Cancer Diagnosis, Treatment Selection And Treatment
Szallasi; Zoltan I. ; et al.
U.S. patent application number 16/404204 was filed with the patent office on 2019-10-10 for cancer diagnosis, treatment selection and treatment. This patent application is currently assigned to CHILDREN'S MEDICAL CENTER CORPORATION. The applicant listed for this patent is THE BRIGHAM AND WOMEN'S HOSPITAL, INC., CHILDREN'S MEDICAL CENTER CORPORATION, DANA-FARBER CANCER INSTITUTE, INC., THE TECHNICAL UNIVERSITY OF DENMARK. Invention is credited to Nicolai J. Birkbak, Andrea L. Richardson, Zoltan I. Szallasi, Zhigang Wang.
| Application Number | 20190309373 16/404204 |
| Document ID | / |
| Family ID | 51625329 |
| Filed Date | 2019-10-10 |









View All Diagrams
| United States Patent Application | 20190309373 |
| Kind Code | A1 |
| Szallasi; Zoltan I. ; et al. | October 10, 2019 |
CANCER DIAGNOSIS, TREATMENT SELECTION AND TREATMENT
Abstract
The present invention provides assays, methods and systems for selecting an effective therapy for a subset of cancer patients having cancer cells with increased expression of BML and FANCI genes and/or having copy number increase in chromosome location 15q26 in the cancer cells and for treatment of such patients with the effective therapy of cancer patients based on the personalized cancer cell expression profile.
| Inventors: | Szallasi; Zoltan I.; (Boston, MA) ; Richardson; Andrea L.; (Chestnut Hill, MA) ; Birkbak; Nicolai J.; (Kobenhavn O, DK) ; Wang; Zhigang; (Newton, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | CHILDREN'S MEDICAL CENTER
CORPORATION Boston MA THE BRIGHAM AND WOMEN'S HOSPITAL, INC. Boston MA DANA-FARBER CANCER INSTITUTE, INC. Boston MA THE TECHNICAL UNIVERSITY OF DENMARK Lyngby |
||||||||||
| Family ID: | 51625329 | ||||||||||
| Appl. No.: | 16/404204 | ||||||||||
| Filed: | May 6, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14774772 | Sep 11, 2015 | 10308986 | ||
| PCT/US2014/025774 | Mar 13, 2014 | |||
| 16404204 | ||||
| 61784234 | Mar 14, 2013 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12Q 1/6886 20130101; A61K 33/24 20130101; C12Q 2600/106 20130101; C12Q 2600/156 20130101; C12Q 2600/158 20130101; A61K 31/704 20130101; G01N 33/57415 20130101 |
| International Class: | C12Q 1/6886 20060101 C12Q001/6886; A61K 31/704 20060101 A61K031/704; G01N 33/574 20060101 G01N033/574; A61K 33/24 20060101 A61K033/24 |
Goverment Interests
STATEMENT REGARDING FEDERALLY-SPONSORED RESEARCH
[0002] This invention was made with government support under Grant No. 2P50 CA89393-06 awarded by the National Cancer Institute. The Government has certain rights in the invention.
Claims
1. A method for treating cancer in a human subject in need thereof comprising the step of administering an anthracycline-comprising cancer therapy to the human subject when the cancer in the human subject has been determined to have increased expression of BLM and FANCI compared to a reference value.
2. A method of treating cancer in a human subject, comprising administering an anthracycline-comprising cancer therapy to a human subject when a cancer cell taken from the human subject has been or is determined to carry a chromosome 15q26 copy number gain compared to a reference value.
3. The method of claim 2, further comprising the steps of detecting a chromosome 15q26 copy number in a sample comprising a cancer cell taken from the subject; comparing the chromosome 15q26 copy number to a reference value.
4. The method of claim 2, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer.
5. The method of claim 2, wherein the subject's cancer is known to not express a detectable quantity of ER, PgR, and HER2 receptor.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a Divisional of U.S. patent Ser. No. 14/774,772, filed Sep. 11, 2015, which is a 35 U.S.C. .sctn. 371 National Phase Entry Application of International Application No. PCT/US2014/025774, filed Mar. 13, 2014, which designates the U.S., and which claims benefit under 35 U.S.C. .sctn. 119(e) of U.S. Provisional Application No. 61/784,234, filed Mar. 14, 2013, the contents of each of which are incorporated herein in their entireties.
SEQUENCE LISTING
[0003] The instant application contains a Sequence Listing which has been submitted electronically in ASCII format and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Nov. 18, 2015, is named 701039-075882-US_SL.txt and is 512 bytes in size.
BACKGROUND
[0004] All publications herein are incorporated by reference to the same extent as if each individual publication or patent application was specifically and individually indicated to be incorporated by reference. The following description includes information that may be useful in understanding the present invention. It is not an admission that any of the information provided herein is prior art or relevant to the presently claimed invention, or that any publication specifically or implicitly referenced is prior art.
[0005] Breast cancer is recognized as a collection of malignancies that all arise in the breast but are remarkably heterogeneous. Gene expression array experiments have defined at least 4 major subtypes: 1) Luminal A; 2) Luminal B; 3) HER2+; and 4) Basal. There is a strong correlation between clinically defined triple negative breast cancer "TNBC", as defined by the absence of ER, PgR, and HER2 by standard immunohistochemical staining, and the molecularly defined basal subtype.
[0006] TN breast cancer accounts for only 10-15% of all incident breast cancer cases in the U.S., but results in a disproportionate number of breast cancer deaths. Women who are destined to develop metastatic TN disease typically experience a short disease free interval and have a higher degree of lung and brain involvement than patients with luminal breast cancers. Furthermore, TN breast cancer is overrepresented among patients who carry a deleterious BRCA1 germline alteration, and among women of African ancestry.
[0007] From a clinical perspective, despite substantial efforts to develop novel targeted agents, chemotherapy remains the mainstay of therapy for TN breast cancer, as trials evaluating a number of agents either in lieu of chemotherapy or in addition to chemotherapy have failed to produce any new agent that is capable of convincingly changing the natural history of the disease. In the adjuvant setting, polychemotherapy regimens have been demonstrated to improve both disease-free survival and overall survival. In the neoadjuvant setting, a favorable response to chemotherapy is associated with a low chance of relapse at 5 years. In contrast, women who have a significant amount of residual disease after a course of neoadjuvant chemotherapy have a particularly poor prognosis, with at least half experiencing a recurrence and death from TN breast cancer within 5 years. In the metastatic setting, although patients may respond to chemotherapy, the responses tend to be brief, and resistance tends to appear quickly.
[0008] Given the curative potential of chemotherapy in patients presenting with stage I-III TN breast cancer, and the initial responses (albeit often brief) seen with chemotherapy in the metastatic setting, the general direction of targeted therapy development for TN breast cancer has been to combine targeted agents with chemotherapy. Thus, even with a growing number of targeted therapies under investigation for TN breast cancer, chemotherapy is likely to be a significant component of the treatment of TN breast cancer for many years to come.
[0009] Across breast cancers as a whole, hormone receptor dependence and tumor proliferation appear to be associated with generic "chemosensitivity". Using the 70 gene signature as an example, the current classifiers typically assign a "high-risk" status to the vast majority of TN tumors, and are also unable to define whether there may be differential benefit with one class of chemotherapeutic agents over another. Thus, in both clinical practice and in conventionally designed clinical trials, the tendency is to layer new therapies directly atop existing standards, increasing the risk of both overtreatment (because some, but not all of the administered therapies are efficacious) or undertreatment (because the tumor is not sensitive to any of the specific chemotherapeutic agents contained in the treatment regimen).
[0010] Accordingly, there exists a need in the art to identify the best chemotherapy for each patient and to eliminate agents that are inert and result in toxicity without benefit.
SUMMARY OF THE INVENTION
[0011] We provide novel methods, assays, systems and kits for determining if a cancer patient is responsive to platinum-comprising therapy or anthracyclin therapy. These methods, assays, systems and kits provide a significant improvement to the "trial-and-error"-therapies used in cancer therapy. The methods allow one to personalize the treatment of a cancer patient based on the cancer cells' specific protein/gene expression profile. In other words, the methods apply the novel findings of a cancer cells responses to cancer treatment methods and allow selection of the most likely effective therapy without delay and thus significantly improve the patient's quality of life. Avoiding use of ineffective drugs will also provide a significant saving for the cost of treatment for the cancer patient, as well avoiding exposure to side effects of ineffective drugs.
[0012] The invention is based, at least in part, on the discovery that patients having increased expression of BLM and/or FANCI genes compared to a housekeeping gene or wild-type BRCA1 gene in their cancer cells are likely to respond to platinum-comprising or anthracyclin-comprising cancer therapy.
[0013] The following embodiments and aspects thereof are described and illustrated in conjunction with compositions and methods which are meant to be exemplary and illustrative, not limiting in scope.
[0014] We provide an assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; comparing the BLM and FANCI expression to a reference value; and selecting a platinum-comprising cancer therapy for the subject when the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI, or selecting a non-platinum-comprising cancer therapy for the subject when the BLM and FANCI expression is not increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is not effective in subjects whose cancer does not have increased BLM and FANCI expression compared to the reference value.
[0015] The assay may further comprise assaying the BRCA1 and/or BRCA2 status of the subject; and selecting the platinum-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI and who are negative for BRCA1 and/or BRCA2 mutations.
[0016] In some aspects of all the embodiments of the invention, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations.
[0017] In some aspects of all the embodiments of the invention, the assay further comprises assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and selecting the platinum-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor.
[0018] In some aspects of all the embodiments of the invention, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor.
[0019] In some aspects of all the embodiments of the invention, the assay further comprises administering the selected therapy to the subject.
[0020] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0021] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0022] In some aspects of all the embodiments of the invention, the reference value is based on at least BRCA1 gene expression in the cancer cell.
[0023] In some aspects of all the embodiments of the invention, the reference value is based on at least one housekeeping gene expression in the cancer cell.
[0024] We also provide a method for selecting platinum-comprising therapy for a subject having cancer, and optionally administering the platinum-comprising therapy, the method comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; detecting the BLM and FANCI expression in the sample compared to a reference value; and electing a platinum-comprising cancer therapy for the subject when the BLM and FANCI expression compared to a reference value is increased based on the recognition that platinum-comprising cancer therapy is effective in patients whose cancer has increased BLM and FANCI expression compared to the reference value.
[0025] In some aspects of all the embodiments of the invention, the method further comprises administering to the subject the platinum-comprising cancer therapy when the platinum-comprising cancer therapy is selected. One can further select a therapy other than platinum-comprising therapy when it is determined using the assay that the cancer is not likely responsive to platinum-comprising therapy.
[0026] In some aspects of all the embodiments of the invention, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations.
[0027] In some aspects of all the embodiments of the invention, the subject's cancer or cancer cell is known to not or determined to not express a detectable quantity of ER, PgR, and HER2 receptor.
[0028] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0029] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0030] In some aspects of all the embodiments of the invention, the reference value is based on at least BRCA1 gene expression in the cancer cell.
[0031] In some aspects of all the embodiments of the invention, the reference value is based on at least one housekeeping gene expression in the cancer cell.
[0032] In some aspects of all the embodiments of the invention, the housekeeping gene is selected from beta-actin, GAPDH, RPLP0, GUS, TFRC and any combination thereof.
[0033] In some aspects of all the embodiments of the invention, the housekeeping gene is RPLP0, and the BLM and/or FANCI expression is increased by at least six-fold.
[0034] We further provide a method for selecting a non-platinum-comprising therapy, and optionally administering the non-platinum-comprising therapy, for a subject having cancer comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; detecting the BLM and FANCI expression in the sample compared to a reference value; and selecting the non-platinum-comprising cancer therapy for the subject when the BLM and FANCI expression compared to the reference value is not increased based on the recognition that non-platinum-comprising cancer therapy is effective in patients whose cancer does not have increased gene expression of BLM and FANCI compared to the reference value.
[0035] In some aspects of all the embodiments of the invention, the method further comprises administering to the subject the non-platinum-comprising cancer therapy when non-platinum-comprising cancer therapy is selected.
[0036] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0037] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0038] In some aspects of all the embodiments of the invention, the reference value is based on at least BRCA1 gene expression in the cancer cell.
[0039] In some aspects of all the embodiments of the invention, the reference value is based on at least one housekeeping gene expression in the cancer cell.
[0040] We provide an assay for selecting a therapy for a subject having cancer, comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; comparing the BLM and FANCI expression to a reference value; and selecting an anthracycline-comprising cancer therapy for the subject when the BLM and FANCI expression is increased compared to a reference value based on the recognition that anthracycline-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI, or selecting a non-anthracycline-comprising cancer therapy for the subject when the BLM and FANCI expression is not increased compared to a reference value based on the recognition that anthracycline-comprising cancer therapy is not effective in subjects whose cancer does not have increased BLM expression compared to a reference value.
[0041] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0042] We provide a method for selecting an anthracycline-comprising cancer therapy for a subject having cancer and determined to be negative for BRCA1 and/or BRCA2 mutations, comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; comparing the BLM and FANCI expression to a reference value; and selecting the anthracycline-compri sing cancer therapy for the subject when the BLM and FANCI expression compared to the reference value is increased based on the recognition that anthracycline-comprising cancer therapy is effective in patients whose cancer has increased expression of BLM and FANCI compared to the reference value.
[0043] In some aspects of all the embodiments of the invention, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0044] We further provide a method of treating cancer in a human subject, comprising: detecting BLM and FANCI expression in a sample comprising a cancer cell taken from the human subject; and comparing the BLM and FANCI expression to a reference value; and administering a platinum-comprising cancer therapy to the human subject wherein an increase of BLM and FANCI expression compared to the reference value is detected.
[0045] In some aspects of all the embodiments of the invention, the human subject's cancer or cancer cell is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor.
[0046] In some aspects of all the embodiments of the invention, the cancer is selected from breast, ovarian, and lung cancers.
[0047] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0048] In some aspects of all the embodiments of the invention, the reference value is based on BRCA1 gene expression in the cancer cell.
[0049] In some aspects of all the embodiments of the invention, the reference value is based on a housekeeping gene expression in the cancer cell.
[0050] We provide method of treating cancer in a human subject, comprising: detecting BLM and FANCI expression in a sample comprising a cancer cell taken from the human subject; and comparing the BLM and FANCI expression to a reference value; and administering an anthracycline-comprising cancer therapy to the human subject wherein an increase of BLM and FANCI expression compared to the reference value is detected.
[0051] In some aspects of all the embodiments of the invention, the human subject's cancer or cancer cell is known to not or is determined to not to express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor.
[0052] In some aspects of all the embodiments of the invention, the cancer is selected from breast, ovarian, and lung cancers.
[0053] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0054] In some aspects of all the embodiments of the invention, the reference value is based on BRCA1 gene expression in the cancer cell.
[0055] In some aspects of all the embodiments of the invention, the reference value is based on a housekeeping gene expression in the cancer cell.
[0056] We provide a method for assessing responsiveness of a cancer cell to cancer therapy, comprising: assaying, in a cancer cell or mRNA derived therefrom, BLM and FANCI expression; and comparing said BLM and FANCI expression to a reference value, wherein the cancer cell is assessed as responsive to a platinum-comprising therapy if the BLM and FANCI expression is increased compared to the reference value, or wherein the cancer cell is assessed as poorly or not responsive to platinum-comprising cancer therapy cancer if the BLM and FANCI expression is not increased.
[0057] In some aspects of all the embodiments of the invention, the step of assaying comprises: contacting the cancer cell or mRNA derived therefrom with at least one detectably labeled probe capable of specifically binding to BLM mRNA, at least one detectably probe capable of specifically binding to FANCI, at least one detectably labeled probe capable of specifically binding to BRCA1 and/or at least one housekeeping gene; and measuring the expression of BLM and FANCI compared to the BRCA1 and/or the at least one housekeeping gene.
[0058] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0059] We provide a method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: determining, in a cancer cell from the cancer patient, BLM and FANCI expression; and correlating the expression to a reference value, wherein when the expression is increased the patient is predicted to respond well to a cancer treatment regimen comprising platinum or anthracycline.
[0060] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0061] We also provide a method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: determining, in a cancer cell or mRNA derived therefrom from said cancer patient, BLM and FANCI expression; and correlating the expression to a reference value, wherein when the expression is not increased the patient is predicted to respond poorly to a cancer treatment regimen comprising platinum or anthracycline.
[0062] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0063] We provide a method of treating cancer, comprising: assaying, in a cancer cell from a cancer patient or mRNA obtained therefrom, the BLM and FANCI expression compared to a reference value; and administering to the cancer patient a cancer treatment regimen comprising platinum or anthracycline if the BLM and FANCI expression is increased compared to the reference value.
[0064] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0065] We also provide a use of platinum comprising cancer therapy for treating a cancer patient that has been determined to have a tumor comprising cancer cells wherein BLM and FANCI expression is increased compared to a reference value.
[0066] In some aspects of all the embodiments of the invention, the cancer patient has been determined to be negative for BRCA1 and/or BRCA2 mutations.
[0067] In some aspects of all the embodiments of the invention, the cancer patient's cancer or cancer cell is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor.
[0068] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0069] We provide a system for determining responsiveness of a cancer cell to platinum-comprising therapy from a cancer cell of a cancer patient, comprising: a sample analyzer configured to produce a signal for the mRNA from each one of BLM and FANCI from a cancer cell sample of a cancer patient; and a computer sub-system programmed to calculate, based on the mRNA whether the signal is greater or not than a reference value.
[0070] In some aspects of all the embodiments of the invention, said computer sub-system is programmed to compare the mRNA to determine a likelihood of responsiveness of said cancer cell to platinum-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a platinum-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the platinum-comprising therapy if the BLM and FANCI expression is not increased; or a likelihood of responsiveness of said cancer cell to anthracycline-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a anthracycline-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the anthracycline-comprising therapy if the BLM and FANCI expression is not increased.
[0071] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0072] We provide a computer program product embodied in a computer readable medium that, when executing on a computer, performs steps comprising: detecting the BLM and FANCI gene expression in sample comprising a cancer cell from a cancer patient; and comparing the BLM and FANCI expression to a reference value.
[0073] In some aspects of all the embodiments of the invention, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0074] We provide a diagnostic kit for detecting a likelihood of a cancer patient to respond to platinum- or anthracycline-comprising comprising cancer therapy, comprising: no more than 10 probes comprising a combination of detectably labeled probes or primers for BLM and FANCI, and optionally for BRCA1 and/or at least one housekeeping gene; and the computer program product of Claim 59.
[0075] We provide use of a plurality of oligonucleotides comprising no more than 10 oligonucleotides capable of hybridizing to BLM and FANCI, and optionally to BRCA1 and/or at least one housekeeping gene, in a diagnostic kit for determining an increased likelihood that a cancer patient will respond to cancer treatment regimen comprising a platinum and/or anthracycline.
[0076] In some aspects of all the embodiments of the invention, said anthracycline is epirubincin or doxorubicin.
[0077] In some aspects of all the embodiments of the invention, said platinum comprising cancer therapy comprises cisplatinum or cis-diamminedichloroplatinum, phenanthriplatin, carboplatin, oxaliplatin, or a platinum complex that is activated by ultraviolet A light.
[0078] We provide an assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; comparing the chromosome 15q26 copy number to a reference value; and selecting a platinum-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or selecting a non-platinum-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0079] In some aspects of all the embodiments of the invention, the assay further comprises: assaying the BRCA1 and/or BRCA2 status of the subject; and selecting the platinum-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and there is a chromosome 15q26 copy number gain based on the recognition that platinum-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain who are negative for BRCA1 and/or BRCA2 mutations.
[0080] In some aspects of all the embodiments of the invention, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations.
[0081] In some aspects of all the embodiments of the invention, the assay further comprises: assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and selecting the platinum-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when there is a chromosome 15q26 copy number gain based on the recognition that platinum-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor.
[0082] In some aspects of all the embodiments of the invention, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor.
[0083] In some aspects of all the embodiments of the invention, the assay further comprises administering the selected therapy to the subject.
[0084] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0085] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0086] We provide an assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; comparing the chromosome 15q26 copy number to a reference value; and selecting an anthracycline-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or selecting a non-anthracycline-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0087] In some aspects of all the embodiments of the invention, the assay further comprises: assaying the BRCA1 and/or BRCA2 status of the subject; and selecting the anthracycline-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and there is a chromosome 15q26 copy number gain based on the recognition that anthracycline-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain who are negative for BRCA1 and/or BRCA2 mutations.
[0088] In some aspects of all the embodiments of the invention, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations.
[0089] In some aspects of all the embodiments of the invention, the assay further comprises assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and selecting the anthracycline-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when there is a chromosome 15q26 copy number gain based on the recognition that anthracycline-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor.
[0090] In some aspects of all the embodiments of the invention, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor.
[0091] In some aspects of all the embodiments of the invention, the assay further comprises administering the selected therapy to the subject.
[0092] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0093] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0094] We provide a method of treating cancer in a human subject, comprising: detecting a chromosome 15q26 copy number in a sample comprising a cancer cell taken from the subject; comparing the chromosome 15q26 copy number to a reference value; and administering an platinum-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or administering a non-platinum-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0095] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0096] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0097] In some aspects of all the embodiments of the invention, the subject's cancer is known to not express a detectable quantity of ER, PgR, and HER2 receptor.
[0098] We provide a method of treating cancer in a human subject, comprising: detecting a chromosome 15q26 copy number in a sample comprising a cancer cell taken from the subject; comparing the chromosome 15q26 copy number to a reference value; and administering an anthracycline-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or administering a non-anthracycline-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0099] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0100] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0101] In some aspects of all the embodiments of the invention, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor.
[0102] We provide a method for assessing responsiveness of a cancer cell to a cancer therapy, and optionally administering the cancer therapy, comprising: assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; and comparing the chromosome 15q26 copy number to a reference value, wherein the cancer cell is assessed as responsive to a platinum-comprising therapy if there is a chromosome 15q26 copy number gain compared to the reference value, or wherein the cancer cell is assessed as poorly or not responsive to platinum-comprising cancer therapy cancer if there is not a chromosome 15q26 copy number gain or if there is a chromosome 15q26 copy number loss.
[0103] In some aspects of all the embodiments of the invention, the reference value is copy number of chromosome 15.
[0104] In some aspects of all the embodiments of the invention, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0105] In some aspects of all the embodiments of the invention, the method further comprises administering the platinum-comprising therapy if there is a chromosome 15q26 copy number gain.
[0106] We provide a method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: determining, in a cancer cell from the cancer patient, chromosome 15q26 copy number; and correlating the chromosome 15q26 copy number to a reference value, wherein when there is a chromosome 15q26 copy number gain, the patient is predicted to respond well to a cancer treatment comprising platinum or anthracycline, or wherein when there is not a chromosome 15q26 copy number gain or a chromosome 15q26 copy number loss, the patient is predicted respond poorly to a cancer treatment comprising platinum or anthracycline.
[0107] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0108] We provide use of platinum comprising cancer therapy for treating a cancer patient that has been determined to have a tumor comprising cancer cells wherein a chromosome 15q26 copy gain is detected compared to a reference value.
[0109] In some aspects of all the embodiments of the invention, the cancer patient is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations.
[0110] In some aspects of all the embodiments of the invention, the cancer patient's cancer or cancer cell is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor.
[0111] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0112] We provide a system for determining responsiveness of a cancer cell to platinum-comprising therapy from a cancer cell of a cancer patient, comprising: a sample analyzer configured to produce a signal for chromosome 15q26 copy number from a cancer cell sample of a cancer patient; and a computer sub-system programmed to calculate, based on the mRNA whether the signal is greater or not than a reference value.
[0113] In some aspects of all the embodiments of the invention, said computer sub-system is programmed to compare the mRNA to determine a likelihood of responsiveness of said cancer cell to platinum-comprising cancer therapy and/or or a anthracycline-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a platinum-comprising therapy if there is a chromosome 15q26 copy number gain and as unlikely to respond to the platinum-comprising therapy if there is not a chromosome 15q26 copy number gain or if there is a chromosome 15q26 copy number loss.
[0114] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0115] We provide a computer program product embodied in a computer readable medium that, when executing on a computer, performs steps comprising: detecting chromosome 15q26 copy number in sample comprising a cancer cell from a cancer patient; and comparing the chromosome 15q26 copy number to a reference value.
[0116] In some aspects of all the embodiments of the invention, the reference value is chromosome 15 centromere copy number in the sample.
[0117] We provide a diagnostic kit for detecting a likelihood of a cancer patient to respond to platinum- or anthracycline-comprising cancer therapy, comprising: no more than 10 probes comprising a combination of detectably labeled probes or primers for chromosome 15q26, and optionally for chromosome 15 centromere; and a computer program as described herein.
[0118] We provide us of a plurality of oligonucleotides comprising no more than 10 oligonucleotides capable of hybridizing to chromosome 15q26, and optionally for chromosome 15 centromere, in a diagnostic kit for determining an increased likelihood that a cancer patient will respond to cancer treatment regimen comprising a platinum and/or anthracycline.
[0119] In some aspects of all the embodiments of the invention, said anthracycline is epirubincin or doxorubicin.
[0120] In some aspects of all the embodiments of the invention, said platinum comprising cancer therapy comprises cisplatinum or cis-diamminedichloroplatinum, phenanthriplatin, carboplatin, oxaliplatin, or a platinum complex that is activated by ultraviolet A light.
[0121] Other features and advantages of the invention will become apparent from the following detailed description, taken in conjunction with the accompanying drawings, which illustrate, by way of example, various features of embodiments of the invention.
BRIEF DESCRIPTION OF THE FIGURES
[0122] This patent or application file contains at least one drawing executed in color. Copies of this patent or patent application publication with color drawings will be provided by the Office upon request and payment of the necessary fee.
[0123] Exemplary embodiments are illustrated in referenced figures. It is intended that the embodiments and figures disclosed herein are to be considered illustrative rather than restrictive.
[0124] FIGS. 1A and 1B show that GISTIC identifies 169 chromosomal regions as either gained or lost. Several contained genes with significantly different copy number between sensitive and resistant cases. FIG. 1A shows data relating to the Cisplatin-1 clinical trial. FIG. 1B shows data relating to the Cisplatin-2 clinical trial.
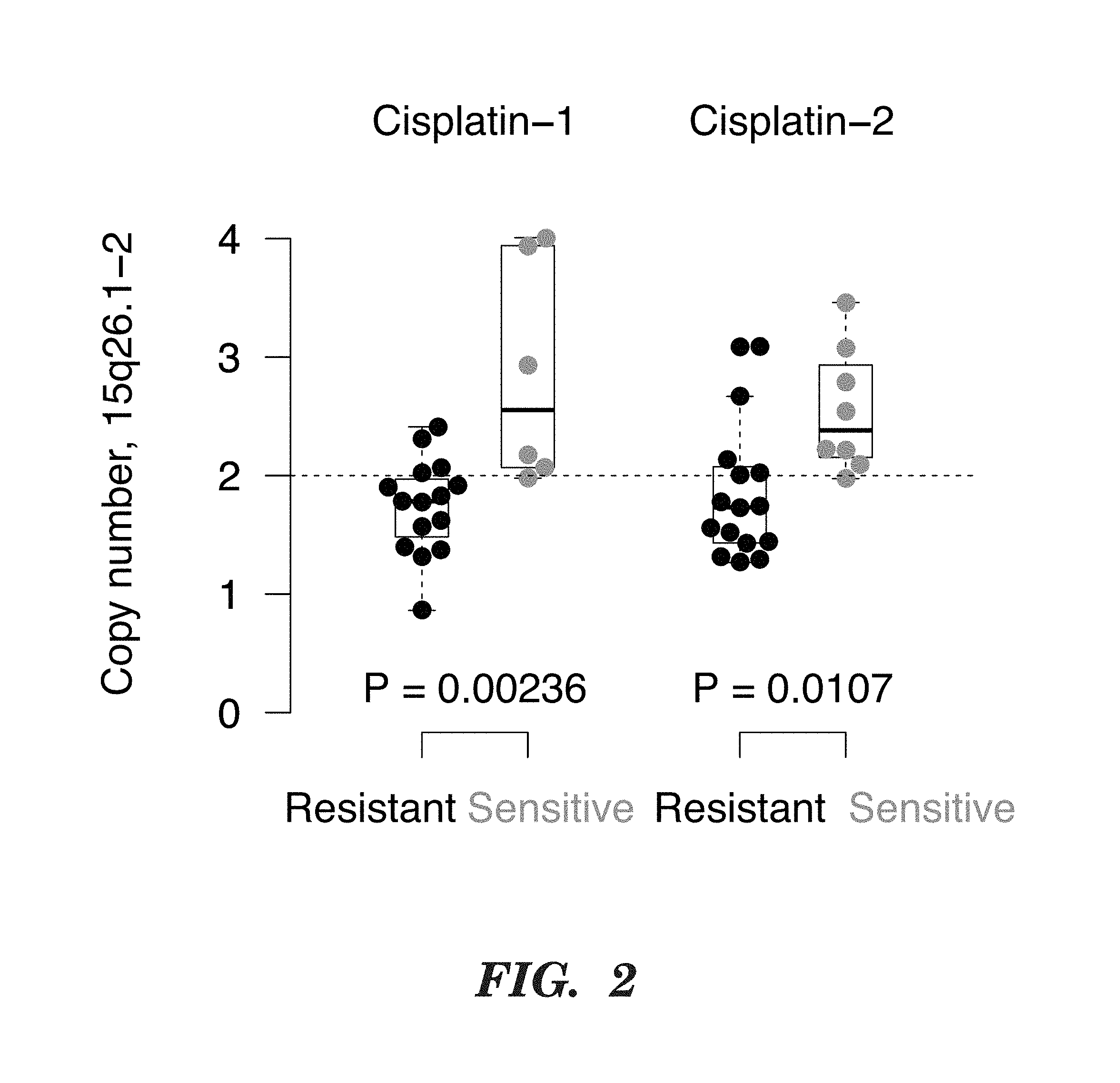
[0125] FIG. 2 shows a 10 MB region with 59 genes on chr. 15q26.1-2 showed significant gain in sensitive samples.
