Antibodies Against Human Csf-1r And Uses Thereof
DIMOUDIS; Nikolaos ; et al.
U.S. patent application number 16/362962 was filed with the patent office on 2019-10-03 for antibodies against human csf-1r and uses thereof. This patent application is currently assigned to HOFFMANN-LA ROCHE INC.. The applicant listed for this patent is HOFFMANN-LA ROCHE INC.. Invention is credited to Nikolaos DIMOUDIS, Georg FERTIG, Alexander FIDLER, Guy GEORGES, Klaus KALUZA, Martin LANZENDOERFER, Carola RIES, Stefan SEEBER, Marlene THOMAS.
| Application Number | 20190300614 16/362962 |
| Document ID | / |
| Family ID | 43735019 |
| Filed Date | 2019-10-03 |









| United States Patent Application | 20190300614 |
| Kind Code | A1 |
| DIMOUDIS; Nikolaos ; et al. | October 3, 2019 |
ANTIBODIES AGAINST HUMAN CSF-1R AND USES THEREOF
Abstract
The present invention relates to antibodies against human CSF-1R (anti-CSF-1R antibody), methods for their production, pharmaceutical compositions containing said antibodies, and uses thereof.
| Inventors: | DIMOUDIS; Nikolaos; (Wielenbach, DE) ; FERTIG; Georg; (Penzberg, DE) ; FIDLER; Alexander; (Penzberg, DE) ; GEORGES; Guy; (Habach, DE) ; KALUZA; Klaus; (Weilheim, DE) ; LANZENDOERFER; Martin; (Tutzing, DE) ; RIES; Carola; (Penzberg, DE) ; SEEBER; Stefan; (Penzberg, DE) ; THOMAS; Marlene; (Penzberg, DE) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | HOFFMANN-LA ROCHE INC. Little Falls NJ |
||||||||||
| Family ID: | 43735019 | ||||||||||
| Appl. No.: | 16/362962 | ||||||||||
| Filed: | March 25, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 16055947 | Aug 6, 2018 | 10287358 | ||
| 16362962 | ||||
| 15875530 | Jan 19, 2018 | 10077314 | ||
| 16055947 | ||||
| 15489519 | Apr 17, 2017 | 9879085 | ||
| 15875530 | ||||
| 15292710 | Oct 13, 2016 | 9663580 | ||
| 15489519 | ||||
| 14640183 | Mar 6, 2015 | 9499625 | ||
| 15292710 | ||||
| 12962445 | Dec 7, 2010 | 8999327 | ||
| 14640183 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61P 35/00 20180101; C07K 2317/567 20130101; A61P 29/00 20180101; C07K 2317/56 20130101; C07K 2317/622 20130101; A61P 19/08 20180101; C07K 2317/565 20130101; C07K 2317/92 20130101; A61P 19/10 20180101; A61P 35/04 20180101; C07K 2317/31 20130101; C07K 16/2866 20130101; C07K 2317/76 20130101; C07K 2317/24 20130101; C07K 2317/21 20130101; A61P 37/00 20180101; C07K 2317/73 20130101; A61K 2039/505 20130101 |
| International Class: | C07K 16/28 20060101 C07K016/28 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Dec 10, 2009 | EP | 09015310.7 |
| Aug 19, 2010 | EP | 10173407.7 |
Claims
1. An isolated antibody binding to human CSF-1R, wherein the antibody comprises a heavy chain variable domain and a light chain variable domain, and wherein the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO: 43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO: 45, and a CDR1 region comprising SEQ ID NO: 46.
2. A pharmaceutical composition comprising the antibody according to claim 1.
3. The pharmaceutical composition according to claim 2, wherein the antibody is formulated in a pharmaceutically acceptable carrier.
4. The antibody according to claim 1, wherein the antibody is of human IgG1 subclass or is of human IgG4 subclass.
5. The antibody according to claim 1, wherein the antibody is a chimeric, single chain, multispecific, or humanized antibody.
6. A pharmaceutical composition comprising the antibody according to claim 5.
7. The antibody according to claim 1, wherein the antibody is an antigen binding fragment.
8. A pharmaceutical composition comprising the antibody according to claim 7.
9. The antibody according to claim 1, wherein the antibody is a monoclonal antibody.
10. A pharmaceutical composition comprising the antibody according to claim 9.
11. An isolated antibody binding to human CSF-1R, wherein the antibody comprises a heavy chain variable domain and a light chain variable domain, and wherein the heavy chain variable domain comprises SEQ ID NO: 47 and the light chain variable domain comprises SEQ ID NO: 48.
12. A pharmaceutical composition comprising the antibody according to claim 11.
13. The pharmaceutical composition according to claim 12, wherein the antibody is formulated in a pharmaceutically acceptable carrier.
14. The antibody according to claim 11, wherein the antibody is of human IgG1 subclass or is of human IgG4 subclass.
15. The antibody according to claim 11, wherein the antibody is a chimeric, single chain, multispecific, or humanized antibody.
16. A pharmaceutical composition comprising the antibody according to claim 15.
17. The antibody according to claim 11, wherein the antibody is an antigen binding fragment.
18. A pharmaceutical composition comprising the antibody according to claim 17.
19. The antibody according to claim 11, wherein the antibody is a monoclonal antibody.
20. A pharmaceutical composition comprising the antibody according to claim 19.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a Divisional of U.S. patent application Ser. No. 15/292,710, filed Oct. 13, 2016, which is a Divisional of U.S. patent application Ser. No. 14/640,183, filed Mar. 6, 2015, now U.S. Pat. No. 9,499,625, issued Nov. 22, 2016, which is a Divisional of U.S. patent application Ser. No. 12/962,445, filed Dec. 7, 2010, now U.S. Pat. No. 8,999,327, issued Apr. 7, 2015, which claims the benefit of EP Patent Application Nos. 09 015 310.7, filed Dec. 10, 2009, and 10 173 407.7, filed Aug. 19, 2010, the entire disclosures of which are expressly incorporated by reference herein.
SUBMISSION OF SEQUENCE LISTING ON ASCII TEXT FILE
[0002] The content of the following submission on ASCII text file is incorporated herein by reference in its entirety: a computer readable form (CRF) of the Sequence Listing (file name: 146392029613SEQLIST.txt, date recorded: Apr. 14, 2017, size: 68 KB).
FIELD OF THE INVENTION
[0003] The present invention relates to antibodies against human CSF-1R (anti-CSF-1R antibodies), methods for their production, pharmaceutical compositions containing said antibodies, and uses thereof.
BACKGROUND OF THE INVENTION
[0004] The human CSF-1 receptor (CSF-1R; colony stimulating factor 1 receptor; synonyms: M-CSF receptor; Macrophage colony-stimulating factor 1 receptor. Fins proto-oncogene, c-fins. SEQ ID NO: 62) is known since 1986 (Coussens, L., et al., Nature 320 (1986) 277-280). CSF-1R is a growth factor and encoded by the c-fms proto-oncogene (reviewed e.g. in Roth, P., and Stanley. E. R., Curr. Top. Microbiol. Immunol. 181 (1992) 141-67).
[0005] CSF-1R is the receptor for CSF-1 (colony stimulating factor 1, also called M-CSF, macrophage colony-stimulating factor) and mediates the biological effects of this cytokine (Sherr, C. J., et al., Cell 41 (1985) 665-676). The cloning of the colony stimulating factor-1 receptor (CSF-1R) (also called c-fms) was described for the first time in Roussel, M. F., et al., Nature 325 (1987) 549-552. In that publication, it was shown that CSF-1R had transforming potential dependent on changes in the Cterminal tail of the protein including the loss of the inhibitory tyrosine 969 phosphorylation which binds Cbl and thereby regulates receptor down regulation (Lee, P. S., et al., Embo J. 18 (1999) 3616-3628). Recently a second ligand for CSF-1R termed interleukin-34 (IL-34) was identified (Lin. H., et al. Science 320 (2008) 807-811).
[0006] The cytokine CSF-1 (colony stimulating factor 1, also called M-CSF, macrophage) is found extracellularly as a disulfide-linked homodimer (Stanley, E. R. et al.,
[0007] Journal of Cellular Biochemistry 21 (1983) 151-159; Stanley, E. R. et al., Stem Cells 12 Suppl. 1 (1995) 15-24).
[0008] The main biological effects of CSF-1R signaling are the differentiation, proliferation, migration, and survival of hematopoietic precursor cells to the macrophage lineage (including osteoclast). Activation of CSF-1R is mediated by its ligands, CSF-1 (M-CSF) and IL-34. Binding of CSF-1 (M-CSF) to CSF-1R induces the formation of homodimers and activation of the kinase by tyrosine phosphorylation (Li, W. et al, EMBO Journal. 10 (1991) 277-288; Stanley, E. R., et al., Mol. Reprod. Dev. 46 (1997) 4-10).
[0009] The biologically active homodimer CSF-1 binds to the CSF-1R within the subdomains D1 to D3 of the extracellular domain of the CSF-1 receptor (CSF-1R-ECD). The CSF-1R-ECD comprises five immunoglobulin-like subdomains (designated D1 to D5). The subdomains D4 to D5 of the extracellular domain (CSF-1R-ECD) are not involved in the CSF-1 binding. (Wang, Z., et al Molecular and Cellular Biology 13 (1993) 5348-5359). The subdomain D4 is involved in dimerization (Yeung, Y-G., et al Molecular & Cellular Proteomics 2 (2003) 1143-1155; Pixley, F. J., et al., Trends Cell Biol 14 (2004) 628-638).
[0010] Further signaling is mediated by the p85 subunit of PI3K and Grb2 connecting to the PI3K/AKT and Ras/MAPK pathways, respectively. These two important signaling pathways can regulate proliferation, survival and apoptosis. Other signaling molecules that bind the phosphorylated intracellular domain of CSF-1R include STAT1, STAT3, PLCy, and Cbl (Bourette, R. P. and Rohrschneider, L. R., Growth Factors 17 (2000) 155-166).
[0011] CSF-1R signaling has a physiological role in immune responses, in bone remodeling and in the reproductive system. The knockout animals for either CSF-1 (Pollard, J. W., Mol. Reprod. Dev. 46 (1997) 54-61) or CSF-1R (Dai, X. M., et al., Blood 99 (2002) 111-120) have been shown to have osteopetrotic, hematopoietic, tissue macrophage, and reproductive phenotypes consistent with a role for CSF-1R in the respective cell types.
[0012] Sherr, C. J., et al., Blood 73 (1989) 1786-1793 relates to some antibodies against CSF-1R that inhibit the CSF-1 activity (see Sherr, C. J. et al., Blood 73 (1989) 1786-1793). Ashmun, R. A., et al., Blood 73 (1989) 827-837 relates to CSF-1R antibodies. Lenda, D., et al., Journal of Immunology 170 (2003) 3254-3262 relates to reduced macrophage recruitment, proliferation, and activation in CSF-1-deficient mice results in decreased tubular apoptosis during renal inflammation. Kitaura, H., et al., Journal of Dental Research 87 (2008) 396-400 refers to an anti-CSF-1 antibody which inhibits orthodontic tooth movement. WO 2001/030381 mentions CSF-1 activity inhibitors including antisense nucleotides and antibodies while disclosing only CSF-1 antisense nucleotides. WO 2004/045532 relates to metastases and bone loss prevention and treatment of metastatic cancer by a CSF-1 antagonist disclosing as antagonist anti-CSF-1-antibodies only. WO 2005/046657 relates to the treatment of inflammatory bowel disease by anti-CSF-1-antibodies. US 2002/0141994 relates to inhibitors of colony stimulating factors. WO 2006/096489 relates to the treatment of rheumatoid arthritis by anti-CSF-1-antibodies. WO 2009/026303 and WO 2009/112245 relate to certain anti-CSF-1R antibodies binding to CSF-1R within the first three subdomains (D1 to D3) of the Extracellular Domain (CSF-1R-ECD).
SUMMARY OF THE INVENTION
[0013] The invention comprises an isolated antibody binding to human CSF-1R, wherein the antibody binds to human CSF-1R fragment delD4 (SEQ ID NO: 65) and to human CSF-1R Extracellular Domain (SEQ ID NO: 64) with a ratio of 1:50 or lower.
[0014] The invention further comprises an isolated antibody wherein [0015] a) the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, [0016] b) the heavy chain variable domain comprises SEQ ID NO:15 and the light chain variable domain comprises SEQ ID NO: 16; [0017] c) the heavy chain variable domain comprises SEQ ID NO:75 and the light chain variable domain comprises SEQ ID NO:76; [0018] d) the heavy chain variable domain comprises SEQ ID NO:83 and the light chain variable domain comprises SEQ ID NO:84;
[0019] or a humanized version thereof.
[0020] The invention further comprises an isolated antibody wherein [0021] a) the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, [0022] b) the heavy chain variable domain comprises SEQ TD NO:15 and the light chain variable domain comprises SEQ ID NO: 16;
[0023] or a humanized version thereof.
[0024] In one embodiment the isolated antibody comprises [0025] a) a heavy chain variable domain comprising SEQ ID NO:23 and a light chain variable domain comprising SEQ ID NO:24, or [0026] b) a heavy chain variable domain comprising SEQ ID NO:31 and a light chain variable domain comprising SEQ ID NO:32, or [0027] c) a heavy chain variable domain comprising SEQ ID NO:39 and a light chain variable domain comprising SEQ ID NO:40, or [0028] d) a heavy chain variable domain comprising SEQ ID NO:47 and a light chain variable domain comprising SEQ ID NO:48, or [0029] e) a heavy chain variable domain comprising SEQ ID NO:55 and a light chain variable domain comprising SEQ ID NO:56.
[0030] The invention further comprises an isolated antibody, wherein [0031] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 1, a CDR2 region comprising SEQ ID NO: 2, and a CDR1 region comprising SEQ ID NO:3, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 4, a CDR2 region comprising SEQ ID NO:5, and a CDR1 region comprising SEQ ID NO:6, or [0032] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 9, a CDR2 region comprising SEQ ID NO: 10, and a CDR1 region comprising SEQ ID NO: 11, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:12, a CDR2 region comprising SEQ ID NO: 13, and a CDR1 region comprising SEQ ID NO: 14, or [0033] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0034] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0035] e) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0036] f) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46, or [0037] g) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 49, a CDR2 region comprising SEQ ID NO: 50, and a CDR1 region comprising SEQ ID NO: 51, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:52, a CDR2 region comprising SEQ ID NO: 53, and a CDR1 region comprising SEQ ID NO: 54; or [0038] h) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:69, a CDR2 region comprising SEQ ID NO: 70, and a CDR1 region comprising SEQ ID NO:71, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 72, a CDR2 region comprising SEQ ID NO:73, and a CDR1 region comprising SEQ ID NO:74, or [0039] i) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 77, a CDR2 region comprising SEQ ID NO: 78, and a CDR1 region comprising SEQ ID NO: 79, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:80, a CDR2 region comprising SEQ ID NO: 81, and a CDR1 region comprising SEQ ID NO: 82.
[0040] Preferably the antibody according to the invention is of human IgG1 subclass or of human IgG4 subclass.
[0041] A further embodiment of the invention is a pharmaceutical composition comprising any of the antibodies described herein.
[0042] The invention further comprises the use an of an antibody according to the invention for the manufacture of a medicament for treatment of a CSF-1R mediated disease.
[0043] The invention further comprises the use an of an antibody according to the invention for the manufacture of a medicament for treatment of cancer.
[0044] The invention further comprises the use an of an antibody according to the invention for the manufacture of a medicament for treatment of bone loss.
[0045] The invention further comprises the use an of an antibody according to the invention for the manufacture of a medicament for treatment of metastasis.
[0046] The invention further comprises the use an of an antibody according to the invention for the manufacture of a medicament for treatment of inflammatory diseases.
[0047] The invention further comprises an antibody according to the invention for treatment of a CSF-1R mediated disease.
[0048] The invention further comprises an antibody according to the invention for treatment of cancer.
[0049] The invention further comprises an antibody according to the invention for treatment of bone loss.
[0050] The invention further comprises an antibody according to the invention for treatment of metastasis.
[0051] The invention further comprises an antibody according to the invention for treatment of inflammatory diseases.
[0052] A further embodiment of the invention is a nucleic acid encoding an antibody wherein [0053] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 1, a CDR2 region comprising SEQ ID NO: 2, and a CDR1 region comprising SEQ ID NO:3, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 4, a CDR2 region comprising SEQ ID NO:5, and a CDR1 region comprising SEQ ID NO:6, or, [0054] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 9, a CDR2 region comprising SEQ ID NO: 10, and a CDR1 region comprising SEQ ID NO: 11, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:12, a CDR2 region comprising SEQ ID NO: 13, and a CDR1 region comprising SEQ ID NO: 14, or [0055] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0056] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0057] e) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0058] f) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46, or [0059] g) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 49, a CDR2 region comprising SEQ ID NO: 50, and a CDR1 region comprising SEQ ID NO: 51, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:52, a CDR2 region comprising SEQ ID NO: 53, and a CDR1 region comprising SEQ ID NO: 54, or [0060] h) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:69, a CDR2 region comprising SEQ ID NO: 70, and a CDR1 region comprising SEQ ID NO:71, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 72, a CDR2 region comprising SEQ ID NO:73, and a CDR1 region comprising SEQ ID NO:74, or [0061] i) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 77, a CDR2 region comprising SEQ ID NO: 78, and a CDR1 region comprising SEQ ID NO: 79, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:80, a CDR2 region comprising SEQ ID NO: 81, and a CDR1 region comprising SEQ ID NO: 82.
[0062] A further embodiment of the invention is a nucleic acid encoding an antibody wherein [0063] a) the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, [0064] b) the heavy chain variable domain comprises SEQ ID NO:15 and the light chain variable domain comprises SEQ ID NO: 16; [0065] c) the heavy chain variable domain comprises SEQ ID NO:75 and the light chain variable domain comprises SEQ ID NO:76; [0066] d) the heavy chain variable domain comprises SEQ ID NO:83 and the light chain variable domain comprises SEQ ID NO:84;
[0067] or a humanized version thereof.
[0068] A further embodiment of the invention is a nucleic acid encoding an antibody wherein [0069] a) the heavy chain variable domain comprises SEQ ID NO:23 and the light chain variable domain comprises SEQ ID) NO:24, or [0070] b) the heavy chain variable domain comprises SEQ ID NO:31 and the light chain variable domain comprises SEQ ID NO:32, or [0071] c) the heavy chain variable domain comprises SEQ ID NO:39 and the light chain variable domain comprises SEQ ID NO:40, or [0072] d) the heavy chain variable domain comprises SEQ ID NO:47 and the light chain variable domain comprises SEQ ID NO:48, or [0073] e) the heavy chain variable domain comprises SEQ ID NO:55 and the light chain variable domain comprises SEQ ID NO:56.
[0074] The invention further provides expression vectors containing nucleic acid according to the invention capable of expressing said nucleic acid in a prokaryotic or eukaryotic host cell, and host cells containing such vectors for the recombinant production of an antibody according to the invention.
[0075] The invention further comprises a prokaryotic or eukaryotic host cell comprising a vector according to the invention.
[0076] The invention further comprises a method for the production of a recombinant human or humanized antibody according to the invention, the method comprising expressing a nucleic acid according to the invention in a prokaryotic or eukaryotic host cell and recovering said antibody from said cell or the cell culture supernatant.
[0077] The invention further comprises the antibody obtained by such a recombinant method.
[0078] Antibodies according to the invention show benefits for patients in need of a CSF-1R targeting therapy. The antibodies according to the invention show efficient antiproliferative activity against ligand-independent and ligand-dependant proliferation and are therefore especially useful in the treatment of cancer and metastasis.
[0079] The invention further provides a method for treating a patient suffering from cancer, the method comprising administering to a patient diagnosed as having such a disease (and therefore being in need of such a therapy) an effective amount of an antibody according to the invention. The antibody is administered preferably in a pharmaceutical composition.
[0080] A further embodiment of the invention is a method for treating a patient suffering from cancer the method comprising administering to the patient an antibody according to the invention.
[0081] These and other embodiments of the invention are further described in the detailed description that follows.
DESCRIPTION OF THE SEQUENCES
[0082] SEQ ID NO: 1 heavy chain CDR3, Mab 2F11 [0083] SEQ ID NO: 2 heavy chain CDR2, Mab 2F11 [0084] SEQ ID NO: 3 heavy chain CDR1, Mab 2F11 [0085] SEQ ID NO: 4 light chain CDR3, Mab 2F11 [0086] SEQ ID NO: 5 light chain CDR2, Mab 2F11 [0087] SEQ ID NO: 6 light chain CDR1, Mab 2F11 [0088] SEQ ID NO: 7 heavy chain variable domain, Mab 2F11 [0089] SEQ ID NO: 8 light chain variable domain, Mab 2F11 [0090] SEQ ID NO: 9 heavy chain CDR3, Mab 2E10 [0091] SEQ ID NO: 10 heavy chain CDR2, Mab 2E10 [0092] SEQ ID NO: 11 heavy chain CDR1, Mab 2E10 [0093] SEQ ID NO: 12 light chain CDR3, Mab 2E10 [0094] SEQ ID NO: 13 light chain CDR2, Mab 2E10 [0095] SEQ ID NO: 14 light chain CDR1, Mab 2E10 [0096] SEQ ID NO: 15 heavy chain variable domain, Mab 2E10 [0097] SEQ ID NO: 16 light chain variable domain, Mab 2E10 [0098] SEQ ID NO: 17 heavy chain CDR3, hMab 2F11-c11 [0099] SEQ ID NO: 18 heavy chain CDR2, hMab 2F11-c11 [0100] SEQ ID NO: 19 heavy chain CDR1, hMab 2F11-c11 [0101] SEQ ID NO: 20 light chain CDR3, hMab 2F11-c11 [0102] SEQ ID NO: 21 light chain CDR2, hMab 2F11-c11 [0103] SEQ ID NO: 22 light chain CDR1, hMab 2F11-c11 [0104] SEQ ID NO: 23 heavy chain variable domain, hMab 2F11-c11 [0105] SEQ ID NO: 24 light chain variable domain, hMab 2F11-c11 [0106] SEQ ID NO: 25 heavy chain CDR3, hMab 2F11-d8 [0107] SEQ ID NO: 26 heavy chain CDR2, hMab 2F11-d8 [0108] SEQ ID NO: 27 heavy chain CDR1, hMab 2F11-d8 [0109] SEQ ID NO: 28 light chain CDR3, hMab 2F11-d8 [0110] SEQ ID NO: 29 light chain CDR2, hMab 2F11-d8 [0111] SEQ ID NO: 30 light chain CDR1, hMab 2F11-d8 [0112] SEQ ID NO: 31 heavy chain variable domain, hMab 2F11-d8 [0113] SEQ ID NO: 32 light chain variable domain, hMab 2F11-d8 [0114] SEQ ID NO: 33 heavy chain CDR3, hMab 2F11-e7 [0115] SEQ ID NO: 34 heavy chain CDR2, hMab 2F11-e7 [0116] SEQ ID NO: 35 heavy chain CDR1, hMab 2F11-e7 [0117] SEQ ID NO: 36 light chain CDR3, hMab 2F11-e7 [0118] SEQ ID NO: 37 light chain CDR2, hMab 2F11-e7 [0119] SEQ ID NO: 38 light chain CDR1, hMab 2F11-e7 [0120] SEQ ID NO: 39 heavy chain variable domain, hMab 2F11-e7 [0121] SEQ ID NO: 40 light chain variable domain, hMab 2F11-e7 [0122] SEQ ID NO: 41 heavy chain CDR3, hMab 2F11-f12 [0123] SEQ ID NO: 42 heavy chain CDR2, hMab 2F11-f12 [0124] SEQ TD NO: 43 heavy chain CDR1, hMab 2F11-f12 [0125] SEQ ID NO: 44 light chain CDR3, hMab 2F11-f12 [0126] SEQ ID NO: 45 light chain CDR2, hMab 2F11-f12 [0127] SEQ ID NO: 46 light chain CDR1, hMab 2F11-f12 [0128] SEQ ID NO: 47 heavy chain variable domain, hMab 2F11-f12 [0129] SEQ ID NO: 48 light chain variable domain, hMab 2F11-f12 [0130] SEQ ID NO: 49 heavy chain CDR3, hMab 2F11-g1 [0131] SEQ ID NO: 50 heavy chain CDR2, hMab 2F11-g1 [0132] SEQ ID NO: 51 heavy chain CDR1, hMab 2F11-g1 [0133] SEQ ID NO: 52 light chain CDR3, hMab 2F11-g1 [0134] SEQ ID NO: 53 light chain CDR2, hMab 2F11-g1 [0135] SEQ ID NO: 54 light chain CDR1, hMab 2F11-g1 [0136] SEQ ID NO: 55 heavy chain variable domain, hMab 2F11-g1 [0137] SEQ ID NO: 56 light chain variable domain, hMab 2F11-g1 [0138] SEQ ID NO: 57 human kappa light chain constant region [0139] SEQ ID NO: 58 human heavy chain constant region derived from IgG [0140] SEQ ID NO: 59 human heavy chain constant region derived from IgG1 mutated on L234A and L235A [0141] SEQ ID NO: 60 human heavy chain constant region derived from IgG4 [0142] SEQ ID NO: 61 human heavy chain constant region derived from IgG4 mutated on S228P [0143] SEQ ID NO: 62 human wildtype CSF-1R (wt CSF-1R) [0144] SEQ ID NO: 63 human mutant CSF-1R L3O1S Y969F [0145] SEQ ID NO: 64 human CSF-1R Extracellular Domain [0146] SEQ ID NO: 65 human CSF-1R fragment delD4 [0147] SEQ ID NO: 66 human CSF-1R fragment D1-D3 [0148] SEQ ID NO: 67 signal peptide [0149] SEQ ID NO: 68 Primer [0150] SEQ ID NO: 69 heavy chain CDR3, Mab 1G10 [0151] SEQ ID NO: 70 heavy chain CDR2, Mab 1G10 [0152] SEQ ID NO: 71 heavy chain CDR1, Mab 1G10 [0153] SEQ ID NO: 72 light chain CDR3, Mab 1G10 [0154] SEQ ID NO: 73 light chain CDR2, Mab 1G10 [0155] SEQ ID NO: 74 light chain CDR1, Mab 1G10 [0156] SEQ ID NO: 75 heavy chain variable domain, Mab 1G10 [0157] SEQ ID NO: 76 light chain variable domain, Mab 1G10 [0158] SEQ ID NO: 77 heavy chain CDR3, Mab 2H7 [0159] SEQ ID NO: 78 heavy chain CDR2, Mab 2H7 [0160] SEQ ID NO: 79 heavy chain CDR1, Mab 2H7 [0161] SEQ ID NO: 80 light chain CDR3, Mab 2H7 [0162] SEQ ID NO: 81 light chain CDR2, Mab 2H7 [0163] SEQ ID NO: 82 light chain CDR1, Mab 2H7 [0164] SEQ ID NO: 83 heavy chain variable domain, Mab 2H7 [0165] SEQ ID NO: 84 light chain variable domain, Mab 2H7
DESCRIPTION OF THE FIGURES
[0166] FIG. 1A-1B Growth inhibition of BeWo tumor cells in 3D culture under treatment with different anti-CSF-1R monoclonal antibodies at a concentration of 10 .mu.g/ml. [0167] X axis: viability normalized mean relative light units (RLU) corresponding to the ATP-content of the cells (CellTiterGlo assay). [0168] Y axis: tested probes: Minimal Medium (0.5% FBS), mouse IgG1 (mIgG1, 10 .mu.g/ml), mouse IgG2a (mIgG2a 10 .mu.g/ml), CSF-1 only, Mab 2F11, Mab 2E10, Mab2H7. Mab IG10 and SC 2-4A5. Highest inhibition of CSF-1 induced growth was observed with the anti-CSF-1R antibodies according to the invention.
