Mitochondrial Proteins Constructs And Uses Thereof
Lorberboum-Galski; Haya ; et al.
U.S. patent application number 16/395055 was filed with the patent office on 2019-08-15 for mitochondrial proteins constructs and uses thereof. The applicant listed for this patent is Yissum Research Development Company of the Hebrew University of Jerusalem Ltd.. Invention is credited to Hagar Greif, Haya Lorberboum-Galski.
| Application Number | 20190248846 16/395055 |
| Document ID | / |
| Family ID | 55454113 |
| Filed Date | 2019-08-15 |





View All Diagrams
| United States Patent Application | 20190248846 |
| Kind Code | A1 |
| Lorberboum-Galski; Haya ; et al. | August 15, 2019 |
MITOCHONDRIAL PROTEINS CONSTRUCTS AND USES THEREOF
Abstract
Disclosed are novel fusion protein constructs comprising a functional mitochondrial protein, that can enter mitochondria within intact cells. Further disclosed are methods of treating mitochondrial disorders by the disclosed fusion proteins and compositions therefor.
| Inventors: | Lorberboum-Galski; Haya; (Jerusalem, IL) ; Greif; Hagar; (Ness Ziona, IL) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 55454113 | ||||||||||
| Appl. No.: | 16/395055 | ||||||||||
| Filed: | April 25, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14784253 | Oct 13, 2015 | 10287331 | ||
| PCT/IL2014/050354 | Apr 10, 2014 | |||
| 16395055 | ||||
| 14034224 | Sep 23, 2013 | 8912147 | ||
| 14784253 | ||||
| 61869981 | Aug 26, 2013 | |||
| 61811934 | Apr 15, 2013 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 14/005 20130101; Y10S 530/826 20130101; C12N 2740/16033 20130101; C07K 2319/10 20130101; C12Y 108/01004 20130101; C07K 14/435 20130101; C07K 14/47 20130101; Y10S 530/827 20130101; A61K 9/0019 20130101; C12Y 203/03001 20130101; C07K 2319/21 20130101; A61K 38/00 20130101; C07K 2319/07 20130101; C12N 9/1018 20130101 |
| International Class: | C07K 14/435 20060101 C07K014/435; C07K 14/005 20060101 C07K014/005; C07K 14/47 20060101 C07K014/47; C12N 9/10 20060101 C12N009/10; A61K 9/00 20060101 A61K009/00 |
Claims
1. A fusion protein comprising an HIV-1 transactivator of transcription (TAT) domain fused to a human pyruvate dehydrogenase E1 component alpha subunit (PDHE1a) and a human mitochondria targeting sequence (MTS), wherein the MTS is situated between the TAT domain and the PDHE1a, the PDHE1a is C-terminal to the MTS, and the MTS is heterologous to the PDHE1a.
2. The fusion protein of claim 1, wherein the MTS is a human citrate synthase (CS) MTS.
3. The fusion protein of claim 2, wherein the MTS comprises the amino acid sequence of SEQ ID NO. 23.
4. The fusion protein of claim 1, further comprising a linker covalently linking the TAT domain to the MTS.
5. A pharmaceutical composition comprising a pharmaceutically acceptable carrier and the fusion protein according to claim 1 as an active ingredient.
6. A method for introducing a human pyruvate dehydrogenase E1 component alpha subunit (PDHE1a) into a subject, comprising administering to the subject the fusion protein of claim 1.
7. A method for treating or alleviating a mitochondrial disorder, comprising administering to a human subject in need thereof the fusion protein of claim 1.
8. The method of claim 7, wherein the fusion protein is intravenously administered to the subject.
Description
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] The present application is a continuation application of U.S. Ser. No. 14/784,253, filed Oct. 13, 2015, which is US national stage application of International Application PCT/IL2014/050354, filed Apr. 10, 2014, which is a continuation-in-part of U.S. Ser. No. 14/034,224, filed Sep. 23, 2013, now U.S. Pat. No. 8,912,147, issued Dec. 16, 2014, which claims the benefit under 35 U.S.C. .sctn. 119(e) of U.S. Provisional Application Ser. No. 61/869,981, filed Aug. 26, 2013, and, which claims the benefit under 35 U.S.C. .sctn. 119(e) of U.S. Provisional Application Ser. No. 61/811,934, filed Apr. 15, 2013 the contents of each of which are incorporated by reference in their entireties into the present disclosure.
SEQUENCE LISTING
[0002] The present application contains a Sequence Listing which has been submitted in ASCII format via EFS-Web and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Oct. 13, 2015, is named BIOB-001NO1US_ST25.txt and is 76,765 bytes in size.
TECHNOLOGICAL FIELD
[0003] Disclosed are novel fusion protein constructs comprising a functional mitochondrial protein. Further disclosed are methods of treating mitochondrial disorders by the disclosed fusion proteins and compositions therefor.
PRIOR ART
[0004] References considered to be relevant as background to the presently disclosed subject matter are listed below: [0005] [1] Chinnery P F, Schon E A (2003) Mitochondria. J Neurol Neurosurg Psychiatry 74: 1188-1199. [0006] [2] DiMauro S, Schon E A (2003) Mitochondrial respiratory-chain diseases. N Engl J Med 348: 2656-2668. [0007] [3] Brautigam C A, Chuang J L, Tomchick D R, Machius M, Chuang D T (2005) Crystal structure of human dihydrolipoamide dehydrogenase: NAD+/NADH binding and the structural basis of disease-causing mutations. J Mol Biol 350: 543-552. [0008] [4] Brady R O, Schiffmann R (2004) Enzyme-replacement therapy for metabolic storage disorders. Lancet Neurol 3: 752-756. [0009] [5] Wang D, Bonten E J, Yogalingam G, Mann L, d'Azzo A (2005) Short-term, high dose enzyme replacement therapy in sialidosis mice. Mol Genet Metab 85: 181-189. [0010] [6] Luft F C (2003) Transducing proteins to manipulate intracellular targets. J Mol Med (Berl) 81: 521-523. [0011] [7] Kabouridis P S (2003) Biological applications of protein transduction technology. Trends
[0012] Biotechnol 21: 498-503. [0013] [8] Green M, Loewenstein P M (1988) Autonomous functional domains of chemically synthesized human immunodeficiency virus tat trans-activator protein. Cell 55: 1179-1188. [0014] [9] Frankel A D, Pabo C O (1988) Cellular uptake of the tat protein from human immunodeficiency virus. Cell 55: 1189-1193. [0015] [10] Futaki S, Suzuki T, Ohashi W, Yagami T, Tanaka S, et al. (2001) Arginine-rich peptides. An abundant source of membrane-permeable peptides having potential as carriers for intracellular protein delivery. J Biol Chem 276: 5836-5840. [0016] [11] Schwarze S R, Ho A, Vocero-Akbani A, Dowdy S F (1999) In vivo protein transduction: delivery of a biologically active protein into the mouse. Science 285: 1569-1572. [0017] [12] Guo X, Hutcheon A E, Zieske J D (2004) Transduction of functionally active TAT fusion proteins into cornea. Exp Eye Res 78: 997-1005. [0018] [13] Del Gaizo V, MacKenzie J A, Payne R M (2003) Targeting proteins to mitochondria using TAT. Mol Genet Metab 80: 170-180. [0019] [14] Harding A E (1981) Friedreich's ataxia: a clinical and genetic study of 90 families with an analysis of early diagnostic criteria and intrafamilial clustering of clinical features. Brain 104: 589-620. [0020] [15] Schulz J B, Boesch S, Burk K, Dun A, Giunti P, et al. (2009) Diagnosis and treatment of Friedreich ataxia: a European perspective. Nat Rev Neurol 5: 222-234. [0021] [16] Dun A, Cossee M, Agid Y, Campuzano V, Mignard C, et al. (1996) Clinical and genetic abnormalities in patients with Friedreich's ataxia. N Engl J Med 335: 1169-1175. [0022] [17] Campuzano V, Montermini L, Lutz Y, Cova L, Hindelang C, et al. (1997) Frataxin is reduced in Friedreich ataxia patients and is associated with mitochondrial membranes. Hum Mol Genet 6: 1771-1780. [0023] [18] Rotig A, de Lonlay P, Chretien D, Foury F, Koenig M, et al. (1997) Aconitase and mitochondrial iron-sulphur protein deficiency in Friedreich ataxia. Nat Genet 17: 215-217. [0024] [19] Lodi R, Cooper J M, Bradley J L, Manners D, Styles P, et al. (1999) Deficit of in vivo mitochondrial ATP production in patients with Friedreich ataxia. Proc Natl Acad Sci USA 96: 11492-11495. [0025] [20] Delatycki M B, Camakaris J, Brooks H, Evans-Whipp T, Thorburn D R, et al. (1999) Direct evidence that mitochondrial iron accumulation occurs in Friedreich ataxia. Ann Neurol 45: 673-675. [0026] [21] Tsou A Y, Friedman L S, Wilson R B, Lynch D R (2009) Pharmacotherapy for Friedreich ataxia. CNS Drugs 23: 213-223. [0027] [22] Perlman S L (2012) A review of Friedreich ataxia clinical trial results. J Child Neurol 27: 1217-1222. [0028] [23] Rapoport M, Saada A, Elpeleg O, Lorberboum-Galski H (2008) TAT-mediated delivery of LAD restores pyruvate dehydrogenase complex activity in the mitochondria of patients with LAD deficiency. Mol Ther 16: 691-697. [0029] [24] Rapoport M, Salman L, Sabag O, Patel M S, Lorberboum-Galski H (2011) Successful TAT-mediated enzyme replacement therapy in a mouse model of mitochondrial E3 deficiency. J Mol Med (Berl) 89: 161-170. [0030] [25] Vyas P M, Tomamichel W J, Pride P M, Babbey C M, Wang Q, et al. (2012) A TAT-frataxin fusion protein increases lifespan and cardiac function in a conditional Friedreich's ataxia mouse model. Hum Mol Genet 21: 1230-1247. [0031] [26] Gakh O, Cavadini P, Isaya G (2002) Mitochondrial processing peptidases. Biochim Biophys Acta 1592: 63-77. [0032] [27] Cavadini P, Adamec J, Taroni F, Gakh O, Isaya G (2000) Two-step processing of human frataxin by mitochondrial processing peptidase. Precursor and intermediate forms are cleaved at different rates. J Biol Chem 275: 41469-41475. [0033] [28] Schmucker S, Argentini M, Carelle-Calmels N, Martelli A, Puccio H (2008) The in vivo mitochondrial two-step maturation of human frataxin. Hum Mol Genet 17: 3521-3531. [0034] [29] Gakh O, Bedekovics T, Duncan S F, Smith D Yt, Berkholz D S, et al. (2010) Normal and Friedreich ataxia cells express different isoforms of frataxin with complementary roles in iron-sulfur cluster assembly. J Biol Chem 285: 38486-38501. [0035] [30] Gavel Y, von Heijne G (1990) Cleavage-site motifs in mitochondrial targeting peptides. Protein Eng 4: 33-37. [0036] [31] Braun H P, Schmitz U K (1997) The mitochondrial processing peptidase. Int J Biochem Cell Biol 29: 1043-1045. [0037] [32] Horwich A (1990) Protein import into mitochondria and peroxisomes. Curr Opin Cell Biol 2: 625-633. [0038] [33] Saada A, Edvardson S, Rapoport M, Shaag A, Amry K, et al. (2008) C6ORF66 is an assembly factor of mitochondrial complex I. Am J Hum Genet 82: 32-38. [0039] [34] Cheng T L, Liao C C, Tsai W H, Lin C C, Yeh C W, et al. (2009) Identification and characterization of the mitochondrial targeting sequence and mechanism in human citrate synthase. J Cell Biochem 107: 1002-1015. [0040] [35] Santos R, Lefevre S, Sliwa D, Seguin A, Camadro J M, et al. (2010) Friedreich ataxia: molecular mechanisms, redox considerations, and therapeutic opportunities. Antioxid Redox Signal 13: 651-690. [0041] [36] Bulteau A L, O'Neill H A, Kennedy M C, Ikeda-Saito M, Isaya G, et al. (2004) Frataxin acts as an iron chaperone protein to modulate mitochondrial aconitase activity. Science 305: 242-245. [0042] [37] Richardson T E, Yu A E, Wen Y, Yang S H, Simpkins J W (2012) Estrogen prevents oxidative damage to the mitochondria in Friedreich's ataxia skin fibroblasts. PLoS One 7: e34600. [0043] [38] WO 2009/098682. [0044] [39] http://www.curefa.org/pipeline.html. [0045] [40] Tang N, et al. (2012) Stable Overexpression of Arginase I and Ornithine Transcarbamylase in HepG2 Cells Improves Its Ammonia Detoxification. Journal of Cellular Biochemistry 113: 518-527. [0046] [41] Cunningham, S. C. et al. (2011) Induction and Prevention of Severe Hyperammonemia in the spf.sup.ash Mouse Model of Ornithine Transcarbamylase Deficiency Using shRNA and rAAV-mediated Gene Delivery. Molecular Therapy 19(5):854-9.
[0047] Acknowledgement of the above references herein is not to be inferred as meaning that these are in any way relevant to the patentability of the presently disclosed subject matter.
BACKGROUND
[0048] Mitochondria play a major and critical role in cellular homeostasis--they participate in intracellular signaling and apoptosis, and perform numerous biochemical tasks, such as in pyruvate oxidation, in the citric acid cycle (also referred to as the Krebs cycle), and in metabolism of amino acids, fatty acids, nucleotides and steroids. However, the most crucial task of mitochondria is their role in cellular energy metabolism. This includes .beta.-oxidation of fatty acids and production of ATP by means of the electron-transport chain and the oxidative-phosphorylation system [1, 2].
[0049] Most of the approximately 900 gene products in the mitochondria are encoded by nuclear DNA (nDNA) while mitochondrial DNA (mtDNA) only contains 13 protein encoding genes. Most of the polypeptides encoded by nDNA genes are synthesized with a mitochondrial targeting sequence (MTS), allowing their import from the cytoplasm into mitochondria through the translocation machinery (TOM/TIM). Upon entering the mitochondria, the MTS is recognized and cleaved off, allowing for proper processing and, if necessary, assembly into mitochondrial enzymatic complexes [3].
[0050] Currently, there is no cure for genetic mitochondrial metabolic disorders and treatment is mostly palliative.
[0051] Enzyme or Protein Replacement Therapy (E/PRT) is a therapeutic approach for metabolic disorders, whereby the deficient or absent protein/enzyme is artificially manufactured, purified and administered intravenously to the patient in need thereof on a regular basis.
[0052] After many years of extensive research, E/PRT has been successfully accepted as the treatment of choice for metabolic lysosomal storage diseases, including Gaucher disease, Fabry disease and attenuated variants of mucopolysaccaridoses type 1 (MPS 1). However, the inability of the intravenously administered enzymes to penetrate the blood-brain barrier severely limits the application of this approach for the treatment of other metabolic disorders that involve the central nervous system (CNS) [4, 5].
[0053] One approach for delivering proteins into cells is their fusion with protein transduction domains (PTDs), a group of short peptides that serve as delivery vectors for large molecules. Generally, PTDs are defined as short, water-soluble and partly hydrophobic, and/or polybasic peptides (at most 30-35 amino acids residues) with a net positive charge at physiological pH [6, 7]. The main feature of PTDs is that they are able to penetrate the cell membrane at low micromolar concentrations both in vitro and in vivo without using any chiral receptors and without causing significant membrane damage. Furthermore, and even more importantly, these peptides are capable of internalizing electrostatically or covalently bound biologically active cargoes (such as drugs) with high efficiency and with low toxicity. The mechanism(s) by which PTDs enter the cells has not been completely understood. One of the well-characterized PTDs is the transactivator of transcription (TAT) peptide originating from the HIV-1 virus. TAT is an 11-amino-acid (residues 47-57) arginine- and lysine-rich portion of the Tat protein encoded by HIV-1 virus [8, 9]. TAT-fusion proteins have been previously shown to be rapidly and efficiently introduced into cultured cells, intact tissue and live tissues when injected into mice [10-12]. It has also been demonstrated that TAT fusion proteins traverse mitochondrial membranes [13, 38].
[0054] There has been great progress in the use of PTD-fusion proteins for the delivery of different macromolecules into cells both in vitro and in vivo. This system can be used even for the delivery of cargoes into the brain across the blood-brain barrier. In addition, the ability to target specific intracellular sub-localizations, such as the nuclei, the mitochondria and lysosomes, further expands the possibilities of this delivery system to the development of sub-cellular organelle-targeted therapy. The therapeutic applications seem almost unlimited, and the use of the TAT-based delivery system has extended from proteins to a large variety of cargoes such as oligonucleotides, imaging agents, low molecular mass drugs, nanoparticles, micelles and liposomes. As will be shown, this PTD system is used for developing fusion constructs of functional mitochondrial proteins, for treatment of mitochondrial disorders, for example Friedreich ataxia.
[0055] Friedreich ataxia is an autosomal recessive degenerative disorder characterized by ataxia, areflexia, sensory loss, weakness, scoliosis, and cardiomyopathy. Diabetes mellitus, optic neuropathy, and hearing loss are also seen in patients suffering from this disease [14, 15]. Most patients with Friedreich ataxia (97%) have expansions of a GAA repeat in the first intron on both alleles of the gene encoding the mitochondrial protein frataxin [15, 16] whose expression is reduced in Friedreich ataxia [17]. The size of the GAA repeat expansion inversely correlates with frataxin expression and with the age of disease onset [16]. A deficiency of frataxin in cells leads to decreased activities of mitochondrial iron-sulfur cluster-containing enzymes, to an accumulation of iron in the mitochondrial matrix, increased sensitivity to oxidative stress, as well as to impaired adenosine triphosphate (ATP) production [18-20].
[0056] Current targets for disease-modifying drug development include agents targeting the mitochondria, aimed to (1) reduce oxidative stress and free-radical generation; (2) improve ATP production; (3) reduce iron accumulation; and (4) increase frataxin production and the assembly of iron-sulfur clusters [21]. There are presently 21 agents or classes of therapeutic agents enrolled in the research pipeline of Friedreich ataxia disease [39]. Millions of dollars from public, private, and industry-based initiatives have been dedicated to research of Friedreich ataxia therapeutics. Despite this vigorous international effort, there is as yet no proven disease-modifying therapy for Friedreich ataxia [22].
[0057] Development of E/PRT using the TAT delivery system in mitochondrial disorders was previously reported for lipoamide dehydrogenase (LAD) mitochondrial deficiency [23, 38]. LAD is the E3 subunit of the three .alpha.-ketoacid dehydrogenase complexes in the mitochondrial matrix, which are crucial for the metabolism of carbohydrates and amino acids. These complexes are the pyruvate dehydrogenase complex (PDHC), the .alpha.-ketoglutarate dehydrogenase complex (KGDHC) and the branched chain ketoacid dehydrogenase complex (BCKDHC). This previously reported TAT delivery system was based on a TAT-LAD fusion protein comprising the natural precursor sequence of the human LAD containing the N-terminal 35 amino acid mitochondrial targeting sequence (MTS). The natural MTS of LAD was used to facilitate processing of the TAT-LAD construct upon delivery into the mitochondria, thus allowing the incorporation of the delivered LAD into the a-ketoacid dehydrogenase complexes. This TAT-LAD construct was demonstrated to enter patients' cells rapidly, and efficiently reaching the mitochondria. Inside the mitochondria, TAT-LAD was shown to be processed and to restore LAD activity [23]. Moreover, delivery of TAT-LAD into E3-deficient mice tissues was also demonstrated [24]. In mice tissues, a single administration of TAT-LAD resulted in a significant increase in the enzymatic activity of the mitochondrial multienzyme complex pyruvate-dehydrogenase complex within the liver, heart and, most importantly, brain of TAT-LAD-treated E3-deficient mice [24].
[0058] Notably, TAT-LAD was shown to be able to restore the activity of the pyruvate dehydrogenase complex (PDHC) within treated patients' cells almost back to its normal levels. PDHC is a 9.5.times.10.sup.6 Da macromolecular machine whose multipart structure assembly process involves numerous different subunits: a pentagonal core of 60 units of the E2 component (dihydrolipoamide), attached to 30 tetramers of the E1 component (.alpha.2.beta.2) (pyruvate decarboxylase), 12 dimers of the E3 (LAD, dihydrolipoamide) component and 12 units of the E3 binding protein. The structure of all a-ketoacid dehydrogenase complexes is similar to that of PDHC. The complexity of this structure emphasizes the significance in showing that TAT-mediated replacement of one mutated component restores the activity of an essential mitochondrial multi-component enzymatic complex in cells of enzyme-deficient patients.
[0059] Previous studies of mitochondria delivery system primarily used the native MTS of mitochondrial proteins (e.g. LAD) and showed that the native MTS was necessary for maximal restoration of LAD enzymatic function. Deleting the MTS restored a significantly smaller amount of LAD activity within the mitochondria. Since TAT can move both ways across a membrane and thus pull the therapeutic cargo out of the mitochondria, when MTS is included, the matrix processing peptidases recognizes the sequence and clips it, and the cargo (e.g. mature LAD) is left in the mitochondrial matrix while the TAT peptide can transduce out of the mitochondrion. Repeated dosing should therefore result in accumulating amounts of cargo in the mitochondria over time.
[0060] A TAT-FRATAXIN (TAT-FRA, also referred to as TAT-FXN) fusion protein for putative treatment of Friedreich's ataxia was recently reported [25]. This TAT-FXN fusion protein was shown to bind iron in vitro, transduce into the mitochondria of Friedreich ataxia deficient fibroblasts and also reduce caspase-3 activation in response to an exogenous iron-oxidant stress. In this TAT-FXN fusion protein, the authors used the native MTS of frataxin that consists of 80 amino acid residues (aa) for preparing their TAT-FXN fusion protein [26].
[0061] It is known that FXN mRNA is translated to a precursor polypeptide that is transported to the mitochondrial matrix and processed to at least two forms, namely FXN42-210 and FXN81-210. FXN42-210 is a transient processing intermediate, whereas FXN81-210 represents the mature protein [27, 28]. However, it was found that both FXN42-210 and FXN81-210 are present in control cell lines and tissues at steady-state, and that FXN42-210 is consistently more depleted than FXN81-210 in samples from Friedreich's ataxia patients [29].
[0062] Most nuclear-encoded mitochondrial proteins contain a cleavable N-terminal MTS that directs mitochondrial targeting of the protein; as detailed above, the N-terminal MTS is cleaved off by matrix processing proteases at a well-conserved RXY .dwnarw. (S/A) motif, which is a three amino acid (aa) motif, where X can be any aa, followed by serine or alanine and cleavage is performed after the three first amino acids [26, 30-31]. These N-terminal MTSs are typically 15-30 amino acids in length including 3-5 nonconsecutive basic amino acid (arginine/lysine) residues, often with several serine/threonine residues but without acidic amino acid (asparate/glutamate) residues. In their molecular structure, these MTSs are able to form strong basic amphipathic .alpha.-helices that are essential for efficient mitochondrial transportation [32]. Thus, by way of example, the long 80-aa native MTS of frataxin as well as its two-step processing can reduce its efficiency in the delivery of cargos into the matrix of the mitochondria.
SUMMARY OF THE INVENTION
[0063] Provided is a fusion protein comprising a HIV-1 transactivator of transcription (TAT) domain, a functional human mitochondrial protein and a human mitochondria targeting sequence (MTS) situated between said TAT domain and said functional human mitochondrial protein and wherein said human MTS is heterologous to said functional human mitochondrial protein.
[0064] In the disclosed fusion protein said functional human mitochondrial protein can be situated C-terminal to said human MTS.
[0065] In the disclosed fusion protein said human mitochondrial protein can be a functional human mitochondrial protein per se and/or a component of a mitochondrial multi-component complex, for example human frataxin and ornithine transcarbamoylase (OTC).
[0066] In the disclosed fusion protein, said MTS can comprise from about 15 to about 40 amino acid residues, including from about 3 to about 5 nonconsecutive basic amino acid residues, and optionally from about 1 to about 3 or 4 or 5 serine/threonine residues.
[0067] Non-limiting examples of the MTS comprised in the disclosed fusion protein are any one of human mitochondrial citrate synthase MTS (the amino acid and the nucleic acid sequence encoding therefor are denoted by SEQ ID NO. 23 and SEQ ID NO. 3, respectively), the human lipoamide dehydrogenase MTS (the amino acid and the nucleic acid sequence encoding therefor are denoted by SEQ ID NO. 24 and SEQ ID NO. 5, respectively), the MTS of the human C6ORF66 gene product (the amino acid and the nucleic acid sequence encoding therefor are denoted by SEQ ID NO. 25 and SEQ ID NO. 4, respectively) and the MTS of human mitochondrial GLUD2 (encoded by the nucleic acid sequence denoted by SEQ ID NO. 16).
[0068] Further provided is a fusion protein comprising a HIV-1 transactivator of transcription (TAT) domain fused to a functional human mitochondrial protein and a human mitochondria targeting sequence (MTS) of a human mitochondrial protein selected from citrate synthase (CS) and lipoamide dehydrogenase (LAD) situated between said TAT domain and said functional human mitochondrial protein, wherein said functional human mitochondrial protein is C-terminal to said MTS of human lipoamide dehydrogenase or human citrate synthase.
[0069] The disclosed fusion protein as herein defined may further comprise a linker covalently linking said TAT domain to said MTS sequence.
[0070] In the disclosed fusion protein, the fusion protein may have the amino acid sequence denoted by SEQ ID NO. 30, comprising a HIV-1 transactivator of transcription (TAT) domain having the amino acid sequence denoted by SEQ ID NO. 27 fused to human frataxin having the amino acid sequence denoted by SEQ ID NO. 26 and a mitochondria targeting sequence (MTS) of human lipoamide dehydrogenase having the amino acid sequence denoted by SEQ ID NO. 24, said MTS situated between said TAT domain and said frataxin, and is linked to said TAT domain via a linker having the amino acid sequence denoted by SEQ ID NO. 32, and wherein said frataxin is C-terminal to said MTS of human lipoamide dehydrogenase.
[0071] In further embodiments of the disclosed fusion protein, the fusion protein may have the amino acid sequence denoted by SEQ ID NO. 28, comprising a HIV-1 transactivator of transcription (TAT) domain having the amino acid sequence denoted by SEQ ID NO. 27 fused to human frataxin having the amino acid sequence denoted by SEQ ID NO. 26 and a mitochondria targeting sequence (MTS) of human citrate synthase having the amino acid sequence denoted by SEQ ID NO. 23, said MTS situated between said TAT domain and said frataxin, and is linked to said TAT domain via a linker having the amino acid sequence denoted by SEQ ID NO. 32, and wherein said frataxin is C-terminal to said MTS of human citrate synthase.
[0072] Further disclosed is a composition comprising a physiologically acceptable carrier and as an active ingredient a fusion protein as disclosed herein, and a pharmaceutical composition comprising a physiologically or a pharmaceutically acceptable carrier and as an active ingredient a fusion protein as disclosed herein.
[0073] Further disclosed is a pharmaceutical composition for restoring, at least in part, activity of a defective or deficient or unfunctional human mitochondrial protein in a subject in need. The said human mitochondrial protein can be active per se, or can be a member of a functional mitochondrial protein complex.
[0074] Further disclosed is a composition comprising a physiologically acceptable carrier and as an active ingredient a fusion protein comprising a HIV-1 transactivator of transcription (TAT) domain fused to human frataxin and a human mitochondria targeting sequence (MTS) of a human mitochondrial protein selected from citrate synthase (CS) and lipoamide dehydrogenase (LAD) situated between said TAT domain and said frataxin, wherein said frataxin is C-terminal to said MTS of human lipoamide dehydrogenase or human citrate synthase, as disclosed herein.
[0075] The present disclosure further provides a composition comprising as an active ingredient a fusion protein having the amino acid sequence denoted by SEQ ID NO. 30 or a fusion protein having the amino acid sequence denoted by SEQ ID NO. 28 and a physiologically acceptable carrier.
[0076] The pharmaceutical composition disclosed herein can be intended for treating or alleviating a mitochondrial disorder, such as but not limited to Friedreich's ataxia or any other disorder associated with deficiency of frataxin or defective frataxin or a disorder associated with a deficiency of OTC or with defective OTC.
[0077] Further disclosed is a pharmaceutical composition for the treatment of Friedreich's Ataxia by intravenous administration to a subject in need thereof, said composition comprising a therapeutically effective amount of a fusion protein having the amino acid sequence denoted by SEQ ID NO. 30 or a fusion protein having the amino acid sequence denoted by SEQ ID NO. 28 and at least one of pharmaceutically acceptable carrier, diluent, additive and excipient.
[0078] A non-limiting example of a pharmaceutical composition as herein defined is wherein the therapeutically effective amount administered is from about 0.5 mg/Kg to about 2 mg/Kg body weight of the subject.
[0079] Thus, the disclosed fusion protein can be used in a method for the treatment of a mitochondrial disorder, such as but not limited to Friedreich's ataxia or any other disorder associated with deficiency of frataxin or defective frataxin or, respectively a disorder associated with a deficiency of OTC or with defective OTC.
[0080] Further provided is a method for treating or alleviating a mitochondrial disorder, said method comprising the step of administering to a subject in need of such treatment a therapeutically effective amount of the fusion protein disclosed herein, thereby treating a mitochondria disorder.
[0081] In some embodiments of the disclosed method, the functional protein is frataxin, respectively OTC, and the mitochondrial disorder is Friedreich's ataxia or any other disorder associated with deficiency of frataxin or defective frataxin or, respectively, a disorder associated with a deficiency of OTC or with defective OTC.
[0082] The disclosed method for treating or alleviating a mitochondrial disorder can further comprises administering an additional therapeutic agent.
[0083] Further provided is a method for introducing a functional mitochondrial protein into mitochondria of a subject, said method comprising the step of administering to a subject in need of such treatment a therapeutically effective amount of the fusion protein as disclosed herein, thereby introducing a functional human mitochondrial protein into the mitochondria of a subject in need thereof.
[0084] In the disclosed method for introducing a functional mitochondrial protein into mitochondria of a subject, said introduced functional human mitochondrial protein may restore at least partial activity of a wild type human mitochondrial protein, at least 5%, at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90% or up to 100% of the activity of a wild type human mitochondrial protein.
[0085] Further disclosed is a method for restoring, at least in part, activity of a defective or deficient or un-functional human mitochondrial protein in a subject in need, by administering to said subject a therapeutically effective amount of a fusion protein according to the present disclosure. The said human mitochondrial protein can be active per se, or can be a member of a functional mitochondrial protein complex.
[0086] Still further, provided is a method for alleviating oxidative stress in a subject in need thereof, said method comprising the step of administering to a subject in need of such treatment a therapeutically effective amount of the fusion protein as disclosed herein, thereby alleviating oxidative stress in said subject.
BRIEF DESCRIPTION OF THE DRAWINGS
[0087] In order to better understand the subject matter that is disclosed herein and to exemplify how it may be carried out in practice, embodiments will now be described, by way of non-limiting example only, with reference to the accompanying drawings, in which:
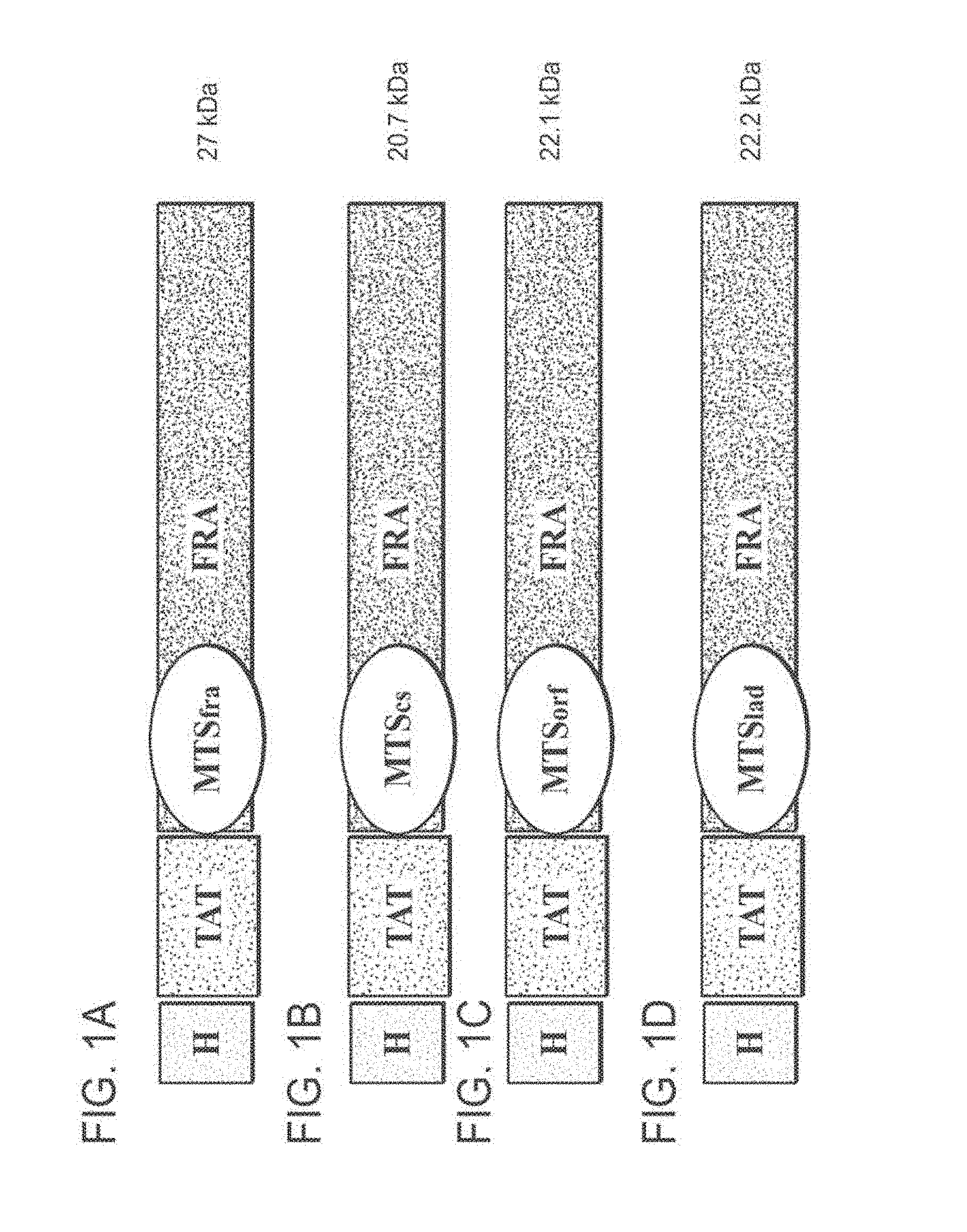
[0088] FIG. 1A-1D: Schematic structures of the various TAT-MTS-FRA fusion proteins and the expected molecular weights thereof. Abbreviations: H, His tag; TAT, transactivator of transcription; FRA (or fra), Frataxin; MTS, mitochondrial translocation sequence; cs, Citrate synthase; orf, C6ORF66; lad, LAD and kDa, kilo Dalton.
[0089] FIG. 2A-2D: Expression and sub-cellular localization of TAT-MTSfra-FRA and of TAT-MTSorf-FRA fusion proteins
[0090] FIG. 2 presents an image of SDS-PAGE analysis of bacterial sub-fractions expressing the TAT-MTSfra-FRA fusion protein (FIG. 2A) and an immunoblot of Western blot analysis thereof (FIG. 2B) using anti-His antibody; an image of SDS-PAGE analysis of bacterial sub-fractions expressing TAT-MTSorf-FRA fusion protein (FIG. 2C) and an immunoblot of Western blot analysis thereof using anti-His antibody is presented in FIG. 2D.
[0091] Abbreviations (for FIGS. 2A-2D): 1, whole cell extract; 2, soluble fraction; 3, insoluble fraction; kDa, kilo Dalton; and M=marker. Arrow heads indicate the fusion proteins.
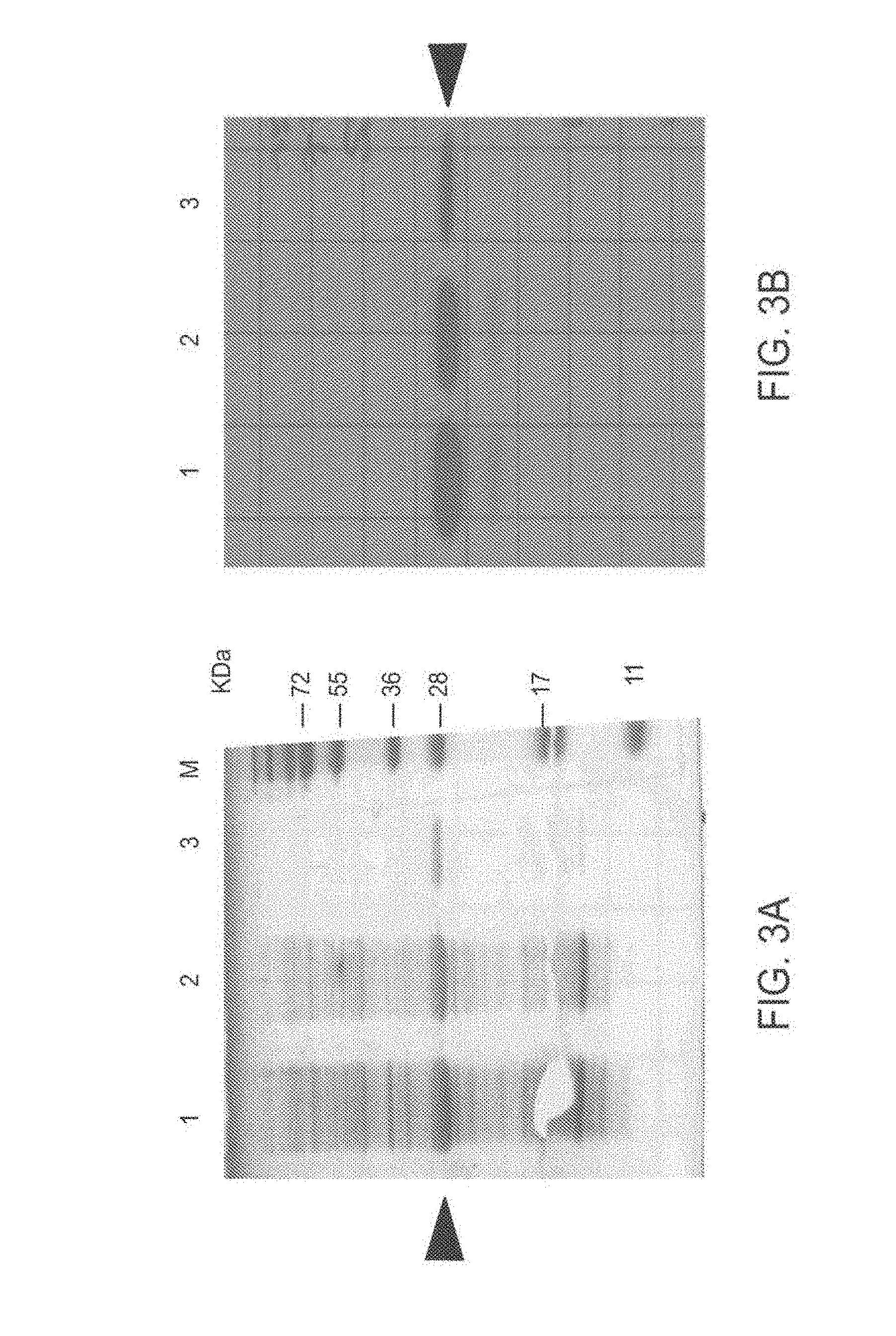
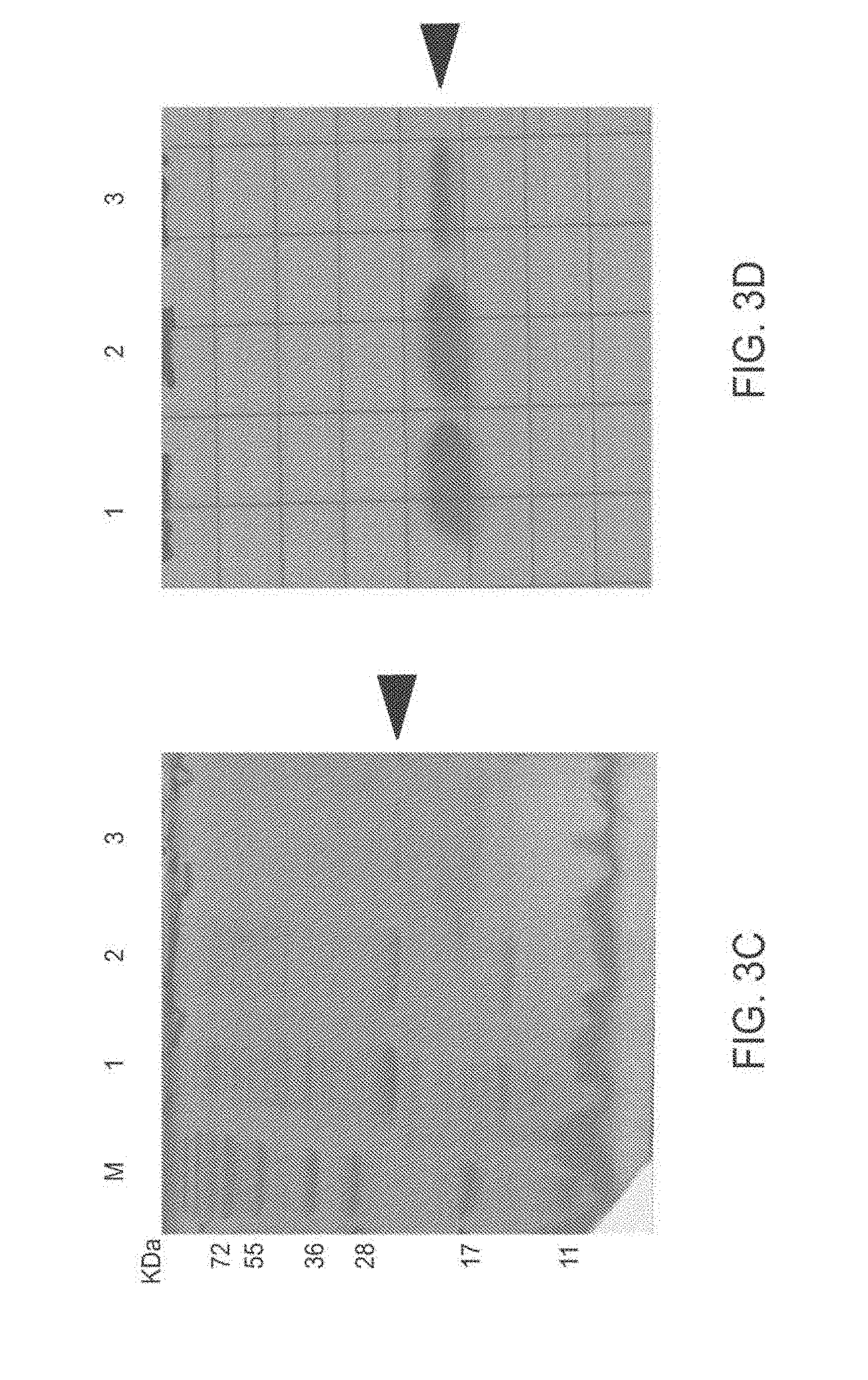
[0092] FIG. 3A-3D: Expression and sub-cellular localization of TAT-MTSlad-FRA and TAT-MTScs-FRA fusion proteins
[0093] FIG. 3 presents an image of SDS-PAGE analysis of bacteria sub-fractions expressing the TAT-MTSlad-FRA fusion protein (FIG. 3A) and an immunoblot of Western blot analysis thereof (FIG. 3B) using anti-His antibody; an image of SDS-PAGE analysis of bacterial sub-fractions expressing TAT-MTScs-FRA fusion protein (FIG. 3C) and an immunoblot of Western blot analysis thereof using anti-His antibody is presented in FIG. 3D.
