Acyl-acp Thioesterases And Mutants Thereof
Casolari; Jason ; et al.
U.S. patent application number 16/185705 was filed with the patent office on 2019-06-27 for acyl-acp thioesterases and mutants thereof. The applicant listed for this patent is Corbion Biotech, Inc.. Invention is credited to Jason Casolari, Scott Franklin, George N. Rudenko, Xinhua Zhao.
| Application Number | 20190194703 16/185705 |
| Document ID | / |
| Family ID | 55525191 |
| Filed Date | 2019-06-27 |












View All Diagrams
| United States Patent Application | 20190194703 |
| Kind Code | A1 |
| Casolari; Jason ; et al. | June 27, 2019 |
ACYL-ACP THIOESTERASES AND MUTANTS THEREOF
Abstract
Novel plant acyl-ACP thioesterase genes of the FatB and FatA classes and proteins encoded by these genes are disclosed. The genes are useful for constructing recombinant host cells having altered fatty acid profiles. Expression of the novel and/or mutated FATB and FATA genes is demonstrated in oleaginous microalga host cells. Furthermore, a method for producing an oil elevated in one or more of C12:0, C14:0, C16:0, C18:0 and/or C18:1 fatty acids includes transforming a cell with novel and/or mutated FATB and/or FATA genes, e.g., having an N-terminal deletion. The cells produce triglycerides with altered and useful fatty acid profiles.
| Inventors: | Casolari; Jason; (Palo Alto, CA) ; Rudenko; George N.; (Mountain View, CA) ; Franklin; Scott; (Woodside, CA) ; Zhao; Xinhua; (Dublin, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 55525191 | ||||||||||
| Appl. No.: | 16/185705 | ||||||||||
| Filed: | November 9, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14858527 | Sep 18, 2015 | 10125382 | ||
| 16185705 | ||||
| 62075168 | Nov 4, 2014 | |||
| 62052440 | Sep 18, 2014 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 9/0006 20130101; C12P 7/44 20130101; C12P 7/6463 20130101; C12N 9/16 20130101; C12N 15/52 20130101; C12N 15/8247 20130101; C12N 9/2402 20130101; C12Y 301/02014 20130101 |
| International Class: | C12P 7/64 20060101 C12P007/64; C12P 7/44 20060101 C12P007/44; C12N 15/52 20060101 C12N015/52; C12N 9/04 20060101 C12N009/04; C12N 9/16 20060101 C12N009/16; C12N 9/24 20060101 C12N009/24; C12N 15/82 20060101 C12N015/82 |
Claims
1. A cDNA or nucleic acid construct comprising a polynucleotide sequence encoding a heterologous regulatory element and a FatB acyl-ACP thioesterase gene operable to produce an altered fatty acid profile in an oil produced by a cell expressing the nucleic acid construct, wherein the FatB gene expresses a protein having an amino acid sequence having at least 65% identity to SEQ ID NO: 40 or SEQ ID NO: 42.
2. The cDNA or nucleic acid construct of claim 1, wherein the acyl-ACP thioesterase coding sequence comprises at least 65% identity to SEQ ID NO: 44 or any equivalent sequences by virtue of the degeneracy of the genetic code.
3. (canceled)
4. The nucleic acid construct of claim 1, further comprising a polynucleotide encoding a plastid targeting peptide with at least 65% identity to SEQ ID NO: 37.
5. (canceled)
6. A host cell capable of expressing the cDNA or nucleic acid construct of claim 1 so as to produce a triglyceride oil having an altered composition relative to a control cell without the construct, the oil optionally having an increase in C8-C12 fatty acids.
7. A host cell capable of expressing the cDNA or nucleic acid construct of claim 1 so as to produce a triglyceride oil having an altered composition relative to a control cell without the construct, the oil optionally having an increase in C10 and C12 fatty acids.
8. The host cell of claim 6, wherein the host cell is selected from the group consisting of a plant cell, a microbial cell, and a microalgal cell.
9. A method of producing a recombinant cell that produces an altered fatty acid profile, the method comprising transforming a cell with a cDNA or nucleic acid construct of claim 1.
10. A host cell produced by the method of claim 9.
11. The host cell of claim 10, wherein the host cell is selected from the group consisting of a plant cell, a microbial cell, and a microalgal cell.
12. A method for producing an oil or oil-derived product, the method comprising cultivating a host cell of claim 6 and extracting oil produced thereby, optionally wherein the cultivation is by heterotrophic growth on sugar.
13. The method of claim 12, further comprising producing a fatty acid, fuel, chemical, food, or other oil-derived product from the oil.
14. (canceled)
15. An oil produced by the method of claim 12, optionally having a fatty acid profile comprising at least 20% C8, C10, C12, C14, C16 or C18 fatty acids.
16. The oil of claim 15, wherein the oil is produced by a microalgae, has a microalgal sterol profile, and optionally, lacks C24-alpha sterols.
17. (canceled)
18. A method for producing an oil, the method comprising: (a) providing a plastidic, oleaginous cell, optionally a microbial cell, the cell expressing a functional, acyl-ACP thioesterase gene encoded by a FATB gene having a deletion mutation in a region corresponding to the region coding for amino acids 66-98 of SEQ ID NO: 8; (b) cultivating the cell to produce a cell-oil; and (c) isolating the cell-oil from the cell.
19. The method of claim 17, wherein the cell-oil is enriched in C12 due to the deletion.
20. The method of claim 18 wherein the FATB gene encodes a protein with at least 65% sequence identity to SEQ ID NO: 42.
21. (canceled)
22.-38. (canceled)
39. The oil produced by the method of claim 19, wherein the fatty acid profile of the cell-oil comprises at least 5% C12 fatty acids.
40. The oil produced by the method of claim 39, wherein the fatty acid profile of the cell-oil comprises at least 10% C12 fatty acids.
41. The oil produced by the method of claim 40, wherein the fatty acid profile of the cell-oil comprises at least 15% C12 fatty acids.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of U.S. application Ser. No. 14/858,527, filed on Sep. 18, 2015, which claims the benefit under 35 U.S.C. .sctn. 119(e) of U.S. Provisional Application No. 62/052,440, filed on Sep. 18, 2014, and U.S. Provisional Application No. 62/075,168, filed on Nov. 4, 2014, all of which are hereby incorporated herein in their entireties.
SEQUENCE LISTING
[0002] This instant application contains a Sequence Listing which has been submitted electronically in ASCCII format and is hereby incorporated by reference in its entirety. Said ASCII copy, created on Nov. 20, 2015, is named SOLAP0281412A01_SL.txt and is 163,325 bytes in size.
Background
[0003] Certain organisms including plants and some microalgae use a type II fatty acid biosynthetic pathway, characterized by the use of discrete, monofunctional enzymes for fatty acid synthesis. In contrast, mammals and fungi use a single, large, multifunctional protein.
[0004] Type II fatty acid biosynthesis typically involves extension of a growing acyl-ACP (acyl-carrier protein) chain by two carbon units followed by cleavage by an acyl-ACP thioesterase. In plants, two main classes of acyl-ACP thioesterases have been identified: (i) those encoded by genes of the FatA class, which tend to hydrolyze oleoyl-ACP into oleate (an 18:1 fatty acid) and ACP, and (ii) those encoded by genes of the FatB class, which liberate C8-C16 fatty acids from corresponding acyl-ACP molecules.
[0005] Different FatB genes from various plants have specificities for different acyl chain lengths. As a result, different gene products will produce different fatty acid profiles in plant seeds. See, U.S. Pat. Nos. 5,850,022; 5,723,761; 5,639,790; 5,807,893; 5,455,167; 5,654,495; 5,512,482;5,298,421;5,667,997; and 5,344,771; 5,304,481. Recently, FatB genes have been cloned into oleaginous microalgae to produce triglycerides with altered fatty acid profiles. See, WO2010/063032, WO2011/150411, WO2012/106560, and WO2013/158938.
SUMMARY
[0006] According to an embodiment, there is a nucleic acid construct comprising a polynucleotide sequence encoding a heterologous regulatory element and a FatB acyl-ACP thioesterase gene operable to produce an altered fatty acid profile in an oil produced by a cell expressing the nucleic acid construct. The FatB gene expresses a protein having an amino acid sequence having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% identity to any of SEQ ID NOS: 1-18 or an amino acid sequence encoding a plastid targeting peptide fused upstream of any of SEQ ID NOS: 10-18.
[0007] Optionally, the acyl-ACP thioesterase coding sequence of the nucleic acid construct comprises at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% identity to any of SEQ ID NOS: 19-36 or any equivalent sequences by virtue of the degeneracy of the genetic code. In varying embodiments, the protein further comprises an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A).
[0008] In varying embodiments, the construct can have a plastid targeting peptide with at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% identity to SEQ ID NO: 37. In varying embodiments, the construct can have a plastid targeting peptide with at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% identity to SEQ ID NO: 40.
[0009] The FatB gene can express an active acyl-ACP-thioesterase protein having an amino acid sequence having:
[0010] (a) greater than 94.5, 94.6, 94.7, 94.8, 94.9, 95, or 95.1% identity to SEQ ID NO: 5;
[0011] (b) greater than 95.7, 95.8, 95.9, 96, 96.1 or 96.2% identity to SEQ ID NO: 14;
[0012] (c) greater than 95.4, 95.5, 95.6, 95.7, 95.8, 95.9, or 96% identity to SEQ ID NO: 3;
[0013] (d) greater than 94.5, 94.6, 94.7, 94.8, 94.9, 95, or 95.1% identity to SEQ ID NO: 12;
[0014] (e) greater than 94.8, 94.9, 95, 95.1, 95.2, 95.3, or 95.4% identity to SEQ ID NO: 1;
[0015] (f) greater than 95.9, 96.0, 96.1, 96.2, 96.3 or 96.4% identity to SEQ ID NO: 10;
[0016] (g) greater than 94.5, 94.6, 94.7, 94.8, 94.9, 95, or 95.1% identity to SEQ ID NO: 6;
[0017] (h) greater than 95.7, 95.8, 95.9, 96, 96.1 or 96.2% identity to SEQ ID NO: 15;
[0018] (i) greater than 94.5, 94.6, 94.7, 94.8, 94.9, 95, or 95.1% identity to SEQ ID NO: 4;
[0019] (j) greater than 95.7, 95.8, 95.9, 96, 96.1 or 96.2% identity to SEQ ID NO: 13;
[0020] (k) greater than 94.3, 94.4, 94.5, 94.6, 94.7, 94.8, or 94.9% identity to SEQ ID NO: 2;
[0021] (l) greater than 94.9, 95, 95.1, 95.2, 95.3, 95.4, or 95.5% identity to SEQ ID NO: 11;
[0022] (m) greater than 93.5, 93.6, 93.7, 93.8, 93.9, 94.0, or 94.1% identity to SEQ ID NO: 7;
[0023] (n) greater than 92.8, 92.9, 93.0, 93.1, 93.2, 93.3, or 93.4% identity to SEQ ID NO: 16;
[0024] (o) greater than 86.5, 86.6, 86.7, 86.8, 86.9, 87, or 87.1% identity to SEQ ID NO: 8;
[0025] (p) greater than 85.1, 85.2, 85.3, 85.4, 85.5, 85.6 or 85.7% identity to SEQ ID NO: 17;
[0026] (q) greater than 88, 88.1, 88.2, 88.3, 88.4, 88.5, or 88.6% identity to SEQ ID NO: 9; or
[0027] (r) greater than 87.6, 87.7, 87.8, 87.9, 88, 88.1, or 88.2% identity to SEQ ID NO: 18.
[0028] In another embodiment, a host cell is capable of expressing the nucleic acid construct so as to produce a triglyceride oil having an altered composition relative to a control cell without the construct. Optionally the oil has an increase in C8-C12 fatty acids.
[0029] The host cell can be selected, without limitation, from a plant cell, a microbial cell, and a microalgal cell.
[0030] In a third embodiment, a recombinant host cell produces an altered fatty acid profile, using a method comprising transforming the host cell with the nucleic acid construct. The host cell can, without limitation, be a microbial cell, a plant cell, or a microalgal cell. In varying embodiments, the host cell expresses a nucleic acid encoding a protein having an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A), and produces at least 2-fold the amount of C18:0 and/or C18:1 fatty acids compared to a host cell that expresses the wild-type protein.
[0031] In a fourth embodiment, a method produces an oil or oil-derived product, by cultivating a host cell as mentioned above and extracting the oil produced. Optionally, the cultivation is by heterotrophic growth on sugar. Optionally, the method also includes producing a fatty acid, fuel, chemical, food, or other oil-derived product from the oil. Optionally, an oil is produced having a fatty acid profile comprising at least 20% C8, C10, C12, C14, C16 or C18 (e.g., C18:0 and/or C18:1) fatty acids. Where the oil is produced by a microalgae, the oil can have a microalgal sterol profile and optionally lack C24-alpha sterols. The oil can be used to produce an oil-derived product, optionally a fatty acid, fuel, chemical, food, or other oil-derived product from the oil produced by the above method.
[0032] In a fifth embodiment, there is a method for producing an oil. The method includes providing a plastidic, oleaginous cell, optionally a microbial cell expressing a functional, acyl-ACP thioesterase gene encoded by a FATB gene having a deletion mutation in a region corresponding to the region coding for amino acids 66-98 of SEQ ID NO: 8. cultivating the cell to produce a cell-oil, and isolating the cell-oil from the cell. The cell-oil can be enriched in C12 due to the deletion. The FATB gene can encode a protein with at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NOS: 40 to 43. The FATB gene can have least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NOS: 44 or 45 or equivalent sequence by virtue of the degeneracy of the genetic code.
[0033] In a sixth embodiment, a cDNA, gene, expression cassette or host cell comprising a polynucleotide encoding a FATB protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to any of SEQ ID NOS: 40 to 43.
[0034] In a seventh embodiment, a cDNA, gene, expression cassette or host cell comprises a polynucleotide having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NOS 44 or 45, or equivalent sequence by virtue of the degeneracy of the genetic code.
[0035] In an eight embodiment, a method of genetically engineering a cell includes expressing in the cell, a polynucleotide that encodes a protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to any of SEQ ID NOS: 40 to 43; or has at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NOS: 47 or 48, or equivalent sequence by virtue of the degeneracy of the genetic code.
[0036] In a further aspect, provided is a cDNA, gene, expression cassette or host cell comprising a polynucleotide encoding a FATA protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61 and wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A). In a further aspect, provided is a cDNA, gene, expression cassette or host cell comprising a polynucleotide encoding a FATA protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61, or equivalent sequence by virtue of the degeneracy of the genetic code and wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A). In a further aspect, provided is a method of genetically engineering a cell comprising expressing in the cell, a polynucleotide that (a) encodes a protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61, wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A).
[0037] In a further aspect, provided is a host cell capable of expressing the nucleic acid construct encoding a FATA protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61 and wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A) so as to produce a triglyceride oil having an altered composition relative to a control cell without the construct, the oil optionally having an increase in C18 fatty acids, including C18:0 and C18:1 fatty acids. In varying embodiments, the host cell is selected from a plant cell, a microbial cell, and a microalgal cell. In a further aspect, provided is a method of producing a recombinant cell that produces an altered fatty acid profile, the method comprising transforming the cell with a nucleic acid encoding a FATA protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61 and wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A). In a further aspect, provided is a host cell produced according to such a method. In some embodiments, the host cell is selected from a plant cell, a microbial cell, and a microalgal cell. In a further aspect, provided is a method for producing an oil or oil-derived product, the method comprising cultivating a host cell encoding a FATA protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61 and wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A) and extracting oil produced thereby, optionally wherein the cultivation is by heterotrophic growth on sugar. In varying embodiments, the methods further comprise producing a fatty acid, fuel, chemical, food, or other oil-derived product from the oil. In varying embodiments, the host cell produces at least 2-fold the amount of C18:0 and/or C18:1 fatty acids compared to a host cell that expresses the wild-type protein. In a further aspect, further provided is an oil produced by the method of expressing in a host cell a polynucleotide encoding a FATA protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61 and wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A), optionally having a fatty acid profile comprising at least 20% C8, C10, C12, C14, C16 or C18 (e.g., C18:0 and/or C18:1) fatty acids. In varying embodiments, the oil is produced by a microalgae, has a microalgal sterol profile, and/or optionally, lacks C24-alpha sterols. Further provided is an oil-derived product, optionally a fatty acid, fuel, chemical, food, or other oil-derived product from the oil produced by the method of expressing in a host cell a polynucleotide encoding a FATA protein having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO:61 and wherein the protein has an alanine (A) at one or both positions corresponding to position 126 of SEQ ID NO: 61 (D124A) and 211 of SEQ ID NO: 61 (D209A).
Description of Illustrative Embodiments
Definitions
[0038] As used with respect to nucleic acids, the term "isolated" refers to a nucleic acid that is free of at least one other component that is typically present with the naturally occurring nucleic acid. Thus, a naturally occurring nucleic acid is isolated if it has been purified away from at least one other component that occurs naturally with the nucleic acid.
[0039] A "cell oil" or "natural fat" shall mean a predominantly triglyceride oil obtained from an organism, where the oil has not undergone blending with another natural or synthetic oil, or fractionation so as to substantially alter the fatty acid profile of the triglyceride. In connection with an oil comprising triglycerides of a particular regiospecificity, the cell oil or natural fat has not been subjected to interesterification or other synthetic process to obtain that regiospecific triglyceride profile, rather the regiospecificity is produced naturally, by a cell or population of cells. In connection with a cell oil or natural fat, and as used generally throughout the present disclosure, the terms oil and fat are used interchangeably, except where otherwise noted. Thus, an "oil" or a "fat" can be liquid, solid, or partially solid at room temperature, depending on the makeup of the substance and other conditions. Here, the term "fractionation" means removing material from the oil in a way that changes its fatty acid profile relative to the profile produced by the organism, however accomplished. The terms "cell oil" and "natural fat" encompass such oils obtained from an organism, where the oil has undergone minimal processing, including refining, bleaching and/or degumming, which does not substantially change its triglyceride profile. A cell oil can also be a "noninteresterified cell oil", which means that the cell oil has not undergone a process in which fatty acids have been redistributed in their acyl linkages to glycerol and remain essentially in the same configuration as when recovered from the organism.
[0040] "Exogenous gene" shall mean a nucleic acid that codes for the expression of an RNA and/or protein that has been introduced into a cell (e.g. by transformation/transfection), and is also referred to as a "transgene". A cell comprising an exogenous gene may be referred to as a recombinant cell, into which additional exogenous gene(s) may be introduced. The exogenous gene may be from a different species (and so heterologous), or from the same species (and so homologous), relative to the cell being transformed. Thus, an exogenous gene can include a homologous gene that occupies a different location in the genome of the cell or is under different control, relative to the endogenous copy of the gene. An exogenous gene may be present in more than one copy in the cell. An exogenous gene may be maintained in a cell, for example, as an insertion into the genome (nuclear or plastid) or as an episomal molecule.
[0041] "Fatty acids" shall mean free fatty acids, fatty acid salts, or fatty acyl moieties in a glycerolipid. It will be understood that fatty acyl groups of glycerolipids can be described in terms of the carboxylic acid or anion of a carboxylic acid that is produced when the triglyceride is hydrolyzed or saponified.
[0042] "Microalgae" are microbial organisms that contain a chloroplast or other plastid, and optionally that are capable of performing photosynthesis, or a prokaryotic microbial organism capable of performing photosynthesis. Microalgae include obligate photoautotrophs, which cannot metabolize a fixed carbon source as energy, as well as heterotrophs, which can live solely off of a fixed carbon source. Microalgae include unicellular organisms that separate from sister cells shortly after cell division, such as Chlamydomonas, as well as microbes such as, for example, Volvox, which is a simple multicellular photosynthetic microbe of two distinct cell types. Microalgae include cells such as Chlorella, Dunaliella, and Prototheca. Microalgae also include other microbial photosynthetic organisms that exhibit cell-cell adhesion, such as Agmenellum, Anabaena, and Pyrobotrys. Microalgae also include obligate heterotrophic microorganisms that have lost the ability to perform photosynthesis, such as certain dinoflagellate algae species and species of the genus Prototheca.
[0043] An "oleaginous" cell is a cell capable of producing at least 20% lipid by dry cell weight, naturally or through recombinant or classical strain improvement. An "oleaginous microbe" or "oleaginous microorganism" is a microbe, including a microalga that is oleaginous.
[0044] The term "percent sequence identity," in the context of two or more amino acid or nucleic acid sequences, refers to two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues or nucleotides that are the same, when compared and aligned for maximum correspondence, as measured using a sequence comparison algorithm or by visual inspection. For sequence comparison to determine percent nucleotide or amino acid identity, typically one sequence acts as a reference sequence, to which test sequences are compared. When using a sequence comparison algorithm, test and reference sequences are input into a computer, subsequence coordinates are designated, if necessary, and sequence algorithm program parameters are designated. The sequence comparison algorithm then calculates the percent sequence identity for the test sequence(s) relative to the reference sequence, based on the designated program parameters. Optimal alignment of sequences for comparison can be conducted using the NCBI BLAST software (ncbi.nlm.nih.gov/BLAST/) set to default parameters. For example, to compare two nucleic acid sequences, one may use blastn with the "BLAST 2 Sequences" tool Version 2.0.12 (Apr. 21, 2000) set at the following default parameters: Matrix: BLOSUM62; Reward for match: 1; Penalty for mismatch: -2; Open Gap: 5 and Extension Gap: 2 penalties; Gap x drop-off: 50; Expect: 10; Word Size: 11; Filter: on. For a pairwise comparison of two amino acid sequences, one may use the "BLAST 2 Sequences" tool Version 2.0.12 (Apr. 21, 2000) with blastp set, for example, at the following default parameters: Matrix: BLOSUM62; Open Gap: 11 and Extension Gap: 1 penalties; Gap x drop-off 50; Expect: 10; Word Size: 3; Filter: on.
[0045] Numbering of a given amino acid polymer or nucleic acid polymer "corresponds to" or is "relative to" the numbering of a selected amino acid polymer or nucleic acid polymer when the position of any given polymer component (e.g., amino acid, nucleotide, also referred to generically as a "residue") is designated by reference to the same or to an equivalent position (e.g., based on an optimal alignment or a consensus sequence) in the selected amino acid or nucleic acid polymer, rather than by the actual numerical position of the component in the given polymer.
[0046] A "variant" is a polypeptide comprising a sequence which differs in one or more amino acid position(s) from that of a parent polypeptide sequence (e.g., by substitution, deletion, or insertion). A variant may comprise a sequence which differs from the parent polypeptides sequence in up to 40% of the total number of residues of the parent polypeptide sequence, such as in up to 40%, 35%, 30%, 25%, 20%, 15%, 10%, 9%, 8%, 7%, 6%, 5%, 4%, 3% 2% or 1% of the total number of residues of the parent polypeptide sequence. For example, a variant of a 400 amino acid polypeptide sequence comprises a sequence which differs in up to 40% of the total number of residues of the parent polypeptide sequence, that is, in up to 160 amino acid positions within the 400 amino acid polypeptide sequence (such as in 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 78, 79, 80, 85, 90, 95, 100, 105, 110, 115, 120, 125, 130, 135, 140, 145, 150, 155, or 160 amino acid positions within the reference sequence.
[0047] "Naturally occurring" as applied to a composition that can be found in nature as distinct from being artificially produced by man. For example, a polypeptide or polynucleotide that is present in an organism (including viruses, bacteria, protozoa, insects, plants or mammalian tissue) that can be isolated from a source in nature and which has not been intentionally modified by man in the laboratory is naturally occurring. "Non-naturally occurring" (also termed "synthetic" or "artificial") as applied to an object means that the object is not naturally-occurring--i.e., the object cannot be found in nature as distinct from being artificially produced by man.
[0048] In connection with a cell oil, a "profile" is the distribution of particular species or triglycerides or fatty acyl groups within the oil. A "fatty acid profile" is the distribution of fatty acyl groups in the triglycerides of the oil without reference to attachment to a glycerol backbone. Fatty acid profiles are typically determined by conversion to a fatty acid methyl ester (FAME), followed by gas chromatography (GC) analysis with flame ionization detection (FID). The fatty acid profile can be expressed as one or more percent of a fatty acid in the total fatty acid signal determined from the area under the curve for that fatty acid. FAME-GC-FID measurement approximate weight percentages of the fatty acids.