[0169] FIG. 2A Biacore sensogram of binding of different anti-CSF-1R antibodies to immobilized human CSF-1R fragment delD4 (comprising the extracellular subdomains D1-D3 and D5) (SEQ ID NO: 65) (y-axis: binding signal in Response Units (RU), baseline=0 RU, x-axis: time in seconds (s)): While the antibodies Mab 3291 and sc 2-4A5 clearly show binding to this delD4 fragment, the antibodies according to the invention e.g. Mab 2F11, and Mab 2E10, did not bind to the CSF-1R fragment delD4. The control anti-CCR5 antibody m<CCR5>Pz03.1C5 did also not bind to the CSF-1R fragment delD4.
[0170] FIG. 2B Biacore sensogram of binding of different anti-CSF-1R antibodies to immobilized human CSF-1R Extracellular Domain (CSF-1R-ECD) (comprising the extracellular subdomains D1-D5) (SEQ ID NO: 64) (y-axis: binding signal in Response Units (RU), baseline=0 RU, x-axis: time in seconds (s)): All anti-CSF-1R antibodies show binding to CSF-1R-ECD. The control anti-CCR5 antibody m<CCR5>Pz03.1C5 did not bind to the CSF-1R-ECD.
[0171] FIG. 2C Biacore sensogram of binding of different anti-CSF-1R antibodies to immobilized human CSF-1R fragment delD4 (comprising the extracellular subdomains D1-D3 and D5) (SEQ ID NO: 65) (y-axis: binding signal in Response Units (RU), baseline=0 RU, x-axis: time in seconds (s)): Mab 1G10, Mab 2H7 and humanized hMab 2F11-e7 did not bind to the CSF-1R fragment delD4. The control anti-CCR5 antibody m<CCR5>Pz03.1C5 did also not bind to the CSF-1R fragment delD4.
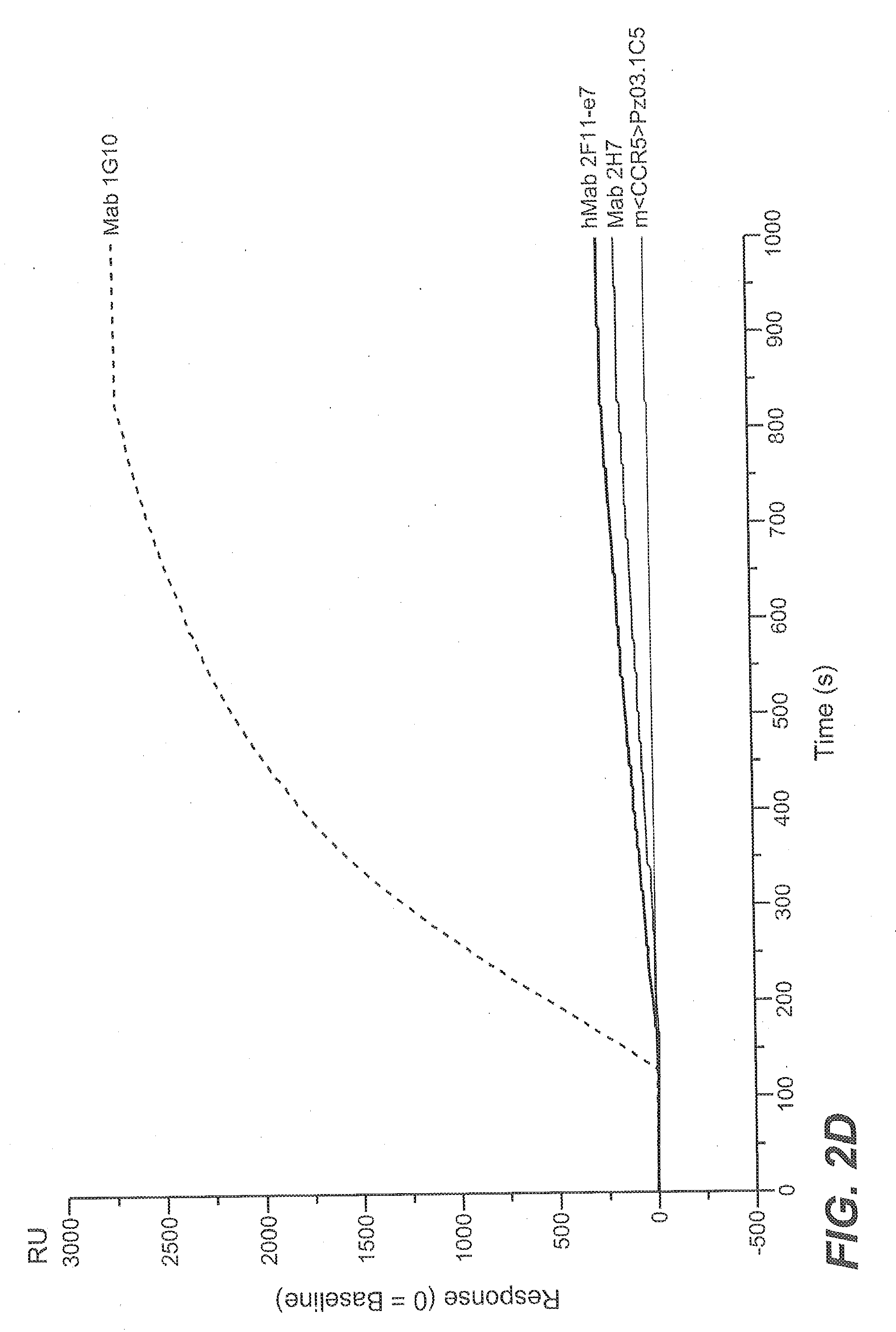
[0172] FIG. 2D Biacore sensogram of binding of different anti-CSF-1R antibodies to immobilized human CSF-1R Extracellular Domain (CSF-1R-ECD) (comprising the extracellular subdomains D1-D5) (SEQ ID NO: 64) (y-axis: binding signal in Response Units (RU), baseline=0 RU, x-axis: time in seconds (s)): All anti-CSF-1R antibodies Mab 1G10. Mab 2H7 and humanized hMab 2F11-e7 showed binding to CSF-1R-ECD. The control anti-CCR5 antibody m<CCR5>Pz03.1C5 did not bind to the CSF-1R-ECD.
[0173] FIG. 2E Biacore sensogram of binding of different anti-CSF-1R antibodies to immobilized human CSF-1R fragment delD4 (comprising the extracellular subdomains D1-D3 and D5) (SEQ ID NO: 65) (y-axis: binding signal in Response Units (RU), baseline=0 RU, x-axis: time in seconds (s)): All anti-CSF-1R antibodies 1.2.SM, CXIIG6, ab10676 and MAB3291 show binding to to the CSF-1R fragment delD4. The control anti-CCR5 antibody m<CCR5>Pz03.1C5 did also not bind to the CSF-1R fragment delD4.
[0174] FIG. 2F Biacore sensogram of binding of different anti-CSF-1R antibodies to immobilized human CSF-1R Extracellular Domain (CSF-1R-ECD) (comprising the extracellular subdomains D1-D5) (SEQ ID NO: 64) (y-axis: binding signal in Response Units (RU), baseline=0 RU, x-axis: time in seconds (s)): All anti-CSF-1R antibodies 1.2.SM, CXIIG6, ab10676 and MAB3291 show binding to CSF-1R-ECD. The control anti-CCR5 antibody m<CCR5>Pz03.1C5 did not bind to the CSF-1R-ECD.
[0175] FIG. 3A-3D CSF-1 levels in Cynomolgus monkey after application of different dosages of anti-CSF-1R antibody according to the invention
[0176] FIG. 4 In vivo efficacy--tumor growth inhibition of anti-CSF-1R antibodies according to the invention in breast cancer BT20 xenograft
DETAILED DESCRIPTION OF THE INVENTION
I. Introduction
[0177] The present invention provides compositions that bind human CSF-1R or a portion thereof, kits and articles of manufacture comprising such compositions, and methods of using such compositions, including, e.g., methods for modulating ligand binding to CSF-1R and for modulating biological/physiological activities associated with ligand binding to CSF-1R. The invention is based in part on the identification of a variety of anti-CSF-1R antibodies that bind CSF-1R. The anti-CSF-1R antibodies of the invention can conveniently be used as therapeutic and diagnostic agents for use in targeting CSF-1R-mediated diseases including, e.g., cancer (including, for example, metastatic cancer), bone disorders (including, for example, bone loss), and inflammatory disorders.
[0178] Surprisingly it has been found that, using a human CSF-1R fragment delD4 in which the D4 subdomain of human CSF-1R-ECD was deleted (SEQ ID NO:65), the new anti-CSF-1R antibodies according to the invention could be selected. These antibodies show valuable properties like excellent ligand-dependant cell growth inhibition and at the same time ligand independent cell growth inhibition of NIH 3T3 cell, retrovirally infected with either an expression vector for full-length wildtype CSF-1R (SEQ ID NO:62) or mutant CSF-1R L301S Y969F (SEQ ID NO:63) whereby mutant CSF-1R recombinant cells are able to form spheroids independent of the CSF-1 ligand. Furthermore the antibodies according to the invention inhibit (both) human and cynomolgous macrophage differentiation, as they inhibit survival of human and cynomolgous monocytes.
II. Definitions
[0179] As used herein, the following terms have the meanings ascribed to them below unless otherwise specified.
[0180] The human CSF-1R (CSF-1 receptor; synonyms: M-CSF receptor; Macrophage colony-stimulating factor 1 receptor. Fms proto-oncogene, c-fms, SEQ ID NO: 22)) is known since 1986 (Coussens, L., et al., Nature 320 (1986) 277-280). CSF-1R is a growth factor and encoded by the c-fms proto-oncogene (reviewed e.g. in Roth, P. and Stanley, E. R., Curr. Top. Microbiol. Immunol. 181 (1992) 141-67).
[0181] CSF-1R is the receptor for CSF-1 (macrophage colony stimulating factor, also called M-CSF) and IL-34 and mediates the biological effects of these cytokines (Sherr, C. J., et al., Cell 41 (1985) 665-676 (Lin, H., et al., Science 320 (2008) 807-811). The cloning of the colony stimulating factor-1 receptor (also called c-fms) was described for the first time in Roussel, M. F., et al., Nature 325 (1987) 549-552. In that publication, it was shown that CSF-1R had transforming potential dependent on changes in the C-terminal tail of the protein including the loss of the inhibitory tyrosine 969 phosphorylation which binds Cbl and thereby regulates receptor down regulation (Lee, P. S., et al., Embo J. 18 (1999) 3616-3628). CSF-1R is a single chain, transmembrane receptor tyrosine kinase (RTK) and a member of the family of immunoglobulin (Ig) motif containing RTKs characterized by 5 repeated Ig-like subdomains D1-D5 in the extracellular domain (ECD) of the receptor (Wang, Z., et al Molecular and Cellular Biology 13 (1993) 5348-5359). The human CSF-1R Extracellular Domain (CSF-1R-ECD) (SEQ ID NO: 64) comprises all five extracellular Ig-like subdomains D1-D5. The human CSF-1R fragment delD4 (SEQ ID NO: 65) comprises the extracellular Ig-like subdomains D1-D3 and D5, but is missing the D4 subdomain. The human CSF-1R fragment D1-D3 (SEQ ID NO: 66) comprises the respective subdomains D1-D3. The sequences are listed without the signal peptide MGSGPGVLLL LLVATAWHGQ G (SEQ ID NO: 67).
[0182] The intracellular protein tyrosine kinase domain is interrupted by a unique insert domain that is also present in the other related RTK class III family members that include the platelet derived growth factor receptors (PDGFR), stem cell growth factor receptor (c-Kit) and fins-like cytokine receptor (FLT3). In spite of the structural homology among this family of growth factor receptors, they have distinct tissue-specific functions.
[0183] CSF-1R is mainly expressed on cells of the monocytic lineage and in the female reproductive tract and placenta. In addition expression of CSF-1R has been reported in Langerhans cells in skin, a subset of smooth muscle cells (Inaba, T., et al., J. Biol. Chem. 267 (1992) 5693-5699), B cells (Baker, A. H., et al., Oncogene 8 (1993) 371-378) and microglia (Sawada, M., et al., Brain Res. 509 (1990) 119-124). Cells with mutant human CSF-1R ((SEQ ID NO: 23) are known to proliferate independently of ligand stimulation.
[0184] As used herein, "binding to human CSF-1R" or "specifically binding to human CSF-1R" refers to an antibody specifically binding to the human CSF-1R antigen with a binding affinity of KD-value of 1.0.times.10.sup.-8 mol/l or lower at 35.degree. C., in one embodiment of a KD-value of 1.0.times.10.sup.-9 mol/l or lower at 35.degree. C. The binding affinity is determined with a standard binding assay at 35.degree. C., such as surface plasmon resonance technique (BIAcore.RTM., GE-Healthcare Uppsala, Sweden) A method for determining the KD-value of the binding affinity is described in Example 9. Thus an "antibody binding to human CSF-1R" as used herein refers to an antibody specifically binding to the human CSF-1R antigen with a binding affinity of KD 1.0.times.10.sup.-8 mol/l or lower (preferably 1.0.times.10.sup.0.8 mol/l-1.0.times.10.sup.-12 mol/1) at 35.degree. C., preferably of a KD 1.0.times.10.sup.-9 mol/l or lower at 35.degree. C. (preferably 1.0.times.10.sup.-9 mol/l-1.0.times.10.sup.-2 mol/l).
[0185] The "binding to human CSF-1R fragment delD4 (SEQ ID NO: 65) and to human CSF-1R Extracellular Domain (SEQ ID NO: 64)" as used herein is measured by a Surface Plasmon Resonance assay (Biacore assay) as described in Example 4. The human CSF-1R fragment delD4 (SEQ ID NO: 65) or human CSF-1R Extracellular Domain (SEQ ID NO: 64), respectively, are captured to the surface (each to a separate surface) and the test antibodies were added (each in a separate measurement) and the respective binding signals (Response Units (RU)) were determined. Reference signals (blank surface) were subtracted. If signals of nonbinding test antibodies were slightly below 0 the values were set as 0. Then the ratio of the respective binding signals (binding signal (RU) to human CSF-1R fragment delD4/binding signal (RU) to human CSF-1R Extracellular Domain (CSF-1R-ECD)) is determined. The antibodies according to the invention have a ratio of the binding signals (RU(delD4)/RU(CSF-1R-ECD) of 1:50 or lower, preferably of 1:100 or lower (the lower included end is 0 (e.g. if the RU is 0, then the ratio is 0:50 or 0:100)).
[0186] This means that such anti-CSF-1R antibodies according to the invention do not bind to the human CSF-1R fragment delD4 (like the anti-CCR5 antibody m<CCR5>Pz03.1C5 (deposited as DSM ACC 2683 on 18 Aug. 2004 at DSMZ) and have binding signals for binding to the human CSF-1R fragment delD4 in the range of the anti-CCR5 antibody m<CCR5>Pz03.1C5, which are below 20 RU (Response Units), preferably below 10 RU in a Surface Plasmon Resonance (BIAcore) assay as shown in Example 4.
[0187] The term "binding to human CSF-1R fragment D1-D3" refers to a binding affinity determination by a Surface Plasmon Resonance assay (Biacore assay). The test antibody is captured to the surface and the human CSF-1R fragment D1-D3 (SEQ ID NO: 66) was added and the respective binding affinities were determined. The term "not binding to human CSF-1R fragment D1-D3" denotes that in such an assay the detected signal was in the area of no more than 1.2 fold of background signal and therefore no significant binding could be detected and no binding affinity could be determined (see Example 10).
[0188] The term "antibody" encompasses the various forms of antibodies including but not being limited to whole antibodies, antibody fragments, human antibodies, humanized antibodies, chimeric antibodies, T cell epitope depleted antibodies, and further genetically engineered antibodies as long as the characteristic properties according to the invention are retained. "Antibody fragments" comprise a portion of a full length antibody, preferably the variable domain thereof, or at least the antigen binding site thereof. Examples of antibody fragments include diabodies, single-chain antibody molecules, and multispecific antibodies formed from antibody fragments, scFv antibodies are, e.g., described in Houston, J. S., Methods in Enzymol. 203 (1991) 46-88). In addition, antibody fragments comprise single chain polypeptides having the characteristics of a V.sub.H domain binding to CSF-1R, namely being able to assemble together with a V.sub.L domain, or of a V.sub.L domain binding to CSF-1R, namely being able to assemble together with a V.sub.H domain to a functional antigen binding site and thereby providing the property.
[0189] The terms "monoclonal antibody" or "monoclonal antibody composition" as used herein refer to a preparation of antibody molecules of a single amino acid composition.
[0190] The term "chimeric antibody" refers to a monoclonal antibody comprising a variable region, i.e., binding region, from mouse and at least a portion of a constant region derived from a different source or species, usually prepared by recombinant DNA techniques. Chimeric antibodies comprising a mouse variable region and a human constant region are especially preferred. Such rat/human chimeric antibodies are the product of expressed immunoglobulin genes comprising DNA segments encoding rat immunoglobulin variable regions and DNA segments encoding human immunoglobulin constant regions. Other forms of "chimeric antibodies" encompassed by the present invention are those in which the class or subclass has been modified or changed from that of the original antibody. Such "chimeric" antibodies are also referred to as "class-switched antibodies." Methods for producing chimeric antibodies involve conventional recombinant DNA and gene transfection techniques now well known in the art. See, e.g., Morrison, S. L., et al., Proc. Natl. Acad Sci. USA 81 (1984) 6851-6855; U.S. Pat. Nos. 5,202,238 and 5,204,244.
[0191] The term "humanized antibody" refers to antibodies in which the framework or "complementarity determining regions" (CDR) have been modified to comprise the CDR of an immunoglobulin of different specificity as compared to that of the parent immunoglobulin. In a preferred embodiment, a murine CDR is grafted into the framework region of a human antibody to prepare the "humanized antibody."
[0192] See e.g. Riechmann, L., et al., Nature 332 (1988) 323-327; and Neuberger, M. S., et al., Nature 314 (1985) 268-270. Optionally the framework region can be modified by further mutations. Also the CDRs can be modified by one or more mutations to generate antibodies according to the invention e.g. by mutagenesis based upon molecular modeling as described by Riechmann, L., et al., Nature 332 (1988) 323-327 and Queen, C., et al., Proc. Natl. Acad. Sci. USA 86 (1989) 10029-10033, or others. Particularly preferred CDRs correspond to those representing sequences recognizing the antigens noted above for chimeric antibodies. A "humanized version of an antibody according to the invention" (which is e.g. of mouse origin) refers to an antibody, which is based on the mouse antibody sequences in which the V.sub.H and V.sub.L are humanized by standard techniques (including CDR grafting and optionally subsequent mutagenesis of certain amino acids in the framework region and the CDRs). Preferably such humanized version is chimerized with a human constant region (see e.g. Sequences SEQ ID NO:57-61).
[0193] Other forms of "humanized antibodies" encompassed by the present invention are those in which the constant region has been additionally modified or changed from that of the original antibody to generate the properties according to the invention, especially in regard to C1q binding and/or Fc receptor (FcR) binding.
[0194] In the following examples the terms "Mab" or "muMab" refer to murine monoclonal antibodies such as Mab 2F11 or Mab 2E10, whereas the term "hMab" refers to humanized monoclonal versions of such murine antibodies such as hMab 2F11-c11, hMab 2F11-d8, hMab 2F11-e7, hMab 2F11-f12, etc.
[0195] The term "human antibody", as used herein, is intended to include antibodies having variable and constant regions derived from human germ line immunoglobulin sequences. Human antibodies are well-known in the state of the art (van Dijk, M. A., and van de Winkel, J. G., Curr. Opin. Chem. Biol. 5 (2001) 368-374). Human antibodies can also be produced in transgenic animals (e.g., mice) that are capable, upon immunization, of producing a full repertoire or a selection of human antibodies in the absence of endogenous immunoglobulin production. Transfer of the human germ-line immunoglobulin gene array in such germ-line mutant mice will result in the production of human antibodies upon antigen challenge (see, e.g., Jakobovits, A., et al., Proc. Natl. Acad. Sci. USA 90 (1993) 2551-2555; Jakobovits, A., et al., Nature 362 (1993) 255-258; Brueggemann, M., et al., Year Immunol. 7 (1993) 33-40). Human antibodies can also be produced in phage display libraries (Hoogenboom, H. R., and Winter, G. J. Mol. Biol. 227 (1992) 381-388; Marks, J. D., et al., J. Mol. Biol. 222 (1991) 581-597). The techniques of Cole, et al., and Boerner, et al., are also available for the preparation of human monoclonal antibodies (Cole, S. P. C., et al., Monoclonal Antibodies and Cancer Therapy, Alan R. Liss, p. 77 (1985); and Boerner, P., et al., J. Immunol. 147 (1991) 86-95). As already mentioned for chimeric and humanized antibodies according to the invention the term "human antibody" as used herein also comprises such antibodies which are modified in the constant region to generate the properties according to the invention, especially in regard to C1q binding and/or FcR binding, e.g. by "class switching" i.e. change or mutation of Fc parts (e.g. from IgG1 to IgG4 and/or IgG1/IgG4 mutation).
[0196] The term "recombinant human antibody", as used herein, is intended to include all human antibodies that are prepared, expressed, created or isolated by recombinant means, such as antibodies isolated from a host cell such as a NS0 or CHO cell or from an animal (e.g. a mouse) that is transgenic for human immunoglobulin genes or antibodies expressed using a recombinant expression vector transfected into a host cell. Such recombinant human antibodies have variable and constant regions in a rearranged form. The recombinant human antibodies according to the invention have been subjected to in vivo somatic hypermutation. Thus, the amino acid sequences of the VH and VL regions of the recombinant antibodies are sequences that, while derived from and related to human germ line VH and VL sequences, may not naturally exist within the human antibody germ line repertoire in vivo.
