Cellular Targeted Pharmaceutically Active Substance Or Label Delivery System
KROL; Magdalena ; et al.
U.S. patent application number 16/312726 was filed with the patent office on 2019-10-31 for cellular targeted pharmaceutically active substance or label delivery system. The applicant listed for this patent is CELLIS AG. Invention is credited to Paola BAIOCCO, Irene BENN, Alberto BOFFI, Lukasz KIRAGA, Magdalena KROL, Zofia PLICH, Tomasz RYGIEL, Aleksandra SZULC, Bartlomiej TACIAK, Katarzyna TONECKA, Katarzyna ULEWICZ.
| Application Number | 20190328911 16/312726 |
| Document ID | / |
| Family ID | 56235808 |
| Filed Date | 2019-10-31 |









View All Diagrams
| United States Patent Application | 20190328911 |
| Kind Code | A1 |
| KROL; Magdalena ; et al. | October 31, 2019 |
CELLULAR TARGETED PHARMACEUTICALLY ACTIVE SUBSTANCE OR LABEL DELIVERY SYSTEM
Abstract
The present invention relates to an isolated cellular targeted delivery system comprising a CD45+ leukocyte cell comprising within said cell a complex of one or more iron binding proteins and an active pharmaceutically active substance and/or label as well as methods for producing such isolated cellular targeted delivery system and uses of such system for prophylaxis, therapy, diagnosis or theragnosis, in particular for prophylactic or therapeutic vaccination, therapy of cancer, particularly metastatic cancer or inflammatory diseases.
| Inventors: | KROL; Magdalena; (Warsaw, PL) ; BENN; Irene; (Rome, IT) ; BAIOCCO; Paola; (Rome, IT) ; RYGIEL; Tomasz; (Warsaw, PL) ; BOFFI; Alberto; (Rome, IT) ; SZULC; Aleksandra; (Zurich, PL) ; PLICH; Zofia; (Zurich, PL) ; TONECKA; Katarzyna; (Zurich, PL) ; ULEWICZ; Katarzyna; (Zurich, PL) ; TACIAK; Bartlomiej; (Zurich, PL) ; KIRAGA; Lukasz; (Zurich, PL) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 56235808 | ||||||||||
| Appl. No.: | 16/312726 | ||||||||||
| Filed: | December 21, 2016 | ||||||||||
| PCT Filed: | December 21, 2016 | ||||||||||
| PCT NO: | PCT/PL2016/050057 | ||||||||||
| 371 Date: | December 21, 2018 |
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61P 13/10 20180101; C12N 2501/2313 20130101; C12N 2501/90 20130101; C12N 2501/21 20130101; A61K 38/42 20130101; A61P 11/00 20180101; A61K 45/06 20130101; A61K 47/64 20170801; C12N 5/0646 20130101; A61K 38/40 20130101; C12N 5/0636 20130101; A61P 31/00 20180101; A61P 35/00 20180101; A61K 35/17 20130101; C12N 2501/2304 20130101; A61K 51/1203 20130101; A61P 35/04 20180101; A61K 47/6901 20170801; A61K 47/6445 20170801; A61K 47/6811 20170801; A61P 9/10 20180101; A61P 15/00 20180101; C12N 2501/231 20130101; A61K 49/0056 20130101; A61K 49/1896 20130101; A61P 1/18 20180101; C12N 5/0642 20130101; A61P 27/02 20180101; A61P 29/00 20180101; C12N 5/0645 20130101; A61P 1/04 20180101; C12N 2501/15 20130101; A61K 51/0497 20130101; A61P 43/00 20180101; C12N 2501/999 20130101; A61K 35/15 20130101; A61P 37/02 20180101; A61K 47/644 20170801; A61P 17/02 20180101; C12N 2501/25 20130101; C12N 2501/24 20130101 |
| International Class: | A61K 51/12 20060101 A61K051/12; A61K 47/64 20060101 A61K047/64; A61K 35/15 20060101 A61K035/15; A61K 35/17 20060101 A61K035/17; A61K 51/04 20060101 A61K051/04; A61K 47/69 20060101 A61K047/69; A61K 45/06 20060101 A61K045/06; A61K 47/68 20060101 A61K047/68; A61K 49/18 20060101 A61K049/18; A61K 49/00 20060101 A61K049/00; C12N 5/0786 20060101 C12N005/0786; C12N 5/0783 20060101 C12N005/0783; C12N 5/0787 20060101 C12N005/0787 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Jun 22, 2016 | EP | PCT/EP2016/064483 |
| Jun 22, 2016 | EP | PCT/EP2016/064484 |
Claims
1. An isolated targeted delivery system comprising a CD45.sup.+ monocyte, CD45.sup.+ monocyte-macrophage, CD45.sup.+ lymphocyte and/or CD45.sup.+ granulocyte (commonly referred to as "CD45.sup.+ leukocyte cell") comprising within said cell a complex of one or more iron binding proteins and a pharmaceutically active substance, label or pharmaceutically active substance and label.
2. An isolated targeted delivery system comprising a CD45.sup.+ leukocyte cell comprising one or more label.
3. The isolated targeted delivery system according to claim 1, wherein the CD45.sup.+ leukocyte cell is producible from a CD34.sup.+ hematopoietic precursor cell.
4. The isolated targeted delivery system according to claim 3, wherein (i) the monocyte is a CD11b.sup.+ monocyte, but is not a dendritic cell which differentiation is controlled by following transcription factors: IFN-regulatory factor 8 (IRF8), nuclear factor interleukin (IL)-3-regulated protein (NFIL3), basic leucine zipper transcriptional factor ATF-like 3 (BATF3) or Transcription Factor RelB (NF-KB Subunit)-RELB, Spi 1 Proto-Oncogene (PU/1), recombining binding protein suppressor of hairless (RBPJ), IFN-regulatory factor 4 (IRF4) or transcription factor E2-2 (also known as (TCF4); (ii) the differentiated monocyte is selected from the group consisting of a macrophage, an activated macrophage, preferably a CD11b.sup.+ macrophage, more preferably a CD11b.sup.+ CD16.sup.+ macrophage, CD11b.sup.+ CD32.sup.+ macrophage, CD11b.sup.+ CD64.sup.+ macrophage, CD11b.sup.+ CD68.sup.+ macrophage, preferably a CD11b.sup.+ CD86.sup.+ M1 macrophage, preferably producing iNOS and/or secreting interleukin 12 (IL-12) or preferably CD11b.sup.+ CCR2.sup.+ M2 macrophage, CD11b.sup.+ CD204.sup.+ M2 macrophage, CD11b.sup.+ CD206.sup.+ M2 macrophage, CD11b.sup.+ CD204.sup.+ CD206.sup.+ M2 macrophage, CD11b.sup.+ Mayor Histocompatibility Complex II.sup.+ (MHCII.sup.+) (low or hi expression) M2 macrophage, CD11b.sup.+ CD200R.sup.+ M2 macrophage, CD11b.sup.+ CD163.sup.+ M2 macrophage or activated macrophage producing arginase and/or secreting interleukin 10 (IL-10); preferably the differentiated monocyte-macrophage is not a Lox1.sup.+, CXCR7.sup.+ and NRF2.sup.+ foam cell; (iii) monocyte-macrophage or activated monocyte-macrophage expressing of at least one chemokine receptor, or at least one growth factor receptor; (iv) the lymphocyte is selected from the group consisting of a CD3.sup.+ and CD4.sup.+ or CD8.sup.+ T lymphocyte, or a CD19.sup.+, CD20.sup.+, CD21.sup.+, CD19.sup.+ CD20.sup.+, CD19.sup.+ CD21.sup.+, CD20.sup.+ CD21.sup.+, or CD19.sup.+ CD20.sup.+ CD21.sup.+ B lymphocyte, and a natural killer (NK) cell; or (v) the granulocyte is selected from the group consisting of a neutrophil, an eosinophil and a basophil.
5. The isolated targeted delivery system of claim 4, wherein the activated macrophage: (i) is producible by in vitro incubation of a monocyte or macrophage with a factor capable of altering expression markers on macrophages; (ii) is characterized by expression of at least one of following antigens: CD64, CD86, CD16, CD32, high expression of WICK and/or production of iNOS and/or IL-12; (iii) is producible by in vitro incubation of a monocyte or macrophage with a factor capable of inducing the ability of the macrophage to phagocytose; (iv) is characterized by expression of at least one of following antigens: CD204, CD206, CD200R; CCR2, transferrin receptor (TfR), CXC-motive chemokine receptor 4 (CXCR4), CD163, and/or T cell immunoglobulin-domain and mucin-domain 2 (TIM-2), and/or show low expression of MHCII; (v) has the ability to phagocytose; and/or (vi) is capable of cytokine secretion, or production of inducible nitric oxide synthetase (iNOS) or other pro-inflammatory compounds, arginase or other immunosuppressive/anti-inflammatory compounds.
6. The targeted delivery system according to claim 5, wherein: (i) the M1 inducer is selected from the group consisting of LPS, INF-.gamma., and viral and bacterial infection; or (ii) the M2 inducer is selected from the group consisting of IL-4, IL-10, IL-13, immune complex of an antigen and antibody, IgG, heat activated gamma-globulin, glucocorticosteroid, TGF-.beta., IL-1R, CCL-2, IL-6, M-CSF, PPAR.gamma. agonist, Leukocyte inhibitory factor, adenosine, helminth and fungal infection.
7. The isolated targeted delivery system of claim 4, wherein the monocyte-macrophage cell: (i) is producible from a CD34.sup.+ hematopoietic precursor cell; (ii) is producible by in vitro incubation of monocytes/monocyte-macrophage with at least one inducer; (iii) is characterized by expression of at least one of the following antigens: TfR.sup.+, CD163.sup.+, TIM-2.sup.+, CD14.sup.+, CD16.sup.+, CD33.sup.+, and/or CD115.sup.+; (iv) is characterized by expression of at least one of the following antigens: TfR.sup.+, CD163.sup.+, TIM-2.sup.+, CXCR4.sup.+, CD14.sup.+, and/or CD16.sup.+; and/or (v) has the ability to phagocytose.
8. The targeted delivery system according to claim 7, wherein: (i) the M1 inducer is selected from the group consisting of LPS, INF-.gamma., or viral or bacterial infection; (ii) the M2 inducer is selected from the group consisting of IL-4, IL-10, IL-13, immune complex of an antigen and antibody, IgG, heat activated gamma-globulins, Glucocorticosteroids, TGF-.beta., IL-1R, CCL-2, IL-6, M-CSF, PPAR.gamma. agonist, Leukocyte inhibitory factor, cancer-conditioned medium, cancer cells, adenosine and helminth or fungal infection.
9. The isolated targeted delivery system of claim 4, wherein the lymphocyte: (i) is obtainable from blood, spleen, or bone marrow or is producible from a CD34.sup.+ precursor cell; (ii) is an immunologically competent lymphocyte; (iii) expresses antigen specific T cell receptors; and/or (iv) is characterized by expression of at least one of the following antigens: (a) CD3.sup.+ and CD4.sup.+ or CD8.sup.+ or (b): CD19.sup.+, CD20.sup.+, CD21.sup.+, CD19.sup.+ CD20.sup.+, CD19.sup.+ CD21.sup.+, CD20.sup.+ CD21.sup.+, or CD19.sup.+ CD20.sup.+ CD21.sup.+ antigen.
10. The isolated targeted delivery system of claim 4, wherein the granulocyte: (i) is obtainable from blood, spleen or bone marrow or producible from a CD34.sup.+ precursor cell; (ii) is characterized by expression of at least one of the following CD66b.sup.+ and/or CD193.sup.+; (iii) is a polymorphonuclear leukocyte characterized by the presence of granules in their cytoplasm; and/or (iv) is characterized by expression of at least one of the following: TfR.sup.+, CD163.sup.+, TIM-2.sup.+, and/or CXCR4.sup.+.
11. The isolated targeted delivery system of claim 4, wherein the NK cell: (i) is obtainable from blood, spleen or bone marrow or producible from a CD34.sup.+ precursor cell; and/or (ii) is characterized by the lack of CD3 expression and expression of at least one of the following CD56.sup.+ and/or CD94.sup.+, CD158a.sup.+ CD158f.sup.+ CD314.sup.+ CD335.sup.+.
12. The isolated targeted delivery system of claim 1, wherein the iron binding protein is selected from the group consisting of ferritin, haemoglobin, haemoglobin-haptoglobin complex, hemopexin, transferrin, and lactoferrin.
13. The isolated targeted delivery system according to claim 1, wherein the pharmaceutically active substance is selected from the group consisting of a protein, a nucleic acid, a non-protein non-nucleic acid compound with a molecular weight of less than 1.5 kD, an anti-arteriosclerotic drug, an anti-inflammatory drug, a photosensitizing compound, a virus, and a .alpha. or radiation emitting radioisotope, which also emit a cell damaging amount of .gamma. radiation or a cell damaging amount of .alpha. radiation, or a complex of the compound or isotope linked to a nanoparticle.
14. The isolated targeted delivery system according to claim 13, wherein the pharmaceutically active substance is an anticancer drug, wherein the anticancer drug is selected from the group consisting of an apoptosis-inducing drug, an alkylating substance, anti-metabolites, antibiotics, epothilones, nuclear receptor agonists and antagonists, an anti-androgene, an anti-estrogen, a platinum compound, a hormone, a antihormone, an interferon, an inhibitor of cell cycle-dependent protein kinases (CDKs), an inhibitor of cyclooxygenases and/or lipoxygenases, a biogeneic fatty acid, a biogenic fatty acid derivative, including prostanoids and leukotrienes, an inhibitor of protein kinases, an inhibitor of protein phosphatases, an inhibitor of lipid kinases, a platinum coordination complex, an ethyleneimine, a methylmelamine, a triazine, a vinca alkaloid, a pyrimidine analog, a purine analog, an alkylsulfonate, a folic acid analog, an anthracendione, a substituted urea, and a methylhydrazin derivative, an ene-diyne antibiotic, a maytansinoid an auristatine derivate, immune check-point inhibitor, and an inhibitor of tumour-specific protein or marker, preferably a Rho-GDP-dissociation inhibitor, more preferably Grp94 or AXL inhibitor.
15. The isolated targeted delivery system according to claim 13, wherein the pharmaceutically active substance is an anticancer drug, wherein the anticancer drug is selected from the group consisting of a acediasulfone, aclarubicine, ambazone, aminoglutethimide, L-asparaginase, azathioprine, banoxantrone, bendamustine, bleomycin, busulfan, calcium folinate, carboplatin, carpecitabine, carmustine, celecoxib, chlorambucil, cis-platin, cladribine, cyclophosphamide, cytarabine, dacarbazine, dactinomycin dapsone, daunorubicin, dibrompropamidine, diethylstilbestrole, docetaxel, doxorubicin, enediynes, epirubicin, epothilone B, epothilone D, estramucin phosphate, estrogen, ethinylestradiole, etoposide, flavopiridol, floxuridine, fludarabine, fluorouracil, fluoxymesterone, flutamide fosfestrol, furazolidone, gemcitabine, gonadotropin releasing hormone analog, hexamethylmelamine, hydroxycarbamide, hydroxymethylnitrofurantoin, hydroxyprogesteronecaproat, hydroxyurea, idarubicin, idoxuridine, ifosfamide, interferon .alpha., irinotecan, leuprolide, lomustine, lurtotecan, mafenide sulfate olamide, mechlorethamine, medroxyprogesterone acetate, megastrolacetate, melphalan, mepacrine, mercaptopurine, methotrexate, metronidazole, mitomycin C, mitopodozide, mitotane, mitoxantrone, mithramycin, nalidixic acid, nifuratel, nifuroxazide, nifuralazine, nifurtimox, nimustine, ninorazole, nitrofurantoin, nitrogen mustards, oleomucin, oxolinic acid, pentamidine, pentostatin, phenazopyridine, phthalyl sulfathiazole, pipobroman, prednimustine, prednisone, preussin, procarbazine, pyrimethamine, raltitrexed, rapamycin, rofecoxib, rosiglitazone, salazosulfapyridine, scriflavinium chloride, semustine streptozocine, sulfacarbamide, sulfacetamide, sulfachlopyridazine, sulfadiazine, sulfadicramide, sulfadimethoxine, sulfaethidole, sulfafurazole, sulfaguanidine, sulfaguanole, sulfamethizole, sulfamethoxazole, co-trimoxazole, sulfamethoxydiazine, sulfamethoxypyridazine, sulfamoxole, sulfanilamide, sulfaperin, sulfaphenazole, sulfathiazole, sulfisomidine, staurosporin, tamoxifen, taxol, teniposide, tertiposide, testolactone, testosteronpropionate, thioguanine, thiotepa, tinidazole, topotecan, triaziquone, treosulfan, trimethoprim, trofosfamide, UCN-01, vinblastine, vincristine, vindesine, vinblastine, vinorelbine, and zorubicin, preferably selected from the group consisting of auristatin, banoxantrone, bendamustine, chlorambucil, chaliceamycin, dynemycin A, maytansine, melphalan, mertansine, and neocazinostatin.
16. The isolated targeted delivery system according to claim 13, wherein the immunomodulatory drugs activate or inhibit activity of immune cells, preferably the immunomodulatory drugs are ligands or antagonists of Pattern Recognition Receptors, particularly Toll-like Receptors, NOD-like receptors (NLR), RIG-I-like receptors (RLR).
17. The isolated targeted delivery system according to claim 13, wherein the pharmaceutically active substance is an anticancer drug, wherein the anticancer drug is a proliferation inhibiting protein, or an antibody or antibody like binding protein that specifically binds to a proliferation promoting protein or a nucleic acid.
18. The isolated targeted delivery system according to claim 1 comprising a pharmaceutically active substance, wherein the pharmaceutically active substance is a hypoxia-activated prodrug.
19. The isolated targeted delivery system according to claim 1 comprising a pharmaceutically active substance, wherein the pharmaceutically active substance is an antigen or a nucleic acid encoding an antigen.
20. The isolated targeted delivery system of claim 1 comprising a label, wherein the label is selected from the group consisting of a fluorescent dye, a fluorescence emitting isotope, a radioisotope, a detectable polypeptide or nucleic acid encoding a detectable polypeptide and a contrast agent.
21. The isolated targeted delivery system of claim 1 comprising a label, wherein the label comprises a chelating agent which forms a complex with divalent or trivalent metal cations.
22. The isolated targeted delivery system of claim 21, wherein the chelating agent is selected from the group consisting of 1,4,7,10-tetraazacyclododecane-N,N',N,N'-tetraacetic acid (DOTA), ethylenediaminetetraacetic acid (EDTA), 1,4,7-triazacyclononane-1,4,7-triacetic acid (NOTA), triethylenetetramine (TETA), iminodiacetic acid, Diethylenetriamine-N,N,N',N',N''-pentaacetic acid (DTPA) and 6-Hydrazinopyridine-3-carboxylic acid (HYNIC).
23. The isolated targeted delivery system of claim 20, wherein the label comprises a contrast agent, wherein the contrast agent comprises a paramagnetic agent.
24. The isolated targeted delivery system of claim 20, wherein the label comprises a radioisotope/fluorescence emitting isotope, wherein the radioisotope/fluorescence emitting isotope is selected from the group consisting of alpha radiation emitting isotopes, gamma radiation emitting isotopes, Auger electron emitting isotopes, X-ray emitting isotopes, fluorescent isotopes, fluorescence emitting isotopes, as well as conjugates and combinations of the above with proteins, peptides, small molecular inhibitors, antibodies or other compounds.
25. The isolated targeted delivery system of claim 20, wherein the label comprises a fluorescent dye, wherein the fluorescence dye is selected from the group consisting of the following classes of fluorescent dyes: Xanthens, Acridines, Oxazines, Cynines, Styryl dyes, Coumarines, Porphines, Metal-Ligand-Complexes, Fluorescent proteins, Nanocrystals, Perylenes and Phtalocyanines as well as conjugates and combinations of these classes of dyes.
26. The isolated targeted delivery system according to claim 20, wherein the label comprises a detectable polypeptide, wherein the detectable polypeptide is an autofluorescent protein.
27. The isolated targeted delivery system according to claim 1, wherein: (i) the bond(s) between the iron binding protein(s) and the pharmaceutically active substance, label or pharmaceutically active substance and label comprised in the complex are covalent and/or non-covalent; and/or (ii) the pharmaceutically active substance, label or pharmaceutically active substance and label comprised in the complex is entrapped/encapsulated by the iron binding protein or multimers thereof.
28. Method of preparation of the isolated targeted delivery system of claim 1 comprising steps of a) providing purified iron binding protein; b) covalently or non-covalently linking a pharmaceutically active substance, label or pharmaceutically active substance and label to and/or encapsulating a pharmaceutically active substance, label or pharmaceutically active substance and label in an iron binding protein; c) providing a CD45.sup.+ leukocyte cell; and d1) incubating the CD45.sup.+ leukocyte cell in the presence of the iron binding protein produced in step b) until the CD45.sup.+ leukocyte cell is at least partially loaded with the complex of the iron binding protein and the a pharmaceutically active substance, label or pharmaceutically active substance and label produced in step b); and/or d2) incubating CD45.sup.+ leukocyte cell in the presence of the label until the CD45.sup.+ leukocyte cell is at least partially labelled with the label.
29. The isolated targeted delivery system of claim 1 for use as a medicament or diagnostic.
30. A pharmaceutical composition comprising the isolated targeted delivery system of claim 1 and a pharmaceutically acceptable carrier and/or suitable excipient(s).
31. The isolated targeted delivery system of claim 1 for use in preventing, treating or diagnosing a tumour, an inflammatory disease or ischemic areas, or for prophylactic of therapeutic vaccination, in particular to prevent or treat an infectious disease or cancer.
Description
[0001] The present invention relates to an isolated cellular targeted delivery system comprising a CD45.sup.+ leukocyte cell comprising within said cell a complex of one or more iron binding proteins and an active pharmaceutically active substance and/or label as well as methods for producing such isolated cellular targeted delivery system and uses of such system for prophylaxis, therapy, diagnosis or theragnosis, in particular for prophylactic or therapeutic vaccination, therapy of cancer, particularly metastatic cancer or inflammatory diseases.
BACKGROUND OF THE INVENTION
[0002] Current imaging tools are capable of detecting large metastases (larger than 0.5-1 cm in size). However, they rarely detect the early spread of metastatic tumour cells. Human metastases smaller than 0.5 cm are avascular so without proper blood and oxygen supply. It means that the delivery of contrast agents through the blood circulation for the purpose of labelling these metastases and imaging them is not possible. The presence of hypoxia is a common characteristic of micrometastases where hypoxic fraction may be as high as 90% with little or no blood perfusion (Li, et al. 2012, Journal of Solid Tumours, 2(2): 28-33). Thus, severe hypoxia is considered as a general feature of micro-metastases.
[0003] The targeting of one or more micrometastases hidden within a large population of normal cells presents a unique challenge since access to the micrometastases is impeded by several bio-barriers, poor blood supply, further obstacles are presented by small size of the micrometastases and their dispersion to organs.
[0004] For the same reason micrometastases are often refractive to therapy. While the solid tumours from which the micrometastases have originated often respond well to conventional therapy there is often regrowth at the site of the primary tumour or at sites of metastasis. This constitutes a serious problem in clinical oncology (Muthana, et al. 2012, Cancer Res; 73(2); 490-495). It is related to characteristics of the microenvironment of solid tumours that limit drug penetration, thereby exposing the tumour to lower than efficacious concentrations of drugs (Hobbs, et al. 1998, Proc Natl Acad Sci USA: 4607-4612). This is caused by inadequate vasculature resulting in: high heterogeneity of cancer cells, low oxygen tension (hypoxia), low pH and low glucose concentration within the mass (Kizaka-Kondoh, et al., 2003, Cancer Sci 94(12):1021-1028). Additionally, rapid tumour cell proliferation in some areas might outpace the rate of new blood vessel growth, promoting formation of hypoxic area (Lewis and Murdoch, 2005, Am J Pathol 167(3):627-635). This abnormal vessel architecture and, subsequently, their impaired function resulting in tumour hypoxia is associated with a more malignant phenotype and poor survival in patients suffering from solid tumours and results in both treatment failure due to decreased drug uptake and hypoxiainducible changes in cancer cells (Sun, et al., 2012, Clin Cancer Res 18(3):758-770; Sullivan, et al., 2008 Mol Cancer Ther 7(7):1961-1973; Kizaka-Kondoh, et al., 2003, Cancer Sci 94(12):1021-1028). Moreover, chemotherapy or radiotherapy causes additional formation of large areas of tumour hypoxia thus making the treatment of tumour even more difficult. The fact that the efficacy of anticancer therapy is limited by the presence of hypoxic tumour cells has resulted in the introduction of variety of therapeutic approaches aimed at overcoming this problem.
[0005] The present inventors have discovered that CD45.sup.+ leukocyte cells, in particular activated macrophages, their precursors monocytes, lymphocytes and granulocytes can uptake pharmaceutically active substances, labels or pharmaceutically active substances and labels complexed with one or more iron binding proteins in vitro and deliver these complexes to or into cells, preferably to or into tumour cells in vivo or other cells undergoing stress (e.g. oxidative stress). Based on this observation the present inventors have overcome one or more of the above stated problems of the prior art. Thus, the targeted delivery system of the present invention provides inta alia one or more of the following advantages: (i) specific delivery of one or more pharmaceutically active substances, labels or pharmaceutically active substances and labels to tissues that attract above mentioned CD45.sup.+ leukocytes, preferably into diseased cells, (ii) protection of pharmaceutically active substances, labels or pharmaceutically active substances and labels from inactivation in the blood circulation or clearance from the body, (iii) delivery of pharmaceutically active substances, labels or pharmaceutically active substances and labels to, preferably into cells of poorly or non-vascularized areas of disease, e.g. metastases, hypoxic areas within larger tumours, rheumatic lesions, inflamed lymph nodes, avascular wounds, skin, (iv) reduced toxicity of pharmaceutically active substances, labels or pharmaceutically active substances and labels, (v) delivery of pharmaceutically active substances, labels or pharmaceutically active substances and labels with poor pharmacokinetics, (vi) reduced side effects of the drugs due to their targeted delivery, (vii) higher treatment efficacy with lower doses of the drugs due to targeted delivery; and/or (viii) lower risk of local tissue injury at the site of drug administration due to administration of the drug linked with iron-binding protein, which is loaded inside the CD45.sup.+ leukocyte; and/or (ix) possibility to detect highly hypoxic small metastases; and/or (x) early detection of inflammatory diseases.
SUMMARY OF THE INVENTION
[0006] In a first aspect the present invention relates to an isolated targeted delivery system comprising a CD45.sup.+ monocyte-macrophage, CD45.sup.+ lymphocyte, in particular CD45.sup.+ natural killer cells (NK cells), precursors of any of these cell types, preferably MDSC, and/or CD45.sup.+ granulocyte, ("CD45.sup.+ leukocyte cell"), or their precursors, preferably MDSC, comprising within said cell a complex of one or more iron binding proteins and a pharmaceutically active substance, label or pharmaceutically active substance and label.
[0007] In a second aspect the present invention relates to an isolated targeted delivery label system comprising a CD45.sup.+ leukocyte cell, preferably capable of being labelled, comprising one or more labels, preferably radiolabels or their conjugates and combinations.
[0008] In a third aspect the present invention relates to a method of preparation of the isolated targeted delivery system of claims 1 to 25 comprising steps of [0009] a) providing purified iron binding protein; [0010] b) covalently or non-covalently linking a pharmaceutically active substance, label or pharmaceutically active substance and label to and/or encapsulating a pharmaceutically active substance, label or pharmaceutically active substance and label in an iron binding protein; [0011] c) providing a CD45.sup.+ leukocyte cell; and [0012] d1) incubating the CD45.sup.+ leukocyte cell in the presence of the iron binding protein produced in step b) until the CD45.sup.+ leukocyte cell is at least partially loaded with the complex of the iron binding protein and the a pharmaceutically active substance, label or pharmaceutically active substance and label produced in step b); and/or [0013] d2) incubating CD45.sup.+ leukocyte cell in the presence of the label until the CD45.sup.+ leukocyte cell is at least partially labelled with the label.
[0014] In a fourth aspect the present invention relates to an isolated targeted delivery system of the first aspect or second aspect of the invention or producible according to the method of the third aspect of the invention for use as a medicament or diagnostic.
[0015] In a fifth aspect the present invention relates to a pharmaceutical or diagnostic composition comprising the isolated targeted delivery system of the first or second aspect of the present invention or producible according to the method of the third aspect of the invention and a pharmaceutically acceptable carrier and/or suitable excipient(s).
[0016] In a sixth aspect the present invention relates to an isolated targeted delivery system of the first aspect or second aspect of the present invention or producible according to the method of the third aspect of the invention for use in preventing, treating or diagnosing a disease characterized by hypoxic areas within the diseases tissue and/or by areas of oxidative stress; tumours, preferably solid tumours and/or its metastases, preferably breast cancer, pancreatic cancer, bladder cancer, lung cancer, colon cancer, or a tumour having hypoxic areas, an inflammatory disease, inflamed tissue, preferably inflamed joints or arthritic joints, inflamed lung, inflamed intestine or other inflamed tissue; lymph nodes, preferably inflamed lymph nodes, or other non-physiological lymph nodes that develop during disease preferably but not only during infection, cancer, or autoimmune disease; or ischemic areas, in particular in skin wounds or after organ infarctus (heart) or ischemic retina or for prophylactic or therapeutic vaccination, in particular to prevent or treat an infectious disease or cancer. This aspect also includes also of antigens to physiological or non-physiological lymph nodes in order to vaccinate an individual or to induce immune memory.
DETAILED DESCRIPTION OF PREFERRED EMBODIMENTS
[0017] Before the present invention is described in detail below, it is to be understood that this invention is not limited to the particular methodology, protocols and reagents described herein as these may vary. It is also to be understood that the terminology used herein is for the purpose of describing particular embodiments only, and is not intended to limit the scope of the present invention which will be limited only by the appended claims. Unless defined otherwise, all technical and scientific terms used herein have the same meanings as commonly understood by one of ordinary skill in the art.
[0018] In the following, the elements of the present invention will be described. These elements are listed with specific embodiments, however, it should be understood that they may be combined in any manner and in any number to create additional embodiments. The variously described examples and preferred embodiments should not be construed to limit the present invention to only the explicitly described embodiments. This description should be understood to support and encompass embodiments which combine the explicitly described embodiments with any number of the disclosed and/or preferred elements. Furthermore, any permutations and combinations of all described elements in this application should be considered disclosed by the description of the present application unless the context indicates otherwise.
[0019] Several documents are cited throughout the text of this specification. Each of the documents cited herein (including all patents, patent applications, scientific publications, manufacturer's specifications, instructions, etc.), whether supra or infra, are hereby incorporated by reference in their entirety. Nothing herein is to be construed as an admission that the invention is not entitled to antedate such disclosure by virtue of prior invention.
Definitions
[0020] To practice the present invention, unless otherwise indicated, conventional methods of chemistry, biochemistry, and recombinant DNA techniques are employed which are explained in the literature in the field (cf., e.g., Molecular Cloning: A Laboratory Manual, 2.sup.nd Edition, J. Sambrook et al. eds., Cold Spring Harbor Laboratory Press, Cold Spring Harbor 1989).
[0021] Throughout this specification and the claims which follow, unless the context requires otherwise, the word "comprise", and variations such as "comprises" and "comprising", will be understood to imply the inclusion of a stated integer or step or group of integers or steps but not the exclusion of any other integer or step or group of integers or steps. As used in this specification and the appended claims, the singular forms "a", "an", and "the" include plural referents, unless the content clearly dictates otherwise.
[0022] The term "targeted pharmaceutically active substance delivery" refers to the delivery of a pharmaceutically active substance to a patient which results in an increased concentration of the pharmaceutically active substance in a particular region of the body when compared to other regions of the body of that patient. Preferably, the relative concentrations are compared between the diseased region(s) of the body and other regions of the body having similar access to the blood circulation. In preferred embodiments the concentration of the pharmaceutically active substance in a given number of cells or a given biopsy volume from the diseased region is at least 10% higher, if compared to the identical number of cells or biopsy volume from a non-diseased region after administration of the targeted pharmaceutically active substance delivery system of the present invention, preferably after 2-24 hrs. More preferably, the concentration of the pharmaceutically active substance in the diseased region of the body of a patient is at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70% at least 80%, at least 90%, at least 100%, at least 150%, at least 200%, at least 250%, at least 300%, at least 350%, at least 400%, at least 450%, at least 500%, more preferably at least 1000% higher than in a non-diseased region of the body after administration of the targeted pharmaceutically active substance delivery system of the present invention, preferably after 2-24 hrs. When assessed on the basis of total body distribution it is preferred that at least 5% of the pharmaceutically active substance administered to a patient is delivered to the diseased region of the body, preferably at least 10%, more preferably at least 15%. The targeted delivery of the pharmaceutically active substance limits the potential deleterious effects of a pharmaceutically active substance to the diseased region of the body.
[0023] The term "targeted label delivery" or "targeted contrast agent delivery" refers to the delivery of a label, in particular a contrast agent to a patient or a person to be diagnosed which results in an increased concentration of the label, in particular the contrast agent in a particular region of the body when compared to other regions of the body of that patient. Preferably, the relative concentrations are compared between the diseased region(s) of the body and other regions of the body having similar access to the blood circulation. In preferred embodiments the concentration of the label, in particular a contrast agent in a given number of cells or a given biopsy volume from the diseased region is at least 10% higher, if compared to the identical number of cells or biopsy volume from a non-diseased region after administration of the targeted delivery system of the present invention, preferably after 1-24 hrs. More preferably, the concentration of the label, in particular the contrast agent in the diseased region of the body of a patient is at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70% at least 80%, at least 90%, at least 100%, at least 150%, at least 200%, at least 250%, at least 300%, at least 350%, at least 400%, at least 450%, at least 500%, more preferably at least 1000% higher than in a non-diseased region of the body after administration of the targeted delivery system of the present invention, preferably after 2-24 hrs. When assessed on the basis of total body distribution it is preferred that at least 5% of the label, in particular the contrast agent administered to a patient or a person to be diagnosed is delivered to the diseased region of the body, preferably at least 10%, more preferably at least 15%. The targeted delivery of the label, in particular the contrast agent limits the potential deleterious effects of an label, in particular the contrast agent to the diseased region of the body.
[0024] The term "targeted theragnostic delivery" has a similar meaning as set out above for "targeted pharmaceutically active substance" and "targeted label delivery, however instead of only delivering a pharmaceutical active substance or a label the complex of the iron binding protein comprises in this embodiment both a pharmaceutically active substance and a label, thereby allowing the concomitant delivery of both a pharmaceutically active substance and a label.
[0025] The term "targeted pharmaceutically active substance delivery system" is used in the present application to refer to a system that is capable of delivering a pharmaceutically active substance to the targeted region, i.e. capable of targeted delivery within the body of a patient, preferably to a diseased region.
[0026] The term "targeted label delivery system" is used in the present application to refer to a system that is capable of delivering a label to the targeted region, i.e. capable of targeted delivery within the body of a patient, preferably to a diseased region.
[0027] The term "targeted theragnostic delivery system" is used in the present application to refer to a system that is capable of delivering a complex of a pharmaceutically active substance and at the same time a label to the targeted region, i.e. capable of targeted delivery within the body of a patient, preferably to a diseased region and thus allows simultaneous treatment and diagnosis and/or treatment monitoring.
[0028] The term "targeted delivery system" is used to commonly refer to "targeted pharmaceutically active substance delivery system", "targeted label delivery system" and "targeted theragnostic delivery system"
[0029] The term "pharmaceutically active substance" is used in the context of the present invention to refer to a substance that modifies or modulates cell activity, preferably a cell activity that leads to a disease in a patient. The term encompasses both substances which are already pharmaceutically active or which are capable of being activated, i.e. a prodrug. Examples of such pharmaceutically active substances include so called (i) "small molecules", (ii) polynucleotides and (iii) peptide or proteins. The term "small molecule" is used in the context of the present invention to refer to a hydrocarbon with a molecular mass of below 1.500 g/mol or to pharmaceutically active radioactive isotopes. Preferred, drugs that can be used comprise anti-cancer drugs, pharmaceutically active radioactive isotopes or ferrihydrite.
[0030] The term "prodrug" as used in the context of the present invention refers to any active ingredient that, after administration, is metabolized or otherwise converted to a biologically active or more active ingredient (or drug) with respect to at least one property. In comparison to the drug, a prodrug is modified chemically in a manner that makes it, relative to the drug, less active or inactive, but the chemical modification is such that the corresponding drug is generated by metabolic or other biological processes after the prodrug is administered to the patient. A prodrug may for example have, relative to the active drug, altered metabolic stability or transport characteristics, fewer side effects or lower toxicity, or improved flavor (for example, see the reference Nogrady, 1985, Medicinal Chemistry A Biochemical Approach, Oxford University Press, New York, pages 388-392, incorporated herein by reference). A prodrug may be synthesized using reactants other than the corresponding drug.
[0031] The terms "polynucleotide" and "nucleic acid" are used interchangeably herein and are understood as a polymeric or oligomeric macromolecule made from nucleotide monomers. Nucleotide monomers are composed of a nucleobase, a five-carbon sugar (such as but not limited to ribose or 2'-deoxyribose), and one to three phosphate groups. Typically, a polynucleotide is formed through phosphodiester bonds between the individual nucleotide monomers. In the context of the present invention referred to nucleic acid molecules include but are not limited to ribonucleic acid (RNA), deoxyribonucleic acid (DNA), and mixtures thereof such as e.g. RNA-DNA hybrids. The nucleic acids, can e.g. be synthesized chemically, e.g. in accordance with the phosphotriester method (see, for example, Uhlmann, E. & Peyman, A. (1990) Chemical Reviews, 90, 543-584). "Aptamers" are nucleic acids which bind with high affinity to a polypeptide. Aptamers can be isolated by selection methods such as SELEmir146-a (see e.g. Jayasena (1999) Clin. Chem., 45, 1628-50; Klug and Famulok (1994) M. Mol. Biol. Rep., 20, 97-107; U.S. Pat. No. 5,582,981) from a large pool of different single-stranded RNA molecules. Aptamers can also be synthesized and selected in their mirror-image form, for example as the L-ribonucleotide (Nolte et al. (1996) Nat. Biotechnol., 14, 1116-9; Klussmann et al. (1996) Nat. Biotechnol., 14, 1112-5). Forms which have been isolated in this way enjoy the advantage that they are not degraded by naturally occurring ribonucleases and, therefore, possess greater stability.
[0032] The term "peptide" or "polypeptide" is used interchangeably in the context of the present invention to refer to a chain of at least two amino acids linked by peptide bonds. Thus, the term "peptide" in the context of the present invention is also used to refer to amino acid chains with more than 50, more than 100 or more than 150 amino acids.
[0033] The term "antibody" as used in the context of the present invention refers to a glycoprotein belonging to the immunoglobulin superfamily; the terms antibody and immunoglobulin are often used interchangeably. An antibody refers to a protein molecule produced by plasma cells and is used by the immune system to identify and neutralize foreign objects such as bacteria and viruses. The antibody recognizes a unique part of the foreign target, its antigen.
[0034] The term "antibody fragment" as used herein, refers to one or more fragments of an antibody that retain the ability to specifically bind to an antigen. Examples of binding fragments encompassed within the term "antibody fragment" include a fragment antigen binding (Fab) fragment, a Fab' fragment, a F(ab').sub.2 fragment, a heavy chain antibody, a single-domain antibody (sdAb), a single-chain fragment variable (scFv), a fragment variable (Fv), a V.sub.H domain, a V.sub.L domain, a single domain antibody, a nanobody, an IgNAR (immunoglobulin new antigen receptor), a di-scFv, a bispecific T-cell engager (BITEs), a dual affinity re-targeting (DART) molecule, a triple body, a diabody, a single-chain diabody, an alternative scaffold protein, and a fusion protein thereof.
[0035] The term "diabody" as used within this specification refers to a fusion protein or a bivalent antibody which can bind different antigens. A diabody is composed of two single protein chains which comprise fragments of an antibody, namely variable fragments. Diabodies comprise a heavy chain variable domain (V.sub.H) connected to a light-chain variable domain (V.sub.L) on the same polypeptide chain (V.sub.H-V.sub.L, or V.sub.L-V.sub.H). By using a short peptide connecting the two variable domains, the domains are forced to pair with the complementary domain of another chain and thus, create two antigen-binding sites. Diabodies can target the same (monospecific) or different antigens (bispecific).
[0036] A "single domain antibody", refers to antibody fragments consisting of a single, monomeric variable domain of an antibody. Simply, they only comprise the monomeric heavy chain variable regions of heavy chain antibodies produced by camelids or cartilaginous fish. Due to their different origins they are also referred to VHH or VNAR (variable new antigen receptor)-fragments. Alternatively, single-domain antibodies can be obtained by monomerization of variable domains of conventional mouse or human antibodies by the use of genetic engineering. They show a molecular mass of approximately 12-15 kDa and thus, are the smallest antibody fragments capable of antigen recognition. Further examples include nanobodies or nanoantibodies.
[0037] The term "antibody mimetic" as used within the context of the present specification refers to compounds which can specifically bind antigens, similar to an antibody, but are not structurally related to antibodies. Usually, antibody mimetics are artificial peptides or proteins with a molar mass of about 3 to 20 kDa which comprise one, two or more exposed domains specifically binding to an antigen. Examples include inter alia the LACI-D1 (lipoprotein-associated coagulation inhibitor); affilins, e.g. human-.gamma. B crystalline or human ubiquitin; cystatin; Sac7D from Sulfolobus acidocaldarius; lipocalin and anticalins derived from lipocalins; DARPins (designed ankyrin repeat domains); SH3 domain of Fyn; Kunits domain of protease inhibitors; monobodies, e.g. the 10.sup.th type III domain of fibronectin; adnectins: knottins (cysteine knot miniproteins); atrimers; evibodies, e.g. CTLA4-based binders, affibodies, e.g. three-helix bundle from Z-domain of protein A from Staphylococcus aureus; Trans-bodies, e.g. human transferrin; tetranectins, e.g. monomeric or trimeric human C-type lectin domain; microbodies, e.g. trypsin-inhibitor-II; affilins; armadillo repeat proteins. Nucleic acids and small molecules are sometimes considered antibody mimetics as well (aptamers), but not artificial antibodies, antibody fragments and fusion proteins composed from these. Common advantages over antibodies are better solubility, tissue penetration, stability towards heat and enzymes, and comparatively low production costs.
[0038] The term "antigen" is used to refer to a substance, preferably an immunogenic peptide that comprises at least one epitope, preferably an epitope that elicits a B or T cell response or B cell and T cell response.
[0039] An "epitope", also known as antigenic determinant, is that part of a substance, e.g. an immunogenic polypeptide, which is recognized by the immune system. Preferably, this recognition is mediated by the binding of antibodies, B cells, or T cells to the epitope in question. In this context, the term "binding" preferably relates to a specific binding. Epitopes usually consist of chemically active surface groupings of molecules such as amino acids or sugar side chains and usually have specific three-dimensional structural characteristics, as well as specific charge characteristics. The term "epitope" comprises both conformational and non-conformational epitopes. Conformational and non-conformational epitopes are distinguished in that the binding to the former but not the latter is lost in the presence of denaturing solvents.
[0040] An immunogenic polypeptide according to the present invention is, preferably, derived from a pathogen selected from the group consisting of viruses, bacteria and protozoa. In particular embodiments. However, in an alternative embodiment of the present invention the immunogenic polypeptide is a tumor antigen, i.e. polypeptide or fragment of a polypeptide specifically expressed by a cancer.
