Antibody Constant Region Variant
Igawa; Tomoyuki ; et al.
U.S. patent application number 16/298032 was filed with the patent office on 2019-07-11 for antibody constant region variant. This patent application is currently assigned to Chugai Seiyaku Kabushiki Kaisha. The applicant listed for this patent is Chugai Seiyaku Kabushiki Kaisha. Invention is credited to Tomoyuki Igawa, Taichi Kuramochi, Atsuhiko Maeda, Hirotake Shiraiwa.
| Application Number | 20190211081 16/298032 |
| Document ID | / |
| Family ID | 42739765 |
| Filed Date | 2019-07-11 |

View All Diagrams
| United States Patent Application | 20190211081 |
| Kind Code | A1 |
| Igawa; Tomoyuki ; et al. | July 11, 2019 |
ANTIBODY CONSTANT REGION VARIANT
Abstract
By altering amino acid sequences, the present inventors successfully produced constant regions that can confer antibodies with particularly favorable properties for pharmaceutical agents. When used to produce antibodies, the altered constant regions produced according to the present invention significantly reduce heterogeneity. Specifically, the antibody homogeneity can be achieved by using antibody heavy chain and light chain constant regions introduced with alterations provided by the present invention. More specifically, the alterations can prevent the loss of homogeneity of antibody molecules due to disulfide bond differences in the heavy chain. Furthermore, in a preferred embodiment, the present invention can improve antibody pharmacokinetics as well as prevent the loss of homogeneity due to C-terminal deletion in antibody constant region.
| Inventors: | Igawa; Tomoyuki; (Shizuoka, JP) ; Kuramochi; Taichi; (Shizuoka, JP) ; Maeda; Atsuhiko; (Shizuoka, JP) ; Shiraiwa; Hirotake; (Shizuoka, JP) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | Chugai Seiyaku Kabushiki
Kaisha Tokyo JP |
||||||||||
| Family ID: | 42739765 | ||||||||||
| Appl. No.: | 16/298032 | ||||||||||
| Filed: | March 11, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14680250 | Apr 7, 2015 | 10253091 | ||
| 16298032 | ||||
| 13257145 | Nov 22, 2011 | |||
| PCT/JP2010/054767 | Mar 19, 2010 | |||
| 14680250 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 16/2875 20130101; C07K 2317/522 20130101; C07K 2317/94 20130101; C07K 16/2866 20130101; C07K 2317/71 20130101; C07K 2317/72 20130101; C07K 2317/92 20130101; C07K 16/00 20130101; C07K 2317/526 20130101; C07K 2317/515 20130101; C07K 2317/52 20130101; C07K 2317/53 20130101; C07K 2317/528 20130101; C07K 2317/76 20130101; C07K 2317/524 20130101 |
| International Class: | C07K 16/00 20060101 C07K016/00; C07K 16/28 20060101 C07K016/28 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Mar 19, 2009 | JP | 2009-068631 |
| Sep 16, 2009 | JP | 2009-213901 |
Claims
1-21. (canceled)
22. A human .lamda. chain constant region variant comprising the amino acid sequence of SEQ ID NO: 37 in which either (a) at least one amino acid at any of positions 99 to 103 in the amino acid sequence of SEQ ID NO: 37 is deleted, and optionally any other amino acid at any of positions 1 to 103 of SEQ ID NO: 37 is substituted with another amino acid, wherein the total number of deleted amino acids is one to five and the combined total number of deleted amino acids and substituted amino acids is 20 or fewer; or (b) Cys at position 104 of SEQ ID NO: 37 is deleted or substituted with another amino acid, and optionally at least one amino acid at any of positions 1 to 103 of SEQ ID NO: 37 is deleted or substituted with another amino acid, provided that at least one amino acid at any of positions 99 to 103 of SEQ ID NO: 37 is substituted with Cys, wherein the total number of deleted amino acids is up to five and the combined total number of deleted amino acids and substituted amino acids is 20 or fewer.
23. The human .lamda. chain constant region variant of claim 22, wherein the amino acid at position 102 in the amino acid sequence of SEQ ID NO: 37 is deleted.
24. The human .lamda. chain constant region variant of claim 22, wherein the amino acid at position 103 in the amino acid sequence of SEQ ID NO: 37 is deleted.
25. A human .lamda. chain constant region variant comprising the amino acid sequence of SEQ ID NO: 37 in which at least one amino acid at any of positions 99 to 103 is substituted with Cys, and Cys at position 104 of SEQ ID NO: 37 is deleted or substituted with another amino acid.
26. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 22.
27. A pharmaceutical composition comprising the antibody of claim 26.
28. The human .lamda. chain constant region variant of claim 22, comprising the amino acid sequence of SEQ ID NO: 37 with Cys at position 104 deleted and a Cys substituted at one or more of positions 99 to 103.
29. The human .lamda. chain constant region variant of claim 22, wherein only one of positions 99 to 103 of SEQ ID NO: 37 is a Cys.
30. The human .lamda. chain constant region variant of claim 28, wherein only one of positions 99 to 103 of SEQ ID NO: 37 is a Cys.
31. The human .lamda. chain constant region variant of claim 22, wherein the amino acids at two of positions 99 to 103 of SEQ ID NO: 37 are substituted with Cys.
32. The human .lamda. chain constant region variant of claim 28, wherein the amino acids at two of positions 99 to 103 of SEQ ID NO: 37 are substituted with Cys.
33. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 23.
34. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 24.
35. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 25.
36. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 28.
37. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 29.
38. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 30.
39. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 31.
40. An antibody comprising a light chain comprising the human .lamda. chain constant region variant of claim 32.
41. The antibody of claim 26, wherein disulfide bond pattern heterogeneity of the human .lamda. chain constant region variant in the antibody is reduced compared to disulfide bond pattern heterogeneity of a reference human .lamda. chain constant region in a reference antibody that is identical to the antibody of claim 26 except that the reference human .lamda. chain constant regions in the reference antibody each consists of the amino acid sequence of SEQ ID NO: 37.
42. The human .lamda. chain constant region variant of claim 22, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
43. The human .lamda. chain constant region variant of claim 23, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
44. The human .lamda. chain constant region variant of claim 24, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
45. The antibody of claim 26, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
46. The pharmaceutical composition of claim 27, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
47. The human .lamda. chain constant region variant of claim 29, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
48. The human .lamda. chain constant region variant of claim 31, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
49. The antibody of claim 33, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
50. The antibody of claim 34, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
51. The antibody of claim 37, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
52. The antibody of claim 39, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
53. The antibody of claim 41, wherein the human .lamda. chain constant region variant comprises the amino acid sequence of SEQ ID NO: 37 except at one or more positions within positions 99 to 103 of SEQ ID NO: 37.
54. The human .lamda. chain constant region variant of claim 22, wherein the combined total number of deleted or substituted amino acids is 10 or fewer.
55. A human .lamda. chain constant region variant that is 99-104 amino acid residues in length, comprises at least one Cys residue at a position or positions selected from positions 99-103, and does not comprise a Cys residue at position 104, wherein the variant comprises SEQ ID NO: 37 with a combined total of up to 20 substitutions or deletions.
56. The variant of claim 55, wherein the variant comprises SEQ ID NO: 37 with a combined total of up to 10 substitutions or deletions.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a divisional application of U.S. application Ser. No. 14/680,250, filed on Apr. 7, 2015, which is a divisional of U.S. application Ser. No. 13/257,145, having a 371 (c) date of Nov. 22, 2011, which is the National Stage of International Application Serial No. PCT/JP2010/054767, filed on Mar. 19, 2010, which claims priority to Japanese Application Serial Nos. 2009-068631 and 2009-213901, filed on Mar. 19, 2009, and Sep. 16, 2009, respectively.
TECHNICAL FIELD
[0002] The present invention relates to antibody constant regions with an altered amino acid sequence, and antibodies comprising these constant regions.
BACKGROUND
[0003] Antibodies are drawing attention as pharmaceuticals as they are highly stable in plasma (blood) and have few side effects. Of these, a number of IgG-type antibody pharmaceuticals are available on the market and many antibody pharmaceuticals are currently under development (Non-patent Documents 1 and 2).
[0004] Almost all antibody pharmaceuticals currently available on the market are of the IgG1 subclass. IgG1 type antibodies are expected to be useful as anti-cancer antibody pharmaceuticals since they can bind to Fc.gamma. receptor and exert ADCC activity. However, when it comes to antibody pharmaceuticals intended for neutralizing biological activity of an antigen, the binding of the Fc domain to Fc.gamma. receptor, which is important for effector functions such as ADCC, can cause unnecessary side effects, and thus it is preferable to eliminate such binding activity (Non-patent Document 3). Furthermore, since Fc.gamma. receptor is expressed in antigen-presenting cells, molecules that bind to Fc.gamma. receptor tend to be presented as antigens. It has been reported that immunogenicity is and can be enhanced by linking a protein or peptide to the Fc domain of IgG1 (Non-patent Document 4 and Patent Document 1). Interaction between the antibody Fc domain and Fc.gamma. receptor is thought to be a cause of the serious side effects encountered in phase-I clinical trials of TGN1412 (Non-patent Document 5). Thus, binding to Fc.gamma. receptor is considered unfavorable in antibody pharmaceuticals intended for neutralizing the biological activity of an antigen from the perspective of side effects and immunogenicity.
[0005] A method for impairing the binding to Fc.gamma. receptor is to alter the subtype of the IgG antibody from IgG1 to IgG2 or IgG4; however, this method cannot completely inhibit the binding (Non-patent Document 6). One of the methods reported for completely inhibiting the binding to Fc.gamma. receptor is to artificially alter the Fc domain. For example, the effector functions of anti-CD3 antibodies and anti-CD4 antibodies cause side effects. Thus, amino acids that are not present in the wild type sequence were introduced into the Fc.gamma.-receptor-binding domain of Fc (Non-patent Documents 3 and 7), and clinical trials are currently being conducted to assess anti-CD3 antibodies and anti-CD4 antibodies that have a mutated Fc domain and do not bind to Fc.gamma. receptor (Non-patent Documents 5 and 8). Alternatively, Fc.gamma. receptor-nonbinding antibodies can be prepared by altering the Fc.gamma.R-binding domain of IgG1 (positions 233, 234, 235, 236, 327, 330, and 331 in the EU numbering; hereinafter abbreviated as position X (EU numbering)) to an IgG2 or IgG4 sequence (Non-patent Document 9 and Patent Document 2). However, these molecules contain new non-native peptide sequences of nine to twelve amino acids, which may constitute a T-cell epitope peptide and thus pose an immunogenicity risk. There is no previous report on Fc.gamma. receptor-nonbinding antibodies that have overcome these problems.
[0006] Furthermore, for heterogeneity of the C-terminal sequence of an antibody, deletion of C-terminal amino acid lysine residue, and amidation of the C-terminal amino group due to deletion of both of the two C-terminal amino acids, glycine and lysine, have been reported (Non-patent Document 12). It is preferable to eliminate such heterogeneity when developing antibodies into pharmaceuticals.
[0007] Furthermore, in general, it is necessary that subcutaneous formulations are high-concentration formulations. From the perspective of stability and such, the concentration limit of IgG-type antibody formulations is generally thought to be about 100 mg/ml (Non-patent Document 13). Thus, it was a challenge to secure stability at high concentrations. However, there has been no report published on the improvement of the stability of IgG at high concentrations by introducing amino acid substitutions into its constant region. Meanwhile, instead of increasing the antibody concentration, methods that reduce the antibody dose by improving antibody kinetics in blood can be thought. A method for prolonging the antibody half-life in plasma has been reported and it substitutes amino acids in the constant region (Non-patent Documents 14 and 15); however, introduction of non-native sequences into the constant region is unpreferable from the perspective of immunogenicity risk.
[0008] Furthermore, physicochemical properties of antibody proteins, in particular, homogeneity, are very crucial in the development of antibody pharmaceuticals. For the IgG2 subtype, heterogeneity caused by disulfide bonds in the hinge region has been reported (Non-patent Documents 10, 16, 17, and 18 and Patent Document 3). It is not easy to manufacture them as a pharmaceutical in a large scale while maintaining differences of objective substance/related substance-related heterogeneity between productions. Thus, single substances are desirable as much as possible for antibody molecules developed as pharmaceuticals. In the present invention, differences in heterogeneity among productions can be understood, for example, as differences in heterogeneity among production lots. Heterogeneity in the production lots can be evaluated quantitatively by determining the diversity of molecular weight and structure of the produced antibody molecules.
[0009] As described above, it is desirable that the constant region sequences of antibody pharmaceuticals that are intended for neutralizing an antigen meet all the requirements in terms of the stability, C-terminal heterogeneity, immunogenicity (antigenicity), blood pharmacokinetics, and heterogeneity of hinge region. In particular, constant regions that do not have the heterogeneity of hinge region, which are more superior in blood pharmacokinetics than the natural constant regions such as of IgG1, are expected to be very useful as a constant region of antibody pharmaceuticals. However, altered constant regions that meet all of the above requirements have not yet been reported. Thus, there is a demand for antibody constant regions that have overcome the problems described above.
[0010] Documents of related prior arts for the present invention are described below.
PRIOR ART DOCUMENTS
Non-Patent Documents
[0011] [Non-patent Document 1] Monoclonal antibody successes in the clinic, Janice M Reichert, Clark J Rosensweig, Laura B Faden & Matthew C Dewitz, Nature Biotechnology 23, 1073-1078 (2005) [0012] [Non-patent Document 2] Pavlou A K, Belsey M J., The therapeutic antibodies market to 2008., Eur J Pharm Biopharm. 2005 April; 59(3): 389-96 [0013] [Non-patent Document 3] Reddy M P, Kinney C A, Chaikin M A, Payne A, Fishman-Lobell J, Tsui P, Dal Monte P R, Doyle M L, Brigham-Burke M R, Anderson D, Reff M, Newman R, Hanna N, Sweet R W, Truneh A. Elimination of Fc receptor-dependent effector functions of a modified IgG4 monoclonal antibody to human CD4. J Immunol. 2000 Feb. 15; 164(4): 1925-33 [0014] [Non-patent Document 4] Guyre P M, Graziano R F, Goldstein J, Wallace P K, Morganelli P M, Wardwell K, Howell A L. Increased potency of Fc-receptor-targeted antigens. Cancer Immunol Immunother. 1997 November-December; 45(3-4): 146-8 [0015] [Non-patent Document 5] Strand V, Kimberly R, Isaacs J D. Biologic therapies in rheumatology: lessons learned, future directions. Nat Rev Drug Discov. 2007 January; 6(1): 75-92 [0016] [Non-patent Document 6] Gessner J E, Heiken H, Tamm A, Schmidt R E. The IgG Fc receptor family. Ann Hematol. 1998 June; 76(6): 231-48 [0017] [Non-patent Document 7] Cole M S, Anasetti C, Tso J Y. Human IgG2 variants of chimeric anti-CD3 are nonmitogenic to T cells. J Immunol. 1997 Oct. 1; 159(7): 3613-21 [0018] [Non-patent Document 8] Chau L A, Tso J Y, Melrose J, Madrenas J. HuM291 (Nuvion), a humanized Fc receptor-nonbinding antibody against CD3, anergizes peripheral blood T cells as partial agonist of the T cell receptor. Transplantation. 2001 Apr. 15; 71(7): 941-50 [0019] [Non-patent Document 9] Armour K L, Clark M R, Hadley A G, Williamson L M., Recombinant human IgG molecules lacking Fcgamma receptor I binding and monocyte triggering activities. Eur J Immunol. 1999 August; 29(8): 2613-24 [0020] [Non-patent Document 10] Chu G C, Chelius D, Xiao G, Khor H K, Coulibaly S, Bondarenko P V. Accumulation of Succinimide in a Recombinant Monoclonal Antibody in Mildly Acidic Buffers Under Elevated Temperatures. Pharm Res. 2007 Mar. 24; 24(6): 1145-56 [0021] [Non-patent Document 11] A. J. Cordoba, B. J. Shyong, D. Breen, R. J. Harris, Nonenzymatic hinge region fragmentation of antibodies in solution, J. Chromatogr., B, Anal. Technol. Biomed. Life Sci. (2005) 818: 115-121 [0022] [Non-patent Document 12] Johnson K A, Paisley-Flango K, Tangarone B S, Porter T J, Rouse J C. Cation exchange-HPLC and mass spectrometry reveal C-terminal amidation of an IgG1 heavy chain. Anal Biochem. 2007 Jan. 1; 360(1): 75-83 [0023] [Non-patent Document 13] Shire S J, Shahrokh Z, Liu J. Challenges in the development of high protein concentration formulations., J Pharm Sci. 2004 June; 93(6): 1390-402 [0024] [Non-patent Document 14] Hinton P R, Xiong J M, Johlfs M G, Tang M T, Keller S, Tsurushita N., An engineered human IgG1 antibody with longer serum half-life., J Immunol. 2006 Jan. 1; 176(1): 346-56 [0025] [Non-patent Document 15] Ghetie V, Popov S, Borvak J, Radu C, Matesoi D, Medesan C, Ober R J, Ward E S., Increasing the serum persistence of an IgG fragment by random mutagenesis., Nat Biotechnol. 1997 July; 15(7): 637-40 [0026] [Non-patent Document 16] [0027] Wypych J, Li M, Guo A, Zhang Z, Martinez T, Allen M J, Fodor S, Kelner D N, Flynn G C, Liu Y D, Bondarenko P V, Ricci M S, Dillon T M, Balland A., Human IgG2 antibodies display disulfide-mediated structural isoforms., J Biol Chem. 2008 Jun. 6; 283(23): 16194-205 [0028] [Non-patent Document 17] [0029] Dillon T M, Ricci M S, Vezina C, Flynn G C, Liu Y D, Rehder D S, Plant M, Henkle B, Li Y, Deechongkit S, Varnum B, Wypych J, Balland A, Bondarenko P V., Structural and functional characterization of disulfide isoforms of the human IgG2 subclass., J Biol Chem. 2008 Jun. 6; 283(23): 16206-15 [0030] [Non-patent Document 18] [0031] Martinez T, Guo A, Allen M J, Han M, Pace D, Jones J, Gillespie R, Ketchem R R, Zhang Y, [0032] Balland A., Disulfide connectivity of human immunoglobulin G2 structural isoforms., Biochemistry. 2008 Jul. 15; 47(28): 7496-508
Patent Documents
[0032] [0033] [Patent Document 1] US 20050261229 A1 [0034] [Patent Document 2] WO 99/58572 [0035] [Patent Document 3] US 2006/0194280
SUMMARY
Problems to be Solved by the Invention
[0036] The present invention was achieved in view of the above circumstances. An objective of the present invention is to provide constant regions that can confer antibodies with properties preferable for pharmaceuticals by altering amino acids in the antibody constant regions, and antibodies comprising these constant regions and variable regions.
Means for Solving the Problems
[0037] The present inventors conducted dedicated studies to generate antibody constant regions that have been improved by altering their amino acid sequences, which have improved homogeneity (C-terminal and hinge region), immunogenicity, stability, and pharmacokinetics. As a result, the present inventors successfully produced antibody constant regions with improved heterogeneity, immunogenicity, and stability. The present inventors also successfully produced antibody constant regions with a reduced Fc.gamma. receptor-binding activity by further altering amino acids in the constant regions described above. The resulting antibody constant regions are excellent constant regions that are superior to the native IgG1 constant region in pharmacokinetics and have an improved heterogeneity of hinge region.
[0038] The present invention relates to antibody constant regions that are superior in terms of safety, immunogenicity risk, physicochemical properties (stability and homogeneity), and more superior in terms of pharmacokinetics and heterogeneity of hinge region through improvement by amino acid alterations; antibodies comprising such antibody constant region; pharmaceutical compositions comprising such antibody; and methods for producing them. More specifically, the present invention provides:
[1] an antibody constant region comprising an amino acid sequence in which Cys at position 14 (position 131 in the EU numbering), Arg at position 16 (position 133 in the EU numbering), Cys at position 103 (position 220 in the EU numbering), Glu at position 20 (position 137 in the EU numbering), Ser at position 21 (position 138 in the EU numbering), His at position 147 (position 268 in the EU numbering), Arg at position 234 (position 355 in the EU numbering), and Gln at position 298 (position 419 in the EU numbering) in the amino acid sequence of SEQ ID NO: 24 (IgG2 constant region) are substituted with other amino acids; [2] the antibody constant region of [1], wherein Ser is substituted for Cys at position 14, Lys is substituted for Arg at position 16, Ser is substituted for Cys at position 103, Gly is substituted for Glu at position 20, Gly is substituted for Ser at position 21, Gln is substituted for His at position 147, Gln is substituted for Arg at position 234, and Glu is substituted for Gln at position 298; [3] the antibody constant region of [1] or [2], which comprises an amino acid sequence additionally having deletion of Gly at position 325 (position 446 in the EU numbering) and Lys at position 326 (position 447 in the EU numbering); [4] an antibody constant region comprising an amino acid sequence in which Cys at position 14 (position 131 in the EU numbering), Arg at position 16 (position 133 in the EU numbering), Cys at position 103 (position 220 in the EU numbering), Glu at position 20 (position 137 in the EU numbering), Ser at position 21 (position 138 in the EU numbering), His at position 147 (position 268 in the EU numbering), Arg at position 234 (position 355 in the EU numbering), Gln at position 298 (position 419 in the EU numbering), Ala at position 209 (position 330 in the EU numbering), Pro at position 210 (position 331 in the EU numbering), and Thr at position 218 (position 339 in the EU numbering) in the amino acid sequence of SEQ ID NO: 24 (IgG2 constant region) are substituted with other amino acids; [5] the antibody constant region of [4], wherein Ser is substituted for Cys at position 14, Lys is substituted for Arg at position 16, Ser is substituted for Cys at position 103, Gly is substituted for Glu at position 20, Gly is substituted for Ser at position 21, Gln is substituted for His at position 147, Gln is substituted for Arg at position 234, Glu is substituted for Gln at position 298, Ser is substituted for Ala at position 209, Ser is substituted for Pro at position 210, and Ala is substituted for Thr at position 218; [6] the antibody constant region of [4] or [5], which also comprises an amino acid sequence in which Gly at position 325 (position 446 in the EU numbering) and Lys at position 326 (position 447 in the EU numbering) are deleted; [7] an antibody comprising the constant region of any one of [1] to [6]; [8] a pharmaceutical composition comprising the antibody of [7]; [9] a human .kappa. chain constant region comprising at least one Cys at positions 102 to 106; [10] a human .kappa. chain constant region which does not comprise Cys at position 107; [11] a human .kappa. chain constant region which comprises at least one Cys at positions 102 to 106 but does not comprise Cys at position 107; [12] the human .kappa. chain constant region of any one of [9] to [11], in which at least one amino acid at positions 1 to 106 in the amino acid sequence of SEQ ID NO: 32 is deleted; [13] the human .kappa. chain constant region of [12], in which at least one amino acid at positions 102 to 106 is deleted; [14] the human .kappa. chain constant region of [13], in which the amino acid at position 105 is deleted; [15] the human .kappa. chain constant region of [13], in which the amino acid at position 106 is deleted; [16] the human .kappa. chain constant region of [9], in which at least one amino acid at position 102 to 106 is substituted with Cys; [17] the human .kappa. chain constant region of any one of [9] to [11], in which at least one amino acid at positions 102 to 106 is substituted with Cys, and Cys at position 107 is deleted or substituted with another amino acid; [18] an antibody comprising the human .kappa. chain constant region of any one of [9] to [17]; [19] a pharmaceutical composition comprising the antibody of [18]; [20] an antibody comprising the heavy chain constant region of any one of [1] to [6] and the light chain constant region of any one of [9] to [17]; and [21] a pharmaceutical composition comprising the antibody of [20].
Effects of the Invention
[0039] The present invention provides constant regions that can confer to antibodies properties desirable for pharmaceutical agents. By means of amino acid sequence alterations, the constant regions of the present invention can improve the following antibody properties to conditions favorable for pharmaceutical agents.
[0040] Decrease in Antibody Heterogeneity:
[0041] Polypeptides obtainable by expressing a DNA encoding a certain amino acid sequence should theoretically be homogeneous polypeptide molecules consisting of the same amino acid sequence. However, in practice, when a DNA encoding an antibody is expressed in suitable hosts, heterogeneous polypeptides with different structures may be formed due to various factors.
[0042] In the production of antibodies, an antibody population comprising many heterogeneous polypeptides can be referred to as having high heterogeneity. The constant regions of the present invention have the causes of heterogeneity removed by amino acid sequence alteration. Therefore, constructing antibodies using the constant regions of the present invention enables production of antibodies with low heterogeneity. Specifically, by introducing alterations provided by the present invention into heavy chain constant regions of antibodies, the homogeneity of the antibodies can be maintained at a high level. Suppressing the antibody heterogeneity to a low level means ameliorating the heterogeneity and this is an important objective in maintaining the quality of pharmaceuticals. Therefore, the constant regions of the present invention contribute to the maintenance of the quality of antibody-containing pharmaceuticals.
[0043] Improvement of Pharmacokinetics:
[0044] In a preferred embodiment, the present invention contributes to improvement of antibody pharmacokinetics. Specifically, when specific amino acid residues are altered in an antibody constant region of the present invention, blood concentration of the antibody composed of this constant region is maintained for a longer time than an antibody without amino acid sequence alterations. Maintaining blood concentration for as long a time as possible means that, when an antibody is administered as a pharmaceutical, its therapeutic effect can be maintained for a long time with a smaller amount of antibody. Alternatively, the antibody can be administered with wider intervals and smaller number of administrations.
BRIEF DESCRIPTION OF THE DRAWINGS
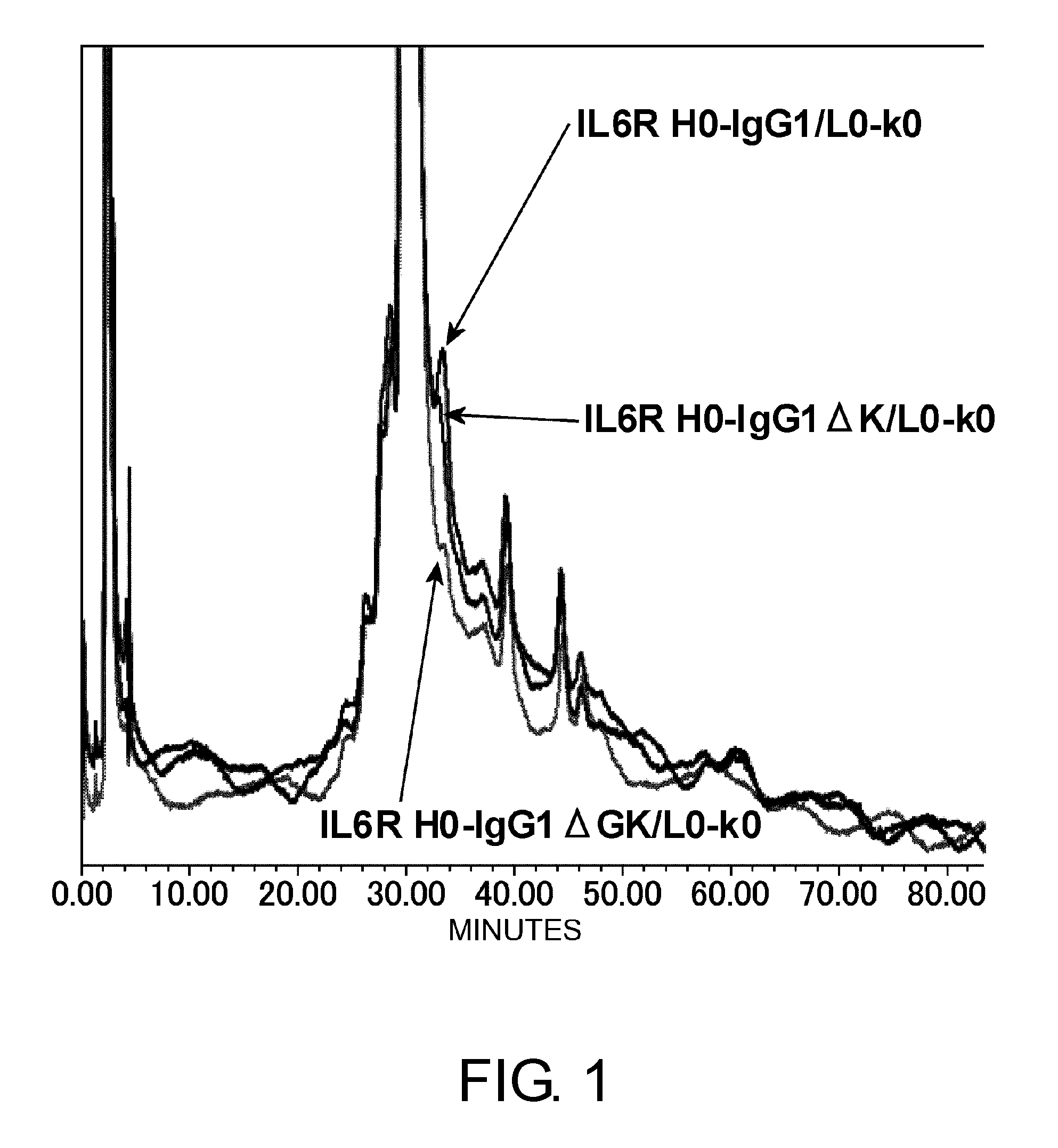
[0045] FIG. 1 shows in a graph the results of performing cation exchange chromatography on IL6R H0-IgG1/L0-k0, IL6R H0-IgG1.DELTA.K/L0-k0, and IL6R H0-IgG1.DELTA.GK/L0-k0 to evaluate the heterogeneity derived from the C terminus. In the figure, the vertical axis shows absorbance at 280 nm and the horizontal axis shows elution time (minutes).
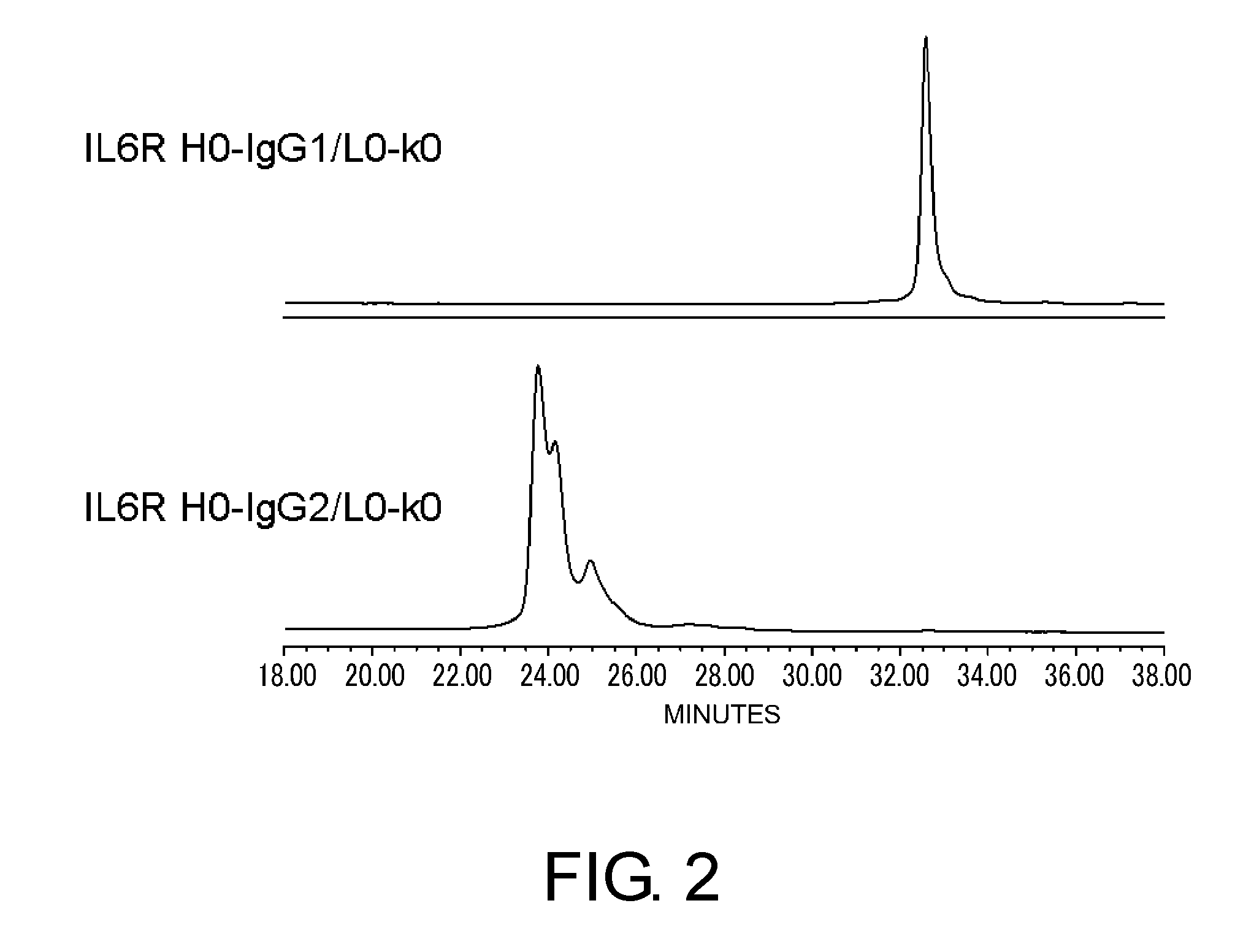
[0046] FIG. 2 shows in graphs the results of performing cation exchange chromatography on IL6R H0-IgG1/L0-k0 and IL6R H0-IgG2/L0-k0 to evaluate the heterogeneity derived from disulfide bonds. In the figure, the vertical axis shows absorbance at 280 nm and the horizontal axis shows elution time (minutes).
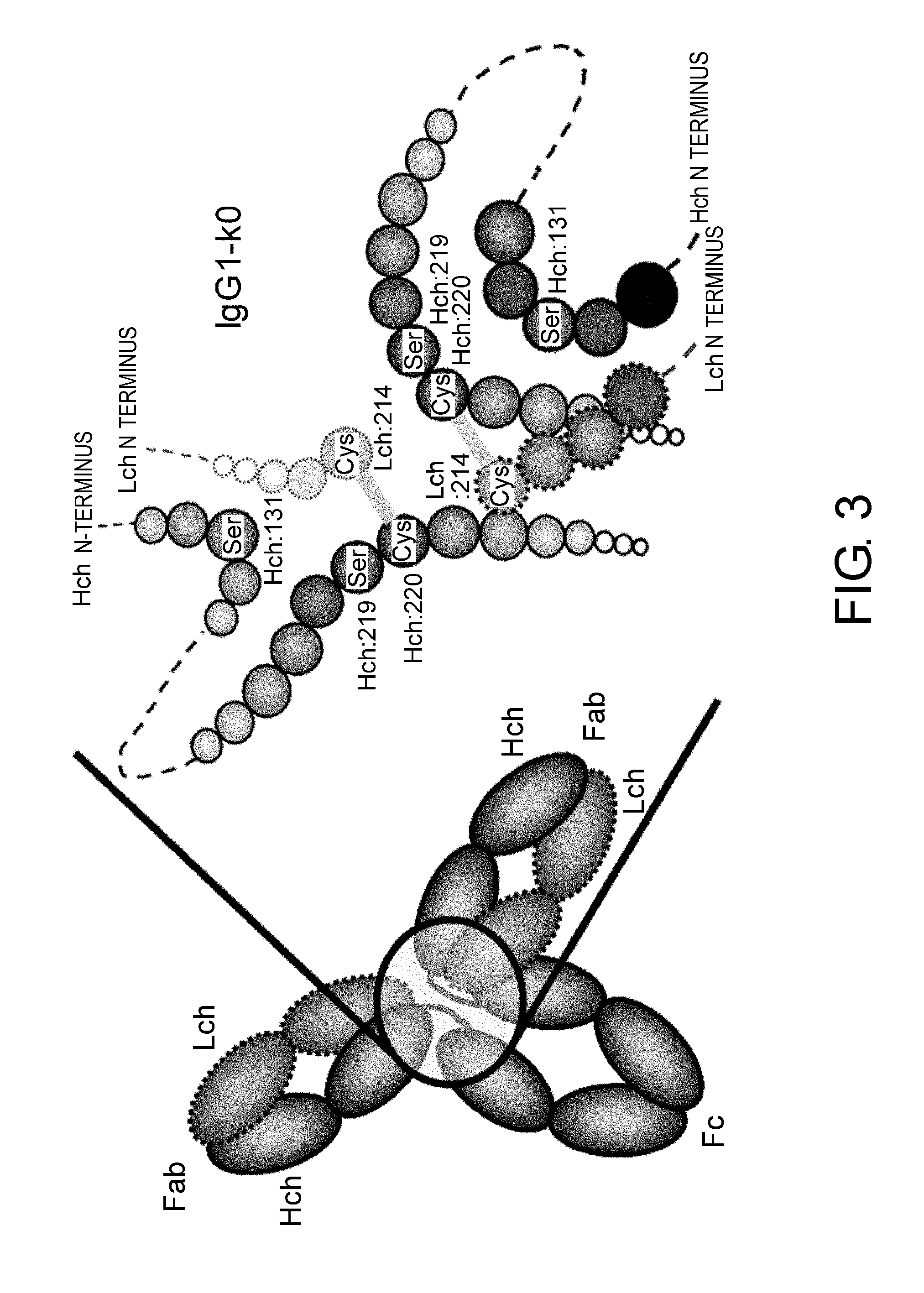
[0047] FIG. 3 shows in diagrams the IgG-type antibody and details of the structure around its hinge region (positioning of the heavy chain (H chain) and light chain (L chain) and disulfide bonds between them; detailed drawing represents IgG1-k0).