[0049] As used herein, an oil is said to be "enriched" in one or more particular fatty acids if there is at least a 10% increase in the mass of that fatty acid in the oil relative to the non-enriched oil. For example, in the case of a cell expressing a heterologous FatB gene described herein, the oil produced by the cell is said to be enriched in, e.g., C8 and C16 fatty acids if the mass of these fatty acids in the oil is at least 10% greater than in oil produced by a cell of the same type that does not express the heterologous FatB gene (e.g., wild type oil).
[0050] "Recombinant" is a cell, nucleic acid, protein or vector that has been modified due to the introduction of an exogenous nucleic acid or the alteration of a native nucleic acid. Thus, e.g., recombinant (host) cells can express genes that are not found within the native (non-recombinant) form of the cell or express native genes differently than those genes are expressed by a non-recombinant cell. Recombinant cells can, without limitation, include recombinant nucleic acids that encode a gene product or suppression elements such as mutations, knockouts, antisense, interfering RNA (RNAi) or dsRNA that reduce the levels of active gene product in a cell. A "recombinant nucleic acid" is a nucleic acid originally formed in vitro, in general, by the manipulation of nucleic acid, e.g., using polymerases, ligases, exonucleases, and endonucleases, using chemical synthesis, or otherwise is in a form not normally found in nature. Recombinant nucleic acids may be produced, for example, to place two or more nucleic acids in operable linkage. Thus, an isolated nucleic acid or an expression vector formed in vitro by nucleic by ligating DNA molecules that are not normally joined in nature, are both considered recombinant herein. Recombinant nucleic acids can also be produced in other ways; e.g., using chemical DNA synthesis. Once a recombinant nucleic acid is made and introduced into a host cell or organism, it may replicate using the in vivo cellular machinery of the host cell; however, such nucleic acids, once produced recombinantly, although subsequently replicated intracellularly, are still considered recombinant herein. Similarly, a "recombinant protein" is a protein made using recombinant techniques, i.e., through the expression of a recombinant nucleic acid.
[0051] Embodiments relate to the use of novel FatB acyl-ACP thioesterase genes (e.g. in the form of cDNA, vectors, and constructs in vitro or in host cells) gene-variants, and peptides isolated from plants which can be expressed in a host cell in order to alter the fatty acid profile of an oil produced by the cell. The genes were discovered by obtaining cDNA from various plant species and transforming a model organism --the obligate heterotrophic microalga, Prototheca moriformis. Although P. moriformis was used to screen the FatB genes for ability to the alter fatty acid profile, the genes and corresponding gene-products are useful in a wide variety of host cells. For example, the genes can be expressed in bacteria, other microalgae, or higher plants. The genes can be expressed in higher plants according to the methods of U.S. Pat. Nos. 5,850,022; 5,723,761; 5,639,790; 5,807,893; 5,455,167; 5,654,495; 5,512,482; 5,298,421; 5,667,997; 5,344,771; and 5,304,481. The fatty acids can be further converted to triglycerides, fatty aldehydes, fatty alcohols and other oleochemicals either synthetically or biosynthetically.
[0052] Additionally, in the course of obtaining the novel FatB sequences, we discovered that certain N-terminal deletions in the FatB cDNAs led to desirably altered fatty acid profiles in the microalgal model.
[0053] In an embodiment, there is a polynucleotide comprising a nucleic acid sequence operably linked to a heterologous expression control sequence, wherein said nucleic acid sequence is selected from the group consisting of SEQ ID NOs: 19-36, a sequence encoding the amino acid sequence of the group consisting of SEQ ID NOs: 1-18, or a variant thereof with acyl-ACP thioesterase activity when expressed in a plastidic oleaginous cell.
[0054] In an embodiment, triglycerides are produced by a host cell expressing a novel FatB gene of Table 1. A triglyceride-containing cell oil can be recovered from the host cell. The cell oil can be refined, degummed, bleached and/or deodorized. The oil, in its natural or processed form, can be used for foods, chemicals, fuels, cosmetics, plastics, and other uses.
[0055] The genes can be used in a variety of genetic constructs including plasmids or other vectors for expression or recombination in a host cell. The genes can be codon optimized for expression in a target host cell (e.g., using the codon usage tables of Tables 2-5.) For example, at least 60, 65, 70, 75, 80, 85, 90, 95 or 100% of the codons used can be the most preferred codon according to Table 2, 3, 4 or 5. Alternately, at least 60, 65, 70, 75, 80, 85, 90, 95 or 100% of the codons used can be the first or second most preferred codon according to Table 2, 3, or 5. The proteins produced by the genes can be used in vivo or in purified form.
[0056] For example, the gene can be prepared in an expression vector comprising an operably linked promoter and 5'UTR. Where a plastidic cell is used as the host, a suitably active plastid targeting peptide (also "transit peptide") can be fused to the FATB gene, as in the examples below. Transit peptides are denoted by underlined or outlined text in some of the FATB peptide sequences that appear below. Generally, for the newly identified FATB genes, there are roughly 50 amino acids at the N-terminal that constitute a plastid transit peptide, which are responsible for transporting the enzyme to the chloroplast. In the examples below, this transit peptide is replaced with a 38 amino acid sequence (SEQ ID NO: 37) that is effective in Prototheca moriformis host cells for transporting the enzyme to the plastids of those cells. Thus, we contemplate deletions and fusion proteins in order to optimize enzyme activity in a given host cell. For example, a transit peptide from the host or related species may be used instead of that of the newly discovered plant genes described here. In general, plastid targeting peptides are less conserved than the enzymatic domains of FATB genes. Plastid targeting peptides can be substituted with other sequences such as those found in plant-derived sequences of plastid targeting genes (e.g., those for FATA, FATB, SAD or KAS genes) in the ThYme database of thioesters-active enzymes hosted by Iowa State University/NSF Engineering Research Center for Biorenewable Chemicals. Accordingly, certain embodiments describe percent identity to gene or protein sequences to FATB genes lacking the plastid targeting peptide.
[0057] A selectable marker gene may be included in the vector to assist in isolating a transformed cell. Examples of selectable markers useful in microalgae include sucrose invertase, antibiotic resistance, and thiamine synthesis genes.
[0058] The gene sequences disclosed can also be used to prepare antisense, or inhibitory RNA (e.g., RNAi or hairpin RNA) to inhibit complementary genes in a plant or other organism.
[0059] FatB genes found to be useful in producing desired fatty acid profiles in a cell are summarized below in Table 1. Nucleic acids or proteins having the sequence of SEQ ID NOS: 19-36 can be used to alter the fatty acid profile of a recombinant cell. Variant nucleic acids can also be used; e.g., variants having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NOS: 19-36. Codon optimization of the genes for a variety of host organisms is contemplated, as is the use of gene fragments. Preferred codons for Prototheca strains and for Chlorella protothecoides are shown below in Tables 2 and 3, respectively. Codon usage for Cuphea wrightii is shown in Table 4. Codon usage for Arabidopsis is shown in Table 5; for example, the most preferred of codon for each amino acid can be selected. Codon tables for other organisms including microalgae and higher plants are known in the art. In some embodiments, the first and/or second most preferred Prototheca codons are employed for codon optimization. In specific embodiments, the novel amino acid sequences contained in the sequence listings below are converted into nucleic acid sequences according to the preferred codon usage in Prototheca, Chlorella, Cuphea wrightii, or Arabidopsis as set forth in tables 2 through 5 or nucleic acid sequences having at least 65, 70, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to these derived nucleic acid sequences.
[0060] In embodiments, there is protein or a nucleic acid encoding a protein having any of SEQ ID NOS: 1-18. In an embodiment, there is protein or a nucleic acid encoding a protein having at least 80, 85, 85.1, 85.2, 85.3,85.4, 85.5, 86, 86.5, 86.6, 86.7, 87, 87.5, 87.6, 87.7, 87.8, 87.9, 88, 89, 90, 91, 92, 92.5, 92.6, 92.7, 92.8, 92.9, 93, 93.5, 93.6, 93.7, 93.8, 94, 94.1, 94.2, 94.3, 94.4, 94.5, 94.6, 94.7, 94.8, 94.9, 95, 95.1, 95.2, 95.3, 95.4, 95.6, 95.7, 95.8, 95.9, 96, 96.1, 96.2, 96.3, 96.4, 96.5, 97, 98, 99, or 100% sequence identity with any of SEQ ID NOS: 1-18. An embodiment comprises a fragment of any of the above-described proteins or nucleic acids (including fragments of protein or nucleic acid variants), wherein the protein fragment has acyl-ACP thioesterase activity or the nucleic acid fragment encodes such a protein fragment. In other embodiments, the fragment includes a domain of an acyl-ACP thioesterase that mediates a particular function, e.g., a specificity-determining domain. Illustrative fragments can be produced by C-terminal and/or N-terminal truncations and include at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% of the full-length sequences disclosed herein.
[0061] In certain embodiments, percent sequence identity for variants of the nucleic acids or proteins discussed above can be calculated by using the full-length nucleic acid sequence (e.g., one of SEQ ID NOS: 19-36) or full-length amino acid sequence (e.g., one of SEQ ID NOS: 1-18) as the reference sequence and comparing the full-length test sequence to this reference sequence. In some embodiments relating to fragments, percent sequence identity for variants of nucleic acid or protein fragments can be calculated over the entire length of the fragment.
[0062] The nucleic acids can be in isolated form, or part of a vector or other construct, chromosome or host cell. It has been found that is many cases the full length gene (and protein) is not needed; for example, deletion of some or all of the N-terminal hydrophobic domain (typically an 18 amino acid domain starting with LPDW) (SEQ ID NO:62) yields a still-functional gene. In addition, fusions of the specificity determining regions of the genes in Table 1 with catalytic domains of other acyl-ACP thioesterases can yield functional genes. Certain embodiments encompass functional fragments (e.g., specificity determining regions) of the disclosed nucleic acid or amino acids fused to heterologous acyl-ACP thioesterase nucleic acid or amino acid sequences, respectively.
TABLE-US-00001 TABLE 1 FatB genes and proteins according to embodiments Plant nucleic Amino acid Amino acid Native sequence acid sequence plant codon- sequence (without nucleic optimized for GENE SEQ ID targeting acid Prototheca (species, abbreviation) NO: peptide) sequence moriformis Cuphea crassiflora 1 10 19 28 (CcrasFATB1) Cuphea koehneana 2 11 20 29 (CkoeFATB3) Cuphea leptopoda 3 12 21 30 (CleptFATB1) Cuphea angustifolia 4 13 22 31 (CangFATB1) Cuphea llavea 5 14 23 32 (CllaFATB1) Cuphea lophostoma 6 15 24 33 (ClopFATB1) Sassafras albidum 7 16 25 34 FATB1 (SalFATB1) Sassafras albidum 8 17 26 35 FATB2 (SalFATB2) Lindera benzoin 9 18 27 36 FATB1 (LbeFATB1)
TABLE-US-00002 TABLE 2 Preferred codon usage in Prototheca strains Ala GCG 345 (0.36) Asn AAT 8 (0.04) GCA 66 (0.07) AAC 201 (0.96) GCT 101 (0.11) Pro CCG 161 (0.29) GCC 442 (0.46) CCA 49 (0.09) Cys TGT 12 (0.10) CCT 71 (0.13) TGC 105 (0.90) CCC 267 (0.49) Asp GAT 43 (0.12) Gln CAG 226 (0.82) GAC 316 (0.88) CAA 48 (0.18) Glu GAG 377 (0.96) Arg AGG 33 (0.06) GAA 14 (0.04) AGA 14 (0.02) Phe TTT 89 (0.29) CGG 102 (0.18) TTC 216 (0.71) CGA 49 (0.08) Gly GGG 92 (0.12) CGT 51 (0.09) GGA 56 (0.07) CGC 331 (0.57) GGT 76 (0.10) Ser AGT 16 (0.03) GGC 559 (0.71) AGC 123 (0.22) His CAT 42 (0.21) TCG 152 (0.28) CAC 154 (0.79) TCA 31 (0.06) Ile ATA 4 (0.01) TCT 55 (0.10) ATT 30 (0.08) TCC 173 (0.31) ATC 338 (0.91) Thr ACG 184 (0.38) Lys AAG 284 (0.98) ACA 24 (0.05) AAA 7 (0.02) ACT 21 (0.05) Leu TTG 26 (0.04) ACC 249 (0.52) TTA 3 (0.00) Val GTG 308 (0.50) CTG 447 (0.61) GTA 9 (0.01) CTA 20 (0.03) GTT 35 (0.06) CTT 45 (0.06) GTC 262 (0.43) CTC 190 (0.26) Trp TGG 107 (1.00) Met ATG 191 (1.00) Tyr TAT 10 (0.05) TAC 180 (0.95) Stop TGA/TAG/TAA
TABLE-US-00003 TABLE 3 Preferred codon usage in Chlorella protothecoides TTC (Phe) TAC (Tyr) TGC (Cys) TGA (Stop) TGG (Trp) CCC (Pro) CAC (His) CGC (Arg) CTG (Leu) CAG (Gin) ATC (Ile) ACC (Thr) GAC (Asp) TCC (Ser) ATG (Met) AAG (Lys) GCC (Ala) AAC (Asn) GGC (Gly) GTG (Val) GAG (Glu)
TABLE-US-00004 TABLE 4 Codon usage for Cuphea wrightii (codon, amino acid, frequency, per thousand, number) UUU F 0.48 19.5 (52) UCU S 0.21 19.5 (52) UAU Y 0.45 6.4 (17) UGU C 0.41 10.5 (28) UUC F 0.52 21.3 (57) UCC S 0.26 23.6 (63) UAC Y 0.55 7.9 (21) UGC C 0.59 15.0 (40) UUA L 0.07 5.2 (14) UCA S 0.18 16.8 (45) UAA * 0.33 0.7 (2) UGA * 0.33 0.7 (2) UUG L 0.19 14.6 (39) UCG S 0.11 9.7 (26) UAG * 0.33 0.7 (2) UGG W 1.00 15.4 (41) CUU L 0.27 21.0 (56) CCU P 0.48 21.7 (58) CAU H 0.60 11.2 (30) CGU R 0.09 5.6 (15) CUC L 0.22 17.2 (46) CCC P 0.16 7.1 (19) CAC H 0.40 7.5 (20) CGC R 0.13 7.9 (21) CUA L 0.13 10.1 (27) CCA P 0.21 9.7 (26) CAA Q 0.31 8.6 (23) CGA R 0.11 6.7 (18) CUG L 0.12 9.7 (26) CCG P 0.16 7.1 (19) CAG Q 0.69 19.5 (52) CGG R 0.16 9.4 (25) AUU I 0.44 22.8 (61) ACU T 0.33 16.8 (45) AAU N 0.66 31.4 (84) AGU S 0.18 16.1 (43) AUC I 0.29 15.4 (41) ACC T 0.27 13.9 (37) AAC N 0.34 16.5 (44) AGC S 0.07 6.0 (16) AUA I 10.27 13.9 (37) ACA T 0.26 13.5 (36) AAA K 0.42 21.0 (56) AGA R 0.24 14.2 (38) AUG M 1.00 28.1 (75) ACG T 0.14 7.1 (19) AAG K 0.58 29.2 (78) AGG R 0.27 16.1 (43) GUU V 0.28 19.8 (53) GCU A 0.35 31.4 (84) GAU D 0.63 35.9 (96) GGU G 0.29 26.6 (71) GUC V 0.21 15.0 (40) GCC A 0.20 18.0 (48) GAC D 0.37 21.0 (56) GGC G 0.20 18.0 (48) GUA V 0.14 10.1 (27) GCA A 0.33 29.6 (79) GAA E 0.41 18.3 (49) GGA G 0.35 31.4 (84) GUG V 0.36 25.1 (67) GCG A 0.11 9.7 (26) GAG E 0.59 26.2 (70) GGG G 0.16 14.2 (38)
TABLE-US-00005 TABLE 5 Codon usage for Arabidopsis (codon, amino acid, frequency, per thousand) UUU F 0.51 21.8 UCU S 0.28 25.2 UAU Y 0.52 14.6 UGU C 0.60 10.5 UUC F 0.49 20.7 UCC S 0.13 11.2 UAC Y 0.48 13.7 UGC C 0.40 7.2 UUA L 0.14 12.7 UCA S 0.20 18.3 UAA * 0.36 0.9 UGA * 0.44 1.2 UUG L 0.22 20.9 UCG S 0.10 9.3 UAG * 0.20 0.5 UGG W 1.00 12.5 CUU L 0.26 24.1 CCU P 0.38 18.7 CAU H 0.61 13.8 CGU R 0.17 9.0 CUC L 0.17 16.1 CCC P 0.11 5.3 CAC H 0.39 8.7 CGC R 0.07 3.8 CUA L 0.11 9.9 CCA P 0.33 16.1 CAA Q 0.56 19.4 CGA R 0.12 6.3 CUG L 0.11 9.8 CCG P 0.18 8.6 CAG Q 0.44 15.2 CGG R 0.09 4.9 AUU I 0.41 21.5 ACU T 0.34 17.5 AAU N 0.52 22.3 AGU S 0.16 14.0 AUC I 0.35 18.5 ACC T 0.20 10.3 AAC N 0.48 20.9 AGC S 0.13 11.3 AUA I 0.24 12.6 ACA T 0.31 15.7 AAA K 0.49 30.8 AGA R 0.35 19.0 AUG M 1.00 24.5 ACG T 0.15 7.7 AAG K 0.51 32.7 AGG R 0.20 11.0 GUU V 0.40 27.2 GCU A 0.43 28.3 GAU D 0.68 36.6 GGU G 0.34 22.2 GUC V 0.19 12.8 GCC A 0.16 10.3 GAC D 0.32 17.2 GGC G 0.14 9.2 GUA V 0.15 9.9 GCA A 0.27 17.5 GAA E 0.52 34.3 GGA G 0.37 24.2 GUG V 0.26 17.4 GCG A 0.14 9.0 GAG E 0.48 32.2 GGG G 0.16 10.2
Host Cells
[0063] The host cell can be a single cell (e.g., microalga, bacteria, yeast) or part of a multicellular organism such as a plant or fungus. Methods for expressing Fatb genes in a plant are described, e.g., in U.S. Pat. Nos. 5,850,022; 5,723,761; 5,639,790; 5,807,893; 5,455,167; 5,654,495; 5,512,482; 5,298,421; 5,667,997; 5,344,771; and 5,304,481, or can be obtained using other techniques generally known in plant biotechnology. Engineering of oleaginous microbes including those of Chlorophyta is disclosed in WO2010/063032, WO2011/150411, and WO2012/106560 and in the examples below.
[0064] Examples of oleaginous host cells include plant cells and microbial cells having a type II fatty acid biosynthetic pathway, including plastidic oleaginous cells such as those of oleaginous algae. Specific examples of microalgal cells include heterotrophic or obligate heterotrophic microalgae of the phylum Chlorophtya, the class Trebouxiophytae, the order Chlorellales, or the family Chlorellacae. Examples of oleaginous microalgae are provided in Published PCT Patent Applications WO2008/151149, WO2010/06032, WO2011/150410, and WO2011/150411, including species of Chlorella and Prototheca, a genus comprising obligate heterotrophs. The oleaginous cells can be, for example, capable of producing 20, 25, 30, 40, 50, 60, 70, 80, 85, or about 90% oil by cell weight, .+-.5%. Optionally, the oils produced can be low in DHA or EPA fatty acids. For example, the oils can comprise less than 5%, 2%, or 1% DHA and/or EPA. The above-mentioned publications also disclose methods for cultivating such cells and extracting oil, especially from microalgal cells; such methods are applicable to the cells disclosed herein and incorporated by reference for these teachings. When microalgal cells are used they can be cultivated autotrophically (unless an obligate heterotroph) or in the dark using a sugar (e.g., glucose, fructose and/or sucrose). In any of the embodiments described herein, the cells can be heterotrophic cells comprising an exogenous invertase gene so as to allow the cells to produce oil from a sucrose feedstock. Alternately, or in addition, the cells can metabolize xylose from cellulosic feedstocks. For example, the cells can be genetically engineered to express one or more xylose metabolism genes such as those encoding an active xylose transporter, a xylulose-5-phosphate transporter, a xylose isomerase, a xylulokinase, a xylitol dehydrogenase and a xylose reductase. See WO2012/154626, "GENETICALLY ENGINEERED MICROORGANISMS THAT METABOLIZE XYLOSE", published Nov. 15, 2012. The cells can be cultivated on a depolymerized cellulosic feedstock such as acid or enzyme hydrolyzed bagasse, sugar beet pulp, corn stover, wood chips, sawdust or switchgrass. Optionally, the cells can be cultivated on a depolymerized cellulosic feedstock comprising glucose and at least 5, 10, 20, 30 or 40% xylose, while producing at least 20% lipid by dry weight. Optionally, the lipid comprises triglycerides having a fatty acid profile characterized by at least 10, 15 or 20% C12:0
Oils and Related Products
[0065] The oleaginous cells express one or more exogenous genes encoding fatty acid biosynthesis enzymes. As a result, some embodiments feature cell oils that were not obtainable from a non-plant or non-seed oil, or not obtainable at all.
[0066] The oleaginous cells produce a storage oil, which is primarily triacylglyceride and may be stored in storage bodies of the cell. A raw oil may be obtained from the cells by disrupting the cells and isolating the oil. WO2008/151149, WO2010/06032, WO2011/150410, and WO2011/1504 disclose heterotrophic cultivation and oil isolation techniques. For example, oil may be obtained by cultivating, drying and pressing the cells. Methods for pressing cells are given in WO2010/120939. The oils produced may be refined, bleached and deodorized (RBD) as known in the art or as described in WO2010/120939. The raw or RBD oils may be used in a variety of food, chemical, and industrial products or processes. After recovery of the oil, a valuable residual biomass remains. Uses for the residual biomass include the production of paper, plastics, absorbents, adsorbents, as animal feed, for human nutrition, or for fertilizer.
[0067] Where a fatty acid profile of a triglyceride (also referred to as a "triacylglyceride" or "TAG") cell oil is given here, it will be understood that this refers to a nonfractionated sample of the storage oil extracted from the cell analyzed under conditions in which phospholipids have been removed or with an analysis method that is substantially insensitive to the fatty acids of the phospholipids (e.g. using chromatography and mass spectrometry). The oil may be subjected to an RBD process to remove phospholipids, free fatty acids and odors yet have only minor or negligible changes to the fatty acid profile of the triglycerides in the oil. Because the cells are oleaginous, in some cases the storage oil will constitute the bulk of all the TAGs in the cell.
[0068] The stable carbon isotope value .delta.13C is an expression of the ratio of 13C/12C relative to a standard (e.g. PDB, carbonite of fossil skeleton of Belemnite americana from Peedee formation of South Carolina). The stable carbon isotope value .delta.13C (0/00) of the oils can be related to the .delta.13C value of the feedstock used. In some embodiments, the oils are derived from oleaginous organisms heterotrophically grown on sugar derived from a C4 plant such as corn or sugarcane. In some embodiments the .delta.13C (0/00) of the oil is from -10 to -17 0/00 or from -13 to -16 0/00.
[0069] The oils produced according to the above methods in some cases are made using a microalgal host cell. As described above, the microalga can be, without limitation, fall in the classification of Chlorophyta, Trebouxiophyceae, Chlorellales, Chlorellaceae, or Chlorophyceae. It has been found that microalgae of Trebouxiophyceae can be distinguished from vegetable oils based on their sterol profiles. Oil produced by Chlorella protothecoides was found to produce sterols that appeared to be brassicasterol, ergosterol, campesterol, stigmasterol, and .beta.-sitosterol, when detected by GC-MS. However, it is believed that all sterols produced by Chlorella have C24.beta. stereochemistry. Thus, it is believed that the molecules detected as campesterol, stigmasterol, and .beta.-sitosterol, are actually 22,23-dihydrobrassicasterol, proferasterol and clionasterol, respectively. Thus, the oils produced by the microalgae described above can be distinguished from plant oils by the presence of sterols with C24.beta. stereochemistry and the absence of C24.alpha. stereochemistry in the sterols present. For example, the oils produced may contain 22, 23-dihydrobrassicasterol while lacking campesterol; contain clionasterol, while lacking in .beta.-sitosterol, and/or contain poriferasterol while lacking stigmasterol. Alternately, or in addition, the oils may contain significant amounts of .DELTA..sup.7-poriferasterol.