[0041] The term "sequence identity" is used throughout the specification with regard to polypeptide and nucleotide sequence comparisons. In case where two sequences are compared and the reference sequence is not specified in comparison to which the sequence identity percentage is to be calculated, the sequence identity is to be calculated with reference to the longer of the two sequences to be compared, if not specifically indicated otherwise. If the reference sequence is indicated, the sequence identity is determined on the basis of the full length of the reference sequence indicated by SEQ ID, if not specifically indicated otherwise. For example, a polypeptide sequence consisting of 200 amino acids compared to a reference 300 amino acid long polypeptide sequence may exhibit a maximum percentage of sequence identity of 66.6% (200/300) while a sequence with a length of 150 amino acids may exhibit a maximum percentage of sequence identity of 50% (150/300). If 15 out of those 150 amino acids are different from the respective amino acids of the 300 amino acid long reference sequence, the level of sequence identity decreases to 45%. The similarity of nucleotide and amino acid sequences, i.e. the percentage of sequence identity, can be determined via sequence alignments. Such alignments can be carried out with several art-known algorithms, preferably with the mathematical algorithm of Karlin and Altschul (Karlin & Altschul (1993) Proc. Natl. Acad. Sci. USA 90: 5873-5877), with hmmalign (HMMER package, http://hmmer.wustl.edu/) or with the CLUSTAL algorithm (Thompson, J. D., Higgins, D. G. & Gibson, T. J. (1994) Nucleic Acids Res. 22, 4673-80) available e.g. on http://www.ebi.ac.uk/Tools/clustalw/ or on http://www.ebi.ac.uk/Tools/clustalw2/index.html or on http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_clustalw- .html. Preferred parameters used are the default parameters as they are set on http://www.ebi.ac.uk/Tools/clustalw/ or http://www.ebi.ac.uk/Tools/clustalw2/index.html. The grade of sequence identity (sequence matching) may be calculated using e.g. BLAST, BLAT or BlastZ (or BlastX). BLAST protein searches are performed with the BLASTP program, score=50, word length=3. To obtain gapped alignments for comparative purposes, Gapped BLAST is utilized as described in Altschul et al. (1997) Nucleic Acids Res. 25: 3389-3402. When utilizing BLAST and Gapped BLAST programs, the default parameters of the respective programs are used. Sequence matching analysis may be supplemented by established homology mapping techniques like Shuffle-LAGAN (Brudno M., Bioinformatics 2003b, 19 Suppl 1:154-162) or Markov random fields. Structure based alignments for multiple protein sequences and/or structures using information from sequence database searches, available homologs with 3D structures and user-defined constraints may also be used (Pei J, Grishin N V: PROMALS: towards accurate multiple sequence alignments of distantly related proteins. Bioinformatics 2007, 23:802-808; 3DCoffee@igs: a web server for combining sequences and structures into a multiple sequence alignment. Poirot O, Suhre K, Abergel C, O'Toole E, Notredame C. Nucleic Acids Res. 2004 Jul. 1; 32:W37-40). When percentages of sequence identity are referred to in the present application, these percentages are calculated in relation to the full length of the longer sequence, if not specifically indicated otherwise.
[0042] The term "label" as used in the context of the present invention refers to any kind of compound being suitable for diagnostic purposes. Preferred compounds are selected from a fluorescent dye, a radioisotope and a contrast agent. A contrast agent is a dye or other substance that helps to show abnormal areas inside the body. In one embodiment the term label refers to a compound that comprises a chelating agent which forms a complex with divalent or trivalent metal cations. Preferred radioisotopes/fluorescence emitting isotopes are selected from the group consisting of alpha radiation emitting isotopes, gamma radiation emitting isotopes, Auger electron emitting isotopes, X-ray emitting isotopes, fluorescent isotopes, such as .sup.65Tb, fluorescence emitting isotopes, such as .sup.18F, .sup.51Cr, .sup.67Ga, .sup.68Ga, .sup.111In, .sup.99mTc, .sup.140La, .sup.175Yb, .sup.153Sm, .sup.166Ho, .sup.88Y, .sup.89Zr, .sup.90Y, .sup.149Pm, .sup.177Lu, .sup.47Sc, .sup.142Pr, .sup.159Gd, .sup.212Bi, .sup.72As, .sup.72Se, .sup.97Ru, .sup.109Pd, .sup.105Rh, .sup.101m15Rh, .sup.119Sb, .sup.128Ba, .sup.123I, .sup.124I, .sup.131I, .sup.197Hg, .sup.211At, .sup.169Eu, .sup.203Pb, .sup.212Pb, .sup.64Cu, .sup.67Cu, .sup.188Re, .sup.186Re, .sup.198Au and .sup.199Ag as well as conjugates, compounds comprising such isotopes and combinations of above with proteins, peptides, small molecular inhibitors, antibodies or other compounds, e.g. .sup.18F fluorodeoxyglucose (.sup.18F-FDG), .sup.89Zr-oxide or .sup.64Cu-porfirin. Preferred fluorescent dyes are selected from the following classes of dyes: Xanthens (e.g. Fluorescein), Acridines (e.g. Acridine Yellow), Oxazines (e.g. Oxazine 1), Cynines (e.g. Cy7/Cy 3), Styryl dyes (e.g. Dye-28), Coumarines (e.g. Alexa Fluor 350), Porphines (e.g. Chlorophyll B), Metal-Ligand-Complexes (e.g. PtOEPK), Fluorescent proteins (e.g APC, R-Phycoerythrin), Nanocrystals (e.g QuantumDot 705), Perylenes (e.g. Lumogen Red F300) and Phtalocyanines (e.g. IRDYE.TM.700DX) as well as conjugates and combinations of these classes of dyes or fluorescent .sup.65Tb emitting. Preferred contrast agents are selected from paramagnetic agents, e.g. Gd, Eu, W and Mn, preferably complexed with a chelating agent. Further options are superparamagnetic iron (Fe) complexes and particles, compounds containing atoms of high atomic number, i.e. iodine for computer tomography (CT), microbubbles and carriers such as liposomes that contain these contrast agents.
[0043] The term "leukocyte" is used in the context of the present invention to refer to cells of the immune system that are involved in protecting the body against both infectious disease and foreign invaders. All leukocytes are produced and derived from multipotent cells in the bone marrow known as a hematopoietic stem cells. Leukocytes are found throughout the body, including the blood and lymphatic system. All leukocytes have nuclei, which distinguishes them from the other blood cells, the anucleated red blood cells (RBCs) and platelets. Types of leukocyte can be classified in standard ways. Two pairs of the broadest categories classify them either by structure (granulocytes or agranulocytes) or by cell division lineage (myeloid cells or lymphoid cells). These broadest categories can be further divided into the five main types: neutrophils, eosinophils, basophils, lymphocytes, and monocytes. These types are distinguished by their physical and functional characteristics. Monocytes and neutrophils are phagocytic. Further subtypes can be classified; for example, among lymphocytes, there are B cells, T cells, and NK cells. Granulocytes are distinguished from agranulocytes by their nucleus shape (lobed versus round, that is, polymorphonuclear versus mononuclear) and by their cytoplasm granules (present or absent, or more precisely, visible on light microscopy or not thus visible). The other dichotomy is by lineage: Myeloid cells (neutrophils, monocytes, eosinophils and basophils) are distinguished from lymphoid cells (lymphocytes) by hematopoietic lineage (cellular differentiation lineage).
[0044] The present inventors have observed that CD45.sup.+ expression is characteristic of a subgroup of leukocyte cells, i.e. monocyte, monocyte-macrophages, lymphocytes, granulocytes, NK cells that are suitable to be used in the context of the targeted delivery system of the present invention, in particular since CD45.sup.+ leukocyte cells are attracted to particular tissues and cells within the body and are capable of delivering complexes of one or more iron binding proteins and one or more pharmaceutically active substances, labels or pharmaceutically active substances and labels to or into cells. This subgroup of leukocytes is in the following referred to as "CD45.sup.+ leukocyte cells" or "CD45.sup.+ leukocytes". Preferably the monocyte is not a dendritic cell which differentiation is controlled by one or more of the following transcription factors: IFN-regulatory factor 8 (IRF8), nuclear factor interleukin (IL)-3-regulated protein (NFIL3), basic leucine zipper transcriptional factor ATF-like 3 (BATF3) or Transcription Factor RelB (NF-KB Subunit)-RELB, Spi-1 Proto-Oncogene (PU/1), recombining binding protein suppressor of hairless (RBPJ), IFN-regulatory factor 4 (IRF4) or transcription factor E2-2 (also known as (TCF4).
[0045] It is understood by the skilled person that CD45.sup.+ leukocyte cells as defined above unless of clonal origin are a mixed population of different leukocytes which share the common property of expressing CD45.sup.+ surface antigen. Accordingly, subpopulations of cells within the diverse group of CD45.sup.+ leukocyte cells as defined above are characterized throughout the specification by further functional and/or structural characteristics. The term "CD45.sup.+" indicates that the majority of cells within a population of cells or essentially all cells express the CD45.sup.+ surface antigen. In this context and also with reference to other cellular surface antigens, the term "expresses" indicates that the surface antigen is produced within the cell and detectably exposed on the surface of a cell. The level of expression and, thus the number of surface antigens detectably exposed on the surface of a cell can vary greatly among different leukocytes. Generally, a cell is considered to be positive, i.e. is indicated to be "+", for a cellular surface antigen, if at least 5, preferably at least 10 copies of the surface antigen are detectably exposed on the surface of the cell. The skilled person is well aware of how to detect, quantify and select for cells, which are positive (or negative) for a given cellular surface antigen. Preferred methods include Fluorescence Activated Cell Sorting (FACS). In this technology fluorescently labelled antibodies are used to bind to cellular surface antigens of a population of cells, the cells are subsequently isolated into single cells and based on fluorescence intensity measured for the single cell, characterized as being positive or negative for the given cellular surface antigen. In some embodiments of the present invention it is indicated that the expression of a given protein is high or low. This means that the protein is detectably expressed in both instances, i.e. is "+", however, at different levels. High and low expression, respectively, will mean different absolute numbers of proteins per cell for different proteins. Thus, a given protein may be considered to be expressed at high levels if there are more than 500 detectable copies of that protein per cell and to be expressed at low levels if there are between 1 to 50 detectable copies of that protein per cell. However, another protein may be considered to be expressed at high levels, if there are more than 5000 detectable copies and expressed at low levels, if there are between 1 to 500 detectable copies per cell. It is well known in the art how to quantify the number of proteins expressed or produced in a cell using flow cytometry and Becton Dickinson Quantibrite.TM. bead method (see e.g. Pannu, K. K., 2001, Cytometry. 2001 Dec. 1; 45(4):250-8) or mass spectrometry (see, e.g. Milo, R., 2013, Bioessays, 35(12): 1050-1055). For the purpose of the present invention the term "high expression" of a given protein refers to detectable expression of that protein that is at least 70% of the highest expression level found, i.e. number of copies per cell, in a population of healthy CD45.sup.+ leukocytes. The term "low expression" of a given protein refers to detectable expression of that protein that is 30% or less of the highest expression level found, i.e. number of copies of that protein per cell, in a population of healthy CD45.sup.+ leukocytes. Preferably, the "highest expression level" is determined as the average of the highest expression levels found in healthy CD45.sup.+ leukocytes of different subjects. In some embodiments preferred subpopulations of cells are characterized as "producing" a given protein. This is understood to mean that the protein is not necessarily detectable on the surface of the cell but may only be present inside the cell. The skilled person is well aware how to detect and/or quantify production of a protein inside a cell and/or select cells producing such proteins. Alternatively, cell populations can be defined by expression of specific transcription factors. It is well known in the art how to determine expression of a given protein or its encoding mRNA in a population of cells or even in single cells, e.g. using in vivo labeling with antibodies, FISH assays, in vivo single molecule fluorescent microscopy (Crawford, R. et al. Biophys J. (2013) 105(11): 2439) alone or in combination with Fluorescent Activated Cell Sorting (FACS), or by the PrimeFlow technique (e Bioscience), (Adam S. Venable, et. al., (2015) Methods in Molecular Biology).
[0046] The term "differentiated monocyte" is used in the context of the present invention to refer to a monocyte differentiated from the committed precursor termed macrophage-DC precursor (MDP) mainly resident in bone marrow (but could be also in the spleen) and differentiate into either dendritic cells or macrophages. In mice they consist of two main subpopulations: (i) CD11b.sup.+ cell with high expression of CX3CR1, low expression of CCR2 and Ly6C.sup.- and (ii) CD11b.sup.+ cell with low expression of CX3CR1, high expression of CCR2 and Ly6C.sup.+. After leaving the bone marrow, mouse Ly6C.sup.+ monocytes differentiate into Ly6C.sup.- monocytes in circulation. Similarly, in human monocyte differentiation, it is accepted that CD14.sup.++ classical monocytes leave bone marrow and differentiate into CD14.sup.++CD16.sup.+ intermediate monocytes and sequentially to CD14.sup.+CD16.sup.++ non-classical monocytes in peripheral blood circulation (Yang et al. 2014; Biomark Res 2(1) doi. 10.1186/2050-7771-2-1). Preferably the differentiated monocyte is not a dendritic cell, which differentiation is controlled by one or more of the following transcription factors: IRF8, NFIL3, BATF3, RELB, PU/1, RBPJ, IIRF4, and/or TCF4, and more preferably is not a dendritic cell.
[0047] Macrophages are tissue-resident professional phagocytes and antigen-presenting cells (APC), which differentiate from circulating peripheral blood monocytes (PBMs). The term "activated macrophage" is used in the context of the present invention to refer to any macrophage that is polarized. Macrophage activation is in general achieved by incubation with interleukins, cytokines and/or growth factors. In particular IL-4 and M-CSF can be used as activating agents. Activated macrophages of different phenotypes are classified into M1-macrophages, classically activated macrophages (CAM) and M2-macrophages, alternatively activated macrophages (AAM). The classically activated M1-macrophages comprise immune effector cells with an acute inflammatory phenotype. These are highly aggressive against bacteria and produce large amounts of lymphokines (Murray, and Wynn, 2011, J Leukoc Biol, 89(4):557-63). The alternatively activated, anti-inflammatory M2-macrophages can be separated into at least three subgroups. These subtypes have various different functions, including regulation of immunity, maintenance of tolerance and tissue repair/wound healing. The term "M1 inducer" is used in the context of the present invention to refer to a compound that directs differentiation of PBMs to macrophages of the M1 type. The term "M2 inducer" is used in the context of the present invention to refer to a compound that directs differentiation of PBMs to macrophages of the M2 type. The skilled person is aware of a large number of ways to promote differentiation into either M1 or M2 macrophages.
[0048] The term "phagocytosis by macrophages" is the process by which a macrophage engulfs a solid particle to form an internal vesicle known as a phagosome.
[0049] The term "iron binding protein" as used refers to a protein that non-covalently binds an iron ion. Examples of such proteins comprise ferritin, haemoglobin, transferrin; and lactoferrin. Iron binding proteins are bound by cellular surface receptors which facilitate the internalization of these proteins into cells.
[0050] In a first aspect the present invention relates to an isolated targeted delivery system comprising a CD45.sup.+ monocyte, CD45.sup.+ monocyte-macrophage, CD45.sup.+ lymphocyte, CD45.sup.+ NK cell and/or CD45.sup.+ granulocyte (hereinafter referred to as "CD45.sup.+ leukocyte cell") comprising within said cell a complex of one or more iron binding proteins and a pharmaceutically active substance, label or pharmaceutically active substance and label. Preferably the CD45.sup.+ monocyte is not a dendritic cell, which differentiation is controlled by one or more of the following transcription factors: IRF8, NFIL3, BATF3, RELB, PU/1, RBPJ, IIRF4, and/or TCF4, and more preferably is not a dendritic cell.
[0051] It has been surprisingly found by the present inventors that CD45.sup.+ leukocyte cells, preferably activated macrophages, preferably M1 macrophages, more preferably M2 macrophages acquire pharmaceutically active substances, labels or pharmaceutically active substances and labels and deliver the pharmaceutically active substances, labels or pharmaceutically active substances and labels in the amount sufficient (i) to be detected using imaging systems to allowing them to be used to trace their location in the body. In case of .sup.18F-FDG this was particularly preferable in case of monocytes and activated monocytes, macrophages, activated macrophages, preferably M1 macrophages and most preferably M2 macrophages. Administration of cells-loaded with a label (.sup.18F-FDG, .sup.89Zr-oxide), preferably radioactive label can visualize organ with accumulation of the labelled cells preferably by positron emission tomography (PET) imaging, (ii) to elicit a prophylactic or therapeutic effect or to allow a therapy and diagnosis of a disease at the same time.
[0052] In a second aspect the present invention relates to an isolated targeted label delivery system comprising a CD45.sup.+ leukocyte cell, preferably capable of being labelled with one or more labels, preferably radiolabels or their conjugates and combinations. It is a preferred embodiment of the second aspect of the present invention that the cell is directly linked to or labelled with label. The label may be within said cell, or on its surface, preferably it is on the surface of the cell.
[0053] The following preferred embodiments in each case further specify both the first and second aspect of the present invention.
[0054] The ability of a given CD45.sup.+ leukocyte cell or cell population to internalize iron binding proteins depends on the expression of receptors involved in this internalization process. Receptors that lead to internalization of ferritin comprise, e.g. TfR, CXCR4, CD163, and TIM-2. The skilled person is well aware how to measure the amount of uptake of an iron binding protein and preferred method of measuring the uptake are described in the Example Section below. The present inventors also noted that subpopulations of CD45.sup.+ leukocyte cells have a certain propensity to internalize one iron binding protein over another iron binding protein and, thus can attain higher complex concentrations and/or show less leakage of the complex from the cells. Such CD45.sup.+ leukocyte subpopulations are described in more detail below.
[0055] The phrase "complex of one or more iron binding proteins and a pharmaceutically active substance, label or pharmaceutically active substance and label" as used in the context of the present invention refers to a composition in which one or more molecules of the pharmaceutically active substance, label or pharmaceutically active substance and label are covalently or non-covalently bound to one or more iron binding proteins. The covalent or non-covalent binding between the one or more iron binding proteins and the one more pharmaceutically active substance, label or pharmaceutically active substance and label can be direct or indirect. In the latter case the pharmaceutically active substance, label or pharmaceutically active substance and label is linked to the iron binding protein via a linker or spacer. Linker or spacers are known to the skilled artisan, such as polyalanine, polyglycin, carbohydrates, (CH.sub.2)n groups or polypeptide linkers. The skilled artisan will, thus, be able to select the respective suitable linker(s) or spacer(s) depending on the respective application. If the iron binding proteins form cages like, e.g. ferritin, than the term "complex" also encompasses the enclosure of pharmaceutically active substances, labels or pharmaceutically active substances and labels within the cage even in the absence of a covalent or non-covalent bond between the protein(s) and the active compound(s). The formation of the complex allows the transport of the pharmaceutically active substance, label or pharmaceutically active substance and label into the cell when the cell is internalizing the iron binding protein. Thus, it is preferred that the pharmaceutically active substances, labels or pharmaceutically active substances and labels are bound to the iron binding protein in a way that does not interfere with the transport mechanism. This can be easily tested by the skilled person using uptake assays known in the art and described in the Example Section below. It is preferred that the complex is sufficiently stable to survive the transport within the cell to the target region within the body. Thus, it is preferred that the complex rather than the pharmaceutically active substance, label or pharmaceutically active substance and label alone is delivered to the cells or into the cells in the target region. This property also reduces possible deleterious effects, e.g. cytotoxicity, of the pharmaceutically active substance, label or pharmaceutically active substance and label to the CD45.sup.+ leukocyte cell. If pharmaceutically active substances, labels or pharmaceutically active substances and labels are covalently coupled to the iron binding proteins such coupling is preferably through amino acids residues known to be located in surface areas that are not involved in binding to the cellular receptors required for cellular uptake of the iron binding proteins. Iron binding proteins used in the context of the present invention can form stable non-covalently bound complexes with a wide variety of pharmaceutically active substances, labels or pharmaceutically active substances and labels. If the pharmaceutically active substance or label is a peptide, e.g. an antigenic peptide it is preferred that it is not expressed as a fusion with the iron binding protein, since in this case release of the peptide from the iron binding protein will require endosomal processing of the entire iron binding protein peptide fusion protein.
[0056] The CD45.sup.+ leukocyte cells originate from the patient to be treated in such case the cell loaded with the complex would be autologous to the patient. It is also envisioned that patients are MHC typed prior to treatment with the targeted delivery of the present invention and that the leukocyte cell type used for a given patient is MHC matched to the patient. In these two preferred embodiments the CD45.sup.+ leukocyte cell is a primary cell or derived by a low number of differentiation steps from a primary cell. Alternatively, the CD45.sup.+ leukocyte cell may be from an immortalized but preferably non-transformed CD45.sup.+ leukocyte cell line. Thus the blood used for CD45.sup.+ leukocyte cells, i.e. CD45.sup.+ monocyte, CD45.sup.+ monocyte-macrophage, CD45.sup.+ granulocyte, or CD45.sup.+ lymphocyte, in particular CD45.sup.+ NK cell, isolation is preferably obtained from the patient to be treated or from a healthy donor. Alternatively the blood can be obtained from the blood bank. Use of umbilical cord blood is also considered herein.
[0057] The present inventors noted that a subpopulation of CD45.sup.+ leukocyte cells, which are producible from a CD34.sup.+ hematopoietic precursor cell are particular suitable for target specific delivery of the pharmaceutically active substance, label or pharmaceutically active substance and label. Accordingly, it is preferred that the leukocytes used to produce the target delivery system are derived from CD34.sup.+ hematopoietic precursor cells. The skilled person is well aware how to select CD34.sup.+ hematopoietic precursor cells and how to differentiate such cells into leukocytes.
[0058] As indicated above, the term "CD45.sup.+ leukocyte cell" is used throughout this specification to refer to a CD45.sup.+ monocyte, CD45.sup.+ monocyte-macrophage, CD45.sup.+ lymphocyte and/or CD45.sup.+. Preferably, the monocyte is not a dendritic cell, which differentiation is controlled by one or more of the following transcription factors: IRF8, NFIL3, BATF3, RELB, PU/1, RBPJ, IIRF4, and/or TCF4, and more preferably is not a dendritic cell. Preferred subpopulations in these general categories of leukocytes are defined in the following by structural parameters, e.g. presence or absence of a given protein, functional properties and/or method of their production/differentiation. As outlined above, the targeted delivery system of the present invention still provides the advantages outlined above, if in a population of cells not every cell has a particular property in as long as the majority of cells within that population has that property. Thus, in the following the property of one preferred cell of the targeted delivery system of the present invention is described. It is appreciated by the skilled person that a pharmaceutical composition of the present invention will comprises millions of cells and that not every cell within the population will have the functional and/or structural properties outlined herein but that the pharmaceutical composition can nevertheless be used to treat a disease, if the majority of cells share the respective functional and/or structural properties.
[0059] The present inventors have recognized that subpopulations of CD45.sup.+ leukocyte cells have particular advantageous properties including among others efficiency and/amount of complex uptake in general, ability to retain the complex within the cell, i.e. to avoid leakage and of target release of the pharmaceutically active substance, label or pharmaceutically active substance and label, efficiency of uptake of a particular iron binding protein and/or targeting to particular tissues or cells and, thus suitability to treat or ameliorate a particular disease. The present inventors have, e.g. observed that CD45.sup.+ leukocyte cells, which express one or more of the following antigens: CD204, CD206, CD200R, CCR2 have a preference for ferritin uptake over the uptake of other iron binding proteins. Thus, if the iron binding protein in the complex is ferritin it is preferred to select CD45.sup.+ leukocyte cells that express one or more of the following antigens: CD204, CD206, CD200R, CCR2. Accordingly, in a preferred embodiment of the present invention [0060] (i) the monocyte is a CD11b.sup.+ monocyte, preferably selected from the group consisting of a CD11b.sup.+ CD14.sup.+ monocyte, a CD11b.sup.+ CD16.sup.+ monocyte, a CD11b.sup.+ CD14.sup.+ CD16.sup.+ monocyte, a CD11b.sup.+ CD14.sup.+ MHCII.sup.+ monocyte, a CD11b.sup.+ CD14.sup.+ CD115.sup.+ monocyte, CD11b.sup.+ CD114.sup.+ monocyte, CD11b.sup.+ CD116.sup.+ monocyte, CD11b.sup.+ CCR1.sup.+ monocyte, CD11b.sup.+ CCR2.sup.+ monocyte, CD11b.sup.+ CX3CR.sup.+ monocyte, CD11b.sup.+ CXR4.sup.+ monocyte, CD11b.sup.+ CXR6.sup.+ monocyte and a CD11b.sup.+ CD14.sup.+ CD33.sup.+ monocyte, preferably the monocyte is not a dendritic cell, which differentiation is controlled by one or more of the following transcription factors: IRF8, NFIL3, BATF3, RELB, PU/1, RBPJ, IIRF4, and/or TCF4, and more preferably is not a dendritic cell; [0061] (ii) the differentiated monocyte or monocyte-macrophage is differentiated by M-CSF and selected from the group consisting of a macrophage, an activated macrophage, preferably a CD11b.sup.+ macrophage, more preferably a CD11b.sup.+ CD16.sup.+ macrophage, CD11b.sup.+ CD32.sup.+ macrophage, CD11b.sup.+ CD64.sup.+ macrophage, CD11b.sup.+ CD68.sup.+ macrophage, preferably a CD11b.sup.+ CD86.sup.+ M1 macrophage, preferably producing inducible nitric oxide synthetase (iNOS) and/or secreting interleukin 12 (IL-12) or preferably CD11b.sup.+ CCR2.sup.+ M2 macrophage, CD11b.sup.+ CD204.sup.+ M2 macrophage, CD11b.sup.+ CD206.sup.+ M2 macrophage, CD11b.sup.+ CD204.sup.+ CD206.sup.+ M2 macrophage, CD11b.sup.+ Major Histocompatibility Complex II.sup.+ (MHCII.sup.+) (low or hi expression) M2 macrophage, CD11b.sup.+ CD200R.sup.+ M2 macrophage, CD11b.sup.+ CD163.sup.+ M2 macrophage or activated macrophage producing and/or secreting Arginase-1 and/or interleukin 10 (IL10); preferably the differentiated monocyte is not a foam cell expressing Lectin-like oxidized low-density lipoprotein receptor-1 (Lox1.sup.+), C--X--C chemokine receptor type 7 (CXCR7.sup.+) and Nuclear factor (erythroid-derived 2)-like 2 (NRF2.sup.+). A foam cell is a type of macrophage that localize to fatty deposits on blood vessel walls, where they ingest low-density lipoproteins and become loaded with lipids giving them a foamy appearance. These cells secrete various substances involved in plaque growth and their death promotes inflammation, thereby contributing to cardiovascular disease; [0062] (iii) monocyte-macrophage or activated monocyte-macrophage is differentiated by M-CSF and is preferably expressing at least one chemokine receptor, preferably selected from the group consisting of CCR1, CCR2, CXCR4, and CXCR6, or at least one growth factor receptor, preferably selected from the group consisting of macrophage colony stimulating factor Receptor (CD115), granulocyte colony stimulating factor Receptor (CD114), and granulocyte-macrophage colony stimulating factor Receptor (consisting of CD116 and CD131); monocytes of these characteristics are particular suitable to treat inflammatory conditions and cancer; [0063] (iv) the lymphocyte is selected from the group consisting of a CD3.sup.+ and CD4.sup.+ or CD8.sup.+ T lymphocyte, or a CD19.sup.+, CD20.sup.+, CD21.sup.+, CD19.sup.+ CD20.sup.+, CD19.sup.+ CD21.sup.+, CD20.sup.+ CD21.sup.+, or CD19.sup.+ CD20.sup.+ CD21.sup.+ B lymphocyte; or a natural killer (NK) cell, preferably the NK cell is selected from the group consisting of CD56.sup.+ and without CD3 expression, or CD16.sup.+ CD56.sup.+, CD56.sup.+ CD94.sup.+, CD56.sup.+ CD158a.sup.+, CD56.sup.+ CD158f.sup.+, CD56.sup.+CD314.sup.+, CD56.sup.+CD335.sup.+ cell; or [0064] (v) the granulocyte is selected from the group consisting of a neutrophil, preferably a CD66b.sup.+ neutrophil, an eosinophil and a basophil, preferably a CD193.sup.+ eosinophil.
[0065] In a preferred embodiment of the targeted delivery system of the present invention the activated macrophage: [0066] (i) is producible by in vitro incubation of a monocyte or macrophage or their precursors with a factor capable of altering expression markers on macrophages, preferably [0067] (a) with at least one M1 inducer, [0068] (b) with at least one M2 inducer, [0069] (c) or with a factor capable of altering the macrophages ability to secrete cytokines, preferably IL-10 and IL-12, chemokines and/or to produce iNOS, arginase or other immunomodulating enzymes; examples of such factors are: activated platelets, IL-4, IL-10, IL-13, immune complex of an antigen and antibody, IgG, heat activated gamma-globulin, glucocorticosteroid, tumour growth factor-.beta. (TGF-.beta.), IL-1R, CC-chemokine ligand 2 (CCL-2), IL-6, Macrophage colony-stimulating factor (M-CSF), peroxisome proliferator-activated receptor .gamma. (PPAR.gamma.) agonist, leukocyte inhibitory factor (LIF), adenosine, helminth and fungal infection, lipopolysaccharide (LPS), interferon .gamma. (INF-.gamma.), viral and bacterial infection; in this respect it was observed that activation of a monocyte with a M1 inducer, particularly LPS will cause cell to express iNOS, that activation of a monocyte with a M1 inducer, particularly LPS will cause cell not to express Arginase-1, that activation of a monocyte with a M2 inducer, particularly IL-4 will cause cell to express Arginase-1, and that activation of a monocyte with a M2 inducer, particularly IL-4 will cause cell not to express iNOS, [0070] (ii) is characterized by expression of at least one of following antigens: CD64, CD86, CD16, CD32, high expression of MCHII, and/or production of iNOS and/or IL-12; [0071] (iii) is producible by in vitro incubation of a monocyte or macrophage with a factor capable of inducing the ability of the macrophage to phagocytose, e.g. IL-18, opsonins (for example complement-derived proteins such as iC3b, immunoglobulin G), calcitonin gene-related peptide (CGRP), lipopolysaccharide (LPS), interferon .gamma. (INF-.gamma.), viral infection and/or bacterial infection; [0072] (iv) is characterized by expression of at least one of following antigens: CD204, CD206, CD200R; CCR2, transferrin receptor (TfR), CXC-motive chemokine receptor 4 (CXCR4), CD163, and/or T cell immunoglobulin-domain and mucin-domain 2 (TIM-2), and/or show low expression of MHCII; activated macrophages having these properties are particularly suitable for complexes comprising ferritin as the iron binding protein; [0073] (v) has the ability to phagocytose; and/or [0074] (vi) is capable of cytokine secretion, preferably of IL-12, or IL-10, or production of inducible nitric oxide synthetase (iNOS) (or other pro-inflammatory compounds), arginase or other immunosuppressive/anti-inflammatory compounds.
[0075] In a preferred embodiment of the targeted delivery system of the present invention the M1 inducer for differentiating macrophages into M1 macrophages is selected from the group consisting of lipopolysaccharide (LPS), interferon .gamma. (INF-.gamma.), and viral and bacterial infection and the M2 inducer for differentiating macrophages into M2 macrophages is selected from the group consisting of IL-4, IL-10, IL-13, immune complex of an antigen and antibody, IgG, heat activated gamma-globulin, glucocorticosteroid, tumour growth factor-0 (TGF-.beta.), IL-1R, CC-chemokine ligand 2 (CCL-2), IL-6, Macrophage colony-stimulating factor (M-CSF), peroxisome proliferator-activated receptor .gamma. (PPAR.gamma.) agonist, leukocyte inhibitory factor (LIF), adenosine, helminth and fungal infection.
[0076] It has been surprisingly found by the present inventors that both complex loaded M1 macrophages and M2 macrophages are suitable for targeted active agent delivery into hypoxic tissues, preferably a tumour or its metastasis. In general we observed that 3 to 5% of the administered M1 macrophages localized at the tumour site while about 35% of the M2 macrophages showed tumour specific targeting. However, this general advantage of M2 macrophages were offset when using a complex comprising haemoglobin and drug, since significantly larger amounts of this complex could be loaded into M1 macrophages than into M2 macrophages. Generally this specific tropisms makes M2 macrophages more suitable for treating tumour and diseases characterized by hypoxic tissue.
[0077] The present inventors also found CCL2-activated macrophages (bone marrow derived) are particularly suitable to target lymph nodes, preferably inflamed lymph nodes. Thus, the use of such macrophages is preferred in all uses in which the targeting of lymph nodes is desired, in particular for prophylactic or therapeutic vaccination as well as for diagnosis of inflamed lymph nodes. Accordingly, the loading of CCL2-activated macrophages with complexes of iron binding proteins and antigens is preferred.
[0078] In a preferred embodiment of the targeted delivery system of the present invention the monocyte-macrophage: [0079] (i) is producible from a CD34.sup.+ hematopoietic precursor cell; [0080] (ii) is producible by in vitro incubation of monocytes with at least one inducer, preferably M1 or M2 inducer, more preferably at least one M2 inducer; [0081] (iii) is characterized by expression of at least one of the following antigens: TfR, CD163, TIM-2, CD14, CD16, CD33, and/or CD115; [0082] (iv) is characterized by expression of at least one of the following antigens: TfR, CD163, TIM-2, CXCR4, CD14, and/or CD16; and/or [0083] (v) has the ability to phagocytose; and/or [0084] (vi) is not a dendritic cell which differentiation is controlled by one or more of the following transcription factors: IRF8, NFIL3, BATF3 or RELB, PU/1, RBPJ, IRF4 or TCF4.
[0085] In this embodiment of the targeted delivery system of the present invention the M1 inducer for differentiating monocyte-macrophage cells is selected from the group consisting of LPS, INF-.gamma. or viral or bacterial infection or the M2 inducer for differentiating monocytes is selected from the group consisting of IL-4, IL-10, IL-13, immune complex of an antigen and antibody, IgG, heat activated gamma-globulins, Glucocorticosteroids, TGF-.beta., IL-1R, CCL-2, IL-6, M-CSF, PPAR.gamma. agonist, Leukocyte inhibitory factor (LIF), cancer-conditioned medium, cancer cells, adenosine and helminth or fungal infection.
[0086] It has been surprisingly found by the present inventors that monocytes are suitable for targeted active agent delivery into hypoxic tissues, preferably the tumour or its metastasis while monocytes treated with M2 activators are more suitable for targeted active agent delivery into hypoxic tissues, preferably the tumour or its metastasis. This specific tropisms make monocytes treated with M2 activators more suitable to targeting tumour and hypoxic sites, e.g. sites of inflammation.
[0087] In a preferred embodiment of the targeted delivery system of the present invention the lymphocyte: [0088] (i) is obtainable from blood, spleen, or bone marrow or is producible from a CD34.sup.+ precursor cell as known to the skilled person and also described in the, e.g. Lefort and Kim, 2010, J Vis Exp 40: 2017; Tassone and Fidler, 2012, Methods in Molecular Biology 882: 351-357; Kouro et al. 2005, Current Protocols in Immunology, 66:F22F.1:22F.1.1-22F.1.9; [0089] (ii) is an immunologically competent lymphocyte; [0090] (iii) expresses antigen specific T cell receptors; and/or [0091] (iv) is characterized by expression of at least one of the following antigens: (a) CD3 and CD4 or CD8 or (b): CD19, CD20, CD21, CD19 CD20, CD19 CD21, CD20 CD21, or CD19 CD20 CD21 antigen, and is preferably capable of producing immunoglobulins In a particularly preferred embodiment the CD45+ lymphocytes is a NK cell, which [0092] (i) is obtainable from blood, spleen or bone marrow or producible from a CD34.sup.+ precursor cell; and/or [0093] (ii) is characterized by the lack of CD3 expression and expression of at least one of the following CD56.sup.+ and/or CD94.sup.+, CD158a.sup.+ CD158f.sub.+ CD314.sup.+ CD335.sup.+.
[0094] In a preferred embodiment of the targeted delivery system of the present invention the granulocyte: [0095] (i) is obtainable from blood, spleen or bone marrow or producible from a CD34.sup.+ precursor cell as described, e.g. in Kuhs et al. 2015, Curr Protoc Immunol 111:7.23-1-7.23.16; Coquery et al. 2012, Cytometry A 81(9): 806-814; Swemydas and Lionakis 2013, J Vis Exp 77: 50586; [0096] (ii) is characterized by expression of at least one of the following CD66b and/or CD193; [0097] (iii) is a polymorphonuclear leukocyte characterized by the presence of granules in its cytoplasm; and/or [0098] (iii) is characterized by expression of at least one of the following: TfR, CD163, TIM-2, and/or CXCR4.
[0099] In a preferred embodiment of the targeted delivery system of the present invention the iron binding protein is selected from the group consisting of ferritin, preferably heavy (H) type ferritin, light (L) ferritin and/or mitochondrial ferritin; haemoglobin, preferably haemoglobin A, haemoglobin AS, haemoglobin SC, haemoglobin C, haemoglobin D, haemoglobin E, haemoglobin F, haemoglobin H; haemoglobin-haptoglobin complex, haemopexin, transferrin; and lactoferrin. The terms ferritin; haemoglobin, preferably haemoglobin A, haemoglobin AS, haemoglobin SC, haemoglobin C, haemoglobin D, haemoglobin E, haemoglobin F, haemoglobin H; haemoglobin-haptoglobin complex, hemopexin, transferrin; and lactoferrin encompass structural variants of the naturally occurring proteins and, thus relates to proteins that have at least 70%, preferably at least 75%, more preferably at least 80%, more preferably at least 85%, more preferably at least 90%, more preferably at least 95% more preferably at least 100% of the ability of the respective wild-type protein to bind iron ion(s). The iron binding proteins used in the context of the present invention are preferably of mammalian, more preferably mouse, rat, dog, ape, in particular, chimpanzee, or human, most preferably of human origin. Consensus sequences of the preferred iron binding proteins used in the context of the present invention are shown in FIG. 1 below. Preferred structural variants are based on the sequences indicated in FIG. 1. The residues marked with X vary among different mammalian ferritins, transferrins, and haemoglobins. The alteration of these residues is not crucial for the ability of the proteins to bind to iron ions. Accordingly, it is preferred that amino acid mutations or deletions effect one or more of the residues marked with an X.
[0100] Plasma proteins have always been privileged carriers for the delivery of active pharrrra ingredients in cancer therapy. Thus, albumin, the most abundant plasma protein, is currently used in therapeutic protocols for the delivery of taxane molecules and doxorubicin derivatives (Larsen M T et al. 2016, Mol Cell Ther 27; 4:3).
[0101] Human transferrin and ferritin proteins have been considered as effective carriers for the delivery of small molecules or toxin-conjugates to specifically target cancer cells. To date; in spite of considerable efforts, no successful transferrin or ferritin drug complexes have however reached the clinic (Luck A N et al. 2013, Adv Drug Deliv Rev 65(8):1012-9).
[0102] Ferritin has a cage architecture and capability of iron-binding which could be used to encapsulate pharmaceutically active substances and/or labels inside its cavity. Ferritins are not abundant in plasma, but can be readily produced in high yield as recombinant proteins in common protein expression vectors such as Escherichia coli cells. Ferritins H- or L-chains are encoded as small protein monomer (21 kDa and 19 KDa for H and L chains, respectively) capable of a 24-mer assembly into a cage-like structure, delimiting a 8 nm diameter cavity. The present inventors noted in the context of working on the present invention that H-ferritin homopolymers, represent a preferred protein construct in order to specifically deliver encapsulated drugs to CD45.sup.+ leukocyte cells expressing TfR. Furthermore H-ferritin targets complex pharmaceutically active substances, labels or pharmaceutically active substances and labels to the cell nucleus (and therefore directly delivers DNA-binding proteins into the nucleus.
[0103] Purified transferrin can be efficiently conjugated to various molecules including anticancer drugs through covalent linkers that are appropriately released inside the cells (Beyer U et al. 1998, J Med Chem 41(15):2701-2708). In case of transferrin, only lysine groups on the protein surface are ready available for covalent attachment.
[0104] Haemoglobin has been considered in the past as a possible drug carrier, due to its versatility in chemical conjugation with drugs, its abundance and relative stability in the blood (Somatogen, 1993, WO1993008842 A1). Nevertheless, the lack of receptor targeting properties did not foster biomedical applications other than blood substitutes or antisickling agent. As a matter of fact, Hb can only be recognized by CD163 (haptoglobin/haemoglobin receptor) epitopes on the leukocytes, especially monocyte-macrophage origin. The CD45.sup.+ leukocyte, in particular macrophage based protein delivery, described in this application moved haemoglobin center stage as a target specific carrier of pharmaceutically active substances and/or labels. Haemoglobin can be readily covalently linked to appropriate pharmaceutically active substances and/or labels, host hydrophobic pharmaceutically active substances and/or labels within the heme binding pocket or even transport small molecules, e.g. cytotoxic molecules linked to the heme iron. Hb can be easily modified by selective attachment of the appropriate drug conjugate to the beta93 cysteine residue, the only titratable cystein on the protein surface. Maleimido functionalized drugs, such as the tubuline inhibitor MonomethylAuristatin (MMAE) or the DNA crosslinking drug Pyrrolobenzodiazepine dimer (PBD) are most notable examples of extremely potent cytotoxics that can be readily and specifically attached to the relevant cys beta93 residue.
[0105] Alternatively, lysine residues on the Hb surface (at least 10 titratable lysine residues per Hb tetramer) may be easily amenable to drug conjugation through cleavable succinimide linkers. Haemoglobin also offers a unique capability of releasing non covalently bound heme group at acidic pH values. Apo-haemoglobin thus obtained is capable of hosting several hydrophobic molecules within the empty heme pocket, as shown in the case of paclitaxel (Meng Z et al. 2015 J Pharm Sci 104(3):1045-55) or for labels with fluorescent properties (e.g. chlorine e6, hyperycin, phtalocyanine derivatives) (Bong J et al. J Photochem Photobiol B 2014, 140:163-172).
[0106] Whatever the conjugation/adsorption/binding method, haemoglobin (Fib), transferrin (Tf) and ferritins were shown by the present inventors to be privileged carriers of pharmaceutically active substances and labels, once loaded into appropriate cell systems with tumour targeting properties, e.g. activated macrophages. The easy, fast, cheap and safe purification procedure of these protein also provide a tremendous added value.
[0107] Based on sequences of mammalian H-type ferritins, L-type ferritins, haemoglobin alpha chains, haemoglobin beta chains and transferrins a consensus sequence was determined for each of these proteins. These are shown in FIG. 1 in SEQ ID NO: 2, 7, 9, 14, 16, 20, and 25, respectively). On this basis but also on the basis of deletion and structural analysis disclosed in the prior art a minimal fragment was determined for H-type ferritins, L-type ferritins, haemoglobin alpha chains, and haemoglobin beta chains sufficient for uptake by CD45.sup.+ leukocytes. These are shown in SEQ ID NO: 1, 3, 5 (H-type ferritin); 8, 10, 12 (L-type ferritin), 15 and 17 (haemoglobin alpha chain) and 19 and 21 (haemoglobin beta chain. Transferrin comprises a N-terminally located domain and a C-terminally located domain that are necessary for binding iron and uptake by CD45.sup.+ leukocytes, if comprised in one polypeptide and positioned between 100 to 450 amino acids apart, preferably between 150 to 400, more preferably between 200 to 350 amino acids and more preferably 250 to 320 amino acids apart. The N-terminal domain comprises amino acids 1 to 82 of full length consensus transferrin (SEQ ID NO: 25) or full length human transferrin (SEQ ID NO: 28). The C-terminal domain comprises amino acids 396 to 510 of full length consensus transferrin (SEQ ID NO: 25) or full length human transferrin (SEQ ID NO: 28). In each case an X is indicated in the consensus sequence it independently stands for any amino acid and characterizes an amino acid not or only poorly conserved among mammalian H-type ferritins, which can be mutated without or little detriment to the iron binding properties of the respective iron binding protein. It is preferred that X in each case takes on the meaning of the amino acid of the respective human iron binding protein aligning with X. This information can be taken, e.g. from FIG. 1, which shows alignments of the consensus sequences with human an in some instances mouse proteins.
[0108] Preferred H-type ferritins comprise or consist of the amino acid sequence indicated in SEQ ID NO: 1 and variants thereof having at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a H-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 1 to be taken up by CD45.sup.+ leukocytes, preferably M2 leukocytes. Within SEQ ID NO: 1 X at position 5 may be present or absent, if present it means any amino acid, preferably Ile, X at position 6 means any amino acid, preferably Asn, X at position 14 means any amino acid, preferably Tyr, X at position 24 means any amino acid, preferably Tyr or Cys, X at position 66 means any amino acid, preferably Phe, X at position 68 means any amino acid, preferably Gln, X at position 75 means any amino acid, preferably Arg or Cys, X at position 90 means any amino acid, preferably His, X at position 94 means any amino acid, preferably Ser or Asn, X at position 120 may be present or absent, if present it means any amino acid, preferably His or Tyr, more preferably His, X at position 123 means any amino acid, preferably Asn or Ser, more preferably Asn, X at position 128 means any amino acid, preferably Ala or Ser, more preferably Ala.