[0094] Abbreviations (for FIG. 3A-3D): 1, whole cell extract; 2. soluble fraction; 3, insoluble fraction; KDa, kilo Dalton; and M=marker. Arrow heads indicate the fusion proteins.
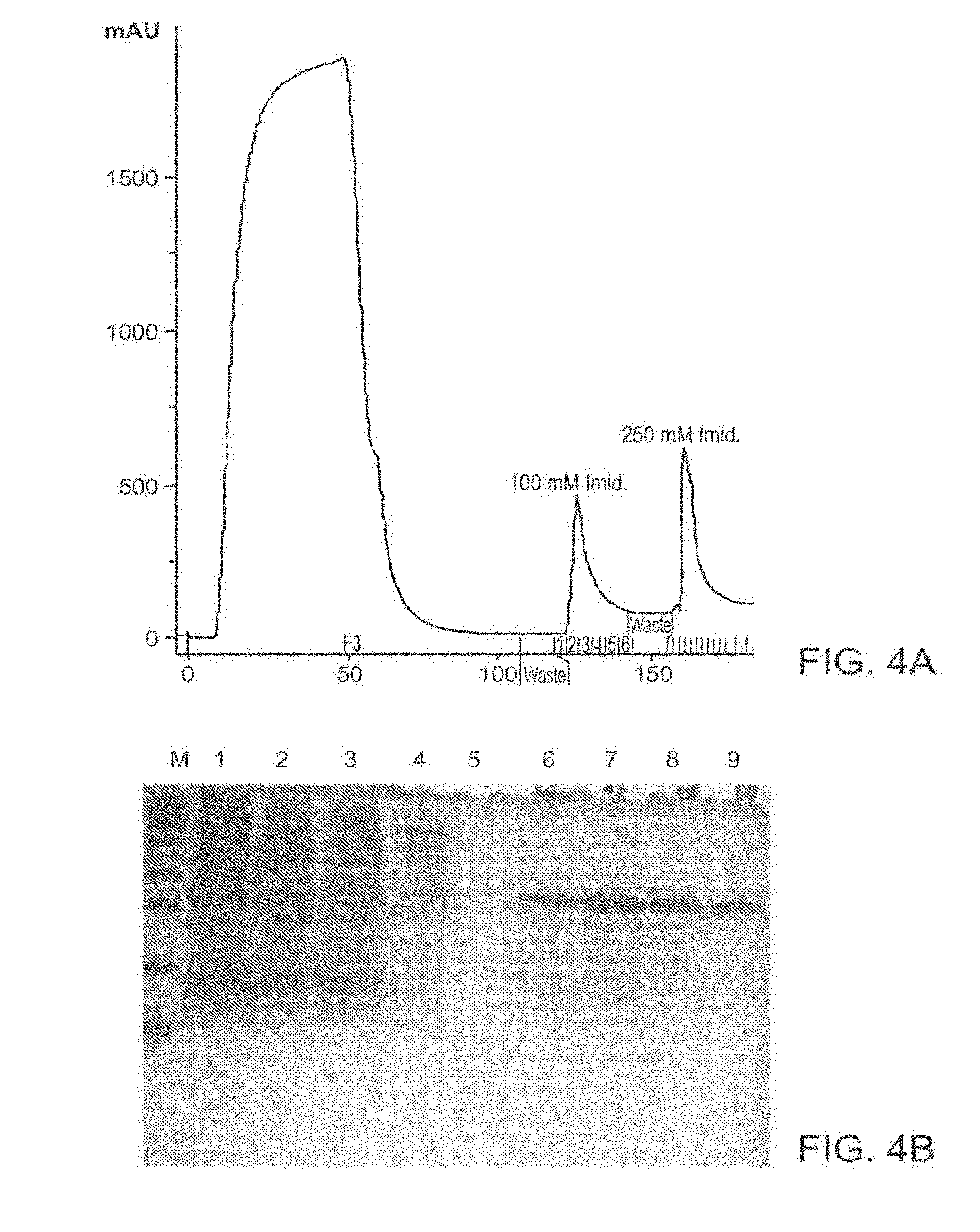
[0095] FIG. 4A-4D: Purification of TAT-MTSfra-FRA and of TAT-MTSorf-FRA fusion proteins
[0096] FIG. 4 presents an image of an affinity chromatography purification profile of the fusion protein TAT-MTSfra-FRA (FIG. 4A) and an image of SDS-PAGE analysis of the purification steps obtained for TAT-MTSfra-FRA is presented in FIG. 4B. An image of an affinity chromatography purification profile of the fusion protein TAT-MTSorf-FRA is presented in FIG. 4C and an image of SDS-PAGE analysis of the purification steps obtained for TAT-MTSorf-FRA is presented in FIG. 4D.
[0097] Abbreviations: M, marker; 1, whole cell extract; 2, pre-run fraction (the soluble sub-fraction of bacterial cells expressing the fusion protein); 3, flow through; 4, elution with 100 mM imidazole; and 5-9, elution with 250 mM imidazole.
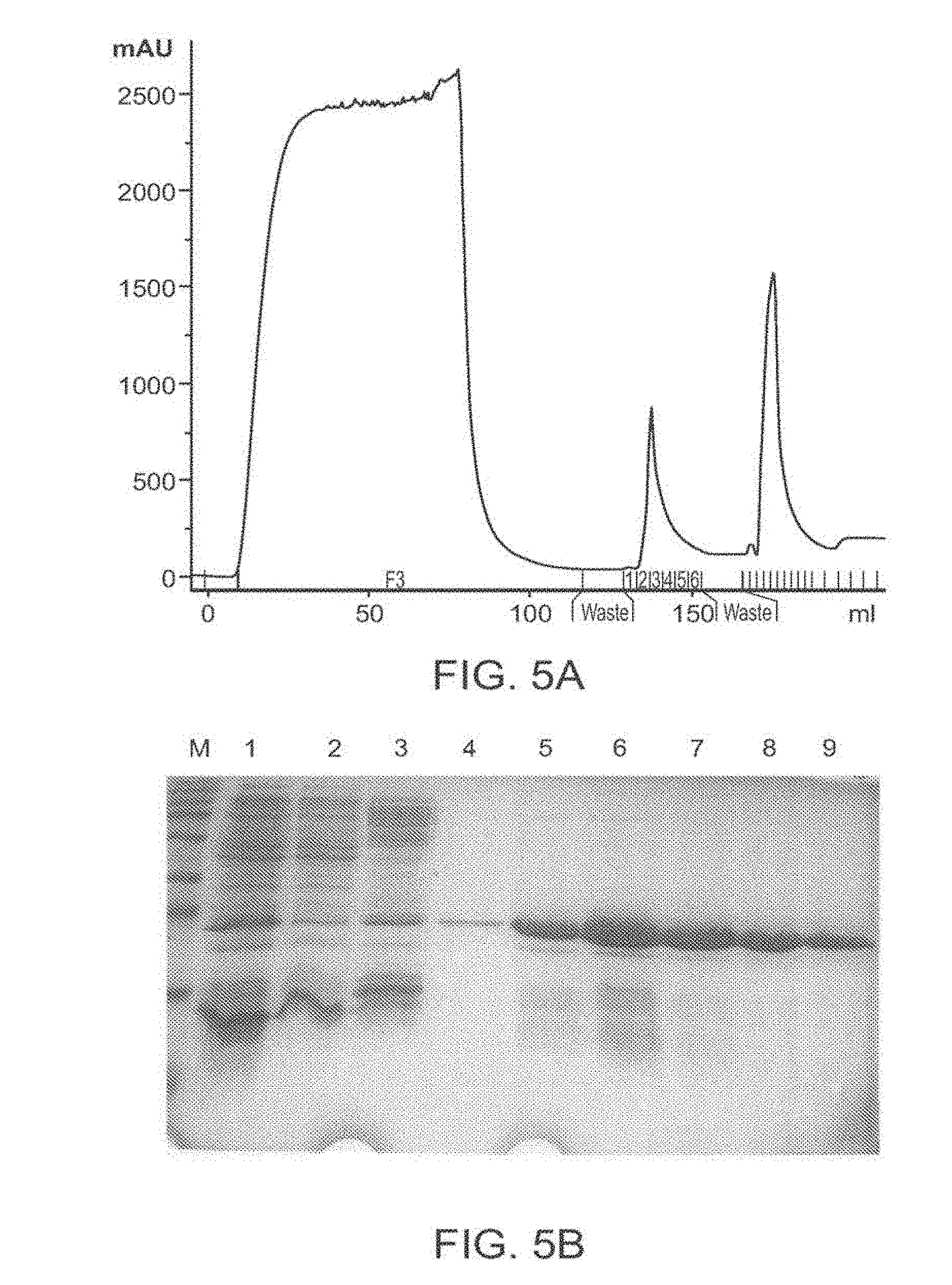
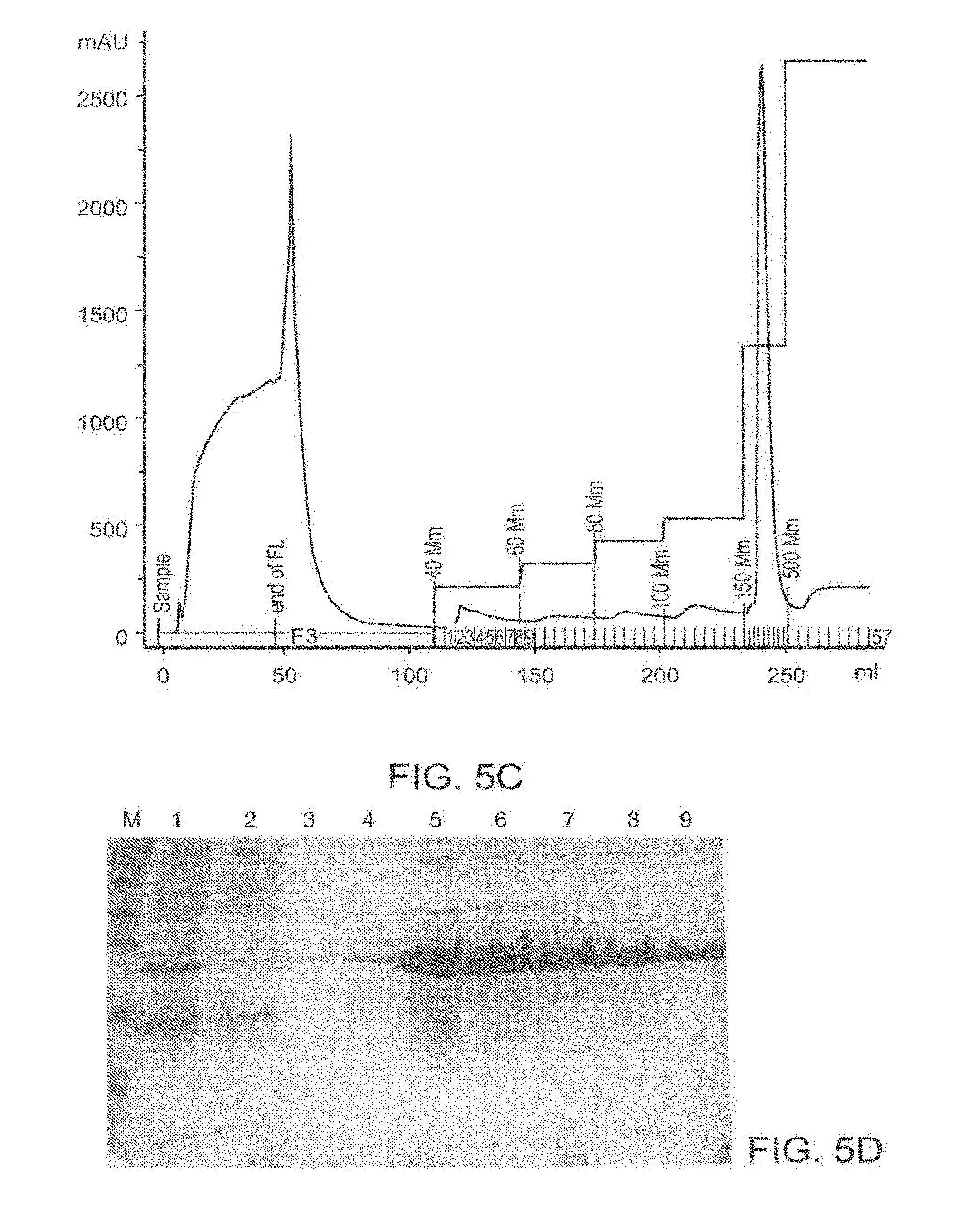
[0098] FIG. 5A-5D: Purification of TAT-MTSlad-FRA and of TAT-MTScs-FRA fusion proteins FIG. 5 presents an image of an affinity chromatography purification profile of the fusion protein TAT-MTSlad-FRA (FIG. 5A) and an image of SDS-PAGE analysis of the purification steps obtained for TAT-MTSlad-FRA is presented in FIG. 5B. An image of an affinity chromatography purification profile of the fusion protein TAT-MTScs-FRA is presented in FIG. 5C and an image of SDS-PAGE analysis of the purification steps obtained for TAT-MTScs-FRA is presented in FIG. 5D.
[0099] Abbreviations: M, marker; 1, whole cell extract; 2, pre-run fraction (the soluble sub-fraction of bacterial cells expressing the fusion protein); 3, flow through; 4, elution with 100 mM imidazole; and 5-9, elution with 250 mM imidazole.
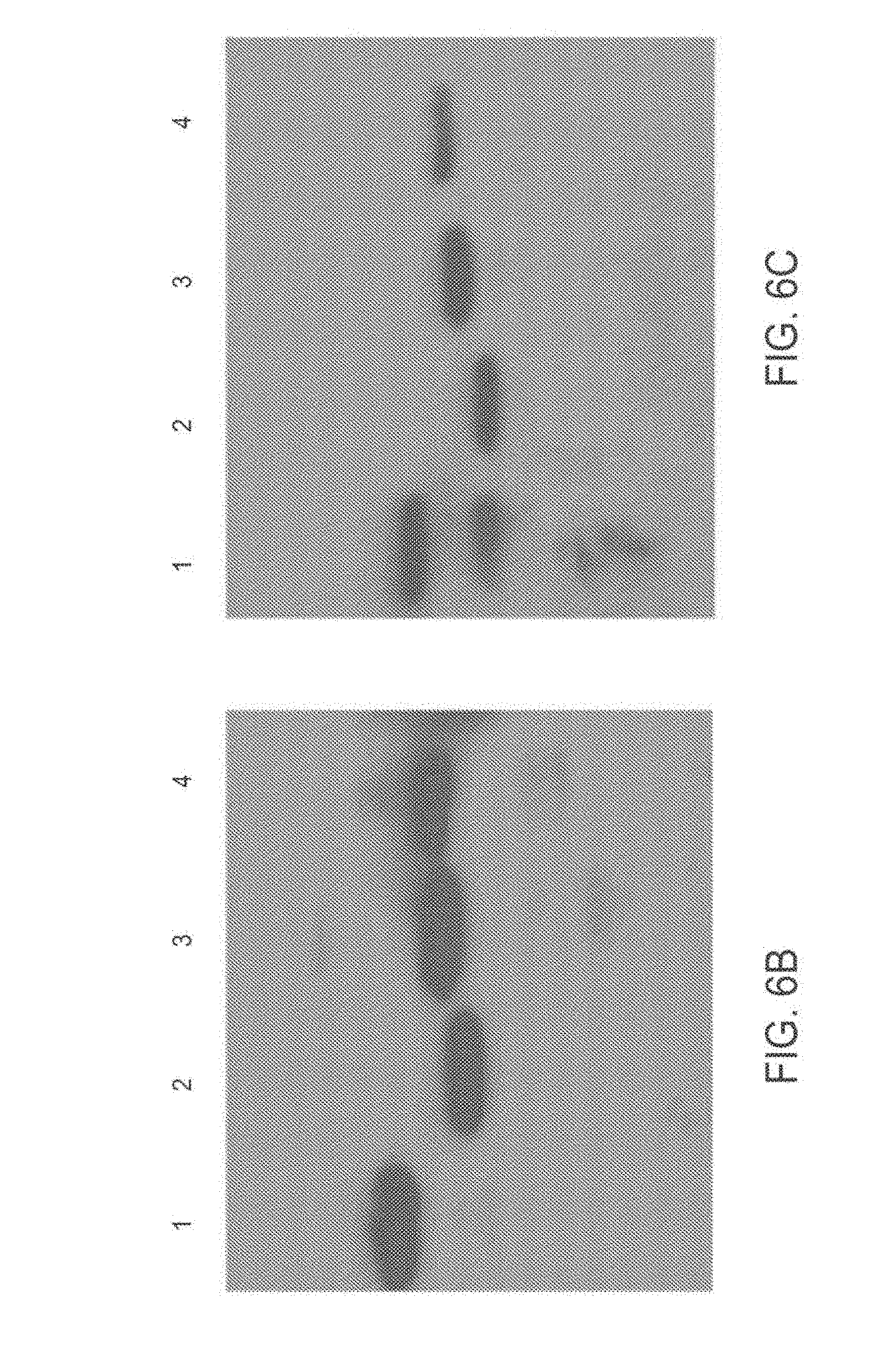
[0100] FIG. 6A-6C: Characterization of TAT-MTS-FRA highly purified fusion proteins The four highly purified TAT-MTS-FRA fusion proteins were characterized by a SDS-PAGE gel (FIG. 6A) and by Western blot analyses using anti-His (FIG. 6B) or anti-frataxin (FIG. 6C) antibodies.
[0101] Abbreviations: 1, TAT-MTSfra-FRA; 2, TAT-MTScs-FRA; 3, TAT-TSlad-FRA; 4, TAT-MTSorf-FRA; and M, Marker.
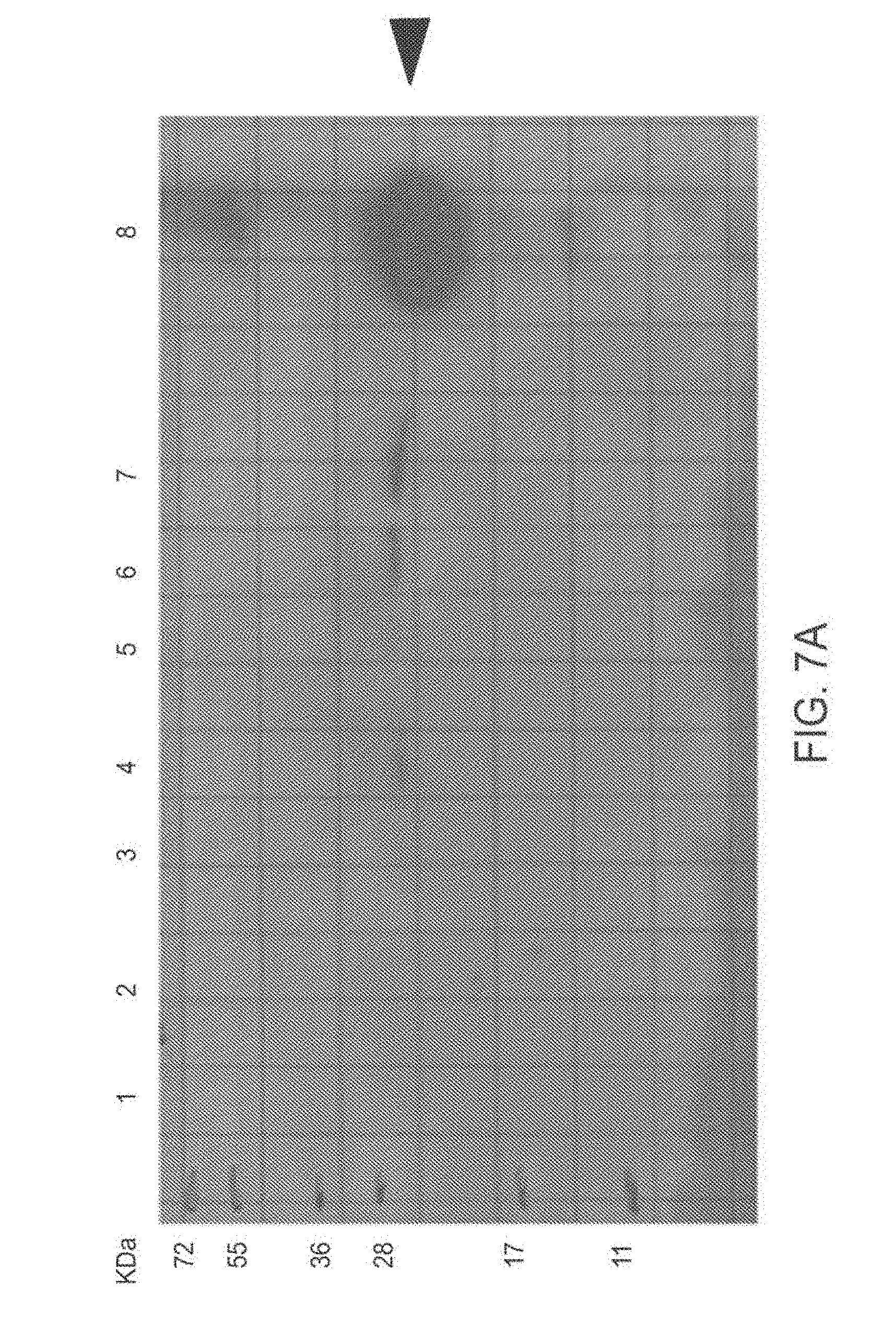
[0102] FIG. 7A-7B: Internalization of TAT-MTSlad-FRA into cells and their mitochondria FIG. 7A presents an immunoblot of a Western blot analysis using anti-His antibodies and FIG. 7B presents an immunoblot of a Western blot analysis using anti frataxin antibodies performed with BJAB cells incubated in the absence (lanes 1 & 2) or in the presence of TAT-MTSlad-FRA (lanes 3-7). At the end of the incubation period, sub-fractionation was performed, obtaining the cytoplasmic and mitochondrial fractions. Fractions were separated by SDS-PAGE and subjected to Western blot analysis.
[0103] Abbreviations: control, untreated cells: cytoplasm (1), mitochondria (2); cells treated for 1 hr with the fusion protein: cytoplasm (3), mitochondria (4); cells treated for 5 hr: cytoplasm (5), mitochondria (6, 7; from two separate experiments); highly purified TAT-MTSlad-FRA fusion protein as a positive control is shown in (8). Arrow-heads indicate the fusion protein (or its processing products, indicated by the lower arrow-head in FIG. 7B).
[0104] FIG. 8A-8B: Internalization of TAT-MTS-FRA fusion proteins into mitochondria of cells FIG. 8A presents a Western blot analysis using anti-frataxin antibodies of cells incubated for 3 hours with TAT-MTS-FRA fusion proteins, each fusion protein at a final concentration of 0.02 .mu.g/.mu.l. The cells were washed and their mitochondria were isolated.
[0105] FIG. 8B presents a Western blot analysis using anti-E1.alpha. antibodies of cells as detailed above. Abbreviations: M, marker; mitochondria isolated from control cells without any treatment (1), cells incubated with TAT-MTSfra-FRA (2), cells incubated with TAT-MTScs-FRA (3), cells incubated with TAT-MTSlad-FRA (4), cells incubated with TAT-MTSorf-FRA (5), purified TAT-MTSfra-FRA fusion protein (6), and purified TAT-MTScs-FRA fusion protein (7).
[0106] FIG. 9: Internalization of TAT-MTScs-FRA to fibroblasts of FRA patients
[0107] An immunoblot of a Western blot analysis using an anti-Frataxin antibody performed for mitochondrial (M) and cytosolic (C) fractions of fibroblasts (F816) obtained from Friedreich's ataxia patients that were incubated in the presence of 20 .mu.g/ml TAT-MTScs-FRA (or vehicle) for 2, 6 and 48 hours, with fresh addition of TAT-MTScs-FRA (at 20 .mu.g/ml) after 24 hours. TAT-MTScs-FRA and processed Frataxin are marked with arrows. Abbreviations: kDa, kilodalton; TAT, transactivator of transcription; MTS, mitochondria targeting sequence; CS, citrate synthase; FXN, frataxin; Std, standard.
[0108] FIG. 10: Aconitase activity in fibroblasts obtained from Friedreich's ataxia patients following 48 hours incubation with TAT-MTScs-FRA
[0109] A bar graph showing aconitase activity (mOD/min) of mitochondrial fractions obtained from Friedreich's ataxia patients' fibroblasts (F816) incubated for 48 hours with either 20 .mu.g/ml TAT-MTScs-FRA protein or vehicle (FXN or VEH, respectively). HepG2 whole cells homogenate served as positive control (POS.CON). Abbreviations: kDa, kilodalton; TAT, transactivator of transcription; MTS, mitochondria targeting sequence; CS, citrate synthase; FXN, frataxin; Std, standard; VEH, vehicle, POS. CON., positive control.
[0110] FIG. 11A-11B: TAT-MTS-FRA fusion proteins partially rescue cells from BSO-induced oxidative stress
[0111] FIG. 11A presents a bar diagram showing percentage of cell death induced by L-Buthionine-sulfoximine (BSO). Normal lymphocytes or lymphocytes obtained from Friedreich's ataxia (FRDA) patients (Lym 43) were seeded, incubated for 5 hr with the various TAT-MTS-FRA fusion proteins, after which BSO at different concentrations was added for additional 48 hr. At the end of the incubation time, cell cultures were subjected to cell proliferation assays.
[0112] FIG. 11B presents a bar diagram showing percentage of cell death of Lym 43 induced by L-Buthionine-sulfoximine (BSO) correlated with caspase 3 activity within the cells, assessed using the Apo-ONE Homogeneous Caspase 3/7 Assay Kit (Promega). Experiments were carried in parallel with cell viability assays.
[0113] Abbreviations: %, percent; YC, normal lymphocytes; FRA lym, lymphocytes obtained from Friedreich's ataxia patients; HTFRA=TAT-MTSfra-FRA, HT(LAD)FRA=TAT-MTSlad-FRA, HT(ORF)FRA=TAT-MTSorf-FRA, HT(CS)FRA=TAT-MTScs-FRA; BSO, L-Buthionine-sulfoximine; and PBS, Phosphate buffered saline.
[0114] FIG. 12A-12D: TAT-MTS-FRA fusion proteins partially rescue fibroblasts obtained from patients from BSO-induced oxidative stress, a comparison
[0115] FIG. 12A to FIG. 12D present bar diagrams of percentage of cell death induced by BSO in fibroblasts obtained from FRDA patients (Fib. 78). Cells were seeded, incubated for 24 hr with the TAT-MTSfra-FRA (FIG. 12A), TAT-MTScs-FRA (FIG. 12B), TAT-MTSlad-FRA (FIG. 12C) and with the TAT-MTSorf-FRA (FIG. 12D) fusion protein, after which BSO at different concentrations was added for additional 48 hr. At the end of the incubation time, cell cultures were subjected to cell proliferation assays. Abbreviations: %, percent; PBS, Phosphate buffered saline; BSO, L-Buthionine-sulfoximine; HTF, TAT-MTSfra-FRA; MTS (cs), TAT-MTScs-FRA; MTS (LAD), TAT-MTSlad-FRA; MTS (ORF), TAT-MTSorf-FRA.
[0116] FIG. 13A-13B: TAT-MTSorf-FRA rescues lymphocytes and fibroblasts obtained from Friedreich's ataxia patients from BSO-induced oxidative stress
[0117] FIG. 13A presents a bar diagram of percentage of cell death induced by BSO in fibroblasts obtained from FRDA patients (Fib. 78) and FIG. 13B presents a bar diagram of percentage of cell death induced by BSO in lymphocytes obtained from patients (Lym 43). Cells were seeded, incubated for 24 hr with the TAT-MTSorf-FRA fusion protein, after which BSO at different concentrations was added for additional 48 hr. At the of the incubation time, cell cultures were subjected to cell proliferation assays. Abbreviations: PBS, Phosphate buffered saline; %, percent; BSO, L-Buthionine-sulfoximine; and ORF, C6ORF66.
[0118] FIG. 14A-14B: TAT-MTSlad-FRA rescues fibroblasts obtained from patients from BSO-induced oxidative stress
[0119] FIG. 14A and FIG. 14B present bar diagrams of percentage of cell death induced by BSO in fibroblasts obtained from FRDA patients (Fib. 78) of two representative experiments. Cells were seeded, incubated for 24 hr with the TAT-MTSlad-FRA fusion protein, after which BSO at different concentrations was added for additional 48 hr. At the of the incubation time, cell cultures were subjected to cell proliferation assays. Abbreviations: %, percent; PBS, Phosphate buffered saline; BSO, L-Buthionine-sulfoximine; LAD, lipoamide dehydrogenase.
[0120] FIG. 15A-15D: TAT-MTS-FRA fusion proteins constructs
[0121] FIG. 15A to FIG. 15D are schematic presentations of TAT-MTS-FRA fusion protein constructs comprising a HIV-1 transactivator of transcription (TAT) domain (boxed) fused to a GSDP linker (colored in grey) fused to a human mitochondria targeting sequence (MTS) of a human mitochondrial protein (double-underlined) selected from frataxin (MTSfra, FIG. 15A), citrate synthase (MTScs, FIG. 15B), lipoamide dehydrogenase (MTSlad, FIG. 15C) and C6ORF66 (MTSorf, FIG. 15D) fused to human frataxin (underlined).
[0122] FIG. 16A-16E: Expression and purification of TAT-MTS-OTC protein constructs
[0123] FIG. 16A to FIG. 16D are images of sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS PAGE) analysis of bacterially expressed TAT-MTS-OTC fusion protein constructs comprising ornithine transcarbamoylase (OTC, FIG. 16A), citrate synthase (CS, FIG. 16B), C6ORF66 (ORF, FIG. 16C) or lipoamide dehydrogenase (LAD FIG. 16D) as their MTS. The fusion protein construct is indicated with arrows. About 2 .mu.g of each protein and BSA were analyzed on 4-20% gels, followed by Coomassie blue staining. Marker sizes are presented in FIG. 16E. Abbreviations: M1, protein marker; kDa, kilodalton.
[0124] FIG. 17A-17B: Internalization of TAT-MTS-OTC protein constructs into mitochondria FIG. 17A is an image of a Western blot analysis performed for mitochondrial fractions of HepG2 cells incubated with 12 .mu.g/ml of the TAT-MTS-OTC fusion protein TAT-MTSotc-OTC, TAT-MTScs-OTC, TAT-MTSlad-OTC or vehicle for 20 min, 1 and 3 hours in either Eagle's Minimum Essential Medium (EMEM) alone or EMEM supplemented with DMSO (D), Trehalose (T), or Ornithine (O), as indicated. The protein standards TAT-MTSlad-OTC or TAT-MTSorf-OTC (Std.) were run in parallel, sizes of the protein markers (M) are presented in FIG. 17B. Migration of the OTC fusion protein is marked by an arrow. Abbreviations: EMEM, Eagle's Minimum Essential Medium; TAT, transactivator of transcription; MTS, mitochondria targeting sequence; OTC, ornithine transcarbamoylase; CS, citrate synthase; LAD, lipoamide dehydrogenase; Std., standard; T/D, Trehalose/DMSO; T/D/O, Trehalose/DMSO/Ornithine; M, marker.
[0125] FIG. 18A-18B: Western blot analysis of cytosolic fractions of HepG2 cells incubated with TAT-MTS-OTC protein constructs
[0126] FIG. 18A and FIG. 18B are images of Western blot analyses performed for cytosolic fractions of HepG2 cells incubated with 12 .mu.g/ml of the TAT-MTS-OTC fusion protein TAT-MTSotc-OTC, TAT-MTScs-OTC, TAT-MTSlad-OTC or vehicle for 20 min, 1 and 3 hours in either EMEM alone or EMEM supplemented with DMSO (D), Trehalose (T), or Ornithine (0), as indicated. The protein standards TAT-MTScs-OTC or TAT-MTSorf-OTC (Std.) were run in parallel. Abbreviations: OTC, ornithine transcarbamoylase; CS, citrate synthase; LAD, lipoamide dehydrogenase; ORF, C6ORF66; EMEM, Eagle's Minimum Essential Medium; T/D, Trehalose/DMSO; T/D/O, Trehalose/DMSO/Ornithine; M, marker.
[0127] FIG. 19: In vitro enzymatic activity of OTC
[0128] A bar graph showing the level of in vitro enzymatic activities of the OTC protein constructs TAT-MTScs-OTC and TAT-MTSotc-OTC measured by detecting the net absorbance of citrulline at an optical density (O.D.) of 492 nm compared to a control. Abbreviations: TAT, transactivator of transcription; MTS, mitochondria targeting sequence; OTC, ornithine transcarbamoylase; CS, citrate synthase.
[0129] FIG. 20: Rescue from ammonia stress by OTC fusion proteins
[0130] A bar graph showing cell viability of HepG2 cells suffering from ammonia stress in the presence of 14 .mu.g/ml of the OTC protein constructs TAT-MTScs-OTC, and TAT-MTSotc-OTC, compared to vehicle-treated and untreated cells. Florescence signal was calculated as the ratio between Ammonium chloride treated and non-treated cells. Abbreviations: TAT, transactivator of transcription; MTS, mitochondria targeting sequence; CS, citrate synthase; OTC, ornithine transcarbamoylase.
DETAILED DESCRIPTION OF EMBODIMENTS
[0131] The presently disclosed subject matter relates to the preparation of various plasmid constructs encoding TAT-MTS-Frataxin fusion proteins, providing a basis of a wide-range therapeutic tool for delivering mitochondrial proteins into mitochondria.
[0132] The protein constructs described herein, comprising a mitochondrial protein as well as TAT and a specific mitochondrial targeting sequence (MTS), enabling the mitochondrial protein to cross both cellular and mitochondrial membranes, were expressed and purified and their biological activity was verified. Remarkably, the protein yield obtained for fusion protein constructs comprising an MTS which was other than the native MTS of the mitochondrial protein present in the fusion construct (e.g., a frataxin fusion protein construct with MTS heterologous to frataxin), was superior to the yield obtained for a frataxin fusion protein construct comprising the native MTS of frataxin.
[0133] As demonstrated below, the inventors show that various fusion proteins are able to enter the mitochondria within intact cells. In addition, the inventors show that the fusion proteins exhibit biological activity. For example, the various TAT-MTS-FRA fusion proteins were shown to rescue cells obtained from Friedreich ataxia patients as well as normal cells from oxidative stress, as demonstrated in the Examples below. Surprisingly, a superior protective effect was observed for fusion proteins carrying an MTS, which was other than the native MTS of the functional mitochondrial protein present in the fusion construct (i.e. a heterologous MTS) as compared to the effect demonstrated by the fusion protein constructs carrying the native MTS.
[0134] The presently disclosed subject matter provides fusion protein constructs comprising heterologous MTSs of human nuclear-encoded mitochondrial proteins that are classical MTS sequences, which are known to be removed upon entry to the mitochondria. By a non-limiting example, a delivery system as herein described comprising frataxin may be used for the treatment or alleviation of Friedreich's ataxia or any other disorder associated with a deficiency of frataxin or defective frataxin.
[0135] Thus, the presently disclosed subject matter provides a fusion protein comprising a HIV-1 transactivator of transcription (TAT) domain, a functional human mitochondrial protein and a human mitochondria targeting sequence (MTS) situated between said TAT domain and said functional human mitochondrial protein and wherein said human MTS is heterologous to said functional human mitochondrial protein.
[0136] The term "functional human mitochondrial protein" as used herein refers to any protein which is essential for a biological activity of mitochondria. A functional human mitochondrial protein may be a protein, which is active when present in the mitochondria by itself (per se) or a protein that when present in the mitochondria functions as a component of a mitochondrial multi-component complex (i.e. with other enzymes, co-factors, or proteins). Typically, a functional human mitochondrial protein is a protein, which, when absent, deficient or mutated, causes a mitochondrial disorder or is associated with a mitochondrial disorder.
[0137] In some specific embodiments, the functional mitochondrial protein refers to the full-length amino acid sequence of the protein. In other embodiments, the functional mitochondrial protein is a fragment of the full-length amino acid sequence, sufficient to provide the mitochondrial protein activity, either alone or as part of a multi-component complex, as appropriate.
[0138] In further embodiments, the functional human mitochondrial protein is a mutated derivative of said protein, wherein one or more of the native amino acid residues has been deleted, replaced or modified while still maintaining the mitochondrial functionally of the protein (alone or as part of a multi-component complex).
[0139] In the above and other embodiments, the functional human mitochondrial protein (also denoted "mature" protein) refers to a protein devoid of its mitochondrial targeting sequence (MTS). In other words, the fusion protein construct herein provided comprises a functional mitochondrial protein, which, upon entry to the mitochondria is cleaved off from the fusion protein construct in its mature, active (functional) state.
[0140] By way of non-limiting example, in the above and other embodiments of the disclosed subject matter, the functional human mitochondrial protein whose activity is supplied by a fusion protein of the present invention may be any one of human frataxin (the mature protein having the amino acid sequence denoted by SEQ ID NO. 26 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 6), ornithine transcarbamoylase (OTC, the mature protein having the amino acid sequence denoted by SEQ ID NO. 39 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 15), human Lipoamide Dehydrogenase (LAD), 2-oxoisovalerate dehydrogenase alpha subunit (Branched-Chain Keto Acid Dehydrogenase E1a) (NCBI Protein Database Accession No. P12694; OMIM:248600), 2-oxoisovalerate dehydrogenase beta subunit (Branched-Chain Keto Acid Dehydrogenase E1.beta.; P21953), Acyl-CoA dehydrogenase, medium-chain specific (P1 1310; OMIM:201450), Acyl-CoA dehydrogenase, very-long-chain specific (P49748; OMIM:201475), Trifunctional enzyme alpha subunit (Long-chain 3 hydroxyacyl CoA Dehydrogenase or LCHAD) (P40939; OMIM:609015) (HADHA), Trifunctional enzyme beta subunit (Hydroxyacyl-CoA Dehydrogenase/3-Ketoacyl-CoA Thiolase/Enoyl-CoA Hydratase (P55084) (HADHB)), Pyruvate dehydrogenase E1 component beta subunit (P1 1177; OMIM:208800), and Pyruvate dehydrogenase E1 component alpha subunit (P08559; OMIM:312170).
[0141] In some embodiments, the human mitochondrial protein is a functional mitochondrial protein per se and/or is a component of a mitochondrial multi-component complex.
[0142] As indicated above, the functional human mitochondrial protein of the disclosed subject matter may be a protein which is active when present in the mitochondria by itself (i.e. the protein per se is active) or a protein that when present in the mitochondria functions as a component of a mitochondrial multi-component complex (i.e. with other enzymes, co-factors, or proteins). The term "mitochondrial multi-component complex" as used herein refers to an enzyme that forms a complex with other enzymes or proteins that is essential for a biological activity of mitochondria.
