.g., IGHD6-19*01), IGHD3-10 (e.g., IGHD3-10*01) or IGHD3-9 (e.g., IGHD3-9*01); and/or [1159] the J gene segment is IGHJ6 (e.g., IGHJ6*02), IGHJ4 (e.g., IGHJ4*02) or IGHJ3 (e.g., IGHJ3*02), or [1160] (ii) comprises framework regions FR1, FR2, FR3 and FR4, wherein [1161] FR1 aligns with human germline V gene segment IGHV1-18 (e.g., V1-18*01), IGVH3-20 (e.g. V3-20*d01), IGVH3-11 (e.g, V3-11*01) or IGVH2-5 (e.g., V2-5*10), optionally with 1, 2, 3, 4 or 5 amino acid alterations, [1162] FR2 aligns with human germline V gene segment IGHV1-18 (e.g., V1-18*01), IGVH3-20 (e.g. V3-20*d01), IGVH3-11 (e.g, V3-11*01) or IGVH2-5 (e.g., V2-5*10), optionally with 1, 2, 3, 4 or 5 amino acid alterations, [1163] FR3 aligns with human germline V gene segment IGHV1-18 (e.g., V1-18*01), IGVH3-20 (e.g. V3-20*d01), IGVH3-11 (e.g, V3-11*01) or IGVH2-5 (e.g., V2-5*10), optionally with 1, 2, 3, 4 or 5 amino acid alterations, and/or [1164] FR4 aligns with human germline J gene segment IGJH6 (e.g., JH6*02), IGJH4 (e.g., JH4*02) or IGJH3 (e.g., JH3*02), optionally with 1, 2, 3, 4 or 5 amino acid alterations. Sentence 9. An antibody according to any of the preceding sentences, comprising an antibody V.sub.L domain which [1165] (i) is derived from recombination of a human light chain V gene segment and a human light chain J gene segment, wherein [1166] the V segment is IGKV2-28 (e.g., IGKV2-28*01), IGKV3-20 (e.g., IGKV3-20*01), IGKV1D-39 (e.g., IGKV1D-39*01) or IGKV3-11 (e.g., IGKV3-11*01), and/or [1167] the J gene segment is IGKJ4 (e.g., IGKJ4*01), IGKJ2 (e.g., IGKJ2*04), IGLJ3 (e.g., IGKJ3*01) or IGKJ1 (e.g., IGKJ1*01); or [1168] (ii) comprises framework regions FR1, FR2, FR3 and FR4, wherein [1169] FR1 aligns with human germline V gene segment IGKV2-28 (e.g., IGKV2-28*01), IGKV3-20 (e.g., IGKV3-20*01), IGKV1D-39 (e.g., IGKV1D-39*01) or IGKV3-11 (e.g., IGKV3-11*01), optionally with 1, 2, 3, 4 or 5 amino acid alterations, [1170] FR2 aligns with human germline V gene segment IGKV2-28 (e.g., IGKV2-28*01), IGKV3-20 (e.g., IGKV3-20*01), IGKV1D-39 (e.g., IGKV1D-39*01) or IGKV3-11 (e.g., IGKV3-11*01), optionally with 1, 2, 3, 4 or 5 amino acid alterations, [1171] FR3 aligns with human germline V gene segment IGKV2-28 (e.g., IGKV2-28*01), IGKV3-20 (e.g., IGKV3-20*01), IGKV1D-39 (e.g., IGKV1D-39*01) or IGKV3-11 (e.g., IGKV3-11*01), optionally with 1, 2, 3, 4 or 5 amino acid alterations, and/or [1172] FR4 aligns with human germline J gene segment IGKJ4 (e.g., IGKJ4*01), IGKJ2 (e.g., IGKJ2*04), IGKJ3 (e.g., IGKJ3*01) or IGKJ1 (e.g., IGKJ1*01), optionally with 1, 2, 3, 4 or 5 amino acid alterations. Sentence 10. An antibody according to any of the preceding sentences, comprising an antibody VH domain which is the V.sub.H domain of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, or which has an amino acid sequence at least 90% identical to the antibody VH domain sequence of STIM001, STIM002, STIM002-13, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009. Sentence 11. An antibody according to any of the preceding sentences, comprising an antibody VL domain which is the V.sub.L domain of STIM001, STIM002, STIM002-13, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, or which has an amino acid sequence at least 90% identical to the antibody V.sub.L domain sequence of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009. Sentence 12. An antibody according to sentence 11, comprising [1173] an antibody V.sub.H domain which is selected from the VH domain of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, or which has an amino acid sequence at least 90% identical to the antibody VH domain sequence of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009, and [1174] an antibody V.sub.L domain which is the V.sub.L domain of said selected antibody, or which has an amino acid sequence at least 90% identical to the antibody V.sub.L domain sequence of said selected antibody. Sentence 13. An antibody according to sentence 12, comprising the STIM003 V.sub.H domain and the STIM003 V.sub.L domain. Sentence 14. An antibody according to any of the preceding sentences, comprising an antibody constant region. Sentence 15. An antibody according to sentence 14, wherein the constant region comprises a human heavy and/or light chain constant region. Sentence 16. An antibody according to sentence 14 or sentence 15, wherein the constant region is Fc effector positive. Sentence 17. An antibody according to sentence 16, comprising an Fc region that has enhanced ADCC, ADCP and/or CDC function compared with a native human Fc region. Sentence 18. An antibody according to any of sentences 14 to 17, wherein the antibody is an IgG1. Sentence 19. An antibody according to sentence 17 or sentence 18, wherein the antibody is afucosylated. Sentence 20. An antibody according to any of the preceding sentences which is conjugated to a cytotoxic drug or pro-drug. Sentence 21. An antibody according to any of the preceding sentences, which is a multispecific antibody. Sentence 22. An isolated antibody that competes for binding to human ICOS with a human IgG1 antibody comprising the heavy and light chain complementarity determining regions of STIM001, STIM002, STIM002-B, STIM003, STIM004, STIM005, STIM006, STIM007, STIM008 or STIM009. Sentence 23. An isolated antibody that binds the extracellular domain of human and mouse ICOS with an affinity (KD) of less than 50 nM as determined by surface plasmon resonance. Sentence 24. An antibody according to sentence 23, wherein the antibody binds the extracellular domain of human and mouse ICOS with an affinity (KD) of less than 5 nM as determined by surface plasmon resonance. Sentence 25. An antibody according to sentence 23 or sentence 24, wherein the KD of binding the extracellular domain of human ICOS is within 10-fold of the KD of binding the extracellular domain of mouse ICOS. Sentence 26. A composition comprising an isolated antibody according to any of the preceding sentences and a pharmaceutically acceptable excipient. Sentence 27. A composition comprising isolated nucleic acid encoding an antibody according to any of sentences 1 to 25 and a pharmaceutically acceptable excipient. Sentence 28. A method of depleting regulatory T-cells and/or increasing effector T-cell response in a patient comprising administering a composition according to sentence 26 to the patient. Sentence 29. A method of treating a disease or condition amenable to therapy by depleting regulatory T-cells and/or increasing effector T-cell response in a patient, the method comprising administering a composition according to sentence 26 to the patient. Sentence 30. A composition according to sentence 26 for use in a method of treatment of the human body by therapy. Sentence 31. A composition for use according to sentence 30, for use in depleting regulatory T-cells and/or increasing effector T-cell response in a patient. Sentence 32. A composition for use according to sentence 30, for use in treating a disease or condition amenable to therapy by depleting regulatory T-cells and/or increasing effector T-cell response in a patient. Sentence 33. A method according to sentence 29, or a composition for use according to sentence 32, wherein the disease is a cancer or a solid tumour. Sentence 34. A method or a composition for use according to any of sentences 29 to 33, wherein the method comprises administering the antibody and another therapeutic agent to the patient. Sentence 35. A method or composition for use according to sentence 34, wherein the therapeutic agent is an anti-PDL1 antibody. Sentence 36. A method or composition for use according to sentence 35, wherein the anti-ICOS antibody and the anti-PDL1 antibody are each able to mediate ADCC, ADCP and/or CDC. Sentence 37. A method or composition for use according to sentence 35, wherein the anti-ICOS antibody is a human IgG1 antibody and the anti-PDL1 antibody is a human IgG1 antibody. Sentence 38. A method or composition for use according to sentence 34, wherein the other therapeutic agent is IL-2. Sentence 39. A method or composition for use according to any of sentences 34 to 38, wherein the method comprises administering the anti-ICOS antibody after administering the other therapeutic agent. Sentence 40. A method or a composition for use according to any of sentences 28 to 39, wherein the anti-ICOS antibody is conjugated to a pro-drug, and wherein the method or use comprises administering the anti-ICOS antibody to a patient and selectively activating the pro-drug at a target tissue site. Sentence 41. A method or a composition for use according to sentence 40, wherein the patient has a solid tumour and the method comprises selectively activating the pro-drug in the tumour. Sentence 42. A method or a composition for use according to sentence 40 or sentence 41, comprising selectively activating the pro-drug through photoactivation. Sentence 43. Combination of anti-ICOS human IgG1 antibody and anti-PDL1 human IgG1 antibody for use in a method of treating cancer. Sentence 44. Combination according to sentence 43, wherein the anti-ICOS antibody and the anti-PDL1 antibody are provided in separate compositions for administration. Sentence 45. A method or composition for use according to sentence 37, or a combination according to sentence 43 or sentence 44, wherein the human IgG1 constant region has a wild type amino acid sequence shown in the appended sequence listing. Sentence 46. Anti-ICOS antibody for use in a method of reducing or reversing a surge in ICOS-positive regulatory T-cells in a patient, wherein the surge results from treatment of the patient with another therapeutic agent. Sentence 47. A method of treating a patient, the method comprising reducing or reversing a surge in ICOS-positive regulatory T-cells in the patient, wherein the surge results from treatment of the patient with another therapeutic agent. Sentence 48. Anti-ICOS antibody for use in a method of treating a patient, the method comprising comprising administering the anti-ICOS antibody to a patient who has an increased level of ICOS-positive regulatory T-cells following treatment with another therapeutic agent. Sentence 49. A method of treating a patient, the method comprising administering an anti-ICOS antibody to a patient who has an increased level of ICOS-positive regulatory T-cells following treatment with another therapeutic agent. Sentence 50. An anti-ICOS antibody for use according to sentence 46 or sentence 48, or a method according to sentence 47 or sentence 49, wherein the method comprises administering a therapeutic agent to the patient, determining that the patient has an increased level of ICOS-positive regulatory T-cells following the treatment with said agent, and administering an anti-ICOS antibody to the patient to reduce the level of regulatory T-cells. Sentence 51. An anti-ICOS antibody for use or a method according to any of sentences 46 to 50, wherein the therapeutic agent is IL-2 or an immunomodulatory antibody (e.g., anti-PDL-1, anti-PD-1 or anti-CTLA-4). Sentence 52. An anti-ICOS antibody for use or a method according to any of sentences 46 to 51, wherein the method comprises treating a tumour, e.g., melanoma, such as metastatic melanoma. Sentence 53. Anti-ICOS antibody for use in a method of treating cancer in a patient by in vivo vaccination of the patient against their cancer cells, the method comprising
[1175] treating the patient with a therapy that causes immunological cell death of the cancer cells, resulting in presentation of antigen to antigen-specific effector T-cells, and
[1176] administering an anti-ICOS antibody to the patient, wherein the anti-ICOS antibody enhances the antigen-specific effector T-cell response.
Sentence 54. A method of treating cancer in a patient by in vivo vaccination of the patient against their cancer cells, the method comprising
[1177] treating the patient with a therapy that causes immunological cell death of the cancer cells, resulting in presentation of antigen to antigen-specific effector T-cells, and
[1178] administering an anti-ICOS antibody to the patient, wherein the anti-ICOS antibody enhances the antigen-specific effector T-cell response.
Sentence 55. A method of treating cancer in a patient by in vivo vaccination of the patient against their cancer cells, the method comprising administering an anti-ICOS antibody to the patient, wherein [1179] the patient is one who has been previously treated with a therapy that causes immunological cell death of the cancer cells, resulting in presentation of antigen to antigen-specific effector T-cells, and wherein [1180] the anti-ICOS antibody enhances the antigen-specific effector T-cell response. Sentence 56. Anti-ICOS antibody for use or a method according to any of sentences 53 to 55, wherein the therapy that causes immunological cell death is radiation of the cancer cells, administration of a chemotherapeutic agent and/or administration of an antibody directed to a tumour-associated antigen. Sentence 57. Anti-ICOS antibody for use or a method according to sentence 56, wherein the chemotherapeutic agent is oxaliplatin. Sentence 58. Anti-ICOS antibody for use or a method according to sentence 56, wherein the tumour-associated antigen is HER2 or CD20. Sentence 59. Anti-ICOS antibody for use in a method of vaccinating a patient, the method comprising administering the antibody and a vaccine composition to the patient. Sentence 60. A method of vaccinating a patient, the method comprising administering an anti-ICOS antibody and a vaccine composition to the patient. Sentence 61. Anti-ICOS antibody for use according to sentence 59, or a method according to sentence 60, wherein the vaccine composition is a vaccine against hepatitis B, malaria or HIV. Sentence 62. Anti-ICOS antibody for use in a method of treating a cancer in a patient, wherein the cancer is or has been characterised as being positive for expression of ICOS ligand and/or FOXP3. Sentence 63. A method of treating a cancer in a patient, wherein the cancer is or has been characterised as being positive for expression of ICOS ligand and/or FOXP3, the method comprising administering an anti-ICOS antibody to the patient. Sentence 64. Anti-ICOS antibody for use according to sentence 62, or a method according to sentence 63, wherein the method comprises: [1181] testing a sample from a patient to determine that the cancer expresses ICOS ligand and/or FOXP3; [1182] selecting the patient for treatment with the anti-ICOS antibody; and [1183] administering the anti-ICOS antibody to the patient. Sentence 65. Anti-ICOS antibody for use according to sentence 62, or a method according to sentence 63, wherein the method comprises administering an anti-ICOS antibody to a patient from whom a test sample has indicated that the cancer is positive for expression of ICOS ligand and/or FOXP3. Sentence 66. Anti-ICOS antibody for use or a method according to sentence 64 or sentence 65, wherein the sample is biopsy sample of a solid tumour. Sentence 67. Anti-ICOS antibody for use in a method of treating a cancer in a patient, wherein the cancer is or has been characterised as being refractory to treatment with an immunooncology drug, e.g., anti-CTLA-4 antibody, anti-PD1 antibody, anti-PD-L1 antibody, anti-CD137 antibody or anti-GITR antibody. Sentence 68. A method of treating a cancer in a patient, wherein the cancer is or has been characterised as being refractory to treatment with an immunooncology drug, e.g., anti-CRA-4 antibody, anti-PD1 antibody, anti-PD-L1 antibody, anti-CD137 antibody or anti-GITR antibody, the method comprising administering an anti-ICOS antibody to the patient. Sentence 69. Anti-ICOS antibody for use according to sentence 67 or a method according to sentence 68, wherein the method comprises:
[1184] treating the patient with the immunooncology drug;
[1185] determining that the cancer is not responsive to the drug;
[1186] selecting the patient for treatment with the anti-ICOS antibody; and
[1187] administering the anti-ICOS antibody to the patient.
Sentence 70. Anti-ICOS antibody for use according to sentence 67, or a method according to sentence 68, wherein the method comprises administering an anti-ICOS antibody to a patient whose cancer was not responsive to prior treatment with the immunooncology drug. Sentence 71. Anti-ICOS antibody for use or a method according to any of sentences 62 to 70, wherein the cancer is a tumour derived from cells that have acquired ability to express ICOS ligand. Sentence 72. Anti-ICOS antibody for use or a method according to sentence 71, wherein the cancer is melanoma. Sentence 73. Anti-ICOS antibody for use or a method according to any of sentences 62 to 70, wherein the cancer is derived from an antigen-presenting cell, such as a B lymphocyte (e.g., B cell lymphoma, such as diffused large B cell lymphoma) or a T lymphocyte. Sentence 74. Anti-ICOS antibody for use or a method according to any of sentences 62 to 70, wherein the cancer is resistant to treatment with an anti-CD20 antibody. Sentence 75. Anti-ICOS antibody for use or a method according to sentence 74, wherein the cancer is B cell lymphoma. Sentence 76. Anti-ICOS antibody for use or a method according to sentence 75, wherein the anti-CD20 antibody is rituximab. Sentence 77. Anti-ICOS antibody for use or a method according to any of sentences 74 to 76, wherein the method comprises treating the patient with the anti-CD20 antibody;
[1188] determining that the cancer is not responsive to the anti-CD20 antibody;
[1189] testing a sample from a patient to determine that the cancer expresses ICOS ligand;
[1190] selecting the patient for treatment with the anti-ICOS antibody; and
[1191] administering the anti-ICOS antibody to the patient.
Sentence 78. Anti-ICOS antibody for use or a method according to any of sentences 74 to 76, wherein the method comprises administering an anti-ICOS antibody to a patient whose cancer was not responsive to prior treatment with anti-CD20 antibody. Sentence 79. Anti-ICOS antibody for use or a method according to any of sentences 52 to 78, wherein the cancer is a solid tumour. Sentence 80. Anti-ICOS antibody for use or a method according to any of sentences 52 to 78, wherein the cancer is a haemotological liquid tumour. Sentence 81. Anti-ICOS antibody for use or a method according to sentence 79 or 80, wherein the tumour is high in regulatory T-cells. Sentence 82. Anti-ICOS antibody for use or a method according to any of sentences 43 to 81, wherein the anti-ICOS antibody is as defined in any of sentences 1 to 25 or is provided in a composition according to sentence 26. Sentence 83. A transgenic non-human mammal having a genome comprising a human or humanised immunoglobulin locus encoding human variable region gene segments, wherein the mammal does not express ICOS. Sentence 84. A method of producing an antibody that binds the extracellular domain of human and non-human ICOS, comprising
[1192] (a) immunising a mammal according to sentence 83 with human ICOS antigen;
[1193] (b) isolating antibodies generated by the mammal;
[1194] (c) testing the antibodies for ability to bind human ICOS and non-human ICOS; and
[1195] (d) selecting one or more antibodies that binds both human and non-human ICOS.
Sentence 85. A method according to sentence 84, comprising immunising the mammal with cells expressing human ICOS. Sentence 86. A method according to sentence 84 or sentence 85, comprising [1196] (c) testing the antibodies for ability to bind human ICOS and non-human ICOS using surface plasmon resonance and determining binding affinities; and [1197] (d) selecting one or more antibodies for which the KD of binding to human ICOS is less than 50 nM and the KD of binding to non-human ICOS is less than 500 nM. Sentence 87. A method according to sentence 86, comprising [1198] (d) selecting one or more antibodies for which the KD of binding to human ICOS is less than 10 nM and the KD of binding to non-human ICOS is less than 100 nM. Sentence 88. A method according to any of sentences 84 to 87, comprising [1199] (c) testing the antibodies for ability to bind human ICOS and non-human ICOS using surface plasmon resonance and determining binding affinities; and [1200] (d) selecting one or more antibodies for which the KD of binding to human ICOS is within 10-fold of the KD of binding to non-human ICOS. Sentence 89. A method according to sentence 88, comprising [1201] (d) selecting one or more antibodies for which the KD of binding to human ICOS is within 5-fold of the KD of binding to non-human ICOS. Sentence 90. A method according to any of sentences 84 to 89, comprising testing the antibodies for ability to bind non-human ICOS from the same species as the mammal. Sentence 91. A method according to any of sentences 84 to 90, comprising testing the antibodies for ability to bind non-human ICOS from a different species as the mammal. Sentence 92. A method according to any of sentences 84 to 91, wherein the mammal is a mouse or a rat. Sentence 93. A method according to any of sentences 84 to 92, wherein the non-human ICOS is mouse ICOS or rat ICOS. Sentence 94. A method according to any of sentences 84 to 93, wherein the human or humanised immunoglobulin locus comprises human variable region gene segments upstream of an endogenous constant region. Sentence 95. A method according to sentence 94, comprising (a) immunising a mammal according to sentence 83 with human ICOS antigen, wherein the mammal is a mouse; (b) isolating antibodies generated by the mouse; (c) testing the antibodies for ability to bind human ICOS and mouse ICOS; and (d) selecting one or more antibodies that binds both human and mouse ICOS. Sentence 96. A method according to any of sentences 84 to 95, comprising isolating nucleic acid encoding an antibody heavy chain variable domain and/or an antibody light chain variable domain. Sentence 97. A method according to any of sentences 84 to 96, wherein the mammal generates antibodies through recombination of human variable region gene segments and an endogenous constant region. Sentence 98. A method according to sentence 96 or sentence 97, comprising conjugating the nucleic acid encoding the heavy and/or light chain variable domain to a nucleotide sequence encoding a human heavy chain constant region and/or human light chain constant region respectively. Sentence 99. A method according to any of sentences 96 to 98, comprising introducing the nucleic acid into a host cell. Sentence 100. A method according to sentence 99, comprising culturing the host cell under conditions for expression of the antibody, or of the antibody heavy and/or light chain variable domain. Sentence 101. An antibody, or antibody heavy and/or light chain variable domain, produced by the method according to any of sentences 84 to 100. Sentence 102. A method of selecting an antibody that binds ICOS, optionally for selecting an ICOS agonist antibody, the assay comprising:
[1202] providing an array of antibodies immobilised (attached or adhered) to a substrate in a test well;
[1203] adding ICOS-expressing cells (e.g., activated primary T-cells, or MJ cells) to the test well;
[1204] observing morphology of the cells;
[1205] detecting shape change in the cells from rounded to flattened against the substrate within the well; wherein the shape change indicates that the antibody is an antibody that binds ICOS, optionally an ICOS agonist antibody;
[1206] selecting the antibody from the test well;
expressing nucleic acid encoding the CDRs of the selected antibody; and formulating the antibody into a composition comprising one or more additional components.
[1207] Alternative sentences describing anti-ICOS antibodies are described below:
Sentence 1a. An antibody or a fragment thereof which specifically binds to human ICOS (hICOS) (SEQ ID NO: 508, 507 and/or 506), and: [1208] a) competes for binding to said hICOS with the antibody STIM001, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:365, or the CDRH3 sequence of SEQ ID NO:365 comprising 3, 2 or 1 amino acid substitution(s); [1209] b) competes for binding to said hICOS with the antibody STIM002, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:379, or the CDRH3 sequence of SEQ ID NO:379 comprising 3, 2 or 1 amino acid substitution(s); [1210] c) competes for binding to said hICOS with the antibody STIM002-B, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:393, or the CDRH3 sequence of SEQ ID NO:393 comprising 3, 2 or 1 amino acid substitution(s); [1211] d) competes for binding to said hICOS with the antibody STIM003, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:407, or the CDRH3 sequence of SEQ ID NO:407 comprising 3, 2 or 1 amino acid substitution(s); [1212] e) competes for binding to said hICOS with the antibody STIM004, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:421, or the CDRH3 sequence of SEQ ID NO:421 comprising 3, 2 or 1 amino acid substitution(s); [1213] f) competes for binding to said hICOS with the antibody STIM005, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:437, or the CDRH3 sequence of SEQ ID NO:437 comprising 3, 2 or 1 amino acid substitution(s); [1214] g) competes for binding to said hICOS with the antibody STIM006, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:451, or the CDRH3 sequence of SEQ ID NO:451 comprising 3, 2 or 1 amino acid substitution(s); [1215] h) competes for binding to said hICOS with the antibody STIM007, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:465, or the CDRH3 sequence of SEQ ID NO:465 comprising 3, 2 or 1 amino acid substitution(s); [1216] i) competes for binding to said hICOS with the antibody STIM008, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:479, or the CDRH3 sequence of SEQ ID NO:479 comprising 3, 2 or 1 amino acid substitution(s); or [1217] j) competes for binding to said hICOS with the antibody STIM009, wherein the antibody or fragment comprises a V.sub.H domain which comprises the CDRH3 sequence of SEQ ID NO:493, or the CDRH3 sequence of SEQ ID NO:493 comprising 3, 2 or 1 amino acid substitution(s). Sentence 2a. The antibody or a fragment thereof according to sentence 1a, wherein the V.sub.H domain comprises the CDRH1 sequence of: [1218] a) SEQ ID NO:363, or the CDRH1 sequence of SEQ ID NO:363 comprising 1 amino acid substitution; [1219] b) SEQ ID NO:377, or the CDRH1 sequence of SEQ ID NO:377 comprising 1 amino acid substitution; [1220] c) SEQ ID NO:391, or the CDRH1 sequence of SEQ ID NO:391 comprising 1 amino acid substitution; [1221] d) SEQ ID NO:405, or the CDRH1 sequence of SEQ ID NO:405 comprising 1 amino acid substitution; [1222] e) SEQ ID NO:419, or the CDRH1 sequence of SEQ ID NO:419 comprising 1 amino acid substitution; [1223] f) SEQ ID NO:435, or the CDRH1 sequence of SEQ ID NO:435 comprising 1 amino acid substitution; [1224] g) SEQ ID NO:449, or the CDRH1 sequence of SEQ ID NO:449 comprising 1 amino acid substitution; [1225] h) SEQ ID NO:463, or the CDRH1 sequence of SEQ ID NO:463 comprising 1 amino acid substitution; or [1226] i) SEQ ID NO:477, or the CDRH1 sequence of SEQ ID NO:477 comprising 1 amino acid substitution. [1227] j) SEQ ID NO:491, or the CDRH1 sequence of SEQ ID NO:491 comprising 1 amino acid substitution. Sentence 3a. The antibody or a fragment thereof according to sentence 1a or sentence 2a, wherein the V.sub.H domain comprises the CDRH2 sequence of: [1228] a) SEQ ID NO:364, or the CDRH2 sequence of SEQ ID NO:364 comprising 2 or 1 amino acid substitution(s); [1229] b) SEQ ID NO:378, or the CDRH2 sequence of SEQ ID NO:378 comprising 2 or 1 amino acid substitution(s); [1230] c) SEQ ID NO:392, or the CDRH2 sequence of SEQ ID NO:392 comprising 2 or 1 amino acid substitution(s); [1231] d) SEQ ID NO:406, or the CDRH2 sequence of SEQ ID NO:406 comprising 2 or 1 amino acid substitution(s); [1232] e) SEQ ID NO:420, or the CDRH2 sequence of SEQ ID NO:420 comprising 2 or 1 amino acid substitution(s); [1233] f) SEQ ID NO:436, or the CDRH2 sequence of SEQ ID NO:436 comprising 2 or 1 amino acid substitution(s); [1234] g) SEQ ID NO:450, or the CDRH2 sequence of SEQ ID NO:450 comprising 2 or 1 amino acid substitution(s); [1235] h) SEQ ID NO:464, or the CDRH2 sequence of SEQ ID NO:464 comprising 2 or 1 amino acid substitution(s); [1236] i) SEQ ID NO:478, or the CDRH2 sequence of SEQ ID NO:478 comprising 2 or 1 amino acid substitution(s); or [1237] j) SEQ ID NO:492, or the CDRH2 sequence of SEQ ID NO:492 comprising 2 or 1 amino acid substitution(s). Sentence 4a. The antibody or a fragment thereof according to any preceding sentence, wherein the V.sub.H domain comprises: [1238] a) an amino acid sequence of SEQ ID NO:366, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:366; [1239] b) an amino acid sequence of SEQ ID NO:380, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:380; [1240] c) an amino acid sequence of SEQ ID NO:394, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:394; [1241] d) an amino acid sequence of SEQ ID NO:408, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:408; [1242] e) an amino acid sequence of SEQ ID NO:422, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:422; [1243] f) an amino acid sequence of SEQ ID NO:438, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:438; [1244] g) an amino acid sequence of SEQ ID NO:452, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:452; [1245] h) an amino acid sequence of SEQ ID NO:466, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:466; [1246] i) an amino acid sequence of SEQ ID NO:480, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:480; or [1247] j) an amino acid sequence of SEQ ID NO:494, or a heavy chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:494. Sentence 5a. The antibody or fragment according to any preceding sentence comprising first and second copies of said V.sub.H domain.
[1248] Sentence 6a. The antibody or a fragment thereof according to any preceding sentence comprising a V.sub.L domain, which comprises the CDRL1 sequence of: [1249] a) SEQ ID NO:370, or the CDRL1 sequence of SEQ ID NO:370 comprising one amino acid substitution; [1250] b) SEQ ID NO:384, or the CDRL1 sequence of SEQ ID NO:384 comprising one amino acid substitution; [1251] c) SEQ ID NO:398, or the CDRL1 sequence of SEQ ID NO:398 comprising one amino acid substitution; [1252] d) SEQ ID NO:412, or the CDRL1 sequence of SEQ ID NO:412 comprising one amino acid substitution; [1253] e) SEQ ID NO:426, or the CDRL1 sequence of SEQ ID NO:426 comprising one amino acid substitution; [1254] f) SEQ ID NO:442, or the CDRL1 sequence of SEQ ID NO:442 comprising one amino acid substitution; [1255] g) SEQ ID NO:456, or the CDRL1 sequence of SEQ ID NO:456 comprising one amino acid substitution; [1256] h) SEQ ID NO:470, or the CDRL1 sequence of SEQ ID NO:470 comprising one amino acid substitution; or [1257] i) SEQ ID NO:484, or the CDRL1 sequence of SEQ ID NO:484 comprising one amino acid substitution. [1258] j) SEQ ID NO:498, or the CDRL1 sequence of SEQ ID NO:498 comprising one amino acid substitution. Sentence 7a. The antibody or a fragment thereof according to any preceding sentence comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL2 sequence of: [1259] a) SEQ ID NO:371, or the CDRL2 sequence of SEQ ID NO:371 comprising 1 amino acid substitution; [1260] b) SEQ ID NO:385, or the CDRL2 sequence of SEQ ID NO:385 comprising 1 amino acid substitution; [1261] c) SEQ ID NO:399, or the CDRL2 sequence of SEQ ID NO:399 comprising 1 amino acid substitution; [1262] d) SEQ ID NO:413, or the CDRL2 sequence of SEQ ID NO:413 comprising 1 amino acid substitution; [1263] e) SEQ ID NO:427, or the CDRL2 sequence of SEQ ID NO:427 comprising 1 amino acid substitution; [1264] f) SEQ ID NO:443, or the CDRL2 sequence of SEQ ID NO:443 comprising 1 amino acid substitution; [1265] g) SEQ ID NO:457, or the CDRL2 sequence of SEQ ID NO:457 comprising 1 amino acid substitution; [1266] h) SEQ ID NO:471, or the CDRL2 sequence of SEQ ID NO:471 comprising 1 amino acid substitution; [1267] i) SEQ ID NO:485, or the CDRL2 sequence of SEQ ID NO:485 comprising 1 amino acid substitution; or [1268] j) SEQ ID NO:499, or the CDRL2 sequence of SEQ ID NO:499 comprising 1 amino acid substitution. Sentence 8a. The antibody or a fragment thereof according to any preceding sentence comprising a or said V.sub.L domain, which V.sub.L domain comprises the CDRL3 sequence of: [1269] a) SEQ ID NO:372, or the CDRL3 sequence of SEQ ID NO:372 comprising 2 or 1 amino acid substitution(s); [1270] b) SEQ ID NO:386, or the CDRL3 sequence of SEQ ID NO:386 comprising 2 or 1 amino acid substitution(s); [1271] c) SEQ ID NO:400, or the CDRL3 sequence of SEQ ID NO:400 comprising 2 or 1 amino acid substitution(s); [1272] d) SEQ ID NO:414, or the CDRL3 sequence of SEQ ID NO:414 comprising 2 or 1 amino acid substitution(s); [1273] e) SEQ ID NO:428, or the CDRL3 sequence of SEQ ID NO:428 comprising 2 or 1 amino acid substitution(s); [1274] t) SEQ ID NO:444, or the CDRL3 sequence of SEQ ID NO:444 comprising 2 or 1 amino acid substitution(s); [1275] g) SEQ ID NO:458, or the CDRL3 sequence of SEQ ID NO:458 comprising 2 or 1 amino acid substitution(s); [1276] h) SEQ ID NO:472, or the CDRL3 sequence of SEQ ID NO:472 comprising 2 or 1 amino acid substitution(s); [1277] i) SEQ ID NO:486, or the CDRL3 sequence of SEQ ID NO:486 comprising 2 or 1 amino acid substitution(s); or [1278] j) SEQ ID NO:500, or the CDRL3 sequence of SEQ ID NO:500 comprising 2 or 1 amino acid substitution(s). Sentence 9a. The antibody or a fragment thereof according to any preceding sentence, comprising a or said V.sub.L domain, wherein the V.sub.L domain comprises an amino acid sequence of: [1279] a) SEQ ID NO:373, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:373; [1280] b) SEQ ID NO:387, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:387; [1281] c) SEQ ID NO:401, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:401; [1282] d) SEQ ID NO:415, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:415; [1283] e) SEQ ID NO:429, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:429; [1284] f) SEQ ID NO:445, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:445; [1285] g) SEQ ID NO:459, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:459; [1286] h) SEQ ID NO:473, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:473; [1287] i) SEQ ID NO:487, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:487; or [1288] j) SEQ ID NO:501, or a light chain variable domain amino acid sequence that is at least 98% identical to SEQ ID NO:501. Sentence 10a. The antibody or fragment according to any one of sentences 6a to 9a, comprising first and second copies of the a or said V.sub.L domain. Sentence 11. The antibody or fragment according to any preceding sentence, wherein the amino acid substitutions are conservative amino acid substitutions, optionally wherein the conservative substitutions are from one of six groups (each group containing amino acids that are conservative substitutions for one another) selected from:
[1289] 1) Alanine (A), Serine (S), Threonine (T);
[1290] 2) Aspartic acid (D), Glutamic acid (E);
[1291] 3) Asparagine (N), Glutamine (Q);
[1292] 4) Arginine (R), Lysine (K);
[1293] 5) Isoleucine (I), Leucine (L), Methionine (M), Valine (V); and
[1294] 6) Phenylalanine (F), Tyrosine (Y), Tryptophan (W).
Sentence 12a. An antibody or fragment thereof which specifically binds to an epitope that is:
[1295] a) Identical to an epitope to which the antibody STIM001 specifically binds;
[1296] b) Identical to an epitope to which the antibody STIM002 specifically binds;
[1297] c) Identical to an epitope to which the antibody STIM002-B specifically binds;
[1298] d) Identical to an epitope to which the antibody STIM003 specifically binds;
[1299] e) Identical to an epitope to which the antibody STIM004 specifically binds;
[1300] f) Identical to an epitope to which the antibody STIM005 specifically binds;
[1301] g) Identical to an epitope to which the antibody STIM006 specifically binds;
[1302] h) Identical to an epitope to which the antibody STIM007 specifically binds;
[1303] i) Identical to an epitope to which the antibody STIM008 specifically binds; or [1304] j) Identical to an epitope to which the antibody STIM009 specifically binds. Sentence 13a. The antibody or fragment according to sentence 12a, wherein the epitope is identified by unrelated amino acid scanning, or by X-ray crystallography. Sentence 14a. The antibody or fragment according to sentence 13a, wherein the contact residues of the epitope are defined by a reduction in affinity of at least 10-fold in an unrelated amino acid scan, e.g. an alanine scan as determined by SPR. Sentence 15a. An antibody or fragment thereof which:
[1305] a) Competes for binding to hICOS with the antibody STIM001;
[1306] b) Competes for binding to hICOS with the antibody STIM002;
[1307] c) Competes for binding to hICOS with the antibody STIM002-B;
[1308] d) Competes for binding to hICOS with the antibody STIM003;
[1309] e) Competes for binding to hICOS with the antibody STIM004;
[1310] f) Competes for binding to hICOS with the antibody STIM005;
[1311] g) Competes for binding to hICOS with the antibody STIM006;
[1312] h) Competes for binding to hICOS with the antibody STIM007;
[1313] i) Competes for binding to hICOS with the antibody STIM008; or [1314] j) Competes for binding to hICOS with the antibody STIM009. Sentence 16a. The antibody or fragment according to any preceding sentence which specifically binds to cynomolgus ICOS (Seq ID No:513, Seq ID NO: 513 or Seq ID No: 514) and/or mouse ICOS (Seq ID No:510, Seq ID No:511 or Seq ID No:512). Sentence 17a. The antibody or fragment according to any preceding sentence which specifically binds to a hICOS isoform or natural variant, a mouse ICOS isoform or natural variant and/or a cynomolgus ICOS isoform or natural variant. Sentence 18a. The antibody or fragment according to sentence 17a, wherein the hICOS isoform comprises an amino acid sequence as defined by Seq ID no:509. Sentence 19a. The antibody or fragment according to any preceding sentence, wherein the antibody or fragment comprises a constant region, such as a human constant region, for example an effector-null human constant region, e.g. an IgG4 constant region or an IgG1 constant region, optionally wherein the constant region is IgG4-PE (Seq ID No:199), or a disabled IgG1 (Seq ID No:205). Sentence 20a. The antibody or fragment according to sentence 19a, wherein the constant region is a murine constant region. Sentence 21a. The antibody or fragment according to sentence 19a or sentence 20a, wherein the constant region has CDC and/or ADCC activity.
5. Anti-ICOS Bispecific Antibodies
[1315] As previously described, the PD-L1 antibodies as provided herein, may be formatted as a multispecific (e.g. bispecific) antibody, as disclosed hereinabove in concepts 37 to 40. In one embodiment disclosed therein, the PD-L1 antibodies as disclosed herein may be formatted in a bispecific antibody which has specificity for both PD-L1 (e.g. human PD-L1) and for ICOS (e.g. an agonist to ICOS, such as human ICOS).
[1316] Thus, there is provided a multispecific (e.g. bispecific antibody or a dual-binding antibody) which has specificity for PD-L1 (e.g. human PD-L1) and ICOS (e.g. human ICOS). In one embodiment the multispecific (e.g. bispecific or dual-binding) antibody has agonistic activity against ICOS (e.g. human ICOS).
[1317] Various ICOS-containing mutispecific antibodies are described in the arrangements below:
Arrangement 1. A multispecific antibody (e.g. bispecific antibody or a dual-binding antibody) which binds (and optionally has specificity for) ICOS (e.g. human ICOS) and another target antigen.
[1318] In one embodiment, there is provided a bispecific antibody or a dual-binding antibody which binds ICOS (e.g. human ICOS) and another target antigen. In one embodiment, there is provided a bispecific antibody or a dual-binding antibody which has specificity for ICOS (e.g. human ICOS) and another target antigen. In one embodiment, there is provided a bispecific antibody antibody which binds ICOS (e.g. human ICOS) and another target antigen, and wherein the bispecific antibody format is a mAb.sup.2. In one embodiment, there is provided a bispecific antibody antibody which binds ICOS (e.g. human ICOS) and another target antigen, and wherein the bispecific antibody format is a mAb.sup.2, and the binding to another target antigen is provided by a modified constant region (i.e. an Fcab). In one embodiment, there is provided a bispecific antibody antibody which binds ICOS (e.g. human ICOS) and another target antigen which is PD-L1 (e.g. human PD-L1), and wherein the bispecific antibody format is a mAb2, and the binding to ICOS is provided by a modified constant region (i.e. an Fcab). In one embodiment, there is provided a bispecific antibody antibody which binds ICOS (e.g. human ICOS) and another target antigen which is PD-L1 (e.g. human PD-L1), and wherein the bispecific antibody format is a mAb2, and the binding to ICOS is provided by a modified constant region (i.e. an Fcab) and the binding to PD-L1 is provided by any of the antibodies described in concepts 1 to 70, or by any of the PD-L1 antibodies described in arrangement 5 or 5a below. In one embodiment, there is provided a bispecific antibody antibody which binds ICOS (e.g. human ICOS) and another target antigen which is PD-L1 (e.g. human PD-L1), and wherein the bispecific antibody format is a mAb.sup.2, and the binding to PD-L1 is provided by a modified constant region (i.e. an Fcab). In one embodiment, there is provided a bispecific antibody antibody which binds ICOS (e.g. human ICOS) and another target antigen which is PD-L1 (e.g. human PD-L1), and wherein the bispecific antibody format is a mAb.sup.2, and the binding to PD-L1 is provided by a modified constant region (i.e. an Fcab) and the binding to ICOS is provided by any of the antibodies described in sentences 1 to 102 or sentences 1a to 21a.
[1319] In one embodiment, the multispecific (e.g. bispecific or dual-binding) antibody has agonistic activity against ICOS (e.g. human ICOS). The another target antigen may be any of the target antigens specified in concept 39. In one embodiment, the another target antigen is an immune checkpoint inhibitor, such as PD-1, PD-L1, CTLA-4, TIGIT, TIM-3, LAG-3 and VISTA, e.g. PD-L1, TIGIT, CTLA-4, TIM-3 and LAG-3. In one embodiment, the another target antigen is an immune modulator, such as BTLA, hHVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10 and CD155, e.g. GARP, SIRP.alpha., CXCR4, BTLA, hVEM and CSF1R. In one embodiment, the another target antigen is an immune activator, such as CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies), CD27 and CD3, or CD137, GITR, OX40, CD40, CXCR3 (e.g. agonistic anti-CXCR3 antibodies) and CD3, for example CD137, GITR and OX40). In one embodiment, the another target antigen is PD-L1. In one embodiment, the another target antigen is CTLA-4. In one embodiment, the another target antigen is TIGIT. In one embodiment, the another target antigen is TIM-3. In one embodiment, the another target antigen is LAG-3. In one embodiment, the another target antigen is GITR. In one embodiment, the another target antigen is VISTA. In one embodiment, the another target antigen is CD137. In one embodiment, the another target antigen is SIRP.alpha.. In one embodiment, the another target antigen is CXCL10. In one embodiment, the another target antigen is CD155. In one embodiment, the another target antigen is CD40. The antibodies against these another target antigens may be any of those described in aspect 1a hereinabove.
[1320] The format of the multispecific, bispecific or dual-binding antibody may be any of the formats disclosed herein, for example as set out in concepts 37 to 40. In particular, the binding and/or specificity for ICOS may be provided by a non-immunoglobulin format, for example, a T-cell receptor binding domain; an immunoglobulin superfamily domain; an agnathan variable lymphocyte receptor; a fibronectin domain (e.g., an Adnectin.TM.); an antibody constant domain (e.g., a CH3 domain, e.g., a CH2 and/or CH3 of an Fcab.TM.) wherein the constant domain is not a functional CH1 domain; an scFv; an (scFv)2; an sc-diabody; an scFab; a centyrin and an epitope binding domain derived from a scaffold selected from CTLA-4 (Evibody.TM.); a lipocalin domain; Protein A such as Z-domain of Protein A (e.g., an Affibody.TM. or SpA); an A-domain (e.g., an Avimer.TM. or Maxibody.TM.); a heat shock protein (such as and epitope binding domain derived from GroEI and GroES); a transferrin domain (e.g., a trans-body); ankyrin repeat protein (e.g., a DARPin.TM.); peptide aptamer; C-type lectin domain (e.g., Tetranectin.TM.); human .gamma.-crystallin or human ubiquitin (an affilin); a PDZ domain; scorpion toxin; and a kunitz type domain of a human protease inhibitor. The binding and/or specificity for another target antigen may be provided by an immunoglobulin-derived antigen-binding protein.
[1321] "Specifically binds" has the meaning provided hereinabove. Binding constants, e.g. K.sub.D may be determined as described elsewhere herein, and particular K.sub.Ds of interest are described in arrangement 2 below, and in concept 1 hereinabove (although specified for PD-L1 binding, the values of K.sub.D may be equally applied to anti-ICOS binding).
Arrangement 2. A multispecific antibody according to arrangement 1, wherein the ICOS is human ICOS.
[1322] Sequences of human ICOS are provided in Seq ID Nos:506, 507 and 508. In one embodiment, the multispecific antibody is specific for wild type human ICOS. In another embodiment, the multispecific antibody is cross-reactive to an isoform or natural variant of hICOS, for example the isoform of Seq ID No:509. Other isoforms and natural variants are well known to those skilled in the art. In another embodiment, the multispecific antibody is specific for the isoform or natural variant (e.g. the ICOS isoform having the amino acid sequence of Seq ID No:509) over wild type hICOS.
[1323] One way to quantify the extent of species cross-reactivity of an antibody, e.g. a multispecific, bispecific or dual-binding antibody is as the fold-difference in its affinity for antigen compared with a different antigen (e.g. fold difference in affinity for human ICOS vs mouse ICOS or fold difference in affinity for wild-type hICOS vs an isoform of hICOS). Affinity may be quantified as K.sub.D, referring to the equilibrium dissociation constant of the antibody-antigen reaction as determined by SPR (optionally with the antibody in Fab format as described elsewhere herein). A species or isoform cross-reactive anti-ICOS antibody may have a fold-difference in affinity for binding human and mouse ICOS that is 30-fold or less, 25-fold or less, 20-fold or less, 15-fold or less, 10-fold or less or 5-fold or less. To put it another way, the K.sub.D of binding the extracellular domain of hICOS may be within 30-fold, 25-fold, 20-fold, 15-fold, 10-fold or 5-fold of the K.sub.D of binding the extracellular domain of mouse ICOS. Antibodies can also be considered cross-reactive if the K.sub.D for binding antigen of both species meets a threshold value, e.g., if the K.sub.D of binding hICOS and the K.sub.D of binding mouse ICOS are both 10 mM or less, preferably 5 mM or less, more preferably 1 mM or less. The K.sub.D may be 10 nM or less, 5 nM or less, 2 nM or less, or 1 nM or less. The K.sub.D may be 0.9 nM or less, 0.8 nM or less, 0.7 nM or less, 0.6 nM or less, 0.5 nM or less, 0.4 nM or less, 0.3 nM or less, 0.2 nM or less, or 0.1 nM or less.
[1324] An alternative measure of cross-reactivity for binding hICOS and mouse ICOS, or WT hICOS and an isoform of hICOS is the ability of an antibody to neutralise ICOS ligand binding to ICOS receptor, such as in an HTRF assay (as described elsewhere herein). Examples of species cross-reactive antibodies are provided herein, including STIM001, STIM002, STIM002-B, STIM003, STIM005 and STIM006, each of which was confirmed as neutralising binding of human B7-H2 (ICOS ligand) to hICOS and neutralising binding of mouse B7-H2 to mouse ICOS in an HTRF assay. Any of these antibodies or their variants may be selected when an antibody cross-reactive for human and mouse ICOS is desired. A species cross-reactive anti-ICOS antibody may have an IC.sub.50 for inhibiting binding of hICOS to human ICOS receptor that is within 25-fold, 20-fold, 15-fold, 10-fold or 5-fold of the IC.sub.50 for inhibiting mouse ICOS to mouse ICOS receptor as determined in an HTRF assay. Antibodies can also be considered cross-reactive if the IC.sub.50 for inhibiting binding of hICOS to human ICOS receptor and the IC.sub.50 for inhibiting binding of mouse ICOS to mouse ICOS receptor are both 1 mM or less, preferably 0.5 mM or less, e.g., 30 nM or less, 20 nM or less, 10 nM or less. The IC.sub.50s may be 5 nM or less, 4 nM or less, 3 nM or less or 2 nM or less. In some cases, the IC.sub.50s will be at least 0.1 nM, at least 0.5 nM or at least 1 nM.
[1325] Affinities may also be as disclosed in concept 27 hereinabove.
Arrangement 3. A multispecific antibody according to arrangement 2, which comprises a V.sub.H domain comprising a CDRH1, a CDRH2 and a CDRH3 which V.sub.H domain binds (and optionally has specificity for) hICOS.
[1326] In one embodiment, the multispecific antibody comprises at least one V.sub.H domain which binds to hICOS. For example, the multispecific antibody may comprise a single-chain Fv (scFv), single-chain antibody, a single domain antibody or a domain antibody compositing only the V.sub.H region which binds to (and optionally has specificity for) hICOS.
Arrangement 4. A multispecific antibody according to arrangement 2 or arrangement 3, which comprises a V.sub.L domain comprising a CDRL1, a CDRL2 and a CDRL3, which V.sub.L domain binds (and optionally has specificity for) hICOS.
[1327] In one embodiment, the multispecific antibody comprises at least one V.sub.L domain which binds to hICOS. For example, the multispecific antibody may comprise a single-chain Fv (scFv), single-chain antibody, a single domain antibody or a domain antibody compositing only the V.sub.L region which binds to (and optionally has specificity for) hICOS.
[1328] In another embodiment, the multispecific antibody comprises a paired V.sub.H and V.sub.L domain, including, but not limited to, an intact or full-length antibody, a Fab fragment, a Fab' fragment, a F(ab').sub.2 fragment or a Fv fragment.
[1329] Arrangement 5. A multispecific antibody according to arrangement 3 or 4, wherein the V.sub.H and/or V.sub.L domain is any of V.sub.H and/or V.sub.L domains: [1330] a. of the antibody 7F12, 37A10, 35A9, 36E10, 16G10, 37A10S713, 37A10S714, 37A10S715, 37A10S716, 37A10S717, 37A10S718, 16G10S71, 16G10S72, 16G10S73, 16G10S83, 35A9S79, 35A9S710, 35A9S89 or any other antibody described in WO2016/154177 and US2016/0304610; [1331] b. of the antibody 422.2, H2L5, or any other antibody described in WO2016/120789 and US2016/0215059; [1332] c. of the antibody 314-8, the antibody produced from hybridoma CNCM 1-4180, or any other antibody described in WO2014/033327 and US2015/0239978; [1333] d. of the antibody Icos145-1, the antibody produced by hybridoma CNCM 1-4179, or any other antibody described in WO2012/131004, U.S. Pat. No. 9,376,493 and US2016/0264666; [1334] e. of the antibody JMAb 136, "136", or any other antibody described in WO2010/056804; [1335] f. of the antibody MIC-944, 9F3 or any other antibody described in WO99/15553, U.S. Pat. Nos. 7,259,247, 7,132,099, 7,125,551, 7,306,800, 7,722,872, WO05/103086, US8.318.905 and U.S. Pat. No. 8,916,155; [1336] g. of any JMAb antibody, e.g., any of JMAb-124, JMAb-126, JMAb-127, JMAb-128, JMAb-135, JMAb-136, JMAb-137, JMAb-138, JMAb-139, JMAb-140, JMAb-141, e.g., JMAb136, or any other antibody described in WO98/3821, U.S. Pat. No. 7,932,358B2, US2002/156242, U.S. Pat. Nos. 7,030,225, 7,045,615, 7,279,560, 7,226,909, 7,196,175, 7,932,358, 8,389,690, WO02/070010, U.S. Pat. Nos. 7,438,905, 7,438,905, WO01/87981, U.S. Pat. Nos. 6,803,039, 7,166,283, 7,988,965, WO01/15732, U.S. Pat. Nos. 7,465,445 and 7,998,478; [1337] h. of the antibody 17G9 or any other antibody described in WO2014/08911; [1338] i. of any antibody described in WO2012/174338; [1339] j. of any antibody described in US2016/0145344; [1340] k. of any antibody described in WO2011/020024, US2016/002336, US2016/024211 and U.S. Pat. No. 8,840,889; [1341] l. of any antibody described in U.S. Pat. No. 8,497,244; [1342] m. of the antibody known as GSK3359609; [1343] n. of the antibody known as JTX-2011; or [1344] o. of antibody clone ISA-3 (eBioscience), clone SP98 (Novus Biologicals), clone 1 G1, clone 3G4 (Abnova Corporation), clone 669222 (R&D Systems), clone TQ09 (Creative Diagnostics), or clone C398.4A (BioLegend). Arrangement 5a. A multispecific antibody according to any preceding arrangement, which comprises the CDRH1, CDRH2, CDR3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L region: [1345] a. of the antibody 7F12, 37A10, 35A9, 36E10, 16G10, 37A10S713, 37A10S714, 37A10S715, 37A10S716, 37A10S717, 37A10S718, 16G10S71, 16G10S72, 16G10S73, 16G10S83, 35A9S79, 35A9S710, 35A9S89 or any other antibody described in WO2016/154177 and US2016/0304610; [1346] b. of the antibody 422.2, H2L5, or any other antibody described in WO2016/120789 and US2016/0215059; [1347] c. of the antibody 314-8, the antibody produced from hybridoma CNCM 1-4180, or any other antibody described in WO2014/033327 and US2015/0239978; [1348] d. of the antibody Icos145-1, the antibody produced by hybridoma CNCM 1-4179, or any other antibody described in WO2012/131004, U.S. Pat. No. 9,376,493 and US2016/0264666; [1349] e. of the antibody JMAb 136, "136", or any other antibody described in WO2010/056804; [1350] f. of the antibody MIC-944, 9F3 or any other antibody described in WO99/15553, U.S. Pat. Nos. 7,259,247, 7,132,099, 7,125,551, 7,306,800, 7,722,872, WO05/103086, US8.318.905 and U.S. Pat. No. 8,916,155; [1351] g. of any JMAb antibody, e.g., any of JMAb-124, JMAb-126, JMAb-127, JMAb-128, JMAb-135, JMAb-136, JMAb-137, JMAb-138, JMAb-139, JMAb-140, JMAb-141, e.g., JMAb136, or any other antibody described in WO98/3821, U.S. Pat. No. 7,932,358B2, US2002/156242, U.S. Pat. Nos. 7,030,225, 7,045,615, 7,279,560, 7,226,909, 7,196,175, 7,932,358, 8,389,690, WO02/070010, U.S. Pat. Nos. 7,438,905, 7,438,905, WO01/87981, U.S. Pat. Nos. 6,803,039, 7,166,283, 7,988,965, WO01/15732, U.S. Pat. Nos. 7,465,445 and 7,998,478; [1352] h. of the antibody 17G9 or any other antibody described in WO2014/08911; [1353] i. of any antibody described in WO2012/174338; [1354] j. of any antibody described in US2016/0145344; [1355] k. of any antibody described in WO2011/020024, US2016/002336, US2016/024211 and U.S. Pat. No. 8,840,889; [1356] I. of any antibody described in U.S. Pat. No. 8,497,244; [1357] m. of the antibody known as GSK3359609; [1358] n. of the antibody known as JTX-2011; or [1359] o. of antibody clone ISA-3 (eBioscience), clone SP98 (Novus Biologicals), clone 1 G1, clone 3G4 (Abnova Corporation), clone 669222 (R&D Systems), clone TQ09 (Creative Diagnostics), or clone C398.4A (BioLegend). Arrangement 6. A multispecific antibody according to arrangement 3 or 4, wherein the V.sub.H and/or V.sub.L domain is any of V.sub.H and/or V.sub.L domains defined in sentences 1 to 102 or sentences 1a to 21a.
[1360] In one embodiment, the anti-ICOS V.sub.H and/or V.sub.L is as described in GB patent application 1620414.1 (filed 1 Dec. 2016), the contents of which are incorporated herein by reference.
Arrangement 7. A multispecific antibody according to any preceding arrangement, which has agonistic activity against ICOS.
[1361] Agonism can be tested for in an in vitro T-cell activation assays, using antibody in soluble form (e.g. in immunoglobulin format or other antibody format comprising two spatially separated antigen-binding sites, e.g., two V.sub.H-V.sub.L pairs), either including or excluding a cross-linking agent, or using antibody (e.g. multispecific antibody) bound to a solid surface to provide a tethered array of antigen-binding sites. Agonism assays may use a hICOS positive T-lymphocyte cell line such as MJ cells (ATCC CRL-8294) as the target T-cell for activation in such assays. One or more measures of T-cell activation can be determined for a test antibody and compared with a reference molecule or a negative control to determine whether there is a statistically significant (p<0.05) difference in T-cell activation effected by the test antibody (e.g. multispecific antibody) compared with the reference molecule or the control. One suitable measure of T-cell activation is production of cytokines, e.g., IFN.gamma., TNF.alpha. or IL-2. A skilled person will include suitable controls as appropriate, standardising assay conditions between test antibody and control. A suitable negative control is an antibody in the same format (e.g., isotype control) that does not bind ICOS, e.g., an antibody (e.g. multispecific antibody) specific for an antigen that is not present in the assay system. A significant difference is observed for test antibody relative to a cognate isotype control within the dynamic range of the assay is indicative that the antibody acts as an agonist of the ICOS receptor in that assay.
[1362] An agonist antibody may be defined as one which, when tested in a T-cell activation assay:
[1363] has a significantly lower EC.sub.50 for induction of IFN.gamma. production compared with control antibody;
[1364] induces significantly higher maximal IFN.gamma. production compared with control antibody;
[1365] has a significantly lower EC.sub.50 for induction of IFN.gamma. production compared with ICOSL-Fc;
[1366] induces significantly higher maximal IFN.gamma. production compared with ICOSL-Fc;
[1367] has a significantly lower EC.sub.50 for induction of IFN.gamma. production compared with reference antibody C398.4A; and/or
[1368] induces significantly higher maximal IFN.gamma. production compared with reference antibody C398.4A.
[1369] A significantly lower or significantly higher value may for example be up to 0.5-fold different, up to 0.75-fold different, up to 2-fold different, up to 3-fold different, up to 4-fold different or up to 5-fold different, compared with the reference or control value.
[1370] Thus, in one example, an antibody (e.g. a multispecific antibody) provided herein has a significantly lower, e.g., at least 2-fold lower, EC.sub.50 for induction of IFN.gamma. in an MJ cell activation assay using the antibody in bead-bound format, compared with control.
[1371] The bead-bound assay uses the antibody (e.g. multispecific antibody) (and, for control or reference experiments, the control antibody, reference antibody or ICOSL-Fc) bound to the surface of beads. Magnetic beads may be used, and various kinds are commercially available, e.g., Tosyl-activated DYNABEADS M-450 (DYNAL Inc, 5 Delaware Drive, Lake Success, N.Y. 11042 Prod No. 140.03, 140.04). Beads may be coated (coating methods are well-known to those skilled in the art), or generally by dissolving the coating material in carbonate buffer (pH 9.6, 0.2 M) or other method known in the art. Use of beads conveniently allows the quantity of protein bound to the bead surface to be determined with a good degree of accuracy. Standard Fc-protein quantification methods can be used for coupled protein quantification on beads. Any suitable method can be used, with reference to a relevant standard within the dynamic range of the assay. DELFIA, ELISA or other methods could be used.
[1372] Agonism activity of an antibody can also be measured in primary human T-lymphocytes ex vivo. The ability of an antibody (e.g. multispecific antibody) to induce expression of IFN.gamma. in such T-cells is indicative of ICOS agonism. Preferably, an antibody will show significant (p<0.05) induction of IFN.gamma. at 5 .mu.g/mL compared with control antibody in a T-cell activation assay. An anti-ICOS antibody may stimulate T-cell activation to a greater degree than ICOS-L or C398.4 in such an assay. Thus, the antibody may show significantly (p<0.05) greater induction of IFN.gamma. at 5 .mu.g/mL compared with the control or reference antibody in a T-cell activation assay. TNF.alpha. or IL-2 induction may be measured as an alternative assay readout.
[1373] Agonism of an anti-ICOS antibody may contribute to its ability to change the balance between populations of T.sub.Reg and T.sub.Eff cells in vivo, e.g., in a site of pathology such as a tumour microenvironment, in favour of T cells. The ability of an antibody to enhance tumour cell killing by activated ICOS-positive effector T-cells may be determined, as discussed elsewhere herein.
Arrangement 8. A multispecific antibody according to any preceding arrangement, which binds (and optionally has specificity for) mouse ICOS and/or cynomolgus ICOS.
[1374] The multispecific antibodies described herein may be cross-reactive, and may for example bind the extracellular domain of mouse ICOS as well as human ICOS. The multispecific antibodies may bind other non-human ICOS, including ICOS of primates, such as cynomolgus monkey. An anti-ICOS multispecific antibody intended for therapeutic use in humans must bind human ICOS, whereas binding to ICOS of other species would not have direct therapeutic relevance in the human clinical context. Regardless of the underlying theory, however, cross-reactive antibodies are of high value and are excellent candidates as therapeutic molecules for pre-clinical and clinical studies. Cross-reactivity may be determined as set out for arrangement 2 hereinabove.
Arrangement 9. A multispecific antibody according to any preceding arrangement which is a bispecific antibody.
[1375] A bispecific antibody has any of the meanings set out hereinabove.
Arrangement 10. A bispecifc antibody according to arrangement 9, wherein the bispecific antibody format is selected from DVD-Ig, mAb.sup.2, FIT-Ig, mAb-dAb, dock and lock, SEEDbody, scDiabody-Fc, diabody-Fc, tandem scFv-Fc, Fab-scFv-Fc, Fab-scFv, intrabody, BiTE, diabody, DART, TandAb, scDiabody, scDiabody-CH3, Diabody-CH3, minibody, knobs-in-holes, knobs-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs, charge pairs, charge pairs with common light chain, in particular mAb.sup.2, knob-in-holes, knob-in-holes with common light chain, knobs-in-holes with common light chain and charge pairs and FIT-Ig, e.g. mAb.sup.2 and FIT-Ig.
[1376] In one embodiment, the bispecific antibody format is as described in any of concepts 37 to 40 described hereinabove, or as described in the definitions section. In one embodiment, the bispecific antibody format is a mAb.sup.2, wherein the ICOS binding is provided by the Fcab portion of the bispecific antibody. In another embodiment, the the bispecific antibody format is a mAb.sup.2, wherein the ICOS binding is provided by the Fab portion of the bispecific antibody.
[1377] In another embodiment, the bispecific antibody is not a mAb.sup.2 bispecific antibody.
Arrangement 11. A multispecific antibody according to any one of arrangements 1 to 8 which is a dual binding antibody.
[1378] A dual-binding antibody has any of the meanings set out hereinabove.
Arrangement 12. A multispecific, bispecific or dual binding antibody according to any one of arrangements 1 to 11, wherein the another target antigen is selected from immune checkpoint inhibitors, immune modulators and immune activators. Arrangement 13. A multispecific, bispecific or dual-binding antibody according to arrangement 12, wherein the another target antigen is selected from PD-1, PD-L1, CTLA-4, TIGIT, TIM-3, LAG-3, VISTA, BTLA, HVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10, CD155, CD137, GITR, OX40, CXCR3, CD27 and CD3. Arrangement 13a. A multispecific, bispecific or dual-binding antibody according to arrangement 12, wherein the another target antigen is selected from PD-1, PD-L1, CTLA-4, TIGIT, TIM-3, LAG-3, VISTA, BTLA, HVEM, CSF1R, CCR4, CD39, CD40, CD73, CD96, CXCR2, CXCR4, CD200, GARP, SIRP.alpha., CXCL9, CXCL10, CD155, CD137, GITR, OX40, CXCR3 and CD3.
[1379] In one embodiment, the antigen-binding site which binds the another target antigen is provided for by any of the CDRH1, CDRH2, CDR3, CDRL1, CDRL2 and CDRL3, or the V.sub.H, or the V.sub.L or the V.sub.H and V.sub.L regions from any one of the antibodies against the targets listed in arrangement 13 which are described in more detail in aspect 1a hereinabove.
Arrangement 14. A multispecific, bispecific or dual-binding antibody according to arrangement 13, wherein the another target antigen is selected from PD-L1, TIGIT, TIM-3, LAG-3, GARP, SIRP.alpha., CXCR4, BTLA, HVEM, CSF1R, agonistic anti-CXCR3 antibodies), CD137, GITR and OX40. Arrangement 15. A multispecific, bispecific or dual-binding antibody according to arrangement 14, wherein the another target antigen is PD-L1 (e.g. human PD-L1). Arrangement 16. A multispecific, bispecific or dual-binding antibody according to arrangement 15, wherein the binding (and optionally specificity for) PD-L1 is provided by any of the antibodies or fragments as defined in concepts 1 to 70. Arrangement 17. A multispecific, bispecific or dual-binding antibody according to arrangement 15 or arrangement 16, which comprises a V.sub.H domain comprising a CDRH1, a CDRH2 and a CDRH3 which V.sub.H domain has specificity for human PD-L1. Arrangement 18. A multispecific, bispecific or dual-binding antibody according to any one of arrangements 15 to 17, which comprises a V.sub.L domain comprising a CDRL1, a CDRL2 and a CDRL3, which V.sub.L domain as specificity for human PD-L1. Arrangement 19. A multispecific, bispecific or dual-binding antibody according to arrangement 17 or arrangement 18, wherein the V.sub.H and/or V.sub.L domain is any of V.sub.H and/or V.sub.L domains from atezolizumab (Roche), avelumab (Merck), BMS-936559 (BMS), durvalumab (Medimmune) or from any of the PD-L1 antibodies disclosed in WO2016/061142, WO2016/022630, WO2016/007235, WO2015/173267, WO2015/181342, WO2015/109124, WO2015/112805, WO2015/061668, WO2014/159562, WO2014/165082, WO2014/100079, WO2014/055897, WO2013/181634, WO2013/173223, WO2013/079174, WO2012/145493, WO2011/066389, WO2010/077634, WO2010/036959 or WO2007/005874. Arrangement 20. A multispecific, bispecific or dual-binding antibody according to arrangement 17 or arrangement 18, wherein the V.sub.H and/or V.sub.L domain is any of V.sub.H and/or V.sub.L domains described in concepts 1 to 70. Arrangement 21. A multispecific, bispecific or dual-binding antibody according to any one of arrangements 15 to 20, which binds (and optionally has specificity for) mouse PD-L1 and/or cynomolgus PD-L1.
[1380] Cross reactivity may be as described hereinabove for arrangement 2 or concept 27.
Arrangement 22. A composition comprising a multispecific, bispecific or dual-binding antibody as defined in any preceding arrangement and a pharmaceutically acceptable excipient, diluent or carrier and optionally further comprising a further therapeutic agent independently selected from the group consisting of:
[1381] a) other immune checkpoint inhibitors (such as anti-TIM-3 antibodies, anti-PD-1 antibodies, anti-CRA-4 antibodies, anti-TIGIT antibodies and anti-LAG-3 antibodies);
[1382] b) immune stimulators (such as anti-OX40 antibodies, anti-GITR antibodies, anti-CD137 antibodies, anti-ICOS antibodies and anti-CD40 antibodies);
[1383] c) chemokine receptor antagonists (such as CXCR4, CCR4 and CXCR2);
[1384] d) targeted kinase inhibitors (such as CSF-1R or VEGFR inhibitors);
[1385] e) angiogenesis inhibitors (such as anti-VEGF-A or Delta-like Ligand-4);
[1386] f) immune stimulating peptides or chemokines (such as CXCL9 or CXCL10);
[1387] g) cytokines (such as IL-15 and IL-21);
[1388] h) bispecific T-cell engagers (BiTEs) having at least one specificity against CD3 (e.g. CD3/CD19 BiTE);
[1389] i) other bi-specific molecules (for example IL-15-containing molecules targeted towards tumour associated antigens, for example Epidermal growth factor receptors such as EGFR, Her-2, New York Esophageal Cancer-1 (NY-ESO-1), GD2, EpCAM or Melanoma. Associated Antigen-3 (MAGE-A3));
[1390] j) oncolytic viruses (such as HSV virus (optionally which secretes GMCSF), Newcastle disease virus and Vaccinia virus);
[1391] k) vaccination with tumour associated antigens (such as New York Esophageal Cancer-1 [NY-ESO-1], Melanoma Associated Antigen-3 [MAGE-3]);
[1392] l) cell-based therapies (such as chimeric Antigen Receptor-T-cells (CAR-T) for example expressing anti-CD19, anti-EpCam or anti-mesothelin);
[1393] m) bi-specific NK cell engagers having a specificity against an activating MK receptor such as NKG2D or CD16a; and
[1394] n) adoptive transfer of tumour specific T-cells or LAK cells.
[1395] The antibodies may be any of the sequences or antibodies described in arrangement 5 or detailed in aspect 1a. Other features of this arrangement may be as described in concept 49.
[1396] Arrangement 22a. A pharmaceutical composition according to arrangement 22, or a kit comprising a pharmaceutical composition as defined in arrangement 22, wherein the composition is for treating and/or preventing a condition or disease selected from neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease, diffuse large B-cell lymphoma.
[1397] Arrangement 22b. A pharmaceutical composition according to arrangement 22 or arrangement 22a in combination with, or kit according to arrangement 22a comprising, a label or instructions for use to treat and/or prevent said disease or condition in a human; optionally wherein the label or instructions comprise a marketing authorisation number (e.g., an FDA or EMA authorisation number); optionally wherein the kit comprises an IV or injection device that comprises the multispecific, bispecific or dual-binding antibody.
[1398] Arrangement 23. A multispecific, bispecific or dual-binding antibody as defined in any one of arrangements. 1 to 21 for use in treating or preventing a disease or condition, selected from neurological disease, neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours; such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas).
Arrangement 24. Use of a multispecific, bispecific or dual-binding antibody as defined in any one of arrangements 1 to 21 in the manufacture of a medicament for administration to a human for treating or preventing a disease or condition in the human selected from neurological disease, neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas). Arrangement 25. A method of treating or preventing a disease or condition selected from neurological disease, neoplastic or non-neoplastic disease, chronic viral infections, and malignant tumours, such as melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma, mesothelioma, virally induced cancers (such as cervical cancer and nasopharyngeal cancer), soft tissue sarcomas, haematological malignancies such as Hodgkin's and non-Hodgkin's disease and diffuse large B-cell lymphoma (for example melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or for example virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas) in a human, comprising administering to said human a therapeutically effective amount of a multispecific, bispecific or dual-binding antibody as defined in any one of arrangements 1 to 21, wherein the disease or condition is thereby treated or prevented.
[1399] The diseases and conditions which may be treated or prevented by the multispecific, bisecific or dual-binding antibodies provided for in these arrangements may be any of the diseases provided for in, for example concepts 41 to 45, aspects 51 to 55, or in any of the sentences described herein.
Arrangement 26. The multispecific, bispecific or dual-binding antibody according to arrangement 23, the use according to arrangement 24 or the method according to arrangement 25, wherein the neurological disease is a neurodegenerative disease, disorder or condition, optionally wherein the neurodegenerative disease, disorder or condition is selected from Alzheimer's disease, amyotrophic lateral sclerosis, Parkinson's disease, Huntington's disease, primary progressive multiple sclerosis, secondary progressive multiple sclerosis, corticobasal degeneration, Rett syndrome, a retinal degeneration disorder selected from age-related macular degeneration and retinitis pigmentosa; anterior ischemic optic neuropathy, glaucoma, uveitis, depression, trauma-associated stress or post-traumatic stress disorder, frontotemporal dementia, Lewy body dementias, mild cognitive impairments, posterior cortical atrophy, primary progressive aphasia and progressive supranuclear palsy or aged-related dementia, in particular Alzheimer's disease, amyotrophic lateral sclerosis, Parkinson's disease and Huntington's disease, and e.g. Alzheimer's disease. Arrangement 27. The multispecific, bispecific or dual-binding antibody according to arrangement 23, the use according to arrangement 24 or the method according to arrangement 25, wherein the cancer is selected from melanoma, Merkel cell carcinoma, non-small cell lung cancer (squamous and non-squamous), renal cell cancer, bladder cancer, head and neck squamous cell carcinoma and mesothelioma or is selected from virally induced cancers (such as cervical cancer and nasopharyngeal cancer) and soft tissue sarcomas. Arrangement 28. The multispecific, bispecific or dual-binding antibody, the use or the method according to any one of arrangements 23 to 27, further comprising administering to the human a further therapy, for example a further therapeutic agent, optionally wherein the further therapeutic agent is independently selected from the group consisting of: [1400] a. other immune checkpoint inhibitors (such as anti-TIM-3 antibodies, anti-PD-1 antibodies, anti-CTLA-4 antibodies, anti-TIGIT antibodies and anti-LAG-3 antibodies); [1401] b. immune stimulators (such as anti-OX40 antibodies, anti-GITR antibodies, anti-CD137 antibodies and anti-CD40 antibodies); [1402] c. chemokine receptor antagonists (such as CXCR4, CCR4 and CXCR2); [1403] d. targeted kinase inhibitors (such as CSF-1R or VEGFR inhibitors); [1404] e. angiogenesis inhibitors (such as anti-VEGF-A or Delta-like Ligand-4); [1405] f. immune stimulating peptides or chemokines (such as CXCL9 or CXCL10); [1406] g. cytokines (such as IL-15 and IL-21); [1407] h. bispecific T-cell engagers (BiTEs) having at least one specificity against CD3 (e.g. CD3/CD19 BiTE); [1408] i. other bi-specific molecules (for example IL-15-containing molecules targeted towards tumour associated antigens, for example Epidermal growth factor receptors such as EGFR, Her-2, New York Esophageal Cancer-1 (NY-ESO-1), GD2, EpCAM or Melanoma Associated Antigen-3 (MAGE-A3)); [1409] j. oncolytic viruses (such as HSV virus (optionally which secretes GMCSF), Newcastle disease virus and Vaccinia virus); [1410] k. vaccination with tumour associated antigens (such as New York Esophageal Cancer-1 [NY-ESO-1], Melanoma Associated Antigen-3 [MAGE-3]); [1411] l. cell-based therapies (such as chimeric Antigen Receptor-T-cells (CAR-T) for example expressing anti-CD19, anti-EpCam or anti-mesothelin); [1412] m. bi-specific NK cell engagers having a specificity against an activating MK receptor such as NKG2D or CD16a; and [1413] n. adoptive transfer of tumour specific T-cells or LAK cells,
[1414] or optionally wherein the further therapy is chemotherapy, radiotherapy and surgical removal of tumours.
[1415] Radiotherapy may be single dose or in fractionated doses, either delivered to affected tissues directly or to the whole body.
[1416] In this arrangement, any of the features and embodiments of concept 46 apply mutatis mutandis.
[1417] In this aspect, the bispecific molecules include "bispecific antibodies" and antibody fusion proteins, including those formats and molecules described in concepts 37 to 40.
Arrangement 29. A nucleic acid that encodes a heavy chain and/or a light chain of a multispecific, bispecific or dual-binding antibody as defined in any one of arrangements 1 to 21. Arrangement 30. A vector comprising the nucleic acid as defined in arrangement 29; optionally wherein the vector is a CHO or HEK293 vector. Arrangement 31. A host comprising the nucleic acid as defined in arrangement 29 or the vector as defined in arrangement 30.
6. Uses for Antibodies and Immunocytokines
[1418] Unless otherwise apparent from the context, the uses for antibodies or fragments applies mutatis mutandis to the immunocytokines and multispecific (e.g. bispecific or dual-binding antibodies) of the invention.
[1419] Therapeutic
[1420] In one embodiment, the PD-L1 specific antibodies described herein and antigen binding fragments thereof can be used for therapeutic modulation of the PD-1/PD-L1 pathway. In one embodiment, the PD-L1 specific antibody or fragment thereof is as described in any concept, aspect or embodiment herein.
[1421] In one embodiment, the antibody or antibody binding fragment specifically binds to PD-L1 and thereby inhibits PD-L1 activity. In another embodiment, the antibody or antibody binding fragment specifically binds to PD-L1 and thereby inhibits binding of PD-L1 to PD-1. In another embodiment, the antibody or antibody binding fragment specifically binds to PD-L1 and thereby inhibits binding of PD-L1 to B7-1. In yet another embodiment, the antibody or antigen binding fragment thereof blocks PD-L1 induced T-cell suppression and thereby enhance anti-tumour immunity.
[1422] In yet another embodiment, the antibody or antigen binding fragment thereof is capable of stimulating one or more of the following activities: T-cell proliferation, IFN-.gamma., CD25 and/or IL-2 secretion in mixed lymphocyte reactions.
[1423] In one embodiment, the antibody or antigen binding fragment thereof specifically binds PD-L1 and inhibits PD-L1 induced cell proliferation, for example, tumour cell proliferation and/or inhibits, tumour cell survival. In another embodiment, the antibody or antigen binding fragment thereof specifically binds PD-L1 and thereby inhibits PD-L1 mediated suppression of T-cells, including, but not limited to, tumour reactive T-cells, thereby enhancing anti-tumour cytolytic T-cell activity. In other embodiments, the antibodies or binding fragments thereof as described herein inhibit tumour cell adhesion, motility, invasion and cellular metastasis, and reduce tumour growth. In other embodiments, the antibodies or binding fragments thereof can bind to cells expressing PD-L1, including tumour and non-tumour cells, and recruit, by means of interaction with the Fc portion of the antibody, cellular effector functions against the target cells by mechanisms including but not limited to antibody dependent cellular cytotoxicity (ADCC) and antibody dependent cellular phagocytosis (ADCP).
[1424] Still further embodiments include methods of treating a proliferative or invasion-related disease in a mammal by administering to the animal a therapeutically effective dose of an antibody or antigen binding fragment thereof. In another embodiment, the antibodies or antigen binding fragments thereof can be used in a method for treating a mammal suffering from a disease selected from: neoplastic or non-neoplastic disease, chronic viral infection; and a malignant tumour, wherein the method includes administering to the mammal a therapeutically effective dose of an antibody or antigen binding fragment thereof.
[1425] Still further embodiments include methods of treating a disease of immunological dysfunction in a mammal by administering to the animal a therapeutically effective dose of an antibody or antigen binding fragment thereof as described herein. Exemplary immunological dysfunction in humans includes diseases of neurological deficit, such as Alzheimer's disease.
[1426] It has further been proposed that an immune response, particularly an IFN.gamma.-dependent systemic immune response, could be beneficial for treatment of Alzheimer's disease and other CNS pathologies that share a neuroinflammatory component. WO2015/136541 (incorporated herein by reference) proposes treatment of Alzheimer's disease using an anti-PD-1 antibody (also see Baruch K. et al., PD-1 immune checkpoint blockade reduces pathology and improves memory in mouse models of Alzheimer's disease, Nature Medicine, 2016, 22(2):137-137).
[1427] Thus, in one embodiment, the antibody or antigen binding fragment thereof specifically binds PD-L1 and reduces the level of systemic immunosuppression in an individual by release of a restraint imposed on the immune system by PD-1/PD-L1 immune checkpoint pathway. In an aspect, PD-1/PD-L1 inhibitory immune checkpoint pathway blockade results in transient relief the systemic adaptive immune activity from suppression, which results in a transiently augmented immune response in the periphery, mainly manifested by elevation of IFN-.gamma. secretion by IFN-.gamma.-producing cells. Increased IFN-.gamma. activity may enable the brain's choroid plexus to allow selective leukocyte trafficking and infiltration of T-cells and monocytes into the damaged CNS, homing of these immune cells to sites of neurodegenerative pathology and neuroinflammation, and may modulate the environment to become less toxic and more permissive for clearance of toxic agents, rescue of neurons, regeneration and repair.
[1428] Thus, the PD-L1 mediated disease or condition is a neurodegenerative disease, disorder or condition. In one embodiment, the neurodegenerative disease, disorder or condition is Alzheimer's disease. In another embodiment, the neurodegenerative disease, disorder or condition is selected from amyotrophic lateral sclerosis, Parkinson's disease, Huntington's disease, primary progressive multiple sclerosis, secondary progressive multiple sclerosis, corticobasal degeneration, Rett syndrome, a retinal degeneration disorder selected from age-related macular degeneration and retinitis pigmentosa; anterior ischemic optic neuropathy, glaucoma, uveitis, depression, trauma-associated stress or post-traumatic stress disorder, frontotemporal dementia, Lewy body dementias, mild cognitive impairments, posterior cortical atrophy, primary progressive aphasia and progressive supranuclear palsy or aged-related dementia. In another embodiment, the neurodegenerative disease, disorder or condition is selected from Alzheimer's disease, amyotrophic lateral sclerosis, Parkinson's disease and Huntington's disease.
[1429] Anti-PD-L1 antibodies as described herein may be used in the treatment of Alzheimer's disease or other neurodegenerative diseases, optionally in combination with one or more other immune checkpoint inhibitors (such as anti-TIM-3 antibodies, anti-CTLA-4 antibodies, anti-TIGIT antibodies and anti-LAG-3 antibodies) or one or more other immune stimulators (such as anti-OX40 antibodies, anti-GITR antibodies, anti-CD137 antibodies, anti-ICOS antibodies and anti-CD40 antibodies, including those which are specifically described in Aspect 1a herein). Other combination partners include any of the the active agents as listed in claim 10 of WO2015/136541, which is incorporated herein by reference.
[1430] Any of the PD-L1 antibodies described herein (including at least the antibodies described in any of concepts 1 to 40, and the PD-L1 antibodies described in aspect 1a) may be used for the treatment of the neurodegenerative diseases, disorders or conditions described above.
[1431] Exemplary cancers in humans include a Merkel cell carcinoma, breast cancer, prostate cancer, basal cell carcinoma, biliary tract cancer, bladder cancer, bone cancer, brain and CNS cancer (e.g. glioblastoma), cervical cancer, choriocarcinoma, colon and rectum cancer, connective tissue cancer, cancer of the digestive system; endometrial cancer, esophageal cancer; eye cancer; cancer of the head and neck; nasopharyngeal cancer; gastric cancer; intra-epithelial neoplasm; kidney cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g. small cell and non-small cell); lymphoma including Hodgkin's and Non-Hodgkin's lymphoma including but not limited to DLBCL; Chronic lymphocytic leukaemia, melanoma; uveal melanoma, myeloma, neuroblastoma, oral cavity cancer (e.g., lip, tongue, mouth, and pharynx); ovarian cancer; pancreatic cancer, retinoblastoma; rhabdomyosarcoma; rectal cancer, renal cancer (renal cell carcinoma (RCC)), cancer of the respiratory system; sarcoma, skin cancer; stomach cancer, testicular cancer, thyroid cancer; uterine cancer, cancer of the urinary system, as well as other carcinomas and sarcomas. Further examples of virally induced cancers including; Nasopharyngeal carcinoma, certain Types of NHL (for example but not limited to EBV+ CNS lymphomas, DLBCL and BL, Hodgkins lymphoma (thought to be EBV driven) HPV-related cervical and head and neck squamous cell carcinomas); HBV hepatocellular carcinoma.
[1432] Exemplary chronic infections in humans include HIV, hepatitis B virus (HBV), and hepatitis C virus (HCV).
[1433] Proliferative or invasion-related diseases that can be treated with the antibodies or antigen binding fragments described herein include neoplastic diseases, and the metastasis associated with such neoplastic disease, such as, melanoma, uveal melanoma, skin cancer, small cell lung cancer, non-small cell lung cancer, salivary gland, glioma, hepatocellular (liver) carcinoma, gallbladder cancer, thyroid tumour, bone cancer, gastric (stomach) cancer, prostate cancer, breast cancer (including triple negative breast cancer), ovarian cancer, cervical cancer, uterine cancer, vulval cancer, endometrial cancer, testicular cancer, bladder cancer, lung cancer, glioblastoma, thyroid cancer, endometrial cancer, kidney cancer, colon cancer, colorectal cancer, pancreatic cancer, esophageal carcinoma, brain/CNS cancers, neuronal cancers, head and neck cancers (including but not limited to squamous cell carcinoma of the head and neck (SCCHN)), mesothelioma, sarcomas, biliary (cholangiocarcinoma), small bowel adenocarcinoma, pediatric malignancies, epidermoid carcinoma, sarcomas, cancer of the pleural/peritoneal membranes and leukaemia, including acute myeloid leukaemia, acute lymphoblastic leukaemia, and multiple myeloma. Treatable chronic viral infections include HIV, hepatitis B virus (HBV), and hepatitis C virus (HCV) in humans, simian immunodeficiency virus (SW) in monkeys, and lymphocytic choriomeningitis virus (LCMV) in mice.
[1434] The antibody or antigen binding fragment thereof can be administered alone, or in combination with other antibodies or chemo therapeutic drugs, radiation therapy or therapeutic vaccines. In one embodiment, the antibody or antigen binding fragment thereof is administered as an antibody-drug conjugate in which the antibody or antigen binding fragment thereof is linked to a drug moiety such as a cytotoxic or cytostatic agent. The use of antibody-drug conjugates for the local delivery of cytotoxic or cytostatic agents in the treatment of cancer allows targeted delivery of the drug moiety to tumours, and intracellular accumulation therein, where systemic administration of unconjugated drug may result in unacceptable levels of toxicity. Drugs in antibody drug conjugates can include, but are not limited to, daunomycin, doxorubicin, methotrexate, and vindesine. Toxins can also be used in antibody-toxin conjugates, including, for example, bacterial toxins such as diphtheria toxin, plant toxins such as ricin, small molecule toxins such as geldanamycin. The toxins may effect their cytotoxic and cytostatic effects by mechanisms including tubulin binding, DNA binding, or topoisomerase.
[1435] Detection
[1436] In another embodiment, the antibodies or antigen binding fragments can be used to detect the presence, absence and/or level of surface expressed PD-L1 expression in a sample. PD-L1 surface expression can be detected in vivo and/or in vitro and is useful in helping diagnose diseases or conditions that involve expression and/or overexpression of PD-L1.
[1437] In Vitro Diagnostic
[1438] In another embodiment, the PD-L1 specific antibodies or antigen binding fragments thereof can be used for the assessment of expression and localization of PD-L1 in a biological sample from a patient. In one embodiment, the biological sample is a tissue sample and PD-L1 expression is detected using known methods such as FLOW cytometry, IHC in fresh tissue, IHC in FFPE tissue and/or IHC in frozen tissue. In other embodiments, the biological sample is blood, plasma or serum.
[1439] In one embodiment, the antibody or antibody fragment described herein is labeled with a detectable moiety, for example, a radiolabel, fluorescent label, enzymatic label chemiluminescent labeled or a biotinyl group. Radioisotopes or radionuclides may include .sup.3H, .sup.14C, .sup.15N, .sup.35S, .sup.90Y, .sup.99Tc, .sup.115In, .sup.125I, .sup.131I, fluorescent labels may include rhodamine, lanthanide phosphors or FITC and enzymatic labels may include horseradish peroxidase, .beta.-galactosidase, luciferase, alkaline phosphatase. Additional labels include, by way of illustration and not limitation: enzymes, such as glucose-6-phosphate dehydrogenase ("G6PDH"), alpha-D-galactosidase, glucose oxydase, glucose amylase, carbonic anhydrase, acetylcholinesterase, lysozyme, malate dehydrogenase and peroxidase; dyes; additional fluorescent labels or fluorescers include, such as fluorescein and its derivatives, fluorochrome, GFP (GFP for "Green Fluorescent Protein"), dansyl, umbelliferone, phycoerythrin, phycocyanin, allophycocyanin, o-phthaldehyde, and fluorescamine; fluorophores such as lanthanide cryptates and chelates e.g. Europium etc (Perkin Elmer and Cisbio Assays); chemoluminescent labels or chemiluminescers, such as isoluminol, luminol and the dioxetanes; sensitisers; coenzymes; enzyme substrates; particles, such as latex or carbon particles; metal sol; crystallite; liposomes; cells, etc., which may be further labelled with a dye, catalyst or other detectable group; molecules such as biotin, digoxygenin or 5-bromodeoxyuridine; toxin moieties, such as for example a toxin moiety selected from a group of Pseudomonas exotoxin (PE or a cytotoxic fragment or mutant thereof), Diptheria toxin or a cytotoxic fragment or mutant thereof, a botulinum toxin A, B, C, D, E or F, ricin or a cytotoxic fragment thereof e.g. ricin A, abrin or a cytotoxic fragment thereof, saporin or a cytotoxic fragment thereof, pokeweed antiviral toxin or a cytotoxic fragment thereof and bryodin 1 or a cytotoxic fragment thereof.
[1440] In Vivo Diagnostic
[1441] In one embodiment, the antibody or antigen binding fragment thereof can be administered to a patient, wherein the antibody or antigen binding fragment is conjugated to a label. The presence of the label in the patient can be measured or observed, wherein a relatively high amount of the label may indicate a high risk of disease and a relatively low amount of the label may indicate a relatively low risk of the disease. In one embodiment, the label is a contrast agent, isotopic tag, or fluorescent marker, such as green fluorescent protein.
[1442] In one embodiment, the antibody or antigen binding fragment is used to monitor therapy that involves the use of other therapeutic agents, including, for example, chemotherapeutic agents or other antibodies that specifically bind PD-L1. In one embodiment, the antibody does not compete with the therapeutic PD-L1 antibodies.
[1443] Guide Patient Selection
[1444] In one embodiment, detection of PD-L1 expression can be used to guide patient selection. In one embodiment, the antibodies or antigen binding fragments thereof can be used to assist in patient selection for therapeutic antibody treatment with an anti-PD-L1 antibody, including, but not limited to anti-PD-L1 antibodies disclosed in WO2011/066389, entitled "Targeted Binding Agents Against B7-H1", which antibodies and sequences are incorporated herein by reference. In another embodiment, the antibodies or antigen binding fragments thereof can be used to assist in patient selection for treatment with immunotherapies such as anti-PD-L1, anti-CTLA4, anti-OX40, anti-PD-1, vaccines etc. In some cases, higher levels of PD-L1 may be indicative of successful therapy, whereas lower levels may indicate a reduced likelihood of success. Preferential expression of splice variants and/or protein processing may produce unique protein mixture profiles which may impact a patient's response to treatment or may change following treatment. These profiles may help to identify patients and define patient subsets who should receive treatment, continue to receive treatment or who should receive an alternative treatment. In another embodiment, the antibodies or antigen binding fragments thereof can be used for detection of PD-L1 isoforms. Patient samples can include, for example, blood, plasma, serum, sputum, saliva, urine, CSF, tears, exhaled exogenous particle samples, cell supernatant, cell or tissue lysate or tissue samples.
[1445] In one embodiment, the antibodies or antigen binding fragments thereof can be used to identify the presence, absence and/or level of PD-L1 expression at baseline, i.e., before treatment.
[1446] In another embodiment, the PD-L1 specific antibodies or antigen binding fragments thereof can be used as an exclusion marker to suggest treatment with therapies that do not target PD-L1. In another embodiment, the PD-L1 specific antibodies or antigen binding fragments thereof can be used as a prognostic marker for life expectancy. In particular, PD-L1 expression on tumours is linked to poor prognosis and life expectancy can be estimated based on historical data within tumour types.
[1447] Methods for detection of proteins are known, and include, for example, IHC, FLOW cytometry, Western blotting and Mass Spectroscopy, Immunoprecipitation, aptamers, immuno-PCR, and protein array.
[1448] Guide Therapy
[1449] The antibodies can be used to guide therapy. For example, the antibodies or antigen binding fragments thereof can be used to identify the presence, absence and/or level of PD-L1 expression during or after treatment. In one embodiment, the antibodies or antigen binding fragments thereof can be used as early response biomarkers to assist in patient management, drug approval and reimbursement. In another embodiment, the antibodies or antigen binding fragments thereof can be used to identify the presence, absence and/or level of PD-L1 expression to help guide therapy. For example, PD-L1 expression can help determine whether the treatment is effective, and hence, whether or not treatment should be continued, or whether the dose should be adjusted (increased or decreased) and whether a combination regimen should be changed. For example, in one embodiment, the PD-L1 specific antibodies or antigen binding fragments thereof can be used for determining receptor occupancy of PD-L1 on cells in a patient treated with anti-PD-L1 therapy for dose setting (PK/PD). In particular, receptor occupancy can be used as a measure of target engagement or target coverage. Estimates of the amount or duration of target engagement needed to elicit a biological or clinical response could be used to determine if a patient has been dosed sufficiently or not. In particular, the antibodies can be used to assist in evaluating the relationship between, dose, exposure, receptor occupancy, pharmacodynamic response and clinical benefit.
[1450] Monitor Efficacy of Therapy
[1451] In another embodiment, the PD-L1 specific antibodies or antigen binding fragments thereof can be used for patient monitoring, to help evaluate whether a course of treatment is effective and whether or not treatment should be continued. For example, in one embodiment, the antibodies or antigen binding fragments thereof can be used detect expression before a patient receives therapeutic treatment that targets PD-L1. In another embodiment, the antibodies or antigen binding fragments thereof can be used to detect expression during therapy or after a patient has received therapeutic anti-PD-L1 treatment. In another embodiment, the antibodies or antigen binding fragments thereof can be used as an early response marker to assist in the determination as to whether or not a course of therapy is effective and should be continued or discontinued. In one embodiment, the expression of PD-L1 is detected after washout, wherein the term "washout" refers to a period of time after which the administered drug has been eliminated from the body. In particular, expression of PD-L1 may be detected after washout if the patient is treated with anti-PD-L1 therapy that competes with the detection antibody. However, if the patient is treated with an antibody that does not compete with an anti-PD-L1 antibody, such as anti-CTLA-4 or anti-PD-1, detection can be performed without waiting for washout. In another embodiment, the detection antibody can bind to PD-L1 but not compete with a therapeutic antibody that binds to PD-L1. In this situation, washout may not be necessary. The washout period can vary depending upon many factors, but is generally a period of at least about 1, 2, 3, 4, 5, or 6 weeks and up to about 1, 2, 3, 4, 5 or 6 months from the most recent chemotherapy or immunotherapy treatment. The antibodies or antigen binding fragments thereof can be used to determine expression of PD-L1 on biopsy samples or on circulating tumour cells (CTC).
[1452] In one embodiment, labelled antibodies or antigen binding fragments thereof can be used to identify a peripheral correlate to enable non-invasive assessment of tumour status pre, during and post treatment.
[1453] Methods for detection of proteins are known, and include, for example, IHC, flow cytometry, Western blotting and Mass Spectroscopy, immunoprecipitation, aptamers, immuno-PCR., and protein array.
[1454] Identify Protein Binding Partners for PD-L1
[1455] In another embodiment, antibodies or antigen binding fragments thereof can be used as a capture reagent or detection reagent for examination of the protein binding partners of PD-L1 protein species in the context of a protein "pull-down." A protein "pull down" refers to immunoprecipitation of intact protein complexes, such as antigen along with any proteins or ligands that are bound to it--also known as co-immunoprecipitation (Co-IP). Co-IP works by selecting an antibody that targets a known protein that is believed to be a member of a larger complex of proteins. By targeting the known member with an antibody it may become possible to pull the entire protein complex out of solution and thereby identify unknown members of the complex. Complete understanding of the regulation of immune recognition through and PD-1 axis vs. CTLA-4 etc. is not fully understood. As such, antibodies and antigen binding fragments could improve knowledge of the interplay among accessory proteins and factors, which may determine a patient's propensity to respond to specific therapies or immunotherapy in general.
7. Pharmaceutical Compositions
[1456] Unless otherwise apparent from the context, the compositions for antibodies or fragments applies mutatis mutandis to the immunocytokines and multispecific (e.g. bispecific or dual-binding antibodies) of the invention.
[1457] In one embodiment, there is provided a pharmaceutical composition comprising an effective amount of an antibody or antigen binding fragment and a pharmaceutically acceptable carrier. An effective amount of antibody to be employed therapeutically will depend, for example, upon the therapeutic objectives, the route of administration, and the condition of the patient. In one embodiment, the composition includes other excipients or stabilizers.
[1458] Pharmaceutically acceptable carriers are known and include carriers, excipients, or stabilizers that are nontoxic to the cell or mammal being exposed thereto at the dosages and concentrations employed. Often the physiologically acceptable carrier is an aqueous pH buffered solution. Examples of physiologically acceptable carriers include buffers such as phosphate, citrate, and other organic acids; antioxidants including ascorbic acid; low molecular weight (less than about 10 residues) polypeptide; proteins, such as serum albumin, gelatin, or immunoglobulins; hydrophilic polymers such as polyvinylpyrrolidone; amino acids such as glycine, glutamine, asparagine, arginine or lysine; monosaccharides, disaccharides, and other carbohydrates including glucose, mannose, or dextrins; chelating agents such as Ethylenediaminetetraacetic acid (EDTA); sugar alcohols such as mannitol or sorbitol; salt-forming counterions such as sodium; and/or nonionic surfactants such as TWEEN.TM., polyethylene glycol (PEG), and PLURONICS.TM..
[1459] The antibodies or antigen binding fragments can be administered intravenously or through the nose, lung, for example, as a liquid or powder aerosol (lyophilized). The composition can also be administered parenterally or subcutaneously. When administered systemically, the composition should be sterile, pyrogen-free and in a physiologically acceptable solution, having due regard for pH, isotonicity and stability. These conditions are known to those skilled in the art.
[1460] Methods of administering a prophylactic or therapeutic agent (e.g., an antibody as disclosed herein), or pharmaceutical composition include, but are not limited to, parenteral administration (e.g., intradermal, intramuscular, intraperitoneal, intravenous and subcutaneous), epidural, and mucosal (e.g., intranasal and oral routes). In a specific embodiment, a prophylactic or therapeutic agent (e.g., an antibody as disclosed herein), or a pharmaceutical composition is administered intranasally, intramuscularly, intravenously, or subcutaneously. The prophylactic or therapeutic agents, or compositions may be administered by any convenient route, for example by infusion or bolus injection, by absorption through epithelial or mucocutaneous linings (e.g., oral mucosa, intranasal mucosa, rectal and intestinal mucosa, etc.) and may be administered together with other biologically active agents. Administration can be systemic or local. Each dose may or may not be administered by, an identical route of administration. In one embodiment, an anti-PD-L1 antibody or fragment as disclosed herein may be administered via multiple routes of administration simultaneously or subsequently to other doses of the same or a different anti-PD-L1 antibody or fragment as disclosed herein.
[1461] Various delivery systems are known and can be used to administer a prophylactic or therapeutic agent (e.g., an antibody or fragment as disclosed herein), including, but not limited to, encapsulation in liposomes, microparticles, microcapsules, recombinant cells capable of expressing the antibody, receptor-mediated endocytosis (see, e.g., Wu and Wu, J. Biol. Chem. 262:4429-4432 (1987)), construction of a nucleic acid as part of a retroviral or other vector, etc. In addition, pulmonary administration can also be employed, e.g., by use of an inhaler or nebulizer, and formulation with an aerosolizing agent. See, e.g., U.S. Pat. Nos. 6,019,968, 5,985,320, 5,985,309, 5,934,272, 5,874,064, 5,855,913, 5,290,540, and 4,880,078; and PCT Publication Nos. WO92/19244, WO97/32572, WO97/44013, WO98/31346, and WO99/66903, each of which is incorporated herein by reference their entirety.
[1462] In a specific embodiment, it may be desirable to administer a prophylactic or therapeutic agent, or a pharmaceutical composition as described herein locally to the area in need of treatment. This may be achieved by, for example, local infusion, by topical administration (e.g., by intranasal spray), by injection, or by means of an implant, said implant being of a porous, non-porous, or gelatinous material, including membranes, such as sialastic membranes, or fibres. When administering an anti-PD-L1 antibody or fragment, care must be taken to use materials to which the antibody does not absorb.
8. Kits and Articles of Manufacture
[1463] Unless otherwise apparent from the context, the kits and articles of manufacture for antibodies or fragments applies mutatis mutandis to the immunocytokines and multispecific (e.g. bispecific or dual-binding antibodies) of the invention.
[1464] In one embodiment, the invention provides a kit for detecting PD-L1 in a biological sample. The kit can be used to screen for PD-L1 related diseases. In one embodiment, the kit includes an antibody or antigen binding fragment and a means for determining whether the antibody or antigen binding fragment is bound to PD-L1 in a sample. In one embodiment, the antibody or antigen binding fragment is labelled. In another embodiment, the antibody or antigen binding fragment is an unlabelled primary antibody and the kit includes means for detecting the primary antibody. In one embodiment, the means for detecting includes a labelled secondary antibody that is an anti-immunoglobulin antibody. The antibody may be labelled with any suitable marker, including, for example, a fluorochrome, an enzyme, a radionuclide and a radiopaque material. Suitable antibodies and antigen binding fragments are described in detail above.
[1465] In one embodiment, a kit for detecting PD-L1 is provided, wherein the kit includes an antibody or antigen binding fragment described herein. In one embodiment, the kit may also include instructions and one or more reagents for detecting PD-L1. In one embodiment, the kit includes an antigen or antigen binding fragment described herein, along with instructions for preparing a formalin-fixed paraffin-embedded (FFPE) tissue sample for IHC and/or one or more reagents for IHC. In one embodiment, the kit includes an antigen or antigen binding fragment described herein as a primary antibody and a secondary antibody that specifically binds thereto. In one embodiment, the kit includes a labeled antigen or antigen binding fragment described herein, wherein the label includes a fluorescent label such as fluoroscein or rhodamine or an enzymatic reporter such as horseradish peroxidase (HRP) or alkaline phosphatase (AP). In one embodiment, the kit includes a blocking reagent that includes at least about 1% and up to about 5%, or between about 2% and 3%, or about 2% cold water fish skin gelatin protein (CWF) in a buffer, such as phosphate buffered saline (PBS). In one embodiment, the kit includes buffer for antigen retrieval, such as a citrate buffer, for example sodium citrate, at a concentration of at least about 1, 2, 5, or 10 mM and up to about 10, 15 or 20 mM and at a pH between about 5.5 and 9, or a pH of about 6.
[1466] In another embodiment, a kit for treating diseases involving the expression of PD-L1 is provided, wherein the kit includes an antibody or antigen binding fragment described herein and instructions to administer the antibody or antigen binding fragment to a subject in need of treatment. There is also provided a pharmaceutical or diagnostic pack or kit comprising one or more containers filled with one or more of the ingredients of the pharmaceutical compositions as disclosed herein, such as one or more anti-PD-L1 antibodies or fragments provided herein. Optionally associated with such container(s) can be a notice in the form prescribed by a governmental agency regulating the manufacture, use or sale of pharmaceuticals or biological products, which notice reflects approval by the agency of manufacture, use or sale for human administration, e.g., an authorisation number.
[1467] In another embodiment, an article of manufacture that includes a container in which a composition containing an antibody or antigen binding fragment described herein and a package insert or label indicating that the composition can be used to treat diseases characterized by the expression or overexpression of PD-L1 is provided. In one embodiment, there is provided a kit for treating and/or preventing a PD-L1-mediated condition or disease, the kit comprising an antibody or fragment as disclosed herein in any embodiment or combination of embodiments (and optionally a further therapeutic agent as described elsewhere herein) optionally in combination with a label or instructions for use to treat and/or prevent said disease or condition in a human; optionally wherein the label or instructions comprise a marketing authorisation number (e.g., an FDA or EMA authorisation number); optionally wherein the kit comprises an IV or injection device that comprises the antibody or fragment. In another embodiment, the kit comprises an antibody or antigen binding fragment thereof contained within a container or an IV bag. In another embodiment, the container or IV bag is a sterile container or a sterile IV bag. In another embodiment, the antibody or antigen binding fragment therefore is formulated into a pharmaceutical composition contained within a (sterile) container or contained within a (sterile) IV bag. In a further embodiment, the kit further comprises instructions for use.
9. Examples
Example 1--Antigen Preparation, Immunization Procedures, and Hybridoma Generation
[1468] The following example provides a detailed description of the generation and identification of a panel of anti-human PD-L1 monoclonal antibodies using the KyMouse.TM. system (see, e.g., WO2011/004192, WO2011/158009 and WO2013/061098). To this end, genetically engineered mice containing a large number of human immunoglobulin genes were immunized with soluble recombinant human PD-L1 or surface expressed human PD-L1 displayed on mouse embryonic fibroblast (MEF) cells. Various immunization regimens, including conventional intraperitoneal injections as well as a rapid immunisation at multiple sites (RIMMS) regimen were set up, boosting animals over several weeks (see detailed methods below). At the end of each regimen, secondary lymphoid tissue such as the spleen, and in some cases, the lymph nodes were removed. Tissues were prepared into a single cell suspension and fused with SP2/0 cells to generate a stable hybridoma cell line.
[1469] Materials and Methods
[1470] a) Generation of Stably Transfected MEF and CHO-S Cells Expressing Human PD-L1:
[1471] Full length human PD-L1 sequence (SEQ ID No:1 also known as 137-H1) was codon optimized for mammalian expression and cloned into an expression vector under the CMV promoter flanked by 3' and 5' piggyBac specific terminal repeat sequences, facilitating stable integration into the cell genome (see: "A hyperactive piggyBac transposase for mammalian applications"; Yusa K., et al., Proc. Natl. Acad. Sci. USA., 108(4): 1531-6, 2011 Jan. 25). Furthermore, the expression vector contained a puromycin selection cassette to facilitate stable cell line generation. The human PD-L1 expression plasmid was co-transfected with a plasmid encoding piggyBac transposase into an in-house derived mouse embryonic fibroblast (MEF) cell line (embryos used to generate this line were obtained from a 12955 crossed to C57/BL6 female mouse) and CHO-S cells using the FreeStyle Max transfection reagent (Invitrogen) according to manufacturer instructions. 24 hours after transfection, the media was supplemented with puromycin and grown for at least two weeks to select a stable cell line with complete medium being exchanged every 3 to 4 days. The expression of hPD-L1 was assessed by flow cytometry using an anti-human PD-L1-PE conjugated antibody (eBioscience). Complete MEF media was made up of Dulbecco's Modified Eagle's Medium (Gibco) supplemented with 10% v/v fetal bovine serum (Gibco). Complete CHO-S media was made up of CD-CHO media (Gibco) supplemented with 8 mM Glutamax (Gibco). Transfected CHO cells were used for screening purposes (see Example 2).
[1472] b) Preparation of MEF Cells for Mouse Immunizations:
[1473] Cell culture medium was removed and cells washed once with 1.times.PBS. Cells were treated for 5 minutes with trypsin to loosen cells from tissue culture surface. Cells were collected and trypsin neutralized by the addition of complete MEF media. Cells were then centrifuged at 300 g for 10 minutes and washed with 25 mL of 1.times.PBS. Cells were counted and resuspended at the appropriate concentration in 1.times.PBS.
[1474] c) Immunisations with PD-L1
[1475] Genetically engineered Kymouse.TM. HK strain, containing human immunoglobulin genes producing human kappa (HK) light chain antibodies (Lee et al, Nature Biotechnology, 32, 6-363, 2014) were immunized by various immunisation regimens for the generation of human anti-PD-L1 antibodies.
[1476] Mice were immunised either with soluble recombinant hPD-L1 (R&D Systems, 156-67, Fc chimera) using a modified sub-cutaneous immunisation procedure (RIMMS; modified after Kilpatrick et al., "Rapid development of affinity matured monoclonal antibodies using RIMMS"; Hybridoma. 1997 August; 16(4):381-9, hereafter referred to as KM031), or by using soluble recombinant hPD-L1 in a prime-rest-boost regime by sub-cutaneous administration (hereafter referred to as KM032) or by combination of soluble recombinant hPD-L1 and stably transfected MEF cells expressing hPD-L1 administered intra-peritoneally (hereafter referred to as KM033). Sigma Adjuvant System was used for all immunisations and rest intervals were usually between 2 and 3 weeks. Where protein was used as the immunogen, CpG (Hokkaido System Science) was also administered. Serum from serial or terminal blood samples were analysed for the presence of specific antibodies by ELISA and flow cytometry and the titre data was used (where possible) to select mice to be used for hybridoma fusions. A further regimen, KM042 immunising with MEF-PD-L1 cells alone, or protein alone in a prime-rest-boost setting, was also performed, but out of six antibodies confirmed to bind to hPD-L1, no neutralising antibodies were identified.
[1477] d) Cloning and Expression of Recombinant Proteins
[1478] DNA sequences encoding PD-L1 were purchased as synthetic DNA strings and cloned into appropriate mammalian expression vectors for transient expression in Expi293 and CHO cells. The sequence listing shows the sequences of the antigens, where available, and affinity tags, for purification/labelling (shown in bold and underlined), see Seq ID Nos:3 to 6.
[1479] e) Determining Serum Titre by Reverse PD-L1 ELISA Protocol
[1480] Titres in mouse serum samples were determined using a reverse PD-L1 ELISA protocol. Anti-mouse IgG capture antibody (Southern Biotech) (4 .mu.g/mL diluted in PBS, 50 .mu.L/well) was adsorbed to 96 well low auto-fluorescent, high protein binding plates (Costar) overnight at 4.degree. C. Excess IgG was removed by washing three times with PBS-Tween (0.1% v/v) and the wells were blocked with 1% w/v bovine serum albumin (BSA, Sigma) in PBS for 1 hour at room temperature, after which plates were washed three times with PBS-Tween (0.1% v/v). Serial ten-fold dilutions of mouse serum were prepared, diluting samples in reagent diluent (0.1% w/v BSA/PBS). 50 .mu.L/well of this titration was then added to ELISA plates. To determine the change in activity level due to immunization, serum from each animal prior to immunization was diluted to 1/100 in reagent diluent and 50 .mu.L/well added to the ELISA plate. Following incubation, plates were washed as before to remove unbound proteins. Biotinylated hPD-L1-his (in-house generated protein, Seq ID No: 3, labelled in-house using Sulfo-NHS-LC-Biotin (Thermo)), used at 100 ng/mL in reagent diluent; 50 .mu.L/well) was then added to the plates and incubated at room temperature for 1 hour. Unbound biotinylated hPD-L1 was removed by washing with PBS-Tween (0.1% v/v), while the remaining biotinylated hPD-L1 was detected by addition of streptavidin-HRP (Sigma) diluted 1/10,000 in reagent diluent. Following incubation for 1 hour at room temperature, plates were washed as described before and 50 .mu.L TMB (Sigma) was added to the plate. The reaction was stopped by adding 50 .mu.L 1M sulphuric acid (Fluka Analytical). The OD at 450 nm was measured on an Envision plate reader (PerkinElmer). Titres were not performed for KM032 as only one mouse was immunised. For KM031, titres were performed on terminal bleeds only.
[1481] f) Determination of Serum Titres by Flow Cytometry Using CHO-S Expressed hPD-L1
[1482] CHO-S cells expressing hPD-L1, suspended in FACS buffer (PBS+1% w/v BSA+0.1% w/v sodium azide) were distributed to a 96-well, V-bottom plate (Greiner) at a density of 10.sup.5 cells per well. A titration of mouse serum was prepared, diluting samples in FACS buffer. 25 .mu.L/well of this titration was then added to the cell plate. To determine the change in activity level due to immunization, serum from each animal prior to immunization was diluted to 1/100 in FACS buffer and 25 .mu.L/well added to the cells. Cells were incubated at 4.degree. C. for 1 hour. Cells were washed twice with 150 .mu.L PBS, centrifuging after each wash step and aspirating supernatant (centrifuged at 300.times.g for 3 minutes). To detect antibody binding, PE goat-anti-mouse IgG (Jackson ImmunoResearch) was diluted 1/500 in FACS buffer and 50 .mu.L was added to the cells. Cells were incubated 1 hour at 4.degree. C. in the dark, then washed twice with 150 .mu.L PBS as above. To fix cells, 100 .mu.L 2% v/v paraformaldehyde was added and cells incubated for 30 minutes at 4.degree. C. Cells were then pelleted by centrifugation at 300.times.g and the plates resuspended in 100 .mu.L of FACS buffer. PE signal intensity (geometric mean) was measured by flow cytometry using a BD FACS Array instrument. Titres were performed by this method for KM033 only.
[1483] g) Murine Tissue Isolation and Preparation
[1484] Following final boost, mice were culled and spleens were excised from immunized mice, washed in 1.times.PBS and kept on ice until further processing. Tissues were prepared in buffer containing 1.times.PBS (Invitrogen) and 3% heat-inactivated FBS (Invitrogen). Splenocytes were dispersed by mashing the tissue through a 45 .mu.m strainer (BD Falcon) and rinsing with 30 mL 3% FBS/PBS buffer before centrifugation at 700 g for 10 minutes at 4.degree. C. To remove red blood cells, the pelleted splenocytes were resuspended in 4 mL Red Blood Cell Lysis Buffer (Sigma). After 4 minutes of incubation, the lysis reaction was stopped by addition of 3% FBS/1.times.PBS buffer. Cell clumps were filtered out with a 45 .mu.m strainer. The remaining splenocytes were pelleted for further procedures. For KM031 and KM032, axillary, inguinal and mesenteric lymph nodes were also removed and placed in sterile 1.times.PBS on ice until further processing. The lymph nodes were processed separately from splenocytes. Lymph node cells were prepared as above, but did not undergo red blood cell lysis. The remaining lymph node cells were pelleted for further procedures.
[1485] h) Hybridoma Fusion
[1486] Spleen and lymph node cells were pooled from KM031 and also from KM032 and subjected to a negative selection method using the MACS.RTM. Separation system. Briefly, where lymph nodes were used those cells were pooled with the splenocytes from the corresponding mice after red blood cell lysis and total cell number determined. Cells were resuspended in 100 .mu.L 3% FBS/PBS buffer per 10.sup.7 cells, before adding 10 .mu.L of Pan B Cell Biotin-Antibody Cocktail (Cat#130-095-813) per 10.sup.7 total cells and 10 .mu.L of anti-IgD-Biotin antibody (Cat#130-096-979) and incubated for 10 minutes at 4.degree. C. 2 mL FBS/PBS buffer was added and the cells were spun down at 700 g for 10 minutes. The supernatant was aspirated completely and 100 uL fresh buffer was added, then 30 uL Anti-Biotin MicroBeads (Cat#130-047-302) was added per 10.sup.7 cells along with 7 .mu.L Anti-Mouse IgM MicroBeads (Miltenyi Biotec). The cells were incubated for 15 minutes in the refrigerator. The cells/MicroBeads mixture was then applied to a pre-wetted LD column (Miltenyi Biotec) placed in a magnetic MACS Separator and washed with 3% FBS/PBS buffer. The unlabelled cells that flowed through the column were collected in 3% FBS/PBS buffer.
[1487] KM033 cells were subjected to a positive selection method using the MACS.RTM. Separation system. After red blood cell lysis, splenocytes were resuspended in 80 .mu.L 3% FBS/PBS buffer per 10.sup.7 cells, before adding anti-mouse IgG1 (Cat#130-047-101) plus anti-mouse IgG2a+b MicroBeads (Cat#130-047-201) and incubated for 15 minutes at 4.degree. C. The cell/MicroBead mixture was then applied to a pre-wetted LS column (Miltenyi Biotec) placed in a magnetic MACS Separator and washed with 3% FBS/PBS buffer. IgG positive cells were collected in the labelled, column-bound fraction in 3% FBS/PBS buffer.
[1488] Enriched B-cells were treated with CpG (Hokkaido System Science) overnight (final concentration 25 .mu.M) and the following day washed once in BSA fusion buffer (0.3 M D-Sorbitol, 0.11 mM calcium acetate hydrate, 0.5 mM magnesium acetate tetrahydrate and 0.1% BSA (v/w), adjusted to pH 7.2). Washed cells were resuspended in 200 .mu.L BSA fusion buffer and cell count determined. SP2/0 cells were treated in the same way, but washed twice instead of once with BSA fusion buffer. B-cells fused at a ratio of 3:1 with SP2/0 myeloma cells by electrofusion using a BTX ECM 2001 Electro Cell Manipulator (Harvard Apparatus). Each fusion was left overnight in recovery medium (Dulbecco's Modified Eagle's Medium (high glucose, no phenol red) supplemented with OPI (Sigma), 1.times.L-Glutamax (Gibco), 20% FBS (Gibco, batch-tested for hybridoma) and 0.05 mM 2-mercaptoethanol), then resuspended in 1 part recovery medium and 9 parts semi-solid medium (ClonaCell-HY Hybridoma Selection Medium D, Stemcell Technologies) and seeded onto 10 cm petri dishes. Visible colonies were picked 12 days later into 96-well plates and cultured for another 2 to 3 days prior to screening.
Example 2--Hybridoma Supernatant Screening
[1489] After generation of hybridoma clones, the hybridoma supernatant was assessed in a sequential primary and secondary screen and appropriate hybridoma clones selected based on criteria of antibody binding to human PD-L1 and receptor neutralization activity. In the screening cascades described, 9317 hybridoma clones were tested and 120 identified as primary hits. Thereafter, 36 hybridoma clones were confirmed by using secondary screening criteria (see details in Materials and Methods and Table 1). Among the clones identified by secondary screen, four clones were selected by the inventors to be part of the antibody shortlist, dependent upon desired selection criteria (see details in Example 3).
[1490] Materials and Methods
[1491] a) Primary Screen--Binding to Cell-Expressed Human PD-L1
[1492] Supernatants collected from hybridoma cells were screened for the ability of secreted antibodies to bind to hPD-L1 expressed on the surface of CHO-S cells. To determine CHO-S hPD-L1 binding, cells were plated in black-walled, clear-bottom tissue culture treated 384-well plates (Costar) at 1.times.10.sup.4/well in 80 .mu.L F12 media (Gibco) supplemented with 10% FBS (Gibco) and cultured overnight at 37.degree. C., 5% CO.sub.2. Culture media was removed from 384-well assay plates. At least 5 .mu.L of hybridoma supernatant or 5 .mu.L MIH1 at 2 .mu.g/mL in hybridoma maintaining media (HMM) or isotype IgG1 control antibody (referred to in some instances as Cm7, Sigma M9269, at a final concentration of 1 .mu.g/mL) diluted in HMM were added to each well. HMM was made up of Advanced DMEM (Gibco) supplemented with 1.times. Glutamax (Gibco), 20% v/v FBS (Gibco), 0.05 mM .beta.-Mercaptoethanol, 1.times.HT supplement (Gibco), and 1.times. penicillin/streptomycin (Gibco). 45 .mu.L FACS buffer containing 500 ng/mL IRDye 800CW anti-Mouse Ab (LICOR) and 0.2 .mu.M DRAQ5 (Biostatus) was added to each well. DRAQ5 was not added to background wells. Plates were incubated for 1 hour at 4.degree. C. Supernatant was aspirated and 25 .mu.L 4% v/v paraformaldehyde added and plates were incubated for 15 minutes at room temperature. Plates were washed twice with 100 .mu.L PBS and then the wash buffer was completely removed. Fluorescence intensity was read by scanning plates using an Odyssey Infrared Imaging System (LI-COR.RTM.). Anti-mouse binding (800 nm channel) was normalised to cell number (700 nm channel) according to the LI-COR.RTM. recommended algorithm. Percent effect was calculated as detailed below (Equation 1). Total binding was defined using reference antibody at a final assay concentration of 0.2 .mu.g/mL. Non-specific binding was defined using mouse IgG1 isotype control (Sigma) at a final assay concentration of 0.2 .mu.g/mL. Criteria for hit selection were based on assay signal and visual inspection of scanned plates.
Equation 1 Calculation of Percentage Effect from Primary Screen ( LI - COR ) and HTRF Using 800 % Resp values ( LI - COR ) or 665 / 620 nm ratio ( see Equation 2 ) ( HTRF ) Percent effect = ( sample well - non - specific binding ) .times. 100 ( total binding - non - specific binding ) Non - specific binding = values from wells containing isotype control mouse IgG 1 Total Binding = values from wells containing reference antibody ##EQU00001##
[1493] b) Primary Screen: Binding to Recombinant Human PD-L1
[1494] In parallel to screening for binding to CHO-S expressed PD-L1, supernatants collected from hybridoma wells were screened for the ability of secreted antibodies to bind to hPD-L1 expressed as a recombinant protein (produced in-house). Binding of secreted antibodies to recombinant PD-L1 were identified by HTRF.RTM. (Homogeneous Time-Resolved Fluorescence, Cisbio) assay format using biotinylated hPD-L1. 10 .mu.L hybridoma supernatant was transferred to a white 384 well, low-volume, non-binding surface polystyrene plate (Greiner). 5 .mu.L 230 nM biotinylated hPD-L1 his diluted in HTRF assay buffer (PBS (Sigma)+0.53 M KF (Sigma)+0.1% w/v BSA (Sigma)) was pre-incubated with 10 .mu.L hybridoma supernatant or 10 .mu.L reference antibody diluted to 3.3 nM working concentration for 1 hour at room temperature. For negative control wells, 10 .mu.L HMM was added. Streptavidin D2 (Cisbio), and goat anti-mouse IgG (Southern Biotech) labelled with Europium cryptate (Cisbio) were both diluted 1/100 in HTRF assay buffer, and 5 .mu.L of this mixture added to all wells. The plate was left to incubate in the dark for 2 hours prior to reading time resolved fluorescence at 620 nm and 665 nm emission wavelengths using an EnVision plate reader (Perkin Elmer). More details of the HTRF.RTM. assay technology can be found in Mathis (1995) Clinical Chemistry 41(9), 1391-1397.
[1495] Data were analysed by calculating 665/620 ratio and percent effect for each sample according to Equation 2 and Equation 1 respectively.
665/620 ratio=(sample 665/620 nm value).times.10000 Emotion 2: Calculation of 665/620 ratio
[1496] In general, criteria for hit selection were based on greater than or equal to 10 percent effect. In some instances, hit selection was based on greater than or equal to 20 percent effect.
[1497] Progression to secondary screen was based on a combination of data from recombinant PD-L1 binding hits and binding to human PD-L1 expressed on CHO cells.
[1498] c) Secondary Screen: Binding to Cell Expressed Recombinant Human PD-L1 or Natively Expressed hPD-L1 and Binding Affinity
[1499] To determine whether wells selected using the primary screen selection criteria had the required characteristics set by the inventors, a number of assays were performed. Hybridoma clones selected as hits from primary screening were cultured for 3 days and the supernatants collected from hybridoma cells were tested to assess whether the secreted antibodies that bind to in some cases CHO-S expressed hPD-L1, or in some cases ES2 cells. In addition, the ability to neutralise recombinant hPD-1 Fc, binding to CHO-S hPD-L1 or ES2 cells was also assessed. Binding of antibodies to human PD-L1 by SPR was also tested.
[1500] d) Binding to Cell Expressed hPD-L1 and Neutralisation and hPD-L1 Binding to PD-1
[1501] Binding of hybridoma supernatants was tested for ability to bind to either CHO-S cells expressing hPD-L1 or ES2 cells. CHO-S cells expressing hPD-L1 (generated in-house), or ES2 cells (ATCC CRL-1978) natively expressing hPD-L1 were diluted in FACS buffer and were distributed to a 96-well, V-bottom plate (Greiner) at a density of 0.5 to 1.times.10.sup.5 cells per well. Cells were washed with 150 .mu.L PBS and centrifuged at 300 g for 3 minutes. Supernatant was aspirated and 150 .mu.L PBS added. This wash step was repeated.
[1502] 50 .mu.L hybridoma supernatant or purified hybridoma material was added to the washed cells, to which 500 ng/mL human PD-1 Fc (in-house, Seq ID No:6) was added. Reference antibody was added to medium at 2 .mu.g/mL. Where purified material was used, titrations were prepared from a top concentration of 600 nM before addition to cells. When supernatants were used, neat supernatant, and three serial two-fold dilutions were added to cells. Cells were incubated at 4.degree. C. for 30 minutes. Cells were washed twice with 150 .mu.L FACS buffer, centrifuging at 300 g for 3 minutes after each wash step and aspirating supernatant.
[1503] To detect antibody and receptor binding, 50 .mu.L goat anti-human IgG-PE (Jackson ImmunoResearch) and APC anti-mouse IgG (Jackson ImmunoResearch) diluted 1/500 in FACS buffer was added to the cells. Cells were incubated for 30 minutes at 4.degree. C. in the dark. Cells were washed twice as above and resuspended in FACS buffer for analysis. PE and APC signal intensity (geometric mean) was measured by flow cytometry using a BD FACS Array instrument. Data was plotted as geometric mean values without further calculation.
[1504] e) Determination of Affinity by Surface Plasmon Resonance
[1505] Label-free surface plasmon resonance (SPR) analysis was carried out on the ProteOn XPR36 (BioRad) array SPR machine. An anti-mouse IgG capture surface was created on a GLC biosensor chip using amine coupling of an anti-mouse IgG from GE Healthcare. Test antibodies were captured on this surface and human PD-L1 (in-house) was used as the analyte at 256 nM, 64 nM, 16 nM, 4 nM and 1 nM. The assay was carried out at 25.degree. C. using HBS-EP (Teknova H8022). Buffer alone was used to reference the binding sensorgrams. The data was analysed using the 1:1 model inherent to the ProteOn XPR36 analysis software. In some instances, hybridoma supernatants were used as the source of antibody; in other instances, antibody was purified from hybridoma supernatant prior to analysis (see below). In some instances, a Protein A/G capture surface was used. This was created on a GLM biosensor chip using amine coupling of Protein A/G from Biorbyt [1506] f) Purification of Antibodies from Hybridoma Supernatant
[1507] Protein G resin in a gravity-flow column was first washed with water, then 50 mM sodium hydroxide or IgG Elute (Pierce) and was then equilibrated with tissue culture grade PBS. Clarified hybridoma supernatant containing 10% v/v 10.times. tissue culture grade PBS was applied several times to the equilibrated protein G column. Resin was washed with tissue culture grade PBS to remove unbound material. Antibody was then eluted with IgG Elute (Pierce) and the eluted fraction was then neutralized with 100 mM final TRIS, at pH 8.0. The eluted fraction was then concentrated down to <1.5 mL by centrifugation in a 10 kDa cut-off centrifugal filter unit. Tissue culture grade PBS was then added and the sample was concentrated down again to <1.5 mL. Protein concentration was quantified at OD.sub.280 using the molar extinction coefficient inherent to the Nanodrop for IgG. Finally, sample was analysed on a SDS-PAGE to assess purity.
TABLE-US-00001 TABLE 1 Summary of hybridoma clone screening Number of Number of Number of Number of Primary hits secondary Lead Experiment hybridoma cherry hits Candidate ID screened picked confirmed mAbs KM031 1872 41 4 0 KM032 115 14 6 1 KM033 7330 66 26 1
Example 3--Antibody Shortlist Selection Criteria
[1508] Binding to hPD-L1 natively expressed on ES2 cells, and neutralisation of recombinant human PD-1 binding to ES2 cells were used as criteria for secondary screen hit selection. Hits to progress to purification and further characterisation were determined by a combination of high affinity for human PD-L1 and neutralisation capacity.
[1509] After the selection and characterization of shortlisted antibodies, their fully-human variable domains were recovered using RT-PCR using a mixture of forward and reverse primers. Antibodies were reformatted into a human IgG1 backbone and expressed using a transient expression system in CHO-S cells.
[1510] Materials and Methods
[1511] a) RNA Isolation from Hybridoma Cells
[1512] Total RNA was extracted from hybridoma cells using TRIzol.TM. Reagent (Invitrogen). The quantity and quality of the isolated RNA was analysed spectrophotometrically.
[1513] b) Antibody Variable Domain Recovery by RT-PCR
[1514] Selected clones were used to prepare total RNA, which was used in an RT-PCR reaction to recover the heavy and light chain V-regions. Murine IgG-specific reverse primers and human Ig-leader sequence-specific forward primer sets were used for the heavy chains. Murine kappa constant region specific reverse primers and human kappa-leader sequence specific forward primer sets were used for the kappa light chains. The RT-PCR products were separated by agarose gel electrophoresis with the DNA of the predicted size being gel purified and sequenced in the forward and reverse directions. Alternatively, the RT-PCR products were subcloned into a cloning vector and DNA of individual colonies submitted for, sequencing.
