Generation of Inner Ear Cells
Edge; Albert
U.S. patent application number 16/273071 was filed with the patent office on 2019-08-01 for generation of inner ear cells. The applicant listed for this patent is Massachusetts Eye & Ear Infirmary. Invention is credited to Albert Edge.
| Application Number | 20190233796 16/273071 |
| Document ID | / |
| Family ID | 39536941 |
| Filed Date | 2019-08-01 |








| United States Patent Application | 20190233796 |
| Kind Code | A1 |
| Edge; Albert | August 1, 2019 |
Generation of Inner Ear Cells
Abstract
Methods for generating cells of the inner ear, e.g., hair cells and supporting cells, from stem cells, e.g., mesenchymal stem cells, are provided, as well as compositions including the inner ear cells. Methods for the therapeutic use of the inner ear cells for the treatment of hearing loss are also described.
| Inventors: | Edge; Albert; (Brookline, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 39536941 | ||||||||||
| Appl. No.: | 16/273071 | ||||||||||
| Filed: | February 11, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15876899 | Jan 22, 2018 | |||
| 16273071 | ||||
| 14833919 | Aug 24, 2015 | 9896658 | ||
| 15876899 | ||||
| 13759441 | Feb 5, 2013 | |||
| 14833919 | ||||
| 12233017 | Sep 18, 2008 | |||
| 13759441 | ||||
| PCT/US2007/084654 | Nov 14, 2007 | |||
| 12233017 | ||||
| 60859041 | Nov 15, 2006 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 5/062 20130101; A61P 27/16 20180101; C12N 5/0627 20130101; C12N 2506/1353 20130101; C12N 2501/42 20130101 |
| International Class: | C12N 5/071 20060101 C12N005/071; C12N 5/0793 20060101 C12N005/0793 |
Goverment Interests
FEDERALLY SPONSORED RESEARCH OR DEVELOPMENT
[0002] This invention was made with Government support under Grant No. F33 DC006789, RO1 DC007174, and P30 DC05209 from the National Institute on Deafness and other Communicative Disorders (NIDCD) of the National Institutes of Health. The Government has certain rights in the invention.
Claims
1. (canceled)
2. A method of increasing numbers of differentiated inner ear cells in the ear of a subject, the method comprising: generating a differentiated inner ear cell, using a method comprising: (i) providing a population of mesenchymal stem cells obtained from mammalian bone marrow; (ii) culturing the mesenchymal stem cell in serum-free medium containing insulin-like growth factor-1 (IGF-1), epidermal growth factor (EGF), and basic fibroblast growth factor (bFGF); (iii) inducing the stem cell by maintaining said cell in medium comprising neurotrophin-3 (NT-3) or brain derived neurotrophic factor (BDNF) for time sufficient to differentiate into a progenitor cell that expresses Sox2, Pax6, nestin, and musashih; and (iv) culturing the progenitor cell in the presence of a gamma secretase inhibitor, wherein the gamma secretase inhibitor is selected from the group consisting of an arylsulfonamide, a dibenzazepine, a benzodiazepine, N--[N-(3,5-difluorophenacetyl)-L-alanyl]-(S)-phenylglycine t-butyl ester (DAPT), L-685,458, and MK0752, in an amount and for a time sufficient to produce a differentiated inner ear auditory hair cell that expresses myosin VIIa, and espin and optionally expresses one or more of atonal homolog 1 (Atoh1), or jagged 2, thereby producing a differentiated inner ear auditory hair cell, and implanting the differentiated inner ear auditory hair cell into the ear of the subject.
3. The method of claim 2, wherein the mesenchymal stem cell is obtained from a subject who has sensorineural hair cell loss.
4. The method of claim 2, wherein the differentiated cell is an inner ear auditory hair cell.
5. The method of claim 4, wherein the inner ear auditory hair cell expresses Atoh1, myosin7a, and epsin.
Description
CLAIM OF PRIORITY
[0001] This application is continuation application of U.S. patent application Ser. No. 15/876,899, filed Jan. 22, 2018, which is a continuation of U.S. patent application Ser. No. 14/833,919, filed Aug. 24, 2015, now U.S. Pat. No. 9,896,658, which is a continuation of U.S. patent application Ser. No. 13/759,441, filed Feb. 5, 2013, which is a continuation application of U.S. patent application Ser. No. 12/233,017, filed Sep. 18, 2008, which is a continuation of International Patent Application No. PCT/US2007/084654, filed on Nov. 14, 2007, which claims the benefit of U.S. Provisional Patent Application Ser. No. 60/859,041, filed on Nov. 15, 2006; the entire contents of each of the foregoing applications are hereby incorporated by reference.
TECHNICAL FIELD
[0003] This invention relates to methods using bone marrow mesenchymal stem cells to regenerate inner ear cells, e.g., hair cells and supporting cells, to treat inner ear damage.
BACKGROUND
[0004] A source of sensory cells and neurons for regeneration of inner ear cells would provide a valuable tool for clinical application because neurons and hair cells could be employed in cell replacement therapy for hearing loss. Recent work has shown that hair cells and neurons can be differentiated from endogenous stem cells of the inner ear (Li et al., Nat Med 9, 1293-1299 (2003); Rask-Andersen et al., Hear Res 203, 180-191 (2005)) and other work has shown that endogenous cells of the sensory epithelium can be converted to hair cells when the proneural transcription factor, Atoh1, is expressed exogenously (Izumikawa et al., Nat Med 11, 271-276 (2005); Zheng and Gao, Nat Neurosci 3, 580-586 (2000)) and yet the endogenous stem cells of the inner ear do not spontaneously generate hair cells. Injection of whole bone marrow to reconstitute a lethally irradiated mouse resulted in engraftment of these cells in areas occupied by inner ear mesenchymal cells and fibrocytes but did not yield hair cells (Lang et al., J Comp Neurol 496, 187-201 (2006)).
SUMMARY
[0005] The present invention is based, at least in part, on the discovery of methods that can be used to induce stem cells to differentiate into hair cells and supporting cells. Thus, described herein are methods for providing populations of hair cells and/or supporting cells, compositions comprising said cells, and methods of use thereof, e.g., for the treatment of subjects who have or are at risk of developing a hearing loss.
[0006] In one aspect, the invention provides methods for providing populations of hair cells and/or supporting cells. The methods include:
[0007] obtaining a population of stem cells with neurogenic potential;
[0008] culturing the stem cells under conditions sufficient to induce the differentiation of at least some of the stem cells into inner ear progenitor cells, and doing one (or more) of the following: [0009] (i) inducing the expression of Atoh1 in the inner ear progenitor cells, in an amount and for a time sufficient to induce at least some of the inner ear progenitor cells to differentiate into hair cells; [0010] (ii) contacting the inner ear progenitor cells with an inhibitor of Notch signalling (e.g., a gamma-secretase inhibitor or inhibitory nucleic acid), in an amount and for a time sufficient to induce at least some of the inner ear progenitor cells to differentiate into hair cells; or [0011] (iii) culturing the inner ear progenitor cells in the presence of chick otocyst cells for a time and under conditions sufficient for at least some of the inner ear progenitor cells to differentiate into hair cells, thereby providing populations of hair cells and/or supporting cells.
[0012] In some embodiments, the methods include isolating the inner ear progenitor cells, hair cells, and/or supporting cells, e.g., to provide a purified population thereof.
[0013] In some embodiments, the inner ear progenitor cells express nestin, sox2, musashi, Brn3C, Pax2, and Atoh1.
[0014] In some embodiments, the hair cells express one or more genes selected from the group consisting of Atoh1, jagged 2, Brn3c, p27Kip, Ngn1, NeuroD, myosin VIIa and espin. In some embodiments, the hair cells express jagged 2, Brn3c, myosin VIIa and espin. In some embodiments, the hair cells express F-actin in a V pattern on the apical surface of the cells.
[0015] In some embodiments, the supporting cells express one or more of claudin14, connexin 26, p75.sup.Trk Notch 1, and S100A.
[0016] In some embodiments, the methods further include transplanting the hair cells or supporting cells into a subject in need thereof, e.g., into or near the sensory epithelium of the subject. In some embodiments, the population of stem cells is obtained from a subject in need of the transplant.
[0017] Also described herein are isolated populations of hair cells, supporting cells, and inner ear progenitor cells obtained by a method described herein.
[0018] In another aspect, the invention features methods for treating a subject who has or is at risk for developing a disorder, e.g., a hearing disorder or vestibular disorder, wherein the disorder is treatable with a transplant of hair cells and/or supporting cells, the method comprising transplanting cells obtained by a method described herein into the cochlea of the subject, thereby treating the subject. In these embodiments, it is preferable if the population of stem cells was obtained from the subject in need of the transplant.
[0019] In some embodiments, inducing the expression of Atoh1 in the cells comprises inducing the expression of exogenous Atoh1 in the cells, e.g., by transducing the cells with a vector encoding a Atoh1 polypeptide, e.g., a plasmid vector or a viral vector, e.g., an adenovirus, lentivirus, or retrovirus.
[0020] In some embodiments, inducing the expression of exogenous Atoh1 in the stem cells comprises increasing expression of endogenous Atoh1, e.g., by increasing activity of the Atoh1 promoter or by replacing the endogenous Atoh1 promoter with a more highly active promoter.
[0021] In some embodiments, culturing the stem cells in the presence of chick otocyst cells for a time and under conditions sufficient for at least some of the stem cells to differentiate into hair cells comprises culturing the stem cells in medium comprising IGF, EGF, and FGF.
[0022] In some embodiments, the stem cells used in the methods described herein are mesenchymal stem cells. In some embodiments, the stem cells used in the methods described herein are human stem cells.
[0023] As noted, the invention also features cells isolated by a method described herein, as well as compositions containing them.
[0024] Methods for treating subjects (e.g., mammals such as humans) who have, or who are at risk for developing, a hearing loss, are also described herein. These methods include administering a cell or population of cells (as described herein; e.g., a population of hair cells obtained by differentiating a population of stem cells) to the ear of the patient, e.g., to the cochlea. The administered cells may be obtained by the methods described herein, and the starting material may be stem cells obtained from the patient to be treated.
[0025] There may be certain advantages to the use of the cells described herein for the treatment of hearing loss. For example, the stem cells can be obtained from humans for clinical applications. Because the stem cells can be harvested from a human, and in particular can be harvested from the human in need of treatment, the immunological hurdles common in xeno- and allotransplantation experiments can be largely avoided.
[0026] Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Methods and materials are described herein for use in the present invention; other, suitable methods and materials known in the art can also be used. The materials, methods, and examples are illustrative only and not intended to be limiting. All publications, patent applications, patents, sequences, database entries, and other references mentioned herein are incorporated by reference in their entirety. In case of conflict, the present specification, including definitions, will control.
[0027] Other features and advantages of the invention will be apparent from the following detailed description and figures, and from the claims.
DESCRIPTION OF DRAWINGS
[0028] FIG. 1A is a row of four photomicrographs of bone marrow MSCs from passage 3 immunostained with antibodies against CD44, CD45, CD34 and Sca-1 followed by secondary antibodies against mouse immunoglobulins labeled with TRITC (medium gray, shown in red in the original). Staining for CD34 and CD45 was negative, but CD44 and Sca-1 were expressed. Nuclei were stained with DAPI (darker gray, blue in the original).
[0029] FIG. 1B is a row of four photomicrographs of bone marrow MSCs from passage 3 immunolabeled for CD44 (first panel, medium gray, shown in red in the original) and nestin (second panel, lighter gray, shown in green in the original). The third panel is a DAPI nuclear stain (blue in original). The merged image in the right-most panel shows co-staining of a population of cells with both markers (lightest gray, yellow/orange in the original)
[0030] FIG. 1C is a row of four photomicrographs of bone marrow MSCs from passage 3 stained for co-expression of Sca-1 (first panel, red in the original) and nestin (second panel, green in the original). Merged image in the right-most panel shows co-staining.
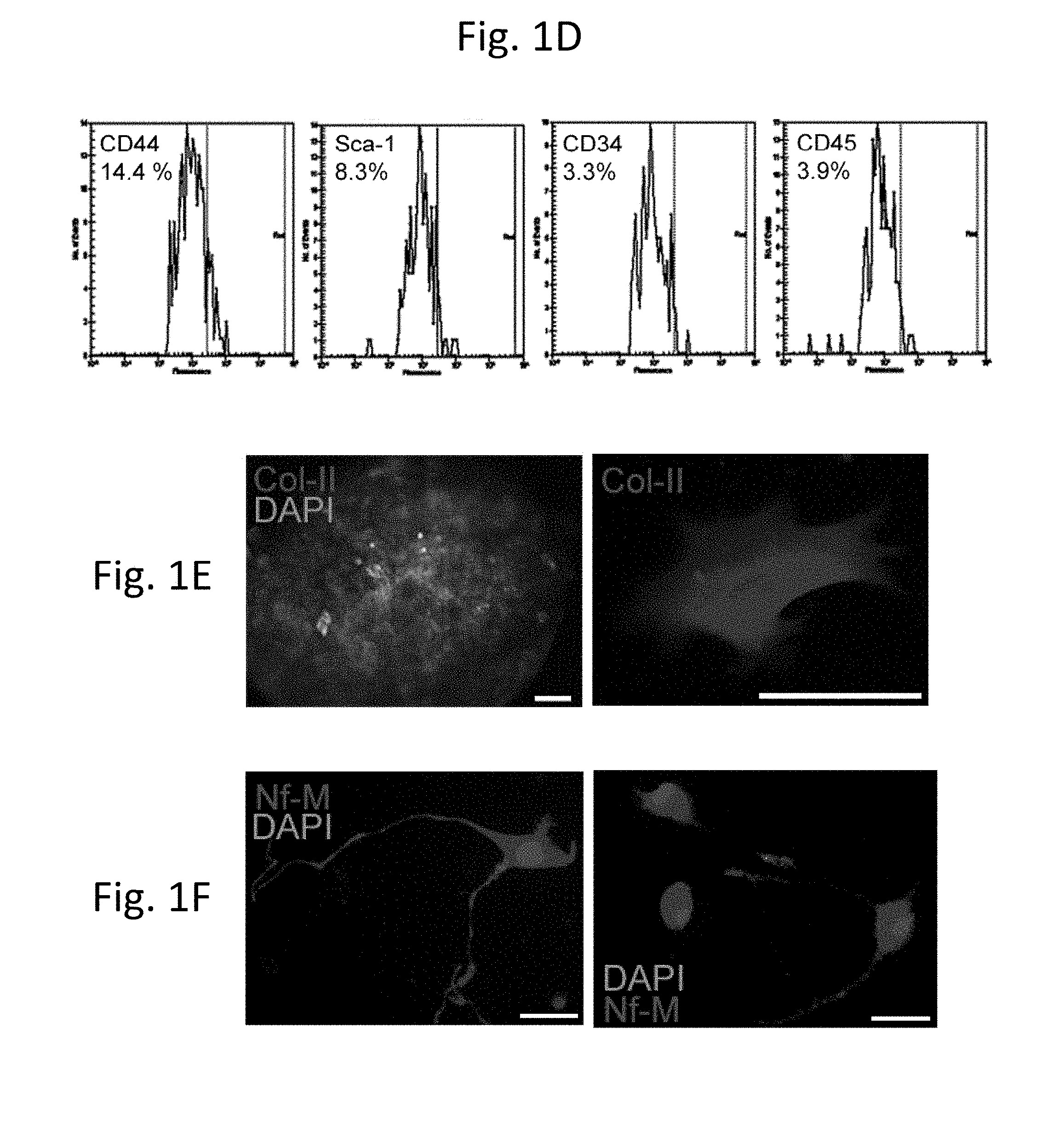
[0031] FIG. 1D is a row of four plots showing the results of analysis of bone marrow MSCs by chip flow cytometry indicating the ratio of immunopositive cells for each of the listed antibodies (CD44, first panel; Sca-1, second panel; CD34, third panel; and CD45, last panel); axes are "Fluorescence" and "No. of events."
[0032] FIG. 1E is a pair of photomicrographs showing the potential for lineage differentiation, as demonstrated by formation of chondrocytes and extracellular matrix after treatment of bone marrow MSCs with TGF-.beta.. Cells that grew out from a micro-aggregate (left) were stained for type II collagen (right).
[0033] FIG. 1F is a pair of photomicrographs showing the differentiation of bone marrow MSCs to neurons by differentiation in serum-free medium containing neuronal growth supplements and bFGF. Staining for neurofilament (NF-M) is shown in these cells.