[0143] As shown in the Examples below (FIG. 7), the fusion protein comprising a TAT and MTS sequences is cleaved upon entry into the mitochondria, and a mature active protein is obtained. The protein constructs provided by the presently disclosed subject matter thus allow a human mitochondrial protein, which is first covalently attached to TAT and MTS domains, to cross both cellular and mitochondrial membranes, and once inside the mitochondria, be processed by mitochondrial peptidases while retaining their biological activity and proper conformation. The delivery system described herein thus enables a human mitochondrial protein to retain its mitochondrial function per se or the integration thereof in a mitochondrial multi-component complex.
[0144] The mitochondrial multi-component complex encompassed by the present disclosure refers to a group of at least two different proteins assembled together in a specific ratio that functions in a coordinated fashion to catalyze a series of reactions. The function of a mitochondrial multi-component complex is dependent on its structure; thus, the proteins that compose the complex must properly fold and physically fit together in the proper configuration in order to efficiently catalyze the series of reactions.
[0145] In all embodiments, the functional human mitochondrial protein according to presently disclosed subject matter is cleaved off from the fusion protein construct upon entry to the mitochondria and resides therein at its mature, properly-folded active state. In some embodiments, the functional human mitochondrial protein may readily then integrate into a conformationally-sensitive mitochondrial multi-component complex.
[0146] By way of non-limiting example, the presently disclosed subject matter encompasses a mitochondrial multi-component complex which is any one of pyruvate dehydrogenase complex (PDHC), .alpha.-ketoglutarate dehydrogenase complex (KGDHC), and branched-chain keto-acid dehydrogenase complex (BCKDHC), the complexes of the respiratory chain, and those involved in fatty acid .beta.-oxidation and the urea cycle. The complexes of the respiratory chain are complex I (NADH-ubiquinone oxidoreductase), complex II (succinate-ubiquinone oxidoreductase), complex III (ubiquinol-ferricytochrome C oxidoreductase), complex IV (Cytochrome C oxidoreductase), and complex V (FIFO ATPase) where each mitochondrial multicomponent complex represents a separate embodiment of the present invention.
[0147] As shown in Examples 1-3 below, the inventors have cloned, expressed and purified fusion protein constructs comprising the protein frataxin.
[0148] The mitochondrial protein human frataxin (FXN) is an essential and highly conserved protein expressed in most eukaryotic organisms that appears to function in mitochondrial iron homeostasis, notably the de novo biosynthesis of iron-sulfur (Fe--S) cluster proteins and heme biosynthesis. The exact function of FXN has not been defined but recent studies suggest that FXN functions as an allosteric activator with Fe.sup.2+ for Fe--S cluster biosynthesis. The absence of FXN is associated with a loss of activity in Fe--S-containing proteins, such as aconitase as well as with the disease Friedreich ataxia.
[0149] Precursor FXN protein (23.1 kDa, 210 amino acids) comprises an 80 amino acid mitochondrial targeting sequence (MTS) at its amino (N) terminus. Within mitochondria, the precursor FXN protein is processed in two steps by the mitochondrial matrix processing peptidase (MPP). It has been shown that the intermediate form of FXN is formed by cleavage at residue 42 by the MPP, and the resulting form of FXN (FXN42-210) has been shown to be cleaved at amino acid 81, yielding a mature, 130 amino acid protein, with a predicted molecular weight of 14.2 kDa.
[0150] As described above, Friedreich ataxia is an autosomal recessive degenerative disorder characterized by ataxia, areflexia, sensory loss, weakness, scoliosis, and cardiomyopathy. A deficiency of frataxin in cells leads to decreased activities of mitochondrial iron-sulfur cluster-containing enzymes, to an accumulation of iron in the mitochondrial matrix, increased sensitivity to oxidative stress, as well as to impaired adenosine triphosphate (ATP) production.
[0151] In the above and other embodiments of the presently disclosed subject matter, frataxin refers to human frataxin and any biologically active fragments and derivatives thereof, which is devoid of its natural (native) MTS sequence. Non limiting examples for mature human frataxin are given by the accession number Q16595[81-210] and as indicated in Table 1 below, where the amino acid sequence of mature human frataxin is as set forth in SEQ ID NO. 26 and the nucleic acid sequence encoding therefor is as set forth in SEQ ID NO. 6.
[0152] Notably, as shown in Example 4 below, a TAT-MTS-frataxin fusion protein was demonstrated by the inventors to enter mitochondria of human intact BJAB cells. Analysis of sub-cellular fractions of these cells, in order to separate the mitochondria from the cytosol, verified that the various frataxin fusion protein constructs (i.e. TAT-MTSlad-FRA, TAT-MTSfra-FRA, TAT-MTScs-FRA, and TAT-MTSorf-FRA) were indeed successfully delivered into the mitochondria. Surprisingly, among the fusion proteins carrying a heterologous MTS, the MTS of citrate synthase (MTScs) was shown by the inventors to be delivered into the mitochondria in the most efficient manner.
[0153] Delivery of fusion proteins comprising a mitochondrial protein into the mitochondria has far-reaching therapeutic beneficial implications for treatment of mitochondrial disorders in general, and delivery of frataxin into mitochondria has specific therapeutic benefit for treatment of Friedreich's ataxia in particular.
[0154] As noted above, ornithine transcarbamoylase (OTC, also called ornithine carbamoyltransferase) is also encompassed by the presently disclosed subject matter. OTC is a protein having enzymatic activity that catalyzes the reaction between carbamoyl phosphate (CP) and ornithine (Orn) to form citrulline (Cit) and phosphate (Pt). In mammals OTC is located in the mitochondria and is part of the urea cycle.
[0155] OTC is a trimer, and the active sites thereof are located at the interface between the protein monomers, emphasizing the importance of proper folding to the mitochondrial activity of the protein. Deficiency in OTC results in an increase in ammonia level, leading to neurological problems.
[0156] As demonstrated in the appended Examples, fusion protein constructs according to the invention comprising OTC and its native MTS as well as fusion protein constructs comprising OTC and a heterologous MTS were able to internalize into the mitochondria in HepG2 cells (Example 9). Interestingly, the internalization ability of the fusion protein constructs comprising an MTS that is heterologous to OTC was slightly higher than the internalization ability of the fusion protein construct comprising the native MTS of OTC. Fusion protein constructs comprising the protein OTC were also shown to be active. As shown in FIG. 20, the level of cell viability in the presence of fusion protein constructs comprising OTC and citrate synthase as the MTS was similar to the level of cell viability for cells which were not exposed to ammonium chloride (which served as a model for ammonia stress conferred by defective or missing OTC).
[0157] Furthermore, the level of cell viability in the presence of a fusion protein construct comprising OTC and citrate synthase as the MTS was higher from the level of cell viability in cells treated with the fusion protein construct comprising the native MTS of OTC.
[0158] Thus, in the above and other embodiments of the disclosed subject matter, the functional human mitochondrial protein is specifically any one of frataxin and ornithine transcarbamoylase (OTC). In some embodiments, the nucleic acid encoding the mature OTC protein is denoted by SEQ ID NO. 15. In other embodiments the OTC mature protein of the present disclosure has the amino acid sequence denoted by SEQ ID NO. 39.
[0159] As indicated above, the presently disclosed subject matter provides a fusion protein comprising a HIV-1 transactivator of transcription (TAT) domain, a functional human mitochondrial protein and a human mitochondria targeting sequence (MTS) situated between said TAT domain and said functional mitochondrial protein and wherein said human MTS is heterologous to said functional protein.
[0160] Most of the proteins directed to the mitochondria are synthesized with a mitochondrial targeting (or translocation) sequence (MTS), which allows their import from the cytoplasm into mitochondria through the translocation machinery. Once entering the mitochondria, the MTS is recognized and cleaved off, allowing for proper processing and, if necessary, assembly into mitochondrial enzymatic complexes.
[0161] Thus, as used herein, the term "mitochondria targeting sequence", MTS or "mitochondria translocation sequence" refers to an amino acid sequence capable of causing the transport into the mitochondria of a protein, peptide, amino acid sequence, or compound attached thereto, and any biologically active fragments thereof. MTSs used in the fusion protein constructs in accordance with the presently disclosed subject matter, which are situated N-terminal to the functional mitochondrial protein, are typically from about 15 to about 40 amino acids in length, including from about 3 to about 5 nonconsecutive basic amino acid (arginine/lysine) residues, often with several serine/threonine residues but without acidic amino acid (asparate/glutamate) residues. In their molecular structure, these MTS s are able to form strong basic amphipathic .alpha.-helices that are essential for efficient mitochondrial transportation.
[0162] In other words, presently disclosed is a fusion protein as herein defined, wherein said functional human mitochondrial protein is C-terminal to said human mitochondria targeting sequence (MTS).
[0163] The term "about" as used herein indicates values that may deviate up to 1%, more specifically 5%, more specifically 10%, more specifically 15%, and in some cases up to 20% higher or lower than the value referred to, the deviation range including integer values, and, if applicable, non-integer values as well, constituting a continuous range.
[0164] In the above and other embodiments, the MTS is human MTS, namely MTS of a human mitochondrial protein.
[0165] In the above and other embodiments of the presently disclosed subject matter the MTS comprises from about 15 to about 40 amino acid residues, including from about 3 to about 5 nonconsecutive (i.e. which are not covalently linked one to the other in a sequential manner) basic amino acid residues, and optionally from about 1 to about 3 or 4 or 5 serine/threonine residues.
[0166] The term "amino acid residues" as used herein refers to naturally occurring and synthetic amino acids, as well as amino acid analogs and amino acid mimetics that can function in a manner similar to the naturally occurring amino acids. Naturally occurring amino acids are those encoded by the genetic code, as well as those amino acids that are later modified, e.g., hydroxyproline, .gamma.-carboxyglutamate, and O-phosphoserine. "amino acid analogs and amino acid mimetics" refers to compounds that have the same fundamental chemical structure as a naturally occurring amino acid. Such analogs have modified R groups or modified peptide backbones, but retain the same basic chemical structure as a naturally occurring amino acid. Amino acids may be referred to herein by either their commonly known three letter symbols or by the one-letter symbols recommended by the IUPAC-IUB Biochemical Nomenclature Commission.
[0167] It is well known in the art that amino acid residues may be divided according to their chemical properties to various groups, inter alia, on the basis of similarity in polarity, charge, solubility, hydrophobicity, hydrophilicity, and/or the amphipathic nature of the residues involved.
[0168] For example, nonpolar "hydrophobic" amino acids are selected from the group consisting of Valine (V), Isoleucine (I), Leucine (L), Methionine (M), Phenylalanine (F), Tryptophan (W), Cysteine (C), Alanine (A), Tyrosine (Y), Histidine (H), Threonine (T), Serine (S), Proline (P), Glycine (G), Arginine (R) and Lysine (K); "polar" amino acids are selected from the group consisting of Arginine (R), Lysine (K), Aspartic acid (D), Glutamic acid (E), Asparagine (N), Glutamine (Q); "positively charged" amino acids are selected form the group consisting of Arginine (R), Lysine (K) and Histidine (H) and wherein "acidic" amino acids are selected from the group consisting of Aspartic acid (D), Asparagine (N), Glutamic acid (E) and Glutamine (Q). "Basic" amino acids are selected from the group consisting of Histidine (H), lysine (K) and Arginine (R), which are polar and positively charged at pH values below their pKa's, and are very hydrophilic.
[0169] As indicated above, the presently disclosed subject matter encompasses human mature frataxin and any biologically active fragments and derivatives thereof, which is devoid of its natural (native) MTS sequence. By the term "biologically active fragments and derivatives" it is meant any variations, including deletion, substitution and/or insertion of one or more amino acid residues in the amino acid sequences of mature frataxin (or in the nucleic acid encoding therefor), for example 1, 2, 3, 4, 5 or more amino acid residues, in accordance with the presently disclosed subject matter which would not alter the biological activity of frataxin.
[0170] The invention further relates to DNA constructs comprising the nucleic acid sequence of the presently disclosed subject matter or biologically functional fragments and derivatives thereof. The constructs of the presently disclosed subject matter may further comprise additional elements such as promoters, regulatory and control elements, translation, expression and other signals, operably linked to the nucleic acid sequence of the invention.
[0171] It is known that each mitochondrial enzyme produced in the cytoplasm and transported into the mitochondria is produced as a precursor protein, carrying its natural MTS. Thus, the precursor mitochondrial protein already has its native MTS. However, this naturally occurring sequence in the precursor protein may be exchanged with any other known MTS.
[0172] As exemplified herein, the fusion protein constructs comprising frataxin or ornithine transcarbamoylase prepared by the inventors further comprised the MTS of lipoamide dehydrogenase (MTSlad, of the amino acid sequence denoted by SEQ ID NO. 24 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 5), C6ORF66 (MTSorf, of the amino acid sequence denoted by SEQ ID NO. 25 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 4), or of citrate synthase (MTScs, of the amino acid sequence denoted by SEQ ID NO. 23 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 3).
[0173] Alternatively, fusion protein constructs comprising frataxin or ornithine transcarbamoylase prepared by the inventors comprised respectively the native MTS of frataxin (MTSfra, of the amino acid sequence denoted by SEQ ID NO. 22 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 2) or the native MTS of ornithine transcarbamoylase (MTSotc, of the amino acid sequence denoted by SEQ ID NO. 38 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 37).
[0174] In addition, the inventors showed that fusion protein constructs comprising frataxin and a MTS sequence that is not the native MTS of frataxin (i.e. heterologous MTS) were superior to the frataxin fusion protein construct comprising the native MTS, based on the higher yield obtained for fusion protein constructs comprising heterologous MTS during the expression and purification stages.
[0175] Surprisingly, fusion protein constructs comprising frataxin and a heterologous MTS were also demonstrated by the inventors to have an enhanced biological activity as compared to the frataxin fusion protein construct comprising the native MTS (Example 5). In particular, a fusion protein construct comprising frataxin and citrate synthase MTS showed the highest effect in reducing toxicity of BSO (FIG. 11A and FIG. 11B). As shown in Example 4 below, the fusion protein construct comprising frataxin and citrate synthase MTS also showed the highest ability of being delivered into mitochondria among the exemplified constructs comprising heterologous MTS.
[0176] Consistent with the above results, a fusion protein construct comprising frataxin and another heterologous MTS, namely the MTS of lipoamide dehydrogenase (LAD), was also demonstrated by the inventors to have an enhanced biological activity as compared to the frataxin fusion protein construct comprising the frataxin native MTS, as demonstrated in FIG. 12C. As evident from FIG. 12, the biological activity of this fusion protein construct was comparable to the biological activity of the fusion protein construct comprising the citrate synthase MTS.
[0177] As detailed above, it is known that FXN mRNA is translated into a precursor polypeptide that is transported to the mitochondrial matrix and processed to at least two forms, namely FXN42-210 and FXN81-210, where FXN42-210 is the transient processing intermediate and FXN81-210 represents the mature protein. In other words, the transient frataxin polypeptide FXN42-210 includes a portion of the native frataxin MTS, whereas the FXN81-210 is the mature protein per se, devoid of its native MTS. Without wishing to be bound by theory, by using a heterologous MTS in fusion protein constructs comprising frataxin, mature frataxin is expected to be released from the fusion protein at a single step, thereby raising the biological availability of this protein in the mitochondria compared to fusion protein constructs comprising frataxin and its native MTS.
[0178] Thus, the MTS encompassed by the presently disclosed subject matter is any human MTS that is encoded by the nuclear DNA, translated (produced) in the cytoplasm and transported into the mitochondria and which is not the native N-terminal MTS sequence of the functional protein present in the fusion protein construct according to the invention. In other words, the MTS sequence is other than the native N-terminal MTS sequence of the functional protein, i.e. is heterologous thereto. The various MTS may be exchangeable for each mitochondrial enzyme among themselves. Each possibility represents a separate embodiment of the present invention.
[0179] As used herein, the term "heterologous" when referring to MTS fused to the functional human mitochondrial protein according to the present disclosure, is to be taken to mean MTS obtained from another (distinct) mitochondrial protein, i.e. MTS which is not the naturally occurring MTS of the said functional protein.
[0180] By way of a non-binding example, the heterologous MTS according to the present disclosure is a MTS which is heterologous to the mitochondrial protein frataxin, or to the mitochondrial protein ornithine transcarbamoylase (OTC) as exemplified herein, which may be, but is not limited to, the MTS of any human mitochondrial protein, e.g. lipoamide dehydrogenase (MTSlad, of the amino acid sequence denoted by SEQ ID NO. 24 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 5), C6ORF66 (MTSorf, of the amino acid sequence denoted by SEQ ID NO. 25 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 4), and of citrate synthase (MTScs, of the amino acid sequence denoted by SEQ ID NO. 23 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 3), which are non-limiting examples.
[0181] Thus, in embodiments of the presently disclosed fusion protein constructs, the MTS can be any one of human mitochondrial citrate synthase MTS having the amino acid sequence denoted by SEQ ID NO. 23, the human lipoamide dehydrogenase MTS having the amino acid sequence denoted by SEQ ID NO. 24, the MTS of the human C6ORF66 gene product having the amino acid sequence denoted by SEQ ID NO. 25 and the MTS of human mitochondrial GLUD2 encoded by the nucleic acid sequence denoted by SEQ ID NO. 16.
[0182] In other embodiments of the presently disclosed fusion protein construct, the MTS is any one of human mitochondrial citrate synthase MTS having the amino acid sequence denoted by SEQ ID NO. 23 and the human lipoamide dehydrogenase MTS having the amino acid sequence denoted by SEQ ID NO. 24.
[0183] In some embodiments disclosed is a fusion protein comprising a HIV-1 transactivator of transcription (TAT) domain fused to human frataxin and a human mitochondria targeting sequence (MTS) of a human mitochondrial protein selected from lipoamide dehydrogenase (LAD) and citrate synthase (CS) situated between said TAT domain and said frataxin, wherein said frataxin is C-terminal to said MTS of human lipoamide dehydrogenase or human citrate synthase.
[0184] In other embodiments disclosed is a fusion protein comprising a HIV-1 transactivator of transcription (TAT) domain fused to human ornithine transcarbamoylase (OTC) and a human mitochondria targeting sequence (MTS) of a human mitochondrial protein selected from lipoamide dehydrogenase (LAD) and citrate synthase (CS) situated between said TAT domain and said OTC, wherein said OTC is C-terminal to said MTS of human lipoamide dehydrogenase or human citrate synthase.
[0185] As indicated above, the fusion protein according to the presently disclosed subject matter comprises a HIV-1 transactivator of transcription (TAT) domain, located at the N-terminus of the fusion protein, N-terminal to the MTS as defined above, which in turn is situated N-terminal to the functional human mitochondrial protein (see FIG. 1 for a schematic presentation).
[0186] As used herein, the term HIV-1 transactivator of transcription (TAT) domain refers to a protein transduction domain which is an 11-amino-acid (residues 47-57) arginine- and lysine-rich portion of the MV-I Tat protein having the amino acid sequence YGRKKRRQRRR as set forth in SEQ ID NO. 21. TAT-fusion protein constructs are known in the art to be introduced into cultured cells, intact tissue, and live tissues and cross the blood-brain barrier (BBB). TAT fusion proteins are also known to traverse mitochondrial membranes [13].
[0187] The presently disclosed subject matter also encompasses any fragments of the above defined TAT domain. For example, a TAT domain according to the presently disclosed subject matter may comprise from about 3 to about 11 (e.g. 4-11, 5-11, 6-11, 7-11, 8-11, 9, 10 or 11) sequential amino acid residues of the HIV-I Tat protein having the amino acid sequence YGRKKRRQRRR (SEQ ID NO. 21).
[0188] In some embodiments, the fragment of the above defined TAT domain comprise 9 sequential amino acid residues of the HIV-I Tat protein, having the amino acid sequence of RKKRRQRRR, as set forth in SEQ ID NO. 27 and encoded by the nucleic acid sequence denoted by SEQ ID NO. 1, which was used in the preparation of the fusion protein constructs exemplified below.
[0189] Thus, in this and other embodiments of the presently disclosed subject matter, the fusion protein comprises a TAT domain at its N-terminus and a functional mitochondrial protein at its C-terminus, both covalently linked (fused) to an MTS that is situated between said TAT domain and said functional mitochondrial protein. In other words, the disclosure provides a protein construct comprising an N-terminal TAT fused to N-terminal of MTS fused to N-terminal of functional protein, as schematically presented in FIG. 1 for frataxin.
[0190] The fusion protein according to the presently disclosed subject matter may be prepared by any method known to a skilled artisan. By example, the fusion protein as herein defined may be prepared as exemplified below, by standard molecular biology and cloning techniques.
[0191] The term "fusion protein" in the context of the invention concerns a sequence of amino acids, predominantly (but not necessarily) connected to each other by peptidic bonds. The term "fused" in accordance with the fusion protein of the invention refers to the fact that the amino acid sequences of at least three different origins, namely, the TAT domain, the sequence of the heterologous mitochondrial targeting domain (MTS) and the functional mitochondrial protein, are linked to each other by covalent bonds either directly or via an amino acid linker joining (bridging, conjugating, covalently binding) the amino acid sequences. The fusion may be by chemical conjugation such as by using state of the art methodologies used for conjugating peptides.
[0192] The fusion protein in the context of the invention may also optionally cc p se at least one linker covalently joining different domains of the fusion protein construct.
[0193] The term "linker" in the context of the invention concerns an amino acid sequence of from about 4 to about 20 amino acid residues positioned between the different fusion protein domains and covalently joining them together. For example, a linker in accordance with the invention may be 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 amino acid long. Linkers are often composed of flexible amino acid residues, for example but not limited to glycine and serine so that the adjacent protein domains are free to move relative to one another. The term "linker" can be interchageably used with "spacer".
[0194] The design of a linker that enables proper folding of the various domains of a protein is well known in the art. A non-binding example of a linker is the amino acid sequence GSDP (Gly-Ser-Asp-Pro) as denoted by SEQ ID NO. 32, which was used in the Examples below to construct the fusion proteins His TAT MTS(cs) 81-210 FRA (denoted by SEQ ID NO. 17), His TAT MTS(fra) 81-210 FRA (denoted by SEQ ID NO. 18), His TAT MTS(lad) 81-210 FRA (denoted by SEQ ID NO. 19) and His TAT MTS(orf) 81-210 FRA (denoted by SEQ ID NO. 20) as well as several of the fusion proteins comprising OTC.
[0195] Thus in some embodiments the present disclosure relates to a fusion protein as herein defined further comprising a linker covalently linking said TAT domain to said MTS sequence.
[0196] The fusion protein in the context of the invention may also optionally comprise at least one methionine (M) residue at its N-terminus, as in the case of the exemplified fusion proteins below. The methionine is positioned N-terminal to the TAT domain.
[0197] Fusion may also be achieved by recombinant techniques, i.e. by construction of a nucleic acid. sequence coding for the entire the fusion protein (coding for all segments) so that essentially all the bonds are peptidic bonds.
[0198] In order to facilitate purification of the protein constructs described herein, fusion protein constructs in accordance with the invention may also comprise an N-terminal tag (e.g. His tag as exemplified below, Glutathione S-transferase (GST), Maltose-Binding Protein (MBP), FLAG octapeptide, to name but few), which may be removed or retained in the final fusion construct. Such tags are normally cleaved off from the fusion protein upon entry to the mitochondria, along with the TAT and MTS sequences.
[0199] In some embodiments, the amino acid sequence of a fusion protein according to the invention is as set forth in SEQ ID NO. 17, namely His TAT MTS(cs) FRA, SEQ ID NO. 19, namely His TAT MTS(lad) FRA, SEQ ID NO. 20, namely, His TAT MTS(orf) FRA, SEQ ID NO. 45, namely, His TAT MTS(cs) OTC, SEQ ID NO. 46, namely, His TAT MTS(orf) OTC as well as SEQ ID NO. 47, namely, His TAT MTS(lad) OTC.
[0200] Fusion protein constructs in accordance with the invention may also be prepared without an N-terminal tag. In some embodiments, the amino acid sequence of a fusion protein according to the invention is as set forth in SEQ ID NO. 28, namely TAT MTS(cs) FRA, SEQ ID NO. 30, namely TAT MTS(lad) FRA, SEQ ID NO. 31, namely, TAT MTS(orf) FRA, SEQ ID NO. 49, namely TAT MTS(cs) OTC, SEQ ID NO. 50, namely TAT MTS(orf) OTC, as well as in SEQ ID NO. 51, namely TAT MTS(lad) OTC.
[0201] Therefore the present disclosure further encompasses a fusion protein having the amino acid sequence denoted by SEQ ID NO. 30, comprising a HIV-1 transactivator of transcription (TAT) domain having the amino acid sequence denoted by SEQ ID NO. 27 fused to human frataxin having the amino acid sequence denoted by SEQ ID NO. 26 and a mitochondria targeting sequence (MTS) of human lipoamide dehydrogenase having the amino acid sequence denoted by SEQ ID NO. 24, said MTS situated between said TAT domain and said frataxin and is linked to said TAT domain via a linker having the amino acid sequence denoted by SEQ ID NO. 32, and wherein said frataxin is C-terminal to said MTS of human lipoamide dehydrogenase.
[0202] In some embodiments the fusion protein as herein defined has the amino acid sequence denoted by SEQ ID NO. 28, comprising a HIV-1 transactivator of transcription (TAT) domain having the amino acid sequence denoted by SEQ ID NO. 27 fused to human frataxin having the amino acid sequence denoted by SEQ ID NO. 26 and a mitochondria targeting sequence (MTS) of human citrate synthase having the amino acid sequence denoted by SEQ ID NO. 23, said MTS situated between said TAT domain and said frataxin, and is linked to said TAT domain via a linker having the amino acid sequence denoted by SEQ ID NO. 32, and wherein said frataxin is C-terminal to said MTS of human citrate synthase.
[0203] In some embodiments the fusion protein as herein defined has the amino acid sequence denoted by SEQ ID NO. 45, comprising a HIV-1 transactivator of transcription (TAT) domain having the amino acid sequence denoted by SEQ ID NO. 27 fused to human ornithine transcarbamoylase (OTC) having the amino acid sequence denoted by SEQ ID NO. 39 and a mitochondria targeting sequence (MTS) of human citrate synthase having the amino acid sequence denoted by SEQ ID NO. 23, said MTS situated between said TAT domain and said OTC, and is linked to said TAT domain via a linker having the amino acid sequence denoted by SEQ ID NO. 32, and wherein said OTC is C-terminal to said MTS of human citrate synthase.
[0204] The presently disclosed subject matter further provides a composition comprising a physiologically acceptable carrier and as an active ingredient a fusion protein as herein defined.
[0205] In specific embodiments the presently disclosed subject matter provides a composition comprising as an active ingredient a fusion protein having the amino acid sequence denoted by SEQ ID NO. 45, denoted by SEQ ID NO. 30, or having the amino acid sequence denoted by SEQ ID NO. 28 and a physiologically acceptable carrier.
[0206] Also provided by the presently disclosed subject matter is a pharmaceutical composition comprising a pharmaceutically acceptable carrier and as an active ingredient a fusion protein construct as herein defined.
[0207] The "composition" as herein defined generally comprises a buffering agent, an agent which adjusts the osmolarity thereof, and optionally, one or more pharmaceutically (or physiologically) acceptable carriers, diluents, additives and excipients as known in the art. Supplementary active ingredients can also be incorporated into the compositions. The pharmaceutically acceptable carrier can be solvent or dispersion medium containing, for example, water, ethanol, polyol (for example, glycerol, propylene glycol, and liquid polyethylene glycol, and the like), suitable mixtures thereof, and vegetable oils. Each carrier should be physiologically or pharmaceutically acceptable, as the case may be, in the sense of being compatible with the other ingredients and not injurious to the patient.
[0208] The additives may be but are not limited to at least one of a protease inhibitor, for example phenylmethanesulfonylfluoride or phenylmethylsulfonyl fluoride (PMSF), Nafamostat Mesylate, 4-(2-Aminoethyl)benzenesulfonyl fluoride hydrochloride (AEBSF), Bestatin, Pepstatin A, E-64, Leupeptin, 1, 10-Phenanthroline and any other protease inhibitor known in the art.
[0209] The "pharmaceutical compositions" of the presently disclosed subject matter are compositions as described above, comprising pharmaceutically acceptable carriers, diluent, adjuvant and/or excipients and/or additives as known in the art.
[0210] The presently disclosed subject matter further provides a pharmaceutical composition as herein defined for treating or alleviating a mitochondrial disorder.
[0211] The term "mitochondrial disorder" as encompassed by the presently disclosed subject matter refers to a group of systemic diseases caused by inherited or acquired damage to the mitochondria causing an energy shortage within those areas of the body that consume large amounts of energy such as the liver, muscles, brain, and the heart. The result is often liver failure, muscle weakness, fatigue, and problems with the heart, eyes, and various other systems.
[0212] The mitochondrial disorder may be any one of frataxin deficiency which causes or is associated with Friedreich's ataxia; a deficiency in OTC (X-linked recessive genetic disorder caused by non-conservative mutations in the OTC gene); disorder associated with LAD deficiency; or the mitochondrial metabolic disorder is Complex I deficiency (OMIM:252010). Complex I deficiency can be caused by a mutation in any of the subunits thereof. Alternatively, the Complex I deficiency is caused by a mutation in a gene selected from NDUFV1 (OMIM: 161015), NDUFV2 (OMIM:600532), NDUFS1 (OMIM: 157655), NDUFS2 (OMIM:602985), NDUFS3 (OMIM:603846), NDUFS4 (OMIM:602694), NDUFS6 (OMIM:603848), NDUFS7 (OMIM:601825), NDUFS8 (OMIM:602141), and NDUF A2 (OMIM: 602137).
[0213] In other embodiments, the mitochondrial disorder is Complex IV deficiency (Cytochrome C oxidase; OMIM:220110). Complex IV deficiency can be caused by a mutation in any of the subunits thereof. In another embodiment, the Complex IV deficiency is caused by a mutation in a gene selected from the group consisting of MTCO1 (0MIM:516030), MTCO2 (0MIM:516040), MTCO3 (0MIM:516050), COX1O (0MIM:602125), COX6B1 (OMIM: 124089), SCO1 (OMIM:603644), FASTKD2 (0MIM:612322), and SC02 (OMIM:604272).
[0214] In other embodiments, the mitochondrial disorder is a neurodegenerative disease. As provided herein, compositions of the present invention exhibit the ability to traverse the blood-brain barrier (BBB).
[0215] In further embodiments of the presently disclosed subject matter, the mitochondrial disorder is selected from the group consisting of encephalopathy and liver failure that is accompanied by stormy lactic acidosis, hyperammonemia and coagulopathy. In other embodiments, the mitochondrial disorder is selected from the group consisting of Ornithine Transcarbamoylase deficiency (hyperammonemia) (OTCD), Carnitine O-palmitoyltransferase II deficiency (CPT2), Fumarase deficiency, Cytochrome c oxidase deficiency associated with Leigh syndrome, Maple Syrup Urine Disease (MSUD), Medium-Chain Acyl-CoA Dehydrogenase deficiency (MCAD), Acyl-CoA Dehydrogenase Very Long-Chain deficiency (LCAD), Trifunctional Protein deficiency, Progressive External Ophthalmoplegia with Mitochondrial DNA Deletions (POLG), DGUOK, TK2, Pyruvate Decarboxylase deficiency, and Leigh Syndrome (LS). In another embodiment, the mitochondrial metabolic disorder is selected from the group consisting of Alpers Disease; Barth syndrome; beta-oxidation defects; carnitine-acyl-carnitine deficiency; carnitine deficiency; co-enzyme Q10 deficiency; Complex II deficiency (OMIM:252011), Complex III deficiency (OMIM: 124000), Complex V deficiency (OMIM:604273), LHON-Leber Hereditary Optic Neuropathy; MM--Mitochondrial Myopathy; LIMM-Lethal Infantile Mitochondrial Myopathy; MMC--Maternal Myopathy and Cardiomyopathy; NARP-Neurogenic muscle weakness, Ataxia, and Retinitis Pigmentosa; Leigh Disease; FICP-Fatal Infantile Cardiomyopathy Plus, a MELAS-associated cardiomyopathy; MELAS-Mitochondrial Encephalomyopathy with Lactic Acidosis and Strokelike episodes; LDYT-Leber's hereditary optic neuropathy and Dystonia; MERRF-Myoclonic Epilepsy and Ragged Red Muscle Fibers; MHCM-Maternally inherited Hypertrophic CardioMyopathy; CPEO-Chronic Progressive External Ophthalmoplegia; KSS-Kearns Sayre Syndrome; DM-Diabetes Mellitus; DMDF Diabetes Mellitus+Deafness; CIPO-Chronic Intestinal Pseudoobstruction with myopathy and Ophthalmoplegia; DEAF-Maternally inherited DEAFness or aminoglycoside-induced DEAFness; PEM-Progres sive encephalopathy; SNHL-SensoriNeural Hearing Loss; Encephalomyopathy; Mitochondrial cytopathy; Dilated Cardiomyopathy; GER-Gastro intestinal Reflux; DEMCHO-Dementia and Chorea; AMDF-Ataxia, Myoclonus; Exercise Intolerance; ESOC Epilepsy, Strokes, Optic atrophy, and Cognitive decline; FBSN Familial Bilateral Striatal Necrosis; FSGS Focal Segmental Glomerulosclerosis; LIMM Lethal Infantile Mitochondrial Myopathy; MDM Myopathy and Diabetes Mellitus; MEPR Myoclonic Epilepsy and Psychomotor Regression; MERME MERRF/MELAS overlap disease; MHCM Maternally Inherited Hypertrophic CardioMyopathy; MICM Maternally Inherited Cardiomyopathy; MILS Maternally Inherited Leigh Syndrome; Mitochondrial Encephalocardio myopathy; Multisystem Mitochondrial Disorder (myopathy, encephalopathy, blindness, hearing loss, peripheral neuropathy); NAION Nonarteritic Anterior Ischemic Optic Neuropathy; NIDDM Non-Insulin Dependent Diabetes Mellitus; PEM Progressive Encephalopathy; PME Progressive Myoclonus Epilepsy; RTT Rett Syndrome; SIDS Sudden Infant Death Syndrome; MIDD Maternally Inherited Diabetes and Deafness; and MODY Maturity-Onset Diabetes of the Young, and MNGIE.
[0216] Each mitochondrial disease represents an embodiment of the present invention.
[0217] In the above and other embodiments, the presently disclosed subject matter provides a pharmaceutical composition as herein defined for use in the treatment of Friedreich's ataxia or any other disorder associated with deficiency of frataxin or defective frataxin or for use in the treatment of a disorder associated with a deficiency of OTC or with defective OTC.
[0218] In still further embodiments, the presently disclosed subject matter provides a pharmaceutical composition for the treatment of Friedreich's Ataxia by intravenous administration to a subject in need thereof, said composition comprising a therapeutically effective amount of a fusion protein as herein defined.
[0219] In specific embodiments the presently disclosed subject matter provides a pharmaceutical composition for the treatment of Friedreich's Ataxia by intravenous administration to a subject in need thereof, said composition comprising a therapeutically effective amount of a fusion protein having the amino acid sequence denoted by SEQ ID NO. 30 or having the amino acid sequence denoted by SEQ ID NO. 28.
[0220] In further specific embodiments the presently disclosed subject matter provides a pharmaceutical composition for the treatment of Friedreich's Ataxia by intravenous administration to a subject in need thereof, said composition comprising a therapeutically effective amount of a fusion protein having the amino acid sequence denoted by SEQ ID NO. 30.
[0221] In still further specific embodiments the presently disclosed subject matter provides a pharmaceutical composition for the treatment of Friedreich's Ataxia by intravenous administration to a subject in need thereof, said composition comprising a therapeutically effective amount of a fusion protein having the amino acid sequence denoted by SEQ ID NO. 28.
[0222] The presently disclosed subject matter further provides a fusion protein according to the invention for use in a method for the treatment of a mitochondrial disorder.
[0223] In the above and other embodiments of the presently disclosed subject matter the functional human mitochondrial protein is frataxin for use in a method for the treatment or alleviation of Friedreich's ataxia or any other disorder associated with deficiency of frataxin or defective frataxin.
[0224] In the above and other embodiments of the presently disclosed subject matter the functional human mitochondrial protein is OTC for use in a method for the treatment or alleviation of a disorder associated with a deficiency of OTC or with defective OTC.
[0225] As shown below in Example 7, the inventors have shown that fusion proteins comprising TAT, MTS and frataxin (TAT-MTS-FRA) were able to partially rescue cells obtained from Friedreich's ataxia patients, as well as normal cells, from oxidative stress, exhibiting a clear biological activity of the fusion protein constructs of the presently disclosed subject matter.
[0226] L-Buthionine sulphoximine (BSO) is an inhibitor of gamma-glutamylcysteine synthetase (gamma-GCS) and, consequently lowers tissue glutathione (GSH) concentrations. GSH plays an important role in cellular defense against a wide variety of toxic electrophiles via the formation of thioether conjugates. Therefore, BSO was used by the inventors to model oxidative stress, through its ability to inhibit de novo glutathione synthesis, thereby depleting an important component of these cells' intrinsic defenses against reactive oxygen species (ROS) and allowing for the accumulation of ROS produced by natural cell processes, known to result in cell death.
[0227] It is known that Friedreich ataxia cells are extremely sensitive to BSO-induced oxidative stress compared with normal cells because they lack Frataxin, and thus are used as an in vitro model of the long-term consequences of absent Frataxin.
[0228] As shown in the Examples below, oxidative stress was induced with various concentrations of BSO in cells obtained from patients as well as in normal healthy cells and the effect of the various TAT-MTS-FRA fusion proteins on cell death was measured. As can be seen in FIG. 11, BSO caused cell death of normal lymphocytes as well as of cells obtained from Friedreich ataxia patients. However, cells obtained from patients were more sensitive to BSO-induced oxidative stress, consistent with previous findings. Most importantly, the various TAT-MTS-FRA fusion proteins, which were added a few hours before oxidative stress induction, were demonstrated to partially rescue both normal lymphocytes as well as patients' cells from cell death. This partial rescue was determined by both reduction in cell death and by reduction in caspase 3 activity.
[0229] Surprisingly, as also shown in FIG. 11, at least two out of the three fusion proteins carrying a heterologous MTS (namely, MTSorf and MTScs) demonstrated a superior protective effect with respect to the effect demonstrated by the fusion protein carrying the native MTS.
[0230] In addition, in an independent comparative experiment shown in FIG. 12, in which the effect of the various TAT-MTS-FRA fusion proteins on oxidative stress was examined, a fusion protein carrying another heterologous MTS, namely, MTSlad also demonstrated a superior protective effect with respect to the effect demonstrated by the fusion protein carrying the native MTS (FIG. 12C).
[0231] Thus the presently disclosed subject matter further provides a method for treating or alleviating a mitochondrial disorder, said method comprising the step of administering to a subject in need of such treatment a therapeutically effective amount of the fusion protein as defined herein, thereby treating a mitochondria disorder.
[0232] In the above and other embodiments, the method for treating or alleviating a mitochondrial disorder according to the invention is wherein said functional protein is frataxin or OTC, and the mitochondrial disorder is Friedreich's ataxia or any other disorder associated with deficiency of frataxin or defective frataxin or respectively, a disorder associated with a deficiency of OTC or with defective OTC.
[0233] The term "treat" or "treatment" or forms thereof as herein defined means to prevent worsening or arrest or alleviate or cure the disease or condition in a subject in need thereof. Thus the term "treatment", "treating" or "alleviating" in the context of the intention does not refers to complete curing of the diseases, as it does not change the mutated genetics causing the disease. This term refers to alleviating at least one of the undesired symptoms associated with the disease, improving the quality of life of the subject, decreasing disease-caused mortality, or (if the treatment in administered early enough) preventing the full manifestation of the mitochondrial disorder before it occurs, mainly to organs and tissues that have a high energy demand. The treatment may be a continuous prolonged treatment for a chronic disease or a single, several or multiple administrations for the treatment of an acute condition such as encephalopathy and liver failure that is accompanied by stormy lactic acidosis, hyperammonemia and coagulopathy.
[0234] Notably, in the case of metabolic or mitochondrial disorders there is no need to restore protein activity back to 100%, but rather raise it above a certain energetic threshold which can vary from patient to patient depending on basal protein activity.
[0235] In addition, treatment of mitochondrial disorders using replacement therapy is necessarily more complex than replacement of a cytosolic gene product and must consider not only the need to target and cross multiple membranes in mitochondria, but also the fact that many proteins in the mitochondria act as multi-component complexes which require appropriate assembly in order to integrate properly. Additionally, many of the mitochondrial gene defects cause severe neurologic symptoms as the primary or most prominent phenotype, and, as mentioned above, drug delivery across the BBB is difficult.
[0236] Therapeutic formulations may be administered in any conventional route and dosage formulation. Formulations typically comprise at least one active ingredient, as defined above, together with one or more acceptable carriers thereof. Thus, administration can be any one of intravenous, intraperitoneal, intramuscular and intratechal administration. Oral administration is also contemplated.
[0237] The term "therapeutically effective amount" (or amounts) of the peptide for purposes herein defined is determined by such considerations as are known in the art in order to cure or at least arrest or at least alleviate the medical condition.
[0238] As indicated above, in the case of metabolic or mitochondrial disorders it is not required to restore protein activity back to 100%. Rather, raising the protein activity to above a certain energetic threshold, which can vary from patient to patient depending on basal protein activity, can be sufficient to provide a therapeutic effect.