Example 4--Selection of Final Lead Panel
[1515] Recombinantly expressed antibodies were analysed by SPR to confirm binding to cynomolgus monkey PD-L1, as well as human PD-L1. Antibodies were also tested in a dendritic cell-T-cell mixed lymphocyte reaction (MLR) for ability to enhance IFN.gamma. production (FIG. 1). Antibodies with consistent immune-stimulatory effects in the MLR, and binding to both human and cynomolgus PD-L1 were selected as the final lead panel--these were designated as clone 84G09 and clone 1D05. Data in FIG. 1 is from a single experiment. A further five experiments were conducted and showed similar results (84G09 showed activity in 3 out of 5 experiments, 1D05 showed activity in 3 out of 4 experiments, 1A01 showed activity is 1 out of 3 experiments and 8609 showed activity in 0 out of 3 experiments). One further experiment failed (including positive control).
[1516] Materials and Methods
[1517] a) Surface Plasmon Resonance for Analysis of Antibodies with Human Constant Region
[1518] Label-free surface plasmon resonance (SPR) analysis was carried out on the ProteOn XPR36 (BioRad) array SPR machine. An anti-human IgG capture surface was created on a GLC biosensor chip using a combination of anti-human Fc antibodies (Jackson Labs 09-005-008,109-006-008 and 309-006-008) by amine coupling. Test antibodies were captured on this surface and human PD-L1-his and cynomolgus monkey PD-L1-FLAG (in-house, Seq ID No: 5) was used as the analyte at 128 nM, 32 nM, 8 nM, 2 nM, 0.5 nM and 0 nM. The data was analysed using the 1:1 model inherent to the ProteOn XPR36 analysis software.
[1519] b) Dendritic Cell--T-Cell MLR (Mixed Lymphocyte Reaction)
[1520] Dendritic cells were generated from monocytic precursors. Monocytic precursors were isolated from peripheral blood mononuclear cells (PBMCs) isolated using Ficoll-Paque plus (GE Healthcare) density gradient centrifugation from leukoreduction system chambers, (NHSBT). Monocytes were isolated from PBMCs using negative selection magnetic separation beads (Miltenyi Biotec). Monocytes were plated in 96-well, flat-bottom TC plates at 5.times.10.sup.4/well and 1.times.10.sup.4/well and cultured with cytokines GM-CSF and IL-4 (both Peprotech) at 100 ng/mL for 7 days in culture media (Advanced RPMI (Gibco) supplemented with 10% v/v FBS and 2 nM glutamine (culture medium).
[1521] After 7 days, T-cells were purified from allogeneic PBMC using negative selection magnetic separation beads (Miltenyi). After purification, the isolation buffer was removed by centrifugation and aspiration. The cells were resuspended at 1.times.10.sup.6 cells/mL in culture medium, and 100 .mu.L of T-cells were added to all wells with the exception of the DC-only wells. An additional 100 .mu.L of culture medium was added to the DC-only and T-cell-only wells. Serial three-fold dilutions of antibodies were prepared in culture medium (top concentration 60 nM final). 10 .mu.L of each dilution was added to cells.
[1522] The cells were incubated for five days at 37.degree. C. After this period IFN-.gamma. was measured by Duoset ELISA (R&D Systems) according to manufacturer's instructions.
Example 5--in Depth Characterisation of Lead Antibodies
[1523] Lead antibodies 84G09 and 1D05 were subjected to in-depth characterisation, including SPR at 37.degree. C., full titrations of antibodies in neutralisation assays, and confirmation of binding to PD-L1 but not PD-L2. Antibodies were also expressed with a human IgG4(PE) constant region (Seq ID No:199) for analysis by mixed lymphocyte reaction. Lead antibodies retain sub-nanomolar affinity at 37.degree. C., and show potent neutralisation of PD-L1 binding to both PD-1 and CD80. Antibodies do not cross-react with PD-L2, bind natively expressed PD-L1 on dendritic cells, and are potent stimulators of IFN.gamma. production in an MLR.
[1524] a) Human PD-L1/PD-1 Neutralisation Assay (ELISA)
[1525] PD-1 Fc (in house, Seq ID No:6) diluted to 1 .mu.g/mL was adsorbed to 96-well, low auto-fluorescent, high protein binding plates (Costar) overnight at 4.degree. C. Excess protein was removed by washing with PBS-Tween (0.1% v/v) and the wells were blocked with 1% w/v bovine serum albumin (BSA, Sigma) in PBS for 1 hour at room temperature, after which plates were washed as described previously. 30 .mu.L titration (1/3 dilution) of antibody was added to a 96-well non-binding plate diluted in ELISA assay buffer (PBS+0.1% BSA). 30 .mu.L biotinylated PD-L1 his (in-house, Seq ID No:3) at 50 nM working concentration (25 nM final assay concentration [FAC]) was added to the plate excluding control wells where 30 .mu.L ELISA assay buffer was added. The plate was incubated for 30 minutes before transferring 50 .mu.L to the coated plates. The coated plates were incubated for 1 hour at room temperature. Excess protein was removed by washing with PBS-Tween (0.1% v/v). PD-L1 binding was detected using streptavidin labelled Europium (Perkin Elmer) diluted 1/1000 in DELFIA assay buffer (Perkin Elmer). Plates were washed with TBS (Tris buffered saline)-Tween (0.1% v/v) and 50 .mu.L/well of DELFIA Enhancement solution (Perkin Elmer) was added to the plate. Time-resolved fluorescence was measured at 615 nm on an Envision plate reader (PerkinElmer). Percentage of specific binding was calculated using Equation 3. IC.sub.50 values were determined using GraphPad Prism software by curve fitting using a four-parameter logistic equation (Equation 4). Results are shown in FIG. 2 and summarised in Table 2.
Equation 3 Percentage of receptor binding ( ELISA ) Based on fluorescence at 615 nm % of specific binding = sample value - non - specific binding total binding - non - specific binding .times. 100 Total binding = biotinylated PD - L 1 ( no antibody ) Non - specific binding = no biotinylated PD - L 1 Equation 4 Four Parameter logistic calculation Y = Bottom + ( Top - Bottom ) / ( 1 + 10 ^ ( ( Log IC 50 - X ) * HillSlope ) ) X = logarithm of concentration . Y = specific binding ( Equation 3 ) Top and Bottom = Plateaus in same units as Y ( specific binding ) ##EQU00002##
[1526] Log IC.sub.50 in same units as X. Y starts at Bottom and goes to Top with a sigmoid shape. Specific binding decreases as X increases.
[1527] c) CHO Human PD-L1/PD-1 or CD80 Neutralisation Assay (Flow Cytometry)
[1528] CHO-S cells untransfected (referred to as WT) or transfected with hPD-L1 were diluted in FACS buffer and were distributed to a 96-well V-bottom plate (Greiner) at a density of 1.times.10.sup.5 cells per well in 50 .mu.L. Biotinylated human PD-1-Fc (in-house expressed, Seq ID No:6) or CD80-Fc (R&D Systems) were prepared as a titration from 1 .mu.M final assay concentration (FAC), 1/2 dilution series in FACS buffer. Antibody titrations were prepared from 300 nM working concentration, 150 nM FAC, as a 1/3 dilution series in FACS buffer. Biotinylated PD-1 or CD80 were diluted in FACS buffer to 60 nM working concentration, 30 nM FAC. Plates were centrifuged at 300.times.g for 3 minutes to supernatant aspirated. 25 .mu.L ligand and 25 .mu.L antibody solution (or 50 .mu.L of ligand titration) were added to cells and incubated at 4.degree. C. for 1 hour. Cells were washed with 150 .mu.L of PBS and centrifuged at 300 g for 3 minutes. Supernatant was aspirated and 150 .mu.L PBS added. This wash step was repeated. Presence of bound CD80 or PD-1 was detected by addition of 50 .mu.L of streptavidin-AlexaFluor 647 (Jackson ImmunoResearch) diluted 1/500 in FACS buffer. Cells were incubated 30 minutes at 4.degree. C. in the dark. Cells were washed as described above. To fix cells, 100 .mu.L 2% v/v paraformaldehyde was added and cells incubated for 30 minutes at 4.degree. C., cells were pelleted by centrifugation at 300.times.g and the plates resuspended in 100 .mu.L FACS buffer. AlexaFluor 647 signal intensity (geometric mean) was measured by flow cytometry using a BD FACS Array instrument. Results are shown in FIGS. 3 and 4 and summarised in Table 2.
Equation 5 Percentage of receptor binding ( flow cytometry ) Based on geometric mean fluorescence % of specific binding = sample value - non - specific binding total binding - non - specific binding .times. 100 Total binding = biotinylated PD - 1 or CD 80 only ( no antibody ) Non - specific binding = no ligand streptavidin AlexaFluor 647 only ##EQU00003##
TABLE-US-00002 TABLE 2 Summary of lead antibody binding and neutralisation of PD-L1 binding to PD-1 or CD80 Receptor Neutralisation (mean of n = 3) PD-L1/PD-1 PD-L1/CD80 PD-L1/PD-1 neutralisation neutralisation neutralisation Human PD-L1 Cyno PD-L1 (FACS) (FACS) (ELISA) Clone ID (nM at 37.degree. C.) (nM at 37.degree. C.) IC.sub.50 (nM) IC.sub.50 (nM) IC.sub.50 (nM) 1D05 0.42 0.43 2.21 1.18 5.21 K.sub.on = 1.85 .mu.M K.sub.on = 1.89 .mu.M K.sub.off = 0.779 mM K.sub.off = 0.813 mM 84G09 0.43 0.52 1.82 1.60 7.90 K.sub.on = 2.43 .mu.M K.sub.on = 2.61 .mu.M K.sub.off = 1.05 mM K.sub.off = 1.35 mM Benchmark 0.25 4.79 1.85 1.42 14.1
[1529] d) PD-L1/PD-L2 Binding
[1530] PD-L1-Fc (R&D Systems) and PD-L2-Fc (R&D Systems) were diluted to 2 .mu.g/mL and separately adsorbed to 96-well, high protein binding plates (Greiner) overnight at 4.degree. C., 50 .mu.L/well. Excess protein was removed by washing with PBS-Tween (0.1% v/v) and the wells were blocked with 250 .mu.l/well Pierce Protein Free Blocking Buffer (Thermo, 37572) for 1 hour, after which plates were washed as described previously. Biotinylated anti-PD-L1 antibodies (in-house) or anti-PD-L2 control antibody (R&D Systems) were diluted in blocking buffer and three-fold serial dilutions performed from 10 .mu.g/mL. 100 .mu.L each antibody dilution was added to the plates in duplicate and incubated for 1 hour at room temperature, before washing as stated above. Antibody binding was detected using streptavidin labelled Europium (Perkin Elmer) diluted 1/1000 in DELFIA Assay buffer (Perkin Elmer). Plates were washed with TBS (Tris buffered saline)-Tween (0.1% v/v) and 50 .mu.L/well of DELFIA Enhancement solution (Perkin Elmer) was added to the plate. Time-resolved fluorescence was measured at 615 nm on an Envision plate reader (PerkinElmer). Results are shown in FIG. 5.
[1531] e) SPR Analysis
[1532] Label-free surface plasmon resonance (SPR) analysis was performed as per Example 4, except the assay was performed at 37.degree. C. Additionally, due to artefacts of running the assay at 37.degree. C., the best referencing of the binding sensorgrams was found to be using a sensorgrams from a negative control antibody using the same concentrations of human PD-L1. Results are shown in Table 2.
[1533] f) Mixed Lymphocyte Reaction
[1534] Expanded CD4.sup.+ T-cells were thawed and rested in AIM V.COPYRGT. medium (Gibco) at 37.degree. C., 5% CO.sub.2 overnight prior to the assay day. Serial dilutions of anti-human PD-L1 mAbs were prepared in the AIM medium at 4.times. final concentration. 50 .mu.L of diluted mAbs was added to 96-well, U-bottom plates. 1.times.10.sup.4 immature dendritic cells (iDC) in 50 .mu.L AIM medium and 1.times.10.sup.5 expanded CD4.sup.+ T-cells (expanded using Dynabeads Human T-Activator CD3/CD28 by Life Technologies (Invitrogen/Applied Biosystems; Cat No: 11131D), according to manufacturer's instructions) in 100 .mu.L AIM medium were added to the antibody dilutions in each well. Control wells include: CD4.sup.+ T-cells alone, iDC alone, CD4.sup.+ T-cell and iDC with or without IgG isotype control antibodies in 200 .mu.L AIM medium. Reaction plates were incubated for 5 days in a humidified incubator (37.degree. C. in 5% CO.sub.2). At the end of the assay, the plate was spun down (528.times.g for 3 minutes) and 100 .mu.L of supernatant was collected from the wells by gentle pipetting. Supernatants were analysed using human IFN.gamma. Quantikine ELISA kit (R&D Systems) according to manufacturer's instructions. Results are shown in FIG. 6.
[1535] g) Sequencing and Characterisation of Gene Segment Usage of 1D05 and 84G09
[1536] Antibodies were sequenced by Source Bioscience, and V-genes were compared to germline sequences.
TABLE-US-00003 TABLE 3 V region usage of lead antibodies Anti- non- Non- body CDRH3 germline CDRL3 germline clone V D J length CDRH3 V J length CDRL3 ID gene gene gene (aa) (aa) gene gene (aa) (aa) 1D05 IGHV3- IGHD3- IGHJ5 16 6 IGKV1D- IGKJ5*1 9 0 9*01 10*01 *02 39*01 84G09 IGHV3- IGHD3- IGHJ5 15 4 IGKV1D- IGKJ5*1 9 1 9*01 10*01 *02 39*01
[1537] h) Binding of Lead Antibodies to Natively Expressed PD-L1
[1538] 84G09 and 1D05 were labelled with AlexaFluor647 and used to stain dendritic cells derived from monocytic precursors. This shows that lead antibodies bind PD-L1 that is natively expressed on human dendritic cells. Data is shown in FIG. 7.
[1539] Materials and Methods
[1540] PBMC were suspended in RPMI 1640 medium without additives and allowed to adhere to a tissue culture flask for two hours at 37.degree. C. Non-adherent cells were removed and the flask washed three times with PBS. PBS was removed and replaced with RPMI 10% hiFBS (Gibco) containing 100 ng/mL GM-CSF and IL-4 (both Peprotech). Cells were cultured at 37.degree. C. for 7 days, and then removed from flask using a cell scraper.
[1541] Cells were resuspended in FACS buffer (PBS 1% w/v BSA 0.1% w/v sodium azide) and plated at 10.sup.5 cells/well, and incubated with Trustain FcX (Biolegend) for 10 min to prevent antibody binding to Fc.gamma.R. AlexaFluor647 labelled antibodies were added at a final concentration of 5 .mu.g/mL and incubated at 4.degree. C. for 1 hour. Cells were then washed three times in FACS buffer and fixed for 20 min in 4% paraformaldehyde (Affymetrix). After fixation, cells were washed three times as before and resuspended in FACS buffer for analysis by flow cytometry. Data was acquired using the MACSQuant flow cytometer (Miltenyi Biotec) and analysed in FlowJo v10.
Example 6--Antigen Preparation, Immunization Procedures, and Antigen-Specific B Cell Sorting and V-Region Recovery
[1542] Additional anti-human PD-L1 monoclonal antibodies were generated using the KyMouse.TM. system previously described. Genetically engineered HK mice were immunized with soluble recombinant human and mouse PD-L1 or surface expressed human and mouse PD-L1 displayed on mouse embryonic fibroblast (MEF) cells. Serum titres were performed by reverse ELISA and mice with the highest titres were selected for processing. At the end of each regime, spleen and lymph nodes were removed. Tissues were prepared into a single cell suspension and stained for sorting antigen-specific B-cells by FACS.
[1543] Materials and Methods
[1544] a) Immunisation of Mice
[1545] Mice were immunised with soluble recombinant human PD-L1 or a combination of human and mouse PD-L1 protein (in-house) as per the schedule described in Example 1 for KM032 (hereafter described as KM121). Mice were also immunised with human PD-L1 protein, and MEF cells expressing human or mouse PD-L1, as per the schedule described in Example 1 for KM033 (hereafter described as KM122). MEF cells expressing mouse PD-L1 were generated as per Example 1, but substituting mouse PD-L1 sequences for the human PD-L1 sequences, and substituting anti-mouse PD-L1 detection antibody (eBioscience) for the anti-human PD-L1 detection antibody.
[1546] b) Determining Serum Titre by Reverse PD-L1 ELISA Protocol
[1547] Titres in mouse serum samples were determined using a reverse PD-L1 ELISA protocol as per Example 1, with the following changes. In-house generated hPD-L1-his was labelled in-house using Lightning Link kit (Innova Biosciences), and used at 1 .mu.g/mL in reagent diluent; 50 .mu.L/well). Bound hPD-L1 was detected by addition of streptavidin-Europium (Perkin Elmer) diluted 1/1000 in DELFIA assay buffer (Perkin Elmer). Following incubation for 1 hour at room temperature in the dark, plates were washed using TBS (Tris buffered saline)-Tween (0.1% v/v) and 50 .mu.L/well of DELFIA Enhancement solution (Perkin Elmer) was added to the plate. Time-resolved fluorescence was measured at 615 nm on an Envision plate reader (PerkinElmer). Fluorescence data was plotted as Europium counts.
[1548] c) Sorting of Antigen-Specific B Cells and Retrieval of V-Regions
[1549] The methods used were substantially as described in Example 1 of PCT application WO2015/040401, which is incorporated herein by reference. In brief, splenocytes and lymph node cells isolated from KM121 and KM122 immunisation regimes were stained with an antibody cocktail containing markers for the selection of cells of interest (CD19), whereas unwanted cells were excluded from the final sorted population (IgM, IgD, 7AAD). CD19.sup.+ B-cells were further labelled with human PD-L1 (Seq ID No: i) and mouse PD-L1 (Seq ID No:325, labelled with AlexaFluor647 and AlexaFluor488, respectively, in-house using Lightning Link kits) to detect B-cells producing specific antibodies--cells binding human PD-L1, or both human and mouse PD-L1 were selected. These cells were single cell sorted by FACS into lysis buffer. V-region sequences were recovered using RT-PCR and two further rounds of PCR, then bridged to mouse IgG1 constant region and expressed in HEK293 cells. Supernatants from HEK293 cells were screened for the presence of PD-L1 binding antibodies. This method is hereafter referred to as BCT.
Example 7--Supernatant Screening
[1550] BCT supernatants were screened by HTRF, and selected primary hits further screened for binding to cell-expressed recombinant hPD-L1 and neutralisation of PD-1 binding, and for affinity of binding to human, cynomolgus and mouse PD-L1 recombinant protein by SPR, as described in this Example. KM121 antibodies, with an affinity of 1 nM or better for human and in some cases also cynomolgus PD-L1 were taken forward for further characterisation. For KM122, antibodies with the capacity to neutralise PD-1 binding to cell-expressed PD-L1 were taken forward, along with high affinity (<1 nM) binding to both human and cynomolgus PD-L1. Antibodies did not bind to mouse PD-L1.
[1551] a) Primary Screen--Binding to Recombinant Human PD-L1 (BCT Supernatants)
[1552] Supernatants collected from BCT expression were screened for the ability of secreted antibodies to bind to hPD-L1 expressed as a recombinant protein (produced in-house). Binding of secreted antibodies to recombinant human and mouse. PD-L1 were identified by HTRF.RTM. (Homogeneous Time-Resolved Fluorescence, Cisbio) assay format using FluoProbes.RTM.647H (Innova Biosciences) labelled PD-L1 (referred to herein as 647 hPD-L1 or 647 mPD-L1 for human PD-L1 and mouse PD-L1 labelled with FluoProbes.RTM.647H respectively). 5 .mu.L BCT supernatant was transferred to a white 384-well, low-volume, non-binding surface polystyrene plate (Greiner). 5 .mu.L of 25 nM 647 hPD-L1 or 647 mPD-L1 diluted in HTRF assay buffer was added to all wells. Reference antibody was diluted in BCT media (Gibco #A14351-01) to 40 nM and 5 .mu.L added to plate. For negative control wells, 5 .mu.L of mouse IgG1 (Sigma M9269 in some instances referred to as CM7) diluted to 40 nM in BCT media was added. Binding of secreted antibodies to PD-L1 was detected by addition of 10 .mu.L of goat anti-mouse IgG (Southern Biotech) directly labelled with Europium cryptate (Cisbio) diluted 1/2000 in HTRF assay buffer. The plate was left to incubate in the dark for 2 hours prior to reading time resolved fluorescence at 620 nm and 665 nm emission wavelengths using an EnVision plate reader (Perkin Elmer).
[1553] Data were analysed by calculating 665/620 ratio and percent effect for each sample according to Equation 2 and Equation 1 respectively.
[1554] For KM121, primary hits were selected based on greater than or equal to 30 percent effect whereas for KM122 primary hits were selected based on greater than or equal to 40 percent effect.
[1555] Progression to secondary screen was based on data from recombinant PD-L1 binding.
[1556] b) Secondary Screen--Binding to Cell Expressed hPD-L1 and Neutralisation of hPD-L1 Binding to PD-1 (BCT Supernatants)
[1557] Binding of BCT supernatants were tested for ability to bind to CHO-S cells expressing hPD-L1. CHO-S cells expressing hPD-L1 (generated in-house), were diluted in FACS buffer (PBS 1% BSA 0.1% sodium azide) and were distributed to a 96-well, V-bottom plate (Greiner) at a density of 0.5-1.times.10.sup.5 cells per well. Cells were washed with 150 .mu.L PBS and centrifuged at 300 g for 3 minutes. Supernatant was aspirated and 150 .mu.L PBS added. This wash step was repeated.
[1558] 25 .mu.L BCT neat supernatant, reference antibody or control antibody diluted to 300 nM in BCT media was added to the washed cells. 25 .mu.L of 30 nM biotinylated human PD-1 (in-house) was added and cells were incubated at 4.degree. C. for 60 minutes. 150 .mu.L FACS buffer was added and cells washed as described above. To detect biotinylated PD-1 and anti-PD-L1 antibody binding, Streptavidin-647 (Jackson ImmunoResearch) and anti-Mouse PE (Jackson ImmunoResearch) were each diluted 1/500 in FACS buffer and 50 .mu.L of this mixture added to cells. Cells were incubated 4.degree. C. for 60 minutes. Cells were washed twice with 150 .mu.L FACS buffer, centrifuging at 300 g for 3 minutes after each wash step and aspirating supernatant. Cells were fixed by addition of 50 .mu.L 4% paraformaldehyde overnight. Cells were washed once as above and resuspended in FACS buffer for analysis. PE and APC signal intensity (geometric mean) was measured by flow cytometry using a BD FACS Array instrument. Data was plotted as geometric mean values without further calculation.
[1559] For KM121, secondary hits were selected based on high affinity (<1 nM) binding to human PD-L1. For KM122, secondary hits were selected based on comparable high affinity (<1 nM) binding human and cynomolgus PD-L1 and ability to neutralise PD-1 binding to cell-expressed PD-L1. Results are summarised in Table 4.
TABLE-US-00004 TABLE 4 Summary of BCT clone screening Number of Number of Number of BCT Primary hits secondary Experiment supernatants cherry hits ID screened picked confirmed KM121 984 162 7* KM122 1312 263 45** *three of these secondary hits were not included in the primary screen and were screened by SPR and neutralisation only **one hit was identified by primary screen but insufficient material was available for secondary screen. After re-expression, clone was shown to bind human and cynomolgus PD-L1 with affinity of <1 nM and carried forward
[1560] c) Analysis of Binding by Surface Plasmon Resonance
[1561] SPR analysis was carried out on the ProteOn XPR36 Array system. Anti-mouse IgG (GE Healthcare BR-1008-38) was immobilised on a GLM chip by primary amine coupling. Antibodies were directly captured from BCT supernatants. Human, mouse and cynomolgus PD-L1 were used as analytes and passed over the captured antibodies at a single concentration. The binding sensorgrams are double referenced with a 0 nM (ie buffer alone) injection, and the data is analysed using the 1:1 model inherent to the ProteOn analysis software. The assay is carried out at 25.degree. C. and used HBS-EP as running buffer.
Example 8--Characterisation of Selected Antibodies
[1562] Selected hits were re-expressed with a human IgG1 constant region and sent for sequencing at Source Bioscience. V region usage is listed in Table 5. Hits were then analysed in an ELISA to determine their ability to neutralise PD-L1/PD-1 interactions, and PD-L1/CD80 interactions. All seven KM121 hits neutralised PD-L1/CD80 interactions; however, four antibodies did not neutralise PD-L1/PD-1. Four out of five KM122 hits neutralised both PD-L1/PD-1 and PD-L1/CD80 internations. Results are shown in FIGS. 8 and 9. Antibodies shown to neutralise both PD-1 and CD80 interactions with PD-L1 were further screened for their ability to increase IFN.gamma. in an autologous monocyte-T-cell co-culture assay.
[1563] Materials and Methods
[1564] a) PD-L1/PD-1 and PD-L1/CD80 Neutralisation ELISA
[1565] CD80 (R&D Systems) or PD-1 (in-house) diluted to 2.5 .mu.g/mL were adsorbed to 96-well, low auto-fluorescent, high protein binding plates (Costar) overnight at 4.degree. C. Excess protein was removed by washing with PBS-Tween (0.1% v/v) and the wells were blocked with 1% w/v bovine serum albumin (BSA, Sigma) in PBS for 1 hour at room temperature, after which plates were washed with PBS-Tween as above. 60 .mu.L of a titration (three-fold serial dilution) of antibody was added to a 96-well, non-binding plate diluted in ELISA assay buffer (PBS+0.1% BSA). 60 .mu.L of biotin labelled PD-L1 at 16 nM working concentration (8 nM FAC) was added to the plate excluding control wells where 60 .mu.L ELISA assay buffer was added. The plate was incubated for 30 minutes before transferring 50 .mu.L to the coated plates. The coated, plates were incubated for 1 hour at room temperature. Excess protein was removed by washing with PBS-Tween (0.1% v/v). PD-L1 binding was detected using streptavidin labelled europium (Perkin Elmer) diluted 1/1000 in DELFIA assay buffer (Perkin Elmer). The plates were washed with TBS (Tris buffered saline)-Tween (0.1% v/v) and 50 .mu.L/well of DELFIA Enhancement solution (Perkin Elmer) was added to the plate. Time-resolved fluorescence was measured at 615 nm on an Envision plate reader (PerkinElmer). Percentage specific binding was calculated as defined in Equation 3. IC.sub.50 values were determined using GraphPad Prism software by curve fitting using a four-parameter logistic equation (Equation 4). Results are shown in Table 4a below. Values for KM121 antibodies are a mean of three independent experiments. Values for KM122 are from a single experiment. ND indicates IC.sub.50 value not determined, as a complete curve could not be generated.
TABLE-US-00005 TABLE 4a IC.sub.50 values for neutralisation of PD- L1 interactions with PD-1 and CD80 Antibody PD-1/PD-L1 CD80/PD-L1 Regime clone ID IC.sub.50 (nM) IC.sub.50 (nM) KM121 411B08 2.22 1.60 KM121 411C04 2.45 1.93 KM121 411D07 ND 9.06 KM121 385F01 2.25 1.76 KM121 386H03 ND 0.74 KM121 389A03 ND 13.18 KM122 416E01 1.72 0.98 KM122 413G05 2.02 1.10 KM122 414B06 1.84 1.00 KM122 413F09 ND ND KM122 413D08 1.20 0.67
[1566] Selected lead antibodies active in the monocyte-T-cell co-culture assay (see Example 9) were analysed by SPR at 25 and 37.degree. C. Lead antibodies retained sub-nanomolar affinity binding to PD-L1 even at 37..degree. C. Antibodies did not bind mouse PD-L1. Results are shown in Table 4b.
[1567] Materials and Methods
[1568] SPR analysis was performed as per Example 4 with the following amendments: analysis was performed at 37.degree. C. as well as 25.degree. C. to increase the stringency of the assay. Human, cynomolgus and mouse PD-L1 (his-tagged) were generated in house (Seq ID Nos 3, 5 and 326, respectively).
TABLE-US-00006 TABLE 4b Binding affinities of selected lead antibodies Clone ID Temperature Antigen KD (nM) 413G05 25.degree. C. Human 0.024 Kon = 2.57 .mu.M Koff = 62.3 .mu.M 414B06 25.degree. C. Human 0.172 Kon = 4.09 .mu.M Koff = 0.702 mM 416E01 25.degree. C. Human 0.193 Kon = 2.34 .mu.M Koff = 45.1 mM 413G05 25.degree. C. Cyno 0.015 Kon = 2.66 .mu.M Koff = 38.9 mM 414B06 25.degree. C. Cyno 0.192 Kon = 3.78 .mu.M Koff = 0.726 mM 416E01 25.degree. C. Cyno 0.411 Kon = 2.44 .mu.M Koff = 1.0 mM 413G05 37.degree. C. Human 0.050 Kon = 4.67 .mu.M Koff = 0.235 mM 414B06 37.degree. C. Human 0.778 Kon = 5.88 .mu.M Koff = 4.57 mM 416E01 37.degree. C. Human 0.511 Kon = 4.34 .mu.M Koff = 2.22 mM 413G05 37.degree. C. Cyno 0.046 Kon = 4.31 .mu.M Koff = 0.197 mM 414B06 37.degree. C. Cyno 0.794 Kon = 5.02 .mu.M Koff = 3.98 mM 416E01 37.degree. C. Cyno 0.998 Kon = 4.03 .mu.M Koff = 4.02 mM
Example 9--Testing of Lead Anti-PD-L1 Antibodies in an Autologous Co-Culture Assay
[1569] The effects of anti-PD-L1 antibodies on IFN.gamma. production are analysed in a co-culture of purified peripheral blood monocytes and CD45RO.sup.+ memory T-cells from the same donor. In brief, monocytes are isolated by negative selection using magnetic separation beads (Miltenyi Biotec). CD45RO.sup.+ T-cells are isolated by a first round of negative selection for CD3.sup.+ T-cells, and one round of positive selection for CD45RO.sup.+ cells (Miltenyi Biotec). Cell subsets are co-cultured at a 1:1 ratio in RPMI 10% hiFBS in the presence of anti-CD3 (UCHT1, eBioscience) to provide TCR stimulation, and antibodies under investigation. Supernatants are taken after 4 days for analysis of IFN.gamma. by MSD (Meso Scale Discovery).
[1570] The experiments were performed as described, except IFN.gamma. production was measured with the R&D systems.TM. Human IFN.gamma. Duoset.RTM. ELISA, using DELFIA.RTM. Eu-N1 Streptavidin detection.
[1571] Response for IFN.gamma. standard (pg/mL) was plotted versus relative fluorescence response at 615 nM. IFN.gamma. concentration was interpolated from standard curve in pg/mL using a 4-parameter logistic fit as defined by Equation 4. Antibody-induced IFN.gamma. is represented as fold induction compared to assay signal of wells showing background levels of response as defined in Equation 6. Each plot represents mean fold induction for individual donors with at least, 2 different donors represented versus antibody concentration Log (M). Results are shown in FIGS. 22 and 37.
Fold induction=assay response (pg/mL)/background response (pg/mL)
Background IFN.gamma. response=IFN.gamma. concentration (pg/mL) from wells containing monocyte-T-cell co-culture with anti-CD3 stimulation, without antibody. Equation 6
[1572] All five antibodies, in human IgG1 format, induced a specific, dose-dependent increase in IFN.gamma. production by T-cells after 4 days of co-culture with autologous monocytes and anti-CD3 (see FIGS. 22a and 22b). The two antibodies that induced the highest increase in cytokine production, 413G05 and 414E106, were selected for repeat characterisation by SPR (see Example 8). Antibody 416E01, in human IgG4(PE) format (Seq ID No:199), also induced specific dose-dependent increase in IFN.gamma. production in the co-culture assay. This antibody was also selected for repeat SPR analysis.
[1573] The three selected antibodies were also analysed alongside the two lead antibodies selected in Example 4 (1D05 and 84G09), and a commercial effector enabled benchmark antibody. Antibodies were formatted as human IgG1. All antibodies induced dose-dependent IFN.gamma. production in this assay (FIG. 37 and Table 22).
TABLE-US-00007 TABLE 5 V Gene usage for antibody leads Non- Non- CDHR3 germline CDRL3 germline Antibody length CDRH3 length CDRL3 Regime clone ID V gene D gene J gene (aa) (aa) V gene J gene (aa) (aa) KM121 411B08 IGHV3-7*01 IGHD4-11*01 IGHJ4*02 12 7 IGKV1D-12*02 IGKJ3*01 9 1 KM121 411C04 IGHV3-7*01 IGHD4-11*01 IGHJ4*02 12 6 IGKV1D-12*02 IGKJ3*01 9 1 KM121 411D07 IGHV4-4*02 IGHD3-10*01 IGHJ4*02 8 1 IGKV4-1*01 IGKJ2*04 8 2 KM121 386H03 IGHV4-4*02 IGHD3-10*01 IGHJ4*02 8 2 IGKV4-1*01 IGKJ2*04 8 1 KM121 389A03 IGHV4-39*01 IGHD6-13*01 IGHJ1*01 13 6 IGKV4-1*01 IGKJ1*01 9 1 KM121 385F01 IGHV3-7*01 IGHD4-11*01 IGHJ4*02 12 7 IGKV1D-12*02 IGKJ3*01 9 1 KM122 413D08 IGHV3-33*01 IGHD5-18*01 IGHJ6*02 11 3 IGKV1-17*0 IGKJ1*01 9 1 KM122 413G05 IGHV3-11*01 IGHD1-20*01 IGHJ6*02 16 5 IGKV1D-12*02 IGKJ4*01 9 1 KM122 413F09 IGHV3-23*04 IGHD5-18*01 IGHJ4*02 16 8 IGKV1-9*d0 IGKJ5*01 9 3 KM122 414B06 IGHV3-7*01 IGHD5-24*01 IGHJ4*02 12 6 IGKV1D-12*02 IGKJ3*01 9 0 KM122 416E01 IGHV3-23*04 IGHD6-13*01 IGHJ4*02 14 10 IGKV1D-12*02 IGKJ5*01 9 2
TABLE-US-00008 TABLE 22 Summary of data from monocyte-T cell co-culture experiments. Antibody mean EC50 mean fold increase name (nM) IFN.gamma. 1D05 0.21 3.04 84G09 0.081 3.60 413G05 0.082 2.85 414B06 0.012 3.33 416E01 0.064 2.23 benchmark 2 0.057 2.30
Example 10--Bispecific FIT-Ig Molecules Targeting PD-L1 and TIGIT
[1574] Bispecific FIT-Ig constructs were constructed substantially as described in Example 1 of International Application WO2015/103072 (in the name of EpiMab Biotherapeutics, and is incorporated herein by reference).
[1575] The bispecific constructs, having a FIT-Ig structure, as described in FIG. 1 of WO2015/103072 were expressed in CHO cells with a vector ratio of: Construct 1 DNA: 50%, Construct 2: DNA 25%: Construct 3 DNA 25% of total DNA in the transient transfection. The bispecific molecules were purified by standard Protein A and size exclusion chromatography. In this regard, Construct 1 is the polypeptide chain made up of VL.sub.A-CL-VH.sub.B-CH1-CH2-CH3 in FIG. 1 of WO2015/103072. Construct 2 is the polypeptide chain made up of VH.sub.A-CH1 in FIG. 1 of WO2015/103072, and Construct 3 is the polypeptide chain made up of VL.sub.B-CL in FIG. 1 of WO2015/103072.
[1576] SPR analysis was used to determine affinities of the various arms of the bispecific and the parental monospecific antibodies were used to determine if the affinities had been altered in the bispecific molecule. Sequential binding of antigens were used to test whether the bispecific constructs were capable of binding on both arms of the bispecific.
TABLE-US-00009 TABLE 6 Bispecific antibody constructs and control monospecific antibodies Native variable Additional Full name Alias domain.sup.1 Domain.sup.2 1D05/in-house anti-TIGIT Bispecific 1 1D05 (anti-PD-L1)* Kymab TIGIT In-house anti-TIGIT/1D05 Bispecific 2 Kymab TIGIT 1D05 (anti-PD-L1)* Tool anti-TIGIT/Tool anti-PD-L1 Bispecific 3 Tool anti-TIGIT Tool anti-PD-L1 Tool anti-PD-L1/Tool anti-TIGIT Bispecific 4 Tool anti-PD-L1 Tool anti-TIGIT 1D05 Antibody 1 In-house anti-PD- na L1* Kymab TIGIT Antibody 2 In-house anti- na TIGIT Tool PD-L1 Antibody 3 Tool anti-PD-L1 na Tool TIGIT Antibody 4 Tool anti-TIGIT na *1D05 has the V.sub.H sequence of Seq ID No: 33 and the V.sub.L sequence of Seq ID No: 43, and a heavy chain constant region of Seq ID No: 205 .sup.1"Native Variable domain" corresponds to the antigen-binding site formed by VH.sub.B and VL.sub.B in FIG. 1 of WO2015/103072 .sup.2"Additional domain" corresponds to the antigen binding site formed by VHA and VLA in FIG. 1 of WO2015/103072
[1577] a) Kinetic Analysis
[1578] An anti-human IgG capture surface was created by a mix of 3 anti-human Fc antibodies (Jackson Labs 109-005-008, 109-006-008 and 309-006-008) immobilised on a GLC chip by primary amine coupling. Control monospecific antibodies or Bispecific antibody constructs were captured on this surface and human PD-L1 or TIGIT was used as analyte at 512 nM, 128 nM, 32 nM, 8 nM and 2 nM with 0 nM (i.e. buffer alone) used to double reference the binding sensorgrams. The assay was run at 25.degree. C., using HBS-EP as running buffer. The sensorgrams were fitted to the 1:1 model inherent to the ProteOn analysis software.
TABLE-US-00010 TABLE 7 TIGIT Binding KD Full name Alias Ka Kd (nM) 1D05/in-house anti-TIGIT Bispecific 1 2.38E+06 2.65E-03 1.11 In-house anti-TIGIT/1D05 Bispecific 2 1.12E+06 2.02E-03 1.8 Tool anti-TIGIT/Tool Bispecific 3 2.10E+06 3.69E-03 1.75 anti-PD-L1 Tool anti-PD-L1/Tool Bispecific 4 3.22E+06 2.98E-03 0.93 anti-TIGIT 1D05 Antibody 1 nbs nbs nbs Kymab TIGIT Antibody 2 1.58E+06 2.27E-03 1.44 Tool PD-L1 Antibody 3 nbs nbs nbs Tool TIGIT Antibody 4 3.16E+06 5.42E-03 1.72
TABLE-US-00011 TABLE 8 PD-L1 Binding KD Full name Alias Ka Kd (nM) lD05/in-house anti-TIGIT Bispecific 1 6.03E+05 1.61E-04 0.27 In-house anti-TIGIT/1D05 Bispecific 2 1.04E+06 2.14E-04 0.21 Tool anti-TIGIT/Tool Bispecific 3 1.25E+06 1.22E-04 0.1 anti-PD-L1 Tool anti-PD-L1/Tool Bispecific 4 7.36E+05 1.57E-04 0.21 anti-TIGIT 1D05 Antibody 1 9.71E+05 3.36E-04 0.35 Kymab TIGIT Antibody 2 nbs nbs nbs Tool PD-L1 Antibody 3 1.05E+06 2.08E-04 0.2 Tool TIGIT Antibody 4 nbs nbs nbs
[1579] b) Bispecific Binding
[1580] Using the same anti-human IgG capture surface created for kinetic analysis, the bispecific antibody constructs were captured on this surface and recombinant PD-L1 or TIGIT was used as analyte at 512 nM, 128 nM, 32 nM, 8 nM and 2 nM with 0 nM (i.e. buffer alone) used to double reference the binding sensorgrams. The assay was carried out by injecting PD-L1 followed by TIGIT with no regeneration between analyte injections, and also with TIGIT followed by PD-L1. The sensorgrams for the double referenced 512 nM are shown in FIGS. 10 and 11.
[1581] c) Characterisation of Bispecific FIT-Ig Molecules Binding to PD-L1 and TIGIT by AlphaScreen.RTM.
[1582] An AlphaScreen.RTM. binding assay was developed to assess the bispecific binding of PD-L1/TIGIT FIT-Ig molecules. The assay was set up using biotinylated His-PD-L1 (SEQ ID No:3) and His-FLAG-TIGIT (SEQ ID No:539) detected respectively with streptavidin donor beads and anti-FLAG acceptor beads (both Perkin Elmer, 6760613). Human IgG1 (Sigma 15154) and parental monospecific antibodies alone or in combination were used as negative controls, while an anti-His antibody (Qiagen 34660) was used as positive control.
[1583] Two protocols were created to investigate the ability of FIT-Ig molecules to promote proximity of TIGIT and PD-L1 coated beads with a distinct stringency. Antibodies were either incubated with PD-L1 and TIGIT proteins before adding the AlphaScreen.RTM. detection beads (Method one), or incubated with the detection beads pre-coated with their respective TIGIT and PD-L1 proteins (Method two). Method two was designed to mimic the cell recruitment by bispecific antibodies.
[1584] i) Method One
[1585] Bispecific antibodies, parental monospecific antibodies and control antibodies were prepared in buffer (PBS pH 7.4 (Gibco) and 0.1% w/v BSA (Sigma)) at 150 nM and diluted as per 1:3 series, 8 points. 5 .mu.L of each serial dilution of antibody were mixed in a 384-well AlphaLISA.RTM. assay plate (Perkin Elmer 6005350) to 5 .mu.L of biotinylated His-PD-L1 and 5 .mu.L of His-FLAG-TIGIT at 50 nM in buffer. Parental monospecific antibodies were also prepared as described above starting from 300 nM to be tested in combination. 2.5 .mu.L of the first antibody was added to the same volume of the second antibody, then 5 .mu.L of each combination of parental monospecific antibodies were mixed in assay plates to 5 .mu.L of biotinylated His-PD-L1 and 5 .mu.L of His-FLAG-TIGIT at 50 nM in buffer. Assay plates were incubated for 1 hour at room temperature before adding 5 .mu.L of anti-FLAG acceptor beads at 0.1 g/L for an additional hour at room temperature in the dark. Finally, 5 .mu.L of streptavidin donor beads at 0.1 g/L were added to assay plates for 2 hours and 30 minutes. Assay plates were read using an EnVision plate reader (Perkin Elmer) with excitation/emission wavelengths of 680/615 nm. The fluorescent counts measured (Alpha signal) were plotted in Prism against antibody titrations. Results are shown in FIG. 25. Binding of FIT-Ig molecules to PD-L1 and TIGIT increases with the concentration of antibody up to 10 nM. No binding is observed for the monospecific parental antibodies and the isotype control.
[1586] ii) Method Two
[1587] Streptavidin donor beads prepared at 0.05 g/L in buffer (PBS pH 7.4 (Gibco 14190169) and 0.1% w/v BSA (Sigma)) were coated with biotinylated His-PD-L1 (Seq ID No:3) at 25 nM, while His-FLAG-TIGIT (Seq ID No:539) at 25 nM was used to label anti-FLAG acceptor beads at 0.05 g/L in buffer. Both acceptor and donor beads were incubated for 1 hour at room temperature in the dark.
[1588] Bispecific antibodies, parental monospecific antibodies, alone and combined, and control antibodies were prepared in buffer at 300 nM and diluted as per 1:3 series, 8 points. 5 .mu.L of each serial dilution of antibody were mixed in a 384-well AlphaLISA.RTM. assay plate (Perkin Elmer 6005350) to 10 .mu.L of pre-coated donor beads and 10 .mu.L of pre-coated acceptor beads. Assay plates were incubated at room temperature for 4 hours in the dark and then read as described for method one. The fluorescent counts measured (Alpha signal) were plotted in Prism against antibody titrations. Results are shown in FIG. 26. Binding of FIT-Ig molecules to PD-L1 and TIGIT increases with the concentration of antibody up to 20 nM. No binding is observed for the monospecific parental antibodies and the isotype control.
[1589] d) Characterisation of Bispecific FIT-Ig Molecules Binding to PD-L1 and TIGIT by Flow Cytometry
[1590] A flow cytometry protocol was developed to assess the ability of the FIT-Ig molecules to promote the recruitment of cells expressing TIGIT and PD-L1. For this purpose, CHO cells transfected with human PD-L1 were stained with Celllrace.TM. Far Red (Invitrogen C34572) which emits maximally at 661 nm while HEK cells transfected with human TIGIT were stained with CellTrace.TM. Violet (Invitrogen C34571) which emits maximally at 450 nm.
[1591] CHO human PD-L1 and HEK human TIGIT cells were harvested, counted, washed, and re-suspended in PBS (Gibco 14190169) at 1 million of cells per mL. CellTrace.TM. Far Red and CellTrace.TM. Violet dyes were diluted 1:2000 and incubated with cells for 20 min at 37.degree. C. in the dark, according to manufacturer's recommendations. Buffer (PBS (Gibco 14190169), 1% BSA (Sigma) 0.1% Na azide (Severn Biotech 40-2010-01)) was then added in excess for an additional 5-minute incubation step. Cells were spun down, re-suspended in buffer at 0.5 million of cells per mL and incubated for at least 10 minutes at 37.degree. C. before proceeding with binding protocol. Unstained cells were kept and used to set up the gating strategy.
[1592] Bispecific antibodies and human IgG1 were prepared in buffer at 150 nM and diluted as per 1:3 series, 8 points. 50 .mu.L of each serial dilution of antibody, 50 .mu.L of CHO human PD-L1 cells labelled with CellTrace.TM. Far Red and 50 .mu.L of HEK human TIGIT labelled with Celllrace.TM. Violet were added to a 96-well, V-bottom PS plate (Greiner 651901). Assay plates were incubated at room temperature for 1 hour under gentle agitation (450 rpm) before being read using the Attune N.times.T flow cytometer (Thermo Fisher). CellTrace.TM. Violet was excited using the Violet laser and detected in the VL1 channel with a 440/50 bandpass filter. CellTrace.TM. Far Red was excited using the Red laser and detected in the RL1 channel with a 670/14 bandpass filter. Sample collection was performed without vortexing samples. FCS files were analysed with FlowJo.RTM. software. Single cells and duplets were gated based on the forward and side scatter dot plot.
[1593] Data analysis resulted in the identification of four different gates: a double negative quadrant corresponding to unstained CHO human PD-L1 and unstained HEK human TIGIT; two quadrants positive for single staining (in VL1 or RL1 channel); and a quadrant positive for dual staining (in both VL1 and RL1 channels) composed of stained CHO human PD-L1 and stained HEK human TIGIT recruited by FIT-Ig molecules. Percentages of double positive cells were plotted into Prism against antibody titrations. Results are shown in FIG. 27. Percentage of double positive cells increases with the concentration of FIT-Ig molecules up to 1 nM.
[1594] The monospecific binding of test molecules to target was confirmed on stained cells using monospecific antibodies labelled with R-Phycoerythrin (PE) which emits maximally at 590 nm. PE-labelled Antibody 1, Antibody 2 and human IgG1 were diluted in buffer at 150 nM. 50 .mu.L of each antibody were mixed with 50 .mu.L of stained CHO human PD-L1 and 50 .mu.L of stained HEK human TIGIT in a 96-well, V-bottom PS plate (Greiner 651901). Following a 1 hour incubation at room temperature, cells were washed 3 times with 200 .mu.L/well of PBS and re-suspended in 150 .mu.L/well of buffer. Assay plates were read using the Attune N.times.T flow cytometer (Thermo Fisher) to record fluorescence. Cell Trace.TM. Violet and Far Red were detected as stated above. PE was excited using the Yellow laser and detected in the YL1 channel with a 585/16 bandpass filter. GeoMean values in the YL1 channel were used to determine monospecific binding to stained CHO human PD-L1 or stained HEK human TIGIT.
Example 11--Generation and Expression of Anti-PD-L1-IL-2 Immunocytokine Constructs
[1595] Immunocytokines were generated by fusing wild type IL-2 (SEQ ID No:301), or IL-2 containing deletions in the first nine amino acids (see SEQ ID Nos:303 to 323, fused to Seq ID No:324), to the light chain of anti-PD-L1 antibody 1D05 (see Seq ID No:45). These were paired with an IgG1 effector-disabled variant of 1D05 heavy chain (Seq ID No:205). Wild type IL-2 fused to the heavy chain of 1D05 was generated for use as a control (SEQ ID No:302) and paired with the unmodified light chain of 1D05 (Seq ID No:45). Twenty-two immunocytokines were successfully expressed and characterised further. One light chain construct, 1D05 D1 did not express successfully.
[1596] Materials and Methods
[1597] The DNA sequences encoding the anti-PD-L1 (antibody 1D05) immunocytokine (C-terminal IL-2 fusion to light chain) were purchased as synthetic DNA strings and cloned into the pTT5 expression vector using the Golden Gate cloning strategy. The heavy chain sequence of 1D05, includes a constant region which is a disabled IgG1 variant with changes from wild-type shown in bold (Seq ID No:299). The light chain of antibody 1D05 has full length wild type IL-2 sequence (underlined) fused to the C-terminus of the Kappa constant region (Seq ID No:300). Overlap PCR using appropriate oligonucleotide primers were used to generate variants of N-terminal of IL-2 (see Seq ID No:300 where IL-2 the sequence is underlined and the region to be varied is shown in bold). Variant sequences were cloned into the pTT5 expression vector using the Golden Gate method. The wild type and variant constructs were transfected to Expi293.TM. cells for expression.
Example 12--Generation of IL2R Transfectant Cells for Screening
[1598] In order to differentiate between immunocytokine activity on the high affinity (.alpha..beta..gamma.) and intermediate affinity (.beta..gamma.) IL-2 receptors, IL-2R transfectants were generated. TF-1 cells, expressing endogenous common .gamma. chain, were transfected with .beta., or .alpha. and .beta. receptor subunits, to impart responsiveness to IL-2. The proliferative response to immunocytokines was then analysed using these cells (see Example 13).
[1599] Materials and Methods
[1600] Two recombinant cell lines were generated to distinguish between signalling through high affinity (.alpha..beta..gamma.) and intermediate affinity (.beta..gamma.) IL-2R. The erythroleukemia cell line TF-1 (European Collection of Authenticated Cell Cultures) shows complete growth dependency on granulocyte-macrophage colony-stimulating factor (GM-CSF) or interleukin-3 (IL-3). The first cell line generated was transfected with full length human IL-2R.beta. (CD122) only. The second cell line was generated by transfecting the full length human IL-2R.alpha. (CD25) into the first cell line.
[1601] The transfected sequences were codon optimized for mammalian expression and cloned into an expression vector under the CMV promoter flanked by 3' and 5' piggyBac specific terminal repeat sequences facilitating stable integration into the cell genome (see: "A hyperactive piggyBac transposase for mammalian applications"; Yusa K., et al., Proc. Natl. Acad. Sci. USA., 108(4): 1531-6, 2011 Jan. 25). Furthermore, the expression vector for each subunit contained a different selection cassette to facilitate stable cell line generation. The .beta. subunit was selected using puromycin (Sigma) and the .alpha. subunit using geneticin (Gibco). The a subunit was transfected into cells already expressing the .beta. subunit.
[1602] The expression plasmids were co-transfected with a plasmid encoding piggyBac transposase into the TF1 cell line by electroporation using the Lonza 4-D nucleofector transfection X kit system according to manufacturer instructions. 24 hours after transfection, complete media was supplemented with the appropriate selection and cells grown for at least 3 weeks to select a stable line, with media being exchanged every 3 to 4 days. The expression of the recombinant human subunits was assessed by flow cytometry using anti-human CD122 (IL-2R.beta.) APC conjugated antibody (eBioscience) and anti-human CD25 (IL-2R.alpha.) PE conjugated antibody (eBioscience). Endogenous common .gamma. chain expression was confirmed with anti-human CD132 (common .gamma. chain) PE conjugated antibody (eBioscience). As expression was low, CD122.sup.+ cells were sorted by fluorescence activated cell sorting (FACS) and further cultured under selection. There was uniform expression of a chain after transfection, and therefore these cells were not sorted.
[1603] Complete TF1 media was made up of RPMI medium 1640 (Gibco) plus GM-CSF (2 ng/mL) and supplemented with 10% v/v heat inactivated fetal bovine serum (hiFBS, Gibco). Once responsiveness to IL-2 was confirmed, transfected cell lines were maintained in RPMI 1640, 10% hiFBS and 5 ng/mL recombinant human IL-2 with (.alpha..beta.) or without (.beta.) geneticin.
Example 13--Assessing Ability of Immunocytokine Constructs to Signal Through IL-2R
[1604] Immunocytokines were assessed for their ability to induce proliferation of TF1 cell lines transfected with the .beta. subunit, or with both the .alpha. and .beta. subunits of IL-2R. Cells were starved of cytokines overnight, then stimulated with titrations of each immunocytokine. CellTiter-Glo.RTM. was used to determine the number of viable cells in culture after 3 days, based on quantitation of the ATP present. There was a broad range of activities of the immunocytokines on IL-2R.beta..gamma., with the largest IL-2 deletions having the greatest reduction on proliferation, compared with equimolar amounts of free IL-2. The effect on .alpha..beta..gamma. activity is not as pronounced, but again the greatest reduction in proliferation is seen with the largest IL-2 deletions. Deletions in the first few N-terminal amino acids of IL-2 allow for fine tuning of cytokine activity. A representative experiment is shown in FIGS. 12(a) and (b).
[1605] Materials and Methods
[1606] IL-2R transfected TF1 cells were routinely cultured in RPMI+10% fetal bovine serum (culture medium) with the addition of IL-2 (Peprotech) at 5 ng/mL for the .beta. transfected cell line and IL-2 at 5 ng/mL and Geneticin (Gibco) at 350 .mu.g/mL for the .alpha..beta. transfected cell line. Prior to testing of immunocytokine constructs, the cells were harvested by centrifugation and aspirated to remove the supernatant. The cells were washed in PBS to remove cytokines and antibiotics. Cells were resuspended in fresh culture medium at 10.sup.5 cells/mL, without supplements and returned to the incubator overnight.
[1607] The cells were harvested by centrifugation and aspirated to remove the supernatant. Cells were resuspended in complete medium and 30 .mu.L of cell solution was added to the plate (white walled tissue culture treated 384-well plate) wells to achieve an initial cell concentration of 1250 cells/well.
[1608] The IL-2 ligand was prepared as serial four-fold dilutions from 300 ng/mL final assay concentration (FAC) (600 ng/mL working) in culture media. The immunocytokine constructs were titrated from 0.1 .mu.g/mL (three-fold dilutions) for testing on the .alpha..beta..gamma. cell line and 10 .mu.g/mL (three-fold dilutions) for the .beta..gamma. cell line. 30 .mu.L of titrations were added to the cell plate. To control wells, 30 .mu.L of culture media without IL-2 was added. To reduce evaporation effects, the outermost rows/columns of the plate were filled with 80 .mu.L of culture media. The plates were then incubated for 3 days at 37.degree. C., 5% CO.sub.2. Following the culture period proliferation of TF-1 cells was assessed by addition of 30 .mu.L of Cell Titre Glo (Promega) to all wells. The plate was incubated at room temperature for 10 minutes then read using ultrasensitive luminescence filter.
Equation 7 Calculation of fold over background from TF - 1 proliferation assay fold over background = sample RLU mean over background RLU RLU = relative luminescence units ##EQU00004##
[1609] Data expressed as fold over background. Background was defined as wells containing cells but no cytokine
Example 14--Binding of Immunocytokines to PD-L1
[1610] Surface plasmon resonance was used to confirm the ability of the immunocytokine constructs to bind PD-L1. The presence of the IL-2 on the light chain does not have any detrimental effect on binding (Table 9). Four constructs with a range of IL-2 activities were shortlisted for further characterisation--these were 1D05 D1-9 ICK, 1D05 D1-8 ICK, 1D05 D9-2 ICK and 1D05 D9-7 ICK.
TABLE-US-00012 TABLE 9 Affinity of 1D05 binding to PD-L1 is unaffected by the fusion of IL-2 to the antibody, as measured by surface plasmon resonance. Data shown is from a single experiment KD Sample Name (nM) 1D05 0.171 1D05 HC-IL2 0.240 1D05 LC-IL2 0.207 1D05 IC45 (D5-9) 0.203 1D05 IC46 (D1-9) 0.195 1D05 IC64 (D5-7) 0.214 1D05 D1-2 0.187 1D05 D1-3 0.199 1D05 D1-4 0.186 1D05 D1-5 0.203 1D05 D1-6 0.211 1D05 D1-7 0.178 1D05 D1-8 0.190 1D05 D9 0.205 1D05 D9-8 0.225 1D05 D9-7 0.200 1D05 D9-6 0.211 1D05 D9-4 0.175 1D05 D9-3 0.171 1D05 D9-2 0.189 1D05 D2-6 0.201 1D05 D3-7 0.203 1D05 D4-8 0.208 benchmark 0.099
[1611] Materials and Methods
[1612] Analysis of Immunocytokines by Surface Plasmon Resonance
[1613] Label-free surface plasmon resonance (SPR) analysis was carried out on the ProteOn XPR36 (BioRad) array SPR machine. An anti-human IgG capture surface was created on a GLC biosensor chip using amine coupling of an anti-human IgG from GE Healthcare. Test antibodies were captured on this surface and human PD-L1 (in-house) was used as the analyte at 64 nM, 16 nM, 4 nM, 1 nM and 0.25 nM. The assay was carried out at 25.degree. C. using HBS-EP (Teknova H8022). Buffer alone was used to reference the binding sensorgrams. The data was analysed using the 1:1 model inherent to the ProteOn XPR36 analysis software.
Example 15--Assessing Capacity of Immunocytokines to Neutralise the Interaction of PD-L1 and PD-1/CD80
[1614] To ensure that fusion of the IL-2 molecule to the antibody did not disrupt its neutralisation capacity, shortlisted immunocytokines were tested in a neutralisation ELISA. The shortlisted immunocytokines tested did not differ from wild type antibody in their ability to neutralise interactions between PD-L1 and PD-1, and PD-L1 and CD80. Results are shown in FIG. 13 and Table 10. Values in the table are the means of three independent experiments.
TABLE-US-00013 TABLE 10 Summary of neutralisation ELISA data, expressed as mean of three independent experiments PD1-PD-L1 CD80-PD-L1 Clone Neutralisation IC.sub.50 (nM) Neutralisation IC.sub.50 (nM) 1D05 1.41 0.882 1D05 LC-IL-2 0.833 0.505 1D05 IC46 (D1-9) 1.75 1.07 1D05 D1-8 1.16 0.745 1D05 D9-2 1.55 0.947 1D05 D9-7 1.15 0.70 Hybrid Control N/A N/A
[1615] Materials and Methods
[1616] a) PD-L1/PD-1 or PD-L1/CD80 Neutralisation ELISA
[1617] CD80 (R&D Systems) or PD-1 (in house) diluted to 2.5 .mu.g/mL were adsorbed to 96-well, low auto-fluorescent, high protein binding plates (Costar) overnight at 4.degree. C. Excess protein was removed by washing with PBS-Tween (0.1% v/v) and the wells were blocked with 1% w/v bovine serum albumin (BSA, Sigma) in PBS for 1 hour at room temperature, after which plates were washed as described previously. 60 .mu.L of a titration (three-fold dilutions from 100 nM) of antibody was added to a 96-well, non-binding plate diluted in ELISA assay buffer (PBS+0.1% BSA). 60 .mu.L of biotinylated PD-L1 (in house, labelled with Lightning Link Biotinylation kit) at 16 nM working concentration (8 nM FAC) was added to the plate excluding control wells where 60 .mu.L ELISA assay buffer was added. The plate was incubated for 30 min before transferring 50 .mu.L to the coated plates.
[1618] The coated plates were incubated for 1 hour at room temperature. Excess protein was removed by washing with PBS-Tween (0.1% v/v). PD-L1 binding was detected using streptavidin labelled Europium (Perkin Elmer) diluted 1/1000 in DELFIA assay buffer (Perkin Elmer). The plates were washed with TBS (Tris buffered saline)-Tween (0.1% v/v) and 50 .mu.L/well of DELFIA Enhancement solution (Perkin Elmer) was added to the plate. The time-resolved fluorescence was measured at 615 nm on an Envision plate reader (PerkinElmer). Percentage specific binding was calculated as defined in Equation 3.
[1619] IC.sub.50 values were determined using GraphPad Prism software by curve fitting using a four-parameter logistic equation (Equation 4) from the percentage specific binding (Equation 3).
Example 16--De-Immunisation of Anti-PD-L1 Antibody
[1620] To reduce the possibility of adverse immunological reactions based around the anti-PD-L1-immunocytokine, a series of 1D05 antibody mutants (Seq ID Nos:47 to 51) was created with anticipated lower potential of immunogenicity, as determined by T-cell epitope analysis software. The mutations can be single or in combination. Mutants were assessed for their ability to bind PD-L1 with the same affinity as the wild-type molecule by SPR as described in Example 14, with the addition of human PD-L1 analyte at 256 nM. Mutations under investigation are included as Seq ID Nos:47 to 51, indicated by underlined and bold text. The V.sub.H framework mutations (Seq ID Nos:47 and 48) do not have any detrimental effects on binding. The V to A mutation in CDRH2 (Seq ID No:50) was detrimental to binding, and so an alternative mutation will be analysed (V to Y, Seq ID No:298). Results are shown in Table 11.
Example 17--Inhibition of Tumour Growth by an Anti-PD-L1 Antibody in NOD/SCID: Xenograft T-Cell Model
[1621] Inhibition of melanoma tumour growth by lead antibody 1D05 in the hIgG1 LAGA (Seq ID No: 205) format was demonstrated in a NOD/SCID:xenograft T-cell model. T-cells were expanded in the presence of A375, a melanoma cell line, for 20 days in the presence of IL-2 and IL-7. T-cells were co-implanted subcutaneously with fresh A375 cells, then the antibody administered intraperitoneally after 1 hour. Tumour size and animal survival were monitored. Tumours in mice treated with antibody 1D05 were smaller than in animals treated with isotype control. Survival time in 1D05-treated mice was also increased.
[1622] Materials and Methods
[1623] Efficacy studies were performed using a T-cell/Xenograft model in NOD/SCID mice employing a refinement of the methods outlined in Stewart R et al. (Cancer Immunol. Res., 2015 September; 3(9):1052-62). Leukoreduction system chambers were obtained from NHSBT. HLA-A2 positive donors were selected by staining unfractionated blood using a PE-labelled anti-human HLA-A2 (Biolegend, Clone:
TABLE-US-00014 TABLE 11 Deimmunisation mutations to lead 1D05 antibody KD Heavy Chain Light Chain (nM) 1D05 - IgG1 disabled (LAGA) 1D05 kappa (Seq ID No: 45) 0.29 Seq ID No: 299 1D05 V to A change in V.sub.H (Seq 1D05 kappa (Seq ID No: 45) 0.33 ID No: 47), IgG1 disabled (LAGA, Seq ID No: 205) 1D05 F to S change in V.sub.H (Seq 1D05 kappa (Seq ID No: 45) 0.23 ID No: 48), IgG1 disabled (LAGA, Seq ID No: 205) 1D05 V to A & F to S change in 1D05 kappa (Seq ID No: 45) 0.23 V.sub.H (Seq ID No: 342), IgG1 disabled (LAGA, Seq ID No: 205) 1D05 - IgG1 disabled (LAGA) 1D05 kappa, V to A change 2.66 Seq ID No: 299 (Seq ID No: 50) 1D05 V to A change in V.sub.H (Seq 1D05 kappa, V to A change 2.8 ID No: 47), IgG1 disabled (LAGA, (Seq ID No: 50) Seq ID No: 205) 1D05 F to S change in V.sub.H (Seq 1D05 kappa, V to A change 1.94 ID No: 48), IgG1 disabled (LAGA, (Seq ID No: 50) Seq ID No: 205) 1D05 V to A & F to S change in 1D05 kappa, V to A change 1.94 V.sub.H (Seq ID No: 342), IgG1 (Seq ID No: 50) disabled (LAGA, Seq ID No: 205)
BB7.2), the red blood cells were then lysed, followed by fixation with 4% PFA, prior to acquisition on the Attune flow cytometer. PBMCs were isolated by density gradient centrifugation over Ficoll. Primary human CD4.sup.+ and CD8.sup.+ T-cells were then isolated using an EasySep human CD4.sup.+ and CD8.sup.+ T-cell enrichment kit (Stemcell Technologies, Cat 19052 and 19053). The CD4.sup.+ and CD8.sup.+ T-cells were then cultured separately for 20 days on a monolayer of mitomycin C treated A375 cells (at day 10, T-cells were re-plated on a fresh A375 monolayer) in the presence of recombinant human IL-2 and IL-7 (Peprotech). On day 20 the cells were frozen in 90% hiFBS/10% DMSO at -80.degree. C. in a "Mr Frosty" (Nalgene) and stored in liquid nitrogen until required. The day, before starting an in vivo experiment the cells were thawed and placed in culture.
[1624] On the day of implantation, the CD4.sup.+ and CD8.sup.+ T-cells were counted and mixed together in a 1:1 ratio. The CD4.sup.+/CD8.sup.+ mixture was then added to A375 tumour cells and injected subcutaneously into mice on the rear right flank. Treated groups received their first dose of antibody or isotype control (all dosed intraperitoneally at 10 mg/kg) one-hour post implantation of the cells. The animals received further doses 3, 6, 8 and 10 days post-implantation. Tumour development was monitored three times a week using digital callipers measuring in two dimensions until end of the study. Tumour volumes (mm.sup.3) were estimated using a standard formula (L.times.W.sup.2)/2 (with L being the larger diameter, and W the smaller diameter of the tumour). Mice were kept on studies until their tumours developed to a mean diameter of 12 mm or they reached one of the humane endpoints outlined in the study protocol. The humane endpoint survival statistics were calculated using the Kaplan-Meier method with Prism. This approach was used to determine if PD-L1 treatment was associated with improved survival.
TABLE-US-00015 TABLE 12 Treatment Groups Number of Groups animals Cell Line 1 9 2 .times. 10.sup.6 A375 Cells 2 9 1:6 ratio T-cells: A375 Cells (2 .times. 10.sup.6 A375 Cells) 3 8 10 mg/kg isotype control hIgG1 1:6 ratio T-cells: A375 Cells (2 .times. 10.sup.6 A375 Cells) 4 8 10 mg/kg anti-PD-L1 1D05 hIgG1 LAGA (Seq ID No: 205) 1:6 ratio T-cells: A375 Cells (2 .times. 10.sup.6 A375 Cells)
[1625] Treatment with the isotype control had no effect on tumour development when compared to the group where the CD4.sup.+/8.sup.+ T-cells are co-injected with the tumour cells. Whilst treatment with the anti-PDL1 antibody 1D05 delayed the tumour development when compared to the Isotype Control. This is shown in FIG. 14.
[1626] All groups with T-cells co-injected with the tumours showed an increase in time on study when compared to the tumour alone group. Treatment with the isotype control had no effect on time on study, whilst treatment with the anti-PDL1 antibody 1D05 increased time on study when compared to all the other groups including the isotype control groups. Results are shown in FIG. 15.
Example 18: Single Dose Study of Immunocytokines in Cynomolgus Monkeys
[1627] To assess pharmacodynamic and pharmacokinetic (PK) parameters in the most relevant animal model, male cynomolgus monkeys received a single dose of immunocytokine (ICK) at 1 mg/kg. Animals were observed for clinical manifestations of toxicity, and blood samples were taken over the course of 7 days for the analysis of PK, production of cytokines and characterisation of leukocyte subsets. The in-life phase of the study, and haematology, flow cytometry and cytokine analysis was performed at Envigo UK (study number GF13YC). Pharmacokinetic analysis was performed in-house.
[1628] Materials and Methods
[1629] Male cynomolgus monkeys of at least 2 years of age were used for the study and body weights were recorded at 7 days and 4 days before the start of the study. Immunocytokine constructs were formulated in 50 mM sodium acetate pH 5.5, at 1 mg/mL and were diluted to 0.2 mg/mL in physiological saline for intravenous infusion at a rate of 5 mL/kg/hour. Blood pressure and body temperature were monitored pre-treatment, 1 hour and 4 hours post end-of-dose. Animals were observed twice daily for signs of ill-health. The study was performed in two phases--initial doses of 1D05 HC IL-2 ICK and 1D05 LC D9-7 ICK to ensure dose level and PK timepoints were suitable, then dosing of 1D05 LC D9-7 ICK was repeated, alongside four further constructs (see Table 1). Phase 2 dosing of 1D05 LC D9-7 ICK is indicated by a (2) next to the construct name.
TABLE-US-00016 TABLE 13 Treatment groups and animal numbers Phase Animal Test Item 1 134 1D05 HC IL-2 ICK 1 135 1D05 LC D9-7 ICK 2 136 1D05 LC IL-2 ICK 2 137 1D05 LC D9-7 ICK (2) 2 138 1D05 LC D9-2 ICK 2 139 1D05 LC D1-8 ICK 2 140 1D05 LC D1-9 ICK
[1630] For haematological analysis, fasting blood samples were taken into EDTA treated tubes pre-treatment, and 2, 5 and 7 days post-treatment. Routine haematology parameters were measured by the Bayer Advia 120. Results are shown in FIGS. 16 and 17.
[1631] For analysis of cytokines and soluble CD25, blood samples were taken into EDTA-treated tubes pre-treatment and 3 days post-treatment, and plasma extracted by centrifugation at 2000 g for 10 minutes. Samples were frozen until analysis by multiplex MSD (cytokines) or commercial ELISA (soluble CD25). Results are shown in FIGS. 18 and 19.
[1632] For immunophenotyping, blood samples were taken into EDTA-treated tubes pre-treatment and 5 days post-treatment. Blood samples were stained with cocktails of directly conjugated monoclonal antibodies, then red blood cells were lysed and the samples fixed by re-suspension in phosphate buffered saline containing 1% formaldehyde prior to analysis. Results are shown in FIG. 20.
[1633] For PK analysis, blood samples were taken into untreated tubes pre-treatment, end of infusion (EOI), 2, 4, 8, 16, 24, 32, 40 and 48 hours after EOI, extended to 72 hours and 96 hours for Phase 2) and serum prepared by allowing the blood to clot, then centrifugation at 2000 g for 10 min. Serum samples were frozen on dry ice for shipment to Kymab. Results are shown in FIG. 21.
[1634] Pharmacokinetic Analysis of Serum Samples
[1635] a) PK Assay for Detection of Anti-PD-L1 Antibody
[1636] 50 .mu.L/well of human PD-L1 Flag His (Seq ID No:505, in house) diluted to 2 .mu.g/mL in PBS (Sigma, P3813-10PAK) was adsorbed to 96-well, high protein binding fluorescent plates (Greiner) overnight at 4.degree. C. Excess protein was removed by washing 3.times. with 300 .mu.L/well PBS-Tween (0.1% v/v) and the wells were blocked with 1% w/v bovine serum albumin (BSA, Sigma) in PBS for 1 hour at room temperature, after which plates were washed as described previously. Antibodies were diluted from 10,000 ng/mL to 9.77 ng/mL (1/2 dilution) in pooled cynomolgus serum (Seralab, CYNSRM) to give 12 standards including a blank. Standards, quality controls and samples were diluted at 1 in 50 MRD (minimum required dilution) in ELISA assay buffer (PBS+0.1% BSA) and were added to the coated 96-well high-binding plates at 50 .mu.L/well. The plate was incubated for 1 hour at room temperature, after which plates were washed 3.times. with PBS-Tween. 50 .mu.L biotinylated goat anti-human IgG (Southern Biotech) at 1 .mu.g/mL was added to the plate. The plate was incubated for 1 hour at room temperature, after which plates were washed 3.times. with PBS-Tween. PD-L1 binding was detected using streptavidin labelled Europium (Perkin Elmer) diluted 1/1000 in DELFIA assay buffer (Perkin Elmer). Plates were washed with TBS (Tris buffered saline)-Tween (0.1% v/v) and 50 .mu.L/well of DELFIA Enhancement solution (Perkin Elmer) was added to the plate. Time-resolved fluorescence was measured at 615 nm on an Envision plate reader (PerkinElmer). Concentrations were determined using GraphPad Prism software by interpolating from a standard curve fitted using a four-parameter logistic equation (Equation 4). Results are shown in FIGS. 21a and 21b.
[1637] b) PK Assay for Detection of Intact Immunocytokine (Antibody Fused to IL-2)
[1638] 50 .mu.L/well of human PD-L1 Flag His (Seq ID No:505 in house) diluted to 3 .mu.g/mL in PBS (Sigma, P3813-10PAK) was adsorbed to 96-well, low auto-fluorescent, high protein binding plates (Costar) overnight at 4.degree. C. Excess protein was removed by washing 3.times. with 300 .mu.L/well PBS-Tween (0.1% v/v) and the wells were blocked with 1% w/v bovine serum albumin (BSA, Sigma) in PBS for 1 hour at room temperature, after which plates were washed 3.times. with PBS-Tween. Antibodies were diluted from 50,000 ng/mL to 617.3 ng/mL in pooled cynomolgus serum (Seralab, CYNSRM) to give 10 standards including a blank. Standards, quality controls and samples were diluted at 1 in 20 MRD in ELISA assay buffer (PBS+0.1% BSA) and were added to the coated 96-well high-binding plates at 50 .mu.L/well. The plate was incubated for 1 hour at room temperature, after which plates were washed 3.times. with PBS-Tween. 50 .mu.L biotinylated anti-human IL-2 (Peprotech) at 2 .mu.g/mL was added to the plate. The plate was incubated for 1 hour at room temperature, after which plates were washed as described previously. Binding was detected using streptavidin labelled Europium (Perkin Elmer) diluted 1/1000 in DELFIA assay buffer (Perkin Elmer). Plates were washed with TBS (Tris buffered saline)-Tween (0.1% v/v) and 50 .mu.L/well of DELFIA Enhancement solution (Perkin Elmer) was added to the plate. Time-resolved fluorescence was measured at 615 nm on an Envision plate reader (PerkinElmer). Concentrations were determined using GraphPad Prism software by interpolating from a standard curve fitted using a four-parameter logistic equation. Results are shown in FIGS. 21c and 21d.
[1639] Results Summary
[1640] No signs of overt IL-2 mediated toxicity (fever, vascular leak, diarrhea) were observed after dosing. Lymphocyte numbers increased over the duration of the study with the different immunocytokine constructs. The constructs with the greatest truncations induced the lowest levels of lymphocyte expansion; little expansion was observed with 1D05 LC D1-9 ICK or 1D05 LC D1-8 ICK over the seven-day period, whereas 1D05 LC D9-7 ICK and the full-length IL-2 induced significant expansion. The lymphopenia observed at day 2 with some constructs is indicative of lymphocyte margination out of the circulation. This is followed by a rebound lymphocytosis which can be seen at day 5 (FIG. 16).
[1641] Administration of immunocytokine constructs did not cause significant anemia (FIG. 17). Around a 20% reduction in haemoglobin, haematocrit and red blood cell levels was observed at day 7 with the most active constructs (1D05 HC IL-2 ICK, 1D05 LC IL2 ICK and 1D05 LC D9-7 ICK), and around a 10% reduction with the other constructs. This agrees with anecdotal evidence from studies with IL-2 heavy chain immunocytokines. Thrombocytopenia (reduced platelet count) was not observed.
[1642] IL-2 was strongly increased 3 days post-dosing, indicative of production by activated T-cells. However, there is a possibility that the assay is cross-reactive for human IL-2 and so could also detect the immunocytokine. There was no clear up- or down-regulation of any of the other cytokines post-dosing, although there was a trend for down-regulation of IL-8 levels (FIG. 18). Levels of soluble CD25, which is a biomarker of T-cell activation, were strongly increased 3 days after dosing with immunocytokines (FIG. 19). Levels of soluble CD25 correlated with the in vitro stimulatory activity of the immunocytokines described in Example 13.
[1643] Dosing with immunocytokines increases the number of activated T-cells in the blood (FIG. 20). When dosed with 1D05 LC IL-2 ICK, total CD4.sup.+ and CD8.sup.+ cell numbers are increased, but CD69+(early activation) and CD25.sup.+ (later activation) subsets are greatly increased, compared with pre-treatment levels. The increase in cell numbers is less striking for the truncated constructs. No significant changes in B-cell, NK cell or neutrophil numbers were observed, with a moderate increase in monocyte numbers. Data for the animal dosed with 1D05 LC D9-7 ICK is not available, due to clotting of the sample.
[1644] The light chain (LC) fusions have a longer half-life than the heavy chain (HC) fusion, which agrees with previous data in mouse (Gillies S D, Protein Engineering, Design and Selection, 26:10: 561-569, 2013). The half-life of 1D05 LC IL-2 ICK was around 8 hours, and the half-life of the truncated IL-2 constructs was around two-fold longer (FIGS. 21a and 21b). The increased half-life of immunocytokines with truncated IL-2, compared with full-length IL-2, may reflect reduced binding to IL-2 receptors.
[1645] A modified assay was used to detect intact immunocytokine i.e. antibody fused to IL-2 (FIGS. 21c and 21d). This result shows that the IL-2 part of the molecule remains fused in vivo and is not cleaved.
Example 19--Extended Single Dose Study in Cynomolgus Monkeys
[1646] To determine the duration of lymphocytosis, and obtain more detailed analysis of T-cell subsets, an extended single dose study will be performed (study number HQ52PV). Female cynomolgus monkeys are dosed with 1 mg/kg immunocytokine as per Example 18 and monitored over at least 14 days. Cytokines will be analysed on days 1, 3, 7, 10 and 14, and pre-treatment. Haematology measurements will be performed on days 2, 5, 7, 10 and 14, and pre-treatment. Detection of soluble CD25 will be performed on days 3, 7 and 10, and pre-treatment. CD127 will be added to the immunophenotyping panel, to allow for detection of regulatory T-cells (CD3.sup.+CD4.sup.+CD25.sup.hi CD127.sup.lo), and analysis will be performed on days 1, 5, 7, 10 and 14, and pre-treatment. PK analysis will be performed as before. Treatment groups are shown in Table 14.
TABLE-US-00017 TABLE 14 Treatment groups and animal numbers Phase Animal Test Item 3 378 1D05 LC IL-2 ICK 3 379 1D05 LC D9-7 ICK 3 380 1D05 LC D9-2 ICK 3 381 1D05 LC D1-8 ICK 3 382 1D05 LC D5-9 ICK
[1647] No signs of overt IL-2 mediated toxicity (fever, vascular leak, diarrhea) were observed after dosing. Lymphocyte numbers peaked at day 7 with all immunocytokine constructs. The constructs with the greatest truncations induced the lowest levels of lymphocyte expansion; the least expansion was observed with 1D05 LC D1-8 ICK, whereas 1D05 LC D9-7 ICK and the full-length IL-2 induced the greatest expansion. The lymphopenia observed at day 2 with some constructs is indicative of lymphocyte margination out of the circulation. This is followed by a rebound lymphocytosis (FIG. 28). Levels of soluble CD25, which is a biomarker of T-cell activation, peaked 3 days after dosing with immunocytokines (FIG. 29). Levels of soluble CD25 correlated with the in vitro stimulatory activity of the immunocytokines described in Example 13.
[1648] Administration of immunocytokine constructs did not cause significant anemia (FIG. 30). A 10-20% reduction in haemoglobin, haematocrit and red blood cell levels was observed with the immunocytokine constructs. Haemoglobin levels remained lower over the entire time course in the animal dosed with 1D05 LC D1-8 ICK. Mild thrombocytopenia was observed with the two most active constructs at day 5, but levels recovered after this timepoint. IL-2 was strongly increased 3 days post-dosing, indicative of production by activated T-cells. However, there is a possibility that the assay is cross-reactive for human IL-2 and so these levels reflect the presence of the immunocytokine. There was no clear up- or down-regulation of any of the other cytokines post-dosing (FIG. 31).
[1649] As observed previously in Example 18, the half-life of 1D05 LC IL-2 ICK was around 8 hours, and the half-life of the truncated IL-2 constructs correlated with the size of the truncation (FIG. 32). The immunocytokine construct containing the longest truncation, D1-8, had the longest half-life, of approximately 24 hours. The increased half-life of immunocytokines with truncated IL-2, compared with full-length IL-2, may reflect reduced binding to IL-2 receptors.
[1650] Expansion of CD4.sup.+ and CD8.sup.+ T-cells is shown in FIG. 33. As observed for the automated cell counts, the degree of expansion correlates well with the size of the IL-2 truncation, the greatest expansion of both T-cell subsets observed in the animal dosed with 1D05 LC ICK D9-7.
Example 20--Binding to Cell Endogenously Expressed hPD-L1 and Neutralisation of hPD-L1 Binding to PD-1 and CD80
[1651] Lead antibodies are tested for ability to bind to ES2 cells endogenously expressing hPD-L1 as well as the neutralisation of PD-L1/PD-1 interaction and PD-L1/CD80 interactions. ES2 cells endogenously expressing hPD-L1 (ATCC) are diluted in FACS buffer (PBS 1% BSA 0.1% sodium azide) and distributed to three 96-well, V-bottom plate (Greiner) at a density of 0.5-1.times.10.sup.5 cells per well. Cells are washed with 150 .mu.L PBS and centrifuged at 300 g for 3 minutes. Supernatant is aspirated and 150 .mu.L PBS added. This wash step is repeated.
[1652] To plate 1 (PD-L1 binding), 25 .mu.L lead antibody, reference antibody or control antibody diluted in FACS buffer is added to the washed cells. 25 .mu.L FACS buffer is added and cells are incubated at 4.degree. C. for 60 minutes. 150 .mu.L FACS buffer is added and cells washed as described above. To detect anti-PD-L1 antibody binding, anti-human PE (Jackson ImmunoResearch) is diluted 1/500 in FACS buffer and 50 .mu.L of this mixture added to cells. Cells are incubated 4.degree. C. for 60 minutes. Cells are washed twice with 150 .mu.L FACS buffer, centrifuging at 300 g for 3 minutes after each wash step and aspirating supernatant. Cells are fixed by addition of 50 .mu.L 4% paraformaldehyde and overnight incubation at 4.degree. C. Cells are washed once as above and resuspended in FACS buffer for analysis. PE signal intensity (geometric mean) is measured by flow cytometry using a Beckman Coulter Cytoflex instrument. Data is plotted as geometric mean values without further calculation.
[1653] To plate 2 (PD-1 neutralisation) 25 .mu.L lead antibody, reference antibody or control antibody diluted in FACS buffer is added to the washed cells. 25 .mu.L of biotinylated human PD-1 (in-house, Fc-tagged, SEQ ID No:6) is added and cells are incubated at 4.degree. C. for 60 minutes. Biotinylation is performed in-house using Lightning Link conjugation kit (Innova Biosciences) according to manufacturer's instructions. 150 .mu.L FACS buffer is added and cells washed as described above. To detect biotinylated PD-1, Streptavidin-Alexa Fluor 647 (AF647, Jackson ImmunoResearch) is diluted 1/500 in FACS buffer and 50 .mu.L of this mixture added to cells. Cells are incubated at 4.degree. C. for 60 minutes. Cells are washed twice with 150 .mu.L FACS buffer, centrifuging at 300 g for 3 minutes after each wash step and aspirating supernatant. Cells are fixed, washed and resuspended for analysis as above. APC signal intensity (geometric mean) is measured by flow cytometry. Data is plotted as geometric mean values without further calculation.
[1654] To plate 3 (CD80 neutralisation) 25 .mu.L lead antibody, reference antibody or control antibody diluted in FACS buffer is added to the washed cells. 25 .mu.L biotinylated human CD80 (Fc tagged, R&D Systems, 140-B1) is added and cells are incubated at 4.degree. C. for 60 minutes. All other steps are performed as per plate 2.
[1655] Alternatively, to simultaneously detect binding and neutralisation, ES2 cells expressing hPD-L1 are diluted in FACS buffer and are distributed to two 96-well, V-bottom plate (Greiner) at a density of 0.5-1.times.10.sup.5 cells per well. Cells are washed with 150 .mu.L PBS and centrifuged at 300 g for 3 minutes. Supernatant is aspirated and 150 .mu.L PBS added. This wash step is repeated.
[1656] 25 .mu.L lead antibody, reference antibody or control antibody diluted in FACS buffer is added to the washed cells. 25 .mu.L biotinylated human PD-1 (R&D Systems, 8986-PD-100, his-tagged) or CD80 (R&D Systems, 9050-B1-100, his-tagged) is added and cells are incubated at 4.degree. C. for 60 minutes. 150 .mu.L FACS buffer is added and cells washed as described above. To detect biotinylated PD-1 or CD80 and anti-PD-L1 antibody binding, streptavidin-AF647 and anti-human PE are each diluted 1:500 in FACS buffer and 50 .mu.L of this mixture added to cells. Cells are incubated at 4.degree. C. for 60 minutes. Cells are washed twice with 150 .mu.L FACS buffer, centrifuging at 300 g for 3 minutes after each wash step and aspirating supernatant. Cells are fixed, washed and resuspended for analysis as above. PE and APC signal intensity (geometric mean) are measured by flow cytometry. Data is plotted as geometric mean values without further calculation. Alternatively, an anti-his tag antibody conjugated to APC (R&D Systems) may be used to detect PD-1 or CD80, or PD-1 and CD80 may be directly labelled with AF647.
Example 21--Testing of Lead Anti-PD-L1 Antibodies in Reporter Cell Bioassay
[1657] The ability of anti-PDL1 antibodies to neutralise PD-L1/PD-1 interaction on cells will be determined using a bioluminescence cell based assay (Promega.RTM.). PD-L1 aAPC/CHO-K1 cells, transfected with PD-L1 and a cell surface protein designed to promote TCR activation, are co-cultured with PD-1 expressing Jurkat cells. These cells also present a NFAT induced luciferase responsive element. Co-culture of the two cell types in the presence of an antibody able to block PD-1-PD-L1 interaction activates TCR signaling and NFAT-mediated luciferase activity.
[1658] The assay is run according to manufacturer's recommendations. Briefly, PD-L1 aAPC/CHO-K1 cells are cultured overnight in Hams F12 medium supplemented with 10% hiFBS. The next day, media is removed, effector PD-1 Jurkat cells and anti-PD-L1 antibodies are added to assay plates for 6 hours at 37.degree. C. in RPMI 1640 supplemented with 1% hiFBS. Plates are read following 10 minutes of incubation with Bio-Glo.TM. on the Envision plate reader using luminescence settings. Antibody-induced luciferase activity is represented as fold induction compared to assay signal of wells showing background levels of response as defined by Equation 8. EC.sub.50 values are calculated using a 4-parameter logistic fit (Equation 4).
Fold induction=sample well/basal luciferase response
Basal luciferase response=value from wells containing PD-L1 CHO-K1 cells and PD-1 Jurkat cells Equation 8
Example 22--Pharmacokinetic Study of Lead Antibodies in hPD-L1 Expressing Mice
[1659] Lead antibodies, in human IgG1 effector enabled format (i.e. having a constant region of wild type IgG1, Seq ID No:341), are dosed intra-peritoneally at 10 mg/kg in mice expressing human PD-L1, eight mice per antibody. Blood samples are taken pre-treatment and at 2, 4, 8, 12, 24, 48, 72, 96, 192, 336, 508 and 672 hours. Serum is prepared and samples frozen until analysis. Samples will be analysed according to the method described for detection of antibody in Example 18 with the following exception: serum from C57BL/6 mice will be used as the vehicle in which to prepare standard curves and blanks. Minimum required dilution will differ from Example 18 due to the larger dose administered; this will be determined empirically.
Example 23--Pharmacokinetic Study of Lead Antibodies in Non-Human Primates
[1660] Lead antibodies, in human IgG1 effector enabled format (i.e. having a constant region of wild type IgG1, Seq ID No:341), are dosed intravenously at 10 mg/kg in male cynomolgus monkeys, three animals per antibody. Blood samples are taken pre-treatment and at 2, 4, 8, 12, 24, 48, 72, 96, 192, 336, 508 and 672 hours. Serum is prepared and samples frozen until analysis. Samples will be analysed according to the method described for detection of antibody in Example 18. Minimum required dilution will differ from Example 18 due to the larger dose administered; this will be determined empirically.
Example 24: Lead Antibody Activity in a Mouse B Cell:T-Cell Hybridoma Assay
[1661] Antibodies were tested in a murine B cell:T-cell hybridoma co-culture assay to assess induction of IL-2. 50 .mu.L of human PD-L1 (SEQ ID No:1) transfected LK35.2 mouse B-lymphocyte hybridoma cells (ATCC) prepared in DMEM (Gibco) supplemented with 1% Foetal Bovine Serum (Gibco) were treated with 10 .mu.M Ovalbumin323-329 peptide (Thermo Scientific) and dispensed at a density of 2.times.10.sup.4 cells/well in a 96-well tissue culture treated plate (Costar). Ovalbumin peptide loaded cells were then mixed with 50 .mu.L 1:3 titration series of anti-PD-L1 antibodies or anti-ICOS/PD-L1 bi-specific antibodies in a mAb.sup.2.TM. format from 30 nM for 9 concentration points in DMEM supplemented with 1% Foetal Bovine Serum.
[1662] Following 1 hour incubation at 37.degree. C. 5% CO.sub.2, 100 .mu.L of murine T-helper hybridoma cell line DO-11-10 (National Jewish Health) cultured overnight in DMEM (Gibco) supplemented with 1% Foetal Bovine Serum (Gibco) were added to assay plate at 2.times.10.sup.4 cells/well. LK35.2/DO-11-10 co-culture was incubated overnight at 37.degree. C. 5% CO.sub.2 before supernatant was collected to assess production of mouse IL-2. Cells treated with 1 or 0.1.times. working stock of cell stimulation cocktail (eBioscience) were used as positive control for murine IL-2 production.
[1663] Mouse IL-2 quantification was performed using the mouse IL-2 Duoset ELISA kit (R&D Systems) following manufacturer's protocol, modified to include streptavidin-Europium as the detection reagent (DELFIA.RTM.). Briefly, assay plates were coated overnight at 4.degree. C. with provided capture antibody prepared in PBS at 1 .mu.g/mL. Plates were washed three times with PBS-Tween (0.1% v/v) before adding 200 .mu.L of 1% w/v bovine serum albumin (BSA, Sigma) in PBS for 1 hour at room temperature. 50 .mu.L cell supernatants were added to assay plates following a washing step performed as described previously. Following one hour incubation, 50 .mu.L of provided detection antibody at 200 .mu.g/mL prepared in 0.1% w/v BSA in PBS was added and plates were incubated for a further hour. Plates were washed as described above and 50 .mu.L of DELFIA.RTM. Eu-N1 streptavidin diluted 1:500 from stock solution in DELFIA.RTM. assay buffer (Perkin Elmer) were added to all wells for 1 hour. An additional washing step was performed using DELFIA wash buffer (0.5 M Tris HCL (Gibco), 1% Tween v/v (Sigma)) before the addition of 50 .mu.L DELFIA.RTM. Enhancement Solution (Perkin Elmer). The plate was incubated for 5 minutes at room temperature protected from the light and read at 615 nm using appropriate settings for DELFIA.RTM. time resolved fluorescence on an Envision plate reader (Perkin Elmer). The concentration of mouse IL-2 was interpolated from a standard curve run alongside test samples. Final plotted values were calculated using Equation 9, where background signal was calculated using assay signal of co-culture cells treated with 50 .mu.L of media only. Results are shown in FIG. 23. All antibodies potently enhance production of IL-2 in this co-culture system.
Equation 9=Mouse IL-2 (pg/mL)-Background
TABLE-US-00018 TABLE 15 EC.sub.50 values for induction of IL-2 by of PD-L1 antibodies in a murine T cell hybridoma assay EC.sub.50 (nM) Antibody name n = 1 n = 2 n = 3 1D05 0.33 0.21 0.21 84G09 0.24 0.14 0.17 413G05 0.29 0.11 0.29 414B06 0.59 0.30 0.18 416E01 0.19 0.15 0.14 benchmark 2 0.26 0.13 0.13
TABLE-US-00019 TABLE 16 EC.sub.50 values for induction of IL-2 by ICOS/PD- L1 bi-specific mAb.sup.2 .TM. antibodies in a murine T cell hybridoma assay Bispecific EC.sub.50 (nM) antibody name n = 1 n = 2 n = 3 ICOS/PD-L1 bispecific 1 0.386 n/a 0.888 Hybrid Control/PD-L1 bispecific 0.247 0.311 0.162 ICOS/PD-L1 bispecific 2 0.338 0.217 0.229 PD-L1 benchmark 0.122 0.128 0.123
Example 25--Testing of Lead Antibodies in an Activated DC--T-Cell Mixed Lymphocyte Reaction
[1664] Monocytes were isolated from cryopreserved PBMCs by negative selection methods using a Monocyte Isolation Kit and the MACS.TM. magnetic separation system (Miltenyi Biotec). Monocytes were resuspended in RPMI 1640 medium containing 10% hiFBS and 100 ng/mL GM-CSF and IL-4 (both Peprotech). Cells were cultured for 5 days in non-TC treated 6-well plates (Greiner) to induce differentiation of DCs, before addition of 100 ng/mL lipopolysaccharide from E. coli O55:B5 (Sigma) to activate the DCs. Cells were harvested after 24 hours of activation, and washed once with PBS to remove LPS, and resuspended at 10.sup.6/mL in RPMI 10% hiFBS. Allogeneic CD3.sup.+ T-cells were isolated from cryopreserved PBMC using a Pan T-Cell Isolation kit and the MACS system as above, and resuspended at 2.times.10.sup.6/mL in RPMI 10% hiFBS. Serial dilutions of selected antibodies (1:3) from 10 nM were prepared in RPMI 10% hiFBS and 50 .mu.L added to 96-well, flat-bottomed TC plates plates in triplicate. DCs (100 .mu.L) and T-cells (50 .mu.L) were added to plates and incubated at 37.degree. C., 5% CO.sub.2 for five days. Supematants were removed after three days for measurement of IL-2, and five days for measurement of IFN.gamma.. Supernatants were stored at -20.degree. C. until use. Cytokine production was measured with the R&D Systems Human IFN.gamma. and IL-2 Duoset.RTM. ELISA, using DELFIA.RTM. Eu-N1 Streptavidin detection. Results are shown in FIG. 24.
Example 26--Multi-Dose Study of Immunocytokines in Cynomolgus Monkeys
[1665] Pharmacology and toxicity of two immunocytokines, 1D05 D9-7 ICK and 1D05 D1-8 ICK (as described in Example 14), will be assessed in a multi-dose study in cynomolgus monkeys. Male juvenile monkeys are dosed with 1 mg/kg/dose according to two different regimens: Regimen 1--animals dosed on day 0 and day 14; regimen 2--animals dosed on day 0, 2, 14 and 16. Two animals will be dosed per group and monitored for 28 days. Treatment groups are shown in Table 17.
TABLE-US-00020 TABLE 17 Treatment groups for multi-dose study Treatment Animals group Construct Regimen per group 1 1D05 D9-7 ICK 1 2 2 1D05 D9-7 ICK 2 2 3 1D05 D1-8 ICK 1 2 4 1D05 D1-8 ICK 2 2
[1666] Heart rate, body temperature, respiration rate and blood pressure will be measured 1 hour and 4 hours after dosing. Body weight will be monitored daily. Cytokines will be analysed on days 2, 5, 7, 10, 14, 16, 19, 21, 24 and 28, and pre-treatment. Haematology measurements will be performed on days 2, 5, 7, 10, 14, 16, 19, 21, 24 and 28, and pre-treatment. Detection of soluble CD25 will be performed on days 3, 7 and 10, 17, 21, 24, and pre-treatment. Immunophenotyping will be performed on days 7, 10, 14, 24 and 28, and pre-treatment, according to the panel described in Example 19. Samples for pharmacokinetic (PK) analysis will be taken at the following timepoints at each infusion: pre-treatment, end of infusion, and at 8, 16, 24, 32, 40, 48, 72, and 96 hours.
Example 27: Immunocytokine Efficacy Study in a Syngeneic Tumour Model
[1667] An efficacy study will be performed using a CT-26 mouse tumour model, to compare surrogate immunocytokine activity with unmodified antibody, and to assess the role of effector function. On the day of implantation, BALB/c mice are injected subcutaneously into mice on the rear right flank with 1.times.10.sup.5 CT-26 cells/animal. Treated groups will receive their first dose of antibody or relevant control (all dosed intraperitoneally at 10 mg/kg) 6 days post implantation of the tumour cells and will be dosed three times a week for a total of two weeks. Tumour development will be monitored three times a week using digital callipers measuring in two dimensions until end of the study. Tumour volumes (mm.sup.3) will be estimated using a standard formula (L.times.W.sup.2)/2 (with L being the larger diameter, and W the smaller diameter of the tumour). Mice are kept on studies until their tumours developed to a mean diameter of 12 mm or they reached the one humane endpoints outlined in the study protocol. The humane endpoint survival statistics will be calculated using the Kaplan-Meier method with Prism.
TABLE-US-00021 TABLE 18 Treatment groups for efficacy study Number of Groups animals Treatment 1 10 10 mg/kg hybrid control huIgG1.sub.LAGA (constant region SEQ ID No: 205) 2 10 10 mg/kg hybrid control huIgG1.sub.LAGA IL-2 3 10 10 mg/kg anti-PD-L1 huIgG1.sub.LAGA 4 10 10 mg/kg anti-PD-L1 huIgG1.sub.LAGA IL-2 5 10 10 mg/kg anti-PD-L1 huIgG1 (constant region SEQ ID No: 340) 6 10 10 mg/kg anti-PD-L1 huIgG1 IL-2
Example 28: Immunocytokine Efficacy Study in a T-Cell: Melanoma Cell Line Xenograft Model
[1668] An efficacy study will be performed using a T-cell: A375 cell line xenograft model in NOD/SCID mice employing a refinement of the methods outlined in R. Stewart et al. Briefly HLA-A2 positive donors are selected by staining whole blood using a PE labelled anti-human HLA-A2 antibody (Biolegend), followed by red blood cell lysis and analysis by flow cytometry. Primary human CD4.sup.+ and CD8.sup.+ T-cells will then be isolated, using an EasySep human CD4.sup.+ or CD8.sup.+ T-cell enrichment kit, Stemcell Technologies, Cat 19052/3). The CD4.sup.+ and CD8.sup.+ cells are then cultured separately for 20 days on a monolayer of mitomycin C treated A375 cells in the presence of IL-2 and IL-7. T-cells are plated on a fresh feeder layer of A375 at day 10. On day 20, the cells are cryopreserved and stored in liquid nitrogen till required. The day before implantation, T-cells are thawed and cultured in medium plus cytokines overnight. On the day of implantation, the CD4.sup.+ and CD8.sup.+ cells are counted and mixed together in a 1:1 ratio. The T-cells are mixed with fresh A375 tumour cells at 1:6 ratio and injected subcutaneously into mice on the rear right flank. Treated groups will receive their dose of antibody, immunocytokine or relevant control (all dosed intraperitoneally at 10 mg/kg) one-hour post implantation of the T-cells and tumour cells. Tumour development will be monitored three times a week using digital callipers measuring in two dimensions until end of the study. Tumour volumes (mm.sup.3) will be estimated using a standard formula (L.times.W.sup.2)/2 (with L being the larger diameter, and W the smaller diameter of the tumour). Mice are kept on studies until their tumours developed to a mean diameter of 12 mm or they reached the one humane endpoints outlined in the study protocol. The humane endpoint survival statistics will be calculated using the Kaplan-Meier method with Prism. This approach will be used to determine which treatment is/are associated with improved survival. Subsequent studies will compare the immunocytokine constructs with different IL-2 activities.
TABLE-US-00022 TABLE 19 Treatment groups for efficacy study Number of Groups animals Treatment 1 10 2 .times. 10.sup.6 A375 Cells 2 10 CD4.sup.+/CD8.sup.+ T cells: A375 Cells 3 10 10 mg/kg hybrid control CD4.sup.+/CD8.sup.+ T cells: A375 Cells 4 10 10 mg/kg hybrid control IL-2 CD4.sup.+/CD8.sup.+ T cells: A375 Cells 5 10 200,000 IU recombinant human IL-2 CD4.sup.+/CD8.sup.+ T cells: A375 Cells (2 .times. 10.sup.6 A375 Cells) 6 10 10 mg/kg 1D05 CD4.sup.+/CD8.sup.+ T cells: A375 Cells (2 .times. 10.sup.6 A375 Cells) 7 10 10 mg/kg 1D05 LC IL-2 CD4.sup.+/CD8.sup.+ T cells: A375 Cells (2 .times. 10.sup.6 A375 Cells)
Example 29: Activity of Lead Antibodies in a Reporter Assay of Effector Function
[1669] The antibody-dependent cell-mediated cytotoxicity (ADCC) activity of selected antibodies was evaluated using an ADCC Reporter Bioassay. ES2 cells (ATCC CRL-1978) endogenously expressing human PD-L1 were co-incubated with effector cells (engineered Jurkat cells stably expressing human Fc.gamma.RIIIa receptor--V158, Promega) that produce luciferase in a concentration-dependent manner in the presence of an ADCC-enabled antibody. The soluble luciferase activity is assessed by measuring the luminescence produced as the luciferase transformed a luminogenic substrate into a luminescent product.
[1670] Immediately prior to the assay, target cells (ES2) were centrifuged and resuspended in RPMI 1640+10% Ultra low IgG FBS (Thermo Fisher Scientific) and plated at 30,000 cells/well (10 .mu.L/well) in 384-well white bottom plates. Jurkat NFAT luciferase reporter (effector) cells were resuspended in RPMI 1640+10% Ultra low IgG FBS and added to the target cells at 10,000 cells per well (10 .mu.L/well). Eleven three-fold serial dilutions of antibodies were prepared from 2.2 nM in RPMI 1640+10% Ultra low IgG FBS and added to the target cells (10 .mu.L/well). The plates were incubated overnight at 37.degree. C., 5% CO.sub.2, after which a luminogenic BioGlo substrate was added directly to the wells (30 .mu.L/well) and luminescence quantified on an Envision (Perkin Elmer) plate reader.
[1671] Relative light unit (RLU) values from the raw data (Envision reads) were normalised to `Fold of induction using the following equation:
fold of induction = ( RLU ( induced ) - RLU ( background ) ) RLU ( background ) Equation 10 ##EQU00005##
[1672] Data was plotted in GraphPad Prism, using a 4-parameter logistic fit, and a representative experiment is shown in FIG. 34. Results are summarised in Table 20. All antibodies tested induced luminescence, suggesting that all have the capacity to induce killing of target cells by ADCC. Although EC.sub.50 values are generally similar, 416E01 induces a highest maximum level of luminescence.
TABLE-US-00023 TABLE 20 Summary of data from reporter cell assay Antibody EC.sub.50 (pM) max fold induction name n =1 n = 2 n = 3 n = 1 n = 2 n = 3 1D05 24.14 21.53 34.71 13.70 16.69 25.04 84G09 24.95 15.30 25.90 16.33 22.21 33.82 413G05 21.92 20.87 27.52 13.22 16.41 23.45 414B06 19.08 15.02 24.24 10.94 20.15 31.51 416E01 79.98 10.58 22.86 32.11 33.76 60.16 benchmark 2 24.81 10.56 24.93 19.55 31.88 43.23
Example 30: Binding of Lead Antibodies to Cell-Expressed Cynomolgus PD-L1
[1673] CHO-S cells transfected with cynomolgus PD-L1 were diluted in FACS buffer (PBS+1% w/v BSA+0.1% w/v sodium azide) and were distributed to a 96-well V-bottom plate (Greiner) at a density of 1.times.10.sup.5 cells per well. Antibody titrations were prepared from 133 nM working concentration as a 1/3 dilution series in FACS buffer. Plates were centrifuged at 300.times.g for 3 minutes and supernatant aspirated. 50 .mu.L per well of antibody titrations were added to cells and incubated at 4.degree. C. for 1 hour. Cells were washed with 150 .mu.L of PBS and centrifuged at 300 g for 3 minutes. Supernatant was aspirated and 150 .mu.L PBS added per well. This wash step was repeated. Presence of bound antibody was detected by addition of 50 .mu.L per well of anti-Human IgG AlexaFluor 647 (Jackson ImmunoResearch) diluted 1/500 in FACS buffer. Cells were incubated for 1 hour at 4.degree. C. in the dark. Cells were washed as previously described. To fix cells, 50 .mu.L per well of 4% v/v paraformaldehyde was added and cells incubated for 20 minutes at 4.degree. C., cells were pelleted by centrifugation at 300.times.g and the plates resuspended in 75 .mu.L PBS. Geometric mean was measured by flow cytometry using a Beckman Coulter CytoFLEX instrument. Alexa Fluor 647 was excited by a 637 nm laser and detected in the Red channel with a 660/20 bandpass filter. Data was analysed using FlowJo software and is shown in FIG. 35. All antibodies bind to cynomolgus PD-L1 expressed on cells.
Example 31: Binding to CHO-Expressed hPD-L1 and Neutralisation of hPD-L1 Binding to PD-1 and CD80
[1674] CHO cells untransfected (referred to as WT) or transfected with, hPD-L1 expressing recombinant human PD-L1 were diluted in FACS buffer (PBS 1% BSA 0.1% sodium azide) and distributed to three 96-well, V-bottom plate (Greiner) at a density of 1.times.10.sup.5 cells per well. Cells are washed with 150 .mu.L PBS and centrifuged at 300 g for 3 minutes. Supernatant is aspirated and 150 .mu.L PBS added. This wash step is repeated.
[1675] To plate 1 (PD-L1 binding), lead antibody, reference antibody or control antibody titrations were prepared from 150 nM working concentration as a 1/3 dilution series in FACS buffer. 50 .mu.L of antibody diluted in FACS buffer is added to the washed cells and incubated at 4.degree. C. for 60 minutes. 150 .mu.L FACS buffer is added and cells washed as described above. To detect anti-PD-L1 antibody binding, anti-human PE (Jackson ImmunoResearch) is diluted 1/500 in FACS buffer and 50 .mu.L of this mixture added to cells. Cells are incubated 4.degree. C. for 60 minutes. Cells are washed twice with 150 .mu.L FACS buffer, centrifuging at 300 g for 3 minutes after each wash step and aspirating supernatant. Cells are fixed by addition of 100 .mu.L 4% paraformaldehyde and 30 mins at 4.degree. C. Cells are washed once as above and resuspended in 100 .mu.L FACS buffer for analysis. PE signal intensity (geometric mean) is measured by flow cytometry using a Beckman Coulter Cytoflex instrument. Data is plotted as geometric mean values without further calculation.
[1676] To plate 2 (PD-1 neutralisation), biotinylated human PD-1-Fc (in-house expressed, Seq ID No:6) were prepared as a titration from 1 .mu.M final assay concentration (FAC), 1/2 dilution series in FACS buffer. Lead antibody, reference antibody or control antibody titrations were prepared from 300 nM working concentration, 150 nM FAC, as a 1/3 dilution series in FACS buffer. Biotinylated PD-1 were diluted in FACS buffer to 60 nM working concentration, 30 nM FAC. 25 .mu.L PD-1 and 25 .mu.L antibody solution (or 50 .mu.L of PD1 titration) were added to cells and incubated at 4.degree. C. for 1 hour. Biotinylation is performed in-house using Lightning Link conjugation kit (Innova Biosciences) according to manufacturer's instructions. 150 .mu.L FACS buffer is added and cells washed as described above. To detect biotinylated PD-1, Streptavidin-Alexa Fluor 647 (AF647, Jackson ImmunoResearch) is diluted 1/500 in FACS buffer and 50 .mu.L of this mixture added to cells. Cells are incubated at 4.degree. C. for 60 minutes. Cells are washed twice with 150 .mu.L FACS buffer, centrifuging at 300 g for 3 minutes after each wash step and aspirating supernatant. Cells are fixed, washed and resuspended for analysis as above. APC signal intensity (geometric mean) is measured by flow cytometry using a Beckman Coulter CYTOFLEX instrument. Data is plotted as percentage of receptor binding.
[1677] To plate 3 (CD80 neutralisation) Biotinylated human CD80 (Fc tagged, R&D Systems, 140-B1) were prepared as a titration from 1 .mu.M final assay concentration (FAC), 1/2 dilution series in FACS buffer. Lead antibody, reference antibody or control antibody titrations were prepared from 300 nM working concentration, 150 nM FAC, as a 1/3 dilution series in FACS buffer. Biotinylated CD80 were diluted in FACS buffer to 60 nM working concentration, 30 nM FAC. 25 .mu.L CD80 and 25 .mu.L antibody solution (or 50 .mu.L of CD80 titration) were added to cells and incubated at 4.degree. C. for 1 hour. All other steps are performed as per plate 2.
Equation 11 Percentage of receptor binding ( flow cytometry ) Based on geometric mean fluorescence % of specific binding = sample value - non - specific binding total binding - non - specific binding .times. 100 Total binding = biotinylated PD - 1 or CD 80 only ( Hybrid Control at 150 nM FAC ) Non - specific binding = no PDL 1 binding , ( benchmark 2 at 150 nM FAC ) ##EQU00006##
[1678] Results are shown in FIG. 36 and in Table 21. All lead antibodies neutralise interactions of PD-L1 with both PD-1 and CD80.
TABLE-US-00024 TABLE 21 Summary of neutralisation of PD-1 and CD80 interactions with PD-L1 PD-1 IC.sub.50 (nM) CD80 IC.sub.50 (nM) Antibody name n = 1 n = 2 n = 1 n = 2 1D05 1.27 1.46 1.15 1.46 84G09 0.89 1.41 1.11 1.41 413G05 1.07 1.17 1.09 1.17 414B06 0.95 1.22 1.26 1.22 416E01 0.81 0.89 1.04 0.89 benchmark 2 1.36 1.46 1.27 1.46
Example 32: Effector Function of Lead Antibodies in a Primary NK Cell ADCC Assay
[1679] Activity of antibodies to kill PD-L1 expressing target cells via ADCC (antibody-dependent cell-mediated cytotoxicity) is measured by DELFIA cytotoxicity assay (Perkin Elmer) using human primary NK cells as effectors and ES2 as PD-L1.sup.+ target cells.
[1680] This method is based on loading target cells with an acetoxymethyl ester of fluorescence enhancing ligand (BATDA) which quickly penetrates the cell membrane. Within the cell the ester bonds are hydrolysed to form a hydrophilic ligand (TDA) which no longer passes the membrane. After cytolysis the ligand is released and can be detected by addition of Europium which forms with the BATDA a highly fluorescent and stable chelate (EuTDA). The measured signal correlates directly with the degree of cell lysis.
[1681] ES2 cells are resuspended at 10.sup.6/mL in assay medium (RPMI+10% ultra-low IgG FBS, from Gibco) and loaded with 5 .mu.L/mL of BATDA reagent (Perkin Elmer) for 30 min at 37.degree. C. Cells were then washed 3 times with 50 mL PBS (300.times.g for 5 min) and resuspended at 8.times.10.sup.5/mL in assay medium supplemented with, 2 mM Probenecid (Life technologies) to reduce BATDA spontaneous release from the cells. Supernatant from ES2 cells immediately after final resuspension in assay medium is used as background control.
[1682] Seven serial three-fold dilutions of PD-L1 antibodies and isotype controls are prepared in assay media+2 mM Probenecid from 4 .mu.g/mL (4.times. final concentration). NK cells are negatively isolated from fresh PBMC using Human NK Cell Isolation Kit (Miltenyi Biotec) as per manufacturer's instructions and resuspended at 4.times.10.sup.6/mL in assay medium+2 mM Probenecid. 50 .mu.L of diluted Ab, 50 .mu.L of BATDA loaded target cells, 50 .mu.L of NK cells and 50 .mu.L of assay medium+2 mM Probenecid (final volume of 200 .mu.L/well) are added in each well to give an effector: target ratio of 5:1. Wells containing ES cells only or ES2 cells+DELFIA lysis buffer (Perkin Elmer) are used to determine spontaneous and maximum release, respectively.
[1683] Cells are incubated at 37.degree. C., 5% CO.sub.2 for 4 hours before centrifugation of plates for 5 min at 500.times.g, and transfer of 50 .mu.L of cell-free supernatant into a DELFIA microtitration Plates (Perkin Elmer). 200 .mu.L of DELFIA Europium solution (Perkin Elmer) was added to the supernatants and incubated for 15 min at Room Temperature. Fluorescent signal was then quantified with an EnVision plate reader (PerkinElmer).
[1684] Background counts are subtracted from all experimental counts. Specific release is calculated according to the following equation:
% specific release = ( experimental release - spontaneous release ) ( maximum release - spontaneous release ) .times. 100 Equation 12 ##EQU00007##
INCORPORATION BY REFERENCE
[1685] All references cited herein, including patents, patent applications, papers, text books and the like, and the references cited therein, to the extent that they are not already, are hereby incorporated herein by reference in their entirety.
EQUIVALENTS
[1686] The foregoing written specification is considered to be sufficient to enable one skilled in the art to practice the invention. The foregoing description and Examples detail certain preferred embodiments of the invention. It will be appreciated, however, that the invention may be practiced in many ways and the invention should be construed in accordance with the appended claims and any equivalents thereof.
[1687] It will be understood that particular configurations, aspects, examples, clauses and embodiments described herein are shown by way of illustration and not as limitations of the invention. Any part of this disclosure may be read in combination with any other part of the disclosure, unless otherwise apparent from the context.