[0070] In one embodiment, the oils provided herein are not vegetable oils. Vegetable oils are oils extracted from plants and plant seeds. Vegetable oils can be distinguished from the non-plant oils provided herein on the basis of their oil content. A variety of methods for analyzing the oil content can be employed to determine the source of the oil or whether adulteration of an oil provided herein with an oil of a different (e.g. plant) origin has occurred. The determination can be made on the basis of one or a combination of the analytical methods. These tests include but are not limited to analysis of one or more of free fatty acids, fatty acid profile, total triacylglycerol content, diacylglycerol content, peroxide values, spectroscopic properties (e.g. UV absorption), sterol profile, sterol degradation products, antioxidants (e.g. tocopherols), pigments (e.g. chlorophyll), d13C values and sensory analysis (e.g. taste, odor, and mouth feel). Many such tests have been standardized for commercial oils such as the Codex Alimentarius standards for edible fats and oils.
[0071] Sterol profile analysis is a particularly well-known method for determining the biological source of organic matter. Campesterol, b-sitosterol, and stigmasterol are common plant sterols, with b-sitosterol being a principle plant sterol. For example, b-sitosterol was found to be in greatest abundance in an analysis of certain seed oils, approximately 64% in corn, 29% in rapeseed, 64% in sunflower, 74% in cottonseed, 26% in soybean, and 79% in olive oil (Gul et al. J. Cell and Molecular Biology 5:71-79, 2006).
[0072] Oil isolated from Prototheca moriformis strain UTEX1435 were separately clarified (CL), refined and bleached (RB), or refined, bleached and deodorized (RBD) and were tested for sterol content according to the procedure described in JAOCS vol. 60, no. 8, August 1983. Results of the analysis are shown below (units in mg/100 g):
TABLE-US-00006 TABLE 6 Refined, Refined & bleached, & Sterol Crude Clarified bleached deodorized 1 Ergosterol 384 398 293 302 (56%) (55%) (50%) (50%) 2 5,22-cholestadien- 14.6 18.8 14 15.2 24-methyl-3-ol (2.1%) (2.6%) (2.4%) (2.5%) (Brassicasterol) 3 24-methylcholest-5- 10.7 11.9 10.9 10.8 en-3-ol (Campesterol (1.6%) (1.6%) (1.8%) (1.8%) or 22,23- dihydrobrassicasterol) 4 5,22-cholestadien- 57.7 59.2 46.8 49.9 24-ethy1-3-ol (8.4%) (8.2%) (7.9%) (8.3%) (Stigmasterol or poriferasterol) 5 24-ethylcholest-5- 9.64 9.92 9.26 10.2 en-3-ol (.beta.-Sitosterol (1.4%) (1.4%) (1.6%) (1.7%) or clionasterol) 6 Other sterols 209 221 216 213 Total sterols 685.64 718.82 589.96 601.1
[0073] These results show three striking features. First, ergosterol was found to be the most abundant of all the sterols, accounting for about 50% or more of the total sterols. The amount of ergosterol is greater than that of campesterol, .beta.-sitosterol, and stigmasterol combined. Ergosterol is steroid commonly found in fungus and not commonly found in plants, and its presence particularly in significant amounts serves as a useful marker for non-plant oils. Secondly, the oil was found to contain brassicasterol. With the exception of rapeseed oil, brassicasterol is not commonly found in plant based oils. Thirdly, less than 2% .beta.-sitosterol was found to be present. .beta.-sitosterol is a prominent plant sterol not commonly found in microalgae, and its presence particularly in significant amounts serves as a useful marker for oils of plant origin. In summary, Prototheca moriformis strain UTEX1435 has been found to contain both significant amounts of ergosterol and only trace amounts of .beta.-sitosterol as a percentage of total sterol content. Accordingly, the ratio of ergosterol: .beta.-sitosterol or in combination with the presence of brassicasterol can be used to distinguish this oil from plant oils.
[0074] In some embodiments, the oil content of an oil provided herein contains, as a percentage of total sterols, less than 20%, 15%, 10%, 5%, 4%, 3%, 2%, or 1% .beta.-sitosterol. In other embodiments the oil is free from .beta.-sitosterol.
[0075] In some embodiments, the oil is free from one or more of .beta.-sitosterol, campesterol, or stigmasterol. In some embodiments the oil is free from .beta.-sitosterol, campesterol, and stigmasterol. In some embodiments the oil is free from campesterol. In some embodiments the oil is free from stigmasterol.
[0076] In some embodiments, the oil content of an oil provided herein comprises, as a percentage of total sterols, less than 20%, 15%, 10%, 5%, 4%, 3%, 2%, or 1% 24-ethylcholest-5-en-3-ol. In some embodiments, the 24-ethylcholest-5-en-3-ol is clionasterol. In some embodiments, the oil content of an oil provided herein comprises, as a percentage of total sterols, at least 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, or 10% clionasterol.
[0077] In some embodiments, the oil content of an oil provided herein contains, as a percentage of total sterols, less than 20%, 15%, 10%, 5%, 4%, 3%, 2%, or 1% 24-methylcholest-5-en-3-ol. In some embodiments, the 24-methylcholest-5-en-3-ol is 22, 23-dihydrobrassicasterol. In some embodiments, the oil content of an oil provided herein comprises, as a percentage of total sterols, at least 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, or 10% 22,23-dihydrobrassicasterol.
[0078] In some embodiments, the oil content of an oil provided herein contains, as a percentage of total sterols, less than 20%, 15%, 10%, 5%, 4%, 3%, 2%, or 1% 5,22-cholestadien-24-ethyl-3-ol. In some embodiments, the 5, 22-cholestadien-24-ethyl-3-ol is poriferasterol. In some embodiments, the oil content of an oil provided herein comprises, as a percentage of total sterols, at least 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, or 10% poriferasterol.
[0079] In some embodiments, the oil content of an oil provided herein contains ergosterol or brassicasterol or a combination of the two. In some embodiments, the oil content contains, as a percentage of total sterols, at least 5%, 10%, 20%, 25%, 35%, 40%, 45%, 50%, 55%, 60%, or 65% ergosterol. In some embodiments, the oil content contains, as a percentage of total sterols, at least 25% ergosterol. In some embodiments, the oil content contains, as a percentage of total sterols, at least 40% ergosterol. In some embodiments, the oil content contains, as a percentage of total sterols, at least 5%, 10%, 20%, 25%, 35%, 40%, 45%, 50%, 55%, 60%, or 65% of a combination of ergosterol and brassicasterol.
[0080] In some embodiments, the oil content contains, as a percentage of total sterols, at least 1%, 2%, 3%, 4% or 5% brassicasterol. In some embodiments, the oil content contains, as a percentage of total sterols less than 10%, 9%, 8%, 7%, 6%, or 5% brassicasterol.
[0081] In some embodiments the ratio of ergosterol to brassicasterol is at least 5:1, 10:1, 15:1, or 20:1.
[0082] In some embodiments, the oil content contains, as a percentage of total sterols, at least 5%, 10%, 20%, 25%, 35%, 40%, 45%, 50%, 55%, 60%, or 65% ergosterol and less than 20%, 15%, 10%, 5%, 4%, 3%, 2%, or 1% .beta.-sitosterol. In some embodiments, the oil content contains, as a percentage of total sterols, at least 25% ergosterol and less than 5% .beta.-sitosterol. In some embodiments, the oil content further comprises brassicasterol.
[0083] Sterols contain from 27 to 29 carbon atoms (C27 to C29) and are found in all eukaryotes. Animals exclusively make C27 sterols as they lack the ability to further modify the C27 sterols to produce C28 and C29 sterols. Plants however are able to synthesize C28 and C29 sterols, and C28/C29 plant sterols are often referred to as phytosterols. The sterol profile of a given plant is high in C29 sterols, and the primary sterols in plants are typically the C29 sterols b-sitosterol and stigmasterol. In contrast, the sterol profile of non-plant organisms contain greater percentages of C27 and C28 sterols. For example the sterols in fungi and in many microalgae are principally C28 sterols. The sterol profile and particularly the striking predominance of C29 sterols over C28 sterols in plants has been exploited for determining the proportion of plant and marine matter in soil samples (Huang, Wen-Yen, Meinschein W. G., "Sterols as ecological indicators"; Geochimica et Cosmochimia Acta. Vol 43. pp 739-745).
[0084] In some embodiments the primary sterols in the microalgal oils provided herein are sterols other than b-sitosterol and stigmasterol. In some embodiments of the microalgal oils, C29 sterols make up less than 50%, 40%, 30%, 20%, 10%, or 5% by weight of the total sterol content.
[0085] In some embodiments the microalgal oils provided herein contain C28 sterols in excess of C29 sterols. In some embodiments of the microalgal oils, C28 sterols make up greater than 50%, 60%, 70%, 80%, 90%, or 95% by weight of the total sterol content. In some embodiments the C28 sterol is ergosterol. In some embodiments the C28 sterol is brassicasterol.
[0086] In embodiments, oleaginous cells expressing one or more of the genes of Table 1 can produce an oil with at least 20, 40, 60 or 70% of C8, C10, C12, C14 or C16 fatty acids. In a specific embodiment, the level of myristate (C14:0) in the oil is greater than 30%.
[0087] Thus, in embodiments, there is a process for producing an oil, triglyceride, fatty acid, or derivative of any of these, comprising transforming a cell with any of the nucleic acids discussed herein. In another embodiment, the transformed cell is cultivated to produce an oil and, optionally, the oil is extracted. Oil extracted in this way can be used to produce food, oleochemicals or other products.
[0088] The oils discussed above alone or in combination are useful in the production of foods, fuels and chemicals (including plastics, foams, films, etc). The oils, triglycerides, fatty acids from the oils may be subjected to C--H activation, hydroamino methylation, methoxy-carbonation, ozonolysis, enzymatic transformations, epoxidation, methylation, dimerization, thiolation, metathesis, hydro-alkylation, lactonization, or other chemical processes.
[0089] After extracting the oil, a residual biomass may be left, which may have use as a fuel, as an animal feed, or as an ingredient in paper, plastic, or other product. For example, residual biomass from heterotrophic algae can be used in such products.
Deletion Mutants of FATB Genes that Enhance Production of Mid-Chain Fatty Acids in Host Cells
[0090] In another embodiment, there is a method for increasing the production of C12 or C10 fatty acids. The method comprises producing a polynucleotide having a sequence encoding a FATB acyl-ACP thioesterase but encoding a deletion mutation in the region corresponding to amino acids 66-98 of the SalFATB2 gene (SEQ ID NO: 8); i.e., a deletion in the FATB region corresponding to that characterized by SEQ ID NO: 42. In some cases, the region of the deletion mutant for the starting FATB already contains gaps; in this case, further residues in the region can be removed. For example, UcFATB2 has a 2-residue gap at positions 95-96 relative to SalFatB2, UcFatB1 has a 6-residue gap at positions 92-97 relative to SalFatB2, and LbeFatB1 has a 4-residue gap at positions 94-97 relative to SalFatB2. The full 32 amino acid deletion or shorter deletions (i.e., of 1, 2, 3, 4, 5, 6, 7, 8, 9,10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 or 31 amino acid residues) may also be effective in increasing C12 fatty acids in the FATB enzymes disclosed here or others known in the art (e.g., those with at least 80, 85, 90 or 95% identity to one of SEQ ID NOs: 1-18); this can readily be determined using the techniques disclosed here including the Examples.
[0091] Vectors containing genes that encode the deletion mutants can be expressed in an oleaginous host cell (single or multicellular) and compared to an untransformed cell to select mutants that increase the production of mid-chain fatty acids by the cell. This can be determined by extracting the oil and using has chromatography techniques.
[0092] Accordingly, in an embodiment, there is a method for increasing the production of C10-C14 fatty acids in a cell. The method comprises producing or providing an exogenous polynucleotide, the exogenous polynucleotide comprising an, optionally heterologous, control sequence fused to a coding region that encodes a plastid targeting sequence and a mutant FATB acyl-ACP thioesterase enzyme domain. The FATB acyl-ACP thioesterase enzyme domain has a deletion in the region corresponding to amino acids 66-98 of SEQ ID NO: 8. The exogenous polynucleotide is expressed in an oleaginous host cell. As a result of the expression, the host cell produces an oil that is enriched in C12 fatty acids, relative to a control cell lacking the exogenous polynucleotide. In specific embodiments, the sum of C10 and C12 fatty acids in the fatty acid profile of the oil is increased by at least 10, 20, 30, 50, 100, 150, or 200%. For example, the amount of C12 fatty acids in the oil is increased by at least 2-fold relative to the control cell. The starting FATB gene is not the CcFATB4 gene (SEQ ID NO: 46), because this gene already has a gap spanning the domain in which the deletion is made. In a related embodiment, the deletion leads to an increase in C8 and/or C10 fatty acids.
[0093] In an embodiment, there is a polynucleotide encoding a protein sequence having at least 75, 80, 85, 85.5, 86, 86.5, 87, 87.5, 88, 88.5, 89, 89.5, 90, 90.5, 91, 91.5, 92, 92.5, 93, 93.5, 94, 94.5, 95, 95.5, 96, 96.5, 97, 97.5, 98, 98.5 or 99% amino acid identity to any of SEQ ID NOs: 43-46, 50, 51, 54 or 55. The polynucleotide can comprise at least 60,65,70, 75, 80, 85, 85.5, 86, 86.5, 87, 87.5, 88, 88.5, 89, 89.5, 90, 90.5, 91, 91.5, 92, 92.5, 93, 93.5, 94, 94.5, 95, 95.5, 96, 96.5, 97, 97.5, 98, 98.5 or 99% sequence identity to any of SEQ ID NOs 47, 48, 52, or 56, or equivalent sequence by virtue of the degeneracy of the genetic code. The sequence has a deletion in the region corresponding to amino acids 66-98, and is not that of CcFatB4 (SEQ ID NO: 46). In related embodiments, there is a protein encoded by one of the above sequences, a vector for transforming a host cell, or a host cell expressing one of the sequences. There is also a method of producing an oil comprising expressing one of these sequences in an oleaginous host cell, cultivating the cell, and isolating an oil from the cell. The oil recovered can be elevated in C12 fatty acids 10, 20, 50, 100, 150, 200% or more relative to a control cell lacking the polynucleotide. Example 3 demonstrates the increase in C12:0 fatty acids resulting from expression of the deletion mutants in a Eukaryotic microalga, relative to controls lacking the deletion.
[0094] The polynucleotide sequence can be codon optimized for a variety of organisms including according to Tables 2-5.
TABLE-US-00007 TABLE 7 FatB Deletion mutant sequences Mature Amino amino acid Plant nucleic acid sequence acid sequence sequence (without codon-optimized GENE SEQ ID targeting for Prototheca (species, abbreviation) NO: peptide) moriformis Sassafras albidum 40 42 44 FATB1a (SalFATB1a) Lindera benzoin FATB1a 41 43 45 (LbeFATB1a) CpauFATB1.DELTA.28 (deletion 50 51 52 mutant of Cuphea paucipetala FATB1) ChFATB2.DELTA.27 (deletion 54 55 56 mutant of Cuphea hookeriana FATB1)
[0095] In accordance with an embodiment, a method of genetically engineering a cell includes expressing in the cell, a polynucleotide that encodes a protein having at least 65, 70, 80, 85, 86, 86, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to any of SEQ ID NOS: 40 to 43, 50, 51, 54 or 55; or has at least 65, 70, 80, 85, 86, 86, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% sequence identity to SEQ ID NO: 44, 45, 52 or 56, or equivalent sequence by virtue of the degeneracy of the genetic code. In a specific embodiment, a method of genetically engineering a cell includes expressing in the cell, a polynucleotide that encodes a protein having at least 86.7% sequence identity to 42, at least 80.7% sequence identity to 43, at least 88.2% sequence identity SEQ ID NOS: 51.
EXAMPLE 1
Discovery of Novel FATB Sequences
[0096] RNA was extracted from dried plant seeds and submitted for paired-end sequencing using the Illumina Hiseq 2000 platform. RNA sequence reads were assembled into corresponding seed transcriptomes using the Trinity software package and putative thioesterase-containing cDNA contigs were identified by mining transcriptomes for sequences with homology to known thioesterases. In some cases, these in silico identified putative thioesterase cDNAs were verified by direct reverse transcription PCR analysis using seed RNA and primer pairs targeting full-length thioesterase cDNAs. The resulting amplified products were cloned and sequenced de novo to confirm authenticity of identified thioesterase genes and to identify sequence variants arising from expression of different gene alleles or diversity of sequences within a population of seeds. For some sequences, a high-confidence, full-length transcript was assembled using Trinity and reverse transcription was not deemed to be necessary. The resulting amino acid sequences of all new putative FATB thioesterases were subjected to phylogenetic analyses using published full-length (Mayer and Shanklin, 2007) and truncated (THYME database) sequences as well as an extensive in-house phylogeny developed at Solazyme from FATB sequences identified in numerous oilseed transcriptomes. The in-house phylogeny comprising the acyl-ACP FATB thioesterases allows for prediction, in many cases, of the midchain specificity for each thioesterase; the FATBs predicted to be involved in biosynthesis of C8-C12 fatty acids were pursued.
[0097] The amino acid sequence and nucleic acid CDSs (native to the plant and codon optimized for Prototheca moriformis) of the novel FatB genes with and without their N-terminal plastid targeting peptides are shown in Table 1, above.
EXAMPLE 2
Expression of Transforming Vectors Expressing Acyl-ACP FATB Thioesterases
[0098] The nine acyl-ACP FATB thioesterase genes of Example 1 were synthesized in a codon-optimized form to reflect Prototheca moriformis (UTEX 1435) codon usage. A representative transforming construct and the sequence of the FATB enzymes is provided in SEQ ID NO: 38, using CcrasFATB1 as an example. The new thioesterases were synthesized with a modified transit peptide from Chlorella protothecoides (Cp) (SEQ ID NO: 40) in place of the native transit peptide. The modified transit peptide derived from the CpSAD1 gene, "CpSAD1tp_trimmed", was synthesized as an in-frame, N-terminal fusion to the FATB thioesterases in place of the native transit peptide.
[0099] Transgenic strains were generated via transformation of the base strain S7485 with a construct encoding 1 of the 12 FatB thioesterases. The construct pSZ5342/D4219 encoding CcrasFATB1 is shown as an example, but identical methods were used to generate each of the remaining 11 constructs encoding the different respective thioesterases. Construct pSZ5342 can be written as THI4A_5'::CrTUB2-ScSUC2-PmPGH:PmSAD2-2ver3-CpSAD1tp_trmd:CcrasFATB1-CvNR-- THI4A_3'. The relevant restriction sites in the construct from 5'-3', BspQ1, KpnI, BamHI, EcoRV, SpeI, XhoI, SacI, BspQ1, respectively, are indicated in lowercase, bold, and underlined. BspQ1 sites delimit the 5' and 3' ends of the transforming DNA. Bold, lowercase sequences at the 5' and 3' end of the construct represent genomic DNA from UTEX 1435 that target integration to the THI4A locus via homologous recombination. Proceeding in the 5' to 3' direction, the selection cassette has the C. reinhardtii .beta.-tubulin promoter driving expression of the S. cerevisiae gene SUC2 (conferring the ability to grow on sucrose) and the P.moriformis PGH gene 3' UTR. The promoter is indicated by lowercase, boxed text. The initiator ATG and terminator TGA for ScSUC2 are indicated by bold, uppercase italics, while the coding region is indicated with lowercase italics. The 3' UTR is indicated by lowercase underlined text. The spacer region between the two cassettes is indicated by upper case text. The second cassette containing the codon optimized CcrasFATB1 gene from Cuphea crassiflora fused to the heterologous C. protothecoides SAD1 plastid-targeting transit peptide, CpSAD1tp_trimmed, is driven by the P.moriformis SAD2-2ver3 pH5-responsive promoter and has the Chlorella vulgaris Nitrate Reductase (NR) gene 3' UTR. In this cassette, the PmSAD2-2ver3 promoter is indicated by lowercase, boxed text. The initiator ATG and terminator TGA for the CcrasFATB1 gene are indicated in bold, uppercase italics, while the coding region is indicated by lowercase italics. The 3' UTR is indicated by lowercase underlined text.
[0100] The sequence for all of the thioesterase constructs is identical with the exception of the encoded thioesterase. The full sequence for pSZ5342/D4219 integrating construct (SEQ ID NO: 38) is provided.
[0101] Constructs encoding heterologous FATB genes were transformed into a high-lipid-producing Prototheca strain and selected for the ability to grow on sucrose. Transformations, cell culture, lipid production and fatty acid analysis were all carried out as in WO2013/158938. Multiple transformations were performed. The fatty acid profiles of the strain with the highest C10 (for the first 6 genes listed), or C12 production (for the remaining genes) is reported in Table 8.
TABLE-US-00008 TABLE 8 Fatty acid profiles of top performing strain from each transformation (%; primary lipid) Species Gene Name C8:0 C10:0 C12:0 C14:0 C16:0 C18:0 C18:1 C18:2 C18:3.alpha. Cuphea crassiflora CcrasFATB1 0 4 1 3 35 3 47 5 0 Cuphea koehneana CkoeFATB3 0 9 2 3 32 3 45 5 0 Cuphea leptopoda CleptFATB1 0 6 1 3 34 4 46 5 0 Cuphea angustifolia CangFATB1 0 4 1 3 34 3 48 5 1 Cuphea llavea CllaFATB1 0 9 1 4 33 3 43 5 1 Cuphea lophostoma ClopFATB1 0 7 1 4 33 3 45 5 1 Sassafras albidum SalFATB1 0 0 7 3 32 4 47 5 1 Sassafras albidum SalFATB2 0 0 0 2 36 3 52 5 1 Lindera benzoin LbeFATB1 0 1 11 3 23 2 53 6 1 None (Parent strain) None 0 0 0 2 38 4 48 5 1
[0102] The six thioesterases from the Lythraceae cluster all display specificity towards C10:0 fatty acids: CcrasFATB1, which exhibits 4% C10:0 and 1% C12:0 fatty acid levels; CkoeFATB3, which exhibits 9% C10:0 and 2% C12:0 fatty acid levels; CleptFATB1, which exhibits 6% C10:0 and 1% C12:0 fatty acid levels; CangFATB1, which exhibits 4% C10:0 and 1% C12:0 fatty acid levels; CllaFATB1, which exhibits 9% C10:0 and 1% C12:0 fatty acid levels; and, ClopFATB1, which exhibits 7% C10:0 and 1% C12:0 fatty acid levels.
[0103] SalFATB1 and LbeFATB1, both of the Lauraceae family, exhibit substantial activity towards C12:0 fatty acids.
EXAMPLE 3
FATB Deletion Mutants of Lauraceae FATB Genes
[0104] Transforming vectors for deletion variants, of SalFATB1, and LbeFATB1, known respectively as SalFATB 1 a and LbeFATB1a, were synthesized, using the expression cassette and transit-peptide described in Example 2. The deletion variants had deletions in the region corresponding to amino acids 66-98 of the SalFATB2 gene (SEQ ID NO: 8). The constructs were codon-optimized to reflect UTEX 1435 codon usage. Transformations, cell culture, lipid production and fatty acid analysis were carried out as in Example 2. Constructs encoding heterologous FATB genes were transformed into a Prototheca moriformis strain and selected for the ability to grow on sucrose. The results for the two novel FATB thioesterases are displayed in Table 9.
