Cns Targeting Aav Vectors And Methods Of Use Thereof
Gao; Guangping ; et al.
U.S. patent application number 16/364763 was filed with the patent office on 2019-09-12 for cns targeting aav vectors and methods of use thereof. This patent application is currently assigned to University of Massachusetts. The applicant listed for this patent is University of Massachusetts. Invention is credited to Guangping Gao, Hongyan Wang, Zuoshang Xu, Hongwei Zhang.
| Application Number | 20190276848 16/364763 |
| Document ID | / |
| Family ID | 44834531 |
| Filed Date | 2019-09-12 |











View All Diagrams
| United States Patent Application | 20190276848 |
| Kind Code | A1 |
| Gao; Guangping ; et al. | September 12, 2019 |
CNS TARGETING AAV VECTORS AND METHODS OF USE THEREOF
Abstract
The invention in some aspects relates to recombinant adeno-associated viruses useful for targeting transgenes to CNS tissue, and compositions comprising the same, and methods of use thereof. In some aspects, the invention provides methods and compositions for treating CNS-related disorders.
| Inventors: | Gao; Guangping; (Westborough, MA) ; Zhang; Hongwei; (Worcester, MA) ; Wang; Hongyan; (Worcester, MA) ; Xu; Zuoshang; (Wellesley, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | University of Massachusetts Boston MA |
||||||||||
| Family ID: | 44834531 | ||||||||||
| Appl. No.: | 16/364763 | ||||||||||
| Filed: | March 26, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 16234819 | Dec 28, 2018 | |||
| 16364763 | ||||
| 15613646 | Jun 5, 2017 | |||
| 16234819 | ||||
| 16234834 | Dec 28, 2018 | |||
| 15613646 | ||||
| 15613646 | Jun 5, 2017 | |||
| 16234834 | ||||
| 14445670 | Jul 29, 2014 | 9701984 | ||
| 15613646 | ||||
| 13642719 | Jan 7, 2013 | 9102949 | ||
| PCT/US2011/033616 | Apr 22, 2011 | |||
| 14445670 | ||||
| 61327627 | Apr 23, 2010 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 15/8645 20130101; C12N 15/1137 20130101; C12N 2750/14141 20130101; C12N 2840/007 20130101; C12N 15/635 20130101; C12Y 305/01015 20130101; C12N 9/80 20130101; C12N 2750/14145 20130101; C12N 2750/14143 20130101; C12N 2750/14162 20130101; A61K 48/0058 20130101; A61K 48/0075 20130101; C12N 2810/10 20130101; C12N 2310/141 20130101; A61P 25/00 20180101; C12N 2750/14133 20130101; A61K 31/713 20130101; C12N 7/00 20130101; C12N 15/86 20130101; A61K 48/00 20130101; A61K 38/50 20130101 |
| International Class: | C12N 15/86 20060101 C12N015/86; A61K 38/50 20060101 A61K038/50; C12N 7/00 20060101 C12N007/00; A61K 48/00 20060101 A61K048/00; C12N 9/80 20060101 C12N009/80; C12N 15/864 20060101 C12N015/864; C12N 15/113 20060101 C12N015/113; A61K 31/713 20060101 A61K031/713 |
Claims
1-55. (canceled)
56. A method for treating Canavan disease in a subject, the method comprising: intrathecally, intraventricularly, or intravascularly administering rAAV to the subject in an amount effective for transducing oligodendrocytes of the subject with the rAAV, wherein the rAAV comprises (i) a capsid protein having the amino acid sequence of SEQ ID NO: 8 and (ii) a nucleic acid comprising a promoter operably linked with a region encoding aspartoacylase (ASPA), wherein the ASPA is expressed from the nucleic acid in oligodendrocytes transduced by the rAAV.
57. The method of claim 56, wherein the nucleic acid expresses an aspartoacylase (ASPA) mRNA comprising one or more miRNA binding sites for one or more miRNAs that are more abundant in one or more non-CNS tissues in comparison to a CNS tissue.
58. The method of claim 57, wherein the one or more miRNAs that are more abundant in one or more non-CNS tissues in comparison to the CNS tissue are at least twofold more abundant.
59. The method of claim 57, wherein the one or more non-CNS tissue is not kidney tissue or retinal tissue.
60. The method of claim 56 further comprising evaluating kidney function in the subject at least once after the administration.
61. The method of claim 56 further comprising evaluating vision of the subject at least once after the administration.
62. The method of claim 56, wherein the rAAV is administered intrathecally to the subject.
63. The method of claim 56, wherein the rAAV is administered intraventricularly to the subject.
64. The method of claim 56, wherein the rAAV is administered intravascularly to the subject.
Description
RELATED APPLICATIONS
[0001] This application is a continuation under 35 U.S.C. .sctn. 120 of U.S. application Ser. Nos. 16/234,819 and 16/234,834, filed Dec. 28, 2018, which are each a continuation under 35 U.S.C. .sctn. 120 of U.S. application Ser. No. 15/613,646, filed Jun. 5, 2017, which is a continuation of U.S. application Ser. No. 14/445,670, filed Jul. 29, 2014, which is a continuation of U.S. application Ser. No. 13/642,719, filed Jan. 7, 2013, which is a national stage filing under 35 U.S.C. .sctn. 371 of International Patent Application Serial No. PCT/US2011/033616, filed Apr. 22, 2011, which claims the benefit under 35 U.S.C. .sctn. 119(e) of U.S. Application Ser. No. 61/327,627, filed Apr. 23, 2010. The entire content of each of the above-referenced applications is incorporated herein by reference.
FIELD OF THE INVENTION
[0002] The invention in some aspects relates to recombinant adeno-associated viruses useful for targeting transgenes to CNS tissue, and compositions comprising the same, and methods of use thereof.
BACKGROUND OF THE INVENTION
[0003] Gene therapy has been investigated for delivery of therapeutic genes to the CNS cells for treatment of various CNS disease, e.g., Canavan disease, ALS, Parkinson disease (PD), etc. In some limited cases, therapeutic benefits have been observed using certain viruses, e.g., recombinant adenovirus (rAd), lentivirus (LV) and adeno-associated virus (AAV) to express a variety of therapeutic genes. AAV2 has been used in clinical trials for treatment of PD and Leber congenital amaurosis (an eye disease) and preliminary findings suggest symptomatic improvements without noticeable toxicity [2-4].
[0004] However, AAV-based gene therapy to treat CNS disease has still faced major obstacle. Many CNS diseases including, for example, ALS affect both cortical and spinal motor neurons that are distributed in a very broad area in the CNS. It has frequently been the case that viral vectors injected into CNS tissue transduce cells only in the vicinity of the injection site, have a very limited spread and generally have not impacted the lifespan in CNS disease animal models [See, e.g., Ref. 5]. Still, a variety of other viral administration methods have been tested. One example, involves injecting the viral particles into skeletal muscle and allowing the nerve terminals to internalize the viral genome, which is then retrogradely transported back to the spinal motor neurons. This approach has shown some positive results in certain mouse models [68]. However, to apply this method in larger mammals, like adult humans, would be impractical. Overall, the transduction efficiency observed with muscle injection is relatively low. Some investigators have tried to improve this efficiency by modifying viral capsid proteins with the nerve binding domains of tetanus toxin or botulinum toxin. These efforts have not been fruitful due to various technical difficulties. Another problem with muscle injection in larger mammals, is a need for large doses, which is technically challenging, expensive, and carries a high risk for adverse effects, ranging from immune reaction to transduction of unintended cells (e.g., germ cells).
[0005] Another method that has been evaluated for delivering transgenes into motor neurons is to inject the virus into large nerves, which maximizes that exposure of the virus to motor axons, allowing the motor neurons to internalize the viral genome and retrogradely transport them back to the cell body. This method has been demonstrated to be more efficient in transducing motor neurons than muscle injection [9]. Still, to implement a method such as this in larger mammals would be challenging.
SUMMARY OF THE INVENTION
[0006] Aspects of the invention, are based on the discovery of recombinant AAVs that achieve wide-spread distribution throughout CNS tissue of a subject. In some embodiments, the rAAVs spread throughout CNS tissue following direct administration into the cerebrospinal fluid (CSF), e.g., via intrathecal and/or intracerebral injection. In other embodiments, the rAAVs cross the blood-brain-barrier and achieve wide-spread distribution throughout CNS tissue of a subject following intravenous administration. In some aspects the invention relates to rAAVs having distinct central nervous system tissue targeting capabilities (e.g., CNS tissue tropisms), which achieve stable and nontoxic gene transfer at high efficiencies. Methods involving co-administration via intrathecal and intracerebral (e.g., intraventricular) injection of rAAVs are provided in some aspects. For example, it has been discovered that rAAVs having a capsid protein comprising a sequence as set forth in SEQ ID NO: 9 achieves wide-spread distribution following intrathecal injection throughout the CNS, and thus, are particularly useful for treating CNS-associated disorders such as, for example, ALS. In still further aspects of the invention methods are provided for treating Canavan disease.
[0007] According to some aspects of the invention, methods for delivering a transgene to CNS tissue in a subject are provided. In some embodiments, the methods comprise administering an effective amount of a rAAV by intrathecal administration, wherein the rAAV comprises (i) a capsid protein comprising a sequence as set forth in SEQ ID NO: 9 and (ii) a nucleic acid comprising a promoter operably linked with a transgene. In some embodiments, the methods further comprise administering an effective amount of the rAAV by intracerebral administration. In some embodiments, the methods comprise administering an effective amount of a rAAV by intrathecal administration and by intracerebral administration, wherein the rAAV infects cells of CNS tissue in the subject and comprises a nucleic acid comprising a promoter operably linked with a transgene. In certain embodiments, the intracerebral administration is an intraventricular administration. In one embodiment, the intraventricular administration is an administration into a ventricular region of the forebrain of the subject. In certain embodiments, the intrathecal administration is in the lumbar region of the subject. In some embodiments, the dose of the rAAV for intrathecal administration is in a range of 10.sup.10 genome copies/subject to 10.sup.11 genome copies/subject. In some embodiments, the dose of the rAAV for intrathecal administration is in a range of 10.sup.11 genome copies/subject to 10.sup.12 genome copies/subject. In some embodiments, the dose of the rAAV for intrathecal administration is in a range of 10.sup.12 genome copies/subject to 10.sup.13 genome copies/subject. In some embodiments, the dose of the rAAV for intrathecal administration is in a range of 10.sup.13 genome copies/subject to 10.sup.14 genome copies/subject. In some embodiments, the dose of the rAAV for intracerebral administration is in a range of 10.sup.10 genome copies/subject to 10.sup.11 genome copies/subject. In some embodiments, the dose of the rAAV for intracerebral administration is in a range of 10.sup.11 genome copies/subject to 10.sup.12 genome copies/subject. In some embodiments, the dose of the rAAV for intracerebral administration is in a range of 10.sup.12 genome copies/subject to 10.sup.13 genome copies/subject. In some embodiments, the dose of the rAAV for intracerebral administration is in a range of 10.sup.13 genome copies/subject to 10.sup.14 genome copies/subject. In some embodiments, the dose of the rAAV for intracerebral or intrathecal administration is formulated for injection of a volume in a range of 1 .mu.l to 10 .mu.l. In some embodiments, the dose of the rAAV for intracerebral or intrathecal administration is formulated for injection of a volume in a range of 10 .mu.l to 100 .mu.l. In some embodiments, the rAAV for the intracerebral or intrathecal administration is formulated for injection of a volume in a range of 100 .mu.l to 1 ml. In some embodiments, the rAAV for the intracerebral or intrathecal administration is formulated for injection of a volume of 1 ml or more. In some embodiments, the transgene encodes a reporter protein. In certain embodiments, the reporter protein is a fluorescent protein, an enzyme that catalyzes a reaction yielding a detectable product, or a cell surface antigen. In certain embodiments, the enzyme is a luciferase, a beta-glucuronidase, a chloramphenicol acetyltransferase, an aminoglycoside phosphotransferase, an aminocyclitol phosphotransferase, or a Puromycin N-acetyl-tranferase. In some embodiments, the transgene is a CNS-associated gene. In some embodiments, the CNS-associated gene is neuronal apoptosis inhibitory protein (NAIP), nerve growth factor (NGF), glial-derived growth factor (GDNF), brain-derived growth factor (BDNF), ciliary neurotrophic factor (CNTF), tyrosine hydroxlase (TH), GTP-cyclohydrolase (GTPCH), amino acid decorboxylase (AADC) or aspartoacylase (ASPA). In some embodiments, the transgene encodes an inhibitory RNA that binds specifically to SOD1 mRNA and inhibits expression of SOD1 in the subject. In some embodiments, the inhibitory RNA is an antisense RNA, a shRNA or a miRNA. In some embodiments, the inhibitory RNA has a sequence as set forth in SEQ ID NO: 26. Thus, according to some aspects of the invention a nucleic acid comprising a sequence as set forth in SEQ ID NO: 26 is provided. In some embodiments, a nucleic acid comprising a promoter operably linked with a region having a sequence as set forth in SEQ ID NO: 26 is provided.
[0008] In further aspects of the invention a recombinant AAV comprising a nucleic acid comprising a sequence as set forth in SEQ ID NO: 26 is provided. In some aspects of the invention a recombinant AAV comprising a nucleic acid comprising a promoter operably linked with a region having a sequence as set forth in SEQ ID NO: 26 is provided. In some embodiments the recombinant AAV further comprises a capsid protein comprising a sequence as set forth in SEQ ID NO: 9.
[0009] According to some aspects of the invention, methods for treating amyotrophic lateral sclerosis (ALS) in a subject in need thereof are provided. In some embodiments, the methods comprise administering an effective amount of a rAAV to CNS tissue of the subject, wherein the rAAV comprises (i) a capsid protein comprising a sequence as set forth in SEQ ID NO: 9 and (ii) a nucleic acid comprising a promoter operably linked with a region encoding an inhibitory RNA that binds specifically to SOD1 mRNA and inhibits expression of SOD1 in the subject. In some embodiments, the inhibitory RNA is an antisense RNA, a shRNA or a miRNA. In some embodiments, the inhibitory RNA has a sequence as set forth in SEQ ID NO: 26. In some embodiments, the methods comprise administering an effective amount of a rAAV to the subject, wherein the rAAV comprises a nucleic acid comprising a promoter operably linked with a region encoding a sequence as set forth in SEQ ID NO: 26 and wherein the rAAV infects cells of CNS tissue in the subject.
[0010] According to some aspects of the invention, methods for delivering a transgene to a CNS tissue in a subject are provided that comprise administering an effective amount of a rAAV by intravenous administration, wherein the rAAV infects cells of CNS tissue in the subject and comprises a nucleic acid comprising a promoter operably linked with a transgene. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.10 genome copies/subject to 10.sup.11 genome copies/subject. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.11 genome copies/subject to 10.sup.12 genome copies/subject. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.12 genome copies/subject to 10.sup.13 genome copies/subject. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.13 genome copies/subject to 10.sup.14 genome copies/subject. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.14 genome copies/subject to 10.sup.15 genome copies/subject. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.10 genome copies/kg to 10.sup.11 genome copies/kg. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.11 genome copies/kg to 10.sup.12 genome copies/kg. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.12 copies/kg to 10.sup.13 genome copies/kg. In some embodiments, the dose of the rAAV for intravenous administration is in a range of 10.sup.13 genome copies/kg to 10.sup.14 genome copies/kg.
[0011] According to some aspects of the invention, methods for delivering a transgene to a CNS tissue in a subject are provided that comprise administering to the subject an effective amount of a rAAV that comprises (i) a capsid protein having a sequence as set forth in any one of SEQ ID NO: 10 to 12 and (ii) a nucleic acid comprising a promoter operably linked with a transgene. In some embodiments, the methods comprise administering to the subject an effective amount of a rAAV comprising a transgene to a subject, wherein the rAAV comprises a capsid protein of a AAV serotype, or serotype variant, selected from the group consisting of: AAV1, AAV2, AAV5, AAV6, AV6.2, AAV7, AAV8, AAV9, rh.10, rh.39, rh.43 and CSp3, and wherein: (a) if the AAV serotype is AAV1, the administration route is not intracerebral, intramuscular, intranerve, or intraventricular and/or the subject is not a mouse, rat or feline; (b) if the AAV serotype is AAV2, the administration route is not intracerebral or intraventricular administration and/or the subject is not a rat, mouse, feline, marmoset, or macaque; (c) if the AAV serotype is AAV5, the administration route is not intracerebral or intraventricular administration and/or the subject is not a rat, mouse, or marmoset; (d) if the AAV serotype is AAV6, the subject is not a mouse; (e) if the AAV serotype is AAV7, the administration route is not intracerebral administration and/or the subject is not a mouse or macaque; (f) if the AAV serotype is AAV8, the administration route is not intracerebral, intraperitoneal, or intravascular administration and/or the subject is not a mouse or macaque; (g) if the AAV serotype is AAV9, the administration route is not intracerebral or intravascular administration and/or the subject is not a rat or mouse; and (h) if the AAV serotype is AAVrh.10, the administration route is not intracerebral or intravascular administration and/or the subject is not a rat or mouse. In some embodiments, the AAV serotype, or serotype variant, is selected from AAV1, AAV6, AAV7, rh.39, rh.43, and CSp3, and the administration route is intravascular administration. In some embodiments, the AAV serotype is AAV7 and the administration route is intravascular administration. In some embodiments, the CNS tissue is selected from cortex, hippocampus, thalamus, hypothalamus, cerebellum, brain stem, cervical spinal cord, thoracic spinal cord, and lumbar spinal cord. In some embodiments, the transgene encodes a reporter protein. In certain embodiments, the reporter protein is a fluorescent protein, an enzyme that catalyzes a reaction yielding a detectable product, or a cell surface antigen. In certain embodiments, the enzyme is a luciferase, a beta-glucuronidase, a chloramphenicol acetyltransferase, an aminoglycoside phosphotransferase, an aminocyclitol phosphotransferase, or a Puromycin N-acetyl-tranferase. In some embodiments, the transgene is a CNS-associated gene. In certain embodiments, the CNS-associated gene is neuronal apoptosis inhibitory protein (NAIP), nerve growth factor (NGF), glial-derived growth factor (GDNF), brain-derived growth factor (BDNF), ciliary neurotrophic factor (CNTF), tyrosine hydroxlase (TH), GTP-cyclohydrolase (GTPCH), amino acid decorboxylase (AADC) or aspartoacylase (ASPA). In some embodiments, the rAAV is administered by intravenous injection.
[0012] According to some aspects of the invention a rAAV that comprises (i) a capsid protein having a sequence as set forth in any one of SEQ ID NO: 10 to 12 and (ii) a nucleic acid comprising a promoter operably linked with a CNS-associated gene is provided. In certain embodiments, the CNS-associated gene is neuronal apoptosis inhibitory protein (NAIP), nerve growth factor (NGF), glial-derived growth factor (GDNF), brain-derived growth factor (BDNF), ciliary neurotrophic factor (CNTF), tyrosine hydroxlase (TH), GTP-cyclohydrolase (GTPCH), amino acid decorboxylase (AADC) or aspartoacylase (ASPA). In some embodiments, mRNA expressed from the CNS-associated gene comprises a miRNA binding site of a miRNA that is preferentially expressed in non-CNS tissue. In certain embodiments, the miRNA binding site is a binding site for miR-122. In certain embodiments, the miRNA binding site is a binding site for miR-1. In some embodiments, mRNA expressed from the CNS-associated gene does not comprise a miRNA binding site of a miRNA that is preferentially expressed in CNS tissue. In some embodiments, the promoter is a CNS tissue specific promoter. In certain embodiments, the promoter is a promoter of a gene selected from: neuronal nuclei (NeuN), glial fibrillary acidic protein (GFAP), adenomatous polyposis coli (APC), and ionized calcium-binding adapter molecule 1 (Iba-1).
[0013] According to some aspects of the invention, a composition comprising a rAAV that comprises (i) a capsid protein having a sequence as set forth in SEQ ID NO: 10 to 12 and (ii) a nucleic acid comprising a promoter operably linked with a CNS-associated gene is provided. In certain embodiments the composition further comprises a pharmaceutically acceptable carrier. According to some aspects of the invention, a kit comprising a container housing the composition is provided. In some embodiments, the container is a sealed vial or ampule. In some embodiments, the container is a syringe.
[0014] According to some aspects of the invention, an isolated mammalian cell is provided that comprises a nucleic acid encoding a capsid protein having a sequence as set forth in any one of SEQ ID NO: 10 to 12 and a rAAV vector comprising a nucleic acid encoding a CNS-disease associated gene. In some embodiments, the isolated mammalian cell further comprises an AAV helper function vector. In some embodiments, isolated mammalian cell further comprises an accessory function vector. In certain embodiments, the CNS-associated gene is neuronal apoptosis inhibitory protein (NAIP), nerve growth factor (NGF), glial-derived growth factor (GDNF), brain-derived growth factor (BDNF), ciliary neurotrophic factor (CNTF), tyrosine hydroxlase (TH), GTP-cyclohydrolase (GTPCH), amino acid decorboxylase (AADC) or aspartoacylase (ASPA).
[0015] According to further aspects of the invention, a method for treating Canavan disease in a subject in need thereof is provided. In some embodiments, the methods comprise administering an effective amount of a rAAV to CNS tissue of the subject, wherein the rAAV comprises (i) a capsid protein other than a capsid protein of AAV serotype 2 and (ii) a nucleic acid comprising a promoter operably linked with a region encoding aspartoacylase (ASPA). Any of the rAAV serotypes disclosed herein may be used in the methods for treating Canavan disease. In some embodiments, the rAAV has a capsid protein having an amino acid sequence as set forth in SEQ ID NO: 8 or 9 or a variant thereof. In some embodiments, administering is performed intrathecally or intracerebrally. In some embodiments, administering is performed intravascularly.
[0016] In some embodiments, the methods comprise administering an effective amount of a rAAV to CNS tissue of the subject by a route other than intracerebral administration, wherein the rAAV comprises a nucleic acid comprising a promoter operably linked with a region encoding aspartoacylase (ASPA). In some embodiments, the methods comprise administering an effective amount of a rAAV to CNS tissue of the subject, wherein the rAAV comprises a nucleic acid comprising a promoter operably linked with a region encoding aspartoacylase (ASPA); and evaluating kidney function in the subject at least once after the administration. Any suitable method known in the art may be used to evaluate a subject's kidney function. The evaluation may involve, for example, an examination of blood or urine urea nitrogen levels, an examination of blood or urine creatinine levels, a creatinine clearance rate examination, a glomerular filtration rate examination, a filtration fraction examination, a renal plasma flow examination, an ultrasound examination, a microscopic examination of a kidney tissue biopsy or any other suitable kidney function test. It should be appreciated that in some embodiments an improvement in a subject's kidney function following treatment with an rAAV-mediated gene therapy is indicative of efficacy of the gene therapy for treating Canavan disease.
[0017] In some embodiments, the methods comprise administering an effective amount of a rAAV to CNS tissue of the subject, wherein the rAAV comprises a nucleic acid comprising a promoter operably linked with a region encoding aspartoacylase (ASPA); and evaluating vision of the subject at least once after the administration. Any suitable method known in the art may be used to evaluate a subject's vision. The evaluation may involve, for example, an external examination of the eye, a visual acuity examination, an examination of pupil function, a retinal examination, an ocular motility examination, an intraocular pressure test, or an ophthalmoscopic examination. The evaluation may involve a determination regarding a subject's ability to discriminate colors, objects or shapes or the ability of a subject to discern colors, objects or shapes from a particular distance. It should be appreciated that in some embodiments an improvement in a subject's vision following treatment with an rAAV-mediated gene therapy is indicative of efficacy of the gene therapy for treating Canavan disease.
[0018] In some embodiments, the nucleic acid expresses an aspartoacylase (ASPA) mRNA comprising one or more miRNA binds sites for one or more miRNAs that are more abundant in one or more non-CNS tissues in comparison to CNS tissue. Accordingly, in some embodiments, the mRNA is targeted for degradation by an miRNA in one or more non-CNS tissues. In some embodiments, the one or more non-CNS tissue is not kidney tissue or retinal tissue. In some embodiments, the one or more miRNAs that are more abundant in non-CNS tissues in comparison to CNS tissue are at least two-fold, at least three-fold, at least four-fold, at least five-fold, or at least ten-fold more abundant. MiRNAs that are more abundant in non-CNS tissue versus CNS tissue are known in the art. For example, one study discloses the expression levels of more than three-hundred different human miRNAs in 40 different tissues, including CNS tissue, kidney tissue. (See Liang Y, et al., Characterization of microRNA expression profiles in normal human tissues. BMC Genomics. 2007 Jun. 12; 8:166, the contents of which relating to miRNAs are incorporated herein by reference). Thus, in some embodiments, the skilled artisan could readily select (e.g., based on data such as are disclosed in Liang et al.) a suitable miRNA that is more abundant in non-CNS tissue and incorporate a binding site for that miRNA into the encoded mRNA.
[0019] Each of the limitations of the invention can encompass various embodiments of the invention. It is, therefore, anticipated that each of the limitations of the invention involving any one element or combinations of elements can be included in each aspect of the invention. This invention is not limited in its application to the details of construction and the arrangement of components set forth in the following description or illustrated in the drawings. The invention is capable of other embodiments and of being practiced or of being carried out in various ways.
BRIEF DESCRIPTION OF THE DRAWINGS
[0020] FIGS. 1A to 1B depict quantitative results of EGFP intensities from fluorescence microscopic images of a panel of CNS tissue sections from neonatal mice infected with various rAAVs harboring EGFP expression vectors. Neonatal mice were administered the rAAVs by intravenous administration (superfacial temporal vein injection).
[0021] FIGS. 2A to 2B depict quantitative results of EGFP intensities from fluorescence microscopic images of a panel of CNS tissue sections from adult mice infected with various rAAVs harboring EGFP expression vectors. Adult mice were administered the rAAVs by intravenous administration (tail vein injection).
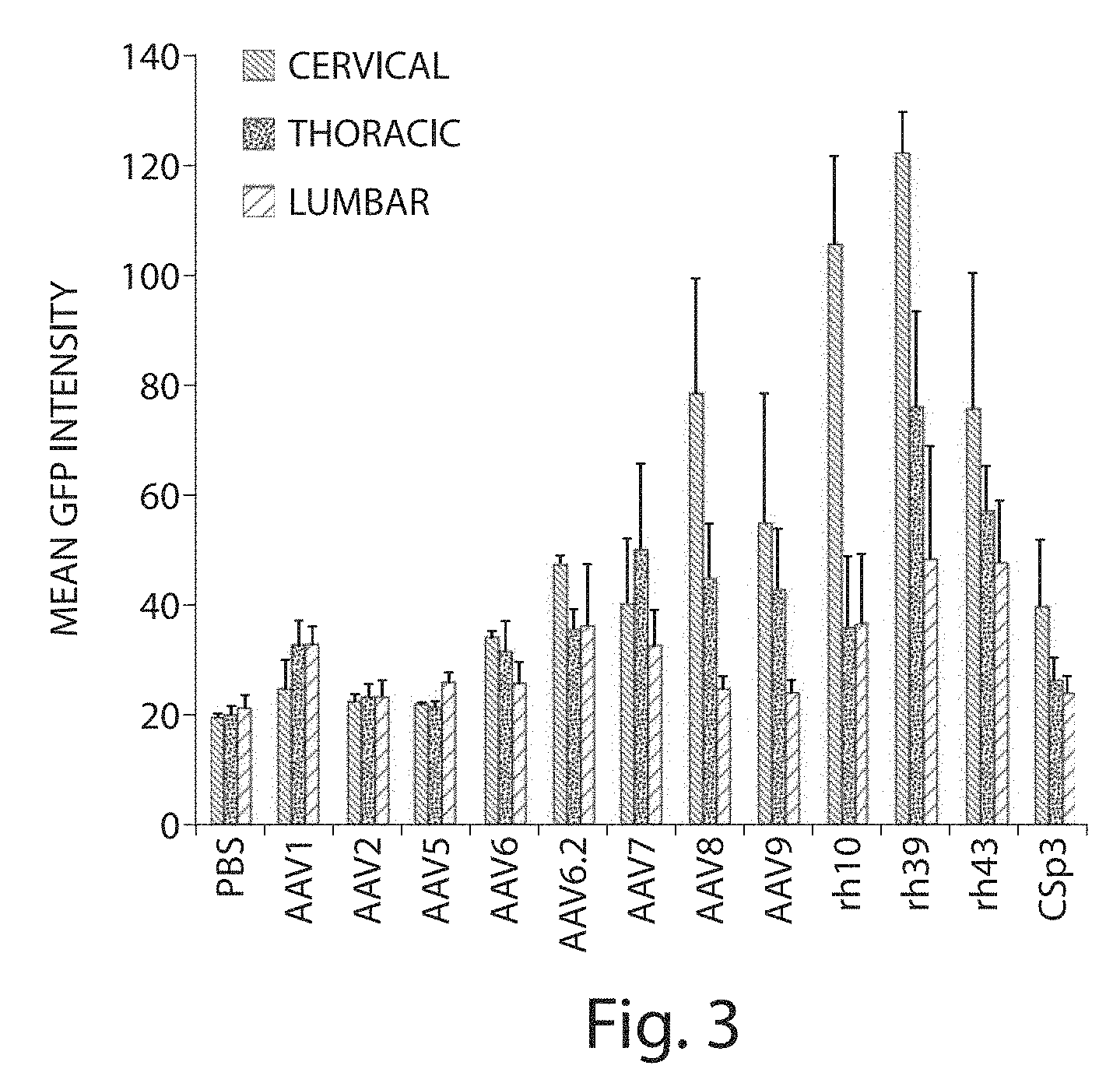
[0022] FIG. 3 depicts quantitation of EGFP expression in neonatal mice spinal cord (cervical, thoracic and lumber regions) 21 days post IV injection (5 mice per group). Neonatal mice were administered the rAAVs by intravenous administration (superfacial temporal vein injection).
[0023] FIG. 4A depicts results showing that direct CSF injection of AAVrh.10 harboring a EGFP gene leads to EGFP expression in broad areas of the CNS. Tissue sections, prepared 60 days post virus injection, from brainstem, cervical spinal cord, thoracic spinal cord and lumbar spinal cord are shown. Gray/black pixels correspond with EGFP expression.
[0024] FIG. 4B depicts results showing that direct CSF injection of AAVrh.10 harboring a EGFP gene leads to EGFP expression in astrocytes. Gray/black pixels correspond with EGFP expression.
[0025] FIG. 5A depicts a rAAVrh.10 vector that expresses a microRNA targeting SOD1. The construct employs CAG (chicken -actin promoter with a CMV enhancer) to drive the expression of EGFP and miR-SOD1 that is located in an intron in the 3'-UTR. pA stands for poly A signal. ITRs mark the inverted repeats of the AAV.
[0026] FIG. 5B depicts results of experiments that test the silencing potency of 9 different miRNA constructs, miR-SOD1#5 was found to silence SOD1 expression most potently.
[0027] FIG. 5C depicts results of experiments in which miR-SOD1#5 was packaged into AAVrh.10 and used to infect HEK293 cells. Total cellular protein was extracted 43 hours after the infection and blotted to detect SOD1. Scr stands for scrambled miRNA; Sod stands for miR-SOD1#5; and C stands for a control that expresses EGFP only.
[0028] FIG. 5D depicts a plasmid map of pAAVscCB6 EGFPmir SOD5 (5243 bp) (SEQ ID NO: 21).
[0029] FIG. 6A depicts results of gene transfer studies in SOD1 (G93A) mutant mice showing that rAAV rh.10-SOD1 miRNA knockdowns levels of mutant SOD1 in astrocytes. Staining in motor neurons was also observed.
[0030] FIG. 6B depicts results of gene transfer studies in SOD1 (G93A) mutant mice showing that rAAV rh.10-SOD1 shRNA increases live span, compared with a rAAV rh.10-scrambled miRNA.
[0031] FIG. 7A depicts quantitation of EGFP expression in cervical, thoracic, and lumber spinal cord tissue compared with life spans individual mice infected with rAAV rh.10-SOD1 miRNA; rAAV rh.10-SOD1 was administered directly to the CSF.
[0032] FIG. 7B depicts quantitation of EGFP expression in cervical, thoracic, and lumber spinal cord tissue compared with life spans of individual mice infected with rAAV rh.10-scrambled miRNA; rAAV rh.10-scrambled miRNA was administered directly to the CSF.
[0033] FIG. 8 depicts fluorescence microscopy analysis of mice that have been administered intrathecal injections of various AAVs. In this experiment, both AAV9 and AAVrh10 transduce cells along the full length of the spinal cord after a single injection into the CSF in lumbar subarachnoid space.
[0034] FIG. 9 depicts the effects of AAV10-miR-SOD1 treatment. AAV10-miR-SOD1 treatment slows disease progression as indicated by the slower loss of body weight in treated compared with the control G93A mice.
[0035] FIG. 10 depicts fluorescence microscopy analysis of mice that have been administered intrathecal injections of various AAVs. In this experiment, AAV9 and AAVrh10 can transduce cells in the broad forebrain areas after a single injection into the CSF in the third ventricle.
[0036] FIG. 11 depicts fluorescence microscopy analysis of tissue sections from AAV9-injected mice. A single injection of AAV9 and AAVrh10 into the third ventricle can transduce cells in the broad forebrain areas, including cortex, hippocampus, striatum, thalamus, cerebellum and some scattered cells in the spinal cord. The same general pattern is also observed in AAV10-injected mice.
[0037] FIGS. 12A to 12C depict an in vitro validation of artificial miRNA-binding sites for reporter silencing. Plasmids harboring the rAAVCBnLacZ genome with or without miR-1 or miR-122-binding sites were transfected into human hepatoma (HuH7) cells (FIG. 12A) which express miR-122 or cotransfected into 293 cells, together with a plasmid expressing either pri-miR-122 (FIG. 12B) or pri-miR-1 (FIG. 12C) at molar ratios of 1:3 (low) or 1:10 (high). 0.times.: no miRNA-binding site; 1.times.: one miRNA-binding site; 3.times.: three miRNA-binding sites. The cells were fixed and stained histochemically with X-gal 48 hours after transfection and blue cells counted. The percentage of nLacZ-positive cells in each transfection were compared to transfection of the control plasmid (prAAVCBnLacZ). CB, chicken .beta.-actin; miR, microRNA; nLacZ, .beta.-galactosidase reporter transgene; rAAV, recombinant adeno-associated viruses.
[0038] FIGS. 13A to 13D depict an in vivo evaluation of endogenous miRNA-mediated transgene silencing in an rAAV9 transduction. (FIGS. 13A to 13C) Adult male C588L/6 mice were injected intravenously with 5.times.10.sup.13 genome copies per kg (GC/kg) each of rAAV9CBnLacZ (no binding site), (FIG. 13A) rAAVCB9nLacZmiR-122BS (one miR-122-binding site) and rAAV9C8nlacZ-(miR-122BS).sub.3 (three miR-122-binding sites), (FIG. 13B) rAAV9CBnLacZ-miR-1 BS (one miR-1 binding site) and rAAV9CBnLacZ-(miR-1BS).sub.3 (three miR-1-binding sites, (FIG. 13C) rAAV9CBnLacZ-miR-1BS-miR-122BS (1.times. each binding site) and rAAV9CBnLacZ-(miR-1BS).sub.3-(miR-122BS).sub.3 (three miR-1 and three miR-122-binding sites). The animals were necropsied 4 weeks after vector administration, and appropriate tissues were harvested for cryosectioning and X-gal histochemical staining. miR, microRNA; nLacZ, .beta.-galactosidase reporter transgene; rAAV, recombinant adeno-associated viruses, and (FIG. 13D) quantification of .beta.-galactosidase activities in liver tissue from animals that received rAAVnLacZ vectors with and without miRNA-binding sites.
[0039] FIGS. 14A to 14E depict an analysis of expression levels of cognate miRNA, mRNA, and protein of endogenous miRNA target genes in mice transduced with rAAV9CBnLacZ with or without miRNA-binding sites. Total cellular RNA or protein was prepared from (FIGS. 14A to 14C) liver or (FIG. 14D) heart. (FIG. 14A) Northern blot detection of miRNAs. U6 small nuclear RNA provided a loading control. (FIG. 14B) Quantitative reverse-transcription PCR measuring cyclin G1 mRNA. The data are presented as relative cyclin Gl mRNA levels normalized to .beta.-actin. (FIGS. 14C to 14D) Western blot analyses of protein levels of endogenous targets of miR-122 and miR-1. Total cellular protein prepared from (FIG. 14C) liver or (FIG. 14D) heart was analyzed for cyclin G1 and calmodulin. (FIG. 14E) Serum cholesterol levels. Serum samples from mice that received rAAV9 with or without miRNA-binding sites were collected after 4 weeks and measured for total cholesterol, high-density lipoprotein (HDL) and low-density lipoprotein (LDL). miR, microRNA; nLacZ, .beta.-galactosidase reporter transgene; rAAV, recombinant adeno-associated viruses.
[0040] FIGS. 15A to 15F depict a molecular characterization of transgene mRNAs with or without miRNA binding sites. (FIG. 15A) Locations of the probes and primers, the sequences of mature miR-122 and its perfectly complementary binding site in the transgene mRNA are presented. (FIG. 15B) Total cellular RNA from liver was analyzed either by conventional reverse-transcription PCR (RT-PCR) by using primers that span a region between the 3' end of nLacZ and the 5' end of poly(A) signal (FIG. 15C) or by quantitative RT-PCR; data are presented as relative nLacZ mRNA levels normalized to .beta.-actin. (FIG. 15D) For the northern blot analysis of nLacZ mRNA, 18S RNA served as a loading control, and the blots were hybridized with either a transgene DNA (FIG. 15E) or RNA probe. (FIG. 15F) In addition, poly(A) bearing mRNA from the liver of an animal received rAAV containing three miR-1- and three miR-122-binding sites was analyzed by 5' RACE; the PCR product was resolved on an ethidium bromide-stained agarose gel. miR, microRNA; nLacZ, .beta.-galactosidase reporter transgene; rAAV, recombinant adeno-associated viruses.
[0041] FIGS. 16A to 16B depict an alignment of sequences spanning the miRNA-binding sites and poly(A) signal regions recovered by 5 RACE. Poly(A)-containing mRNA was isolated from the (FIG. 16A) liver and (FIG. 16B) heart of an animal injected with rAAV9CBnLacZ-(miR-1BS).sub.3-(miR-122BS).sub.3. Twenty-one liver-derived and twenty-two heart-derived clones were sequenced. The putative cleavage sites in each clone are identified by arrows; the frequencies of miRNA-directed, site-specific cleavage for each miRNA-binding site are reported; triangles point to the positions of the expected miRNA-directed cleavage sites (FIGS. 16A to 16B). miRNA, microRNA, nLacZ, .beta.-galactosidase reporter transgene; rAAV, recombinant adeno-associated viruses.
[0042] FIGS. 17A to 17B depict an endogenous miRNA-repressed, CNS-directed EGFP gene transfer by systemically delivered rAAV9. Ten-week-old male C57BL/6 mice were injected intravenously with scAAV9CBEGFP or scAAV9CBnLacZ(miR-1BS).sub.3-(miR-122BS).sub.3 at a dose of 2.times.10.sup.14 genome copies per kg (GC/kg) body weight. The animals were necropsied 3 weeks later for whole body fixation by transcardiac perfusion. (FIG. 17A) Brain, spinal cord, liver, heart, and muscle were harvested for cryosectioning, immunofluorescent staining for EGFP (brain and cervical spinal cord), and fluorescence microscopy to detect EGFP. Total cellular DNA and RNA were extracted from brain, liver, heart and muscle to measure the amount of persistent vector genome by qPCR and EGFP mRNA by qRT-PCR. (FIG. 17B) For each tissue, the relative abundance of the EGFP mRNA containing miRNA-binding sites was compared to that of the EGFP mRNA lacking miRNA-binding sites. For each sample, mRNA abundance was normalized to the amount of vector genome detected in the tissue. EGFP, enhanced green fluorescent protein; miRNA, microRNA; nLacZ, .beta.-galactosidase reporter transgene; qRT-PCR, quantitative reverse-transcription PCR; rAAV, recombinant adeno-associated viruses.
[0043] FIG. 18 depicts a molecular model for endogenous miRNA-regulated rAAV expression. miRNA, microRNA; rAAV, recombinant adeno-associated viruses.
[0044] FIGS. 19A to 19D depict a quantification of GFP intensity levels in the brain and spinal cord of neonatal mice transduced with various AAV vectors. 4.times.10.sup.11 genome copies (GCs) of ten different AAV vectors were injected into neonatal P1 pups through superfacial vein. The mice were sacrificed 21 days after injection. The brain tissues were extracted and 40 .mu.m thick cryosections were prepared. The sections were stained against anti-EGFP antibody. The images were analyzed and the intensity/pixel values of all AAV serotypes in various regions in brain and spinal cord (FIGS. 19A to 19C) were calculated by using Nikon NIS elements AR software version 3.2. Average intensities of the brain and spinal cord regions for different rAAVs were also presented (FIG. 19D). Region of interest (ROI) of each anatomical structure was fixed for all vectors to ensure the parallel comparison.
[0045] FIG. 20 depicts a strong and widespread EGFP expression in neonatal mouse brain after intravenous injection of rAAVs. 4.times.10.sup.11 genome copies (GCs) of rAAVs 7, 9, rh.10, rh.39 and rh.43 were injected into neonatal P1 pups through superfacial vein. The mice were sacrificed 21 days after injection. The brain tissues were extracted and 40 .mu.m thick cryosections were prepared. The sections were stained against anti-EGFP antibody. Bars represent 100 .mu.m. The regions shown are: olfactory bulb, striatum, hippocampus, cortex, hypothalamus, cerebellum and medulla.
[0046] FIG. 21 depicts EGFP expression in neonatal mouse spinal cord after intravenous injection of rAAVs. 4.times.10.sup.11 GCs of rAAVs 7, 9, rh.10, rh.39 and rh.43 were injected into neonatal P1 pups through superfacial vein. The mice were sacrificed 21 days after injection. The spinal cord tissues were extracted and 40 .mu.m thick cryosections were prepared. The sections from cervical, thoracic and lumbar regions were stained against anti-EGFP antibody. Bars represent 100 .mu.m.
[0047] FIG. 22 depicts EGFP expression in dorsal root ganglia transduced by intravascularly delivered rAAVs1, 2, 6, 6.2, 7, 9, rh.10 and rh.39. Neonatal pups received 4.times.10.sup.11 GCs of rAAVs at P1 and were necropsied 21 days after injection. Forty .mu.m thick cryosections were processed for double immunohistochemical staining for EGFP (green) and Neurons (NeuN, red). Bars represent 75 .mu.m.
[0048] FIG. 23 depicts confocal microscopic analysis of the transduced cell types in mouse CNS after systemic delivery of rAAVs to P1 neonates. The 40 .mu.m thick brain and spinal cord sections of the animals treated with different rAAVs were co-strained against anti-EGFP antibody and cell-type specific markers. Anti-NeuN was used to stain neuronal cells; anti-GFAP was used to stain astrocytes; anti-Calbindin was used to stain Purkinje cells; anti-ChAT was used to stain motor neurons; anti-DARPP was used to stain dopaminergic neurons in the substantia nigra. All rAAVs were examined, but for each cell type, only one representative picture was shown here.
[0049] FIG. 24 depicts a transduction of the brain ventricular structures by intravascularly delivered rAAVs. Neonatal pups received 4.times.10.sup.11 GCs of rAAVs at P1 and were necropsied 21 days after injection. The choroid plexuses in different ventricles were well preserved during tissue process. Forty .mu.m thick cryosections were stained against anti-EGFP antibody. Bars represent 100 .mu.m.
[0050] FIGS. 25A to 25B depict an analysis of purity and morphological integrity of rAAV vectors. FIG. 25A. Silver stained SDS-Page analysis of CsCl gradient purified rAAVCBEGFP vectors used in this study. Approximately 1.5.times.1010 virus particles each of rAAVs 1, 2, 5, 6, 6.2, 7, 9, rh10, rh39 and rh43 were loaded in the corresponding lane. FIG. 25B. Transmission electron microscopy of negative stained recombinant AAV virions. rAAV virions were spread on a freshly prepared carbon coated--Formvar support film and stained with 1% uranyl acetate for transmission microscopy. The images of virus particles from representative vector lots were taken at 92,000.times. and presented.
[0051] FIG. 26 depicts a transduction of neonatal mouse dorsal root ganglia by systemically delivered rAAVs 1, 6, 6.2 and rh43. Neonatal pups received 4.times.10.sup.11 GCs of rAAVs at P1 were necropsied 21 days after injection. Forty .mu.m thick cryosections were stained against anti-EGFP antibody. Bars represent 75 .mu.m.
[0052] FIGS. 27A to 27B depict a transduction of the brain capillary vessels by intravascularly delivered rAAVs. Neonatal pups received 4.times.10.sup.11 GCs of rAAVs at P1 were necropsied 21 days after injection. Forty .mu.m thick cryosections of the brains were stained against: (FIG. 27A) anti-EGFP antibody (AAV1, AAV6, AAV6.2, AAV7, AAV9, AAVrh.10, AAVrh.39 and AAVrh.43); (FIG. 27B) anti-EGFP and anti-CD34 antibodies (rh.10 only). Bars represent 100 .mu.m.
[0053] FIG. 28 depicts an evaluation of microgliosis in mice brain after systemic delivery of rAAVs to P1 neonates. The 40 .mu.m thick brain sections of the animals treated with different rAAVs were co-strained against anti-EGFP antibody and anti-IBa-1. Only the staining result of rAAVrh.10 was shown.
[0054] FIG. 29 depicts native EGFP expression in mice CNS after systemic delivery of rAAVs to P1 neonates. Neonatal pups received 4.times.10.sup.11 GCs of rAAVs at P1 were necropsied 21 days after injection. Forty .mu.m thick cryosections were mounted and observed under microscope without immunostaining. The exposure times for each image were indicated.
[0055] FIGS. 30A to 30E depict results showing the effects of rAAV based gene therapy in the treatment of Canavan disease. FIG. 30A shows that treatment corrected gait and motor function of the CD mice. FIG. 30B shows that treatment mitigated retinopathy and restored vision in CD mice. FIG. 30C shows that NAA levels in the treated CD mice approach those in the normal mice. FIG. 30D indicates that APSA activity is detected in the brains of CD mice. FIG. 30E indicates APSA expression is detected in the brains of CD mice.
[0056] FIG. 31A depicts that vacuolation in both brain and spinal cord of the treated mice is more patchy and variable with generally smaller-sized vacuoles and that some areas of the cerebral cortex show almost no vacuolation. FIG. 31B shows ASPA expression in the cerebral cortex in situ.
[0057] FIGS. 32A to 32B depict results of a quantitative analysis of vacuolation in various brain regions. FIG. 32A shows that olfactory bulb had a dramatic mitigation in the white matter degeneration after gene therapy and that the large vacuoles were essentially eliminated in other tissues. FIG. 32B shows results from a similar analysis on spinal cord sections.
[0058] FIGS. 33A to 33D depict results from a histopathological evaluation of kidneys in the CD mice. FIG. 33A shows that the renal tubular epithelium of the kidney was diffusely attenuated and exhibited enlargement of the tubular lumens in untreated CD mice. FIG. 33B shows that treated CD mouse had normal glomeruli. FIGS. 33C and 33D depict results of an analysis of two lead candidate vectors, rAAV9 and rh.10, respectively, for efficiency of kidney transduction after IV delivery.
DETAILED DESCRIPTION OF CERTAIN EMBODIMENTS OF THE INVENTION