TABLE-US-00025 SEOUENCE LISTING SEQ ID NO: Name Description Sequence 1 Human PD-L1 NCBI number: NP_054862.1 MRIFAVFIFMTYWHLLNAFTVTVPKDLYVVEYGSNMTIECKFPVEKQLDLAA (ECD highlighted in BOLD, LIVYWEMEDKNIIQFVHGEEDLKVQHSSYRQRARLLKDQLSLGNAALQI cytoplasmic domain underlined) TDVKLQDAGVYRCMISYGGADYKRITVINNAPYNKINQRILVVDPVTSE HELTCQAEGYPKAEVIWTSSDHQVLSGIMTTNSKREEKLFNVTSTLRIN TTTNEIFYCTFRRLDPEENHTAELVIPELPLAHPPNERTHLVILGAILLCLGVA LTFIFRLRKGRMMDVKKCGIQDTNSKKQSDTHLEET 2 Cyno PD-L1 NCBI number: XP_014973154.1 MGWSCIILFLVATATGVHSMFTVTVPKDLYVVEYGSNMTIECKFPVEKQLDL (ECD highlighted in BOLD) TSLIVYWEMEDKNIIQFVHGEEDLINQHSNYRQRAQLLKDQLSLGNAAL RITDVLQDAGVYRCMISYGGADYKRTTVKVNAPYNKINQRILVVDPVT SEHELTCQAEGYPKAEVIWTSSDHQVLSGKTTTTNSKREEKLLNYTSTLR INTTANEIFYCIFRRLDPEENHTAELVIPELPLALPPNERT 3 Human PD-L1 His Human PD-L1 ECD with C- MRIFAVFIFMTYWHLLNAFTVTVPKDLYVVEYGSNMTIECKFPVEKQLDLAALIVYW terminal His tag EMEDKNIIQFVHGEEDLKVQHSSYRQRARLLKDQLSLGNAALQITDVKLQDAGVYR CMISYGGADYKRITVKVNAPYNKINQRILVVDPVTSEHELTCQAEGYPKAEVIWTSS DHQVLSGKTTTTNSKREEKLFNVTSTLRINTTTNEIFYCTFRRLDPEENHTAELVIPE LPLAHPPNERTHHHHHH 4 Human PD-L1 Fc Human PD-L1 ECD with C-term MRIFAVFIFMTYWHLLNAFTVTVPKDLYVVEYGSNMTIECKFPVEKQLDLAALIVYW Fc fusion (in bold) EMEDKNITQFVHGEEDLKVQHSSYRQRARLLKDQLSLGNAALQITDVKLQDAGVYR CMISYGGADYKRITVINNAPYNKINQRILWDPVTSEHELTCQAEGYPKAEVIWTSS DHQVLSGKTTTTNSKREEKLFNVISTLRINTTTNEIFYCTFRRLDPEENHTAELVIPE LPLAHPPNERTIEGREPKSCDKTHTCPPCPAPELLGGPSVFLFPPICPKDTLM ISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYT LPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLD SDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 5 Cyno PD-L1 FLAG Cynomolgus PD-L1 ECD with N- MGWSCIILFLVATATGVHSMFVTVPKDLYVVEYGSNMTIECKFPVEKQLDLTSLIV term FLAG tag YWEMEDKNITQFVHGEEDLKVQHSNYRQRAQLLKDQLSLGNAALRITDVKLQDAG VYRCMISYGGADYKRITVINNAPYNICENQRILVVDPVTSEHELTCQAEGYPKAEVI WTSSDHQVLSGMTTNSKREEKLLNVTSTLRINTTANEIFYCIFRRLDPEENHTAE LVIPELPLALPPNERTDYKDDDDK 6 Human PD-1 Fc Human PD-1 full length MGWSCIILFLVATATGVHSLDSPDRPWNPPTFSPALLVVTEGDNATFTCSFSNTSE sequence derived from cDNA as SFVLNWYRMSPSNQTDKLAAFPEDRSQPGQDCRFRVTQLPNGRDFHMSVVRARR human Fc fusion NDSGTYLCGAISLAPKAQIKESLRAELRVTERRAEVPTAHPSPSPRPAGQKLENLY FQGIEGRPIDEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRT PEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRANS VLTVLHQDWLNGKEYKKVSNKALPAPIEKTISKAKGQPREPQVYTLPP SRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDG SFFLYSKLTVDKSRWQQGNFSCSVMHEALHNHYTQKSLSLSP 7 84G09-CDRH1 Amino acid sequence of CDRH1 GFTFDDYA (IMGT) of 84G09 using IMGT 8 84G09-CDRH2 Amino acid sequence of CDRH2 ISWKSNII (IMGT) of 84G09 using IMGT 9 84G09-CDRH3 Amino acid sequence of CDRH3 ARDITGSGSYGWFDP (IMGT) of 84G09 using IMGT 10 84G09-CDRH1 Amino acid sequence of CDRH1 DYAMH (Kabat) of 84G09 using Kabat 11 84G09-CDRH2 Amino acid sequence of CDRH2 GISWKSNIIGYADSVKG (Kabat) of 84G09 using Kabat 12 84G09-CDRH3 Amino acid sequence of CDRH3 DITGSGSYGWFDP (Kabat) of 84G09 using Kabat 13 84G09-Heavy chain Amino acid sequence of V.sub.H of EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQTPGKGLEWVSGISWK variable region 84G09 (mutations from germline SNIIGYADSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCARDITGSGSYGWFD are shown in bold letters) PWGQGTLVTVSS 14 84G09-Heavy chain Nucleic acid sequence of V.sub.H of CAaGAAAAAGCTTGCCGCCACCATGGAGTTTGGGCTGAGCTGGATTTTCCTTTT variable reg ion 84G09 GGCTATTTTAAAAGGTGTCCAGTGTGAAGTACAATTGGTGGAGTCCGGGGGAG GCTTGGTACAGCCTGGCAGGTCCCTGAGACTCTCCTGTGCAGCCTCTGGATTCA CCTTTGATGATTATGCCATGCACTGGGTCCGACAAACTCCAGGGAAGGGCCTGG AGTGGGTCTCAGGTATAAGTTGGAAGAGTAATATCATAGGCTATGCGGACTCTG TGAAGGGCCGATTCACCATCTCCAGAGACAACGCCAAGAACTCCCTGTATCTGC AAATGAACAGTCTGAGAGCTGAGGACACGGCCTTGTATTATTGTGCAAGAGATA TAACGGGTTCGGGGAGTTATGGCTGGTTCGACCCCTGGGGCCAGGGAACCCTG GTCACCGTCTCCTCAGCCAAAACGACACCCCCATCTGTCTATCCACTGGCCCCTG AATCTGCTAAAACTCAGCCTCCG 15 84G09-full heavy Amino acid sequence of 84G09 EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQTPGKGLEWVSGISWK chain sequence heavy chain (mutations from SNIIGYADSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCARDITGSGSYGWFD germline are shown in bold PWGQGTLVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSG letters) ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESK YGPPCPPCPAPEFEGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNW YVDGVEVHNAKTKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIE KTISKAKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPEN NYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSL GK 16 84G09-full heavy Nucleic acid sequence of 84G09 GAAGTGCAGCTGGTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCAGATCCCT chain sequence heavy chain GAGACTGTCTTGTGCCGCCTCCGGCTTCACCTTCGACGACTACGCTATGCACTG GGTGCGACAGACCCCTGGCAAGGGCCTGGAATGGGTGTCCGGCATCTCCTGGA AGTCCAACATCATCGGCTACGCCGACTCCGTGAAGGGCCGGTTCACCATCTCCC GGGACAACGCCAAGAACTCCCTGTACCTGCAGATGAACAGCCTGCGGGCCGAGG ACACCGCCCTGTACTACTGCGCCAGAGACATCACCGGCTCCGGCTCCTACGGAT GGTTCGATCCTTGGGGCCAGGGCACCCTCGTGACCGTGTCCTCTGCCAGCACCA AGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAA CAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGT CCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGC AGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCT GGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGT GGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTG TCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCC CAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGA TGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGA AGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCG GGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGT ACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCT CCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCA GGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCT ACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACT ACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAA GCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGT GATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGCCC CGGCAAG 17 84G09-CDRL1 Amino acid sequence of CDRL1 QSISSY (IMGT) of 84G09 using IMGT 18 84G09-CDRL2 Amino acid sequence of CDRL2 VAS (IMGT) of 84G09 using IMGT 19 84G09-CDRL3 Amino acid sequence of CDRL3 QQSYSNPIT (IMGT) of 84G09 using IMGT 20 84G09-CDRL1 Amino acid sequence of CDRL1 RASQSISSYLN (Kabat) of 84G09 using Kabat 21 84G09-CDRL2 Amino acid sequence of CDRL2 VASSLQS (Kabat) of 84G09 using Kabat 22 84G09-CDRL3 Amino acid sequence of CDRL3 QQSYSNPIT (Kabat) of 84G09 using Kabat 23 84G09-Light chain Amino acid sequence of V.sub.L of DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKPLPNASSLQS variable region 84G09 GVPSSFSGSGSGTDFTLTISSLQPEDFATYYCQQSYSNPITFGQGTRLEIK 24 84G09-Light chain Nucleic acid sequence of V.sub.L of GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGA variable region 84G09 GTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCCCCTGATCTATGTTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGTTTCAGTGGCAGTGGATCTGGGACAGATTTCACT CTCACCATCAGCAGTCTGCAACCTGAAGATTTTGCAACTTACTACTGTCAACAGA GTTACAGTAATCCGATCACCTTCGGCCAAGGGACACGACTGGAGATCAAA 25 84G09-full light Amino acid sequence of 84G09 DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKPLIYVASSLQS chain sequence light chain GVPSSFSGSGSGTDFTLTISSLQPEDFATYYCQQSYSNPITFGQGTRLEIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 26 84G09-full light Nucleic acid sequence of 84G09 GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGA chain sequence light chain GTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCCCCTGATCTATGTTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGTTTCAGTGGCAGTGGATCTGGGACAGATTTCACT CTCACCATCAGCAGTCTGCAACCTGAAGATTTTGCAACTTACTACTGTCAACAGA GTTACAGTAATCCGATCACCTTCGGCCAAGGGACACGACTGGAGATCAAACGTA CGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGT CCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCA AGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCG TGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCC TGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACC AGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 27 1D05-CDRH1 Amino acid sequence of CDRH1 GFTFDDYA (IMGT) of 1D05 using IMGT 28 1D05-CDRH2 Amino acid sequence of CDRH2 ISWIRTGI (IMGT) of 1D05 using IMGT 29 1D05-CDRH3 Amino acid sequence of CDRH3 AKDMKGSGTYGGWFDT (IMGT) of 1D05 using IMGT 30 1D05-CDRH1 Amino acid sequence of CDRH1 DYAMH (Kabat) of 1D05 using Kabat 31 1D05-CDRH2 Amino acid sequence of CDRH2 GISVVIRTGIGYADSVKG (Kabat) of 1D05 using Kabat 32 1D05-CDRH3 Amino acid sequence of CDRH3 DMKGSGTYGGWFDT (Kabat) of 1D05 using Kabat 33 1D05-Heavy chain Amino acid sequence of V.sub.H of EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQBPGKGLEWVSGISWB variable region 1D05 (mutations from germline BIGYADSVKGRFTIBRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGG are shown in bold letters) WFDTWGQGTLVTVSS 34 1D05-Heavy chain Nucleic acid sequence of V.sub.H of AAGCTTGCCGCCACCATGGAGTTTGGGCTGAGCTGGATTTTCCTTTTGGCTATT variable region 1D05 TTAAAAGGTGTCCAGTGTGAAGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGT GCAGCCTGGCAGGTCCCTGAGACTCTCCTGTGCAGCCTCTGGATTCACCTTTGA TGATTATGCCATGCACTGGGTCCGGCAAGTTCCAGGGAAGGGCCTGGAATGGG TCTCAGGCATTAGTTGGATTCGTACTGGCATAGGCTATGCGGACTCTGTGAAGG GCCGATTCACCATTTTCAGAGACAACGCCAAGAATTCCCTGTATCTGCAAATGAA CAGTCTGAGAGCTGAGGACACGGCCTTGTATTACTGTGCAAAAGATATGAAGGG TTCGGGGACTTATGGGGGGTGGTTCGACACCTGGGGCCAGGGAACCCTGGTCA CCGTCTCCTCAGCCAAAACAACAGCCCCATCGGTCTATCCACTGGCCCCTGC 35 1D05-full heavy Amino acid sequence of 1D05 EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQVPGKGLEWVSGISWI chain sequence heavy chain RTGIGYADSVKGRFTIFRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGGW FDTWGQGTLVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWN SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRV ESKYGPPCPPCPAPEFEGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQF NWYVDGVEVHNAKTKPREEQFNSTYRVVSLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNG QPENNYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKS LSLSLGK 36 1D05-full heavy Nucleic acid sequence of 1D05 GAAGTGCAGCTGGTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCAGATCCCT chain sequence heavy chain GAGACTGTCTTGTGCCGCCTCCGGCTTCACCTTCGACGACTACGCTATGCACTG GGTGCGACAGGTGCCAGGCAAGGGCCTGGAATGGGTGTCCGGCATCTCTTGGA TCCGGACCGGCATCGGCTACGCCGACTCTGTGAAGGGCCGGTTCACCATCTTCC
GGGACAACGCCAAGAACTCCCTGTACCTGCAGATGAACAGCCTGCGGGCCGAGG ACACCGCCCTGTACTACTGCGCCAAGGACATGAAGGGCTCCGGCACCTACGGCG GATGGTTCGATACTTGGGGCCAGGGCACCCTCGTGACCGTGTCCTCTGCCAGCA CCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCG GAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCG TGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGC TGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCT CTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAA GGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCC TTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAA GCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGT GGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGT GGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTA CCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAG AGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCA TCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTA GCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCT TCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACA ACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAG CAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTC CGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAG CCCCGGCAAG 37 1D05-CDRL1 (IMGT) Amino acid sequence of CDRL1 QSISSY of 1D05 using IMGT 38 1D05-CDRL2 (IMGT) Amino acid sequence of CDRL2 VAS of 1D05 using IMGT 39 1D05-CDRL3 (IMGT) Amino acid sequence of CDRL3 QQSYSTPIT of 1D05 using IMGT 40 1D05-CDRL1 (Kabat) Amino acid sequence of CDRL1 RASQSISSYLN of 1D05 using Kabat 41 1D05-CDRL2 (Kabat) Amino acid sequence of CDRL2 VASSLQS of 1D05 using Kabat 42 1D05-CDRL3 (Kabat) Amino acid sequence of CDRL3 QQSYSTPIT of 1D05 using Kabat 43 1D05-Light chain Amino acid sequence of V.sub.L of DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYVASSLQS variable region 1D05 (mutations from germline GVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQSYSTPITFGQGTRLEIK are shown in bold letters) 44 1D05-Light chain Nucleic acid sequence of V.sub.L of AAAGCTTGCCGCCACCATGAGGCTCCCTGCTCAGCTTCTGGGGCTCCTGCTACT variable region 1D05 CTGGCTCCGAGGTGCCAGATGTGACATCCAGATGACCCAGTCTCCATCCTCCCT GTCTGCATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCAT TAGCAGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAACTCCT GATCTATGTTGCATCCAGTTTGCAAAGTGGGGTCCCATCAAGGTTCAGTGGCAG TGGATCTGGGACAGATTTCACTCTCACTATCAGCAGTCTGCAACCTGAAGATTTT GCAACTTACTACTGTCAACAGAGTTACAGTACCCCGATCACCTTCGGCCAAGGG ACACGTCTGGAGATCAAACGTACGGATGCTGCACCAACT 45 1D05-full light chain Amino acid sequence of 1D05 DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYVASSLQS sequence light chain GVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQSYSTPITFGQGTRLEIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 46 1D05-full light chain Nucleic acid sequence of 1D05 GACATCCAGATGACCCAGTCCCCCTCCAGCCTGTCTGCTTCCGTGGGCGACAGA sequence light chain GTGACCATCACCIGTCGGGCCTCCCAGTCCATCTCCTCCTACCTGAACTGGTATC AGCAGAAGCCCGGCAAGGCCCCCAAGCTGCTGATCTACGTGGCCAGCTCTCTGC AGTCCGGCGTGCCCTCTAGATTCTCCGGCTCTGGCTCTGGCACCGACTTTACCC TGACCATCAGCTCCCTGCAGCCCGAGGACTTCGCCACCTACTACTGCCAGCAGT CCTACTCCACCCCTATCACCTTCGGCCAGGGCACCCGGCTGGAAATCAAACGTA CGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGT CCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCA AGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCG TGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCC TGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACC AGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 47 Mutated 1D05-HC Amino acid sequence of 1D05 EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWI mutant 1 heavy chain with V to A back- RTGIGYADSVKGRFTIFRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGGW mutation in framework region to FDTWGQGTLVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWN germline highlighted with IgG1 SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRV disabled (LAGA) constant region ESKYGPPCPPCPAPELAGAPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQF NWYVDGVEVHNAKTKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNG QPENNYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKS LSLSLGK 48 Mutated 1D05-HC Amino acid sequence of 1D05 EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQVPGKGLEWVSGISWI mutant 2 heavy chain with F to S back- RTGIGYADSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGGW mutation in framework region to FDTWGQGTLVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWN germline highlighted with IgG1 SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRV disabled (LAGA) constant region ESKYGPPCPPCPAPELAGAPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQF NWYVDGVEVHNAKTKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISMKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNG QPENNYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKS LSLSLGK 49 Mutated 1D05-HC Amino acid sequence of 1D05 EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQVPGKGLEWVSGISWI mutant 3 heavy chain with ELLG to -PVA RTGIGYADSVKGRFTIFRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGGW back-mutation in constant FDTWGQGTLVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWN region to germline highlighted SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRV ESKYGPPCPPCPAP- PVAGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAK TKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISMKGQPR EPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDS DGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK 50 Mutated 1D05-LC Amino acid sequence of 1D05 DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQS mutant 1 kappa light chain with V to A GVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQSYSTPITFGQGTRLEIKRTVAAP back-mutation in CDRL2 to SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK germline highlighted DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 51 Mutated 1D05-LC Amino acid sequence of 1D05 DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLFIYVASSLQS mutant 2 kappa light chain with L to F GVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQSYSTPITFGQGTRLEIKRTVAAP back-mutation in framework to SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK germline highlighted DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVYKSFNRGEC 52 411B08-CDRH1 Amino acid sequence of CDRH1 GFTFSSYW (IMGT) of 411B08 using IMGT 53 411B08-CDRH2 Amino acid sequence of CDRH2 IKEDGSEK (IMGT) of 411B08 using IMGT 54 411B08-CDRH3 Amino acid sequence of CDRH3 ARNRLYSDFLDN (IMGT) of 411B08 using IMGT 55 411B08-CDRH1 Amino acid sequence of CDRH1 SYWMS (Kabat) of 411B08 using Kabat 56 411B08-CDRH2 Amino acid sequence of CDRH2 NIKEDGSEKYYVDSVKG (Kabat) of 411B08 using Kabat 57 411B08-CDRH3 Amino acid sequence of CDRH3 NRLYSDFLDN (Kabat) of 411B08 using Kabat 58 411B08-Heavy chain Amino acid sequence of V.sub.H of EVQLVESGGGLVQPGGSLRLSCAASGFTFSSYWMSWVRQAPGKGLEWVANIKED variable region 411B08 GSEKMDSVKGRFTISRDNAKNSLYLQMNSLRAEDTSVYYCARNRLYSDFLDNW GQGTLVTVSS 59 411B08-Heavy chain Nucleic acid sequence of V.sub.H of GAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT variable region 411B08 GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGTAGCTATTGGATGAGTTG GGTCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTGGCCAACATCAAAGAAG ATGGAAGTGAGAAATACTATGTCGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGTCTGTGTATTACTGTGCGAGAAATCGACTCTACAGTGACTTCCTTGACAA CTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAG 60 411B08-full heavy Amino acid sequence of 411B08 EVQLVESGGGLVQPGGSLRLSCAASGFTFSSYWMSWVRQAPGKGLEWVANIKED chain sequence heavy chain GSEKYYVDSVKGRFTISRDNAKNSLYLQMNSLRAEDTSVYYCARNRLYSDFLDNW GQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGAL TSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSC DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAP IEKTISMKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK 61 411B08-full heavy Nucleic acid sequence of 411B08 GAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT chain sequence heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGTAGCTATTGGATGAGTTG GGTCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTGGCCAACATCAAAGAAG ATGGAAGTGAGAAATACTATGTCGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGTCTGTGTATTACTGTGCGAGAAATCGACTCTACAGTGACTTCCTTGACAA CTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCTC TGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTCT GGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTGGAACTC TGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCGG CCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCCA GACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGAA GGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCCC TGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACAC CCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCCA CGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACAA CGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGTC CGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCAA GGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCAA GGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGCT GACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCGA TATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCAC CCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAGT GGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATGCACGA GGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGCAAG 62 411B08-CDRL1 Amino acid sequence of CDRL1 QGVSSW (IMGT) of 411B08 using IMGT 63 411B08-CDRL2 Amino acid sequence of CDRL2 GAS (IMGT) of 411B08 using IMGT 64 411B08-CDRL3 Amino acid sequence of CDRL3 QQANSIPFT (IMGT) of 411B08 using IMGT 65 411B08-CDRL1 Amino acid sequence of CDRL1 RASQGVSSWLA (Kabat) of 411B08 using Kabat 66 411B08-CDRL2 Amino acid sequence of CDRL2 GASSLQS (Kabat) of 411B08 using Kabat 67 411B08-CDRL3 Amino acid sequence of CDRL3 QQANSIPFT (Kabat) of 411B08 using Kabat 68 411B08-Light chain Amino acid sequence of V.sub.L of DIQMTQSPSSVSASVGDRVTITCRASQGVSSWLAVVYQQKSGKAPKLLIYGASSLQ variable region 411B08 SGVPSRFSGSGSGTEFILTISSLQPEDFATYYCQQANSIPFTFGPGTKVDIK 69 411B08-Light chain Nucleic acid sequence of V.sub.L of GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTCGGAGACAGA variable region 411B08 GTCACCATCACTTGTCGGGCGAGTCAGGGTGTTAGCAGCTGGTTAGCCTGGTAT CAGCAGAAATCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGATTCAGCGGCAGTGGATCTGGGACAGAGTICATT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTATCCCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAAC 70 411B08-full light Amino acid sequence of 411B08 DIQMTQSPSSVSASVGDRVTITCRASQGVSSWLAWYQQKSGKAPKLLIYGASSLQ chain sequence light chain SGVPSRFSGSGSGTEFILTISSLQPEDFATYYCQQANSIPFTFGPGTKVDIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAINQWKVDNALQSGNSQESVTEQDSK
DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 71 411B08-full light Nucleic acid sequence of 411B08 GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTCGGAGACAGA chain sequence light chain GTCACCATCACTTGTCGGGCGAGTCAGGGTGTTAGCAGCTGGTTAGCCTGGTAT CAGCAGAAATCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGATTCAGCGGCAGTGGATCTGGGACAGAGTTCATT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTATCCCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAACGTAC GGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGTC CGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCAA GGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCGT GACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCCT GTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACCA GGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 72 411C04-CDRH1 Amino acid sequence of CDRH1 GFTFSSYW (IMGT) of 411C04 using IMGT 73 411C04-CDRH2 Amino acid sequence of CDRH2 IKEDGSEK (IMGT) of 411C04 using IMGT 74 411C04-CDRH3 Amino acid sequence of CDRH3 ARVRLYSDFLDY (IMGT) of 411C04 using IMGT 75 411C04-CDRH1 Amino acid sequence of CDRH1 SYWMS (Kabat) of 411C04 using Kabat 76 411C04-CDRH2 Amino acid sequence of CDRH2 NIKEDGSEKYYVDSLKG (Kabat) of 411C04 using Kabat 77 411C04-CDRH3 Amino acid sequence of CDRH3 VRLYSDFLDY (Kabat) of 411C04 using Kabat 78 411C04-Heavy chain Amino acid sequence of V.sub.H of EVQLVDSGGGLVQPGGSLRLSCAASGFTFSSYWMSWVRQAPGKGLEWVANIKED variable region 411C04 GSEKYYVDSLKGRFTISRDNAKNSLYLQMNSLRAEDTSVYYCARVRLYSDFLDYWG QGTLVTVSS 79 411C04-Heavy chain Nucleic acid sequence of V.sub.H of GAGGTGCAGCTGGTGGACTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT variable region 411C04 GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGTAGCTATTGGATGAGTTG GGTCCGCCAGGCTCCAGGAAAGGGGCTGGAGTGGGTGGCCAACATAAAAGAAG ATGGAAGTGAGAAATACTATGTAGACTCTTTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGTCTGTGTATTACTGTGCGAGAGTTCGACTCTACAGTGACTTCCTTGACTA CTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAG 80 411C04-full heavy Amino acid sequence of 411C04 EVQLVDSGGGLVQPGGSLRLSCAASGFTFSSYWMSWVRQAPGKGLEWVANIKED chain sequence heavy chain GSEKYYVDSLKGRFTISRDNAKNSLYLQMNSLRAEDTSVYYCARVRLYSDFLDYWG QGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALT SGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCD KTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRWSVLTVLHQDWLNGKEYKCKVSNKALPAPIE KTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPEN NYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGK 81 411C04-full heavy Nucleic acid sequence of 411C04 GAGGTGCAGCTGGTGGACTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT chain sequence heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGTAGCTATTGGATGAGTTG GGTCCGCCAGGCTCCAGGAAAGGGGCTGGAGTGGGTGGCCAACATAAAAGAAG ATGGAAGTGAGAAATACTATGTAGACTCTTTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGTCTGTGTATTACTGTGCGAGAGTTCGACTCTACAGTGACTTCCTTGACTA CTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCTC TGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTCT GGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTGGAACTC TGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCGG CCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCCA GACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGAA GGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCCC TGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACAC CCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCCA CGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACAA CGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGTC CGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCAA GGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCAA GGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGCT GACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCGA TATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCAC CCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAGT GGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATGCACGA GGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGCAAG 82 411C04-CDRL1 Amino acid sequence of CDRL1 QGVSSW (IMGT) of 411C04 using IMGT 83 411C04-CDRL2 Amino acid sequence of CDRL2 GAS (IMGT) of 411C04 using IMGT 84 411C04-CDRL3 Amino acid sequence of CDRL3 QQANSIPFT (IMGT) of 411C04 using IMGT 85 411C04-CDRL1 Amino acid sequence of CDRL1 RASQGVSSWLA (Kabat) of 411C04 using Kabat 86 411C04-CDRL2 Amino acid sequence of CDRL2 GASSLQS (Kabat) of 411C04 using Kabat 87 411C04-CDRL3 Amino acid sequence of CDRL3 QQANSIPFT (Kabat) of 411C04 using Kabat 88 411C04-Light chain Amino acid sequence of V.sub.L of DIQMTQSPSSVSASVGDRVTITCRASQGVSSWLAWYQQKSGKAPKLLIYGASSLQ variable region 411C04 SGVPSRFSGSGSGTEFILSISSLQPEDFATYYCQQANSIPFTFGPGTKVDIK 89 411C04-Light chain Nucleic acid sequence of V.sub.L of GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTCGGAGACAGA variable region 411C04 GTCACCATCACTTGTCGGGCGAGTCAGGGTGTTAGCAGTTGGTTAGCCTGGTAT CAGCAGAAATCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTGCCTCCAGTTTG CAAAGTGGGGTCCCATCAAGATTCAGCGGCAGTGGATCTGGGACAGAGTTCATT CTCAGCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTATCCCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAAC 90 411C04-full light Amino acid sequence of 411C04 DIQMTQSPSSVSASVGDRVTITCRASQGVSSWLAWYQQKSGKAPKLLIYGASSLQ chain sequence light chain SGVPSRFSGSGSGTEFILSISSLQPEDFATYYCQQANSIPFTFGPGTINDIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 91 411C04-full light Nucleic acid sequence of 411C04 GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTCGGAGACAGA chain sequence light chain GTCACCATCACTTGTCGGGCGAGTCAGGGTGTTAGCAGTTGGTTAGCCTGGTAT CAGCAGAAATCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTGCCTCCAGTTTG CAAAGTGGGGTCCCATCAAGATTCAGCGGCAGTGGATCTGGGACAGAGTTCATT CTCAGCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTATCCCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAACGTAC GGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGTC CGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCAA GGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCGT GACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCCT GTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACCA GGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 92 411D07-CDRH1 Amino acid sequence of CDRH1 GGSIISSDW (IMGT) of 411D07 using IMGT 93 411D07-CDRH2 Amino acid sequence of CDRH2 IFHSGRT (IMGT) of 411D07 using IMGT 94 411D07-CDRH3 Amino acid sequence of CDRH3 ARDGSGSY (IMGT) of 411D07 using IMGT 95 411D07-CDRH1 Amino acid sequence of CDRH1 SSDWWN (Kabat) of 411D07 using Kabat 96 411D07-CDRH2 Amino acid sequence of CDRH2 EIFHSGRTNYNPSLKS (Kabat) of 411D07 using Kabat 97 411D07-CDRH3 Amino acid sequence of CDRH3 DGSGSY (Kabat) of 411D07 using Kabat 98 411D07-Heavy chain Amino acid sequence of V.sub.H of QVQLQESGPGLVKPSGTLSLTCIVSGGSIISSDWWNWVRQPPGKGLEWIGEIFHS variable region 411D07 GRTNYNPSLKSRVTISIDKSKNQFSLRLSSVTAADTAVYYCARDGSGSYWGQGTLV TVSS 99 411D07-Heavy chain Nucleic acid sequence of V.sub.H of CAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGGGACCCT variable region 411D07 GTCCCTCACCTGCATTGTCTCTGGTGGCTCCATCATCAGTAGTGACTGGTGGAA TTGGGTCCGCCAGCCCCCAGGGAAGGGGCTGGAGTGGATTGGAGAAATCTTTC ATAGTGGGAGGACCAACTACAACCCGTCCCTCAAGAGTCGAGTCACCATATCAA TAGACAAGTCCAAGAATCAGTTCTCCCTGAGGCTGAGCTCTGTGACCGCCGCGG ACACGGCCGTGTATTACTGTGCGAGAGATGGTTCGGGGAGTTACTGGGGCCAG GGAACCCTGGTCACCGTCTCCTCAG 100 411D07-full heavy Amino acid sequence of 411D07 QVQLQESGPGLVKPSGTLSLTCIVSGGSIISSDWWNWVRQPPGKGLEWIGEIFHS chain sequence heavy chain GRTNYNPSLKSRVTISIDKSKNQFSLRLSSVTAADTAVYYCARDGSGSYWGQGTLV TVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHT FPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC PPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGV EVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISK AKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTT PPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 101 411D07-full heavy Nucleic acid sequence of CAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGGGACCCT chain sequence 411D07 heavy chain GTCCCTCACCTGCATTGTCTCTGGTGGCTCCATCATCAGTAGTGACTGGTGGAA TTGGGTCCGCCAGCCCCCAGGGAAGGGGCTGGAGTGGATTGGAGAAATCTTTC ATAGTGGGAGGACCAACTACAACCCGTCCCTCAAGAGTCGAGTCACCATATCAA TAGACAAGTCCAAGAATCAGTTCTCCCTGAGGCTGAGCTCTGTGACCGCCGCGG ACACGGCCGTGTATTACTGTGCGAGAGATGGTTCGGGGAGTTACTGGGGCCAG GGAACCCTGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCTCTGTGTTCCCT CTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTCTGGGCTGCCTC GTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCIGGAACTCTGGCGCTCTG ACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCGGCCTGTACTCC CTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCCAGACCTACATC TGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGAAGGTGGAACCC AAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCCCTGAACTGCTG GGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACACCCTGATGATC TCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCCACGAGGACCCT GAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACAACGCCAAGACC AAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGTCCGTGCTGACC GTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCAAGGTGTCCAAC AAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCAAGGGCCAGCCC CGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGCTGACCAAGAAC CAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCGATATCGCCGTG GAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCACCCCCCCTGTG CTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAGTGGACAAGTCC CGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATGCACGAGGCCCTGCAC AACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGCAAG 102 411D07-CDRL1 Amino acid sequence of CDRL1 QSVLYSSNNKNY (IMGT) of 411D07 using IMGT 103 411D07-CDRL2 Amino acid sequence of CDRL2 WAS (IMGT) of 411D07 using IMGT 104 411D07-CDRL3 Amino acid sequence of CDRL3 QQYYSNRS (IMGT) of 411D07 using IMGT 105 411D07-CDRL1 Amino acid sequence of CDRL1 KSSQSVLYSSNNKNYLA (Kabat) of 411D07 using Kabat 106 411D07-CDRL2 Amino acid sequence of CDRL2 WASTRES (Kabat) of 411D07 using Kabat 107 411D07-CDRL3 Amino acid sequence of CDRL3 QQYYSNRS (Kabat) of 411D07 using Kabat 108 411D07-Light chain Amino acid sequence of V.sub.L of DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNNKNYLAWYQQKSGQPPKWYW variable region 411D07 ASTRESGVPDRFSGSGSGTDFTLTISSLQTEDVAVYYCQQYYSNRSFGQGTKLEIK 109 411D07-Light chain Nucleic acid sequence of V.sub.L of GACATCGTGATGACCCAGTCTCCAGACTCCCTGGCTGIGTCTCTGGGCGAGAGG variable region 411D07 GCCACCATCAACTGCAAGTCCAGCCAGAGTGTTTTATACAGCTCCAACAATAAGA ATTACTTAGCTTGGTACCAGCAGAAATCAGGACAGCCTCCTAAGTTGCTCATTTA CTGGGCATCTACCCGGGAATCCGGGGTCCCTGACCGATTCAGTGGCAGCGGGT CTGGGACAGATTTCACTCTCACCATCAGCAGCCTGCAGACTGAAGATGTGGCAG TTTATTACTGTCAGCAATATTATAGTAATCGCAGTTTTGGCCAGGGGACCAAGCT GGAGATCAAAC
110 411D07-full light Amino acid sequence of 411D07 DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNNKNYLAWYQQKSGQPPKWYW chain sequence light chain ASTRESGVPDRFSGSGSGTDFTLTISSLQTEDVAVYYCQQYYSNRSFGQGTKLEIK RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESV TEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 111 411D07-full light Nucleic acid sequence of GACATCGTGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAGG chain sequence 411D07 light chain GCCACCATCAACTGCAAGTCCAGCCAGAGTGTTTTATACAGCTCCAACAATAAGA ATTACTTAGCTTGGTACCAGCAGAAATCAGGACAGCCTCCTAAGTTGCTCATTTA CTGGGCATCTACCCGGGAATCCGGGGTCCCTGACCGATTCAGTGGCAGCGGGT CTGGGACAGATTTCACTCTCACCATCAGCAGCCTGCAGACTGAAGATGTGGCAG TTTATTACTGTCAGCAATATTATAGTAATCGCAGTTTTGGCCAGGGGACCAAGCT GGAGATCAAACGTACGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGA CGAGCAGCTGAAGTCCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTA CCCCCGCGAGGCCAAGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCA ACTCCCAGGAATCCGTGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGT CCTCCACCCTGACCCTGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCT GCGAAGTGACCCACCAGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGG GCGAGTGT 112 385F01-CDRH1 Amino acid sequence of CDRH1 GFTFSSYW (IMGT) of 385F01 using IMGT 113 385F01-CDRH2 Amino acid sequence of CDRH2 IKEDGSEK (IMGT) of 385F01 using IMGT 114 385F01-CDRH3 Amino acid sequence of CDRH3 ARNRLYSDFLDN (IMGT) of 385F01 using IMGT 115 385F01-CDRH1 Amino acid sequence of CDRH1 SYWMS (Kabat) of 385F01 using Kabat 116 385F01-CDRH2 Amino acid sequence of CDRH2 NIKEDGSEKYYVDSVKG (Kabat) of 385F01 using Kabat 117 385F01-CDRH3 Amino acid sequence of CDRH3 NRLYSDFLDN (Kabat) of 385F01 using Kabat 118 385F01-Heavy chain Amino acid sequence of V.sub.H of EVQLVESGGGLVQPGGSLRLSCAASGFTFSSYWMSWVRQAPGKGLEWVANIKED variable region 385F01 GSEKYYVDSVKGRFTISRDNAKNSLYLQMNSLRAEDTSVYYCARNRLYSDFLDNW GQGTLVTVSS 119 385F01-Heavy chain Nucleic acid sequence of V.sub.H of GAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT variable region 385F01 GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGTAGCTATTGGATGAGTTG GGTCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTGGCCAACATCAAAGAAG ATGGAAGTGAGAAATACTATGTCGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGTCTGTGTATTACTGTGCGAGAAATCGACTCTACAGTGACTTCCTTGACAA CTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAG 120 385F01-full heavy Amino acid sequence of 385F01 EVQLVESGGGLVQPGGSLRLSCAASGFTFSSYWMSWVRQAPGKGLEWVANIKED chain sequence heavy chain GSEKYYVDSVKGRFTISRDNAKNSLYLQMNSLRAEDTSVYYCARNRLYSDFLDNW GQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGAL TSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSC DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKOWSNKALPAP IEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYIMPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK 121 385F01-full heavy Nucleic acid sequence of 385F01 GAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT chain sequence heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGTAGCTATTGGATGAGTTG GGTCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTGGCCAACATCAAAGAAG ATGGAAGTGAGAAATACTATGTCGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGTCTGTGTATTACTGTGCGAGAAATCGACTCTACAGTGACTTCCTTGACAA CTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCTC TGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTCT GGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTGGAACTC TGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCGG CCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCCA GACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGAA GGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCCC TGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACAC CCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCCA CGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACAA CGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGTC CGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCAA GGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCAA GGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGCT GACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCGA TATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCAC CCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAGT GGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATGCACGA GGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGCAAG 122 385F01-CDRL1 Amino acid sequence of CDRL1 QGVSSW (IMGT) of 385F01 using IMGT 123 385F01-CDRL2 Amino acid sequence of CDRL2 GAS (IMGT) of 385F01 using IMGT 124 385F01-CDRL3 Amino acid sequence of CDRL3 QQANSIPFT (IMGT) of 385F01 using IMGT 125 385F01-CDRL1 Amino acid sequence of CDRL1 RASQGVSSWLA (Kabat) of 385F01 using Kabat 126 385F01-CDRL2 Amino acid sequence of CDRL2 GASSLQS (Kabat) of 385F01 using Kabat 127 385F01-CDRL3 Amino acid sequence of CDRL3 QQANSIPFT (Kabat) of 385F01 using Kabat 128 385F01-Light chain Amino acid sequence of V.sub.L of DIQMTQSPSSVSASVGDRVTITCRASQGVSSWLAWYQQKSGKAPKLLIYGASSLQ variable region 385F01 SGVPSRFSGSGSGTEFILTISSLQPEDFATYYCQQANSIPFTFGPGTINDIK 129 385F01-Light chain Nucleic acid sequence of V.sub.L of GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTCGGAGACAGA variable region 385F01 GTCACCATCACTTGTCGGGCGAGTCAGGGTGTTAGCAGCTGGTTAGCCTGGTAT CAGCAGAAATCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGATTCAGCGGCAGTGGATCTGGGACAGAGTTCATT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTATCCCATTCACTTTCGGCCCTGGGACCAAAGIGGATATCAAAC 130 385F01-full light Amino acid sequence of 385F01 DIQMTQSPSSVSASVGDRVTITCRASQGVSSWLAWYQQKSGKAPKLLIYGASSLQ chain sequence light chain SGVPSRFSGSGSGTEFILTISSLQPEDFATYYCQQANSIPFTFGPGTKVDIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAINQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 131 385F01-full light Nucleic acid sequence of 385F01 GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTCGGAGACAGA chain sequence light chain GTCACCATCACTTGTCGGGCGAGTCAGGGTGTTAGCAGCTGGTTAGCCTGGTAT CAGCAGAAATCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGATTCAGCGGCAGTGGATCTGGGACAGAGTTCATT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTATCCCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAACGTAC GGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGTC CGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCAA GGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCGT GACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCCT GTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACCA GGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 132 413D08-CDRH1 Amino acid sequence of CDRH1 GFTFRIYG (IMGT) of 413008 using IMGT 133 413D08-CDRH2 Amino acid sequence of CDRH2 IWYDGSNK (IMGT) of 413D08 using IMGT 134 413D08-CDRH3 Amino acid sequence of CDRH3 ARDMDYFGMDV (IMGT) of 413D08 using IMGT 135 413D08-CDRH1 Amino acid sequence of CDRH1 IYGMH (Kabat) of 413D08 using Kabat 136 413D08-CDRH2 Amino acid sequence of CDRH2 VIWYDGSNKYYADSVKG (Kabat) of 413D08 using Kabat 137 413D08-CDRH3 Amino acid sequence of CDRH3 DMDYFGMDV (Kabat) of 413D08 using Kabat 138 413D08-Heavy chain Amino acid sequence of V.sub.H of QVQLVESGGGVVQPGRSLRLSCAASGFTFRIYGMHWVRQAPGKGLEWVAVIWYD variable region 413D08 GSNKYYADSVKGRFTISRDNSDNTLYLQMNSLRAEDTAVYYCARDMDYFGMDVW GQGTTVTVSS 139 413D08-Heavy chain Nucleic acid sequence of V.sub.H of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCCAGCCTGGGAGGTCCCT variable region 413D08 GAGACTCTCCTGTGCAGCGTCTGGATTCACCTTCCGTATTTATGGCATGCACTG GGTCCGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCAGTTATATGGTATG ATGGAAGTAATAAATACTATGCTGACTCCGTGAAGGGCCGATTCACCATCTCCA GAGACAATTCCGACAACACGCTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGGCTGTGTATTACTGTGCGAGAGATATGGACTACTTCGGTATGGACGTCT GGGGCCAAGGGACCACGGTCACCGTCTCCTCAG 140 413D08-full heavy Amino acid sequence of 413D08 QVQLVESGGGVVQPGRSLRLSCAASGFTFRIYGMHWVRQAPGKGLEWVAVIWYD chain sequence heavy chain GSNKYYADSVKGRFTISRDNSDNTLYLQMNSLRAEDTAVYYCARDMDYFGMDVW GQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGAL TSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKINEPKSC DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAP IEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK 141 413D08-full heavy Nucleic acid sequence of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCGTGGTCCAGCCTGGGAGGTCCCT chain sequence 413D08 heavy chain GAGACTCTCCTGTGCAGCGTCTGGATTCACCTTCCGTATTTATGGCATGCACTG GGTCCGCCAGGCTCCAGGCAAGGGGCTGGAGTGGGTGGCAGTTATATGGTATG ATGGAAGTAATAAATACTATGCTGACTCCGTGAAGGGCCGATTCACCATCTCCA GAGACAATTCCGACAACACGCTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGGCTGTGTATTACTGTGCGAGAGATATGGACTACTTCGGTATGGACGTCT GGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCTCTG TGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTCTGG GCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTGGAACTCTG GCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCGGCC TGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCCAGA CCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGAAGG TGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCCCTGA ACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACACCCT GATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCCACGA GGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACAACGC CAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGTCCGT GCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCAAGGT GTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCAAGGG CCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGCTGAC CAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCGATAT CGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCACCCC CCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAGTGGA CAAGTCCCGGTGGCAGCAGGGCMCGTGTTCTCCTGCTCCGTGATGCACGAGGC CCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGCAAG 142 413D08-CDRL1 Amino acid sequence of CDRL1 QGIRND (IMGT) of 413D08 using IMGT 143 413D08-CDRL2 Amino acid sequence of CDRL2 AAS (IMGT) of 413D08 using IMGT 144 413D08-CDRL3 Amino acid sequence of CDRL3 LQHNSYPRT (IMGT) of 413D08 using IMGT 145 413D08-CDRL1 Amino acid sequence of CDRL1 RASQGIRNDLG (Kabat) of 413D08 using Kabat 146 413D08-CDRL2 Amino acid sequence of CDRL2 AASSLQS (Kabat) of 413D08 using Kabat 147 413D08-CDRL3 Amino acid sequence of CDRL3 LQHNSYPRT (Kabat) of 413D08 using Kabat 148 413D08-Light chain Amino acid sequence of V.sub.L of DLQMTQSPSSLSASVGDRVTITCRASQGIRNDLGWYQQKPGKAPKRLIYAASSLQS variable region 413D08 GVPSRFSGSGSGTEFTLTISSLQPEDFATYYCLQHNSYPRTFGQGTINEK 149 413D08-Light chain Nucleic acid sequence of V.sub.L of GACCTCCAGATGACCCAGTCTCCATCCTCCCTGICTGCATCTGTAGGAGACAGA variable region 413D08
GTCACCATCACTTGCCGGGCAAGTCAGGGCATTAGAAATGATTTAGGCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCGCCTGATCTATGCTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGAATTCACT CTCACAATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTATTACTGTCTACAGC ATAATAGTTACCCTCGGACGTTCGGCCAAGGGACCAAGGIGGAAATCAAAC 150 413D08-full light Amino acid sequence of 413D08 DLQMTQSPSSLSASVGDRVTITCRASQGIRNDLGWYQQKPGKAPKRLIYAASSLQS chain sequence light chain GVPSRFSGSGSGTEFTLTISSLQPEDFATYYCLQHNSYPRTFGQGTKVEIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 151 413D08-full light Nucleic acid sequence of GACCTCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGA chain sequence 413D08 light chain GTCACCATCACTTGCCGGGCAAGTCAGGGCATTAGAAATGATTTAGGCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCGCCTGATCTATGCTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGAATTCACT CTCACAATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTATTACTGTCTACAGC ATAATAGTTACCCTCGGACGTTCGGCCAAGGGACCAAGGTGGAAATCAAACGTA CGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGT CCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCA AGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCG TGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCC TGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACC AGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 152 386H03-CDRH1 Amino acid sequence of CDRH1 GGSISSSDW (IMGT) of 386H03 using IMGT 153 386H03-CDRH2 Amino acid sequence of CDRH2 IFHSGNT (IMGT) of 386H03 using IMGT 154 386H03-CDRH3 Amino acid sequence of CDRH3 VRDGSGSY (IMGT) of 386H03 using IMGT 155 386H03-CDRH1 Amino acid sequence of CDRH1 SSDWWS (Kabat) of 386H03 using Kabat 156 386H03-CDRH2 Amino acid sequence of CDRH2 EIFHSGNTNYNPSLKS (Kabat) of 386H03 using Kabat 157 386H03-CDRH3 Amino acid sequence of CDRH3 DGSGSY (Kabat) of 386H03 using Kabat 158 386H03-Heavy chain Amino acid sequence of V.sub.H of QVQLQESGPGLVKPSGTLSLTCAVSGGSISSSDWWSWVRQPPGKGLEWIGEIFHS variable region 386H03 GNTNYNPSLKSRVTISVDKSKNQISLRLNSVTAADTAVYYCVRDGSGSYWGQGTL VTVSS 159 386H03-Heavy chain Nucleic acid sequence of V.sub.H of CAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGGGACCCT variable region 386H03 GTCCCTCACCTGCGCTGTCTCTGGTGGCTCCATCAGCAGTAGTGACTGGTGGAG TTGGGTCCGCCAGCCCCCAGGGAAGGGGCTGGAGTGGATTGGGGAAATCTTTC ATAGTGGGAACACCAACTACAACCCGTCCCTCAAGAGTCGAGTCACCATATCAGT AGACAAGTCCAAGAACCAGATCTCCCTGAGGCTGAACTCTGTGACCGCCGCGGA CACGGCCGTGTATTACTGTGTGAGAGATGGTTCGGGGAGTTACTGGGGCCAGG GAACCCTGGTCACCGTCTCCTCAG 160 386H03-full heavy Amino acid sequence of 386H03 QVQLQESGPGLVKPSGTLSLTCAVSGGSISSSDWWSWVRQPPGKGLEWIGEIFHS chain sequence heavy chain GNTNYNPSLKSRVTISVDKSKNQISLRLNSVTAADTAVYYCVRDGSGSYWGQGTL VWSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVWSWNSGALTSGVH TFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTINDKKVEPKSCDKTHT CPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTIS KAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKT TPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 161 386H03-full heavy Nucleic acid sequence of CAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGGGACCCT chain sequence 386H03 heavy chain GTCCCTCACCTGCGCTGTCTCTGGTGGCTCCATCAGCAGTAGTGACTGGTGGAG TTGGGTCCGCCAGCCCCCAGGGAAGGGGCTGGAGTGGATTGGGGAAATCTTTC ATAGTGGGAACACCAACTACAACCCGTCCCTCAAGAGTCGAGTCACCATATCAGT AGACAAGTCCAAGAACCAGATCTCCCTGAGGCTGAACTCTGTGACCGCCGCGGA CACGGCCGTGTATTACTGTGTGAGAGATGGTTCGGGGAGTTACTGGGGCCAGG GAACCCTGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCTCTGTGTTCCCTC TGGCCCCTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTCTGGGCTGCCTCG TGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTGGAACTCTGGCGCTCTGA CCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCGGCCTGTACTCCC TGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCCAGACCTACATCT GCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGAAGGTGGAACCCA AGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCCCTGAACTGCTGG GCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACACCCTGATGATCT CCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCCACGAGGACCCT GAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACAACGCCAAGACC AAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGTCCGTGCTGACC GTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCAAGGTGTCCAAC AAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCAAGGGCCAGCCC CGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGCTGACCAAGAAC CAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCGATATCGCCGTG GAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCACCCCCCCTGTG CTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAGTGGACAAGTCC CGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATGCACGAGGCCCTGCAC AACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGCAAG 162 386H03-CDRL1 Amino acid sequence of CDRL1 QSVLYSSNNKNY (IMGT) of 386H03 using IMGT 163 386H03-CDRL2 Amino acid sequence of CDRL2 WAS (IMGT) of 3861103 using IMGT 164 386H03-CDRL3 Amino acid sequence of CDRL3 QQYYSTRS (IMGT) of 386H03 using IMGT 165 386H03-CDRL1 Amino acid sequence of CDRL1 KSSQSVLYSSNNKNYLA (Kabat) of 386H03 using Kabat 166 386H03-CDRL2 Amino acid sequence of CDRL2 WASTRES (Kabat) of 386H03 using Kabat 167 386H03-CDRL3 Amino acid sequence of CDRL3 QQYYSTRS (Kabat) of 386H03 using Kabat 168 386H03-Light chain Amino acid sequence of V.sub.L of DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNNKNYLAWYQQKPGQPPKLLIYW variable region 386H03 ASTRESGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSTRSFGQGTKLEIK 169 386H03-Light chain Nucleic acid sequence of V.sub.L of GACATCGTGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAGG variable region 386H03 GCCACCATCAACTGCAAGTCCAGCCAGAGTGTTTTATACAGCTCCAACAATAAGA ACTACTTAGCTTGGTACCAGCAGAAACCAGGACAGCCTCCTAAACTGCTCATTTA CTGGGCATCTACCCGGGAATCCGGGGTCCCTGACCGATTCAGTGGCAGCGGGT CTGGGACAGATTTCACTCTCACCATCAGCAGCCTGCAGGCTGAAGATGTGGCAG TTTATTACTGTCAGCAATATTATAGTACTCGCAGTTTTGGCCAGGGGACCAAGCT GGAGATCAAAC 170 386H03-full light Amino acid sequence of 386H03 DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNNKNYLAWYQQKPGQPPKWYW chain sequence light chain ASTRESGVPDRFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYSTRSFGQGTKLEIK RTVAAPSVFIFPPSDEQLKSGTASWCLLNNFYPREAKVQWKVDNALQSGNSQESV TEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 171 386H03-full light Nucleic acid sequence of GACATCGTGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAGG chain sequence 386H03 light chain GCCACCATCAACTGCAAGTCCAGCCAGAGTGTTTTATACAGCTCCAACAATAAGA ACTACTTAGCTTGGTACCAGCAGAAACCAGGACAGCCTCCTAAACTGCTCATTTA CTGGGCATCTACCCGGGAATCCGGGGTCCCTGACCGATTCAGTGGCAGCGGGT CTGGGACAGATTTCACTCTCACCATCAGCAGCCTGCAGGCTGAAGATGTGGCAG TTTATTACTGTCAGCAATATTATAGTACTCGCAGTTTTGGCCAGGGGACCAAGCT GGAGATCAAACGTACGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGA CGAGCAGCTGAAGTCCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTA CCCCCGCGAGGCCAAGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCA ACTCCCAGGAATCCGTGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGT CCTCCACCCTGACCCTGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCT GCGAAGTGACCCACCAGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGG GCGAGTGT 172 389A03-CDRH1 Amino acid sequence of CDRH1 GGSISSSSYY (IMGT) of 389A03 using IMGT 173 389A03-CDRH2 Amino acid sequence of CDRH2 IYSTGYT (IMGT) of 389A03 using IMGT 174 389A03-CDRH3 Amino acid sequence of CDRH3 AISTAAGPEYFHR (IMGT) of 389A03 using IMGT 175 389A03-CDRH1 Amino acid sequence of CDRH1 SSSYYCG (Kabat) of 389A03 using Kabat 176 389A03-CDRH2 Amino acid sequence of CDRH2 SIYSTGYTYYNPSLKS (Kabat) of 389A03 using Kabat 177 389A03-CDRH3 Amino acid sequence of CDRH3 STAAGPEYFHR (Kabat) of 389A03 using Kabat 178 389A03-Heavy chain Amino acid sequence of V.sub.H of QLQESGPGLVKPSETLSLTCTVSGGSISSSSYYCGWIRQPPGKGLDWIGSTYSTGYT variable region 389A03 YYNPSLKSRVTISIDTSKNQFSCLILTSVTAADTAVYYCAISTAAGPEYFHRWGQGT LVTVSS 179 389A03-Heavy chain Nucleic acid sequence of V.sub.H of CAGCTGCAGGAGTCGGGCCCAGGCCTGGTGAAGCCTTCGGAGACCCTGTCCCTC variable region 389A03 ACCTGCACTGTCTCTGGTGGCTCCATCAGCAGTAGTAGTTATTACTGCGGCTGG ATCCGCCAGCCCCCTGGGAAGGGGCTGGACTGGATTGGGAGTATCTATTCTACT GGGTACACCTACTACAACCCGTCCCTCAAGAGTCGAGTCACCATTICCATAGACA CGTCCAAGAACCAGTTCTCATGCCTGATACTGACCTCTGTGACCGCCGCAGACA CGGCTGTGTATTACTGTGCGATAAGTACAGCAGCTGGCCCTGAATACTTCCATC GCTGGGGCCAGGGCACCCTGGTCACCGTCTCCTCAG 180 389A03-full heavy Amino acid sequence of 389A03 QLQESGPGLVKPSETLSLTCTVSGGSISSSSYYCGWIRQPPGKGLDWIGSTYSTGYT chain sequence heavy chain YYNPSLKSRVTISIDTSKNQFSCLILTSVTAADTAVYYCAISTAAGPEYFHRWGQGT LVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVH TFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTINDKINEPKSCDKTHT CPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTIS KAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKT TPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 181 389A03-full heavy Nucleic acid sequence of 389A03 CAGCTGCAGGAGTCGGGCCCAGGCCTGGTGAAGCCTTCGGAGACCCTGTCCCTC chain sequence heavy chain ACCTGCACTGTCTCTGGTGGCTCCATCAGCAGTAGTAGTTATTACTGCGGCTGG ATCCGCCAGCCCCCTGGGAAGGGGCTGGACTGGATTGGGAGTATCTATTCTACT GGGTACACCTACTACAACCCGTCCCTCAAGAGTCGAGTCACCATTTCCATAGACA CGTCCAAGAACCAGTTCTCATGCCTGATACTGACCTCTGTGACCGCCGCAGACA CGGCTGTGTATTACTGTGCGATAAGTACAGCAGCTGGCCCTGAATACTTCCATC GCTGGGGCCAGGGCACCCTGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCT CTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTC TGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTGGAACT CTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCG GCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCC AGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGA AGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCC CTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACA CCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCC ACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACA ACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGT CCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCA AGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCA AGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGC TGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCG ATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCA CCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAG TGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATGCACG AGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGCAAG 182 389A03-CDRL1 Amino acid sequence of CDRL1 QSVLYSSNSKNF (IMGT) of 389A03 using IMGT 183 389A03-CDRL2 Amino acid sequence of CDRL2 WAS (IMGT) of 389A03 using IMGT 184 389A03-CDRL3 Amino acid sequence of CDRL3 QQYYSTPRT (IMGT) of 389A03 using IMGT 185 389A03-CDRL1 Amino acid sequence of CDRL1 KSSQSVLYSSNSKNFLA (Kabat) of 389A03 using Kabat 186 389A03-CDRL2 Amino acid sequence of CDRL2 WASTRGS (Kabat) of 389A03 using Kabat 187 389A03-CDRL3 Amino acid sequence of CDRL3 QQYYSTPRT (Kabat) of 389A03 using Kabat 188 389A03-Light chain Amino acid sequence of V.sub.L of DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNSKNFLAWYQQKPGQPPKLFIYW variable region 389A03
ASTRGSGVPDRISGSGSGTDFNLTISSLQAEDVAVYYCQQYYSTPRTFGQGTKVEI K 189 389A03-Light chain Nucleic acid sequence of V.sub.L of GACATCGTGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAGG variable region 389A03 GCCACCATCAACTGCAAGTCCAGCCAGAGTGTTTTATACAGCTCCAACAGTAAGA ACTTCTTAGCTTGGTACCAGCAGAAACCGGGACAGCCTCCTAAGCTGTTCATTTA CTGGGCATCTACCCGGGGATCCGGGGTCCCTGACCGAATCAGTGGCAGCGGGT CTGGGACAGATTTCAATCTCACCATCAGCAGCCTGCAGGCTGAAGATGTGGCAG TTTATTACTGTCAACAATATTATAGTACTCCTCGGACGTTCGGCCAAGGGACCAA GGTGGAGATCAAAC 190 389A03-full light Amino acid sequence of 389A03 DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNSKNFLAWYQQKPGQPPKLFIYW chain sequence light chain ASTRGSGVPDRISGSGSGTDFNLTISSLQAEDVAVYYCQQYYSTPRTFGQGTKVEI KRTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQES VTEQDSKIDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 191 389A03-full light Nucleic acid sequence of 389A03 GACATCGTGATGACCCAGTCTCCAGACTCCCTGGCTGTGTCTCTGGGCGAGAGG chain sequence light chain GCCACCATCAACTGCAAGTCCAGCCAGAGTGTTTTATACAGCTCCAACAGTAAGA ACTTCTTAGCTTGGTACCAGCAGAAACCGGGACAGCCTCCTAAGCTGTTCATTTA CTGGGCATCTACCCGGGGATCCGGGGTCCCTGACCGAATCAGTGGCAGCGGGT CTGGGACAGATTTCAATCTCACCATCAGCAGCCTGCAGGCTGAAGATGTGGCAG TTTATTACTGTCAACAATATTATAGTACTCCTCGGACGTTCGGCCAAGGGACCAA GGTGGAGATCAAACGTACGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTC CGACGAGCAGCTGAAGTCCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTT CTACCCCCGCGAGGCCAAGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCG GCAACTCCCAGGAATCCGTGACCGAGCAGGACTCCAAGGACAGCACCTACTCCC TGTCCTCCACCCTGACCCTGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACG CCTGCGAAGTGACCCACCAGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACC GGGGCGAGTGT 192 Human IGHG4* Heavy Chain Constant Region gcttccaccaagggcccatccgtcttccccctggcgccctgctccaggagcacctccgagagcacagccg IgG4 heavy 01 & Nucleotide Sequence ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgccctgac chain IGHG4* cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg constant 04 tgccctccagcagcttgggcacgaagacctacacctgcaacgtagatcacaagcccagcaacaccaagg region #1 tggacaagagagttgagtccaaatatggtcccccatgcccatcatgcccagcacctgagttcctgggggg accatcagtcttcctgttccccccaaaacccaaggacactctcatgatctcccggacccctgaggtcacg- tg cgtggtggtggacgtgagccaggaagaccccgaggtccagttcaactggiacgtggatggcgtggagg tgcataatgccaagacaaagccgcgggaggagcagttcaacagcacgtaccgtgtggtcagcgtcctca ccgtcctgcaccaggactggctgaacggcaaggagtacaagtgcaaggtctccaacaaaggcctcccgt cctccatcgagaaaaccatctccaaagccaaagggcagccccgagagccacaggtgtacaccctgcccc catcccaggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctaccccagcg acatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtgctg gactccgacggctccttcttcctctacagcaggctaaccgtggacaagagcaggtggcaggaggggaat gtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacacagaagagcctctccctgtctc- tg ggtaaa 193 Heavy Chain Constant Region ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Amino Acid Sequence QSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNKVNDKRVESKYGPPCPSCPAPEF LGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTK PREEQFNSTYRVVSVLTVLHQDWLNGKEYKOWSNKGLPSSIEKTISKAKGQPREP QVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTIPPVLDSD GSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK 194 Human IGHG4* Heavy Chain Constant Region gcttccaccaagggcccatccgtcttccccctggcgccctgctccaggagcacctccgagagcacagccg IgG4 heavy 02 Nucleotide Sequence ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgccctgac chain cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtgg- tgaccg constant tgccctccagcagcttgggcacgaagacctacacctgcaacgtagatcacaagcccagcaacaccaagg region #2 tggacaagagagttgagtccaaatatggtcccccgtgcccatcatgcccagcacctgagttcctgggggg accatcagtcttcctgttccccccaaaacccaaggacactctcatgatctcccggacccctgaggtcacg- tg cgtggtggtggacgtgagccaggaagaccccgaggtccagttcaactggtacgtggatggcgtggagg tgcataatgccaagacaaagccgcgggaggagcagttcaacagcacgtaccgtgtggtcagcgtcctca ccgtcgtgcaccaggactggctgaacggcaaggagtacaagtgcaaggtctccaacaaaggcctcccg tcctccatcgagaaaaccatctccaaagccaaagggcagccccgagagccacaggtgtacaccctgccc ccatcccaggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctaccccagc gacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtgct ggactccgacggctccttcttcctctacagcaggctaaccgtggacaagagcaggtggcaggagggga atgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcctctccctgtc- tct gggtaaa 195 Heavy Chain Constant Region ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Amino Acid Sequence QSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPSCPAPEF LGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTK PREEQFNSTYRVVSVLTVVHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPREP QVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSD GSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK 196 Human IGHG4* Heavy Chain Constant Region gcttccaccaagggcccatccgtcttccccctggcgccctgctccaggagcacctccgagagcacagccg IgG4 heavy 03 Nucleotide Sequence ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgccctgac chain cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtgg- tgaccg constant tgccctccagcagcttgggcacgaagacctacacctgcaacgtagatcacaagcccagcaacaccaagg region #3 tggacaagagagttgagtccaaatatggtcccccatgcccatcatgcccagcacctgagttcctgggggg accatcagtcttcctgttccccccaaaacccaaggacactctcatgatctcccggacccctgaggtcacg- tg cgtggtggtggacgtgagccaggaagaccccgaggtccagttcaactggtacgtggatggcgtggagg tgcataatgccaagacaaagccgcgggaggagcagttcaacagcacgtaccgtgtggtcagcgtcctca ccgtcctgcaccaggactggctgaacggcaaggagtacaagtgcaaggtctccaacaaaggcctcccgt cctccatcgagaaaaccatctccaaagccaaagggcagccccgagagccacaggtgtacaccctgcccc catcccaggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctaccccagcg acatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgtgctg gactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtggcaggaggggaac gtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcctctccctgtctc- tg ggtaaa 197 Heavy Chain Constant Region ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Amino Acid Sequence QSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTINDKRVESKYGPPCPSCPAPEF LGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTK PREEQFNSTYRVVSVLTVLHQDWLNGKEYKCVSNKGLPSSIEKTISKAKGQPREP QVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSD GSFFLYSKLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK 198 IgG4 heavy IGHG4- Heavy Chain Constant Region gcctccaccaagggcccatccgtcttccccctggcgccctgctccaggagcacctccgagagcacggccg chain PE Nucleotide Sequence-Synthetic ccctgggctgcctggtcaaggactacttccccgaaccagtgacggtgtcgtggaactcaggcgccctgac constant Version A cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg region- tgccctccagcagcttgggcacgaagacctacacctgcaacgtagatcacaagcccagcaaca- ccaagg IgG4-PE tggacaagagagttgagtccaaatatggtcccccatgcccaccatgcccagcgcctgaatttg- aggggg gaccatcagtcttcctgttccccccaaaacccaaggacactctcatgatctcccggacccctgaggtcac- gt gcgtggtggtggacgtgagccaggaagaccccgaggtccagttcaactggtacgtggatggcgtggag gtgcataatgccaagacaaagccgcgggaggagcagttcaacagcacgtaccgtgtggtcagcgtcctc accgtcctgcaccaggactggctgaacggcaaggagtacaagtgcaaggtctccaacaaaggcctccc gtcatcgatcgagaaaaccatctccaaagccaaagggcagccctgagagccacaggtgtacaccctgc ccccatcccaggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctacccca gcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgt gctggactccgacggatccttcttcctctacagcaggctaaccgtggacaagagcaggtggcaggaggg gaatgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacacagaagagcctctccctg- tc tctgggtaaa 199 IgG4 heavy Heavy Chain Constant Region ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPV7VSWNSGALTSGVHTFPAVL chain Amino Acid Sequence- QSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNT1NDKRVESKYGPPCPPCPAPEF constant Encoded by Synthetic Version A, EGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKT region- B & C (Two residues that differ KPREEQFNSTYRVVSVLTVLHQDWLNGKEYKOWSNKGLPSSIEKTISKAKGQPRE IgG4-PE from the wild-type sequence are PQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSD identified in bold) GSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK 200 IgG4 heavy Heavy Chain Constant Region Gcctccaccaagggacctagcgtgttccctctcgccccctgttccaggtccacaagcgagtccaccgctgc chain Nucleotide Sequence-Synthetic cctcggctgtctggtgaaagactactttcccgagcccgtgaccgtctcctggaatagcggagccctgacct constant Version B ccggcgtgcacacatttcccgccgtgctgcagagcagcggactgtatagcctgagcagcgtggtgaccgt region- gcccagctccagcctcggcaccaaaacctacacctgcaacgtggaccacaagccctccaacac- caaggt IgG4-PE ggacaagcgggtggagagcaagtacggccccccttgccctccttgtcctgcccctgagttcga- gggagg accctccgtgttcctgtttccccccaaacccaaggacaccctgatgatctcccggacacccgaggtgacc- t gtgtggtcgtggacgtcagccaggaggaccccgaggtgcagttcaactggtatgtggacggcgtggag gtgcacaatgccaaaaccaagcccagggaggagcagttcaattccacctacagggtggtgagcgtgct gaccgtcctgcatcaggattggctgaacggcaaggagtacaagtgcaaggtgtccaacaagggactgc ccagctccatcgagaagaccatcagcaaggctaagggccagccgagggagccccaggtgtataccctg cctcctagccaggaagagatgaccaagaaccaagtgtccctgacctgcctggtgaagggattctacccct ccgacatcgccgtggagtgggagagcaatggccagcccgagaacaactacaaaacaacccctcccgtg ctcgatagcgacggcagcttctttctctacagccggctgacagtggacaagagcaggtggcaggagggc aacgtgttctcctgttccgtgatgcacgaggccctgcacaatcactacacccagaagagcctctccctgt- cc ctgggcaag 201 IgG4 heavy Heavy Chain Constant Region gccagcaccaagggcccttccgtgttccccctggccccttgcagcaggagcacctccgaatccacagctg chain Nucleotide Sequence-Synthetic ccctgggctgtctggtgaaggactactttcccgagcccgtgaccgtgagctggaacagcggcgctctgac constant Version C atccggcgtccacacctttcctgccgtcctgcagtcctccggcctctactccctgtcctccgtggtgaccgtg region- cctagctcctccctcggcaccaagacctacacctgtaacgtggaccacaaaccctccaacacc- aaggtgg IgG4-PE acaaacgggtcgagagcaagtacggccctccctgccctccttgtcctgcccccgagttcgaag- gcggacc cagcgtgttcctgttccctcctaagcccaaggacaccctcatgatcagccggacacccgaggtgacctgc gtggtggtggatgtgagccaggaggaccctgaggtccagttcaactggtatgtggatggcgtggaggtg cacaacgccaagacaaagccccgggaagagcagttcaactccacctacagggtggtcagcgtgctgac cgtgctgcatcaggactggctgaacggcaaggagtacaagtgcaaggtcagcaataagggactgccca gcagcatcgagaagaccatctccaaggctaaaggccagccccgggaacctcaggtgtacaccctgcctc ccagccaggaggagatgaccaagaaccaggtgagcctgacctgcctggtgaagggattctacccttccg acatcgccgtggagtgggagtccaacggccagcccgagaacaattataagaccacccctcccgtcctcg acagcgacggatccttcdtctgtactccaggctgaccgtggataagtccaggtggcaggaaggcaacgt gttcagctgctccgtgatgcacgaggccctgcacaatcactacacccagaagtccctgagcctgtccctg- g gaaag 202 IgG4 heavy Heavy Chain Constant Region gcctccaccaagggcccatccgtcttccccctggcgccctgctccaggagcacctccgagagcacggccg chain Nucleotide Sequence ccctgggctgcctggtcaaggactacttccccgaaccagtgacggtgtcgtggaactcaggcgccctgac constant cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg region- tgccctccagcagcttgggcacgaagacctacacctgcaacgtagatcacaagcccagcaaca- ccaagg in- tggacaagagagttgagtccaaatatggtcccccatgcccaccatgcccagcgcctccagttgcggg- gg activated gaccatcagtcttcctgttccccccaaaacccaaggacactctcatgatctcccggacccctgaggtcacgt gcgtggtggtggacgtgagccaggaagaccccgaggtccagttcaactggtacgtggatggcgtggag gtgcataatgccaagacaaagccgcgggaggagcagttcaacagcacgtaccgtgtggtcagcgtcctc accgtcctgcaccaggactggctgaacggcaaggagtacaagtgcaaggtctccaacaaaggcctccc gtcatcgatcgagaaaaccatctccaaagccaaagggcagccccgagagccacaggtgtacaccctgc ccccatcccaggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctacccca gcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacgcctcccgt gctggactccgacggatccttcttcctctacagcaggctaaccgtggacaagagcaggtggcaggaggg gaatgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacacagaagagcctctccctg- tc tctgggtaaa 203 Heavy Chain Constant Region ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Amino Acid Sequence QSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPAPP (inactivating mutations from VAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAK human WT IgG4 shown in bold) TKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSSIEKTISKAKGQPR EPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTYPPVLDS DGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLGK 204 Disabled Disabled Heavy Chain Constant Region gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctgggggcacagcgg Human IGHG1 Nucleotide Sequence ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgccctgac IgG1 heavy cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg
chain tgccctccagcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacacc- aagg constant tggacaagaaagtggagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaac region tcgcgggggcaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccg- gacccct gaggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactggtacgtggac ggcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaacagcacgtaccgtgtgg tcagcgtcctcaccgtcctgcaccaggactggctgaatggcaaggagtacaagtgcaaggtctccaaca aagccctcccagcccccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtg tacaccctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggct tctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccac gcctcccgtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtgg- c agcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcct ctccctgtctccgggtaaa 205 Heavy Chain Constant Region ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Amino Acid Sequence (Two QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCP residues that differ from the APELAGAPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN wild-type sequence are AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISMKGQ identified in bold) PREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVL DSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 206 Human C.kappa. IGKC*01 C.kappa. Light Chain Constant Region cgtacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccggcaccgcttct constant Nucleotide Sequence gtcgtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtggacaacgccctg region cagtccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcacctactccctgt- cctcca ccctgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacccaccagggc ctgtctagccccgtgaccaagtctttcaaccggggcgagtgt 207 C.kappa. Light Chain Constant Region RTVAAPSVFIFPPSDEQLKSGTASWCLLNNFYPREAKVQWKVDNALQSGNSQESV Amino Acid Sequence TEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 208 Human C.kappa. IGKC*02 C.kappa. Light Chain Constant Region cgaactgtggctgcaccatctgtcttcatcttcccgccatctgatgagcagttgaaatctggaactgcctctg constant Nucleotide Sequence ttgtgtgcctgctgaataacttctatcccagagaggccaaagtacagtggaaggtggataacgccctcca region atcgggtaactcccaggagagtgtcacagagcaggagagcaaggacagcacctacagcctcagc- agc accctgacgctgagcaaagcagactacgagaaacacaaagtctacgccggcgaagtcacccatcaggg cctgagctcgcccgtcacaaagagcttcaacaggggagagtgt 209 C.kappa. Light Chain Constant Region RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESV Amino Acid Sequence TEQESKDSTYSLSSTLTLSKADYEKHKVYAGEVTHQGLSSPVTKSFNRGEC 210 Human C.kappa. IGKC*03 C.kappa. Light Chain Constant Region cgaactgtggctgcaccatctgtcttcatcttcccgccatctgatgagcagttgaaatctggaactgcctctg constant Nucleotide Sequence ttgtgtgcctgctgaataacttctatcccagagaggccaaagtacagcggaaggtggataacgccctcca region atcgggtaactcccaggagagtgtcacagagcaggagagcaaggacagcacctacagcctcagc- agc accctgacgctgagcaaagcagactacgagaaacacaaagtctacgcctgcgaagtcacccatcaggg cctgagctcgcccgtcacaaagagcttcaacaggggagagtgt 211 C.kappa. Light Chain Constant Region RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQRKVDNALQSGNSQESV Amino Acid Sequence TEQESKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 212 Human C.kappa. IGKC*04 C.kappa. Light Chain Constant Region cgaactgtggctgcaccatctgtcttcatcttcccgccatctgatgagcagttgaaatctggaactgcctctg constant Nucleotide Sequence ttgtgtgcctgctgaataacttctatcccagagaggccaaagtacagtggaaggtggataacgccctcca region atcgggtaactcccaggagagtgtcacagagcaggacagcaaggacagcacctacagcctcagc- agc accctgacgctgagcaaagcagactacgagaaacacaaactctacgcctgcgaagtcacccatcaggg cctgagctcgcccgtcacaaagagcttcaacaggggagagtgt 213 C.kappa. Light Chain Constant Region RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESV Amino Acid Sequence TEQDSKDSTYSLSSTLTLSKADYEKHKLYACEVTHQGLSSPVTKSFNRGEC 214 Human C.kappa. IGKC*05 C.kappa. Light Chain Constant Region cgaactgtggctgcaccatctgtcttcatcttcccgccatctgatgagcagttgaaatctggaactgcctctg constant Nucleotide Sequence ttgtgtgcctgctgaataacttctatcccagagaggccaaagtacagtggaaggtggataacgccctcca region atcgggtaactcccaggagagtgtcacagagcaggacagcaaggacagcacctacagcctcagc- aac accctgacgctgagcaaagcagactacgagaaacacaaagtctacgcctgcgaagtcacccatcaggg cctgagctcgcccgtcacaaagagcttcaacaggggagagtgc 215 C.kappa. Light Chain Constant Region RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWINDNALQSGNSQESV Amino Acid Sequence TEQDSKDSTYSLSNTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 216 Human C.lamda. IGLC1* C.lamda. Light Chain Constant Region cccaaggccaaccccacggtcactctgttcccgccctcctctgaggagctccaagccaacaaggccacac constant 01 Nucleotide Sequence tagtgtgtctgatcagtgacttctacccgggagctgtgacagtggcttggaaggcagatggcagccccgt region caaggcgggagtggagacgaccaaaccctccaaacagagcaacaacaagtacgcggccagcagc- ta cctgagcctgacgcccgagcagtggaagtcccacagaagctacagctgccaggtcacgcatgaaggga gcaccgtggagaagacagtggcccctacagaatgttca 217 C.lamda. Light Chain Constant Region PKANPTVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADGSPVKAGVETTKPS Amino Acid Sequence KQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS 218 Human C.lamda. IGLC1* C.lamda. Light Chain Constant Region ggtcagcccaaggccaaccccactgtcactctgttcccgccctcctctgaggagctccaagccaacaagg constant 02 Nucleotide Sequence-version A ccacactagtgtgtctgatcagtgacttctacccgggagctgtgacagtggcctggaaggcagatggcag region- ccccgtcaaggcgggagtggagaccaccaaaccctccaaacagagcaacaacaagtacgcggc- cagc agctacctgagcctgacgcccgagcagtggaagtcccacagaagctacagctgccaggtcacgcatga agggagcaccgtggagaagacagtggcccctacagaatgttca 219 C.lamda. Light Chain Constant Region GQPKANPTVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADGSPVKAGVETT Amino Acid Sequence-encoded KPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS by version A or B 220 C.lamda. Light Chain Constant Region ggtcagcccaaggccaaccccactgtcactctgttcccgccctcctctgaggagctccaagccaacaagg Nucleotide Sequence-Version B ccacactagtgtgtctgatcagtgacttctacccgggagctgtgacagtggcctggaaggcagatggcag ccccgtcaaggcgggagtggagaccaccaaaccctccaaacagagcaacaacaagtacgcggccagc agctacctgagcctgacgcccgagcagtggaagtcccacagaagctacagctgccaggtcacgcatga agggagcaccgtggagaagacagtggcccctacagaatgttca 221 Human C.lamda. IGLC2* C.lamda. Light Chain Constant Region ggccagcctaaggccgctccttctgtgaccctgttccccccatcctccgaggaactgcaggctaacaaggc constant 01 Nucleotide Sequence-Version A caccctcgtgtgcctgatcagcgacttctaccctggcgccgtgaccgtggcctggaaggctgatagctctc region ctgtgaaggccggcgtggaaaccaccaccccttccaagcagtccaacaacaaatacgccgcctc- ctccta cctgtccctgacccctgagcagtggaagtcccaccggtcctacagctgccaagtgacccacgagggctcc accgtggaaaagaccgtggctcctaccgagtgctcc 222 C.lamda. Light Chain Constant Region ggccagcctaaagctgcccccagcgtcaccctgtttcctccctccagcgaggagctccaggccaacaagg Nucleotide Sequence-Version B ccaccctcgtgtgcctgatctccgacttctatcccggcgctgtgaccgtggcttggaaagccgactccagcc ctgtcaaagccggcgtggagaccaccacaccctccaagcagtccaacaacaagtacgccgcctccagct atctctccctgacccctgagcagtggaagtcccaccggtcctactcctgtcaggtgacccacgagggctc- c accgtggaaaagaccgtcgcccccaccgagtgctcc 223 C.lamda. Light Chain Constant Region GQPKANPTVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADGSPVKAGVETT Amino Acid Sequence-Encoded KPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS by Version A or B 224 Human C.lamda. IGLC2* C.lamda. Light Chain Constant Region ggtcagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagccaacaaggc constant 02 & Nucleotide Sequence cacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggcctggaaggcagatagcag region IGLC2* ccccgtcaaggcgggagtggagaccaccacaccctccaaacaaagcaacaacaagtacgcggccagc 03 agctatctgagcctgacgcctgagcagtggaagtcccacagaagctacagctgccaggtcacgcatga- a gggagcaccgtggagaagacagtggcccctacagaatgttca 225 C.lamda. Light Chain Constant Region GQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKAGVETTT Amino Acid Sequence PSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS 226 Human C.lamda. IGLC3* C.lamda. Light Chain Constant Region cccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggccacact constant 01 Nucleotide Sequence ggtgtgtctcataagtgacttctacccgggagccgtgacagttgcctggaaggcagatagcagccccgtc region aaggcgggggtggagaccaccacaccctccaaacaaagcaacaacaagtacgcggccagcagct- acc tgagcctgacgcctgagcagtggaagtcccacaaaagctacagctgccaggtcacgcatgaagggagc accgtggagaagacagttgcccctacggaatgttca 227 C.lamda. Light Chain Constant Region PKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKAGVETTTPS Amino Acid Sequence KQSNNKYAASSYLSLTPEQWKSHKSYSCQVTHEGSTVEKTVAPTECS 228 Human C.lamda. IGLC3* C.lamda. Light Chain Constant Region ggtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggc constant 02 Nucleotide Sequence cacactggtgtgtctcataagtgacttctacccggggccagtgacagttgcctggaaggcagatagcagc region cccgtcaaggcgggggtggagaccaccacaccctccaaacaaagcaacaacaagtacgcggcca- gca gctacctgagcctgacgcctgagcagtggaagtcccacaaaagctacagctgccaggtcacgcatgaag ggagcaccgtggagaagacagtggcccctacqgaatgttca 229 C.lamda. Light Chain Constant Region GQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGPVTVAWKADSSPVKAGVETTT Amino Acid Sequence PSKQSNNKYAASSYLSLTPEQWKSHKSYSCQVTHEGSTVEKTVAPTECS 230 Human C.lamda. IGLC3* C.lamda. Light Chain Constant Region ggtcagcccaaggctgccccctcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggc constant 03 Nucleotide Sequence cacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggcctggaaggcagatagcag region ccccgtcaaggcgggagtggagaccaccacaccctccaaacaaagcaacaacaagtacgcggcc- agc agctacctgagcctgacgcctgagcagtggaagtcccacaaaagctacagctgccaggtcacgcatgaa gggagcaccgtggagaagacagtggcccctacagaatgttca 231 C.lamda. Light Chain Constant Region GQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKAGVETTT Amino Acid Sequence PSKQSNNKYAASSYLSLTPEQWKSHKSYSCQVTHEGSTVEKTVAPTECS 232 Human C.lamda. IGLC3* C.lamda. Light Chain Constant Region ggtcagcccaaggctgccccctcggtcactctgttcccgccctcctctgaggagcttcaagccaacaaggc constant 04 Nucleotide Sequence cacactggtgtgtctcataagtgacttctacccgggagccgtgacagtggcctggaaggcagatagcag region ccccgtcaaggcgggagtggagaccaccacaccctccaaacaaagcaacaacaagtacgcggcc- agc agctacctgagcctgacgcctgagcagtggaagtcccacagaagctacagctgccaggtcacgcatgaa gggagcaccgtggagaagacagtggcccctacagaatgttca 233 C.lamda. Light Chain Constant Region GQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKAGVETTT Amino Acid Sequence PSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS 234 Human C.lamda. IGLC6* C.lamda. Light Chain Constant Region ggtcagcccaaggctgccccatcggtcactctgttcccgccctcctctgaggagcttcaagccaacaaggc constant 01 Nucleotide Sequence cacactggtgtgcctgatcagtgacttctacccgggagcbgtgaaagtggcctggaaggcagatggcag region ccccgtcaacacgggagtggagaccaccacaccctccaaacagagcaacaacaagtacgcggcc- agc agctacctgagcctgacgcctgagcagtggaagtcccacagaagctacagctgccaggtcacgcatgaa gggagcaccgtggagaagacagtggcccctgcagaatgttca 235 C.lamda. Light Chain Constant Region GQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVKVAWKADGSPVNTGVETT Amino Acid Sequence TPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPAECS 236 Human C.lamda. IGLC7* C.lamda. Light Chain Constant Region ggtcagcccaaggctgccccatcggtcactctgttcccaccctcctctgaggagcttcaagccaacaaggc constant 01 & Nucleotide Sequence cacactggtgtgtctcgtaagtgacttctacccgggagccgtgacagtggcctggaaggcagatggcag region IGLC7* ccccgtcaaggtgggagtggagaccaccaaaccctccaaacaaagcaacaacaagtatgcggccagc 02 agctacctgagcctgacgcccgagcagtggaagtcccacagaagctacagctgccgggtcacgcatga agggagcaccgtggagaagacagtggcccctgcagaatgctct 237 C.lamda. Light Chain Constant Region GQPKAAPSVTLFPPSSEELQANKATLVCLVSDFYPGAVTVAWKADGSPVKVGVETT Amino Acid Sequence KPSQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAPAECS 238 413G05-CDRH1 Amino acid sequence of CDRH1 GFTFSDYY (IMGT) of 413G05 using IMGT 239 413G05-CDRH2 Amino acid sequence of CDRH2 ISTSGSTI (IMGT) of 413G05 using IMGT 240 413G05-CDRH3 Amino acid sequence of CDRH3 ARGITGTNFYHYGLGV (IMGT) of 413G05 using IMGT 241 413G05-CDRH1 Amino acid sequence of CDRH1 DYYMS