[0197] The antibodies according to the invention include, in addition, such antibodies having "conservative sequence modifications", nucleotide and amino acid sequence modifications which do not affect or alter the above-mentioned characteristics of the antibody according to the invention. Modifications can be introduced by standard techniques known in the art, such as site-directed mutagenesis and PCR-mediated mutagenesis. Conservative amino acid substitutions include ones in which the amino acid residue is replaced with an amino acid residue having a similar side chain. Families of amino acid residues having similar side chains have been defined in the art. These families include amino acids with basic side chains (e.g., lysine, arginine, histidine), acidic side chains (e.g., aspartic acid, glutamic acid), uncharged polar side chains (e.g. glycine, asparagine, glutamine, serine, threonine, tyrosine, cysteine, tryptophan), nonpolar side chains (e.g., alanine, valine, leucine, isoleucine, proline, phenylalanine, methionine), beta-branched side chains (e.g., threonine, valine, isoleucine) and aromatic side chains (e.g., tyrosine, phenylalanine, tryptophan, histidine). Thus, a predicted nonessential amino acid residue in a human anti-CSF-1R antibody can be preferably replaced with another amino acid residue from the same side chain family.
[0198] Amino acid substitutions can be performed by mutagenesis based upon molecular modeling as described by Riechmann, L., et al., Nature 332 (1988) 323-327 and Queen, C., et al., Proc. Natl. Acad. Sci. USA 86 (1989) 10029-10033.
[0199] The term "epitope" denotes a protein determinant of human CSF-1R capable of specifically binding to an antibody. Epitopes usually consist of chemically active surface groupings of molecules such as amino acids or sugar side chains and usually epitopes have specific three dimensional structural characteristics, as well as specific charge characteristics. Conformational and nonconformational epitopes are distinguished in that the binding to the former but not the latter is lost in the presence of denaturing solvents. Preferably an antibody according to the invention binds specifically to native and to denatured CSF-1R.
[0200] The "variable domain" (variable domain of a light chain (V.sub.L), variable domain of a heavy chain (V.sub.H)) as used herein denotes each of the pair of light and heavy chain domains which are involved directly in binding the antibody to the antigen. The variable light and heavy chain domains have the same general structure and each domain comprises four framework (FR) regions whose sequences are widely conserved, connected by three "hypervariable regions" (or complementary determining regions, CDRs). The framework regions adopt a .beta.-sheet conformation and the CDRs may form loops connecting the .beta.-sheet structure. The CDRs in each chain are held in their three-dimensional structure by the framework regions and form together with the CDRs from the other chain the antigen binding site. The antibody's heavy and light chain CDR3 regions play a particularly important role in the binding specificity/affinity of the antibodies according to the invention and therefore provide a further object of the invention.
[0201] The term "antigen-binding portion of an antibody" when used herein refer to the amino acid residues of an antibody which are responsible for antigen-binding. The antigen-binding portion of an antibody comprises amino acid residues from the "complementary determining regions" or "CDRs". "Framework" or "FR" regions are those variable domain regions other than the hypervariable region residues as herein defined. Therefore, the light and heavy chain variable domains of an antibody comprise from N- to C-terminus the domains FR1, CDR1, FR2, CDR2, FR3, CDR3, and FR4. Especially, CDR3 of the heavy chain is the region which contributes most to antigen binding and defines the antibody's properties. CDR and FR regions are determined according to the standard definition of Kabat et al., Sequences of Proteins of Immunological Interest, 5th ed., Public Health Service, National Institutes of Health, Bethesda, Md. (1991) and/or those residues from a "hypervariable loop".
[0202] The "Fc part" or "Fc portion" of an antibody is not involved directly in binding of an antibody to an antigen, but exhibit various effector functions. A "Fc part of an antibody" is a term well known to the skilled artisan and defined on the basis of papain cleavage of antibodies. Depending on the amino acid sequence of the constant region of their heavy chains, antibodies or immunoglobulins are divided in the classes: IgA, IgD, IgE, IgG and IgM, and several of these may be further divided into subclasses (isotypes), e.g. IgG1, IgG2, IgG3, and IgG4, IgA1, and IgA2. According to the heavy chain constant regions the different classes of immunoglobulins are called .alpha., .delta., .epsilon., .gamma., and .mu., respectively. The Fc part of an antibody is directly involved in ADCC (antibody-dependent cell-mediated cytotoxicity) and CDC (complement-dependent cytotoxicity) based on complement activation, C1q binding and Fc receptor binding. Complement activation (CDC) is initiated by binding of complement factor C1q to the Fc part of most IgG antibody subclasses. While the influence of an antibody on the complement system is dependent on certain conditions, binding to C1q is caused by defined binding sites in the Fc part. Such binding sites are known in the state of the art and described e.g. by Boackle, R. J., et al., Nature 282 (1979) 742-743, Lukas, T. J., et al., J. Immunol. 127 (1981) 2555-2560, Brunhouse, R., and Cebra, J. J., Mol. Immunol. 16 (1979) 907-917, Burton, D. R., et al., Nature 288 (1980) 338-344, Thommesen, J. E., et al., Mol. Immunol. 37 (2000) 995-1004, Idusogie, E. E., et al., J. Immunol. 164 (2000) 4178-4184, Hezareh, M., et al., J. Virology 75 (2001) 12161-12168, Morgan, A., et al., Immunology 86 (1995) 319-324, EP 0307434. Such binding sites are e.g. L234, L235, D270, N297, E318, K320, K322, P331 and P329 (numbering according to EU index of Kabat, E. A., see below). Antibodies of subclass IgG1, IgG2 and IgG3 usually show complement activation and C1q and C3 binding, whereas IgG4 do not activate the complement system and do not bind C1q and C3.
[0203] In one embodiment the antibody according to the invention comprises a Fc part derived from human origin and preferably all other parts of the human constant regions. As used herein the term "Fe part derived from human origin" denotes a Fc part which is either a Fc part of a human antibody of the subclass IgG1, IgG2, IgG3 or IgG4, preferably a Fc part from human IgG1 subclass, a mutated Fe part from human IgG1 subclass (preferably with a mutation on L234A+L235A), a Fc part from human IgG4 subclass or a mutated Fc part from human IgG4 subclass (preferably with a mutation on S228P). Mostly preferred are the human heavy chain constant regions of SEQ ID NO: 58 (human IgG1 subclass), SEQ ID NO: 59 (human IgG1 subclass with mutations L234A and L235A), SEQ ID NO: 60 human IgG4 subclass), or SEQ ID NO: 61 (human TgG4 subclass with mutation S228P).
[0204] The terms "nucleic acid" or "nucleic acid molecule", as used herein, are intended to include DNA molecules and RNA molecules. A nucleic acid molecule may be single-stranded or double-stranded, but preferably is double-stranded DNA.
[0205] Nucleic acid is "operably linked" when it is plated into a functional relationship with another nucleic acid sequence. For example, DNA for a presequence or secretory leader is operably linked to DNA for a polypeptide if it is expressed as a preprotein that participates in the secretion of the polypeptide; a promoter or enhancer is operably linked to a coding sequence if it affects the transcription of the sequence; or a ribosome binding site is operably linked to a coding sequence if it is positioned so as to facilitate translation. Generally, "operably linked" means that the DNA sequences being linked are contiguous, and, in the case of a secretory leader, contiguous and in reading frame. However, enhancers do not have to be contiguous. Linking is accomplished by ligation at convenient restriction sites. If such sites do not exist, the synthetic oligonucleotide adaptors or linkers are used in accordance with conventional practice.
[0206] The term "amino acid" as used within this application denotes the group of naturally occurring carboxy .alpha.-amino acids comprising alanine (three letter code: ala, one letter code: A), arginine (arg, R), asparagine (asn, N), aspartic acid (asp, D), cysteine (cys, C), glutamine (gin, Q), glutamic acid (glu, E), glycine (gly, G), histidine (his, H), isoleucine (ile, I), leucine (leu, L), lysine (lys, K), methionine (met, M), phenylalanine (phe, F), proline (pro, P), serine (ser, S), threonine (thr, T), tryptophan (trp, W), tyrosine (tyr, Y), and valine (val, V).
[0207] As used herein, the expressions "cell", "cell line", and "cell culture" are used interchangeably and all such designations include progeny. Thus, the words "transformants" and "transformed cells" include the primary subject cell and cultures derived therefrom without regard for the number of transfers. It is also understood that all progeny may not be precisely identical in DNA content, due to deliberate or inadvertent mutations. Variant progeny that have the same function or biological activity as screened for in the originally transformed cell are included.
[0208] An "effective amount" refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired therapeutic or prophylactic result.
[0209] A "therapeutically effective amount" of a substance/molecule of the invention may vary according to factors such as the disease state, age, sex, and weight of the individual, and the ability of the substance/molecule, to elicit a desired response in the individual. A therapeutically effective amount encompasses an amount in which any toxic or detrimental effects of the substance/molecule are outweighed by the therapeutically beneficial effects. A therapeutically effective amount also encompasses an amount sufficient to confer benefit, e.g., clinical benefit.
[0210] A "prophylactically effective amount" refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired prophylactic result. Typically, but not necessarily, since a prophylactic dose is used in subjects prior to or at an earlier stage of disease, the prophylactically effective amount would be less than the therapeutically effective amount. A prophylactically effective amount encompasses an amount sufficient to confer benefit, e.g., clinical benefit.
[0211] In the case of pre-cancerous, benign, early or late-stage tumors, the therapeutically effective amount of the angiogenic inhibitor may reduce the number of cancer cells; reduce the primary tumor size; inhibit (i.e., slow to some extent and preferably stop) cancer cell infiltration into peripheral organs; inhibit (i.e., slow to some extent and preferably stop) tumor metastasis; inhibit or delay, to some extent, tumor growth or tumor progression; and/or relieve to some extent one or more of the symptoms associated with the disorder. To the extent the drug may prevent growth and/or kill existing cancer cells, it may be cytostatic and/or cytotoxic. For cancer therapy, efficacy in vivo can, for example, be measured by assessing the duration of survival, time to disease progression (TTP), the response rates (RR), duration of response, and/or quality of life.
[0212] To "reduce" or "inhibit" is to decrease or reduce an activity, function, and/or amount as compared to a reference. In certain embodiments, by "reduce" or "inhibit" is meant the ability to cause an overall decrease of 20% or greater. In another embodiment, by "reduce" or "inhibit" is meant the ability to cause an overall decrease of 50% or greater. In yet another embodiment, by "reduce" or "inhibit" is meant the ability to cause an overall decrease of 75%, 85%, 90%, 95%, or greater. Reduce or inhibit can refer to the symptoms of the disorder being treated, the presence or size of metastases, the size of the primary tumor, or the size or number of the blood vessels in angiogenic disorders.
[0213] A "disorder" is any condition that would benefit from treatment including, but not limited to, chronic and acute disorders or diseases including those pathological conditions which predispose the mammal to the disorder in question. Disorders include cancer (including metastatic cancer), bone disorders (including, e.g., bone loss such as osteoporosis), and inflammatory disorders.
III. Antibodies and Methods of the Invention
[0214] The invention comprises an antibody binding to human CSF-1R, wherein the antibody binds to human CSF-1R fragment delD4 (comprising the extracellular subdomains D1-D3 and D5) (SEQ ID NO: 65) and to human CSF-1R Extracellular Domain (CSF-1R-ECD) (comprising the extracellular subdomains D1-D5) (SEQ ID NO: 64) with a ratio of 1:50 or lower.
[0215] The invention further comprises an antibody comprising a heavy chain variable domain CDR3 region comprising SEQ ID NO: 1, SEQ ID NO: 9, SEQ ID NO:23, SEQ ID NO:31, SEQ ID NO:39, SEQ ID NO:47 or SEQ ID NO:55.
[0216] The invention further comprises an antibody wherein [0217] a) the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, [0218] b) the heavy chain variable domain comprises SEQ ID NO:15 and the light chain variable domain comprises SEQ ID NO: 16; [0219] or a humanized version thereof.
[0220] The invention further comprises an antibody wherein [0221] a) the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, [0222] b) the heavy chain variable domain comprises SEQ ID NO:15 and the light chain variable domain comprises SEQ ID NO: 16; [0223] c) the heavy chain variable domain comprises SEQ ID NO:75 and the light chain variable domain comprises SEQ ID NO:76; [0224] d) the heavy chain variable domain comprises SEQ ID NO:83 and the light chain variable domain comprises SEQ ID NO:84;
[0225] or a humanized version thereof.
[0226] The invention further comprises an antibody wherein [0227] the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, or a humanized version thereof.
[0228] In one embodiment the antibody comprises [0229] a) a heavy chain variable domain comprising SEQ ID NO:23 and a light chain variable domain comprising SEQ ID NO:24, or [0230] b) a heavy chain variable domain comprising SEQ ID NO:31 and a light chain variable domain comprising SEQ ID NO:32, or [0231] c) a heavy chain variable domain comprising SEQ ID NO:39 and a light chain variable domain comprising SEQ ID NO:40, or [0232] d) a heavy chain variable domain comprising SEQ ID NO:47 and a light chain variable domain comprising SEQ ID NO:48, or [0233] e) a heavy chain variable domain comprising SEQ ID NO:55 and a light chain variable domain comprising SEQ ID NO:56.
[0234] In one embodiment the antibody comprises [0235] a) a heavy chain variable domain comprising SEQ ID NO:23 and a light chain variable domain comprising SEQ ID NO:24, or [0236] b) a heavy chain variable domain comprising SEQ ID NO:31 and a light chain variable domain comprising SEQ ID NO:32, or [0237] c) a heavy chain variable domain comprising SEQ ID NO:39 and a light chain variable domain comprising SEQ ID NO:40, or [0238] d) a heavy chain variable domain comprising SEQ ID NO:47 and a light chain variable domain comprising SEQ ID NO:48.
[0239] In one embodiment the antibody comprises [0240] a heavy chain variable domain comprising SEQ ID NO:23 and a light chain variable domain comprising SEQ ID NO:24, or
[0241] In one embodiment the antibody comprises [0242] a heavy chain variable domain comprising SEQ ID NO:31 and a light chain variable domain comprising SEQ ID NO:32.
[0243] In one embodiment the antibody comprises [0244] a heavy chain variable domain comprising SEQ ID NO:39 and a light chain variable domain comprising SEQ ID NO:40.
[0245] In one embodiment the antibody comprises [0246] a heavy chain variable domain comprising SEQ ID NO:47 and a light chain variable domain comprising SEQ ID NO:48.
[0247] The invention further comprises an antibody wherein [0248] the heavy chain variable domain comprises SEQ ID NO:15 and the light chain variable domain comprises SEQ ID NO: 16, or a humanized version thereof.
[0249] The invention further comprises an antibody wherein [0250] the heavy chain variable domain comprises SEQ ID NO:75 and the light chain variable domain comprises SEQ ID NO:76;
[0251] or a humanized version thereof.
[0252] The invention further comprises an antibody wherein [0253] the heavy chain variable domain comprises SEQ ID NO:83 and the light chain variable domain comprises SEQ ID NO:84;
[0254] or a humanized version thereof.
[0255] The invention further comprises an antibody, wherein [0256] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:1, a CDR2 region comprising SEQ ID NO: 2, and a CDR1 region comprising SEQ ID NO:3, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 4, a CDR2 region comprising SEQ ID NO:5, and a CDR1 region comprising SEQ ID NO:6, or, [0257] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 9, a CDR2 region comprising SEQ ID NO: 10, and a CDR1 region comprising SEQ ID NO: 11, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:12, a CDR2 region comprising SEQ ID NO: 13, and a CDR1 region comprising SEQ ID NO: 14, or [0258] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0259] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0260] e) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0261] f) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46, or [0262] g) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 49, a CDR2 region comprising SEQ ID NO: 50, and a CDR1 region comprising SEQ ID NO: 51, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:52, a CDR2 region comprising SEQ ID NO: 53, and a CDR1 region comprising SEQ ID NO: 54.
[0263] The invention further comprises an antibody, wherein [0264] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:1, a CDR2 region comprising SEQ ID NO: 2, and a CDR1 region comprising SEQ ID NO:3, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 4, a CDR2 region comprising SEQ ID NO:5, and a CDR1 region comprising SEQ ID NO:6, or, [0265] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 9, a CDR2 region comprising SEQ ID NO: 10, and a CDR1 region comprising SEQ ID NO: 11, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:12, a CDR2 region comprising SEQ ID NO: 13, and a CDR1 region comprising SEQ ID NO: 14, or [0266] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0267] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0268] e) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0269] f) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46, [0270] g) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 49, a CDR2 region comprising SEQ ID NO: 50, and a CDR1 region comprising SEQ ID NO: 51, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:52, a CDR2 region comprising SEQ ID NO: 53, and a CDR1 region comprising SEQ ID NO: 54; [0271] h) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:69, a CDR2 region comprising SEQ ID NO: 70, and a CDR1 region comprising SEQ ID NO:71, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 72, a CDR2 region comprising SEQ ID NO:73, and a CDR1 region comprising SEQ ID NO:74, or [0272] i) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 77, a CDR2 region comprising SEQ ID NO: 78, and a CDR1 region comprising SEQ ID NO: 79, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:80, a CDR2 region comprising SEQ ID NO: 81, and a CDR1 region comprising SEQ ID NO: 82.
[0273] In one embodiment the antibody comprises [0274] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:69, a CDR2 region comprising SEQ ID NO: 70, and a CDR1 region comprising SEQ ID NO:71, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 72, a CDR2 region comprising SEQ ID NO:73, and a CDR1 region comprising SEQ ID NO:74, or [0275] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 77, a CDR2 region comprising SEQ ID NO: 78, and a CDR1 region comprising SEQ ID NO: 79, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:80, a CDR2 region comprising SEQ ID NO: 81, and a CDR1 region comprising SEQ ID NO: 82.
[0276] In one embodiment the antibody comprises [0277] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0278] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ TD NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0279] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0280] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46, or [0281] e) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 49, a CDR2 region comprising SEQ ID NO: 50, and a CDR1 region comprising SEQ ID NO: 51, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:52, a CDR2 region comprising SEQ ID NO: 53, and a CDR1 region comprising SEQ ID NO: 54.
[0282] In one embodiment the antibody comprises [0283] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0284] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0285] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ 1D NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0286] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46.
[0287] In one embodiment the antibody comprises [0288] the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO: 19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22.
[0289] In one embodiment the antibody comprises [0290] the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30.
[0291] In one embodiment the antibody comprises [0292] the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38.
[0293] In one embodiment the antibody comprises [0294] the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46.
[0295] In one embodiment the antibody binding to human CSF-1R binds to human CSF-1R fragment delD4 (SEQ ID NO: 65) and to human CSF-1R-ECD (SEQ ID NO: 64) with a ratio of 1:50 or lower, further does not bind to human CSF-1R fragment D1-D3 (SEQ ID NO: 66).
[0296] One embodiment of the invention is a screening method for selecting antibodies according to the invention comprising the following steps: [0297] a) determining the binding signal (Response Units (RU)) of anti-CSF-1R antibodies to human CSF-1R fragment delD4 (SEQ ID NO: 65) and to human CSF-1R Extracellular Domain (CSF-1R-ECD) (SEQ ID NO: 64) by a Surface Plasmon Resonance assay (Biacore assay). [0298] b) selecting antibodies with ratio of the binding signals (human CSF-1R fragment delD4/human CSF-1R Extracellular Domain (CSF-1R-ECD)) of 50:1 or lower.
[0299] In one embodiment the determination is performed at 25.degree. C.
[0300] In one embodiment the screening method comprises as further steps the measuring of the binding of anti-CSF-1R antibodies to human CSF-1R fragment D1-D3 (SEQ ID NO: 66) (D1-D3) and the selecting of antibodies which show no binding to said fragment.
[0301] In one embodiment the antibodies according to the invention inhibit CSF-1 binding to CSF-1R. In one embodiment with an IC50 of 200 ng/ml or lower, in one embodiment with an IC50 of 50 ng/ml or lower. The 1C50 of inhibition of CSF-1 binding to CSF-1R can be determined as shown in Example 2.
[0302] In one embodiment the antibodies according to the invention inhibit CSF-1-induced CSF-1R phosphorylation (in NIH3T3-CSF-1R recombinant cells).
[0303] In one embodiment with an IC50 of 800 ng/ml or lower, in one embodiment with an IC50 of 600 ng/ml or lower, in one embodiment with an IC50 of 250 ng/ml or lower. The IC50 of CSF-1-induced CSF-1R phosphorylation can be determined as shown in Example 3.
[0304] In one embodiment the antibodies according to the invention inhibit the growth of recombinant NIH3T3 cells expressing human CSF-1R (SEQ ID No: 62). In one embodiment with an IC50 of 10 .mu.g/ml or lower, in one embodiment with an IC50 of 5 .mu.g/ml or lower, in one embodiment with an IC50 of 2 .mu.g/ml or lower. In one embodiment with an IC30 of 10 .mu.g/ml or lower, in one embodiment with an IC30 of 5 .mu.g/ml or lower, in one embodiment with an IC30 of 2 .mu.g/ml or lower. The IC50 value, the IC30 value or the % growth inhibition is determined as shown in Example 5.
[0305] In one embodiment the antibodies according to the invention inhibit the growth of recombinant NIH3T3 cells expressing human mutant CSF-1R L301S Y969F (SEQ ID No: 63). In one embodiment with an IC50 of 15 gig/ml or lower, in one embodiment with an IC50 of 10 .mu.g/ml or lower. In one embodiment with an IC30 of 10 .mu.g/ml or lower, in one embodiment with an IC50 of 5 .mu.g/ml ng/ml or lower; in one embodiment with an IC50 of 2 .mu.g/ml or lower. The IC50 value, the IC30 value or the % growth inhibition is determined as shown in Example 5.
[0306] In one embodiment the antibodies according to the invention inhibit the growth of BeWo tumor cells (ATCC CCL-98) by 65% or more (at an antibody concentration of 10 .mu.g/ml; and as compared to the absence of antibody). The % growth inhibition is determined as shown in Example 8. E.g. Mab 2F11 shows a growth inhibition of BeWo tumor cells of 70%.
[0307] In one embodiment the antibodies according to the invention inhibit (both) human and cynomolgous macrophage differentiation (which is indicated by the inhibition of the survival of human and cynomolgous monocytes as shown in Examples 7 and 8). In one embodiment the antibodies according to the invention inhibit the survival of human monocytes with an IC50 of 0.15 .mu.g/ml or lower, in on embodiment with an IC50 of 0.10 .mu.g/ml or lower. The inhibition of the survival of human monocytes is determined as shown in Example 7. In one embodiment the antibodies according to the invention inhibit the survival of cynomolgous monocytes by 80% or more, in one embodiment by 90% or more (at an antibody concentration of 5 .mu.g/ml; and as compared to the absence of antibody). The inhibition of the survival of human monocytes is determined as shown in Example 8.
[0308] A further embodiment of the invention is a method for the production of an antibody against CSF-1R wherein the sequence of a nucleic acid encoding the heavy chain of a human IgG1 class antibody binding to human CSF-1R according to the invention said modified nucleic acid and the nucleic acid encoding the light chain of said antibody are inserted into an expression vector, said vector is inserted in a eukaryotic host cell, the encoded protein is expressed and recovered from the host cell or the supernatant.
[0309] The antibodies according to the invention are preferably produced by recombinant means. Therefore the antibody is preferably an isolated monoclonal antibody. Such recombinant methods are widely known in the state of the art and comprise protein expression in prokaryotic and eukaryotic cells with subsequent isolation of the antibody polypeptide and usually purification to a pharmaceutically acceptable purity. For the protein expression, nucleic acids encoding light and heavy chains or fragments thereof are inserted into expression vectors by standard methods. Expression is performed in appropriate prokaryotic or eukaryotic host cells like CHO cells, NS0 cells, SP2/0 cells, HEK293 cells, COS cells, yeast, or E. coli cells, and the antibody is recovered from the cells (supernatant or cells after lysis).
[0310] Recombinant production of antibodies is well-known in the state of the art and described, for example, in the review articles of Makrides, S. C., Protein Expr. Purif. 17 (1999) 183-202; Geisse, S., et al., Protein Expr. Purif. 8 (1996) 271-282; Kaufman, R. J., Mol. Biotechnol. 16 (2000) 151-161; Werner, R. G., Drug Res. 48 (1998) 870-880.
[0311] The antibodies may be present in whole cells, in a cell lysate, or in a partially purified or substantially pure form. Purification is performed in order to eliminate other cellular components or other contaminants, e.g. other cellular nucleic acids or proteins, by standard techniques, including alkaline/SDS treatment, CsCl banding, column chromatography, agarose gel electrophoresis, and others well known in the art. See Ausubel, F., et al., ed. Current Protocols in Molecular Biology, Greene Publishing and Wiley Interscience, New York (1987).
[0312] The monoclonal antibodies are suitably separated from the culture medium by conventional immunoglobulin purification procedures such as, for example, protein A-Sepharose, hydroxylapatite chromatography, gel electrophoresis, dialysis, or affinity chromatography. DNA and RNA encoding the monoclonal antibodies are readily isolated and sequenced using conventional procedures. The hybridoma cells can serve as a source of such DNA and RNA. Once isolated, the DNA may be inserted into expression vectors, which are then transfected into host cells such as HEK 293 cells. CHO cells, or myeloma cells that do not otherwise produce immunoglobulin protein, to obtain the synthesis of recombinant monoclonal antibodies in the host cells.