[0239] In some embodiments the therapeutically effective amount may be determined for each patient individually, based on the patient's basal protein activity. The patient's basal protein activity, or the level of protein activity may in turn be determined using any method known in the art, for example, by subjecting a biological sample obtained from a patient (e.g. blood (white cells), skin fibroblast cells, whole tissue biopsies) to an in vitro enzymatic assay or immunohistochemistry or any specific imaging such as PET.
[0240] In some embodiments the functional human mitochondrial protein is OTC and the patient's basal OTC activity is determined by liver biopsy.
[0241] In some embodiments the therapeutically effective amount according to the presently disclosed subject matter is between about 0.5 mg/kg to about 2.0 mg/kg body weight of the subject in need thereof which is the Human Equivalent Dose (HED) of an effective amount in mice of between about 7 mg/kg to about 25 mg/kg.
[0242] Thus in specific embodiments disclosed is a pharmaceutical composition as herein defined wherein the therapeutically effective amount administered is from about 0.5 mg/Kg to about 2 mg/Kg body weight of said subject.
[0243] As used herein, the term "subject in need" is to be taken to mean a human suffering from a mitochondrial disorder as herein defined.
[0244] In the above and other embodiments, the method for treating or alleviating a mitochondrial disorder according to the invention further comprises administering an additional therapeutic agent.
[0245] The term "additional therapeutic agent" as herein defined refers to any agent that is administered in addition to the fusion protein according to the invention in order to alleviate the symptoms associated with the disease or disorder the treatment of which is desirable. In the above and other embodiments of the disclosed subject matter, the fusion protein of the invention and said additional therapeutic agent are administered simultaneously. Alternatively or additionally, said fusion protein and said additional therapeutic agent are administered at different time points, at different intervals between administrations, for different durations of time, or in a different order.
[0246] The disclosed subject matter further provides a method for introducing a functional mitochondrial protein into mitochondria of a subject, said method comprising the step of administering to a subject in need of such treatment a therapeutically effective amount of the fusion protein as defined herein, thereby introducing a functional mitochondrial protein into the mitochondria of a subject in need thereof.
[0247] In some embodiments the functional human mitochondrial protein thereby introduced restores at least partial activity of the wild type human mitochondrial protein, for example at least 5%, at least 10%, at least 15%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90% or up to 100% of the activity of a wild type human mitochondrial protein.
[0248] Thus, the disclosed subject matter further provides a method for restoring at least in part activity of a defective or deficient or unfunctional human mitochondrial protein in a subject in need, by administering a fusion as herein disclosed to said subject. The human mitochondrial protein may be active per se, or may be a member of a mitochondrial protein complex.
[0249] By another one of its aspects, the disclosed subject matter further provides a method for alleviating oxidative stress in a subject in need thereof, said method comprising the step of administering to a subject in need of such treatment a therapeutically effective amount of the fusion protein as herein defined, thereby alleviating oxidative stress in said subject.
[0250] The term "oxidative stress" as herein defined refers to an imbalance between the systemic manifestations of reactive oxygen species (ROS) and an ability of a biological system to readily detoxify the reactive intermediates or to repair the resulting damage. Disturbances in the normal redox state of cells can cause toxic effects through the production of peroxides and free radicals that damage all components of the cell. In addition, since some reactive oxidative species act as cellular messengers in redox signaling, accumulation of ROS can cause disruptions in normal mechanisms of cellular signaling. In humans, oxidative stress is thought to be involved in the development of various disorders and conditions, among which are cancer, Parkinson's disease, Alzheimer disease to name but a few.
[0251] It is appreciated that certain features of the presently disclosed subject matter, which are, for clarity, described in the context of separate embodiments, may also be provided in combination in a single embodiment. Conversely, various features of the invention, which are, for brevity, described in the context of a single embodiment, may also be provided separately or in any suitable sub combination or as suitable in any other described embodiment of the invention. Certain features described in the context of various embodiments are not to be considered essential features of those embodiments, unless the embodiment is inoperative without those elements.
[0252] Disclosed and described, it is to be understood that this invention is not limited to the particular examples, process steps, and materials disclosed herein as such process steps and materials may vary somewhat. It is also to be understood that the terminology used herein is used for the purpose of describing particular embodiments only and not intended to be limiting since the scope of the present invention will be limited only by the appended claims and equivalents thereof.
[0253] It must be noted that, as used in this specification and the appended claims, the singular forms "a", "an" and "the" include plural referents unless the content clearly dictates otherwise.
[0254] Throughout this specification and the claims which follow, unless the context requires otherwise, the word "comprise", and variations such as "comprises" and "comprising", will be understood to imply the inclusion of a stated integer or step or group of integers or steps but not the exclusion of any other integer or step or group of integers or steps.
[0255] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention is related. The following terms are defined for purposes of the invention as described herein.
[0256] The following Examples are representative of techniques employed by the inventors in carrying out aspects of the present invention. It should be appreciated that while these techniques are exemplary of preferred embodiments for the practice of the invention, those of skill in the art, in light of the present disclosure, will recognize that numerous modifications can be made without departing from the spirit and intended scope of the invention.
EXAMPLES
[0257] Without further elaboration, it is believed that one skilled in the art can, using the preceding description, utilize the present invention to its fullest extent. The following preferred specific embodiments are, therefore, to be construed as merely illustrative, and not limitative of the claimed invention in any way.
[0258] Standard molecular biology protocols known in the art not specifically described herein are generally followed essentially as in Sambrook & Russell, 2001.
Experimental Procedures
Cell Cultures
[0259] Lymphocytes (Lym 43) and fibroblasts (Fib. 78 and Fib. 65, F816) from Friedreich's ataxia. patients were obtained from Coriell Cell Repositories (Camden, N.J.) and grown at the recommended medium (fibroblasts in Eagle's Minimum Essential Medium (EMEM) with Earle's salts and non-essential amino acids, and lymphocytes in Roswell Park Memorial Institute (RPMI) Medium 1640 with 2m M L-glutamine, all supplemented with 15% fetal bovine serum not inactivated). Human BJ AB cells (EBV-negative Burkitt's lymphoma cells; provided by Hanna Ben-Bassat, Hadassah Medical Center, Jerusalem, Israel) were grown in RPMI medium supplemented with 20% heat-inactivated fetal calf serum (FCS), 2 mM. L-glutamine, 100 U/mL penicillin, and 100 .mu.g/mL streptomycin. Normal lymphocytes [20] were grown in RPMI 1640 supplemented with 10% fetal calf serum (FCS) and antibiotics as above. HepG2 cells (ATCC: HB-8065) were grown in EMEM supplemented with 10% FCS, 2 mM L-Glutamine, non-essential amino acids, antibiotics and sodium pyruvate. All culture cells were grown in 37.degree. C. with 5% CO.sub.2. All media and supplements were purchased from Biological Industries (Bet Ha'emek, Israel). The cells were kept in a humidified atmosphere with 5% CO.sub.2 at 37.degree. C. All cultures were tested for mycoplasma contamination and were found to be negative.
Cloning of the Plasmids Encoding the Fusion Proteins
His-TAT-MTSfra-FRA
[0260] The plasmid His-TAT-LAD was prepared as previously reported [23] and was cut with BamHI and XhoII to remove the LAD sequence, thus obtaining a vector fragment encoding the His-TAT domain. The full-length frataxin was generated by PCR using a frataxin clone (purchased from Open Biosystem. Ltd., clone no. 4842134) as a template and a pair primers covering the whole sequence, including its native MTS, 5'-CGCGGATCCGTGGACTCTCGGGCGCCG-3' (forward) and 5'-ACGCTCGAGTCAAGCATCTTTCCGGAATAGGC-3' (reverse), as denoted by SEQ ID NO. 7 and SEQ ID NO. 8, respectively. The PCR fragment was cut with BamHI and XhoII and ligated with the vector fragment, thus obtaining the plasmid encoding the His-TAT-MTSfra-FRA fusion protein.
His-MTSlad-FRA
[0261] The plasmid encoding the His-TAT-MTSfra-FRA was cut with BamHI and BsaI to remove the MTSfra sequence. The MTSlad (the native MTS of lipoamide dehydrogenase) was obtained by PCR using the plasmid His-TAT-LAD [23] as a template and the following pair of primers: 5'-CGCGGATCCACAGAGCTGGAGTCGTGTGTA-3' (forward) and 5'-CATAGG-TGGTCTCATCTAGAGAGCCTGGGTGGCCCAAAGTTCCAGATGCGTAAGTTCTCAGAG GCA-3' (reverse) as denoted by SEQ ID NO. 9 and SEQ ID NO. 10, respectively. The PCR product was cut with BamHI and BsaI and ligated to the vector fragment thus obtaining the plasmid encoding the His-TAT-MTSlad-FRA fusion protein.
His-TAT-MTSorf-FRA
[0262] The plasmid encoding the His-TAT-MTSfra-FRA was cut with BamHI and BsaI to remove the MTSfra sequence as described above. The MTSorf (the native MTS of the protein C6ORF66 assembly factor) was obtained by PCR using the plasmid His-TAT-ORF as a template and the following pair of primers: 5'-CGCGGATCCGGGAGCACTAGTGATTCGC-3' (forward) and 5'-CATAGGTG GTCTCATCTAGAGAGCCTGGGTGGCCCAAAGTTCCAGAAGAGGGGTGTCTGGGAGC GA-3' (reverse), as denoted by SEQ ID NO. 11 and SEQ ID NO. 12, respectively. The PCR product was cut with BamHI and BsaI and ligated to the vector fragment thus obtaining the plasmid encoding the His-TAT-MTSorf-FRA fusion protein.
His-TAT-MTScs-FRA
[0263] The plasmid encoding the His-TAT-MTSfra-FRA was cut with BamHI and BsaI to remove the MTSfra sequence as described above. The MTScs (the native MTS of the protein citrate synthase) was generated by synthesizing two oligonucleotides covering the MTScs sequence and the BamHI and BsaI sites at the ends: 5'-GATCCGGCTTACTTACTGCGGCCGCCCGGCTCTTGGGAACCAAGAATGCATCTTGT CTTGTTCTTGCAGCCCGGCATGCCAGTTCTGGAACTTrGGGCCACCCAGGCTCTC-3' (forward) and 5'-TCTAGAGAGCCTGGGTGGCCCAAAGTTCCAG AACTGGCATGCCGGGCTGCAAGAACAAGACAAGATGCATTCTTGGTTCCCAAGAGC CGGGCGGCCGCAGTAAGTAAAGCCG-3' (reverse), as denoted by SEQ ID NO. 13 and SEQ ID NO. 14, respectively. The oligonucleotides were ligated to the vector fragment thus obtaining the plasmid encoding the His-TAT-MTScs-FRA fusion protein. In some of the assays described below the His-TAT-MTScs-FRA fusion protein construct used was obtained from Genscript (Lot #342511S03/P10011312; 2.5 mg/ml stock in 50 mM Tris-HCl, 300 mM NaCl, 10% Glycerol, pH 8.0).
[0264] All plasmids were confirmed by restriction enzymes and sequencing analyses.
TABLE-US-00001 TABLE 1 Nucleic acid and Amino acid sequences SEQ ID NO. Sequence Name 1 aggaagaagcggagacagcgacgaaga nt seq. encoding TAT fragment 2 TGGACTCTCGGGCGCCGCgcagtagccggcctcctggcgtcacccagc nt seq. of ccggcccaggcccagaccctcacccgggtcccgcggccggcagagttggccccactc MTSfra tgcggccgccgtggcctgcgcaccgacatcgatgcgacctgcacgccccgccgcgca agttcgaaccaacgtggcctcaaccagatttggaatgtcaaaaagcagagtgtctat ttgatgaatttgaggaaa 3 GCTTTACTTACTGCGGCCGCCCGGCTCTTGGGAACCA nt seq. of AGAATGCATCTTGTCTTGTTCTTGCAGCCCGGCATGCC MTScs AGT 4 ggagcactagtgattcgcggtatcaggaatttcaacctagagaaccgagcggaacgg nt seq. of gaaatcagcaagatgaagccctctgtcgctcccagacacccctct MTSorf 5 cagagctggagtcgtgtgtactgctccttggccaagagaggccatttcaatcgaata nt seq. of tctcatggcctacagggactttctgcagtgcctctgagaacttacgca MTSlad 6 tctggaactttgggccacccaggctctctagatgagaccacctatgaaagactagcag nt seq. of aggaaacgctggactctttagcagagttattgaagaccttgcagacaagccatacacgt Mature ttgaggactatgatgtctcctttgggagtggtgtcttaactgtcaaactgggtggagatc Frataxin taggaacctatgtgatcaacaagcagacgccaaacaagcaaatctggctatcttctccat (FRA) ccagtggacctaagcgttatgactggactgggaaaaactgggtgtactcccacgacggcg tgtccctccatgagctgctggccgcagagctcactaaagccttaaaaaccaaactggact tgtcttCCTTGGCCTATTCCGGAAAAGATGCTTGA 7 CGCGGATCCGTGGACTCTCGGGCGCCG forward primer for precursor frataxin cloning 8 ACGCTCGAGTCAAGCATCTTTTCCGGAATAGGC reverse primer for precursor frataxin cloning 9 CGCGGATCCACAGAGCTGGAGTCGTGTGTA forward primer for MTS lad cloning 10 CATAGGTGGTCTCATCTAGAGAGCCTGGGTGGCCCAA reverse AGTTCCAGATGCGTAAGTTCTCAGAGGCA primer for MTS lad cloning 11 CGCGGATCCGGGAGCACTAGTGATTCGC forward primer for MTS orf cloning 12 CATAGGTGGTCTCATCTAGAGAGCCTGGGTGGCCCAA reverse AGTTCCAGAAGAGGGGTGTCTGGGAGCGA primer for MTS orf cloning 13 GATCCGGCTTTACTTACTGCGGCCGCCCGGCTCTTGGG forward AACCAAGAATGCATCTTGTCTTGTTCTTGCAGCCCGGC oligo for ATGCCAGTTCTGGAACTTTGGGCCACCCAGGCTCTC MTS cs cloning 14 TCTAGAGAGCCTGGGTGGCCCAAAGTTCCAGAACTGG reverse CATGCCGGGCTGCAAGAACAAGACAAGATGCATTCTT primer for GGTTCCCAAGAGCCGGGCGGCCGCAGTAAGTAAAGCC MTS cs G cloning 15 CTGAAGGGCCGTGACCTTCTCACTCTAAGAAACTTTA nt seq. of CCGGAGAAGAAATTAAATATATGCTATGGCTATCAGC Mature AGATCTGAAATTTAGGATAAAACAGAAAGGAGAGTA OTC TTTGCCTTTATTGCAAGGGAAGTCCTTAGGCATGATTT TTGAGAAAAGAAGTACTCGAACAAGATTGTCTACAGA AACAGGCTTTGCACTTCTGGGAGGACATCCTTGTTTTC TTACCACACAAGATATTCATTTGGGTGTGAATGAAAG TCTCACGGACACGGCCCGTGTATTGTCTAGCATGGCA GATGCAGTATTGGCTCGAGTGTATAAACAATCAGATT TGGACACCCTGGCTAAAGAAGCATCCATCCCAATTAT CAATGGGCTGTCAGATTTGTACCATCCTATCCAGATCC TGGCTGATTACCTCACGCTCCAGGAACACTATAGCTCT CTGAAAGGTCTTACCCTCAGCTGGATCGGGGATGGGA ACAATATCCTGCACTCCATCATGATGAGCGCAGCGAA ATTCGGAATGCACCTTCAGGCAGCTACTCCAAAGGGT TATGAGCCGGATGCTAGTGTAACCAAGTTGGCAGAGC AGTATGCCAAAGAGAATGGTACCAAGCTGTTGCTGAC AAATGATCCATTGGAAGCAGCGCATGGAGGCAATGTA TTAATTACAGACACTTGGATAAGCATGGGACAAGAAG AGGAGAAGAAAAAGCGGCTCCAGGCTTTCCAAGGTTA CCAGGTTACAATGAAGACTGCTAAAGTTGCTGCCTCT GACTGGACATTTTTACACTGCTTGCCCAGAAAGCCAG AAGAAGTGGATGATGAAGTCTTTTATTCTCCTCGATCA CTAGTGTTCCCAGAGGCAGAAAACAGAAAGTGGACA ATCATGGCTGTCATGGTGTCCCTGCTGACAGATTACTC ACCTCAGCTCCAGAAGCCTAAATTTTGA 16 ATGTACCGCTACCTGGCCAAAGCGCTGCTGCCGTCCC nt seq. of GGGCCGGGCCCGCTGCCCTGGGCTCCGCGGCCAACCA MTS of CTCGGCCGCGTTGCTGGGCCGGGGCCGCGGACAGCCC human GCCGCCGCCTCGCAGCCGGGGCTCGCATTGGCCGCCC mitochon. GGCGCCACTAC GLUD2 17 MGSSHHHHHHSSGLVPRGSHMRKKRRQRRRGSDPALLT aa seq. of AAARLLGTKNASCLVLAARHASSGTLGHPGSLDETTYE His TAT RLAEETLDSLAEFFEDLADKPYTFEDYDVSFGSGVLTVK MTS(cs) LGGDLGTYVINKQTPNKQIWLSSPSSGPKRYDWTGKNW 81-210 VYSHDGVSLHELLAAELTKALKTKLDLSSLAYSGKDAFRA 18 MGSSHHHHHHSSGLVPRGSHMRKKRRQRRRGSDPWTL aa seq. of GRRAVAGLLASPSPAQAQTLTRVPRPAELAPLCGRRGLR HTFrataxin TDIDATCTPRRASSNQRGLNQIWNVKKQSVYLMNLRKS [His GTLGHPGSLDETTYERLAEETLDSLAEFFEDLADKPYTFE TAT DYDVSFGSGVLTVKLGGDLGTYVINKQTPNKQIWLSSPS MTS(fra) SGPKRYDWTGKNWVYSHDGVSLHELLAAELTKALKTK 81-210 LDLSSLAYSGKDA FRA] 19 MGSSHHHHHHSSGLVPRGSHMRKKRRQRRRGSDPQSW aa seq. of SRVYCSLAKRGHFNRISHGLQGLSAVPLRTYASGTLGHP His TAT GSLDETTYERLAEETLDSLAEFFEDLADKPYTFEDYDVS MTS(lad) FGSGVLTVKLGGDLGTYVINKQTPNKQIWLSSPSSGPKR 81-210 YDWTGKNWVYSHDGVSLHELLAAELTKALKTKLDLSS FRA LAYSGKDA 20 MGSSHHHHHHSSGLVPRGSHMRKKRRQRRRGSDPGAL aa seq. of VIRGRNFNLENRAEREISKMKPSVAPRHPSSGTLGHPGS His TAT LDETTYERLAEETLDSLAEFFEDLADKPYTFEDYDVSFG MTS(orf) SGVLTVKLGGDLGTYVINKQTPNKQIWLSSPSSGPKRYD 81-210 WTGKNWVYSHDGVSLHELLAAELTKALKTKLDLSSLA FRA YSGKDA 21 YGRKKRRQRRR aa seq. of TAT 22 WTLGRRAVAGLLASPSPAQAQTLTRVPRPAELAPLCGR aa seq. of RGLRTDIDATCTPRRASSNQRGLNQIWNVKKQSVYLMN MTS(fra) LRK 23 ALLTAAARLLGTKNASCLVLAARHAS aa seq. of MTS(cs) 24 QSWSRVYCSLAKRGHFNRISHGLQGLSAVPLRTYA aa seq. of MTS(lad) 25 GALVIRGIRNFNLENRAEREISKMKPSVAPRHPS aa seq. of MTS(orf) 26 SGTLGHPGSLDETTYERLAEETLDSLAEFFEDLADKPYTF aa seq. of EDYDVSFGSGVLTVKLGGDLGTYVINKQTPNKQIWLSSP Mature SSGPKRYDWTGKNWVYSHDGVSLHELLAAELTKALKT frataxin KLDLSSLAYSGKDA 27 RKKRRQRRR aa seq. of TAT fragment 28 MRKKRRQRRRGSDPALLTAAARLLGTKNASCLVLAAR aa seq. of HASSGTLGHPGSLDETTYERLAEETLDSLAEFFEDLADK TAT PYTFEDYDVSFGSGVLTVKLGGDLGTYVINKQTPNKQI MTS(cs) WLSSPSSGPKRYDWTGKNWVYSHDGVSLHELLAAELT 81-210 KALKTKLDLSSLAYSGKDA FRA 29 MRKKRRQRRRGSDPWTLGRRAVAGLLASPSPAQAQTLT aa seq. of RVPRPAELAPLCGRRGLRTDIDATCTPRRASSNQRGLNQI TAT WNVKKQSVYLMNLRKSGTLGHPGSLDETTYERLAEETL MTS(fra) DSLAEFFEDLADKPYTFEDYDVSFGSGVLTVKLGGDLGT 81-210 YVINKQTPNKQIWLSSPSSGPKRYDWTGKNWVYSHDGV FRA SLHELLAAELTKALKTKLDLSSLAYSGKDA 30 MRKKRRQRRRGSDPQSWSRVYCSLAKRGHFNRISHGLQ aa seq. of GLSAVPLRTYASGTLGHPGSLDETTYERLAEETLDSLAE TAT FFEDLADKPYTFEDYDVSFGSGVLTVKLGGDLGTYVINK MTS(lad) QTPNKQIWLSSPSSGPKRYDWTGKNWVYSHDGVSLHEL 81-210 LAAELTKALKTKLDLSSLAYSGKDA FRA 31 MRKKRRQRRRGSDPGALVIRGIRNFNLENRAEREISKMK aa seq. of PSVAPRHPSSGTLGHPGSLDETTYERLAEETLDSLAEFFE TAT DLADKPYTFEDYDVSFGSGVLTVKLGGDLGTYVINKQT MTS(orf) PNKQIWLSSPSSGPKRYDWTGKNWVYSHDGVSLHELLA 81-210 AELTKALKTKLDLSSLAYSGKDA FRA 32 GSDP aa seq. of linker 33 MRKKRRQRRRALLTAAARLLGTKNASCLVLAARHASS aa seq. of GTLGHPGSLDETTYERLAEETLDSLAEFFEDLADKPYTFE TAT DYDVSFGSGVLTVKLGGDLGTYVINKQTPNKQIWLSSPS MTS(cs) SGPKRYDWTGKNWVYSHDGVSLHELLAAELTKALKTK 81-210 LDLSSLAYSGKDA FRA A linker 34 MRKKRRQRRRWTLGRRAVAGLLASPSPAQAQTLTRVPR aa seq. of PAELAPLCGRRGLRTDIDATCTPRRASSNQRGLNQIWNV TAT KKQSVYLMNLRKSGTLGHPGSLDETTYERLAEETLDSL MTS(fra) AEFFEDLADKPYTFEDYDVSFGSGVLTVKLGGDLGTYVI 81-210 NKQTPNKQIWLSSPSSGPKRYDWTGKNWVYSHDGVSL FRA HELLAAELTKALKTKLDLSSLAYSGKDA .DELTA. linker 35 MRKKRRQRRRQSWSRVYCSLAKRGHFNRISHGLQGLSA aa seq. of VPLRTYASGTLGHPGSLDETTYERLAEETLDSLAEFFEDL TAT ADKPYTFEDYDVSFGSGVLTVKLGGDLGTYVINKQTPN MTS(lad) KQIWLSSPSSGPKRYDWTGKNWVYSHDGVSLHELLAAE 81-210 LTKALKTKLDLSSLAYSGKDA FRA A linker 36 MRKKRRQRRRGALVIRORNFNLENRAEREISKMKPSVA aa seq. of PRHPSSGTLGHPGSLDETTYERLAEETLDSLAEFFEDLAD TAT KPYTFEDYDVSFGSGVLTVKLGGDLGTYVINKQTPNKQI MTS(orf) WLSSPSSGPKRYDWTGKNWVYSHDGVSLHELLAAELT 81-210 KALKTKLDLSSLAYSGKDA FRA .DELTA. linker
[0265] Table 1 above summarizes the nucleic acid sequences (nt) of the primers used as described above (i.e. the primers denoted by SEQ ID NO. 7-SEQ ID NO. 14), the amino acid sequence (aa) of the TAT domain (denoted by SEQ ID NO. 21), the amino acid sequence of a fragment of the TAT domain used in the present disclosure (denoted by SEQ ID NO. 27) and the nucleic acid sequence encoding therefor (denoted by SEQ ID NO. 1), the amino acid sequences of the various MTSs, namely MTS fra, MTS cs, MTS orf and MTS lad (denoted by SEQ ID NO. 22, SEQ ID NO. 23, SEQ ID NO. 25 and SEQ ID NO. 24, respectively) and the nucleic acid sequences encoding therefor (denoted by SEQ ID NO. 2, SEQ ID NO. 3, SEQ ID NO. 4 and SEQ ID NO. 5, respectively), the amino acid sequence of mature frataxin (denoted by SEQ ID NO. 26) and the nucleic acid sequence encoding therefor (denoted by SEQ ID NO. 6), as well as the nucleic acid sequences encoding the OTC protein and the MTS of human mitochondrial GLUD2 (denoted by SEQ ID NO. 15 and SEQ ID NO. 16, respectively). A four-amino acid long linker, having the amino acid sequence of GSDP is also listed in Table 1 above and is denoted by SEQ ID NO. 32.
[0266] In addition Table 1 above indicates the amino acid sequences of the various His TAT MTS 81-210 FRA constructs, which comprise the mature frataxin, namely, His TAT MTS(cs) 81-210 FRA, His TAT MTS(fra) 81-210 FRA (also denoted herein as "HTFrataxin"), His TAT MTS(lad) 81-210 FRA and His TAT MTS(orf) 81-210 FRA (denoted by SEQ ID NO. 17, SEQ ID NO. 18, SEQ ID NO. 19 and SEQ ID NO. 20, respectively). The various His TAT MTS 81-210 FRA constructs denoted by SEQ ID NO. 17, SEQ ID NO. 18, SEQ ID NO. 19 and SEQ ID NO. 20 all comprise a short peptide linker denoted by SEQ ID NO. 32, which joins the TAT and MTS domains.
[0267] Table 1 above further indicates the amino acid sequences of the various fusion constructs constructed without a His tag at their N-termini and with a short peptide linker denoted by SEQ ID NO. 32 (which joins the TAT and MTS domains). These constructs comprise TAT, a linker, MTS and mature frataxin (81-210 FRA), namely TAT MTS(cs) 81-210 FRA, TAT MTS(fra) 81-210 FRA, TAT MTS(lad) 81-210 FRA and TAT MTS(orf) 81-210 FRA, denoted by SEQ ID NO. 28, SEQ ID NO. 29, SEQ ID NO. 30 and SEQ ID NO. 31, respectively.
[0268] In addition Table 1 above indicates the amino acid sequences of the various fusion constructs constructed without a His tag and without a linker. These fusion constructs comprise TAT, MTS and mature frataxin (81-210 FRA), namely TAT MTS(cs) 81-210 FRA A linker, TAT MTS(fra) 81-210 FRA A linker, TAT MTS(lad) 81-210 FRA A linker and TAT MTS(orf) 81-210 FRA A linker, denoted by SEQ ID NO. 33, SEQ ID NO. 34, SEQ ID NO. 35 and SEQ ID NO. 36, respectively.
Proteins Expression and Purification
[0269] E. coli BL21-CodonPlus (.lamda.DE3) or HMS competent cells transformed with plasmids encoding the fusion proteins were incubated at 37.degree. C. in a saline lactose broth (SLB medium) containing kanamycine (50 .mu.g/ml), tetracycline (12.5 .mu.g/ml) and chloramphenicol (34 .mu.g/ml). At an OD.sub.600 of 0.2-0.3, 0.1% glycerol and 0.1 mM potassium glutamate were added to the culture which was then heat-shocked for 20-30 min at 42.degree. C., after which the bacteria were grown at 37.degree. C. until an OD.sub.600 of 0.8. Protein expression was induced by adding isopropyl-beta-D-thiogalactopyranoside (IPTG, for final concentrations see Table 2 below). After 18 hrs of incubation at 22.degree. C., the cells were harvested by centrifugation (500 g for 15 min at 4.degree. C.).
[0270] For the purification procedure, bacteria pellets from 0.5 L culture of expressed cells were sonicated in binding buffer (PBS, pH 7.4, 0.4 M NaCl, 10% Glycerol, 1 mM phenylmethylsulphonylfluoride (PMSF) and 30 mM imidazole (Sigma-Aldrich, St. Louis, Mo., USA)). The suspensions were clarified by centrifugation (35,000 g for 30 min at 4.degree. C.), and the supernatants containing the fusion proteins were purified under native conditions, using binding buffer pre-equilibrated HiTrap Chelating HP columns (Amersham-Pharmacia Biotech, Uppsala, Sweden). Columns were washed by stepwise addition of increasing imidazole concentrations. Finally, the target proteins were eluted with elution buffer (PBS, pH 7.4, 0.4 M NaCl, 10% Glycerol, 250 mM imidazole). All purification procedures were carried out using the FPLC system AKTA (Amersham-Pharmacia Biotech). Imidazole, NaCl and glycerol were removed by transferring the purified proteins to PBS using PD-10 desalting columns (GE Healthcare, Piscataway, N.J., USA). Aliquots of the proteins were kept frozen at -80.degree. C. until use.
TABLE-US-00002 TABLE 2 Expression conditions of TAT-MTS-FRA fusion proteins Heat Temp, for IPTG Protein Bacterial Host Shock induction (.degree. C.) (mM) TAT-MTSfra-FRA codonPlus yes 22 over night 0.5 TAT-MTScs-FRA codonPlus yes 22 over night 0.5 TAT-MTSorf-FRA HMS yes 22 over night 1.0 TAT-MTSlad-FRA HMS yes 22 over night 1.0
Characterization of the Fusion Proteins
Determination of Protein Concentration
[0271] Protein concentration was measured according to the Bradford method, using the Bradford reagent and the standard curve of BSA. Protein concentration was determined at a wavelength of 595 nm.
Separation of Proteins by Electrophoresis
[0272] Samples from the various protein fractions (5-20 .mu.g protein/lane) were loaded on 12% or 15% (w/v) SDS-PAGE gels. The separation of proteins was done using Sturdier Slab Gel Electrophoresis apparatus according to the manufacturer's instructions (Hoefer Sci Instruments, San Francisco, Calif., USA).
Western Blot Analysis
[0273] Proteins (5-20 .mu.g protein/lane) were resolved on 12%-15% SDS-PAGE gels and transferred onto an Immobilon-P Transfer membrane (Millipore, Bradford, Pa., USA). Western blot analysis was performed using either anti-frataxin (Abcam), or anti-His (Amersham-Pharmacia Biotech) antibodies at dilutions of 1:600 and 1:30,000, respectively, to identify the relevant proteins. Primary antibody binding was detected by blotting with a suitable secondary antibody conjugated to horseradish peroxidase (HRP) (1:10,000). Band visualization was done using an enhanced chemiluminescence kit (EZ-ECL, Biological Industries, Beit-Haemek, Israel).
Internalization of TAT-MTS-FRA into the Mitochondria of Patients' Fibroblasts
[0274] FA patient fibroblasts (Coriell repository, #GM03816, termed F816) were thawed and passed at least 2-3 times prior to assay performance and then incubated in the presence of the fusion protein construct TAT-MTS(cs)-FRA (20 .mu.g/ml) comprising frataxin and citrate synthase as the mitochondrial targeting sequence (MTS) for 2, 6 and 48 hours, with fresh addition of TAT-MTS(CS)-FXN (at 20 .mu.g/ml) after 24 hours. Vehicle was added as negative control. Cells were fractionated to cytosolic (C) and mitochondrial (M) fractions using Mitochondria isolation kit (Millipore, MIT1000). All fractions were analyzed in Western blot analysis using a monoclonal anti-frataxin antibody directed against the C-terminal region of frataxin (Abnova, cat. no. H00002395-M02, lot 08078-3F3) diluted 1:500 in blocking solution. Detection was performed using 1 ml at 1 mg/ml of a goat anti mouse IgG-h+1 alkaline phosphatase conjugated antibody (BETHYL, Cat#A90-116AP-16) diluted to 1:20,000 in blocking solution. Signal was developed using the ready-to-use buffered alkaline phosphatase substrate for use in immunoblotting BCIP /NBT-Blue (Sigma, B3804).
Aconitase Activity Assay
[0275] Aconitase activity assay was performed using a commercial kit (Abcam, ab109712, microplate assay kit) according to the manufacturer's instructions. The conversion of isocitrate to cis-aconitate is measured as an increase in absorbance at OD 240 nm at specific time points and calculated as activity rate (mOD/min).
BSO Experiments for Inducing Oxidative Stress
[0276] Normal or patients' lymphocytes (4.times.10.sup.3 cells/100 .mu.l) were seeded in Dulbecco's Modified Eagle Medium (DMEM, Bet Ha'emek, Israel) medium without phenol red and sodium pyruvate (experimental medium) and after 3-4 hr the tested TAT-MTS-FRA fusion protein (at a final concentration of 0.1 .mu.g/.mu.l) was added to the cells for 5 or 24 hr. Following the incubation time with the fusion protein, L-Buthionine-sulfoximine (BSO, Sigma B2640) at different concentrations was added for an additional period of 48 hr. At the end of the incubation time, cell cultures were subjected to cell viability or cell proliferation assays, using the CellTiter-Blue.TM. kit (Promega, Madison, Wis.) according to manufacturer's instructions, as detailed below. In experiments conducted in fibroblasts, cells (3.times.10.sup.3 cells/100 .mu.l) were seeded in the growth medium as specified above and left for 24 hr to allow the cells to adhere. After 24 hr, the medium was changed to the experimental medium and the experiment was continued as described for lymphocytes above.
Cell Proliferation Assay
[0277] 10.sup.4/100 .mu.l/well cells growing in suspension or 5.times.10.sup.3/100 .mu.l/well adherent cells were seeded and treated with increasing concentrations of the fusion protein for 48-72 h, after which cellTiter-Blue.RTM. reagent (Promega, Madison, Wis., USA) was added according to the manufacturer's instructions to determine cell survival. All treatments were performed in triplicates.
In Vitro Caspase 3 Activity Assay
[0278] 10.sup.4/100 .mu.l/well cells growing in suspension or 5.times.10.sup.3/100 .mu.l/well adherent cells were treated with the fusion protein (0.15 .mu.g/.mu.l, final concentration). Caspase3 activity within the cells was assessed by Apo-ONE.RTM. Homogeneous Caspase3/7 Assay Kit (Promega). Caspase 3 activity assays were carried in parallel to cell viability assays.
Preparation of his-TAT-MTS-OTC Constructs
[0279] Fusion protein constructs comprising ornithine transcarbamoylase (OTC) conjugated to a His-TAT-MTS fragment, namely, His-TAT-MTS(otc)-OTC, His-TAT-MTS(lad)-OTC, His-TAT-MTS(cs)-OTC and His-TAT-MTS(orf)-OTC were obtained from GenScript. These fusion protein constructs were prepared by expressing vectors harboring these constructs in E. coli cells and purified from inclusion bodies to about 80% purity using Ni column chromatography.