[0126] FIG. 3 shows correlation between copy number and mRNA expression, Genes on 15q26.1-2. 23 genes show significant correlation.
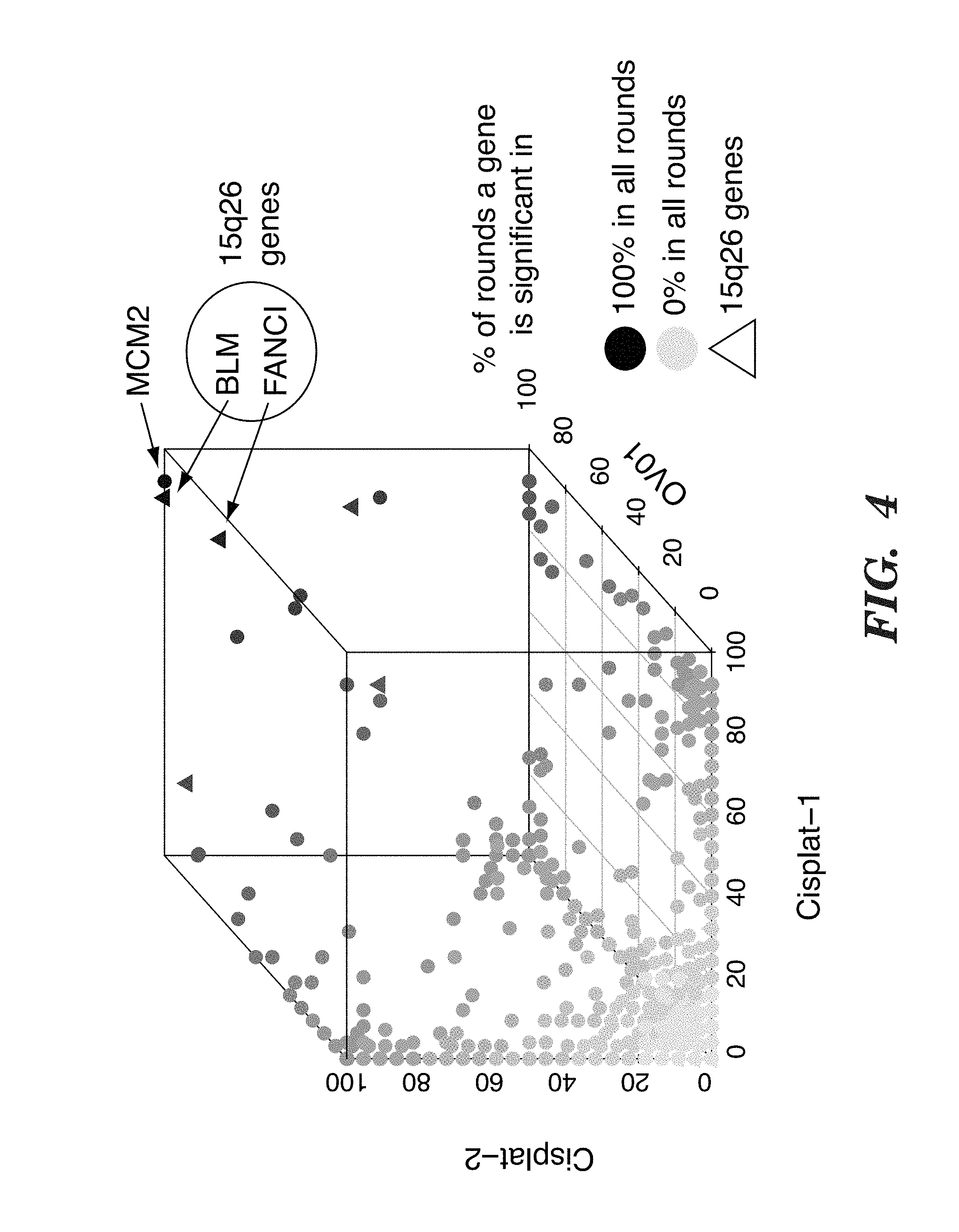
[0127] FIG. 4 depicts leave-one-out analysis of gene expression data.
[0128] FIGS. 5A-5B shows that expression of BLM and FANCI is only associated with response to carboplatin, but not paclitaxel in OVO1. (FIG. 5A) BLM expression; (FIG. 5B) FANCI Expression. Ahmed et al., Cancer Cell 2007.
[0129] FIGS. 6A-6B depicts expression of BLM or FANCI is not associated with response to multi-agent chemotherapy regimens in triple negative breast cancer (TNBC). (FIG. 6A) BLM expression; (FIG. 6B) FANCI Expression. .sup.1Hess et al., JCO 2006; Popovici et al.; .sup.2Breast Cancer Res 2010; .sup.3Andre et al., Clin Cancer Res 2009.
[0130] FIG. 7 shows that BLM is also associated with response to single agent epirubicin. TOP trial, single agent epirubicin in ER-negative breast cancer. .sup.1Desmedt et al., JCO 2011.
[0131] FIG. 8 depicts repair of cisplatin induced damage.
[0132] FIG. 9 depicts the metaphase spread, BRCA1 knock-out in MEFs. BRCA1 is an essential part of the homologous repair system. BRCA1 knock out cause high levels of quadrahelical chromosomes, which is a prime substrate for the Bloom helicase. Gain of BLM is a possible compensation mechanism for deficient HR. Silver D P et al, Cell 2007.
[0133] FIGS. 10A-10B show gene signatures. FIG. 10A shows receiver operating characteristic curves showing the ability of the BRCA1-BLM-FANCI RNA 3-gene signature to predict for response to cisplatin in the two cisplatin trials. FIG. 10B shows ROC curves showing the ability of number of telomeric allelic imbalance (NtAI)+3-gene mRNA signature to predict for sensitivity to cisplatin in the combined trials.
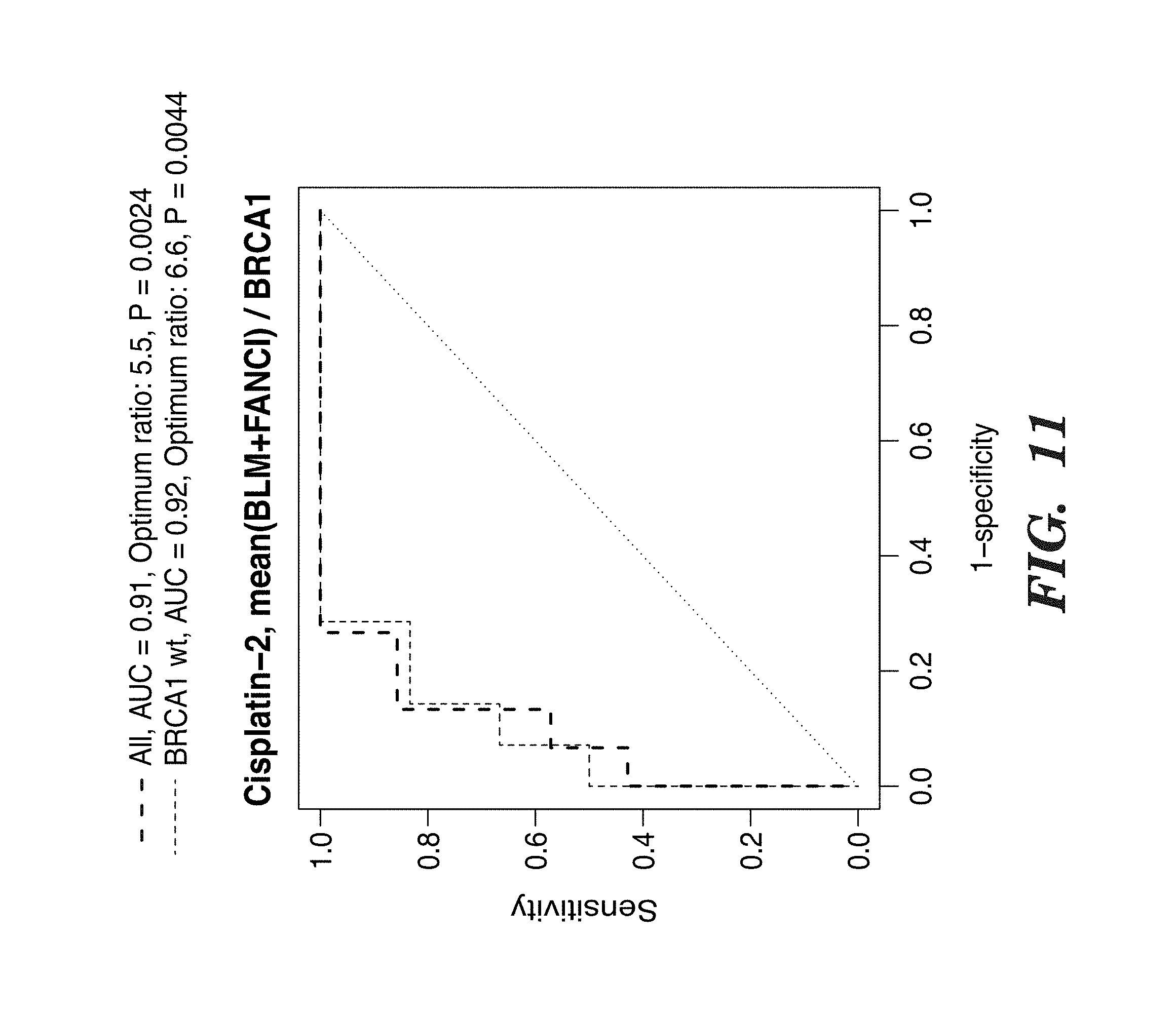
[0134] FIG. 11 depicts ROC analysis of 3 gene signature to determine cisplatin sensitivity in cisplatin 2 trial. Blue: all cases; red; BRCA normal cases only.
[0135] FIG. 12 depicts functional modules of various embodiments of the invention.
[0136] FIG. 13 depicts survival time with Kaplan-Meier plot, showing over-all survival of patients from the TCGA squamous cell lung cancer cohort, split by the median of the three gene signature calculated as the ratio of BRCA1 expression over the mean of FANI and BLM expression.
[0137] FIG. 14 shows the BLM+FANCI/BRCA1 signature. The Cisplatin-2 trial, separating resistant (MP 1-2-3) from sensitive (MP 4-5) cases is optimized for qPCR data. Blue line is based on all samples, while red is based on wtBRCA1. The optimum ratio is calculated by ROC curve, and shows an optimum ratio of 5.5 for all cases, on a log 2 scale. In un-logged values, this is equivalent of 45 fold.
[0138] FIG. 15 shows BLM/BRCA1 signature. The Cisplatin-2 trial, separating resistant (MP 1-2-3) from sensitive (MP 4-5) cases is optimized for qPCR data. The optimum ratio is 7.5 for all cases, on a log 2 scale, which is equivalent of 180 fold or 7.5 cycles on a PCR machine. This is a very large difference.
[0139] FIG. 16 shows FANCI/BRCA1 signature. The Cisplatin-2 trial, separating resistant (MP 1-2-3) from sensitive (MP 4-5) cases is optimized for qPCR data. The optimum ratio is 5.1 for all cases, on a log 2 scale, which is equivalent of 34 fold.
[0140] FIG. 17 shows the BRCA1/RPLP0 signature. BRCA1 is compared to a housekeeping gene, RPLP0. The optimum ratio here is -14 for all cases, on a log 2 scale, which is approximately equivalent to 1/1600, or 14 cycles on a PCR machine. This is a very large difference, but HK genes are expressed in very high numbers.
[0141] FIG. 18 shows BLM/RPLP0 signature. BLM is compared to a housekeeping gene, RPLP0 The optimum ratio is higher, and separation of sensitive/resistant cases is good.
[0142] FIG. 19 shows FANCI/RPLP0 signature. FANCI is compared to a housekeeping gene, RPLP0. The optimum ratio is higher than BRCA1 alone.
[0143] FIGS. 20A and 20B show how the expression of each of the three genes looks like in a large cohort of breast cancer patients from the TCGA, and how their ratios is to RPLP0 and BRCA1. These patients were not treated with platinum. This was focused on DNBC cases (ER/HER2 negative), and the normal breast samples. The ratio is much lower than using PCR data, yet there still appear to be a clear increase in some cases. The normal samples have lower BLM and BRCA1 expression, but that the ratio between these is inversed, with higher BRCA1 than BLM, probably reflecting the normal balance between these genes. FIG. 20A shows the expression of ESR1, BLM, ERBB2, FANCI, BRCA1, and RPLP0 in DNBC, estrogen receptor negative (ER), and HER2 receptor negative breast cancer patients and in patients with metastases compared to normal breast samples. FIG. 20B shows data relating to (i) the 3-gene signature, (ii) BRCA1/RPLP0 ratio, (iii) BLM/BRCA1 ratios, (iv) BLMIRPLP0 ratios, (v) FANCI/BRCA1 ratios, and (vi) FANCI/RPLP0 ratios in the same cohorts as FIG. 20A.
DETAILED DESCRIPTION
[0144] All references cited herein are incorporated by reference in their entirety as though fully set forth. Unless defined otherwise, technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Singleton et al., Dictionary of Microbiology and Molecular Biology 3.sup.rd ed., J. Wiley & Sons (New York, N.Y. 2001); March, Advanced Organic Chemistry Reactions, Mechanisms and Structure 5.sup.th ed., J. Wiley & Sons (New York, N.Y. 2001); and Sambrook and Russel, Molecular Cloning: A Laboratory Manual 3rd ed., Cold Spring Harbor Laboratory Press (Cold Spring Harbor, N.Y. 2001), provide one skilled in the art with a general guide to many of the terms used in the present application. For references on how to prepare antibodies, see D. Lane, Antibodies: A Laboratory Manual (Cold Spring Harbor Press, Cold Spring Harbor N.Y., 1988); Kohler and Milstein, (1976) Eur. J. Immunol. 6: 511; Queen et al. U.S. Pat. No. 5,585,089; and Riechmann et al., Nature 332: 323 (1988).
[0145] One skilled in the art will recognize many methods and materials similar or equivalent to those described herein, which could be used in the practice of the present invention. Indeed, the present invention is in no way limited to the methods and materials described.
[0146] To investigate whether specific genomic aberrations may affect cancer sensitivity to cisplatin, the inventors generated tumor DNA copy number profiles of 21 and 24 TNBC patients who received pre-operative cisplatin-based chemotherapy in two separate clinical trials. Using the GISTIC algorithm (1), the inventors found that only a single region on chromosome 15q26 showed consistent significant differential copy number in responders versus non-responders, being preferentially lost in non-responders, but preferentially gained in responders in both trials.
[0147] To see if genes on 15q26 were associated with platinum sensitivity, the inventors acquired gene expression data from the cisplatin TNBC trial (2), and from the carboplatin-only arm of an ovarian cancer trial (3). The inventors then performed a leave-one-out analysis, and found 9 genes significantly associated with platinum response in at least 75% of all rounds in both cohorts. These included BLM and FANCI located in the 15q26 region, both showing higher expression in sensitive tumors, and known to be involved in related DNA repair processes. To investigate if BLM and FANCI were specifically associated with genotoxic chemotherapy sensitivity, the inventors analyzed their expression in TNBCs from three neoadjuvant trials of epirubicin alone (4) or taxane-containing combination therapy (5, 6) and in ovarian cancers from the taxane-only treatment arm (3). In the epirubicin trial, BLM and FANCI expression was again significantly associated with increased sensitivity to therapy. In contrast, there was no association between either BLM or FANCI expression and TNBC response to the taxane-containing regimen or ovarian cancer response to single agent taxane treatment. These data suggest that high expression of BLM and FANCI are associated with improved response to DNA damaging agents, but not with response to other types of chemotherapeutics. Furthermore, it suggests that the patient subpopulations that respond to drugs such as anthracyclines and taxanes are not overlapping, and that it will therefore be difficult to robustly identify predictors of single agent response based on multi-drug trials.
[0148] While not wishing to be bound by any particular theory, the inventors believe that FANCI and BLM functions in multiple DNA repair processes; increased FANCI and BLM is associated with response to platinum-comprising therapy and anthracycline-comprising therapy; low copy gain might be a compensatory mechanism of HR deficient cells, trying to rescue some DNA repair capacity; and if so, FANCI/BLM expression is a marker for DNA repair deficiency, and increased sensitivity to genotoxic chemotherapy drugs, such as platinum-comprising therapy and anthracycline-comprising therapy.
[0149] Therefore, while the cancers specifically investigated for their responses in the particular studies, such as breast cancer, such as triple negative breast cancer, ovarian cancer and lung cancer, Applicants believe that the finding of the association between increased FANCI and BLM expression and responsiveness to platinum-comprising therapy and anthracycline-comprising therapy is applicable to most cancers.
[0150] The present invention is based, at least in part, on these findings, and those further described herein and in the figures and examples.
Selecting Therapy
[0151] Various embodiments provide for assays, methods and systems for selecting an appropriate therapy for a subject based on an analysis of the subject's BLM and FANCI expression, or based on the subject's 15q26 copy number.
[0152] In various embodiments, the invention provide for an assay for selecting a therapy, and optionally administering the therapy, for a subject having cancer, the assay comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; comparing the BLM and FANCI expression to a reference value; and selecting a platinum-comprising cancer therapy for the subject when the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI, or selecting a non-platinum-comprising cancer therapy for the subject when the BLM and FANCI expression is not increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is not effective in subjects whose cancer does not have increased BLM and FANCI expression compared to the reference value.
[0153] In various embodiments, the assay further comprises: assaying the BRCA1 and/or BRCA2 status of the subject; and selecting the platinum-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI and who are negative for BRCA1 and/or BRCA2 mutations. In various embodiments, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. The determination can be made before, concurrently with, or after the analysis for BLM and FANCI expression.
[0154] In some aspects of all the embodiments of the invention, a mutation that inactivates BRCA2 is highly predictive of response.
[0155] In various embodiments, the assay further comprises: assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and selecting the platinum-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor. In various embodiments, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. The determination can be made before, concurrently with, or after analysis for BLM and FANCI expression.
[0156] In various embodiments, the assay further comprises administering the selected therapy to the subject.
[0157] In various embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0158] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0159] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold, 2.8-fold, 2.9-fold, 3-fold, 4-fold, 5-fold, or 6-fod, 10-20 fold, 20-50 fold or higher, depending on the expression level of the gene used as a standard, such as one or more housekeeping genes or BRCA1.
[0160] Typically, an increase in expression of at least 1.5 or at least 2 fold is considered as a cut-off point for increased expression if BRCA1 gene expression is used as a standard. Thus, in certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0161] For example, in our examples, we calculated the ratios, and in cells expressing wild type BRCA1 (wtBRCA1) the ratio was around 6.6 for BLM+FANCI/BRCA1. Thus, is some aspects, the expression level can be over 5, or over 6 or over 7 times that of BRCA1.
[0162] For analysis with qPCR, each of the analyzed genes is first normalized to a housekeeping gene, such as RPLP0. For example, with 6 cycles of PCT, we calculated the optimum for BRCA1/BLM/FANCI normalized to RPLP0, expression of which is typically very low. All values are log 2, which means that a ratio of 6 reflects an amount of 64 times higher than the reference gene, namely RPLP0.
[0163] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Examples of useful housekeeping genes are described herein, e.g., in Table 1.
[0164] Various embodiments of the present invention provide for a method for selecting platinum-comprising therapy, and optionally administering the platinum-comprising therapy, for a subject having cancer, comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; detecting the BLM and FANCI expression in the sample compared to a reference value; and selecting a platinum-comprising cancer therapy for the subject when the BLM and FANCI expression compared to a reference value is increased based on the recognition that platinum-comprising cancer therapy is effective in patients whose cancer has increased BLM and FANCI expression compared to the reference value.
[0165] In various embodiments, the method further comprises administering to the subject the platinum-comprising cancer therapy when the platinum-comprising cancer therapy is selected.
[0166] In various embodiments, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. The determination can be made before, concurrently with, or after the analysis for BLM and FANCI expression. In various embodiments, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. The determination can be made before, concurrently with, or after analysis for BLM and FANCI expression.
[0167] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0168] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about two-fold compared to BRCA1 expression. In certain embodiments, the BLM and/or FANCI expression is increased by at least or about 6-fold compared to BRCA1 expression.
[0169] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Housekeeping genes are described herein.
[0170] In certain embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0171] Various embodiments of the present invention provide for a method for selecting a non-platinum-comprising therapy, and optionally administering the non-platinum-comprising therapy, for a subject having cancer comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; detecting the BLM and FANCI expression in the sample compared to a reference value; and selecting the non-platinum-comprising cancer therapy for the subject when the BLM and FANCI expression compared to the reference value is not increased based on the recognition that non-platinum-comprising cancer therapy is effective in patients whose cancer does not have increased gene expression of BLM and FANCI compared to the reference value.
[0172] In some aspects of all the embodiments of the invention a dual assay allowing analysis of both BLM and FANCI expression to be compared in the same assay is used. The dual assay may be based on detecting RNA or protein. The assay may be specific for the dual analysis of BLM and FANCI or may comprise reagents for assaying one, two, three or more other biomolecules as well. In some aspects of all the embodiments of the invention the one other biomolecule is BRCA1.
[0173] In various embodiments, the method further comprises administering to the subject the non-platinum-comprising cancer therapy when non-platinum-comprising cancer therapy is selected.
[0174] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0175] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0176] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Examples of housekeeping genes are described herein although these genes are well known to one of ordinary skill in the art.
[0177] In various embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0178] Various embodiments of the present invention provide for an assay for selecting a therapy for a subject having cancer, comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; comparing the BLM and FANCI expression, optionally in a dual assay, to a reference value; and selecting an anthracycline-comprising cancer therapy for the subject when the BLM and FANCI expression is increased compared to a reference value based on the recognition that anthracycline-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI, or selecting a non-anthracycline-comprising cancer therapy for the subject when the BLM and FANCI expression is not increased compared to a reference value based on the recognition that anthracycline-comprising cancer therapy is not effective in subjects whose cancer does not have increased BLM expression compared to a reference value.
[0179] Various embodiments of the present invention provide for a method for selecting an anthracycline-comprising cancer therapy for a subject having cancer and determined to be negative for BRCA1 and BRCA2 mutations, comprising: subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression comparing the BLM and FANCI expression to a reference value; and selecting the anthracycline-comprising cancer therapy for the subject when the BLM and FANCI expression compared to the reference value is increased based on the recognition that anthracycline-comprising cancer therapy is effective in patients whose cancer has increased expression of BLM and FANCI compared to the reference value.
[0180] Various embodiments provide for an assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; comparing the chromosome 15q26 copy number to a reference value; and selecting a platinum-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or selecting a non-platinum-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0181] In various embodiments, the assay further comprises assaying the BRCA1 and/or BRCA2 status of the subject; and selecting the platinum-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and there is a chromosome 15q26 copy number gain based on the recognition that platinum-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain who are negative for BRCA1 and/or BRCA2 mutations. In various embodiments, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. The determination can be made before, concurrently with, or after the analysis for BLM and FANCI expression.
[0182] In various embodiments, the assay further comprises assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and selecting the platinum-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when there is a chromosome 15q26 copy number gain based on the recognition that platinum-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor. In various embodiments, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. The determination can be made before, concurrently with, or after analysis for chromosome 15q26 copy number.
[0183] In various embodiments, the assay further comprises administering the selected therapy to the subject.
[0184] In various embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0185] In various embodiments, the reference value is chromosome 15 centromere copy number.
[0186] Various embodiments of the present invention provide for an assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; comparing the chromosome 15q26 copy number to a reference value; and selecting an anthracycline-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or selecting a non-anthracycline-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0187] In various embodiments, the assay further comprises assaying the BRCA1 and/or BRCA2 status of the subject; and selecting the anthracycline-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and there is a chromosome 15q26 copy number gain based on the recognition that anthracycline-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain and who are negative for BRCA1 and/or BRCA2 mutations. In various embodiments, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. In some aspects of all the embodiments of the invention, a mutation that inactivates BRCA2 is highly predictive of response to platinum-comprising cancer therapy. The determination can be made before, concurrently with, or after analysis for chromosome 15q26 copy number.
[0188] In various embodiments, the assay further comprises assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and selecting the anthracycline-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when there is a chromosome 15q26 copy number gain based on the recognition that anthracycline-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor. In various embodiments, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. The determination can be made before, concurrently with, or after analysis for chromosome 15q26 copy number.
[0189] In various embodiments, the assay further comprises administering the selected therapy to the subject.
[0190] In various embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0191] In various embodiments, the reference value is chromosome 15 centromere copy number.
Cancer Treatment
[0192] Various embodiments of the present invention provide for a method of treating cancer in a human subject, comprising: detecting BLM and FANCI expression in a sample comprising a cancer cell taken from the human subject; and comparing the BLM and FANCI expression to a reference value; and administering a platinum-comprising cancer therapy to the human subject wherein an increase of BLM and FANCI expression compared to the reference value is detected.
[0193] In various embodiments, the cancer is selected from breast, ovarian, and lung cancers.
[0194] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0195] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0196] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Housekeeping genes are described herein.
[0197] In certain embodiments, the human subject's cancer or cancer cell is known to not or determined not to express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. The determination can be made before, concurrently with, or after analysis for chromosome 15q26 copy number.
[0198] Various embodiments of the present invention provide for a method of treating cancer in a human subject, comprising: detecting BLM and FANCI expression in a sample comprising a cancer cell taken from the human subject; and comparing the BLM and FANCI expression to a reference value; and administering an anthracycline-comprising cancer therapy to the human subject wherein an increase of BLM and FANCI expression compared to the reference value is detected.
[0199] In various embodiments, the human subject's cancer is known to not or determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. The determination can be made before, concurrently with, or after analysis for chromosome 15q26 copy number.
[0200] In various embodiments, the cancer is selected from breast, ovarian, and lung cancers.
[0201] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0202] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0203] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Housekeeping genes are described herein, e.g., in Table 1.
[0204] Various embodiments of the present invention provide for a method of treating cancer, comprising: assaying, in a cancer cell from a cancer patient or mRNA obtained therefrom, the BLM and FANCI expression compared to a reference value; and administering to the cancer patient a cancer treatment regimen comprising platinum or anthracycline if the BLM and FANCI expression is increased compared to the reference value.
[0205] Various embodiments provide for a use of platinum comprising cancer therapy for treating a cancer patient that has been determined to have a tumor comprising cancer cells wherein BLM and FANCI expression is increased compared to a reference value.
[0206] In various embodiments, the cancer patient is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. The determination can be made before, concurrently with, or after the analysis for BLM and FANCI expression. In certain embodiments, the cancer patient's cancer or cancer cell is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. The determination can be made before, concurrently with, or after analysis of BLM and FANCI expression.
[0207] Various embodiments of the present invention provide for a method of treating cancer in a human subject whose cancer has increased BLM and FANCI expression, comprising: identifying the human subject whose cancer has increased BLM and FANCI expression; and administering a platinum-comprising cancer therapy or an anthracycline-comprising therapy to the human subject. In certain embodiments, the human subject's cancer is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. In various embodiments, the human subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. These determinations can be made before, concurrently with, or after analysis of BLM and FANCI expression.
[0208] Various embodiments of the present invention provide for a method of treating cancer in a human subject, comprising: detecting a chromosome 15q26 copy number in a sample comprising a cancer cell taken from the subject; comparing the chromosome 15q26 copy number to a reference value; and administering an platinum-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or administering a non-platinum-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0209] In various embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer. In various embodiments, the reference value is chromosome 15 centromere copy number. In various embodiments, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. The determination can be made before, concurrently with, or after analysis of chromosome 15q26 copy number.
[0210] Various embodiments of the present invention provide for a method of treating cancer in a human subject, comprising: detecting a chromosome 15q26 copy number in a sample comprising a cancer cell taken from the subject; comparing the chromosome 15q26 copy number to a reference value; and administering an anthracycline-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or administering a non-anthracycline-compri sing cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss.
[0211] In various embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer. In various embodiments, the reference value is chromosome 15 centromere copy number. In various embodiments, the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. The determination can be made before, concurrently with, or after analysis of chromosome 15q26 copy number.
Assessing and Predicting Responsiveness to Cancer Therapy
[0212] Various embodiments of the present invention provide for a method for assessing responsiveness of a cancer cell to cancer therapy, comprising: assaying, in a cancer cell or mRNA derived therefrom, BLM and FANCI expression; and comparing said BLM and FANCI expression to a reference value, wherein the cancer cell is assessed as responsive to a platinum-comprising therapy if the BLM and FANCI expression is increased compared to the reference value, or wherein the cancer cell is assessed as poorly or not responsive to platinum-comprising cancer therapy cancer if the BLM and FANCI expression is not increased.
[0213] In various embodiments, the step of assaying comprises: contacting the cancer cell or mRNA derived therefrom with at least one detectably labeled probe capable of specifically binding to BLM mRNA, at least one detectably probe capable of specifically binding to FANCI, at least one detectably labeled probe capable of specifically binding to BRCA1 and/or at least one housekeeping gene; and measuring the expression of BLM and FANCI compared to the BRCA1 and/or the at least one housekeeping gene.
[0214] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0215] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0216] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Housekeeping genes are described herein, e.g., in Table 1.
[0217] Various embodiments of the present invention provide for a method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: determining, in a cancer cell from the cancer patient, BLM and FANCI expression; and correlating the expression to a reference value, wherein when the expression is increased the patient is predicted to respond well to a cancer treatment regimen comprising platinum or anthracycline.
[0218] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0219] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0220] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Housekeeping genes are described herein, e.g., in Table 1.
[0221] Various embodiments of the present invention provide for a method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: determining, in a cancer cell or mRNA derived therefrom from said cancer patient, BLM and FANCI expression; and correlating the expression to a reference value, wherein when the expression is not increased the patient is predicted to respond poorly to a cancer treatment regimen comprising platinum or anthracycline.
[0222] In various embodiments, the reference value is based on BRCA1 gene expression in the cancer cell. In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0223] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression.
[0224] In various embodiments, the reference value is based on a housekeeping gene expression in the cancer cell. Housekeeping genes are described herein.