[0109] In a preferred embodiment the H-type ferritin comprises or consists of murine ferritin according to SEQ ID NO: 3. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a H-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 3 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0110] In a preferred embodiment the H-type ferritin comprises or consists of human ferritin according to SEQ ID NO: 5. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a H-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 5 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0111] In a preferred embodiment the H-type ferritin comprises or consists of a mammalian consensus sequence derived from aligning full length H-type ferritins according to SEQ ID NO: 2 or 7, with 2 being preferred. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a H-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 2 or 7, with 2 being preferred to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages. In SEQ ID NO: 2 X at position 6 can be any naturally occurring amino acid, preferably Pro, X at position 14 can be any naturally occurring amino acid, preferably His, X at position 16 can be any naturally occurring amino acid, preferably Asp, X at position 21 may be present or absent, if present it means any amino acid, preferably Ile, X at position 22 means any amino acid, preferably Asn, X at position 30 can be any naturally occurring amino acid, preferably Tyr, X at position 40 can be any naturally occurring amino acid, preferably Tyr or Cys, more preferably Tyr, X at position 82 can be any naturally occurring amino acid, preferably Phe, X at position 84 can be any naturally occurring amino acid, preferably Gln, X at position 91 can be any naturally occurring amino acid, preferably Arg or Cys, more preferably Cys, X at position 106 can be any naturally occurring amino acid, preferably His, X at position 110 can be any naturally occurring amino acid, preferably Asn or Ser, more preferably Asn, X at position 137 can be any naturally occurring amino acid, preferably His or Tyr, more preferably His, X at position 140 can be any naturally occurring amino acid, preferably Asn or Ser, more preferably Asn, X at position 145 can be any naturally occurring amino acid, preferably Ala or Ser, more preferably Ala, X at position 164 can be any naturally occurring amino acid, preferably Ala or Ser, more preferably Ser, X at position 166 can be any naturally occurring amino acid, preferably Met or Leu, preferably Leu, X at position 178 can be any naturally occurring amino acid, preferably Asp or His, more preferably Asp, X at position 181 is absent or any naturally occurring amino acid, preferably Asn, X at position 182 is absent or any naturally occurring amino acid, preferably Glu, X at position 183 is absent or any naturally occurring amino acid, preferably Ser. In SEQ ID NO: 7 X at position 6 can be any naturally occurring amino acid, preferably Pro X at position 14 can be any naturally occurring amino acid, preferably His, X at position 16 can be any naturally occurring amino acid, preferably Asp, X at position 21 may be present or absent, if present it means any amino acid, preferably Ile, X at position 29 can be any naturally occurring amino acid, preferably Tyr, X at position 81 can be any naturally occurring amino acid, preferably Phe, X at position 83 can be any naturally occurring amino acid, preferably Gln, X at position 105 can be any naturally occurring amino acid, preferably His, X at position 144 can be any naturally occurring amino acid, preferably Ala or Ser, more preferably Ala, X at position 180 is absent or any naturally occurring amino acid, preferably Asn, X at position 181 is absent or any naturally occurring amino acid, preferably Glu, X at position 182 is absent or any naturally occurring amino acid, preferably Ser.
[0112] In a preferred embodiment the H-type ferritin comprises or consists of murine full length ferritin according to SEQ ID NO: 4 being preferred. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a murine H-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 4 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0113] In a preferred embodiment the H-type ferritin comprises or consists of human full length ferritin according to SEQ ID NO: 6 being preferred. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a human H-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 6 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0114] Preferred L-type ferritins comprise or consist of the amino acid sequence indicated in SEQ ID NO: 8 and variants thereof having at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a L-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 8 to be taken up by CD45.sup.+ leukocytes, preferably M2 leukocytes. In SEQ ID NO: 8 X at position at position 5 can be any naturally occurring amino acid, preferably Asp or Glu, more preferably Asp, X at position 12 can be any naturally occurring amino acid, preferably Arg or Ser, more preferably Ser, X at position 17 can be any naturally occurring amino acid, preferably Ser or Arg, more preferably Ser, X at position 19 can be any naturally occurring amino acid, preferably Arg or Gln, more preferably Gln, X at position 29 can be any naturally occurring amino acid, preferably Phe, X at position 30 can be any naturally occurring amino acid, preferably Tyr or Phe, more preferably Tyr, X at position 42 can be any naturally occurring amino acid, preferably Ser or Gly, more preferably Ser, X at position 56 can be any naturally occurring amino acid, preferably Ala or Tyr, more preferably Tyr, X at position 61 can be any naturally occurring amino acid, preferably Glu or Lys, more preferably Lys, X at position 62 can be any naturally occurring amino acid, preferably Met or Phe, more preferably Met, X at position 65 can be any naturally occurring amino acid, preferably Asp or Gln, more preferably Gln, X at position 75 can be any naturally occurring amino acid, preferably Ile or Val, more preferably Ile, X at position 76 can be any naturally occurring amino acid, preferably Lys or Gln, more preferably Lys, X at position 79 can be any naturally occurring amino acid, preferably Ala or Ser, more preferably Ala, X at position 80 can be any naturally occurring amino acid, preferably Glu or Gln, more preferably Gln, X at position 87 can be any naturally occurring amino acid, preferably Pro or Gln, more preferably Pro, X at position 88 can be any naturally occurring amino acid, preferably Glu or Asp, more preferably Asp, X at position 91 can be any naturally occurring amino acid, preferably Glu or Lys, more preferably Lys, X at position 94 can be any naturally occurring amino acid, preferably Met or Leu, more preferably Leu, X at position 96 can be any naturally occurring amino acid, preferably Met or Leu, more preferably Met, X at position 99 can be any naturally occurring amino acid, preferably Lys or Asn, preferably Lys, X at position 115 can be any naturally occurring amino acid, preferably Thr or Ala, more preferably Thr, X at position 119 can be any naturally occurring amino acid, preferably Leu, X at position 125 can be any naturally occurring amino acid, preferably Ser or Thr, more preferably Thr, X at position 127 can be any naturally occurring amino acid, preferably Tyr or Phe, more preferably Phe, X at position 130 can be any naturally occurring amino acid, preferably Lys or Glu, more preferably Glu, X at position 140 can be any naturally occurring amino acid, preferably Asp or Asn, more preferably Asp, X at position 146 can be any naturally occurring amino acid, preferably Arg or His, more preferably His, and X at position 148 can be any naturally occurring amino acid, preferably Leu or Val, more preferably Leu.
[0115] In a preferred embodiment the L-type ferritin comprises or consists of murine L-type ferritin according to SEQ ID NO: 10. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a L-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 10 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0116] In a preferred embodiment the L-type ferritin comprises or consists of human ferritin according to SEQ ID NO: 12. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a L-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 12 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0117] In a preferred embodiment the L-type ferritin comprises or consists of a mammalian consensus sequence derived from aligning full length H-type ferritins according to SEQ ID NO: 9 or 14, with 9 being preferred. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a L-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 9 or 14, with 9 being preferred to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages. In SEQ ID NO: 9 X at position 12 X at position 12 can be any naturally occurring amino acid, preferably Asp or Glu, more preferably Asp, X at position 19 can be any naturally occurring amino acid, preferably Ser or Arg, more preferably Ser, X at position 24 can be any naturally occurring amino acid, preferably Ser or Arg, more preferably Ser, X at position 26 can be any naturally occurring amino acid, preferably Arg or Gln, more preferably Gln, X at position 36 can be any naturally occurring amino acid, preferably Phe, X at position 37 can be any naturally occurring amino acid, preferably Tyr or Phe, more preferably Tyr, X at position 47 can be any naturally occurring amino acid, preferably Ser or Gly, more preferably Ser, X at position 63 can be any naturally occurring amino acid, preferably Ala or Tyr, more preferably Tyr, X at position 68 can be any naturally occurring amino acid, preferably Glu or Lys, more preferably Lys, X at position 69 can be any naturally occurring amino acid, preferably Met or Phe, more preferably Met, X at position 72 can be any naturally occurring amino acid, preferably Asp or Gln, more preferably Gln, X at position 82 can be any naturally occurring amino acid, preferably Ile or Val, more preferably Ile, X at position 83 can be any naturally occurring amino acid, preferably Lys or Gln, more preferably Lys, X at position 86 can be any naturally occurring amino acid, preferably Ala or Ser, more preferably Ala, X at position 87 can be any naturally occurring amino acid, preferably Glu or Gln, more preferably Gln, X at position 94 can be any naturally occurring amino acid, preferably Pro or Gln, more preferably Pro, X at position 95 can be any naturally occurring amino acid, preferably Glu or Asp, more preferably Asp, X at position 98 can be any naturally occurring amino acid, preferably Glu or Lys, more preferably Lys, X at position 101 can be any naturally occurring amino acid, preferably Met or Leu, more preferably Leu, X at position 103 can be any naturally occurring amino acid, preferably Met or Leu, more preferably Met, X at position 106 can be any naturally occurring amino acid, preferably Lys or Asn, preferably Lys, X can be any naturally occurring amino acid, X at position 126 can be any naturally occurring amino acid, preferably Leu, X at position 132 can be any naturally occurring amino acid, preferably Ser or Thr, more preferably Thr, X at position 134 can be any naturally occurring amino acid, preferably Tyr or Phe, more preferably Phe, X at position 137 can be any naturally occurring amino acid, preferably Lys or Glu, more preferably Glu, X at position 147 can be any naturally occurring amino acid, preferably Asp or Asn, more preferably Asp, X at position 153 can be any naturally occurring amino acid, preferably Arg or His, more preferably His, X at position 155 can be any naturally occurring amino acid, preferably Leu or Val, more preferably Leu, X at position 161 can be absent or any naturally occurring amino acid, preferably Ala, X at position 163 can be absent or any naturally occurring amino acid, preferably Thr, X at position 166 can be absent or any naturally occurring amino acid, preferably Pro, and X at position 168 can be any naturally occurring amino acid, preferably Gly or Ala, more preferably Ala. In SEQ ID NO: 14 X at position 36 can be any naturally occurring amino acid, preferably Phe, X at position 37 can be any naturally occurring amino acid, preferably Tyr or Phe, more preferably Tyr, X at position 94 can be any naturally occurring amino acid, preferably Pro or Gln, more preferably Pro, X at position 126 can be any naturally occurring amino acid, preferably Leu, X at position 137 can be any naturally occurring amino acid, preferably Lys or Glu, more preferably Glu, X at position 147 can be any naturally occurring amino acid, preferably Asp or Asn, more preferably Asp, X can be any naturally occurring amino acid, X at position 163 can be absent or any naturally occurring amino acid, preferably Thr, X at position 166 can be absent or any naturally occurring amino acid, preferably Pro, X at position 168 can be any naturally occurring amino acid, preferably Gly or Ala, more preferably Ala.
[0118] In a preferred embodiment the L-type ferritin comprises or consists of murine full length ferritin according to SEQ ID NO: 11 being preferred. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a murine L-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 11 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0119] In a preferred embodiment the L-type ferritin comprises or consists of human full length ferritin according to SEQ ID NO: 13 being preferred. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a human L-type ferritin consisting of the amino acid sequence according to SEQ ID NO: 13 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0120] In a preferred embodiment the alpha haemoglobin comprises or consists of a minimal mammalian consensus sequence derived from aligning full length alpha haemoglobins according to SEQ ID NO: 15 Preferred comprise or consist of the amino acid sequence indicated in SEQ ID NO: 15 and variants thereof having at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of an alpha haemoglobin consisting of the amino acid sequence according to SEQ ID NO: 15 to be taken up by CD45.sup.+ leukocytes, preferably M1 macrophages.
[0121] In a preferred embodiment the alpha haemoglobin comprises or consists of a minimal human amino acid sequence derived of human alpha haemoglobin according to SEQ ID NO: 17. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of an alpha haemoglobin consisting of the amino acid sequence according to SEQ ID NO: 17 to be taken up by CD45.sup.+ leukocytes, preferably M1 macrophages.
[0122] In a preferred embodiment the alpha haemoglobin comprises or consists of a mammalian consensus sequence derived from aligning full length alpha haemoglobins according to SEQ ID NO: 16. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of an alpha haemoglobins consisting of the amino acid sequence according to SEQ ID NO: 16 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0123] In a preferred embodiment the alpha haemoglobin comprises or consists of human full length alpha haemoglobin according to SEQ ID NO: 18. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a human full length alpha haemoglobin consisting of the amino acid sequence according to SEQ ID NO: 18 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0124] In a preferred embodiment the alpha haemoglobin comprises or consists of a minimal mammalian consensus sequence derived from aligning full length beta haemoglobins according to SEQ ID NO: 19 and variants thereof having at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a beta haemoglobin consisting of the amino acid sequence according to SEQ ID NO: 19 to be taken up by CD45.sup.+ leukocytes, preferably M1 macrophages.
[0125] In a preferred embodiment the alpha haemoglobin comprises or consists of a minimal human amino acid sequence derived of human beta haemoglobin according to SEQ ID NO: 21. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a beta haemoglobin consisting of the amino acid sequence according to SEQ ID NO: 21 to be taken up by CD45.sup.+ leukocytes, preferably M1 macrophages.
[0126] In a preferred embodiment the beta haemoglobin comprises or consists of a mammalian consensus sequence derived from aligning full length beta haemoglobins according to SEQ ID NO: 20. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of an beta haemoglobins consisting of the amino acid sequence according to SEQ ID NO: 20 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0127] In a preferred embodiment the beta haemoglobin comprises or consists of human full length beta haemoglobin according to SEQ ID NO: 22. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a human full length beta haemoglobin consisting of the amino acid sequence according to SEQ ID NO: 22 to be taken up by CD45.sup.+ leukocytes, preferably M2 macrophages.
[0128] In a preferred embodiment the transferrin comprises or consists of a mammalian consensus sequence derived from aligning full length alpha haemoglobins according to SEQ ID NO: 25. Thus, particularly, preferred transferrins comprise or consist of the amino acid sequence indicated in SEQ ID NO: 25 and of variants thereof having at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a transferrin consisting of the amino acid sequence according to SEQ ID NO: 25 to be taken up by CD45.sup.+ leukocytes preferably M1 macrophages.
[0129] In a preferred embodiment the transferrin comprises or consists of human transferrin according to SEQ ID NO: 28. Accordingly, preferred structural variants have at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a transferrin consisting of the amino acid sequence according to SEQ ID NO: 28 to be taken up by CD45.sup.+ leukocytes, preferably M1 macrophages.
[0130] The iron binding properties of transferrins are dependent on a N-terminally and C-terminally located domain. Thus, in a preferred embodiment the transferrin used in the present invention comprises at least the N-terminal domain according to SEQ ID NO: 23 and the C-terminal domain according to SEQ ID NO: 24. Preferred transferrin comprise proteins that comprise the amino acid sequence indicated in SEQ ID NO: 23 and 24 as well as variants thereof having at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a transferrin consisting of the amino acid sequence according to SEQ ID NO: 23 and 24 to be taken up by CD45.sup.+ leukocytes, preferably M1 macrophages. SEQ ID NO: 23 or 24 indicates a consensus sequence of mammalian transferrins.
[0131] Thus, in a preferred embodiment the transferrin used in the present invention comprises at least the N-terminal domain according to SEQ ID NO: 26 and the C-terminal domain according to SEQ ID NO: 27. Preferred transferrin comprise proteins that comprise the amino acid sequence indicated in SEQ ID NO: 26 and 27 as well as variants thereof having at least 70% amino acid identity, more preferably at least 75% amino acid identity, more preferably at least 80% amino acid identity, more preferably at least 85% amino acid identity, more preferably at least 90% amino acid identity, more preferably at least 95% amino acid identity and in each case at least 70% of the ability of a transferrin consisting of the amino acid sequence according to SEQ ID NO: 26 and 27, respectively, to be taken up by CD45.sup.+ leukocytes, preferably M1 macrophages.
[0132] Preferred ferritins, also comprise proteins that, irrespective of the given amino acid sequence, conform to the 24-mer subunit assembly of a four helix bundle protein module, falling within given sequence alignments of distantly related proteins as defined by 3D structure based alignments.
[0133] It is a surprising observation of the present inventors that monocytes, macrophages and preferably M2 macrophages are able to uptake the amount of label enough to make them visible using imaging methods at the site of their accumulation (e.g. in the tumour or its metastasis, hypoxia site). Surprisingly, inventors found out that lymphocytes and M2 macrophages are better in uptake of complexes comprising one or more ferritin and one or more pharmaceutically active substance, label or pharmaceutically active substance and label, that M1 macrophages are better in uptake of complexes comprising one or more haemoglobin and one or more pharmaceutically active substance, label or pharmaceutically active substance and label and that macrophages are better in uptake of complexes comprising one or more transferrin and one or more pharmaceutically active substance, label or pharmaceutically active substance and label. Accordingly, based on the tissue and cellular tropism of CD45.sup.+ leukocytes: monocytes, M1 and M2 macrophages, granulocytes and lymphocytes, described above complexes comprising one or more ferritin and one or more pharmaceutically active substance, label or pharmaceutically active substance and label are used to load M2 macrophages, lymphocytes or monocytes if the tropism of M2 macrophages, lymphocytes or monocytes is desired and complexes comprising one or more haemoglobin and one or more pharmaceutically active substance, label or pharmaceutically active substance and label are used to load M1 macrophages, if the tropism of M1 macrophages is desired.
[0134] In a preferred embodiment of the targeted delivery system of the present invention the pharmaceutically active substance, label or pharmaceutically active substance and label is selected from the group consisting of a protein, a nucleic acid, a chemical non-protein non-nucleic acid compound with a molecular weight of less than 1.5 kD, more preferably less than 1 kD, preferably an anticancer drug, in particular a cytostatic drug, cytotoxic drug and prodrugs thereof; an anti arteriosclerotic drug; and anti-inflammatory drug; and photosensitizing compound; a virus, in particular oncolytic virus; and .alpha. a or radiation emitting radioisotope, which also emit a cell damaging amount of .gamma. radiation, preferably selected from the group consisting of lutetium-177, ytterbium-90, iodine-131, samarium-153, phosphorus-32, caesium-131, palladium-103, radium-233, iodine-125, and boron-10 or a cell damaging amount of a radiation, preferably selected from the group consisting of actinium-225, bismuth-213, lead-212, and polonium-212. Also preferred is a complex of above mentioned compounds and isotopes linked to the nanoparticles (e.g. gold, argentum, graphen) or these nanoparticles.
[0135] Preferred anticancer drugs are selected from an apoptosis/autophagy or necrosis-inducing drug. An apoptosis/autophagy or necrosis-inducing drug can be any drug that is able to induce apoptosis/autophagy or necrosis effectively even in cells having an abnormality in cell proliferation. These drugs are preferably used in complexes with one or more ferritins.
[0136] Preferred anticancer drugs are selected from the group consisting of an apoptosis-inducing drug, an alkylating substance, anti-metabolites, antibiotics, epothilones, nuclear receptor agonists and antagonists, an anti-androgene, an anti-estrogen, a platinum compound, a hormone, a antihormone, an interferon, an inhibitor of cell cycle-dependent protein kinases (CDKs), an inhibitor of cyclooxygenases and/or lipoxygenases, a biogeneic fatty acid, a biogenic fatty acid derivative, including prostanoids and leukotrienes, an inhibitor of protein kinases, an inhibitor of protein phosphatases, an inhibitor of lipid kinases, a platinum coordination complex, an ethyleneimine, a methylmelamine, a triazine, a vinca alkaloid, a pyrimidine analog, a purine analog, an alkylsulfonate, a folic acid analog, an anthracendione, a substituted urea, and a methylhydrazin derivative, an ene-diyne antibiotic, a tubulin polymerization inhibitor such as a maytansinoid or an auristatine derivate, immune check-point inhibitor, and an inhibitor of tumour-specific protein or marker, preferably a Rho-GDP-dissociation inhibitor, more preferably Grp94, or AXL inhibitor.
[0137] Other preferred anticancer drugs are selected from the group consisting of acediasulfone, aclarubicine, ambazone, aminoglutethimide, L-asparaginase, azathioprine, banoxantrone, bendamustine, bleomycin, busulfan, calcium folinate, carboplatin, carpecitabine, carmustine, celecoxib, chlorambucil, cis-platin, cladribine, cyclophosphamide, cytarabine, dacarbazine, dactinomycin dapsone, daunorubicin, dibrompropamidine, diethylstilbestrole, docetaxel, doxorubicin, enediynes, epirubicin, epothilone B, epothilone D, estramucin phosphate, estrogen, ethinylestradiole, etoposide, flavopiridol, floxuridine, fludarabine, fluorouracil, fluoxymesterone, flutamide fosfestrol, furazolidone, gemcitabine, gonadotropin releasing hormone analog, hexamethylmelamine, hydroxycarbamide, hydroxymethylnitrofurantoin, hydroxyprogesteronecaproat, hydroxyurea, idarubicin, idoxuridine, ifosfamide, interferon .alpha., irinotecan, leuprolide, lomustine, lurtotecan, mafenide sulfate olamide, mechlorethamine, medroxyprogesterone acetate, megastrolacetate, melphalan, mepacrine, mercaptopurine, methotrexate, metronidazole, mitomycin C, mitopodozide, mitotane, mitoxantrone, mithramycin, nalidixic acid, nifuratel, nifuroxazide, nifuralazine, nifurtimox, nimustine, ninorazole, nitrofurantoin, nitrogen mustards, oleomucin, oxolinic acid, pentamidine, pentostatin, phenazopyridine, phthalylsulfathiazole, pipobroman, prednimustine, prednisone, preussin, procarbazine, pyrimethamine, raltitrexed, rapamycin, rofecoxib, rosiglitazone, salazosulfapyridine, scriflavinium chloride, semustine streptozocine, sulfacarbamide, sulfacetamide, sulfachlopyridazine, sulfadiazine, sulfadicramide, sulfadimethoxine, sulfaethidole, sulfafurazole, sulfaguanidine, sulfaguanole, sulfamethizole, sulfamethoxazole, co-trimoxazole, sulfamethoxydiazine, sulfamethoxypyridazine, sulfamoxole, sulfanilamide, sulfaperin, sulfaphenazole, sulfathiazole, sulfisomidine, staurosporin, tamoxifen, taxol, teniposide, tertiposide, testolactone, testosteronpropionate, thioguanine, thiotepa, tinidazole, topotecan, triaziquone, treosulfan, trimethoprim, trofosfamide, UCN-01, vinblastine, vincristine, vindesine, vinblastine, vinorelbine, and zorubicin, preferably selected from the group consisting of auristatin, banoxantrone, bendamustine, chlorambucil, chaliceamycin, cyclophosphamide dynemycin A, maytansine, melphalan, mertansine, and neocazinostatin, most preferably banoxantrone, bendamustine, chlorambucil, cyclophosphamide, pyrrolobenzodiazepine and melphalan.
[0138] It is particularly preferred that the anticancer drug is a proliferation inhibiting protein, preferably a cell cycle inhibitor or an antibody or antibody like binding protein that specifically binds to a proliferation promoting protein or a nucleic acid, preferably encoding a proliferation inhibiting protein or an antibody or antibody like binding protein that specifically binds to a proliferation promoting protein or a siRNA or DNAzym.
[0139] Preferred immunomodulatory drugs that activate or inhibit activity of immune cells. These can be natural or synthetic ligands or antagonists of Pattern Recognition Receptors, particularly Toll-like Receptors, NOD-like receptors (NLR), RIG-I-like receptors (RLR). Physiologically, these receptors recognize class of signals known as pathogen-associated molecular patterns (PAMPs) and damage-associated molecular patterns (DAMPs).
[0140] Preferred examples of antibodies to be used in the context of the present invention are single chain antibodies, antibody fragments, nanobodies, light or heavy chains, variable light or variable heavy chains, or diabodies. Preferred antibody fragments comprise a fragment antigen binding (Fab) fragment, a Fab' fragment, a F(ab')2 fragment, a heavy chain antibody, a single-domain antibody (sdAb), a single-chain fragment variable (scFv), a fragment variable (Fv), a VH domain, a VL domain, a single domain antibody, a nanobody, an IgNAR (immunoglobulin new antigen receptor), a di-scFv, a bispecific T-cell engager (BITEs), a dual affinity re-targeting (DART) molecule, a triple body, a diabody, a single-chain diabody, and a fusion protein thereof.
[0141] It has been surprisingly found by the present inventors that inflamed joints are targeted by macrophage-monocytes, preferably activated macrophages. Thus, in diagnostic applications in which inflamed joints are to be detected macrophage-monocytes are preferably loaded with a label or a complex comprising an iron binding protein and a label. In therapeutic applications, it is preferred that the complex comprises a pharmaceutically active substance with anti-inflammatory activity to deliver the anti-inflammatory substance to the inflamed tissue.
[0142] It has been surprisingly found by the present inventors that the targeted delivery system of the present invention targets lymph nodes, which makes it particularly suitable for delivery of antigens to dendritic cells residing in the lymph nodes. The lymph node targeting is particularly pronounced, if the cells loaded with the complex are macrophages in particular activated macrophages, even more preferably CCL-2 activated bone marrow derived activated macrophages, or lymphocytes, in particular B cells or T cells. Thus in a preferred embodiment the targeted delivery system is used to deliver one or more antigens in order to elicit a prophylactic and/or therapeutic immune response against the one or more antigens. Preferred antigens are derived from pathogens, i.e. bacteria or viruses or are tumour specific antigens. The term "tumour specific antigens" refers to proteins or epitopes (including peptides with altered glycosylation patterns) that are higher expressed on tumour cells in comparison to non-tumour cells, preferably to antigens or epitopes only expressed on tumour cells. Preferred antigens are selected from the group consisting of epidermal growth factor receptor (EGFR, ErbB-1, HER1), ErbB-2 (HER2/neu), ErbB-3/HER3, ErbB-4/HER4, EGFR ligand family; insulin-like growth factor receptor (IGFR) family, IGF-binding proteins (IGFBPs), IGFR ligand family; platelet derived growth factor receptor (PDGFR) family, PDGFR ligand family; fibroblast growth factor receptor (FGFR) family, FGFR ligand family, vascular endothelial growth factor receptor (VEGFR) family, VEGF family; HGF receptor family; TRK receptor family; ephrin (EPH) receptor family; AXL receptor family; leukocyte tyrosine kinase (LTK) receptor family; TIE receptor family, angiopoietin 1,2; receptor tyrosine kinase-like orphan receptor (ROR) receptor family; discoidin domain receptor (DDR) family; RET receptor family; KLG receptor family; RYK receptor family; MuSK receptor family; Transforming growth factor .alpha. (TGF-.alpha.) receptors, TGF-.beta.; Cytokine receptors, Class I (hematopoietin family) and Class II (interferon/IL-10 family) receptors, tumor necrosis factor (TNF) receptor superfamily (TNFRSF), death receptor family; cancertestis (CT) antigens, lineage-specific antigens, differentiation antigens, alpha-actinin-4, ARTC1, breakpoint cluster region-Abelson (Bcr-abl) fusion products, B-RAF, caspase-5 (CASP-5), caspase-8 (CASP-8), .beta.-catenin (CTNNB1), cell division cycle 27 (CDC27), cyclin-dependent kinase 4 (CDK4), CDKN2A, COA-1, dek-can fusion protein, EFTUD-2, Elongation factor 2 (ELF2), Ets variant gene 6/acute myeloid leukemia 1 gene ETS (ETC6-AML1) fusion protein, fibronectin (FN), GPNMB, low density lipid receptor/GDP-L fucose: .beta.-Dgalactose 2-.alpha.-Lfucosyltransferase (LDLR/FUT) fusion protein, HLA-A2. arginine to isoleucine exchange at residue 170 of the .alpha.-helix of the .alpha.2-domain in the HLA-A2 gene (HLA-A*201-R170I), HLA-A11, heat shock protein 70-2 mutated (HSP70-2M), KIAA0205, MART2, melanoma ubiquitous mutated 1, 2, 3 (MUM-1, 2, 3), prostatic acid phosphatase (PAP), neo-PAP, Myosin class I, NFYC, OGT, OS-9, pml-RARalpha fusion protein, PRDX5, PTPRK, K-ras (KRAS2), N-ras (NRAS), HRAS, RBAF600, SIRT2, SNRPD1, SYT-SSX1 or -SSX2 fusion protein, Triosephosphate Isomerase, BAGE, BAGE-1, BAGE-2,3,4,5, GAGE-1,2,3,4,5,6,7,8, GnT-V (aberrant N-acetyl glucosaminyl transferase V, MGAT5), HERV-K-MEL, KK-LC, KM-HN-1, LAGE, LAGE-1, CTL-recognized antigen on melanoma (CAMEL), MAGE-A1 (MAGE-1), MAGE-A2, MAGE-A3, MAGE-A4, MAGE-A5, MAGE-A6, MAGE-A8, MAGE-A9, MAGE-A10, MAGE-A11, MAGE-A12, MAGE-3, MAGE-B1, MAGE-B2, MAGE-B5, MAGE-B6, MAGE-C1, MAGE-C2, mucin 1 (MUC1), MART-1/Melan-A (MLANA), gp100, gp100/Pme117 (SILV), tyrosinase (TYR), TRP-1, HAGE, NA-88, NY-ESO-1, NY-ESO-1/LAGE-2, SAGE, Sp17, SSX-1,2,3,4, TRP2-INT2, carcino-embryonic antigen (CEA), Kallikrein 4, mammaglobin-A, OA1, prostate specific antigen (PSA), TRP-1/gp75, TRP-2, adipophilin, interferon inducible protein absent in melanoma 2 (AIM-2), BING-4, CPSF, cyclin D1, epithelial cell adhesion molecule (EpCAM), EphA3, fibroblast growth factor-5 (FGF-5), glycoprotein 250 (gp250), EGFR (ERBB1), HER-2/neu (ERBB2), interleukin 13 receptor .alpha.2 chain (IL13Ralpha2), IL-6 receptor, intestinal carboxyl esterase (iCE), alpha-feto protein (AFP), M-CSF, mdm-2, MUC1, p53 (TP53), PBF, PRAME, PSMA, RAGE-1, RNF43, RU2AS, SOX10, STEAP1, survivin (BIRC5), human telomerase reverse transcriptase (hTERT), telomerase, Wilms' tumor gene (WT1), SYCP1, BRDT, SPANX, XAGE, ADAM2, PAGE-5, LIP1, CTAGE-1, CSAGE, MMA1, CAGE, BORIS, HOM-TES-85, AF15q14, HCA661, LDHC, MORC, SGY-1, SPO11, TPX1, NY-SAR-35, FTHL17, NXF2, TDRD1, TEX15, FATE, TPTE, immunoglobulin idiotypes, Bence-Jones protein, estrogen receptors (ER), androgen receptors (AR), CD40, CD30, CD20, CD19, CD33, cancer antigen 72-4 (CA 72-4), cancer antigen 15-3 (CA 15-3), cancer antigen 27-29 (CA 27-29), cancer antigen 125 (CA 125), cancer antigen 19-9 (CA 19-9), .beta.-human chorionic gonadotropin, 1-2 microglobulin, squamous cell carcinoma antigen, neuron-specific enolase, heat shock protein gp96, GM2, sargramostim, CTLA-4, 707 alanine proline (707-AP), adenocarcinoma antigen recognized by T cells 4 (ART-4), carcinoembryogenic antigen peptide-1 (CAP-1), calcium-activated chloride channel-2 (CLCA2), cyclophilin B (Cyp-B), human signet ring tumor-2 (HST-2), Human papilloma virus (HPV) proteins (HPV-E6, HPV-E7, major or minor capsid antigens, others), Epstein-Barr virus (EBV) proteins (EBV latent membrane proteins-LMPL LMP2; others), Hepatitis B or C virus proteins, and HIV proteins.
[0143] If the pharmaceutically active substance is a nucleic acid it is preferred that it is a miRNA, siRNA, chemically modified-RNA, DNAzyme or a nucleic acid encoding a pharmaceutically active protein, e.g. an antibody, an antibody mimetic, a cytokine, a prodrug-converting enzyme, an immunogenic peptide or the like.
[0144] In a preferred embodiment of the targeted delivery system of the present invention the label is selected from a fluorescent dye, a fluorescence emitting isotope, a radioisotope, a detectable polypeptide or nucleic acid encoding such a detectable polypeptide and a contrast agent.
[0145] In a preferred embodiment of the targeted delivery system of the present invention the label comprises a chelating agent which forms a complex with divalent or trivalent metal cations.
[0146] In a preferred embodiment of the targeted delivery system of the present invention the chelating agent is selected from the group consisting of 1,4,7,10-tetraazacyclododecane-N,N',N,N'-tetraacetic acid (DOTA), ethylenediaminetetraacetic acid (EDTA), 1,4,7-triazacyclononane-1,4,7-triacetic acid (NOTA), triethylenetetramine (TETA), iminodiacetic acid, Diethylenetriamine-N,N,N',N',N''-pentaacetic acid (DTPA) and 6-Hydrazinopyridine-3-carboxylic acid (HYNIC).
[0147] In a preferred embodiment of the targeted delivery system of the present invention the contrast agent comprises a paramagnetic agent, preferably selected from Gd, Eu, W and Mn, or ferrihydride.
[0148] In a preferred embodiment of the targeted delivery system of the present invention the radioisotope/fluorescence emitting isotope is selected from the group consisting of alpha radiation emitting isotopes, gamma radiation emitting isotopes, Auger electron emitting isotopes, X-ray emitting isotopes, fluorescence emitting isotopes, such as .sup.18F, .sup.51Cr, fluorescent .sup.65Tb, .sup.67Ga, .sup.68Ga, .sup.111In, .sup.99mTc, .sup.140La, .sup.175Yb, .sup.153Sm, .sup.166Ho, .sup.88Y, .sup.90Y, .sup.149Pm, .sup.177Lu, .sup.47Sc, .sup.142Pr, .sup.159Gd, .sup.212Bi, .sup.72As, .sup.72Se, .sup.97Ru, .sup.109Pd, .sup.105Rh, .sup.101m15Rh, .sup.119Sb, .sup.128Ba, .sup.123I, .sup.124I, .sup.131I, .sup.197Hg, .sup.211At, .sup.169Eu, .sup.203Pb, .sup.212Pb, .sup.64Cu, .sup.89Zr, .sup.67Cu, .sup.188Re, .sup.186Re, .sup.198Au and .sup.199Ag as well as conjugates and combinations of above with proteins, peptides, small molecular inhibitors, antibodies or other compounds (e.g. .sup.18F-FDG, .sup.89Zr-oxide or .sup.64Cu-porfirin).
[0149] In a preferred embodiment of the targeted delivery system of the present invention the fluorescence dye is selected from the group consisting of the following classes of fluorescent dyes: Xanthens, Acridines, Oxazines, Cynines, Styryl dyes, Coumarines, Porphines, Metal-Ligand-Complexes, Fluorescent proteins, Nanocrystals, Perylenes and Phtalocyanines as well as conjugates and combinations of these classes of dyes.
[0150] In a preferred embodiment of the targeted delivery system of the present invention the detectable polypeptide is an autofluorescent protein, preferably green fluorescent protein or any structural variant thereof with an altered adsorption and/or emission spectrum.
[0151] As has been outlined above, the targeted delivery system of the present invention has particular suitability to deliver pharmaceutically active substances, labels or pharmaceutically active substances and labels to hypoxic areas. Hypoxy is characteristic of various disease including cancer and inflammatory diseases and thus allows targeting such diseases.
[0152] In addition to the targeting the use of pharmaceutically active substances, which are activated under hypoxic conditions adds a further specificity to the targeting and/or further reduces adverse effects of the pharmaceutically active substances, labels or pharmaceutically active substances and labels. Thus, in particularly preferred embodiments the pharmaceutically active substance, label or pharmaceutically active substance and label is a hypoxia-activated prodrug. The backbone of all the hypoxia-activated prodrugs is the presence of one of five different chemical moieties (nitro groups, quinines, aromatic and aliphatic N-oxides and transition metals) that are enzymatically reduced under hypoxic conditions in tissue. Hypoxia-activated prodrugs are any prodrug that is less active or inactive, relative to the corresponding drug, and comprises the drug and one or more bioreducible groups. Such hypoxia-activated prodrugs include all prodrugs activated by a variety of reducing agents and reducing enzymes, including without limitation single electron transferring enzymes (such as cytochrome P450 reductases) and two electron transferring (or hydride transferring) enzymes. According to preferred embodiment of the invention hypoxia-activated prodrug is TH-302. Methods of synthesizing TH-302 are described in PCT application WO 07/002931 and WO 08/083101. Preferably examples of such prodrugs are selected from the class I group consisting of: benzotriazine N-oxides, apaziquone (E09), tirapazamine (TPN) and SN30000; or class II group consisting of: nitro compounds PR104A, TH-302, TH-4000, and AQ4N.
[0153] In a preferred embodiment of the isolated targeted delivery system of the present invention the bond(s) between the iron binding protein(s) and the pharmaceutically active substance, label or pharmaceutically active substance and label comprised in the complex are covalent and/or non-covalent; and/or the pharmaceutically active substance, label or pharmaceutically active substance and label comprised in the complex is entrapped/encapsulated by the iron binding protein, preferably ferritin or multimers thereof. In one embodiment the covalent and/or non-covalent coupling is indirect through a linker or spacer. If the formation of covalent bonds is desired, relevant thiol, amino or carboxyl groups of the iron binding proteins are used to covalently couple pharmaceutically active substances, labels or pharmaceutically active substances and labels directly or indirectly to the one or more iron binding protein.
[0154] It is also envisioned that different pharmaceutically active substances, labels or pharmaceutically active substances and labels are comprised in the complex. For example, one type of pharmaceutically active substance, label or pharmaceutically active substance and label may be bound to an iron binding protein (non-covalently bound), while another type is encapsulated. This approach utilizes different release rates of the pharmaceutically active substances, labels or pharmaceutically active substances and labels from the iron binding protein once delivered to the targeted tissue and/or cells. For example, pharmaceutically active substance, label or pharmaceutically active substance and label can be covalently attached to ferritin molecule either on the surface of the 24-mer or within the internal cavity by exploiting the reactivity of relevant thiol, amino or carboxyl groups. The types of such useful reactions are well known in the art and can be adopted by the person skilled in the art to the particular pharmaceutically active substance, label or pharmaceutically active substance and label without any additional work. Examples of such reactions are described in Behrens C R, Liu B. Methods for site-specific drug conjugation to antibodies. MAbs. 2014 Jan-Feb; 6(1):46-53.
[0155] In theragnostic applications, i.e. in which the complex comprises both a label and a pharmaceutically active substance, it is preferred that the label is covalently attached to the iron binding protein and the pharmaceutically active substance is non-covalently bound to the iron binding protein and/or entrapped in the internal cavity formed upon assembly of the iron binding proteins.
[0156] In a third aspect the present invention relates to a method of preparation of the isolated targeted delivery system of the first or second aspect comprising the steps of [0157] a) providing purified iron binding protein; [0158] b) covalently or non-covalently linking a pharmaceutically active substance, label or pharmaceutically active substance and label to and/or encapsulating a pharmaceutically active substance, label or pharmaceutically active substance and label in an iron binding protein; [0159] c) providing a CD45.sup.+ leukocyte cell; and [0160] d1) incubating the CD45.sup.+ leukocyte cell in the presence of the iron binding protein produced in step b) until the CD45.sup.+ leukocyte cell is at least partially loaded with the complex of the iron binding protein and the a pharmaceutically active substance, label or pharmaceutically active substance and label produced in step b); and/or [0161] d2) incubating CD45.sup.+ leukocyte cell in the presence of the label until the CD45.sup.+ leukocyte cell is at least partially labelled with the label.
[0162] In a further aspect the present invention relates to the isolated targeted delivery system producible by the method according to the third aspect of the present invention.
[0163] The formation of the adduct between the protein and the pharmaceutically active substance and/or label is preferably by a non-covalent binding to the iron binding protein and can be described as follows: In the case of ferritin, pharmaceutically active substances and/or labels can be typically encapsulated within the internal cavity (physical confinement) by exploiting the association dissociation properties of the ferritin macromolecule itself. pharmaceutically active substance and/or label molecules are held in place by non-covalent interactions with amino acid residues within the cavity internal surface. Haemoglobin macromolecules also offer the possibility of non-covalent binding of selected pharmaceutically active substances and/or labels molecules that may be hosted within the heme binding pocket of haemoglobin itself. The heme in the pocket can be displaced and be replaced by pharmaceutically active substances and/or labels with appropriate hydrophobicity profile. In a further aspect, all proteins considered in the present invention may be covalently attached to pharmaceutically active substance and/or label molecules modified by specific active linker moieties reactive towards thiol or amino groups of the protein itself. As such, ferritins or haemoglobin may be linked to cysteine thiol reactive pharmaceutically active substances and/or labels bearing a peptide based cleavable linker (e.g. cathepsin sensitive valine-citrulline sequence and para-aminobenzylcarbamate spacer). As a notable example, the antimitotic agent monomethyl auristatin E (MMAE) has been used. The peptide-based linker binds the protein to the cytotoxic compound in a stable manner so the drug is not easily released from the protein under physiologic conditions and help prevent toxicity to healthy cells and ensure dosage efficiency. The protein pharmaceutically active substance and/or label adduct thus generated is capable of attaching to the selected receptor types, i.e. CD163 for haemoglobin and TfR for ferritin or transferrin, respectively. Once bound the protein pharmaceutically active substance and/or label adduct is internalised by endocytosis and thus selectively taken up by targeted cells. The vesicle containing the drug is fused with lysosomes and lysosomal cysteine proteases, particularly cathepsin B start to break down valine-citrulline linker and MMAE is no longer bound to the iron binding protein and is released directly into the tumour environment.
[0164] Alternatively, DM1-SMCC is an efficient mertansine derivative bearing a linker that specifically binds to lysine residues generating a covalent complex with ferritin, haemoglobin or transferrin in a reaction that has been successfully described for antibodies. In particular, haemoglobin, ferritin or transferrin can be reacted with DM1-SMCC thus providing a covalent protein-drug adduct that can be cleaved inside cells and releases the active drug in a time-dependent manner. The suppression of microtubule dynamics by DM1 induces mitotic arrest and cell death.
[0165] The term "full load" is used in the context of the present invention to refer to the maximum amount of iron binding protein, preferably ferritin, complexed with a pharmaceutically active substance, label or pharmaceutically active substance and label that can be taken up by the CD45+ leukocyte cell, preferably macrophage more preferably activated macrophage.
[0166] In a fourth aspect the present invention relates to an isolated targeted delivery system of the first aspect or second aspect of the invention or producible according to the method of the third aspect of the invention for use as a medicament or diagnostic.
[0167] In a fifth aspect the present invention relates to a pharmaceutical or diagnostic composition comprising the isolated targeted delivery system of the first or second aspect of the present invention or producible according to the method of the third aspect of the invention and a pharmaceutically acceptable carrier and/or suitable excipient(s). Since the isolated targeted delivery system comprises living cells, it is preferred that carriers and excipients are chosen in such to keep the cells alive.
[0168] "Pharmaceutically acceptable" means approved by a regulatory agency of the Federal or a state government or listed in the U.S. Pharmacopeia or other generally recognized pharmacopeia for use in animals, and more particularly in humans.
[0169] The term "carrier", as used herein, refers to a pharmacologically inactive substance such as but not limited to a diluent, excipient, surfactants, stabilizers, physiological buffer solutions or vehicles with which the pharmaceutically active substance is administered. Such pharmaceutical carriers can be liquid or solid. Liquid carrier include but are not limited to sterile liquids, such as saline solutions in water and oils, including but not limited to those of petroleum, animal, vegetable or synthetic origin, such as peanut oil, soybean oil, mineral oil, sesame oil and the like. Saline solutions and aqueous dextrose and glycerol solutions can also be employed as liquid carriers, particularly for injectable solutions. A saline solution is a preferred carrier when the pharmaceutical composition is administered intravenously. Examples of suitable pharmaceutical carriers are described in "Remington's Pharmaceutical Sciences" by E. W. Martin.
[0170] Suitable pharmaceutical "excipients" include starch, glucose, lactose, sucrose, gelatine, malt, rice, flour, chalk, silica gel, sodium stearate, glycerol monostearate, talc, sodium chloride, dried skim milk, glycerol, propylene, glycol, water, ethanol and the like.