(Kabat) of 413G05 using Kabat 242 413G05-CDRH2 Amino acid sequence of CDRH2 YISTSGSTIYYADSVKG (Kabat) of 413G05 using Kabat 243 413G05-CDRH3 Amino acid sequence of CDRH3 GITGTNFYHYGLGV (Kabat) of 413G05 using Kabat 244 413G05-Heavy chain Amino acid sequence of V.sub.H of QVQLVESGGGLVKPGGSLRLSCAASGFTFSDYYMSWIRQVPGKGLEWVSYISTSG variable region 413G05 STIYYADSVKGRFTISRDNAKNSLYLQMNSLRAEDAAVYHCARGITGTNFYHYGLG VWGQGTFVTVSS 245 413G05-Heavy chain Nucleic acid sequence of V.sub.H of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGGTCCCT variable region 413G05 GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTCAGTGACTACTACATGAGCTG GATCCGCCAGGTTCCAGGGAAGGGGCTGGAGTGGGTTTCATACATTAGTACTAG TGGTAGTACCATATACTACGCAGACTCTGTGAAGGGCCGATTCACCATCTCCAG GGACAACGCCAAGAACTCACTGTATCTACAAATGAACAGCCTGAGAGCCGAGGA CGCGGCCGTGTATCACTGTGCGAGAGGTATAACTGGAACTAACTTCTACCACTA CGGTTTGGGCGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAG 246 413G05-full heavy Amino acid sequence of 413G05 QVQLVESGGGLVKPGGSLRLSCAASGFIFSDYYMSWIRQVPGKGLEWVSYISTSG chain sequence heavy chain STIYYADSVKGRFTISRDNAKNSLYLQMNSLRAEDAAVYHCARGITGTNFYHYGLG VWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNS GALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKINEP KSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVK FNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQ PENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSL SLSPGK 247 413G05-full heavy Nucleic acid sequence of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGGTCCCT chain sequence 413G05 heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTCAGTGACTACTACATGAGCTG GATCCGCCAGGTTCCAGGGAAGGGGCTGGAGTGGGTTTCATACATTAGTACTAG TGGTAGTACCATATACTACGCAGACTCTGTGAAGGGCCGATTCACCATCTCCAG GGACAACGCCAAGAACTCACTGTATCTACAAATGAACAGCCTGAGAGCCGAGGA CGCGGCCGTGTATCACTGTGCGAGAGGTATAACTGGAACTAACTTCTACCACTA CGGTTTGGGCGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCCAGCAC CAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGG AACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGT GTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCT GCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTC TCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAG GTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCT TGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAG CCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTG GATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTG GAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTAC CGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGA GTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCAT CTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAG CAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTT CTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAA CTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGC AAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCC GTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGC CCCGGCAAG 248 413G05-CDRL1 Amino acid sequence of CDRL1 QGINSW (IMGT) of 413G05 using IMGT 249 413G05-CDRL2 Amino acid sequence of CDRL2 MS (IMGT) of 413G05 using IMGT 250 413G05-CDRL3 Amino acid sequence of CDRL3 QQVNSFPLT (IMGT) of 413G05 using IMGT 251 413G05-CDRL1 Amino acid sequence of CDRL1 RASQGINSWLA (Kabat) of 413G05 using Kabat 252 413G05-CDRL2 Amino acid sequence of CDRL2 AASTLQS (Kabat) of 413G05 using Kabat 253 413G05-CDRL3 Amino acid sequence of CDRL3 QQVNSFPLT (Kabat) of 413G05 using Kabat 254 413G05-Light chain Amino acid sequence of V.sub.L of DIQMTQSPSSVSASVGDRVTITCRASQGINSWLAWYQQKPGKAPKLLIYAASTLQS variable region 413G05 GVPSRFSGSGSGADFTLTISSLQPEDFATYYCQQVNSFPLTFGGGMEIK 255 413G05-Light chain Nucleic acid sequence of V.sub.L of GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTAGGAGACAGA variable region 413G05 GTCACCATCACTTGTCGGGCGAGTCAGGGTATFAACAGCTGGTTAGCCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCACTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGGTCTGGGGCAGATTTCACT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG TTAACAGTTTCCCGCTCACTTTCGGCGGAGGGACCAAGGTGGAGATCAAAC 256 413G05-full light Amino acid sequence of 413G05 DIQMTQSPSSVSASVGDRVTITCRASQGINSWLAWYQQKPGKAPKLLIYAASTLQS chain sequence light chain GVPSRFSGSGSGADFTLTISSLQPEDFATYYCQQVNSFPLTFGGGTKVEIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 257 413G05-full light Nucleic acid sequence of GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTAGGAGACAGA chain sequence 413G05 light chain GTCACCATCACTTGTCGGGCGAGTCAGGGTATTAACAGCTGGTTAGCCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCACTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGGTCTGGGGCAGATTTCACT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG TTAACAGTTTCCCGCTCACTTTCGGCGGAGGGACCAAGGTGGAGATCAAACGTA CGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGT CCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCA AGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCG TGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCC TGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACC AGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 258 413F09-CDRH1 Amino acid sequence of CDRH1 GFTFSYYA (IMGT) of 413F09 using IMGT 259 413F09-CDRH2 Amino acid sequence of CDRH2 ISGGGGNT (IMGT) of 413F09 using IMGT 260 413F09-CDRH3 Amino acid sequence of CDRH3 AKDRMKQLVRAYYFDY (IMGT) of 413F09 using IMGT 261 413F09-CDRH1 Amino acid sequence of CDRH1 YYAMS (Kabat) of 413F09 using Kabat 262 413F09-CDRH2 Amino acid sequence of CDRH2 TISGGGGNTHYADSVKG (Kabat) of 413F09 using Kabat 263 413F09-CDRH3 Amino acid sequence of CDRH3 DRMKQLVRAYYFDY (Kabat) of 413F09 using Kabat 264 413F09-Heavy chain Amino acid sequence of V.sub.H of EVPLVESGGGLVQPGGSLRLSCAASGFTFSYYAMSWVRQAPGKGLDWVSTISGGG variable region 413F09 GNTHYADSVKGRFTISRDNSKNTLYLHMNSLRAEDTAVYYCAKDRMKQLVRAYYF DYWGQGTLVTVSS 265 413F09-Heavy chain Nucleic acid sequence of V.sub.H of GAGGTGCCGCTGGTGGAGTCTGGGGGAGGCTTGGTACAGCCTGGGGGGTCCCT variable region 413F09 GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGCTACTATGCCATGAGCTG GGTCCGTCAGGCTCCAGGGAAGGGGCTGGACTGGGTCTCAACTATTAGTGGTG GIGGTGGTAACACACACTACGCAGACTCCGTGAAGGGCCGATTCACTATATCCA GAGACAATTCCAAGAACACGCTGTATCTGCACATGAACAGCCTGAGAGCCGAAG ACACGGCCGTCTATTACTGTGCGAAGGATCGGATGAAACAGCTCGTCCGGGCCT ACTACTTTGACTACTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAG 266 413F09-full heavy Amino acid sequence of 413F09 EVPLVESGGGLVQPGGSLRLSCAASGFTFSYYAMSWVRQAPGKGLDWVSTISGGG chain sequence heavy chain GNTHYADSVKGRFTISRDNSKNTLYLHMNSLRAEDTAVYYCAKDRMKQLVRAYYF DYWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNS GALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTINDKKVEP KSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVK FNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLICLVKGFYPSDIAVEWESNGQ PENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSL SLSPGK 267 413F09-full heavy Nucleic acid sequence of 413F09 GAGGTGCCGCTGGTGGAGTCTGGGGGAGGCTTGGTACAGCCTGGGGGGTCCCT chain sequence heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACGTTTAGCTACTATGCCATGAGCTG GGTCCGTCAGGCTCCAGGGAAGGGGCTGGACTGGGTCTCAACTATTAGTGGTG GTGGTGGTAACACACACTACGCAGACTCCGTGAAGGGCCGATTCACTATATCCA GAGACAATTCCAAGAACACGCTGTATCTGCACATGAACAGCCTGAGAGCCGAAG ACACGGCCGTCTATTACTGTGCGAAGGATCGGATGAAACAGCTCGTCCGGGCCT ACTACTTTGACTACTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAGCCAGCA CCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCG GAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCG TGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGC TGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCT CTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAA GGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCC TTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAA GCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGT GGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGT GGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTA CCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAG AGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCA TCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTA GCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCT TCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACA ACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAG CAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTC CGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAG CCCCGGCAAG 268 413F09-CDRL1 Amino acid sequence of CDRL1 QDISTY (IMGT) of 413F09 using IMGT 269 413F09-CDRL2 Amino acid sequence of CDRL2 GTS (IMGT) of 413F09 using IMGT 270 413F09-CDRL3 Amino acid sequence of CDRL3 QQLHTDPIT (IMGT) of 413F09 using IMGT 271 413F09-CDRL1 Amino acid sequence of CDRL1 WASQDISTYLG (Kabat) of 413F09 using Kabat 272 413F09-CDRL2 Amino acid sequence of CDRL2 GTSSLQS (Kabat) of 413F09 using Kabat 273 413F09-CDRL3 Amino acid sequence of CDRL3 QQLHTDPIT (Kabat) of 413F09 using Kabat 274 413F09-Light chain Amino acid sequence of V.sub.L of DIQLTQSPSFLSASVGDRVTITCWASQDISTYLGWYQQKPGKAPKLIYGTSSLQS variable region 413F09 GVPSRFSGSGSGTEFTLTISSLQPEDFATYYCQQLHTDPITFGQGTRLEIK 275 413F09-Light chain Nucleic acid sequence of V.sub.L of GACATCCAGTTGACCCAGTCTCCATCCTTCCTGTCTGCATCTGTAGGAGACAGA variable region 413F09 GTCACCATCACTTGCTGGGCCAGTCAGGACATTAGCACTTATTTAGGCTGGTAT CAGCAAAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTACATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGAATTCACT CTCACAATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTATTACTGTCAACAGC TTCATACTGACCCGATCACCTTCGGCCAAGGGACACGACTGGAGATCAAAC 276 413F09-full light Amino acid sequence of 413F09 DIQLTQSPSFLSASVGDRVTITCWASQDISTYLGWYQQKPGKAPKWYGTSSLQS chain sequence light chain GVPSRFSGSGSGTEFTLTISSLQPEDFATYYCQQLHTDPITFGQGTRLEIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 277 413F09-full light Nucleic acid sequence of 413F09 GACATCCAGTTGACCCAGTCTCCATCCTTCCTGTCTGCATCTGTAGGAGACAGA chain sequence light chain GTCACCATCACTTGCTGGGCCAGTCAGGACATTAGCACTTATTTAGGCTGGTAT CAGCAAAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGGTACATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGAATTCACT CTCACAATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTATTACTGTCAACAGC TTCATACTGACCCGATCACCTTCGGCCAAGGGACACGACTGGAGATCAAACGTA CGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGT CCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCA AGGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCG TGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCC TGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACC AGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 278 414B06-CDRH1 Amino acid sequence of CDRH1 GFTFSSYW (IMGT) of 414B06 using IMGT 279 414B06-CDRH2 Amino acid sequence of CDRH2 IKQDGSEK
(IMGT) of 414B06 using IMGT 280 414B06-CDRH3 Amino acid sequence of CDRH3 ARVRQWSDYSDY (IMGT) of 414B06 using IMGT 281 414B06-CDRH1 Amino acid sequence of CDRH1 SYWMN (Kabat) of 414B06 using Kabat 282 414B06-CDRH2 Amino acid sequence of CDRH2 NIKQDGSEKYYVDSVKG (Kabat) of 414B06 using Kabat 283 414B06-CDRH3 Amino acid sequence of CDRH3 VRQWSDYSDY (Kabat) of 414B06 using Kabat 284 414B06-Heavy chain Amino acid sequence of V.sub.H of EVHLVESGGGLVQPGGSLRLSCAASGFTFSSYWMNWVRQAPGKGLEWVANIKQD variable region 414B06 GSEKYYVDSVKGRFTVSRDNAKNSLYLQMNSLRAEDTAVYYCARVRQWSDYSDY WGQGTPVTVSS 285 414B06-Heavy chain Nucleic acid sequence of V.sub.H GAGGTGCACCTGGTGGAGTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT variable region of 414B06 GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTTAGTAGCTATTGGATGAACTG GGTCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTGGCCAACATAAAGCAAG ATGGAAGTGAGAAATACTATGTGGACTCTGTGAAGGGCCGCTTCACCGTCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGGCTGTGTATTACTGTGCGAGAGTTCGACAATGGTCCGACTACTCTGACT ACTGGGGCCAGGGAACCCCGGTCACCGTCTCCTCAG 286 414B06-full heavy Amino acid sequence of 414B06 EVHLVESGGGLVQPGGSLRLSCAASGFTFSSYWMNWVRQAPGKGLEWVANIKQD chain sequence heavy chain GSEKYYVDSVKGRFTVSRDNAKNSLYLQMNSLRAEDTAVYYCARVRQWSDYSDY WGQGTPVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGA LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKS CDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN INYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCINSNKALPAP IEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL SPGK 287 414B06-full heavy Nucleic acid sequence of 414B06 GAGGTGCACCTGGTGGAGTCTGGGGGAGGCTTGGTCCAGCCTGGGGGGTCCCT chain sequence heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTTAGTAGCTATTGGATGAACTG GGTCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTGGCCAACATAAAGCAAG ATGGAAGTGAGAAATACTATGTGGACTCTGTGAAGGGCCGCTTCACCGTCTCCA GAGACAACGCCAAGAACTCACTGTATCTGCAAATGAACAGCCTGAGAGCCGAGG ACACGGCTGTGTATTACTGTGCGAGAGTTCGACAATGGTCCGACTACTCTGACT ACTGGGGCCAGGGAACCCCGGTCACCGTCTCCTCAGCCAGCACCAAGGGCCCCT CTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGCCGCTC TGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTGGAACT CTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTCCTCCG GCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGGCACCC AGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGACAAGA AGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCTGCCC CTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAGGACA CCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTGTCCC ACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTGCACA ACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTGGTGT CCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAGTGCA AGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAGGCCA AGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGACGAGC TGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCCTCCG ATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAGACCA CCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTGACAG TGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATGCACG AGGCCCTGCACAACCACTACACCCAGAAGTtCCTGTCCCTGAGCCCCGGCAAG 288 414B06-CDRL1 Amino acid sequence of CDRL1 QGISSW (IMGT) of 414606 using IMGT 289 414B06-CDRL2 Amino acid sequence of CDRL2 MS (IMGT) of 414B06 using IMGT 290 414B06-CDRL3 Amino acid sequence of CDRL3 QQANSFPFT (IMGT) of 414B06 using IMGT 291 414B06-CDRL1 Amino acid sequence of CDRL1 RASQGISSWLA (Kabat) of 414B06 using Kabat 292 414B06-CDRL2 Amino acid sequence of CDRL2 AASSLQS (Kabat) of 414B06 using Kabat 293 414B06-CDRL3 Amino acid sequence of CDRL3 QQANSFPFT (Kabat) of 414B06 using Kabat 294 414B06-Light chain Amino acid sequence of V.sub.L of DIQMTQSPSSVSASVGDRVTITCRASQGISSWLAWYQQKPGKAPKLLIYAASSLQS variable region 414B06 GVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQANSFPFTFGPGTKVDIK 295 414B06-Light chain Nucleic acid sequence of V.sub.L of GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTAGGAGACAGA variable region 414B06 GTCACCATCACTTGTCGGGCGAGTCAGGGTATTAGCAGCTGGTTAGCCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGATTTCACT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTTTCCCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAAC 296 414B06-full light Amino acid sequence of 414B06 DIQMTQSPSSVSASVGDRVTITCRASQGISSWLAWYQQKPGKAPKLIYAASSLQS chain sequence light chain GVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQANSFPFTFGPGTKVDIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAMIQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 297 414B06-full light Nucleic acid sequence of 414B06 GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTAGGAGACAGA chain sequence light chain GTCACCATCACTTGTCGGGCGAGTCAGGGTATTAGCAGCTGGTTAGCCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGATTTCACT CTCACCATCAGCAGCCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTTTCCCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAACGTAC GGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGTC CGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCAA GGTGCAGTGGAAGGTGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCGT GACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCCT GICCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACCA GGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 298 Mutated 1D05-LC Amino acid sequence of 1D05 DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYYASSLQS mutant 3 kappa light chain with V to Y GVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQSYSTPITFGQGTRLEIKRIVAAP mutation in CDRL2 highlighted SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 299 1D05-heavy chain Amino acid sequence of IgG1 EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQVPGKGLEWVSGISWI disabled IgG1 Fc disabled variant of 1D05 RTGIGYADSVKGRFTIFRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGGW FDTVVGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWN SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTINDKINE PKSCDKTHTCPPCPAPELAGAPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPE VKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK 300 1D05-light chain IL- 1D05 Light chain sequence DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNVVYQQKPGKAPKLLIYVASSLQS 2 fusion fused to wild-type human IL-2 GVPSRFSGSGSGTDFILTISSLQPEDFATYYCQQSYSTPITFGQGTRLEIKRIVAAP sequence (IL-2 amino acid SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK sequence is underlined and DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGECAPTSSSTIGCT region to be varied is shown in QLQLEHLLLDLQMILNGINNYKNPKLTRMLTFKFYMPKKATELKHLQCLEEELKPL bold) EEVLNLAQSKNFHLRPRDLISNINVIVLELKGSETTFMCEYADETATIVEFLNRWITF CQSIISTLT 301 Human IL-2 Uniprot number: P60568 APTSSSTKKTQLQLEHLLLDLQMILNGINNYKNPKLTRMLTFKFYMPKKATELK Full length amino acid sequence HLQCLEEELKPLEEVLNLAQSKNFHLRPRDLISNINVIVLELKGSETTFMCEYADETA of human IL-2 (minus signal TIVEFLNRWITFCQSIISTLT sequence) 302 Control 1D05 Heavy chain 1D05 IgG1 variant EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQVPGKGLEWVSGISWI immunocytokine HC fused at the N-terminus to RTGIGYADSVKGRFTIFRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGGW C-terminal fusion wild-type human IL2 sequence FDTWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWN (control) SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKIWE PKSCDKTHTCPPCPAPELAGAPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPE VITNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGKAPTSSSTKKTQLQLEHLLLDLQMILNGINNYKNPKLTRMLTFKFYMP KKATELKHLQCLEEELKPLEEVLNLAQSKNFHLRPRDLISNINVIVLELKGSETTFMC EYADETATIVEFLNRWITFCQSIISTLT 303 IL-2 D5-9 IL-2 IC45 (Del 5-9) N terminal APTSTQLQLELLLD IL-2 sequence 304 IL-2 D1-9 IL-2 IC46 (Del 1-9) N terminal TQLQLEHLLLD IL-2 sequence 305 IL-2 D5-7 IL-2 IC64 (Del 5-7) N terminal APTSKKTQLQLEHLLLD IL-2 sequence 306 IL-2 D1 IL-2 D1 N terminal IL-2 PTSSSTKKTQLQLEHLLLD sequence 307 IL-2 D1-2 IL-2 D1-2 N terminal IL-2 TSSSTKICTQLQLEHLLLD sequence 308 IL-2 D1-3 IL-2 D1-3 N terminal IL-2 SSSTKKTQLQLEHLLLD sequence 309 IL-2 D1-4 IL-2 D1-4 N terminal IL-2 SSTKKTQLQLEHLLLD sequence 310 IL-2 D1-5 IL-2 D1-5 N terminal IL-2 STKKTQLQLEHLLLD sequence 311 IL-2 D1-6 IL-2 D1-6 N terminal IL-2 TKKTQLQLEHLLLD sequence 312 IL-2 D1-7 IL-2 D1-7 N terminal IL-2 KKTQLQLEHLLLD sequence 313 IL-2 D1-8 IL-2 D1-8 N terminal IL-2 KTQLQLEHLLLD sequence 314 IL-2 D9 IL-2 D9 N terminal IL-2 APTSSSTKTQLQLEHLLLD sequence 315 IL-2 D9-8 IL-2 D9-8 N terminal IL-2 APTSSSTTQLQLEHLLLD sequence 316 IL-2 D9-7 IL-2 D9-7 N terminal IL-2 APTSSSTQLQLEHLLLD sequence 317 IL-2 D9-6 IL-2 D9-6 N terminal IL-2 APTSSTQLQLEHLLLD sequence 318 IL-2 D9-4 IL-2 D9-4 N terminal IL-2 APTTQLQLEHLLLD sequence 319 IL-2 D9-3 IL-2 D9-3 N terminal IL-2 APTQLQLEHLLLD sequence 320 IL-2 D9-2 IL-2 D9-2 N terminal IL-2 ATQLQLEHLLLD sequence 321 IL-2 D2-6 IL-2 D2-6 N terminal IL-2 ATKKTQLQLEHLLLD sequence 322 IL-2 D3-7 IL-2 D3-7 N terminal IL-2 APKKTQLQLEHLLLD sequence 323 IL-2 D4-8 IL-2 D4-8 N terminal IL-2 APTKTQLQLEHLLLD sequence 324 C-terminal amino acid Amino acids 21 to 133 of hIL-2 LQMILNGINNYKNPKLTRMLTFKFYMPKKATELKHLQCLEEELKPLEEVLNLAQSKN sequence of hIL-2 FHLRPRDLISNINVIVLELKGSETTFMCEYADETATIVEFLNRWITFCQSIISTLT 325 Mouse PD-L1 Uniprot number: Q9EP73 MRIFAGIIFTACCHLLRAFTITAPKDLYVVEYGSNVTMECRFPVERELDLLAL (ECD highlighted in BOLD, and VVYWEKEDEQVIQFVAGEEDLKPQHSNFRGRASLPKDQLLKGNAALQIT cytoplasmic domain underlined)
DVKLQDAGVYCCIISYGGADYKRITLKVNAPYRKINQRISVDPATSEHEL ICQAEGYPEAEVIWTNSDHQPVSGKRSVTTSRTEGMLLNVTSSLRVNAT ANDVFYCTFWRSQPGQNHTAELIIPELPATHPPQNRTHWVLLGSILLFLIW STVLLFLRKQVRMLDVEKCGVEDTSSKNRNDTQFEET 326 Mouse PD-L1 ECD His Mouse PD-L1 extracellular FTITAPKDLYVVEYGSNVTMECRFPVERELDLLALWYWEKEDEQVIQFVAGEEDLK domain with his tag PQHSNFRGRASLPKDQLLKGNAALQITDVKLQDAGVYCCIISYGGADYKRITLKVN APYRKINQRISVDPATSEHELICQAEGYPEAEVIWTNSDHQPVSGKRSVTTSRTEG MLLNVTSSLRVNATANDVFYCTFWRSQPGQNHTAELIIPELPATHPPQNRTHHHH HH 327 Human IL-2R.alpha. chain Human IL-2 receptor alpha ELCDDDPPEIPHATFKAMAYKEGTMLNCECKRGFRRIKSGSLYMLCIGNSSHSSW chain DNQCQCTSSAIRNTTKQVTPQPEEQKERKTTEMQSPMQPVDQASLPGHCREPPP WENEATERIYHFVVGQMVYYQCVQGYRALHRGPAESVCKMTHGKTRWTQPQLIC TGEMETSQFPGEEKPQASPEGRPESETSCLVTTTDFQIQTEMAATMETSIFTTEYQ VAVAGCVFLLISVLLLSGLTVVQRRQRKSRRTI 328 Human IL-2R.beta. chain Human IL-2 receptor beta chain AVNGTSQFTCFYNSRANISCVWSQDGALQDTSCQVHAWPDRRRWNQTCELLPVS QASWACNLILGAPDSQKLTTVDIVTLRVLCREGVRWRVMAIQDFKPFENLRLMAPI SLQVVHVETHRCNISWEISQASHYFERHLEFEARTLSPGHIWEEAPLLTLKQKQE WICLETLTPDTQYEFQVRVKPLQGEFTTWSPWSQPLAFRTKPAALGKDTIPWLGH LLVGLSGAFGFIILVYLLINCRNTGPWLKKVLKCNTPDPSKFFSQLSSEHGGDVQKW LSSPFPSSSFSPGGLAPEISPLEVLERDINTQLLLQQDKVPEPASLSSNHSLTSCFTN QGYFFFHLPDALEIEACQVYFTYDPYSEEDPDEGVAGAPTGSSPQPLQPLSGEDDA YCTFPSRDDLLLFSPSLLGGPSPPSTAPGGSGAGEERMPPSLQERVPRDWDPQPLG PPTPGVPDLVDFQPPPELVLREAGEEVPDAGPREGVSFPWSRPPGQGEFRALNARL PLNTDAYLSLQELQGQDPTHLV 329 Human IL-2R.gamma. chain Human IL-2 receptor common LNTTILTPNGNEDTTADFFLTTMPTDSLSVSTLPLPEVQCFVFNVEYMNCTWNSSS gamma chain EPQPTNLTLHYWYKNSDNDINQKCSHYLFSEEITSGCQLQKKEIHLYQTFVVQLQD PREPRRQATQMLKLQNLVIPWAPENLTLHKLSESQLELNWNNRFLNHCLEHLVQY RTDWDHSWTEQSVDYRHKFSLPSVDGQKRYTFRVRSRFNPLCGSAQHWSEWSH PIHWGSNTSKENPFLFALEAVVISVGSMGLIISLLCVYFWLERTMPRIPTLKNLEDLV TEYHGNFSAWSGVSKGLAESLQPDYSERLCLVSEIPPKGGALGEGPGASPCNQHSP YWAPPCYTLKPET 330 IL-7 Human IL-7 amino acid DCDIEGKDGKQYESVLMVSIDQLLDSMKEIGSNCLNNEFNFFKRHICDANKEGMFL sequence FRAARKLRQFLKMNSTGDFDLHLMSEGTTILLNCTGQVKGRKPAALGEAQPTKS LEENKSLKEQKKLNDLCFLKRLLQEIKTCWNKILMGTKEH 331 IL-15 Human IL-15 amino acid GIHVFILGCFSAGLPKTEANINVNVISDLKKIEDLIQSMHIDATLYTESDVHPSCKVT sequence AMKCFLLELQVISLESGDASIHDTVENLIILANNSLSSNGNVTESGCKECEELEEKNI KEFLQSFVHIVQMFINTS 332 IL-21 Human IL-21 amino acid QGQDRHMIRMRQLIDIVDQLKNYVNDLVPEFLPAPEDVETNCEWSAFSCFQKAQL sequence KSANTGNNERIINVSIKKLKRKPPSTNAGRRQKHRLTCPSCDSYEKKPPKEFLERFK SLLQKMIHQHLSSRTHGSEDS 333 GM-CSF Human GM-CSF amino acid APARSPSPSTQPWEHVNAIQEARRLLNLSRDTAAEMNETVEVISEMFDLQEPTCLQ sequence TRLELYKQGLRGSLTKLKGPLTMMASHYKQHCPPTPETSCATQIITFESFKENLKDF LLVIPFDCWEPVQE 334 IFN.alpha. Human IFN-.alpha. amino acid CDLPQNHGLLSRNTLVLLHQMRRISPFLCLKDRRDFRFPQEMVKGSQLQKAHVMS sequence VLHEMLQQIFSLFHTERSSAAWNMTLLDQLHTELHQQLQHLETCLLQVVGEGESA GAISSPALTLRRYFQGIRVYLKEKKYSDCAWEVVRMEIMKSLFLSTNMQERLRSKD RDLGS 335 TNF.alpha. Extracellular portion of human GPQREEFPRDLSLISPLAQAVRSSSRTPSDKPVAHVVANPQAEGQLQWLNRRANAL TNF-.alpha. amino acid sequence LANGVELRDNQLVVPSEGLYLIYSQVLFKGQGCPSTHVLLTHTISRIAVSYQTWNL LSAIKSPCQRETPEGAEAKPWYEPIYLGGVFQLEKGDRLSAEINRPDYLDFAESGQV YFGIIAL 336 IL-12.alpha. Alpha chain of human IL-12 RNLPVATPDPGMFPCLHHSQNLLRAVSNMLQKARQTLEFYPCTSEEDHEDITKDK amino acid sequence TSTVEACLPLELTKNESCLNSRETSFITNGSCLASRKTSFMMALCLSSIYEDLKMYQ VEFKTMNAKLLMDPKRQIFLDQNMLAVIDELMQALNFNSETVPQKSSLEEPDFYKT KIKLCILLHAFRIRAVTIDRVMSYLNAS 337 IL-12.beta. Beta chain of human IL-12 IWELKKDVYVVELDWYPDAPGEMVVLTCDTPEEDGMATTLDQSSEVLGSGKILTI amino acid sequence QVKEFGDAGQYTCHKGGEVLSHSLLLLHKKEDGIWSTDILKDQKEPKNKTFLRCEA KNYSGRFTCWWLTTISTDLTFSVIGSRGSSDPQGVTCGAATLSAERVRGDNKEYE YSVECQEDSACPAAEESLPIEVMVDAVHKLKYENYTSSFFIRDIIKPDPPKNLQLKPL KNSRQVEVSWEYPDTTSTPHSYFSLTFCVQVQGKSKREKKDRVFTDKTSATVICR KNASISVRAQDRYYSSSWSEWASVPCS 338 CXCL9 Human CXCL-9 amino acid TPWRKGRCSCISTNQGTIHLQSLKDLKQFAPSPSCEMEIIATLKNGVQTCLNPDSA sequence DVKELIKKWEKQVSQKKKQKNGKKHQKKKVLIMKSQRSRQKKTT 339 CXCL10 Human CXCL-10 amino acid VPLSRTVRCTCISISNQPVNPRSLEKLEIIPASQFCPRVEIIATMKKKGEKRCLNPESK sequence AIKNLLKAVSKERSKRSP 340 Human WT IGHG1* WT human IgG1 amino acid ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL IgG1 01 & sequence QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCP constant IGHG1* APELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN region 02 & AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQ IGHG1* PREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVL 05 DSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (IgG1) 341 WT human IgG1 nucleic acid GCCAGCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACC sequence TCTGGCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCT GTGACCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCT GCTGTGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCT TCCAGCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCA ACACCAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCT GTCCCCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCC CCCCAAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCG TGGTGGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGG ACGGCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACT CCACCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACG GCAAAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAA AGACCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGC CCCCTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGA AAGGCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTG AGAACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCT GTACAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTC CTGCTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTC CCTGAGCCCCGGCAAGTGATGA 342 Mutated 1D05-HC Amino acid sequence of 1D05 EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWI mutant 2 heavy chain with V to A and F to RTGIGYADSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCAKDMKGSGTYGGW S back-mutation in framework FDTWGQGTLVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWN region to germline highlighted SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRV with IgG1 disabled (LAGA) ESKYGPPCPPCPAPELAGAPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQF constant region NWYVDGVEVHNAKTKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCINSNKGLP SSIEKTISMKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNG QPENNYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKS LSLSLGK 343 416E01-CDRH1 Amino acid sequence of CDRH1 GFTFSNYA (IMGT) of 416E01 using IMGT 344 416E01-CDRH2 Amino acid sequence of CDRH2 ISFSGGTT (IMGT) of 416E01 using IMGT 345 416E01-CDRH3 Amino acid sequence of CDRH3 AKDEAPAGATFFDS (IMGT) of 416E01 using IMGT 346 416E01-CDRH1 Amino acid sequence of CDRH1 NYAMS (Kabat) of 416E01 using Kabat 347 416E01-CDRH2 Amino acid sequence of CDRH2 AISFSGGTTYYADSVKG (Kabat) of 416E01 using Kabat 348 416E01-CDRH3 Amino acid sequence of CDRH3 DEAPAGATFFDS (Kabat) of 416E01 using Kabat 349 416E01-Heavy chain Amino acid sequence of V.sub.H of EVQLAESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQTPGKGLEWVSAISFSG variable region 416E01 (mutations from GTTYYADSVKGRFTISRDNSKNTLYLHMNSLRADDTAVYYCAKDEAPAGATFFD germline are shown in SWGQGTLVTVSS boldletters) 350 416E01-Heavy chain Nucleic acid sequence of V.sub.H of GAAGTGCAACTGGCGGAGTCTGGGGGAGGCTTGGTACAGCCGGGGGGGTCCCT variable region 416E01 GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTTAGCAACTATGCCATGAGTTG GGTCCGCCAGACTCCAGGAAAGGGGCTGGAGTGGGTCTCAGCTATTAGTTTTAG TGGTGGTACTACATACTACGCTGACTCCGTGAAGGGCCGGTTCACCATCTCCAG AGACAATTCCAAGAACACGCTGTATTTGCACATGAACAGCCTGAGAGCCGATGA CACGGCCGTATATTACTGTGCGAAAGATGAGGCACCAGCTGGCGCAACCTTCTT TGACTCCTGGGGCCAGGGAACGCTGGTCACCGTCTCCTCAG 351 416E01-full heavy Amino acid sequence of 416E01 EVQLAESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQTPGKGLEWVSAISFSG chain sequence heavy chain GTTNADSVKGRFTISRDNSKNTLYLHMNSLRADDTAVYYCAKDEAPAGATFFDS WGQGTLVTVSSASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGA LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTKTYTCNVDHKPSNTKVDKRVESKY GPPCPPCPAPEFEGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQFNWY VDGVEVHNAKTKPREEQFNSTYRWSVLTVLHQDWLNGKEYKCINSNKGLPSSIEK TISKAKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENN YKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSLSLSLG K 352 416E01-full heavy Nucleic acid sequence of 416E01 GAAGTGCAACTGGCGGAGTCTGGGGGAGGCTTGGTACAGCCGGGGGGGTCCCT chain sequence heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTTAGCAACTATGCCATGAGTTG GGTCCGCCAGACTCCAGGAAAGGGGCTGGAGTGGGTCTCAGCTATTAGTTTTAG TGGTGGTACTACATACTACGCTGACTCCGTGAAGGGCCGGTTCACCATCTCCAG AGACAATTCCAAGAACACGCTGTATTTGCACATGAACAGCCTGAGAGCCGATGA CACGGCCGTATATTACTGTGCGAAAGATGAGGCACCAGCTGGCGCAACCTTCTT TGACTCCTGGGGCCAGGGAACGCTGGTCACCGTCTCCTCAGCCAGCACCAAGGG CCCTTCCGTGTTCCCCCTGGCCCCTTGCAGCAGGAGCACCTCCGAATCCACAGC TGCCCTGGGCTGTCTGGTGAAGGACTACTTTCCCGAGCCCGTGACCGTGAGCTG GAACAGCGGCGCTCTGACATCCGGCGTCCACACCTTTCCTGCCGTCCTGCAGTC CTCCGGCCTCTACTCCCTGTCCTCCGTGGTGACCGTGCCTAGCTCCTCCCTCGG CACCAAGACCTACACCTGTAACGTGGACCACAAACCCTCCAACACCAAGGTGGAC AAACGGGTCGAGAGCAAGTACGGCCCTCCCTGCCCTCCTTGTCCTGCCCCCGAG TTCGAAGGCGGACCCAGCGTGTTCCTGTTCCCTCCTAAGCCCAAGGACACCCTC ATGATCAGCCGGACACCCGAGGTGACCTGCGTGGTGGTGGATGTGAGCCAGGA GGACCCTGAGGTCCAGTTCAACTGGTATGTGGATGGCGTGGAGGTGCACAACG CCAAGACAAAGCCCCGGGAAGAGCAGTTCAACTCCACCTACAGGGTGGTCAGCG TGCTGACCGTGCTGCATCAGGACTGGCTGAACGGCAAGGAGTACAAGTGCAAG GTCAGCAATAAGGGACTGCCCAGCAGCATCGAGAAGACCATCTCCAAGGCTAAA GGCCAGCCCCGGGAACCTCAGGTGTACACCCTGCCTCCCAGCCAGGAGGAGATG ACCAAGAACCAGGTGAGCCTGACCTGCCTGGTGAAGGGATTCTACCCTTCCGAC ATCGCCGTGGAGTGGGAGTCCAACGGCCAGCCCGAGAACAATTATAAGACCACC CCTCCCGTCCTCGACAGCGACGGATCCTTCTTTCTGTACTCCAGGCTGACCGTG GATAAGTCCAGGTGGCAGGAAGGCAACGTGTTCAGCTGCTCCGTGATGCACGAG GCCCTGCACAATCACTACACCCAGAAGTCCCTGAGCCTGTCCCTGGGAAAG 353 416E01-CDRL1 Amino acid sequence of CDRL1 QGIRRW (IMGT) of 416E01 using IMGT 354 416E01-CDRL2 Amino acid sequence of CDRL2 GAS (IMGT) of 416E01 using IMGT 355 416E01-CDRL3 Amino acid sequence of CDRL3 QQANSFPIT (IMGT) of 416E01 using IMGT 356 416E01-CDRL1 Amino acid sequence of CDRL1 RASQGIRRWLA (Kabat) of 416E01 using Kabat 357 416E01-CDRL2 Amino acid sequence of CDRL2 GASSLQS (Kabat) of 416E01 using Kabat 358 416E01-CDRL3 Amino acid sequence of CDRL3 QQANSFPIT (Kabat) of 416E01 using Kabat 359 416E01-Light chain Amino acid sequence of VL of DIQMTQSPSSVSASVGDRVTITCRASQGIRRWLAWYQQKPGKAPKLISGASSLQ variable region 416E01 (mutations from SGVPSRFSGSGSGTDFTLIITSLQPEDFATYYCQQANSFPITFGQGTRLEIK germline are shown in bold letters) 360 416E01-Light chain Nucleic acid sequence of VL of GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTAGGAGACAGA variable region 416E01 GTCACCATCACTTGTCGGGCGAGTCAGGGTATTAGGAGGTGGTTAGCCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAACTCCTGATCTCTGGTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGATTTCACT CTCATCATTACCAGTCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG
CTAACAGTTTCCCGATCACCTTCGGCCAAGGGACACGACTGGAGATCAAAC 361 416E01-full light Amino acid sequence of 416E01 DIQMTQSPSSVSASVGDRVTITCRASQGIRRWLAWYQQKPGKAPKLISGASSLQS chain sequence light chain GVPSRFSGSGSGTDFTLIITSLQPEDFATYYCQQANSFPITFGQGTRLEIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 362 416E01-full light Nucleic acid sequence of 416E01 GACATCCAGATGACCCAGTCTCCATCTTCCGTGTCTGCATCTGTAGGAGACAGA chain sequence light chain GTCACCATCACTTGTCGGGCGAGTCAGGGTATTAGGAGGTGGTTAGCCTGGTAT CAGCAGAAACCAGGGAAAGCCCCTAAACTCCTGATCTCTGGTGCATCCAGTTTG CAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGATTTCACT CTCATCATTACCAGTCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAACAGG CTAACAGTTTCCCGATCACCTTCGGCCAAGGGACACGACTGGAGATCAAACGTA CGGTGGCCGCTCCCTCCGTGTTCATCTTCCCACCTTCCGACGAGCAGCTGAAGT CCGGCACCGCTTCTGTCGTGTGCCTGCTGAACAACTTCTACCCCCGCGAGGCCA AGGTGCAGTGGAAGGIGGACAACGCCCTGCAGTCCGGCAACTCCCAGGAATCCG TGACCGAGCAGGACTCCAAGGACAGCACCTACTCCCTGTCCTCCACCCTGACCC TGTCCAAGGCCGACTACGAGAAGCACAAGGTGTACGCCTGCGAAGTGACCCACC AGGGCCTGTCTAGCCCCGTGACCAAGTCTTTCAACCGGGGCGAGTGT 363 STIM001-CDRH1 Amino acid sequence of CDRH1 GYTFSTFG of STIM001 using IMGT 364 STIM001-CDRH2 Amino acid sequence of CDRH2 ISAYNGDT of STIM001 using IMGT 365 STIM001-CDRH3 Amino acid sequence of CDRH3 ARSSGHYYYYGMDV of STIM001 using IMGT 366 STIM001-Heavy Amino acid sequence of V.sub.H of QVQVVQSGAEVKKPGASVKVSCKASGYTFSTFGITWVRQAPGQGLEWMGWISAY chain variable region STIM001 NGDTNYAQNLQGRVIMTTDTSTSTAYMELRSLRSDDTAVYYCARSSGHYYYYGM DVWGQGTTVTVSS 367 STIM001-Heavy Nucleic acid sequence of V.sub.H of CAGGTTCAGGTGGTGCAGTCTGGAGCTGAGGTGAAGAAGCCTGGGGCCTCAGT chain variable region STIM001 GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTTCCACCTTTGGTATCACCTGG GTGCGACAGGCCCCTGGACAAGGGCTTGAATGGATGGGATGGATCAGCGCTTA CAATGGTGACACAAACTATGCACAGAATCTCCAGGGCAGAGTCATCATGACCAC AGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCCTGAGATCTGACGA CACGGCCGTTTATTACTGTGCGAGGAGCAGTGGCCACTACTACTACTACGGTAT GGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCA 368 STIM001-full heavy Amino acid sequence of QVQVVQSGAEVKKPGASMSCKASGYTFSTFGITWVRQAPGQGLEWMGWISAY chain sequence STIM001 heavy chain NGDTNYAQNLQGRVIMTTDTSTSTAYMELRSLRSDDTAVYYCARSSGHYYYYGM DVWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWN SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHITSNTKVDKIWE PKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEV KFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKOWSNKA LPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQ KSLSLSPGK 369 STIM001-full heavy Nucleic acid sequence of CAGGTTCAGGTGGTGCAGTCTGGAGCTGAGGTGAAGAAGCCTGGGGCCTCAGT chain sequence STIM001 heavy chain GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTTCCACCTTTGGTATCACCTGG GTGCGACAGGCCCCTGGACAAGGGCTTGAATGGATGGGATGGATCAGCGCTTA CAATGGTGACACAAACTATGCACAGAATCTCCAGGGCAGAGTCATCATGACCAC AGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCCTGAGATCTGACGA CACGGCCGTTTATTACTGTGCGAGGAGCAGTGGCCACTACTACTACTACGGTAT GGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCCAGCACCAAGGG CCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCGGAACAGC CGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCGTGTCCTG GAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGCTGCAGTC CTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCTCTCTGGG CACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAAGGTGGAC AAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCCTTGTCCT GCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAAGCCCAAG GACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGTGGATGTG TCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGTGGAAGTG CACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTACCGGGTG GTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAGAGTACAAG TGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCATCTCCAAG GCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTAGCAGGGAC GAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCTTCTACCCC TCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACAACTACAAG ACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAGCAAGCTG ACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTCCGTGATG CACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAGCCCCGGC AAGTGATGA 370 STIM001-CDRL1 Amino acid sequence of CDRL1 QSLLHSNEYNY of STIM001 using IMGT 371 STIM001-CDRL2 Amino acid sequence of CDRL2 LGS of STIM001 using IMGT 372 STIM001-CDRL3 Amino acid sequence of CDRL3 MQSLQTPLT of STIM001 using IMGT 373 STIM001-Light chain Amino acid sequence of VL of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSNEYNYLDWYLQKPGQSPQLLIFLGS variable region STIM001 NRASGVPDRFSGSGSGTDFTLKITRVEAEDVGIYYCMQSLQTPLTFGGGTKVEIK 374 STIM001-Light chain Nucleic acid sequence of VL of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG variable region STIM001 GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTAATGAATACAACT ATTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTTTT TGGGTTCTAATCGGGCCTCCGGGGTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCACCAGAGTGGAGGCTGAGGATGTTGGAATT TATTACTGCATGCAATCTCTACAAACTCCGCTCACTTTCGGCGGAGGGACCAAG GTGGAGATCAAA 375 STIM001-full light Amino acid sequence of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSNEYNYLDWYLQKPGQSPQLLIFLGS chain sequence STIM001 light chain NRASGVPDRFSGSGSGTDFTLKITRVEAEDVGIYYCMQSLQTPLTFGGGTKVEIK RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWINDNALQSGNSQESV TEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 376 STIM001-full light Nucleic acid sequence of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG chain sequence STIM001 light chain GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTAATGAATACAACT ATTIGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTTTT TGGGTICTAATCGGGCCTCCGGGGICCCTGACAGGTICAGIGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCACCAGAGTGGAGGCTGAGGATGTTGGAATT TATTACTGCATGCAATCTCTACAAACTCCGCTCACTTTCGGCGGAGGGACCAAG GIGGAGATCAAAcgtacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaa gtccggcaccgcttctgtcgtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaag gtggacaacgccctgcagtccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcac ctactccctgtcctccaccctgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaa gtgacccaccaqggcctgtctagccccgtgaccaatctttcctgggcgagtgt 377 STIM002-CDRH1 Amino acid sequence of CDRH1 GYTFTSYG of STIM002 using IMGT 378 STIM002-CDRH2 Amino acid sequence of CDRH2 ISAYNGNT of STIM002 using IMGT 379 STIM002-CDRH3 Amino acid sequence of CDRH3 ARSTYFYGSGTLYGMDV of STIM002 using IMGT 380 STIM002-Heavy Amino acid sequence of V.sub.H of QVQLVQSGGEVKKPGASVKVSCKASGYTFTSYGFSWVRQAPGQGLEWMGWISAY chain variable region STIM002 NGNTNYAQKLQGRVTMTTDTSTSTAYMELRSLRSDDTAVYYCARSTYFYGSGTLY GMDVWGQGTTVTVSS 381 STIM002-Heavy Nucleic acid sequence of V.sub.H of CAGGTTCAACTGGTGCAGTCTGGAGGTGAGGTGAAGAAGCCTGGGGCCTCAGT chain variable region STIM002 GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTACCAGCTATGGTTTCAGCTG GGTGCGACAGGCCCCTGGACAAGGACTAGAGTGGATGGGATGGATCAGCGCTT ACAATGGTAACACAAACTATGCACAGAAGCTCCAGGGCAGAGTCACCATGACCA CAGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCTTGAGATCTGACG ACACGGCCGTGTATTACTGTGCGAGATCTACGTATTTCTATGGTTCGGGGACCC TCTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCA 382 STIM002-full heavy Amino acid sequence of QVQLVQSGGEVKKPGASVKVSCKASGYTFTSYGFSWVRQAPGQGLEWMGWISAY chain sequence STIM002 heavy chain NGNTNYAQKLQGRVTMTTDTSTSTAYMELRSLRSDDTAVYYCARSTYFYGSGTLY GMDVWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVS WNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDK KVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHED PEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCINS NKALPAPIEKTISMKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEW ESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHY TQKSLSLSPGK 383 STIM002-full heavy Nucleic acid sequence of CAGGTTCAACTGGTGCAGTCTGGAGGTGAGGTGAAGAAGCCTGGGGCCTCAGT chain sequence STIM002 heavy chain GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTACCAGCTATGGTTTCAGCTG GGTGCGACAGGCCCCTGGACAAGGACTAGAGTGGATGGGATGGATCAGCGCTT ACAATGGTAACACAAACTATGCACAGAAGCTCCAGGGCAGAGTCACCATGACCA CAGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCTTGAGATCTGACG ACACGGCCGTGTATTACTGTGCGAGATCTACGTATTTCTATGGTTCGGGGACCC TCTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCA GCCAGCACCAAGGGCCCCTCTGTGTTCCCCTGGCCCCTTCCAGCAAGTCCACC TCTGGCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCT GTGACCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCT GCTGTGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCT TCCAGCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCA ACACCAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCT GTCCCCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCC CCCCAAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCG TGGTGGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGG ACGGCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACT CCACCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACG GCAAAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAA AGACCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGC CCCCTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGA AAGGCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTG AGAACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCT GTACAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTC CTGCTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTC CCTGAGCCCCGGCAAGTGATGA 384 STIM002-CDRL1 Amino acid sequence of CDRL1 QSLLHSDGYNY of STIM002 using IMGT 385 STIM002-CDRL2 Amino acid sequence of CDRL2 LGS of STIM002 using IMGT 386 STIM002-CDRL3 Amino acid sequence of CDRL3 MQALQTPLS of STIM002 using IMGT 387 STIM002-Light chain Amino acid sequence of VL of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSDGYNYLDWYLQKPGQSPQLLIYLGS variable region STIM002 TRASGFPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPLSFGQGTKLEIK 388 STIM002-Light chain Nucleic acid sequence of VL of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG variable region STIM002 GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTGATGGATACAAC TGTTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTAT TTGGGTTCTACTCGGGCCTCCGGGTTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCGTGCAGTTTTGGCCAGGGGACCAAG CTGGAGATCAAA 389 STIM002-full light Amino acid sequence of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSDGYNYLDWYLQKPGQSPQLLIYLGS chain sequence STIM002 light chain TRASGFPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPLSFGQGTKLEIK RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAINQWKVDNALQSGNSQESV TEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 390 STIM002-full light Nucleic acid sequence of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG chain sequence STIM002 light chain GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTGATGGATACAAC TGTTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTAT TTGGGTTCTACTCGGGCCTCCGGGTTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCGTGCAGTTTTGGCCAGGGGACCAAG CTGGAGATCAAAcgtacggtggccgctccctccgtgtcatcttcccaccttccgacgagcagctgaa gtccggcaccgcttctgtcgtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaag gtggacaacgccctgcagtccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcac ctactccctgtcctccaccctgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaa gtgacccaccagggcctgtctagccccgtgaccaagtctttcaaccggggcgagtgt 391 STIM002-B-CDRH1 Amino acid sequence of CDRH1 GYTFTSYG of STIM002-B using IMGT 392 STIM002-B-CDRH2 Amino acid sequence of CDRH2 ISAYNGNT of STIM002-B using IMGT
393 STIM002-B-CDRH3 Amino acid sequence of CDRH3 ARSTYFYGSGTLYGMDV of STIM002-B using IMGT 394 STIM002-B-Heavy Amino acid sequence of VH of QVQLVQSGGEVKKPGASMSCKASGYTFTSYGFSWVRQAPGQGLEWMGWISAY chain variable region STIM002-B NGNTNYAQKLQGRVTMTIDTSTSTAYMELRSLRSDDTAVYYCARSTYFYGSGTLY GMDVWGQGTTVTVSS 395 STIM002-B-Heavy Nucleic acid sequence of VH of CAGGTTCAACTGGTGCAGTCTGGAGGTGAGGTGAAGAAGCCTGGGGCCTCAGT chain variable region STIM002-B GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTACCAGCTATGGTTTCAGCTG GGTGCGACAGGCCCCTGGACAAGGACTAGAGTGGATGGGATGGATCAGCGCTT ACAATGGTAACACAAACTATGCACAGAAGCTCCAGGGCAGAGTCACCATGACCA CAGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCTTGAGATCTGACG ACACGGCCGTGTATTACTGTGCGAGATCTACGTATTTCTATGGTTCGGGGACCC TCTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCA 396 STIM002-B-full Amino acid sequence of QVQLVQSGGEVKKPGASMSCKASGYTFTSYGFSWVRQAPGQGLEWMGWISAY heavy chain sequence STIM002-B heavy chain NGNTNYAQKLQGRVTMTTDTSTSTAYMELRSLRSDDTAVYYCARSTYFYGSGTLY GMDVWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVS WNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDK WEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHED PEVKFNWYVDGVEVHNAKTKPREEQYNSTYRWSVLTVLHQDWLNGKEYKCKVS NKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEW ESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHY TQKSLSLSPGK 397 STIM002-B-full Nucleic acid sequence of CAGGTTCAACTGGTGCAGTCTGGAGGTGAGGTGAAGAAGCCTGGGGCCTCAGT heavy chain sequence STIM002-B heavy chain GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTACCAGCTATGGTTTCAGCTG GGTGCGACAGGCCCCTGGACAAGGACTAGAGTGGATGGGATGGATCAGCGCTT ACAATGGTAACACAAACTATGCACAGAAGCTCCAGGGCAGAGTCACCATGACCA CAGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCTTGAGATCTGACG ACACGGCCGTGTATTACTGTGCGAGATCTACGTATTTCTATGGTTCGGGGACCC TCTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCCA GCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTG GCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGA CCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTG TGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCA GCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACAC CAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCC CCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCC AAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGT GGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACG GCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCA CCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCA AAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGA CCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCC CTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAG GCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGA ACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTA CAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTG CTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCT GAGCCCCGGCAAGTGATGA 398 STIM002-B-CDRL1 Amino acid sequence of CDRL1 QSLLHSDGYNC of STIM002-B using IMGT 399 STIM002-B-CDRL2 Amino acid sequence of CDRL2 LGS of STIM002-B using IMGT 400 STIM002-B-CDRL3 Amino acid sequence of CDRL3 MQALQTPCS of STIM002-B using IMGT 401 STIM002-B-Light Amino acid sequence of V.sub.L of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSDGYNCLDWYLQKPGQSPQLLIYLGS chain variable region STIM002-B TRASGFPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPCSFGQGTKLEIK 402 STIM002-B-Light Nucleic acid sequence of V.sub.L of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG chain variable region STIM002-B GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTGATGGATACAAC TGTTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTAT TTGGGTTCTACTCGGGCCTCCGGGTTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCGTGCAGTTTTGGCCAGGGGACCAAG CTGGAGATCAAA 403 STIM002-B-full light Amino acid sequence of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSDGYNCLDWYLQKPGQSPQLLIYLGS chain sequence STIM002-B light chain TRASGFPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPCSFGQGTKLEIK RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESV TEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 404 STIM002-B-full light Nucleic acid sequence of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG chain sequence STIM002-B light chain GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTGATGGATACAAC TGTTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTAT TTGGGTTCTACTCGGGCCTCCGGGTTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCGTGCAGTTTTGGCCAGGGGACCAAG CTGGAGATCAAAcgtacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaa gtccggcaccgcttctgtcgtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaag gtggacaacgccctgcagtccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcac ctactccctgtcctccaccctgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaa gtgacccaccagggcctgtctagccccgtgaccaagtctttcaaccqgggcgaqtgt 405 STIM003-CDRH1 Amino acid sequence of CDRH1 GVTFDDYG of STIM003 using IMGT 406 STIM003-CDRH2 Amino acid sequence of CDRH2 INWNGGDT of STIM003 using IMGT 407 STIM003-CDRH3 Amino acid sequence of CDRH3 ARDFYGSGSYYHVPFDY of STIM003 using IMGT 408 STIM003-Heavy Amino acid sequence of VH of EVQLVESGGGWRPGGSLRLSCVASGVTFDDYGMSWVRQAPGKGLEWVSGINWN chain variable region STIM003 GGDTDYSDSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCARDFYGSGSYYHV PFDYWGQGILVTVSS 409 STIM003-Heavy Nucleic acid sequence of VH of GAGGTGCAGCTGGTGGAGTCTGGGGGAGGTGTGGTACGGCCTGGGGGGTCCCT chain variable region STIM003 GAGACTCTCCTGTGTAGCCTCTGGAGTCACCTTTGATGATTATGGCATGAGCTG GGTCCGCCAAGCTCCAGGGAAGGGGCTGGATTGGGTCTCTGGTATTAATTGGA ATGGTGGCGACACAGATTATTCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCCCTGTATCTACAAATGAATAGTCTGAGAGCCGAGG ACACGGCCTTGTATTACTGTGCGAGGGATTTCTATGGTTCGGGGAGTTATTATC ACGTTCCTTTTGACTACTGGGGCCAGGGAATCCTGGTCACCGTCTCCTCA 410 STIM003-full heavy Amino acid sequence of EVQLVESGGGWRPGGSLRLSCVASGVTFDDYGMSWVRQAPGKGLEWVSGINWN chain sequence STIM003 heavy chain GGDTDYSDSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCARDFYGSGSYYHV PFDYWGQGILVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSW NSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKK VEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDP EVKFNWYVDGVEVHNAKTKPREEQYNSTYRWSVLTVLHQDWLNGKEYKCKVSN KALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK 411 STIM003-full heavy Nucleic acid sequence of GAGGTGCAGCTGGTGGAGTCTGGGGGAGGTGTGGTACGGCCTGGGGGGTCCCT chain sequence STIM003 heavy chain GAGACTCTCCTGTGTAGCCTCTGGAGTCACCTTTGATGATTATGGCATGAGCTG GGTCCGCCAAGCTCCAGGGAAGGGGCTGGARTGGGTCTCTGGTATTAATTGGA ATGGTGGCGACACAGATTATTCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCCCTGTATCTACAAATGAATAGTCTGAGAGCCGAGG ACACGGCCTTGTATTACTGTGCGAGGGATTTCTATGGTTCGGGGAGTTATTATC ACGTTCCTTTTGACTACTGGGGCCAGGGAATCCTGGTCACCGTCTCCTCAGCCA GCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTG GCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGA CCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTG TGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCA GCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACAC CAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCC CCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCC AAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGT GGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACG GCGTGGAAGTGCACMCGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCA CCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCA AAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGA CCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCC CTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAG GCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGA ACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTA CAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTG CTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCT GAGCCCCGGCAAGTGATGA 412 STIM003-CDRL1 Amino acid sequence of CDRL1 QSVSRSY of STIM003 using IMGT 413 STIM003-CDRL2 Amino acid sequence of CDRL2 GAS of STIM003 using IMGT 414 STIM003-CDRL3 Amino acid sequence of CDRL3 HQYDMSPFT of STIM003 using IMGT 415 STIM003-Light chain Amino acid sequence of V.sub.L of EIVLIQSPGTLSLSPGERATLSCRASQSVSRSYLAWYQQKRGQAPRLLIYGASSRAT variable region STIM003 GIPDRFSGDGSGTDFTLSISRLEPEDFAVYYCHQYDMSPFTFGPGTKVDIK 416 STIM003-Light chain Nucleic acid sequence of V.sub.L of GAAATTGTGTTGACGCAGTCTCCAGGGACCCTGTCTTTGTCTCCAGGGGAAAGA variable region STIM003 GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGAAGCTACTTAGCCTGG TACCAGCAGAAACGTGGCCAGGCTCCCAGGCTCCTCATCTATGGTGCATCCAGC AGGGCCACTGGCATCCCAGACAGGTTCAGTGGCGATGGGTCTGGGACAGACTTC ACTCTCTCCATCAGCAGACTGGAGCCTGAAGATTTTGCAGTGTATTACTGTCACC AGTATGATATGTCACCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAA 417 STIM003-full light Amino acid sequence of EIVLIQSPGTLSLSPGERATLSCRASQSVSRSYLAWYQQKRGQAPRLLTYGASSRAT chain sequence STIM003 light chain GIPDRFSGDGSGTDFTLSISRLEPEDFAVYYCHQYDMSPFTFGPGTINDIKRTVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAINQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 418 STIM003-full light Nucleic acid sequence of GAAATTGTGTTGACGCAGTCTCCAGGGACCCTGTCTTTGTCTCCAGGGGAAAGA chain sequence STIM003 light chain GCCACCUCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGAAGCTACTTAGCCTGG TACCAGCAGAAACGTGGCCAGGCTCCCAGGCTCCTCATCTATGGTGCATCCAGC AGGGCCACTGGCATCCCAGACAGGTTCAGTGGCGATGGGTCTGGGACAGACTTC ACTCTCTCCATCAGCAGACTGGAGCCTGAAGATTTGCAGTGTATTACTGTCACC AGTATGATATGTCACCATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAAcg tacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccggcaccgcttctg- tc gtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtggacaacgccctgcag tccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcacctactccctgtcctccaccc tgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacccaccagggcctgt ctagccccgtgaccaagtctttcaaccggggcgagtgt 419 STIM004-CDRH1 Amino acid sequence of CDRH1 GLTFDDYG of STIM004 using IMGT 420 STIM004-CDRH2 Amino acid sequence of CDRH2 INWNGDNT of STIM004 using IMGT 421 STIM004-CDRH3 Amino acid sequence of CDRH3 ARDYYGSGSYYNVPFDY of STIM004 using IMGT 422 STIM004-Heavy Amino acid sequence of VH of EVQLVESGGGWRPGGSLRLSCAASGLTFDDYGMSWVRQVPGKGLEWVSGINWN chain variable region STIM004 GDNTDYADSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCARDYYGSGSYYNV PFDYWGQGTLVTVSS 423 STIM004-Heavy Nucleic acid sequence of VH of GAGGTGCAGCTGGTGGAGTCTGGGGGAGGTGTGGTACGGCCTGGGGGGTCCCT chain variable region STIM004 GAGACTCTCCTGTGCAGCCTCTGGACTCACCTTTGATGATTATGGCATGAGCTG GGTCCGCCAAGTTCCAGGGAAGGGGCTGGAGTGGGTCTCTGGTATTAATTGGA ATGGTGATAACACAGATTATGCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCCCTGTATCTGCAAATGAACAGTCTGAGAGCCGAGG ACACGGCCTTGTATTACTGTGCGAGGGATTACTATGGTTCGGGGAGTTATTATA ACGTTCCTTTTGACTACTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCA 424 STIM004-full heavy Amino acid sequence of EVQLVESGGGVVRPGGSLRLSCAASGLTFDDYGMSWVRQVPGKGLEWVSGINWN chain sequence STIM004 heavy chain GDNTDYADSVKGRFTISRDNAKNSLYLQMNSLRAEDTALYYCARDYYGSGSYYNV PFDYWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSW NSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTINDKK VEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDP EVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSN KALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLNDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK 425 STIM004-full heavy Nucleic acid sequence of
GAGGTGCAGCTGGTGGAGTCTGGGGGAGGTGTGGTACGGCCTGGGGGGTCCCT chain sequence STIM004 heavy chain GAGACTCTCCTGTGCAGCCTCTGGACTCACCTTTGATGATTATGGCATGAGCTG GGTCCGCCAAGTTCCAGGGAAGGGGCTGGAGTGGGTCTCTGGTATTAATTGGA ATGGTGATAACACAGATTATGCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCCCTGTATCTGCAAATGAACAGTCTGAGAGCCGAGG ACACGGCCTTGTATTACTGTGCGAGGGATTACTATGGTTCGGGGAGTTATTATA ACGTTCCTTTTGACTACTGGGGCCAGGGAACCCTGGTCACCGTCTCCTCAGCCA GCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTG GCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGA CCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTG TGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCA GCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACAC CAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCC CCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCC AAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGT GGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACG GCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCA CCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCA AAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGA CCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCC CTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAG GCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGA ACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTA CAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTG CTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCT GAGCCCCGGCAAGTGATGA 426 STIM004-CDRL1 Amino acid sequence of CDRL1 QSVSSSY of STIM004 using IMGT 427 STIM004-CDRL2 Amino acid sequence of CDRL2 GAS of STIM004 using IMGT 428 STIM004-CDRL3 Amino acid sequence of CDRL3 QQYGSSPF of STIM004 using IMGT 429 STIM004-Corrected Amino acid sequence of EIVLTQSPGTLSLSPGERATLSCRASQSVSSSYLAWYQQKPGQAPRLLIYGASSRAT light chain variable corrected V.sub.L of STIM004 GIPDRFSGSGSGTDFTLTIRRLEPEDFAVYYCQQYGSSPFFGPGTKVDIK region 430 STIM004-Corrected Nucleic acid sequence of GAAATTGTGTTGACGCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAGA light chain variable corrected V.sub.L of STIM004 GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAGCTACTTAGCCTGG region TACCAGCAGAAACCTGGCCAGGCTCCCAGGCTCCTCATATATGGTGCATCCAGC AGGGCCACTGGCATCCCAGACAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTC ACTCTCACCATCAGAAGACTGGAGCCTGAAGATTTTGCAGTGTATTACTGTCAG CAGTATGGTAGTTCACCATTCTTCGGCCCTGGGACCAAAGTGGATATCAAA 431 STIM004-Light chain Nucleic acid sequence of V.sub.L of GAAATTGTGTTGACGCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAGA variable region STIM004 GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAGCTACTTAGCCTGG TACCAGCAGAAACCTGGCCAGGCTCCCAGGCTCCTCATATATGGTGCATCCAGC AGGGCCACTGGCATCCCAGACAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTC ACTCTCACCATCAGAAGACTGGAGCCTGAAGATTTTGCAGTGTATTACTGTCAG CAGTATGGTAGTTCACCATTCACTTCGGCCCTGGGACCAAAGTGGATATCAAA 432 STIM004-full Amino acid sequence of EIVLTQSPGTLSLSPGERATLSCRASQSVSSSYLAWYQQKPGQAPRLLIYGASSRAT corrected light chain STIM004 light chain GIPDRFSGSGSGTDFTLTIRRLEPEDFAVYYCQQYGSSPFFGPGTKVDIKRTVAAPS sequence VFIFPPSDEQLKSGTASVVCLLNNFYPREAINQWINDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 433 STIM004-full Nucleic acid sequence of GAAATTGTGTTGACGCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAGA corrected light chain corrected STIM004 light chain GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAGCTACTTAGCCTGG sequence TACCAGCAGAAACCTGGCCAGGCTCCCAGGCTCCTCATATATGGTGCATCCAGC AGGGCCACTGGCATCCCAGACAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTC ACTCTCACCATCAGAAGACTGGAGCCTGAAGATTTTGCAGTGTATTACTGTCAG CAGTATGGTAGTTCACCATTCTTCGGCCCTGGGACCAAAGTGGATATCAAAcgta cggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccggcaccgcttctgtc- g tgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtggacaacgccctgcagt ccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcacctactccctgtcctccaccct gaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacccaccagggcctgt ctagccccgtgaccaagtctttcaaccggggcgagtgt 434 STIM004-full light Nucleic acid sequence of GAAATTGTGTTGACGCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAGA chain sequence STIM004 light chain GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAGCTACTTAGCCTGG TACCAGCAGAAACCTGGCCAGGCTCCCAGGCTCCTCATATATGGTGCATCCAGC AGGGCCACTGGCATCCCAGACAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTC ACTCTCACCATCAGAAGACTGGAGCCTGAAGATTTTGCAGTGTATTACTGTCAG CAGTATGGTAGTTCACCATTCACTTCGGCCCTGGGACCAAAGTGGATATCAAAcg tacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccggcaccgcttctg- tc gtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtggacaacgccctgcag tccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcacctactccctgtcctccaccc tgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacccaccagggcctgt ctagccccgtgaccaagtctttcaaccggggcgagtgt 435 STIM005-CDRH1 Amino acid sequence of CDRH1 GYTFNSYG of STIM005 using IMGT 436 STIM005-CDRH2 Amino acid sequence of CDRH2 ISVHNGNT of STIM005 using IMGT 437 STIM005-CDRH3 Amino acid sequence of CDRH3 ARAGYDILTDFSDAFDI of STIM005 using IMGT 438 STIM005-Heavy Amino acid sequence of V.sub.H of QVQLVQSGAEVKKPGASVKVSCKASGYTFNSYGIIWVRQAPGQGLEWMGVVISVH chain variable region STIM005 NGNTNCAQKLQGRVIMTTDTSTSTAYMELRSLRTDDTAVYYCARAGYDILTDFSD AFDIWGHGTMVTVSS 439 STIM005-Heavy Nucleic acid sequence of V.sub.H of CAGGTTCAGTTGGTGCAGTCTGGAGCTGAGGTGAAGAAGCCTGGGGCCTCAGT chain variable region STIM005 GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTAATAGTTATGGTATCATCTG GGTGCGACAGGCCCCTGGACAAGGGCTTGAGTGGATGGGATGGATCAGCGTTC ACAATGGTAACACAAACTGTGCACAGAAGCTCCAGGGTAGAGTCACCATGACCA CAGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCCTGAGAACTGACG ACACGGCCGTGTATTACTGTGCGAGAGCGGGTTACGATATTTTGACTGATTTTT CCGATGCTTTTGATATCTGGGGCCACGGGACAATGGTCACCGTCTCTTCA 440 STIM005-full heavy Amino acid sequence of QVQLVQSGAEVKKPGASVKVSCKASGYTFNSYGIIWVRQAPGQGLEWMGWISVH chain sequence STIM005 heavy chain NGNTNCAQKLQGRVTMTTDTSTSTAYMELRSLRTDDTAVYYCARAGYDILTDFSD AFDIWGHGTMVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSW NSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKK VEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDILMISRTPEVTCVVVDVSHEDP EVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLIVLHQDWLNGKEYKOWSN KALPAPIEKTISMKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK 441 STIM005-full heavy Nucleic acid sequence of CAGGTTCAGTTGGTGCAGTCTGGAGCTGAGGTGAAGAAGCCTGGGGCCTCAGT chain sequence STIM005 heavy chain GAAGGTCTCCTGCAAGGCTTCTGGTTACACCTTTAATAGTTATGGTATCATCTG GGTGCGACAGGCCCCTGGACAAGGGCTTGAGTGGATGGGATGGATCAGCGTTC ACAATGGTAACACAAACTGTGCACAGAAGCTCCAGGGTAGAGTCACCATGACCA CAGACACATCCACGAGCACAGCCTACATGGAGCTGAGGAGCCTGAGAACTGACG ACACGGCCGTGTATTACTGTGCGAGAGCGGGTTACGATATTTTGACTGATTTTT CCGATGCTTTTGATATCTGGGGCCACGGGACAATGGTCACCGTCTCTTCA GCCAGCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACC TCTGGCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCT GTGACCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCT GCTGTGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCT TCCAGCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCA ACACCAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCT GTCCCCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCC CCCCAAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCG TGGTGGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGG ACGGCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACT CCACCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACG GCAAAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAA AGACCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGC CCCCTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGA AAGGCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTG AGAACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCT GTACAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTC CTGCTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTC CCTGAGCCCCGGCAAGTGATGA 442 STIM005-CDRL1 Amino acid sequence of CDRL1 QNINNF of STIM005 using IMGT 443 STIM005-CDRL2 Amino acid sequence of CDRL2 AAS of STIM005 using IMGT 444 STIM005-CDRL3 Amino acid sequence of CDRL3 QQSYGIPW of STIM005 using IMGT 445 STIM005-Light chain Amino acid sequence of VL of DIQMTQSPSSLSASVGDRVTITCRASQNINNFLNWYQQKEGKGPKLLIYAASSLQR variable region STIM005 GIPSTFSGSGSGTDFTLTISSLQPEDFATYICQQSYGIPWVGQGTKVEIK 446 STIM005-Light chain Nucleic acid sequence of VL of GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGA variable region STIM005 GTCACCATCACTTGCCGGGCAAGTCAGAACATTAATAACTTTTTAAATTGGTATC AGCAGAAAGAAGGGAAAGGCCCTAAGCTCCTGATCTATGCAGCATCCAGTTTGC AAAGAGGGATACCATCAACGTTCAGTGGCAGTGGATCTGGGACAGACTTCACTC TCACCATCAGCAGTCTGCAACCTGAAGATTTTGCAACTTACATCTGTCAACAGAG CTACGGTATCCCGTGGGTCGGCCAAGGGACCAAGGTGGAAATCAAA 447 STIM005-full light Amino acid sequence of DIQMTQSPSSLSASVGDRVTITCRASQNINNFLNWYQQKEGKGPKLLIYAASSLQR chain sequence STIM005 light chain GIPSTFSGSGSGTDFTLTISSLQPEDFATYICQQSYGIPWVGQGTKVEIK RTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESV TEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 448 STIM005-full light Nucleic acid sequence of GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGA chain sequence STIM005 light chain GTCACCATCACTTGCCGGGCAAGTCAGAACATTAATAACTTTTTAAATTGGTATC AGCAGAAAGAAGGGAAAGGCCCTAAGCTCCTGATCTATGCAGCATCCAGTTTGC AAAGAGGGATACCATCAACGTTCAGTGGCAGTGGATCTGGGACAGACTTCACTC TCACCATCAGCAGTCTGCAACCTGAAGATTTTGCAACTTACATCTGTCAACAGAG CTACGGTATCCCGTGGGTCGGCCAAGGGACCAAGGTGGAAATCAAAcgtacggtgg ccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccggcaccgcttctgtcgtgtgc- ct gctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtggacaacgccctgcagtccggca actcccaggaatccgtgaccgagcaggactccaaggacagcacctactccctgtcctccaccctgaccct gtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacccaccagggcctgtctagcc ccgtgaccaagtctttcaaccggggcgagtgt 449 STIM006-CDRH1 Amino acid sequence of CDRH1 GFTFSDYF of STIM006 using IMGT 450 STIM006-CDRH2 Amino acid sequence of CDRH2 ISSSGSTI of STIM006 using IMGT 451 STIM006-CDRH3 Amino acid sequence of CDRH3 ARDHYDGSGIYPLYYYYGLDV of STIM006 using IMGT 452 STIM006-Heavy Amino acid sequence of VH of QVQLVESGGGLVKPGGSLRLSCAASGFTFSDYFMSWIRQAPGKGLEWISYISSSGS chain variable region STIM006 TIYYADSVRGRFTISRDNAKYSLYLQMNSLRSEDTAVYYCARDHYDGSGIYPLYYYY GLDVWGQGTTVTVSS 453 STIM006-Heavy Nucleic acid sequence of VH of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGGTCCCT chain variable region STIM006 GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTCAGTGACTACTTCATGAGCTG GATCCGCCAGGCGCCAGGGAAGGGGCTGGAGTGGATTTCATACATTAGTTCTAG TGGTAGTACCATATACTACGCAGACTCTGTGAGGGGCCGATTCACCATCTCCAG GGACAACGCCAAGTACTCACTGTATCTGCAAATGAACAGCCTGAGATCCGAGGA CACGGCCGTGTATTACTGTGCGAGAGATCACTACGATGGTTCGGGGATTTATCC CCTCTACTACTATTACGGTTTGGACGTCTGGGGCCAGGGGACCACGGICACCGT CTCCTCA 454 STIM006-full heavy Amino acid sequence of QVQLVESGGGLVKPGGSLRLSCAASGFTFSDYFMSWIRQAPGKGLEWISYISSSGS chain sequence STIM006 heavy chain TIYYADSVRGRFTISRDNAKYSLYLQMNSLRSEDTAVYYCARDHYDGSGIYPLYYYY GLDVWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSW NSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTINDKK VEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDP EVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCINSN KALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK 455 STIM006-full heavy Nucleic acid sequence of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGGTCCCT chain sequence STIM006 heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTCAGTGACTACTTCATGAGCTG
GATCCGCCAGGCGCCAGGGAAGGGGCTGGAGTGGATTTCATACATTAGTTCTAG TGGTAGTACCATATACTACGCAGACTCTGTGAGGGGCCGATTCACCATCTCCAG GGACAACGCCAAGTACTCACTGTATCTGCAAATGAACAGCCTGAGATCCGAGGA CACGGCCGTGTATTACTGTGCGAGAGATCACTACGATGGTTCGGGGATTTATCC CCTCTACTACTATTACGGTTTGGACGTCTGGGGCCAGGGGACCACGGTCACCGT CTCCTCAGCCAGCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAA GTCCACCTCTGGCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCC CGAGCCTGTGACCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACAC CTTCCCTGCTGTGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGAC CGTGCCTTCCAGCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAA GCCCTCCAACACCAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGAC CCACACCTGTCCCCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTT CCTGTTCCCCCCAAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGT GACCTGCGTGGTGGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTG GTACGTGGACGGCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAAC AGTACAACTCCACCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATT GGCTGAACGGCAAAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCC CCATCGAAAAGACCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGT ACACACTGCCCCCTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCT GTCTCGTGAAAGGCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACG GCCAGCCTGAGAACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCT CATTCTTCCTGTACAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCA ACGTGTTCTCCTGCTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGA AGTCCCTGTCCCTGAGCCCCGGCAAGTGATGA 456 STIM006-CDRL1 Amino acid sequence of CDRL1 QSLLHSNGYNY of STIM006 using IMGT 457 STIM006-CDRL2 Amino acid sequence of CDRL2 LGS of STIM006 using IMGT 458 STIM006-CDRL3 Amino acid sequence of CDRL3 MQALQTPRS of STIM006 using IMGT 459 STIM006-Light chain Amino acid sequence of VL of IVMTQSPLSLPVTPGEPASISCRSSQSLLHSNGYNYLDYYLQKPGQSPQLLIYLGSY variable region STIM006 RASGVPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPRSFGQGTTLEIK 460 STIM006-Light chain Nucleic acid sequence of VL of ATTGTGATGACTCAGTCTCCACTCTCCCTACCCGTCACCCCTGGAGAGCCGGCC variable region STIM006 TCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTAATGGATACAACTATT TGGATTATTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTATTTGG GTTCTTATCGGGCCTCCGGGGTCCCTGACAGGTTCAGTGGCAGTGGATCAGGCA CAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTTTATT ACTGCATGCAAGCTCTACAAACTCCTCGCAGTTTTGGCCAGGGGACCACGCTGG AGATCAAA 461 STIM006-full light Amino acid sequence of IVMTQSPLSLPVTPGEPASISCRSSQSLLHSNGYNYLDYYLQKPGQSPQLLTYLGSY chain sequence STIM006 light chain RASGVPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPRSFGQGTTLEIKRT VAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAWQWINDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 462 STIM006-full light Nucleic acid sequence of ATTGTGATGACTCAGTCTCCACTCTCCCTACCCGTCACCCCTGGAGAGCCGGCC chain sequence STIM006 light chain TCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTAATGGATACAACTATT TGGATTATTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTATTTGG GTTCTTATCGGGCCTCCGGGGTCCCTGACAGGTTCAGTGGCAGTGGATCAGGCA CAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTTTATT ACTGCATGCAAGCTCTACAAACTCCTCGCAGTTTTGGCCAGGGGACCACGCTGG AGATCAAAcgtacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccg gcaccgcttctgtcgtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtgga caacgccctgcagtccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcacctactc cctgtcctccaccctgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacc caccagggcctgtctagccccgtgaccaagtctttcaaccggggcgagtgt 463 STIM007-CDRH1 Amino acid sequence of CDRH1 GFSLSTTGVG of STIM007 using IMGT 464 STIM007-CDRH2 Amino acid sequence of CDRH2 IYWDDDK of STIM007 using IMGT 465 STIM007-CDRH3 Amino acid sequence of CDRH3 THGYGSASYYHYGMDV of STIM007 using IMGT 466 STIM007-Heavy Amino acid sequence of V.sub.H of QITLKESGPTLVKPTQTLTLTCTFSGFSLSTTGVGVGWIRQPPGKALEWLAVIYWD chain variable region STIM007 DDKRYSPSLKSRLTITKDTSKNQVVLTMTNMDPVDTATYFCTHGYGSASYYHYGM DVWGQGTTVTVSS 467 STIM007-Heavy Nucleic acid sequence of V.sub.H of CAGATCACCTTGAAGGAGTCTGGTCCTACGCTGGTGAAACCCACACAGACCCTC chain variable region STIM007 ACGCTGACCTGCACCTTCTCTGGGTTCTCACTCAGCACTACTGGAGTGGGTGTG GGCTGGATCCGTCAGCCCCCAGGAAAGGCCCTGGAGTGGCTTGCAGTCATTTAT TGGGATGATGATAAGCGCTACAGCCCATCTCTGAAGAGCAGACTCACCATCACC AAGGACACCTCCAAAAACCAGGTGGTCCTTACAATGACCAACATGGACCCTGTG GACACAGCCACATATTTCTGTACACACGGATATGGTTCGGCGAGTTATTACCACT ACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCA 468 STIM007-full heavy Amino acid sequence of QITLKESGPTLVKPTQTLTLTCTFSGFSLSTTGVGVGWIRQPPGKALEWLAVIYWD chain sequence STIM007 heavy chain DDKRYSPSLKSRLTITKDTSKNQVVLTMTNMDPVDTATYFCTHGYGSASYYHYGM DVWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWN SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVE PKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVWDVSHEDPEV KFNWYVDGVEVHNAKTKPREEQYNSTYRWSVLTVLHQDWLNGKEYKCVSNKA LPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQ KSLSLSPGK 469 STIM007-full heavy Nucleic acid sequence of CAGATCACCTTGAAGGAGTCTGGTCCTACGCTGGTGAAACCCACACAGACCCTC chain sequence STIM007 heavy chain ACGCTGACCTGCACCTTCTCTGGGTTCTCACTCAGCACTACTGGAGTGGGTGTG GGCTGGATCCGTCAGCCCCCAGGAAAGGCCCTGGAGTGGCTTGCAGTCATTTAT TGGGATGATGATAAGCGCTACAGCCCATCTCTGAAGAGCAGACTCACCATCACC AAGGACACCTCCAAAAACCAGGTGGTCCTTACAATGACCAACATGGACCCTGTG GACACAGCCACATATTTCTGTACACACGGATATGGTTCGGCGAGTTATTACCACT ACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCA GCCAGCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACC TCTGGCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCT GTGACCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCT GCTGTGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCT TCCAGCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCA ACACCAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCT GTCCCCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCC CCCCAAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCG TGGTGGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGG ACGGCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACT CCACCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACG GCAAAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAA AGACCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGC CCCCTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGA AAGGCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTG AGAACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCT GTACAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTC CTGCTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTC CCTGAGCCCCGGCAAGTGATGA 470 STIM007-CDRL1 Amino acid sequence of CDRL1 QSVTNY of STIM007 using IMGT 471 STIM007-CDRL2 Amino acid sequence of CDRL2 DAS of STIM007 using IMGT 472 STIM007-CDRL3 Amino acid sequence of CDRL3 QHRSNWPLT of STIM007 using IMGT 473 STIM007-Light chain Amino acid sequence of V.sub.L of EIVLTQSPATLSLSPGERATLSCRASQSVTNYLAWHQQKPGQAPRLLIYDASNRAT variable region STIM007 GIPARFSGSGSGTDFTLTISSLEPEDFAVYYCQHRSNWPLTFGGGTINEIK 474 STIM007-Light chain Nucleic acid sequence of V.sub.L of GAAATTGTATTGACACAGTCTCCAGCCACCCTGTCTTTGTCTCCAGGGGAAAGA variable region STIM007 GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTACCAACTACTTAGCCTGGCAC CAACAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGATGCATCCAACAGG GCCACTGGCATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACT CTCACCATCAGCAGCCTAGAGCCTGAAGATTTTGCAGTTTATTACTGTCAGCACC GTAGCAACTGGCCTCTCACTTTCGGCGGAGGGACCAAGGTGGAGATCAAAC 475 STIM007-full light Amino acid sequence of EIVLTQSPATLSLSPGERATLSCRASQSVTNYLAWHQQKPGQAPRLLIYDASNRAT chain sequence STIM007 light chain GIPARFSGSGSGTDFTLTISSLEPEDFAVYYCQHRSNWPLTFGGGTKVEIKRIVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 476 STIM007-full light Nucleic acid sequence of GAAATTGTATTGACACAGTCTCCAGCCACCCTGTCTTTGTCTCCAGGGGAAAGA chain sequence STIM007 light chain GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTACCAACTACTTAGCCTGGCAC CAACAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGATGCATCCAACAGG GCCACTGGCATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACT CTCACCATCAGCAGCCTAGAGCCTGAAGATTTTGCAGTTTATTACTGTCAGCACC GTAGCAACTGGCCTCTCACTTTCGGCGGAGGGACCAAGGTGGAGATCAAACcgta cggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccggcaccgcttctgtc- g tgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtggacaacgccctgcagt ccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcacctactccctgtcctccaccct gaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacccaccagggcctgt ctagccccgtgaccaagtctttcaaccggggcgagtgt 477 STIM008-CDRH1 Amino acid sequence of CDRH1 GFSLSTSGVG of STIM008 using IMGT 478 STIM008-CDRH2 Amino acid sequence of CDRH2 IYWDDDK of STIM008 using IMGT 479 STIM008-CDRH3 Amino acid sequence of CDRH3 THGYGSASYYHYGMDV of STIM008 using IMGT 480 STIM008-Heavy Amino acid sequence of V.sub.H of QITLKESGPTLVKPTQTLTLTCTFSGFSLSTSGVGVGWIRQPPGKALEWLAVIYWD chain variable region STIM008 DDKRYSPSLKSRLITTKDTSKNQVVLTMTNMDPVDTATYFCTHGYGSASYYHYGM DVWGQGTTVTVSS 481 STIM008-Heavy Nucleic acid sequence of V.sub.H of CAGATCACCTTGAAGGAGTCTGGTCCTACGCTGGTGAAACCCACACAGACCCTC chain variable region STIM008 ACGCTGACCTGCACCTTCTCTGGGTTCTCACTCAGCACTAGTGGAGTGGGTGTG GGCTGGATCCGTCAGCCCCCAGGAAAGGCCCTGGAGTGGCTTGCAGTCATTTAT TGGGATGATGATAAGCGCTACAGCCCATCTCTGAAGAGCAGGCTCACCATCACC AAGGACACCTCCAAAAACCAGGTGGTCCTTACAATGACCAACATGGACCCTGTG GACACAGCCACATATTTCTGTACACACGGATATGGTTCGGCGAGTTATTACCACT ACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCA 482 STIM008-full heavy Amino acid sequence of QITLKESGPTLVKPTQTLTLTCTFSGFSLSTSGVGVGWIRQPPGKALEWLAVIYWD chain sequence STIM008 heavy chain DDKRYSPSLKSRLTITKDTSKNQWLTMTNMDPVDTATYFCTHGYGSASYYHYGM DVWGQGTTVWSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWN SGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKINE PKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEV KFNWYVDGVEVHNAKTKPREEQYNSTYRWSVLTVLHQDWLNGKEYKCKVSNKA LPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQ KSLSLSPGK 483 STIM008-full heavy Nucleic acid sequence of CAGATCACCTTGAAGGAGTCTGGTCCTACGCTGGTGAAACCCACACAGACCCTC chain sequence STIM008 heavy chain ACGCTGACCTGCACCTTCTCTGGGTTCTCACTCAGCACTAGTGGAGTGGGTGTG GGCTGGATCCGTCAGCCCCCAGGAAAGGCCCTGGAGTGGCTTGCAGTCATTTAT TGGGATGATGATAAGCGCTACAGCCCATCTCTGAAGAGCAGGCTCACCATCACC AAGGACACCTCCAAAAACCAGGTGGTCCTTACAATGACCAACATGGACCCTGTG GACACAGCCACATATTTCTGTACACACGGATATGGTTCGGCGAGTTATTACCACT ACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCCAGCA CCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTGGCG GAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGACCG TGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTGTGC TGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCAGCT CTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACACCAA GGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCCCCC TTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCCAAA GCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGTGGT GGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACGGCGT GGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCACCTA CCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCAAAG AGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGACCA TCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCCCTA GCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAGGCT TCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGAACA ACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTACAG CAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTGCTC CGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCTGAG CCCCGGCAAGTGATGA 484 STIM008-CDRL1 Amino acid sequence of CDRL1 QSVTNY of STIM008 using IMGT 485 STIM008-CDRL2 Amino acid sequence of CDRL2 DAS