[0313] Expression in NS0 cells is described by, e.g., Barnes, L. M., et al., Cytotechnology 32 (2000) 109-123; and Barnes, L. M., et al., Biotech. Bioeng. 73 (2001) 261-270. Transient expression is described by, e.g., Durocher, Y., et al., Nucl. Acids. Res. 30 (2002) E9. Cloning of variable domains is described by Orlandi, R., et al., Proc. Natl. Acad. Sci. USA 86 (1989) 3833-3837; Carter, P., et al., Proc. Natl. Acad. Sci. USA 89 (1992) 4285-4289; and Norderhaug, L., et al., J. Immunol. Methods 204 (1997) 77-87. A preferred transient expression system (HEK 293) is described by Schlaeger, E.-J., and Christensen, K., in Cytotechnology 30 (1999) 71-83 and by Schlaeger, E.-J., in J. Immunol. Methods 194 (1996) 191-199.
[0314] The control sequences that are suitable for prokaryotes, for example, include a promoter, optionally an operator sequence, and a ribosome binding site. Eukaryotic cells are known to utilize promoters, enhancers and polyadenylation signals.
[0315] In some embodiments, the antibody according to the invention is of human IgG1 subclass or of human IgG4 subclass. In one embodiment the antibody according to the invention is of human IgG1 subclass. In one embodiment the antibody according to the invention is of human IgG4 subclass.
[0316] In one embodiment the constant chains are of human origin. Such constant chains are well known in the state of the art and e.g. described by Kabat, E. A., (see e.g. Johnson, G. and Wu, T. T., Nucleic Acids Res. 28 (2000) 214-218). For example, a useful human heavy chain constant region comprises an amino acid sequence of SEQ ID NO: 58. For example, a useful human light chain constant region comprises an amino acid sequence of a kappa-light chain constant region of SEQ ID NO: 57.
[0317] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0318] a) the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, [0319] b) the heavy chain variable domain comprises SEQ ID NO: 15 and the light chain variable domain comprises SEQ ID NO: 16; [0320] or a humanized version thereof.
[0321] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0322] a) the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, [0323] b) the heavy chain variable domain comprises SEQ ID NO: 15 and the light chain variable domain comprises SEQ ID NO: 16; [0324] c) the heavy chain variable domain comprises SEQ ID NO:75 and the light chain variable domain comprises SEQ ID NO:76; [0325] d) the heavy chain variable domain comprises SEQ ID NO:83 and the light chain variable domain comprises SEQ ID NO:84; [0326] or a humanized version thereof.
[0327] Another aspect of the invention is an antibody binding to human CSF-1R, wherein the heavy chain variable domain comprises SEQ ID NO:7 and the light chain variable domain comprises SEQ ID NO:8, or a humanized version thereof.
[0328] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0329] a) the heavy chain variable domain comprises SEQ ID NO:23 and the light chain variable domain comprises SEQ ID NO:24, or [0330] b) the heavy chain variable domain comprises SEQ ID NO:31 and the light chain variable domain comprises SEQ ID NO:32, or [0331] c) the heavy chain variable domain comprises SEQ ID NO:39 and the light chain variable domain comprises SEQ ID NO:40, or [0332] d) the heavy chain variable domain comprises SEQ ID NO:47 and the light chain variable domain comprises SEQ ID NO:48, or [0333] e) the heavy chain variable domain comprises SEQ ID NO:55 and the light chain variable domain comprises SEQ ID NO:56.
[0334] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0335] a) the heavy chain variable domain comprises SEQ ID NO:23 and the light chain variable domain comprises SEQ ID NO:24, or [0336] b) the heavy chain variable domain comprises SEQ ID NO:31 and the light chain variable domain comprises SEQ ID NO:32, or [0337] c) the heavy chain variable domain comprises SEQ ID NO:39 and the light chain variable domain comprises SEQ ID NO:40, or [0338] d) the heavy chain variable domain comprises SEQ ID NO:47 and the light chain variable domain comprises SEQ ID NO:48.
[0339] Another aspect of the invention is an antibody binding to human CSF-1R, wherein the heavy chain variable domain comprises SEQ ID NO:23 and the light chain variable domain comprises SEQ ID NO:24, or Another aspect of the invention is an antibody binding to human CSF-1R, wherein the heavy chain variable domain comprises SEQ ID NO:31 and the light chain variable domain comprises SEQ ID NO:32.
[0340] Another aspect of the invention is an antibody binding to human CSF-1R, wherein the heavy chain variable domain comprises SEQ ID NO:39 and the light chain variable domain comprises SEQ ID NO:40.
[0341] Another aspect of the invention is an antibody binding to human CSF-1R, wherein the heavy chain variable domain comprises SEQ ID NO:47 and the light chain variable domain comprises SEQ ID NO:48.
[0342] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0343] the heavy chain variable domain comprises SEQ ID NO:15 and the light chain variable domain comprises SEQ ID NO:16, or a humanized version thereof.
[0344] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0345] the heavy chain variable domain comprises SEQ ID NO:75 and the light chain variable domain comprises SEQ ID NO:76;
[0346] or a humanized version thereof.
[0347] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0348] the heavy chain variable domain comprises SEQ ID NO:83 and the light chain variable domain comprises SEQ ID NO:84;
[0349] or a humanized version thereof.
[0350] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0351] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:1, a CDR2 region comprising SEQ ID NO: 2, and a CDR1 region comprising SEQ ID NO:3, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 4, a CDR2 region comprising SEQ ID NO:5, and a CDR1 region comprising SEQ ID NO:6, or, [0352] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 9, a CDR2 region comprising SEQ ID NO: 10, and a CDR1 region comprising SEQ ID NO: 11, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:12, a CDR2 region comprising SEQ ID NO: 13, and a CDR1 region comprising SEQ ID NO: 14, or [0353] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0354] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0355] e) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0356] f) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46, [0357] g) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 49, a CDR2 region comprising SEQ ID NO: 50, and a CDR1 region comprising SEQ ID NO: 51, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:52, a CDR2 region comprising SEQ ID NO: 53, and a CDR1 region comprising SEQ ID NO: 54; [0358] h) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:69, a CDR2 region comprising SEQ ID NO: 70, and a CDR1 region comprising SEQ ID NO:71, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 72, a CDR2 region comprising SEQ ID NO:73, and a CDR1 region comprising SEQ ID NO:74, or [0359] i) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 77, a CDR2 region comprising SEQ ID NO: 78, and a CDR1 region comprising SEQ ID NO: 79, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:80, a CDR2 region comprising SEQ ID NO: 81, and a CDR1 region comprising SEQ ID NO: 82.
[0360] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0361] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0362] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0363] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0364] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46, or [0365] e) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 49, a CDR2 region comprising SEQ ID NO: 50, and a CDR1 region comprising SEQ ID NO: 51, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:52, a CDR2 region comprising SEQ ID NO: 53, and a CDR1 region comprising SEQ ID NO: 54.
[0366] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0367] a) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO:19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22, or [0368] b) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30, or [0369] c) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38, or [0370] d) the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46.
[0371] Another aspect of the invention is an antibody binding to human CSF-1R, wherein the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 17, a CDR2 region comprising SEQ ID NO: 18, and a CDR1 region comprising SEQ ID NO: 19, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 20, a CDR2 region comprising SEQ ID NO:21, and a CDR1 region comprising SEQ ID NO:22.
[0372] Another aspect of the invention is an antibody binding to human CSF-1R, wherein the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 25, a CDR2 region comprising SEQ ID NO: 26, and a CDR1 region comprising SEQ ID NO: 27, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:28, a CDR2 region comprising SEQ ID NO: 29, and a CDR1 region comprising SEQ ID NO: 30.
[0373] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0374] the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO: 33, a CDR2 region comprising SEQ ID NO: 34, and a CDR1 region comprising SEQ ID NO: 35, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO:36, a CDR2 region comprising SEQ ID NO: 37, and a CDR1 region comprising SEQ ID NO: 38.
[0375] Another aspect of the invention is an antibody binding to human CSF-1R, wherein [0376] the heavy chain variable domain comprises a CDR3 region comprising SEQ ID NO:41, a CDR2 region comprising SEQ ID NO: 42, and a CDR1 region comprising SEQ ID NO:43, and the light chain variable domain comprises a CDR3 region comprising SEQ ID NO: 44, a CDR2 region comprising SEQ ID NO:45, and a CDR1 region comprising SEQ ID NO:46.
[0377] The invention comprises a method for the treatment of a patient in need of therapy, comprising administering to the patient a therapeutically effective amount of an antibody according to the invention.
[0378] The invention comprises the use of an antibody according to the invention for therapy.
[0379] One preferred embodiment of the invention are the CSF-1R antibodies of the present invention for use in the treatment of "CSF-1R mediated diseases" or the CSF-1R antibodies of the present invention for use for the manufacture of a medicament in the treatment of "CSF-1R mediated diseases", which can be described as follows:
[0380] There are 3 distinct mechanisms by which CSF-1R signaling is likely involved in tumor growth and metastasis. The first is that expression of CSF-ligand and receptor has been found in tumor cells originating in the female reproductive system (breast, ovarian, endometrium, cervical) (Scholl, S. M., et al., J. Natl. Cancer Inst. 86 (1994) 120-126; Kacinski, B. M., Mol. Reprod. Dev. 46 (1997) 71-74; Ngan, H. Y., et al., Eur. J. Cancer 35 (1999) 1546-1550; Kirma, N., et al., Cancer Res 67 (2007) 1918-1926) and the expression has been associated with breast cancer xenograft growth as well as poor prognosis in breast cancer patients. Two point mutations were seen in CSF-1R in about 10-20% of acute myelocytic leukemia, chronic myelocytic leukemia and myelodysplasia patients tested in one study, and one of the mutations was found to disrupt receptor turnover (Ridge, S. A., et al., Proc. Natl. Acad. Sci USA 87 (1990) 1377-1380). However the incidence of the mutations could not be confirmed in later studies (Abu-Duhier, F. M., et al., Br. J. Haematol. 120 (2003) 464-470). Mutations were also found in some cases of hepatocellular cancer (Yang, D. H., et al., Hepatobiliary Pancreat. Dis. Int. 3 (2004) 86-89) and idiopathic myelofibrosis (Abu-Duhier, F. M., et al., Br. J. Haematol. 120 (2003) 464-470). Recently, in the GDM-1 cell line derived from a patient with myelomonoblastic leukemia the Y571D mutation in CSF-1R was identified (Chase, A., et al., Leukemia 23 (2009) 358-364).
[0381] Pigmented villonodular synovitis (PVNS) and Tenosynovial Giant cell tumors (TGCT) can occur as a result of a translocation that fuses the M-CSF gene to a collagen gene COL6A3 and results in overexpression of M-CSF (West, R. B., et al., Proc. Natl. Acad. Sci. USA 103 (2006) 690-695). A landscape effect is proposed to be responsible for the resulting tumor mass that consists of monocytic cells attracted by cells that express M-CSF. TGCTs are smaller tumors that can be relatively easily removed from fingers where they mostly occur. PVNS is more aggressive as it can recur in large joints and is not as easily controlled surgically.
[0382] The second mechanism is based on blocking signaling through M-CSF/CSF-1R at metastatic sites in bone which induces osteoclastogenesis, bone resorption and osteolytic bone lesions. Breast, multiple myeloma and lung cancers are examples of cancers that have been found to metastasize to the bone and cause osteolytic bone disease resulting in skeletal complications. M-CSF released by tumor cells and stroma induces the differentiation of hematopoietic myeloid monocyte progenitors to mature osteoclasts in collaboration with the receptor activator of nuclear factor kappa-B ligand-RANKL. During this process, M-CSF acts as a permissive factor by giving the survival signal to osteoclasts (Tanaka, S., et al., J. Clin. Invest. 91 (1993) 257-263). Inhibition of CSF-1R activity during osteoclast differentiation and maturation with a anti-CSF-1R antibody is likely to prevent unbalanced activity of osteoclasts that cause osteolytic disease and the associated skeletal related events in metastatic disease. Whereas breast, lung cancer and multiple myeloma typically result in osteolytic lesions, metastasis to the bone in prostate cancer initially has an osteoblastic appearance in which increased bone forming activity results in `woven bone` which is different from typical lamellar structure of normal bone. During disease progression bone lesions display a significant osteolytic component as well as high serum levels of bone resorption and suggests that anti-resorptive therapy may be useful. Bisphosphonates have been shown to inhibit the formation of osteolytic lesions and reduced the number of skeletal-related events only in men with hormone-refractory metastatic prostate cancer but at this point their effect on osteoblastic lesions is controversial and bisphosphonates have not been beneficial in preventing bone metastasis or hormone responsive prostate cancer to date. The effect of anti-resorptive agents in mixed osteolytic/osteoblastic prostate cancer is still being studied in the clinic (Choueiri, M. B., et al., Cancer Metastasis Rev. 25 (2006) 601-609; Vessella, R. L. and Corey, E., Clin. Cancer Res. 12 (20 Pt 2) (2006) 6285s-6290s).
[0383] The third mechanism is based on the recent observation that tumor associated macrophages (TAM) found in solid tumors of the breast, prostate, ovarian and cervical cancers correlated with poor prognosis (Bingle, L., et al., J. Pathol. 196 (2002) 254-265; Pollard, J. W., Nat. Rev. Cancer 4 (2004) 71-78). Macrophages are recruited to the tumor by M-CSF and other chemokines. The macrophages can then contribute to tumor progression through the secretion of angiogenic factors, proteases and other growth factors and cytokines and may be blocked by inhibition of CSF-1R signaling. Recently it was shown by Zins et al (Zins, K., et al., Cancer Res. 67 (2007) 1038-1045) that expression of siRNA of Tumor necrosis factor alpha (TNF alpha), M-CSF or the combination of both would reduce tumor growth in a mouse xenograft model between 34% and 50% after intratumoral injection of the respective siRNA. SiRNA targeting the TNF alpha secreted by the human SW620 cells reduced mouse M-CSF levels and led to reduction of macrophages in the tumor. In addition treatment of MCF7 tumor xenografts with an antigen binding fragment directed against M-CSF did result in 40% tumor growth inhibition, reversed the resistance to chemotherapeutics and improved survival of the mice when given in combination with chemotherapeutics (Paulus, P., et al., Cancer Res. 66 (2006) 4349-4356).
[0384] TAMs are only one example of an emerging link between chronic inflammation and cancer. There is additional evidence for a link between inflammation and cancer as many chronic diseases are associated with an increased risk of cancer, cancers arise at sites of chronic inflammation, chemical mediators of inflammation are found in many cancers; deletion of the cellular or chemical mediators of inflammation inhibits development of experimental cancers and long-term use of anti-inflammatory agents reduce the risk of some cancers. A link to cancer exists for a number of inflammatory conditions among-those H. pylori induced gastritis for gastric cancer, Schistosomiasis for bladder cancer, HHVX for Kaposi's sarcoma, endometriosis for ovarian cancer and prostatitis for prostate cancer (Balkwill, F., et al., Cancer Cell 7 (2005) 211-217). Macrophages are key cells in chronic inflammation and respond differentially to their microenvironment. There are two types of macrophages that are considered extremes in a continuum of functional states: M1 macrophages are involved in Type 1 reactions. These reactions involve the activation by microbial products and consequent killing of pathogenic microorganisms that result in reactive oxygen intermediates. On the other end of the extreme are M2 macrophages involved in Type 2 reactions that promote cell proliferation, tune inflammation and adaptive immunity and promote tissue remodeling, angiogenesis and repair (Mantovani, A., et al., Trends Immunol. 25 (2004) 677-686). Chronic inflammation resulting in established neoplasia is usually associated with M2 macrophages. A pivotal cytokine that mediates inflammatory reactions is TNF alpha that true to its name can stimulate anti-tumor immunity and hemorrhagic necrosis at high doses but has also recently been found to be expressed by tumor cells and acting as a tumor promoter (Zins, K., et al., Cancer Res. 67 (2007) 1038-1045; Balkwill, F., Cancer Metastasis Rev. 25 (2006) 409-416). The specific role of macrophages with respect to the tumor still needs to be better understood including the potential spatial and temporal dependence on their function and the relevance to specific tumor types.
[0385] Thus one embodiment of the invention are the CSF-1R antibodies of the present invention for use in the treatment of cancer. The term "cancer" as used herein may be, for example, lung cancer, non small cell lung (NSCL) cancer, bronchioloalviolar cell lung cancer, bone cancer, pancreatic cancer, skin cancer, cancer of the head or neck, cutaneous or intraocular melanoma, uterine cancer, ovarian cancer, rectal cancer, cancer of the anal region, stomach cancer, gastric cancer, colon cancer, breast cancer, uterine cancer, carcinoma of the fallopian tubes, carcinoma of the endometrium, carcinoma of the cervix, carcinoma of the vagina, carcinoma of the vulva, Hodgkin's Disease, cancer of the esophagus, cancer of the small intestine, cancer of the endocrine system, cancer of the thyroid gland, cancer of the parathyroid gland, cancer of the adrenal gland, sarcoma of soft tissue, cancer of the urethra, cancer of the penis, prostate cancer, cancer of the bladder, cancer of the kidney or ureter, renal cell carcinoma, carcinoma of the renal pelvis, mesothelioma, hepatocellular cancer, biliary cancer, neoplasms of the central nervous system (CNS), spinal axis tumors, brain stem glioma, glioblastoma multiforme, astrocytomas, schwanomas, ependymonas, medulloblastomas, meningiomas, squamous cell carcinomas, pituitary adenoma, lymphoma, lymphocytic leukemia, including refractory versions of any of the above cancers, or a combination of one or more of the above cancers. Preferably such cancer is a breast cancer, ovarian cancer, cervical cancer, lung cancer or prostate cancer. Preferably such cancers are further characterized by CSF-1 or CSF-1R expression or overexpression. One further embodiment the invention are the CSF-1R antibodies of the present invention for use in the simultaneous treatment of primary tumors and new metastases.
[0386] Thus another embodiment of the invention are the CSF-1R antibodies of the present invention for use in the treatment of periodontitis, histiocytosis X, osteoporosis, Paget's disease of bone (PDB), bone loss due to cancer therapy, periprosthetic osteolysis, glucocorticoid-induced osteoporosis, rheumatoid arthritis, psiratic arthritis, osteoarthritis, inflammatory arthridities, and inflammation.
[0387] Rabello, D., et al., Biochem. Biophys. Res. Commun. 347 (2006) 791-796 has demonstrated that SNPs in the CSF1 gene exhibited a positive association with aggressive periodontitis: an inflammatory disease of the periodontal tissues that causes tooth loss due to resorption of the alveolar bone.
[0388] Histiocytosis X (also called Langerhans cell histiocytosis, LCH) is a proliferative disease of Langerhans dendritic cells that appear to differentiate into osteoclasts in bone and extra osseous LCH lesions. Langerhans cells are derived from circulating monocytes. Increased levels of M-CSF that have been measured in scra and lesions where found to correlate with disease severity (da Costa, C. E., et al., J. Exp. Med. 201 (2005) 687-693). The disease occurs primarily in a pediatric patient population and has to be treated with chemotherapy when the disease becomes systemic or is recurrent.
[0389] The pathophysiology of osteoporosis is mediated by loss of bone forming osteoblasts and increased osteoclast dependent bone resorption. Supporting data has been described by Cenci et al showing that an anti-M-CSF antibody injection preserves bone density and inhibits bone resorption in ovariectomized mice (Cenci, S., et al., J. Clin. Invest. 105 (2000) 1279-1287). Recently a potential link between postmenopausal bone loss due to estrogen deficiency was identified and found that the presence of TNF alpha producing T-cell affected bone metabolism (Roggia, C., et al., Minerva Med. 95 (2004) 125-132). A possible mechanism could be the induction of M-CSF by TNF alpha in vivo. An important role for M-CSF in TNF-alpha-induced osteoclastogenesis was confirmed by the effect of an antibody directed against M-CSF that blocked the TNF alpha induced osteolysis in mice and thereby making inhibitors of CSF-1R signaling potential targets for inflammatory arthritis (Kitaura, H., et al., J. Clin. Invest. 115 (2005) 3418-3427).
[0390] Paget's disease of bone (PDB) is the second most common bone metabolism disorder after osteoporosis in which focal abnormalities of increased bone turnover lead to complications such as bone pain, deformity, pathological fractures and deafness. Mutations in four genes have been identified that regulate normal osteoclast function and predispose individuals to PDB and related disorders: insertion mutations in TNFRSF11A, which encodes receptor activator of nuclear factor (NF) kappaB (RANK)--a critical regulator of osteoclast function, inactivating mutations of TNFRSF11B which encodes osteoprotegerin (a decoy receptor for RANK ligand), mutations of the sequestosome 1 gene (SQSTM1), which encodes an important scaffold protein in the NFkappaB pathway and mutations in the valosin-containing protein (VCP) gene. This gene encodes VCP, which has a role in targeting the inhibitor of NFkappaB for degradation by the proteasome (Daroszewska, A. and Ralston, S. H., Nat. Clin. Pract. Rheumatol. 2 (2006) 270-277). Targeted CSF-1R inhibitors provide an opportunity to block the deregulation of the RANKL signaling indirectly and add an additional treatment option to the currently used bisphosphonates.
[0391] Cancer therapy induced bone loss especially in breast and prostate cancer patients is an additional indication where a targeted CSF-1R inhibitor could prevent bone loss (Lester, J. E., et al., Br. J. Cancer 94 (2006) 30-35). With the improved prognosis for early breast cancer the long-term consequences of the adjuvant therapies become more important as some of the therapies including chemotherapy, irradiation, aromatase inhibitors and ovary ablation affect bone metabolism by decreasing the bone mineral density, resulting in increased risk for osteoporosis and associated fractures (Lester, J. E., et al., Br. J. Cancer 94 (2006) 30-35). The equivalent to adjuvant aromatase inhibitor therapy in breast cancer is androgen ablation therapy in prostate cancer which leads to loss of bone mineral density and significantly increases the risk of osteoporosis-related fractures (Stoch, S. A., et al., J. Clin. Endocrinol. Metab. 86 (2001) 2787-2791).
[0392] Targeted inhibition of CSF-1R signaling is likely to be beneficial in other indications as well when targeted cell types include osteoclasts and macrophages e.g. treatment of specific complications in response to joint replacement as a consequence of rheumatoid arthritis. Implant failure due to periprosthetic bone loss and consequent loosing of prostheses is a major complication of joint replacement and requires repeated surgery with high socioeconomic burdens for the individual patient and the health-care system. To date, there is no approved drug therapy to prevent or inhibit periprosthetic osteolysis (Drees, P., et al., Nat. Clin. Pract. Rheumatol. 3 (2007) 165-171).
[0393] Glucocorticoid-induced osteoporosis (GIOP) is another indication in which a CSF-1R inhibitor could prevent bone loss after longterm glucocorticocosteroid use that is given as a result of various conditions among those chronic obstructive pulmonary disease, asthma and rheumatoid arthritis (Guzman-Clark, J. R., et al., Arthritis Rheum. 57 (2007) 140-146; Feldstein, A. C., et al., Osteoporos. Int. 16 (2005) 2168-2174).
[0394] Rheumatoid arthritis, psioratic arthritis and inflammatory arthridities are in itself potential indications for CSF-1R signaling inhibitors in that they consist of a macrophage component and to a varying degree bone destruction (Ritchlin, C. T., et al., J. Clin. Invest. 111 (2003) 821-831). Osteoarthritis and rheumatoid arthritis are inflammatory autoimmune disease caused by the accumulation of macrophages in the connective tissue and infiltration of macrophages into the synovial fluid, which is at least partially mediated by M-CSF. Campbell, I., K., et al., J. Leukoc. Biol. 68 (2000) 144-150, demonstrated that M-CSF is produced by human-joint tissue cells (chondrocytes, synovial fibroblasts) in vitro and is found in synovial fluid of patients with rheumatoid arthritis, suggesting that it contributes to the synovial tissue proliferation and macrophage infiltration which is associated with the pathogenesis of the disease. Inhibition of CSF-1R signaling is likely to control the number of macrophages in the joint and alleviate the pain from the associated bone destruction. In order to minimize adverse affects and to further understand the impact of the CSF-1R signaling in these indications, one method is to specifically inhibit CSF-1R without targeting a myriad other kinases, such as Raf kinase.
[0395] Recent literature reports correlate increased circulating M-CSF with poor prognosis and atherosclerotic progression in chronic coronary artery disease (Saitoh, T., et al., J. Am. Coll. Cardiol. 35 (2000) 655-665; Ikonomidis, I., et al., Eur. Heart. J. 26 (2005) p. 1618-1624); M-CSF influences the atherosclerotic process by aiding the formation of foam cells (macrophages with ingested oxidized LDL) that express CSF-1R and represent the initial plaque (Murayama, T., et al., Circulation 99 (1999) 1740-1746).