[0059] Adeno-associated virus (AAV) is a small (26 nm) replication-defective, nonenveloped virus, that depends on the presence of a second virus, such as adenovirus or herpes virus, for its growth in cells. AAV is not known to cause disease and induces a very mild immune response. AAV can infect both dividing and non-dividing cells and may incorporate its genome into that of the host cell. Aspects of the invention provide methods for delivering a transgene to a CNS tissue in a subject using recombinant AAV-based gene transfer. Accordingly, methods and compositions for treating CNS-related disorders are provided herein. Further aspects of the invention, are based on the discovery of rAAVs that achieve wide-spread distribution throughout CNS tissue. In some embodiments, the rAAVs spread throughout CNS tissue following direct administration into the cerebrospinal fluid (CSF), e.g., via intrathecal and/or intracerebral injection. In other embodiments, the rAAVs cross the blood-brain-barrier and achieve wide-spread distribution throughout CNS tissue of a subject following intravenous administration. Such rAAVs are useful for the treatment of CNS-related disorders, including, for example, amyotrophic lateral sclerosis (ALS) and Canavan disease (CD).
Methods and Compositions for Targeting CNS Tissue
[0060] Methods for delivering a transgene to central nervous system (CNS) tissue in a subject are provided herein. The methods typically involve administering to a subject an effective amount of a rAAV comprising a nucleic acid vector for expressing a transgene in the subject. An "effective amount" of a rAAV is an amount sufficient to infect a sufficient number of cells of a target tissue in a subject. An effective amount of a rAAV may be an amount sufficient to have a therapeutic benefit in a subject, e.g., to extend the lifespan of a subject, to improve in the subject one or more symptoms of disease, e.g., a symptom of ALS, a symptom of Canavan disease, etc. In some cases, an effective amount of a rAAV may be an amount sufficient to produce a stable somatic transgenic animal model. The effective amount will depend on a variety of factors such as, for example, the species, age, weight, health of the subject, and the CNS tissue to be targeted, and may thus vary among subject and tissue. An effective amount may also depend on the mode of administration. For example, targeting a CNS tissue by intravascular injection may require different (e.g., higher) doses, in some cases, than targeting CNS tissue by intrathecal or intracerebral injection. In some cases, multiple doses of a rAAV are administered. An effective amount may also depend on the rAAV used. For example, dosages for targeting a CNS tissue may depend on the serotype (e.g., the capsid protein) of the rAAV. For example, the rAAV may have a capsid protein of a AAV serotype selected from the group consisting of: AAV1, AAV2, AAV5, AAV6, AAV6.2, AAV7, AAV8, AAV9, rh.10, rh.39, rh.43 and CSp3. In certain embodiments, the effective amount of rAAV is 10.sup.10, 10.sup.11, 10.sup.12, 10.sup.13, or 10.sup.14 genome copies per kg. In certain embodiments, the effective amount of rAAV is 10.sup.10, 10.sup.11, 10.sup.12, 10.sup.13, 10.sup.14, or 10.sup.15 genome copies per subject.
[0061] A method for delivering a transgene to CNS tissue in a subject may comprise administering a rAAV by a single route or by multiple routes. For example, delivering a transgene to CNS tissue in a subject may comprise administering to the subject, by intravenous administration, an effective amount of a rAAV that crosses the blood-brain-barrier. Delivering a transgene to CNS tissue in a subject may comprise administering to the subject an effective amount of a rAAV by intrathecal administration or intracerebral administration, e.g., by intraventricular injection. A method for delivering a transgene to CNS tissue in a subject may comprise co-administering of an effective amount of a rAAV by two different administration routes, e.g., by intrathecal administration and by intracerebral administration. Co-administration may be performed at approximately the same time, or different times.
[0062] The CNS tissue to be targeted may be selected from cortex, hippocampus, thalamus, hypothalamus, cerebellum, brain stem, cervical spinal cord, thoracic spinal cord, and lumbar spinal cord, for example. The administration route for targeting CNS tissue typically depends on the AAV serotype. For example, in certain instances where the AAV serotype is selected from AAV1, AAV6, AAV6.2, AAV7, AAV8, AAV9, rh.10, rh.39, rh.43 and CSp3, the administration route may be intravascular injection. In some instances, for example where the AAV serotype is selected from AAV1, AAV2, AAV5, AAV6, AAV6.2, AAV7, AAV8, AAV9, rh.10, rh.39, rh.43 and CSp3, the administration route may be intrathecal and/or intracerebral injection.
[0063] Intravascular Administration
[0064] As used herein the term "intravascular administration" refers to the administration of an agent, e.g., a composition comprising a rAAV, into the vasculature of a subject, including the venous and arterial circulatory systems of the subject. Typically, rAAVs that cross the blood-brain-barrier may be delivered by intravascular administration for targeting CNS tissue. In some cases, intravascular (e.g., intravenous) administration facilitates the use of larger volumes than other forms of administration (e.g., intrathecal, intracerebral). Thus, large doses of rAAVs (e.g., up to 10.sup.15 GC/subject) can be delivered at one time by intravascular (e.g., intravenous) administration. Methods for intravascular administration are well known in the art and include for example, use of a hypodermic needle, peripheral cannula, central venous line, etc.
[0065] Intrathecal and/or Intracerebral Administration
[0066] As used herein the term "intrathecal administration" refers to the administration of an agent, e.g., a composition comprising a rAAV, into the spinal canal. For example, intrathecal administration may comprise injection in the cervical region of the spinal canal, in the thoracic region of the spinal canal, or in the lumbar region of the spinal canal. Typically, intrathecal administration is performed by injecting an agent, e.g., a composition comprising a rAAV, into the subarachnoid cavity (subarachnoid space) of the spinal canal, which is the region between the arachnoid membrane and pia mater of the spinal canal. The subarchnoid space is occupied by spongy tissue consisting of trabeculae (delicate connective tissue filaments that extend from the arachnoid mater and blend into the pia mater) and intercommunicating channels in which the cerebrospinal fluid is contained. In some embodiments, intrathecal administration is not administration into the spinal vasculature.
[0067] As used herein, the term "intracerebral administration" refers to administration of an agent into and/or around the brain. Intracerebral administration includes, but is not limited to, administration of an agent into the cerebrum, medulla, pons, cerebellum, intracranial cavity, and meninges surrounding the brain. Intracerebral administration may include administration into the dura mater, arachnoid mater, and pia mater of the brain. Intracerebral administration may include, in some embodiments, administration of an agent into the cerebrospinal fluid (CSF) of the subarachnoid space surrounding the brain. Intracerebral administration may include, in some embodiments, administration of an agent into ventricles of the brain, e.g., the right lateral ventricle, the left lateral ventricle, the third ventricle, the fourth ventricle. In some embodiments, intracerebral administration is not administration into the brain vasculature.
[0068] Intracerebral administration may involve direct injection into and/or around the brain. In some embodiments, intracerebral administration involves injection using stereotaxic procedures. Stereotaxic procedures are well know in the art and typically involve the use of a computer and a 3-dimensional scanning device that are used together to guide injection to a particular intracerebral region, e.g., a ventricular region. Micro-injection pumps (e.g., from World Precision Instruments) may also be used. In some embodiments, a microinjection pump is used to deliver a composition comprising a rAAV. In some embodiments, the infusion rate of the composition is in a range of 1 .mu.l/minute to 100 .mu.l/minute. As will be appreciated by the skilled artisan, infusion rates will depend on a variety of factors, including, for example, species of the subject, age of the subject, weight/size of the subject, serotype of the AAV, dosage required, intracerebral region targeted, etc. Thus, other infusion rates may be deemed by a skilled artisan to be appropriate in certain circumstances.
Methods and Compositions for Treating CNS-Related Disorders
[0069] Methods and compositions for treating CNS-related disorders are also provided herein. As used herein, a "CNS-related disorder" is a disease or condition of the central nervous system. A CNS-related disorder may affect the spinal cord (e.g., a myelopathy), brain (e.g., a encephalopathy) or tissues surrounding the brain and spinal cord. A CNS-related disorder may be of a genetic origin, either inherited or acquired through a somatic mutation. A CNS-related disorder may be a psychological condition or disorder, e.g., Attention Deficient Hyperactivity Disorder, Autism Spectrum Disorder, Mood Disorder, Schizophrenia, Depression, Rett Syndrome, etc. A CNS-related disorder may be an autoimmune disorder. A CNS-related disorder may also be a cancer of the CNS, e.g., brain cancer. A CNS-related disorder that is a cancer may be a primary cancer of the CNS, e.g., an astrocytoma, glioblastomas, etc., or may be a cancer that has metastasized to CNS tissue, e.g., a lung cancer that has metastasized to the brain. Further non-limiting examples of CNS-related disorders, include Parkinson's Disease, Lysosomal Storage Disease, Ischemia, Neuropathic Pain, Amyotrophic lateral sclerosis (ALS), Multiple Sclerosis (MS), and Canavan disease (CD).
[0070] Methods for treating amyotrophic lateral sclerosis (ALS) in a subject in need thereof are provided herein. A subject in need of a treatment for ALS is a subject having or suspected of having ALS. In some cases, ALS has been linked to a mutation in the gene coding for superoxide dismutase (SOD1). Elevated levels of SOD1 appear to be associated with ALS in some instances. It has been shown that transgenic expression of shRNA against SOD1 can knockdown mutant SOD1 expression, delay disease onset and extend survival (Xia et al. 2006, Neurobiol Dis 23: 578). Intrathecal infusion of siRNA against SOD1 at disease onset has also been found to knockdown mutant SOD1 expression and extend survival (Wang et al. 2008, JBC 283: 15845). Furthermore, nerve injection of adenovirus expressing shRNA against SOD1 at the disease onset can knockdown mutant SOD1 expression and extend survival (Wu et al. 2009, Antiox Redox Sig 11: 1523).
[0071] Aspects of the invention, are based on the discovery of AAV-based therapies that achieve, with low-toxicity, long-term inhibition of SOD1 expression that is wide-spread throughout CNS tissue of the subject. Methods for treating ALS that are provided herein, typically involve administering to CNS tissue of a subject an effective amount of a rAAV that harbors a nucleic acid comprising a promoter operably linked with a region encoding an inhibitory RNA that binds specifically to SOD1 mRNA (e.g., that hybridizes specifically to a nucleic acid having a sequence as set forth in SEQ ID NO 17 or 19) and inhibits expression of SOD1 in the subject. It has been discovered that rAAVs having a capsid protein comprising a sequence as set forth in SEQ ID NO: 9 achieve wide-spread distribution throughout the CNS following intrathecal injection and/or intracerebral injection, and thus, are particularly useful for treating ALS. This result is surprising in light of certain other rAAVs that infect cells only within the immediate vicinity of the injection site, or the achieve only a limited distribution, following intrathecal injection. Thus, rAAVs that achieve wide-spread distribution throughout the CNS are particularly useful as gene transfer vectors for treating ALS.
[0072] In some embodiments, it has been discovered that co-administration by intrathecal injection and intracerebral injection, e.g., intraventricular injection, of rAAVs having a capsid protein comprising a sequence as set forth in SEQ ID NO: 9 and a nucleic acid comprising a promoter operably linked with a region encoding an inhibitory RNA that binds specifically to SOD1 mRNA and inhibits expression of SOD1, achieves long-term inhibition of SOD1 and improves outcome (e.g., lifespan) in an animal model of ALS (See, e.g., FIG. 6A). In some embodiments, the inhibitory RNA is an antisense RNA, a shRNA or a miRNA. The inhibitory RNA may have a sequence as set forth in SEQ ID NO: 26. The inhibitory RNA may have a sequence as set forth in any one of SEQ ID NO: 22 to 30. Thus, in some embodiments, a nucleic acid comprising a promoter operably linked with a nucleic acid having a sequence as set forth in any one of SEQ ID NO: 22 to 30 is provided. In some embodiments, a recombinant AAV that harbors a nucleic acid comprising a sequence as set forth in any one of SEQ ID NO: 22 to 30 is provided. The recombinant AAV may have a capsid protein comprising a sequence as set forth in SEQ ID NO: 9. The recombinant AAV may have a capsid protein comprising a sequence as set forth in any one of SEQ ID NO: 1 to 12.
[0073] Methods for treating Canavan disease (CD) in a subject in need thereof are provided herein. A subject in need of a treatment for CD is a subject having or suspected of having CD. Canavan disease is caused by a defective ASPA gene which is responsible for the production of the enzyme aspartoacylase. This enzyme normally breaks down the concentrated brain molecule N-acetyl aspartate. Decreased aspartoacylase activity in subjects with CD prevents the normal breakdown of N-acetyl aspartate, and the lack of breakdown appears to interfere with growth of the myelin sheath of the nerve fibers in the brain. Symptoms of Canavan disease, which may appear in early infancy and progress rapidly, may include mental retardation, loss of previously acquired motor skills, feeding difficulties, abnormal muscle tone (i.e., floppiness or stiffness), poor head control, and megalocephaly (abnormally enlarged head). Paralysis, blindness, or seizures may also occur. Aspects of the invention improve one or more symptoms of CD in a subject by administering to the subject a recombinant AAV harboring a nucleic acid that expresses aspartoacylase (ASPA). For example, a method for treating Canavan disease in a subject in need thereof may comprise administering an effective amount of a rAAV to CNS tissue of the subject by intravascular administration, wherein the rAAV comprises a nucleic acid comprising a promoter operably linked with a region encoding ASPA (e.g., a region having a sequence as set forth in SEQ ID NO: 14 or 16). A method for treating Canavan disease in a subject in need thereof may comprise administering an effective amount of a rAAV to CNS tissue of the subject by intrathecal administration, wherein the rAAV comprises a nucleic acid comprising a promoter operably linked with a region encoding ASPA. In some cases, methods for treating CD involve administering, to CNS tissue of the subject, an effective amount of a rAAV that comprises a capsid protein other than a capsid protein of AAV serotype 2 (e.g., other than a protein having an amino acid sequence as set forth in SEQ ID NO: 2) and a nucleic acid comprising a promoter operably linked with a region encoding ASPA. In another example, a method for treating Canavan disease in a subject in need thereof comprises administering an effective amount of a rAAV to CNS tissue of the subject by a route other than intracerebral administration, wherein the rAAV comprises a nucleic acid comprising a promoter operably linked with a region encoding ASPA. In some embodiments, ASPA expressed in CNS tissue following administration of the rAAV results in a decrease in aspartoacylase activity and breakdown of N-acetyl aspartate in the CNS tissue. Thus, in some embodiments, a recombinant AAV vector is provided that comprises a nucleic acid encoding a sequence as set forth in SEQ ID NO: 14 or 16. In some embodiments, a recombinant AAV is provided that harbors a nucleic acid comprising a promoter operably linked with a region having a sequence as set forth in SEQ ID NO: 14 or 16. In some embodiments, a recombinant AAV is provided that harbors a nucleic acid comprising a promoter operably linked with a region encoding a protein having a sequence as set forth in SEQ ID NO: 13 or 15. The recombinant AAV may have a capsid protein comprising an amino acid sequence as set forth in any one of SEQ ID NO: 1 to 12. The recombinant AAV may have a capsid protein comprising a sequence as set forth in any one of SEQ ID NO: 1 and 3 to 12.
Recombinant AAVs
[0074] In some aspects, the invention provides isolated AAVs. As used herein with respect to AAVs, the term "isolated" refers to an AAV that has been isolated from its natural environment (e.g., from a host cell, tissue, or subject) or artificially produced. Isolated AAVs may be produced using recombinant methods. Such AAVs are referred to herein as "recombinant AAVs". Recombinant AAVs (rAAVs) preferably have tissue-specific targeting capabilities, such that a transgene of the rAAV will be delivered specifically to one or more predetermined tissue(s). The AAV capsid is an important element in determining these tissue-specific targeting capabilities. Thus, a rAAV having a capsid appropriate for the tissue being targeted can be selected. In some embodiments, the rAAV comprises a capsid protein having an amino acid sequence as set forth in any one of SEQ ID NOs 1 to 12, or a protein having substantial homology thereto.
[0075] Methods for obtaining recombinant AAVs having a desired capsid protein are well known in the art (See, for example, US 2003/0138772, the contents of which are incorporated herein by reference in their entirety). AAVs capsid protein that may be used in the rAAVs of the invention a include, for example, those disclosed in G. Gao, et al., J. Virol, 78(12):6381-6388 (June 2004); G. Gao, et al, Proc Natl Acad Sci USA, 100(10):6081-6086 (May 13, 2003); US 2003-0138772, US 2007/0036760, US 2009/0197338, and U.S. provisional application Ser. No. 61/182,084, filed May 28, 2009, the contents of which relating to AAVs capsid proteins and associated nucleotide and amino acid sequences are incorporated herein by reference. Typically the methods involve culturing a host cell which contains a nucleic acid sequence encoding an AAV capsid protein (e.g., a nucleic acid encoding a protein having a sequence as set forth in any one of SEQ ID NOs 1-12) or fragment thereof; a functional rep gene; a recombinant AAV vector composed of, AAV inverted terminal repeats (ITRs) and a transgene; and sufficient helper functions to permit packaging of the recombinant AAV vector into the AAV capsid proteins.
[0076] The components to be cultured in the host cell to package a rAAV vector in an AAV capsid may be provided to the host cell in trans. Alternatively, any one or more of the required components (e.g., recombinant AAV vector, rep sequences, cap sequences, and/or helper functions) may be provided by a stable host cell which has been engineered to contain one or more of the required components using methods known to those of skill in the art. Most suitably, such a stable host cell will contain the required component(s) under the control of an inducible promoter. However, the required component(s) may be under the control of a constitutive promoter. Examples of suitable inducible and constitutive promoters are provided herein, in the discussion of regulatory elements suitable for use with the transgene. In still another alternative, a selected stable host cell may contain selected component(s) under the control of a constitutive promoter and other selected component(s) under the control of one or more inducible promoters. For example, a stable host cell may be generated which is derived from 293 cells (which contain E1 helper functions under the control of a constitutive promoter), but which contain the rep and/or cap proteins under the control of inducible promoters. Still other stable host cells may be generated by one of skill in the art.
[0077] The recombinant AAV vector, rep sequences, cap sequences, and helper functions required for producing the rAAV of the invention may be delivered to the packaging host cell using any appropriate genetic element (vector). The selected genetic element may be delivered by any suitable method, including those described herein. The methods used to construct any embodiment of this invention are known to those with skill in nucleic acid manipulation and include genetic engineering, recombinant engineering, and synthetic techniques. See, e.g., Sambrook et al, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Press, Cold Spring Harbor, N.Y. Similarly, methods of generating rAAV virions are well known and the selection of a suitable method is not a limitation on the present invention. See, e.g., K. Fisher et al, J. Virol., 70:520-532 (1993) and U.S. Pat. No. 5,478,745.
[0078] In some embodiments, recombinant AAVs may be produced using the triple transfection method (e.g., as described in detail in U.S. Pat. No. 6,001,650, the contents of which relating to the triple transfection method are incorporated herein by reference). Typically, the recombinant AAVs are produced by transfecting a host cell with a recombinant AAV vector (comprising a transgene) to be packaged into AAV particles, an AAV helper function vector, and an accessory function vector. An AAV helper function vector encodes the "AAV helper function" sequences (i.e., rep and cap), which function in trans for productive AAV replication and encapsidation. Preferably, the AAV helper function vector supports efficient AAV vector production without generating any detectable wild-type AAV virions (i.e., AAV virions containing functional rep and cap genes). Non-limiting examples of vectors suitable for use with the present invention include pHLP19, described in U.S. Pat. No. 6,001,650 and pRep6cap6 vector, described in U.S. Pat. No. 6,156,303, the entirety of both incorporated by reference herein. The accessory function vector encodes nucleotide sequences for non-AAV derived viral and/or cellular functions upon which AAV is dependent for replication (i.e., "accessory functions"). The accessory functions include those functions required for AAV replication, including, without limitation, those moieties involved in activation of AAV gene transcription, stage specific AAV mRNA splicing, AAV DNA replication, synthesis of cap expression products, and AAV capsid assembly. Viral-based accessory functions can be derived from any of the known helper viruses such as adenovirus, herpesvirus (other than herpes simplex virus type-1), and vaccinia virus.
[0079] In some aspects, the invention provides transfected host cells. The term "transfection" is used to refer to the uptake of foreign DNA by a cell, and a cell has been "transfected" when exogenous DNA has been introduced inside the cell membrane. A number of transfection techniques are generally known in the art. See, e.g., Graham et al. (1973) Virology, 52:456, Sambrook et al. (1989) Molecular Cloning, a laboratory manual, Cold Spring Harbor Laboratories, New York, Davis et al. (1986) Basic Methods in Molecular Biology, Elsevier, and Chu et al. (1981) Gene 13:197. Such techniques can be used to introduce one or more exogenous nucleic acids, such as a nucleotide integration vector and other nucleic acid molecules, into suitable host cells.
[0080] A "host cell" refers to any cell that harbors, or is capable of harboring, a substance of interest. Often a host cell is a mammalian cell. A host cell may be used as a recipient of an AAV helper construct, an AAV minigene plasmid, an accessory function vector, or other transfer DNA associated with the production of recombinant AAVs. The term includes the progeny of the original cell which has been transfected. Thus, a "host cell" as used herein may refer to a cell which has been transfected with an exogenous DNA sequence. It is understood that the progeny of a single parental cell may not necessarily be completely identical in morphology or in genomic or total DNA complement as the original parent, due to natural, accidental, or deliberate mutation.
[0081] In some aspects, the invention provides isolated cells. As used herein with respect to cell, the term "isolated" refers to a cell that has been isolated from its natural environment (e.g., from a tissue or subject). As used herein, the term "cell line" refers to a population of cells capable of continuous or prolonged growth and division in vitro. Often, cell lines are clonal populations derived from a single progenitor cell. It is further known in the art that spontaneous or induced changes can occur in karyotype during storage or transfer of such clonal populations. Therefore, cells derived from the cell line referred to may not be precisely identical to the ancestral cells or cultures, and the cell line referred to includes such variants. As used herein, the terms "recombinant cell" refers to a cell into which an exogenous DNA segment, such as DNA segment that leads to the transcription of a biologically-active polypeptide or production of a biologically active nucleic acid such as an RNA, has been introduced.
[0082] As used herein, the term "vector" includes any genetic element, such as a plasmid, phage, transposon, cosmid, chromosome, artificial chromosome, virus, virion, etc., which is capable of replication when associated with the proper control elements and which can transfer gene sequences between cells. Thus, the term includes cloning and expression vehicles, as well as viral vectors. In some embodiments, useful vectors are contemplated to be those vectors in which the nucleic acid segment to be transcribed is positioned under the transcriptional control of a promoter. A "promoter" refers to a DNA sequence recognized by the synthetic machinery of the cell, or introduced synthetic machinery, required to initiate the specific transcription of a gene. The phrases "operatively positioned," "under control" or "under transcriptional control" means that the promoter is in the correct location and orientation in relation to the nucleic acid to control RNA polymerase initiation and expression of the gene. The term "expression vector or construct" means any type of genetic construct containing a nucleic acid in which part or all of the nucleic acid encoding sequence is capable of being transcribed. In some embodiments, expression includes transcription of the nucleic acid, for example, to generate a biologically-active polypeptide product or inhibitory RNA (e.g., shRNA, miRNA) from a transcribed gene.
[0083] The foregoing methods for packaging recombinant vectors in desired AAV capsids to produce the rAAVs of the invention are not meant to be limiting and other suitable methods will be apparent to the skilled artisan.
Recombinant AAV Vectors
[0084] "Recombinant AAV (rAAV) vectors" of the invention are typically composed of, at a minimum, a transgene and its regulatory sequences, and 5' and 3' AAV inverted terminal repeats (ITRs). It is this recombinant AAV vector which is packaged into a capsid protein and delivered to a selected target cell. In some embodiments, the transgene is a nucleic acid sequence, heterologous to the vector sequences, which encodes a polypeptide, protein, functional RNA molecule (e.g., miRNA, miRNA inhibitor) or other gene product, of interest. The nucleic acid coding sequence is operatively linked to regulatory components in a manner which permits transgene transcription, translation, and/or expression in a cell of a target tissue.
[0085] The AAV sequences of the vector typically comprise the cis-acting 5' and 3' inverted terminal repeat sequences (See, e.g., B. J. Carter, in "Handbook of Parvoviruses", ed., P. Tijsser, CRC Press, pp. 155 168 (1990)). The ITR sequences are about 145 bp in length. Preferably, substantially the entire sequences encoding the ITRs are used in the molecule, although some degree of minor modification of these sequences is permissible. The ability to modify these ITR sequences is within the skill of the art. (See, e.g., texts such as Sambrook et al, "Molecular Cloning. A Laboratory Manual", 2d ed., Cold Spring Harbor Laboratory, New York (1989); and K. Fisher et al., J Virol., 70:520 532 (1996)). An example of such a molecule employed in the present invention is a "cis-acting" plasmid containing the transgene, in which the selected transgene sequence and associated regulatory elements are flanked by the 5' and 3' AAV ITR sequences. The AAV ITR sequences may be obtained from any known AAV, including presently identified mammalian AAV types.
[0086] In addition to the major elements identified above for the recombinant AAV vector, the vector also includes conventional control elements which are operably linked to the transgene in a manner which permits its transcription, translation and/or expression in a cell transfected with the plasmid vector or infected with the virus produced by the invention. As used herein, "operably linked" sequences include both expression control sequences that are contiguous with the gene of interest and expression control sequences that act in trans or at a distance to control the gene of interest. Expression control sequences include appropriate transcription initiation, termination, promoter and enhancer sequences; efficient RNA processing signals such as splicing and polyadenylation (polyA) signals; sequences that stabilize cytoplasmic mRNA; sequences that enhance translation efficiency (i.e., Kozak consensus sequence); sequences that enhance protein stability; and when desired, sequences that enhance secretion of the encoded product. A great number of expression control sequences, including promoters which are native, constitutive, inducible and/or tissue-specific, are known in the art and may be utilized.
[0087] As used herein, a nucleic acid sequence (e.g., coding sequence) and regulatory sequences are said to be operably linked when they are covalently linked in such a way as to place the expression or transcription of the nucleic acid sequence under the influence or control of the regulatory sequences. If it is desired that the nucleic acid sequences be translated into a functional protein, two DNA sequences are said to be operably linked if induction of a promoter in the 5' regulatory sequences results in the transcription of the coding sequence and if the nature of the linkage between the two DNA sequences does not (1) result in the introduction of a frame-shift mutation, (2) interfere with the ability of the promoter region to direct the transcription of the coding sequences, or (3) interfere with the ability of the corresponding RNA transcript to be translated into a protein. Thus, a promoter region would be operably linked to a nucleic acid sequence if the promoter region were capable of effecting transcription of that DNA sequence such that the resulting transcript might be translated into the desired protein or polypeptide. Similarly two or more coding regions are operably linked when they are linked in such a way that their transcription from a common promoter results in the expression of two or more proteins having been translated in frame. In some embodiments, operably linked coding sequences yield a fusion protein. In some embodiments, operably linked coding sequences yield a functional RNA (e.g., shRNA, miRNA).
[0088] For nucleic acids encoding proteins, a polyadenylation sequence generally is inserted following the transgene sequences and before the 3' AAV ITR sequence. A rAAV construct useful in the present invention may also contain an intron, desirably located between the promoter/enhancer sequence and the transgene. One possible intron sequence is derived from SV-40, and is referred to as the SV-40 T intron sequence. Another vector element that may be used is an internal ribosome entry site (IRES). An IRES sequence is used to produce more than one polypeptide from a single gene transcript. An IRES sequence would be used to produce a protein that contain more than one polypeptide chains. Selection of these and other common vector elements are conventional and many such sequences are available [see, e.g., Sambrook et al, and references cited therein at, for example, pages 3.18 3.26 and 16.17 16.27 and Ausubel et al., Current Protocols in Molecular Biology, John Wiley & Sons, New York, 1989]. In some embodiments, a Foot and Mouth Disease Virus 2A sequence is included in polyprotein; this is a small peptide (approximately 18 amino acids in length) that has been shown to mediate the cleavage of polyproteins (Ryan, M D et al., EMBO, 1994; 4: 928-933; Mattion, N M et al., J Virology, November 1996; p. 8124-8127; Furler, S et al., Gene Therapy, 2001; 8: 864-873; and Halpin, C et al., The Plant Journal, 1999; 4: 453-459). The cleavage activity of the 2A sequence has previously been demonstrated in artificial systems including plasmids and gene therapy vectors (AAV and retroviruses) (Ryan, M D et al., EMBO, 1994; 4: 928-933; Mattion, N M et al., J Virology, November 1996; p. 8124-8127; Furler, S et al., Gene Therapy, 2001; 8: 864-873; and Halpin, C et al., The Plant Journal, 1999; 4: 453-459; de Felipe, P et al., Gene Therapy, 1999; 6: 198-208; de Felipe, P et al., Human Gene Therapy, 2000; 11: 1921-1931.; and Klump, H et al., Gene Therapy, 2001; 8: 811-817).
[0089] The precise nature of the regulatory sequences needed for gene expression in host cells may vary between species, tissues or cell types, but shall in general include, as necessary, 5' non-transcribed and 5' non-translated sequences involved with the initiation of transcription and translation respectively, such as a TATA box, capping sequence, CAAT sequence, enhancer elements, and the like. Especially, such 5' non-transcribed regulatory sequences will include a promoter region that includes a promoter sequence for transcriptional control of the operably joined gene. Regulatory sequences may also include enhancer sequences or upstream activator sequences as desired. The vectors of the invention may optionally include 5' leader or signal sequences. The choice and design of an appropriate vector is within the ability and discretion of one of ordinary skill in the art.
[0090] Examples of constitutive promoters include, without limitation, the retroviral Rous sarcoma virus (RSV) LTR promoter (optionally with the RSV enhancer), the cytomegalovirus (CMV) promoter (optionally with the CMV enhancer) [see, e.g., Boshart et al, Cell, 41:521-530 (1985)], the SV40 promoter, the dihydrofolate reductase promoter, the .beta.-actin promoter, the phosphoglycerol kinase (PGK) promoter, and the EF1.alpha. promoter [Invitrogen].
[0091] Inducible promoters allow regulation of gene expression and can be regulated by exogenously supplied compounds, environmental factors such as temperature, or the presence of a specific physiological state, e.g., acute phase, a particular differentiation state of the cell, or in replicating cells only. Inducible promoters and inducible systems are available from a variety of commercial sources, including, without limitation, Invitrogen, Clontech and Ariad. Many other systems have been described and can be readily selected by one of skill in the art. Examples of inducible promoters regulated by exogenously supplied promoters include the zinc-inducible sheep metallothionine (MT) promoter, the dexamethasone (Dex)-inducible mouse mammary tumor virus (MMTV) promoter, the T7 polymerase promoter system (WO 98/10088); the ecdysone insect promoter (No et al, Proc. Natl. Acad. Sci. USA, 93:3346-3351 (1996)), the tetracycline-repressible system (Gossen et al, Proc. Natl. Acad. Sci. USA, 89:5547-5551 (1992)), the tetracycline-inducible system (Gossen et al, Science, 268:1766-1769 (1995), see also Harvey et al, Curr. Opin. Chem. Biol., 2:512-518 (1998)), the RU486-inducible system (Wang et al, Nat. Biotech., 15:239-243 (1997) and Wang et al, Gene Ther., 4:432-441 (1997)) and the rapamycin-inducible system (Magari et al, J. Clin. Invest., 100:2865-2872 (1997)). Still other types of inducible promoters which may be useful in this context are those which are regulated by a specific physiological state, e.g., temperature, acute phase, a particular differentiation state of the cell, or in replicating cells only.
[0092] In another embodiment, the native promoter, or fragment thereof, for the transgene will be used. The native promoter may be preferred when it is desired that expression of the transgene should mimic the native expression. The native promoter may be used when expression of the transgene must be regulated temporally or developmentally, or in a tissue-specific manner, or in response to specific transcriptional stimuli. In a further embodiment, other native expression control elements, such as enhancer elements, polyadenylation sites or Kozak consensus sequences may also be used to mimic the native expression.
[0093] In some embodiments, the regulatory sequences impart tissue-specific gene expression capabilities. In some cases, the tissue-specific regulatory sequences bind tissue-specific transcription factors that induce transcription in a tissue specific manner. Such tissue-specific regulatory sequences (e.g., promoters, enhancers, etc.) are well known in the art. Exemplary tissue-specific regulatory sequences include, but are not limited to the following tissue specific promoters: neuronal such as neuron-specific enolase (NSE) promoter (Andersen et al., Cell. Mol. Neurobiol., 13:503-15 (1993)), neurofilament light-chain gene promoter (Piccioli et al., Proc. Natl. Acad. Sci. USA, 88:5611-5 (1991)), and the neuron-specific vgf gene promoter (Piccioli et al., Neuron, 15:373-84 (1995)). In some embodiments, the tissue-specific promoter is a promoter of a gene selected from: neuronal nuclei (NeuN), glial fibrillary acidic protein (GFAP), adenomatous polyposis coli (APC), and ionized calcium-binding adapter molecule 1 (Iba-1). Other appropriate tissue specific promoters will be apparent to the skilled artisan. In some embodiments, the promoter is a chicken Beta-actin promoter.
[0094] In some embodiments, one or more bindings sites for one or more of miRNAs are incorporated in a transgene of a rAAV vector, to inhibit the expression of the transgene in one or more tissues of a subject harboring the transgenes, e.g., non-CNS tissues. The skilled artisan will appreciate that binding sites may be selected to control the expression of a transgene in a tissue specific manner. For example, expression of a transgene in the liver may be inhibited by incorporating a binding site for miR-122 such that mRNA expressed from the transgene binds to and is inhibited by miR-122 in the liver. Expression of a transgene in the heart may be inhibited by incorporating a binding site for miR-133a or miR-1, such that mRNA expressed from the transgene binds to and is inhibited by miR-133a or miR-1 in the heart. The miRNA target sites in the mRNA may be in the 5' UTR, the 3' UTR or in the coding region. Typically, the target site is in the 3' UTR of the mRNA. Furthermore, the transgene may be designed such that multiple miRNAs regulate the mRNA by recognizing the same or multiple sites. The presence of multiple miRNA binding sites may result in the cooperative action of multiple RISCs and provide highly efficient inhibition of expression. The target site sequence may comprise a total of 5-100, 10-60, or more nucleotides. The target site sequence may comprise at least 5 nucleotides of the sequence of a target gene binding site.
Transgene Coding Sequences: CNS-Related Genes
[0095] The composition of the transgene sequence of a rAAV vector will depend upon the use to which the resulting vector will be put. For example, one type of transgene sequence includes a reporter sequence, which upon expression produces a detectable signal. In another example, the transgene encodes a therapeutic protein or therapeutic functional RNA. In another example, the transgene encodes a protein or functional RNA that is intended to be used for research purposes, e.g., to create a somatic transgenic animal model harboring the transgene, e.g., to study the function of the transgene product. In another example, the transgene encodes a protein or functional RNA that is intended to be used to create an animal model of disease. Appropriate transgene coding sequences will be apparent to the skilled artisan.
[0096] In some aspects, the invention provides rAAV vectors for use in methods of preventing or treating one or more gene defects (e.g., heritable gene defects, somatic gene alterations) in a mammal, such as for example, a gene defect that results in a polypeptide deficiency or polypeptide excess in a subject, and particularly for treating or reducing the severity or extent of deficiency in a subject manifesting a CNS-associated disorder linked to a deficiency in such polypeptides in cells and tissues. In some embodiments, methods involve administration of a rAAV vector that encodes one or more therapeutic peptides, polypeptides, shRNAs, microRNAs, antisense nucleotides, etc. in a pharmaceutically-acceptable carrier to the subject in an amount and for a period of time sufficient to treat the CNS-associated disorder in the subject having or suspected of having such a disorder.
[0097] A rAAV vector may comprise as a transgene, a nucleic acid encoding a protein or functional RNA that modulates or treats a CNS-associated disorder. The following is a non-limiting list of genes associated with CNS-associated disorders: neuronal apoptosis inhibitory protein (NAIP), nerve growth factor (NGF), glial-derived growth factor (GDNF), brain-derived growth factor (BDNF), ciliary neurotrophic factor (CNTF), tyrosine hydroxlase (TH), GTP-cyclohydrolase (GTPCH), aspartoacylase (ASPA), superoxide dismutase (SOD1) and amino acid decorboxylase (AADC). For example, a useful transgene in the treatment of Parkinson's disease encodes TH, which is a rate limiting enzyme in the synthesis of dopamine. A transgene encoding GTPCH, which generates the TH cofactor tetrahydrobiopterin, may also be used in the treatment of Parkinson's disease. A transgene encoding GDNF or BDNF, or AADC, which facilitates conversion of L-Dopa to DA, may also be used for the treatment of Parkinson's disease. For the treatment of ALS, a useful transgene may encode: GDNF, BDNF or CNTF. Also for the treatment of ALS, a useful transgene may encode a functional RNA, e.g., shRNA, miRNA, that inhibits the expression of SOD1. For the treatment of ischemia a useful transgene may encode NAIP or NGF. A transgene encoding Beta-glucuronidase (GUS) may be useful for the treatment of certain lysosomal storage diseases (e.g., Mucopolysacharidosis type VII (MPS VII)). A transgene encoding a prodrug activation gene, e.g., HSV-Thymidine kinase which converts ganciclovir to a toxic nucleotide which disrupts DNA synthesis and leads to cell death, may be useful for treating certain cancers, e.g., when administered in combination with the prodrug. A transgene encoding an endogenous opioid, such a .beta.-endorphin may be useful for treating pain. Other examples of transgenes that may be used in the rAAV vectors of the invention will be apparent to the skilled artisan (See, e.g., Costantini L C, et al., Gene Therapy (2000) 7, 93-109).
[0098] In some embodiments, the cloning capacity of the recombinant RNA vector may be limited and a desired coding sequence may involve the complete replacement of the virus's 4.8 kilobase genome. Large genes may, therefore, not be suitable for use in a standard recombinant AAV vector, in some cases. The skilled artisan will appreciate that options are available in the art for overcoming a limited coding capacity. For example, the AAV ITRs of two genomes can anneal to form head to tail concatamers, almost doubling the capacity of the vector. Insertion of splice sites allows for the removal of the ITRs from the transcript. Other options for overcoming a limited cloning capacity will be apparent to the skilled artisan.
Recombinant AAV Administration
[0099] rAAVS are administered in sufficient amounts to transfect the cells of a desired tissue and to provide sufficient levels of gene transfer and expression without undue adverse effects. Conventional and pharmaceutically acceptable routes of administration include, but are not limited to, direct delivery to the selected tissue (e.g., intracerebral administration, intrathecal administration), intravenous, oral, inhalation (including intranasal and intratracheal delivery), intraocular, intravenous, intramuscular, subcutaneous, intradermal, intratumoral, and other parental routes of administration. Routes of administration may be combined, if desired.
[0100] Delivery of certain rAAVs to a subject may be, for example, by administration into the bloodstream of the subject. Administration into the bloodstream may be by injection into a vein, an artery, or any other vascular conduit. Moreover, in certain instances, it may be desirable to deliver the rAAVs to brain tissue, meninges, neuronal cells, glial cells, astrocytes, oligodendrocytes, cereobrospinal fluid (CSF), interstitial spaces and the like. In some embodiments, recombinant AAVs may be delivered directly to the spinal cord or brain by injection into the ventricular region, as well as to the striatum (e.g., the caudate nucleus or putamen of the striatum), and neuromuscular junction, or cerebellar lobule, with a needle, catheter or related device, using neurosurgical techniques known in the art, such as by stereotactic injection (see, e.g., Stein et al., J Virol 73:3424-3429, 1999; Davidson et al., PNAS 97:3428-3432, 2000; Davidson et al., Nat. Genet. 3:219-223, 1993; and Alisky and Davidson, Hum. Gene Ther. 11:2315-2329, 2000). In certain circumstances it will be desirable to deliver the rAAV-based therapeutic constructs in suitably formulated pharmaceutical compositions disclosed herein either subcutaneously, intrapancreatically, intranasally, parenterally, intravenously, intramuscularly, intracerebrally, intrathecally, intracerebrally, orally, intraperitoneally, or by inhalation. In some embodiments, the administration modalities as described in U.S. Pat. Nos. 5,543,158; 5,641,515 and 5,399,363 (each specifically incorporated herein by reference in its entirety) may be used to deliver rAAVs.