of STIM008 using IMGT 486 STIM008-CDRL3 Amino acid sequence of CDRL3 QQRSNWPLT of STIM008 using IMGT 487 STIM008-Light chain Amino acid sequence of V.sub.L of EIVLTQSPATLSLSPGERATLSCRASQSVTNYLAWHQQKPGQAPRLLIYDASNRAT variable region STIM008 GIPARFSGSGSGTDFTLTISSLEPEDFAVYYCQQRSNWPLIFGGGTWEIK 488 STIM008-Light chain Nucleic acid sequence of V.sub.L of GAAATTGTGTTGACACAGTCTCCAGCCACCCTGTCTTTGTCTCCAGGGGAAAGA variable region STIM008 GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTACCAACTACTTAGCCTGGCAC CAACAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGATGCATCCAACAGG GCCACTGGCATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACT CTCACCATCAGCAGCCTAGAGCCTGAAGATTTTGCAGTTTATTACTGTCAGCAGC GTAGCAACTGGCCTCTCACTTTCGGCGGAGGGACCAAGGTGGAGATCAAA 489 STIM008-full light Amino acid sequence of EIVLTQSPATLSLSPGERATLSCRASQSVTNYLAWHQQKPGQAPRLLIYDASNRAT chain sequence STIM008 light chain GIPARFSGSGSGTDFTLTISSLEPEDFAVYYCQQRSNWPLIFGGGTKVEIKRIVAAP SVFIFPPSDEQLKSGTASVVCLLNNFYPREAINQWKVDNALQSGNSQESVTEQDSK DSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 490 STIM008-full light Nucleic acid sequence of GAAATTGTGTTGACACAGTCTCCAGCCACCCTGTCTTTGTCTCCAGGGGAAAGA chain sequence STIM008 light chain GCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTACCAACTACTTAGCCTGGCAC CAACAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGATGCATCCAACAGG GCCACTGGCATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACT CTCACCATCAGCAGCCTAGAGCCTGAAGATTTTGCAGTTTATTACTGTCAGCAGC GTAGCAACTGGCCTCTCACTTTCGGCGGAGGGACCAAGGTGGAGATCAAAcgtac ggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaagtccggcaccgcttctgtcg- t gtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaaggtggacaacgccctgcagtc cggcaactcccaggaatccgtgaccgagcaggactccaaggacagcacctactccctgtcctccaccctg accctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaagtgacccaccagggcctgtct agccccgtgaccaagtctttcaaccggggcgagtgt 491 STIM009-CDRH1 Amino acid sequence of CDRH1 GFTFSDYY of STIM009 using IMGT 492 STIM009-CDRH2 Amino acid sequence of CDRH2 ISSSGSTI of STIM009 using IMGT 493 STIM009-CDRH3 Amino acid sequence of CDRH3 ARDFYDILTDSPYFYYGVDV of STIM009 using IMGT 494 STIM009-Heavy Amino acid sequence of V.sub.H of QVQLVESGGGLVKPGGSLRLSCAASGFTFSDYYMSWIRQAPGKGLEWVSYISSSGS chain variable region STIM009 TIYYADSVKGRFTISRDNAKNSLYLQINSLRAEDTAVYYCARDFYDILTDSPYFYYG VDVWGQGTTVTVSS 495 STIM009-Heavy Nucleic acid sequence of V.sub.H of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGGTCCCT chain variable region STIM009 GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTCAGTGACTACTACATGAGCTG GATCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTTTCATACATTAGTAGTA GTGGTAGTACCATATACTACGCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GGGACAACGCCAAGAACTCACTGTATCTGCAAATTAACAGCCTGAGAGCCGAGG ACACGGCCGTGTATTACTGTGCGAGAGATTTTTACGATATTTTGACTGATAGTC CGTACTTCTACTACGGTGTGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCT CCTCA 496 STIM009-full heavy Amino acid sequence of QVQLVESGGGLVKPGGSLRLSCAASGFTFSDYYMSWIRQAPGKGLEWVSYISSSGS chain sequence STIM009 heavy chain TIYYADSVKGRFTISRDNAKNSLYLQINSLRAEDTAVYYCARDFYDILTDSPYFYYG VDVWGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSW NSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKK VEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDP EVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSN KALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK 497 STIM009-full heavy Nucleic acid sequence of CAGGTGCAGCTGGTGGAGTCTGGGGGAGGCTTGGTCAAGCCTGGAGGGTCCCT chain sequence STIM009 heavy chain GAGACTCTCCTGTGCAGCCTCTGGATTCACCTTCAGTGACTACTACATGAGCTG GATCCGCCAGGCTCCAGGGAAGGGGCTGGAGTGGGTTTCATACATTAGTAGTA GTGGTAGTACCATATACTACGCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GGGACAACGCCAAGAACTCACTGTATCTGCAAATTAACAGCCTGAGAGCCGAGG ACACGGCCGTGTATTACTGTGCGAGAGATTTTTACGATATTTTGACTGATAGTC CGTACTTCTACTACGGTGTGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCT CCTCAGCCAGCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGT CCACCTCTGGCGGAACAGCCTGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCG AGCCTGTGACCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCT TCCCTGCTGTGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCG TGCCTTCCAGCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCC CTCCAACACCAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCA CACCTGTCCCCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCT GTTCCCCCCAAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGAC CTGCGTGGTGGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTA CGTGGACGGCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGT ACAACTCCACCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGC TGAACGGCAAAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCA TCGAAAAGACCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACA CACTGCCCCCTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTC TCGTGAAAGGCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCC AGCCTGAGAACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCAT TCTTCCTGTACAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACG TGTTCTCCTGCTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGT CCCTGTCCCTGAGCCCCGGCAAGTGATGA 498 STIM009-CDRL1 Amino acid sequence of CDRL1 QSLLHSNGYNY of STIM009 using IMGT 499 STIM009-CDRL2 Amino acid sequence of CDRL2 LGS of STIM009 using IMGT 500 STIM009-CDRL3 Amino acid sequence of CDRL3 MQALQTPRT of STIM009 using IMGT 501 STIM009-Light chain Amino acid sequence of V.sub.L of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSNGYNYLDWYLQKPGQSPQLLIYLGS variable region STIM009 NRASGVPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPRTFGQGTWEIK 502 STIM009-Light chain Nucleic acid sequence of V.sub.L of GATATTGTGATGACTCAGTTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG variable region STIM009 GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTAATGGATACAACT ATTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTATT TGGGTTCTAATCGGGCCTCCGGGGTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCTCGGACGTTCGGCCAAGGGACCAAG GTGGAAATCAAA 503 STIM009-full light Amino acid sequence of DIVMTQSPLSLPVTPGEPASISCRSSQSLLHSNGYNYLDWYLQKPGQSPQLLIYLGS chain sequence STIM009 light chain NRASGVPDRFSGSGSGTDFTLKISRVEAEDVGVYYCMQALQTPRTFGQGTKVEIKR TVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVT EQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC 504 STIM009-full light Nucleic acid sequence of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG chain sequence STIM009 light chain GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTAATGGATACAACT ATTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTATT TGGGTTCTAATCGGGCCTCCGGGGTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCTCGGACGTTCGGCCAAGGGACCAAG GTGGAAATCAAAcgtacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaa gtccggcaccgcttctgtcgtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaag gtggacaacgccctgcagtccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcac ctactccctgtcctccaccctgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaa gtgacccaccagggcctgtctagccccgtgaccaagtctttcaaccggggcgagtgt 505 Human PD-L1 Flag His Amino acid sequence of FTVTVPKDLYVVEYGSNMTIECKFPVEKQLDLAALIVYWEMEDKNIIQFVHGEEDLK (KYPROT286) KWROT286 with FLAG taq in VQHSSYRQRARLLKDQLSLGNAALQITDVKLQDAGVYRCMISYGGADYKRITVKV bold and underlined and NAPYNKINQRILVVDPVTSEHELTCQAEGYPKAEVIWTSSDHQVLSGKTTTTNSKR histidine tag in bold EEKLFNVTSTLRINTTTNEIFYCTFRRLDPEENHTAELVIPELPLAHPPNERTIEGRD YKDDDDKHHHHHH 506 Mature human ICOS Mature amino acid sequence of EINGSANYEMFIFHNGGVQILCKYPDIVQQFKMQLLKGGQILCDLIKTKGSGNTVSI human ICOS KSLKFCHSQLSNNSVSFFLYNLDHSHANYYFCNLSIFDPPPFKVTLIGGYLHIYESQL CCQLXFWLPIGCAAFVVVCILGCILICWLIKKKYSSSVHDPNGEYMFMRAVNTAKK SRLTDVTL 507 Human ICOS Amino acid sequence of human EINGSANYEMFIFHNGGVQILCKYPDIVQQFKMQLLKGGQILCDLIKTKGSGNTVSI extracellular domain ICOS extracellular domain KSLKFCHSQLSNNSVSFFLYNLDHSHANYYFCNLSIFDPPPFKVTLTGGYLHIYESQL CCQLKF 508 Human ICOS with Amino acid sequence of human MKSGLWYFFLFCLRIKVLTGEINGSANYEMFIFHNGGVQILC1MDIVQQFKMQLLK signal peptide ICOS (signal peptide is GGQILCDLTKTKGSGNTVSIKSLKFCHSQLSNNSVSFFLYNLDHSHANYYFCNLSIF underlined) DPPPFKVTLTGGYLHIYESQLCCQLKFWLPIGCAAFVVVCILGCILICWLTKKKYSSS VHDPNGEYMFMRAVNTAKKSRLTDVTL 509 Isoform of human Amino acid sequence of a The sequence of this isoform differs from the canonical sequence in its ICOS (Q9Y6W8-2) human ICOS isoform cytoplasmic domain as follows: 168-199: KYSSSVHDPNGEYMFMRAVNTAKKSRLTDVTLM 510 Mature mouse ICOS Amino acid sequence of mature EINGSADHRMFSFHNGGVQISCKYPETVQQLKMRLFREREVLCELTKTKGSGNAVS mouse ICOS IKNPMLCLYHLSNNSVSFFLNNPDSSQGSYYFCSLSIFDPPPFQERNLSGGYLHIYES QLCCQLKIVVQVTE 511 Mouse ICOS Amino acid sequence of the EINGSADHRMFSFHNGGVQISCKYPETVQQLKMRLFREREVLCELTKTKGSGNAVS extracellular domain extracellular domain of mouse IKNPMLCLYHLSNNSVSFFLNNPDSSQGSYYFCSLSIFDPPPFQERNLSGGYLHIYES ICOS QLCCQLK 512 Mouse ICOS with Amino acid sequence of mouse MGWSCIILFLVATATGVHSEINGSADHRMFSFHNGGVQISCKYPETVQQLKMRLFR signal peptide ICOS (signal peptide is EREVLCELTKTKGSGNAVSIKNPMLCLYHLSNNSVSFFLNNPDSSQGSYYFCSLSIF underlined) DPPPFQERNLSGGYLHIYESQLCCQLKIVVQVTE 513 Cynomolgus ICOS with Amino acid sequence of MKSGLWYFFL FCLHMKVLTG EINGSANYEM FIFHNGGVQI LCKYPDIVQQ signal peptide cynomolgus ICOS (signal FKMQLLKGGQILCDLTKTKGSGNINSIKSLKFCHSQLSNNSVSFFLYNLD peptide is underlined) RSHANYYFCNLSIFDPPPFKVTLTGGYLHIYESQLCCQLKFWLPIGCATF VVVCIFGCILICWLTKKKYSSTVHDPNGEYMFMRAVNTAKKSRLTGTTP 514 Cynomolgus ICOS Amino acid sequence of EINGSANYEMFIFHNGGVQILCKYPDIVQQFKMQLLKGGQILCDLTKTKG extracellular domain cynomolgus ICOS extracellular SGNINSIKSLKFCHSQLSNNSVSFFLYNLDRSHANYYFCNLSIFDPPPFK domain VTLTGGYLHIYESQLCCQLK 515 Human ICOS ligand Amino acid sequence of human DTQEKEVRAMVGSDVELSCACPEGSRFDLNDVYVYWQTSESKTVVTYHIPQNSSL ICOS ligand comprising ENVDSRYRNRALMSPAGMLRGDFSLRLFNVIPQDEQKFHCLVLSQSLGFQEVLSVE extracellular domain VTLHVAANFSVPVVSAPHSPSQDELTFTCTSINGYPRPNVYWINKTDNSLLDQALQ NDTVFLNMRGLYDVVSVLRIARTPSVNIGCCIENVLLQQNLTVGSQTGNDIGERDK ITENPVSTGEKNAATWS 516 Human ICOS ligand MRLGSPGLLFLLFSSLRADTQEKEVRAMVGSDVELSCACPEGSRFDLNDVYVYWQT SESKTWTYHIPQNSSLENVDSRYRNRALMSPAGMLRGDFSLRLFNVTPQDEQKFH CLVLSQSLGFQEVLSVEVTLHVAANFSVPWSAPHSPSQDELTFTCTSINGYPRPNV YWINKTDNSLLDQALQNDTVFLNMRGLYDVVSVLRIARTPSVNIGCCIENVLLQQN LTVGSQTGNDIGERDKITENPVSTGEKNAATWSILAVLCLLVVVAVAIGWVCRDRC LQHSYAGAWAVSPETELTGHV 517 C-terminal amino acid Amino acids 21 to 133 of hIL-2 LQMILNGINNYKNPKLTAMLIFITYMPKKATELKHLQCLEEELKPLEEVLNLAQSKN sequence of hIL-2 with R38W mutation (bold & FHLRPRDLISNINVIVLELKGSETTFMCEYADETATIVEFLNRWITFCQSIISTLT underlined) 518 C-terminal amino acid Amino acids 21 to 133 of hIL-2 LQMILNGINNYKNPKLTQMLIFKFYMPKKATELKHLQCLEEELKPLEEVLNLAQSKN sequence of hIL-2 with R38Q mutation (bold & FHLRPRDLISNINVIVLELKGSETTFMCEYADETATIVEFLNRWITFCQSIISTLT underlined) 519 STIM002-Corrected Nucleic acid sequence of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG Light chain variable corrected VL of STIM002
GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTGATGGATACAAC region TATTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTAT TTGGGTTCTACTCGGGCCTCCGGGTTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCGCTCAGTTTTGGCCAGGGGACCAAG CTGGAGATCAAA 520 STIM002-Corrected Nucleic acid sequence of GATATTGTGATGACTCAGTCTCCACTCTCCCTGCCCGTCACCCCTGGAGAGCCG full light chain corrected STIM002 light chain GCCTCCATCTCCTGCAGGTCTAGTCAGAGCCTCCTGCATAGTGATGGATACAAC sequence TATTTGGATTGGTACCTGCAGAAGCCAGGGCAGTCTCCACAGCTCCTGATCTAT TTGGGTTCTACTCGGGCCTCCGGGTTCCCTGACAGGTTCAGTGGCAGTGGATCA GGCACAGATTTTACACTGAAAATCAGCAGAGTGGAGGCTGAGGATGTTGGGGTT TATTACTGCATGCAAGCTCTACAAACTCCGCTCAGTTTTGGCCAGGGGACCAAG CTGGAGATCAAAcgtacggtggccgctccctccgtgttcatcttcccaccttccgacgagcagctgaa gtccggcaccgcttctgtcgtgtgcctgctgaacaacttctacccccgcgaggccaaggtgcagtggaag gtggacaacgccctgcagtccggcaactcccaggaatccgtgaccgagcaggactccaaggacagcac ctactccctgtcctccaccctgaccctgtccaaggccgactacgagaagcacaaggtgtacgcctgcgaa gtgacccaccagggcctgtctagccccgtgaccaagtctttcaaccggggcgagtgt 521 STIM003-Corrected Nucleic acid sequence of GAGGTGCAGCTGGTGGAGTCTGGGGGAGGTGTGGTACGGCCTGGGGGGTCCCT heavy chain variable corrected VH of STIM003 GAGACTCTCCTGTGTAGCCTCTGGAGTCACCTTTGATGATTATGGCATGAGCTG region GGTCCGCCAAGCTCCAGGGAAGGGGCTGGAGTGGGTCTCTGGTATTAATTGGA ATGGTGGCGACACAGATTATTCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCCCTGTATCTACAAATGAATAGTCTGAGAGCCGAGG ACACGGCCTTGTATTACTGTGCGAGGGATTTCTATGGTTCGGGGAGTTATTATC ACGTTCCTTTTGACTACTGGGGCCAGGGAATCCTGGTCACCGTCTCCTCA 522 STIM003-Corrected Nucleic acid sequence of GAGGTGCAGCTGGTGGAGTCTGGGGGAGGTGTGGTACGGCCTGGGGGGTCCCT full heavy chain corrected STIM003 heavy chain GAGACTCTCCTGTGTAGCCTCTGGAGTCACCTTTGATGATTATGGCATGAGCTG sequence GGTCCGCCAAGCTCCAGGGAAGGGGCTGGAGTGGGTCTCTGGTATTAATTGGA ATGGTGGCGACACAGATTATTCAGACTCTGTGAAGGGCCGATTCACCATCTCCA GAGACAACGCCAAGAACTCCCTGTATCTACAAATGAATAGTCTGAGAGCCGAGG ACACGGCCTTGTATTACTGTGCGAGGGATTTCTATGGTTCGGGGAGTTATTATC ACGTTCCTTTTGACTACTGGGGCCAGGGAATCCTGGTCACCGTCTCCTCAGCCA GCACCAAGGGCCCCTCTGTGTTCCCTCTGGCCCCTTCCAGCAAGTCCACCTCTG GCGGAACAGCCGCTCTGGGCTGCCTCGTGAAGGACTACTTCCCCGAGCCTGTGA CCGTGTCCTGGAACTCTGGCGCTCTGACCAGCGGAGTGCACACCTTCCCTGCTG TGCTGCAGTCCTCCGGCCTGTACTCCCTGTCCTCCGTCGTGACCGTGCCTTCCA GCTCTCTGGGCACCCAGACCTACATCTGCAACGTGAACCACAAGCCCTCCAACAC CAAGGTGGACAAGAAGGTGGAACCCAAGTCCTGCGACAAGACCCACACCTGTCC CCCTTGTCCTGCCCCTGAACTGCTGGGCGGACCTTCCGTGTTCCTGTTCCCCCC AAAGCCCAAGGACACCCTGATGATCTCCCGGACCCCCGAAGTGACCTGCGTGGT GGTGGATGTGTCCCACGAGGACCCTGAAGTGAAGTTCAATTGGTACGTGGACG GCGTGGAAGTGCACAACGCCAAGACCAAGCCTAGAGAGGAACAGTACAACTCCA CCTACCGGGTGGTGTCCGTGCTGACCGTGCTGCACCAGGATTGGCTGAACGGCA AAGAGTACAAGTGCAAGGTGTCCAACAAGGCCCTGCCTGCCCCCATCGAAAAGA CCATCTCCAAGGCCAAGGGCCAGCCCCGGGAACCCCAGGTGTACACACTGCCCC CTAGCAGGGACGAGCTGACCAAGAACCAGGTGTCCCTGACCTGTCTCGTGAAAG GCTTCTACCCCTCCGATATCGCCGTGGAATGGGAGTCCAACGGCCAGCCTGAGA ACAACTACAAGACCACCCCCCCTGTGCTGGACTCCGACGGCTCATTCTTCCTGTA CAGCAAGCTGACAGTGGACAAGTCCCGGTGGCAGCAGGGCAACGTGTTCTCCTG CTCCGTGATGCACGAGGCCCTGCACAACCACTACACCCAGAAGTCCCTGTCCCT GAGCCCCGGCAAGTGATGA 523 Human IGHG1*03 Human Heavy Chain Constant gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctgggggcacagcgg IgG1 Region (IGHG1*03) Nucleotide ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgccctgac constant Sequence cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg region tgccctccagcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacac- caagg tggacaagagagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaact cctggggggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct- g aggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactggtacgtggacg gcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaacagcacgtaccgtgtggt cagcgtcctcaccgtcctgcaccaggactggctgaatggcaaggagtacaagtgcaaggtctccaacaa agccctcccagcccccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgt acaccctgcccccatcccgggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggct tctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccac gcctcccgtgctggactccgacggctccttcttcctctatagcaagctcaccgtggacaagagcaggtgg- c agcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcct ctccctgtccccgggtaaa 524 Human Heavy Chain Constant A S T K G P S V F P L A P S S K S T S G G T A A L G C L V K D Y F P E P V T Region (IGHG1*03) Protein V S W N S G A L T S G V H T F P A V L Q S S G L Y S L S S V V T V P S S S Sequence L G T Q T Y I C N V N H K P S N T K V D K R V E P K S C D K T H T C P P C P A P E L L G G P S V F L F P P K P K D T L M I S R T P E V T C V V V D V S H E D P E V K F N W Y V D G V E V H N A K T K P R E E Q Y N S T Y R V V S V L T V L H Q D W L N G K E Y K C K V S N K A L P A P I E K T I S K A K G Q P R E P Q V Y T L P P S R E E M T K N Q V S L T C L V K G F Y P S D I A V E W E S N G Q P E N N Y K T T P P V L D S D G S F F L Y S K L T V D K S R W Q Q G N V F S C S V M H E A L H N H Y T Q K S L S L S P G K 525 Human IGHG1*04 Human Heavy Chain Constant gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctgggggcacagcgg IgG1 Region (IGHG1*04) Nucleotide ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgccctgac constant Sequence cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg region tgccctccagcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacac- caagg tggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaact cctggggggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct- g aggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactggtacgtggacg gcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaacagcacgtaccgtgtggt cagcgtcctcaccgtcctgcaccaggactggctgaatggcaaggagtacaagtgcaaggtctccaacaa agccctcccagcccccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgt acaccctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggctt ctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacg cctcccgtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtggc- a gcaggggaacatcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcctc tccctgtctccgggtaaa 526 Human Heavy Chain Constant ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Region (IGHG1*04) Protein QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKKVEPKSCDKTHTCPPCP Sequence APELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQ PREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVL DSDGSFFLYSKLTVDKSRWQQGNIFSCSVMHEALHNHYTQKSLSLSPGK 527 Human IGHG2*01 Human Heavy Chain Constant gcctccaccaagggcccatcggtcttccccctggcgccctgctccaggagcacctccgagagcacagccg IgG2 & Region (IGHG2*01) Nucleotide ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgctctgac constant IGHG2*03 Sequence cagcggcgtgcacaccttcccagctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg region & tgccctccagcaacttcggcacccagacctacacctgcaacgtagatcacaagcccagcaaca- ccaagg IGHG2*05 tggacaagacagttgagcgcaaatgttgtgtcgagtgcccaccgtgcccagcaccacctgtggcaggac cgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccctgaggtcacgtg- cg tggtggtggacgtgagccacgaagaccccgaggtccagttcaactggtacgtggacggcgtggaggtg cataatgccaagacaaagccacgggaggagcagttcaacagcacgttccgtgtggtcagcgtcctcacc gttgtgcaccaggactggctgaacggcaaggagtacaagtgcaaggtctccaacaaaggcctcccagc ccccatcgagaaaaccatctccaaaaccaaagggcagccccgagaaccacaggtgtacaccctgcccc catcccgggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctaccccagcg acatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacacctcccatgctg gactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtggcagcaggggaac gtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcctctccctgtctc- cg ggtaaa 528 Human Heavy Chain Constant ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Region (IGHG2*01) Protein QSSGLYSLSSVVTVPSSNFGTQTYTCNVDHKPSNTKVDKTVERKCCVECPPCPAPP Sequence VAGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVQFNWYVDGVEVHNAKTK PREEQFNSTFRWSVLTVVHQDWLNGKEYKCKVSNKGLPAPIEKTISKTKGQPREP QVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPMLDSD GSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 529 Human IGHG2*02 Human Heavy Chain Constant GCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCGCCCTGCTCCAGGAGCACC IgG2 Region (IGHG2*02) Nucleotide TCCGAGAGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCG constant Sequence GTGACGGTGTCGTGGAACTCAGGCGCTCTGACCAGCGGCGTGCACACCTTCCCG region GCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGTGACCGTGACC TCCAGCAACTTCGGCACCCAGACCTACACCTGCAACGTAGATCACAAGCCCAGCA ACACCAAGGTGGACAAGACAGTTGAGCGCAAATGTTGTGTCGAGTGCCCACCGT GCCCAGCACCACCTGTGGCAGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACGTGCGTGGTGGTGGACGT GAGCCACGAAGACCCCGAGGTCCAGTTCAACTGGTACGTGGACGGCATGGAGG TGCATAATGCCAAGACAAAGCCACGGGAGGAGCAGTTCAACAGCACGTTCCGTG TGGTCAGCGTCCTCACCGTCGTGCACCAGGACTGGCTGAACGGCAAGGAGTACA AGTGCAAGGTCTCCAACAAAGGCCTCCCAGCCCCCATCGAGAAAACCATCTCCAA AACCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGGGA GGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTACCC CAGCGACATCGCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACA AGACCACACCTCCCATGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCT CACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGICTTCTCATGCTCCGTGAT GCATGAGGCTCTGCACAACCACTACACACAGAAGAGCCTCTCCCTGTCTCCGGG TAAA 530 Human Heavy Chain Constant ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Region (IGHG2*02) Protein QSSGLYSLSSVVTVTSSNFGTQTYTCNVDHKPSNTKVDKTVERKCCVECPPCPAPP Sequence VAGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVQFNWYVDGMEVHNAKT KPREEQFNSTFRVVSVLTVVHQDWLNGKEYKCKVSNKGLPAPIEKTISKTKGQPRE PQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPMLDS DGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 531 Human IGHG2*04 Human Heavy Chain Constant- gcctccaccaagggcccatcggtcttccccctggcgccctgctccaggagcacctccgagagcacagcg IgG2 Region (IGHG2*04) Nucleotide gccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgctctga constant Sequence ccagcggcgtgcacaccttcccagctgtcctacagtcctcaggactctactccctcagcagcgtggtgacc region gtgccctccagcagcttgggcacccagacctacacctgcaacgtagatcacaagcccagcaaca- ccaag gtggacaagacagttgagcgcaaatgttgtgtcgagtgcccaccgtgcccagcaccacctgtggcagga ccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccctgaggtcacgt- gc gtggtggtggacgtgagccacgaagaccccgaggtccagttcaactggtacgtggacggcgtggaggt gcataatgccaagacaaagccacgggaggagcagttcaacagcacgttccgtgtggtcagcgtcctcac cgttgtgcaccaggactggctgaacggcaaggagtacaagtgcaaggtctccaacaaaggcctcccag cccccatcgagaaaaccatctccaaaaccaaagggcagccccgagaaccacaggtgtacaccctgccc ccatcccgggaggagatgaccaagaaccaggtcagcctgacctgcctggtcaaaggcttctaccccagc gacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacacctcccatgct ggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtggcagcaggggaa cgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcctctccctgtct- cc gggtaaa 532 Human Heavy Chain Constant ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Region (IGHG2*04) Protein QSSGLYSLSSWTVPSSSLGTQTYTCNVDHKPSNTINDKTVERKCCVECPPCPAPP Sequence VAGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVQFNWYVDGVEVHNAKTK PREEQFNSTFRVVSVLTVVHQDWLNGKEYKCKVSNKGLPAPIEKTISKTKGQPREP QVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPMLDSD GSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 533 Human IGHG2*06 Human Heavy Chain Constant GCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCGCCCTGCTCCAGGAGCACC IgG2 Region (IGHG2*06) Nucleotide TCCGAGAGCACAGCGGCCCTGGGCTGCCTGGTCAAGGACTACTTCCCCGAACCG constant Sequence GTGACGGTGTCGTGGAACTCAGGCGCTCTGACCAGCGGCGTGCACACCTTCCCG region GCTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGTGACCGTGCCC TCCAGCAACTTCGGCACCCAGACCTACACCTGCAACGTAGATCACAAGCCCAGCA ACACCAAGGTGGACAAGACAGTTGAGCGCAAATGTTGTGTCGAGTGCCCACCGT GCCCAGCACCACCTGTGGCAGGACCGTCAGTCTTCCTCTTCCCCCCAAAACCCAA GGACACCCTCATGATCTCCCGGACCCCTGAGGTCACGTGCGTGGTGGTGGACGT GAGCCACGAAGACCCCGAGGTCCAGTTCAACTGGTACGTGGACGGCGTGGAGG TGCATAATGCCAAGACAAAGCCACGGGAGGAGCAGTTCAACAGCACGTTCCGTG TGGTCAGCGTCCTCACCGTCGTGCACCAGGACTGGCTGAACGGCAAGGAGTACA AGTGCAAGGTCTCCAACAAAGGCCTCCCAGCCCCCATCGAGAAAACCATCTCCAA AACCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGGGA GGAGATGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTACCC CAGCGACATCTCCGTGGAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAA GACCACACCTCCCATGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTC ACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCATGCTCCGTGATG CATGAGGCTCTGCACAACCACTACACACAGAAGAGCCTCTCCCTGTCTCCGGGT AAA 534 Human Heavy Chain Constant ASTKGPSVFPLAPCSRSTSESTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL Region (IGHG2*06) Protein
QSSGLYSLSSWTVPSSNFGTQTYTCNVDHKPSNTINDKTVERKCCVECPPCPAPP Sequence VAGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVQFNWYVDGVEVHNAKTK PREEQFNSTFRVVSVLTVVHQDWLNGKEYKCKVSNKGLPAPIEKTISKTKGQPREP QVYTLPPSREEMTKNQVSLTCLVKGFYPSDISVEWESNGQPENNYKTTPPMLDSD GSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 535 Human C.lamda. IGLC7*03 C.lamda. Light Chain Constant Region GGTCAGCCCAAGGCTGCCCCCTCGGTCACTCTGTTCCCACCCTCCTCTGAGGAG constant (IGLC7*03) Nucleotide CTTCAAGCCAACAAGGCCACACTGGTGTGTCTCGTAAGTGACTTCAACCCGGGA region Sequence GCCGTGACAGTGGCCTGGAAGGCAGATGGCAGCCCCGTCAAGGTGGGAGTGGA GACCACCAAACCCTCCAAACAAAGCAACAACAAGTATGCGGCCAGCAGCTACCT GAGCCTGACGCCCGAGCAGTGGAAGTCCCACAGAAGCTACAGCTGCCGGGTCAC GCATGAAGGGAGCACCGTGGAGAAGACAGTGGCCCCTGCAGAATGCTCT 536 C.lamda. Light Chain Constant Region GQPKAAPSVTLFPPSSEELQANKATLVCLVSDFNPGAVTVAWKADGSPVIWGVETT (IGLC7*03) Amino Acid KPSKQSNNKYAASSYLSLTPEQWKSHRSYSCRVTHEGSTVEKTVAPAECS Sequence 537 Human IGHG1*01 WT human IgG1 nucleotide gcctccaccaagggcccatcggtcttccccctggcaccctcctccaagagcacctctgggggcacagcgg WT IgG1 & sequence #2 ccctgggctgcctggtcaaggactacttccccgaaccggtgacggtgtcgtggaactcaggcgccctgac constant IGHG1*05 cagcggcgtgcacaccttcccggctgtcctacagtcctcaggactctactccctcagcagcgtggtgaccg region (IgG1) tgccctccagcagcttgggcacccagacctacatctgcaacgtgaatcacaagcccagcaacaccaagg tggacaagaaagttgagcccaaatcttgtgacaaaactcacacatgcccaccgtgcccagcacctgaact cctggggggaccgtcagtcttcctcttccccccaaaacccaaggacaccctcatgatctcccggacccct- g aggtcacatgcgtggtggtggacgtgagccacgaagaccctgaggtcaagttcaactggtacgtggacg gcgtggaggtgcataatgccaagacaaagccgcgggaggagcagtacaacagcacgtaccgggtggt cagcgtcctcaccgtcctgcaccaggactggctgaatggcaaggagtacaagtgcaaggtctccaacaa agccctcccagcccccatcgagaaaaccatctccaaagccaaagggcagccccgagaaccacaggtgt acaccctgcccccatcccgggatgagctgaccaagaaccaggtcagcctgacctgcctggtcaaaggctt ctatcccagcgacatcgccgtggagtgggagagcaatgggcagccggagaacaactacaagaccacg cctcccgtgctggactccgacggctccttcttcctctacagcaagctcaccgtggacaagagcaggtggc- a gcaggggaacgtcttctcatgctccgtgatgcatgaggctctgcacaaccactacacgcagaagagcctc tccctgtctccgggtaaa 538 Human C.lamda. IGLC2*01 C.lamda. Light Chain Constant Region GOPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKAGVETTTPSK constant Amino Acid Sequence #2- QSNNKYAASSYLSLTPECIWKSHRSYSCCIVTHEGSTVEKTVAPTECS region Encoded by nucleotide sequence version A & B 539 Human TIGIT-His/Flag NCBI accession number MGWSCIILFLVATATGVHSMMTGTIETTGNISAEKGGSIILQCHLSSTTAQV NM_173799.3 TQVNWEQQDQLLAICNADLGWHISPSFKDRVAPGPGLGLTLQSLTVND Uniprot accession number TGEYFCIYHTYPDGTYTGRIFLEVLESSVAEHGARFQIPIEGRDYKDDDDKH Q495A1-1 HHHHH Expressed protein corresponds to ECD only: Met 22-Pro 141 as per full length sequence Protein domains annotated: Campath leader (double line), extracellular domain (bold), linker & Flag/His tag (italic)
Sequence CWU 1
1
5391290PRTHomo Sapien 1Met Arg Ile Phe Ala Val Phe Ile Phe Met Thr
Tyr Trp His Leu Leu1 5 10 15Asn Ala Phe Thr Val Thr Val Pro Lys Asp
Leu Tyr Val Val Glu Tyr 20 25 30Gly Ser Asn Met Thr Ile Glu Cys Lys
Phe Pro Val Glu Lys Gln Leu 35 40 45Asp Leu Ala Ala Leu Ile Val Tyr
Trp Glu Met Glu Asp Lys Asn Ile 50 55 60Ile Gln Phe Val His Gly Glu
Glu Asp Leu Lys Val Gln His Ser Ser65 70 75 80Tyr Arg Gln Arg Ala
Arg Leu Leu Lys Asp Gln Leu Ser Leu Gly Asn 85 90 95Ala Ala Leu Gln
Ile Thr Asp Val Lys Leu Gln Asp Ala Gly Val Tyr 100 105 110Arg Cys
Met Ile Ser Tyr Gly Gly Ala Asp Tyr Lys Arg Ile Thr Val 115 120
125Lys Val Asn Ala Pro Tyr Asn Lys Ile Asn Gln Arg Ile Leu Val Val
130 135 140Asp Pro Val Thr Ser Glu His Glu Leu Thr Cys Gln Ala Glu
Gly Tyr145 150 155 160Pro Lys Ala Glu Val Ile Trp Thr Ser Ser Asp
His Gln Val Leu Ser 165 170 175Gly Lys Thr Thr Thr Thr Asn Ser Lys
Arg Glu Glu Lys Leu Phe Asn 180 185 190Val Thr Ser Thr Leu Arg Ile
Asn Thr Thr Thr Asn Glu Ile Phe Tyr 195 200 205Cys Thr Phe Arg Arg
Leu Asp Pro Glu Glu Asn His Thr Ala Glu Leu 210 215 220Val Ile Pro
Glu Leu Pro Leu Ala His Pro Pro Asn Glu Arg Thr His225 230 235
240Leu Val Ile Leu Gly Ala Ile Leu Leu Cys Leu Gly Val Ala Leu Thr
245 250 255Phe Ile Phe Arg Leu Arg Lys Gly Arg Met Met Asp Val Lys
Lys Cys 260 265 270Gly Ile Gln Asp Thr Asn Ser Lys Lys Gln Ser Asp
Thr His Leu Glu 275 280 285Glu Thr 2902241PRTCynomologus 2Met Gly
Trp Ser Cys Ile Ile Leu Phe Leu Val Ala Thr Ala Thr Gly1 5 10 15Val
His Ser Met Phe Thr Val Thr Val Pro Lys Asp Leu Tyr Val Val 20 25
30Glu Tyr Gly Ser Asn Met Thr Ile Glu Cys Lys Phe Pro Val Glu Lys
35 40 45Gln Leu Asp Leu Thr Ser Leu Ile Val Tyr Trp Glu Met Glu Asp
Lys 50 55 60Asn Ile Ile Gln Phe Val His Gly Glu Glu Asp Leu Lys Val
Gln His65 70 75 80Ser Asn Tyr Arg Gln Arg Ala Gln Leu Leu Lys Asp
Gln Leu Ser Leu 85 90 95Gly Asn Ala Ala Leu Arg Ile Thr Asp Val Lys
Leu Gln Asp Ala Gly 100 105 110Val Tyr Arg Cys Met Ile Ser Tyr Gly
Gly Ala Asp Tyr Lys Arg Ile 115 120 125Thr Val Lys Val Asn Ala Pro
Tyr Asn Lys Ile Asn Gln Arg Ile Leu 130 135 140Val Val Asp Pro Val
Thr Ser Glu His Glu Leu Thr Cys Gln Ala Glu145 150 155 160Gly Tyr
Pro Lys Ala Glu Val Ile Trp Thr Ser Ser Asp His Gln Val 165 170
175Leu Ser Gly Lys Thr Thr Thr Thr Asn Ser Lys Arg Glu Glu Lys Leu
180 185 190Leu Asn Val Thr Ser Thr Leu Arg Ile Asn Thr Thr Ala Asn
Glu Ile 195 200 205Phe Tyr Cys Ile Phe Arg Arg Leu Asp Pro Glu Glu
Asn His Thr Ala 210 215 220Glu Leu Val Ile Pro Glu Leu Pro Leu Ala
Leu Pro Pro Asn Glu Arg225 230 235 240Thr3245PRTHomo Sapien 3Met
Arg Ile Phe Ala Val Phe Ile Phe Met Thr Tyr Trp His Leu Leu1 5 10
15Asn Ala Phe Thr Val Thr Val Pro Lys Asp Leu Tyr Val Val Glu Tyr
20 25 30Gly Ser Asn Met Thr Ile Glu Cys Lys Phe Pro Val Glu Lys Gln
Leu 35 40 45Asp Leu Ala Ala Leu Ile Val Tyr Trp Glu Met Glu Asp Lys
Asn Ile 50 55 60Ile Gln Phe Val His Gly Glu Glu Asp Leu Lys Val Gln
His Ser Ser65 70 75 80Tyr Arg Gln Arg Ala Arg Leu Leu Lys Asp Gln
Leu Ser Leu Gly Asn 85 90 95Ala Ala Leu Gln Ile Thr Asp Val Lys Leu
Gln Asp Ala Gly Val Tyr 100 105 110Arg Cys Met Ile Ser Tyr Gly Gly
Ala Asp Tyr Lys Arg Ile Thr Val 115 120 125Lys Val Asn Ala Pro Tyr
Asn Lys Ile Asn Gln Arg Ile Leu Val Val 130 135 140Asp Pro Val Thr
Ser Glu His Glu Leu Thr Cys Gln Ala Glu Gly Tyr145 150 155 160Pro
Lys Ala Glu Val Ile Trp Thr Ser Ser Asp His Gln Val Leu Ser 165 170
175Gly Lys Thr Thr Thr Thr Asn Ser Lys Arg Glu Glu Lys Leu Phe Asn
180 185 190Val Thr Ser Thr Leu Arg Ile Asn Thr Thr Thr Asn Glu Ile
Phe Tyr 195 200 205Cys Thr Phe Arg Arg Leu Asp Pro Glu Glu Asn His
Thr Ala Glu Leu 210 215 220Val Ile Pro Glu Leu Pro Leu Ala His Pro
Pro Asn Glu Arg Thr His225 230 235 240His His His His His
2454475PRTHomo Sapien 4Met Arg Ile Phe Ala Val Phe Ile Phe Met Thr
Tyr Trp His Leu Leu1 5 10 15Asn Ala Phe Thr Val Thr Val Pro Lys Asp
Leu Tyr Val Val Glu Tyr 20 25 30Gly Ser Asn Met Thr Ile Glu Cys Lys
Phe Pro Val Glu Lys Gln Leu 35 40 45Asp Leu Ala Ala Leu Ile Val Tyr
Trp Glu Met Glu Asp Lys Asn Ile 50 55 60Ile Gln Phe Val His Gly Glu
Glu Asp Leu Lys Val Gln His Ser Ser65 70 75 80Tyr Arg Gln Arg Ala
Arg Leu Leu Lys Asp Gln Leu Ser Leu Gly Asn 85 90 95Ala Ala Leu Gln
Ile Thr Asp Val Lys Leu Gln Asp Ala Gly Val Tyr 100 105 110Arg Cys
Met Ile Ser Tyr Gly Gly Ala Asp Tyr Lys Arg Ile Thr Val 115 120
125Lys Val Asn Ala Pro Tyr Asn Lys Ile Asn Gln Arg Ile Leu Val Val
130 135 140Asp Pro Val Thr Ser Glu His Glu Leu Thr Cys Gln Ala Glu
Gly Tyr145 150 155 160Pro Lys Ala Glu Val Ile Trp Thr Ser Ser Asp
His Gln Val Leu Ser 165 170 175Gly Lys Thr Thr Thr Thr Asn Ser Lys
Arg Glu Glu Lys Leu Phe Asn 180 185 190Val Thr Ser Thr Leu Arg Ile
Asn Thr Thr Thr Asn Glu Ile Phe Tyr 195 200 205Cys Thr Phe Arg Arg
Leu Asp Pro Glu Glu Asn His Thr Ala Glu Leu 210 215 220Val Ile Pro
Glu Leu Pro Leu Ala His Pro Pro Asn Glu Arg Thr Ile225 230 235
240Glu Gly Arg Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro
245 250 255Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu
Phe Pro 260 265 270Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr
Pro Glu Val Thr 275 280 285Cys Val Val Val Asp Val Ser His Glu Asp
Pro Glu Val Lys Phe Asn 290 295 300Trp Tyr Val Asp Gly Val Glu Val
His Asn Ala Lys Thr Lys Pro Arg305 310 315 320Glu Glu Gln Tyr Asn
Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val 325 330 335Leu His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser 340 345 350Asn
Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys 355 360
365Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp
370 375 380Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys
Gly Phe385 390 395 400Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Asn Gly Gln Pro Glu 405 410 415Asn Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp Ser Asp Gly Ser Phe 420 425 430Phe Leu Tyr Ser Lys Leu Thr
Val Asp Lys Ser Arg Trp Gln Gln Gly 435 440 445Asn Val Phe Ser Cys
Ser Val Met His Glu Ala Leu His Asn His Tyr 450 455 460Thr Gln Lys
Ser Leu Ser Leu Ser Pro Gly Lys465 470 4755249PRTCynomologus 5Met
Gly Trp Ser Cys Ile Ile Leu Phe Leu Val Ala Thr Ala Thr Gly1 5 10
15Val His Ser Met Phe Thr Val Thr Val Pro Lys Asp Leu Tyr Val Val
20 25 30Glu Tyr Gly Ser Asn Met Thr Ile Glu Cys Lys Phe Pro Val Glu
Lys 35 40 45Gln Leu Asp Leu Thr Ser Leu Ile Val Tyr Trp Glu Met Glu
Asp Lys 50 55 60Asn Ile Ile Gln Phe Val His Gly Glu Glu Asp Leu Lys
Val Gln His65 70 75 80Ser Asn Tyr Arg Gln Arg Ala Gln Leu Leu Lys
Asp Gln Leu Ser Leu 85 90 95Gly Asn Ala Ala Leu Arg Ile Thr Asp Val
Lys Leu Gln Asp Ala Gly 100 105 110Val Tyr Arg Cys Met Ile Ser Tyr
Gly Gly Ala Asp Tyr Lys Arg Ile 115 120 125Thr Val Lys Val Asn Ala
Pro Tyr Asn Lys Ile Asn Gln Arg Ile Leu 130 135 140Val Val Asp Pro
Val Thr Ser Glu His Glu Leu Thr Cys Gln Ala Glu145 150 155 160Gly
Tyr Pro Lys Ala Glu Val Ile Trp Thr Ser Ser Asp His Gln Val 165 170
175Leu Ser Gly Lys Thr Thr Thr Thr Asn Ser Lys Arg Glu Glu Lys Leu
180 185 190Leu Asn Val Thr Ser Thr Leu Arg Ile Asn Thr Thr Ala Asn
Glu Ile 195 200 205Phe Tyr Cys Ile Phe Arg Arg Leu Asp Pro Glu Glu
Asn His Thr Ala 210 215 220Glu Leu Val Ile Pro Glu Leu Pro Leu Ala
Leu Pro Pro Asn Glu Arg225 230 235 240Thr Asp Tyr Lys Asp Asp Asp
Asp Lys 2456405PRTHomo Sapien 6Met Gly Trp Ser Cys Ile Ile Leu Phe
Leu Val Ala Thr Ala Thr Gly1 5 10 15Val His Ser Leu Asp Ser Pro Asp
Arg Pro Trp Asn Pro Pro Thr Phe 20 25 30Ser Pro Ala Leu Leu Val Val
Thr Glu Gly Asp Asn Ala Thr Phe Thr 35 40 45Cys Ser Phe Ser Asn Thr
Ser Glu Ser Phe Val Leu Asn Trp Tyr Arg 50 55 60Met Ser Pro Ser Asn
Gln Thr Asp Lys Leu Ala Ala Phe Pro Glu Asp65 70 75 80Arg Ser Gln
Pro Gly Gln Asp Cys Arg Phe Arg Val Thr Gln Leu Pro 85 90 95Asn Gly
Arg Asp Phe His Met Ser Val Val Arg Ala Arg Arg Asn Asp 100 105
110Ser Gly Thr Tyr Leu Cys Gly Ala Ile Ser Leu Ala Pro Lys Ala Gln
115 120 125Ile Lys Glu Ser Leu Arg Ala Glu Leu Arg Val Thr Glu Arg
Arg Ala 130 135 140Glu Val Pro Thr Ala His Pro Ser Pro Ser Pro Arg
Pro Ala Gly Gln145 150 155 160Lys Leu Glu Asn Leu Tyr Phe Gln Gly
Ile Glu Gly Arg Met Asp Glu 165 170 175Pro Lys Ser Cys Asp Lys Thr
His Thr Cys Pro Pro Cys Pro Ala Pro 180 185 190Glu Leu Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys 195 200 205Asp Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val 210 215 220Asp
Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp225 230
235 240Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr 245 250 255Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp 260 265 270Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser Asn Lys Ala Leu 275 280 285Pro Ala Pro Ile Glu Lys Thr Ile Ser
Lys Ala Lys Gly Gln Pro Arg 290 295 300Glu Pro Gln Val Tyr Thr Leu
Pro Pro Ser Arg Asp Glu Leu Thr Lys305 310 315 320Asn Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp 325 330 335Ile Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys 340 345
350Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
355 360 365Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser 370 375 380Cys Ser Val Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser385 390 395 400Leu Ser Leu Ser Pro 40578PRTHomo
Sapien 7Gly Phe Thr Phe Asp Asp Tyr Ala1 588PRTHomo Sapien 8Ile Ser
Trp Lys Ser Asn Ile Ile1 5915PRTHomo Sapien 9Ala Arg Asp Ile Thr
Gly Ser Gly Ser Tyr Gly Trp Phe Asp Pro1 5 10 15105PRTHomo Sapien
10Asp Tyr Ala Met His1 51117PRTHomo Sapien 11Gly Ile Ser Trp Lys
Ser Asn Ile Ile Gly Tyr Ala Asp Ser Val Lys1 5 10 15Gly1213PRTHomo
Sapien 12Asp Ile Thr Gly Ser Gly Ser Tyr Gly Trp Phe Asp Pro1 5
1013122PRTHomo Sapien 13Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
Phe Thr Phe Asp Asp Tyr 20 25 30Ala Met His Trp Val Arg Gln Thr Pro
Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Trp Lys Ser Asn
Ile Ile Gly Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser
Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser
Leu Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Arg Asp Ile
Thr Gly Ser Gly Ser Tyr Gly Trp Phe Asp Pro Trp 100 105 110Gly Gln
Gly Thr Leu Val Thr Val Ser Ser 115 12014508DNAHomo Sapien
14caagaaaaag cttgccgcca ccatggagtt tgggctgagc tggattttcc ttttggctat
60tttaaaaggt gtccagtgtg aagtacaatt ggtggagtcc gggggaggct tggtacagcc
120tggcaggtcc ctgagactct cctgtgcagc ctctggattc acctttgatg
attatgccat 180gcactgggtc cgacaaactc cagggaaggg cctggagtgg
gtctcaggta taagttggaa 240gagtaatatc ataggctatg cggactctgt
gaagggccga ttcaccatct ccagagacaa 300cgccaagaac tccctgtatc
tgcaaatgaa cagtctgaga gctgaggaca cggccttgta 360ttattgtgca
agagatataa cgggttcggg gagttatggc tggttcgacc cctggggcca
420gggaaccctg gtcaccgtct cctcagccaa aacgacaccc ccatctgtct
atccactggc 480ccctgaatct gctaaaactc agcctccg 50815449PRTHomo Sapien
15Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Arg1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Asp Asp
Tyr 20 25 30Ala Met His Trp Val Arg Gln Thr Pro Gly Lys Gly Leu Glu
Trp Val 35 40 45Ser Gly Ile Ser Trp Lys Ser Asn Ile Ile Gly Tyr Ala
Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys
Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Arg Asp Ile Thr Gly Ser Gly Ser
Tyr Gly Trp Phe Asp Pro Trp 100 105 110Gly Gln Gly Thr Leu Val Thr
Val Ser Ser Ala Ser Thr Lys Gly Pro 115 120 125Ser Val Phe Pro Leu
Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr 130 135 140Ala Ala Leu
Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr145 150 155
160Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro
165 170 175Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val
Val Thr 180 185 190Val Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr
Cys Asn Val Asp 195 200 205His Lys Pro Ser Asn Thr Lys Val Asp Lys
Arg Val Glu Ser Lys Tyr 210 215 220Gly Pro Pro Cys Pro Pro Cys Pro
Ala Pro Glu Phe Glu Gly Gly Pro225 230 235 240Ser Val Phe Leu Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser 245 250 255Arg Thr Pro
Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp 260 265 270Pro Glu Val
Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn 275 280 285Ala
Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val 290 295
300Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys
Glu305 310 315 320Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys 325 330 335Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg
Glu Pro Gln Val Tyr Thr 340 345 350Leu Pro Pro Ser Gln Glu Glu Met
Thr Lys Asn Gln Val Ser Leu Thr 355 360 365Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu 370 375 380Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu385 390 395 400Asp
Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys 405 410
415Ser Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu
420 425 430Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
Leu Gly 435 440 445Lys161356DNAHomo Sapien 16gaagtgcagc tggtggaatc
tggcggcgga ctggtgcagc ctggcagatc cctgagactg 60tcttgtgccg cctccggctt
caccttcgac gactacgcta tgcactgggt gcgacagacc 120cctggcaagg
gcctggaatg ggtgtccggc atctcctgga agtccaacat catcggctac
180gccgactccg tgaagggccg gttcaccatc tcccgggaca acgccaagaa
ctccctgtac 240ctgcagatga acagcctgcg ggccgaggac accgccctgt
actactgcgc cagagacatc 300accggctccg gctcctacgg atggttcgat
ccttggggcc agggcaccct cgtgaccgtg 360tcctctgcca gcaccaaggg
cccctctgtg ttccctctgg ccccttccag caagtccacc 420tctggcggaa
cagccgctct gggctgcctc gtgaaggact acttccccga gcctgtgacc
480gtgtcctgga actctggcgc tctgaccagc ggagtgcaca ccttccctgc
tgtgctgcag 540tcctccggcc tgtactccct gtcctccgtc gtgaccgtgc
cttccagctc tctgggcacc 600cagacctaca tctgcaacgt gaaccacaag
ccctccaaca ccaaggtgga caagaaggtg 660gaacccaagt cctgcgacaa
gacccacacc tgtccccctt gtcctgcccc tgaactgctg 720ggcggacctt
ccgtgttcct gttcccccca aagcccaagg acaccctgat gatctcccgg
780acccccgaag tgacctgcgt ggtggtggat gtgtcccacg aggaccctga
agtgaagttc 840aattggtacg tggacggcgt ggaagtgcac aacgccaaga
ccaagcctag agaggaacag 900tacaactcca cctaccgggt ggtgtccgtg
ctgaccgtgc tgcaccagga ttggctgaac 960ggcaaagagt acaagtgcaa
ggtgtccaac aaggccctgc ctgcccccat cgaaaagacc 1020atctccaagg
ccaagggcca gccccgggaa ccccaggtgt acacactgcc ccctagcagg
1080gacgagctga ccaagaacca ggtgtccctg acctgtctcg tgaaaggctt
ctacccctcc 1140gatatcgccg tggaatggga gtccaacggc cagcctgaga
acaactacaa gaccaccccc 1200cctgtgctgg actccgacgg ctcattcttc
ctgtacagca agctgacagt ggacaagtcc 1260cggtggcagc agggcaacgt
gttctcctgc tccgtgatgc acgaggccct gcacaaccac 1320tacacccaga
agtccctgtc cctgagcccc ggcaag 1356176PRTHomo Sapien 17Gln Ser Ile
Ser Ser Tyr1 5183PRTHomo Sapien 18Val Ala Ser1199PRTHomo Sapien
19Gln Gln Ser Tyr Ser Asn Pro Ile Thr1 52011PRTHomo Sapien 20Arg
Ala Ser Gln Ser Ile Ser Ser Tyr Leu Asn1 5 10217PRTHomo Sapien
21Val Ala Ser Ser Leu Gln Ser1 5229PRTHomo Sapien 22Gln Gln Ser Tyr
Ser Asn Pro Ile Thr1 523107PRTHomo Sapien 23Asp Ile Gln Met Thr Gln
Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Tyr 20 25 30Leu Asn Trp Tyr
Gln Gln Lys Pro Gly Lys Ala Pro Lys Pro Leu Ile 35 40 45Tyr Val Ala
Ser Ser Leu Gln Ser Gly Val Pro Ser Ser Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ser Tyr Ser Asn Pro Ile
85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys 100
10524321DNAHomo Sapien 24gacatccaga tgacccagtc tccatcctcc
ctgtctgcat ctgtaggaga cagagtcacc 60atcacttgcc gggcaagtca gagcattagc
agctatttaa attggtatca gcagaaacca 120gggaaagccc ctaagcccct
gatctatgtt gcatccagtt tgcaaagtgg ggtcccatca 180agtttcagtg
gcagtggatc tgggacagat ttcactctca ccatcagcag tctgcaacct
240gaagattttg caacttacta ctgtcaacag agttacagta atccgatcac
cttcggccaa 300gggacacgac tggagatcaa a 32125214PRTHomo Sapien 25Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Tyr
20 25 30Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Pro Leu
Ile 35 40 45Tyr Val Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Ser Phe
Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser
Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ser
Tyr Ser Asn Pro Ile 85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile
Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro
Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys
Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp
Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu
Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170
175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr
180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr
Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys 21026642DNAHomo Sapien
26gacatccaga tgacccagtc tccatcctcc ctgtctgcat ctgtaggaga cagagtcacc
60atcacttgcc gggcaagtca gagcattagc agctatttaa attggtatca gcagaaacca
120gggaaagccc ctaagcccct gatctatgtt gcatccagtt tgcaaagtgg
ggtcccatca 180agtttcagtg gcagtggatc tgggacagat ttcactctca
ccatcagcag tctgcaacct 240gaagattttg caacttacta ctgtcaacag
agttacagta atccgatcac cttcggccaa 300gggacacgac tggagatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc tgcagtccgg
caactcccag 480gaatccgtga ccgagcagga ctccaaggac agcacctact
ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga gaagcacaag
gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc ccgtgaccaa
gtctttcaac cggggcgagt gt 642278PRTHomo Sapien 27Gly Phe Thr Phe Asp
Asp Tyr Ala1 5288PRTHomo Sapien 28Ile Ser Trp Ile Arg Thr Gly Ile1
52916PRTHomo Sapien 29Ala Lys Asp Met Lys Gly Ser Gly Thr Tyr Gly
Gly Trp Phe Asp Thr1 5 10 15305PRTHomo Sapien 30Asp Tyr Ala Met
His1 53117PRTHomo Sapien 31Gly Ile Ser Trp Ile Arg Thr Gly Ile Gly
Tyr Ala Asp Ser Val Lys1 5 10 15Gly3214PRTHomo Sapien 32Asp Met Lys
Gly Ser Gly Thr Tyr Gly Gly Trp Phe Asp Thr1 5 1033123PRTHomo
Sapien 33Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro
Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Asp Asp Tyr 20 25 30Ala Met His Trp Val Arg Gln Val Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Trp Ile Arg Thr Gly Ile Gly
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Phe Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Lys Asp Met Lys Gly Ser
Gly Thr Tyr Gly Gly Trp Phe Asp Thr 100 105 110Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Ser 115 12034482DNAHomo Sapien 34aagcttgccg
ccaccatgga gtttgggctg agctggattt tccttttggc tattttaaaa 60ggtgtccagt
gtgaagtgca gctggtggag tctgggggag gcttggtgca gcctggcagg
120tccctgagac tctcctgtgc agcctctgga ttcacctttg atgattatgc
catgcactgg 180gtccggcaag ttccagggaa gggcctggaa tgggtctcag
gcattagttg gattcgtact 240ggcataggct atgcggactc tgtgaagggc
cgattcacca ttttcagaga caacgccaag 300aattccctgt atctgcaaat
gaacagtctg agagctgagg acacggcctt gtattactgt 360gcaaaagata
tgaagggttc ggggacttat ggggggtggt tcgacacctg gggccaggga
420accctggtca ccgtctcctc agccaaaaca acagccccat cggtctatcc
actggcccct 480gc 48235450PRTHomo Sapien 35Glu Val Gln Leu Val Glu
Ser Gly Gly Gly Leu Val Gln Pro Gly Arg1 5 10 15Ser Leu Arg Leu Ser
Cys Ala Ala Ser Gly Phe Thr Phe Asp Asp Tyr 20 25 30Ala Met His Trp
Val Arg Gln Val Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile
Ser Trp Ile Arg Thr Gly Ile Gly Tyr Ala Asp Ser Val 50 55 60Lys Gly
Arg Phe Thr Ile Phe Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys
85 90 95Ala Lys Asp Met Lys Gly Ser Gly Thr Tyr Gly Gly Trp Phe Asp
Thr 100 105 110Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser
Thr Lys Gly 115 120 125Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg
Ser Thr Ser Glu Ser 130 135 140Thr Ala Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val145 150 155 160Thr Val Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe 165 170 175Pro Ala Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val 180 185 190Thr Val
Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val 195 200
205Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys
210 215 220Tyr Gly Pro Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Glu
Gly Gly225 230 235 240Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys
Asp Thr Leu Met Ile 245 250 255Ser Arg Thr Pro Glu Val Thr Cys Val
Val Val Asp Val Ser Gln Glu 260 265 270Asp Pro Glu Val Gln Phe Asn
Trp Tyr Val Asp Gly Val Glu Val His 275 280 285Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg 290 295 300Val Val Ser
Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys305 310 315
320Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu
325 330 335Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr 340 345 350Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys Asn
Gln Val Ser Leu 355 360 365Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu Trp 370 375 380Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr Lys Thr Thr Pro Pro Val385 390 395 400Leu Asp Ser Asp Gly
Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp 405 410 415Lys Ser Arg
Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His 420 425 430Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu 435 440
445Gly Lys 450361359DNAHomo Sapien 36gaagtgcagc tggtggaatc
tggcggcgga ctggtgcagc ctggcagatc cctgagactg 60tcttgtgccg cctccggctt
caccttcgac gactacgcta tgcactgggt gcgacaggtg 120ccaggcaagg
gcctggaatg ggtgtccggc atctcttgga tccggaccgg catcggctac
180gccgactctg tgaagggccg gttcaccatc ttccgggaca acgccaagaa
ctccctgtac 240ctgcagatga acagcctgcg ggccgaggac accgccctgt
actactgcgc caaggacatg 300aagggctccg gcacctacgg cggatggttc
gatacttggg gccagggcac cctcgtgacc 360gtgtcctctg ccagcaccaa
gggcccctct gtgttccctc tggccccttc cagcaagtcc 420acctctggcg
gaacagccgc tctgggctgc ctcgtgaagg actacttccc cgagcctgtg
480accgtgtcct ggaactctgg cgctctgacc agcggagtgc acaccttccc
tgctgtgctg 540cagtcctccg gcctgtactc cctgtcctcc gtcgtgaccg
tgccttccag ctctctgggc 600acccagacct acatctgcaa cgtgaaccac
aagccctcca acaccaaggt ggacaagaag 660gtggaaccca agtcctgcga
caagacccac acctgtcccc cttgtcctgc ccctgaactg 720ctgggcggac
cttccgtgtt cctgttcccc ccaaagccca aggacaccct gatgatctcc
780cggacccccg aagtgacctg cgtggtggtg gatgtgtccc acgaggaccc
tgaagtgaag 840ttcaattggt acgtggacgg cgtggaagtg cacaacgcca
agaccaagcc tagagaggaa 900cagtacaact ccacctaccg ggtggtgtcc
gtgctgaccg tgctgcacca ggattggctg 960aacggcaaag agtacaagtg
caaggtgtcc aacaaggccc tgcctgcccc catcgaaaag 1020accatctcca
aggccaaggg ccagccccgg gaaccccagg tgtacacact gccccctagc
1080agggacgagc tgaccaagaa ccaggtgtcc ctgacctgtc tcgtgaaagg
cttctacccc 1140tccgatatcg ccgtggaatg ggagtccaac ggccagcctg
agaacaacta caagaccacc 1200ccccctgtgc tggactccga cggctcattc
ttcctgtaca gcaagctgac agtggacaag 1260tcccggtggc agcagggcaa
cgtgttctcc tgctccgtga tgcacgaggc cctgcacaac 1320cactacaccc
agaagtccct gtccctgagc cccggcaag 1359376PRTHomo Sapien 37Gln Ser Ile
Ser Ser Tyr1 5383PRTHomo Sapien 38Val Ala Ser1399PRTHomo Sapien
39Gln Gln Ser Tyr Ser Thr Pro Ile Thr1 54011PRTHomo Sapien 40Arg
Ala Ser Gln Ser Ile Ser Ser Tyr Leu Asn1 5 10417PRTHomo Sapien
41Val Ala Ser Ser Leu Gln Ser1 5429PRTHomo Sapien 42Gln Gln Ser Tyr
Ser Thr Pro Ile Thr1 543107PRTHomo Sapien 43Asp Ile Gln Met Thr Gln
Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Tyr 20 25 30Leu Asn Trp Tyr
Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Val Ala
Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ser Tyr Ser Thr Pro Ile
85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys 100
10544418DNAHomo Sapien 44aaagcttgcc gccaccatga ggctccctgc
tcagcttctg gggctcctgc tactctggct 60ccgaggtgcc agatgtgaca tccagatgac
ccagtctcca tcctccctgt ctgcatctgt 120aggagacaga gtcaccatca
cttgccgggc aagtcagagc attagcagct atttaaattg 180gtatcagcag
aaaccaggga aagcccctaa actcctgatc tatgttgcat ccagtttgca
240aagtggggtc ccatcaaggt tcagtggcag tggatctggg acagatttca
ctctcactat 300cagcagtctg caacctgaag attttgcaac ttactactgt
caacagagtt acagtacccc 360gatcaccttc ggccaaggga cacgtctgga
gatcaaacgt acggatgctg caccaact 41845214PRTHomo Sapien 45Asp Ile Gln
Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg
Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Tyr 20 25 30Leu
Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40
45Tyr Val Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly
50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln
Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ser Tyr Ser
Thr Pro Ile 85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys Arg
Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp
Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu
Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val
Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val
Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser
Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185
190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser
195 200 205Phe Asn Arg Gly Glu Cys 21046642DNAHomo Sapien
46gacatccaga tgacccagtc cccctccagc ctgtctgctt ccgtgggcga cagagtgacc
60atcacctgtc gggcctccca gtccatctcc tcctacctga actggtatca gcagaagccc
120ggcaaggccc ccaagctgct gatctacgtg gccagctctc tgcagtccgg
cgtgccctct 180agattctccg gctctggctc tggcaccgac tttaccctga
ccatcagctc cctgcagccc 240gaggacttcg ccacctacta ctgccagcag
tcctactcca cccctatcac cttcggccag 300ggcacccggc tggaaatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca
gtggaaggtg gacaacgccc tgcagtccgg caactcccag 480gaatccgtga
ccgagcagga ctccaaggac agcacctact ccctgtcctc caccctgacc
540ctgtccaagg ccgactacga gaagcacaag gtgtacgcct gcgaagtgac
ccaccagggc 600ctgtctagcc ccgtgaccaa gtctttcaac cggggcgagt gt
64247450PRTHomo Sapien 47Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Asp Asp Tyr 20 25 30Ala Met His Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Trp Ile Arg
Thr Gly Ile Gly Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile
Phe Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn
Ser Leu Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Lys Asp
Met Lys Gly Ser Gly Thr Tyr Gly Gly Trp Phe Asp Thr 100 105 110Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly 115 120
125Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser
130 135 140Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu
Pro Val145 150 155 160Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser
Gly Val His Thr Phe 165 170 175Pro Ala Val Leu Gln Ser Ser Gly Leu
Tyr Ser Leu Ser Ser Val Val 180 185 190Thr Val Pro Ser Ser Ser Leu
Gly Thr Lys Thr Tyr Thr Cys Asn Val 195 200 205Asp His Lys Pro Ser
Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys 210 215 220Tyr Gly Pro
Pro Cys Pro Pro Cys Pro Ala Pro Glu Leu Ala Gly Ala225 230 235
240Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
245 250 255Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser
Gln Glu 260 265 270Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly
Val Glu Val His 275 280 285Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Phe Asn Ser Thr Tyr Arg 290 295 300Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu Asn Gly Lys305 310 315 320Glu Tyr Lys Cys Lys
Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu 325 330 335Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr 340 345 350Thr
Leu Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu 355 360
365Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp
370 375 380Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro
Pro Val385 390 395 400Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Arg Leu Thr Val Asp 405 410 415Lys Ser Arg Trp Gln Glu Gly Asn Val
Phe Ser Cys Ser Val Met His 420 425 430Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu Ser Leu Ser Leu 435 440 445Gly Lys
45048450PRTHomo Sapien 48Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Asp Asp Tyr 20 25 30Ala Met His Trp Val Arg Gln Val
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Trp Ile Arg
Thr Gly Ile Gly Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn
Ser Leu Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Lys Asp
Met Lys Gly Ser Gly Thr Tyr Gly Gly Trp Phe Asp Thr 100 105 110Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly 115 120
125Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser
130 135 140Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu
Pro Val145 150 155 160Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser
Gly Val His Thr Phe 165 170 175Pro Ala Val Leu Gln Ser Ser Gly Leu
Tyr Ser Leu Ser Ser Val Val 180 185 190Thr Val Pro Ser Ser Ser Leu
Gly Thr Lys Thr Tyr Thr Cys Asn Val 195 200 205Asp His Lys Pro Ser
Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys 210 215 220Tyr Gly Pro
Pro Cys Pro Pro Cys Pro Ala Pro Glu Leu Ala Gly Ala225 230 235
240Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
245 250 255Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser
Gln Glu 260 265 270Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly
Val Glu Val His 275 280 285Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Phe Asn Ser Thr Tyr Arg 290 295 300Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu Asn Gly Lys305 310 315 320Glu Tyr Lys Cys Lys
Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu 325 330 335Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr 340 345 350Thr
Leu Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu 355 360
365Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp
370 375 380Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro
Pro Val385 390 395 400Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Arg Leu Thr Val Asp 405 410 415Lys Ser Arg Trp Gln Glu Gly Asn Val
Phe Ser Cys Ser Val Met His 420 425 430Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu Ser Leu Ser Leu 435 440 445Gly Lys
45049448PRTHomo Sapien 49Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Asp Asp Tyr 20 25 30Ala Met His Trp Val Arg Gln Val
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Trp Ile Arg
Thr Gly Ile Gly Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile
Phe Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn
Ser Leu Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Lys Asp
Met Lys Gly Ser Gly Thr Tyr Gly Gly Trp Phe Asp Thr 100 105 110Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly 115 120
125Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser
130 135 140Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu
Pro Val145 150 155 160Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser
Gly Val His Thr Phe 165 170 175Pro Ala Val Leu Gln Ser Ser Gly Leu
Tyr Ser Leu Ser Ser Val Val 180 185 190Thr Val Pro Ser Ser Ser Leu
Gly Thr Lys Thr Tyr Thr Cys Asn Val 195 200 205Asp His Lys Pro Ser
Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys 210 215 220Tyr Gly Pro
Pro Cys Pro Pro Cys Pro Ala Pro Val Ala Gly Pro Ser225 230 235
240Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
245 250 255Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu
Asp Pro 260 265 270Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His Asn Ala 275 280 285Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn
Ser Thr Tyr Arg Val Val 290 295 300Ser Val Leu Thr Val Leu His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr305 310 315 320Lys Cys Lys Val Ser
Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr 325 330 335Ile Ser Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu 340 345 350Pro
Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360
365Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
370 375 380Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp385 390 395 400Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu
Thr Val Asp Lys Ser 405 410 415Arg Trp Gln Glu Gly Asn Val Phe Ser
Cys Ser Val Met His Glu Ala 420 425 430Leu His Asn His Tyr Thr Gln
Lys Ser Leu Ser Leu Ser Leu Gly Lys 435 440 44550214PRTHomo Sapien
50Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser
Tyr 20 25 30Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
Leu Ile 35 40 45Tyr Ala Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln
Ser Tyr Ser Thr Pro Ile 85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu
Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro
Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val
Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln
Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155
160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys 21051214PRTHomo
Sapien 51Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser
Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile
Ser Ser Tyr 20 25 30Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro
Lys Leu Phe Ile 35 40 45Tyr Val Ala Ser Ser Leu Gln Ser Gly Val Pro
Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr
Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys
Gln Gln Ser Tyr Ser Thr Pro Ile 85 90 95Thr Phe Gly Gln Gly Thr Arg
Leu Glu Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile
Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser
Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys
Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150
155 160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu
Ser 165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His
Lys Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser
Pro Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys
210528PRTHomo Sapien 52Gly Phe Thr Phe Ser Ser Tyr Trp1 5538PRTHomo
Sapien 53Ile Lys Glu Asp Gly Ser Glu Lys1 55412PRTHomo Sapien 54Ala
Arg Asn Arg Leu Tyr Ser Asp Phe Leu Asp Asn1 5 10555PRTHomo Sapien
55Ser Tyr Trp Met Ser1 55617PRTHomo Sapien 56Asn Ile Lys Glu Asp
Gly Ser Glu Lys Tyr Tyr Val Asp Ser Val Lys1 5 10 15Gly5710PRTHomo
Sapien 57Asn Arg Leu Tyr Ser Asp Phe Leu Asp Asn1 5 1058119PRTHomo
Sapien 58Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Ser Ser Tyr 20 25 30Trp Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ala Asn Ile Lys Glu Asp Gly Ser Glu Lys Tyr
Tyr Val Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ser Val Tyr Tyr Cys 85 90 95Ala Arg Asn Arg Leu Tyr Ser
Asp Phe Leu Asp Asn Trp Gly Gln Gly 100 105 110Thr Leu Val Thr Val
Ser Ser 11559358DNAHomo Sapien 59gaggtgcagc tggtggagtc tgggggaggc
ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacgtttagt
agctattgga tgagttgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtggccaac atcaaagaag atggaagtga gaaatactat 180gtcgactctg
tgaagggccg attcaccatc tccagagaca acgccaagaa ctcactgtat
240ctgcaaatga acagcctgag agccgaggac acgtctgtgt attactgtgc
gagaaatcga 300ctctacagtg acttccttga caactggggc cagggaaccc
tggtcaccgt ctcctcag 35860449PRTHomo Sapien 60Glu Val Gln Leu Val
Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu
Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30Trp Met Ser
Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Asn
Ile Lys Glu Asp Gly Ser Glu Lys Tyr Tyr Val Asp Ser Val 50 55 60Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ser Val Tyr Tyr Cys
85 90 95Ala Arg Asn Arg Leu Tyr Ser Asp Phe Leu Asp Asn Trp Gly Gln
Gly 100 105 110Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro
Ser Val Phe 115 120 125Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly
Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu Val Lys Asp Tyr Phe Pro
Glu Pro Val Thr Val Ser Trp145 150 155 160Asn Ser Gly Ala Leu Thr
Ser Gly Val His Thr Phe Pro Ala Val Leu 165 170 175Gln Ser Ser Gly
Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro Ser 180 185 190Ser Ser
Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys Pro 195 200
205Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Asp Lys
210 215 220Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly
Gly Pro225 230 235 240Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser 245 250 255Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser His Glu Asp 260 265 270Pro Glu Val Lys Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn 275 280 285Ala Lys Thr Lys Pro
Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val 290 295 300Val Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu305 310 315
320Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys
325 330 335Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val
Tyr Thr 340 345 350Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln
Val Ser Leu Thr 355 360 365Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu 370 375 380Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Val Leu385 390 395
400Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
405 410 415Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu 420 425 430Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly 435 440 445Lys611347DNAHomo Sapien 61gaggtgcagc
tggtggagtc tgggggaggc ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag
cctctggatt cacgtttagt agctattgga tgagttgggt ccgccaggct
120ccagggaagg ggctggagtg ggtggccaac atcaaagaag atggaagtga
gaaatactat 180gtcgactctg tgaagggccg attcaccatc tccagagaca
acgccaagaa ctcactgtat 240ctgcaaatga acagcctgag agccgaggac
acgtctgtgt attactgtgc gagaaatcga 300ctctacagtg acttccttga
caactggggc cagggaaccc tggtcaccgt ctcctcagcc 360agcaccaagg
gcccctctgt gttccctctg gccccttcca gcaagtccac ctctggcgga
420acagccgctc tgggctgcct cgtgaaggac tacttccccg agcctgtgac
cgtgtcctgg 480aactctggcg ctctgaccag cggagtgcac accttccctg
ctgtgctgca gtcctccggc 540ctgtactccc tgtcctccgt cgtgaccgtg
ccttccagct ctctgggcac ccagacctac 600atctgcaacg tgaaccacaa
gccctccaac accaaggtgg acaagaaggt ggaacccaag 660tcctgcgaca
agacccacac ctgtccccct tgtcctgccc ctgaactgct gggcggacct
720tccgtgttcc tgttcccccc aaagcccaag gacaccctga tgatctcccg
gacccccgaa 780gtgacctgcg tggtggtgga tgtgtcccac gaggaccctg
aagtgaagtt caattggtac 840gtggacggcg tggaagtgca caacgccaag
accaagccta gagaggaaca gtacaactcc 900acctaccggg tggtgtccgt
gctgaccgtg ctgcaccagg attggctgaa cggcaaagag 960tacaagtgca
aggtgtccaa caaggccctg cctgccccca tcgaaaagac catctccaag
1020gccaagggcc agccccggga accccaggtg tacacactgc cccctagcag
ggacgagctg 1080accaagaacc aggtgtccct gacctgtctc gtgaaaggct
tctacccctc cgatatcgcc 1140gtggaatggg agtccaacgg ccagcctgag
aacaactaca agaccacccc ccctgtgctg 1200gactccgacg gctcattctt
cctgtacagc aagctgacag tggacaagtc ccggtggcag 1260cagggcaacg
tgttctcctg ctccgtgatg cacgaggccc tgcacaacca ctacacccag
1320aagtccctgt ccctgagccc cggcaag 1347626PRTHomo Sapien 62Gln Gly
Val Ser Ser Trp1 5633PRTHomo Sapien 63Gly Ala Ser1649PRTHomo Sapien
64Gln Gln Ala Asn Ser Ile Pro Phe Thr1 56511PRTHomo Sapien 65Arg
Ala Ser Gln Gly Val Ser Ser Trp Leu Ala1 5 10667PRTHomo Sapien
66Gly Ala Ser Ser Leu Gln Ser1 5679PRTHomo Sapien 67Gln Gln Ala Asn
Ser Ile Pro Phe Thr1 568107PRTHomo Sapien 68Asp Ile Gln Met Thr Gln
Ser Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Gly Val Ser Ser Trp 20 25 30Leu Ala Trp Tyr
Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Ala
Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Glu Phe Ile Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Ile Pro Phe
85 90 95Thr Phe Gly Pro Gly Thr Lys Val Asp Ile Lys 100
10569322DNAHomo Sapien 69gacatccaga tgacccagtc tccatcttcc
gtgtctgcat ctgtcggaga cagagtcacc 60atcacttgtc gggcgagtca gggtgttagc
agctggttag cctggtatca gcagaaatca 120gggaaagccc ctaagctcct
gatctatggt gcatccagtt tgcaaagtgg ggtcccatca 180agattcagcg
gcagtggatc tgggacagag ttcattctca ccatcagcag cctgcagcct
240gaagattttg caacttacta ttgtcaacag gctaacagta tcccattcac
tttcggccct 300gggaccaaag tggatatcaa ac 32270214PRTHomo Sapien 70Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Gly Val Ser Ser Trp
20 25 30Leu Ala Trp Tyr Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu
Ile 35 40 45Tyr Gly Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe
Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu Phe Ile Leu Thr Ile Ser Ser
Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ala
Asn Ser Ile Pro Phe 85 90 95Thr Phe Gly Pro Gly Thr Lys Val Asp Ile
Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro
Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys
Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp
Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu
Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170
175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr
180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr
Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys 21071642DNAHomo Sapien
71gacatccaga tgacccagtc tccatcttcc gtgtctgcat ctgtcggaga cagagtcacc
60atcacttgtc gggcgagtca gggtgttagc agctggttag cctggtatca gcagaaatca
120gggaaagccc ctaagctcct gatctatggt gcatccagtt tgcaaagtgg
ggtcccatca 180agattcagcg gcagtggatc tgggacagag ttcattctca
ccatcagcag cctgcagcct 240gaagattttg caacttacta ttgtcaacag
gctaacagta tcccattcac tttcggccct 300gggaccaaag tggatatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc tgcagtccgg
caactcccag 480gaatccgtga ccgagcagga ctccaaggac agcacctact
ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga gaagcacaag
gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc ccgtgaccaa
gtctttcaac cggggcgagt gt 642728PRTHomo Sapien 72Gly Phe Thr Phe Ser
Ser Tyr Trp1 5738PRTHomo Sapien 73Ile Lys Glu Asp Gly Ser Glu Lys1
57412PRTHomo Sapien 74Ala Arg Val Arg Leu Tyr Ser Asp Phe Leu Asp
Tyr1 5 10755PRTHomo Sapien 75Ser Tyr Trp Met Ser1 57617PRTHomo
Sapien 76Asn Ile Lys Glu Asp Gly Ser Glu Lys Tyr Tyr Val Asp Ser
Leu Lys1 5 10 15Gly7710PRTHomo Sapien 77Val Arg Leu Tyr Ser Asp Phe
Leu Asp Tyr1 5 1078119PRTHomo Sapien 78Glu Val Gln Leu Val Asp Ser
Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30Trp Met Ser Trp Val
Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Asn Ile Lys
Glu Asp Gly Ser Glu Lys Tyr Tyr Val Asp Ser Leu 50 55 60Lys Gly Arg
Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu
Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ser Val Tyr Tyr Cys 85 90
95Ala Arg Val Arg Leu Tyr Ser Asp Phe Leu Asp Tyr Trp Gly Gln Gly
100 105 110Thr Leu Val Thr Val Ser Ser 11579358DNAHomo Sapien
79gaggtgcagc tggtggactc tgggggaggc ttggtccagc ctggggggtc cctgagactc
60tcctgtgcag cctctggatt cacgtttagt agctattgga tgagttgggt ccgccaggct
120ccaggaaagg ggctggagtg ggtggccaac ataaaagaag atggaagtga
gaaatactat 180gtagactctt tgaagggccg attcaccatc tccagagaca
acgccaagaa ctcactgtat 240ctgcaaatga acagcctgag agccgaggac
acgtctgtgt attactgtgc gagagttcga 300ctctacagtg acttccttga
ctactggggc cagggaaccc tggtcaccgt ctcctcag 35880449PRTHomo Sapien
80Glu Val Gln Leu Val Asp Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser
Tyr 20 25 30Trp Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu
Trp Val 35 40 45Ala Asn Ile Lys Glu Asp Gly Ser Glu Lys Tyr Tyr Val
Asp Ser Leu 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys
Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr Ser Val Tyr Tyr Cys 85 90 95Ala Arg Val Arg Leu Tyr Ser Asp Phe
Leu Asp Tyr Trp Gly Gln Gly 100 105 110Thr Leu Val Thr Val Ser Ser
Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala Pro Ser
Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150 155
160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu
165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro Ser 180 185 190Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val
Asn His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Lys Val Glu
Pro Lys Ser Cys Asp Lys 210 215 220Thr His Thr Cys Pro Pro Cys Pro
Ala Pro Glu Leu Leu Gly Gly Pro225 230 235 240Ser Val Phe Leu Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser 245 250 255Arg Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp 260 265 270Pro
Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn 275 280
285Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val
290 295 300Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly
Lys Glu305 310 315 320Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro
Ala Pro Ile Glu Lys 325 330 335Thr Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr 340 345 350Leu Pro Pro Ser Arg Asp Glu
Leu Thr Lys Asn Gln Val Ser Leu Thr 355 360 365Cys Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu 370 375 380Ser Asn Gly
Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu385 390 395
400Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
405 410 415Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu 420 425 430Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly 435 440 445Lys811347DNAHomo Sapien 81gaggtgcagc
tggtggactc tgggggaggc ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag
cctctggatt cacgtttagt agctattgga tgagttgggt ccgccaggct
120ccaggaaagg ggctggagtg ggtggccaac ataaaagaag atggaagtga
gaaatactat 180gtagactctt tgaagggccg attcaccatc tccagagaca
acgccaagaa ctcactgtat 240ctgcaaatga acagcctgag agccgaggac
acgtctgtgt attactgtgc gagagttcga 300ctctacagtg acttccttga
ctactggggc cagggaaccc tggtcaccgt ctcctcagcc 360agcaccaagg
gcccctctgt gttccctctg gccccttcca gcaagtccac ctctggcgga
420acagccgctc tgggctgcct cgtgaaggac tacttccccg agcctgtgac
cgtgtcctgg 480aactctggcg ctctgaccag cggagtgcac accttccctg
ctgtgctgca gtcctccggc 540ctgtactccc tgtcctccgt cgtgaccgtg
ccttccagct ctctgggcac ccagacctac 600atctgcaacg tgaaccacaa
gccctccaac accaaggtgg acaagaaggt ggaacccaag 660tcctgcgaca
agacccacac ctgtccccct tgtcctgccc ctgaactgct gggcggacct
720tccgtgttcc tgttcccccc aaagcccaag gacaccctga tgatctcccg
gacccccgaa 780gtgacctgcg tggtggtgga tgtgtcccac gaggaccctg
aagtgaagtt caattggtac 840gtggacggcg tggaagtgca caacgccaag
accaagccta gagaggaaca gtacaactcc 900acctaccggg tggtgtccgt
gctgaccgtg ctgcaccagg attggctgaa cggcaaagag 960tacaagtgca
aggtgtccaa caaggccctg cctgccccca tcgaaaagac catctccaag
1020gccaagggcc agccccggga accccaggtg tacacactgc cccctagcag
ggacgagctg 1080accaagaacc aggtgtccct gacctgtctc gtgaaaggct
tctacccctc cgatatcgcc 1140gtggaatggg agtccaacgg ccagcctgag
aacaactaca agaccacccc ccctgtgctg 1200gactccgacg gctcattctt
cctgtacagc aagctgacag tggacaagtc ccggtggcag 1260cagggcaacg
tgttctcctg ctccgtgatg cacgaggccc tgcacaacca ctacacccag
1320aagtccctgt ccctgagccc cggcaag 1347826PRTHomo Sapien 82Gln Gly
Val Ser Ser Trp1 5833PRTHomo Sapien 83Gly Ala Ser1849PRTHomo Sapien
84Gln Gln Ala Asn Ser Ile Pro Phe Thr1 58511PRTHomo Sapien 85Arg
Ala Ser Gln Gly Val Ser Ser Trp Leu Ala1 5 10867PRTHomo Sapien
86Gly Ala Ser Ser Leu Gln Ser1 5879PRTHomo Sapien 87Gln Gln Ala Asn
Ser Ile Pro Phe Thr1 588107PRTHomo Sapien 88Asp Ile Gln Met Thr Gln
Ser Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Gly Val Ser Ser Trp 20 25 30Leu Ala Trp Tyr
Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Ala
Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Glu Phe Ile Leu Ser Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Ile Pro Phe
85 90 95Thr Phe Gly Pro Gly Thr Lys Val Asp Ile Lys 100
10589322DNAHomo Sapien 89gacatccaga tgacccagtc tccatcttcc
gtgtctgcat ctgtcggaga cagagtcacc 60atcacttgtc gggcgagtca gggtgttagc
agttggttag cctggtatca gcagaaatca 120gggaaagccc ctaagctcct
gatctatggt gcctccagtt tgcaaagtgg ggtcccatca 180agattcagcg
gcagtggatc tgggacagag ttcattctca gcatcagcag cctgcagcct
240gaagattttg caacttacta ttgtcaacag gctaacagta tcccattcac
tttcggccct 300gggaccaaag tggatatcaa ac 32290214PRTHomo Sapien 90Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Gly Val Ser Ser Trp
20 25 30Leu Ala Trp Tyr Gln Gln Lys Ser Gly Lys Ala Pro Lys Leu Leu
Ile 35 40 45Tyr Gly Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe
Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu Phe Ile Leu Ser Ile Ser Ser
Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ala
Asn Ser Ile Pro Phe 85 90 95Thr Phe Gly Pro Gly Thr Lys Val Asp Ile
Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro
Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys
Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp
Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu
Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170
175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr
180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr
Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys 21091642DNAHomo Sapien
91gacatccaga tgacccagtc tccatcttcc gtgtctgcat ctgtcggaga cagagtcacc
60atcacttgtc gggcgagtca gggtgttagc agttggttag cctggtatca gcagaaatca
120gggaaagccc ctaagctcct gatctatggt gcctccagtt tgcaaagtgg
ggtcccatca 180agattcagcg gcagtggatc tgggacagag ttcattctca
gcatcagcag cctgcagcct 240gaagattttg caacttacta ttgtcaacag
gctaacagta tcccattcac tttcggccct 300gggaccaaag tggatatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc tgcagtccgg
caactcccag 480gaatccgtga ccgagcagga ctccaaggac agcacctact
ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga gaagcacaag
gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc ccgtgaccaa
gtctttcaac cggggcgagt gt 642929PRTHomo Sapien 92Gly Gly Ser Ile Ile
Ser Ser Asp Trp1 5937PRTHomo Sapien 93Ile Phe His Ser Gly Arg Thr1
5948PRTHomo Sapien 94Ala Arg Asp Gly Ser Gly Ser Tyr1 5956PRTHomo
Sapien 95Ser Ser Asp Trp Trp Asn1 59616PRTHomo Sapien 96Glu Ile Phe
His Ser Gly Arg Thr Asn Tyr Asn Pro Ser Leu Lys Ser1 5 10
15976PRTHomo Sapien 97Asp Gly Ser Gly Ser Tyr1 598115PRTHomo Sapien
98Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Gly1
5 10 15Thr Leu Ser Leu Thr Cys Ile Val Ser Gly Gly Ser Ile Ile Ser
Ser 20 25 30Asp Trp Trp Asn Trp Val Arg Gln Pro Pro Gly Lys Gly Leu
Glu Trp 35 40 45Ile Gly Glu Ile Phe His Ser Gly Arg Thr Asn Tyr Asn
Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Ile Ser Ile Asp Lys Ser Lys
Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala Ala Asp
Thr
Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Gly Ser Gly Ser Tyr Trp Gly
Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ser 11599346DNAHomo
Sapien 99caggtgcagc tgcaggagtc gggcccagga ctggtgaagc cttcggggac
cctgtccctc 60acctgcattg tctctggtgg ctccatcatc agtagtgact ggtggaattg
ggtccgccag 120cccccaggga aggggctgga gtggattgga gaaatctttc
atagtgggag gaccaactac 180aacccgtccc tcaagagtcg agtcaccata
tcaatagaca agtccaagaa tcagttctcc 240ctgaggctga gctctgtgac
cgccgcggac acggccgtgt attactgtgc gagagatggt 300tcggggagtt
actggggcca gggaaccctg gtcaccgtct cctcag 346100445PRTHomo Sapien
100Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Gly1
5 10 15Thr Leu Ser Leu Thr Cys Ile Val Ser Gly Gly Ser Ile Ile Ser
Ser 20 25 30Asp Trp Trp Asn Trp Val Arg Gln Pro Pro Gly Lys Gly Leu
Glu Trp 35 40 45Ile Gly Glu Ile Phe His Ser Gly Arg Thr Asn Tyr Asn
Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Ile Ser Ile Asp Lys Ser Lys
Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Gly Ser Gly Ser Tyr Trp
Gly Gln Gly Thr Leu Val Thr 100 105 110Val Ser Ser Ala Ser Thr Lys
Gly Pro Ser Val Phe Pro Leu Ala Pro 115 120 125Ser Ser Lys Ser Thr
Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val 130 135 140Lys Asp Tyr
Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala145 150 155
160Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly
165 170 175Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser
Leu Gly 180 185 190Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys Pro
Ser Asn Thr Lys 195 200 205Val Asp Lys Lys Val Glu Pro Lys Ser Cys
Asp Lys Thr His Thr Cys 210 215 220Pro Pro Cys Pro Ala Pro Glu Leu
Leu Gly Gly Pro Ser Val Phe Leu225 230 235 240Phe Pro Pro Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu 245 250 255Val Thr Cys
Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys 260 265 270Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys 275 280
285Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu
290 295 300Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys305 310 315 320Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu
Lys Thr Ile Ser Lys 325 330 335Ala Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser 340 345 350Arg Asp Glu Leu Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys 355 360 365Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln 370 375 380Pro Glu Asn
Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly385 390 395
400Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln
405 410 415Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn 420 425 430His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly
Lys 435 440 4451011335DNAHomo Sapien 101caggtgcagc tgcaggagtc
gggcccagga ctggtgaagc cttcggggac cctgtccctc 60acctgcattg tctctggtgg
ctccatcatc agtagtgact ggtggaattg ggtccgccag 120cccccaggga
aggggctgga gtggattgga gaaatctttc atagtgggag gaccaactac
180aacccgtccc tcaagagtcg agtcaccata tcaatagaca agtccaagaa
tcagttctcc 240ctgaggctga gctctgtgac cgccgcggac acggccgtgt
attactgtgc gagagatggt 300tcggggagtt actggggcca gggaaccctg
gtcaccgtct cctcagccag caccaagggc 360ccctctgtgt tccctctggc
cccttccagc aagtccacct ctggcggaac agccgctctg 420ggctgcctcg
tgaaggacta cttccccgag cctgtgaccg tgtcctggaa ctctggcgct
480ctgaccagcg gagtgcacac cttccctgct gtgctgcagt cctccggcct
gtactccctg 540tcctccgtcg tgaccgtgcc ttccagctct ctgggcaccc
agacctacat ctgcaacgtg 600aaccacaagc cctccaacac caaggtggac
aagaaggtgg aacccaagtc ctgcgacaag 660acccacacct gtcccccttg
tcctgcccct gaactgctgg gcggaccttc cgtgttcctg 720ttccccccaa
agcccaagga caccctgatg atctcccgga cccccgaagt gacctgcgtg
780gtggtggatg tgtcccacga ggaccctgaa gtgaagttca attggtacgt
ggacggcgtg 840gaagtgcaca acgccaagac caagcctaga gaggaacagt
acaactccac ctaccgggtg 900gtgtccgtgc tgaccgtgct gcaccaggat
tggctgaacg gcaaagagta caagtgcaag 960gtgtccaaca aggccctgcc
tgcccccatc gaaaagacca tctccaaggc caagggccag 1020ccccgggaac
cccaggtgta cacactgccc cctagcaggg acgagctgac caagaaccag
1080gtgtccctga cctgtctcgt gaaaggcttc tacccctccg atatcgccgt
ggaatgggag 1140tccaacggcc agcctgagaa caactacaag accacccccc
ctgtgctgga ctccgacggc 1200tcattcttcc tgtacagcaa gctgacagtg
gacaagtccc ggtggcagca gggcaacgtg 1260ttctcctgct ccgtgatgca
cgaggccctg cacaaccact acacccagaa gtccctgtcc 1320ctgagccccg gcaag
133510212PRTHomo Sapien 102Gln Ser Val Leu Tyr Ser Ser Asn Asn Lys
Asn Tyr1 5 101033PRTHomo Sapien 103Trp Ala Ser11048PRTHomo Sapien
104Gln Gln Tyr Tyr Ser Asn Arg Ser1 510517PRTHomo Sapien 105Lys Ser
Ser Gln Ser Val Leu Tyr Ser Ser Asn Asn Lys Asn Tyr Leu1 5 10
15Ala1067PRTHomo Sapien 106Trp Ala Ser Thr Arg Glu Ser1
51078PRTHomo Sapien 107Gln Gln Tyr Tyr Ser Asn Arg Ser1
5108112PRTHomo Sapien 108Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Ser Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Thr Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser
Asn Arg Ser Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100 105
110109337DNAHomo Sapien 109gacatcgtga tgacccagtc tccagactcc
ctggctgtgt ctctgggcga gagggccacc 60atcaactgca agtccagcca gagtgtttta
tacagctcca acaataagaa ttacttagct 120tggtaccagc agaaatcagg
acagcctcct aagttgctca tttactgggc atctacccgg 180gaatccgggg
tccctgaccg attcagtggc agcgggtctg ggacagattt cactctcacc
240atcagcagcc tgcagactga agatgtggca gtttattact gtcagcaata
ttatagtaat 300cgcagttttg gccaggggac caagctggag atcaaac
337110219PRTHomo Sapien 110Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Ser Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Thr Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser
Asn Arg Ser Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100 105 110Arg
Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu 115 120
125Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
130 135 140Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln145 150 155 160Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser 165 170 175Thr Tyr Ser Leu Ser Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu 180 185 190Lys His Lys Val Tyr Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser 195 200 205Pro Val Thr Lys Ser
Phe Asn Arg Gly Glu Cys 210 215111657DNAHomo Sapien 111gacatcgtga
tgacccagtc tccagactcc ctggctgtgt ctctgggcga gagggccacc 60atcaactgca
agtccagcca gagtgtttta tacagctcca acaataagaa ttacttagct
120tggtaccagc agaaatcagg acagcctcct aagttgctca tttactgggc
atctacccgg 180gaatccgggg tccctgaccg attcagtggc agcgggtctg
ggacagattt cactctcacc 240atcagcagcc tgcagactga agatgtggca
gtttattact gtcagcaata ttatagtaat 300cgcagttttg gccaggggac
caagctggag atcaaacgta cggtggccgc tccctccgtg 360ttcatcttcc
caccttccga cgagcagctg aagtccggca ccgcttctgt cgtgtgcctg
420ctgaacaact tctacccccg cgaggccaag gtgcagtgga aggtggacaa
cgccctgcag 480tccggcaact cccaggaatc cgtgaccgag caggactcca
aggacagcac ctactccctg 540tcctccaccc tgaccctgtc caaggccgac
tacgagaagc acaaggtgta cgcctgcgaa 600gtgacccacc agggcctgtc
tagccccgtg accaagtctt tcaaccgggg cgagtgt 6571128PRTHomo Sapien
112Gly Phe Thr Phe Ser Ser Tyr Trp1 51138PRTHomo Sapien 113Ile Lys
Glu Asp Gly Ser Glu Lys1 511412PRTHomo Sapien 114Ala Arg Asn Arg
Leu Tyr Ser Asp Phe Leu Asp Asn1 5 101155PRTHomo Sapien 115Ser Tyr
Trp Met Ser1 511617PRTHomo Sapien 116Asn Ile Lys Glu Asp Gly Ser
Glu Lys Tyr Tyr Val Asp Ser Val Lys1 5 10 15Gly11710PRTHomo Sapien
117Asn Arg Leu Tyr Ser Asp Phe Leu Asp Asn1 5 10118119PRTHomo
Sapien 118Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Ser Ser Tyr 20 25 30Trp Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ala Asn Ile Lys Glu Asp Gly Ser Glu Lys Tyr