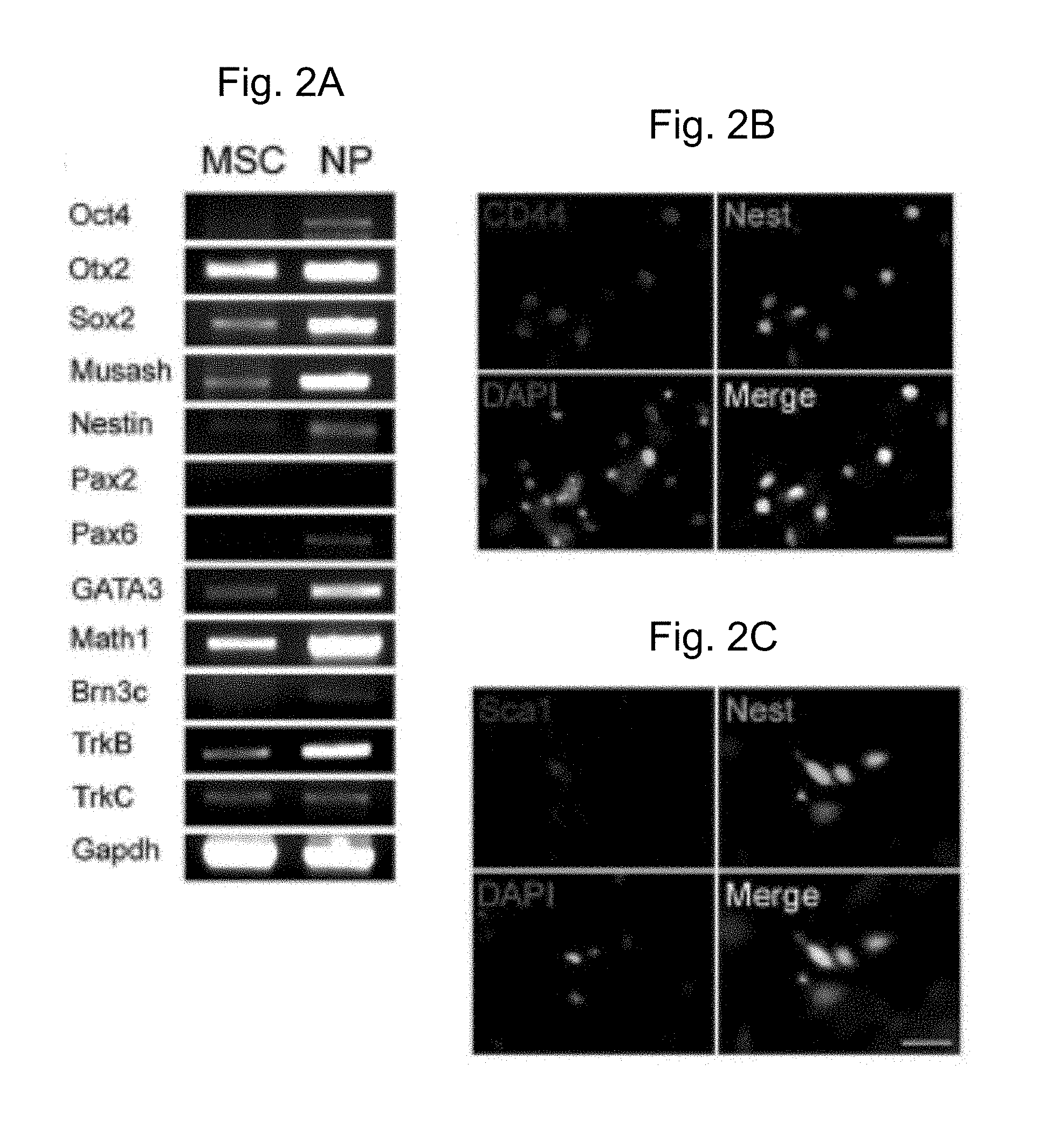
[0034] FIG. 2A is a gel showing the results of genetic analysis for neural progenitor markers by RT-PCR of MSCs treated with IGF-1, EGF and bFGF for 14 days followed by analysis. MSC (bone marrow MSCs), NP (neural progenitors at 2 wks after induction of progenitor formation). The genes analyzed are shown to the left of the gel.
[0035] FIGS. 2B-C are two sets of four photomicrographs showing that the neural progenitor marker, nestin, visualized by immunohistochemistry using a secondary antibody labeled with FITC (top right panel in 2B and 2C, shown in green in the original), was co-expressed with CD44 (2B, top left panel, shown in red in the original) and with Sca-1 (2C, top left panel, shown in red in the original). DAPI is shown in blue in the original (lower left panel in each figure). Scale bars are 50 .mu.m. Merged images in the lower right panel of each figure show coexpression of nestin and CD44 (2B) or Sca1 (2C) (all of the cells appeared green in the original, indicating coexpression).
[0036] FIG. 3A is a gel showing the results of genetic analysis by RT-PCR of precursor cells incubated in NT3, FGF and BDNF (which support neuronal and sensory cell progenitors in the inner ear). The gene profiles included expression of Oct4, nestin, Otx2, and Musashi, as well as proneural transcription factors, GATA3, NeuroD, Ngn1, Atoh1, Brn3c, and Zic2. These cells did not express hair cells genes, myosin VIIa and espin.
[0037] FIG. 3B is a gel showing the results of genetic analysis by RT-PCR of the cells obtained after induction with NT3, FGF, and BDNF. Genes characteristic of supporting cells (claudin14, connexin 26, p75.sup.Trk, Notch 1, and S100A) were also observed. These progenitor cells thus had expression profiles characteristic of neuronal or sensory progenitors. Genes analyzed are shown to the left of the gels.
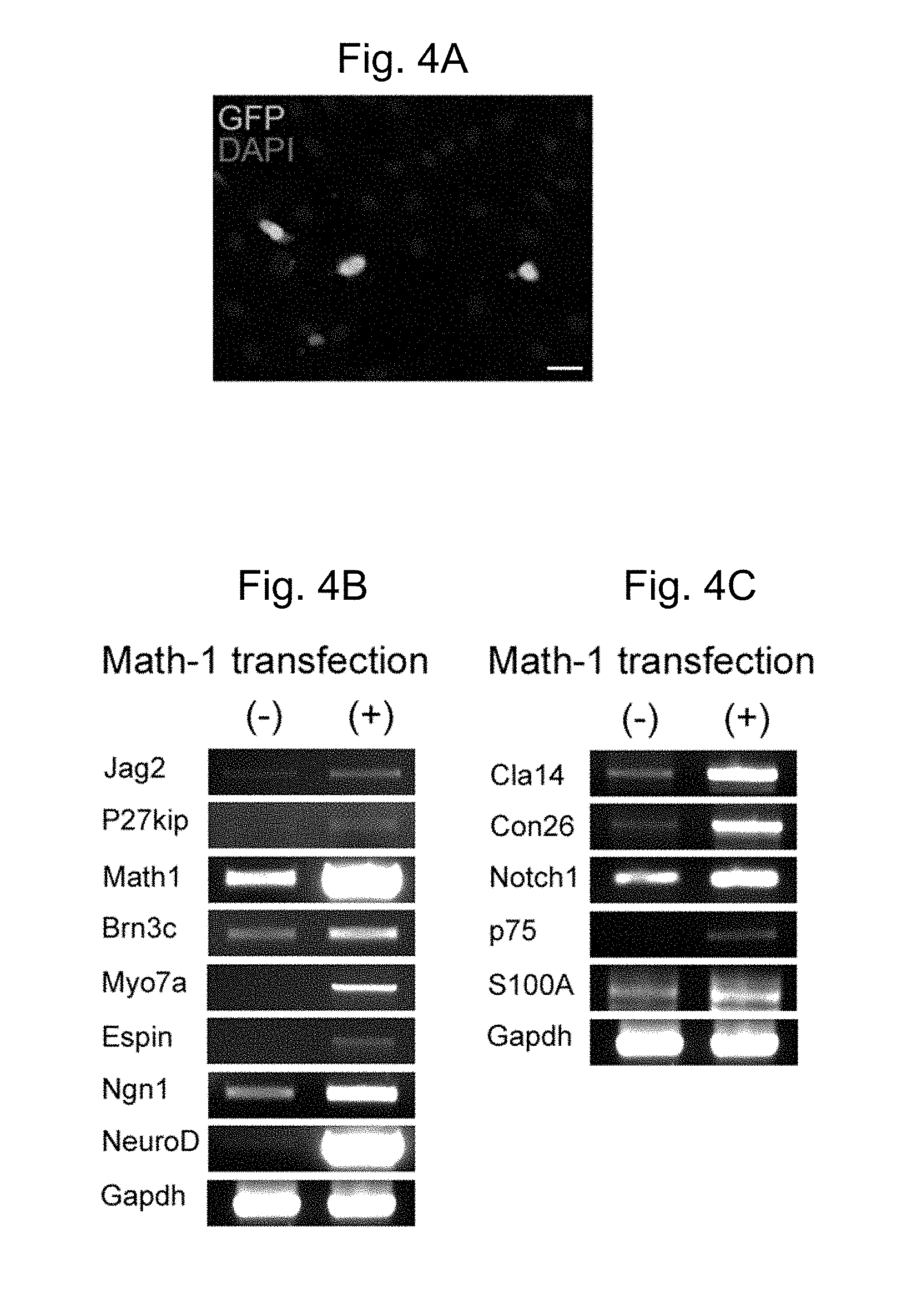
[0038] FIG. 4A is a photomicrograph showing exogenous expression of Atoh1 in bone marrow MSCs; expression was observed in cells and nuclei (green in the original) due to the expression of GFP from the vector.
[0039] FIG. 4B is a gel showing the results of gene expression in cells transfected with Atoh1 followed by treatment of the cells with NT3, FGF and BDNF. The results indicate that this protocol gave rise to progenitor cells that subsequently matured into cells expressing hair cell genes, including espin, myosin VIIa, jagged 2, and Brn3c, and p27Kip, in addition to the proneural genes, Ngn1 and NeuroD.
[0040] FIG. 4C is a gel showing the results of further genetic analysis of the cells under the differentiating conditions described in 4B; the results showed that the cells also expressed S100A, p75.sup.Trk, claudin 14, connexin 26, and Notch1, consistent with some cells having a supporting cell phenotype.
[0041] FIG. 4D is a photomicrograph of an MSC cell line selected in Zeocin; the cells had a high percentage of GFP expression when cultured in serum (green in original).
[0042] FIG. 4E is a row of 4 photomicrographs of cells stained for Myo7a (first panel), Math1/Atoh1 (second panel), or DAPI (third panel); the last panel is a merged image. After differentiation, the number of hair cell-like cells per DAPI nucleus rose and these cells stained for myosin VIIa (shown in red in the first panel) and Atoh1 (shown in green in the second panel; arrows in the second and last panels).
[0043] FIG. 4F is two rows of 4 photomicrographs of an Atoh1 expressing cell line differentiated to cells with nuclei that were immunopositive for Brn3c (second column, green in original; indicated by arrowheads) and cytoplasm positive for myosin VIIa (first column, red in original; indicated by arrows). Nuclei were stained with DAPI (third column, blue in original).
[0044] FIG. 4G is a row of three photomicrographs showing that the differentiated cells were positive for F-actin which protruded from the apex of the cell in the shape of a stereocilia bundle (arrow).
[0045] FIG. 4H is a row of three photomicrographs showing that F-actin staining was arranged in a characteristic V pattern on the apical surface.
[0046] FIG. 5A is a gel showing the results of genetic analysis of bone marrow MSC derived progenitors were co-cultured for 21 days with chick otocyst cells that had been treated with mitomycin C (Mito C); the results showed that expression of jagged 2, p27Kip, Atoh1, Brn3c, myosin VIIa and espin was increased, whereas the expression of these genes in chick cells was undetectable. Chick otocyst cells that had been fixed by incubation with paraformaldehyde were less effective (PFA) than the unfixed cells but did cause differentiation of the progenitors. Conditioned medium from the chick cells (Cnd Med) had no effect (levels of expression of these markers similar to previously shown data for differentiating conditions).
[0047] FIG. 5B is a set of three photomicrographs showing that expression of Atoh1 (Math-1, middle panel, green in original) and myosin VIIa (top panel, red in original) in cells from a Atoh1-GFP mouse showed green fluorescence corresponding to the induction of this marker in the nucleus and had expression of myosin VIIa in the cytoplasm.
[0048] FIG. 6A is a set of four photomicrographs showing an increase in fluorescence (green in original) indicating the conversion of bone marrow cells to cells expressing Atoh1. The cells stained for Atoh1 (Math1, bottom left, green in original), myosin VIIa (top left, red in original) and DAPI (top right, blue in original). A merged image is shown in on the bottom right panel.
[0049] FIG. 6B is a photomicrograph showing that Atoh1-expressing cells were found incorporated into the tissue of the chick otic epithelium. The hair cells of the chick were stained with the chick-specific marker, HCA (white in original) and myosin VIIa (red in original), whereas the Atoh-1 expressing mouse cells were green due to expression of GFP (arrows).
[0050] FIG. 6C is a set of four photomicrographs showing a lack of cell fusion, demonstrated by the presence of HCA (arrowhead, lower panels) in cells that did not have green fluorescence and of Atoh1-GFP (arrow, right column) exclusively in cells that did not stain for HCA, a marker for chicken cells. No cells with both GFP and HCA were observed in these experiments. Scale bars are 100 .mu.m.
[0051] FIG. 7 is a gel showing the results of genetic analysis of cells after inhibition of Notch signaling with an inhibitor of .gamma.-secretase increases expression of hair cell markers. Gene expression in MSCs treated with a .gamma.-secretase inhibitor showed that loss of Notch signaling increased Atoh1 expression. The timing of inhibition was critical: .gamma.-secretase inhibitor added at dl of differentiation in vitro for a total of 10 days led to an increase in hair cell markers, myosin VIIa and espin, whereas inhibitor added at d3 did not induce hair cell markers.
DETAILED DESCRIPTION
[0052] Although stem cells are present in the inner ear (Li et al., Trends Mol Med 10, 309-315 (2004); Li et al., Nat Med 9, 1293-1299 (2003); Rask-Andersen et al., Hear Res 203, 180-191 (2005)), hair cells do not regenerate after damage, and, therefore, a source of cells that could potentially be used for cell transplantation in a therapeutic replacement of these sensory cells has important implications for treatment of sensorineural hearing loss. Bone marrow has been harvested and used extensively in clinical applications and is a highly desirable source, because cells from a patient's bone marrow could potentially be transplanted without the problem of immune rejection. The present methods include a treatment regimen for hearing loss including transplantation of hair cells obtained by methods described herein.
[0053] By a combination of growth factor stimulation and expression of the transcription factor, Atoh1, that is required for hair cell formation in the inner ear, the present inventors demonstrate herein that stem cells, e.g., mesenchymal stem cells derived from bone marrow, can be induced to differentiate into hair cells. In addition, the neurosensory progenitors obtained from bone marrow can be converted to sensory cells by co-culture with cells of the developing sensory epithelium, even in the absence of Atoh1 expression.
[0054] Stem cells in bone marrow are known to be the precursors for all lymphoid and erythroid cells, but mesenchymal stem cells in bone marrow also act as precursors to bone, cartilage, and fat cells (Colter et al., Proc Natl Acad Sci USA 97, 3213-3218 (2000); Pittenger et al., Science 284, 143-147 (1999)). In addition to mesenchymal tissues, these stem cells have been shown to give rise to cells of other lineages including pancreatic cells (Hess et al., Nat Biotechnol 21, 763-770 (2003)), muscle cells (Doyonnas et al., Proc Natl Acad Sci USA 101, 13507-13512 (2004)) and neurons (Dezawa et al., J Clin Invest 113, 1701-1710 (2004); Hermann et al., J Cell Sci 117, 4411-4422 (2004); Jiang et al., Proc Natl Acad Sci USA 100 Suppl 1, 11854-11860 (2003)). The evidence provided herein demonstrates an extended range of cell fates available for these bone marrow-derived cells that includes cells of the neurosensory lineage, even including differentiation to inner ear hair cells.
Methods for Generating Cells of the Inner Ear
[0055] Methods of generating cells of the inner ear are provided, including progenitor cells and differentiated inner ear cells including hair cells and supporting cells. Stem cells are unspecialized cells capable of extensive proliferation. Stem cells are pluripotent and are believed to have the capacity to differentiate into most cell types in the body (Pedersen, Scientif. Am. 280:68 (1999)), including neural cells, muscle cells, blood cells, epithelial cells, skin cells, and cells of the inner ear (e.g., hair cells and cells of the spiral ganglion). Stem cells are capable of ongoing proliferation in vitro without differentiating. As they divide, they retain a normal karyotype, and they retain the capacity to differentiate to produce adult cell types.