[0396] Expression and signaling of M-CSF and CSF-1R is found in activated microglia. Microglia, which are resident macrophages of the central nervous system, can be activated by various insults, including infection and traumatic injury. M-CSF is considered a key regulator of inflammatory responses in the brain and M-CSF levels increase in HIV-1, encephalitis, Alzheimer's disease (AD) and brain tumors. Microgliosis as a consequence of autocrine signaling by M-CSF/CSF-1R results in induction of inflammatory cytokines and nitric oxides being released as demonstrated by e.g. using an experimental neuronal damage model (Hao, A. J., et al., Neuroscience 112 (2002) 889-900; Murphy, G. M., Jr., et al., J. Biol. Chem. 273 (1998) 20967-20971). Microglia that have increased expression of CSF-1R are found to surround plaques in AD and in the amyloid precursor protein V717F transgenic mouse model of AD (Murphy, G. M., Jr., et al., Am. J. Pathol. 157 (2000) 895-904). On the other hand op/op mice with fewer microglia in the brain resulted in fibrilar deposition of A-beta and neuronal loss compared to normal control suggesting that microglia do have a neuroprotective function in the development of AD lacking in the op/op mice (Kaku, M., et al., Brain Res. Brain Res. Protoc. 12 (2003) 104-108).
[0397] Expression and signaling of M-CSF and CSF-1R is associated with inflammatory bowel disease (IBD) (WO 2005/046657). The term "inflammatory bowel disease" refers to serious, chronic disorders of the intestinal tract characterized by chronic inflammation at various sites in the gastrointestinal tract, and specifically includes ulcerative colitis (UC) and Crohn's disease. [0398] The invention comprises an antibody binding to human CSF-1R comprising the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for the treatment of cancer. [0399] The invention comprises an antibody binding to human CSF-1R comprising the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for the treatment of bone loss. [0400] The invention comprises an antibody binding to human CSF-1R comprising the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for the prevention or treatment of metastasis. [0401] The invention comprises an antibody binding to human CSF-1R comprising the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for treatment of inflammatory diseases. [0402] The invention comprises the use of an antibody comprising the antibody binding to human CSF-1R, wherein the antibody comprises the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for the treatment of cancer or alternatively for the manufacture of a medicament for the treatment of cancer. [0403] The invention comprises the use of an antibody comprising the antibody binding to human CSF-1R, wherein the antibody comprises the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for the treatment of bone loss or alternatively for the manufacture of a medicament for the treatment of bone loss. [0404] The invention comprises the use of an antibody comprising the antibody binding to human CSF-1R, wherein the antibody comprises the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for the prevention or treatment of metastasis or alternatively for the manufacture of a medicament for the prevention or treatment of metastasis. [0405] The invention comprises the use of an antibody comprising the antibody binding to human CSF-1R, wherein the antibody comprises the above mentioned epitope binding properties or alternatively by the above mentioned amino acid sequences and amino acid sequence fragments for treatment of inflammatory diseases or alternatively for the manufacture of a medicament for the treatment of inflammatory diseases.
[0406] A further embodiment of the invention is a method for the production of an antibody against CSF-1R wherein the sequence of a nucleic acid encoding the heavy chain of a human IgG1 class antibody binding to human CSF-1R according to the invention said modified nucleic acid and the nucleic acid encoding the light chain of said antibody are inserted into an expression vector, said vector is inserted in a eukaryotic host cell, the encoded protein is expressed and recovered from the host cell or the supernatant.
[0407] The antibodies according to the invention are preferably produced by recombinant means. Such methods are widely known in the state of the art and comprise protein expression in prokaryotic and eukaryotic cells with subsequent isolation of the antibody polypeptide and usually purification to a pharmaceutically acceptable purity. For the protein expression nucleic acids encoding light and heavy chains or fragments thereof are inserted into expression vectors by standard methods. Expression is performed in appropriate prokaryotic or eukaryotic host cells, such as CHO cells, NS0 cells, SP2/0 cells, HEK293 cells, COS cells, yeast, or E. coli cells, and the antibody is recovered from the cells (from the supernatant or after cells lysis).
[0408] Recombinant production of antibodies is well-known in the state of the art and described, for example, in the review articles of Makrides, S. C., Protein Expr. Purif. 17 (1999) 183-202; Geisse, S., et al., Protein Expr. Purif. 8 (1996) 271-282; Kaufman, R. J., Mol. Biotechnol. 16 (2000) 151-161; Werner, R. G., Drug Res. 48 (1998) 870-880.
[0409] The antibodies may be present in whole cells, in a cell lysate, or in a partially purified, or substantially pure form. Purification is performed in order to eliminate other cellular components or other contaminants, e.g. other cellular nucleic acids or proteins, by standard techniques, including alkaline/SDS treatment, CsCl banding, column chromatography, agarose gel electrophoresis, and others well known in the art. See Ausubel, F., et al., ed. Current Protocols in Molecular Biology, Greene Publishing and Wiley Interscience, New York (1987).
[0410] Expression in NS0 cells is described by, e.g., Barnes, L. M., et al., Cytotechnology 32 (2000) 109-123; Barnes, L. M., et al., Biotech. Bioeng. 73 (2001) 261-270. Transient expression is described by, e.g., Durocher, Y., et al., Nucl. Acids. Res. 30 (2002) E9. Cloning of variable domains is described by Orlandi, R., et al., Proc. Natl. Acad. Sci. USA 86 (1989) 3833-3837; Carter, P., et al., Proc. Natl. Acad. Sci. USA 89 (1992) 4285-4289; Norderhaug, L., et al., J. Immunol. Methods 204 (1997) 77-87. A preferred transient expression system (HEK 293) is described by Schlaeger, E.-J. and Christensen, K., in Cytotechnology 30 (1999) 71-83, and by Schlaeger, E.-J., in J. Immunol. Methods 194 (1996) 191-199.
[0411] Nucleic acid molecules encoding amino acid sequence variants of anti-CSF-1R antibody are prepared by a variety of methods known in the art. These methods include, but are not limited to, isolation from a natural source (in the case of naturally occurring amino acid sequence variants) or preparation by oligonucleotide-mediated (or site-directed) mutagenesis, PCR mutagenesis, and cassette mutagenesis of an earlier prepared variant or a non-variant version of humanized anti-CSF-1R antibody.
[0412] The heavy and light chain variable domains according to the invention are combined with sequences of promoter, translation initiation, constant region. 3' untranslated region, polyadenylation, and transcription termination to form expression vector constructs. The heavy and light chain expression constructs can be combined into a single vector, co-transfected, serially transfected, or separately transfected into host cells which are then fused to form a single host cell expressing both chains.
[0413] In another aspect, the present invention provides a composition, e.g. a pharmaceutical composition, containing one or a combination of monoclonal antibodies, or the antigen-binding portion thereof, of the present invention, formulated together with a pharmaceutically acceptable carrier.
[0414] As used herein, "pharmaceutically acceptable carrier" includes any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption/resorption delaying agents, and the like that are physiologically compatible. Preferably, the carrier is suitable for injection or infusion.
[0415] A composition of the present invention can be administered by a variety of methods known in the art. As will be appreciated by the skilled artisan, the route and/or mode of administration will vary depending upon the desired results.
[0416] Pharmaceutically acceptable carriers include sterile aqueous solutions or dispersions and sterile powders for the preparation of sterile injectable solutions or dispersion. The use of such media and agents for pharmaceutically active substances is known in the art. In addition to water, the carrier can be, for example, an isotonic buffered saline solution.
[0417] Regardless of the route of administration selected, the compounds of the present invention, which may be used in a suitable hydrated form, and/or the pharmaceutical compositions of the present invention, are formulated into pharmaceutically acceptable dosage forms by conventional methods known to those of skill in the art.
[0418] Actual dosage levels of the active ingredients in the pharmaceutical compositions of the present invention may be varied so as to obtain an amount of the active ingredient which is effective to achieve the desired therapeutic response for a particular patient, composition, and mode of administration, without being toxic to the patient (effective amount). The selected dosage level will depend upon a variety of pharmacokinetic factors including the activity of the particular compositions of the present invention employed, or the ester, salt or amide thereof, the route of administration, the time of administration, the rate of excretion of the particular compound being employed, other drugs, compounds and/or materials used in combination with the particular compositions employed, the age, sex, weight, condition, general health and prior medical history of the patient being treated, and like factors well known in the medical arts.
[0419] The invention comprises the use of the antibodies according to the invention for the treatment of a patient suffering from cancer, especially from colon, lung or pancreas cancer.
[0420] The invention comprises also a method for the treatment of a patient suffering from such disease.
[0421] The invention further provides a method for the manufacture of a pharmaceutical composition comprising an effective amount of an antibody according to the invention together with a pharmaceutically acceptable carrier and the use of the antibody according to the invention for such a method.
[0422] The invention further provides the use of an antibody according to the invention in an effective amount for the manufacture of a pharmaceutical agent, preferably together with a pharmaceutically acceptable carrier, for the treatment of a patient suffering from cancer.
[0423] The invention also provides the use of an antibody according to the invention in an effective amount for the manufacture of a pharmaceutical agent, preferably together with a pharmaceutically acceptable carrier, for the treatment of a patient suffering from cancer.
EXAMPLES
[0424] The examples and figures are provided to aid the understanding of the present invention, the true scope of which is set forth in the appended claims. It is understood that modifications can be made in the procedures set forth without departing from the spirit of the invention.
Example 1
[0425] Generation of a Hybridoma Cell Line Producing Anti-CSF-1R Antibodies
[0426] Immunization Procedure of NMR1 Mice
[0427] NMR1 mice were immunized with an expression vector pDisplay.TM. (Invitrogen, USA) encoding the extracellular domain of huCSF-1R by utilizing electroporation. Every mouse was 4 times immunized with 100 g DNA. When serum titers of anti-huCSF-1R were found to be sufficient, mice were additionally boosted once with 50 .mu.g of a 1:1 mixture huCSF-1R ECD/huCSF-1R ECDhuFc chimera in 200 .mu.l PBS intravenously (i.v.) 4 and 3 days before fusion.
[0428] Antigen Specific ELISA
[0429] Anti-CSF-1R titers in sera of immunized mice were determined by antigen specific ELISA.
[0430] 0.3 .mu.g/ml huCSF-1R-huFc chimera (soluble extracellular domain) was captured on a streptavidin plate (MaxiSorb; MicroCoat, DE, Cat. No. 11974998/MC1099) with 0.1 mg/ml biotinylated anti Fc.gamma. (Jackson ImmunoResearch., Cat. No. 109-066-098) and horse radish peroxidase (HRP)-conjugated F(ab').sub.2 anti mouse IgG (GE Healthcare, UK, Cat. No. NA9310V) diluted 1/800 in PBS/0.05% Tween20/0.5% BSA was added. Sera from all taps were diluted 1/40 in PBS/0.05% Tween20/0.5% BSA and serially diluted up to 1/1638400. Diluted sera were added to the wells. Pre-tap serum was used as negative control. A dilution series of mouse anti-human CSF-1R Mab3291 (R&D Systems, UK) from 500 ng/ml to 0.25 ng/ml was used as positive control. All components were incubated together for 1.5 hours, Wells were washed 6 times with PBST (PBS/0.2% Tween20) and assays were developed with freshly prepared ABTSo solution (1 mg/ml) (ABTS: 2,2'-azino bis (3-ethylbenzthiazoline-6-sulfonic acid) for 10 minutes at RT. Absorbance was measured at 405 nm.
[0431] Hybridoma Generation
[0432] The mouse lymphocytes can be isolated and fused with a mouse myeloma cell line using PEG based standard protocols to generate hybridomas. The resulting hybridomas are then screened for the production of antigen-specific antibodies. For example, single cell suspensions of splenic derived lymphocytes from immunized mice are fused to Ag8 non-secreting mouse myeloma cells P3X63Ag8.653 (ATCC, CRL-1580) with 50% PEG. Cells are plated at approximately 10.sup.4 in flat bottom 96 well micro titer plate, followed by about two weeks incubation in selective medium. Individual wells are then screened by ELISA for human anti-CSF-1R monoclonal IgM and IgG antibodies. Once extensive hybridoma growth occurs, the antibody secreting hybridomas are replated, screened again, and if still positive for human IgG, anti-CSF-1R monoclonal antibodies, can be subcloned by FACS. The stable subclones are then cultured in vitro to produce antibody in tissue culture medium for characterization. Antibodies according to the invention could be selected using the determination of the binding of anti-CSF-1R antibodies to human CSF-1R fragment delD4 and to human CSF-1R Extracellular Domain (CSF-1R-ECD) as described in Example 4, as well as the determination of growth inhibition of NIH3T3 cells transfected with wildtype CSF-1R (ligand dependent signalling) or mutant CSF-1R L301S Y969F (ligand independent signalling) under treatment with anti-CSF-1R monoclonal antibodies as described in Example 5.
[0433] Culture of Hybridomas
[0434] Generated muMAb hybridomas were cultured in RPMI 1640 (PAN--Catalogue No. (Cat. No.) PO4-17500) supplemented with 2 mM L-glutamine (GIBCO--Cat. No. 35050-038), 1 mM Na-Pyruvat (GIBCO--Cat. No. 11360-039). 1.times.NEAA (GIBCO--Cat. No. 11140-035), 10% FCS (PAA--Cat. No. A15-649), 1.times. Pen Strep (Roche--Cat. No. 1074440), 1.times. Nutridoma CS (Roche--Cat. No. 1363743), 50 .mu.M Mercaptoethanol (GIBCO--Cat. No. 31350-010) and 50 U/ml IL 6 mouse (Roche--Cat. No. 1 444 581) at 37.degree. C. and 5% CO.sub.2. Some of the resulting mouse antibodies have been humanized (e.g. Mab 2F11) and been expressed recombinantly.
Example 2
[0435] Inhibition of CSF-1 Binding to CSF-1R (ELISA)
[0436] By setting-up this assay to first allow for anti-CSF-1R antibody binding to the CSF-1R-ECD followed by detection of ligand not bound to the receptor both-ligand displacing antibodies and dimerization inhibitor anti-CSF-1R antibodies--can be tested. The test was performed on 384 well microtiter plates (MicroCoat, DE, Cat. No. 464718) at RT. After each incubation step plates were washed 3 times with PBST.
[0437] At the beginning, plates were coated with 0.5 mg/ml goat F(ab')2 biotinylated anti Fc.gamma. (Jackson ImmunoResearch., Cat. No. 109-006-170) for 1 hour (h).
[0438] Thereafter the wells were blocked with PBS supplemented with 0.2% Tween.RTM.-20 and 2% BSA (Roche Diagnostics GmbH, DE) for 0.5 h. 75 ng/ml of huCSF-1R-huFc chimera (which forms the dimeric soluble extracellular domain of huCSF-1R) was immobilized to plate for 1 h. Then dilutions of purified antibodies in PBS/0.05% Tween20/0.5% BSA were incubated for 1 h. After adding a mixture of 3 ng/ml CSF-1 (Biomol, DE. Cat. No. 60530), 50 ng/ml biotinylated anti CSF-1 clone BAF216 (R&D Systems, UK) and 1:5000 diluted streptavidin HRP (Roche Diagnostics GmbH, DE, Cat. No. 11089153001) for 1 h the plates were washed 6 times with PBST. Anti CSF-1R SC 2-4A5 (Santa Cruz Biotechnology, US), which inhibits the ligand-receptor interaction, was used as positive control. Plates were developed with freshly prepared BM Blue.RTM. POD substrate solution (BM Blue.RTM.: 3,3'-5,5'-Tetramethylbenzidine, Roche Diagnostics GmbH, DE, Cat. No. 11484281001) for 30 minutes at RT. Absorbance was measured at 370 nm. A decrease of absorbance is found, if the anti-CSF-1R antibody causes a release of CSF-1 from the dimeric complex. All anti-CSF-1R antibodies showed significant inhibition of the CSF-1 interaction with CSF-1R (see Table 1). Anti CSF-1R SC 2-4A5 (Santa Cruz Biotechnology, US see also Sherr, C. J. et al., Blood 73 (1989) 1786-1793), which inhibits the ligand-receptor interaction, was used as reference control.
TABLE-US-00001 TABLE 1 Calculated IC50 values for the inhibition of the CSF-1/CSF-1R interaction IC50 CSF-1/CSF-1R CSF-1R Mab Inhibition [ng/ml] Mab 2F11 19.3 Mab 2E10 20.6 Mab 2H7 18.2 Mab 1G10 11.8 SC-2-4A5 35.2
Example 3
[0439] Inhibition of CSF-1-Induced CSF-1R Phosphorylation in NIH3T3-CSF-1R Recombinant Cells
[0440] 4.5.times.10.sup.3 NIH 3T3 cells, retrovirally infected with an expression vector for full-length CSF-1R, were cultured in DMEM (PAA Cat. No. E15-011), 2 mM L-glutamine (Sigma, Cat. No. G7513, 2 mM Sodium pyruvate, 1.times. nonessential aminoacids, 10% FKS (PAA, Cat. No. A15-649) and 100 g/ml PenStrep (Sigma, Cat. No. P4333 [10 mg/ml]) until they reached confluency. Thereafter cells were washed with serum-free DMEM media (PAA Cat. No. E15-011) supplemented with sodium selenite [5 ng/ml] (Sigma, Cat. No. S9133), transferrin [10 .mu.g/ml] (Sigma, Cat. No. T8158), BSA [400 g/ml] (Roche Diagnostics GmbH, Cat. No. 10735078), 4 mM L-glutamine (Sigma, Cat. No. G7513), 2 mM sodium pyruvate (Gibco, Cat. No. 11360), 1.times. nonessential aminoacids (Gibco, Cat: 11140-035), 2-mercaptoethanol [0.05 mM] (Merck, Cat. No. M7522), 100 .mu.g/ml and PenStrep (Sigma, Cat. No. P4333) and incubated in 30 .mu.l of the same medium for 16 hours to allow for receptor up-regulation. 10 .mu.l of diluted anti-CSR-1R antibodies were added to the cells for 1.5 h. Then cells were stimulated with 10 .mu.l of 100 ng/ml huM-CSF-1 (Biomol Cat. No. 60530) for 5 min. After the incubation, supernatant was removed, cells were washed twice with 80 .mu.l of ice-cold PBS and 50 .mu.l of freshly prepared ice-cold lysis buffer (150 mM NaCl/20 mM Tris pH 7.5/1 mM EDTA/1 mM EGTA/1% Triton X-100/1 protease inhibitor tablet (Roche Diagnostics GmbH Cat. No. 1 836 170) per 10 ml buffer, 10 .mu.l/ml phosphatase inhibitor cocktail 1 (Sigma Cat. No. P-2850, 100.times. Stock)/10 .mu.l/ml protease inhibitor 1 (Sigma Cat. No. P-5726, 100.times. Stock)/10 .mu.l/ml 1 M NaF) was added. After 30 minutes on ice the plates were shaken vigourously on a plateshaker for 3 minutes and then centrifuged 10 minutes at 2200 rpm (Heraeus Megafuge 10).
[0441] The presence of phosphorylated and total CSF-1 receptor in the cell lysate was analyzed with Elisa. For detection of the phosphorylated receptor the kit from R&D Systems (Cat. No. DYC3268-2) was used according to the instructions of the supplier. For detection of total CSF-1R 10 .mu.l of the lysate was immobilized on plate by use of the capture antibody contained in the kit. Thereafter 1:750 diluted biotinylated anti CSF-1R antibody BAF329 (R&D Systems) and 1:1000 diluted streptavidin-HRP conjugate was added. After 60 minutes plates were developed with freshly prepared ABTS.RTM. solution and the absorbance was detected. Data were calculated as % of positive control without antibody and the ratio value phospho/total receptor expressed. The negative control was defined without addition of M-CSF-1. Anti CSF-1R SC 2-4A5 (Santa Cruz Biotechnology, US, see also Sherr, C. J. et al., Blood 73 (1989) 1786-1793), which inhibits the ligand-receptor interaction, was used as reference control.
TABLE-US-00002 TABLE 2 Calculated IC50 values for the inhibition of CSF-1 receptor phosphorylation. IC50 CSF-1R Phosphorylation CSF-1R Mab [ng/ml] Mab 2F11 219.4 Mab 2E10 752.0 Mab 2H7 703.4 Mab 1G10 56.6 SC-2-4A5 1006.6
Example 4
[0442] Determination of the Binding of Anti-CSF-1R Antibodies to Human CSF-1R Fragment delD4 and to Human CSF-1R Extracellular Domain (CSF-1R-ECD)
[0443] Preparation of Human CSF-1R Extracellular Domain (CSF-1R-ECD) (Comprising the Extracellular Subdomains D1-D5, hCSF-1R-ECD) of SEQ ID NO: 64:
[0444] pCMV-preS-Fc-hCSF-1R-ECD (7836 bp) encodes the complete ECD of human CSF-1R (SEQ ID NO: 64) C-terminally fused to a PreScission protease cleavage site, followed by aa100-330 of human IgG1 and a 6.times.His-Tag, under the control of CMV promoter. The natural signal peptide has been varied by insertion of amino acids G and S after the first M, in order to create a BamHI restriction site.
[0445] Preparation of Human CSF-1R Fragment delD4 (Comprising the Extracellular Subdomains D1-D3 and D5, hCSF-1R-delD4) of SEQ ID NO: 65:
[0446] hCSF1R-delD4-V1-PreSc-hFc-His was cloned from pCMV-preS-Fc-hCSF-1R-ECD by means of the Stratagene QuikChange XL site-directed mutagenesis protocol, using delD4--for with sequence CACCTCCATGTTCTTCCGGTACCCCCCAGAGGTAAG (SEQ ID NO: 68) as the forward primer and delD4-rev with the reverse complement sequence as the reverse primer. A protocol variation published in BioTechniques 26 (1999) 680 was used to extend both primers in separate reactions in three cycles preceeding the regular Stratagene protocol:
[0447] Two separate 50 .mu.l reaction mixtures were set up according to the manufacturer's manual, each containing 10 ng plasmid pCMV-preS-Fc-hCSF1R-ECD as the template and 10 .mu.M of one of the primers delD4--for or delD4-rev, and 0.5 .mu.l Pfu DNA polymerase as provided with the kit. Three PCR cycles 95.degree. C. 30 sec/55.degree. C. 60 sec/68.degree. C. 8 min were run, then 25 .mu.l each of both reaction mixtures were combined in a new tube and 0.5 .mu.l fresh Pfu DNA polymerase were added. The regular PCR protocol with 18 temperature cycles as specified by Stratagene in the kit manual was carried out, followed by 2 hrs final digestion with the Dpn restriction enzyme provided with the kit. Clones bearing the deletion were detected by digestion with Cel II and Not I and verified by sequencing.
[0448] Protein was prepared by transient transfection in the Hek293 FreeStyle suspension cell system (Invitrogen) according to the manufacturer's specifications. After 1 week 500 ml supernatant was filtered and loaded onto a 1 ml HiTrap MabSelect Xtra (GE healthcare) protein A column (0.2 ml/min). The column was washed first with PBS, then with 50 mM Tris/150 mM NaCl/1 mM EDTA/pH 7.3. 75 .mu.l PreScission Protease (GE #27-0843-01) diluted in 375 .mu.l of the same buffer were loaded onto the column and the closed column was incubated over night at 4.degree. C. with rolling. The column was mounted on top of a 1 ml GSTrap FF column (GE helthcare) and the desired protein was eluted (0.2 ml/min, 0.2 ml fractions). Pooled fractions were concentrated from 1.8 ml to 0.4 ml by centrifugal ultrafiltration via a 3 k Nanosep and chromatographed over an S200 HR SEC in PBS (0.5 ml/min).
[0449] Human CSF-1R fragment delD4 was obtained in two fractions as a dimeric molecule (pool1, V=1.5 ml; c=0.30 mg/ml; apparent mass on SDS page 83 kDa, reduced 62 kDa) and as the monomer (pool 2, V=1.4 ml; c=0.25 mg/ml apparent mass on SDS page 62 kDa). The dimeric form was used for all experiments.
[0450] Determination of the Binding of Anti-CSF-1R Antibodies to Human CSF-1R Fragment delD4 and to Human CSF-1R Extracellular Domain (CSF-1R-ECD) (Binding Signals as Response Units (RU):
[0451] Instrument: Biacore T00 (GE Healthcare) [0452] Software: T100 Control, Version 2.0.1 [0453] T100 Evaluation, Version 2.0.2
[0454] Assayformat Chip: CM5
[0455] Temperature: 25.degree. C.
[0456] CSF-1R fragments were immobilized via amine coupling. To compare the binding of different anti-CSF-1R antibodies according to the invention one concentration of the test antibody was injected. Anti CSF-1R Mab3291 (R&D-Systems) and SC 2-4A5 (Santa Cruz Biotechnology, US-- see also Sherr, C. J. et al., Blood 73 (1989) 1786-1793), was used as reference control, anti-CCR5 m<CCR5>Pz03.1C5 (deposited as DSM ACC 2683 on 18 Aug. 2004 at DSMZ) as negative control, all under the same conditions as the anti-CSF-1R antibodies according to the invention.