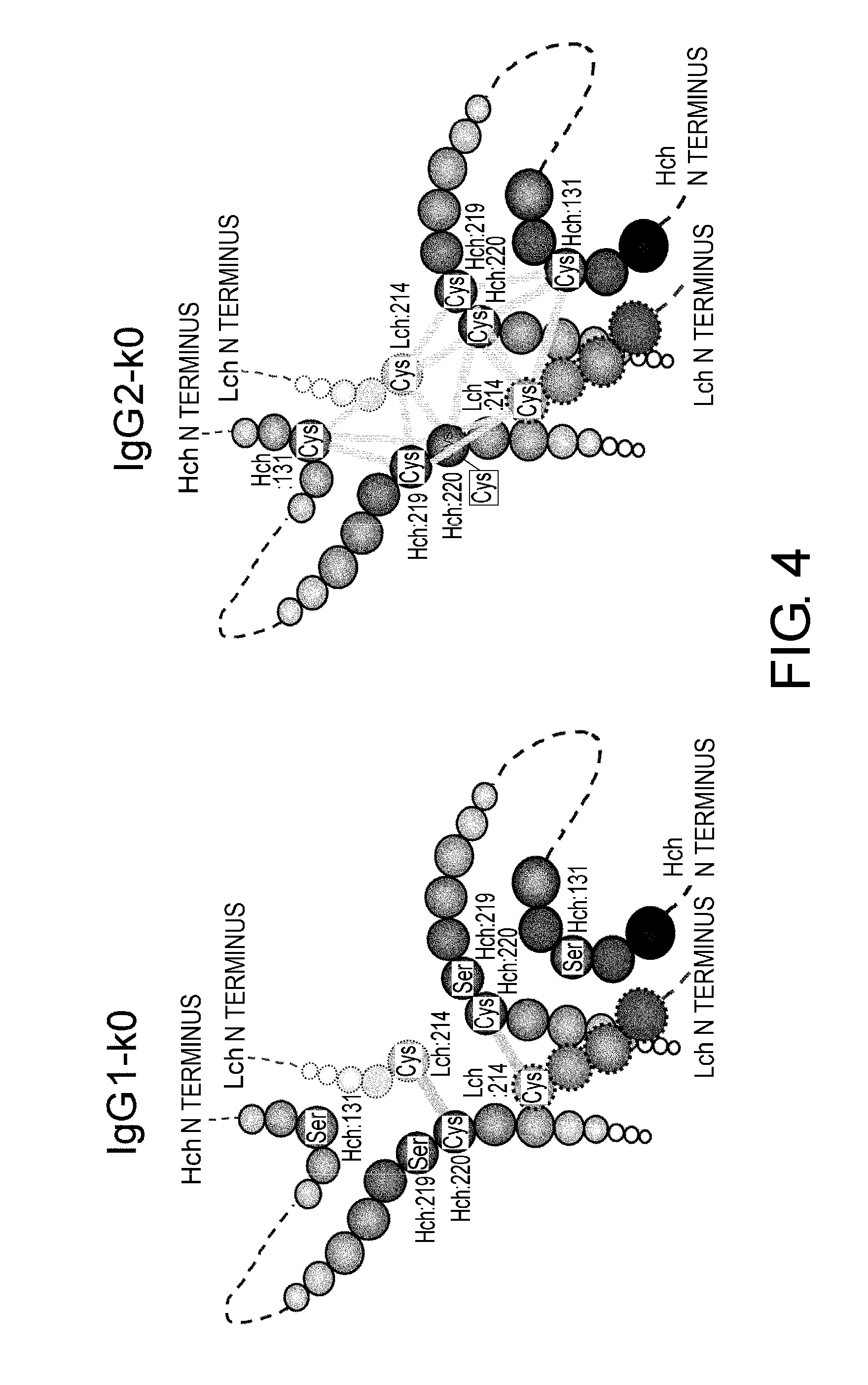
[0048] FIG. 4 shows in diagrams predicted disulfide bond patterns around the hinge region of the constant regions IgG1-k0 and IgG2-k0. Various conceivable disulfide bond patterns in IgG2-k0 are indicated by bold lines.
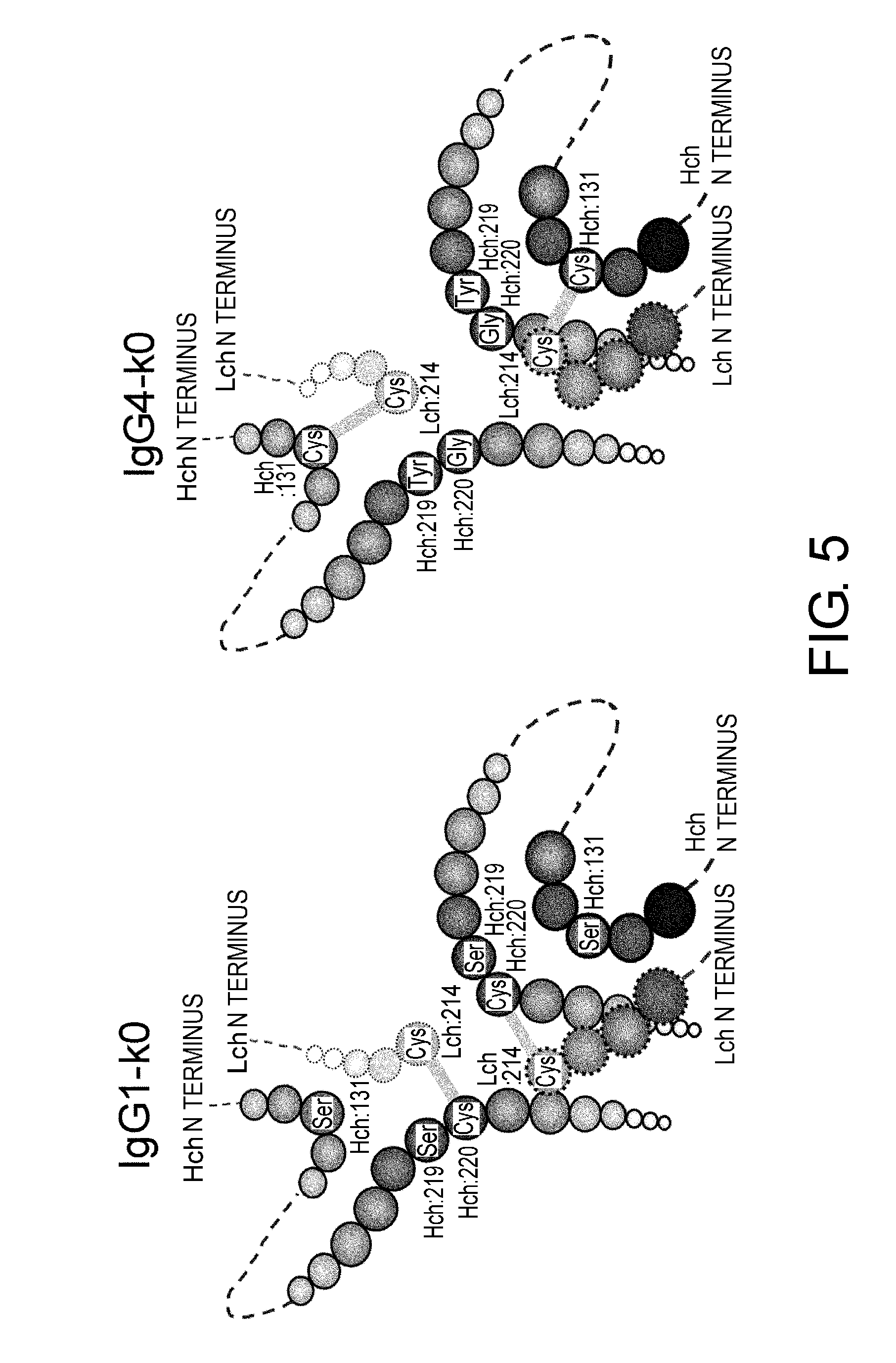
[0049] FIG. 5 shows in diagrams predicted disulfide bond patterns around the hinge region of the constant regions IgG1-k0 and IgG4-k0. Disulfide bond patterns linking H and L chains are different between IgG1-k0 and IgG4-k0.
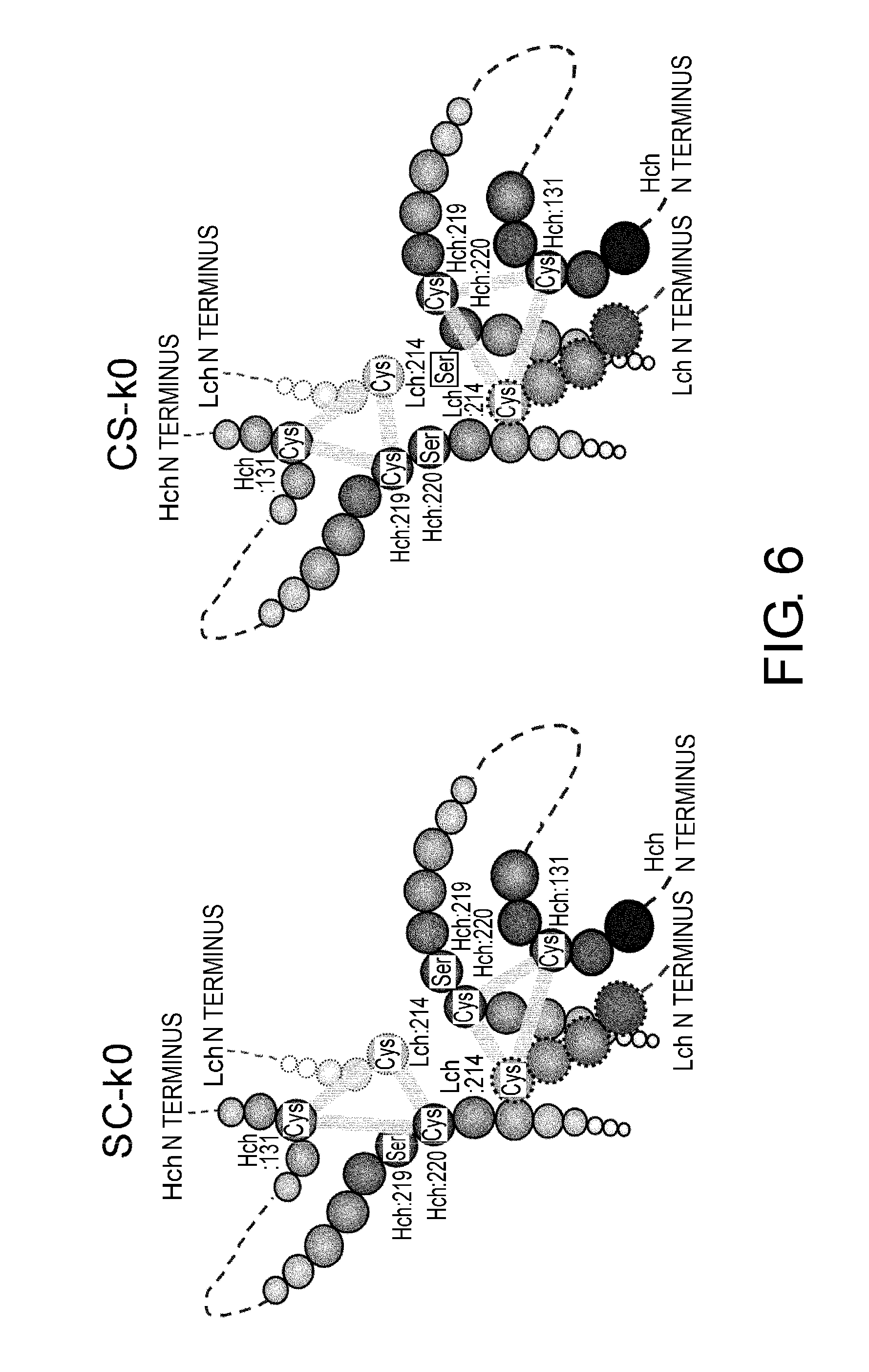
[0050] FIG. 6 shows in diagrams predicted disulfide bond patterns around the hinge region of the constant regions SC-k0 and CS-k0. Various conceivable disulfide bond patterns in SC-k0 and CS-k0 are indicated by bold lines.
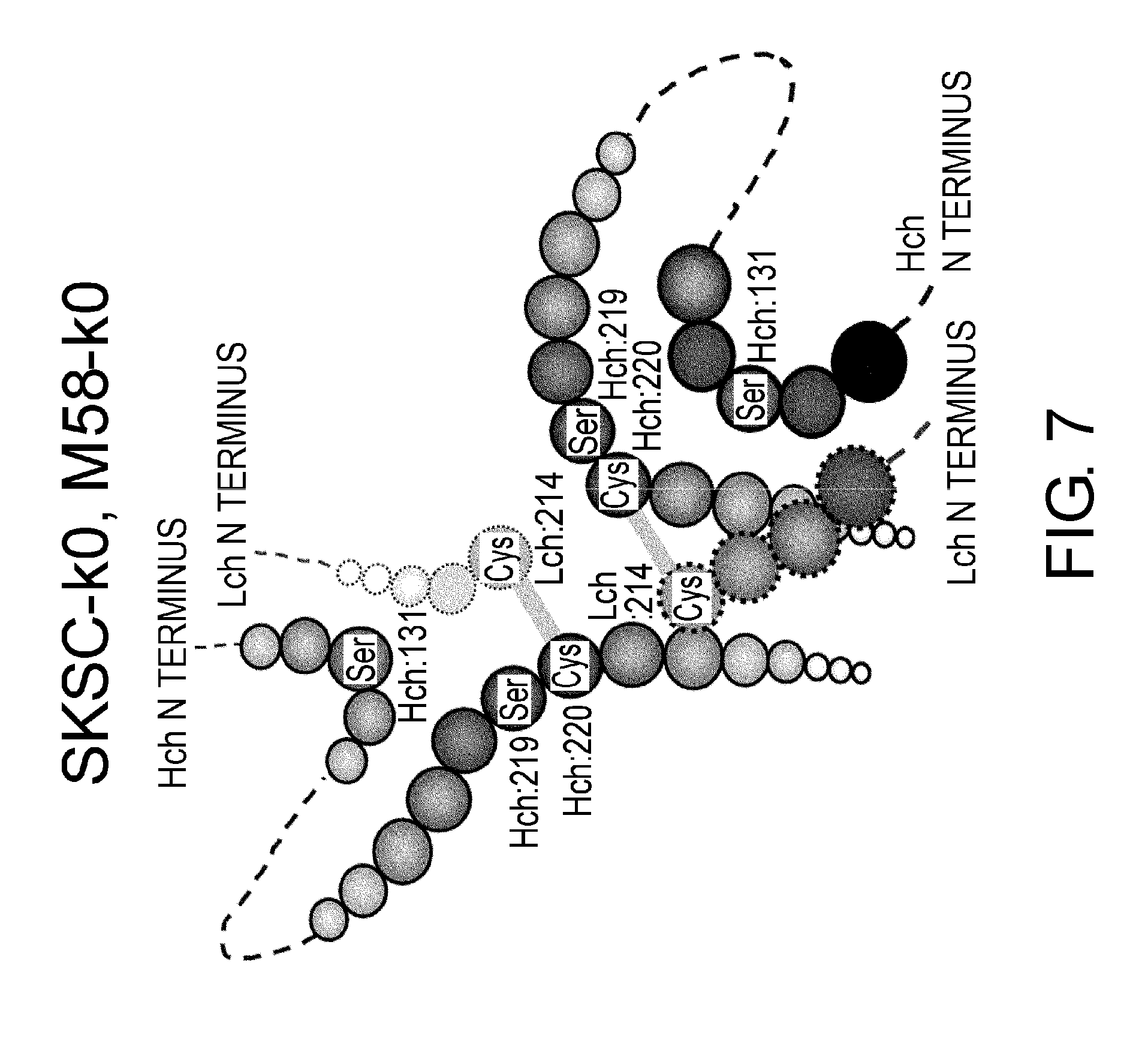
[0051] FIG. 7 shows in a diagram a predicted disulfide bond pattern around the hinge region of constant regions SKSC-k0 and M58-k0.
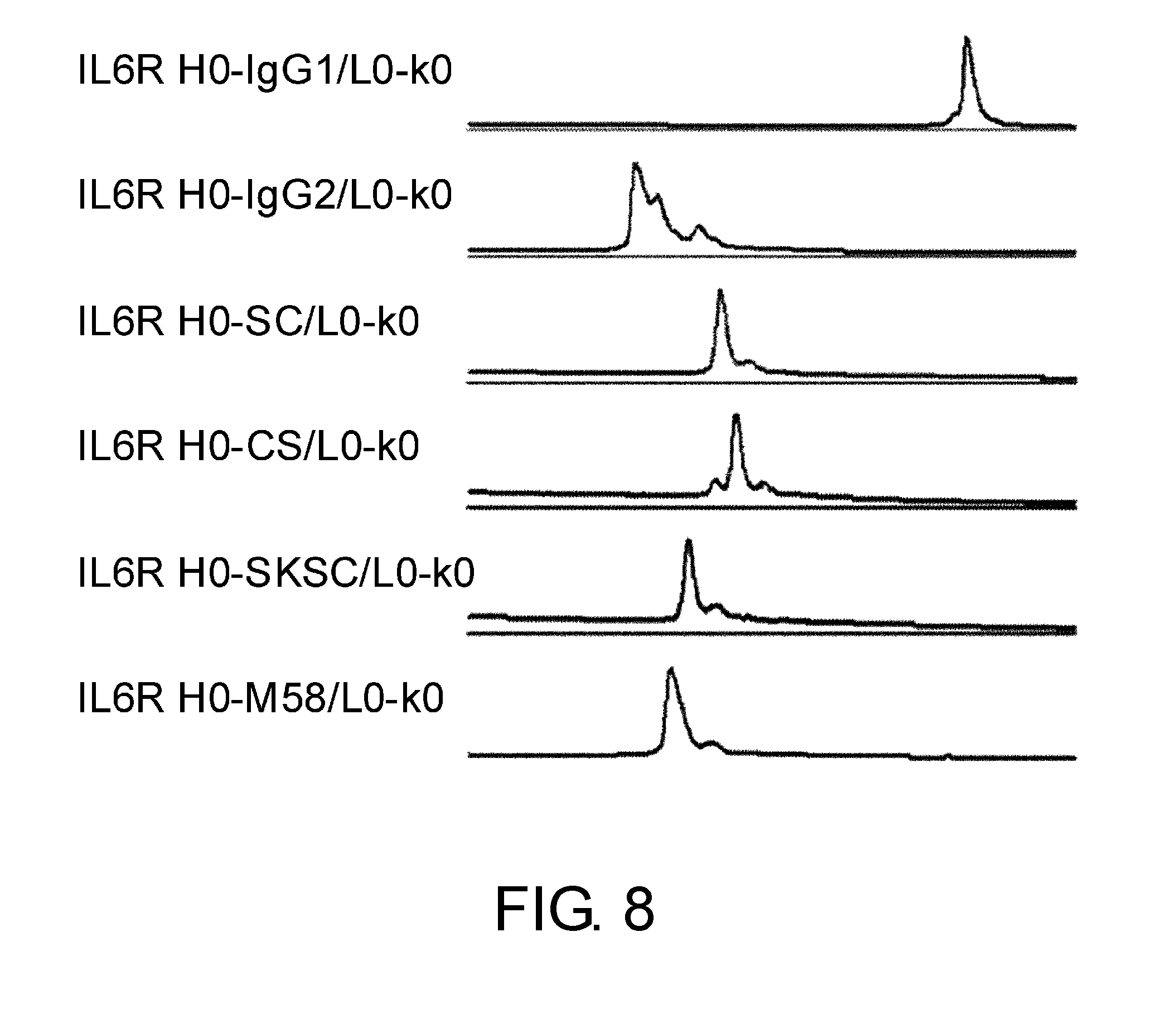
[0052] FIG. 8 shows in graphs the results of assessing the heterogeneity of IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-SC/L0-k0, IL6R H0-CS/L0-k0, IL6R H0-SKSC/L0-k0, and IL6R H0-M58/L0-k0 by cation exchange chromatography based on their disulfide bond differences. In these graphs, the vertical axis shows absorbance at 280 nm and the horizontal axis shows elution time (minutes).
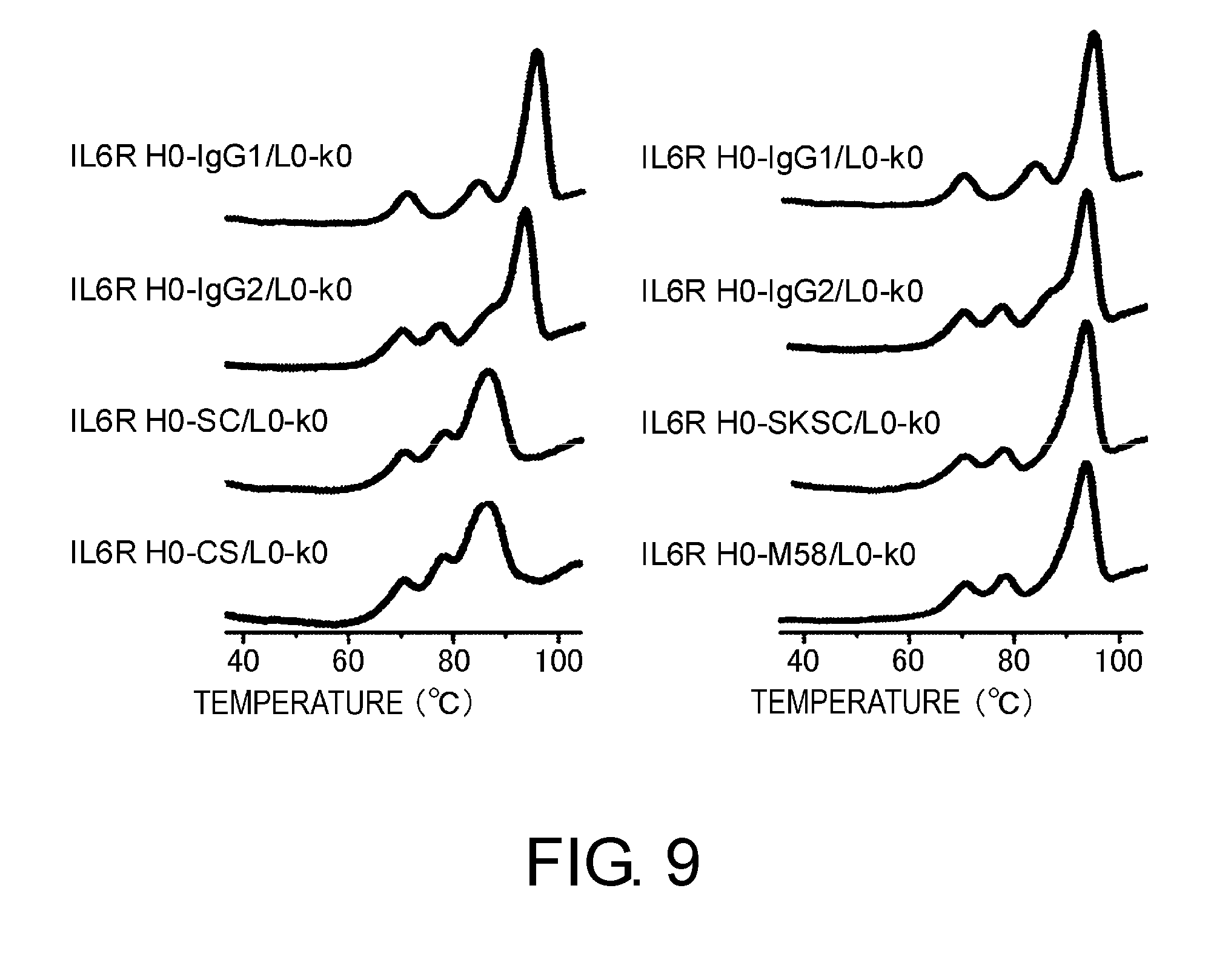
[0053] FIG. 9 shows denaturation curves determined by measuring IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-SC/L0-k0, IL6R H0-CS/L0-k0, IL6R H0-SKSC/L0-k0, and IL6R H0-M58/L0-k0 by differential scanning calorimetry (DSC).
[0054] FIG. 10 shows in graphs that the heterogeneity is greatly improved in the anti-IL-6 receptor antibody, anti-IL-31 receptor antibody, and anti-RANKL antibody by converting their constant region from IgG2-k0 into M58-k0. In these graphs, the vertical axis shows absorbance at 280 nm and the horizontal axis shows elution time (minutes).
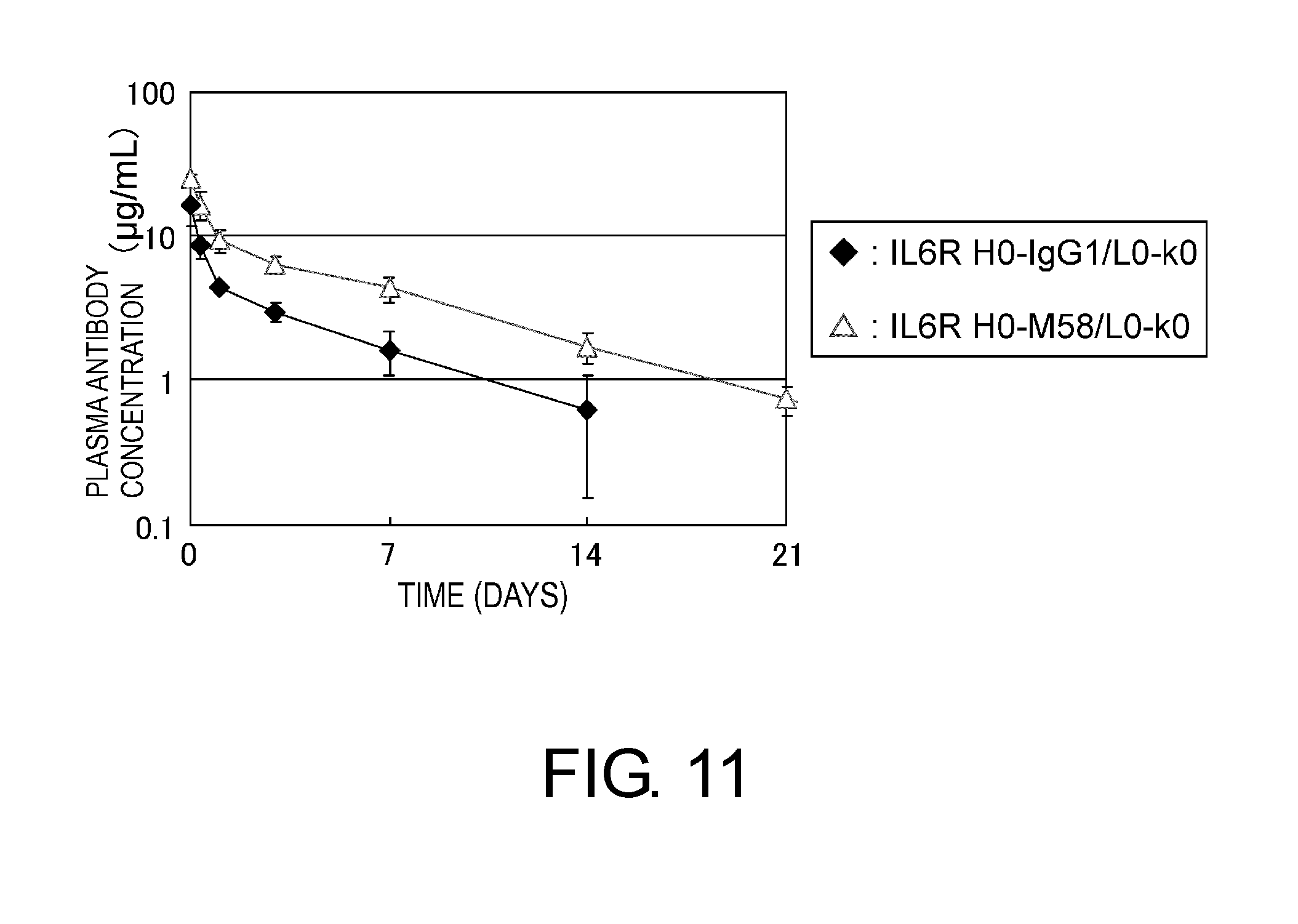
[0055] FIG. 11 shows in a graph a time course of plasma antibody concentration after administration of IL6R H0-IgG1/L0-k0 or IL6R H0-M58/L0-k0 at 1 mg/kg to human FcRn transgenic mice. In this graph, the vertical axis shows plasma antibody concentration (.mu.g/ml), and the horizontal axis shows time after administration (days). The antibodies administered were IL6R H0-IgG1/L0-k0 (filled diamond, antibody with unaltered constant region) and IL6R H0-M58/L0-k0 (open diamond, antibody with altered constant region).
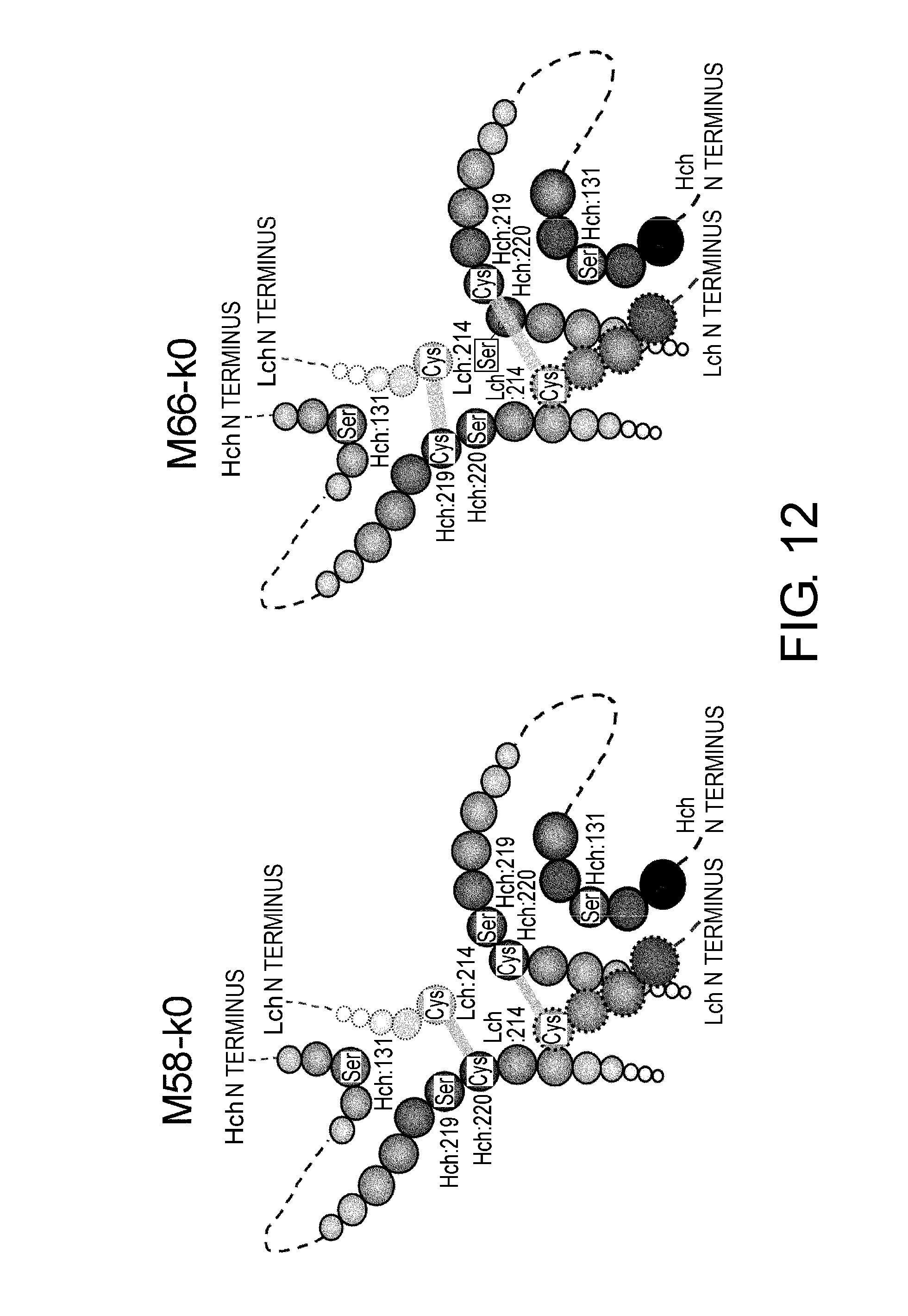
[0056] FIG. 12 shows in diagrams predicted disulfide bond patterns around the hinge regions of constant regions M58-k0 and M66-k0.
[0057] FIG. 13 shows in a graph a time course of plasma antibody concentration after administration of IL6R H0-IgG1/L0-k0, IL6R H0-M58/L0-k0, or IL6R H0-M66/L0-k0 at 1 mg/kg to human FcRn transgenic mice. In this graph, the vertical axis shows plasma antibody concentration (.mu.g/ml), and the horizontal axis shows time after administration (days). The antibodies administered were IL6R H0-IgG1/L0-k0 (filled diamond, antibody with unaltered constant region), IL6R H0-M58/L0-k0 (open diamond, antibody with altered constant region), and IL6R H0-M66/L0-k0 (filled circle, antibody with altered constant region).
[0058] FIG. 14 shows in graphs the results of assessing the heterogeneity of IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-M58/L0-k0, and IL6R H0-M66/L0-k0 by cation exchange chromatography based on their disulfide bond differences. In these graphs, the vertical axis shows absorbance at 280 nm and the horizontal axis shows elution time (minutes).
[0059] FIG. 15 shows in diagrams two types of predicted disulfide bond patterns in the constant region M66-k0.
[0060] FIG. 16 shows in diagrams predicted disulfide bond patterns around the hinge region of constant region M66-k0, M66-k3, or M66-k4. The disulfide bonds which correspond to the two peaks of M66 are indicated by bold lines.
[0061] FIG. 17 shows in graphs the results of assessing the heterogeneity of IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-M58/L0-k0, IL6R H0-M66/L0-k0, IL6R H0-M66/L0-k3, and IL6R H0-M66/L0-k4 by cation exchange chromatography based on their disulfide bond differences. In these graphs, the vertical axis shows absorbance at 280 nm and the horizontal axis shows elution time (minutes).
[0062] FIG. 18 shows in graphs the results of assessing the heterogeneity of IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-M106/L0-k0, IL6R H0-M106/L0-k3, and IL6R H0-M106/L0-k4 by cation exchange chromatography based on their disulfide bond differences. In these graphs, the vertical axis shows absorbance at 280 nm and the horizontal axis shows elution time (minutes).
[0063] FIG. 19 shows in a diagram the binding of H0-IgG1/L0-k0, H0-IgG2/L0-k0, or H0-M106/L0-k3 to various types of Fc.gamma. receptors.
[0064] FIG. 20 shows in graphs the results of assessing the heterogeneity of IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, and IL6R H0-IgG2/L0-k3 by cation exchange chromatography based on their disulfide bond differences. In these graphs, the vertical axis indicates absorbance at 280 nm and the horizontal axis indicates elution time (minutes).
[0065] FIG. 21 shows in a graph a time course of plasma antibody concentration after administration of IL6R H0-M66/L0-k0, IL6R H0-M66/L0-k3, IL6R H0-M106/L0-k3, or IL6R H0-IgG2/L0-k3 at 1 mg/kg to human FcRn transgenic mice. In this graph, the vertical axis shows plasma antibody concentration (.mu.g/ml), and the horizontal axis shows time after administration (days).