TABLE-US-00009 TABLE 9 Fatty acid profiles of strains expressing deletion mutants of fatty acyl-ACP FATB genes (FATB1a) compared to wildtype genes lacking the deletion (FATB1). Gene C8:0 C10:0 C12:0 C14:0 C16:0 C18:0 C18:1 C18:2 C18:3 SalFATB1 0 0 7 3 32 4 47 5 1 SalFATB1a 0 0 15 3 27 3 45 5 1 LbeFATB1 0 1 11 3 23 2 53 6 1 LbeFATB1a 0 3 28 5 18 2 37 4 0
[0105] SalFATB1 and LbeFATB1, both of the Lauraceae family, exhibit substantial activity towards C12:0 fatty acids. SalFATB1a, which has a deletion of the 32 amino acids LFAVITTIFSVAEKQWTNLEWKPKPKPRLPQL (SEQ ID NO: 47), produced up to 15% C12:0 compared to 7% produced by the wild-type SalFATB1. The mean C12:0 level in SalFATB1a was 8.3% compared to 3.7% in SalFATB1, demonstrating a greater than 2-fold increase in activity upon deletion of the 32 amino acids. LbeFATB1a, which had a deletion of the 28 amino acids LLTVITTIFSAAEKQWTNLERKPKPPHL (SEQ ID NO: 48), produced up to 28% C12:0 compared to 11% produced by the wild-type LbeFATB1. The mean C12:0 level in LbeFATB1a is 17.2% compared to just 5.7% in LbeFATB1, demonstrating a greater than 3.0-fold increase in activity upon deletion of the 28 amino acids. The data suggest that deletion of those amino acids significantly improves (e.g., by 2-3 fold) the C12 activity of two other Lauraceae family thioesterases, SalFATB1 and LbeFATB1.
EXAMPLE 4
Additional Deletion Mutants from FATB Genes from Lythraceae
[0106] P. moriformis was transformed with additional deletion mutants of Lythraceae FATB genes above for Lauraceae FATB genes. Two deletion mutants were identified that showed elevated midchain (C8-14) fatty acid levels in cell-oil extracted from the microalga relative an equivalent transformation lacking the deletion. These are listed in Table 9, above in which they appear as CpauFATB1.DELTA.28 and ChFATB2.DELTA.27. Fatty acid profiles obtained in the P. moriformis model system are reported below in Table 10. ChFATB2.DELTA.27 demonstrated an increase in C8 and C10 fatty acids when compared to the wild-type, elaborating an average of 3.8% C8:0 and 11.5% C10:0 compared to 2.7% C8:0 and 8.0% C10:0, respectively. CpauFATB1.DELTA.28 demonstrates an increase in C10, C12 and C14 fatty acids when compared to the wild-type, elaborating an average of 7.6% C10:0 compared to 4.1% C10:0, respectively.
TABLE-US-00010 TABLE 10 Fatty acid profiles of cell-oil from P. moriformis transformed with Lythraceae FATB deletion mutants for top performing transformants (mean given in parenthesis). Mutant C8:0 C10:0 C12:0 C14:0 C16:0 C18:0 C18:1 C18:2 C18:3 CpauFATB1 0 (0.0) 9 (4.1) 1 3 31 2 45 6 1 (0.6) (2.7) CpauFATB1.DELTA.28 0 (0.0) 14 (7.6) 2 4 30 3 42 5 1 (1.1) (3.0) ChFATB2 7 (2.7) 16 (8.0) 0 2 21 3 44 5 1 (0.2) (2.0) ChFATB2.DELTA.27 9 (3.8) 20 (11.5) 0 1 17 2 45 5 0 (0.2) (1.8)
EXAMPLE 5
Modify Brassica napus Thioesterase (BnOTE) Enzyme Specificity by Site Directed Mutagenesis
[0107] In the example below, we demonstrate the ability of modifying the enzyme specificity of a FATA thioesterase originally isolated from Brassica napus (BnOTE, accession CAA52070), by site directed mutagenesis targeting two amino acids positions (D124 and D209).
[0108] To determine the impact of each amino acid substitution on the enzyme specificity of the BnOTE, the wild-type and the mutant BnOTE genes were cloned into a vector enabling expression within the lower palmitate P. moriformis strain S8588. The Saccharomyces carlsbergensis MEL1 gene (Accession no: AAA34770) was utilized as the selectable marker to introduce the wild-type and mutant BnOTE genes into FAD2-2 locus of P. moriformis strain S8588 by homologous recombination using previously described transformation methods (biolistics). The constructs that have been expressed in S8588 are listed in Table 11. S8588 is a recombinant P. moriformis strain having a FATA knockout and expressing an exogenous SUC2 gene and an exogenous P. moriformis KASII gene in the FATA locus. FATA knockouts that express sucrose invertase and/or KASII are described in co-owned applications WO2012/106560, WO2013/158938, WO2015/051319 and their respective priority applications thereof, all of which are herein incorporated by reference.
TABLE-US-00011 TABLE 11 DNA lot# and plasmid ID of DNA constructs that expressing wild-type and mutant BnOTE genes SEQ DNA Solazyme ID Lot# Plasmid NO: Construct D5309 pSZ6315 57 FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE-PmSAD2-1 utr::FAD2-2 D5310 pSZ6316 58 FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE(D124A)-PmSAD2-1 utr::FAD2-2 D5311 pSZ6317 59 FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE(D209A)-PmSAD2-1 utr::FAD2-2 D5312 pSZ6318 60 FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE(D124A, D209A)-PmSAD2-1 utr::FAD2-2
Construct pSZ6315: FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE-PmSAD2-1 utr::FAD2-2
[0109] The sequence of the pSZ6315 transforming DNA is provided in SEQ ID NO: 57. Relevant restriction sites in pSZ6315 are indicated in lowercase, bold and underlining and are 5'-3' SgrAI, Kpn I, SnaBI, AvrII, SpeI, AscI, ClaI, Sac I, SbfI, respectively. SgrAI and SbfI sites delimit the 5' and 3' ends of the transforming DNA. Bold, lowercase sequences represent FAD2-2 genomic DNA that permit targeted integration at FAD2-2 locus via homologous recombination. Proceeding in the 5' to 3' direction, the P. moriformis HXT1 promoter driving the expression of the Saccharomyces carlsbergensis MEL1 gene is indicated by boxed text. The initiator ATG and terminator TGA for MEL1 gene are indicated by uppercase, bold italics while the coding region is indicated in lowercase italics. The P. moriformis PGK 3' UTR is indicated by lowercase underlined text followed by the P. moriformis SAD2-2 V3 promoter, indicated by boxed italics text. The Initiator ATG and terminator TGA codons of the wild-type BnOTE are indicated by uppercase, bold italics, while the remainder of the coding region is indicated by bold italics. The three-nucleotide codon corresponding to the target two amino acids, D124 and D209, are double underlined. The P. moriformis SAD2-1 3'UTR is again indicated by lowercase underlined text followed by the FAD2-2 genomic region indicated by bold, lowercase text.
Construct pSZ6316: FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE (D124A)-PmSAD2-1 utr::FAD2-2
[0110] The sequence of the pSZ6316 transforming DNA is same as pSZ6315 except the D124A point mutation, the BnOTE D124A DNA sequence is provided in SEQ ID NO: 58.
Construct pSZ6317: FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE (D209A)-PmSAD2-1 utr::FAD2-2
[0111] The sequence of the pSZ6317 transforming DNA is same as pSZ6315 except the D209A point mutation, the BnOTE D209A DNA sequence is provided in SEQ ID NO: 59.
Construct pSZ6318: FAD2-2::PmHXT1-ScarMEL1-PmPGK:PmSAD2-2 V3-CpSADtp-BnOTE (D124A, D209A)-PmSAD2-1 utr::FAD2-2
[0112] The sequence of the pSZ6318 transforming DNA is same as pSZ6315 except two point mutations, D124A and D209A, the BnOTE (D124A, D209A) DNA sequence is provided in SEQ ID NO:60.
Results
[0113] The DNA constructs containing the wild-type and mutant BnOTE genes were transformed into the low palmitate parental strain S8588, primary transformants were clonally purified and grown under standard lipid production conditions at pH5.0. The resulting profiles from representative clones arising from transformations with pSZ6315, pSZ6316, pSZ6317, and pSZ6318 into S8588 are shown in Table 12. The parental strain S8588 produces 5.4% C18:0, when transformed with the DNA cassette expressing wild-type BnOTE, the transgenic lines produce approximately 11% C18:0. The BnOTE mutant (D124A) increased the amount of C18:0 by at least 2 fold compared to the wild-type protein. In contrast, the BnOTE D209A mutation appears to have no impact on the enzyme activity/specificity of the BnOTE thioesterase. Finally, expression of the BnOTE (D124A, D209A) resulted in very similar fatty acid profile to what we observed in the transformants from S8588 expressing BnOTE (S124A), again indicating that D209A has no significant impact on the enzyme activity.
TABLE-US-00012 TABLE 12 Fatty acid profiles in S8588 and derivative transgenic lines transformed with wild-type and mutant BnOTE genes Fatty Acid Area % Transforming DNA Sample ID C16:0 C18:0 C18:1 C18:2 pH5; S8588 (parental strain) 3.00 5.43 81.75 6.47 D5309, pSZ6315, pH5; S8588, D5309-6; 3.86 11.68 76.51 5.06 wild-type BnOTE pH5; S8588, D5309-2; 3.50 11.00 77.80 4.95 pH5; S8588, D5309-9; 3.51 10.72 78.03 5.00 pH5; S8588, D5309-10; 3.55 10.69 78.06 4.96 pH5; S8588, D5309-11; 3.61 10.69 78.05 4.95 D5310, pSZ6316, pH5; S8588, D5310-6; 4.27 31.55 55.31 5.30 BnOTE (D124A) pH5; S8588, D5310-1; 4.53 30.85 54.71 6.03 pH5; S8588, D5310-5; 5.21 20.75 65.43 5.02 pH5; S8588, D5310-10; 4.99 19.18 67.75 5.00 pH5; S8588, D5310-2; 4.90 18.92 68.17 4.98 D5311, pSZ6317, pH5; S8588, D5311-3; 3.50 11.90 76.95 4.98 BnOTE (D209A) pH5; S8588, D5311-4; 3.63 11.35 77.44 4.94 pH5; S8588, D5311-14; 3.47 11.23 77.68 4.98 pH5; S8588, D5311-10; 3.60 11.20 77.53 5.00 pH5; S8588, D5311-12; 3.53 11.12 77.59 5.09 D5312, pSZ6318, pH5; S8588, D5312-20; 4.79 37.97 47.74 6.01 BnOTE (D127A, pH5; S8588, D5312-40; 5.97 22.94 62.20 5.11 D212A) pH5; S8588, D5312-39; 6.07 22.75 62.24 5.17 pH5; S8588, D5312-16; 5.25 18.81 67.36 5.09 pH5; S8588, D5312-26; 4.93 18.70 68.37 4.96
[0114] It is understood that the examples and embodiments described herein are for illustrative purposes only and that various modifications or changes in light thereof will be suggested to persons skilled in the art and are to be included within the spirit and purview of this application and scope of the appended claims. All publications, patents, and patent applications cited herein are hereby incorporated by reference in their entirety for all purposes.
Informal Sequence Listing
TABLE-US-00013 [0115] Informal Sequence Listing Cuphea crassiflora FATB amino acid sequence (CcrasFATB1) SEQ ID NO: 1 MVAAAASSAFFPVPAPGTSTKPRKSGNWPSRLSPSSKPKSIPNGGFQVK ANASAHPKANGSAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLTA ITIVFVAAEKQWTMLDRKSKRPDMLVDSVGLKSIVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPTWGDTVEINTWFSQSGKIGMGSDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHFVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSKLESVTAMDPSEEDGVRSQYNHLLRLEDGTDVVKGR TEWRPKNAGTNGAISTGKTSNGNSVS Cuphea koehneana FATB amino acid sequence (CkoeFATB3) SEQ ID NO: 2 MVTAAASSAFFPVPAPGTSPKPGKSWPSSLSPSFKPKSIPNAGFQVKAN ASAHPKANGSAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLTAIT TVFVAAEKQWTMRDRKSKRPDMLVDSVGSKSIVLDGLVSRQIFSIRSYE IGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIWVL TKMQIMVNRYPTWGDTVEINTWFSHSGKIGMASDWLITDCNTGEILIRA TSVWAMMNQKTRRFSRLPYEVRQELTPHYVDSPHVIEDNDRKLHKFDVK TGDSIRKGLTPKWNDLDVNQHVNNVKYIGWILESMPIEVLETQELCSLT VEYRRECGMDSVLESVTAMDPSEDGGLSQYKHLLRLEDGTDIVKGRTEW RPKNAGTNGAISTAKPSNGNSVS Cuphea leptopoda FATB amino acid sequence (C1eptFATB1) SEQ ID NO: 3 MVGAAASSAFFPAPAPGTSPKPGKSGNWPSSLSPSLKPKSIPNGGFQVK ANASAHPKANGAAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLTA ITTVFVAAEKQWTMLDRKSKRPDMLVDSVGLKNIVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQILVNRYPAWGDTVEINTWFSQSGKIGMGSDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHFVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTARDPSEDGGRSQYNHLLRLEDGTDVVKGRT EWRSKNAGTNGATSTAKTSNGNSVS Cuphea angustifolia FATB amino acid sequence (CangFATB1) SEQ ID NO: 4 MVAAAASSAFFPVPAPGTSLKPGKSGNWPSSLSPSFKPKTIPSGGLQVK ANASAHPKANGSAVNLKSGSLDTQEDTSSSPPPRAFLNQLPDWSMLLTA ITTVFVAAEKQWTMLDRKSKRPEMLVDSVGLKSSVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPTWGDTVEVNTWFSQSGKIGMASDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHYVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTAMDPSEDGGVSQYKHLLRLEDGTDIVKGRT EWRPKNAGTNGATSKAKTSNGNSVS Cuphea llavea FATB1 amino acid sequence (CllaFATB1) SEQ ID NO: 5 MVAAAASSAFFPAPAPGSSPKPGKPGNWPSSLSPSFKPKSIPNGRFQVK ANASAHPKANGSAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLSA ITTVFVAAEKQWTMLDRKSKRPDMLVDSVGLKNIVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPAWGDTVEINTWFSQSGKIGMGSDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHFVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTAIDPSEDGGRSQYNHLLRLDDGTDVVKGRT EWRPKNAGTNGAISTGKTSNGNSVS Cuphea lophostoma FATB1 amino acid sequence (ClopFATB1) SEQ ID NO: 6 MVAAAASSAFFPVPAPGTSLKPWKSGNWPSSLSPSFKPKTIPSGGFQVK ANASAQPKANGSAVNLKSGSLNTQEDTTSSPPPRAFLNQLPDWSMLLTA ITTVFVAAEKQWTMLDRKSKRPEKLVDSVGLKSSVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPTWGDTVEINTWFSQSGKIGMASDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHYVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTAMDPSEDEGRSQYKHLLRLEDGTDIVKGRT EWRPKNAGTNGAISTAKNSNGNSVS Sassafras albidum FATB1 amino acid sequence (SalFATB1) SEQ ID NO: 7 MATTSLASAFCSMKAVMLARDGRGMKPRSSDLQLRAGNAQTPLKMINGT KFSYTESLKRLPDWSMLFAVITTIFSVAEKQWTNLEWKPKPKPRLPQLL DDHFGLHGLVERRTFAIRSYEVGPDRSTSIVAVMNHLQEATLNHAKSVG ILGDGFGTTLEMSKRDLAWVVRRTHVAVERYPAWGDTVEVECWIGASGN NGMRRDFLVRDCKTGEILTRCTSLSVMMNTRTRRLSKIPEEVRGEIGPL FIDNVAVKDEEIKKLQKLNDSSADYIQGGLTPRWNDLDVNQHVNNIKYV GWILETVPDSIFESHHISSITLEYRRECTRDSVLQSLTTVSGGSLEAGL VCDHLLQLEGGSEVLRARTEWRPKLTDSFRGIIVIPAEPSV Sassafras albidum FATB2 amino acid sequence (SalFATB2) SEQ ID NO: 8 MATTSLASAFCSMKAVMLARDGRGMKPRSSDLQLRAGNAQTPLKMINGT KESYTESLKRLPDWSMLFAVITTIFSVAEKQWTNLEWKPKPKPRLPQLL DDHFGLHGLVFRRTFAIRSYEVGPDRSTSIVAVMNHLQEATLNHAKSVG ILGDGFGTTLEMSKRDLAWVVRRTHVAVERYPAWGDTVEVEAWVGASGN IGMRRDFLVRDCKTGHILARCTSVSVMMNARTRRLSKIPQEVRAEIDPL FIEKVAVKEGEIKKLQKFNDSTADYIQGGWTPRWNDLDVNQHVNNIKYI GWIFKSVPDSISENHYLSSITLEYRRECTRGSALQSLTTVCGDSSEAGI ICEHLLQLEDGPEVLRARTEWRPKLTDSFRGIIVIPAEPSV Lindera benzoin FATB1 amino acid sequence (LbeFATB1) SEQ ID NO: 9 MVATSLASAFCSMKAVMLADDGRGMKPRSSDLQLRAGNAQTSLKMIDGT KFSYTESLKRLPDWSKLLTVITTIFSAAEKQWTNLERKPKPPHLLDDRF GLHGLVFRRTFAIRSYEVGPDRSASILAVLNHLQEATLNHAESVGILGD RFGETLEMSKRDLMWVVRRTYVAVERYPAWGDTVEIESWIGASGNNGMR REFLVRDFKTGEILTRCTSLSVMMNTRTRRLSKIPEEVRGEIGPVFIDN VAVKDEEIKKLQKLNDSTADYIQGGLIPRWNDLDLNQHVNNIKYVSWIL ETVPDSILESYHMSSITLEYRRECTRDSVLQSLTTVSGGSSEAGLVCEH SLLLEGGSEVLRARTEWRPKLTDSFRGISVIPAEQSV Cuphea crassiflora FATB amino acid sequence (CcrasFATB1), without targeting peptide SEQ ID NO: 10 MVAAAASSAFFPVPAPGTSTKPRKSGNWPSRLSPSSKPKSIPNGGFQVK ANASAHPKANGSAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLTA ITTVFVAAEKQWTMLDRKSKRPDMLVDSVGLKSIVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPTWGDTVEINTWFSQSGKIGMGSDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHFVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSKLESVTAMDPSEEDGVRSQYNHLLRLEDGTDVVKGR TEWRPKNAGTNGAISTGKTSNGNSVS Cuphea koehneana FATB amino acid sequence (CkoeFATB3), without targeting peptide SEQ ID NO: 11 MVTAAASSAFFPVPAPGTSPKPGKSWPSSLSPSFKPKSIPNAGFQVKAN ASAHPKANGSAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLTAIT TVFVAAEKQWTMRDRKSKRPDMLVDSVGSKSIVLDGLVSRQIFSIRSYE IGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIWVL TKMQIMVNRYPTWGDTVEINTWFSHSGKIGMASDWLITDCNTGEILIRA TSVWAMMNQKTRRFSRLPYEVRQELTPHYVDSPHVIEDNDRKLHKFDVK TGDSIRKGLTPKWNDLDVNQHVNNVKYIGWILESMPIEVLETQELCSLT VEYRRECGMDSVLESVTAMDPSEDGGLSQYKHLLRLEDGTDIVKGRTEW RPKNAGTNGAISTAKPSNGNSVS Cuphea leptopoda FATB amino acid sequence (CleptFATB1), without targeting peptide SEQ ID NO: 12 MVGAAASSAFFPAPAPGTSPKPGKSGNWPSSLSPSLKPKSIPNGGFQVK ANASAHPKANGAAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLTA ITTVFVAAEKQWTMLDRKSKRPDMLVDSVGLKNIVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQILVNRYPAWGDTVEINTWFSQSGKIGMGSDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHFVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTARDPSEDGGRSQYNHLLRLEDGTDVVKGRT EWRSKNAGTNGATSTAKTSNGNSVS Cuphea angustifolia FATB amino acid sequence (CangFATB1), without targeting peptide SEQ ID NO: 13