[0056] Hematopoietic stem cells resident in bone marrow are the source of blood cells, but in addition to these hematopoietic stem cells, the bone marrow contains mesenchymal stem cells (MSCs) that can differentiate into cell types of all three embryonic germ layers (Colter et al., Proc Natl Acad Sci USA 97, 3213-3218 (2000); Doyonnas et al., Proc Natl Acad Sci USA 101, 13507-13512 (2004); Herzog et al., Blood 102, 3483-3493 (2003); Hess et al., Nat Biotechnol 21, 763-770 (2003); Jiang et al., Nature 418, 41-49 (2002); Pittenger et al., Science 284, 143-147 (1999)). This has been demonstrated in vivo in studies that track transplanted bone marrow cells to specific tissues where they differentiate into the resident tissue type (Mezey et al., Proc Natl Acad Sci USA 100, 1364-1369 (2003); Weimann et al., Proc Natl Acad Sci USA 100, 2088-2093 (2003)).
[0057] Many of these cells have been used for transplantation and are a preferred source of new cells for therapies because the transplanted cells are immunologically matched when harvested from a patient to be treated and because they have been extensively used in clinical applications so that their safety is known.
[0058] Stem cells can differentiate to varying degrees. For example, stem cells can form cell aggregates called embryoid bodies in hanging drop cultures. The embryoid bodies contain neural progenitor cells that can be selected by their expression of an early marker gene such as Sox1 and the nestin gene, which encodes an intermediate filament protein (Lee et al., Nat. Biotech. 18:675-9, 2000).
[0059] Neurogenic Stem Cells
[0060] Inner ear cells or inner ear cell progenitors can be generated from mammalian stem cells. As described herein, stem cells suitable for use in the present methods can be any stem cell that has neurogenic potential, i.e., any stem cell that has the potential to differentiate into a neural cell, e.g., neurons, glia, astrocytes, retinal photoreceptors, oligodendrocytes, olfactory cells, hair cells, supporting cells, and the like. Neurogenic stem cells, including human adult stem cells such as bone marrow mesenchymal stem cells, can be induced to differentiate into inner ear progenitor cells that are capable of giving rise to mature inner ear cells including hair cells and supporting cells. Neurogenic stem cells useful in the methods described herein can be identified by the expression of certain neurogenic stem cell markers, such as nestin, sox1, sox2, and musashi. Alternatively or in addition, these cells express high levels of helix-loop-helix transcription factors NeuroD, Atoh1, and neurogenin1.
[0061] Examples of neurogenic stem cells include embryonic stem cells or stem cells derived from mature (e.g., adult) tissue, such as the ear (e.g., inner ear), central nervous system, blood, skin, eye or bone marrow. In some embodiments, the stem cells are mesenchymal stem cells. Any of the methods described herein for culturing stem cells and inducing differentiation into inner ear cells (e.g., hair cells or supporting cells) can be used.
[0062] Stem cells useful for generating cells of the inner ear can be derived from a mammal, such as a human, mouse, rat, pig, sheep, goat, or non-human primate. For example, stem cells have been identified and isolated from the mouse utricular macula (Li et al., Nature Medicine 9:1293-1299, 2003).
[0063] Generation of Neural Progenitor Cells
[0064] There are a number of induction protocols known in the art for inducing differentiation of stem cells with neurogenic potential into neural progenitor cells, including growth factor treatment (e.g., treatment with EGF, FGF, and IGF, as described herein) and neurotrophin treatment (e.g., treatment with NT3 and BDNF, as described herein). Other differentiation protocols are known in the art; see, e.g., Corrales et al., J. Neurobiol. 66(13):1489-500 (2006); Kim et al., Nature 418, 50-6 (2002); Lee et al., Nat Biotechnol 18, 675-9 (2000); and Li et al., Nat Biotechnol 23, 215-21 (2005).
[0065] As one example of an induction protocol, the stem cells are grown in the presence of supplemental growth factors that induce differentiation into progenitor cells. These supplemental growth factors are added to the culture medium. The type and concentration of the supplemental growth factors is be adjusted to modulate the growth characteristics of the cells (e.g., to stimulate or sensitize the cells to differentiate) and to permit the survival of the differentiated cells such as neurons, glial cells, supporting cells or hair cells.
[0066] Exemplary supplementary ng/mL, about 50 ng/mL, about 40 ng/mL, about 30 ng/mL, about 20 ng/mL, about 10 ng/mL, or about 5 ng/mL).
[0067] Neural progenitor cells produced by these methods include inner ear progenitor cells, i.e., cells that can give rise to inner ear cells such as hair cells and supporting cells. Inner ear progenitor cells can be identified by the expression of marker genes such as nestin, sox2, and musashi, in addition to certain inner-ear specific marker genes Brn3C, Pax2, and Atoh1. The invention includes purified populations of inner ear progenitor cells expressing nestin, sox2, musashi, Brn3C, Pax2, and Atoh1. These inner ear progenitor cells are lineage committed, and can be induced to further differentiate into hair cells and supporting cells by a method described herein.
[0068] Progenitor cells prepared by a method described herein can optionally be frozen for future use.
[0069] Cell Culture Methods
[0070] In general, standard culture methods are used in the methods described herein. Appropriate culture medium is described in the art, such as in Li et al. (supra). For example, stem cells can be cultured in serum free DMEM/high-glucose and F12 media (mixed 1:1), and supplemented with N2 and B27 solutions and growth factors. Growth factors such as EGF, IGF-1, and bFGF have been demonstrated to augment sphere formation in culture. In vitro, stem cells often show a distinct potential for forming spheres by proliferation of single cells. Thus, the identification and isolation of spheres can aid in the process of isolating stem cells from mature tissue for use in making differentiated cells of the inner ear. The growth medium for cultured stem cells can contain one or more or any combination of growth factors. This includes leukemia inhibitory factor (LIF) which prevents the stem cells from differentiating. To induce the cells (and the cells of the spheres) to differentiate, the medium can be exchanged for medium lacking growth factors. For example, the medium can be serum-free DMEM/high glucose and F12 media (mixed 1:1) supplemented with N2 and B27 solutions. Equivalent alternative media and nutrients can also be used. Culture conditions can be optimized using methods known in the art.
[0071] growth factors are discussed in detail below, and include, but are not limited to basic fibroblast growth factor (bFGF), insulin-like growth factor (IGF), and epidermal growth factor (EGF). Alternatively, the supplemental growth factors can include the neurotrophic factors neurotrophin-3 (NT3) and brain derived neurotrophic factor (BDNF). Concentrations of growth factors can range from about 100 ng/mL to about 0.5 ng/mL (e.g., from about 80 ng/mL to about 3 ng/mL, such as about 60
[0072] Differentiation by Expression of Atoh1
[0073] As described herein, expression of Atoh1 in stem-cell derived progenitor cells was sufficient to drive them into adopting hair cell markers. Studies of Atoh1 expression in the ear have indicated that this helix-loop-helix transcription factor occupies a key place in the hierarchy of inner ear transcription factors for differentiation of hair cells.
[0074] Atoh1 nucleic acids and polypeptides are known in the art, and described in, for example, U.S. Pat. Nos. 6,838,444 and 7,053,200, and P.G PUB. Nos. 2004/0237127 and 2004/0231009, all to Zoghbi et al., all incorporated by reference in their entirety. In some embodiments, the Atoh1 is, or is at least 80%, 85%, 90%, 93%, or 95% identical to, human atonal homolog 1 (ATOH1); ATH1; and HATH1 (for additional information see Ben-Arie et al., Molec. Genet. 5: 1207-1216 (1996); Bermingham et al., Science 284: 1837-1841 (1999); OMIM*601461; UniGene Hs.532680; GenBank Accession Nos. NM_005172.1 (nucleic acid) and NP 005163.1 (polypeptide)). Other species can also be used, e.g., Mouse Atoh1 (also known as Math1, GenBankAcc. No. NM_007500.2), chicken Atoh1 (also known as Cath1; GenBankAcc. No. AF467292.1).
[0075] The human Atoh1 mRNA (CDS=-1065) and polypeptide sequences are as follows:
TABLE-US-00001 (SEQ ID NO: 1) 1 atgtcccgcc tgctgcatgc agaagagtgg gctgaagtga aggagttggg agaccaccat 61 cgccagcccc agccgcatca tctcccgcaa ccgccgccgc cgccgcagcc acctgcaact 121 ttgcaggcga gagagcatcc cgtctacccg cctgagctgt ccctcctgga cagcaccgac 181 ccacgcgcct ggctggctcc cactttgcag ggcatctgca cggcacgcgc cgcccagtat 241 ttgctacatt ccccggagct gggtgcctca gaggccgctg cgccccggga cgaggtggac 301 ggccgggggg agctggtaag gaggagcagc ggcggtgcca gcagcagcaa gagccccggg 361 ccggtgaaag tgcgggaaca gctgtgcaag ctgaaaggcg gggtggtggt agacgagctg 421 ggctgcagcc gccaacgggc cccttccagc aaacaggtga atggggtgca gaagcagaga 481 cggctagcag ccaacgccag ggagcggcgc aggatgcatg ggctgaacca cgccttcgac 541 cagctgcgca atgttatccc gtcgttcaac aacgacaaga agctgtccaa atatgagacc 601 ctgcagatgg cccaaatcta catcaacgcc ttgtccgagc tgctacaaac gcccagcgga 661 ggggaacagc caccgccgcc tccagcctcc tgcaaaagcg accaccacca ccttcgcacc 721 gcggcctcct atgaaggggg cgcgggcaac gcgaccgcag ctggggctca gcaggcttcc 781 ggagggagcc agcggccgac cccgcccggg agttgccgga ctcgcttctc agccccagct 841 tctgcgggag ggtactcggt gcagctggac gctctgcact tctcgacttt cgaggacagc 901 gccctgacag cgatgatggc gcaaaagaat ttgtctcctt ctctccccgg gagcatcttg 961 cagccagtgc aggaggaaaa cagcaaaact tcgcctcggt cccacagaag cgacggggaa 1021 ttttcccccc attcccatta cagtgactcg gatgaggcaa gttag (SEQ ID NO: 2) MSRLLHAEEWAEVKELGDHHRQPQPHHLPQPPPPPQPPATLQAREHPVYP PELSLLDSTDPRAWLAPTLQGICTARAAQYLLHSPELGASEAAAPRDEVD GRGELVRRSSGGASSSKSPGPVKVREQLCKLKGGVVVDELGCSRQRAPSS KQVNGVQKQRRLAANARERRRMHGLNHAFDQLRNVIPSFNNDKKLSKYET LQMAQIYINALSELLQTPSGGEQPPPPPASCKSDHHHLRTAASYEGGAGN ATAAGAQQASGGSQRPTPPGSCRTRFSAPASAGGYSVQLDALHFSTFEDS ALTAMMAQKNLSPSLPGSILQPVQEENSKTSPRSHRSDGEFSPHSHYSDS DEAS
[0076] The mouse Atoh1 mRNA (CDS=196-1251) and polypeptide sequences are as follows:
TABLE-US-00002 (SEQ ID NO: 3) 1 tcgacccacg cgtccgccca cgcgtccgga tctccgagtg agagggggag ggtcagagga 61 ggaaggaaaa aaaaatcaga ccttgcagaa gagactagga aggtttttgt tgttgttgtt 121 cggggcttat ccccttcgtt gaactgggtt gccagcacct cctctaacac ggcacctccg 181 agccattgca gtgcgatgtc ccgcctgctg catgcagaag agtgggctga ggtaaaagag 241 ttgggggacc accatcgcca tccccagccg caccacgtcc cgccgctgac gccacagcca 301 cctgctaccc tgcaggcgag agaccttccc gtctacccgg cagaactgtc cctcctggat 361 agcaccgacc cacgcgcctg gctgactccc actttgcagg gcctctgcac ggcacgcgcc 421 gcccagtatc tgctgcattc tcccgagctg ggtgcctccg aggccgcggc gccccgggac 481 gaggctgaca gccagggtga gctggtaagg agaagcggct gtggcggcct cagcaagagc 541 cccgggcccg tcaaagtacg ggaacagctg tgcaagctga agggtggggt tgtagtggac 601 gagcttggct gcagccgcca gcgagcccct tccagcaaac aggtgaatgg ggtacagaag 661 caaaggaggc tggcagcaaa cgcaagggaa cggcgcagga tgcacgggct gaaccacgcc 721 ttcgaccagc tgcgcaacgt tatcccgtcc ttcaacaacg acaagaagct gtccaaatat 781 gagaccctac agatggccca gatctacatc aacgctctgt cggagttgct gcagactccc 841 aatgtcggag agcaaccgcc gccgcccaca gcttcctgca aaaatgacca ccatcacctt 901 cgcaccgcct cctcctatga aggaggtgcg ggcgcctctg cggtagctgg ggctcagcca 961 gccccgggag ggggcccgag acctaccccg cccgggcctt gccggactcg cttctcaggc 1021 ccagcttcct ctgggggtta ctcggtgcag ctggacgctt tgcacttccc agccttcgag 1081 gacagggccc taacagcgat gatggcacag aaggacctgt cgccttcgct gcccgggggc 1141 atcctgcagc ctgtacagga ggacaacagc aaaacatctc ccagatccca cagaagtgac 1201 ggagagtttt ccccccactc tcattacagt gactctgatg aggccagtta ggaaggcaac 1261 agctccctga aaactgagac aaccaaatgc ccttcctagc gcgcgggaag ccccgtgaca 1321 aatatccctg caccctttaa tttttggtct gtggtgatcg ttgttagcaa cgacttgact 1381 tcggacggct gcagctcttc caatcccctt cctcctacct tctccttcct ctgtatgtag 1441 atactgtatc attatatgta cctttacgtg gcatcgtttc atggtccatg ctgccaatat 1501 gctgctaaaa tgtcgtatct ctgcctctgg tctgggtttc acttatttta taccttggga 1561 gttcatcctt gcgtgttgcg ctcactcaca aataagggag ttagtcaatg aagttgtttc 1621 cccaactgct tgagacccgc attgggtact ttactgaaca cggactattg tgttgttaaa 1681 atgcaggggc agataagagt atctgtagag cttagacacc aagtgtgtcc agcagtgtgt 1741 ctagcggacc cagaatacac gcacttcatc actggccgct gcgccgcctt gaagaaactc 1801 aactgccaat gcagagcaac ttttgatttt aaaaacagcc actcataatc attaaactct 1861 ttgcaaatgt ttgtttttgc aaatgaaaat taaaaaaaaa catgtagtgt caaaggcatt 1921 tggtcaattt tattttgctt tgttaacatt agaaaagtta tttattattg cgtatttgga 1981 cccatttcta cttaattgcc ttttttttac attttctact cgagatcgtt ttattttgat 2041 ttagcaaatc cagttgccat tgctttatgt atgtatgctc ttttacaaat gataaaataa 2101 actcggaaaa aaaaaaaaaa aaaaaaaaaa aaaaaaaaaa aaaa (SEQ ID NO: 4) MSRLLHAEEWAEVKELGDHHRHPQPHHVPPLTPQPPATLQARDLLVRRSG CGGLSKSPGPVKVREQLCKLKGGVVVDELGCSRQRAPSSKQVNGVQKQRR LAANARERRRMHGLNHAFDQLRNVIPSENNDKKLSKYETLQMAQIYINAL SELLQTPNVGASSGGYSVQLDALHEPAFEDRALTAMMAQKDLSPSLPGGI LQPVQEDNSKTSPRSHRSDGEFSPHSHYSDSDEAS
[0077] The chicken Cath1 mRNA (CDS=1-717) and polypeptide sequences are as follows:
TABLE-US-00003 (SEQ ID NO: 5) 1 atggccccag gaggtagcga gtgttgttgc agtgatgccg cgcacatcac ttggaggcag 61 tgggagtaca cgcacgagaa ccaactgtgc gtggcaggaa ctgtcagcag gatgaggccc 121 aggacgtggg tctgcaccgg atctttgtgg gaccaggaag cgggaattac tttgatgggc 181 ccccaaatac ccaaagtgga tgaggcagga gtgatgaccc acccggcaag gtcgctttgc 241 agcactgggg cacatccgtg tcccggggtg gtcgtgctgc ccacgggtgg gatagggcag 301 ccttcaaaga agctctccaa gtacgagacg ctgcagatgg cgcaaatcta catcagcgcc 361 ctcgccgagc ttctgcacgg gccgcccgcg ccccccgagc cgcccgccaa ggccgagctc 421 cgcggggccc ccttcgagcc tcccccgccg ccccctcctc cgccgccccg cgcctcgccc 481 cccgcgcccg ccaggactcg cttccccccg gcggcggccg cgggcggttt cgcggcgctt 541 ctcgagccgc tgcgcttccc ttctttcccg gcgcagaaag cgccttctcc cgcgctgctc 601 ctggggccgc ccgcgccgca gcagcccgag aggagcaaag cgtcgccgcg ctctcaccgc 661 agcgacgggg agttctcgcc gcgctcccac tacagtgact cggacgaggc cagctag (SEQ ID NO: 6) MAPGGSECCCSDAAHITWRQWEYTHENQLCVAGTVSRMRPRTWVCTGSLWDQEAGI TLMGPQIPKVDEAGVMTHPARSLCSTGAHPCPGVVVLPTGGIGQPSKKLSKYETLQ MAQIYISALAELLHGPPAPPEPPAKAELRGAPFEPPPPPPPPPPRASPPAPARTRF PPAAAAGGFAALLEPLRFPSFPAQKAPSPALLLGPPAPQQPERSKASPRSHRSDGE FSPRSHYSDSDEAS
[0078] To determine the percent identity of two amino acid sequences, or of two nucleic acid sequences, the sequences are aligned for optimal comparison purposes (e.g., gaps can be introduced in one or both of a first and a second amino acid or nucleic acid sequence for optimal alignment and non-homologous sequences can be disregarded for comparison purposes). The length of a reference sequence aligned for comparison purposes is at least 80% of the length of the reference sequence, and in some embodiments is at least 90% or 100%. The amino acid residues or nucleotides at corresponding amino acid positions or nucleotide positions are then compared. When a position in the first sequence is occupied by the same amino acid residue or nucleotide as the corresponding position in the second sequence, then the molecules are identical at that position (as used herein amino acid or nucleic acid "identity" is equivalent to amino acid or nucleic acid "homology"). The percent identity between the two sequences is a function of the number of identical positions shared by the sequences, taking into account the number of gaps, and the length of each gap, which need to be introduced for optimal alignment of the two sequences.