[0457] Amine Coupling of CSF-1R Fragments
[0458] Standard amine coupling according to the manufacturer's instructions: running buffer: PBS-T (Roche: 11 666 789+0.05% Tween20: 11 332 465), activation by mixture of EDC/NHS, injection of human CSF-1R fragment delD4 (comprising the extracellular subdomains D1-D3 and D5) (SEQ ID NO: 65) and human CSF-1R Extracellular Domain (CSF-1R-ECD) (comprising the extracellular subdomains D1-D5) (SEQ ID NO: 64) for 600 seconds at flow rate 10 .mu.l/min; diluted in coupling buffer NaAc, pH 5.0, c=10 .mu.g/mL; finally remaining activated carboxyl groups were blocked by injection of 1 M Ethanolamin.
[0459] Binding of <CSF-1R>Mab 2F11, Mab 2E10, Mab 3291 and Sc2-4A5 and Other Anti-CSF-1R Antibodies to Human CSF-1R Fragment delD4 and Human CSF-1R Extracelular Domain (CSF-1R-ECD) at 25.degree. C.
[0460] Running buffer: PBS-T (Roche: 11 666 789+0.05% Tween20: 11 332 465)
[0461] Analyte Sample:
[0462] Binding was measured at a flow rate of 30 l.mu.L/min by one injection of the analyte with concentration c=10 nM. (for Mab 1G10, Mab 2H7 and humanized hMab 2F11-e7 in second experiment) Each injection was 700 seconds long, followed by a dissociation phase of 180 seconds. Final regeneration was performed after each cycle using 50 mM NaOH, contact time 60 seconds, flow rate 30 .mu.L/min.
[0463] Signals were measured by a report point 10 seconds after end of injection. Reference signals (signals from a blank reference flow cell (treated with EDC/NHS and ethanolamine, only) were subtracted to give the binding signals (as RU). If binding signals of nonbinding antibodies were slightly below 0 (Mab 2F11=-3; Mab 2E10=-2; Mab 1G10=-6, Mab 2H7=-9; and humanized hMab 2F11-e7=-7) the values were set as 0.
TABLE-US-00003 TABLE 3a Binding of <CSF-1R> MAbs to human CSF-1R fragment delD4 and CSF-1R-ECD and ratio at 25.degree. C., measured by SPR Binding Ratio of binding of Binding to to CSF- anti-CSF1R antibodies delD4 1R-ECD to CSF1R fragment delD4/ [RU] [RU] to CSF-1R-ECD Mab 3291 1015 627 1015/627 = 1.61 sc2-4A5 374 249 374/249 = 1.50 Mab 2F11 0 176 0/176 = 0 hMab 2F11-e7 0 237 0/237 = 0 Mab 2E10 0 120 0/120 = 0 Mab 1G10 0 2708 0/2708 = 0 Mab 2H7 0 147 0/147 = 0 m<CCR5>Pz03.1C5 2 5 --
[0464] Mab 2F11 and Mab 2E10 showed binding to the human CSF-1R Extracellular Domain (CSF-1R-ECD) (see FIG. 2b); however no binding was detected to CSF-1R fragment delD4. (see FIG. 2a).
[0465] Sc2-4A5 and MAB3291 showed binding to CSF-1R-ECD and to del D4 (see FIGS. 2b and 2a).
[0466] Thus the ratio of binding of anti-CSF1R antibodies Mab 2F11 and Mab 2E10 to CSF1R fragment delD4/to CSF-1R-ECD was clearly below 1:50 (=0.02), while the binding ratio of MAB3291 and Sc2-4A5 were 1.61 and 1.50, respectively and were highly above 1:50 (=0.02). Negative control antibody m<CCR5>Pz03.1C5 did not show any binding (as expected).
[0467] Mab 1G10, Mab 2H7 and humanized hMab 2F11-e7 showed binding to the human CSF-1R Extracellular Domain (CSF-1R-ECD) (see FIG. 2d); however no binding was detected to CSF-1R fragment delD4. (see FIG. 2c). Thus the ratio of binding of anti-CSF1R antibodies Mab 1G10, Mab 2H7 and humanized hMab 2F11-e7 to CSF1R fragment delD4/to CSF-1R-ECD was clearly below 1:50 (=0.02).
[0468] In a further experiment anti-CSF-1R antibodies 1.2.SM (ligand displacing CSF-1R antibody described in WO2009026303), CXIIG6 (ligand displacing CSF-1R antibody described in WO 2009/112245), the goat polyclonal anti-CSF-1R antibody ab10676 (abcam) were investigated. Anti-CSF-1R antibody Mab3291 (R&D-Systems) was used as reference control. Anti-CCR5 m<CCR5>Pz03.1C5 (deposited as DSM ACC 2683 on 18 Aug. 2004 at DSMZ) was used as negative control.
TABLE-US-00004 TABLE 3b Binding of <CSF-1R> MAbs to human CSF-1R fragment delD4 and CSF-1R-ECD and ratio at 25.degree. C., measured by SPR Binding Ratio of binding of Binding to to CSF- anti-CSF1R antibodies delD4 1R-ECD to CSF1R fragment delD4/ [RU] [RU] to CSF-1R-ECD MAB3291 1790 1222 1790/1222 = 1.47 1.2.SM 469 704 469/704 = 0.67 CXIIG6 1983 1356 1983/1356 = 1.46 ab10676 787 547 787/547 = 1.44 m<CCR5>Pz03.1C5 0 0 --
[0469] 1.2.SM, CXIIG6, ab10676 and MAB3291 showed binding to CSF-1R-ECD and to del D4 (see FIGS. 2f and 2e).
[0470] The binding ratio of 1.2.SM, CXIIG6, ab10676 and MAB3291 was highly above 1:50 (=0.02). Negative control antibody m<CCR5>Pz03.1C5 did not show any binding (as expected).
Example 5
[0471] Growth Inhibition of NIH3T3-CSF-1R Recombinant Cells in 3D Culture Under Treatment with Anti-CSF-1R Monoclonal Antibodies (CellTiterGlo-Assay)
[0472] NIH 3T3 cells, retrovirally infected with either an expression vector for full-length wildtype CSF-1R (SEQ ID NO: 62) or mutant CSF-1R L301S Y969F (SEQ ID NO: 63), were cultured in DMEM high glucose media (PAA, Pasching, Austria) supplemented with 2 mM L-glutamine, 2 mM sodium pyruvate and non-essential amino acids and 10% fetal bovine serum (Sigma, Taufkirchen, Germany) on poly-HEMA (poly(2-hydroxyethylmethacrylatc)) (Polysciences, Warrington, Pa., USA)) coated dishes to prevent adherence to the plastic surface. Cells are seeded in medium replacing serum with 5 ng/ml sodium selenite, 10 mg/nl transferrin, 400 g/ml BSA and 0.05 mM 2-mercaptoethanol. When treated with 100 ng/ml huCSF-1 (Biomol, Hamburg, Germany) wtCSF-1R (expressing cells form dense spheroids that grow three dimensionally, a property that is called anchorage independence. These spheroids resemble closely the three dimensional architecture and organization of solid tumors in situ. Mutant CSF-1R recombinant cells are able to form spheroids independent of the CSF-1 ligand. Spheroid cultures were incubated for 3 days in the presence of different concentrations of antibody in order to determine an IC50 (concentration with 50 percent inhibition of cell viability). The CellTiterGlo assay was used to detect cell viability by measuring the ATP-content of the cells.
TABLE-US-00005 TABLE 5a wtCSF-1R Mutant CSF-1R CSF-1R Mab IC.sub.50 [.mu.g/ml] IC.sub.50 [.mu.g/ml] Mab 2F11 1.1 8.0 Mab 2E10 0.49 4.9 Mab 2H7 0.31 5.3 Mab 1G10 0.29 14.2 SC 2-4A5 10.0 10.0
[0473] Reference control Mab R&D-Systems 3291 did not show inhibition of mutant CSF-1R recombinant cell proliferation.
[0474] In a further experiment the anti-CSF-1R antibody according to the invention hMab 2F11-e7 and the anti-CSF-1R antibodies 1.2.SM (ligand displacing CSF-1R antibody described in WO2009026303), CXIIG6 (ligand displacing CSF-1R antibody described in WO 2009/112245), the goat polyclonal anti-CSF-1R antibody ab10676 (abcam), and SC 2-4A5 (Santa Cruz Biotechnology, US-- see also Sherr, C. J. et al., Blood 73 (1989) 1786-1793) were investigated.
[0475] Spheroid cultures were incubated for 3 days in the presence of different concentrations of antibody in order to determine an IC30 (concentration with 30 percent inhibition of cell viability). Maximum concentration was 20 .mu.g/ml The CellTiterGlo assay was used to detect cell viability by measuring the ATP-content of the cells.
TABLE-US-00006 TABLE 5b wtCSF-1R Mutant CSF-1R CSF-1R Mab IC.sub.30 [.mu.g/ml] IC.sub.30 [.mu.g/ml] hMab 2F11-e7 4.91 0.54 1.2.SM 1.19 >20 .mu.g/ml (-19% inhibition at 20 .mu.g/ml = 19% stimulation) CXIIG6 >20 .mu.g/ml (21% >20 .mu.g/ml (-36% inhibition at 20 .mu.g/ml) inhibition at 20 .mu.g/ml = 36% stimulation) ab10676 14.15 >20 .mu.g/ml (0% inhibition at 20 .mu.g/ml) SC 2-4A5 16.62 2.56
Example 6
[0476] Growth Inhibition of BeWo Tumor Cells in 3D Culture Under Treatment with Anti-CSF-1R Monoclonal Antibodies (CellTiterGlo-Assay)
[0477] BeWo choriocarcinoma cells (ATCC CCL-98) were cultured in F12K media (Sigma, Steinheim, Germany) supplemented with 10% FBS (Sigma) and 2 mM L-glutamine. 5.times.10.sup.4 cells/well were seeded in 96-well poly-HEMA (poly(2-hydroxyethylmethacrylate)) coated plates containing F12K medium supplemented with 0.5% FBS and 5% BSA. Concomitantly, 200 ng/ml huCSF-1 and 10 .mu.g/ml of different anti-CSF-1R monoclonal antibodies were added and incubated for 6 days. The CellTiterGlo assay was used to detect cell viability by measuring the ATP-content of the cells in relative light units (RLU). When BeWo spheroid cultures were treated with different anti-CSF-1R antibodies (10 .mu.g/ml) inhibition of CSF-1 induced growth was observed. To calculate antibody-mediated inhibition the mean RLU value of unstimulated BeWo cells was subtracted from all samples. Mean RLU value of CSF-1 stimulated cells was set arbitrarily to 100%. Mean RLU values of cells stimulated with CSF-1 and treated with anti-CSF-1R antibodies were calculated in % of CSF-1 stimulated RLUs. The Table 6 shows the calculated data of growth inhibition of BeWo tumor cells in 3D culture under treatment with anti-CSF-1R monoclonal antibodies; FIGS. 1a and b depicts normalized mean RLU values.
TABLE-US-00007 TABLE 6 % inhibition 10 .mu.g/ml CSF-1R Mab antibody concentration CSF-1 only 0 Mab 2F11 70 Mab 2E10 102 Mab 2H7 103 Mab 1G10 99 SC 2-4A5 39
Example 7
[0478] Inhibition of Human Macrophage Differentiation Under Treatment with Anti-CSF-1R Monoclonal Antibodies (CellTiterGlo-Assay)
[0479] Human monocytes were isolated from peripheral blood using the RosettcSep.TM. Human Monocyte Enrichment Cocktail (StemCell Tech. --Cat. No. 15028). Enriched monocyte populations were seeded into 96 well microtiterplates (2.5.times.10.sup.4 cells/well) in 100 .mu.l RPMI 1640 (Gibco--Cat. No. 31870) supplemented with 10% FCS (GIBCO--Cat. No. 011-090014M), 4 mM L-glutamine (GIBCO--Cat. No. 25030) and 1.times. PenStrep (Roche Cat. No. 1 074 440) at 37.degree. C. and 5% CO.sub.2 in a humidified atmosphere. When 150 ng/ml huCSF-1 was added to the medium, a clear differentiation into adherent macrophages could be observed. This differentiation could be inhibited by addition of anti-CSF-1R antibodies. Furthermore, the monocyte survival is affected and could be analyzed by CellTiterGlo (CTG) analysis. From the concentration dependent inhibition of the survival of monocytes by antibody treatment, an IC.sub.50 was calculated (see Table 7).
TABLE-US-00008 TABLE 7 CSF-1R Mab IC.sub.50 [.mu.g/ml] Mab 2F11 0.08 Mab 2E10 0.06 Mab 2H7 0.03 Mab 1G10 0.06 SC 2-4A5 0.36
[0480] In a separate test series humanized versions of Mab 2 F11, e.g. hMab 2F11-c11, hMab 2F11-d8, hMab 2F11-e7, hMab 2F11-f12, showed TC50 values of 0.07 .mu.g/ml (hMab 2F11-c11), 0.07 .mu.g/ml (hMab 2F11-d8), 0.04 g g/ml (hMab 2F11-e7) and 0.09 .mu.g/ml (hMab 2F11-f12).
Example 8
[0481] Inhibition of Cynomolgous Macrophage Differentiation Under Treatment with Anti-CSF-1R Monoclonal Antibodies (CellTiterGlo-Assay)
[0482] Cynomolgous monocytes were isolated from peripheral blood using the CD14 MicroBeads non-human primate kit (Miltenyi Biotec--Cat. No. 130-091-097) according to the manufacturers description. Enriched monocyte populations were seeded into 96 well microtiterplates (1-3.times.10.sup.4 cells/well) in 100 .mu.l RPMI 1640 (Gibco--Cat. No. 31870) supplemented with 10% FCS (GIBCO--Cat. No. 011-090014M), 4 mM L-glutamine (GIBCO--Cat. No. 25030) and 1.times. PenStrep (Roche Cat. No. 1 074 440) at 37.degree. C. and 5% CO.sub.2 in a humidified atmosphere. When 150 ng/ml huCSF-1 was added to the medium, a clear differentiation into adherent macrophages could be observed. This differentiation could be inhibited by addition of anti-CSF-1R antibodies. Furthermore, the monocyte survival is affected and could be analyzed by CellTiterGlo (CTG) analysis. The viability was analyzed at a concentration of 5 .mu.g/ml antibody treatment (see Table 8).
TABLE-US-00009 TABLE 8 % inhibition (of survival) = CSF-1R Mab % survival (100% - % survival) Mab 2F11 4* 96 Mab 2E10 17** 83 Mab 2H7 8 92 Mab 1G10 2 98 SC 2-4A5 31 69 *mean of four experiments (3 expts. using the murine, 1 expt. using the chimeric mAb) **mean of two experiments using the murine mAb only
Example 9
[0483] Determination of the Binding Affinity of Anti-CSF-1R Antibodies to Human CSF-1R
[0484] Instrument: BIACORE.RTM. A100
[0485] Chip: CM5 (Biacore BR-1006-68)
[0486] Coupling: amine coupling
[0487] Buffer: PBS (Biacore BR-1006-72), pH 7.4, 35.degree. C.
[0488] For affinity measurements 36 .mu.g/ml anti mouse Fc.gamma. antibodies (from goat, Jackson Immuno Reasearch JIR115-005-071) have been coupled to the chip surface for capturing the antibodies against CSF-1R. Human CSF-1R Extracellular Domain (CSF-1R-ECD) (comprising the extracellular subdomains D1-D5) (SEQ ID NO: 64) (R&D-Systems 329-MR or subcloned pCMV-presS-HisAvitag-hCSF-1R-ECD) was added in various concentrations in solution. Association was measured by an CSF-1R-injection of 1.5 minutes at 35.degree. C.; dissociation was measured by washing the chip surface with buffer for 10 minutes at 35.degree. C. For calculation of kinetic parameters the Langmuir 1:1 model was used.
TABLE-US-00010 TABLE 9 Affinity data measured by SPR CSF-1R Mab K.sub.D (nM) k.sub.a (1/Ms) k.sub.d (1/s) t.sub.1/2 (min) Mab 2F11 0.29 1.77E.sup.+05 5.18E.sup.-05 223 Mab 2E10 0.2 1.52E.sup.+05 2.97E.sup.-05 389 Mab 2H7 0.21 1.47E.sup.+05 3.12E.sup.-05 370 Mab 1G10 0.36 1.75E.sup.+05 6.28E.sup.-05 184
[0489] In a separate biacore binding assay using the CSF-1R ECD (data not shown) some competition of the antibodies Mab 2F11 and Mab 2E10 with the antibody Ab SC-2-4A5 was shown. However Mab 2F11/Mab 2E10 do not bind to the human CSF-1R fragment delD4, whereas Ab SC-2-4A5 binds to this delD4 fragment (see Example 4 and FIG. 2a). Thus the binding region of Mab 2F11/Mab 2E10 is clearly distinct from the binding region of Ab SC-2-4A5, but probably located in a vicinity area. In such competition assay both antibodies Mab 2F11 and Mab 2E10 did not compete with Mab3291 from R&D-Systems (data not shown).
Example 10
[0490] Determination of the Binding of Anti-CSF-1R Antibodies to Human CSF-1R Fragment D1-D3
[0491] Instrument: Biacore T100 (GE Healthcare) [0492] Software: T100 Control, Version 1.1.11 [0493] B3000 Evaluation, Version 4.01 [0494] Scrubber, Version 2.0a
[0495] Assayformat Chip: CM5-Chip
[0496] Antibodies against CSF-1R were captured via amine coupled capture molecules. Using the single cycle kinetics five increasing concentrations of human CSF-1R fragment D1-D3 (SEQ ID NO: 66) were injected. Human CSF-1R fragment D1-D3 was subcloned into pCMV-presS-HisAvitag expression vector.
[0497] Anti CSF-1R SC 2-4A5 (Santa Cruz Biotechnology, US; Sherr. C. J. et al., Blood 73 (1989) 1786-1793) which inhibits the ligand-receptor interaction, and Mab 3291 (R&D-Systems) were used as reference controls.
[0498] Capture molecules: Anti mouse Fc.gamma. antibodies (from goat, Jackson Immuno Reasearch JIR115-005-071) for antibodies according to the invention and the R&D-Systems control Mab 3291 and Anti rat Fc.gamma. antibodies (from goat, Jackson Immuno Reasearch JIR112-005-071) for the reference control anti CSF-1R SC 2-4A5.
[0499] Amine Coupling of Capture Molecules
[0500] Standard amine coupling according to the manufacturer's instructions: running buffer: HBS--N buffer, activation by mixture of EDC/NHS, aim for ligand density of 2000 RU; the capture-Abs were diluted in coupling buffer NaAc, pH 4.5, c=10 g/mL; finally remaining activated carboxyl groups were blocked by injection of 1 M Ethanolamin.
[0501] Kinetic Characterization of Human CSF-1R Fragments D1-D3 Binding to MAbs <CSF-1R> at 37.degree. C.
[0502] Running buffer: PBS (Biacore BR-1006-72)
[0503] Capturing of Mabs <CSF-1R> on flow cells 2 to 4: Flow 20 .mu.L/min, contact time 90 seconds, c(Abs<CSF-1R>)=50 nM, diluted with running buffer+1 mg/mL BSA;
[0504] Analyte Sample:
[0505] Single Cycle Kinetics was measured at a flow rate of 30 .mu.L/min by five consecutive injections of the analyte with concentrations, c=7.8, 31.25, 125 500 and 2000 nM, without regeneration. Each injection was 30 seconds long and followed by a dissociation phase of 120 Seconds for the first four injections, and finally 1200 seconds for the highest concentration (=last injection).
[0506] Final regeneration was performed after each cycle using 10 mM Glycin pH 1.5 (Biacore BR-1003-54), contact time 60 seconds, flow rate 30 .mu.L/min.
[0507] Kinetic parameters were calculated by using the usual double referencing (control reference: binding of analyte to capture molecule; Flow Cell: subdomain CSF-1R concentration "0" as Blank) and calculation with model `titration kinetics 1:1 binding with draft`.
TABLE-US-00011 TABLE 10 Affinity data for binding of human CSF-1R fragment D1-D3 measured by SPR Sub CSF-1R Mab domain K.sub.D (nM) k.sub.a (1/Ms) k.sub.d (1/s) t.sub.1/2 (min) Mab 2F11 D1-D3 no binding Mab 2E10 D1-D3 no binding Mab 2H7 D1-D3 not determined Mab 1G10 D1-D3 no binding SC-2-4A5 D1-D3 no binding R&D-Systems D1-D3 5.4 2.2E.sup.+5 1.2E.sup.-3 9.6 3291
[0508] The antibodies Mab 2F11, Mab 2E10 and Mab 1G10 showed no binding to human CSF-1R fragment D1-D3
[0509] Also reference control-Ab SC-2-4A5 did not bind to human CSF-1R fragment D1-D3.
[0510] The reference control Mab R&D-Systems 3291 showed binding to the human CSF-1R fragment D1-D3.
Example 11
[0511] CSF-1 level increase during CSF-1R inhibition in Cynomolgus monkey Serum CSF-1 levels provide a pharmacodynamic marker of CSF-1R neutralizing activity of anti-human CSF-1R dimerization inhibitor hMab 2F11-e7. One male and one female cynomolgus monkey per dosage group (1 and 10 mg/kg) were intravenously administered anti-CSF1R antibody hMab 2F11-e7. Blood samples for analysis of CSF-1 levels were collected 1 week before treatment (pre-dose), 2, 24, 48, 72, 96, 168 hours post-dose and weekly for two additional weeks. CSF-1 levels were determined using a commercially available ELISA kit (Quantikine.RTM. human M-CSF) according to the manufacturer's instructions (R&D Systems, UK). Monkey CSF-1 level were determined by comparison with CSF-1 standard curve samples provided in the kit.
[0512] Administration of hMab 2F11-e7 induced a dramatic increase in CSF-1 by .about.1000-fold, which depending on the dose administered lasted for 48 hr (1 mg/kg) or 15 days (10 mg/kg). Hence, a dimerization inhibitor for CSF-1R offers the advantage to not directly compete with the dramatically upregulated ligand for binding to the receptor in contrast to a ligand displacing antibody.
Example 12
[0513] In Vivo Efficacy--Tumor Growth Inhibition of Anti-CSF-1R Antibodies in Breast Cancer BT20 Xenograft Tumor Cells in SCID Beige Mice
[0514] The human breast cancer cell line BT-20 expresses human CSF-1R but lacks CSF-1 expression (Sapi, E. et al Cancer Res 59 (1999) 5578-5585). Since the mouse derived CSF-1 fails to activate human CSF-1R on the tumor cells recombinant human CSF-1 (Biomol, Hamburg, Germany) was supplemented via osmotic minipumps (ALZET, Cupertino, Calif.) providing a continuous CSF-1 infusion rate of 2 .mu.g/day (Martin, T. A., Carcinogenesis 24 (2003) 1317-1323).
[0515] To directly compare the efficacy of an antibody interfering with dimerization of CSF-1R with a ligand displacing CSF-1R antibody we tested the chimeric anti-CSF-1R Mab 2F1I (antibody interfering with dimerization of CSF-1R) and 1.2.SM (ligand displacing CSF-1R antibody described in WO2009026303) in the BT-20 xenograft model.
[0516] SCID beige mice (Charles River, Sulzfeld, Germany) were subcutaneously coinjected with 1.times.107 cells BT-20 cells (ATCC HTB-19) and 100l of Matrigel. Treatment of animals started at day of randomization at a mean tumor volume of 100 mm3. Mice are treated once weekly i.p. with the respective antibodies (see FIG. 4) in 20 mM Histidine, 140 mM NaCl pH 6.0 buffer. The tumor dimensions are measured by caliper beginning on the staging day and subsequently 2 times per week during the whole treatment period. Tumor volume is calculated according to NCI protocol (Tumor weight=1/2ab2, where "a" and "b" are the long and the short diameters of the tumor, respectively).
[0517] Tumor growth analysis is shown in FIG. 4. Inhibition of human CSF-1R on tumor cells with the chimeric anti-CSF-1R Mab 2F1I was statistically more efficacious in mediating tumor growth inhibition than anti-CSF-1R antibody 1.2.SM (CSF-1R antibody described in WO2009026303).
[0518] Although the foregoing invention has been described in some detail by way of illustration and example for purposes of clarity of understanding, the descriptions and examples should not be construed as limiting the scope of the invention. The disclosures of all patents, patent applications, scientific references, and Genbank Accession Nos. cited herein are expressly incorporated by reference in their entirety for all purposes as if each patent, patent application, scientific reference, and Genbank Accession No. were specifically and individually incorporated by reference.