MVAAAASSAFFPVPAPGTSLKPGKSGNWPSSLSPSFKPKTIPSGGLQVK ANASAHPKANGSAVNLKSGSLDTQEDTSSSPPPRAFLNQLPDWSMLLTA ITTVFVAAEKQWTMLDRKSKRPEMLVDSVGLKSSVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPTWGDTVEVNTWFSQSGKIGMASDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHYVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTAMDPSEDGGVSQYKHLLRLEDGTDIVKGRT EWRPKNAGTNGATSKAKTSNGNSVS Cuphea llavea FATB1 amino acid sequence (CllaFATB1), without targeting peptide SEQ ID NO: 14 MVAAAASSAFFPAPAPGSSPKPGKPGNWPSSLSPSFKPKSIPNGRFQVK ANASAHPKANGSAVNLKSGSLNIQEDTSSSPPPRAFLNQLPDWSMLLSA ITTVFVAAEKQWTMLDRKSKRPDMLVDSVGLKNIVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPAWGDTVEINTWFSQSGKIGMGSDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHFVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTAIDPSEDGGRSQYNHLLRLDDGTDVVKGRT EWRPKNAGTNGAISTGKTSNGNSVS Cuphea lophostoma FATB1 amino acid sequence (ClopFATB1), without targeting peptide SEQ ID NO: 15 MVAAAASSAFFPVPAPGTSLKPWKSGNWPSSLSPSFKPKTIPSGGFQVK ANASAQPKANGSAVNLKSGSLNTQEDTTSSPPPRAFLNQLPDWSMLLTA ITTVFVAAEKQWTMLDRKSKRPEKLVDSVGLKSSVRDGLVSRQSFSIRS YEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIW VLTKMQIMVNRYPTWGDTVEINTWFSQSGKIGMASDWLISDCNTGEILI RATSVWAMMNQKTRRFSRLPYEVRQELTPHYVDSPHVIEDNDRKLHKFD VKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCS LTVEYRRECGMDSVLESVTAMDPSEDEGRSQYKHLLRLEDGTDIVKGRT EWRPKNAGTNGAISTAKNSNGNSVS Sassafras albidum FATB1 amino acid sequence (SalFATB1), without targeting peptide SEQ ID NO: 16 MATTSLASAFCSMKAVMLARDGRGMKPRSSDLQLRAGNAQTPLKMINGT KFSYTESLKRLPDWSMLFAVITTIFSVAEKQWTNLEWKPKPKPRLPQLL DDHFGLHGLVERRTFAIRSYEVGPDRSTSIVAVMNHLQEATLNHAKSVG ILGDGFGTTLEMSKRDLAWVVRRTHVAVERYPAWGDTVEVECWIGASGN NGMRRDFLVRDCKTGEILTRCTSLSVMMNTRTRRLSKIPEEVRGEIGPL FIDNVAVKDEEIKKLQKLNDSSADYIQGGLTPRWNDLDVNQHVNNIKYV GWILETVPDSIFESHHISSITLEYRRECTRDSVLQSLTTVSGGSLEAGL VCDHLLQLEGGSEVLRARTEWRPKLTDSFRGIIVIPAEPSV Sassafras albidum FATB2 amino acid sequence (SalFATB2), without targeting peptide SEQ ID NO: 17 MATTSLASAFCSMKAVMLARDGRGMKPRSSDLQLRAGNAQTPLKMINGT KFSYTESLKRLPDWSMLFAVITTIFSVAEKQWTNLEWKPKPKPRLPQLL DDHFGLHGLVFRRTFAIRSYEVGPDRSTSIVAVMNHLQEATLNHAKSVG ILGDGFGTTLEMSKRDLAWVVRRTHVAVERYPAWGDTVEVEAWVGASGN IGMRRDFLVRDCKTGHILARCTSVSVMMNARTRRLSKIPQEVRAEIDPL FIEKVAVKEGEIKKLQKFNDSTADYIQGGWTPRWNDLDVNQHVNNIKYI GWIFKSVPDSISENHYLSSITLEYRRECTRGSALQSLTTVCGDSSEAGI ICEHLLQLEDGPEVLRARTEWRPKLTDSFRGIIVIPAEPSV Lindera benzoin FATB1 amino acid sequence (LbeFATB1), without targeting peptide SEQ ID NO: 18 MVATSLASAFCSMKAVMLADDGRGMKPRSSDLQLRAGNAQTSLKMIDGI KESYTESLKRLPDWSKLLTVITTIFSAAEKQWTNLERKPKPPHLLDDRF GLHGLVFRRTFAIRSYEVGPDRSASILAVLNHLQEATLNHAESVGILGD RFGETLEMSKRDLMWVVRRTYVAVERYPAWGDTVEIESWIGASGNNGMR REFLVRDFKTGEILTRCTSLSVMMNTRTRRLSKIPEEVRGEIGPVFIDN VAVKDEEIKKLQKLNDSTADYIQGGLIPRWNDLDLNQHVNNIKYVSWIL ETVPDSILESYHMSSITLEYRRECTRDSVLQSLTTVSGGSSEAGLVCEH SLLLEGGSEVLRARTEWRPKLTDSFRGISVIPAEQSV Cuphea crassiflora FATB native CDS nucleic acid sequence (CcrasFATB1) SEQ ID NO: 19 ATGGTGGCTGCTGCAGCAAGTTCTGCATTCTTCCCTGTTCCTGCCCCAG GAACCTCCACTAAACCCAGGAAGTCCGGCAATTGGCCATCGAGATTGAG CCCTTCCTCCAAGCCCAAGTCAATCCCCAATGGCGGATTTCAGGTTAAG GCAAATGCCAGTGCCCATCCTAAGGCTAACGGTTCTGCAGTAAATCTAA AGTCTGGCAGCCTCAACACTCAGGAGGACACTTCGTCGTCCCCTCCTCC TCGGGCTTTCCTTAACCAGTTGCCTGATTGGAGTATGCTTCTGACTGCA ATCACGACCGTTTTCGTGGCGGCAGAGAAGCAGTGGACAATGCTTGATC GGAAATCTAAGAGGCCTGACATGCTCGTGGACTCGGTTGGGTTGAAGAG TATTGTTCGGGATGGGCTCGTGTCCAGACAAAGTTTTTCGATCAGGTCT TATGAAATAGGCGCTGATCGAACAGCCTCTATAGAGACGCTGATGAACC ACTTGCAGGAAACATCTATTAATCATTGTAAGAGTTTGGGCCTTCTCAA TGACGGCTTTGGTCGGACTCCTGGGATGTGTAAAAACGACCTCATTTGG GTGCTTACAAAAATGCAGATCATGGTGAATCGCTACCCAACTTGGGGCG ATACTGTTGAGATCAATACCTGGTTCTCCCAGTCGGGGAAAATCGGTAT GGGTAGCGATTGGCTAATAAGTGATTGCAATACAGGAGAAATTCTTATA AGGGCAACGAGCGTGTGGGCCATGATGAATCAAAAGACGAGAAGATTCT CAAGACTTCCATACGAGGTTCGCCAGGAGTTAACGCCTCATTTTGTGGA CTCTCCTCATGTCATTGAAGACAATGATCGGAAATTGCATAAGTTTGAT GTGAAGACTGGCGATTCTATTCGCAAGGGTCTAACTCCGAGGTGGAATG ATTTGGATGTCAATCAGCACGTAAGCAACGTGAAGTACATTGGGTGGAT TCTCGAGAGTATGCCAATAGAAGTTCTGGAGACCCAGGAGCTATGCTCT CTGACAGTTGAATATAGGCGGGAATGCGGAATGGACAGTAAGCTGGAGT CCGTGACTGCTATGGATCCCTCAGAAGAAGATGGAGTCCGGTCTCAGTA CAATCACCTTCTGCGGCTTGAGGATGGGACTGATGTCGTGAAGGGCAGA ACTGAGTGGCGACCGAAGAATGCAGGAACTAACGGGGCGATATCAACAG GAAAGACTTCAAATGGAAACTCGGTTTCTTAG Cuphea koehneana FATB FATB native CDS nucleic acid sequence (CkoeFATB3) SEQ ID NO: 20 ATGGTCACTGCTGCAGCAAGTTCTGCATTCTTCCCTGTTCCAGCCCCGG GAACCTCCCCTAAACCCGGGAAGTCCTGGCCATCGAGCTTGAGCCCTTC CTTCAAGCCCAAGTCAATCCCCAATGCCGGATTTCAGGTTAAGGCAAAT GCCAGTGCCCATCCTAAGGCTAACGGTTCTGCAGTAAATCTAAAGTCTG GCAGCCTCAACACTCAGGAGGACACTTCGTCGTCCCCTCCTCCTCGGGC TTTCCTTAACCAGTTGCCTGATTGGAGTATGCTTCTGACTGCAATCACG ACCGTCTTCGTGGCGGCAGAGAAGCAGTGGACTATGCGTGATCGGAAAT CTAAGAGGCCTGACATGCTCGTGGACTCGGTTGGATCGAAGAGTATTGT TCTGGATGGGCTCGTGTCCAGACAGATTTTTTCGATTAGATCTTATGAA ATAGGCGCTGATCGAACAGCCTCTATAGAGACGCTGATGAACCACTTGC AGGAAACATCTATCAATCATTGTAAGAGTTTGGGTCTTCTCAATGACGG CTTTGGTCGTACTCCTGGGATGTGTAAAAACGACCTCATTTGGGTGCTT ACAAAAATGCAGATCATGGTGAATCGCTACCCAACTTGGGGCGATACTG TTGAGATCAATACCTGGTTCTCCCATTCGGGGAAAATCGGTATGGCTAG CGATTGGCTAATAACTGATTGCAACACAGGAGAAATTCTTATAAGAGCA ACGAGCGTGTGGGCCATGATGAATCAAAAGACGAGAAGATTCTCAAGAC TTCCATACGAGGTTCGCCAGGAGTTAACGCCTCATTATGTGGACTCTCC TCATGTCATTGAAGATAATGATCGGAAATTGCATAAGTTTGATGTGAAG ACTGGTGATTCCATTCGTAAGGGTCTAACTCCGAAGTGGAATGACTTGG ATGTCAATCAGCACGTCAACAACGTGAAGTACATCGGGTGGATTCTCGA GAGTATGCCAATAGAAGTTTTGGAGACTCAGGAGCTATGCTCTCTCACC GTTGAATATAGGCGGGAATGCGGAATGGACAGTGTGCTGGAGTCCGTGA CTGCTATGGATCCCTCAGAAGATGGAGGCCTATCTCAGTACAAGCACCT TCTGCGGCTTGAGGATGGGACTGACATCGTGAAGGGCAGAACTGAGTGG CGACCGAAGAATGCAGGAACTAACGGGGCGATATCAACAGCAAAGCCTT CAAATGGAAACTCGGTCTCTTAG Cuphea leptopoda FATB native CDS nucleic acid sequence (CleptFATB1) SEQ ID NO: 21 ATGGTGGGTGCTGCAGCAAGTTCTGCATTCTTCCCTGCTCCAGCCCCGG GAACCTCCCCTAAACCCGGGAAGTCCGGCAATTGGCCATCAAGCTTGAG CCCTTCCTTAAAGCCCAAGTCAATCCCCAATGGCGGATTTCAGGTTAAG GCAAATGCCAGTGCCCATCCTAAGGCTAACGGTGCTGCAGTAAATCTAA AGTCTGGCAGCCTCAACACTCAGGAGGACACTTCGTCGTCCCCTCCTCC TCGGGCTTTCCTTAACCAGTTGCCTGATTGGAGTATGCTTCTGACTGCA ATCACGACCGTCTTCGTGGCGGCAGAGAAGCAGTGGACTATGCTTGATC GGAAATCTAAGAGGCCTGACATGCTCGTGGACTCGGTTGGGTTGAAGAA TATTGTTCGGGATGGGCTCGTGTCCAGACAGAGTTTTTCGATCAGGTCT TATGAAATAGGCGCTGATCGAACAGCCTCTATAGAGACGCTGATGAACC ACTTGCAGGAAACATCTATCAATCATTGTAAGAGTTTGGGTCTTCTCAA
TGACGGCTTTGGTCGTACTCCTGGGATGTGTAAAAACGACCTCATTTGG GTGCTTACAAAAATGCAGATCCTGGTGAATCGCTACCCAGCTTGGGGAG ATACTGTTGAGATCAATACCTGGTTCTCTCAGTCGGGGAAAATCGGCAT GGGTAGTGATTGGCTAATAAGTGATTGCAACACAGGAGAAATTCTTATA AGAGCAACGAGCGTGTGGGCAATGATGAATCAAAAGACGAGAAGATTCT CAAGACTTCCATACGAGGTTCGCCAGGAGTTAACGCCTCATTTTGTAGA CTCACCTCATGTCATTGAAGACAATGATCGGAAATTGCATAAGTTTGAT GTGAAGACTGGTGATTCTATTCGCAAGGGTCTAACTCCGAGGTGGAATG ACTTGGATGTCAATCAACACGTAAGCAACGTGAAGTACATTGGGTGGAT TCTCGAGAGTATGCCAATAGAAGTTTTGGAGACTCAGGAGCTATGCTCT CTCACCGTTGAATATAGGCGGGAATGCGGAATGGACAGTGTGCTGGAGT CCGTGACTGCTAGGGATCCCTCAGAAGATGGAGGCCGGTCTCAGTACAA TCACCTTCTGCGGCTTGAGGATGGGACTGATGTCGTGAAGGGCAGAACT GAGTGGCGATCGAAGAATGCAGGAACTAACGGGGCGACATCAACAGCAA AGACTTCAAATGGAAACTCGGTCTCTTAG Cuphea angustifolia FATB native CDS nucleic acid sequence (CangFATB1) SEQ ID NO: 22 ATGGTGGCTGCTGCAGCAAGTTCTGCATTCTTCCCTGTTCCAGCCCCGG GAACATCCCTTAAACCCGGGAAGTCCGGCAATTGGCCATCGAGCTTGAG CCCTTCCTTCAAGCCCAAGACAATCCCCAGTGGCGGACTTCAGGTTAAG GCAAATGCCAGTGCCCATCCTAAGGCTAACGGTTCTGCAGTAAATCTAA AGTCTGGCAGCCTCGACACTCAGGAGGACACTTCGTCGTCCCCTCCTCC TCGGGCTTTCCTTAACCAGTTGCCTGATTGGAGTATGCTTCTGACTGCA ATCACGACCGTCTTCGTGGCGGCAGAGAAGCAGTGGACTATGCTTGATA GGAAATCTAAGAGGCCTGAAATGCTCGTGGACTCGGTTGGGTTGAAGAG TAGTGTTCGGGATGGGCTCGTGTCCAGACAGAGTTTTTCGATTAGGTCT TATGAAATAGGCGCTGATCGAACAGCCTCTATAGAGACGCTGATGAACC ACTTGCAGGAAACATCTATCAATCATTGTAAGAGTTTGGGTCTTCTCAA CGATGGCTTTGGTCGTACTCCTGGGATGTGTAAAAACGACCTCATTTGG GTGCTTACAAAAATGCAGATCATGGTGAATCGCTACCCAACTTGGGGCG ATACTGTTGAGGTCAATACCTGGTTCTCCCAGTCGGGGAAAATCGGTAT GGCTAGCGATTGGCTAATCAGTGATTGCAACACAGGAGAAATTCTTATA AGAGCAACAAGCGTGTGGGCCATGATGAATCAAAAGACGAGAAGATTCT CAAGACTTCCATACGAGGTTCGCCAGGAGCTAACACCTCATTATGTGGA CTCTCCTCATGTCATTGAAGATAATGATCGGAAATTGCATAAGTTTGAT GTGAAGACTGGTGATTCCATTCGCAAGGGTCTAACTCCGAGGTGGAATG ACTTGGATGTCAATCAGCACGTAAGCAACGTGAAGTACATTGGGTGGAT TCTTGAGAGTATGCCAATAGAAGTTTTGGAGACCCAGGAGCTATGCTCT CTCACCGTTGAATATAGGCGGGAATGCGGAATGGACAGTGTGCTGGAGT CCGTGACTGCTATGGATCCCTCAGAAGATGGAGGCGTGTCTCAGTACAA GCACCTTCTGCGGCTTGAGGATGGGACTGATATCGTGAAGGGCAGAACT GAATGGCGACCGAAGAATGCAGGAACTAATGGGGCGACATCAAAAGCAA AGACTTCAAATGGAAACTCGGTCTCTTAG Cuphea llavea FATB1 native CDS nucleic acid sequence (CllaFATB1) SEQ ID NO: 23 ATGGTGGCTGCTGCAGCAAGTTCTGCATTCTTCCCTGCTCCAGCCCCGG GATCCTCACCTAAACCCGGGAAGCCCGGTAATTGGCCATCGAGCTTGAG CCCTTCCTTCAAGCCCAAGTCAATCCCCAATGGCCGATTTCAGGTTAAG GCAAATGCGAGTGCCCATCCTAAGGCTAACGGTTCTGCAGTAAATCTAA AGTCTGGCAGCCTCAACACTCAGGAGGACACTTCGTCGTCCCCTCCTCC TCGGGCTTTCCTTAACCAGTTGCCTGATTGGAGTATGCTTCTGTCTGCA ATCACGACTGTATTCGTGGCGGCAGAGAAGCAGTGGACTATGCTTGATC GGAAATCTAAGAGGCCTGACATGCTTGTGGACTCGGTTGGGTTGAAGAA TATTGTTCGGGATGGGCTCGTGTCCAGACAGAGTTTTTCGATTAGATCT TATGAAATAGGCGCTGATCGAACAGCTTCTATAGAGACACTGATGAACC ACTTGCAGGAAACATCTATCAATCATTGTAAGAGTTTGGGTCTTCTCAA TGACGGCTTTGGTCGTACTCCTGGGATGTGTAAAAACGACCTCATTTGG GTGCTTACAAAAATGCAGATCATGGTGAATCGCTACCCAGCTTGGGGCG ATACTGTTGAGATCAATACATGGTTCTCCCAGTCGGGGAAAATCGGTAT GGGTAGCGATTGGCTAATAAGTGATTGCAACACAGGAGAAATTCTTATA AGAGCAACGAGCGTGTGGGCCATGATGAATCAAAAGACGAGAAGATTCT CAAGACTTCCATATGAGGTTCGCCAGGAGTTAACGCCTCATTTTGTGGA CTCTCCTCATGTCATTGAAGACAATGATCGGAAATTGCATAAGTTCGAT GTGAAGACTGGTGATTCTATTCGCAAGGGTCTAACTCCGAGGTGGAATG ACTTGGATGTCAATCAACACGTAAGCAACGTGAAGTACATTGGGTGGAT TCTCGAGAGTATGCCAATAGAAGTTTTGGAGACCCAGGAACTATGCTCT CTCACAGTTGAATATAGGCGGGAATGCGGAATGGACAGTGTGCTGGAGT CCGTGACTGCTATAGATCCCTCAGAAGATGGAGGGCGGTCTCAGTACAA TCACCTTCTGCGGCTTGATGATGGGACTGATGTCGTGAAGGGCAGAACA GAGTGGCGACCGAAGAATGCAGGAACTAACGGGGCGATATCAACAGGAA AGACTTCAAATGGGAACTCGGTCTCCTAG Cuphea lophostoma FATB1 native CDS nucleic acid sequence (ClopFATB1) SEQ ID NO: 24 ATGGTGGCTGCTGCAGCAAGTTCTGCATTCTTCCCTGTTCCAGCCCCGG GAACCTCCCTTAAACCCTGGAAGTCCGGAAATTGGCCATCGAGCTTGAG CCCTTCCTTCAAGCCCAAGACAATCCCCAGTGGCGGATTTCAGGTTAAG GCAAATGCCAGTGCCCAGCCTAAGGCTAACGGTTCTGCAGTAAATCTAA AGTCTGGCAGCCTCAACACTCAGGAGGACACAACGTCGTCGCCTCCTCC TCGGGCTTTCCTTAACCAGTTGCCTGATTGGAGTATGCTTCTGACTGCA ATCACGACCGTCTTCGTGGCGGCGGAGAAGCAGTGGACAATGCTTGATA GGAAATCTAAGAGGCCTGAAAAGCTCGTGGACTCGGTTGGGTTGAAGAG TAGTGTTCGGGATGGGCTCGTGTCCAGACAGAGTTTTTCGATTAGGTCT TATGAAATAGGCGCTGATCGAACAGCCTCTATAGAGACGTTGATGAACC ACTTGCAGGAAACATCTATCAATCATTGTAAGAGTTTGGGTCTTCTCAA CGACGGCTTTGGTCGTACTCCTGGGATGTGTAAAAACGACCTCATTTGG GTGCTTACGAAAATGCAGATCATGGTGAATCGCTACCCAACTTGGGGCG ATACTGTTGAGATCAATACCTGGTTCTCCCAGTCGGGGAAAATCGGTAT GGCTAGCGATTGGCTAATAAGTGATTGCAACACAGGAGAAATTCTTATA AGAGCAACGAGCGTGTGGGCCATGATGAATCAAAAGACGAGAAGGTTCT CAAGACTTCCATACGAGGTTCGCCAGGAGTTAACGCCTCATTATGTGGA CTCTCCTCATGTCATTGAAGACAATGATCGGAAATTGCATAAGTTTGAT GTGAAGACTGGTGATTCCATTCGCAAGGGTCTGACTCCGAGGTGGAATG ACTTGGATGTCAATCAGCACGTAAGCAACGTGAAGTACATTGGGTGGAT TCTGGAGAGTATGCCAATAGAAGTTTTGGAGACCCAGGAGCTATGCTCT CTCACCGTTGAATATAGGCGGGAATGCGGGATGGACAGTGTGCTGGAGT CCGTGACTGCTATGGATCCCTCAGAAGATGAAGGCCGGTCTCAGTACAA GCACCTTCTGCGGCTTGAGGATGGGACTGATATCGTGAAGGGCAGAACT GAGTGGCGACCGAAGAATGCAGGAACTAACGGGGCGATATCAACAGCAA AGAATTCAAATGGAAACTCGGTCTCTTAG Sassafras albidum FATB1 native CDS nucleic acid sequence (SalFATB1) SEQ ID NO: 25 ATGGCCACCACCTCTTTAGCTTCTGCTTTCTGCTCGATGAAAGCTGTAA TGTTGGCTCGTGATGGCAGGGGCATGAAACCCAGGAGCAGTGATTTGCA GCTGAGGGCGGGAAATGCACAAACCCCTTTGAAGATGATCAATGGGACC AAGTTCAGTTACACGGAGAGCTTGAAAAGGTTGCCTGACTGGAGCATGC TCTTTGCAGTGATCACAACCATCTTTTCGGTTGCTGAGAAGCAGTGGAC CAATCTAGAGTGGAAGCCGAAGCCGAAGCCGAGGCTACCCCAGTTGCTT GATGACCATTTTGGACTGCATGGGTTAGTTTTCAGGCGCACCTTTGCCA TCAGATCTTATGAGGTCGGACCTGACCGCTCCACATCTATAGTGGCTGT TATGAATCACTTGCAGGAGGCTACACTTAATCATGCGAAGAGTGTGGGA ATTCTAGGAGATGGATTCGGTACGACGCTAGAGATGAGTAAGAGAGATC TGGCGTGGGTTGTGAGACGCACGCATGTTGCTGTGGAACGGTACCCTGC TTGGGGTGATACTGTTGAAGTAGAGTGCTGGATTGGTGCATCTGGAAAT AATGGCATGCGCCGTGATTTCCTTGTCCGGGACTGCAAAACAGGCGAAA TTCTTACAAGATGTACCAGTCTTTCGGTGATGATGAATACAAGGACAAG GAGGTTGTCCAAAATCCCTGAAGAAGTTAGAGGGGAGATAGGGCCTCTA TTCATTGATAATGTGGCTGTCAAGGACGAGGAAATTAAGAAACTACAGA AGCTCAATGACAGCTCTGCAGATTACATCCAAGGAGGTTTGACTCCTCG ATGGAATGATTTGGATGTCAATCAGCATGTTAACAACATCAAATACGTT GGCTGGATTCTTGAGACTGTCCCAGACTCCATCTTTGAGAGTCATCATA TTTCCAGCATCACTCTTGAATACAGGAGAGAGTGCACCAGGGATAGCGT GCTGCAGTCCCTGACCACTGTCTCCGGTGGCTCGTTGGAGGCTGGGTTA GTGTGCGATCACTTGCTCCAGCTTGAAGGTGGGTCTGAGGTATTGAGGG CAAGAACAGAGTGGAGGCCTAAGCTTACCGATAGTTTCAGAGGGATTAT TGTGATACCCGCAGAACCGAGTGTGTAA Sassafras albidum FATB2 native CDS nucleic acid sequence (SalFATB2) SEQ ID NO: 26 ATGGCCACCACCTCTTTAGCTTCTGCTTTCTGCTCGATGAAAGCTGTAA