Tyr Val Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ser Val Tyr Tyr Cys 85 90 95Ala Arg Asn Arg Leu Tyr Ser
Asp Phe Leu Asp Asn Trp Gly Gln Gly 100 105 110Thr Leu Val Thr Val
Ser Ser 115119358DNAHomo Sapien 119gaggtgcagc tggtggagtc tgggggaggc
ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacgtttagt
agctattgga tgagttgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtggccaac atcaaagaag atggaagtga gaaatactat 180gtcgactctg
tgaagggccg attcaccatc tccagagaca acgccaagaa ctcactgtat
240ctgcaaatga acagcctgag agccgaggac acgtctgtgt attactgtgc
gagaaatcga 300ctctacagtg acttccttga caactggggc cagggaaccc
tggtcaccgt ctcctcag 358120449PRTHomo Sapien 120Glu Val Gln Leu Val
Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu
Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30Trp Met Ser
Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Asn
Ile Lys Glu Asp Gly Ser Glu Lys Tyr Tyr Val Asp Ser Val 50 55 60Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ser Val Tyr Tyr Cys
85 90 95Ala Arg Asn Arg Leu Tyr Ser Asp Phe Leu Asp Asn Trp Gly Gln
Gly 100 105 110Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro
Ser Val Phe 115 120 125Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly
Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu Val Lys Asp Tyr Phe Pro
Glu Pro Val Thr Val Ser Trp145 150 155 160Asn Ser Gly Ala Leu Thr
Ser Gly Val His Thr Phe Pro Ala Val Leu 165 170 175Gln Ser Ser Gly
Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro Ser 180 185 190Ser Ser
Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys Pro 195 200
205Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Asp Lys
210 215 220Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly
Gly Pro225 230 235 240Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser 245 250 255Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser His Glu Asp 260 265 270Pro Glu Val Lys Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn 275 280 285Ala Lys Thr Lys Pro
Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val 290 295 300Val Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu305 310 315
320Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys
325 330 335Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val
Tyr Thr 340 345 350Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln
Val Ser Leu Thr 355 360 365Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu 370 375 380Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Val Leu385 390 395 400Asp Ser Asp Gly Ser
Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys 405 410 415Ser Arg Trp
Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu 420 425 430Ala
Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly 435 440
445Lys1211347DNAHomo Sapien 121gaggtgcagc tggtggagtc tgggggaggc
ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacgtttagt
agctattgga tgagttgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtggccaac atcaaagaag atggaagtga gaaatactat 180gtcgactctg
tgaagggccg attcaccatc tccagagaca acgccaagaa ctcactgtat
240ctgcaaatga acagcctgag agccgaggac acgtctgtgt attactgtgc
gagaaatcga 300ctctacagtg acttccttga caactggggc cagggaaccc
tggtcaccgt ctcctcagcc 360agcaccaagg gcccctctgt gttccctctg
gccccttcca gcaagtccac ctctggcgga 420acagccgctc tgggctgcct
cgtgaaggac tacttccccg agcctgtgac cgtgtcctgg 480aactctggcg
ctctgaccag cggagtgcac accttccctg ctgtgctgca gtcctccggc
540ctgtactccc tgtcctccgt cgtgaccgtg ccttccagct ctctgggcac
ccagacctac 600atctgcaacg tgaaccacaa gccctccaac accaaggtgg
acaagaaggt ggaacccaag 660tcctgcgaca agacccacac ctgtccccct
tgtcctgccc ctgaactgct gggcggacct 720tccgtgttcc tgttcccccc
aaagcccaag gacaccctga tgatctcccg gacccccgaa 780gtgacctgcg
tggtggtgga tgtgtcccac gaggaccctg aagtgaagtt caattggtac
840gtggacggcg tggaagtgca caacgccaag accaagccta gagaggaaca
gtacaactcc 900acctaccggg tggtgtccgt gctgaccgtg ctgcaccagg
attggctgaa cggcaaagag 960tacaagtgca aggtgtccaa caaggccctg
cctgccccca tcgaaaagac catctccaag 1020gccaagggcc agccccggga
accccaggtg tacacactgc cccctagcag ggacgagctg 1080accaagaacc
aggtgtccct gacctgtctc gtgaaaggct tctacccctc cgatatcgcc
1140gtggaatggg agtccaacgg ccagcctgag aacaactaca agaccacccc
ccctgtgctg 1200gactccgacg gctcattctt cctgtacagc aagctgacag
tggacaagtc ccggtggcag 1260cagggcaacg tgttctcctg ctccgtgatg
cacgaggccc tgcacaacca ctacacccag 1320aagtccctgt ccctgagccc cggcaag
13471226PRTHomo Sapien 122Gln Gly Val Ser Ser Trp1 51233PRTHomo
Sapien 123Gly Ala Ser11249PRTHomo Sapien 124Gln Gln Ala Asn Ser Ile
Pro Phe Thr1 512511PRTHomo Sapien 125Arg Ala Ser Gln Gly Val Ser
Ser Trp Leu Ala1 5 101267PRTHomo Sapien 126Gly Ala Ser Ser Leu Gln
Ser1 51279PRTHomo Sapien 127Gln Gln Ala Asn Ser Ile Pro Phe Thr1
5128107PRTHomo Sapien 128Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala
Ser Gln Gly Val Ser Ser Trp 20 25 30Leu Ala Trp Tyr Gln Gln Lys Ser
Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Ala Ser Ser Leu Gln
Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu
Phe Ile Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Ile Pro Phe 85 90 95Thr Phe Gly
Pro Gly Thr Lys Val Asp Ile Lys 100 105129322DNAHomo Sapien
129gacatccaga tgacccagtc tccatcttcc gtgtctgcat ctgtcggaga
cagagtcacc 60atcacttgtc
gggcgagtca gggtgttagc agctggttag cctggtatca gcagaaatca
120gggaaagccc ctaagctcct gatctatggt gcatccagtt tgcaaagtgg
ggtcccatca 180agattcagcg gcagtggatc tgggacagag ttcattctca
ccatcagcag cctgcagcct 240gaagattttg caacttacta ttgtcaacag
gctaacagta tcccattcac tttcggccct 300gggaccaaag tggatatcaa ac
322130214PRTHomo Sapien 130Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala
Ser Gln Gly Val Ser Ser Trp 20 25 30Leu Ala Trp Tyr Gln Gln Lys Ser
Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Ala Ser Ser Leu Gln
Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu
Phe Ile Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Ile Pro Phe 85 90 95Thr Phe Gly
Pro Gly Thr Lys Val Asp Ile Lys Arg Thr Val Ala Ala 100 105 110Pro
Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120
125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala
130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn
Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser
Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp
Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln
Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu
Cys 210131642DNAHomo Sapien 131gacatccaga tgacccagtc tccatcttcc
gtgtctgcat ctgtcggaga cagagtcacc 60atcacttgtc gggcgagtca gggtgttagc
agctggttag cctggtatca gcagaaatca 120gggaaagccc ctaagctcct
gatctatggt gcatccagtt tgcaaagtgg ggtcccatca 180agattcagcg
gcagtggatc tgggacagag ttcattctca ccatcagcag cctgcagcct
240gaagattttg caacttacta ttgtcaacag gctaacagta tcccattcac
tttcggccct 300gggaccaaag tggatatcaa acgtacggtg gccgctccct
ccgtgttcat cttcccacct 360tccgacgagc agctgaagtc cggcaccgct
tctgtcgtgt gcctgctgaa caacttctac 420ccccgcgagg ccaaggtgca
gtggaaggtg gacaacgccc tgcagtccgg caactcccag 480gaatccgtga
ccgagcagga ctccaaggac agcacctact ccctgtcctc caccctgacc
540ctgtccaagg ccgactacga gaagcacaag gtgtacgcct gcgaagtgac
ccaccagggc 600ctgtctagcc ccgtgaccaa gtctttcaac cggggcgagt gt
6421328PRTHomo Sapien 132Gly Phe Thr Phe Arg Ile Tyr Gly1
51338PRTHomo Sapien 133Ile Trp Tyr Asp Gly Ser Asn Lys1
513411PRTHomo Sapien 134Ala Arg Asp Met Asp Tyr Phe Gly Met Asp
Val1 5 101355PRTHomo Sapien 135Ile Tyr Gly Met His1 513617PRTHomo
Sapien 136Val Ile Trp Tyr Asp Gly Ser Asn Lys Tyr Tyr Ala Asp Ser
Val Lys1 5 10 15Gly1379PRTHomo Sapien 137Asp Met Asp Tyr Phe Gly
Met Asp Val1 5138118PRTHomo Sapien 138Gln Val Gln Leu Val Glu Ser
Gly Gly Gly Val Val Gln Pro Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Arg Ile Tyr 20 25 30Gly Met His Trp Val
Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Val Ile Trp
Tyr Asp Gly Ser Asn Lys Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg
Phe Thr Ile Ser Arg Asp Asn Ser Asp Asn Thr Leu Tyr65 70 75 80Leu
Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85 90
95Ala Arg Asp Met Asp Tyr Phe Gly Met Asp Val Trp Gly Gln Gly Thr
100 105 110Thr Val Thr Val Ser Ser 115139355DNAHomo Sapien
139caggtgcagc tggtggagtc tgggggaggc gtggtccagc ctgggaggtc
cctgagactc 60tcctgtgcag cgtctggatt caccttccgt atttatggca tgcactgggt
ccgccaggct 120ccaggcaagg ggctggagtg ggtggcagtt atatggtatg
atggaagtaa taaatactat 180gctgactccg tgaagggccg attcaccatc
tccagagaca attccgacaa cacgctgtat 240ctgcaaatga acagcctgag
agccgaggac acggctgtgt attactgtgc gagagatatg 300gactacttcg
gtatggacgt ctggggccaa gggaccacgg tcaccgtctc ctcag 355140448PRTHomo
Sapien 140Gln Val Gln Leu Val Glu Ser Gly Gly Gly Val Val Gln Pro
Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Arg Ile Tyr 20 25 30Gly Met His Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ala Val Ile Trp Tyr Asp Gly Ser Asn Lys Tyr
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ser Asp Asn Thr Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Met Asp Tyr Phe
Gly Met Asp Val Trp Gly Gln Gly Thr 100 105 110Thr Val Thr Val Ser
Ser Ala Ser Thr Lys Gly Pro Ser Val Phe Pro 115 120 125Leu Ala Pro
Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly 130 135 140Cys
Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp Asn145 150
155 160Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu
Gln 165 170 175Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro Ser Ser 180 185 190Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val
Asn His Lys Pro Ser 195 200 205Asn Thr Lys Val Asp Lys Lys Val Glu
Pro Lys Ser Cys Asp Lys Thr 210 215 220His Thr Cys Pro Pro Cys Pro
Ala Pro Glu Leu Leu Gly Gly Pro Ser225 230 235 240Val Phe Leu Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro 260 265
270Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
275 280 285Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg
Val Val 290 295 300Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn
Gly Lys Glu Tyr305 310 315 320Lys Cys Lys Val Ser Asn Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr 325 330 335Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu 340 345 350Pro Pro Ser Arg Asp
Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380Asn
Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp385 390
395 400Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser 405 410 415Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala 420 425 430Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 435 440 4451411344DNAHomo Sapien 141caggtgcagc
tggtggagtc tgggggaggc gtggtccagc ctgggaggtc cctgagactc 60tcctgtgcag
cgtctggatt caccttccgt atttatggca tgcactgggt ccgccaggct
120ccaggcaagg ggctggagtg ggtggcagtt atatggtatg atggaagtaa
taaatactat 180gctgactccg tgaagggccg attcaccatc tccagagaca
attccgacaa cacgctgtat 240ctgcaaatga acagcctgag agccgaggac
acggctgtgt attactgtgc gagagatatg 300gactacttcg gtatggacgt
ctggggccaa gggaccacgg tcaccgtctc ctcagccagc 360accaagggcc
cctctgtgtt ccctctggcc ccttccagca agtccacctc tggcggaaca
420gccgctctgg gctgcctcgt gaaggactac ttccccgagc ctgtgaccgt
gtcctggaac 480tctggcgctc tgaccagcgg agtgcacacc ttccctgctg
tgctgcagtc ctccggcctg 540tactccctgt cctccgtcgt gaccgtgcct
tccagctctc tgggcaccca gacctacatc 600tgcaacgtga accacaagcc
ctccaacacc aaggtggaca agaaggtgga acccaagtcc 660tgcgacaaga
cccacacctg tcccccttgt cctgcccctg aactgctggg cggaccttcc
720gtgttcctgt tccccccaaa gcccaaggac accctgatga tctcccggac
ccccgaagtg 780acctgcgtgg tggtggatgt gtcccacgag gaccctgaag
tgaagttcaa ttggtacgtg 840gacggcgtgg aagtgcacaa cgccaagacc
aagcctagag aggaacagta caactccacc 900taccgggtgg tgtccgtgct
gaccgtgctg caccaggatt ggctgaacgg caaagagtac 960aagtgcaagg
tgtccaacaa ggccctgcct gcccccatcg aaaagaccat ctccaaggcc
1020aagggccagc cccgggaacc ccaggtgtac acactgcccc ctagcaggga
cgagctgacc 1080aagaaccagg tgtccctgac ctgtctcgtg aaaggcttct
acccctccga tatcgccgtg 1140gaatgggagt ccaacggcca gcctgagaac
aactacaaga ccaccccccc tgtgctggac 1200tccgacggct cattcttcct
gtacagcaag ctgacagtgg acaagtcccg gtggcagcag 1260ggcaacgtgt
tctcctgctc cgtgatgcac gaggccctgc acaaccacta cacccagaag
1320tccctgtccc tgagccccgg caag 13441426PRTHomo Sapien 142Gln Gly
Ile Arg Asn Asp1 51433PRTHomo Sapien 143Ala Ala Ser11449PRTHomo
Sapien 144Leu Gln His Asn Ser Tyr Pro Arg Thr1 514511PRTHomo Sapien
145Arg Ala Ser Gln Gly Ile Arg Asn Asp Leu Gly1 5 101467PRTHomo
Sapien 146Ala Ala Ser Ser Leu Gln Ser1 51479PRTHomo Sapien 147Leu
Gln His Asn Ser Tyr Pro Arg Thr1 5148107PRTHomo Sapien 148Asp Leu
Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp
Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Gly Ile Arg Asn Asp 20 25
30Leu Gly Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Arg Leu Ile
35 40 45Tyr Ala Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser
Gly 50 55 60Ser Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu
Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Leu Gln His Asn
Ser Tyr Pro Arg 85 90 95Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys
100 105149322DNAHomo Sapien 149gacctccaga tgacccagtc tccatcctcc
ctgtctgcat ctgtaggaga cagagtcacc 60atcacttgcc gggcaagtca gggcattaga
aatgatttag gctggtatca gcagaaacca 120gggaaagccc ctaagcgcct
gatctatgct gcatccagtt tgcaaagtgg ggtcccatca 180aggttcagcg
gcagtggatc tgggacagaa ttcactctca caatcagcag cctgcagcct
240gaagattttg caacttatta ctgtctacag cataatagtt accctcggac
gttcggccaa 300gggaccaagg tggaaatcaa ac 322150214PRTHomo Sapien
150Asp Leu Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Gly Ile Arg Asn
Asp 20 25 30Leu Gly Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Arg
Leu Ile 35 40 45Tyr Ala Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Leu Gln
His Asn Ser Tyr Pro Arg 85 90 95Thr Phe Gly Gln Gly Thr Lys Val Glu
Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro
Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val
Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln
Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155
160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys 210151642DNAHomo
Sapien 151gacctccaga tgacccagtc tccatcctcc ctgtctgcat ctgtaggaga
cagagtcacc 60atcacttgcc gggcaagtca gggcattaga aatgatttag gctggtatca
gcagaaacca 120gggaaagccc ctaagcgcct gatctatgct gcatccagtt
tgcaaagtgg ggtcccatca 180aggttcagcg gcagtggatc tgggacagaa
ttcactctca caatcagcag cctgcagcct 240gaagattttg caacttatta
ctgtctacag cataatagtt accctcggac gttcggccaa 300gggaccaagg
tggaaatcaa acgtacggtg gccgctccct ccgtgttcat cttcccacct
360tccgacgagc agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa
caacttctac 420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc
tgcagtccgg caactcccag 480gaatccgtga ccgagcagga ctccaaggac
agcacctact ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga
gaagcacaag gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc
ccgtgaccaa gtctttcaac cggggcgagt gt 6421529PRTHomo Sapien 152Gly
Gly Ser Ile Ser Ser Ser Asp Trp1 51537PRTHomo Sapien 153Ile Phe His
Ser Gly Asn Thr1 51548PRTHomo Sapien 154Val Arg Asp Gly Ser Gly Ser
Tyr1 51556PRTHomo Sapien 155Ser Ser Asp Trp Trp Ser1 515616PRTHomo
Sapien 156Glu Ile Phe His Ser Gly Asn Thr Asn Tyr Asn Pro Ser Leu
Lys Ser1 5 10 151576PRTHomo Sapien 157Asp Gly Ser Gly Ser Tyr1
5158115PRTHomo Sapien 158Gln Val Gln Leu Gln Glu Ser Gly Pro Gly
Leu Val Lys Pro Ser Gly1 5 10 15Thr Leu Ser Leu Thr Cys Ala Val Ser
Gly Gly Ser Ile Ser Ser Ser 20 25 30Asp Trp Trp Ser Trp Val Arg Gln
Pro Pro Gly Lys Gly Leu Glu Trp 35 40 45Ile Gly Glu Ile Phe His Ser
Gly Asn Thr Asn Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Ile
Ser Val Asp Lys Ser Lys Asn Gln Ile Ser65 70 75 80Leu Arg Leu Asn
Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp
Gly Ser Gly Ser Tyr Trp Gly Gln Gly Thr Leu Val Thr 100 105 110Val
Ser Ser 115159346DNAHomo Sapien 159caggtgcagc tgcaggagtc gggcccagga
ctggtgaagc cttcggggac cctgtccctc 60acctgcgctg tctctggtgg ctccatcagc
agtagtgact ggtggagttg ggtccgccag 120cccccaggga aggggctgga
gtggattggg gaaatctttc atagtgggaa caccaactac 180aacccgtccc
tcaagagtcg agtcaccata tcagtagaca agtccaagaa ccagatctcc
240ctgaggctga actctgtgac cgccgcggac acggccgtgt attactgtgt
gagagatggt 300tcggggagtt actggggcca gggaaccctg gtcaccgtct cctcag
346160445PRTHomo Sapien 160Gln Val Gln Leu Gln Glu Ser Gly Pro Gly
Leu Val Lys Pro Ser Gly1 5 10 15Thr Leu Ser Leu Thr Cys Ala Val Ser
Gly Gly Ser Ile Ser Ser Ser 20 25 30Asp Trp Trp Ser Trp Val Arg Gln
Pro Pro Gly Lys Gly Leu Glu Trp 35 40 45Ile Gly Glu Ile Phe His Ser
Gly Asn Thr Asn Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Ile
Ser Val Asp Lys Ser Lys Asn Gln Ile Ser65 70 75 80Leu Arg Leu Asn
Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Val Arg Asp
Gly Ser Gly Ser Tyr Trp Gly Gln Gly Thr Leu Val Thr 100 105 110Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro 115 120
125Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val
130 135 140Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser
Gly Ala145 150 155 160Leu Thr Ser Gly Val His Thr Phe Pro Ala Val
Leu Gln Ser Ser Gly 165 170 175Leu Tyr Ser Leu Ser Ser Val Val Thr
Val Pro Ser Ser Ser Leu Gly 180 185 190Thr Gln Thr Tyr Ile Cys Asn
Val Asn His Lys Pro Ser Asn Thr Lys 195 200 205Val Asp Lys Lys Val
Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys 210 215 220Pro Pro Cys
Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu225 230 235
240Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu
245 250 255Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu
Val Lys 260 265 270Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn
Ala Lys Thr Lys 275 280 285Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val Ser Val Leu 290 295 300Thr Val Leu His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr Lys Cys Lys305 310 315 320Val Ser Asn Lys Ala
Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys 325 330 335Ala Lys Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser 340 345 350Arg
Asp Glu Leu Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys 355 360 365Gly Phe Tyr Pro Ser Asp Ile Ala Val
Glu Trp Glu Ser Asn Gly Gln 370 375 380Pro Glu Asn Asn Tyr Lys Thr
Thr Pro Pro Val Leu Asp Ser Asp Gly385 390 395 400Ser Phe Phe Leu
Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln 405 410 415Gln Gly
Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn 420 425
430His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 435 440
4451611335DNAHomo Sapien 161caggtgcagc tgcaggagtc gggcccagga
ctggtgaagc cttcggggac cctgtccctc 60acctgcgctg tctctggtgg ctccatcagc
agtagtgact ggtggagttg ggtccgccag 120cccccaggga aggggctgga
gtggattggg gaaatctttc atagtgggaa caccaactac 180aacccgtccc
tcaagagtcg agtcaccata tcagtagaca agtccaagaa ccagatctcc
240ctgaggctga actctgtgac cgccgcggac acggccgtgt attactgtgt
gagagatggt 300tcggggagtt actggggcca gggaaccctg gtcaccgtct
cctcagccag caccaagggc 360ccctctgtgt tccctctggc cccttccagc
aagtccacct ctggcggaac agccgctctg 420ggctgcctcg tgaaggacta
cttccccgag cctgtgaccg tgtcctggaa ctctggcgct 480ctgaccagcg
gagtgcacac cttccctgct gtgctgcagt cctccggcct gtactccctg
540tcctccgtcg tgaccgtgcc ttccagctct ctgggcaccc agacctacat
ctgcaacgtg 600aaccacaagc cctccaacac caaggtggac aagaaggtgg
aacccaagtc ctgcgacaag 660acccacacct gtcccccttg tcctgcccct
gaactgctgg gcggaccttc cgtgttcctg 720ttccccccaa agcccaagga
caccctgatg atctcccgga cccccgaagt gacctgcgtg 780gtggtggatg
tgtcccacga ggaccctgaa gtgaagttca attggtacgt ggacggcgtg
840gaagtgcaca acgccaagac caagcctaga gaggaacagt acaactccac
ctaccgggtg 900gtgtccgtgc tgaccgtgct gcaccaggat tggctgaacg
gcaaagagta caagtgcaag 960gtgtccaaca aggccctgcc tgcccccatc
gaaaagacca tctccaaggc caagggccag 1020ccccgggaac cccaggtgta
cacactgccc cctagcaggg acgagctgac caagaaccag 1080gtgtccctga
cctgtctcgt gaaaggcttc tacccctccg atatcgccgt ggaatgggag
1140tccaacggcc agcctgagaa caactacaag accacccccc ctgtgctgga
ctccgacggc 1200tcattcttcc tgtacagcaa gctgacagtg gacaagtccc
ggtggcagca gggcaacgtg 1260ttctcctgct ccgtgatgca cgaggccctg
cacaaccact acacccagaa gtccctgtcc 1320ctgagccccg gcaag
133516212PRTHomo Sapien 162Gln Ser Val Leu Tyr Ser Ser Asn Asn Lys
Asn Tyr1 5 101633PRTHomo Sapien 163Trp Ala Ser11648PRTHomo Sapien
164Gln Gln Tyr Tyr Ser Thr Arg Ser1 516517PRTHomo Sapien 165Lys Ser
Ser Gln Ser Val Leu Tyr Ser Ser Asn Asn Lys Asn Tyr Leu1 5 10
15Ala1667PRTHomo Sapien 166Trp Ala Ser Thr Arg Glu Ser1
51678PRTHomo Sapien 167Gln Gln Tyr Tyr Ser Thr Arg Ser1
5168112PRTHomo Sapien 168Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser
Thr Arg Ser Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100 105
110169337DNAHomo Sapien 169gacatcgtga tgacccagtc tccagactcc
ctggctgtgt ctctgggcga gagggccacc 60atcaactgca agtccagcca gagtgtttta
tacagctcca acaataagaa ctacttagct 120tggtaccagc agaaaccagg
acagcctcct aaactgctca tttactgggc atctacccgg 180gaatccgggg
tccctgaccg attcagtggc agcgggtctg ggacagattt cactctcacc
240atcagcagcc tgcaggctga agatgtggca gtttattact gtcagcaata
ttatagtact 300cgcagttttg gccaggggac caagctggag atcaaac
337170219PRTHomo Sapien 170Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Asn Lys Asn Tyr Leu Ala
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60Pro Asp Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser
Thr Arg Ser Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100 105 110Arg
Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu 115 120
125Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
130 135 140Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln145 150 155 160Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser 165 170 175Thr Tyr Ser Leu Ser Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu 180 185 190Lys His Lys Val Tyr Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser 195 200 205Pro Val Thr Lys Ser
Phe Asn Arg Gly Glu Cys 210 215171657DNAHomo Sapien 171gacatcgtga
tgacccagtc tccagactcc ctggctgtgt ctctgggcga gagggccacc 60atcaactgca
agtccagcca gagtgtttta tacagctcca acaataagaa ctacttagct
120tggtaccagc agaaaccagg acagcctcct aaactgctca tttactgggc
atctacccgg 180gaatccgggg tccctgaccg attcagtggc agcgggtctg
ggacagattt cactctcacc 240atcagcagcc tgcaggctga agatgtggca
gtttattact gtcagcaata ttatagtact 300cgcagttttg gccaggggac
caagctggag atcaaacgta cggtggccgc tccctccgtg 360ttcatcttcc
caccttccga cgagcagctg aagtccggca ccgcttctgt cgtgtgcctg
420ctgaacaact tctacccccg cgaggccaag gtgcagtgga aggtggacaa
cgccctgcag 480tccggcaact cccaggaatc cgtgaccgag caggactcca
aggacagcac ctactccctg 540tcctccaccc tgaccctgtc caaggccgac
tacgagaagc acaaggtgta cgcctgcgaa 600gtgacccacc agggcctgtc
tagccccgtg accaagtctt tcaaccgggg cgagtgt 65717210PRTHomo Sapien
172Gly Gly Ser Ile Ser Ser Ser Ser Tyr Tyr1 5 101737PRTHomo Sapien
173Ile Tyr Ser Thr Gly Tyr Thr1 517413PRTHomo Sapien 174Ala Ile Ser
Thr Ala Ala Gly Pro Glu Tyr Phe His Arg1 5 101757PRTHomo Sapien
175Ser Ser Ser Tyr Tyr Cys Gly1 517616PRTHomo Sapien 176Ser Ile Tyr
Ser Thr Gly Tyr Thr Tyr Tyr Asn Pro Ser Leu Lys Ser1 5 10
1517711PRTHomo Sapien 177Ser Thr Ala Ala Gly Pro Glu Tyr Phe His
Arg1 5 10178120PRTHomo Sapien 178Gln Leu Gln Glu Ser Gly Pro Gly
Leu Val Lys Pro Ser Glu Thr Leu1 5 10 15Ser Leu Thr Cys Thr Val Ser
Gly Gly Ser Ile Ser Ser Ser Ser Tyr 20 25 30Tyr Cys Gly Trp Ile Arg
Gln Pro Pro Gly Lys Gly Leu Asp Trp Ile 35 40 45Gly Ser Ile Tyr Ser
Thr Gly Tyr Thr Tyr Tyr Asn Pro Ser Leu Lys 50 55 60Ser Arg Val Thr
Ile Ser Ile Asp Thr Ser Lys Asn Gln Phe Ser Cys65 70 75 80Leu Ile
Leu Thr Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala
Ile Ser Thr Ala Ala Gly Pro Glu Tyr Phe His Arg Trp Gly Gln 100 105
110Gly Thr Leu Val Thr Val Ser Ser 115 120179361DNAHomo Sapien
179cagctgcagg agtcgggccc aggcctggtg aagccttcgg agaccctgtc
cctcacctgc 60actgtctctg gtggctccat cagcagtagt agttattact gcggctggat
ccgccagccc 120cctgggaagg ggctggactg gattgggagt atctattcta
ctgggtacac ctactacaac 180ccgtccctca agagtcgagt caccatttcc
atagacacgt ccaagaacca gttctcatgc 240ctgatactga cctctgtgac
cgccgcagac acggctgtgt attactgtgc gataagtaca 300gcagctggcc
ctgaatactt ccatcgctgg ggccagggca ccctggtcac cgtctcctca 360g
361180450PRTHomo Sapien 180Gln Leu Gln Glu Ser Gly Pro Gly Leu Val
Lys Pro Ser Glu Thr Leu1 5 10 15Ser Leu Thr Cys Thr Val Ser Gly Gly
Ser Ile Ser Ser Ser Ser Tyr 20 25 30Tyr Cys Gly Trp Ile Arg Gln Pro
Pro Gly Lys Gly Leu Asp Trp Ile 35 40 45Gly Ser Ile Tyr Ser Thr Gly
Tyr Thr Tyr Tyr Asn Pro Ser Leu Lys 50 55 60Ser Arg Val Thr Ile Ser
Ile Asp Thr Ser Lys Asn Gln Phe Ser Cys65 70 75 80Leu Ile Leu Thr
Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Ile Ser
Thr Ala Ala Gly Pro Glu Tyr Phe His Arg Trp Gly Gln 100 105 110Gly
Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val 115 120
125Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala
130 135 140Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr
Val Ser145 150 155 160Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His
Thr Phe Pro Ala Val 165 170 175Leu Gln Ser Ser Gly Leu Tyr Ser Leu
Ser Ser Val Val Thr Val Pro 180 185 190Ser Ser Ser Leu Gly Thr Gln
Thr Tyr Ile Cys Asn Val Asn His Lys 195 200 205Pro Ser Asn Thr Lys
Val Asp Lys Lys Val Glu Pro Lys Ser Cys Asp 210 215 220Lys Thr His
Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly Gly225 230 235
240Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
245 250 255Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser
His Glu 260 265 270Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
Val Glu Val His 275 280 285Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn Ser Thr Tyr Arg 290 295 300Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu Asn Gly Lys305 310 315 320Glu Tyr Lys Cys Lys
Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu 325 330 335Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr 340 345 350Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu 355 360
365Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp
370 375 380Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro
Pro Val385 390 395 400Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys Leu Thr Val Asp 405 410 415Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys Ser Val Met His 420 425 430Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu Ser Leu Ser Pro 435 440 445Gly Lys
4501811350DNAHomo Sapien 181cagctgcagg agtcgggccc aggcctggtg
aagccttcgg agaccctgtc cctcacctgc 60actgtctctg gtggctccat cagcagtagt
agttattact gcggctggat ccgccagccc 120cctgggaagg ggctggactg
gattgggagt atctattcta ctgggtacac ctactacaac 180ccgtccctca
agagtcgagt caccatttcc atagacacgt ccaagaacca gttctcatgc
240ctgatactga cctctgtgac cgccgcagac acggctgtgt attactgtgc
gataagtaca 300gcagctggcc ctgaatactt ccatcgctgg ggccagggca
ccctggtcac cgtctcctca 360gccagcacca agggcccctc tgtgttccct
ctggcccctt ccagcaagtc cacctctggc 420ggaacagccg ctctgggctg
cctcgtgaag gactacttcc ccgagcctgt gaccgtgtcc 480tggaactctg
gcgctctgac cagcggagtg cacaccttcc ctgctgtgct gcagtcctcc
540ggcctgtact ccctgtcctc cgtcgtgacc gtgccttcca gctctctggg
cacccagacc 600tacatctgca acgtgaacca caagccctcc aacaccaagg
tggacaagaa ggtggaaccc 660aagtcctgcg acaagaccca cacctgtccc
ccttgtcctg cccctgaact gctgggcgga 720ccttccgtgt tcctgttccc
cccaaagccc aaggacaccc tgatgatctc ccggaccccc 780gaagtgacct
gcgtggtggt ggatgtgtcc cacgaggacc ctgaagtgaa gttcaattgg
840tacgtggacg gcgtggaagt gcacaacgcc aagaccaagc ctagagagga
acagtacaac 900tccacctacc gggtggtgtc cgtgctgacc gtgctgcacc
aggattggct gaacggcaaa 960gagtacaagt gcaaggtgtc caacaaggcc
ctgcctgccc ccatcgaaaa gaccatctcc 1020aaggccaagg gccagccccg
ggaaccccag gtgtacacac tgccccctag cagggacgag 1080ctgaccaaga
accaggtgtc cctgacctgt ctcgtgaaag gcttctaccc ctccgatatc
1140gccgtggaat gggagtccaa cggccagcct gagaacaact acaagaccac
cccccctgtg 1200ctggactccg acggctcatt cttcctgtac agcaagctga
cagtggacaa gtcccggtgg 1260cagcagggca acgtgttctc ctgctccgtg
atgcacgagg ccctgcacaa ccactacacc 1320cagaagtccc tgtccctgag
ccccggcaag 135018212PRTHomo Sapien 182Gln Ser Val Leu Tyr Ser Ser
Asn Ser Lys Asn Phe1 5 101833PRTHomo Sapien 183Trp Ala
Ser11849PRTHomo Sapien 184Gln Gln Tyr Tyr Ser Thr Pro Arg Thr1
518517PRTHomo Sapien 185Lys Ser Ser Gln Ser Val Leu Tyr Ser Ser Asn
Ser Lys Asn Phe Leu1 5 10 15Ala1867PRTHomo Sapien 186Trp Ala Ser
Thr Arg Gly Ser1 51879PRTHomo Sapien 187Gln Gln Tyr Tyr Ser Thr Pro
Arg Thr1 5188113PRTHomo Sapien 188Asp Ile Val Met Thr Gln Ser Pro
Asp Ser Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Ser Lys Asn Phe
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Phe
Ile Tyr Trp Ala Ser Thr Arg Gly Ser Gly Val 50 55 60Pro Asp Arg Ile
Ser Gly Ser Gly Ser Gly Thr Asp Phe Asn Leu Thr65 70 75 80Ile Ser
Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr
Tyr Ser Thr Pro Arg Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105
110Lys189340DNAHomo Sapien 189gacatcgtga tgacccagtc tccagactcc
ctggctgtgt ctctgggcga gagggccacc 60atcaactgca agtccagcca gagtgtttta
tacagctcca acagtaagaa cttcttagct 120tggtaccagc agaaaccggg
acagcctcct aagctgttca tttactgggc atctacccgg 180ggatccgggg
tccctgaccg aatcagtggc agcgggtctg ggacagattt caatctcacc
240atcagcagcc tgcaggctga agatgtggca gtttattact gtcaacaata
ttatagtact 300cctcggacgt tcggccaagg gaccaaggtg gagatcaaac
340190220PRTHomo Sapien 190Asp Ile Val Met Thr Gln Ser Pro Asp Ser
Leu Ala Val Ser Leu Gly1 5 10 15Glu Arg Ala Thr Ile Asn Cys Lys Ser
Ser Gln Ser Val Leu Tyr Ser 20 25 30Ser Asn Ser Lys Asn Phe Leu Ala
Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45Pro Pro Lys Leu Phe Ile Tyr
Trp Ala Ser Thr Arg Gly Ser Gly Val 50 55 60Pro Asp Arg Ile Ser Gly
Ser Gly Ser Gly Thr Asp Phe Asn Leu Thr65 70 75 80Ile Ser Ser Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95Tyr Tyr Ser
Thr Pro Arg Thr Phe Gly Gln Gly Thr Lys Val Glu Ile 100 105 110Lys
Arg Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp 115 120
125Glu Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn
130 135 140Phe Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn
Ala Leu145 150 155 160Gln Ser Gly Asn Ser Gln Glu Ser Val Thr Glu
Gln Asp Ser Lys Asp 165 170 175Ser Thr Tyr Ser Leu Ser Ser Thr Leu
Thr Leu Ser Lys Ala Asp Tyr 180 185 190Glu Lys His Lys Val Tyr Ala
Cys Glu Val Thr His Gln Gly Leu Ser 195 200 205Ser Pro Val Thr Lys
Ser Phe Asn Arg Gly Glu Cys 210 215 220191660DNAHomo Sapien
191gacatcgtga tgacccagtc tccagactcc ctggctgtgt ctctgggcga
gagggccacc 60atcaactgca agtccagcca gagtgtttta tacagctcca acagtaagaa
cttcttagct 120tggtaccagc agaaaccggg acagcctcct aagctgttca
tttactgggc atctacccgg 180ggatccgggg tccctgaccg aatcagtggc
agcgggtctg ggacagattt caatctcacc 240atcagcagcc tgcaggctga
agatgtggca gtttattact gtcaacaata ttatagtact 300cctcggacgt
tcggccaagg gaccaaggtg gagatcaaac gtacggtggc cgctccctcc
360gtgttcatct tcccaccttc cgacgagcag ctgaagtccg gcaccgcttc
tgtcgtgtgc 420ctgctgaaca acttctaccc ccgcgaggcc aaggtgcagt
ggaaggtgga caacgccctg 480cagtccggca actcccagga atccgtgacc
gagcaggact ccaaggacag cacctactcc 540ctgtcctcca ccctgaccct
gtccaaggcc gactacgaga agcacaaggt gtacgcctgc 600gaagtgaccc
accagggcct gtctagcccc gtgaccaagt ctttcaaccg gggcgagtgt
660192981DNAHomo Sapien 192gcttccacca agggcccatc cgtcttcccc
ctggcgccct gctccaggag cacctccgag 60agcacagccg ccctgggctg cctggtcaag
gactacttcc ccgaaccggt gacggtgtcg 120tggaactcag gcgccctgac
cagcggcgtg cacaccttcc cggctgtcct acagtcctca 180ggactctact
ccctcagcag cgtggtgacc gtgccctcca
gcagcttggg cacgaagacc 240tacacctgca acgtagatca caagcccagc
aacaccaagg tggacaagag agttgagtcc 300aaatatggtc ccccatgccc
atcatgccca gcacctgagt tcctgggggg accatcagtc 360ttcctgttcc
ccccaaaacc caaggacact ctcatgatct cccggacccc tgaggtcacg
420tgcgtggtgg tggacgtgag ccaggaagac cccgaggtcc agttcaactg
gtacgtggat 480ggcgtggagg tgcataatgc caagacaaag ccgcgggagg
agcagttcaa cagcacgtac 540cgtgtggtca gcgtcctcac cgtcctgcac
caggactggc tgaacggcaa ggagtacaag 600tgcaaggtct ccaacaaagg
cctcccgtcc tccatcgaga aaaccatctc caaagccaaa 660gggcagcccc
gagagccaca ggtgtacacc ctgcccccat cccaggagga gatgaccaag
720aaccaggtca gcctgacctg cctggtcaaa ggcttctacc ccagcgacat
cgccgtggag 780tgggagagca atgggcagcc ggagaacaac tacaagacca
cgcctcccgt gctggactcc 840gacggctcct tcttcctcta cagcaggcta
accgtggaca agagcaggtg gcaggagggg 900aatgtcttct catgctccgt
gatgcatgag gctctgcaca accactacac acagaagagc 960ctctccctgt
ctctgggtaa a 981193327PRTHomo Sapien 193Ala Ser Thr Lys Gly Pro Ser
Val Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr
Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val
Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr
Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser
Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr
Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90
95Arg Val Glu Ser Lys Tyr Gly Pro Pro Cys Pro Ser Cys Pro Ala Pro
100 105 110Glu Phe Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys 115 120 125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val 130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln
Phe Asn Trp Tyr Val Asp145 150 155 160Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn
Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro
Ser Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215
220Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr
Lys225 230 235 240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp 245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys
Ser Arg Trp Gln Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu
Ser Leu Ser Leu Gly Lys 325194981DNAHomo Sapien 194gcttccacca
agggcccatc cgtcttcccc ctggcgccct gctccaggag cacctccgag 60agcacagccg
ccctgggctg cctggtcaag gactacttcc ccgaaccggt gacggtgtcg
120tggaactcag gcgccctgac cagcggcgtg cacaccttcc cggctgtcct
acagtcctca 180ggactctact ccctcagcag cgtggtgacc gtgccctcca
gcagcttggg cacgaagacc 240tacacctgca acgtagatca caagcccagc
aacaccaagg tggacaagag agttgagtcc 300aaatatggtc ccccgtgccc
atcatgccca gcacctgagt tcctgggggg accatcagtc 360ttcctgttcc
ccccaaaacc caaggacact ctcatgatct cccggacccc tgaggtcacg
420tgcgtggtgg tggacgtgag ccaggaagac cccgaggtcc agttcaactg
gtacgtggat 480ggcgtggagg tgcataatgc caagacaaag ccgcgggagg
agcagttcaa cagcacgtac 540cgtgtggtca gcgtcctcac cgtcgtgcac
caggactggc tgaacggcaa ggagtacaag 600tgcaaggtct ccaacaaagg
cctcccgtcc tccatcgaga aaaccatctc caaagccaaa 660gggcagcccc
gagagccaca ggtgtacacc ctgcccccat cccaggagga gatgaccaag
720aaccaggtca gcctgacctg cctggtcaaa ggcttctacc ccagcgacat
cgccgtggag 780tgggagagca atgggcagcc ggagaacaac tacaagacca
cgcctcccgt gctggactcc 840gacggctcct tcttcctcta cagcaggcta
accgtggaca agagcaggtg gcaggagggg 900aatgtcttct catgctccgt
gatgcatgag gctctgcaca accactacac gcagaagagc 960ctctccctgt
ctctgggtaa a 981195327PRTHomo Sapien 195Ala Ser Thr Lys Gly Pro Ser
Val Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr
Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val
Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr
Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser
Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr
Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90
95Arg Val Glu Ser Lys Tyr Gly Pro Pro Cys Pro Ser Cys Pro Ala Pro
100 105 110Glu Phe Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys 115 120 125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val 130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln
Phe Asn Trp Tyr Val Asp145 150 155 160Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val Val His Gln Asp 180 185 190Trp Leu Asn
Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro
Ser Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215
220Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr
Lys225 230 235 240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp 245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys
Ser Arg Trp Gln Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu
Ser Leu Ser Leu Gly Lys 325196981DNAHomo Sapien 196gcttccacca
agggcccatc cgtcttcccc ctggcgccct gctccaggag cacctccgag 60agcacagccg
ccctgggctg cctggtcaag gactacttcc ccgaaccggt gacggtgtcg
120tggaactcag gcgccctgac cagcggcgtg cacaccttcc cggctgtcct
acagtcctca 180ggactctact ccctcagcag cgtggtgacc gtgccctcca
gcagcttggg cacgaagacc 240tacacctgca acgtagatca caagcccagc
aacaccaagg tggacaagag agttgagtcc 300aaatatggtc ccccatgccc
atcatgccca gcacctgagt tcctgggggg accatcagtc 360ttcctgttcc
ccccaaaacc caaggacact ctcatgatct cccggacccc tgaggtcacg
420tgcgtggtgg tggacgtgag ccaggaagac cccgaggtcc agttcaactg
gtacgtggat 480ggcgtggagg tgcataatgc caagacaaag ccgcgggagg
agcagttcaa cagcacgtac 540cgtgtggtca gcgtcctcac cgtcctgcac
caggactggc tgaacggcaa ggagtacaag 600tgcaaggtct ccaacaaagg
cctcccgtcc tccatcgaga aaaccatctc caaagccaaa 660gggcagcccc
gagagccaca ggtgtacacc ctgcccccat cccaggagga gatgaccaag
720aaccaggtca gcctgacctg cctggtcaaa ggcttctacc ccagcgacat
cgccgtggag 780tgggagagca atgggcagcc ggagaacaac tacaagacca
cgcctcccgt gctggactcc 840gacggctcct tcttcctcta cagcaagctc
accgtggaca agagcaggtg gcaggagggg 900aacgtcttct catgctccgt
gatgcatgag gctctgcaca accactacac gcagaagagc 960ctctccctgt
ctctgggtaa a 981197327PRTHomo Sapien 197Ala Ser Thr Lys Gly Pro Ser
Val Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr
Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val
Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr
Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser
Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr
Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90
95Arg Val Glu Ser Lys Tyr Gly Pro Pro Cys Pro Ser Cys Pro Ala Pro
100 105 110Glu Phe Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys 115 120 125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val 130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln
Phe Asn Trp Tyr Val Asp145 150 155 160Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn
Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro
Ser Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215
220Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr
Lys225 230 235 240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp 245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser 275 280 285Lys Leu Thr Val Asp Lys
Ser Arg Trp Gln Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu
Ser Leu Ser Leu Gly Lys 325198981DNAHomo Sapien 198gcctccacca
agggcccatc cgtcttcccc ctggcgccct gctccaggag cacctccgag 60agcacggccg
ccctgggctg cctggtcaag gactacttcc ccgaaccagt gacggtgtcg
120tggaactcag gcgccctgac cagcggcgtg cacaccttcc cggctgtcct
acagtcctca 180ggactctact ccctcagcag cgtggtgacc gtgccctcca
gcagcttggg cacgaagacc 240tacacctgca acgtagatca caagcccagc
aacaccaagg tggacaagag agttgagtcc 300aaatatggtc ccccatgccc
accatgccca gcgcctgaat ttgagggggg accatcagtc 360ttcctgttcc
ccccaaaacc caaggacact ctcatgatct cccggacccc tgaggtcacg
420tgcgtggtgg tggacgtgag ccaggaagac cccgaggtcc agttcaactg
gtacgtggat 480ggcgtggagg tgcataatgc caagacaaag ccgcgggagg
agcagttcaa cagcacgtac 540cgtgtggtca gcgtcctcac cgtcctgcac
caggactggc tgaacggcaa ggagtacaag 600tgcaaggtct ccaacaaagg
cctcccgtca tcgatcgaga aaaccatctc caaagccaaa 660gggcagcccc
gagagccaca ggtgtacacc ctgcccccat cccaggagga gatgaccaag
720aaccaggtca gcctgacctg cctggtcaaa ggcttctacc ccagcgacat
cgccgtggag 780tgggagagca atgggcagcc ggagaacaac tacaagacca
cgcctcccgt gctggactcc 840gacggatcct tcttcctcta cagcaggcta
accgtggaca agagcaggtg gcaggagggg 900aatgtcttct catgctccgt
gatgcatgag gctctgcaca accactacac acagaagagc 960ctctccctgt
ctctgggtaa a 981199327PRTHomo Sapien 199Ala Ser Thr Lys Gly Pro Ser
Val Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr
Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val
Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr
Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser
Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr
Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90
95Arg Val Glu Ser Lys Tyr Gly Pro Pro Cys Pro Pro Cys Pro Ala Pro
100 105 110Glu Phe Glu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys 115 120 125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val 130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln
Phe Asn Trp Tyr Val Asp145 150 155 160Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn
Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro
Ser Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215
220Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr
Lys225 230 235 240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp 245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys
Ser Arg Trp Gln Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu
Ser Leu Ser Leu Gly Lys 325200981DNAHomo Sapien 200gcctccacca
agggacctag cgtgttccct ctcgccccct gttccaggtc cacaagcgag 60tccaccgctg
ccctcggctg tctggtgaaa gactactttc ccgagcccgt gaccgtctcc
120tggaatagcg gagccctgac ctccggcgtg cacacatttc ccgccgtgct
gcagagcagc 180ggactgtata gcctgagcag cgtggtgacc gtgcccagct
ccagcctcgg caccaaaacc 240tacacctgca acgtggacca caagccctcc
aacaccaagg tggacaagcg ggtggagagc 300aagtacggcc ccccttgccc
tccttgtcct gcccctgagt tcgagggagg accctccgtg 360ttcctgtttc
cccccaaacc caaggacacc ctgatgatct cccggacacc cgaggtgacc
420tgtgtggtcg tggacgtcag ccaggaggac cccgaggtgc agttcaactg
gtatgtggac 480ggcgtggagg tgcacaatgc caaaaccaag cccagggagg
agcagttcaa ttccacctac 540agggtggtga gcgtgctgac cgtcctgcat
caggattggc tgaacggcaa ggagtacaag 600tgcaaggtgt ccaacaaggg
actgcccagc tccatcgaga agaccatcag caaggctaag 660ggccagccga
gggagcccca ggtgtatacc ctgcctccta gccaggaaga gatgaccaag
720aaccaagtgt ccctgacctg cctggtgaag ggattctacc cctccgacat
cgccgtggag 780tgggagagca atggccagcc cgagaacaac tacaaaacaa
cccctcccgt gctcgatagc 840gacggcagct tctttctcta cagccggctg
acagtggaca agagcaggtg gcaggagggc 900aacgtgttct cctgttccgt
gatgcacgag gccctgcaca atcactacac ccagaagagc 960ctctccctgt
ccctgggcaa g 981201981DNAHomo Sapien 201gccagcacca agggcccttc
cgtgttcccc ctggcccctt gcagcaggag cacctccgaa 60tccacagctg ccctgggctg
tctggtgaag gactactttc ccgagcccgt gaccgtgagc 120tggaacagcg
gcgctctgac atccggcgtc cacacctttc ctgccgtcct gcagtcctcc
180ggcctctact ccctgtcctc cgtggtgacc gtgcctagct cctccctcgg
caccaagacc 240tacacctgta acgtggacca caaaccctcc aacaccaagg
tggacaaacg ggtcgagagc 300aagtacggcc ctccctgccc tccttgtcct
gcccccgagt tcgaaggcgg acccagcgtg 360ttcctgttcc ctcctaagcc
caaggacacc ctcatgatca gccggacacc cgaggtgacc 420tgcgtggtgg
tggatgtgag ccaggaggac cctgaggtcc agttcaactg gtatgtggat
480ggcgtggagg tgcacaacgc caagacaaag ccccgggaag agcagttcaa
ctccacctac 540agggtggtca gcgtgctgac cgtgctgcat caggactggc
tgaacggcaa ggagtacaag 600tgcaaggtca gcaataaggg actgcccagc
agcatcgaga agaccatctc caaggctaaa 660ggccagcccc gggaacctca
ggtgtacacc ctgcctccca gccaggagga gatgaccaag 720aaccaggtga
gcctgacctg cctggtgaag ggattctacc cttccgacat cgccgtggag
780tgggagtcca acggccagcc cgagaacaat tataagacca cccctcccgt
cctcgacagc 840gacggatcct tctttctgta ctccaggctg accgtggata
agtccaggtg gcaggaaggc 900aacgtgttca gctgctccgt gatgcacgag
gccctgcaca atcactacac ccagaagtcc 960ctgagcctgt ccctgggaaa g
981202981DNAHomo Sapien 202gcctccacca agggcccatc cgtcttcccc
ctggcgccct gctccaggag cacctccgag 60agcacggccg ccctgggctg cctggtcaag
gactacttcc ccgaaccagt gacggtgtcg 120tggaactcag gcgccctgac
cagcggcgtg cacaccttcc cggctgtcct acagtcctca 180ggactctact
ccctcagcag cgtggtgacc gtgccctcca gcagcttggg cacgaagacc
240tacacctgca acgtagatca caagcccagc aacaccaagg tggacaagag
agttgagtcc 300aaatatggtc ccccatgccc accatgccca gcgcctccag
ttgcgggggg accatcagtc 360ttcctgttcc ccccaaaacc caaggacact
ctcatgatct cccggacccc tgaggtcacg 420tgcgtggtgg tggacgtgag
ccaggaagac cccgaggtcc agttcaactg gtacgtggat 480ggcgtggagg
tgcataatgc caagacaaag ccgcgggagg agcagttcaa cagcacgtac
540cgtgtggtca gcgtcctcac cgtcctgcac caggactggc tgaacggcaa
ggagtacaag 600tgcaaggtct ccaacaaagg cctcccgtca tcgatcgaga
aaaccatctc caaagccaaa 660gggcagcccc gagagccaca ggtgtacacc
ctgcccccat cccaggagga gatgaccaag 720aaccaggtca gcctgacctg
cctggtcaaa ggcttctacc ccagcgacat cgccgtggag 780tgggagagca
atgggcagcc ggagaacaac tacaagacca cgcctcccgt gctggactcc
840gacggatcct tcttcctcta cagcaggcta accgtggaca agagcaggtg
gcaggagggg 900aatgtcttct catgctccgt gatgcatgag gctctgcaca
accactacac acagaagagc 960ctctccctgt ctctgggtaa a
981203327PRTHomo Sapien 203Ala Ser Thr Lys Gly Pro Ser Val Phe Pro
Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu
Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser
Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala
Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr
Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr Thr Cys Asn
Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Arg Val Glu
Ser Lys Tyr Gly Pro Pro Cys Pro Pro Cys Pro Ala Pro 100 105 110Pro
Val Ala Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys 115 120
125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val
130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr
Val Asp145 150 155 160Gly Val Glu Val His Asn Ala Lys Thr Lys Pro
Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg Val Val Ser Val
Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn Gly Lys Glu Tyr
Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro Ser Ser Ile Glu
Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215 220Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys225 230 235
240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys Ser Arg Trp Gln
Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu Ser Leu Ser Leu
Gly Lys 325204990DNAHomo Sapien 204gcctccacca agggcccatc ggtcttcccc
ctggcaccct cctccaagag cacctctggg 60ggcacagcgg ccctgggctg cctggtcaag
gactacttcc ccgaaccggt gacggtgtcg 120tggaactcag gcgccctgac
cagcggcgtg cacaccttcc cggctgtcct acagtcctca 180ggactctact
ccctcagcag cgtggtgacc gtgccctcca gcagcttggg cacccagacc
240tacatctgca acgtgaatca caagcccagc aacaccaagg tggacaagaa
agtggagccc 300aaatcttgtg acaaaactca cacatgccca ccgtgcccag
cacctgaact cgcgggggca 360ccgtcagtct tcctcttccc cccaaaaccc
aaggacaccc tcatgatctc ccggacccct 420gaggtcacat gcgtggtggt
ggacgtgagc cacgaagacc ctgaggtcaa gttcaactgg 480tacgtggacg
gcgtggaggt gcataatgcc aagacaaagc cgcgggagga gcagtacaac
540agcacgtacc gtgtggtcag cgtcctcacc gtcctgcacc aggactggct
gaatggcaag 600gagtacaagt gcaaggtctc caacaaagcc ctcccagccc
ccatcgagaa aaccatctcc 660aaagccaaag ggcagccccg agaaccacag
gtgtacaccc tgcccccatc ccgggatgag 720ctgaccaaga accaggtcag
cctgacctgc ctggtcaaag gcttctatcc cagcgacatc 780gccgtggagt
gggagagcaa tgggcagccg gagaacaact acaagaccac gcctcccgtg
840ctggactccg acggctcctt cttcctctac agcaagctca ccgtggacaa
gagcaggtgg 900cagcagggga acgtcttctc atgctccgtg atgcatgagg
ctctgcacaa ccactacacg 960cagaagagcc tctccctgtc tccgggtaaa
990205330PRTHomo Sapien 205Ala Ser Thr Lys Gly Pro Ser Val Phe Pro
Leu Ala Pro Ser Ser Lys1 5 10 15Ser Thr Ser Gly Gly Thr Ala Ala Leu
Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser
Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala
Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr
Val Pro Ser Ser Ser Leu Gly Thr Gln Thr65 70 75 80Tyr Ile Cys Asn
Val Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Lys Val Glu
Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys 100 105 110Pro
Ala Pro Glu Leu Ala Gly Ala Pro Ser Val Phe Leu Phe Pro Pro 115 120
125Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys
130 135 140Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe
Asn Trp145 150 155 160Tyr Val Asp Gly Val Glu Val His Asn Ala Lys
Thr Lys Pro Arg Glu 165 170 175Glu Gln Tyr Asn Ser Thr Tyr Arg Val
Val Ser Val Leu Thr Val Leu 180 185 190His Gln Asp Trp Leu Asn Gly
Lys Glu Tyr Lys Cys Lys Val Ser Asn 195 200 205Lys Ala Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly 210 215 220Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu225 230 235
240Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr
245 250 255Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn 260 265 270Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp
Gly Ser Phe Phe 275 280 285Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp Gln Gln Gly Asn 290 295 300Val Phe Ser Cys Ser Val Met His
Glu Ala Leu His Asn His Tyr Thr305 310 315 320Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 325 330206321DNAHomo Sapien 206cgtacggtgg
ccgctccctc cgtgttcatc ttcccacctt ccgacgagca gctgaagtcc 60ggcaccgctt
ctgtcgtgtg cctgctgaac aacttctacc cccgcgaggc caaggtgcag
120tggaaggtgg acaacgccct gcagtccggc aactcccagg aatccgtgac
cgagcaggac 180tccaaggaca gcacctactc cctgtcctcc accctgaccc
tgtccaaggc cgactacgag 240aagcacaagg tgtacgcctg cgaagtgacc
caccagggcc tgtctagccc cgtgaccaag 300tctttcaacc ggggcgagtg t
321207107PRTHomo Sapien 207Arg Thr Val Ala Ala Pro Ser Val Phe Ile
Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr Ala Ser Val
Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala Lys Val Gln
Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser Gln Glu Ser
Val Thr Glu Gln Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser Leu Ser Ser
Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75 80Lys His Lys Val
Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser 85 90 95Pro Val Thr
Lys Ser Phe Asn Arg Gly Glu Cys 100 105208321DNAHomo Sapien
208cgaactgtgg ctgcaccatc tgtcttcatc ttcccgccat ctgatgagca
gttgaaatct 60ggaactgcct ctgttgtgtg cctgctgaat aacttctatc ccagagaggc
caaagtacag 120tggaaggtgg ataacgccct ccaatcgggt aactcccagg
agagtgtcac agagcaggag 180agcaaggaca gcacctacag cctcagcagc
accctgacgc tgagcaaagc agactacgag 240aaacacaaag tctacgccgg
cgaagtcacc catcagggcc tgagctcgcc cgtcacaaag 300agcttcaaca
ggggagagtg t 321209107PRTHomo Sapien 209Arg Thr Val Ala Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr
Ala Ser Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala
Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser
Gln Glu Ser Val Thr Glu Gln Glu Ser Lys Asp Ser 50 55 60Thr Tyr Ser
Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75 80Lys
His Lys Val Tyr Ala Gly Glu Val Thr His Gln Gly Leu Ser Ser 85 90
95Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 100 105210321DNAHomo
Sapien 210cgaactgtgg ctgcaccatc tgtcttcatc ttcccgccat ctgatgagca
gttgaaatct 60ggaactgcct ctgttgtgtg cctgctgaat aacttctatc ccagagaggc
caaagtacag 120cggaaggtgg ataacgccct ccaatcgggt aactcccagg
agagtgtcac agagcaggag 180agcaaggaca gcacctacag cctcagcagc
accctgacgc tgagcaaagc agactacgag 240aaacacaaag tctacgcctg
cgaagtcacc catcagggcc tgagctcgcc cgtcacaaag 300agcttcaaca
ggggagagtg t 321211107PRTHomo Sapien 211Arg Thr Val Ala Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr
Ala Ser Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala
Lys Val Gln Arg Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser
Gln Glu Ser Val Thr Glu Gln Glu Ser Lys Asp Ser 50 55 60Thr Tyr Ser
Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75 80Lys
His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser 85 90
95Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 100 105212321DNAHomo
Sapien 212cgaactgtgg ctgcaccatc tgtcttcatc ttcccgccat ctgatgagca
gttgaaatct 60ggaactgcct ctgttgtgtg cctgctgaat aacttctatc ccagagaggc
caaagtacag 120tggaaggtgg ataacgccct ccaatcgggt aactcccagg
agagtgtcac agagcaggac 180agcaaggaca gcacctacag cctcagcagc
accctgacgc tgagcaaagc agactacgag 240aaacacaaac tctacgcctg
cgaagtcacc catcagggcc tgagctcgcc cgtcacaaag 300agcttcaaca
ggggagagtg t 321213107PRTHomo Sapien 213Arg Thr Val Ala Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr
Ala Ser Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala
Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser
Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser
Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75 80Lys
His Lys Leu Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser 85 90
95Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 100 105214321DNAHomo
Sapien 214cgaactgtgg ctgcaccatc tgtcttcatc ttcccgccat ctgatgagca
gttgaaatct 60ggaactgcct ctgttgtgtg cctgctgaat aacttctatc ccagagaggc
caaagtacag 120tggaaggtgg ataacgccct ccaatcgggt aactcccagg
agagtgtcac agagcaggac 180agcaaggaca gcacctacag cctcagcaac
accctgacgc tgagcaaagc agactacgag 240aaacacaaag tctacgcctg
cgaagtcacc catcagggcc tgagctcgcc cgtcacaaag 300agcttcaaca
ggggagagtg c 321215107PRTHomo Sapien 215Arg Thr Val Ala Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr
Ala Ser Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala
Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser
Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser
Leu Ser Asn Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75 80Lys
His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser 85 90
95Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 100 105216312DNAHomo
Sapien 216cccaaggcca accccacggt cactctgttc ccgccctcct ctgaggagct
ccaagccaac 60aaggccacac tagtgtgtct gatcagtgac ttctacccgg gagctgtgac
agtggcttgg 120aaggcagatg gcagccccgt caaggcggga gtggagacga
ccaaaccctc caaacagagc 180aacaacaagt acgcggccag cagctacctg
agcctgacgc ccgagcagtg gaagtcccac 240agaagctaca gctgccaggt
cacgcatgaa gggagcaccg tggagaagac agtggcccct 300acagaatgtt ca
312217104PRTHomo Sapien 217Pro Lys Ala Asn Pro Thr Val Thr Leu Phe
Pro Pro Ser Ser Glu Glu1 5 10 15Leu Gln Ala Asn Lys Ala Thr Leu Val
Cys Leu Ile Ser Asp Phe Tyr 20 25 30Pro Gly Ala Val Thr Val Ala Trp
Lys Ala Asp Gly Ser Pro Val Lys 35 40 45Ala Gly Val Glu Thr Thr Lys
Pro Ser Lys Gln Ser Asn Asn Lys Tyr 50 55 60Ala Ala Ser Ser Tyr Leu
Ser Leu Thr Pro Glu Gln Trp Lys Ser His65 70 75 80Arg Ser Tyr Ser
Cys Gln Val Thr His Glu Gly Ser Thr Val Glu Lys 85 90 95Thr Val Ala
Pro Thr Glu Cys Ser 100218318DNAHomo Sapien 218ggtcagccca
aggccaaccc cactgtcact ctgttcccgc cctcctctga ggagctccaa 60gccaacaagg
ccacactagt gtgtctgatc agtgacttct acccgggagc tgtgacagtg
120gcctggaagg cagatggcag ccccgtcaag gcgggagtgg agaccaccaa
accctccaaa 180cagagcaaca acaagtacgc ggccagcagc tacctgagcc
tgacgcccga gcagtggaag 240tcccacagaa gctacagctg ccaggtcacg
catgaaggga gcaccgtgga gaagacagtg 300gcccctacag aatgttca
318219106PRTHomo Sapien 219Gly Gln Pro Lys Ala Asn Pro Thr Val Thr
Leu Phe Pro Pro Ser Ser1 5 10 15Glu Glu Leu Gln Ala Asn Lys Ala Thr
Leu Val Cys Leu Ile Ser Asp 20 25 30Phe Tyr Pro Gly Ala Val Thr Val
Ala Trp Lys Ala Asp Gly Ser Pro 35 40 45Val Lys Ala Gly Val Glu Thr
Thr Lys Pro Ser Lys Gln Ser Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser
Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys65 70 75 80Ser His Arg Ser
Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr Val 85 90 95Glu Lys Thr
Val Ala Pro Thr Glu Cys Ser 100 105220318DNAHomo Sapien
220ggtcagccca aggccaaccc cactgtcact ctgttcccgc cctcctctga
ggagctccaa 60gccaacaagg ccacactagt gtgtctgatc agtgacttct acccgggagc
tgtgacagtg 120gcctggaagg cagatggcag ccccgtcaag gcgggagtgg
agaccaccaa accctccaaa 180cagagcaaca acaagtacgc ggccagcagc
tacctgagcc tgacgcccga gcagtggaag 240tcccacagaa gctacagctg
ccaggtcacg catgaaggga gcaccgtgga gaagacagtg 300gcccctacag aatgttca
318221318DNAHomo Sapien 221ggccagccta aggccgctcc ttctgtgacc
ctgttccccc catcctccga ggaactgcag 60gctaacaagg ccaccctcgt gtgcctgatc
agcgacttct accctggcgc cgtgaccgtg 120gcctggaagg ctgatagctc
tcctgtgaag gccggcgtgg aaaccaccac cccttccaag 180cagtccaaca
acaaatacgc cgcctcctcc tacctgtccc tgacccctga gcagtggaag
240tcccaccggt cctacagctg ccaagtgacc cacgagggct ccaccgtgga
aaagaccgtg 300gctcctaccg agtgctcc 318222318DNAHomo Sapien
222ggccagccta aagctgcccc cagcgtcacc ctgtttcctc cctccagcga
ggagctccag 60gccaacaagg ccaccctcgt gtgcctgatc tccgacttct atcccggcgc
tgtgaccgtg 120gcttggaaag ccgactccag ccctgtcaaa gccggcgtgg
agaccaccac accctccaag 180cagtccaaca acaagtacgc cgcctccagc
tatctctccc tgacccctga gcagtggaag 240tcccaccggt cctactcctg
tcaggtgacc cacgagggct ccaccgtgga aaagaccgtc 300gcccccaccg agtgctcc
318223106PRTHomo Sapien 223Gly Gln Pro Lys Ala Asn Pro Thr Val Thr
Leu Phe Pro Pro Ser Ser1 5 10 15Glu Glu Leu Gln Ala Asn Lys Ala Thr
Leu Val Cys Leu Ile Ser Asp 20 25 30Phe Tyr Pro Gly Ala Val Thr Val
Ala Trp Lys Ala Asp Gly Ser Pro 35 40 45Val Lys Ala Gly Val Glu Thr
Thr Lys Pro Ser Lys Gln Ser Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser
Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys65 70 75 80Ser His Arg Ser
Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr Val 85 90 95Glu Lys Thr
Val Ala Pro Thr Glu Cys Ser 100 105224318DNAHomo Sapien
224ggtcagccca aggctgcccc ctcggtcact ctgttcccgc cctcctctga
ggagcttcaa 60gccaacaagg ccacactggt gtgtctcata agtgacttct acccgggagc
cgtgacagtg 120gcctggaagg cagatagcag ccccgtcaag gcgggagtgg
agaccaccac accctccaaa 180caaagcaaca acaagtacgc ggccagcagc
tatctgagcc tgacgcctga gcagtggaag 240tcccacagaa gctacagctg
ccaggtcacg catgaaggga gcaccgtgga gaagacagtg 300gcccctacag aatgttca
318225106PRTHomo Sapien 225Gly Gln Pro Lys Ala Ala Pro Ser Val Thr
Leu Phe Pro Pro Ser Ser1 5 10 15Glu Glu Leu Gln Ala Asn Lys Ala Thr
Leu Val Cys Leu Ile Ser Asp 20 25 30Phe Tyr Pro Gly Ala Val Thr Val
Ala Trp Lys Ala Asp Ser Ser Pro 35 40 45Val Lys Ala Gly Val Glu Thr
Thr Thr Pro Ser Lys Gln Ser Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser
Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys65
70 75 80Ser His Arg Ser Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr
Val 85 90 95Glu Lys Thr Val Ala Pro Thr Glu Cys Ser 100
105226312DNAHomo Sapien 226cccaaggctg ccccctcggt cactctgttc
ccaccctcct ctgaggagct tcaagccaac 60aaggccacac tggtgtgtct cataagtgac
ttctacccgg gagccgtgac agttgcctgg 120aaggcagata gcagccccgt
caaggcgggg gtggagacca ccacaccctc caaacaaagc 180aacaacaagt
acgcggccag cagctacctg agcctgacgc ctgagcagtg gaagtcccac
240aaaagctaca gctgccaggt cacgcatgaa gggagcaccg tggagaagac
agttgcccct 300acggaatgtt ca 312227104PRTHomo Sapien 227Pro Lys Ala
Ala Pro Ser Val Thr Leu Phe Pro Pro Ser Ser Glu Glu1 5 10 15Leu Gln
Ala Asn Lys Ala Thr Leu Val Cys Leu Ile Ser Asp Phe Tyr 20 25 30Pro
Gly Ala Val Thr Val Ala Trp Lys Ala Asp Ser Ser Pro Val Lys 35 40
45Ala Gly Val Glu Thr Thr Thr Pro Ser Lys Gln Ser Asn Asn Lys Tyr
50 55 60Ala Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys Ser
His65 70 75 80Lys Ser Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr
Val Glu Lys 85 90 95Thr Val Ala Pro Thr Glu Cys Ser
100228318DNAHomo Sapien 228ggtcagccca aggctgcccc ctcggtcact
ctgttcccac cctcctctga ggagcttcaa 60gccaacaagg ccacactggt gtgtctcata
agtgacttct acccggggcc agtgacagtt 120gcctggaagg cagatagcag
ccccgtcaag gcgggggtgg agaccaccac accctccaaa 180caaagcaaca
acaagtacgc ggccagcagc tacctgagcc tgacgcctga gcagtggaag
240tcccacaaaa gctacagctg ccaggtcacg catgaaggga gcaccgtgga
gaagacagtg 300gcccctacgg aatgttca 318229106PRTHomo Sapien 229Gly
Gln Pro Lys Ala Ala Pro Ser Val Thr Leu Phe Pro Pro Ser Ser1 5 10
15Glu Glu Leu Gln Ala Asn Lys Ala Thr Leu Val Cys Leu Ile Ser Asp
20 25 30Phe Tyr Pro Gly Pro Val Thr Val Ala Trp Lys Ala Asp Ser Ser
Pro 35 40 45Val Lys Ala Gly Val Glu Thr Thr Thr Pro Ser Lys Gln Ser
Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu
Gln Trp Lys65 70 75 80Ser His Lys Ser Tyr Ser Cys Gln Val Thr His
Glu Gly Ser Thr Val 85 90 95Glu Lys Thr Val Ala Pro Thr Glu Cys Ser
100 105230318DNAHomo Sapien 230ggtcagccca aggctgcccc ctcggtcact
ctgttcccac cctcctctga ggagcttcaa 60gccaacaagg ccacactggt gtgtctcata
agtgacttct acccgggagc cgtgacagtg 120gcctggaagg cagatagcag
ccccgtcaag gcgggagtgg agaccaccac accctccaaa 180caaagcaaca
acaagtacgc ggccagcagc tacctgagcc tgacgcctga gcagtggaag
240tcccacaaaa gctacagctg ccaggtcacg catgaaggga gcaccgtgga
gaagacagtg 300gcccctacag aatgttca 318231106PRTHomo Sapien 231Gly
Gln Pro Lys Ala Ala Pro Ser Val Thr Leu Phe Pro Pro Ser Ser1 5 10
15Glu Glu Leu Gln Ala Asn Lys Ala Thr Leu Val Cys Leu Ile Ser Asp
20 25 30Phe Tyr Pro Gly Ala Val Thr Val Ala Trp Lys Ala Asp Ser Ser
Pro 35 40 45Val Lys Ala Gly Val Glu Thr Thr Thr Pro Ser Lys Gln Ser
Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu
Gln Trp Lys65 70 75 80Ser His Lys Ser Tyr Ser Cys Gln Val Thr His
Glu Gly Ser Thr Val 85 90 95Glu Lys Thr Val Ala Pro Thr Glu Cys Ser
100 105232318DNAHomo Sapien 232ggtcagccca aggctgcccc ctcggtcact
ctgttcccgc cctcctctga ggagcttcaa 60gccaacaagg ccacactggt gtgtctcata
agtgacttct acccgggagc cgtgacagtg 120gcctggaagg cagatagcag
ccccgtcaag gcgggagtgg agaccaccac accctccaaa 180caaagcaaca
acaagtacgc ggccagcagc tacctgagcc tgacgcctga gcagtggaag
240tcccacagaa gctacagctg ccaggtcacg catgaaggga gcaccgtgga
gaagacagtg 300gcccctacag aatgttca 318233106PRTHomo Sapien 233Gly
Gln Pro Lys Ala Ala Pro Ser Val Thr Leu Phe Pro Pro Ser Ser1 5 10
15Glu Glu Leu Gln Ala Asn Lys Ala Thr Leu Val Cys Leu Ile Ser Asp
20 25 30Phe Tyr Pro Gly Ala Val Thr Val Ala Trp Lys Ala Asp Ser Ser
Pro 35 40 45Val Lys Ala Gly Val Glu Thr Thr Thr Pro Ser Lys Gln Ser
Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu
Gln Trp Lys65 70 75 80Ser His Arg Ser Tyr Ser Cys Gln Val Thr His
Glu Gly Ser Thr Val 85 90 95Glu Lys Thr Val Ala Pro Thr Glu Cys Ser
100 105234318DNAHomo Sapien 234ggtcagccca aggctgcccc atcggtcact
ctgttcccgc cctcctctga ggagcttcaa 60gccaacaagg ccacactggt gtgcctgatc
agtgacttct acccgggagc tgtgaaagtg 120gcctggaagg cagatggcag
ccccgtcaac acgggagtgg agaccaccac accctccaaa 180cagagcaaca
acaagtacgc ggccagcagc tacctgagcc tgacgcctga gcagtggaag
240tcccacagaa gctacagctg ccaggtcacg catgaaggga gcaccgtgga
gaagacagtg 300gcccctgcag aatgttca 318235106PRTHomo Sapien 235Gly
Gln Pro Lys Ala Ala Pro Ser Val Thr Leu Phe Pro Pro Ser Ser1 5 10
15Glu Glu Leu Gln Ala Asn Lys Ala Thr Leu Val Cys Leu Ile Ser Asp
20 25 30Phe Tyr Pro Gly Ala Val Lys Val Ala Trp Lys Ala Asp Gly Ser
Pro 35 40 45Val Asn Thr Gly Val Glu Thr Thr Thr Pro Ser Lys Gln Ser
Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu
Gln Trp Lys65 70 75 80Ser His Arg Ser Tyr Ser Cys Gln Val Thr His
Glu Gly Ser Thr Val 85 90 95Glu Lys Thr Val Ala Pro Ala Glu Cys Ser
100 105236318DNAHomo Sapien 236ggtcagccca aggctgcccc atcggtcact
ctgttcccac cctcctctga ggagcttcaa 60gccaacaagg ccacactggt gtgtctcgta
agtgacttct acccgggagc cgtgacagtg 120gcctggaagg cagatggcag
ccccgtcaag gtgggagtgg agaccaccaa accctccaaa 180caaagcaaca
acaagtatgc ggccagcagc tacctgagcc tgacgcccga gcagtggaag
240tcccacagaa gctacagctg ccgggtcacg catgaaggga gcaccgtgga
gaagacagtg 300gcccctgcag aatgctct 318237106PRTHomo Sapien 237Gly
Gln Pro Lys Ala Ala Pro Ser Val Thr Leu Phe Pro Pro Ser Ser1 5 10
15Glu Glu Leu Gln Ala Asn Lys Ala Thr Leu Val Cys Leu Val Ser Asp
20 25 30Phe Tyr Pro Gly Ala Val Thr Val Ala Trp Lys Ala Asp Gly Ser
Pro 35 40 45Val Lys Val Gly Val Glu Thr Thr Lys Pro Ser Lys Gln Ser
Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu
Gln Trp Lys65 70 75 80Ser His Arg Ser Tyr Ser Cys Arg Val Thr His
Glu Gly Ser Thr Val 85 90 95Glu Lys Thr Val Ala Pro Ala Glu Cys Ser
100 1052388PRTHomo Sapien 238Gly Phe Thr Phe Ser Asp Tyr Tyr1
52398PRTHomo Sapien 239Ile Ser Thr Ser Gly Ser Thr Ile1
524016PRTHomo Sapien 240Ala Arg Gly Ile Thr Gly Thr Asn Phe Tyr His
Tyr Gly Leu Gly Val1 5 10 152415PRTHomo Sapien 241Asp Tyr Tyr Met
Ser1 524217PRTHomo Sapien 242Tyr Ile Ser Thr Ser Gly Ser Thr Ile
Tyr Tyr Ala Asp Ser Val Lys1 5 10 15Gly24314PRTHomo Sapien 243Gly
Ile Thr Gly Thr Asn Phe Tyr His Tyr Gly Leu Gly Val1 5
10244123PRTHomo Sapien 244Gln Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Asp Tyr 20 25 30Tyr Met Ser Trp Ile Arg Gln Val
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Tyr Ile Ser Thr Ser Gly
Ser Thr Ile Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn
Ser Leu Arg Ala Glu Asp Ala Ala Val Tyr His Cys 85 90 95Ala Arg Gly
Ile Thr Gly Thr Asn Phe Tyr His Tyr Gly Leu Gly Val 100 105 110Trp
Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120245370DNAHomo Sapien
245caggtgcagc tggtggagtc tgggggaggc ttggtcaagc ctggagggtc
cctgagactc 60tcctgtgcag cctctggatt caccttcagt gactactaca tgagctggat
ccgccaggtt 120ccagggaagg ggctggagtg ggtttcatac attagtacta
gtggtagtac catatactac 180gcagactctg tgaagggccg attcaccatc
tccagggaca acgccaagaa ctcactgtat 240ctacaaatga acagcctgag
agccgaggac gcggccgtgt atcactgtgc gagaggtata 300actggaacta
acttctacca ctacggtttg ggcgtctggg gccaagggac cacggtcacc
360gtctcctcag 370246453PRTHomo Sapien 246Gln Val Gln Leu Val Glu
Ser Gly Gly Gly Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser
Cys Ala Ala Ser Gly Phe Thr Phe Ser Asp Tyr 20 25 30Tyr Met Ser Trp
Ile Arg Gln Val Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Tyr Ile
Ser Thr Ser Gly Ser Thr Ile Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly
Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Ala Ala Val Tyr His Cys
85 90 95Ala Arg Gly Ile Thr Gly Thr Asn Phe Tyr His Tyr Gly Leu Gly
Val 100 105 110Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser Ala Ser
Thr Lys Gly 115 120 125Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys
Ser Thr Ser Gly Gly 130 135 140Thr Ala Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val145 150 155 160Thr Val Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe 165 170 175Pro Ala Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val 180 185 190Thr Val
Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val 195 200
205Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys
210 215 220Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro
Glu Leu225 230 235 240Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr 245 250 255Leu Met Ile Ser Arg Thr Pro Glu Val
Thr Cys Val Val Val Asp Val 260 265 270Ser His Glu Asp Pro Glu Val
Lys Phe Asn Trp Tyr Val Asp Gly Val 275 280 285Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser 290 295 300Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu305 310 315
320Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala
325 330 335Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg
Glu Pro 340 345 350Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu
Thr Lys Asn Gln 355 360 365Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile Ala 370 375 380Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr Thr385 390 395 400Pro Pro Val Leu Asp
Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu 405 410 415Thr Val Asp
Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser 420 425 430Val
Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser 435 440
445Leu Ser Pro Gly Lys 4502471359DNAHomo Sapien 247caggtgcagc
tggtggagtc tgggggaggc ttggtcaagc ctggagggtc cctgagactc 60tcctgtgcag
cctctggatt caccttcagt gactactaca tgagctggat ccgccaggtt
120ccagggaagg ggctggagtg ggtttcatac attagtacta gtggtagtac
catatactac 180gcagactctg tgaagggccg attcaccatc tccagggaca
acgccaagaa ctcactgtat 240ctacaaatga acagcctgag agccgaggac
gcggccgtgt atcactgtgc gagaggtata 300actggaacta acttctacca
ctacggtttg ggcgtctggg gccaagggac cacggtcacc 360gtctcctcag
ccagcaccaa gggcccctct gtgttccctc tggccccttc cagcaagtcc
420acctctggcg gaacagccgc tctgggctgc ctcgtgaagg actacttccc
cgagcctgtg 480accgtgtcct ggaactctgg cgctctgacc agcggagtgc
acaccttccc tgctgtgctg 540cagtcctccg gcctgtactc cctgtcctcc
gtcgtgaccg tgccttccag ctctctgggc 600acccagacct acatctgcaa
cgtgaaccac aagccctcca acaccaaggt ggacaagaag 660gtggaaccca
agtcctgcga caagacccac acctgtcccc cttgtcctgc ccctgaactg
720ctgggcggac cttccgtgtt cctgttcccc ccaaagccca aggacaccct
gatgatctcc 780cggacccccg aagtgacctg cgtggtggtg gatgtgtccc
acgaggaccc tgaagtgaag 840ttcaattggt acgtggacgg cgtggaagtg
cacaacgcca agaccaagcc tagagaggaa 900cagtacaact ccacctaccg
ggtggtgtcc gtgctgaccg tgctgcacca ggattggctg 960aacggcaaag
agtacaagtg caaggtgtcc aacaaggccc tgcctgcccc catcgaaaag
1020accatctcca aggccaaggg ccagccccgg gaaccccagg tgtacacact
gccccctagc 1080agggacgagc tgaccaagaa ccaggtgtcc ctgacctgtc
tcgtgaaagg cttctacccc 1140tccgatatcg ccgtggaatg ggagtccaac
ggccagcctg agaacaacta caagaccacc 1200ccccctgtgc tggactccga
cggctcattc ttcctgtaca gcaagctgac agtggacaag 1260tcccggtggc
agcagggcaa cgtgttctcc tgctccgtga tgcacgaggc cctgcacaac
1320cactacaccc agaagtccct gtccctgagc cccggcaag 13592486PRTHomo
Sapien 248Gln Gly Ile Asn Ser Trp1 52493PRTHomo Sapien 249Ala Ala
Ser12509PRTHomo Sapien 250Gln Gln Val Asn Ser Phe Pro Leu Thr1
525111PRTHomo Sapien 251Arg Ala Ser Gln Gly Ile Asn Ser Trp Leu
Ala1 5 102527PRTHomo Sapien 252Ala Ala Ser Thr Leu Gln Ser1
52539PRTHomo Sapien 253Gln Gln Val Asn Ser Phe Pro Leu Thr1
5254107PRTHomo Sapien 254Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala
Ser Gln Gly Ile Asn Ser Trp 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro
Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Ala Ala Ser Thr Leu Gln
Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Ala Asp
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Gln Gln Val Asn Ser Phe Pro Leu 85 90 95Thr Phe Gly
Gly Gly Thr Lys Val Glu Ile Lys 100 105255322DNAHomo Sapien
255gacatccaga tgacccagtc tccatcttcc gtgtctgcat ctgtaggaga
cagagtcacc 60atcacttgtc gggcgagtca gggtattaac agctggttag cctggtatca
gcagaaacca 120gggaaagccc ctaagctcct gatctatgct gcatccactt
tgcaaagtgg ggtcccatca 180aggttcagcg gcagtgggtc tggggcagat
ttcactctca ccatcagcag cctgcagcct 240gaagattttg caacttacta
ttgtcaacag gttaacagtt tcccgctcac tttcggcgga 300gggaccaagg
tggagatcaa ac 322256214PRTHomo Sapien 256Asp Ile Gln Met Thr Gln
Ser Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Gly Ile Asn Ser Trp 20 25 30Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Ala Ala
Ser Thr Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Ala Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Val Asn Ser Phe Pro Leu
85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala
Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu
Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 210257642DNAHomo Sapien 257gacatccaga
tgacccagtc tccatcttcc
gtgtctgcat ctgtaggaga cagagtcacc 60atcacttgtc gggcgagtca gggtattaac
agctggttag cctggtatca gcagaaacca 120gggaaagccc ctaagctcct
gatctatgct gcatccactt tgcaaagtgg ggtcccatca 180aggttcagcg
gcagtgggtc tggggcagat ttcactctca ccatcagcag cctgcagcct
240gaagattttg caacttacta ttgtcaacag gttaacagtt tcccgctcac
tttcggcgga 300gggaccaagg tggagatcaa acgtacggtg gccgctccct
ccgtgttcat cttcccacct 360tccgacgagc agctgaagtc cggcaccgct
tctgtcgtgt gcctgctgaa caacttctac 420ccccgcgagg ccaaggtgca
gtggaaggtg gacaacgccc tgcagtccgg caactcccag 480gaatccgtga
ccgagcagga ctccaaggac agcacctact ccctgtcctc caccctgacc
540ctgtccaagg ccgactacga gaagcacaag gtgtacgcct gcgaagtgac
ccaccagggc 600ctgtctagcc ccgtgaccaa gtctttcaac cggggcgagt gt
6422588PRTHomo Sapien 258Gly Phe Thr Phe Ser Tyr Tyr Ala1
52598PRTHomo Sapien 259Ile Ser Gly Gly Gly Gly Asn Thr1
526016PRTHomo Sapien 260Ala Lys Asp Arg Met Lys Gln Leu Val Arg Ala
Tyr Tyr Phe Asp Tyr1 5 10 152615PRTHomo Sapien 261Tyr Tyr Ala Met
Ser1 526217PRTHomo Sapien 262Thr Ile Ser Gly Gly Gly Gly Asn Thr
His Tyr Ala Asp Ser Val Lys1 5 10 15Gly26314PRTHomo Sapien 263Asp
Arg Met Lys Gln Leu Val Arg Ala Tyr Tyr Phe Asp Tyr1 5
10264123PRTHomo Sapien 264Glu Val Pro Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Tyr Tyr 20 25 30Ala Met Ser Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Asp Trp Val 35 40 45Ser Thr Ile Ser Gly Gly Gly
Gly Asn Thr His Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65 70 75 80Leu His Met Asn
Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Lys Asp
Arg Met Lys Gln Leu Val Arg Ala Tyr Tyr Phe Asp Tyr 100 105 110Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser 115 120265370DNAHomo Sapien
265gaggtgccgc tggtggagtc tgggggaggc ttggtacagc ctggggggtc
cctgagactc 60tcctgtgcag cctctggatt cacgtttagc tactatgcca tgagctgggt
ccgtcaggct 120ccagggaagg ggctggactg ggtctcaact attagtggtg
gtggtggtaa cacacactac 180gcagactccg tgaagggccg attcactata
tccagagaca attccaagaa cacgctgtat 240ctgcacatga acagcctgag
agccgaagac acggccgtct attactgtgc gaaggatcgg 300atgaaacagc
tcgtccgggc ctactacttt gactactggg gccagggaac cctggtcacc
360gtctcctcag 370266453PRTHomo Sapien 266Glu Val Pro Leu Val Glu
Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser
Cys Ala Ala Ser Gly Phe Thr Phe Ser Tyr Tyr 20 25 30Ala Met Ser Trp
Val Arg Gln Ala Pro Gly Lys Gly Leu Asp Trp Val 35 40 45Ser Thr Ile
Ser Gly Gly Gly Gly Asn Thr His Tyr Ala Asp Ser Val 50 55 60Lys Gly
Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65 70 75
80Leu His Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Lys Asp Arg Met Lys Gln Leu Val Arg Ala Tyr Tyr Phe Asp
Tyr 100 105 110Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser
Thr Lys Gly 115 120 125Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys
Ser Thr Ser Gly Gly 130 135 140Thr Ala Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val145 150 155 160Thr Val Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe 165 170 175Pro Ala Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val 180 185 190Thr Val
Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val 195 200
205Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys
210 215 220Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro
Glu Leu225 230 235 240Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr 245 250 255Leu Met Ile Ser Arg Thr Pro Glu Val
Thr Cys Val Val Val Asp Val 260 265 270Ser His Glu Asp Pro Glu Val
Lys Phe Asn Trp Tyr Val Asp Gly Val 275 280 285Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser 290 295 300Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu305 310 315
320Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala
325 330 335Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg
Glu Pro 340 345 350Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu
Thr Lys Asn Gln 355 360 365Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile Ala 370 375 380Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr Thr385 390 395 400Pro Pro Val Leu Asp
Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu 405 410 415Thr Val Asp
Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser 420 425 430Val
Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser 435 440
445Leu Ser Pro Gly Lys 4502671359DNAHomo Sapien 267gaggtgccgc
tggtggagtc tgggggaggc ttggtacagc ctggggggtc cctgagactc 60tcctgtgcag
cctctggatt cacgtttagc tactatgcca tgagctgggt ccgtcaggct
120ccagggaagg ggctggactg ggtctcaact attagtggtg gtggtggtaa
cacacactac 180gcagactccg tgaagggccg attcactata tccagagaca
attccaagaa cacgctgtat 240ctgcacatga acagcctgag agccgaagac
acggccgtct attactgtgc gaaggatcgg 300atgaaacagc tcgtccgggc
ctactacttt gactactggg gccagggaac cctggtcacc 360gtctcctcag
ccagcaccaa gggcccctct gtgttccctc tggccccttc cagcaagtcc
420acctctggcg gaacagccgc tctgggctgc ctcgtgaagg actacttccc
cgagcctgtg 480accgtgtcct ggaactctgg cgctctgacc agcggagtgc
acaccttccc tgctgtgctg 540cagtcctccg gcctgtactc cctgtcctcc
gtcgtgaccg tgccttccag ctctctgggc 600acccagacct acatctgcaa
cgtgaaccac aagccctcca acaccaaggt ggacaagaag 660gtggaaccca
agtcctgcga caagacccac acctgtcccc cttgtcctgc ccctgaactg
720ctgggcggac cttccgtgtt cctgttcccc ccaaagccca aggacaccct
gatgatctcc 780cggacccccg aagtgacctg cgtggtggtg gatgtgtccc
acgaggaccc tgaagtgaag 840ttcaattggt acgtggacgg cgtggaagtg
cacaacgcca agaccaagcc tagagaggaa 900cagtacaact ccacctaccg
ggtggtgtcc gtgctgaccg tgctgcacca ggattggctg 960aacggcaaag
agtacaagtg caaggtgtcc aacaaggccc tgcctgcccc catcgaaaag
1020accatctcca aggccaaggg ccagccccgg gaaccccagg tgtacacact
gccccctagc 1080agggacgagc tgaccaagaa ccaggtgtcc ctgacctgtc
tcgtgaaagg cttctacccc 1140tccgatatcg ccgtggaatg ggagtccaac
ggccagcctg agaacaacta caagaccacc 1200ccccctgtgc tggactccga
cggctcattc ttcctgtaca gcaagctgac agtggacaag 1260tcccggtggc
agcagggcaa cgtgttctcc tgctccgtga tgcacgaggc cctgcacaac
1320cactacaccc agaagtccct gtccctgagc cccggcaag 13592686PRTHomo
Sapien 268Gln Asp Ile Ser Thr Tyr1 52693PRTHomo Sapien 269Gly Thr
Ser12709PRTHomo Sapien 270Gln Gln Leu His Thr Asp Pro Ile Thr1
527111PRTHomo Sapien 271Trp Ala Ser Gln Asp Ile Ser Thr Tyr Leu
Gly1 5 102727PRTHomo Sapien 272Gly Thr Ser Ser Leu Gln Ser1
52739PRTHomo Sapien 273Gln Gln Leu His Thr Asp Pro Ile Thr1
5274107PRTHomo Sapien 274Asp Ile Gln Leu Thr Gln Ser Pro Ser Phe
Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Trp Ala
Ser Gln Asp Ile Ser Thr Tyr 20 25 30Leu Gly Trp Tyr Gln Gln Lys Pro
Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Thr Ser Ser Leu Gln
Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Glu
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Gln Gln Leu His Thr Asp Pro Ile 85 90 95Thr Phe Gly
Gln Gly Thr Arg Leu Glu Ile Lys 100 105275322DNAHomo Sapien
275gacatccagt tgacccagtc tccatccttc ctgtctgcat ctgtaggaga
cagagtcacc 60atcacttgct gggccagtca ggacattagc acttatttag gctggtatca
gcaaaaacca 120gggaaagccc ctaagctcct gatctatggt acatccagtt
tgcaaagtgg ggtcccatca 180aggttcagcg gcagtggatc tgggacagaa
ttcactctca caatcagcag cctgcagcct 240gaagattttg caacttatta
ctgtcaacag cttcatactg acccgatcac cttcggccaa 300gggacacgac
tggagatcaa ac 322276214PRTHomo Sapien 276Asp Ile Gln Leu Thr Gln
Ser Pro Ser Phe Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Trp Ala Ser Gln Asp Ile Ser Thr Tyr 20 25 30Leu Gly Trp Tyr
Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Thr
Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Leu His Thr Asp Pro Ile
85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys Arg Thr Val Ala
Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu
Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 210277642DNAHomo Sapien 277gacatccagt
tgacccagtc tccatccttc ctgtctgcat ctgtaggaga cagagtcacc 60atcacttgct
gggccagtca ggacattagc acttatttag gctggtatca gcaaaaacca
120gggaaagccc ctaagctcct gatctatggt acatccagtt tgcaaagtgg
ggtcccatca 180aggttcagcg gcagtggatc tgggacagaa ttcactctca
caatcagcag cctgcagcct 240gaagattttg caacttatta ctgtcaacag
cttcatactg acccgatcac cttcggccaa 300gggacacgac tggagatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc tgcagtccgg
caactcccag 480gaatccgtga ccgagcagga ctccaaggac agcacctact
ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga gaagcacaag
gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc ccgtgaccaa
gtctttcaac cggggcgagt gt 6422788PRTHomo Sapien 278Gly Phe Thr Phe
Ser Ser Tyr Trp1 52798PRTHomo Sapien 279Ile Lys Gln Asp Gly Ser Glu
Lys1 528012PRTHomo Sapien 280Ala Arg Val Arg Gln Trp Ser Asp Tyr
Ser Asp Tyr1 5 102815PRTHomo Sapien 281Ser Tyr Trp Met Asn1
528217PRTHomo Sapien 282Asn Ile Lys Gln Asp Gly Ser Glu Lys Tyr Tyr
Val Asp Ser Val Lys1 5 10 15Gly28310PRTHomo Sapien 283Val Arg Gln
Trp Ser Asp Tyr Ser Asp Tyr1 5 10284119PRTHomo Sapien 284Glu Val
His Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser
Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25
30Trp Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Ala Asn Ile Lys Gln Asp Gly Ser Glu Lys Tyr Tyr Val Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Val Ser Arg Asp Asn Ala Lys Asn Ser
Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala
Val Tyr Tyr Cys 85 90 95Ala Arg Val Arg Gln Trp Ser Asp Tyr Ser Asp
Tyr Trp Gly Gln Gly 100 105 110Thr Pro Val Thr Val Ser Ser
115285358DNAHomo Sapien 285gaggtgcacc tggtggagtc tgggggaggc
ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagt
agctattgga tgaactgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtggccaac ataaagcaag atggaagtga gaaatactat 180gtggactctg
tgaagggccg cttcaccgtc tccagagaca acgccaagaa ctcactgtat
240ctgcaaatga acagcctgag agccgaggac acggctgtgt attactgtgc
gagagttcga 300caatggtccg actactctga ctactggggc cagggaaccc
cggtcaccgt ctcctcag 358286449PRTHomo Sapien 286Glu Val His Leu Val
Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu
Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30Trp Met Asn
Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ala Asn
Ile Lys Gln Asp Gly Ser Glu Lys Tyr Tyr Val Asp Ser Val 50 55 60Lys
Gly Arg Phe Thr Val Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Val Arg Gln Trp Ser Asp Tyr Ser Asp Tyr Trp Gly Gln
Gly 100 105 110Thr Pro Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro
Ser Val Phe 115 120 125Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly
Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu Val Lys Asp Tyr Phe Pro
Glu Pro Val Thr Val Ser Trp145 150 155 160Asn Ser Gly Ala Leu Thr
Ser Gly Val His Thr Phe Pro Ala Val Leu 165 170 175Gln Ser Ser Gly
Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro Ser 180 185 190Ser Ser
Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His Lys Pro 195 200
205Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Asp Lys
210 215 220Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly
Gly Pro225 230 235 240Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser 245 250 255Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser His Glu Asp 260 265 270Pro Glu Val Lys Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn 275 280 285Ala Lys Thr Lys Pro
Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val 290 295 300Val Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu305 310 315
320Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys
325 330 335Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val
Tyr Thr 340 345 350Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln
Val Ser Leu Thr 355 360 365Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu 370 375 380Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Val Leu385 390 395 400Asp Ser Asp Gly Ser
Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys 405 410 415Ser Arg Trp
Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu 420 425 430Ala
Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly 435 440
445Lys2871347DNAHomo Sapien 287gaggtgcacc tggtggagtc tgggggaggc
ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagt
agctattgga tgaactgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtggccaac ataaagcaag atggaagtga gaaatactat 180gtggactctg
tgaagggccg cttcaccgtc tccagagaca acgccaagaa ctcactgtat
240ctgcaaatga acagcctgag agccgaggac acggctgtgt attactgtgc
gagagttcga 300caatggtccg actactctga ctactggggc cagggaaccc
cggtcaccgt ctcctcagcc 360agcaccaagg gcccctctgt gttccctctg
gccccttcca gcaagtccac ctctggcgga 420acagccgctc tgggctgcct
cgtgaaggac tacttccccg agcctgtgac cgtgtcctgg 480aactctggcg
ctctgaccag cggagtgcac accttccctg ctgtgctgca gtcctccggc
540ctgtactccc tgtcctccgt cgtgaccgtg ccttccagct ctctgggcac
ccagacctac 600atctgcaacg tgaaccacaa gccctccaac accaaggtgg
acaagaaggt ggaacccaag 660tcctgcgaca agacccacac ctgtccccct
tgtcctgccc ctgaactgct gggcggacct 720tccgtgttcc tgttcccccc
aaagcccaag gacaccctga tgatctcccg gacccccgaa 780gtgacctgcg
tggtggtgga tgtgtcccac
gaggaccctg aagtgaagtt caattggtac 840gtggacggcg tggaagtgca
caacgccaag accaagccta gagaggaaca gtacaactcc 900acctaccggg
tggtgtccgt gctgaccgtg ctgcaccagg attggctgaa cggcaaagag
960tacaagtgca aggtgtccaa caaggccctg cctgccccca tcgaaaagac
catctccaag 1020gccaagggcc agccccggga accccaggtg tacacactgc
cccctagcag ggacgagctg 1080accaagaacc aggtgtccct gacctgtctc
gtgaaaggct tctacccctc cgatatcgcc 1140gtggaatggg agtccaacgg
ccagcctgag aacaactaca agaccacccc ccctgtgctg 1200gactccgacg
gctcattctt cctgtacagc aagctgacag tggacaagtc ccggtggcag
1260cagggcaacg tgttctcctg ctccgtgatg cacgaggccc tgcacaacca
ctacacccag 1320aagtccctgt ccctgagccc cggcaag 13472886PRTHomo Sapien
288Gln Gly Ile Ser Ser Trp1 52893PRTHomo Sapien 289Ala Ala
Ser12909PRTHomo Sapien 290Gln Gln Ala Asn Ser Phe Pro Phe Thr1
529111PRTHomo Sapien 291Arg Ala Ser Gln Gly Ile Ser Ser Trp Leu
Ala1 5 102927PRTHomo Sapien 292Ala Ala Ser Ser Leu Gln Ser1
52939PRTHomo Sapien 293Gln Gln Ala Asn Ser Phe Pro Phe Thr1
5294107PRTHomo Sapien 294Asp Ile Gln Met Thr Gln Ser Pro Ser Ser
Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala
Ser Gln Gly Ile Ser Ser Trp 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro
Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Ala Ala Ser Ser Leu Gln
Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala
Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Phe Pro Phe 85 90 95Thr Phe Gly
Pro Gly Thr Lys Val Asp Ile Lys 100 105295322DNAHomo Sapien
295gacatccaga tgacccagtc tccatcttcc gtgtctgcat ctgtaggaga
cagagtcacc 60atcacttgtc gggcgagtca gggtattagc agctggttag cctggtatca
gcagaaacca 120gggaaagccc ctaagctcct gatctatgct gcatccagtt
tgcaaagtgg ggtcccatca 180aggttcagcg gcagtggatc tgggacagat
ttcactctca ccatcagcag cctgcagcct 240gaagattttg caacttacta
ttgtcaacag gctaacagtt tcccattcac tttcggccct 300gggaccaaag
tggatatcaa ac 322296214PRTHomo Sapien 296Asp Ile Gln Met Thr Gln
Ser Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Gly Ile Ser Ser Trp 20 25 30Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Ala Ala
Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Phe Pro Phe
85 90 95Thr Phe Gly Pro Gly Thr Lys Val Asp Ile Lys Arg Thr Val Ala
Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu
Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 210297642DNAHomo Sapien 297gacatccaga
tgacccagtc tccatcttcc gtgtctgcat ctgtaggaga cagagtcacc 60atcacttgtc
gggcgagtca gggtattagc agctggttag cctggtatca gcagaaacca
120gggaaagccc ctaagctcct gatctatgct gcatccagtt tgcaaagtgg
ggtcccatca 180aggttcagcg gcagtggatc tgggacagat ttcactctca
ccatcagcag cctgcagcct 240gaagattttg caacttacta ttgtcaacag
gctaacagtt tcccattcac tttcggccct 300gggaccaaag tggatatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc tgcagtccgg
caactcccag 480gaatccgtga ccgagcagga ctccaaggac agcacctact
ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga gaagcacaag
gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc ccgtgaccaa
gtctttcaac cggggcgagt gt 642298214PRTHomo Sapien 298Asp Ile Gln Met
Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val
Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Tyr 20 25 30Leu Asn
Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr
Tyr Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55
60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65
70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ser Tyr Ser Thr Pro
Ile 85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys Arg Thr Val
Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln
Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn
Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn
Ala Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu
Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu
Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala
Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 210299453PRTHomo Sapien 299Glu Val Gln
Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Arg1 5 10 15Ser Leu
Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Asp Asp Tyr 20 25 30Ala
Met His Trp Val Arg Gln Val Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Gly Ile Ser Trp Ile Arg Thr Gly Ile Gly Tyr Ala Asp Ser Val
50 55 60Lys Gly Arg Phe Thr Ile Phe Arg Asp Asn Ala Lys Asn Ser Leu
Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Leu
Tyr Tyr Cys 85 90 95Ala Lys Asp Met Lys Gly Ser Gly Thr Tyr Gly Gly
Trp Phe Asp Thr 100 105 110Trp Gly Gln Gly Thr Leu Val Thr Val Ser
Ser Ala Ser Thr Lys Gly 115 120 125Pro Ser Val Phe Pro Leu Ala Pro
Ser Ser Lys Ser Thr Ser Gly Gly 130 135 140Thr Ala Ala Leu Gly Cys
Leu Val Lys Asp Tyr Phe Pro Glu Pro Val145 150 155 160Thr Val Ser
Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe 165 170 175Pro
Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val 180 185
190Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val
195 200 205Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu
Pro Lys 210 215 220Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro
Ala Pro Glu Leu225 230 235 240Ala Gly Ala Pro Ser Val Phe Leu Phe
Pro Pro Lys Pro Lys Asp Thr 245 250 255Leu Met Ile Ser Arg Thr Pro
Glu Val Thr Cys Val Val Val Asp Val 260 265 270Ser His Glu Asp Pro
Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val 275 280 285Glu Val His
Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser 290 295 300Thr
Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu305 310
315 320Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro
Ala 325 330 335Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro 340 345 350Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu
Leu Thr Lys Asn Gln 355 360 365Val Ser Leu Thr Cys Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala 370 375 380Val Glu Trp Glu Ser Asn Gly
Gln Pro Glu Asn Asn Tyr Lys Thr Thr385 390 395 400Pro Pro Val Leu
Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu 405 410 415Thr Val
Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser 420 425
430Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
435 440 445Leu Ser Pro Gly Lys 450300347PRTHomo Sapien 300Asp Ile
Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp
Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Tyr 20 25
30Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile
35 40 45Tyr Val Ala Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser
Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu
Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ser Tyr
Ser Thr Pro Ile 85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys
Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser
Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu
Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys
Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser
Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170
175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr
180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr
Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys Ala Pro Thr Ser Ser Ser
Thr Lys Lys Thr 210 215 220Gln Leu Gln Leu Glu His Leu Leu Leu Asp
Leu Gln Met Ile Leu Asn225 230 235 240Gly Ile Asn Asn Tyr Lys Asn
Pro Lys Leu Thr Arg Met Leu Thr Phe 245 250 255Lys Phe Tyr Met Pro
Lys Lys Ala Thr Glu Leu Lys His Leu Gln Cys 260 265 270Leu Glu Glu
Glu Leu Lys Pro Leu Glu Glu Val Leu Asn Leu Ala Gln 275 280 285Ser
Lys Asn Phe His Leu Arg Pro Arg Asp Leu Ile Ser Asn Ile Asn 290 295
300Val Ile Val Leu Glu Leu Lys Gly Ser Glu Thr Thr Phe Met Cys
Glu305 310 315 320Tyr Ala Asp Glu Thr Ala Thr Ile Val Glu Phe Leu
Asn Arg Trp Ile 325 330 335Thr Phe Cys Gln Ser Ile Ile Ser Thr Leu
Thr 340 345301133PRTHomo Sapien 301Ala Pro Thr Ser Ser Ser Thr Lys
Lys Thr Gln Leu Gln Leu Glu His1 5 10 15Leu Leu Leu Asp Leu Gln Met
Ile Leu Asn Gly Ile Asn Asn Tyr Lys 20 25 30Asn Pro Lys Leu Thr Arg
Met Leu Thr Phe Lys Phe Tyr Met Pro Lys 35 40 45Lys Ala Thr Glu Leu
Lys His Leu Gln Cys Leu Glu Glu Glu Leu Lys 50 55 60Pro Leu Glu Glu
Val Leu Asn Leu Ala Gln Ser Lys Asn Phe His Leu65 70 75 80Arg Pro
Arg Asp Leu Ile Ser Asn Ile Asn Val Ile Val Leu Glu Leu 85 90 95Lys
Gly Ser Glu Thr Thr Phe Met Cys Glu Tyr Ala Asp Glu Thr Ala 100 105
110Thr Ile Val Glu Phe Leu Asn Arg Trp Ile Thr Phe Cys Gln Ser Ile
115 120 125Ile Ser Thr Leu Thr 130302586PRTHomo Sapien 302Glu Val
Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Arg1 5 10 15Ser
Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Asp Asp Tyr 20 25
30Ala Met His Trp Val Arg Gln Val Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Ser Gly Ile Ser Trp Ile Arg Thr Gly Ile Gly Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile Phe Arg Asp Asn Ala Lys Asn Ser
Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala
Leu Tyr Tyr Cys 85 90 95Ala Lys Asp Met Lys Gly Ser Gly Thr Tyr Gly
Gly Trp Phe Asp Thr 100 105 110Trp Gly Gln Gly Thr Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly 115 120 125Pro Ser Val Phe Pro Leu Ala
Pro Ser Ser Lys Ser Thr Ser Gly Gly 130 135 140Thr Ala Ala Leu Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val145 150 155 160Thr Val
Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe 165 170
175Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val
180 185 190Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys
Asn Val 195 200 205Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys Lys
Val Glu Pro Lys 210 215 220Ser Cys Asp Lys Thr His Thr Cys Pro Pro
Cys Pro Ala Pro Glu Leu225 230 235 240Ala Gly Ala Pro Ser Val Phe
Leu Phe Pro Pro Lys Pro Lys Asp Thr 245 250 255Leu Met Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val Asp Val 260 265 270Ser His Glu
Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val 275 280 285Glu
Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser 290 295
300Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp
Leu305 310 315 320Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Ala Leu Pro Ala 325 330 335Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys
Gly Gln Pro Arg Glu Pro 340 345 350Gln Val Tyr Thr Leu Pro Pro Ser
Arg Asp Glu Leu Thr Lys Asn Gln 355 360 365Val Ser Leu Thr Cys Leu
Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala 370 375 380Val Glu Trp Glu
Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr385 390 395 400Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu 405 410
415Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser
420 425 430Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser
Leu Ser 435 440 445Leu Ser Pro Gly Lys Ala Pro Thr Ser Ser Ser Thr
Lys Lys Thr Gln 450 455 460Leu Gln Leu Glu His Leu Leu Leu Asp Leu
Gln Met Ile Leu Asn Gly465 470 475 480Ile Asn Asn Tyr Lys Asn Pro
Lys Leu Thr Arg Met Leu Thr Phe Lys 485 490 495Phe Tyr Met Pro Lys
Lys Ala Thr Glu Leu Lys His Leu Gln Cys Leu 500 505 510Glu Glu Glu
Leu Lys Pro Leu Glu Glu Val Leu Asn Leu Ala Gln Ser 515 520 525Lys
Asn Phe His Leu Arg Pro Arg Asp Leu Ile Ser Asn Ile Asn Val 530 535
540Ile Val Leu Glu Leu Lys Gly Ser Glu Thr Thr Phe Met Cys Glu
Tyr545 550 555 560Ala Asp Glu Thr Ala Thr Ile Val Glu Phe Leu Asn
Arg Trp Ile Thr 565 570 575Phe Cys Gln Ser Ile Ile Ser Thr Leu Thr
580 58530314PRTHomo Sapien 303Ala Pro Thr Ser Thr Gln Leu Gln Leu
Glu Leu Leu Leu Asp1 5 1030411PRTHomo Sapien 304Thr Gln Leu Gln Leu
Glu His Leu Leu Leu Asp1 5 1030517PRTHomo Sapien 305Ala Pro Thr Ser
Lys Lys Thr Gln Leu Gln Leu Glu His Leu Leu Leu1 5 10
15Asp30619PRTHomo Sapien 306Pro Thr Ser Ser Ser Thr Lys Lys Thr Gln
Leu Gln Leu Glu His Leu1 5 10 15Leu Leu Asp30718PRTHomo Sapien
307Thr Ser Ser Ser Thr Lys Lys Thr Gln Leu Gln Leu Glu His Leu Leu1
5
10 15Leu Asp30817PRTHomo Sapien 308Ser Ser Ser Thr Lys Lys Thr Gln
Leu Gln Leu Glu His Leu Leu Leu1 5 10 15Asp30916PRTHomo Sapien
309Ser Ser Thr Lys Lys Thr Gln Leu Gln Leu Glu His Leu Leu Leu Asp1
5 10 1531015PRTHomo Sapien 310Ser Thr Lys Lys Thr Gln Leu Gln Leu
Glu His Leu Leu Leu Asp1 5 10 1531114PRTHomo Sapien 311Thr Lys Lys
Thr Gln Leu Gln Leu Glu His Leu Leu Leu Asp1 5 1031213PRTHomo
Sapien 312Lys Lys Thr Gln Leu Gln Leu Glu His Leu Leu Leu Asp1 5
1031312PRTHomo Sapien 313Lys Thr Gln Leu Gln Leu Glu His Leu Leu
Leu Asp1 5 1031419PRTHomo Sapien 314Ala Pro Thr Ser Ser Ser Thr Lys
Thr Gln Leu Gln Leu Glu His Leu1 5 10 15Leu Leu Asp31518PRTHomo
Sapien 315Ala Pro Thr Ser Ser Ser Thr Thr Gln Leu Gln Leu Glu His
Leu Leu1 5 10 15Leu Asp31617PRTHomo Sapien 316Ala Pro Thr Ser Ser
Ser Thr Gln Leu Gln Leu Glu His Leu Leu Leu1 5 10 15Asp31716PRTHomo
Sapien 317Ala Pro Thr Ser Ser Thr Gln Leu Gln Leu Glu His Leu Leu
Leu Asp1 5 10 1531814PRTHomo Sapien 318Ala Pro Thr Thr Gln Leu Gln
Leu Glu His Leu Leu Leu Asp1 5 1031913PRTHomo Sapien 319Ala Pro Thr
Gln Leu Gln Leu Glu His Leu Leu Leu Asp1 5 1032012PRTHomo Sapien
320Ala Thr Gln Leu Gln Leu Glu His Leu Leu Leu Asp1 5
1032115PRTHomo Sapien 321Ala Thr Lys Lys Thr Gln Leu Gln Leu Glu
His Leu Leu Leu Asp1 5 10 1532215PRTHomo Sapien 322Ala Pro Lys Lys
Thr Gln Leu Gln Leu Glu His Leu Leu Leu Asp1 5 10 1532315PRTHomo
Sapien 323Ala Pro Thr Lys Thr Gln Leu Gln Leu Glu His Leu Leu Leu
Asp1 5 10 15324113PRTHomo Sapien 324Leu Gln Met Ile Leu Asn Gly Ile
Asn Asn Tyr Lys Asn Pro Lys Leu1 5 10 15Thr Arg Met Leu Thr Phe Lys
Phe Tyr Met Pro Lys Lys Ala Thr Glu 20 25 30Leu Lys His Leu Gln Cys
Leu Glu Glu Glu Leu Lys Pro Leu Glu Glu 35 40 45Val Leu Asn Leu Ala
Gln Ser Lys Asn Phe His Leu Arg Pro Arg Asp 50 55 60Leu Ile Ser Asn
Ile Asn Val Ile Val Leu Glu Leu Lys Gly Ser Glu65 70 75 80Thr Thr
Phe Met Cys Glu Tyr Ala Asp Glu Thr Ala Thr Ile Val Glu 85 90 95Phe
Leu Asn Arg Trp Ile Thr Phe Cys Gln Ser Ile Ile Ser Thr Leu 100 105
110Thr325290PRTMus Musculus 325Met Arg Ile Phe Ala Gly Ile Ile Phe
Thr Ala Cys Cys His Leu Leu1 5 10 15Arg Ala Phe Thr Ile Thr Ala Pro
Lys Asp Leu Tyr Val Val Glu Tyr 20 25 30Gly Ser Asn Val Thr Met Glu
Cys Arg Phe Pro Val Glu Arg Glu Leu 35 40 45Asp Leu Leu Ala Leu Val
Val Tyr Trp Glu Lys Glu Asp Glu Gln Val 50 55 60Ile Gln Phe Val Ala
Gly Glu Glu Asp Leu Lys Pro Gln His Ser Asn65 70 75 80Phe Arg Gly
Arg Ala Ser Leu Pro Lys Asp Gln Leu Leu Lys Gly Asn 85 90 95Ala Ala
Leu Gln Ile Thr Asp Val Lys Leu Gln Asp Ala Gly Val Tyr 100 105
110Cys Cys Ile Ile Ser Tyr Gly Gly Ala Asp Tyr Lys Arg Ile Thr Leu
115 120 125Lys Val Asn Ala Pro Tyr Arg Lys Ile Asn Gln Arg Ile Ser
Val Asp 130 135 140Pro Ala Thr Ser Glu His Glu Leu Ile Cys Gln Ala
Glu Gly Tyr Pro145 150 155 160Glu Ala Glu Val Ile Trp Thr Asn Ser
Asp His Gln Pro Val Ser Gly 165 170 175Lys Arg Ser Val Thr Thr Ser
Arg Thr Glu Gly Met Leu Leu Asn Val 180 185 190Thr Ser Ser Leu Arg
Val Asn Ala Thr Ala Asn Asp Val Phe Tyr Cys 195 200 205Thr Phe Trp
Arg Ser Gln Pro Gly Gln Asn His Thr Ala Glu Leu Ile 210 215 220Ile
Pro Glu Leu Pro Ala Thr His Pro Pro Gln Asn Arg Thr His Trp225 230
235 240Val Leu Leu Gly Ser Ile Leu Leu Phe Leu Ile Val Val Ser Thr
Val 245 250 255Leu Leu Phe Leu Arg Lys Gln Val Arg Met Leu Asp Val
Glu Lys Cys 260 265 270Gly Val Glu Asp Thr Ser Ser Lys Asn Arg Asn
Asp Thr Gln Phe Glu 275 280 285Glu Thr 290326226PRTMus Musculus
326Phe Thr Ile Thr Ala Pro Lys Asp Leu Tyr Val Val Glu Tyr Gly Ser1
5 10 15Asn Val Thr Met Glu Cys Arg Phe Pro Val Glu Arg Glu Leu Asp
Leu 20 25 30Leu Ala Leu Val Val Tyr Trp Glu Lys Glu Asp Glu Gln Val
Ile Gln 35 40 45Phe Val Ala Gly Glu Glu Asp Leu Lys Pro Gln His Ser
Asn Phe Arg 50 55 60Gly Arg Ala Ser Leu Pro Lys Asp Gln Leu Leu Lys
Gly Asn Ala Ala65 70 75 80Leu Gln Ile Thr Asp Val Lys Leu Gln Asp
Ala Gly Val Tyr Cys Cys 85 90 95Ile Ile Ser Tyr Gly Gly Ala Asp Tyr
Lys Arg Ile Thr Leu Lys Val 100 105 110Asn Ala Pro Tyr Arg Lys Ile
Asn Gln Arg Ile Ser Val Asp Pro Ala 115 120 125Thr Ser Glu His Glu
Leu Ile Cys Gln Ala Glu Gly Tyr Pro Glu Ala 130 135 140Glu Val Ile
Trp Thr Asn Ser Asp His Gln Pro Val Ser Gly Lys Arg145 150 155
160Ser Val Thr Thr Ser Arg Thr Glu Gly Met Leu Leu Asn Val Thr Ser
165 170 175Ser Leu Arg Val Asn Ala Thr Ala Asn Asp Val Phe Tyr Cys
Thr Phe 180 185 190Trp Arg Ser Gln Pro Gly Gln Asn His Thr Ala Glu
Leu Ile Ile Pro 195 200 205Glu Leu Pro Ala Thr His Pro Pro Gln Asn
Arg Thr His His His His 210 215 220His His225327251PRTHomo Sapien
327Glu Leu Cys Asp Asp Asp Pro Pro Glu Ile Pro His Ala Thr Phe Lys1
5 10 15Ala Met Ala Tyr Lys Glu Gly Thr Met Leu Asn Cys Glu Cys Lys
Arg 20 25 30Gly Phe Arg Arg Ile Lys Ser Gly Ser Leu Tyr Met Leu Cys
Thr Gly 35 40 45Asn Ser Ser His Ser Ser Trp Asp Asn Gln Cys Gln Cys
Thr Ser Ser 50 55 60Ala Thr Arg Asn Thr Thr Lys Gln Val Thr Pro Gln
Pro Glu Glu Gln65 70 75 80Lys Glu Arg Lys Thr Thr Glu Met Gln Ser
Pro Met Gln Pro Val Asp 85 90 95Gln Ala Ser Leu Pro Gly His Cys Arg
Glu Pro Pro Pro Trp Glu Asn 100 105 110Glu Ala Thr Glu Arg Ile Tyr
His Phe Val Val Gly Gln Met Val Tyr 115 120 125Tyr Gln Cys Val Gln
Gly Tyr Arg Ala Leu His Arg Gly Pro Ala Glu 130 135 140Ser Val Cys
Lys Met Thr His Gly Lys Thr Arg Trp Thr Gln Pro Gln145 150 155
160Leu Ile Cys Thr Gly Glu Met Glu Thr Ser Gln Phe Pro Gly Glu Glu
165 170 175Lys Pro Gln Ala Ser Pro Glu Gly Arg Pro Glu Ser Glu Thr
Ser Cys 180 185 190Leu Val Thr Thr Thr Asp Phe Gln Ile Gln Thr Glu
Met Ala Ala Thr 195 200 205Met Glu Thr Ser Ile Phe Thr Thr Glu Tyr
Gln Val Ala Val Ala Gly 210 215 220Cys Val Phe Leu Leu Ile Ser Val
Leu Leu Leu Ser Gly Leu Thr Trp225 230 235 240Gln Arg Arg Gln Arg
Lys Ser Arg Arg Thr Ile 245 250328525PRTHomo Sapien 328Ala Val Asn
Gly Thr Ser Gln Phe Thr Cys Phe Tyr Asn Ser Arg Ala1 5 10 15Asn Ile
Ser Cys Val Trp Ser Gln Asp Gly Ala Leu Gln Asp Thr Ser 20 25 30Cys
Gln Val His Ala Trp Pro Asp Arg Arg Arg Trp Asn Gln Thr Cys 35 40
45Glu Leu Leu Pro Val Ser Gln Ala Ser Trp Ala Cys Asn Leu Ile Leu
50 55 60Gly Ala Pro Asp Ser Gln Lys Leu Thr Thr Val Asp Ile Val Thr
Leu65 70 75 80Arg Val Leu Cys Arg Glu Gly Val Arg Trp Arg Val Met
Ala Ile Gln 85 90 95Asp Phe Lys Pro Phe Glu Asn Leu Arg Leu Met Ala
Pro Ile Ser Leu 100 105 110Gln Val Val His Val Glu Thr His Arg Cys
Asn Ile Ser Trp Glu Ile 115 120 125Ser Gln Ala Ser His Tyr Phe Glu
Arg His Leu Glu Phe Glu Ala Arg 130 135 140Thr Leu Ser Pro Gly His
Thr Trp Glu Glu Ala Pro Leu Leu Thr Leu145 150 155 160Lys Gln Lys
Gln Glu Trp Ile Cys Leu Glu Thr Leu Thr Pro Asp Thr 165 170 175Gln
Tyr Glu Phe Gln Val Arg Val Lys Pro Leu Gln Gly Glu Phe Thr 180 185
190Thr Trp Ser Pro Trp Ser Gln Pro Leu Ala Phe Arg Thr Lys Pro Ala
195 200 205Ala Leu Gly Lys Asp Thr Ile Pro Trp Leu Gly His Leu Leu
Val Gly 210 215 220Leu Ser Gly Ala Phe Gly Phe Ile Ile Leu Val Tyr
Leu Leu Ile Asn225 230 235 240Cys Arg Asn Thr Gly Pro Trp Leu Lys
Lys Val Leu Lys Cys Asn Thr 245 250 255Pro Asp Pro Ser Lys Phe Phe
Ser Gln Leu Ser Ser Glu His Gly Gly 260 265 270Asp Val Gln Lys Trp
Leu Ser Ser Pro Phe Pro Ser Ser Ser Phe Ser 275 280 285Pro Gly Gly
Leu Ala Pro Glu Ile Ser Pro Leu Glu Val Leu Glu Arg 290 295 300Asp
Lys Val Thr Gln Leu Leu Leu Gln Gln Asp Lys Val Pro Glu Pro305 310
315 320Ala Ser Leu Ser Ser Asn His Ser Leu Thr Ser Cys Phe Thr Asn
Gln 325 330 335Gly Tyr Phe Phe Phe His Leu Pro Asp Ala Leu Glu Ile
Glu Ala Cys 340 345 350Gln Val Tyr Phe Thr Tyr Asp Pro Tyr Ser Glu
Glu Asp Pro Asp Glu 355 360 365Gly Val Ala Gly Ala Pro Thr Gly Ser
Ser Pro Gln Pro Leu Gln Pro 370 375 380Leu Ser Gly Glu Asp Asp Ala
Tyr Cys Thr Phe Pro Ser Arg Asp Asp385 390 395 400Leu Leu Leu Phe
Ser Pro Ser Leu Leu Gly Gly Pro Ser Pro Pro Ser 405 410 415Thr Ala
Pro Gly Gly Ser Gly Ala Gly Glu Glu Arg Met Pro Pro Ser 420 425
430Leu Gln Glu Arg Val Pro Arg Asp Trp Asp Pro Gln Pro Leu Gly Pro
435 440 445Pro Thr Pro Gly Val Pro Asp Leu Val Asp Phe Gln Pro Pro
Pro Glu 450 455 460Leu Val Leu Arg Glu Ala Gly Glu Glu Val Pro Asp
Ala Gly Pro Arg465 470 475 480Glu Gly Val Ser Phe Pro Trp Ser Arg
Pro Pro Gly Gln Gly Glu Phe 485 490 495Arg Ala Leu Asn Ala Arg Leu
Pro Leu Asn Thr Asp Ala Tyr Leu Ser 500 505 510Leu Gln Glu Leu Gln
Gly Gln Asp Pro Thr His Leu Val 515 520 525329347PRTHomo Sapien
329Leu Asn Thr Thr Ile Leu Thr Pro Asn Gly Asn Glu Asp Thr Thr Ala1
5 10 15Asp Phe Phe Leu Thr Thr Met Pro Thr Asp Ser Leu Ser Val Ser
Thr 20 25 30Leu Pro Leu Pro Glu Val Gln Cys Phe Val Phe Asn Val Glu
Tyr Met 35 40 45Asn Cys Thr Trp Asn Ser Ser Ser Glu Pro Gln Pro Thr
Asn Leu Thr 50 55 60Leu His Tyr Trp Tyr Lys Asn Ser Asp Asn Asp Lys
Val Gln Lys Cys65 70 75 80Ser His Tyr Leu Phe Ser Glu Glu Ile Thr
Ser Gly Cys Gln Leu Gln 85 90 95Lys Lys Glu Ile His Leu Tyr Gln Thr
Phe Val Val Gln Leu Gln Asp 100 105 110Pro Arg Glu Pro Arg Arg Gln
Ala Thr Gln Met Leu Lys Leu Gln Asn 115 120 125Leu Val Ile Pro Trp
Ala Pro Glu Asn Leu Thr Leu His Lys Leu Ser 130 135 140Glu Ser Gln
Leu Glu Leu Asn Trp Asn Asn Arg Phe Leu Asn His Cys145 150 155
160Leu Glu His Leu Val Gln Tyr Arg Thr Asp Trp Asp His Ser Trp Thr
165 170 175Glu Gln Ser Val Asp Tyr Arg His Lys Phe Ser Leu Pro Ser
Val Asp 180 185 190Gly Gln Lys Arg Tyr Thr Phe Arg Val Arg Ser Arg
Phe Asn Pro Leu 195 200 205Cys Gly Ser Ala Gln His Trp Ser Glu Trp
Ser His Pro Ile His Trp 210 215 220Gly Ser Asn Thr Ser Lys Glu Asn
Pro Phe Leu Phe Ala Leu Glu Ala225 230 235 240Val Val Ile Ser Val
Gly Ser Met Gly Leu Ile Ile Ser Leu Leu Cys 245 250 255Val Tyr Phe
Trp Leu Glu Arg Thr Met Pro Arg Ile Pro Thr Leu Lys 260 265 270Asn
Leu Glu Asp Leu Val Thr Glu Tyr His Gly Asn Phe Ser Ala Trp 275 280
285Ser Gly Val Ser Lys Gly Leu Ala Glu Ser Leu Gln Pro Asp Tyr Ser
290 295 300Glu Arg Leu Cys Leu Val Ser Glu Ile Pro Pro Lys Gly Gly
Ala Leu305 310 315 320Gly Glu Gly Pro Gly Ala Ser Pro Cys Asn Gln
His Ser Pro Tyr Trp 325 330 335Ala Pro Pro Cys Tyr Thr Leu Lys Pro
Glu Thr 340 345330152PRTHomo Sapien 330Asp Cys Asp Ile Glu Gly Lys
Asp Gly Lys Gln Tyr Glu Ser Val Leu1 5 10 15Met Val Ser Ile Asp Gln
Leu Leu Asp Ser Met Lys Glu Ile Gly Ser 20 25 30Asn Cys Leu Asn Asn
Glu Phe Asn Phe Phe Lys Arg His Ile Cys Asp 35 40 45Ala Asn Lys Glu
Gly Met Phe Leu Phe Arg Ala Ala Arg Lys Leu Arg 50 55 60Gln Phe Leu
Lys Met Asn Ser Thr Gly Asp Phe Asp Leu His Leu Leu65 70 75 80Lys
Val Ser Glu Gly Thr Thr Ile Leu Leu Asn Cys Thr Gly Gln Val 85 90
95Lys Gly Arg Lys Pro Ala Ala Leu Gly Glu Ala Gln Pro Thr Lys Ser
100 105 110Leu Glu Glu Asn Lys Ser Leu Lys Glu Gln Lys Lys Leu Asn
Asp Leu 115 120 125Cys Phe Leu Lys Arg Leu Leu Gln Glu Ile Lys Thr
Cys Trp Asn Lys 130 135 140Ile Leu Met Gly Thr Lys Glu His145
150331133PRTHomo Sapien 331Gly Ile His Val Phe Ile Leu Gly Cys Phe
Ser Ala Gly Leu Pro Lys1 5 10 15Thr Glu Ala Asn Trp Val Asn Val Ile
Ser Asp Leu Lys Lys Ile Glu 20 25 30Asp Leu Ile Gln Ser Met His Ile
Asp Ala Thr Leu Tyr Thr Glu Ser 35 40 45Asp Val His Pro Ser Cys Lys
Val Thr Ala Met Lys Cys Phe Leu Leu 50 55 60Glu Leu Gln Val Ile Ser
Leu Glu Ser Gly Asp Ala Ser Ile His Asp65 70 75 80Thr Val Glu Asn
Leu Ile Ile Leu Ala Asn Asn Ser Leu Ser Ser Asn 85 90 95Gly Asn Val
Thr Glu Ser Gly Cys Lys Glu Cys Glu Glu Leu Glu Glu 100 105 110Lys
Asn Ile Lys Glu Phe Leu Gln Ser Phe Val His Ile Val Gln Met 115 120
125Phe Ile Asn Thr Ser 130332133PRTHomo Sapien 332Gln Gly Gln Asp
Arg His Met Ile Arg Met Arg Gln Leu Ile Asp Ile1 5 10 15Val Asp Gln
Leu Lys Asn Tyr Val Asn Asp Leu Val Pro Glu Phe Leu 20 25 30Pro Ala
Pro Glu Asp Val Glu Thr Asn Cys Glu Trp Ser Ala Phe Ser 35 40 45Cys
Phe Gln Lys Ala Gln Leu Lys Ser Ala Asn Thr Gly Asn Asn Glu 50 55
60Arg Ile Ile Asn Val Ser Ile Lys Lys Leu Lys Arg Lys Pro Pro Ser65
70 75 80Thr Asn Ala Gly Arg Arg Gln Lys His Arg Leu Thr Cys Pro Ser
Cys 85 90 95Asp Ser Tyr Glu Lys Lys Pro Pro Lys Glu Phe Leu Glu Arg
Phe Lys 100 105 110Ser Leu Leu Gln Lys Met Ile His Gln His Leu Ser
Ser Arg Thr His 115
120 125Gly Ser Glu Asp Ser 130333127PRTHomo Sapien 333Ala Pro Ala
Arg Ser Pro Ser Pro Ser Thr Gln Pro Trp Glu His Val1 5 10 15Asn Ala
Ile Gln Glu Ala Arg Arg Leu Leu Asn Leu Ser Arg Asp Thr 20 25 30Ala
Ala Glu Met Asn Glu Thr Val Glu Val Ile Ser Glu Met Phe Asp 35 40
45Leu Gln Glu Pro Thr Cys Leu Gln Thr Arg Leu Glu Leu Tyr Lys Gln
50 55 60Gly Leu Arg Gly Ser Leu Thr Lys Leu Lys Gly Pro Leu Thr Met
Met65 70 75 80Ala Ser His Tyr Lys Gln His Cys Pro Pro Thr Pro Glu
Thr Ser Cys 85 90 95Ala Thr Gln Ile Ile Thr Phe Glu Ser Phe Lys Glu
Asn Leu Lys Asp 100 105 110Phe Leu Leu Val Ile Pro Phe Asp Cys Trp
Glu Pro Val Gln Glu 115 120 125334171PRTHomo Sapien 334Cys Asp Leu
Pro Gln Asn His Gly Leu Leu Ser Arg Asn Thr Leu Val1 5 10 15Leu Leu
His Gln Met Arg Arg Ile Ser Pro Phe Leu Cys Leu Lys Asp 20 25 30Arg
Arg Asp Phe Arg Phe Pro Gln Glu Met Val Lys Gly Ser Gln Leu 35 40
45Gln Lys Ala His Val Met Ser Val Leu His Glu Met Leu Gln Gln Ile
50 55 60Phe Ser Leu Phe His Thr Glu Arg Ser Ser Ala Ala Trp Asn Met
Thr65 70 75 80Leu Leu Asp Gln Leu His Thr Glu Leu His Gln Gln Leu
Gln His Leu 85 90 95Glu Thr Cys Leu Leu Gln Val Val Gly Glu Gly Glu
Ser Ala Gly Ala 100 105 110Ile Ser Ser Pro Ala Leu Thr Leu Arg Arg
Tyr Phe Gln Gly Ile Arg 115 120 125Val Tyr Leu Lys Glu Lys Lys Tyr
Ser Asp Cys Ala Trp Glu Val Val 130 135 140Arg Met Glu Ile Met Lys
Ser Leu Phe Leu Ser Thr Asn Met Gln Glu145 150 155 160Arg Leu Arg
Ser Lys Asp Arg Asp Leu Gly Ser 165 170335177PRTHomo Sapien 335Gly
Pro Gln Arg Glu Glu Phe Pro Arg Asp Leu Ser Leu Ile Ser Pro1 5 10
15Leu Ala Gln Ala Val Arg Ser Ser Ser Arg Thr Pro Ser Asp Lys Pro
20 25 30Val Ala His Val Val Ala Asn Pro Gln Ala Glu Gly Gln Leu Gln
Trp 35 40 45Leu Asn Arg Arg Ala Asn Ala Leu Leu Ala Asn Gly Val Glu
Leu Arg 50 55 60Asp Asn Gln Leu Val Val Pro Ser Glu Gly Leu Tyr Leu
Ile Tyr Ser65 70 75 80Gln Val Leu Phe Lys Gly Gln Gly Cys Pro Ser
Thr His Val Leu Leu 85 90 95Thr His Thr Ile Ser Arg Ile Ala Val Ser
Tyr Gln Thr Lys Val Asn 100 105 110Leu Leu Ser Ala Ile Lys Ser Pro
Cys Gln Arg Glu Thr Pro Glu Gly 115 120 125Ala Glu Ala Lys Pro Trp
Tyr Glu Pro Ile Tyr Leu Gly Gly Val Phe 130 135 140Gln Leu Glu Lys
Gly Asp Arg Leu Ser Ala Glu Ile Asn Arg Pro Asp145 150 155 160Tyr
Leu Asp Phe Ala Glu Ser Gly Gln Val Tyr Phe Gly Ile Ile Ala 165 170
175Leu336197PRTHomo Sapien 336Arg Asn Leu Pro Val Ala Thr Pro Asp
Pro Gly Met Phe Pro Cys Leu1 5 10 15His His Ser Gln Asn Leu Leu Arg
Ala Val Ser Asn Met Leu Gln Lys 20 25 30Ala Arg Gln Thr Leu Glu Phe
Tyr Pro Cys Thr Ser Glu Glu Ile Asp 35 40 45His Glu Asp Ile Thr Lys
Asp Lys Thr Ser Thr Val Glu Ala Cys Leu 50 55 60Pro Leu Glu Leu Thr
Lys Asn Glu Ser Cys Leu Asn Ser Arg Glu Thr65 70 75 80Ser Phe Ile
Thr Asn Gly Ser Cys Leu Ala Ser Arg Lys Thr Ser Phe 85 90 95Met Met
Ala Leu Cys Leu Ser Ser Ile Tyr Glu Asp Leu Lys Met Tyr 100 105
110Gln Val Glu Phe Lys Thr Met Asn Ala Lys Leu Leu Met Asp Pro Lys
115 120 125Arg Gln Ile Phe Leu Asp Gln Asn Met Leu Ala Val Ile Asp
Glu Leu 130 135 140Met Gln Ala Leu Asn Phe Asn Ser Glu Thr Val Pro
Gln Lys Ser Ser145 150 155 160Leu Glu Glu Pro Asp Phe Tyr Lys Thr
Lys Ile Lys Leu Cys Ile Leu 165 170 175Leu His Ala Phe Arg Ile Arg
Ala Val Thr Ile Asp Arg Val Met Ser 180 185 190Tyr Leu Asn Ala Ser
195337306PRTHomo Sapien 337Ile Trp Glu Leu Lys Lys Asp Val Tyr Val
Val Glu Leu Asp Trp Tyr1 5 10 15Pro Asp Ala Pro Gly Glu Met Val Val
Leu Thr Cys Asp Thr Pro Glu 20 25 30Glu Asp Gly Ile Thr Trp Thr Leu
Asp Gln Ser Ser Glu Val Leu Gly 35 40 45Ser Gly Lys Thr Leu Thr Ile
Gln Val Lys Glu Phe Gly Asp Ala Gly 50 55 60Gln Tyr Thr Cys His Lys
Gly Gly Glu Val Leu Ser His Ser Leu Leu65 70 75 80Leu Leu His Lys
Lys Glu Asp Gly Ile Trp Ser Thr Asp Ile Leu Lys 85 90 95Asp Gln Lys
Glu Pro Lys Asn Lys Thr Phe Leu Arg Cys Glu Ala Lys 100 105 110Asn
Tyr Ser Gly Arg Phe Thr Cys Trp Trp Leu Thr Thr Ile Ser Thr 115 120
125Asp Leu Thr Phe Ser Val Lys Ser Ser Arg Gly Ser Ser Asp Pro Gln
130 135 140Gly Val Thr Cys Gly Ala Ala Thr Leu Ser Ala Glu Arg Val
Arg Gly145 150 155 160Asp Asn Lys Glu Tyr Glu Tyr Ser Val Glu Cys
Gln Glu Asp Ser Ala 165 170 175Cys Pro Ala Ala Glu Glu Ser Leu Pro
Ile Glu Val Met Val Asp Ala 180 185 190Val His Lys Leu Lys Tyr Glu
Asn Tyr Thr Ser Ser Phe Phe Ile Arg 195 200 205Asp Ile Ile Lys Pro
Asp Pro Pro Lys Asn Leu Gln Leu Lys Pro Leu 210 215 220Lys Asn Ser
Arg Gln Val Glu Val Ser Trp Glu Tyr Pro Asp Thr Trp225 230 235
240Ser Thr Pro His Ser Tyr Phe Ser Leu Thr Phe Cys Val Gln Val Gln
245 250 255Gly Lys Ser Lys Arg Glu Lys Lys Asp Arg Val Phe Thr Asp
Lys Thr 260 265 270Ser Ala Thr Val Ile Cys Arg Lys Asn Ala Ser Ile
Ser Val Arg Ala 275 280 285Gln Asp Arg Tyr Tyr Ser Ser Ser Trp Ser
Glu Trp Ala Ser Val Pro 290 295 300Cys Ser305338103PRTHomo Sapien
338Thr Pro Val Val Arg Lys Gly Arg Cys Ser Cys Ile Ser Thr Asn Gln1
5 10 15Gly Thr Ile His Leu Gln Ser Leu Lys Asp Leu Lys Gln Phe Ala
Pro 20 25 30Ser Pro Ser Cys Glu Lys Ile Glu Ile Ile Ala Thr Leu Lys
Asn Gly 35 40 45Val Gln Thr Cys Leu Asn Pro Asp Ser Ala Asp Val Lys
Glu Leu Ile 50 55 60Lys Lys Trp Glu Lys Gln Val Ser Gln Lys Lys Lys
Gln Lys Asn Gly65 70 75 80Lys Lys His Gln Lys Lys Lys Val Leu Lys
Val Arg Lys Ser Gln Arg 85 90 95Ser Arg Gln Lys Lys Thr Thr
10033977PRTHomo Sapien 339Val Pro Leu Ser Arg Thr Val Arg Cys Thr
Cys Ile Ser Ile Ser Asn1 5 10 15Gln Pro Val Asn Pro Arg Ser Leu Glu
Lys Leu Glu Ile Ile Pro Ala 20 25 30Ser Gln Phe Cys Pro Arg Val Glu
Ile Ile Ala Thr Met Lys Lys Lys 35 40 45Gly Glu Lys Arg Cys Leu Asn
Pro Glu Ser Lys Ala Ile Lys Asn Leu 50 55 60Leu Lys Ala Val Ser Lys
Glu Arg Ser Lys Arg Ser Pro65 70 75340330PRTHomo Sapien 340Ala Ser
Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys1 5 10 15Ser
Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25
30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser
35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr
Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr
Gln Thr65 70 75 80Tyr Ile Cys Asn Val Asn His Lys Pro Ser Asn Thr
Lys Val Asp Lys 85 90 95Lys Val Glu Pro Lys Ser Cys Asp Lys Thr His
Thr Cys Pro Pro Cys 100 105 110Pro Ala Pro Glu Leu Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro Pro 115 120 125Lys Pro Lys Asp Thr Leu Met
Ile Ser Arg Thr Pro Glu Val Thr Cys 130 135 140Val Val Val Asp Val
Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp145 150 155 160Tyr Val
Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu 165 170
175Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
180 185 190His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser Asn 195 200 205Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser
Lys Ala Lys Gly 210 215 220Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu
Pro Pro Ser Arg Asp Glu225 230 235 240Leu Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr 245 250 255Pro Ser Asp Ile Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn 260 265 270Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe 275 280 285Leu
Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn 290 295
300Val Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn His Tyr
Thr305 310 315 320Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 325
330341996DNAHomo Sapien 341gccagcacca agggcccctc tgtgttccct
ctggcccctt ccagcaagtc cacctctggc 60ggaacagccg ctctgggctg cctcgtgaag
gactacttcc ccgagcctgt gaccgtgtcc 120tggaactctg gcgctctgac
cagcggagtg cacaccttcc ctgctgtgct gcagtcctcc 180ggcctgtact
ccctgtcctc cgtcgtgacc gtgccttcca gctctctggg cacccagacc
240tacatctgca acgtgaacca caagccctcc aacaccaagg tggacaagaa
ggtggaaccc 300aagtcctgcg acaagaccca cacctgtccc ccttgtcctg
cccctgaact gctgggcgga 360ccttccgtgt tcctgttccc cccaaagccc
aaggacaccc tgatgatctc ccggaccccc 420gaagtgacct gcgtggtggt
ggatgtgtcc cacgaggacc ctgaagtgaa gttcaattgg 480tacgtggacg
gcgtggaagt gcacaacgcc aagaccaagc ctagagagga acagtacaac
540tccacctacc gggtggtgtc cgtgctgacc gtgctgcacc aggattggct
gaacggcaaa 600gagtacaagt gcaaggtgtc caacaaggcc ctgcctgccc
ccatcgaaaa gaccatctcc 660aaggccaagg gccagccccg ggaaccccag
gtgtacacac tgccccctag cagggacgag 720ctgaccaaga accaggtgtc
cctgacctgt ctcgtgaaag gcttctaccc ctccgatatc 780gccgtggaat
gggagtccaa cggccagcct gagaacaact acaagaccac cccccctgtg
840ctggactccg acggctcatt cttcctgtac agcaagctga cagtggacaa
gtcccggtgg 900cagcagggca acgtgttctc ctgctccgtg atgcacgagg
ccctgcacaa ccactacacc 960cagaagtccc tgtccctgag ccccggcaag tgatga
996342450PRTHomo Sapien 342Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Arg1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Asp Asp Tyr 20 25 30Ala Met His Trp Val Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile Ser Trp Ile Arg
Thr Gly Ile Gly Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn
Ser Leu Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Lys Asp
Met Lys Gly Ser Gly Thr Tyr Gly Gly Trp Phe Asp Thr 100 105 110Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly 115 120
125Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser
130 135 140Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu
Pro Val145 150 155 160Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser
Gly Val His Thr Phe 165 170 175Pro Ala Val Leu Gln Ser Ser Gly Leu
Tyr Ser Leu Ser Ser Val Val 180 185 190Thr Val Pro Ser Ser Ser Leu
Gly Thr Lys Thr Tyr Thr Cys Asn Val 195 200 205Asp His Lys Pro Ser
Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys 210 215 220Tyr Gly Pro
Pro Cys Pro Pro Cys Pro Ala Pro Glu Leu Ala Gly Ala225 230 235
240Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
245 250 255Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser
Gln Glu 260 265 270Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly
Val Glu Val His 275 280 285Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Phe Asn Ser Thr Tyr Arg 290 295 300Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu Asn Gly Lys305 310 315 320Glu Tyr Lys Cys Lys
Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu 325 330 335Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr 340 345 350Thr
Leu Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu 355 360
365Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp
370 375 380Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro
Pro Val385 390 395 400Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Arg Leu Thr Val Asp 405 410 415Lys Ser Arg Trp Gln Glu Gly Asn Val
Phe Ser Cys Ser Val Met His 420 425 430Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu Ser Leu Ser Leu 435 440 445Gly Lys
4503438PRTHomo Sapien 343Gly Phe Thr Phe Ser Asn Tyr Ala1
53448PRTHomo Sapien 344Ile Ser Phe Ser Gly Gly Thr Thr1
534514PRTHomo Sapien 345Ala Lys Asp Glu Ala Pro Ala Gly Ala Thr Phe
Phe Asp Ser1 5 103465PRTHomo Sapien 346Asn Tyr Ala Met Ser1
534717PRTHomo Sapien 347Ala Ile Ser Phe Ser Gly Gly Thr Thr Tyr Tyr
Ala Asp Ser Val Lys1 5 10 15Gly34812PRTHomo Sapien 348Asp Glu Ala
Pro Ala Gly Ala Thr Phe Phe Asp Ser1 5 10349121PRTHomo Sapien
349Glu Val Gln Leu Ala Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn
Tyr 20 25 30Ala Met Ser Trp Val Arg Gln Thr Pro Gly Lys Gly Leu Glu
Trp Val 35 40 45Ser Ala Ile Ser Phe Ser Gly Gly Thr Thr Tyr Tyr Ala
Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys
Asn Thr Leu Tyr65 70 75 80Leu His Met Asn Ser Leu Arg Ala Asp Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Lys Asp Glu Ala Pro Ala Gly Ala
Thr Phe Phe Asp Ser Trp Gly 100 105 110Gln Gly Thr Leu Val Thr Val
Ser Ser 115 120350364DNAHomo Sapien 350gaagtgcaac tggcggagtc
tgggggaggc ttggtacagc cgggggggtc cctgagactc 60tcctgtgcag cctctggatt
cacctttagc aactatgcca tgagttgggt ccgccagact 120ccaggaaagg
ggctggagtg ggtctcagct attagtttta gtggtggtac tacatactac
180gctgactccg tgaagggccg gttcaccatc tccagagaca attccaagaa
cacgctgtat 240ttgcacatga acagcctgag agccgatgac acggccgtat
attactgtgc gaaagatgag 300gcaccagctg gcgcaacctt ctttgactcc
tggggccagg gaacgctggt caccgtctcc 360tcag 364351448PRTHomo Sapien
351Glu Val Gln Leu Ala Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn
Tyr 20 25 30Ala Met Ser Trp Val Arg Gln Thr Pro Gly Lys Gly Leu Glu
Trp Val 35 40 45Ser Ala Ile Ser Phe Ser Gly Gly Thr Thr Tyr Tyr Ala
Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr
Leu Tyr65 70 75 80Leu His Met Asn Ser Leu Arg Ala Asp Asp Thr Ala
Val Tyr Tyr Cys 85 90 95Ala Lys Asp Glu Ala Pro Ala Gly Ala Thr Phe
Phe Asp Ser Trp Gly 100 105 110Gln Gly Thr Leu Val Thr Val Ser Ser
Ala Ser Thr Lys Gly Pro Ser 115 120 125Val Phe Pro Leu Ala Pro Cys
Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140Ala Leu Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val145 150 155 160Ser Trp
Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170
175Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
180 185 190Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val
Asp His 195 200 205Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu
Ser Lys Tyr Gly 210 215 220Pro Pro Cys Pro Pro Cys Pro Ala Pro Glu
Phe Glu Gly Gly Pro Ser225 230 235 240Val Phe Leu Phe Pro Pro Lys
Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255Thr Pro Glu Val Thr
Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270Glu Val Gln
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285Lys
Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290 295
300Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr305 310 315 320Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser
Ile Glu Lys Thr 325 330 335Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu 340 345 350Pro Pro Ser Gln Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys 355 360 365Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp385 390 395 400Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410
415Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala
420 425 430Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu
Gly Lys 435 440 4453521344DNAHomo Sapien 352gaagtgcaac tggcggagtc
tgggggaggc ttggtacagc cgggggggtc cctgagactc 60tcctgtgcag cctctggatt
cacctttagc aactatgcca tgagttgggt ccgccagact 120ccaggaaagg
ggctggagtg ggtctcagct attagtttta gtggtggtac tacatactac
180gctgactccg tgaagggccg gttcaccatc tccagagaca attccaagaa
cacgctgtat 240ttgcacatga acagcctgag agccgatgac acggccgtat
attactgtgc gaaagatgag 300gcaccagctg gcgcaacctt ctttgactcc
tggggccagg gaacgctggt caccgtctcc 360tcagccagca ccaagggccc
ttccgtgttc cccctggccc cttgcagcag gagcacctcc 420gaatccacag
ctgccctggg ctgtctggtg aaggactact ttcccgagcc cgtgaccgtg
480agctggaaca gcggcgctct gacatccggc gtccacacct ttcctgccgt
cctgcagtcc 540tccggcctct actccctgtc ctccgtggtg accgtgccta
gctcctccct cggcaccaag 600acctacacct gtaacgtgga ccacaaaccc
tccaacacca aggtggacaa acgggtcgag 660agcaagtacg gccctccctg
ccctccttgt cctgcccccg agttcgaagg cggacccagc 720gtgttcctgt
tccctcctaa gcccaaggac accctcatga tcagccggac acccgaggtg
780acctgcgtgg tggtggatgt gagccaggag gaccctgagg tccagttcaa
ctggtatgtg 840gatggcgtgg aggtgcacaa cgccaagaca aagccccggg
aagagcagtt caactccacc 900tacagggtgg tcagcgtgct gaccgtgctg
catcaggact ggctgaacgg caaggagtac 960aagtgcaagg tcagcaataa
gggactgccc agcagcatcg agaagaccat ctccaaggct 1020aaaggccagc
cccgggaacc tcaggtgtac accctgcctc ccagccagga ggagatgacc
1080aagaaccagg tgagcctgac ctgcctggtg aagggattct acccttccga
catcgccgtg 1140gagtgggagt ccaacggcca gcccgagaac aattataaga
ccacccctcc cgtcctcgac 1200agcgacggat ccttctttct gtactccagg
ctgaccgtgg ataagtccag gtggcaggaa 1260ggcaacgtgt tcagctgctc
cgtgatgcac gaggccctgc acaatcacta cacccagaag 1320tccctgagcc
tgtccctggg aaag 13443536PRTHomo Sapien 353Gln Gly Ile Arg Arg Trp1
53543PRTHomo Sapien 354Gly Ala Ser13559PRTHomo Sapien 355Gln Gln
Ala Asn Ser Phe Pro Ile Thr1 535611PRTHomo Sapien 356Arg Ala Ser
Gln Gly Ile Arg Arg Trp Leu Ala1 5 103577PRTHomo Sapien 357Gly Ala
Ser Ser Leu Gln Ser1 53589PRTHomo Sapien 358Gln Gln Ala Asn Ser Phe
Pro Ile Thr1 5359107PRTHomo Sapien 359Asp Ile Gln Met Thr Gln Ser
Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr
Cys Arg Ala Ser Gln Gly Ile Arg Arg Trp 20 25 30Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Ser Gly Ala Ser
Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser
Gly Thr Asp Phe Thr Leu Ile Ile Thr Ser Leu Gln Pro65 70 75 80Glu
Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Phe Pro Ile 85 90
95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys 100 105360322DNAHomo
Sapien 360gacatccaga tgacccagtc tccatcttcc gtgtctgcat ctgtaggaga
cagagtcacc 60atcacttgtc gggcgagtca gggtattagg aggtggttag cctggtatca
gcagaaacca 120gggaaagccc ctaaactcct gatctctggt gcatccagtt
tgcaaagtgg ggtcccatca 180aggttcagcg gcagtggatc tgggacagat
ttcactctca tcattaccag tctgcagcct 240gaagattttg caacttacta
ttgtcaacag gctaacagtt tcccgatcac cttcggccaa 300gggacacgac
tggagatcaa ac 322361214PRTHomo Sapien 361Asp Ile Gln Met Thr Gln
Ser Pro Ser Ser Val Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Gln Gly Ile Arg Arg Trp 20 25 30Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Ser Gly Ala
Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly
Ser Gly Thr Asp Phe Thr Leu Ile Ile Thr Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ala Asn Ser Phe Pro Ile
85 90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys Arg Thr Val Ala
Ala 100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu
Lys Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200
205Phe Asn Arg Gly Glu Cys 210362642DNAHomo Sapien 362gacatccaga
tgacccagtc tccatcttcc gtgtctgcat ctgtaggaga cagagtcacc 60atcacttgtc
gggcgagtca gggtattagg aggtggttag cctggtatca gcagaaacca
120gggaaagccc ctaaactcct gatctctggt gcatccagtt tgcaaagtgg
ggtcccatca 180aggttcagcg gcagtggatc tgggacagat ttcactctca
tcattaccag tctgcagcct 240gaagattttg caacttacta ttgtcaacag
gctaacagtt tcccgatcac cttcggccaa 300gggacacgac tggagatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc tgcagtccgg
caactcccag 480gaatccgtga ccgagcagga ctccaaggac agcacctact
ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga gaagcacaag
gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc ccgtgaccaa
gtctttcaac cggggcgagt gt 6423638PRTHomo Sapien 363Gly Tyr Thr Phe
Ser Thr Phe Gly1 53648PRTHomo Sapien 364Ile Ser Ala Tyr Asn Gly Asp
Thr1 536514PRTHomo Sapien 365Ala Arg Ser Ser Gly His Tyr Tyr Tyr
Tyr Gly Met Asp Val1 5 10366121PRTHomo Sapien 366Gln Val Gln Val
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys
Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Ser Thr Phe 20 25 30Gly Ile
Thr Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly
Trp Ile Ser Ala Tyr Asn Gly Asp Thr Asn Tyr Ala Gln Asn Leu 50 55
60Gln Gly Arg Val Ile Met Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65
70 75 80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr
Cys 85 90 95Ala Arg Ser Ser Gly His Tyr Tyr Tyr Tyr Gly Met Asp Val
Trp Gly 100 105 110Gln Gly Thr Thr Val Thr Val Ser Ser 115
120367363DNAHomo Sapien 367caggttcagg tggtgcagtc tggagctgag
gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta caccttttcc
acctttggta tcacctgggt gcgacaggcc 120cctggacaag ggcttgaatg
gatgggatgg atcagcgctt acaatggtga cacaaactat 180gcacagaatc
tccagggcag agtcatcatg accacagaca catccacgag cacagcctac
240atggagctga ggagcctgag atctgacgac acggccgttt attactgtgc
gaggagcagt 300ggccactact actactacgg tatggacgtc tggggccaag
ggaccacggt caccgtctcc 360tca 363368451PRTHomo Sapien 368Gln Val Gln
Val Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Ser Thr Phe 20 25 30Gly
Ile Thr Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Ser Ala Tyr Asn Gly Asp Thr Asn Tyr Ala Gln Asn Leu
50 55 60Gln Gly Arg Val Ile Met Thr Thr Asp Thr Ser Thr Ser Thr Ala
Tyr65 70 75 80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr Ala Val
Tyr Tyr Cys 85 90 95Ala Arg Ser Ser Gly His Tyr Tyr Tyr Tyr Gly Met
Asp Val Trp Gly 100 105 110Gln Gly Thr Thr Val Thr Val Ser Ser Ala
Ser Thr Lys Gly Pro Ser 115 120 125Val Phe Pro Leu Ala Pro Ser Ser
Lys Ser Thr Ser Gly Gly Thr Ala 130 135 140Ala Leu Gly Cys Leu Val
Lys Asp Tyr Phe Pro Glu Pro Val Thr Val145 150 155 160Ser Trp Asn
Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170 175Val
Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val 180 185
190Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His
195 200 205Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys
Ser Cys 210 215 220Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro
Glu Leu Leu Gly225 230 235 240Gly Pro Ser Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met 245 250 255Ile Ser Arg Thr Pro Glu Val
Thr Cys Val Val Val Asp Val Ser His 260 265 270Glu Asp Pro Glu Val
Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val 275 280 285His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr 290 295 300Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly305 310
315 320Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro
Ile 325 330 335Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val 340 345 350Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr
Lys Asn Gln Val Ser 355 360 365Leu Thr Cys Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu 370 375 380Trp Glu Ser Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro385 390 395 400Val Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val 405 410 415Asp Lys
Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met 420 425
430His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
435 440 445Pro Gly Lys 4503691359DNAHomo Sapien 369caggttcagg
tggtgcagtc tggagctgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg
cttctggtta caccttttcc acctttggta tcacctgggt gcgacaggcc
120cctggacaag ggcttgaatg gatgggatgg atcagcgctt acaatggtga
cacaaactat 180gcacagaatc tccagggcag agtcatcatg accacagaca
catccacgag cacagcctac 240atggagctga ggagcctgag atctgacgac
acggccgttt attactgtgc gaggagcagt 300ggccactact actactacgg
tatggacgtc tggggccaag ggaccacggt caccgtctcc 360tcagccagca
ccaagggccc ctctgtgttc cctctggccc cttccagcaa gtccacctct
420ggcggaacag ccgctctggg ctgcctcgtg aaggactact tccccgagcc
tgtgaccgtg 480tcctggaact ctggcgctct gaccagcgga gtgcacacct
tccctgctgt gctgcagtcc 540tccggcctgt actccctgtc ctccgtcgtg
accgtgcctt ccagctctct gggcacccag 600acctacatct gcaacgtgaa
ccacaagccc tccaacacca aggtggacaa gaaggtggaa 660cccaagtcct
gcgacaagac ccacacctgt cccccttgtc ctgcccctga actgctgggc
720ggaccttccg tgttcctgtt ccccccaaag cccaaggaca ccctgatgat
ctcccggacc 780cccgaagtga cctgcgtggt ggtggatgtg tcccacgagg
accctgaagt gaagttcaat 840tggtacgtgg acggcgtgga agtgcacaac
gccaagacca agcctagaga ggaacagtac 900aactccacct accgggtggt
gtccgtgctg accgtgctgc accaggattg gctgaacggc 960aaagagtaca
agtgcaaggt gtccaacaag gccctgcctg cccccatcga aaagaccatc
1020tccaaggcca agggccagcc ccgggaaccc caggtgtaca cactgccccc
tagcagggac 1080gagctgacca agaaccaggt gtccctgacc tgtctcgtga
aaggcttcta cccctccgat 1140atcgccgtgg aatgggagtc caacggccag
cctgagaaca actacaagac caccccccct 1200gtgctggact ccgacggctc
attcttcctg tacagcaagc tgacagtgga caagtcccgg 1260tggcagcagg
gcaacgtgtt ctcctgctcc gtgatgcacg aggccctgca caaccactac
1320acccagaagt ccctgtccct gagccccggc aagtgatga 135937011PRTHomo
Sapien 370Gln Ser Leu Leu His Ser Asn Glu Tyr Asn Tyr1 5
103713PRTHomo Sapien 371Leu Gly Ser13729PRTHomo Sapien 372Met Gln
Ser Leu Gln Thr Pro Leu Thr1 5373112PRTHomo Sapien 373Asp Ile Val
Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro
Ala Ser Ile Ser Cys Arg Ser Ser Gln Ser Leu Leu His Ser 20 25 30Asn
Glu Tyr Asn Tyr Leu Asp Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40
45Pro Gln Leu Leu Ile Phe Leu Gly Ser Asn Arg Ala Ser Gly Val Pro
50 55 60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys
Ile65 70 75 80Thr Arg Val Glu Ala Glu Asp Val Gly Ile Tyr Tyr Cys
Met Gln Ser 85 90 95Leu Gln Thr Pro Leu Thr Phe Gly Gly Gly Thr Lys
Val Glu Ile Lys 100 105 110374336DNAHomo Sapien 374gatattgtga
tgactcagtc tccactctcc ctgcccgtca cccctggaga gccggcctcc 60atctcctgca
ggtctagtca gagcctcctg catagtaatg aatacaacta tttggattgg
120tacctgcaga agccagggca gtctccacag ctcctgatct ttttgggttc
taatcgggcc 180tccggggtcc ctgacaggtt cagtggcagt ggatcaggca
cagattttac actgaaaatc 240accagagtgg aggctgagga tgttggaatt
tattactgca tgcaatctct acaaactccg 300ctcactttcg gcggagggac
caaggtggag atcaaa 336375219PRTHomo Sapien 375Asp Ile Val Met Thr
Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser
Ile Ser Cys Arg Ser Ser Gln Ser Leu Leu His Ser 20 25 30Asn Glu Tyr
Asn Tyr Leu Asp Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Gln
Leu Leu Ile Phe Leu Gly Ser Asn Arg Ala Ser Gly Val Pro 50 55 60Asp
Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75
80Thr Arg Val Glu Ala Glu Asp Val Gly Ile Tyr Tyr Cys Met Gln Ser
85 90 95Leu Gln Thr Pro Leu Thr Phe Gly Gly Gly Thr Lys Val Glu Ile
Lys 100 105 110Arg Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro
Ser Asp Glu 115 120 125Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys
Leu Leu Asn Asn Phe 130 135 140Tyr Pro Arg Glu Ala Lys Val Gln Trp
Lys Val Asp Asn Ala Leu Gln145 150 155 160Ser Gly Asn Ser Gln Glu
Ser Val Thr Glu Gln Asp Ser Lys Asp Ser
165 170 175Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp
Tyr Glu 180 185 190Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln
Gly Leu Ser Ser 195 200 205Pro Val Thr Lys Ser Phe Asn Arg Gly Glu
Cys 210 215376657DNAHomo Sapien 376gatattgtga tgactcagtc tccactctcc
ctgcccgtca cccctggaga gccggcctcc 60atctcctgca ggtctagtca gagcctcctg
catagtaatg aatacaacta tttggattgg 120tacctgcaga agccagggca
gtctccacag ctcctgatct ttttgggttc taatcgggcc 180tccggggtcc
ctgacaggtt cagtggcagt ggatcaggca cagattttac actgaaaatc
240accagagtgg aggctgagga tgttggaatt tattactgca tgcaatctct
acaaactccg 300ctcactttcg gcggagggac caaggtggag atcaaacgta
cggtggccgc tccctccgtg 360ttcatcttcc caccttccga cgagcagctg
aagtccggca ccgcttctgt cgtgtgcctg 420ctgaacaact tctacccccg
cgaggccaag gtgcagtgga aggtggacaa cgccctgcag 480tccggcaact
cccaggaatc cgtgaccgag caggactcca aggacagcac ctactccctg
540tcctccaccc tgaccctgtc caaggccgac tacgagaagc acaaggtgta
cgcctgcgaa 600gtgacccacc agggcctgtc tagccccgtg accaagtctt
tcaaccgggg cgagtgt 6573778PRTHomo Sapien 377Gly Tyr Thr Phe Thr Ser
Tyr Gly1 53788PRTHomo Sapien 378Ile Ser Ala Tyr Asn Gly Asn Thr1
537917PRTHomo Sapien 379Ala Arg Ser Thr Tyr Phe Tyr Gly Ser Gly Thr
Leu Tyr Gly Met Asp1 5 10 15Val380124PRTHomo Sapien 380Gln Val Gln
Leu Val Gln Ser Gly Gly Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30Gly
Phe Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys Leu
50 55 60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr Ser Thr Ala
Tyr65 70 75 80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr Ala Val
Tyr Tyr Cys 85 90 95Ala Arg Ser Thr Tyr Phe Tyr Gly Ser Gly Thr Leu
Tyr Gly Met Asp 100 105 110Val Trp Gly Gln Gly Thr Thr Val Thr Val
Ser Ser 115 120381372DNAHomo Sapien 381caggttcaac tggtgcagtc
tggaggtgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta
cacctttacc agctatggtt tcagctgggt gcgacaggcc 120cctggacaag
gactagagtg gatgggatgg atcagcgctt acaatggtaa cacaaactat
180gcacagaagc tccagggcag agtcaccatg accacagaca catccacgag
cacagcctac 240atggagctga ggagcttgag atctgacgac acggccgtgt
attactgtgc gagatctacg 300tatttctatg gttcggggac cctctacggt
atggacgtct ggggccaagg gaccacggtc 360accgtctcct ca 372382454PRTHomo
Sapien 382Gln Val Gln Leu Val Gln Ser Gly Gly Glu Val Lys Lys Pro
Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe
Thr Ser Tyr 20 25 30Gly Phe Ser Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn
Tyr Ala Gln Lys Leu 50 55 60Gln Gly Arg Val Thr Met Thr Thr Asp Thr
Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu Arg Ser Leu Arg Ser
Asp Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Thr Tyr Phe Tyr
Gly Ser Gly Thr Leu Tyr Gly Met Asp 100 105 110Val Trp Gly Gln Gly
Thr Thr Val Thr Val Ser Ser Ala Ser Thr Lys 115 120 125Gly Pro Ser
Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly 130 135 140Gly
Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro145 150
155 160Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His
Thr 165 170 175Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu
Ser Ser Val 180 185 190Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln
Thr Tyr Ile Cys Asn 195 200 205Val Asn His Lys Pro Ser Asn Thr Lys
Val Asp Lys Lys Val Glu Pro 210 215 220Lys Ser Cys Asp Lys Thr His
Thr Cys Pro Pro Cys Pro Ala Pro Glu225 230 235 240Leu Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 245 250 255Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 260 265
270Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
275 280 285Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn 290 295 300Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp305 310 315 320Leu Asn Gly Lys Glu Tyr Lys Cys Lys
Val Ser Asn Lys Ala Leu Pro 325 330 335Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu 340 345 350Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn 355 360 365Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 370 375 380Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr385 390
395 400Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys 405 410 415Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys 420 425 430Ser Val Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu 435 440 445Ser Leu Ser Pro Gly Lys
4503831368DNAHomo Sapien 383caggttcaac tggtgcagtc tggaggtgag
gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttacc
agctatggtt tcagctgggt gcgacaggcc 120cctggacaag gactagagtg
gatgggatgg atcagcgctt acaatggtaa cacaaactat 180gcacagaagc
tccagggcag agtcaccatg accacagaca catccacgag cacagcctac
240atggagctga ggagcttgag atctgacgac acggccgtgt attactgtgc
gagatctacg 300tatttctatg gttcggggac cctctacggt atggacgtct
ggggccaagg gaccacggtc 360accgtctcct cagccagcac caagggcccc
tctgtgttcc ctctggcccc ttccagcaag 420tccacctctg gcggaacagc
cgctctgggc tgcctcgtga aggactactt ccccgagcct 480gtgaccgtgt
cctggaactc tggcgctctg accagcggag tgcacacctt ccctgctgtg
540ctgcagtcct ccggcctgta ctccctgtcc tccgtcgtga ccgtgccttc
cagctctctg 600ggcacccaga cctacatctg caacgtgaac cacaagccct
ccaacaccaa ggtggacaag 660aaggtggaac ccaagtcctg cgacaagacc
cacacctgtc ccccttgtcc tgcccctgaa 720ctgctgggcg gaccttccgt
gttcctgttc cccccaaagc ccaaggacac cctgatgatc 780tcccggaccc
ccgaagtgac ctgcgtggtg gtggatgtgt cccacgagga ccctgaagtg
840aagttcaatt ggtacgtgga cggcgtggaa gtgcacaacg ccaagaccaa
gcctagagag 900gaacagtaca actccaccta ccgggtggtg tccgtgctga
ccgtgctgca ccaggattgg 960ctgaacggca aagagtacaa gtgcaaggtg
tccaacaagg ccctgcctgc ccccatcgaa 1020aagaccatct ccaaggccaa
gggccagccc cgggaacccc aggtgtacac actgccccct 1080agcagggacg
agctgaccaa gaaccaggtg tccctgacct gtctcgtgaa aggcttctac
1140ccctccgata tcgccgtgga atgggagtcc aacggccagc ctgagaacaa
ctacaagacc 1200accccccctg tgctggactc cgacggctca ttcttcctgt
acagcaagct gacagtggac 1260aagtcccggt ggcagcaggg caacgtgttc
tcctgctccg tgatgcacga ggccctgcac 1320aaccactaca cccagaagtc
cctgtccctg agccccggca agtgatga 136838411PRTHomo Sapien 384Gln Ser
Leu Leu His Ser Asp Gly Tyr Asn Tyr1 5 103853PRTHomo Sapien 385Leu
Gly Ser13869PRTHomo Sapien 386Met Gln Ala Leu Gln Thr Pro Leu Ser1
5387112PRTHomo Sapien 387Asp Ile Val Met Thr Gln Ser Pro Leu Ser
Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile Ser Cys Arg Ser
Ser Gln Ser Leu Leu His Ser 20 25 30Asp Gly Tyr Asn Tyr Leu Asp Trp
Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Gln Leu Leu Ile Tyr Leu
Gly Ser Thr Arg Ala Ser Gly Phe Pro 50 55 60Asp Arg Phe Ser Gly Ser
Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu
Ala Glu Asp Val Gly Val Tyr Tyr Cys Met Gln Ala 85 90 95Leu Gln Thr
Pro Leu Ser Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100 105
110388336DNAHomo Sapien 388gatattgtga tgactcagtc tccactctcc
ctgcccgtca cccctggaga gccggcctcc 60atctcctgca ggtctagtca gagcctcctg
catagtgatg gatacaactg tttggattgg 120tacctgcaga agccagggca
gtctccacag ctcctgatct atttgggttc tactcgggcc 180tccgggttcc
ctgacaggtt cagtggcagt ggatcaggca cagattttac actgaaaatc
240agcagagtgg aggctgagga tgttggggtt tattactgca tgcaagctct
acaaactccg 300tgcagttttg gccaggggac caagctggag atcaaa
336389219PRTHomo Sapien 389Asp Ile Val Met Thr Gln Ser Pro Leu Ser
Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile Ser Cys Arg Ser
Ser Gln Ser Leu Leu His Ser 20 25 30Asp Gly Tyr Asn Tyr Leu Asp Trp
Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Gln Leu Leu Ile Tyr Leu
Gly Ser Thr Arg Ala Ser Gly Phe Pro 50 55 60Asp Arg Phe Ser Gly Ser
Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu
Ala Glu Asp Val Gly Val Tyr Tyr Cys Met Gln Ala 85 90 95Leu Gln Thr
Pro Leu Ser Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100 105 110Arg
Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu 115 120
125Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
130 135 140Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln145 150 155 160Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser 165 170 175Thr Tyr Ser Leu Ser Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu 180 185 190Lys His Lys Val Tyr Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser 195 200 205Pro Val Thr Lys Ser
Phe Asn Arg Gly Glu Cys 210 215390657DNAHomo Sapien 390gatattgtga
tgactcagtc tccactctcc ctgcccgtca cccctggaga gccggcctcc 60atctcctgca
ggtctagtca gagcctcctg catagtgatg gatacaactg tttggattgg
120tacctgcaga agccagggca gtctccacag ctcctgatct atttgggttc
tactcgggcc 180tccgggttcc ctgacaggtt cagtggcagt ggatcaggca
cagattttac actgaaaatc 240agcagagtgg aggctgagga tgttggggtt
tattactgca tgcaagctct acaaactccg 300tgcagttttg gccaggggac
caagctggag atcaaacgta cggtggccgc tccctccgtg 360ttcatcttcc
caccttccga cgagcagctg aagtccggca ccgcttctgt cgtgtgcctg
420ctgaacaact tctacccccg cgaggccaag gtgcagtgga aggtggacaa
cgccctgcag 480tccggcaact cccaggaatc cgtgaccgag caggactcca
aggacagcac ctactccctg 540tcctccaccc tgaccctgtc caaggccgac
tacgagaagc acaaggtgta cgcctgcgaa 600gtgacccacc agggcctgtc
tagccccgtg accaagtctt tcaaccgggg cgagtgt 6573918PRTHomo Sapien
391Gly Tyr Thr Phe Thr Ser Tyr Gly1 53928PRTHomo Sapien 392Ile Ser
Ala Tyr Asn Gly Asn Thr1 539317PRTHomo Sapien 393Ala Arg Ser Thr
Tyr Phe Tyr Gly Ser Gly Thr Leu Tyr Gly Met Asp1 5 10
15Val394124PRTHomo Sapien 394Gln Val Gln Leu Val Gln Ser Gly Gly
Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala
Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30Gly Phe Ser Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Trp Ile Ser Ala Tyr
Asn Gly Asn Thr Asn Tyr Ala Gln Lys Leu 50 55 60Gln Gly Arg Val Thr
Met Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu
Arg Ser Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg
Ser Thr Tyr Phe Tyr Gly Ser Gly Thr Leu Tyr Gly Met Asp 100 105
110Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115
120395372DNAHomo Sapien 395caggttcaac tggtgcagtc tggaggtgag
gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttacc
agctatggtt tcagctgggt gcgacaggcc 120cctggacaag gactagagtg
gatgggatgg atcagcgctt acaatggtaa cacaaactat 180gcacagaagc
tccagggcag agtcaccatg accacagaca catccacgag cacagcctac
240atggagctga ggagcttgag atctgacgac acggccgtgt attactgtgc
gagatctacg 300tatttctatg gttcggggac cctctacggt atggacgtct
ggggccaagg gaccacggtc 360accgtctcct ca 372396454PRTHomo Sapien
396Gln Val Gln Leu Val Gln Ser Gly Gly Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser
Tyr 20 25 30Gly Phe Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala
Gln Lys Leu 50 55 60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Thr Tyr Phe Tyr Gly Ser
Gly Thr Leu Tyr Gly Met Asp 100 105 110Val Trp Gly Gln Gly Thr Thr
Val Thr Val Ser Ser Ala Ser Thr Lys 115 120 125Gly Pro Ser Val Phe
Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly 130 135 140Gly Thr Ala
Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro145 150 155
160Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr
165 170 175Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser
Ser Val 180 185 190Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr
Tyr Ile Cys Asn 195 200 205Val Asn His Lys Pro Ser Asn Thr Lys Val
Asp Lys Lys Val Glu Pro 210 215 220Lys Ser Cys Asp Lys Thr His Thr
Cys Pro Pro Cys Pro Ala Pro Glu225 230 235 240Leu Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 245 250 255Thr Leu Met
Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 260 265 270Val
Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly 275 280
285Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn
290 295 300Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln
Asp Trp305 310 315 320Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro 325 330 335Ala Pro Ile Glu Lys Thr Ile Ser Lys
Ala Lys Gly Gln Pro Arg Glu 340 345 350Pro Gln Val Tyr Thr Leu Pro
Pro Ser Arg Asp Glu Leu Thr Lys Asn 355 360 365Gln Val Ser Leu Thr
Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 370 375 380Ala Val Glu
Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr385 390 395
400Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
405 410 415Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe
Ser Cys 420 425 430Ser Val Met His Glu Ala Leu His Asn His Tyr Thr
Gln Lys Ser Leu 435 440 445Ser Leu Ser Pro Gly Lys
4503971368DNAHomo Sapien 397caggttcaac tggtgcagtc tggaggtgag
gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttacc
agctatggtt tcagctgggt gcgacaggcc 120cctggacaag gactagagtg
gatgggatgg atcagcgctt acaatggtaa cacaaactat 180gcacagaagc
tccagggcag agtcaccatg accacagaca catccacgag cacagcctac
240atggagctga ggagcttgag atctgacgac acggccgtgt attactgtgc
gagatctacg 300tatttctatg gttcggggac cctctacggt atggacgtct
ggggccaagg gaccacggtc 360accgtctcct cagccagcac caagggcccc
tctgtgttcc ctctggcccc ttccagcaag 420tccacctctg gcggaacagc
cgctctgggc tgcctcgtga aggactactt ccccgagcct 480gtgaccgtgt
cctggaactc tggcgctctg accagcggag tgcacacctt ccctgctgtg
540ctgcagtcct ccggcctgta ctccctgtcc tccgtcgtga ccgtgccttc
cagctctctg 600ggcacccaga cctacatctg caacgtgaac cacaagccct
ccaacaccaa ggtggacaag 660aaggtggaac ccaagtcctg cgacaagacc
cacacctgtc ccccttgtcc tgcccctgaa 720ctgctgggcg gaccttccgt
gttcctgttc cccccaaagc ccaaggacac cctgatgatc 780tcccggaccc
ccgaagtgac ctgcgtggtg gtggatgtgt cccacgagga ccctgaagtg
840aagttcaatt ggtacgtgga cggcgtggaa gtgcacaacg ccaagaccaa
gcctagagag 900gaacagtaca actccaccta ccgggtggtg tccgtgctga
ccgtgctgca ccaggattgg 960ctgaacggca
aagagtacaa gtgcaaggtg tccaacaagg ccctgcctgc ccccatcgaa
1020aagaccatct ccaaggccaa gggccagccc cgggaacccc aggtgtacac
actgccccct 1080agcagggacg agctgaccaa gaaccaggtg tccctgacct
gtctcgtgaa aggcttctac 1140ccctccgata tcgccgtgga atgggagtcc
aacggccagc ctgagaacaa ctacaagacc 1200accccccctg tgctggactc
cgacggctca ttcttcctgt acagcaagct gacagtggac 1260aagtcccggt
ggcagcaggg caacgtgttc tcctgctccg tgatgcacga ggccctgcac
1320aaccactaca cccagaagtc cctgtccctg agccccggca agtgatga
136839811PRTHomo Sapien 398Gln Ser Leu Leu His Ser Asp Gly Tyr Asn
Cys1 5 103993PRTHomo Sapien 399Leu Gly Ser14009PRTHomo Sapien
400Met Gln Ala Leu Gln Thr Pro Cys Ser1 5401112PRTHomo Sapien
401Asp Ile Val Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1
5 10 15Glu Pro Ala Ser Ile Ser Cys Arg Ser Ser Gln Ser Leu Leu His
Ser 20 25 30Asp Gly Tyr Asn Cys Leu Asp Trp Tyr Leu Gln Lys Pro Gly
Gln Ser 35 40 45Pro Gln Leu Leu Ile Tyr Leu Gly Ser Thr Arg Ala Ser
Gly Phe Pro 50 55 60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu Ala Glu Asp Val Gly Val
Tyr Tyr Cys Met Gln Ala 85 90 95Leu Gln Thr Pro Cys Ser Phe Gly Gln
Gly Thr Lys Leu Glu Ile Lys 100 105 110402336DNAHomo Sapien
402gatattgtga tgactcagtc tccactctcc ctgcccgtca cccctggaga
gccggcctcc 60atctcctgca ggtctagtca gagcctcctg catagtgatg gatacaactg
tttggattgg 120tacctgcaga agccagggca gtctccacag ctcctgatct
atttgggttc tactcgggcc 180tccgggttcc ctgacaggtt cagtggcagt
ggatcaggca cagattttac actgaaaatc 240agcagagtgg aggctgagga
tgttggggtt tattactgca tgcaagctct acaaactccg 300tgcagttttg
gccaggggac caagctggag atcaaa 336403219PRTHomo Sapien 403Asp Ile Val
Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro
Ala Ser Ile Ser Cys Arg Ser Ser Gln Ser Leu Leu His Ser 20 25 30Asp
Gly Tyr Asn Cys Leu Asp Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40
45Pro Gln Leu Leu Ile Tyr Leu Gly Ser Thr Arg Ala Ser Gly Phe Pro
50 55 60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys
Ile65 70 75 80Ser Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys
Met Gln Ala 85 90 95Leu Gln Thr Pro Cys Ser Phe Gly Gln Gly Thr Lys
Leu Glu Ile Lys 100 105 110Arg Thr Val Ala Ala Pro Ser Val Phe Ile
Phe Pro Pro Ser Asp Glu 115 120 125Gln Leu Lys Ser Gly Thr Ala Ser
Val Val Cys Leu Leu Asn Asn Phe 130 135 140Tyr Pro Arg Glu Ala Lys
Val Gln Trp Lys Val Asp Asn Ala Leu Gln145 150 155 160Ser Gly Asn
Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 165 170 175Thr
Tyr Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu 180 185
190Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser
195 200 205Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 210
215404657DNAHomo Sapien 404gatattgtga tgactcagtc tccactctcc
ctgcccgtca cccctggaga gccggcctcc 60atctcctgca ggtctagtca gagcctcctg
catagtgatg gatacaactg tttggattgg 120tacctgcaga agccagggca
gtctccacag ctcctgatct atttgggttc tactcgggcc 180tccgggttcc
ctgacaggtt cagtggcagt ggatcaggca cagattttac actgaaaatc
240agcagagtgg aggctgagga tgttggggtt tattactgca tgcaagctct
acaaactccg 300tgcagttttg gccaggggac caagctggag atcaaacgta
cggtggccgc tccctccgtg 360ttcatcttcc caccttccga cgagcagctg
aagtccggca ccgcttctgt cgtgtgcctg 420ctgaacaact tctacccccg
cgaggccaag gtgcagtgga aggtggacaa cgccctgcag 480tccggcaact
cccaggaatc cgtgaccgag caggactcca aggacagcac ctactccctg
540tcctccaccc tgaccctgtc caaggccgac tacgagaagc acaaggtgta
cgcctgcgaa 600gtgacccacc agggcctgtc tagccccgtg accaagtctt
tcaaccgggg cgagtgt 6574058PRTHomo Sapien 405Gly Val Thr Phe Asp Asp
Tyr Gly1 54068PRTHomo Sapien 406Ile Asn Trp Asn Gly Gly Asp Thr1
540717PRTHomo Sapien 407Ala Arg Asp Phe Tyr Gly Ser Gly Ser Tyr Tyr
His Val Pro Phe Asp1 5 10 15Tyr408124PRTHomo Sapien 408Glu Val Gln
Leu Val Glu Ser Gly Gly Gly Val Val Arg Pro Gly Gly1 5 10 15Ser Leu
Arg Leu Ser Cys Val Ala Ser Gly Val Thr Phe Asp Asp Tyr 20 25 30Gly
Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Gly Ile Asn Trp Asn Gly Gly Asp Thr Asp Tyr Ser Asp Ser Val
50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu
Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Leu
Tyr Tyr Cys 85 90 95Ala Arg Asp Phe Tyr Gly Ser Gly Ser Tyr Tyr His
Val Pro Phe Asp 100 105 110Tyr Trp Gly Gln Gly Ile Leu Val Thr Val
Ser Ser 115 120409372DNAHomo Sapien 409gaggtgcagc tggtggagtc
tgggggaggt gtggtacggc ctggggggtc cctgagactc 60tcctgtgtag cctctggagt
cacctttgat gattatggca tgagctgggt ccgccaagct 120ccagggaagg
ggctggartg ggtctctggt attaattgga atggtggcga cacagattat
180tcagactctg tgaagggccg attcaccatc tccagagaca acgccaagaa
ctccctgtat 240ctacaaatga atagtctgag agccgaggac acggccttgt
attactgtgc gagggatttc 300tatggttcgg ggagttatta tcacgttcct
tttgactact ggggccaggg aatcctggtc 360accgtctcct ca 372410454PRTHomo
Sapien 410Glu Val Gln Leu Val Glu Ser Gly Gly Gly Val Val Arg Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Val Ala Ser Gly Val Thr Phe
Asp Asp Tyr 20 25 30Gly Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ser Gly Ile Asn Trp Asn Gly Gly Asp Thr Asp
Tyr Ser Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Arg Asp Phe Tyr Gly Ser
Gly Ser Tyr Tyr His Val Pro Phe Asp 100 105 110Tyr Trp Gly Gln Gly
Ile Leu Val Thr Val Ser Ser Ala Ser Thr Lys 115 120 125Gly Pro Ser
Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly 130 135 140Gly
Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro145 150
155 160Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His
Thr 165 170 175Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu
Ser Ser Val 180 185 190Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln
Thr Tyr Ile Cys Asn 195 200 205Val Asn His Lys Pro Ser Asn Thr Lys
Val Asp Lys Lys Val Glu Pro 210 215 220Lys Ser Cys Asp Lys Thr His
Thr Cys Pro Pro Cys Pro Ala Pro Glu225 230 235 240Leu Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 245 250 255Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 260 265
270Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
275 280 285Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn 290 295 300Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp305 310 315 320Leu Asn Gly Lys Glu Tyr Lys Cys Lys
Val Ser Asn Lys Ala Leu Pro 325 330 335Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu 340 345 350Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn 355 360 365Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 370 375 380Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr385 390
395 400Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys 405 410 415Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys 420 425 430Ser Val Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu 435 440 445Ser Leu Ser Pro Gly Lys
4504111368DNAHomo Sapien 411gaggtgcagc tggtggagtc tgggggaggt
gtggtacggc ctggggggtc cctgagactc 60tcctgtgtag cctctggagt cacctttgat
gattatggca tgagctgggt ccgccaagct 120ccagggaagg ggctggartg
ggtctctggt attaattgga atggtggcga cacagattat 180tcagactctg
tgaagggccg attcaccatc tccagagaca acgccaagaa ctccctgtat
240ctacaaatga atagtctgag agccgaggac acggccttgt attactgtgc
gagggatttc 300tatggttcgg ggagttatta tcacgttcct tttgactact
ggggccaggg aatcctggtc 360accgtctcct cagccagcac caagggcccc
tctgtgttcc ctctggcccc ttccagcaag 420tccacctctg gcggaacagc
cgctctgggc tgcctcgtga aggactactt ccccgagcct 480gtgaccgtgt
cctggaactc tggcgctctg accagcggag tgcacacctt ccctgctgtg
540ctgcagtcct ccggcctgta ctccctgtcc tccgtcgtga ccgtgccttc
cagctctctg 600ggcacccaga cctacatctg caacgtgaac cacaagccct
ccaacaccaa ggtggacaag 660aaggtggaac ccaagtcctg cgacaagacc
cacacctgtc ccccttgtcc tgcccctgaa 720ctgctgggcg gaccttccgt
gttcctgttc cccccaaagc ccaaggacac cctgatgatc 780tcccggaccc
ccgaagtgac ctgcgtggtg gtggatgtgt cccacgagga ccctgaagtg
840aagttcaatt ggtacgtgga cggcgtggaa gtgcacaacg ccaagaccaa
gcctagagag 900gaacagtaca actccaccta ccgggtggtg tccgtgctga
ccgtgctgca ccaggattgg 960ctgaacggca aagagtacaa gtgcaaggtg
tccaacaagg ccctgcctgc ccccatcgaa 1020aagaccatct ccaaggccaa
gggccagccc cgggaacccc aggtgtacac actgccccct 1080agcagggacg
agctgaccaa gaaccaggtg tccctgacct gtctcgtgaa aggcttctac
1140ccctccgata tcgccgtgga atgggagtcc aacggccagc ctgagaacaa
ctacaagacc 1200accccccctg tgctggactc cgacggctca ttcttcctgt
acagcaagct gacagtggac 1260aagtcccggt ggcagcaggg caacgtgttc
tcctgctccg tgatgcacga ggccctgcac 1320aaccactaca cccagaagtc
cctgtccctg agccccggca agtgatga 13684127PRTHomo Sapien 412Gln Ser
Val Ser Arg Ser Tyr1 54133PRTHomo Sapien 413Gly Ala Ser14149PRTHomo
Sapien 414His Gln Tyr Asp Met Ser Pro Phe Thr1 5415108PRTHomo
Sapien 415Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser
Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val
Ser Arg Ser 20 25 30Tyr Leu Ala Trp Tyr Gln Gln Lys Arg Gly Gln Ala
Pro Arg Leu Leu 35 40 45Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly Ile
Pro Asp Arg Phe Ser 50 55 60Gly Asp Gly Ser Gly Thr Asp Phe Thr Leu
Ser Ile Ser Arg Leu Glu65 70 75 80Pro Glu Asp Phe Ala Val Tyr Tyr
Cys His Gln Tyr Asp Met Ser Pro 85 90 95Phe Thr Phe Gly Pro Gly Thr
Lys Val Asp Ile Lys 100 105416324DNAHomo Sapien 416gaaattgtgt
tgacgcagtc tccagggacc ctgtctttgt ctccagggga aagagccacc 60ctctcctgca
gggccagtca gagtgttagc agaagctact tagcctggta ccagcagaaa
120cgtggccagg ctcccaggct cctcatctat ggtgcatcca gcagggccac
tggcatccca 180gacaggttca gtggcgatgg gtctgggaca gacttcactc
tctccatcag cagactggag 240cctgaagatt ttgcagtgta ttactgtcac
cagtatgata tgtcaccatt cactttcggc 300cctgggacca aagtggatat caaa
324417215PRTHomo Sapien 417Glu Ile Val Leu Thr Gln Ser Pro Gly Thr
Leu Ser Leu Ser Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala
Ser Gln Ser Val Ser Arg Ser 20 25 30Tyr Leu Ala Trp Tyr Gln Gln Lys
Arg Gly Gln Ala Pro Arg Leu Leu 35 40 45Ile Tyr Gly Ala Ser Ser Arg
Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60Gly Asp Gly Ser Gly Thr
Asp Phe Thr Leu Ser Ile Ser Arg Leu Glu65 70 75 80Pro Glu Asp Phe
Ala Val Tyr Tyr Cys His Gln Tyr Asp Met Ser Pro 85 90 95Phe Thr Phe
Gly Pro Gly Thr Lys Val Asp Ile Lys Arg Thr Val Ala 100 105 110Ala
Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser 115 120
125Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu
130 135 140Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly
Asn Ser145 150 155 160Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp
Ser Thr Tyr Ser Leu 165 170 175Ser Ser Thr Leu Thr Leu Ser Lys Ala
Asp Tyr Glu Lys His Lys Val 180 185 190Tyr Ala Cys Glu Val Thr His
Gln Gly Leu Ser Ser Pro Val Thr Lys 195 200 205Ser Phe Asn Arg Gly
Glu Cys 210 215418645DNAHomo Sapien 418gaaattgtgt tgacgcagtc
tccagggacc ctgtctttgt ctccagggga aagagccacc 60ctctcctgca gggccagtca
gagtgttagc agaagctact tagcctggta ccagcagaaa 120cgtggccagg
ctcccaggct cctcatctat ggtgcatcca gcagggccac tggcatccca
180gacaggttca gtggcgatgg gtctgggaca gacttcactc tctccatcag
cagactggag 240cctgaagatt ttgcagtgta ttactgtcac cagtatgata
tgtcaccatt cactttcggc 300cctgggacca aagtggatat caaacgtacg
gtggccgctc cctccgtgtt catcttccca 360ccttccgacg agcagctgaa
gtccggcacc gcttctgtcg tgtgcctgct gaacaacttc 420tacccccgcg
aggccaaggt gcagtggaag gtggacaacg ccctgcagtc cggcaactcc
480caggaatccg tgaccgagca ggactccaag gacagcacct actccctgtc
ctccaccctg 540accctgtcca aggccgacta cgagaagcac aaggtgtacg
cctgcgaagt gacccaccag 600ggcctgtcta gccccgtgac caagtctttc
aaccggggcg agtgt 6454198PRTHomo Sapien 419Gly Leu Thr Phe Asp Asp
Tyr Gly1 54208PRTHomo Sapien 420Ile Asn Trp Asn Gly Asp Asn Thr1
542117PRTHomo Sapien 421Ala Arg Asp Tyr Tyr Gly Ser Gly Ser Tyr Tyr
Asn Val Pro Phe Asp1 5 10 15Tyr422124PRTHomo Sapien 422Glu Val Gln
Leu Val Glu Ser Gly Gly Gly Val Val Arg Pro Gly Gly1 5 10 15Ser Leu
Arg Leu Ser Cys Ala Ala Ser Gly Leu Thr Phe Asp Asp Tyr 20 25 30Gly
Met Ser Trp Val Arg Gln Val Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Gly Ile Asn Trp Asn Gly Asp Asn Thr Asp Tyr Ala Asp Ser Val
50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu
Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Leu
Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ser Gly Ser Tyr Tyr Asn
Val Pro Phe Asp 100 105 110Tyr Trp Gly Gln Gly Thr Leu Val Thr Val
Ser Ser 115 120423372DNAHomo Sapien 423gaggtgcagc tggtggagtc
tgggggaggt gtggtacggc ctggggggtc cctgagactc 60tcctgtgcag cctctggact
cacctttgat gattatggca tgagctgggt ccgccaagtt 120ccagggaagg
ggctggagtg ggtctctggt attaattgga atggtgataa cacagattat
180gcagactctg tgaagggccg attcaccatc tccagagaca acgccaagaa
ctccctgtat 240ctgcaaatga acagtctgag agccgaggac acggccttgt
attactgtgc gagggattac 300tatggttcgg ggagttatta taacgttcct
tttgactact ggggccaggg aaccctggtc 360accgtctcct ca 372424454PRTHomo
Sapien 424Glu Val Gln Leu Val Glu Ser Gly Gly Gly Val Val Arg Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Leu Thr Phe
Asp Asp Tyr 20 25 30Gly Met Ser Trp Val Arg Gln Val Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ser Gly Ile Asn Trp Asn Gly Asp Asn Thr Asp
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90 95Ala Arg Asp Tyr Tyr Gly Ser
Gly Ser Tyr Tyr Asn Val Pro Phe Asp 100 105 110Tyr Trp Gly Gln Gly
Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys 115 120 125Gly Pro Ser
Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly 130 135 140Gly
Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro145 150
155 160Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His
Thr 165 170 175Phe Pro Ala Val Leu Gln Ser
Ser Gly Leu Tyr Ser Leu Ser Ser Val 180 185 190Val Thr Val Pro Ser
Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn 195 200 205Val Asn His
Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro 210 215 220Lys
Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu225 230
235 240Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys
Asp 245 250 255Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val
Val Val Asp 260 265 270Val Ser His Glu Asp Pro Glu Val Lys Phe Asn
Trp Tyr Val Asp Gly 275 280 285Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Tyr Asn 290 295 300Ser Thr Tyr Arg Val Val Ser
Val Leu Thr Val Leu His Gln Asp Trp305 310 315 320Leu Asn Gly Lys
Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro 325 330 335Ala Pro
Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu 340 345
350Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn
355 360 365Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp Ile 370 375 380Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr Lys Thr385 390 395 400Thr Pro Pro Val Leu Asp Ser Asp Gly
Ser Phe Phe Leu Tyr Ser Lys 405 410 415Leu Thr Val Asp Lys Ser Arg
Trp Gln Gln Gly Asn Val Phe Ser Cys 420 425 430Ser Val Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu 435 440 445Ser Leu Ser
Pro Gly Lys 4504251368DNAHomo Sapien 425gaggtgcagc tggtggagtc
tgggggaggt gtggtacggc ctggggggtc cctgagactc 60tcctgtgcag cctctggact
cacctttgat gattatggca tgagctgggt ccgccaagtt 120ccagggaagg
ggctggagtg ggtctctggt attaattgga atggtgataa cacagattat
180gcagactctg tgaagggccg attcaccatc tccagagaca acgccaagaa
ctccctgtat 240ctgcaaatga acagtctgag agccgaggac acggccttgt
attactgtgc gagggattac 300tatggttcgg ggagttatta taacgttcct
tttgactact ggggccaggg aaccctggtc 360accgtctcct cagccagcac
caagggcccc tctgtgttcc ctctggcccc ttccagcaag 420tccacctctg
gcggaacagc cgctctgggc tgcctcgtga aggactactt ccccgagcct
480gtgaccgtgt cctggaactc tggcgctctg accagcggag tgcacacctt
ccctgctgtg 540ctgcagtcct ccggcctgta ctccctgtcc tccgtcgtga
ccgtgccttc cagctctctg 600ggcacccaga cctacatctg caacgtgaac
cacaagccct ccaacaccaa ggtggacaag 660aaggtggaac ccaagtcctg
cgacaagacc cacacctgtc ccccttgtcc tgcccctgaa 720ctgctgggcg
gaccttccgt gttcctgttc cccccaaagc ccaaggacac cctgatgatc
780tcccggaccc ccgaagtgac ctgcgtggtg gtggatgtgt cccacgagga
ccctgaagtg 840aagttcaatt ggtacgtgga cggcgtggaa gtgcacaacg
ccaagaccaa gcctagagag 900gaacagtaca actccaccta ccgggtggtg
tccgtgctga ccgtgctgca ccaggattgg 960ctgaacggca aagagtacaa
gtgcaaggtg tccaacaagg ccctgcctgc ccccatcgaa 1020aagaccatct
ccaaggccaa gggccagccc cgggaacccc aggtgtacac actgccccct
1080agcagggacg agctgaccaa gaaccaggtg tccctgacct gtctcgtgaa
aggcttctac 1140ccctccgata tcgccgtgga atgggagtcc aacggccagc
ctgagaacaa ctacaagacc 1200accccccctg tgctggactc cgacggctca
ttcttcctgt acagcaagct gacagtggac 1260aagtcccggt ggcagcaggg
caacgtgttc tcctgctccg tgatgcacga ggccctgcac 1320aaccactaca
cccagaagtc cctgtccctg agccccggca agtgatga 13684267PRTHomo Sapien
426Gln Ser Val Ser Ser Ser Tyr1 54273PRTHomo Sapien 427Gly Ala
Ser14288PRTHomo Sapien 428Gln Gln Tyr Gly Ser Ser Pro Phe1
5429107PRTHomo Sapien 429Glu Ile Val Leu Thr Gln Ser Pro Gly Thr
Leu Ser Leu Ser Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala
Ser Gln Ser Val Ser Ser Ser 20 25 30Tyr Leu Ala Trp Tyr Gln Gln Lys
Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45Ile Tyr Gly Ala Ser Ser Arg
Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60Gly Ser Gly Ser Gly Thr
Asp Phe Thr Leu Thr Ile Arg Arg Leu Glu65 70 75 80Pro Glu Asp Phe
Ala Val Tyr Tyr Cys Gln Gln Tyr Gly Ser Ser Pro 85 90 95Phe Phe Gly
Pro Gly Thr Lys Val Asp Ile Lys 100 105430321DNAHomo Sapien
430gaaattgtgt tgacgcagtc tccaggcacc ctgtctttgt ctccagggga
aagagccacc 60ctctcctgca gggccagtca gagtgttagc agcagctact tagcctggta
ccagcagaaa 120cctggccagg ctcccaggct cctcatatat ggtgcatcca
gcagggccac tggcatccca 180gacaggttca gtggcagtgg gtctgggaca
gacttcactc tcaccatcag aagactggag 240cctgaagatt ttgcagtgta
ttactgtcag cagtatggta gttcaccatt cttcggccct 300gggaccaaag
tggatatcaa a 321431323DNAHomo Sapien 431gaaattgtgt tgacgcagtc
tccaggcacc ctgtctttgt ctccagggga aagagccacc 60ctctcctgca gggccagtca
gagtgttagc agcagctact tagcctggta ccagcagaaa 120cctggccagg
ctcccaggct cctcatatat ggtgcatcca gcagggccac tggcatccca
180gacaggttca gtggcagtgg gtctgggaca gacttcactc tcaccatcag
aagactggag 240cctgaagatt ttgcagtgta ttactgtcag cagtatggta
gttcaccatt cacttcggcc 300ctgggaccaa agtggatatc aaa 323432214PRTHomo
Sapien 432Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser
Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val
Ser Ser Ser 20 25 30Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala
Pro Arg Leu Leu 35 40 45Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly Ile
Pro Asp Arg Phe Ser 50 55 60Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu
Thr Ile Arg Arg Leu Glu65 70 75 80Pro Glu Asp Phe Ala Val Tyr Tyr
Cys Gln Gln Tyr Gly Ser Ser Pro 85 90 95Phe Phe Gly Pro Gly Thr Lys
Val Asp Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile
Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser
Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys
Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150
155 160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu
Ser 165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His
Lys Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser
Pro Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys
210433642DNAHomo Sapien 433gaaattgtgt tgacgcagtc tccaggcacc
ctgtctttgt ctccagggga aagagccacc 60ctctcctgca gggccagtca gagtgttagc
agcagctact tagcctggta ccagcagaaa 120cctggccagg ctcccaggct
cctcatatat ggtgcatcca gcagggccac tggcatccca 180gacaggttca
gtggcagtgg gtctgggaca gacttcactc tcaccatcag aagactggag
240cctgaagatt ttgcagtgta ttactgtcag cagtatggta gttcaccatt
cttcggccct 300gggaccaaag tggatatcaa acgtacggtg gccgctccct
ccgtgttcat cttcccacct 360tccgacgagc agctgaagtc cggcaccgct
tctgtcgtgt gcctgctgaa caacttctac 420ccccgcgagg ccaaggtgca
gtggaaggtg gacaacgccc tgcagtccgg caactcccag 480gaatccgtga
ccgagcagga ctccaaggac agcacctact ccctgtcctc caccctgacc
540ctgtccaagg ccgactacga gaagcacaag gtgtacgcct gcgaagtgac
ccaccagggc 600ctgtctagcc ccgtgaccaa gtctttcaac cggggcgagt gt
642434644DNAHomo Sapien 434gaaattgtgt tgacgcagtc tccaggcacc
ctgtctttgt ctccagggga aagagccacc 60ctctcctgca gggccagtca gagtgttagc
agcagctact tagcctggta ccagcagaaa 120cctggccagg ctcccaggct
cctcatatat ggtgcatcca gcagggccac tggcatccca 180gacaggttca
gtggcagtgg gtctgggaca gacttcactc tcaccatcag aagactggag
240cctgaagatt ttgcagtgta ttactgtcag cagtatggta gttcaccatt
cacttcggcc 300ctgggaccaa agtggatatc aaacgtacgg tggccgctcc
ctccgtgttc atcttcccac 360cttccgacga gcagctgaag tccggcaccg
cttctgtcgt gtgcctgctg aacaacttct 420acccccgcga ggccaaggtg
cagtggaagg tggacaacgc cctgcagtcc ggcaactccc 480aggaatccgt
gaccgagcag gactccaagg acagcaccta ctccctgtcc tccaccctga
540ccctgtccaa ggccgactac gagaagcaca aggtgtacgc ctgcgaagtg
acccaccagg 600gcctgtctag ccccgtgacc aagtctttca accggggcga gtgt
6444358PRTHomo Sapien 435Gly Tyr Thr Phe Asn Ser Tyr Gly1
54368PRTHomo Sapien 436Ile Ser Val His Asn Gly Asn Thr1
543717PRTHomo Sapien 437Ala Arg Ala Gly Tyr Asp Ile Leu Thr Asp Phe
Ser Asp Ala Phe Asp1 5 10 15Ile438124PRTHomo Sapien 438Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Asn Ser Tyr 20 25 30Gly
Ile Ile Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Ser Val His Asn Gly Asn Thr Asn Cys Ala Gln Lys Leu
50 55 60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr Ser Thr Ala
Tyr65 70 75 80Met Glu Leu Arg Ser Leu Arg Thr Asp Asp Thr Ala Val
Tyr Tyr Cys 85 90 95Ala Arg Ala Gly Tyr Asp Ile Leu Thr Asp Phe Ser
Asp Ala Phe Asp 100 105 110Ile Trp Gly His Gly Thr Met Val Thr Val
Ser Ser 115 120439372DNAHomo Sapien 439caggttcagt tggtgcagtc
tggagctgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta
cacctttaat agttatggta tcatctgggt gcgacaggcc 120cctggacaag
ggcttgagtg gatgggatgg atcagcgttc acaatggtaa cacaaactgt
180gcacagaagc tccagggtag agtcaccatg accacagaca catccacgag
cacagcctac 240atggagctga ggagcctgag aactgacgac acggccgtgt
attactgtgc gagagcgggt 300tacgatattt tgactgattt ttccgatgct
tttgatatct ggggccacgg gacaatggtc 360accgtctctt ca 372440454PRTHomo
Sapien 440Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro
Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe
Asn Ser Tyr 20 25 30Gly Ile Ile Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45Gly Trp Ile Ser Val His Asn Gly Asn Thr Asn
Cys Ala Gln Lys Leu 50 55 60Gln Gly Arg Val Thr Met Thr Thr Asp Thr
Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu Arg Ser Leu Arg Thr
Asp Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ala Gly Tyr Asp Ile
Leu Thr Asp Phe Ser Asp Ala Phe Asp 100 105 110Ile Trp Gly His Gly
Thr Met Val Thr Val Ser Ser Ala Ser Thr Lys 115 120 125Gly Pro Ser
Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly 130 135 140Gly
Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro145 150
155 160Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His
Thr 165 170 175Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu
Ser Ser Val 180 185 190Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln
Thr Tyr Ile Cys Asn 195 200 205Val Asn His Lys Pro Ser Asn Thr Lys
Val Asp Lys Lys Val Glu Pro 210 215 220Lys Ser Cys Asp Lys Thr His
Thr Cys Pro Pro Cys Pro Ala Pro Glu225 230 235 240Leu Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 245 250 255Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 260 265
270Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
275 280 285Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn 290 295 300Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp305 310 315 320Leu Asn Gly Lys Glu Tyr Lys Cys Lys
Val Ser Asn Lys Ala Leu Pro 325 330 335Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu 340 345 350Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn 355 360 365Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 370 375 380Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr385 390
395 400Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys 405 410 415Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys 420 425 430Ser Val Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu 435 440 445Ser Leu Ser Pro Gly Lys
4504411368DNAHomo Sapien 441caggttcagt tggtgcagtc tggagctgag
gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttaat
agttatggta tcatctgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcagcgttc acaatggtaa cacaaactgt 180gcacagaagc
tccagggtag agtcaccatg accacagaca catccacgag cacagcctac
240atggagctga ggagcctgag aactgacgac acggccgtgt attactgtgc
gagagcgggt 300tacgatattt tgactgattt ttccgatgct tttgatatct
ggggccacgg gacaatggtc 360accgtctctt cagccagcac caagggcccc
tctgtgttcc ctctggcccc ttccagcaag 420tccacctctg gcggaacagc
cgctctgggc tgcctcgtga aggactactt ccccgagcct 480gtgaccgtgt
cctggaactc tggcgctctg accagcggag tgcacacctt ccctgctgtg
540ctgcagtcct ccggcctgta ctccctgtcc tccgtcgtga ccgtgccttc
cagctctctg 600ggcacccaga cctacatctg caacgtgaac cacaagccct
ccaacaccaa ggtggacaag 660aaggtggaac ccaagtcctg cgacaagacc
cacacctgtc ccccttgtcc tgcccctgaa 720ctgctgggcg gaccttccgt
gttcctgttc cccccaaagc ccaaggacac cctgatgatc 780tcccggaccc
ccgaagtgac ctgcgtggtg gtggatgtgt cccacgagga ccctgaagtg
840aagttcaatt ggtacgtgga cggcgtggaa gtgcacaacg ccaagaccaa
gcctagagag 900gaacagtaca actccaccta ccgggtggtg tccgtgctga
ccgtgctgca ccaggattgg 960ctgaacggca aagagtacaa gtgcaaggtg
tccaacaagg ccctgcctgc ccccatcgaa 1020aagaccatct ccaaggccaa
gggccagccc cgggaacccc aggtgtacac actgccccct 1080agcagggacg
agctgaccaa gaaccaggtg tccctgacct gtctcgtgaa aggcttctac
1140ccctccgata tcgccgtgga atgggagtcc aacggccagc ctgagaacaa
ctacaagacc 1200accccccctg tgctggactc cgacggctca ttcttcctgt
acagcaagct gacagtggac 1260aagtcccggt ggcagcaggg caacgtgttc
tcctgctccg tgatgcacga ggccctgcac 1320aaccactaca cccagaagtc
cctgtccctg agccccggca agtgatga 13684426PRTHomo Sapien 442Gln Asn
Ile Asn Asn Phe1 54433PRTHomo Sapien 443Ala Ala Ser14448PRTHomo
Sapien 444Gln Gln Ser Tyr Gly Ile Pro Trp1 5445106PRTHomo Sapien
445Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Asn Ile Asn Asn
Phe 20 25 30Leu Asn Trp Tyr Gln Gln Lys Glu Gly Lys Gly Pro Lys Leu
Leu Ile 35 40 45Tyr Ala Ala Ser Ser Leu Gln Arg Gly Ile Pro Ser Thr
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Ile Cys Gln Gln
Ser Tyr Gly Ile Pro Trp 85 90 95Val Gly Gln Gly Thr Lys Val Glu Ile
Lys 100 105446318DNAHomo Sapien 446gacatccaga tgacccagtc tccatcctcc
ctgtctgcat ctgtaggaga cagagtcacc 60atcacttgcc gggcaagtca gaacattaat
aactttttaa attggtatca gcagaaagaa 120gggaaaggcc ctaagctcct
gatctatgca gcatccagtt tgcaaagagg gataccatca 180acgttcagtg
gcagtggatc tgggacagac ttcactctca ccatcagcag tctgcaacct
240gaagattttg caacttacat ctgtcaacag agctacggta tcccgtgggt
cggccaaggg 300accaaggtgg aaatcaaa 318447213PRTHomo Sapien 447Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Asn Ile Asn Asn Phe
20 25 30Leu Asn Trp Tyr Gln Gln Lys Glu Gly Lys Gly Pro Lys Leu Leu
Ile 35 40 45Tyr Ala Ala Ser Ser Leu Gln Arg Gly Ile Pro Ser Thr Phe
Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser
Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Ile Cys Gln Gln Ser
Tyr Gly Ile Pro Trp 85 90 95Val Gly Gln Gly Thr Lys Val Glu Ile Lys
Arg Thr Val Ala Ala Pro 100 105 110Ser Val Phe Ile Phe Pro Pro Ser
Asp Glu Gln Leu Lys Ser Gly Thr 115 120 125Ala Ser Val Val Cys Leu
Leu Asn Asn Phe Tyr Pro Arg Glu Ala Lys 130 135 140Val Gln Trp Lys
Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln Glu145 150 155 160Ser
Val
Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser Ser 165 170
175Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr Ala
180 185 190Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys
Ser Phe 195 200 205Asn Arg Gly Glu Cys 210448639DNAHomo Sapien
448gacatccaga tgacccagtc tccatcctcc ctgtctgcat ctgtaggaga
cagagtcacc 60atcacttgcc gggcaagtca gaacattaat aactttttaa attggtatca
gcagaaagaa 120gggaaaggcc ctaagctcct gatctatgca gcatccagtt
tgcaaagagg gataccatca 180acgttcagtg gcagtggatc tgggacagac
ttcactctca ccatcagcag tctgcaacct 240gaagattttg caacttacat
ctgtcaacag agctacggta tcccgtgggt cggccaaggg 300accaaggtgg
aaatcaaacg tacggtggcc gctccctccg tgttcatctt cccaccttcc
360gacgagcagc tgaagtccgg caccgcttct gtcgtgtgcc tgctgaacaa
cttctacccc 420cgcgaggcca aggtgcagtg gaaggtggac aacgccctgc
agtccggcaa ctcccaggaa 480tccgtgaccg agcaggactc caaggacagc
acctactccc tgtcctccac cctgaccctg 540tccaaggccg actacgagaa
gcacaaggtg tacgcctgcg aagtgaccca ccagggcctg 600tctagccccg
tgaccaagtc tttcaaccgg ggcgagtgt 6394498PRTHomo Sapien 449Gly Phe
Thr Phe Ser Asp Tyr Phe1 54508PRTHomo Sapien 450Ile Ser Ser Ser Gly
Ser Thr Ile1 545121PRTHomo Sapien 451Ala Arg Asp His Tyr Asp Gly
Ser Gly Ile Tyr Pro Leu Tyr Tyr Tyr1 5 10 15Tyr Gly Leu Asp Val
20452128PRTHomo Sapien 452Gln Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Asp Tyr 20 25 30Phe Met Ser Trp Ile Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Ile 35 40 45Ser Tyr Ile Ser Ser Ser Gly
Ser Thr Ile Tyr Tyr Ala Asp Ser Val 50 55 60Arg Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Tyr Ser Leu Tyr65 70 75 80Leu Gln Met Asn
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp
His Tyr Asp Gly Ser Gly Ile Tyr Pro Leu Tyr Tyr Tyr 100 105 110Tyr
Gly Leu Asp Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120
125453384DNAHomo Sapien 453caggtgcagc tggtggagtc tgggggaggc
ttggtcaagc ctggagggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt
gactacttca tgagctggat ccgccaggcg 120ccagggaagg ggctggagtg
gatttcatac attagttcta gtggtagtac catatactac 180gcagactctg
tgaggggccg attcaccatc tccagggaca acgccaagta ctcactgtat
240ctgcaaatga acagcctgag atccgaggac acggccgtgt attactgtgc
gagagatcac 300tacgatggtt cggggattta tcccctctac tactattacg
gtttggacgt ctggggccag 360gggaccacgg tcaccgtctc ctca
384454458PRTHomo Sapien 454Gln Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Asp Tyr 20 25 30Phe Met Ser Trp Ile Arg Gln Ala
Pro Gly Lys Gly Leu Glu Trp Ile 35 40 45Ser Tyr Ile Ser Ser Ser Gly
Ser Thr Ile Tyr Tyr Ala Asp Ser Val 50 55 60Arg Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Tyr Ser Leu Tyr65 70 75 80Leu Gln Met Asn
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp
His Tyr Asp Gly Ser Gly Ile Tyr Pro Leu Tyr Tyr Tyr 100 105 110Tyr
Gly Leu Asp Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120
125Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys
130 135 140Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val Lys
Asp Tyr145 150 155 160Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser
Gly Ala Leu Thr Ser 165 170 175Gly Val His Thr Phe Pro Ala Val Leu
Gln Ser Ser Gly Leu Tyr Ser 180 185 190Leu Ser Ser Val Val Thr Val
Pro Ser Ser Ser Leu Gly Thr Gln Thr 195 200 205Tyr Ile Cys Asn Val
Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys 210 215 220Lys Val Glu
Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys225 230 235
240Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro
245 250 255Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val
Thr Cys 260 265 270Val Val Val Asp Val Ser His Glu Asp Pro Glu Val
Lys Phe Asn Trp 275 280 285Tyr Val Asp Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu 290 295 300Glu Gln Tyr Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu305 310 315 320His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn 325 330 335Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly 340 345 350Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu 355 360
365Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr
370 375 380Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn385 390 395 400Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe 405 410 415Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn 420 425 430Val Phe Ser Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr 435 440 445Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 450 4554551380DNAHomo Sapien 455caggtgcagc
tggtggagtc tgggggaggc ttggtcaagc ctggagggtc cctgagactc 60tcctgtgcag
cctctggatt caccttcagt gactacttca tgagctggat ccgccaggcg
120ccagggaagg ggctggagtg gatttcatac attagttcta gtggtagtac
catatactac 180gcagactctg tgaggggccg attcaccatc tccagggaca
acgccaagta ctcactgtat 240ctgcaaatga acagcctgag atccgaggac
acggccgtgt attactgtgc gagagatcac 300tacgatggtt cggggattta
tcccctctac tactattacg gtttggacgt ctggggccag 360gggaccacgg
tcaccgtctc ctcagccagc accaagggcc cctctgtgtt ccctctggcc
420ccttccagca agtccacctc tggcggaaca gccgctctgg gctgcctcgt
gaaggactac 480ttccccgagc ctgtgaccgt gtcctggaac tctggcgctc
tgaccagcgg agtgcacacc 540ttccctgctg tgctgcagtc ctccggcctg
tactccctgt cctccgtcgt gaccgtgcct 600tccagctctc tgggcaccca
gacctacatc tgcaacgtga accacaagcc ctccaacacc 660aaggtggaca
agaaggtgga acccaagtcc tgcgacaaga cccacacctg tcccccttgt
720cctgcccctg aactgctggg cggaccttcc gtgttcctgt tccccccaaa
gcccaaggac 780accctgatga tctcccggac ccccgaagtg acctgcgtgg
tggtggatgt gtcccacgag 840gaccctgaag tgaagttcaa ttggtacgtg
gacggcgtgg aagtgcacaa cgccaagacc 900aagcctagag aggaacagta
caactccacc taccgggtgg tgtccgtgct gaccgtgctg 960caccaggatt
ggctgaacgg caaagagtac aagtgcaagg tgtccaacaa ggccctgcct
1020gcccccatcg aaaagaccat ctccaaggcc aagggccagc cccgggaacc
ccaggtgtac 1080acactgcccc ctagcaggga cgagctgacc aagaaccagg
tgtccctgac ctgtctcgtg 1140aaaggcttct acccctccga tatcgccgtg
gaatgggagt ccaacggcca gcctgagaac 1200aactacaaga ccaccccccc
tgtgctggac tccgacggct cattcttcct gtacagcaag 1260ctgacagtgg
acaagtcccg gtggcagcag ggcaacgtgt tctcctgctc cgtgatgcac
1320gaggccctgc acaaccacta cacccagaag tccctgtccc tgagccccgg
caagtgatga 138045611PRTHomo Sapien 456Gln Ser Leu Leu His Ser Asn
Gly Tyr Asn Tyr1 5 104573PRTHomo Sapien 457Leu Gly Ser14589PRTHomo
Sapien 458Met Gln Ala Leu Gln Thr Pro Arg Ser1 5459111PRTHomo
Sapien 459Ile Val Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro
Gly Glu1 5 10 15Pro Ala Ser Ile Ser Cys Arg Ser Ser Gln Ser Leu Leu
His Ser Asn 20 25 30Gly Tyr Asn Tyr Leu Asp Tyr Tyr Leu Gln Lys Pro
Gly Gln Ser Pro 35 40 45Gln Leu Leu Ile Tyr Leu Gly Ser Tyr Arg Ala
Ser Gly Val Pro Asp 50 55 60Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp
Phe Thr Leu Lys Ile Ser65 70 75 80Arg Val Glu Ala Glu Asp Val Gly
Val Tyr Tyr Cys Met Gln Ala Leu 85 90 95Gln Thr Pro Arg Ser Phe Gly
Gln Gly Thr Thr Leu Glu Ile Lys 100 105 110460333DNAHomo Sapien
460attgtgatga ctcagtctcc actctcccta cccgtcaccc ctggagagcc
ggcctccatc 60tcctgcaggt ctagtcagag cctcctgcat agtaatggat acaactattt
ggattattac 120ctgcagaagc cagggcagtc tccacagctc ctgatctatt
tgggttctta tcgggcctcc 180ggggtccctg acaggttcag tggcagtgga
tcaggcacag attttacact gaaaatcagc 240agagtggagg ctgaggatgt
tggggtttat tactgcatgc aagctctaca aactcctcgc 300agttttggcc
aggggaccac gctggagatc aaa 333461218PRTHomo Sapien 461Ile Val Met
Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly Glu1 5 10 15Pro Ala
Ser Ile Ser Cys Arg Ser Ser Gln Ser Leu Leu His Ser Asn 20 25 30Gly
Tyr Asn Tyr Leu Asp Tyr Tyr Leu Gln Lys Pro Gly Gln Ser Pro 35 40
45Gln Leu Leu Ile Tyr Leu Gly Ser Tyr Arg Ala Ser Gly Val Pro Asp
50 55 60Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile
Ser65 70 75 80Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Met
Gln Ala Leu 85 90 95Gln Thr Pro Arg Ser Phe Gly Gln Gly Thr Thr Leu
Glu Ile Lys Arg 100 105 110Thr Val Ala Ala Pro Ser Val Phe Ile Phe
Pro Pro Ser Asp Glu Gln 115 120 125Leu Lys Ser Gly Thr Ala Ser Val
Val Cys Leu Leu Asn Asn Phe Tyr 130 135 140Pro Arg Glu Ala Lys Val
Gln Trp Lys Val Asp Asn Ala Leu Gln Ser145 150 155 160Gly Asn Ser
Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr 165 170 175Tyr
Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys 180 185
190His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
195 200 205Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 210
215462654DNAHomo Sapien 462attgtgatga ctcagtctcc actctcccta
cccgtcaccc ctggagagcc ggcctccatc 60tcctgcaggt ctagtcagag cctcctgcat
agtaatggat acaactattt ggattattac 120ctgcagaagc cagggcagtc
tccacagctc ctgatctatt tgggttctta tcgggcctcc 180ggggtccctg
acaggttcag tggcagtgga tcaggcacag attttacact gaaaatcagc
240agagtggagg ctgaggatgt tggggtttat tactgcatgc aagctctaca
aactcctcgc 300agttttggcc aggggaccac gctggagatc aaacgtacgg
tggccgctcc ctccgtgttc 360atcttcccac cttccgacga gcagctgaag
tccggcaccg cttctgtcgt gtgcctgctg 420aacaacttct acccccgcga
ggccaaggtg cagtggaagg tggacaacgc cctgcagtcc 480ggcaactccc
aggaatccgt gaccgagcag gactccaagg acagcaccta ctccctgtcc
540tccaccctga ccctgtccaa ggccgactac gagaagcaca aggtgtacgc
ctgcgaagtg 600acccaccagg gcctgtctag ccccgtgacc aagtctttca
accggggcga gtgt 65446310PRTHomo Sapien 463Gly Phe Ser Leu Ser Thr
Thr Gly Val Gly1 5 104647PRTHomo Sapien 464Ile Tyr Trp Asp Asp Asp
Lys1 546516PRTHomo Sapien 465Thr His Gly Tyr Gly Ser Ala Ser Tyr
Tyr His Tyr Gly Met Asp Val1 5 10 15466124PRTHomo Sapien 466Gln Ile
Thr Leu Lys Glu Ser Gly Pro Thr Leu Val Lys Pro Thr Gln1 5 10 15Thr
Leu Thr Leu Thr Cys Thr Phe Ser Gly Phe Ser Leu Ser Thr Thr 20 25
30Gly Val Gly Val Gly Trp Ile Arg Gln Pro Pro Gly Lys Ala Leu Glu
35 40 45Trp Leu Ala Val Ile Tyr Trp Asp Asp Asp Lys Arg Tyr Ser Pro
Ser 50 55 60Leu Lys Ser Arg Leu Thr Ile Thr Lys Asp Thr Ser Lys Asn
Gln Val65 70 75 80Val Leu Thr Met Thr Asn Met Asp Pro Val Asp Thr
Ala Thr Tyr Phe 85 90 95Cys Thr His Gly Tyr Gly Ser Ala Ser Tyr Tyr
His Tyr Gly Met Asp 100 105 110Val Trp Gly Gln Gly Thr Thr Val Thr
Val Ser Ser 115 120467372DNAHomo Sapien 467cagatcacct tgaaggagtc
tggtcctacg ctggtgaaac ccacacagac cctcacgctg 60acctgcacct tctctgggtt
ctcactcagc actactggag tgggtgtggg ctggatccgt 120cagcccccag
gaaaggccct ggagtggctt gcagtcattt attgggatga tgataagcgc
180tacagcccat ctctgaagag cagactcacc atcaccaagg acacctccaa
aaaccaggtg 240gtccttacaa tgaccaacat ggaccctgtg gacacagcca
catatttctg tacacacgga 300tatggttcgg cgagttatta ccactacggt
atggacgtct ggggccaagg gaccacggtc 360accgtctcct ca 372468454PRTHomo
Sapien 468Gln Ile Thr Leu Lys Glu Ser Gly Pro Thr Leu Val Lys Pro
Thr Gln1 5 10 15Thr Leu Thr Leu Thr Cys Thr Phe Ser Gly Phe Ser Leu
Ser Thr Thr 20 25 30Gly Val Gly Val Gly Trp Ile Arg Gln Pro Pro Gly
Lys Ala Leu Glu 35 40 45Trp Leu Ala Val Ile Tyr Trp Asp Asp Asp Lys
Arg Tyr Ser Pro Ser 50 55 60Leu Lys Ser Arg Leu Thr Ile Thr Lys Asp
Thr Ser Lys Asn Gln Val65 70 75 80Val Leu Thr Met Thr Asn Met Asp
Pro Val Asp Thr Ala Thr Tyr Phe 85 90 95Cys Thr His Gly Tyr Gly Ser
Ala Ser Tyr Tyr His Tyr Gly Met Asp 100 105 110Val Trp Gly Gln Gly
Thr Thr Val Thr Val Ser Ser Ala Ser Thr Lys 115 120 125Gly Pro Ser
Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly 130 135 140Gly
Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro145 150
155 160Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His
Thr 165 170 175Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu
Ser Ser Val 180 185 190Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln
Thr Tyr Ile Cys Asn 195 200 205Val Asn His Lys Pro Ser Asn Thr Lys
Val Asp Lys Lys Val Glu Pro 210 215 220Lys Ser Cys Asp Lys Thr His
Thr Cys Pro Pro Cys Pro Ala Pro Glu225 230 235 240Leu Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 245 250 255Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 260 265
270Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
275 280 285Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn 290 295 300Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp305 310 315 320Leu Asn Gly Lys Glu Tyr Lys Cys Lys
Val Ser Asn Lys Ala Leu Pro 325 330 335Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu 340 345 350Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn 355 360 365Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 370 375 380Ala
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr385 390
395 400Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys 405 410 415Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys 420 425 430Ser Val Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu 435 440 445Ser Leu Ser Pro Gly Lys
4504691368DNAHomo Sapien 469cagatcacct tgaaggagtc tggtcctacg
ctggtgaaac ccacacagac cctcacgctg 60acctgcacct tctctgggtt ctcactcagc
actactggag tgggtgtggg ctggatccgt 120cagcccccag gaaaggccct
ggagtggctt gcagtcattt attgggatga tgataagcgc 180tacagcccat
ctctgaagag cagactcacc atcaccaagg acacctccaa aaaccaggtg
240gtccttacaa tgaccaacat ggaccctgtg gacacagcca catatttctg
tacacacgga 300tatggttcgg cgagttatta ccactacggt atggacgtct
ggggccaagg gaccacggtc 360accgtctcct cagccagcac caagggcccc
tctgtgttcc ctctggcccc ttccagcaag 420tccacctctg gcggaacagc
cgctctgggc tgcctcgtga aggactactt ccccgagcct 480gtgaccgtgt
cctggaactc tggcgctctg accagcggag tgcacacctt ccctgctgtg
540ctgcagtcct ccggcctgta ctccctgtcc tccgtcgtga ccgtgccttc
cagctctctg 600ggcacccaga cctacatctg caacgtgaac cacaagccct
ccaacaccaa ggtggacaag 660aaggtggaac ccaagtcctg cgacaagacc
cacacctgtc ccccttgtcc tgcccctgaa 720ctgctgggcg gaccttccgt
gttcctgttc cccccaaagc ccaaggacac cctgatgatc 780tcccggaccc
ccgaagtgac ctgcgtggtg gtggatgtgt cccacgagga ccctgaagtg
840aagttcaatt
ggtacgtgga cggcgtggaa gtgcacaacg ccaagaccaa gcctagagag
900gaacagtaca actccaccta ccgggtggtg tccgtgctga ccgtgctgca
ccaggattgg 960ctgaacggca aagagtacaa gtgcaaggtg tccaacaagg
ccctgcctgc ccccatcgaa 1020aagaccatct ccaaggccaa gggccagccc
cgggaacccc aggtgtacac actgccccct 1080agcagggacg agctgaccaa
gaaccaggtg tccctgacct gtctcgtgaa aggcttctac 1140ccctccgata
tcgccgtgga atgggagtcc aacggccagc ctgagaacaa ctacaagacc
1200accccccctg tgctggactc cgacggctca ttcttcctgt acagcaagct
gacagtggac 1260aagtcccggt ggcagcaggg caacgtgttc tcctgctccg
tgatgcacga ggccctgcac 1320aaccactaca cccagaagtc cctgtccctg
agccccggca agtgatga 13684706PRTHomo Sapien 470Gln Ser Val Thr Asn
Tyr1 54713PRTHomo Sapien 471Asp Ala Ser14729PRTHomo Sapien 472Gln
His Arg Ser Asn Trp Pro Leu Thr1 5473107PRTHomo Sapien 473Glu Ile
Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly1 5 10 15Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Thr Asn Tyr 20 25
30Leu Ala Trp His Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45Tyr Asp Ala Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser
Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu
Glu Pro65 70 75 80Glu Asp Phe Ala Val Tyr Tyr Cys Gln His Arg Ser
Asn Trp Pro Leu 85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys
100 105474322DNAHomo Sapien 474gaaattgtat tgacacagtc tccagccacc
ctgtctttgt ctccagggga aagagccacc 60ctctcctgca gggccagtca gagtgttacc
aactacttag cctggcacca acagaaacct 120ggccaggctc ccaggctcct
catctatgat gcatccaaca gggccactgg catcccagcc 180aggttcagtg
gcagtgggtc tgggacagac ttcactctca ccatcagcag cctagagcct
240gaagattttg cagtttatta ctgtcagcac cgtagcaact ggcctctcac
tttcggcgga 300gggaccaagg tggagatcaa ac 322475214PRTHomo Sapien
475Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly1
5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Thr Asn
Tyr 20 25 30Leu Ala Trp His Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu
Leu Ile 35 40 45Tyr Asp Ala Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Glu Pro65 70 75 80Glu Asp Phe Ala Val Tyr Tyr Cys Gln His
Arg Ser Asn Trp Pro Leu 85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro
Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val
Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln
Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155
160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys 210476643DNAHomo
Sapien 476gaaattgtat tgacacagtc tccagccacc ctgtctttgt ctccagggga
aagagccacc 60ctctcctgca gggccagtca gagtgttacc aactacttag cctggcacca
acagaaacct 120ggccaggctc ccaggctcct catctatgat gcatccaaca
gggccactgg catcccagcc 180aggttcagtg gcagtgggtc tgggacagac
ttcactctca ccatcagcag cctagagcct 240gaagattttg cagtttatta
ctgtcagcac cgtagcaact ggcctctcac tttcggcgga 300gggaccaagg
tggagatcaa accgtacggt ggccgctccc tccgtgttca tcttcccacc
360ttccgacgag cagctgaagt ccggcaccgc ttctgtcgtg tgcctgctga
acaacttcta 420cccccgcgag gccaaggtgc agtggaaggt ggacaacgcc
ctgcagtccg gcaactccca 480ggaatccgtg accgagcagg actccaagga
cagcacctac tccctgtcct ccaccctgac 540cctgtccaag gccgactacg
agaagcacaa ggtgtacgcc tgcgaagtga cccaccaggg 600cctgtctagc
cccgtgacca agtctttcaa ccggggcgag tgt 64347710PRTHomo Sapien 477Gly
Phe Ser Leu Ser Thr Ser Gly Val Gly1 5 104787PRTHomo Sapien 478Ile
Tyr Trp Asp Asp Asp Lys1 547916PRTHomo Sapien 479Thr His Gly Tyr
Gly Ser Ala Ser Tyr Tyr His Tyr Gly Met Asp Val1 5 10
15480124PRTHomo Sapien 480Gln Ile Thr Leu Lys Glu Ser Gly Pro Thr
Leu Val Lys Pro Thr Gln1 5 10 15Thr Leu Thr Leu Thr Cys Thr Phe Ser
Gly Phe Ser Leu Ser Thr Ser 20 25 30Gly Val Gly Val Gly Trp Ile Arg
Gln Pro Pro Gly Lys Ala Leu Glu 35 40 45Trp Leu Ala Val Ile Tyr Trp
Asp Asp Asp Lys Arg Tyr Ser Pro Ser 50 55 60Leu Lys Ser Arg Leu Thr
Ile Thr Lys Asp Thr Ser Lys Asn Gln Val65 70 75 80Val Leu Thr Met
Thr Asn Met Asp Pro Val Asp Thr Ala Thr Tyr Phe 85 90 95Cys Thr His
Gly Tyr Gly Ser Ala Ser Tyr Tyr His Tyr Gly Met Asp 100 105 110Val
Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120481372DNAHomo
Sapien 481cagatcacct tgaaggagtc tggtcctacg ctggtgaaac ccacacagac
cctcacgctg 60acctgcacct tctctgggtt ctcactcagc actagtggag tgggtgtggg
ctggatccgt 120cagcccccag gaaaggccct ggagtggctt gcagtcattt
attgggatga tgataagcgc 180tacagcccat ctctgaagag caggctcacc
atcaccaagg acacctccaa aaaccaggtg 240gtccttacaa tgaccaacat
ggaccctgtg gacacagcca catatttctg tacacacgga 300tatggttcgg
cgagttatta ccactacggt atggacgtct ggggccaagg gaccacggtc
360accgtctcct ca 372482454PRTHomo Sapien 482Gln Ile Thr Leu Lys Glu
Ser Gly Pro Thr Leu Val Lys Pro Thr Gln1 5 10 15Thr Leu Thr Leu Thr
Cys Thr Phe Ser Gly Phe Ser Leu Ser Thr Ser 20 25 30Gly Val Gly Val
Gly Trp Ile Arg Gln Pro Pro Gly Lys Ala Leu Glu 35 40 45Trp Leu Ala
Val Ile Tyr Trp Asp Asp Asp Lys Arg Tyr Ser Pro Ser 50 55 60Leu Lys
Ser Arg Leu Thr Ile Thr Lys Asp Thr Ser Lys Asn Gln Val65 70 75
80Val Leu Thr Met Thr Asn Met Asp Pro Val Asp Thr Ala Thr Tyr Phe
85 90 95Cys Thr His Gly Tyr Gly Ser Ala Ser Tyr Tyr His Tyr Gly Met
Asp 100 105 110Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser Ala
Ser Thr Lys 115 120 125Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser
Lys Ser Thr Ser Gly 130 135 140Gly Thr Ala Ala Leu Gly Cys Leu Val
Lys Asp Tyr Phe Pro Glu Pro145 150 155 160Val Thr Val Ser Trp Asn
Ser Gly Ala Leu Thr Ser Gly Val His Thr 165 170 175Phe Pro Ala Val
Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val 180 185 190Val Thr
Val Pro Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn 195 200
205Val Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro
210 215 220Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala
Pro Glu225 230 235 240Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp 245 250 255Thr Leu Met Ile Ser Arg Thr Pro Glu
Val Thr Cys Val Val Val Asp 260 265 270Val Ser His Glu Asp Pro Glu
Val Lys Phe Asn Trp Tyr Val Asp Gly 275 280 285Val Glu Val His Asn
Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn 290 295 300Ser Thr Tyr
Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp305 310 315
320Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro
325 330 335Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu 340 345 350Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu
Leu Thr Lys Asn 355 360 365Gln Val Ser Leu Thr Cys Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile 370 375 380Ala Val Glu Trp Glu Ser Asn Gly
Gln Pro Glu Asn Asn Tyr Lys Thr385 390 395 400Thr Pro Pro Val Leu
Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys 405 410 415Leu Thr Val
Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys 420 425 430Ser
Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu 435 440
445Ser Leu Ser Pro Gly Lys 4504831368DNAHomo Sapien 483cagatcacct
tgaaggagtc tggtcctacg ctggtgaaac ccacacagac cctcacgctg 60acctgcacct
tctctgggtt ctcactcagc actagtggag tgggtgtggg ctggatccgt
120cagcccccag gaaaggccct ggagtggctt gcagtcattt attgggatga
tgataagcgc 180tacagcccat ctctgaagag caggctcacc atcaccaagg
acacctccaa aaaccaggtg 240gtccttacaa tgaccaacat ggaccctgtg
gacacagcca catatttctg tacacacgga 300tatggttcgg cgagttatta
ccactacggt atggacgtct ggggccaagg gaccacggtc 360accgtctcct
cagccagcac caagggcccc tctgtgttcc ctctggcccc ttccagcaag
420tccacctctg gcggaacagc cgctctgggc tgcctcgtga aggactactt
ccccgagcct 480gtgaccgtgt cctggaactc tggcgctctg accagcggag
tgcacacctt ccctgctgtg 540ctgcagtcct ccggcctgta ctccctgtcc
tccgtcgtga ccgtgccttc cagctctctg 600ggcacccaga cctacatctg
caacgtgaac cacaagccct ccaacaccaa ggtggacaag 660aaggtggaac
ccaagtcctg cgacaagacc cacacctgtc ccccttgtcc tgcccctgaa
720ctgctgggcg gaccttccgt gttcctgttc cccccaaagc ccaaggacac
cctgatgatc 780tcccggaccc ccgaagtgac ctgcgtggtg gtggatgtgt
cccacgagga ccctgaagtg 840aagttcaatt ggtacgtgga cggcgtggaa
gtgcacaacg ccaagaccaa gcctagagag 900gaacagtaca actccaccta
ccgggtggtg tccgtgctga ccgtgctgca ccaggattgg 960ctgaacggca
aagagtacaa gtgcaaggtg tccaacaagg ccctgcctgc ccccatcgaa
1020aagaccatct ccaaggccaa gggccagccc cgggaacccc aggtgtacac
actgccccct 1080agcagggacg agctgaccaa gaaccaggtg tccctgacct
gtctcgtgaa aggcttctac 1140ccctccgata tcgccgtgga atgggagtcc
aacggccagc ctgagaacaa ctacaagacc 1200accccccctg tgctggactc
cgacggctca ttcttcctgt acagcaagct gacagtggac 1260aagtcccggt
ggcagcaggg caacgtgttc tcctgctccg tgatgcacga ggccctgcac
1320aaccactaca cccagaagtc cctgtccctg agccccggca agtgatga
13684846PRTHomo Sapien 484Gln Ser Val Thr Asn Tyr1 54853PRTHomo
Sapien 485Asp Ala Ser14869PRTHomo Sapien 486Gln Gln Arg Ser Asn Trp
Pro Leu Thr1 5487107PRTHomo Sapien 487Glu Ile Val Leu Thr Gln Ser
Pro Ala Thr Leu Ser Leu Ser Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser
Cys Arg Ala Ser Gln Ser Val Thr Asn Tyr 20 25 30Leu Ala Trp His Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45Tyr Asp Ala Ser
Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60Ser Gly Ser
Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro65 70 75 80Glu
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Arg Ser Asn Trp Pro Leu 85 90
95Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105488321DNAHomo
Sapien 488gaaattgtgt tgacacagtc tccagccacc ctgtctttgt ctccagggga
aagagccacc 60ctctcctgca gggccagtca gagtgttacc aactacttag cctggcacca
acagaaacct 120ggccaggctc ccaggctcct catctatgat gcatccaaca
gggccactgg catcccagcc 180aggttcagtg gcagtgggtc tgggacagac
ttcactctca ccatcagcag cctagagcct 240gaagattttg cagtttatta
ctgtcagcag cgtagcaact ggcctctcac tttcggcgga 300gggaccaagg
tggagatcaa a 321489214PRTHomo Sapien 489Glu Ile Val Leu Thr Gln Ser
Pro Ala Thr Leu Ser Leu Ser Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser
Cys Arg Ala Ser Gln Ser Val Thr Asn Tyr 20 25 30Leu Ala Trp His Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45Tyr Asp Ala Ser
Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60Ser Gly Ser
Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro65 70 75 80Glu
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Arg Ser Asn Trp Pro Leu 85 90
95Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala
100 105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys
Ser Gly 115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr
Pro Arg Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala Leu
Gln Ser Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln Asp
Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr Leu
Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys Glu
Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200 205Phe
Asn Arg Gly Glu Cys 210490642DNAHomo Sapien 490gaaattgtgt
tgacacagtc tccagccacc ctgtctttgt ctccagggga aagagccacc 60ctctcctgca
gggccagtca gagtgttacc aactacttag cctggcacca acagaaacct
120ggccaggctc ccaggctcct catctatgat gcatccaaca gggccactgg
catcccagcc 180aggttcagtg gcagtgggtc tgggacagac ttcactctca
ccatcagcag cctagagcct 240gaagattttg cagtttatta ctgtcagcag
cgtagcaact ggcctctcac tttcggcgga 300gggaccaagg tggagatcaa
acgtacggtg gccgctccct ccgtgttcat cttcccacct 360tccgacgagc
agctgaagtc cggcaccgct tctgtcgtgt gcctgctgaa caacttctac
420ccccgcgagg ccaaggtgca gtggaaggtg gacaacgccc tgcagtccgg
caactcccag 480gaatccgtga ccgagcagga ctccaaggac agcacctact
ccctgtcctc caccctgacc 540ctgtccaagg ccgactacga gaagcacaag
gtgtacgcct gcgaagtgac ccaccagggc 600ctgtctagcc ccgtgaccaa
gtctttcaac cggggcgagt gt 6424918PRTHomo Sapien 491Gly Phe Thr Phe
Ser Asp Tyr Tyr1 54928PRTHomo Sapien 492Ile Ser Ser Ser Gly Ser Thr
Ile1 549320PRTHomo Sapien 493Ala Arg Asp Phe Tyr Asp Ile Leu Thr
Asp Ser Pro Tyr Phe Tyr Tyr1 5 10 15Gly Val Asp Val 20494127PRTHomo
Sapien 494Gln Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Lys Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Ser Asp Tyr 20 25 30Tyr Met Ser Trp Ile Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ser Tyr Ile Ser Ser Ser Gly Ser Thr Ile Tyr
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala Lys Asn Ser Leu Tyr65 70 75 80Leu Gln Ile Asn Ser Leu Arg Ala
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Phe Tyr Asp Ile
Leu Thr Asp Ser Pro Tyr Phe Tyr Tyr 100 105 110Gly Val Asp Val Trp
Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120 125495381DNAHomo
Sapien 495caggtgcagc tggtggagtc tgggggaggc ttggtcaagc ctggagggtc
cctgagactc 60tcctgtgcag cctctggatt caccttcagt gactactaca tgagctggat
ccgccaggct 120ccagggaagg ggctggagtg ggtttcatac attagtagta
gtggtagtac catatactac 180gcagactctg tgaagggccg attcaccatc
tccagggaca acgccaagaa ctcactgtat 240ctgcaaatta acagcctgag
agccgaggac acggccgtgt attactgtgc gagagatttt 300tacgatattt
tgactgatag tccgtacttc tactacggtg tggacgtctg gggccaaggg
360accacggtca ccgtctcctc a 381496457PRTHomo Sapien 496Gln Val Gln
Leu Val Glu Ser Gly Gly Gly Leu Val Lys Pro Gly Gly1 5 10 15Ser Leu
Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asp Tyr 20 25 30Tyr
Met Ser Trp Ile Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Tyr Ile Ser Ser Ser Gly Ser Thr Ile Tyr Tyr Ala Asp Ser Val
50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu
Tyr65 70 75 80Leu Gln Ile Asn Ser Leu Arg Ala Glu Asp Thr Ala Val
Tyr Tyr Cys 85 90 95Ala Arg Asp Phe Tyr Asp Ile Leu Thr Asp Ser Pro
Tyr Phe Tyr Tyr 100 105 110Gly Val Asp Val Trp Gly Gln Gly Thr Thr
Val Thr Val Ser Ser Ala 115 120 125Ser Thr Lys Gly Pro Ser Val Phe
Pro Leu Ala Pro Ser Ser Lys Ser 130 135 140Thr Ser Gly Gly Thr Ala
Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe145 150 155 160Pro Glu Pro
Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly
165 170 175Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr
Ser Leu 180 185 190Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu Gly
Thr Gln Thr Tyr 195 200 205Ile Cys Asn Val Asn His Lys Pro Ser Asn
Thr Lys Val Asp Lys Lys 210 215 220Val Glu Pro Lys Ser Cys Asp Lys
Thr His Thr Cys Pro Pro Cys Pro225 230 235 240Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys 245 250 255Pro Lys Asp
Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val 260 265 270Val
Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr 275 280
285Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu
290 295 300Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val
Leu His305 310 315 320Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn Lys 325 330 335Ala Leu Pro Ala Pro Ile Glu Lys Thr
Ile Ser Lys Ala Lys Gly Gln 340 345 350Pro Arg Glu Pro Gln Val Tyr
Thr Leu Pro Pro Ser Arg Asp Glu Leu 355 360 365Thr Lys Asn Gln Val
Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro 370 375 380Ser Asp Ile
Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn385 390 395
400Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu
405 410 415Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly
Asn Val 420 425 430Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn
His Tyr Thr Gln 435 440 445Lys Ser Leu Ser Leu Ser Pro Gly Lys 450
4554971377DNAHomo Sapien 497caggtgcagc tggtggagtc tgggggaggc
ttggtcaagc ctggagggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt
gactactaca tgagctggat ccgccaggct 120ccagggaagg ggctggagtg
ggtttcatac attagtagta gtggtagtac catatactac 180gcagactctg
tgaagggccg attcaccatc tccagggaca acgccaagaa ctcactgtat
240ctgcaaatta acagcctgag agccgaggac acggccgtgt attactgtgc
gagagatttt 300tacgatattt tgactgatag tccgtacttc tactacggtg
tggacgtctg gggccaaggg 360accacggtca ccgtctcctc agccagcacc
aagggcccct ctgtgttccc tctggcccct 420tccagcaagt ccacctctgg
cggaacagcc gctctgggct gcctcgtgaa ggactacttc 480cccgagcctg
tgaccgtgtc ctggaactct ggcgctctga ccagcggagt gcacaccttc
540cctgctgtgc tgcagtcctc cggcctgtac tccctgtcct ccgtcgtgac
cgtgccttcc 600agctctctgg gcacccagac ctacatctgc aacgtgaacc
acaagccctc caacaccaag 660gtggacaaga aggtggaacc caagtcctgc
gacaagaccc acacctgtcc cccttgtcct 720gcccctgaac tgctgggcgg
accttccgtg ttcctgttcc ccccaaagcc caaggacacc 780ctgatgatct
cccggacccc cgaagtgacc tgcgtggtgg tggatgtgtc ccacgaggac
840cctgaagtga agttcaattg gtacgtggac ggcgtggaag tgcacaacgc
caagaccaag 900cctagagagg aacagtacaa ctccacctac cgggtggtgt
ccgtgctgac cgtgctgcac 960caggattggc tgaacggcaa agagtacaag
tgcaaggtgt ccaacaaggc cctgcctgcc 1020cccatcgaaa agaccatctc
caaggccaag ggccagcccc gggaacccca ggtgtacaca 1080ctgcccccta
gcagggacga gctgaccaag aaccaggtgt ccctgacctg tctcgtgaaa
1140ggcttctacc cctccgatat cgccgtggaa tgggagtcca acggccagcc
tgagaacaac 1200tacaagacca ccccccctgt gctggactcc gacggctcat
tcttcctgta cagcaagctg 1260acagtggaca agtcccggtg gcagcagggc
aacgtgttct cctgctccgt gatgcacgag 1320gccctgcaca accactacac
ccagaagtcc ctgtccctga gccccggcaa gtgatga 137749811PRTHomo Sapien
498Gln Ser Leu Leu His Ser Asn Gly Tyr Asn Tyr1 5 104993PRTHomo
Sapien 499Leu Gly Ser15009PRTHomo Sapien 500Met Gln Ala Leu Gln Thr
Pro Arg Thr1 5501112PRTHomo Sapien 501Asp Ile Val Met Thr Gln Ser
Pro Leu Ser Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile Ser
Cys Arg Ser Ser Gln Ser Leu Leu His Ser 20 25 30Asn Gly Tyr Asn Tyr
Leu Asp Trp Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Gln Leu Leu
Ile Tyr Leu Gly Ser Asn Arg Ala Ser Gly Val Pro 50 55 60Asp Arg Phe
Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75 80Ser
Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Met Gln Ala 85 90
95Leu Gln Thr Pro Arg Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys
100 105 110502336DNAHomo Sapien 502gatattgtga tgactcagtc tccactctcc
ctgcccgtca cccctggaga gccggcctcc 60atctcctgca ggtctagtca gagcctcctg
catagtaatg gatacaacta tttggattgg 120tacctgcaga agccagggca
gtctccacag ctcctgatct atttgggttc taatcgggcc 180tccggggtcc
ctgacaggtt cagtggcagt ggatcaggca cagattttac actgaaaatc
240agcagagtgg aggctgagga tgttggggtt tattactgca tgcaagctct
acaaactcct 300cggacgttcg gccaagggac caaggtggaa atcaaa
336503219PRTHomo Sapien 503Asp Ile Val Met Thr Gln Ser Pro Leu Ser
Leu Pro Val Thr Pro Gly1 5 10 15Glu Pro Ala Ser Ile Ser Cys Arg Ser
Ser Gln Ser Leu Leu His Ser 20 25 30Asn Gly Tyr Asn Tyr Leu Asp Trp
Tyr Leu Gln Lys Pro Gly Gln Ser 35 40 45Pro Gln Leu Leu Ile Tyr Leu
Gly Ser Asn Arg Ala Ser Gly Val Pro 50 55 60Asp Arg Phe Ser Gly Ser
Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65 70 75 80Ser Arg Val Glu
Ala Glu Asp Val Gly Val Tyr Tyr Cys Met Gln Ala 85 90 95Leu Gln Thr
Pro Arg Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105 110Arg
Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu 115 120
125Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe
130 135 140Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala
Leu Gln145 150 155 160Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser 165 170 175Thr Tyr Ser Leu Ser Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr Glu 180 185 190Lys His Lys Val Tyr Ala Cys
Glu Val Thr His Gln Gly Leu Ser Ser 195 200 205Pro Val Thr Lys Ser
Phe Asn Arg Gly Glu Cys 210 215504657DNAHomo Sapien 504gatattgtga
tgactcagtc tccactctcc ctgcccgtca cccctggaga gccggcctcc 60atctcctgca
ggtctagtca gagcctcctg catagtaatg gatacaacta tttggattgg
120tacctgcaga agccagggca gtctccacag ctcctgatct atttgggttc
taatcgggcc 180tccggggtcc ctgacaggtt cagtggcagt ggatcaggca
cagattttac actgaaaatc 240agcagagtgg aggctgagga tgttggggtt
tattactgca tgcaagctct acaaactcct 300cggacgttcg gccaagggac
caaggtggaa atcaaacgta cggtggccgc tccctccgtg 360ttcatcttcc
caccttccga cgagcagctg aagtccggca ccgcttctgt cgtgtgcctg
420ctgaacaact tctacccccg cgaggccaag gtgcagtgga aggtggacaa
cgccctgcag 480tccggcaact cccaggaatc cgtgaccgag caggactcca
aggacagcac ctactccctg 540tcctccaccc tgaccctgtc caaggccgac
tacgagaagc acaaggtgta cgcctgcgaa 600gtgacccacc agggcctgtc
tagccccgtg accaagtctt tcaaccgggg cgagtgt 657505239PRTHomo Sapien
505Phe Thr Val Thr Val Pro Lys Asp Leu Tyr Val Val Glu Tyr Gly Ser1
5 10 15Asn Met Thr Ile Glu Cys Lys Phe Pro Val Glu Lys Gln Leu Asp
Leu 20 25 30Ala Ala Leu Ile Val Tyr Trp Glu Met Glu Asp Lys Asn Ile
Ile Gln 35 40 45Phe Val His Gly Glu Glu Asp Leu Lys Val Gln His Ser
Ser Tyr Arg 50 55 60Gln Arg Ala Arg Leu Leu Lys Asp Gln Leu Ser Leu
Gly Asn Ala Ala65 70 75 80Leu Gln Ile Thr Asp Val Lys Leu Gln Asp
Ala Gly Val Tyr Arg Cys 85 90 95Met Ile Ser Tyr Gly Gly Ala Asp Tyr
Lys Arg Ile Thr Val Lys Val 100 105 110Asn Ala Pro Tyr Asn Lys Ile
Asn Gln Arg Ile Leu Val Val Asp Pro 115 120 125Val Thr Ser Glu His
Glu Leu Thr Cys Gln Ala Glu Gly Tyr Pro Lys 130 135 140Ala Glu Val
Ile Trp Thr Ser Ser Asp His Gln Val Leu Ser Gly Lys145 150 155
160Thr Thr Thr Thr Asn Ser Lys Arg Glu Glu Lys Leu Phe Asn Val Thr
165 170 175Ser Thr Leu Arg Ile Asn Thr Thr Thr Asn Glu Ile Phe Tyr
Cys Thr 180 185 190Phe Arg Arg Leu Asp Pro Glu Glu Asn His Thr Ala
Glu Leu Val Ile 195 200 205Pro Glu Leu Pro Leu Ala His Pro Pro Asn
Glu Arg Thr Ile Glu Gly 210 215 220Arg Asp Tyr Lys Asp Asp Asp Asp
Lys His His His His His His225 230 235506179PRTHomo Sapien 506Glu
Ile Asn Gly Ser Ala Asn Tyr Glu Met Phe Ile Phe His Asn Gly1 5 10
15Gly Val Gln Ile Leu Cys Lys Tyr Pro Asp Ile Val Gln Gln Phe Lys
20 25 30Met Gln Leu Leu Lys Gly Gly Gln Ile Leu Cys Asp Leu Thr Lys
Thr 35 40 45Lys Gly Ser Gly Asn Thr Val Ser Ile Lys Ser Leu Lys Phe
Cys His 50 55 60Ser Gln Leu Ser Asn Asn Ser Val Ser Phe Phe Leu Tyr
Asn Leu Asp65 70 75 80His Ser His Ala Asn Tyr Tyr Phe Cys Asn Leu
Ser Ile Phe Asp Pro 85 90 95Pro Pro Phe Lys Val Thr Leu Thr Gly Gly
Tyr Leu His Ile Tyr Glu 100 105 110Ser Gln Leu Cys Cys Gln Leu Lys
Phe Trp Leu Pro Ile Gly Cys Ala 115 120 125Ala Phe Val Val Val Cys
Ile Leu Gly Cys Ile Leu Ile Cys Trp Leu 130 135 140Thr Lys Lys Lys
Tyr Ser Ser Ser Val His Asp Pro Asn Gly Glu Tyr145 150 155 160Met
Phe Met Arg Ala Val Asn Thr Ala Lys Lys Ser Arg Leu Thr Asp 165 170
175Val Thr Leu507121PRTHomo Sapien 507Glu Ile Asn Gly Ser Ala Asn
Tyr Glu Met Phe Ile Phe His Asn Gly1 5 10 15Gly Val Gln Ile Leu Cys
Lys Tyr Pro Asp Ile Val Gln Gln Phe Lys 20 25 30Met Gln Leu Leu Lys
Gly Gly Gln Ile Leu Cys Asp Leu Thr Lys Thr 35 40 45Lys Gly Ser Gly
Asn Thr Val Ser Ile Lys Ser Leu Lys Phe Cys His 50 55 60Ser Gln Leu
Ser Asn Asn Ser Val Ser Phe Phe Leu Tyr Asn Leu Asp65 70 75 80His
Ser His Ala Asn Tyr Tyr Phe Cys Asn Leu Ser Ile Phe Asp Pro 85 90
95Pro Pro Phe Lys Val Thr Leu Thr Gly Gly Tyr Leu His Ile Tyr Glu
100 105 110Ser Gln Leu Cys Cys Gln Leu Lys Phe 115 120508199PRTHomo
Sapien 508Met Lys Ser Gly Leu Trp Tyr Phe Phe Leu Phe Cys Leu Arg
Ile Lys1 5 10 15Val Leu Thr Gly Glu Ile Asn Gly Ser Ala Asn Tyr Glu
Met Phe Ile 20 25 30Phe His Asn Gly Gly Val Gln Ile Leu Cys Lys Tyr
Pro Asp Ile Val 35 40 45Gln Gln Phe Lys Met Gln Leu Leu Lys Gly Gly
Gln Ile Leu Cys Asp 50 55 60Leu Thr Lys Thr Lys Gly Ser Gly Asn Thr
Val Ser Ile Lys Ser Leu65 70 75 80Lys Phe Cys His Ser Gln Leu Ser
Asn Asn Ser Val Ser Phe Phe Leu 85 90 95Tyr Asn Leu Asp His Ser His
Ala Asn Tyr Tyr Phe Cys Asn Leu Ser 100 105 110Ile Phe Asp Pro Pro
Pro Phe Lys Val Thr Leu Thr Gly Gly Tyr Leu 115 120 125His Ile Tyr
Glu Ser Gln Leu Cys Cys Gln Leu Lys Phe Trp Leu Pro 130 135 140Ile
Gly Cys Ala Ala Phe Val Val Val Cys Ile Leu Gly Cys Ile Leu145 150
155 160Ile Cys Trp Leu Thr Lys Lys Lys Tyr Ser Ser Ser Val His Asp
Pro 165 170 175Asn Gly Glu Tyr Met Phe Met Arg Ala Val Asn Thr Ala
Lys Lys Ser 180 185 190Arg Leu Thr Asp Val Thr Leu 19550933PRTHomo
Sapien 509Lys Tyr Ser Ser Ser Val His Asp Pro Asn Gly Glu Tyr Met
Phe Met1 5 10 15Arg Ala Val Asn Thr Ala Lys Lys Ser Arg Leu Thr Asp
Val Thr Leu 20 25 30Met510128PRTMus Musculus 510Glu Ile Asn Gly Ser
Ala Asp His Arg Met Phe Ser Phe His Asn Gly1 5 10 15Gly Val Gln Ile
Ser Cys Lys Tyr Pro Glu Thr Val Gln Gln Leu Lys 20 25 30Met Arg Leu
Phe Arg Glu Arg Glu Val Leu Cys Glu Leu Thr Lys Thr 35 40 45Lys Gly
Ser Gly Asn Ala Val Ser Ile Lys Asn Pro Met Leu Cys Leu 50 55 60Tyr
His Leu Ser Asn Asn Ser Val Ser Phe Phe Leu Asn Asn Pro Asp65 70 75
80Ser Ser Gln Gly Ser Tyr Tyr Phe Cys Ser Leu Ser Ile Phe Asp Pro
85 90 95Pro Pro Phe Gln Glu Arg Asn Leu Ser Gly Gly Tyr Leu His Ile
Tyr 100 105 110Glu Ser Gln Leu Cys Cys Gln Leu Lys Ile Val Val Gln
Val Thr Glu 115 120 125511121PRTMus Musculus 511Glu Ile Asn Gly Ser
Ala Asp His Arg Met Phe Ser Phe His Asn Gly1 5 10 15Gly Val Gln Ile
Ser Cys Lys Tyr Pro Glu Thr Val Gln Gln Leu Lys 20 25 30Met Arg Leu
Phe Arg Glu Arg Glu Val Leu Cys Glu Leu Thr Lys Thr 35 40 45Lys Gly
Ser Gly Asn Ala Val Ser Ile Lys Asn Pro Met Leu Cys Leu 50 55 60Tyr
His Leu Ser Asn Asn Ser Val Ser Phe Phe Leu Asn Asn Pro Asp65 70 75
80Ser Ser Gln Gly Ser Tyr Tyr Phe Cys Ser Leu Ser Ile Phe Asp Pro
85 90 95Pro Pro Phe Gln Glu Arg Asn Leu Ser Gly Gly Tyr Leu His Ile
Tyr 100 105 110Glu Ser Gln Leu Cys Cys Gln Leu Lys 115
120512147PRTMus Musculus 512Met Gly Trp Ser Cys Ile Ile Leu Phe Leu
Val Ala Thr Ala Thr Gly1 5 10 15Val His Ser Glu Ile Asn Gly Ser Ala
Asp His Arg Met Phe Ser Phe 20 25 30His Asn Gly Gly Val Gln Ile Ser
Cys Lys Tyr Pro Glu Thr Val Gln 35 40 45Gln Leu Lys Met Arg Leu Phe
Arg Glu Arg Glu Val Leu Cys Glu Leu 50 55 60Thr Lys Thr Lys Gly Ser
Gly Asn Ala Val Ser Ile Lys Asn Pro Met65 70 75 80Leu Cys Leu Tyr
His Leu Ser Asn Asn Ser Val Ser Phe Phe Leu Asn 85 90 95Asn Pro Asp
Ser Ser Gln Gly Ser Tyr Tyr Phe Cys Ser Leu Ser Ile 100 105 110Phe
Asp Pro Pro Pro Phe Gln Glu Arg Asn Leu Ser Gly Gly Tyr Leu 115 120
125His Ile Tyr Glu Ser Gln Leu Cys Cys Gln Leu Lys Ile Val Val Gln
130 135 140Val Thr Glu145513199PRTCynomologus 513Met Lys Ser Gly
Leu Trp Tyr Phe Phe Leu Phe Cys Leu His Met Lys1 5 10 15Val Leu Thr
Gly Glu Ile Asn Gly Ser Ala Asn Tyr Glu Met Phe Ile 20 25 30Phe His
Asn Gly Gly Val Gln Ile Leu Cys Lys Tyr Pro Asp Ile Val 35 40 45Gln
Gln Phe Lys Met Gln Leu Leu Lys Gly Gly Gln Ile Leu Cys Asp 50 55
60Leu Thr Lys Thr Lys Gly Ser Gly Asn Lys Val Ser Ile Lys Ser Leu65
70 75 80Lys Phe Cys His Ser Gln Leu Ser Asn Asn Ser Val Ser Phe Phe
Leu 85 90 95Tyr Asn Leu Asp Arg Ser His Ala Asn Tyr Tyr Phe Cys Asn
Leu Ser 100 105 110Ile Phe Asp Pro Pro Pro Phe Lys Val Thr Leu Thr
Gly Gly Tyr Leu 115 120 125His Ile Tyr Glu Ser Gln Leu Cys Cys Gln
Leu Lys Phe Trp Leu Pro 130 135 140Ile Gly Cys Ala Thr Phe Val Val
Val Cys Ile Phe Gly Cys Ile Leu145 150 155 160Ile Cys Trp Leu Thr
Lys Lys Lys Tyr Ser Ser Thr Val His Asp Pro 165 170 175Asn Gly Glu
Tyr Met Phe Met Arg Ala Val Asn Thr Ala Lys Lys Ser 180 185 190Arg
Leu Thr Gly Thr Thr Pro 195514120PRTCynomologus 514Glu Ile Asn Gly
Ser Ala Asn Tyr Glu Met Phe Ile Phe His Asn Gly1 5 10 15Gly Val Gln
Ile Leu Cys Lys Tyr Pro Asp Ile Val Gln Gln Phe Lys 20 25
30Met Gln Leu Leu Lys Gly Gly Gln Ile Leu Cys Asp Leu Thr Lys Thr
35 40 45Lys Gly Ser Gly Asn Lys Val Ser Ile Lys Ser Leu Lys Phe Cys
His 50 55 60Ser Gln Leu Ser Asn Asn Ser Val Ser Phe Phe Leu Tyr Asn
Leu Asp65 70 75 80Arg Ser His Ala Asn Tyr Tyr Phe Cys Asn Leu Ser
Ile Phe Asp Pro 85 90 95Pro Pro Phe Lys Val Thr Leu Thr Gly Gly Tyr
Leu His Ile Tyr Glu 100 105 110Ser Gln Leu Cys Cys Gln Leu Lys 115
120515240PRTHomo Sapien 515Asp Thr Gln Glu Lys Glu Val Arg Ala Met
Val Gly Ser Asp Val Glu1 5 10 15Leu Ser Cys Ala Cys Pro Glu Gly Ser
Arg Phe Asp Leu Asn Asp Val 20 25 30Tyr Val Tyr Trp Gln Thr Ser Glu
Ser Lys Thr Val Val Thr Tyr His 35 40 45Ile Pro Gln Asn Ser Ser Leu
Glu Asn Val Asp Ser Arg Tyr Arg Asn 50 55 60Arg Ala Leu Met Ser Pro
Ala Gly Met Leu Arg Gly Asp Phe Ser Leu65 70 75 80Arg Leu Phe Asn
Val Thr Pro Gln Asp Glu Gln Lys Phe His Cys Leu 85 90 95Val Leu Ser
Gln Ser Leu Gly Phe Gln Glu Val Leu Ser Val Glu Val 100 105 110Thr
Leu His Val Ala Ala Asn Phe Ser Val Pro Val Val Ser Ala Pro 115 120
125His Ser Pro Ser Gln Asp Glu Leu Thr Phe Thr Cys Thr Ser Ile Asn
130 135 140Gly Tyr Pro Arg Pro Asn Val Tyr Trp Ile Asn Lys Thr Asp
Asn Ser145 150 155 160Leu Leu Asp Gln Ala Leu Gln Asn Asp Thr Val
Phe Leu Asn Met Arg 165 170 175Gly Leu Tyr Asp Val Val Ser Val Leu
Arg Ile Ala Arg Thr Pro Ser 180 185 190Val Asn Ile Gly Cys Cys Ile
Glu Asn Val Leu Leu Gln Gln Asn Leu 195 200 205Thr Val Gly Ser Gln
Thr Gly Asn Asp Ile Gly Glu Arg Asp Lys Ile 210 215 220Thr Glu Asn
Pro Val Ser Thr Gly Glu Lys Asn Ala Ala Thr Trp Ser225 230 235
240516302PRTHomo Sapien 516Met Arg Leu Gly Ser Pro Gly Leu Leu Phe
Leu Leu Phe Ser Ser Leu1 5 10 15Arg Ala Asp Thr Gln Glu Lys Glu Val
Arg Ala Met Val Gly Ser Asp 20 25 30Val Glu Leu Ser Cys Ala Cys Pro
Glu Gly Ser Arg Phe Asp Leu Asn 35 40 45Asp Val Tyr Val Tyr Trp Gln
Thr Ser Glu Ser Lys Thr Val Val Thr 50 55 60Tyr His Ile Pro Gln Asn
Ser Ser Leu Glu Asn Val Asp Ser Arg Tyr65 70 75 80Arg Asn Arg Ala
Leu Met Ser Pro Ala Gly Met Leu Arg Gly Asp Phe 85 90 95Ser Leu Arg
Leu Phe Asn Val Thr Pro Gln Asp Glu Gln Lys Phe His 100 105 110Cys
Leu Val Leu Ser Gln Ser Leu Gly Phe Gln Glu Val Leu Ser Val 115 120
125Glu Val Thr Leu His Val Ala Ala Asn Phe Ser Val Pro Val Val Ser
130 135 140Ala Pro His Ser Pro Ser Gln Asp Glu Leu Thr Phe Thr Cys
Thr Ser145 150 155 160Ile Asn Gly Tyr Pro Arg Pro Asn Val Tyr Trp
Ile Asn Lys Thr Asp 165 170 175Asn Ser Leu Leu Asp Gln Ala Leu Gln
Asn Asp Thr Val Phe Leu Asn 180 185 190Met Arg Gly Leu Tyr Asp Val
Val Ser Val Leu Arg Ile Ala Arg Thr 195 200 205Pro Ser Val Asn Ile
Gly Cys Cys Ile Glu Asn Val Leu Leu Gln Gln 210 215 220Asn Leu Thr
Val Gly Ser Gln Thr Gly Asn Asp Ile Gly Glu Arg Asp225 230 235
240Lys Ile Thr Glu Asn Pro Val Ser Thr Gly Glu Lys Asn Ala Ala Thr
245 250 255Trp Ser Ile Leu Ala Val Leu Cys Leu Leu Val Val Val Ala
Val Ala 260 265 270Ile Gly Trp Val Cys Arg Asp Arg Cys Leu Gln His
Ser Tyr Ala Gly 275 280 285Ala Trp Ala Val Ser Pro Glu Thr Glu Leu
Thr Gly His Val 290 295 300517113PRTHomo Sapien 517Leu Gln Met Ile
Leu Asn Gly Ile Asn Asn Tyr Lys Asn Pro Lys Leu1 5 10 15Thr Ala Met
Leu Thr Phe Lys Phe Tyr Met Pro Lys Lys Ala Thr Glu 20 25 30Leu Lys
His Leu Gln Cys Leu Glu Glu Glu Leu Lys Pro Leu Glu Glu 35 40 45Val
Leu Asn Leu Ala Gln Ser Lys Asn Phe His Leu Arg Pro Arg Asp 50 55
60Leu Ile Ser Asn Ile Asn Val Ile Val Leu Glu Leu Lys Gly Ser Glu65
70 75 80Thr Thr Phe Met Cys Glu Tyr Ala Asp Glu Thr Ala Thr Ile Val
Glu 85 90 95Phe Leu Asn Arg Trp Ile Thr Phe Cys Gln Ser Ile Ile Ser
Thr Leu 100 105 110Thr518113PRTHomo Sapien 518Leu Gln Met Ile Leu
Asn Gly Ile Asn Asn Tyr Lys Asn Pro Lys Leu1 5 10 15Thr Gln Met Leu
Thr Phe Lys Phe Tyr Met Pro Lys Lys Ala Thr Glu 20 25 30Leu Lys His
Leu Gln Cys Leu Glu Glu Glu Leu Lys Pro Leu Glu Glu 35 40 45Val Leu
Asn Leu Ala Gln Ser Lys Asn Phe His Leu Arg Pro Arg Asp 50 55 60Leu
Ile Ser Asn Ile Asn Val Ile Val Leu Glu Leu Lys Gly Ser Glu65 70 75
80Thr Thr Phe Met Cys Glu Tyr Ala Asp Glu Thr Ala Thr Ile Val Glu
85 90 95Phe Leu Asn Arg Trp Ile Thr Phe Cys Gln Ser Ile Ile Ser Thr
Leu 100 105 110Thr519336DNAHomo Sapien 519gatattgtga tgactcagtc
tccactctcc ctgcccgtca cccctggaga gccggcctcc 60atctcctgca ggtctagtca
gagcctcctg catagtgatg gatacaacta tttggattgg 120tacctgcaga
agccagggca gtctccacag ctcctgatct atttgggttc tactcgggcc
180tccgggttcc ctgacaggtt cagtggcagt ggatcaggca cagattttac
actgaaaatc 240agcagagtgg aggctgagga tgttggggtt tattactgca
tgcaagctct acaaactccg 300ctcagttttg gccaggggac caagctggag atcaaa
336520657DNAHomo Sapien 520gatattgtga tgactcagtc tccactctcc
ctgcccgtca cccctggaga gccggcctcc 60atctcctgca ggtctagtca gagcctcctg
catagtgatg gatacaacta tttggattgg 120tacctgcaga agccagggca
gtctccacag ctcctgatct atttgggttc tactcgggcc 180tccgggttcc
ctgacaggtt cagtggcagt ggatcaggca cagattttac actgaaaatc
240agcagagtgg aggctgagga tgttggggtt tattactgca tgcaagctct
acaaactccg 300ctcagttttg gccaggggac caagctggag atcaaacgta
cggtggccgc tccctccgtg 360ttcatcttcc caccttccga cgagcagctg
aagtccggca ccgcttctgt cgtgtgcctg 420ctgaacaact tctacccccg
cgaggccaag gtgcagtgga aggtggacaa cgccctgcag 480tccggcaact
cccaggaatc cgtgaccgag caggactcca aggacagcac ctactccctg
540tcctccaccc tgaccctgtc caaggccgac tacgagaagc acaaggtgta
cgcctgcgaa 600gtgacccacc agggcctgtc tagccccgtg accaagtctt
tcaaccgggg cgagtgt 657521372DNAHomo Sapien 521gaggtgcagc tggtggagtc
tgggggaggt gtggtacggc ctggggggtc cctgagactc 60tcctgtgtag cctctggagt
cacctttgat gattatggca tgagctgggt ccgccaagct 120ccagggaagg
ggctggagtg ggtctctggt attaattgga atggtggcga cacagattat
180tcagactctg tgaagggccg attcaccatc tccagagaca acgccaagaa
ctccctgtat 240ctacaaatga atagtctgag agccgaggac acggccttgt
attactgtgc gagggatttc 300tatggttcgg ggagttatta tcacgttcct
tttgactact ggggccaggg aatcctggtc 360accgtctcct ca 3725221368DNAHomo
Sapien 522gaggtgcagc tggtggagtc tgggggaggt gtggtacggc ctggggggtc
cctgagactc 60tcctgtgtag cctctggagt cacctttgat gattatggca tgagctgggt
ccgccaagct 120ccagggaagg ggctggagtg ggtctctggt attaattgga
atggtggcga cacagattat 180tcagactctg tgaagggccg attcaccatc
tccagagaca acgccaagaa ctccctgtat 240ctacaaatga atagtctgag
agccgaggac acggccttgt attactgtgc gagggatttc 300tatggttcgg
ggagttatta tcacgttcct tttgactact ggggccaggg aatcctggtc
360accgtctcct cagccagcac caagggcccc tctgtgttcc ctctggcccc
ttccagcaag 420tccacctctg gcggaacagc cgctctgggc tgcctcgtga
aggactactt ccccgagcct 480gtgaccgtgt cctggaactc tggcgctctg
accagcggag tgcacacctt ccctgctgtg 540ctgcagtcct ccggcctgta
ctccctgtcc tccgtcgtga ccgtgccttc cagctctctg 600ggcacccaga
cctacatctg caacgtgaac cacaagccct ccaacaccaa ggtggacaag
660aaggtggaac ccaagtcctg cgacaagacc cacacctgtc ccccttgtcc
tgcccctgaa 720ctgctgggcg gaccttccgt gttcctgttc cccccaaagc
ccaaggacac cctgatgatc 780tcccggaccc ccgaagtgac ctgcgtggtg
gtggatgtgt cccacgagga ccctgaagtg 840aagttcaatt ggtacgtgga
cggcgtggaa gtgcacaacg ccaagaccaa gcctagagag 900gaacagtaca
actccaccta ccgggtggtg tccgtgctga ccgtgctgca ccaggattgg
960ctgaacggca aagagtacaa gtgcaaggtg tccaacaagg ccctgcctgc
ccccatcgaa 1020aagaccatct ccaaggccaa gggccagccc cgggaacccc
aggtgtacac actgccccct 1080agcagggacg agctgaccaa gaaccaggtg
tccctgacct gtctcgtgaa aggcttctac 1140ccctccgata tcgccgtgga
atgggagtcc aacggccagc ctgagaacaa ctacaagacc 1200accccccctg
tgctggactc cgacggctca ttcttcctgt acagcaagct gacagtggac
1260aagtcccggt ggcagcaggg caacgtgttc tcctgctccg tgatgcacga
ggccctgcac 1320aaccactaca cccagaagtc cctgtccctg agccccggca agtgatga
1368523990DNAHomo Sapien 523gcctccacca agggcccatc ggtcttcccc
ctggcaccct cctccaagag cacctctggg 60ggcacagcgg ccctgggctg cctggtcaag
gactacttcc ccgaaccggt gacggtgtcg 120tggaactcag gcgccctgac
cagcggcgtg cacaccttcc cggctgtcct acagtcctca 180ggactctact
ccctcagcag cgtggtgacc gtgccctcca gcagcttggg cacccagacc
240tacatctgca acgtgaatca caagcccagc aacaccaagg tggacaagag
agttgagccc 300aaatcttgtg acaaaactca cacatgccca ccgtgcccag
cacctgaact cctgggggga 360ccgtcagtct tcctcttccc cccaaaaccc
aaggacaccc tcatgatctc ccggacccct 420gaggtcacat gcgtggtggt
ggacgtgagc cacgaagacc ctgaggtcaa gttcaactgg 480tacgtggacg
gcgtggaggt gcataatgcc aagacaaagc cgcgggagga gcagtacaac
540agcacgtacc gtgtggtcag cgtcctcacc gtcctgcacc aggactggct
gaatggcaag 600gagtacaagt gcaaggtctc caacaaagcc ctcccagccc
ccatcgagaa aaccatctcc 660aaagccaaag ggcagccccg agaaccacag
gtgtacaccc tgcccccatc ccgggaggag 720atgaccaaga accaggtcag
cctgacctgc ctggtcaaag gcttctatcc cagcgacatc 780gccgtggagt
gggagagcaa tgggcagccg gagaacaact acaagaccac gcctcccgtg
840ctggactccg acggctcctt cttcctctat agcaagctca ccgtggacaa
gagcaggtgg 900cagcagggga acgtcttctc atgctccgtg atgcatgagg
ctctgcacaa ccactacacg 960cagaagagcc tctccctgtc cccgggtaaa
990524330PRTHomo Sapien 524Ala Ser Thr Lys Gly Pro Ser Val Phe Pro
Leu Ala Pro Ser Ser Lys1 5 10 15Ser Thr Ser Gly Gly Thr Ala Ala Leu
Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser
Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala
Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr
Val Pro Ser Ser Ser Leu Gly Thr Gln Thr65 70 75 80Tyr Ile Cys Asn
Val Asn His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Arg Val Glu
Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys 100 105 110Pro
Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro 115 120
125Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys
130 135 140Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe
Asn Trp145 150 155 160Tyr Val Asp Gly Val Glu Val His Asn Ala Lys
Thr Lys Pro Arg Glu 165 170 175Glu Gln Tyr Asn Ser Thr Tyr Arg Val
Val Ser Val Leu Thr Val Leu 180 185 190His Gln Asp Trp Leu Asn Gly
Lys Glu Tyr Lys Cys Lys Val Ser Asn 195 200 205Lys Ala Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly 210 215 220Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu225 230 235
240Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr
245 250 255Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn 260 265 270Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp
Gly Ser Phe Phe 275 280 285Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp Gln Gln Gly Asn 290 295 300Val Phe Ser Cys Ser Val Met His
Glu Ala Leu His Asn His Tyr Thr305 310 315 320Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 325 330525990DNAHomo Sapien 525gcctccacca
agggcccatc ggtcttcccc ctggcaccct cctccaagag cacctctggg 60ggcacagcgg
ccctgggctg cctggtcaag gactacttcc ccgaaccggt gacggtgtcg
120tggaactcag gcgccctgac cagcggcgtg cacaccttcc cggctgtcct
acagtcctca 180ggactctact ccctcagcag cgtggtgacc gtgccctcca
gcagcttggg cacccagacc 240tacatctgca acgtgaatca caagcccagc
aacaccaagg tggacaagaa agttgagccc 300aaatcttgtg acaaaactca
cacatgccca ccgtgcccag cacctgaact cctgggggga 360ccgtcagtct
tcctcttccc cccaaaaccc aaggacaccc tcatgatctc ccggacccct
420gaggtcacat gcgtggtggt ggacgtgagc cacgaagacc ctgaggtcaa
gttcaactgg 480tacgtggacg gcgtggaggt gcataatgcc aagacaaagc
cgcgggagga gcagtacaac 540agcacgtacc gtgtggtcag cgtcctcacc
gtcctgcacc aggactggct gaatggcaag 600gagtacaagt gcaaggtctc
caacaaagcc ctcccagccc ccatcgagaa aaccatctcc 660aaagccaaag
ggcagccccg agaaccacag gtgtacaccc tgcccccatc ccgggatgag
720ctgaccaaga accaggtcag cctgacctgc ctggtcaaag gcttctatcc
cagcgacatc 780gccgtggagt gggagagcaa tgggcagccg gagaacaact
acaagaccac gcctcccgtg 840ctggactccg acggctcctt cttcctctac
agcaagctca ccgtggacaa gagcaggtgg 900cagcagggga acatcttctc
atgctccgtg atgcatgagg ctctgcacaa ccactacacg 960cagaagagcc
tctccctgtc tccgggtaaa 990526330PRTHomo Sapien 526Ala Ser Thr Lys
Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys1 5 10 15Ser Thr Ser
Gly Gly Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro
Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly
Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55
60Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Gln Thr65
70 75 80Tyr Ile Cys Asn Val Asn His Lys Pro Ser Asn Thr Lys Val Asp
Lys 85 90 95Lys Val Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro
Pro Cys 100 105 110Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe
Leu Phe Pro Pro 115 120 125Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
Thr Pro Glu Val Thr Cys 130 135 140Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn Trp145 150 155 160Tyr Val Asp Gly Val
Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu 165 170 175Glu Gln Tyr
Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu 180 185 190His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn 195 200
205Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly
210 215 220Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg
Asp Glu225 230 235 240Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe Tyr 245 250 255Pro Ser Asp Ile Ala Val Glu Trp Glu
Ser Asn Gly Gln Pro Glu Asn 260 265 270Asn Tyr Lys Thr Thr Pro Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe 275 280 285Leu Tyr Ser Lys Leu
Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn 290 295 300Ile Phe Ser
Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr305 310 315
320Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 325 330527978DNAHomo
Sapien 527gcctccacca agggcccatc ggtcttcccc ctggcgccct gctccaggag
cacctccgag 60agcacagccg ccctgggctg cctggtcaag gactacttcc ccgaaccggt
gacggtgtcg 120tggaactcag gcgctctgac cagcggcgtg cacaccttcc
cagctgtcct acagtcctca 180ggactctact ccctcagcag cgtggtgacc
gtgccctcca gcaacttcgg cacccagacc 240tacacctgca acgtagatca
caagcccagc aacaccaagg tggacaagac agttgagcgc 300aaatgttgtg
tcgagtgccc accgtgccca gcaccacctg tggcaggacc gtcagtcttc
360ctcttccccc caaaacccaa ggacaccctc atgatctccc ggacccctga
ggtcacgtgc 420gtggtggtgg acgtgagcca cgaagacccc gaggtccagt
tcaactggta cgtggacggc 480gtggaggtgc ataatgccaa gacaaagcca
cgggaggagc agttcaacag cacgttccgt 540gtggtcagcg tcctcaccgt
tgtgcaccag gactggctga acggcaagga gtacaagtgc 600aaggtctcca
acaaaggcct cccagccccc atcgagaaaa ccatctccaa aaccaaaggg
660cagccccgag aaccacaggt gtacaccctg cccccatccc gggaggagat
gaccaagaac 720caggtcagcc tgacctgcct ggtcaaaggc
ttctacccca gcgacatcgc cgtggagtgg 780gagagcaatg ggcagccgga
gaacaactac aagaccacac ctcccatgct ggactccgac 840ggctccttct
tcctctacag caagctcacc gtggacaaga gcaggtggca gcaggggaac
900gtcttctcat gctccgtgat gcatgaggct ctgcacaacc actacacgca
gaagagcctc 960tccctgtctc cgggtaaa 978528326PRTHomo Sapien 528Ala
Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg1 5 10
15Ser Thr Ser Glu Ser Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr
20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr
Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu
Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser Asn Phe Gly
Thr Gln Thr65 70 75 80Tyr Thr Cys Asn Val Asp His Lys Pro Ser Asn
Thr Lys Val Asp Lys 85 90 95Thr Val Glu Arg Lys Cys Cys Val Glu Cys
Pro Pro Cys Pro Ala Pro 100 105 110Pro Val Ala Gly Pro Ser Val Phe
Leu Phe Pro Pro Lys Pro Lys Asp 115 120 125Thr Leu Met Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val Asp 130 135 140Val Ser His Glu
Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly145 150 155 160Val
Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn 165 170
175Ser Thr Phe Arg Val Val Ser Val Leu Thr Val Val His Gln Asp Trp
180 185 190Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly
Leu Pro 195 200 205Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly
Gln Pro Arg Glu 210 215 220Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg
Glu Glu Met Thr Lys Asn225 230 235 240Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe Tyr Pro Ser Asp Ile 245 250 255Ala Val Glu Trp Glu
Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr 260 265 270Thr Pro Pro
Met Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys 275 280 285Leu
Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys 290 295
300Ser Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser
Leu305 310 315 320Ser Leu Ser Pro Gly Lys 325529978DNAHomo Sapien
529gcctccacca agggcccatc ggtcttcccc ctggcgccct gctccaggag
cacctccgag 60agcacagcgg ccctgggctg cctggtcaag gactacttcc ccgaaccggt
gacggtgtcg 120tggaactcag gcgctctgac cagcggcgtg cacaccttcc
cggctgtcct acagtcctca 180ggactctact ccctcagcag cgtggtgacc
gtgacctcca gcaacttcgg cacccagacc 240tacacctgca acgtagatca
caagcccagc aacaccaagg tggacaagac agttgagcgc 300aaatgttgtg
tcgagtgccc accgtgccca gcaccacctg tggcaggacc gtcagtcttc
360ctcttccccc caaaacccaa ggacaccctc atgatctccc ggacccctga
ggtcacgtgc 420gtggtggtgg acgtgagcca cgaagacccc gaggtccagt
tcaactggta cgtggacggc 480atggaggtgc ataatgccaa gacaaagcca
cgggaggagc agttcaacag cacgttccgt 540gtggtcagcg tcctcaccgt
cgtgcaccag gactggctga acggcaagga gtacaagtgc 600aaggtctcca
acaaaggcct cccagccccc atcgagaaaa ccatctccaa aaccaaaggg
660cagccccgag aaccacaggt gtacaccctg cccccatccc gggaggagat
gaccaagaac 720caggtcagcc tgacctgcct ggtcaaaggc ttctacccca
gcgacatcgc cgtggagtgg 780gagagcaatg ggcagccgga gaacaactac
aagaccacac ctcccatgct ggactccgac 840ggctccttct tcctctacag
caagctcacc gtggacaaga gcaggtggca gcaggggaac 900gtcttctcat
gctccgtgat gcatgaggct ctgcacaacc actacacaca gaagagcctc
960tccctgtctc cgggtaaa 978530326PRTHomo Sapien 530Ala Ser Thr Lys
Gly Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser
Glu Ser Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro
Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly
Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55
60Leu Ser Ser Val Val Thr Val Thr Ser Ser Asn Phe Gly Thr Gln Thr65
70 75 80Tyr Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp
Lys 85 90 95Thr Val Glu Arg Lys Cys Cys Val Glu Cys Pro Pro Cys Pro
Ala Pro 100 105 110Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp 115 120 125Thr Leu Met Ile Ser Arg Thr Pro Glu Val
Thr Cys Val Val Val Asp 130 135 140Val Ser His Glu Asp Pro Glu Val
Gln Phe Asn Trp Tyr Val Asp Gly145 150 155 160Met Glu Val His Asn
Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn 165 170 175Ser Thr Phe
Arg Val Val Ser Val Leu Thr Val Val His Gln Asp Trp 180 185 190Leu
Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro 195 200
205Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu
210 215 220Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr
Lys Asn225 230 235 240Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile 245 250 255Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr 260 265 270Thr Pro Pro Met Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Lys 275 280 285Leu Thr Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys 290 295 300Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu305 310 315
320Ser Leu Ser Pro Gly Lys 325531978DNAHomo Sapien 531gcctccacca
agggcccatc ggtcttcccc ctggcgccct gctccaggag cacctccgag 60agcacagcgg
ccctgggctg cctggtcaag gactacttcc ccgaaccggt gacggtgtcg
120tggaactcag gcgctctgac cagcggcgtg cacaccttcc cagctgtcct
acagtcctca 180ggactctact ccctcagcag cgtggtgacc gtgccctcca
gcagcttggg cacccagacc 240tacacctgca acgtagatca caagcccagc
aacaccaagg tggacaagac agttgagcgc 300aaatgttgtg tcgagtgccc
accgtgccca gcaccacctg tggcaggacc gtcagtcttc 360ctcttccccc
caaaacccaa ggacaccctc atgatctccc ggacccctga ggtcacgtgc
420gtggtggtgg acgtgagcca cgaagacccc gaggtccagt tcaactggta
cgtggacggc 480gtggaggtgc ataatgccaa gacaaagcca cgggaggagc
agttcaacag cacgttccgt 540gtggtcagcg tcctcaccgt tgtgcaccag
gactggctga acggcaagga gtacaagtgc 600aaggtctcca acaaaggcct
cccagccccc atcgagaaaa ccatctccaa aaccaaaggg 660cagccccgag
aaccacaggt gtacaccctg cccccatccc gggaggagat gaccaagaac
720caggtcagcc tgacctgcct ggtcaaaggc ttctacccca gcgacatcgc
cgtggagtgg 780gagagcaatg ggcagccgga gaacaactac aagaccacac
ctcccatgct ggactccgac 840ggctccttct tcctctacag caagctcacc
gtggacaaga gcaggtggca gcaggggaac 900gtcttctcat gctccgtgat
gcatgaggct ctgcacaacc actacacgca gaagagcctc 960tccctgtctc cgggtaaa
978532326PRTHomo Sapien 532Ala Ser Thr Lys Gly Pro Ser Val Phe Pro
Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu
Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser
Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala
Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr
Val Pro Ser Ser Ser Leu Gly Thr Gln Thr65 70 75 80Tyr Thr Cys Asn
Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Thr Val Glu
Arg Lys Cys Cys Val Glu Cys Pro Pro Cys Pro Ala Pro 100 105 110Pro
Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 115 120
125Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp
130 135 140Val Ser His Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val
Asp Gly145 150 155 160Val Glu Val His Asn Ala Lys Thr Lys Pro Arg
Glu Glu Gln Phe Asn 165 170 175Ser Thr Phe Arg Val Val Ser Val Leu
Thr Val Val His Gln Asp Trp 180 185 190Leu Asn Gly Lys Glu Tyr Lys
Cys Lys Val Ser Asn Lys Gly Leu Pro 195 200 205Ala Pro Ile Glu Lys
Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu 210 215 220Pro Gln Val
Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn225 230 235
240Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
245 250 255Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr 260 265 270Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe
Leu Tyr Ser Lys 275 280 285Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly Asn Val Phe Ser Cys 290 295 300Ser Val Met His Glu Ala Leu His
Asn His Tyr Thr Gln Lys Ser Leu305 310 315 320Ser Leu Ser Pro Gly
Lys 325533978DNAHomo Sapien 533gcctccacca agggcccatc ggtcttcccc
ctggcgccct gctccaggag cacctccgag 60agcacagcgg ccctgggctg cctggtcaag
gactacttcc ccgaaccggt gacggtgtcg 120tggaactcag gcgctctgac
cagcggcgtg cacaccttcc cggctgtcct acagtcctca 180ggactctact
ccctcagcag cgtggtgacc gtgccctcca gcaacttcgg cacccagacc
240tacacctgca acgtagatca caagcccagc aacaccaagg tggacaagac
agttgagcgc 300aaatgttgtg tcgagtgccc accgtgccca gcaccacctg
tggcaggacc gtcagtcttc 360ctcttccccc caaaacccaa ggacaccctc
atgatctccc ggacccctga ggtcacgtgc 420gtggtggtgg acgtgagcca
cgaagacccc gaggtccagt tcaactggta cgtggacggc 480gtggaggtgc
ataatgccaa gacaaagcca cgggaggagc agttcaacag cacgttccgt
540gtggtcagcg tcctcaccgt cgtgcaccag gactggctga acggcaagga
gtacaagtgc 600aaggtctcca acaaaggcct cccagccccc atcgagaaaa
ccatctccaa aaccaaaggg 660cagccccgag aaccacaggt gtacaccctg
cccccatccc gggaggagat gaccaagaac 720caggtcagcc tgacctgcct
ggtcaaaggc ttctacccca gcgacatctc cgtggagtgg 780gagagcaatg
ggcagccgga gaacaactac aagaccacac ctcccatgct ggactccgac
840ggctccttct tcctctacag caagctcacc gtggacaaga gcaggtggca
gcaggggaac 900gtcttctcat gctccgtgat gcatgaggct ctgcacaacc
actacacaca gaagagcctc 960tccctgtctc cgggtaaa 978534326PRTHomo
Sapien 534Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Cys
Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu Gly Cys Leu Val
Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly
Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser
Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser
Asn Phe Gly Thr Gln Thr65 70 75 80Tyr Thr Cys Asn Val Asp His Lys
Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Thr Val Glu Arg Lys Cys Cys
Val Glu Cys Pro Pro Cys Pro Ala Pro 100 105 110Pro Val Ala Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 115 120 125Thr Leu Met
Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 130 135 140Val
Ser His Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly145 150
155 160Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe
Asn 165 170 175Ser Thr Phe Arg Val Val Ser Val Leu Thr Val Val His
Gln Asp Trp 180 185 190Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Gly Leu Pro 195 200 205Ala Pro Ile Glu Lys Thr Ile Ser Lys
Thr Lys Gly Gln Pro Arg Glu 210 215 220Pro Gln Val Tyr Thr Leu Pro
Pro Ser Arg Glu Glu Met Thr Lys Asn225 230 235 240Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 245 250 255Ser Val
Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr 260 265
270Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
275 280 285Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe
Ser Cys 290 295 300Ser Val Met His Glu Ala Leu His Asn His Tyr Thr
Gln Lys Ser Leu305 310 315 320Ser Leu Ser Pro Gly Lys
325535318DNAHomo Sapien 535ggtcagccca aggctgcccc ctcggtcact
ctgttcccac cctcctctga ggagcttcaa 60gccaacaagg ccacactggt gtgtctcgta
agtgacttca acccgggagc cgtgacagtg 120gcctggaagg cagatggcag
ccccgtcaag gtgggagtgg agaccaccaa accctccaaa 180caaagcaaca
acaagtatgc ggccagcagc tacctgagcc tgacgcccga gcagtggaag
240tcccacagaa gctacagctg ccgggtcacg catgaaggga gcaccgtgga
gaagacagtg 300gcccctgcag aatgctct 318536106PRTHomo Sapien 536Gly
Gln Pro Lys Ala Ala Pro Ser Val Thr Leu Phe Pro Pro Ser Ser1 5 10
15Glu Glu Leu Gln Ala Asn Lys Ala Thr Leu Val Cys Leu Val Ser Asp
20 25 30Phe Asn Pro Gly Ala Val Thr Val Ala Trp Lys Ala Asp Gly Ser
Pro 35 40 45Val Lys Val Gly Val Glu Thr Thr Lys Pro Ser Lys Gln Ser
Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu
Gln Trp Lys65 70 75 80Ser His Arg Ser Tyr Ser Cys Arg Val Thr His
Glu Gly Ser Thr Val 85 90 95Glu Lys Thr Val Ala Pro Ala Glu Cys Ser
100 105537990DNAHomo Sapien 537gcctccacca agggcccatc ggtcttcccc
ctggcaccct cctccaagag cacctctggg 60ggcacagcgg ccctgggctg cctggtcaag
gactacttcc ccgaaccggt gacggtgtcg 120tggaactcag gcgccctgac
cagcggcgtg cacaccttcc cggctgtcct acagtcctca 180ggactctact
ccctcagcag cgtggtgacc gtgccctcca gcagcttggg cacccagacc
240tacatctgca acgtgaatca caagcccagc aacaccaagg tggacaagaa
agttgagccc 300aaatcttgtg acaaaactca cacatgccca ccgtgcccag
cacctgaact cctgggggga 360ccgtcagtct tcctcttccc cccaaaaccc
aaggacaccc tcatgatctc ccggacccct 420gaggtcacat gcgtggtggt
ggacgtgagc cacgaagacc ctgaggtcaa gttcaactgg 480tacgtggacg
gcgtggaggt gcataatgcc aagacaaagc cgcgggagga gcagtacaac
540agcacgtacc gggtggtcag cgtcctcacc gtcctgcacc aggactggct
gaatggcaag 600gagtacaagt gcaaggtctc caacaaagcc ctcccagccc
ccatcgagaa aaccatctcc 660aaagccaaag ggcagccccg agaaccacag
gtgtacaccc tgcccccatc ccgggatgag 720ctgaccaaga accaggtcag
cctgacctgc ctggtcaaag gcttctatcc cagcgacatc 780gccgtggagt
gggagagcaa tgggcagccg gagaacaact acaagaccac gcctcccgtg
840ctggactccg acggctcctt cttcctctac agcaagctca ccgtggacaa
gagcaggtgg 900cagcagggga acgtcttctc atgctccgtg atgcatgagg
ctctgcacaa ccactacacg 960cagaagagcc tctccctgtc tccgggtaaa
990538106PRTHomo Sapien 538Gly Gln Pro Lys Ala Ala Pro Ser Val Thr
Leu Phe Pro Pro Ser Ser1 5 10 15Glu Glu Leu Gln Ala Asn Lys Ala Thr
Leu Val Cys Leu Ile Ser Asp 20 25 30Phe Tyr Pro Gly Ala Val Thr Val
Ala Trp Lys Ala Asp Ser Ser Pro 35 40 45Val Lys Ala Gly Val Glu Thr
Thr Thr Pro Ser Lys Gln Ser Asn Asn 50 55 60Lys Tyr Ala Ala Ser Ser
Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys65 70 75 80Ser His Arg Ser
Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr Val 85 90 95Glu Lys Thr
Val Ala Pro Thr Glu Cys Ser 100 105539157PRTHomo Sapien 539Met Gly
Trp Ser Cys Ile Ile Leu Phe Leu Val Ala Thr Ala Thr Gly1 5 10 15Val
His Ser Met Met Thr Gly Thr Ile Glu Thr Thr Gly Asn Ile Ser 20 25
30Ala Glu Lys Gly Gly Ser Ile Ile Leu Gln Cys His Leu Ser Ser Thr
35 40 45Thr Ala Gln Val Thr Gln Val Asn Trp Glu Gln Gln Asp Gln Leu
Leu 50 55 60Ala Ile Cys Asn Ala Asp Leu Gly Trp His Ile Ser Pro Ser
Phe Lys65 70 75 80Asp Arg Val Ala Pro Gly Pro Gly Leu Gly Leu Thr
Leu Gln Ser Leu 85 90 95Thr Val Asn Asp Thr Gly Glu Tyr Phe Cys Ile
Tyr His Thr Tyr Pro 100 105 110Asp Gly Thr Tyr Thr Gly Arg Ile Phe
Leu Glu Val Leu Glu Ser Ser 115 120 125Val Ala Glu His Gly Ala Arg
Phe Gln Ile Pro Ile Glu Gly Arg Asp 130 135 140Tyr Lys Asp Asp Asp
Asp Lys His His His His His His145
150 155
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
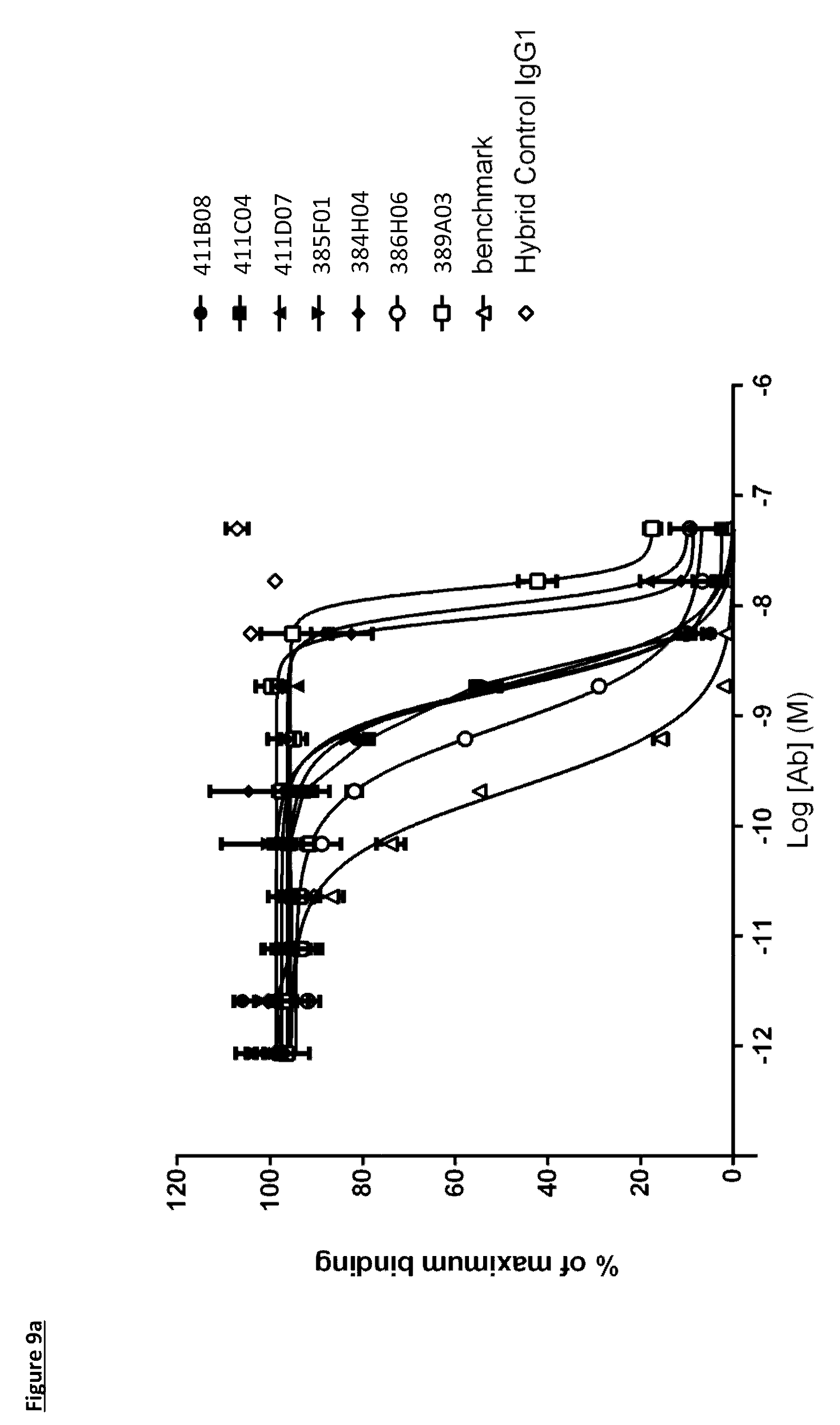
D00014
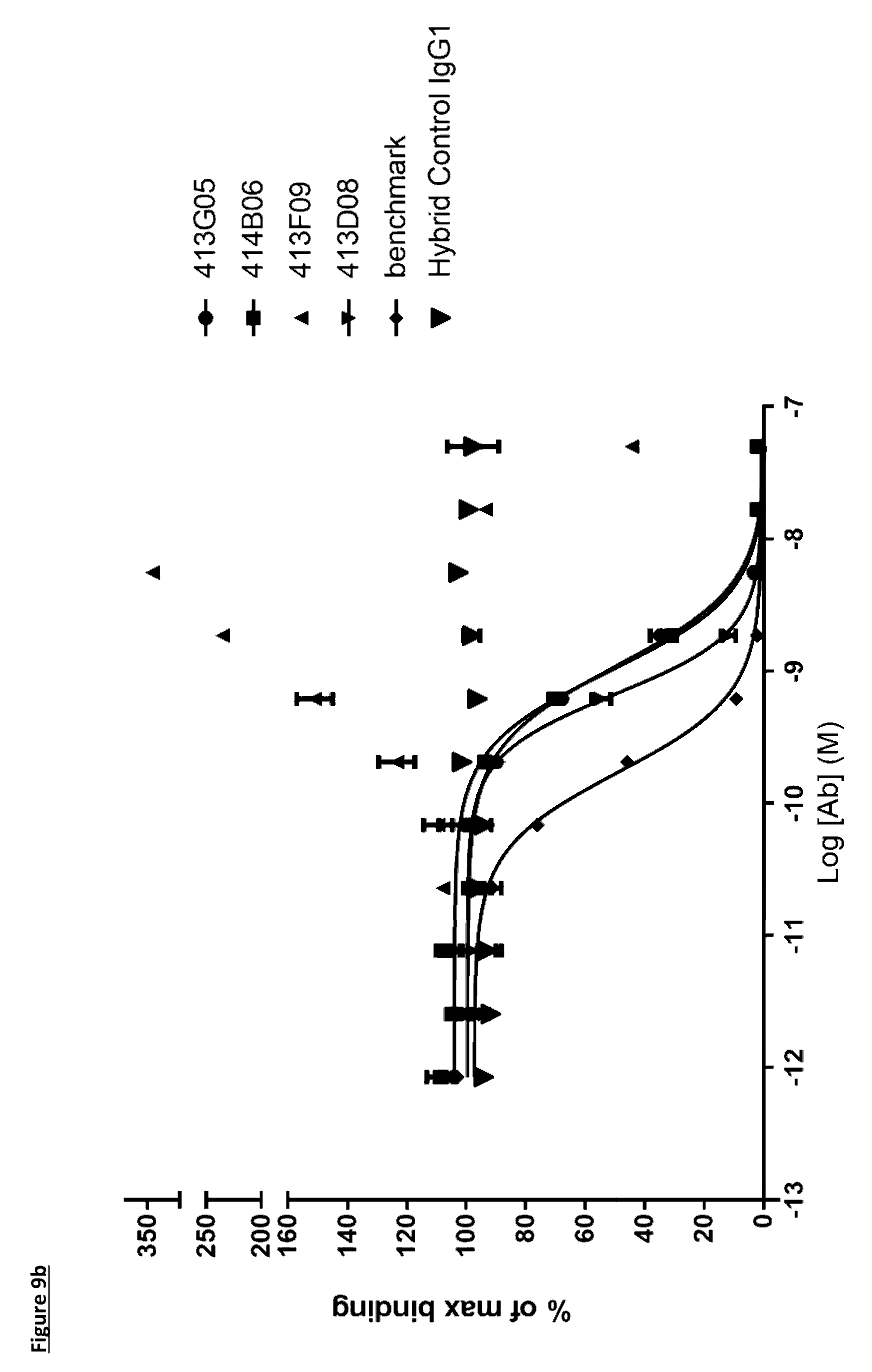
D00015
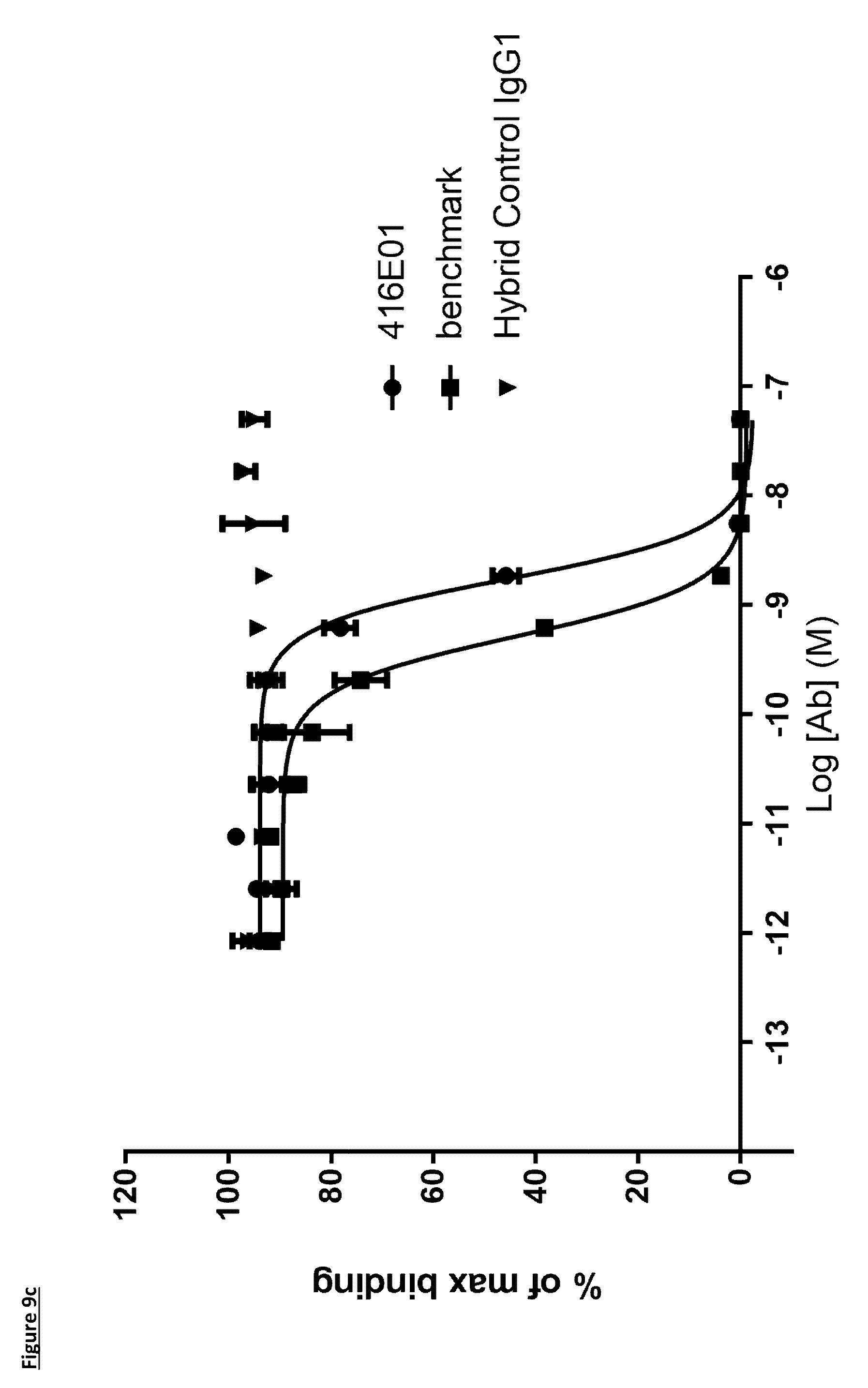
D00016
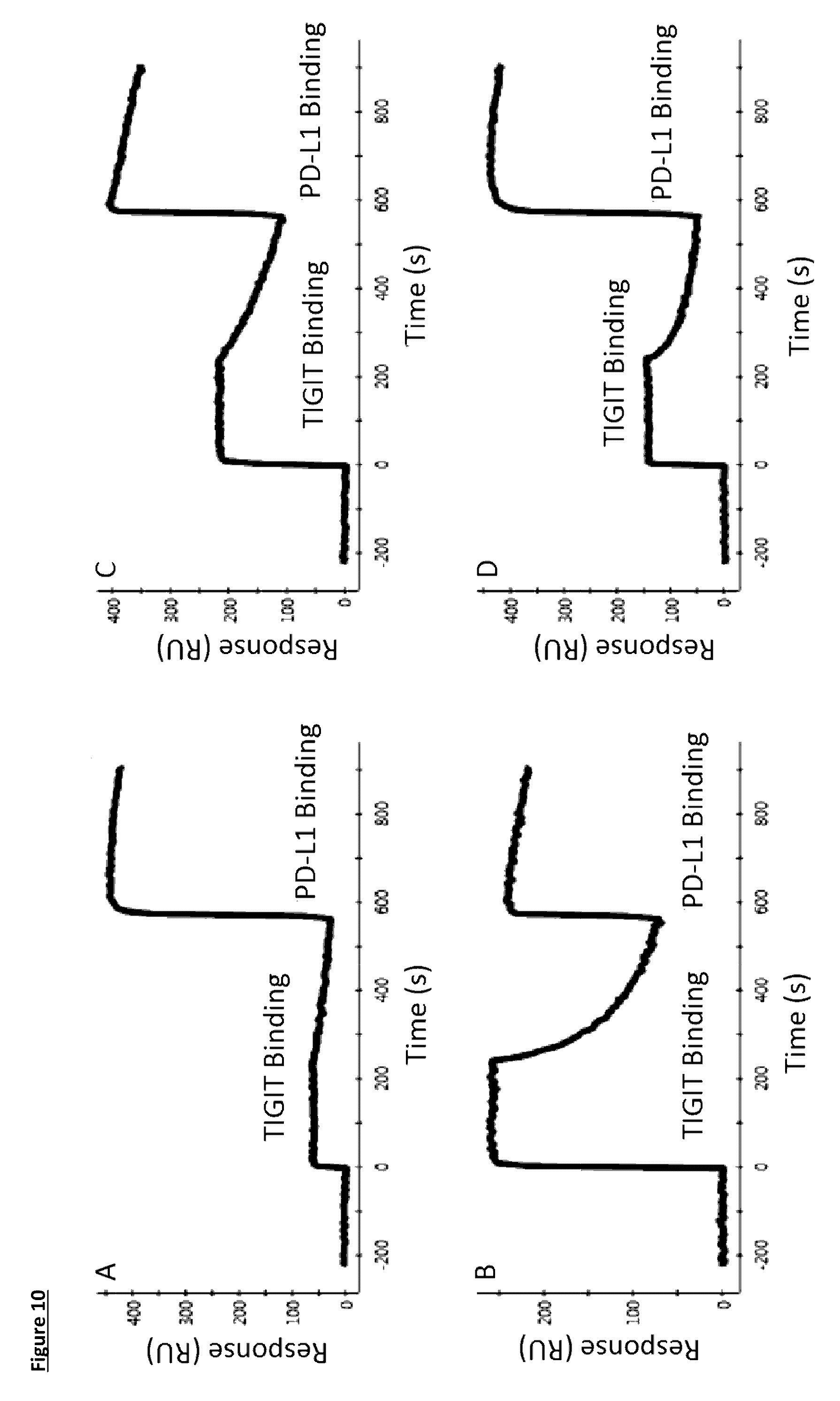
D00017
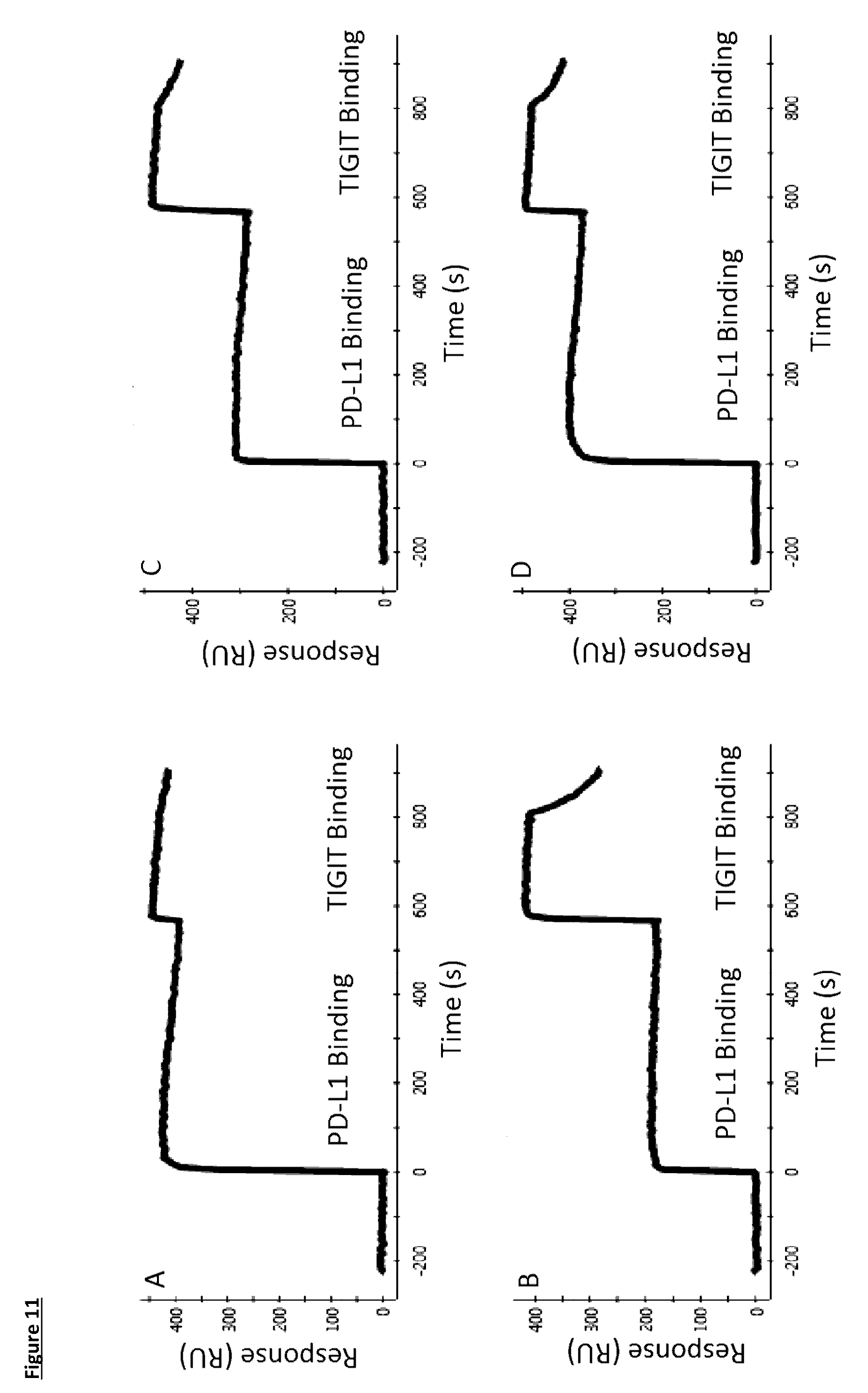
D00018
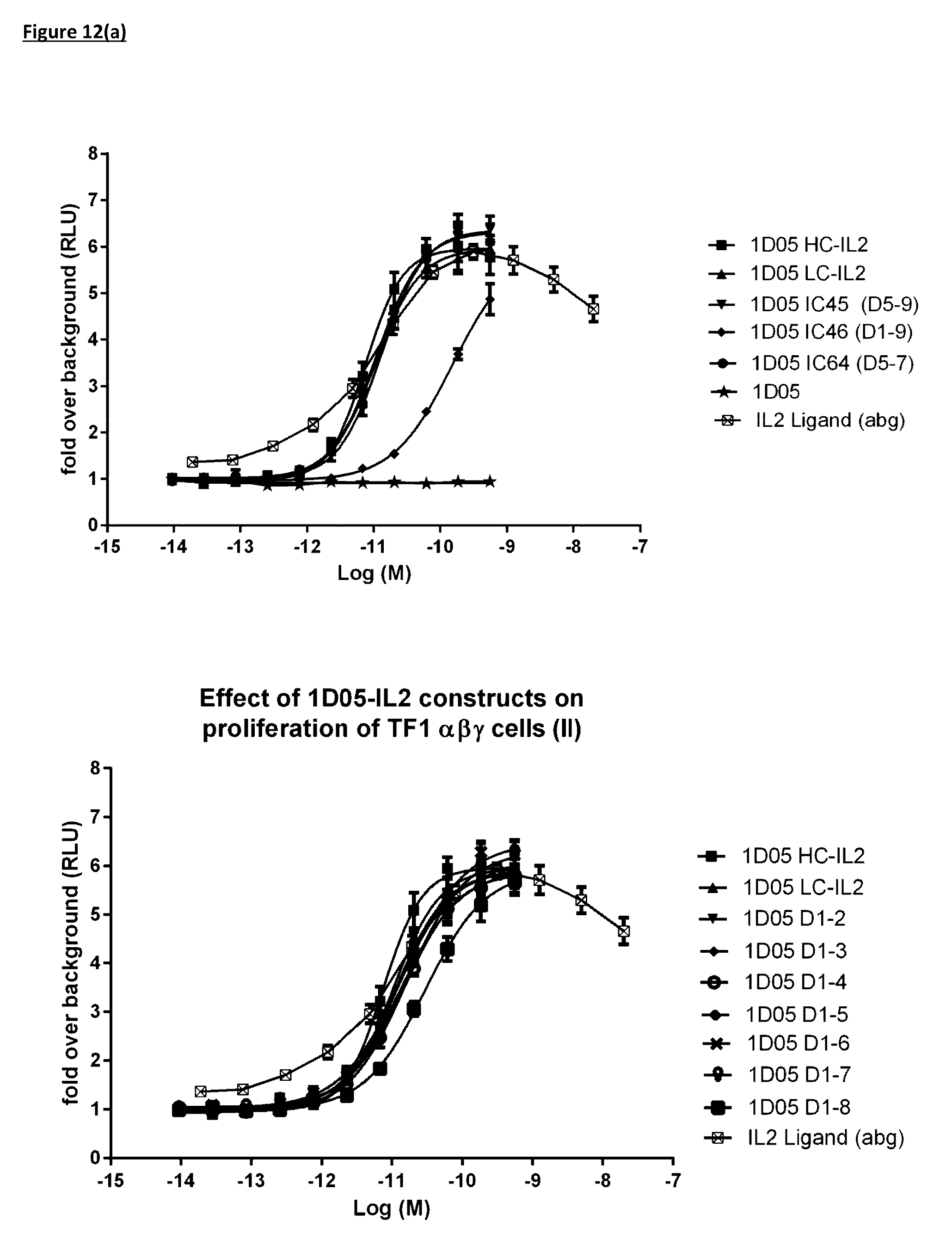
D00019
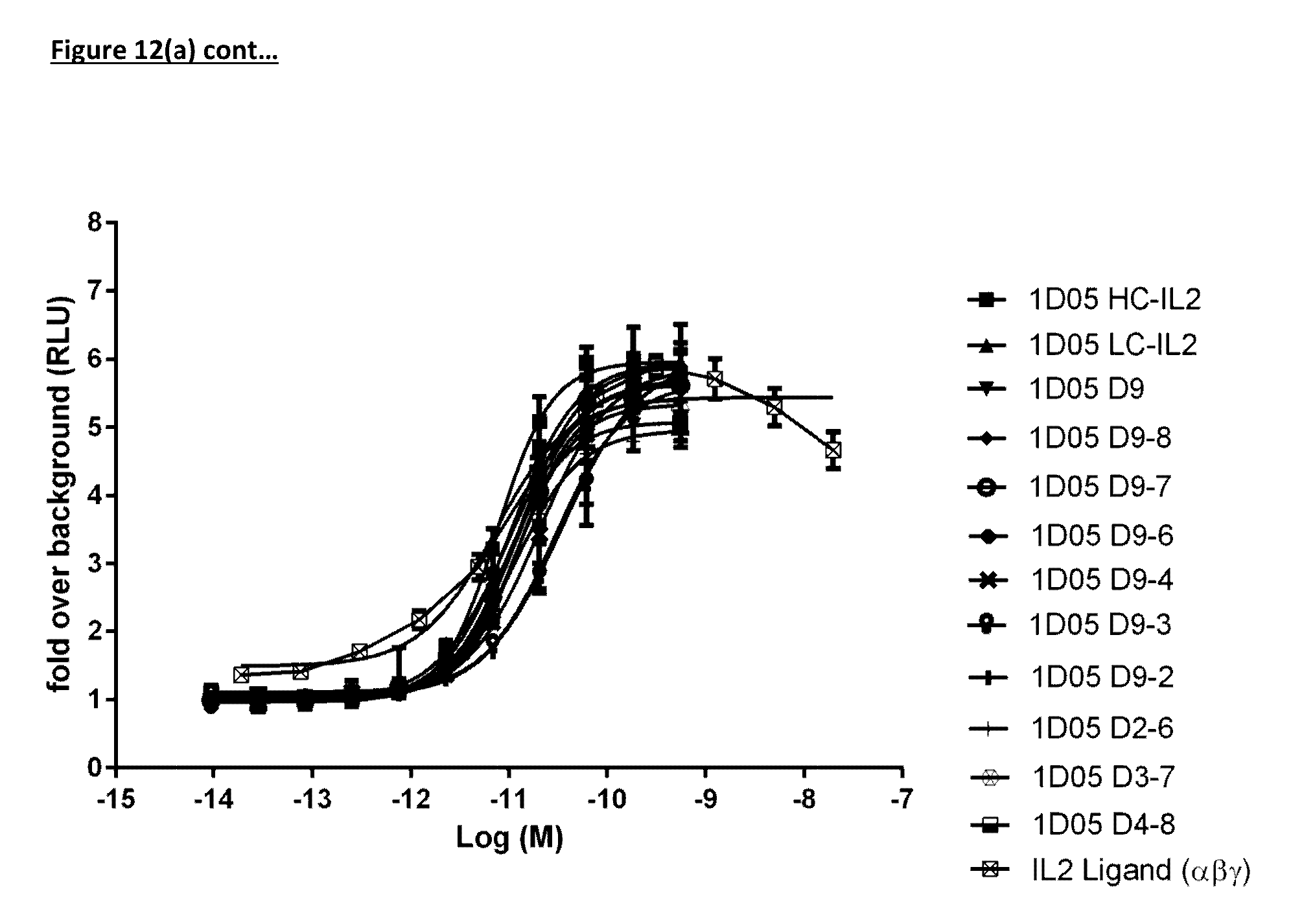
D00020
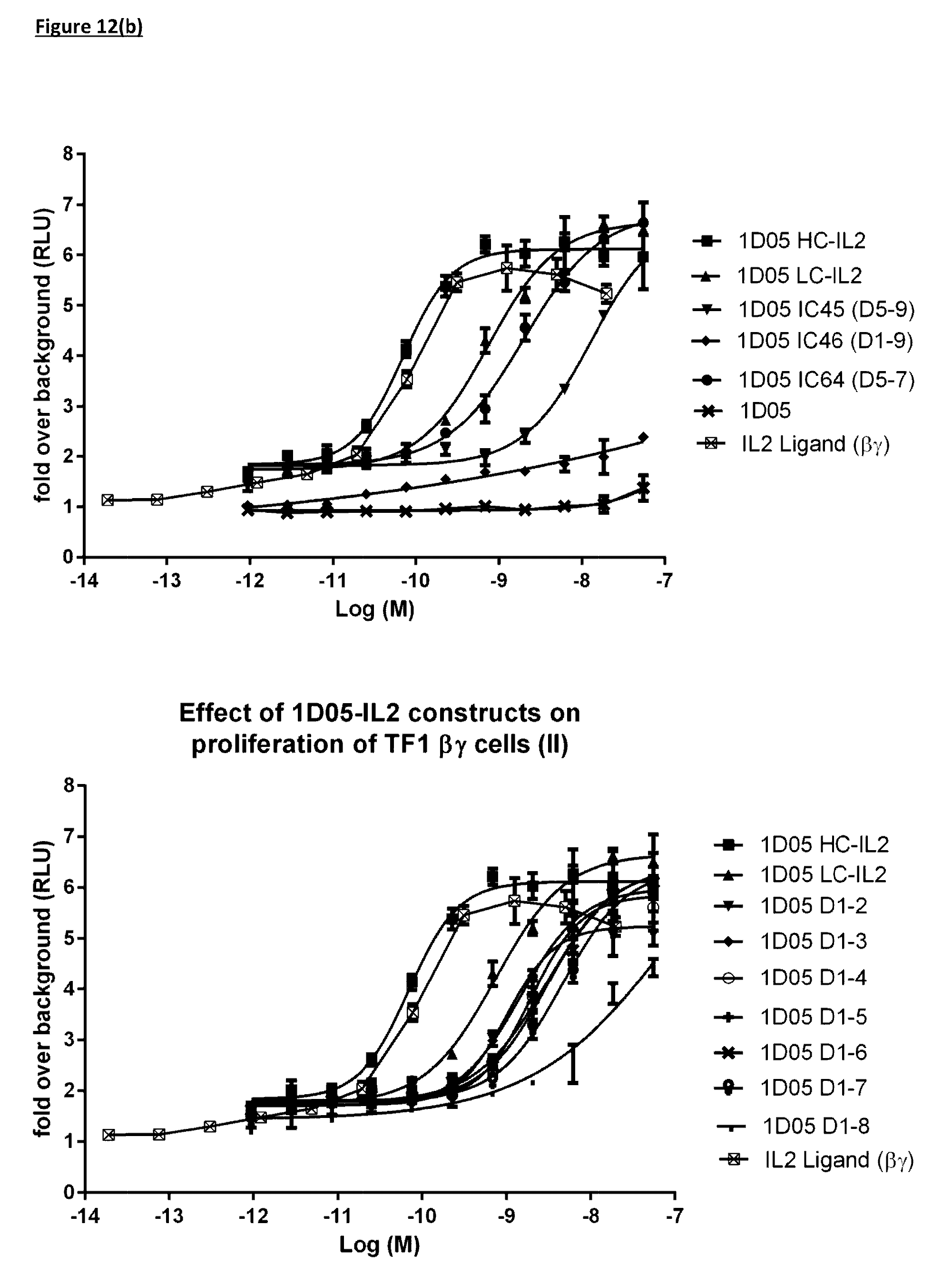
D00021
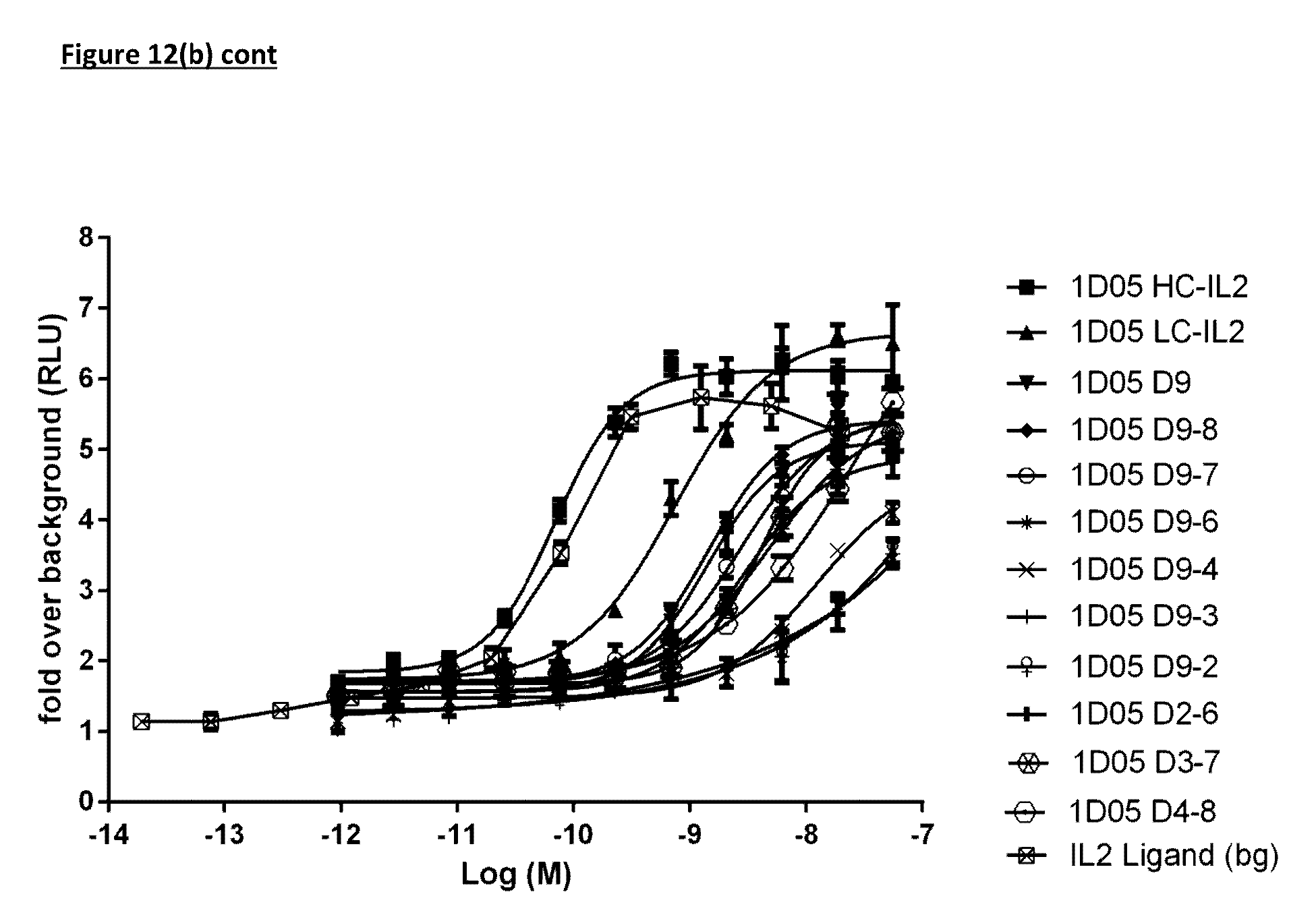
D00022
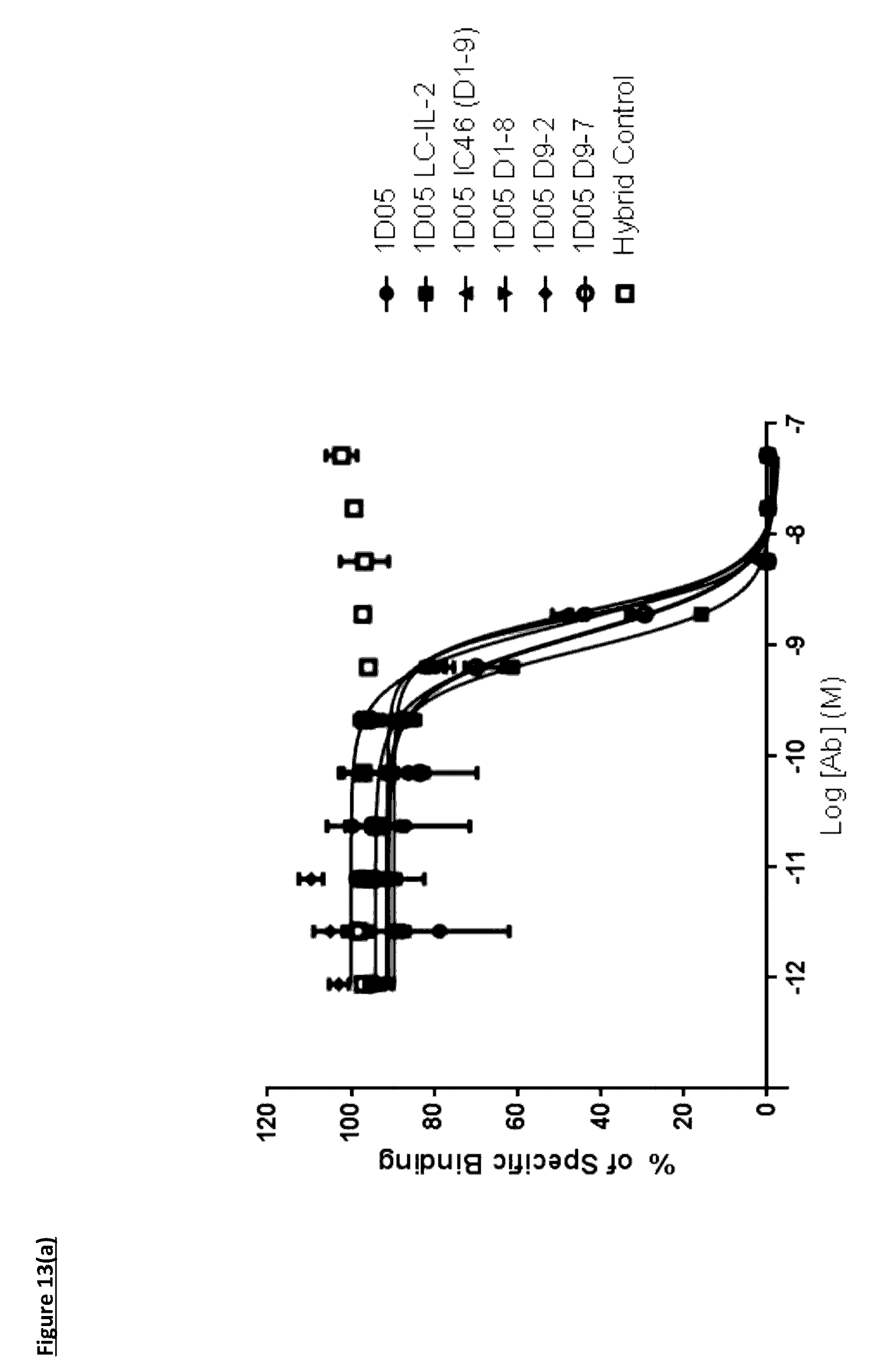
D00023
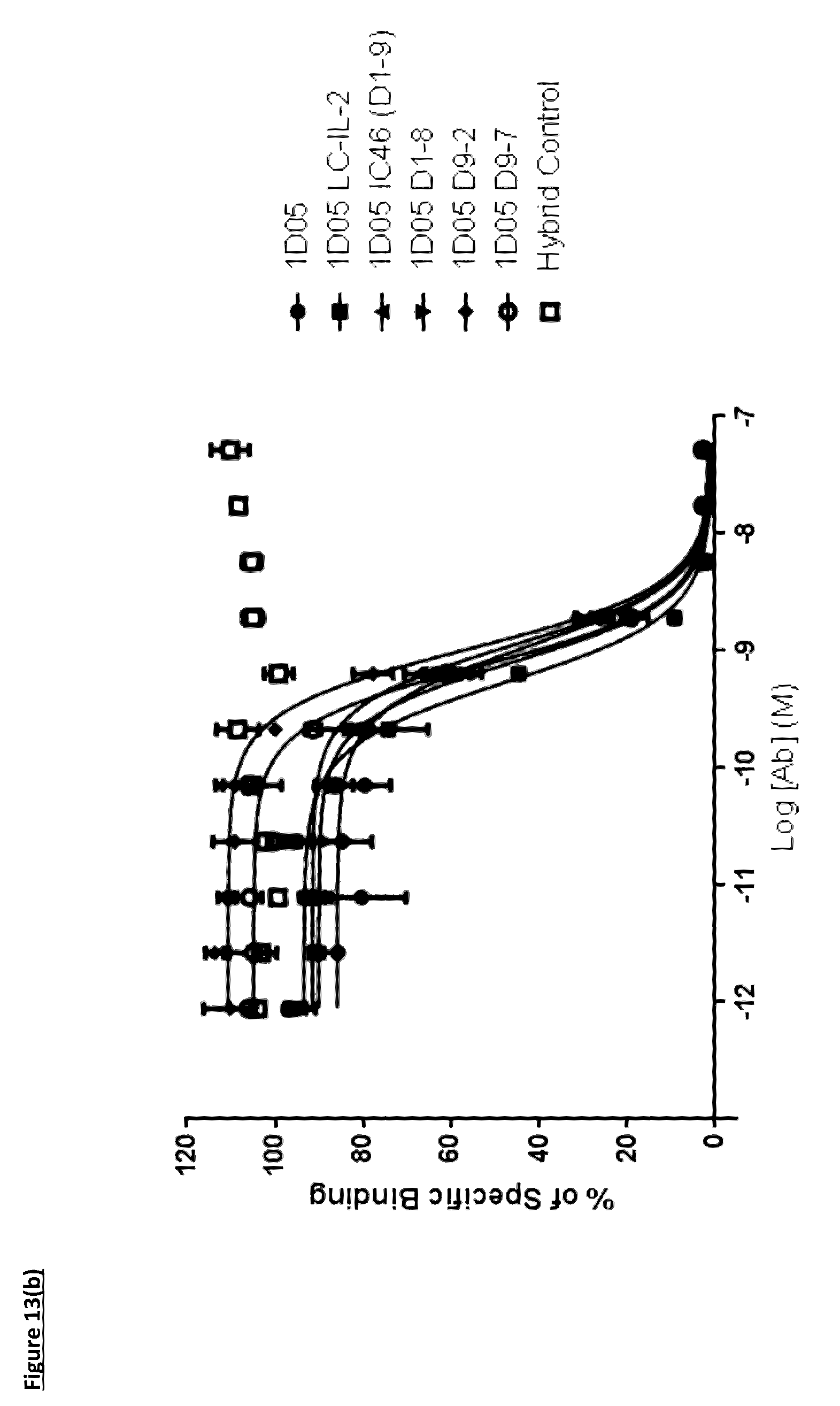
D00024
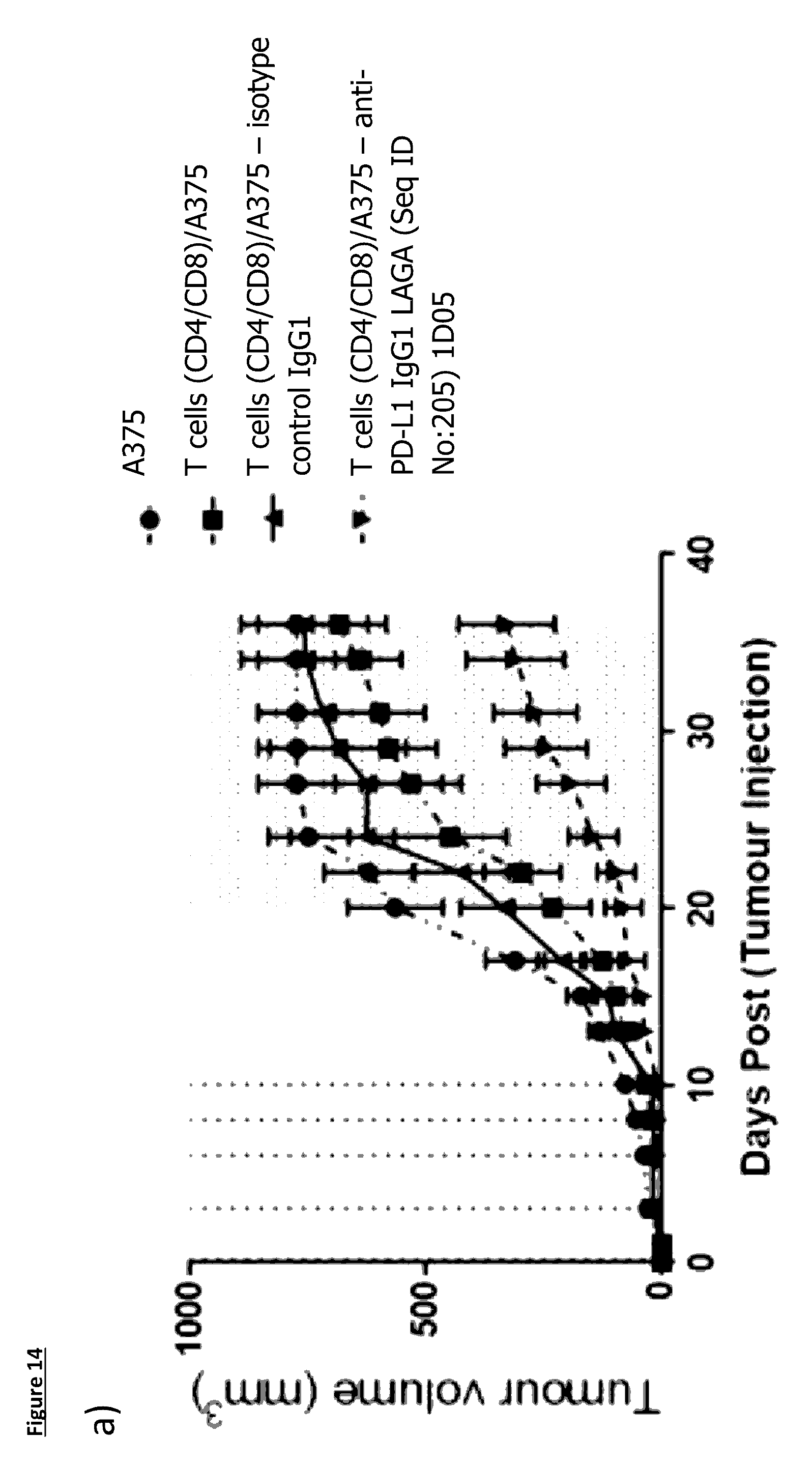
D00025
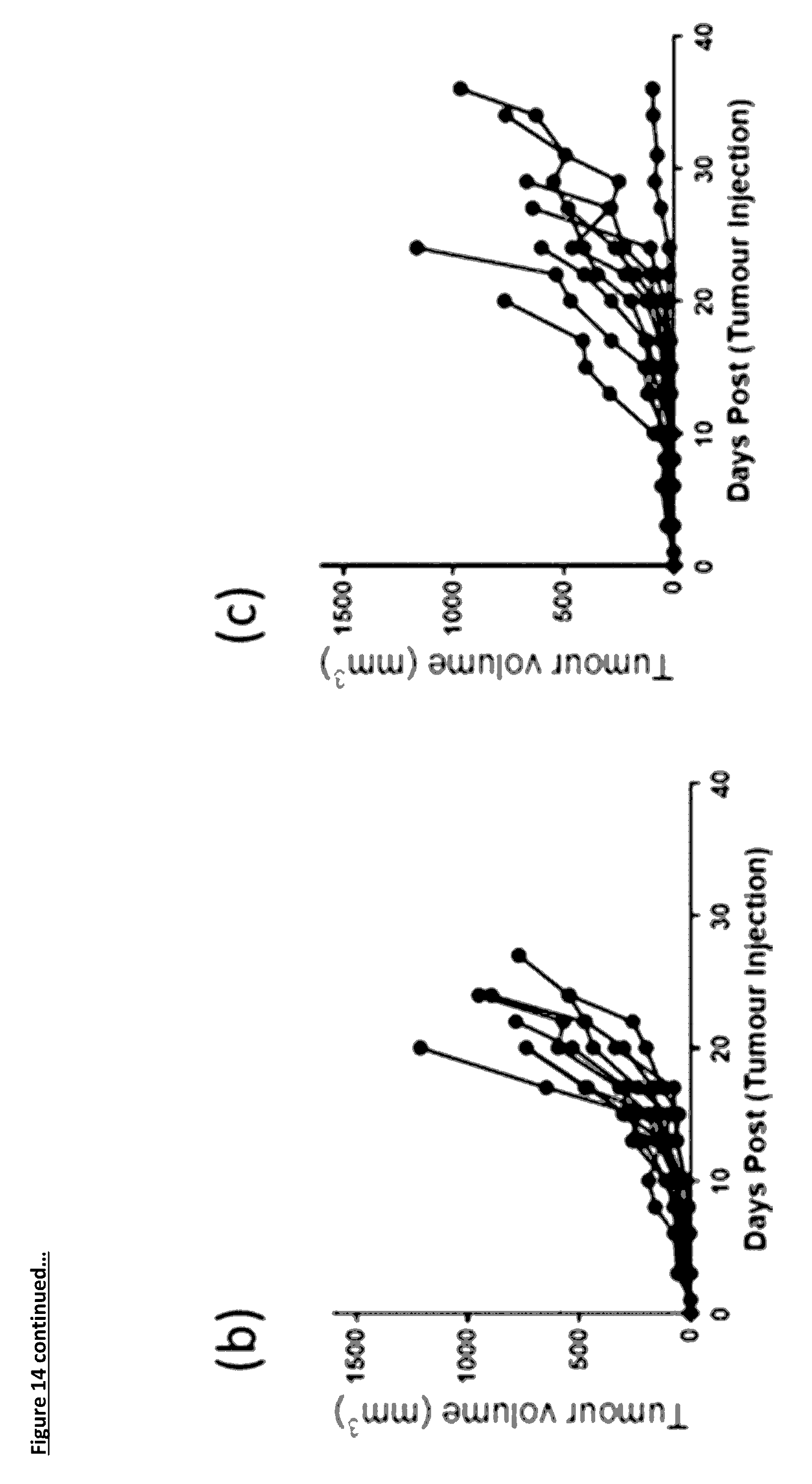
D00026
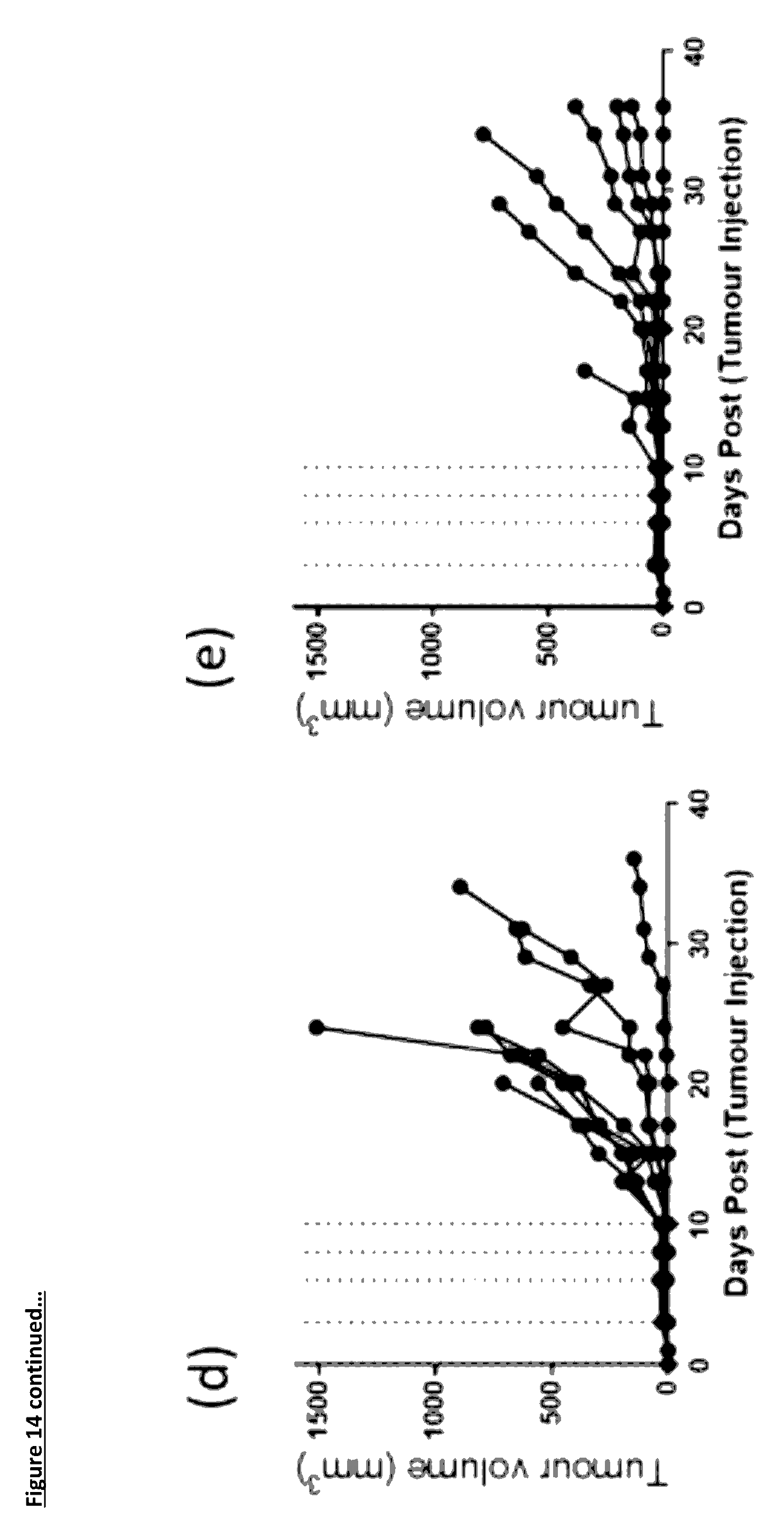
D00027
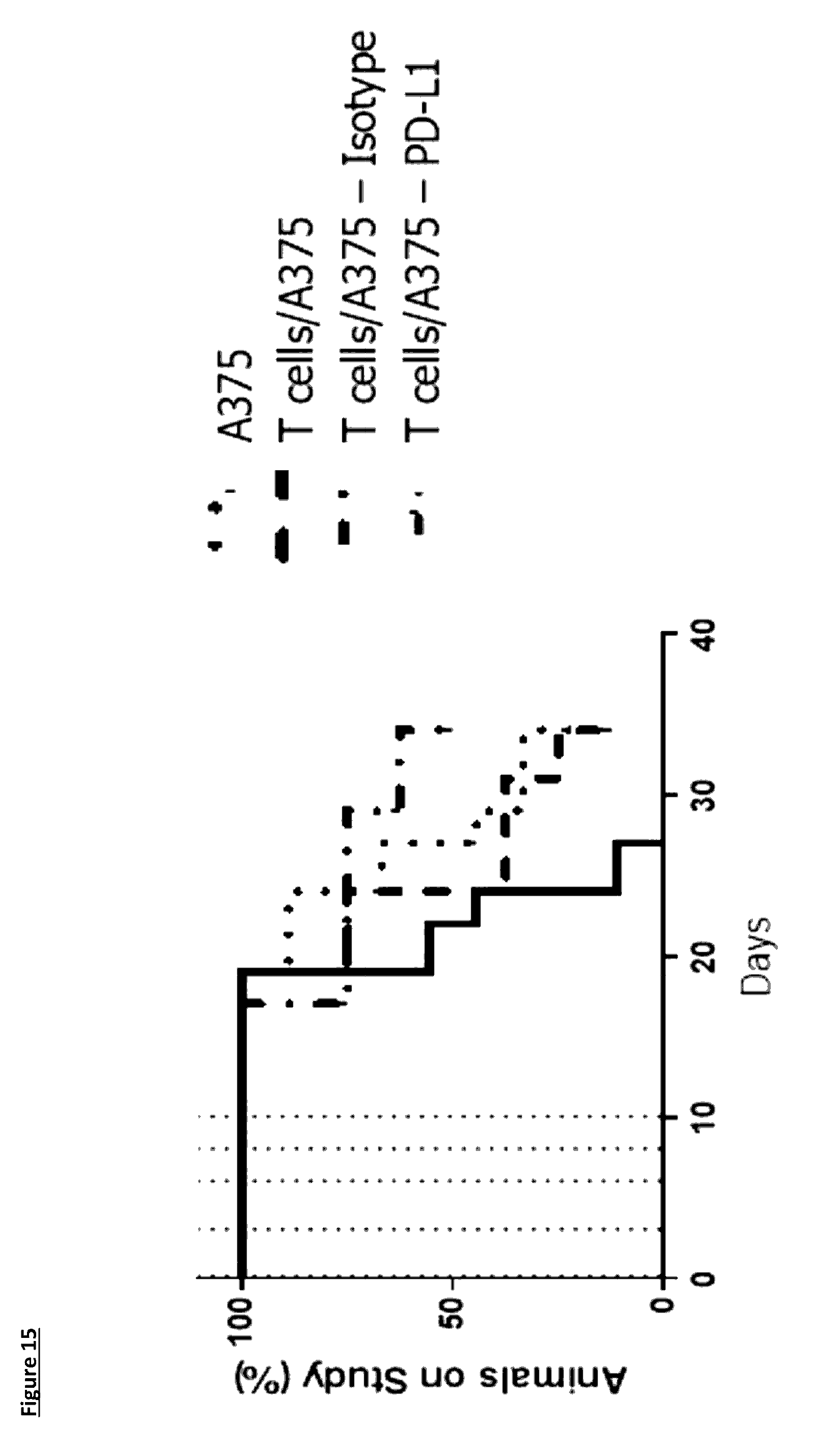
D00028
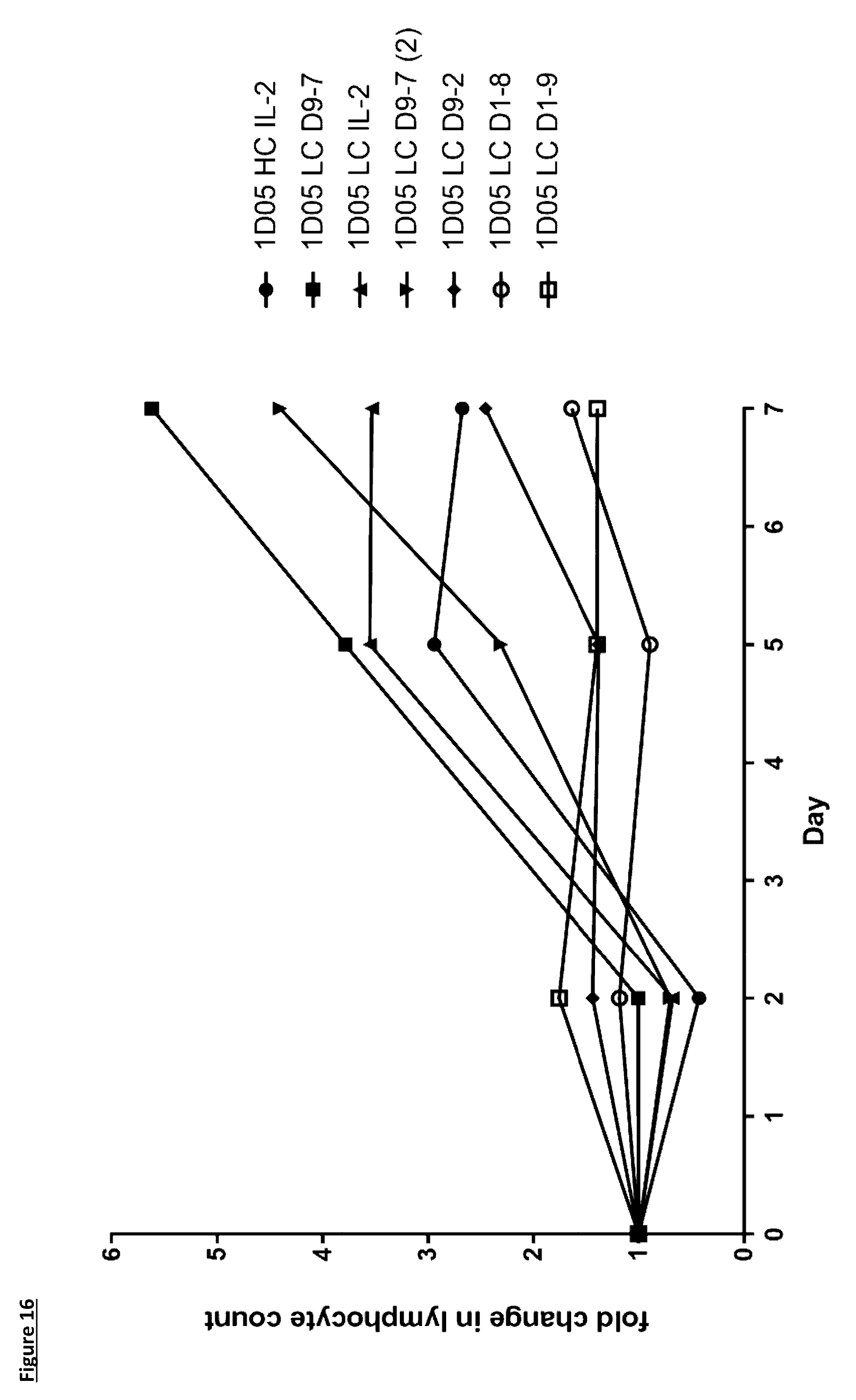
D00029
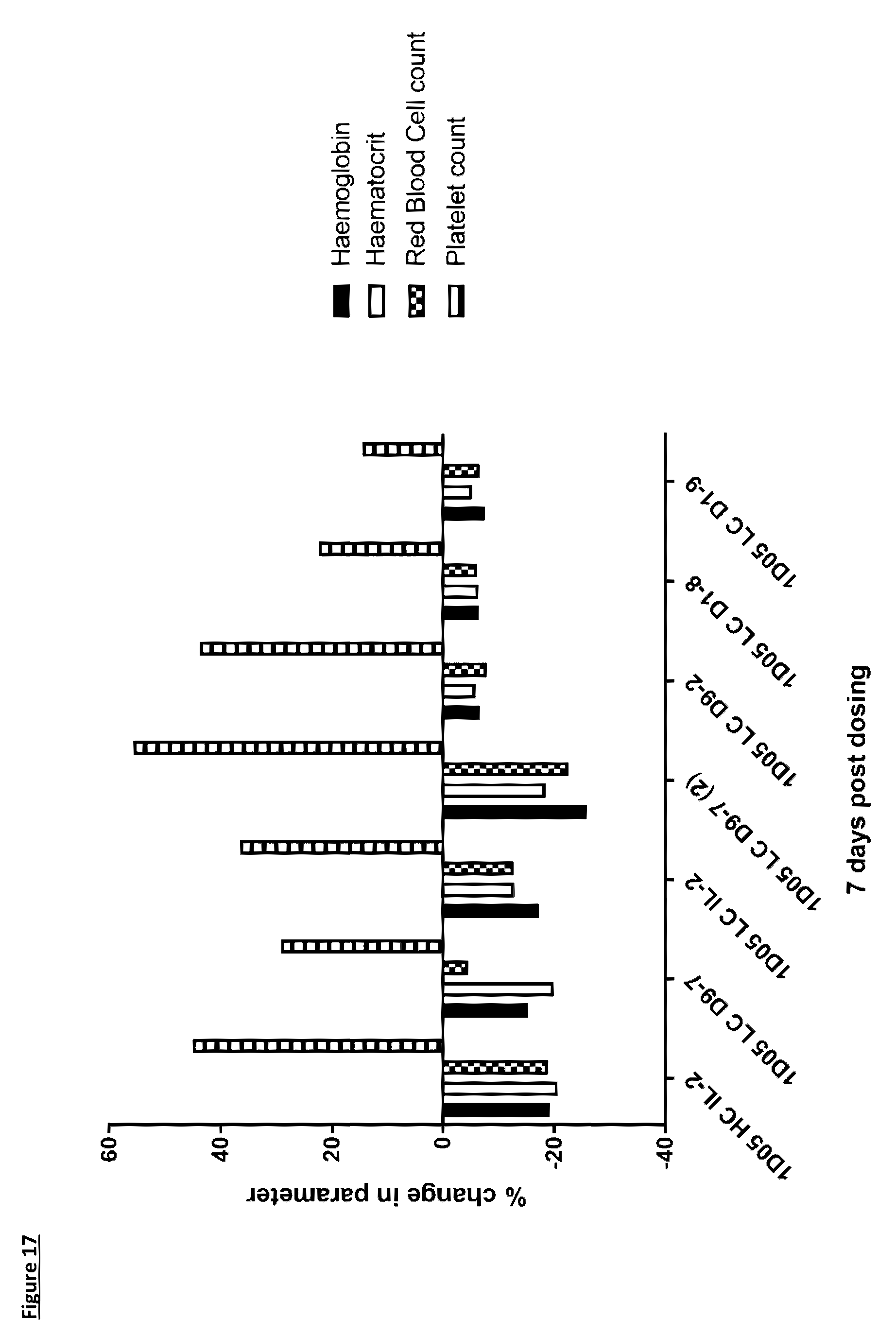
D00030

D00031
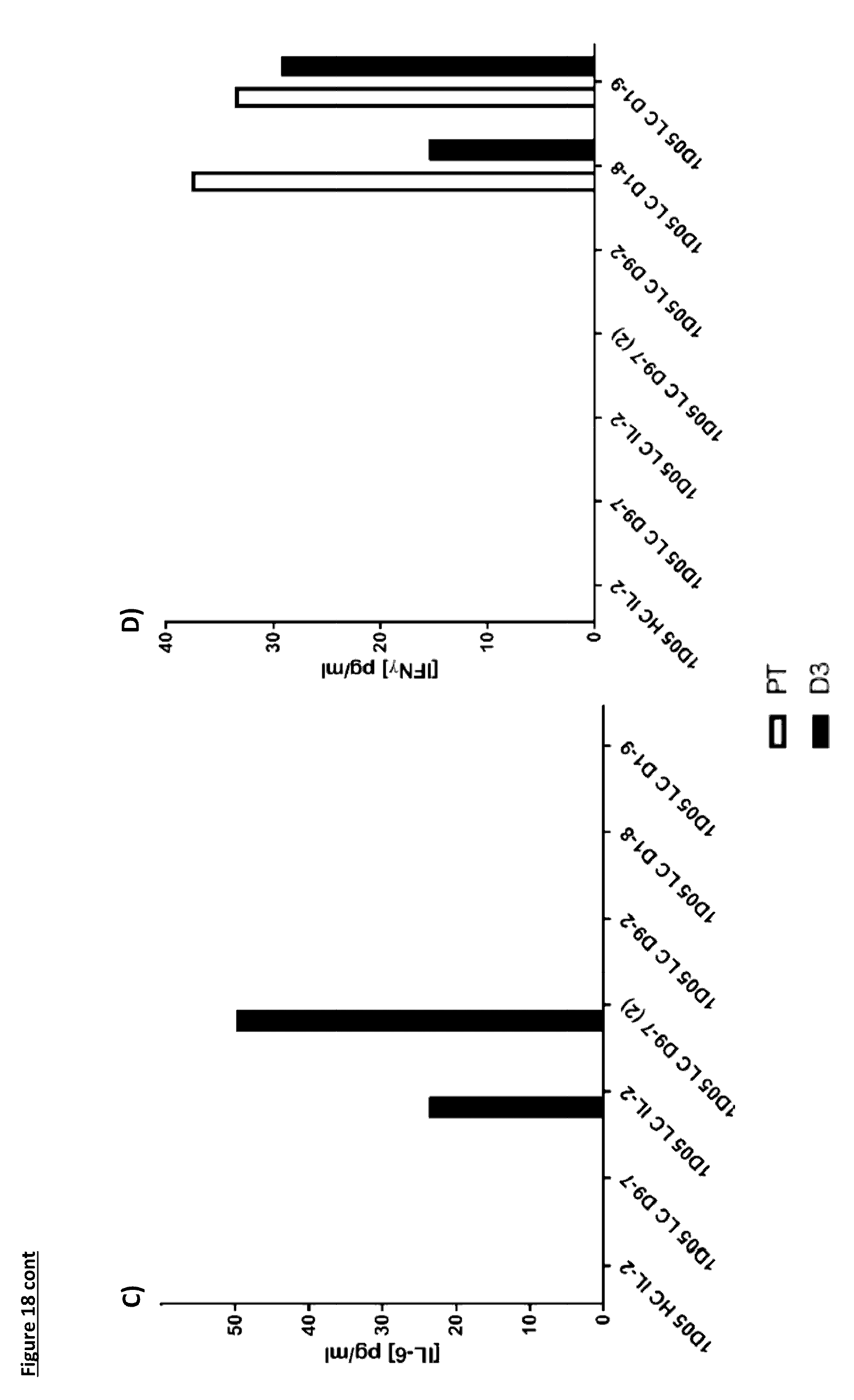

D00032
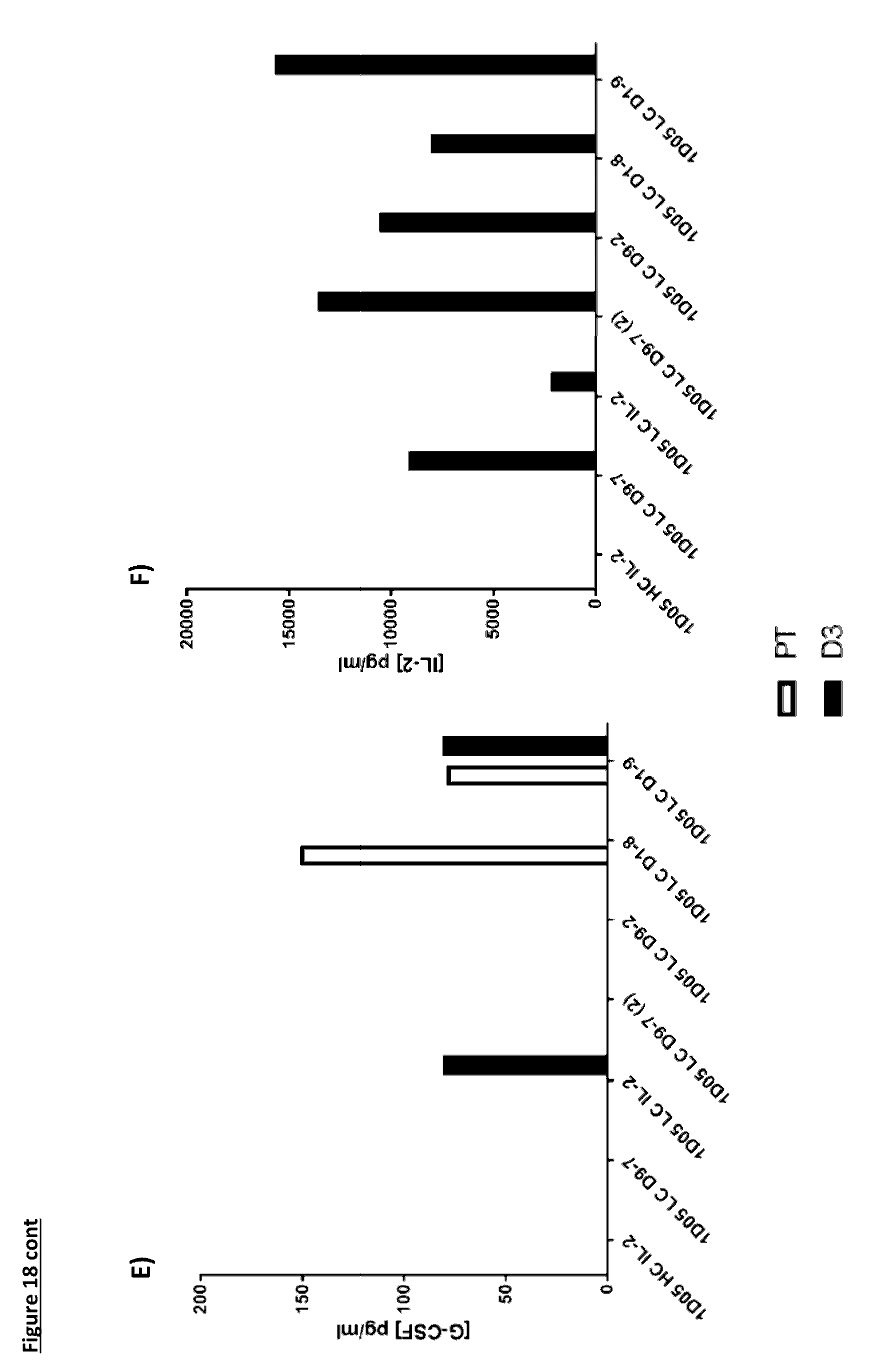
D00033
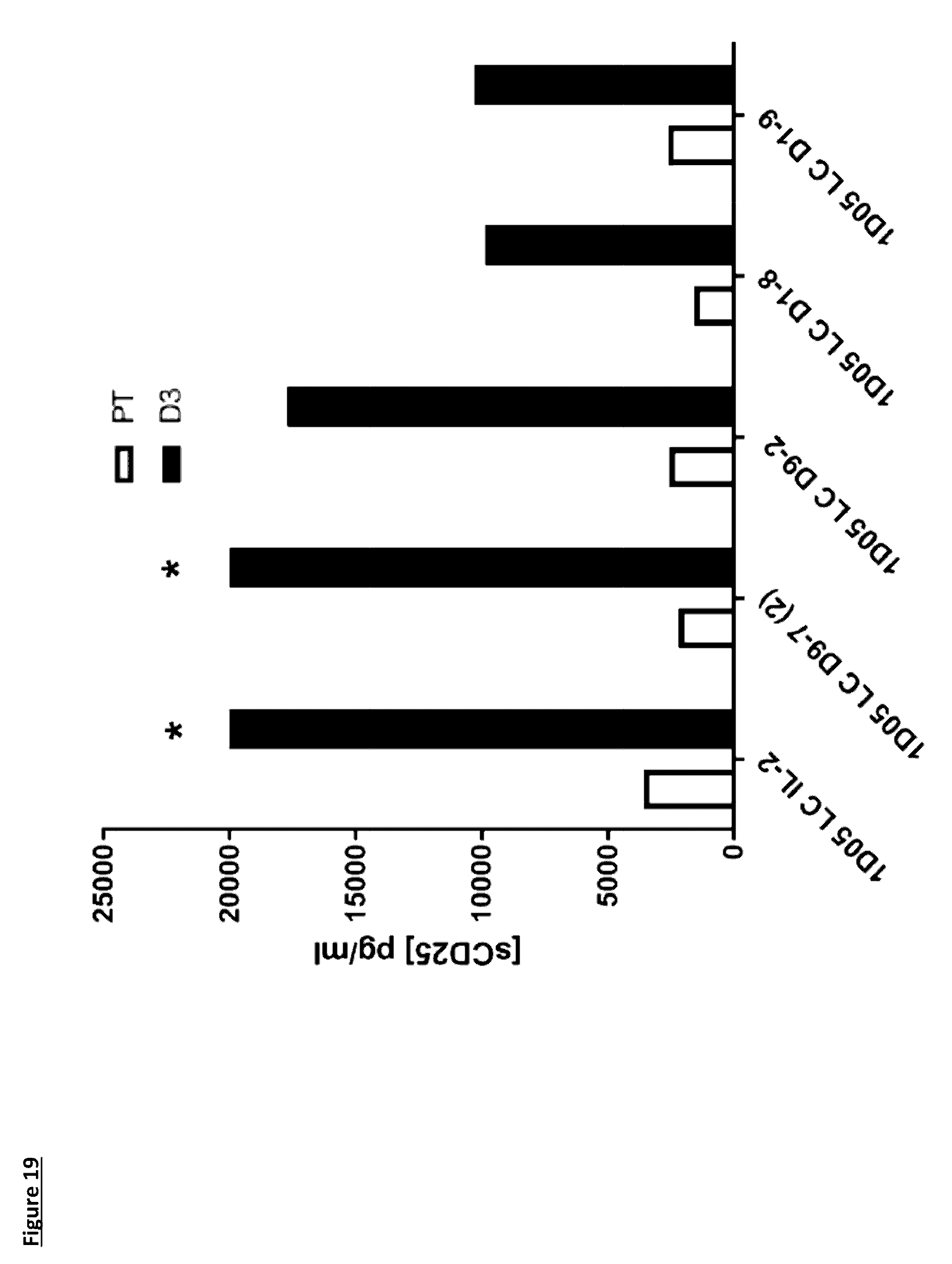
D00034
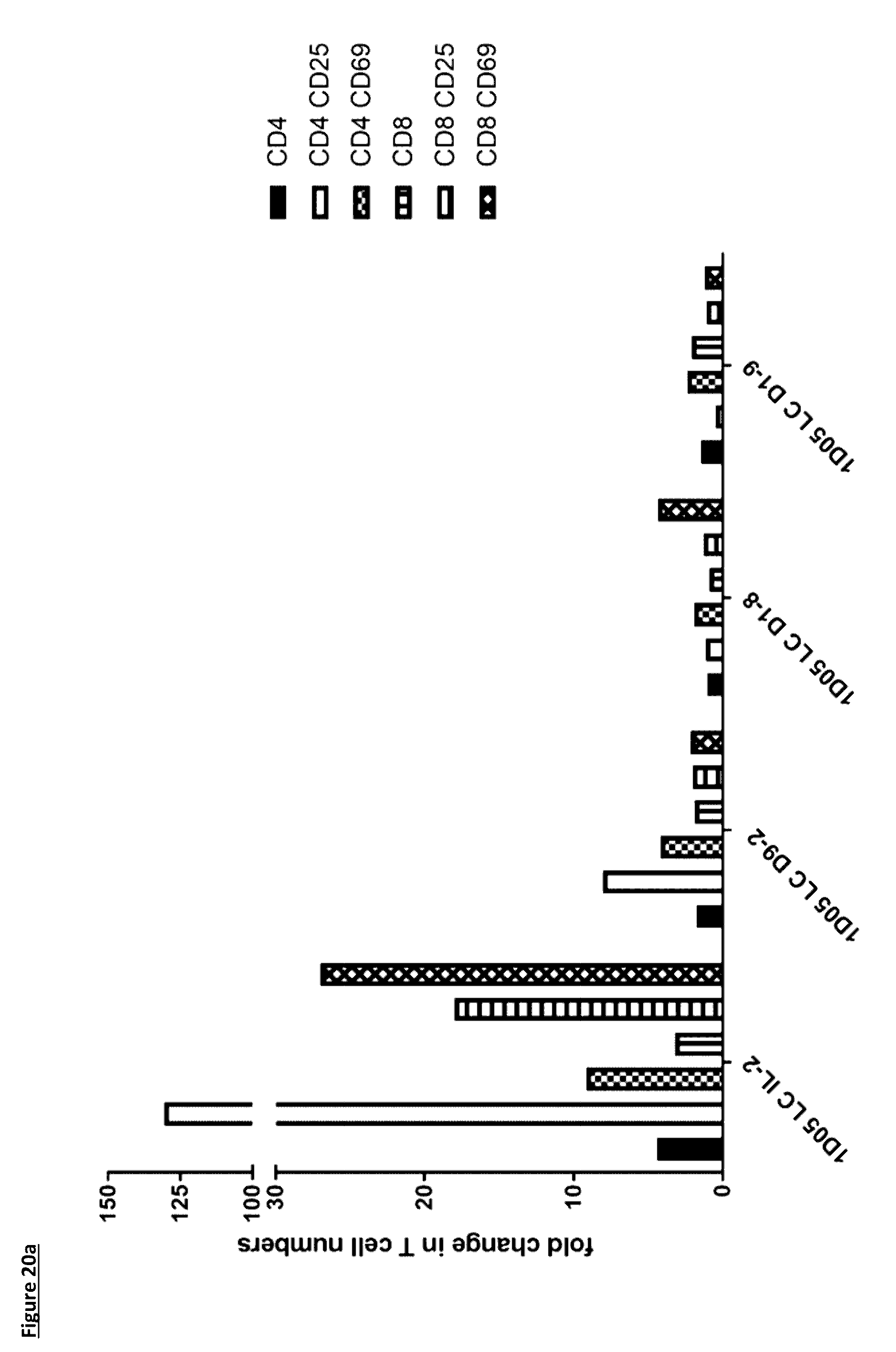
D00035
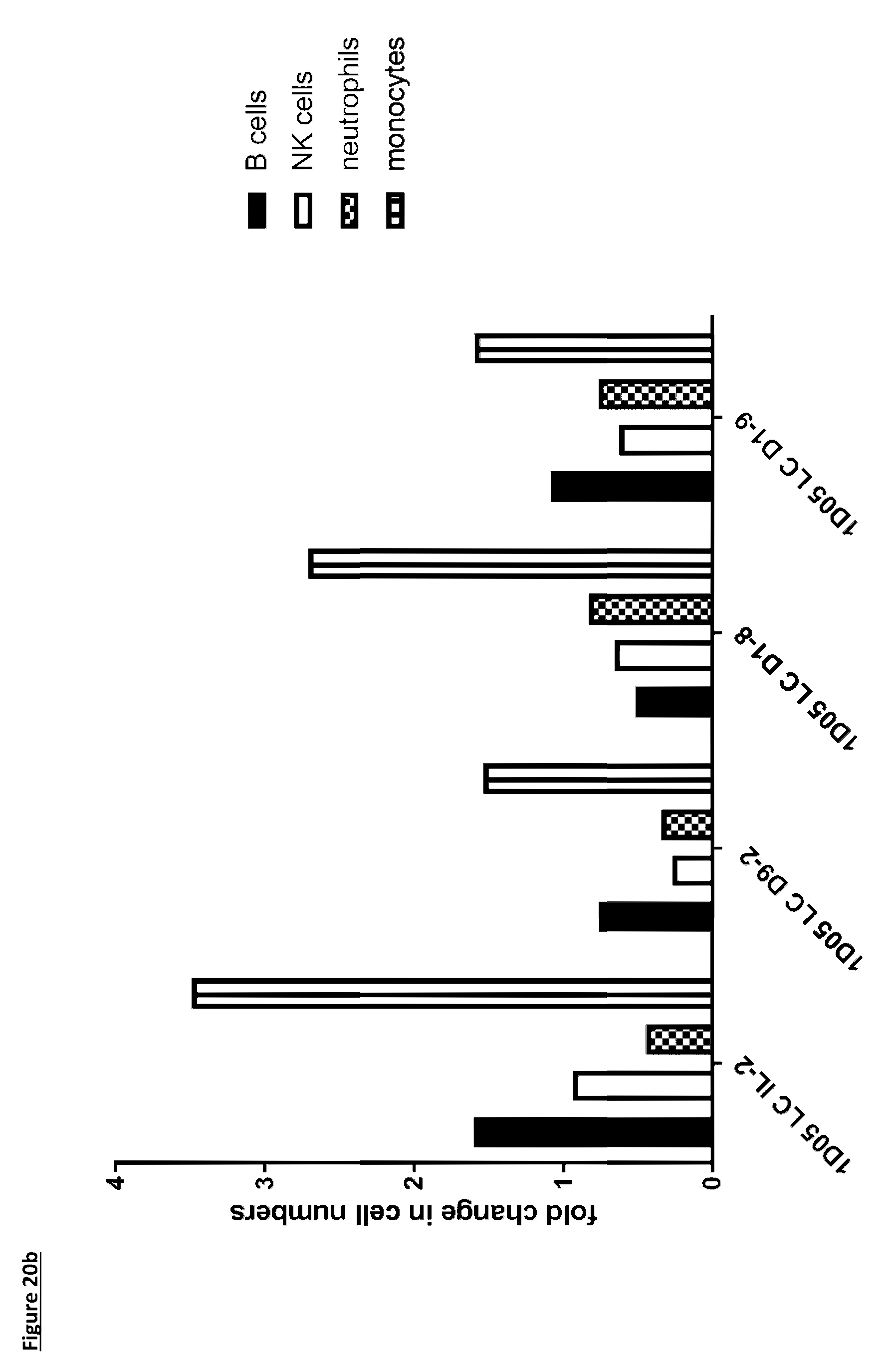
D00036
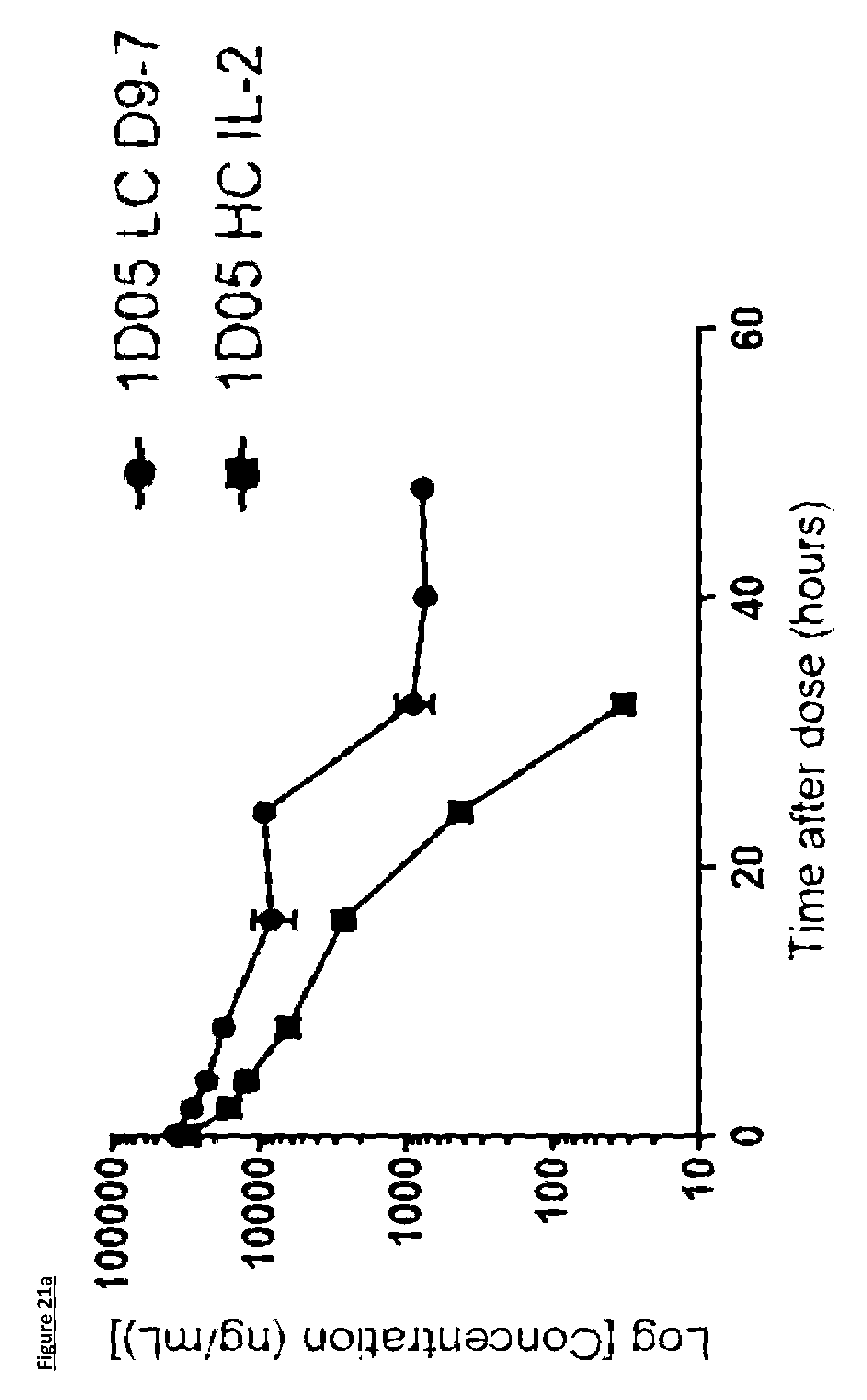
D00037
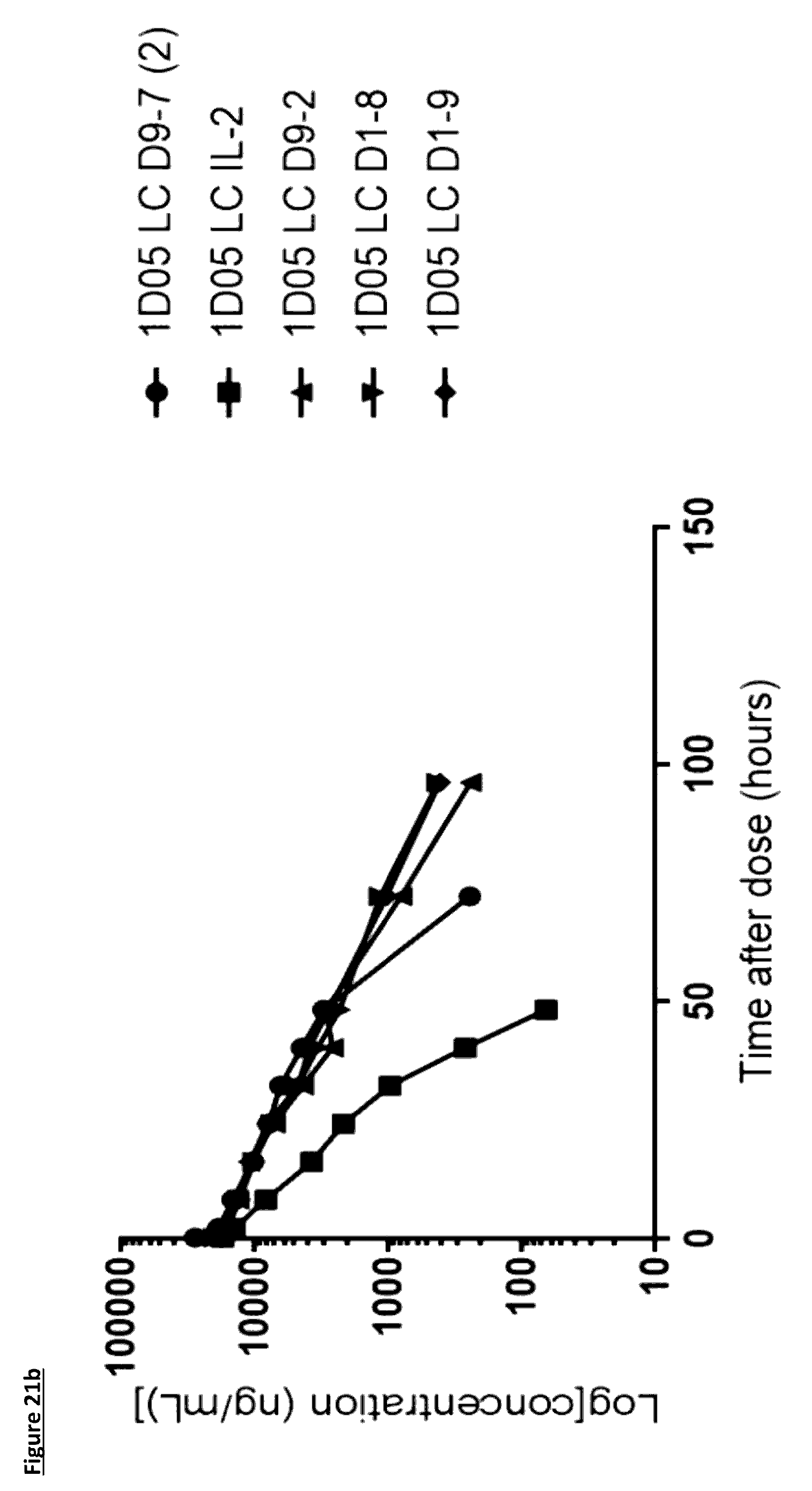
D00038
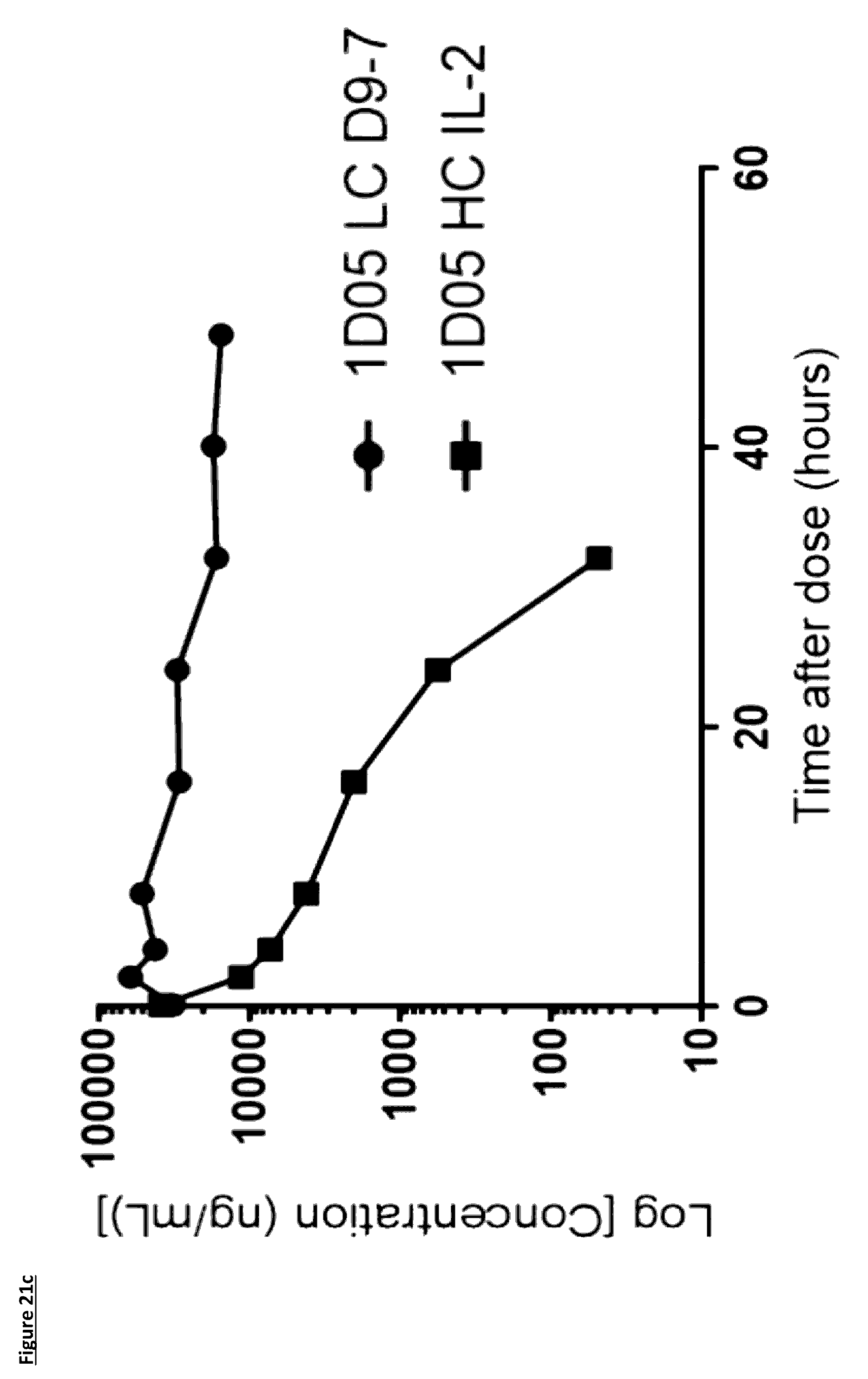
D00039
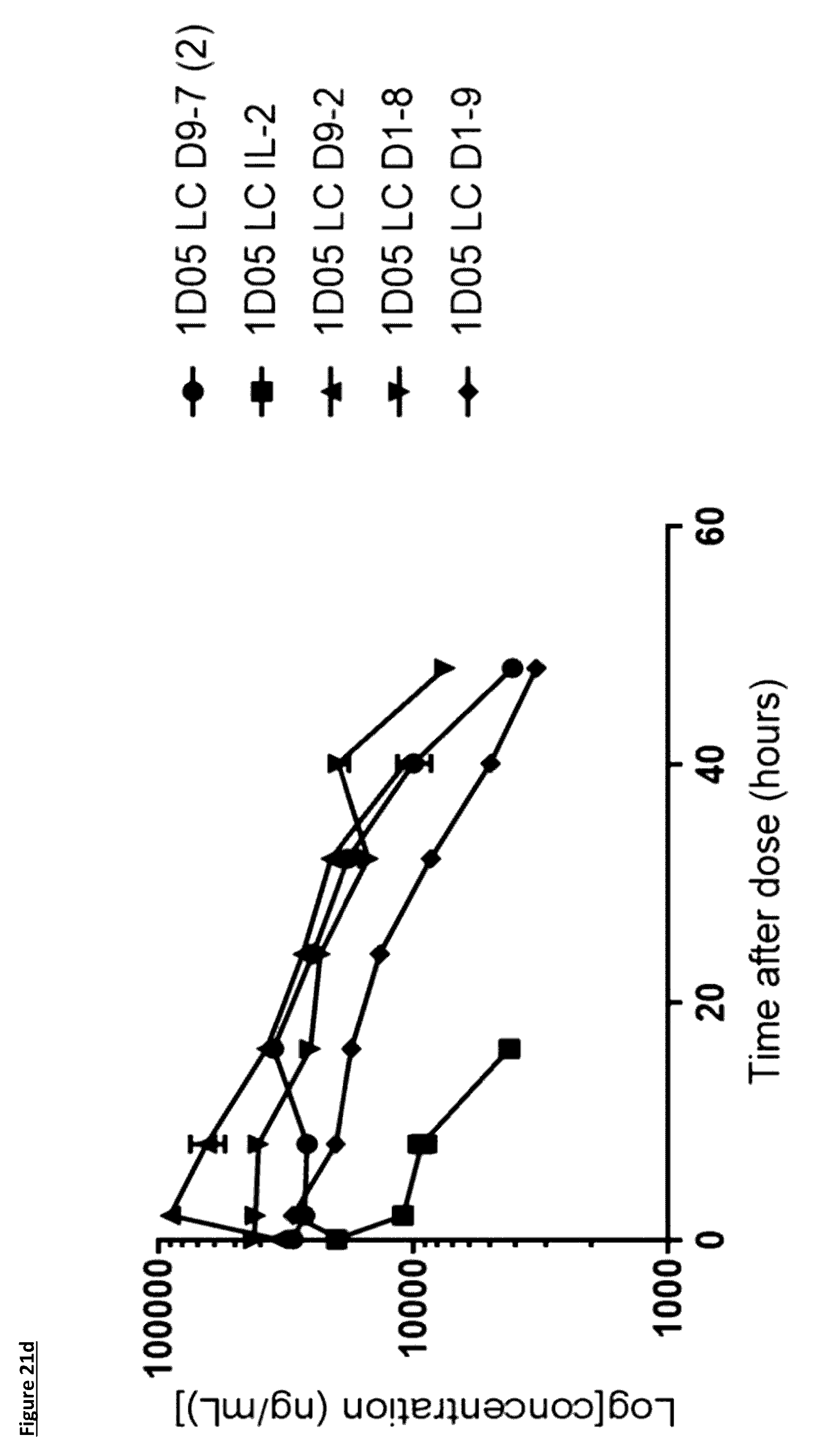
D00040
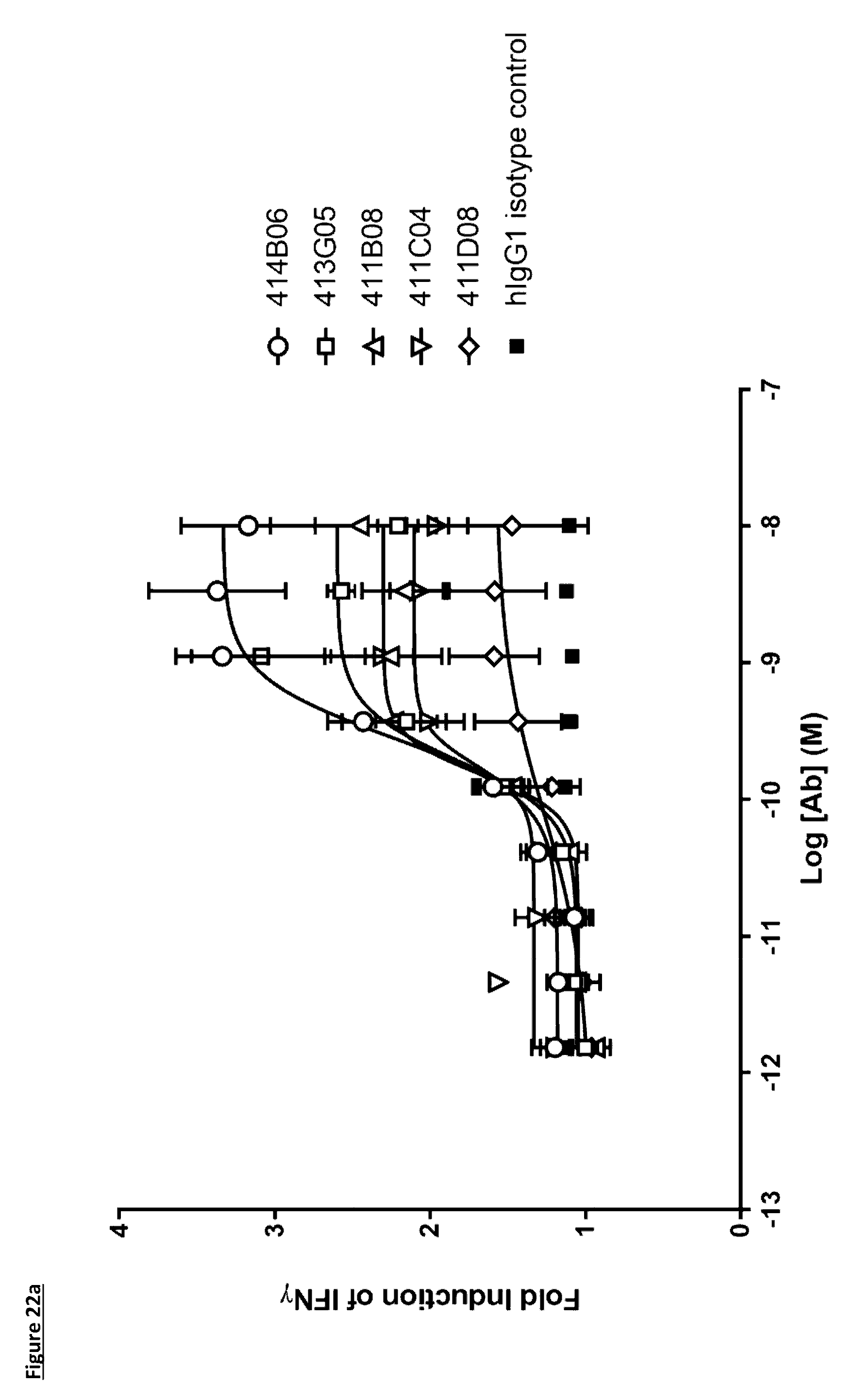
D00041
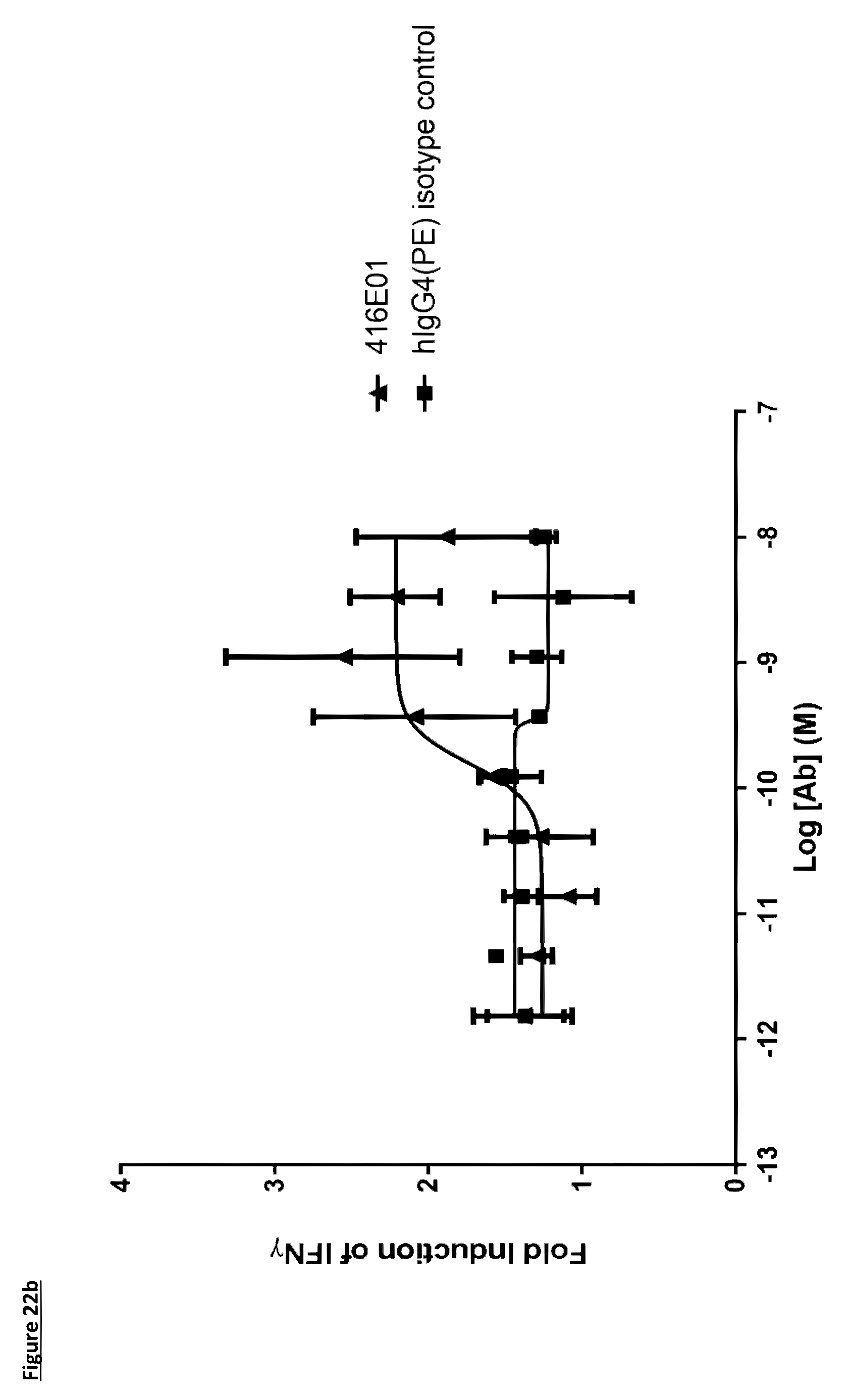
D00042
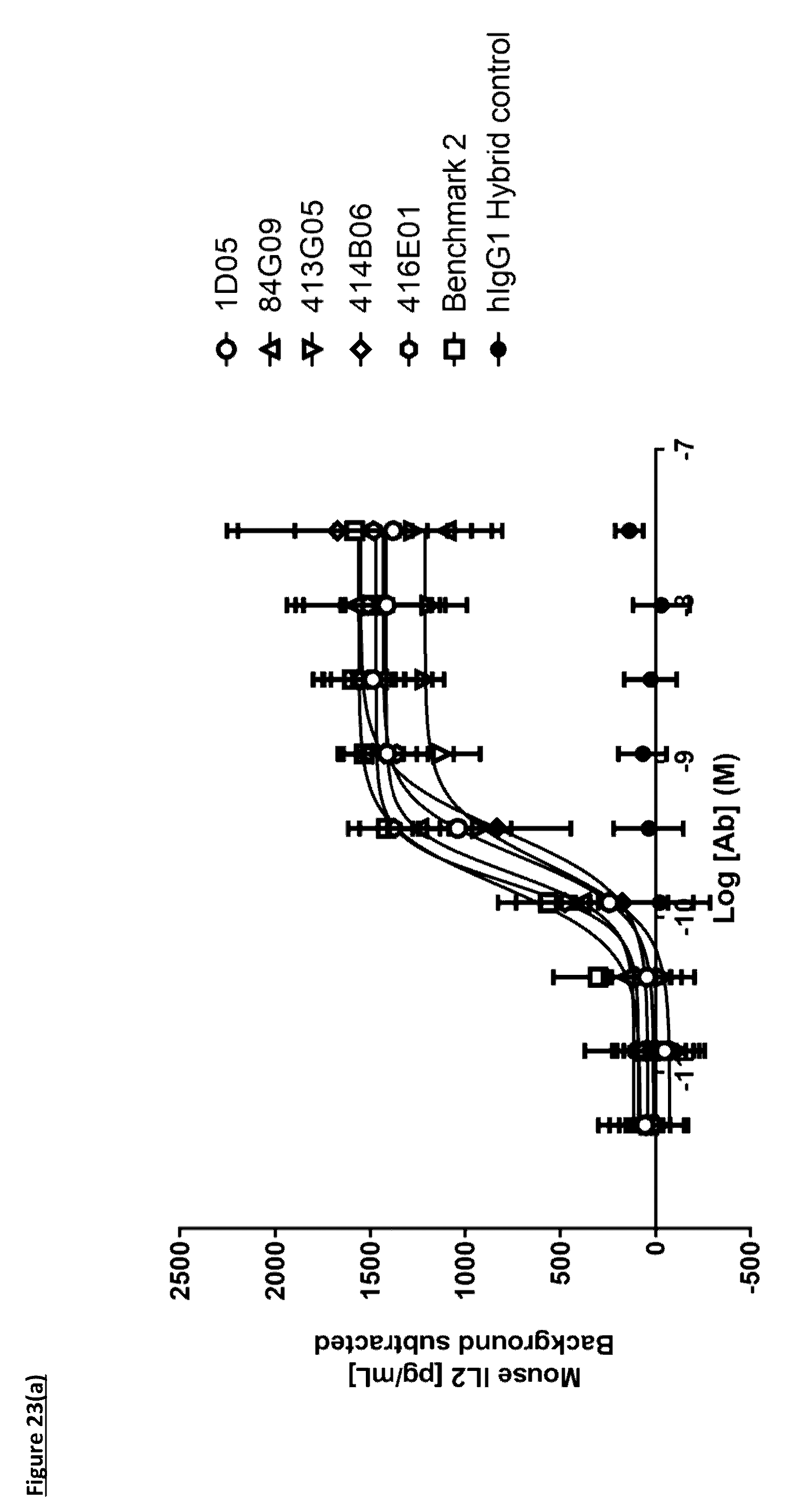
D00043
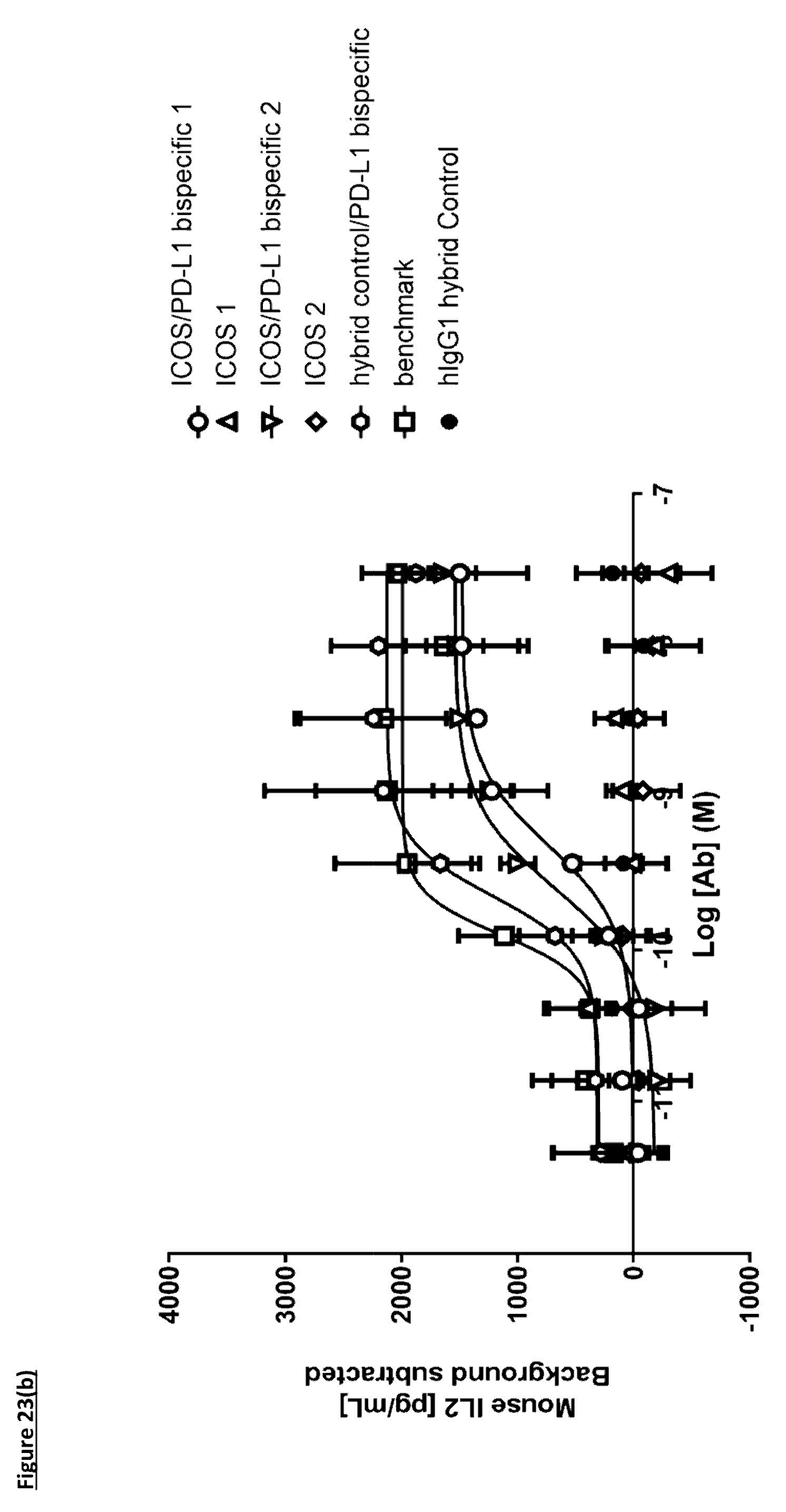
D00044
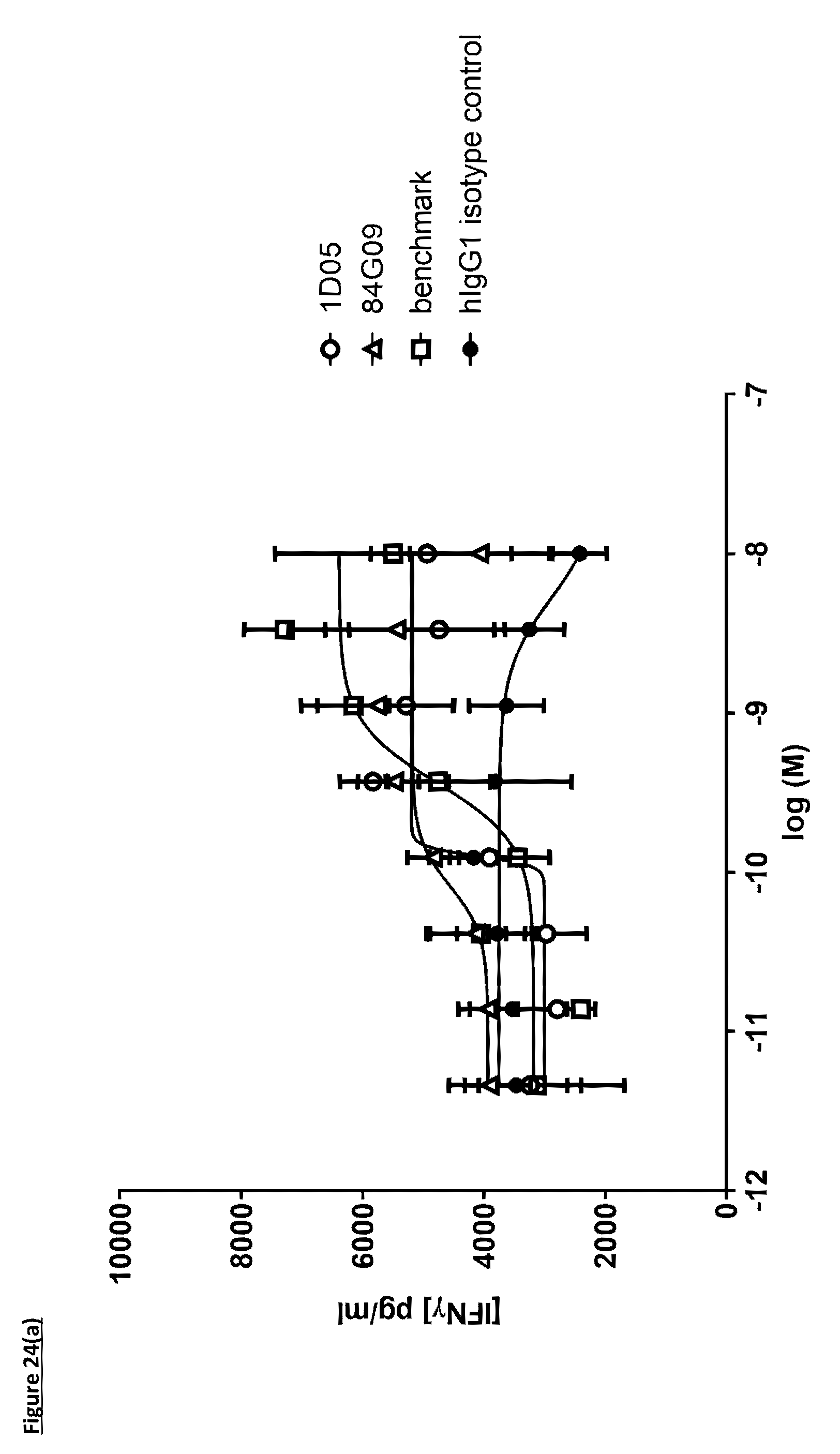
D00045
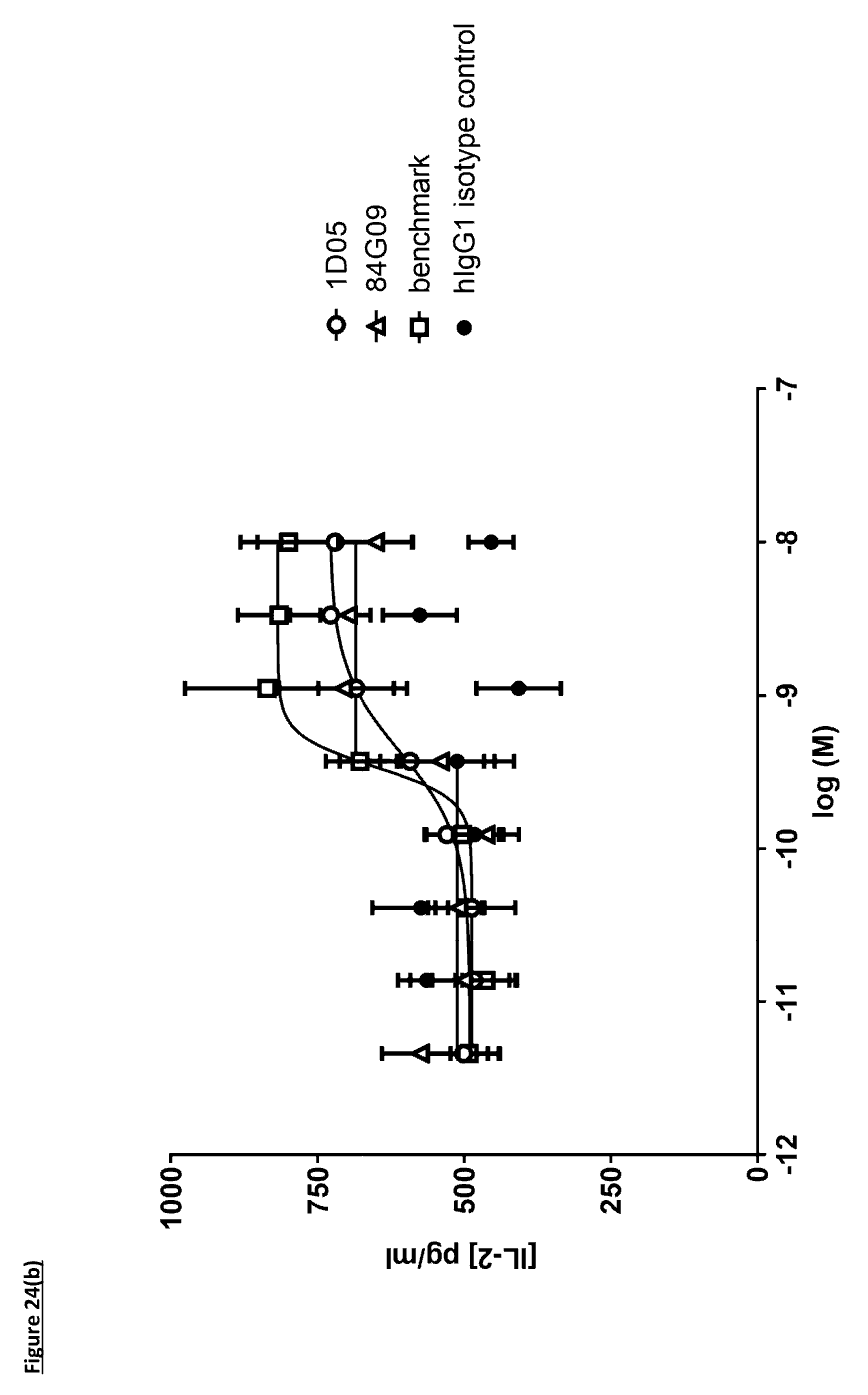
D00046
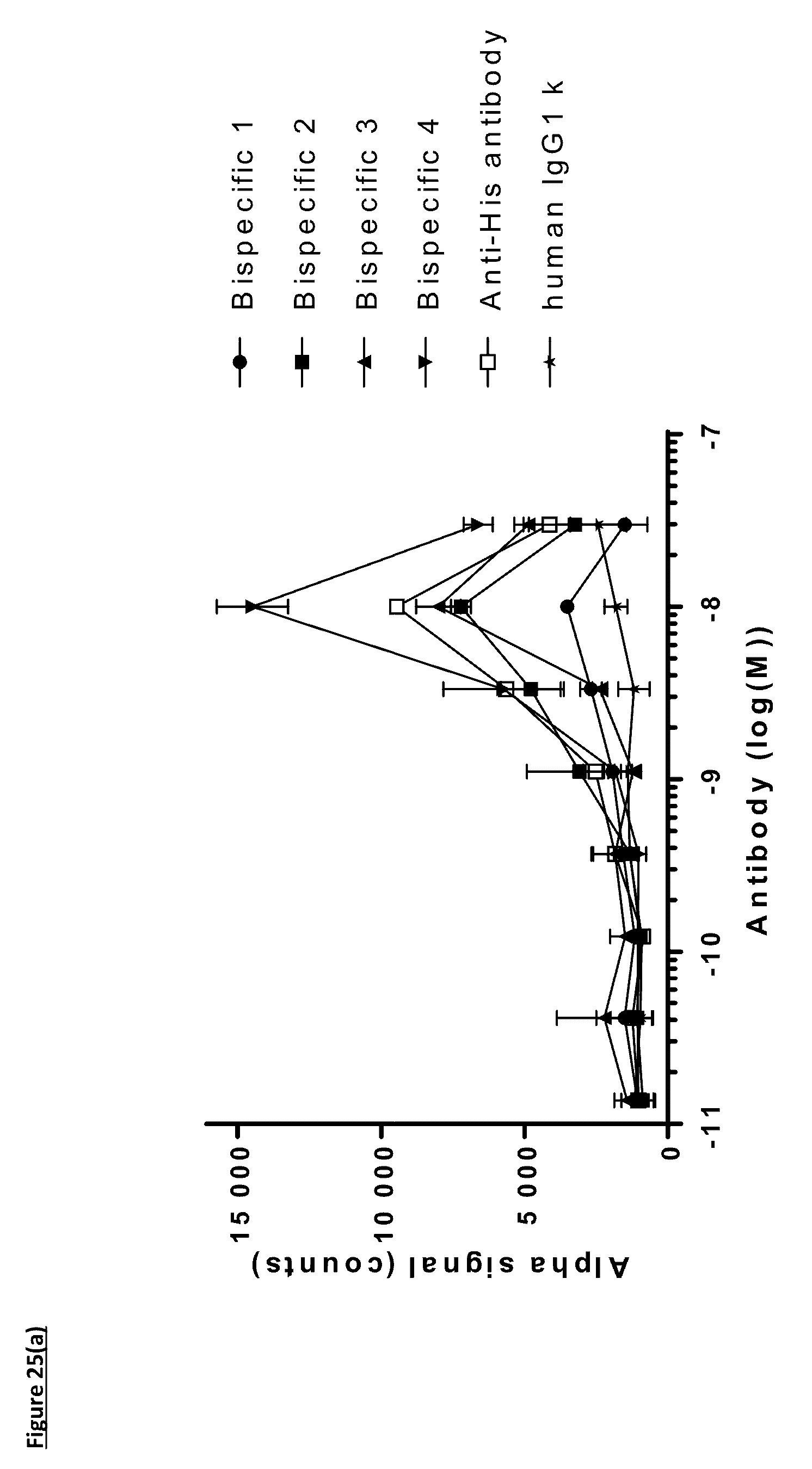
D00047
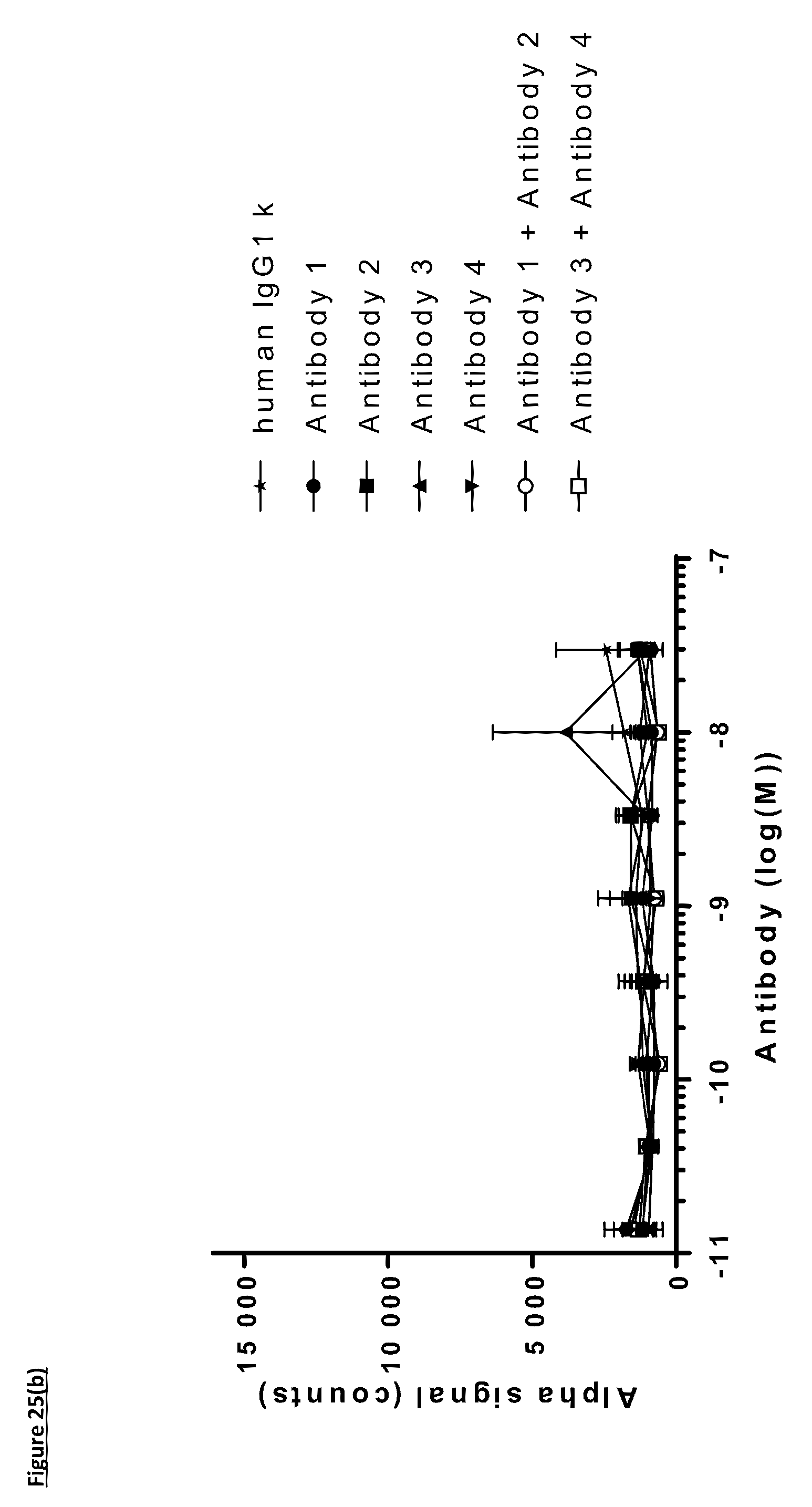
D00048
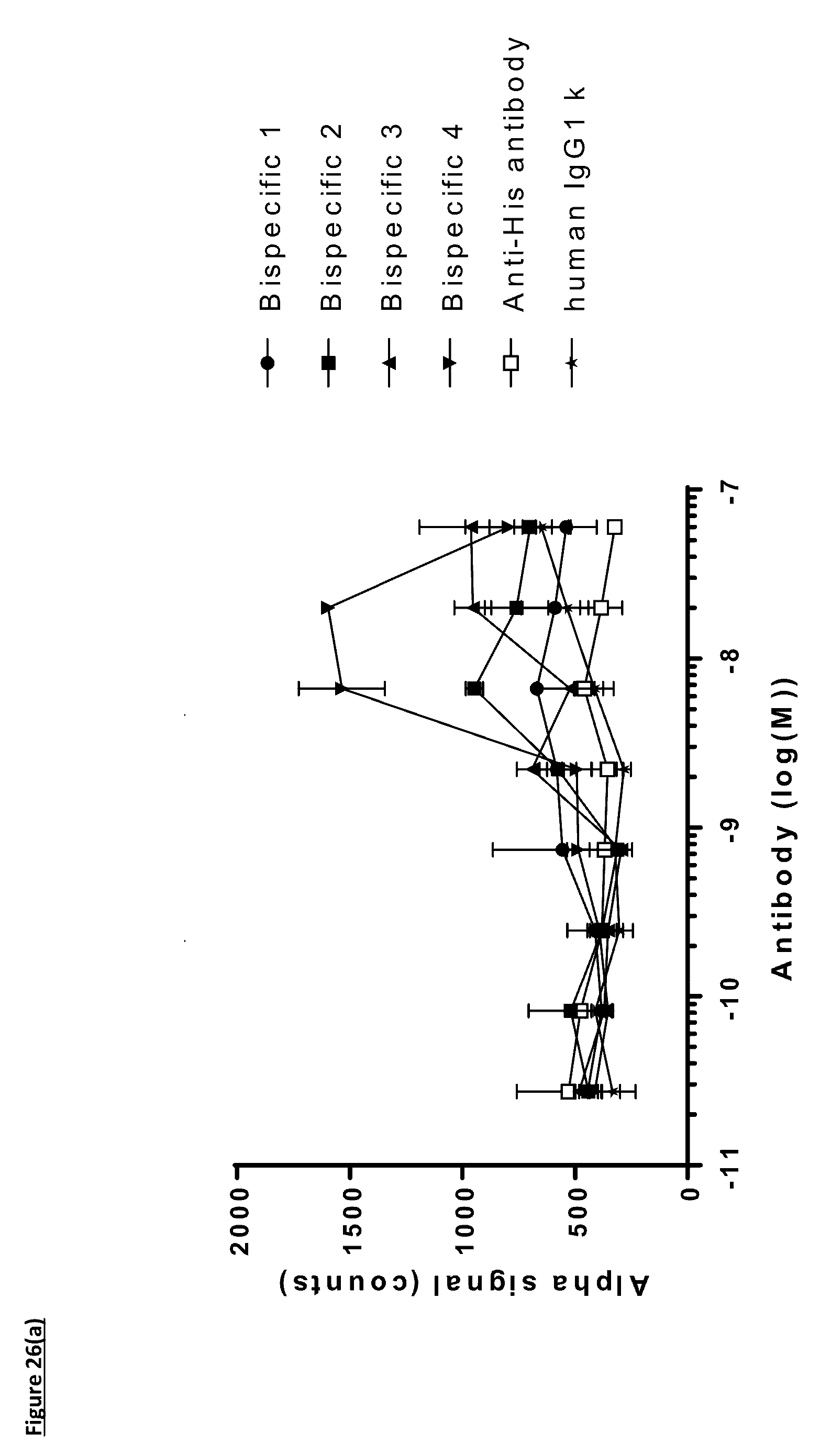
D00049
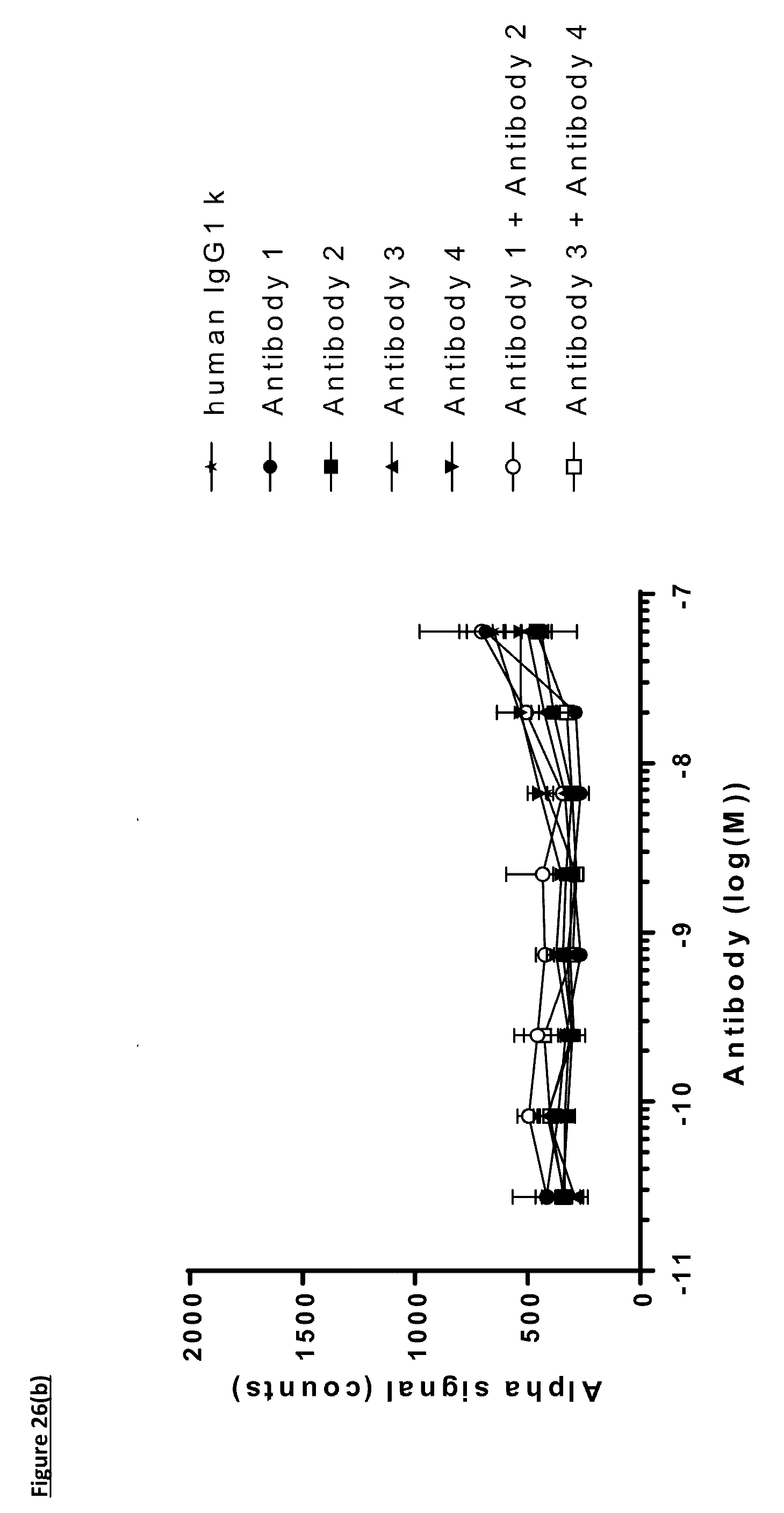

D00050
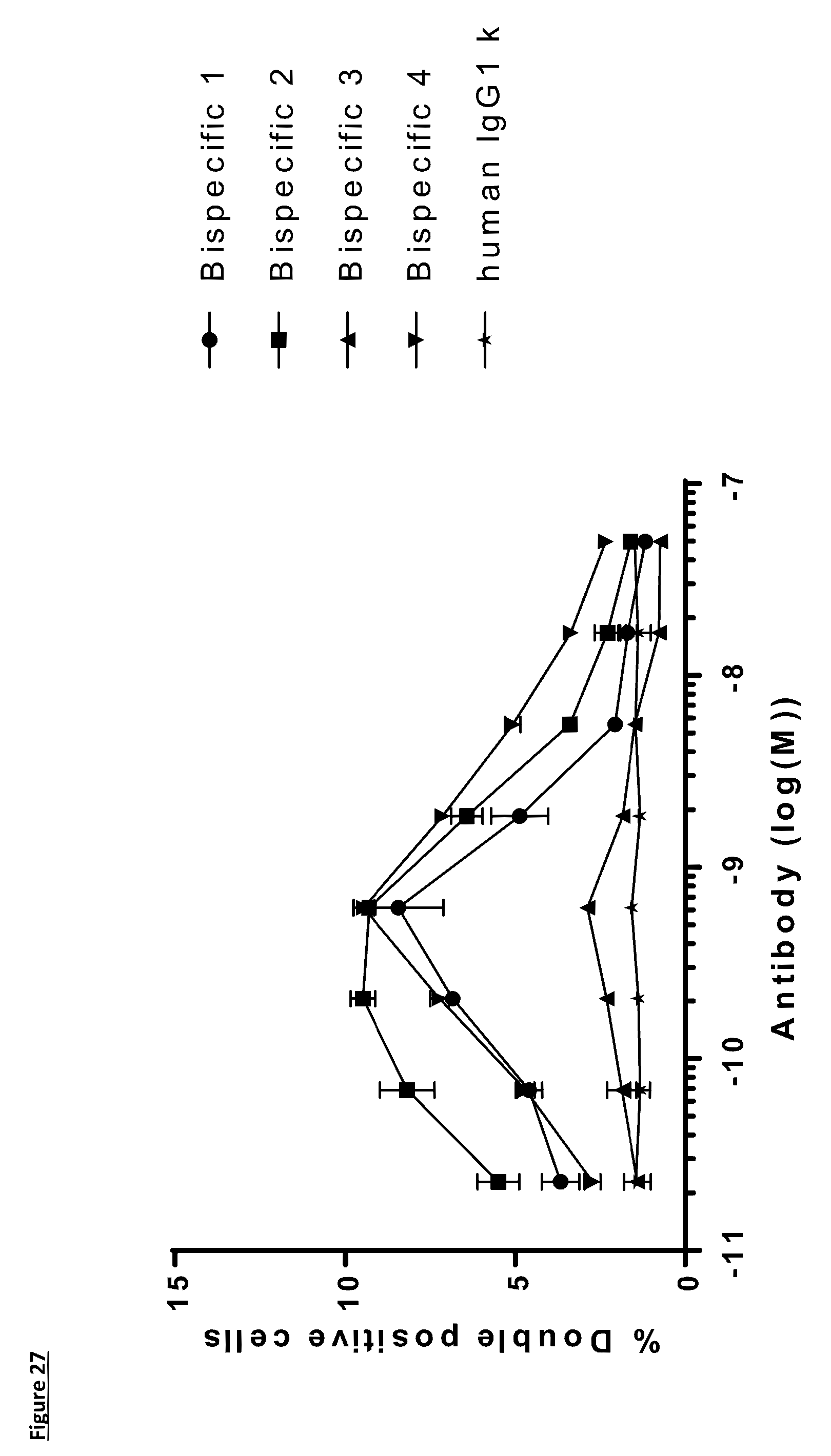
D00051
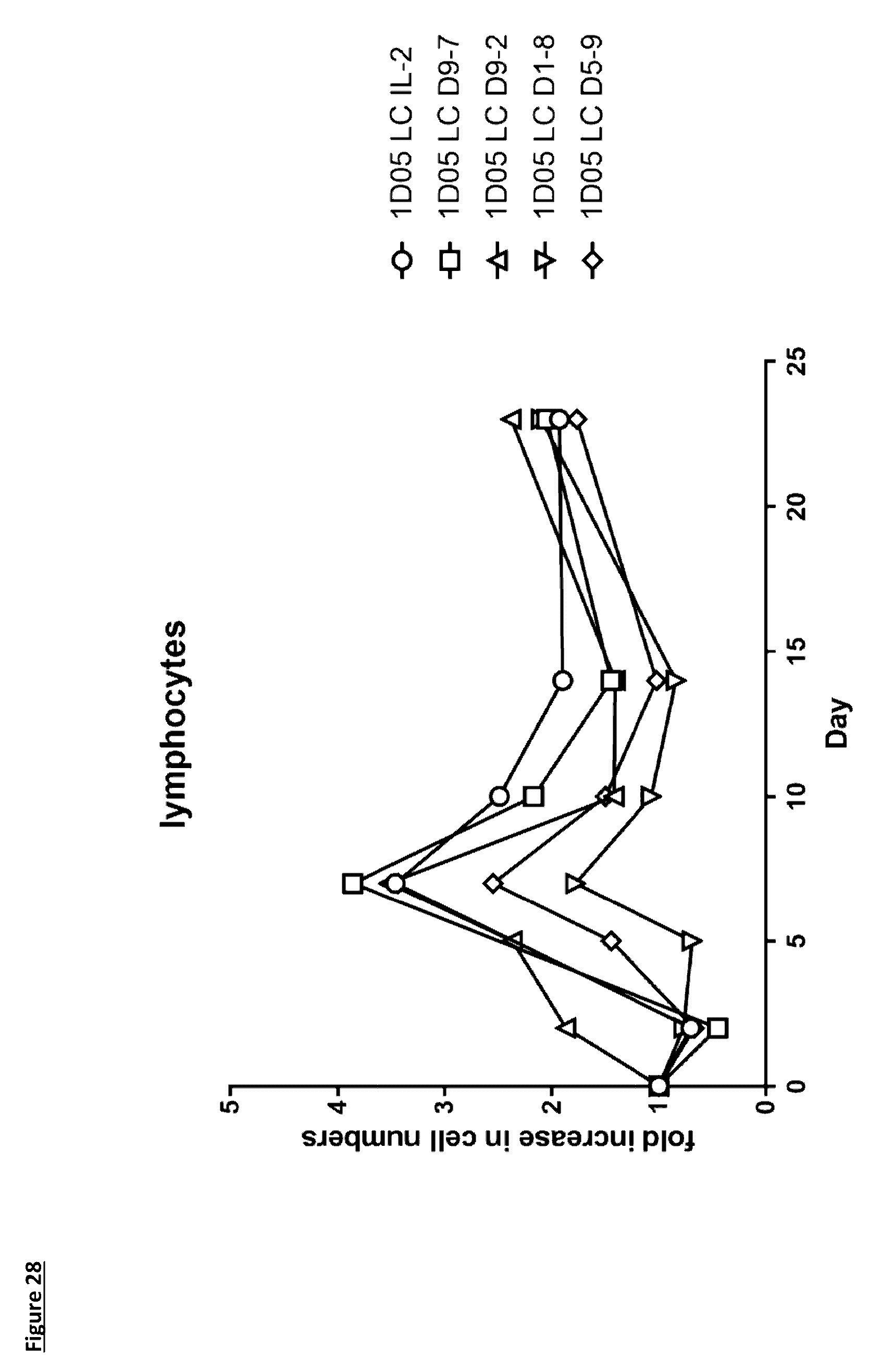
D00052
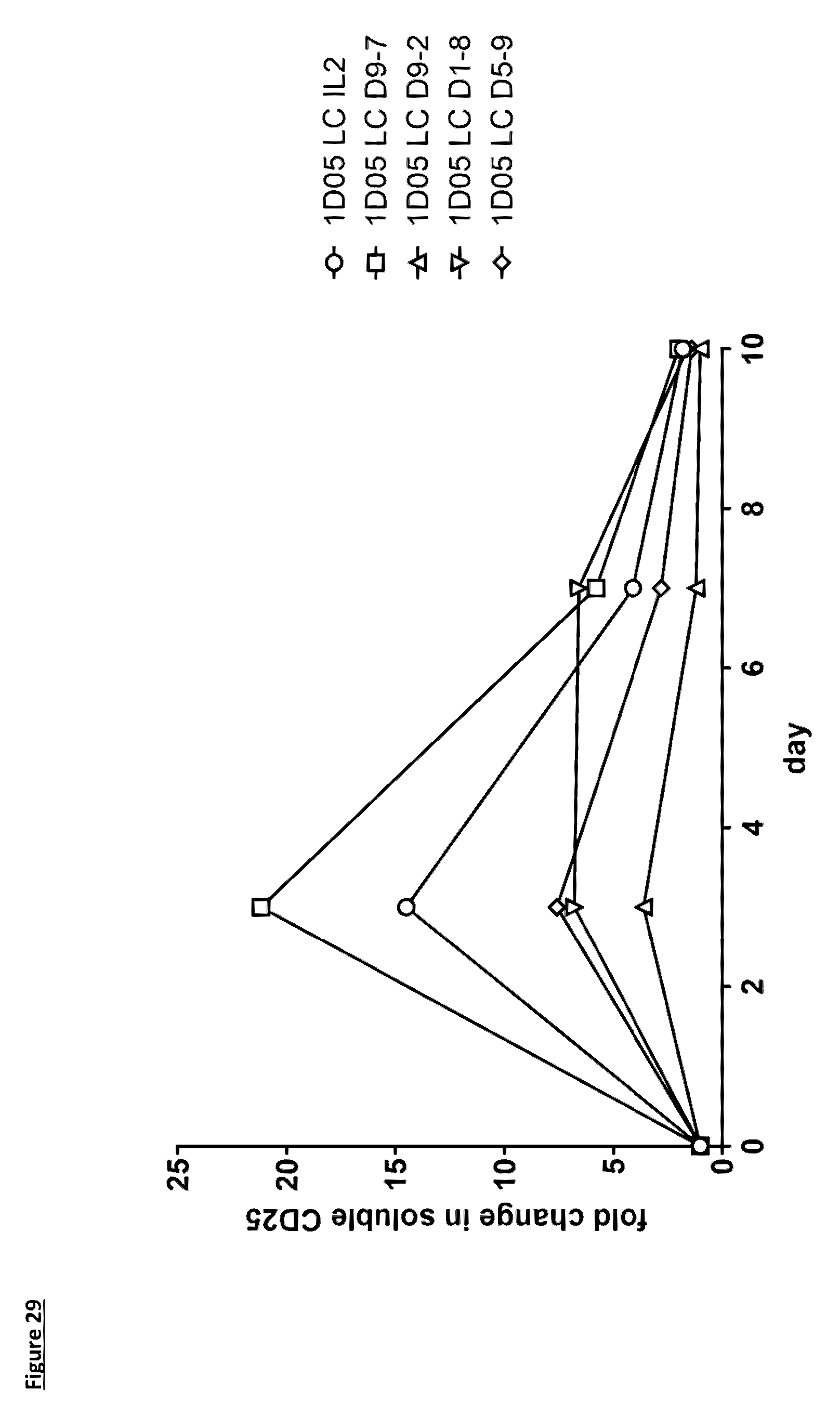
D00053
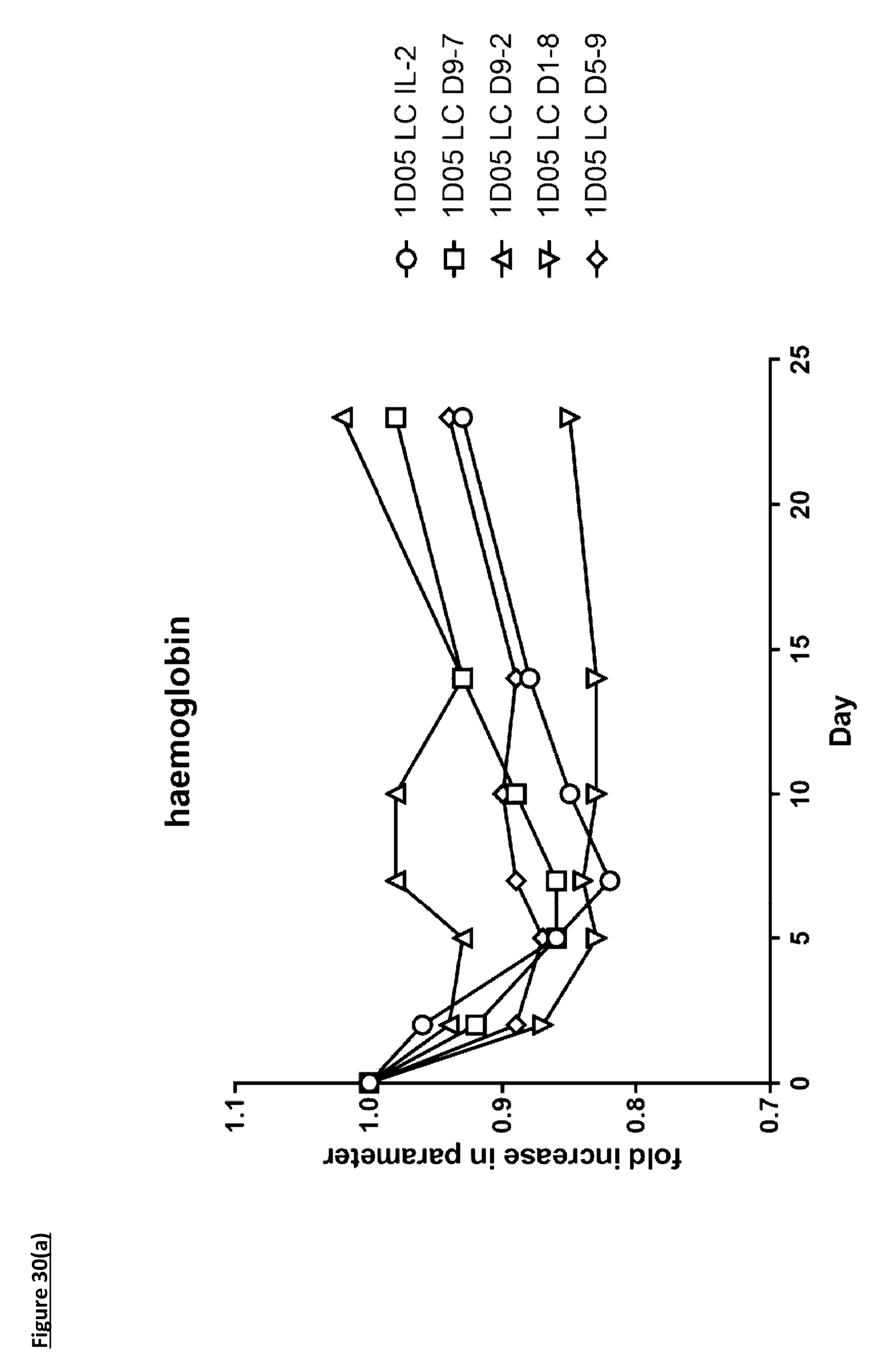
D00054
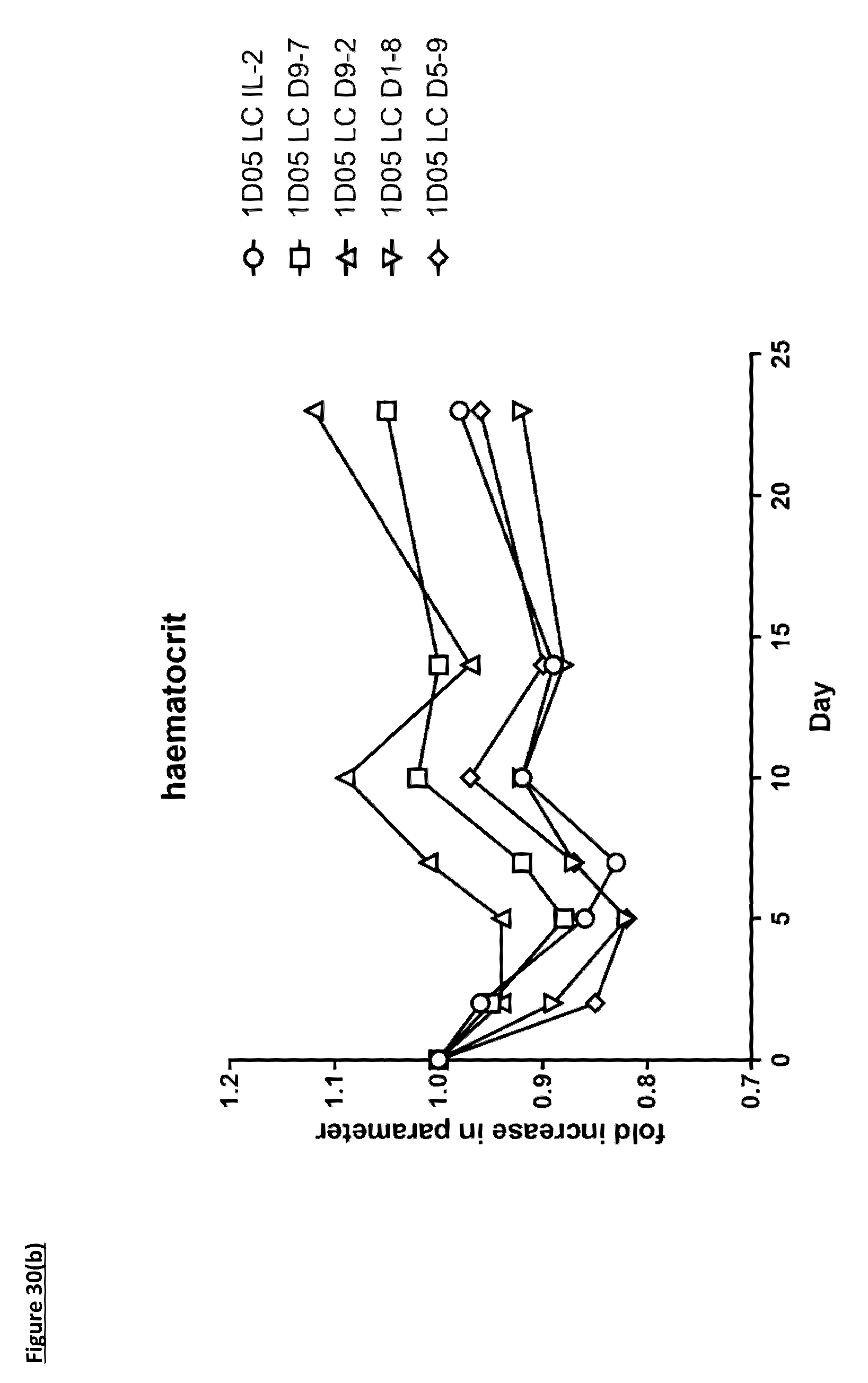
D00055
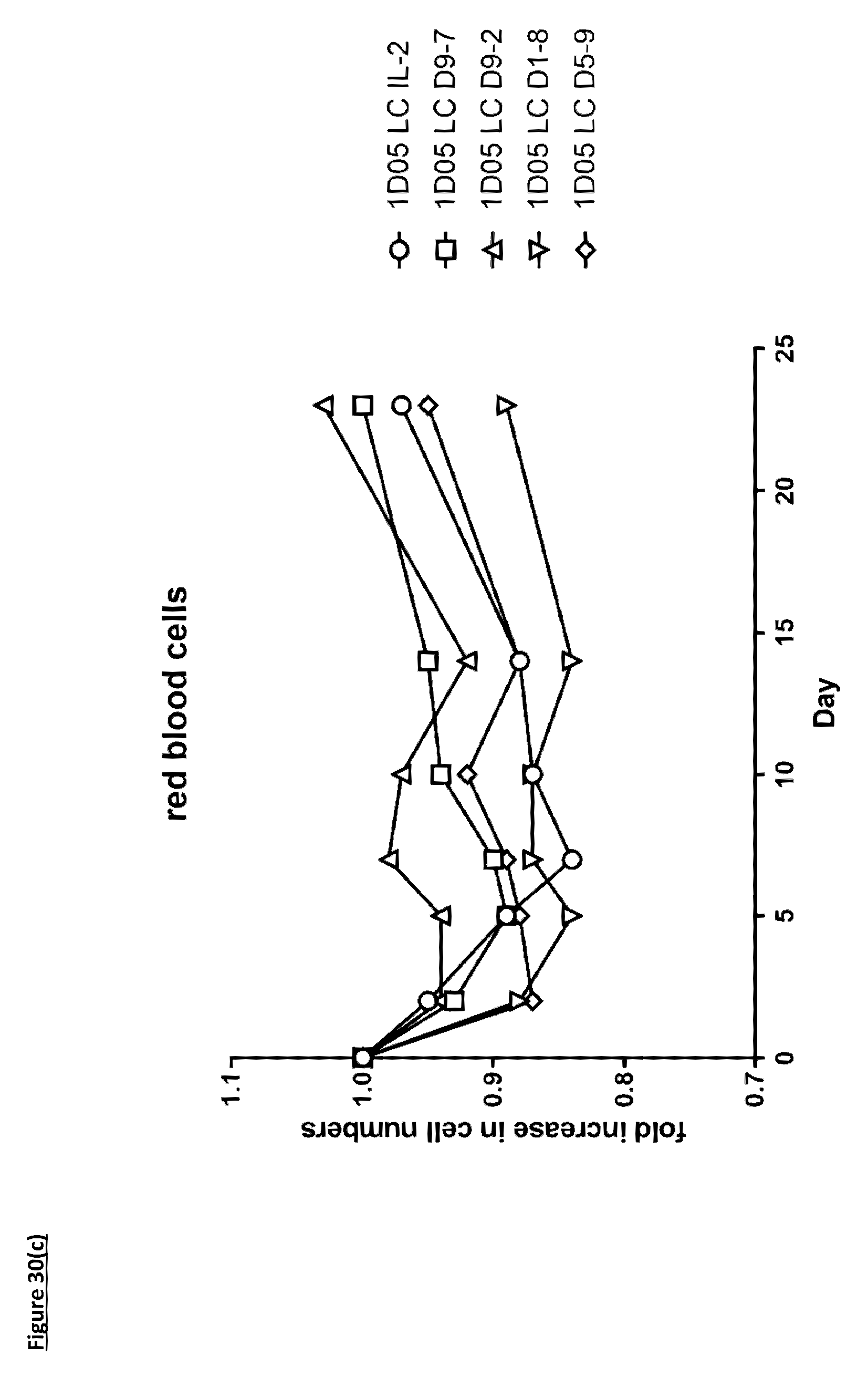
D00056
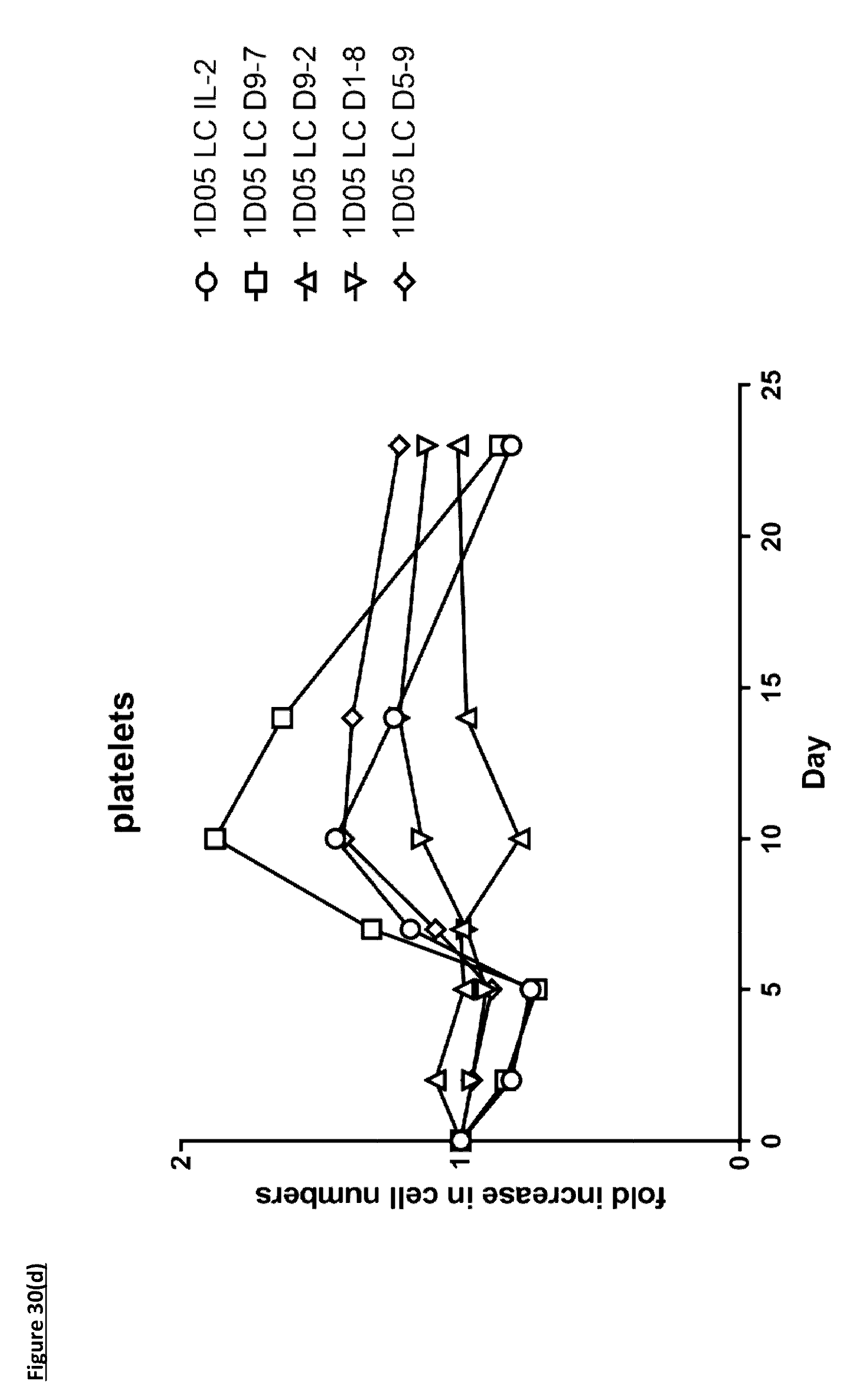
D00057
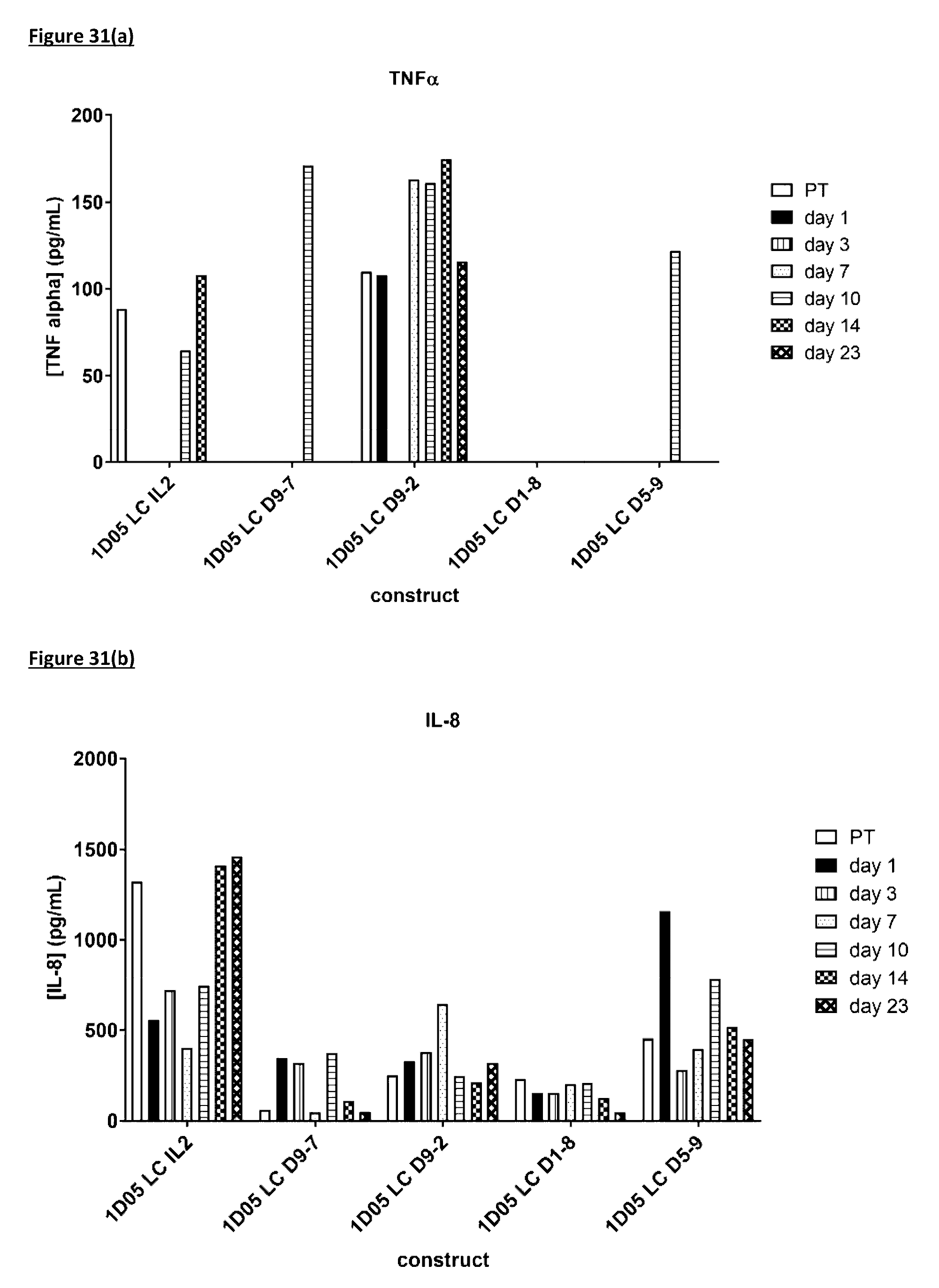
D00058
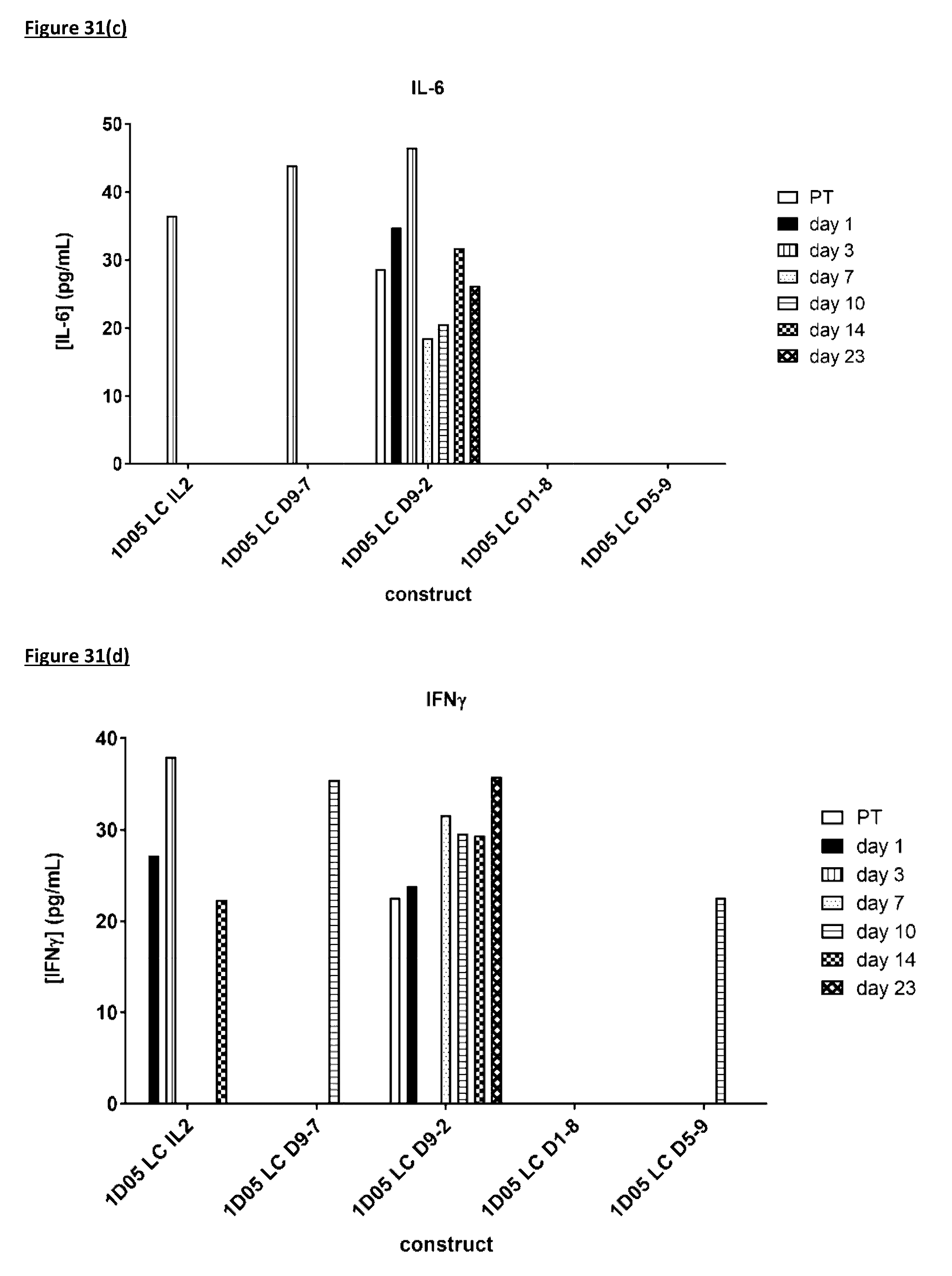
D00059
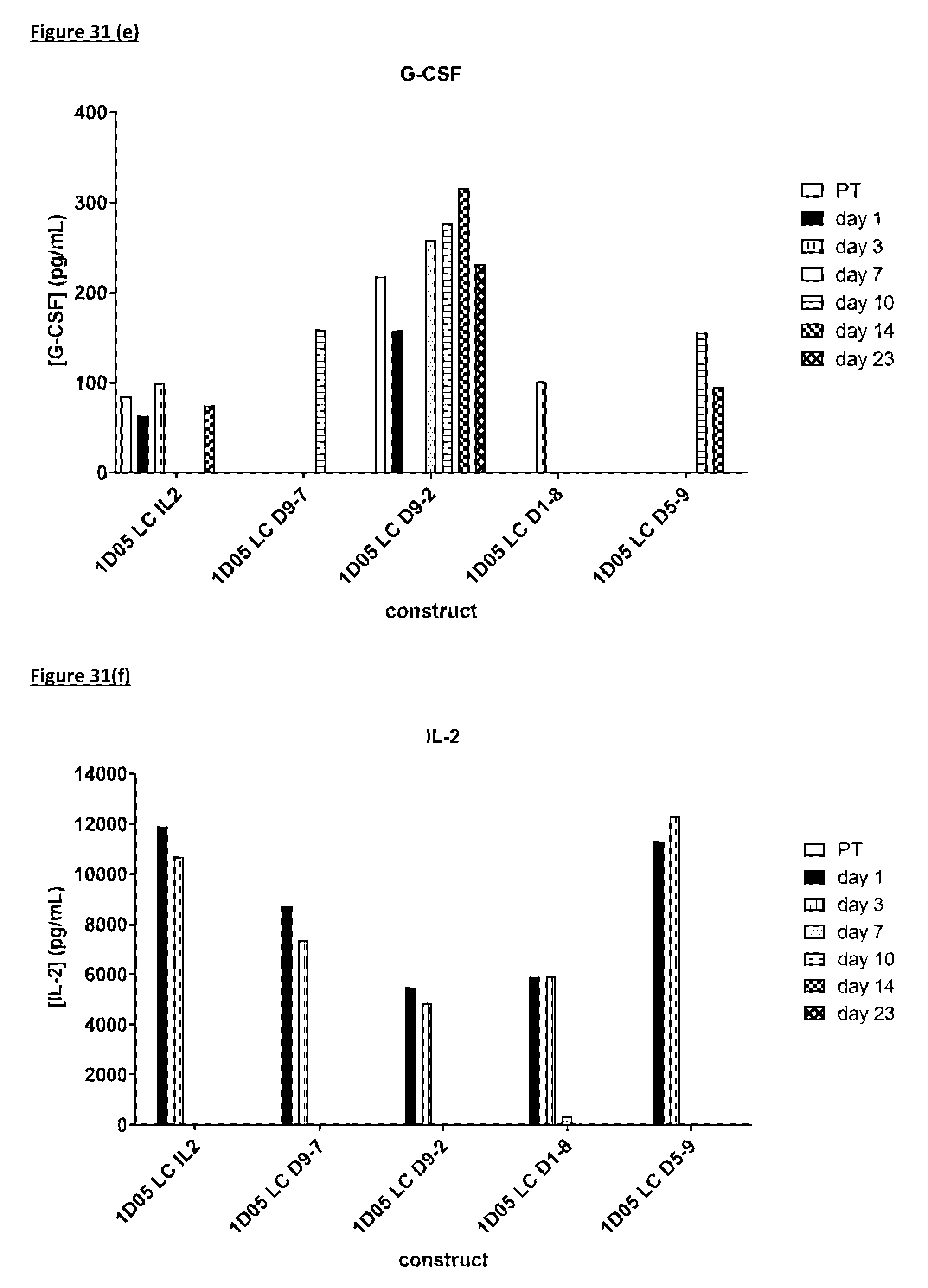
D00060
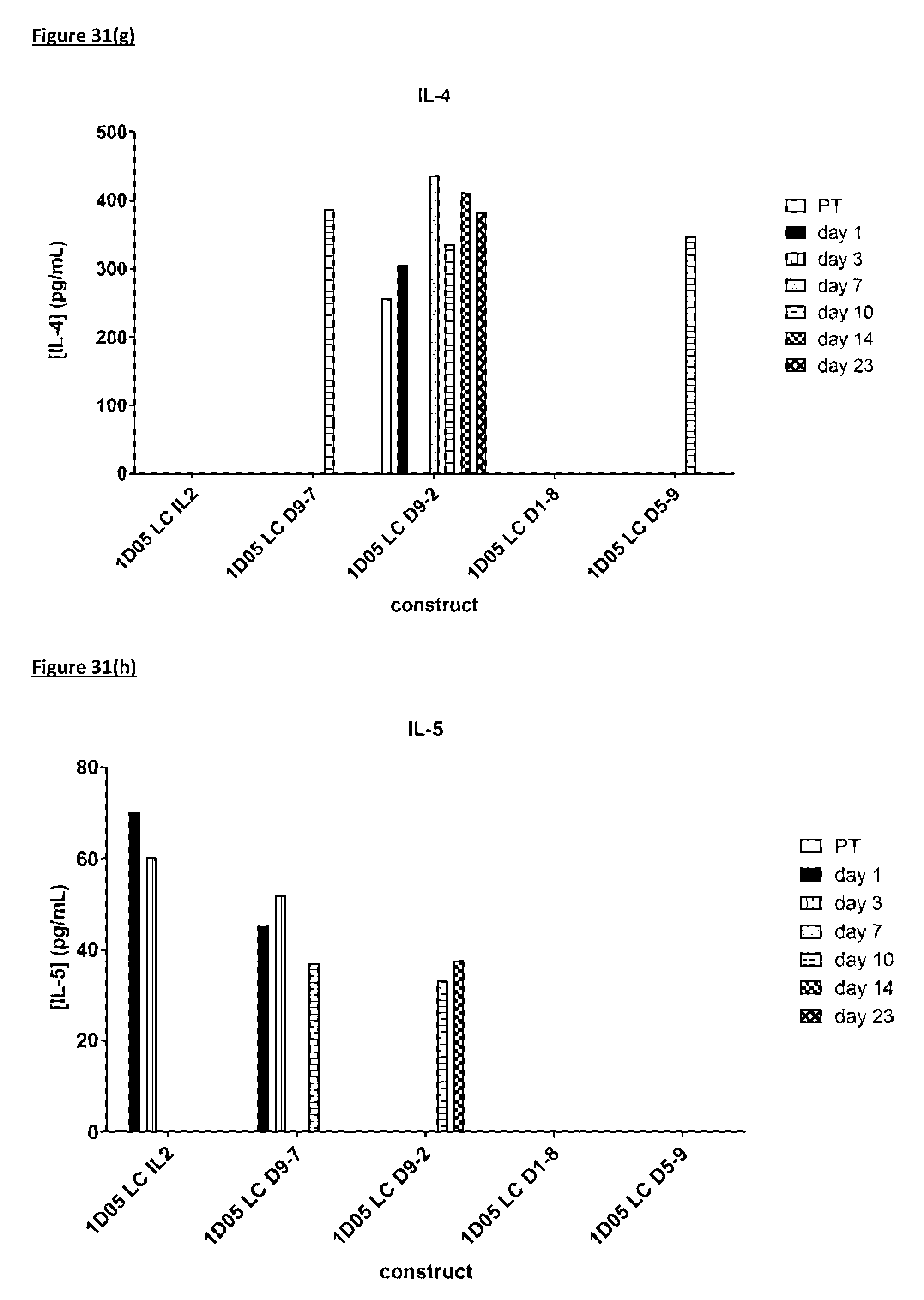
D00061
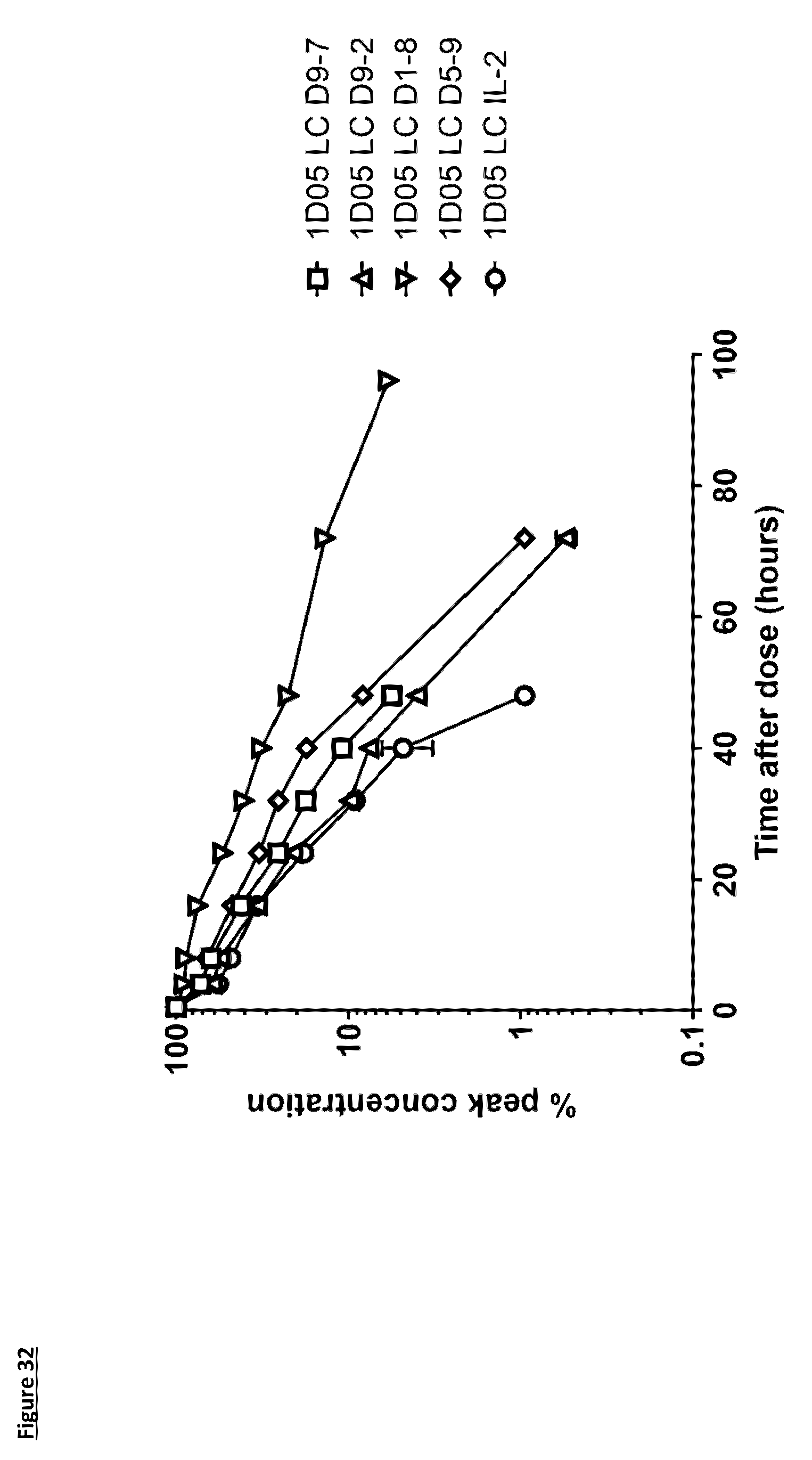
D00062
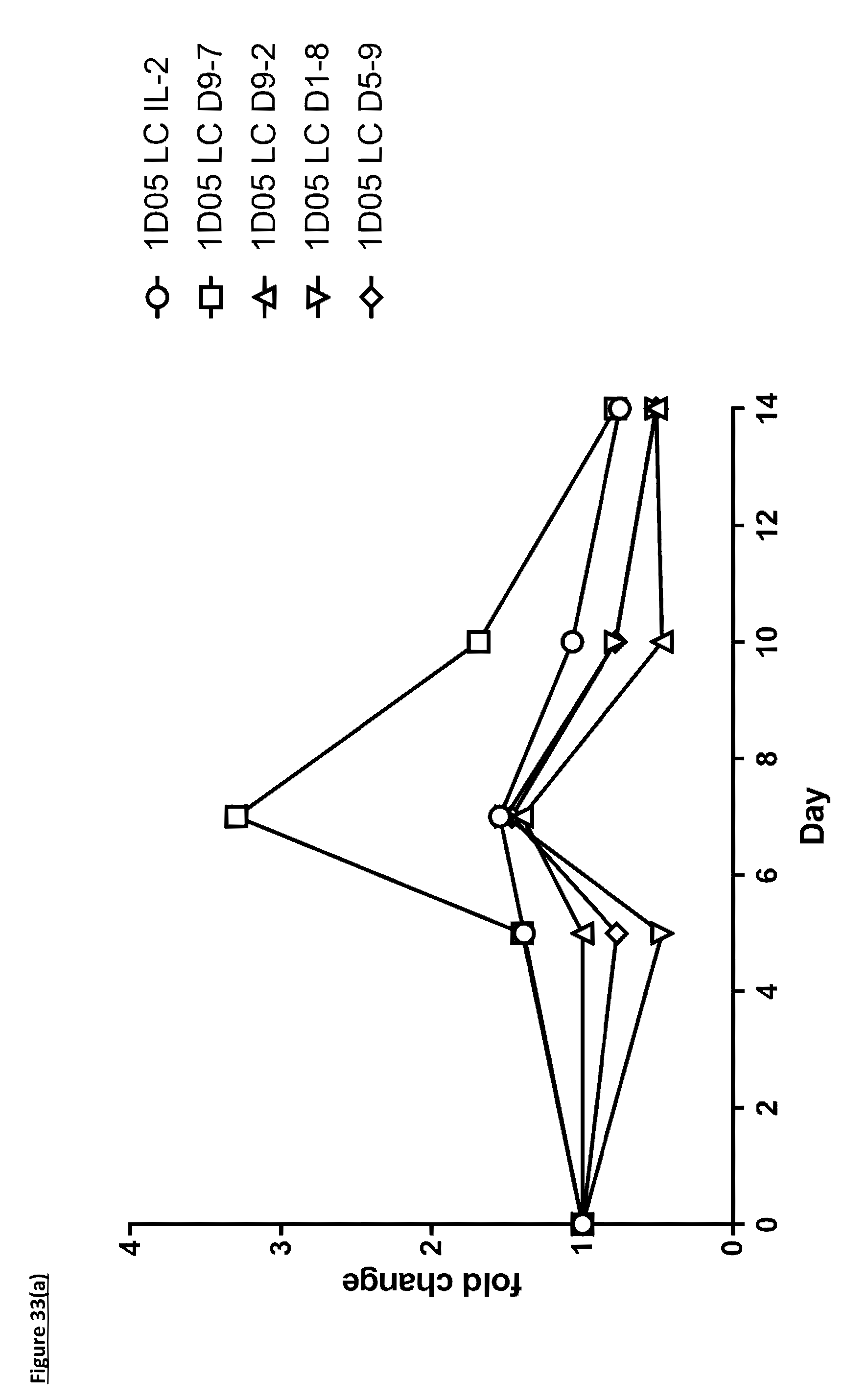
D00063
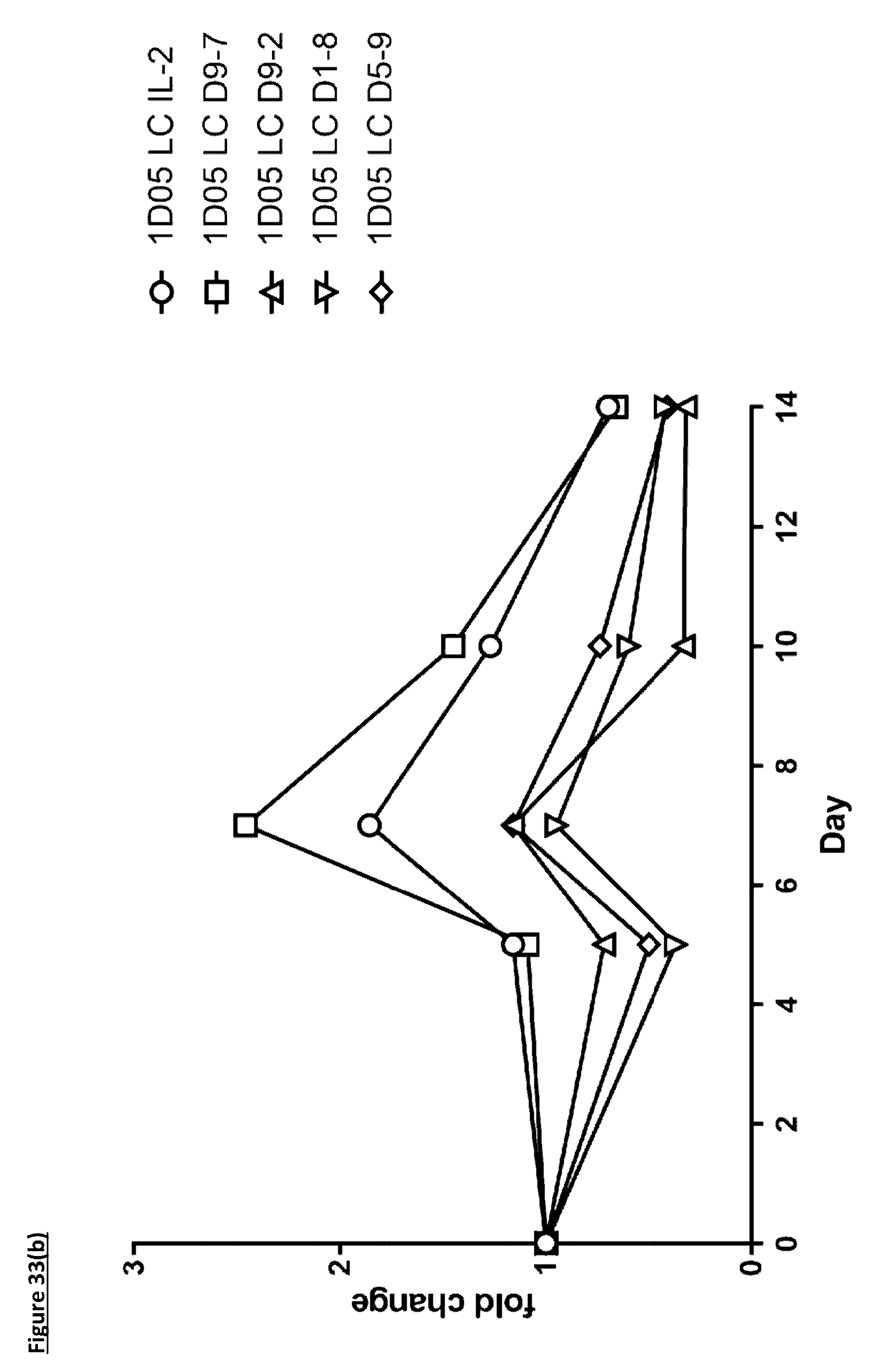
D00064
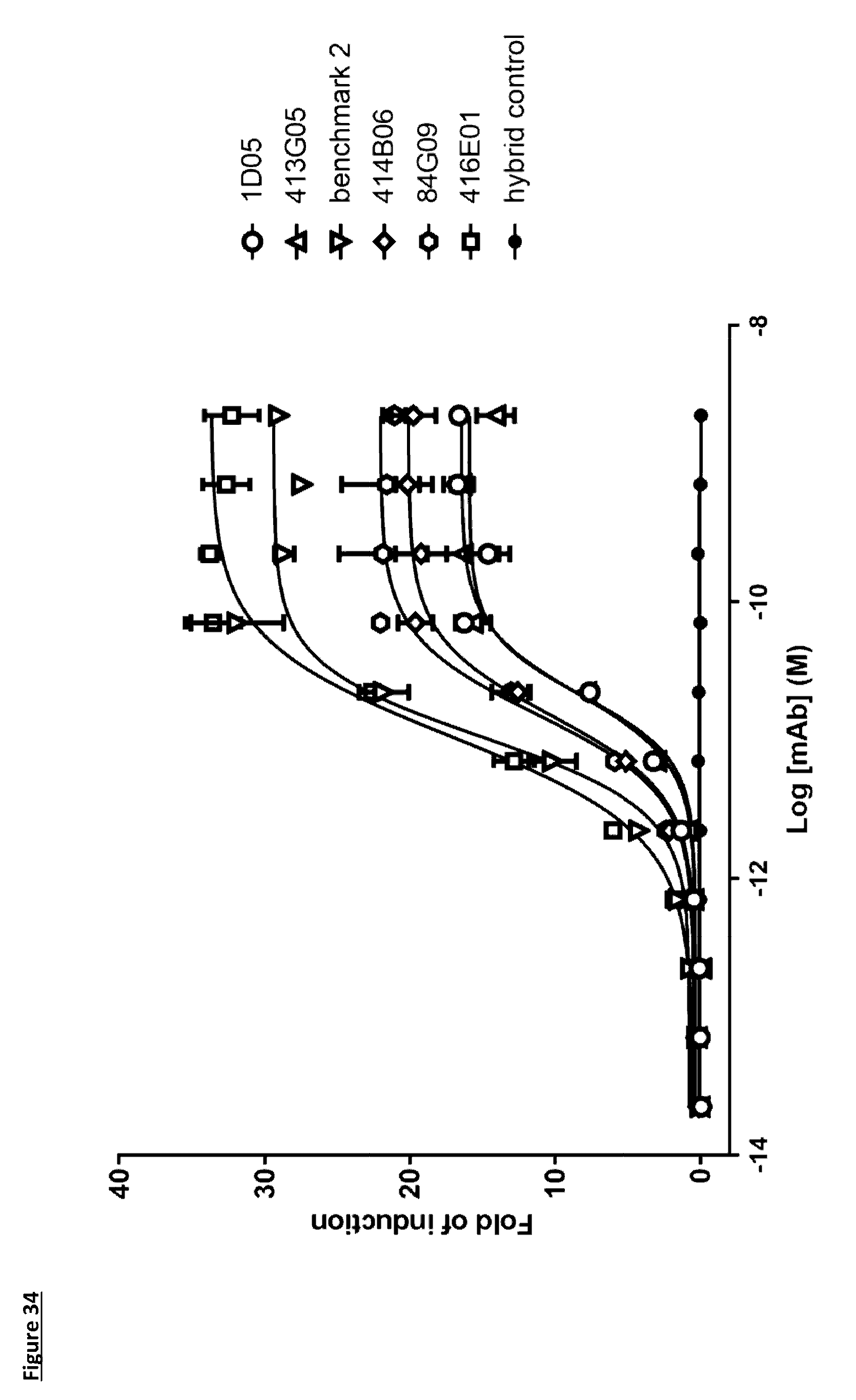
D00065
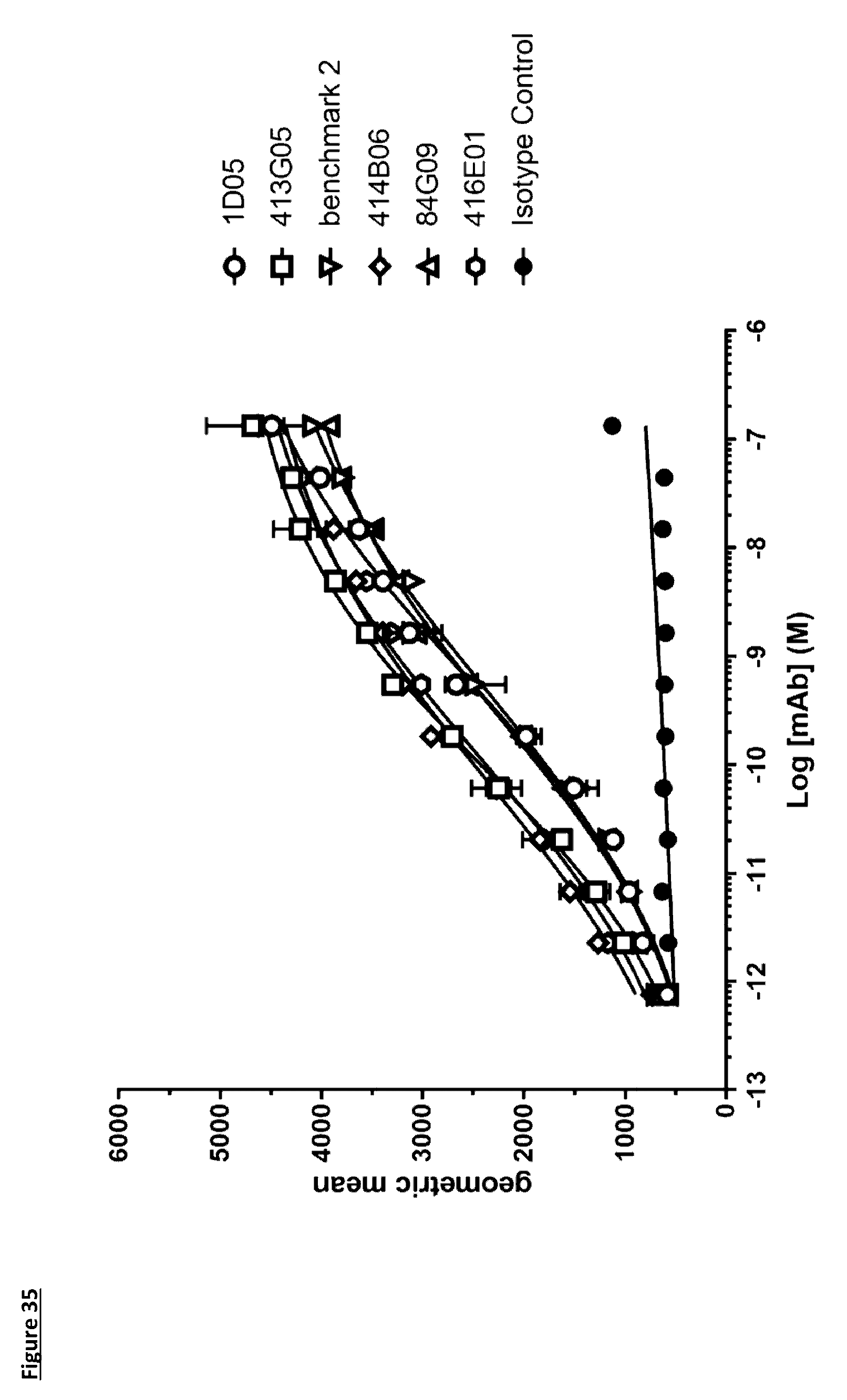
D00066
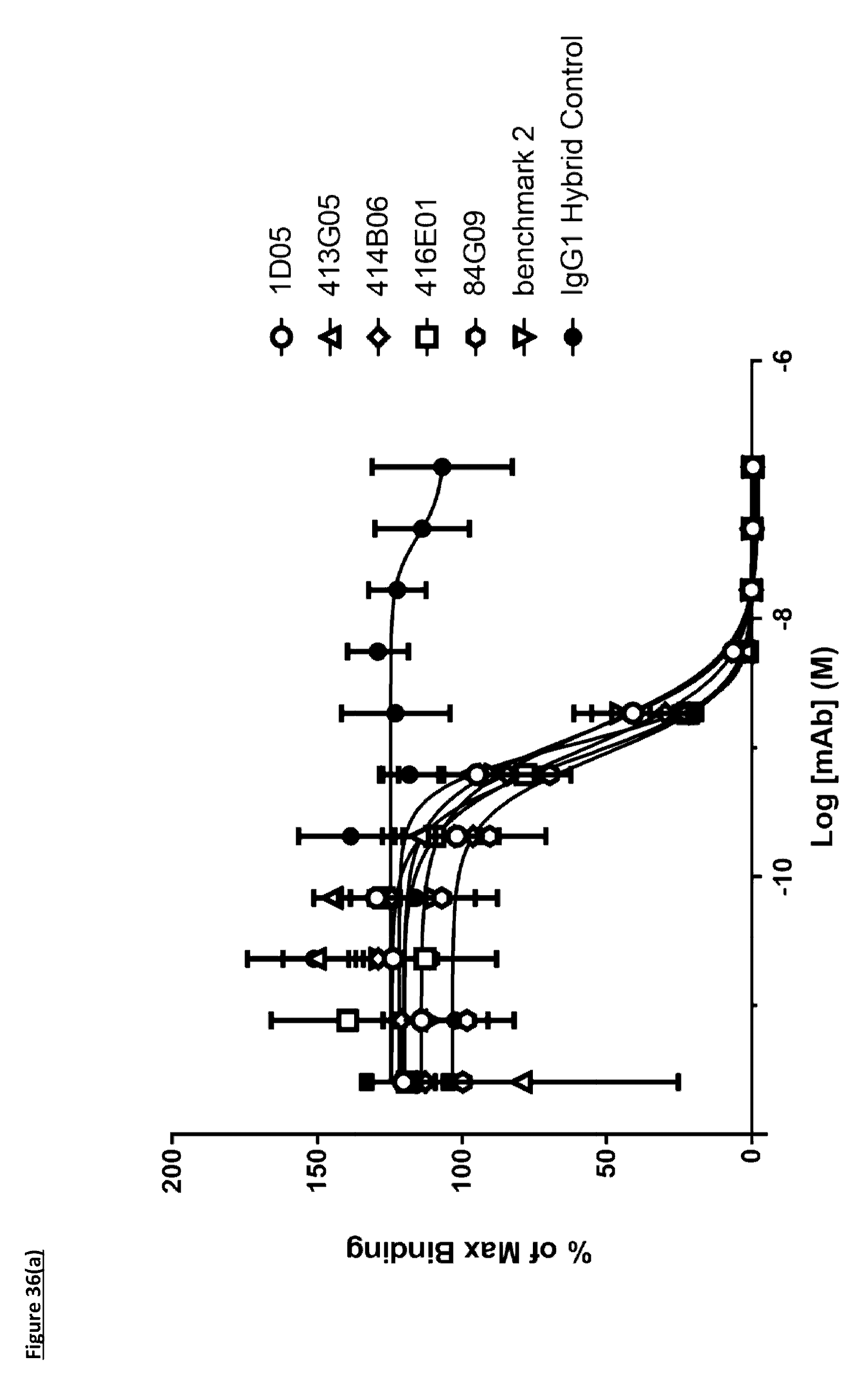
D00067
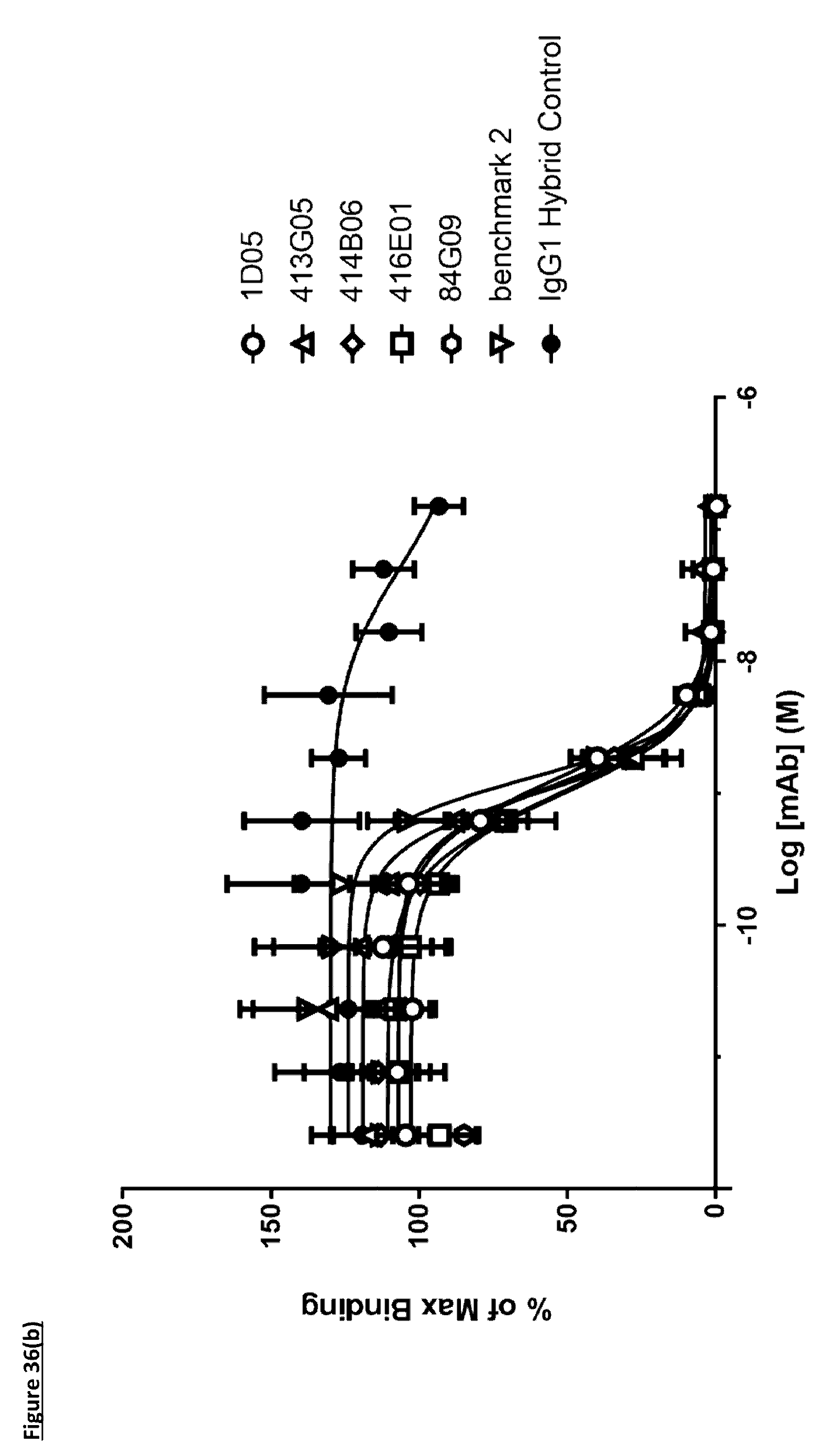
D00068
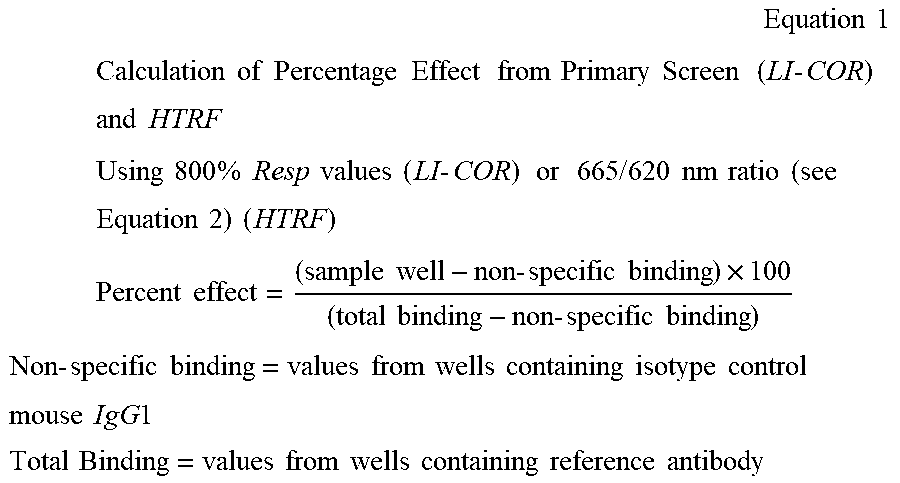
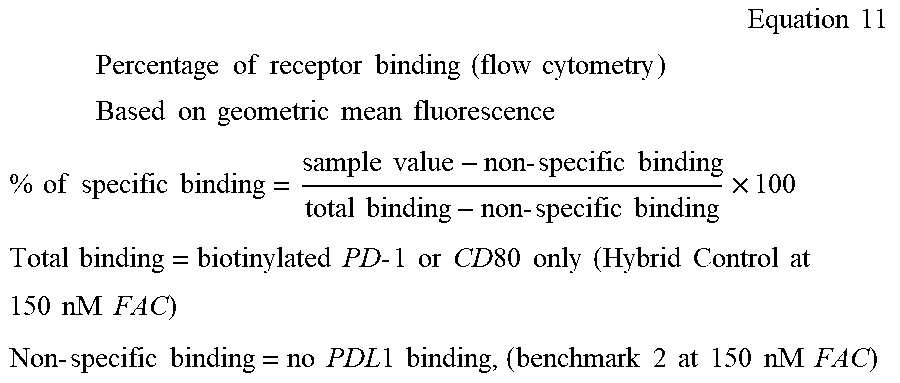
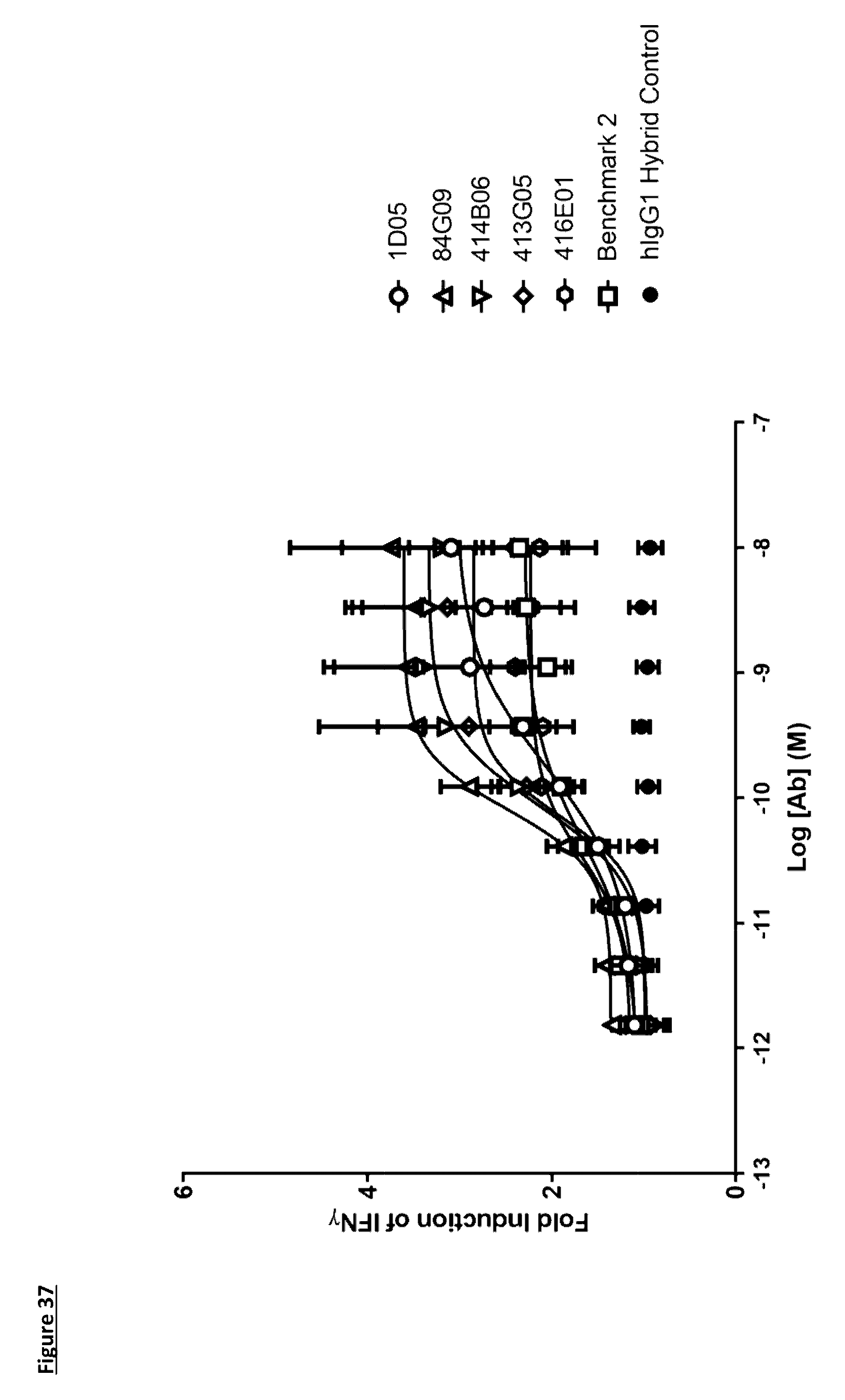



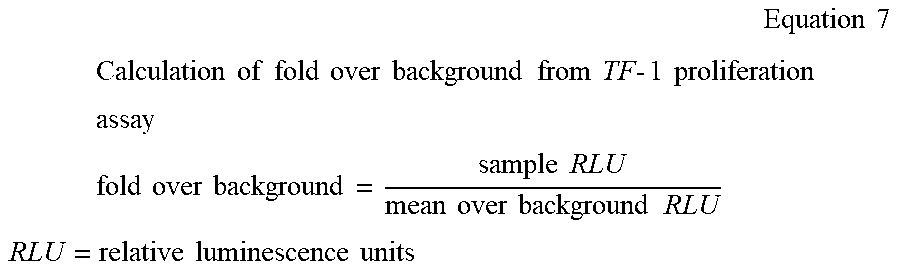
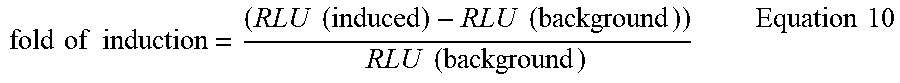

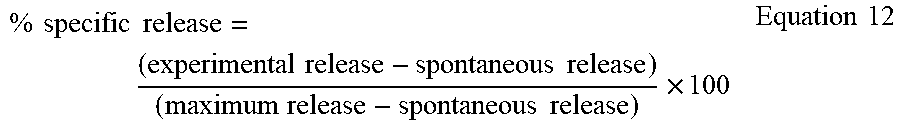
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.