[0171] "Surfactants" include anionic, cationic, and non-ionic surfactants such as but not limited to sodium deoxycholate, sodium dodecylsulfate, Triton X-100, and polysorbates such as polysorbate 20, polysorbate 40, polysorbate 60, polysorbate 65 and polysorbate 80.
[0172] "Stabilizers" include but are not limited to mannitol, sucrose, trehalose, albumin, as well as protease and/or nuclease antagonists.
[0173] "Physiological buffer solution" include but are not limited to sodium chloride solution, demineralized water, as well as suitable organic or inorganic buffer solutions such as but not limited to phosphate buffer, citrate buffer, tris buffer (tris(hydroxymethyl)aminomethane), HEPES buffer ([4 (2 hydroxyethyl)piperazino]ethanesulphonic acid) or MOPS buffer (3 morpholino-1 propanesulphonic acid). The choice of the respective buffer in general depends on the desired buffer molarity. Phosphate buffer are suitable, for example, for injection and infusion solutions.
[0174] The term "adjuvant" refers to agents that augment, stimulate, activate, potentiate, or modulate the immune response to the pharmaceutically active substance comprised in the composition at either the cellular or humoral level, e.g. immunologic adjuvants stimulate the response of the immune system to the actual antigen, but have no immunological effect themselves. Examples of such adjuvants include but are not limited to inorganic adjuvants (e.g. inorganic metal salts such as aluminium phosphate or aluminium hydroxide), organic adjuvants (e.g. saponins or squalene), oil-based adjuvants (e.g. Freund's complete adjuvant and Freund's incomplete adjuvant), cytokines (e.g. IL-1.beta., IL-2, IL-7, IL-12, IL-18, GM-CFS, and INF-.gamma.) particulate adjuvants (e.g. immuno-stimulatory complexes (ISCOMS), liposomes, or biodegradable microspheres), virosomes, bacterial adjuvants (e.g. monophosphoryl lipid A, or muramyl peptides), synthetic adjuvants (e.g. non-ionic block copolymers, muramyl peptide analogues, or synthetic lipid A), or synthetic polynucleotides adjuvants (e.g polyarginine or polylysine).
[0175] A striking observation was the disease specific homing of the targeted delivery system of the present invention. The CD45.sup.+ leukocyte cells appear to have a tropism for hypoxic areas and areas of oxidative stress. Hypoxia is a hallmark of various diseases as is oxidative stress. Accordingly, in a sixth aspect the present invention relates to an isolated targeted delivery system of the first aspect or second aspect of the present invention or producible according to the method of the third aspect of the invention for use in preventing, treating or diagnosing a disease characterized by hypoxic areas within the diseases tissue and/or by areas of oxidative stress (FIG. 27); tumours, preferably solid tumours and/or its metastases, preferably breast cancer, pancreatic cancer, bladder cancer, lung cancer, colon cancer, or a tumour having hypoxic areas, an inflammatory disease, inflamed tissue, preferably inflamed joints or arthritic joints, inflamed lung, inflamed intestine or other inflamed tissue; lymph nodes, preferably inflamed lymph nodes, or other non-physiological lymph nodes that develop during disease preferably but not only during infection, cancer, or autoimmune disease; or ischemic areas, in particular in skin wounds or after organ infarctus (heart) or ischemic retina or for prophylactic or therapeutic vaccination, in particular to prevent or treat an infectious disease or cancer. This aspect also includes also of antigens to physiological or non-physiological lymph nodes in order to vaccinate an individual or to induce immune memory.
[0176] The term "treatment" as used herein includes all types of preventive and/or therapeutic interventions medically allowed for the purpose of cure, temporary remission, prevention, etc. for different purposes including delaying or stopping the progress of a disease, making a lesion regress or disappear, preventing onset, or inhibiting recurrence.
[0177] The targeted delivery system according to the present invention enables tumour delivery of the pharmaceutically active substances, labels or pharmaceutically active substances and labels, which normally would not be able to reach the tumour (for example, due to solubility problems). It also enables the delivery of pharmaceutically active substances, labels or pharmaceutically active substances and labels to the hypoxic tumours or to the hypoxic areas of the tumour. This system also provides for delivery of pharmaceutically active substance, label or pharmaceutically active substance and label to any area within an organism subjected to hypoxic conditions, for example during ischaemic incidents, or undergoing an inflammatory process.
[0178] As mentioned above the present invention provides also the method for targeted delivery into the tumour mass. This method comprises preparation of CD45.sup.+ leukocytes, preferably activated macrophages which enables highly efficient iron-binding protein (ferritin, haemoglobin and/or transferring) uptake by the macrophages, wherein said ferritin, haemoglobin and/or transferrin carry a pharmaceutically active substance, label or pharmaceutically active substance and label (for example a drug/prodrug), tumour targeting and iron-binding protein transfer to the cancer cell, where the pharmaceutically active substance, label or pharmaceutically active substance and label is released.
[0179] The present invention exploits CD45.sup.+ leukocytes, preferably activated macrophages loaded with iron-binding proteins linked with a drug/prodrug as a delivery system to target the tumour. Unsatisfactory response of the tumours to chemotherapy or difficulties in their detection using imaging methods are mainly related to an altered penetration of the anticancer drugs to the hypoxic areas due to poor vasculature. However, these avascular regions attract CD45.sup.+ leukocytes, preferably activated macrophages to migrate even in areas far away from blood vessels. Therefore, they constitute a delivery system of particles to the tumour mass. A promising example of such particles is iron-binding protein. However, when used as single agents they do not reach hypoxic regions, similarly to other compounds and accumulate in other organs.
[0180] The present inventors linked anticancer drugs, hypoxia activated prodrugs (for treatment purposes), labels or isotopes to haemoglobin or transferrin using chemical methods and loaded it into CD45.sup.+ leukocytes (monocytes, macrophages, lymphocytes and/or granulocytes), preferably activated macrophages treating cells with iron-binding protein solution as it is described in examples. The inventors observed that upon administration to the animal, loaded CD45.sup.+ leukocytes, preferably activated macrophages migrate to the tumour hypoxic sites and release iron-binding protein with encapsulated pharmaceutically active substances, labels or pharmaceutically active substances and labels into the cancer cells. This method allows precise administration of the pharmaceutically active substances, labels or pharmaceutically active substances and labels to the tumour site (especially to the hypoxic regions), avoiding their accumulation in other organs.
BRIEF DESCRIPTION OF DRAWINGS
[0181] FIG. 1: Panel (A) shows a minimal active fragment of a consensus amino acid sequence among mammalian ferritin H chains and two full length consensus sequences based on several mammalian ferritin H chains (see SEQ ID NO: 1, 2 and 7, respectively) as well as a minimal and full length amino acid sequence of mouse (SEQ ID NO: 3 and 4) and human (SEQ ID NO: 5 and 6) ferritin H chain. Panel (B) shows a minimal active fragment of a consensus amino acid sequence among mammalian ferritin L chains and two full length consensus sequences based on several mammalian ferritin L chains (see SEQ ID NO: 8, 9 and 14, respectively) as well as a minimal and full length amino acid sequence of mouse (SEQ ID NO: 10 and 11) and human (SEQ ID NO: 12 and 13) ferritin L chain. Panel (C) shows a minimal active fragment of a consensus amino acid sequence among mammalian haemoglobin alpha chains and one full length consensus sequences based on several mammalian haemoglobin alpha chain (see SEQ ID NO: 15 and 16, respectively) as well as a minimal and full length amino acid sequence of human (SEQ ID NO: 17 and 18) haemoglobin alpha chain. Panel (D) shows a minimal active fragment of a consensus amino acid sequence among mammalian haemoglobin beta chains and a full length consensus sequences based on several mammalian haemoglobin beta chain (see SEQ ID NO: 19 and 20, respectively) as well as a minimal and full length amino acid sequence of human (SEQ ID NO: 21 and 22) haemoglobin beta chain. Panel (E) shows a N- and C-terminal minimal active fragment of a consensus amino acid sequence among mammalian transferrins (SEQ ID NO: 23 and 24) and a full length consensus sequences based on several mammalian transferrins (SEQ ID NO: 25) as well as a N- and C-terminal minimal active fragment of a human transferrin (SEQ ID NO: 26 and 27) and full length amino acid sequence of human transferrin (SEQ ID NO: 28). In the consensus sequences X indicates a position that is variable and stands for any natural amino acid. Preferably, in each case X in dependently of other X stands for the amino acid present in the human protein.
[0182] FIG. 2: Shows macrophage inside the mouse tumour mass (TRITC stained before injection, loaded with FITC-decorated ferritin).
[0183] FIG. 3: Shows confocal microscopy image of tumour tissue mouse injected with mammary cancer cells and given i.v. macrophages loaded with FITC-ferritin (asterix)--it is clearly observed, not only in macrophages but also in cancer cells, that ferritin-FITC spread within all tumour mass.
[0184] FIG. 4: Shows snapshots of one channel (in original green channel converted to the grey-scale picture) recording using confocal microscopy of macrophages (indicated with *; loaded with FITC-ferritin) and cancer cell (indicated with arrow; stained with red label and therefore not observed in green channel before ferritin uptake) in vitro taken at the starting time point (A) and after time long enough to fill cancer cell with ferritin (B). FITC-ferritin was dynamically transported to the cancer cell (accumulating firstly in the vesicles; then spreading to the whole cytoplasm as seen at this image (the cell appeared in green channel).
[0185] FIG. 5: Shows survival of mice receiving placebo and macrophages loaded with ferritin-coupled melphalan and ferritin-coupled chlorambucil.
[0186] FIG. 6: Shows tumour cells apoptosis caused by treatment with cyclophosphamide and cyclophosphamide encapsulated in ferritins loaded to macrophages (given at the same doses).
[0187] FIG. 7: Shows MRI images of mouse mammary tumour. The mouse was treated (at time point 0 h) with macrophages (i.v. injection) loaded with ferritin Fh. Then we observed increased diameter of blood vessels (arrow) filled with injected macrophages (giving significant T2-signal reduction) and afterword macrophages spread to the tissue (spot-like pattern; arrows). These changes (in the same time points) were observed in all examined mice.
[0188] FIG. 8: Shows ferritin, haemoglobin and transferrin uptake by macrophages, ferritin and haemoglobin uptake by monocytes and ferritin uptake by lymphocytes and granulocytes.
[0189] FIG. 9: Shows the stability of the ferritin storage by macrophages.
[0190] FIG. 10: Shows transfer of ferritin, haemoglobin and transferrin from macrophage to various cancer cells.
[0191] FIG. 11: Shows the transfer of ferritin from macrophage to cancer and non-cancer cells.
[0192] FIG. 12: Shows the transfer of ferritin encapsulated with hypoxia activated prodrug from macrophage to cancer cells.
[0193] FIG. 13: Shows the apoptosis in cancer cells that received ferritin with encapsulated various anticancer agents from co-cultured macrophages or soluble ferritin with the same agents.
[0194] FIG. 14: Shows the transfer of ferritin, haemoglobin and transferrin from monocyte to various cancer cells.
[0195] FIG. 15: Shows the transfer of ferritin and haemoglobin from granulocyte to various cancer cells.
[0196] FIG. 16: Shows the transfer of ferritin and haemoglobin from lymphocyte to various cancer cells.
[0197] FIG. 17: Shows the picture from two-photon microscopy showing tumour from a mouse that received pre-labeled (before administration) macrophages containing Ferritin-FITC.
[0198] FIG. 18: Shows the whole body imaging of mice that intravenously received labeled macrophages, showing their accumulation in the tumour site and their distribution in other organs.
[0199] FIG. 19: Shows the migration of macrophages to hypoxic tissue, a cross-section of the tumour from a mouse that was administered intravenously with pre-labeled macrophages, tumour hypoxic areas are visualized with a hypoxia marker--pimonidazolone.
[0200] FIG. 20: Shows the presents localization of vesicles containing FITC-laded ferritin (round objects) in the microenvironment inside the tumour mass. Macrophages containing FITC-laded ferritin were administered intravenously to the mouse.
[0201] FIG. 21: Shows the signal recorded by PET from a whole-body analysis of mice with the metastatic 4T1 cancer. Mice received intravenously macrophages loaded with .sup.18F-FDG. Signal accumulation is increased in the lungs of mice with micrometastases (confirmed by pathology examination). Mice receiving plain .sup.18F-FDG, or mice without 4T1 cancer had lower PET signal. MQ-FDG: indicates mice with metastatic 4T1 tumour+intravenous macrophages loaded with .sup.18F-FDG. FDG: indicates mice with metastatic 4T1 tumour+intravenous free .sup.18F-FDG. Naive mice MQ-FDG: Indicates naive mice without tumour+intravenous macrophages loaded with .sup.18F-FDG.
[0202] FIG. 22: Shows the tumour homing by different CD45+ leukocyte cell types usable in the targeted delivers systems of the invention. Mice with CT26 tumors (marked by the arrows) were intravenously injected with PBS or fluorescently labelled cells. Panel A. --PBS, no cells added, Panel B.--shows homing of a CD45.sup.+ leukocyte subpopulation, Panel C. shows homing of activated CD4.sup.+ T lymphocytes, Panel D. shows homing of a macrophage-monocytic cell line. 24 hours after injection mice were anaesthetized and imaged. Panel E. Shows the quantification of the tumour homing by different CD45+ leukocyte cells.
[0203] FIG. 23: Shows Survival of mice with CT26 tumor receiving therapy with monocytes loaded with ferritin-Cisplatin. n=6-8 mice/group. Survival of the mice was calculated by reaching (1500 mm3) maximum-allowed tumour volume (measured by caliper, according to formula A.times.B2).
[0204] FIG. 24: Shows targeting of intravenously administered cells to inflamed (arthritic) joints Control mouse (panel A.) or a mouse with severe inflammation of the joints (panel B.) have been intravenously injected with macrophage-monocyte cell line. Before injection cells were labelled with fluorescent tracer. 24 hours after injection joints were removed and single cell suspension generated. Cells have been analyzed by flow cytometry. Fluorescently labelled cells are presented in the P2 area. Panel C. shows quantification of the fluorescent cells in the inflamed than in control joints. Macrophage-monocyte cells are present with greater number in the inflamed than in control joints.
[0205] FIG. 25: Shows targeting of intravenously administered cells to the lymph nodes. Control mouse (panel A.) or a mouse with severe inflammation of the joints (panel B.) have been intravenously injected with activated macrophages (with M-CSF and CCL2). Before injection cells were labelled with fluorescent tracer. 24 hours after injection mice were anaesthetized and imaged. Mice with inflammatory condition have positive signal derived from cell-tracer, from the lymph nodes (marked by white arrows).
[0206] FIG. 26: Shows radiolabeling of monocyte-macrophage cell line with plain .sup.89Zr-oxinate, i.e. without iron binding protein (.about.1.85 MBq) for PET imaging. Radioactivity of cells after loading with .sup.89Zr is indicated in MBq.
[0207] FIG. 27: Ferritin transfer was examined in cancer cells (EMT6) co-cultured with ferritin-loaded macrophages (RAW264.7) for 2 hrs. Three experimental conditions were used: "ctrl" (normal co-culture, no stress); "EMT6 OS" when EMT6 cells were subjected to the oxidative stress before co-culture and then co-cultured with macrophages at normoxic conditions and "co-culture OS" when both cell types were subjected to the oxidative stress before the co-culture. Results showed that cancer cells subjected to the oxidative stress received (from macrophages) more ferritins than control cells. The uptake was even more efficient when both cell types were subjected to the oxidative stress. It can be related with higher load of ferritin due to oxidative stress and therefore more efficient transfer, or oxidative stress triggers macrophages to more efficiently deliver ferritins to cancer cells in order to reduce oxidative stress in neighboring cells. Panel A shows: Ferritin uptake from macrophages (2 hrs co-culture) in oxidative stress (oxidative glucose treatment for 16 hrs). Panel B shows: Ferritin uptake from macrophages (2 hrs co-culture) in oxidative stress (oxidative glucose treatment for 16 hrs).
EXAMPLE SECTION
Example 1--Activation of Macrophages
[0208] Macrophages for use according to the present invention were obtained, differentiated and activated as follows. In order to activate macrophages, they are obtained firstly from bone marrow precursors (for example see paper: Weischenfeld and Porse, 2008, CSH Protoc, doi. 10.1101/pdb.prot.5080) or blood monocytes. Alternatively, they can be obtained from peritoneum. The methods of macrophage isolation, culture, differentiation and polarization/activation are well known for those skilled in the art. For example, they have been described in details by Murray et al. (Immunity, 2014, 41(1):14-20).
[0209] In this practical realization of the invention bone marrow derived macrophages were obtained from BALB/c or C57Bl/6 mouse, however canine blood-monocyte-derived macrophages or commercially available macrophage cell lines (monocyte-macrophage lineage mouse cells: RAW 264.7, J744, human: THP-1, U937, or canine DH82 cell line).
[0210] Shortly, such bone marrow derived macrophages are seeded in plastic Petri dish in 5 ml medium (3 ml cells per plate): DMEM:F12+glutamine/glutamax+10% FBS+Penicillin/Streptomycin and 20% of L929 conditioned medium or M-CSF (50 ng/ml). In the next five days the medium is supplemented in growth factor and one of the activating compounds or their combinations as one cytokine cocktail.
[0211] Alternatively, macrophages have been cultured in "M1/M2 Macrophage Generation Medium" (Promocell) or equivalent commercially available or self-made medium containing all the necessary cytokines and interleukins to consider them as activated.
[0212] In order to obtain macrophages from blood monocytes, fresh blood (not older than 12 hours) cells were purified by centrifugation using Histopaque system 1.077 g/ml or equivalent and white blood cells (or alternatively, only white blood cells collected from the blood bank) in an appropriate amount of pre-warmed Monocyte Attachment Medium (or equivalent, e.g. DMEM/RPMI supplemented with M-CSF), e.g. 15 ml Medium per T-75 flask. A seeding density should be of 1-2 million/cm.sup.2 for mononuclear cells with a monocyte content of .gtoreq.2 5% and 1.5-3 million/cm.sup.2 for a monocyte content of <25%. Then, cells are incubated for 1-1.5 hours at 5% CO.sub.2 and 37.degree. C. in the incubator without any further manipulation.
[0213] After cell attachment, they are washed at least twice, and then an appropriate amount of complete "M1- or M2-Macrophage Generation Medium DXF" is added to the cells (e.g. 20 ml per T-75 flask) and cells are incubated for 6 days at 37.degree. C. and 5% CO.sub.2 without medium change. In order to activate macrophages, the whole medium is replaced with medium supplemented with activating compound.
[0214] Activating compounds used in this invention (for bone-marrow derived cells or to activate cells from monocyte-macrophage cell lines) are as follows: IL-4 (20 ng/ml), IFN-.gamma. (at least 20 ng/ml), LPS (at least 10 ng/ml), IL-13 (at least 20 ng/ml), IL-10 (at least 20 ng/ml), dexamethason (at least 20 .mu.g/ml), CCL2 (at least 20 ng/ml), oxLDL (at least 20 ng/ml), TNF-.alpha. (20 ng/ml), TGF-.beta. (20 ng/ml), cortisol (150-300 ng/ml) or their combinations as one cytokine cocktail. In order to obtain non-activated macrophages, the activating compound has not been added.
[0215] Reverse of the polarization/activation of macrophages (from classically activated to alternatively activated) can be reached for example by culture of macrophages in appropriate cytokines listed above for at least 48 hours.
Example 2--Monocyte Isolation
[0216] In order to obtain monocytes in this practical realization of the invention bone marrow derived or spleen-derived monocytes were obtained from BALB/c or C57Bl/6 mouse, however canine blood monocyte or commercially available monocyte cell lines were used (monocyte-macrophage lineage mouse cells: RAW 264.7, J744, human cells: THP-1, U937, or canine cells DH82).
[0217] To obtain blood monocytes, fresh blood (not older than 12 hours) cells were purified by centrifugation using Histopaque system 1.077 g/ml or equivalent and white blood cells are seeded in an appropriate amount of pre-warmed Monocyte Attachment Medium (or equivalent, e.g. DMEM/RPMI supplemented with 20 ng/ml M-CSF), e.g. 15 ml Medium per T-75 flask. Alternatively, only white blood cells collected from the blood bank (buffy coat) may be used. After cell attachment, they are washed at least twice, and adherent cells are considered as monocytes.
[0218] In order to obtain bone-marrow derived monocytes, in this practical realization of the invention monocytes were obtained from BALB/c or C57Bl/6 mouse. Shortly, such bone marrow derived precursors were isolated using Mouse Monocyte Enrichment Kit (StemCells) according to manufacturer's instructions. In order to obtain spleen derived monocytes, in this practical realization of the invention, the spleen has been mechanically dissociated to obtain single cell suspension and passed through the 70 .mu.m cell strainer. Cells were centrifuged and supernatant was removed. Optionally erythrocyte lysis was performed prior to monocyte isolation by EasySep.TM. Mouse Monocyte Enrichment Kit (StemCells) according to manufacturer's instructions.
[0219] To obtain better effects of their protein loads and migration before use they may be pre-treated with activation stimuli: IL-4 (20 ng/ml), IFN-.gamma. (at least 20 ng/ml), LPS (at least 10 ng/ml), IL-13 (at least 20 ng/ml), IL-10 (at least 20 ng/ml), dexamethason (at least 20 .mu.g/ml), CCL-2 (at least 20 ng/ml), oxLDL (at least 20 ng/ml), TNF-.alpha. (20 ng/ml), TGF-.beta. (20 ng/ml), cortisol (150-300 ng/ml) or their combinations as one cytokine cocktail.
Example 3--Granulocyte Isolation
[0220] To obtain granulocyte cells from blood, 9 parts of blood were diluted with 1 part of ACD buffer (containing 0.17 M d-glucose, 0.10 M citric acid, 0.11 M trisodium citrate). Blood from this step was further diluted with PBS at the 1:1 ratio and centrifuged. After removing plasma and buffy coat, remaining cells were mixed with PBS to 80% of the original volume from the first step (ACD-blood) and then diluted with cold distillated water at the ratio of 4:12. Then, 6 parts of 2.7% of NaCl solution were added and centrifuged. After removal of supernatant cells were resuspended in RPMI-1640 medium. These cell were considered as granulocytes.
Example 4--Lymphocyte Isolation
[0221] In order to obtain spleen derived lymphocytes, in this practical realization of the invention, the spleen has been mechanically dissociated to obtain single cell suspension and passed through the 70 .mu.m cell strainer. Cells were centrifuged and supernatant was removed. After erythrocyte lysis, lymphocytes were isolated using a negative cell isolation strategy with a help of EasySep.TM. CD4.sup.+ Enrichment Kit (Stemcell) according to manufacturer s instructions.
[0222] Lymphocytes were activated by the 3-5 day culture in RPMI medium supplemented with glutamine/glutamax, 10% FBS, Penicillin/Streptomycin, in the presence of beads covered with anti-CD3 and anti-CD28 (Gibco) according to manufacturer's instructions.
[0223] Activation of lymphocytes was confirmed with upregulation of CD25 and CD69 cell surface expression, that was monitored by flow cytometry.
Example 5--Leukocyte (CD45.sup.+ Cells) Isolation
[0224] In order to obtain macrophages from blood monocytes, fresh blood (not older than 12 hours) is purified by centrifugation using Histopaque system 1.077 g/ml or equivalent and white blood cells (or alternatively, only white blood cells collected from the blood bank) are used for further steps of the procedure. Alternatively, leukocytes are obtained after erythrocyte lysis of the whole blood, as it was done in this practical realization of the invention.
Example 6--Preparation of Ferritin Complexes
[0225] In order to incorporate ferritins with the anticancer drug (e.g. classic drugs like cyclophosphamide, chlorambucil, melphalan, bendamustine, banoxantrone or hypoxia-activated prodrug like TH-302) or with the "imaging contrast agents" (e.g. ferrihydride or isotope) ferritins have to be prepared before macrophage treatment. Shortly, recombinant mouse proteins according to SEQ ID NO: 4 (FIG. 1) are obtained as follows. The expression vector pET-22b containing a synthetic gene encoding ferritin protein of SEQ ID NO: 4 was transformed into E. coli BL21 (DE3). E. coli culture was grown at 37.degree. C. to OD600 0.6 in 1 L of Luria-Bertani broth (LB) added with ampicillin (100 mg/L). Protein expression was induced by addition of 1 mM isopropyl thio-b-D-galactoside (IPTG) and the culture was incubated overnight. Cells were harvested by centrifugation (15000 g for 15 min) and suspended in 20 mM Hepes (pH 7.5), 150 mM NaCl, 0.1 mg/mL DNase, 10 mM MgCl.sub.2 and disrupted by sonication. The lysate was centrifuged at 15000 g for 30 min and the supernatant was treated 10 min at 50.degree. C., centrifuged to remove denatured proteins and then at 70.degree. C. for 10 min and centrifuged again. The supernatant was added with 30% (NH).sub.4SO.sub.4 at 4.degree. C. stirring for 1 h and centrifuged at 15000 g for 30 min. The supernatant was added with 70% (NH).sub.4SO.sub.4 at 4.degree. C. stirring for 1 h and centrifuged at 15000 g for 30 min. The pellet was resuspended in 20 mM Hepes (pH 7.5), 150 mM NaCl and dialysed overnight at 4.degree. C. against the same buffer. The protein was loaded on a HILOAD 26/600 SUPERDEX 200 gel-filtration column (GE-Healthcare) and then sterile filtered and stored at 4.degree. C. (FIG. 9) Protein concentration was determined spectrophotometrically at 280 nm using a molar extinction coefficient of 21000 M.sup.-1 m.sup.-1 and by Bradford assay measuring the absorbance at 595 nm.
[0226] Said ferritins include recombinant mammalian ferritin proteins H and/or L homopolymers.
[0227] Ferritins, obtained as previously described, are purified by standard methods in order to obtain an endotoxin free, pre-clinical grade product (see, for example: Ceci et al. 2011, Extremophiles 15(3):431-439; Vanucci et al. 2012, Int J Nanomed 7:1489-1509). Shortly, the ferritin conserved sterile in a storage solution containing 20 mM Hepes pH 7.5 is diluted to a final concentration of 4 uM in 24-mer in acidic solution (final pH<3.0) or, alternatively, at highly basic pH values (pH>9.5) (see for example Pontillo et al., 2016), thus allowing the dissociation of multimer. Drugs are dissolved at very high concentrations in the appropriate solvent and then a small volume is added to the ferritin solution with a 200 molar excess. PH is then brought to neutrality by addition of appropriate amounts of NaOH/HCl solutions in order to allow multimer reconstitution. Current experimental methods indicate that three/four washings using PBS (concentration steps) in 100 kDa cut off concentrators allows rapid and complete elimination both the co-solvents as well as nonencapsulated drugs and full recovery of drug loaded ferritin nanocages. The ferritin-drug complex thus obtained was then flash freezed in liquid nitrogen and lyophilised.
[0228] Depending on the choice of co-solvent and on the intrinsic chemical properties of the drug molecule, it can be estimated that up to 150-180 drug molecules can be entrapped/adsorbed within the 24-mer ferritin cage.
[0229] Pharmaceutically active substances or labels may also be covalently coupled to ferritin amino acid side chains (lysines or cysteines) by appropriate choice of phenylhydrazone, succinimide or maleimide activated drugs. Accordingly, i) phenylhydrazone derivative may breaks and liberates the drug from the ferritin surface, ii) lysine bound derivatives may become active after full protein degradation into aminoacids or iii) cysteine bound derivative may be liberated within the cell through reductive hydrolysis of the maleimede thioether link.
Example 7--Preparation of Haemoglobin-Compound Complex
[0230] Human haemoglobin is prepared from fresh red cells as described in Rossi-Fanelli et al. (Archives of biochemistry and biophysics 77:478-492, 1958). Shortly, the heparinized blood, obtained from healthy donors, was centrifuged at 1600 rpm for 30 minutes (4.degree. C.) to sediment the RBCs. Buffy coat was accurately removed by needle aspiration on the surface of the pellet. The plasma supernatant was discarded and the RBC pellet was washed three times by resuspending the RBCs in isotonic 0.9% saline solution and centrifuging at 1600 rpm for 30 minutes at 4.degree. C. After the final saline wash and centrifugation step, the RBC pellet was resuspended in distilled water buffered at pH 7.2 with 5 mM potassium phosphate buffer (PB, pH=7.2) and allowed to lyse at 4.degree. C. overnight under gentle stirring. Dialyzed RBC lysate was subsequently centrifuged at 13.000 rpm for 30 min at 4.degree. C. and supernatant was directly loaded on an AKTA Explorer system equipped with an XK 26/40 column packed with Q-sepharose XL resin (GE Healthcare) at room temperature. Columns were equilibrated with buffer A (20 mM Tris-HCl, pH=8.2) at a flow rate of 12 mL/min and washed three times with the same buffer. A linear gradient elution was generated by changing from 100% buffer A to 75% buffer B (20 mM Tris-Cl, plus 0.2 M NaCl pH8.20) followed by a step gradient of 100% buffer B. Upon elution, a fraction collector was used to collect protein fractions. Protein thus obtained was analyzed by SDS page and stored frozen at -80.degree. C.
[0231] Human Haemoglobin (SEQ ID NO: 18 or 22, see FIG. 1) can be readily covalently linked to appropriate drug conjugates, host hydrophobic drug molecules within the heme binding pocket or even transport small cytotoxic molecules linked to the heme iron. Hb can be easily modified by selective attachment of the appropriate drug conjugate to the cysteine residue in position 93 of the beta chains, the only titratable cystein on the protein surface. Maleimido functionalized drugs, such as the tubuline inhibitor MonomethylAuristatin (MMAE) or the succinimide functionalized mertansine analogue (DM1-SMCC) are most notable examples of extremely potent cytotoxics that can be readily and specifically attached to the relevant cys beta93 residue (for maleimido functionalized drugs) or to one or more lysine residues (succinimide functionalized drug), respectively. These drugs have been conveniently conjugated to human haemoglobin according to the following procedures:
[0232] The auristatin E analogue, maleimidocaproyl-valine-citrulline-paminobenzoyloxycarbonyl-monomethyl auristatin E (vcMMAE) was obtained from MedChem Express (Princeton, N.J.). The Haemoglobin vcMMAE adduct was prepared as follows. Human haemoglobin solution was adjusted to a concentration of 120 .mu.M heme with reaction buffer (50 mM phosphate buffer pH 6,8, containing 0.1 mM EDTA) and conjugated with 10-fold molar excess of vcMMAE in the presence of 20% v/v acetonitrile solution at 4.degree. C. overnight. Maleimide groups react efficiently and specifically with free (reduced) sulfhydryls at pH 6.5-7.5 to form stable thioether bonds. The excess vcMMAE was purified and buffer-exchanged with D-PBS using PM 100 ultrafiltration concentrator. The yield of conjugation was approximately 80% of the total cysteines. Formation of vcMMAE conjugate was confirmed by LC-MS analysis and by titration of residual free thiol group with p-chloromercuribenzoate. The concentrations of Hb-vcMMAE conjugates were determined by UV-vis spectroscopy analysis.
[0233] The mertansin analogue DM1 SMCC (Alb Technology Ltd, Hederson, Nev., USA), functionalized for lysine covalent attachment, was prepared as follows. Human haemoglobin solution was adjusted to a concentration of 400 .mu.M heme with reaction buffer (0.1 mM phosphate buffer pH 7.4, containing 0.5 mM EDTA) and conjugated with 20-fold molar excess of DM1-SMCC in the presence of 10% v/v DMSO solution at 4.degree. C. for 16 hours. The amine-reactive succinimidyl ester couples to amines thus yielding a covalent adduct with lysine groups on the surface of the protein. The excess DM1-SMCC was eliminated and buffer-exchanged with D-PBS using PM 100 ultrafiltration concentrator. The yield of conjugation was approximately 2.4 mertansine molecules per haemoglobin tetramer. Formation of DM1-SMCC conjugate was confirmed by LC-MS analysis. The concentrations of Hb-DM1-SMCC conjugates were determined by UV-vis spectroscopy analysis.
[0234] Alternatively, the apo-protein solution must be kept on an ice bath for the duration of the reconstitution process. A 1.5-fold molar excess of CuT-CPP (Cu-TCPP 4,4',4'',4'''-(Porphine-5,10,15,20-tetrayl)tetrakis(benzoic acid)--Cu.sup.67) in 0.1 M NaOH must be added dropwise to the apo-globin solution in 0.2 M KPi buffer at pH 7.0 vortexed quickly at room temperature and then placed back in ice bath for 30 minutes. The protein solution must then be filtered in a syringe filter before use.
Example 8--Preparation of Transferrin-Compound Complex
[0235] The serum was obtained from healthy donor and excess iron was added in the presence of citrate ions as a chelator and bicarbonate, which is facilitates for iron binding to transferrin. The reaction mixture contained 6.5 mg sodium bicarbonate and 153.16 ferric citrate in pH=8, 4.degree. C., 1 hour per 100 mL of serum. Albumin was subsequently precipitated by Rivanol (4%) by adding the alcohol solution to the serum sample in a 3.5 V/V ratio at 4.degree. C., and pH=9.4 for 2 h. Then, the solution was centrifuged at 3000 rpm for 20 min and finally filtered by filter on a 0.8 mm syringe filter. Excess Rivanol was subsequently removed by gel-filtration on a Sephadex G-25 column in ammonium sulfate 0.025 M. A first precipitation of by saturated ammonium sulfate 50% at pH=6.5 was subsequently carried out followed by centrifugation at 3000 rpm for 10 min (immunoglobulin removal). A second precipitation at 80% saturated ammonium sulfate was then carried out thus allowing recovery of transferrin the precipitate. Solid precipitate was then dissolved in buffer of 0.06 M Tris HCl buffer, pH=8, containing 1 M NaCl. The solution was dialyzed in the same buffer to allow full removal of ammonium sulfate. Protein solution was then concentrated with a centricon PM50 centrifugal concentrator up to 10-15 mg/ml (as estimated by Bradford method) and loaded on a Sephadex G-100 gel-filtration column (2,4.times.80 cm) equilibrated in 1M NaCl, flow rate of 15 ml/h. Transferrin thus obtained was estimated to be 88-90% pure by SDS page. Ion-exchange chromatography by anion exchanger DEAE Sephadex A-50 was then used as a final polishing step. The transferrin sample was loaded in the column equilibrated with 0.06 M Tris HCl at pH=8 and eluted by a linear concentration gradient with elution buffer, 0.3 M Tris HCl, pH=8. Protein purity was higher than 98% with a yield of about 150 mg per 100 mL of serum.
[0236] Human Holo-transferrin, (SEQ ID NO: 28, FIG. 1) similarly to haemoglobin can be readily covalently linked to appropriate drug conjugates, although there is only availability for lysine modifications, due to the absence of freely titratable cysteine groups. Thus the succinimide functionalized mertansine analogue (DM1-SMCC) has been used to covalently attach to one or more lysine residues (succinimide functionalized drug). The drug has been conveniently conjugated to transferrin according to the following procedure:
[0237] The mertansin analogue DM1 SMCC (Alb Technology Ltd, Hederson, Nev., USA), functionalized for lysine covalent attachment, was prepared as follows. Transferrin solution was adjusted to a concentration of 100 .mu.M heme with reaction buffer (0.1 mM phosphate buffer pH 7.4, no EDTA in this case due to possible iron chelation effects) and conjugated with 20-fold molar excess of DM1-SMCC in the presence of 8% v/v DMSO solution at 4.degree. C. for 16 hours. The amine-reactive succinimidyl ester couples to amines thus yielding a covalent adduct with lysine groups on the surface of the protein. The excess DM1-SMCC was eliminated and buffer-exchanged with D-PBS using PM 100 ultrafiltration concentrator. The yield of conjugation was approximately 1.5 mertansine molecule per transferring dimer. Formation of DM1-SMCC conjugate was confirmed by LC-MS analysis. The concentrations of Transferrin-DM1-SMCC conjugates were determined by UV-vis spectroscopy analysis.
[0238] Alternatively, the apo-protein solution is kept in an ice bath for the duration of the reconstitution process. A 1.5-fold molar excess of CuT-CPP (Cu-TCPP 4,4',4'',4'''-(Porphine-5,10,15,20-tetrayl)tetrakis(benzoic acid)--Cu.sup.67) in 0.1 M NaOH must be added dropwise to the apo-globin solution in 0.2 M KPi buffer at pH 7.0 vortexed quickly at room temperature and then placed back in ice bath for 30 minutes. The protein solution must then be filtered in a syringe filter before use.
Example 9--Obtaining Ferritin Loaded Cells
[0239] Obtained cells are incubated in ferritin solution for a time and at the concentration sufficient to ensure proper ratio of ferritin/cell for their full load and also to ensure proper drug content to obtain therapeutic effect or contrast for proper imaging. The time and concentration may vary depending on the number of molecules encapsulated/adsorbed into the ferritin cage, status of cell activation, condition and number of their intended administration.
[0240] For example, to ensure proper load with ferritins, cells are incubated for 1-4 hours in ferritin solution 0.8 mg/ml in standard culture conditions. The frame of ferritin concentration may vary at least between 0.01 and 4 mg/ml as well as incubation time (5 min-6 hours or more). Adjusting time and concentration of ferritin load to cells, the influence of ferritin and treatment conditions on cell viability should be minded. Cells obtained as stated above very easily uptake ferritins in a relatively short time (in minutes; FIG. 8). Once they absorb ferritins, they do not release it to the culture medium (FIG. 9).
[0241] Nevertheless, the person skilled in the art is able to re-adjust the above conditions and optimize the protocol for the own purposes in the own laboratory.
Example 10--Obtaining Haemoglobin Loaded Cells
[0242] Obtained cells are incubated in haemoglobin solution for a time and at the concentration sufficient to ensure proper ratio of haemoglobin/cell for their full load and also to ensure proper drug content to obtain therapeutic effect or contrast for proper imaging. The time and concentration may vary depending on the number of molecules linked with the haemoglobin molecule, status of cell activation, condition and number of their intended administration.
[0243] For example, to ensure proper load with haemoglobins, cells are incubated for 1-4 hours in haemoglobin solution 0.1 mg/ml in standard culture conditions. The frame of haemoglobin concentration may vary at least between 0.01 and 0.2 mg/ml as well as incubation time (5 min-4 hours or more). Adjusting time and concentration of haemoglobin load to cells, the influence of ferritin and treatment conditions on cell viability should be minded. Cells obtained as stated above very easily uptake haemoglobins in a relatively short time (in minutes; FIG. 8).
[0244] Nevertheless, the person skilled in the art is able to re-adjust the above conditions and optimize the protocol for the own purposes in the own laboratory.
Example 11--Obtaining Transferrin Loaded Cells
[0245] Obtained cells are incubated in transferrin solution for a time and at the concentration sufficient to ensure proper ratio of transferrin/cell for their full load and also to ensure proper drug content to obtain therapeutic effect or contrast for proper imaging. The time and concentration may vary depending on the number of molecules linked with the transferrin molecule, status of cell activation, condition and number of their intended administration.
[0246] For example, to ensure proper load with transferrins, cells are incubated for 1-4 hours in transferrin solution 0.1 mg/ml in standard culture conditions. The frame of transferrin concentration may vary at least between 0.01 and 0.2 mg/ml as well as incubation time (5 min-4 hours or more). Adjusting time and concentration of transferrin load to cells, the influence of transferrin and treatment conditions on cell viability should be minded. Cells obtained as stated above very easily uptake transferrin in a relatively short time (in minutes; FIG. 8).
[0247] Nevertheless, the person skilled in the art is able to re-adjust the above conditions and optimize the protocol for the own purposes in the own laboratory.
Example 11--Ferritin/Haemoglobin/Transferrin-Macrophage Complex as Useful Delivery Tool to Cancer Cells
[0248] The macrophages from Example 1 prepared as described in Examples 8, 9 and 10, very easily transport ferritins, haemoglobins, transferrins to the cancer cells: mouse mammary cancer, colon cancer, canine mammary cancer, human breast, pancreatic, and bladder cancer (FIGS. 4, 10). Moreover, this transfer is much more specific to cancer cells than to non-cancer cells (FIG. 11). However, in case of cancer cells the ratio of both cell types is crucial. The more macrophages the better and faster the transport is. The most efficient transfer to the cancer cells was observed when ratio of macrophages to cancer cells was 1:1 or more.
[0249] This transfer occurred not only when the protein carriers are conjugated with fluorescent label (e.g. FITC or Alexa610), but also when they were conjugated/encapsulated with other compounds, e.g. anticancer drugs (FIG. 12 shows this transfer of ferritin encapsulated with fluorescent hypoxia activated prodrug--banoxantrone). This transfer of compounds conjugated with anticancer drugs made the effect inducing apoptosis in cancer cells (FIG. 6, 13).
Example 12--Ferritin/Haemoglobin/Transferrin-Monocyte Complex as Useful Delivery Tool to Cancer Cells
[0250] The monocytes from Example 2 prepared as described in Examples 8, 9 and 10, very easily transport ferritins, haemoglobins, transferrins to the cancer cells (FIG. 14). However, the ratio of both cell types is important. The more monocytes the better and faster the transport is. The most efficient transfer to the cancer cells was observed when ratio of monocytes to cancer cells was 1:1 or more.
Example 13--Ferritin/Haemoglobin/Transferrin-Granulocyte Complex as Useful Delivery Tool to Cancer Cells
[0251] The granulocytes from Example 3 prepared as described in Examples 8, 9 and 10, very easily transport ferritins, haemoglobins, transferrins to the cancer cells (FIG. 15). However, the ratio of both cell types is crucial. The more granulocytes the better and faster the transport is. The most efficient transfer to the cancer cells was observed when ratio of granulocytes to cancer cells was 1:1 or more.
Example 14--Ferritin/Haemoglobin/Transferrin-Lymphocyte Complex as Useful Delivery Tool to Cancer Cells
[0252] The lymphocytes from Example 4 prepared as described in Examples 8, 9 and 10, very easily transport ferritins, haemoglobins, transferrins to the cancer cells (FIG. 16). However, the ratio of both cell types is crucial. The more lymphocytes the better and faster the transport is. The most efficient transfer to the cancer cells was observed when ratio of lymphocytes to cancer cells was 1:1 or more.
Example 15--Cell Labeling with Radioisotope
[0253] In this invention cells were labeled with .sup.18F-FDG or .sup.89Zr-oxinate in order to become imaged on PET. Cells have been detached from the plate and incubated with .sup.18F-FDG or .sup.89Zr-oxinate solution in adequate concentration to ensure the most optimal .sup.18F-FDG or .sup.89Zr-oxinate uptake by cells allowing their radio-detection at the site of their accumulation. In this practice of the invention cells were incubated at room temperature for at least 90 min. in .sup.89Zr-oxinate solution containing 3-9 MBq per 5 min of cells. However, the ratio of radioisotope and cells influences significantly the reaction efficacy (FIG. 26). After labeling, cells are be centrifuged and supernatant is removed. This step should be repeated until no radioactivity is detected in the supernatant.
Example 16--Ferritin/Haemoglobin/Transferrin/Label-Leukocyte Complex as Useful Delivery Tool to Tumor, Arthritic Joints and Lymph Nodes
[0254] The macrophages from Example 1 prepared as described in Examples 9, 10, 11 and 11; monocytes from Example 2 prepared as described in Examples 9, 10, 11 and 12, granulocytes from Example 3 prepared as described in Examples 9, 10, 11 and 12, and lymphocytes from Example 4 prepared as described in Examples 9, 10, 11 and 12, highly efficiently migrate to the tumors (FIG. 22 and FIG. 23), arthritic joints (FIG. 24), in this particular example monocytic-macrophage cell line was used) and inflamed lymph nodes (FIG. 25), in this particular example monocyte/young macrophages activated using M-CSF and CCL-2 were used). Once they migrate to the tumors or other tissues they very easily transport ferritins, haemoglobins, transferrins and labels to the tumor in the enough amount to be detected using imaging systems (FIGS. 2, 3, 7, 17, and 18).
Example 17--Leukocyte-Protein Carrier Complex as Useful Targeted Pharmaceutically Active Substance Delivery System to Hypoxic Regions
[0255] Macrophages prepared as above are injected into the tail vein of animal with the tumour (appropriate number of macrophages should be adjusted to the tumour size, stage of development and presence of metastases). As it is shown on FIGS. 2, and 17 they specifically reach the tumour (after a few hrs) and also disperse in other organs of the whole animal (FIG. 18). Moreover, as it is shown on FIG. 19, in hypoxic model they are also able to migrate to the avascular and hypoxic sites and to transfer carrier proteins to cancer cells (FIGS. 3, and 20).