TABLE-US-00003 TABLE 3 Nucleic acid and amino acid sequences of TAT-MTS-OTC constructs SEQ ID NO. Sequence Name 37 CTGTTTAACCTGCGCATTCTGCTGAACAATGCGGCCTTCCG nt seq. of TAACGGCCATAATTTTATGGTCCGCAACTTCCGTTGCGGTCA MTS of GCCGCTGCAAAATAAAGTGCAG OTC 38 LFNLRILLNNAAFRNGHNFMVRNFRCGQPLQNKVQ aa seq. of MTS of OTC 39 LKGRDLLTLRNFTGEEIKYMLWLSADLKFRIKQKGEYLPLLQ aa seq. of G KSLGMIFEKRSTRTRLSTETGFALLGGHPCFLTTQDIHLG mature VNESLTDTARVLSSMADAVLARVYKQSDLDTLAKEASIPIIN OTC GLSDLYHPIQILADYLTLQEHYSSLKGLTLSWIGDGNNILHS IMMSAAKFGMHLQAATPKGYEPDASVTKLAEQYAKENGTKLL LTNDPLEAAHGGNVLITDTWISMGQEEEKKKRLQAFQGYQVT MKTAKVAASDWTFLHCLPRKPEEVDDEVFYSPRSLVFPEAEN RKWTIMAVMVSLLTDYSPQLQKPKF 40 CATATGGGCT CATCGCATCA TCATCATCAT nt seq. of CACTCATCAG GTCTGGTTCC GCGTGGCTCG His-TAT- CACATGTATG GTCGCAAAAA ACGTCGTCAA MTS(otc)- CGTCGCCGTC TGTTTAACCT GCGCATTCTG OTC CTGAACAATG CGGCCTTCCG TAACGGCCAT .DELTA. linker AATTTTATGG TCCGCAACTT CCGTTGCGGT CAGCCGCTGC AAAATAAAGT GCAGCTGAAA GGCCGCGATC TGCTGACCCT GCGTAACTTC ACGGGTGAAG AAATCAAATA CATGCTGTGG CTGAGCGCAG ACCTGAAATT CCGCATCAAA CAAAAAGGCG AATACCTGCC GCTGCTGCAG GGCAAATCTC TGGGTATGAT TTTTGAAAAA CGTAGTACCC GCACGCGTCT GTCCACCGAA ACGGGCTTTG CCCTGCTGGG CGGTCATCCG TGTTTCCTGA CCACGCAAGA TATCCACCTG GGTGTGAACG AAAGTCTGAC CGATACGGCA CGCGTTCTGA GCTCTATGGC AGACGCTGTG CTGGCTCGTG TTTATAAACA GTCCGATCTG GACACCCTGG CGAAAGAAGC CTCAATTCCG ATTATCAATG GCCTGTCGGA TCTGTACCAT CCGATTCAAA TCCTGGCGGA CTATCTGACC CTGCAGGAAC ACTACAGTTC CCTGAAAGGT CTGACCCTGA GTTGGATCGG CGATGGTAAC AATATTCTGC ATAGCATCAT GATGTCTGCA GCTAAATTTG GCATGCACCT GCAAGCGGCC ACCCCGAAAG GTTATGAACC GGATGCCAGC GTTACGAAAC TGGCAGAACA GTACGCTAAA GAAAACGGTA CCAAACTGCT GCTGACGAAT GATCCGCTGG AAGCAGCTCA TGGCGGTAAC GTCCTGATTA CCGACACGTG GATCTCTATG GGCCAGGAAG AAGAAAAGAA AAAACGTCTG CAGGCGTTTC AAGGTTATCA GGTTACCATG AAAACGGCCA AAGTCGCGGC CAGCGATTGG ACCTTCCTGC ACTGCCTGCC GCGTAAACCG GAAGAAGTCG ATGACGAAGT GTTTTACTCA CCGCGCTCGC TGGTGTTCCC GGAAGCAGAA AATCGTAAAT GGACCATCAT GGCTGTTATG GTGTCCCTGC TGACCGACTA TTCCCCGCAA CTGCAAAAAC CGAAATTCTA ATGAAAGCTT 41 CATATGGGCT CATCTCATCA TCATCATCAT nt seq. of CATTCGTCAG GTCTGGTCCC GCGTGGCTCT His-TAT- CACATGCGTA AAAAACGTCG TCAGCGTCGT MTS(cs)- CGTGGCAGTG ATCCGGCACT GCTGACCGCA OTC GCAGCACGTC TGCTGGGTAC GAAAAACGCT AGCTGCCTGG TGCTGGCTGC GCGTCATGCG TCTGAATTTC TGAAAGGCCG TGATCTGCTG ACCCTGCGCA ACTTCACGGG TGAAGAAATC AAATACATGC TGTGGCTGAG TGCCGACCTG AAATTTCGTA TCAAACAAAA AGGCGAATAC CTGCCGCTGC TGCAGGGCAA ATCCCTGGGT ATGATTTTCG AAAAACGCAG TACCCGTACG CGCCTGTCCA CCGAAACGGG CTTTGCACTG CTGGGCGGTC ATCCGTGTTT CCTGACCACG CAAGATATCC ACCTGGGTGT GAACGAATCA CTGACCGATA CGGCTCGTGT TCTGAGCTCT ATGGCAGACG CAGTGCTGGC ACGTGTTTAT AAACAGTCGG ATCTGGACAC CCTGGCTAAA GAAGCGTCAA TTCCGATTAT CAATGGCCTG TCGGATCTGT ACCATCCGAT TCAAATCCTG GCGGACTATC TGACCCTGCA GGAACACTAC AGTTCCCTGA AAGGTCTGAC CCTGAGCTGG ATCGGCGATG GTAACAATAT TCTGCATAGC ATCATGATGT CTGCCGCAAA ATTTGGCATG CACCTGCAAG CTGCGACCCC GAAAGGTTAT GAACCGGACG CCAGCGTCAC GAAACTGGCC GAACAGTACG CAAAAGAAAA CGGTACCAAA CTGCTGCTGA CGAATGATCC GCTGGAAGCC GCACATGGCG GTAACGTTCT GATTACCGAC ACGTGGATCA GCATGGGCCA GGAAGAAGAA AAGAAAAAAC GTCTGCAGGC CTTTCAAGGT TATCAGGTTA CCATGAAAAC GGCAAAAGTC GCTGCGTCTG ATTGGACCTT CCTGCACTGC CTGCCGCGCA AACCGGAAGA AGTCGATGAC GAAGTGTTTT ACTCACCGCG TTCGCTGGTT TTCCCGGAAG CGGAAAATCG CAAATGGACC ATTATGGCTG TGATGGTCTC TCTGCTGACG GACTACTCGC CGCAACTGCA AAAACCGAAA TTCTAATGAA AGCTT 42 CATATGGGTT CATCACATCA TCATCATCAT nt seq. of CATTCATCAG GTCTGGTCCC GCGTGGTTCA His-TAT- CACATGCGTA AAAAACGTCG TCAGCGTCGT MTS(orf)- CGTGGCAGTG ATCCGGGTGC GCTGGTCATT OTC CGTGGCATCC GCAACTTTAA TCTGGAAAAC CGTGCGGAAC GCGAAATTAG TAAAATGAAA CCGTCCGTGG CACCGCGTCA TCCGTCTGAA TTTCTGAAAG GCCGTGATCT GCTGACCCTG CGCAACTTCA CGGGTGAAGA AATCAAATAC ATGCTGTGGC TGAGTGCAGA CCTGAAATTC CGTATCAAAC AAAAGGGTGA ATACCTGCCG CTGCTGCAGG GCAAATCCCT GGGTATGATT TTCGAAAAAC GCTCAACCCG TACGCGCCTG TCGACCGAAA CGGGCTTTGC CCTGCTGGGC GGTCATCCGT GCTTCCTGAC CACGCAAGAT ATCCACCTGG GTGTGAACGA ATCACTGACC GATACGGCAC GTGTTCTGAG CTCTATGGCA GACGCAGTGC TGGCTCGTGT TTATAAACAG TCGGATCTGG ACACCCTGGC AAAAGAAGCT AGCATTCCGA TTATCAATGG CCTGTCTGAT CTGTACCATC CGATTCAAAT CCTGGCGGAC TATCTGACCC TGCAGGAACA CTACAGTTCC CTGAAAGGTC TGACCCTGAG CTGGATCGGC GATGGTAACA ATATTCTGCA TAGCATCATG ATGTCTGCGG CCAAATTCGG CATGCACCTG CAAGCAGCTA CCCCGAAAGG TTATGAACCG GACGCCTCCG TTACGAAACT GGCGGAACAG TACGCCAAAG AAAACGGCAC CAAACTGCTG CTGACGAATG ATCCGCTGGA AGCGGCCCAT GGCGGTAACG TCCTGATTAC CGACACGTGG ATCAGCATGG GCCAGGAAGA AGAAAAGAAA AAACGTCTGC AGGCATTTCA AGGTTATCAG GTTACCATGA AAACGGCTAA AGTCGCAGCT TCTGATTGGA CCTTCCTGCA CTGTCTGCCG CGCAAACCGG AAGAAGTCGA TGACGAAGTG TTTTACTCAC CGCGTTCGCT GGTGTTCCCG GAAGCGGAAA ATCGCAAATG GACCATTATG GCTGTGATGG TGTCGCTGCT GACGGACTAC TCGCCGCAAC TGCAAAAACC GAAATTCTAA TGAAAGCTT 43 CATATGGGTA GTTCACATCA TCATCATCAT nt seq. of CACTCGTCGG GTCTGGTGCC GCGTGGCTCA His-TAT- CACATGCGTA AAAAACGTCG TCAGCGTCGT MTS(lad)- CGTGGCTCAG ATCCGCAATC ATGGTCGCGC OTC GTCTATTGCT CGCTGGCGAA ACGTGGTCAT TTTAACCGCA TTAGCCACGG CCTGCAGGGT CTGTCTGCAG TGCCGCTGCG TACCTACGCT GAATTTCTGA AAGGCCGTGA TCTGCTGACC CTGCGCAACT TCACGGGTGA AGAAATCAAA TACATGCTGT GGCTGAGCGC AGACCTGAAA TTTCGTATCA AACAAAAAGG CGAATACCTG CCGCTGCTGC AGGGCAAATC TCTGGGTATG ATTTTCGAAA AACGCTCAAC CCGTACGCGC CTGTCGACCG AAACGGGCTT TGCCCTGCTG GGCGGTCATC CGTGTTTCCT GACCACGCAG GATATCCACC TGGGTGTGAA CGAAAGTCTG ACCGATACGG CACGTGTTCT GAGCTCTATG GCAGACGCAG TGCTGGCTCG TGTTTATAAA CAGTCCGATC TGGACACCCT GGCAAAAGAA GCTAGTATTC CGATTATCAA TGGCCTGTCC GATCTGTACC ATCCGATTCA AATCCTGGCG GACTATCTGA CCCTGCAGGA ACACTACAGT TCCCTGAAAG GTCTGACGCT GAGCTGGATC GGCGATGGTA ACAATATTCT GCATAGTATC ATGATGTCCG CGGCCAAATT CGGCATGCAC CTGCAAGCAG CTACCCCGAA AGGTTATGAA CCGGACGCCT CTGTTACGAA ACTGGCGGAA CAGTACGCCA AAGAAAACGG TACCAAACTG CTGCTGACGA ATGATCCGCT GGAAGCGGCC CATGGCGGTA ACGTCCTGAT TACCGACACG TGGATCAGTA TGGGCCAGGA AGAAGAAAAG AAAAAACGTC TGCAGGCGTT TCAAGGTTAT CAGGTTACCA TGAAAACGGC CAAAGTCGCA GCTAGCGATT GGACCTTCCT GCACTGCCTG CCGCGCAAAC CGGAAGAAGT CGATGACGAA GTGTTTTATA GCCCGCGTTC TCTGGTGTTC CCGGAAGCGG AAAATCGCAA ATGGACCATC ATGGCCGTTA TGGTGTCGCT GCTGACCGAT TACTCCCCGC AACTGCAAAA ACCGAAATTC TAATGAAAGC TT 44 MGSSHHHHHH SSGLVPRGSH MYGRKKRRQR aa seq. of RRLFNLRILL NNAAFRNGHN FMVRNFRCGQ His-TAT- PLQNKVQLKG RDLLTLRNFT GEEIKYMLWL MTS(otc)- SADLKFRIKQ KGEYLPLLQG KSLGMIFEKR OTC STRTRLSTET GFALLGGHPC FLTTQDIHLG .DELTA. linker VNESLTDTAR VLSSMADAVL ARVYKQSDLD TLAKEASIPI INGLSDLYHP IQILADYLTL QEHYSSLKGL TLSWIGDGNN ILHSIMMSAA KFGMHLQAAT PKGYEPDASV TKLAEQYAKE NGTKLLLTND PLEAAHGGNV LITDTWISMG QEEEKKKRLQ AFQGYQVTMK TAKVAASDWT FLHCLPRKPE EVDDEVFYSP RSLVFPEAEN RKWTIMAVMV SLLTDYSPQL QKPKF 45 MGSSHHHHHH SSGLVPRGSH MRKKRRQRRR aa seq. of GSDPALLTAA ARLLGTKNAS CLVLAARHAS His-TAT- EFLKGRDLLT LRNFTGEEIK YMLWLSADLK MTS(cs)- FRIKQKGEYL PLLQGKSLGM IFEKRSTRTR OTC LSTETGFALL GGHPCFLTTQ DIHLGVNESL TDTARVLSSM ADAVLARVYK QSDLDTLAKE ASIPIINGLS DLYHPIQILA DYLTLQEHYS SLKGLTLSWI GDGNNILHSI MMSAAKFGMH LQAATPKGYE PDASVTKLAE QYAKENGTKL LLTNDPLEAA HGGNVLITDT WISMGQEEEK KKRLQAFQGY QVTMKTAKVA ASDWTFLHCL PRKPEEVDDE VFYSPRSLVF PEAENRKWTI MAVMVSLLTD YSPQLQKPKF 46 MGSSHHHHHH SSGLVPRGSH MRKKRRQRRR aa seq. of GSDPGALVIR GIRNFNLENR AEREISKMKP His-TAT- SVAPRHPSEF LKGRDLLTLR NFTGEEIKYM MTS(orf)- LWLSADLKFR IKQKGEYLPL LQGKSLGMIF OTC EKRSTRTRLS TETGFALLGG HPCFLTTQDI HLGVNESLTD TARVLSSMAD AVLARVYKQS DLDTLAKEAS IPIINGLSDL YHPIQILADY LTLQEHYSSL KGLTLSWIGD GNNILHSIMM SAAKFGMHLQ AATPKGYEPD ASVTKLAEQY AKENGTKLLL TNDPLEAAHG GNVLITDTWI SMGQEEEKKK RLQAFQGYQV TMKTAKVAAS DWTFLHCLPR KPEEVDDEVF YSPRSLVFPE AENRKWTIMA VMVSLLTDYS PQLQKPKF 47 MGSSHHHHHH SSGLVPRGSH MRKKRRQRRR aa seq. of GSDPQSWSRV YCSLAKRGHF NRISHGLQGL His-TAT- SAVPLRTYAE FLKGRDLLTL RNFTGEEIKY MTS(lad)- MLWLSADLKF RIKQKGEYLP LLQGKSLGMI OTC FEKRSTRTRL STETGFALLG GHPCFLTTQD IHLGVNESLT DTARVLSSMA DAVLARVYKQ SDLDTLAKEA SIPIINGLSD LYHPIQILAD YLTLQEHYSS LKGLTLSWIG DGNNILHSIM MSAAKFGMHL QAATPKGYEP DASVTKLAEQ YAKENGTKLL LTNDPLEAAH GGNVLITDTW ISMGQEEEKK KRLQAFQGYQ VTMKTAKVAA SDWTFLHCLP RKPEEVDDEV FYSPRSLVFP EAENRKWTIM AVMVSLLTDY SPQLQKPKF 48 RKKRRQRRRL FNLRILLNNA AFRNGHNFMV aa seq. of RNFRCGQPLQ NKVQLKGRDL LTLRNFTGEE TAT- IKYMLWLSAD LKFRIKQKGE YLPLLQGKSL MTS(otc)- GMIFEKRSTR TRLSTETGFA LLGGHPCFLT OTC TQDIHLGVNE SLTDTARVLS SMADAVLARV .DELTA. linker YKQSDLDTLA KEASIPIING LSDLYHPIQI LADYLTLQEH YSSLKGLTLS WIGDGNNILH SIMMSAAKFG MHLQAATPKG YEPDASVTKL AEQYAKENGT KLLLTNDPLE AAHGGNVLIT DTWISMGQEE EKKKRLQAFQ GYQVTMKTAK
VAASDWTFLH CLPRKPEEVD DEVFYSPRSL VFPEAENRKW TIMAVMVSLL TDYSPQLQKPKF 49 MRKKRRQRRR GSDPALLTAA ARLLGTKNAS aa seq. of CLVLAARHAS EFLKGRDLLT LRNFTGEEIK TAT- YMLWLSADLK FRIKQKGEYL PLLQGKSLGM MTS(cs)- IFEKRSTRTR LSTETGFALL GGHPCFLTTQ OTC DIHLGVNESL TDTARVLSSM ADAVLARVYK QSDLDTLAKE ASIPIINGLS DLYHPIQILA DYLTLQEHYS SLKGLTLSWI GDGNNILHSI MMSAAKFGMH LQAATPKGYE PDASVTKLAE QYAKENGTKL LLTNDPLEAA HGGNVLITDT WISMGQEEEK KKRLQAFQGY QVTMKTAKVA ASDWTFLHCL PRKPEEVDDE VFYSPRSLVF PEAENRKWTI MAVMVSLLTD YSPQLQKPKF 50 MRKKRRQRRR GSDPGALVIR GIRNFNLENR aa seq. of AEREISKMKP TAT- SVAPRHPSEF LKGRDLLTLR NFTGEEIKYM MTS(orf)- LWLSADLKFR IKQKGEYLPL LQGKSLGMIF OTC EKRSTRTRLS TETGFALLGG HPCFLTTQDI HLGVNESLTD TARVLSSMAD AVLARVYKQS DLDTLAKEAS IPIINGLSDL YHPIQILADY LTLQEHYSSL KGLTLSWIGD GNNILHSIMM SAAKFGMHLQ AATPKGYEPD ASVTKLAEQY AKENGTKLLL TNDPLEAAHG GNVLITDTWI SMGQEEEKKK RLQAFQGYQV TMKTAKVAAS DWTFLHCLPR KPEEVDDEVF YSPRSLVFPE AENRKWTIMA VMVSLLTDYS PQLQKPKF 51 MRKKRRQRRR GSDPQSWSRV YCSLAKRGHF aa seq. of NRISHGLQGL TAT- SAVPLRTYAE FLKGRDLLTL RNFTGEEIKY MTS(lad)- MLWLSADLKF RIKQKGEYLP LLQGKSLGMI OTC FEKRSTRTRL STETGFALLG GHPCFLTTQD IHLGVNESLT DTARVLSSMA DAVLARVYKQ SDLDTLAKEA SIPIINGLSD LYHPIQILAD YLTLQEHYSS LKGLTLSWIG DGNNILHSIM MSAAKFGMHL QAATPKGYEP DASVTKLAEQ YAKENGTKLL LTNDPLEAAH GGNVLITDTW ISMGQEEEKK KRLQAFQGYQ VTMKTAKVAA SDWTFLHCLP RKPEEVDDEV FYSPRSLVFP EAENRKWTIM AVMVSLLTDY SPQLQKPKF 52 MRKKRRQRRR ALLTAA ARLLGTKNAS CLVLAARHAS aa seq. of EFLKGRDLLT LRNFTGEEIK YMLWLSADLK TAT- FRIKQKGEYL PLLQGKSLGM IFEKRSTRTR MTS(cs)- LSTETGFALL GGHPCFLTTQ DIHLGVNESL OTC TDTARVLSSM ADAVLARVYK QSDLDTLAKE linker ASIPIINGLS DLYHPIQILA DYLTLQEHYS SLKGLTLSWI GDGNNILHSI MMSAAKFGMH LQAATPKGYE PDASVTKLAE QYAKENGTKL LLTNDPLEAA HGGNVLITDT WISMGQEEEK KKRLQAFQGY QVTMKTAKVA ASDWTFLHCL PRKPEEVDDE VFYSPRSLVF PEAENRKWTI MAVMVSLLTD YSPQLQKPKF 53 MRKKRRQRRR GALVIR GIRNFNLENR AEREISKMKP aa seq. of SVAPRHPSEF LKGRDLLTLR NFTGEEIKYM TAT- LWLSADLKFR IKQKGEYLPL LQGKSLGMIF MTS(orf)- EKRSTRTRLS TETGFALLGG HPCFLTTQDI OTC .DELTA. HLGVNESLTD TARVLSSMAD AVLARVYKQS linker DLDTLAKEAS IPIINGLSDL YHPIQILADY LTLQEHYSSL KGLTLSWIGD GNNILHSIMM SAAKFGMHLQ AATPKGYEPD ASVTKLAEQY AKENGTKLLL TNDPLEAAHG GNVLITDTWI SMGQEEEKKK RLQAFQGYQV TMKTAKVAAS DWTFLHCLPR KPEEVDDEVF YSPRSLVFPE AENRKWTIMA VMVSLLTDYS PQLQKPKF 54 MRKKRRQRRR QSWSRV YCSLAKRGHF NRISHGLQGL aa seq. of SAVPLRTYAE FLKGRDLLTL RNFTGEEIKY TAT- MLWLSADLKF RIKQKGEYLP LLQGKSLGMI MTS(lad)- FEKRSTRTRL STETGFALLG GHPCFLTTQD OTC .DELTA. IHLGVNESLT DTARVLSSMA DAVLARVYKQ linker SDLDTLAKEA SIPIINGLSD LYHPIQILAD YLTLQEHYSS LKGLTLSWIG DGNNILHSIM MSAAKFGMHL QAATPKGYEP DASVTKLAEQ YAKENGTKLL LTNDPLEAAH GGNVLITDTW ISMGQEEEKK KRLQAFQGYQ VTMKTAKVAA SDWTFLHCLP RKPEEVDDEV FYSPRSLVFP EAENRKWTIM AVMVSLLTDY SPQLQKPKF
[0280] Table 3 recites the amino acid sequence of the MTS of the OTC protein and the nucleotide sequence encoding therefor (denoted by SEQ ID NO. 38 and by SEQ ID NO. 37, respectively), the amino acid sequence of mature OTC (denoted by SEQ ID NO. 39), and the nucleotide sequences encoding the fusion protein constructs His-TAT-MTS(otc)-OTC .DELTA. linker, His-TAT-MTS(cs)-OTC, His-TAT-MTS(orf)-OTC and His-TAT-MTS(lad)-OTC (denoted by SEQ ID NO. 40, denoted by SEQ ID NO. 41, SEQ ID NO. 42 and by SEQ ID NO. 43, respectively). Table 3 also indicates the amino acid sequences of the fusion protein constructs His-TAT-MTS(otc)-OTC .DELTA. linker, His-TAT-MTS(cs)-OTC, His-TAT-MTS(orf)-OTC and His-TAT-MTS(lad)-OTC (denoted by SEQ ID NO. 44, SEQ ID NO. 45, SEQ ID NO. 46 and by SEQ ID NO. 47, respectively).
[0281] Also recited are fusion protein constructs comprising the mitochondrial active protein OTC where the fusion protein construct does not include a His tag, namely the constructs TAT-MTS(otc)-OTC .DELTA. linker having the amino acid sequences denoted by SEQ ID NO. 48, TAT-MTS(cs)-OTC (denoted by SEQ ID NO. 49), TAT-MTS(orf)-OTC (denoted by SEQ ID NO. 50) and TAT-MTS(lad)-OTC (denoted by SEQ ID NO. 51).
[0282] The fusion protein constructs according to the present disclosure may be prepared in the presence of a linker situated between the TAT and the MTS fragments or in its absence. Thus table 3 also recites the sequences of the fusion protein constructs termed His-TAT-MTS(otc)-OTC .DELTA. linker, TAT-MTS(cs)-OTC .DELTA. linker, TAT-MTS(orf)-OTC .DELTA. linker, TAT-MTS(lad)-OTC .DELTA. linker (having the amino acid sequences denoted by SEQ ID NO. 44, SEQ ID NO. 52, SEQ ID NO. 53 and SEQ ID NO. 54, respectively).
Internalization of his-TAT-MTS-OTC Constructs into the Mitochondria
[0283] A culture of 10.sup.7 HepG2 cells (ATCC, #HB-8065) was seeded in T-75 flasks. The next day, TAT-MTS-OTC fusion protein constructs comprising either OTC, CS or LAD at 12 .mu.g/ml prepared as described above were added to the cells in Eagle's Minimum Essential Medium (EMEM) complete medium, or in EMEM medium supplemented with DMSO (1%) and Trehalose (30 mM) or in EMEM medium supplemented with DMSO (1%), Trehalose (30 mM) and Ornithine (1 mM) for an incubation of 20 min, 1 or 3 hours. Cells were then fractionated to cytosolic and mitochondrial fractions using Mitochondria isolation kit (Millipore, MIT1000) according to the manufacturer's instructions. Fractions were analyzed in Western blot analysis using a monoclonal antibody directed to the C terminal region of OTC (Aviva Systems Biology, cat. no. ARP41767_P050), at 1 mg/ml diluted 1:1000. Detection was performed with the goat anti rabbit IgG (H_L) secondary antibody peroxidase conjugated (Jackson AffiniPure, Code 111-035-003) at 0.8 mg/ml, diluted 1:20,000 in blocking buffer ( ). Signal was developed with chemiluminescence mixture (SC-2048 Santa Cruz) according to the manufacturer's instructions.
In Vitro Production of Citrulline by the Enzymatic Activity of OTC
[0284] Purified TAT-MTS-OTC fusion protein constructs comprising the MTS of OTC or CS prepared as described above were tested for their enzymatic activity as described in [40]. Briefly, proteins were diluted in a mitochondria lysis buffer (0.5% Triton, 10 mM HEPES, pH 7.2 and 2 mM dithiothreitol) to a concentration of 20 .mu.g in a volume of 80 .mu.l. Then the volume of each of the reaction mixtures was completed to 600 .mu.l by adding 520 .mu.l reaction mixture (5 mM ornithine, 15 mM carbamoyl phosphate, and 270 mM triethanolamine, pH 7.7). Following vortexing, each reaction mixture was divided into two separate tubes, the first tube was incubated at 37.degree. C. for 30 min and the second tube served as "no-reaction" background and used to subtract endogenous signal of OTC proteins. Reactions were terminated by adding 150 .mu.l of 1:3 sulfuric acid/phosphoric acid (by volume). Citrulline production was then determined by adding 20 .mu.l of 3% 2,3-butanedione monoxime, incubating at 100.degree. C. in the dark for 15 min, and measuring absorbance at 490 nm. Commercially available OTC from Bacillus subtilis (Sigma, 5 ng) was used as Positive Control (data not shown).
Enzymatic Activity of OTC in HepG2 Cells in the Presence of Ammonia
[0285] HepG2 cells 2.5.times.10.sup.5 (ATCC, #HB-8065) per well were seeded in two 6-well plates. The next day, TAT-MTS-OTC fusion protein constructs comprising the MTS of OTC or CS (prepared as described above) were applied to the cells for 72 hours with replenishment after each 24 hours. The final concentration of the fusion proteins was 14 .mu.g/ml, in 3 ml complete medium and the final buffer dilution was 1:85. Proteins storage buffer (1.times.PBS, 0.5% Sodium Lauroyl Sarcosine at pH 7.4) served as vehicle control and untreated cells as negative control. At 48 hours prior to the termination of the assay, cells were treated with 0.5% serum growth medium in the absence or in the presence of 5 mM Ammonium chloride. Cells viability was monitored for 4 hours with alamarBlue Cell Viability Assay Reagent (Pierce). Fluorescence signal was obtained by excitation at 544 nm and emission at 590 nm.
Example 1
Cloning of Plasmids Encoding TA T-MTS-FRA Fusion Proteins
[0286] Expression plasmids encoding the TAT-fusion proteins were cloned and prepared by standard molecular biology tools known in the art. For a general reference see Molecular Cloning: A Laboratory manual (2001) Joseph Sambrook and David William Russell. TAT [9] having the amino acid sequence as denoted by SEQ ID NO. 27 (encoded by the nucleic acid sequence denoted by SEQ ID NO. 1) was fused (N-terminal) to the mature human frataxin protein (having the amino acid sequence as denoted by SEQ ID NO. 26 and encoded by the nucleic acid sequence as denoted by SEQ ID NO. 6).
[0287] Various fusion constructs were prepared, which differ in their mitochondrial targeting sequence present at the N terminus of human frataxin (and thus located between the TAT and mature human frataxin described above), being either the native mitochondrial targeting sequence of frataxin (referred to herein as MTSfra, having the amino acid sequence as denoted by SEQ ID NO. 22 and encoded by the nucleic acid sequence as denoted by SEQ ID NO. 2) or other defined MTSs of human mitochondrial proteins, including lipoamide dehydrogenase (referred to herein as MTSlad, having the amino acid sequence as denoted by SEQ ID NO. 24 and encoded by the nucleic acid sequence as denoted by SEQ ID NO. 5), C6ORF66 (referred to herein as MTSorf, having the amino acid sequence as denoted by SEQ ID NO. 25 and encoded by the nucleic acid sequence as denoted by SEQ ID NO. 4) and citrate synthase (referred to herein as MTScs, having the amino acid sequence as denoted by SEQ ID NO. 23 and encoded by the nucleic acid sequence as denoted by SEQ ID NO. 3). The various TAT-MTS-FRA fusion proteins are summarized in Table 4 below and schematically presented in FIG. 1.
[0288] All plasmids were cloned with His-tag at the 5'-terminus of the coding sequence and all coding sequences were under the control of the T7 promotor. All clones were confirmed by restriction enzymes and sequencing analyses.
TABLE-US-00004 TABLE 4 The cloned plasmid constructs No. Plasmid name Abbreviated name 1 His-TAT-MTSfra-FRA FRA 2 His-TAT-MTSlad-FRA (lad)FRA 3 His-TAT-MTSorf-FRA (orf)FRA 4 His-TAT-MTScs-FRA (cs)FRA Abbreviations: lad, Lipoamide dehydrogenase protein (E3 subunit); orf, C6ORF66 assembly factor; and cs, citrate synthase.
[0289] As indicated above, the amino acid sequences of the fusion proteins obtained from the cloned plasmid constructs indicated as 1-4 in Table 4 above are denoted by SEQ ID NO. 18, SEQ ID NO. 19, SEQ ID NO. 20 and SEQ ID NO. 17, respectively.
Example 2
[0290] Expression of the Fusion Proteins in E. coli Hosts
[0291] Expression of the fusion proteins was performed in E. coli hosts as described below and was calibrated for optimal expression conditions. As known in the art, there are several different bacterial expression systems. Successful expression of recombinant proteins is often dependent on the strain of the bacteria expression system used. Thus, for each fusion protein prepared as described above, four different E. coli bacterial stains were tested: BL21-CodonPlus, BL21, Rosetta and HMS (Invitrogen, USA) and the host for expression was thereby selected. The conditions for expression were also calibrated for each of the TAT-fusion proteins, by changing several parameters, including the concentration of the inducer Isopropyl .beta.-D-1-thiogalactopyranoside (IPTG) and length of induction growth conditions (i.e. temperature, addition of chemicals, etc.)
[0292] Upon expression, bacterial cells were disrupted and cellular sub-fractions were prepared, separating the soluble and non-soluble fractions. Analysis was performed for the whole-cell bacteria (W.C. or whole-cell extract), the soluble fraction (Sol) and insoluble fraction (Insol) on sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gels in order to examine whether the fusion protein was expressed and at which sub-cellular fraction it accumulates. The goal was to obtain high expression levels of the different TAT-fusion proteins in the soluble sub-fraction of the expressing bacteria, for future purification. The different TAT-fusion proteins were also characterized by Western blots analyses using both anti-His and anti-frataxin antibodies. Table 2 above summarizes the bacterial host, IPTG concentration and temperature for production of each TAT-MTS-FRA fusion proteins carrying a different MTS sequence.
[0293] Typical expression and sub-cellular localization of each of the four fusion proteins as characterized by SDS-PAGE gels and Western blots using anti-His antibodies are demonstrated in FIG. 2 and in FIG. 3. Expression of the His-TAT-MTSfra-FRA fusion protein is shown in FIG. 2A-2B, Expression of the His-TAT-MTSorf-FRA fusion protein is shown in FIG. 2C-2D, Expression of the His-TAT-MTSlad-FRA fusion protein is shown in FIG. 3A-3B and Expression of the His-TAT-MTScs-FRA fusion protein is shown in FIG. 3C-3D. These experiments confirmed the full-length expression of the different TAT-fusion proteins and their identity.
[0294] It should be pointed out the anti-His antibodies recognize, most probably, the various fusion proteins with different efficacy, depending on the exposure or availability of the His sequence in the final protein preparation, for antibody interactions. Thus, expression levels of the various fusion proteins are determined based on the SDS-PAGE gels.
[0295] As can be seen in FIG. 2 and in FIG. 3, the TAT-MTSfra-FRA fusion protein (FIG. 2A & FIG. 2B) was expressed at low levels in the bacterial hosts, as compared to the other three fusion proteins carrying an heterologous MTS (FIG. 2C & FIG. 2D and FIG. 3A-3D), even under the best calibrated conditions. Thus, using a heterologous MTS instead of the native frataxin-MTS has an advantage in its expression levels in a bacterial host. This has major implications on its future development for human use, where large quantities of the fusion protein are needed to be produced (see also below).
Example 3
Purification of the TA T-MTS-FRA Fusion Proteins
[0296] The soluble fractions of the expressed TAT-MTS-FRA fusion proteins were loaded onto a nickel-chelating column to affinity-purify these proteins, as detailed above. Calibration experiments were performed for each of the fusion proteins, including specific conditions for binding of the fusion protein onto the affinity column, its elution and removal of the imidazole from the final protein preparations. One typical purification run of each of the fusion proteins is demonstrated in FIG. 4 (for His-TAT-MTSfra-FRA and His-TAT-MTSorf-FRA) and in FIG. 5 (for His-TAT-MTSlad-FRA and His-TAT-MTScs-FRA).
[0297] For example, FIG. 4A shows an image of an affinity chromatography purification profile obtained for the fusion protein TAT-MTSfra-FRA. As can be seen in FIG. 4B, which is an image of SDS-PAGE analysis of the purification steps of this protein construct, the protein fraction shown in FIG. 4B, lane 7, represents the fusion protein TAT-MTSfra-FRA that was eluted from the column at 250 mM imidazole.
[0298] Corresponding analyses are shown for TAT-MTSorf-FRA in FIG. 4C and FIG. 4D. for TAT-MTSlad-FRA in FIG. 5A and FIG. 5B and for TAT-MTScs-FRA corresponding analyses are shown in FIG. 5C and FIG. 5D.
[0299] As shown in FIG. 6A, eluted proteins showed a major band of the expected size (approximately 20-27 kDa) and were >95% pure, as determined by SDS-PAGE analysis and by Western blot analyses using both anti-His (FIG. 6B) and anti-Fra antibodies (FIG. 6C). As can be seen in FIG. 6, TAT-MTS-FRA fusion proteins carrying a heterologous MTS were full-length, intact proteins with no evidence for protein degradation (see lane 2 for TAT-MTScs-FRA, lane 3 for TAT-TSlad-FRA and lane 4 for TAT-MTSorf-FRA). However, TAT-MTSfra-FRA carrying the native MTS sequence was partially degraded (see lane 1 in FIG. 6B and in FIG. 6C). Thus, using a heterologous MTS sequence for the TAT-FRA fusion protein has an advantage also for keeping the fusion protein intact and stable.
[0300] In addition, when comparing the total amounts and concentrations of each of the fusion proteins, which were produced from the same starting volume of bacterial cultures, fusion proteins carrying an heterologous MTS were produced in larger amounts and at higher concentration as compared to that of fusion protein carrying the native MTS (Table 5, below). This has again major implications on its future development for human use, where large quantities of the fusion protein are needed and at high concentrations. Moreover, the stability of the produced fusion protein has an additional advantage.
TABLE-US-00005 TABLE 5 TAT-MTS-FRA fusion proteins Final concentration Amount purified from 0.5 L Protein (mg/ml) bacterial culture (mg) TAT-MTSfra-FRA 0.2 0.6-0.7 TAT-MTScs-FRA 1.0 3-4 TAT-MTSorf-FRA 1.0 3-4 TAT-MTSlad-FRA 0.8 2.4-3.2
Example 4
Internalization of TA T-MTS-FRA Fusion Proteins
[0301] In order to test the ability of the fusion protein to reach the mitochondria within intact cells, human BJAB cells were incubated with one of the purified fusion proteins, namely, TAT-MTSlad-FRA. After incubation, sub-cellular fractions were prepared. to separate the mitochondria and the cytosol. The mitochondria were then treated with proteinase K (Sigma P2308) to digest proteins nonspecifically adsorbed to the outer membrane, thereby ensuring that the mitochondrial extract represented only proteins within the mitochondria. Samples were then analyzed by Western blot assay for the presence of the FRA-based fusion protein, using both anti-His and anti-FRA antibodies (see FIG. 7).
[0302] As shown in FIG. 7A, using anti-His antibodies the results indicated the presence of the His-tagged TAT-MTSlad-FRA fusion protein within the mitochondrial fractions of the treated cells after 1 and 5 hours of incubation (lane 4 and lanes 6 & 7, respectively).
[0303] As shown in FIG. 7B, using anti-FRA antibodies, the presence of a frataxin fusion protein construct inside the mitochondria is indicated. From lane 2 of FIG. 7B it is evident that control cells that were not treated with the fusion protein have endogenous FRA protein, most probably, both the mature isoform and the intermediate isoform (indicated by the two arrows).
[0304] As shown in FIG. 7B (lanes 4. 6, 7), upon treatment with the TAT-MTSlad-FRA fusion protein, the relative amounts of both the unprocessed fusion protein as well as of the mature protein increased with the incubation time (FIG. 7B; upper and lower bands, respectively). Without wishing to be bound by theory, the increase in the upper band is explained by fusion protein construct that entered the mitochondria and was not processed.
[0305] Internalization of fusion proteins comprising frataxin into the mitochondria of cells was examined also for other TAT-MTS-FRA fusion proteins, namely, TAT-MTSfra-FRA, TAT-MTScs-FRA, and TAT-MTSorf-FRA (as well as for TAT-MTSlad-FRA), as shown in FIG. 8A. In this experiment, cells were incubated for 3 hours with each of the TAT-MTS-FRA fusion proteins (at a final concentration of 0.02 .mu.g/.mu.l). The cells were then washed, their mitochondria were isolated and then submitted to Western blot analysis using monoclonal anti-FRA antibodies (Abnova, Taiwan; 1 .mu.g/ml) that are directed to the C-terminus of the frataxin protein.
[0306] As demonstrated in FIG. 8A, lanes 2-5, a slight size variation was observed among the various fusion protein constructs, resulting from the variation in the length of the various MTS polypeptides (see FIG. 1).
[0307] As shown in FIG. 8A, internalization into mitochondria was observed for all four TAT-MTS-FRA fusion proteins. Remarkably, as shown in lane 3 of FIG. 8A, among the fusion proteins with a MTS that is heterologous to frataxin, the citrate synthase MTS (MTScs) was demonstrated to be delivered most efficiently into the mitochondria.
[0308] As indicated above, there is a slight variation in the lengths of the fusion protein constructs that entered the mitochondria, indicating that under the present experimental conditions (namely incubation of cells for 3 hours with each of the TAT-MTS-FRA fusion proteins) the frataxin protein was not cleaved from its respective His-TAT-MTS-frataxin construct inside the mitochondria (protein constructs that still include intact His-TAT-MTS-frataxin are termed below as "un-processed fusion construct").
[0309] Notably, FIG. 8A also shows that while for the fusion proteins comprising an MTS which is heterologous to frataxin, namely the MTS of citrate synthase (cs), lipoamide dehydrogenase (lad) and C6ORF66 (orf), only a single band (representing the un-processed fusion construct) was demonstrated inside the mitochondria, two distinct bands appeared for the fusion protein comprising the native MTS of frataxin (FIG. 8A, lane 2). Without wishing to be bound by theory, this may be the result of instability of the intact fusion protein comprising the TAT and native frataxin MTS regions.
[0310] The antibodies used to detect frataxin in the internalization assay presented in FIG. 7B were polyclonal antibodies directed against human frataxin. These antibodies recognize the full-length, unprocessed human frataxin very well, as well as its processed products. In FIG. 8 referred to below, detection was performed using monoclonal antibodies specifically directed against the C-terminus of the frataxin protein. As these monoclonal antibodies are less efficient in recognizing the protein, particularly at the amounts of protein detected in FIG. 8, the results of FIG. 7 and FIG. 8 cannot be compared.
[0311] Control Western blot analysis using anti-E1.alpha. antibodies (Molecular Probes, Eugene, Oreg.) at a dilution of 1:1,000) are demonstrated in FIG. 8B.
Example 5
His-TA T-MTS(Cs)-FRA Enter the Mitochondria of Patients' Flbroblasts
[0312] Further to the above results demonstrating the ability of the frataxin fusion proteins to enter mitochondria of human BJAB cells, the internalization of a frataxin fusion protein comprising the MTS of citrate synthase into the mitochondria of fibroblasts obtained from Friedreich's ataxia patients was examined, as detailed below.
[0313] Fibroblasts obtained from Friedreich's ataxia patients were treated as described above and incubated in the presence of His-TAT-MTS(cs)-FRA (at 20 .mu.g/ml) for 2, 6 and 48 hours, with fresh addition after 24 hours. Mitochondrial and cytosolic fractions of the fibroblasts were analyzed by Western blot analysis as described above.
[0314] As demonstrated in FIG. 9, the fusion protein construct His-TAT-MTS(cs)-FRA was detected in mitochondrial fractions of fibroblasts after an incubation of 2, 6 and 48 hours. The two bands detected by the anti-frataxin antibody (indicated by the upper and lower arrows in FIG. 9) indicate that the fusion protein was processed inside mitochondria. Frataxin was not detected in the cytosolic fractions of the fibroblasts indicating that all of the His-TAT-MTS(cs)-FRA fusion protein construct has entered mitochondria.
Example 6
[0315] Aconitase Activity in Fibroblasts Obtained from Patients Following Administration of his-TAT-MTS(Cs)-FRA
[0316] It has been reported that the reduction in the levels of frataxin within the mitochondria has two direct effects on several tissue types, namely impaired formation of iron-sulfur (Fe--S) clusters and a rise in intracellular reactive oxygen species (ROS) [35]. The decrease in Fe--S containing proteins, such as heme, electron transport chain (ETC) complexes I-III and the Kreb's cycle protein aconitase severely impairs cellular respiration [36], which is further complicated by simultaneous oxidative damage to these mitochondrial proteins. These events all culminate in an inability of the mitochondria to fulfill the cell's energy requirements resulting in cell death [35].
[0317] In order to assess the activity of frataxin inside mitochondria, the level of aconitase activity in mitochondria obtained from Friedreich's ataxia patients was examined upon administration of His-TAT-MTS(cs)-FRA.
[0318] Aconitase activity (mOD/min) was measured by following the conversion of isocitrate to cis-aconitate, as described above, in mitochondrial fractions of fibroblasts (F816) obtained from Friedreich's ataxia patients that were treated as described above and incubated in the presence of His-TAT-MTS(cs)-FRA or vehicle for 48 hours (FXN or VEH, respectively). His-TAT-MTS(cs)-FRA was administered at 20 .mu.g/ml and an additional dose of 20 .mu.g/ml was administered 24 hours after the first administration). HepG2 whole cells homogenate served as positive control (POS.CON).
[0319] As shown in FIG. 10, aconitase activity was higher in fibroblasts that were incubated with His-TAT-MTS(cs)-FRA (FXN, 14.5) compared to fibroblasts that were incubated in the presence of the vehicle control (VEH, 10.5). This finding is a clear demonstration that the fusion protein not only enters mitochondria, but is also active and available for its various cellular activities.
[0320] HepG2 whole cells homogenate served as positive control (POS.CON).
Example 7
[0321] TAT-MTS-FRA Fusion Proteins Partially Rescue FA-Patients' Cells as Well as Normal Cells from Oxidative Stress
[0322] As indicated above, the reduction in the levels of frataxin within the mitochondria leads to a rise in intracellular reactive oxygen species (ROS) [35].
[0323] L-Buthionine sulphoximine (BSO) is an inhibitor of gamma-glutamylcysteine synthetase (gamma-GCS) and, consequently lowers tissue glutathione (GSH) concentrations. GSH plays an important role in cellular defense or protection against a wide variety of toxic electrophiles via the formation of thioether conjugates. Therefore, BSO assay was used to inhibit de novo glutathione synthesis, depleting an important component of these cells' intrinsic defenses against reactive oxygen species (ROS) and allowing for the accumulation of ROS produced by natural cell processes, known to result in cell death [37]. The mechanism by which BSO inhibits production of GSH and results in cell death was described by Richardson, T. E. et al. [37]. Because they are lacking in Frataxin, Friedreich ataxia cells are extremely sensitive to BSO-induced oxidative stress compared with normal cells [37], and thus are used as an in vitro model of the long-term consequences of absent Frataxin.
[0324] Oxidative stress was induced with various concentrations of BSO in patients' cells as well as in normal healthy cells and the effect of the various TAT-MTS-FRA fusion proteins on cell death was measured. As can be seen in FIG. 11, BSO caused cell death of normal lymphocytes as well as of cells obtained from Friedreich ataxia patients. However, patients' cells were more sensitive to BSO-induced oxidative stress, consistent with previous findings, showing higher percentages of cell death. Most importantly, the various TAT-MTS-FRA fusion proteins, which were added a few hours before oxidative stress induction, were demonstrated to partially rescue both normal lymphocytes as well as patients' cells from cell death. This partial rescue was determined by both reduction in cell death and by reduction in caspase 3 activity, as demonstrated in FIG. 11B.
[0325] As shown in FIG. 11A and FIG. 11B, at least two out of the three fusion proteins carrying a heterologous MTS (namely, MTSorf and MTScs) demonstrated a superior protective effect with respect to the effect demonstrated by the fusion protein carrying the native MTS, in both patients' cells as well as in healthy cells, from BSO-induced oxidative stress.
[0326] A comparative study of the ability of the various TAT-MTS-FRA fusion proteins to rescue BSO-induced oxidative stress of patients' fibroblasts performed as detailed above is shown in FIG. 12A-FIG. 12D for the fusion protein constructs TAT-MTSfra-FRA, TAT-MTScs-FRA, TAT-MTSlad-FRA and TAT-MTSorf-FRA, respectively.
[0327] Interestingly, as demonstrated in FIG. 12B and in FIG. 12C, respectively, two of the protein constructs comprising heterologous MTS, namely the TAT-MTScs-FRA and TAT-MTSlad fusion proteins were more efficient than the fusion protein comprising the native frataxin MTS (TAT-MTSfra-FRA) in partially rescuing BSO-induced oxidative stress of patients' fibroblasts.
[0328] Similar protective effects in patients' cells induced with various concentrations of BSO were also observed for TAT-MTSorf-FRA and TAT-MTSlad fusion proteins when assayed alone (FIG. 13 and FIG. 14, respectively).