[0225] Various embodiments provide for a method for assessing responsiveness of a cancer cell to a cancer therapy, and optionally administering the cancer therapy, comprising: assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; and comparing the chromosome 15q26 copy number to a reference value, wherein the cancer cell is assessed as responsive to a platinum-comprising therapy if there is a chromosome 15q26 copy number gain compared to the reference value, or wherein the cancer cell is assessed as poorly or not responsive to platinum-comprising cancer therapy cancer if there is not a chromosome 15q26 copy number gain or if there is a chromosome 15q26 copy number loss.
[0226] In various embodiments, the reference value is chromosome 15 centromere copy number. In various embodiments, the cancer is selected from breast cancer, ovarian cancer and lung cancer.
[0227] In various embodiments, the method further comprises administering the platinum-comprising therapy if there is a chromosome 15q26 copy number gain.
[0228] Various embodiments provide for a method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: determining, in a cancer cell from the cancer patient, chromosome 15q26 copy number; and correlating the chromosome 15q26 copy number to a reference value, wherein when there is a chromosome 15q26 copy number gain, the patient is predicted to respond well to a cancer treatment comprising platinum or anthracycline, or wherein when there is not a chromosome 15q26 copy number gain or a chromosome 15q26 copy number loss, the patient is predicted respond poorly to a cancer treatment comprising platinum or anthracycline.
[0229] In various embodiments, the reference value is chromosome 15 centromere copy number.
[0230] Various embodiments provide for the use of platinum comprising cancer therapy for treating a cancer patient that has been determined to have a tumor comprising cancer cells wherein a chromosome 15q26 copy gain is detected compared to a reference value. In various embodiments, cancer patient is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. The determination can be made before, concurrently with, or after the analysis for BLM and FANCI expression. In various embodiments, the cancer patient is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. In various embodiments, the reference value is chromosome 15 centromere copy number.
Systems, Computers, Kits and Uses
[0231] Various embodiments of the present invention provide for a system for determining responsiveness of a cancer cell to platinum-comprising therapy from a cancer cell of a cancer patient, comprising: a sample analyzer configured to produce a signal for the mRNA from each one of BLM and FANCI from a cancer cell sample of a cancer patient; and a computer sub-system programmed to calculate, based on the mRNA whether the signal is greater or not than a reference value.
[0232] In various embodiments, said computer sub-system is programmed to compare the mRNA to determine a likelihood of responsiveness of said cancer cell to platinum-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a platinum-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the platinum-comprising therapy if the BLM and FANCI expression is not increased; or a likelihood of responsiveness of said cancer cell to anthracycline-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a anthracycline-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the anthracycline-comprising therapy if the BLM and FANCI expression is not increased.
[0233] Various embodiments of the present invention provide for a system for determining responsiveness of a cancer cell to platinum-comprising therapy from a cancer cell of a cancer patient, comprising: a sample analyzer configured to produce a signal for the protein from each one of BLM and FANCI from a cancer cell sample of a cancer patient; and a computer sub-system programmed to calculate, based on the protein whether the signal is greater or not than a reference value.
[0234] In various embodiments, said computer sub-system is programmed to compare the protein to determine a likelihood of responsiveness of said cancer cell to platinum-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a platinum-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the platinum-comprising therapy if the BLM and FANCI expression is not increased; or a likelihood of responsiveness of said cancer cell to anthracycline-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a anthracycline-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the anthracycline-comprising therapy if the BLM and FANCI expression is not increased.
[0235] Various embodiments of the present invention provide for a computer program product embodied in a computer readable medium that, when executing on a computer, performs steps comprising: detecting the BLM and FANCI expression in sample comprising a cancer cell from a cancer patient; and comparing the BLM and FANCI expression to a reference value.
[0236] Various embodiments of the present invention provide for a diagnostic kit for detecting a likelihood of a cancer patient to respond to platinum- or anthracycline-compri sing comprising cancer therapy, comprising: no more than 10 probes comprising a combination of detectably labeled probes or primers for BLM and FANCI, and optionally for BRCA1 and/or at least one housekeeping gene; and the computer program product.
[0237] Various embodiments of the present invention provide for the use of a plurality of oligonucleotides comprising no more than 10 oligonucleotides capable of hybridizing to BLM and FANCI, and optionally to BRCA1 and/or at least one housekeeping gene, in a diagnostic kit for determining an increased likelihood that a cancer patient will respond to cancer treatment regimen comprising a platinum and/or anthracycline.
[0238] Various embodiments provide for a system for determining responsiveness of a cancer cell to platinum-comprising therapy from a cancer cell of a cancer patient, comprising: a sample analyzer configured to produce a signal for chromosome 15q26 copy number from a cancer cell sample of a cancer patient; and a computer sub-system programmed to calculate, based on the mRNA whether the signal is greater or not than a reference value.
[0239] In various embodiments, the computer sub-system is programmed to compare the mRNA to determine a likelihood of responsiveness of said cancer cell to platinum-comprising cancer therapy and/or or a anthracycline-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a platinum-comprising therapy if there is a chromosome 15q26 copy number gain and as unlikely to respond to the platinum-comprising therapy if there is not a chromosome 15q26 copy number gain or if there is a chromosome 15q26 copy number loss. In various embodiments, the reference value is chromosome 15 centromere copy number.
[0240] Various embodiments of the present invention provide for a computer program product embodied in a computer readable medium that, when executing on a computer, performs steps comprising: detecting chromosome 15q26 copy number in sample comprising a cancer cell from a cancer patient; and comparing the chromosome 15q26 copy number to a reference value. In various embodiments, the reference value is chromosome 15 centromere copy number.
[0241] Various embodiments of the present invention provide for a diagnostic kit for detecting a likelihood of a cancer patient to respond to platinum- or anthracycline-compri sing cancer therapy, comprising: no more than 10 probes comprising a combination of detectably labeled probes or primers for chromosome 15q26, and optionally for chromosome 15 centromere; and the computer program product of described herein.
[0242] Various embodiments of the present invention provide for the use of a plurality of oligonucleotides comprising no more than 10 oligonucleotides capable of hybridizing to chromosome 15q26, and optionally for chromosome 15 centromere, in a diagnostic kit for determining an increased likelihood that a cancer patient will respond to cancer treatment regimen comprising a platinum and/or anthracycline.
Reference Values
[0243] In various embodiments of the present invention, the reference value is based on BRCA1 gene expression in the cancer cell.
[0244] In various embodiments, the reference value for BLM expression or FANCI expression is BRCA1 expression.
[0245] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to BRCA1 expression.
[0246] In various embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold, 2.8-fold, 2.9-fold, or 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold or 10-fold compared to BRCA1 expression. In certain embodiments, the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least or about two-fold to about 7-fold compared to BRCA1 expression.
[0247] In various embodiments of the present invention, the reference value can be one or more housekeeping gene as described herein. In certain embodiments, one or more housekeeping gene as described herein, and the BLM and/or FANCI expression is increased by at least or about 10, 20, 30, 40, 50, 60, 70, 80, or 90% compared to the one or more housekeeping gene expression.
[0248] In certain embodiments, one or more housekeeping gene as described herein, and the BLM and/or FANCI expression is increased by at least or about 1-fold, 1.1-fold, 1.2-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.8-fold, 1.9-fold, 2-fold, 2.1-fold 2.2-fold 2.3-fold 2.4-fold 2.5-fold, 2.6-fold, 2.7-fold 2.8-fold 2.9-fold or 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 30-fold, 40-fold, 50-fold, 60-fold or higher compared to the one or more housekeeping gene expression depending on the level of the expression of the housekeeping gene in the cell.
[0249] In various embodiments, the housekeeping gene can be selected from the group consisting of beta-actin, GAPDH, RPLP0, GUS, TFRC and combinations thereof. (See e.g., Cronin et al., Clinical Chemistry 53:6, 1084-1091 (2007), and ONCOTYPE DX ASSAY.RTM., herein incorporated by reference in its entirety.) Thus, in some embodiments, the house keeping gene is one of these genes, and in other embodiments, the housekeeping gene is a combination of 2, 3, 4, or all 5 of these genes. In particular embodiments, the housekeeping gene is RPLP0, also called 36b4.
[0250] In various embodiments, the house keeping gene can be selected from the genes listed in Table 1. Accordingly, in some embodiments, the housekeeping gene is one of the genes from Table 1, and in other embodiments, the housekeeping gene is a combination of any number or all of the genes from Table 1. (See e.g., Eisenberg and Levanon, Human housekeeping genes are compact. Trends in Genetics, Volume 19, Issue 7, 362-365, 1 July 2003). Each gene name/description is followed by its geometric average expression level according to the data published by Su et al. Thus, based on our experimental data on the RPLP0 housekeeping gene as a reference gene, one of ordinary skill in the art can easily determine what the cut-off points for increased expression for any one of these genes is. For example, genes designated by asterisk are in popular use as reference in real-time PCR or quantitative PCR (qPCR), which is most often used in gene expression analysis.
[0251] In some aspects of all the embodiments on the invention, the assays, methods, kits, and systems incorporate qPCR as the gene expression analysis method to determine the amount compared to a reference value.
TABLE-US-00001 TABLE 1 Housekeeping Genes. Accession No. Description *NM_001101 Homo sapiens actin, beta (ACTB), mRNA 6988 *NM_000034 Homo sapiens aldolase A,fructose-bisphosphate (ALDOA), mRNA 3425 *NM_002046 Homo sapiens glyceraldehyde-3-phosphate dehydrogenase (GAPD), mRNA 828 *NM_000291 Homo sapiens phosphoglycerate kinase 1 (PGK1), mRNA 2727 *NM_005566 Homo sapiens lactate dehydrogenase A (LDHA), mRNA 2105 *NM_002954 Homo sapiens ribosomal protein S27a (RPS27A), mRNA 4156 *NM_000981 Homo sapiens ribosomal protein L19 (RPL19), mRNA 6997 *NM_000975 Homo sapiens ribosomal protein L11 (RPL11), mRNA 6060 *NM_007363 Homo sapiens non-POU domain containing, octamer- binding (NONO), mRNA 1708 *NM_004309 Homo sapiens Rho GDP dissociation inhibitor (GDI) alpha (ARHGDIA), mRNA 1358 *NM_000994 Homo sapiens ribosomal protein L32 (RPL32), mRNA 9523 *NM_022551 Homo sapiens ribosomal protein S18 (RPS18), mRNA 11261 *NM_007355 Homo sapiens heat shock 90 kDa protein 1, beta (HSPCB), mRNA 4119 NM_004515 Homo sapiens interleukin enhancer binding factor 2, 45 kDa (ILF2), mRNA 1000 NM_004651 Homo sapiens ubiquitin specific protease 11 (USP11), mRNA 1950 NM_004888 Homo sapiens ATPase, H+ transporting, lysosomal 13 kDa, V1 subunit G isoform 1 (ATP6V1G1), mRNA 928 NM_003334 Homo sapiens ubiquitin-activating enzyme El (A1S9T and BN75 temperature sensitivity complementing) (UBE1), transcript variant 1, mRNA 1351 NM_001320 Homo sapiens casein kinase 2, beta polypeptide (CSNK2B), mRNA 1233 NM_003915 Homo sapiens copine I (CPNE1), transcript variant 3, mRNA 698 NM_001250 Homo sapiens tumor necrosis factor receptor superfamily, member 5 (TNFRSF5), transcript variant 1, mRNA 524 NM_001904 Homo sapiens catenin (cadherin-associated protein), beta 1, 88 kDa (CTNNB1), mRNA 1165 NM_003753 Homo sapiens eukaryotic translation initiation factor 3, subunit 7 zeta, 66/67 kDa (EIF3S7), mRNA 1363 NM_004541 Homo sapiens NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 1, 7.5 kDa (NDUFA1), nuclear gene encoding mitochondrial protein, mRNA 2307 NM_001654 Homo sapiens v-raf murine sarcoma 3611 viral oncogene homolog 1 (ARAF1), mRNA 846 NM_002967 Homo sapiens scaffold attachment factor B (SAFB), mRNA 545 NM_001183 Homo sapiens ATPase, H+ transporting, lysosomal interacting protein 1 (ATP6IP1), mRNA 1089 NM_003526 Homo sapiens H2B histone family, member L (H2BFL), mRNA 2232 NM_004718 Homo sapiens cytochrome c oxidase subunit VIIa polypeptide 2 like (COX7A2L), nuclear gene encoding mitochondrial protein, mRNA 1013 NM_004436 Homo sapiens endosulfine alpha (ENSA), mRNA 659 NM_001207 Homo sapiens basic transcription factor 3 (BTF3), mRNA 3348 NM_004907 Homo sapiens immediate early protein (ETR101), mRNA 1249 NM_004889 Homo sapiens ATP synthase, H+ transporting, mitochondrial F0 complex, subunit f, isoform 2 (ATP5J2), mRNA 1513 NM_003769 Homo sapiens splicing factor, arginine/serine-rich 9 (SFRS9), mRNA 1207 NM_003910 Homo sapiens maternal G10 transcript (G10), mRNA 696 NM_000100 Homo sapiens cystatin B (stefin B) (CSTB), mRNA 1422 NM_004785 Homo sapiens solute carrier family 9 (sodium/hydrogen exchanger), isoform 3 regulatory factor 2 (SLC9A3R2), mRNA 455 NM_001120 Homo sapiens tetracycline transporter-like protein (TETRAN), mRNA 935 NM_000182 Homo sapiens hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl- Coenzyme A hydratase (trifunctional protein), alpha subunit (HADHA), mRNA 609 NM_003377 Homo sapiens vascular endothelial growth factor B (VEGFB), mRNA 649 NM_003576 Homo sapiens serine/threonine kinase 24 (STE20 homolog, yeast) (STK24), mRNA 769 NM_000918 Homo sapiens procollagen-proline, 2-oxoglutarate 4- dioxygenase (proline 4-hydroxylase), beta polypeptide (protein disulfide isomerase; thyroid hormone binding protein p55) (P4HB), mRNA 2719 NM_004584 Homo sapiens RAD9 homolog (S. pombe) (RAD9), mRNA 860 NM_004952 Homo sapiens ephrin-A3 (EFNA3), mRNA 505 NM_004308 Homo sapiens Rho GTPase activating protein 1 (ARHGAP1), mRNA 893 NM_003190 Homo sapiens TAP binding protein (tapasin) (TAPBP), mRNA 1213 NM_004640 Homo sapiens HLA-B associated transcript 1 (BAT1), transcript variant 1, mRNA 1022 NM_001064 Homo sapiens transketolase (Wernicke-Korsakoff syndrome) (TKT), mRNA 1139 NM_002117 Homo sapiens major histocompatibility complex, class I, C (HLA-C), mRNA 4696 NM_004161 Homo sapiens RAB1A, member RAS oncogene family (RAB1A), mRNA 2163 NM_003339 Homo sapiens ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) (UBE2D2), mRNA 514 NM_003969 Homo sapiens ubiquitin-conjugating enzyme E2M (UBC12 homolog, yeast) (UBE2M), mRNA 1197 NM_000516 Homo sapiens GNAS complex locus (GNAS), transcript variant 1, mRNA 4358 NM_002819 Homo sapiens polypyrimidine tract binding protein 1 (PTBP1), transcript variant 1, mRNA 1138 NM_001001 Homo sapiens ribosomal protein L36a-like (RPL36AL), mRNA 1740 NM_004649 Homo sapiens chromosome 21 open reading frame 33 (C21orf33), mRNA 585 NM_000175 Homo sapiens glucose phosphate isomerase (GPI), mRNA 1633 NM_001867 Homo sapiens cytochrome c oxidase subunit VIIc (COX7C), nuclear gene encoding mitochondrial protein, mRNA 3004 NM_001967 Homo sapiens eukaryotic translation initiation factor 4A, isoform 2 (EIF4A2), mRNA 2935 NM_001863 Homo sapiens cytochrome c oxidase subunit VIb (COX6B), nuclear gene encoding mitochondrial protein, mRNA 1719 NM_001997 Homo sapiens Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed (fox derived); ribosomal protein S30 (FAU), mRNA 3898 NM_002088 Homo sapiens glutamate receptor, ionotropic, kainate 5 (GRIK5), mRNA 535 NM_001862 Homo sapiens cytochrome c oxidase subunit Yb (COX5B), nuclear gene encoding mitochondrial protein, mRNA 1087 NM_004255 Homo sapiens cytochrome c oxidase subunit Va (COX5A), nuclear gene encoding mitochondrial protein, mRNA 853 NM_001788 Homo sapiens CDC10 cell division cycle 10 homolog (S. cerevisiae) (CDC10), mRNA 1340 NM_004781 Homo sapiens vesicle-associated membrane protein 3 (cellubrevin) (VAMP3), mRNA 684 NM_003801 Homo sapiens GPAA1P anchor attachment protein 1 homolog (yeast) (GPAA1), mRNA 776 NM_004643 Homo sapiens poly(A) binding protein, nuclear 1 (PABPN1), mRNA 502 NM_001537 Homo sapiens heat shock factor binding protein 1 (HSBP1), mRNA 874 NM_003680 Homo sapiens tyrosyl-tRNA synthetase (YARS), mRNA 535 NM_003345 Homo sapiens ubiquitin-conjugating enzyme E21 (UBC9 homolog, yeast) (UBE2I), mRNA 1283 NM_002568 Homo sapiens poly(A) binding protein, cytoplasmic 1 (PABPC1), mRNA 3199 NM_001487 Homo sapiens GCN5 general control of amino-acid synthesis 5-like 1 (yeast) (GCN5L1), mRNA 313 NM_001861 Homo sapiens cytochrome c oxidase subunit IV isoform 1 (COX4I1), nuclear gene encoding mitochondrial protein, mRNA2 738 NM_004890 Homo sapiens sperm associated antigen 7 (SPAG7), mRNA 618 NM_002812 Homo sapiens proteasome (prosome, macropain) 26S subunit, non-ATPase, 8 (PSMD8), mRNA 942 NM_004926 Homo sapiens zinc finger protein 36, C3H type-like 1 (ZFP36L1), mRNA 1168 NM_002539 Homo sapiens ornithine decarboxylase 1 (ODC1), mRNA 1361 NM_000979 Homo sapiens ribosomal protein L18 (RPL18), mRNA 4417 NM_000977 Homo sapiens ribosomal protein L13 (RPL13), transcript variant 1, mRNA 6407 NM_001015 Homo sapiens ribosomal protein S11 (RPS11), mRNA 7614 NM_001760 Homo sapiens cyclin D3 (CCND3), mRNA 676 NM_003973 Homo sapiens ribosomal protein L14 (RPL14), mRNA 3135 NM_002815 Homo sapiens proteasome (prosome, macropain) 26S subunit, non-ATPase, 11 (PSMD11), mRNA 536 NM_000367 Homo sapiens thiopurine S-methyltransferase (TPMT), mRNA 1574 NM_000973 Homo sapiens ribosomal protein L8 (RPL8), transcript variant 1, mRNA 8138 NM_004689 Homo sapiens metastasis associated 1 (MTA1), mRNA 506 NM_001848 Homo sapiens collagen, type VI, alpha 1 (COL6A1), mRNA 757 NM_004068 Homo sapiens adaptor-related protein complex 2, mu 1 subunit (AP2M1), mRNA 2188 NM_001687 Homo sapiens ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit (ATP5D), mRNA 1167 NM_004197 Homo sapiens serine/threonine kinase 19 (STK19), transcript variant 1, mRNA 574 NM_001028 Homo sapiens ribosomal protein S25 (RPS25), mRNA 4683 NM_001022 Homo sapiens ribosomal protein S19 (RPS19), mRNA 6683 NM_004759 Homo sapiens mitogen-activated protein kinase-activated protein kinase 2 (MAPKAPK2), transcript variant 1, mRNA 641 NM_001623 Homo sapiens allograft inflammatory factor 1 (AIF1), transcript variant 3, mRNA 497 NM_004894 Homo sapiens chromosome 14 open reading frame 2 (C14orf2), mRNA 704 NM_002375 Homo sapiens microtubule-associated protein 4 (MAP4), transcript variant 1, mRNA 717 NM_001013 Homo sapiens ribosomal protein S9 (RPS9), mRNA 6868 NM_003779 Homo sapiens UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 3 (B4GALT3), mRNA 565 NM_001296 Homo sapiens chemokine binding protein 2 (CCBP2), mRNA 394 NM_001009 Homo sapiens ribosomal protein S5 (RPS5), mRNA 6739 NM_003021 Homo sapiens small glutamine-rich tetratricopeptide repeat (TPR)-containing (SGT), mRNA 488 NM_004285 Homo sapiens hexose-6-phosphate dehydrogenase (glucose 1-dehydrogenase) (H6PD), mRNA 646 NM_004142 Homo sapiens matrix metalloproteinase-like 1 (MMPL1), mRNA 695 NM_001950 Homo sapiens E2F transcription factor 4, p107/p130- binding (E2F4), mRNA 956 NM_003815 Homo sapiens a disintegrin and metalloproteinase domain 15 (metargidin) (ADAM15), mRNA 771 NM_001119 Homo sapiens adducin 1 (alpha) (ADD1), transcript variant 1, mRNA 1356 NM_001111 Homo sapiens adenosine deaminase, RNA-specific (ADAR), transcript variant ADAR-a, mRNA 1036 NM_003466 Homo sapiens paired box gene 8 (PAX8), transcript variant PAX8A, mRNA 901 NM_001155 Homo sapiens annexin A6 (ANXA6), transcript variant 1, mRNA 718 NM_003465 Homo sapiens chitinase 1 (chitotriosidase) (CHIT1), mRNA 561 NM_003186 Homo sapiens transgelin (TAGLN), mRNA 1209 NM_000802 Homo sapiens folate receptor 1 (adult) (FOLR1), transcript variant 2, mRNA 514 NM_004924 Homo sapiens actinin, alpha 4 (ACTN4), mRNA 1187 NM_002931 Homo sapiens ring finger protein 1 (RING 1), mRNA 576 NM_000020 Homo sapiens activin A receptor type II-like 1 (ACVRL1), mRNA 849 NM_001785 Homo sapiens cytidine deaminase (CDA), mRNA 391 NM_004339 Homo sapiens pituitary tumor-transforming 1 interacting protein (PTTG1 IP), mRNA 1200 NM_003860 Homo sapiens Breakpoint cluster region protein, uterine leiomyoma, 1; barrier to autointegration factor (BCRP1), mRNA 1303 NM_000214 Homo sapiens jagged 1 (Alagille syndrome) (JAGO, mRNA 536
NM_002167 Homo sapiens inhibitor of DNA binding 3, dominant negative helix-loop-helix protein (ID3), mRNA 1192 NM_001664 Homo sapiens ras homolog gene family, member A (ARHA), mRNA 4050 NM_003166 Homo sapiens sulfotransferase family, cytosolic, 1A, phenol-preferring, member 3 (SULT1A3), mRNA 461 NM_001746 Homo sapiens calnexin (CANX), mRNA 1923 NM_001662 Homo sapiens ADP-ribosylation factor 5 (ARF5), mRNA 724 NM_001660 Homo sapiens ADP-ribosylation factor 4 (ARF4), mRNA 1014 NM_001658 Homo sapiens ADP-ribosylation factor 1 (ARF1), mRNA 2195 NM_003313 Homo sapiens tissue specific transplantation antigen P35B (TSTA3), mRNA 440 NM_001494 Homo sapiens GDP dissociation inhibitor 2 (GDI2), mRNA 1352 NM_003145 Homo sapiens signal sequence receptor, beta (translocon- associated protein beta) (SSR2), mRNA 1265 NM_001619 Homo sapiens adrenergic, beta, receptor kinase 1 (ADRBK1), mRNA 695 NM_001420 Homo sapiens ELAV (embryonic lethal, abnormal vision, Drosophila)-like 3 (Hu antigen C) (ELAVL3), mRNA 1070 NM_004930 Homo sapiens capping protein (actin filament) muscle Z- line, beta (CAPZB), mRNA 1183 NM_004596 Homo sapiens small nuclear ribonucleoprotein polypeptide A (SNRPA), mRNA 734 NM_004168 Homo sapiens succinate dehydrogenase complex, subunit A, flavoprotein (Fp) (SDHA), nuclear gene encoding mitochondrial protein, mRNA 1106 NM_004156 Homo sapiens protein phosphatase 2 (formerly 2A), catalytic subunit, beta isoform (PPP2CB), mRNA 1195 NM_004910 Homo sapiens phosphatidylinositol transfer protein, membrane-associated (PITPNM), mRNA 869 NM_004517 Homo sapiens integrin-linked kinase (ILK), mRNA 654 NM_004494 Homo sapiens hepatoma-derived growth factor (high- mobility group protein 1-like) (HDGF), mRNA 1385 NM_004121 Homo sapiens gamma-glutamyltransferase-like activity 1 (GGTLA1), mRNA 412 NM_004404 Homo sapiens neural precursor cell expressed, developmentally down-regulated 5 (NEDD5), mRNA 1571 NM_004394 Homo sapiens death-associated protein (DAP), mRNA 623 NM_004383 Homo sapiens c-src tyrosine kinase (CSK), mRNA 899 NM_004074 Homo sapiens cytochrome c oxidase subunit VIII (COX8), nuclear gene encoding mitochondrial protein, mRNA 3188 NM_004039 Homo sapiens annexin A2 (ANXA2), mRNA 2417 NM_001053 Homo sapiens somatostatin receptor 5 (SSTR5), mRNA 423 NM_001328 Homo sapiens C-terminal binding protein 1 (CTBP1), mRNA 798 NM_001273 Homo sapiens chromodomain helicase DNA binding protein 4 (CHD4), mRNA 775 NM_003430 Homo sapiens zinc finger protein 91 (HPF7, HTF10) (ZNF91), mRNA 716 NM_003314 Homo sapiens tetratricopeptide repeat domain 1 (TTC1), mRNA 651 NM_003217 Homo sapiens testis enhanced gene transcript (TEGT), mRNA 1766 NM_003132 Homo sapiens spermidine synthase (SRM), mRNA 1093 NM_000199 Homo sapiens N-sulfoglucosamine sulfohydrolase (sulfamidase) (SGSH), mRNA 565 NM_002818 Homo sapiens proteasome (prosome, macropain) activator subunit 2 (PA28 beta) (PSME2), mRNA 697 NM_002733 Homo sapiens protein kinase, AMP-activated, gamma 1 non-catalytic subunit (PRKAG1), mRNA 497 NM_002631 Homo sapiens phosphogluconate dehydrogenase (PGD), mRNA 1009 NM_002574 Homo sapiens peroxiredoxin 1 (PRDX1), mRNA 2241 NM_002512 Homo sapiens non-metastatic cells 2, protein (NM23B) expressed in (NME2), nuclear gene encoding mitochondrial protein, mRNA 2553 NM_002455 Homo sapiens metaxin 1 (MTX1), mRNA 845 NM_002444 Homo sapiens moesin (MSN), mRNA 1798 NM_000529 Homo sapiens melanocortin 2 receptor (adrenocorticotropic hormone) (MC2R), mRNA 420 NM_003573 Homo sapiens latent transforming growth factor beta binding protein 4 (LTBP4), mRNA 740 NM_002315 Homo sapiens LIM_domain only 1 (rhombotin 1) (LMO1), mRNA 469 NM_000884 Homo sapiens IMP (inosine monophosphate) dehydrogenase 2 (IMPDH2), mRNA 1288 NM_003641 Homo sapiens interferon induced transmembrane protein 1 (9-27) (IFITM1), mRNA 4898 NM_000841 Homo sapiens glutamate receptor, metabotropic 4 (GRM4), mRNA 1369 NM_002070 Homo sapiens guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 2 (GNAI2), mRNA 1877 NM_001493 Homo sapiens GDP dissociation inhibitor 1 (GDI1), mRNA 1387 NM_002048 Homo sapiens growth arrest-specific 1 (GAS1), mRNA 940 NM_002032 Homo sapiens ferritin, heavy polypeptide 1 (FTH1), mRNA 2616 NM_001418 Homo sapiens eukaryotic translation initiation factor 4 gamma, 2 (EIF4G2), mRNA 3646 NM_001350 Homo sapiens death-associated protein 6 (DAXX), mRNA 510 NM_001843 Homo sapiens contactin 1 (CNTN1), mRNA 535 NM_001728 Homo sapiens basigin (BSG), mRNA 1508 NM_001667 Homo sapiens ADP-ribosylation factor-like 2 (ARL2), mRNA 965 NM_001659 Homo sapiens ADP-ribosylation factor 3 (ARF3), mRNA 1327 NM_003746 Homo