[0256] For the imaging purposes, 1-50 millions of macrophages were injected into the tail vein of mammary or colon cancer tumour-bearing animal. Before, macrophages were pre-labeled with Cell Tracker and loaded with ferritin-FITC (as shown in Example 8). Using two-photon of the tumour mass 8 hours after administration of macrophages the presence of macrophages carrying Ferritin-FITC was detected (FIG. 17). Their specific targeting of tumour but also their migration to other organs was shown using whole animal body imaging (IVIS) after macrophage pre-labeling using DIR cytoplasmic dye (FIG. 18).
[0257] The 1-10 millions of macrophages loaded with ferritin encapsulated cyclophosphamide, melphalan and ferritin encapsulated chlorambucil were injected i.v. into the tumour-bearing mice (300 000-500 000 of EMT6 cells injected into the skin flank). We made 3 injections of macrophages every third day (on the day 5, 8 and 11 after cancer cells injection or on the day 7, 10 and 13 after cancer cells injection) or five consecutive injections every day and we observed increased mouse survival (FIG. 5).
Example 18--Leukocyte-Protein Carrier Complex or Labeled Leukocyte as Useful Targeted Label Deliver Agent
[0258] The targeting of the targeted delivery system described in present invention can be followed by coupling the ferritin to a contrast agent. As it is presented on FIG. 21, after injection of 1-50 ml of macrophages loaded with ferritin coupled as described in Example 8 with a contrast agent (in this case: ferrihydrite, however the same results are obtained with isotope, e.g. .sup.123I) or labeled with isotope (in this case .sup.89Zr-oxide or .sup.18F-FDG) (FIG. 21) they can be easily detected by MRI, PET or SPECT. In this example (FIG. 7), mammary-tumour bearing mice were imaged using MRI at 3, 22 and 24 hours after i.v. injection of macrophages loaded with ferritin Fh. The mouse was treated (at time point 0 h) with macrophages. Then increased diameter of blood vessels (arrow) filled with injected macrophages (giving significant T2-signal reduction) has been observed and afterword macrophages spread to the tissue (spot-like pattern; arrows). These changes (in the same time points) were observed in all examined mice.
[0259] Macrophages were also labeled with .sup.89Zr-oxide or .sup.18F-FDG (5-50 min) and their radioactivity was confirmed to be imaged using PET after i.v. administration to the tumour-bearing mice. These mice were inoculated with 4T1 metastatic cell line 3 weeks before the experiment and metastases in the lungs, liver and spleen were histopatologically confirmed. At FIG. 21 it is seen that macrophages migrated to the regions with metastatic tumours allowing their visualization at PET.
[0260] While the foregoing written description of the invention enables one of ordinary skill to make and use what is considered presently to be the best mode thereof, those of ordinary skill will understand and appreciate the existence of variations, combinations, and equivalents of the specific embodiment, method, and examples herein. The invention should therefore not be limited by the above described embodiment, method, and examples, but by all embodiments and methods within the scope and spirit of the invention.
Sequence CWU 1 SEQUENCE LISTING <160> NUMBER OF SEQ ID
NOS: 28 <210> SEQ ID NO 1 <211> LENGTH: 132 <212>
TYPE: PRT <213> ORGANISM: Artificial Sequence <220>
FEATURE: <223> OTHER INFORMATION: Minimal consensus sequence
of mammalian ferritin H chain <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (5)..(5) <223>
OTHER INFORMATION: Xaa at position 5 may be present or absent, if
present it means any naturally occurring amino acid, preferably Ile
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (6)..(6) <223> OTHER INFORMATION: Xaa at position 6
means any naturally occurring amino acid, preferably Asn
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (14)..(14) <223> OTHER INFORMATION: Xaa at position
14 means any naturally occurring amino acid, preferably Tyr
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (24)..(24) <223> OTHER INFORMATION: Xaa at position
24 means any naturally occurring amino acid, preferably Tyr or Cys,
more preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (66)..(66) <223> OTHER
INFORMATION: Xaa at position 66 means any naturally occurring amino
acid, preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa at position 68 means any naturally occurring amino
acid, preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (75)..(75) <223> OTHER
INFORMATION: Xaa at position 75 means any naturally occurring amino
acid, preferably Arg or Cys, more preferably Cys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(90)..(90) <223> OTHER INFORMATION: Xaa at position 90 means
any naturally occurring amino acid, preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(94)..(94) <223> OTHER INFORMATION: Xaa at position 94 means
any naturally occurring amino acid, preferably Ser or Asn, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (120)..(120) <223> OTHER
INFORMATION: Xaa at position 120 may be present or absent, if
present it means any naturally occurring amino acid, preferably His
or Tyr, more preferably His <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Xaa at position 123 means any
naturally occurring amino acid, preferably Asn or Ser, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (128)..(128) <223> OTHER
INFORMATION: Xaa at position 128 means any naturally occurring
amino acid, preferably Ala or Ser, more preferably Ala <400>
SEQUENCE: 1 Ser Glu Ala Ala Xaa Xaa Arg Gln Ile Asn Leu Glu Leu Xaa
Ala Ser 1 5 10 15 Tyr Val Tyr Leu Ser Met Ser Xaa Tyr Phe Asp Arg
Asp Asp Val Ala 20 25 30 Leu Lys Asn Phe Ala Lys Tyr Phe Leu His
Gln Ser His Glu Glu Arg 35 40 45 Glu His Ala Glu Lys Leu Met Lys
Leu Gln Asn Gln Arg Gly Gly Arg 50 55 60 Ile Xaa Leu Xaa Asp Ile
Lys Lys Pro Asp Xaa Asp Asp Trp Glu Ser 65 70 75 80 Gly Leu Asn Ala
Met Glu Cys Ala Leu Xaa Leu Glu Lys Xaa Asn Gln 85 90 95 Ser Leu
Leu Glu Leu His Lys Leu Ala Thr Asp Lys Asn Asp Pro His 100 105 110
Leu Cys Asp Phe Ile Glu Thr Xaa Tyr Leu Xaa Glu Gln Val Lys Xaa 115
120 125 Ile Lys Glu Leu 130 <210> SEQ ID NO 2 <211>
LENGTH: 183 <212> TYPE: PRT <213> ORGANISM: Artificial
Sequence <220> FEATURE: <223> OTHER INFORMATION:
Consensus sequence of mammalian ferritin H chain <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(6)..(6) <223> OTHER INFORMATION: Xaa at position 6 can be
any naturally occurring amino acid, preferably Pro <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(14)..(14) <223> OTHER INFORMATION: Xaa at position 14 can be
any naturally occurring amino acid, preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(16)..(16) <223> OTHER INFORMATION: Xaa at position 16 can be
any naturally occurring amino acid, preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(21)..(21) <223> OTHER INFORMATION: Xaa at position 21 may be
present or absent, if present it means any naturally occurring
amino acid, preferably Ile <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (22)..(22) <223>
OTHER INFORMATION: Xaa at position 22 means any naturally occurring
amino acid, preferably Asn <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (30)..(30) <223>
OTHER INFORMATION: Xaa at position 30 can be any naturally
occurring amino acid, preferably Tyr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Xaa at position 40 can be any
naturally occurring amino acid, preferably Tyr or Cys, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (82)..(82) <223> OTHER
INFORMATION: Xaa at position 82 can be any naturally occurring
amino acid, preferably Phe <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (84)..(84) <223>
OTHER INFORMATION: Xaa at position 84 can be any naturally
occurring amino acid, preferably Gln <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (91)..(91)
<223> OTHER INFORMATION: Xaa at position 91 can be any
naturally occurring amino acid, preferably Arg or Cys, more
preferably Cys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (106)..(106) <223> OTHER
INFORMATION: Xaa at position 106 can be any naturally occurring
amino acid, preferably His <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (110)..(110)
<223> OTHER INFORMATION: Xaa at position 110 can be any
naturally occurring amino acid, preferably Asn or Ser, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (137)..(137) <223> OTHER
INFORMATION: Xaa at position 137 can be any naturally occurring
amino acid, preferably His or Tyr, more preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(140)..(140) <223> OTHER INFORMATION: Xaa at position 140 can
be any naturally occurring amino acid, preferably Asn or Ser, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (145)..(145) <223> OTHER
INFORMATION: Xaa at position 145 can be any naturally occurring
amino acid, preferably Ala or Ser, more preferably Ala <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(164)..(164) <223> OTHER INFORMATION: Xaa at position 164 can
be any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (166)..(166) <223> OTHER
INFORMATION: Xaa at position 166 can be any naturally occurring
amino acid, preferably Met or Leu, prerferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(178)..(178) <223> OTHER INFORMATION: Xaa at position 178 can
be any naturally occurring amino acid, preferably Asp or His, more
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (181)..(181) <223> OTHER
INFORMATION: Xaa at position 181 is absent or any naturally
occurring amino acid, preferably Asn <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(182)..(182) <223> OTHER INFORMATION: Xaa at position 182 is
absent or any naturally occurring amino acid, preferably Glu
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (183)..(183) <223> OTHER INFORMATION: Xaa at
position 183 is absent or any naturally occurring amino acid,
preferably Ser <400> SEQUENCE: 2 Met Thr Thr Ala Ser Xaa Ser
Gln Val Arg Gln Asn Tyr Xaa Gln Xaa 1 5 10 15 Ser Glu Ala Ala Xaa
Xaa Arg Gln Ile Asn Leu Glu Leu Xaa Ala Ser 20 25 30 Tyr Val Tyr
Leu Ser Met Ser Xaa Tyr Phe Asp Arg Asp Asp Val Ala 35 40 45 Leu
Lys Asn Phe Ala Lys Tyr Phe Leu His Gln Ser His Glu Glu Arg 50 55
60 Glu His Ala Glu Lys Leu Met Lys Leu Gln Asn Gln Arg Gly Gly Arg
65 70 75 80 Ile Xaa Leu Xaa Asp Ile Lys Lys Pro Asp Xaa Asp Asp Trp
Glu Ser 85 90 95 Gly Leu Asn Ala Met Glu Cys Ala Leu Xaa Leu Glu
Lys Xaa Val Asn 100 105 110 Gln Ser Leu Leu Glu Leu His Lys Leu Ala
Thr Asp Lys Asn Asp Pro 115 120 125 His Leu Cys Asp Phe Ile Glu Thr
Xaa Tyr Leu Xaa Glu Gln Val Lys 130 135 140 Xaa Ile Lys Glu Leu Gly
Asp His Val Thr Asn Leu Arg Lys Met Gly 145 150 155 160 Ala Pro Glu
Xaa Gly Xaa Ala Glu Tyr Leu Phe Asp Lys His Thr Leu 165 170 175 Gly
Xaa Ser Asp Xaa Xaa Xaa 180 <210> SEQ ID NO 3 <211>
LENGTH: 133 <212> TYPE: PRT <213> ORGANISM: Mus
musculus <400> SEQUENCE: 3 Ala Glu Ala Ala Ile Asn Arg Gln
Ile Asn Leu Glu Leu Tyr Ala Ser 1 5 10 15 Tyr Val Tyr Leu Ser Met
Ser Cys Tyr Phe Asp Arg Asp Asp Val Ala 20 25 30 Leu Lys Asn Phe
Ala Lys Tyr Phe Leu His Gln Ser His Glu Glu Arg 35 40 45 Glu His
Ala Glu Lys Leu Met Lys Leu Gln Asn Gln Arg Gly Gly Arg 50 55 60
Ile Phe Leu Gln Asp Ile Lys Lys Pro Asp Arg Asp Asp Trp Glu Ser 65
70 75 80 Gly Leu Asn Ala Met Glu Cys Ala Leu His Leu Glu Lys Ser
Val Asn 85 90 95 Gln Ser Leu Leu Glu Leu His Lys Leu Ala Thr Asp
Lys Asn Asp Pro 100 105 110 His Leu Cys Asp Phe Ile Glu Thr Tyr Tyr
Leu Ser Glu Gln Val Lys 115 120 125 Ser Ile Lys Glu Leu 130
<210> SEQ ID NO 4 <211> LENGTH: 180 <212> TYPE:
PRT <213> ORGANISM: Mus musculus <400> SEQUENCE: 4 Met
Thr Thr Ala Ser Pro Ser Gln Val Arg Gln Asn Tyr His Gln Asp 1 5 10
15 Ala Glu Ala Ala Ile Asn Arg Gln Ile Asn Leu Glu Leu Tyr Ala Ser
20 25 30 Tyr Val Tyr Leu Ser Met Ser Cys Tyr Phe Asp Arg Asp Asp
Val Ala 35 40 45 Leu Lys Asn Phe Ala Lys Tyr Phe Leu His Gln Ser
His Glu Glu Arg 50 55 60 Glu His Ala Glu Lys Leu Met Lys Leu Gln
Asn Gln Arg Gly Gly Arg 65 70 75 80 Ile Phe Leu Gln Asp Ile Lys Lys
Pro Asp Arg Asp Asp Trp Glu Ser 85 90 95 Gly Leu Asn Ala Met Glu
Cys Ala Leu His Leu Glu Lys Ser Val Asn 100 105 110 Gln Ser Leu Leu
Glu Leu His Lys Leu Ala Thr Asp Lys Asn Asp Pro 115 120 125 His Leu
Cys Asp Phe Ile Glu Thr Tyr Tyr Leu Ser Glu Gln Val Lys 130 135 140
Ser Ile Lys Glu Leu Gly Asp His Val Thr Asn Leu Arg Lys Met Gly 145
150 155 160 Ala Pro Glu Ala Gly Met Ala Glu Tyr Leu Phe Asp Lys His
Thr Leu 165 170 175 Gly His Gly Asp 180 <210> SEQ ID NO 5
<211> LENGTH: 132 <212> TYPE: PRT <213> ORGANISM:
Homo sapiens <400> SEQUENCE: 5 Ser Glu Ala Ala Asn Arg Gln
Ile Asn Leu Glu Leu Tyr Ala Ser Tyr 1 5 10 15 Val Tyr Leu Ser Met
Ser Tyr Tyr Phe Asp Arg Asp Asp Val Ala Leu 20 25 30 Lys Asn Phe
Ala Lys Tyr Phe Leu His Gln Ser His Glu Glu Arg Glu 35 40 45 His
Ala Glu Lys Leu Met Lys Leu Gln Asn Gln Arg Gly Gly Arg Ile 50 55
60 Phe Leu Gln Asp Ile Lys Lys Pro Asp Cys Asp Asp Trp Glu Ser Gly
65 70 75 80 Leu Asn Ala Met Glu Cys Ala Leu His Leu Glu Lys Asn Val
Asn Gln 85 90 95 Ser Leu Leu Glu Leu His Lys Leu Ala Thr Asp Lys
Asn Asp Pro His 100 105 110 Leu Cys Asp Phe Ile Glu Thr His Tyr Leu
Asn Glu Gln Val Lys Ala 115 120 125 Ile Lys Glu Leu 130 <210>
SEQ ID NO 6 <211> LENGTH: 179 <212> TYPE: PRT
<213> ORGANISM: Homo sapiens <400> SEQUENCE: 6 Met Thr
Thr Ala Ser Thr Ser Gln Val Arg Gln Asn Tyr His Gln Asp 1 5 10 15
Ser Glu Ala Ala Asn Arg Gln Ile Asn Leu Glu Leu Tyr Ala Ser Tyr 20
25 30 Val Tyr Leu Ser Met Ser Tyr Tyr Phe Asp Arg Asp Asp Val Ala
Leu 35 40 45 Lys Asn Phe Ala Lys Tyr Phe Leu His Gln Ser His Glu
Glu Arg Glu 50 55 60 His Ala Glu Lys Leu Met Lys Leu Gln Asn Gln
Arg Gly Gly Arg Ile 65 70 75 80 Phe Leu Gln Asp Ile Lys Lys Pro Asp
Cys Asp Asp Trp Glu Ser Gly 85 90 95 Leu Asn Ala Met Glu Cys Ala
Leu His Leu Glu Lys Asn Val Asn Gln 100 105 110 Ser Leu Leu Glu Leu
His Lys Leu Ala Thr Asp Lys Asn Asp Pro His 115 120 125 Leu Cys Asp
Phe Ile Glu Thr His Tyr Leu Asn Glu Gln Val Lys Ala 130 135 140 Ile
Lys Glu Leu Gly Asp His Val Thr Asn Leu Arg Lys Met Gly Ala 145 150
155 160 Pro Glu Ser Gly Leu Ala Glu Tyr Leu Phe Asp Lys His Thr Leu
Gly 165 170 175 Asp Ser Asp <210> SEQ ID NO 7 <211>
LENGTH: 182 <212> TYPE: PRT <213> ORGANISM: Artificial
Sequence <220> FEATURE: <223> OTHER INFORMATION:
Consensus sequence of mammalian ferritin H chains 2nd version
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (6)..(6) <223> OTHER INFORMATION: Xaa at position 6
can be any naturally occurring amino acid, preferably Pro
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (14)..(14) <223> OTHER INFORMATION: Xaa at position
14 can be any naturally occurring amino acid, preferably His
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (16)..(16) <223> OTHER INFORMATION: Xaa at position
16 can be any naturally occurring amino acid, preferably Asp
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (21)..(21) <223> OTHER INFORMATION: Xaa at position
21 may be present or absent, if present it means any naturally
occurring amino acid, preferably Ile <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (29)..(29)
<223> OTHER INFORMATION: Xaa at position 29 can be any
naturally occurring amino acid, preferably Tyr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (81)..(81)
<223> OTHER INFORMATION: Xaa at position 81 can be any
naturally occurring amino acid, preferably Phe <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (83)..(83)
<223> OTHER INFORMATION: Xaa at position 83 can be any
naturally occurring amino acid, preferably Gln <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(105)..(105) <223> OTHER INFORMATION: Xaa at position 105 can
be any naturally occurring amino acid, preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(144)..(144) <223> OTHER INFORMATION: Xaa at position 144 can
be any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (180)..(180) <223> OTHER
INFORMATION: Xaa at position 180 is absent or any naturally
occuring amino acid, preferably Asn <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(181)..(181) <223> OTHER INFORMATION: Xaa at position 181 is
absent or any naturally occuring amino acid, preferably Glu
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (182)..(182) <223> OTHER INFORMATION: Xaa at
position 182 is absent or any naturally occuring amino acid,
preferably Ser <400> SEQUENCE: 7 Met Thr Thr Ala Ser Xaa Ser
Gln Val Arg Gln Asn Tyr Xaa Gln Xaa 1 5 10 15 Ser Glu Ala Ala Xaa
Arg Gln Ile Asn Leu Glu Leu Xaa Ala Ser Tyr 20 25 30 Val Tyr Leu
Ser Met Ser Tyr Tyr Phe Asp Arg Asp Asp Val Ala Leu 35 40 45 Lys
Asn Phe Ala Lys Tyr Phe Leu His Gln Ser His Glu Glu Arg Glu 50 55
60 His Ala Glu Lys Leu Met Lys Leu Gln Asn Gln Arg Gly Gly Arg Ile
65 70 75 80 Xaa Leu Xaa Asp Ile Lys Lys Pro Asp Cys Asp Asp Trp Glu
Ser Gly 85 90 95 Leu Asn Ala Met Glu Cys Ala Leu Xaa Leu Glu Lys
Asn Val Asn Gln 100 105 110 Ser Leu Leu Glu Leu His Lys Leu Ala Thr
Asp Lys Asn Asp Pro His 115 120 125 Leu Cys Asp Phe Ile Glu Thr His
Tyr Leu Asn Glu Gln Val Lys Xaa 130 135 140 Ile Lys Glu Leu Gly Asp
His Val Thr Asn Leu Arg Lys Met Gly Ala 145 150 155 160 Pro Glu Ser
Gly Leu Ala Glu Tyr Leu Phe Asp Lys His Thr Leu Gly 165 170 175 Asp
Ser Asp Xaa Xaa Xaa 180 <210> SEQ ID NO 8 <211> LENGTH:
150 <212> TYPE: PRT <213> ORGANISM: Artificial Sequence
<220> FEATURE: <223> OTHER INFORMATION: Minimal
consensus sequence of mammalian ferritin L chain <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(5)..(5) <223> OTHER INFORMATION: Xaa at position 5 can be
any naturally occurring amino acid, preferably Asp or Glu, more
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (12)..(12) <223> OTHER
INFORMATION: Xaa at position 12 can be any nnaturally occurring
amino acid, preferably Arg or Ser, more preferably Ser <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(17)..(17) <223> OTHER INFORMATION: Xaa at position 17 can be
any naturally occurring amino acid, preferably Ser or Arg, more
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (19)..(19) <223> OTHER
INFORMATION: Xaa at position 19 can be any naturally occurring
amino acid, preferably Arg or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(29)..(29) <223> OTHER INFORMATION: Xaa at position 29 can be
any naturally occurring amino acid, preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(30)..(30) <223> OTHER INFORMATION: Xaa at position 30 can be
any naturally occurring amino acid, preferably Tyr or Phe, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (42)..(42) <223> OTHER
INFORMATION: Xaa at position 42 can be any naturally occurring
amino acid, preferably Ser or Gly, more preferably Ser <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(56)..(56) <223> OTHER INFORMATION: Xaa at position 56 can be
any naturally occurirng amino acid, preferably Ala or Tyr, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (61)..(61) <223> OTHER
INFORMATION: Xaa at position 61 can be any naturally occurring
amino acid, preferably Glu or Lys, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(62)..(62) <223> OTHER INFORMATION: Xaa at position 62 can be
any naturally occurring amino acid, preferably Met or Phe, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa at position 65 can be any naturally occurring
amino acid, preferably Asp or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(75)..(75) <223> OTHER INFORMATION: Xaa at position 75 can be
any naturally occurring amino acid, preferably Ile or Val, more
preferably Ile <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (76)..(76) <223> OTHER
INFORMATION: Xaa at position 76 can be any naturally occurring
amino acid, preferably Lys or Gln, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(79)..(79) <223> OTHER INFORMATION: Xaa at position 79 can be
any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (80)..(80) <223> OTHER
INFORMATION: Xaa at position 80 can be any naturally occurring
amino acid, preferably Glu or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(87)..(87) <223> OTHER INFORMATION: Xaa at position 87 can be
any naturally occurring amino acid, preferably Pro or Gln, more
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (88)..(88) <223> OTHER
INFORMATION: Xaa at position 88 can be any naturally occurring
amino acid, preferably Glu or Asp, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(91)..(91) <223> OTHER INFORMATION: Xaa at position 91 can be
any naturally occurring amino acid, preferably Glu or Lys, more
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (94)..(94) <223> OTHER
INFORMATION: Xaa at position 94 can be any naturally occurring
amino acid, preferably Met or Leu, more preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(96)..(96) <223> OTHER INFORMATION: Xaa at position 96 can be
any naturally occurring amino acid, preferably Met or Leu, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (99)..(99) <223> OTHER
INFORMATION: Xaa at position 99 can be any naturally occurring
amino acid, preferably Lys or Asn, preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(115)..(115) <223> OTHER INFORMATION: Xaa at position 115 can
be any naturally occurring amino acid, preferably Thr or Ala, more
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (119)..(119) <223> OTHER
INFORMATION: Xaa at position 119 can be any naturally occurring
amino acid, preferably Leu <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (125)..(125)
<223> OTHER INFORMATION: Xaa at position 125 can be any
naturally occurring amino acid, preferably Ser or Thr, more
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (127)..(127) <223> OTHER
INFORMATION: Xaa at position 127 can be any naturally occurring
amino acid, preferably Tyr or Phe, more preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(130)..(130) <223> OTHER INFORMATION: Xaa at position 130 can
be any naturally occurring amino acid, preferably Lys or Glu, more
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (140)..(140) <223> OTHER
INFORMATION: Xaa at position 140 can be any naturally occurring
amino acid, preferably Asp or Asn, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(146)..(146) <223> OTHER INFORMATION: Xaa at position 146 can
be any naturally occurring amino acid, preferably Arg or His, more
preferably His <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (148)..(148) <223> OTHER
INFORMATION: Xaa at position 148 can be any naturally occurring
amino acid, preferably Leu or Val, more preferably Leu <400>
SEQUENCE: 8 Asn Tyr Ser Thr Xaa Val Glu Ala Ala Val Asn Xaa Leu Val
Asn Leu 1 5 10 15 Xaa Leu Xaa Ala Ser Tyr Thr Tyr Leu Ser Leu Gly
Xaa Xaa Phe Asp 20 25 30 Arg Asp Asp Val Ala Leu Glu Gly Val Xaa
His Phe Phe Arg Glu Leu 35 40 45 Ala Glu Glu Lys Arg Glu Gly Xaa
Glu Arg Leu Leu Xaa Xaa Gln Asn 50 55 60 Xaa Arg Gly Gly Arg Ala
Leu Phe Gln Asp Xaa Xaa Lys Pro Xaa Xaa 65 70 75 80 Asp Glu Trp Gly
Lys Thr Xaa Xaa Ala Met Xaa Ala Ala Xaa Ala Xaa 85 90 95 Glu Lys
Xaa Leu Asn Gln Ala Leu Leu Asp Leu His Ala Leu Gly Ser 100 105 110
Ala Arg Xaa Asp Pro His Xaa Cys Asp Phe Leu Glu Xaa His Xaa Leu 115
120 125 Asp Xaa Glu Val Lys Leu Ile Lys Lys Met Gly Xaa His Leu Thr
Asn 130 135 140 Leu Xaa Arg Xaa Ala Gly 145 150 <210> SEQ ID
NO 9 <211> LENGTH: 183 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Consensus sequence of mammalian ferritin L chain
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (12)..(12) <223> OTHER INFORMATION: Xaa at position
12 can be any naturally occurring amino acid, preferably Asp or
Glu, more preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (19)..(19) <223> OTHER
INFORMATION: Xaa at position 19 can be any naturally occurring
amino acid, preferably Ser or Arg, more preferably Ser <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(24)..(24) <223> OTHER INFORMATION: Xaa at position 24 can be
any naturally occurring amino acid, preferably Ser or Arg, more
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (26)..(26) <223> OTHER
INFORMATION: Xaa at position 26 can be any naturally occurring
amino acid, preferably Arg or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(36)..(36) <223> OTHER INFORMATION: Xaa at position 36 can be
any naturally occurring amino acid, preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(37)..(37) <223> OTHER INFORMATION: Xaa at position 37 can be
any naturally occurring amino acid, preferably Tyr or Phe, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (49)..(49) <223> OTHER
INFORMATION: Xaa at position 47 can be any naturally occurring
amino acid, preferably Ser or Gly, more preferably Ser <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(63)..(63) <223> OTHER INFORMATION: Xaa at position 63 can be
any naturally occurring amino acid, preferably Ala or Tyr, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa at position 68 can be any naturally occurring
amino acid, preferably Glu or Lys, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(69)..(69) <223> OTHER INFORMATION: Xaa at position 69 can be
any naturally occurring amino acid, preferably Met or Phe, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (72)..(72) <223> OTHER
INFORMATION: Xaa at position 72 can be any naturally occurring
amino acid, preferably Asp or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(82)..(82) <223> OTHER INFORMATION: Xaa at position 82 can be
any naturally occurring amino acid, preferably Ile or Val, more
preferably Ile <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (83)..(83) <223> OTHER
INFORMATION: Xaa at position 83 can be any naturally occurring
amino acid, preferably Lys or Gln, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(86)..(86) <223> OTHER INFORMATION: Xaa at position 86 can be
any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (87)..(87) <223> OTHER
INFORMATION: Xaa at position 87 can be any naturally occurring
amino acid, preferably Glu or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(94)..(94) <223> OTHER INFORMATION: Xaa at position 94 can be
any naturally occurring amino acid, preferably Pro or Gln, more
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (95)..(95) <223> OTHER
INFORMATION: Xaa at position 95 can be any naturally occurring
amino acid, preferably Glu or Asp, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(98)..(98) <223> OTHER INFORMATION: Xaa at position 98 can be
any naturally occurring amino acid, preferably Glu or Lys, more
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (101)..(101) <223> OTHER
INFORMATION: Xaa at position 101 can be any naturally occurring
amino acid, preferably Met or Leu, more preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(103)..(103) <223> OTHER INFORMATION: Xaa at position 103 can
be any naturally occurring amino acid, preferably Met or Leu, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (106)..(106) <223> OTHER
INFORMATION: Xaa at position 106 can be any naturally occurring
amino acid, preferably Lys or Asn, preferably Lys <220>
FEATURE: <221> NAME/KEY: misc_feature <222> LOCATION:
(122)..(122) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (126)..(126)
<223> OTHER INFORMATION: Xaa at position 126 can be any
naturally occurring amino acid, preferably Leu <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(132)..(132) <223> OTHER INFORMATION: Xaa at position 132 can
be any naturally occurring amino acid, preferably Ser or Thr, more
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (134)..(134) <223> OTHER
INFORMATION: Xaa at position 134 can be any naturally occurring
amino acid, preferably Tyr or Phe, more preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(137)..(137) <223> OTHER INFORMATION: Xaa at position 137 can
be any naturally occurring amino acid, preferably Lys or Glu, more
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (147)..(147) <223> OTHER
INFORMATION: Xaa at position 147 can be any naturally occurring
amino acid, preferably Asp or Asn, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(153)..(153) <223> OTHER INFORMATION: Xaa at position 153 can
be any naturally occurring amino acid, preferably Arg or His, more
preferably His <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (155)..(155) <223> OTHER
INFORMATION: Xaa at position 155 can be any naturally occurring
amino acid, preferably Leu or Val, more preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(161)..(161) <223> OTHER INFORMATION: Xaa at position 161 can
be absent or any naturally occurring amino acid, preferably Ala
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (163)..(163) <223> OTHER INFORMATION: Xaa at
position 163 can be absent or any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (166)..(166) <223> OTHER
INFORMATION: Xaa at position 166 can be absent or any naturally
occurring amino acid, preferably Pro <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(168)..(168) <223> OTHER INFORMATION: Xaa at position 168 can
be any naturally occurring amino acid, preferably Gly or Ala, more
preferably Ala <400> SEQUENCE: 9 Met Thr Ser Gln Ile Arg Gln
Asn Tyr Ser Thr Xaa Val Glu Ala Ala 1 5 10 15 Val Asn Xaa Leu Val
Asn Leu Xaa Leu Xaa Ala Ser Tyr Thr Tyr Leu 20 25 30 Ser Leu Gly
Xaa Xaa Phe Asp Arg Asp Asp Val Ala Leu Glu Gly Val 35 40 45 Xaa
His Phe Phe Arg Glu Leu Ala Glu Glu Lys Arg Glu Gly Xaa Glu 50 55
60 Arg Leu Leu Xaa Xaa Gln Asn Xaa Arg Gly Gly Arg Ala Leu Phe Gln
65 70 75 80 Asp Xaa Xaa Lys Pro Xaa Xaa Asp Glu Trp Gly Lys Thr Xaa
Xaa Ala 85 90 95 Met Xaa Ala Ala Xaa Ala Xaa Glu Lys Xaa Leu Asn
Gln Ala Leu Leu 100 105 110 Asp Leu His Ala Leu Gly Ser Ala Arg Xaa
Asp Pro His Xaa Cys Asp 115 120 125 Phe Leu Glu Xaa His Xaa Leu Asp
Xaa Glu Val Lys Leu Ile Lys Lys 130 135 140 Met Gly Xaa His Leu Thr
Asn Leu Xaa Arg Xaa Ala Gly Pro Gln Pro 145 150 155 160 Xaa Gln Xaa
Gly Val Xaa Gln Xaa Ser Leu Gly Glu Tyr Leu Phe Glu 165 170 175 Arg
Leu Thr Leu Lys His Asp 180 <210> SEQ ID NO 10 <211>
LENGTH: 150 <212> TYPE: PRT <213> ORGANISM: Mus
musculus <220> FEATURE: <221> NAME/KEY: MISC_FEATURE
<222> LOCATION: (12)..(12) <223> OTHER INFORMATION: Xaa
at position 5 can be any naturally occuring amino acid, preferably
Asp or Glu, more preferably Asp <400> SEQUENCE: 10 Asn Tyr
Ser Thr Glu Val Glu Ala Ala Val Asn Arg Leu Val Asn Leu 1 5 10 15
His Leu Arg Ala Ser Tyr Thr Tyr Leu Ser Leu Gly Phe Phe Phe Asp 20
25 30 Arg Asp Asp Val Ala Leu Glu Gly Val Gly His Phe Phe Arg Glu
Leu 35 40 45 Ala Glu Glu Lys Arg Glu Gly Ala Glu Arg Leu Leu Glu
Phe Gln Asn 50 55 60 Asp Arg Gly Gly Arg Ala Leu Phe Gln Asp Val
Gln Lys Pro Ser Gln 65 70 75 80 Asp Glu Trp Gly Lys Thr Gln Glu Ala
Met Glu Ala Ala Leu Ala Met 85 90 95 Glu Lys Asn Leu Asn Gln Ala
Leu Leu Asp Leu His Ala Leu Gly Ser 100 105 110 Ala Arg Ala Asp Pro
His Leu Cys Asp Phe Leu Glu Ser His Tyr Leu 115 120 125 Asp Lys Glu
Val Lys Leu Ile Lys Lys Met Gly Asn His Leu Thr Asn 130 135 140 Leu
Arg Arg Val Ala Gly 145 150 <210> SEQ ID NO 11 <400>
SEQUENCE: 11 000 <210> SEQ ID NO 12 <211> LENGTH: 149
<212> TYPE: PRT <213> ORGANISM: Homo sapiens
<400> SEQUENCE: 12 Asn Tyr Ser Thr Asp Val Glu Ala Ala Val
Asn Ser Leu Val Asn Leu 1 5 10 15 Tyr Leu Gln Ala Ser Tyr Thr Tyr
Leu Ser Leu Gly Phe Tyr Phe Asp 20 25 30 Arg Asp Asp Val Ala Leu
Glu Gly Val Ser His Phe Phe Arg Glu Leu 35 40 45 Ala Glu Glu Lys
Arg Glu Gly Tyr Glu Arg Leu Leu Lys Met Gln Asn 50 55 60 Gln Arg
Gly Gly Arg Ala Leu Phe Gln Asp Ile Lys Lys Pro Ala Glu 65 70 75 80
Asp Glu Trp Gly Lys Thr Pro Asp Ala Met Lys Ala Ala Met Ala Leu 85
90 95 Glu Lys Lys Leu Asn Gln Ala Leu Leu Asp Leu His Ala Leu Gly
Ser 100 105 110 Ala Arg Thr Asp Pro His Leu Cys Asp Phe Leu Glu Thr
His Phe Leu 115 120 125 Asp Glu Glu Val Lys Leu Ile Lys Lys Met Gly
Asp His Leu Thr Asn 130 135 140 Leu His Arg Leu Gly 145 <210>
SEQ ID NO 13 <211> LENGTH: 175 <212> TYPE: PRT
<213> ORGANISM: Homo sapiens <400> SEQUENCE: 13 Met Ser
Ser Gln Ile Arg Gln Asn Tyr Ser Thr Asp Val Glu Ala Ala 1 5 10 15
Val Asn Ser Leu Val Asn Leu Tyr Leu Gln Ala Ser Tyr Thr Tyr Leu 20
25 30 Ser Leu Gly Phe Tyr Phe Asp Arg Asp Asp Val Ala Leu Glu Gly
Val 35 40 45 Ser His Phe Phe Arg Glu Leu Ala Glu Glu Lys Arg Glu
Gly Tyr Glu 50 55 60 Arg Leu Leu Lys Met Gln Asn Gln Arg Gly Gly
Arg Ala Leu Phe Gln 65 70 75 80 Asp Ile Lys Lys Pro Ala Glu Asp Glu
Trp Gly Lys Thr Pro Asp Ala 85 90 95 Met Lys Ala Ala Met Ala Leu
Glu Lys Lys Leu Asn Gln Ala Leu Leu 100 105 110 Asp Leu His Ala Leu
Gly Ser Ala Arg Thr Asp Pro His Leu Cys Asp 115 120 125 Phe Leu Glu
Thr His Phe Leu Asp Glu Glu Val Lys Leu Ile Lys Lys 130 135 140 Met