DETAILED DESCRIPTION
[0066] The present invention provides antibody constant regions with an altered amino acid sequence, antibodies comprising such constant regions, pharmaceutical compositions comprising such antibodies, and methods for producing them.
[0067] The constant regions of antibody heavy chain include IgG1-, IgG2-, IgG3-, and IgG4-type constant regions. The heavy-chain constant region of the present invention is not particularly limited; however, it is preferably a human heavy-chain constant region. A human IgG2 constant region is particularly preferred in the present invention. The amino acid sequence of human IgG2 constant region is known in the art (SEQ ID NO: 24). A number of allotype sequences of human IgG2 constant regions due to gene polymorphisms are described in "Sequences of proteins of immunological interest", NIH Publication No. 91-3242. Any of such sequences may be used in the present invention.
[0068] Meanwhile, antibody light-chain constant regions include .kappa.- and .lamda.-chain type constant regions. The light-chain constant region of the present invention is not particularly limited; however, it is preferably a human light-chain constant region. A human .kappa. chain constant region is preferred in the present invention. The amino acid sequence of human .kappa. chain constant region is known (SEQ ID NO: 32). A number of allotype sequences of human .kappa. and .lamda. chain constant regions due to gene polymorphisms are described in "Sequences of proteins of immunological interest", NIH Publication No. 91-3242. Any of such sequences may be used in the present invention.
[0069] The antibody constant regions of the present invention with an amino acid alteration (substitution, deletion, addition and/or insertion) may additionally contain other amino acid alterations and modifications, as long as they comprise amino acid alternations of the present invention.
[0070] Specifically, constant regions with the following alterations are all included in the present invention. [0071] An alteration of the present invention is introduced into the amino acid sequence of SEQ ID NO: 24 (human IgG2 constant region). [0072] An alteration of the present invention is introduced into an altered amino acid sequence of SEQ ID NO: 24 (human IgG2 constant region). [0073] An alteration of the present invention plus an additional alteration is introduced into the amino acid sequence of SEQ ID NO: 24 (human IgG2 constant region).
[0074] Furthermore, constant regions with the following alterations are also included in the present invention. [0075] An alteration of the present invention is introduced into the amino acid sequence of SEQ ID NO: 32 (human .kappa. chain constant region). [0076] An alteration of the present invention is introduced into an altered amino acid sequence of SEQ ID NO: 32 (human .kappa. chain constant region). [0077] An alteration of the present invention plus an additional alteration is introduced into the amino acid sequence of SEQ ID NO: 32 (human .kappa. chain constant region).
[0078] Furthermore, constant regions with the following alterations are also included in the present invention. [0079] An alteration of the present invention is introduced into the amino acid sequence of SEQ ID NO: 37 (human .lamda. chain constant region). [0080] An alteration of the present invention is introduced into an altered amino acid sequence of SEQ ID NO: 37 (human .lamda. chain constant region). [0081] An alteration of the present invention plus an additional alteration is introduced into the amino acid sequence of SEQ ID NO: 37 (human .lamda. chain constant region).
[0082] Furthermore, when sugar chains bind to the constant regions, they may have any structure. For example, the sugar chain bound at position 297 (EU numbering) may have any sugar chain structure (fucosylated sugar chains are preferred). Alternatively, it is acceptable that the constant regions have no sugar chain (for example, such constant regions can be produced in E. coli).
<Amino Acid-Altered IgG2 Constant Regions and Antibodies Comprising Such Constant Regions>
[0083] The present invention provides heavy-chain constant regions with improved stability, heterogeneity, immunogenicity, and/or pharmacokinetics. The present invention also provides antibodies comprising the heavy-chain constant region.
[0084] More specifically, the present invention provides heavy chain constant regions that comprise an amino acid sequence in which Cys at position 14 (position 131 (EU numbering)), Arg at position 16 (position 133 (EU numbering)), Cys at position 103 (position 220 (EU numbering)), Glu at position 20 (position 137 (EU numbering)), Ser at position 21 (position 138 (EU numbering)), His at position 147 (position 268 (EU numbering)), Arg at position 234 (position 355 (EU numbering)), and Gln at position 298 (position 419 (EU numbering)) in a heavy-chain constant region (IgG2 constant region) having the amino acid sequence of SEQ ID NO: 24 has been substituted with other amino acids, and antibodies comprising the heavy-chain constant regions.
[0085] Amino acids after substitution are not particularly limited; however, substitutions of Ser for Cys at position 14; Lys for Arg at position 16; Ser for Cys at position 103; Gly for Glu at position 20; Gly for Ser at position 21; Gln for His at position 147; Gln for Arg at position 234; and Glu for Gln at position 298 are preferred.
[0086] Such substitutions can improve antibody stability, immunogenicity, and/or pharmacokinetics. In particular, such substitutions enable one to provide excellent heavy-chain constant regions that are superior to IgG1 in pharmacokinetics, stability, and immunogenicity as well as to provide antibodies comprising such heavy-chain constant regions.
[0087] As long as heavy-chain constant regions of the present invention comprise at least the above-described amino acid substitution, they may comprise other amino acid alterations (substitutions, deletions, additions, insertions, and/or such) or modifications.
[0088] Furthermore, the present invention provides heavy-chain constant regions whose Fc.gamma. receptor-binding activity has been reduced by additionally substituting amino acids in the above-described heavy-chain constant regions. The present invention also provides antibodies comprising the heavy-chain constant regions.
[0089] More specifically, the present invention provides heavy-chain constant regions having an amino acid sequence in which Cys at position 14 (position 131 (EU numbering)), Arg at position 16 (position 133 (EU numbering)), Cys at position 103 (position 220 (EU numbering)), Glu at position 20 (position 137 (EU numbering)), Ser at position 21 (position 138 (EU numbering)), His at position 147 (position 268 (EU numbering)), Arg at position 234 (position 355 (EU numbering)), Gln at position 298 (position 419 (EU numbering)), Ala at position 209 (position 330 (EU numbering)), Pro at position 210 (position 331 (EU numbering)), and Thr at position 218 (position 339 (EU numbering)) in a heavy-chain constant region (IgG2 constant region) comprising the amino acid sequence of SEQ ID NO: 24 have been substituted with other amino acids. The present invention also provides antibodies comprising such heavy-chain constant regions.
[0090] Amino acids after substitution are not particularly limited; however, substitutions of Ser for Cys at position 14; Lys for Arg at position 16; Ser for Cys at position 103; Gly for Glu at position 20; Gly for Ser at position 21; Gln for His at position 147; Gln for Arg at position 234; and Glu for Gln at position 298; Ser for Ala at position 209; Ser for Pro at position 210; and Ala for Thr at position 218 are preferred.
[0091] Such substitutions can improve antibody stability, heterogeneity, immunogenicity, safety, and/or pharmacokinetics.
[0092] As long as heavy-chain constant regions of the present invention comprise at least the above-described amino acid substitution, they may comprise other amino acid alterations (substitutions, deletions, additions, insertions, and/or such) or modifications.
[0093] Furthermore, the present invention provides heavy-chain constant regions comprising an amino acid sequence additionally having deletion of Gly and Lys at positions 325 and 326 (positions 446 and 447 (EU numbering)), respectively, in the above-described heavy-chain constant regions. The present invention also provides antibodies comprising the heavy-chain constant regions. The C-terminal heterogeneity can be improved by deleting these amino acids.
[0094] Specifically, such heavy-chain constant regions with altered amino acids include, for example, heavy-chain constant regions comprising the amino acid sequence of SEQ ID NO: 30 (M66) or 31 (M106).
[0095] Those described above are optimized heavy-chain constant regions with reduced Fc.gamma. receptor-binding activity, reduced immunogenicity risk, reduced hinge-region heterogeneity, reduced C-terminal heterogeneity, and/or improved pharmacokinetics.
[0096] Furthermore, in the present invention, the heavy-chain constant regions of the present invention may comprise amino acid alterations to improve the stability under acidic conditions, in addition to the above-described amino acid alterations.
[0097] Specifically, the amino acid alterations to improve stability under acidic conditions include, for example, substitution of Met at position 276 (position 397 (EU numbering)) in the IgG2 constant region having the amino acid sequence of SEQ ID NO: 24 with another amino acid. The other amino acid is not particularly limited; however, Val is preferred. The substitution of Met at position 276 (position 397 (EU numbering)) in the amino acid sequence of SEQ ID NO: 24 with another amino acid can improve antibody stability under acidic conditions.
<.kappa. Chain Constant Regions with Altered Amino Acids and Antibodies Comprising Such .kappa. Chain Constant Regions>
[0098] Furthermore, the present invention provides light-chain constant regions that can be used to improve the heterogeneity of hinge region. The present invention also provides antibodies comprising such light-chain constant regions.
[0099] More specifically, the present invention provides human .kappa. chain constant regions having at least one Cys at positions 102 to 106, and antibodies comprising the human .kappa. chain constant regions. For example, "a human .kappa. chain comprising the amino acid sequence of SEQ ID NO: 32 has at least one Cys at positions 102 to 106" means that there is at least one Cys in the region between Phe at position 102 and Glu at position 106.
[0100] The number of Cys present in the region of positions 102 to 106 in the human .kappa. chain is not particularly limited; however, the number is five or less, preferably three or less, more preferably two or less, and still more preferably one.
[0101] The position of Cys is not particularly limited; however, Cys is preferably located at position 104, 105, or 106, more preferably at position 105 or 106, and particularly preferably at position 106.
[0102] The number of amino acids in the human .kappa. chain constant region that has at least one Cys at positions 102 to 106 is not particularly limited; however, the number is preferably 102 to 107 amino acids, more preferably 105 or 106 amino acids, and still more preferably 106 amino acids.
[0103] Methods for producing a human .kappa. chain constant region that has at least one Cys at positions 102 to 106 are not particularly limited, and include, for example, the methods described below. It is also possible to use a combination of the insertion, substitution, and deletion described below.
[0104] Insertion of at least one Cys at positions 102 to 106;
[0105] Substitution of Cys for at least one amino acid at positions 102 to 106;
[0106] Deletion of one to five amino acids at positions 1 to 106.
[0107] Furthermore, the present invention provides human .kappa. chain constant regions that do not have Cys at position 107, and antibodies comprising such human .kappa. chain constant regions. For example, "a human .kappa. chain comprising the amino acid sequence of SEQ ID NO: 32 does not have Cys at position 107" means deletion of Cys at position 107, substitution of another amino acid for Cys at position 107, insertion of other amino acids, relocation of Cys from position 107 to a different position, etc. Preferred human .kappa. chain constant regions include those having deletion of Cys at position 107, relocation of Cys from position 107 to a different position, or substitution of Cys at position 107 with another amino acid.
[0108] In a preferred embodiment, human .kappa. chain constant regions of the present invention include those which have at least one Cys at positions 102 to 106 but do not have Cys at position 107.
[0109] Preferred human .kappa. chain constant regions include, for example, those having deletion of at least one amino acid at positions 1 to 106. For example, in the human .kappa. chain constant region having the amino acid sequence of SEQ ID NO: 32, deletion of Glu at position 106 leads to relocation of Cys from position 107 to 106, resulting in a human .kappa. chain constant region having Cys at position 106 but not at position 107. The position of amino acid deletion is not particularly limited; however, a human .kappa. chain constant region preferably comprises deletion of at least one amino acid at positions 102 to 106, more preferably deletion of the amino acid at position 105 or 106.
[0110] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0111] Such preferred human .kappa. chain constant regions include, for example, those having deletion of the amino acid at position 105 or 106.
[0112] In another preferred embodiment, the above-described human .kappa. chain constant regions include those having substitution of Cys for at least one amino acid at positions 102 to 106 and additionally having deletion of Cys at position 107 or substitution of Cys at position 107 with another amino acid. The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0113] The position of substitution by Cys is not particularly limited; however, such preferred substitution positions include positions 105 and 106.
[0114] Such preferred human .kappa. chain constant regions include, for example, those having substitution of Cys for Gly at position 105, and deletion of Cys at position 107 or substitution of another amino acid for Cys at position 107, and those having substitution of Cys for Glu at position 106, and deletion of Cys at position 107 or substitution of Cys at position 107 with another amino acid. Specific examples of the human .kappa. chain constant regions of the present invention include those comprising the amino acid sequence of SEQ ID NO: 33 (k3) or 34 (k4).
[0115] The human .kappa. chain constant regions of the present invention may comprise other amino acid alterations in addition to the above-described amino acid alterations. Such human .kappa. chain constant regions additionally comprising other amino acid alterations and modifications are also included in the human .kappa. chain constant regions of the present invention, as long as they comprise the above-described amino acid alteration.
[0116] The heterogeneity of hinge region can be reduced by using the human .kappa. chain constant region of the present invention. In particular, the .kappa. chain constant region of the present invention is efficient when used in combination with a heavy-chain constant region having Cys either at position 219 or 220 in the EU numbering (for example, heavy-chain constant regions having Cys only at position 219 in the EU numbering) such as a heavy-chain constant region comprising the amino acid sequence of SEQ ID NO: 30 (M66) or 31 (M106).
<.lamda. Chain Constant Region with Altered Amino Acids and Antibodies Comprising the .lamda. Chain Constant Regions>
[0117] Furthermore, the present invention provides light-chain constant regions that can be used to reduce the heterogeneity of hinge region. The present invention also provides antibodies comprising the above-described light-chain constant regions.
[0118] More specifically, the present invention provides human .lamda. chain constant regions having at least one Cys at positions 99 to 103, and antibodies comprising such human .lamda. chain constant regions. For example, "a human .lamda. chain comprising the amino acid sequence of SEQ ID NO: 37 has at least one Cys at positions 99 to 103" means that there is at least one Cys in the region between Val at position 99 and Glu at position 103.
[0119] The number of Cys present in the region at positions 99 to 103 in the human .lamda. chains is not particularly limited; however, the number is five or less, preferably three or less, more preferably two or one, and still more preferably one.
[0120] The position of Cys is not particularly limited; however, Cys is preferably located at position 101, 102, or 103, more preferably at position 102 or 103, and particularly preferably at position 103.
[0121] The number of amino acids in a human .lamda. chain that has at least one Cys at positions 99 to 103 is not particularly limited; however, the number is preferably 100 to 103 amino acids, more preferably 102 or 103 amino acids, and still more preferably 103 amino acids.
[0122] Methods for producing a .lamda. chain constant region that has at least one Cys at positions 99 to 103 are not particularly limited, and include, for example, the methods described below. It is also possible to use a combination of the insertion, substitution, and deletion described below.
[0123] Insertion of at least one Cys at positions 99 to 103;
[0124] Substitution of Cys at least one amino acid at positions 99 to 103;
[0125] Deletion of one to five amino acids at positions 1 to 103.
[0126] Furthermore, the present invention provides human .lamda. chain constant regions that do not have Cys at position 104, and antibodies comprising such human .lamda. chain constant regions. For example, "a human .lamda. chain comprising the amino acid sequence of SEQ ID NO: 37 does not have Cys at position 104" means deletion of Cys at position 104, substitution of another amino acid for Cys at position 104, insertion of another amino acid, relocation of Cys from position 104 to another position; etc. Preferred human .lamda. chain constant regions include those having deletion of Cys at position 104, relocation of Cys from position 104 to a different position, or substitution of another amino acid for Cys at position 104.
[0127] In a preferred embodiment, human .lamda. chain constant region of the present invention include those that have at least one Cys at positions 99 to 103 but do not have Cys at position 104.
[0128] Such preferred human .lamda. chain constant regions include, for example, those having deletion of at least one amino acid at positions 1 to 103. For example, in the human .lamda. chain constant region comprising the amino acid sequence of SEQ ID NO: 37, the deletion of Glu at position 103 leads to relocation of Cys at position 104 to position 103, resulting in a human .lamda. chain constant region having Cys at position 103 but not at position 104. The position of amino acid deletion is not particularly limited; however, a human .kappa. chain constant region preferably comprises deletion of at least one amino acid at positions 99 to 103, more preferably deletion of the amino acid at position 102 or 103.
[0129] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0130] Preferred examples of human .lamda. chain constant regions include those having an amino acid deletion at position 102 or 103.
[0131] In another preferred embodiment, the above-described human .lamda. chain constant regions include those having substitution of Cys for at least one amino acid at positions 99 to 103, and additionally having deletion of Cys at position 104 or substitution of Cys at position 104 with another amino acid. The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0132] The position of substitution with Cys is not particularly limited; however, such preferred substitution positions include 102 and 103.
[0133] Such preferred human .lamda. chain constant regions include, for example, those having substitution of Cys for Thr at position 102, and deletion of Cys at position 104 or substitution of Cys at position 104 with another amino acid, and those having substitution of Cys for Glu at position 103, and deletion of Cys at position 104 or substitution of Cys at position 104 with another amino acid. Specifically, the human .lamda. chain constant regions of the present invention include, for example, those comprising the amino acid sequence of SEQ ID NO: 38 or 39.
[0134] The human .lamda. chain constant regions of the present invention may comprise other amino acid alterations in addition to the above-described amino acid alterations. The human .lamda. chain constant regions additionally comprising other amino acid alterations and modifications are also included in the human .lamda. chain constant regions of the present invention, as long as they comprise the above-described amino acid alteration.
[0135] The heterogeneity of hinge region can be reduced by using the human .lamda. chain constant regions of the present invention. In particular, the human .lamda. chain constant region of the present invention is efficient when used in combination with a heavy-chain constant region comprising Cys either at position 219 or 220 in the EU numbering (for example, heavy-chain constant regions comprising Cys only at position 219 in the EU numbering) such as a heavy-chain constant region comprising the amino acid sequence of SEQ ID NO: 30 (M66) or 31 (M106).
[0136] Without being restricted to a particular theory, the reason why the human .kappa. or .lamda. chain constant regions of the present invention reduces the heterogeneity of hinge region can be described below using .kappa. chain as an example.
[0137] As shown in FIG. 15, cysteine at position 107 in the human .kappa. chain constant region can form a disulfide bond with cysteine at position 219 (EU numbering) in both of the two H chains of an antibody. It is thought that the resulting two types of disulfide bonds cause heterogeneity in hinge region.
[0138] On the other hand, as in the human .kappa. chain constant region of the present invention, by relocating cysteine to its N-terminal side, the distance between this cysteine and cysteine at position 219 (EU numbering) in one H chain becomes greater, and as a result cysteine in the chain constant region can only form a disulfide bond with cysteine at position 219 (EU numbering) in one of the two H chains. This causes reduction of heterogeneity in hinge region (see FIG. 16). Specifically, the heterogeneity of hinge region can be reduced by increasing the distance between cysteine in the human .kappa. chain constant region and cysteine at position 219 (EU numbering) in one H chain. In the same manner, the heterogeneity of hinge region in a human .lamda. chain constant region can also be reduced by increasing the distance from cysteine at position 219 (EU numbering) in one H chain.
<.kappa. Chain Constant Regions Derived from Nonhuman Animals and Antibodies Comprising Such Chain Constant Regions>
[0139] The present invention can also be used to alter light-chain constant regions derived from nonhuman animals. Examples of light-chain constant regions derived from nonhuman animals include mouse antibody .kappa. chain constant region (SEQ ID NO: 40), rat antibody .kappa. chain constant region (SEQ ID NO: 41), and rabbit antibody .kappa. chain constant regions (SEQ ID NOs: 42 and 43), but are not limited thereto.
[0140] Thus, the present invention provides mouse and rat antibody .kappa. chain constant regions having at least one Cys at positions 102 to 106, and antibodies comprising such light-chain constant regions. For example, "there is at least one Cys at positions 102 to 106 in the mouse chain constant region comprising the amino acid sequence of SEQ ID NO: 40 or the rat .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 41" means that there is at least one Cys in the region between Phe at position 102 and Glu at position 106.
[0141] Furthermore, the present invention provides rabbit .kappa. chain constant regions having at least one Cys at positions 99 to 103 in the rabbit antibody .kappa. chain constant region (SEQ ID NO: 42), and antibodies comprising such rabbit .kappa. chain constant regions. For example, "there is at least one Cys at positions 99 to 103 in the rabbit .kappa. chain comprising the amino acid sequence of SEQ ID NO: 42" means that there is at least one Cys in the region between Phe at position 99 and Asp at position 103.
[0142] The present invention also provides rabbit .kappa. chain constant regions having at least one Cys at positions 101 to 105 in the rabbit antibody .kappa. chain constant region (SEQ ID NO: 43), and antibodies comprising the rabbit .kappa. chain constant region. For example, "there is at least one Cys at positions 101 to 105 in the rabbit .kappa. chain comprising the amino acid sequence of SEQ ID NO: 43" means that there is at least one Cys in the region between Phe at position 101 and Asp at position 105.
[0143] The number of Cys present in the region of positions 102 to 106 in a mouse .kappa. chain constant region, positions 102 to 106 in a rat .kappa. chain constant region, positions 99 to 103 in the rabbit .kappa. chain constant region (SEQ ID NO: 42), or positions 101 to 105 in the rabbit .kappa. chain constant region (SEQ ID NO: 43) is not particularly limited; however, the number is five or less, preferably three or less, more preferably two or less, and still more preferably one.
[0144] The position of Cys is not particularly limited; however,
in a mouse or rat .kappa. chain constant region, the position is preferably 104, 105, or 106, more preferably 105 or 106, and particularly preferably 106; in the rabbit .kappa. chain constant region (SEQ ID NO: 42), the position is preferably 101, 102, or 103, more preferably 102 or 103, and particularly preferably 103; and in the rabbit .kappa. chain constant region (SEQ ID NO: 43), the position is preferably 103, 104, or 105, more preferably 104 or 105, and particularly preferably 105.
[0145] The number of amino acids in such .kappa. chain constant region is not particularly limited; however,
in mouse or rat .kappa. chain constant regions, the number is preferably 102 to 107 amino acids, more preferably 105 or 106 amino acids, and still more preferably 106 amino acids; in the rabbit .kappa. chain constant region (SEQ ID NO: 42), the number is preferably 99 to 104 amino acids, more preferably 102 or 103 amino acids, and still more preferably 103 amino acids; and in the rabbit .kappa. chain constant region (SEQ ID NO: 43), the number is preferably 101 to 106 amino acids, more preferably 104 or 105 amino acids, and still more preferably 105 amino acids.
[0146] Methods for producing a mouse or rat .kappa. chain constant region that has at least one Cys at positions 102 to 106 are not particularly limited, and include, for example, the methods described below. It is also possible to use in combination the insertion, substitution, and deletion described below.
[0147] Insertion of at least one Cys at positions 102 to 106.
[0148] Substitution of Cys for at least one amino acid at positions 102 to 106.
[0149] Deletion of one to five amino acids at positions 1 to 106.
[0150] Meanwhile, methods for producing a mouse or rabbit .kappa. chain constant region that has at least one Cys at positions 99 to 103 are not particularly limited, and include, for example, the methods described below. It is also possible to use in combination the insertion, substitution, and deletion described below.
[0151] Insertion of at least one Cys at positions 99 to 103.
[0152] Substitution of Cys for at least one amino acid at positions 99 to 103.
[0153] Deletion of one to five amino acids at positions 1 to 103.
[0154] Methods for producing a rabbit .kappa. chain constant region that has at least one Cys at positions 101 to 105 are not particularly limited, and include, for example, the methods described below. It is also possible to use in combination the insertion, substitution, and deletion described below.
[0155] Insertion of at least one Cys at positions 101 to 105.
[0156] Substitution of Cys for at least one amino acid at positions 101 to 105.
[0157] Deletion of one to five amino acids at positions 1 to 105.
[0158] Furthermore, the present invention provides mouse and rat .kappa. chain constant regions that do not have Cys at position 107, and antibodies comprising such .kappa. chain constant regions. For example, "there is no Cys at position 107 in a mouse .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 40 or a rat .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 41" means deletion of Cys at position 107, substitution of Cys at position 107 with another amino acid, insertion of other amino acids, relocation of Cys from position 107 to a different position; etc. Preferred .kappa. chain constant regions include those having deletion of Cys at position 107, relocation of Cys from position 107 to another position, or substitution of Cys at position 107 with another amino acid.
[0159] In a preferred embodiment, .kappa. chain constant regions of the present invention include those which have at least one Cys at positions 102 to 106 but do not have Cys at position 107. Such preferred .kappa. chain constant regions include, for example, .kappa. chain constant regions having deletion of at least one amino acid at positions 1 to 106. For example, in a .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 40 or 41, deletion of Glu at position 106 leads to relocation of Cys from position 107 to position 106, resulting in a .kappa. chain constant region having Cys at position 106 but not at position 107. The position of amino acid deletion is not particularly limited; however, the .kappa. chain constant region preferably comprises deletion of at least one amino acid at positions 102 to 106, more preferably deletion of the amino acid at position 105 or 106.
[0160] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0161] Such preferred .kappa. chain constant regions include, for example, those having deletion of the amino acid at position 105 or 106.
[0162] In another preferred embodiment, the above-described mouse and rat .kappa. chain constant regions which do not have Cys at position 107 include those having substitution of Cys for at least one amino acid at positions 102 to 106, and additionally having deletion of Cys at position 107 or substitution of Cys at position 107 with another amino acid. The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0163] The position of Cys substitution is not particularly limited; however, such preferred substitution positions include positions 105 and 106.
[0164] Such preferred .kappa. chain constant regions include, for example, those having substitution of Cys for Asn at position 105, and deletion of Cys at position 107 or substitution of Cys at position 107 with another amino acid, and those having substitution of Cys for Glu at position 106, and deletion of Cys at position 107 or substitution of Cys at position 107 with another amino acid.
[0165] The mouse or rat .kappa. chain constant regions of the present invention may comprise other amino acid alterations in addition to the above-described amino acid alterations. Such .kappa. chain constant regions comprising other amino acid alterations and modifications are also included in the .kappa. chain constant regions of the present invention, as long as they comprise the above-described amino acid alteration.
[0166] Furthermore, the present invention provides rabbit .kappa. chain constant regions which do not have Cys at position 104 in the rabbit .kappa. chain constant region (SEQ ID NO: 42) and antibodies comprising such .kappa. chain constant regions. For example, "there is no Cys as position 104 in the rabbit .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 42" means deletion of Cys at position 104, substitution of Cys at position 104 with another amino acid, insertion of other amino acids, relocation of Cys from position 104 to a different position; etc. Preferred .kappa. chain constant regions include those having deletion of Cys at position 104, relocation of Cys from position 104 to a different position, or substitution of Cys at position 104 with another amino acid.
[0167] In a preferred embodiment of the present invention, .kappa. chain constant regions include those which have at least one Cys at positions 99 to 103 but do not have Cys at position 104.
[0168] Such preferred .kappa. chain constant regions include, for example, those having deletion of at least one amino acid at positions 1 to 103. For example, in a .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 42, deletion of Asp at position 103 leads to relocation of Cys from position 104 to position 103, resulting in a .kappa. chain constant region having Cys at position 103 but not at position 104. The position of amino acid deletion is not particularly limited; however, a .kappa. chain constant region preferably comprises deletion of at least one amino acid at positions 99 to 103, more preferably deletion of the amino acid at position 102 or 103.
[0169] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0170] Such preferred .kappa. chain constant regions include, for example, those having deletion of the amino acid at position 102 or 103.
[0171] In another preferred embodiment, the above-described rabbit .kappa. chain constant regions which do not have Cys at position 104 include those having substitution of Cys for at least one amino acid at positions 99 to 103 and additionally having deletion of Cys at position 104 or substitution of Cys at position 104 with another amino acid. The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0172] The position of Cys substitution is not particularly limited; however, such preferred positions of substitution include positions 102 and 103.
[0173] Such preferred .kappa. chain constant regions include, for example, those having substitution of Cys for Gly at position 102, and deletion of Cys at position 104 or substitution of Cys at position 104 with another amino acid, and those having substitution of Cys for Asp at position 103, and deletion of Cys at position 104 or substitution of Cys at position 104 with another amino acid.
[0174] The rabbit .kappa. chain constant regions of the present invention may comprise other amino acid alterations in addition to the above-described amino acid alterations. .kappa. chain constant regions comprising other amino acid alterations and modifications are also included in the chain constant regions of the present invention, as long as they comprise the above-described amino acid alterations.
[0175] Furthermore, the present invention provides rabbit .kappa. chain constant regions which do not have Cys at position 106 in the rabbit .kappa. chain constant region (SEQ ID NO: 43) and antibodies comprising such .kappa. chain constant regions. For example, "there is no Cys at position 106 in the rabbit .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 43" means deletion of Cys at position 106, substitution of Cys at position 106 with another amino acid, insertion of other amino acids, relocation of Cys from position 106 to a different position; etc. Preferred .kappa. chain constant regions include those having deletion of Cys at position 106, relocation of Cys from position 106 to a different position, or substitution of Cys at position 106 with another amino acid.
[0176] In a preferred embodiment, .kappa. chain constant regions of the present invention include those which have at least one Cys at positions 101 to 105 but do not have Cys at position 106.
[0177] Such preferred .kappa. chain constant regions include, for example, those having deletion of at least one amino acid at positions 1 to 105. For example, in a .kappa. chain constant region comprising the amino acid sequence of SEQ ID NO: 43, deletion of Asp at position 105 leads to relocation of Cys from position 106 to 105, resulting in a .kappa. chain constant region having Cys at position 105 but not at position 106. The position of amino acid deletion is not particularly limited; however, a human .kappa. chain constant region preferably comprises deletion of at least one amino acid at positions 101 to 105, more preferably deletion of the amino acid at position 104 or 105.
[0178] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0179] Such preferred .kappa. chain constant regions include, for example, those having deletion of the amino acid at position 104 or 105.
[0180] In another preferred embodiment, the above-described rabbit .kappa. chain constant regions which do not have Cys at position 106 include those which have substitution of Cys for at least one amino acid at positions 101 to 105, and additionally have deletion of Cys at position 106 or substitution of Cys at position 106 with another amino acid. The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0181] The position of Cys substitution is not particularly limited; however, such preferred positions of substitution include positions 104 and 105.
[0182] Such preferred .kappa. chain constant regions include, for example, those having substitution of Cys for Gly at position 104, and deletion of Cys at position 106 or substitution of Cys at position 106 with another amino acid, and those having substitution of Cys for Asp at position 105, and deletion of Cys at position 106 or substitution of Cys at position 106 with another amino acid.
[0183] The rabbit .kappa. chain constant regions of the present invention may comprise other amino acid alterations in addition to the above-described amino acid alterations. .kappa. chain constant regions comprising other amino acid alterations and modifications are also included in the chain constant regions of the present invention, as long as they comprise the above-described amino acid alterations.
[0184] Furthermore, the present invention provides antibodies comprising heavy-chain constant regions having the above-described amino acid alterations. The present invention also provides antibodies comprising light chain constant regions having the above-described amino acid alterations. The present invention also provides antibodies comprising heavy chain constant regions having the above-described amino acid alterations and light chain constant regions having the above-described amino acid alterations. Amino acid alterations in the antibodies of the present invention include all possible alterations specified by the description herein and combinations thereof.
[0185] The present invention also provides antibodies comprising light chains comprising light chain constant regions having the above-described amino acid alterations and heavy chain constant regions in which at least one Cys is substituted with another amino acid. Such heavy chain constant regions are not particularly limited; however, IgG2 heavy chain constant regions are preferred. When the heavy chain constant region is an IgG2 constant region, Cys to be substituted is not particularly limited; however, the constant region includes, for example, those having substitution of another amino acid for at least one of:
Cys at position 131 in the EU numbering (position 14 in SEQ ID NO: 24), Cys at position 219 in the EU numbering (position 102 in SEQ ID NO: 24), and Cys at position 220 in the EU numbering (position 103 in SEQ ID NO: 24). When two Cys are substituted with other amino acids, the combination is not particularly limited and includes the combination of substitutions at positions 131 and 219 (EU numbering) and the combination of substitutions at positions 131 and 220.
[0186] Furthermore, the present invention provides antibodies comprising light chains comprising light chain constant regions having the above-described amino acid alterations, and heavy chains comprising heavy chain constant regions which have Cys at position 219 in the EU numbering (position 102 in SEQ ID NO: 24) but not at position 220 in the EU numbering (position 103 in SEQ ID NO: 24). Such heavy chain constant regions are not particularly limited; however, they are preferably IgG2 constant regions, more preferably M66 and M106. The antibody constant regions may have one or more amino acid substitutions, deletions, additions, and/or insertions (for example, 20 amino acids or less, or 10 amino acids or less).
[0187] Variable regions that constitute the antibodies of the present invention may recognize any antigen. Preferred variable regions of the present invention include antibody variable regions having an antigen-neutralizing activity. The variable regions that constitute the antibodies of the present invention include, for example, antibody variable regions having an activity of neutralizing IL6 receptor, IL31 receptor, or RANKL.
[0188] The antibodies of the present invention are not particularly limited in type, origin, or such; any antibody may be used in the present invention as long as it has the above-described antibody constant region. The origin of antibodies is not particularly limited. The antibodies include human, mouse, rat, and rabbit antibodies. The antibodies of the present invention may be chimeric, humanized, fully humanized antibodies, or such. In a preferred embodiment, the antibodies of the present invention are humanized antibodies.
[0189] Antibody molecules of the present invention usually include heavy chains and light chains. Heavy chains may include variable regions in addition to constant regions. Variable regions may include variable portions derived not only from humans but also from nonhuman animal species. Furthermore, CDRs from variable portions derived from non-human species such as mice can be transplanted to humanize the variable portions. Antibody molecules composed of heavy chains and light chains may be oligomers. Specifically, they may be monomers, dimers, or larger oligomers.
[0190] Alternatively, the above-described antigen constant regions may be linked with various molecules such as bioactive peptides or antigen-binding peptides to be fusion proteins.
[0191] The antibodies of the present invention also include modification products of an antibody comprising any one of the constant regions described above.
[0192] Such antibody modification products include, for example, antibodies linked with various molecules such as polyethylene glycol (PEG) and cytotoxic substances. Such antibody modification products can be obtained by chemically modifying antibodies of the present invention. Methods for modifying antibodies are already established in this field.
[0193] The antibodies of the present invention may also be bispecific antibodies. "Bispecific antibody" refers to an antibody that has in a single molecule variable regions that recognize different epitopes. The epitopes may be present in a single molecule or in separate molecules.
[0194] The antibody constant regions described above can be used as a constant region in an antibody against an arbitrary antigen. The antigen is not particularly limited.
[0195] The antibodies of the present invention can also be obtained by methods known to one skilled in the art. Methods for substituting or deleting one or more amino acid residues with amino acids of interest include, for example, site-directed mutagenesis (Hashimoto-Gotoh, T., Mizuno, T., Ogasahara, Y., and Nakagawa, M. An oligodeoxyribonucleotide-directed dual amber method for site-directed mutagenesis. Gene (1995) 152: 271-275; Zoller, M. J., and Smith, M. Oligonucleotide-directed mutagenesis of DNA fragments cloned into M13 vectors. Methods Enzymol. (1983) 100: 468-500; Kramer, W., Drutsa, V., Jansen, H. W., Kramer, B., Pflugfelder, M., and Fritz, H. J. The gapped duplex DNA approach to oligonucleotide-directed mutation construction. Nucleic Acids Res. (1984) 12: 9441-9456; Kramer W., and Fritz H. J. Oligonucleotide-directed construction of mutations via gapped duplex DNA Methods. Enzymol. (1987) 154: 350-367; Kunkel, T. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc. Natl. Acad. Sci. USA (1985) 82: 488-492). These methods can be used to substitute target amino acids in the constant region of an antibody with amino acids of interest. Furthermore, one or more amino acid residues can be deleted.
[0196] In another embodiment to obtain antibodies, an antibody that binds to an antigen of interest is first prepared by methods known to those skilled in the art. When the prepared antibody is derived from a nonhuman animal, it can be humanized. The binding activity of the antibody can be determined by known methods. Next, one or more amino acid residues in the constant region of the antibody are deleted or substituted with amino acids of interest.
[0197] The present invention relates to methods for producing antibodies with altered amino acid residues in the heavy chain and/or light chain constant regions, which comprise the steps of:
(a) expressing a DNA encoding a heavy chain having deletion or substitution of one or more amino acid residues in a constant region with amino acids of interest, and/or a light chain having deletion or substitution of one or more amino acid residues in a constant region with amino acids of interest; and (b) collecting the expression product of (a).
[0198] Such alterations of amino acid residues in the heavy chain constant region include, but are not limited to, for example, those described below.