Sequence CWU 1
1
8418PRTMus musculus 1Asp Gln Arg Leu Tyr Phe Asp Val1 5216PRTMus
musculus 2Val Ile Trp Thr Asp Gly Gly Thr Asn Tyr Asn Ser Pro Phe
Met Ser1 5 10 1535PRTMus musculus 3Thr Tyr Asp Ile Ser1 548PRTMus
musculus 4Gly Gln Ser Phe Ser Tyr Pro Thr1 557PRTMus musculus 5Gly
Ala Ser Asn Arg Tyr Thr1 5611PRTMus musculus 6Lys Ala Ser Glu Asp
Val Asn Thr Tyr Val Ser1 5 107116PRTMus musculus 7Gln Val Gln Leu
Lys Glu Ser Gly Pro Gly Leu Val Ala Pro Ser Gln1 5 10 15Ser Leu Ser
Ile Thr Cys Thr Val Ser Gly Phe Ser Leu Thr Thr Tyr 20 25 30Asp Ile
Ser Trp Ile Arg Gln Ser Pro Gly Lys Gly Leu Glu Trp Leu 35 40 45Gly
Val Ile Trp Thr Asp Gly Gly Thr Asn Tyr Asn Ser Pro Phe Met 50 55
60Ser Arg Leu Ser Ile Arg Lys Asp Asn Ser Lys Ser Gln Val Phe Leu65
70 75 80Lys Met Asn Arg Leu Gln Thr Asp Asp Thr Ala Ile Tyr Tyr Cys
Val 85 90 95Arg Asp Gln Arg Leu Tyr Phe Asp Val Trp Gly Ala Gly Thr
Thr Val 100 105 110Thr Val Ser Ser 1158106PRTMus musculus 8Asn Ile
Val Met Thr Gln Ser Pro Lys Ser Met Ser Met Ser Val Gly1 5 10 15Glu
Arg Val Thr Leu Asn Cys Lys Ala Ser Glu Asp Val Asn Thr Tyr 20 25
30Val Ser Trp Tyr Gln Gln Gln Pro Glu Gln Ser Pro Lys Leu Leu Ile
35 40 45Tyr Gly Ala Ser Asn Arg Tyr Thr Gly Val Pro Asp Arg Phe Thr
Gly 50 55 60Gly Gly Ser Thr Thr Asp Phe Thr Leu Thr Ile Ser Ser Val
Gln Ala65 70 75 80Glu Asp Leu Ala Asp Tyr Phe Cys Gly Gln Ser Phe
Ser Tyr Pro Thr 85 90 95Phe Gly Thr Gly Thr Lys Leu Glu Ile Lys 100
10597PRTMus musculus 9Asp Pro Arg Leu Tyr Phe Asp1 51016PRTMus
musculus 10Val Ile Trp Thr Gly Gly Gly Thr Asn Tyr Asn Ser Gly Phe
Met Ser1 5 10 15115PRTMus musculus 11Ser Phe Asp Ile Ser1
5128PRTMus musculus 12Gly Gln Thr Phe Ser Tyr Pro Thr1 5137PRTMus
musculus 13Gly Ala Ser Asn Arg Tyr Thr1 51411PRTMus musculus 14Lys
Ala Ser Glu Asp Val Val Thr Tyr Val Ser1 5 1015116PRTMus musculus
15Gln Val Gln Leu Lys Glu Ser Gly Pro Gly Leu Val Ala Pro Ser Lys1
5 10 15Ser Leu Ser Ile Thr Cys Thr Val Ser Gly Ser Ser Leu Asp Ser
Phe 20 25 30Asp Ile Ser Trp Ile Arg Gln Ser Pro Gly Lys Gly Leu Glu
Trp Leu 35 40 45Gly Val Ile Trp Thr Gly Gly Gly Thr Asn Tyr Asn Ser
Gly Phe Met 50 55 60Ser Arg Leu Arg Ile Thr Lys Asp Asn Ser Lys Ser
Gln Val Leu Leu65 70 75 80Lys Met Asn Ser Leu Gln Ser Asp Asp Thr
Ala Ile Tyr Tyr Cys Val 85 90 95Arg Asp Pro Arg Leu Tyr Phe Asp Val
Trp Gly Ala Gly Thr Thr Val 100 105 110Thr Val Ser Ser
11516106PRTMus musculus 16Asn Ile Val Met Thr Gln Ser Pro Lys Ser
Met Ser Met Ser Val Gly1 5 10 15Glu Arg Val Thr Leu Ser Cys Lys Ala
Ser Glu Asp Val Val Thr Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro
Asp Gln Ser Pro Lys Leu Leu Ile 35 40 45Tyr Gly Ala Ser Asn Arg Tyr
Thr Gly Val Pro Asp Arg Phe Thr Gly 50 55 60Ser Gly Ser Ala Thr Asp
Phe Thr Leu Thr Ile Ser Ser Val Gln Ala65 70 75 80Glu Asp Leu Ala
Asp Tyr Tyr Cys Gly Gln Thr Phe Ser Tyr Pro Thr 85 90 95Phe Gly Thr
Gly Thr Lys Leu Glu Ile Lys 100 105178PRTArtificialheavy chain
CDR3, hMab 2F11-c11 17Asp Gln Arg Leu Tyr Phe Asp Val1
51816PRTArtificialheavy chain CDR2, hMab 2F11-c11 18Val Ile Trp Thr
Asp Gly Gly Thr Asn Tyr Asn Ser Pro Phe Met Ser1 5 10
15195PRTArtificialheavy chain CDR1, hMab 2F11-c11 19Thr Tyr Asp Ile
Ser1 5208PRTArtificiallight chain CDR3, hMab 2F11-c11 20Gly Gln Ser
Phe Ser Tyr Pro Thr1 5217PRTArtificiallight chain CDR2, hMab
2F11-c11 21Gly Ala Ser Asn Arg Tyr Thr1 52211PRTArtificiallight
chain CDR1, hMab 2F11-c11 22Arg Ala Ser Glu Asp Val Asn Thr Tyr Val
Ser1 5 1023116PRTArtificialheavy chain variable domain, hMab
2F11-c11 23Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro
Gly Ser1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Phe Ser Leu
Thr Thr Tyr 20 25 30Asp Ile Ser Trp Ile Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45Gly Val Ile Trp Thr Asp Gly Gly Thr Asn Tyr
Asn Ser Pro Phe Met 50 55 60Ser Arg Val Thr Ile Thr Lys Asp Glu Ser
Thr Ser Thr Ala Tyr Met65 70 75 80Glu Leu Ser Ser Leu Arg Ser Glu
Asp Thr Ala Val Tyr Tyr Cys Val 85 90 95Arg Asp Gln Arg Leu Tyr Phe
Asp Val Trp Gly Gln Gly Thr Thr Val 100 105 110Thr Val Ser Ser
11524106PRTArtificiallight chain variable domain, hMab 2F11-c11
24Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu Asp Val Asn Thr
Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
Leu Ile 35 40 45Tyr Gly Ala Ser Asn Arg Tyr Thr Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gly Gln
Ser Phe Ser Tyr Pro Thr 85 90 95Phe Gly Gln Gly Thr Lys Leu Glu Ile
Lys 100 105258PRTArtificialheavy chain CDR3, hMab 2F11-d8 25Asp Gln
Arg Leu Tyr Phe Asp Val1 52616PRTArtificialheavy chain CDR2, hMab
2F11-d8 26Val Ile Trp Thr Asp Gly Gly Ala Asn Tyr Ala Gln Lys Phe
Gln Gly1 5 10 15275PRTArtificialheavy chain CDR1, hMab 2F11-d8
27Thr Tyr Asp Ile Ser1 5288PRTArtificiallight chain CDR3, hMab
2F11-d8 28Gly Gln Ser Phe Ser Tyr Pro Thr1 5297PRTArtificiallight
chain CDR2, hMab 2F11-d8 29Gly Ala Ser Asn Arg Tyr Thr1
53011PRTArtificiallight chain CDR1, hMab 2F11-d8 30Lys Ala Ser Glu
Asp Val Asn Thr Tyr Val Ser1 5 1031116PRTArtificialheavy chain
variable domain, hMab 2F11-d8 31Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser1 5 10 15Ser Val Lys Val Ser Cys Lys Ala
Ser Gly Phe Ser Leu Thr Thr Tyr 20 25 30Asp Ile Ser Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Val Ile Trp Thr Asp
Gly Gly Ala Asn Tyr Ala Gln Lys Phe Gln 50 55 60Gly Arg Val Thr Ile
Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr Met65 70 75 80Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys Ala 85 90 95Arg Asp
Gln Arg Leu Tyr Phe Asp Val Trp Gly Gln Gly Thr Thr Val 100 105
110Thr Val Ser Ser 11532106PRTArtificiallight chain variable
domain, hMab 2F11-d8 32Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu
Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Lys Ala Ser
Glu Asp Val Asn Thr Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro Gly
Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Ala Ser Asn Arg Tyr Thr
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr
Tyr Tyr Cys Gly Gln Ser Phe Ser Tyr Pro Thr 85 90 95Phe Gly Gln Gly
Thr Lys Leu Glu Ile Lys 100 105338PRTArtificialheavy chain CDR3,
hMab 2F11-e7 33Asp Gln Arg Leu Tyr Phe Asp Val1
53416PRTArtificialheavy chain CDR2, hMab 2F11-e7 34Val Ile Trp Thr
Asp Gly Gly Thr Asn Tyr Ala Gln Lys Leu Gln Gly1 5 10
15355PRTArtificialheavy chain CDR1, hMab 2F11-e7 35Ser Tyr Asp Ile
Ser1 5368PRTArtificiallight chain CDR3, hMab 2F11-e7 36Gln Gln Ser
Phe Ser Tyr Pro Thr1 5377PRTArtificiallight chain CDR2, hMab
2F11-e7 37Ala Ala Ser Asn Arg Tyr Thr1 53811PRTArtificiallight
chain CDR1, hMab 2F11-e7 38Arg Ala Ser Glu Asp Val Asn Thr Tyr Val
Ser1 5 1039116PRTArtificialheavy chain variable domain, hMab
2F11-e7 39Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro
Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe
Thr Ser Tyr 20 25 30Asp Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45Gly Val Ile Trp Thr Asp Gly Gly Thr Asn Tyr
Ala Gln Lys Leu Gln 50 55 60Gly Arg Val Thr Met Thr Thr Asp Thr Ser
Thr Ser Thr Ala Tyr Met65 70 75 80Glu Leu Arg Ser Leu Arg Ser Asp
Asp Thr Ala Val Tyr Tyr Cys Ala 85 90 95Arg Asp Gln Arg Leu Tyr Phe
Asp Val Trp Gly Gln Gly Thr Thr Val 100 105 110Thr Val Ser Ser
11540106PRTArtificiallight chain variable domain, hMab 2F11-e7
40Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu Asp Val Asn Thr
Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
Leu Ile 35 40 45Tyr Ala Ala Ser Asn Arg Tyr Thr Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln
Ser Phe Ser Tyr Pro Thr 85 90 95Phe Gly Gln Gly Thr Lys Leu Glu Ile
Lys 100 105418PRTArtificialheavy chain CDR3, hMab 2F11-f12 41Asp
Gln Arg Leu Tyr Phe Asp Val1 54216PRTArtificialheavy chain CDR2,
hMab 2F11-f12 42Val Ile Trp Thr Asp Gly Gly Thr Asn Tyr Asn Ser Pro
Phe Met Ser1 5 10 15435PRTArtificialheavy chain CDR1, hMab 2F11-f12
43Thr Tyr Asp Ile Ser1 5448PRTArtificiallight chain CDR3, hMab
2F11-f12 44Gly Gln Ser Phe Ser Tyr Pro Thr1 5457PRTArtificiallight
chain CDR2, hMab 2F11-f12 45Gly Ala Ser Ser Leu Gln Ser1
54611PRTArtificiallight chain CDR1, hMab 2F11-f12 46Arg Ala Ser Glu
Asp Val Asn Thr Tyr Val Ser1 5 1047116PRTArtificialheavy chain
variable domain, hMab 2F11-f12 47Gln Val Gln Leu Val Gln Ser Gly
Ala Glu Val Lys Lys Pro Gly Ser1 5 10 15Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Phe Ser Leu Thr Thr Tyr 20 25 30Asp Ile Ser Trp Val Arg
Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Val Ile Trp Thr
Asp Gly Gly Thr Asn Tyr Asn Ser Pro Phe Met 50 55 60Ser Arg Val Thr
Ile Thr Lys Asp Glu Ser Thr Ser Thr Ala Tyr Met65 70 75 80Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys Val 85 90 95Arg
Asp Gln Arg Leu Tyr Phe Asp Val Trp Gly Gln Gly Thr Thr Val 100 105
110Thr Val Ser Ser 11548106PRTArtificiallight chain variable
domain, hMab 2F11-f12 48Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu
Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser
Glu Asp Val Asn Thr Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro Gly
Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Gly Ala Ser Ser Leu Gln Ser
Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr
Tyr Tyr Cys Gly Gln Ser Phe Ser Tyr Pro Thr 85 90 95Phe Gly Gln Gly
Thr Lys Leu Glu Ile Lys 100 105498PRTArtificialheavy chain CDR3,
hMab 2F11-g1 49Asp Gln Arg Leu Tyr Phe Asp Val1
55016PRTArtificialheavy chain CDR2, hMab 2F11-g1 50Val Ile Trp Thr
Asp Gly Gly Thr Asn Tyr Asn Ser Pro Leu Lys Ser1 5 10
15515PRTArtificialheavy chain CDR1, hMab 2F11-g1 51Thr Tyr Asp Ile
Ser1 5528PRTArtificiallight chain CDR3, hMab 2F11-g1 52Gly Gln Ser
Phe Ser Tyr Pro Thr1 5537PRTArtificiallight chain CDR2, hMab
2F11-g1 53Gly Ala Ser Ser Arg Ala Thr1 55411PRTArtificiallight
chain CDR1, hMab 2F11-g1 54Arg Ala Ser Glu Asp Val Asn Thr Tyr Leu
Ala1 5 1055116PRTArtificialheavy chain variable domain, hMab
2F11-g1 55Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro
Ser Glu1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Phe Ser Leu
Thr Thr Tyr 20 25 30Asp Ile Ser Trp Ile Arg Gln Pro Pro Gly Lys Gly
Leu Glu Trp Ile 35 40 45Gly Val Ile Trp Thr Asp Gly Gly Thr Asn Tyr
Asn Ser Pro Leu Lys 50 55 60Ser Arg Val Thr Ile Ser Val Asp Thr Ser
Lys Asn Gln Phe Ser Leu65 70 75 80Lys Leu Ser Ser Val Thr Ala Ala
Asp Thr Ala Val Tyr Tyr Cys Ala 85 90 95Arg Asp Gln Arg Leu Tyr Phe
Asp Val Trp Gly Gln Gly Thr Thr Val 100 105 110Thr Val Ser Ser
11556106PRTArtificiallight chain variable domain, hMab 2F11-g1
56Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly1
5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Glu Asp Val Asn Thr
Tyr 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu
Leu Ile 35 40 45Tyr Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Arg Leu Glu Pro65 70 75 80Glu Asp Phe Ala Val Tyr Tyr Cys Gly Gln
Ser Phe Ser Tyr Pro Thr 85 90 95Phe Gly Gln Gly Thr Lys Leu Glu Ile
Lys 100 10557107PRTHomo sapiens 57Arg Thr Val Ala Ala Pro Ser Val
Phe Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr Ala
Ser Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala Lys
Val Gln Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser Gln
Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser Leu
Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75 80Lys His
Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser 85 90 95Pro
Val Thr Lys Ser Phe Asn Arg Gly Glu Cys 100 10558330PRTHomo sapiens
58Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys1
5 10 15Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val Lys Asp
Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu
Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly
Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu
Gly Thr Gln Thr65 70 75 80Tyr Ile Cys Asn Val Asn His Lys Pro Ser
Asn Thr Lys Val Asp Lys 85 90
95Lys Val Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys
100 105 110Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe
Pro Pro 115 120 125Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro
Glu Val Thr Cys 130 135 140Val Val Val Asp Val Ser His Glu Asp Pro
Glu Val Lys Phe Asn Trp145 150 155 160Tyr Val Asp Gly Val Glu Val
His Asn Ala Lys Thr Lys Pro Arg Glu 165 170 175Glu Gln Tyr Asn Ser
Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu 180 185 190His Gln Asp
Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn 195 200 205Lys
Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly 210 215
220Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp
Glu225 230 235 240Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
Lys Gly Phe Tyr 245 250 255Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Asn Gly Gln Pro Glu Asn 260 265 270Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp Ser Asp Gly Ser Phe Phe 275 280 285Leu Tyr Ser Lys Leu Thr
Val Asp Lys Ser Arg Trp Gln Gln Gly Asn 290 295 300Val Phe Ser Cys
Ser Val Met His Glu Ala Leu His Asn His Tyr Thr305 310 315 320Gln
Lys Ser Leu Ser Leu Ser Pro Gly Lys 325 33059330PRTArtificialhuman
heavy chain constant region derived from IgG1 mutated on L234A and
L235A 59Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser
Lys1 5 10 15Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val Lys
Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala
Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser
Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser
Leu Gly Thr Gln Thr65 70 75 80Tyr Ile Cys Asn Val Asn His Lys Pro
Ser Asn Thr Lys Val Asp Lys 85 90 95Lys Val Glu Pro Lys Ser Cys Asp
Lys Thr His Thr Cys Pro Pro Cys 100 105 110Pro Ala Pro Glu Ala Ala
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro 115 120 125Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys 130 135 140Val Val
Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp145 150 155
160Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu
165 170 175Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr
Val Leu 180 185 190His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn 195 200 205Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr
Ile Ser Lys Ala Lys Gly 210 215 220Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu Pro Pro Ser Arg Asp Glu225 230 235 240Leu Thr Lys Asn Gln
Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr 245 250 255Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn 260 265 270Asn
Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe 275 280
285Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn
290 295 300Val Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn His
Tyr Thr305 310 315 320Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 325
33060327PRTHomo sapiens 60Ala Ser Thr Lys Gly Pro Ser Val Phe Pro
Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu
Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser
Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala
Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr
Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr Thr Cys Asn
Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Arg Val Glu
Ser Lys Tyr Gly Pro Pro Cys Pro Ser Cys Pro Ala Pro 100 105 110Glu
Phe Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys 115 120
125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val
130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr
Val Asp145 150 155 160Gly Val Glu Val His Asn Ala Lys Thr Lys Pro
Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg Val Val Ser Val
Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn Gly Lys Glu Tyr
Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro Ser Ser Ile Glu
Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215 220Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys225 230 235
240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys Ser Arg Trp Gln
Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu Ser Leu Ser Leu
Gly Lys 32561327PRTArtificialhuman heavy chain constant region
derived from IgG4 mutated onS228P 61Ala Ser Thr Lys Gly Pro Ser Val
Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala
Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr
Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe
Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val
Val Thr Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr Thr
Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Arg
Val Glu Ser Lys Tyr Gly Pro Pro Cys Pro Pro Cys Pro Ala Pro 100 105
110Glu Phe Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys
115 120 125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val
Val Val 130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn
Trp Tyr Val Asp145 150 155 160Gly Val Glu Val His Asn Ala Lys Thr
Lys Pro Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg Val Val
Ser Val Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn Gly Lys
Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro Ser Ser
Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215 220Glu
Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys225 230
235 240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp 245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser
Phe Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys Ser Arg Trp
Gln Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met His Glu Ala
Leu His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu Ser Leu Ser
Leu Gly Lys 32562972PRTHomo sapiens 62Met Gly Pro Gly Val Leu Leu
Leu Leu Leu Val Ala Thr Ala Trp His1 5 10 15Gly Gln Gly Ile Pro Val
Ile Glu Pro Ser Val Pro Glu Leu Val Val 20 25 30Lys Pro Gly Ala Thr
Val Thr Leu Arg Cys Val Gly Asn Gly Ser Val 35 40 45Glu Trp Asp Gly
Pro Pro Ser Pro His Trp Thr Leu Tyr Ser Asp Gly 50 55 60Ser Ser Ser
Ile Leu Ser Thr Asn Asn Ala Thr Phe Gln Asn Thr Gly65 70 75 80Thr
Tyr Arg Cys Thr Glu Pro Gly Asp Pro Leu Gly Gly Ser Ala Ala 85 90