Example 8
Pharmacodynamic and Pharmacokinetic Assessment of TAT-MTS-FRA Fusion Proteins in Friedreich's Ataxia Mice
[0329] In an ongoing study, in order to evaluate pharmacodynamic, pharmacokinetic (PK/PD) and safety of TAT-MTS-FRA fusion proteins in a Friedreich's ataxia preclinical mouse model, the mouse model JR#18299 FVB; B6-Tg(FXN)1Sars Fxn.sub.tm1MKn/J is used in an ongoing study. The information obtained from these studies will be used for determining dose and time interval for further clinical studies. Briefly, the study requires 45 female mice, 6-8 weeks of age that are homozygous for the targeted mutation at the mouse Frataxin locus and hemizygous for the transgene. It is noted that strain 18299 is the original Sarsero mouse on a mixed genetic background, which was selected over the original C57BL/6J congenic strain due to its superior breeding performance and since no behavioral phenotyping is performed in this study.
[0330] In order to perform the preclinical study, two doses of TAT-MTScs-FRA fusion protein (at 100 .mu.g and 400 .mu.g) are administered twice per week for a maximal period of three weeks along with a vehicle, via tail vein injection into 18229 FVB;B6-Tg(FXN)1Sars Fxntm1Mkn/J mice. Mice are sacrificed 1, 4, 7, 14 or 21 days post injection and then blood is collected by cardiac puncture and various tissues are harvested (e.g. brain, heart, liver and kidney). Brain and heart tissue are processed to measure aconitase activity. Brain and heart tissue are also processed to measure frataxin levels by ELISA assay in mitochondrial extracts. Blood, liver and kidney are flash-frozen and stored.
[0331] The effect of the treatment is then assessed in the harvested tissues.
Example 9
[0332] TAT-MTS-OTC Protein Constructs Internalize into Mitochondria
[0333] Fusion protein constructs comprising OTC fused to a His-TAT-MTS were prepared as described above. As shown in FIG. 16, purified fusion protein constructs comprising the native MTS of OTC, the MTS of citrate synthase, the MTS of C6ORF66 and the MTS of lipoamide dehydrogenase were obtained (FIG. 16A, FIG. 16B, FIG. 16C and FIG. 16D, respectively). The size of the purified fusion protein constructs obtained was about 40 kDa as determined based on comparing their gel migration to the migration of the protein marker shown in FIG. 16E). Similar expression yields were obtained for all four fusion protein constructs that were prepared.
[0334] In order to examine the ability of the various OTC fusion protein constructs to enter mitochondria, HepG2 cells were incubated in the presence of the OTC fusion proteins constructs comprising either the native MTS of OTC or an MTS that is heterologous to OTC, namely, His-TAT-MTSotc-OTC, His-TAT-MTScs-OTC and TAT-MTSlad-OTC as described above.
[0335] First, as shown in FIG. 17A, it is noteworthy that the fusion protein constructs His-TAT-MTSotc-OTC, His-TAT-MTScs-OTC and His-TAT-MTSlad-OTC (and His-TAT-MTSorf-OTC, data not shown) were soluble under the various assay conditions based on the fact that no precipitation or degradation products were detected.
[0336] Analysis of mitochondrial fractions of HepG2 cells incubated with the above constructs revealed that the OTC fusion protein construct comprising the native MTS of OTC (His-TAT-MTSotc-OTC) as well as the OTC fusion protein constructs comprising an MTS that is heterologous to the OTC protein, namely citrate synthase (cs) and lipoamide dehydrogenase (lad) were able to enter mitochondria after an incubation period of 1 hour (FIG. 17A). FIG. 17A also shows that prolonging the incubation period to 3 hours resulted in an increase in the level of the fusion protein construct inside the mitochondria.
[0337] Interestingly, by comparing the ability of the protein construct comprising the native MTS of OTC (His-TAT-MTSotc-OTC) to the ability of OTC fusion protein constructs comprising an MTS that is heterologous to the OTC protein (namely His-TAT-MTScs-OTC and His-TAT-MTSlad-OTC) for example upon incubation of 3 hours in EMEM, it appears that the internalization ability of the fusion protein constructs comprising an MTS that is heterologous to OTC was slightly higher than the internalization ability of the fusion protein construct comprising the native MTS of OTC.
[0338] In order to further confirm the internalization of the OTC fusion protein constructs into mitochondria, the cytosolic fractions of the cells incubated with the various His-TAT-MTS-OTC protein constructs were also analyzed. As shown in FIG. 18A and FIG. 18B, no fusion protein constructs were observed in any of the assayed cytosolic fractions.
Example 10
In Vitro Enzymatic Activity of OTC
[0339] As indicated above, OTC is a protein having enzymatic activity that catalyzes the reaction between carbamoyl phosphate and ornithine to form citrulline and phosphate. In order to assess the activity of the purified fusion protein constructs comprising OTC described herein above, the production of citrulline from ornithine and carbamoyl phosphate by various fusion protein constructs was evaluated, as described above.
[0340] As shown in FIG. 19, the fusion protein construct His-TAT-MTScs-OTC assayed had relatively low enzymatic activity, where the fusion protein construct His-TAT-MTSotc-OTC had none. Without wishing to be bound by theory, this may be due to the fact that OTC is probably not present in the reaction mixture at its native trimeric state, which is necessary for its enzymatic activity, since it is conjugated to a His-TAT-MTS fragment.
[0341] However, as shown in FIG. 19 a low level of enzymatic activity was indeed observed for the fusion protein construct comprising the MTS of CS.
Example 11
[0342] Enzymatic Activity of OTC in HepG2 Cells Suffering from Ammonia Stress
[0343] OTC deficiency is the most common urea cycle disorder in humans. OTC, the defective enzyme in this disorder, is the final enzyme in the proximal portion of the urea cycle, and is responsible for converting carbamoyl phosphate and ornithine into citrulline, as indicated above. In severely affected individuals, ammonia concentrations increase rapidly causing ataxia, lethargy and death without rapid intervention.
[0344] In order to test the ability of the OTC fusion protein constructs to rescue ammonia stress in cells, HepG2 cells were administered daily (for 3 days) with a fusion protein construct comprising the native MTS of OTC or with fusion protein construct comprising the MTS of CS then treated with ammonium chloride, and finally, their cell viability was evaluated using the alamarBlue indicator, as described above.
[0345] As shown in FIG. 20, the level of cell viability in the presence of fusion protein constructs comprising OTC and citrate synthase as the MTS was similar to the level of cell viability for cells which were not exposed to ammonium chloride (untreated cells). Furthermore, the level of cell viability in the presence of a fusion protein construct comprising OTC and citrate synthase as the MTS was higher from the level of cell viability in cells treated with the fusion protein construct comprising the native MTS of OTC.
[0346] Taken together, the above results demonstrate that fusion protein constructs comprising TAT, MTS and OTC and in particular such fusion protein which comprise an MTS which is heterologous to OTC are able to enter mitochondria, the OTC is then processed, thereby enabling its enzymatic activity.
[0347] The results also indicated that the fusion protein constructs without ammonia were not cytotoxic (data not shown).
Example 12
Kinetics of Liver Entry and Dose Determination of TAT-MTS-OTC Fusion Protein Constructs in Mice
[0348] In order to assess the kinetics of liver entry of OTC, liver levels of any of the TAT-MTS-OTC fusion protein constructs described herein are determined in OTC-deficient spf.sup.ash mice (3 hemizygous males and 3 homozygous females at group) 4, 8, 24 and 48 hours after administration of a fusion protein construct comprising OTC (500 .mu.g/mouse). Liver pieces and other tissues (e.g. skeletal muscle, brain) are then harvested and analyzed for their OTC protein levels by Western blot analysis. In addition, the activity of OTC is examined in harvested liver, by using an OTC enzymatic assay (for example the citrulline assay described above).
[0349] Blood is collected by cardiac puncture and plasma is isolated for liver function tests (ALT/AST), amino acids (including citrulline), ammonia determination and OTC protein levels (Western blot). Urine is collected for orotic acid analysis.
[0350] Male mice are used for further studies in case the above experiments do not indicate any differences between males and females.
[0351] Then, protein levels are determined in tissues of mice administered with various doses of a TAT-MTS-OTC fusion protein construct (100-500 .mu.g/mouse) at the time point for which the highest OTC protein levels were detected in the above assay.
Example 13
Phenotype Protection by TAT-MTS-OTC Fusion Protein Constructs
[0352] OTC-deficient spf.sup.ash mice (6 males or 3 males and 3 females) are administered with OTC-shRNA knockdown rAAV according to Cunningham, S. C. et al. [41]. At a designated time-point (based on the activity of the OTC-shRNA batch and results obtained from the above experiments) dosing with a TAT-MTS-OTC fusion protein construct is commenced by administering 0, 1, 2 or 3 dose(s) per week to each tested group of mice.
[0353] Analysis is performed by first measuring baseline plasma levels of ammonia, amino acids (including citrulline) and ALT/AST as well as urinary orotic acid prior to injection of OTC-shRNA knockdown rAAV (blood is collected via tail vein nicking). Blood and urine are collected again prior to administration of a TAT-MTS-OTC fusion protein construct for the above analyses and then weekly for the duration of the experiment.
[0354] Mice are monitored and sacrificed upon observation of clinical signs of hyperammonaemia (lethargy, tremors, ataxia) or 1 month post-injection of a TAT-MTS-OTC fusion protein construct. At the termination of the experiment, liver, muscle and brain are frozen in liquid nitrogen or fixed in 4% PFA for vector copy analysis (quantitative PCR), OTC activity (in liver lysate and frozen sections) and protein levels (Western blot).
Sequence CWU 1
1
54127DNAHuman immunodeficiency virus type 1 1aggaagaagc ggagacagcg
acgaaga 272237DNAHomo sapiens 2tggactctcg ggcgccgcgc agtagccggc
ctcctggcgt cacccagccc ggcccaggcc 60cagaccctca cccgggtccc gcggccggca
gagttggccc cactctgcgg ccgccgtggc 120ctgcgcaccg acatcgatgc
gacctgcacg ccccgccgcg caagttcgaa ccaacgtggc 180ctcaaccaga
tttggaatgt caaaaagcag agtgtctatt tgatgaattt gaggaaa 237378DNAHomo
sapiens 3gctttactta ctgcggccgc ccggctcttg ggaaccaaga atgcatcttg
tcttgttctt 60gcagcccggc atgccagt 784102DNAHomo sapiens 4ggagcactag
tgattcgcgg tatcaggaat ttcaacctag agaaccgagc ggaacgggaa 60atcagcaaga
tgaagccctc tgtcgctccc agacacccct ct 1025105DNAHomo sapiens
5cagagctgga gtcgtgtgta ctgctccttg gccaagagag gccatttcaa tcgaatatct
60catggcctac agggactttc tgcagtgcct ctgagaactt acgca 1056393DNAHomo
sapiens 6tctggaactt tgggccaccc aggctctcta gatgagacca cctatgaaag
actagcagag 60gaaacgctgg actctttagc agagtttttt gaagaccttg cagacaagcc
atacacgttt 120gaggactatg atgtctcctt tgggagtggt gtcttaactg
tcaaactggg tggagatcta 180ggaacctatg tgatcaacaa gcagacgcca
aacaagcaaa tctggctatc ttctccatcc 240agtggaccta agcgttatga
ctggactggg aaaaactggg tgtactccca cgacggcgtg 300tccctccatg
agctgctggc cgcagagctc actaaagcct taaaaaccaa actggacttg
360tcttccttgg cctattccgg aaaagatgct tga 393727DNAArtificial
Sequenceforward primer for precursor frataxin cloning 7cgcggatccg
tggactctcg ggcgccg 27833DNAArtificial Sequencereverse primer for
precursor frataxin cloning 8acgctcgagt caagcatctt ttccggaata ggc
33930DNAArtificial Sequenceforward primer for MTS lad cloning
9cgcggatcca cagagctgga gtcgtgtgta 301066DNAArtificial
Sequencereverse primer for MTS lad cloning 10cataggtggt ctcatctaga
gagcctgggt ggcccaaagt tccagatgcg taagttctca 60gaggca
661128DNAArtificial Sequenceforward primer for MTS orf cloning
11cgcggatccg ggagcactag tgattcgc 281266DNAArtificial
Sequencereverse primer for MTS orf cloning 12cataggtggt ctcatctaga
gagcctgggt ggcccaaagt tccagaagag gggtgtctgg 60gagcga
6613112DNAArtificial Sequenceforward oligo for MTS cs cloning
13gatccggctt tacttactgc ggccgcccgg ctcttgggaa ccaagaatgc atcttgtctt
60gttcttgcag cccggcatgc cagttctgga actttgggcc acccaggctc tc
11214112DNAArtificial Sequencereverse primer for MTS cs cloning
14tctagagagc ctgggtggcc caaagttcca gaactggcat gccgggctgc aagaacaaga
60caagatgcat tcttggttcc caagagccgg gcggccgcag taagtaaagc cg
11215957DNAHomo sapiens 15ctgaagggcc gtgaccttct cactctaaga
aactttaccg gagaagaaat taaatatatg 60ctatggctat cagcagatct gaaatttagg
ataaaacaga aaggagagta tttgccttta 120ttgcaaggga agtccttagg
catgattttt gagaaaagaa gtactcgaac aagattgtct 180acagaaacag
gctttgcact tctgggagga catccttgtt ttcttaccac acaagatatt
240catttgggtg tgaatgaaag tctcacggac acggcccgtg tattgtctag
catggcagat 300gcagtattgg ctcgagtgta taaacaatca gatttggaca
ccctggctaa agaagcatcc 360atcccaatta tcaatgggct gtcagatttg
taccatccta tccagatcct ggctgattac 420ctcacgctcc aggaacacta
tagctctctg aaaggtctta ccctcagctg gatcggggat 480gggaacaata
tcctgcactc catcatgatg agcgcagcga aattcggaat gcaccttcag
540gcagctactc caaagggtta tgagccggat gctagtgtaa ccaagttggc
agagcagtat 600gccaaagaga atggtaccaa gctgttgctg acaaatgatc
cattggaagc agcgcatgga 660ggcaatgtat taattacaga cacttggata
agcatgggac aagaagagga gaagaaaaag 720cggctccagg ctttccaagg
ttaccaggtt acaatgaaga ctgctaaagt tgctgcctct 780gactggacat
ttttacactg cttgcccaga aagccagaag aagtggatga tgaagtcttt
840tattctcctc gatcactagt gttcccagag gcagaaaaca gaaagtggac
aatcatggct 900gtcatggtgt ccctgctgac agattactca cctcagctcc
agaagcctaa attttga 95716159DNAHomo sapiens 16atgtaccgct acctggccaa
agcgctgctg ccgtcccggg ccgggcccgc tgccctgggc 60tccgcggcca accactcggc
cgcgttgctg ggccggggcc gcggacagcc cgccgccgcc 120tcgcagccgg
ggctcgcatt ggccgcccgg cgccactac 15917190PRTArtificial SequenceHis
TAT MTS(cs) 81-210 FRA 17Met Gly Ser Ser His His His His His His
Ser Ser Gly Leu Val Pro1 5 10 15Arg Gly Ser His Met Arg Lys Lys Arg
Arg Gln Arg Arg Arg Gly Ser 20 25 30Asp Pro Ala Leu Leu Thr Ala Ala
Ala Arg Leu Leu Gly Thr Lys Asn 35 40 45Ala Ser Cys Leu Val Leu Ala
Ala Arg His Ala Ser Ser Gly Thr Leu 50 55 60Gly His Pro Gly Ser Leu
Asp Glu Thr Thr Tyr Glu Arg Leu Ala Glu65 70 75 80Glu Thr Leu Asp
Ser Leu Ala Glu Phe Phe Glu Asp Leu Ala Asp Lys 85 90 95Pro Tyr Thr
Phe Glu Asp Tyr Asp Val Ser Phe Gly Ser Gly Val Leu 100 105 110Thr
Val Lys Leu Gly Gly Asp Leu Gly Thr Tyr Val Ile Asn Lys Gln 115 120
125Thr Pro Asn Lys Gln Ile Trp Leu Ser Ser Pro Ser Ser Gly Pro Lys
130 135 140Arg Tyr Asp Trp Thr Gly Lys Asn Trp Val Tyr Ser His Asp
Gly Val145 150 155 160Ser Leu His Glu Leu Leu Ala Ala Glu Leu Thr
Lys Ala Leu Lys Thr 165 170 175Lys Leu Asp Leu Ser Ser Leu Ala Tyr
Ser Gly Lys Asp Ala 180 185 19018243PRTArtificial
SequenceHTFrataxin [His TAT MTS(fra)81-210 FRA] 18Met Gly Ser Ser
His His His His His His Ser Ser Gly Leu Val Pro1 5 10 15Arg Gly Ser
His Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Ser 20 25 30Asp Pro
Trp Thr Leu Gly Arg Arg Ala Val Ala Gly Leu Leu Ala Ser 35 40 45Pro
Ser Pro Ala Gln Ala Gln Thr Leu Thr Arg Val Pro Arg Pro Ala 50 55
60Glu Leu Ala Pro Leu Cys Gly Arg Arg Gly Leu Arg Thr Asp Ile Asp65
70 75 80Ala Thr Cys Thr Pro Arg Arg Ala Ser Ser Asn Gln Arg Gly Leu
Asn 85 90 95Gln Ile Trp Asn Val Lys Lys Gln Ser Val Tyr Leu Met Asn
Leu Arg 100 105 110Lys Ser Gly Thr Leu Gly His Pro Gly Ser Leu Asp
Glu Thr Thr Tyr 115 120 125Glu Arg Leu Ala Glu Glu Thr Leu Asp Ser
Leu Ala Glu Phe Phe Glu 130 135 140Asp Leu Ala Asp Lys Pro Tyr Thr
Phe Glu Asp Tyr Asp Val Ser Phe145 150 155 160Gly Ser Gly Val Leu
Thr Val Lys Leu Gly Gly Asp Leu Gly Thr Tyr 165 170 175Val Ile Asn
Lys Gln Thr Pro Asn Lys Gln Ile Trp Leu Ser Ser Pro 180 185 190Ser
Ser Gly Pro Lys Arg Tyr Asp Trp Thr Gly Lys Asn Trp Val Tyr 195 200
205Ser His Asp Gly Val Ser Leu His Glu Leu Leu Ala Ala Glu Leu Thr
210 215 220Lys Ala Leu Lys Thr Lys Leu Asp Leu Ser Ser Leu Ala Tyr
Ser Gly225 230 235 240Lys Asp Ala19199PRTArtificial SequenceHis TAT
MTS(lad) 81-210 FRA 19Met Gly Ser Ser His His His His His His Ser
Ser Gly Leu Val Pro1 5 10 15Arg Gly Ser His Met Arg Lys Lys Arg Arg
Gln Arg Arg Arg Gly Ser 20 25 30Asp Pro Gln Ser Trp Ser Arg Val Tyr
Cys Ser Leu Ala Lys Arg Gly 35 40 45His Phe Asn Arg Ile Ser His Gly
Leu Gln Gly Leu Ser Ala Val Pro 50 55 60Leu Arg Thr Tyr Ala Ser Gly
Thr Leu Gly His Pro Gly Ser Leu Asp65 70 75 80Glu Thr Thr Tyr Glu
Arg Leu Ala Glu Glu Thr Leu Asp Ser Leu Ala 85 90 95Glu Phe Phe Glu
Asp Leu Ala Asp Lys Pro Tyr Thr Phe Glu Asp Tyr 100 105 110Asp Val
Ser Phe Gly Ser Gly Val Leu Thr Val Lys Leu Gly Gly Asp 115 120
125Leu Gly Thr Tyr Val Ile Asn Lys Gln Thr Pro Asn Lys Gln Ile Trp
130 135 140Leu Ser Ser Pro Ser Ser Gly Pro Lys Arg Tyr Asp Trp Thr
Gly Lys145 150 155 160Asn Trp Val Tyr Ser His Asp Gly Val Ser Leu
His Glu Leu Leu Ala 165 170 175Ala Glu Leu Thr Lys Ala Leu Lys Thr
Lys Leu Asp Leu Ser Ser Leu 180 185 190Ala Tyr Ser Gly Lys Asp Ala
19520198PRTArtificial SequenceHis TAT MTS(orf) 81-210 FRA 20Met Gly
Ser Ser His His His His His His Ser Ser Gly Leu Val Pro1 5 10 15Arg
Gly Ser His Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Ser 20 25
30Asp Pro Gly Ala Leu Val Ile Arg Gly Ile Arg Asn Phe Asn Leu Glu
35 40 45Asn Arg Ala Glu Arg Glu Ile Ser Lys Met Lys Pro Ser Val Ala
Pro 50 55 60Arg His Pro Ser Ser Gly Thr Leu Gly His Pro Gly Ser Leu
Asp Glu65 70 75 80Thr Thr Tyr Glu Arg Leu Ala Glu Glu Thr Leu Asp
Ser Leu Ala Glu 85 90 95Phe Phe Glu Asp Leu Ala Asp Lys Pro Tyr Thr
Phe Glu Asp Tyr Asp 100 105 110Val Ser Phe Gly Ser Gly Val Leu Thr
Val Lys Leu Gly Gly Asp Leu 115 120 125Gly Thr Tyr Val Ile Asn Lys
Gln Thr Pro Asn Lys Gln Ile Trp Leu 130 135 140Ser Ser Pro Ser Ser
Gly Pro Lys Arg Tyr Asp Trp Thr Gly Lys Asn145 150 155 160Trp Val
Tyr Ser His Asp Gly Val Ser Leu His Glu Leu Leu Ala Ala 165 170
175Glu Leu Thr Lys Ala Leu Lys Thr Lys Leu Asp Leu Ser Ser Leu Ala
180 185 190Tyr Ser Gly Lys Asp Ala 1952111PRTHuman immunodeficiency
virus type 1 21Tyr Gly Arg Lys Lys Arg Arg Gln Arg Arg Arg1 5
102279PRTHomo sapiens 22Trp Thr Leu Gly Arg Arg Ala Val Ala Gly Leu
Leu Ala Ser Pro Ser1 5 10 15Pro Ala Gln Ala Gln Thr Leu Thr Arg Val