Gly Asp His Leu Thr Asn Leu His Arg Leu Gly Gly Pro Glu Ala 145 150
155 160 Gly Leu Gly Glu Tyr Leu Phe Glu Arg Leu Thr Leu Lys His Asp
165 170 175 <210> SEQ ID NO 14 <211> LENGTH: 183
<212> TYPE: PRT <213> ORGANISM: Artificial Sequence
<220> FEATURE: <223> OTHER INFORMATION: Consensus
seqeuence of mammaliam ferritin L chain 2nd version <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(36)..(36) <223> OTHER INFORMATION: Xaa at position 36 can be
any naturally occuring amino acid, preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(37)..(37) <223> OTHER INFORMATION: Xaa at position 37 can be
any naturally occuring amino acid, preferably Tyr or Phe, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (94)..(94) <223> OTHER
INFORMATION: Xaa at position 94 can be any naturally occurring
amino acid, preferably Pro or Gln, more preferably Pro <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(126)..(126) <223> OTHER INFORMATION: Xaa at position 126 can
be any naturally occurring amino acid, preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(137)..(137) <223> OTHER INFORMATION: Xaa at position 137 can
be any naturally occurring amino acid, preferably Lys or Glu, more
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (147)..(147) <223> OTHER
INFORMATION: Xaa at position 147 can be any naturally occurring
amino acid, preferably Asp or Asn, more preferably Asp <220>
FEATURE: <221> NAME/KEY: misc_feature <222> LOCATION:
(161)..(161) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (163)..(163)
<223> OTHER INFORMATION: Xaa at position 163 can be absent or
any naturally occurring amino acid, preferably Thr <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(166)..(166) <223> OTHER INFORMATION: Xaa at position 166 can
be absent or any naturally occurring amino acid, preferably Pro
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (168)..(168) <223> OTHER INFORMATION: Xaa at
position 168 can be any naturally occurring amino acid, preferably
Gly or Ala, more preferably Ala <400> SEQUENCE: 14 Met Thr
Ser Gln Ile Arg Gln Asn Tyr Ser Thr Glu Val Glu Ala Ala 1 5 10 15
Val Asn Arg Leu Val Asn Leu His Leu Arg Ala Ser Tyr Thr Tyr Leu 20
25 30 Ser Leu Gly Xaa Xaa Phe Asp Arg Asp Asp Val Ala Leu Glu Gly
Val 35 40 45 Gly His Phe Phe Arg Glu Leu Ala Glu Glu Lys Arg Glu
Gly Ala Glu 50 55 60 Arg Leu Leu Lys Leu Gln Asn Gln Arg Gly Gly
Arg Ala Leu Phe Gln 65 70 75 80 Asp Val Gln Lys Pro Ala Gln Asp Glu
Trp Gly Lys Thr Xaa Glu Ala 85 90 95 Met Glu Ala Ala Leu Ala Leu
Glu Lys Asn Leu Asn Gln Ala Leu Leu 100 105 110 Asp Leu His Ala Leu
Gly Ser Ala Arg Thr Asp Pro His Xaa Cys Asp 115 120 125 Phe Leu Glu
Asn His Phe Leu Asp Xaa Glu Val Lys Leu Ile Lys Lys 130 135 140 Met
Gly Xaa His Leu Thr Asn Leu Arg Arg Leu Ala Gly Pro Gln Pro 145 150
155 160 Xaa Gln Xaa Gly Val Xaa Gln Xaa Ser Leu Gly Glu Tyr Leu Phe
Glu 165 170 175 Arg Leu Thr Leu Lys His Asp 180 <210> SEQ ID
NO 15 <211> LENGTH: 117 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Minimal consensus sequence of mammalian
haemoglobin alpha chain <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (5)..(5) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (9)..(9) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (21)..(21) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (22)..(22) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (38)..(38) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (41)..(41) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (48)..(48) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (49)..(49) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (55)..(55) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (59)..(59) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Aap <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (60)..(60) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (76)..(76) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (79)..(79) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (90)..(90) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (91)..(91) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (94)..(94) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (102)..(102) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (108)..(108) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (117)..(117) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <400> SEQUENCE: 15 Val Gly Ala His Xaa Gly Glu
Tyr Xaa Ala Glu Ala Leu Glu Xaa Met 1 5 10 15 Phe Leu Ser Phe Xaa
Xaa Thr Lys Thr Tyr Phe Pro His Phe Asp Leu 20 25 30 Ser His Gly
Ser Ala Xaa Val Lys Xaa His Gly Lys Lys Val Ala Xaa 35 40 45 Xaa
Leu Thr Asn Ala Val Xaa His Val Asp Xaa Xaa Pro Asn Ala Leu 50 55
60 Ser Ala Leu Xaa Asp Leu His Ala His Lys Leu Xaa Val Asp Xaa Val
65 70 75 80 Asn Phe Lys Leu Leu Ser His Cys Leu Xaa Xaa Thr Leu Xaa
Ala His 85 90 95 Leu Pro Ala Glu Phe Xaa Pro Ala Val His Ala Xaa
Leu Asp Lys Phe 100 105 110 Leu Ala Ser Val Xaa 115 <210> SEQ
ID NO 16 <211> LENGTH: 142 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Consensus sequence of mammalian alpha chain
haemoglobins <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (1)..(1) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (2)..(2) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (14)..(14) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (17)..(17) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (22)..(22) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (26)..(26) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (32)..(32) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (38)..(38) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (39)..(39) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (55)..(55) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (58)..(58) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (66)..(66) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (72)..(72) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (76)..(76) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (77)..(77) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (85)..(85) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (93)..(93) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (96)..(96) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (107)..(107) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (108)..(108) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (111)..(111) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (119)..(119) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (125)..(125) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (134)..(134) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (142)..(142) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <400> SEQUENCE: 16 Xaa Xaa Leu Ser Pro Ala Asp
Lys Thr Asn Val Lys Ala Xaa Trp Gly 1 5 10 15 Xaa Val Gly Ala His
Xaa Gly Glu Tyr Xaa Ala Glu Ala Leu Glu Xaa 20 25 30 Met Phe Leu
Ser Phe Xaa Xaa Thr Lys Thr Tyr Phe Pro His Phe Asp 35 40 45 Leu
Ser His Gly Ser Ala Xaa Val Lys Xaa His Gly Lys Lys Val Ala 50 55
60 Xaa Xaa Leu Thr Asn Ala Val Xaa His Val Asp Xaa Xaa Pro Asn Ala
65 70 75 80 Leu Ser Ala Leu Xaa Asp Leu His Ala His Lys Leu Xaa Val
Asp Xaa 85 90 95 Val Asn Phe Lys Leu Leu Ser His Cys Leu Xaa Xaa
Thr Leu Xaa Ala 100 105 110 His Leu Pro Ala Glu Phe Xaa Pro Ala Val
His Ala Xaa Leu Asp Lys 115 120 125 Phe Leu Ala Ser Val Xaa Thr Val
Leu Thr Ser Lys Tyr Xaa 130 135 140 <210> SEQ ID NO 17
<211> LENGTH: 117 <212> TYPE: PRT <213> ORGANISM:
Homo sapiens <400> SEQUENCE: 17 Val Gly Ala His Ala Gly Glu
Tyr Gly Ala Glu Ala Leu Glu Arg Met 1 5 10 15 Phe Leu Ser Phe Pro
Thr Thr Lys Thr Tyr Phe Pro His Phe Asp Leu 20 25 30 Ser His Gly
Ser Ala Gln Val Lys Gly His Gly Lys Lys Val Ala Asp 35 40 45 Ala
Leu Thr Asn Ala Val Ala His Val Asp Asp Met Pro Asn Ala Leu 50 55
60 Ser Ala Leu Ser Asp Leu His Ala His Lys Leu Arg Val Asp Pro Val
65 70 75 80 Asn Phe Lys Leu Leu Ser His Cys Leu Leu Val Thr Leu Ala
Ala His 85 90 95 Leu Pro Ala Glu Phe Thr Pro Ala Val His Ala Ser
Leu Asp Lys Phe 100 105 110 Leu Ala Ser Val Ser 115 <210> SEQ
ID NO 18 <211> LENGTH: 142 <212> TYPE: PRT <213>
ORGANISM: Homo sapiens <400> SEQUENCE: 18 Met Val Leu Ser Pro
Ala Asp Lys Thr Asn Val Lys Ala Ala Trp Gly 1 5 10 15 Lys Val Gly
Ala His Ala Gly Glu Tyr Gly Ala Glu Ala Leu Glu Arg 20 25 30 Met
Phe Leu Ser Phe Pro Thr Thr Lys Thr Tyr Phe Pro His Phe Asp 35 40
45 Leu Ser His Gly Ser Ala Gln Val Lys Gly His Gly Lys Lys Val Ala
50 55 60 Asp Ala Leu Thr Asn Ala Val Ala His Val Asp Asp Met Pro
Asn Ala 65 70 75 80 Leu Ser Ala Leu Ser Asp Leu His Ala His Lys Leu
Arg Val Asp Pro 85 90 95 Val Asn Phe Lys Leu Leu Ser His Cys Leu
Leu Val Thr Leu Ala Ala 100 105 110 His Leu Pro Ala Glu Phe Thr Pro
Ala Val His Ala Ser Leu Asp Lys 115 120 125 Phe Leu Ala Ser Val Ser
Thr Val Leu Thr Ser Lys Tyr Arg 130 135 140 <210> SEQ ID NO
19 <211> LENGTH: 124 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Minimal consensus sequence of mammalian
haemoglobin beta chain <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (4)..(4) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (9)..(9) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (10)..(10) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (12)..(12) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (13)..(13) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (14)..(14) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (27)..(27) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (35)..(35) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (36)..(36) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (49)..(49) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (52)..(52) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (66)..(66) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (69)..(69) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (86)..(86) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Glu <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (121)..(121) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (122)..(122) <223> OTHER INFORMATION: Xaa can be
absent or any naturally occurring amino acid, preferably Val
<400> SEQUENCE: 19 Gly Lys Val Xaa Val Asp Glu Val Xaa Xaa
Glu Xaa Xaa Xaa Xaa Leu 1 5 10 15 Leu Val Val Tyr Pro Trp Thr Gln
Arg Phe Xaa Glu Ser Phe Gly Asp 20 25 30 Leu Ser Xaa Xaa Asp Ala
Val Met Gly Asn Pro Lys Val Lys Ala His 35 40 45 Xaa Lys Lys Xaa
Leu Gly Ala Phe Ser Asp Gly Leu Ala His Leu Asp 50 55 60 Asn Xaa
Lys Xaa Xaa Phe Ala Thr Leu Ser Glu Leu His Cys Asp Lys 65 70 75 80
Leu His Val Asp Pro Xaa Asn Phe Arg Leu Leu Gly Asn Val Leu Val 85
90 95 Cys Val Leu Ala His His Phe Gly Lys Glu Phe Thr Pro Pro Val
Gln 100 105 110 Ala Ala Tyr Gln Lys Val Val Ala Xaa Xaa Ala Asn 115
120 <210> SEQ ID NO 20 <211> LENGTH: 147 <212>
TYPE: PRT <213> ORGANISM: Artificial Sequence <220>
FEATURE: <223> OTHER INFORMATION: Consensus sequence of
mammalian haemoglobin beta chain <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (10)..(10) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (11)..(11) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (20)..(20) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (25)..(25) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (26)..(26) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (28)..(28) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (29)..(29) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (30)..(30) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (31)..(31) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (43)..(43) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (51)..(51) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (52)..(52) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (82)..(82) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (84)..(84) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (85)..(85) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (102)..(102) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (137)..(137) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (138)..(138) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (147)..(147) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably His <400> SEQUENCE: 20 Met Val His Leu Thr Pro Glu
Glu Lys Xaa Xaa Val Thr Ala Xaa Trp 1 5 10 15 Gly Lys Val Xaa Val
Asp Glu Val Xaa Xaa Glu Xaa Xaa Xaa Xaa Leu 20 25 30 Leu Val Val
Tyr Pro Trp Thr Gln Arg Phe Xaa Glu Ser Phe Gly Asp 35 40 45 Leu
Ser Xaa Xaa Asp Ala Val Met Gly Asn Pro Lys Val Lys Ala His 50 55
60 Xaa Lys Lys Xaa Leu Gly Ala Phe Ser Asp Gly Leu Ala His Leu Asp
65 70 75 80 Asn Xaa Lys Xaa Xaa Phe Ala Thr Leu Ser Glu Leu His Cys
Asp Lys 85 90 95 Leu His Val Asp Pro Xaa Asn Phe Arg Leu Leu Gly
Asn Val Leu Val 100 105 110 Cys Val Leu Ala His His Phe Gly Lys Glu
Phe Thr Pro Pro Val Gln 115 120 125 Ala Ala Tyr Gln Lys Val Val Ala
Xaa Xaa Ala Asn Ala Leu Ala His 130 135 140 Lys Tyr Xaa 145
<210> SEQ ID NO 21 <211> LENGTH: 124 <212> TYPE:
PRT <213> ORGANISM: Homo sapiens <400> SEQUENCE: 21 Gly
Lys Val Asn Val Asp Glu Val Gly Gly Glu Ala Leu Gly Arg Leu 1 5 10
15 Leu Val Val Tyr Pro Trp Thr Gln Arg Phe Leu Glu Ser Phe Gly Asp
20 25 30 Leu Ser Thr Pro Asp Ala Val Met Gly Asn Pro Lys Val Lys
Ala His 35 40 45 Gly Lys Lys Val Leu Gly Ala Phe Ser Asp Gly Leu
Ala His Leu Asp 50 55 60 Asn Leu Lys Gly Thr Phe Ala Thr Leu Ser
Glu Leu His Cys Asp Lys 65 70 75 80 Leu His Val Asp Pro Glu Asn Phe
Arg Leu Leu Gly Asn Val Leu Val 85 90 95 Cys Val Leu Ala His His
Phe Gly Lys Glu Phe Thr Pro Pro Val Gln 100 105 110 Ala Ala Tyr Gln
Lys Val Val Ala Gly Val Ala Asn 115 120 <210> SEQ ID NO 22
<211> LENGTH: 147 <212> TYPE: PRT <213> ORGANISM:
Homo sapiens <400> SEQUENCE: 22 Met Val His Leu Thr Pro Glu
Glu Lys Ser Ala Val Thr Ala Leu Trp 1 5 10 15 Gly Lys Val Asn Val
Asp Glu Val Gly Gly Glu Ala Leu Gly Arg Leu 20 25 30 Leu Val Val
Tyr Pro Trp Thr Gln Arg Phe Leu Glu Ser Phe Gly Asp 35 40 45 Leu
Ser Thr Pro Asp Ala Val Met Gly Asn Pro Lys Val Lys Ala His 50 55
60 Gly Lys Lys Val Leu Gly Ala Phe Ser Asp Gly Leu Ala His Leu Asp
65 70 75 80 Asn Leu Lys Gly Thr Phe Ala Thr Leu Ser Glu Leu His Cys
Asp Lys 85 90 95 Leu His Val Asp Pro Glu Asn Phe Arg Leu Leu Gly
Asn Val Leu Val 100 105 110 Cys Val Leu Ala His His Phe Gly Lys Glu
Phe Thr Pro Pro Val Gln 115 120 125 Ala Ala Tyr Gln Lys Val Val Ala
Gly Val Ala Asn Ala Leu Ala His 130 135 140 Lys Tyr His 145
<210> SEQ ID NO 23 <211> LENGTH: 80 <212> TYPE:
PRT <213> ORGANISM: Artificial Sequence <220> FEATURE:
<223> OTHER INFORMATION: N-terminal region of a consensus
sequence of mammalian transferrins <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (2)..(2) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (9)..(9) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (10)..(10) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (24)..(24) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (25)..(25) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (31)..(31) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (32)..(32) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (42)..(42) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (47)..(47) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (59)..(59) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (64)..(64) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Tyr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (65)..(65)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Leu <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (71)..(71) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ile <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (73)..(73) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (77)..(77) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (78)..(78) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <400> SEQUENCE: 23 Met Xaa Leu Ala Val Gly Ala
Leu Xaa Xaa Cys Ala Val Leu Xaa Leu 1 5 10 15 Cys Leu Ala Val Pro
Asp Lys Xaa Xaa Arg Trp Cys Ala Val Xaa Xaa 20 25 30 His Glu Ala
Thr Lys Cys Gln Ser Phe Xaa Asp His Met Lys Xaa Val 35 40 45 Ile
Pro Ser Asp Gly Pro Ser Val Ala Cys Xaa Lys Lys Ala Ser Xaa 50 55
60 Xaa Asp Cys Ile Arg Ala Xaa Ala Xaa Asn Glu Ala Xaa Xaa Val Thr
65 70 75 80 <210> SEQ ID NO 24 <211> LENGTH: 114
<212> TYPE: PRT <213> ORGANISM: Artificial Sequence
<220> FEATURE: <223> OTHER INFORMATION: C-terminal
region of a consensus sequence of mammalian transferrins
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (3)..(3) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Thr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (16)..(16)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Asp <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (18)..(18) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (19)..(19) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (23)..(23) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (25)..(25) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (28)..(28) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (30)..(30) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (61)..(61) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (62)..(62) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (75)..(75) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (78)..(78) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (96)..(96) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (97)..(97) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (105)..(105) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (107)..(107) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (110)..(110) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <400> SEQUENCE: 24 Glu Thr Xaa Glu Asp Cys Ile
Ala Lys Ile Met Asn Gly Glu Ala Xaa 1 5 10 15 Ala Xaa Xaa Leu Asp
Gly Xaa Phe Xaa Tyr Ile Xaa Gly Xaa Cys Gly 20 25 30 Leu Val Pro
Val Leu Ala Glu Asn Tyr Asn Lys Ser Asp Asn Cys Glu 35 40 45 Asp
Thr Pro Glu Ala Gly Tyr Phe Ala Val Ala Val Xaa Xaa Lys Ser 50 55
60 Ala Ser Asp Leu Thr Trp Asp Asn Leu Lys Xaa Lys Lys Xaa Cys His
65 70 75 80 Thr Ala Val Gly Arg Thr Ala Gly Trp Asn Ile Pro Met Gly
Leu Xaa 85 90 95 Xaa Asn Lys Ile Asn His Cys Arg Xaa Asp Xaa Phe
Phe Xaa Glu Gly 100 105 110 Cys Ala <210> SEQ ID NO 25
<211> LENGTH: 698 <212> TYPE: PRT <213> ORGANISM:
Artificial Sequence <220> FEATURE: <223> OTHER
INFORMATION: Consensus sequence of mammalian transferrins
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (2)..(2) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Arg <220> FEATURE:
<221> NAME/KEY: misc_feature <222> LOCATION: (9)..(9)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid <220> FEATURE: <221> NAME/KEY: MISC_FEATURE
<222> LOCATION: (10)..(10) <223> OTHER INFORMATION: Xaa
can be any naturally occurring amino acid, preferably Val
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (15)..(15) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Gly <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (24)..(24)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Thr <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (25)..(25) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (31)..(31) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (32)..(32) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (42)..(42) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (47)..(47) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (59)..(59) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (64)..(64) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (71)..(71) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (73)..(73) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Ile <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Asp <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (78)..(78) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (84)..(84) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (87)..(87) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (88)..(88) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (94)..(94) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (98)..(98) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (104)..(104) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (107)..(107) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (108)..(108) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (115)..(115) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (116)..(116) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (121)..(121) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (124)..(124) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (129)..(129) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (146)..(146) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (152)..(152) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (153)..(153) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (161)..(161) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (179)..(179) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (188)..(188) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (204)..(204) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (224)..(224) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (238)..(238) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (245)..(245) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (255)..(255) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (268)..(268) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably His <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (275)..(275) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (276)..(276) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (286)..(286) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (293)..(293) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (309)..(309) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (310)..(310) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (326)..(326) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (327)..(327) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (357)..(357) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (381)..(381) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (393)..(393) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (406)..(406) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (408)..(408) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (409)..(409) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (413)..(413) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (415)..(415) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (418)..(418) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (420)..(420) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (451)..(451) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (452)..(452) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (465)..(465) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (468)..(468) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (486)..(486) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (487)..(487) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (495)..(495) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (497)..(497) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (500)..(500) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (505)..(505) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (509)..(509) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (529)..(529) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (536)..(536) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (558)..(558) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (559)..(559) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (563)..(563) <223> OTHER INFORMATION: Xaa can be
any naturally occurring amino acid, preferably Gly <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(586)..(586) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Thr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(592)..(592) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Glu <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(603)..(603) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Asn <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(607)..(607) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Val <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(622)..(622) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Gln <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(635)..(635) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Ser <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(648)..(648) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Leu <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(652)..(652) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Asp <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(678)..(678) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Val <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(692)..(692) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Ala <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(696)..(696) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Arg <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(698)..(698) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Pro <400>
SEQUENCE: 25 Met Xaa Leu Ala Val Gly Ala Leu Xaa Xaa Cys Ala Val
Leu Xaa Leu 1 5 10 15 Cys Leu Ala Val Pro Asp Lys Xaa Xaa Arg Trp
Cys Ala Val Xaa Xaa 20 25 30 His Glu Ala Thr Lys Cys Gln Ser Phe
Xaa Asp His Met Lys Xaa Val 35 40 45 Ile Pro Ser Asp Gly Pro Ser
Val Ala Cys Xaa Lys Lys Ala Ser Xaa 50 55 60 Xaa Asp Cys Ile Arg
Ala Xaa Ala Xaa Asn Glu Ala Xaa Xaa Val Thr 65 70 75 80 Leu Asp Ala
Xaa Leu Val Xaa Xaa Ala Tyr Leu Ala Pro Xaa Asn Leu 85 90 95 Lys
Xaa Val Val Ala Glu Phe Xaa Gly Ser Xaa Xaa Asp Pro Gln Thr 100 105
110 Phe Tyr Xaa Xaa Val Ala Val Val Xaa Lys Asp Xaa Gly Phe Gln Met
115 120 125 Xaa Gln Leu Arg Gly Lys Lys Ser Cys His Thr Gly Leu Gly
Arg Ser 130 135 140 Ala Xaa Trp Asn Ile Pro Ile Xaa Xaa Leu Tyr Cys
Asp Leu Pro Glu 145 150 155 160 Xaa Arg Lys Pro Leu Glu Lys Ala Val
Ala Asn Phe Phe Ser Gly Ser 165 170 175 Cys Ala Xaa Cys Ala Asp Gly
Thr Asp Phe Pro Xaa Leu Cys Gln Leu 180 185 190 Cys Pro Gly Cys Gly
Cys Ser Thr Leu Asn Gln Xaa Phe Gly Tyr Ser 195 200 205 Gly Ala Phe
Lys Cys Leu Lys Asp Gly Ala Gly Asp Val Ala Phe Xaa 210 215 220 Lys
His Ser Thr Ile Phe Glu Asn Leu Ala Asn Lys Ala Xaa Arg Asp 225 230
235 240 Gln Tyr Glu Leu Xaa Cys Leu Asp Asn Thr Arg Lys Pro Val Xaa
Glu 245 250 255 Tyr Lys Asp Cys His Leu Ala Gln Val Pro Ser Xaa Thr
Val Val Ala 260 265 270 Arg Ser Xaa Xaa Gly Lys Glu Asp Leu Ile Trp
Glu Leu Xaa Asn Gln 275 280 285 Ala Gln Glu His Xaa Gly Lys Asp Lys
Ser Lys Glu Phe Gln Leu Phe 290 295 300 Ser Ser Pro His Xaa Xaa Asp
Leu Leu Phe Lys Asp Ser Ala His Gly 305 310 315 320 Phe Leu Lys Val
Pro Xaa Xaa Met Asp Ala Lys Met Tyr Leu Gly Tyr 325 330 335 Glu Tyr
Val Thr Ala Ile Arg Asn Leu Arg Glu Gly Thr Cys Pro Glu 340 345 350
Ala Pro Thr Asp Xaa Cys Lys Pro Val Lys Trp Cys Ala Leu Ser His 355
360 365 His Glu Arg Leu Lys Cys Asp Glu Trp Ser Val Asn Xaa Val Gly
Lys 370 375 380 Ile Glu Cys Val Ser Ala Glu Thr Xaa Glu Asp Cys Ile
Ala Lys Ile 385 390 395 400 Met Asn Gly Glu Ala Xaa Ala Xaa Xaa Leu
Asp Gly Xaa Phe Xaa Tyr 405 410 415 Ile Xaa Gly Xaa Cys Gly Leu Val
Pro Val Leu Ala Glu Asn Tyr Asn 420 425 430 Lys Ser Asp Asn Cys Glu
Asp Thr Pro Glu Ala Gly Tyr Phe Ala Val 435 440 445 Ala Val Xaa Xaa
Lys Ser Ala Ser Asp Leu Thr Trp Asp Asn Leu Lys 450 455 460 Xaa Lys
Lys Xaa Cys His Thr Ala Val Gly Arg Thr Ala Gly Trp Asn 465 470 475
480 Ile Pro Met Gly Leu Xaa Xaa Asn Lys Ile Asn His Cys Arg Xaa Asp
485 490 495 Xaa Phe Phe Xaa Glu Gly Cys Ala Xaa Gly Ser Lys Xaa Asp
Ser Ser 500 505 510 Leu Cys Lys Leu Cys Met Gly Ser Gly Leu Asn Leu
Cys Glu Pro Asn 515 520 525 Xaa Lys Glu Gly Tyr Tyr Gly Xaa Thr Gly
Ala Phe Arg Cys Leu Val 530 535 540 Glu Lys Gly Asp Val Ala Phe Val
Lys His Gln Thr Val Xaa Xaa Asn 545 550 555 560 Thr Gly Xaa Lys Asn
Pro Asp Pro Trp Ala Lys Asn Leu Asn Glu Lys 565 570 575 Asp Tyr Glu
Leu Leu Cys Leu Asp Gly Xaa Arg Lys Pro Val Glu Xaa 580 585 590 Tyr
Ala Asn Cys His Leu Ala Arg Ala Pro Xaa His Ala Val Xaa Thr 595 600
605 Arg Lys Asp Lys Glu Ala Cys Val His Lys Ile Leu Arg Xaa Gln Gln
610 615 620 His Leu Phe Gly Ser Asn Val Thr Asp Cys Xaa Gly Asn Phe
Cys Leu 625 630 635 640 Phe Arg Ser Glu Thr Lys Asp Xaa Leu Phe Arg
Xaa Asp Thr Val Cys 645 650 655 Leu Ala Lys Leu His Asp Arg Asn Thr
Tyr Glu Lys Tyr Leu Gly Glu 660 665 670 Glu Tyr Val Lys Ala Xaa Gly
Asn Leu Arg Lys Cys Ser Thr Ser Ser 675 680 685 Leu Leu Glu Xaa Cys
Thr Phe Xaa Arg Xaa 690 695 <210> SEQ ID NO 26 <211>
LENGTH: 80 <212> TYPE: PRT <213> ORGANISM: Homo sapiens
<400> SEQUENCE: 26 Met Arg Leu Ala Val Gly Ala Leu Leu Val
Cys Ala Val Leu Gly Leu 1 5 10 15 Cys Leu Ala Val Pro Asp Lys Thr
Val Arg Trp Cys Ala Val Ser Glu 20 25 30 His Glu Ala Thr Lys Cys
Gln Ser Phe Arg Asp His Met Lys Ser Val 35 40 45 Ile Pro Ser Asp
Gly Pro Ser Val Ala Cys Val Lys Lys Ala Ser Tyr 50 55 60 Leu Asp
Cys Ile Arg Ala Ile Ala Ala Asn Glu Ala Asp Ala Val Thr 65 70 75 80
<210> SEQ ID NO 27 <211> LENGTH: 114 <212> TYPE:
PRT <213> ORGANISM: Homo sapiens <400> SEQUENCE: 27 Glu
Thr Thr Glu Asp Cys Ile Ala Lys Ile Met Asn Gly Glu Ala Asp 1 5 10
15 Ala Met Ser Leu Asp Gly Gly Phe Val Tyr Ile Ala Gly Lys Cys Gly
20 25 30 Leu Val Pro Val Leu Ala Glu Asn Tyr Asn Lys Ser Asp Asn
Cys Glu 35 40 45 Asp Thr Pro Glu Ala Gly Tyr Phe Ala Val Ala Val
Val Lys Lys Ser 50 55 60 Ala Ser Asp Leu Thr Trp Asp Asn Leu Lys
Gly Lys Lys Ser Cys His 65 70 75 80 Thr Ala Val Gly Arg Thr Ala Gly
Trp Asn Ile Pro Met Gly Leu Leu 85 90 95 Tyr Asn Lys Ile Asn His
Cys Arg Phe Asp Glu Phe Phe Ser Glu Gly 100 105 110 Cys Ala
<210> SEQ ID NO 28 <211> LENGTH: 698 <212> TYPE:
PRT <213> ORGANISM: Homo sapiens <400> SEQUENCE: 28 Met
Arg Leu Ala Val Gly Ala Leu Leu Val Cys Ala Val Leu Gly Leu 1 5 10
15 Cys Leu Ala Val Pro Asp Lys Thr Val Arg Trp Cys Ala Val Ser Glu
20 25 30 His Glu Ala Thr Lys Cys Gln Ser Phe Arg Asp His Met Lys
Ser Val 35 40 45 Ile Pro Ser Asp Gly Pro Ser Val Ala Cys Val Lys
Lys Ala Ser Tyr 50 55 60 Leu Asp Cys Ile Arg Ala Ile Ala Ala Asn
Glu Ala Asp Ala Val Thr 65 70 75 80 Leu Asp Ala Gly Leu Val Tyr Asp
Ala Tyr Leu Ala Pro Asn Asn Leu 85 90 95 Lys Pro Val Val Ala Glu
Phe Tyr Gly Ser Lys Glu Asp Pro Gln Thr 100 105 110 Phe Tyr Tyr Ala
Val Ala Val Val Lys Lys Asp Ser Gly Phe Gln Met 115 120 125 Asn Gln
Leu Arg Gly Lys Lys Ser Cys His Thr Gly Leu Gly Arg Ser 130 135 140
Ala Gly Trp Asn Ile Pro Ile Gly Leu Leu Tyr Cys Asp Leu Pro Glu 145
150 155 160 Pro Arg Lys Pro Leu Glu Lys Ala Val Ala Asn Phe Phe Ser
Gly Ser 165 170 175 Cys Ala Pro Cys Ala Asp Gly Thr Asp Phe Pro Gln
Leu Cys Gln Leu 180 185 190 Cys Pro Gly Cys Gly Cys Ser Thr Leu Asn
Gln Tyr Phe Gly Tyr Ser 195 200 205 Gly Ala Phe Lys Cys Leu Lys Asp
Gly Ala Gly Asp Val Ala Phe Val 210 215 220 Lys His Ser Thr Ile Phe
Glu Asn Leu Ala Asn Lys Ala Asp Arg Asp 225 230 235 240 Gln Tyr Glu
Leu Leu Cys Leu Asp Asn Thr Arg Lys Pro Val Asp Glu 245 250 255 Tyr
Lys Asp Cys His Leu Ala Gln Val Pro Ser His Thr Val Val Ala 260 265
270 Arg Ser Met Gly Gly Lys Glu Asp Leu Ile Trp Glu Leu Leu Asn Gln
275 280 285 Ala Gln Glu His Phe Gly Lys Asp Lys Ser Lys Glu Phe Gln
Leu Phe 290 295 300 Ser Ser Pro His Gly Lys Asp Leu Leu Phe Lys Asp
Ser Ala His Gly 305 310 315 320 Phe Leu Lys Val Pro Pro Arg Met Asp
Ala Lys Met Tyr Leu Gly Tyr 325 330 335 Glu Tyr Val Thr Ala Ile Arg
Asn Leu Arg Glu Gly Thr Cys Pro Glu 340 345 350 Ala Pro Thr Asp Glu
Cys Lys Pro Val Lys Trp Cys Ala Leu Ser His 355 360 365 His Glu Arg
Leu Lys Cys Asp Glu Trp Ser Val Asn Ser Val Gly Lys 370 375 380 Ile
Glu Cys Val Ser Ala Glu Thr Thr Glu Asp Cys Ile Ala Lys Ile 385 390
395 400 Met Asn Gly Glu Ala Asp Ala Met Ser Leu Asp Gly Gly Phe Val
Tyr 405 410 415 Ile Ala Gly Lys Cys Gly Leu Val Pro Val Leu Ala Glu
Asn Tyr Asn 420 425 430 Lys Ser Asp Asn Cys Glu Asp Thr Pro Glu Ala
Gly Tyr Phe Ala Val 435 440 445 Ala Val Val Lys Lys Ser Ala Ser Asp
Leu Thr Trp Asp Asn Leu Lys 450 455 460 Gly Lys Lys Ser Cys His Thr
Ala Val Gly Arg Thr Ala Gly Trp Asn 465 470 475 480 Ile Pro Met Gly
Leu Leu Tyr Asn Lys Ile Asn His Cys Arg Phe Asp 485 490 495 Glu Phe
Phe Ser Glu Gly Cys Ala Pro Gly Ser Lys Lys Asp Ser Ser 500 505 510
Leu Cys Lys Leu Cys Met Gly Ser Gly Leu Asn Leu Cys Glu Pro Asn 515
520 525 Asn Lys Glu Gly Tyr Tyr Gly Tyr Thr Gly Ala Phe Arg Cys Leu
Val 530 535 540 Glu Lys Gly Asp Val Ala Phe Val Lys His Gln Thr Val
Pro Gln Asn 545 550 555 560 Thr Gly Gly Lys Asn Pro Asp Pro Trp Ala
Lys Asn Leu Asn Glu Lys 565 570 575 Asp Tyr Glu Leu Leu Cys Leu Asp
Gly Thr Arg Lys Pro Val Glu Glu 580 585 590 Tyr Ala Asn Cys His Leu