[0199] (1) In an IgG2 constant region (the amino acid sequence of SEQ ID NO: 24),
Ser is substituted for Cys at position 14 (position 131 (EU numbering)); Lys is substituted for Arg at position 16 (position 133 (EU numbering)); Ser is substituted for Cys at position 103 (position 220 (EU numbering)); Gly is substituted for Glu at position 20 (position 137 (EU numbering)); Gly is substituted for Ser at position 21 (position 138 (EU numbering)); Gln is substituted for His at position 147 (position 268 (EU numbering)); Gln is substituted for Arg at position 234 (position 355 (EU numbering)); and Glu is substituted for Gln at position 298 (position 419 (EU numbering)). Antibodies with improved stability, immunogenicity, and/or pharmacokinetics can be produced using such substitutions.
[0200] (2) In an IgG2 constant region (the amino acid sequence of SEQ ID NO: 24),
Ser is substituted for Cys at position 14 (position 131 (EU numbering)); Lys is substituted for Arg at position 16 (position 133 (EU numbering)); Ser is substituted for Cys at position 103 (position 220 (EU numbering)); Gly is substituted for Glu at position 20 (position 137 (EU numbering)); Gly is substituted for Ser at position 21 (position 138 (EU numbering)); Gln is substituted for His at position 147 (position 268 (EU numbering)); Gln is substituted for Arg at position 234 (position 355 (EU numbering)); Glu is substituted for Gln at position 298 (position 419 (EU numbering)); Ser is substituted for Ala at position 209 (position 330 (EU numbering)); Ser is substituted for Pro at position 210 (position 331 (EU numbering)); and Ala is substituted for Thr at position 218 (position 339 (EU numbering)). Antibodies with reduced Fc.gamma. receptor-binding activity can be produced using such substitutions.
[0201] (3) In an IgG2 constant region (the amino acid sequence of SEQ ID NO: 24), Gly at position 325 (position 446 (EU numbering)) and Lys at position 326 (position 447 (EU numbering)) are deleted. Antibodies with reduced C-terminal heterogeneity can be produced using such deletions.
[0202] (4) In an IgG2 constant region (the amino acid sequence of SEQ ID NO: 24), Val is substituted for Met at position 276 (position 397 (EU numbering)). Antibodies with improved stability under acidic conditions can be produced using such substitutions.
[0203] Meanwhile, amino acid alterations of light chain constant regions include, for example, those described below; but are not limited thereto. The alterations described below can reduce the heterogeneity of hinge region.
[0204] (1) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), amino acids are substituted or deleted so that it has at least one Cys at positions 102 to 106.
[0205] (2) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), amino acids are substituted or deleted so that it does not have Cys at position 107.
[0206] (3) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), amino acids are substituted or deleted so that it has at least one Cys at positions 102 to 106 but does not have Cys at position 107.
[0207] (4) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), at least one amino acid at positions 1 to 106 is deleted.
[0208] (5) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), at least one amino acid at positions 102 to 106 is deleted.
[0209] (6) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), the amino acid at position 105 is deleted.
[0210] (7) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), the amino acid at position 106 is deleted.
[0211] (8) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), Cys is substituted for at least one amino acid at positions 102 to 106.
[0212] (9) In a human .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 32), mouse .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 40), or rat .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 41), Cys is substituted for at least one amino acid at positions 102 to 106, and Cys at position 107 is deleted or substituted with another amino acid.
[0213] (10) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), amino acids are substituted or deleted so that it has at least one Cys at positions 99 to 103.
[0214] (11) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), amino acids are substituted or deleted so that it does not have Cys at position 104.
[0215] (12) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), amino acids are substituted or deleted so that it has at least one Cys at positions 99 to 103 but does not have Cys at position 104.
[0216] (13) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), at least one amino acid at positions 1 to 103 is deleted.
[0217] (14) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), at least one amino acid at positions 99 to 103 is deleted.
[0218] (15) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), the amino acid at position 102 is deleted.
[0219] (16) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), the amino acid at position 103 is deleted.
[0220] (17) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), Cys is substituted for at least one amino acid at positions 99 to 103.
[0221] (18) In a human .lamda. chain constant region (the amino acid sequence of SEQ ID NO: 37) or rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 42), Cys is substituted for at least one amino acid at positions 99 to 103, and Cys at position 104 is deleted or substituted with another amino acid.
[0222] (19) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), amino acids are substituted or deleted so that it has at least one Cys at potions 101 to 105.
[0223] (20) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), amino acids are substituted or deleted so that it does not have Cys at position 106.
[0224] (21) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), amino acids are substituted or deleted so that it has at least one Cys at positions 101 to 105 but does not have Cys at position 106.
[0225] (22) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), at least one amino acid at positions 1 to 105 is deleted.
[0226] (23) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), at least one amino acid at positions 101 to 105 is deleted.
[0227] (24) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), the amino acid at position 104 is deleted.
[0228] (25) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), the amino acid at position 105 is deleted.
[0229] (26) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), Cys is substituted for at least one amino acid at positions 101 to 105.
[0230] (27) In a rabbit .kappa. chain constant region (the amino acid sequence of SEQ ID NO: 43), Cys is substituted for at least one amino acid at positions 101 to 105, and Cys at position 106 is deleted or substituted with another amino acid.
[0231] Furthermore, the present invention provides methods for producing antibodies, which comprise the step of culturing host cells that comprise a vector introduced with a polynucleotide encoding an antibody heavy chain comprising the heavy chain constant region with the amino acid alterations of the present invention and/or a polynucleotide encoding an antibody light chain comprising the light chain constant region with the amino acid alterations of the present invention.
[0232] More specifically, the present invention provides methods for producing antibodies that comprise a heavy chain constant region with the amino acid alterations of the present invention and/or a light chain constant region with the amino acid alterations of the present invention, which comprise the steps of:
(a) culturing host cells that comprise a vector introduced with a polynucleotide encoding an antibody heavy chain comprising the heavy chain constant region with the amino acid alterations of the present invention and/or a polynucleotide encoding an antibody light chain comprising the light chain constant region with the amino acid alterations of the present invention; and (b) collecting the antibody heavy chain and/or light chain encoded by the gene.
[0233] Such amino acid alterations of the heavy chain constant region include the amino acid substitutions or deletions described above in (1) to (4), but are not limited thereto.
[0234] Such amino acid alterations of the light chain constant region include the amino acid substitutions or deletions described above (1) to (27), but are not limited thereto.
[0235] In the methods of the present invention for producing antibodies, first, the following DNAs are expressed: a DNA encoding an antibody heavy chain in which one or more amino acid residues in the constant region have been deleted or substituted with amino acids of interest, and/or a DNA encoding an antibody light chain in which one or more amino acid residues in the constant region have been deleted or substituted with amino acids of interest. A DNA encoding a heavy chain in which one or more amino acid residues in the constant region are deleted or substituted with amino acids of interest and/or a DNA encoding a light chain in which one or more amino acid residues in the constant region are deleted or substituted with amino acids of interest can be prepared, for example, by obtaining a DNA encoding the constant region of a wild type heavy chain and/or light chain, and introducing an appropriate substitution so that a codon encoding a particular amino acid in the constant region encodes an amino acid of interest.
[0236] Alternatively, a DNA encoding a heavy chain in which one or more amino acid residues in the constant region are deleted or substituted with amino acids of interest and/or a DNA encoding a light chain in which one or more amino acid residues in the constant region are deleted or substituted with amino acids of interest can also be prepared by designing and then chemically synthesizing a DNA encoding a protein in which one or more amino acid residues in the constant region of the wild type heavy chain are deleted or substituted with amino acids of interest.
[0237] The types of amino acid substitution and deletion include the substitutions and deletions described herein, but are not limited thereto.
[0238] Alternatively, a DNA encoding a heavy chain in which one or more amino acid residues in the constant region are deleted or substituted with amino acids of interest and/or a DNA encoding a light chain in which one or more amino acid residues in the constant region are deleted or substituted with amino acids of interest can also be prepared as a combination of partial DNAs. Such combinations of partial DNAs include, for example, the combination of a DNA encoding a variable region and a DNA encoding a constant region, and the combination of a DNA encoding an Fab region and a DNA encoding an Fc region, but are not limited thereto.
[0239] Methods for expressing the above-described DNAs include the methods described below. For example, a heavy chain expression vector is constructed by inserting a DNA encoding a heavy chain variable region into an expression vector along with a DNA encoding a heavy chain constant region. Likewise, a light chain expression vector is constructed by inserting a DNA encoding a light chain variable region into an expression vector along with a DNA encoding a light chain constant region. Alternatively, these heavy and light chain genes may be inserted into a single vector. Expression vectors include, for example, SV40 virus-based vectors, EB virus-based vectors, and BPV (papilloma virus)-based vectors, but are not limited thereto.
[0240] Host cells are co-transformed with an antibody expression vector constructed by the methods described above. Such host cells include the above-described cells such as Chinese hamster ovary (CHO) cells as well as microorganisms such as E. coli, yeast, and Bacillus subtilis, and plants and animals (Nature Biotechnology (2007) 25: 563-565; Nature Biotechnology (1998) 16: 773-777; Biochemical and Biophysical Research Communications (1999) 255: 444-450; Nature Biotechnology (2005) 23: 1159-1169; Journal of Virology (2001) 75: 2803-2809; Biochemical and Biophysical Research Communications (2003) 308: 94-100). Such host cells also include human embryonic kinder cancer cell-derived HEK298H cells. The transformation can be preferably achieved by using electroporation, the lipofectin method (R. W. Malone et al., Proc. Natl. Acad. Sci. USA (1989) 86: 6077; P. L. Felgner et al., Proc. Natl. Acad. Sci. USA (1987) 84: 7413), calcium phosphate method (F. L. Graham & A. J. van der Eb, Virology (1973) 52: 456-467), DEAE-Dextran method, and the like.
[0241] In the next step of antibody production, the expression products are collected. The expression products can be collected, for example, by culturing the transformants and then separating and purifying the antibody from the transformed cells or culture media. Separation and purification of antibodies can be achieved by an appropriate combination of methods such as centrifugation, ammonium sulfate fractionation, salting out, ultrafiltration, columns of 1 q, FcRn, Protein A, and Protein G, affinity chromatography, ion exchange chromatography, and gel filtration chromatography.
[0242] The present invention provides antibodies produced as described above. More specifically, the present invention relates to antibodies that can be produced by the following steps:
(a) expressing in host cells, DNAs encoding an antibody heavy chain which comprises variable and constant regions, and a light chain; and (b) collecting the antibodies expressed in (a).
[0243] A characteristic of the above-mentioned method is that the amino acid sequences of the constant regions of heavy and light chains are the above-mentioned constant regions provided by the present invention. In a preferred embodiment of the present invention, the heavy chain constant region consists of, for example, the amino acid sequence of SEQ ID NOs: 24 and 26 to 31. Linking a DNA consisting of the nucleotide sequence encoding this amino acid sequence with a DNA encoding the heavy chain variable region can produce a DNA encoding the antibody heavy chain. Meanwhile, the light chain constant region consists of, for example, the amino acid sequence of SEQ ID NOs: 32 to 34 and 37 to 39. Linking a DNA consisting of the nucleotide sequence encoding this amino acid sequence with a DNA encoding the light chain variable region can produce a DNA encoding the antibody light chain.
[0244] As described above, the variable regions that constitute the antibodies of the present invention may recognize any antigen. The variable regions that constitute the antibodies of the present invention are not particularly limited, and include, for example, those described below.
[0245] For an antibody that has IL6 receptor-neutralizing activity, the heavy-chain variable region may be, for example, variable regions having the CDR1, CDR2, and CDR3 of the humanized antibody heavy chain comprising the amino acid sequence of SEQ ID NO: 5. Meanwhile, the light-chain variable region may be variable regions having the CDR1, CDR2, and CDR3 of the humanized antibody light chain comprising the amino acid sequence of SEQ ID NO: 2.
[0246] For an antibody that has IL31 receptor-neutralizing activity, the heavy-chain variable region may be, for example, variable regions having the CDR1, CDR2, and CDR3 of the humanized antibody heavy chain comprising the amino acid sequence of SEQ ID NO: 13. Meanwhile, the light-chain variable region may be variable regions having the CDR1, CDR2, and CDR3 of the humanized antibody light chain comprising the amino acid sequence of SEQ ID NO: 12.
[0247] For an antibody that has RANKL-neutralizing activity, the heavy-chain variable region may be variable regions having the CDR1, CDR2, and CDR3 of the humanized antibody heavy chain comprising the amino acid sequence of SEQ ID NO: 17. Meanwhile, the light-chain variable region may be variable regions having the CDR1, CDR2, and CDR3 of the humanized antibody light chain comprising the amino acid sequence of SEQ ID NO: 16.
[0248] The amino acid sequences that constitute heavy chain and light chain variable regions may have one or more amino acid residue alterations and/or modifications, as long as they retain antigen-binding activity. In the present invention, when altering antibody variable regions, it is preferable to conserve the amino acid sequences of CDRs. The antigen-binding activity of the variable regions can be maintained by conserving the amino acid sequences of CDRs. The acceptable number of altered amino acid residues in a variable region is generally one to ten, for example, one to five, and preferably one or two amino acids.
[0249] For example, the modification of the N-terminal glutamine of a variable region into pyroglutamic acid by pyroglutamylation is a modification well known to those skilled in the art. Thus, when the heavy-chain N terminus is glutamine, the antibodies of the present invention comprise the variable regions in which the glutamine is modified to pyroglutamic acid.
[0250] The alterations of the present invention include substitutions, deletions, additions, and/or insertions of the above-described amino acids, and combinations thereof.
[0251] Furthermore, the present invention provides genes encoding antibody constant regions comprising the amino acid alterations of the present invention. Such genes encoding the antibody constant regions of the present invention may be any gene such as DNA or RNA.
[0252] The present invention also provides vectors carrying the genes. The type of vector can be appropriately selected by those skilled in the art depending on the host cells to be introduced with the vector. The vectors include, for example, those described above.
[0253] Furthermore, the present invention relates to host cells transformed with the vectors. Appropriate host cells can be selected by those skilled in the art. The host cells include, for example, those described above.
[0254] The present invention also relates to methods for producing the constant regions of the present invention, which comprise the steps of culturing the host cells and collecting the expressed constant regions of the present invention.
<Improvement of Antibody Function by Amino Acid Alterations of Heavy Chain Constant Region>
[0255] Furthermore, the present invention also relates to methods for improving antibody functions, which comprise the step of altering amino acids in the human IgG2 constant region of SEQ ID NO: 24. The present invention also relates to antibodies produced by methods comprising the above-described step. The improvement of antibody functions includes improvement of antibody stability, reduction of immunogenicity, and improvement of pharmacokinetics, but is not limited thereto. The methods of the present invention comprise the steps of:
(a) substituting another amino acid for Cys at position 14 (position 131 (EU numbering)) in SEQ ID NO: 24; (b) substituting another amino acid for Arg at position 16 (position 133 (EU numbering)) in SEQ ID NO: 24; (c) substituting another amino acid for Cys at position 103 (position 220 (EU numbering)) in SEQ ID NO: 24; (d) substituting another amino acid for Glu at position 20 (position 137 (EU numbering)) in SEQ ID NO: 24; (e) substituting another amino acid for Ser at position 21 (position 138 (EU numbering)) in SEQ ID NO: 24; (f) substituting another amino acid for His at position 147 (position 268 (EU numbering)) in SEQ ID NO: 24; (g) substituting another amino acid for Arg at position 234 (position 355 (EU numbering)) in SEQ ID NO: 24; and (h) substituting another amino acid for Gln at position 298 (position 419 (EU numbering)) in SEQ ID NO: 24.
[0256] The amino acids after substitution are not particularly limited; however,
Cys at position 14 is preferably substituted with Ser; Arg at position 16 is preferably substituted with Lys; Cys at position 103 is preferably substituted with Ser; Glu at position 20 is preferably substituted with Gly; Ser at position 21 is preferably substituted with Gly; His at position 147 is preferably substituted with Gln; Arg at position 234 is preferably substituted with Gln; and Gln at position 298 is preferably substituted with Glu.
[0257] The methods of the present invention may additionally comprise the step of altering (substituting, deleting, adding, and/or inserting) other amino acids, modifying amino acids, and such, as long as they comprise the steps described above. The methods for altering or modifying amino acids are not particularly limited, and include, for example, the above-described site-specific mutagenesis and the methods described in the Examples herein.
[0258] The methods of the present invention may also comprise, in addition to the above-described steps, the step of deleting Gly at position 325 (position 446 (EU numbering)) and Lys at position 326 (position 447 (EU numbering)) to reduce the C-terminal heterogeneity.
[0259] Furthermore, the present invention relates to methods for improving antibody functions, which comprise the step of altering amino acids of the human IgG2 constant region of SEQ ID NO: 24. The present invention also relates to antibodies produced by methods comprising the step. Such improvement of antibody functions includes, but is not limited to, improvement of antibody stability, reduction of immunogenicity, improvement of safety, and improvement of pharmacokinetics. The methods of the present invention comprise the steps of:
(a) substituting another amino acid for Cys at position 14 (position 131 (EU numbering)) in SEQ ID NO: 24; (b) substituting another amino acid for Arg at position 16 (position 133 (EU numbering)) in SEQ ID NO: 24; (c) substituting another amino acid for Cys at position 103 (position 220 (EU numbering)) in SEQ ID NO: 24; (d) substituting another amino acid for Glu at position 20 (position 137 (EU numbering)) in SEQ ID NO: 24; (e) substituting another amino acid for Ser at position 21 (position 138 (EU numbering)) in SEQ ID NO: 24; (f) substituting another amino acid for His at position 147 (position 268 (EU numbering)) in SEQ ID NO: 24; (g) substituting another amino acid for Arg at position 234 (position 355 (EU numbering)) in SEQ ID NO: 24; (h) substituting another amino acid for Gln at position 298 (position 419 (EU numbering)) in SEQ ID NO: 24; (i) substituting another amino acid for Ala at position 209 (position 330 (EU numbering)) in SEQ ID NO: 24; (j) substituting another amino acid for Pro at position 210 (position 331 (EU numbering)) in SEQ ID NO: 24; and (k) substituting another amino acid for Thr at position 218 (position 339 (EU numbering)) in SEQ ID NO: 24.
[0260] The amino acids after substitution are not particularly limited; however,
Cys at position 14 is preferably substituted with Ser; Arg at position 16 is preferably substituted with Lys; Cys at position 103 is preferably substituted with Ser; Glu at position 20 is preferably substituted with Gly; Ser at position 21 is preferably substituted with Gly; His at position 147 is preferably substituted with Gln; Arg at position 234 is preferably substituted with Gln; Gln at position 298 is preferably substituted with Glu; Ala at position 209 is preferably substituted with Ser; Pro at position 210 is preferably substituted with Ser; and Thr at position 218 is preferably substituted with Ala.
[0261] The methods of the present invention may additionally comprise the step of altering (substituting, deleting, adding, and/or inserting) other amino acids, modifying amino acids, and such, as long as they comprise the above-described steps. The methods for altering or modifying amino acids are not particularly limited, and include, for example, the above-described site-specific mutagenesis and the methods described in the Examples herein.
[0262] The methods of the present invention may also comprise, in addition to the above-described steps, the step of deleting Gly at position 325 (position 446 (EU numbering)) and Lys at position 326 (position 447 (EU numbering)) to reduce the C-terminal heterogeneity.
[0263] Furthermore, the present invention relates to methods for improving the blood kinetics (pharmacokinetics) of an antibody by controlling (or changing) the disulfide bond pattern in antibody constant regions. The antibody constant regions are not particularly limited; however, it is preferable to control the pattern of disulfide bonds between an antibody light chain constant region (.kappa. chain constant region or .lamda. chain constant region) and IgG2 constant region.
[0264] Specifically, the present invention relates to methods for improving pharmacokinetics of an antibody, which comprise the step of allowing specific formation of a disulfide bond between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy-chain constant region.
[0265] In the above-described step, it is preferable to avoid formation of an additional disulfide bond between Cys in the light chain C-terminal region and Cys at position 220 (EU numbering) in the heavy-chain constant region.
[0266] In the methods of the present invention, it is not necessary that every antibody forms a disulfide bond between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy-chain constant region. For example, it is acceptable that 80% or more, preferably 90% or more, more preferably 95% or more, and still more preferably 99% or more of the antibody forms a disulfide bond between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy-chain constant region.
[0267] The step of allowing formation of a disulfide bond between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy-chain constant region may be achieved by any method, for example, by substituting another amino acid for Cys at position 220 (EU numbering) in the heavy chain (position 103 in SEQ ID NO: 24). As a result of the substitution of another amino acid for Cys at position 220 (EU numbering) in the heavy chain, a disulfide bond is formed between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy-chain constant region (position 102 in SEQ ID NO: 24), instead of between Cys in the light chain C-terminal region and Cys at position 220 (EU numbering) in the heavy-chain constant region.
[0268] In the present invention, not only Cys at position 220 (EU numbering) but also Cys at position 131 (EU numbering) may be substituted with another amino acid. The amino acids after substitution are not particularly limited, and include, for example, Ser.
[0269] Herein, for example, in the case of a human .kappa. chain constant region, Cys in the light chain C-terminal region is typically Cys in the region of positions 102 to 106 (for example, positions 102 to 106 in the human .kappa. chain constant region of SEQ ID NO: 32), preferably Cys in the region of positions 104 to 106. Alternatively, for example, in the case of a human .lamda. chain constant region, the Cys is typically Cys in the region of positions 99 to 105 (for example, positions 99 to 105 in the human .lamda. chain constant region of SEQ ID NO: 37), preferably Cys in the region of positions 102 to 104.
[0270] An IgG2 constant region used in the methods of the present invention may comprise one or more amino acid deletions, substitutions, additions, and/or insertions (for example, 20 amino acids or less, or 10 amino acids or less) in the amino acid sequence of SEQ ID NO: 24. An IgG2 constant region used in the methods of the present invention may comprise alterations specified by the description herein and combinations thereof.
[0271] Meanwhile, a human .kappa. chain constant region used in the methods of the present invention may comprise one or more amino acid deletions, substitutions, additions, and/or insertions (for example, 20 amino acids or less, or 10 amino acids or less) in the amino acid sequence of SEQ ID NO: 32.
[0272] On the other hand, a human .lamda. chain constant region used in the methods of the present invention may comprise one or more amino acid deletions, substitutions, additions, and/or insertions (for example, 20 amino acids or less, or 10 amino acids or less) in the amino acid sequence of SEQ ID NO: 37.
[0273] Furthermore, human .kappa. and .lamda. chain constant regions used in the methods of the present invention may comprise alterations specified by the description herein and combinations thereof.
[0274] Antibodies in which a disulfide bond between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy-chain constant region has been formed are demonstrated to be superior in pharmacokinetics as compared to antibodies having a disulfide bond between Cys in the light chain C-terminal region and Cys at position 220 (EU numbering) in the heavy-chain constant region. Thus, the present invention provides antibodies with improved pharmacokinetics by allowing formation of a disulfide bond between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy-chain constant region. The methods of the present invention are useful in producing antibodies that are superior in pharmacokinetics and excellent as pharmaceuticals.
<Functional Improvement of Antibodies by Altering Amino Acids in Light Chain Constant Regions>
[0275] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the step of introducing at least one Cys at positions 102 to 106 into the human .kappa. constant region of SEQ ID NO: 32, the mouse .kappa. chain constant region of SEQ ID NO: 40, or the rat .kappa. chain constant region of SEQ ID NO: 41. The present invention also relates to antibodies produced by methods comprising the above-described step.
[0276] Herein, "introducing at least one Cys at positions 102 to 106" means achieving the state where there is at least one Cys at positions 102 to 106 in a human .kappa. chain constant region.
[0277] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the step of removing (deleting) Cys at position 107 from the human .kappa. constant region of SEQ ID NO: 32, the mouse .kappa. chain constant region of SEQ ID NO: 40, or the rat .kappa. chain constant region of SEQ ID NO: 41. The present invention also relates to antibodies produced by methods comprising the above-described step.
[0278] Herein, "removing Cys at position 107" means achieving the state where there is no Cys at position 107 in a human .kappa. chain constant region.
[0279] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region in the human .kappa. constant region of SEQ ID NO: 32, the mouse .kappa. chain constant region of SEQ ID NO: 40, or the rat .kappa. chain constant region of SEQ ID NO: 41, which comprise the steps of:
(a) introducing at least one Cys at positions 102 to 106; and (b) removing (deleting) Cys at position 107. The present invention also relates to antibodies produced by methods comprising the above-described steps.
[0280] In the present invention, the steps of introducing at least one Cys at positions 102 to 106 and the step of removing Cys at position 107 may be carried out in a single step.
[0281] The number of Cys to be introduced in the step of introducing at least one Cys at positions 102 to 106 is not particularly limited; however, the number is five or less, preferably three or less, more preferably two or one, and still more preferably one.
[0282] The position of Cys introduction is not particularly limited; however, the position is preferably 104, 105, or 106, more preferably 105 or 106, and particularly preferably 106.
[0283] Specifically, the step of introducing at least one Cys at positions 102 to 106 includes, for example, the steps described below. Such steps may be used in combination.
[0284] The step of inserting at least one Cys at positions 102 to 106.
[0285] The step of substituting Cys for at least one amino acid at positions 102 to 106.
[0286] The step of deleting one to five amino acids at positions 1 to 106.
[0287] Specifically, the step of removing Cys at position 107 includes, for example, the following steps.
[0288] The step of deleting Cys at position 107.
[0289] The step of substituting another amino acid for Cys at position 107.
[0290] The step of inserting another amino acid at position 107.
[0291] The step of relocating Cys from position 107 to another position by deleting at least one amino acid at positions 1 to 106.
[0292] In a preferred embodiment, the methods of the present invention (i.e., methods for reducing the heterogeneity of hinge region and methods for enhancing FcRn binding, which comprise the steps of:
(a) introducing at least one Cys at positions 102 to 106, and (b) deleting Cys at position 107 in the human .kappa. constant region of SEQ ID NO: 32, mouse .kappa. chain constant region of SEQ ID NO: 40, or rat .kappa. chain constant region of SEQ ID NO: 41) include, for example, the step of deleting one to five amino acids at positions 1 to 106 from the human .kappa. chain constant region. When one to five amino acids are deleted at positions 1 to 106 from the human .kappa. chain constant region, Cys at position 107 is relocated to one of positions 102 to 106. Thus, the steps of introducing at least one Cys at positions 102 to 106 and deleting Cys at position 107 can be achieved simultaneously by this method.
[0293] The position of amino acid deletion is not particularly limited; however, the human .kappa. chain constant region preferably comprises deletion of at least one amino acid at positions 102 to 106, more preferably deletion of the amino acid at position 105 or 106.
[0294] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0295] Such preferred steps include, but are not limited to, for example, the following steps.
[0296] The step of deleting the amino acid at position 105.
[0297] The step of deleting the amino acid at position 106.
[0298] In another preferred embodiment, the methods of the present invention include methods comprising the steps of:
(a) substituting Cys for at least one amino acid at positions 102 to 106; and (b) deleting Cys at position 107 or substituting another amino acid for Cys at position 107. The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one. The position of Cys substitution is not particularly limited; however, the position is preferably 105 or 106.
[0299] Specifically, in the case of a human .kappa. chain constant region, such steps include, for example, the steps of:
(a) substituting Cys for Gly at position 105 or Glu at position 106; and (b) deleting Cys at position 107, or substituting another amino acid for Cys at position 107.
[0300] Alternatively, in the case of a mouse or rat .kappa. chain constant region, such steps include the steps of:
(a) substituting Cys for Asn at position 105 or Glu at position 106; and (b) deleting Cys at position 107 or substituting another amino acid for Cys at position 107.
[0301] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the step of introducing at least one Cys at positions 99 to 103 in the human .lamda. constant region of SEQ ID NO: 37 or the rabbit .kappa. chain constant region of SEQ ID NO: 42. The present invention also relates to antibodies produced by methods comprising the above-described steps.
[0302] Herein, "introducing at least one Cys at positions 99 to 103" means achieving the state where there is at least one Cys at positions 99 to 103 in the human .lamda. chain constant region.
[0303] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the step of removing (deleting) Cys at position 104 in the human .lamda. constant region of SEQ ID NO: 37 or rabbit .kappa. chain constant region of SEQ ID NO: 42. The present invention also relates to antibodies produced by methods comprising the above-described steps.
[0304] Herein, "removing Cys at position 104" means achieving the state where there is no Cys at position 104 in the human .lamda. chain constant region.
[0305] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the steps of:
(a) introducing at least one Cys at positions 99 to 103, and (b) removing Cys at position 104 in the human .lamda. constant region of SEQ ID NO: 37 or rabbit .kappa. chain constant region of SEQ ID NO: 42.
[0306] In the present invention, the steps of introducing at least one Cys at positions 99 to 103 and removing Cys at position 104 may be achieved in a single step.
[0307] The number of Cys introduced in the step of introducing at least one Cys at positions 99 to 103 is not particularly limited; however, the number is five or less, preferably three or less, more preferably two or one, and still more preferably one.
[0308] The position of Cys introduction is not particularly limited; however, the position is preferably 101, 102, or 103, more preferably 102 or 103, and particularly preferably 103.
[0309] Specifically, the step of introducing at least one Cys at positions 99 to 103 includes, for example, those described below. The steps described below may be used in combination.
[0310] The step of inserting at least one Cys at positions 99 to 103.
[0311] The step of substituting Cys for at least one amino acid at positions 99 to 103.
[0312] The step of deleting one to five amino acids at positions 1 to 103.
[0313] Specifically, the step of removing Cys at position 104 includes, but is not limited to, for example, those described below.
[0314] The step of deleting Cys at position 104.
[0315] The step of substituting another amino acid for Cys at position 104.
[0316] The step of inserting another amino acid at position 104.
[0317] The step of relocating Cys from position 105 to another position by deleting at least one amino acid at positions 1 to 103.
[0318] In a preferred embodiment, the methods of the present invention (i.e., methods for reducing the heterogeneity of hinge region, which comprise the steps of:
(a) introducing at least one Cys at positions 99 to 103, and (b) removing Cys at position 104 in the human .lamda. constant region of SEQ ID NO: 37 or rabbit .kappa. chain constant region of SEQ ID NO: 42) include, for example, methods comprising the step of deleting one to five amino acids at positions 1 to 103 in the human .lamda. chain constant region. When one to five amino acids are deleted at positions 1 to 103 in the human .kappa. chain constant region, Cys at position 104 is relocated to one of positions 99 to 103. Thus, the steps of introducing at least one Cys at positions 99 to 103 and deleting Cys at position 104 can be achieved simultaneously.
[0319] The position of amino acid deletion is not particularly limited; however, a human .lamda. chain constant region preferably comprises deletion of at least one amino acid at positions 99 to 103, more preferably deletion of the amino acid at position 102 or 103.
[0320] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0321] Such preferred steps include, but are not limited to, for example, those described below.
[0322] The step of deleting the amino acid at position 102.
[0323] The step of deleting the amino acid at position 103.
[0324] In another preferred embodiment, the methods of the present invention include methods comprising the steps of:
(a) substituting Cys for at least one amino acid at positions 99 to 103; and (b) deleting Cys at position 104 or substituting another amino acid for Cys at position 104.
[0325] The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one. The position of Cys substitution is not particularly limited; however, the position is preferably 102 or 103.
[0326] Specifically, in the case of a human .lamda. chain constant region, the steps include, but are not limited to, for example, the steps of:
(a) substituting Cys for Thr at position 102 or Glu at position 103; and (b) deleting Cys at position 104, or substituting another amino acid for Cys at position 104.
[0327] Alternatively, in the rabbit .kappa. chain constant region of SEQ ID NO: 42, the steps include, but are not limited to, the steps of:
(a) substituting Cys for Gly at position 102 or Asp at position 103; and (b) deleting Cys at position 104, or substituting another amino acid for Cys at position 104.
[0328] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the step of introducing at least one Cys at positions 101 to 105 in the rabbit .kappa. chain constant region of SEQ ID NO: 43. The present invention also relates to antibodies produced by methods comprising the above-described steps.
[0329] Herein, "introducing at least one Cys at positions 101 to 105" means achieving the state where there is at least one Cys at positions 101 to 105 in the rat .kappa. chain constant region.
[0330] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the step of removing (deleting) Cys at position 106 in the rabbit chain constant region of SEQ ID NO: 43. The present invention also relates to antibodies produced by methods comprising the above-described steps.
[0331] Herein, "removing Cys at position 106" means achieving the state where there is no Cys at position 106 in the rabbit .kappa. chain constant region.
[0332] Furthermore, the present invention relates to methods for reducing the heterogeneity of hinge region, which comprise the steps of:
(a) introducing at least one Cys at positions 101 to 105, and (b) removing Cys at position 106 in the rabbit .kappa. chain constant region of SEQ ID NO: 43.
[0333] In the present invention, the steps of introducing at least one Cys at positions 101 to 105 and removing Cys at position 106 may be achieved in a single step.
[0334] The number of Cys introduced in the step of introducing at least one Cys at positions 101 to 105 is not particularly limited; however, the number is five or less, preferably three or less, more preferably two or one, and still more preferably one.
[0335] The position of Cys introduction is not particularly limited; however, the position is preferably 103, 104, or 105, more preferably 104 or 105, and particularly preferably 105.
[0336] Specifically, the step of introducing at least one Cys at positions 101 to 105 includes, for example, those described below. The steps described below may be used in combination.
[0337] The step of inserting at least one Cys at positions 101 to 105.
[0338] The step of substituting Cys for at least one amino acid at positions 101 to 105.
[0339] The step of deleting one to five amino acids at positions 1 to 105.
[0340] Specifically, the step of removing Cys at position 106 includes, but is not limited to, for example, those described below.
[0341] The step of deleting Cys at position 106.
[0342] The step of substituting another amino acid for Cys at position 106.
[0343] The step of inserting another amino acid at position 106.
[0344] The step of relocating Cys from position 106 to another position by deleting at least one amino acid at positions 1 to 105.
[0345] In a preferred embodiment, the methods of the present invention (i.e., methods for reducing the heterogeneity of hinge region, which comprise the steps of:
(a) introducing at least one Cys at positions 101 to 105, and (b) removing Cys at position 106 in the rabbit .kappa. chain constant region of SEQ ID NO: 43) include, for example, methods comprising the step of deleting one to five amino acids at positions 1 to 105 from the rabbit .kappa. chain constant region. When one to five amino acids are deleted at positions 1 to 105, Cys at position 106 is relocated to one of positions 101 to 105. Thus, the steps of introducing at least one Cys at positions 101 to 105 and deleting Cys at position 106 can be achieved simultaneously.
[0346] The position of amino acid deletion is not particularly limited; however, a human chain constant region preferably comprises deletion of at least one amino acid at positions 101 to 105, more preferably deletion of the amino acid at position 104 or 105.
[0347] The number of deleted amino acids is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one.
[0348] Such preferred steps include, but are not limited to, for example, those described below.
[0349] The step of deleting the amino acid at position 104.
[0350] The step of deleting the amino acid at position 105.
[0351] In another preferred embodiment, the methods of the present invention include methods comprising the steps of:
(a) substituting Cys for at least one amino acid at positions 101 to 105; and (b) deleting Cys at position 106, or substituting another amino acid for Cys at position 106. The number of amino acids substituted with Cys is not particularly limited; however, the number is typically one to five, preferably one to three, more preferably one or two, and still more preferably one. The position of Cys substitution is not particularly limited; however, the position is preferably 104 or 105.
[0352] Specifically, such steps include, but are not limited to, for example, the steps of:
(a) substituting Cys for Gly at position 104 or Asp at position 105; and (b) deleting Cys at position 106, or substituting another amino acid for Cys at position 106.
[0353] As long as the methods of the present invention comprise the above-described steps, they may additionally comprise other steps, for example, the step of altering (substituting, deleting, adding, and/or inserting) other amino acids or modifying amino acids.
<Pharmaceutical Compositions Comprising Antibodies>
[0354] The present invention provides pharmaceutical compositions comprising the antibodies or constant regions of the present invention.
[0355] The present invention also provides antibody pharmaceutical compositions in which the ratio of antibody having a disulfide bond between Cys in the light chain C-terminal region and Cys at position 219 (EU numbering) in the heavy chain constant region is 80% or more, preferably 90% or more, more preferably 95% or more, and still more preferably 99% or more.