TGTTGGCTCGTGATGGCAGGGGCATGAAACCCAGGAGCAGTGATTTGCA GCTGAGGGCGGGAAATGCACAAACCCCTTTGAAGATGATCAATGGGACC AAGTTCAGTTACACGGAGAGCTTGAAAAGGTTGCCTGACTGGAGCATGC TCTTTGCAGTGATCACAACCATCTTTTCGGTTGCTGAGAAGCAGTGGAC CAATCTAGAGTGGAAGCCGAAGCCGAAGCCGAGGCTACCCCAGTTGCTT GATGACCATTTTGGACTGCATGGGTTAGTTTTCAGGCGCACCTTTGCCA TCAGATCTTATGAGGTCGGACCTGACCGCTCCACATCTATAGTGGCTGT TATGAATCACTTGCAGGAGGCTACACTTAATCATGCGAAGAGTGTGGGA ATTCTAGGAGATGGATTCGGTACGACGCTAGAGATGAGTAAGAGAGATC TGGCGTGGGTTGTGAGACGCACGCATGTTGCTGTGGAACGGTACCCCGC TTGGGGCGATACTGTTGAAGTCGAGGCCTGGGTCGGTGCATCTGGAAAC ATTGGCATGCGCCGCGATTTTCTTGTCCGCGACTGCAAAACTGGCCACA TTCTTGCAAGATGTACCAGTGTTTCAGTGATGATGAATGCGAGGACACG GAGATTGTCCAAAATTCCCCAAGAAGTTAGAGCCGAGATTGACCCTCTT TTCATTGAAAAGGTTGCGGTCAAGGAAGGGGAAATTAAGAAATTACAGA AGTTCAATGATAGCACTGCAGATTACATTCAAGGGGGTTGGACTCCTCG ATGGAATGATTTGGATGTCAATCAGCACGTGAACAATATCAAATACATT GGCTGGATTTTTAAGAGCGTCCCAGACTCTATCTCTGAGAATCATTATC TTTCTAGCATCACTCTCGAATACAGGAGAGAGTGCACAAGGGGCAGCGC GCTGCAGTCCCTGACCACTGTTTGTGGTGACTCGTCGGAAGCTGGGATC ATATGTGAGCACCTACTCCAGCTTGAGGATGGGCCTGAGGTTTTGAGGG CAAGAACAGAGTGGAGGCCTAAGCTTACCGATAGTTTCAGAGGGATTAT TGTGATACCCGCAGAACCGAGTGTGTAA Lindera benzoin FATB1 native CDS nucleic acid sequence (LbeFATB1) SEQ ID NO: 27 ATGGTCACTGCTGCAGCAAGTTCTGCATTCTTCCCTGTTCCAGCCCCGG GAACCTCCCCTAAACCCGGGAAGTCCTGGCCATCGAGCTTGAGCCCTTC CTTCAAGCCCAAGTCAATCCCCAATGCCGGATTTCAGGTTAAGGCAAAT GCCAGTGCCCATCCTAAGGCTAACGGTTCTGCAGTAAATCTAAAGTCTG GCAGCCTCAACACTCAGGAGGACACTTCGTCGTCCCCTCCTCCTCGGGC TTTCCTTAACCAGTTGCCTGATTGGAGTATGCTTCTGACTGCAATCACG ACCGTCTTCGTGGCGGCAGAGAAGCAGTGGACTATGCGTGATCGGAAAT CTAAGAGGCCTGACATGCTCGTGGACTCGGTTGGATCGAAGAGTATTGT TCTGGATGGGCTCGTGTCCAGACAGATTTTTTCGATTAGATCTTATGAA ATAGGCGCTGATCGAACAGCCTCTATAGAGACGCTGATGAACCACTTGC AGGAAACATCTATCAATCATTGTAAGAGTTTGGGTCTTCTCAATGACGG CTTTGGTCGTACTCCTGGGATGTGTAAAAACGACCTCATTTGGGTGCTT ACAAAAATGCAGATCATGGTGAATCGCTACCCAACTTGGGGCGATACTG TTGAGATCAATACCTGGTTCTCCCATTCGGGGAAAATCGGTATGGCTAG CGATTGGCTAATAACTGATTGCAACACAGGAGAAATTCTTATAAGAGCA ACGAGCGTGTGGGCCATGATGAATCAAAAGACGAGAAGATTCTCAAGAC TTCCATACGAGGTTCGCCAGGAGTTAACGCCTCATTATGTGGACTCTCC TCATGTCATTGAAGATAATGATCGGAAATTGCATAAGTTTGATGTGAAG ACTGGTGATTCCATTCGTAAGGGTCTAACTCCGAAGTGGAATGACTTGG ATGTCAATCAGCACGTCAACAACGTGAAGTACATCGGGTGGATTCTCGA GAGTATGCCAATAGAAGTTTTGGAGACTCAGGAGCTATGCTCTCTCACC GTTGAATATAGGCGGGAATGCGGAATGGACAGTGTGCTGGAGTCCGTGA CTGCTATGGATCCCTCAGAAGATGGAGGCCTATCTCAGTACAAGCACCT TCTGCGGCTTGAGGATGGGACTGACATCGTGAAGGGCAGAACTGAGTGG CGACCGAAGAATGCAGGAACTAACGGGGCGATATCAACAGCAAAGCCTT CAAATGGAAACTCGGTCTCTTAG Cuphea crassiflora FATB native CDS codon optimized acid sequence (CcrasFATB1) SEQ ID NO: 28 ATGGTGGCCGCCGCCGCCTCCTCCGCCTTCTTCCCCGTGCCCGCCCCCG GCACCTCCACCAAGCCCCGCAAGTCCGGCAACTGGCCCTCCCGCCTGTC CCCCTCCTCCAAGCCCAAGTCCATCCCCAACGGCGGCTTCCAGGTGAAG GCCAACGCCTCCGCCCACCCCAAGGCCAACGGCTCCGCCGTGAACCTGA AGTCCGGCTCCCTGAACACCCAGGAGGACACCTCCTCCTCCCCCCCCCC CCGCGCCTTCCTGAACCAGCTGCCCGACTGGTCCATGCTGCTGACCGCC ATCACCACCGTGTTCGTGGCCGCCGAGAAGCAGTGGACCATGCTGGACC GCAAGTCCAAGCGCCCCGACATGCTGGTGGACTCCGTGGGCCTGAAGTC CATCGTGCGCGACGGCCTGGTGTCCCGCCAGTCCTTCTCCATCCGCTCC TACGAGATCGGCGCCGACCGCACCGCCTCCATCGAGACCCTGATGAACC ACCTGCAGGAGACCTCCATCAACCACTGCAAGTCCCTGGGCCTGCTGAA CGACGGCTTCGGCCGCACCCCCGGCATGTGCAAGAACGACCTGATCTGG GTGCTGACCAAGATGCAGATCATGGTGAACCGCTACCCCACCTGGGGCG ACACCGTGGAGATCAACACCTGGTTCTCCCAGTCCGGCAAGATCGGCAT GGGCTCCGACTGGCTGATCTCCGACTGCAACACCGGCGAGATCCTGATC CGCGCCACCTCCGTGTGGGCCATGATGAACCAGAAGACCCGCCGCTTCT CCCGCCTGCCCTACGAGGTGCGCCAGGAGCTGACCCCCCACTTCGTGGA CTCCCCCCACGTGATCGAGGACAACGACCGCAAGCTGCACAAGTTCGAC GTGAAGACCGGCGACTCCATCCGCAAGGGCCTGACCCCCCGCTGGAACG ACCTGGACGTGAACCAGCACGTGTCCAACGTGAAGTACATCGGCTGGAT CCTGGAGTCCATGCCCATCGAGGTGCTGGAGACCCAGGAGCTGTGCTCC CTGACCGTGGAGTACCGCCGCGAGTGCGGCATGGACTCCAAGCTGGAGT CCGTGACCGCCATGGACCCCTCCGAGGAGGACGGCGTGCGCTCCCAGTA CAACCACCTGCTGCGCCTGGAGGACGGCACCGACGTGGTGAAGGGCCGC ACCGAGTGGCGCCCCAAGAACGCCGGCACCAACGGCGCCATCTCCACCG GCAAGACCTCCAACGGCAACTCCGTGTCCTGA Cuphea koehneana FATB FATB codon optimized CDS nucleic acid sequence (CkoeFATB3) SEQ ID NO: 29 ATGGTGACCGCCGCCGCCTCCTCCGCCTTCTTCCCCGTGCCCGCCCCCG GCACCTCCCCCAAGCCCGGCAAGTCCTGGCCCTCCTCCCTGTCCCCCTC CTTCAAGCCCAAGTCCATCCCCAACGCCGGCTTCCAGGTGAAGGCCAAC GCCTCCGCCCACCCCAAGGCCAACGGCTCCGCCGTGAACCTGAAGTCCG GCTCCCTGAACACCCAGGAGGACACCTCCTCCTCCCCCCCCCCCCGCGC CTTCCTGAACCAGCTGCCCGACTGGTCCATGCTGCTGACCGCCATCACC ACCGTGTTCGTGGCCGCCGAGAAGCAGTGGACCATGCGCGACCGCAAGT CCAAGCGCCCCGACATGCTGGTGGACTCCGTGGGCTCCAAGTCCATCGT GCTGGACGGCCTGGTGTCCCGCCAGATCTTCTCCATCCGCTCCTACGAG ATCGGCGCCGACCGCACCGCCTCCATCGAGACCCTGATGAACCACCTGC AGGAGACCTCCATCAACCACTGCAAGTCCCTGGGCCTGCTGAACGACGG CTTCGGCCGCACCCCCGGCATGTGCAAGAACGACCTGATCTGGGTGCTG ACCAAGATGCAGATCATGGTGAACCGCTACCCCACCTGGGGCGACACCG TGGAGATCAACACCTGGTTCTCCCACTCCGGCAAGATCGGCATGGCCTC CGACTGGCTGATCACCGACTGCAACACCGGCGAGATCCTGATCCGCGCC ACCTCCGTGTGGGCCATGATGAACCAGAAGACCCGCCGCTTCTCCCGCC TGCCCTACGAGGTGCGCCAGGAGCTGACCCCCCACTACGTGGACTCCCC CCACGTGATCGAGGACAACGACCGCAAGCTGCACAAGTTCGACGTGAAG ACCGGCGACTCCATCCGCAAGGGCCTGACCCCCAAGTGGAACGACCTGG ACGTGAACCAGCACGTGAACAACGTGAAGTACATCGGCTGGATCCTGGA GTCCATGCCCATCGAGGTGCTGGAGACCCAGGAGCTGTGCTCCCTGACC GTGGAGTACCGCCGCGAGTGCGGCATGGACTCCGTGCTGGAGTCCGTGA CCGCCATGGACCCCTCCGAGGACGGCGGCCTGTCCCAGTACAAGCACCT GCTGCGCCTGGAGGACGGCACCGACATCGTGAAGGGCCGCACCGAGTGG CGCCCCAAGAACGCCGGCACCAACGGCGCCATCTCCACCGCCAAGCCCT CCAACGGCAACTCCGTGTCCTGA Cuphea leptopoda FATB codon optimized CDS nucleic acid sequence (CleptFATB1) SEQ ID NO: 30 ATGGTGGGCGCCGCCGCCTCCTCCGCCTTCTTCCCCGCCCCCGCCCCCG GCACCTCCCCCAAGCCCGGCAAGTCCGGCAACTGGCCCTCCTCCCTGTC CCCCTCCCTGAAGCCCAAGTCCATCCCCAACGGCGGCTTCCAGGTGAAG GCCAACGCCTCCGCCCACCCCAAGGCCAACGGCGCCGCCGTGAACCTGA AGTCCGGCTCCCTGAACACCCAGGAGGACACCTCCTCCTCCCCCCCCCC CCGCGCCTTCCTGAACCAGCTGCCCGACTGGTCCATGCTGCTGACCGCC ATCACCACCGTGTTCGTGGCCGCCGAGAAGCAGTGGACCATGCTGGACC GCAAGTCCAAGCGCCCCGACATGCTGGTGGACTCCGTGGGCCTGAAGAA CATCGTGCGCGACGGCCTGGTGTCCCGCCAGTCCTTCTCCATCCGCTCC TACGAGATCGGCGCCGACCGCACCGCCTCCATCGAGACCCTGATGAACC ACCTGCAGGAGACCTCCATCAACCACTGCAAGTCCCTGGGCCTGCTGAA CGACGGCTTCGGCCGCACCCCCGGCATGTGCAAGAACGACCTGATCTGG GTGCTGACCAAGATGCAGATCCTGGTGAACCGCTACCCCGCCTGGGGCG ACACCGTGGAGATCAACACCTGGTTCTCCCAGTCCGGCAAGATCGGCAT GGGCTCCGACTGGCTGATCTCCGACTGCAACACCGGCGAGATCCTGATC CGCGCCACCTCCGTGTGGGCCATGATGAACCAGAAGACCCGCCGCTTCT CCCGCCTGCCCTACGAGGTGCGCCAGGAGCTGACCCCCCACTTCGTGGA CTCCCCCCACGTGATCGAGGACAACGACCGCAAGCTGCACAAGTTCGAC
GTGAAGACCGGCGACTCCATCCGCAAGGGCCTGACCCCCCGCTGGAACG ACCTGGACGTGAACCAGCACGTGTCCAACGTGAAGTACATCGGCTGGAT CCTGGAGTCCATGCCCATCGAGGTGCTGGAGACCCAGGAGCTGTGCTCC CTGACCGTGGAGTACCGCCGCGAGTGCGGCATGGACTCCGTGCTGGAGT CCGTGACCGCCCGCGACCCCTCCGAGGACGGCGGCCGCTCCCAGTACAA CCACCTGCTGCGCCTGGAGGACGGCACCGACGTGGTGAAGGGCCGCACC GAGTGGCGCTCCAAGAACGCCGGCACCAACGGCGCCACCTCCACCGCCA AGACCTCCAACGGCAACTCCGTGTCCTGA Cuphea angustifolia FATB codon optimized CDS nucleic acid sequence (CangFATB1) SEQ ID NO: 31 ATGGTGGCCGCCGCCGCCTCCTCCGCCTTCTTCCCCGTGCCCGCCCCCG GCACCTCCCTGAAGCCCGGCAAGTCCGGCAACTGGCCCTCCTCCCTGTC CCCCTCCTTCAAGCCCAAGACCATCCCCTCCGGCGGCCTGCAGGTGAAG GCCAACGCCTCCGCCCACCCCAAGGCCAACGGCTCCGCCGTGAACCTGA AGTCCGGCTCCCTGGACACCCAGGAGGACACCTCCTCCTCCCCCCCCCC CCGCGCCTTCCTGAACCAGCTGCCCGACTGGTCCATGCTGCTGACCGCC ATCACCACCGTGTTCGTGGCCGCCGAGAAGCAGTGGACCATGCTGGACC GCAAGTCCAAGCGCCCCGAGATGCTGGTGGACTCCGTGGGCCTGAAGTC CTCCGTGCGCGACGGCCTGGTGTCCCGCCAGTCCTTCTCCATCCGCTCC TACGAGATCGGCGCCGACCGCACCGCCTCCATCGAGACCCTGATGAACC ACCTGCAGGAGACCTCCATCAACCACTGCAAGTCCCTGGGCCTGCTGAA CGACGGCTTCGGCCGCACCCCCGGCATGTGCAAGAACGACCTGATCTGG GTGCTGACCAAGATGCAGATCATGGTGAACCGCTACCCCACCTGGGGCG ACACCGTGGAGGTGAACACCTGGTTCTCCCAGTCCGGCAAGATCGGCAT GGCCTCCGACTGGCTGATCTCCGACTGCAACACCGGCGAGATCCTGATC CGCGCCACCTCCGTGTGGGCCATGATGAACCAGAAGACCCGCCGCTTCT CCCGCCTGCCCTACGAGGTGCGCCAGGAGCTGACCCCCCACTACGTGGA CTCCCCCCACGTGATCGAGGACAACGACCGCAAGCTGCACAAGTTCGAC GTGAAGACCGGCGACTCCATCCGCAAGGGCCTGACCCCCCGCTGGAACG ACCTGGACGTGAACCAGCACGTGTCCAACGTGAAGTACATCGGCTGGAT CCTGGAGTCCATGCCCATCGAGGTGCTGGAGACCCAGGAGCTGTGCTCC CTGACCGTGGAGTACCGCCGCGAGTGCGGCATGGACTCCGTGCTGGAGT CCGTGACCGCCATGGACCCCTCCGAGGACGGCGGCGTGTCCCAGTACAA GCACCTGCTGCGCCTGGAGGACGGCACCGACATCGTGAAGGGCCGCACC GAGTGGCGCCCCAAGAACGCCGGCACCAACGGCGCCACCTCCAAGGCCA AGACCTCCAACGGCAACTCCGTGTCCTGA Cuphea llavea FATB1 codon optimized CDS nucleic acid sequence (CllaFATB1) SEQ ID NO: 32 ATGGTGGCCGCCGCCGCCTCCTCCGCCTTCTTCCCCGCCCCCGCCCCCG GCTCCTCCCCCAAGCCCGGCAAGCCCGGCAACTGGCCCTCCTCCCTGTC CCCCTCCTTCAAGCCCAAGTCCATCCCCAACGGCCGCTTCCAGGTGAAG GCCAACGCCTCCGCCCACCCCAAGGCCAACGGCTCCGCCGTGAACCTGA AGTCCGGCTCCCTGAACACCCAGGAGGACACCTCCTCCTCCCCCCCCCC CCGCGCCTTCCTGAACCAGCTGCCCGACTGGTCCATGCTGCTGTCCGCC ATCACCACCGTGTTCGTGGCCGCCGAGAAGCAGTGGACCATGCTGGACC GCAAGTCCAAGCGCCCCGACATGCTGGTGGACTCCGTGGGCCTGAAGAA CATCGTGCGCGACGGCCTGGTGTCCCGCCAGTCCTTCTCCATCCGCTCC TACGAGATCGGCGCCGACCGCACCGCCTCCATCGAGACCCTGATGAACC ACCTGCAGGAGACCTCCATCAACCACTGCAAGTCCCTGGGCCTGCTGAA CGACGGCTTCGGCCGCACCCCCGGCATGTGCAAGAACGACCTGATCTGG GTGCTGACCAAGATGCAGATCATGGTGAACCGCTACCCCGCCTGGGGCG ACACCGTGGAGATCAACACCTGGTTCTCCCAGTCCGGCAAGATCGGCAT GGGCTCCGACTGGCTGATCTCCGACTGCAACACCGGCGAGATCCTGATC CGCGCCACCTCCGTGTGGGCCATGATGAACCAGAAGACCCGCCGCTTCT CCCGCCTGCCCTACGAGGTGCGCCAGGAGCTGACCCCCCACTTCGTGGA CTCCCCCCACGTGATCGAGGACAACGACCGCAAGCTGCACAAGTTCGAC GTGAAGACCGGCGACTCCATCCGCAAGGGCCTGACCCCCCGCTGGAACG ACCTGGACGTGAACCAGCACGTGTCCAACGTGAAGTACATCGGCTGGAT CCTGGAGTCCATGCCCATCGAGGTGCTGGAGACCCAGGAGCTGTGCTCC CTGACCGTGGAGTACCGCCGCGAGTGCGGCATGGACTCCGTGCTGGAGT CCGTGACCGCCATCGACCCCTCCGAGGACGGCGGCCGCTCCCAGTACAA CCACCTGCTGCGCCTGGACGACGGCACCGACGTGGTGAAGGGCCGCACC GAGTGGCGCCCCAAGAACGCCGGCACCAACGGCGCCATCTCCACCGGCA AGACCTCCAACGGCAACTCCGTGTCCTGA Cuphea lophostoma FATB1 codon optimized CDS nucleic acid sequence (ClopFATB1) SEQ ID NO: 33 ATGGTGGCCGCCGCCGCCTCCTCCGCCTTCTTCCCCGTGCCCGCCCCCG GCACCTCCCTGAAGCCCTGGAAGTCCGGCAACTGGCCCTCCTCCCTGTC CCCCTCCTTCAAGCCCAAGACCATCCCCTCCGGCGGCTTCCAGGTGAAG GCCAACGCCTCCGCCCAGCCCAAGGCCAACGGCTCCGCCGTGAACCTGA AGTCCGGCTCCCTGAACACCCAGGAGGACACCACCTCCTCCCCCCCCCC CCGCGCCTTCCTGAACCAGCTGCCCGACTGGTCCATGCTGCTGACCGCC ATCACCACCGTGTTCGTGGCCGCCGAGAAGCAGTGGACCATGCTGGACC GCAAGTCCAAGCGCCCCGAGAAGCTGGTGGACTCCGTGGGCCTGAAGTC CTCCGTGCGCGACGGCCTGGTGTCCCGCCAGTCCTTCTCCATCCGCTCC TACGAGATCGGCGCCGACCGCACCGCCTCCATCGAGACCCTGATGAACC ACCTGCAGGAGACCTCCATCAACCACTGCAAGTCCCTGGGCCTGCTGAA CGACGGCTTCGGCCGCACCCCCGGCATGTGCAAGAACGACCTGATCTGG GTGCTGACCAAGATGCAGATCATGGTGAACCGCTACCCCACCTGGGGCG ACACCGTGGAGATCAACACCTGGTTCTCCCAGTCCGGCAAGATCGGCAT GGCCTCCGACTGGCTGATCTCCGACTGCAACACCGGCGAGATCCTGATC CGCGCCACCTCCGTGTGGGCCATGATGAACCAGAAGACCCGCCGCTTCT CCCGCCTGCCCTACGAGGTGCGCCAGGAGCTGACCCCCCACTACGTGGA CTCCCCCCACGTGATCGAGGACAACGACCGCAAGCTGCACAAGTTCGAC GTGAAGACCGGCGACTCCATCCGCAAGGGCCTGACCCCCCGCTGGAACG ACCTGGACGTGAACCAGCACGTGTCCAACGTGAAGTACATCGGCTGGAT CCTGGAGTCCATGCCCATCGAGGTGCTGGAGACCCAGGAGCTGTGCTCC CTGACCGTGGAGTACCGCCGCGAGTGCGGCATGGACTCCGTGCTGGAGT CCGTGACCGCCATGGACCCCTCCGAGGACGAGGGCCGCTCCCAGTACAA GCACCTGCTGCGCCTGGAGGACGGCACCGACATCGTGAAGGGCCGCACC GAGTGGCGCCCCAAGAACGCCGGCACCAACGGCGCCATCTCCACCGCCA AGAACTCCAACGGCAACTCCGTGTCCTGA Sassafras albidum FATB1 codon optimized CDS nucleic acid sequence (SalFATB1) SEQ ID NO: 34 ATGGCCACCACCTCCCTGGCCTCCGCCTTCTGCTCCATGAAGGCCGTGA TGCTGGCCCGCGACGGCCGCGGCATGAAGCCCCGCTCCTCCGACCTGCA GCTGCGCGCCGGCAACGCCCAGACCCCCCTGAAGATGATCAACGGCACC AAGTTCTCCTACACCGAGTCCCTGAAGCGCCTGCCCGACTGGTCCATGC TGTTCGCCGTGATCACCACCATCTTCTCCGTGGCCGAGAAGCAGTGGAC CAACCTGGAGTGGAAGCCCAAGCCCAAGCCCCGCCTGCCCCAGCTGCTG GACGACCACTTCGGCCTGCACGGCCTGGTGTTCCGCCGCACCTTCGCCA TCCGCTCCTACGAGGTGGGCCCCGACCGCTCCACCTCCATCGTGGCCGT GATGAACCACCTGCAGGAGGCCACCCTGAACCACGCCAAGTCCGTGGGC ATCCTGGGCGACGGCTTCGGCACCACCCTGGAGATGTCCAAGCGCGACC TGGCCTGGGTGGTGCGCCGCACCCACGTGGCCGTGGAGCGCTACCCCGC CTGGGGCGACACCGTGGAGGTGGAGTGCTGGATCGGCGCCTCCGGCAAC AACGGCATGCGCCGCGACTTCCTGGTGCGCGACTGCAAGACCGGCGAGA TCCTGACCCGCTGCACCTCCCTGTCCGTGATGATGAACACCCGCACCCG CCGCCTGTCCAAGATCCCCGAGGAGGTGCGCGGCGAGATCGGCCCCCTG TTCATCGACAACGTGGCCGTGAAGGACGAGGAGATCAAGAAGCTGCAGA AGCTGAACGACTCCTCCGCCGACTACATCCAGGGCGGCCTGACCCCCCG CTGGAACGACCTGGACGTGAACCAGCACGTGAACAACATCAAGTACGTG GGCTGGATCCTGGAGACCGTGCCCGACTCCATCTTCGAGTCCCACCACA TCTCCTCCATCACCCTGGAGTACCGCCGCGAGTGCACCCGCGACTCCGT GCTGCAGTCCCTGACCACCGTGTCCGGCGGCTCCCTGGAGGCCGGCCTG GTGTGCGACCACCTGCTGCAGCTGGAGGGCGGCTCCGAGGTGCTGCGCG CCCGCACCGAGTGGCGCCCCAAGCTGACCGACTCCTTCCGCGGCATCAT CGTGATCCCCGCCGAGCCCTCCGTGTGA Sassafras albidum FATB2 codon optimized CDS nucleic acid sequence (SalFATB2) SEQ ID NO: 35 ATGGCCACCACCTCCCTGGCCTCCGCCTTCTGCTCCATGAAGGCCGTGA TGCTGGCCCGCGACGGCCGCGGCATGAAGCCCCGCTCCTCCGACCTGCA GCTGCGCGCCGGCAACGCCCAGACCCCCCTGAAGATGATCAACGGCACC AAGTTCTCCTACACCGAGTCCCTGAAGCGCCTGCCCGACTGGTCCATGC TGTTCGCCGTGATCACCACCATCTTCTCCGTGGCCGAGAAGCAGTGGAC CAACCTGGAGTGGAAGCCCAAGCCCAAGCCCCGCCTGCCCCAGCTGCTG GACGACCACTTCGGCCTGCACGGCCTGGTGTTCCGCCGCACCTTCGCCA TCCGCTCCTACGAGGTGGGCCCCGACCGCTCCACCTCCATCGTGGCCGT