Recombinant AAV Compositions
[0101] The rAAVs may be delivered to a subject in compositions according to any appropriate methods known in the art. The rAAV, preferably suspended in a physiologically compatible carrier (e.g., in a composition), may be administered to a subject, e.g., a human, mouse, rat, cat, dog, sheep, rabbit, horse, cow, goat, pig, guinea pig, hamster, chicken, turkey, or a non-human primate (e.g, Macaque). The compositions of the invention may comprise a rAAV alone, or in combination with one or more other viruses (e.g., a second rAAV encoding having one or more different transgenes). In some embodiments, a compositions comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more different rAAVs each having one or more different transgenes.
[0102] Suitable carriers may be readily selected by one of skill in the art in view of the indication for which the rAAV is directed. For example, one suitable carrier includes saline, which may be formulated with a variety of buffering solutions (e.g., phosphate buffered saline). Other exemplary carriers include sterile saline, lactose, sucrose, calcium phosphate, gelatin, dextran, agar, pectin, peanut oil, sesame oil, and water. The selection of the carrier is not a limitation of the present invention.
[0103] Optionally, the compositions of the invention may contain, in addition to the rAAV and carrier(s), other conventional pharmaceutical ingredients, such as preservatives, or chemical stabilizers. Suitable exemplary preservatives include chlorobutanol, potassium sorbate, sorbic acid, sulfur dioxide, propyl gallate, the parabens, ethyl vanillin, glycerin, phenol, and parachlorophenol. Suitable chemical stabilizers include gelatin and albumin.
[0104] The dose of rAAV virions required to achieve a desired effect or "therapeutic effect," e.g., the units of dose in vector genomes/per kilogram of body weight (vg/kg), will vary based on several factors including, but not limited to: the route of rAAV administration, the level of gene or RNA expression required to achieve a therapeutic effect, the specific disease or disorder being treated, and the stability of the gene or RNA product. One of skill in the art can readily determine a rAAV virion dose range to treat a subject having a particular disease or disorder based on the aforementioned factors, as well as other factors that are well known in the art. An effective amount of the rAAV is generally in the range of from about 10 .mu.l to about 100 ml of solution containing from about 10.sup.9 to 10.sup.16 genome copies per subject. Other volumes of solution may be used. The volume used will typically depend, among other things, on the size of the subject, the dose of the rAAV, and the route of administration. For example, for intrathecal or intracerebral administration a volume in range of 1 .mu.l to 10 .mu.l or 10 .mu.l to 100 .mu.l may be used. For intravenous administration a volume in range of 10 .mu.l to 100 .mu.l, 100 .mu.l to 1 ml, 1 ml to 10 ml, or more may be used. In some cases, a dosage between about 10.sup.10 to 10.sup.12 rAAV genome copies per subject is appropriate. In certain embodiments, 10.sup.12 rAAV genome copies per subject is effective to target CNS tissues. In some embodiments the rAAV is administered at a dose of 10.sup.10, 10.sup.11, 10.sup.12, 10.sup.13, 10.sup.14, or 10.sup.15 genome copies per subject. In some embodiments the rAAV is administered at a dose of 10.sup.10, 10.sup.11, 10.sup.12, 10.sup.13, or 10.sup.14 genome copies per kg.
[0105] In some embodiments, rAAV compositions are formulated to reduce aggregation of AAV particles in the composition, particularly where high rAAV concentrations are present (e.g., .about.10.sup.13 GC/ml or more). Methods for reducing aggregation of rAAVs are well known in the art and, include, for example, addition of surfactants, pH adjustment, salt concentration adjustment, etc. (See, e.g., Wright F R, et al., Molecular Therapy (2005) 12, 171-178, the contents of which are incorporated herein by reference.)
[0106] Formulation of pharmaceutically-acceptable excipients and carrier solutions is well-known to those of skill in the art, as is the development of suitable dosing and treatment regimens for using the particular compositions described herein in a variety of treatment regimens. Typically, these formulations may contain at least about 0.1% of the active ingredient or more, although the percentage of the active ingredient(s) may, of course, be varied and may conveniently be between about 1 or 2% and about 70% or 80% or more of the weight or volume of the total formulation. Naturally, the amount of active ingredient in each therapeutically-useful composition may be prepared is such a way that a suitable dosage will be obtained in any given unit dose of the compound. Factors such as solubility, bioavailability, biological half-life, route of administration, product shelf life, as well as other pharmacological considerations will be contemplated by one skilled in the art of preparing such pharmaceutical formulations, and as such, a variety of dosages and treatment regimens may be desirable.
[0107] The pharmaceutical forms suitable for injectable use include sterile aqueous solutions or dispersions and sterile powders for the extemporaneous preparation of sterile injectable solutions or dispersions. Dispersions may also be prepared in glycerol, liquid polyethylene glycols, and mixtures thereof and in oils. Under ordinary conditions of storage and use, these preparations contain a preservative to prevent the growth of microorganisms. In many cases the form is sterile and fluid to the extent that easy syringability exists. It must be stable under the conditions of manufacture and storage and must be preserved against the contaminating action of microorganisms, such as bacteria and fungi. The carrier can be a solvent or dispersion medium containing, for example, water, ethanol, polyol (e.g., glycerol, propylene glycol, and liquid polyethylene glycol, and the like), suitable mixtures thereof, and/or vegetable oils. Proper fluidity may be maintained, for example, by the use of a coating, such as lecithin, by the maintenance of the required particle size in the case of dispersion and by the use of surfactants. The prevention of the action of microorganisms can be brought about by various antibacterial and antifungal agents, for example, parabens, chlorobutanol, phenol, sorbic acid, thimerosal, and the like. In many cases, it will be preferable to include isotonic agents, for example, sugars or sodium chloride. Prolonged absorption of the injectable compositions can be brought about by the use in the compositions of agents delaying absorption, for example, aluminum monostearate and gelatin.
[0108] For administration of an injectable aqueous solution, for example, the solution may be suitably buffered, if necessary, and the liquid diluent first rendered isotonic with sufficient saline or glucose. These particular aqueous solutions are especially suitable for intravenous, intramuscular, subcutaneous and intraperitoneal administration. In this connection, a sterile aqueous medium that can be employed will be known to those of skill in the art. For example, one dosage may be dissolved in 1 ml of isotonic NaCl solution and either added to 1000 ml of hypodermoclysis fluid or injected at the proposed site of infusion, (see for example, "Remington's Pharmaceutical Sciences" 15th Edition, pages 1035-1038 and 1570-1580). Some variation in dosage will necessarily occur depending on the condition of the host. The person responsible for administration will, in any event, determine the appropriate dose for the individual host.
[0109] Sterile injectable solutions are prepared by incorporating the active rAAV in the required amount in the appropriate solvent with various of the other ingredients enumerated herein, as required, followed by filtered sterilization. Generally, dispersions are prepared by incorporating the various sterilized active ingredients into a sterile vehicle which contains the basic dispersion medium and the required other ingredients from those enumerated above. In the case of sterile powders for the preparation of sterile injectable solutions, the preferred methods of preparation are vacuum-drying and freeze-drying techniques which yield a powder of the active ingredient plus any additional desired ingredient from a previously sterile-filtered solution thereof.
[0110] The rAAV compositions disclosed herein may also be formulated in a neutral or salt form. Pharmaceutically-acceptable salts, include the acid addition salts (formed with the free amino groups of the protein) and which are formed with inorganic acids such as, for example, hydrochloric or phosphoric acids, or such organic acids as acetic, oxalic, tartaric, mandelic, and the like. Salts formed with the free carboxyl groups can also be derived from inorganic bases such as, for example, sodium, potassium, ammonium, calcium, or ferric hydroxides, and such organic bases as isopropylamine, trimethylamine, histidine, procaine and the like. Upon formulation, solutions will be administered in a manner compatible with the dosage formulation and in such amount as is therapeutically effective. The formulations are easily administered in a variety of dosage forms such as injectable solutions, drug-release capsules, and the like.
[0111] As used herein, "carrier" includes any and all solvents, dispersion media, vehicles, coatings, diluents, antibacterial and antifungal agents, isotonic and absorption delaying agents, buffers, carrier solutions, suspensions, colloids, and the like. The use of such media and agents for pharmaceutical active substances is well known in the art. Supplementary active ingredients can also be incorporated into the compositions. The phrase "pharmaceutically-acceptable" refers to molecular entities and compositions that do not produce an allergic or similar untoward reaction when administered to a host.
[0112] Delivery vehicles such as liposomes, nanocapsules, microparticles, microspheres, lipid particles, vesicles, and the like, may be used for the introduction of the compositions of the present invention into suitable host cells. In particular, the rAAV vector delivered trans genes may be formulated for delivery either encapsulated in a lipid particle, a liposome, a vesicle, a nanosphere, or a nanoparticle or the like.
[0113] Such formulations may be preferred for the introduction of pharmaceutically acceptable formulations of the nucleic acids or the rAAV constructs disclosed herein. The formation and use of liposomes is generally known to those of skill in the art. Recently, liposomes were developed with improved serum stability and circulation half-times (U.S. Pat. No. 5,741,516). Further, various methods of liposome and liposome like preparations as potential drug carriers have been described (U.S. Pat. Nos. 5,567,434; 5,552,157; 5,565,213; 5,738,868 and 5,795,587).
[0114] Liposomes have been used successfully with a number of cell types that are normally resistant to transfection by other procedures. In addition, liposomes are free of the DNA length constraints that are typical of viral-based delivery systems. Liposomes have been used effectively to introduce genes, drugs, radiotherapeutic agents, viruses, transcription factors and allosteric effectors into a variety of cultured cell lines and animals. In addition, several successful clinical trails examining the effectiveness of liposome-mediated drug delivery have been completed.
[0115] Liposomes are formed from phospholipids that are dispersed in an aqueous medium and spontaneously form multilamellar concentric bilayer vesicles (also termed multilamellar vesicles (MLVs). MLVs generally have diameters of from 25 nm to 4 .mu.m. Sonication of MLVs results in the formation of small unilamellar vesicles (SUVs) with diameters in the range of 200 to 500 .ANG., containing an aqueous solution in the core.
[0116] Alternatively, nanocapsule formulations of the rAAV may be used. Nanocapsules can generally entrap substances in a stable and reproducible way. To avoid side effects due to intracellular polymeric overloading, such ultrafine particles (sized around 0.1 .mu.m) should be designed using polymers able to be degraded in vivo. Biodegradable polyalkyl-cyanoacrylate nanoparticles that meet these requirements are contemplated for use.
[0117] In addition to the methods of delivery described above, the following techniques are also contemplated as alternative methods of delivering the rAAV compositions to a host. Sonophoresis (ie., ultrasound) has been used and described in U.S. Pat. No. 5,656,016 as a device for enhancing the rate and efficacy of drug permeation into and through the circulatory system. Other drug delivery alternatives contemplated are intraosseous injection (U.S. Pat. No. 5,779,708), microchip devices (U.S. Pat. No. 5,797,898), ophthalmic formulations (Bourlais et al., 1998), transdermal matrices (U.S. Pat. Nos. 5,770,219 and 5,783,208) and feedback-controlled delivery (U.S. Pat. No. 5,697,899).
Kits and Related Compositions
[0118] The agents described herein may, in some embodiments, be assembled into pharmaceutical or diagnostic or research kits to facilitate their use in therapeutic, diagnostic or research applications. A kit may include one or more containers housing the components of the invention and instructions for use. Specifically, such kits may include one or more agents described herein, along with instructions describing the intended application and the proper use of these agents. In certain embodiments agents in a kit may be in a pharmaceutical formulation and dosage suitable for a particular application and for a method of administration of the agents. Kits for research purposes may contain the components in appropriate concentrations or quantities for running various experiments.
[0119] The kit may be designed to facilitate use of the methods described herein by researchers and can take many forms. Each of the compositions of the kit, where applicable, may be provided in liquid form (e.g., in solution), or in solid form, (e.g., a dry powder). In certain cases, some of the compositions may be constitutable or otherwise processable (e.g., to an active form), for example, by the addition of a suitable solvent or other species (for example, water or a cell culture medium), which may or may not be provided with the kit. As used herein, "instructions" can define a component of instruction and/or promotion, and typically involve written instructions on or associated with packaging of the invention. Instructions also can include any oral or electronic instructions provided in any manner such that a user will clearly recognize that the instructions are to be associated with the kit, for example, audiovisual (e.g., videotape, DVD, etc.), Internet, and/or web-based communications, etc. The written instructions may be in a form prescribed by a governmental agency regulating the manufacture, use or sale of pharmaceuticals or biological products, which instructions can also reflects approval by the agency of manufacture, use or sale for animal administration.
[0120] The kit may contain any one or more of the components described herein in one or more containers. As an example, in one embodiment, the kit may include instructions for mixing one or more components of the kit and/or isolating and mixing a sample and applying to a subject. The kit may include a container housing agents described herein. The agents may be in the form of a liquid, gel or solid (powder). The agents may be prepared sterilely, packaged in syringe and shipped refrigerated. Alternatively it may be housed in a vial or other container for storage. A second container may have other agents prepared sterilely. Alternatively the kit may include the active agents premixed and shipped in a syringe, vial, tube, or other container. The kit may have one or more or all of the components required to administer the agents to a subject, such as a syringe, topical application devices, or IV needle tubing and bag.
EXAMPLES
Example 1: Characterization of 12 AAV Vectors for Intravascular Delivery to Target CNS and Detarget Non-CNS Tissues by miRNA Regulation
[0121] The CNS gene transfer properties of 12 scAAVEGFP vectors of different serotypes, or natural variants were evaluated. RAAVs that cross the blood-brain-barrier (BBB) and target oligodendrocytes were discovered. Experiments were performed in neonatal mice (1 day old) and in adult mice (10 week old) (C57BL/6). The following AAV serotypes were tested: AAV1, AAV2, AAV5, AAV6, AAV6.2, AAV7, AAV8, AAV9, rh.10 (also referred to herein as AAVrh.10), rh.39, rh.43, CSp3.
[0122] The recombinant AAV vectors expressed an enhanced GFP reporter gene under the CMV-enhanced chicken .beta.-actin hybrid promoter and were produced by transient transfection in 293 cells. The neonatal day 1 pups were anesthetized with isoflurane. Then 100 .mu.L of rAAV vectors (4.times.10.sup.11 GC per mouse) was injected to the pups via superfacial temporal vein under a dissection microscope. In adult mice, rAAV was administered by tail vein injection (two different doses were evaluated 4.times.10.sup.11 GC per mouse or 4.times.10.sup.12 GC per mouse). Twenty-one days post injection, the treated animals were anesthetized and transcardially perfused with cold PBS and 4% (v/v) paraformaldehyde. Brains were extracted, immersed in 20% sucrose, and embedded in Tissue-Tek OCT. 40 .mu.m thick sections were cut and stained in 12-well plate with primary antibodies, e.g., anti-NeuN, anti-EGFP and anti-GFAP, overnight at 4.degree. C., then with secondary antibodies for 2 h at room temperature. Control mice received PBS injections.
[0123] In the neonatal study, the distribution of EGFP (+) cells throughout the brain at 3 wks post-infusion was observed. Large numbers of EGFP (+) cells with variable intensities were visible in different regions of the brains from the animals treated with 10 out of 12 vectors. In many instances the choroid plexus showed very strong EGFP expression, and transduced brain parenchyma cells appeared predominantly in periventricular regions. This indicates that a fraction of IV delivered vectors may enter the CNS via the choroid plexus-blood interface. In adults, substantial staining of brain vasculature was observed. Overall targeting efficiencies by AAVs to different regions of the brain was ranked as hypothalamus>medulla>cortex>hippocampus>cerebellum&- gt;thalamus. EGFP expression was not detected at high levels in neonatal mice that were administered rAAV2 or rAAV5 harboring the EGFP reporter gene by injection of 4.times.10.sup.11 GC per mouse in the superfacial temporal vein. (See Table 1 and FIGS. 1A to 1B and 2A to 2B for summary data).
[0124] Tissue sections were also immunofluorescently stained with anti-EGFP and -cell type specific marker antibodies to classify EGFP (+) cell types in the CNS. Detection sensitivity for EGFP (+) cells, particularly neurons and oligodendrocytes, was improved dramatically. Although different vectors transduced neurons at variable efficiencies, all 10 vectors (including AAV9) exhibited stronger tropisms to non-neuronal cells, especially astrocytes. One vector (AAV7) targeted oligodendrocytes more efficiently than the other 9 vectors. Several rAAVs transduced both neurons and/or astrocytes at higher efficiencies as compared to rAAV9 (AAVrh.10, rh.34, and rh.43). Extensive astrocyte transduction was observed in hypothalamus and medulla. Injection of certain vectors resulted in substantial neuron transduction in different regions of the brain, including neocortex, hippocampus, and hypothalamus. Some vectors appeared to transduce Purkinje cells in cerebella cortex (e.g., CSp3), while others effectively transduced blood vessel in neocortex, thalamus and hypothalamus. In addition, choroid plexuses in 3.sup.rd ventricle, lateral ventricle and 4.sup.th ventricle showed strong EGFP expression. EGFP expression was also evaluated in different spinal cord regions of neonatal and adult mice (results for neonatal studies are shown in FIG. 3).
[0125] Transduction of non-CNS tissues such as heart and skeletal muscle was observed (e.g., for AAV9, AAV8, and CSp3). In some cases, this may lead to some undesirable side effects. To address this issue, miRNA binding sites were incorporated into the 3' UTR of the transgene cassette and achieved highly specific and effective detargeting of AAV transduction from non-CNS tissues. To inhibit expression in liver, miRNA binding(s) for mR-122 were used. To inhibit expression in skeletal muscle and heart, miRNA binding(s) for mR-1 were used.
TABLE-US-00001 TABLE 1 AAV CNS TROPISMS AAV1 AAV2 AAV5 AAV6 AAV6.2 AAV7 AAV8 AAV9 rh.10 rh.39 rh.43 CSp3 Adult Cortex + + + ++ ++ +++ ++ + - + Hippocampus + + + ++ ++ +++ ++ + - + Thalamus + + + ++ ++ ++++ +++ ++ + + Hypothalamus + ++ + +++ ++ ++ +++ +++ + ++ Cerebellum + ++ + ++ +++ +++ ++++ + + + Brain Stem + ++ + ++ ++ ++++ +++ ++ - + Cervical +++ + + +++ +++ ++++ +++ +++ - + Thoracic +++ + + +++ +++ ++++ +++ ++ - + Lumbar +++ + + +++ +++ ++++++ +++ ++ - + Neo-Natal Cortex ++ + - ++ + + +++ ++ ++ ++ ++ ++ Hippocampus + + - - - +++ ++ + + ++ ++ + Thalamus + + - - + ++ + + + ++ + + Hypothalamus ++ - - + + ++++++ ++++++ + + ++++++ +++ - Cerebellum ++ - - + - + + + + ++ + + Brain Stem ++ - - + - ++ + + + +++++ +++ +++ Cervical - - - + ++ ++ +++ ++ +++++ ++++ +++ ++ Thoracic + - - + ++ +++ ++ ++ ++ ++++ +++ + Lumbar + - - + ++ ++ + + ++ +++ +++ + Extent of Tissue Tropsim (- no tropism; ++++++ high tropism) Based on Data in FIGS. 1A to 1B and 2A to 2B.
Example 2: Construction and Evaluation of a Recombinant AAVrh.10 Vector to Treat CD
[0126] Canavan disease (CD) is an inherited neurodegenerative disorder caused by mutations in the aspartoacylase gene (ASPA), leading to accumulation of N-acetyl-aspartic acid (NAA) in oligodendrocytes with resultant spongy degeneration of white matter in the brain. An initial clinical study on rAAV2-based ASPA gene therapy for CD achieved very limited success. It is believed, without wishing to be bound by theory, that an effective CD gene therapy will transduce oligodendrocytes throughout the CNS.
[0127] A rAAV vector is constructed that comprises a promoter operably linked with a region encoding ASPA protein (SEQ ID NO: 13 or 15) as a gene therapy vector for CD. The construct employs CAG (chicken -actin promoter with CMV enhancer) to drive the expression of ASPA having a coding sequence as set forth in SEQ ID NO: 14 or 16. The rAAV vector is package into rAAV particles using the triple transfection method. To evaluate its effectiveness, rAAV-ASPA is examined in an ASAP knock-out mouse model of CD for its ability to eliminate or attenuate the CD-like phenotypic of homozygous ASPA knock-out mice (Matalon R et al. The Journal of Gene Medicine, Volume 2 Issue 3, Pages 165-175). Homozygous ASPA knock-out mice exhibit neurological impairment, macrocephaly, generalized white matter disease, deficient ASPA activity and high levels of NAA in urine. Magnetic resonance imaging (MRI) and spectroscopy (MRS) of the brain of the homozygous mice show white matter changes characteristic of Canavan disease and elevated NAA levels. Heterozygous ASPA knock-out mice, which have no overt phenotype at birth, serve as controls.
Example 3: Therapeutic Efficacy and Safety Evaluation of an AAV Vector to Treat CD
[0128] The mouse model of CD is a C57BL/6 derived ASPA gene KO strain. The homozygous KO animals present biochemical and neurological defects similar to those observed in CD patients. CD mice provide an animal model for evaluating gene therapy and other therapeutics for the treatment of CD. CD mice are used to study the efficacy and safety of the novel gene therapy strategies for the treatment of CD.
[0129] Experiment Design
[0130] To examine therapeutic efficacy and safety, scAAV vectors (e.g., AAV7, AAV8, CSp3 and AAV9) carrying an optimized ASPA expression cassette are investigated in a preclinical gene therapy trial of CD. The vectors include miRNA binding site(s) to inhibit ASPA expression in non-CNS tissues. Both postnatal day-1 and 3-month-old adult animals are treated with each vector at two doses, 1 and 3.times.10.sup.14 GC/kg by intravenous administration. For the neonatal CD mice, two litters of animals receive each vector at each dose via temporal vein injections for necropsy of one litter each at 1- and 3-month time points. For the 3-month-old adult CD mice, 12 male animals are treated with each vector at each dose via tail vein injections. Six each of the treated animals are necropsied 1 and 3 months later. In further experiments, both postnatal day-1 and 3-month-old adult animals are treated with vectors at a dose in a range of 10.sup.11 to 10.sup.12 GC/subject by direct intraventricular administration.
[0131] Functional and Neurological Measurements During the Live Phase of the Study
[0132] 1). NAA metabolism. Urine samples are collected from the treated, untreated control, and wild type animals at days 14, 30, 45, 60, 75, and 90. The samples are analyzed by HPLC to determine the NAA levels.
[0133] 2). NAA accumulation and NAA-induced water retention in brain. MRI/MRS-based neuroimaging studies are performed on the live animals in all study groups at 1, 2, and 3 months after the vector treatment to measure spectral peak integrals for creatine/phosphocreatine and NAA as well as abnormal hyperintense areas in the brain.
[0134] 3). Liver function tests. Serum samples are collected from the animals in all study groups at days 14, 30, 60, and 90 to measure the levels of alanine transaminase (ALT) and aspartate aminotransferase (AST) as indicators of vector-related liver toxicity.
[0135] 4). Neurological tests. Tremors, walking with splayed legs at a slow and shaky pace, and ataxia are among the prominent neurological features of the CD mice. At 1, 2, and 3 months after the gene therapy treatment, the animals in all study groups are subjected to a walking-pattern analysis by staining their feet with color ink and then recording their walking patterns as footprints on white paper. The animals also are tested and scored on a rotarod test for their ability to maintain balance.
[0136] Enzymatic and Histopathological Analyses at the Endpoints of the Study
[0137] 1). ASPA activities in the brain and non-CNS tissues. On-target and off-target expression of ASPA are analyzed by collecting brain, liver, heart and pancreatic tissues at necropsy to measure ASAP activities in the respective tissue homogenates.
[0138] 2). Brain white matter and liver pathologies. To examine potential improvement in brain white-matter pathology and vector-related liver toxicity resulting from the gene therapy, brain and liver tissues are harvested and fixed, paraffin-embedded and sectioned, and stained with hematoxylin and eosin. Histopathological examination is performed by a pathologist.
Example 4: Delivery of Therapeutic Genes to the CNS Cells by AAVrh.10
[0139] A screen of different AAV serotypes, was developed to identify candidates for a therapeutic gene transfer to the CNS. A recombinant AAV vector was constructed that expresses EGFP. The rAAV vector was packaged into four different AAVs: AAV1, 8, 9 and 10. Adult mice were injected with the AAVs into the CSF in the lumbar position. AAV1, 8 and 9 transduced cells only in the vicinity of the injection site at the lumber region of the spinal cord following administration of .about.4.8.times.10.sup.10 particles. Surprisingly, AAVrh.10 transduced cells in the gray matter along the entire spinal cord and brainstem following the same injection protocol and dosage as AAV1, AAV8 and AAV9 (FIG. 4A). Recently, AAV9 has been shown to cross the blood brain barrier (BBB) and transduce spinal cord cells after intravenous injection. A weak signal was observed in the cerebellum and strong signals in the brainstem and spinal cord. A weak signal (similar to the cerebellum) in the forebrain was also observed. Without wishing to be bound by theory, it is believed that CSF flow and diffusion allows the virus spread along the entire spinal cord, but that the ability of a virus to flow and diffuse depends on the structure of the viral capsid. The transduced cell types include neurons and oligodendrocytes. But the majority appears to be astrocytes (FIG. 4B), as indicated by overlap of EGFP with GFAP-positive cells. Substantial overlap with the microglia marker, Iba-1 was not observed. A number of motor neurons were transduced as indicated by overlap of EGFP expression and NeuN staining. It was surprising that among the astrocytes, only those situated in the gray matter were transduced and those that were situated in the white matter and beneath the pia matter were not transduced. This was striking because the virus is likely to be exposed to astrocytes in these areas since it was administered in the subarachnoid space.
Example 5: Construction of a Recombinant AAVrh.10 Vector to Treat ALS
[0140] An recombinant AAV system was developed as a treatment for ALS. A rAAVrh.10 vector was constructed that expresses a microRNA targeting SOD1 (FIG. 5A). This microRNA was identified as miR-SOD1. The construct employed CAG (chicken -actin promoter with CMV enhancer) to drive the expression of EGFP and miR-SOD1 that was located in an intron in the 3'-UTR.
[0141] The silencing potency of 9 miRNA constructs was evaluated. The constructs were transfected into HEK293 cells. After 48 hours, RNA was isolated and Northern blot was carried out to detect SOD1 mRNA (FIG. 5B). MiR-SOD1#5 (SEQ ID NO: 26) silenced SOD1 expression most potently. Next, miR-SOD1#5 was packaged into AAVrh.10 (FIG. 5D), which was used to infect HEK293 cells. Total cellular protein was extracted 43 hours after the infection and blotted to detect SOD1 (FIG. 5C). Inhibition of expression of SOD1 at the protein level was observed.
Example 6: Delivery of Therapeutic Genes to the CNS Cells to Treat ALS
[0142] Large batches of AAVrh.10-miR-SOD1 and AAVrh.10-miR-Scr (scrambled miRNA) were produced using standard techniques. Self-complementary AAV (scAAV) was made because it mediates transduction with higher efficiency than conventional single stranded AAV [14]. A scAAVrh.10 was tested and found to express EGFP more rapidly (within 1 week) and stronger than a single stranded AAV.
[0143] AAVrh.10-miR-SOD1 was administered to one group of G93A mice (high SOD1 expressers) and AAVrh.10-miR-Scr to another group of G93A mice (n=15). The AAVrh.10 was injected intrathecally into the CSF in the lumbar area and injected intraventricularly into the forebrain in mice of 60 days of age (.about.4.8.times.10.sup.10 particles in 8 ul).
[0144] The animals were allowed to live their natural lifespan before succumbing to ALS. The lifespan was compared between the two groups. It was found that mice receiving the AAVrh.10-miR-SOD1 virus, which expresses a SOD1miR5 (SEQ ID NO: 26), lived on average 135 days (.+-.14 days), whereas mice receiving the AAVrh.10-miR-Scr, which expresses a scrambled miRNA (SEQ ID NO: 31), lived on average 122 days (.+-.6 days) (FIG. 6B). Moreover, by examining the extent of EGFP expression in cervical, thoracic, and lumber spinal cord tissue, a correlation in the levels of expression in these tissues, particularly with cervical tissue, and lifespan was observed in AAVrh.10-miR-SOD1 treated mice (FIG. 7A), but not AAVrh.10-miR-Scr treated mice (FIG. 7B). These results suggest that silencing mutant SOD1 expression in the cervical spinal cord is particularly beneficial in extending survival. A subset of the animals from each group were perfused with fixative, sectioned and stained for SOD1 in the spinal cord. SOD1 was detected using standard techniques [9]. SOD1 staining intensity in EGFP expressing cells was reduced compared with the non-EGFP cells that are transduced with AAVrh.10-miR-SOD1 (FIG. 6A, showing knockdown of SOD1 expression in astrocytes). Reduction of expression of SOD1 was not observed in cells transduced with AAVrh.10-miR-Scr.
[0145] Tissues from another subset of animals in both groups were dissected to estimate transduction levels. The levels of transduction were estimated by determining the viral genome content using PCR on DNA samples obtained from different CNS and non-CNS regions. Measurements in non-CNS tissues (e.g. liver) provided an indication of whether virus had leaked to the periphery. Northern and Western analysis was performed to measure the SOD1 levels in the spinal cord. The antibody used for SOD1 detection was polyclonal, sheep anti-human SOD1, by Biodesign International, catalog#K90077C.
Example 7: Combined Intrathecal/Intraventricular Administration Protocol
[0146] AAV viruses were injected into mouse CSF by lumbar intrathecal injection and/or brain third ventricle injection. Injection into mice lumbar subarachnoid space was carried out using a method modified from Wu et al. [22]. A thin catheter (about 5 cm) was made by stretching PE10 tube to the inner diameter 0.12 mm. The stretched section was cut to 1.7 to 1.9 mm, and two beads (1 mm apart) were made between the thin and the thick sections by heating and pressing the tube. To implant the catheter, the mouse was anesthetized by injection of Avertin (1.2% 2,2,2-tribromoethanol in 2% tert-amyl alcohol and PBS) intraperitoneally at 0.23 ml/10 g of body weight [23]. The catheter was then implanted between the L5 and L6 vertebra. The catheter was stitched to the surface muscle at the beaded area. Viruses of dose from 4.80E+10 Genome Copy (for virus screening, in 6 ul) to 2.40E+10 Genome copy (for therapy, in 8 ul) were injected via the catheter by a Hamilton syringe at a speed of 2 ul/minute. The catheter was sealed at the end by heat and left in place for one day. Wound was closed by clips. Injection into brain third ventricle was carried out using a Stoelting Stereotaxic Instrument and micro-injection pumps from World Precision Instruments following standard stereotaxic procedure. Same doses of virus were injected into the third ventricle at a rate of 1 ul/minutes.
[0147] Estimated doses for human and monkeys and comparison with IV injection are shown below. The two types of monkey are similar in size.
TABLE-US-00002 TABLE 2 Estimated Doses for Human and Monkeys Estimated CSF Avg CSF production rate Estimate particles/g of Species ml ml/hour dose (GC) body weight mouse 0.035 0.018 2.40E+10 1.2E+09 human 140 21 9.6E+13 1.3E+09 Macaca mulatta 14 2.5 9.6E+12 1.7E+09 (rhesus monkeys) Macaca See, Foust KD, et al., 1.00E+14 2.20E+11 fascicularis Nature Biotechnology, (cynomolgus Volume 28, Number 3, macaque) March 2010, 271-274
REFERENCES FOR BACKGROUND AND EXAMPLES 1-7
[0148] 1. Daya S, Berns K I: Gene Therapy Using Adeno-Associated Virus Vectors. Clin Microbiol Rev 2008, 21:583-593. [0149] 2. Eberling J L, Jagust W J, Christine C W, Starr P, Larson P, Bankiewicz K S, Aminoff M J: Results from a phase I safety trial of hAADC gene therapy for Parkinson disease. Neurology 2008, 70:1980-1983. [0150] 3. Feigin A, Kaplitt M G, Tang C, Lin T, Mattis P, Dhawan V, During M J, Eidelberg D: Modulation of metabolic brain networks after subthalamic gene therapy for Parkinson's disease. Proceedings of the National Academy of Sciences 2007, 104:19559-19564. [0151] 4. Cideciyan A V, Hauswirth W W, Aleman T S, Kaushal S, Schwartz S B, Boye S L, Windsor E A M, Conlon T J, Sumaroka A, Pang J-j, et al: Human RPE65 Gene Therapy for Leber Congenital Amaurosis: Persistence of Early Visual Improvements and Safety at 1 Year. Hum Gen Ther, 0. [0152] 5. Raoul C, Abbas-Terki T, Bensadoun J-C, Guillot S, Haase G, Szulc J, Henderson C E, Aebischer P: Lentiviral-mediated silencing of SOD1 through RNA interference retards disease onset and progression in a mouse model of ALS. Nat Med 2005, 11:423-428. [0153] 6. Kaspar B K, Llado J, Sherkat N, Rothstein J D, Gage F H: Retrograde Viral Delivery of IGF-1 Prolongs Survival in a Mouse ALS Model. Science 2003, 301:839-842. [0154] 7. Azzouz M, Ralph G S, Storkebaum E, Walmsley L E, Mitrophanous K A, Kingsman S M, Carmeliet P, Mazarakis N D: VEGF delivery with retrogradely transported lentivector prolongs survival in a mouse ALS model. Nature 2004, 429:413-417. [0155] 8. Ralph G S, Radcliffe P A, Day D M, Carthy J M, Leroux M A, Lee D C P, Wong L-F, Bilsland L G, Greensmith L, Kingsman S M, et al: Silencing mutant SOD1 using RNAi protects against neurodegeneration and extends survival in an ALS model. Nat Med 2005, 11:429-433. [0156] 9. Wu R, Wang H, Xia X, Zhou H, Liu C, Castro M, Xu Z: Nerve Injection of Viral Vectors Efficiently Transfers Transgenes into Motor Neurons and Delivers RNAi Therapy Against ALS. Antioxidants & Redox Signaling 2009, 11:1523-1534. [0157] 10. Foust K D, Nurre E, Montgomery C L, Hernandez A, Chan C M, Kaspar B K: Intravascular AAV9 preferentially targets neonatal neurons and adult astrocytes. Nat Biotechnol 2009, 27:59-65. [0158] 11. Duque S, Joussemet B, Riviere C, Marais T, Dubreil L, Douar A M, Fyfe J, Moullier P, Colle M A, Barkats M: Intravenous administration of self-complementary AAV9 enables transgene delivery to adult motor neurons. Mol Ther 2009, 17:1187-1196. [0159] 12. Vandenberghe L H, Wilson J M, Gao G: Tailoring the AAV vector capsid for gene therapy. Gene Ther 2009, 16:311-319. [0160] 13. McBride J L, Boudreau R L, Harper S Q, Staber P D, Monteys A M, Martins I, Gilmore B L, Burstein H, Peluso R W, Polisky B, et al: Artificial miRNAs mitigate shRNA-mediated toxicity in the brain: Implications for the therapeutic development of RNAi. Proceedings of the National Academy of Sciences 2008, 105:5868-5873. [0161] 14. McCarty D M: Self-complementary AAV Vectors; Advances and Applications. Mol Ther 2008, 16:1648-1656. [0162] 15. Boillee S, Yamanaka K, Lobsiger C S, Copeland N G, Jenkins N A, Kassiotis G, Kollias G, Cleveland D W: Onset and progression in inherited ALS determined by motor neurons and microglia. Science 2006, 312:1389-1392. [0163] 16. Xia X, Zhou H, Huang Y, Xu Z: Allele-specific RNAi selectively silences mutant SOD1 and achieves significant therapeutic benefit in vivo. Neurobiol Dis 2006, 23:578-586. [0164] 17. Yamanaka K, Chun S J, Boillee S, Fujimori-Tonou N, Yamashita H, Gutmann D H, Takahashi R, Misawa H, Cleveland D W: Astrocytes as determinants of disease progression in inherited amyotrophic lateral sclerosis. Nat Neurosci 2008, 11:251253. [0165] 18. Di Giorgio F P, Carrasco M A, Siao M C, Maniatis T, Eggan K: Non-cell autonomous effect of glia on motor neurons in an embryonic stem cell-based ALS model. Nat Neurosci 2007, 10:608-614. [0166] 19. Nagai M, Re D B, Nagata T, Chalazonitis A, Jessell T M, Wichterle H, Przedborski S: Astrocytes expressing ALS-linked mutated SOD1 release factors selectively toxic to motor neurons. Nat Neurosci 2007, 10:615-622. [0167] 20. Wang Y, Ou Mao X, Xie L, Banwait S, Marti H H, Greenberg D A, Jin K: Vascular Endothelial Growth Factor Overexpression Delays Neurodegeneration and Prolongs Survival in Amyotrophic Lateral Sclerosis Mice. J Neurosci 2007, 27:304-307. [0168] 21. Storkebaum E, Lambrechts D, Dewerchin M, Moreno-Murciano M-P, Appelmans S, Oh H, Van Damme P, Rutten B, Man W Y, De Mol M, et al: Treatment of motoneuron degeneration by intracerebroventricular delivery of VEGF in a rat model of ALS. Nat Neurosci 2005, 8:85-92. [0169] 22. Wu, W. P., Xu, X. J., and Hao, J. X. (2004) J. Neurosci. Methods 133, 65-69 [0170] 23. Papaioannou, V. E., and Fox, J. G. (1993) Lab. Anim. Sci. 43, 189-192
Example 8. MicroRNA-Regulated, Systemically Delivered rAAV9 Introduction to the Example
[0171] This example involves the use of tissue-specific, endogenous microRNAs (miRNAs) to repress rAAV expression outside the CNS, by engineering perfectly complementary miRNA-binding sites into the rAAV9 genome. The example describes recombinant adeno-associated viruses (rAAVs) that can cross the blood-brain-barrier and achieve efficient and stable transvascular gene transfer to the central nervous system (CNS), while de-targeting certain other tissues (e.g., liver, heart, skeletal muscle and other tissues) The approaches described in this example allowed simultaneous multi-tissue regulation and CNS-directed stable transgene expression without detectably perturbing the endogenous miRNA pathway. Regulation of rAAV expression by miRNA was primarily via site-specific cleavage of the transgene mRNA, generating specific 5' and 3' mRNA fragments.
[0172] Gene transfer mediated by recombinant adeno-associated virus (rAAV), as disclosed herein, is useful for treatment of a large number of neurological disorders. It has been found that rAAV vectors disclosed herein cross the blood-brain barrier and are specifically expressed in the CNS. Thus, the vectors may be used for intravascular delivery of rAAV for gene therapy of CNS diseases, including those that affect large areas of the brain and spinal cord.
[0173] This example describes the use of endogenous microRNAs (miRNAs) to suppress transgene expression outside the CNS. miRNAs are small, noncoding RNAs that regulate gene expression by post-transcriptional silencing. In general, miRNAs may silence genes by two mechanisms. When partially complementary to mRNA sequences, they typically reduce target mRNA stability and protein expression (e.g., by two- to fourfold or less), a mode of regulation thought to tune mRNA expression. In contrast, when miRNAs are nearly perfectly complementary to their mRNA targets, they typically bring about cleavage of the mRNA, triggering its wholesale destruction.
[0174] In particular, this example describes the use of miRNAs to detarget rAAV9 expression both separately and concurrently in the liver, heart, and skeletal muscle, the three tissues that are most efficiently targeted by intravenously delivered rAAV9. Silencing of transgene expression in liver, heart, and muscle exploited the natural expression of the abundant (60,000 copies/cell) miRNAs, miR-122, which is expressed in hepatocytes, and miR-1, a miRNA found in the heart and skeletal muscle of virtually all animals. miR-122-binding sites have been successfully used to prevent hepatotoxicity of a transgene from an adenovirus vector. Perfectly complementary sites for miR-1, miR-122, or both were engineered into the 3' untranslated region (UTR) of a nuclear-targeted, .beta.-galactosidase (nLacZ) reporter transgene whose expression was driven by a cytomegalovirus-enhancer, chicken .beta.-actin (CB) promoter. This example presents multiple independent results indicating that the miRNAs repress nLacZ expression by cleaving the transgene mRNA at exactly the same site as by all Argonaute-bound small RNAs in eukaryotic cells. When delivered systemically in vivo, the miRNA-detargeted rAAV9 vector successfully expressed the reporter transgene in the CNS, but not the liver or heart or skeletal muscle.
Results
[0175] miRNAs Efficiently Repress Reporter Gene Expression in Cultured Cells
[0176] To evaluate a strategy for rAAV-mediated transduction, one or three tandem copies of a perfectly complementary binding site for miR-1 or miR-122 were introduced into the 3' UTR of nLacZ in a rAAV plasmid vector. The constructs were transfected into HuH7 cells, a human hepatoma cell line expressing--16,000 copies of miR-122 per cell, and measured the number of nLacZ-positive cells. The number of nLacZ-expres sing HuH7 cells for the one-site plasmid was about half that of the no site control; three sites reduced the number of nLacZ-expressing cells more than sevenfold (FIG. 12A).
[0177] Next, expression of the nLacZ constructs was analyzed in human embryonic kidney 293 cells, which naturally express low levels of both miR-122 and miR-1, when miR-1 or miR122 was introduced as a pri-miRNA from a second plasmid. 293 cells were transfected with the nLacZ reporter plasmids carrying 0, 1, or 3 miR-122 or miR-1-binding sites, together with a plasmid expressing either pri-miR-122 (FIG. 12B) or pri-miR-1 (FIG. 12C). To vary the concentration of the miRNA, either a low (1:3) or a high (1:10) molar ratio of the nLacZ-binding site plasmid to the miRNA expression plasmid was used. When miR-122 or miR-1 was introduced into the cells, nLacZ expression was repressed only when the nLacZ reporter mRNA contained the corresponding miRNA-binding sites; there was no reduction of nLacZ-positive cells when miR-1 was coexpressed with nLacZ containing miR-122-binding sites or when miR-122 was coexpressed with nLacZ containing miR-1-binding sites (FIGS. 12B and 12C).
Tissue-Specific Endogenous miRNAs Regulate Expression of rAAV9 Delivered Systemically in Adult Mice
[0178] To evaluate miRNA regulation of systemically delivered AAV9CBnLacZ vectors in vivo, AAV9CBnLacZ vectors carrying 0, 1, or 3 miRNA-binding sites perfectly complementary to either miR-122 or miR-1 were produced. The vectors were administered by tail vein injection to adult male C56BL/6 mice at a dose of 5.times.10.sup.13 genome copies per kg (GC/kg) body weight. Four weeks later, the liver and heart of the transduced animals were examined. LacZ staining revealed that the nLacZ transgene was silenced by the endogenous miRNAs in the cell type and organ in which they are predominantly expressed: the transgene was specifically silenced by miR-122 in the liver and by miR-1 in the heart (FIGS. 13A and 13B). While nLacZ positive cells were reduced in the livers of the animals treated with rAAV9CBnLacZ bearing one or three miR-122-binding sites, nLacZ expression levels in the hearts of the same animals were similar to those in the animals treated with AAV9CBnLacZ bearing no sites (FIG. 13A). Similarly, nLacZ expression was not detected in the hearts of the animals that received AAV9CBnLacZ containing one or three miR-1-binding sites, but nLacZ expression in the livers of the same animals was not affected as compared to that in the control animal (FIG. 13B). These data suggest that the greater the number of sites for a miRNA in rAAV, the lower the nLacZ expression in the tissue where the corresponding miRNA was expressed (FIGS. 13A and 13B).