Pro Arg Pro Ala Glu Leu 20 25 30Ala Pro Leu Cys Gly Arg Arg Gly Leu
Arg Thr Asp Ile Asp Ala Thr 35 40 45Cys Thr Pro Arg Arg Ala Ser Ser
Asn Gln Arg Gly Leu Asn Gln Ile 50 55 60Trp Asn Val Lys Lys Gln Ser
Val Tyr Leu Met Asn Leu Arg Lys65 70 752326PRTHomo sapiens 23Ala
Leu Leu Thr Ala Ala Ala Arg Leu Leu Gly Thr Lys Asn Ala Ser1 5 10
15Cys Leu Val Leu Ala Ala Arg His Ala Ser 20 252435PRTHomo sapiens
24Gln Ser Trp Ser Arg Val Tyr Cys Ser Leu Ala Lys Arg Gly His Phe1
5 10 15Asn Arg Ile Ser His Gly Leu Gln Gly Leu Ser Ala Val Pro Leu
Arg 20 25 30Thr Tyr Ala 352534PRTHomo sapiens 25Gly Ala Leu Val Ile
Arg Gly Ile Arg Asn Phe Asn Leu Glu Asn Arg1 5 10 15Ala Glu Arg Glu
Ile Ser Lys Met Lys Pro Ser Val Ala Pro Arg His 20 25 30Pro
Ser26130PRTHomo sapiens 26Ser Gly Thr Leu Gly His Pro Gly Ser Leu
Asp Glu Thr Thr Tyr Glu1 5 10 15Arg Leu Ala Glu Glu Thr Leu Asp Ser
Leu Ala Glu Phe Phe Glu Asp 20 25 30Leu Ala Asp Lys Pro Tyr Thr Phe
Glu Asp Tyr Asp Val Ser Phe Gly 35 40 45Ser Gly Val Leu Thr Val Lys
Leu Gly Gly Asp Leu Gly Thr Tyr Val 50 55 60Ile Asn Lys Gln Thr Pro
Asn Lys Gln Ile Trp Leu Ser Ser Pro Ser65 70 75 80Ser Gly Pro Lys
Arg Tyr Asp Trp Thr Gly Lys Asn Trp Val Tyr Ser 85 90 95His Asp Gly
Val Ser Leu His Glu Leu Leu Ala Ala Glu Leu Thr Lys 100 105 110Ala
Leu Lys Thr Lys Leu Asp Leu Ser Ser Leu Ala Tyr Ser Gly Lys 115 120
125Asp Ala 130279PRTHuman immunodeficiency virus type 2 27Arg Lys
Lys Arg Arg Gln Arg Arg Arg1 528170PRTArtificial SequenceTAT
MTS(cs) 81-210 FRA 28Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly
Ser Asp Pro Ala Leu1 5 10 15Leu Thr Ala Ala Ala Arg Leu Leu Gly Thr
Lys Asn Ala Ser Cys Leu 20 25 30Val Leu Ala Ala Arg His Ala Ser Ser
Gly Thr Leu Gly His Pro Gly 35 40 45Ser Leu Asp Glu Thr Thr Tyr Glu
Arg Leu Ala Glu Glu Thr Leu Asp 50 55 60Ser Leu Ala Glu Phe Phe Glu
Asp Leu Ala Asp Lys Pro Tyr Thr Phe65 70 75 80Glu Asp Tyr Asp Val
Ser Phe Gly Ser Gly Val Leu Thr Val Lys Leu 85 90 95Gly Gly Asp Leu
Gly Thr Tyr Val Ile Asn Lys Gln Thr Pro Asn Lys 100 105 110Gln Ile
Trp Leu Ser Ser Pro Ser Ser Gly Pro Lys Arg Tyr Asp Trp 115 120
125Thr Gly Lys Asn Trp Val Tyr Ser His Asp Gly Val Ser Leu His Glu
130 135 140Leu Leu Ala Ala Glu Leu Thr Lys Ala Leu Lys Thr Lys Leu
Asp Leu145 150 155 160Ser Ser Leu Ala Tyr Ser Gly Lys Asp Ala 165
17029223PRTArtificial SequenceTAT MTS(fra)81-210 FRA 29Met Arg Lys
Lys Arg Arg Gln Arg Arg Arg Gly Ser Asp Pro Trp Thr1 5 10 15Leu Gly
Arg Arg Ala Val Ala Gly Leu Leu Ala Ser Pro Ser Pro Ala 20 25 30Gln
Ala Gln Thr Leu Thr Arg Val Pro Arg Pro Ala Glu Leu Ala Pro 35 40
45Leu Cys Gly Arg Arg Gly Leu Arg Thr Asp Ile Asp Ala Thr Cys Thr
50 55 60Pro Arg Arg Ala Ser Ser Asn Gln Arg Gly Leu Asn Gln Ile Trp
Asn65 70 75 80Val Lys Lys Gln Ser Val Tyr Leu Met Asn Leu Arg Lys
Ser Gly Thr 85 90 95Leu Gly His Pro Gly Ser Leu Asp Glu Thr Thr Tyr
Glu Arg Leu Ala 100 105 110Glu Glu Thr Leu Asp Ser Leu Ala Glu Phe
Phe Glu Asp Leu Ala Asp 115 120 125Lys Pro Tyr Thr Phe Glu Asp Tyr
Asp Val Ser Phe Gly Ser Gly Val 130 135 140Leu Thr Val Lys Leu Gly
Gly Asp Leu Gly Thr Tyr Val Ile Asn Lys145 150 155 160Gln Thr Pro
Asn Lys Gln Ile Trp Leu Ser Ser Pro Ser Ser Gly Pro 165 170 175Lys
Arg Tyr Asp Trp Thr Gly Lys Asn Trp Val Tyr Ser His Asp Gly 180 185
190Val Ser Leu His Glu Leu Leu Ala Ala Glu Leu Thr Lys Ala Leu Lys
195 200 205Thr Lys Leu Asp Leu Ser Ser Leu Ala Tyr Ser Gly Lys Asp
Ala 210 215 22030179PRTArtificial SequenceTAT MTS(lad) 81-210 FRA
30Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Ser Asp Pro Gln Ser1
5 10 15Trp Ser Arg Val Tyr Cys Ser Leu Ala Lys Arg Gly His Phe Asn
Arg 20 25 30Ile Ser His Gly Leu Gln Gly Leu Ser Ala Val Pro Leu Arg
Thr Tyr 35 40 45Ala Ser Gly Thr Leu Gly His Pro Gly Ser Leu Asp Glu
Thr Thr Tyr 50 55 60Glu Arg Leu Ala Glu Glu Thr Leu Asp Ser Leu Ala
Glu Phe Phe Glu65 70 75 80Asp Leu Ala Asp Lys Pro Tyr Thr Phe Glu
Asp Tyr Asp Val Ser Phe 85 90 95Gly Ser Gly Val Leu Thr Val Lys Leu
Gly Gly Asp Leu Gly Thr Tyr 100 105 110Val Ile Asn Lys Gln Thr Pro
Asn Lys Gln Ile Trp Leu Ser Ser Pro 115 120 125Ser Ser Gly Pro Lys
Arg Tyr Asp Trp Thr Gly Lys Asn Trp Val Tyr 130 135 140Ser His Asp
Gly Val Ser Leu His Glu Leu Leu Ala Ala Glu Leu Thr145 150 155
160Lys Ala Leu Lys Thr Lys Leu Asp Leu Ser Ser Leu Ala Tyr Ser Gly
165 170 175Lys Asp Ala31178PRTArtificial SequenceTAT MTS(orf)
81-210 FRA 31Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Ser Asp
Pro Gly Ala1 5 10 15Leu Val Ile Arg Gly Ile Arg Asn Phe Asn Leu Glu
Asn Arg Ala Glu 20 25 30Arg Glu Ile Ser Lys Met Lys Pro Ser Val Ala
Pro Arg His Pro Ser 35 40 45Ser Gly Thr Leu Gly His Pro Gly Ser Leu
Asp Glu Thr Thr Tyr Glu 50 55 60Arg Leu Ala Glu Glu Thr Leu Asp Ser
Leu Ala Glu Phe Phe Glu Asp65 70 75 80Leu Ala Asp Lys Pro
Tyr Thr Phe Glu Asp Tyr Asp Val Ser Phe Gly 85 90 95Ser Gly Val Leu
Thr Val Lys Leu Gly Gly Asp Leu Gly Thr Tyr Val 100 105 110Ile Asn
Lys Gln Thr Pro Asn Lys Gln Ile Trp Leu Ser Ser Pro Ser 115 120
125Ser Gly Pro Lys Arg Tyr Asp Trp Thr Gly Lys Asn Trp Val Tyr Ser
130 135 140His Asp Gly Val Ser Leu His Glu Leu Leu Ala Ala Glu Leu
Thr Lys145 150 155 160Ala Leu Lys Thr Lys Leu Asp Leu Ser Ser Leu
Ala Tyr Ser Gly Lys 165 170 175Asp Ala324PRTArtificial
Sequencepolypeptide linker 32Gly Ser Asp Pro133166PRTArtificial
SequenceTAT MTS(cs) 81-210 FRA delta linker 33Met Arg Lys Lys Arg
Arg Gln Arg Arg Arg Ala Leu Leu Thr Ala Ala1 5 10 15Ala Arg Leu Leu
Gly Thr Lys Asn Ala Ser Cys Leu Val Leu Ala Ala 20 25 30Arg His Ala
Ser Ser Gly Thr Leu Gly His Pro Gly Ser Leu Asp Glu 35 40 45Thr Thr
Tyr Glu Arg Leu Ala Glu Glu Thr Leu Asp Ser Leu Ala Glu 50 55 60Phe
Phe Glu Asp Leu Ala Asp Lys Pro Tyr Thr Phe Glu Asp Tyr Asp65 70 75
80Val Ser Phe Gly Ser Gly Val Leu Thr Val Lys Leu Gly Gly Asp Leu
85 90 95Gly Thr Tyr Val Ile Asn Lys Gln Thr Pro Asn Lys Gln Ile Trp
Leu 100 105 110Ser Ser Pro Ser Ser Gly Pro Lys Arg Tyr Asp Trp Thr
Gly Lys Asn 115 120 125Trp Val Tyr Ser His Asp Gly Val Ser Leu His
Glu Leu Leu Ala Ala 130 135 140Glu Leu Thr Lys Ala Leu Lys Thr Lys
Leu Asp Leu Ser Ser Leu Ala145 150 155 160Tyr Ser Gly Lys Asp Ala
16534219PRTArtificial SequenceTAT MTS(fra)81-210 FRA delta linker
34Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Trp Thr Leu Gly Arg Arg1
5 10 15Ala Val Ala Gly Leu Leu Ala Ser Pro Ser Pro Ala Gln Ala Gln
Thr 20 25 30Leu Thr Arg Val Pro Arg Pro Ala Glu Leu Ala Pro Leu Cys
Gly Arg 35 40 45Arg Gly Leu Arg Thr Asp Ile Asp Ala Thr Cys Thr Pro
Arg Arg Ala 50 55 60Ser Ser Asn Gln Arg Gly Leu Asn Gln Ile Trp Asn
Val Lys Lys Gln65 70 75 80Ser Val Tyr Leu Met Asn Leu Arg Lys Ser
Gly Thr Leu Gly His Pro 85 90 95Gly Ser Leu Asp Glu Thr Thr Tyr Glu
Arg Leu Ala Glu Glu Thr Leu 100 105 110Asp Ser Leu Ala Glu Phe Phe
Glu Asp Leu Ala Asp Lys Pro Tyr Thr 115 120 125Phe Glu Asp Tyr Asp
Val Ser Phe Gly Ser Gly Val Leu Thr Val Lys 130 135 140Leu Gly Gly
Asp Leu Gly Thr Tyr Val Ile Asn Lys Gln Thr Pro Asn145 150 155
160Lys Gln Ile Trp Leu Ser Ser Pro Ser Ser Gly Pro Lys Arg Tyr Asp
165 170 175Trp Thr Gly Lys Asn Trp Val Tyr Ser His Asp Gly Val Ser
Leu His 180 185 190Glu Leu Leu Ala Ala Glu Leu Thr Lys Ala Leu Lys
Thr Lys Leu Asp 195 200 205Leu Ser Ser Leu Ala Tyr Ser Gly Lys Asp
Ala 210 21535175PRTArtificial SequenceTAT MTS(lad) 81-210 FRA delta
linker 35Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gln Ser Trp Ser
Arg Val1 5 10 15Tyr Cys Ser Leu Ala Lys Arg Gly His Phe Asn Arg Ile
Ser His Gly 20 25 30Leu Gln Gly Leu Ser Ala Val Pro Leu Arg Thr Tyr
Ala Ser Gly Thr 35 40 45Leu Gly His Pro Gly Ser Leu Asp Glu Thr Thr
Tyr Glu Arg Leu Ala 50 55 60Glu Glu Thr Leu Asp Ser Leu Ala Glu Phe
Phe Glu Asp Leu Ala Asp65 70 75 80Lys Pro Tyr Thr Phe Glu Asp Tyr
Asp Val Ser Phe Gly Ser Gly Val 85 90 95Leu Thr Val Lys Leu Gly Gly
Asp Leu Gly Thr Tyr Val Ile Asn Lys 100 105 110Gln Thr Pro Asn Lys
Gln Ile Trp Leu Ser Ser Pro Ser Ser Gly Pro 115 120 125Lys Arg Tyr
Asp Trp Thr Gly Lys Asn Trp Val Tyr Ser His Asp Gly 130 135 140Val
Ser Leu His Glu Leu Leu Ala Ala Glu Leu Thr Lys Ala Leu Lys145 150
155 160Thr Lys Leu Asp Leu Ser Ser Leu Ala Tyr Ser Gly Lys Asp Ala
165 170 17536174PRTArtificial SequenceTAT MTS(orf) 81-210 FRA delta
linker 36Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Ala Leu Val
Ile Arg1 5 10 15Gly Ile Arg Asn Phe Asn Leu Glu Asn Arg Ala Glu Arg
Glu Ile Ser 20 25 30Lys Met Lys Pro Ser Val Ala Pro Arg His Pro Ser
Ser Gly Thr Leu 35 40 45Gly His Pro Gly Ser Leu Asp Glu Thr Thr Tyr
Glu Arg Leu Ala Glu 50 55 60Glu Thr Leu Asp Ser Leu Ala Glu Phe Phe
Glu Asp Leu Ala Asp Lys65 70 75 80Pro Tyr Thr Phe Glu Asp Tyr Asp
Val Ser Phe Gly Ser Gly Val Leu 85 90 95Thr Val Lys Leu Gly Gly Asp
Leu Gly Thr Tyr Val Ile Asn Lys Gln 100 105 110Thr Pro Asn Lys Gln
Ile Trp Leu Ser Ser Pro Ser Ser Gly Pro Lys 115 120 125Arg Tyr Asp
Trp Thr Gly Lys Asn Trp Val Tyr Ser His Asp Gly Val 130 135 140Ser
Leu His Glu Leu Leu Ala Ala Glu Leu Thr Lys Ala Leu Lys Thr145 150
155 160Lys Leu Asp Leu Ser Ser Leu Ala Tyr Ser Gly Lys Asp Ala 165
17037105DNAHomo sapiens 37ctgtttaacc tgcgcattct gctgaacaat
gcggccttcc gtaacggcca taattttatg 60gtccgcaact tccgttgcgg tcagccgctg
caaaataaag tgcag 1053835PRTHomo sapiens 38Leu Phe Asn Leu Arg Ile
Leu Leu Asn Asn Ala Ala Phe Arg Asn Gly1 5 10 15His Asn Phe Met Val
Arg Asn Phe Arg Cys Gly Gln Pro Leu Gln Asn 20 25 30Lys Val Gln
3539318PRTHomo sapiens 39Leu Lys Gly Arg Asp Leu Leu Thr Leu Arg
Asn Phe Thr Gly Glu Glu1 5 10 15Ile Lys Tyr Met Leu Trp Leu Ser Ala
Asp Leu Lys Phe Arg Ile Lys 20 25 30Gln Lys Gly Glu Tyr Leu Pro Leu
Leu Gln Gly Lys Ser Leu Gly Met 35 40 45Ile Phe Glu Lys Arg Ser Thr
Arg Thr Arg Leu Ser Thr Glu Thr Gly 50 55 60Phe Ala Leu Leu Gly Gly
His Pro Cys Phe Leu Thr Thr Gln Asp Ile65 70 75 80His Leu Gly Val
Asn Glu Ser Leu Thr Asp Thr Ala Arg Val Leu Ser 85 90 95Ser Met Ala
Asp Ala Val Leu Ala Arg Val Tyr Lys Gln Ser Asp Leu 100 105 110Asp
Thr Leu Ala Lys Glu Ala Ser Ile Pro Ile Ile Asn Gly Leu Ser 115 120
125Asp Leu Tyr His Pro Ile Gln Ile Leu Ala Asp Tyr Leu Thr Leu Gln
130 135 140Glu His Tyr Ser Ser Leu Lys Gly Leu Thr Leu Ser Trp Ile
Gly Asp145 150 155 160Gly Asn Asn Ile Leu His Ser Ile Met Met Ser
Ala Ala Lys Phe Gly 165 170 175Met His Leu Gln Ala Ala Thr Pro Lys
Gly Tyr Glu Pro Asp Ala Ser 180 185 190Val Thr Lys Leu Ala Glu Gln
Tyr Ala Lys Glu Asn Gly Thr Lys Leu 195 200 205Leu Leu Thr Asn Asp
Pro Leu Glu Ala Ala His Gly Gly Asn Val Leu 210 215 220Ile Thr Asp
Thr Trp Ile Ser Met Gly Gln Glu Glu Glu Lys Lys Lys225 230 235
240Arg Leu Gln Ala Phe Gln Gly Tyr Gln Val Thr Met Lys Thr Ala Lys
245 250 255Val Ala Ala Ser Asp Trp Thr Phe Leu His Cys Leu Pro Arg
Lys Pro 260 265 270Glu Glu Val Asp Asp Glu Val Phe Tyr Ser Pro Arg
Ser Leu Val Phe 275 280 285Pro Glu Ala Glu Asn Arg Lys Trp Thr Ile
Met Ala Val Met Val Ser 290 295 300Leu Leu Thr Asp Tyr Ser Pro Gln
Leu Gln Lys Pro Lys Phe305 310 315401170DNAArtificial
SequenceHis-TAT-MTS(otc)-OTC delta inker 40catatgggct catcgcatca
tcatcatcat cactcatcag gtctggttcc gcgtggctcg 60cacatgtatg gtcgcaaaaa
acgtcgtcaa cgtcgccgtc tgtttaacct gcgcattctg 120ctgaacaatg
cggccttccg taacggccat aattttatgg tccgcaactt ccgttgcggt
180cagccgctgc aaaataaagt gcagctgaaa ggccgcgatc tgctgaccct
gcgtaacttc 240acgggtgaag aaatcaaata catgctgtgg ctgagcgcag
acctgaaatt ccgcatcaaa 300caaaaaggcg aatacctgcc gctgctgcag
ggcaaatctc tgggtatgat ttttgaaaaa 360cgtagtaccc gcacgcgtct
gtccaccgaa acgggctttg ccctgctggg cggtcatccg 420tgtttcctga
ccacgcaaga tatccacctg ggtgtgaacg aaagtctgac cgatacggca
480cgcgttctga gctctatggc agacgctgtg ctggctcgtg tttataaaca
gtccgatctg 540gacaccctgg cgaaagaagc ctcaattccg attatcaatg
gcctgtcgga tctgtaccat 600ccgattcaaa tcctggcgga ctatctgacc
ctgcaggaac actacagttc cctgaaaggt 660ctgaccctga gttggatcgg
cgatggtaac aatattctgc atagcatcat gatgtctgca 720gctaaatttg
gcatgcacct gcaagcggcc accccgaaag gttatgaacc ggatgccagc
780gttacgaaac tggcagaaca gtacgctaaa gaaaacggta ccaaactgct
gctgacgaat 840gatccgctgg aagcagctca tggcggtaac gtcctgatta
ccgacacgtg gatctctatg 900ggccaggaag aagaaaagaa aaaacgtctg
caggcgtttc aaggttatca ggttaccatg 960aaaacggcca aagtcgcggc
cagcgattgg accttcctgc actgcctgcc gcgtaaaccg 1020gaagaagtcg
atgacgaagt gttttactca ccgcgctcgc tggtgttccc ggaagcagaa
1080aatcgtaaat ggaccatcat ggctgttatg gtgtccctgc tgaccgacta
ttccccgcaa 1140ctgcaaaaac cgaaattcta atgaaagctt
1170411155DNAArtificial SequenceHis-TAT-MTS(cs)-OTC 41catatgggct
catctcatca tcatcatcat cattcgtcag gtctggtccc gcgtggctct 60cacatgcgta
aaaaacgtcg tcagcgtcgt cgtggcagtg atccggcact gctgaccgca
120gcagcacgtc tgctgggtac gaaaaacgct agctgcctgg tgctggctgc
gcgtcatgcg 180tctgaatttc tgaaaggccg tgatctgctg accctgcgca
acttcacggg tgaagaaatc 240aaatacatgc tgtggctgag tgccgacctg
aaatttcgta tcaaacaaaa aggcgaatac 300ctgccgctgc tgcagggcaa
atccctgggt atgattttcg aaaaacgcag tacccgtacg 360cgcctgtcca
ccgaaacggg ctttgcactg ctgggcggtc atccgtgttt cctgaccacg
420caagatatcc acctgggtgt gaacgaatca ctgaccgata cggctcgtgt
tctgagctct 480atggcagacg cagtgctggc acgtgtttat aaacagtcgg
atctggacac cctggctaaa 540gaagcgtcaa ttccgattat caatggcctg
tcggatctgt accatccgat tcaaatcctg 600gcggactatc tgaccctgca
ggaacactac agttccctga aaggtctgac cctgagctgg 660atcggcgatg
gtaacaatat tctgcatagc atcatgatgt ctgccgcaaa atttggcatg
720cacctgcaag ctgcgacccc gaaaggttat gaaccggacg ccagcgtcac
gaaactggcc 780gaacagtacg caaaagaaaa cggtaccaaa ctgctgctga
cgaatgatcc gctggaagcc 840gcacatggcg gtaacgttct gattaccgac
acgtggatca gcatgggcca ggaagaagaa 900aagaaaaaac gtctgcaggc
ctttcaaggt tatcaggtta ccatgaaaac ggcaaaagtc 960gctgcgtctg
attggacctt cctgcactgc ctgccgcgca aaccggaaga agtcgatgac
1020gaagtgtttt actcaccgcg ttcgctggtt ttcccggaag cggaaaatcg
caaatggacc 1080attatggctg tgatggtctc tctgctgacg gactactcgc
cgcaactgca aaaaccgaaa 1140ttctaatgaa agctt 1155421179DNAArtificial
SequenceHis-TAT-MTS(orf)-OTC 42catatgggtt catcacatca tcatcatcat
cattcatcag gtctggtccc gcgtggttca 60cacatgcgta aaaaacgtcg tcagcgtcgt
cgtggcagtg atccgggtgc gctggtcatt 120cgtggcatcc gcaactttaa
tctggaaaac cgtgcggaac gcgaaattag taaaatgaaa 180ccgtccgtgg
caccgcgtca tccgtctgaa tttctgaaag gccgtgatct gctgaccctg
240cgcaacttca cgggtgaaga aatcaaatac atgctgtggc tgagtgcaga
cctgaaattc 300cgtatcaaac aaaagggtga atacctgccg ctgctgcagg
gcaaatccct gggtatgatt 360ttcgaaaaac gctcaacccg tacgcgcctg
tcgaccgaaa cgggctttgc cctgctgggc 420ggtcatccgt gcttcctgac
cacgcaagat atccacctgg gtgtgaacga atcactgacc 480gatacggcac
gtgttctgag ctctatggca gacgcagtgc tggctcgtgt ttataaacag
540tcggatctgg acaccctggc aaaagaagct agcattccga ttatcaatgg
cctgtctgat 600ctgtaccatc cgattcaaat cctggcggac tatctgaccc
tgcaggaaca ctacagttcc 660ctgaaaggtc tgaccctgag ctggatcggc
gatggtaaca atattctgca tagcatcatg 720atgtctgcgg ccaaattcgg
catgcacctg caagcagcta ccccgaaagg ttatgaaccg 780gacgcctccg
ttacgaaact ggcggaacag tacgccaaag aaaacggcac caaactgctg
840ctgacgaatg atccgctgga agcggcccat ggcggtaacg tcctgattac
cgacacgtgg 900atcagcatgg gccaggaaga agaaaagaaa aaacgtctgc
aggcatttca aggttatcag 960gttaccatga aaacggctaa agtcgcagct
tctgattgga ccttcctgca ctgtctgccg 1020cgcaaaccgg aagaagtcga
tgacgaagtg ttttactcac cgcgttcgct ggtgttcccg 1080gaagcggaaa
atcgcaaatg gaccattatg gctgtgatgg tgtcgctgct gacggactac
1140tcgccgcaac tgcaaaaacc gaaattctaa tgaaagctt
1179431182DNAArtificial SequenceHis-TAT-MTS(lad)-OTC 43catatgggta
gttcacatca tcatcatcat cactcgtcgg gtctggtgcc gcgtggctca 60cacatgcgta
aaaaacgtcg tcagcgtcgt cgtggctcag atccgcaatc atggtcgcgc
120gtctattgct cgctggcgaa acgtggtcat tttaaccgca ttagccacgg
cctgcagggt 180ctgtctgcag tgccgctgcg tacctacgct gaatttctga
aaggccgtga tctgctgacc 240ctgcgcaact tcacgggtga agaaatcaaa
tacatgctgt ggctgagcgc agacctgaaa 300tttcgtatca aacaaaaagg
cgaatacctg ccgctgctgc agggcaaatc tctgggtatg 360attttcgaaa
aacgctcaac ccgtacgcgc ctgtcgaccg aaacgggctt tgccctgctg
420ggcggtcatc cgtgtttcct gaccacgcag gatatccacc tgggtgtgaa
cgaaagtctg 480accgatacgg cacgtgttct gagctctatg gcagacgcag
tgctggctcg tgtttataaa 540cagtccgatc tggacaccct ggcaaaagaa
gctagtattc cgattatcaa tggcctgtcc 600gatctgtacc atccgattca
aatcctggcg gactatctga ccctgcagga acactacagt 660tccctgaaag
gtctgacgct gagctggatc ggcgatggta acaatattct gcatagtatc
720atgatgtccg cggccaaatt cggcatgcac ctgcaagcag ctaccccgaa
aggttatgaa 780ccggacgcct ctgttacgaa actggcggaa cagtacgcca
aagaaaacgg taccaaactg 840ctgctgacga atgatccgct ggaagcggcc
catggcggta acgtcctgat taccgacacg 900tggatcagta tgggccagga
agaagaaaag aaaaaacgtc tgcaggcgtt tcaaggttat 960caggttacca
tgaaaacggc caaagtcgca gctagcgatt ggaccttcct gcactgcctg
1020ccgcgcaaac cggaagaagt cgatgacgaa gtgttttata gcccgcgttc
tctggtgttc 1080ccggaagcgg aaaatcgcaa atggaccatc atggccgtta
tggtgtcgct gctgaccgat 1140tactccccgc aactgcaaaa accgaaattc
taatgaaagc tt 118244385PRTArtificial SequenceHis-TAT-MTS(otc)-OTC
delta linker 44Met Gly Ser Ser His His His His His His Ser Ser Gly
Leu Val Pro1 5 10 15Arg Gly Ser His Met Tyr Gly Arg Lys Lys Arg Arg
Gln Arg Arg Arg 20 25 30Leu Phe Asn Leu Arg Ile Leu Leu Asn Asn Ala
Ala Phe Arg Asn Gly 35 40 45His Asn Phe Met Val Arg Asn Phe Arg Cys
Gly Gln Pro Leu Gln Asn 50 55 60Lys Val Gln Leu Lys Gly Arg Asp Leu
Leu Thr Leu Arg Asn Phe Thr65 70 75 80Gly Glu Glu Ile Lys Tyr Met
Leu Trp Leu Ser Ala Asp Leu Lys Phe 85 90 95Arg Ile Lys Gln Lys Gly
Glu Tyr Leu Pro Leu Leu Gln Gly Lys Ser 100 105 110Leu Gly Met Ile
Phe Glu Lys Arg Ser Thr Arg Thr Arg Leu Ser Thr 115 120 125Glu Thr
Gly Phe Ala Leu Leu Gly Gly His Pro Cys Phe Leu Thr Thr 130 135
140Gln Asp Ile His Leu Gly Val Asn Glu Ser Leu Thr Asp Thr Ala
Arg145 150 155 160Val Leu Ser Ser Met Ala Asp Ala Val Leu Ala Arg
Val Tyr Lys Gln 165 170 175Ser Asp Leu Asp Thr Leu Ala Lys Glu Ala
Ser Ile Pro Ile Ile Asn 180 185 190Gly Leu Ser Asp Leu Tyr His Pro
Ile Gln Ile Leu Ala Asp Tyr Leu 195 200 205Thr Leu Gln Glu His Tyr
Ser Ser Leu Lys Gly Leu Thr Leu Ser Trp 210 215 220Ile Gly Asp Gly
Asn Asn Ile Leu His Ser Ile Met Met Ser Ala Ala225 230 235 240Lys
Phe Gly Met His Leu Gln Ala Ala Thr Pro Lys Gly Tyr Glu Pro 245 250
255Asp Ala Ser Val Thr Lys Leu Ala Glu Gln Tyr Ala Lys Glu Asn Gly
260 265 270Thr Lys Leu Leu Leu Thr Asn Asp Pro Leu Glu Ala Ala His
Gly Gly 275 280 285Asn Val Leu Ile Thr Asp Thr Trp Ile Ser Met Gly
Gln Glu Glu Glu 290 295 300Lys Lys Lys Arg Leu Gln Ala Phe Gln Gly
Tyr Gln Val Thr Met Lys305 310 315 320Thr Ala Lys Val Ala Ala Ser
Asp Trp Thr Phe Leu His Cys Leu Pro 325 330 335Arg Lys Pro Glu Glu
Val Asp Asp Glu Val Phe Tyr Ser Pro Arg Ser 340 345 350Leu Val Phe
Pro Glu Ala Glu Asn Arg Lys Trp Thr Ile Met Ala Val 355 360 365Met
Val Ser Leu Leu Thr Asp Tyr Ser Pro Gln Leu Gln Lys Pro Lys 370
375
380Phe38545380PRTArtificial SequenceHis-TAT-MTS(cs)-OTC 45Met Gly
Ser Ser His His His His His His Ser Ser Gly Leu Val Pro1 5 10 15Arg