95Ile His Leu Tyr Val Lys Asp Pro Ala Arg Pro Trp Asn Val Leu Ala
100 105 110Gln Glu Val Val Val Phe Glu Asp Gln Asp Ala Leu Leu Pro
Cys Leu 115 120 125Leu Thr Asp Pro Val Leu Glu Ala Gly Val Ser Leu
Val Arg Val Arg 130 135 140Gly Arg Pro Leu Met Arg His Thr Asn Tyr
Ser Phe Ser Pro Trp His145 150 155 160Gly Phe Thr Ile His Arg Ala
Lys Phe Ile Gln Ser Gln Asp Tyr Gln 165 170 175Cys Ser Ala Leu Met
Gly Gly Arg Lys Val Met Ser Ile Ser Ile Arg 180 185 190Leu Lys Val
Gln Lys Val Ile Pro Gly Pro Pro Ala Leu Thr Leu Val 195 200 205Pro
Ala Glu Leu Val Arg Ile Arg Gly Glu Ala Ala Gln Ile Val Cys 210 215
220Ser Ala Ser Ser Val Asp Val Asn Phe Asp Val Phe Leu Gln His
Asn225 230 235 240Asn Thr Lys Leu Ala Ile Pro Gln Gln Ser Asp Phe
His Asn Asn Arg 245 250 255Tyr Gln Lys Val Leu Thr Leu Asn Leu Asp
Gln Val Asp Phe Gln His 260 265 270Ala Gly Asn Tyr Ser Cys Val Ala
Ser Asn Val Gln Gly Lys His Ser 275 280 285Thr Ser Met Phe Phe Arg
Val Val Glu Ser Ala Tyr Leu Asn Leu Ser 290 295 300Ser Glu Gln Asn
Leu Ile Gln Glu Val Thr Val Gly Glu Gly Leu Asn305 310 315 320Leu
Lys Val Met Val Glu Ala Tyr Pro Gly Leu Gln Gly Phe Asn Trp 325 330
335Thr Tyr Leu Gly Pro Phe Ser Asp His Gln Pro Glu Pro Lys Leu Ala
340 345 350Asn Ala Thr Thr Lys Asp Thr Tyr Arg His Thr Phe Thr Leu
Ser Leu 355 360 365Pro Arg Leu Lys Pro Ser Glu Ala Gly Arg Tyr Ser
Phe Leu Ala Arg 370 375 380Asn Pro Gly Gly Trp Arg Ala Leu Thr Phe
Glu Leu Thr Leu Arg Tyr385 390 395 400Pro Pro Glu Val Ser Val Ile
Trp Thr Phe Ile Asn Gly Ser Gly Thr 405 410 415Leu Leu Cys Ala Ala
Ser Gly Tyr Pro Gln Pro Asn Val Thr Trp Leu 420 425 430Gln Cys Ser
Gly His Thr Asp Arg Cys Asp Glu Ala Gln Val Leu Gln 435 440 445Val
Trp Asp Asp Pro Tyr Pro Glu Val Leu Ser Gln Glu Pro Phe His 450 455
460Lys Val Thr Val Gln Ser Leu Leu Thr Val Glu Thr Leu Glu His
Asn465 470 475 480Gln Thr Tyr Glu Cys Arg Ala His Asn Ser Val Gly
Ser Gly Ser Trp 485 490 495Ala Phe Ile Pro Ile Ser Ala Gly Ala His
Thr His Pro Pro Asp Glu 500 505 510Phe Leu Phe Thr Pro Val Val Val
Ala Cys Met Ser Ile Met Ala Leu 515 520 525Leu Leu Leu Leu Leu Leu
Leu Leu Leu Tyr Lys Tyr Lys Gln Lys Pro 530 535 540Lys Tyr Gln Val
Arg Trp Lys Ile Ile Glu Ser Tyr Glu Gly Asn Ser545 550 555 560Tyr
Thr Phe Ile Asp Pro Thr Gln Leu Pro Tyr Asn Glu Lys Trp Glu 565 570
575Phe Pro Arg Asn Asn Leu Gln Phe Gly Lys Thr Leu Gly Ala Gly Ala
580 585 590Phe Gly Lys Val Val Glu Ala Thr Ala Phe Gly Leu Gly Lys
Glu Asp 595 600 605Ala Val Leu Lys Val Ala Val Lys Met Leu Lys Ser
Thr Ala His Ala 610 615 620Asp Glu Lys Glu Ala Leu Met Ser Glu Leu
Lys Ile Met Ser His Leu625 630 635 640Gly Gln His Glu Asn Ile Val
Asn Leu Leu Gly Ala Cys Thr His Gly 645 650 655Gly Pro Val Leu Val
Ile Thr Glu Tyr Cys Cys Tyr Gly Asp Leu Leu 660 665 670Asn Phe Leu
Arg Arg Lys Ala Glu Ala Met Leu Gly Pro Ser Leu Ser 675 680 685Pro
Gly Gln Asp Pro Glu Gly Gly Val Asp Tyr Lys Asn Ile His Leu 690 695
700Glu Lys Lys Tyr Val Arg Arg Asp Ser Gly Phe Ser Ser Gln Gly
Val705 710 715 720Asp Thr Tyr Val Glu Met Arg Pro Val Ser Thr Ser
Ser Asn Asp Ser 725 730 735Phe Ser Glu Gln Asp Leu Asp Lys Glu Asp
Gly Arg Pro Leu Glu Leu 740 745 750Arg Asp Leu Leu His Phe Ser Ser
Gln Val Ala Gln Gly Met Ala Phe 755 760 765Leu Ala Ser Lys Asn Cys
Ile His Arg Asp Val Ala Ala Arg Asn Val 770 775 780Leu Leu Thr Asn
Gly His Val Ala Lys Ile Gly Asp Phe Gly Leu Ala785 790 795 800Arg
Asp Ile Met Asn Asp Ser Asn Tyr Ile Val Lys Gly Asn Ala Arg 805 810
815Leu Pro Val Lys Trp Met Ala Pro Glu Ser Ile Phe Asp Cys Val Tyr
820 825 830Thr Val Gln Ser Asp Val Trp Ser Tyr Gly Ile Leu Leu Trp
Glu Ile 835 840 845Phe Ser Leu Gly Leu Asn Pro Tyr Pro Gly Ile Leu
Val Asn Ser Lys 850 855 860Phe Tyr Lys Leu Val Lys Asp Gly Tyr Gln
Met Ala Gln Pro Ala Phe865 870 875 880Ala Pro Lys Asn Ile Tyr Ser
Ile Met Gln Ala Cys Trp Ala Leu Glu 885 890 895Pro Thr His Arg Pro
Thr Phe Gln Gln Ile Cys Ser Phe Leu Gln Glu 900 905 910Gln Ala Gln
Glu Asp Arg Arg Glu Arg Asp Tyr Thr Asn Leu Pro Ser 915 920 925Ser
Ser Arg Ser Gly Gly Ser Gly Ser Ser Ser Ser Glu Leu Glu Glu 930 935
940Glu Ser Ser Ser Glu His Leu Thr Cys Cys Glu Gln Gly Asp Ile
Ala945 950 955 960Gln Pro Leu Leu Gln Pro Asn Asn Tyr Gln Phe Cys
965 97063972PRTArtificialmutant CSF-1R L301S Y969F 63Met Gly Pro
Gly Val Leu Leu Leu Leu Leu Val Ala Thr Ala Trp His1 5 10 15Gly Gln
Gly Ile Pro Val Ile Glu Pro Ser Val Pro Glu Leu Val Val 20 25 30Lys
Pro Gly Ala Thr Val Thr Leu Arg Cys Val Gly Asn Gly Ser Val 35 40
45Glu Trp Asp Gly Pro Pro Ser Pro His Trp Thr Leu Tyr Ser Asp Gly
50 55 60Ser Ser Ser Ile Leu Ser Thr Asn Asn Ala Thr Phe Gln Asn Thr
Gly65 70 75 80Thr Tyr Arg Cys Thr Glu Pro Gly Asp Pro Leu Gly Gly
Ser Ala Ala 85 90 95Ile His Leu Tyr Val Lys Asp Pro Ala Arg Pro Trp
Asn Val Leu Ala 100 105 110Gln Glu Val Val Val Phe Glu Asp Gln Asp
Ala Leu Leu Pro Cys Leu 115 120 125Leu Thr Asp Pro Val Leu Glu Ala
Gly Val Ser Leu Val Arg Val Arg 130 135 140Gly Arg Pro Leu Met Arg
His Thr Asn Tyr Ser Phe Ser Pro Trp His145 150 155 160Gly Phe Thr
Ile His Arg Ala Lys Phe Ile Gln Ser Gln Asp Tyr Gln 165 170 175Cys
Ser Ala Leu Met Gly Gly Arg Lys Val Met Ser Ile Ser Ile Arg 180 185
190Leu Lys Val Gln Lys Val Ile Pro Gly Pro Pro Ala Leu Thr Leu Val
195 200 205Pro Ala Glu Leu Val Arg Ile Arg Gly Glu Ala Ala Gln Ile
Val Cys 210 215 220Ser Ala Ser Ser Val Asp Val Asn Phe Asp Val Phe
Leu Gln His Asn225 230 235
240Asn Thr Lys Leu Ala Ile Pro Gln Gln Ser Asp Phe His Asn Asn Arg
245 250 255Tyr Gln Lys Val Leu Thr Leu Asn Leu Asp Gln Val Asp Phe
Gln His 260 265 270Ala Gly Asn Tyr Ser Cys Val Ala Ser Asn Val Gln
Gly Lys His Ser 275 280 285Thr Ser Met Phe Phe Arg Val Val Glu Ser
Ala Tyr Ser Asn Leu Ser 290 295 300Ser Glu Gln Asn Leu Ile Gln Glu
Val Thr Val Gly Glu Gly Leu Asn305 310 315 320Leu Lys Val Met Val
Glu Ala Tyr Pro Gly Leu Gln Gly Phe Asn Trp 325 330 335Thr Tyr Leu
Gly Pro Phe Ser Asp His Gln Pro Glu Pro Lys Leu Ala 340 345 350Asn
Ala Thr Thr Lys Asp Thr Tyr Arg His Thr Phe Thr Leu Ser Leu 355 360
365Pro Arg Leu Lys Pro Ser Glu Ala Gly Arg Tyr Ser Phe Leu Ala Arg
370 375 380Asn Pro Gly Gly Trp Arg Ala Leu Thr Phe Glu Leu Thr Leu
Arg Tyr385 390 395 400Pro Pro Glu Val Ser Val Ile Trp Thr Phe Ile
Asn Gly Ser Gly Thr 405 410 415Leu Leu Cys Ala Ala Ser Gly Tyr Pro
Gln Pro Asn Val Thr Trp Leu 420 425 430Gln Cys Ser Gly His Thr Asp
Arg Cys Asp Glu Ala Gln Val Leu Gln 435 440 445Val Trp Asp Asp Pro
Tyr Pro Glu Val Leu Ser Gln Glu Pro Phe His 450 455 460Lys Val Thr
Val Gln Ser Leu Leu Thr Val Glu Thr Leu Glu His Asn465 470 475
480Gln Thr Tyr Glu Cys Arg Ala His Asn Ser Val Gly Ser Gly Ser Trp
485 490 495Ala Phe Ile Pro Ile Ser Ala Gly Ala His Thr His Pro Pro
Asp Glu 500 505 510Phe Leu Phe Thr Pro Val Val Val Ala Cys Met Ser
Ile Met Ala Leu 515 520 525Leu Leu Leu Leu Leu Leu Leu Leu Leu Tyr
Lys Tyr Lys Gln Lys Pro 530 535 540Lys Tyr Gln Val Arg Trp Lys Ile
Ile Glu Ser Tyr Glu Gly Asn Ser545 550 555 560Tyr Thr Phe Ile Asp
Pro Thr Gln Leu Pro Tyr Asn Glu Lys Trp Glu 565 570 575Phe Pro Arg
Asn Asn Leu Gln Phe Gly Lys Thr Leu Gly Ala Gly Ala 580 585 590Phe
Gly Lys Val Val Glu Ala Thr Ala Phe Gly Leu Gly Lys Glu Asp 595 600
605Ala Val Leu Lys Val Ala Val Lys Met Leu Lys Ser Thr Ala His Ala
610 615 620Asp Glu Lys Glu Ala Leu Met Ser Glu Leu Lys Ile Met Ser
His Leu625 630 635 640Gly Gln His Glu Asn Ile Val Asn Leu Leu Gly
Ala Cys Thr His Gly 645 650 655Gly Pro Val Leu Val Ile Thr Glu Tyr
Cys Cys Tyr Gly Asp Leu Leu 660 665 670Asn Phe Leu Arg Arg Lys Ala
Glu Ala Met Leu Gly Pro Ser Leu Ser 675 680 685Pro Gly Gln Asp Pro
Glu Gly Gly Val Asp Tyr Lys Asn Ile His Leu 690 695 700Glu Lys Lys
Tyr Val Arg Arg Asp Ser Gly Phe Ser Ser Gln Gly Val705 710 715
720Asp Thr Tyr Val Glu Met Arg Pro Val Ser Thr Ser Ser Asn Asp Ser
725 730 735Phe Ser Glu Gln Asp Leu Asp Lys Glu Asp Gly Arg Pro Leu
Glu Leu 740 745 750Arg Asp Leu Leu His Phe Ser Ser Gln Val Ala Gln
Gly Met Ala Phe 755 760 765Leu Ala Ser Lys Asn Cys Ile His Arg Asp
Val Ala Ala Arg Asn Val 770 775 780Leu Leu Thr Asn Gly His Val Ala
Lys Ile Gly Asp Phe Gly Leu Ala785 790 795 800Arg Asp Ile Met Asn
Asp Ser Asn Tyr Ile Val Lys Gly Asn Ala Arg 805 810 815Leu Pro Val
Lys Trp Met Ala Pro Glu Ser Ile Phe Asp Cys Val Tyr 820 825 830Thr
Val Gln Ser Asp Val Trp Ser Tyr Gly Ile Leu Leu Trp Glu Ile 835 840
845Phe Ser Leu Gly Leu Asn Pro Tyr Pro Gly Ile Leu Val Asn Ser Lys
850 855 860Phe Tyr Lys Leu Val Lys Asp Gly Tyr Gln Met Ala Gln Pro
Ala Phe865 870 875 880Ala Pro Lys Asn Ile Tyr Ser Ile Met Gln Ala
Cys Trp Ala Leu Glu 885 890 895Pro Thr His Arg Pro Thr Phe Gln Gln
Ile Cys Ser Phe Leu Gln Glu 900 905 910Gln Ala Gln Glu Asp Arg Arg
Glu Arg Asp Tyr Thr Asn Leu Pro Ser 915 920 925Ser Ser Arg Ser Gly
Gly Ser Gly Ser Ser Ser Ser Glu Leu Glu Glu 930 935 940Glu Ser Ser
Ser Glu His Leu Thr Cys Cys Glu Gln Gly Asp Ile Ala945 950 955
960Gln Pro Leu Leu Gln Pro Asn Asn Phe Gln Phe Cys 965
97064493PRTArtificialhuman CSF-1R Extracellular Domain 64Ile Pro
Val Ile Glu Pro Ser Val Pro Glu Leu Val Val Lys Pro Gly1 5 10 15Ala
Thr Val Thr Leu Arg Cys Val Gly Asn Gly Ser Val Glu Trp Asp 20 25
30Gly Pro Pro Ser Pro His Trp Thr Leu Tyr Ser Asp Gly Ser Ser Ser
35 40 45Ile Leu Ser Thr Asn Asn Ala Thr Phe Gln Asn Thr Gly Thr Tyr
Arg 50 55 60Cys Thr Glu Pro Gly Asp Pro Leu Gly Gly Ser Ala Ala Ile
His Leu65 70 75 80Tyr Val Lys Asp Pro Ala Arg Pro Trp Asn Val Leu
Ala Gln Glu Val 85 90 95Val Val Phe Glu Asp Gln Asp Ala Leu Leu Pro
Cys Leu Leu Thr Asp 100 105 110Pro Val Leu Glu Ala Gly Val Ser Leu
Val Arg Val Arg Gly Arg Pro 115 120 125Leu Met Arg His Thr Asn Tyr
Ser Phe Ser Pro Trp His Gly Phe Thr 130 135 140Ile His Arg Ala Lys
Phe Ile Gln Ser Gln Asp Tyr Gln Cys Ser Ala145 150 155 160Leu Met
Gly Gly Arg Lys Val Met Ser Ile Ser Ile Arg Leu Lys Val 165 170
175Gln Lys Val Ile Pro Gly Pro Pro Ala Leu Thr Leu Val Pro Ala Glu
180 185 190Leu Val Arg Ile Arg Gly Glu Ala Ala Gln Ile Val Cys Ser
Ala Ser 195 200 205Ser Val Asp Val Asn Phe Asp Val Phe Leu Gln His
Asn Asn Thr Lys 210 215 220Leu Ala Ile Pro Gln Gln Ser Asp Phe His
Asn Asn Arg Tyr Gln Lys225 230 235 240Val Leu Thr Leu Asn Leu Asp
Gln Val Asp Phe Gln His Ala Gly Asn 245 250 255Tyr Ser Cys Val Ala
Ser Asn Val Gln Gly Lys His Ser Thr Ser Met 260 265 270Phe Phe Arg
Val Val Glu Ser Ala Tyr Leu Asn Leu Ser Ser Glu Gln 275 280 285Asn
Leu Ile Gln Glu Val Thr Val Gly Glu Gly Leu Asn Leu Lys Val 290 295
300Met Val Glu Ala Tyr Pro Gly Leu Gln Gly Phe Asn Trp Thr Tyr
Leu305 310 315 320Gly Pro Phe Ser Asp His Gln Pro Glu Pro Lys Leu
Ala Asn Ala Thr 325 330 335Thr Lys Asp Thr Tyr Arg His Thr Phe Thr
Leu Ser Leu Pro Arg Leu 340 345 350Lys Pro Ser Glu Ala Gly Arg Tyr
Ser Phe Leu Ala Arg Asn Pro Gly 355 360 365Gly Trp Arg Ala Leu Thr
Phe Glu Leu Thr Leu Arg Tyr Pro Pro Glu 370 375 380Val Ser Val Ile
Trp Thr Phe Ile Asn Gly Ser Gly Thr Leu Leu Cys385 390 395 400Ala
Ala Ser Gly Tyr Pro Gln Pro Asn Val Thr Trp Leu Gln Cys Ser 405 410
415Gly His Thr Asp Arg Cys Asp Glu Ala Gln Val Leu Gln Val Trp Asp
420 425 430Asp Pro Tyr Pro Glu Val Leu Ser Gln Glu Pro Phe His Lys
Val Thr 435 440 445Val Gln Ser Leu Leu Thr Val Glu Thr Leu Glu His
Asn Gln Thr Tyr 450 455 460Glu Cys Arg Ala His Asn Ser Val Gly Ser
Gly Ser Trp Ala Phe Ile465 470 475 480Pro Ile Ser Ala Gly Ala His
Thr His Pro Pro Asp Glu 485 49065388PRTArtificialhuman CSF-1R
fragment delD4 65Ile Pro Val Ile Glu Pro Ser Val Pro Glu Leu Val
Val Lys Pro Gly1 5 10 15Ala Thr Val Thr Leu Arg Cys Val Gly Asn Gly
Ser Val Glu Trp Asp 20 25 30Gly Pro Pro Ser Pro His Trp Thr Leu Tyr
Ser Asp Gly Ser Ser Ser 35 40 45Ile Leu Ser Thr Asn Asn Ala Thr Phe
Gln Asn Thr Gly Thr Tyr Arg 50 55 60Cys Thr Glu Pro Gly Asp Pro Leu
Gly Gly Ser Ala Ala Ile His Leu65 70 75 80Tyr Val Lys Asp Pro Ala
Arg Pro Trp Asn Val Leu Ala Gln Glu Val 85 90 95Val Val Phe Glu Asp
Gln Asp Ala Leu Leu Pro Cys Leu Leu Thr Asp 100 105 110Pro Val Leu
Glu Ala Gly Val Ser Leu Val Arg Val Arg Gly Arg Pro 115 120 125Leu
Met Arg His Thr Asn Tyr Ser Phe Ser Pro Trp His Gly Phe Thr 130 135
140Ile His Arg Ala Lys Phe Ile Gln Ser Gln Asp Tyr Gln Cys Ser
Ala145 150 155 160Leu Met Gly Gly Arg Lys Val Met Ser Ile Ser Ile
Arg Leu Lys Val 165 170 175Gln Lys Val Ile Pro Gly Pro Pro Ala Leu
Thr Leu Val Pro Ala Glu 180 185 190Leu Val Arg Ile Arg Gly Glu Ala
Ala Gln Ile Val Cys Ser Ala Ser 195 200 205Ser Val Asp Val Asn Phe
Asp Val Phe Leu Gln His Asn Asn Thr Lys 210 215 220Leu Ala Ile Pro
Gln Gln Ser Asp Phe His Asn Asn Arg Tyr Gln Lys225 230 235 240Val
Leu Thr Leu Asn Leu Asp Gln Val Asp Phe Gln His Ala Gly Asn 245 250
255Tyr Ser Cys Val Ala Ser Asn Val Gln Gly Lys His Ser Thr Ser Met
260 265 270Phe Phe Arg Tyr Pro Pro Glu Val Ser Val Ile Trp Thr Phe
Ile Asn 275 280 285Gly Ser Gly Thr Leu Leu Cys Ala Ala Ser Gly Tyr
Pro Gln Pro Asn 290 295 300Val Thr Trp Leu Gln Cys Ser Gly His Thr
Asp Arg Cys Asp Glu Ala305 310 315 320Gln Val Leu Gln Val Trp Asp
Asp Pro Tyr Pro Glu Val Leu Ser Gln 325 330 335Glu Pro Phe His Lys
Val Thr Val Gln Ser Leu Leu Thr Val Glu Thr 340 345 350Leu Glu His
Asn Gln Thr Tyr Glu Cys Arg Ala His Asn Ser Val Gly 355 360 365Ser
Gly Ser Trp Ala Phe Ile Pro Ile Ser Ala Gly Ala His Thr His 370 375
380Pro Pro Asp Glu38566292PRTArtificialhuman CSF-1R fragment D1-D3
66Ile Pro Val Ile Glu Pro Ser Val Pro Glu Leu Val Val Lys Pro Gly1
5 10 15Ala Thr Val Thr Leu Arg Cys Val Gly Asn Gly Ser Val Glu Trp
Asp 20 25 30Gly Pro Pro Ser Pro His Trp Thr Leu Tyr Ser Asp Gly Ser
Ser Ser 35 40 45Ile Leu Ser Thr Asn Asn Ala Thr Phe Gln Asn Thr Gly
Thr Tyr Arg 50 55 60Cys Thr Glu Pro Gly Asp Pro Leu Gly Gly Ser Ala
Ala Ile His Leu65 70 75 80Tyr Val Lys Asp Pro Ala Arg Pro Trp Asn
Val Leu Ala Gln Glu Val 85 90 95Val Val Phe Glu Asp Gln Asp Ala Leu
Leu Pro Cys Leu Leu Thr Asp 100 105 110Pro Val Leu Glu Ala Gly Val
Ser Leu Val Arg Val Arg Gly Arg Pro 115 120 125Leu Met Arg His Thr
Asn Tyr Ser Phe Ser Pro Trp His Gly Phe Thr 130 135 140Ile His Arg
Ala Lys Phe Ile Gln Ser Gln Asp Tyr Gln Cys Ser Ala145 150 155
160Leu Met Gly Gly Arg Lys Val Met Ser Ile Ser Ile Arg Leu Lys Val
165 170 175Gln Lys Val Ile Pro Gly Pro Pro Ala Leu Thr Leu Val Pro
Ala Glu 180 185 190Leu Val Arg Ile Arg Gly Glu Ala Ala Gln Ile Val
Cys Ser Ala Ser 195 200 205Ser Val Asp Val Asn Phe Asp Val Phe Leu
Gln His Asn Asn Thr Lys 210 215 220Leu Ala Ile Pro Gln Gln Ser Asp
Phe His Asn Asn Arg Tyr Gln Lys225 230 235 240Val Leu Thr Leu Asn
Leu Asp Gln Val Asp Phe Gln His Ala Gly Asn 245 250 255Tyr Ser Cys
Val Ala Ser Asn Val Gln Gly Lys His Ser Thr Ser Met 260 265 270Phe
Phe Arg Val Val Glu Ser Ala Tyr Leu Asn Leu Ser Ser Glu Gln 275 280
285Asn Leu Ile Gln 2906721PRTArtificialsignal peptide 67Met Gly Ser
Gly Pro Gly Val Leu Leu Leu Leu Leu Val Ala Thr Ala1 5 10 15Trp His
Gly Gln Gly 206836DNAArtificialPrimer 68cacctccatg ttcttccggt
accccccaga ggtaag 36698PRTMus musculus 69Asp Leu Arg Leu Tyr Phe
Asp Val1 57016PRTMus musculus 70Val Ile Trp Ser Gly Gly Gly Thr Asn
Tyr Asn Ser Pro Phe Met Ser1 5 10 157110PRTMus musculus 71Gly Phe
Ser Leu Thr Ser Tyr Asp Ile Ser1 5 10728PRTMus musculus 72Gly Gln
Ser Phe Thr Tyr Pro Thr1 5737PRTMus musculus 73Gly Ser Ser Asn Arg
Tyr Thr1 57411PRTMus musculus 74Lys Ala Ser Glu Asp Val Gly Thr Tyr
Val Ser1 5 1075116PRTMus musculus 75Arg Val Gln Leu Lys Glu Ser Gly
Pro Gly Leu Val Ala Pro Ser Gln1 5 10 15Ser Leu Ser Ile Thr Cys Thr
Val Ser Gly Phe Ser Leu Thr Ser Tyr 20 25 30Asp Ile Ser Trp Ile Arg
Gln Ser Pro Gly Lys Gly Leu Glu Trp Leu 35 40 45Gly Val Ile Trp Ser
Gly Gly Gly Thr Asn Tyr Asn Ser Pro Phe Met 50 55 60Ser Arg Leu Arg
Ile Ser Lys Asp Asp Ser Arg Ser Gln Val Phe Leu65 70 75 80Lys Val
Asn Arg Leu Gln Thr Asp Asp Thr Ala Ile Tyr Tyr Cys Val 85 90 95Arg
Asp Leu Arg Leu Tyr Phe Asp Val Trp Gly Ala Gly Thr Thr Val 100 105
110Thr Val Ser Ser 11576106PRTMus musculus 76Lys Ile Val Met Thr
Gln Ser Pro Lys Ser Met Ser Val Ser Val Gly1 5 10 15Glu Arg Val Ser
Leu Ser Cys Lys Ala Ser Glu Asp Val Gly Thr Tyr 20 25 30Val Ser Trp
Tyr Gln Gln Lys Pro Glu Gln Ser Pro Lys Leu Leu Ile 35 40 45Tyr Gly
Ser Ser Asn Arg Tyr Thr Gly Val Pro Asp Arg Phe Thr Gly 50 55 60Ser
Gly Ser Ala Thr Asp Phe Thr Leu Thr Ile Ser Ser Val Gln Ala65 70 75
80Glu Asp Leu Ala Asp Tyr Ser Cys Gly Gln Ser Phe Thr Tyr Pro Thr
85 90 95Phe Gly Thr Gly Thr Lys Leu Glu Ile Lys 100 105778PRTMus
musculus 77Asp Pro Arg Leu Tyr Phe Asp Val1 57816PRTMus musculus
78Val Ile Trp Thr Gly Gly Gly Thr Asn Tyr Asn Ser Gly Phe Met Ser1
5 10 157910PRTMus musculus 79Gly Ser Ser Leu Asp Ser Phe Asp Ile
Ser1 5 10808PRTMus musculus 80Gly Gln Thr Phe Ser Tyr Pro Thr1
5817PRTMus musculus 81Gly Ala Ser Asn Arg Tyr Thr1 58211PRTMus
musculus 82Lys Ala Ser Glu Asp Val Val Thr Tyr Val Ser1 5
1083116PRTMus musculus 83Gln Val Gln Leu Lys Glu Ser Gly Pro Gly
Leu Val Ala Pro Ser Lys1 5 10 15Ser Leu Ser Ile Thr Cys Thr Val Ser
Gly Ser Ser Leu Asp Ser Phe 20 25 30Asp Ile Ser Trp Ile Arg Gln Pro
Pro Gly Lys Gly Leu Glu Trp Leu 35 40 45Gly Val Ile Trp Thr Gly Gly
Gly Thr Asn Tyr Asn Ser Gly Phe Met 50 55 60Ser Arg Leu Arg Ile Ser
Lys Asp Asn Ser Lys Ser Gln Val Phe Leu65 70 75 80Lys Met Ser Ser
Leu Gln Ser Asp Asp Thr Ala Ile Tyr Tyr Cys Val 85 90 95Arg Asp Pro
Arg Leu Tyr Phe Asp Val Trp Gly Ala Gly Thr Thr Val 100 105 110Thr
Val Ser Ser 11584106PRTMus musculus 84Asn Ile Val Met Thr Gln Ser
Pro Lys Ser Met Ser Met Ser Val Gly1 5
10 15Glu Arg Val Thr Leu Ser Cys Lys Ala Ser Glu Asp Val Val Thr
Tyr 20 25 30Val Ser Trp Tyr Gln Gln Lys Pro Glu Gln Ser Pro Lys Leu
Leu Ile 35 40 45Tyr Gly Ala Ser Asn Arg Tyr Thr Gly Val Pro Asp Arg
Phe Thr Gly 50 55 60Ser Gly Ser Ala Thr Asp Phe Thr Leu Thr Ile Ser
Ser Ile Gln Ala65 70 75 80Glu Asp Leu Ala Asp Tyr Tyr Cys Gly Gln
Thr Phe Ser Tyr Pro Thr 85 90 95Phe Gly Thr Gly Thr Lys Leu Glu Ile
Lys 100 105
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.