[0179] Next, to evaluate whether transgene silencing could be achieved simultaneously in multiple tissues, different numbers of both miR-122- and miR-1-binding sites were inserted in the 3' UTR of the rAAV9CBnLacZ genome and examined for their expression in rAAV9 transduced mice. Histochemical staining of tissue sections showed that nLacZ expression was suppressed in both heart and liver for rAAV9CBnLac containing one or three copies each of the miR-1- and miR-122-binding sites, but nLacZ was readily detectable in pancreas, where expression of both miR-122 and miR-1 was low (FIG. 13C). Quantitative, .beta.-galactosidase assays of homogenized liver tissue similarly showed that nLacZ expression was significantly lower when the transgene contained the miRNA-binding sites (one miR-122-binding site: 7.8.+-.7.4%, P value=0.005; three miR-122-binding sites: 1.6.+-.1.0%, P value=0.005; one miR-1-plus one miR-122-binding site: 8.6.+-.5.7%, P value=0.005; three miR-1-plus three miR-122-binding sites: 3.1.+-.1.2%, P value=0.005; three miR-1-binding sites: 105.7.+-.11.6%) (FIG. 13D).
miRNA Repression of rAAV Expression does not Perturb Endogenous miRNA Pathways
[0180] Highly expressed transgenes bearing miRNA-complementary sites have been reported to promote degradation of the corresponding miRNA. The levels of miR-122, miR-22, miR-26a, and let-7 were determined in rAAV transduced liver. No difference in abundance of the four miRNAs was detected among the three study groups (FIG. 14A). Moreover, data from high throughput sequencing analyses of small RNA from the livers of one animal each from the three study groups show no change in miRNA levels.
[0181] In order to determine whether the miRNA-binding sites in the transgene transcripts would deregulate the expression of the known endogenous target mRNAs of miR-122 or miR-1, the expression of cyclin G1, a miR-122 target in liver (FIGS. 14B and 14C) and calmodulin, a miR-1 target in heart (FIG. 14D) were analyzed. No significant alteration in cyclin G1 or calmodulin expression was detected. miR-122 regulates cholesterol biosynthesis in the liver, and agents that block miR-122 function may produce readily detectable changes in serum cholesterol levels. No change in total cholesterol, high-density lipoprotein, or low-density lipoprotein levels was detected in mice 4 weeks after transduction with either control rAAV9 or rAAV9 expressing a transgene bearing miR-122-binding sites (FIG. 14E). It was concluded that in this example miRNA-mediated detargeting of rAAV expression had no detectable effect on endogenous miRNA expression or function.
Endogenous miRNAs Silence rAAV Transduction by Site-Specific Cleavage of Transgene mRNA
[0182] To determine how miRNAs suppress expression of transgenes delivered by rAAV in vivo, the transgene mRNA in liver was characterized by conventional PCR (FIG. 15B), quantitative reverse transcription PCR (qRT-PCR) (FIG. 15C), Northern hybridization (FIGS. 15D and 15E), and rapid amplification of 5' complimentary DNA (cDNA) ends (5' RACE; FIG. 15F). When primers were used that amplify the region between the 3' end of nLacZ (A.sup.+F primer) and the 5' end of the poly(A) signal (APR primer), an amplicon that spans the miRNA-biding sites, a 145 basepair (bp) product was detected after 26 cycles of amplification for the samples that received control rAAV. An additional six cycles of amplification were required to detect a weak 220 bp band for the samples transduced by rAAV containing three miR-122-binding sites. These data are consistent with low levels of intact nLacZ mRNA (FIGS. 15A and 15B).
[0183] To quantitatively assess the extent of the miRNA-directed repression of the transgene transcripts, qRT-PCR was performed using either oligo(dT) or random hexamer primers for reverse-transcription and PCR primer pairs that span either a 5' (nLacZ5'F/5'R), or 3' (nLacZ 3'F13'R) region of the nLacZ coding sequence (FIG. 15A). The levels of nLacZ mRNA were examined with intact 5' and 3' ends in total liver RNA extracted from four animals that received the control rAAV9CBnLacZ and four that received rAAV9CBnLacZ containing three miR-122-binding sites in the 3' UTR. Reductions ranging from 3.+-.1 (random hexamer) to 7.+-.1 (oligo[dT])-fold in nLacZ mRNA with an intact 3' end were observed in the animals that had received rAAV9 containing miR-122-binding sites, relative to the control. In contrast, little or no decrease in nLacZ mRNA with an intact 5' end were detected for the same samples using the 5'F/5'R primer pair (FIG. 15C). These results indicate that the primary mode of turnover of the mRNA that has been cleaved by a miRNA was 3'-to-5' exonucleolytic degradation.
[0184] To further characterize the fate of the transgene mRNA targeted by miR-1 or miR-122, Northern blot analyses was performed. A transgene probe binding to the 5' end of nLacZ mRNA detected a .about.3.4 kb RNA in an animal injected with control rAAV9CbnLacZ, the expected size of the of the full-length nLacZ transcript; a slightly larger band was detected in the liver sample from a mouse treated with rAAV9CBnLacZ bearing three miR1-binding sites (FIGS. 15A and 15D). In contrast to the single transcript detected for the rAAV9 expressing nLacZ bearing three miR-1-binding sites, two RNAs of different sizes were detected for the rAAV expressing nLacZ bearing three miR-122 sites (FIG. 15D).
[0185] The lengths of these transcripts indicate that the longer transcript likely represents the full-length mRNA, whereas the shorter, more abundant transcript corresponds to 5' fragments of nLacZ RNA cleaved by miR-122 at the corresponding miR-122-binding sites in the 3' UTR (FIG. 15D).
[0186] To confirm this observation, the Northern analysis was repeated using an RNA probe spanning a portion of 3' UTR of the transgene mRNA. In addition to detecting full-length nLacZ transcripts in the samples transduced by rAAV9 lacking miRNA-binding sites, two closely migrating species smaller than the 281 nucleotide RNA marker were detected. The size of these fragments was consistent with miRNA-directed 3' cleavage products of the nLacZ mRNA (FIG. 15E). These two 3' cleavage products were also detected by gel electrophoresis of the product from the 5' RACE experiment described below (FIG. 15F).
[0187] To determine whether such target cleavage occurs in vivo when the nLacZ transcript contained miR-1 or miR-122-binding sites, rapid amplification of 5' cDNA ends (5' RACE) was performed. FIGS. 16A to 16B present the sequences of 21 clones recovered using 5' RACE from liver RNA (FIG. 16A) and 22 clones isolated from heart RNA (FIG. 16B) from the animals injected with rAAV9 in which the nLacZ 3' UTR contained three miR-1 and three miR-122-binding sites. In liver, the sequence signatures for miR-122-directed cleavage of the transgene mRNA were detected at each miR-122-binding site: 5% for the first binding site, 48% for the second binding site, and 43% for the third binding site. All 5' ends mapped to the phosphate that lies between the target nucleotides that pair with positions 10 and 11 of the sequence perfectly complementary to miR-122, the precise site cleaved by small RNAs bound to Argonaute proteins in all eukaryotes (FIG. 17A). Similar results were obtained in the heart for the rniR-1 sites (FIG. 17B).
[0188] Table 3 presents an expanded 5' RACE analysis for additional vector groups. It was noted that none of the 5' RACE products sequenced corresponded to miR-1-directed site-specific cleavage in liver or miR-122-directed site-specific cleavage in heart (Table 3). Although no cleavage was detected within miR-1-binding sites in the liver, some clones from heart were cleaved within the miR-122-binding sites, but not at the hallmark position for miRNA-directed cleavage.
Intravascularly Delivered rAAV9 can be Efficiently Controlled by Endogenous miRNAs
[0189] MiRNA-1 and miRNA-122-binding sites were added into the scAAV9CB enhanced GFP (EGFP) vector genome and injected 10-week-old C57BL/6 male mice with 2.times.10.sup.14 GC/kg. After 3 weeks, 40 .mu.m sections of brain and spinal cord and 8 .mu.m sections of liver, heart, and skeletal muscle were prepared and examined for EGFP protein expression. It was found that intravenously delivered scAAV9CBEGFP efficiently transduced the CNS; EGFP was readily detectable in the thalamus region of the brain and the cervical region of the spinal cord, but also in non-CNS tissues such as liver, heart, and muscle (FIG. 17A). In contrast, transgene expression in those non-CNS tissues was reduced when miR-1 and miR122-binding sites were included in the transgene; EGFP expression was unaltered in the CNS, where miR-1 and miR-122 were not present (FIG. 17A). Quantitative RT-PCR was used to measure the differential expression of the rniRNA-repressed EGFP transgene in brain (41.2.+-.7.7%), liver (3.0.+-.0.5%), heart (0.4.+-.0.1%), and muscle (1.3.+-.0.4%), relative to the EGFP transgene lacking miRNA-binding sites (FIG. 17B). To eliminate changes associated with transduction efficiency between experiments, the data were normalized to the number of vector genomes detected in the experimental and control samples. Similar to the microscopic analyses of native EGFP expression, the qRT-PCR data show that the presence of miR-122- or miR-1-binding sites reduced transgene expression in liver (20-fold), heart (100-fold), and muscle (50-fold), but did not detectably alter transgene expression in brain.
Discussion of Results
[0190] This example shows that rAAV9 can be engineered so that endogenous miRNAs repress transgene expression outside the CNS. The results indicate that such engineered rAAV9s may be used in therapies for the degenerating neurons associated with Parkinson's disease, Alzheimer's disease and amyotrophic lateral sclerosis, by expressing neurotrophic growth factors such as insulin-like growth factor, brain-derived neurotrophic factor or glial-derived neurotrophic factor in the transduced astrocytes. This approaches eliminates or lessens non-CNS expression derived from the peripheral tissues transduced by systemically delivered rAAV9.
[0191] Achieving transgene expression in primarily only the target tissues is a consideration for the clinical development of safe CNS gene delivery. The results in this example indicate that endogenous miRNAs can be harnessed to restrict the tissue- and cell-type specificity of rAAV expression, as was initially shown for lentiviral vectors. The data demonstrate that endogenous miRNAs can effectively repress transgene expression from rAAV. In both heart and liver, the miRNAs repressed transgene expression by directing endonucleolytic cleavage of the transgene mRNA (FIG. 18). MiRNA regulation of rAAV expression did not perturb the expression or function of the corresponding endogenous miRNA, allowing transgene expression to be restricted to the CNS in mice. The example indicates that a strategy that combines multiple binding sites for miRNAs expressed in the periphery but not the CNS is useful for the development of safer, CNS-specific gene therapy vectors.
Materials And Methods
[0192] Vector design, construction, and production. Perfectly complementary miRNA-binding sites were designed based on the annotated miR-1 and miR-122 sequences in miRBase and inserted into the BstBI restriction site in the 3' UTR of the nLacZ expression cassette of the ubiquitously expressed pAAVCB nuclear-targeted .beta.-galactosidase (nLacZ) plasmid using synthetic oligonucleotides (FIG. 15A and Table 3). This vector uses a hybrid cytomegalovirus enhancer/CB promoter cassette that is active in most cells and tissues. To express miR-122 and miR-1, pri-miR-122 and pri-miR-1 fragments were amplified by PCR from C57/B6 mouse genomic DNA (Table 4) and inserted into the XbaI restriction site 3' to a firefly luciferase cDNA in the pAAVCBELuc plasmid. The identity of each pri-miRNA was verified by sequencing. AAV9 vectors used in this study were generated, purified, and tittered.
[0193] Cell Culture and Transfection.
[0194] HEK-293 and HuH7 cells were cultured in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum and 100 mg/l of penicillin-streptomycin (Hyclone, South Logan, Utah). Cells were maintained in a humidified incubator at 37.degree. C. and 5% CO.sub.2. Plasmids were transiently transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, Calif.) according to the manufacturer's instructions.
[0195] Mouse Studies.
[0196] Male C57BL/6 mice (Charles River Laboratories, Wilmington, Mass.) were obtained and maintained. To monitor lipid profiles of the study animals, serum samples were collected 4 weeks after rAAV9 injection and analyzed for total cholesterol, high-density lipoprotein and low-density lipoprotein on a COBAS C 111 analyzer (Roche Diagnostics, Lewes, UK). To evaluate endogenous miRNA-mediated, CNS-restricted EGFP gene transfer, 10-week-old male C57BL/6 mice were injected intravenously (tail vein) with AAV9CBnLacZ-[miR-122-binding site (BS).sub.1]. AAV9CBnLacZ-(miR-122BS).sub.3. AAV9CBnLacZ-(miR-1BS).sub.1. AAV9CBnLacZ-(miR-1BS).sub.3. AAV9CBnLacZ-(miR-1BS)i-(miR-122BS).sub.1, and AAV9CBnLacZ-(miR-1BS).sub.3-(miR-122BS).sub.3, respectively, at 5.times.10.sup.13 GC/kg body weight) or scAAV9CBEGFP at 2.times.10.sup.14 GC/kg body weight). Animals receiving nLacZ vectors were necropsied 4 weeks later; 8 .mu.m cryosections of liver, heart, and pancreas tissues were prepared for X-gal-histochemical staining. Animals that received EGFP vectors were necropsied 3 weeks later and fixed by transcardial perfusion with 4% (wt/vol) paraformaldehyde. Brain, spinal cord, liver, heart, and muscle were harvested for cryosectioning. Brain and cervical spinal cord tissue were stained as floating sections in a 12-well plate using rabbit anti-EGFP antibody (Invitrogen) diluted 1:500, followed by goat anti-rabbit secondary antibody (Invitrogen) diluted 1:400. Outside the CNS, EGFP expression was detected directly by fluorescence. EGFP and antibody fluorescence was recorded using a Nikon TE-20005 inverted microscope at .times.10 magnification and an exposure time of 3 seconds for liver, heart, and muscle, and 5 seconds for thalamus (brain) and cervical spinal cord.
[0197] Vector Genome Quantification by qPCR.
[0198] Genome DNA was extracted from the selected tissues using QIAamp DNA Mini Kit (Qiagen, West Sussex, UK), according to the manufacturer's instructions. Quantitative PCR were carried out in triplicate using Ring DNA and 0.3 .mu.mol/l EGFP-specific primers (EGFP-F and EGFP-R) using GoTaq qPCR master mix (Promega, Madison, Wis.) in a StepOne Plus real-time PCR instrument (Applied Biosystems, Foster City, Calif.).
[0199] qRT-PCR Analysis.
[0200] RNA was extracted using Trizol (Invitrogen), according to the manufacturer's instructions. Total RNA (0.5-1.0 .mu.g) was primed with random hexamers or oligo(dT) and reverse-transcribed with MultiScribe Reverse Transcriptase (Applied Biosystems). Quantitative PCR were performed in triplicate with 0.3 .mu.mol/l gene-specific primer pairs (nLacZ5'F/5'R, nLacZ 3'F/3'R, cyclinG1F/R and EGFP-F/EGFP-R) using the GoTaq qPCR master mix in a StepOne Plus Real-time PCR device. The specificity of qRT-PCR products derived from the 5' and 3' ends of nlacZ mRNA was confirmed by gel electrophoresis.
[0201] Northern Blot Analysis.
[0202] Total RNA was extracted from mouse liver and analyzed by Northern hybridization. To detect nLacZ mRNA, a 618 bp fragment of nLacZ cDNA was isolated by NcoI and PciI digestion of pAAVCBnLacZ and labeled with .alpha.-.sup.32P dCTP by random priming (Takara, Shiga, Japan). To detect 3' fragments of the cleaved nLacZ mRNA, an 111 bp fragment of the poly(A) sequence in the vector genome was cloned into pCR4-TOPO (Invitrogen) for preparation of antisense RNA probe labeled with .alpha.-.sup.32P CTP during in vitro transcription using the Riboprobe System T7 kit (Promega). To detect miR-122, miR-26a, miR-22, and let-7 or U6 in total liver RNA, small RNAs were resolved by denaturing 15% polyacrylamide gels, transferred to Hybond N+ membrane (Amersham BioSciences, Pittsburgh, Pa.), and crosslinked with 254 nm light (Stratagene, La Jolla, Calif.). Synthetic oligonucleotides, 5' end-labeled with .gamma.-.sup.32P ATP using T4 polynucleotide kinase (New England Biolabs, Beverly, Mass.), were used as DNA probes (Table 4) and hybridized in Church buffer (0.5 mol/l NaHPO.sub.4, pH 7.2, 1 mmol/l EDTA, 7% (w/v) sodium dodecyl sulphate) at 37.degree. C. Membranes were washed using 1.times.SSC (150 mM sodium chloride, 15 mM sodium citrate), 0.1% sodium dodecyl sulphate buffer, and then visualized using an FLA-5100 Imager (Fujifilm, Tokyo, Japan).
[0203] Western Blot Analysis.
[0204] Proteins were extracted with radioimmunoprecipitation assay buffer [25 mmol/l Tris-HCl, pH 7.6, 150 mmol/l NaCl, 1% (vol/vol) NP-40, 1% (wt/vol) sodium deoxycholate, 0.1% (w/v) sodium dodecyl sulphate] containing a protease inhibitor mixture (Boston BP, Boston, Mass.). Protein concentration was determined using the Bradford method (Bio-Rad, Melville, N.Y.). Protein samples, 50 ug each, were loaded onto 12% polyacrylamide gels, electrophoresed, and transferred to nitrocellulose membrane (Amersham BioSciences). Briefly, membranes were blocked with blocking buffer (LI-COR Biosciences, Lincoln, Nebr.) at room temperature for 2 hours, followed by incubation with either anti-GAPDH (Millipore, Billerica, Mass.), anti-cyclin GI (Santa Cruz Biotechnology, Santa Cruz, Calif.) or anti-calmodulin (Millipore) for 2 hours at room temperature. After three washes with PBS containing 0.1% (vol/vol) Tween-20, membranes were incubated with secondary antibodies conjugated to LI-COR IRDye for 1 hour at room temperature, and then antibodies detected using the Odyssey Imager (LI-COR).
[0205] .beta.-Galactosidase Assay.
[0206] Proteins were extracted with radioimmunoprecipitation assay buffer and quantified as described above. Fifty micrograms of protein was used for each .beta.-galactosidase assay using the Galacto-Star System (Applied Biosystems), according to the manufacturer's instructions.
[0207] 5' RACE.
[0208] 5' RACE was performed as described. The 5' RACE Outer Primer and the nLacZ gene-specific primer bGHpolyAR (Table 4) were used for the first round of nested PCR. The 5' RACE Inner Primer and the nLacZ gene-specific primer nLacZpolyR, which is located near the stop codon of nLacZ cDNA, were used for the second round of nested PCR (Table 4). PCR products were TOPO-cloned into pCR-4.0 (Invitrogen) and sequenced.
[0209] Statistical Analysis.
[0210] All results are reported as mean.+-.SD and compared between groups using the two-tailed Student's t-test.
TABLE-US-00003 TABLE 3 Summary of microRNA-guided transgene mRNA cleavage in mouse liver and heart Cleavage site Between Between Between Random miR BS cleavage Position 10 and 11 nt 17 and 18 nt 18 and 19 nt site Liver 1 Copy of miR-122 BS (21 clones) 1 17/21 81% ND ND 19% 3 Copies of miR-122 BS (11 clones) 1 ND 100% ND ND 0% 2 4/11 3 7/11 3 Copies each of miR-1 and miR 122 1 ND ND ND ND 0% miR 1 3 .times. BS BS in a single vector (21 clones) 2 ND 3 ND miR-122 3 .times. BS 1 1/21 95% ND ND 5% 2 10/21 3 9/21 Heart 1 Copy of miR-1BS (12 clones) 1 12/12 100% ND ND 0% 3 Copies of miR 1BS (21 clones) 1 ND 80% 4/21 20% ND 0% 2 16/21 ND 3 1/21 ND 3 Copies each of miR 1 and miR 122 1 ND ND ND 1/22 14% 4% miR-122 3 .times. BS BS in a single vector (22 clones) 2 ND 1/22 3 ND ND miR 1 3 .times. BS 1 1/22 73% ND 9% ND 0% 2 7/22 1/22 3 8/22 1/22
TABLE-US-00004 TABLE 4 Oligonucleotide primers and probes used in Example 8. Oligo nucleotides Sequence SEQ ID NO (miR-1).sub.1 sense [PHOS]CGAAATACATACTTCTTTACATTCCATT SEQ ID NO: 32 (miR-1).sub.1 anti-sense [PHOS]CGAATGGAATGTAAAGAAGTATGTATTT SEQ ID NO: 33 (miR-122).sub.1 sense [PHOS]CGAAACAAACACCATTGTCACACTCCATT SEQ ID NO: 34 (miR-122).sub.1 anti-sense [PHOS]CGAATGGAGTGTGACAATGGTGTTTGTTT SEQ ID NO: 35 (miR-1).sub.3 sense [PHOS]CGAAATACATACTTCTTTACATTCCAATA SEQ ID NO: 36 CATACTTCTTTACATTCCAATACATACTTCTTTAC ATTCCATT (miR-1).sub.3 anti-sense [PHOS]CGAATGGAATGTAAAGAAGTATGTATTGG SEQ ID NO: 37 AATGTAAAGAAGTATGTATTGGAATGTAAAGAAGT ATGTATTT (miR-122).sub.3 sense [PHOS]CGAAACAAACACCATTGTCACACTCCAAC SEQ ID NO: 38 AAACACCATTGTCACACTCCAACAAACACCATTGT CACACTCCATT (miR-122).sub.3 anti-sense [PHOS]CGAATGGAGTGTGACAATGGTGTTTGTTG SEQ ID NO: 39 GAGTGTGACAATGGTGTTTGTTGGAGTGTGACAAT GGTGTTTGTTT (miR-1).sub.1-(miR-122).sub.1 [PHOS]CGAAATACATACTTCTTTACATTCCAACA SEQ ID NO: 40 sense AACACCATTGTCACACTCCATT (miR-1).sub.1-(miR-122).sub.1 [PHOS]CGAATGGAGTGTGACAATGGTGTTTGTTG SEQ ID NO: 41 anti-sense GAATGTAAAGAAGTATGTATTT Synthesized (miR-1).sub.3- TTCGAACTCGAGATACATACTTCTTTACATTCCAA SEQ ID NO: 42 (miR-122).sub.3 fragment TACATACTTCTTTACATTCCAATACATACTTCTTT ACATTCCACCATGGACTAGTACAAACACCATTGTC ACACTCCAACAAACACCATTGTCACACTCCAACAA ACACCATTGTCACACTCCAGCGGCCGCTTCGAA Pri-miR-122F ATCGGGCCCGACTGCAGTTTCAGCGTTTG SEQ ID NO: 43 Pri-miR-122R CGCGGGCCCGACTTTACATTACACACAAT SEQ ID NO: 44 Pri-miR-1F CGCGGGCCCGACTGATGTGTGAGAGAGAC SEQ ID NO: 45 Pri-miR-1R CGCGGGCCCGACTTTCGGCCTCCCGAGGC SEQ ID NO: 46 nLacZ5 F(5 F) TGAAGCTGAAGCCTGTGATG SEQ ID NO: 47 nLacZ5 R(5 R) GAGCACCTGACAGCATTGAA SEQ ID NO: 48 nLacZ3 F(3 F) CTCAGCAACAGCTCATGGAA SEQ ID NO: 49 nLacZ3 R(3 R) TTACTTCTGGCACCACACCA SEQ ID NO: 50 nLacZpolyF(A .sup.+F) TGGTGTGGTGCCAGAAGTAA SEQ ID NO: 51 nLacZpolyR(A .sup.+R) CAACAGATGGCTGGCAACTA SEQ ID NO: 52 bGHpolyAR(bGH.sup.+ AR) TGGGAGTGGCACCTTCCA SEQ ID NO: 53 EGFP-F CGACCACTACCAGCAGAACA SEQ ID NO: 54 EGFP-R CTTGTACAGCTCGTCCATGC SEQ ID NO: 55 CyclinG1F AATGGCCTCAGAATGACTGC SEQ ID NO: 56 CyclinG1R AGTCGCTTTCACAGCCAAAT SEQ ID NO: 57 MM-ActinF ATGCCAACACAGTGCTGTCTGG SEQ ID NO: 58 MM-ActinR TGCTTGCTGATCCACATCTGCT SEQ ID NO: 59 miR-122 probe TGGAGTGTGACAATGGTGTTTG SEQ ID NO: 60 Let-7 probe AACTATACAACCTACTACCTCA SEQ ID NO: 61 miR-26a probe AGCCTATCCTGGATTACTTGAA SEQ ID NO: 62 miR-22 Probe ACA GTT CTT CAA CTG GCA GCTT SEQ ID NO: 63 U6 probe CTCTGTATCGTTCCAATTTTAGTATA SEQ ID NO: 64
Example 9: Intravenous Injection of rAAVs Mediated Widespread Transduction in Neonatal Mouse CNS
Introduction to the Example
[0211] This example describes an analysis of nine scAAV vectors for CNS gene transfer properties after systemic administration. This study involved identifying more effective vectors for the CNS gene transfer, In some aspects the study examined serotypes or natural variants of rAAVs for enhanced-permeation of the BBB. In some cases, the study sought to identify rAAV vectors with improved delivery of enhanced green fluorescent protein (EGFP) to the CNS following facial vein injection on postnatal day 1 (P1). AAV9 was included in the study. Except for rAAV2 and rAAV5, all other 7 vectors crossed the BBB with varied transduction efficiency, among which rAAVrh.10, rAAVrh.39, rAAVrh.43, rAAV9 and rhAAV7 rank in the top 5, mediating robust EGFP expression in both neuronal and glial cells throughout the CNS in this study. The performance of rAAVrh.10 was comparable to that of rAAV9 and in some case better. Several rAAVs efficiently transduce neurons, motor neurons, astrocytes and Purkinje cells; among them, rAAVrh.10 is at least as efficient as rAAV9 in many of the regions examined. Intravenously delivered rAAVs did not cause abnormal microgliosis in the CNS. The rAAVs that achieve stable widespread gene transfer in the CNS are useful as therapeutic vectors for neurological disorders affecting large regions of the CNS as well as convenient biological tools for neuroscience research.
Results
[0212] Twenty one days after vector administration in P1 mice, the CNS transduction profiles of the following recombinant AAV vectors encoding EGFP: rAAV1, rAAV2, rAAV5, rAAV6, rAAV6.2, rAAV7, rAAV9, rAAVrh.10, rAAVrh.39 and rhAAVrh.43 were compared. The vectors used in this study were comparable in purity and morphological integrity (FIGS. 19A to 19D). As assessed by the scoring system described in the methods, rAAV9 was among the top performers; most other rAAVs tested (rAAV1, rAAV6, rAAV6.2, rAAV7, rAAVrh.10, rAAVrh.39 and rAAVrh.43) also gave rise to EGFP expression throughout the CNS (Table 2). The number of apparent EGFP positive cells (Table 5) among sub-anatomical structures was influenced by the particular vector used. For these seven rAAVs, and rAAV9 (total of eight rAAVs), that permeated the BBB and accomplished CNS transduction after i.v. delivery, EGFP positive cells were found in hypothalamus followed by medulla, striatum, hippocampus, cortex and cerebellum. In contrast, the transduction efficiency in olfactory bulb and thalamus was relatively low (Table 5). A quantitative assessment of EGFP gene transfer efficiency was made of each rAAV. 12 sub-anatomically and functionally important regions in the brain were selected for quantitative analysis of the mean EGFP intensity/pixel in each region for each rAAV by using Nikon NIS elements AR software V. 32 (FIGS. 19A to 19C) (see Methods). For the eight vectors that achieved CNS transduction after i.v. injection, the mean EGFP intensity/pixel was relatively low in cortex, habenular nucleus, cornu ammonis, dentate gyrus, thalamus, cerebellum and olfactory bulb, moderate in choroid plexus and caudate-putamen, but high in hypothalamus, medulla and amygdale (FIGS. 19A to 19C). The average EGFP intensities of all 12 regions for different rAAVs were compared in FIG. 19D. AAVrh.10, AAVrh.39 and AAVrh.43 were noted for gene transduction efficiency in brain, followed by AAV7, AAV9, and AAV1 (FIGS. 19A to 19D). Those eight effective serotypes also mediated EGFP expression throughout the spinal cord, to different degrees. The same quantitative analysis was performed for each rAAV in the cervical, thoracic and lumbar sections of the spinal cord (FIGS. 19A to 19C); the average EGFP intensities of the three sections for different rAAVs were also compared (FIG. 19D). AAV1, AAV9, AAVrh10, AAV.rh39 and AAV.rh43 displayed strong transduction in the spinal cord with the high EGFP intensity observed in the cervix, followed by thoracic and lumbar sections of the spinal cord (FIGS. 19A to 19D). For rAAV2 there were few EGFP-positive cells in hippocampus, cortex and hypothalamus. EGFP-positive cells were observed in the hypothalamus in AAV5-injected mice. A description of the observations made in different CNS structures is provided below. The subanatomic CNS structures may serve as a target for CNS gene therapy. In some cases, the subanatomic CNS structures are associated with pathological changes in one or more neurological disorders. In some cases, the subanatomic CNS structure have distinct transduction profiles for one or more rAAVs.
[0213] Striatum.
[0214] Pathology of the striatum is associated with Huntington's disease, choreas, choreoathetosis, and dyskinesias. Addiction may involve plasticity at striatal synapses. Systemic injection of rAAV9 in neonatal mice tranduces striatal tissue. In this study, a large number of cells with neuronal morphology in this region were also transduced by rAAVrh.10 (FIG. 20), which was confirmed by co-staining with a neuronal marker as described below. Other vectors, including rAAVrh.39 and rAAV7, also mediated moderate transduction in striatum (FIG. 20). In contrast, rAAV6, rAAV6.2, and rAAV1 resulted in relatively lower EGFP expression in this structure (FIG. 20).
[0215] Hippocampus.
[0216] The hippocampus is a region associated with long-term memory and spatial navigation, which is usually damaged by stress and pathogenesis of diseases such as epilepsy and Schizophrenia. Large numbers of EGFP-positive neurons were observed bilaterally in all regions of the hippocampus, namely dentate gyrus, hilus, CA1, CA2 and CA3 for the mice received intravenous rAAVrh.10, rAAV9, rAAV7, rAAVrh.39, and rAAVrh.43 (ranked by transduction efficiency in this structure, Table 5 and FIGS. 19A-19D and FIG. 20). In addition to the neuronal transduction pattern, EGFP-positive cells had morphologic appearance of astrocytes (FIG. 20). This was further confirmed by double staining with antibodies against EGFP and astrocytic marker as described below. For intravenously delivered rAAV1, rAAV6 and rAAV6.2 vectors there were small numbers of EGFP-positive cells in the hippocampus (FIG. 20).
[0217] Cortex.
[0218] Pathological changes in the cortex have been implicated in Alzheimer's and Parkinson's diseases. AAV7, AAV9, AAVrh.10, AAVrh.39 and AAVrh.43 vectors achieved moderate EGFP transduction in cortex (Table 5 and FIGS. 19A to 19D and 20). The morphology of transduced cells was consistent with both neurons and astrocytes as further confirmed by cellular marker staining and confocal microscopic analysis described below. Prominent EGFP-positive cells were typically observed in the ventrolateral regions of the cortex, including posterior agranular insular cortex, piriform cortex, lateral entorhinal cortex, posterolateral cortical amygdaloid nucleus and posteromedial cortical amygdaloid nucleus (FIG. 20). Strong EGFP signals spread from +1.5 to -3.3 mm in relation to the Bregma (0.0 mm). The cortical transduction efficiency of rAAVrh.10, rAAV9, rAAVrh.39 and rAAVrh.43 was comparable (Table 5 and FIGS. 19A to 19D and 20). AAV1, AAV6 and AAV6.2 vectors also transduced cells in the cortex (FIG. 20).
[0219] Hypothalamus.
[0220] A role for the hypothalamus is to secret neurohormones to control certain metabolic processes. The hypothalamus is also indicated in the etiology of diabetes. EGFP signal was observed in the hypothalamus for eight vectors. Intravenous administration of rAAVrh.10 resulted in the highest EGFP expression in the entire hypothalamus, followed by rAAVrh.39, rAAV7, rAAV6.2, rAAVrh.43, rAAV9, rAAV1 and rAAV6 (FIGS. 19A to 19D and 20 and Table 5). Interestingly most EGFP-positive cells in this structure have an astrocytic morphology which was ascertained by immunostaining for an astrocytic cell type specific marker as described below. The astrocytic EGFP signal tended to obscure direct examination of morphological details of other transduced cells. However, this was clarified by double immunofluorescent staining of tissue sections with antibodies for EGFP and neuronal cell markers as described below.
[0221] Cerebellum.
[0222] The pathological lesions in cerebellum are often found in diseases such as cerebellar-cognitive affective syndrome, developmental coordination disorder, posterior fossa syndrome, linguistic deficits, aging, attention deficit hyperreactivity disorder, autism, dementia and schizophrenia. EGFP-positive cells and fibers were detected in cerebellum for most rAAV vectors (Table 5 and FIGS. 19A to 19D and 20). A large number of EGFP-expressing cells were found in the Purkinje and granule cell layers for rAAV7, rAAV9, rAAVrh.10, rAAVrh.39 and rAAVrh.43 (FIG. 20). The transduction profile of rAAV1 vector indicated expression in cells in the granule cell layer, while rAAV6 and rAAV6.2 were localized in cells in the Purkinje cell layer (FIG. 20).
[0223] Medulla.
[0224] The medulla is a potential gene therapy target for treating chronic pain. Most rAAVs mediated moderate to robust EGFP expression in medulla with most green cells being present in the outer rim (FIG. 20). Transduction efficiencies of these rAAV in this region are ranked in the following order: rAAVrh.39=rAAVrh.43>rAAV.rh10>rAAV1>rAAV9>rAAV7>rAAV6.2>- ;rAAV6 (Table 5 and FIG. 19A). The morphology of most EGFP-transduced cells was consistent with the cells being astrocytes.
[0225] Spinal Cord.
[0226] The spinal cord is involved with motor neurons diseases. rAAVrh.10, rAAV9, rAAVrh.39 and rAAVrh.43 gave rise to very robust EGFP expression in cervical gray and white matter, while rAAV1, rAAV6.2 and rAAV7 showed moderate EGFP intensity (Table 5 and FIGS. 19A to 19D and 21). For rAAV1 the EGFP signal was observed in white matter. The transduction ability of all effective rAAVs decreased from cervical to lumbar spinal cord. EGFP-positive cells were visible in the latter region. Large populations of EGFP-positive cells with astrocytic morphology were observed throughout the spinal cord (FIG. 21). In addition, rAAVrh.10, rAAV9, rAAVrh.39, rAAVrh.43 and rAAV7 also transduced cells with motor neuron morphology in the ventral regions of spinal cord (FIG. 21). Ascending dorsal column fibers showed clear EGFP signal. In addition, dorsal root ganglia (DRG) displayed remarkable transduction with strong EGFP expression in DRG neurons (FIG. 22 and FIG. 26). The identities of rAAV transduced cell types in the spinal cord were characterized by co-immunofluorescence staining with antibodies against EGFP and cell type specific markers as described below.
IV Administration of AAV Vectors Leads to Transduction of Different Cell Types in the CNS
[0227] To confirm the identity of transduced cells in different regions of the CNS, double immunofluorescent staining was performed with antibodies for EGFP and NeuN (generic neuronal marker), glial fibrillary acid protein (GFAP; astrocyte marker), calbindin-D28K (Purkinje cell marker), and choline acetyl transferase (ChAT; motor neuron marker) (FIG. 23). The immunostaining results showed that a large number of NeuN positive cells expressed EGFP throughout the mouse brain, which indicated widespread neuronal transduction. The regions with high density of transduced neurons included striatum, hippocampus, cortex and hypothalamus. rAAVrh.10, rAAV9, rAAV7 and rAAVrh.39 vectors were efficient in mediating neuronal transduction, followed by AAV6.2, AAV1 and AAV6 (FIGS. 19A to 19D and 23). In addition, dopaminergic neurons in substantia nigra were transduced by AAV.rh10 (FIG. 23). Transduced cells in the CNS included GFAP-positive astrocytes with small cell bodies and highly ramified processes (FIG. 23). The calbindin-D28K immunostaining confirmed the identity of a number of transduced cells in the cerebellum as Purkinje cells, with EGFP expression in both cell body and their tree-like processes (FIG. 23). The rAAVs proficient in transducing Purkinje cells include: rAAVrh.10, rAAV9, rAAVrh.39, rAAV7, rAAV6.2 and rAAVrh.43. rAAV1 and rAAV6 transduced a portion of Purkinje cells with relatively low EGFP intensity (FIGS. 19A to 19D). Transduction of motor neurons was confirmed by the presence of large EGFP+/ChAT+ cells in the ventral spinal cord for several rAAV vectors (FIG. 23). rAAVrh.10, rAAV9, rAAV7, rAAVrh.39 showed comparable efficiency transduction of motor neurons (FIG. 21).
IV Administration of AAV Vectors Mediated Robust Transduction in Ventricles and Brain Blood Vessels
[0228] EGFP expression was observed in the choroid plexus cells in lateral, 3.sup.rd and 4.sup.th ventricles of the animals infused with rAAVrh.39, rAAVrh.10, rAAVrh.43, rAAV7 and rAAV9 (ranked by transduction efficiency, Table 5 and FIGS. 19A to 19D and 24). EGFP expression in different ventricles of the same mouse brain was similar (FIG. 24). Ependymal cells lining the ventricles were also transduced. An observation regarding the distribution of EGFP-positive cells was the apparent gradient with the highest number of transduced cells in peri-ventricular regions and progressively lower numbers with increasing distance to the ventricles. This was apparent in areas around the 3.sup.rd and 4.sup.th ventricles than the lateral ventricles (FIG. 24). Extensive EGFP signal was also found with blood vessels throughout mouse brain and spinal cord. This was verified by dual immunofluorescent staining with antibodies directed to EGFP and a blood vessel endothelium specific marker, CD34 (FIGS. 27A and 27B). Unlike the rAAV transduction profiles in different regions of the brain parenchyma, the EGFP transduction of the blood vessels throughout the CNS was relatively uniform for any given vector. However, transduction of blood vessels was influenced by the particular rAAV used. A majority of rAAVs mediated moderate (e.g., rAAV6) to highly efficient (e.g. rAAVrh.10) blood vessel transduction in the CNS.
IV Injection of AAV Vectors Did not Cause Microgliosis
[0229] Brain sections were also stained with antibody against Iba-1 to label microgial cells. The Iba-1-positive cells in the sections from mice received rAAVrh.10 was no more than those in naive or PBS-injected mice (FIG. 28). This result indicated that intravascularly delivered rAAVs do not cause sustained inflammation in the CNS of mice 3 weeks after the injection of P1 neonates.
Discussion of Results
[0230] In this study, the CNS transduction profile was evaluated for 10 different rAAV vectors delivered by intravascular infusion in neonatal mice. Most of the rAAVs can cross the BBB and mediate gene transfer to the neonatal mouse CNS with varying degrees of efficiency (FIGS. 19A to 19D, 20 and 21 and Table 5). After systemic administration, rAAVrh.10, rAAVrh.39, rAAVrh.43, and rAAV9 are the effective rAAVs with similar transduction capabilities and cellular tropism, as assessed by overall EGFP expression in the CNS. Specifically, a number of regions in the mouse CNS, including striatum, hippocampus, cortex, hypothalamus, cerebellum, medulla, and cervical spinal cord, revealed substantial EGFP expression. In addition, rAAV6.2 and rAAV7 were also effective. AAV1 and AAV6, achieved CNS transduction (Table 5). Native EGFP expression was detectable in brain and spinal cord sections for most of the rAAVs without immunostaining (FIG. 29).
[0231] This example has clinical significance for gene therapy of CNS-related disorders, including for young patients. For a variety of neurological diseases, early treatment during infancy may be necessary to prevent irreversible CNS injury. The capacity of rAAVs to transduce large numbers of neuronal cells in different regions is relevant for treating neurological diseases such as spinal muscular atrophies, neuronal ceroid lipofuscinoses, and spinocerebellar degenerations. The efficiency of some rAAV vectors in transducing Purkinje and granule layer cells indicates that the vectors may be used for treating spinocerebellar ataxias. Transduction of astrocytes by rAAVs expressing secreted neurotrophic factors may be also beneficial for a number of neurodegenerative diseases such as Canavan's disease and amyotrophic lateral sclerosis. The vascular transduction in the CNS may be relevant for treating brain ischemia and stroke. The clinical application of intravascular rAAV-mediated gene delivery may also extend to the peripheral nervous system (PNS). Efficient transduction of DRG provides new therapeutic strategies for patients suffering from chronic pain.
[0232] Systemic gene delivery to the CNS is also useful as a method to manipulate gene expression in research. Effective and stable transgene expression in the CNS by intravenous administration of rAAVs may be applied to establish somatic transgenic animal models, which is a potentially cheaper, faster and simpler method than conventional transgenesis. Somatic CNS gene knock-down animal models may also be created using the method described herein.
[0233] Some rAAVs indeed demonstrated unique transduction profiles in the CNS. For instance, rAAV1 displayed transduced granule cells in the cerebellum, while rAAV6 and rAAV6.2 transduced Purkinje cells, and others transduced both types of cells (FIG. 9). This indicates that once across the BBB, the rAAVs have distinct tropisms, which can be attributed to the capsid because that the vector genome used in all vectors was the same.
[0234] AAV serotypes disclosed herein can efficiently transduce brain capillary endothelial cells, neurons and astrocytes. This indicates that these vectors may extravasate from the circulation and reach the CNS parenchyma, possibly by crossing the BBB. AAV may cross the endothelial barrier by a transcytosis pathway. In this study, choroid plexuses and their surrounding parenchymal tissue were efficiently transduced. In addition, there was an apparent gradient of EGFP intensity from peri-ventricular (higher) to deep parenchymal (lower) tissue. These observations indicate that AAV may enter the neonatal mouse CNS through the choroid plexus, followed by widespread distribution via CSF and/or interstitial fluid flow to transduce neuronal and glial cells.
[0235] Neuronal- or glial-specific promoters, such as synapsin-1, and GFAP promoters may be used to restrict gene expression to a specific cell type. A further method to achieve targeted CNS gene delivery is to utilize RNA interference to detarget the peripheral tissues by post-transcriptional regulatory mechanisms. By adding microRNA binding sites into the 3' end of the transgene cassettes, transgene expression after systemic administration of AAV vectors may be reduced or eliminated in tissues such as liver, heart and skeletal muscle, while maintaining CNS transduction.
Materials And Methods
[0236] AAV Production
[0237] ScAAV vectors were produced by trans-encapsidation of rAAV vector genome flanking by inverted terminal repeats (ITRs) from AAV2 with the capsids of different AAVs using the method transient transfection of 293 cells and CsCl gradient sedimentation as previously described. Vector preparations were titered by quantitative PCR. Purity of vectors was assessed by 4-12% SDS-acrylamide gel electrophoresis and silver staining (Invitrogen, Carlsbad, Calif.). Morphological integrity of each vector used in the study was examined by transmission electron microscopy of negative stained recombinant AAV virions. The expression of EGFP in the scAAV vector genome is directed by a hybrid CMV enhancer/chicken .beta.-actin promoter.
Neonatal Mouse Injections
[0238] Wild-type C57BL/6 mice littermates were used. Mice breeding were conducted using programmatic timing method. Pregnant mice were monitored daily from embryonic day 17 to 21 to ensure the newborn pups could be dosed with vectors on P1. The mother (singly housed) of each litter to be injected was removed from the cage. Vectors were diluted to concentration of 4.times.10.sup.12 GCs/mL in PBS and 100 .mu.l of solution was subsequently drawn into 31G insulin syringes (BD Ultra-Fine II U-100 Insulin Syringes). P1 pups of C57BL/6 mice were anesthetized using isoflurane and rested on ice. For intravenous injections, a dissection microscope was used to visualize the temporal vein (located just anterior to the ear). The needle was inserted into the vein and the plunger was manually depressed. Correct injection was verified by noting blanching of the vein. Each pup received 4.times.10.sup.11 GCs of different scrAAVCBEGFP vectors (rAAV1, rAAV2, rAAV5, rAAV6, rAAV6.2, rAAV7, rAAV9, rAAVrh.10, rAAVrh.39, rAAVrh.43; n=6-8 mice per group) via the superficial temporal vein. After the injection pups were carefully cleaned, rubbed with their original bedding, and then returned to their original cage. The mother was then reintroduced to the cage after brief nose numbing using ethanol pads.
Histological Processing
[0239] The study animals were anesthetized 21 days post-injection, then transcardially perfused with 15 mL of cold PBS followed by 15 mL of fixation solution containing 4% paraformaldehyde (v/v) with 0.2% of glutaraldehyde (v/v) in PBS. Then the whole carcasses were post-fixed in fixation solution for 5 days. Spinal cords and brains were extracted under a bright-field dissecting microscope, rinsed in PBS, and then cryoprotected in 30% sucrose (w/v) in PBS at 4.degree. C. Once the tissues sank to the bottom of the sucrose solution, they were embedded in Tissue-Tek OCT compound (Sakura Finetek, Torrance, Calif.) and frozen in a dry ice/ethanol bath. The tissue blocks were stored at -80.degree. C. until sectioning. Serial 40 .mu.m floating sections of the entire brain were cut in a Cryostat (Thermo Microm HM 550). For the spinal cord, 3 mm length sections were taken from cervical, thoracic and lumbar regions, and then serial 40 .mu.m transverse sections prepared as above.
Immunostaining and Microscopy Imaging Analysis
[0240] Brain and spinal cord sections were stained as floating sections in 12-well plates. Sections were washed 3 times in PBS for 5 min each time, and then incubated in blocking solution containing 1% Triton-X100 (v/v) (Fisher, Pittsburgh, Pa.), 5% dry-milk (w/v) and 10% goat serum (v/v) (Invitrogen) for 2 h at room temperature. Then the sections were incubated with primary antibodies diluted in blocking solution at 4.degree. C. overnight. The following day tissue sections were washed twice in 0.05% Tween-20 (v/v) in PBS (PBST) and once with PBS, with each washing step lasting 10 min. Afterwards sections were incubated with appropriate secondary antibodies in blocking solution at room temperature for 2 h. Sections were washed again as above before mounting on glass slides. Vectashield with DAPI (Vector Laboratories, Burlingame, Calif.) was used to coverslip all slides, and then they were analyzed using a fluorescent inverted microscope (Nikon Eclipse Ti) or a Leica TSC-SP2 AOBS confocal microscope equipped with a 63.times. oil lens and a DM-IRE2 inverted microscope. The primary antibodies used in this study were as follows: rabbit anti-GFP (Invitrogen), goat anti-ChAT and mouse anti-NeuN (both from Millipore, Billerica, Mass.), mouse anti-GFAP (Cell signaling, Danvers, Mass.), rat anti-CD34 (Abcam, Cambridge, Mass.), mouse anti-Calbindin D-28k (Sigma, St Louis, Mo.) and rabbit anti-DARPP (Abcam, Cambridge, Mass.). The secondary antibodies used in the study included: DyLight 488 AffiniPure Donkey Anti-rabbit IgG (Jackson ImmunoResearch, West Grove, Pa.); DyLight 549 AffiniPure Donkey Anti-Goat IgG (Jackson ImmunoResearch); DyLight 549 Affinipure Goat Anti-Rat IgG (Jackson ImmunoResearch); DyLight 594 AffiniPure Goat Anti-Mouse IgG (Jackson ImmunoResearch); goat anti-rabbit IgG-Alexa fluro 488 (Invitrogen) and goat anti-mouse IgG-Alexa fluro 568 (Invitrogen).
Semi-Quantitative and Quantitative Comparison of EGFP Transduction by Different Vectors
[0241] To generate a quantifiable and comparable data format, a semi-quantitative scoring system was develop to estimate transduction efficiency of different rAAV vectors in different regions of the mouse CNS. Briefly, regions with no EGFP positive cells were marked as (-). Regions with very few EGFP positive cells were scored (+), regions with some EGFP positive cells were ranked as (++), regions with many EGFP positive cells were marked as (+++). Finally, regions filled with EGFP positive cells were marked as (++++).