[0356] The pharmaceutical compositions of the present invention can be formulated, in addition to the antibodies or constant regions, with pharmaceutically acceptable carriers by known methods. For example, the compositions can be used parenterally, when the antibodies are formulated in a sterile solution or suspension for injection using water or any other pharmaceutically acceptable liquid. For example, the compositions can be formulated by appropriately combining the antibodies with pharmaceutically acceptable carriers or media, specifically, sterile water or physiological saline, vegetable oils, emulsifiers, suspending agents, surfactants, stabilizers, flavoring agents, excipients, vehicles, preservatives, binding agents, and such, by mixing them at a unit dose and form required by generally accepted pharmaceutical implementations. The content of the active ingredient in such a formulation is adjusted so that an appropriate dose within the required range can be obtained.
[0357] Sterile compositions for injection can be formulated using vehicles such as distilled water for injection, according to standard protocols.
[0358] Aqueous solutions used for injection include, for example, physiological saline and isotonic solutions containing glucose or other adjuvants such as D-sorbitol, D-mannose, D-mannitol, and sodium chloride. These can be used in conjunction with suitable solubilizers such as alcohol, specifically ethanol, polyalcohols such as propylene glycol and polyethylene glycol, and non-ionic surfactants such as Polysorbate 80.TM. and HCO-50.
[0359] Oils include sesame oils and soybean oils, and can be combined with solubilizers such as benzyl benzoate or benzyl alcohol. These may also be formulated with buffers, for example, phosphate buffers or sodium acetate buffers; analgesics, for example, procaine hydrochloride; stabilizers, for example, benzyl alcohol or phenol; or antioxidants. The prepared injections are typically aliquoted into appropriate ampules.
[0360] The administration is preferably carried out parenterally, and specifically includes injection, intranasal administration, intrapulmonary administration, and percutaneous administration. For example, injections can be administered systemically or locally by intravenous injection, intramuscular injection, intraperitoneal injection, or subcutaneous injection.
[0361] Furthermore, the method of administration can be appropriately selected according to the age and symptoms of the patient. A single dose of the pharmaceutical composition containing an antibody or a polynucleotide encoding an antibody can be selected, for example, from the range of 0.0001 to 1,000 mg per kg of body weight. Alternatively, the dose may be, for example, in the range of 0.001 to 100,000 mg/patient. However, the dose is not limited to these values. The dose and method of administration vary depending on the patient's body weight, age, and symptoms, and can be appropriately selected by those skilled in the art.
[0362] As used herein, the three-letter and single-letter codes for respective amino acids are as follows:
Alanine: Ala (A)
Arginine: Arg (R)
Asparagine: Asn (N)
[0363] Aspartic acid: Asp (D)
Cysteine: Cys (C)
Glutamine: Gln (Q)
[0364] Glutamic acid: Glu (E)
Glycine: Gly (G)
Histidine: His (H)
Isoleucine: Ile (I)
Leucine: Leu (L)
Lysine: Lys (K)
Methionine: Met (M)
Phenylalanine: Phe (F)
Proline: Pro (P)
Serine: Ser (S)
Threonine: Thr (T)
Tryptophan: Trp (W)
Tyrosine: Tyr (Y)
Valine: Val (V)
[0365] All prior art documents cited herein are incorporated by reference in their entirety.
EXAMPLES
[Example 1] Improvement of C-Terminal Heterogeneities of IgG Molecules
[0366] Construction of an expression vector for H-chain C-terminal .DELTA.GK antibody
[0367] Heterogeneities of the C-terminal sequence of the IgG antibody H chain that have been reported are deletion of the C-terminal amino acid lysine residue, and amidation of the C-terminal carboxyl group due to deletion of both of the two C-terminal amino acids, glycine and lysine residues (Anal Biochem. 2007 Jan. 1; 360(1): 75-83). In TOCILIZUMAB which is an anti-IL-6 receptor antibody, the main component is a sequence in which the C-terminal amino acid lysine present on the nucleotide sequence is deleted by post-translational modification, but an accessory component with remnant lysine and an accessory component with an amidated C-terminal carboxyl group produced by deletion of both glycine and lysine are also present as heterogeneities. It is not easy to manufacture such an antibody as a pharmaceutical in a large scale, while maintaining differences of objective substance/related substance-related heterogeneity between productions, which will lead to increased cost. Thus, single substances are desirable as much as possible, and in developing antibodies as pharmaceuticals, such heterogeneities are desirably reduced. Therefore, in terms of development as pharmaceuticals, absence of heterogeneities of the H-chain C terminal is desirable.
[0368] Thus, the C-terminal amino acids were altered to reduce the C-terminal heterogeneity. Specifically, the present inventors altered the nucleotide sequence of wild type IgG1 to delete the C-terminal lysine and glycine from the H-chain constant region of the IgG1, and assessed whether the amidation of the C-terminal amino group due to deletion of the two C-terminal amino acids glycine and lysine could be suppressed.
[0369] According to the method of Reference Example 1, TOCILIZUMAB (hereinafter abbreviated as IL6R H0/L0-IgG1) consisting of H0-IgG1 (amino acid SEQ ID NO: 1) as an H chain and L0-k0 (amino acid SEQ ID NO: 2) as an L chain was prepared. Furthermore, the nucleotide sequence of the H chain encoding Lys at position 447 and/or Gly at position 446 (EU numbering) was converted into a stop codon. Thus, expression vectors for antibody H chain H0-IgG1.DELTA.K (amino acid SEQ ID NO: 3) engineered to lack the C-terminal amino acid lysine (position 447 (EU numbering)) and antibody H chain H0-IgG1.DELTA.GK (amino acid SEQ ID NO: 4) engineered to lack the two C-terminal amino acids glycine and lysine (positions 446 and 447 (EU numbering), respectively) were constructed.
[0370] IL6R H0-IgG1/L0-k0 consisting of H0-IgG1 (amino acid SEQ ID NO: 1) as the H chain and L0-k0 (amino acid SEQ ID NO: 2) as the L chain, IL6R H0-IgG1.DELTA.K/L0-k0 consisting of H0-IgG1.DELTA.K (amino acid SEQ ID NO: 3) as the H chain and L0-k0 (amino acid SEQ ID NO: 2) as the L chain, and IL6R H0-IgG1.DELTA.GK/L0-k0 consisting of H0-IgG1.DELTA.GK-k0 (amino acid SEQ ID NO: 4) as the H chain and L0-k0 (amino acid SEQ ID NO: 2) as the L chain were expressed and purified by the method described in Reference Example 1.
Cation Exchange Chromatographic Analysis of the H-Chain C-Terminal .DELTA.GK Antibody
[0371] Heterogeneity of the purified antibodies was evaluated by performing cation exchange chromatography. ProPac WCX-10, 4.times.250 mm (Dionex) was used for the column, 25 mmol/L MES/NaOH, pH 6.1 was used as mobile phase A, 25 mmol/L MES/NaOH, 250 mmol/L NaCl, pH 6.1 was used as mobile phase B, and the chromatography was performed using appropriate flow and gradient. The results of performing cation exchange chromatographic evaluations on the purified IL6R H0-IgG1/L0-k0, IL6R H0-IgG1.DELTA.K/L0-k0, and IL6R H0-IgG1.DELTA.GK/L0-k0 are shown in FIG. 1.
[0372] From the results, it was discovered that heterogeneity of the C-terminal amino acid can be decreased for the first time by deleting both the C-terminal lysine and glycine of the H-chain constant region, not only the C-terminal lysine of the H-chain constant region, from the nucleotide sequence. In the human antibody constant regions of IgG1, IgG2, and IgG4, the C-terminal sequence is lysine at position 447 and glycine at position 446 in the EU numbering (see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) in all cases; therefore, the method of reducing C-terminal amino acid heterogeneity discovered in the present examination was considered to be applicable to the IgG2 constant region and IgG4 constant region, or their modified forms.
[Example 2] Novel Constant Regions with Reduced Heterogeneity, which Retain the Stability of Natural IgG2
Heterogeneity of Natural IgG1 and Natural IgG2
[0373] For antibody pharmaceuticals against cancer such as those that kill target cells with effector functions and such, IgG1 constant region (isotype) having effector function is preferred. However, for antibody pharmaceuticals that neutralize the functions of a target antigen or antibody pharmaceuticals that bind to target cells but do not kill them, binding to Fc.gamma. receptors is not preferred.
[0374] As methods for decreasing the binding to Fc.gamma. receptors, the method of changing the IgG antibody isotype from IgG1 to IgG2 or IgG4 has been considered (Ann Hematol. 1998 June; 76(6): 231-48), and from the viewpoint of binding to Fc.gamma. receptor I and pharmacokinetics of each isotype, IgG2 was considered to be more desirable than IgG4 (Nat Biotechnol. 2007 December; 25(12): 1369-72). On the other hand, when developing antibodies into pharmaceuticals, physicochemical properties of the proteins, particularly homogeneity and stability are extremely important. The IgG2 isotype has been reported to have a very large degree of heterogeneity caused by disulfide bond linkage differences in the hinge region (J Biol Chem. 2008 Jun. 6; 283(23): 16194-205; J Biol Chem. 2008 Jun. 6; 283(23): 16206-15; Biochemistry 2008 Jul. 15; 47(28): 7496-508).
[0375] Accordingly, IL6R H0-IgG1/L0-k0 having the constant regions of natural IgG1 and IL6R H0-IgG2/L0-k0 having the constant regions of natural IgG2 were actually produced and heterogeneity evaluations were carried out for both of them. IL6R H0-IgG1/L0-k0 which was produced in Example 1 consisting of IL6R H0-IgG1 (amino acid SEQ ID NO: 1) as the H chain and IL6R L0-k0 (amino acid SEQ ID NO: 2) as the L chain and IL6R H0-IgG2/L0-k0 consisting of IL6R H0-IgG2 (amino acid SEQ ID NO: 5) as the H chain in which the H-chain constant region was converted into IgG2 and IL6R L0-k0 (amino acid SEQ ID NO: 2) as the L chain were expressed and purified by the method described in Reference Example 1.
[0376] Evaluation by cation exchange chromatography was carried out as the method for evaluating the heterogeneity caused by disulfide bonds in IL6R H0-IgG1/L0-k0 having the constant regions of natural IgG1 and IL6R H0-IgG2/L0-k0 having the constant regions of natural IgG2. ProPac WCX-10 (Dionex) was used for the column, 20 mM sodium acetate, pH 5.0 was used as mobile phase A, 20 mM sodium acetate, 1M NaCl, pH 5.0 was used as mobile phase B, and the chromatography was performed using appropriate flow and gradient. As a result, as shown in FIG. 2, IL6R H0-IgG2/L0-k0 having the constant regions of natural IgG2 showed multiple peaks and it was found to have markedly high heterogeneity compared to IL6R H0-IgG1/L0-k0 having the constant regions of natural IgG1 showing only almost single main peak.
[0377] The details of the structure around the hinge region of an IgG-type antibody are shown in FIG. 3. In IgG antibodies, H and L chains (or two H chains) form disulfide bonds around the hinge region. This pattern of disulfide bonds differ depending on the isotypes of the IgG-type antibodies as indicated below. It is considered that since disulfide bonds in the hinge region of natural IgG1 have a single pattern such as that shown in FIG. 4, heterogeneity caused by disulfide bonds does not exist and it was eluted as a nearly single main peak in cation exchange chromatography. In contrast, as shown in FIG. 3, regarding the disulfide bonds in the hinge region of natural IgG2, natural IgG2 has two cysteines in the hinge region (positions 219 and 220 (EU numbering)), and cysteines exist adjacent to these two cysteines of the hinge region, which are cysteine at position 131 (EU numbering), present in the H-chain CH1 domain, cysteine at the L-chain C terminus, and two cysteines of the corresponding hinge region of the H chain of the dimerization partner. Therefore, around the hinge region of IgG2, there are a total of eight neighboring cysteines when an H2L2 assembly is formed. This leads to the presence of a variety of heterogeneity due to disulfide bond linkage differences in natural IgG2, and the heterogeneity is considered to be remarkably high.
[0378] It is not easy to manufacture as a pharmaceutical in large-scale while maintaining the differences of objective substance/related substance-related heterogeneity between productions (the differences originate from these disulfide bond linkage differences), and this leads to increased cost. Thus, single substances are desirable as much as possible. Therefore, in developing antibodies of the IgG2 isotype into pharmaceuticals, it is desirable to reduce heterogeneity derived from disulfide bonds without decreasing stability. In fact, it is reported in US 20060194280 (A1) that a variety of heterogeneous peaks derived from disulfide bonds were observed for natural IgG2 in ion exchange chromatographic analysis, and biological activities were also reported to be different among these peaks. As a method for unifying these heterogeneous peaks, US 20060194280 (A1) reports refolding in the purification steps, but since using these steps in the production will be costly and complicated, preferably, unifying the heterogeneous peaks by producing an IgG2 variant in which disulfide bonds will be formed in a single pattern by means of amino acid substitutions was considered desirable. However, to date, there has been no report on IgG2 variants that will form disulfide bonds in a single pattern, or IgG2 variants that have reduced heterogeneity due to disulfide bonds without having decreased stability.
Influence of the Disulfide Bond Patterns Between H Chain and L Chain on the Stability
[0379] In natural IgG1, cysteine at position 220 (EU numbering) in the sequence of IgG1 H-chain constant region (amino acid SEQ ID NO: 23) bond to cysteine at position 214 (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) in the L chain (Nat Biotechnol. 2007 December; 25(12): 1369-72; Anal Chem. 2008 Mar. 15; 80(6): 2001-9) by a disulfide bond. Meanwhile, in the natural IgG4, cysteine at position 131 (EU numbering) in the sequence of IgG4 H-chain constant region (amino acid SEQ ID NO: 25) bond to cysteine at position 214 (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) by a disulfide bond in the L chain (Nat Biotechnol. 2007 December; 25(12): 1369-72; Protein Sci. 1997 February; 6(2): 407-15). It has been reported that as described above, the pattern of disulfide bonds between the H chain and L chain is different between natural IgG1 and IgG4. The disulfide bond patterns of the natural IgG1 and IgG4 are shown in FIG. 5. However, to date, there has been no report on the effect of the pattern of disulfide bonds between H chain and L chain on stability.
[0380] As prepared in Example 1, IL6R H0-IgG1/L0-k0 which consists of IL6R H0-IgG1 (amino acid SEQ ID NO: 1) as H chain and IL6R L0-k0 (amino acid SEQ ID NO: 2) as L chain, and IL6R H0-IgG4/L0-k0 which consists of IL6R H0-IgG4 (amino acid SEQ ID NO: 6) as an H-chain constant region resulting from IgG4 conversion and IL6R L0-k0 (amino acid SEQ ID NO: 2) as L chain, were expressed and purified by the method described in Reference Example 1.
[0381] To assess the stability, the midpoint of thermal denaturation (Tm value) was determined by differential scanning calorimetry (DSC) (N-DSCII, calorimetry Science Corporation). The midpoint temperature of thermal denaturation (Tm value) is a stability indicator. Thus, higher thermal denaturation midpoints (Tm values) are preferable for producing stable preparations as pharmaceuticals (J Pharm Sci. 2008 April; 97(4): 1414-26). Purified IL6R H0-IgG1/L0-k0 and IL6R H0-IgG4/L0-k0 were dialyzed against a solution (pH 6.0) containing 20 mM sodium acetate and 150 mM NaCl (EasySEP, TOMY). DSC measurements were carried out at a heating rate of 1.degree. C./min in a range of 40 to 100.degree. C., and at a protein concentration of about 0.1 mg/ml. The Tm values of the Fab domains (as listed in Table 1) are calculated based on the denaturation curves obtained by DSC. These data demonstrate that the Tm value of the IgG1 Fab domain is higher than that of the IgG4 Fab domain. The difference in the Tm value is speculated to be due to difference in the pattern of disulfide bonds between H chain and L chain. The thermal stability was demonstrated to be greatly reduced when the pattern of disulfide bonds between H chain and L chain is disulfide bond between cysteine at position 131 (EU numbering) in the H chain and cysteine at position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242), as compared to disulfide bond between cysteine at position 220 (EU numbering) in the H chain and cysteine at position 214 (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) in the L chain.
TABLE-US-00001 TABLE 1 Tm/.degree. C. OF Fab IL6R H0-IgG1/L0-k0 95.degree. C. IL6R H0-IgG4/L0-k0 88.degree. C.
Production of Various Types of Natural IgG2 Variants
[0382] For methods for reducing the heterogeneity as a result of difference in disulfide bonds of natural IgG2, one can consider the method of substituting serine for cysteine only at position 219 (EU numbering) in the H-chain hinge region or method of substituting serine for cysteine only at position 220 (Biochemistry. 2008 Jul. 15; 47(28): 7496-508). Specifically, such H-chain constant regions include SC (SEQ ID NO: 26) and CS (SEQ ID NO: 27). SC is an H-chain constant region having substitution of serine for cysteine at position 219 (EU numbering) in the H-chain constant region sequence (amino acid SEQ ID NO: 24) of natural IgG2, while CS is an H-chain constant region having substitution of cysteine at position 220 (EU numbering). However, as shown in FIG. 6, disulfide bonds pattern of these H-chain constant regions of SC and CS are not a single but plural like natural IgG2. Such disulfide bonds also include the disulfide bond pattern where cysteine at position 131 (EU numbering) in the H chain is linked to cysteine at position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242). This bond is unfavorable because it reduces stability.
[0383] A possible H-chain constant region that consist of a single disulfide bond and do not reduce stability is an H-chain constant region SKSC (SEQ ID NO: 28), in which serine and lysine are substituted for cysteine at position 131 and arginine at position 133 (EU numbering) respectively, in the H chain SC (SEQ ID NO: 26). Another possible H-chain constant region is M58 (SEQ ID NO: 29) which has the following additional alterations: glycine was substituted for glutamic acid at position 137 (EU numbering) in the H chain; glycine was substituted for serine at position 138; glutamine was substituted for histidine at position 268; glutamine was substituted for arginine at position 355; and glutamic acid was substituted for glutamine at position 419 to reduce immunogenicity and improve pharmacokinetics, and the C-terminal lysine and glycine were removed in advance from the H-chain constant region by deletion from the nucleotide sequence to avoid C-terminal heterogeneity. As shown in FIG. 7, the H-chain constant regions SKSC and M58 are thought to form a single disulfide bond and do not reduce stability.
[0384] Then, expression vectors for IL6R H0-SC (amino acid SEQ ID NO: 7), IL6R H0-CS (amino acid SEQ ID NO: 8), IL6R H0-SKSC (amino acid SEQ ID NO: 9), and IL6R H0-M58 (amino acid SEQ ID NO: 10) were constructed by the method described in Reference Example 1. IL6R H0-IgG1/L0-k0 and IL6R H0-IgG2/L0-k0, each of which consists of the original H chain and IL6R L0-k0 (amino acid SEQ ID NO: 2) as L chain, and IL6R H0-SC/L0-k0, IL6R H0-CS/L0-k0, IL6R H0-SKSC/L0-k0, and IL6R H0-M58/L0-k0, which are variants derived from natural IgG2, were expressed and purified by the method described in Reference Example 1.
Analysis of Various Variants from Natural IgG2 by Cation Exchange Chromatography
[0385] Various variants of natural IgG2 were assessed for their heterogeneity by the method described above using cation exchange chromatography. IL6R H0-IgG1/L0-k0 and IL6R H0-IgG2/L0-k0, and IL6R H0-SC/L0-k0, IL6R H0-CS/L0-k0, IL6R H0-SKSC/L0-k0, and IL6R H0-M58/L0-k0, which are the variants of natural IgG2, were assessed using cation exchange chromatography. The result is shown in FIG. 8.
[0386] As shown in FIG. 8, the result revealed that heterogeneity was increased when the H-chain constant region was converted from the IgG1 subtype into IgG2 subtype; however, heterogeneity was greatly reduced when the H-chain constant region was converted into SKSC or M58. On the other hand, when the H-chain constant region was converted into CS, the reduction of heterogeneity was insufficient. However, heterogeneity was reduced when the H-chain constant region was converted into SC and when the H-chain constant region was converted into SKSC.
DSC Analysis of Various Natural IgG2 Variants
[0387] To develop an antibody as pharmaceutical, it is generally desirable to have high stability in addition to low heterogeneity for preparing stable preparations. Thus, to assess the stability of IL6R H0-IgG1/L0-k0 and IL6R H0-IgG2/L0-k0, and IL6R H0-SC/L0-k0, IL6R H0-CS/L0-k0, IL6R H0-SKSC/L0-k0, and IL6R H0-M58/L0-k0, which are variants of natural IgG2, the midpoint of thermal denaturation (Tm value) was determined by differential scanning calorimetry (DSC) (VP-DSC, Microcal) in the same manner as described above. The purified antibodies were dialyzed against a solution (pH 6.0) containing 20 mM sodium acetate and 150 mM NaCl (EasySEP, TOMY). DSC measurements were carried out at a heating rate of 1.degree. C./min in a range of 40 to 100.degree. C., and at a protein concentration of about 0.1 mg/ml. The denaturation curves obtained by DSC are shown in FIG. 9, while the Tm values for the Fab domains are listed in Table 2 below.
TABLE-US-00002 TABLE 2 Tm/.degree. C. OF Fab IL6R H0-IgG1/L0-k0 94.8.degree. C. IL6R H0-IgG2/L0-k0 93.9.degree. C. IL6R H0-SC/L0-k0 86.7.degree. C. IL6R H0-CS/L0-k0 86.4.degree. C. IL6R H0-SKSC/L0-k0 93.7.degree. C. IL6R H0-M58/L0-k0 93.7.degree. C.
[0388] IL6R H0-IgG1/L0-k0 and IL6R H0-IgG2/L0-k0WT-IgG1 exhibited almost the same Tm value for the Fab domain. The Tm values were about 94.degree. C. (the Tm value of IgG2 was lower by about 1.degree. C.). The Tm values of IL6R H0-SC/L0-k0 and IL6R H0-CS/L0-k0 were about 86.degree. C. which was remarkably lower as compared to those of IL6R H0-IgG1/L0-k0 and IL6R H0-IgG2/L0-k0. Meanwhile, the Tm values of IL6R H0-SKSC/L0-k0 and IL6R H0-M58/L0-k0 were about 94.degree. C. which was comparable to those of IL6R H0-IgG1/L0-k0 and IL6R H0-IgG2/L0-k0.
[0389] The stability of H-chain constant regions SC and CS was markedly lower as compared to IgG1 and IgG2. This suggests that SC and CS form disulfide bonds that reduce stability. As described above, the stability of the Fab domain is reduced when a disulfide bond is formed between cysteine at position 131 (EU numbering) in the H chain and cysteine at position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242). The presence of molecular species having such disulfide bonds was speculated to be a cause of reduced stability of SC and CS. Thus, from the standpoint of stability, it is considered that the H-chain constant regions SKSC and M58 in which serine is substituted for cysteine at position 131 (EU numbering) in their H chains are superior as pharmaceuticals.
[0390] Furthermore, comparison of DSC denaturation curves revealed that the Fab domains of IL6R H0-IgG1/L0-k0, IL6R H0-SKSC/L0-k0, and IL6R H0-M58/L0-k0 each showed a single sharp denaturation peak. On the other hand, as compared to these, IL6R H0-SC/L0-k0 and IL6R H0-CS/L0-k0 have a broader denaturation peak for the Fab domain. IL6R H0-IgG2/L0-k0 showed a shoulder peak on the lower temperature side of the Fab domain denaturation peak. In general, the DSC denaturation peak is sharp when it contains only a single component. On the other hand, when there are a plurality of components with different Tm (i.e., heterogeneity), the denaturation peak is expected to be broader. Specifically, it is suggested that the heterogeneity detected in natural IgG2 is not sufficiently reduced in SC and CS, therefore the H-chain constant regions IgG2, SC, and CS contained a plurality of components. The finding described above suggests that not only cysteines at positions 219 and 220 (EU numbering) in the H-chain hinge region but also cysteine at position 131 (EU numbering) in the H-chain CH1 domain contributes to the heterogeneity of wild type IgG2, and it is necessary to alter not only cysteines in the hinge region but also cysteine in the CH1 domain to reduce the heterogeneity detected by DSC. Thus, from the standpoint of heterogeneity, the H-chain constant regions SKSC and M58 which have substitution of serine for cysteine at position 131 (EU numbering) in the H chain are concluded to be superior as pharmaceuticals.
[0391] As described above, SC and CS, which are H-chain constant regions having substitution of serine for cysteine only in the hinge region to reduce heterogeneity derived from the hinge region of IgG2, are assumed to be insufficient from the standpoint of both heterogeneity and stability. Thus, the present inventors demonstrated that while retaining a comparable stability to that of IgG2, the heterogeneity could be greatly reduced only when serine is substituted for cysteine in the hinge region as well as for cysteine at position 131 (EU numbering) in the H chain in the CH1 domain.
Heterogeneity-Improving Effect of Natural IgG2 Variant (M58-k0) on Various Antibodies
[0392] IL31R H0-IgG1/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 11; L-chain amino acid sequence/SEQ ID NO: 12), which is an anti-IL-31 receptor antibody, and RANKL H0-IgG1/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 15; L-chain amino acid sequence/SEQ ID NO: 16), which is an anti-RANKL antibody, were used in addition to the anti-IL-6 receptor antibodies. The H-chain constant region of each antibody was converted from the IgG1 subtype into IgG2 subtype. The resulting antibodies were IL31R H0-IgG2/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 13; L-chain amino acid sequence/SEQ ID NO: 12) and RANKL H0-IgG2/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 17; L-chain amino acid sequence/SEQ ID NO: 16). The H-chain constant region was also converted from the IgG1 subtype into M58. The resulting antibodies were IL31R H0-M58/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 14; L-chain amino acid sequence/SEQ ID NO: 12) and RANKL H0-M58/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 18; L-chain amino acid sequence/SEQ ID NO: 16). Expression vectors were constructed to express these antibodies. The antibodies were expressed and purified by the method described in Reference Example 1.
[0393] Heterogeneity was assessed by cation exchange chromatography (IEC) using the same method as described above. The results are shown in FIG. 10. As shown in FIG. 10, it was confirmed that heterogeneity was increased not only in the anti-IL-6 receptor antibodies but also in the anti-IL-31 receptor antibody and anti-RANKL antibody when the H-chain constant region was converted from the IgG1 subtype into IgG2 subtype, and that heterogeneity could be reduced in every antibody by converting the H-chain constant region from the IgG2 subtype into M58. The finding described above suggests that regardless of the sequences of antibody variable regions and antigen type, heterogeneity derived from natural IgG2 can be reduced by substituting serine for cysteine at position 131 (EU numbering) in the H-chain CH1 domain and serine for cysteine at position 219 (EU numbering) in the H-chain hinge region.
[Example 3] Pharmacokinetics-Improving Effect of Novel Constant Region M58-k0 Pharmacokinetics of IgG-Type Antibodies
[0394] The prolonged retention (slow elimination) of IgG molecule in plasma is known to be due to the function of FcRn which is known as a salvage receptor of IgG molecule (Nat. Rev. Immunol. 2007 September; 7(9): 715-25). When taken up into endosomes via pinocytosis, IgG molecules bind to FcRn expressed in endosomes under the acidic conditions within the endosomes (approx. pH 6.0). While IgG molecules that do not bind to FcRn are transferred and degraded in lysosomes, those bound to FcRn are translocated to the cell surface and then released from FcRn back into plasma again under the neutral conditions in the plasma (approx. pH 7.4).
[0395] Known IgG-type antibodies include the IgG1, IgG2, IgG3, and IgG4 isotypes. The plasma half-lives of these isotypes in human are reported to be about 36 days for IgG1 and IgG2; about 29 days for IgG3; and 16 days for IgG4 (Nat. Biotechnol. 2007 December; 25(12): 1369-72). Thus, the retention of IgG1 and IgG2 in plasma is believed to be the longest. In general, the isotypes of antibodies used as pharmaceuticals are IgG1, IgG2, and IgG4. Methods reported for further improving the pharmacokinetics of these IgG antibodies include methods for improving the above-described binding to human FcRn, and this is achieved by altering the sequence of IgG constant region (J. Biol. Chem. 2007 Jan. 19; 282(3): 1709-17; J. Immunol. 2006 Jan. 1; 176(1): 346-56).
[0396] There are species-specific differences between mouse FcRn and human FcRn (Proc. Natl. Acad. Sci. USA. 2006 Dec. 5; 103(49): 18709-14). Therefore, to predict the plasma retention of IgG antibodies that have an altered constant region sequence in human, it is desirable to assess the binding to human FcRn and retention in plasma in human FcRn transgenic mice (Int. Immunol. 2006 December; 18(12): 1759-69).
Comparison of IgG1-k0 and M58-k0 for the Binding to Human FcRn
[0397] Human FcRn was prepared according to the method described in Reference Example 2. The binding to human FcRn was assessed using Biacore 3000. An antibody was bound to Protein L or rabbit anti-human IgG Kappa chain antibody immobilized onto a sensor chip, human FcRn was added as an analyte for interaction with the antibody, and the affinity (KD) was calculated from the amount of bound human FcRn. Specifically, Protein L or rabbit anti-human IgG Kappa chain antibody was immobilized onto sensor chip CM5 (BIACORE) by the amine coupling method using 50 mM Na-phosphate buffer (pH 6.0) containing 150 mM NaCl as the running buffer. Then, IL6R H0-IgG1/L0-k0 and IL6R H0-M58/L0-k0 was each diluted with a running buffer containing 0.02% Tween20, and injected to be bound to the chip. Human FcRn was then injected and the binding of the human FcRn to antibody was assessed.
[0398] The affinity was computed using BIAevaluation Software. The obtained sensorgram was used to calculate the amount of hFcRn bound to the antibody immediately before the end of human FcRn injection. The affinity of the antibody for human FcRn was calculated by fitting with the steady state affinity method.
[0399] As a result of evaluating the binding of IL6R H0-IgG1/L0-k0 and IL6R H0-M58/L0-k towards human FcRn by BIAcore, as shown in Table 3, the binding of IL6R H0-M58/L0-k0 was found to be increased approximately 1.4 times compared to that of IL6R H0-IgG1/L0-k0.
TABLE-US-00003 TABLE 3 KD/.mu.M IL6R H0-IgG1/L0-k0 1.42 IL6R H0-M58/L0-K0 1.03
Comparison of IgG1-k0 and M58-k0 for Pharmacokinetics in Human FcRn Transgenic Mice
[0400] The pharmacokinetics in human FcRn transgenic mice (B6.mFcRn-/-.hFcRn Tg line 276+/+ mice; Jackson Laboratories) was assessed by the following procedure. IL6R H0-IgG1/L0-k0 and IL6R H0-M58/L0-k0 was each intravenously administered once at a dose of 1 mg/kg to mice, and blood was collected at appropriate time points. The collected blood was immediately centrifuged at 15,000 rpm and 4.degree. C. for 15 minutes to obtain blood plasma. The separated plasma was stored in a freezer at -20.degree. C. or below until use. The plasma concentration was determined by ELISA (see Reference Example 3).
[0401] As a result of evaluating the plasma retention of IL6R H0-IgG1/L0-k0 and IL6R H0-M58/L0-k0 in human FcRn transgenic mice, as shown in FIG. 11, pharmacokinetics of IL6R H0-M58/L0-k0 was confirmed to be improved compared to IL6R H0-IgG1/L0-k0. As indicated above, this was considered to be due to improvement in the binding of IL6R H0-M58/L0-k0 to human FcRn compared to IL6R H0-IgG1/L0-k0.
Comparison of IgG1-k0 or M58-k0 for the Binding to Human FcRn on Various Antibodies
[0402] As described above, it was demonstrated that by converting the H-chain constant region of the anti-IL-6 receptor antibody IL6R H0-IgG1/L0-k0 from IgG1 into M58, human FcRn-binding activity and pharmacokinetics were confirmed to be improved in human FcRn transgenic mice. Then, the present inventors assessed whether the pharmacokinetics of IgG1 antibodies besides anti-IL-6 receptor antibodies could also be improved by converting the H-chain constant region into M58.
[0403] In addition to the anti-IL-6 receptor antibody, IL31R H0-IgG1/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 11; L-chain amino acid sequence/SEQ ID NO: 12), which is an anti-IL-31 receptor antibody, and RANKL H0-IgG1/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 15; L-chain amino acid sequence/SEQ ID NO: 16), which is an anti-RANKL antibody, were used. The H-chain constant regions of the antibodies were converted from the IgG1 subtype into M58 to prepare antibodies IL31R H0-M58/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 14; L-chain amino acid sequence/SEQ ID NO: 12) and RANKL H0-M58/L0-k0 (H-chain amino acid sequence/SEQ ID NO: 18; L-chain amino acid sequence/SEQ ID NO: 16). The resulting antibodies were assessed for their human FcRn-binding activity by the method described above. The results are shown in Table 4.
TABLE-US-00004 TABLE 4 KD/.mu.M ANTI-IL-6 ANTI-IL-31 RECEPTOR RECEPTOR ANTI-RANKL ANTIBODY ANTIBODY ANTIBODY IgG1-k0 1.42 1.07 1.36 M58-k0 1.03 0.91 1.03
[0404] As shown in Table 4, like the anti-IL-6 receptor antibody, the anti-IL-31 receptor antibody and anti-RANKL antibody were also demonstrated to be improved in terms of human FcRn-binding activity by converting the H-chain constant region from the IgG type into M58. This finding suggests the possibility that regardless of the sequences of antibody variable regions and antigen type, the pharmacokinetics in human was improved by converting the H-chain constant region from the IgG1 subtype into M58.
[Example 4] Novel Constant Region M66-k0 Generated by Further Improving the Pharmacokinetics of M58-k0
Preparation of Novel Constant Region M66-k0
[0405] As described in Example 2, the disulfide bond pattern in the hinge region of an IgG molecule was revealed to greatly influence heterogeneity and stability. Meanwhile, Example 3 demonstrates that the H-chain constant region M58 is superior to IgG1 in pharmacokinetics. Then, the present inventors thought that a novel constant region superior to M58 in pharmacokinetics could be produced by optimizing the disulfide bond pattern in the hinge region, and assessed this possibility.
[0406] As shown in FIG. 12, since the constant region M58 has substitutions of serine for cysteine at positions 131 and 219 (EU numbering) in the H chain, cysteine at position 220 (EU numbering) in the H chain is assumed to form a disulfide bond with cysteine at position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242). On the other hand, as described in Example 2, stability is greatly reduced when cysteine at position 131 (EU numbering) in the H chain is linked via a disulfide bond to cysteine at position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242).
[0407] Then, as shown in FIG. 12, an expression vector for IL6R H0-M66 (amino acid SEQ ID NO: 19) was constructed by the method described in Reference Example 1 to assess the novel H-chain constant region M66 (amino acid SEQ ID NO: 30), which has substitutions of serine for cysteine at positions 131 and 220 (EU numbering) in the H chain, so that cysteine at position 219 (EU numbering) in the H chain forms a disulfide bond with cysteine at position 214 in the L chain (for the numbering, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242). IL6R H0-M66/L0-k0 which consists of IL6R H0-M66 (amino acid SEQ ID NO: 19) as H chain and IL6R L0-k0 (amino acid SEQ ID NO: 2) as L chain was expressed and purified by the method described in Reference Example 1.
Comparison of IgG1-k0, M58-k0, and M66-k0 for Pharmacokinetics in Human FcRn Transgenic Mice
[0408] IL6R H0-IgG1/L0-k0, IL6R H0-M58/L0-k0, and IL6R H0-M66/L0-k0 were assessed for pharmacokinetics using human FcRn transgenic mice (B6.mFcRn-/-.hFcRn Tg line 276+/+ mice; Jackson Laboratories) by the method described in Example 3.
[0409] IL6R H0-IgG1/L0-k0, IL6R H0-M58/L0-k0, and IL6R H0-M66/L0-k0 were assessed for plasma retention in human FcRn transgenic mice. As shown in FIG. 13, the result demonstrated that the pharmacokinetics of IL6R H0-M66/L0-k0 was improved as compared to that of IL6R H0-M58/L0-k0.
[0410] The amino acid sequences of IL6R H0-M58/L0-k0 and IL6R H0-M66/L0-k0 are different in that at positions 219 and 220 (EU numbering) in the H chain, IL6R H0-M58/L0-k0 has serine and cysteine, respectively, while IL6R H0-M66/L0-k0 has cysteine and serine, respectively. Specifically, cysteine at position 214 in the L chain (for the numbering, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) forms a disulfide bond with cysteine at position 220 (EU numbering) in the H chain of IL6R H0-M58/L0-k0 while cysteine at position 214 forms a disulfide bond with cysteine at position 219 (EU numbering) in the H chain of IL6R H0-M66/L0-k0. Thus, the two are different in the position of their disulfide bond.