Ala Arg Ala Pro Asn His Ala Val Val Thr 595 600 605 Arg Lys Asp Lys
Glu Ala Cys Val His Lys Ile Leu Arg Gln Gln Gln 610 615 620 His Leu
Phe Gly Ser Asn Val Thr Asp Cys Ser Gly Asn Phe Cys Leu 625 630 635
640 Phe Arg Ser Glu Thr Lys Asp Leu Leu Phe Arg Asp Asp Thr Val Cys
645 650 655 Leu Ala Lys Leu His Asp Arg Asn Thr Tyr Glu Lys Tyr Leu
Gly Glu 660 665 670 Glu Tyr Val Lys Ala Val Gly Asn Leu Arg Lys Cys
Ser Thr Ser Ser 675 680 685 Leu Leu Glu Ala Cys Thr Phe Arg Arg Pro
690 695
1 SEQUENCE LISTING <160> NUMBER OF SEQ ID NOS: 28 <210>
SEQ ID NO 1 <211> LENGTH: 132 <212> TYPE: PRT
<213> ORGANISM: Artificial Sequence <220> FEATURE:
<223> OTHER INFORMATION: Minimal consensus sequence of
mammalian ferritin H chain <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (5)..(5) <223>
OTHER INFORMATION: Xaa at position 5 may be present or absent, if
present it means any naturally occurring amino acid, preferably Ile
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (6)..(6) <223> OTHER INFORMATION: Xaa at position 6
means any naturally occurring amino acid, preferably Asn
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (14)..(14) <223> OTHER INFORMATION: Xaa at position
14 means any naturally occurring amino acid, preferably Tyr
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (24)..(24) <223> OTHER INFORMATION: Xaa at position
24 means any naturally occurring amino acid, preferably Tyr or Cys,
more preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (66)..(66) <223> OTHER
INFORMATION: Xaa at position 66 means any naturally occurring amino
acid, preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa at position 68 means any naturally occurring amino
acid, preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (75)..(75) <223> OTHER
INFORMATION: Xaa at position 75 means any naturally occurring amino
acid, preferably Arg or Cys, more preferably Cys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(90)..(90) <223> OTHER INFORMATION: Xaa at position 90 means
any naturally occurring amino acid, preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(94)..(94) <223> OTHER INFORMATION: Xaa at position 94 means
any naturally occurring amino acid, preferably Ser or Asn, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (120)..(120) <223> OTHER
INFORMATION: Xaa at position 120 may be present or absent, if
present it means any naturally occurring amino acid, preferably His
or Tyr, more preferably His <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Xaa at position 123 means any
naturally occurring amino acid, preferably Asn or Ser, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (128)..(128) <223> OTHER
INFORMATION: Xaa at position 128 means any naturally occurring
amino acid, preferably Ala or Ser, more preferably Ala <400>
SEQUENCE: 1 Ser Glu Ala Ala Xaa Xaa Arg Gln Ile Asn Leu Glu Leu Xaa
Ala Ser 1 5 10 15 Tyr Val Tyr Leu Ser Met Ser Xaa Tyr Phe Asp Arg
Asp Asp Val Ala 20 25 30 Leu Lys Asn Phe Ala Lys Tyr Phe Leu His
Gln Ser His Glu Glu Arg 35 40 45 Glu His Ala Glu Lys Leu Met Lys
Leu Gln Asn Gln Arg Gly Gly Arg 50 55 60 Ile Xaa Leu Xaa Asp Ile
Lys Lys Pro Asp Xaa Asp Asp Trp Glu Ser 65 70 75 80 Gly Leu Asn Ala
Met Glu Cys Ala Leu Xaa Leu Glu Lys Xaa Asn Gln 85 90 95 Ser Leu
Leu Glu Leu His Lys Leu Ala Thr Asp Lys Asn Asp Pro His 100 105 110
Leu Cys Asp Phe Ile Glu Thr Xaa Tyr Leu Xaa Glu Gln Val Lys Xaa 115
120 125 Ile Lys Glu Leu 130 <210> SEQ ID NO 2 <211>
LENGTH: 183 <212> TYPE: PRT <213> ORGANISM: Artificial
Sequence <220> FEATURE: <223> OTHER INFORMATION:
Consensus sequence of mammalian ferritin H chain <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(6)..(6) <223> OTHER INFORMATION: Xaa at position 6 can be
any naturally occurring amino acid, preferably Pro <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(14)..(14) <223> OTHER INFORMATION: Xaa at position 14 can be
any naturally occurring amino acid, preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(16)..(16) <223> OTHER INFORMATION: Xaa at position 16 can be
any naturally occurring amino acid, preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(21)..(21) <223> OTHER INFORMATION: Xaa at position 21 may be
present or absent, if present it means any naturally occurring
amino acid, preferably Ile <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (22)..(22) <223>
OTHER INFORMATION: Xaa at position 22 means any naturally occurring
amino acid, preferably Asn <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (30)..(30) <223>
OTHER INFORMATION: Xaa at position 30 can be any naturally
occurring amino acid, preferably Tyr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Xaa at position 40 can be any
naturally occurring amino acid, preferably Tyr or Cys, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (82)..(82) <223> OTHER
INFORMATION: Xaa at position 82 can be any naturally occurring
amino acid, preferably Phe <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (84)..(84) <223>
OTHER INFORMATION: Xaa at position 84 can be any naturally
occurring amino acid, preferably Gln <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (91)..(91)
<223> OTHER INFORMATION: Xaa at position 91 can be any
naturally occurring amino acid, preferably Arg or Cys, more
preferably Cys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (106)..(106) <223> OTHER
INFORMATION: Xaa at position 106 can be any naturally occurring
amino acid, preferably His <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (110)..(110)
<223> OTHER INFORMATION: Xaa at position 110 can be any
naturally occurring amino acid, preferably Asn or Ser, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (137)..(137) <223> OTHER
INFORMATION: Xaa at position 137 can be any naturally occurring
amino acid, preferably His or Tyr, more preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(140)..(140) <223> OTHER INFORMATION: Xaa at position 140 can
be any naturally occurring amino acid, preferably Asn or Ser, more
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (145)..(145) <223> OTHER
INFORMATION: Xaa at position 145 can be any naturally occurring
amino acid, preferably Ala or Ser, more preferably Ala <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(164)..(164) <223> OTHER INFORMATION: Xaa at position 164 can
be any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (166)..(166) <223> OTHER
INFORMATION: Xaa at position 166 can be any naturally occurring
amino acid, preferably Met or Leu, prerferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(178)..(178) <223> OTHER INFORMATION: Xaa at position 178 can
be any naturally occurring amino acid, preferably Asp or His, more
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (181)..(181) <223> OTHER
INFORMATION: Xaa at position 181 is absent or any naturally
occurring amino acid, preferably Asn <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(182)..(182) <223> OTHER INFORMATION: Xaa at position 182 is
absent or any naturally occurring amino acid, preferably Glu
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (183)..(183) <223> OTHER INFORMATION: Xaa at
position 183 is absent or any naturally occurring amino acid,
preferably Ser <400> SEQUENCE: 2 Met Thr Thr Ala Ser Xaa Ser
Gln Val Arg Gln Asn Tyr Xaa Gln Xaa 1 5 10 15 Ser Glu Ala Ala Xaa
Xaa Arg Gln Ile Asn Leu Glu Leu Xaa Ala Ser 20 25 30 Tyr Val Tyr
Leu Ser Met Ser Xaa Tyr Phe Asp Arg Asp Asp Val Ala 35 40 45 Leu
Lys Asn Phe Ala Lys Tyr Phe Leu His Gln Ser His Glu Glu Arg 50 55
60 Glu His Ala Glu Lys Leu Met Lys Leu Gln Asn Gln Arg Gly Gly Arg
65 70 75 80 Ile Xaa Leu Xaa Asp Ile Lys Lys Pro Asp Xaa Asp Asp Trp
Glu Ser 85 90 95 Gly Leu Asn Ala Met Glu Cys Ala Leu Xaa Leu Glu
Lys Xaa Val Asn 100 105 110
Gln Ser Leu Leu Glu Leu His Lys Leu Ala Thr Asp Lys Asn Asp Pro 115
120 125 His Leu Cys Asp Phe Ile Glu Thr Xaa Tyr Leu Xaa Glu Gln Val
Lys 130 135 140 Xaa Ile Lys Glu Leu Gly Asp His Val Thr Asn Leu Arg
Lys Met Gly 145 150 155 160 Ala Pro Glu Xaa Gly Xaa Ala Glu Tyr Leu
Phe Asp Lys His Thr Leu 165 170 175 Gly Xaa Ser Asp Xaa Xaa Xaa 180
<210> SEQ ID NO 3 <211> LENGTH: 133 <212> TYPE:
PRT <213> ORGANISM: Mus musculus <400> SEQUENCE: 3 Ala
Glu Ala Ala Ile Asn Arg Gln Ile Asn Leu Glu Leu Tyr Ala Ser 1 5 10
15 Tyr Val Tyr Leu Ser Met Ser Cys Tyr Phe Asp Arg Asp Asp Val Ala
20 25 30 Leu Lys Asn Phe Ala Lys Tyr Phe Leu His Gln Ser His Glu
Glu Arg 35 40 45 Glu His Ala Glu Lys Leu Met Lys Leu Gln Asn Gln
Arg Gly Gly Arg 50 55 60 Ile Phe Leu Gln Asp Ile Lys Lys Pro Asp
Arg Asp Asp Trp Glu Ser 65 70 75 80 Gly Leu Asn Ala Met Glu Cys Ala
Leu His Leu Glu Lys Ser Val Asn 85 90 95 Gln Ser Leu Leu Glu Leu
His Lys Leu Ala Thr Asp Lys Asn Asp Pro 100 105 110 His Leu Cys Asp
Phe Ile Glu Thr Tyr Tyr Leu Ser Glu Gln Val Lys 115 120 125 Ser Ile
Lys Glu Leu 130 <210> SEQ ID NO 4 <211> LENGTH: 180
<212> TYPE: PRT <213> ORGANISM: Mus musculus
<400> SEQUENCE: 4 Met Thr Thr Ala Ser Pro Ser Gln Val Arg Gln
Asn Tyr His Gln Asp 1 5 10 15 Ala Glu Ala Ala Ile Asn Arg Gln Ile
Asn Leu Glu Leu Tyr Ala Ser 20 25 30 Tyr Val Tyr Leu Ser Met Ser
Cys Tyr Phe Asp Arg Asp Asp Val Ala 35 40 45 Leu Lys Asn Phe Ala
Lys Tyr Phe Leu His Gln Ser His Glu Glu Arg 50 55 60 Glu His Ala
Glu Lys Leu Met Lys Leu Gln Asn Gln Arg Gly Gly Arg 65 70 75 80 Ile
Phe Leu Gln Asp Ile Lys Lys Pro Asp Arg Asp Asp Trp Glu Ser 85 90
95 Gly Leu Asn Ala Met Glu Cys Ala Leu His Leu Glu Lys Ser Val Asn
100 105 110 Gln Ser Leu Leu Glu Leu His Lys Leu Ala Thr Asp Lys Asn
Asp Pro 115 120 125 His Leu Cys Asp Phe Ile Glu Thr Tyr Tyr Leu Ser
Glu Gln Val Lys 130 135 140 Ser Ile Lys Glu Leu Gly Asp His Val Thr
Asn Leu Arg Lys Met Gly 145 150 155 160 Ala Pro Glu Ala Gly Met Ala
Glu Tyr Leu Phe Asp Lys His Thr Leu 165 170 175 Gly His Gly Asp 180
<210> SEQ ID NO 5 <211> LENGTH: 132 <212> TYPE:
PRT <213> ORGANISM: Homo sapiens <400> SEQUENCE: 5 Ser
Glu Ala Ala Asn Arg Gln Ile Asn Leu Glu Leu Tyr Ala Ser Tyr 1 5 10
15 Val Tyr Leu Ser Met Ser Tyr Tyr Phe Asp Arg Asp Asp Val Ala Leu
20 25 30 Lys Asn Phe Ala Lys Tyr Phe Leu His Gln Ser His Glu Glu
Arg Glu 35 40 45 His Ala Glu Lys Leu Met Lys Leu Gln Asn Gln Arg
Gly Gly Arg Ile 50 55 60 Phe Leu Gln Asp Ile Lys Lys Pro Asp Cys
Asp Asp Trp Glu Ser Gly 65 70 75 80 Leu Asn Ala Met Glu Cys Ala Leu
His Leu Glu Lys Asn Val Asn Gln 85 90 95 Ser Leu Leu Glu Leu His
Lys Leu Ala Thr Asp Lys Asn Asp Pro His 100 105 110 Leu Cys Asp Phe
Ile Glu Thr His Tyr Leu Asn Glu Gln Val Lys Ala 115 120 125 Ile Lys
Glu Leu 130 <210> SEQ ID NO 6 <211> LENGTH: 179
<212> TYPE: PRT <213> ORGANISM: Homo sapiens
<400> SEQUENCE: 6 Met Thr Thr Ala Ser Thr Ser Gln Val Arg Gln
Asn Tyr His Gln Asp 1 5 10 15 Ser Glu Ala Ala Asn Arg Gln Ile Asn
Leu Glu Leu Tyr Ala Ser Tyr 20 25 30 Val Tyr Leu Ser Met Ser Tyr
Tyr Phe Asp Arg Asp Asp Val Ala Leu 35 40 45 Lys Asn Phe Ala Lys
Tyr Phe Leu His Gln Ser His Glu Glu Arg Glu 50 55 60 His Ala Glu
Lys Leu Met Lys Leu Gln Asn Gln Arg Gly Gly Arg Ile 65 70 75 80 Phe
Leu Gln Asp Ile Lys Lys Pro Asp Cys Asp Asp Trp Glu Ser Gly 85 90
95 Leu Asn Ala Met Glu Cys Ala Leu His Leu Glu Lys Asn Val Asn Gln
100 105 110 Ser Leu Leu Glu Leu His Lys Leu Ala Thr Asp Lys Asn Asp
Pro His 115 120 125 Leu Cys Asp Phe Ile Glu Thr His Tyr Leu Asn Glu
Gln Val Lys Ala 130 135 140 Ile Lys Glu Leu Gly Asp His Val Thr Asn
Leu Arg Lys Met Gly Ala 145 150 155 160 Pro Glu Ser Gly Leu Ala Glu
Tyr Leu Phe Asp Lys His Thr Leu Gly 165 170 175 Asp Ser Asp
<210> SEQ ID NO 7 <211> LENGTH: 182 <212> TYPE:
PRT <213> ORGANISM: Artificial Sequence <220> FEATURE:
<223> OTHER INFORMATION: Consensus sequence of mammalian
ferritin H chains 2nd version <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (6)..(6) <223>
OTHER INFORMATION: Xaa at position 6 can be any naturally occurring
amino acid, preferably Pro <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (14)..(14) <223>
OTHER INFORMATION: Xaa at position 14 can be any naturally
occurring amino acid, preferably His <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (16)..(16)
<223> OTHER INFORMATION: Xaa at position 16 can be any
naturally occurring amino acid, preferably Asp <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (21)..(21)
<223> OTHER INFORMATION: Xaa at position 21 may be present or
absent, if present it means any naturally occurring amino acid,
preferably Ile <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (29)..(29) <223> OTHER
INFORMATION: Xaa at position 29 can be any naturally occurring
amino acid, preferably Tyr <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (81)..(81) <223>
OTHER INFORMATION: Xaa at position 81 can be any naturally
occurring amino acid, preferably Phe <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (83)..(83)
<223> OTHER INFORMATION: Xaa at position 83 can be any
naturally occurring amino acid, preferably Gln <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(105)..(105) <223> OTHER INFORMATION: Xaa at position 105 can
be any naturally occurring amino acid, preferably His <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(144)..(144) <223> OTHER INFORMATION: Xaa at position 144 can
be any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (180)..(180) <223> OTHER
INFORMATION: Xaa at position 180 is absent or any naturally
occuring amino acid, preferably Asn <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(181)..(181) <223> OTHER INFORMATION: Xaa at position 181 is
absent or any naturally occuring amino acid, preferably Glu
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (182)..(182) <223> OTHER INFORMATION: Xaa at
position 182 is absent or any naturally occuring amino acid,
preferably Ser <400> SEQUENCE: 7 Met Thr Thr Ala Ser Xaa Ser
Gln Val Arg Gln Asn Tyr Xaa Gln Xaa 1 5 10 15
Ser Glu Ala Ala Xaa Arg Gln Ile Asn Leu Glu Leu Xaa Ala Ser Tyr 20
25 30 Val Tyr Leu Ser Met Ser Tyr Tyr Phe Asp Arg Asp Asp Val Ala
Leu 35 40 45 Lys Asn Phe Ala Lys Tyr Phe Leu His Gln Ser His Glu
Glu Arg Glu 50 55 60 His Ala Glu Lys Leu Met Lys Leu Gln Asn Gln
Arg Gly Gly Arg Ile 65 70 75 80 Xaa Leu Xaa Asp Ile Lys Lys Pro Asp
Cys Asp Asp Trp Glu Ser Gly 85 90 95 Leu Asn Ala Met Glu Cys Ala
Leu Xaa Leu Glu Lys Asn Val Asn Gln 100 105 110 Ser Leu Leu Glu Leu
His Lys Leu Ala Thr Asp Lys Asn Asp Pro His 115 120 125 Leu Cys Asp
Phe Ile Glu Thr His Tyr Leu Asn Glu Gln Val Lys Xaa 130 135 140 Ile
Lys Glu Leu Gly Asp His Val Thr Asn Leu Arg Lys Met Gly Ala 145 150
155 160 Pro Glu Ser Gly Leu Ala Glu Tyr Leu Phe Asp Lys His Thr Leu
Gly 165 170 175 Asp Ser Asp Xaa Xaa Xaa 180 <210> SEQ ID NO 8
<211> LENGTH: 150 <212> TYPE: PRT <213> ORGANISM:
Artificial Sequence <220> FEATURE: <223> OTHER
INFORMATION: Minimal consensus sequence of mammalian ferritin L
chain <220> FEATURE: <221> NAME/KEY: MISC_FEATURE
<222> LOCATION: (5)..(5) <223> OTHER INFORMATION: Xaa
at position 5 can be any naturally occurring amino acid, preferably
Asp or Glu, more preferably Asp <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (12)..(12) <223>
OTHER INFORMATION: Xaa at position 12 can be any nnaturally
occurring amino acid, preferably Arg or Ser, more preferably Ser
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (17)..(17) <223> OTHER INFORMATION: Xaa at position
17 can be any naturally occurring amino acid, preferably Ser or
Arg, more preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (19)..(19) <223> OTHER
INFORMATION: Xaa at position 19 can be any naturally occurring
amino acid, preferably Arg or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(29)..(29) <223> OTHER INFORMATION: Xaa at position 29 can be
any naturally occurring amino acid, preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(30)..(30) <223> OTHER INFORMATION: Xaa at position 30 can be
any naturally occurring amino acid, preferably Tyr or Phe, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (42)..(42) <223> OTHER
INFORMATION: Xaa at position 42 can be any naturally occurring
amino acid, preferably Ser or Gly, more preferably Ser <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(56)..(56) <223> OTHER INFORMATION: Xaa at position 56 can be
any naturally occurirng amino acid, preferably Ala or Tyr, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (61)..(61) <223> OTHER
INFORMATION: Xaa at position 61 can be any naturally occurring
amino acid, preferably Glu or Lys, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(62)..(62) <223> OTHER INFORMATION: Xaa at position 62 can be
any naturally occurring amino acid, preferably Met or Phe, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa at position 65 can be any naturally occurring
amino acid, preferably Asp or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(75)..(75) <223> OTHER INFORMATION: Xaa at position 75 can be
any naturally occurring amino acid, preferably Ile or Val, more
preferably Ile <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (76)..(76) <223> OTHER
INFORMATION: Xaa at position 76 can be any naturally occurring
amino acid, preferably Lys or Gln, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(79)..(79) <223> OTHER INFORMATION: Xaa at position 79 can be
any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (80)..(80) <223> OTHER
INFORMATION: Xaa at position 80 can be any naturally occurring
amino acid, preferably Glu or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(87)..(87) <223> OTHER INFORMATION: Xaa at position 87 can be
any naturally occurring amino acid, preferably Pro or Gln, more
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (88)..(88) <223> OTHER
INFORMATION: Xaa at position 88 can be any naturally occurring
amino acid, preferably Glu or Asp, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(91)..(91) <223> OTHER INFORMATION: Xaa at position 91 can be
any naturally occurring amino acid, preferably Glu or Lys, more
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (94)..(94) <223> OTHER
INFORMATION: Xaa at position 94 can be any naturally occurring
amino acid, preferably Met or Leu, more preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(96)..(96) <223> OTHER INFORMATION: Xaa at position 96 can be
any naturally occurring amino acid, preferably Met or Leu, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (99)..(99) <223> OTHER
INFORMATION: Xaa at position 99 can be any naturally occurring
amino acid, preferably Lys or Asn, preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(115)..(115) <223> OTHER INFORMATION: Xaa at position 115 can
be any naturally occurring amino acid, preferably Thr or Ala, more
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (119)..(119) <223> OTHER
INFORMATION: Xaa at position 119 can be any naturally occurring
amino acid, preferably Leu <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (125)..(125)
<223> OTHER INFORMATION: Xaa at position 125 can be any
naturally occurring amino acid, preferably Ser or Thr, more
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (127)..(127) <223> OTHER
INFORMATION: Xaa at position 127 can be any naturally occurring
amino acid, preferably Tyr or Phe, more preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(130)..(130) <223> OTHER INFORMATION: Xaa at position 130 can
be any naturally occurring amino acid, preferably Lys or Glu, more
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (140)..(140) <223> OTHER
INFORMATION: Xaa at position 140 can be any naturally occurring
amino acid, preferably Asp or Asn, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(146)..(146) <223> OTHER INFORMATION: Xaa at position 146 can
be any naturally occurring amino acid, preferably Arg or His, more
preferably His <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (148)..(148) <223> OTHER
INFORMATION: Xaa at position 148 can be any naturally occurring
amino acid, preferably Leu or Val, more preferably Leu <400>
SEQUENCE: 8 Asn Tyr Ser Thr Xaa Val Glu Ala Ala Val Asn Xaa Leu Val
Asn Leu 1 5 10 15 Xaa Leu Xaa Ala Ser Tyr Thr Tyr Leu Ser Leu Gly
Xaa Xaa Phe Asp 20 25 30 Arg Asp Asp Val Ala Leu Glu Gly Val Xaa
His Phe Phe Arg Glu Leu 35 40 45 Ala Glu Glu Lys Arg Glu Gly Xaa
Glu Arg Leu Leu Xaa Xaa Gln Asn 50 55 60 Xaa Arg Gly Gly Arg Ala
Leu Phe Gln Asp Xaa Xaa Lys Pro Xaa Xaa 65 70 75 80 Asp Glu Trp Gly
Lys Thr Xaa Xaa Ala Met Xaa Ala Ala Xaa Ala Xaa 85 90 95 Glu Lys
Xaa Leu Asn Gln Ala Leu Leu Asp Leu His Ala Leu Gly Ser 100 105 110
Ala Arg Xaa Asp Pro His Xaa Cys Asp Phe Leu Glu Xaa His Xaa Leu 115
120 125 Asp Xaa Glu Val Lys Leu Ile Lys Lys Met Gly Xaa His Leu Thr
Asn 130 135 140 Leu Xaa Arg Xaa Ala Gly 145 150 <210> SEQ ID
NO 9 <211> LENGTH: 183 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Consensus sequence of mammalian ferritin L chain
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (12)..(12) <223> OTHER INFORMATION: Xaa at position
12 can be any naturally occurring amino acid, preferably Asp or
Glu, more preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (19)..(19) <223> OTHER
INFORMATION: Xaa at position 19 can be any naturally occurring
amino acid, preferably Ser or Arg, more preferably Ser <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(24)..(24) <223> OTHER INFORMATION: Xaa at position 24 can be
any naturally occurring amino acid, preferably Ser or Arg, more
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (26)..(26) <223> OTHER
INFORMATION: Xaa at position 26 can be any naturally occurring
amino acid, preferably Arg or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(36)..(36) <223> OTHER INFORMATION: Xaa at position 36 can be
any naturally
occurring amino acid, preferably Phe <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (37)..(37)
<223> OTHER INFORMATION: Xaa at position 37 can be any
naturally occurring amino acid, preferably Tyr or Phe, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (49)..(49) <223> OTHER
INFORMATION: Xaa at position 47 can be any naturally occurring
amino acid, preferably Ser or Gly, more preferably Ser <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(63)..(63) <223> OTHER INFORMATION: Xaa at position 63 can be
any naturally occurring amino acid, preferably Ala or Tyr, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa at position 68 can be any naturally occurring
amino acid, preferably Glu or Lys, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(69)..(69) <223> OTHER INFORMATION: Xaa at position 69 can be
any naturally occurring amino acid, preferably Met or Phe, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (72)..(72) <223> OTHER
INFORMATION: Xaa at position 72 can be any naturally occurring
amino acid, preferably Asp or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(82)..(82) <223> OTHER INFORMATION: Xaa at position 82 can be
any naturally occurring amino acid, preferably Ile or Val, more
preferably Ile <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (83)..(83) <223> OTHER
INFORMATION: Xaa at position 83 can be any naturally occurring
amino acid, preferably Lys or Gln, more preferably Lys <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(86)..(86) <223> OTHER INFORMATION: Xaa at position 86 can be
any naturally occurring amino acid, preferably Ala or Ser, more
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (87)..(87) <223> OTHER
INFORMATION: Xaa at position 87 can be any naturally occurring
amino acid, preferably Glu or Gln, more preferably Gln <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(94)..(94) <223> OTHER INFORMATION: Xaa at position 94 can be
any naturally occurring amino acid, preferably Pro or Gln, more
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (95)..(95) <223> OTHER
INFORMATION: Xaa at position 95 can be any naturally occurring
amino acid, preferably Glu or Asp, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(98)..(98) <223> OTHER INFORMATION: Xaa at position 98 can be
any naturally occurring amino acid, preferably Glu or Lys, more
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (101)..(101) <223> OTHER
INFORMATION: Xaa at position 101 can be any naturally occurring
amino acid, preferably Met or Leu, more preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(103)..(103) <223> OTHER INFORMATION: Xaa at position 103 can
be any naturally occurring amino acid, preferably Met or Leu, more
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (106)..(106) <223> OTHER
INFORMATION: Xaa at position 106 can be any naturally occurring
amino acid, preferably Lys or Asn, preferably Lys <220>
FEATURE: <221> NAME/KEY: misc_feature <222> LOCATION:
(122)..(122) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (126)..(126)
<223> OTHER INFORMATION: Xaa at position 126 can be any
naturally occurring amino acid, preferably Leu <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(132)..(132) <223> OTHER INFORMATION: Xaa at position 132 can
be any naturally occurring amino acid, preferably Ser or Thr, more
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (134)..(134) <223> OTHER
INFORMATION: Xaa at position 134 can be any naturally occurring
amino acid, preferably Tyr or Phe, more preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(137)..(137) <223> OTHER INFORMATION: Xaa at position 137 can
be any naturally occurring amino acid, preferably Lys or Glu, more
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (147)..(147) <223> OTHER
INFORMATION: Xaa at position 147 can be any naturally occurring
amino acid, preferably Asp or Asn, more preferably Asp <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(153)..(153) <223> OTHER INFORMATION: Xaa at position 153 can
be any naturally occurring amino acid, preferably Arg or His, more
preferably His <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (155)..(155) <223> OTHER
INFORMATION: Xaa at position 155 can be any naturally occurring
amino acid, preferably Leu or Val, more preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(161)..(161) <223> OTHER INFORMATION: Xaa at position 161 can
be absent or any naturally occurring amino acid, preferably Ala
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (163)..(163) <223> OTHER INFORMATION: Xaa at
position 163 can be absent or any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (166)..(166) <223> OTHER
INFORMATION: Xaa at position 166 can be absent or any naturally
occurring amino acid, preferably Pro <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(168)..(168) <223> OTHER INFORMATION: Xaa at position 168 can
be any naturally occurring amino acid, preferably Gly or Ala, more
preferably Ala <400> SEQUENCE: 9 Met Thr Ser Gln Ile Arg Gln
Asn Tyr Ser Thr Xaa Val Glu Ala Ala 1 5 10 15 Val Asn Xaa Leu Val
Asn Leu Xaa Leu Xaa Ala Ser Tyr Thr Tyr Leu 20 25 30 Ser Leu Gly
Xaa Xaa Phe Asp Arg Asp Asp Val Ala Leu Glu Gly Val 35 40 45 Xaa
His Phe Phe Arg Glu Leu Ala Glu Glu Lys Arg Glu Gly Xaa Glu 50 55
60 Arg Leu Leu Xaa Xaa Gln Asn Xaa Arg Gly Gly Arg Ala Leu Phe Gln
65 70 75 80 Asp Xaa Xaa Lys Pro Xaa Xaa Asp Glu Trp Gly Lys Thr Xaa
Xaa Ala 85 90 95 Met Xaa Ala Ala Xaa Ala Xaa Glu Lys Xaa Leu Asn
Gln Ala Leu Leu 100 105 110 Asp Leu His Ala Leu Gly Ser Ala Arg Xaa
Asp Pro His Xaa Cys Asp 115 120 125 Phe Leu Glu Xaa His Xaa Leu Asp
Xaa Glu Val Lys Leu Ile Lys Lys 130 135 140 Met Gly Xaa His Leu Thr
Asn Leu Xaa Arg Xaa Ala Gly Pro Gln Pro 145 150 155 160 Xaa Gln Xaa
Gly Val Xaa Gln Xaa Ser Leu Gly Glu Tyr Leu Phe Glu 165 170 175 Arg
Leu Thr Leu Lys His Asp 180 <210> SEQ ID NO 10 <211>
LENGTH: 150 <212> TYPE: PRT <213> ORGANISM: Mus
musculus <220> FEATURE: <221> NAME/KEY: MISC_FEATURE
<222> LOCATION: (12)..(12) <223> OTHER INFORMATION: Xaa
at position 5 can be any naturally occuring amino acid, preferably
Asp or Glu, more preferably Asp <400> SEQUENCE: 10 Asn Tyr
Ser Thr Glu Val Glu Ala Ala Val Asn Arg Leu Val Asn Leu 1 5 10 15
His Leu Arg Ala Ser Tyr Thr Tyr Leu Ser Leu Gly Phe Phe Phe Asp 20
25 30 Arg Asp Asp Val Ala Leu Glu Gly Val Gly His Phe Phe Arg Glu
Leu 35 40 45 Ala Glu Glu Lys Arg Glu Gly Ala Glu Arg Leu Leu Glu
Phe Gln Asn 50 55 60 Asp Arg Gly Gly Arg Ala Leu Phe Gln Asp Val
Gln Lys Pro Ser Gln 65 70 75 80 Asp Glu Trp Gly Lys Thr Gln Glu Ala
Met Glu Ala Ala Leu Ala Met 85 90 95 Glu Lys Asn Leu Asn Gln Ala
Leu Leu Asp Leu His Ala Leu Gly Ser 100 105 110 Ala Arg Ala Asp Pro
His Leu Cys Asp Phe Leu Glu Ser His Tyr Leu 115 120 125 Asp Lys Glu
Val Lys Leu Ile Lys Lys Met Gly Asn His Leu Thr Asn 130 135 140 Leu
Arg Arg Val Ala Gly 145 150 <210> SEQ ID NO 11 <400>
SEQUENCE: 11 000 <210> SEQ ID NO 12 <211> LENGTH: 149
<212> TYPE: PRT <213> ORGANISM: Homo sapiens
<400> SEQUENCE: 12 Asn Tyr Ser Thr Asp Val Glu Ala Ala Val
Asn Ser Leu Val Asn Leu 1 5 10 15 Tyr Leu Gln Ala Ser Tyr Thr Tyr
Leu Ser Leu Gly Phe Tyr Phe Asp 20 25 30 Arg Asp Asp Val Ala Leu
Glu Gly Val Ser His Phe Phe Arg Glu Leu 35 40 45 Ala Glu Glu Lys
Arg Glu Gly Tyr Glu Arg Leu Leu Lys Met Gln Asn 50 55 60
Gln Arg Gly Gly Arg Ala Leu Phe Gln Asp Ile Lys Lys Pro Ala Glu 65
70 75 80 Asp Glu Trp Gly Lys Thr Pro Asp Ala Met Lys Ala Ala Met
Ala Leu 85 90 95 Glu Lys Lys Leu Asn Gln Ala Leu Leu Asp Leu His
Ala Leu Gly Ser 100 105 110 Ala Arg Thr Asp Pro His Leu Cys Asp Phe
Leu Glu Thr His Phe Leu 115 120 125 Asp Glu Glu Val Lys Leu Ile Lys
Lys Met Gly Asp His Leu Thr Asn 130 135 140 Leu His Arg Leu Gly 145
<210> SEQ ID NO 13 <211> LENGTH: 175 <212> TYPE:
PRT <213> ORGANISM: Homo sapiens <400> SEQUENCE: 13 Met
Ser Ser Gln Ile Arg Gln Asn Tyr Ser Thr Asp Val Glu Ala Ala 1 5 10
15 Val Asn Ser Leu Val Asn Leu Tyr Leu Gln Ala Ser Tyr Thr Tyr Leu
20 25 30 Ser Leu Gly Phe Tyr Phe Asp Arg Asp Asp Val Ala Leu Glu
Gly Val 35 40 45 Ser His Phe Phe Arg Glu Leu Ala Glu Glu Lys Arg
Glu Gly Tyr Glu 50 55 60 Arg Leu Leu Lys Met Gln Asn Gln Arg Gly
Gly Arg Ala Leu Phe Gln 65 70 75 80 Asp Ile Lys Lys Pro Ala Glu Asp
Glu Trp Gly Lys Thr Pro Asp Ala 85 90 95 Met Lys Ala Ala Met Ala
Leu Glu Lys Lys Leu Asn Gln Ala Leu Leu 100 105 110 Asp Leu His Ala
Leu Gly Ser Ala Arg Thr Asp Pro His Leu Cys Asp 115 120 125 Phe Leu
Glu Thr His Phe Leu Asp Glu Glu Val Lys Leu Ile Lys Lys 130 135 140
Met Gly Asp His Leu Thr Asn Leu His Arg Leu Gly Gly Pro Glu Ala 145
150 155 160 Gly Leu Gly Glu Tyr Leu Phe Glu Arg Leu Thr Leu Lys His
Asp 165 170 175 <210> SEQ ID NO 14 <211> LENGTH: 183
<212> TYPE: PRT <213> ORGANISM: Artificial Sequence
<220> FEATURE: <223> OTHER INFORMATION: Consensus
seqeuence of mammaliam ferritin L chain 2nd version <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(36)..(36) <223> OTHER INFORMATION: Xaa at position 36 can be
any naturally occuring amino acid, preferably Phe <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(37)..(37) <223> OTHER INFORMATION: Xaa at position 37 can be
any naturally occuring amino acid, preferably Tyr or Phe, more
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (94)..(94) <223> OTHER
INFORMATION: Xaa at position 94 can be any naturally occurring
amino acid, preferably Pro or Gln, more preferably Pro <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(126)..(126) <223> OTHER INFORMATION: Xaa at position 126 can
be any naturally occurring amino acid, preferably Leu <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(137)..(137) <223> OTHER INFORMATION: Xaa at position 137 can
be any naturally occurring amino acid, preferably Lys or Glu, more
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (147)..(147) <223> OTHER
INFORMATION: Xaa at position 147 can be any naturally occurring
amino acid, preferably Asp or Asn, more preferably Asp <220>
FEATURE: <221> NAME/KEY: misc_feature <222> LOCATION:
(161)..(161) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (163)..(163)
<223> OTHER INFORMATION: Xaa at position 163 can be absent or
any naturally occurring amino acid, preferably Thr <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(166)..(166) <223> OTHER INFORMATION: Xaa at position 166 can
be absent or any naturally occurring amino acid, preferably Pro
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (168)..(168) <223> OTHER INFORMATION: Xaa at
position 168 can be any naturally occurring amino acid, preferably
Gly or Ala, more preferably Ala <400> SEQUENCE: 14 Met Thr
Ser Gln Ile Arg Gln Asn Tyr Ser Thr Glu Val Glu Ala Ala 1 5 10 15
Val Asn Arg Leu Val Asn Leu His Leu Arg Ala Ser Tyr Thr Tyr Leu 20
25 30 Ser Leu Gly Xaa Xaa Phe Asp Arg Asp Asp Val Ala Leu Glu Gly
Val 35 40 45 Gly His Phe Phe Arg Glu Leu Ala Glu Glu Lys Arg Glu
Gly Ala Glu 50 55 60 Arg Leu Leu Lys Leu Gln Asn Gln Arg Gly Gly
Arg Ala Leu Phe Gln 65 70 75 80 Asp Val Gln Lys Pro Ala Gln Asp Glu
Trp Gly Lys Thr Xaa Glu Ala 85 90 95 Met Glu Ala Ala Leu Ala Leu
Glu Lys Asn Leu Asn Gln Ala Leu Leu 100 105 110 Asp Leu His Ala Leu
Gly Ser Ala Arg Thr Asp Pro His Xaa Cys Asp 115 120 125 Phe Leu Glu