[0242] Next, 12 sub-anatomically and functionally important regions in the brain as well as cervical, thoracic and lumbar sections of the spinal cord were selected for quantitative analysis of images that were taken on a Nikon Eclipse Ti inverted microscope equipped with a Retiga 2000-RV CCD cooled camera. Nikon NIS elements AR software v. 3.2 was used for intensity quantification. Prior to quantification, optimal light source intensity and exposure times were obtained by plotting an intensity/exposure time curve using fluorescence reference slides (Ted Pella, prod. 2273). It was found that the intensity and exposure times had linear correlation. In addition, overexposure and extreme underexposure distorts the linear correlation. The maximum intensity (ND1) and a 20 ms exposure were used for all sections to avoid overexposure. For quantification, fixed region of interest (ROI) was used to quantify the brightest area of any given brain region. A mean intensity (total intensity/size of ROI) was obtained for each region of all serotypes.
TABLE-US-00005 TABLE 5 Transduction characteristics of AAV serotypes following intravascular injections into neonatal mouse brain score n score n score n score n score n score n Olfactory Bulb Striatum Hippocampus Cortex Thalamus Hypothalamus AAV1 + 3 ++ 3 ++ 3 ++ 3 + 3 +++ 3 AAV2 - 3 - 3 + 3 + 3 + 3 + 3 AAV5 - 3 - 3 - 3 + 3 - 3 + 3 AAV6 + 3 + 3 ++ 3 ++ 3 + 3 +++ 2 ++ 1 AAV6.2 - 3 +++ 2 ++ 3 ++ 3 + 3 ++++ 3 ++ 1 AAV7 +++ 1 +++ 3 +++ 2 ++ 3 + 3 ++++ 3 ++ 2 ++ 1 AAV9 +++ 2 +++ 3 ++ 3 +++ 1 + 3 ++++ 1 ++ 1 ++ 2 +++ 2 rh10 +++ 1 ++++ 1 +++ 3 +++ 2 ++ 2 ++++ 3 ++ 2 ++ 2 ++ 1 + 1 rh39 +++ 1 ++++ 2 +++ 3 +++ 1 + 3 ++++ 3 ++ 2 +++ 1 ++ 2 rh43 ++ 3 +++ 3 +++ 3 +++ 3 + 3 ++++ 1 +++ 2 Choroid Cerebellum Medulla Cervical Thoracic Lumber Plexus AAV1 +++ 3 +++ 3 +++ 3 +++ 3 + 3 +++ 3 AAV2 + 3 + 3 + 3 - 3 - 3 ++ 3 AAV5 - 3 - 3 - 3 - 3 - 3 - 3 AAV6 ++ 3 ++ 3 ++ 3 + 3 + 3 +++ 3 AAV6.2 ++ 3 ++ 3 ++ 3 ++ 3 + 3 ++++ 3 AAV7 +++ 1 ++ 3 ++ 3 + 3 + 3 ++++ 3 ++ 2 AAV9 +++ 1 ++ 3 ++++ 1 ++ 3 + 3 ++++ 3 ++ 2 +++ 2 rh10 +++ 1 +++ 3 ++++ 1 ++ 3 + 3 ++++ 3 ++ 2 +++ 2 rh39 +++ 1 ++++ 1 ++++ 1 +++ 3 + 3 ++++ 3 ++ 2 +++ 2 +++ 2 rh43 ++ 3 ++++ 1 ++++ 2 +++ 3 + 3 ++++ 3 +++ 2 +++ 1 Scoring: (-) no transduction, (+) very few positive cells (++) some positive cells, (+++) many positive cells, and (++++) region is almost saturated with EGFP-positive cells. The number of animals (n) with the particular score is given to the right of the score.
Example 10: Evaluation of an rAAV Based Treatment in a Canavan Disease Model Introduction to the Example
[0243] CD is a rare and fatal childhood leukodystrophy caused by autosomal recessive mutations in the aspartoacylase gene (ASPA) [as established by G.G.'s graduate work (12)]. ASPA deficiency in CD patients leads to elevated N-Acetyl-Aspartic Acid (NAA) in urine (a hallmark of CD) and spongy degeneration of white matter throughout the CNS, producing severe psychomotor retardation and early death. An ASPA.sup.-/- mouse model mimics the neuropathology and clinical manifestations seen in CD patients, i.e., spongy degeneration of white matter, motor deficits, developmental delays, and early death (within 3 weeks after birth).
[0244] In this study, i.v. deliverable rAAVs were used to target the CNS globally to treat diffused WM degeneration in CD mice. Single i.v. injections of ASPA vector to the neonatal CD mice corrected metabolic defect, psychomotor malfunction and other disease phenotypes, and prolonged survival. While untreated CD mice started showing growth retardation, psychomotor malfunction in the 2.sup.nd wk after birth and uniformly died soon after weaning, the treated mice began to gain weight 2 wks after vector injection and nearly caught up with their heterozygous littermates within 7-8 weeks. Unlike CD mice, the mobility of the treated animals was similar to Wt littermates. Data from rotarod test on the treated mice showed no significant differences in the latency time among the treated CD mice and their age-matched Wt littermates, indicating that gene therapy corrected the ataxia, a typical neuromuscular symptom of CD. Biochemical characterization indicated reduction of NAA levels in the urine samples and restoration of ASPA activity in their brain and kidney tissues. Mitigation of the biochemical and clinical phenotypes was well correlated with globally ameliorated histopathology in not only the brain, spinal cord but also in the peripheral tissues such as kidney, indicating that CD is not just a CNS disorder.
Results
[0245] In CD mice were dosed at P1 (facial vein, 4.times.10.sup.11 GCs) with AAV9ASPA. The mice were monitored for growth, gait, motor function on rotarod, NAA levels in urine and ASPA activities in brain. The results showed that i) Untreated CD mice started losing weight at the 2.sup.nd week and died in the 3.sup.rd week after birth; ii) The treated animals recovered their capacity to grow in the 5.sup.th week and caught up with ASPA.sup.+/- animal by the 10.sup.th week; iii). Gene therapy completely corrected gait of CD mice as well as motor function of the CD mice treated at P1 (FIG. 30A) as measured by rotarod test; iv). Gene therapy restored the vision of CD mice. The electroretinography (ERG) tests on the eyes of the CD mice showed non-recordable responses to light, while well-defined ERG responses were readily detectable in the treated CD mice (FIG. 30B). These data indicate a severer retinopathy and loss of vision in CD mice and gene therapy can mitigate the retinopathy and restore the vision of CD mice; v). Gene therapy clearly improved metabolic defects of NAA as the NAA levels in the treated CD mice approach those in the control mice (FIG. 30C); and vi) correction of NAA metabolism is well correlated with restoration of ASPA expression (FIG. 30E) and activities (FIG. 30D) in the brain of the treated CD mice.
[0246] To determine if the phenotypic corrections are correlated with alleviated neuropathology as well as in situ expression of ASPA in the brain sections of the treated CD mice, brain sections were analyzed at 3 months after gene therapy for neuropathology and ASPA immunohistochemistry. While the untreated mouse brain shows marked vacuolation that diffusely involves all regions of the brain and spinal cord, the vacuolation in both brain and spinal cord of the treated animal appears more patchy and variable with generally smaller-sized vacuoles. Some areas of the cerebral cortex show almost no vacuolation (FIG. 31A). In addition, avidin-biotin complex (ABC) system was used to stain brain sections to detect ASPA expression in the cerebral cortex in situ (FIG. 31B). To generate quantitative measurements on the improvement of neuropathology in the treated CD mice, the "vacuoles" in the brain and spinal cord sections caused by the white matter degeneration in the CD mice were quantified before and after gene therapy treatment. For this quantitative analysis, a Nikon Eclipse Ti inverted microscope and Nikon NIS elements AR software V.3.2 were used. Vacuoles that were >3,000 pixels, 1,000-3,000 pixels and 100-1,000 pixels were defined as large, medium and small vacuoles respectively. Among the 5 brain regions evaluated in this experiment, the olfactory bulb had the most dramatic mitigation in the white matter degeneration after gene therapy (FIG. 32A). For the other 3 regions, while the large vacuoles were completely eliminated and the numbers of medium vacuoles were remarkably reduced, the reduction in the numbers of small vacuoles (<100 um) was not as significant in this experiment (FIG. 32A). The same analysis on the spinal cord sections revealed a similar trend (FIG. 32B).
[0247] Histopathology of the kidneys in the CD mice were evaluated. The glomeruli showed normal structure but were associated with dilation of Bowman's spaces. The renal tubular epithelium was diffusely attenuated (or atrophic) in association with enlargement of the tubular lumens (FIG. 33A). In contrast, the treated CD mouse had normal glomeruli. Renal tubular epithelial cells were well-stained and normal in volume (FIG. 33B). These results indicate the involvement of kidney in the pathophysiology of CD and kidney as a peripheral target for CD gene therapy. This result also indicates renal tropism of AAV vectors as a consideration for selection of a vector for CD gene therapy. Two vectors, rAAV9 and rh.10 were evaluated for efficiency of kidney transduction after IV delivery to 10 week old C57BL/6 mice. The results indicate the use of rAAVrh.10 (FIG. 33D) as a useful vector for CD gene therapy because it transduces kidney efficiently in addition to its efficient CNS transduction (FIG. 33C).
TABLE-US-00006 NUCLEIC ACID AND AMINO ACID SEQUENCES >gi|9632548|ref|NP_049542.1| capsid protein [Adeno-associated virus-1] (SEQ ID NO: 1) MAADGYLPDWLEDNLSEGIREWWDLKPGAPKPKANQQKQDDGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPLGLVEEGAKTAPGKKRPVEQSPQEP- DSSSG IGKTGQQPAKKRLNFGQTGDSESVPDPQPLGEPPATPAAVGPTTMASGGGAPMADNNEGADGVGNASGNWHCDS- TWLGD RVITTSTRTWALPTYNNHLYKQISSASTGASNDNHYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPKR- LNFKL FNIQVKEVTTNDGVTTIANNLTSTVQVFSDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQAVGR- SSFYC LEYFPSQMLRTGNNFTFSYTFEEVPFHSSYAHSQSLDRLMNPLIDQYLYYLNRTQNQSGSAQNKDLLFSRGSPA- GMSVQ PKNWLPGPCYRQQRVSKTKTDNNNSNFTWTGASKYNLNGRESIINPGTAMASHKDDEDKFFPMSGVMIFGKESA- GASNT ALDNVMITDEEEIKATNPVATERFGTVAVNFQSSSTDPATGDVHAMGALPGMVWQDRDVYLQGPIWAKIPHTDG- HFHPS PLMGGFGLKNPPPQILIKNTPVPANPPAEFSATKFASFITQYSTGQVSVEIEWELQKENSKRWNPEVQYTSNYA- KSANV DFTVDNNGLYTEPRPIGTRYLTRPL >gi|110645923|ref|YP_680426.1| major coat protein VP1 [Adeno-associated virus-2] (SEQ ID NO: 2) MAADGYLPDWLEDTLSEGIRQWWKLKPGPPPPKPAERHKDDSRGLVLPGYKYLGPFNGLDKGEPVNEADAAALE- HDKAY DRQLDSGDNPYLKYNHADAEFQERLKEDTSFGGNLGRAVFQAKKRVLEPLGLVEEPVKTAPGKKRPVEHSPVEP- DSSSG TGKAGQQPARKRLNFGQTGDADSVPDPQPLGQPPAAPSGLGTNTMATGSGAPMADNNEGADGVGNSSGNWHCDS- TWMGD RVITTSTRTWALPTYNNHLYKQISSQSGASNDNHYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPKRL- NFKLF NIQVKEVTQNDGTTTIANNLTSTVQVFTDSEYQLPYVLGSAHQGCLPPFPADVFMVPQYGYLTLNNGSQAVGRS- SFYCL EYFPSQMLRTGNNFTFSYTFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLSRTNTPSGTTTQSRLQFSQAGASD- IRDQS RNWLPGPCYRQQRVSKTSADNNNSEYSWTGATKYHLNGRDSLVNPGPAMASHKDDEEKFFPQSGVLIFGKQGSE- KTNVD IEKVMITDEEEIRTTNPVATEQYGSVSTNLQRGNRQAATADVNTQGVLPGMVWQDRDVYLQGPIWAKIPHTDGH- FHPSP LMGGFGLKHPPPQILIKNTPVPANPSTTFSAAKFASFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSNYNK- SVNVD FTVDTNGVYSEPRPIGTRYLTRNL >gi|51593838|ref|YP_068409.1| capsid protein [Adeno-associated virus-5] (SEQ ID NO: 3) MSFVDHPPDWLEEVGEGLREFLGLEAGPPKPKPNQQHQDQARGLVLPGYNYLGPGNGLDRGEPVNRADEVAREH- DISYN EQLEAGDNPYLKYNHADAEFQEKLADDTSFGGNLGKAVFQAKKRVLEPFGLVEEGAKTAPTGKRIDDHFPKRKK- ARTEE DSKPSTSSDAEAGPSGSQQLQIPAQPASSLGADTMSAGGGGPLGDNNQGADGVGNASGDWHCDSTWMGDRVVTK- STRTW VLPSYNNHQYREIKSGSVDGSNANAYFGYSTPWGYFDFNRFHSHWSPRDWQRLINNYWGFRPRSLRVKIFNIQV- KEVTV QDSTTTIANNLTSTVQVFTDDDYQLPYVVGNGTEGCLPAFPPQVFTLPQYGYATLNRDNTENPTERSSFFCLEY- FPSKM LRTGNNFEFTYNFEEVPFHSSFAPSQNLFKLANPLVDQYLYRFVSTNNTGGVQFNKNLAGRYANTYKNWFPGPM- GRTQG WNLGSGVNRASVSAFATTNRMELEGASYQVPPQPNGMTNNLQGSNTYALENTMIFNSQPANPGTTATYLEGNML- ITSES ETQPVNRVAYNVGGQMATNNQSSTTAPATGTYNLQEIVPGSVWMERDVYLQGPIWAKIPETGAHFHPSPAMGGF- GLKHP PPMMLIKNTPVPGNITSFSDVPVSSFITQYSTGQVTVEMEWELKKENSKRWNPEIQYTNNYNDPQFVDFAPDST- GEYRT TRPIGTRYLTRPL >gi|2766607|gb|AAB95450.1| capsid protein VP1 [Adeno-associated virus-6] (SEQ ID NO: 4) MAADGYLPDWLEDNLSEGIREWWDLKPGAPKPKANQQKQDDGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPFGLVEEGAKTAPGKKRPVEQSPQEP- DSSSG IGKTGQQPAKKRLNFGQTGDSESVPDPQPLGEPPATPAAVGPTTMASGGGAPMADNNEGADGVGNASGNWHCDS- TWLGD RVITTSTRTWALPTYNNHLYKQISSASTGASNDNHYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPKR- LNFKL FNIQVKEVTTNDGVTTIANNLTSTVQVFSDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQAVGR- SSFYC LEYFPSQMLRTGNNFTFSYTFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLNRTQNQSGSAQNKDLLFSRGSPA- GMSVQ PKNWLPGPCYRQQRVSKTKTDNNNSNFTWTGASKYNLNGRESIINPGTAMASHKDDKDKFFPMSGVMIFGKESA- GASNT ALDNVMITDEEEIKATNPVATERFGTVAVNLQSSSTDPATGDVHVMGALPGMVWQDRDVYLQGPIWAKIPHTDG- HFHPS PLMGGFGLKHPPPQILIKNTPVPANPPAEFSATKFASFITQYSTGQVSVEIEWELQKENSKRWNPEVQYTSNYA- KSANV DFTVDNNGLYTEPRPIGTRYLTRPL >gi|171850125|gb|ACB55302.1| capsid protein VP1 [Adeno-associated virus-6.2] (SEQ ID NO: 5) MAADGYLPDWLEDNLSEGIREWWDLKPGAPKPKANQQKQDDGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPLGLVEEGAKTAPGKKRPVEQSPQEP- DSSSG IGKTGQQPAKKRLNFGQTGDSESVPDPQPLGEPPATPAAVGPTTMASGGGAPMADNNEGADGVGNASGNWHCDS- TWLGD RVITTSTRTWALPTYNNHLYKQISSASTGASNDNHYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPKR- LNFKL FNIQVKEVTTNDGVTTIANNLTSTVQVFSDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQAVGR- SSFYC LEYFPSQMLRTGNNFTFSYTFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLNRTQNQSGSAQNKDLLFSRGSPA- GMSVQ PKNWLPGPCYRQQRVSKTKTDNNNSNFTWTGASKYNLNGRESIINPGTAMASHKDDKDKFFPMSGVMIFGKESA- GASNT ALDNVMITDEEEIKATNPVATERFGTVAVNLQSSSTDPATGDVHVMGALPGMVWQDRDVYLQGPIWAKIPHTDG- HFHPS PLMGGFGLKHPPPQILIKNTPVPANPPAEFSATKFASFITQYSTGQVSVEIEWELQKENSKRWNPEVQYTSNYA- KSANV DFTVDNNGLYTEPRPIGTRYLTRPL >gi|22652861|gb|AAN03855.1|AF513851_2 capsid protein [Adeno-associated virus-7] (SEQ ID NO: 6) MAADGYLPDWLEDNLSEGIREWWDLKPGAPKPKANQQKQDNGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPLGLVEEGAKTAPAKKRPVEPSPQRS- PDSST GIGKKGQQPARKRLNFGQTGDSESVPDPQPLGEPPAAPSSVGSGTVAAGGGAPMADNNEGADGVGNASGNWHCD- STWLG DRVITTSTRTWALPTYNNHLYKQISSETAGSTNDNTYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPK- KLRFK LFNIQVKEVTTNDGVTTIANNLTSTIQVFSDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQSVG- RSSFY CLEYFPSQMLRTGNNFEFSYSFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLARTQSNPGGTAGNRELQFYQGG- PSTMA EQAKNWLPGPCFRQQRVSKTLDQNNNSNFAWTGATKYHLNGRNSLVNPGVAMATHKDDEDRFFPSSGVLIFGKT- GATNK TTLENVLMTNEEEIRPTNPVATEEYGIVSSNLQAANTAAQTQVVNNQGALPGMVWQNRDVYLQGPIWAKIPHTD- GNFHP SPLMGGFGLKHPPPQILIKNTPVPANPPEVFTPAKFASFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSNF- EKQTG VDFAVDSQGVYSEPRPIGTRYLTRNL >gi|22652864|gb|AAN03857.1|AF513852_2 capsid protein [Adeno-associated virus-8] (SEQ ID NO: 7) MAADGYLPDWLEDNLSEGIREWWALKPGAPKPKANQQKQDDGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLQAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPLGLVEEGAKTAPGKKRPVEPSPQRS- PDSST GIGKKGQQPARKRLNFGQTGDSESVPDPQPLGEPPAAPSGVGPNTMAAGGGAPMADNNEGADGVGSSSGNWHCD- STWLG DRVITTSTRTWALPTYNNHLYKQISNGTSGGATNDNTYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRP- KRLSF KLFNIQVKEVTQNEGTKTIANNLTSTIQVFTDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQAV- GRSSF YCLEYFPSQMLRTGNNFQFTYTFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLSRTQTTGGTANTQTLGFSQGG- PNTMA NQAKNWLPGPCYRQQRVSTTTGQNNNSNFAWTAGTKYHLNGRNSLANPGIAMATHKDDEERFFPSNGILIFGKQ- NAARD NADYSDVMLTSEEEIKTTNPVATEEYGIVADNLQQQNTAPQIGTVNSQGALPGMVWQNRDVYLQGPIWAKIPHT- DGNFH PSPLMGGFGLKHPPPQILIKNTPVPADPPTTFNQSKLNSFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSN- YYKST SVDFAVNTEGVYSEPRPIGTRYLTRNL >gi|46487805|gb|AAS99264.1| capsid protein VP1 [Adeno-associated virus 9] (SEQ ID NO: 8) MAADGYLPDWLEDNLSEGIREWWALKPGAPQPKANQQHQDNARGLVLPGYKYLGPGNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLKYNHADAEFQERLKEDTSFGGNLGRAVFQAKKRLLEPLGLVEEAAKTAPGKKRPVEQSPQEP- DSSAG IGKSGAQPAKKRLNFGQTGDTESVPDPQPIGEPPAAPSGVGSLTMASGGGAPVADNNEGADGVGSSSGNWHCDS- QWLGD RVITTSTRTWALPTYNNHLYKQISNSTSGGSSNDNAYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPK- RLNFK LFNIQVKEVTDNNGVKTIANNLTSTVQVFTDSDYQLPYVLGSAHEGCLPPFPADVFMIPQYGYLTLNDGSQAVG- RSSFY CLEYFPSQMLRTGNNFQFSYEFENVPFHSSYAHSQSLDRLMNPLIDQYLYYLSKTINGSGQNQQTLKFSVAGPS- NMAVQ GRNYIPGPSYRQQRVSTTVTQNNNSEFAWPGASSWALNGRNSLMNPGPAMASHKEGEDRFFPLSGSLIFGKQGT- GRDNV DADKVMITNEEEIKTTNPVATESYGQVATNHQSAQAQAQTGWVQNQGILPGMVWQDRDVYLQGPIWAKIPHTDG- NFHPS PLMGGFGMKHPPPQILIKNTPVPADPPTAFNKDKLNSFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSNYY- KSNNV EFAVNTEGVYSEPRPIGTRYLTRNL >gi|29650526|gb|AA088201.1| capsid protein [Non-human primate Adeno-associated virus] (SEQ ID NO: 9) rh-10 MAADGYLPDWLEDNLSEGIREWWDLKPGAPKPKANQQKQDDGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPLGLVEEGAKTAPGKKRPVEPSPQRS- PDSST GIGKKGQQPAKKRLNFGQTGDSESVPDPQPIGEPPAGPSGLGSGTMAAGGGAPMADNNEGADGVGSSSGNWHCD- STWLG DRVITTSTRTWALPTYNNHLYKQISNGTSGGSTNDNTYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRP- KRLNF KLFNIQVKEVTQNEGTKTIANNLTSTIQVFTDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQAV- GRSSF YCLEYFPSQMLRTGNNFEFSYQFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLSRTQSTGGTAGTQQLLFSQAG- PNNMS AQAKNWLPGPCYRQQRVSTTLSQNNNSNFAWTGATKYHLNGRDSLVNPGVAMATHKDDEERFFPSSGVLMFGKQ- GAGKD NVDYSSVMLTSEEEIKTTNPVATEQYGVVADNLQQQNAAPIVGAVNSQGALPGMVWQNRDVYLQGPIWAKIPHT- DGNFH PSPLMGGFGLKHPPPQILIKNTPVPADPPTTFSQAKLASFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSN- YYKST NVDFAVNTDGTYSEPRPIGTRYLTRNL >gi|171850147|gb|ACB55313.1| capsid protein VP1 [Adeno-as sociated virus-rh.39] (SEQ ID NO: 10) MAADGYLPDWLEDNLSEGIREWWALKPGAPKPKANQQKQDDGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPLGLVEEAAKTAPGKKRPVEPSPQRS- PDSST GIGKKGQQPAKKRLNFGQTGDSESVPDPQPIGEPPAGPSGLGSGTMAAGGGAPMADNNEGADGVGSSSGNWHCD- STWLG DRVITTSTRTWALPTYNNHLYKQISNGTSGGSTNDNTYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRP- KRLSF KLFNIQVKEVTQNEGTKTIANNLTSTIQVFTDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQAV- GRSSF YCLEYFPSQMLRTGNNFEFSYTFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLSRTQSTGGTQGTQQLLFSQAG- PANMS AQAKNWLPGPCYRQQRVSTTLSQNNNSNFAWTGATKYHLNGRDSLVNPGVAMATHKDDEERFFPSSGVLMFGKQ- GAGRD NVDYSSVMLTSEEEIKTTNPVATEQYGVVADNLQQTNTGPIVGNVNSQGALPGMVWQNRDVYLQGPIWAKIPHT- DGNFH PSPLMGGFGLKHPPPQILIKNTPVPADPPTTFSQAKLASFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSN- YYKST NVDFAVNTEGTYSEPRPIGTRYLTRNL >gi|46487767|gb|AAS99245.1| capsid protein VP1 [Adeno-associated virus rh.43] (SEQ ID NO: 11) MAADGYLPDWLEDNLSEGIREWWDLKPGAPKPKANQQKQDDGRGLVLPGYKYLGPFNGLDKGEPVNAADAAALE- HDKAY DQQLEAGDNPYLRYNHADAEFQERLQEDTSFGGNLGRAVFQAKKRVLEPLGLVEEGAKTAPGKKRPVEQSPQEP- DSSSG IGKKGQQPARKRLNFGQTGDSESVPDPQPLGEPPAAPSGVGPNTMAAGGGAPMADNNEGADGVGSSSGNWHCDS- TWLGD RVITTSTRTWALPTYNNHLYKQISNGTSGGATNDNTYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPK- RLSFK LFNIQVKEVTQNEGTKTIANNLTSTIQVFTDSEYQLPYVLGSAHQGCLPPFPADVFMIPQYGYLTLNNGSQAVG- RSSFY CLEYFPSQMLRTGNNFQFTYTFEDVPFHSSYAHSQSLDRLMNPLIDQYLYYLSRTQTTGGTANTQTLGFSQGGP- NTMAN
QAKNWLPGPCYRQQRVSTTTGQNNNSNFAWTAGTKYHLNGRNSLANPGIAMATHKDDEERFFPVTGSCFWQQNA- ARDNA DYSDVMLTSEEEIKTTNPVATEEYGIVADNLQQQNTAPQIGTVNSQGALPGMVWQNRDVYLQGPIWAKIPHTDG- NFHPS PLMGGFGLKHPPPQILIKNTPVPADPPTTFNQSKLNSFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSNYY- KSTSV DFAVNTEGVYSEPRPIGTRYLTRNL >capsid protein VP1 [Adeno-associated virus] CSp3 (SEQ ID NO: 12) MAADGYLPDWLEDNLSEGIREWWALKPGAPQPKANQQHQDNARGLVLPGYKYLGPGNGLDKGEPVNAADAAALE- HDKAY DQQLKAGDNPYLKYNHADAEFQERLKEDTSFGGNLGRAVFQAKKRLLEPLGLVEEAAKTAPGKKRPVEQSPQEP- DSSAG IGKSGAQPAKKRLNFGQTGDTESVPDPQPIGEPPAAPSGVGSLTIASGGGAPVADNNEGADGVGSSSGNWHCDS- QWLGD RVITTSTRTWALPTYNNHLYKRISNSTSGGSSNDNAYFGYSTPWGYFDFNRFHCHFSPRDWQRLINNNWGFRPK- RLNFK LFNIRVKEVTDNNGVKTITNNLTSTVQVFTDSDYQLPYVLGSAHEGCLPPFPADVFMIPQYGYLTLNDGSQAVG- RSSFY CLEYFPSQMLRTGNNFQFSYEFENVPFHSSYAHSQSLDRLMNPLIDQYLYYLSKTINGSGQNQQTLKFSVAGPS- NMAVQ GRNYIPGPSYRQQRVSTTVTRNNNSEFAWPGASSWALNGRNSLMNPGPAMASHKEGEDRFFPLSGSLIFGKQGT- GRDNV DADKVMITNEEEIKTTNPVATESYGQVATNHQSAQAQAQTGWVQNQGILPGMVWQDRDVYLQGPIWAKIPHTDG- NFHPS PLMGGFGVKHPPPQILIKNTPVPADPPTAFNKDKLNSFITQYSTGQVSVEIEWELQKENSKRWNPEIQYTSNYY- KSNNV EFAVNTEGVYSEPRPIGTRYLTRNL >gi|189339202|ref|NP_001121557.1| aspartoacylase [Homo sapiens] (SEQ ID NO: 13) MTSCHIAEEHIQKVAIFGGTHGNELTGVFLVKHWLENGAEIQRTGLEVKPFITNPRAVKKCTRYIDCDLNRIFD- LENLG KKMSEDLPYEVRRAQEINHLFGPKDSEDSYDIIFDLHNTTSNMGCTLILEDSRNNFLIQMFHYIKTSLAPLPCY- VYLIE HPSLKYATTRSIAKYPVGIEVGPQPQGVLRADILDQMRKMIKHALDFIHHFNEGKEFPPCAIEVYKIIEKVDYP- RDENG EIAAIIHPNLQDQDWKPLHPGDPMFLTLDGKTIPLGGDCTVYPVFVNEAAYYEKKEAFAKTTKLTLNAKSIRCC- LH >gi|189339201: 92-1033 Homo sapiens aspartoacylase (Canavan disease) (ASPA), transcript variant 2, mRNA (SEQ ID NO: 14) ATGACTTCTTGTCACATTGCTGAAGAACATATACAAAAGGTTGCTATCTTTGGAGGAACCCATGGGAATGAGCT- AACCG GAGTATTTCTGGTTAAGCATTGGCTAGAGAATGGCGCTGAGATTCAGAGAACAGGGCTGGAGGTAAAACCATTT- ATTAC TAACCCCAGAGCAGTGAAGAAGTGTACCAGATATATTGACTGTGACCTGAATCGCATTTTTGACCTTGAAAATC- TTGGC AAAAAAATGTCAGAAGATTTGCCATATGAAGTGAGAAGGGCTCAAGAAATAAATCATTTATTTGGTCCAAAAGA- CAGTG AAGATTCCTATGACATTATTTTTGACCTTCACAACACCACCTCTAACATGGGGTGCACTCTTATTCTTGAGGAT- TCCAG GAATAACTTTTTAATTCAGATGTTTCATTACATTAAGACTTCTCTGGCTCCACTACCCTGCTACGTTTATCTGA- TTGAG CATCCTTCCCTCAAATATGCGACCACTCGTTCCATAGCCAAGTATCCTGTGGGTATAGAAGTTGGTCCTCAGCC- TCAAG GGGTTCTGAGAGCTGATATCTTGGATCAAATGAGAAAAATGATTAAACATGCTCTTGATTTTATACATCATTTC- AATGA AGGAAAAGAATTTCCTCCCTGCGCCATTGAGGTCTATAAAATTATAGAGAAAGTTGATTACCCCCGGGATGAAA- ATGGA GAAATTGCTGCTATCATCCATCCTAATCTGCAGGATCAAGACTGGAAACCACTGCATCCTGGGGATCCCATGTT- TTTAA CTCTTGATGGGAAGACGATCCCACTGGGCGGAGACTGTACCGTGTACCCCGTGTTTGTGAATGAGGCCGCATAT- TACGA AAAGAAAGAAGCTTTTGCAAAGACAACTAAACTAACGCTCAATGCAAAAAGTATTCGCTGCTGTTTACATTAG >gi|31560279|ref|NP_075602.2| aspartoacylase [Mus Musculus] (SEQ ID NO: 15) MTSCVAKEPIKKIAIFGGTHGNELTGVFLVTHWLRNGTEVHRAGLDVKPFITNPRAVEKCTRYIDCDLNRVFDL- ENLSK EMSEDLPYEVRRAQEINHLFGPKNSDDAYDLVFDLHNTTSNMGCTLILEDSRNDFLIQMFHYIKTCMAPLPCSV- YLIEH PSLKYATTRSIAKYPVGIEVGPQPHGVLRADILDQMRKMIKHALDFIQHFNEGKEFPPCSIDVYKIMEKVDYPR- NESGD MAAVIHPNLQDQDWKPLHPGDPVFVSLDGKVIPLGGDCTVYPVFVNEAAYYEKKEAFAKTTKLTLSAKSIRSTL- H >gi|142354273: 148-1086 Mus musculus aspartoacylase (Aspa), mRNA (SEQ ID NO: 16) ATGACCTCTTGTGTTGCTAAAGAACCTATTAAGAAGATTGCCATCTTTGGAGGGACTCATGGAAATGAACTGAC- CGGAG TGTTTCTAGTTACTCACTGGCTAAGGAATGGCACTGAAGTTCACAGAGCAGGGCTGGACGTGAAGCCATTCATT- ACCAA TCCAAGGGCGGTGGAGAAGTGCACCAGATACATTGACTGTGACCTGAATCGTGTTTTTGACCTTGAAAATCTTA- GCAAA GAGATGTCTGAAGACTTGCCATATGAAGTGAGAAGGGCTCAAGAAATAAATCATTTATTTGGTCCAAAAAATAG- TGATG ATGCCTATGACCTTGTTTTTGACCTTCACAACACCACTTCTAACATGGGTTGCACTCTTATTCTTGAGGATTCC- AGGAA TGACTTTTTAATTCAGATGTTTCACTATATTAAGACTTGCATGGCTCCATTACCCTGCTCTGTTTATCTCATTG- AGCAT CCTTCACTCAAATATGCAACCACTCGTTCCATTGCCAAGTATCCTGTTGGTATAGAAGTTGGTCCTCAGCCTCA- CGGTG TCCTTAGAGCTGATATTTTAGACCAAATGAGAAAAATGATAAAACATGCTCTTGATTTTATACAGCATTTCAAT- GAAGG AAAAGAATTTCCTCCCTGTTCTATTGACGTCTATAAAATAATGGAGAAAGTTGATTATCCAAGGAATGAAAGTG- GAGAC ATGGCTGCTGTTATTCATCCTAATCTGCAGGATCAAGACTGGAAACCATTGCACCCTGGAGATCCTGTGTTTGT- GTCTC TTGATGGAAAAGTTATTCCACTGGGTGGAGACTGTACCGTGTACCCAGTGTTTGTGAATGAAGCTGCATATTAT- GAAAA AAAAGAAGCATTTGCAAAGACAACAAAACTAACACTCAGCGCAAAAAGCATCCGCTCCACTTTGCACTAA >gi|48762945: 149-613 Homo sapiens superoxide dismutase 1, soluble (SOD1), mRNA (SEQ ID NO: 17) ATGGCGACGAAGGCCGTGTGCGTGCTGAAGGGCGACGGCCCAGTGCAGGGCATCATCAATTTCGAGCAGAAGGA- AAGTA ATGGACCAGTGAAGGTGTGGGGAAGCATTAAAGGACTGACTGAAGGCCTGCATGGATTCCATGTTCATGAGTTT- GGAGA TAATACAGCAGGCTGTACCAGTGCAGGTCCTCACTTTAATCCTCTATCCAGAAAACACGGTGGGCCAAAGGATG- AAGAG AGGCATGTTGGAGACTTGGGCAATGTGACTGCTGACAAAGATGGTGTGGCCGATGTGTCTATTGAAGATTCTGT- GATCT CACTCTCAGGAGACCATTGCATCATTGGCCGCACACTGGTGGTCCATGAAAAAGCAGATGACTTGGGCAAAGGT- GGAAA TGAAGAAAGTACAAAGACAGGAAACGCTGGAAGTCGTTTGGCTTGTGGTGTAATTGGGATCGCCCAATAA >gi|4507149|ref|NP_000445.1| superoxide dismutase [Homo sapiens] (SEQ ID NO: 18) MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGG- PKDEE RHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIA- Q >gi|45597446: 117-581 Mus musculus superoxide dismutase 1, soluble (Sod1), mRNA (SEQ ID NO: 19) ATGGCGATGAAAGCGGTGTGCGTGCTGAAGGGCGACGGTCCGGTGCAGGGAACCATCCACTTCGAGCAGAAGGC- AAGCG GTGAACCAGTTGTGTTGTCAGGACAAATTACAGGATTAACTGAAGGCCAGCATGGGTTCCACGTCCATCAGTAT- GGGGA CAATACACAAGGCTGTACCAGTGCAGGACCTCATTTTAATCCTCACTCTAAGAAACATGGTGGCCCGGCGGATG- AAGAG AGGCATGTTGGAGACCTGGGCAATGTGACTGCTGGAAAGGACGGTGTGGCCAATGTGTCCATTGAAGATCGTGT- GATCT CACTCTCAGGAGAGCATTCCATCATTGGCCGTACAATGGTGGTCCATGAGAAACAAGATGACTTGGGCAAAGGT- GGAAA TGAAGAAAGTACAAAGACTGGAAATGCTGGGAGCCGCTTGGCCTGTGGAGTGATTGGGATTGCGCAGTAA >gi|45597447|ref|NP035564.1| superoxide dismutase [Mus musculus] (SEQ ID NO: 20) MAMKAVCVLKGDGPVQGTIHFEQKASGEPVVLSGQITGLTEGQHGFHVHQYGDNTQGCTSAGPHFNPHSKKHGG- PADEE RHVGDLGNVTAGKDGVANVSIEDRVISLSGEHSIIGRTMVVHEKQDDLGKGGNEESTKTGNAGSRLACGVIGIA- Q >pAA VscCB6 EGFPmir SOD5 (direct) 5243 bp (SEQ ID NO: 21) CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTC- AGTGA GCGAGCGAGCGCGCAGAGAGGGAGTGTAGCCATGCTCTAGGAAGATCAATTCAATTCACGCGTCGACATTGATT- ATTGA CTAGTTATTAATAGTAATCAATTACGGGGTCATTAGTTCATAGCCCATATATGGAGTTCCGCGTTACATAACTT- ACGGT AAATGGCCCGCCTGGCTGACCGCCCAACGACCCCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAA- CGCCA ATAGGGACTTTCCATTGACGTCAATGGGTGGATATTTACGGTAAACTGCCCACTTGGCAGTACATCAAGTGTAT- CATAT GCCAAGTACGCCCCCTATTGACGTCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTAT- GGGAC TTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTACCATGTCGAGGCCACGTTCTGCTTCACTCTC- CCCAT CTCCCCCCCCTCCCCACCCCCAATTTTGTATTTATTTATTTTTTAATTATTTTGTGCAGCGATGGGGGCGGGGG- GGGGG GGCGCGCGCCAGGCGGGGCGGGGCGGGGCGAGGGGCGGGGCGGGGCGAGGCGGAGAGGTGCGGCGGCAGCCAAT- CAGAG CGGCGCGCTCCGAAAGTTTCCTTTTATGGCGAGGCGGCGGCGGCGGCGGCCCTATAAAAAGCGAAGCGCGCGGC- GGGCG GGAGCAAGCTCTAGCCTCGAGAATTCACGCGTGGTACCTCTAGAGCAGAGCTCGTTTAGTGAACCGTCAGTTCG- AAATC GCCACCATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGT- AAACG GCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGC- ACCAC CGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACC- CCGAC CACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAA- GGACG ACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGC- ATCGA CTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGG- CCGAC AAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGA- CCACT ACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCC- CTGAG CAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCA- TGGAC GAGCTGTACAAGTAAGTAACAGGTAAGTGCGATCGCTAATGCGGGAAAGCTCTTATTCGGGTGAGATGGGCTGG- GGCAC CATCTGGGGACCCTGACGTGAAGTTTGTCACTGACTGGAGAACTCGGTTTGTCGTCTGTTGCGGGGGCGGCAGT- TATGG CGGTGCCGTTGGGCAGTGCACCCGTACCTTTGGGAGCGCGCGCCCTCGTCGTGTCGTGACGTCACCCGTTCTGT- TGGTA CCTGCTGTTGACAGTGAGCGACGCAATGTGACTTCGCTGACAAAGCTGTGAAGCCACAGATGGGCTTTGTCAGC- AGTCA CATTGCGCTGCCTACTGCCTCGGACTTCAAGGGCTCGAGAATTCAGGGTGGGGCCACCTGCCGGTAGGTGTGCG- GTAGG CTTTTCTCCGTCGCAGGACGCAGGGTTCGGGCCTAGGGTAGGCTCTCCTGAATCGACAGGCGCCGGACCTCTGG- CGGCC GCAACAACGCGTTCCTGACCATTCATCCTCTTTCTTTTTCCTGCAGGCTTGTGGAAGAAATGGGATCCGATCTT- TTTCC CTCTGCCAAAAATTATGGGGACATCATGAAGCCCCTTGAGCATCTGACTTCTGGCTAATAAAGGAAATTTATTT- TCATT GCAATAGTGTGTTGGAATTTTTTGTGTCTCTCACTCGGCCTAGGTAGATAAGTAGCATGGCGGGTTAATCATTA- ACTAC AAGGAACCCCTAGTGATGGAGTTGGCCACTCCCTCTCTGCGCGCTCGCTCGCTCACTG]GGCCGGGCGACCAAA- GGTCG CCCGACGCCCGGGCTTTGCCCGGGCGGCCTCAGTGAGCGAGCGAGCGCGCAGCCTTAATTAACCTAATTCACTG- GCCGT CGTTTTACAACGTCGTGACTGGGAAAACCCTGGCGTTACCCAACTTAATCGCCTTGCAGCACATCCCCCTTTCG- CCAGC TGGCGTAATAGCGAAGAGGCCCGCACCGATCGCCCTTCCCAACAGTTGCGCAGCCTGAATGGCGAATGGGACGC- GCCCT GTAGCGGCGCATTAAGCGCGGCGGGTGTGGTGGTTACGCGCAGCGTGACCGCTACACTTGCCAGCGCCCTAGCG- CCCGC TCCTTTCGCTTTCTTCCCTTCCTTTCTCGCCACGTTCGCCGGCTTTCCCCGTCAAGCTCTAAATCGGGGGCTCC- CTTTA GGGTTCCGATTTAGTGCTTTACGGCACCTCGACCCCAAAAAACTTGATTAGGGTGATGGTTCACGTAGTGGGCC- ATCGC CCTGATAGACGGTTTTTCGCCCTTTGACGTTGGAGTCCACGTTCTTTAATAGTGGACTCTTGTTCCAAACTGGA- ACAAC ACTCAACCCTATCTCGGTCTATTCTTTTGATTTATAAGGGATTTTGCCGATTTCGGCCTATTGGTTAAAAAATG- AGCTG ATTTAACAAAAATTTAACGCGAATTTTAACAAAATATTAACGCTTACAATTTAGGTGGCACTTTTCGGGGAAAT- GTGCG CGGAACCCCTATTTGTTTATTTTTCTAAATACATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAA- TGCTT CAATAATATTGAAAAAGGAAGAGTATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATT- TTGCC TTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGT- TACAT CGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTT- TTAAA GTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTC- TCAGA ATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGT- GCTGC CATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTT- TTTTG CACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGA- GCGTG ACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCC- CGGCA ACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGT- TTATT GCTGATAAATCTGGAGCCGGTGAGCGTGGGTCTCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTC- CCGTA
TCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCC- TCACT GATTAAGCATTGGTAACTGTCAGACCAAGTTTACTCATATATACTTTAGATTGATTTAAAACTTCATTTTTAAT- TTAAA AGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAAAATCCCTTAACGTGAGTTTTCGTTCCACTGAGC- GTCAG ACCCCGTAGAAAAGATCAAAGGATCTTCTTGAGATCCTTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAA- AAACC ACCGCTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTCAGCA- GAGCG CAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCTGTAGCACCGCCTAC- ATACC TCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGA- CGATA GTTACCGGATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGAACGACCT- ACACC GAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCC- GGTAA GCGGCAGGGTCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTC- GGGTT TCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAAAACGCCAGCA- ACGCG GCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTTCTTTCCTGCGTTATCCCCTGATTCTGT- GGATA ACCGTATTACCGCCTTTGAGTGAGCTGATACCGCTCGCCGCAGCCGAACGACCGAGCGCAGCGAGTCAGTGAGC- GAGGA AGCGGAAGAGCGCCCAATACGCAAACCGCCTCTCCCCGCGCGTTGGCCGATTCATTAATGCAGCTGGCACGACA- GGTTT CCCGACTGGAAAGCGGGCAGTGAGCGCAACGCAATTAATGTGAGTTAGCTCACTCATTAGGCACCCCAGGCTTT- ACACT TTATGCTTCCGGCTCGTATGTTGTGTGGAATTGTGAGCGGATAACAATTTCACACAGGAAACAGCTATGACCAT- GATTA CGCCAGATTTAATTAAGGCCTTAATTAGG >sod1mir1 (direct) 108 bp (SEQ ID NO: 22) TGCTGTTGACAGTGAGCGACATCATCAATTTTCCGAGCAGAACTGTGAAGCCACAGATGGGTTCTGCTCGAAAT- TGATG ATGCTGCCTACTGCCTCGGACTTCAAGGG >sod1mir2 (direct) 106 bp (SEQ ID NO: 23) TGCTGTTGACAGTGAGCGACGCATTAAAGGATCCTGACTGACTGTGAAGCCACAGATGGGTCAGTCAGTCCTTT- AATGC GCTGCCTACTGCCTCGGACTTCAAGGG >sod1mir3 (direct) 108 bp (SEQ ID NO: 24) TGCTGTTGACAGTGAGCGACTGCATGGATTCTCCATGTTCATCTGTGAAGCCACAGATGGGATGAACATGGAAT- CCATG CAGCTGCCTACTGCCTCGGACTTCAAGGG >sod1mir4 (direct) 106 bp (SEQ ID NO: 25) TGCTGTTGACAGTGAGCGACAAGGATGAAGATCGAGGCATGCTGTGAAGCCACAGATGGGCATGCCTCTCTTCA- TCCTT GCTGCCTACTGCCTCGGACTTCAAGGG >sod lmir5 (direct) 110 bp (SEQ ID NO: 26) TGCTGTTGACAGTGAGCGACGCAATGTGACTTCGCTGACAAAGCTGTGAAGCCACAGATGGGCTTTGTCAGCAG- TCACA TTGCGCTGCCTACTGCCTCGGACTTCAAGGG >sod1mir6 (direct) 108 bp (SEQ ID NO: 27) TGCTGTTGACAGTGAGCGACCGATGTGTCTATCTTGAAGATTCTGTGAAGCCACAGATGGGAATCTTCAATAGA- CACAT CGGCTGCCTACTGCCTCGGACTTCAAGGG >sod1mir7 (direct) 106 bp (SEQ ID NO: 28) TGCTGTTGACAGTGAGCGACGGTGGAAATGATCAGAAAGTACTGTGAAGCCACAGATGGGTACTTTCTTCATTT- CCACC GCTGCCTACTGCCTCGGACTTCAAGGG >sod1mir8 (direct) 110 bp (SEQ ID NO: 29) TGCTGTTGACAGTGAGCGACGCTGTAGAAATTCGTATCCTGATCTGTGAAGCCACAGATGGGATCAGGATACAT- TTCTA CAGCGCTGCCTACTGCCTCGGACTTCAAGGG >sod1mir9 (direct) 106 bp (SEQ ID NO: 30) TGCTGTTGACAGTGAGCGAGGTATTAAACTTGTCAGAATTTAGTGAAGCCACAGATGTAAATTCTGACAAGTTT- AATAC CCTGCCTACTGCCTCGGACTTCAAGGG >pAA VscCB6 EGFPmir scr (1820 bp-1925 bp, direct) 106 bp (SEQ ID NO: 31) TGCTGTTGACAGTGAGCGACGATGCTCTAATCGGTTCTATCAAGTGAAGCCACAGATGTTGATAGAACCTTAGA- GCATC GCTGCCTACTGCCTCGGACTTCAAGGG
[0248] This invention is not limited in its application to the details of construction and the arrangement of components set forth in this description or illustrated in the drawings. The invention is capable of other embodiments and of being practiced or of being carried out in various ways. Also, the phraseology and terminology used herein is for the purpose of description and should not be regarded as limiting. The use of "including," "comprising," or "having," "containing," "involving," and variations thereof herein, is meant to encompass the items listed thereafter and equivalents thereof as well as additional items.
[0249] Having thus described several aspects of at least one embodiment of this invention, it is to be appreciated various alterations, modifications, and improvements will readily occur to those skilled in the art. Such alterations, modifications, and improvements are intended to be part of this disclosure, and are intended to be within the spirit and scope of the invention. Accordingly, the foregoing description and drawings are by way of example only.
Sequence CWU 1
1
641736PRTAdeno associated virus 1 1Met Ala Ala Asp Gly Tyr Leu Pro
Asp Trp Leu Glu Asp Asn Leu Ser1 5 10 15Glu Gly Ile Arg Glu Trp Trp
Asp Leu Lys Pro Gly Ala Pro Lys Pro 20 25 30Lys Ala Asn Gln Gln Lys
Gln Asp Asp Gly Arg Gly Leu Val Leu Pro 35 40 45Gly Tyr Lys Tyr Leu
Gly Pro Phe Asn Gly Leu Asp Lys Gly Glu Pro 50 55 60Val Asn Ala Ala
Asp Ala Ala Ala Leu Glu His Asp Lys Ala Tyr Asp65 70 75 80Gln Gln
Leu Lys Ala Gly Asp Asn Pro Tyr Leu Arg Tyr Asn His Ala 85 90 95Asp
Ala Glu Phe Gln Glu Arg Leu Gln Glu Asp Thr Ser Phe Gly Gly 100 105
110Asn Leu Gly Arg Ala Val Phe Gln Ala Lys Lys Arg Val Leu Glu Pro
115 120 125Leu Gly Leu Val Glu Glu Gly Ala Lys Thr Ala Pro Gly Lys
Lys Arg 130 135 140Pro Val Glu Gln Ser Pro Gln Glu Pro Asp Ser Ser
Ser Gly Ile Gly145 150 155 160Lys Thr Gly Gln Gln Pro Ala Lys Lys
Arg Leu Asn Phe Gly Gln Thr 165 170 175Gly Asp Ser Glu Ser Val Pro
Asp Pro Gln Pro Leu Gly Glu Pro Pro 180 185 190Ala Thr Pro Ala Ala
Val Gly Pro Thr Thr Met Ala Ser Gly Gly Gly 195 200 205Ala Pro Met
Ala Asp Asn Asn Glu Gly Ala Asp Gly Val Gly Asn Ala 210 215 220Ser
Gly Asn Trp His Cys Asp Ser Thr Trp Leu Gly Asp Arg Val Ile225 230
235 240Thr Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr Tyr Asn Asn His
Leu 245 250 255Tyr Lys Gln Ile Ser Ser Ala Ser Thr Gly Ala Ser Asn
Asp Asn His 260 265 270Tyr Phe Gly Tyr Ser Thr Pro Trp Gly Tyr Phe
Asp Phe Asn Arg Phe 275 280 285His Cys His Phe Ser Pro Arg Asp Trp
Gln Arg Leu Ile Asn Asn Asn 290 295 300Trp Gly Phe Arg Pro Lys Arg
Leu Asn Phe Lys Leu Phe Asn Ile Gln305 310 315 320Val Lys Glu Val
Thr Thr Asn Asp Gly Val Thr Thr Ile Ala Asn Asn 325 330 335Leu Thr
Ser Thr Val Gln Val Phe Ser Asp Ser Glu Tyr Gln Leu Pro 340 345
350Tyr Val Leu Gly Ser Ala His Gln Gly Cys Leu Pro Pro Phe Pro Ala
355 360 365Asp Val Phe Met Ile Pro Gln Tyr Gly Tyr Leu Thr Leu Asn
Asn Gly 370 375 380Ser Gln Ala Val Gly Arg Ser Ser Phe Tyr Cys Leu
Glu Tyr Phe Pro385 390 395 400Ser Gln Met Leu Arg Thr Gly Asn Asn
Phe Thr Phe Ser Tyr Thr Phe 405 410 415Glu Glu Val Pro Phe His Ser
Ser Tyr Ala His Ser Gln Ser Leu Asp 420 425 430Arg Leu Met Asn Pro
Leu Ile Asp Gln Tyr Leu Tyr Tyr Leu Asn Arg 435 440 445Thr Gln Asn
Gln Ser Gly Ser Ala Gln Asn Lys Asp Leu Leu Phe Ser 450 455 460Arg
Gly Ser Pro Ala Gly Met Ser Val Gln Pro Lys Asn Trp Leu Pro465 470
475 480Gly Pro Cys Tyr Arg Gln Gln Arg Val Ser Lys Thr Lys Thr Asp
Asn 485 490 495Asn Asn Ser Asn Phe Thr Trp Thr Gly Ala Ser Lys Tyr
Asn Leu Asn 500 505 510Gly Arg Glu Ser Ile Ile Asn Pro Gly Thr Ala
Met Ala Ser His Lys 515 520 525Asp Asp Glu Asp Lys Phe Phe Pro Met
Ser Gly Val Met Ile Phe Gly 530 535 540Lys Glu Ser Ala Gly Ala Ser
Asn Thr Ala Leu Asp Asn Val Met Ile545 550 555 560Thr Asp Glu Glu
Glu Ile Lys Ala Thr Asn Pro Val Ala Thr Glu Arg 565 570 575Phe Gly
Thr Val Ala Val Asn Phe Gln Ser Ser Ser Thr Asp Pro Ala 580 585
590Thr Gly Asp Val His Ala Met Gly Ala Leu Pro Gly Met Val Trp Gln
595 600 605Asp Arg Asp Val Tyr Leu Gln Gly Pro Ile Trp Ala Lys Ile
Pro His 610 615 620Thr Asp Gly His Phe His Pro Ser Pro Leu Met Gly
Gly Phe Gly Leu625 630 635 640Lys Asn Pro Pro Pro Gln Ile Leu Ile
Lys Asn Thr Pro Val Pro Ala 645 650 655Asn Pro Pro Ala Glu Phe Ser
Ala Thr Lys Phe Ala Ser Phe Ile Thr 660 665 670Gln Tyr Ser Thr Gly
Gln Val Ser Val Glu Ile Glu Trp Glu Leu Gln 675 680 685Lys Glu Asn
Ser Lys Arg Trp Asn Pro Glu Val Gln Tyr Thr Ser Asn 690 695 700Tyr
Ala Lys Ser Ala Asn Val Asp Phe Thr Val Asp Asn Asn Gly Leu705 710
715 720Tyr Thr Glu Pro Arg Pro Ile Gly Thr Arg Tyr Leu Thr Arg Pro
Leu 725 730 7352735PRTAdeno associated virus 2 2Met Ala Ala Asp Gly
Tyr Leu Pro Asp Trp Leu Glu Asp Thr Leu Ser1 5 10 15Glu Gly Ile Arg
Gln Trp Trp Lys Leu Lys Pro Gly Pro Pro Pro Pro 20 25 30Lys Pro Ala
Glu Arg His Lys Asp Asp Ser Arg Gly Leu Val Leu Pro 35 40 45Gly Tyr
Lys Tyr Leu Gly Pro Phe Asn Gly Leu Asp Lys Gly Glu Pro 50 55 60Val
Asn Glu Ala Asp Ala Ala Ala Leu Glu His Asp Lys Ala Tyr Asp65 70 75
80Arg Gln Leu Asp Ser Gly Asp Asn Pro Tyr Leu Lys Tyr Asn His Ala
85 90 95Asp Ala Glu Phe Gln Glu Arg Leu Lys Glu Asp Thr Ser Phe Gly
Gly 100 105 110Asn Leu Gly Arg Ala Val Phe Gln Ala Lys Lys Arg Val
Leu Glu Pro 115 120 125Leu Gly Leu Val Glu Glu Pro Val Lys Thr Ala
Pro Gly Lys Lys Arg 130 135 140Pro Val Glu His Ser Pro Val Glu Pro
Asp Ser Ser Ser Gly Thr Gly145 150 155 160Lys Ala Gly Gln Gln Pro
Ala Arg Lys Arg Leu Asn Phe Gly Gln Thr 165 170 175Gly Asp Ala Asp
Ser Val Pro Asp Pro Gln Pro Leu Gly Gln Pro Pro 180 185 190Ala Ala
Pro Ser Gly Leu Gly Thr Asn Thr Met Ala Thr Gly Ser Gly 195 200
205Ala Pro Met Ala Asp Asn Asn Glu Gly Ala Asp Gly Val Gly Asn Ser
210 215 220Ser Gly Asn Trp His Cys Asp Ser Thr Trp Met Gly Asp Arg
Val Ile225 230 235 240Thr Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr
Tyr Asn Asn His Leu 245 250 255Tyr Lys Gln Ile Ser Ser Gln Ser Gly
Ala Ser Asn Asp Asn His Tyr 260 265 270Phe Gly Tyr Ser Thr Pro Trp
Gly Tyr Phe Asp Phe Asn Arg Phe His 275 280 285Cys His Phe Ser Pro
Arg Asp Trp Gln Arg Leu Ile Asn Asn Asn Trp 290 295 300Gly Phe Arg
Pro Lys Arg Leu Asn Phe Lys Leu Phe Asn Ile Gln Val305 310 315
320Lys Glu Val Thr Gln Asn Asp Gly Thr Thr Thr Ile Ala Asn Asn Leu
325 330 335Thr Ser Thr Val Gln Val Phe Thr Asp Ser Glu Tyr Gln Leu
Pro Tyr 340 345 350Val Leu Gly Ser Ala His Gln Gly Cys Leu Pro Pro
Phe Pro Ala Asp 355 360 365Val Phe Met Val Pro Gln Tyr Gly Tyr Leu
Thr Leu Asn Asn Gly Ser 370 375 380Gln Ala Val Gly Arg Ser Ser Phe
Tyr Cys Leu Glu Tyr Phe Pro Ser385 390 395 400Gln Met Leu Arg Thr
Gly Asn Asn Phe Thr Phe Ser Tyr Thr Phe Glu 405 410 415Asp Val Pro
Phe His Ser Ser Tyr Ala His Ser Gln Ser Leu Asp Arg 420 425 430Leu
Met Asn Pro Leu Ile Asp Gln Tyr Leu Tyr Tyr Leu Ser Arg Thr 435 440
445Asn Thr Pro Ser Gly Thr Thr Thr Gln Ser Arg Leu Gln Phe Ser Gln
450 455 460Ala Gly Ala Ser Asp Ile Arg Asp Gln Ser Arg Asn Trp Leu
Pro Gly465 470 475 480Pro Cys Tyr Arg Gln Gln Arg Val Ser Lys Thr
Ser Ala Asp Asn Asn 485 490 495Asn Ser Glu Tyr Ser Trp Thr Gly Ala
Thr Lys Tyr His Leu Asn Gly 500 505 510Arg Asp Ser Leu Val Asn Pro
Gly Pro Ala Met Ala Ser His Lys Asp 515 520 525Asp Glu Glu Lys Phe
Phe Pro Gln Ser Gly Val Leu Ile Phe Gly Lys 530 535 540Gln Gly Ser
Glu Lys Thr Asn Val Asp Ile Glu Lys Val Met Ile Thr545 550 555
560Asp Glu Glu Glu Ile Arg Thr Thr Asn Pro Val Ala Thr Glu Gln Tyr
565 570 575Gly Ser Val Ser Thr Asn Leu Gln Arg Gly Asn Arg Gln Ala
Ala Thr 580 585 590Ala Asp Val Asn Thr Gln Gly Val Leu Pro Gly Met
Val Trp Gln Asp 595 600 605Arg Asp Val Tyr Leu Gln Gly Pro Ile Trp
Ala Lys Ile Pro His Thr 610 615 620Asp Gly His Phe His Pro Ser Pro
Leu Met Gly Gly Phe Gly Leu Lys625 630 635 640His Pro Pro Pro Gln
Ile Leu Ile Lys Asn Thr Pro Val Pro Ala Asn 645 650 655Pro Ser Thr
Thr Phe Ser Ala Ala Lys Phe Ala Ser Phe Ile Thr Gln 660 665 670Tyr
Ser Thr Gly Gln Val Ser Val Glu Ile Glu Trp Glu Leu Gln Lys 675 680
685Glu Asn Ser Lys Arg Trp Asn Pro Glu Ile Gln Tyr Thr Ser Asn Tyr
690 695 700Asn Lys Ser Val Asn Val Asp Phe Thr Val Asp Thr Asn Gly
Val Tyr705 710 715 720Ser Glu Pro Arg Pro Ile Gly Thr Arg Tyr Leu
Thr Arg Asn Leu 725 730 7353724PRTAdeno associated virus 5 3Met Ser
Phe Val Asp His Pro Pro Asp Trp Leu Glu Glu Val Gly Glu1 5 10 15Gly
Leu Arg Glu Phe Leu Gly Leu Glu Ala Gly Pro Pro Lys Pro Lys 20 25
30Pro Asn Gln Gln His Gln Asp Gln Ala Arg Gly Leu Val Leu Pro Gly
35 40 45Tyr Asn Tyr Leu Gly Pro Gly Asn Gly Leu Asp Arg Gly Glu Pro
Val 50 55 60Asn Arg Ala Asp Glu Val Ala Arg Glu His Asp Ile Ser Tyr
Asn Glu65 70 75 80Gln Leu Glu Ala Gly Asp Asn Pro Tyr Leu Lys Tyr
Asn His Ala Asp 85 90 95Ala Glu Phe Gln Glu Lys Leu Ala Asp Asp Thr
Ser Phe Gly Gly Asn 100 105 110Leu Gly Lys Ala Val Phe Gln Ala Lys
Lys Arg Val Leu Glu Pro Phe 115 120 125Gly Leu Val Glu Glu Gly Ala
Lys Thr Ala Pro Thr Gly Lys Arg Ile 130 135 140Asp Asp His Phe Pro
Lys Arg Lys Lys Ala Arg Thr Glu Glu Asp Ser145 150 155 160Lys Pro
Ser Thr Ser Ser Asp Ala Glu Ala Gly Pro Ser Gly Ser Gln 165 170
175Gln Leu Gln Ile Pro Ala Gln Pro Ala Ser Ser Leu Gly Ala Asp Thr
180 185 190Met Ser Ala Gly Gly Gly Gly Pro Leu Gly Asp Asn Asn Gln
Gly Ala 195 200 205Asp Gly Val Gly Asn Ala Ser Gly Asp Trp His Cys
Asp Ser Thr Trp 210 215 220Met Gly Asp Arg Val Val Thr Lys Ser Thr
Arg Thr Trp Val Leu Pro225 230 235 240Ser Tyr Asn Asn His Gln Tyr
Arg Glu Ile Lys Ser Gly Ser Val Asp 245 250 255Gly Ser Asn Ala Asn
Ala Tyr Phe Gly Tyr Ser Thr Pro Trp Gly Tyr 260 265 270Phe Asp Phe
Asn Arg Phe His Ser His Trp Ser Pro Arg Asp Trp Gln 275 280 285Arg
Leu Ile Asn Asn Tyr Trp Gly Phe Arg Pro Arg Ser Leu Arg Val 290 295
300Lys Ile Phe Asn Ile Gln Val Lys Glu Val Thr Val Gln Asp Ser
Thr305 310 315 320Thr Thr Ile Ala Asn Asn Leu Thr Ser Thr Val Gln
Val Phe Thr Asp 325 330 335Asp Asp Tyr Gln Leu Pro Tyr Val Val Gly
Asn Gly Thr Glu Gly Cys 340 345 350Leu Pro Ala Phe Pro Pro Gln Val
Phe Thr Leu Pro Gln Tyr Gly Tyr 355 360 365Ala Thr Leu Asn Arg Asp
Asn Thr Glu Asn Pro Thr Glu Arg Ser Ser 370 375 380Phe Phe Cys Leu
Glu Tyr Phe Pro Ser Lys Met Leu Arg Thr Gly Asn385 390 395 400Asn
Phe Glu Phe Thr Tyr Asn Phe Glu Glu Val Pro Phe His Ser Ser 405 410
415Phe Ala Pro Ser Gln Asn Leu Phe Lys Leu Ala Asn Pro Leu Val Asp
420 425 430Gln Tyr Leu Tyr Arg Phe Val Ser Thr Asn Asn Thr Gly Gly
Val Gln 435 440 445Phe Asn Lys Asn Leu Ala Gly Arg Tyr Ala Asn Thr
Tyr Lys Asn Trp 450 455 460Phe Pro Gly Pro Met Gly Arg Thr Gln Gly
Trp Asn Leu Gly Ser Gly465 470 475 480Val Asn Arg Ala Ser Val Ser
Ala Phe Ala Thr Thr Asn Arg Met Glu 485 490 495Leu Glu Gly Ala Ser
Tyr Gln Val Pro Pro Gln Pro Asn Gly Met Thr 500 505 510Asn Asn Leu
Gln Gly Ser Asn Thr Tyr Ala Leu Glu Asn Thr Met Ile 515 520 525Phe
Asn Ser Gln Pro Ala Asn Pro Gly Thr Thr Ala Thr Tyr Leu Glu 530 535
540Gly Asn Met Leu Ile Thr Ser Glu Ser Glu Thr Gln Pro Val Asn
Arg545 550 555 560Val Ala Tyr Asn Val Gly Gly Gln Met Ala Thr Asn
Asn Gln Ser Ser 565 570 575Thr Thr Ala Pro Ala Thr Gly Thr Tyr Asn
Leu Gln Glu Ile Val Pro 580 585 590Gly Ser Val Trp Met Glu Arg Asp
Val Tyr Leu Gln Gly Pro Ile Trp 595 600 605Ala Lys Ile Pro Glu Thr
Gly Ala His Phe His Pro Ser Pro Ala Met 610 615 620Gly Gly Phe Gly
Leu Lys His Pro Pro Pro Met Met Leu Ile Lys Asn625 630 635 640Thr
Pro Val Pro Gly Asn Ile Thr Ser Phe Ser Asp Val Pro Val Ser 645 650
655Ser Phe Ile Thr Gln Tyr Ser Thr Gly Gln Val Thr Val Glu Met Glu
660 665 670Trp Glu Leu Lys Lys Glu Asn Ser Lys Arg Trp Asn Pro Glu
Ile Gln 675 680 685Tyr Thr Asn Asn Tyr Asn Asp Pro Gln Phe Val Asp
Phe Ala Pro Asp 690 695 700Ser Thr Gly Glu Tyr Arg Thr Thr Arg Pro
Ile Gly Thr Arg Tyr Leu705 710 715 720Thr Arg Pro Leu4736PRTAdeno
associated virus 6 4Met Ala Ala Asp Gly Tyr Leu Pro Asp Trp Leu Glu
Asp Asn Leu Ser1 5 10 15Glu Gly Ile Arg Glu Trp Trp Asp Leu Lys Pro
Gly Ala Pro Lys Pro 20 25 30Lys Ala Asn Gln Gln Lys Gln Asp Asp Gly
Arg Gly Leu Val Leu Pro 35 40 45Gly Tyr Lys Tyr Leu Gly Pro Phe Asn
Gly Leu Asp Lys Gly Glu Pro 50 55 60Val Asn Ala Ala Asp Ala Ala Ala
Leu Glu His Asp Lys Ala Tyr Asp65 70 75 80Gln Gln Leu Lys Ala Gly
Asp Asn Pro Tyr Leu Arg Tyr Asn His Ala 85 90 95Asp Ala Glu Phe Gln
Glu Arg Leu Gln Glu Asp Thr Ser Phe Gly Gly 100 105 110Asn Leu Gly
Arg Ala Val Phe Gln Ala Lys Lys Arg Val Leu Glu Pro 115 120 125Phe
Gly Leu Val Glu Glu Gly Ala Lys Thr Ala Pro Gly Lys Lys Arg 130 135
140Pro Val Glu Gln Ser Pro Gln Glu Pro Asp Ser Ser Ser Gly Ile
Gly145 150 155 160Lys Thr Gly Gln Gln Pro Ala Lys Lys Arg Leu Asn
Phe Gly Gln Thr 165 170 175Gly Asp Ser Glu Ser Val Pro Asp Pro Gln
Pro Leu Gly Glu Pro Pro 180 185 190Ala Thr Pro Ala Ala Val Gly Pro
Thr Thr Met Ala Ser Gly Gly Gly 195 200 205Ala Pro Met Ala Asp Asn
Asn Glu Gly Ala Asp Gly Val Gly Asn Ala 210 215 220Ser Gly Asn Trp
His Cys Asp Ser Thr Trp Leu Gly Asp Arg Val Ile225 230 235 240Thr
Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr Tyr Asn Asn His Leu 245 250
255Tyr Lys Gln Ile Ser Ser Ala Ser Thr Gly Ala Ser Asn Asp Asn
His
260 265 270Tyr Phe Gly Tyr Ser Thr Pro Trp Gly Tyr Phe Asp Phe Asn
Arg Phe 275 280 285His Cys His Phe Ser Pro Arg Asp Trp Gln Arg Leu
Ile Asn Asn Asn 290 295 300Trp Gly Phe Arg Pro Lys Arg Leu Asn Phe
Lys Leu Phe Asn Ile Gln305 310 315 320Val Lys Glu Val Thr Thr Asn
Asp Gly Val Thr Thr Ile Ala Asn Asn 325 330 335Leu Thr Ser Thr Val
Gln Val Phe Ser Asp Ser Glu Tyr Gln Leu Pro 340 345 350Tyr Val Leu
Gly Ser Ala His Gln Gly Cys Leu Pro Pro Phe Pro Ala 355 360 365Asp
Val Phe Met Ile Pro Gln Tyr Gly Tyr Leu Thr Leu Asn Asn Gly 370 375
380Ser Gln Ala Val Gly Arg Ser Ser Phe Tyr Cys Leu Glu Tyr Phe
Pro385 390 395 400Ser Gln Met Leu Arg Thr Gly Asn Asn Phe Thr Phe
Ser Tyr Thr Phe 405 410 415Glu Asp Val Pro Phe His Ser Ser Tyr Ala
His Ser Gln Ser Leu Asp 420 425 430Arg Leu Met Asn Pro Leu Ile Asp
Gln Tyr Leu Tyr Tyr Leu Asn Arg 435 440 445Thr Gln Asn Gln Ser Gly
Ser Ala Gln Asn Lys Asp Leu Leu Phe Ser 450 455 460Arg Gly Ser Pro
Ala Gly Met Ser Val Gln Pro Lys Asn Trp Leu Pro465 470 475 480Gly
Pro Cys Tyr Arg Gln Gln Arg Val Ser Lys Thr Lys Thr Asp Asn 485 490
495Asn Asn Ser Asn Phe Thr Trp Thr Gly Ala Ser Lys Tyr Asn Leu Asn
500 505 510Gly Arg Glu Ser Ile Ile Asn Pro Gly Thr Ala Met Ala Ser
His Lys 515 520 525Asp Asp Lys Asp Lys Phe Phe Pro Met Ser Gly Val
Met Ile Phe Gly 530 535 540Lys Glu Ser Ala Gly Ala Ser Asn Thr Ala
Leu Asp Asn Val Met Ile545 550 555 560Thr Asp Glu Glu Glu Ile Lys
Ala Thr Asn Pro Val Ala Thr Glu Arg 565 570 575Phe Gly Thr Val Ala
Val Asn Leu Gln Ser Ser Ser Thr Asp Pro Ala 580 585 590Thr Gly Asp
Val His Val Met Gly Ala Leu Pro Gly Met Val Trp Gln 595 600 605Asp
Arg Asp Val Tyr Leu Gln Gly Pro Ile Trp Ala Lys Ile Pro His 610 615
620Thr Asp Gly His Phe His Pro Ser Pro Leu Met Gly Gly Phe Gly
Leu625 630 635 640Lys His Pro Pro Pro Gln Ile Leu Ile Lys Asn Thr
Pro Val Pro Ala 645 650 655Asn Pro Pro Ala Glu Phe Ser Ala Thr Lys
Phe Ala Ser Phe Ile Thr 660 665 670Gln Tyr Ser Thr Gly Gln Val Ser
Val Glu Ile Glu Trp Glu Leu Gln 675 680 685Lys Glu Asn Ser Lys Arg
Trp Asn Pro Glu Val Gln Tyr Thr Ser Asn 690 695 700Tyr Ala Lys Ser
Ala Asn Val Asp Phe Thr Val Asp Asn Asn Gly Leu705 710 715 720Tyr
Thr Glu Pro Arg Pro Ile Gly Thr Arg Tyr Leu Thr Arg Pro Leu 725 730
7355736PRTAdeno associated virus 6.2 5Met Ala Ala Asp Gly Tyr Leu
Pro Asp Trp Leu Glu Asp Asn Leu Ser1 5 10 15Glu Gly Ile Arg Glu Trp
Trp Asp Leu Lys Pro Gly Ala Pro Lys Pro 20 25 30Lys Ala Asn Gln Gln
Lys Gln Asp Asp Gly Arg Gly Leu Val Leu Pro 35 40 45Gly Tyr Lys Tyr
Leu Gly Pro Phe Asn Gly Leu Asp Lys Gly Glu Pro 50 55 60Val Asn Ala
Ala Asp Ala Ala Ala Leu Glu His Asp Lys Ala Tyr Asp65 70 75 80Gln
Gln Leu Lys Ala Gly Asp Asn Pro Tyr Leu Arg Tyr Asn His Ala 85 90