sapiens dynein, cytoplasmic, light polypeptide 1 (DNCL1), mRNA 2758 NM_002127 Homo sapiens HLA-G histocompatibility antigen, class I, G (HLA-G), mRNA 1090 NM_004712 Homo sapiens hepatocyte growth factor-regulated tyrosine kinase substrate (HGS), mRNA 505 NM_003475 Homo sapiens chromosome 11 open reading frame 13 (C11orf13), mRNA 413 NM_004046 Homo sapiens ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit, isoform 1, cardiac muscle (ATP5A1), mRNA 1348 NM_001894 Homo sapiens casein kinase 1, epsilon (CSNK1E), transcript variant 2, mRNA 613 NM_003795 Homo sapiens sorting nexin 3 (SNX3), transcript variant 1, mRNA 1426 NM_001909 Homo sapiens cathepsin D (lysosomal aspartyl protease) (CTSD), mRNA 1512 NM_002792 Homo sapiens proteasome (prosome, macropain) subunit, alpha type, 7 (PSMA7), transcript variant 1, mRNA 728 NM_002799 Homo sapiens proteasome (prosome, macropain) subunit, beta type, 7 (PSMB7), mRNA 545 NM_002300 Homo sapiens lactate dehydrogenase B (LDHB), mRNA 4144 NM_004176 Homo sapiens sterol regulatory element binding transcription factor 1 (SREBF1), mRNA 632 NM_002796 Homo sapiens proteasome (prosome, macropain) subunit, beta type, 4 (PSMB4), mRNA 1229 NM_002794 Homo sapiens proteasome (prosome, macropain) subunit, beta type, 2 (PSMB2), mRNA 1304 NM_002793 Homo sapiens proteasome (prosome, macropain) subunit, beta type, 1 (PSMB1), mRNA 2084 NM_002473 Homo sapiens myosin, heavy polypeptide 9, non-muscle (MYH9), mRNA 1381 NM_001810 Homo sapiens centromere protein B, 80 kDa (CENPB), mRNA 532 NM_002624 Homo sapiens prefoldin 5 (PFDN5), transcript variant 1, mRNA 1718 NM_004710 Homo sapiens synaptogyrin 2 (SYNGR2), mRNA 1414 NM_001127 Homo sapiens adaptor-related protein complex 1, beta 1 subunit (AP1B1), transcript variant 1, mRNA 873 NM_002107 Homo sapiens H3 histone, family 3A (H3F3A), mRNA 9328 NM_003899 Homo sapiens Rho guanine nucleotide exchange factor (GEF) 7 (ARHGEF7), transcript variant 1, mRNA 558 NM_003406 Homo sapiens tyrosine 3-monooxygenase/tryptophan 5- monooxygenase activation protein, zeta polypeptide (YWHAZ), transcript variant 1, mRNA 2008 NM_002419 Homo sapiens mitogen-activated protein kinase kinase kinase 11 (MAP3K11), mRNA 535 NM_001130 Homo sapiens amino-terminal enhancer of split (AES), mRNA 2395 NM_003379 Homo sapiens villin 2 (ezrin) (VIL2), mRNA 1356 NM_002636 Homo sapiens PHD finger protein 1 (PHF1), transcript variant 1, mRNA 640 NM_002622 Homo sapiens prefoldin 1 (PFDN1), mRNA 658 NM_001823 Homo sapiens creatine kinase, brain (CKB), mRNA 1407 NM_003405 Homo sapiens tyrosine 3-monooxygenase/tryptophan 5- monooxygenase activation protein, eta polypeptide (YWHAH), mRNA 2195 NM_002939 Homo sapiens ribonuclease/angiogenin inhibitor (RNH), mRNA 726 NM_003562 Homo sapiens solute carrier family 25 (mitochondrial carrier; oxoglutarate carrier), member 11 (SLC25A11), mRNA 517 NM_001916 Homo sapiens cytochrome c-1 (CYC1), mRNA 895 NM_002823 Homo sapiens prothymosin, alpha (gene sequence 28) (PTMA), mRNA 3723 NM_003096 Homo sapiens small nuclear ribonucleoprotein polypeptide G (SNRPG), mRNA 687 NM_003321 Homo sapiens Tu translation elongation factor, mitochondrial (TUFM), mRNA 1543 NM_003404 Homo sapiens tyrosine 3-monooxygenase/tryptophan 5- monooxygenase activation protein, beta polypeptide (YWHAB), transcript variant 1, mRNA 1144 NM_002946 Homo sapiens replication protein A2, 32 kDa (RPA2), mRNA 537 NM_004356 Homo sapiens CD81 antigen (target of antiproliferative antibody 1) (CD81), mRNA 3638 NM_001743 Homo sapiens calmodulin 2 (phosphorylase kinase, delta) (CALM2), mRNA 4675 NM_004231 Homo sapiens ATPase, H+ transporting, lysosomal 14 kDa, V1 subunit F (ATP6V1F), mRNA 997 NM_004893 Homo sapiens H2A histone family, member Y (H2AFY), transcript variant 2, mRNA 788 NM_004146 Homo sapiens NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 7, 18 kDa (NDUFB7), mRNA 967 NM_002128 Homo sapiens high-mobility group box 1 (HMGB1), mRNA 1295 NM_002002 Homo sapiens Fc fragment of IgE, low affinity II, receptor for (CD23A) (FCER2), mRNA 969 NM_000858 Homo sapiens guanylate kinase 1 (GUK1), mRNA 1166 NM_001469 Homo sapiens thyroid autoantigen 70 kDa (Ku antigen) (G22P1), mRNA 1684 NM_003766 Homo sapiens beclin 1 (coiled-coil, myosin-like BCL2 interacting protein) (BECN1), mRNA 948 NM_003906 Homo sapiens MCM3 minichromosome maintenance deficient 3 (S. cerevisiae) associated protein (MCM3AP), mRNA 430 NM_000757 Homo sapiens colony stimulating factor 1 (macrophage) (CSF1), mRNA 797 NM_002149 Homo sapiens hippocalcin-like 1 (HPCAL1), transcript variant 1, mRNA 1026 NM_001694 Homo sapiens ATPase, H+ transporting, lysosomal 16 kDa, V0 subunit c (ATP6V0C), mRNA 2105 NM_004047 Homo sapiens ATPase, H+ transporting, lysosomal 21 kDa, V0 subunit c'' (ATP6V0B), mRNA 1212 NM_001696 Homo sapiens ATPase, H+ transporting, lysosomal 31 kDa, V1 subunit E isoform 1 (ATP6V1E1), mRNA 784 NM_001865 Homo sapiens cytochrome c oxidase subunit VIIa polypeptide 2 (liver) (COX7A2), nuclear gene encoding mitochondrial protein, mRNA 1818 NM_004373 Homo sapiens cytochrome c oxidase subunit VIa polypeptide 1 (COX6A1), nuclear gene encoding mitochondrial protein, mRNA 5529 NM_000801 Homo sapiens FK506 binding protein 1A, 12 kDa (FKBP1A), transcript variant 12B, mRNA 4273 NM_000992 Homo sapiens ribosomal protein L29 (RPL29), mRNA 6060 NM_000988 Homo sapiens ribosomal protein L27 (RPL27), mRNA 6101 NM_001004 Homo sapiens ribosomal protein, large P2 (RPLP2), mRNA 5924 NM_001003 Homo sapiens ribosomal protein, large, P1 (RPLP1), mRNA 10300 NM_000405 Homo sapiens GM2 ganglioside activator protein (GM2A), mRNA 1250 NM_000967 Homo sapiens ribosomal protein L3 (RPL3), mRNA 7416 NM_001428 Homo sapiens enolase 1, (alpha) (ENO1), mRNA 3668 NM_000999 Homo sapiens ribosomal protein L38 (RPL38), mRNA 8302 NM_000997 Homo sapiens ribosomal protein L37 (RPL37), mRNA 6689 NM_000995 Homo sapiens ribosomal protein L34 (RPL34), transcript variant 1, mRNA 5424 NM_002948 Homo sapiens ribosomal protein L15 (RPL15), mRNA 5450 NM_002952 Homo sapiens ribosomal protein S2 (RPS2), mRNA 8825 NM_001026 Homo sapiens ribosomal protein S24 (RP524), transcript variant 2, mRNA 5701 NM_001020 Homo sapiens ribosomal protein S16 (RPS16), mRNA 7477 NM_001018 Homo sapiens ribosomal protein S15 (RPS15), mRNA 6261
NM_001017 Homo sapiens ribosomal protein S13 (RPS13), mRNA 5430 NM_000969 Homo sapiens ribosomal protein L5 (RPL5), mRNA 4653 NM_000985 Homo sapiens ribosomal protein L17 (RPL17), mRNA 4369 NM_000937 Homo sapiens polymerase (RNA) II (DNA directed) polypeptide A, 220 kDa (POLR2A), mRNA 753 NM_001016 Homo sapiens ribosomal protein S12 (RPS12), mRNA 8265 NM_002140 Homo sapiens heterogeneous nuclear ribonucleoprotein K (HNRPK), transcript variant 1, mRNA 2429 NM_002138 Homo sapiens heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37 kDa) (HNRPD), transcript variant 3, mRNA 1157 NM_004499 Homo sapiens heterogeneous nuclear ribonucleoprotein A/B (HNRPAB), transcript variant 2, mRNA 654 NM_001014 Homo sapiens ribosomal protein S10 (RPS10), mRNA 8074 NM_002383 Homo sapiens MYC-associated zinc finger protein (purine-binding transcription factor) (MAZ), mRNA 2580 NM_002467 Homo sapiens v-myc myelocytomatosis viral oncogene homolog (avian) (MYC), mRNA 537 NM_001436 Homo sapiens fibrillarin (FBL), mRNA 1408 NM_004069 Homo sapiens adaptor-related protein complex 2, sigma 1 subunit (AP2S1), transcript variant AP17, mRNA 979 NM_001614 Homo sapiens actin, gamma 1 (ACTG1), mRNA 6560 NM_002355 Homo sapiens mannose-6-phosphate receptor (cation dependent) (M6PR), mRNA 388 NM_004597 Homo sapiens small nuclear ribonucleoprotein D2 polypeptide 16.5 kDa (SNRPD2), mRNA 1136 NM_002308 Homo sapiens lectin, galactoside-binding, soluble, 9 (galectin 9) (LGALS9), transcript variant short, mRNA 888 NM_000398 Homo sapiens diaphorase (NADH) (cytochrome b-5 reductase) (DIA1), nuclear gene encoding mitochondrial protein, transcript variant M, mRNA 2190 NM_000754 Homo sapiens catechol-O-methyltransferase (COMT), transcript variant MB-COMT, mRNA 845 NM_002406 Homo sapiens mannosyl (alpha-1,3-)-glycoprotein beta- 1,2-N-acetylglucosaminyltransferase (MGAT1), mRNA 893 NM_003752 Homo sapiens eukaryotic translation initiation factor 3, subunit 8, 110 kDa (EIF3S8), mRNA 1871 NM_001355 Homo sapiens D-dopachrome tautomerase (DDT), mRNA 631 NM_004960 Homo sapiens fusion, derived from t(12; 16) malignant liposarcoma (FUS), mRNA 1019 NM_004729 Homo sapiens Ac-like transposable element (ALTE), mRNA 963 NM_004587 Homo sapiens ribosome binding protein 1 homolog 180 kDa (dog) (RRBP1), mRNA 658 NM_004552 Homo sapiens NADH dehydrogenase (ubiquinone) Fe--S protein 5, 15 kDa (NADH-coenzyme Q reductase) (NDUFS5), mRNA 936 NM_004450 Homo sapiens enhancer of rudimentary homolog (Drosophila) (ERH), mRNA 1028 NM_004048 Homo sapiens beta-2-microglobulin (B2M), mRNA 4992 NM_000239 Homo sapiens lysozyme (renal amyloidosis) (LYZ), mRNA 796 NM_000269 Homo sapiens non-metastatic cells 1, protein (NM23A) expressed in (NME1), mRNA 887 NM_000431 Homo sapiens mevalonate kinase (mevalonic aciduria) (MVK), mRNA 753 NM_001247 Homo sapiens ectonucleoside triphosphate diphosphohydrolase 6 (putative function) (ENTPD6), mRNA 495 NM_003365 Homo sapiens ubiquinol-cytochrome c reductase core protein I (UQCRC1), mRNA 1026 NM_003329 Homo sapiens thioredoxin (TXN), mRNA 1002 NM_001069 Homo sapiens tubulin, beta polypeptide (TUBB), mRNA 1013 NM_000356 Homo sapiens Treacher Collins-Franceschetti syndrome 1 (TCOF1), mRNA 673 NM_003134 Homo sapiens signal recognition particle 14 kDa (homologous Alu RNA binding protein) (SRP14), mRNA 2911 NM_003131 Homo sapiens serum response factor (c-fos serum response element-binding transcription factor) (SRF), mRNA 664 NM_000454 Homo sapiens superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) (SOD1), mRNA 1739 NM_003091 Homo sapiens small nuclear ribonucleoprotein polypeptides B and B1 (SNRPB), mRNA 1609 NM_003089 Homo sapiens small nuclear ribonucleoprotein 70 kDa polypeptide (RNP antigen) (SNRP70), mRNA 1672 NM_003016 Homo sapiens splicing factor, arginine/serine-rich 2 (SFRS2), mRNA 1476 NM_003952 Homo sapiens ribosomal protein S6 kinase, 70 kDa, polypeptide 2 (RPS6KB2), mRNA 476 NM_002950 Homo sapiens ribophorin I (RPN1), mRNA 495 NM_002743 Homo sapiens protein kinase C substrate 80 K-H (PRKCSH), mRNA 691 NM_002686 Homo sapiens phenylethanolamine N-methyltransferase (PNMT), mRNA 501 NM_002654 Homo sapiens pyruvate kinase, muscle (PKM2), mRNA 2474 NM_002648 Homo sapiens pim-1 oncogene (PIM1), mRNA 1052 NM_002635 Homo sapiens solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3 (SLC25A3), nuclear gene encoding mitochondrial protein, transcript variant 1b, mRNA 2310 NM_002494 Homo sapiens NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 1, 6 kDa (NDUFC1), mRNA 2247 NM_002488 Homo sapiens NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8 kDa (NDUFA2), mRNA 689 NM_002434 Homo sapiens N-methylpurine-DNA glycosylase (MPG), mRNA 1808 NM_002415 Homo sapiens macrophage migration inhibitory factor (glycosylation-inhibiting factor) (MIF), mRNA 3774 NM_002227 Homo sapiens Janus kinase 1 (a protein tyrosine kinase) (JAK1), mRNA 990 NM_001536 Homo sapiens HMT1 hnRNP methyltransferase-like 2 (S. cerevisiae) (HRMT1L2), mRNA 832 NM_000183 Homo sapiens hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl- Coenzyme A hydratase (trifunctional protein), beta subunit (HADHB), mRNA 1087 NM_002085 Homo sapiens glutathione peroxidase 4 (phospholipid hydroperoxidase) (GPX4), mRNA 1279 NM_001502 Homo sapiens glycoprotein 2 (zymogen granule membrane) (GP2), mRNA 572 NM_002080 Homo sapiens glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) (GOT2), nuclear gene encoding mitochondrial protein, mRNA 1000 NM_001440 Homo sapiens exostoses (multiple)-like 3 (EXTL3), mRNA 592 NM_003754 Homo sapiens eukaryotic translation initiation factor 3, subunit 5 epsilon, 47 kDa (EIF3S5), mRNA 1585 NM_003755 Homo sapiens eukaryotic translation initiation factor 3, subunit 4 delta, 44 kDa (EIF3S4), mRNA 1085 NM_003757 Homo sapiens eukaryotic translation initiation factor 3, subunit 2 beta, 36 kDa (EIF3S2), mRNA 622 NM_001360 Homo sapiens 7-dehydrocholesterol reductase (DHCR7), mRNA 653 NM_001344 Homo sapiens defender against cell death 1 (DAD1), mRNA 812 NM_001914 Homo sapiens cytochrome b-5 (CYB5), nuclear gene encoding mitochondrial protein, mRNA 667 NM_001834 Homo sapiens clathrin, light polypeptide (Lcb) (CLTB), transcript variant nonbrain, mRNA 447 NM_001833 Homo sapiens clathrin, light polypeptide (Lca) (CLTA), transcript variant nonbrain, mRNA 1315 NM_001281 Homo sapiens cytoskeleton-associated protein 1 (CKAP1), mRNA 1066 NM_001749 Homo sapiens calpain, small subunit 1 (CAPNS1), mRNA 1308 NM_001697 Homo sapiens ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) (ATP5O), mRNA 1155 NM_001689 Homo sapiens ATP synthase, H+ transporting, mitochondrial F0 complex, subunit c (subunit 9) isoform 3 (ATP5G3), mRNA 1152 NM_001675 Homo sapiens activating transcription factor 4 (tax- responsive enhancer element B67) (ATF4), mRNA 1392 NM_001642 Homo sapiens amyloid beta (A4) precursor-like protein 2 (APLP2), mRNA 1341 NM_014724 Homo sapiens zinc finger protein 305 (ZNF305), mRNA 719 NM_005494 Homo sapiens DnaJ (Hsp40) homolog, subfamily B, member 6 (DNAJB6), transcript variant 2, mRNA 923 NM_006597 Homo sapiens heat shock 70 kDa protein 8 (HSPA8), transcript variant 1, mRNA 1650 NM_006623 Homo sapiens phosphoglycerate dehydrogenase (PHGDH), mRNA 888 NM_015646 Homo sapiens RAP1B, member of RAS oncogene family (RAP1B), mRNA 736 NM_016532 Homo sapiens skeletal muscle and kidney enriched inositol phosphatase (SKIP), transcript variant 1, mRNA 1529 NM_015292 Homo sapiens likely ortholog of mouse membrane bound C2 domain containing protein (MBC2), mRNA 475 NM_005870 Homo sapiens sin3-associated polypeptide, 18 kDa (SAP18), mRNA 1651 NM_006833 Homo sapiens COP9 subunit 6 (MOV34 homolog, 34 kD) (COPS6), mRNA 1338 NM_005718 Homo sapiens actin related protein 2/3 complex, subunit 4, 20 kDa (ARPC4), mRNA 433 NM_005719 Homo sapiens actin related protein 2/3 complex, subunit 3, 21 kDa (ARPC3), mRNA 949 NM_006372 Homo sapiens NS1-associated protein 1 (NSAP1), mRNA 704 NM_005180 Homo sapiens B lymphoma Mo-MLV insertion region (mouse) (BMI1), mRNA 732 NM_018975 Homo sapiens telomeric repeat binding factor 2, interacting protein (TERF2IP), mRNA 684 NM_005731 Homo sapiens actin related protein 2/3 complex, subunit 2, 34 kDa (ARPC2), transcript variant 2, mRNA 2262 NM_020151 Homo sapiens START domain containing 7 (STARD7), transcript variant 1, mRNA 1562 NM_005103 Homo sapiens fasciculation and elongation protein zeta 1 (zygin I) (FEZ1), transcript variant 1, mRNA 672 NM_012179 Homo sapiens F-box only protein 7 (FBXO7), mRNA 865 NM_020360 Homo sapiens phospholipid scramblase 3 (PLSCR3), mRNA 3645 NM_014891 Homo sapiens PDGFA associated protein 1 (PDAP1), mRNA 1045 NM_005745 Homo sapiens accessory protein BAP31 (DX51357E), mRNA 1525 NM_005418 Homo sapiens suppression of tumorigenicity 5 (ST5), transcript variant 1, mRNA 2305 NM_006262 Homo sapiens peripherin (PRPH), mRNA 484 NM_133476 Homo sapiens zinc finger protein 384 (ZNF384), mRNA 448 NM_006570 Homo sapiens Ras-related GTP-binding protein (RAGA), mRNA 903 NM_006333 Homo sapiens nuclear DNA-binding protein (CID), mRNA 740 NM_007285 Homo sapiens GABA(A) receptor-associated protein-like 2 (GABARAPL2), mRNA 1265 NM_006354 Homo sapiens transcriptional adaptor 3-like (TADA3L) transcript variant 1, mRNA 1060 NM_014302 Homo sapiens Sec61 gamma (SEC61G), mRNA 644 NM_006118 Homo sapiens HS1 binding protein (HAX1), mRNA 1493 NM_012100 Homo sapiens aspartyl aminopeptidase (DNPEP), mRNA 419 NM_015680 Homo sapiens hypothetical protein CGI-57 (CGI-57), mRNA 976 NM_030796 Homo sapiens hypothetical protein DKFZp564K0822 (DKFZP564K0822), mRNA 572 NM_024069 Homo sapiens hypothetical protein MGC2749 (MGC2749), mRNA 573 NM_013234 Homo sapiens muscle specific gene (M9), mRNA 1056 NM_013310 Homo sapiens hypothetical protein AF038169 (AF038169), mRNA 1319 NM_018507 Homo sapiens hypothetical protein PRO1843 (PRO1843), mRNA 1081 NM_017670 Homo sapiens hypothetical protein FLJ20113 (FLJ20113), mRNA 999 NM_016292 Homo sapiens heat shock protein 75 (TRAP1), mRNA 615 NM_014916 Homo sapiens KIAA1079 protein (KIAA1079), mRNA 772 NM_014696 Homo sapiens KIAA0514 gene product (KIAA0514), mRNA 746 NM_014630 Homo sapiens KIAA0211 gene product (KIAA0211), mRNA 750 NM_014761 Homo sapiens KIAA0174 gene product (KIAA0174), mRNA 679 NM_007286 Homo sapiens synaptopodin (KIAA1029), mRNA 1349 NM_007263 Homo sapiens coatomer protein complex, subunit epsilon (COPE), mRNA 1272 NM_006349 Homo sapiens putative cyclin G1 interacting protein (CG1I), mRNA 591 NM_006004 Homo sapiens ubiquinol-cytochrome c reductase hinge protein (UQCRH), mRNA 1285 NM_005787 Homo sapiens Not56 (D. melanogaster)-like protein (NOT56L), mRNA 458
NM_012412 Homo sapiens histone H2A.F/Z variant (H2AV), transcript variant 1, mRNA 1070 NM_012401 Homo sapiens plexin B2 (PLXNB2), mRNA 929 NM_007262 Homo sapiens RNA-binding protein regulatory subunit (DJ-1), mRNA 1514 NM_007273 Homo sapiens repressor of estrogen receptor activity (REA), mRNA 1804 NM_021009 Homo sapiens ubiquitin C (UBC), mRNA 8892 NM_023009 Homo sapiens macrophage myristoylated alanine-rich C kinase substrate (MACMARCKS), mRNA 2025 NM_015456 Homo sapiens cofactor of BRCA1 (COBRA1), mRNA 848 NM_005053 Homo sapiens RAD23 homolog A (S. cerevisiae) (RAD23A), mRNA 1015 NM_006830 Homo sapiens ubiquinol-cytochrome c reductase (6.4 kD) subunit (UQCR), mRNA 1489 NM_005682 Homo sapiens G protein-coupled receptor 56 (GPR56), mRNA 742 NM_012102 Homo sapiens arginine-glutamic acid dipeptide (RE) repeats (RERE), mRNA 940 NM_005550 Homo sapiens kinesin family member C3 (KIFC3), mRNA 603 NM_021960 Homo sapiens myeloid cell leukemia sequence 1 (BCL2- related) (MCL1), mRNA 961 NM_021959 Homo sapiens protein phosphatase 1, regulatory (inhibitor) subunit 11 (PPP1R11), mRNA 462 NM_014730 Homo sapiens KIAA0152 gene product (KIAA0152), mRNA 713 NM_014402 Homo sapiens low molecular mass ubiquinone-binding protein (9.5 kD) (QP-C), mRNA 2805 NM_007067 Homo sapiens histone acetyltransferase (HBOA), mRNA 510 NM_006086 Homo sapiens tubulin, beta, 4 (TUBB4), mRNA 2239 NM_014972 Homo sapiens KIAA1049 protein (KIAA1049), mRNA 1011 NM_024092 Homo sapiens hypothetical protein MGC5508 (MGC5508), mRNA 749 NM_021983 Homo sapiens major histocompatibility complex, class II, DR beta 4 (HLA-DRB4), mRNA 1095 NM_006510 Homo sapiens ret finger protein (RFP), transcript variant alpha, mRNA 420 NM_006711 Homo sapiens RNA binding protein Si, serine-rich domain (RNPS1), transcript variant 1, mRNA 1376 NM_006145 Homo sapiens DnaJ (Hsp40) homolog, subfmaily B, member 1 (DNAJB1), mRNA 667 NM_033142 Homo sapiens chorionic gonadotropin, beta polypeptide 7 (CGB7), mRNA 773 NM_006351 Homo sapiens translocase of inner mitochondrial membrane 44 homolog (yeast) (TIM1V144), mRNA 717 NM_014281 Homo sapiens fuse-binding protein-interacting repressor (SIABBP1), transcript variant 2, mRNA 632 NM_012106 Homo sapiens binder of Arl Two (BART1), mRNA 760 NM_021975 Homo sapiens v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) (RELA), mRNA 767 NM_014874 Homo sapiens mitofusin 2 (MFN2), mRNA 453 NM_006796 Homo sapiens AFG3 ATPase family gene 3-like 2 (yeast) (AFG3L2), nuclear gene encoding mitochondrial protein, mRNA 724 NM_006666 Homo sapiens RuvB-like 2 (E. coli) (RUVBL2), mRNA 690 NM_005219 Homo sapiens diaphanous homolog 1 (Drosophila) (DIAPH1), mRNA 592 NM_033546 Homo sapiens myosin regulatory light chain (MLC-B), mRNA 1231 NM_032348 Homo sapiens hypothetical protein MGC3047 (MGC3047), mRNA 553 NM_024798 Homo sapiens hypothetical protein FLJ13952 (FLJ13952), mRNA 2142 NM_021103 Homo sapiens thymosin, beta 10 (TMSB10), mRNA 1890 NM_020195 Homo sapiens HCDI protein (HCDI), mRNA 612 NM_017432 Homo sapiens prostate tumor over expressed gene 1 (PTOV1), mRNA 1509 NM_014901 Homo sapiens KIAA1100 protein (KIAA1100), mRNA 1999 NM_014694 Homo sapiens KIAA0605 gene product (KIAA0605), mRNA 532 NM_007369 Homo sapiens G-protein coupled receptor (RE2), mRNA 963 NM_006156 Homo sapiens neural precursor cell expressed, developmentally down-regulated 8 (NEDD8), mRNA 867 NM_006429 Homo sapiens chaperonin containing TCP1, subunit 7 (eta) (CCT7), mRNA 1434 NM_006513 Homo sapiens seryl-tRNA synthetase (SARS), mRNA 1130 NM_005022 Homo sapiens profilin 1 (PFN1), mRNA 2793 NM_014654 Homo sapiens syndecan 3 (N-syndecan) (SDC3), mRNA 1082 NM_007209 Homo sapiens ribosomal protein L35 (RPL35), mRNA 3463 NM_006082 Homo sapiens tubulin, alpha, ubiquitous (K-ALPHA-1), mRNA 4261 NM_006362 Homo sapiens nuclear RNA export factor 1 (NXF1), mRNA 729 NM_014228 Homo sapiens solute carrier family 6 (neurotransmitter transporter, L-proline), member 7 (SLC6A7), mRNA 1037 NM_006411 Homo sapiens 1-acylglycerol-3-phosphate O- acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) (AGPAT1), mRNA 742 NM_021134 Homo sapiens mitochondrial ribosomal protein L23 (MRPL23), mRNA 771 NM_021974 Homo sapiens polymerase (RNA) II (DNA directed) polypeptide F (POLR2F), mRNA 1169 NM_006808 Homo sapiens protein translocation complex beta (SEC61B), mRNA 679 NM_005617 Homo sapiens ribosomal protein S14 (RPS14), mRNA 7764 NM_005520 Homo sapiens heterogeneous nuclear ribonucleoprotein H1 (H) (HNRPH1), mRNA 2234 NM_006755 Homo sapiens transaldolase 1 (TALDO1), mRNA 874 NM_006010 Homo sapiens arginine-rich, mutated in early stage tumors (ARMET), mRNA 617 NM_005088 Homo sapiens DNA segment on chromosome X and Y (unique) 155 expressed sequence (DXYS155E), mRNA 524 NM_014754 Homo sapiens phosphatidylserine synthase 1 (PTDSS1), mRNA 930 NM_021953 Homo sapiens forkhead box M1 (FOXM1), mRNA 567 NM_006908 Homo sapiens ras-related C3 botulinum toxin substrate 1 (rho family, small GTP binding protein Rac1) (RAC1), transcript variant Rac1, mRNA 1978 NM_014231 Homo sapiens vesicle-associated membrane protein 1 (synaptobrevin 1) (VAMP1), transcript variant VAMP- 1A, mRNA 745 NM_014833 Homo sapiens KIAA0618 gene product (KIAA0618), mRNA 607 NM_005157 Homo sapiens v-abl Abelson murine leukemia viral oncogene homolog 1 (ABL1), transcript variant a, mRNA 777 NM_006325 Homo sapiens RAN, member RAS oncogene family (RAN), mRNA 3100 NM_007245 Homo sapiens ataxin 2 related protein (A2LP), transcript variant 1, mRNA 564 NM_007008 Homo sapiens reticulon 4 (RTN4), mRNA 1267 NM_006782 Homo sapiens zinc finger protein-like 1 (ZFPL1), mRNA 712 NM_006694 Homo sapiens jumping translocation breakpoint (JTB), mRNA 2394 NM_006703 Homo sapiens nudix (nucleoside diphosphate linked moiety X)-type motif 3 (NUDT3), mRNA 566 NM_006032 Homo sapiens copine VI (neuronal) (CPNE6), mRNA 1540 NM_012227 Homo sapiens Pseudoautosomal GTP-binding protein- like (PGPL), mRNA 494 NM_014604 Homo sapiens Tax interaction protein 1 (TIP-1), mRNA 748 NM_021642 Homo sapiens Fc fragment of IgG, low affinity Ha, receptor for (CD32) (FCGR2A), mRNA 772 NM_005354 Homo sapiens jun D proto-oncogene (JUND), mRNA 4967 NM_020529 Homo sapiens nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha (NFKBIA), mRNA 922 NM_005561 Homo sapiens lysosomal-associated membrane protein 1 (LAMP1), mRNA 2865 NM_014774 Homo sapiens KIAA0494 gene product (KIAA0494), mRNA 604 NM_014390 Homo sapiens EBNA-2 co-activator (100 kD) (p100), mRNA 867 NM_014623 Homo sapiens male-enhanced antigen (MEA), mRNA 548 NM_014453 Homo sapiens putative breast adenocarcinoma marker (32 kD) (BC-2), mRNA 597 NM_012127 Homo sapiens Cip1-interacting zinc finger protein (CIZ1), mRNA 1051 NM_012099 Homo sapiens CD3-epsilon-associated protein; antisense to ERCC-1 (ASE-1), mRNA 472 NM_006888 Homo sapiens calmodulin 1 (phosphorylase kinase, delta) (CALM1), mRNA 2161 NM_006867 Homo sapiens RNA-binding protein gene with multiple splicing (RBPMS), mRNA 1255 NM_006743 Homo sapiens RNA binding motif protein 3 (RBM3), mRNA 2949 NM_006688 Homo sapiens C1q-related factor (CRF), mRNA 3604 NM_006295 Homo sapiens valyl-tRNA synthetase 2 (VARS2), mRNA 721 NM_006283 Homo sapiens transforming, acidic coiled-coil containing protein 1 (TACC1), mRNA 867 NM_006221 Homo sapiens protein (peptidyl-prolyl cis/trans isomerase) NIMA-interacting 1 (PIN1), mRNA 561 NM_006148 Homo sapiens LIM_and SH3 protein 1 (LASP1), mRNA 1067 NM_005954 Homo sapiens metallothionein 3 (growth inhibitory factor (neurotrophic)) (MT3), mRNA 2311 NM_006003 Homo sapiens ubiquinol-cytochrome c reductase, Rieske iron-sulfur polypeptide 1 (UQCRFS1), nuclear gene encoding mitochondrial protein, mRNA 951 NM_005998 Homo sapiens chaperonin containing TCP1, subunit 3 (gamma) (CCT3), mRNA 1098 NM_005997 Homo sapiens transcription factor-like 1 (TCFL1), mRNA 505 NM_005629 Homo sapiens solute carrier family 6 (neurotransmitter transporter, creatine), member 8 (SLC6A8), mRNA 628 NM_005548 Homo sapiens lysyl-tRNA synthetase (KARS), mRNA 1193 NM_005545 Homo sapiens immunoglobulin superfamily containing leucine-rich repeat (ISLR), mRNA 796 NM_005507 Homo sapiens cofilin 1 (non-muscle) (CFL1), mRNA 5155 NM_005381 Homo sapiens nucleolin (NCL), mRNA 2043 NM_005439 Homo sapiens myeloid leukemia factor 2 (MLF2), mRNA 697 NM_006445 Homo sapiens PRP8 pre-mRNA processing factor 8 homolog (yeast) (PRPF8), mRNA 1384 NM_019059 Homo sapiens homolog of Tom7 (S. cerevisiae) (TOM7), mRNA 2364 NM_006039 Homo sapiens mannose receptor, C type 2 (MRC2), mRNA 564 NM_006066 Homo sapiens aldo-keto reductase family 1, member A1 (aldehyde reductase) (AKR1A1), transcript variant 1, mRNA 879 NM_013318 Homo sapiens hypothetical protein LQFBS-1 (LQFBS-1), mRNA 485 NM_006098 Homo sapiens guanine nucleotide binding protein (G protein), beta polypeptide 2-like 1 (GNB2L1), mRNA 7556 NM_007359 Homo sapiens M1LN51 protein (M1LN51), mRNA 975 NM_017510 Homo sapiens gp25L2 protein (HSGP25L2G), mRNA 1032 NM_015399 Homo sapiens breast cancer metastasis-suppressor 1 (BRMS1), mRNA 476 NM_014508 Homo sapiens apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3C (APOBEC3C), mRNA 816 NM_018955 Homo sapiens ubiquitin B (UBB), mRNA 6074 NM_006368 Homo sapiens cAMP responsive element binding protein 3 (luman) (CREB3), mRNA 533 NM_015024 Homo sapiens RAN binding protein 16 (RANBP16), mRNA 559 NM_031420 Homo sapiens mitochondrial ribosomal protein L9 (MRPL9), nuclear gene encoding mitochondrial protein, mRNA 476 NM_013232 Homo sapiens programmed cell death 6 (PDCD6), mRNA 816 NM_005917 Homo sapiens malate dehydrogenase 1, NAD (soluble) (MDH1), mRNA 1724 NM_032801 Homo sapiens junctional adhesion molecule 3 (JAM3), mRNA 624 NM_030662 Homo sapiens mitogen-activated protein kinase kinase 2 (MAP2K2), mRNA 663 NM_006268 Homo sapiens requiem, apoptosis response zinc finger gene (REQ), mRNA 784 NM_006826 Homo sapiens tyrosine 3-monooxygenase/tryptophan 5- monooxygenase activation protein, theta polypeptide (YWHAQ), mRNA 1917 NM_144582 Homo sapiens hypothetical protein MGC32043 (MGC32043), mRNA 712 NM_144565 Homo sapiens hypothetical protein FLJ32334 (FLJ32334), mRNA 5319