[0411] There is no previous report demonstrating whether the pharmacokinetics of IgG varies depending on the position of disulfide bond. As described in "J Biol. Chem., 2008 Jun. 6; 283(23): 16194-205; J Biol. Chem., 2008 Jun. 6; 283(23): 16206-15; Biochemistry, 2008 Jul. 15; 47(28): 7496-508", there are many different disulfide bond patterns (isoforms) in natural IgG2, such as form A and form B. According to the report of "J Biol. Chem., 2008 Oct. 24; 283(43): 29266-72", the pharmacokinetics does not change across isoforms having disulfide bonds at different positions.
[0412] By the assessment described above, it was found for the first time that the pharmacokinetics differed greatly between IL6R H0-M58/L0-k0 and IL6R H0-M66/L0-k0 due to difference in the position of disulfide bond. Specifically, it was demonstrated that the pharmacokinetics could be greatly improved by changing the position of disulfide bond from between position 220 (EU numbering) in the H chain and position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) to between position 219 (EU numbering) in the H chain and position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242).
Analysis of Novel Constant Region M66-k0 by DSC
[0413] To assess stability, the midpoint of thermal denaturation (Tm value) was determined by differential scanning calorimetry (DSC) (N-DSCII, calorimetry Science Corporation) in a similar manner as described in Example 2. The purified IL6R H0-IgG1/L0-k0, IL6R H0-M58/L0-k0, and IL6R H0-M66/L0-k0 were dialyzed against a solution (pH 6.0) containing 20 mM sodium acetate and 150 mM NaCl (EasySEP, TOMY). DSC measurements were carried out at a heating rate of 1.degree. C./min in a range of 40 to 100.degree. C., and at a protein concentration of about 0.1 mg/ml. The Tm values for the Fab domains (as listed in Table 5) are calculated based on the denaturation curves obtained by DSC.
TABLE-US-00005 TABLE 5 Tm/.degree. C. OF Fab IL6R H0-IgG1/L0-k0 95.degree. C. IL6R H0-M58/L0-k0 94.degree. C. IL6R H0-M66/L0-k0 93.degree. C.
[0414] As shown in Table 5, the Tm value of IL6R H0-M66/L0-k0 was found to be comparable to that of H0-M58/L0-k0. This demonstrates that when M66 (amino acid SEQ ID NO: 30) is used as an H-chain constant region, the pharmacokinetics can be improved as compared to M58 (SEQ ID NO: 29) without decreasing the stability.
Cation Exchange Chromatography Analysis of Novel Constant Region M66-k0
[0415] IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-M58/L0-k0, and IL6R H0-M66/L0-k0 were assessed by cation exchange chromatography using the method described in Example 2. The result is shown in FIG. 14.
[0416] As described in FIG. 14, the result showed that the H-chain constant region M58 showed a single peak while the H-chain constant region M66 exhibited two major peaks. Likewise, for the anti-IL-31 receptor antibody, M58 showed a single peak while M66 showed two major peaks.
[0417] M58 has serine and cysteine at positions 219 and 220 (EU numbering) in the H chain, respectively, while M66 has cysteine and serine at positions 219 and 220, respectively. This minor difference remarkably improved the pharmacokinetics of M66 compared to M58. However, the new peak in M66, which was not observed with M58, indicated heterogeneity. Thus, M66 was demonstrated to show heterogeneity (isoforms) because of the presence of two types of components.
[Example 5] Novel Constant Regions M66-k3 and M66-k4 Generated by Improvement of M66-k0 in Terms of Heterogeneity by the L-Chain Constant Region Modification
Preparation of Novel Constant Regions M66-k3 and M66-k4
[0418] The heterogeneity (two types of components) of IL6R H0-M66/L0 confirmed as described in Example 4 was speculated to be due to difference in the pattern of disulfide bonds between H chain and L chain. Specifically, it is thought that two types of components corresponding to the two types of disulfide bond patterns shown in FIG. 15 were detected. It is thought that the C-terminal cysteine at position 214 (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) in the L chain (k0 amino acid SEQ ID NO: 32) (cysteine at position 107 in k0 of amino acid SEQ ID NO: 32) can form a disulfide bond with both cysteines at position 219 (EU numbering) in the two H chains. Thus, the present inventors predicted that as shown in FIG. 16, heterogeneity (to generate a single component) could be reduced when cysteine in the L chain was capable of forming a disulfide bond with cysteine at position 219 (EU numbering) in only one of the two H chains. Then, the present inventors conceived that the above-described C-terminal cysteine was moved towards the N-terminal side in the L chain to reduce heterogeneity. This could increase the distance between the C-terminal cysteine in the L chain and cysteine at position 219 (EU numbering) in one of the H chains, and as a result the L-chain C-terminal cysteine could form a disulfide bond with cysteine only at position 219 (EU numbering) in the other H chain. A possible method for relocating the L-chain C-terminal cysteine to a position on the N-terminal side was to shorten the peptide length of L chain at the C terminus. Specifically, the hypothesis was assessed using a novel antibody L-chain constant region k3 (amino acid SEQ ID NO: 33) resulting from deletion of glutamic acid at position 106 from the natural L-chain constant region k0 (amino acid SEQ ID NO: 32) and a novel antibody L-chain constant region k4 (amino acid SEQ ID NO: 34) resulting from deletion of glycine at position 105 from the natural L-chain constant region k0 (amino acid SEQ ID NO: 32). Thus, expression vectors for IL6R L0-k3 (amino acid SEQ ID NO: 21) having k3 as an L-chain constant region and IL6R L0-k4 (amino acid SEQ ID NO: 22) having k4 as an L-chain constant region were constructed by the method described in Reference Example 1.
[0419] IL6R H0-M66/L0-k3, which consists of IL6R H0-M66 (amino acid SEQ ID NO: 19) as H chain and IL6R L0-k3 (amino acid SEQ ID NO: 21) as L chain, and IL6R H0-M66/L0-k4, which consists of IL6R H0-M66 (amino acid SEQ ID NO: 19) as H chain and IL6R L0-k4 (amino acid SEQ ID NO: 22) as L chain were expressed and purified by the method described in Reference Example 1.
Cation Exchange Chromatography Analysis of Novel Constant Region M66-k3 and M66-k4
[0420] IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-M58/L0-k0, IL6R H0-M66/L0-k0, IL6R H0-M66/L0-k3, and IL6R H0-M66/L0-k4 were assessed by cation exchange chromatography using the method described in Example 2. The assessment result is shown in FIG. 17.
[0421] As shown in FIG. 17, the result demonstrated that H0-M66/L0-k0 showed two single peaks, while H0-M66/L0-k3 and H0-M66/L0-k4 each showed a single peak like H0-M58/L0-k0. This finding suggests that heterogeneity can be reduced by relocating the L-chain C-terminal cysteine position toward the N-terminal side by shortening the peptide length of the L chain at the C terminus, so that the L chain cysteine can form a disulfide bond with cysteine only at position 219 (EU numbering) in one of the two H chains.
[0422] A previously assessed method for reducing the heterogeneity (isoforms) of the disulfide bond pattern in the natural IgG2 (J Biol Chem. 2008 Jun. 6, 283(23): 16194-205; J Biol Chem. 2008 Jun. 6, 283(23): 16206-15) is to substitute serine for cysteine in the H-chain constant region. The Examples herein demonstrated for the first time that heterogeneity could be reduced by relocating the C-terminal cysteine in the L-chain constant region. There is no previous report on L-chain constant regions that result from relocation of the L-chain C-terminal cysteine.
[0423] In addition to the method described above, methods for relocating the L-chain C-terminal cysteine toward a position on the N-terminal side include, for example, deletion of arginine at position 104 from the natural L-chain constant region k0 (amino acid SEQ ID NO: 32), deletion of asparagine at position 103 from the natural L-chain constant region k0 (amino acid SEQ ID NO: 32), and substitution of cysteine for glutamic acid at position 106 in the natural L-chain constant region k0 (amino acid SEQ ID NO: 32), in combination with substitution of an amino acid other than cysteine for cysteine at position 107.
[0424] The human L-chain constant region used in this Example is a .kappa. chain (k0, amino acid SEQ ID NO: 32). The same method is expected to be applicable to the .lamda. chain constant region (amino acid SEQ ID NO: 37). The .lamda. chain constant region has cysteine at position 104 in amino acid SEQ ID NO: 37 (at position 214 in the numbering system described in "Sequences of proteins of immunological interest, NIH Publication No. 91-3242"). Thus, the method includes deletion of glutamic acid at position 103, deletion of threonine at position 102, deletion of proline at position 101, deletion of alanine at position 100, and substitution of cysteine for glutamic acid at position 103, in combination with substitution of an amino acid other than cysteine for cysteine at position 104 in the natural .lamda. chain constant region (amino acid SEQ ID NO: 37).
Assessment of Novel Constant Regions M66-k0 and M66-k3 for the Binding to Human FcRn
[0425] IgG is known to bind to FcRn with divalent avidity (Traffic. 2006 September; 7(9): 1127-42). In the method described in Example 3, IgG is immobilized onto a sensor chip and then FcRn is injected as an analyte. IgG binds to FcRn with monovalent affinity. Then, the present inventors assessed in this Example the binding of IgG to FcRn with divalent avidity by immobilizing human FcRn onto a sensor chip and injecting IgG as an analyte to more closely mimic the in vivo case. In Biacore T100 (GE Healthcare), H0-IgG1/L0-k0, H0-M58/L0-k0, H0-M66/L0-k0, and H0-M66/L0-k3 were each injected as an analyte into the sensor chip immobilized with the FcRn to analyze the affinity of the antibody variants for human FcRn at pH 6.0.
[0426] The methods of immobilization and analysis of the interaction are described below. First, about 2,000 RU of human FcRn was immobilized onto sensor chip CM4 (GE Healthcare) by the amino coupling method. Reagents used in the amino coupling were: ethanol amine (GE Healthcare), 50 mM NaOH solution (GE Healthcare), NHS (GE Healthcare), EDC (GE Healthcare). HBS-EP+ solution (10.times.HBS-EP+ solution (GE Healthcare) was used after dilution) was used as the mobile phase. Then, antibodies were each injected as an analyte into the sensor chip immobilized with human FcRn for three minutes to observe the binding of FcRn to the antibodies. After this observation, the mobile phase was injected for five minutes to assess the dissociation of each antibody variants from FcRn. All measurements were carried out at 25.degree. C. The mobile phase used was 10 mM Cit (pH 6.0)/150 mM NaCl/0.05% Tween20.
[0427] From the obtained sensorgrams, the association rate constant ka (1/Ms) and dissociation rate constant kd (1/s) were calculated for the three minutes of binding phase using Biacore T100 Evaluation Software (GE Healthcare). The dissociation constant KD (M) was determined based on these values. KDs are listed in Table 6.
TABLE-US-00006 TABLE 6 KD/nM IL6R H0-IgG1/L0-k0 4.00 IL6R H0-M58/L0-k0 3.85 IL6R H0-M66/L0-k0 3.54 IL6R H0-M66/L0-k3 3.26
[0428] As described in Examples 3 and 4, the pharmacokinetics of H0-IgG1/L0-k0, H0-M58/L0-k0, and H0-M66/L0-k0 in human FcRn transgenic mice was correlated with binding of the antibodies with human FcRn in this assay system. The human FcRn binding activity of H0-M66/L0-k3 was comparable to or greater than that of H0-M66/L0-k0. This suggests that the pharmacokinetics of H0-M66/L0-k3 in human FcRn transgenic mice is comparable or superior to that of H0-M66/L0-k0.
Analysis of Novel Constant Region M66-k3 and M66-k4 by DSC
[0429] To assess the stability, the midpoint of thermal denaturation (Tm value) was determined by differential scanning calorimetry (DSC) (N-DSCII, calorimetry Science Corporation) in a similar manner as described in Example 2. The purified IL6R H0-IgG1/L0-k0, IL6R H0-M66/L0-k0, IL6R H0-M66/L0-k3, and IL6R H0-M66/L0-k4 were dialyzed against a solution (pH 6.0) containing 20 mM sodium acetate and 150 mM NaCl (EasySEP, TOMY). DSC measurements were carried out at a heating rate of 1.degree. C./min in a range of 40 to 100.degree. C., and at a protein concentration of about 0.1 mg/ml. The Tm values for the Fab portions are calculated based on the denaturation curves obtained by DSC, which are listed in Table 7.
TABLE-US-00007 TABLE 7 Tm/.degree. C. OF Fab IL6R H0-IgG1/L0-k0 95.degree. C. IL6R H0-M66/L0-k0 93.degree. C. IL6R H0-M66/L0-k3 94.degree. C. IL6R H0-M66/L0-k4 94.degree. C.
[0430] As shown in Table 7, the Tm values of IL6R H0-M66/L0-k3 and IL6R H0-M66/L0-k4 were found to be comparable to that of H0-M66/L0-k0. This demonstrates that when k3 (amino acid SEQ ID NO: 33) or k4 (amino acid SEQ ID NO: 34) is used as an L-chain constant region, the pharmacokinetics can be improved without decreasing the stability as compared to natural L-chain constant region k0 (SEQ ID NO: 32).
[0431] As shown in FIG. 14, H0-M66/L0-k0 which was designed to form a disulfide bond between position 219 (EU numbering) in the H chain and position 214 in the L chain (for the numbering system, see Sequences of proteins of immunological interest, NIH Publication No. 91-3242) was demonstrated to have two types of isoforms. These two components can be ascribed to the two disulfide bond patterns shown in FIG. 15. Thus, it was found that the heterogeneity derived from natural IgG2 could not be completely eliminated by substitution of serine for only cysteine at positions 137 and 220 (EU numbering) in the H chain. Then, the present inventors demonstrated that the heterogeneity derived from disulfide bonds of natural IgG2 could be eliminated only when using H0-M66/L0-k3 or H0-M66/L0-k4 generating from relocation of the C-terminal cysteine in the L-chain constant region. H0-M66/L0-k3 retains the original stability and human FcRn-binding activity as compared to H0-M66/L0-k0. Furthermore, in H0-M66/L0-k3, the C-terminal heterogeneity of the H chain has been eliminated by alteration of the C-terminal .DELTA.GK in the H chain described in Example 1. Thus, M66/k3 and M66/k4 were concluded to be very useful as antibody H chain/L chain constant regions.
[Example 6] M106-k3 with Reduced Fc.gamma. Receptor-Binding of M66-k3
Preparation of Novel Constant Region M106-k3
[0432] In antibody pharmaceuticals aimed at neutralizing an antigen, effector functions such as the ADCC of Fc domain are unnecessary and therefore the binding to Fc.gamma. receptor is unnecessary. The binding to Fc.gamma. receptor is assumed to be unfavorable from the perspectives of antigenicity and side effect (Nat Rev Drug Discov. 2007 January; 6(1): 75-92; Ann Hematol. 1998 June; 76(6): 231-48). For example, the humanized anti-IL-6 receptor IgG1 antibody TOCILIZUMAB does not need to bind to Fc.gamma. receptor, because it only needs to specifically bind to IL-6 receptor and neutralize its biological activity in order to be used as a therapeutic agent for diseases associated with IL-6, such as rheumatoid arthritis.
[0433] A possible method for reducing the Fc.gamma. receptor binding is to convert the IgG antibody isotype from IgG1 into IgG2 or IgG4 (Ann. Hematol. 1998 June; 76(6): 231-48). As a method for completely eliminating the binding to Fc.gamma. receptor, a method of introducing an artificial modification into Fc domain has been reported. For example, since the effector functions of anti-CD3 antibody and anti-CD4 antibody cause side effects, amino acid mutations that are not present in the wild type sequence have been introduced into the Fc.gamma. receptor-binding region of Fc domain (J Immunol. 2000 Feb. 15; 164(4): 1925-33; J Immunol. 1997 Oct. 1; 159(7): 3613-21.), and the resulting Fc.gamma. receptor-nonbinding anti-CD3 and anti-CD4 antibodies are currently under clinical trials (Nat Rev Drug Discov. 2007 January; 6(1): 75-92, Transplantation. 2001 Apr. 15; 71(7): 941-50). According to another report (US 20050261229A1), Fc.gamma. receptor-nonbinding antibodies can be prepared by converting the Fc.gamma.R-binding domain of IgG1 (at positions 233, 234, 235, 236, 327, 330, and 331 in the EU numbering) into the sequence of IgG2 (at positions 233, 234, 235, and 236 in the EU numbering) or IgG4 (at positions 327, 330, and 331 in the EU numbering). However, if all of the above mutations are introduced into IgG1, novel peptide sequences of nine amino acids, which potentially serve as non-natural T-cell epitope peptides, will be generated, and this increases the immunogenicity risk. The immunogenicity risk should be minimized in developing antibody pharmaceuticals.
[0434] Alterations in the IgG2 constant region were considered to overcome the above-described problem. In the Fc.gamma.R-binding domain of the IgG2 constant region, the amino acids at positions 233, 234, 235 and 236 (EU numbering) are amino acids of nonbinding type; however, the amino acids at positions 330 and 331 (EU numbering) in the Fc.gamma.R-binding domain are different from the nonbinding sequence of IgG4. Therefore, it is necessary to change the amino acids at positions 330 and 331 (EU numbering) to the sequence of IgG4 (G2.DELTA.a described in Eur J Immunol. 1999 August; 29(8): 2613-24). However, since the amino acid at position 339 (EU numbering) in IgG4 is alanine while the corresponding residue in IgG2 is threonine, a simple alteration of the amino acids at positions 330 and 331 (EU numbering) to the sequence of IgG4 unfavorably generates a new peptide sequence of nine amino acids, potentially serving as a non-natural T-cell epitope peptide, and thus increases the immunogenicity risk. Then, the present inventors found that generation of the new peptide sequence could be prevented by introducing the substitution of Ala for Thr at position 339 (EU numbering) in IgG2, in addition to the alteration described above. Thus, the above-described mutations were introduced into the constant region M66 (amino acid SEQ ID NO: 30), and the resulting constant region M106 (amino acid SEQ ID NO: 31) was assessed. Then, an expression vector for IL6R H0-M106 (amino acid SEQ ID NO: 20) containing the H-chain constant region M106 was constructed by the method described in Reference Example 1.
[0435] IL6R H0-M106/L0-k0 which consists of IL6R H0-M106 (amino acid SEQ ID NO: 20) as H chain and IL6R L0-k0 (amino acid SEQ ID NO: 2) as L chain, IL6R H0-M106/L0-k3 which consists of IL6R H0-M106 (amino acid SEQ ID NO: 20) as H chain and IL6R L0-k3 (amino acid SEQ ID NO: 21) as L chain, and IL6R H0-M106/L0-k4 which consists of IL6R H0-M106 (amino acid SEQ ID NO: 20) as H chain and IL6RL0-k4 (amino acid SEQ ID NO: 22) as L chain were expressed and purified by the method described in Reference Example 1.
Cation Chromatography Analysis of Novel Constant Region M106-k3
[0436] IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, IL6R H0-M106/L0-k0, IL6R H0-M106/L0-k3, and IL6R H0-M106/L0-k4 were assessed by cation exchange chromatography using the method described in Example 2. The result is shown in FIG. 18.
[0437] As shown in FIG. 18, the result demonstrated that H0-M106/L0-k0 showed two single peaks, while H0-M106/L0-k3 and H0-M106/L0-k4 each showed a single peak like H0-M66/L0-k3 and H0-M66/L0-k4. This finding suggests that heterogeneity can be reduced by relocating the L-chain C-terminal cysteine toward the N-terminal side by shortening the peptide length of the L chain at the C-terminus, so that the L chain cysteine can form a disulfide bond with cysteine only at position 219 (EU numbering) in one of the two H chains. Thus, heterogeneity has been reduced in the H-chain constant region variant M106 as well as in the H-chain constant region variant M66.
Assessment of Novel Constant Region M106-k3 for the Binding to Various Fc.gamma. Receptors
[0438] H0-IgG1/L0-k0, H0-IgG2/L0-k0, and H0-M106/L0-k3 were assessed for the binding to Fc.gamma. receptor using the active Fc.gamma. receptors, Fc.gamma.RI, Fc.gamma.RIIa, and Fc.gamma.RIIIa.
[0439] The binding to Fc.gamma. receptor was assessed using Biacore T100 (GE Healthcare). The human Fc.gamma. receptors were allowed to interact with the antibodies captured by Protein L immobilized onto a sensor chip. The binding was assessed by comparing the amount of binding. Specifically, Protein L (ACTIgen) was immobilized onto a sensor chip CM5 (Biacore) by the amino coupling method using HBS-EP+(GE Healthcare) as a running buffer. Then, H0-IgG1/L0-k0, H0-IgG2/L0-k0, and H0-M106/L0-k3 were captured by Protein L immobilized onto the sensor chip, and allowed to interact with the following analytes: running buffer, and Fc.gamma.RI, Fc.gamma.RIIa, and Fc.gamma.RIIIa (R&D systems) diluted to 10 .mu.g/ml with running buffer. Since it was difficult to keep the amounts of the respective antibodies captured by Protein L constant in this assay, the amounts were corrected to be constant. Specifically, the binding amount when running buffer alone was used in the interaction was subtracted from the binding amount of each human Fc.gamma. receptor, and the resulting value was divided by the amount of each antibody captured. The obtained value was multiplied by 100, and the resulting value was used as "Normalized response".
[0440] The result of comparison of binding intensities between various antibodies and human Fc.gamma. receptor using "Normalized response" is shown in FIG. 19. The result revealed that the binding activity of H0-M106/L0-k3 to various active Fc.gamma. receptors was significantly lower than that of natural IgG1 and also lower than that of H0-IgG2/L0-k0. Thus, the Fc.gamma. receptor-binding activity of the novel constant region M106-k3 was demonstrated to be lower than that of natural IgG2. This finding suggests that the immunogenicity risk due to Fc.gamma. receptor-mediated internalization into APC and side effects caused by the effector function such as ADCC can be reduced to less than those of natural IgG2 by using H0-M106/L0-k3.
[Example 7] Cation Exchange Chromatography Analysis of IgG2-k3
[0441] IL6R H0-IgG2/L0-k3 which consists of IL6R H0-IgG2 (amino acid SEQ ID NO: 5) as H chain and IL6R L0-k3 (amino acid SEQ ID NO: 21) as L chain was expressed and purified by the method described in Reference Example 1. IL6R H0-IgG1/L0-k0, IL6R H0-IgG2/L0-k0, and IL6R H0-IgG2/L0-k3 were assessed by cation exchange chromatography using the method described in Example 2. The result is shown in FIG. 20.
[0442] As shown in FIG. 20, the result confirmed heterogeneity in IgG2-k0 and showed reduction of heterogeneity in IgG2-k3. IgG2-k0, which is a natural IgG2, exhibits heterogeneity as a result of different disulfide bond patterns. It was demonstrated that heterogeneity could be reduced by merely relocating the L-chain C-terminal cysteine toward a position on the N-terminal side by shortening the peptide length of the L chain at the C terminus (IgG2-k3).
[Example 8] Comparison of IgG1-k0, M66-k0, M66-k3, M106-k3, and IgG2-k3 on Pharmacokinetics in Human FcRn Transgenic Mice
[0443] IL6R H0-IgG1/L0-k0, IL6R H0-M66/L0-k0, IL6R H0-M66/L0-k3, IL6R H0-M106/L0-k3, and IL6R H0-IgG2/L0-k3 were assessed for the pharmacokinetics using human FcRn transgenic mice (B6.mFcRn-/-.hFcRn Tg line 276+/+ mice; Jackson Laboratories) by the method described in Example 3.
[0444] IL6R H0-M66/L0-k0, IL6R H0-M66/L0-k3, IL6R H0-M106/L0-k3, and IL6R H0-IgG2/L0-k3 were assessed for the plasma retention in human FcRn transgenic mice. As shown in FIG. 21, the result demonstrated that IL6R H0-M66/L0-k3 was improved in terms of the pharmacokinetics as compared to IL6R H0-M66/L0-k0. It is thought that this reflects the evaluation results of FcRn binding described in Example 5. L-chain constant region k3 (amino acid SEQ ID NO: 33) generated from deleting glutamic acid at position 106 from the natural L-chain constant region k0 (amino acid SEQ ID NO: 32). Thus, it is thought that the improved plasma retention was due to the substitution of L0-k3 for L0-k0 in the L chain. Since FcRn binds to the Fc domain of the H-chain constant region, the L-chain constant region is in general believed not to affect antibody pharmacokinetics. Indeed, there is no previous report describing that the pharmacokinetics in human FcRn transgenic mice was improved by amino acid substitution in the L-chain constant region. The present inventors for the first time revealed that the pharmacokinetics was improved by amino acid substitution in the L-chain constant region. Furthermore, IL6R H0-M106/L0-k3 exhibited improved plasma retention as compared to IL6R H0-M66/L0-k3.
[0445] The constant regions M66-k3, M106-k3, and IgG2-k3 were each eluted as a single peak in the assay using cation exchange chromatography. The assay demonstrated that the binding of M66-k3, M106-k3, and IgG2-k3 to Fc.gamma. receptor were significantly reduced relative to natural IgG1 and that the pharmacokinetics of constant regions M66-k3, M106-k3, and IgG2-k3 were greatly improved in human FcRn transgenic mice as compared to that of natural IgG1.
[Reference Example 1] Production of Antibody Expression Vectors and Expression and Purification of Antibodies
[0446] Genes encoding the nucleotide sequences of the H chain and L chain of the antibody of interest were amplified using PCR and such by methods known to those skilled in the art. Introduction of amino acid substitutions were carried out by methods known to those skilled in the art using QuikChange Site-Directed Mutagenesis Kit (Stratagene), PCR, or such. The obtained plasmid fragment was inserted into an animal cell expression vector, and the H-chain expression vector and L-chain expression vector of interest were produced. The nucleotide sequence of the obtained expression vector was determined by a method known to those skilled in the art. The antibodies were expressed by the following method. Human embryonic kidney cancer-derived HEK293H cells (Invitrogen) were suspended in DMEM (Invitrogen) supplemented with 10% Fetal Bovine Serum (Invitrogen). The cells (10-ml/plate; cell density of 5 to 6.times.10.sup.5 cells/ml) were plated on dishes for adherent cells (10 cm in diameter; CORNING) and cultured in a CO.sub.2 incubator (37.degree. C., 5% CO.sub.2) for one whole day and night. Then, the medium was removed by aspiration, and 6.9 ml of CHO-S-SFM-II medium (Invitrogen) was added. The prepared plasmids were introduced into cells by lipofection method. The obtained culture supernatants were collected and centrifuged (approx. 2000 g, 5 min, room temperature) to remove the cells, and sterilized through 0.22-.mu.n filter MILLER.RTM.-GV (Millipore) to prepare culture supernatant. Antibodies were purified from the obtained culture supernatant by a method known to those skilled in the art using rProtein A Sepharose.TM. Fast Flow (Amersham Biosciences). Absorbance at 280 nm was measured using a spectrophotometer to know the purified antibody concentrations. Extinction coefficient calculated from the obtained value by the PACE method was used to calculate the antibody concentration (Protein Science (1995) 4: 2411-2423).
[Reference Example 2] Preparation of Human FcRn
[0447] FcRn is a complex of FcRn and .beta.2-microglobulin. Oligo-DNA primers were prepared based on the human FcRn gene sequence disclosed (J. Exp. Med. (1994) 180(6): 2377-2381). A DNA fragment encoding the whole gene was prepared by PCR using human cDNA (Human Placenta Marathon-Ready cDNA, Clontech) as a template and the prepared primers. Using the obtained DNA fragment as a template, a DNA fragment encoding the extracellular domain containing the signal region (Met1-Leu290) was amplified by PCR, and inserted into an animal cell expression vector (the amino acid sequence of human FcRn as set forth in SEQ ID NO: 35). Likewise, oligo-DNA primers were prepared based on the human .beta.2-microglobulin gene sequence disclosed (Proc. Natl. Acad. Sci. USA. (2002) 99(26): 16899-16903). A DNA fragment encoding the whole gene was prepared by PCR using human cDNA (Hu-Placenta Marathon-Ready cDNA, CLONTECH) as a template and the prepared primers. Using the obtained DNA fragment as a template, a DNA fragment encoding the whole .beta.2-microglobulin containing the signal region (Met1-Met119) was amplified by PCR and inserted into an animal cell expression vector (the amino acid sequence of human .beta.2-microglobulin as set forth in SEQ ID NO: 36).
[0448] Soluble human FcRn was expressed by the following procedure. The plasmids constructed for human FcRn and .beta.2-microglobulin were introduced into cells of the human embryonic kidney cancer-derived cell line HEK293H (Invitrogen) using 10% Fetal Bovine Serum (Invitrogen) by lipofection. The resulting culture supernatant was collected, and FcRn was purified using IgG Sepharose 6 Fast Flow (Amersham Biosciences) by the method described in J. Immunol. 2002 Nov. 1; 169(9): 5171-80, followed by further purification using HiTrap Q HP (GE Healthcare).
[Reference Example 3] Measurement of Plasma Antibody Concentration in Mice
[0449] Measurement of the mouse plasma antibody concentration was carried out by the ELISA method using anti-human IgG antibodies and using each of the antibodies as standards according to a method known to those skilled in the art.
INDUSTRIAL APPLICABILITY
[0450] The present invention is useful in the production of antibodies which will be administered to living organisms as pharmaceuticals. More specifically, antibodies comprising the constant regions of the present invention are advantageous in maintaining the quality of the pharmaceuticals since heterogeneity is low. In other words, by using an antibody comprising a constant region of the present invention as a pharmaceutical, a steady supply of homogeneous antibodies will be possible. For example, TOCILIZUMAB (common name) which is an antibody against the IL-6 receptor is a humanized antibody used for treatment of autoimmune diseases and such. Therefore, for example, quality can be kept stable by substituting a constant region provided by the present invention for the constant region of this antibody.
[0451] Furthermore, the present invention provided antibodies with improved pharmacokinetics by altering the amino acid sequence of the constant regions. Antibodies subjected to improvement of pharmacokinetics by the present invention maintain activity for a longer time in a living body. Therefore, for example, by substituting a constant region provided by the present invention for the constant region of TOCILIZUMAB (common name) which is an antibody against the IL-6 receptor, its pharmacokinetics is improved, and it can be an antibody that may maintain the active concentration in a living body for a long time.