[0079] For purposes of the present invention, the comparison of sequences and determination of percent identity between two sequences can be accomplished using a Blossum 62 scoring matrix with a gap penalty of 12, a gap extend penalty of 4, and a frameshift gap penalty of 5.
[0080] In some embodiments, the methods include expressing in the cells a Atoh1 polypeptide encoded by a nucleic acid that hybridizes to the human Atoh1 mRNA under stringent conditions. As used herein, the term "stringent conditions" describes conditions for hybridization and washing. Stringent conditions as used herein are 0.5M sodium phosphate, 7% SDS at 65.degree. C., followed by one or more washes at 0.2.times.SSC, 1% SDS at 65.degree. C. See, e.g., Current Protocols in Molecular Biology, John Wiley & Sons, N.Y. (2006).
[0081] In some embodiments, the methods include expressing exogenous Atoh1 in a stem cell. This can be achieved, for example, by introducing an expression vector in the cell. As used herein, the term "vector" refers to a nucleic acid molecule capable of transporting another nucleic acid to which it has been linked and can include a plasmid, cosmid or viral vector. The vector can be capable of autonomous replication or it can integrate into a host DNA. Viral vectors include, e.g., replication defective retroviruses, adenoviruses and adeno-associated viruses.
[0082] A vector can include a Atoh1 nucleic acid in a form suitable for expression of the nucleic acid in a host cell. Generally, the expression vector includes one or more regulatory sequences operatively linked to the nucleic acid sequence to be expressed. The term "regulatory sequence" includes promoters, enhancers and other expression control elements (e.g., polyadenylation signals). Regulatory sequences include those which direct constitutive expression of a nucleotide sequence, as well as tissue-specific regulatory and/or inducible sequences. The design of the expression vector can depend on such factors as the choice of the host cell to be transformed, the level of expression of protein desired, and the like. The expression vectors can be introduced into host cells using methods known in the art, including calcium phosphate or calcium chloride co-precipitation, DEAE-dextran-mediated transfection, lipofection, or electroporation. See, e.g., Current Protocols in Molecular Biology, John Wiley & Sons, N.Y. (2006).
[0083] In the present methods, the Atoh1 polypeptide expressed in the stem cells will have the ability to induce differentiation of mesenchymal stem cells to hair cells and/or supporting cells, as described herein.
[0084] Differentiation by Culturing with Chick Otocysts
[0085] Also as described herein, the stem cell-derived progenitor cells also responded to physical contact with developing otocyst cells from the chicken embryo by differentiating into sensory epithelial cells, without the requirement for exogenous Atoh1. This was evidenced by nGFP expression from a Atoh1 enhancer-GFP reporter construct and co-expression of myosin VIIa after co-culture and differentiation, as described herein. Neurons that express markers of sensory cells have been induced from bone marrow MSCs in previous work by incubation with otocyst and hindbrain-conditioned medium (Kondo et al., Proc Natl Acad Sci USA 102, 4789-4794 (2005)) from embryonic mice.
[0086] Thus, the methods described herein can include contacting progenitor cells with otocyst cells, e.g., cells isolated from E3 embryonic chicks, as described herein.
[0087] In some embodiments, the methods include culturing the progenitor cells with the otocyst cells in a ratio of about 50,000 cells per confluent layer of otocyst cells, or by injection of 100,000 cells into an intact otocyst (see Examples, below). Alternatively, the stem cells can be cultured in the presence of chick otocyst-conditioned media, which can be produced using methods known in the art, e.g., using media that has been in contact with a culture of chick otocysts for at about four days.
[0088] Differentiation by Inhibition of Notch Signalling
[0089] Notch is a plasma membrane receptor, and the Notch pathway consists of Notch and its ligands, as well as intracellular proteins that transmit the Notch signal to the nucleus. Included in the Notch pathway are the transcription factors that bear the effector function of the pathway.
[0090] Notch signaling plays a role in lateral inhibition, in which one cell is singled out from a cell cluster for a given fate (e.g., differentiation into a hair cell, for example). Differentiation is inhibited in those cells not selected to differentiate, resulting in the prevention of a specified fate commitment on the part of most of the cells of a cluster. Lateral inhibition occurs repeatedly during development. Central to this process is binding to the Notch receptor of one of several ligands, including Delta, Scabrous and Serrate. Ligand binding to Notch ligand triggers a chain of intracellular events resulting in lateral inhibition. A review of the Notch pathway can be found at Artavanis-Tsakonas et al., Science 268: 225-232 (1995). As described herein, inhibition of Notch in the inner ear progenitor cells described herein results in differentiation of the cells into hair cells and supporting cells.
[0091] Thus, in some embodiments of the methods described herein, progenitor cells are grown in the presence of a Notch signalling pathway inhibitor. Exemplary Notch pathway inhibitors include .gamma.-secretase inhibitors, of which a number are known in the art (e.g., aryl sulfonamides (AS), dibenzazepines (DBZ), benzodiazepines (BZ), N--[N-(3,5-difluorophenacetyl)-L-alanyl]-(S)-phenylglycine t-butyl ester (DAPT), L-685,458 (Sigma-Aldrich), and MK0752 (Merck). A useful concentration will depend on the inhibitor chosen.
[0092] Other Notch inhibitors include inhibitory nucleic acids (e.g., small interfering RNAs, antisense oligonucleotides, and morpholino oligos; methods for designing, making, and using them are known in the art, e.g., gene walk methods for selecting and optimizing inhibitory sequences, see, e.g., Engelke, RNA Interference (RNAi): The Nuts & Bolts of siRNA Technology, (DNA Press, 2004); Mol, Antisense Nucleic Acids and Proteins, (CRC, 1994); Sioud, Ribozymes and Sirna Protocols (Methods in Molecular Biology), (Humana Press; 2nd edition 2004); and Philips, Antisense Therapeutics (Methods in Molecular Medicine), (Humana Press 2004)) targeting Notch (see, e.g., Presente et al., Proc. Nat. Acad. Sci. 101(6):1764-1768 (2004); Ivanov et al., Proc. Nat. Acad. Sci. 101(46):16216-16221 (2004)) or its ligands, i.e., Delta or Jagged (see, e.g., Patzel et al., Nature Biotechnology 23, 1440-1444 (2005); Purow et al., Cancer Research 65:2353-2363 (2005); or Stallwood et al., J. Immunol. 177:885-895 (2006)). Alternatively, the cells can be modified to express m-Numb (GenBank Acc. No. NP_001005743.1) or disheveled (Dvl; the human homologs are at GenBank Acc. No. NM_004421.2 (variant 1); NM_004422.2 (variant 2); and NM_004423.3 (variant 3), both endogenous inhibitors of Notch signalling.
[0093] Assaying Differentiation
[0094] A variety of methods can be utilized to determine that a stem cell has differentiated into a progenitor cell, or into a cell of the inner ear, e.g., a hair cell or supporting cell. For example, the cell can be examined for the expression of a cell marker gene. Hair cell marker genes include myosin VIIa (myoVIIa), Atoh1, .alpha.9 acetylcholine receptor, espin, parvalbumin 3, and Brn3c. Supporting cell markers include claudin14, connexin 26, p75Trk, Notch 1, and S100A. Pluripotent stem cells generally do not express these genes. A stem cell that propagates and produces a cell expressing one or more of these genes, has produced a hair cell, i.e., the stem cell has differentiated at least partially into a hair cell. A stem cell that has differentiated into an inner ear progenitor cell (a precursor of hair cells) expresses early ear marker genes such as nestin, sox2, musashi, Brn3C, Pax2, and Atoh1. A progenitor cell can express one or more of these genes. The progenitor cells can be propagated in serum-free medium in the presence of growth factors. Removal of growth factors and expression of Atoh1, or co-culture with chick otocysts, will induce the cells to differentiate further, such as into hair cells and supporting cells.
[0095] Identification of a hair cell or hair cell progenitor (e.g., a hair cell, supporting cell, or progenitor cell that differentiated from a stem cell) can be facilitated by the detection of expression of marker genes as described herein. Detection of the products of gene expression can be by immunocytochemistry. Immunocytochemistry techniques involve the staining of cells or tissues using antibodies against the appropriate antigen. In this case, the appropriate antigen is the protein product of the tissue-specific gene expression. Although, in principle, a first antibody (i.e., the antibody that binds the antigen) can be labeled, it is more common (and improves the visualization) to use a second antibody directed against the first (e.g., an anti-IgG). This second antibody is conjugated either with fluorochromes, or appropriate enzymes for colorimetric reactions, or gold beads (for electron microscopy), or with the biotin-avidin system, so that the location of the primary antibody, and thus the antigen, can be recognized. The protein marker can also be detected by flow cytometry using antibodies against these antigens, or by Western blot analysis of cell extracts.
[0096] Alternatively or in addition, gene expression can be analyzed directly, e.g., using PCR methods known in the art, including quantitative PCR, e.g., quantitative RT-PCR, which can be used to detect and compare levels of expression.
Methods of Treatment
[0097] The methods described herein can be used to generate cells for therapeutic use. Treatment methods include generating cells of the inner ear (e.g., hair cells or supporting cells) from stem cells, using a method described herein, for transplantation into an ear of a human in need thereof. Transplantation of the cells into the inner ear of a subject can be useful for restoring or improving the ability of the subject to hear, or for decreasing the symptoms of vestibular dysfunction. Inner ear cells derived from stem cells according to the methods described herein need not be fully differentiated to be therapeutically useful. A partially differentiated cell that improves any symptom of a hearing disorder in a subject is useful for the therapeutic compositions and methods described herein.
[0098] A human having a disorder of the inner ear, or at risk for developing such a disorder, can be treated with inner ear cells (hair cells or supporting cells) generated from stem cells using a method described herein. In a successful engraftment, at least some transplanted hair cells, for example, will form synaptic contacts with spiral ganglion cells, and integrate into the sensory epithelium of the inner ear. To improve the ability of the cells to engraft, the stem cells can be modified prior to differentiation. For example, the cells can be engineered to overexpress one or more anti-apoptotic genes in the progenitor or differentiated cells. The Fak tyrosine kinase or Akt genes are candidate anti-apoptotic genes that can be useful for this purpose; overexpression of FAK or Akt can prevent cell death in spiral ganglion cells and encourage engraftment when transplanted into another tissue, such as an explanted organ of Corti (see for example, Mangi et al., Nat. Med. 9:1195-201 (2003)). Neural progenitor cells overexpressing .alpha..sub.v.beta..sub.3 integrin may have an enhanced ability to extend neurites into a tissue explant, as the integrin has been shown to mediate neurite extension from spiral ganglion neurons on laminin substrates (Aletsee et al., Audiol. Neurootol. 6:57-65 (2001)). In another example, ephrinB2 and ephrinB3 expression can be altered, such as by silencing with RNAi or overexpression with an exogenously expressed cDNA, to modify EphA4 signaling events. Spiral ganglion neurons have been shown to be guided by signals from EphA4 that are mediated by cell surface expression of ephrin-B2 and -B3 (Brors et al., J. Comp. Neurol. 462:90-100 (2003)). Inactivation of this guidance signal may enhance the number of neurons that reach their target in an adult inner ear. Exogenous factors such as the neurotrophins BDNF and NT3, and LIF can be added to tissue transplants to enhance the extension of neurites and their growth towards a target tissue in vivo and in ex vivo tissue cultures. Neurite extension of sensory neurons can be enhanced by the addition of neurotrophins (BDNF, NT3) and LIF (Gillespie et al., NeuroReport 12:275-279 (2001)). A Sonic hedgehog (Shh) polypeptide or polypeptide fragment (e.g., SHH-N), can also be useful as an endogenous factor to enhance neurite extension. Shh is a developmental modulator for the inner ear and a chemoattractant for axons (Charron et al., Cell 113:11 23 (2003)).
[0099] Any human experiencing or at risk for developing a hearing loss is a candidate for the treatment methods described herein. For example, the human can receive a transplant of inner ear hair cells or supporting cells generated by a method described herein. A human having or at risk for developing a hearing loss can hear less well than the average human being, or less well than a human before experiencing the hearing loss. For example, hearing can be diminished by at least 5, 10, 30, 50% or more. The human can have sensorineural hearing loss, which results from damage or malfunction of the sensory part (the cochlea) or the neural part (the auditory nerve) of the ear, or conductive hearing loss, which is caused by blockage or damage in the outer and/or middle ear, or the human can have mixed hearing loss, which is caused by a problem in both the conductive pathway (in the outer or middle ear) and in the nerve pathway (the inner ear). An example of a mixed hearing loss is a conductive loss due to a middle-ear infection combined with a sensorineural loss due to damage associated with aging.
[0100] The subject can be deaf or have a hearing loss for any reason or as a result of any type of event. For example, a human can be deaf because of a genetic or congenital defect; for example, a human can have been deaf since birth, or can be deaf or hard-of-hearing as a result of a gradual loss of hearing due to a genetic or congenital defect. In another example, a human can be deaf or hard-of-hearing as a result of a traumatic event, such as a physical trauma to a structure of the ear, or a sudden loud noise, or a prolonged exposure to loud noises. For example, prolonged exposures to concert venues, airport runways, and construction areas can cause inner ear damage and subsequent hearing loss. A human can experience chemical-induced ototoxicity, wherein ototoxins include therapeutic drugs including antineoplastic agents, salicylates, quinines, and aminoglycoside antibiotics, contaminants in foods or medicinals, and environmental or industrial pollutants. A human can have a hearing disorder that results from aging, or the human can have tinnitus (characterized by ringing in the ears).
[0101] The cells can be administered by any suitable method. For example, to restore hearing, inner ear cells generated by a method described herein can be transplanted, such as in the form of a cell suspension, into the ear by injection, such as into the luminae of the cochlea. See, e.g., the methods described in Corrales et al., J. Neurobiol. 66(13):1489-500 (2006) and Hu et al., Experimental Cell Research 302:40-47 (2005). Injection can be, for example, through the round window of the ear or through the bony capsule surrounding the cochlea. The cells can be injected through the round window into the auditory nerve trunk in the internal auditory meatus or into the scala tympani. In a preferred embodiment, the cells are administered into or near the sensory epithelium of the subject, e.g., into a fluid (perilymph)-filled space above or below the sensory epithelium, i.e., the scala media, scala tympani, or scala vestibuli.