95Asp Ala Glu Phe Gln Glu Arg Leu Gln Glu Asp Thr Ser Phe Gly Gly
100 105 110Asn Leu Gly Arg Ala Val Phe Gln Ala Lys Lys Arg Val Leu
Glu Pro 115 120 125Leu Gly Leu Val Glu Glu Gly Ala Lys Thr Ala Pro
Gly Lys Lys Arg 130 135 140Pro Val Glu Gln Ser Pro Gln Glu Pro Asp
Ser Ser Ser Gly Ile Gly145 150 155 160Lys Thr Gly Gln Gln Pro Ala
Lys Lys Arg Leu Asn Phe Gly Gln Thr 165 170 175Gly Asp Ser Glu Ser
Val Pro Asp Pro Gln Pro Leu Gly Glu Pro Pro 180 185 190Ala Thr Pro
Ala Ala Val Gly Pro Thr Thr Met Ala Ser Gly Gly Gly 195 200 205Ala
Pro Met Ala Asp Asn Asn Glu Gly Ala Asp Gly Val Gly Asn Ala 210 215
220Ser Gly Asn Trp His Cys Asp Ser Thr Trp Leu Gly Asp Arg Val
Ile225 230 235 240Thr Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr Tyr
Asn Asn His Leu 245 250 255Tyr Lys Gln Ile Ser Ser Ala Ser Thr Gly
Ala Ser Asn Asp Asn His 260 265 270Tyr Phe Gly Tyr Ser Thr Pro Trp
Gly Tyr Phe Asp Phe Asn Arg Phe 275 280 285His Cys His Phe Ser Pro
Arg Asp Trp Gln Arg Leu Ile Asn Asn Asn 290 295 300Trp Gly Phe Arg
Pro Lys Arg Leu Asn Phe Lys Leu Phe Asn Ile Gln305 310 315 320Val
Lys Glu Val Thr Thr Asn Asp Gly Val Thr Thr Ile Ala Asn Asn 325 330
335Leu Thr Ser Thr Val Gln Val Phe Ser Asp Ser Glu Tyr Gln Leu Pro
340 345 350Tyr Val Leu Gly Ser Ala His Gln Gly Cys Leu Pro Pro Phe
Pro Ala 355 360 365Asp Val Phe Met Ile Pro Gln Tyr Gly Tyr Leu Thr
Leu Asn Asn Gly 370 375 380Ser Gln Ala Val Gly Arg Ser Ser Phe Tyr
Cys Leu Glu Tyr Phe Pro385 390 395 400Ser Gln Met Leu Arg Thr Gly
Asn Asn Phe Thr Phe Ser Tyr Thr Phe 405 410 415Glu Asp Val Pro Phe
His Ser Ser Tyr Ala His Ser Gln Ser Leu Asp 420 425 430Arg Leu Met
Asn Pro Leu Ile Asp Gln Tyr Leu Tyr Tyr Leu Asn Arg 435 440 445Thr
Gln Asn Gln Ser Gly Ser Ala Gln Asn Lys Asp Leu Leu Phe Ser 450 455
460Arg Gly Ser Pro Ala Gly Met Ser Val Gln Pro Lys Asn Trp Leu
Pro465 470 475 480Gly Pro Cys Tyr Arg Gln Gln Arg Val Ser Lys Thr
Lys Thr Asp Asn 485 490 495Asn Asn Ser Asn Phe Thr Trp Thr Gly Ala
Ser Lys Tyr Asn Leu Asn 500 505 510Gly Arg Glu Ser Ile Ile Asn Pro
Gly Thr Ala Met Ala Ser His Lys 515 520 525Asp Asp Lys Asp Lys Phe
Phe Pro Met Ser Gly Val Met Ile Phe Gly 530 535 540Lys Glu Ser Ala
Gly Ala Ser Asn Thr Ala Leu Asp Asn Val Met Ile545 550 555 560Thr
Asp Glu Glu Glu Ile Lys Ala Thr Asn Pro Val Ala Thr Glu Arg 565 570
575Phe Gly Thr Val Ala Val Asn Leu Gln Ser Ser Ser Thr Asp Pro Ala
580 585 590Thr Gly Asp Val His Val Met Gly Ala Leu Pro Gly Met Val
Trp Gln 595 600 605Asp Arg Asp Val Tyr Leu Gln Gly Pro Ile Trp Ala
Lys Ile Pro His 610 615 620Thr Asp Gly His Phe His Pro Ser Pro Leu
Met Gly Gly Phe Gly Leu625 630 635 640Lys His Pro Pro Pro Gln Ile
Leu Ile Lys Asn Thr Pro Val Pro Ala 645 650 655Asn Pro Pro Ala Glu
Phe Ser Ala Thr Lys Phe Ala Ser Phe Ile Thr 660 665 670Gln Tyr Ser
Thr Gly Gln Val Ser Val Glu Ile Glu Trp Glu Leu Gln 675 680 685Lys
Glu Asn Ser Lys Arg Trp Asn Pro Glu Val Gln Tyr Thr Ser Asn 690 695
700Tyr Ala Lys Ser Ala Asn Val Asp Phe Thr Val Asp Asn Asn Gly
Leu705 710 715 720Tyr Thr Glu Pro Arg Pro Ile Gly Thr Arg Tyr Leu
Thr Arg Pro Leu 725 730 7356737PRTAdeno associated virus 7 6Met Ala
Ala Asp Gly Tyr Leu Pro Asp Trp Leu Glu Asp Asn Leu Ser1 5 10 15Glu
Gly Ile Arg Glu Trp Trp Asp Leu Lys Pro Gly Ala Pro Lys Pro 20 25
30Lys Ala Asn Gln Gln Lys Gln Asp Asn Gly Arg Gly Leu Val Leu Pro
35 40 45Gly Tyr Lys Tyr Leu Gly Pro Phe Asn Gly Leu Asp Lys Gly Glu
Pro 50 55 60Val Asn Ala Ala Asp Ala Ala Ala Leu Glu His Asp Lys Ala
Tyr Asp65 70 75 80Gln Gln Leu Lys Ala Gly Asp Asn Pro Tyr Leu Arg
Tyr Asn His Ala 85 90 95Asp Ala Glu Phe Gln Glu Arg Leu Gln Glu Asp
Thr Ser Phe Gly Gly 100 105 110Asn Leu Gly Arg Ala Val Phe Gln Ala
Lys Lys Arg Val Leu Glu Pro 115 120 125Leu Gly Leu Val Glu Glu Gly
Ala Lys Thr Ala Pro Ala Lys Lys Arg 130 135 140Pro Val Glu Pro Ser
Pro Gln Arg Ser Pro Asp Ser Ser Thr Gly Ile145 150 155 160Gly Lys
Lys Gly Gln Gln Pro Ala Arg Lys Arg Leu Asn Phe Gly Gln 165 170
175Thr Gly Asp Ser Glu Ser Val Pro Asp Pro Gln Pro Leu Gly Glu Pro
180 185 190Pro Ala Ala Pro Ser Ser Val Gly Ser Gly Thr Val Ala Ala
Gly Gly 195 200 205Gly Ala Pro Met Ala Asp Asn Asn Glu Gly Ala Asp
Gly Val Gly Asn 210 215 220Ala Ser Gly Asn Trp His Cys Asp Ser Thr
Trp Leu Gly Asp Arg Val225 230 235 240Ile Thr Thr Ser Thr Arg Thr
Trp Ala Leu Pro Thr Tyr Asn Asn His 245 250 255Leu Tyr Lys Gln Ile
Ser Ser Glu Thr Ala Gly Ser Thr Asn Asp Asn 260 265 270Thr Tyr Phe
Gly Tyr Ser Thr Pro Trp Gly Tyr Phe Asp Phe Asn Arg 275 280 285Phe
His Cys His Phe Ser Pro Arg Asp Trp Gln Arg Leu Ile Asn Asn 290 295
300Asn Trp Gly Phe Arg Pro Lys Lys Leu Arg Phe Lys Leu Phe Asn
Ile305 310 315 320Gln Val Lys Glu Val Thr Thr Asn Asp Gly Val Thr
Thr Ile Ala Asn 325 330 335Asn Leu Thr Ser Thr Ile Gln Val Phe Ser
Asp Ser Glu Tyr Gln Leu 340 345 350Pro Tyr Val Leu Gly Ser Ala His
Gln Gly Cys Leu Pro Pro Phe Pro 355 360 365Ala Asp Val Phe Met Ile
Pro Gln Tyr Gly Tyr Leu Thr Leu Asn Asn 370 375 380Gly Ser Gln Ser
Val Gly Arg Ser Ser Phe Tyr Cys Leu Glu Tyr Phe385 390 395 400Pro
Ser Gln Met Leu Arg Thr Gly Asn Asn Phe Glu Phe Ser Tyr Ser 405 410
415Phe Glu Asp Val Pro Phe His Ser Ser Tyr Ala His Ser Gln Ser Leu
420 425 430Asp Arg Leu Met Asn Pro Leu Ile Asp Gln Tyr Leu Tyr Tyr
Leu Ala 435 440 445Arg Thr Gln Ser Asn Pro Gly Gly Thr Ala Gly Asn
Arg Glu Leu Gln 450 455 460Phe Tyr Gln Gly Gly Pro Ser Thr Met Ala
Glu Gln Ala Lys Asn Trp465 470 475 480Leu Pro Gly Pro Cys Phe Arg
Gln Gln Arg Val Ser Lys Thr Leu Asp 485 490 495Gln Asn Asn Asn Ser
Asn Phe Ala Trp Thr Gly Ala Thr Lys Tyr His 500 505 510Leu Asn Gly
Arg Asn Ser Leu Val Asn Pro Gly Val Ala Met Ala Thr 515 520 525His
Lys Asp Asp Glu Asp Arg Phe Phe Pro Ser Ser Gly Val Leu Ile 530 535
540Phe Gly Lys Thr Gly Ala Thr Asn Lys Thr Thr Leu Glu Asn Val
Leu545 550 555 560Met Thr Asn Glu Glu Glu Ile Arg Pro Thr Asn Pro
Val Ala Thr Glu 565 570 575Glu Tyr Gly Ile Val Ser Ser Asn Leu Gln
Ala Ala Asn Thr Ala Ala 580 585 590Gln Thr Gln Val Val Asn Asn Gln
Gly Ala Leu Pro Gly Met Val Trp 595 600 605Gln Asn Arg Asp Val Tyr
Leu Gln Gly Pro Ile Trp Ala Lys Ile Pro 610 615 620His Thr Asp Gly
Asn Phe His Pro Ser Pro Leu Met Gly Gly Phe Gly625 630 635 640Leu
Lys His Pro Pro Pro Gln Ile Leu Ile Lys Asn Thr Pro Val Pro 645 650
655Ala Asn Pro Pro Glu Val Phe Thr Pro Ala Lys Phe Ala Ser Phe Ile
660 665 670Thr Gln Tyr Ser Thr Gly Gln Val Ser Val Glu Ile Glu Trp
Glu Leu 675 680 685Gln Lys Glu Asn Ser Lys Arg Trp Asn Pro Glu Ile
Gln Tyr Thr Ser 690 695 700Asn Phe Glu Lys Gln Thr Gly Val Asp Phe
Ala Val Asp Ser Gln Gly705 710 715 720Val Tyr Ser Glu Pro Arg Pro
Ile Gly Thr Arg Tyr Leu Thr Arg Asn 725 730 735Leu7738PRTAdeno
associated virus 8 7Met Ala Ala Asp Gly Tyr Leu Pro Asp Trp Leu Glu
Asp Asn Leu Ser1 5 10 15Glu Gly Ile Arg Glu Trp Trp Ala Leu Lys Pro
Gly Ala Pro Lys Pro 20 25 30Lys Ala Asn Gln Gln Lys Gln Asp Asp Gly
Arg Gly Leu Val Leu Pro 35 40 45Gly Tyr Lys Tyr Leu Gly Pro Phe Asn
Gly Leu Asp Lys Gly Glu Pro 50 55 60Val Asn Ala Ala Asp Ala Ala Ala
Leu Glu His Asp Lys Ala Tyr Asp65 70 75 80Gln Gln Leu Gln Ala Gly
Asp Asn Pro Tyr Leu Arg Tyr Asn His Ala 85 90 95Asp Ala Glu Phe Gln
Glu Arg Leu Gln Glu Asp Thr Ser Phe Gly Gly 100 105 110Asn Leu Gly
Arg Ala Val Phe Gln Ala Lys Lys Arg Val Leu Glu Pro 115 120 125Leu
Gly Leu Val Glu Glu Gly Ala Lys Thr Ala Pro Gly Lys Lys Arg 130 135
140Pro Val Glu Pro Ser Pro Gln Arg Ser Pro Asp Ser Ser Thr Gly
Ile145 150 155 160Gly Lys Lys Gly Gln Gln Pro Ala Arg Lys Arg Leu
Asn Phe Gly Gln 165 170 175Thr Gly Asp Ser Glu Ser Val Pro Asp Pro
Gln Pro Leu Gly Glu Pro 180 185 190Pro Ala Ala Pro Ser Gly Val Gly
Pro Asn Thr Met Ala Ala Gly Gly 195 200 205Gly Ala Pro Met Ala Asp
Asn Asn Glu Gly Ala Asp Gly Val Gly Ser 210 215 220Ser Ser Gly Asn
Trp His Cys Asp Ser Thr Trp Leu Gly Asp Arg Val225 230 235 240Ile
Thr Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr Tyr Asn Asn His 245 250
255Leu Tyr Lys Gln Ile Ser Asn Gly Thr Ser Gly Gly Ala Thr Asn Asp
260 265 270Asn Thr Tyr Phe Gly Tyr Ser Thr Pro Trp Gly Tyr Phe Asp
Phe Asn 275 280 285Arg Phe His Cys His Phe Ser Pro Arg Asp Trp Gln
Arg Leu Ile Asn 290 295 300Asn Asn Trp Gly Phe Arg Pro Lys Arg Leu
Ser Phe Lys Leu Phe Asn305 310 315 320Ile Gln Val Lys Glu Val Thr
Gln Asn Glu Gly Thr Lys Thr Ile Ala 325 330 335Asn Asn Leu Thr Ser
Thr Ile Gln Val Phe Thr Asp Ser Glu Tyr Gln 340 345 350Leu Pro Tyr
Val Leu Gly Ser Ala His Gln Gly Cys Leu Pro Pro Phe 355 360 365Pro
Ala Asp Val Phe Met Ile Pro Gln Tyr Gly Tyr Leu Thr Leu Asn 370 375
380Asn Gly Ser Gln Ala Val Gly Arg Ser Ser Phe Tyr Cys Leu Glu
Tyr385 390 395 400Phe Pro Ser Gln Met Leu Arg Thr Gly Asn Asn Phe
Gln Phe Thr Tyr 405 410 415Thr Phe Glu Asp Val Pro Phe His Ser Ser
Tyr Ala His Ser Gln Ser 420 425 430Leu Asp Arg Leu Met Asn Pro Leu
Ile Asp Gln Tyr Leu Tyr Tyr Leu 435 440 445Ser Arg Thr Gln Thr Thr
Gly Gly Thr Ala Asn Thr Gln Thr Leu Gly 450 455 460Phe Ser Gln Gly
Gly Pro Asn Thr Met Ala Asn Gln Ala Lys Asn Trp465 470 475 480Leu
Pro Gly Pro Cys Tyr Arg Gln Gln Arg Val Ser Thr Thr Thr Gly 485 490
495Gln Asn Asn Asn Ser Asn Phe Ala Trp Thr Ala Gly Thr Lys Tyr His
500 505 510Leu Asn Gly Arg Asn Ser Leu Ala Asn Pro
Gly Ile Ala Met Ala Thr 515 520 525His Lys Asp Asp Glu Glu Arg Phe
Phe Pro Ser Asn Gly Ile Leu Ile 530 535 540Phe Gly Lys Gln Asn Ala
Ala Arg Asp Asn Ala Asp Tyr Ser Asp Val545 550 555 560Met Leu Thr
Ser Glu Glu Glu Ile Lys Thr Thr Asn Pro Val Ala Thr 565 570 575Glu
Glu Tyr Gly Ile Val Ala Asp Asn Leu Gln Gln Gln Asn Thr Ala 580 585
590Pro Gln Ile Gly Thr Val Asn Ser Gln Gly Ala Leu Pro Gly Met Val
595 600 605Trp Gln Asn Arg Asp Val Tyr Leu Gln Gly Pro Ile Trp Ala
Lys Ile 610 615 620Pro His Thr Asp Gly Asn Phe His Pro Ser Pro Leu
Met Gly Gly Phe625 630 635 640Gly Leu Lys His Pro Pro Pro Gln Ile
Leu Ile Lys Asn Thr Pro Val 645 650 655Pro Ala Asp Pro Pro Thr Thr
Phe Asn Gln Ser Lys Leu Asn Ser Phe 660 665 670Ile Thr Gln Tyr Ser
Thr Gly Gln Val Ser Val Glu Ile Glu Trp Glu 675 680 685Leu Gln Lys
Glu Asn Ser Lys Arg Trp Asn Pro Glu Ile Gln Tyr Thr 690 695 700Ser
Asn Tyr Tyr Lys Ser Thr Ser Val Asp Phe Ala Val Asn Thr Glu705 710
715 720Gly Val Tyr Ser Glu Pro Arg Pro Ile Gly Thr Arg Tyr Leu Thr
Arg 725 730 735Asn Leu8736PRTAdeno associated virus 9 8Met Ala Ala
Asp Gly Tyr Leu Pro Asp Trp Leu Glu Asp Asn Leu Ser1 5 10 15Glu Gly
Ile Arg Glu Trp Trp Ala Leu Lys Pro Gly Ala Pro Gln Pro 20 25 30Lys
Ala Asn Gln Gln His Gln Asp Asn Ala Arg Gly Leu Val Leu Pro 35 40
45Gly Tyr Lys Tyr Leu Gly Pro Gly Asn Gly Leu Asp Lys Gly Glu Pro
50 55 60Val Asn Ala Ala Asp Ala Ala Ala Leu Glu His Asp Lys Ala Tyr
Asp65 70 75 80Gln Gln Leu Lys Ala Gly Asp Asn Pro Tyr Leu Lys Tyr
Asn His Ala 85 90 95Asp Ala Glu Phe Gln Glu Arg Leu Lys Glu Asp Thr
Ser Phe Gly Gly 100 105 110Asn Leu Gly Arg Ala Val Phe Gln Ala Lys
Lys Arg Leu Leu Glu Pro 115 120 125Leu Gly Leu Val Glu Glu Ala Ala
Lys Thr Ala Pro Gly Lys Lys Arg 130 135 140Pro Val Glu Gln Ser Pro
Gln Glu Pro Asp Ser Ser Ala Gly Ile Gly145 150 155 160Lys Ser Gly
Ala Gln Pro Ala Lys Lys Arg Leu Asn Phe Gly Gln Thr 165 170 175Gly
Asp Thr Glu Ser Val Pro Asp Pro Gln Pro Ile Gly Glu Pro Pro 180 185
190Ala Ala Pro Ser Gly Val Gly Ser Leu Thr Met Ala Ser Gly Gly Gly
195 200 205Ala Pro Val Ala Asp Asn Asn Glu Gly Ala Asp Gly Val Gly
Ser Ser 210 215 220Ser Gly Asn Trp His Cys Asp Ser Gln Trp Leu Gly
Asp Arg Val Ile225 230 235 240Thr Thr Ser Thr Arg Thr Trp Ala Leu
Pro Thr Tyr Asn Asn His Leu 245 250 255Tyr Lys Gln Ile Ser Asn Ser
Thr Ser Gly Gly Ser Ser Asn Asp Asn 260 265 270Ala Tyr Phe Gly Tyr
Ser Thr Pro Trp Gly Tyr Phe Asp Phe Asn Arg 275 280 285Phe His Cys
His Phe Ser Pro Arg Asp Trp Gln Arg Leu Ile Asn Asn 290 295 300Asn
Trp Gly Phe Arg Pro Lys Arg Leu Asn Phe Lys Leu Phe Asn Ile305 310
315 320Gln Val Lys Glu Val Thr Asp Asn Asn Gly Val Lys Thr Ile Ala
Asn 325 330 335Asn Leu Thr Ser Thr Val Gln Val Phe Thr Asp Ser Asp
Tyr Gln Leu 340 345 350Pro Tyr Val Leu Gly Ser Ala His Glu Gly Cys
Leu Pro Pro Phe Pro 355 360 365Ala Asp Val Phe Met Ile Pro Gln Tyr
Gly Tyr Leu Thr Leu Asn Asp 370 375 380Gly Ser Gln Ala Val Gly Arg
Ser Ser Phe Tyr Cys Leu Glu Tyr Phe385 390 395 400Pro Ser Gln Met
Leu Arg Thr Gly Asn Asn Phe Gln Phe Ser Tyr Glu 405 410 415Phe Glu
Asn Val Pro Phe His Ser Ser Tyr Ala His Ser Gln Ser Leu 420 425
430Asp Arg Leu Met Asn Pro Leu Ile Asp Gln Tyr Leu Tyr Tyr Leu Ser
435 440 445Lys Thr Ile Asn Gly Ser Gly Gln Asn Gln Gln Thr Leu Lys
Phe Ser 450 455 460Val Ala Gly Pro Ser Asn Met Ala Val Gln Gly Arg
Asn Tyr Ile Pro465 470 475 480Gly Pro Ser Tyr Arg Gln Gln Arg Val
Ser Thr Thr Val Thr Gln Asn 485 490 495Asn Asn Ser Glu Phe Ala Trp
Pro Gly Ala Ser Ser Trp Ala Leu Asn 500 505 510Gly Arg Asn Ser Leu
Met Asn Pro Gly Pro Ala Met Ala Ser His Lys 515 520 525Glu Gly Glu
Asp Arg Phe Phe Pro Leu Ser Gly Ser Leu Ile Phe Gly 530 535 540Lys
Gln Gly Thr Gly Arg Asp Asn Val Asp Ala Asp Lys Val Met Ile545 550
555 560Thr Asn Glu Glu Glu Ile Lys Thr Thr Asn Pro Val Ala Thr Glu
Ser 565 570 575Tyr Gly Gln Val Ala Thr Asn His Gln Ser Ala Gln Ala
Gln Ala Gln 580 585 590Thr Gly Trp Val Gln Asn Gln Gly Ile Leu Pro
Gly Met Val Trp Gln 595 600 605Asp Arg Asp Val Tyr Leu Gln Gly Pro
Ile Trp Ala Lys Ile Pro His 610 615 620Thr Asp Gly Asn Phe His Pro
Ser Pro Leu Met Gly Gly Phe Gly Met625 630 635 640Lys His Pro Pro
Pro Gln Ile Leu Ile Lys Asn Thr Pro Val Pro Ala 645 650 655Asp Pro
Pro Thr Ala Phe Asn Lys Asp Lys Leu Asn Ser Phe Ile Thr 660 665
670Gln Tyr Ser Thr Gly Gln Val Ser Val Glu Ile Glu Trp Glu Leu Gln
675 680 685Lys Glu Asn Ser Lys Arg Trp Asn Pro Glu Ile Gln Tyr Thr
Ser Asn 690 695 700Tyr Tyr Lys Ser Asn Asn Val Glu Phe Ala Val Asn
Thr Glu Gly Val705 710 715 720Tyr Ser Glu Pro Arg Pro Ile Gly Thr
Arg Tyr Leu Thr Arg Asn Leu 725 730 7359738PRTAdeno associated
virus rh.10 9Met Ala Ala Asp Gly Tyr Leu Pro Asp Trp Leu Glu Asp
Asn Leu Ser1 5 10 15Glu Gly Ile Arg Glu Trp Trp Asp Leu Lys Pro Gly
Ala Pro Lys Pro 20 25 30Lys Ala Asn Gln Gln Lys Gln Asp Asp Gly Arg
Gly Leu Val Leu Pro 35 40 45Gly Tyr Lys Tyr Leu Gly Pro Phe Asn Gly
Leu Asp Lys Gly Glu Pro 50 55 60Val Asn Ala Ala Asp Ala Ala Ala Leu
Glu His Asp Lys Ala Tyr Asp65 70 75 80Gln Gln Leu Lys Ala Gly Asp
Asn Pro Tyr Leu Arg Tyr Asn His Ala 85 90 95Asp Ala Glu Phe Gln Glu
Arg Leu Gln Glu Asp Thr Ser Phe Gly Gly 100 105 110Asn Leu Gly Arg
Ala Val Phe Gln Ala Lys Lys Arg Val Leu Glu Pro 115 120 125Leu Gly
Leu Val Glu Glu Gly Ala Lys Thr Ala Pro Gly Lys Lys Arg 130 135
140Pro Val Glu Pro Ser Pro Gln Arg Ser Pro Asp Ser Ser Thr Gly
Ile145 150 155 160Gly Lys Lys Gly Gln Gln Pro Ala Lys Lys Arg Leu
Asn Phe Gly Gln 165 170 175Thr Gly Asp Ser Glu Ser Val Pro Asp Pro
Gln Pro Ile Gly Glu Pro 180 185 190Pro Ala Gly Pro Ser Gly Leu Gly
Ser Gly Thr Met Ala Ala Gly Gly 195 200 205Gly Ala Pro Met Ala Asp
Asn Asn Glu Gly Ala Asp Gly Val Gly Ser 210 215 220Ser Ser Gly Asn
Trp His Cys Asp Ser Thr Trp Leu Gly Asp Arg Val225 230 235 240Ile
Thr Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr Tyr Asn Asn His 245 250
255Leu Tyr Lys Gln Ile Ser Asn Gly Thr Ser Gly Gly Ser Thr Asn Asp
260 265 270Asn Thr Tyr Phe Gly Tyr Ser Thr Pro Trp Gly Tyr Phe Asp
Phe Asn 275 280 285Arg Phe His Cys His Phe Ser Pro Arg Asp Trp Gln
Arg Leu Ile Asn 290 295 300Asn Asn Trp Gly Phe Arg Pro Lys Arg Leu
Asn Phe Lys Leu Phe Asn305 310 315 320Ile Gln Val Lys Glu Val Thr
Gln Asn Glu Gly Thr Lys Thr Ile Ala 325 330 335Asn Asn Leu Thr Ser
Thr Ile Gln Val Phe Thr Asp Ser Glu Tyr Gln 340 345 350Leu Pro Tyr
Val Leu Gly Ser Ala His Gln Gly Cys Leu Pro Pro Phe 355 360 365Pro
Ala Asp Val Phe Met Ile Pro Gln Tyr Gly Tyr Leu Thr Leu Asn 370 375
380Asn Gly Ser Gln Ala Val Gly Arg Ser Ser Phe Tyr Cys Leu Glu
Tyr385 390 395 400Phe Pro Ser Gln Met Leu Arg Thr Gly Asn Asn Phe
Glu Phe Ser Tyr 405 410 415Gln Phe Glu Asp Val Pro Phe His Ser Ser
Tyr Ala His Ser Gln Ser 420 425 430Leu Asp Arg Leu Met Asn Pro Leu
Ile Asp Gln Tyr Leu Tyr Tyr Leu 435 440 445Ser Arg Thr Gln Ser Thr
Gly Gly Thr Ala Gly Thr Gln Gln Leu Leu 450 455 460Phe Ser Gln Ala
Gly Pro Asn Asn Met Ser Ala Gln Ala Lys Asn Trp465 470 475 480Leu
Pro Gly Pro Cys Tyr Arg Gln Gln Arg Val Ser Thr Thr Leu Ser 485 490
495Gln Asn Asn Asn Ser Asn Phe Ala Trp Thr Gly Ala Thr Lys Tyr His
500 505 510Leu Asn Gly Arg Asp Ser Leu Val Asn Pro Gly Val Ala Met
Ala Thr 515 520 525His Lys Asp Asp Glu Glu Arg Phe Phe Pro Ser Ser
Gly Val Leu Met 530 535 540Phe Gly Lys Gln Gly Ala Gly Lys Asp Asn
Val Asp Tyr Ser Ser Val545 550 555 560Met Leu Thr Ser Glu Glu Glu
Ile Lys Thr Thr Asn Pro Val Ala Thr 565 570 575Glu Gln Tyr Gly Val
Val Ala Asp Asn Leu Gln Gln Gln Asn Ala Ala 580 585 590Pro Ile Val
Gly Ala Val Asn Ser Gln Gly Ala Leu Pro Gly Met Val 595 600 605Trp
Gln Asn Arg Asp Val Tyr Leu Gln Gly Pro Ile Trp Ala Lys Ile 610 615
620Pro His Thr Asp Gly Asn Phe His Pro Ser Pro Leu Met Gly Gly
Phe625 630 635 640Gly Leu Lys His Pro Pro Pro Gln Ile Leu Ile Lys
Asn Thr Pro Val 645 650 655Pro Ala Asp Pro Pro Thr Thr Phe Ser Gln
Ala Lys Leu Ala Ser Phe 660 665 670Ile Thr Gln Tyr Ser Thr Gly Gln
Val Ser Val Glu Ile Glu Trp Glu 675 680 685Leu Gln Lys Glu Asn Ser
Lys Arg Trp Asn Pro Glu Ile Gln Tyr Thr 690 695 700Ser Asn Tyr Tyr
Lys Ser Thr Asn Val Asp Phe Ala Val Asn Thr Asp705 710 715 720Gly
Thr Tyr Ser Glu Pro Arg Pro Ile Gly Thr Arg Tyr Leu Thr Arg 725 730
735Asn Leu10738PRTAdeno associated virus rh.39 10Met Ala Ala Asp
Gly Tyr Leu Pro Asp Trp Leu Glu Asp Asn Leu Ser1 5 10 15Glu Gly Ile
Arg Glu Trp Trp Ala Leu Lys Pro Gly Ala Pro Lys Pro 20 25 30Lys Ala
Asn Gln Gln Lys Gln Asp Asp Gly Arg Gly Leu Val Leu Pro 35 40 45Gly
Tyr Lys Tyr Leu Gly Pro Phe Asn Gly Leu Asp Lys Gly Glu Pro 50 55
60Val Asn Ala Ala Asp Ala Ala Ala Leu Glu His Asp Lys Ala Tyr Asp65
70 75 80Gln Gln Leu Lys Ala Gly Asp Asn Pro Tyr Leu Arg Tyr Asn His
Ala 85 90 95Asp Ala Glu Phe Gln Glu Arg Leu Gln Glu Asp Thr Ser Phe
Gly Gly 100 105 110Asn Leu Gly Arg Ala Val Phe Gln Ala Lys Lys Arg
Val Leu Glu Pro 115 120 125Leu Gly Leu Val Glu Glu Ala Ala Lys Thr
Ala Pro Gly Lys Lys Arg 130 135 140Pro Val Glu Pro Ser Pro Gln Arg
Ser Pro Asp Ser Ser Thr Gly Ile145 150 155 160Gly Lys Lys Gly Gln
Gln Pro Ala Lys Lys Arg Leu Asn Phe Gly Gln 165 170 175Thr Gly Asp
Ser Glu Ser Val Pro Asp Pro Gln Pro Ile Gly Glu Pro 180 185 190Pro
Ala Gly Pro Ser Gly Leu Gly Ser Gly Thr Met Ala Ala Gly Gly 195 200
205Gly Ala Pro Met Ala Asp Asn Asn Glu Gly Ala Asp Gly Val Gly Ser
210 215 220Ser Ser Gly Asn Trp His Cys Asp Ser Thr Trp Leu Gly Asp
Arg Val225 230 235 240Ile Thr Thr Ser Thr Arg Thr Trp Ala Leu Pro
Thr Tyr Asn Asn His 245 250 255Leu Tyr Lys Gln Ile Ser Asn Gly Thr
Ser Gly Gly Ser Thr Asn Asp 260 265 270Asn Thr Tyr Phe Gly Tyr Ser
Thr Pro Trp Gly Tyr Phe Asp Phe Asn 275 280 285Arg Phe His Cys His
Phe Ser Pro Arg Asp Trp Gln Arg Leu Ile Asn 290 295 300Asn Asn Trp
Gly Phe Arg Pro Lys Arg Leu Ser Phe Lys Leu Phe Asn305 310 315
320Ile Gln Val Lys Glu Val Thr Gln Asn Glu Gly Thr Lys Thr Ile Ala
325 330 335Asn Asn Leu Thr Ser Thr Ile Gln Val Phe Thr Asp Ser Glu
Tyr Gln 340 345 350Leu Pro Tyr Val Leu Gly Ser Ala His Gln Gly Cys
Leu Pro Pro Phe 355 360 365Pro Ala Asp Val Phe Met Ile Pro Gln Tyr
Gly Tyr Leu Thr Leu Asn 370 375 380Asn Gly Ser Gln Ala Val Gly Arg
Ser Ser Phe Tyr Cys Leu Glu Tyr385 390 395 400Phe Pro Ser Gln Met
Leu Arg Thr Gly Asn Asn Phe Glu Phe Ser Tyr 405 410 415Thr Phe Glu
Asp Val Pro Phe His Ser Ser Tyr Ala His Ser Gln Ser 420 425 430Leu
Asp Arg Leu Met Asn Pro Leu Ile Asp Gln Tyr Leu Tyr Tyr Leu 435 440
445Ser Arg Thr Gln Ser Thr Gly Gly Thr Gln Gly Thr Gln Gln Leu Leu
450 455 460Phe Ser Gln Ala Gly Pro Ala Asn Met Ser Ala Gln Ala Lys
Asn Trp465 470 475 480Leu Pro Gly Pro Cys Tyr Arg Gln Gln Arg Val
Ser Thr Thr Leu Ser 485 490 495Gln Asn Asn Asn Ser Asn Phe Ala Trp
Thr Gly Ala Thr Lys Tyr His 500 505 510Leu Asn Gly Arg Asp Ser Leu
Val Asn Pro Gly Val Ala Met Ala Thr 515 520 525His Lys Asp Asp Glu
Glu Arg Phe Phe Pro Ser Ser Gly Val Leu Met 530 535 540Phe Gly Lys
Gln Gly Ala Gly Arg Asp Asn Val Asp Tyr Ser Ser Val545 550 555
560Met Leu Thr Ser Glu Glu Glu Ile Lys Thr Thr Asn Pro Val Ala Thr
565 570 575Glu Gln Tyr Gly Val Val Ala Asp Asn Leu Gln Gln Thr Asn
Thr Gly 580 585 590Pro Ile Val Gly Asn Val Asn Ser Gln Gly Ala Leu
Pro Gly Met Val 595 600 605Trp Gln Asn Arg Asp Val Tyr Leu Gln Gly
Pro Ile Trp Ala Lys Ile 610 615 620Pro His Thr Asp Gly Asn Phe His
Pro Ser Pro Leu Met Gly Gly Phe625 630 635 640Gly Leu Lys His Pro
Pro Pro Gln Ile Leu Ile Lys Asn Thr Pro Val 645 650 655Pro Ala Asp
Pro Pro Thr Thr Phe Ser Gln Ala Lys Leu Ala Ser Phe 660 665 670Ile
Thr Gln Tyr Ser Thr Gly Gln Val Ser Val Glu Ile Glu Trp Glu 675 680
685Leu Gln Lys Glu Asn Ser Lys Arg Trp Asn Pro Glu Ile Gln Tyr Thr
690 695 700Ser Asn Tyr Tyr Lys Ser Thr Asn Val Asp Phe Ala Val Asn
Thr Glu705 710 715 720Gly Thr Tyr Ser Glu Pro Arg Pro Ile Gly Thr
Arg Tyr Leu Thr Arg 725 730 735Asn Leu11736PRTAdeno associated
virus rh.43 11Met Ala Ala Asp Gly Tyr Leu Pro Asp Trp Leu Glu Asp
Asn Leu Ser1 5 10 15Glu Gly Ile Arg Glu Trp Trp Asp Leu Lys Pro
Gly
Ala Pro Lys Pro 20 25 30Lys Ala Asn Gln Gln Lys Gln Asp Asp Gly Arg
Gly Leu Val Leu Pro 35 40 45Gly Tyr Lys Tyr Leu Gly Pro Phe Asn Gly
Leu Asp Lys Gly Glu Pro 50 55 60Val Asn Ala Ala Asp Ala Ala Ala Leu
Glu His Asp Lys Ala Tyr Asp65 70 75 80Gln Gln Leu Glu Ala Gly Asp
Asn Pro Tyr Leu Arg Tyr Asn His Ala 85 90 95Asp Ala Glu Phe Gln Glu
Arg Leu Gln Glu Asp Thr Ser Phe Gly Gly 100 105 110Asn Leu Gly Arg
Ala Val Phe Gln Ala Lys Lys Arg Val Leu Glu Pro 115 120 125Leu Gly
Leu Val Glu Glu Gly Ala Lys Thr Ala Pro Gly Lys Lys Arg 130 135
140Pro Val Glu Gln Ser Pro Gln Glu Pro Asp Ser Ser Ser Gly Ile
Gly145 150 155 160Lys Lys Gly Gln Gln Pro Ala Arg Lys Arg Leu Asn
Phe Gly Gln Thr 165 170 175Gly Asp Ser Glu Ser Val Pro Asp Pro Gln
Pro Leu Gly Glu Pro Pro 180 185 190Ala Ala Pro Ser Gly Val Gly Pro
Asn Thr Met Ala Ala Gly Gly Gly 195 200 205Ala Pro Met Ala Asp Asn
Asn Glu Gly Ala Asp Gly Val Gly Ser Ser 210 215 220Ser Gly Asn Trp
His Cys Asp Ser Thr Trp Leu Gly Asp Arg Val Ile225 230 235 240Thr
Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr Tyr Asn Asn His Leu 245 250
255Tyr Lys Gln Ile Ser Asn Gly Thr Ser Gly Gly Ala Thr Asn Asp Asn
260 265 270Thr Tyr Phe Gly Tyr Ser Thr Pro Trp Gly Tyr Phe Asp Phe
Asn Arg 275 280 285Phe His Cys His Phe Ser Pro Arg Asp Trp Gln Arg
Leu Ile Asn Asn 290 295 300Asn Trp Gly Phe Arg Pro Lys Arg Leu Ser
Phe Lys Leu Phe Asn Ile305 310 315 320Gln Val Lys Glu Val Thr Gln
Asn Glu Gly Thr Lys Thr Ile Ala Asn 325 330 335Asn Leu Thr Ser Thr
Ile Gln Val Phe Thr Asp Ser Glu Tyr Gln Leu 340 345 350Pro Tyr Val
Leu Gly Ser Ala His Gln Gly Cys Leu Pro Pro Phe Pro 355 360 365Ala
Asp Val Phe Met Ile Pro Gln Tyr Gly Tyr Leu Thr Leu Asn Asn 370 375
380Gly Ser Gln Ala Val Gly Arg Ser Ser Phe Tyr Cys Leu Glu Tyr
Phe385 390 395 400Pro Ser Gln Met Leu Arg Thr Gly Asn Asn Phe Gln
Phe Thr Tyr Thr 405 410 415Phe Glu Asp Val Pro Phe His Ser Ser Tyr
Ala His Ser Gln Ser Leu 420 425 430Asp Arg Leu Met Asn Pro Leu Ile
Asp Gln Tyr Leu Tyr Tyr Leu Ser 435 440 445Arg Thr Gln Thr Thr Gly
Gly Thr Ala Asn Thr Gln Thr Leu Gly Phe 450 455 460Ser Gln Gly Gly
Pro Asn Thr Met Ala Asn Gln Ala Lys Asn Trp Leu465 470 475 480Pro
Gly Pro Cys Tyr Arg Gln Gln Arg Val Ser Thr Thr Thr Gly Gln 485 490
495Asn Asn Asn Ser Asn Phe Ala Trp Thr Ala Gly Thr Lys Tyr His Leu
500 505 510Asn Gly Arg Asn Ser Leu Ala Asn Pro Gly Ile Ala Met Ala
Thr His 515 520 525Lys Asp Asp Glu Glu Arg Phe Phe Pro Val Thr Gly
Ser Cys Phe Trp 530 535 540Gln Gln Asn Ala Ala Arg Asp Asn Ala Asp
Tyr Ser Asp Val Met Leu545 550 555 560Thr Ser Glu Glu Glu Ile Lys
Thr Thr Asn Pro Val Ala Thr Glu Glu 565 570 575Tyr Gly Ile Val Ala
Asp Asn Leu Gln Gln Gln Asn Thr Ala Pro Gln 580 585 590Ile Gly Thr
Val Asn Ser Gln Gly Ala Leu Pro Gly Met Val Trp Gln 595 600 605Asn
Arg Asp Val Tyr Leu Gln Gly Pro Ile Trp Ala Lys Ile Pro His 610 615
620Thr Asp Gly Asn Phe His Pro Ser Pro Leu Met Gly Gly Phe Gly
Leu625 630 635 640Lys His Pro Pro Pro Gln Ile Leu Ile Lys Asn Thr
Pro Val Pro Ala 645 650 655Asp Pro Pro Thr Thr Phe Asn Gln Ser Lys
Leu Asn Ser Phe Ile Thr 660 665 670Gln Tyr Ser Thr Gly Gln Val Ser
Val Glu Ile Glu Trp Glu Leu Gln 675 680 685Lys Glu Asn Ser Lys Arg
Trp Asn Pro Glu Ile Gln Tyr Thr Ser Asn 690 695 700Tyr Tyr Lys Ser
Thr Ser Val Asp Phe Ala Val Asn Thr Glu Gly Val705 710 715 720Tyr
Ser Glu Pro Arg Pro Ile Gly Thr Arg Tyr Leu Thr Arg Asn Leu 725 730
73512736PRTAdeno associated virus CSp3 12Met Ala Ala Asp Gly Tyr
Leu Pro Asp Trp Leu Glu Asp Asn Leu Ser1 5 10 15Glu Gly Ile Arg Glu
Trp Trp Ala Leu Lys Pro Gly Ala Pro Gln Pro 20 25 30Lys Ala Asn Gln
Gln His Gln Asp Asn Ala Arg Gly Leu Val Leu Pro 35 40 45Gly Tyr Lys
Tyr Leu Gly Pro Gly Asn Gly Leu Asp Lys Gly Glu Pro 50 55 60Val Asn
Ala Ala Asp Ala Ala Ala Leu Glu His Asp Lys Ala Tyr Asp65 70 75
80Gln Gln Leu Lys Ala Gly Asp Asn Pro Tyr Leu Lys Tyr Asn His Ala
85 90 95Asp Ala Glu Phe Gln Glu Arg Leu Lys Glu Asp Thr Ser Phe Gly
Gly 100 105 110Asn Leu Gly Arg Ala Val Phe Gln Ala Lys Lys Arg Leu
Leu Glu Pro 115 120 125Leu Gly Leu Val Glu Glu Ala Ala Lys Thr Ala
Pro Gly Lys Lys Arg 130 135 140Pro Val Glu Gln Ser Pro Gln Glu Pro
Asp Ser Ser Ala Gly Ile Gly145 150 155 160Lys Ser Gly Ala Gln Pro
Ala Lys Lys Arg Leu Asn Phe Gly Gln Thr 165 170 175Gly Asp Thr Glu
Ser Val Pro Asp Pro Gln Pro Ile Gly Glu Pro Pro 180 185 190Ala Ala
Pro Ser Gly Val Gly Ser Leu Thr Ile Ala Ser Gly Gly Gly 195 200
205Ala Pro Val Ala Asp Asn Asn Glu Gly Ala Asp Gly Val Gly Ser Ser
210 215 220Ser Gly Asn Trp His Cys Asp Ser Gln Trp Leu Gly Asp Arg
Val Ile225 230 235 240Thr Thr Ser Thr Arg Thr Trp Ala Leu Pro Thr
Tyr Asn Asn His Leu 245 250 255Tyr Lys Arg Ile Ser Asn Ser Thr Ser
Gly Gly Ser Ser Asn Asp Asn 260 265 270Ala Tyr Phe Gly Tyr Ser Thr
Pro Trp Gly Tyr Phe Asp Phe Asn Arg 275 280 285Phe His Cys His Phe
Ser Pro Arg Asp Trp Gln Arg Leu Ile Asn Asn 290 295 300Asn Trp Gly
Phe Arg Pro Lys Arg Leu Asn Phe Lys Leu Phe Asn Ile305 310 315
320Arg Val Lys Glu Val Thr Asp Asn Asn Gly Val Lys Thr Ile Thr Asn
325 330 335Asn Leu Thr Ser Thr Val Gln Val Phe Thr Asp Ser Asp Tyr
Gln Leu 340 345 350Pro Tyr Val Leu Gly Ser Ala His Glu Gly Cys Leu
Pro Pro Phe Pro 355 360 365Ala Asp Val Phe Met Ile Pro Gln Tyr Gly
Tyr Leu Thr Leu Asn Asp 370 375 380Gly Ser Gln Ala Val Gly Arg Ser
Ser Phe Tyr Cys Leu Glu Tyr Phe385 390 395 400Pro Ser Gln Met Leu
Arg Thr Gly Asn Asn Phe Gln Phe Ser Tyr Glu 405 410 415Phe Glu Asn
Val Pro Phe His Ser Ser Tyr Ala His Ser Gln Ser Leu 420 425 430Asp
Arg Leu Met Asn Pro Leu Ile Asp Gln Tyr Leu Tyr Tyr Leu Ser 435 440
445Lys Thr Ile Asn Gly Ser Gly Gln Asn Gln Gln Thr Leu Lys Phe Ser
450 455 460Val Ala Gly Pro Ser Asn Met Ala Val Gln Gly Arg Asn Tyr
Ile Pro465 470 475 480Gly Pro Ser Tyr Arg Gln Gln Arg Val Ser Thr
Thr Val Thr Arg Asn 485 490 495Asn Asn Ser Glu Phe Ala Trp Pro Gly
Ala Ser Ser Trp Ala Leu Asn 500 505 510Gly Arg Asn Ser Leu Met Asn
Pro Gly Pro Ala Met Ala Ser His Lys 515 520 525Glu Gly Glu Asp Arg
Phe Phe Pro Leu Ser Gly Ser Leu Ile Phe Gly 530 535 540Lys Gln Gly
Thr Gly Arg Asp Asn Val Asp Ala Asp Lys Val Met Ile545 550 555
560Thr Asn Glu Glu Glu Ile Lys Thr Thr Asn Pro Val Ala Thr Glu Ser
565 570 575Tyr Gly Gln Val Ala Thr Asn His Gln Ser Ala Gln Ala Gln
Ala Gln 580 585 590Thr Gly Trp Val Gln Asn Gln Gly Ile Leu Pro Gly
Met Val Trp Gln 595 600 605Asp Arg Asp Val Tyr Leu Gln Gly Pro Ile
Trp Ala Lys Ile Pro His 610 615 620Thr Asp Gly Asn Phe His Pro Ser
Pro Leu Met Gly Gly Phe Gly Val625 630 635 640Lys His Pro Pro Pro
Gln Ile Leu Ile Lys Asn Thr Pro Val Pro Ala 645 650 655Asp Pro Pro
Thr Ala Phe Asn Lys Asp Lys Leu Asn Ser Phe Ile Thr 660 665 670Gln
Tyr Ser Thr Gly Gln Val Ser Val Glu Ile Glu Trp Glu Leu Gln 675 680
685Lys Glu Asn Ser Lys Arg Trp Asn Pro Glu Ile Gln Tyr Thr Ser Asn
690 695 700Tyr Tyr Lys Ser Asn Asn Val Glu Phe Ala Val Asn Thr Glu
Gly Val705 710 715 720Tyr Ser Glu Pro Arg Pro Ile Gly Thr Arg Tyr
Leu Thr Arg Asn Leu 725 730 73513313PRTHomo sapiens 13Met Thr Ser
Cys His Ile Ala Glu Glu His Ile Gln Lys Val Ala Ile1 5 10 15Phe Gly
Gly Thr His Gly Asn Glu Leu Thr Gly Val Phe Leu Val Lys 20 25 30His
Trp Leu Glu Asn Gly Ala Glu Ile Gln Arg Thr Gly Leu Glu Val 35 40
45Lys Pro Phe Ile Thr Asn Pro Arg Ala Val Lys Lys Cys Thr Arg Tyr
50 55 60Ile Asp Cys Asp Leu Asn Arg Ile Phe Asp Leu Glu Asn Leu Gly
Lys65 70 75 80Lys Met Ser Glu Asp Leu Pro Tyr Glu Val Arg Arg Ala
Gln Glu Ile 85 90 95Asn His Leu Phe Gly Pro Lys Asp Ser Glu Asp Ser
Tyr Asp Ile Ile 100 105 110Phe Asp Leu His Asn Thr Thr Ser Asn Met
Gly Cys Thr Leu Ile Leu 115 120 125Glu Asp Ser Arg Asn Asn Phe Leu
Ile Gln Met Phe His Tyr Ile Lys 130 135 140Thr Ser Leu Ala Pro Leu
Pro Cys Tyr Val Tyr Leu Ile Glu His Pro145 150 155 160Ser Leu Lys
Tyr Ala Thr Thr Arg Ser Ile Ala Lys Tyr Pro Val Gly 165 170 175Ile
Glu Val Gly Pro Gln Pro Gln Gly Val Leu Arg Ala Asp Ile Leu 180 185
190Asp Gln Met Arg Lys Met Ile Lys His Ala Leu Asp Phe Ile His His
195 200 205Phe Asn Glu Gly Lys Glu Phe Pro Pro Cys Ala Ile Glu Val
Tyr Lys 210 215 220Ile Ile Glu Lys Val Asp Tyr Pro Arg Asp Glu Asn
Gly Glu Ile Ala225 230 235 240Ala Ile Ile His Pro Asn Leu Gln Asp
Gln Asp Trp Lys Pro Leu His 245 250 255Pro Gly Asp Pro Met Phe Leu
Thr Leu Asp Gly Lys Thr Ile Pro Leu 260 265 270Gly Gly Asp Cys Thr
Val Tyr Pro Val Phe Val Asn Glu Ala Ala Tyr 275 280 285Tyr Glu Lys
Lys Glu Ala Phe Ala Lys Thr Thr Lys Leu Thr Leu Asn 290 295 300Ala
Lys Ser Ile Arg Cys Cys Leu His305 31014942DNAHomo sapiens
14atgacttctt gtcacattgc tgaagaacat atacaaaagg ttgctatctt tggaggaacc
60catgggaatg agctaaccgg agtatttctg gttaagcatt ggctagagaa tggcgctgag
120attcagagaa cagggctgga ggtaaaacca tttattacta accccagagc
agtgaagaag 180tgtaccagat atattgactg tgacctgaat cgcatttttg
accttgaaaa tcttggcaaa 240aaaatgtcag aagatttgcc atatgaagtg
agaagggctc aagaaataaa tcatttattt 300ggtccaaaag acagtgaaga
ttcctatgac attatttttg accttcacaa caccacctct 360aacatggggt
gcactcttat tcttgaggat tccaggaata actttttaat tcagatgttt
420cattacatta agacttctct ggctccacta ccctgctacg tttatctgat
tgagcatcct 480tccctcaaat atgcgaccac tcgttccata gccaagtatc
ctgtgggtat agaagttggt 540cctcagcctc aaggggttct gagagctgat
atcttggatc aaatgagaaa aatgattaaa 600catgctcttg attttataca
tcatttcaat gaaggaaaag aatttcctcc ctgcgccatt 660gaggtctata
aaattataga gaaagttgat tacccccggg atgaaaatgg agaaattgct
720gctatcatcc atcctaatct gcaggatcaa gactggaaac cactgcatcc
tggggatccc 780atgtttttaa ctcttgatgg gaagacgatc ccactgggcg
gagactgtac cgtgtacccc 840gtgtttgtga atgaggccgc atattacgaa
aagaaagaag cttttgcaaa gacaactaaa 900ctaacgctca atgcaaaaag
tattcgctgc tgtttacatt ag 94215312PRTMus Musculus 15Met Thr Ser Cys
Val Ala Lys Glu Pro Ile Lys Lys Ile Ala Ile Phe1 5 10 15Gly Gly Thr
His Gly Asn Glu Leu Thr Gly Val Phe Leu Val Thr His 20 25 30Trp Leu
Arg Asn Gly Thr Glu Val His Arg Ala Gly Leu Asp Val Lys 35 40 45Pro
Phe Ile Thr Asn Pro Arg Ala Val Glu Lys Cys Thr Arg Tyr Ile 50 55
60Asp Cys Asp Leu Asn Arg Val Phe Asp Leu Glu Asn Leu Ser Lys Glu65
70 75 80Met Ser Glu Asp Leu Pro Tyr Glu Val Arg Arg Ala Gln Glu Ile
Asn 85 90 95His Leu Phe Gly Pro Lys Asn Ser Asp Asp Ala Tyr Asp Leu
Val Phe 100 105 110Asp Leu His Asn Thr Thr Ser Asn Met Gly Cys Thr
Leu Ile Leu Glu 115 120 125Asp Ser Arg Asn Asp Phe Leu Ile Gln Met
Phe His Tyr Ile Lys Thr 130 135 140Cys Met Ala Pro Leu Pro Cys Ser
Val Tyr Leu Ile Glu His Pro Ser145 150 155 160Leu Lys Tyr Ala Thr
Thr Arg Ser Ile Ala Lys Tyr Pro Val Gly Ile 165 170 175Glu Val Gly
Pro Gln Pro His Gly Val Leu Arg Ala Asp Ile Leu Asp 180 185 190Gln
Met Arg Lys Met Ile Lys His Ala Leu Asp Phe Ile Gln His Phe 195 200
205Asn Glu Gly Lys Glu Phe Pro Pro Cys Ser Ile Asp Val Tyr Lys Ile
210 215 220Met Glu Lys Val Asp Tyr Pro Arg Asn Glu Ser Gly Asp Met
Ala Ala225 230 235 240Val Ile His Pro Asn Leu Gln Asp Gln Asp Trp
Lys Pro Leu His Pro 245 250 255Gly Asp Pro Val Phe Val Ser Leu Asp
Gly Lys Val Ile Pro Leu Gly 260 265 270Gly Asp Cys Thr Val Tyr Pro
Val Phe Val Asn Glu Ala Ala Tyr Tyr 275 280 285Glu Lys Lys Glu Ala
Phe Ala Lys Thr Thr Lys Leu Thr Leu Ser Ala 290 295 300Lys Ser Ile
Arg Ser Thr Leu His305 31016939DNAMus Musculus 16atgacctctt
gtgttgctaa agaacctatt aagaagattg ccatctttgg agggactcat 60ggaaatgaac
tgaccggagt gtttctagtt actcactggc taaggaatgg cactgaagtt
120cacagagcag ggctggacgt gaagccattc attaccaatc caagggcggt
ggagaagtgc 180accagataca ttgactgtga cctgaatcgt gtttttgacc
ttgaaaatct tagcaaagag 240atgtctgaag acttgccata tgaagtgaga
agggctcaag aaataaatca tttatttggt 300ccaaaaaata gtgatgatgc
ctatgacctt gtttttgacc ttcacaacac cacttctaac 360atgggttgca
ctcttattct tgaggattcc aggaatgact ttttaattca gatgtttcac
420tatattaaga cttgcatggc tccattaccc tgctctgttt atctcattga
gcatccttca 480ctcaaatatg caaccactcg ttccattgcc aagtatcctg
ttggtataga agttggtcct 540cagcctcacg gtgtccttag agctgatatt
ttagaccaaa tgagaaaaat gataaaacat 600gctcttgatt ttatacagca
tttcaatgaa ggaaaagaat ttcctccctg ttctattgac 660gtctataaaa
taatggagaa agttgattat ccaaggaatg aaagtggaga catggctgct
720gttattcatc ctaatctgca ggatcaagac tggaaaccat tgcaccctgg
agatcctgtg 780tttgtgtctc ttgatggaaa agttattcca ctgggtggag
actgtaccgt gtacccagtg 840tttgtgaatg aagctgcata ttatgaaaaa
aaagaagcat ttgcaaagac aacaaaacta 900acactcagcg caaaaagcat
ccgctccact ttgcactaa 93917465DNAHomo sapiens 17atggcgacga
aggccgtgtg cgtgctgaag ggcgacggcc cagtgcaggg catcatcaat 60ttcgagcaga
aggaaagtaa tggaccagtg aaggtgtggg gaagcattaa aggactgact
120gaaggcctgc atggattcca tgttcatgag tttggagata atacagcagg
ctgtaccagt 180gcaggtcctc actttaatcc tctatccaga aaacacggtg
ggccaaagga tgaagagagg 240catgttggag acttgggcaa tgtgactgct
gacaaagatg gtgtggccga tgtgtctatt 300gaagattctg tgatctcact
ctcaggagac cattgcatca ttggccgcac actggtggtc 360catgaaaaag
cagatgactt gggcaaaggt ggaaatgaag aaagtacaaa gacaggaaac
420gctggaagtc gtttggcttg tggtgtaatt gggatcgccc aataa
46518154PRTHomo sapiens 18Met Ala Thr Lys Ala Val Cys Val Leu Lys
Gly Asp Gly Pro Val Gln1 5 10 15Gly Ile Ile Asn Phe Glu Gln Lys Glu
Ser Asn Gly Pro Val Lys Val 20 25 30Trp Gly Ser Ile Lys Gly Leu Thr
Glu Gly Leu His Gly Phe His Val 35 40 45His Glu Phe Gly Asp Asn Thr
Ala Gly Cys Thr Ser Ala Gly Pro His 50 55 60Phe Asn Pro Leu Ser Arg
Lys His Gly Gly Pro Lys Asp Glu Glu Arg65 70 75 80His Val Gly Asp
Leu Gly Asn Val Thr Ala Asp Lys Asp Gly Val Ala 85 90 95Asp Val Ser
Ile Glu Asp Ser Val Ile Ser Leu Ser Gly Asp His Cys 100 105 110Ile
Ile Gly Arg Thr Leu Val Val His Glu Lys Ala Asp Asp Leu Gly 115 120
125Lys Gly Gly Asn Glu Glu Ser Thr Lys Thr Gly Asn Ala Gly Ser Arg
130 135 140Leu Ala Cys Gly Val Ile Gly Ile Ala Gln145
15019465DNAMus Musculus 19atggcgatga aagcggtgtg cgtgctgaag
ggcgacggtc cggtgcaggg aaccatccac 60ttcgagcaga aggcaagcgg tgaaccagtt
gtgttgtcag gacaaattac aggattaact 120gaaggccagc atgggttcca
cgtccatcag tatggggaca atacacaagg ctgtaccagt 180gcaggacctc
attttaatcc tcactctaag aaacatggtg gcccggcgga tgaagagagg
240catgttggag acctgggcaa tgtgactgct ggaaaggacg gtgtggccaa
tgtgtccatt 300gaagatcgtg tgatctcact ctcaggagag cattccatca
ttggccgtac aatggtggtc 360catgagaaac aagatgactt gggcaaaggt
ggaaatgaag aaagtacaaa gactggaaat 420gctgggagcc gcttggcctg
tggagtgatt gggattgcgc agtaa 46520154PRTMus Musculus 20Met Ala Met
Lys Ala Val Cys Val Leu Lys Gly Asp Gly Pro Val Gln1 5 10 15Gly Thr
Ile His Phe Glu Gln Lys Ala Ser Gly Glu Pro Val Val Leu 20 25 30Ser
Gly Gln Ile Thr Gly Leu Thr Glu Gly Gln His Gly Phe His Val 35 40
45His Gln Tyr Gly Asp Asn Thr Gln Gly Cys Thr Ser Ala Gly Pro His
50 55 60Phe Asn Pro His Ser Lys Lys His Gly Gly Pro Ala Asp Glu Glu
Arg65 70 75 80His Val Gly Asp Leu Gly Asn Val Thr Ala Gly Lys Asp
Gly Val Ala 85 90 95Asn Val Ser Ile Glu Asp Arg Val Ile Ser Leu Ser
Gly Glu His Ser 100 105 110Ile Ile Gly Arg Thr Met Val Val His Glu
Lys Gln Asp Asp Leu Gly 115 120 125Lys Gly Gly Asn Glu Glu Ser Thr
Lys Thr Gly Asn Ala Gly Ser Arg 130 135 140Leu Ala Cys Gly Val Ile
Gly Ile Ala Gln145 150215243DNAArtificial SequenceSynthetic
Oligonucleotide 21ctgcgcgctc gctcgctcac tgaggccgcc cgggcaaagc
ccgggcgtcg ggcgaccttt 60ggtcgcccgg cctcagtgag cgagcgagcg cgcagagagg
gagtgtagcc atgctctagg 120aagatcaatt caattcacgc gtcgacattg
attattgact agttattaat agtaatcaat 180tacggggtca ttagttcata
gcccatatat ggagttccgc gttacataac ttacggtaaa 240tggcccgcct
ggctgaccgc ccaacgaccc ccgcccattg acgtcaataa tgacgtatgt
300tcccatagta acgccaatag ggactttcca ttgacgtcaa tgggtggata
tttacggtaa 360actgcccact tggcagtaca tcaagtgtat catatgccaa
gtacgccccc tattgacgtc 420aatgacggta aatggcccgc ctggcattat
gcccagtaca tgaccttatg ggactttcct 480acttggcagt acatctacgt
attagtcatc gctattacca tgtcgaggcc acgttctgct 540tcactctccc
catctccccc ccctccccac ccccaatttt gtatttattt attttttaat
600tattttgtgc agcgatgggg gcgggggggg ggggcgcgcg ccaggcgggg
cggggcgggg 660cgaggggcgg ggcggggcga ggcggagagg tgcggcggca
gccaatcaga gcggcgcgct 720ccgaaagttt ccttttatgg cgaggcggcg
gcggcggcgg ccctataaaa agcgaagcgc 780gcggcgggcg ggagcaagct
ctagcctcga gaattcacgc gtggtacctc tagagcagag 840ctcgtttagt
gaaccgtcag ttcgaaatcg ccaccatggt gagcaagggc gaggagctgt
900tcaccggggt ggtgcccatc ctggtcgagc tggacggcga cgtaaacggc
cacaagttca 960gcgtgtccgg cgagggcgag ggcgatgcca cctacggcaa
gctgaccctg aagttcatct 1020gcaccaccgg caagctgccc gtgccctggc
ccaccctcgt gaccaccctg acctacggcg 1080tgcagtgctt cagccgctac
cccgaccaca tgaagcagca cgacttcttc aagtccgcca 1140tgcccgaagg
ctacgtccag gagcgcacca tcttcttcaa ggacgacggc aactacaaga
1200cccgcgccga ggtgaagttc gagggcgaca ccctggtgaa ccgcatcgag
ctgaagggca 1260tcgacttcaa ggaggacggc aacatcctgg ggcacaagct
ggagtacaac tacaacagcc 1320acaacgtcta tatcatggcc gacaagcaga
agaacggcat caaggtgaac ttcaagatcc 1380gccacaacat cgaggacggc
agcgtgcagc tcgccgacca ctaccagcag aacaccccca 1440tcggcgacgg
ccccgtgctg ctgcccgaca accactacct gagcacccag tccgccctga
1500gcaaagaccc caacgagaag cgcgatcaca tggtcctgct ggagttcgtg
accgccgccg 1560ggatcactct cggcatggac gagctgtaca agtaagtaac
aggtaagtgc gatcgctaat 1620gcgggaaagc tcttattcgg gtgagatggg
ctggggcacc atctggggac cctgacgtga 1680agtttgtcac tgactggaga
actcggtttg tcgtctgttg cgggggcggc agttatggcg 1740gtgccgttgg
gcagtgcacc cgtacctttg ggagcgcgcg ccctcgtcgt gtcgtgacgt
1800cacccgttct gttggtacct gctgttgaca gtgagcgacg caatgtgact
tcgctgacaa 1860agctgtgaag ccacagatgg gctttgtcag cagtcacatt
gcgctgccta ctgcctcgga 1920cttcaagggc tcgagaattc agggtggggc
cacctgccgg taggtgtgcg gtaggctttt 1980ctccgtcgca ggacgcaggg
ttcgggccta gggtaggctc tcctgaatcg acaggcgccg 2040gacctctggc
ggccgcaaca acgcgttcct gaccattcat cctctttctt tttcctgcag
2100gcttgtggaa gaaatgggat ccgatctttt tccctctgcc aaaaattatg
gggacatcat 2160gaagcccctt gagcatctga cttctggcta ataaaggaaa
tttattttca ttgcaatagt 2220gtgttggaat tttttgtgtc tctcactcgg
cctaggtaga taagtagcat ggcgggttaa 2280tcattaacta caaggaaccc
ctagtgatgg agttggccac tccctctctg cgcgctcgct 2340cgctcactga
ggccgggcga ccaaaggtcg cccgacgccc gggctttgcc cgggcggcct
2400cagtgagcga gcgagcgcgc agccttaatt aacctaattc actggccgtc
gttttacaac 2460gtcgtgactg ggaaaaccct ggcgttaccc aacttaatcg
ccttgcagca catccccctt 2520tcgccagctg gcgtaatagc gaagaggccc
gcaccgatcg cccttcccaa cagttgcgca 2580gcctgaatgg cgaatgggac
gcgccctgta gcggcgcatt aagcgcggcg ggtgtggtgg 2640ttacgcgcag
cgtgaccgct acacttgcca gcgccctagc gcccgctcct ttcgctttct
2700tcccttcctt tctcgccacg ttcgccggct ttccccgtca agctctaaat
cgggggctcc 2760ctttagggtt ccgatttagt gctttacggc acctcgaccc
caaaaaactt gattagggtg 2820atggttcacg tagtgggcca tcgccctgat
agacggtttt tcgccctttg acgttggagt 2880ccacgttctt taatagtgga
ctcttgttcc aaactggaac aacactcaac cctatctcgg 2940tctattcttt
tgatttataa gggattttgc cgatttcggc ctattggtta aaaaatgagc
3000tgatttaaca aaaatttaac gcgaatttta acaaaatatt aacgcttaca
atttaggtgg 3060cacttttcgg ggaaatgtgc gcggaacccc tatttgttta
tttttctaaa tacattcaaa 3120tatgtatccg ctcatgagac aataaccctg
ataaatgctt caataatatt gaaaaaggaa 3180gagtatgagt attcaacatt
tccgtgtcgc ccttattccc ttttttgcgg cattttgcct 3240tcctgttttt
gctcacccag aaacgctggt gaaagtaaaa gatgctgaag atcagttggg
3300tgcacgagtg ggttacatcg aactggatct caacagcggt aagatccttg
agagttttcg 3360ccccgaagaa cgttttccaa tgatgagcac ttttaaagtt
ctgctatgtg gcgcggtatt 3420atcccgtatt gacgccgggc aagagcaact
cggtcgccgc atacactatt ctcagaatga 3480cttggttgag tactcaccag
tcacagaaaa gcatcttacg gatggcatga cagtaagaga 3540attatgcagt
gctgccataa ccatgagtga taacactgcg gccaacttac ttctgacaac
3600gatcggagga ccgaaggagc taaccgcttt tttgcacaac atgggggatc
atgtaactcg 3660ccttgatcgt tgggaaccgg agctgaatga agccatacca
aacgacgagc gtgacaccac 3720gatgcctgta gcaatggcaa caacgttgcg
caaactatta actggcgaac tacttactct 3780agcttcccgg caacaattaa
tagactggat ggaggcggat aaagttgcag gaccacttct 3840gcgctcggcc
cttccggctg gctggtttat tgctgataaa tctggagccg gtgagcgtgg
3900gtctcgcggt atcattgcag cactggggcc agatggtaag ccctcccgta
tcgtagttat 3960ctacacgacg gggagtcagg caactatgga tgaacgaaat
agacagatcg ctgagatagg 4020tgcctcactg attaagcatt ggtaactgtc
agaccaagtt tactcatata tactttagat 4080tgatttaaaa cttcattttt
aatttaaaag gatctaggtg aagatccttt ttgataatct 4140catgaccaaa
atcccttaac gtgagttttc gttccactga gcgtcagacc ccgtagaaaa
4200gatcaaagga tcttcttgag atcctttttt tctgcgcgta atctgctgct
tgcaaacaaa 4260aaaaccaccg ctaccagcgg tggtttgttt gccggatcaa
gagctaccaa ctctttttcc 4320gaaggtaact ggcttcagca gagcgcagat
accaaatact gttcttctag tgtagccgta 4380gttaggccac cacttcaaga
actctgtagc accgcctaca tacctcgctc tgctaatcct 4440gttaccagtg
gctgctgcca gtggcgataa gtcgtgtctt accgggttgg actcaagacg
4500atagttaccg gataaggcgc agcggtcggg ctgaacgggg ggttcgtgca
cacagcccag 4560cttggagcga acgacctaca ccgaactgag atacctacag
cgtgagctat gagaaagcgc 4620cacgcttccc gaagggagaa aggcggacag
gtatccggta agcggcaggg tcggaacagg 4680agagcgcacg agggagcttc
cagggggaaa cgcctggtat ctttatagtc ctgtcgggtt 4740tcgccacctc
tgacttgagc gtcgattttt gtgatgctcg tcaggggggc ggagcctatg
4800gaaaaacgcc agcaacgcgg cctttttacg gttcctggcc ttttgctggc
cttttgctca 4860catgttcttt cctgcgttat cccctgattc tgtggataac
cgtattaccg cctttgagtg 4920agctgatacc gctcgccgca gccgaacgac
cgagcgcagc gagtcagtga gcgaggaagc 4980ggaagagcgc ccaatacgca
aaccgcctct ccccgcgcgt tggccgattc attaatgcag 5040ctggcacgac
aggtttcccg actggaaagc gggcagtgag cgcaacgcaa ttaatgtgag
5100ttagctcact cattaggcac cccaggcttt acactttatg cttccggctc
gtatgttgtg 5160tggaattgtg agcggataac aatttcacac aggaaacagc
tatgaccatg attacgccag 5220atttaattaa ggccttaatt agg
524322108DNAArtificial SequenceSynthetic Oligonucleotide
22tgctgttgac agtgagcgac atcatcaatt ttccgagcag aactgtgaag ccacagatgg
60gttctgctcg aaattgatga tgctgcctac tgcctcggac ttcaaggg
10823106DNAArtificial SequenceSynthetic Oligonucleotide
23tgctgttgac agtgagcgac gcattaaagg atcctgactg actgtgaagc cacagatggg
60tcagtcagtc ctttaatgcg ctgcctactg cctcggactt caaggg
10624108DNAArtificial SequenceSynthetic Oligonucleotide
24tgctgttgac agtgagcgac tgcatggatt ctccatgttc atctgtgaag ccacagatgg
60gatgaacatg gaatccatgc agctgcctac tgcctcggac ttcaaggg
10825106DNAArtificial SequenceSynthetic Oligonucleotide
25tgctgttgac agtgagcgac aaggatgaag atcgaggcat gctgtgaagc cacagatggg
60catgcctctc ttcatccttg ctgcctactg cctcggactt caaggg
10626110DNAArtificial SequenceSynthetic Oligonucleotide
26tgctgttgac agtgagcgac gcaatgtgac ttcgctgaca aagctgtgaa gccacagatg
60ggctttgtca gcagtcacat tgcgctgcct actgcctcgg acttcaaggg
11027108DNAArtificial SequenceSynthetic Oligonucleotide
27tgctgttgac agtgagcgac cgatgtgtct atcttgaaga ttctgtgaag ccacagatgg
60gaatcttcaa tagacacatc ggctgcctac tgcctcggac ttcaaggg
10828106DNAArtificial SequenceSynthetic Oligonucleotide
28tgctgttgac agtgagcgac ggtggaaatg atcagaaagt actgtgaagc cacagatggg
60tactttcttc atttccaccg ctgcctactg cctcggactt caaggg
10629110DNAArtificial SequenceSynthetic Oligonucleotide
29tgctgttgac agtgagcgac gctgtagaaa ttcgtatcct gatctgtgaa gccacagatg
60ggatcaggat acatttctac agcgctgcct actgcctcgg acttcaaggg
11030106DNAArtificial SequenceSynthetic Oligonucleotide
30tgctgttgac agtgagcgag gtattaaact tgtcagaatt tagtgaagcc acagatgtaa
60attctgacaa gtttaatacc ctgcctactg cctcggactt caaggg
10631106DNAArtificial SequenceSynthetic Oligonucleotide
31tgctgttgac agtgagcgac gatgctctaa tcggttctat caagtgaagc cacagatgtt
60gatagaacct tagagcatcg ctgcctactg cctcggactt caaggg
1063228DNAArtificial SequenceSynthetic Oligonucleotide 32cgaaatacat
acttctttac attccatt 283328DNAArtificial SequenceSynthetic
Oligonucleotide 33cgaatggaat gtaaagaagt atgtattt
283429DNAArtificial SequenceSynthetic Oligonucleotide 34cgaaacaaac
accattgtca cactccatt 293529DNAArtificial SequenceSynthetic
Oligonucleotide 35cgaatggagt gtgacaatgg tgtttgttt
293672DNAArtificial SequenceSynthetic Oligonucleotide 36cgaaatacat
acttctttac attccaatac atacttcttt acattccaat acatacttct 60ttacattcca
tt 723772DNAArtificial SequenceSynthetic Oligonucleotide
37cgaatggaat gtaaagaagt atgtattgga atgtaaagaa gtatgtattg gaatgtaaag
60aagtatgtat tt 723875DNAArtificial SequenceSynthetic
Oligonucleotide 38cgaaacaaac accattgtca cactccaaca aacaccattg
tcacactcca acaaacacca 60ttgtcacact ccatt 753975DNAArtificial
SequenceSynthetic Oligonucleotide 39cgaatggagt gtgacaatgg
tgtttgttgg agtgtgacaa tggtgtttgt tggagtgtga 60caatggtgtt tgttt
754051DNAArtificial SequenceSynthetic Oligonucleotide 40cgaaatacat
acttctttac attccaacaa acaccattgt cacactccat t 514151DNAArtificial
SequenceSynthetic Oligonucleotide 41cgaatggagt gtgacaatgg
tgtttgttgg aatgtaaaga agtatgtatt t 5142173DNAArtificial
SequenceSynthetic Oligonucleotide 42ttcgaactcg agatacatac
ttctttacat tccaatacat acttctttac attccaatac 60atacttcttt acattccacc
atggactagt acaaacacca ttgtcacact ccaacaaaca 120ccattgtcac
actccaacaa acaccattgt cacactccag cggccgcttc gaa
1734329DNAArtificial SequenceSynthetic Oligonucleotide 43atcgggcccg
actgcagttt cagcgtttg 294429DNAArtificial SequenceSynthetic
Oligonucleotide 44cgcgggcccg actttacatt acacacaat
294529DNAArtificial SequenceSynthetic Oligonucleotide 45cgcgggcccg
actgatgtgt gagagagac 294629DNAArtificial SequenceSynthetic
Oligonucleotide 46cgcgggcccg actttcggcc tcccgaggc
294720DNAArtificial SequenceSynthetic Oligonucleotide 47tgaagctgaa
gcctgtgatg 204820DNAArtificial SequenceSynthetic Oligonucleotide
48gagcacctga cagcattgaa 204920DNAArtificial SequenceSynthetic
Oligonucleotide 49ctcagcaaca gctcatggaa 205020DNAArtificial
SequenceSynthetic Oligonucleotide 50ttacttctgg caccacacca
205120DNAArtificial SequenceSynthetic Oligonucleotide 51tggtgtggtg
ccagaagtaa 205220DNAArtificial SequenceSynthetic Oligonucleotide
52caacagatgg ctggcaacta 205318DNAArtificial SequenceSynthetic
Oligonucleotide 53tgggagtggc accttcca 185420DNAArtificial
SequenceSynthetic Oligonucleotide 54cgaccactac cagcagaaca
205520DNAArtificial SequenceSynthetic Oligonucleotide 55cttgtacagc
tcgtccatgc 205620DNAArtificial SequenceSynthetic Oligonucleotide
56aatggcctca gaatgactgc 205720DNAArtificial SequenceSynthetic
Oligonucleotide 57agtcgctttc acagccaaat 205822DNAArtificial
SequenceSynthetic Oligonucleotide 58atgccaacac agtgctgtct gg
225922DNAArtificial SequenceSynthetic Oligonucleotide 59tgcttgctga
tccacatctg ct 226022DNAArtificial SequenceSynthetic Oligonucleotide
60tggagtgtga caatggtgtt tg 226122DNAArtificial SequenceSynthetic
Oligonucleotide 61aactatacaa cctactacct ca 226222DNAArtificial
SequenceSynthetic Oligonucleotide 62agcctatcct ggattacttg aa
226322DNAArtificial SequenceSynthetic Oligonucleotide 63acagttcttc
aactggcagc tt 226426DNAArtificial SequenceSynthetic Oligonucleotide
64ctctgtatcg ttccaatttt agtata 26
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014
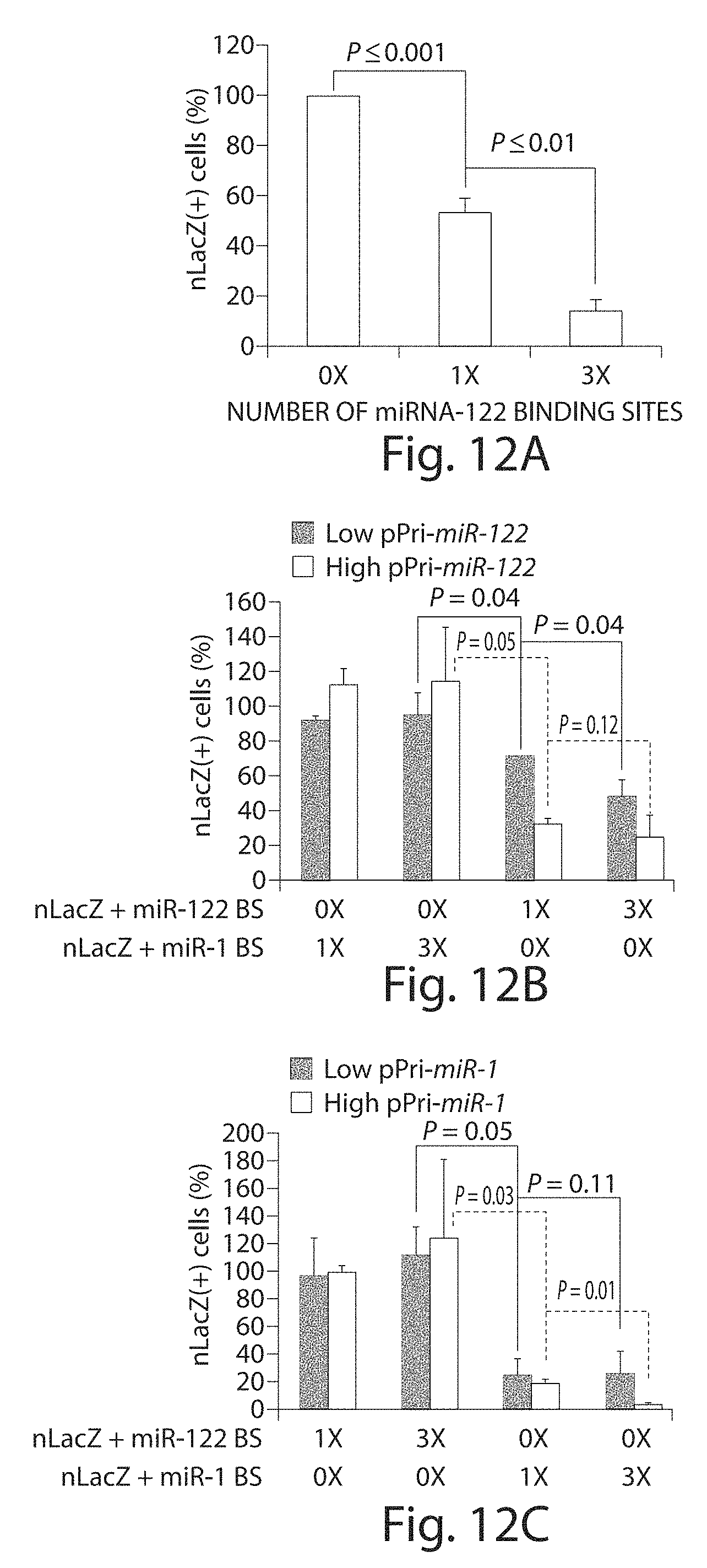
D00015