Gly Ser His Met Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Ser 20 25
30Asp Pro Ala Leu Leu Thr Ala Ala Ala Arg Leu Leu Gly Thr Lys Asn
35 40 45Ala Ser Cys Leu Val Leu Ala Ala Arg His Ala Ser Glu Phe Leu
Lys 50 55 60Gly Arg Asp Leu Leu Thr Leu Arg Asn Phe Thr Gly Glu Glu
Ile Lys65 70 75 80Tyr Met Leu Trp Leu Ser Ala Asp Leu Lys Phe Arg
Ile Lys Gln Lys 85 90 95Gly Glu Tyr Leu Pro Leu Leu Gln Gly Lys Ser
Leu Gly Met Ile Phe 100 105 110Glu Lys Arg Ser Thr Arg Thr Arg Leu
Ser Thr Glu Thr Gly Phe Ala 115 120 125Leu Leu Gly Gly His Pro Cys
Phe Leu Thr Thr Gln Asp Ile His Leu 130 135 140Gly Val Asn Glu Ser
Leu Thr Asp Thr Ala Arg Val Leu Ser Ser Met145 150 155 160Ala Asp
Ala Val Leu Ala Arg Val Tyr Lys Gln Ser Asp Leu Asp Thr 165 170
175Leu Ala Lys Glu Ala Ser Ile Pro Ile Ile Asn Gly Leu Ser Asp Leu
180 185 190Tyr His Pro Ile Gln Ile Leu Ala Asp Tyr Leu Thr Leu Gln
Glu His 195 200 205Tyr Ser Ser Leu Lys Gly Leu Thr Leu Ser Trp Ile
Gly Asp Gly Asn 210 215 220Asn Ile Leu His Ser Ile Met Met Ser Ala
Ala Lys Phe Gly Met His225 230 235 240Leu Gln Ala Ala Thr Pro Lys
Gly Tyr Glu Pro Asp Ala Ser Val Thr 245 250 255Lys Leu Ala Glu Gln
Tyr Ala Lys Glu Asn Gly Thr Lys Leu Leu Leu 260 265 270Thr Asn Asp
Pro Leu Glu Ala Ala His Gly Gly Asn Val Leu Ile Thr 275 280 285Asp
Thr Trp Ile Ser Met Gly Gln Glu Glu Glu Lys Lys Lys Arg Leu 290 295
300Gln Ala Phe Gln Gly Tyr Gln Val Thr Met Lys Thr Ala Lys Val
Ala305 310 315 320Ala Ser Asp Trp Thr Phe Leu His Cys Leu Pro Arg
Lys Pro Glu Glu 325 330 335Val Asp Asp Glu Val Phe Tyr Ser Pro Arg
Ser Leu Val Phe Pro Glu 340 345 350Ala Glu Asn Arg Lys Trp Thr Ile
Met Ala Val Met Val Ser Leu Leu 355 360 365Thr Asp Tyr Ser Pro Gln
Leu Gln Lys Pro Lys Phe 370 375 38046388PRTArtificial
SequenceHis-TAT-MTS(orf)-OTC 46Met Gly Ser Ser His His His His His
His Ser Ser Gly Leu Val Pro1 5 10 15Arg Gly Ser His Met Arg Lys Lys
Arg Arg Gln Arg Arg Arg Gly Ser 20 25 30Asp Pro Gly Ala Leu Val Ile
Arg Gly Ile Arg Asn Phe Asn Leu Glu 35 40 45Asn Arg Ala Glu Arg Glu
Ile Ser Lys Met Lys Pro Ser Val Ala Pro 50 55 60Arg His Pro Ser Glu
Phe Leu Lys Gly Arg Asp Leu Leu Thr Leu Arg65 70 75 80Asn Phe Thr
Gly Glu Glu Ile Lys Tyr Met Leu Trp Leu Ser Ala Asp 85 90 95Leu Lys
Phe Arg Ile Lys Gln Lys Gly Glu Tyr Leu Pro Leu Leu Gln 100 105
110Gly Lys Ser Leu Gly Met Ile Phe Glu Lys Arg Ser Thr Arg Thr Arg
115 120 125Leu Ser Thr Glu Thr Gly Phe Ala Leu Leu Gly Gly His Pro
Cys Phe 130 135 140Leu Thr Thr Gln Asp Ile His Leu Gly Val Asn Glu
Ser Leu Thr Asp145 150 155 160Thr Ala Arg Val Leu Ser Ser Met Ala
Asp Ala Val Leu Ala Arg Val 165 170 175Tyr Lys Gln Ser Asp Leu Asp
Thr Leu Ala Lys Glu Ala Ser Ile Pro 180 185 190Ile Ile Asn Gly Leu
Ser Asp Leu Tyr His Pro Ile Gln Ile Leu Ala 195 200 205Asp Tyr Leu
Thr Leu Gln Glu His Tyr Ser Ser Leu Lys Gly Leu Thr 210 215 220Leu
Ser Trp Ile Gly Asp Gly Asn Asn Ile Leu His Ser Ile Met Met225 230
235 240Ser Ala Ala Lys Phe Gly Met His Leu Gln Ala Ala Thr Pro Lys
Gly 245 250 255Tyr Glu Pro Asp Ala Ser Val Thr Lys Leu Ala Glu Gln
Tyr Ala Lys 260 265 270Glu Asn Gly Thr Lys Leu Leu Leu Thr Asn Asp
Pro Leu Glu Ala Ala 275 280 285His Gly Gly Asn Val Leu Ile Thr Asp
Thr Trp Ile Ser Met Gly Gln 290 295 300Glu Glu Glu Lys Lys Lys Arg
Leu Gln Ala Phe Gln Gly Tyr Gln Val305 310 315 320Thr Met Lys Thr
Ala Lys Val Ala Ala Ser Asp Trp Thr Phe Leu His 325 330 335Cys Leu
Pro Arg Lys Pro Glu Glu Val Asp Asp Glu Val Phe Tyr Ser 340 345
350Pro Arg Ser Leu Val Phe Pro Glu Ala Glu Asn Arg Lys Trp Thr Ile
355 360 365Met Ala Val Met Val Ser Leu Leu Thr Asp Tyr Ser Pro Gln
Leu Gln 370 375 380Lys Pro Lys Phe38547389PRTArtificial
SequenceHis-TAT-MTS(lad)-OTC 47Met Gly Ser Ser His His His His His
His Ser Ser Gly Leu Val Pro1 5 10 15Arg Gly Ser His Met Arg Lys Lys
Arg Arg Gln Arg Arg Arg Gly Ser 20 25 30Asp Pro Gln Ser Trp Ser Arg
Val Tyr Cys Ser Leu Ala Lys Arg Gly 35 40 45His Phe Asn Arg Ile Ser
His Gly Leu Gln Gly Leu Ser Ala Val Pro 50 55 60Leu Arg Thr Tyr Ala
Glu Phe Leu Lys Gly Arg Asp Leu Leu Thr Leu65 70 75 80Arg Asn Phe
Thr Gly Glu Glu Ile Lys Tyr Met Leu Trp Leu Ser Ala 85 90 95Asp Leu
Lys Phe Arg Ile Lys Gln Lys Gly Glu Tyr Leu Pro Leu Leu 100 105
110Gln Gly Lys Ser Leu Gly Met Ile Phe Glu Lys Arg Ser Thr Arg Thr
115 120 125Arg Leu Ser Thr Glu Thr Gly Phe Ala Leu Leu Gly Gly His
Pro Cys 130 135 140Phe Leu Thr Thr Gln Asp Ile His Leu Gly Val Asn
Glu Ser Leu Thr145 150 155 160Asp Thr Ala Arg Val Leu Ser Ser Met
Ala Asp Ala Val Leu Ala Arg 165 170 175Val Tyr Lys Gln Ser Asp Leu
Asp Thr Leu Ala Lys Glu Ala Ser Ile 180 185 190Pro Ile Ile Asn Gly
Leu Ser Asp Leu Tyr His Pro Ile Gln Ile Leu 195 200 205Ala Asp Tyr
Leu Thr Leu Gln Glu His Tyr Ser Ser Leu Lys Gly Leu 210 215 220Thr
Leu Ser Trp Ile Gly Asp Gly Asn Asn Ile Leu His Ser Ile Met225 230
235 240Met Ser Ala Ala Lys Phe Gly Met His Leu Gln Ala Ala Thr Pro
Lys 245 250 255Gly Tyr Glu Pro Asp Ala Ser Val Thr Lys Leu Ala Glu
Gln Tyr Ala 260 265 270Lys Glu Asn Gly Thr Lys Leu Leu Leu Thr Asn
Asp Pro Leu Glu Ala 275 280 285Ala His Gly Gly Asn Val Leu Ile Thr
Asp Thr Trp Ile Ser Met Gly 290 295 300Gln Glu Glu Glu Lys Lys Lys
Arg Leu Gln Ala Phe Gln Gly Tyr Gln305 310 315 320Val Thr Met Lys
Thr Ala Lys Val Ala Ala Ser Asp Trp Thr Phe Leu 325 330 335His Cys
Leu Pro Arg Lys Pro Glu Glu Val Asp Asp Glu Val Phe Tyr 340 345
350Ser Pro Arg Ser Leu Val Phe Pro Glu Ala Glu Asn Arg Lys Trp Thr
355 360 365Ile Met Ala Val Met Val Ser Leu Leu Thr Asp Tyr Ser Pro
Gln Leu 370 375 380Gln Lys Pro Lys Phe38548362PRTArtificial
SequenceTAT-MTS(otc)-OTC delta linker 48Arg Lys Lys Arg Arg Gln Arg
Arg Arg Leu Phe Asn Leu Arg Ile Leu1 5 10 15Leu Asn Asn Ala Ala Phe
Arg Asn Gly His Asn Phe Met Val Arg Asn 20 25 30Phe Arg Cys Gly Gln
Pro Leu Gln Asn Lys Val Gln Leu Lys Gly Arg 35 40 45Asp Leu Leu Thr
Leu Arg Asn Phe Thr Gly Glu Glu Ile Lys Tyr Met 50 55 60Leu Trp Leu
Ser Ala Asp Leu Lys Phe Arg Ile Lys Gln Lys Gly Glu65 70 75 80Tyr
Leu Pro Leu Leu Gln Gly Lys Ser Leu Gly Met Ile Phe Glu Lys 85 90
95Arg Ser Thr Arg Thr Arg Leu Ser Thr Glu Thr Gly Phe Ala Leu Leu
100 105 110Gly Gly His Pro Cys Phe Leu Thr Thr Gln Asp Ile His Leu
Gly Val 115 120 125Asn Glu Ser Leu Thr Asp Thr Ala Arg Val Leu Ser
Ser Met Ala Asp 130 135 140Ala Val Leu Ala Arg Val Tyr Lys Gln Ser
Asp Leu Asp Thr Leu Ala145 150 155 160Lys Glu Ala Ser Ile Pro Ile
Ile Asn Gly Leu Ser Asp Leu Tyr His 165 170 175Pro Ile Gln Ile Leu
Ala Asp Tyr Leu Thr Leu Gln Glu His Tyr Ser 180 185 190Ser Leu Lys
Gly Leu Thr Leu Ser Trp Ile Gly Asp Gly Asn Asn Ile 195 200 205Leu
His Ser Ile Met Met Ser Ala Ala Lys Phe Gly Met His Leu Gln 210 215
220Ala Ala Thr Pro Lys Gly Tyr Glu Pro Asp Ala Ser Val Thr Lys
Leu225 230 235 240Ala Glu Gln Tyr Ala Lys Glu Asn Gly Thr Lys Leu
Leu Leu Thr Asn 245 250 255Asp Pro Leu Glu Ala Ala His Gly Gly Asn
Val Leu Ile Thr Asp Thr 260 265 270Trp Ile Ser Met Gly Gln Glu Glu
Glu Lys Lys Lys Arg Leu Gln Ala 275 280 285Phe Gln Gly Tyr Gln Val
Thr Met Lys Thr Ala Lys Val Ala Ala Ser 290 295 300Asp Trp Thr Phe
Leu His Cys Leu Pro Arg Lys Pro Glu Glu Val Asp305 310 315 320Asp
Glu Val Phe Tyr Ser Pro Arg Ser Leu Val Phe Pro Glu Ala Glu 325 330
335Asn Arg Lys Trp Thr Ile Met Ala Val Met Val Ser Leu Leu Thr Asp
340 345 350Tyr Ser Pro Gln Leu Gln Lys Pro Lys Phe 355
36049360PRTArtificial SequenceTAT-MTS(cs)-OTC 49Met Arg Lys Lys Arg
Arg Gln Arg Arg Arg Gly Ser Asp Pro Ala Leu1 5 10 15Leu Thr Ala Ala
Ala Arg Leu Leu Gly Thr Lys Asn Ala Ser Cys Leu 20 25 30Val Leu Ala
Ala Arg His Ala Ser Glu Phe Leu Lys Gly Arg Asp Leu 35 40 45Leu Thr
Leu Arg Asn Phe Thr Gly Glu Glu Ile Lys Tyr Met Leu Trp 50 55 60Leu
Ser Ala Asp Leu Lys Phe Arg Ile Lys Gln Lys Gly Glu Tyr Leu65 70 75
80Pro Leu Leu Gln Gly Lys Ser Leu Gly Met Ile Phe Glu Lys Arg Ser
85 90 95Thr Arg Thr Arg Leu Ser Thr Glu Thr Gly Phe Ala Leu Leu Gly
Gly 100 105 110His Pro Cys Phe Leu Thr Thr Gln Asp Ile His Leu Gly
Val Asn Glu 115 120 125Ser Leu Thr Asp Thr Ala Arg Val Leu Ser Ser
Met Ala Asp Ala Val 130 135 140Leu Ala Arg Val Tyr Lys Gln Ser Asp
Leu Asp Thr Leu Ala Lys Glu145 150 155 160Ala Ser Ile Pro Ile Ile
Asn Gly Leu Ser Asp Leu Tyr His Pro Ile 165 170 175Gln Ile Leu Ala
Asp Tyr Leu Thr Leu Gln Glu His Tyr Ser Ser Leu 180 185 190Lys Gly
Leu Thr Leu Ser Trp Ile Gly Asp Gly Asn Asn Ile Leu His 195 200
205Ser Ile Met Met Ser Ala Ala Lys Phe Gly Met His Leu Gln Ala Ala
210 215 220Thr Pro Lys Gly Tyr Glu Pro Asp Ala Ser Val Thr Lys Leu
Ala Glu225 230 235 240Gln Tyr Ala Lys Glu Asn Gly Thr Lys Leu Leu
Leu Thr Asn Asp Pro 245 250 255Leu Glu Ala Ala His Gly Gly Asn Val
Leu Ile Thr Asp Thr Trp Ile 260 265 270Ser Met Gly Gln Glu Glu Glu
Lys Lys Lys Arg Leu Gln Ala Phe Gln 275 280 285Gly Tyr Gln Val Thr
Met Lys Thr Ala Lys Val Ala Ala Ser Asp Trp 290 295 300Thr Phe Leu
His Cys Leu Pro Arg Lys Pro Glu Glu Val Asp Asp Glu305 310 315
320Val Phe Tyr Ser Pro Arg Ser Leu Val Phe Pro Glu Ala Glu Asn Arg
325 330 335Lys Trp Thr Ile Met Ala Val Met Val Ser Leu Leu Thr Asp
Tyr Ser 340 345 350Pro Gln Leu Gln Lys Pro Lys Phe 355
36050368PRTArtificial SequenceTAT-MTS(orf)-OTC 50Met Arg Lys Lys
Arg Arg Gln Arg Arg Arg Gly Ser Asp Pro Gly Ala1 5 10 15Leu Val Ile
Arg Gly Ile Arg Asn Phe Asn Leu Glu Asn Arg Ala Glu 20 25 30Arg Glu
Ile Ser Lys Met Lys Pro Ser Val Ala Pro Arg His Pro Ser 35 40 45Glu
Phe Leu Lys Gly Arg Asp Leu Leu Thr Leu Arg Asn Phe Thr Gly 50 55
60Glu Glu Ile Lys Tyr Met Leu Trp Leu Ser Ala Asp Leu Lys Phe Arg65
70 75 80Ile Lys Gln Lys Gly Glu Tyr Leu Pro Leu Leu Gln Gly Lys Ser
Leu 85 90 95Gly Met Ile Phe Glu Lys Arg Ser Thr Arg Thr Arg Leu Ser
Thr Glu 100 105 110Thr Gly Phe Ala Leu Leu Gly Gly His Pro Cys Phe
Leu Thr Thr Gln 115 120 125Asp Ile His Leu Gly Val Asn Glu Ser Leu
Thr Asp Thr Ala Arg Val 130 135 140Leu Ser Ser Met Ala Asp Ala Val
Leu Ala Arg Val Tyr Lys Gln Ser145 150 155 160Asp Leu Asp Thr Leu
Ala Lys Glu Ala Ser Ile Pro Ile Ile Asn Gly 165 170 175Leu Ser Asp
Leu Tyr His Pro Ile Gln Ile Leu Ala Asp Tyr Leu Thr 180 185 190Leu
Gln Glu His Tyr Ser Ser Leu Lys Gly Leu Thr Leu Ser Trp Ile 195 200
205Gly Asp Gly Asn Asn Ile Leu His Ser Ile Met Met Ser Ala Ala Lys
210 215 220Phe Gly Met His Leu Gln Ala Ala Thr Pro Lys Gly Tyr Glu
Pro Asp225 230 235 240Ala Ser Val Thr Lys Leu Ala Glu Gln Tyr Ala
Lys Glu Asn Gly Thr 245 250 255Lys Leu Leu Leu Thr Asn Asp Pro Leu
Glu Ala Ala His Gly Gly Asn 260 265 270Val Leu Ile Thr Asp Thr Trp
Ile Ser Met Gly Gln Glu Glu Glu Lys 275 280 285Lys Lys Arg Leu Gln
Ala Phe Gln Gly Tyr Gln Val Thr Met Lys Thr 290 295 300Ala Lys Val
Ala Ala Ser Asp Trp Thr Phe Leu His Cys Leu Pro Arg305 310 315
320Lys Pro Glu Glu Val Asp Asp Glu Val Phe Tyr Ser Pro Arg Ser Leu
325 330 335Val Phe Pro Glu Ala Glu Asn Arg Lys Trp Thr Ile Met Ala
Val Met 340 345 350Val Ser Leu Leu Thr Asp Tyr Ser Pro Gln Leu Gln
Lys Pro Lys Phe 355 360 36551369PRTArtificial
SequenceTAT-MTS(lad)-OTC 51Met Arg Lys Lys Arg Arg Gln Arg Arg Arg
Gly Ser Asp Pro Gln Ser1 5 10 15Trp Ser Arg Val Tyr Cys Ser Leu Ala
Lys Arg Gly His Phe Asn Arg 20 25 30Ile Ser His Gly Leu Gln Gly Leu
Ser Ala Val Pro Leu Arg Thr Tyr 35 40 45Ala Glu Phe Leu Lys Gly Arg
Asp Leu Leu Thr Leu Arg Asn Phe Thr 50 55 60Gly Glu Glu Ile Lys Tyr
Met Leu Trp Leu Ser Ala Asp Leu Lys Phe65 70 75 80Arg Ile Lys Gln
Lys Gly Glu Tyr Leu Pro Leu Leu Gln Gly Lys Ser 85 90 95Leu Gly Met
Ile Phe Glu Lys Arg Ser Thr Arg Thr Arg Leu Ser Thr 100 105 110Glu
Thr Gly Phe Ala Leu Leu Gly Gly His Pro Cys Phe Leu Thr Thr 115 120
125Gln Asp Ile His Leu Gly Val Asn Glu Ser Leu Thr Asp Thr Ala Arg
130 135 140Val Leu Ser Ser Met Ala Asp Ala Val Leu Ala Arg Val Tyr
Lys Gln145 150 155 160Ser Asp Leu Asp Thr Leu Ala Lys Glu Ala Ser
Ile Pro Ile Ile Asn 165 170 175Gly Leu Ser Asp Leu Tyr His Pro Ile
Gln Ile Leu Ala Asp Tyr Leu 180 185
190Thr Leu Gln Glu His Tyr Ser Ser Leu Lys Gly Leu Thr Leu Ser Trp
195 200 205Ile Gly Asp Gly Asn Asn Ile Leu His Ser Ile Met Met Ser
Ala Ala 210 215 220Lys Phe Gly Met His Leu Gln Ala Ala Thr Pro Lys
Gly Tyr Glu Pro225 230 235 240Asp Ala Ser Val Thr Lys Leu Ala Glu
Gln Tyr Ala Lys Glu Asn Gly 245 250 255Thr Lys Leu Leu Leu Thr Asn
Asp Pro Leu Glu Ala Ala His Gly Gly 260 265 270Asn Val Leu Ile Thr
Asp Thr Trp Ile Ser Met Gly Gln Glu Glu Glu 275 280 285Lys Lys Lys
Arg Leu Gln Ala Phe Gln Gly Tyr Gln Val Thr Met Lys 290 295 300Thr
Ala Lys Val Ala Ala Ser Asp Trp Thr Phe Leu His Cys Leu Pro305 310
315 320Arg Lys Pro Glu Glu Val Asp Asp Glu Val Phe Tyr Ser Pro Arg
Ser 325 330 335Leu Val Phe Pro Glu Ala Glu Asn Arg Lys Trp Thr Ile
Met Ala Val 340 345 350Met Val Ser Leu Leu Thr Asp Tyr Ser Pro Gln
Leu Gln Lys Pro Lys 355 360 365Phe52356PRTArtificial
SequenceTAT-MTS(cs)-OTC delta linker 52Met Arg Lys Lys Arg Arg Gln
Arg Arg Arg Ala Leu Leu Thr Ala Ala1 5 10 15Ala Arg Leu Leu Gly Thr
Lys Asn Ala Ser Cys Leu Val Leu Ala Ala 20 25 30Arg His Ala Ser Glu
Phe Leu Lys Gly Arg Asp Leu Leu Thr Leu Arg 35 40 45Asn Phe Thr Gly
Glu Glu Ile Lys Tyr Met Leu Trp Leu Ser Ala Asp 50 55 60Leu Lys Phe
Arg Ile Lys Gln Lys Gly Glu Tyr Leu Pro Leu Leu Gln65 70 75 80Gly
Lys Ser Leu Gly Met Ile Phe Glu Lys Arg Ser Thr Arg Thr Arg 85 90
95Leu Ser Thr Glu Thr Gly Phe Ala Leu Leu Gly Gly His Pro Cys Phe
100 105 110Leu Thr Thr Gln Asp Ile His Leu Gly Val Asn Glu Ser Leu
Thr Asp 115 120 125Thr Ala Arg Val Leu Ser Ser Met Ala Asp Ala Val
Leu Ala Arg Val 130 135 140Tyr Lys Gln Ser Asp Leu Asp Thr Leu Ala
Lys Glu Ala Ser Ile Pro145 150 155 160Ile Ile Asn Gly Leu Ser Asp
Leu Tyr His Pro Ile Gln Ile Leu Ala 165 170 175Asp Tyr Leu Thr Leu
Gln Glu His Tyr Ser Ser Leu Lys Gly Leu Thr 180 185 190Leu Ser Trp
Ile Gly Asp Gly Asn Asn Ile Leu His Ser Ile Met Met 195 200 205Ser
Ala Ala Lys Phe Gly Met His Leu Gln Ala Ala Thr Pro Lys Gly 210 215
220Tyr Glu Pro Asp Ala Ser Val Thr Lys Leu Ala Glu Gln Tyr Ala
Lys225 230 235 240Glu Asn Gly Thr Lys Leu Leu Leu Thr Asn Asp Pro
Leu Glu Ala Ala 245 250 255His Gly Gly Asn Val Leu Ile Thr Asp Thr
Trp Ile Ser Met Gly Gln 260 265 270Glu Glu Glu Lys Lys Lys Arg Leu
Gln Ala Phe Gln Gly Tyr Gln Val 275 280 285Thr Met Lys Thr Ala Lys
Val Ala Ala Ser Asp Trp Thr Phe Leu His 290 295 300Cys Leu Pro Arg
Lys Pro Glu Glu Val Asp Asp Glu Val Phe Tyr Ser305 310 315 320Pro
Arg Ser Leu Val Phe Pro Glu Ala Glu Asn Arg Lys Trp Thr Ile 325 330
335Met Ala Val Met Val Ser Leu Leu Thr Asp Tyr Ser Pro Gln Leu Gln
340 345 350Lys Pro Lys Phe 35553364PRTArtificial
SequenceTAT-MTS(orf)-OTC delta linker 53Met Arg Lys Lys Arg Arg Gln
Arg Arg Arg Gly Ala Leu Val Ile Arg1 5 10 15Gly Ile Arg Asn Phe Asn
Leu Glu Asn Arg Ala Glu Arg Glu Ile Ser 20 25 30Lys Met Lys Pro Ser
Val Ala Pro Arg His Pro Ser Glu Phe Leu Lys 35 40 45Gly Arg Asp Leu
Leu Thr Leu Arg Asn Phe Thr Gly Glu Glu Ile Lys 50 55 60Tyr Met Leu
Trp Leu Ser Ala Asp Leu Lys Phe Arg Ile Lys Gln Lys65 70 75 80Gly
Glu Tyr Leu Pro Leu Leu Gln Gly Lys Ser Leu Gly Met Ile Phe 85 90
95Glu Lys Arg Ser Thr Arg Thr Arg Leu Ser Thr Glu Thr Gly Phe Ala
100 105 110Leu Leu Gly Gly His Pro Cys Phe Leu Thr Thr Gln Asp Ile
His Leu 115 120 125Gly Val Asn Glu Ser Leu Thr Asp Thr Ala Arg Val
Leu Ser Ser Met 130 135 140Ala Asp Ala Val Leu Ala Arg Val Tyr Lys
Gln Ser Asp Leu Asp Thr145 150 155 160Leu Ala Lys Glu Ala Ser Ile
Pro Ile Ile Asn Gly Leu Ser Asp Leu 165 170 175Tyr His Pro Ile Gln
Ile Leu Ala Asp Tyr Leu Thr Leu Gln Glu His 180 185 190Tyr Ser Ser
Leu Lys Gly Leu Thr Leu Ser Trp Ile Gly Asp Gly Asn 195 200 205Asn
Ile Leu His Ser Ile Met Met Ser Ala Ala Lys Phe Gly Met His 210 215
220Leu Gln Ala Ala Thr Pro Lys Gly Tyr Glu Pro Asp Ala Ser Val
Thr225 230 235 240Lys Leu Ala Glu Gln Tyr Ala Lys Glu Asn Gly Thr
Lys Leu Leu Leu 245 250 255Thr Asn Asp Pro Leu Glu Ala Ala His Gly
Gly Asn Val Leu Ile Thr 260 265 270Asp Thr Trp Ile Ser Met Gly Gln
Glu Glu Glu Lys Lys Lys Arg Leu 275 280 285Gln Ala Phe Gln Gly Tyr
Gln Val Thr Met Lys Thr Ala Lys Val Ala 290 295 300Ala Ser Asp Trp
Thr Phe Leu His Cys Leu Pro Arg Lys Pro Glu Glu305 310 315 320Val
Asp Asp Glu Val Phe Tyr Ser Pro Arg Ser Leu Val Phe Pro Glu 325 330
335Ala Glu Asn Arg Lys Trp Thr Ile Met Ala Val Met Val Ser Leu Leu
340 345 350Thr Asp Tyr Ser Pro Gln Leu Gln Lys Pro Lys Phe 355
36054365PRTArtificial SequenceTAT-MTS(lad)-OTC delta linker 54Met
Arg Lys Lys Arg Arg Gln Arg Arg Arg Gln Ser Trp Ser Arg Val1 5 10
15Tyr Cys Ser Leu Ala Lys Arg Gly His Phe Asn Arg Ile Ser His Gly
20 25 30Leu Gln Gly Leu Ser Ala Val Pro Leu Arg Thr Tyr Ala Glu Phe
Leu 35 40 45Lys Gly Arg Asp Leu Leu Thr Leu Arg Asn Phe Thr Gly Glu
Glu Ile 50 55 60Lys Tyr Met Leu Trp Leu Ser Ala Asp Leu Lys Phe Arg
Ile Lys Gln65 70 75 80Lys Gly Glu Tyr Leu Pro Leu Leu Gln Gly Lys
Ser Leu Gly Met Ile 85 90 95Phe Glu Lys Arg Ser Thr Arg Thr Arg Leu
Ser Thr Glu Thr Gly Phe 100 105 110Ala Leu Leu Gly Gly His Pro Cys
Phe Leu Thr Thr Gln Asp Ile His 115 120 125Leu Gly Val Asn Glu Ser
Leu Thr Asp Thr Ala Arg Val Leu Ser Ser 130 135 140Met Ala Asp Ala
Val Leu Ala Arg Val Tyr Lys Gln Ser Asp Leu Asp145 150 155 160Thr
Leu Ala Lys Glu Ala Ser Ile Pro Ile Ile Asn Gly Leu Ser Asp 165 170
175Leu Tyr His Pro Ile Gln Ile Leu Ala Asp Tyr Leu Thr Leu Gln Glu
180 185 190His Tyr Ser Ser Leu Lys Gly Leu Thr Leu Ser Trp Ile Gly
Asp Gly 195 200 205Asn Asn Ile Leu His Ser Ile Met Met Ser Ala Ala
Lys Phe Gly Met 210 215 220His Leu Gln Ala Ala Thr Pro Lys Gly Tyr
Glu Pro Asp Ala Ser Val225 230 235 240Thr Lys Leu Ala Glu Gln Tyr
Ala Lys Glu Asn Gly Thr Lys Leu Leu 245 250 255Leu Thr Asn Asp Pro
Leu Glu Ala Ala His Gly Gly Asn Val Leu Ile 260 265 270Thr Asp Thr
Trp Ile Ser Met Gly Gln Glu Glu Glu Lys Lys Lys Arg 275 280 285Leu
Gln Ala Phe Gln Gly Tyr Gln Val Thr Met Lys Thr Ala Lys Val 290 295
300Ala Ala Ser Asp Trp Thr Phe Leu His Cys Leu Pro Arg Lys Pro
Glu305 310 315 320Glu Val Asp Asp Glu Val Phe Tyr Ser Pro Arg Ser
Leu Val Phe Pro 325 330 335Glu Ala Glu Asn Arg Lys Trp Thr Ile Met
Ala Val Met Val Ser Leu 340 345 350Leu Thr Asp Tyr Ser Pro Gln Leu
Gln Lys Pro Lys Phe 355 360 365
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014
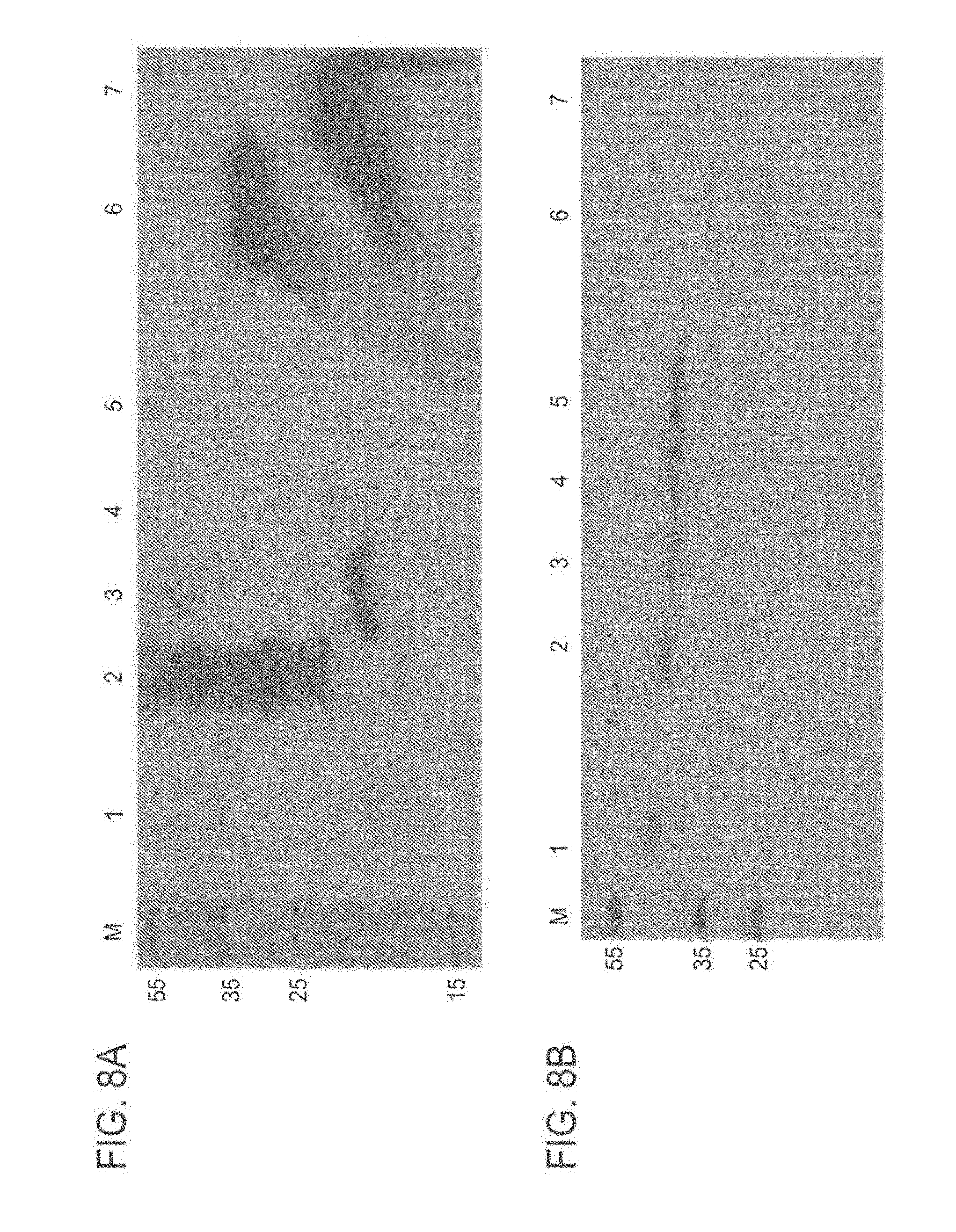
D00015

D00016
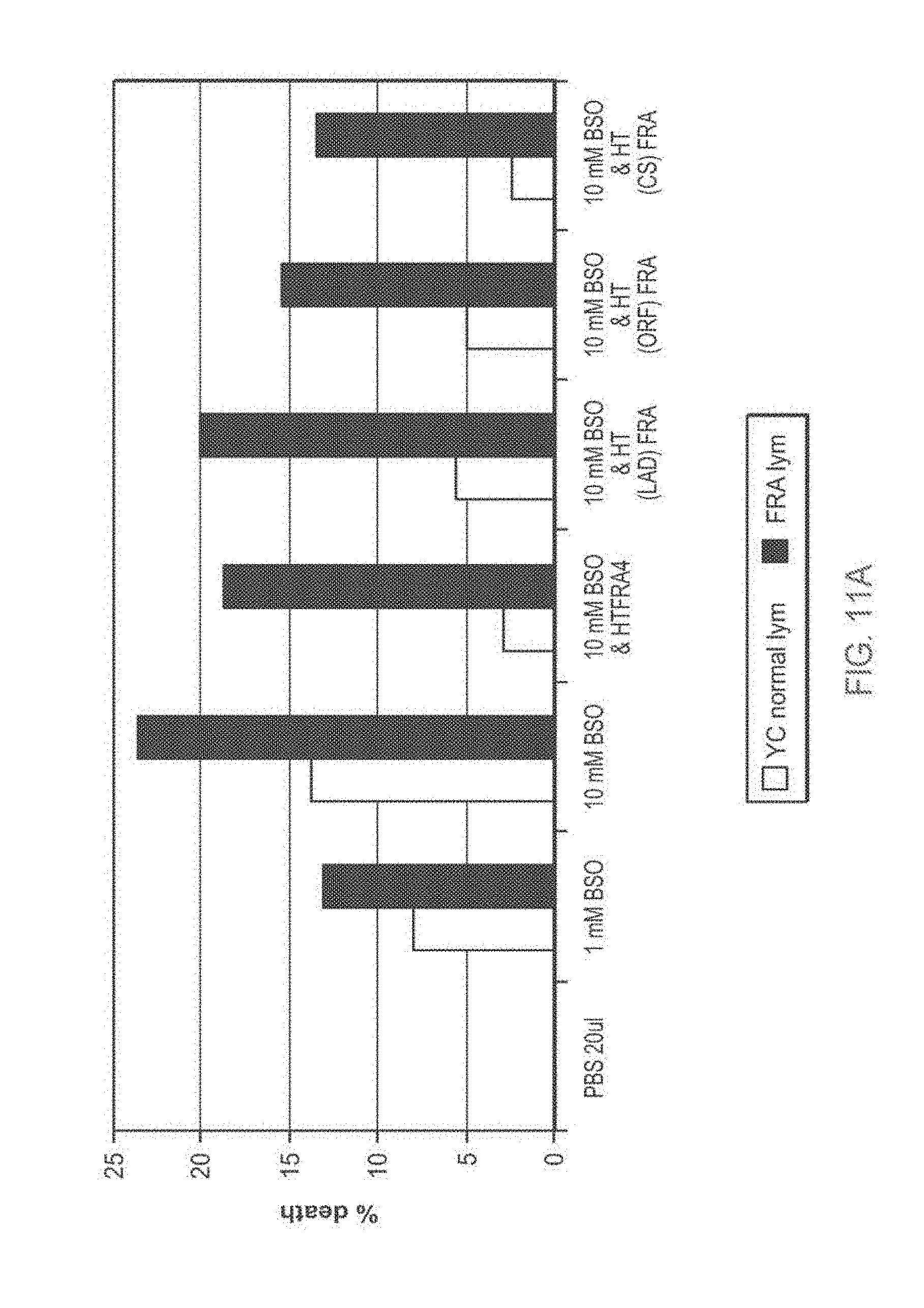

D00017
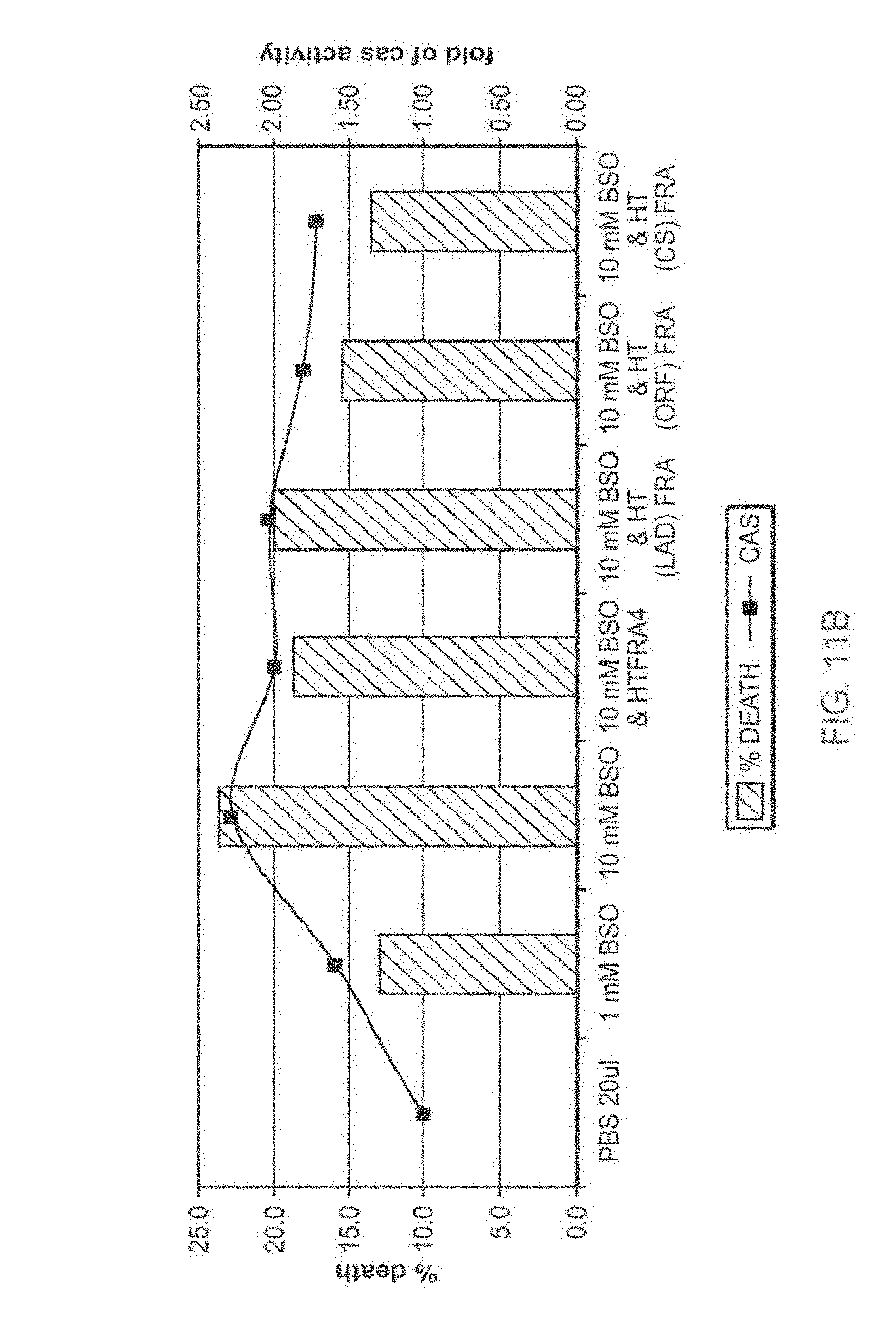
D00018
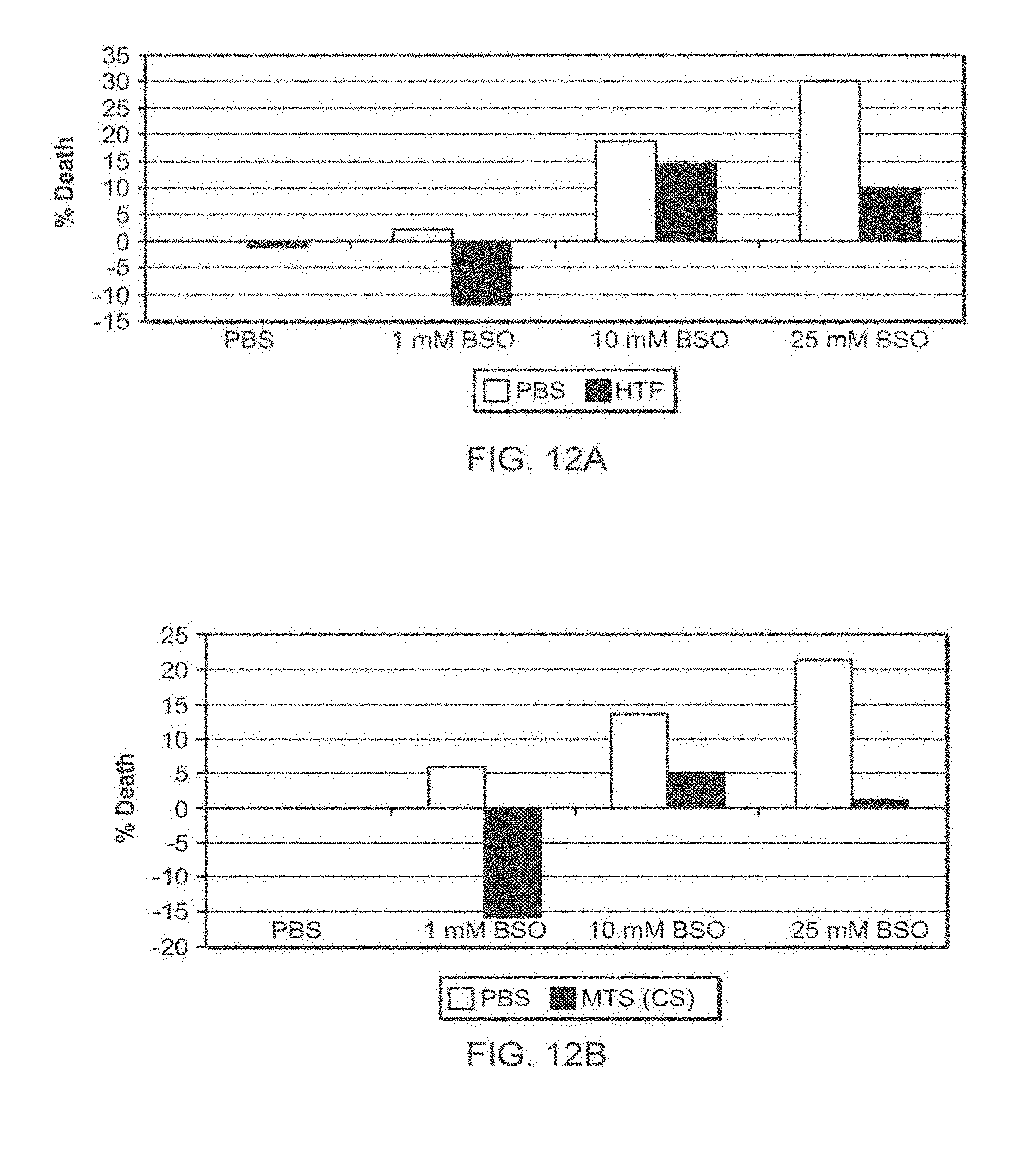
D00019

D00020
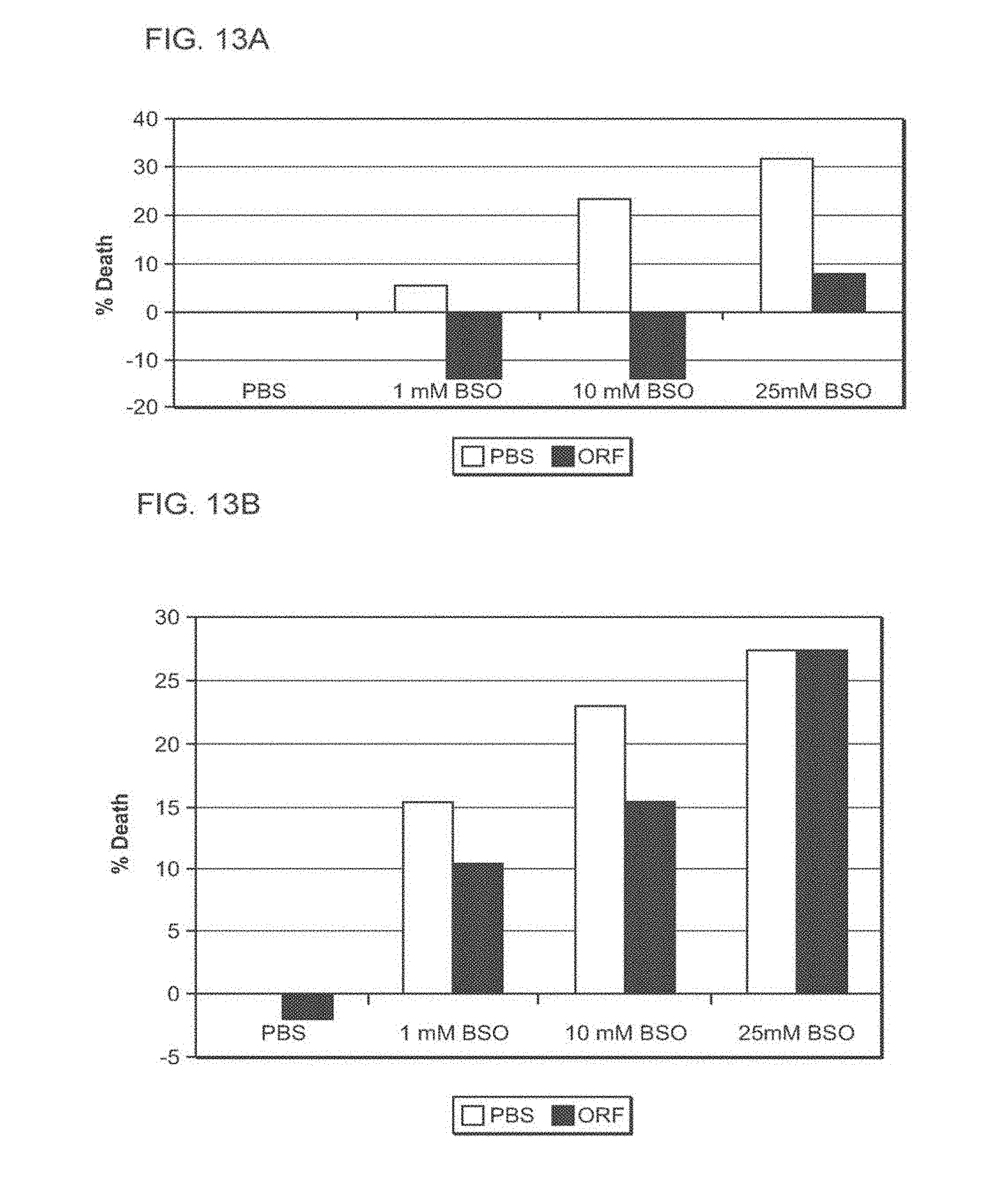
D00021
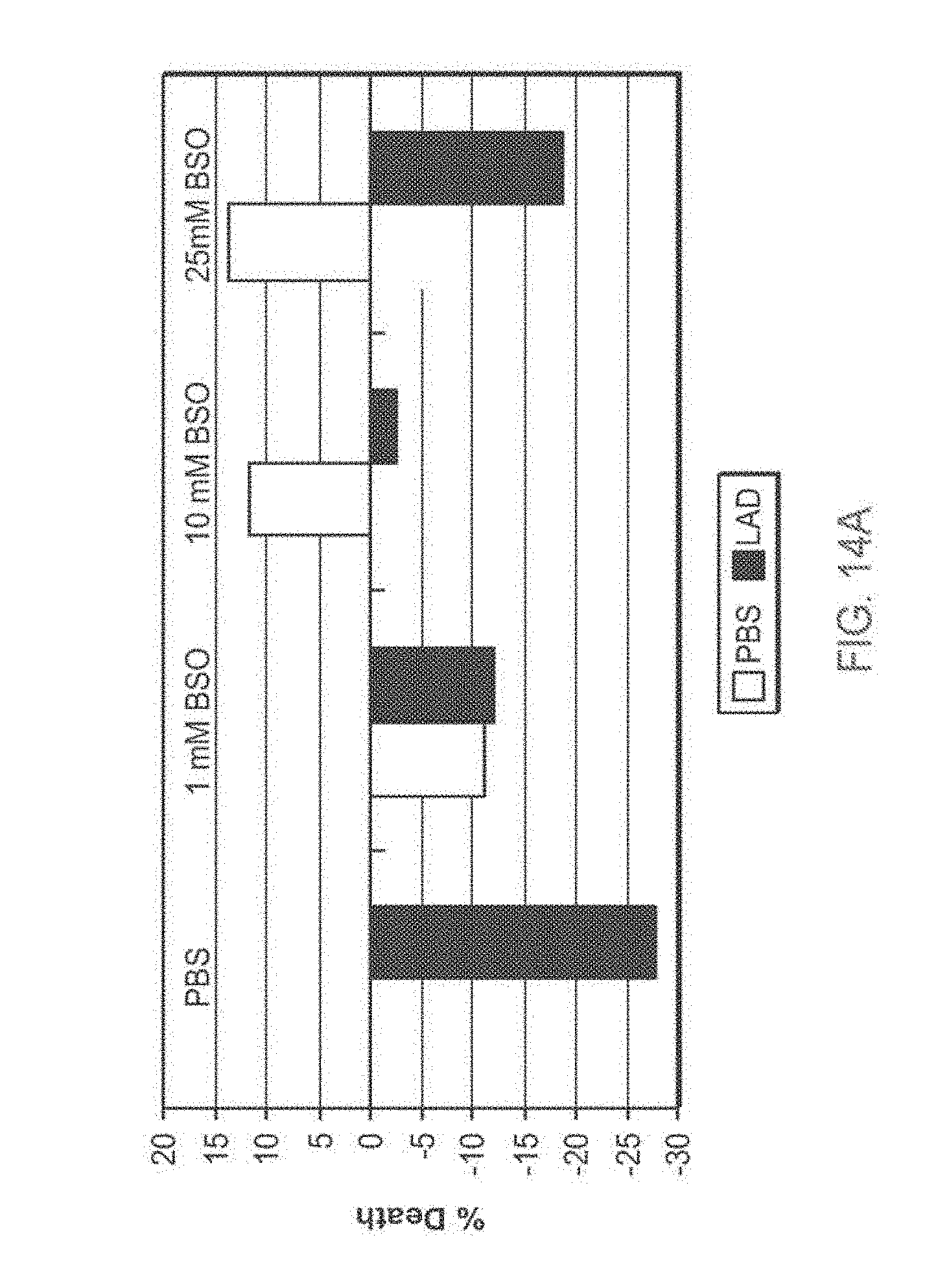
D00022
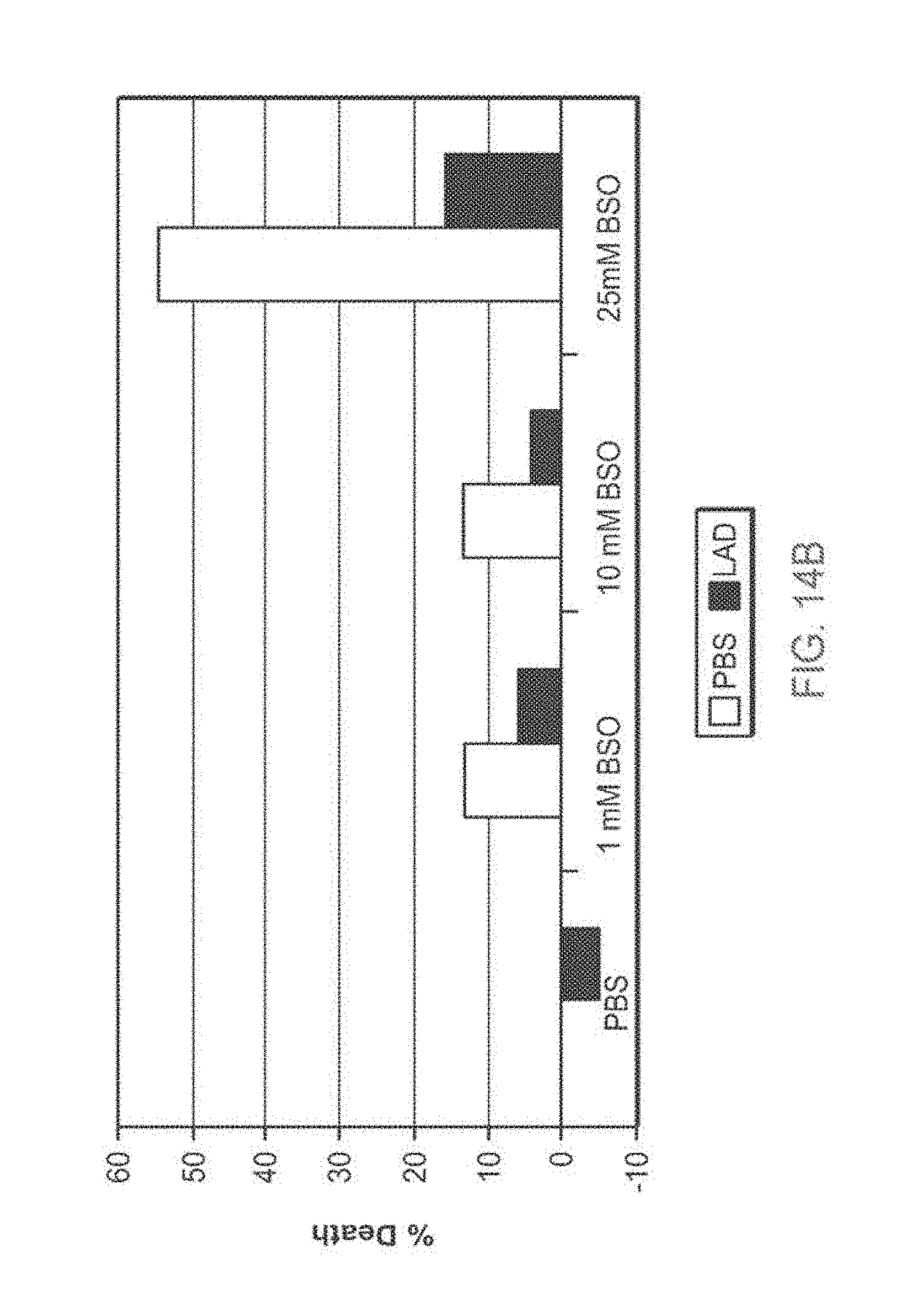
D00023

D00024
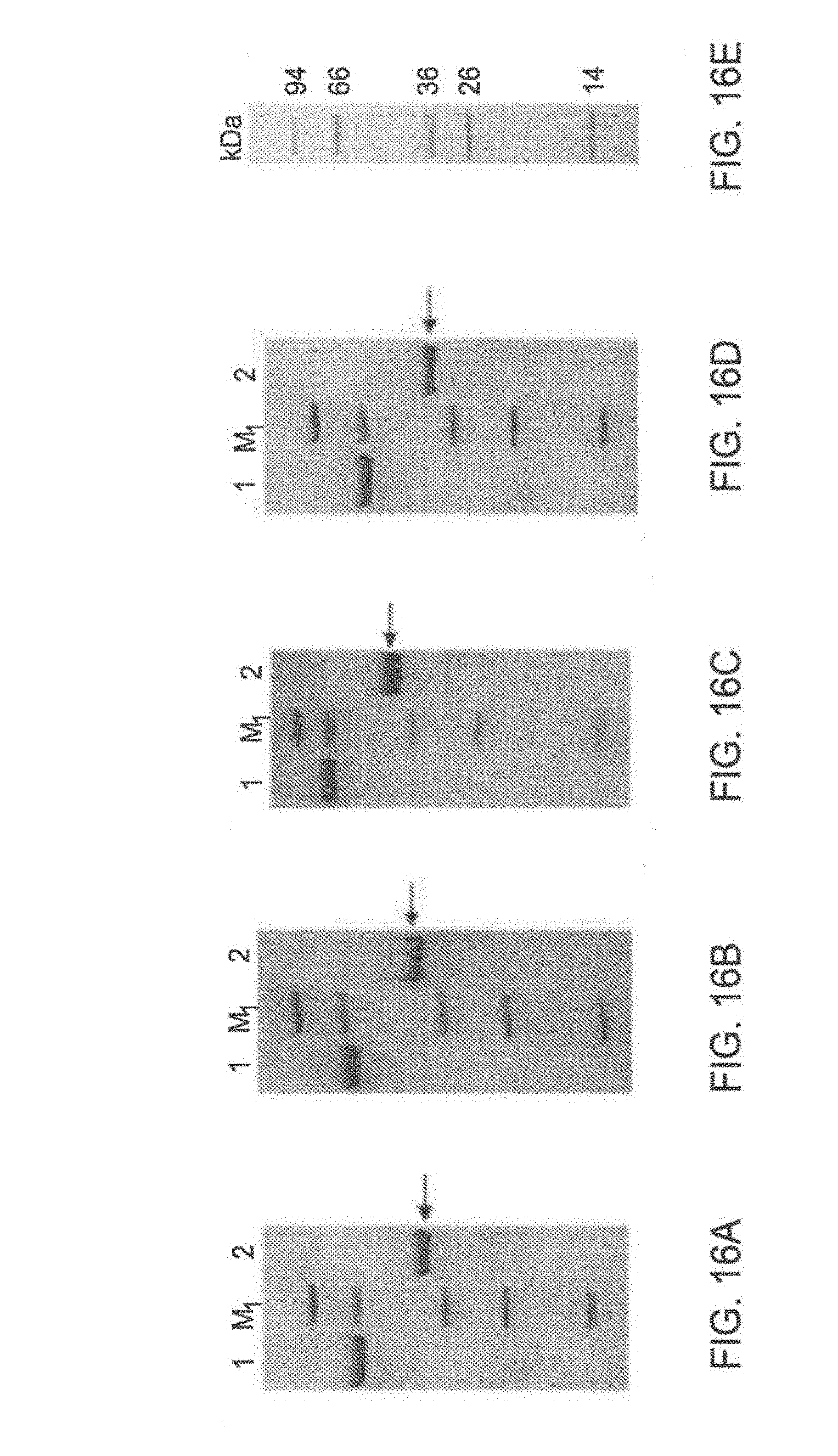
D00025
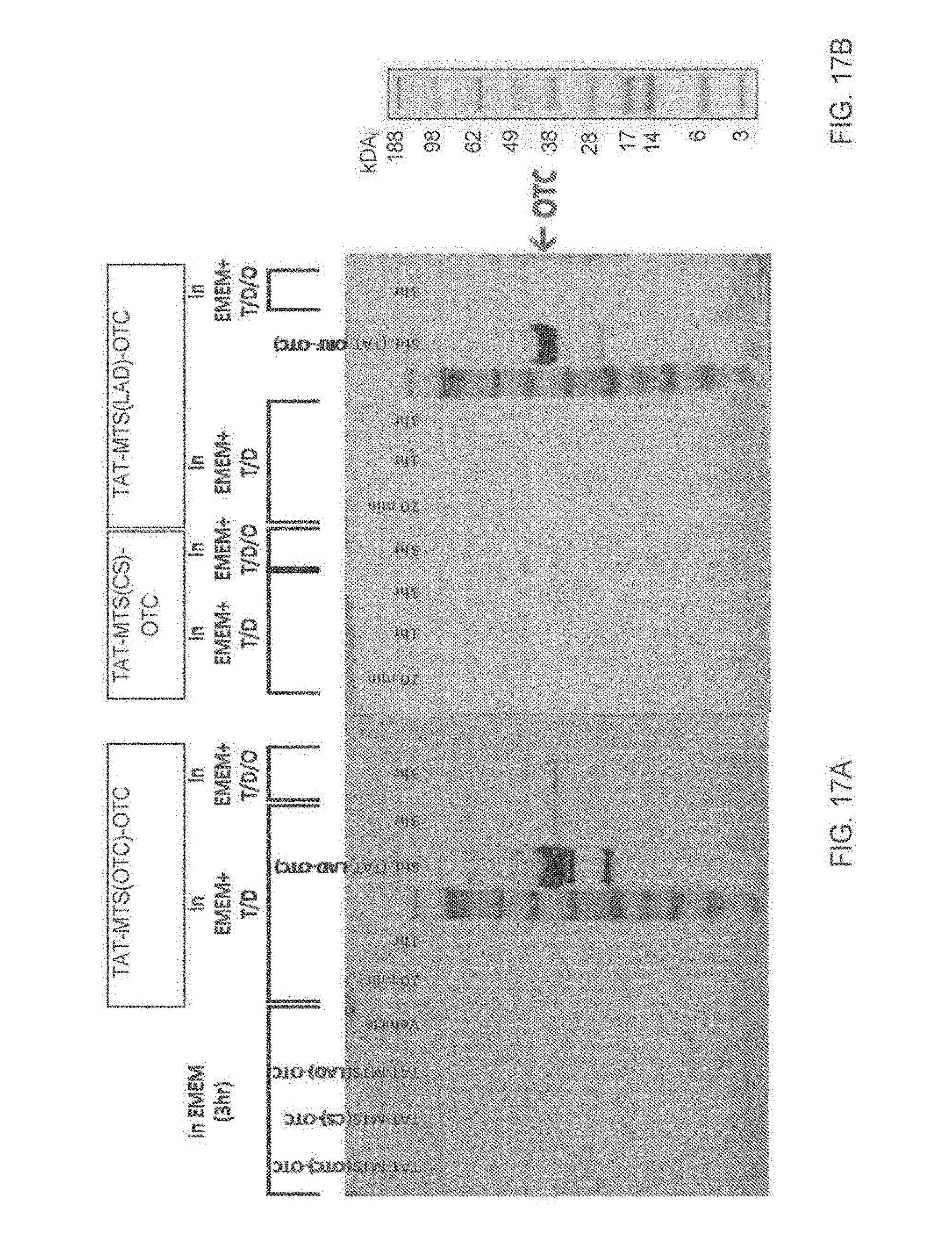
D00026

D00027
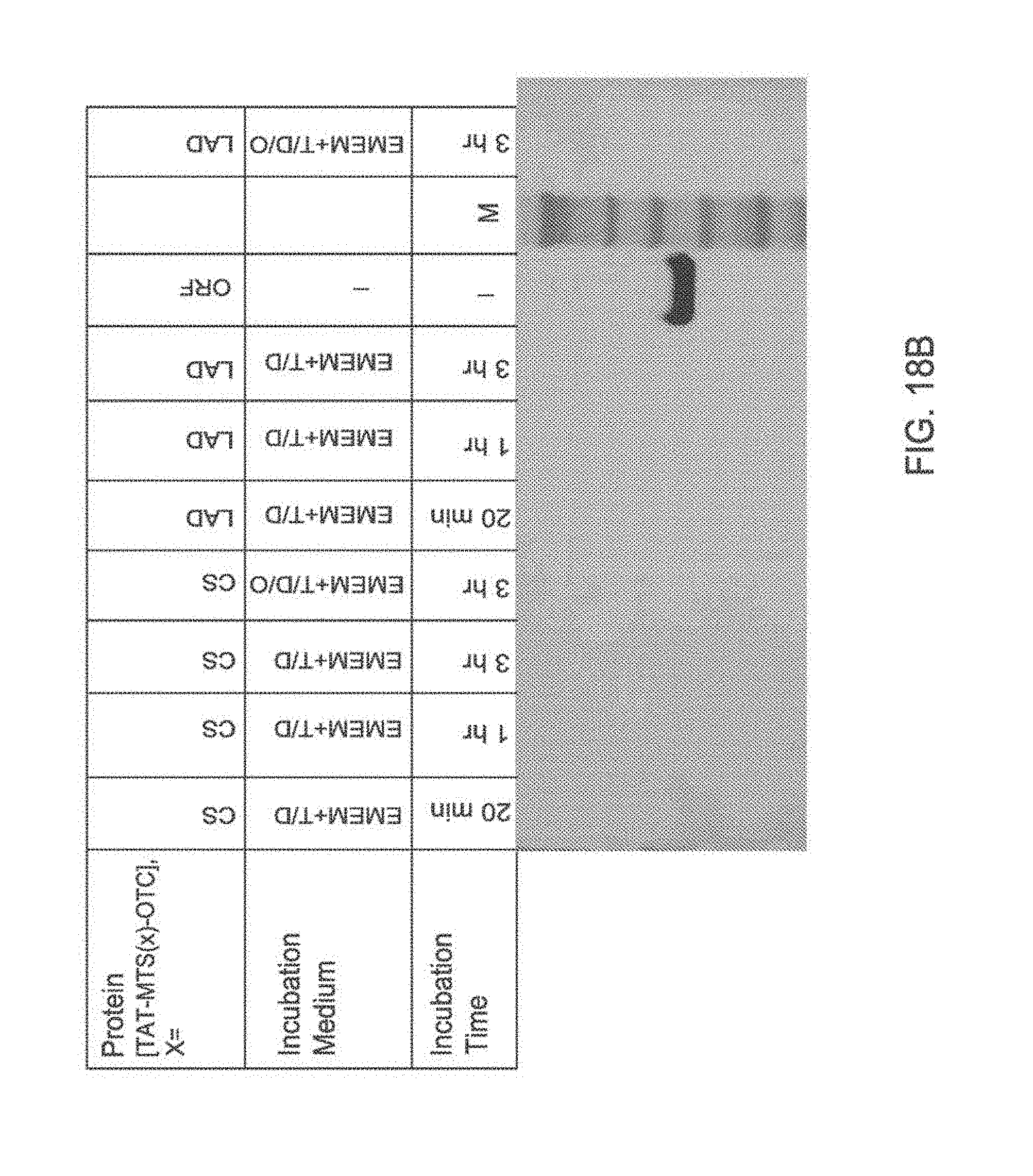
D00028
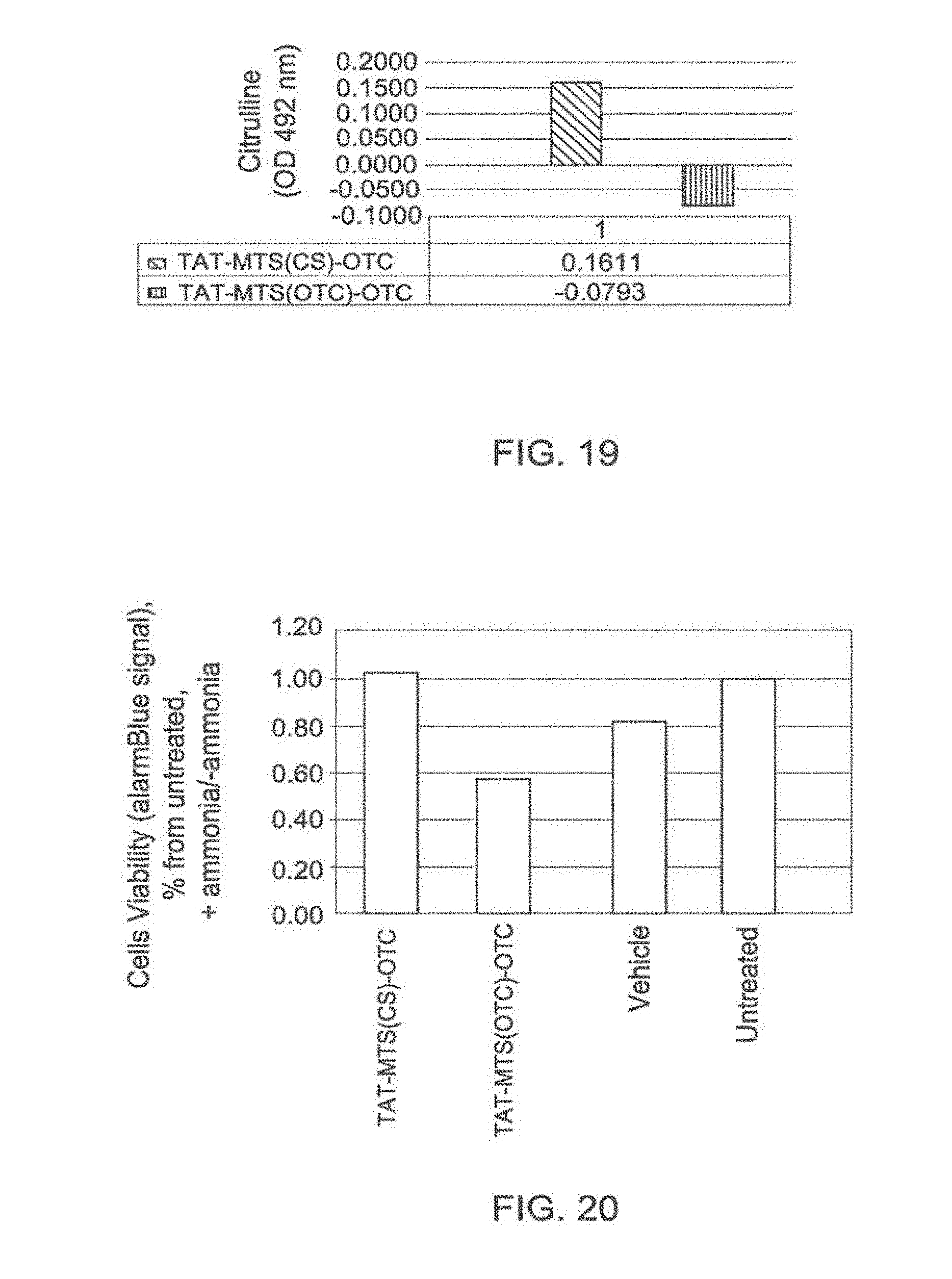
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.