NM_005984 Homo sapiens solute carrier family 25 (mitochondrial carrier; citrate transporter), member 1 (SLC25A1), mRNA 620 NM_017828 Homo sapiens hypothetical protein FLJ20452 (FLJ20452), mRNA 546 NM_020150 Homo sapiens SARI protein (SARI), mRNA 581 NM_014610 Homo sapiens alpha glucosidase II alpha subunit (G2AN), mRNA 1293 NM_014225 Homo sapiens protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), alpha isoform (PPP2R1A), mRNA 1687 NM_006937 Homo sapiens SMT3 suppressor of mif two 3 homolog 2 (yeast) (SMT3H2), mRNA 2940 NM_005726 Homo sapiens Ts translation elongation factor, mitochondrial (TSFM), mRNA 408 NM_005347 Homo sapiens heat shock 70 kDa protein 5 (glucose- regulated protein, 78 kDa) (HSPA5), mRNA 1111 NM_014255 Homo sapiens transmembrane protein 4 (TMEM4), mRNA 505 NM_006815 Homo sapiens coated vesicle membrane protein (RNP24), mRNA 1642 NM_005456 Homo sapiens mitogen-activated protein kinase 8 interacting protein 1 (MAPK8IP1), mRNA 885 NM_005273 Homo sapiens guanine nucleotide binding protein (G protein), beta polypeptide 2 (GNB2), mRNA 1478 NM_007260 Homo sapiens lysophospholipase II (LYPLA2), mRNA 2000 NM_007103 Homo sapiens NADH dehydrogenase (ubiquinone) flavoprotein 1, 51 kDa (NDUFV1), mRNA 803 NM_017797 Homo sapiens BTB (POZ) domain containing 2 (BTBD2), mRNA 1030 NM_016237 Homo sapiens anaphase promoting complex subunit 5 (ANAPC5), mRNA 935 NM_005801 Homo sapiens putative translation initiation factor (SUI1), mRNA 1058 NM_005216 Homo sapiens dolichyl-diphosphooligosaccharide-protein glycosyltransferase (DDOST), mRNA 1421 NM_016457 Homo sapiens protein kinase D2 (PKD2), mRNA 662 NM_006442 Homo sapiens DR1-associated protein 1 (negative cofactor 2 alpha) (DRAP1), mRNA 561 NM_021019 Homo sapiens myosin, light polypeptide 6, alkali, smooth muscle and non-muscle (MYL6), transcript variant 1, mRNA 5933 NM_005112 Homo sapiens WD repeat domain 1 (WDR1), transcript variant 2, mRNA 809 NM_005370 Homo sapiens mel transforming oncogene (derived from cell line NK14)-RAB8 homolog (MEL), mRNA 682 NM_006289 Homo sapiens talin 1 (TLN1), mRNA 1811 NM_005698 Homo sapiens secretory carrier membrane protein 3 (SCAMP3), transcript variant 1, mRNA 602 NM_ 024011 Homo sapiens cell division cycle 2-like 2 (CDC2L2), transcript variant 1, mRNA 430 NM_005105 Homo sapiens RNA binding motif protein 8A (RBM8A), mRNA 785 NM_ 006013 Homo sapiens ribosomal protein L10 (RPL10), mRNA 10721 NM_005786 Homo sapiens serologically defined colon cancer antigen 33 (SDCCAG33), mRNA 1435 NM_007104 Homo sapiens ribosomal protein L10a (RPL10A), mRNA 3006 NM_005762 Homo sapiens tripartite motif-containing 28 (TRIM28), mRNA 1473 NM_012138 Homo sapiens apoptosis antagonizing transcription factor (AATF), mRNA 504 NM_015318 Homo sapiens Rho-specific guanine nucleotide exchange factor p114 (P114-RHO-GEF), mRNA 405 NM_012423 Homo sapiens ribosomal protein L13a (RPL13A), mRNA 9545 NM_021128 Homo sapiens polymerase (RNA) II (DNA directed) polypeptide L, 7.6 kDa (POLR2L), mRNA 1524 NM_032635 Homo sapiens seven transmembrane domain protein (NIFIE14), mRNA 1121 NM_005080 Homo sapiens X-box binding protein 1 (XBP1), mRNA 1772 NM_006389 Homo sapiens hypoxia up-regulated 1 (HYOU1), mRNA 1076 NM_024112 Homo sapiens chromosome 9 open reading frame 16 (C9orf16), mRNA 598 NM_006817 Homo sapiens chromosome 12 open reading frame 8 (C12orf8), mRNA 910 NM_022830 Homo sapiens hypothetical protein FLJ22347 (FLJ22347), mRNA 6021 NM_019884 Homo sapiens glycogen synthase kinase 3 alpha (GSK3A), mRNA 963 NM_021107 Homo sapiens mitochondrial ribosomal protein S12 (MRPS12), nuclear gene encoding mitochondrial protein, transcript variant 1, mRNA 828 NM_021074 Homo sapiens NADH dehydrogenase (ubiquinone) flavoprotein 2, 24 kDa (NDUFV2), mRNA 539 NM_014944 Homo sapiens calsyntenin 1 (CLSTN1), mRNA 1620 NM_014764 Homo sapiens DAZ associated protein 2 (DAZAP2), mRNA 1802 NM_ 015343 Homo sapiens likely ortholog of Xenopus dullard (HSA011916), mRNA 933 NM_014420 Homo sapiens dickkopf homolog 4 (Xenopus laevis) (DKK4), mRNA 426 NM_005884 Homo sapiens p21(CDKN1A)-activated kinase 4 (PAK4), mRNA 375 NM_012407 Homo sapiens protein kinase C, alpha binding protein (PRKCABP), mRNA 553 NM_012111 Homo sapiens chromosome 14 open reading frame 3 (C14orf3), mRNA 717 NM_007144 Homo sapiens zinc finger protein 144 (Mel-18) (ZNF144), mRNA 798 NM_007108 Homo sapiens transcription elongation factor B (SIII), polypeptide 2 (18 kDa, elongin B) (TCEB2), mRNA 1192 NM_007278 Homo sapiens GABA(A) receptor-associated protein (GABARAP), mRNA 2335 NM_007100 Homo sapiens ATP synthase, H+ transporting, mitochondrial F0 complex, subunit e (ATP5I), mRNA 2623 NM_006936 Homo sapiens SMT3 suppressor of mif two 3 homolog 1 (yeast) (SMT3H1), mRNA 1187 NM_006899 Homo sapiens isocitrate dehydrogenase 3 (NAD+) beta (IDH3B), mRNA 565 NM_006801 Homo sapiens KDEL (Lys-Asp-Glu-Leu) (SEQ ID NO: 1) endoplasmic reticulum protein retention receptor 1 (KDELR1), mRNA 1270 NM_006612 Homo sapiens kinesin family member 1C (KIF1C), mRNA 600 NM_006659 Homo sapiens tubulin, gamma complex associated protein 2 (TUBGCP2), mRNA 701 NM_006595 Homo sapiens apoptosis inhibitor 5 (API5), mRNA 1173 NM_006401 Homo sapiens acidic (leucine-rich) nuclear phosphoprotein 32 family, member B (ANP32B), mRNA 2354 NM_006423 Homo sapiens Rab acceptor 1 (prenylated) (RABAC1), mRNA 1706 NM_006460 Homo sapiens HMBA-inducible (HIS1), mRNA 369 NM_006356 Homo sapiens ATP synthase, H+ transporting, mitochondrial F0 complex, subunit d (ATP5H), mRNA 1047 NM_005891 Homo sapiens acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) (ACAT2), mRNA 911 NM_005839 Homo sapiens serine/arginine repetitive matrix 1 (SRRM1), mRNA 745 NM_005594 Homo sapiens nascent-polypeptide-associated complex alpha polypeptide (NACA), mRNA 3678 NM_005340 Homo sapiens histidine triad nucleotide binding protein 1 (HINT1), mRNA 2270 NM_005175 Homo sapiens ATP synthase, H+ transporting, mitochondrial F0 complex, subunit c (subunit 9), isoform 1 (ATP5G1), mRNA 1266 NM_005165 Homo sapiens aldolase C, fructose-bisphosphate (ALDOC), mRNA 1052 NM_005001 Homo sapiens NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 7, 14.5 kDa (NDUFA7), mRNA 806 NM_001402 NM_001958 NM_001961 NM_002714 NM_002808 NM_002809 NM_004898 NM_007182 NM_032378
[0252] In various embodiments of the present invention, the reference value is chromosome 15 centromere copy number. In various embodiments, the reference value for chromosome 15q26 copy number is chromosome 15 centromere copy number.
[0253] The chromosome 15q26 copy number and chromosome 15 centromere copy number can be ascertained by various methods. For example, they can be ascertained through chip based measurements with or without a normal reference, or by using centromeric FISH probes, which is an assay for testing for copy number changes using microscopy.
[0254] In various embodiments, the number of chromosome 15 centromere probes is compared to the number of 15q26.1 probes, in each cell using a microscope. If the numbers match, there is no relative gain of 15q26. Increase in this context can be a numerical increase, e.g., 2-3 copies. In various embodiments, copy gain can be defined by the absolute copy number determined in an interphase FISH assay averaged by counting a minimum of 20 tumor cells. In various embodiments, copy gain can be defined as the ratio of 15q26/centromere 15 determined in an interphase FISH assay counting both spots (15q26 and cent15) in the same cells and averaging over a minimum of 20 cells. In various embodiments, copy gain can be determined using a normalized genome wide assay such as SNP array, genome sequencing and the like, wherein the normalization is done using the ASCAT algorithm or other appropriate algorithms. In various embodiments, the cutpoints can be anything above normal, which is 2 absolute copies of 15q26, or ratio >1 for 15q26/cent15. Due to the typical noise in these assays, in certain embodiments, the cutoff is defined by adding a standard error. Accordingly, copy >2.6 or ratio >1.3 signify copy number gain.
[0255] Thus, in various embodiments, a copy number gain of over 2.6 or a ratio of over 1.3 indicates a copy number gain in the sample.
[0256] In other embodiments, the reference value for chromosome 15q26 is determined from a non-cancer cell sample from the subject or a member of the same species to which the subject belongs. In certain embodiments, the reference value is determined from a non-cancerous cell or tissue sample that is the same type of cell or tissue as the cancer cell from the subject. In certain embodiments, the reference value is determined from a non-cancerous cell or tissue sample that is not the same type of cell or tissue as the cancer cell from the subject. In various embodiments, array-based or sequencing-based technologies can be used wherein the reference can be from patients' normal cells (e.g., blood), or it can be a collection of blood samples.
[0257] Copy number abnormalities can be detected using methods, such as, for example, array CGH using BAC, cDNA and/or oligonucleotide arrays; microsatellite markers; STRs, RFLPS; etc.
[0258] Additional methods for evaluating copy number of nucleic acid in a sample include, but are not limited to, hybridization-based assays. One method for evaluating the copy number of encoding nucleic acid in a sample involves a Southern Blot. In a Southern Blot, the genomic DNA (typically fragmented and separated on an electrophoretic gel) is hybridized to a probe specific for the target region. Comparison of the intensity of the hybridization signal from the probe for the target region with control probe signal from analysis of normal genomic DNA (e.g., a non-amplified portion of the same or related cell, tissue, organ, etc.) provides an estimate of the relative copy number of the target nucleic acid. Alternatively, a Northern blot may be utilized for evaluating the copy number of encoding nucleic acid in a sample. In a Northern blot, mRNA is hybridized to a probe specific for the target region. Comparison of the intensity of the hybridization signal from the probe for the target region with control probe signal from analysis of normal mRNA (e.g., a non-amplified portion of the same or related cell, tissue, organ, etc.) provides an estimate of the relative copy number of the target nucleic acid. Similar methods for determining copy number can be performed using transcriptional arrays, which are well-known in the art.
[0259] An alternative means for determining the copy number is in situ hybridization (e.g., Angerer (1987) Meth. Enzymol 152: 649). Generally, in situ hybridization comprises the following steps: (1) fixation of tissue or biological structure to be analyzed; (2) prehybridization treatment of the biological structure to increase accessibility of target DNA, and to reduce nonspecific binding; (3) hybridization of the mixture of nucleic acids to the nucleic acid in the biological structure or tissue; (4) post-hybridization washes to remove nucleic acid fragments not bound in the hybridization and (5) detection of the hybridized nucleic acid fragments. The reagent used in each of these steps and the conditions for use vary depending on the particular application.
[0260] Preferred hybridization-based assays include, but are not limited to, traditional "direct probe" methods such as Southern blots or in situ hybridization (e.g., FISH and FISH plus SKY), and "comparative probe" methods such as comparative genomic hybridization (CGH), e.g., cDNA-based or oligonucleotide-based CGH. The methods can be used in a wide variety of formats including, but not limited to, substrate (e.g. membrane or glass) bound methods or array-based approaches.
[0261] In a typical in situ hybridization assay, cells are fixed to a solid support, typically a glass slide. If a nucleic acid is to be probed, the cells are typically denatured with heat or alkali. The cells are then contacted with a hybridization solution at a moderate temperature to permit annealing of labeled probes specific to the nucleic acid sequence encoding the protein. The targets (e.g., cells) are then typically washed at a predetermined stringency or at an increasing stringency until an appropriate signal to noise ratio is obtained.
[0262] The probes are typically labeled, e.g., with radioisotopes or fluorescent reporters. Preferred probes are sufficiently long so as to specifically hybridize with the target nucleic acid(s) under stringent conditions. The preferred size range is from about 200 bases to about 1000 bases.
[0263] In some applications it is necessary to block the hybridization capacity of repetitive sequences. Thus, in some embodiments, tRNA, human genomic DNA, or Cot-I DNA is used to block non-specific hybridization.
[0264] In CGH methods, a first collection of nucleic acids (e.g., from a sample, e.g., a possible tumor) is labeled with a first label, while a second collection of nucleic acids (e.g., a control, e.g., from a healthy cell/tissue) is labeled with a second label. The ratio of hybridization of the nucleic acids is determined by the ratio of the two (first and second) labels binding to each fiber in the array. Where there are chromosomal deletions or multiplications, differences in the ratio of the signals from the two labels will be detected and the ratio will provide a measure of the copy number. Array-based CGH may also be performed with single-color labeling (as opposed to labeling the control and the possible tumor sample with two different dyes and mixing them prior to hybridization, which will yield a ratio due to competitive hybridization of probes on the arrays). In single color CGH, the control is labeled and hybridized to one array and absolute signals are read, and the possible tumor sample is labeled and hybridized to a second array (with identical content) and absolute signals are read. Copy number difference is calculated based on absolute signals from the two arrays. Hybridization protocols suitable for use with the methods of the invention are described, e.g., in Albertson (1984) EMBO J. 3: 1227-1234; Pinkel (1988) Proc. Natl. Acad. Sci. USA 85: 9138-9142; EPO Pub. No. 430,402; Methods in Molecular Biology, Vol. 33: In situ Hybridization Protocols, Choo, ed., Humana Press, Totowa, N.J. (1994), etc. In one embodiment, the hybridization protocol of Pinkel, et al. (1998) Nature Genetics 20: 207-211, or of Kallioniemi (1992) Proc. Natl Acad Sci USA 89:5321-5325 (1992) is used.
[0265] The methods of the invention are particularly well suited to array-based hybridization formats. Array-based CGH is described in U.S. Pat. No. 6,455,258, the contents of which are incorporated herein by reference. In still another embodiment, amplification-based assays can be used to measure copy number. In such amplification-based assays, the nucleic acid sequences act as a template in an amplification reaction (e.g., Polymerase Chain Reaction (PCR). In a quantitative amplification, the amount of amplification product will be proportional to the amount of template in the original sample. Comparison to appropriate controls, e.g. healthy tissue, provides a measure of the copy number.
[0266] Methods of "quantitative" amplification are well known to those of skill in the art. For example, quantitative PCR involves simultaneously co-amplifying a known quantity of a control sequence using the same primers. This provides an internal standard that may be used to calibrate the PCR reaction. Detailed protocols for quantitative PCR are provided in Innis, et al. (1990) PCR Protocols, A Guide to Methods and Applications, Academic Press, Inc. N.Y.). Measurement of DNA copy number at microsatellite loci using quantitative PCR anlaysis is described in Ginzonger, et al. (2000) Cancer Research 60:5405-5409. The known nucleic acid sequence for the genes is sufficient to enable one of skill in the art to routinely select primers to amplify any portion of the gene. Fluorogenic quantitative PCR may also be used in the methods of the invention. In fluorogenic quantitative PCR, quantitation is based on amount of fluorescence signals, e.g., TaqMan and sybr green.
[0267] Other suitable amplification methods include, but are not limited to, ligase chain reaction (LCR) (see Wu and Wallace (1989) Genomics 4: 560, Landegren, et al. (1988) Science 241:1077, and Barringer et al. (1990) Gene 89: 117), transcription amplification (Kwoh, et al. (1989) Proc. Natl. Acad. Sci. USA 86: 1173), self-sustained sequence replication (Guatelli, et al. (1990) Proc. Nat. Acad. Sci. USA 87: 1874), dot PCR, and linker adapter PCR, etc.
[0268] In still other embodiments of the methods provided herein, sequencing of individual nucleic molecules (or their amplification products) is performed, as an alternative to hybridization-based assays, using nucleic acid sequencing techniques. In one embodiment, a high throughput parallel sequencing technique that isolates single nucleic acid molecules of a population of nucleic acid molecules prior to sequencing may be used. Such strategies may use so-called "next generation sequencing systems" including, without limitation, sequencing machines and/or strategies well known in the art, such as those developed by Illumina/Solexa (the Genome Analyzer; Bennett et al. (2005) Pharmacogenomics, 6:373-20 382), by Applied Biosystems, Inc. (the SOLiD Sequencer; solid.appliedbiosystems.com), by Roche (e.g., the 454 GS FLX sequencer; Margulies et al. (2005) Nature, 437:376-380; U.S. Pat. Nos. 6,274,320; 6,258,568; 6,210,891), by Heliscope.TM. system from Helicos Biosciences (see, e.g., U.S. Patent App. Pub. No. 2007/0070349), and by others. Other sequencing strategies such as stochastic sequencing (e.g., as developed by Oxford Nanopore) may also be used, e.g., as described in International Application No. PCT/GB2009/001690 (pub. no. WO/2010/004273). All of the copy number determining strategies described herein can similarly be applied to any of other nucleic acid-based analysis described herein, such as for BLM, FANCI, or 15q26 the like described further below.
Therapies
[0269] These therapies can be selected, used, administered, etc., in accordance with various embodiment of the present invention.
[0270] Platinum-comprising therapy, include but are not limited platinum chemotherapeutic agents, such as cisplatin, carboplatin, oxaliplatin, nedaplatin, and iproplatin. Other antineoplastic platinum coordination compounds are well known in the art, can be modified according to well-known methods in the art, and include the compounds disclosed in U.S. Pat. Nos. 4,996,337, 4,946,954, 5,091,521, 5,434,256, 5,527,905, and 5,633,243, all of which are incorporated herein by reference. In various embodiments described herein, the platinum comprising cancer therapy comprises cisplatinum or cis-diamminedichloroplatinum, phenanthriplatin, carboplatin, oxaliplatin, or a platinum complex that is activated by ultraviolet A light.
[0271] In various embodiments, non-platinum comprising therapies include, non-platinum chemotherapy. Non-platinum chemotherapy may be, but is not limited to, those selected from among the following groups of compounds: cytotoxic antibiotics, antimetabolities, anti-mitotic agents, alkylating agents, arsenic compounds, DNA topoisomerase inhibitors, taxanes, nucleoside analogues, plant alkaloids, and toxins; and synthetic derivatives thereof. Exemplary compounds include, but are not limited to, alkylating agents: treosulfan, and trofosfamide; plant alkaloids: vinblastine, paclitaxel, docetaxol; DNA topoisomerase inhibitors: doxorubicin, epirubicin, etoposide, camptothecin, topotecan, irinotecan, teniposide, crisnatol, and mitomycin; anti-folates: methotrexate, mycophenolic acid, and hydroxyurea; pyrimidine analogs: 5-fluorouracil, doxifluridine, and cytosine arabinoside; purine analogs: mercaptopurine and thioguanine; DNA antimetabolites: 2'-deoxy-5-fluorouridine, aphidicolin glycinate, and pyrazoloimidazole; and antimitotic agents: halichondrin, colchicine, and rhizoxin. Compositions comprising one or more chemotherapeutic agents (e.g., FLAG, CHOP) may also be used. FLAG comprises fludarabine, cytosine arabinoside (Ara-C) and G-CSF. CHOP comprises cyclophosphamide, vincristine, doxorubicin, and prednisone. In another embodiments, PARP (e.g., PARP-1 and/or PARP-2) inhibitors are used and such inhibitors are well known in the art (e.g., Olaparib, ABT-888, BSI-201, BGP-15 (N-Gene Research Laboratories, Inc.); INO-1001 (Inotek Pharmaceuticals Inc.); PJ34 (Soriano et al., 2001; Pacher et al., 2002b); 3-aminobenzamide (Trevigen); 4-amino-1,8-naphthalimide; (Trevigen); 6(5H)-phenanthridinone (Trevigen); benzamide (U.S. Pat. Re. 36,397); and NU1025 (Bowman et al.). The foregoing examples of non-platinum chemotherapeutic agents are illustrative, and are not intended to be limiting.
[0272] In various embodiments, non-platinum comprising therapies include, for example, radiation therapy. The radiation used in radiation therapy can be ionizing radiation. Radiation therapy can also be gamma rays, X-rays, or proton beams. Examples of radiation therapy include, but are not limited to, external-beam radiation therapy, interstitial implantation of radioisotopes (I-125, palladium, iridium), radioisotopes such as strontium-89, thoracic radiation therapy, intraperitoneal P-32 radiation therapy, and/or total abdominal and pelvic radiation therapy. For a general overview of radiation therapy, see Hellman, Chapter 16: Principles of Cancer Management: Radiation Therapy, 6th edition, 2001, DeVita et al., eds., J. B. Lippencott Company, Philadelphia. The radiation therapy can be administered as external beam radiation or teletherapy wherein the radiation is directed from a remote source. The radiation treatment can also be administered as internal therapy or brachytherapy wherein a radioactive source is placed inside the body close to cancer cells or a tumor mass. Also encompassed is the use of photodynamic therapy comprising the administration of photosensitizers, such as hematoporphyrin and its derivatives, Vertoporfin (BPD-MA), phthalocyanine, photosensitizer Pc4, demethoxy-hypocrellin A; and 2BA-2-DMHA.
[0273] In various embodiments, non-platinum comprising therapies include, for example, immunotherapy. Immunotherapy may comprise, for example, use of cancer vaccines and/or sensitized antigen presenting cells. The immunotherapy can involve passive immunity for short-term protection of a host, achieved by the administration of pre-formed antibody directed against a cancer antigen or disease antigen (e.g., administration of a monoclonal antibody, optionally linked to a chemotherapeutic agent or toxin, to a tumor antigen). Immunotherapy can also focus on using the cytotoxic lymphocyte-recognized epitopes of cancer cell lines.
[0274] In various embodiments, non-platinum comprising therapies include, for example, hormonal therapy, Hormonal therapeutic treatments can comprise, for example, hormonal agonists, hormonal antagonists (e.g., flutamide, bicalutamide, tamoxifen, raloxifene, leuprolide acetate (LUPRON), LH-RH antagonists), inhibitors of hormone biosynthesis and processing, and steroids (e.g., dexamethasone, retinoids, deltoids, betamethasone, cortisol, cortisone, prednisone, dehydrotestosterone, glucocorticoids, mineralocorticoids, estrogen, testosterone, progestins), vitamin A derivatives (e.g., all-trans retinoic acid (ATRA)); vitamin D3 analogs; antigestagens (e.g., mifepristone, onapristone), or antiandrogens (e.g., cyproterone acetate).
[0275] In various embodiments described herein, the anthracycline is epirubincin or doxorubicin.
[0276] The duration and/or dose of treatment with anti-cancer therapies may vary according to the particular anti-cancer agent or combination thereof. An appropriate treatment time for a particular cancer therapeutic agent will be appreciated by the skilled artisan. The invention contemplates the continued assessment of optimal treatment schedules for each cancer therapeutic agent, where the genetic signature of the cancer of the subject as determined by the methods of the invention is a factor in determining optimal treatment doses and schedules.
Cancers for which the Genetic Signature can be Determined
[0277] The methods of the invention can be used to determine the genetic signature of many different cancers. Specific examples of types of cancers for which the genetic signature can be determined by the methods encompassed by the invention include, but are not limited to, human sarcomas and carcinomas, e.g., fibrosarcoma, myxosarcoma, liposarcoma, chondrosarcoma, osteogenic sarcoma, chordoma, angiosarcoma, endotheliosarcoma, lymphangiosarcoma, lymphangioendotheliosarcoma, synovioma, mesothelioma, Ewing's tumor, leiomyosarcoma, rhabdomyosarcoma, colon carcinoma, colorectal cancer, pancreatic cancer, breast cancer, ovarian cancer, prostate cancer, squamous cell carcinoma, basal cell carcinoma, adenocarcinoma, sweat gland carcinoma, sebaceous gland carcinoma, papillary carcinoma, papillary adenocarcinomas, cystadenocarcinoma, medullary carcinoma, bronchogenic carcinoma, renal cell carcinoma, hepatoma, bile duct carcinoma, liver cancer, choriocarcinoma, seminoma, embryonal carcinoma, Wilms' tumor, cervical cancer, bone cancer, brain tumor, testicular cancer, lung carcinoma, small cell lung carcinoma, bladder carcinoma, epithelial carcinoma, glioma, astrocytoma, medulloblastoma, craniopharyngioma, ependymoma, pinealoma, hemangioblastoma, acoustic neuroma, oligodendroglioma, meningioma, melanoma, neuroblastoma, retinoblastoma; leukemias, e.g., acute lymphocytic leukemia and acute myelocytic leukemia (myeloblastic, promyelocytic, myelomonocytic, monocytic and erythroleukemia); chronic leukemia (chronic myelocytic (granulocytic) leukemia and chronic lymphocytic leukemia); and polycythemia vera, lymphoma (Hodgkin's disease and non-Hodgkin's disease), multiple myeloma, Waldenstrom's macroglobulinemia, and heavy chain disease.