D00016
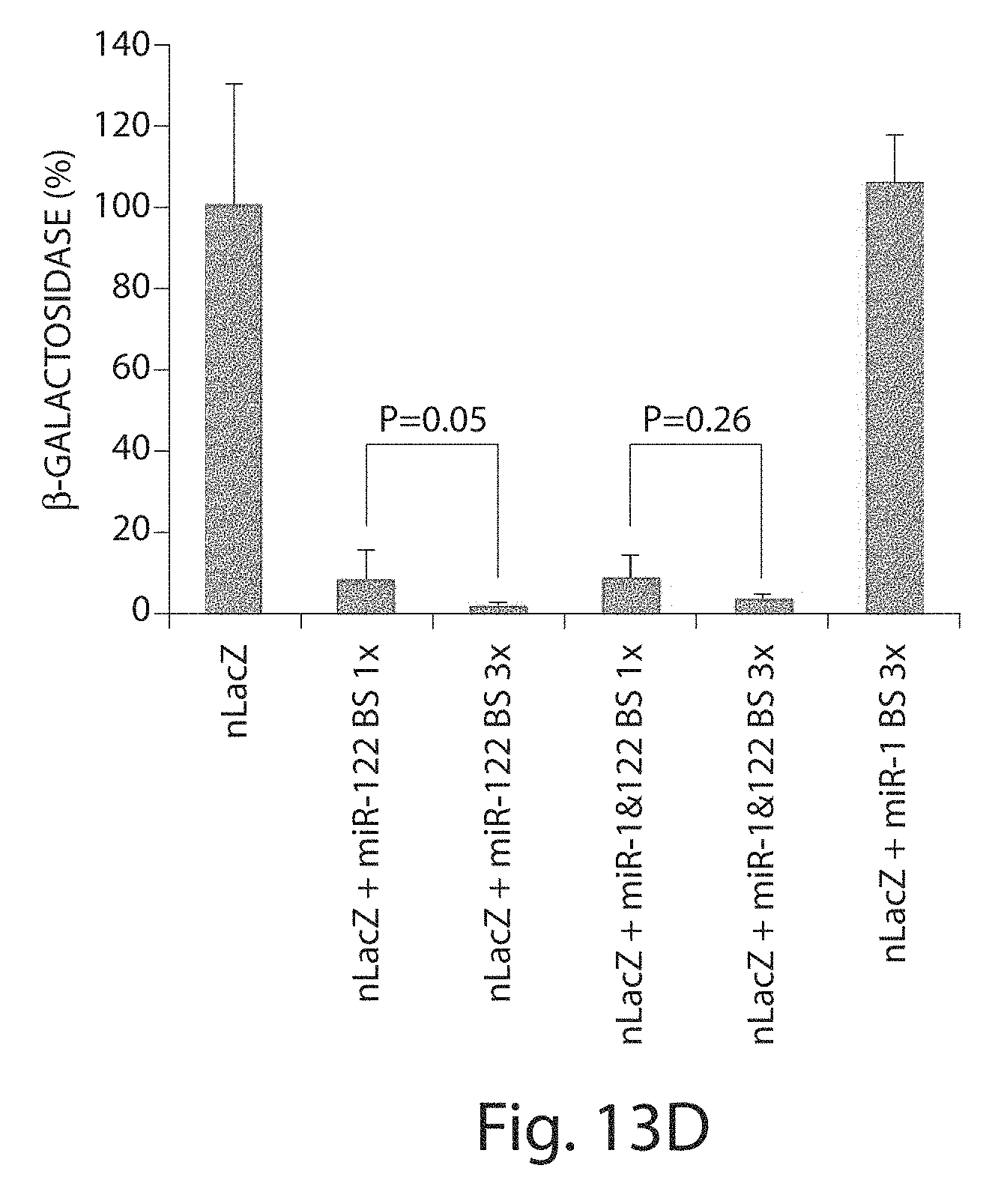
D00017
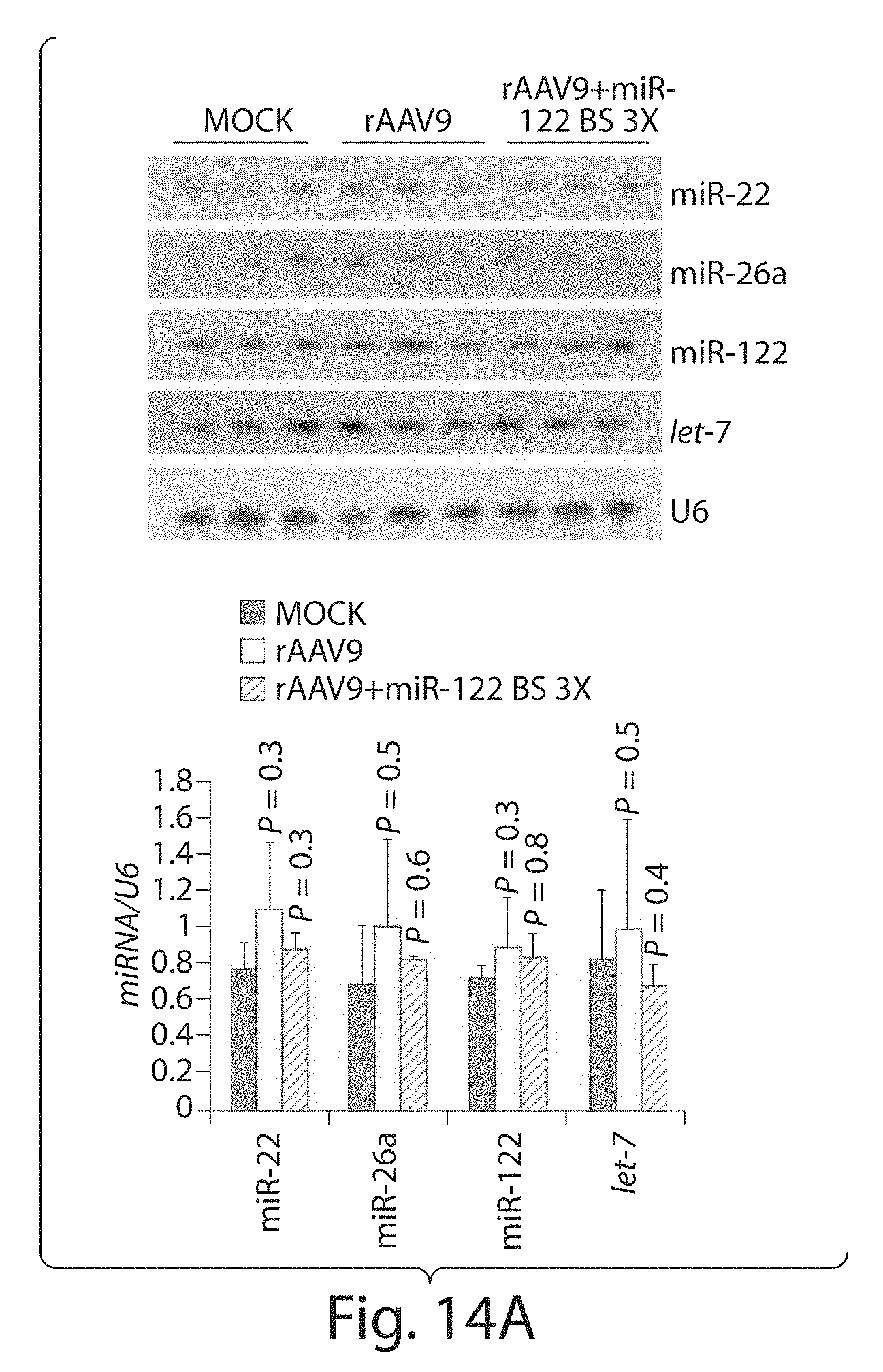