GATGAACCACCTGCAGGAGGCCACCCTGAACCACGCCAAGTCCGTGGGC ATCCTGGGCGACGGCTTCGGCACCACCCTGGAGATGTCCAAGCGCGACC TGGCCTGGGTGGTGCGCCGCACCCACGTGGCCGTGGAGCGCTACCCCGC CTGGGGCGACACCGTGGAGGTGGAGGCCTGGGTGGGCGCCTCCGGCAAC ATCGGCATGCGCCGCGACTTCCTGGTGCGCGACTGCAAGACCGGCCACA TCCTGGCCCGCTGCACCTCCGTGTCCGTGATGATGAACGCCCGCACCCG CCGCCTGTCCAAGATCCCCCAGGAGGTGCGCGCCGAGATCGACCCCCTG TTCATCGAGAAGGTGGCCGTGAAGGAGGGCGAGATCAAGAAGCTGCAGA AGTTCAACGACTCCACCGCCGACTACATCCAGGGCGGCTGGACCCCCCG CTGGAACGACCTGGACGTGAACCAGCACGTGAACAACATCAAGTACATC GGCTGGATCTTCAAGTCCGTGCCCGACTCCATCTCCGAGAACCACTACC TGTCCTCCATCACCCTGGAGTACCGCCGCGAGTGCACCCGCGGCTCCGC CCTGCAGTCCCTGACCACCGTGTGCGGCGACTCCTCCGAGGCCGGCATC ATCTGCGAGCACCTGCTGCAGCTGGAGGACGGCCCCGAGGTGCTGCGCG CCCGCACCGAGTGGCGCCCCAAGCTGACCGACTCCTTCCGCGGCATCAT CGTGATCCCCGCCGAGCCCTCCGTGTGA Lindera benzoin FATB1 codon optimized CDS nucleic acid sequence (LbeFATB1) SEQ ID NO: 36 ATGGTGACCGCCGCCGCCTCCTCCGCCTTCTTCCCCGTGCCCGCCCCCG GCACCTCCCCCAAGCCCGGCAAGTCCTGGCCCTCCTCCCTGTCCCCCTC CTTCAAGCCCAAGTCCATCCCCAACGCCGGCTTCCAGGTGAAGGCCAAC GCCTCCGCCCACCCCAAGGCCAACGGCTCCGCCGTGAACCTGAAGTCCG GCTCCCTGAACACCCAGGAGGACACCTCCTCCTCCCCCCCCCCCCGCGC CTTCCTGAACCAGCTGCCCGACTGGTCCATGCTGCTGACCGCCATCACC ACCGTGTTCGTGGCCGCCGAGAAGCAGTGGACCATGCGCGACCGCAAGT CCAAGCGCCCCGACATGCTGGTGGACTCCGTGGGCTCCAAGTCCATCGT GCTGGACGGCCTGGTGTCCCGCCAGATCTTCTCCATCCGCTCCTACGAG ATCGGCGCCGACCGCACCGCCTCCATCGAGACCCTGATGAACCACCTGC AGGAGACCTCCATCAACCACTGCAAGTCCCTGGGCCTGCTGAACGACGG CTTCGGCCGCACCCCCGGCATGTGCAAGAACGACCTGATCTGGGTGCTG ACCAAGATGCAGATCATGGTGAACCGCTACCCCACCTGGGGCGACACCG TGGAGATCAACACCTGGTTCTCCCACTCCGGCAAGATCGGCATGGCCTC CGACTGGCTGATCACCGACTGCAACACCGGCGAGATCCTGATCCGCGCC ACCTCCGTGTGGGCCATGATGAACCAGAAGACCCGCCGCTTCTCCCGCC TGCCCTACGAGGTGCGCCAGGAGCTGACCCCCCACTACGTGGACTCCCC CCACGTGATCGAGGACAACGACCGCAAGCTGCACAAGTTCGACGTGAAG ACCGGCGACTCCATCCGCAAGGGCCTGACCCCCAAGTGGAACGACCTGG ACGTGAACCAGCACGTGAACAACGTGAAGTACATCGGCTGGATCCTGGA GTCCATGCCCATCGAGGTGCTGGAGACCCAGGAGCTGTGCTCCCTGACC GTGGAGTACCGCCGCGAGTGCGGCATGGACTCCGTGCTGGAGTCCGTGA CCGCCATGGACCCCTCCGAGGACGGCGGCCTGTCCCAGTACAAGCACCT GCTGCGCCTGGAGGACGGCACCGACATCGTGAAGGGCCGCACCGAGTGG CGCCCCAAGAACGCCGGCACCAACGGCGCCATCTCCACCGCCAAGCCCT CCAACGGCAACTCCGTGTCCTGA CpSADtp_trimmed transit (plastid targeting) peptide amino acid sequence SEQ ID NO: 37 MATASTFSAFNARCGDLRRSAGSGPRRPARPLPVRAAI Thioesterase transforming construct SEQ ID NO: 38 gaagagcgcccaatgtttaaacccctcaactgcgacgctgggaaccttc tccgggcaggcgatgtgcgtgggtttgcctccttggcacggctctacac cgtcgagtacgccatgaggcggtgatggctgtgtcggttgccacttcgt ccagagacggcaagtcgtccatcctctgcgtgtgtggcgcgacgctgca gcagtccctctgcagcagatgagcgtgactttggccatttcacgcactc gagtgtacacaatccatttttcttaaagcaaatgactgctgattgacca gatactgtaacgctgatttcgctccagatcgcacagatagcgaccatgt tgctgcgtctgaaaatctggattccgaattcgaccctggcgctccatcc atgcaacagatggcgacacttgttacaattcctgtcacccatcggcatg gagcaggtccacttagattcccgatcacccacgcacatctcgctaatag tcattcgttcgtgtcttcgatcaatctcaagtgagtgtgcatggatctt ggttgacgatgcggtatgggtttgcgccgctggctgcagggtctgccca aggcaagctaacccagctcctctccccgacaatactctcgcaggcaaag ccggtcacttgccttccagattgccaataaactcaattatggcctctgt catgccatccatgggtctgatgaatggtcacgctcgtgtcctgaccgtt ccccagcctctggcgtcccctgccccgcccaccagcccacgccgcgcgg ##STR00001## ##STR00002## tcgccgccaagatcagcgcctccatgacgaacgagacgtccgaccgccc cctggtgcacttcacccccaacaagggctggatgaacgaccccaacggc ctgtggtacgacgagaaggacgccaagtggcacctgtacttccagtaca acccgaacgacaccgtctgggggacgcccttgttctggggccacgccac gtccgacgacctgaccaactgggaggaccagcccatcgccatcgccccg aagcgcaacgactccggcgccttctccggctccatggtggtggactaca acaacacctccggcttcttcaacgacaccatcgacccgcgccagcgctg cgtggccatctggacctacaacaccccggagtccgaggagcagtacatc tcctacagcctggacggcggctacaccttcaccgagtaccagaagaacc ccgtgctggccgccaactccacccagttccgcgacccgaaggtcttctg gtacgagccctcccagaagtggatcatgaccgcggccaagtcccaggac tacaagatcgagatctactcctccgacgacctgaagtcctggaagctgg agtccgcgttcgccaacgagggcttcctcggctaccagtacgagtgccc cggcctgatcgaggtccccaccgagcaggaccccagcaagtcctactgg gtgatgttcatctccatcaaccccggcgccccggccggcggctccttca accagtacttcgtcggcagcttcaacggcacccacttcgaggccttcga caaccagtcccgcgtggtggacttcggcaaggactactacgccctgcag accttcttcaacaccgacccgacctacgggagcgccctgggcatcgcgt gggcctccaactgggagtactccgccttcgtgcccaccaacccctggcg ctcctccatgtccctcgtgcgcaagttctccctcaacaccgagtaccag gccaacccggagacggagctgatcaacctgaaggccgagccgatcctga acatcagcaacgccggcccctggagccggttcgccaccaacaccacgtt gacgaaggccaacagctacaacgtcgacctgtccaacagcaccggcacc ctggagttcgagctggtgtacgccgtcaacaccacccagacgatctcca agtccgtgttcgcggacctctccctctggttcaagggcctggaggaccc cgaggagtacctccgcatgggcttcgaggtgtccgcgtcctccttcttc ctggaccgcgggaacagcaaggtgaagttcgtgaaggagaacccctact tcaccaaccgcatgagcgtgaacaaccagcccttcaagagcgagaacga cctgtcctactacaaggtgtacggcttgctggaccagaacatcctggag ctgtacttcaacgacggcgacgtcgtgtccaccaacacctacttcatga ccaccgggaacgccctgggctccgtgaacatgacgacgggggtggacaa ##STR00003## acgcccgcgcggcgcacctgacctgttctctcgagggcgcctgttctgc cttgcgaaacaagcccctggagcatgcgtgcatgatcgtctctggcgcc ccgccgcgcggtttgtcgccctcgcgggcgccgcggccgcgggggcgca ttgaaattgttgcaaaccccacctgacagattgagggcccaggcaggaa ggcgttgagatggaggtacaggagtcaagtaactgaaagtttttatgat aactaacaacaaagggtcgtttctggccagcgaatgacaagaacaagat tccacatttccgtgtagaggcttgccatcgaatgtgagcgggcgggccg cggacccgacaaaacccttacgacgtggtaagaaaaacgtggcgggcac tgtccctgtagcctgaagaccagcaggagacgatcggaagcatcacagc acaggatccCGCGTCTCGAACAGAGCGCGCAGAGGAACGCTGAAGGTCT CGCCTCTGTCGCACCTCAGCGCGGCATACACCACAATAACCACCTGACG AATGCGCTTGGTTCTTCGTCCATTAGCGAAGCGTCCGGTTCACACACGT GCCACGTTGGCGAGGTGGCAGGTGACAATGATCGGTGGAGCTGATGGTC ##STR00004## gcggcgacctgcgccgctccgccggctccggcccccgccgccccgcccg ccccctgcccgtgcgcgccgccatcaacgcctccgcccaccccaaggcc aacggctccgccgtgaacctgaagtccggctccctgaacacccaggagg acacctcctcctccccccccccccgcgccttcctgaaccagctgcccga ctggtccatgctgctgaccgccatcaccaccgtgttcgtggccgccgag aagcagtggaccatgctggaccgcaagtccaagcgccccgacatgctgg tggactccgtgggcctgaagtccatcgtgcgcgacggcctggtgtcccg ccagtccttctccatccgctcctacgagatcggcgccgaccgcaccgcc tccatcgagaccctgatgaaccacctgcaggagacctccatcaaccact gcaagtccctgggcctgctgaacgacggcttcggccgcacccccggcat gtgcaagaacgacctgatctgggtgctgaccaagatgcagatcatggtg aaccgctaccccacctggggcgacaccgtggagatcaacacctggttct cccagtccggcaagatcggcatgggctccgactggctgatctccgactg caacaccggcgagatcctgatccgcgccacctccgtgtgggccatgatg aaccagaagacccgccgcttctcccgcctgccctacgaggtgcgccagg agctgaccccccacttcgtggactccccccacgtgatcgaggacaacga ccgcaagctgcacaagttcgacgtgaagaccggcgactccatccgcaag
ggcctgaccccccgctggaacgacctggacgtgaaccagcacgtgtcca acgtgaagtacatcggctggatcctggagtccatgcccatcgaggtgct ggagacccaggagctgtgctccctgaccgtggagtaccgccgcgagtgc ggcatggactccaagctggagtccgtgaccgccatggacccctccgagg aggacggcgtgcgctcccagtacaaccacctgctgcgcctggaggacgg caccgacgtggtgaagggccgcaccgagtggcgccccaagaacgccggc accaacggcgccatctccaccggcaagacctccaacggcaactccgtgt ccatggactacaaggaccacgacggcgactacaaggaccacgacatcga ##STR00005## gtatcgacacactctggacgctggtcgtgtgatggactgttgccgccac acttgctgccttgacctgtgaatatccctgccgcttttatcaaacagcc tcagtgtgtttgatcttgtgtgtacgcgcttttgcgagttgctagctgc ttgtgctatttgcgaataccacccccagcatccccttccctcgtttcat atcgcttgcatcccaaccgcaacttatctacgctgtcctgctatccctc agcgctgctcctgctcctgctcactgcccctcgcacagccttggtttgg gctccgcctgtattctcctggtactgcaacctgtaaaccagcactgcaa tgctgatgcacgggaagtagtgggatgggaacacaaatggaaagcttga gctccagcgccatgccacgccctttgatggcttcaagtacgattacggt gttggattgtgtgtttgttgcgtagtgtgcatggtttagaataatacac ttgatttcttgctcacggcaatctcggcttgtccgcaggttcaacccca tttcggagtctcaggtcagccgcgcaatgaccagccgctacttcaagga cttgcacgacaacgccgaggtgagctatgtttaggacttgattggaaat tgtcgtcgacgcatattcgcgctccgcgacagcacccaagcaaaatgtc aagtgcgttccgatttgcgtccgcaggtcgatgttgtgatcgtcggcgc cggatccgccggtctgtcctgcgcttacgagctgaccaagcaccctgac gtccgggtacgcgagctgagattcgattagacataaattgaagattaaa cccgtagaaaaatttgatggtcgcgaaactgtgctcgattgcaagaaat tgatcgtcctccactccgcaggtcgccatcatcgagcagggcgttgctc ccggcggcggcgcctggctggggggacagctgttctcggccatgtgtgt acgtagaaggatgaatttcagctggttttcgttgcacagctgtttgtgc atgatttgtttcagactattgttgaatgtttttagatttcttaggatgc atgatttgtctgcatgcgactgaagagc Amino acid sequence of region of deletion mutant based on SalFATB2 SEQ ID NO: 39 LFAVITTIFSVAEKQWTNLEWKPKPKPRLPQL Amino acid sequence of SalFATB1a (a deletion mutant of Sassafras albidum SalFATB1) SEQ ID NO: 40 MATTSLASAFCSMKAVMLARDGRGMKPRSSDLQLRAGNAQTPLKMINGT KESYTESLKRLPDWSMLDDHFGLHGLVERRTFAIRSYEVGPDRSTSIVA VMNHLQEATLNHAKSVGILGDGFGTTLEMSKRDLAWVVRRTHVAVERYP AWGDTVEVECWIGASGNNGMRRDFLVRDCKTGEILTRCTSLSVMMNTRT RRLSKIPEEVRGEIGPLFIDNVAVKDEEIKKLQKLNDSSADYIQGGLTP RWNDLDVNQHVNNIKYVGWILETVPDSIFESHHISSITLEYRRECTRDS VLQSLTTVSGGSLEAGLVCDHLLQLEGGSEVLRARTEWRPKLTDSFRGI IVIPAEPSV Amino acid sequence of LbeFATB1a(a deletion mutant of LbeFATB1) SEQ ID NO: 41 MVATSLASAFCSMKAVMLADDGRGMKPRSSDLQLRAGNAQTSLKMIDGT KFSYTESLKRLPDWSKLDDRFGLHGLVFRRTFAIRSYEVGPDRSASILA VLNHLQEATLNHAESVGILGDRFGETLEMSKRDLMWVVRRTYVAVERYP AWGDTVEIESWIGASGNNGMRREFLVRDFKTGEILTRCTSLSVMMNTRT RRLSKIPEEVRGEIGPVFIDNVAVKDEEIKKLQKLNDSTADYIQGGLIP RWNDLDLNQHVNNIKYVSWILETVPDSILESYHMSSITLEYRRECTRDS VLQSLTTVSGGSSEAGLVCEHSLLLEGGSEVLRARTEWRPKLTDSFRGI SVIPAEQSV Amino acid sequence of mature SalFATB1a (a deletion mutant of Sassafras albidum SalFATB1) SEQ ID NO: 42 GNAQTPLKMINGTKFSYTESLKRLPDWSMLDDHFGLHGLVFRRTFAIRS YEVGPDRSTSIVAVMNHLQEATLNHAKSVGILGDGFGTTLEMSKRDLAW VVRRTHVAVERYPAWGDTVEVECWIGASGNNGMRRDFLVRDCKTGEILT RCTSLSVMMNTRTRRLSKIPEEVRGEIGPLFIDNVAVKDEEIKKLQKLN DSSADYIQGGLTPRWNDLDVNQHVNNIKYVGWILETVPDSIFESHHISS ITLEYRRECTRDSVLQSLTTVSGGSLEAGLVCDHLLQLEGGSEVLRART EWRPKLTDSFRGIIVIPAEPSV Amino acid sequence of mature LbeFATB1a(a deletion mutant of LbeFATB1) SEQ ID NO: 43 GNAQTSLKMIDGTKFSYTESLKRLPDWSKLDDRFGLHGLVFRRTFAIRS YEVGPDRSASILAVLNHLQEATLNHAESVGILGDRFGETLEMSKRDLMW VVRRTYVAVERYPAWGDTVEIESWIGASGNNGMRREFLVRDFKTGEILT RCTSLSVMMNTRTRRLSKIPEEVRGEIGPVFIDNVAVKDEEIKKLQKLN DSTADYIQGGLIPRWNDLDLNQHVNNIKYVSWILETVPDSILESYHMSS ITLEYRRECTRDSVLQSLTTVSGGSSEAGLVCEHSLLLEGGSEVLRART EWRPKLTDSFRGISVIPAEQSV pSZ5176/D4053 (SalFATB1a) SEQ ID NO: 44 ##STR00006## tgcgccgctccgccggctccggcccccgccgccccgcccgccccctgcc cgtgcgcgccgccatcggcaacgcccagacccccctgaagatgatcaac ggcaccaagttctcctacaccgagtccctgaagcgcctgcccgactggt ccatgctggacgaccacttcggcctgcacggcctggtgttccgccgcac cttcgccatccgctcctacgaggtgggccccgaccgctccacctccatc gtggccgtgatgaaccacctgcaggaggccaccctgaaccacgccaagt ccgtgggcatcctgggcgacggcttcggcaccaccctggagatgtccaa gcgcgacctggcctgggtggtgcgccgcacccacgtggccgtggagcgc taccccgcctggggcgacaccgtggaggtggagtgctggatcggcgcct ccggcaacaacggcatgcgccgcgacttcctggtgcgcgactgcaagac cggcgagatcctgacccgctgcacctccctgtccgtgatgatgaacacc cgcacccgccgcctgtccaagatccccgaggaggtgcgcggcgagatcg gccccctgttcatcgacaacgtggccgtgaaggacgaggagatcaagaa gctgcagaagctgaacgactcctccgccgactacatccagggcggcctg accccccgctggaacgacctggacgtgaaccagcacgtgaacaacatca agtacgtgggctggatcctggagaccgtgcccgactccatcttcgagtc ccaccacatctcctccatcaccctggagtaccgccgcgagtgcacccgc gactccgtgctgcagtccctgaccaccgtgtccggcggctccctggagg ccggcctggtgtgcgaccacctgctgcagctggagggcggctccgaggt gctgcgcgcccgcaccgagtggcgccccaagctgaccgactccttccgc ggcatcatcgtgatccccgccgagccctccgtgatggactacaaggacc acgacggcgactacaaggaccacgacatcgactacaaggacgacgacga ##STR00007## pSZ5179/D4056 (LbeFATB1a) SEQ ID NO: 45 ##STR00008## tgcgccgctccgccggctccggcccccgccgccccgcccgccccctgcc cgtgcgcgccgccatcggcaacgcccagacctccctgaagatgatcgac ggcaccaagttctcctacaccgagtccctgaagcgcctgcccgactggt ccaagctggacgaccgcttcggcctgcacggcctggtgttccgccgcac cttcgccatccgctcctacgaggtgggccccgaccgctccgcctccatc ctggccgtgctgaaccacctgcaggaggccaccctgaaccacgccgagt ccgtgggcatcctgggcgaccgcttcggcgagaccctggagatgtccaa gcgcgacctgatgtgggtggtgcgccgcacctacgtggccgtggagcgc taccccgcctggggcgacaccgtggagatcgagtcctggatcggcgcct ccggcaacaacggcatgcgccgcgagttcctggtgcgcgacttcaagac cggcgagatcctgacccgctgcacctccctgtccgtgatgatgaacacc cgcacccgccgcctgtccaagatccccgaggaggtgcgcggcgagatcg gccccgtgttcatcgacaacgtggccgtgaaggacgaggagatcaagaa gctgcagaagctgaacgactccaccgccgactacatccagggcggcctg atcccccgctggaacgacctggacctgaaccagcacgtgaacaacatca agtacgtgtcctggatcctggagaccgtgcccgactccatcctggagtc ctaccacatgtcctccatcaccctggagtaccgccgcgagtgcacccgc gactccgtgctgcagtccctgaccaccgtgtccggcggctcctccgagg ccggcctggtgtgcgagcactccctgctgctggagggcggctccgaggt gctgcgcgcccgcaccgagtggcgccccaagctgaccgactccttccgc ggcatctccgtgatccccgccgagcagtccgtgatggactacaaggacc acgacggcgactacaaggaccacgacatcgactacaaggacgacgacga ##STR00009## CcFATB4 Cinnamomum camphora acyl-ACP thioesterase CDS. SEQ ID NO: 46 MVTTSLASAYFSMKAVMLAPDGRGIKPRSSGLQVRAGNERNSCKVINGT KVKDTEGLKGCSTLQGQSMLDDHFGLHGLVFRRTFAIRCYEVGPDRSTS IMAVMNHLQEAARNHAESLGLLGDGFGETLEMSKRDLIWVVRRTHVAVE RYPAWGDTVEVEAWVGASGNTGMRRDFLVRDCKTGHILTRCTSVSVMMN MRTRRLSKIPQEVRAEIDPLFIEKVAVKEGEIKKLQKLNDSTADYIQGG WTPRWNDLDVNQHVNNIIYVGWIFKSVPDSISENHHLSSITLEYRRECT
RGNKLQSLTTVCGGSSEAGIICEHLLQLEDGSEVLRARTEWRPKHTDSF QGISERFPQQEPHK Deleted portion of SalFatB1 absent from SalFatB1a SEQ ID NO: 47 LFAVITTIFSVAEKQWTNLEWKPKPKPRLPQL Deleted portion of LbeFATB1 absent from LbeFATB1a SEQ ID NO: 48 LLTVITTIFSAAEKQWTNLERKPKPPHL CpauFATB1 (transit peptide appears in boxed text) SEQ ID NO: 49 ##STR00010## AVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLLTAITTVFVAAEKQW TMRDRKSKRPDMLVDSVGLKSVVLDGLVSRQIFSIRSYEIGADRTASIE TLMNHLQETSINHCKSLGLLNDGFGRTPGMCKNDLIWVLTKMQIMVNRY PTWGDTVEINTWFSHSGKIGMASDWLITDCNTGEILIRATSVWAMMNQK TRRFSRLPYEVRQELTPHYVDSPHVIEDNDRKLHKFDVKTGDSIRKGLT PRWNDLDVNQHVSNVKYIGWILESMPIEVLETQELCSLTVEYRRECGMD SVLESVTAMDPSEDEGRSQYKHLLRLEDGTDIVKGRTEWRPKNAGTNGA ISTAKPSNGNSVS CpauFATB1.DELTA.28 deletion mutant of Cuphea paucipetala FATB1 acyl-ACP thioesterase SEQ ID NO: 50 ##STR00011## AVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLVDSVGLKSVVLDGLV SRQIFSIRSYEIGADRTASIETLMNHLQETSINHCKSLGLLNDGFGRTP GMCKNDLIWVLTKMQIMVNRYPTWGDTVEINTWFSHSGKIGMASDWLIT DCNTGEILIRATSVWAMMNQKTRRFSRLPYEVRQELTPHYVDSPHVIED NDRKLHKFDVKTGDSIRKGLTPRWNDLDVNQHVSNVKYIGWILESMPIE VLETQELCSLTVEYRRECGMDSVLESVTAMDPSEDEGRSQYKHLLRLED GTDIVKGRTEWRPKNAGTNGAISTAKPSNGNSVS Mature CpauFATB1.DELTA.28 deletion mutant of Cuphea paucipetala FATB1 acyl-ACP thioesterase SEQ ID NO: 51 NASAHPKANGSAVNLKSGSLNTQEDTSSSPPPRAFLNQLPDWSMLVDSV GLKSVVLDGLVSRQIFSIRSYEIGADRTASIETLMNHLQETSINHCKSL GLLNDGFGRTPGMCKNDLIWVLTKMQIMVNRYPTWGDTVEINTWFSHSG KIGMASDWLITDCNTGEILIRATSVWAMMNQKTRRFSRLPYEVRQELTP HYVDSPHVIEDNDRKLHKFDVKTGDSIRKGLTPRWNDLDVNQHVSNVKY IGWILESMPIEVLETQELCSLTVEYRRECGMDSVLESVTAMDPSEDEGR SQYKHLLRLEDGTDIVKGRTEWRPKNAGTNGAISTAKPSNGNSVS CpauFATB1.DELTA.28 CDS SEQ ID NO: 52 ##STR00012## tgcgccgctccgccggctccggcccccgccgccccgcccgccccctgcc cgtgcgcgccgccatcaacgcctccgcccaccccaaggccaacggctcc gccgtgaacctgaagtccggctccctgaacacccaggaggacacctcct cctccccccccccccgcgccttcctgaaccagctgcccgactggtccat gctggtggactccgtgggcctgaagtccgtggtgctggacggcctggtg tcccgccagatcttctccatccgctcctacgagatcggcgccgaccgca ccgcctccatcgagaccctgatgaaccacctgcaggagacctccatcaa ccactgcaagtccctgggcctgctgaacgacggcttcggccgcaccccc ggcatgtgcaagaacgacctgatctgggtgctgaccaagatgcagatca tggtgaaccgctaccccacctggggcgacaccgtggagatcaacacctg gttctcccactccggcaagatcggcatggcctccgactggctgatcacc gactgcaacaccggcgagatcctgatccgcgccacctccgtgtgggcca tgatgaaccagaagacccgccgcttctcccgcctgccctacgaggtgcg ccaggagctgaccccccactacgtggactccccccacgtgatcgaggac aacgaccgcaagctgcacaagttcgacgtgaagaccggcgactccatcc gcaagggcctgaccccccgctggaacgacctggacgtgaaccagcacgt gtccaacgtgaagtacatcggctggatcctggagtccatgcccatcgag gtgctggagacccaggagctgtgctccctgaccgtggagtaccgccgcg agtgcggcatggactccgtgctggagtccgtgaccgccatggacccctc cgaggacgagggccgctcccagtacaagcacctgctgcgcctggaggac ggcaccgacatcgtgaagggccgcaccgagtggcgccccaagaacgccg gcaccaacggcgccatctccaccgccaagccctccaacggcaactccgt gtccatggactacaaggaccacgacggcgactacaaggaccacgacatc ##STR00013## ChFATB2 (Uniprot Q39514) SEQ ID NO: 53 