Sequence CWU 1
1
431449PRTArtificialAn artificially synthesized peptide sequence
1Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Arg Pro Ser Gln1 5
10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr Ser Ile Thr Ser
Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro Gly Arg Gly Leu
Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile Thr Thr Tyr Asn
Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg Asp Thr Ser Lys
Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala Arg Thr Thr Ala
Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val Thr Val Ser Ser
Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala Pro Ser
Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150 155
160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu
165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro Ser 180 185 190Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val
Asn His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Lys Val Glu
Pro Lys Ser Cys Asp Lys 210 215 220Thr His Thr Cys Pro Pro Cys Pro
Ala Pro Glu Leu Leu Gly Gly Pro225 230 235 240Ser Val Phe Leu Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser 245 250 255Arg Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp 260 265 270Pro
Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn 275 280
285Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val
290 295 300Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly
Lys Glu305 310 315 320Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro
Ala Pro Ile Glu Lys 325 330 335Thr Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr 340 345 350Leu Pro Pro Ser Arg Asp Glu
Leu Thr Lys Asn Gln Val Ser Leu Thr 355 360 365Cys Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu 370 375 380Ser Asn Gly
Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu385 390 395
400Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
405 410 415Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu 420 425 430Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly 435 440 445Lys2214PRTArtificialAn artificially
synthesized peptide sequence 2Asp Ile Gln Met Thr Gln Ser Pro Ser
Ser Leu Ser Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg
Ala Ser Gln Asp Ile Ser Ser Tyr 20 25 30Leu Asn Trp Tyr Gln Gln Lys
Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40 45Tyr Tyr Thr Ser Arg Leu
His Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr
Asp Phe Thr Phe Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Ile
Ala Thr Tyr Tyr Cys Gln Gln Gly Asn Thr Leu Pro Tyr 85 90 95Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105
110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly
115 120 125Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg
Glu Ala 130 135 140Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser
Gly Asn Ser Gln145 150 155 160Glu Ser Val Thr Glu Gln Asp Ser Lys
Asp Ser Thr Tyr Ser Leu Ser 165 170 175Ser Thr Leu Thr Leu Ser Lys
Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190Ala Cys Glu Val Thr
His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200 205Phe Asn Arg
Gly Glu Cys 2103448PRTArtificialAn artificially synthesized peptide
sequence 3Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Arg Pro
Ser Gln1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr Ser Ile
Thr Ser Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro Gly Arg
Gly Leu Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile Thr Thr
Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg Asp Thr
Ser Lys Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala
Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala Arg Thr
Thr Ala Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala
Pro Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu 130 135 140Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150
155 160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val
Leu 165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val Pro Ser 180 185 190Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn
Val Asn His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Lys Val
Glu Pro Lys Ser Cys Asp Lys 210 215 220Thr His Thr Cys Pro Pro Cys
Pro Ala Pro Glu Leu Leu Gly Gly Pro225 230 235 240Ser Val Phe Leu
Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser 245 250 255Arg Thr
Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp 260 265
270Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn
275 280 285Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val 290 295 300Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu
Asn Gly Lys Glu305 310 315 320Tyr Lys Cys Lys Val Ser Asn Lys Ala
Leu Pro Ala Pro Ile Glu Lys 325 330 335Thr Ile Ser Lys Ala Lys Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr 340 345 350Leu Pro Pro Ser Arg
Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr 355 360 365Cys Leu Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu 370 375 380Ser
Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu385 390
395 400Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp
Lys 405 410 415Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val
Met His Glu 420 425 430Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Pro Gly 435 440 4454447PRTArtificialAn artificially
synthesized peptide sequence 4Gln Val Gln Leu Gln Glu Ser Gly Pro
Gly Leu Val Arg Pro Ser Gln1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val
Ser Gly Tyr Ser Ile Thr Ser Asp 20 25 30His Ala Trp Ser Trp Val Arg
Gln Pro Pro Gly Arg Gly Leu Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr
Ser Gly Ile Thr Thr Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr
Met Leu Arg Asp Thr Ser Lys Asn Gln Phe Ser65 70 75 80Leu Arg Leu
Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg
Ser Leu Ala Arg Thr Thr Ala Met Asp Tyr Trp Gly Gln Gly 100 105
110Ser Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe
115 120 125Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala
Ala Leu 130 135 140Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val
Thr Val Ser Trp145 150 155 160Asn Ser Gly Ala Leu Thr Ser Gly Val
His Thr Phe Pro Ala Val Leu 165 170 175Gln Ser Ser Gly Leu Tyr Ser
Leu Ser Ser Val Val Thr Val Pro Ser 180 185 190Ser Ser Leu Gly Thr
Gln Thr Tyr Ile Cys Asn Val Asn His Lys Pro 195 200 205Ser Asn Thr
Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Asp Lys 210 215 220Thr
His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro225 230
235 240Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
Ser 245 250 255Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser
His Glu Asp 260 265 270Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
Val Glu Val His Asn 275 280 285Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn Ser Thr Tyr Arg Val 290 295 300Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu Asn Gly Lys Glu305 310 315 320Tyr Lys Cys Lys
Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys 325 330 335Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr 340 345
350Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr
355 360 365Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu
Trp Glu 370 375 380Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr
Pro Pro Val Leu385 390 395 400Asp Ser Asp Gly Ser Phe Phe Leu Tyr
Ser Lys Leu Thr Val Asp Lys 405 410 415Ser Arg Trp Gln Gln Gly Asn
Val Phe Ser Cys Ser Val Met His Glu 420 425 430Ala Leu His Asn His
Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro 435 440
4455445PRTArtificialAn artificially synthesized peptide sequence
5Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Arg Pro Ser Gln1 5
10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr Ser Ile Thr Ser
Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro Gly Arg Gly Leu
Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile Thr Thr Tyr Asn
Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg Asp Thr Ser Lys
Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala Arg Thr Thr Ala
Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val Thr Val Ser Ser
Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala Pro Cys
Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala Leu 130 135 140Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150 155
160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu
165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro Ser 180 185 190Ser Asn Phe Gly Thr Gln Thr Tyr Thr Cys Asn Val
Asp His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Thr Val Glu
Arg Lys Cys Cys Val Glu 210 215 220Cys Pro Pro Cys Pro Ala Pro Pro
Val Ala Gly Pro Ser Val Phe Leu225 230 235 240Phe Pro Pro Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu 245 250 255Val Thr Cys
Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Gln 260 265 270Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys 275 280
285Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu
290 295 300Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys305 310 315 320Val Ser Asn Lys Gly Leu Pro Ala Pro Ile Glu
Lys Thr Ile Ser Lys 325 330 335Thr Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser 340 345 350Arg Glu Glu Met Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys 355 360 365Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln 370 375 380Pro Glu Asn
Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp Ser Asp Gly385 390 395
400Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln
405 410 415Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn 420 425 430His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly
Lys 435 440 4456446PRTArtificialAn artificially synthesized peptide
sequence 6Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Arg Pro
Ser Gln1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr Ser Ile
Thr Ser Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro Gly Arg
Gly Leu Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile Thr Thr
Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg Asp Thr
Ser Lys Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala
Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala Arg Thr
Thr Ala Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala Leu 130 135 140Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150
155 160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val
Leu 165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val Pro Ser 180 185 190Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn
Val Asp His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Arg Val
Glu Ser Lys Tyr Gly Pro Pro 210 215 220Cys Pro Pro Cys Pro Ala Pro
Glu Phe Leu Gly Gly Pro Ser Val Phe225 230 235 240Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro 245 250 255Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu Val 260 265
270Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr
275 280 285Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val
Ser Val 290 295 300Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys
Glu Tyr Lys Cys305 310 315 320Lys Val Ser Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys Thr Ile Ser 325 330 335Lys Ala Lys Gly Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro 340 345 350Ser Gln Glu Glu Met
Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val 355 360 365Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly 370 375 380Gln
Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp385 390
395 400Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg
Trp 405 410 415Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu
Ala Leu His
420 425 430Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys
435 440 4457445PRTArtificialAn artificially synthesized peptide
sequence 7Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Arg Pro
Ser Gln1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr Ser Ile
Thr Ser Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro Gly Arg
Gly Leu Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile Thr Thr
Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg Asp Thr
Ser Lys Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala
Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala Arg Thr
Thr Ala Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala Leu 130 135 140Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150
155 160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val
Leu 165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val Pro Ser 180 185 190Ser Asn Phe Gly Thr Gln Thr Tyr Thr Cys Asn
Val Asp His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Thr Val
Glu Arg Lys Ser Cys Val Glu 210 215 220Cys Pro Pro Cys Pro Ala Pro
Pro Val Ala Gly Pro Ser Val Phe Leu225 230 235 240Phe Pro Pro Lys
Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu 245 250 255Val Thr
Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Gln 260 265
270Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys
275 280 285Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser
Val Leu 290 295 300Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr Lys Cys Lys305 310 315 320Val Ser Asn Lys Gly Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys 325 330 335Thr Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu Pro Pro Ser 340 345 350Arg Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys 355 360 365Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln 370 375 380Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp Ser Asp Gly385 390
395 400Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp
Gln 405 410 415Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala
Leu His Asn 420 425 430His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro
Gly Lys 435 440 4458445PRTArtificialAn artificially synthesized
peptide sequence 8Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val
Arg Pro Ser Gln1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr
Ser Ile Thr Ser Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro
Gly Arg Gly Leu Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile
Thr Thr Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg
Asp Thr Ser Lys Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val
Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala
Arg Thr Thr Ala Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val
Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro
Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala Leu 130 135
140Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser
Trp145 150 155 160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe
Pro Ala Val Leu 165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser
Val Val Thr Val Pro Ser 180 185 190Ser Asn Phe Gly Thr Gln Thr Tyr
Thr Cys Asn Val Asp His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp
Lys Thr Val Glu Arg Lys Cys Ser Val Glu 210 215 220Cys Pro Pro Cys
Pro Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu225 230 235 240Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu 245 250
255Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Gln
260 265 270Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys
Thr Lys 275 280 285Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val
Val Ser Val Leu 290 295 300Thr Val Val His Gln Asp Trp Leu Asn Gly
Lys Glu Tyr Lys Cys Lys305 310 315 320Val Ser Asn Lys Gly Leu Pro
Ala Pro Ile Glu Lys Thr Ile Ser Lys 325 330 335Thr Lys Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser 340 345 350Arg Glu Glu
Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys 355 360 365Gly
Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln 370 375
380Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp Ser Asp
Gly385 390 395 400Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser Arg Trp Gln 405 410 415Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala Leu His Asn 420 425 430His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 435 440 4459445PRTArtificialAn artificially
synthesized peptide sequence 9Gln Val Gln Leu Gln Glu Ser Gly Pro
Gly Leu Val Arg Pro Ser Gln1 5 10 15Thr Leu Ser Leu Thr Cys Thr Val
Ser Gly Tyr Ser Ile Thr Ser Asp 20 25 30His Ala Trp Ser Trp Val Arg
Gln Pro Pro Gly Arg Gly Leu Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr
Ser Gly Ile Thr Thr Tyr Asn Pro Ser Leu 50 55 60Lys Ser Arg Val Thr
Met Leu Arg Asp Thr Ser Lys Asn Gln Phe Ser65 70 75 80Leu Arg Leu
Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg
Ser Leu Ala Arg Thr Thr Ala Met Asp Tyr Trp Gly Gln Gly 100 105
110Ser Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe
115 120 125Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Glu Ser Thr Ala
Ala Leu 130 135 140Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val
Thr Val Ser Trp145 150 155 160Asn Ser Gly Ala Leu Thr Ser Gly Val
His Thr Phe Pro Ala Val Leu 165 170 175Gln Ser Ser Gly Leu Tyr Ser
Leu Ser Ser Val Val Thr Val Pro Ser 180 185 190Ser Asn Phe Gly Thr
Gln Thr Tyr Thr Cys Asn Val Asp His Lys Pro 195 200 205Ser Asn Thr
Lys Val Asp Lys Thr Val Glu Arg Lys Ser Cys Val Glu 210 215 220Cys
Pro Pro Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu225 230
235 240Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro
Glu 245 250 255Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro
Glu Val Gln 260 265 270Phe Asn Trp Tyr Val Asp Gly Val Glu Val His
Asn Ala Lys Thr Lys 275 280 285Pro Arg Glu Glu Gln Phe Asn Ser Thr
Phe Arg Val Val Ser Val Leu 290 295 300Thr Val Val His Gln Asp Trp
Leu Asn Gly Lys Glu Tyr Lys Cys Lys305 310 315 320Val Ser Asn Lys
Gly Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys 325 330 335Thr Lys
Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser 340 345
350Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys
355 360 365Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn
Gly Gln 370 375 380Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu
Asp Ser Asp Gly385 390 395 400Ser Phe Phe Leu Tyr Ser Lys Leu Thr
Val Asp Lys Ser Arg Trp Gln 405 410 415Gln Gly Asn Val Phe Ser Cys
Ser Val Met His Glu Ala Leu His Asn 420 425 430His Tyr Thr Gln Lys
Ser Leu Ser Leu Ser Pro Gly Lys 435 440 44510443PRTArtificialAn
artificially synthesized peptide sequence 10Gln Val Gln Leu Gln Glu
Ser Gly Pro Gly Leu Val Arg Pro Ser Gln1 5 10 15Thr Leu Ser Leu Thr
Cys Thr Val Ser Gly Tyr Ser Ile Thr Ser Asp 20 25 30His Ala Trp Ser
Trp Val Arg Gln Pro Pro Gly Arg Gly Leu Glu Trp 35 40 45Ile Gly Tyr
Ile Ser Tyr Ser Gly Ile Thr Thr Tyr Asn Pro Ser Leu 50 55 60Lys Ser
Arg Val Thr Met Leu Arg Asp Thr Ser Lys Asn Gln Phe Ser65 70 75
80Leu Arg Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Ser Leu Ala Arg Thr Thr Ala Met Asp Tyr Trp Gly Gln
Gly 100 105 110Ser Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro
Ser Val Phe 115 120 125Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly
Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu Val Lys Asp Tyr Phe Pro
Glu Pro Val Thr Val Ser Trp145 150 155 160Asn Ser Gly Ala Leu Thr
Ser Gly Val His Thr Phe Pro Ala Val Leu 165 170 175Gln Ser Ser Gly
Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro Ser 180 185 190Ser Asn
Phe Gly Thr Gln Thr Tyr Thr Cys Asn Val Asp His Lys Pro 195 200
205Ser Asn Thr Lys Val Asp Lys Thr Val Glu Arg Lys Ser Cys Val Glu
210 215 220Cys Pro Pro Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val
Phe Leu225 230 235 240Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
Ser Arg Thr Pro Glu 245 250 255Val Thr Cys Val Val Val Asp Val Ser
Gln Glu Asp Pro Glu Val Gln 260 265 270Phe Asn Trp Tyr Val Asp Gly
Val Glu Val His Asn Ala Lys Thr Lys 275 280 285Pro Arg Glu Glu Gln
Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu 290 295 300Thr Val Val
His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys305 310 315
320Val Ser Asn Lys Gly Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys
325 330 335Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro
Pro Ser 340 345 350Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr
Cys Leu Val Lys 355 360 365Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu
Trp Glu Ser Asn Gly Gln 370 375 380Pro Glu Asn Asn Tyr Lys Thr Thr
Pro Pro Met Leu Asp Ser Asp Gly385 390 395 400Ser Phe Phe Leu Tyr
Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln 405 410 415Glu Gly Asn
Val Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn 420 425 430His
Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro 435 44011451PRTArtificialAn
artificially synthesized peptide sequence 11Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser
Cys Lys Ala Ser Gly Tyr Thr Phe Thr Gly Tyr 20 25 30Ile Met Asn Trp
Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Leu Ile
Asn Pro Tyr Asn Gly Gly Thr Ser Tyr Asn Gln Lys Phe 50 55 60Lys Gly
Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Gly Tyr Asp Asp Gly Pro Tyr Thr Met Asp Tyr Trp
Gly 100 105 110Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys
Gly Pro Ser 115 120 125Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr
Ser Gly Gly Thr Ala 130 135 140Ala Leu Gly Cys Leu Val Lys Asp Tyr
Phe Pro Glu Pro Val Thr Val145 150 155 160Ser Trp Asn Ser Gly Ala
Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170 175Val Leu Gln Ser
Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val 180 185 190Pro Ser
Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His 195 200
205Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys
210 215 220Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu
Leu Gly225 230 235 240Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro
Lys Asp Thr Leu Met 245 250 255Ile Ser Arg Thr Pro Glu Val Thr Cys
Val Val Val Asp Val Ser His 260 265 270Glu Asp Pro Glu Val Lys Phe
Asn Trp Tyr Val Asp Gly Val Glu Val 275 280 285His Asn Ala Lys Thr
Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr 290 295 300Arg Val Val
Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly305 310 315
320Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile
325 330 335Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro
Gln Val 340 345 350Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys
Asn Gln Val Ser 355 360 365Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro
Ser Asp Ile Ala Val Glu 370 375 380Trp Glu Ser Asn Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro Pro385 390 395 400Val Leu Asp Ser Asp
Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val 405 410 415Asp Lys Ser
Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met 420 425 430His
Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser 435 440
445Pro Gly Lys 45012214PRTArtificialAn artificially synthesized
peptide sequence 12Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser
Ala Ser Val Gly1 5 10 15Asp Arg Val Thr Ile Thr Cys Arg Thr Ser Glu
Asn Ile Tyr Ser Phe 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Lys
Ala Pro Lys Leu Leu Ile 35 40 45Tyr Asn Ala Lys Thr Leu Ala Lys Gly
Val Pro Ser Arg Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr
Leu Thr Ile Ser Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr
Tyr Cys Gln His His Tyr Glu Ser Pro Leu 85 90 95Thr Phe Gly Gly Gly
Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val
Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr
Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135
140Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser
Gln145 150 155 160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser 165
170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val
Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val
Thr Lys Ser 195 200 205Phe Asn Arg Gly Glu Cys
21013447PRTArtificialAn artificially synthesized peptide sequence
13Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Gly
Tyr 20 25 30Ile Met Asn Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Leu Ile Asn Pro Tyr Asn Gly Gly Thr Ser Tyr Asn
Gln Lys Phe 50 55 60Lys Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr
Ser Thr Ala Tyr65 70 75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Asp Gly Tyr Asp Asp Gly Pro
Tyr Thr Met Asp Tyr Trp Gly 100 105 110Gln Gly Thr Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125Val Phe Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140Ala Leu Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val145 150 155
160Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala
165 170 175Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val
Thr Val 180 185 190Pro Ser Ser Asn Phe Gly Thr Gln Thr Tyr Thr Cys
Asn Val Asp His 195 200 205Lys Pro Ser Asn Thr Lys Val Asp Lys Thr
Val Glu Arg Lys Cys Cys 210 215 220Val Glu Cys Pro Pro Cys Pro Ala
Pro Pro Val Ala Gly Pro Ser Val225 230 235 240Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr 245 250 255Pro Glu Val
Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu 260 265 270Val
Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys 275 280
285Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser
290 295 300Val Leu Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr Lys305 310 315 320Cys Lys Val Ser Asn Lys Gly Leu Pro Ala Pro
Ile Glu Lys Thr Ile 325 330 335Ser Lys Thr Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu Pro 340 345 350Pro Ser Arg Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys Leu 355 360 365Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn 370 375 380Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp Ser385 390 395
400Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg
405 410 415Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu
Ala Leu 420 425 430His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
Pro Gly Lys 435 440 44514445PRTArtificialAn artificially
synthesized peptide sequence 14Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala1 5 10 15Ser Val Lys Val Ser Cys Lys Ala
Ser Gly Tyr Thr Phe Thr Gly Tyr 20 25 30Ile Met Asn Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45Gly Leu Ile Asn Pro Tyr
Asn Gly Gly Thr Ser Tyr Asn Gln Lys Phe 50 55 60Lys Gly Arg Val Thr
Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr65 70 75 80Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg
Asp Gly Tyr Asp Asp Gly Pro Tyr Thr Met Asp Tyr Trp Gly 100 105
110Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser
115 120 125Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly Gly
Thr Ala 130 135 140Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu
Pro Val Thr Val145 150 155 160Ser Trp Asn Ser Gly Ala Leu Thr Ser
Gly Val His Thr Phe Pro Ala 165 170 175Val Leu Gln Ser Ser Gly Leu
Tyr Ser Leu Ser Ser Val Val Thr Val 180 185 190Pro Ser Ser Asn Phe
Gly Thr Gln Thr Tyr Thr Cys Asn Val Asp His 195 200 205Lys Pro Ser
Asn Thr Lys Val Asp Lys Thr Val Glu Arg Lys Ser Cys 210 215 220Val
Glu Cys Pro Pro Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val225 230
235 240Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
Thr 245 250 255Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu
Asp Pro Glu 260 265 270Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His Asn Ala Lys 275 280 285Thr Lys Pro Arg Glu Glu Gln Phe Asn
Ser Thr Phe Arg Val Val Ser 290 295 300Val Leu Thr Val Val His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys305 310 315 320Cys Lys Val Ser
Asn Lys Gly Leu Pro Ala Pro Ile Glu Lys Thr Ile 325 330 335Ser Lys
Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro 340 345
350Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu
355 360 365Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu
Ser Asn 370 375 380Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro
Met Leu Asp Ser385 390 395 400Asp Gly Ser Phe Phe Leu Tyr Ser Lys
Leu Thr Val Asp Lys Ser Arg 405 410 415Trp Gln Glu Gly Asn Val Phe
Ser Cys Ser Val Met His Glu Ala Leu 420 425 430His Asn His Tyr Thr
Gln Lys Ser Leu Ser Leu Ser Pro 435 440 44515452PRTArtificialAn
artificially synthesized peptide sequence 15Glu Val Gln Leu Leu Glu
Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser
Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30Ala Met Ser Trp
Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile
Thr Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly
Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Lys Asp Pro Gly Thr Thr Val Ile Met Ser Trp Phe Asp Pro
Trp 100 105 110Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr
Lys Gly Pro 115 120 125Ser Val Phe Pro Leu Ala Pro Ser Ser Lys Ser
Thr Ser Gly Gly Thr 130 135 140Ala Ala Leu Gly Cys Leu Val Lys Asp
Tyr Phe Pro Glu Pro Val Thr145 150 155 160Val Ser Trp Asn Ser Gly
Ala Leu Thr Ser Gly Val His Thr Phe Pro 165 170 175Ala Val Leu Gln
Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr 180 185 190Val Pro
Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn 195 200
205His Lys Pro Ser Asn Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser
210 215 220Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu
Leu Leu225 230 235 240Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys Asp Thr Leu 245 250 255Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val Asp Val Ser 260 265 270His Glu Asp Pro Glu Val Lys
Phe Asn Trp Tyr Val Asp Gly Val Glu 275 280 285Val His Asn Ala Lys
Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr 290 295 300Tyr Arg Val
Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn305 310 315
320Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro
325 330 335Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln 340 345 350Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr
Lys Asn Gln Val 355 360 365Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val 370 375 380Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro385 390 395 400Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr 405 410 415Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val 420 425 430Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu 435 440
445Ser Pro Gly Lys 45016215PRTArtificialAn artificially synthesized
peptide sequence 16Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser
Leu Ser Pro Gly1 5 10 15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln
Ser Val Arg Gly Arg 20 25 30Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly
Gln Ala Pro Arg Leu Leu 35 40 45Ile Tyr Gly Ala Ser Ser Arg Ala Thr
Gly Ile Pro Asp Arg Phe Ser 50 55 60Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Arg Leu Glu65 70 75 80Pro Glu Asp Phe Ala Val
Phe Tyr Cys Gln Gln Tyr Gly Ser Ser Pro 85 90 95Arg Thr Phe Gly Gln
Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala 100 105 110Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser 115 120 125Gly
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu 130 135
140Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn
Ser145 150 155 160Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser
Thr Tyr Ser Leu 165 170 175Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp
Tyr Glu Lys His Lys Val 180 185 190Tyr Ala Cys Glu Val Thr His Gln
Gly Leu Ser Ser Pro Val Thr Lys 195 200 205Ser Phe Asn Arg Gly Glu
Cys 210 21517448PRTArtificialAn artificially synthesized peptide