Asn His Phe Leu Asp Xaa Glu Val Lys Leu Ile Lys Lys 130 135 140 Met
Gly Xaa His Leu Thr Asn Leu Arg Arg Leu Ala Gly Pro Gln Pro 145 150
155 160 Xaa Gln Xaa Gly Val Xaa Gln Xaa Ser Leu Gly Glu Tyr Leu Phe
Glu 165 170 175 Arg Leu Thr Leu Lys His Asp 180 <210> SEQ ID
NO 15 <211> LENGTH: 117 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Minimal consensus sequence of mammalian
haemoglobin alpha chain <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (5)..(5) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (9)..(9) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (21)..(21) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (22)..(22) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (38)..(38) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (41)..(41) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (48)..(48) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (49)..(49) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (55)..(55) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (59)..(59) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Aap <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (60)..(60) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (76)..(76) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE
<222> LOCATION: (79)..(79) <223> OTHER INFORMATION: Xaa
can be any naturally occurring amino acid, preferably Pro
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (90)..(90) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Leu <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (91)..(91)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Val <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (94)..(94) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (102)..(102) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (108)..(108) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (117)..(117) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <400> SEQUENCE: 15 Val Gly Ala His Xaa Gly Glu
Tyr Xaa Ala Glu Ala Leu Glu Xaa Met 1 5 10 15 Phe Leu Ser Phe Xaa
Xaa Thr Lys Thr Tyr Phe Pro His Phe Asp Leu 20 25 30 Ser His Gly
Ser Ala Xaa Val Lys Xaa His Gly Lys Lys Val Ala Xaa 35 40 45 Xaa
Leu Thr Asn Ala Val Xaa His Val Asp Xaa Xaa Pro Asn Ala Leu 50 55
60 Ser Ala Leu Xaa Asp Leu His Ala His Lys Leu Xaa Val Asp Xaa Val
65 70 75 80 Asn Phe Lys Leu Leu Ser His Cys Leu Xaa Xaa Thr Leu Xaa
Ala His 85 90 95 Leu Pro Ala Glu Phe Xaa Pro Ala Val His Ala Xaa
Leu Asp Lys Phe 100 105 110 Leu Ala Ser Val Xaa 115 <210> SEQ
ID NO 16 <211> LENGTH: 142 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Consensus sequence of mammalian alpha chain
haemoglobins <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (1)..(1) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (2)..(2) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (14)..(14) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (17)..(17) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (22)..(22) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (26)..(26) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (32)..(32) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (38)..(38) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (39)..(39) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (55)..(55) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (58)..(58) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (66)..(66) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (72)..(72) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (76)..(76) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (77)..(77) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (85)..(85) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (93)..(93) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (96)..(96) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (107)..(107) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (108)..(108) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (111)..(111) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (119)..(119) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (125)..(125) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (134)..(134) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (142)..(142) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <400> SEQUENCE: 16 Xaa Xaa Leu Ser Pro Ala Asp
Lys Thr Asn Val Lys Ala Xaa Trp Gly 1 5 10 15 Xaa Val Gly Ala His
Xaa Gly Glu Tyr Xaa Ala Glu Ala Leu Glu Xaa 20 25 30 Met Phe Leu
Ser Phe Xaa Xaa Thr Lys Thr Tyr Phe Pro His Phe Asp 35 40 45 Leu
Ser His Gly Ser Ala Xaa Val Lys Xaa His Gly Lys Lys Val Ala 50 55
60 Xaa Xaa Leu Thr Asn Ala Val Xaa His Val Asp Xaa Xaa Pro Asn Ala
65 70 75 80 Leu Ser Ala Leu Xaa Asp Leu His Ala His Lys Leu Xaa Val
Asp Xaa
85 90 95 Val Asn Phe Lys Leu Leu Ser His Cys Leu Xaa Xaa Thr Leu
Xaa Ala 100 105 110 His Leu Pro Ala Glu Phe Xaa Pro Ala Val His Ala
Xaa Leu Asp Lys 115 120 125 Phe Leu Ala Ser Val Xaa Thr Val Leu Thr
Ser Lys Tyr Xaa 130 135 140 <210> SEQ ID NO 17 <211>
LENGTH: 117 <212> TYPE: PRT <213> ORGANISM: Homo
sapiens <400> SEQUENCE: 17 Val Gly Ala His Ala Gly Glu Tyr
Gly Ala Glu Ala Leu Glu Arg Met 1 5 10 15 Phe Leu Ser Phe Pro Thr
Thr Lys Thr Tyr Phe Pro His Phe Asp Leu 20 25 30 Ser His Gly Ser
Ala Gln Val Lys Gly His Gly Lys Lys Val Ala Asp 35 40 45 Ala Leu
Thr Asn Ala Val Ala His Val Asp Asp Met Pro Asn Ala Leu 50 55 60
Ser Ala Leu Ser Asp Leu His Ala His Lys Leu Arg Val Asp Pro Val 65
70 75 80 Asn Phe Lys Leu Leu Ser His Cys Leu Leu Val Thr Leu Ala
Ala His 85 90 95 Leu Pro Ala Glu Phe Thr Pro Ala Val His Ala Ser
Leu Asp Lys Phe 100 105 110 Leu Ala Ser Val Ser 115 <210> SEQ
ID NO 18 <211> LENGTH: 142 <212> TYPE: PRT <213>
ORGANISM: Homo sapiens <400> SEQUENCE: 18 Met Val Leu Ser Pro
Ala Asp Lys Thr Asn Val Lys Ala Ala Trp Gly 1 5 10 15 Lys Val Gly
Ala His Ala Gly Glu Tyr Gly Ala Glu Ala Leu Glu Arg 20 25 30 Met
Phe Leu Ser Phe Pro Thr Thr Lys Thr Tyr Phe Pro His Phe Asp 35 40
45 Leu Ser His Gly Ser Ala Gln Val Lys Gly His Gly Lys Lys Val Ala
50 55 60 Asp Ala Leu Thr Asn Ala Val Ala His Val Asp Asp Met Pro
Asn Ala 65 70 75 80 Leu Ser Ala Leu Ser Asp Leu His Ala His Lys Leu
Arg Val Asp Pro 85 90 95 Val Asn Phe Lys Leu Leu Ser His Cys Leu
Leu Val Thr Leu Ala Ala 100 105 110 His Leu Pro Ala Glu Phe Thr Pro
Ala Val His Ala Ser Leu Asp Lys 115 120 125 Phe Leu Ala Ser Val Ser
Thr Val Leu Thr Ser Lys Tyr Arg 130 135 140 <210> SEQ ID NO
19 <211> LENGTH: 124 <212> TYPE: PRT <213>
ORGANISM: Artificial Sequence <220> FEATURE: <223>
OTHER INFORMATION: Minimal consensus sequence of mammalian
haemoglobin beta chain <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (4)..(4) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (9)..(9) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (10)..(10) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (12)..(12) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (13)..(13) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (14)..(14) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (27)..(27) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (35)..(35) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (36)..(36) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (49)..(49) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (52)..(52) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (66)..(66) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (69)..(69) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (86)..(86) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Glu <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (121)..(121) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (122)..(122) <223> OTHER INFORMATION: Xaa can be
absent or any naturally occurring amino acid, preferably Val
<400> SEQUENCE: 19 Gly Lys Val Xaa Val Asp Glu Val Xaa Xaa
Glu Xaa Xaa Xaa Xaa Leu 1 5 10 15 Leu Val Val Tyr Pro Trp Thr Gln
Arg Phe Xaa Glu Ser Phe Gly Asp 20 25 30 Leu Ser Xaa Xaa Asp Ala
Val Met Gly Asn Pro Lys Val Lys Ala His 35 40 45 Xaa Lys Lys Xaa
Leu Gly Ala Phe Ser Asp Gly Leu Ala His Leu Asp 50 55 60 Asn Xaa
Lys Xaa Xaa Phe Ala Thr Leu Ser Glu Leu His Cys Asp Lys 65 70 75 80
Leu His Val Asp Pro Xaa Asn Phe Arg Leu Leu Gly Asn Val Leu Val 85
90 95 Cys Val Leu Ala His His Phe Gly Lys Glu Phe Thr Pro Pro Val
Gln 100 105 110 Ala Ala Tyr Gln Lys Val Val Ala Xaa Xaa Ala Asn 115
120 <210> SEQ ID NO 20 <211> LENGTH: 147 <212>
TYPE: PRT <213> ORGANISM: Artificial Sequence <220>
FEATURE: <223> OTHER INFORMATION: Consensus sequence of
mammalian haemoglobin beta chain <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (10)..(10) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (11)..(11) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (20)..(20) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino
acid, preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (25)..(25) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (26)..(26) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (28)..(28) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (29)..(29) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (30)..(30) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (31)..(31) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (43)..(43) <223> OTHER
INFORMATION: Xaa can be absent or any naturally occurring amino
acid, preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (51)..(51) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (52)..(52) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (68)..(68) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (82)..(82) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (84)..(84) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (85)..(85) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (102)..(102) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (137)..(137) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (138)..(138) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (147)..(147) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably His <400> SEQUENCE: 20 Met Val His Leu Thr Pro Glu
Glu Lys Xaa Xaa Val Thr Ala Xaa Trp 1 5 10 15 Gly Lys Val Xaa Val
Asp Glu Val Xaa Xaa Glu Xaa Xaa Xaa Xaa Leu 20 25 30 Leu Val Val
Tyr Pro Trp Thr Gln Arg Phe Xaa Glu Ser Phe Gly Asp 35 40 45 Leu
Ser Xaa Xaa Asp Ala Val Met Gly Asn Pro Lys Val Lys Ala His 50 55
60 Xaa Lys Lys Xaa Leu Gly Ala Phe Ser Asp Gly Leu Ala His Leu Asp
65 70 75 80 Asn Xaa Lys Xaa Xaa Phe Ala Thr Leu Ser Glu Leu His Cys
Asp Lys 85 90 95 Leu His Val Asp Pro Xaa Asn Phe Arg Leu Leu Gly
Asn Val Leu Val 100 105 110 Cys Val Leu Ala His His Phe Gly Lys Glu
Phe Thr Pro Pro Val Gln 115 120 125 Ala Ala Tyr Gln Lys Val Val Ala
Xaa Xaa Ala Asn Ala Leu Ala His 130 135 140 Lys Tyr Xaa 145
<210> SEQ ID NO 21 <211> LENGTH: 124 <212> TYPE:
PRT <213> ORGANISM: Homo sapiens <400> SEQUENCE: 21 Gly
Lys Val Asn Val Asp Glu Val Gly Gly Glu Ala Leu Gly Arg Leu 1 5 10
15 Leu Val Val Tyr Pro Trp Thr Gln Arg Phe Leu Glu Ser Phe Gly Asp
20 25 30 Leu Ser Thr Pro Asp Ala Val Met Gly Asn Pro Lys Val Lys
Ala His 35 40 45 Gly Lys Lys Val Leu Gly Ala Phe Ser Asp Gly Leu
Ala His Leu Asp 50 55 60 Asn Leu Lys Gly Thr Phe Ala Thr Leu Ser
Glu Leu His Cys Asp Lys 65 70 75 80 Leu His Val Asp Pro Glu Asn Phe
Arg Leu Leu Gly Asn Val Leu Val 85 90 95 Cys Val Leu Ala His His
Phe Gly Lys Glu Phe Thr Pro Pro Val Gln 100 105 110 Ala Ala Tyr Gln
Lys Val Val Ala Gly Val Ala Asn 115 120 <210> SEQ ID NO 22
<211> LENGTH: 147 <212> TYPE: PRT <213> ORGANISM:
Homo sapiens <400> SEQUENCE: 22 Met Val His Leu Thr Pro Glu
Glu Lys Ser Ala Val Thr Ala Leu Trp 1 5 10 15 Gly Lys Val Asn Val
Asp Glu Val Gly Gly Glu Ala Leu Gly Arg Leu 20 25 30 Leu Val Val
Tyr Pro Trp Thr Gln Arg Phe Leu Glu Ser Phe Gly Asp 35 40 45 Leu
Ser Thr Pro Asp Ala Val Met Gly Asn Pro Lys Val Lys Ala His 50 55
60 Gly Lys Lys Val Leu Gly Ala Phe Ser Asp Gly Leu Ala His Leu Asp
65 70 75 80 Asn Leu Lys Gly Thr Phe Ala Thr Leu Ser Glu Leu His Cys
Asp Lys 85 90 95 Leu His Val Asp Pro Glu Asn Phe Arg Leu Leu Gly
Asn Val Leu Val 100 105 110 Cys Val Leu Ala His His Phe Gly Lys Glu
Phe Thr Pro Pro Val Gln 115 120 125 Ala Ala Tyr Gln Lys Val Val Ala
Gly Val Ala Asn Ala Leu Ala His 130 135 140 Lys Tyr His 145
<210> SEQ ID NO 23 <211> LENGTH: 80 <212> TYPE:
PRT <213> ORGANISM: Artificial Sequence <220> FEATURE:
<223> OTHER INFORMATION: N-terminal region of a consensus
sequence of mammalian transferrins <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (2)..(2) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (9)..(9) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (10)..(10) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (24)..(24) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (25)..(25) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (31)..(31) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (32)..(32) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (42)..(42) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (47)..(47) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (59)..(59) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (64)..(64) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Tyr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (65)..(65)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Leu <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (71)..(71) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ile <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (73)..(73) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (77)..(77) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (78)..(78) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <400> SEQUENCE: 23 Met Xaa Leu Ala Val Gly Ala
Leu Xaa Xaa Cys Ala Val Leu Xaa Leu 1 5 10 15 Cys Leu Ala Val Pro
Asp Lys Xaa Xaa Arg Trp Cys Ala Val Xaa Xaa 20 25 30 His Glu Ala
Thr Lys Cys Gln Ser Phe Xaa Asp His Met Lys Xaa Val 35 40 45 Ile
Pro Ser Asp Gly Pro Ser Val Ala Cys Xaa Lys Lys Ala Ser Xaa 50 55
60 Xaa Asp Cys Ile Arg Ala Xaa Ala Xaa Asn Glu Ala Xaa Xaa Val Thr
65 70 75 80 <210> SEQ ID NO 24 <211> LENGTH: 114
<212> TYPE: PRT <213> ORGANISM: Artificial Sequence
<220> FEATURE: <223> OTHER INFORMATION: C-terminal
region of a consensus sequence of mammalian transferrins
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (3)..(3) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Thr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (16)..(16)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Asp <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (18)..(18) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (19)..(19) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (23)..(23) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (25)..(25) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (28)..(28) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (30)..(30) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (61)..(61) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (62)..(62) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (75)..(75) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (78)..(78) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (96)..(96) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (97)..(97) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (105)..(105) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (107)..(107) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (110)..(110) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <400> SEQUENCE: 24 Glu Thr Xaa Glu Asp Cys Ile
Ala Lys Ile Met Asn Gly Glu Ala Xaa 1 5 10 15 Ala Xaa Xaa Leu Asp
Gly Xaa Phe Xaa Tyr Ile Xaa Gly Xaa Cys Gly 20 25 30 Leu Val Pro
Val Leu Ala Glu Asn Tyr Asn Lys Ser Asp Asn Cys Glu 35 40 45 Asp
Thr Pro Glu Ala Gly Tyr Phe Ala Val Ala Val Xaa Xaa Lys Ser 50 55
60 Ala Ser Asp Leu Thr Trp Asp Asn Leu Lys Xaa Lys Lys Xaa Cys His
65 70 75 80 Thr Ala Val Gly Arg Thr Ala Gly Trp Asn Ile Pro Met Gly
Leu Xaa 85 90 95 Xaa Asn Lys Ile Asn His Cys Arg Xaa Asp Xaa Phe
Phe Xaa Glu Gly 100 105 110 Cys Ala <210> SEQ ID NO 25
<211> LENGTH: 698 <212> TYPE: PRT <213> ORGANISM:
Artificial Sequence <220> FEATURE: <223> OTHER
INFORMATION: Consensus sequence of mammalian transferrins
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (2)..(2) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Arg <220> FEATURE:
<221> NAME/KEY: misc_feature <222> LOCATION: (9)..(9)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid <220> FEATURE: <221> NAME/KEY: MISC_FEATURE
<222> LOCATION: (10)..(10) <223> OTHER INFORMATION: Xaa
can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (15)..(15) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (24)..(24) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (25)..(25) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (31)..(31) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (32)..(32) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (42)..(42) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (47)..(47) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (59)..(59) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (64)..(64) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (65)..(65) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (71)..(71) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (73)..(73) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Ile <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Xaa can be any naturally occurring
amino acid, preferably Asp <220> FEATURE: <221>
NAME/KEY: MISC_FEATURE <222> LOCATION: (78)..(78) <223>
OTHER INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (84)..(84) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (87)..(87) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (88)..(88) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (94)..(94) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (98)..(98) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (104)..(104) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (107)..(107) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (108)..(108) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (115)..(115) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (116)..(116) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (121)..(121) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (124)..(124) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (129)..(129) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (146)..(146) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (152)..(152) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (153)..(153) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (161)..(161) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (179)..(179) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (188)..(188) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gln <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (204)..(204) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (224)..(224) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (238)..(238) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (245)..(245) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (255)..(255) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (268)..(268) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably His <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (275)..(275) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (276)..(276) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (286)..(286) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (293)..(293) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (309)..(309) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (310)..(310) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (326)..(326) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (327)..(327) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (357)..(357) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (381)..(381) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (393)..(393) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Thr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (406)..(406) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asp <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (408)..(408) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Met <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (409)..(409) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (413)..(413) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (415)..(415) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (418)..(418) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ala <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (420)..(420) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (451)..(451) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Val <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (452)..(452) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (465)..(465) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Gly <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (468)..(468) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (486)..(486) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Leu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (487)..(487) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (495)..(495) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Phe <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (497)..(497) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Glu <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (500)..(500) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Ser <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (505)..(505) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (509)..(509) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Lys <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (529)..(529) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Asn <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (536)..(536) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Tyr <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (558)..(558) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <220> FEATURE: <221> NAME/KEY:
misc_feature <222> LOCATION: (559)..(559) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid
<220> FEATURE: <221> NAME/KEY: MISC_FEATURE <222>
LOCATION: (563)..(563) <223> OTHER INFORMATION: Xaa can be
any naturally occurring amino acid, preferably Gly <220>
FEATURE: <221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(586)..(586) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Thr <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(592)..(592) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Glu <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(603)..(603) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Asn <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(607)..(607) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Val <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(622)..(622) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Gln <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(635)..(635) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Ser <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(648)..(648) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Leu <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(652)..(652) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Asp <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(678)..(678) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Val <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(692)..(692) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid, preferably Ala <220> FEATURE:
<221> NAME/KEY: MISC_FEATURE <222> LOCATION:
(696)..(696) <223> OTHER INFORMATION: Xaa can be any
naturally occurring amino acid,
preferably Arg <220> FEATURE: <221> NAME/KEY:
MISC_FEATURE <222> LOCATION: (698)..(698) <223> OTHER
INFORMATION: Xaa can be any naturally occurring amino acid,
preferably Pro <400> SEQUENCE: 25 Met Xaa Leu Ala Val Gly Ala
Leu Xaa Xaa Cys Ala Val Leu Xaa Leu 1 5 10 15 Cys Leu Ala Val Pro
Asp Lys Xaa Xaa Arg Trp Cys Ala Val Xaa Xaa 20 25 30 His Glu Ala
Thr Lys Cys Gln Ser Phe Xaa Asp His Met Lys Xaa Val 35 40 45 Ile
Pro Ser Asp Gly Pro Ser Val Ala Cys Xaa Lys Lys Ala Ser Xaa 50 55
60 Xaa Asp Cys Ile Arg Ala Xaa Ala Xaa Asn Glu Ala Xaa Xaa Val Thr
65 70 75 80 Leu Asp Ala Xaa Leu Val Xaa Xaa Ala Tyr Leu Ala Pro Xaa
Asn Leu 85 90 95 Lys Xaa Val Val Ala Glu Phe Xaa Gly Ser Xaa Xaa
Asp Pro Gln Thr 100 105 110 Phe Tyr Xaa Xaa Val Ala Val Val Xaa Lys
Asp Xaa Gly Phe Gln Met 115 120 125 Xaa Gln Leu Arg Gly Lys Lys Ser
Cys His Thr Gly Leu Gly Arg Ser 130 135 140 Ala Xaa Trp Asn Ile Pro
Ile Xaa Xaa Leu Tyr Cys Asp Leu Pro Glu 145 150 155 160 Xaa Arg Lys
Pro Leu Glu Lys Ala Val Ala Asn Phe Phe Ser Gly Ser 165 170 175 Cys
Ala Xaa Cys Ala Asp Gly Thr Asp Phe Pro Xaa Leu Cys Gln Leu 180 185
190 Cys Pro Gly Cys Gly Cys Ser Thr Leu Asn Gln Xaa Phe Gly Tyr Ser
195 200 205 Gly Ala Phe Lys Cys Leu Lys Asp Gly Ala Gly Asp Val Ala
Phe Xaa 210 215 220 Lys His Ser Thr Ile Phe Glu Asn Leu Ala Asn Lys
Ala Xaa Arg Asp 225 230 235 240 Gln Tyr Glu Leu Xaa Cys Leu Asp Asn
Thr Arg Lys Pro Val Xaa Glu 245 250 255 Tyr Lys Asp Cys His Leu Ala
Gln Val Pro Ser Xaa Thr Val Val Ala 260 265 270 Arg Ser Xaa Xaa Gly
Lys Glu Asp Leu Ile Trp Glu Leu Xaa Asn Gln 275 280 285 Ala Gln Glu
His Xaa Gly Lys Asp Lys Ser Lys Glu Phe Gln Leu Phe 290 295 300 Ser
Ser Pro His Xaa Xaa Asp Leu Leu Phe Lys Asp Ser Ala His Gly 305 310
315 320 Phe Leu Lys Val Pro Xaa Xaa Met Asp Ala Lys Met Tyr Leu Gly
Tyr 325 330 335 Glu Tyr Val Thr Ala Ile Arg Asn Leu Arg Glu Gly Thr
Cys Pro Glu 340 345 350 Ala Pro Thr Asp Xaa Cys Lys Pro Val Lys Trp
Cys Ala Leu Ser His 355 360 365 His Glu Arg Leu Lys Cys Asp Glu Trp
Ser Val Asn Xaa Val Gly Lys 370 375 380 Ile Glu Cys Val Ser Ala Glu
Thr Xaa Glu Asp Cys Ile Ala Lys Ile 385 390 395 400 Met Asn Gly Glu
Ala Xaa Ala Xaa Xaa Leu Asp Gly Xaa Phe Xaa Tyr 405 410 415 Ile Xaa
Gly Xaa Cys Gly Leu Val Pro Val Leu Ala Glu Asn Tyr Asn 420 425 430
Lys Ser Asp Asn Cys Glu Asp Thr Pro Glu Ala Gly Tyr Phe Ala Val 435
440 445 Ala Val Xaa Xaa Lys Ser Ala Ser Asp Leu Thr Trp Asp Asn Leu
Lys 450 455 460 Xaa Lys Lys Xaa Cys His Thr Ala Val Gly Arg Thr Ala
Gly Trp Asn 465 470 475 480 Ile Pro Met Gly Leu Xaa Xaa Asn Lys Ile
Asn His Cys Arg Xaa Asp 485 490 495 Xaa Phe Phe Xaa Glu Gly Cys Ala
Xaa Gly Ser Lys Xaa Asp Ser Ser 500 505 510 Leu Cys Lys Leu Cys Met
Gly Ser Gly Leu Asn Leu Cys Glu Pro Asn 515 520 525 Xaa Lys Glu Gly
Tyr Tyr Gly Xaa Thr Gly Ala Phe Arg Cys Leu Val 530 535 540 Glu Lys
Gly Asp Val Ala Phe Val Lys His Gln Thr Val Xaa Xaa Asn 545 550 555
560 Thr Gly Xaa Lys Asn Pro Asp Pro Trp Ala Lys Asn Leu Asn Glu Lys
565 570 575 Asp Tyr Glu Leu Leu Cys Leu Asp Gly Xaa Arg Lys Pro Val
Glu Xaa 580 585 590 Tyr Ala Asn Cys His Leu Ala Arg Ala Pro Xaa His
Ala Val Xaa Thr 595 600 605 Arg Lys Asp Lys Glu Ala Cys Val His Lys
Ile Leu Arg Xaa Gln Gln 610 615 620 His Leu Phe Gly Ser Asn Val Thr
Asp Cys Xaa Gly Asn Phe Cys Leu 625 630 635 640 Phe Arg Ser Glu Thr
Lys Asp Xaa Leu Phe Arg Xaa Asp Thr Val Cys 645 650 655 Leu Ala Lys
Leu His Asp Arg Asn Thr Tyr Glu Lys Tyr Leu Gly Glu 660 665 670 Glu
Tyr Val Lys Ala Xaa Gly Asn Leu Arg Lys Cys Ser Thr Ser Ser 675 680
685 Leu Leu Glu Xaa Cys Thr Phe Xaa Arg Xaa 690 695 <210> SEQ
ID NO 26 <211> LENGTH: 80 <212> TYPE: PRT <213>
ORGANISM: Homo sapiens <400> SEQUENCE: 26 Met Arg Leu Ala Val
Gly Ala Leu Leu Val Cys Ala Val Leu Gly Leu 1 5 10 15 Cys Leu Ala
Val Pro Asp Lys Thr Val Arg Trp Cys Ala Val Ser Glu 20 25 30 His
Glu Ala Thr Lys Cys Gln Ser Phe Arg Asp His Met Lys Ser Val 35 40
45 Ile Pro Ser Asp Gly Pro Ser Val Ala Cys Val Lys Lys Ala Ser Tyr
50 55 60 Leu Asp Cys Ile Arg Ala Ile Ala Ala Asn Glu Ala Asp Ala
Val Thr 65 70 75 80 <210> SEQ ID NO 27 <211> LENGTH:
114 <212> TYPE: PRT <213> ORGANISM: Homo sapiens
<400> SEQUENCE: 27 Glu Thr Thr Glu Asp Cys Ile Ala Lys Ile
Met Asn Gly Glu Ala Asp 1 5 10 15 Ala Met Ser Leu Asp Gly Gly Phe
Val Tyr Ile Ala Gly Lys Cys Gly 20 25 30 Leu Val Pro Val Leu Ala
Glu Asn Tyr Asn Lys Ser Asp Asn Cys Glu 35 40 45 Asp Thr Pro Glu
Ala Gly Tyr Phe Ala Val Ala Val Val Lys Lys Ser 50 55 60 Ala Ser
Asp Leu Thr Trp Asp Asn Leu Lys Gly Lys Lys Ser Cys His 65 70 75 80
Thr Ala Val Gly Arg Thr Ala Gly Trp Asn Ile Pro Met Gly Leu Leu 85
90 95 Tyr Asn Lys Ile Asn His Cys Arg Phe Asp Glu Phe Phe Ser Glu
Gly 100 105 110 Cys Ala <210> SEQ ID NO 28 <211>
LENGTH: 698 <212> TYPE: PRT <213> ORGANISM: Homo
sapiens <400> SEQUENCE: 28 Met Arg Leu Ala Val Gly Ala Leu
Leu Val Cys Ala Val Leu Gly Leu 1 5 10 15 Cys Leu Ala Val Pro Asp
Lys Thr Val Arg Trp Cys Ala Val Ser Glu 20 25 30 His Glu Ala Thr
Lys Cys Gln Ser Phe Arg Asp His Met Lys Ser Val 35 40 45 Ile Pro
Ser Asp Gly Pro Ser Val Ala Cys Val Lys Lys Ala Ser Tyr 50 55 60
Leu Asp Cys Ile Arg Ala Ile Ala Ala Asn Glu Ala Asp Ala Val Thr 65
70 75 80 Leu Asp Ala Gly Leu Val Tyr Asp Ala Tyr Leu Ala Pro Asn
Asn Leu 85 90 95 Lys Pro Val Val Ala Glu Phe Tyr Gly Ser Lys Glu
Asp Pro Gln Thr 100 105 110 Phe Tyr Tyr Ala Val Ala Val Val Lys Lys
Asp Ser Gly Phe Gln Met 115 120 125 Asn Gln Leu Arg Gly Lys Lys Ser
Cys His Thr Gly Leu Gly Arg Ser 130 135 140 Ala Gly Trp Asn Ile Pro
Ile Gly Leu Leu Tyr Cys Asp Leu Pro Glu 145 150 155 160 Pro Arg Lys
Pro Leu Glu Lys Ala Val Ala Asn Phe Phe Ser Gly Ser 165 170 175 Cys
Ala Pro Cys Ala Asp Gly Thr Asp Phe Pro Gln Leu Cys Gln Leu 180 185
190 Cys Pro Gly Cys Gly Cys Ser Thr Leu Asn Gln Tyr Phe Gly Tyr Ser
195 200 205 Gly Ala Phe Lys Cys Leu Lys Asp Gly Ala Gly Asp Val Ala
Phe Val 210 215 220 Lys His Ser Thr Ile Phe Glu Asn Leu Ala Asn Lys
Ala Asp Arg Asp 225 230 235 240 Gln Tyr Glu Leu Leu Cys Leu Asp Asn
Thr Arg Lys Pro Val Asp Glu 245 250 255
Tyr Lys Asp Cys His Leu Ala Gln Val Pro Ser His Thr Val Val Ala 260
265 270 Arg Ser Met Gly Gly Lys Glu Asp Leu Ile Trp Glu Leu Leu Asn
Gln 275 280 285 Ala Gln Glu His Phe Gly Lys Asp Lys Ser Lys Glu Phe
Gln Leu Phe 290 295 300 Ser Ser Pro His Gly Lys Asp Leu Leu Phe Lys
Asp Ser Ala His Gly 305 310 315 320 Phe Leu Lys Val Pro Pro Arg Met
Asp Ala Lys Met Tyr Leu Gly Tyr 325 330 335 Glu Tyr Val Thr Ala Ile
Arg Asn Leu Arg Glu Gly Thr Cys Pro Glu 340 345 350 Ala Pro Thr Asp
Glu Cys Lys Pro Val Lys Trp Cys Ala Leu Ser His 355 360 365 His Glu
Arg Leu Lys Cys Asp Glu Trp Ser Val Asn Ser Val Gly Lys 370 375 380
Ile Glu Cys Val Ser Ala Glu Thr Thr Glu Asp Cys Ile Ala Lys Ile 385
390 395 400 Met Asn Gly Glu Ala Asp Ala Met Ser Leu Asp Gly Gly Phe
Val Tyr 405 410 415 Ile Ala Gly Lys Cys Gly Leu Val Pro Val Leu Ala
Glu Asn Tyr Asn 420 425 430 Lys Ser Asp Asn Cys Glu Asp Thr Pro Glu
Ala Gly Tyr Phe Ala Val 435 440 445 Ala Val Val Lys Lys Ser Ala Ser
Asp Leu Thr Trp Asp Asn Leu Lys 450 455 460 Gly Lys Lys Ser Cys His
Thr Ala Val Gly Arg Thr Ala Gly Trp Asn 465 470 475 480 Ile Pro Met
Gly Leu Leu Tyr Asn Lys Ile Asn His Cys Arg Phe Asp 485 490 495 Glu
Phe Phe Ser Glu Gly Cys Ala Pro Gly Ser Lys Lys Asp Ser Ser 500 505
510 Leu Cys Lys Leu Cys Met Gly Ser Gly Leu Asn Leu Cys Glu Pro Asn
515 520 525 Asn Lys Glu Gly Tyr Tyr Gly Tyr Thr Gly Ala Phe Arg Cys
Leu Val 530 535 540 Glu Lys Gly Asp Val Ala Phe Val Lys His Gln Thr
Val Pro Gln Asn 545 550 555 560 Thr Gly Gly Lys Asn Pro Asp Pro Trp
Ala Lys Asn Leu Asn Glu Lys 565 570 575 Asp Tyr Glu Leu Leu Cys Leu
Asp Gly Thr Arg Lys Pro Val Glu Glu 580 585 590 Tyr Ala Asn Cys His
Leu Ala Arg Ala Pro Asn His Ala Val Val Thr 595 600 605 Arg Lys Asp
Lys Glu Ala Cys Val His Lys Ile Leu Arg Gln Gln Gln 610 615 620 His
Leu Phe Gly Ser Asn Val Thr Asp Cys Ser Gly Asn Phe Cys Leu 625 630
635 640 Phe Arg Ser Glu Thr Lys Asp Leu Leu Phe Arg Asp Asp Thr Val
Cys 645 650 655 Leu Ala Lys Leu His Asp Arg Asn Thr Tyr Glu Lys Tyr
Leu Gly Glu 660 665 670 Glu Tyr Val Lys Ala Val Gly Asn Leu Arg Lys
Cys Ser Thr Ser Ser 675 680 685 Leu Leu Glu Ala Cys Thr Phe Arg Arg
Pro 690 695
References
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012
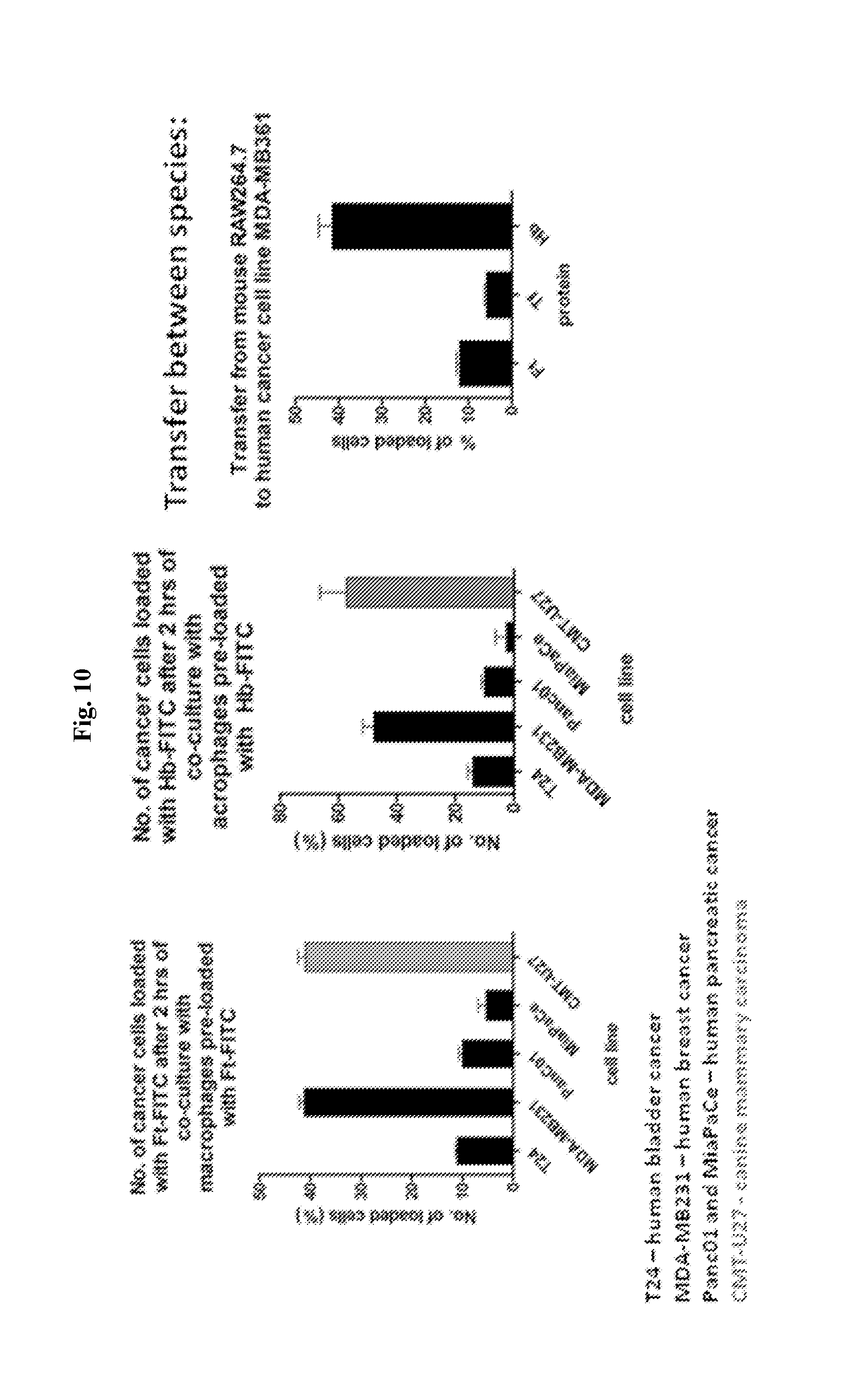

D00013
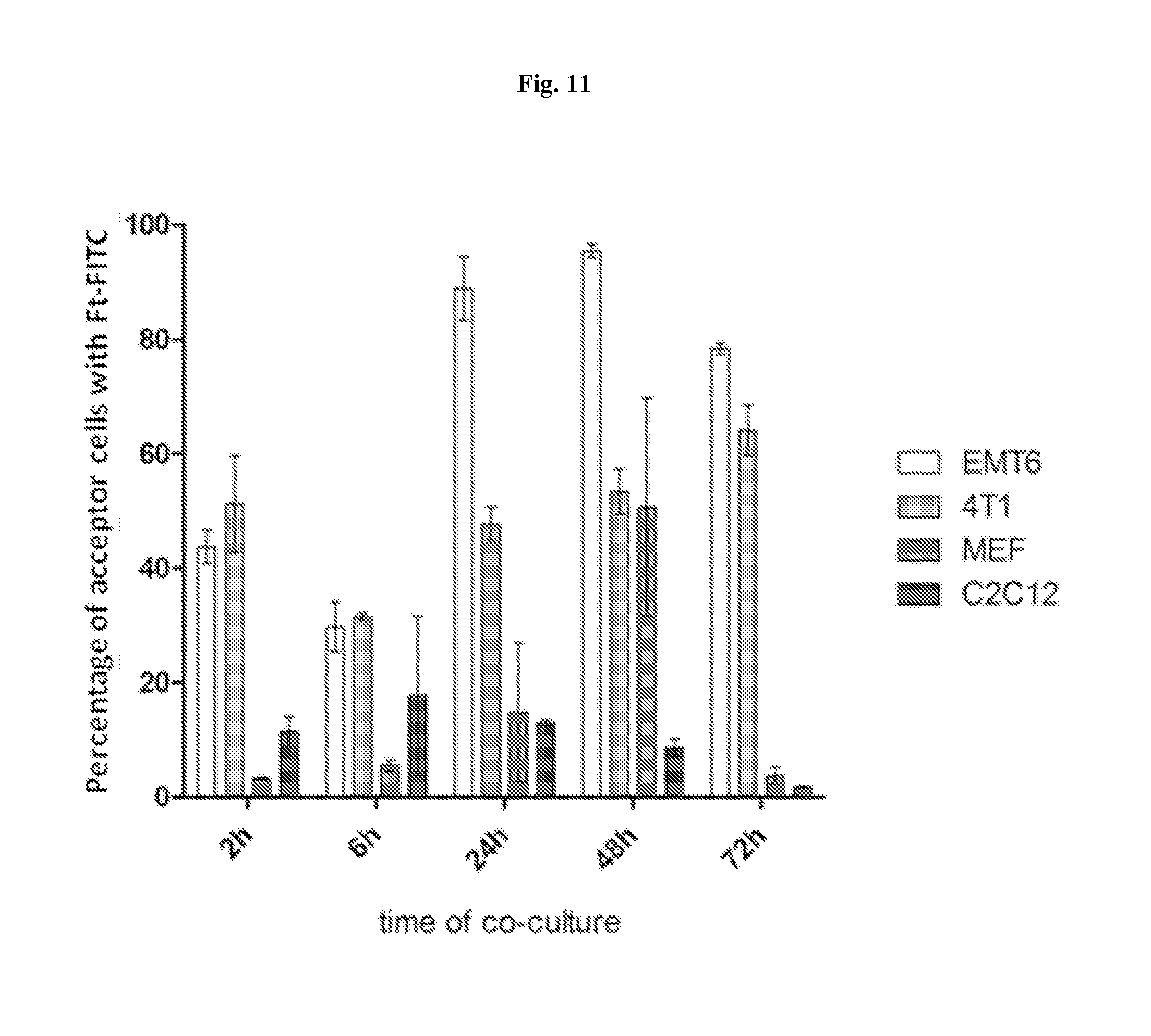
D00014

D00015
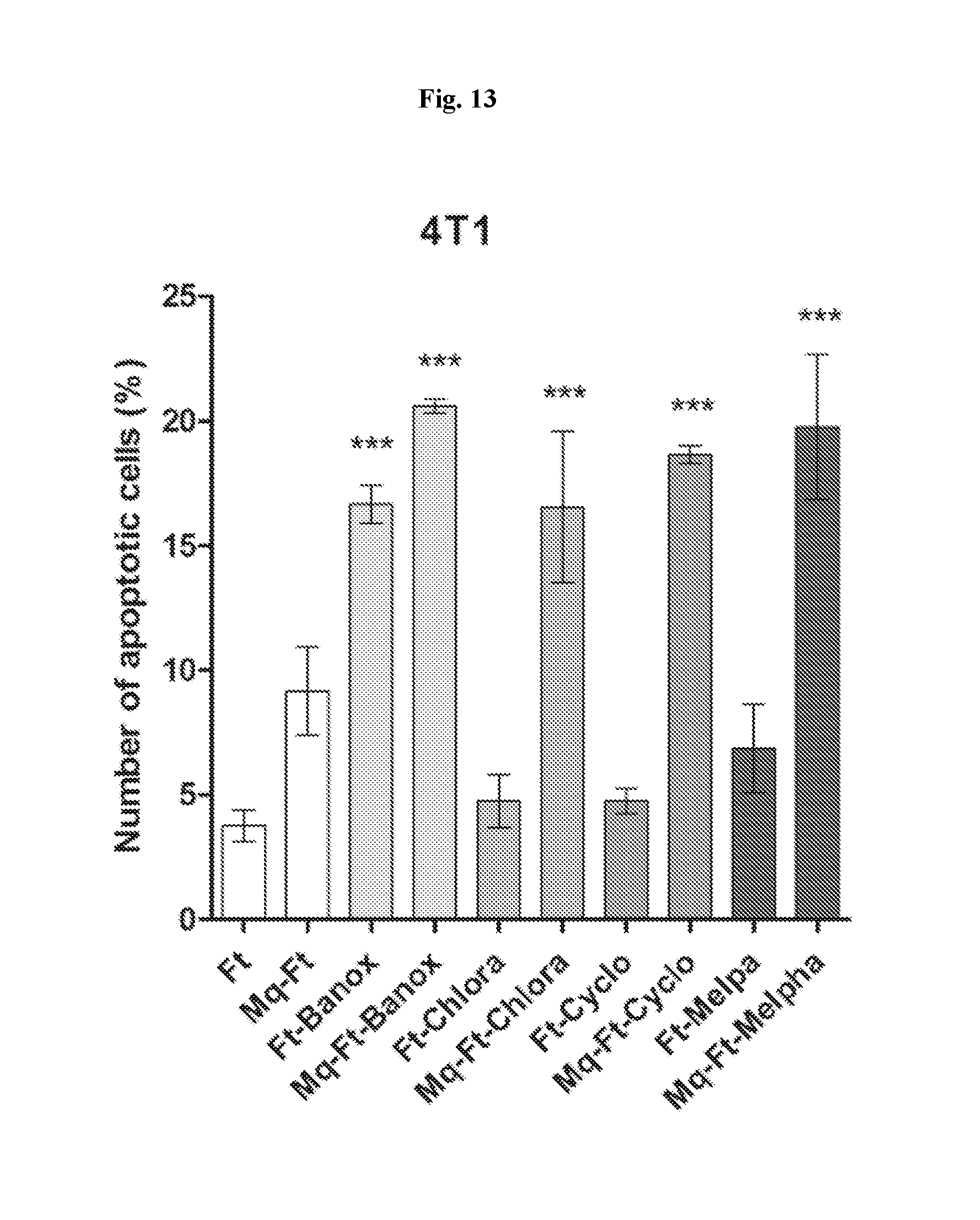
D00016
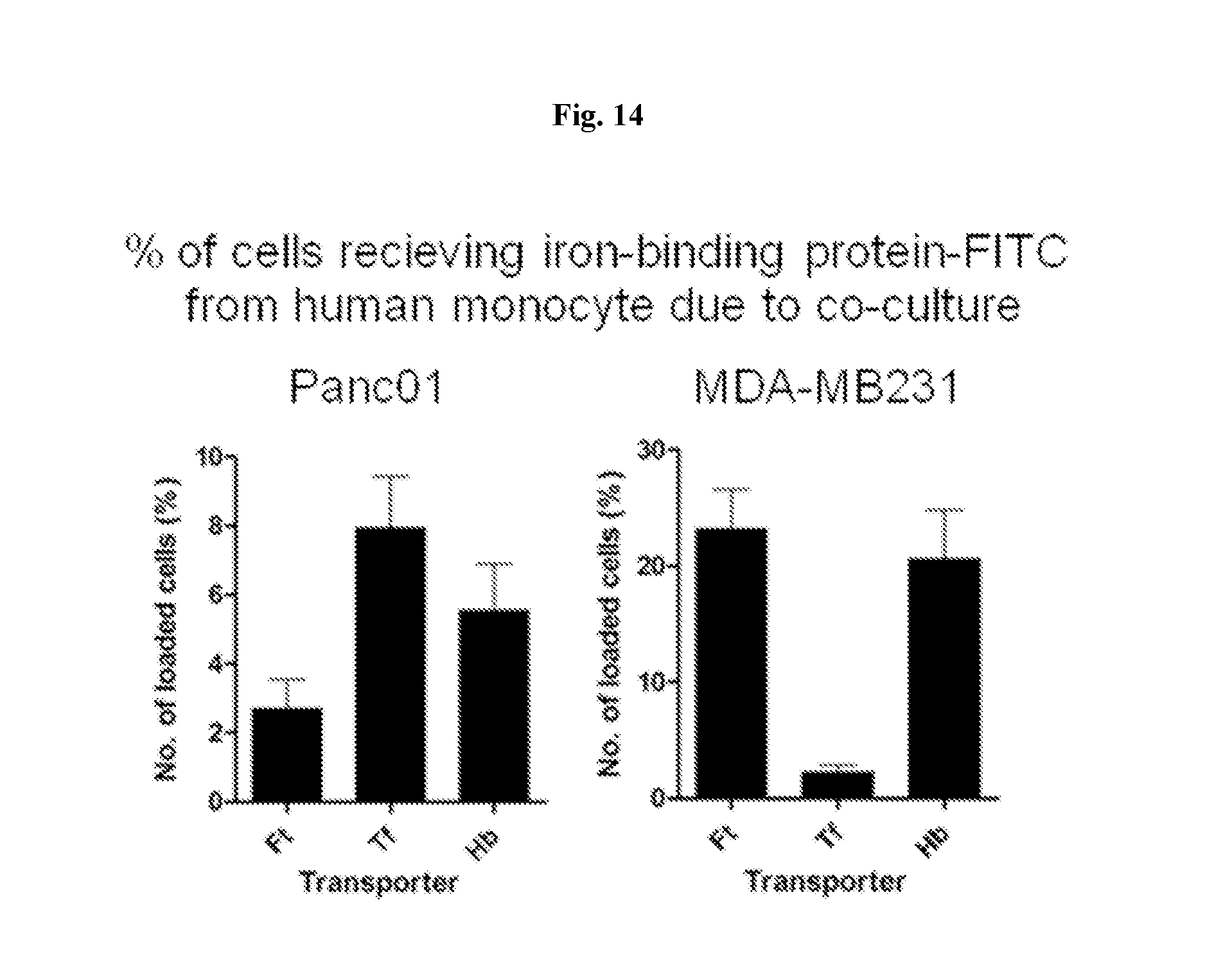
D00017

D00018

D00019

D00020
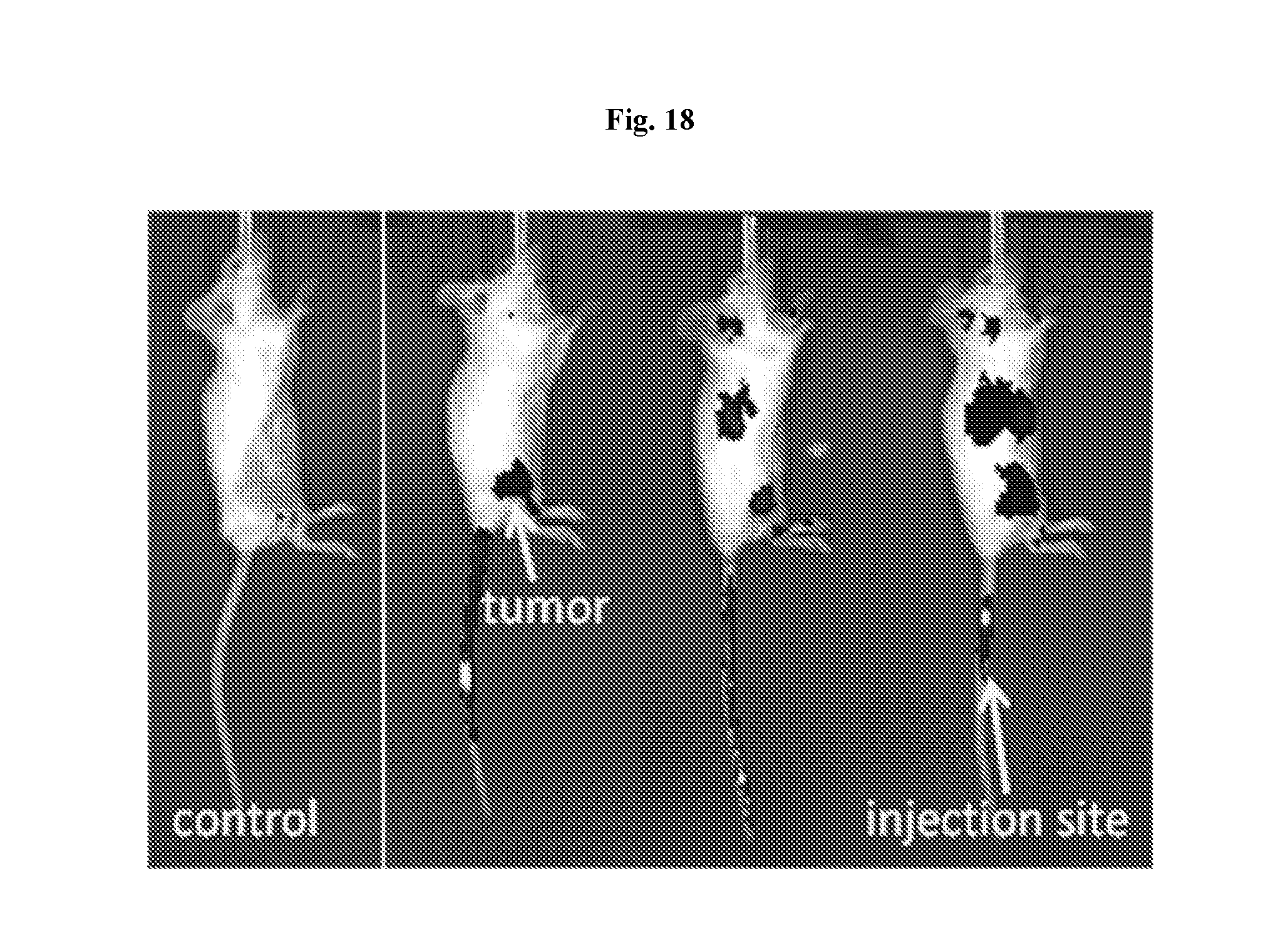

D00021

D00022
D00023
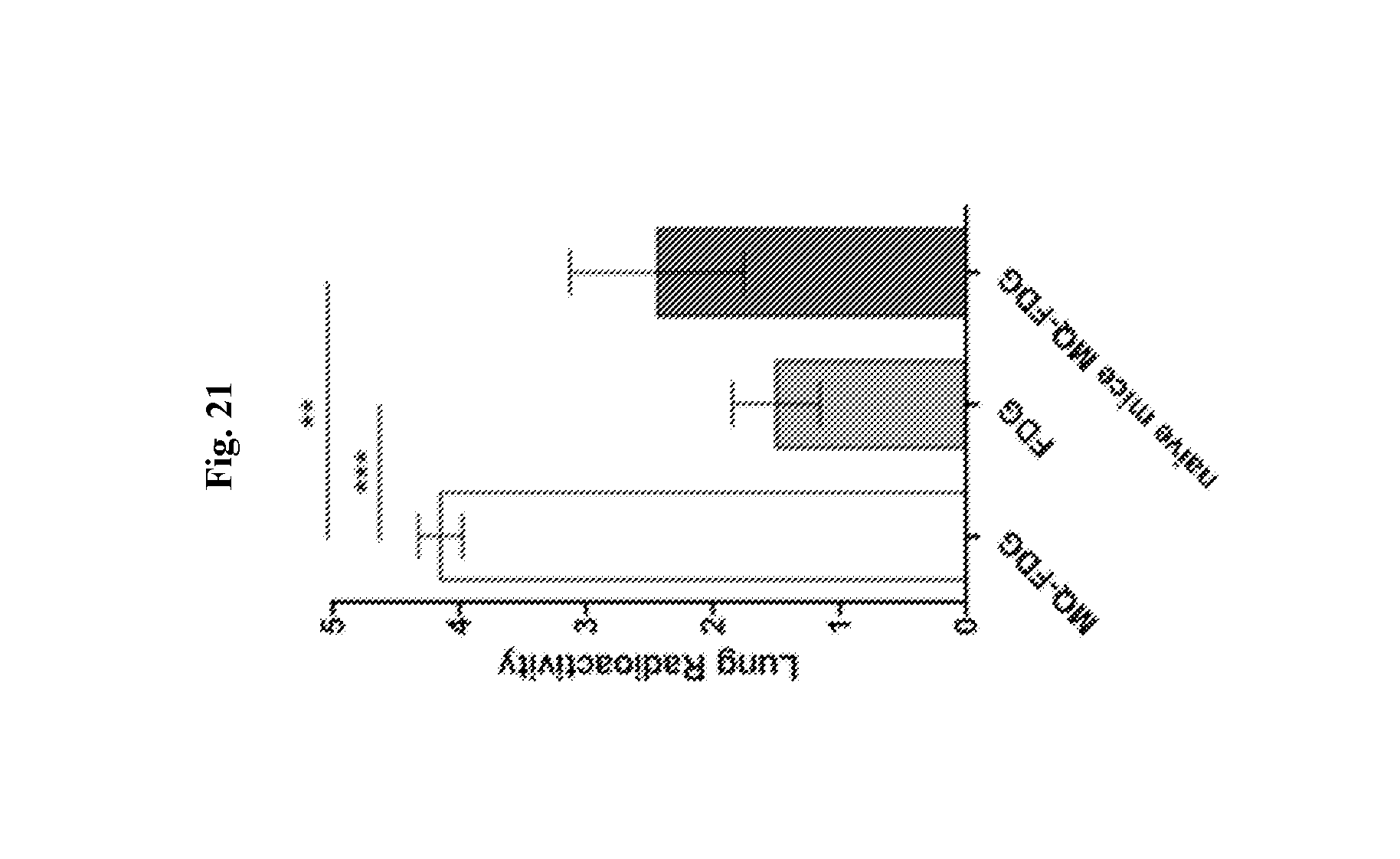
D00024
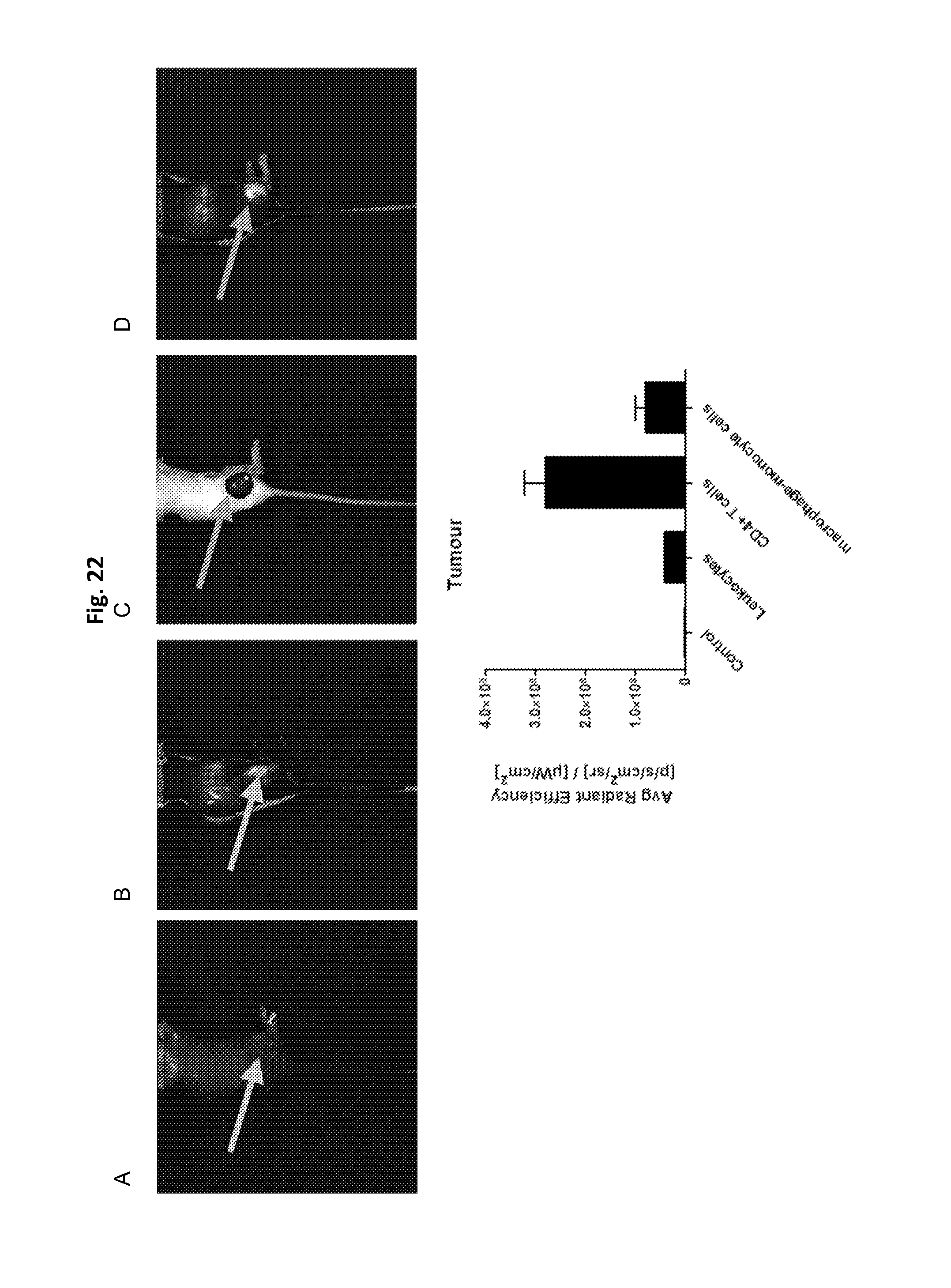

D00025
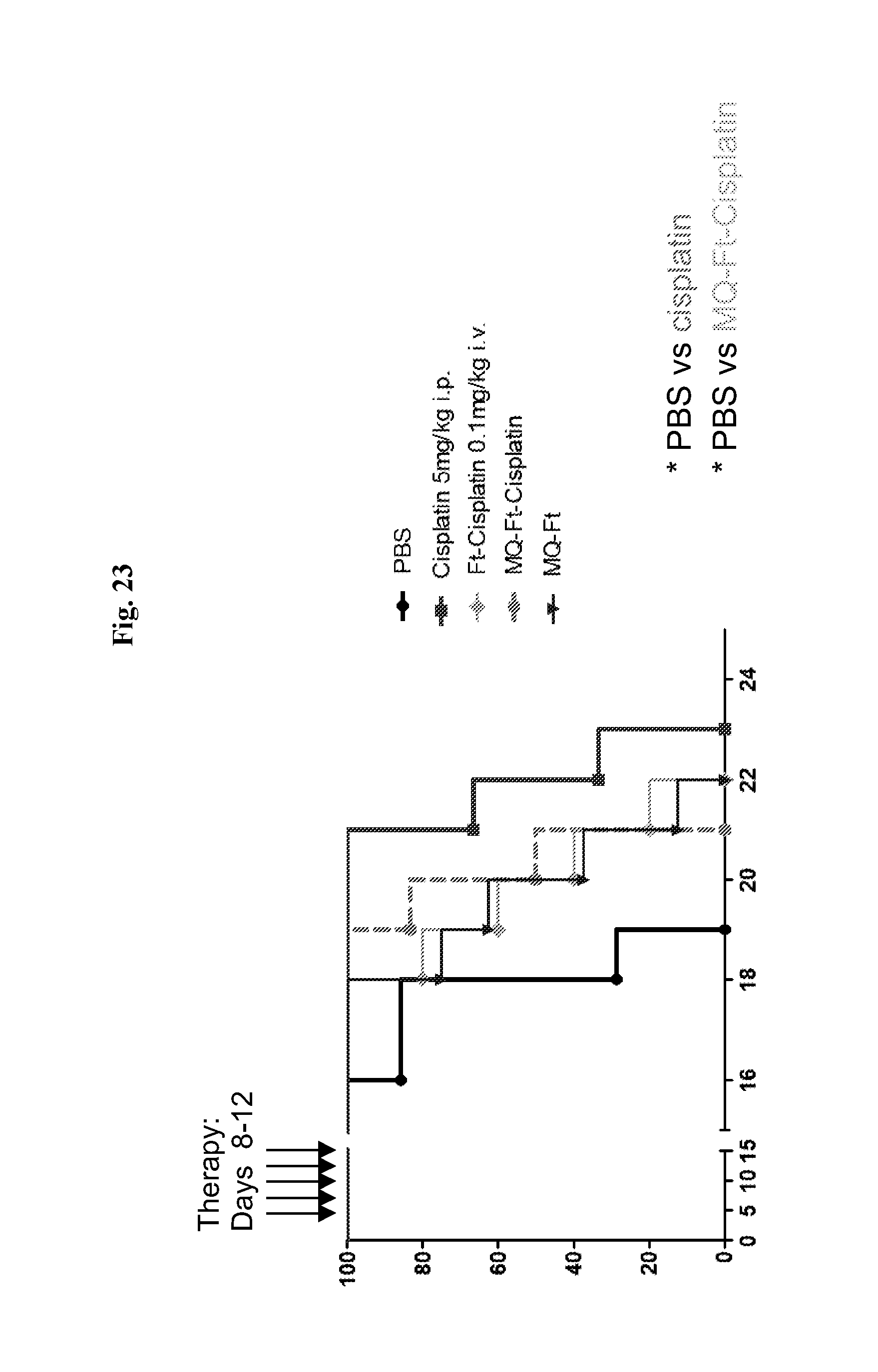
D00026
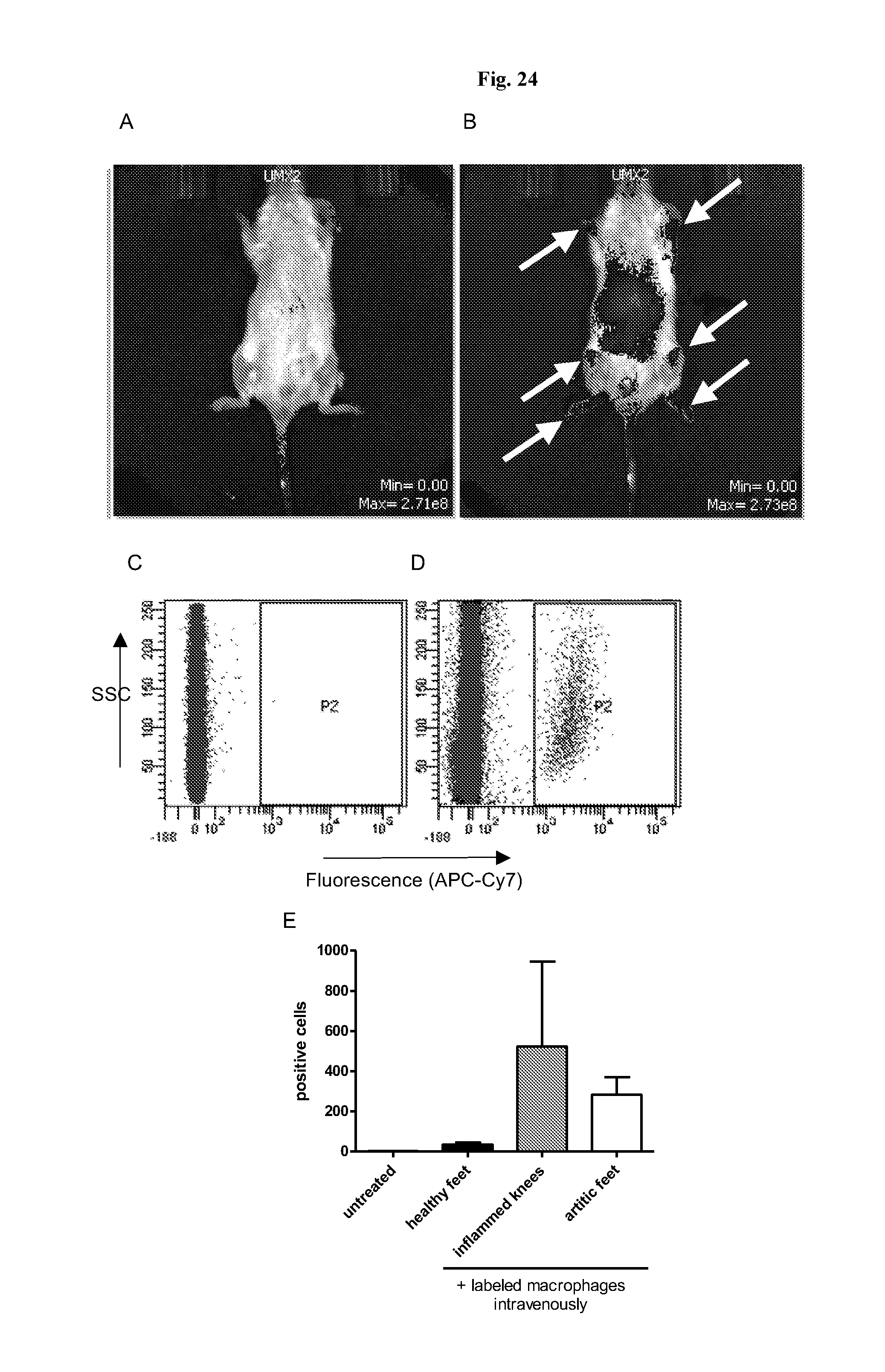
D00027
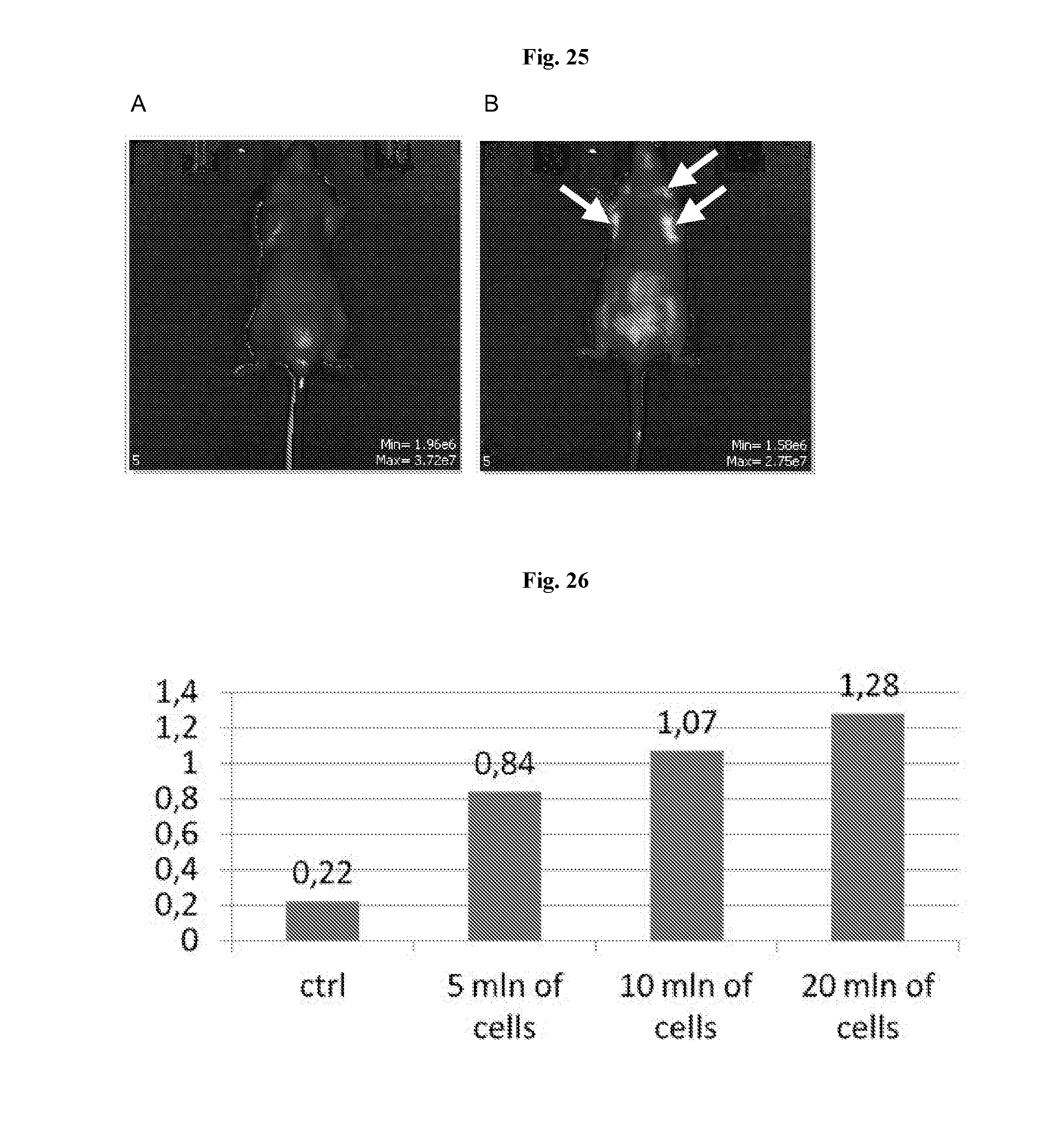
D00028

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.