[0278] In some embodiments, the cancer whose genetic signature is determined by the method of the invention is an epithelial cancer such as, but not limited to, bladder cancer, breast cancer, cervical cancer, colon cancer, gynecologic cancers, renal cancer, laryngeal cancer, lung cancer, oral cancer, head and neck cancer, ovarian cancer, pancreatic cancer, prostate cancer, or skin cancer. In other embodiments, the cancer is breast cancer, prostate cancer, lung cancer, or colon cancer. In still other embodiments, the epithelial cancer is non-small-cell lung cancer, nonpapillary renal cell carcinoma, cervical carcinoma, ovarian carcinoma (e.g., serous ovarian carcinoma), or breast carcinoma. The epithelial cancers may be characterized in various other ways including, but not limited to, serous, endometrioid, mucinous, clear cell, brenner, or undifferentiated. In still other embodiments, the cancer is breast cancer, ovarian cancer or lung cancer. In particular embodiments, the cancer is triple negative breast cancer.
Subjects
[0279] In various embodiments, the subject for whom predicted efficacy of an anti-cancer therapy is determined, is a mammal (e.g., mouse, rat, primate, non-human mammal, domestic animal such as dog, cat, cow, horse), and is preferably a human. In another embodiment of the methods of the invention, the subject has not undergone chemotherapy or radiation therapy. In alternative embodiments, the subject has undergone chemotherapy or radiation therapy (e.g., such as with cisplatin, carboplatin, and/or taxane). In related embodiments, the subject has not been exposed to levels of radiation or chemotoxic agents above those encountered generally or on average by the subjects of a species. In certain embodiments, the subject has had surgery to remove cancerous or precancerous tissue. In other embodiments, the cancerous tissue has not been removed, e.g., the cancerous tissue may be located in an inoperable region of the body, such as in a tissue that is essential for life, or in a region where a surgical procedure would cause considerable risk of harm to the patient, or e.g., the subject is given the anti-cancer therapy prior to removal of the cancerous tissue.
Nucleic Acid Sample Preparation
[0280] A. Nucleic Acid Isolation
[0281] Nucleic acid samples derived from cancerous and non-cancerous cells of a subject that can be used in the methods of the invention to determine the genetic signature of a cancer can be prepared by means well known in the art. For example, surgical procedures or needle biopsy aspiration can be used to collect cancerous samples from a subject. In some embodiments, it is important to enrich and/or purify the cancerous tissue and/or cell samples from the non-cancerous tissue and/or cell samples. In other embodiments, the cancerous tissue and/or cell samples can then be microdissected to reduce the amount of normal tissue contamination prior to extraction of genomic nucleic acid or pre-RNA for use in the methods of the invention. In still another embodiment, the cancerous tissue and/or cell samples are enriched for cancer cells by at least 50%, 55%, 60%, 65%, 70%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or more or any range in between, in cancer cell content. Such enrichment can be accomplished according to methods well-known in the art, such as needle microdissection, laser microdissection, fluorescence activated cell sorting, and immunological cell sorting. In one embodiment, an automated machine performs the hyperproliferative cell enrichment to thereby transform the biological sample into a purified form enriched for the presence of hyperproliferative cells.
[0282] Collecting nucleic acid samples from non-cancerous cells of a subject can also be accomplished with surgery or aspiration. In surgical procedures where cancerous tissue is removed, surgeons often remove non-cancerous tissue and/or cell samples of the same tissue type of the cancer patient for comparison. Nucleic acid samples can be isolated from such non-cancerous tissue of the subject for use in the methods of the invention. In certain embodiments of the methods of the invention, nucleic acid samples from non-cancerous tissues are not derived from the same tissue type as the cancerous tissue and/or cells sampled, and/or are not derived from the cancer patient. The nucleic acid samples from non-cancerous tissues may be derived from any non-cancerous and/or disease-free tissue and/or cells. Such non-cancerous samples can be collected by surgical or non-surgical procedures. In certain embodiments, non-cancerous nucleic acid samples are derived from tumor-free tissues. For example, non-cancerous samples may be collected from lymph nodes, peripheral blood lymphocytes, and/or mononuclear blood cells, or any subpopulation thereof. In a preferred embodiment, the non-cancerous tissue is not pre-cancerous tissue, e.g., it does not exhibit any indicia of a pre-neoplastic condition such as hyperplasia, metaplasia, or dysplasia.
[0283] In one embodiment, the nucleic acid samples used to compute a reference value are taken from at least 1, 2, 5, 10, 20, 30, 40, 50, 100, or 200 different organisms of that species. According to certain aspects of the invention, nucleic acid "derived from" genomic DNA, as used in the methods of the invention, e.g., in hybridization experiments to determine BLM expression, FANCI expression, or 15q26 copy number, chromosome 15 centromere copy number can be fragments of genomic nucleic acid generated by restriction enzyme digestion and/or ligation to other nucleic acid, and/or amplification products of genomic nucleic acids, or pre-messenger RNA (pre-mRNA), amplification products of pre-mRNA, or genomic DNA fragments grown up in cloning vectors generated, e.g., by "shotgun" cloning methods. In certain embodiments, genomic nucleic acid samples are digested with restriction enzymes.
[0284] B. Amplification of Nucleic Acids
[0285] Though the nucleic acid sample need not comprise amplified nucleic acid, in some embodiments, the isolated nucleic acids can be processed in manners requiring and/or taking advantage of amplification. The genomic DNA samples of a subject optionally can be fragmented using restriction endonucleases and/or amplified prior to determining analysis. In one embodiment, the DNA fragments are amplified using polymerase chain reaction (PCR). Methods for practicing PCR are well known to those of skill in the art. One advantage of PCR is that small quantities of DNA can be used. For example, genomic DNA from a subject may be about 150 ng, 175, ng, 200 ng, 225 ng, 250 ng, 275 ng, or 300 ng of DNA.
[0286] In certain embodiments of the methods of the invention, the nucleic acid from a subject is amplified using a single primer pair. For example, genomic DNA samples can be digested with restriction endonucleases to generate fragments of genomic DNA that are then ligated to an adaptor DNA sequence which the primer pair recognizes. In other embodiments of the methods of the invention, the nucleic acid of a subject is amplified using sets of primer pairs specific to BLM, FANCI, 15q26, or chromosome 15 centromere copy and in instances wherein BRCA1, BRCA2, ER, PgR and/or HER2 receptor expression is also to be assessed, sets of primer pairs specific to BRCA1, BRCA2, ER, PgR and/or HER2 receptor. Such sets of primer pairs each recognize genomic DNA sequences flanking BLM, FANCI, 15q26, or chromosome 15 centromere and BRCA1, BRCA2, ER, PgR and/or HER2 receptor wherein the expression is also to be assessed. A DNA sample suitable for hybridization can be obtained, e.g., by polymerase chain reaction (PCR) amplification of genomic DNA, fragments of genomic DNA, fragments of genomic DNA ligated to adaptor sequences or cloned sequences. Computer programs that are well known in the art can be used in the design of primers with the desired specificity and optimal amplification properties, such as Oligo version 5.0 (National Biosciences). PCR methods are well known in the art, and are described, for example, in Innis et al., eds., 1990, PCR Protocols: A Guide to Methods And Applications, Academic Press Inc., San Diego, Calif. It will be apparent to one skilled in the art that controlled robotic systems are useful for isolating and amplifying nucleic acids and can be used.
[0287] In other embodiments, where genomic DNA of a subject is fragmented using restriction endonucleases and amplified prior to analysis, the amplification can comprise cloning regions of genomic DNA of the subject. In such methods, amplification of the DNA regions is achieved through the cloning process. For example, expression vectors can be engineered to express large quantities of particular fragments of genomic DNA of the subject (Sambrook, J. et al., eds., 1989, Molecular Cloning: A Laboratory Manual, 2nd Ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., at pp. 9.47-9.51).
[0288] In yet other embodiments, where the DNA of a subject is fragmented using restriction endonucleases and amplified prior to analysis, the amplification comprises expressing a nucleic acid encoding a gene, or a gene and flanking genomic regions of nucleic acids, from the subject. RNA (pre-messenger RNA) that comprises the entire transcript including introns is then isolated and used in the methods of the invention to analyze and provide a genetic signature of a cancer. In certain embodiments, no amplification is required. In such embodiments, the genomic DNA, or pre-RNA, of a subject may be fragmented using restriction endonucleases or other methods. The resulting fragments may be hybridized to SNP probes. Typically, greater quantities of DNA are needed to be isolated in comparison to the quantity of DNA or pre-mRNA needed where fragments are amplified. For example, where the nucleic acid of a subject is not amplified, a DNA sample of a subject for use in hybridization may be about 400 ng, 500 ng, 600 ng, 700 ng, 800 ng, 900 ng, or 1000 ng of DNA or greater. Alternatively, in other embodiments, methods are used that require very small amounts of nucleic acids for analysis, such as less than 400 ng, 300 ng, 200 ng, 100 ng, 90 ng, 85 ng, 80 ng, 75 ng, 70 ng, 65 ng, 60 ng, 55 ng, 50 ng, or less, such as is used for molecular inversion probe (MIP) assays. These techniques are particularly useful for analyzing clinical samples, such as paraffin embedded formalin-fixed material or small core needle biopsies, characterized as being readily available but generally having reduced DNA quality (e.g., small, fragmented DNA) and/or not providing large amounts of nucleic acids.
[0289] C. Hybridization
[0290] The nucleic acid samples derived from a subject used in the methods of the invention can be hybridized to arrays comprising probes (e.g., oligonucleotide probes) in order to identify BLM, FANCI, 15q26, or chromosome 15 centromere and in instances wherein BRCA1, BRCA2, ER, PgR and/or HER2 receptor expression is also to be assessed, comprising probes in order to identify BRCA1, BRCA2, ER, PgR and/or HER2 receptor. Hybridization can also be used to determine whether the BLM, FANCI, 15q26, or chromosome 15 centromere identified exhibit total copy number change, copy number gain, and copy number loss in nucleic acid samples from cancerous tissues and/or cells of the subject. In preferred embodiments, the probes used in the methods of the invention comprise an array of probes that can be tiled on a DNA chip (e.g., SNP oligonucleotide probes). In some embodiments, BLM expression, FANCI expression, 15q26 copy number, or chromosome 15 centromere copy number is determined by a method that does not comprise detecting a change in size of restriction enzyme-digested nucleic acid fragments. In other embodiments, SNPs are analyzed to identify BLM expression or FANCI expression, 15q26 copy number or chromosome 15 centromere copy number. Hybridization and wash conditions used in the methods of the invention are chosen so that the nucleic acid samples to be analyzed by the invention specifically bind or specifically hybridize to the complementary oligonucleotide sequences of the array, preferably to a specific array site, wherein its complementary DNA is located. In some embodiments, the complementary DNA can be completely matched or mismatched to some degree as used, for example, in Affymetrix oligonucleotide arrays such as those used to analyze SNPs in MIP assays. The single-stranded synthetic oligodeoxyribonucleic acid DNA probes of an array may need to be denatured prior to contact with the nucleic acid samples from a subject, e.g., to remove hairpins or dimers which form due to self-complementary sequences.
[0291] Optimal hybridization conditions will depend on the length of the probes and type of nucleic acid samples from a subject. General parameters for specific (i.e., stringent) hybridization conditions for nucleic acids are described in Sambrook, J. et al., eds., 1989, Molecular Cloning: A Laboratory Manual, 2nd Ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., at pp. 9.47-9.51 and 11.55-11.61; Ausubel et al., eds., 1989, Current Protocols in Molecules Biology, Vol. 1, Green Publishing Associates, Inc., John Wiley & Sons, Inc., New York, at pp. 2.10.1-2.10.16. Exemplary useful hybridization conditions are provided in, e.g., Tijessen, 1993, Hybridization with Nucleic Acid Probes, Elsevier Science Publishers B. V. and Kricka, 1992, Nonisotopic DNA Probe Techniques, Academic Press, San Diego, Calif.
[0292] D. Oligonucleotide Nucleic Acid Arrays
[0293] In some embodiments of the methods of the present invention, DNA arrays can be used to determine total copy number change, copy number gain, and copy number loss by measuring the level of hybridization of the nucleic acid sequence to oligonucleotide probes that comprise complementary sequences. Hybridization can be used to determine the presence or absence of heterozygosity. Various formats of DNA arrays that employ oligonucleotide "probes," (i.e., nucleic acid molecules having defined sequences) are well known to those of skill in the art. Typically, a set of nucleic acid probes, each of which has a defined sequence, is immobilized on a solid support in such a manner that each different probe is immobilized to a predetermined region. In certain embodiments, the set of probes forms an array of positionally-addressable binding (e.g., hybridization) sites on a support. Each of such binding sites comprises a plurality of oligonucleotide molecules of a probe bound to the predetermined region on the support. More specifically, each probe of the array is preferably located at a known, predetermined position on the solid support such that the identity (i.e., the sequence) of each probe can be determined from its position on the array (i.e., on the support or surface). Microarrays can be made in a number of ways, of which several are described herein. However produced, microarrays share certain characteristics, they are reproducible, allowing multiple copies of a given array to be produced and easily compared with each other.
[0294] Preferably, the microarrays are made from materials that are stable under binding (e.g., nucleic acid hybridization) conditions. The microarrays are preferably small, e.g., between about 1 cm.sup.2 and 25 cm.sup.2, preferably about 1 to 3 cm.sup.2. However, both larger and smaller arrays are also contemplated and may be preferable, e.g., for simultaneously evaluating a very large number of different probes. Oligonucleotide probes can be synthesized directly on a support to form the array. The probes can be attached to a solid support or surface, which may be made, e.g., from glass, plastic (e.g., polypropylene, nylon), polyacrylamide, nitrocellulose, gel, or other porous or nonporous material. The set of immobilized probes or the array of immobilized probes is contacted with a sample containing labeled nucleic acid species so that nucleic acids having sequences complementary to an immobilized probe hybridize or bind to the probe. After separation of, e.g., by washing off, any unbound material, the bound, labeled sequences are detected and measured. The measurement is typically conducted with computer assistance. Using DNA array assays, complex mixtures of labeled nucleic acids, e.g., nucleic acid fragments derived a restriction digestion of genomic DNA from non-cancerous tissue, can be analyzed. DNA array technologies have made it possible to determine the expression level or copy number BLM, FANCI, 15q26, or chromosome 15 centromere, or BRCA1, BRCA2, ER, PgR and/or HER2 receptor expression in instances where BRCA1, BRCA2, ER, PgR and/or HER2 receptor expression is also assessed.
[0295] In certain embodiments, high-density oligonucleotide arrays are used in the methods of the invention. These arrays containing thousands of oligonucleotides complementary to defined sequences, at defined locations on a surface can be synthesized in situ on the surface by, for example, photolithographic techniques (see, e.g., Fodor et al., 1991, Science 251:767-773; Pease et al., 1994, Proc. Natl. Acad. Sci. U.S.A. 91:5022-5026; Lockhart et al., 1996, Nature Biotechnology 14:1675; U.S. Pat. Nos. 5,578,832; 5,556,752; 5,510,270; 5,445,934; 5,744,305; and 6,040,138). Methods for generating arrays using inkjet technology for in situ oligonucleotide synthesis are also known in the art (see, e.g., Blanchard, International Patent Publication WO 98/41531, published Sep. 24, 1998; Blanchard et al., 1996, Biosensors And Bioelectronics 11:687-690; Blanchard, 1998, in Synthetic DNA Arrays in Genetic Engineering, Vol. 20, J. K. Setlow, Ed., Plenum Press, New York at pages 111-123). Another method for attaching the nucleic acids to a surface is by printing on glass plates, as is described generally by Schena et al. (1995, Science 270:467-470). Other methods for making microarrays, e.g., by masking (Maskos and Southern, 1992, Nucl. Acids. Res. 20:1679-1684), may also be used. When these methods are used, oligonucleotides (e.g., 15 to 60-mers) of known sequence are synthesized directly on a surface such as a derivatized glass slide. The array produced can be redundant, with several oligonucleotide molecules corresponding to each informative locus of interest (e.g., SNPs, RFLPs, STRs, etc.).
[0296] One exemplary means for generating the oligonucleotide probes of the DNA array is by synthesis of synthetic polynucleotides or oligonucleotides, e.g., using N-phosphonate or phosphoramidite chemistries (Froehler et al., 1986, Nucleic Acid Res. 14:5399-5407; McBride et al., 1983, Tetrahedron Lett. 24:246-248). Synthetic sequences are typically between about 15 and about 600 bases in length, more typically between about 20 and about 100 bases, most preferably between about 40 and about 70 bases in length. In some embodiments, synthetic nucleic acids include non-natural bases, such as, but by no means limited to, inosine. As noted above, nucleic acid analogues may be used as binding sites for hybridization. An example of a suitable nucleic acid analogue is peptide nucleic acid (see, e.g., Egholm et al., 1993, Nature 363:566-568; U.S. Pat. No. 5,539,083). In alternative embodiments, the hybridization sites (i.e., the probes) are made from plasmid or phage clones of regions of genomic DNA corresponding to SNPs or the complement thereof. The size of the oligonucleotide probes used in the methods of the invention can be at least 10, 20, 25, 30, 35, 40, 45, or 50 nucleotides in length. It is well known in the art that although hybridization is selective for complementary sequences, other sequences which are not perfectly complementary may also hybridize to a given probe at some level. Thus, multiple oligonucleotide probes with slight variations can be used, to optimize hybridization of samples. To further optimize hybridization, hybridization stringency condition, e.g., the hybridization temperature and the salt concentrations, may be altered by methods that are well known in the art.
[0297] In preferred embodiments, the high-density oligonucleotide arrays used in the methods of the invention comprise oligonucleotides corresponding to BLM, FANCI, 15q26, or chromosome 15 centromere or in instances wherein BRCA1, BRCA2, ER, PgR and/or HER2 receptor expression is also assessed, the arrays also comprise oligonucleotides corresponding to BRCA1, BRCA2, ER, PgR and/or HER2 receptor. The oligonucleotide probes may comprise DNA or DNA "mimics" (e.g., derivatives and analogues) corresponding to a portion of each informative locus of interest (e.g., SNPs, RFLPs, STRs, etc.) in a subject's genome. The oligonucleotide probes can be modified at the base moiety, at the sugar moiety, or at the phosphate backbone. Exemplary DNA mimics include, e.g., phosphorothioates. For each SNP locus, a plurality of different oligonucleotides may be used that are complementary to the sequences of sample nucleic acids. For example, for a single informative locus of interest (e.g., SNPs, RFLPs, STRs, etc.) about 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, or more different oligonucleotides can be used. Each of the oligonucleotides for a particular informative locus of interest may have a slight variation in perfect matches, mismatches, and flanking sequence around the SNP. In certain embodiments, the probes are generated such that the probes for a particular informative locus of interest comprise overlapping and/or successive overlapping sequences which span or are tiled across a genomic region containing the target site, where all the probes contain the target site. By way of example, overlapping probe sequences can be tiled at steps of a predetermined base interval, e. g. at steps of 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 bases intervals. In certain embodiments, the assays can be performed using arrays suitable for use with molecular inversion probe protocols such as described by Wang et al. (2007) Genome Biol. 8, R246. For oligonucleotide probes targeted at nucleic acid species of closely resembled (i.e., homologous) sequences, "cross-hybridization" among similar probes can significantly contaminate and confuse the results of hybridization measurements. Cross-hybridization is a particularly significant concern in the detection of SNPs since the sequence to be detected (i.e., the particular SNP) must be distinguished from other sequences that differ by only a single nucleotide. Cross-hybridization can be minimized by regulating either the hybridization stringency condition and/or during post-hybridization washings. Highly stringent conditions allow detection of allelic variants of a nucleotide sequence, e.g., about 1 mismatch per 10-30 nucleotides. There is no single hybridization or washing condition which is optimal for all different nucleic acid sequences. For particular arrays of BLM, FANCI, 15q26, or chromosome 15 centromere, or of BRCA1, BRCA2, ER, PgR and/or HER2 receptor these conditions can be identical to those suggested by the manufacturer or can be adjusted by one of skill in the art. In preferred embodiments, the probes used in the methods of the invention are immobilized (i.e., tiled) on a glass slide called a chip. For example, a DNA microarray can comprises a chip on which oligonucleotides (purified single-stranded DNA sequences in solution) have been robotically printed in an (approximately) rectangular array with each spot on the array corresponds to a single DNA sample which encodes an oligonucleotide. In summary the process comprises, flooding the DNA microarray chip with a labeled sample under conditions suitable for hybridization to occur between the slide sequences and the labeled sample, then the array is washed and dried, and the array is scanned with a laser microscope to detect hybridization. In certain embodiments there are at least 250, 500, 1,000, 2,000, 3,000, 4,000, 5,000, 6,000, 7,000, 8,000, 9,000, 10,000, 11,000, 12,000, 13,000, 14,000, 15,000, 16,000, 17,000, 18,000, 19,000, 20,000, 21,000, 22,000, 23,000, 24,000, 25,000, 26,000, 27,000, 28,000, 29,000, 30,000, 31,000, 32,000, 33,000, 34,000, 35,000, 36,000, 37,000, 38,000, 39,000, 40,000, 41,000, 42,000, 43,000, 44,000, 45,000, 50,000, 60,000, 70,000, 80,000, 90,000, 100,000 or more or any range in between, of BLM, FANCI, 15q26, or chromosome 15 centromere, or of BRCA1, BRCA2, ER, PgR and/or HER2 receptor for which probes appear on the array (with match/mismatch probes for a single locus of interest or probes tiled across a single locus of interest counting as one locus of interest). The maximum number of BLM, FANCI, 15q26, or chromosome 15 centromere, or of BRCA1, BRCA2, ER, PgR and/or HER2 receptor being probed per array is determined by the size of the genome and genetic diversity of the subjects species. DNA chips are well known in the art and can be purchased in pre-5 fabricated form with sequences specific to particular species. In some embodiments, the Genome-Wide Human SNP Array 6.0.TM. and/or the 50K XbaI arrays (Affymetrix, Santa Clara, Calif.) are used in the methods of the invention. In other embodiments, SNPs and/or DNA copy number can be detected and quantitated using sequencing methods, such as "next-generation sequencing methods" as described further above.
[0298] E. Signal Detection
[0299] In some embodiments, nucleic acid samples derived from a subject are hybridized to the binding sites of an array described herein. In certain embodiments, nucleic acid samples derived from each of the two sample types of a subject (i.e., cancerous and non-cancerous) are hybridized to separate, though identical, arrays. In certain embodiments, nucleic acid samples derived from one of the two sample types of a subject (i.e., cancerous and non-cancerous) is hybridized to such an array, then following signal detection the chip is washed to remove the first labeled sample and reused to hybridize the remaining sample. In other embodiments, the array is not reused more than once. In certain embodiments, the nucleic acid samples derived from each of the two sample types of a subject (i.e., cancerous and non-cancerous) are differently labeled so that they can be distinguished. When the two samples are mixed and hybridized to the same array, the relative intensity of signal from each sample is determined for each site on the array, and any relative difference in abundance of an allele of BLM, FANCI, 15q26, or chromosome 15 centromere, or of BRCA1, BRCA2, ER, PgR and/or HER2 receptor. Signals can be recorded and, in some embodiments, analyzed by computer. In one embodiment, the scanned image is despeckled using a graphics program (e.g., Hij aak Graphics Suite) and then analyzed using an image gridding program that creates a spreadsheet of the average hybridization at each wavelength at each site. If necessary, an experimentally determined correction for "cross talk" (or overlap) between the channels for the two fluors may be made. For any particular hybridization site on the array, a ratio of the emission of the two fluorophores can be calculated, which may help in eliminating cross hybridization signals to more accurately determining whether a particular SNP locus is heterozygous or homozygous.
[0300] F. Labeling
[0301] In some embodiments, the nucleic acids samples, fragments thereof, or fragments thereof ligated to adaptor regions used in the methods of the invention are detectably labeled. For example, the detectable label can be a fluorescent label, e.g., by incorporation of nucleotide analogues. Other labels suitable for use in the present invention include, but are not limited to, biotin, iminobiotin, antigens, cofactors, dinitrophenol, lipoic acid, olefinic compounds, detectable polypeptides, electron rich molecules, enzymes capable of generating a detectable signal by action upon a substrate, and radioactive isotopes.
[0302] Radioactive isotopes include that can be used in conjunction with the methods of the invention, but are not limited to, 32P and 14C. Fluorescent molecules suitable for the present invention include, but are not limited to, fluorescein and its derivatives, rhodamine and its derivatives, texas red, 5'carboxy-fluorescein ("FAM"), 2',7'-dimethoxy-4',5'-dichloro-6-carboxy-fluorescein ("JOE"), N, N, N',N'-tetramethyl-6-carboxy-rhodamine ("TAMRA"), 6-carboxy-X-rhodamine ("ROX"), HEX, TET, IRD40, and IRD41.
[0303] Fluorescent molecules which are suitable for use according to the invention further include: cyamine dyes, including but not limited to Cy2, Cy3, Cy3.5, CY5, Cy5.5, Cy7 and FLUORX; BODIPY dyes including but not limited to BODIPY-FL, BODIPY-TR, BODIPY-TMR, BODIPY-630/650, and BODIPY-650/670; and ALEXA dyes, including but not limited to ALEXA-488, ALEXA-532, ALEXA-546, ALEXA-568, and ALEXA-594; as well as other fluorescent dyes which will be known to those who are skilled in the art. Electron rich indicator molecules suitable for the present invention include, but are not limited to, ferritin, hemocyanin, and colloidal gold.
[0304] Two-color fluorescence labeling and detection schemes may also be used (Shena et al., 1995, Science 270:467-470). Use of two or more labels can be useful in detecting variations due to minor differences in experimental conditions (e.g., hybridization conditions). In some embodiments of the invention, at least 5, 10, 20, or 100 dyes of different colors can be used for labeling. Such labeling would also permit analysis of multiple samples simultaneously which is encompassed by the invention.
[0305] The labeled nucleic acid samples, fragments thereof, or fragments thereof ligated to adaptor regions that can be used in the methods of the invention are contacted to a plurality of oligonucleotide probes under conditions that allow sample nucleic acids having sequences complementary to the probes to hybridize thereto. Depending on the type of label used, the hybridization signals can be detected using methods well known to those of skill in the art including, but not limited to, X-Ray film, phosphor imager, or CCD camera. When fluorescently labeled probes are used, the fluorescence emissions at each site of a transcript array can be, preferably, detected by scanning confocal laser microscopy. In one embodiment, a separate scan, using the appropriate excitation line, is carried out for each of the two fluorophores used. Alternatively, a laser can be used that allows simultaneous specimen illumination at wavelengths specific to the two fluorophores and emissions from the two fluorophores can be analyzed simultaneously (see Shalon et al. (1996) Genome Res. 6, 639-645). In a preferred embodiment, the arrays are scanned with a laser fluorescence scanner with a computer controlled X-Y stage and a microscope objective. Sequential excitation of the two fluorophores is achieved with a multi-line, mixed gas laser, and the emitted light is split by wavelength and detected with two photomultiplier tubes. Such fluorescence laser scanning devices are described, e.g., in Schena et al. (1996) Genome Res. 6, 639-645. Alternatively, a fiber-optic bundle can be used such as that described by Ferguson et al. (1996) Nat. Biotech. 14, 1681-1684. The resulting signals can then be analyzed to determine the BLM expression, FANCI expression, 15q26 copy number, or chromosome 15 centromere copy number, using computer software.
[0306] G. Algorithms for Analyzing BLM, FANCI, 15q26, or Chromosome 15 Centromere
[0307] Once the hybridization signal has been detected the resulting data can be analyzed using algorithms. In certain embodiments, the algorithm for determining the expression of BLM or FANCI, or copy number 15q26 or chromosome 15 centromere is based on well-known methods. Additional representative illustrations of such well known algorithms are provided in the Examples section below.
[0308] H. Computer Implementation Systems and Methods
[0309] In certain embodiments, the methods of the invention implement a computer program to calculate a copy number, copy number gain, copy number loss, and expression levels. For example, a computer program can be used to perform the algorithms described herein. A computer system can also store and manipulate data generated by the methods of the present invention which comprises a plurality of hybridization signal changes/profiles during approach to equilibrium in different hybridization measurements and which can be used by a computer system in implementing the methods of this invention. In certain embodiments, a computer system receives probe hybridization data; (ii) stores probe hybridization data; and (iii) compares probe hybridization data to determine the state of BLM, FANCI, 15q26, or, chromosome 15 centromere, and/or of BRCA1, BRCA2, ER, PgR and/or HER2 receptor in said nucleic acid sample from cancerous or pre-cancerous tissue. The copy number, copy number gain, copy number loss, or expression levels is then calculated. In some embodiments, a computer system (i) compares the determined copy number, copy number gain, copy number loss, and expression levels to a threshold value or reference value; and (ii) outputs an indication of whether said copy number, copy number gain, copy number loss, and expression levels is above or below a threshold value, or a genetic signature based on said indication. In certain embodiments, such computer systems are also considered part of the present invention.
[0310] Numerous types of computer systems can be used to implement the analytic methods of this invention according to knowledge possessed by a skilled artisan in the bioinformatics and/or computer arts.
[0311] Several software components can be loaded into memory during operation of such a computer system. The software components can comprise both software components that are standard in the art and components that are special to the present invention (e.g., dCHIP software described in Lin et al. (2004) Bioinformatics 20, 1233-1240; CRLMM software described in Silver et al. (2007) Cell 128, 991-1002; Aroma Affymetrix software described in Richardson et al. (2006) Cancer Cell 9, 121-132. The methods of the invention can also be programmed or modeled in mathematical software packages that allow symbolic entry of equations and high-level specification of processing, including specific algorithms to be used, thereby freeing a user of the need to procedurally program individual equations and algorithms. Such packages include, e.g., Matlab from Mathworks (Natick, Mass.), Mathematica from Wolfram Research (Champaign, Ill.) or S-Plus from MathSoft (Seattle, Wash.). In certain embodiments, the computer comprises a database for storage of hybridization signal profiles. Such stored profiles can be accessed and used to calculate a copy number, copy number gain, copy number loss, or expression level. For example, of the hybridization signal profile of a sample derived from the non-cancerous tissue of a subject and/or profiles generated from population-based distributions of BLM, FANCI, 15q26, or chromosome 15 centromere, and/or of BRCA1, BRCA2, ER, PgR and/or HER2 receptor in relevant populations of the same species were stored, it could then be compared to the hybridization signal profile of a sample derived from the cancerous tissue of the subject.