D00018

D00019

D00020

D00021

D00022

D00023
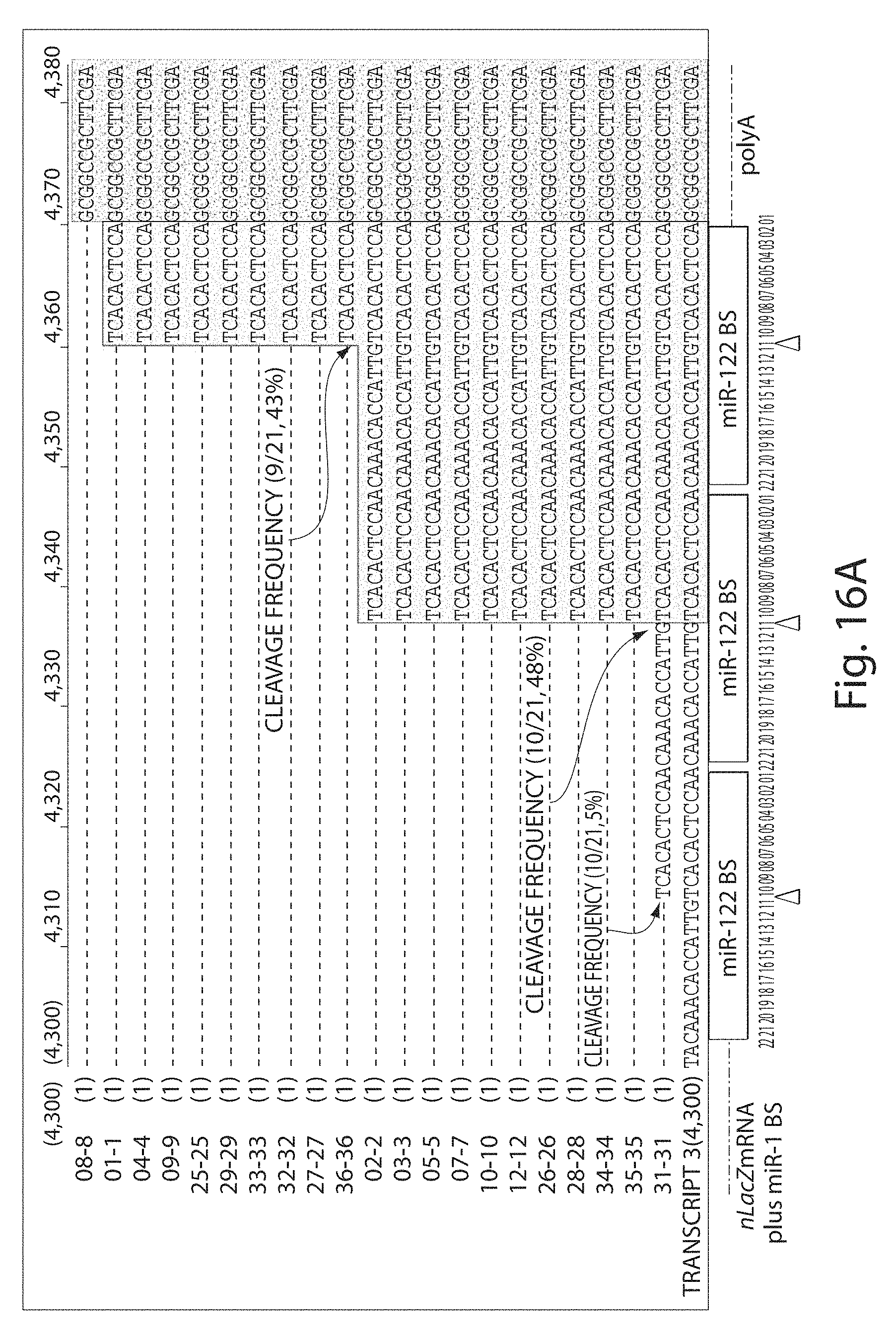

D00024

D00025

D00026

D00027

D00028

D00029

D00030

D00031

D00032

D00033

D00034
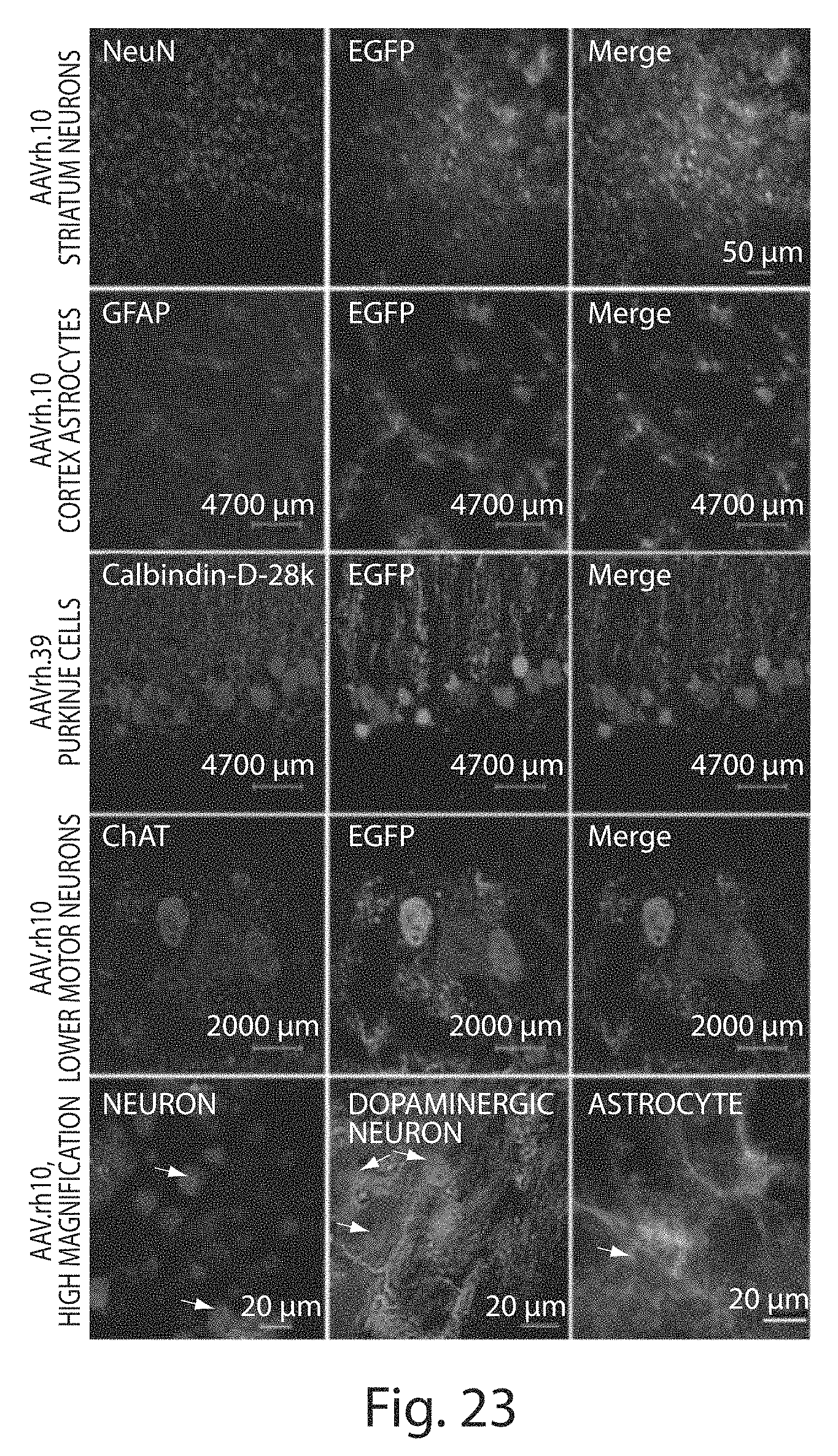
D00035

D00036
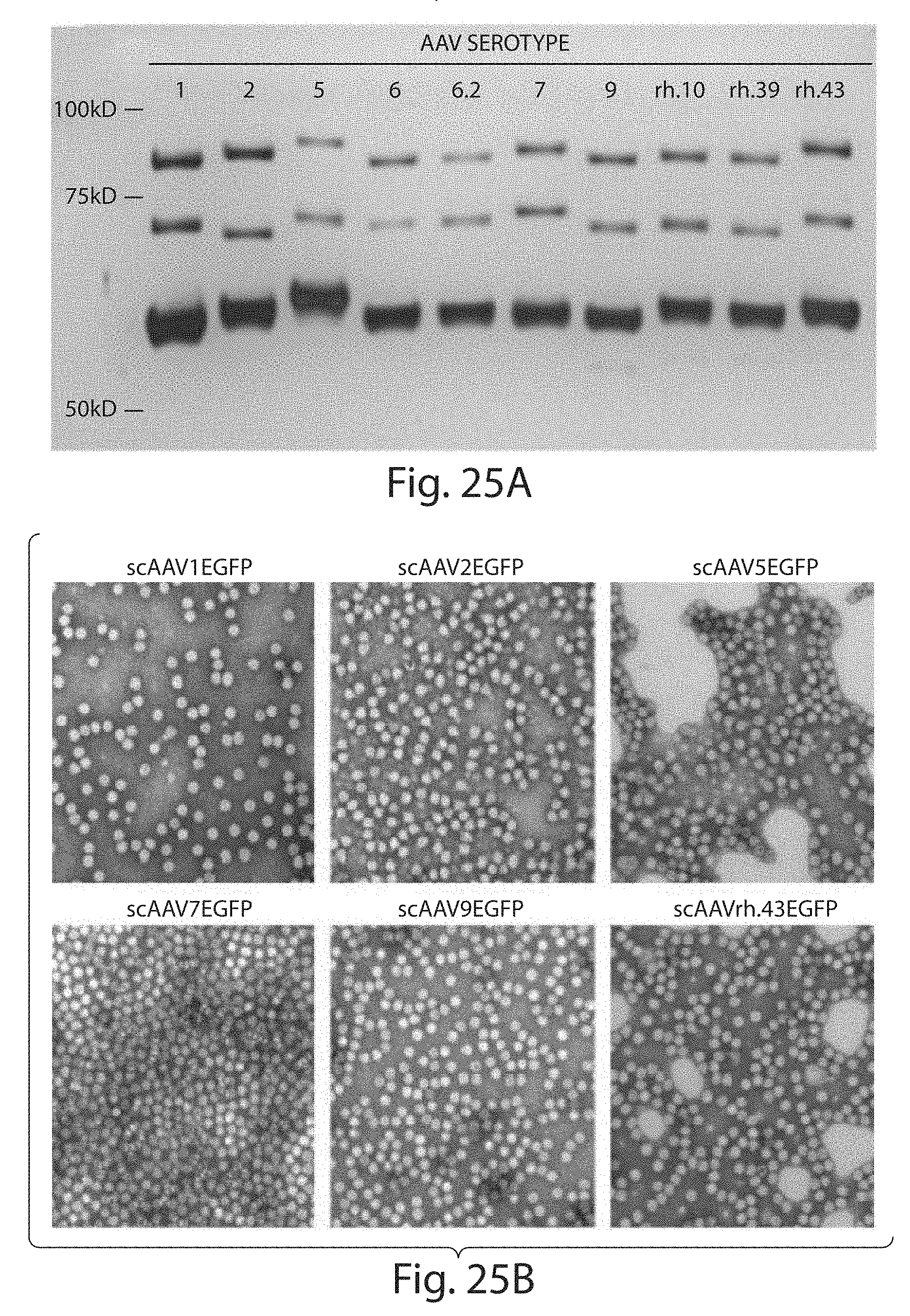
D00037

D00038
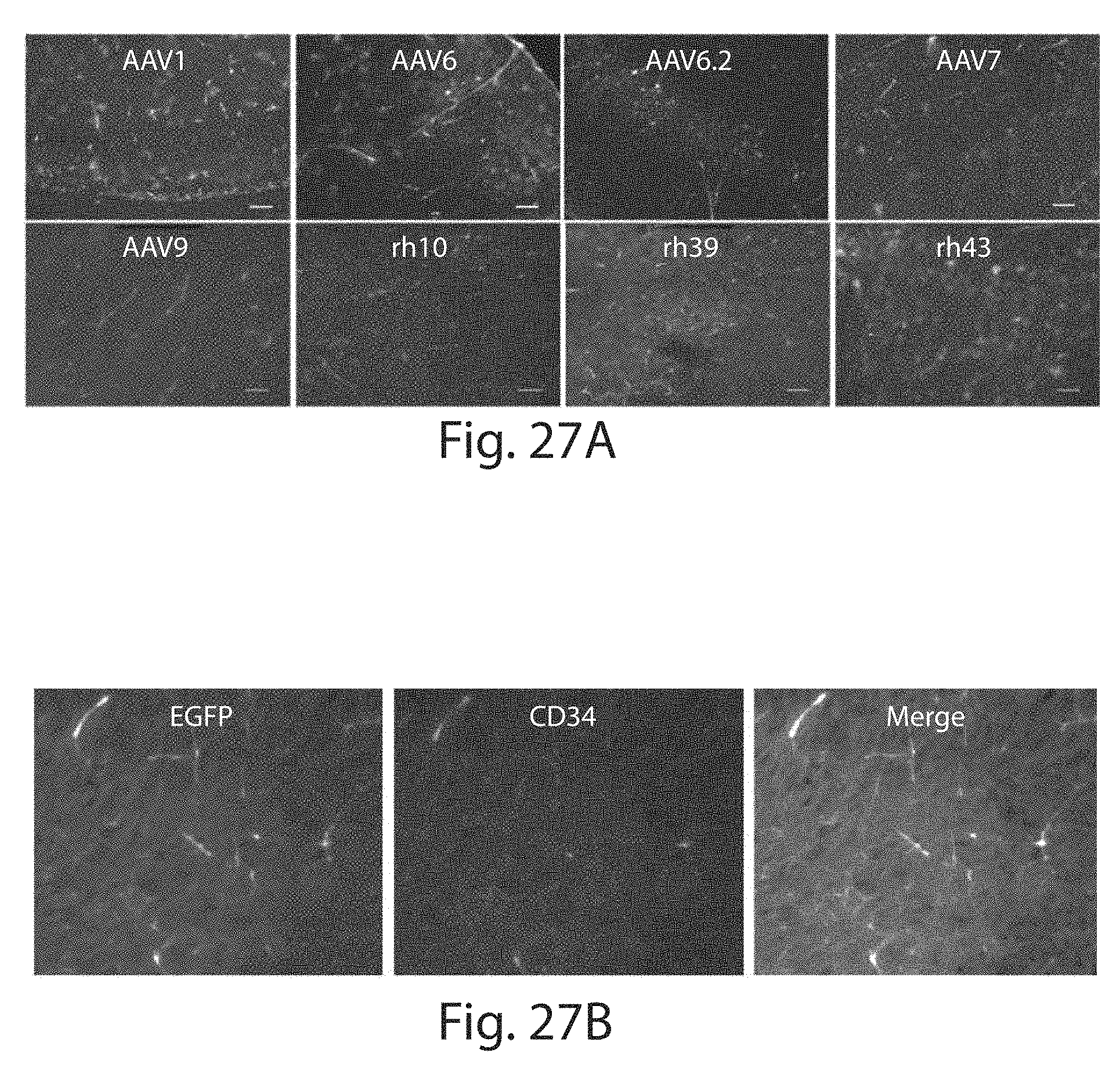
D00039

D00040

D00041

D00042
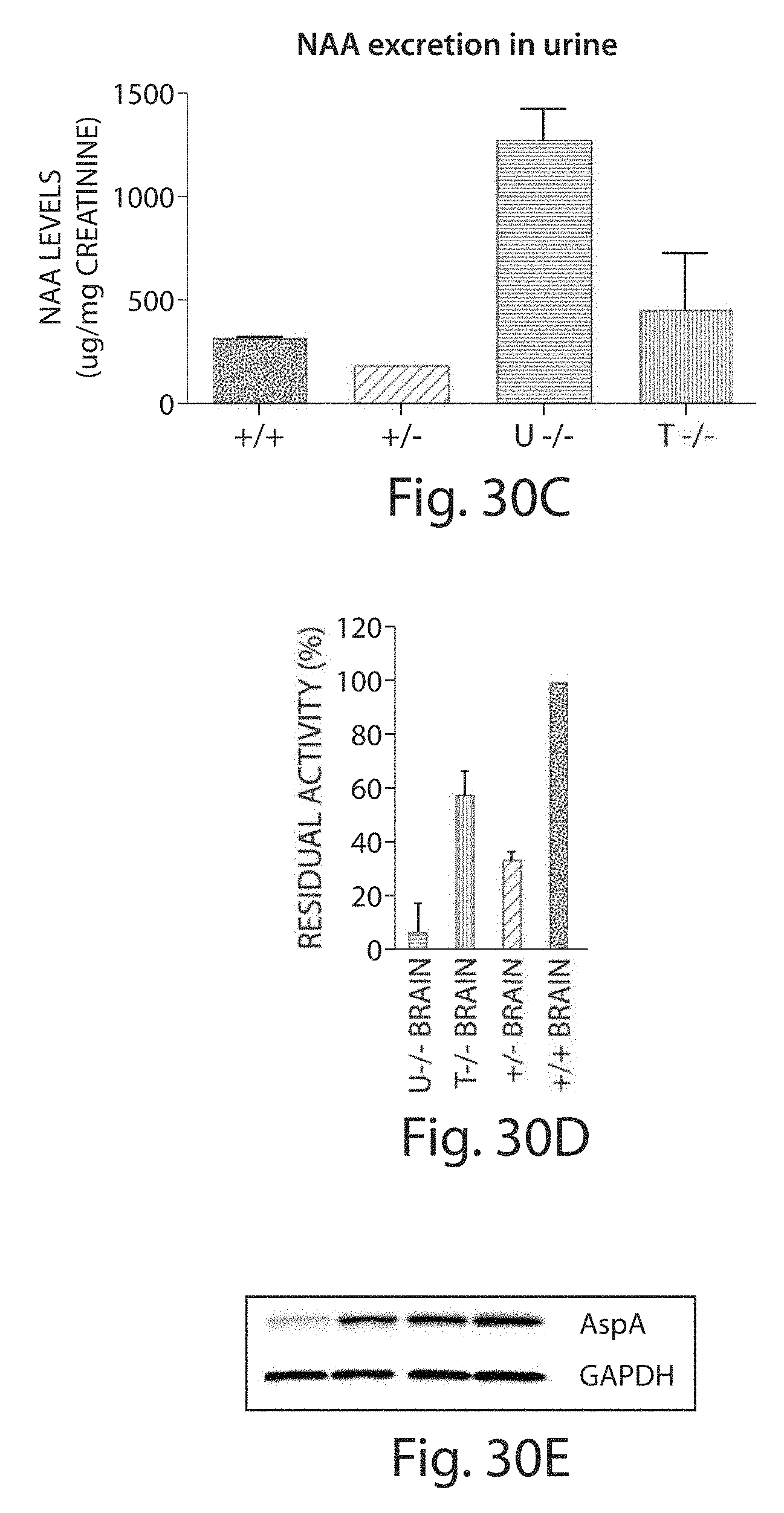
D00043

D00044

D00045

D00046

D00047

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.