sequence 17Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro
Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Ser Ser Tyr 20 25 30Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly
Leu Glu Trp Val 35 40 45Ser Gly Ile Thr Gly Ser Gly Gly Ser Thr Tyr
Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ser Lys Asn Thr Leu Tyr65 70 75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Lys Asp Pro Gly Thr Thr
Val Ile Met Ser Trp Phe Asp Pro Trp 100 105 110Gly Gln Gly Thr Leu
Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro 115 120 125Ser Val Phe
Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr 130 135 140Ala
Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr145 150
155 160Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe
Pro 165 170 175Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser
Val Val Thr 180 185 190Val Pro Ser Ser Asn Phe Gly Thr Gln Thr Tyr
Thr Cys Asn Val Asp 195 200 205His Lys Pro Ser Asn Thr Lys Val Asp
Lys Thr Val Glu Arg Lys Cys 210 215 220Cys Val Glu Cys Pro Pro Cys
Pro Ala Pro Pro Val Ala Gly Pro Ser225 230 235 240Val Phe Leu Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro 260 265
270Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
275 280 285Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg
Val Val 290 295 300Ser Val Leu Thr Val Val His Gln Asp Trp Leu Asn
Gly Lys Glu Tyr305 310 315 320Lys Cys Lys Val Ser Asn Lys Gly Leu
Pro Ala Pro Ile Glu Lys Thr 325 330 335Ile Ser Lys Thr Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu 340 345 350Pro Pro Ser Arg Glu
Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380Asn
Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp385 390
395 400Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser 405 410 415Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala 420 425 430Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys 435 440 44518446PRTArtificialAn artificially
synthesized peptide sequence 18Glu Val Gln Leu Leu Glu Ser Gly Gly
Gly Leu Val Gln Pro Gly Gly1 5 10 15Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30Ala Met Ser Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45Ser Gly Ile Thr Gly Ser
Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val 50 55 60Lys Gly Arg Phe Thr
Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65 70 75 80Leu Gln Met
Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95Ala Lys
Asp Pro Gly Thr Thr Val Ile Met Ser Trp Phe Asp Pro Trp 100 105
110Gly Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro
115 120 125Ser Val Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly
Gly Thr 130 135 140Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro
Glu Pro Val Thr145 150 155 160Val Ser Trp Asn Ser Gly Ala Leu Thr
Ser Gly Val His Thr Phe Pro 165 170 175Ala Val Leu Gln Ser Ser Gly
Leu Tyr Ser Leu Ser Ser Val Val Thr 180 185 190Val Pro Ser Ser Asn
Phe Gly Thr Gln Thr Tyr Thr Cys Asn Val Asp 195 200 205His Lys Pro
Ser Asn Thr Lys Val Asp Lys Thr Val Glu Arg Lys Ser 210 215 220Cys
Val Glu Cys Pro Pro Cys Pro Ala Pro Pro Val Ala Gly Pro Ser225 230
235 240Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser
Arg 245 250 255Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln
Glu Asp Pro 260 265 270Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val
Glu Val His Asn Ala 275 280 285Lys Thr Lys Pro Arg Glu Glu Gln Phe
Asn Ser Thr Phe Arg Val Val 290 295 300Ser Val Leu Thr Val Val His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr305 310 315 320Lys Cys Lys Val
Ser Asn Lys Gly Leu Pro Ala Pro Ile Glu Lys Thr 325 330 335Ile Ser
Lys Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu 340 345
350Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys
355 360 365Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser 370 375 380Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Met Leu Asp385 390 395 400Ser Asp Gly Ser
Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser 405 410 415Arg Trp
Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala 420 425
430Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro 435 440
44519443PRTArtificialAn artificially synthesized peptide sequence
19Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Arg Pro Ser Gln1
5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr Ser Ile Thr Ser
Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro Gly Arg Gly Leu
Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile Thr Thr Tyr Asn
Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg Asp Thr Ser Lys
Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala Arg Thr Thr Ala
Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val Thr Val Ser Ser
Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala Pro Ser
Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150 155
160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu
165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro Ser 180 185 190Ser Asn Phe Gly Thr Gln Thr Tyr Thr Cys Asn Val
Asp His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Thr Val Glu
Arg Lys Cys Ser Val Glu 210 215 220Cys Pro Pro Cys Pro Ala Pro Pro
Val Ala Gly Pro Ser Val Phe Leu225 230 235 240Phe Pro Pro Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu 245 250 255Val Thr Cys
Val Val Val Asp Val Ser Gln Glu Asp Pro Glu Val Gln 260 265 270Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys 275 280
285Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu
290 295 300Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys305 310 315 320Val Ser Asn Lys Gly Leu Pro Ala Pro Ile Glu
Lys Thr Ile Ser Lys 325 330 335Thr Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser 340 345 350Gln Glu Glu Met Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys 355 360 365Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln 370 375 380Pro Glu Asn
Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp Ser Asp Gly385 390 395
400Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln
405 410 415Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn 420 425 430His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro 435
44020443PRTArtificialAn artificially synthesized peptide sequence
20Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Arg Pro Ser Gln1
5 10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Tyr Ser Ile Thr Ser
Asp 20 25 30His Ala Trp Ser Trp Val Arg Gln Pro Pro Gly Arg Gly Leu
Glu Trp 35 40 45Ile Gly Tyr Ile Ser Tyr Ser Gly Ile Thr Thr Tyr Asn
Pro Ser Leu 50 55 60Lys Ser Arg Val Thr Met Leu Arg Asp Thr Ser Lys
Asn Gln Phe Ser65 70 75 80Leu Arg Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr Cys 85 90 95Ala Arg Ser Leu Ala Arg Thr Thr Ala
Met Asp Tyr Trp Gly Gln Gly 100 105 110Ser Leu Val Thr Val Ser Ser
Ala Ser Thr Lys Gly Pro Ser Val Phe 115 120 125Pro Leu Ala Pro Ser
Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu 130 135 140Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp145 150 155
160Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu
165 170 175Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro Ser 180 185 190Ser Asn Phe Gly Thr Gln Thr Tyr Thr Cys Asn Val
Asp His Lys Pro 195 200 205Ser Asn Thr Lys Val Asp Lys Thr Val Glu
Arg Lys Cys Ser Val Glu 210 215 220Cys Pro Pro Cys Pro Ala Pro Pro
Val Ala Gly Pro Ser Val Phe Leu225 230 235 240Phe Pro Pro Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu 245 250 255Val Thr Cys
Val Val Val Asp Val Ser Gln Glu Asp Pro Glu Val Gln 260 265 270Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys 275 280
285Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu
290 295 300Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys305 310 315 320Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu
Lys Thr Ile Ser Lys 325 330 335Ala Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser 340 345 350Gln Glu Glu Met Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys 355 360 365Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln 370 375 380Pro Glu Asn
Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp Ser Asp Gly385 390 395
400Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln
405 410 415Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn 420 425 430His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro 435
44021213PRTArtificialAn artificially synthesized peptide sequence
21Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Thr Ser Glu Asn Ile Tyr Ser
Phe 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
Leu Ile 35 40 45Tyr Asn Ala Lys Thr Leu Ala Lys Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln His
His Tyr Glu Ser Pro Leu 85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro
Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val
Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln
Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155
160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205Phe Asn Arg Gly Cys
21022213PRTArtificialAn artificially synthesized peptide sequence
22Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys Arg Thr Ser Glu Asn Ile Tyr Ser
Phe 20 25 30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
Leu Ile 35 40 45Tyr Asn Ala Lys Thr Leu Ala Lys Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75 80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln His
His Tyr Glu Ser Pro Leu 85 90 95Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys Arg Thr Val Ala Ala 100 105 110Pro Ser Val Phe Ile Phe Pro
Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125Thr Ala Ser Val Val
Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140Lys Val Gln
Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln145 150 155
160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205Phe Asn Arg Glu Cys 21023330PRTHomo
sapiens 23Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser
Ser Lys1 5 10 15Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val
Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly
Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser
Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser
Ser Leu Gly Thr Gln Thr65 70 75 80Tyr Ile Cys Asn Val Asn His Lys
Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Lys Val Glu Pro Lys Ser Cys
Asp Lys Thr His Thr Cys Pro Pro Cys 100 105 110Pro Ala Pro Glu Leu
Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro 115 120 125Lys Pro Lys
Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys 130 135 140Val
Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp145 150
155 160Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg
Glu 165 170 175Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu
Thr Val Leu 180 185 190His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys Val Ser Asn 195 200 205Lys Ala Leu Pro Ala Pro Ile Glu Lys
Thr Ile Ser Lys Ala Lys Gly 210 215 220Gln Pro Arg Glu Pro Gln Val
Tyr Thr Leu Pro Pro Ser Arg Asp Glu225 230 235 240Leu Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr 245 250 255Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn 260 265
270Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe
275 280 285Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly Asn 290 295 300Val Phe Ser Cys Ser Val Met His Glu Ala Leu His
Asn His Tyr Thr305 310 315 320Gln Lys Ser Leu Ser Leu Ser Pro Gly
Lys 325 33024326PRTHomo sapiens 24Ala Ser Thr Lys Gly Pro Ser Val
Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala
Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr
Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe
Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val
Val Thr Val Pro Ser Ser Asn Phe Gly Thr Gln Thr65 70 75 80Tyr Thr
Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Thr
Val Glu Arg Lys Cys Cys Val Glu Cys Pro Pro Cys Pro Ala Pro 100 105
110Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
115 120 125Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp 130 135 140Val Ser His Glu Asp Pro Glu Val Gln Phe Asn Trp
Tyr Val Asp Gly145 150 155 160Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Phe Asn 165 170 175Ser Thr Phe Arg Val Val Ser
Val Leu Thr Val Val His Gln Asp Trp 180 185 190Leu Asn Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro 195 200 205Ala Pro Ile
Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu 210 215 220Pro
Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn225 230
235 240Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile 245 250 255Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr 260 265 270Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys 275 280 285Leu Thr Val Asp Lys Ser Arg Trp Gln
Gln Gly Asn Val Phe Ser Cys 290 295 300Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser Leu305 310 315 320Ser Leu Ser Pro
Gly Lys 32525327PRTHomo sapiens 25Ala Ser Thr Lys Gly Pro Ser Val
Phe Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala
Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr
Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe
Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val
Val Thr Val Pro Ser Ser Ser Leu Gly Thr Lys Thr65 70 75 80Tyr Thr
Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Arg
Val Glu Ser Lys Tyr Gly Pro Pro Cys Pro Pro Cys Pro Ala Pro 100 105
110Glu Phe Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys
115 120 125Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val
Val Val 130 135 140Asp Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn
Trp Tyr Val Asp145 150 155 160Gly Val Glu Val His Asn Ala Lys Thr
Lys Pro Arg Glu Glu Gln Phe 165 170 175Asn Ser Thr Tyr Arg Val Val
Ser Val Leu Thr Val Leu His Gln Asp 180 185 190Trp Leu Asn Gly Lys
Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu 195 200 205Pro Ser Ser
Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 210 215 220Glu
Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys225 230
235 240Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp 245 250 255Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr Lys 260 265 270Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser
Phe Phe Leu Tyr Ser 275 280 285Arg Leu Thr Val Asp Lys Ser Arg Trp
Gln Glu Gly Asn Val Phe Ser 290 295 300Cys Ser Val Met His Glu Ala
Leu His Asn His Tyr Thr Gln Lys Ser305 310 315 320Leu Ser Leu Ser
Leu Gly Lys 32526326PRTArtificialAn artificially synthesized
peptide sequence 26Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala
Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu Gly Cys
Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn
Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu
Gln Ser Ser Gly Leu Tyr Ser 50 55
60Leu Ser Ser Val Val Thr Val Pro Ser Ser Asn Phe Gly Thr Gln Thr65
70 75 80Tyr Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp
Lys 85 90 95Thr Val Glu Arg Lys Ser Cys Val Glu Cys Pro Pro Cys Pro
Ala Pro 100 105 110Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp 115 120 125Thr Leu Met Ile Ser Arg Thr Pro Glu Val
Thr Cys Val Val Val Asp 130 135 140Val Ser His Glu Asp Pro Glu Val
Gln Phe Asn Trp Tyr Val Asp Gly145 150 155 160Val Glu Val His Asn
Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn 165 170 175Ser Thr Phe
Arg Val Val Ser Val Leu Thr Val Val His Gln Asp Trp 180 185 190Leu
Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro 195 200
205Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu
210 215 220Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr
Lys Asn225 230 235 240Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile 245 250 255Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr 260 265 270Thr Pro Pro Met Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Lys 275 280 285Leu Thr Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys 290 295 300Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu305 310 315
320Ser Leu Ser Pro Gly Lys 32527326PRTArtificialAn artificially
synthesized peptide sequence 27Ala Ser Thr Lys Gly Pro Ser Val Phe
Pro Leu Ala Pro Cys Ser Arg1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala
Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val
Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro
Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val
Thr Val Pro Ser Ser Asn Phe Gly Thr Gln Thr65 70 75 80Tyr Thr Cys
Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Thr Val
Glu Arg Lys Cys Ser Val Glu Cys Pro Pro Cys Pro Ala Pro 100 105
110Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
115 120 125Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp 130 135 140Val Ser His Glu Asp Pro Glu Val Gln Phe Asn Trp
Tyr Val Asp Gly145 150 155 160Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Phe Asn 165 170 175Ser Thr Phe Arg Val Val Ser
Val Leu Thr Val Val His Gln Asp Trp 180 185 190Leu Asn Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro 195 200 205Ala Pro Ile
Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu 210 215 220Pro
Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn225 230
235 240Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile 245 250 255Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr 260 265 270Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys 275 280 285Leu Thr Val Asp Lys Ser Arg Trp Gln
Gln Gly Asn Val Phe Ser Cys 290 295 300Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser Leu305 310 315 320Ser Leu Ser Pro
Gly Lys 32528326PRTArtificialAn artificially synthesized peptide
sequence 28Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser
Ser Lys1 5 10 15Ser Thr Ser Glu Ser Thr Ala Ala Leu Gly Cys Leu Val
Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly
Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser
Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser
Asn Phe Gly Thr Gln Thr65 70 75 80Tyr Thr Cys Asn Val Asp His Lys
Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Thr Val Glu Arg Lys Ser Cys
Val Glu Cys Pro Pro Cys Pro Ala Pro 100 105 110Pro Val Ala Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 115 120 125Thr Leu Met
Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 130 135 140Val
Ser His Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly145 150
155 160Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe
Asn 165 170 175Ser Thr Phe Arg Val Val Ser Val Leu Thr Val Val His
Gln Asp Trp 180 185 190Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Gly Leu Pro 195 200 205Ala Pro Ile Glu Lys Thr Ile Ser Lys
Thr Lys Gly Gln Pro Arg Glu 210 215 220Pro Gln Val Tyr Thr Leu Pro
Pro Ser Arg Glu Glu Met Thr Lys Asn225 230 235 240Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 245 250 255Ala Val
Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr 260 265
270Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
275 280 285Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe
Ser Cys 290 295 300Ser Val Met His Glu Ala Leu His Asn His Tyr Thr
Gln Lys Ser Leu305 310 315 320Ser Leu Ser Pro Gly Lys
32529324PRTArtificialAn artificially synthesized peptide sequence
29Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala Pro Ser Ser Lys1
5 10 15Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys Leu Val Lys Asp
Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu
Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly
Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro Ser Ser Asn Phe
Gly Thr Gln Thr65 70 75 80Tyr Thr Cys Asn Val Asp His Lys Pro Ser
Asn Thr Lys Val Asp Lys 85 90 95Thr Val Glu Arg Lys Ser Cys Val Glu
Cys Pro Pro Cys Pro Ala Pro 100 105 110Pro Val Ala Gly Pro Ser Val
Phe Leu Phe Pro Pro Lys Pro Lys Asp 115 120 125Thr Leu Met Ile Ser
Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 130 135 140Val Ser Gln
Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly145 150 155
160Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn
165 170 175Ser Thr Phe Arg Val Val Ser Val Leu Thr Val Val His Gln
Asp Trp 180 185 190Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn
Lys Gly Leu Pro 195 200 205Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr
Lys Gly Gln Pro Arg Glu 210 215 220Pro Gln Val Tyr Thr Leu Pro Pro
Ser Gln Glu Glu Met Thr Lys Asn225 230 235 240Gln Val Ser Leu Thr
Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 245 250 255Ala Val Glu
Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr 260 265 270Thr
Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys 275 280
285Leu Thr Val Asp Lys Ser Arg Trp Gln Glu Gly Asn Val Phe Ser Cys
290 295 300Ser Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys
Ser Leu305 310 315 320Ser Leu Ser Pro30324PRTArtificialAn
artificially synthesized peptide sequence 30Ala Ser Thr Lys Gly Pro
Ser Val Phe Pro Leu Ala Pro Ser Ser Lys1 5 10 15Ser Thr Ser Gly Gly
Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro
Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His
Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser
Ser Val Val Thr Val Pro Ser Ser Asn Phe Gly Thr Gln Thr65 70 75
80Tyr Thr Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val Asp Lys
85 90 95Thr Val Glu Arg Lys Cys Ser Val Glu Cys Pro Pro Cys Pro Ala
Pro 100 105 110Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys Asp 115 120 125Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val Asp 130 135 140Val Ser Gln Glu Asp Pro Glu Val Gln
Phe Asn Trp Tyr Val Asp Gly145 150 155 160Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn 165 170 175Ser Thr Phe Arg
Val Val Ser Val Leu Thr Val Val His Gln Asp Trp 180 185 190Leu Asn
Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro 195 200
205Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu
210 215 220Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr
Lys Asn225 230 235 240Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile 245 250 255Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr 260 265 270Thr Pro Pro Met Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Lys 275 280 285Leu Thr Val Asp Lys
Ser Arg Trp Gln Glu Gly Asn Val Phe Ser Cys 290 295 300Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu305 310 315
320Ser Leu Ser Pro31324PRTArtificialAn artificially synthesized
peptide sequence 31Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu Ala
Pro Ser Ser Lys1 5 10 15Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys
Leu Val Lys Asp Tyr 20 25 30Phe Pro Glu Pro Val Thr Val Ser Trp Asn
Ser Gly Ala Leu Thr Ser 35 40 45Gly Val His Thr Phe Pro Ala Val Leu
Gln Ser Ser Gly Leu Tyr Ser 50 55 60Leu Ser Ser Val Val Thr Val Pro
Ser Ser Asn Phe Gly Thr Gln Thr65 70 75 80Tyr Thr Cys Asn Val Asp
His Lys Pro Ser Asn Thr Lys Val Asp Lys 85 90 95Thr Val Glu Arg Lys
Cys Ser Val Glu Cys Pro Pro Cys Pro Ala Pro 100 105 110Pro Val Ala
Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp 115 120 125Thr
Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp 130 135
140Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp
Gly145 150 155 160Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu
Glu Gln Phe Asn 165 170 175Ser Thr Phe Arg Val Val Ser Val Leu Thr
Val Val His Gln Asp Trp 180 185 190Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn Lys Gly Leu Pro 195 200 205Ser Ser Ile Glu Lys Thr
Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu 210 215 220Pro Gln Val Tyr
Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys Asn225 230 235 240Gln
Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile 245 250
255Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr
260 265 270Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr
Ser Lys 275 280 285Leu Thr Val Asp Lys Ser Arg Trp Gln Glu Gly Asn
Val Phe Ser Cys 290 295 300Ser Val Met His Glu Ala Leu His Asn His
Tyr Thr Gln Lys Ser Leu305 310 315 320Ser Leu Ser Pro32107PRTHomo
sapiens 32Arg Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser
Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu
Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu
Ser Lys Ala Asp Tyr Glu65 70 75 80Lys His Lys Val Tyr Ala Cys Glu
Val Thr His Gln Gly Leu Ser Ser 85 90 95Pro Val Thr Lys Ser Phe Asn
Arg Gly Glu Cys 100 10533106PRTArtificialAn artificially
synthesized peptide sequence 33Arg Thr Val Ala Ala Pro Ser Val Phe
Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly Thr Ala Ser
Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu Ala Lys Val
Gln Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn Ser Gln Glu
Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser Leu Ser
Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75 80Lys His Lys
Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser 85 90 95Pro Val
Thr Lys Ser Phe Asn Arg Gly Cys 100 10534106PRTArtificialAn
artificially synthesized peptide sequence 34Arg Thr Val Ala Ala Pro
Ser Val Phe Ile Phe Pro Pro Ser Asp Glu1 5 10 15Gln Leu Lys Ser Gly
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe 20 25 30Tyr Pro Arg Glu
Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln 35 40 45Ser Gly Asn
Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser 50 55 60Thr Tyr
Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu65 70 75
80Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser
85 90 95Pro Val Thr Lys Ser Phe Asn Arg Glu Cys 100 10535267PRTHomo
sapiens 35Ala Glu Ser His Leu Ser Leu Leu Tyr His Leu Thr Ala Val
Ser Ser1 5 10 15Pro Ala Pro Gly Thr Pro Ala Phe Trp Val Ser Gly Trp
Leu Gly Pro 20 25 30Gln Gln Tyr Leu Ser Tyr Asn Ser Leu Arg Gly Glu
Ala Glu Pro Cys 35 40 45Gly Ala Trp Val Trp Glu Asn Gln Val Ser Trp
Tyr Trp Glu Lys Glu 50 55 60Thr Thr Asp Leu Arg Ile Lys Glu Lys Leu
Phe Leu Glu Ala Phe Lys65 70 75 80Ala Leu Gly Gly Lys Gly Pro Tyr
Thr Leu Gln Gly Leu Leu Gly Cys 85 90 95Glu Leu Gly Pro Asp Asn Thr
Ser Val Pro Thr Ala Lys Phe Ala Leu 100 105 110Asn Gly Glu Glu Phe
Met Asn Phe Asp Leu Lys Gln Gly Thr Trp Gly 115 120 125Gly Asp Trp
Pro Glu Ala Leu Ala Ile Ser Gln Arg Trp Gln Gln Gln 130 135 140Asp
Lys Ala Ala Asn Lys Glu Leu Thr Phe Leu Leu Phe Ser Cys Pro145 150
155 160His Arg Leu Arg Glu His Leu Glu Arg Gly Arg Gly Asn Leu Glu
Trp 165 170 175Lys Glu Pro Pro Ser Met Arg Leu Lys Ala Arg Pro Ser
Ser Pro Gly 180 185 190Phe Ser Val Leu Thr Cys Ser Ala Phe Ser Phe
Tyr Pro Pro Glu Leu 195 200 205Gln
Leu Arg Phe Leu Arg Asn Gly Leu Ala Ala Gly Thr Gly Gln Gly 210 215
220Asp Phe Gly Pro Asn Ser Asp Gly Ser Phe His Ala Ser Ser Ser
Leu225 230 235 240Thr Val Lys Ser Gly Asp Glu His His Tyr Cys Cys
Ile Val Gln His 245 250 255Ala Gly Leu Ala Gln Pro Leu Arg Val Glu
Leu 260 2653699PRTHomo sapiens 36Ile Gln Arg Thr Pro Lys Ile Gln
Val Tyr Ser Arg His Pro Ala Glu1 5 10 15Asn Gly Lys Ser Asn Phe Leu
Asn Cys Tyr Val Ser Gly Phe His Pro 20 25 30Ser Asp Ile Glu Val Asp
Leu Leu Lys Asn Gly Glu Arg Ile Glu Lys 35 40 45Val Glu His Ser Asp
Leu Ser Phe Ser Lys Asp Trp Ser Phe Tyr Leu 50 55 60Leu Tyr Tyr Thr
Glu Phe Thr Pro Thr Glu Lys Asp Glu Tyr Ala Cys65 70 75 80Arg Val
Asn His Val Thr Leu Ser Gln Pro Lys Ile Val Lys Trp Asp 85 90 95Arg
Asp Met37105PRTHomo sapiens 37Gln Pro Lys Ala Ala Pro Ser Val Thr
Leu Phe Pro Pro Ser Ser Glu1 5 10 15Glu Leu Gln Ala Asn Lys Ala Thr
Leu Val Cys Leu Ile Ser Asp Phe 20 25 30Tyr Pro Gly Ala Val Thr Val
Ala Trp Lys Ala Asp Ser Ser Pro Val 35 40 45Lys Ala Gly Val Glu Thr
Thr Thr Pro Ser Lys Gln Ser Asn Asn Lys 50 55 60Tyr Ala Ala Ser Ser
Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys Ser65 70 75 80His Arg Ser
Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr Val Glu 85 90 95Lys Thr
Val Ala Pro Thr Glu Cys Ser 100 10538104PRTArtificialAn
artificially synthesized peptide sequence 38Gln Pro Lys Ala Ala Pro
Ser Val Thr Leu Phe Pro Pro Ser Ser Glu1 5 10 15Glu Leu Gln Ala Asn
Lys Ala Thr Leu Val Cys Leu Ile Ser Asp Phe 20 25 30Tyr Pro Gly Ala
Val Thr Val Ala Trp Lys Ala Asp Ser Ser Pro Val 35 40 45Lys Ala Gly
Val Glu Thr Thr Thr Pro Ser Lys Gln Ser Asn Asn Lys 50 55 60Tyr Ala
Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys Ser65 70 75
80His Arg Ser Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr Val Glu
85 90 95Lys Thr Val Ala Pro Thr Cys Ser 10039104PRTArtificialAn
artificially synthesized peptide sequence 39Gln Pro Lys Ala Ala Pro
Ser Val Thr Leu Phe Pro Pro Ser Ser Glu1 5 10 15Glu Leu Gln Ala Asn
Lys Ala Thr Leu Val Cys Leu Ile Ser Asp Phe 20 25 30Tyr Pro Gly Ala
Val Thr Val Ala Trp Lys Ala Asp Ser Ser Pro Val 35 40 45Lys Ala Gly
Val Glu Thr Thr Thr Pro Ser Lys Gln Ser Asn Asn Lys 50 55 60Tyr Ala
Ala Ser Ser Tyr Leu Ser Leu Thr Pro Glu Gln Trp Lys Ser65 70 75
80His Arg Ser Tyr Ser Cys Gln Val Thr His Glu Gly Ser Thr Val Glu
85 90 95Lys Thr Val Ala Pro Glu Cys Ser 10040107PRTMus spp. 40Arg
Ala Asp Ala Ala Pro Thr Val Ser Ile Phe Pro Pro Ser Ser Glu1 5 10
15Gln Leu Thr Ser Gly Gly Ala Ser Val Val Cys Phe Leu Asn Asn Phe
20 25 30Tyr Pro Lys Asp Ile Asn Val Lys Trp Lys Ile Asp Gly Ser Glu
Arg 35 40 45Gln Asn Gly Val Leu Asn Ser Trp Thr Asp Gln Asp Ser Lys
Asp Ser 50 55 60Thr Tyr Ser Met Ser Ser Thr Leu Thr Leu Thr Lys Asp
Glu Tyr Glu65 70 75 80Arg His Asn Ser Tyr Thr Cys Glu Ala Thr His
Lys Thr Ser Thr Ser 85 90 95Pro Ile Val Lys Ser Phe Asn Arg Asn Glu
Cys 100 10541107PRTRattus spp. 41Arg Ala Asp Ala Ala Pro Thr Val
Ser Ile Phe Pro Pro Ser Thr Glu1 5 10 15Gln Leu Ala Thr Gly Gly Ala
Ser Val Val Cys Leu Met Asn Asn Phe 20 25 30Tyr Pro Arg Asp Ile Ser
Val Lys Trp Lys Ile Asp Gly Thr Glu Arg 35 40 45Arg Asp Gly Val Leu
Asp Ser Val Thr Asp Gln Asp Ser Lys Asp Ser 50 55 60Thr Tyr Ser Met
Ser Ser Thr Leu Ser Leu Thr Lys Ala Asp Tyr Glu65 70 75 80Ser His
Asn Leu Tyr Thr Cys Glu Val Val His Lys Thr Ser Ser Ser 85 90 95Pro
Val Val Lys Ser Phe Asn Arg Asn Glu Cys 100 10542104PRTOryctolagus
cuniculus 42Arg Asp Pro Val Ala Pro Thr Val Leu Ile Phe Pro Pro Ala
Ala Asp1 5 10 15Gln Val Ala Thr Gly Thr Val Thr Ile Val Cys Val Ala
Asn Lys Tyr 20 25 30Phe Pro Asp Val Thr Val Thr Trp Glu Val Asp Gly
Thr Thr Gln Thr 35 40 45Thr Gly Ile Glu Asn Ser Lys Thr Pro Gln Asn
Ser Ala Asp Cys Thr 50 55 60Tyr Asn Leu Ser Ser Thr Leu Thr Leu Thr
Ser Thr Gln Tyr Asn Ser65 70 75 80His Lys Glu Tyr Thr Cys Lys Val
Thr Gln Gly Thr Thr Ser Val Val 85 90 95Gln Ser Phe Asn Arg Gly Asp
Cys 10043106PRTOryctolagus cuniculus 43Arg Asp Pro Val Ala Pro Ser
Val Leu Leu Phe Pro Pro Ser Lys Glu1 5 10 15Glu Leu Thr Thr Gly Thr
Ala Thr Ile Val Cys Val Ala Asn Lys Phe 20 25 30Tyr Pro Ser Asp Ile
Thr Val Thr Trp Lys Val Asp Gly Thr Thr Gln 35 40 45Gln Ser Gly Ile
Glu Asn Ser Lys Thr Pro Gln Ser Pro Glu Asp Asn 50 55 60Thr Tyr Ser
Leu Ser Ser Thr Leu Ser Leu Thr Ser Ala Gln Tyr Asn65 70 75 80Ser
His Ser Val Tyr Thr Cys Glu Val Val Gln Gly Ser Ala Ser Pro 85 90
95Ile Val Gln Ser Phe Asn Arg Gly Asp Cys 100 105
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014
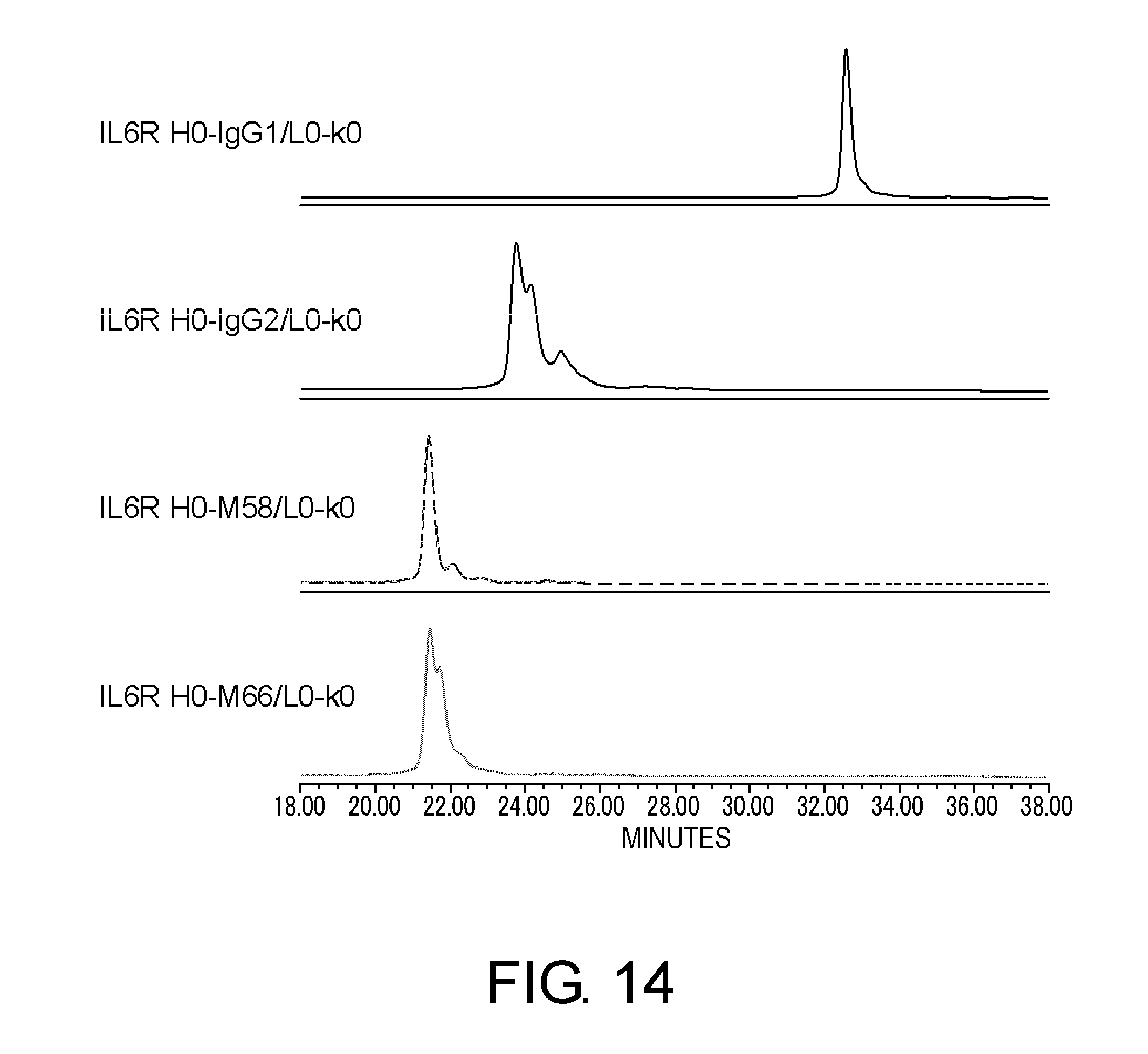
D00015
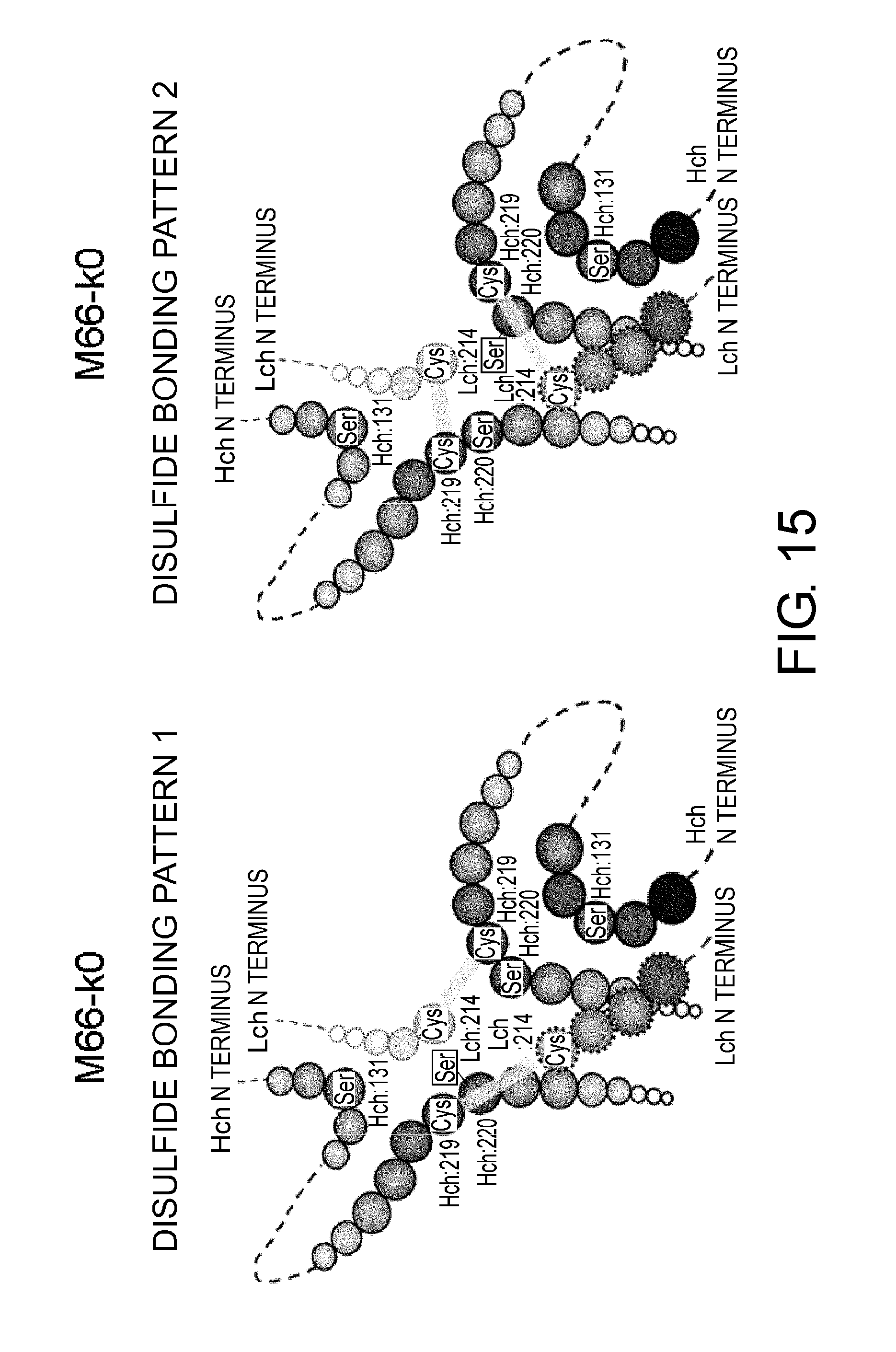
D00016

D00017
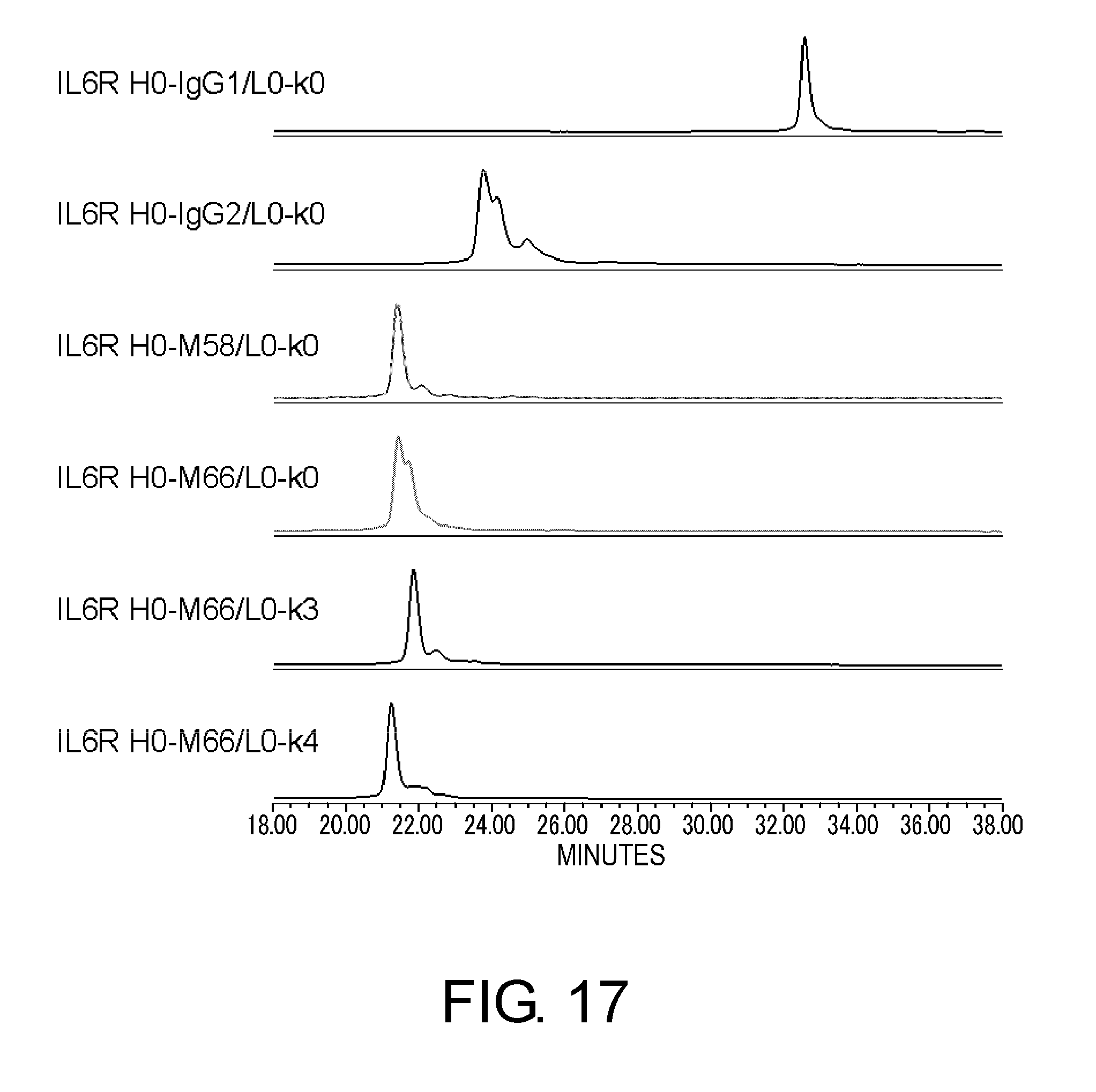
D00018
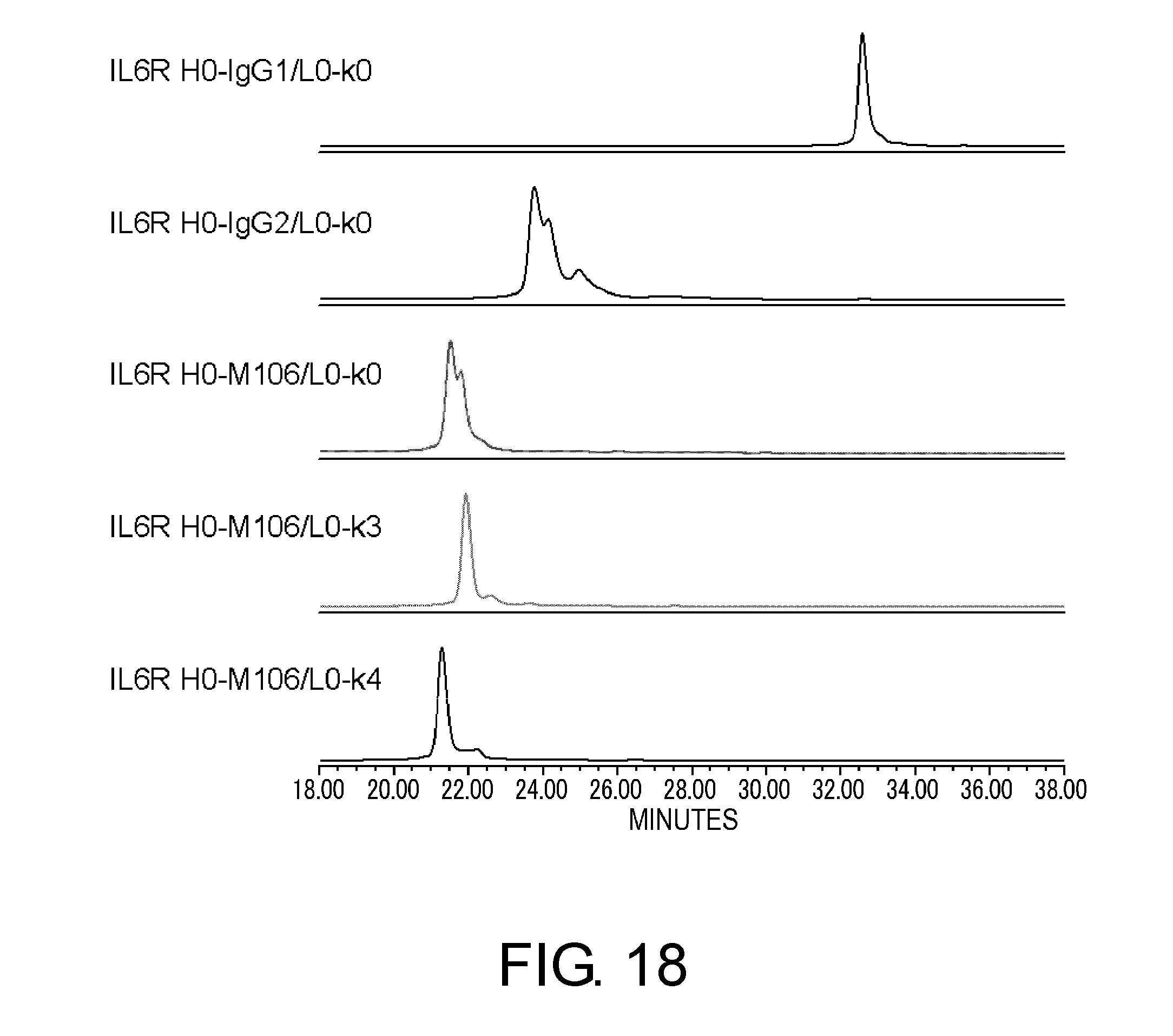
D00019
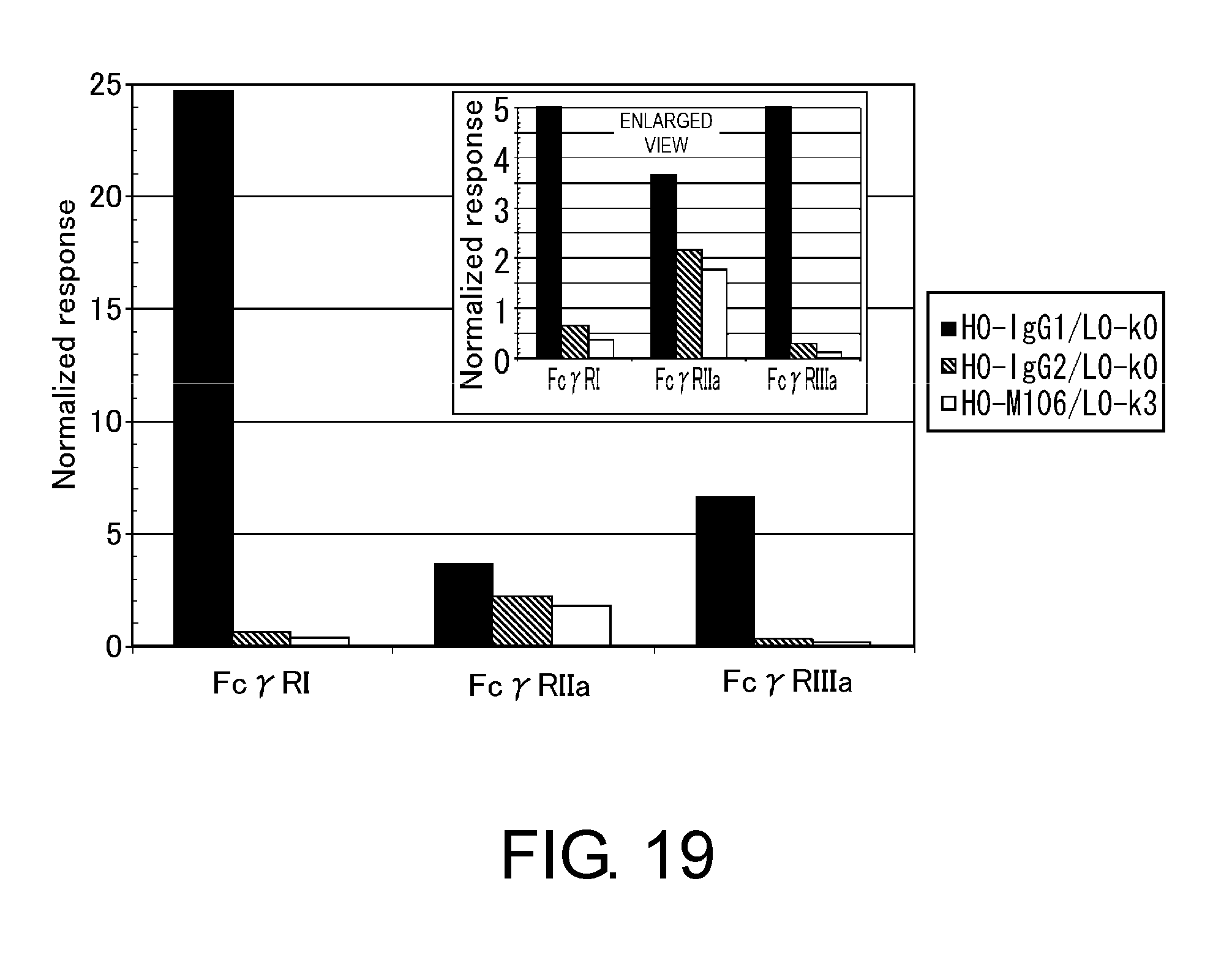
D00020
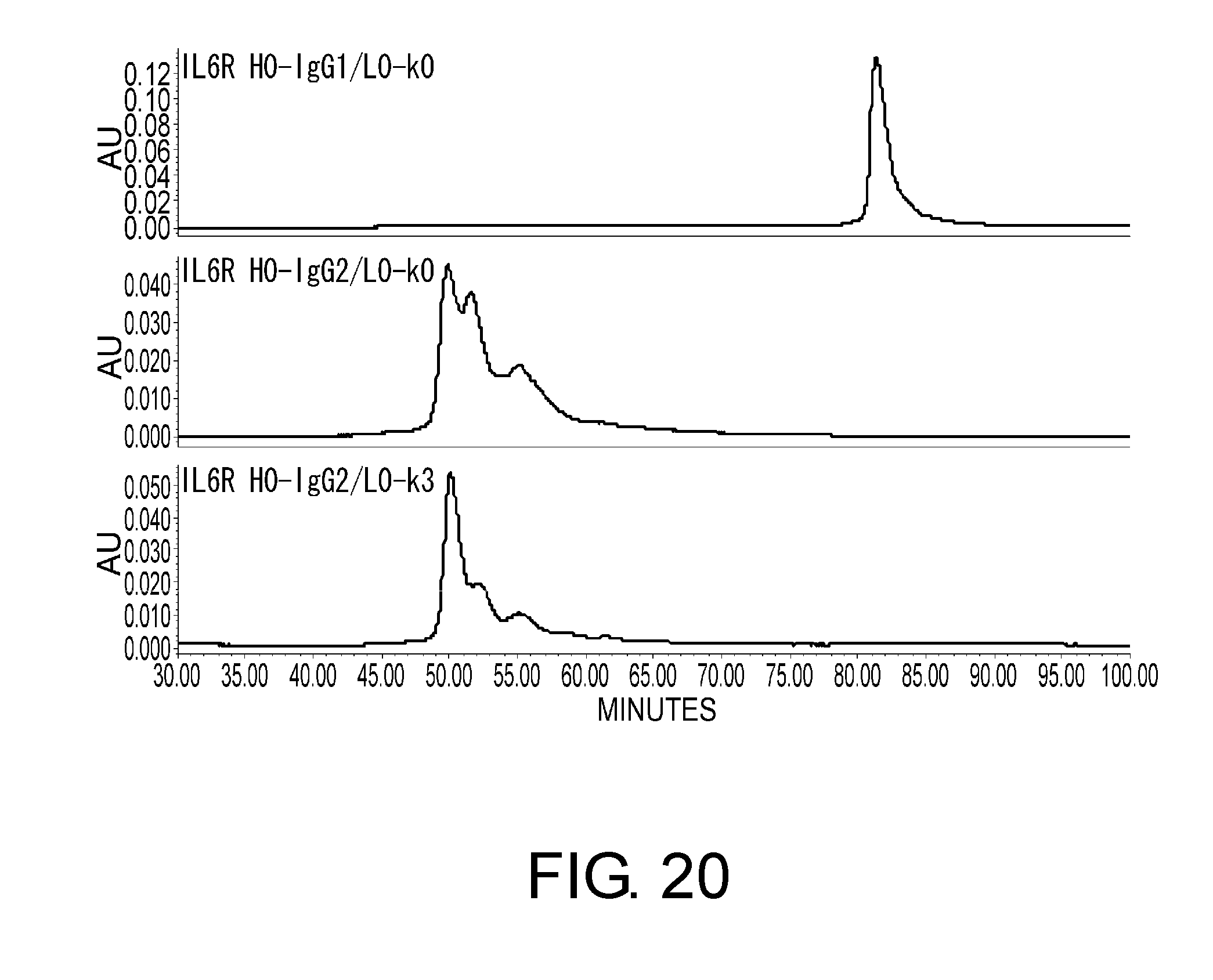
D00021
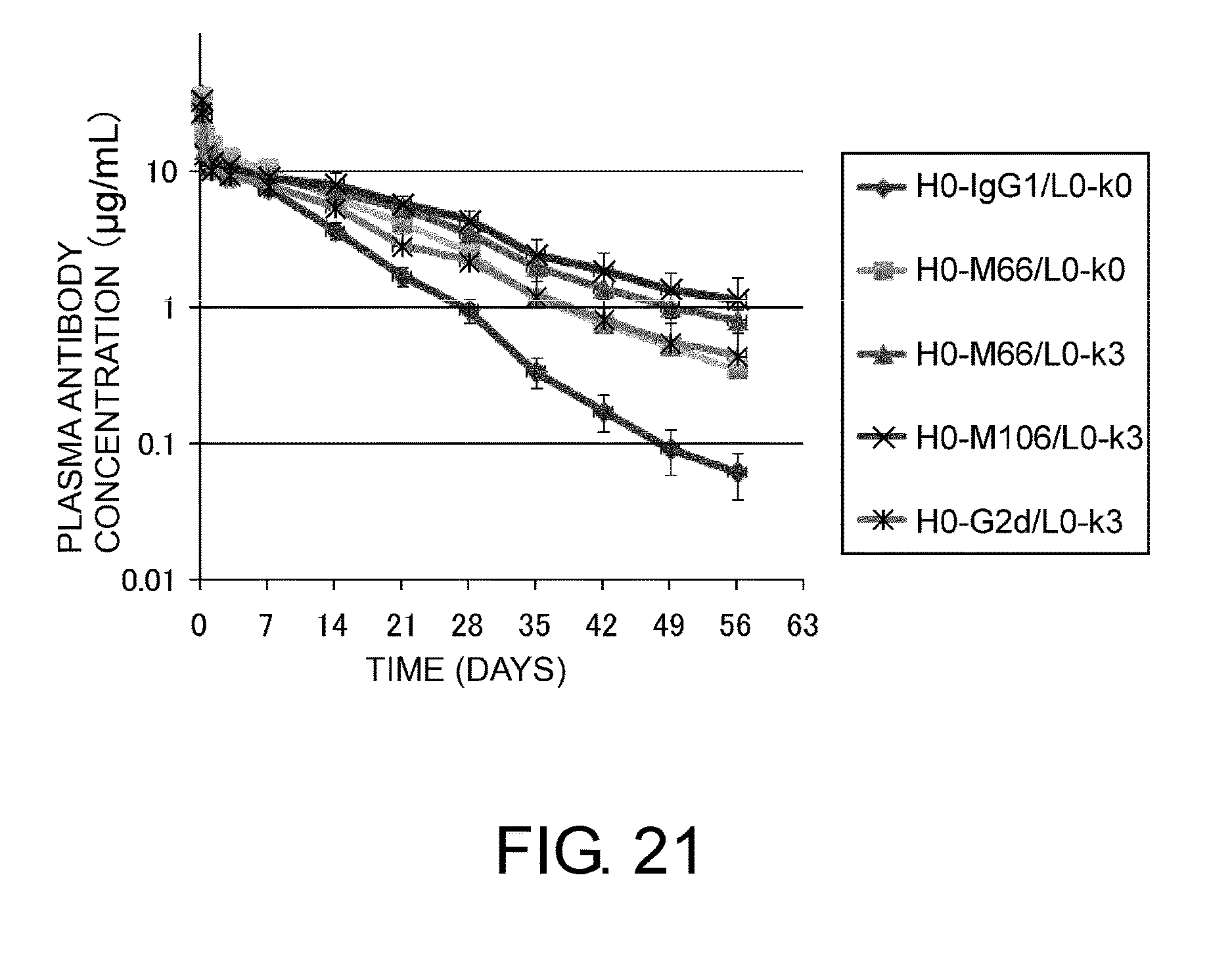
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.