[0312] In addition to the exemplary program structures and computer systems described herein, other, alternative program structures and computer systems will be readily apparent to the skilled artisan. Such alternative systems, which do not depart from the above described computer system and programs structures either in spirit or in scope, are therefore intended to be comprehended within the accompanying claims.
[0313] Once a laboratory technician or laboratory professional or group of laboratory technicians or laboratory professionals determines whether a sample has a copy number, copy number gain, copy number loss, or expression level as described above (e.g., step (1) in many of the methods above), the same or a different laboratory technician or laboratory professional (or group) can analyze a plurality of test BLM, FANCI, 15q26, or chromosome 15 centromere to determine whether there is a copy number, copy number gain, copy number loss, or expression level (e.g., step (2) in many of the methods above). Next, the same or a different laboratory technician or laboratory professional (or group) can combine the a copy number, copy number gain, copy number loss, or expression level data from the test BLM, FANCI, 15q26, or chromosome 15 centromere to derive a copy number, copy number gain, copy number loss, or expression level (e.g., step (3) in many of the methods above). Optionally, the same or a different laboratory technician or laboratory professional (or group) can correlate the copy number, copy number gain, copy number loss, or expression level to an increased or decreased likelihood of response to a particular therapy (e.g., those mentioned above).
[0314] In various embodiments, provided herein is a computer readable storage medium comprising: a storing data module containing data from a sample comprising a cancer cell obtained from a subject that represents an expression level from an assay for BLM and/or FANCI, or copy number of 15q26; a comparison module that compares the data stored on the storing data module with a reference data and/or control data, and to provide a comparison content, and an output module displaying the comparison content for the user, wherein the increased expression of BLM and/or FANCI, or copy number gain of 15q26 indicates that the subject is susceptible to platinum-comprising, or anthracycline-comprising cancer therapy.
[0315] In various embodiments, the control data comprises data from a population of non-cancerous healthy individuals. In various embodiments, the control data comprises data BRAC1 expression, and/or a housekeeping gene expression.
[0316] Embodiments of the invention can be described through functional modules, which are defined by computer executable instructions recorded on computer readable media and which cause a computer to perform method steps when executed. The modules are segregated by function, for the sake of clarity. However, it should be understood that the modules/systems need not correspond to discreet blocks of code and the described functions can be carried out by the execution of various code portions stored on various media and executed at various times. Furthermore, it should be appreciated that the modules may perform other functions, thus the modules are not limited to having any particular functions or set of functions.
[0317] The computer readable storage media 30 can be any available tangible media that can be accessed by a computer. Computer readable storage media includes volatile and nonvolatile, removable and non-removable tangible media implemented in any method or technology for storage of information such as computer readable instructions, data structures, program modules or other data. Computer readable storage media includes, but is not limited to, RAM (random access memory), ROM (read only memory), EPROM (eraseable programmable read only memory), EEPROM (electrically eraseable programmable read only memory), flash memory or other memory technology, CD-ROM (compact disc read only memory), DVDs (digital versatile disks) or other optical storage media, magnetic cassettes, magnetic tape, magnetic disk storage or other magnetic storage media, other types of volatile and non-volatile memory, and any other tangible medium which can be used to store the desired information and which can accessed by a computer including and any suitable combination of the foregoing.
[0318] Computer-readable data embodied on one or more computer-readable media may define instructions, for example, as part of one or more programs that, as a result of being executed by a computer, instruct the computer to perform one or more of the functions described herein, and/or various embodiments, variations and combinations thereof. Such instructions may be written in any of a plurality of programming languages, for example, Java, J#, Visual Basic, C, C#, C++, Fortran, Pascal, Eiffel, Basic, COBOL assembly language, and the like, or any of a variety of combinations thereof. The computer-readable media on which such instructions are embodied may reside on one or more of the components of either of a system, or a computer readable storage medium described herein, may be distributed across one or more of such components.
[0319] The computer-readable media may be transportable such that the instructions stored thereon can be loaded onto any computer resource to implement the aspects of the present invention discussed herein. In addition, it should be appreciated that the instructions stored on the computer-readable medium, described above, are not limited to instructions embodied as part of an application program running on a host computer. Rather, the instructions may be embodied as any type of computer code (e.g., software or microcode) that can be employed to program a computer to implement aspects of the present invention. The computer executable instructions may be written in a suitable computer language or combination of several languages. Basic computational biology methods are known to those of ordinary skill in the art and are described in, for example, Setubal and Meidanis et al., Introduction to Computational Biology Methods (PWS Publishing Company, Boston, 1997); Salzberg, Searles, Kasif, (Ed.), Computational Methods in Molecular Biology, (Elsevier, Amsterdam, 1998); Rashidi and Buehler, Bioinformatics Basics: Application in Biological Science and Medicine (CRC Press, London, 2000) and Ouelette and Bzevanis Bioinformatics: A Practical Guide for Analysis of Gene and Proteins (Wiley & Sons, Inc., 2nd ed., 2001).
[0320] The functional modules of certain embodiments of the invention; for example, as depicted in FIG. 12, include for example, at a measuring module 40, a storage module 30, a comparison module 80, and an output module 110. The functional modules can be executed on one, or multiple, computers, or by using one, or multiple, computer networks. The measuring module has computer executable instructions to provide e.g., expression information in computer readable form.
[0321] The measuring module 40, can comprise any system for detecting the expression of BLM and/or FANCI, the 15Q26 copy number. Such systems can include DNA microarrays, RNA expression arrays, any ELISA detection system and/or any Western blotting detection system.
[0322] The information determined in the determination system can be read by the storage module 30. As used herein the "storage module" is intended to include any suitable computing or processing apparatus or other device configured or adapted for storing data or information. Examples of electronic apparatus suitable for use with the present invention include stand-alone computing apparatus, data telecommunications networks, including local area networks (LAN), wide area networks (WAN), Internet, Intranet, and Extranet, and local and distributed computer processing systems. Storage modules also include, but are not limited to: magnetic storage media, such as floppy discs, hard disc storage media, magnetic tape, optical storage media such as CD-ROM, DVD, electronic storage media such as RAM, ROM, EPROM, EEPROM and the like, general hard disks and hybrids of these categories such as magnetic/optical storage media. The storage module is adapted or configured for having recorded thereon expression level or protein level information. Such information may be provided in digital form that can be transmitted and read electronically, e.g., via the Internet, on diskette, via USB (universal serial bus) or via any other suitable mode of communication.
[0323] As used herein, "stored" refers to a process for encoding information on the storage module. Those skilled in the art can readily adopt any of the presently known methods for recording information on known media to generate manufactures comprising expression level information.
[0324] In one embodiment the reference data stored in the storage module to be read by the comparison module is e.g., expression data obtained from a population of non-cancer subjects, a population of cancer subjects or expression data obtained from the same subject at a prior time point using the measuring module 40.
[0325] The "comparison module" 80 can use a variety of available software programs and formats for the comparison operative to compare expression data determined in the measuring module to reference samples and/or stored reference data. In one embodiment, the comparison module is configured to use pattern recognition techniques to compare information from one or more entries to one or more reference data patterns. The comparison module may be configured using existing commercially-available or freely-available software for comparing patterns, and may be optimized for particular data comparisons that are conducted. The comparison module provides computer readable information related to normalized expression of BLM and/or FANCI, the 15Q26 copy number in an individual, efficacy of treatment in an individual, and/or method for treating an individual.
[0326] The comparison module, or any other module of the invention, may include an operating system (e.g., UNIX) on which runs a relational database management system, a World Wide Web application, and a World Wide Web server. World Wide Web application includes the executable code necessary for generation of database language statements (e.g., Structured Query Language (SQL) statements). Generally, the executables will include embedded SQL statements. In addition, the World Wide Web application may include a configuration file which contains pointers and addresses to the various software entities that comprise the server as well as the various external and internal databases which must be accessed to service user requests. The Configuration file also directs requests for server resources to the appropriate hardware--as may be necessary should the server be distributed over two or more separate computers. In one embodiment, the World Wide Web server supports a TCP/IP protocol. Local networks such as this are sometimes referred to as "Intranets. An advantage of such Intranets is that they allow easy communication with public domain databases residing on the World Wide Web (e.g., the GenBank or Swiss Pro World Wide Web site). Thus, in a particular preferred embodiment of the present invention, users can directly access data (via Hypertext links for example) residing on Internet databases using a HTML interface provided by Web browsers and Web servers.
[0327] The comparison module provides a computer readable comparison result that can be processed in computer readable form by predefined criteria, or criteria defined by a user, to provide a content-based in part on the comparison result that may be stored and output as requested by a user using an output module 110.
[0328] The content based on the comparison result, may be an expression value compared to a reference showing the susceptibility or nonsusceptibility of treatment with platinum-comprising therapy or anthracycline-compri sing therapy.
[0329] In various embodiments of the invention, the content based on the comparison result is displayed on a computer monitor 120. In various embodiments of the invention, the content based on the comparison result is displayed through printable media 130. The display module can be any suitable device configured to receive from a computer and display computer readable information to a user. Non-limiting examples include, for example, general-purpose computers such as those based on Intel PENTIUM-type processor, Motorola PowerPC, Sun UltraSPARC, Hewlett-Packard PA-RISC processors, any of a variety of processors available from Advanced Micro Devices (AMD) of Sunnyvale, Calif., or any other type of processor, visual display devices such as flat panel displays, cathode ray tubes and the like, as well as computer printers of various types.
[0330] In one embodiment, a World Wide Web browser is used for providing a user interface for display of the content based on the comparison result. It should be understood that other modules of the invention can be adapted to have a web browser interface. Through the Web browser, a user may construct requests for retrieving data from the comparison module. Thus, the user will typically point and click to user interface elements such as buttons, pull down menus, scroll bars and the like conventionally employed in graphical user interfaces.
[0331] The present invention therefore provides for systems (and computer readable media for causing computer systems) to perform methods for selecting treatment of cancer in an individual.
[0332] Systems and computer readable media described herein are merely illustrative embodiments of the invention for detecting BLM and/or FANCI expression, or 15Q26 copy number in an individual, and are not intended to limit the scope of the invention. Variations of the systems and computer readable media described herein are possible and are intended to fall within the scope of the invention.
[0333] The modules of the machine, or those used in the computer readable medium, may assume numerous configurations. For example, function may be provided on a single machine or distributed over multiple machines.
[0334] In some cases, a computing system provided herein can include computer-executable instructions or a computer program (e.g., software) containing computer-executable instructions for formatting an output providing an indication BLM and/or FANCI expression, 15q26 copy number, or a likelihood that a cancer patient will respond to a particular cancer treatment regimen (e.g., a regimen as described above), or a combination of these items. In some cases, a computing system provided herein can include computer-executable instructions or a computer program (e.g., software) containing computer-executable instructions for determining a desired cancer treatment regimen for a particular patient based at least in part on increased expression of BLM and/or FANCI, or 15q26 copy number gain.
[0335] In some cases, a computing system provided herein can include a pre-processing device configured to process a sample (e.g., cancer cells) such that a SNP array-based assay or sequencing-based assay can be performed. Examples of pre-processing devices include, without limitation, devices configured to enrich cell populations for cancer cells as opposed to non-cancer cells, devices configured to lyse cells and/or extract genomic nucleic acid, and devices configured to enrich a sample for particular genomic DNA fragments.
[0336] The following numbered paragraphs provide some embodiments and aspects of the invention. [0337] 1. An assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: [0338] subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; [0339] comparing the BLM and FANCI expression to a reference value; and [0340] selecting a platinum-comprising cancer therapy for the subject when the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI, or selecting a non-platinum-comprising cancer therapy for the subject when the BLM and FANCI expression is not increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is not effective in subjects whose cancer does not have increased BLM and FANCI expression compared to the reference value. [0341] 2. The assay of paragraph 1, further comprising: [0342] assaying the BRCA1 and/or BRCA2 status of the subject; and [0343] selecting the platinum-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI and who are negative for BRCA1 and/or BRCA2 mutations. [0344] 3. The assay of paragraph 1, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. [0345] 4. The assay of paragraph 1, further comprising: [0346] assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and [0347] selecting the platinum-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when the BLM and FANCI expression is increased compared to the reference value based on the recognition that platinum-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor. [0348] 5. The assay of paragraph 1, wherein the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. [0349] 6. The assay of paragraphs 1-5, further comprising administering the selected therapy to the subject. [0350] 7. The assay of any one of the paragraphs 1-6, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0351] 8. The assay of paragraphs 1-6, wherein the reference value is BRCA1 expression in the sample and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0352] 9. The assay of paragraphs 1-6, wherein the reference value is based on at least BRCA1 gene expression in the cancer cell. [0353] 10. The assay paragraphs 1-6, wherein the reference value is based on at least one housekeeping gene expression in the cancer cell. [0354] 11. A method for selecting platinum-comprising therapy for a subject having cancer, and optionally administering the platinum-comprising therapy, the method comprising: [0355] subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; [0356] detecting the BLM and FANCI expression in the sample compared to a reference value; and [0357] selecting a platinum-comprising cancer therapy for the subject when the BLM and FANCI expression compared to a reference value is increased based on the recognition that platinum-comprising cancer therapy is effective in patients whose cancer has increased BLM and FANCI expression compared to the reference value. [0358] 12. The method of paragraph 11, further comprising administering to the subject the platinum-comprising cancer therapy when the platinum-comprising cancer therapy is selected. [0359] 13. The method of paragraph 11, wherein the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. [0360] 14. The method of paragraph 11, wherein the subject's cancer or cancer cell is known to not or determined to not express a detectable quantity of ER, PgR, and HER2 receptor. [0361] 15. The method of any one of the paragraphs 11-14, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0362] 16. The method of paragraph 11-15, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0363] 17. The method of any one of the paragraphs 11-15, wherein the reference value is based on at least BRCA1 gene expression in the cancer cell. [0364] 18. The method of any one of the paragraphs 11-15, wherein the reference value is based on at least one housekeeping gene expression in the cancer cell. [0365] 19. The method of paragraph 18, wherein the housekeeping gene is selected from beta-actin, GAPDH, RPLP0, GUS, TFRC and any combination thereof. [0366] 20. The method of paragraph 19, wherein the housekeeping gene is RPLP0, and the BLM and/or FANCI expression is increased by at least six-fold. [0367] 21. A method for selecting a non-platinum-comprising therapy, and optionally administering the non-platinum-comprising therapy, for a subject having cancer comprising: [0368] subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; [0369] detecting the BLM and FANCI expression in the sample compared to a reference value; and [0370] selecting the non-platinum-comprising cancer therapy for the subject when the BLM and FANCI expression compared to the reference value is not increased based on the recognition that non-platinum-comprising cancer therapy is effective in patients whose cancer does not have increased gene expression of BLM and FANCI compared to the reference value. [0371] 22. The method of paragraph 21, further comprising administering to the subject the non-platinum-comprising cancer therapy when non-platinum-comprising cancer therapy is selected. [0372] 23. The method of any one of the paragraphs 21-22, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0373] 24. The method of paragraph 21-23, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0374] 25. The method of any one of the paragraphs 21-24, wherein the reference value is based on at least BRCA1 gene expression in the cancer cell. [0375] 26. The method of any one of the paragraphs 21-22, wherein the reference value is based on at least one housekeeping gene expression in the cancer cell. [0376] 27. An assay for selecting a therapy for a subject having cancer, comprising: [0377] subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; [0378] comparing the BLM and FANCI expression to a reference value; and [0379] selecting an anthracycline-compri sing cancer therapy for the subject when the BLM and FANCI expression is increased compared to a reference value based on the recognition that anthracycline-comprising cancer therapy is effective in subjects whose cancer has increased expression of BLM and FANCI, or [0380] selecting a non-anthracycline-comprising cancer therapy for the subject when the BLM and FANCI expression is not increased compared to a reference value based on the recognition that anthracycline-comprising cancer therapy is not effective in subjects whose cancer does not have increased BLM expression compared to a reference value. [0381] 28. The assay of paragraph 27, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0382] 29. A method for selecting an anthracycline-comprising cancer therapy for a subject having cancer and determined to be negative for BRCA1 and/or BRCA2 mutations, comprising: [0383] subjecting a sample comprising a cancer cell taken from the subject to an analysis for BLM and FANCI expression; [0384] comparing the BLM and FANCI expression to a reference value; and [0385] selecting the anthracycline-comprising cancer therapy for the subject when the BLM and FANCI expression compared to the reference value is increased based on the recognition that anthracycline-comprising cancer therapy is effective in patients whose cancer has increased expression of BLM and FANCI compared to the reference value. [0386] 30. The method of paragraph 29, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0387] 31. A method of treating cancer in a human subject, comprising: [0388] detecting BLM and FANCI expression in a sample comprising a cancer cell taken from the human subject; and [0389] comparing the BLM and FANCI expression to a reference value; and [0390] administering a platinum-comprising cancer therapy to the human subject wherein an increase of BLM and FANCI expression compared to the reference value is detected. [0391] 32. The method of paragraph 31, wherein the human subject's cancer or cancer cell is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. [0392] 33. The method of paragraph 31, wherein the cancer is selected from breast, ovarian, and lung cancers. [0393] 34. The method of paragraph 31-33, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0394] 35. The method of any one of the paragraphs 31-33, wherein the reference value is based on BRCA1 gene expression in the cancer cell. [0395] 36. The method of any one of the paragraphs 31-33, wherein the reference value is based on a housekeeping gene expression in the cancer cell. [0396] 37. A method of treating cancer in a human subject, comprising: [0397] detecting BLM and FANCI expression in a sample comprising a cancer cell taken from the human subject; and [0398] comparing the BLM and FANCI expression to a reference value; and [0399] administering an anthracycline-comprising cancer therapy to the human subject wherein an increase of BLM and FANCI expression compared to the reference value is detected. [0400] 38. The method of paragraph 37, wherein the human subject's cancer or cancer cell is known to not or is determined to not to express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. [0401] 39. The method of paragraph 37, wherein the cancer is selected from breast, ovarian, and lung cancers. [0402] 40. The method of paragraph 37-39, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0403] 41. The method of any one of the paragraphs 37-39, wherein the reference value is based on BRCA1 gene expression in the cancer cell. [0404] 42. The method of any one of the paragraphs 37-39, wherein the reference value is based on a housekeeping gene expression in the cancer cell. [0405] 43. A method for assessing responsiveness of a cancer cell to cancer therapy, comprising: [0406] assaying, in a cancer cell or mRNA derived therefrom, BLM and FANCI expression; and [0407] comparing said BLM and FANCI expression to a reference value, wherein the cancer cell is assessed as responsive to a platinum-comprising therapy if the BLM and FANCI expression is increased compared to the reference value, or wherein the cancer cell is assessed as poorly or not responsive to platinum-comprising cancer therapy cancer if the BLM and FANCI expression is not increased. [0408] 44. The method of paragraph 43, wherein the step of assaying comprises: [0409] contacting the cancer cell or mRNA derived therefrom with at least one detectably labeled probe capable of specifically binding to BLM mRNA, at least one detectably probe capable of specifically binding to FANCI, at least one detectably labeled probe capable of specifically binding to BRCA1 and/or at least one housekeeping gene; and [0410] measuring the expression of BLM and FANCI compared to the BRCA1 and/or the at least one housekeeping gene. [0411] 45. The method of paragraph 43-44, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0412] 46. A method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: [0413] determining, in a cancer cell from the cancer patient, BLM and FANCI expression; and [0414] correlating the expression to a reference value, wherein when the expression is increased the patient is predicted to respond well to a cancer treatment regimen comprising platinum or anthracycline. [0415] 47. The method of paragraph 46, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0416] 48. A method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: [0417] determining, in a cancer cell or mRNA derived therefrom from said cancer patient, BLM and FANCI expression; and [0418] correlating the expression to a reference value, wherein when the expression is not increased the patient is predicted to respond poorly to a cancer treatment regimen comprising platinum or anthracycline. [0419] 49. The method of paragraph 48, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0420] 50. A method of treating cancer, comprising: [0421] assaying, in a cancer cell from a cancer patient or mRNA obtained therefrom, the BLM and FANCI expression compared to a reference value; and [0422] administering to the cancer patient a cancer treatment regimen comprising platinum or anthracycline if the BLM and FANCI expression is increased compared to the reference value. [0423] 51. The method of paragraph 50, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0424] 52. Use of platinum comprising cancer therapy for treating a cancer patient that has been determined to have a tumor comprising cancer cells wherein BLM and FANCI expression is increased compared to a reference value.
[0425] 53. The use of paragraph 52, wherein the cancer patient has been determined to be negative for BRCA1 and/or BRCA2 mutations. [0426] 54. The use of paragraph 52, wherein the cancer patient's cancer or cancer cell is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. [0427] 55. The use of paragraph 52-54, wherein the reference value is BRCA1 expression in the sample, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0428] 56. A system for determining responsiveness of a cancer cell to platinum-comprising therapy from a cancer cell of a cancer patient, comprising: [0429] a sample analyzer configured to produce a signal for the mRNA from each one of BLM and FANCI from a cancer cell sample of a cancer patient; and [0430] a computer sub-system programmed to calculate, based on the mRNA whether the signal is greater or not than a reference value. [0431] 57. The system of paragraph 56, wherein said computer sub-system is programmed to compare the mRNA to determine [0432] a likelihood of responsiveness of said cancer cell to platinum-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a platinum-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the platinum-comprising therapy if the BLM and FANCI expression is not increased; or [0433] a likelihood of responsiveness of said cancer cell to anthracycline-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a anthracycline-comprising therapy if the BLM and FANCI expression is increased and as unlikely to respond to the anthracycline-comprising therapy if the BLM and FANCI expression is not increased. [0434] 58. The system of paragraph 56-57, wherein the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0435] 59. A computer program product embodied in a computer readable medium that, when executing on a computer, performs steps comprising: [0436] detecting the BLM and FANCI gene expression in sample comprising a cancer cell from a cancer patient; and [0437] comparing the BLM and FANCI expression to a reference value. [0438] 60. The computer program of paragraph 59, wherein the reference value is BRCA1 expression, and the BLM and/or FANCI expression is increased by at least two-fold compared to BRCA1 expression. [0439] 61. A diagnostic kit for detecting a likelihood of a cancer patient to respond to platinum- or anthracycline-comprising comprising cancer therapy, comprising: [0440] no more than 10 probes comprising a combination of detectably labeled probes or primers for BLM and FANCI, and optionally for BRCA1 and/or at least one housekeeping gene; and [0441] the computer program product of Paragraph 59. [0442] 62. Use of a plurality of oligonucleotides comprising no more than 10 oligonucleotides capable of hybridizing to BLM and FANCI, and optionally to BRCA1 and/or at least one housekeeping gene, in a diagnostic kit for determining an increased likelihood that a cancer patient will respond to cancer treatment regimen comprising a platinum and/or anthracycline. [0443] 63. The use, method or assay of any one of the preceding paragraphs, wherein said anthracycline is epirubincin or doxorubicin. [0444] 64. The use, method or assay of any one of the preceding paragraphs, wherein said platinum comprising cancer therapy comprises cisplatinum or cis-diamminedichloroplatinum, phenanthriplatin, carboplatin, oxaliplatin, or a platinum complex that is activated by ultraviolet A light. [0445] 65. An assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: [0446] assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; [0447] comparing the chromosome 15q26 copy number to a reference value; and [0448] selecting a platinum-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or selecting a non-platinum-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss. [0449] 66. The assay of paragraph 65, further comprising: [0450] assaying the BRCA1 and/or BRCA2 status of the subject; and [0451] selecting the platinum-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and there is a chromosome 15q26 copy number gain based on the recognition that platinum-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain who are negative for BRCA1 and/or BRCA2 mutations. [0452] 67. The assay of paragraph 65, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. [0453] 68. The assay of paragraph 65, further comprising: [0454] assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and [0455] selecting the platinum-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when there is a chromosome 15q26 copy number gain based on the recognition that platinum-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor. [0456] 69. The assay of paragraph 65, wherein the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. [0457] 70. The assay of paragraphs 65-69, further comprising administering the selected therapy to the subject. [0458] 71. The assay of paragraphs 65-71, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0459] 72. The assay of paragraphs 65-71, wherein the reference value is chromosome 15 centromere copy number in the sample. [0460] 73. An assay for selecting a therapy for a subject having cancer, and optionally administering the therapy, the assay comprising: [0461] assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; [0462] comparing the chromosome 15q26 copy number to a reference value; and [0463] selecting an anthracycline-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or selecting a non-anthracycline-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss. [0464] 74. The assay of paragraph 73, further comprising: [0465] assaying the BRCA1 and/or BRCA2 status of the subject; and [0466] selecting the anthracycline-comprising cancer therapy for the subject when the subject is negative for BRCA1 and/or BRCA2 mutations, and there is a chromosome 15q26 copy number gain based on the recognition that anthracycline-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain who are negative for BRCA1 and/or BRCA2 mutations. [0467] 75. The assay of paragraph 73, the subject is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. [0468] 76. The assay of paragraph 73, further comprising: [0469] assaying the estrogen receptor (ER), progesterone receptor (PgR), and HER2 receptor status of the subject's cancer; and [0470] selecting the anthracycline-comprising cancer therapy for the subject when the subject's cancer does not express a detectable quantity of ER, PgR, and HER2 receptor, and when there is a chromosome 15q26 copy number gain based on the recognition that anthracycline-comprising cancer therapy is effective in subjects who have a chromosome 15q26 copy number gain and whose cancer does not express a detectable quantity of ER, PgR, and HER2 receptor. [0471] 77. The assay of paragraph 73, wherein the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. [0472] 78. The assay of paragraphs 73-77, further comprising administering the selected therapy to the subject. [0473] 79. The assay of paragraphs 73-78, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0474] 80. The assay of paragraphs 73-79, wherein the reference value is chromosome 15 centromere copy number in the sample. [0475] 81. A method of treating cancer in a human subject, comprising: [0476] detecting a chromosome 15q26 copy number in a sample comprising a cancer cell taken from the subject; [0477] comparing the chromosome 15q26 copy number to a reference value; and [0478] administering an platinum-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or administering a non-platinum-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss. [0479] 82. The method of paragraphs 81, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0480] 83. The method of paragraphs 81-82, wherein the reference value is chromosome 15 centromere copy number in the sample. [0481] 84. The method of paragraph 81-83, wherein the subject's cancer is known to not express a detectable quantity of ER, PgR, and HER2 receptor. [0482] 85. A method of treating cancer in a human subject, comprising: [0483] detecting a chromosome 15q26 copy number in a sample comprising a cancer cell taken from the subject; [0484] comparing the chromosome 15q26 copy number to a reference value; and [0485] administering an anthracycline-comprising cancer therapy for the subject if there is a chromosome 15q26 copy number gain compared to the reference value, or administering a non-anthracycline-comprising cancer therapy for the subject if there is not a chromosome 15q26 copy number gain, or if there is a chromosome 15q26 copy number loss. [0486] 86. The method of paragraphs 85, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0487] 87. The method of paragraphs 85-86, wherein the reference value is chromosome 15 centromere copy number in the sample. [0488] 88. The method of paragraph 85-88, wherein the subject's cancer or cancer cell is known to not or is determined to not express a detectable quantity of ER, PgR, and HER2 receptor. [0489] 89. A method for assessing responsiveness of a cancer cell to a cancer therapy, and optionally administering the cancer therapy, comprising: [0490] assaying a sample comprising a cancer cell taken from the subject for a chromosome 15q26 copy number; [0491] and comparing the chromosome 15q26 copy number to a reference value, wherein the cancer cell is assessed as responsive to a platinum-comprising therapy if there is a chromosome 15q26 copy number gain compared to the reference value, or wherein the cancer cell is assessed as poorly or not responsive to platinum-comprising cancer therapy cancer if there is not a chromosome 15q26 copy number gain or if there is a chromosome 15q26 copy number loss. [0492] 90. The method of paragraph 89, wherein the reference value is copy number of chromosome 15. 91. The method of paragraph 89, wherein the cancer is selected from breast cancer, ovarian cancer and lung cancer. [0493] 92. The method of paragraph 89, further comprising administering the platinum-comprising therapy if there is a chromosome 15q26 copy number gain. [0494] 93. A method of predicting a cancer patient's response to a cancer treatment regimen comprising platinum or anthracycline, comprising: [0495] determining, in a cancer cell from the cancer patient, chromosome 15q26 copy number; and [0496] correlating the chromosome 15q26 copy number to a reference value, [0497] wherein when there is a chromosome 15q26 copy number gain, the patient is predicted to respond well to a cancer treatment comprising platinum or anthracycline, or wherein when there is not a chromosome 15q26 copy number gain or a chromosome 15q26 copy number loss, the patient is predicted respond poorly to a cancer treatment comprising platinum or anthracycline. [0498] 94. The method of paragraph 93, wherein the reference value is chromosome 15 centromere copy number in the sample. [0499] 95. Use of platinum comprising cancer therapy for treating a cancer patient that has been determined to have a tumor comprising cancer cells wherein a chromosome 15q26 copy gain is detected compared to a reference value. [0500] 96. The use of paragraph 95, wherein the cancer patient is known to be or is determined to be negative for BRCA1 and/or BRCA2 mutations. [0501] 97. The use of paragraph 95, wherein the cancer patient's cancer or cancer cell is known to not or is determined to not express detectable quantities of estrogen receptor (ER), progesterone receptor (PgR) and HER2 receptor. [0502] 98. The use of paragraph 95-97, wherein the reference value is chromosome 15 centromere copy number in the sample. [0503] 99. A system for determining responsiveness of a cancer cell to platinum-comprising therapy from a cancer cell of a cancer patient, comprising: [0504] a sample analyzer configured to produce a signal for chromosome 15q26 copy number from a cancer cell sample of a cancer patient; and [0505] a computer sub-system programmed to calculate, based on the mRNA whether the signal is greater or not than a reference value. [0506] 100. The system of paragraph 99, wherein said computer sub-system is programmed to compare the mRNA to determine [0507] a likelihood of responsiveness of said cancer cell to platinum-comprising cancer therapy and/or or a anthracycline-comprising cancer therapy based on an algorithm that classifies the patient as likely to respond to a platinum-comprising therapy if there is a chromosome 15q26 copy number gain and as unlikely to respond to the platinum-comprising therapy if there is not a chromosome 15q26 copy number gain or if there is a chromosome 15q26 copy number loss. [0508] 101. The system of paragraph 99-100, wherein the reference value is chromosome 15 centromere copy number in the sample. [0509] 102. A computer program product embodied in a computer readable medium that, when executing on a computer, performs steps comprising: [0510] detecting chromosome 15q26 copy number in sample comprising a cancer cell from a cancer patient; and [0511] comparing the chromosome 15q26 copy number to a reference value. [0512] 103. The computer program of paragraph 102, wherein the reference value is chromosome 15 centromere copy number in the sample. [0513] 104. A diagnostic kit for detecting a likelihood of a cancer patient to respond to platinum- or anthracycline-comprising cancer therapy, comprising:
[0514] no more than 10 probes comprising a combination of detectably labeled probes or primers for chromosome 15q26, and optionally for chromosome 15 centromere; and [0515] the computer program product of Paragraph 102. [0516] 105. Use of a plurality of oligonucleotides comprising no more than 10 oligonucleotides capable of hybridizing to chromosome 15q26, and optionally for chromosome 15 centromere, in a diagnostic kit for determining an increased likelihood that a cancer patient will respond to cancer treatment regimen comprising a platinum and/or anthracycline. [0517] 106. The use, method or assay of paragraphs 65-105, wherein said anthracycline is epirubincin or doxorubicin. [0518] 107. The use, method or assay paragraphs 65-105, wherein said platinum comprising cancer therapy comprises cisplatinum or cis-diamminedichloroplatinum, phenanthriplatin, carboplatin, oxaliplatin, or a platinum complex that is activated by ultraviolet A light.