[0102] Alternatively, a human suitable for the therapeutic compositions and methods described herein can include a human having a vestibular dysfunction, including bilateral and unilateral vestibular dysfunction. Vestibular dysfunction is an inner ear dysfunction characterized by symptoms that include dizziness, imbalance, vertigo, nausea, and fuzzy vision and may be accompanied by hearing problems, fatigue and changes in cognitive functioning. Vestibular dysfunction can be the result of a genetic or congenital defect; an infection, such as a viral or bacterial infection; or an injury, such as a traumatic or nontraumatic injury. Vestibular dysfunction is most commonly tested by measuring individual symptoms of the disorder (e.g., vertigo, nausea, and fuzzy vision). In these embodiments, the inner ear cells generated by a method described herein can be transplanted, such as in the form of a cell suspension, e.g., by injection, into an organ of the vestibular system, e.g., the utricle, ampulla and sacculus. The cells would generally be injected into the perilymph of these organs or into the vestibule (which connects the 3 organs).
[0103] Following treatment with an inner ear cell or inner ear cell progenitor as described herein, the human can be tested for an improvement in hearing or in other symptoms related to inner ear disorders. Methods for measuring hearing are well-known and include pure tone audiometry, air conduction, and bone conduction tests. These exams measure the limits of loudness (intensity) and pitch (frequency) that a human can hear. Hearing tests in humans include behavioral observation audiometry (for infants to seven months), visual reinforcement orientation audiometry (for children 7 months to 3 years) and play audiometry for children older than 3 years. Oto-acoustic emission testing can be used to test the functioning of the cochlear hair cells, and electro-cochleography provides information about the functioning of the cochlea and the first part of the nerve pathway to the brain.
[0104] The therapeutic compositions and methods described herein can be used prophylactically, such as to prevent hearing loss, deafness, or other auditory disorder associated with loss of inner ear function. For example, a composition containing a differentiation agent can be administered with a second therapeutic, such as a therapeutic that may effect a hearing disorder. Such ototoxic drugs include the antibiotics neomycin, kanamycin, amikacin, viomycin, gentamycin, tobramycin, erythromycin, vancomycin, and streptomycin; chemotherapeutics such as cisplatin; nonsteroidal anti-inflammatory drugs (NSAIDs) such as choline magnesium trisalicylate, diclofenac, diflunisal, fenoprofen, flurbiprofen, ibuprofen, indomethacin, ketoprofen, meclofenamate, nabumetone, naproxen, oxaprozin, phenylbutazone, piroxicam, salsalate, sulindac, and tolmetin; diuretics; salicylates such as aspirin; and certain malaria treatments such as quinine and chloroquine.
[0105] For example, a human undergoing chemotherapy can also be administered an inner ear cell or inner ear cell progenitor as described herein, by a method described herein. The chemotherapeutic agent cisplatin, for example, is known to cause hearing loss. Therefore, a composition containing a differentiation agent can be administered with cisplatin therapy to prevent or lessen the severity of the cisplatin side effect. An inner ear cell or inner ear cell progenitor as described herein can be administered before, after and/or simultaneously with the second therapeutic agent. The two treatments generally will be administered by different routes of administration.
[0106] The compositions and methods featured in the invention are appropriate for the treatment of hearing disorders resulting from sensorineural hair cell loss or auditory neuropathy. For example, patients with sensorineural hair cell loss experience the degeneration of cochlear hair cells, which frequently results in the loss of spiral ganglion neurons in regions of hair cell loss, and may also experience loss of supporting cells in the organ of Corti, and degeneration of the limbus, spiral ligament, and stria vascularis in the temporal bone material. Such patients may benefit particularly from administration of supporting cells and/or hair cells into the inner ear.
EXAMPLES
[0107] The invention is further described in the following examples, which do not limit the scope of the invention described in the claims.
Example 1: Sensory Progenitors from Mesenchymal Stem Cells
[0108] Mesenchymal stem cells were obtained from mouse bone marrow by culturing adherent cells from the marrow under high serum conditions.
[0109] Briefly, cells were obtained from bilateral femurs and tibias of 4 week old C57BL/6 or Atoh1-nGFP mice (Helms et al., Development 127, 1185-1196 (2000)) by flushing out the bone marrow with MEM-.alpha. (Gibco/BRL) containing 10% fetal bovine serum (FBS; BioWhittaker, Cambrex, N.Y.) and 1 mM glutamine (Gibco/BRL). Pelleted cells were resuspended and mixed with RBC lysis buffer (Gibco/BRL). Approximately 5.times.10.sup.6 cells were cultured on a 10 cm dish overnight in MEM-.alpha. with 9% horse serum, 9% FBS, 1% Gluta-Max (Invitrogen) and 100 units/ml penicillin and streptomycin (100 .mu.g/ml, Sigma) at 37.degree. C. in a 5% CO.sub.2 atmosphere. Nonadherent hematopoietic stem cells were removed, leaving adherent bone marrow stromal cells. When the cells became confluent, trypsinization was performed and the cells were cultured and passaged three to five times, with media changes every 3-4 days. These cells are referred to as mesenchymal stem cells (MSC).
[0110] Immunohistochemistry was performed as follows. Cells were fixed for 10 min with 4% paraformaldehyde in PBS. Immunostaining was initiated by rehydrating and blocking the sections for 1 h with 0.1% Triton X-100 in PBS supplemented with 1% BSA and 5% goat serum (PBT1). Fixed and permeabilized cells or rehydrated sections were incubated overnight in PBT1. CD34, CD44, CD45, Sca-1 antibodies (BD Biosciences) diluted 1:40 were used for the characterization of extracted bone marrow cells. Hair cells and bone marrow progenitors were characterized using monoclonal antibody to chick hair cell specific antigen diluted 1:500 (gift from Guy Richardson (Bartolami et al., J Comp Neurol 314, 777-788 (1991)); polyclonal antibody to myosin VIIa, 1:500 (Oshima et al., J Assoc Res Otolaryngol. 8(1):18-31 (2007)); monoclonal antibody to nestin, 1,000 (Developmental Studies Hybridoma Bank, Iowa City, Iowa); polyclonal antibody to parvalbumin 3, 1:2,000 (Heller et al., J Assoc Res Otolaryngol 3, 488-498 (2002)); monoclonal antibody to Atoh1, 1:100 (Developmental Studies Hybridoma Bank); monoclonal antibody to neurofilament M, 1:200 (Chemicon); Polyclonal antibody to collagen type II, 1:40 (Chemicon); polyclonal antibody to Brn3c (Covance, Princeton); Cy-5 conjugated F-actin 1:1000 (Molecular probe). Samples were washed three times for 20 min each with PBS. Anti-rabbit, anti-guinea pig and anti-mouse secondary antibodies conjugated with FITC-, TRITC-, and Cy-5-(Jackson ImmunoResearch) were used to detect primary antibodies. The samples were counterstained with DAPI for 10 min (Vector Laboratories) and viewed by epifluorescence microscopy (Axioskop 2 Mot Axiocam, Zeiss) or confocal microscopy (TCS, Leica). The counting of immunopositive cells was performed by counting 300 cells in 20 randomly selected microscopic fields and significance was calculated by Student's t-test.
[0111] Flow cytometric analysis was also performed. MSC were incubated with antibodies to CD34, CD44, CD45 or Sca-1 (BD Biosciences) and further incubated with secondary anti-mouse antibody conjugated to TRITC. Data were acquired and analyzed using an Agilent 2100 Bioanalyzer system and flow cytometry chips (Agilent Technology Inc., Palo Alto, Calif.). The reference window was set so that fluorescence from the secondary antibody alone was less than 2%.
[0112] The MSCs were negative for CD34 and CD45, markers for hematopoietic stem cells in bone marrow (Jiang et al., Nature 418, 41-49 (2002); Pittenger et al., Science 284, 143-147 (1999)) and positive for CD44 and Sca-1, markers for MSCs (Dezawa et al., J Clin Invest 113, 1701-1710 (2004)). Sca-1 was present on 5.2% of the cells and CD44 was present on 11.5% of the cells based on immunohistochemistry and the percentages determined by flow cytometry were similar (FIGS. 1A and 1D and Table 1). We detected co-expression of CD44 and nestin as well as Sca-1 and nestin on a small percentage of the cells (FIGS. 1B and 1C).
TABLE-US-00004 TABLE 1 Co-Expression of CD44 and Sca-1 with Nestin in Mesenchymal Stem Cells pre-induction (%) post-induction (%) Nestin (+) cells 4.7 .+-. 0.8 14.2 .+-. 2.0 CD44 (+) cells 11.5 .+-. 1.6 11.9 .+-. 1.8 Sca-1 (+) cells 5.2 .+-. 1.5 5.0 .+-. 0.4 CD 44 & nestin (+) cells 3.4 .+-. 0.9 9.9 .+-. 0.9 Sca-1 & nestin (+) cells 2.8 .+-. 1.2 4.3 .+-. 0.5 Positive cells were counted in relation to total nuclei stained by DAPI. Data are mean .+-. SE for 10 separate experiments. The increase in cells staining with nestin was significant (p < 0.001) as was the increase in the cells staining for both nestin and CD44 (p < 0.001) and nestin and Sca-1 (p < 0.05).
[0113] We confirmed the previously reported capacity of MSCs to be converted to chondrocytes (Pittenger et al., Science 284, 143-147 (1999)) and neurons (Dezawa et al., J Clin Invest 113, 1701-1710 (2004)). For chondrogenic differentiation, MSC were formed into a micropellet and cultured in DMEM with 10 ng/ml TGFbeta1, 6.25 ug/ml transferrin and 6.25 ug/ml insulin for 2 weeks. Their potential to differentiate into chondrocytes is demonstrated in FIG. 1E. For neuronal differentiation, MSC were cultured in DMEM/F12 1:1 containing N2/B27 supplement with bFGF (10 ng/ml) for 14 days and for 7 days without FGF. This resulted in differentiation to neurons (Dezawa et al., J Clin Invest 113, 1701-1710 (2004)) as shown by neuronal markers (FIG. 1F).
[0114] To determine whether otic vesicle growth factors that are important in the early development of inner ear progenitor cells could have a similar effect on MSCs, we removed the serum from the MSCs after 3-5 passages and cultured the cells in serum-free medium containing IGF-1, EGF and bFGF.
[0115] For the induction of progenitor cells, passage 3-5 MSC were trypsinized and transferred to 6-well plates or 4 well plates (BD Bioscience) coated with poly-L-ornithine and gelatin or fibronectin (Sigma) at 5.times.10.sup.4 cells/ml. Cells were cultured for 5-7 days, and then cultured in serum-free medium composed of DMEM/F12 1:1 containing N2/B27 supplements (Invitrogen). For progenitor cell induction, we used a combination of EGF (20 ng/ml) and IGF (50 ng/ml; R&D Systems, Minneapolis, Minn.) for 2 weeks followed by the addition of bFGF (10 ng/ml) plus the other growth factors for an additional 2 weeks, or a combination of NT3 (30 ng/ml) and bFGF (10 ng/ml) for 4-5 days followed by NT3 (30 ng/ml) and BDNF (10 ng/ml) for 7 days.
[0116] Semiquantitative RT-PCR was performed as follows. Total RNA was extracted with the RNAeasy minikit (Qiagen, Valencia, Calif.) according to the manufacturer's instructions. For reverse transcription, 6 .mu.g of total RNA was used with SuperScript III transcriptase (Invitrogen) and oligo-dT primers. The PCR cycling conditions were optimized in pilot experiments. Specific cycling parameters were: initial denaturation step at 94.degree. C. for 2 minutes, denaturation 94.degree. C. for 30 seconds, annealing temperature optimized between 56-60.degree. C. for 30 seconds, extension 72.degree. C. for 60 seconds, extension 72.degree. C. for 60 seconds, and followed by 7 minutes of terminal extension at 72.degree. C. after the last cycle. The number of cycles was optimized between 30 and 35, and conditions were kept constant for each primer. The presented data are from experiments repeated at least 5 times. Control PCR without reverse transcriptase did not produce specific bands. The primer pairs and cDNA product lengths were as follows:
TABLE-US-00005 TABLE 2 RT-PCR-Primer Pairs and cDNA Product Length Forward SEQ ID Reverse SEQ ID Expected cDNA target primer NO: primer NO: product length Oct4 ATG GCT 7. TTA ACC 8. 1033 bp GGA CAC CCA AAG CTG GCT CTC CAG TCA G GTT C Otx2 CCA TGA 9. GAA GCT 10. 211 bp CCT ATA CCA TAT CTC AGG CCC TGG CTT CAG G GTG GAA AG Sox2 CAC CCG 11. TCC CCT TCT 12. 414 bp GGC CTC CCA GTT AAC GCT CGC AGT CAC G CCA Pax2 CCA AAG 13. GGA TAG 14. 544 bp TGG TGG GAA GGA ACA AGA CGC TCA TTG CC AAG AC Pax6 AGA CTT 15. TAG CCA 16. 589 bp TAA CCA GGT TGC AGG GCG GT GAA GAA CT Nestin AAC AGA 17. CTT CAG 18. 392 bp GAT TGG AAA GGC AAG GCC TGT CAC GCT GGC AGG AG Musashi ATG GAG 19. ATC TTC TTC 20. 332 bp ACT GAC GTC CGA GCG CCC GTG AC CAG GATA3 CCT CCG 21. ACC GTA 22. 319 bp ACG GCA GCC CTG GGA GTC ACG GAG TTT Math1 AGA TCT 23. ACT GGC 24. 449 bp ACA TCA CTC ATC ACG CTC AGA GTC TGT C ACT G Neurogenin- TGG TGT 25. AAG GCC 26. 400 bp 1 CGT CGG GAC CTC GGA AC CAA ACC TC NeuroD ACG GGC 27. TGA AAG 28. 513 bp TGA ACG AGA AGT CGG CGC TGC CAT TGG AC TGA TG Brn3c GCC ATG 29. ATG GCG 30. 714 bp CGC CGA CCT AGA GTT TGT C TGA TGC Espin CAG CCT 31. TGA CCT 32. 475 bp GAG TCA GTC GCT CCG CAG GCC AGG CCT C GCG CG Myo7a CTC CCT 33. AAG CAC 34. 628 bp CTA CAT CTG CTC CGC TCT CTG CTC GTT CG GTC CAC G Zic1 GGC CAA 35. GAG AGC 36. 425 bp CCC CAA TGG GGT AAA GTC GCG TGT GTG AGG A Zic2 GGC GGC 37. TTG CCA 38. 405 bp GCA GCT CAG CCC CCA CAA GGG AAA CCA GTA GGA CAG TrkB TTG CCC 39. CGC TTG 40. 46 bp CTT CCC CTT CTC GCT TTA T CTC GT TrkC ACC CGC 41. TCC CGG 42. 521 bp ATC CCA TGT ACA GTC AT AAG TGC P27Kip CTG GAG 43. CGT CTG 44. 525 BP CGG ATG CTC CAC GAC GCC AGT GCC AGA C AGC Jag2 GTC CTT 45. GTT TCC 46. CCC ACA ACC TTG TGG GAG TT ACC TCG GT Notch1 AGA GAT 47. CAC ACA 48. 306 BP GTG GGA GGG AAC TGC AGG AC TTC ACC CT P75 GTC GTG 49. CTG TGA 50. GGC CTT GTT CAC GTG GCC ACT GGG G S100 GCC AAC 51. ACG TCG 52. 423 bp CGT GTG AGA CTG CTG CTG GGC AAG G Clal4 CCA GCA 53. GGG GCA 54. 664 bp CAG CGG CGG TTG TCC AG TCC TTG TAG Con26 CGG AAC 55. CTA AGC 56. 824 bp CAG AGA ACG GGT TAG GAC TGC CTC CTA C ATC C Gapdh AAC GGG 57. CAG CCT 58. 442 bp AAG CCC TGG CAG ATC ACC CAC CAG
[0117] When the expression of neural progenitor cell markers in the resulting cultures was assessed, Otx2, nestin, Sox2, and Musashi were expressed in increased amounts in these cells, which are subsequently referred to herein as progenitor cells, relative to MSCs based on RT-PCR (FIG. 2A). Pax6 was found in the progenitor cells but not in the MSCs (FIG. 2A). Pax2 was not expressed. A low level of Pax5 was detected but Pax8 was not expressed (data not shown). A similar pattern of expression was seen for the stem cell marker, Oct4, which was expressed in the progenitor cells but interestingly, given its role in maintaining the pluripotency of stem cells, was not found in the MSCs. The increase in expression of nestin in the progenitor cells relative to the MSCs (FIG. 2A) was confirmed by immunohistochemistry (FIGS. 2B and 2C and Table 1) and was significant (p<0.001). Additional markers of the hair cell and neural lineages (Atoh1, Brn3c, GATA3) and neuronal markers (TrkB and TrkC) were also expressed in the progenitors (FIG. 2A).
[0118] Because of the expression of TrkB and TrkC in the progenitor cell populations, we tested whether incubation with NT-3 and BDNF, the neurotrophins that bind to these receptors, would increase the yield of progenitor cells or alter the expression of genes for hair cell or neuronal fate. We found an increase in expression of Otx2, Sox2, nestin, and Musashi under these conditions as well as an increase in Oct4 expression (FIG. 3A), indicating that the cells may have adopted a neural progenitor cell fate. The neurotrophin-mediated conversion to progenitor cells had a more rapid time course that we found for EGF, IGF-1 and bFGF alone. The expression of proneural transcription factors, NeuroD and Ngn1, as well as neural and hair cell lineage markers, GATA3, Atoh1, and Brn3c, were also increased and the expression of Ngn1 and NeuroD, which select for a neural over a hair cell fate in the inner ear (Kim et al., Development 128, 417-426 (2001); Matei et al., Dev Dyn. 234(3):633-50 (2005)) were higher when NT-3 and BDNF were included in the differentiation medium. Other transcription factors expressed in the otic precursors during development, Zic2 and Pax6, were elevated in the progenitor cells relative to the MSCs, and Zic1 expression was not observed. This suggests that NT-3 and BDNF induced the formation of cells of a neural lineage that were potentially destined to become both neurons and hair cells. However, the cells were not converted to hair cells or neurons because markers for these cells were not found (FIG. 3A, hair cell markers myosin VIIa and espin). We also tested for the expression of genes characteristic of other epithelial cells in the cochlea such as supporting cells, because the progenitors for hair cells can include or give rise to these cells and found that the progenitors expressed S100A, p75.sup.trk, claudin 14, connexin 26, and Notch1.
[0119] The observation of supporting cell markers from the MSC-derived progenitor cells after growth factor induction may be correlated to their origin from a common progenitor during in vivo development (Matei et al., Dev Dyn. 234(3):633-50 (2005); Satoh and Fekete, Development 132, 1687-1697 (2005)). Since hair cells can be induced to develop from supporting cells after introduction of the Atoh1 gene (Izumikawa et al., Nat Med 11, 271-276 (2005); Zheng and Gao, Nat Neurosci 3, 580-586 (2000)), the role of supporting cells as potential progenitors for hair cells via transdifferentiation has been discussed (Izumikawa et al., Nat Med 11, 271-276 (2005)). The expression of supporting cell genes may reflect an intermediate or accompanying stage on the way to becoming hair cells; in Atoh1 knockout mice undifferentiated cells with markers of supporting cells have been observed to activate the Atoh1 gene (Fritzsch et al., Dev Dyn 233, 570-583 (2005); Woods et al., Nat Neurosci 7, 1310-1318 (2004)). Alternatively, supporting cells could be induced by the developing hair cells: ectopic hair cells in the greater epithelial ridge induced supporting cell markers in surrounding cells (Woods et al., Nat Neurosci 7, 1310-1318 (2004)). The MSCs could be induced to become hair cell progenitors by bFGF, EGF and IGF-1, factors that potentially stimulate the in vivo formation of these progenitors (Leon et al., Endocrinology 136, 3494-3503 (1995); Pauley et al., Dev Dyn 227, 203-215 (2003); Zheng et al., J Neurosci 17, 216-226 (1997)), and these progenitors were able to give rise to hair cells after overexpression of Atoh1. An increase in expression of neural progenitor markers could be caused by expansion of the cells that express these markers or by differentiation of MSCs to the neural progenitor phenotype.
[0120] As described herein, MSC-derived progenitor cells expressed neurotrophin receptors. BDNF and NT-3 play important roles in maturation of inner ear neurons (Fritzsch et al., J Neurosci 17, 6213-6225 (1997); Pirvola and Ylikoski, Curr Top Dev Biol 57, 207-223 (2003)), and in differentiation of neural stem cells to neurons (Ito et al., J Neurosci Res 71, 648-658 (2003)), and we therefore tested whether the fate of the progenitors could be modulated by neurotrophins. Incubation with these factors resulted in enrichment of progenitors that could be converted to hair cells by subsequent Atoh1 overexpression (Izumikawa et al., Nat Med 11, 271-276 (2005); Zheng and Gao, Nat Neurosci 3, 580-586 (2000)) or co-culture with chick otocyst cells. Since NT-3 and BDNF were found to increase both Atoh1 expression and differentiation in neural stem cells (Ito et al., J Neurosci Res 71, 648-658 (2003)), neurotrophins could directly increase differentiation of MSCs or could increase their competence to respond to overexpressed Atoh1.
[0121] Analysis of the progenitor cells obtained from the MSCs revealed parallels with natural development of the inner ear sensory epithelia. The MSC-derived progenitors expressed Sox2, which must be present for subsequent hair cell differentiation in the developing otocyst (Kiernan et al., Nature 434, 1031-1035 (2005)). The expression of Atoh1 in cells that did not have myosin VIIa and the appearance of myosin VIIa at later time points is consistent with the order of their expression during development based on immunohistochemistry (Chen et al., Development 129, 2495-2505 (2002)). The lack of Pax2 expression was surprising since the paired box transcription factor is ubiquitously expressed in the otocyst (Burton et al., Dev Biol 272, 161-175 (2004); Li et al., J Neurobiol 60, 61-70 (2004)). This may suggest that Pax2 is not required or that it can be replaced by another factor for the conversion of MSCs to hair cells. Pax5 was detected and may substitute for Pax2 based on their functional equivalence (Bouchard et al., Development. 127(5):1017-28 (2000)). This is consistent with the analysis of the Pax2 null mouse (Burton et al., Dev Biol 272, 161-175 (2004)), which appears to develop hair cells despite severe disruption of the normal morphology of the cochlea. The lack of Zic1 expression relative to Zic2 is also found during development of a hair cell phenotype as compared to sensory neurons in the otocyst (Warner et al., Dev Dyn 226, 702-712 (2003)) and is thus consistent with the development of a hair cell phenotype. The identification of inductive molecules on chick otocyst cells that are not present in conditioned media will provide further insights into hair cell differentiation.
[0122] The isolation of progenitor cells that can give rise to the tissue of origin, as observed in the inner ear (Li et al., Trends Mol Med 10, 309-315 (2004); Li et al., Nat Med 9, 1293-1299 (2003a)), might be predicted and yet the cells do not regenerate after damage, possibly because of the decrease in number of inner ear stem cells after birth (Oshima et al., J Assoc Res Otolaryngol. 8(1):18-31 (2007)). Therefore, a source of cells to provide replacements for these sensory cells is highly desirable. The in vivo role of MSCs in regeneration generally remains uncertain although bone marrow could act as a source of new cells in organs with few progenitors. Despite the demonstration that cells from bone marrow migrate into the brain and heart in adults (Oshima et al., J Assoc Res Otolaryngol. 8(1):18-31 (2007)); Weimann et al., Proc Natl Acad Sci USA 100, 2088-2093 (2003)) and differentiate into neurons in the brain, hematopoietic stem cells from bone marrow were not converted to cardiomyocytes after injection (Murry et al., Nature 428, 664-668 (2004)) and conversion to neurons was extremely rare (Wagers et al., Science 297, 2256-2259 (2002); Weimann et al., Proc Natl Acad Sci USA 100, 2088-2093 (2003)). The most successful attempts at regeneration by adult stem cells from other tissues have been obtained after a lesion Doyonnas et al., Proc Natl Acad Sci USA 101, 13507-13512 (2004); Edge, Transplant Proc 32, 1169-1171 (2000); Hess et al., Nat Biotechnol 21, 763-770 (2003); Pagani et al., J Am Coll Cardiol 41, 879-888 (2003)) and tissue damage may be required to see cell replacement by bone marrow-derived cells. Whether bone marrow-derived cells play any regenerative role in the sensory or peripheral nervous system in a spontaneous response to damage in vivo is an unanswered question, but, although low-level replacement of hair cells by bone marrow cells in vivo cannot be ruled out, spontaneous replacement of sensory cells is unlikely to be significant given the lack of hair cell regeneration seen in the adult cochlear and vestibular systems (Hawkins and Lovett, Hum Mol Genet 13(Spec No 2):R289-296 (2004); White et al., Nature 441, 984-987 (2006)).
Example 2: Transfection with an Atoh1 Expression Plasmid Converts Progenitors to Hair Cells
[0123] To test whether the progenitor cells could act as inner ear precursor cells, it was evaluated whether overexpression of Atoh1, a transcription factor that is known to push competent progenitors to a hair cell fate (Izumikawa et al., Nat Med 11, 271-276 (2005); Zheng and Gao, Nat Neurosci 3, 580-586 (2000)), would increase the expression of hair cell markers.
[0124] The efficiency of Atoh1 transfection was tested by counting green fluorescent cells after transfection with a vector coding for GFP expression in addition to Atoh1. We constructed a vector containing the Atoh1 coding sequence under EF1alpha-promoter control in the pTracer-EF vector (Invitrogen) that has a GFP-Zeocin fusion sequence under the CMV promoter. Gene transfection was done in the progenitor cell state or as MSC using LIPOFECTAMINE.TM. transfection reagent (Sigma). Cells were cultured in Zeocin (Invitrogen) to obtain stable transfectants. Transfected MSC were cultured in the serum-free conditions with combinations of growth factors.
[0125] When MSCs were transfected, as many as 2% of the cells were GFP positive at 24 hours (FIG. 4A). RT-PCR at day 14 showed that the transfected cell population expressed markers of developing sensory epithelia, such as p27Kip, Brn3c and jagged2, and mature hair cells markers, myosin VIIa and espin (FIG. 4B) as well as increased expression of Ngn1 and NeuroD. We also detected expression of supporting cell markers, S100A, p75.sup.Trk, claudin 14, connexin 26, and Notch1, indicating that the progenitor cells could give rise to hair cells and supporting cells (FIG. 4C). Selection of MSC transfectants with stable Atoh1 expression increased the percentage of GFP-positive cells (FIG. 4D). Incubation of these cells in the growth factors described above followed by immunohistochemistry yielded cells with expression of Atoh1 and myosin VIIa respectively in 7.7% and 7.1% of the total cells (FIG. 4E). Differentiation under growth factor stimulation gave rise to cells with Brn3c in the nucleus and myosin VIIa in the cytoplasm (FIG. 4F). These cells were positive for both markers in the same cells, with 92% of the Atoh1-positive cells showing staining for myosin VIIa, and 77% of the Brn3c-positive cells showing staining for myosin VIIa. Examination of the myosin VIIa positive cells for F-actin (FIGS. 4G and H) indicated that some of the cells (4.9% of the myosin VIIa-positive cells) had developed protrusions at their apical poles. These protrusion had the polarized appearance of stereociliary bundles and were positive for espin (FIG. 4G).
[0126] Atoh1 expression led to strong expression of helix-loop-helix transcription factors, Ngn1 and NeuroD. Several previous studies have indicated that Atoh1 expression can increase these transcription factors. In mouse cerebellum Atoh1 expression leads to overexpression of NeuroD (Helms et al., Mol Cell Neurosci 17, 671-682 (2001)). In zebrafish NeuroD is not expressed in the absence of Atoh1 (Sarrazin et al., Dev Biol 295, 534-545 (2006)) and is required for hair cell formation. The related mouse achaete-scute (Mash1) upregulates Ngn1 (Cau et al., Development 124, 1611-1621 (1997)). However, Ngn1 was downregulated by overexpression of Atoh1 in chick neural tube (Gowan et al., Neuron 31, 219-232 (2001)).
[0127] These data demonstrate that overexpression of Atoh1 in growth-factor induced progenitor cells induces the differentiation of a percentage of those cells to hair cells.
Example 3: Conversion of Progenitors to Hair Cells is Stimulated by Developing Otocyst Cells
[0128] To test whether the developing otocyst produced factors that would increase the differentiation of MSCs to hair cells, co-culture experiments of E3 chick otocyst cells with MSCs were performed.
[0129] Embryos of the white leghorn strain (Charles River) were harvested 72 hours after placing fertilized eggs onto rocking platforms in a humidified incubator maintained at 38.degree. C. The dissection of otocysts from the extracted embryos was done in cooled PBS, pH 7.2, after removal of periotic mesenchymal tissues. The otocysts were trypsinized and dissociated to single cells for plating and 2.times.10.sup.4 cells were cultured overnight in four-well plates in 10% FBS. One day after plating, the otocyst cells were fixed with 4% paraformaldehyde for 20 minutes, or inactivated with mitomicin C (10 .mu.g/ml) for 3 hours, then washed 4 times with PBS. Conditioned medium from the cultured cells was collected and frozen prior to use on progenitors cells. Progenitor cells (5.times.10.sup.4 cells/ml) induced in serum-free medium with growth factors, were overlaid on the chick otocyst cells and cultured for 5-7 days with EGF/IGF, followed by 10 days with EGF/IGF/FGF and withdrawal of growth factors for 5-10 more days. The cells were analyzed by RT-PCR or immunohistochemistry as described herein.
[0130] After culture in the presence of the chick otocyst cells for 21 days, increased expression of myosin VIIa, jagged2, p27Kip, Brn3c and Atoh1 by RT-PCR was found (FIG. 5A). The factor(s) was unlikely to be a secreted molecule because fixation of the cells did not diminish their ability to promote differentiation after exposure for 14 days, while conditioned medium was ineffective in 14 days (FIG. 5A). Conversion of the stem cells to hair cells could be followed by appearance of green fluorescence in the cultures using MSCs derived from transgenic Atoh1-nGFP mice that express a nuclear version of enhanced GFP when Atoh1 enhancer elements are activated (Chen et al., Development 129, 2495-2505 (2002); Lumpkin et al., Gene Expr Patterns 3, 389-395 (2003)). These green cells were observed in the co-cultures with chick otocyst cells (FIG. 5B) and the cells were co-labeled with antibody to myosin VIIa.
[0131] The otocyst from E3 chick embryos were used for injection of progenitor cells. The dissected otocysts were transferred into 7 ml of serum-free DMEM/F12 1:1 containing N2 and B27 on a gelatin-coated tissue culture dish. After attachment of intact otocysts, progenitor cells from MSC (5.times.10.sup.7 cells/ml) were injected into the otocyst with a micropipette in 2 .mu.l of medium. The left otic vesicles did not receive cell grafts and served as controls. The otocysts were harvested after 10-14 days, fixed 30 min in paraformaldehyde (4% in PBS), cryoprotected overnight in sucrose (30% in PBS), embedded in TissueTek (EMS) and serially sectioned (16 .mu.m) with a cryostat (CM3050, Leica, Nussloch, Germany).
[0132] When the progenitor cells were injected into chick otocysts obtained at E3, conversion of progenitors to cells with hair cell properties (5% of the myosin VIIa-positive cells were positive for nGFP) was observed (FIG. 6A). The murine hair cells were seen to incorporate into the hair cell bearing epithelia of the developing chicken otocyst as detected by expression of GFP (FIG. 6B). One possible explanation for the expression of hair cell genes by the MSC-derived cells in co-culture is fusion with chick cells. To rule this out we labeled the cells with an antibody to chick hair cell antigen (Bartolami et al., J Comp Neurol 314, 777-788 (1991)). Native chick hair cells could be detected lining the internal cavity of the otocyst (51% of 1,352 cells from 15 otocyst injections that stained for myosin VIIa were positive for chick hair cell antigen), and the cells that expressed nGFP and hair cell markers did not co-express chick hair cell antigen (FIG. 6C) and were therefore of mouse origin and not the product of cell fusion.
[0133] These experiments, performed in an attempt to understand how contact of the MSCs with developing otocyst cells provided a signal that induced their differentiation to hair cells, demonstrated that the inductive effect was through a cell surface molecule as opposed to a secreted factor. Injection of the MSC into the developing otocyst in vitro indicated that hair cells that differentiated from the stem cells were integrated into the chick otocyst epithelium, demonstrating that the environment provided by developing chicken otocyst cells could guide differentiation and integration of suitable progenitor cells. The instructive influence has also been seen previously with inner ear-derived stem cells and murine ES cell-derived progenitor cells (Li et al., Trends Mol Med 10, 309-315 (2004); Li et al., Nat Med 9, 1293-1299 (2003); Li et al., Proc Natl Acad Sci USA 100, 13495-13500 (2003).