[0519] The following examples are provided to better illustrate the claimed invention and are not to be interpreted as limiting the scope of the invention. To the extent that specific materials are mentioned, it is merely for purposes of illustration and is not intended to limit the invention. One skilled in the art may develop equivalent means or reactants without the exercise of inventive capacity and without departing from the scope of the invention.
Example 1
[0520] Two clinical trials with cisplatin given to women with triple negative breast cancer were evaluated.
[0521] Cisplatin-1 (DFHCC 04-183; Silver et al., JCO 2010): 28 TNBC patients received neoadjuvant cisplatin as single agent. 10 (36%) achieved at least 90% reduction in tumor size. Good quality RNA and DNA were acquired from 21 tumors. Copy number and gene expression were assayed using AFFYMETRIX arrays.
[0522] Cisplatin-2 (DFHCC 06-202; Ryan et al., JCO 2009): 51 TNBC patients received neoadjuvant cisplatin in combination with the angiogenesis inhibitor bevacizumab. [0523] 44 patients completed therapy. 17 (39%) achieved at least 90% reduction in tumor size. Frozen DNA samples were acquired from 24 tumors, mRNA from 21. Copy number and gene expression assayed using AFFYMETRIX arrays.
Example 2
[0524] In order to find regions or genes where gain or loss were significantly associated with sensitivity of resistance cisplatin, .about.40,000 SNP probes were used in Cisplatin-1 trial and .about.330,000 SNP probes were used in Cisplatin-2 trial to determine sensitivity and resistance. GISTIC identified chromosomal regions that were significantly enriched in either group. Gene copy number comparison showed that genes significantly gained or lost between sensitive and resistant cases, in both cohorts. Comparison to gene expression showed overlap with genes showing differential expression between sensitive and resistant cancers. See e.g., FIGS. 1-3.
Example 3
[0525] In a comparison to gene expression data, the inventors found genes that show both significant gain and higher expression in sensitive sample. In Cisplatin-1 trial, there were 21 cases; Cisplatin-2 trial, there were 21 cases. In an ovarian cancer trial (OV-01 trial; Ahmed et al., Cancer Cell 2007), the two arm carboplatinum had 15 cases and paclitaxel had 19 cases. A leave-one-out analysis of the gene expression data was performed from both platinum breast cancer trials and the carboplatinum arm of OVO1. See e.g., FIGS. 4-5.
Example 4
[0526] In evaluating Hess et al., JCO 2006; Popovici et al., Breast Cancer Res 2010; and Andre et al., Clin Cancer Res 2009, expression of BLM or FANCI is not found to be associated with response to multi-agent chemotherapy regimens in TNBC. See e.g., FIG. 6.
Example 5
[0527] In evaluating Desmedt et al., JCO 2011, BLM is associated with response to single agent epirubicin. See e.g., FIG. 7.
Example 6
Chromosome 15q26 Copy Number and Overexpression of BLM and FANCI Predict for Cisplatin Sensitivity
[0528] To elucidate if particular genomic aberrations may affect cancer cells sensitivity to cisplatin, we compared the tumor DNA copy number profiles in sensitive versus resistant TN breast tumors in the two separate cisplatin clinical trials. We identified several sites of DNA copy number change in each trial, but only a short 15 megabase (MB) region on chromosome 15q26 was significantly different between responders and nonresponders in both trials. In addition to SNP profiles, we have analyzed the pretreatment tumor samples for both Cisplatin1 and Cisplatin2 for mRNA expression profiles on Affymetrix microarrays. In addition, gene expression array data was publically available from the carboplatin arm of a previously published two-arm trial of carboplatin and paclitaxel in serous ovarian cancer[31]. To determine if any specific genes were consistently differentially expressed in platinum sensitive compared to resistant tumors, we performed a leave-one-out analysis in each trial, where we in each round compared the sensitive to the resistant tumors, and scored how many genes showed a significant association. We found that only 3 genes were consistently significantly different in all three platinum-based trials: MCM2, BLM, and FANCI. Interestingly, FANCI and BLM are both located on 15q26.1 and the proteins have been shown to localize to sites of DNA damage. This shows that using two different methods measuring two different biomolecules, high levels of BLM and FANCI DNA and mRNA is associated with greater sensitivity to platinum-based chemotherapy.
[0529] To investigate if higher expression of BLM and FANCI were specifically associated with platinum chemotherapy response, we analyzed the gene expression in the paclitaxel arm of the ovarian trial[31], and across the TN breast cancer subset of three neoadjuvant cohorts that received taxane-containing combination chemotherapy[32-34].
[0530] In the ovarian paclitaxel trial and in the TN subset of the taxane-containing multidrug trials there was no association between either BLM or FANCI and therapy response. These data suggests that high expression of BLM and FANCI are possibly specifically associated with sensitivity to DNA damaging agents like the platinum salts, but not with response to chemotherapeutics that alter microtubules such as taxanes.
[0531] We combined the mRNA results into a predictive gene signature: (BRCA1 mRNA)/(average of FANCI and BLM mRNA). ROC analysis demonstrates that this 3gene mRNA signature is significantly associated with cisplatin response in both TN cisplatin trials (FIG. 10A). A prediction model combining NtAI and the 3 gene signature improved the prediction of cisplatin sensitivity (FIG. 10B). Based on the combined model, 25% of cases are "biomarker positive" with predictive accuracy of 0.86, a positive predictive value of 0.89, a negative predictive value of 0.85, sensitivity of 0.67, and specificity of 0.96 with a p-value of 0.00016.
Example 7
Optimization of BRCA1-BLM-FANCI mRNA Measurement
[0532] Several different RT-PCR primer pair assays will be designed to measure mRNA expression for each of the three genes. Data from the Cisplatin2 trial comparing the BRCA1, BLM, and FANCI mRNA levels on microarray to mRNA levels as measured by quantitative real-time PCR and found good correlation. The 3-gene predictive model of BRCA1/(avg of BLM+FANCI) using the RT-PCR measurements performed very well for prediction of cisplatin response (FIG. 11).
Example 8
[0533] Platinum salts are effective treatment and standard therapy for a number of cancer types including serous ovarian carcinoma, high grade urothelial carcinoma, pancreatic adenocarcinoma, glioblastoma multiforme, and lung squamous cell carcinoma. Breast adenocarcinomas arising in women carrying BRCA1 or BRCA2 mutations are also sensitive to cisplatin chemotherapy (Byrski, 2008). BRCA1 is a tumor suppressor that plays important roles in several aspects of maintenance of genome integrity. Complete absence of functional BRCA1, as occurs in tumors of BRCA1 mutation carriers with loss of the wild-type allele, leads to defective error-free homologous recombination-type double strand break repair. These BRCA1-/- tumors are sensitive to inhibitors of PARP1 (ref) whereas, so far, few or no sporadic BRCA wild-type breast cancers have responded to these agents. Recent studies have shown that in addition to double strand break repair, BRCA1 also plays an important role in response to replication stress and repair of stalled or collapsed replication forks (Pathania, 2011). Preliminary evidence suggests the replication repair functions of BRCA1 may be haploinsufficient in BRCA1 heterozygous cells (Pathania, unpublished data). The platinum salts generate interstrand and intrastrand crosslinks that distort DNA and lead to stalled replication forks. If these stalled forks collapse, double strand breaks will results. It is possible that several BRCA1-dependent pathways are involved in platinum-induced DNA damage response including damage recognition and lesion excision, suppression of translesion synthesis, checkpoint activation, and repair of DS breaks after fork collapse.
[0534] The activity of cisplatin has recently been extended to estrogen, progesterone, and HER2 receptor negative sporadic breast cancers (triple-negative breast cancers, TNBC)(Silver, 2010; Birkbak, 2012; Ryan, 2009). When carboplatin was added to anthracycline and taxane chemotherapy for treatment of TNBC, the response rate was higher but resulted in notably greater toxicity. Predictors of tumor response to platinum salts are needed to determine which patients may derive the greatest benefit from the addition of platinum. Previous molecular studies have shown the platinum-sensitive TNBC and serous ovarian cancers carry high levels of genomic rearrangements and chromosomal allelic imbalance, suggesting these cancers may share similar defects in DNA repair, which may make them particularly sensitive to platinum chemotherapy (Birkbak, 2012). Many of the platinum-sensitive sporadic TNBCs have promoter methylation and reduction in the expression of BRCA1 (Birkbak, 2012; Silver, 2010). These result suggests that platinum sensitivity may be related to a functional defect that occurs when there is insufficient BRCA1 levels, raising the possibility that defects in a haploinsufficient function of BRCA1, such as replication-dependent stalled fork repair, may be indicative of sensitivity to interstrand cross-linkers such as the platinum salts.
[0535] To further explore and define other specific molecular alterations that might be associated with cisplatin sensitivity, we took and integrated genomic approach combining differential analysis of gene expression and DNA copy number in cisplatin sensitive compared to cisplatin resistant triple negative breast cancers. We identify two genes, the Bloom helicase (BLM) and Fanconi anemia complementation group I (FANCI), that have both increased DNA copy number at chr 15q26 and concordant mRNA overexpression in the TNBC with increased sensitivity to cisplatin therapy. In vitro modulation of BLM and FANCI expression suggest these two genes play a functional role and promote DNA damage and sensitivity to platinum salts, but have no effect on taxane sensitivity.
[0536] Identification of genes critical for tumor response to specific chemotherapy drugs is a challenge, but important for tailoring therapy and avoiding unnecessary toxicity in patients. Integrated genomic approaches that combine DNA copy number analysis with gene expression profile analysis has successfully indicated genomic alterations associated with chemotherapy resistance and tumor response (Li, 2010). We have previously published the results of two trials of cisplatin chemotherapy in TNBC in which presurgical treatment with cisplatin resulted in greater than 90% reduction in tumor volume in 36% and 39% of patients respectively (Silver, 2010; Ryan, 2009). Molecular inversion probe SNP copy number profiles of pretreatment tumors from both cisplatin TNBC trials were reported previously (Birkbak, 2012). Gene expression profiles from the pretreatment tumor samples from the first cisplatin TNBC trial were also reported previously (Silver, 2010). For this study, we generated gene expression profiles from the pretreatment tumor biopsies from the second cisplatin TNBC trial.
[0537] To determine genes whose expression is significantly and robustly associated with response to cisplatin, we performed a leave-one-out differential gene expression analysis in each trial, comparing the gene expression of tumors resistant to cisplatin to tumors that were sensitive to cisplatin. For each leave-one-out round, one sample was removed and all genes significantly associated with response were determined. The direction of association for each gene (higher in sensitive vs. lower in sensitive) was also noted. Permutation testing of this gene expression comparison exercise indicated that a gene must be present in 85% or more rounds to be significant. This analysis identified only 12 genes whose expression was significantly associated with platinum response, in the same direction, in at least 85% of all rounds in both cisplatin TNBC cohorts.
[0538] We performed next a similar leave one out comparison analysis of the DNA copy numbers of cisplatin sensitive vs. cisplatin resistant TNBC tumors in the two trials. Permutation testing of the DNA copy number LOO analysis indicated that a gene must be present in 50% or more LOO rounds to be significant. This analysis identified 234 genes from four different chromosomes with differential copy number associated with cisplatin response in at least 50% of all rounds in both cisplatin TNBC cohorts.
[0539] Only two genes were identified in both the DNA copy number and gene expression leave-one-out analyses for association with platinum sensitivity, Bloom helicase (BLM) and Fanconi anemia complementation group I (FANCI), both located at chromosome 15q26. The DNA copy number of the 15q26 region containing these two genes was significantly higher in the cisplatin-sensitive tumors in both TNBC cohorts (p=0.00236 and p=0.0107, respectively. Similarly, cisplatin sensitive tumors had significantly higher BLM expression in both TNBC cohorts (cisplatin-1, mean log 2 expression, 8.26 versus 7.54, p=0.00278; cisplatin-2, mean log 2 expression, 10.67 versus 9.41, p=0.00745; FIG. 1C-D). FANCI expression was higher in the cisplatin-sensitive TNBCs in both cohorts (cisplatin-1, mean log 2 expression 7.87 versus 6.80, p=0.00356; cisplatin-2, mean log 2 expression 10.74 versus 9.77, p=0.0125).
[0540] The gene expression of BLM and FANCI as measured by Affymetrix U133 array was validated by RT-PCR in the same samples and the results showed good correlation (BLM, r=0.866; FANCI, r=0.733). BLM (and FANCI) are reported to have post-translational mechanisms of protein regulation. To determine if overexpression of BLM and FANCI mRNA also results in increased expression of the protein, we measured BLM (and FANCI) protein by western blot analysis in protein extracts from a series of frozen breast tumor samples for which mRNA gene array expression levels were known (Lu, 2008). BLM normalized to actin (and FANCI) showed good correlation between mRNA expression levels and protein expression levels (r=0.70).
[0541] BRCA1 as measured by RT-PCR was identified in our previous studies as significantly associated with cisplatin resistance (Birkbak, 2012; Silver, 2010). In contrast to BLM and FANCI, BRCA1 expression as measured by microarray was poorly correlated with expression as measured by RT-PCR (r=xxx); the microarray BRCA1 probe performance was especially poor in the data from first cisplatin TNBC cohort. Therefore, we used the RT-PCR expression data to test the association of the ratio of BRCA1/average (BLM+FANCI) expression and cisplatin response. The ratio was significantly higher in the cisplatin sensitive tumors in both cohorts (cisplatin-1, median x vs y, p=0.023; cisplatin-2, median 7.69 vs 4.07, p=0.0016).
[0542] To validate these specific gene associations with platinum response, we tested a publically available gene expression data set from a serous ovarian cancer trial of either carboplatin monotherapy or paclitaxel monotherapy and sought associations with response to the therapy received. The average expression of BLM and FANCI was significantly higher in the carboplatin-sensitive ovarian cancers (median 5.46 versus 4.50, p=0.026). The ratio of BLM+FANCI/BRCA1 was also significantly higher in carboplatin sensitive ovarian cancers (median 1.29 vs 0.42, p=0.026). Interestingly, the association of BLM and FANCI with paclitaxel response was not significant and the trend was in the opposite direction (median 5.17 versus 4.79, p=0.27).
[0543] We also showed protein abundance of BRCA1, BLM, FANCI, and Cyclin A was measured by Western blot analysis in protein extracts from a panel of breast cancer cell lines. The bands were quantitated by densitometry and displayed in bar plots. We showed the quantitation of BLM to Actin and BRCA1 to BLM ratio. Three cell lines (BT549, HCC1143, and HCC38) are normal genotype for BRCA1 and have high BLMIActin and low ratio of BRCA1/BLM. Two cell lines (MDA231 and MDA453) are also normal genotype for BRCA1 but have relatively higher expression of BRCA1/BLM and lower BLM/actin. HCC1937 and MDA436 have homozygous mutation in BRCA1 and undetectable BRCA1/BLM, and higher expression of BLM/Actin.
[0544] A panel of cell lines was evaluated for sensitivity to various treatments as indicated by colony formation assay and calculation of IC50 values. We showed a pattern of sensitivity to cisplatin, UV radiation treatment, and Parp inhibitor Olaparib across the panel of cell lines is associated with the pattern of relative expression of BRCA1/BLM and BLM/Actin. The two BRCA1 mutated cell lines and the three cell lines with low BRCA1/BLM and high BLM/actin have greater sensitivity to DNA strand cross-linker cisplatin, PARP inhibitor Olaparib, and UV irradiation. The two cell lines with low BLM and high BRCA1/BLM (MDA231, MDA453) are relatively resistant to these treatments. In contrast, there is no apparent association between BLM and BRCA1 expression with the pattern of sensitivity to the microtubule stabilizer Paclitaxel.
[0545] We also treated U2OS cells treated shRNA to BRCA1. After 1 week, the expression of BRCA1 and BLM were measured by Western blot analysis. Cells treated with the BRCA1 specific shRNA showed increased expression of BLM compared to control cells treated with shRNA to luciferase.
[0546] We further treated BT549 breast cancer cells, with inherent high levels of BLM and FANCI were treated with gene-specific siRNAs to BLM or FANCI or with a scramble control. Gene specific siRNA treatment resulted in reduced mRNA expression as determined by RT-PCR. Sensitivity to cisplatin and to paclitaxel was determined by colony formation assay to calculate the IC50. siRNA knockdown of either BLM or FANCI resulted in increased IC50 (greater resistance) to cisplatin treatment but no significant effect on sensitivity to paclitaxel.
[0547] MDA231 cells have low levels of BLM and relative resistance to cisplatin. These cells were used in experiments to assess the effect of increasing the expression of BLM. A lentivirus expression vector for HA-tagged BLM cDNA or a control vector was transfected into MDA231 cells. BLM expression was assessed by Western blot analysis for endogenous BLM or for the HA-tag in cells treated with control vector, BLM cDNA, BLM cDNA and a small molecule inhibitor of BLM helicase (BLMi), or BLM cDNA and BLM siRNA. We showed that siRNA for BLM reduced the expression of endogeneous and HA-tagged BLM whereas the small molecule inhibitor (BLMi) had no effect on protein expression of BLM.
[0548] IC50 for cisplatin was determined by colony formation assay in MDA231 breast cancer cells treated with control vector, BLM cDNA vector, BLM cDNA plus the BLM small molecule inhibitor, and BLM cDNA plus BLM siRNA. Overexpression of BLM resulted in decrease in the IC50 (greater sensitivity) to cisplatin. Consistent with the other findings, this effect was reversed by treatment with the BLM helicase inhibitor and by siRNA knockdown of BLM.
[0549] We also performed an immunofluorescence assay for markers of DNA damage (H2Ax-p and 53BP1-p) in MDA231 cells treated with control vector, BLM cDNA, BLM cDNA+BLM inhibitor, or BLM cDNA+BLM siRNA. Overexpression of BLM results in increased H2AX and 53BP1 foci in the absence of any cisplatin treatment indicating spontaneous DNA damage. This effect is even greater in cells treated with cisplatin after 4 hours. The quantitation of foci is shown in the lower bar graphs. The addition of a small molecule BLM helicase inhibitor (Bi) or siRNA to BLM (si) blocks the effect of BLM overexpression on DNA damage foci.
INCORPORATION BY REFERENCE
[0550] All publications, patents, and patent applications mentioned herein are hereby incorporated by reference in their entirety as if each individual publication, patent or patent application was specifically and individually indicated to be incorporated by reference. In case of conflict, the present application, including any definitions herein, will control.
[0551] Also incorporated by reference in their entirety are any polynucleotide and polypeptide sequences which reference an accession number correlating to an entry in a public database, such as those maintained by The Institute for Genomic Research (TIGR) on the world wide web and/or the National Center for Biotechnology Information (NCBI) on the world wide web.
EQUIVALENTS
[0552] Those skilled in the art will recognize, or be able to ascertain using no more than routine experimentation, many equivalents to the specific embodiments of the invention described herein. Such equivalents are intended to be encompassed by the following claims.
[0553] Various embodiments of the invention are described above in the Detailed Description. While these descriptions directly describe the above embodiments, it is understood that those skilled in the art may conceive modifications and/or variations to the specific embodiments shown and described herein. Any such modifications or variations that fall within the purview of this description are intended to be included therein as well. Unless specifically noted, it is the intention of the inventors that the words and phrases in the specification and claims be given the ordinary and accustomed meanings to those of ordinary skill in the applicable art(s).
[0554] The foregoing description of various embodiments of the invention known to the applicant at this time of filing the application has been presented and is intended for the purposes of illustration and description. The present description is not intended to be exhaustive nor limit the invention to the precise form disclosed and many modifications and variations are possible in the light of the above teachings. The embodiments described serve to explain the principles of the invention and its practical application and to enable others skilled in the art to utilize the invention in various embodiments and with various modifications as are suited to the particular use contemplated. Therefore, it is intended that the invention not be limited to the particular embodiments disclosed for carrying out the invention.
[0555] While particular embodiments of the present invention have been shown and described, it will be obvious to those skilled in the art that, based upon the teachings herein, changes and modifications may be made without departing from this invention and its broader aspects and, therefore, the appended claims are to encompass within their scope all such changes and modifications as are within the true spirit and scope of this invention. It will be understood by those within the art that, in general, terms used herein are generally intended as "open" terms (e.g., the term "including" should be interpreted as "including but not limited to," the term "having" should be interpreted as "having at least," the term "includes" should be interpreted as "includes but is not limited to," etc.).
Sequence CWU 1
1
114PRTHomo sapiens 1Lys Asp Glu Leu1
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017
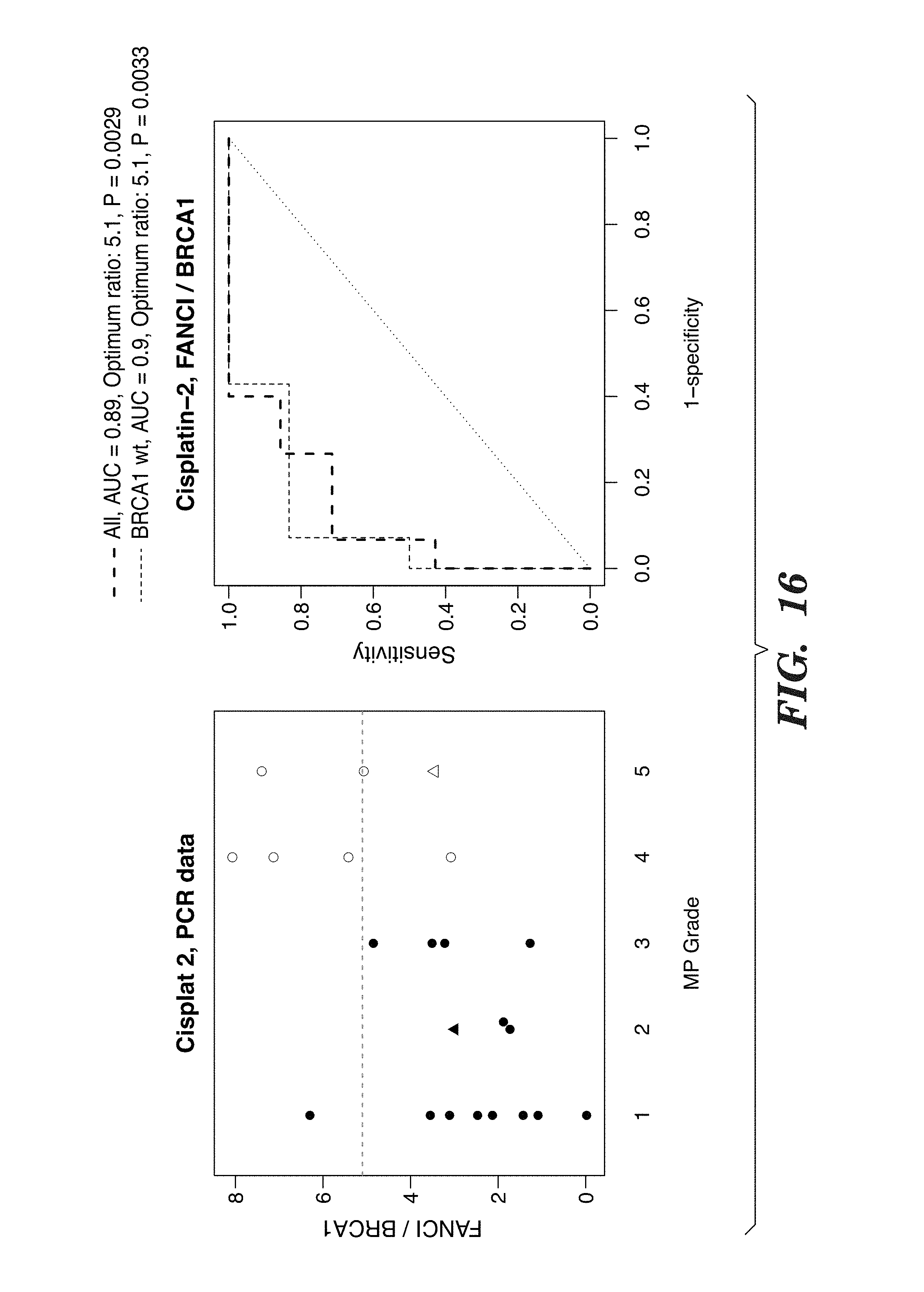
D00018

D00019

D00020

D00021

D00022

D00023

D00024
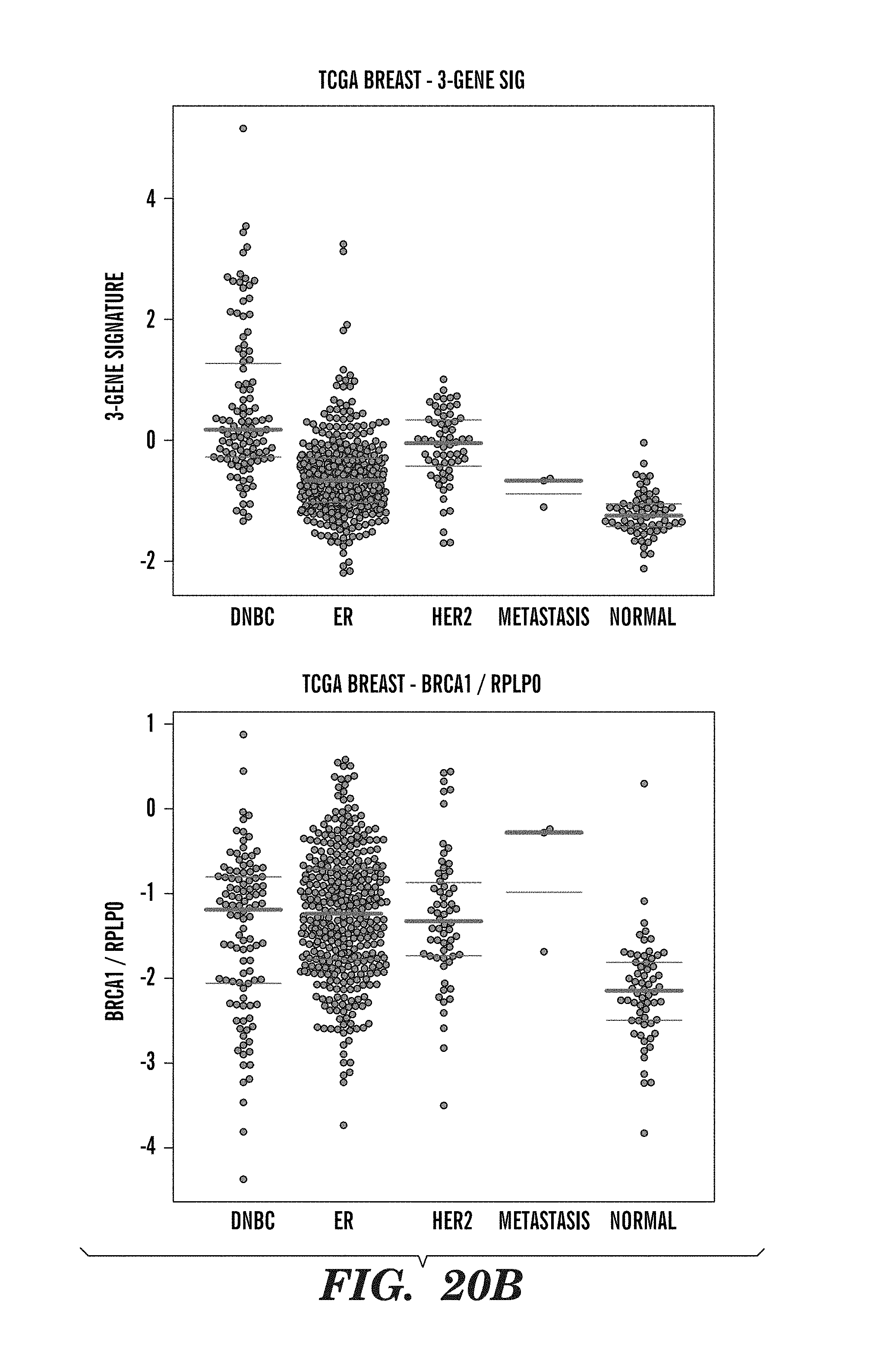

D00025

D00026

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.