[0134] The effect of the co-incubation with otocyst cells may be simply to activate Atoh1 expression and a sufficient amount of Atoh1 may be required to allow hair cell differentiation since the MSCs had low levels of Atoh1 but did not have detectable sensory epithelial cell markers. This type of high level expression could be needed for Atoh1 to overcome the level of preexisting endogenous inhibitors that interact with Atoh1 protein. The murine cells could be clearly distinguished from the chick hair cells that differentiated at the same time by their expression of nGFP and by immunolabeling of the chick hair cells with a species-specific antibody. The cells were never co-stained (based on examination of 1,352 cells), indicating that the mouse hair cells had differentiated from stem cells and did not arise from cell fusion.
Example 4: Inhibition of Notch Signaling Induces Differentiation of Hair Cells
[0135] The Notch pathway maintains the alternating pattern of hair cells and supporting cells in vivo by suppressing the differentiation of hair cells from supporting cells and activation of Notch in the embryo appears to block development of hair cells from progenitors.
[0136] To examine the effect of the Notch pathway on the differentiation of hair cells, the NT3/BDNF treated progenitors were incubated with a .gamma.-secretase inhibitor. Analysis of gene expression in the progenitors made by incubation with NT3, BDNF, FGF and subsequently treated with the .gamma.-secretase inhibitor demonstrated that loss of the notch signaling increased Atoh1 expression. Atoh1 levels rose compared to the treatment with growth factors alone based on RT-PCR when the inhibitor was used at 1 .mu.M (FIG. 7). The timing of the addition of the inhibitor was essential with inhibition at a later stage (after 3 days of differentiation in vitro) causing less induction of hair cell markers than inhibition starting at day 0 and continuing for 10 days. At the low concentration, .gamma.-secretase inhibitor activates ngn1 and NeuroD and causes no increase in Atoh1 or hair cell markers. At higher concentration, the .gamma.-secretase inhibitor increases Atoh1 and Brn3c expression. The increased Atoh1 appeared to be able to produce hair cells as the cells expressed markers for the hair cells such as myosin7a, p27Kip. As HLH transcription factors mediate the effects of the Notch pathway, this result is consistent with the role of Notch and suggests a mechanism for preventing hair cell differentiation under normal conditions.
Example 5: Inhibition of Hair Cell Differentiation in Human Stem Cells
[0137] To determine if human mesenchymal stem cells (hMSCs) can be differentiated into inner ear cell types including hair cells or sensory neurons, human bone marrow cells from healthy adults were evaluated.
[0138] The human bone marrow cells were harvested and cultured as plated on tissue culture plastic for 16 hours, and nonadherent hematopoietic stem cells were aspirated.
[0139] First, the adherent cells were cultured in .alpha.MEM containing 9% horse serum and 9% fetal bovine serum and were negative for blood-forming cell markers, CD34 and CD45. These cells gave rise to chondrocytes expressing type II and IV collagen after culture in the presence of TGF.beta., transferrin and insulin.
[0140] Culture of hMSCs in DMEM/F12 medium containing N2 and B27 without serum in the presence of NT-3, BDNF, Sonic hedgehog and retinoic acid for 10 days gave rise to cells that expressed neurosensory progenitor markers detected by RT-PCR, Musashi, nestin, Pax6, Brn3a, NeuroD, Ngn1, and GATA3, and sensory neuron markers, peripherin and TrkC. These differentiated hMSCs were positive for tubulin (2.1% of the total cells were positive based on immunohistochemistry) and, of these cells, 28% co-stained for peripherin and 31% co-stained for Brn3a.
[0141] For the differentiation to hair cells, hMSCs were transfected with human Atoh1 in an expression vector with a selectable marker for eukaryotic cells. The selected progenitor cells expressed Atoh1 and, after differentiation in DMEM/F12 medium containing N2 and B27 with NT-3 and BDNF for 10 days, expressed hair cell markers, Atoh1, myosin VIIa, p27Kip, Jag2 and espin based on RT-PCR.
[0142] The ability of these cells to engraft in an organ of Corti, the Atoh1-transfected cells were co-cultured with an ex vivo organ of Corti from mouse. This gave rise to cells expressing myosin VIIa and espin that were detected by immunostaining, i.e., differentiated hair cells. When the ex vivo mouse organ of Corti was treated with toxins to induce hair cell degeneration, co-cultured bone marrow-derived cells were observed to engraft in the mouse sensory epithelium, thus demonstrating the ability of cells obtained.
[0143] Thus, human MSCs are a potential alternative for cell-based treatment of hearing loss, as they can be differentiated into inner ear cell types including hair cells or sensory neurons, and can be successfully engrafted into structures of the inner ear.
Example 6: Math1-Estrogen Receptor (ER) Fusion Constructs
[0144] One alternative to constitutive expression of Math1 is to use a conditional or inducible system of gene expression, to upregulate Math1 with an inducer that is added to the cell medium or cochlear environment. An inducible model is particularly useful when investigating the temporal effects of gene expression.
[0145] This Example describes a system in which administration of tamoxifen, a synthetic estrogen agonist, induces expression of Math1. A Math1-estrogen receptor (ER) fusion protein, where the ER has been mutated so that it selectively binds to tamoxifen rather than estrogen, is constitutively expressed. In the absence of tamoxifen, the Math1-ER construct remains quiescent within the cytosol where it is inactivated by heat shock proteins. The addition of tamoxifen to the transfected cells results in a dose-dependent localization of the Math1-ER construct to the nucleus where it is transcribed leading to increased expression of Math1. The sequence of Math1 is given above.
[0146] The sequence of ER used is as follows (SEQ ID NO:59):
TABLE-US-00006 ATGTCCAATTTACTGACCGTACACCAAAATTTGCCTGCATTACCGGTCGA TGCAACGAGTGATGAGGTTCGCAAGAACCTGATGGACATGTTCAGGGATC GCCAGGCGTTTTCTGAGCATACCTGGAAAATGCTTCTGTCCGTTTGCCGG TCGTGGGCGGCATGGTGCAAGTTGAATAACCGGAAATGGTTTCCCGCAGA ACCTGAAGATGTTCGCGATTATCTTCTATATCTTCAGGCGCGCGGTCTGG CAGTAAAAACTATCCAGCAACATTTGGGCCAGCTAAACATGCTTCATCGT CGGTCCGGGCTGCCACGACCAAGTGACAGCAATGCTGTTTCACTGGTTAT GCGGCGGATCCGAAAAGAAAACGTTGATGCCGGTGAACGTGCAAAACAGG CTCTAGCGTTCGAACGCACTGATTTCGACCAGGTTCGTTCACTCATGGAA AATAGCGATCGCTGCCAGGATATACGTAATCTGGCATTTCTGGGGATTGC TTATAACACCCTGTTACGTATAGCCGAAATTGCCAGGATCAGGGTTAAAG ATATCTCACGTACTGACGGTGGGAGAATGTTAATCCATATTGGCAGAACG AAAACGCTGGTTAGCACCGCAGGTGTAGAGAAGGCACTTAGCCTGGGGGT AACTAAACTGGTCGAGCGATGGATTTCCGTCTCTGGTGTAGCTGATGATC CGAATAACTACCTGTTTTGCCGGGTCAGAAAAAATGGTGTTGCCGCGCCA TCTGCCACCAGCCAGCTATCAACTCGCGCCCTGGAAGGGATTTTTGAAGC AACTCATCGATTGATTTACGGCGCTAAGGATGACTCTGGTCAGAGATACC TGGCCTGGTCTGGACACAGTGCCCGTGTCGGAGCCGCGCGAGATATGGCC CGCGCTGGAGTTTCAATACCGGAGATCATGCAAGCTGGTGGCTGGACCAA TGTAAATATTGTCATGAACTATATCCGTAACCTGGATAGTGAAACAGGGG CAATGGTGCGCCTGCTGGAAGATGGGGATCAGGCTGGTGCCATGGGCGAT CCACGAAATGAAATGGGTGCTTCAGGAGACATGAGGGCTGCCAACCTTTG GCCAAGCCCTCTTGTGATTAAGCACACTAAGAAGAATAGCCCTGCCTTGT CCTTGACAGCTGACCAGATGGTCAGTGCCTTGTTGGATGCTGAACCGCCC ATGATCTATTCTGAATATGATCCTTCTAGACCCTTCAGTGAAGCCTCAAT GATGGGCTTATTGACCAACCTAGCAGATAGGGAGCTGGTTCATATGATCA ACTGGGCAAAGAGAGTGCCAGGCTTTGGGGACTTGAATCTCCATGATCAG GTCCACCTTCTCGAGTGTGCCTGGCTGGAGATTCTGATGATTGGTCTCGT CTGGCGCTCCATGGAACACCCGGGGAAGCTCCTGTTTGCTCCTAACTTGC TCCTGGACAGGAATCAAGGTAAATGTGTGGAAGGCATGGTGGAGATCTTT GACATGTTGCTTGCTACGTCAAGTCGGTTCCGCATGATGAACCTGCAGGG TGAAGAGTTTGTGTGCCTCAAATCCATCATTTTGCTTAATTCCGGAGTGT ACACGTTTCTGTCCAGCACCTTGAAGTCTCTGGAAGAGAAGGACCACATC CACCGTGTCCTGGACAAGATCACAGACACTTTGATCCACCTGATGGCCAA AGCTGGCCTGACTCTGCAGCAGCAGCATCGCCGCCTAGCTCAGCTCCTTC TCATTCTTTCCCATATCCGGCACATGAGTAACAAACGCATGGAGCATCTC TACAACATGAAATGCAAGAACGTGGTACCCCTCTATGACCTGCTCCTGGA GATGTTGGATGCCCACCGCCTTCATGCCCCAGCCAGTCGCATGGGAGTGC CCCCAGAGGAGCCCAGCCAGACCCAGCTGGCCACCACCAGCTCCACTTCA GCACATTCCTTACAAACCTACTACATACCCCCGGAAGCAGAGGGCTTCCC CAACACGATCTGA
ADDITIONAL REFERENCES
[0147] Kicic et al., J Neurosci 23, 7742-7749 (2003). [0148] Ma et al., J Assoc Res Otolaryngol 1, 129-143 (2000). [0149] Wang et al., Nature 422, 897-901 (2003).
OTHER EMBODIMENTS
[0150] It is to be understood that while the invention has been described in conjunction with the detailed description thereof, the foregoing description is intended to illustrate and not limit the scope of the invention, which is defined by the scope of the appended claims. Other aspects, advantages, and modifications are within the scope of the following claims.
Sequence CWU 1
1
5911065DNAHomo sapiens 1atgtcccgcc tgctgcatgc agaagagtgg gctgaagtga
aggagttggg agaccaccat 60cgccagcccc agccgcatca tctcccgcaa ccgccgccgc
cgccgcagcc acctgcaact 120ttgcaggcga gagagcatcc cgtctacccg
cctgagctgt ccctcctgga cagcaccgac 180ccacgcgcct ggctggctcc
cactttgcag ggcatctgca cggcacgcgc cgcccagtat 240ttgctacatt
ccccggagct gggtgcctca gaggccgctg cgccccggga cgaggtggac
300ggccgggggg agctggtaag gaggagcagc ggcggtgcca gcagcagcaa
gagccccggg 360ccggtgaaag tgcgggaaca gctgtgcaag ctgaaaggcg
gggtggtggt agacgagctg 420ggctgcagcc gccaacgggc cccttccagc
aaacaggtga atggggtgca gaagcagaga 480cggctagcag ccaacgccag
ggagcggcgc aggatgcatg ggctgaacca cgccttcgac 540cagctgcgca
atgttatccc gtcgttcaac aacgacaaga agctgtccaa atatgagacc
600ctgcagatgg cccaaatcta catcaacgcc ttgtccgagc tgctacaaac
gcccagcgga 660ggggaacagc caccgccgcc tccagcctcc tgcaaaagcg
accaccacca ccttcgcacc 720gcggcctcct atgaaggggg cgcgggcaac
gcgaccgcag ctggggctca gcaggcttcc 780ggagggagcc agcggccgac
cccgcccggg agttgccgga ctcgcttctc agccccagct 840tctgcgggag
ggtactcggt gcagctggac gctctgcact tctcgacttt cgaggacagc
900gccctgacag cgatgatggc gcaaaagaat ttgtctcctt ctctccccgg
gagcatcttg 960cagccagtgc aggaggaaaa cagcaaaact tcgcctcggt
cccacagaag cgacggggaa 1020ttttcccccc attcccatta cagtgactcg
gatgaggcaa gttag 10652354PRTHomo sapiens 2Met Ser Arg Leu Leu His
Ala Glu Glu Trp Ala Glu Val Lys Glu Leu1 5 10 15Gly Asp His His Arg
Gln Pro Gln Pro His His Leu Pro Gln Pro Pro 20 25 30Pro Pro Pro Gln
Pro Pro Ala Thr Leu Gln Ala Arg Glu His Pro Val 35 40 45Tyr Pro Pro
Glu Leu Ser Leu Leu Asp Ser Thr Asp Pro Arg Ala Trp 50 55 60Leu Ala
Pro Thr Leu Gln Gly Ile Cys Thr Ala Arg Ala Ala Gln Tyr65 70 75
80Leu Leu His Ser Pro Glu Leu Gly Ala Ser Glu Ala Ala Ala Pro Arg
85 90 95Asp Glu Val Asp Gly Arg Gly Glu Leu Val Arg Arg Ser Ser Gly
Gly 100 105 110Ala Ser Ser Ser Lys Ser Pro Gly Pro Val Lys Val Arg
Glu Gln Leu 115 120 125Cys Lys Leu Lys Gly Gly Val Val Val Asp Glu
Leu Gly Cys Ser Arg 130 135 140Gln Arg Ala Pro Ser Ser Lys Gln Val
Asn Gly Val Gln Lys Gln Arg145 150 155 160Arg Leu Ala Ala Asn Ala
Arg Glu Arg Arg Arg Met His Gly Leu Asn 165 170 175His Ala Phe Asp
Gln Leu Arg Asn Val Ile Pro Ser Phe Asn Asn Asp 180 185 190Lys Lys
Leu Ser Lys Tyr Glu Thr Leu Gln Met Ala Gln Ile Tyr Ile 195 200
205Asn Ala Leu Ser Glu Leu Leu Gln Thr Pro Ser Gly Gly Glu Gln Pro
210 215 220Pro Pro Pro Pro Ala Ser Cys Lys Ser Asp His His His Leu
Arg Thr225 230 235 240Ala Ala Ser Tyr Glu Gly Gly Ala Gly Asn Ala
Thr Ala Ala Gly Ala 245 250 255Gln Gln Ala Ser Gly Gly Ser Gln Arg
Pro Thr Pro Pro Gly Ser Cys 260 265 270Arg Thr Arg Phe Ser Ala Pro
Ala Ser Ala Gly Gly Tyr Ser Val Gln 275 280 285Leu Asp Ala Leu His
Phe Ser Thr Phe Glu Asp Ser Ala Leu Thr Ala 290 295 300Met Met Ala
Gln Lys Asn Leu Ser Pro Ser Leu Pro Gly Ser Ile Leu305 310 315
320Gln Pro Val Gln Glu Glu Asn Ser Lys Thr Ser Pro Arg Ser His Arg
325 330 335Ser Asp Gly Glu Phe Ser Pro His Ser His Tyr Ser Asp Ser
Asp Glu 340 345 350Ala Ser32144DNAMus musculus 3tcgacccacg
cgtccgccca cgcgtccgga tctccgagtg agagggggag ggtcagagga 60ggaaggaaaa
aaaaatcaga ccttgcagaa gagactagga aggtttttgt tgttgttgtt
120cggggcttat ccccttcgtt gaactgggtt gccagcacct cctctaacac
ggcacctccg 180agccattgca gtgcgatgtc ccgcctgctg catgcagaag
agtgggctga ggtaaaagag 240ttgggggacc accatcgcca tccccagccg
caccacgtcc cgccgctgac gccacagcca 300cctgctaccc tgcaggcgag
agaccttccc gtctacccgg cagaactgtc cctcctggat 360agcaccgacc
cacgcgcctg gctgactccc actttgcagg gcctctgcac ggcacgcgcc
420gcccagtatc tgctgcattc tcccgagctg ggtgcctccg aggccgcggc
gccccgggac 480gaggctgaca gccagggtga gctggtaagg agaagcggct
gtggcggcct cagcaagagc 540cccgggcccg tcaaagtacg ggaacagctg
tgcaagctga agggtggggt tgtagtggac 600gagcttggct gcagccgcca
gcgagcccct tccagcaaac aggtgaatgg ggtacagaag 660caaaggaggc
tggcagcaaa cgcaagggaa cggcgcagga tgcacgggct gaaccacgcc
720ttcgaccagc tgcgcaacgt tatcccgtcc ttcaacaacg acaagaagct
gtccaaatat 780gagaccctac agatggccca gatctacatc aacgctctgt
cggagttgct gcagactccc 840aatgtcggag agcaaccgcc gccgcccaca
gcttcctgca aaaatgacca ccatcacctt 900cgcaccgcct cctcctatga
aggaggtgcg ggcgcctctg cggtagctgg ggctcagcca 960gccccgggag
ggggcccgag acctaccccg cccgggcctt gccggactcg cttctcaggc
1020ccagcttcct ctgggggtta ctcggtgcag ctggacgctt tgcacttccc
agccttcgag 1080gacagggccc taacagcgat gatggcacag aaggacctgt
cgccttcgct gcccgggggc 1140atcctgcagc ctgtacagga ggacaacagc
aaaacatctc ccagatccca cagaagtgac 1200ggagagtttt ccccccactc
tcattacagt gactctgatg aggccagtta ggaaggcaac 1260agctccctga
aaactgagac aaccaaatgc ccttcctagc gcgcgggaag ccccgtgaca
1320aatatccctg caccctttaa tttttggtct gtggtgatcg ttgttagcaa
cgacttgact 1380tcggacggct gcagctcttc caatcccctt cctcctacct
tctccttcct ctgtatgtag 1440atactgtatc attatatgta cctttacgtg
gcatcgtttc atggtccatg ctgccaatat 1500gctgctaaaa tgtcgtatct
ctgcctctgg tctgggtttc acttatttta taccttggga 1560gttcatcctt
gcgtgttgcg ctcactcaca aataagggag ttagtcaatg aagttgtttc
1620cccaactgct tgagacccgc attgggtact ttactgaaca cggactattg
tgttgttaaa 1680atgcaggggc agataagagt atctgtagag cttagacacc
aagtgtgtcc agcagtgtgt 1740ctagcggacc cagaatacac gcacttcatc
actggccgct gcgccgcctt gaagaaactc 1800aactgccaat gcagagcaac
ttttgatttt aaaaacagcc actcataatc attaaactct 1860ttgcaaatgt
ttgtttttgc aaatgaaaat taaaaaaaaa catgtagtgt caaaggcatt
1920tggtcaattt tattttgctt tgttaacatt agaaaagtta tttattattg
cgtatttgga 1980cccatttcta cttaattgcc ttttttttac attttctact
cgagatcgtt ttattttgat 2040ttagcaaatc cagttgccat tgctttatgt
atgtatgctc ttttacaaat gataaaataa 2100actcggaaaa aaaaaaaaaa
aaaaaaaaaa aaaaaaaaaa aaaa 21444235PRTMus musculus 4Met Ser Arg Leu
Leu His Ala Glu Glu Trp Ala Glu Val Lys Glu Leu1 5 10 15Gly Asp His
His Arg His Pro Gln Pro His His Val Pro Pro Leu Thr 20 25 30Pro Gln
Pro Pro Ala Thr Leu Gln Ala Arg Asp Leu Leu Val Arg Arg 35 40 45Ser
Gly Cys Gly Gly Leu Ser Lys Ser Pro Gly Pro Val Lys Val Arg 50 55
60Glu Gln Leu Cys Lys Leu Lys Gly Gly Val Val Val Asp Glu Leu Gly65
70 75 80Cys Ser Arg Gln Arg Ala Pro Ser Ser Lys Gln Val Asn Gly Val
Gln 85 90 95Lys Gln Arg Arg Leu Ala Ala Asn Ala Arg Glu Arg Arg Arg
Met His 100 105 110Gly Leu Asn His Ala Phe Asp Gln Leu Arg Asn Val
Ile Pro Ser Phe 115 120 125Asn Asn Asp Lys Lys Leu Ser Lys Tyr Glu
Thr Leu Gln Met Ala Gln 130 135 140Ile Tyr Ile Asn Ala Leu Ser Glu
Leu Leu Gln Thr Pro Asn Val Gly145 150 155 160Ala Ser Ser Gly Gly
Tyr Ser Val Gln Leu Asp Ala Leu His Phe Pro 165 170 175Ala Phe Glu
Asp Arg Ala Leu Thr Ala Met Met Ala Gln Lys Asp Leu 180 185 190Ser
Pro Ser Leu Pro Gly Gly Ile Leu Gln Pro Val Gln Glu Asp Asn 195 200
205Ser Lys Thr Ser Pro Arg Ser His Arg Ser Asp Gly Glu Phe Ser Pro
210 215 220His Ser His Tyr Ser Asp Ser Asp Glu Ala Ser225 230
2355717DNAGallus gallus 5atggccccag gaggtagcga gtgttgttgc
agtgatgccg cgcacatcac ttggaggcag 60tgggagtaca cgcacgagaa ccaactgtgc
gtggcaggaa ctgtcagcag gatgaggccc 120aggacgtggg tctgcaccgg
atctttgtgg gaccaggaag cgggaattac tttgatgggc 180ccccaaatac
ccaaagtgga tgaggcagga gtgatgaccc acccggcaag gtcgctttgc
240agcactgggg cacatccgtg tcccggggtg gtcgtgctgc ccacgggtgg
gatagggcag 300ccttcaaaga agctctccaa gtacgagacg ctgcagatgg
cgcaaatcta catcagcgcc 360ctcgccgagc ttctgcacgg gccgcccgcg
ccccccgagc cgcccgccaa ggccgagctc 420cgcggggccc ccttcgagcc
tcccccgccg ccccctcctc cgccgccccg cgcctcgccc 480cccgcgcccg
ccaggactcg cttccccccg gcggcggccg cgggcggttt cgcggcgctt
540ctcgagccgc tgcgcttccc ttctttcccg gcgcagaaag cgccttctcc
cgcgctgctc 600ctggggccgc ccgcgccgca gcagcccgag aggagcaaag
cgtcgccgcg ctctcaccgc 660agcgacgggg agttctcgcc gcgctcccac
tacagtgact cggacgaggc cagctag 7176238PRTGallus gallus 6Met Ala Pro
Gly Gly Ser Glu Cys Cys Cys Ser Asp Ala Ala His Ile1 5 10 15Thr Trp
Arg Gln Trp Glu Tyr Thr His Glu Asn Gln Leu Cys Val Ala 20 25 30Gly
Thr Val Ser Arg Met Arg Pro Arg Thr Trp Val Cys Thr Gly Ser 35 40
45Leu Trp Asp Gln Glu Ala Gly Ile Thr Leu Met Gly Pro Gln Ile Pro
50 55 60Lys Val Asp Glu Ala Gly Val Met Thr His Pro Ala Arg Ser Leu
Cys65 70 75 80Ser Thr Gly Ala His Pro Cys Pro Gly Val Val Val Leu
Pro Thr Gly 85 90 95Gly Ile Gly Gln Pro Ser Lys Lys Leu Ser Lys Tyr
Glu Thr Leu Gln 100 105 110Met Ala Gln Ile Tyr Ile Ser Ala Leu Ala
Glu Leu Leu His Gly Pro 115 120 125Pro Ala Pro Pro Glu Pro Pro Ala
Lys Ala Glu Leu Arg Gly Ala Pro 130 135 140Phe Glu Pro Pro Pro Pro
Pro Pro Pro Pro Pro Pro Arg Ala Ser Pro145 150 155 160Pro Ala Pro
Ala Arg Thr Arg Phe Pro Pro Ala Ala Ala Ala Gly Gly 165 170 175Phe
Ala Ala Leu Leu Glu Pro Leu Arg Phe Pro Ser Phe Pro Ala Gln 180 185
190Lys Ala Pro Ser Pro Ala Leu Leu Leu Gly Pro Pro Ala Pro Gln Gln
195 200 205Pro Glu Arg Ser Lys Ala Ser Pro Arg Ser His Arg Ser Asp
Gly Glu 210 215 220Phe Ser Pro Arg Ser His Tyr Ser Asp Ser Asp Glu
Ala Ser225 230 235722DNAArtificial SequencePrimer 7atggctggac
acctggcttc ag 22822DNAArtificial SequencePrimer 8ttaaccccaa
agctccaggt tc 22925DNAArtificial SequencePrimer 9ccatgaccta
tactcaggct tcagg 251026DNAArtificial SequencePrimer 10gaagctccat
atccctgggt ggaaag 261122DNAArtificial SequencePrimer 11cacccgggcc
tcaacgctca cg 221224DNAArtificial SequencePrimer 12tccccttctc
cagttcgcag tcca 241323DNAArtificial SequencePrimer 13ccaaagtggt
ggacaagatt gcc 231423DNAArtificial SequencePrimer 14ggataggaag
gacgctcaaa gac 231520DNAArtificial SequencePrimer 15agactttaac
caagggcggt 201620DNAArtificial SequencePrimer 16tagccaggtt
gcgaagaact 201724DNAArtificial SequencePrimer 17aacagagatt
ggaaggccgc tggc 241823DNAArtificial SequencePrimer 18cttcagaaag
gctgtcacag gag 231921DNAArtificial SequencePrimer 19atggagactg
acgcgcccca g 212020DNAArtificial SequencePrimer 20atcttcttcg
tccgagtgac 202118DNAArtificial SequencePrimer 21cctccgacgg caggagtc
182221DNAArtificial SequencePrimer 22accgtagccc tgacggagtt t
212322DNAArtificial SequencePrimer 23agatctacat caacgctctg tc
222422DNAArtificial SequencePrimer 24actggcctca tcagagtcac tg
222517DNAArtificial SequencePrimer 25tggtgtcgtc ggggaac
172620DNAArtificial SequencePrimer 26aaggccgacc tccaaacctc
202723DNAArtificial SequencePrimer 27acgggctgaa cgcggcgctg gac
232823DNAArtificial SequencePrimer 28tgaaagagaa gttgccattg atg
232919DNAArtificial SequencePrimer 29gccatgcgcc gagtttgtc
193018DNAArtificial SequencePrimer 30atggcgccta gatgatgc
183122DNAArtificial SequencePrimer 31cagcctgagt caccgcagcc tc
223223DNAArtificial SequencePrimer 32tgacctgtcg ctgccagggc gcg
233323DNAArtificial SequencePrimer 33ctccctctac atcgctctgt tcg
233425DNAArtificial SequencePrimer 34aagcacctgc tcctgctcgt ccacg
253521DNAArtificial SequencePrimer 35ggccaacccc aaaaagtcgt g
213622DNAArtificial SequencePrimer 36gagagctggg gtgcgtgtag ga
223724DNAArtificial SequencePrimer 37ggcggcgcag ctccacaacc agta
243824DNAArtificial SequencePrimer 38ttgccacagc ccgggaaagg acag
243919DNAArtificial SequencePrimer 39ttgccccttc cccttttat
194017DNAArtificial SequencePrimer 40cgcttgctcg ctctcgt
174117DNAArtificial SequencePrimer 41acccgcatcc cagtcat
174218DNAArtificial SequencePrimer 42tcccggtgta caaagtgc
184322DNAArtificial SequencePrimer 43ctggagcgga tggacgccag ac
224421DNAArtificial SequencePrimer 44cgtctgctcc acagtgccag c
214520DNAArtificial SequencePrimer 45gtccttccca catgggagtt
204620DNAArtificial SequencePrimer 46gtttccacct tgacctcggt
204720DNAArtificial SequencePrimer 47agagatgtgg gatgcaggac
204820DNAArtificial SequencePrimer 48cacacaggga acttcaccct
204918DNAArtificial SequencePrimer 49gtcgtgggcc ttgtggcc
185019DNAArtificial SequencePrimer 50ctgtgagttc acactgggg
195118DNAArtificial SequencePrimer 51gccaaccgtg tgctgctg
185219DNAArtificial SequencePrimer 52acgtcgagac tgggcaagg
195317DNAArtificial SequencePrimer 53ccagcacagc ggtccag
175421DNAArtificial SequencePrimer 54ggggcacggt tgtccttgta g
215522DNAArtificial SequencePrimer 55cggaaccaga gataggacct ac
225622DNAArtificial SequencePrimer 56ctaagcacgg gttgcctcat cc
225718DNAArtificial SequencePrimer 57aacgggaagc ccatcacc
185818DNAArtificial SequencePrimer 58cagccttggc agcaccag
18592013DNAArtificial SequenceSynthetic construct 59atgtccaatt
tactgaccgt acaccaaaat ttgcctgcat taccggtcga tgcaacgagt 60gatgaggttc
gcaagaacct gatggacatg ttcagggatc gccaggcgtt ttctgagcat
120acctggaaaa tgcttctgtc cgtttgccgg tcgtgggcgg catggtgcaa
gttgaataac 180cggaaatggt ttcccgcaga acctgaagat gttcgcgatt
atcttctata tcttcaggcg 240cgcggtctgg cagtaaaaac tatccagcaa
catttgggcc agctaaacat gcttcatcgt 300cggtccgggc tgccacgacc
aagtgacagc aatgctgttt cactggttat gcggcggatc 360cgaaaagaaa
acgttgatgc cggtgaacgt gcaaaacagg ctctagcgtt cgaacgcact
420gatttcgacc aggttcgttc actcatggaa aatagcgatc gctgccagga
tatacgtaat 480ctggcatttc tggggattgc ttataacacc ctgttacgta
tagccgaaat tgccaggatc 540agggttaaag atatctcacg tactgacggt
gggagaatgt taatccatat tggcagaacg 600aaaacgctgg ttagcaccgc
aggtgtagag aaggcactta gcctgggggt aactaaactg 660gtcgagcgat
ggatttccgt ctctggtgta gctgatgatc cgaataacta cctgttttgc
720cgggtcagaa aaaatggtgt tgccgcgcca tctgccacca gccagctatc
aactcgcgcc 780ctggaaggga tttttgaagc aactcatcga ttgatttacg
gcgctaagga tgactctggt 840cagagatacc tggcctggtc tggacacagt
gcccgtgtcg gagccgcgcg agatatggcc 900cgcgctggag tttcaatacc
ggagatcatg caagctggtg gctggaccaa tgtaaatatt 960gtcatgaact
atatccgtaa cctggatagt gaaacagggg caatggtgcg cctgctggaa
1020gatggggatc aggctggtgc catgggcgat ccacgaaatg aaatgggtgc
ttcaggagac 1080atgagggctg ccaacctttg gccaagccct cttgtgatta
agcacactaa gaagaatagc 1140cctgccttgt ccttgacagc tgaccagatg
gtcagtgcct tgttggatgc tgaaccgccc 1200atgatctatt ctgaatatga
tccttctaga cccttcagtg
aagcctcaat gatgggctta 1260ttgaccaacc tagcagatag ggagctggtt
catatgatca actgggcaaa gagagtgcca 1320ggctttgggg acttgaatct
ccatgatcag gtccaccttc tcgagtgtgc ctggctggag 1380attctgatga
ttggtctcgt ctggcgctcc atggaacacc cggggaagct cctgtttgct
1440cctaacttgc tcctggacag gaatcaaggt aaatgtgtgg aaggcatggt
ggagatcttt 1500gacatgttgc ttgctacgtc aagtcggttc cgcatgatga
acctgcaggg tgaagagttt 1560gtgtgcctca aatccatcat tttgcttaat
tccggagtgt acacgtttct gtccagcacc 1620ttgaagtctc tggaagagaa
ggaccacatc caccgtgtcc tggacaagat cacagacact 1680ttgatccacc
tgatggccaa agctggcctg actctgcagc agcagcatcg ccgcctagct
1740cagctccttc tcattctttc ccatatccgg cacatgagta acaaacgcat
ggagcatctc 1800tacaacatga aatgcaagaa cgtggtaccc ctctatgacc
tgctcctgga gatgttggat 1860gcccaccgcc ttcatgcccc agccagtcgc
atgggagtgc ccccagagga gcccagccag 1920acccagctgg ccaccaccag
ctccacttca gcacattcct tacaaaccta ctacataccc 1980ccggaagcag
agggcttccc caacacgatc tga 2013
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.