MVAAAASSAFFPVPAPGASPKPGKFGNWPSSLSPSFKPKSIPNGGFQVK ANDSAHPKANGSAVSLKSGSLNTQEDTSSSPPPRTFLHQLPDWSRLLTA ITTVFVKSKRPDMHDRKSKRPDMLVDSFGLESTVQDGLVFRQSFSIRSY EIGTDRTASIETLMNHLQETSLNHCKSTGILLDGFGRTLEMCKRDLIWV VIKMQIKVNRYPAWGDTVEINTRFSRLGKIGMGRDWLISDCNTGEILVR ATSAYAMMNQKTRRLSKLPYEVHQEIVPLFVDSPVIEDSDLKVHKFKVK TGDSIQKGLTPGWNDLDVNQHVSNVKYIGWILESMPTEVLETQELCSLA LEYRRECGRDSVLESVTAMDPSKVGVRSQYQHLLRLEDGTAIVNGATEW RPKNAGANGAISTGKTSNGNSVS ChFATB2.DELTA.27 Deletion mutant of Cuphea hookeriana FATB2 SEQ ID NO: 54 ##STR00014## AVSLKSGSLNTQEDTSSSPPPRTFLHQLPDWSRLVDSFGLESTVQDGLV FRQSFSIRSYEIGTDRTASIETLMNHLQETSLNHCKSTGILLDGFGRTL EMCKRDLIWVVIKMQIKVNRYPAWGDTVEINTRFSRLGKIGMGRDWLIS DCNTGEILVRATSAYAMMNQKTRRLSKLPYEVHQEIVPLFVDSPVIEDS DLKVHKFKVKTGDSIQKGLTPGWNDLDVNQHVSNVKYIGWILESMPTEV LETQELCSLALEYRRECGRDSVLESVTAMDPSKVGVRSQYQHLLRLEDG TAIVNGATEWRPKNAGANGAISTGKTSNGNSVS ChFATB2.DELTA.27 Mature deletion mutant of Cuphea hookeriana FATB2 SEQ ID NO: 55 NDSAHPKANGSAVSLKSGSLNTQEDTSSSPPPRTFLHQLPDWSRLVDSF GLESTVQDGLVFRQSFSIRSYEIGTDRTASIETLMNHLQETSLNHCKST GILLDGFGRTLEMCKRDLIWVVIKMQIKVNRYPAWGDTVEINTRFSRLG KIGMGRDWLISDCNTGEILVRATSAYAMMNQKTRRLSKLPYEVHQEIVP LFVDSPVIEDSDLKVHKFKVKTGDSIQKGLTPGWNDLDVNQHVSNVKYI GWILESMPTEVLETQELCSLALEYRRECGRDSVLESVTAMDPSKVGVRS QYQHLLRLEDGTAIVNGATEWRPKNAGANGAISTGKTSNGNSVS ChFATB2.DELTA.27 CDS SEQ ID NO: 56 ##STR00015## tgcgccgctccgccggctccggcccccgccgccccgcccgccccctgcc cgtgcgcgccgccatcaacgactccgcccaccccaaggccaacggctcc gccgtgagcctgaagtccggcagcctgaacacccaggaggacacctcct ccagcccccccccccgcaccttcctgcaccagctgcccgactggagccg cctggtggacagcttcggcctggagtccaccgtgcaggacggcctggtg ttccgccagtccttctccatccgctcctacgagatcggcaccgaccgca ccgccagcatcgagaccctgatgaaccacctgcaggagacctccctgaa ccactgcaagagcaccggcatcctgctggacggcttcggccgcaccctg gagatgtgcaagcgcgacctgatctgggtggtgatcaagatgcagatca aggtgaaccgctaccccgcctggggcgacaccgtggagatcaacacccg cttcagccgcctgggcaagatcggcatgggccgcgactggctgatctcc gactgcaacaccggcgagatcctggtgcgcgccaccagcgcctacgcca tgatgaaccagaagacccgccgcctgtccaagctgccctacgaggtgca ccaggagatcgtgcccctgttcgtggacagccccgtgatcgaggactcc gacctgaaggtgcacaagttcaaggtgaagaccggcgacagcatccaga agggcctgacccccggctggaacgacctggacgtgaaccagcacgtgtc caacgtgaagtacatcggctggatcctggagagcatgcccaccgaggtg ctggagacccaggagctgtgctccctggccctggagtaccgccgcgagt gcggccgcgactccgtgctggagagcgtgaccgccatggaccccagcaa ggtgggcgtgcgctcccagtaccagcacctgctgcgcctggaggacggc accgccatcgtgaacggcgccaccgagtggcgccccaagaacgccggcg ccaacggcgccatctccaccggcaagaccagcaacggcaactccgtgtc catggactacaaggaccacgacggcgactacaaggaccacgacatcgac ##STR00016## Nucleotide sequence of transforming DNA contained in pSZ6315 SEQ ID NO: 57 caccggcgcgctgcttcgcgtgccgggtgcagcaatcagatccaagtct gacgacttgcgcgcacgcgccggatccttcaattccaaagtgtcgtccg cgtgcgcttcttcgccttcgtcctcttgaacatccagcgacgcaagcgc agggcgctgggcggctggcgtcccgaaccggcctcggcgcacgcggctg aaattgccgatgtcggcaatgtagtgccgctccgcccacctctcaatta agtttttcagcgcgtggttgggaatgatctgcgctcatggggcgaaaga aggggttcagaggtgctttattgttactcgactgggcgtaccagcattc gtgcatgactgattatacatacaaaagtacagctcgcttcaatgccctg cgattcctactcccgagcgagcactcctctcaccgtcgggttgcttccc acgaccacgccggtaagagggtctgtggcctcgcgcccctcgcgagcgc atctttccagccacgtctgtatgattttgcgctcatacgtctggcccgt cgaccccaaaatgacgggatcctgcataatatcgcccgaaatgggatcc aggcattcgtcaggaggcgtcagccccgcgggagatgccggtcccgccg
##STR00017## ggcctgcatctccctgaagggcgtgttcggcgtctccccctcctacaac ggcctgggcctgacgccccagatgggctgggacaactggaacacgttcg cctgcgacgtctccgagcagctgctgctggacacggccgaccgcatctc cgacctgggcctgaaggacatgggctacaagtacatcatcctggacgac tgctggtcctccggccgcgactccgacggcttcctggtcgccgacgagc agaagttccccaacggcatgggccacgtcgccgaccacctgcacaacaa ctccttcctgttcggcatgtactcctccgcgggcgagtacacgtgcgcc ggctaccccggctccctgggccgcgaggaggaggacgcccagttcttcg cgaacaaccgcgtggactacctgaagtacgacaactgctacaacaaggg ccagttcggcacgcccgagatctcctaccaccgctacaaggccatgtcc gacgccctgaacaagacgggccgccccatcttctactccctgtgcaact ggggccaggacctgaccttctactggggctccggcatcgcgaactcctg gcgcatgtccggcgacgtcacggcggagttcacgcgccccgactcccgc tgcccctgcgacggcgacgagtacgactgcaagtacgccggcttccact gctccatcatgaacatcctgaacaaggccgcccccatgggccagaacgc gggcgtcggcggctggaacgacctggacaacctggaggtcggcgtcggc aacctgacggacgacgaggagaaggcgcacttctccatgtgggccatgg tgaagtcccccctgatcatcggcgcgaacgtgaacaacctgaaggcctc ctcctactccatctactcccaggcgtccgtcatcgccatcaaccaggac tccaacggcatccccgccacgcgcgtctggcgctactacgtgtccgaca cggacgagtacggccagggcgagatccagatgtggtccggccccctgga caacggcgaccaggtcgtggcgctgctgaacggcggctccgtgtcccgc cccatgaacacgaccctggaggagatcttcttcgactccaacctgggct ccaagaagctgacctccacctgggacatctacgacctgtgggcgaaccg cgtcgacaactccacggcgtccgccatcctgggccgcaacaagaccgcc accggcatcctgtacaacgccaccgagcagtcctacaaggacggcctgt ccaagaacgacacccgcctgttcggccagaagatcggctccctgtcccc caacgcgatcctgaacacgaccgtccccgcccacggcatcgcgttctac ##STR00018## ctgatgtggcgcggacgccgtcgtactctttcagactttactcttgagg aattgaacctttctcgcttgctggcatgtaaacattggcgcaattaatt gtgtgatgaagaaagggtggcacaagatggatcgcgaatgtacgagatc gacaacgatggtgattgttatgaggggccaaacctggctcaatcttgtc gcatgtccggcgcaatgtgatccagcggcgtgactctcgcaacctggta gtgtgtgcgcaccgggtcgctttgattaaaactgatcgcattgccatcc cgtcaactcacaagcctactctagctcccattgcgcactcgggcgcccg gctcgatcaatgttctgagcggagggcgaagcgtcaggaaatcgtctcg gcagctggaagcgcatggaatgcggagcggagatcgaatcaggatcccg cgtctcgaacagagcgcgcagaggaacgctgaaggtctcgcctctgtcg cacctcagcgcggcatacaccacaataaccacctgacgaatgcgcttgg ttcttcgtccattagcgaagcgtccggttcacacacgtgccacgttggc gaggtggcaggtgacaatgatcggtggagctgatggtcgaaacgttcac ##STR00019## ##STR00020## gcgtgcggggctggcgggagtgggacgccctcctcgctcctctctgttc tgaacggaacaatcggccaccccgcgctacgcgccacgcatcgagcaac gaagaaaaccccccgatgataggttgcggtggctgccgggatatagatc cggccgcacatcaaagggcccctccgccagagaagaagctcctttccca gcagactccttctgctgccaaaacacttctctgtccacagcaacaccaa aggatgaacagatcaacttgcgtctccgcgtagcttcctcggctagcgt gcttgcaacaggtccctgcactattatcttcctgctttcctctgaatta tgcggcaggcgagcgctcgctctggcgagcgctccttcgcgccgccctc gctgatcgagtgtacagtcaatgaatggtgagctccgcgcctgcgcgag gacgcagaacaacgctgccgccgtgtcttttgcacgcgcgactccggcg cttcgctggtggcacccccataaagaaaccctcaattctgtttgtggaa gacacggtgtacccccacccacccacctgcacctctattattggtatta ttgacgcgggagtgggcgttgtaccctacaacgtagcttctctagtttt cagctggctcccaccattgtaaattcatgctagaatagtgcgtggttat gtgagaggtatagtgtgtctgagcagacggggcgggatgcatgtcgtgg tggtgatctttggctcaaggcgtcgtcgacgtgacgtgcccgatcatga gagcaataccgcgctcaaagccgacgcatagcctttactccgcaatcca aacgactgtcgctcgtattttttggatatctattttaaagagcgagcac agcgccgggcatgggcctgaaaggcctcgcggccgtgctcgtggtgggg gccgcgagcgcgtggggcatcgcggcagtgcaccaggcgcagacggagg aacgcatggtgcgtgcgcaatataagatacatgtattgttgtcctgcag g Nucleotide sequence of BnOTE (D124A) in pSZ6316 SEQ ID NO: 58 ##STR00021## Nucleotide sequence of BnOTE (D209A) in pSZ6317 SEQ ID NO: 59 ##STR00022## Nucleotide sequence of BnOTE (D124A, D209A) in pSZ6318 SEQ ID NO: 60 ##STR00023## Amino acid sequence of wild-type BnOTE; positions D124 and D209 underlined SEQ ID NO: 61 MATASTFSAFNARCGDLRRSAGSGPRRPARPLPVRGRASQLRKPALDPL RAVISADQGSISPVNSCTPADRLRAGRLMEDGYSYKEKFIVRSYEVGIN KTATVETIANLLQEVACNHVQKCGFSTDGFATTLTMRKLHLIWVTARMH IEIYKYPAWSDVVEIETWCQSEGRIGIRRDWILRDSATNEVIGRATSKW VMMNQDTRRLQRVIDEVRDEYLVFCPREPRLAFPEENNSSLKKIPKLED PAQYSMLELKPRRADLDMNQHVNNVTYIGWVLESIPQEIIDTHELQVIT LDYRRECQQDDIVDSLTTSEIPDDPISKFTGTNGSAMSSIQGHNESQFL HMLRLSENGQEINRGRTQWRKKSSR*
Sequence CWU 1
1
621418PRTCuphea crassiflora 1Met Val Ala Ala Ala Ala Ser Ser Ala
Phe Phe Pro Val Pro Ala Pro1 5 10 15Gly Thr Ser Thr Lys Pro Arg Lys
Ser Gly Asn Trp Pro Ser Arg Leu 20 25 30Ser Pro Ser Ser Lys Pro Lys
Ser Ile Pro Asn Gly Gly Phe Gln Val 35 40 45Lys Ala Asn Ala Ser Ala
His Pro Lys Ala Asn Gly Ser Ala Val Asn 50 55 60Leu Lys Ser Gly Ser
Leu Asn Thr Gln Glu Asp Thr Ser Ser Ser Pro65 70 75 80Pro Pro Arg
Ala Phe Leu Asn Gln Leu Pro Asp Trp Ser Met Leu Leu 85 90 95Thr Ala
Ile Thr Thr Val Phe Val Ala Ala Glu Lys Gln Trp Thr Met 100 105
110Leu Asp Arg Lys Ser Lys Arg Pro Asp Met Leu Val Asp Ser Val Gly
115 120 125Leu Lys Ser Ile Val Arg Asp Gly Leu Val Ser Arg Gln Ser
Phe Ser 130 135 140Ile Arg Ser Tyr Glu Ile Gly Ala Asp Arg Thr Ala
Ser Ile Glu Thr145 150 155 160Leu Met Asn His Leu Gln Glu Thr Ser
Ile Asn His Cys Lys Ser Leu 165 170 175Gly Leu Leu Asn Asp Gly Phe
Gly Arg Thr Pro Gly Met Cys Lys Asn 180 185 190Asp Leu Ile Trp Val
Leu Thr Lys Met Gln Ile Met Val Asn Arg Tyr 195 200 205Pro Thr Trp
Gly Asp Thr Val Glu Ile Asn Thr Trp Phe Ser Gln Ser 210 215 220Gly
Lys Ile Gly Met Gly Ser Asp Trp Leu Ile Ser Asp Cys Asn Thr225 230
235 240Gly Glu Ile Leu Ile Arg Ala Thr Ser Val Trp Ala Met Met Asn
Gln 245 250 255Lys Thr Arg Arg Phe Ser Arg Leu Pro Tyr Glu Val Arg
Gln Glu Leu 260 265 270Thr Pro His Phe Val Asp Ser Pro His Val Ile
Glu Asp Asn Asp Arg 275 280 285Lys Leu His Lys Phe Asp Val Lys Thr
Gly Asp Ser Ile Arg Lys Gly 290 295 300Leu Thr Pro Arg Trp Asn Asp
Leu Asp Val Asn Gln His Val Ser Asn305 310 315 320Val Lys Tyr Ile
Gly Trp Ile Leu Glu Ser Met Pro Ile Glu Val Leu 325 330 335Glu Thr
Gln Glu Leu Cys Ser Leu Thr Val Glu Tyr Arg Arg Glu Cys 340 345
350Gly Met Asp Ser Lys Leu Glu Ser Val Thr Ala Met Asp Pro Ser Glu
355 360 365Glu Asp Gly Val Arg Ser Gln Tyr Asn His Leu Leu Arg Leu
Glu Asp 370 375 380Gly Thr Asp Val Val Lys Gly Arg Thr Glu Trp Arg
Pro Lys Asn Ala385 390 395 400Gly Thr Asn Gly Ala Ile Ser Thr Gly
Lys Thr Ser Asn Gly Asn Ser 405 410 415Val Ser2415PRTCuphea
koehneana 2Met Val Thr Ala Ala Ala Ser Ser Ala Phe Phe Pro Val Pro
Ala Pro1 5 10 15Gly Thr Ser Pro Lys Pro Gly Lys Ser Trp Pro Ser Ser
Leu Ser Pro 20 25 30Ser Phe Lys Pro Lys Ser Ile Pro Asn Ala Gly Phe
Gln Val Lys Ala 35 40 45Asn Ala Ser Ala His Pro Lys Ala Asn Gly Ser
Ala Val Asn Leu Lys 50 55 60Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr
Ser Ser Ser Pro Pro Pro65 70 75 80Arg Ala Phe Leu Asn Gln Leu Pro
Asp Trp Ser Met Leu Leu Thr Ala 85 90 95Ile Thr Thr Val Phe Val Ala
Ala Glu Lys Gln Trp Thr Met Arg Asp 100 105 110Arg Lys Ser Lys Arg
Pro Asp Met Leu Val Asp Ser Val Gly Ser Lys 115 120 125Ser Ile Val
Leu Asp Gly Leu Val Ser Arg Gln Ile Phe Ser Ile Arg 130 135 140Ser
Tyr Glu Ile Gly Ala Asp Arg Thr Ala Ser Ile Glu Thr Leu Met145 150
155 160Asn His Leu Gln Glu Thr Ser Ile Asn His Cys Lys Ser Leu Gly
Leu 165 170 175Leu Asn Asp Gly Phe Gly Arg Thr Pro Gly Met Cys Lys
Asn Asp Leu 180 185 190Ile Trp Val Leu Thr Lys Met Gln Ile Met Val
Asn Arg Tyr Pro Thr 195 200 205Trp Gly Asp Thr Val Glu Ile Asn Thr
Trp Phe Ser His Ser Gly Lys 210 215 220Ile Gly Met Ala Ser Asp Trp
Leu Ile Thr Asp Cys Asn Thr Gly Glu225 230 235 240Ile Leu Ile Arg
Ala Thr Ser Val Trp Ala Met Met Asn Gln Lys Thr 245 250 255Arg Arg
Phe Ser Arg Leu Pro Tyr Glu Val Arg Gln Glu Leu Thr Pro 260 265
270His Tyr Val Asp Ser Pro His Val Ile Glu Asp Asn Asp Arg Lys Leu
275 280 285His Lys Phe Asp Val Lys Thr Gly Asp Ser Ile Arg Lys Gly
Leu Thr 290 295 300Pro Lys Trp Asn Asp Leu Asp Val Asn Gln His Val
Asn Asn Val Lys305 310 315 320Tyr Ile Gly Trp Ile Leu Glu Ser Met
Pro Ile Glu Val Leu Glu Thr 325 330 335Gln Glu Leu Cys Ser Leu Thr
Val Glu Tyr Arg Arg Glu Cys Gly Met 340 345 350Asp Ser Val Leu Glu
Ser Val Thr Ala Met Asp Pro Ser Glu Asp Gly 355 360 365Gly Leu Ser
Gln Tyr Lys His Leu Leu Arg Leu Glu Asp Gly Thr Asp 370 375 380Ile
Val Lys Gly Arg Thr Glu Trp Arg Pro Lys Asn Ala Gly Thr Asn385 390
395 400Gly Ala Ile Ser Thr Ala Lys Pro Ser Asn Gly Asn Ser Val Ser
405 410 4153417PRTCuphea leptopoda 3Met Val Gly Ala Ala Ala Ser Ser
Ala Phe Phe Pro Ala Pro Ala Pro1 5 10 15Gly Thr Ser Pro Lys Pro Gly
Lys Ser Gly Asn Trp Pro Ser Ser Leu 20 25 30Ser Pro Ser Leu Lys Pro
Lys Ser Ile Pro Asn Gly Gly Phe Gln Val 35 40 45Lys Ala Asn Ala Ser
Ala His Pro Lys Ala Asn Gly Ala Ala Val Asn 50 55 60Leu Lys Ser Gly
Ser Leu Asn Thr Gln Glu Asp Thr Ser Ser Ser Pro65 70 75 80Pro Pro
Arg Ala Phe Leu Asn Gln Leu Pro Asp Trp Ser Met Leu Leu 85 90 95Thr
Ala Ile Thr Thr Val Phe Val Ala Ala Glu Lys Gln Trp Thr Met 100 105
110Leu Asp Arg Lys Ser Lys Arg Pro Asp Met Leu Val Asp Ser Val Gly
115 120 125Leu Lys Asn Ile Val Arg Asp Gly Leu Val Ser Arg Gln Ser
Phe Ser 130 135 140Ile Arg Ser Tyr Glu Ile Gly Ala Asp Arg Thr Ala
Ser Ile Glu Thr145 150 155 160Leu Met Asn His Leu Gln Glu Thr Ser
Ile Asn His Cys Lys Ser Leu 165 170 175Gly Leu Leu Asn Asp Gly Phe
Gly Arg Thr Pro Gly Met Cys Lys Asn 180 185 190Asp Leu Ile Trp Val
Leu Thr Lys Met Gln Ile Leu Val Asn Arg Tyr 195 200 205Pro Ala Trp
Gly Asp Thr Val Glu Ile Asn Thr Trp Phe Ser Gln Ser 210 215 220Gly
Lys Ile Gly Met Gly Ser Asp Trp Leu Ile Ser Asp Cys Asn Thr225 230
235 240Gly Glu Ile Leu Ile Arg Ala Thr Ser Val Trp Ala Met Met Asn
Gln 245 250 255Lys Thr Arg Arg Phe Ser Arg Leu Pro Tyr Glu Val Arg
Gln Glu Leu 260 265 270Thr Pro His Phe Val Asp Ser Pro His Val Ile
Glu Asp Asn Asp Arg 275 280 285Lys Leu His Lys Phe Asp Val Lys Thr
Gly Asp Ser Ile Arg Lys Gly 290 295 300Leu Thr Pro Arg Trp Asn Asp
Leu Asp Val Asn Gln His Val Ser Asn305 310 315 320Val Lys Tyr Ile
Gly Trp Ile Leu Glu Ser Met Pro Ile Glu Val Leu 325 330 335Glu Thr
Gln Glu Leu Cys Ser Leu Thr Val Glu Tyr Arg Arg Glu Cys 340 345
350Gly Met Asp Ser Val Leu Glu Ser Val Thr Ala Arg Asp Pro Ser Glu
355 360 365Asp Gly Gly Arg Ser Gln Tyr Asn His Leu Leu Arg Leu Glu
Asp Gly 370 375 380Thr Asp Val Val Lys Gly Arg Thr Glu Trp Arg Ser
Lys Asn Ala Gly385 390 395 400Thr Asn Gly Ala Thr Ser Thr Ala Lys
Thr Ser Asn Gly Asn Ser Val 405 410 415Ser4417PRTCuphea
angustifolia 4Met Val Ala Ala Ala Ala Ser Ser Ala Phe Phe Pro Val
Pro Ala Pro1 5 10 15Gly Thr Ser Leu Lys Pro Gly Lys Ser Gly Asn Trp
Pro Ser Ser Leu 20 25 30Ser Pro Ser Phe Lys Pro Lys Thr Ile Pro Ser
Gly Gly Leu Gln Val 35 40 45Lys Ala Asn Ala Ser Ala His Pro Lys Ala
Asn Gly Ser Ala Val Asn 50 55 60Leu Lys Ser Gly Ser Leu Asp Thr Gln
Glu Asp Thr Ser Ser Ser Pro65 70 75 80Pro Pro Arg Ala Phe Leu Asn
Gln Leu Pro Asp Trp Ser Met Leu Leu 85 90 95Thr Ala Ile Thr Thr Val
Phe Val Ala Ala Glu Lys Gln Trp Thr Met 100 105 110Leu Asp Arg Lys
Ser Lys Arg Pro Glu Met Leu Val Asp Ser Val Gly 115 120 125Leu Lys
Ser Ser Val Arg Asp Gly Leu Val Ser Arg Gln Ser Phe Ser 130 135
140Ile Arg Ser Tyr Glu Ile Gly Ala Asp Arg Thr Ala Ser Ile Glu
Thr145 150 155 160Leu Met Asn His Leu Gln Glu Thr Ser Ile Asn His
Cys Lys Ser Leu 165 170 175Gly Leu Leu Asn Asp Gly Phe Gly Arg Thr
Pro Gly Met Cys Lys Asn 180 185 190Asp Leu Ile Trp Val Leu Thr Lys
Met Gln Ile Met Val Asn Arg Tyr 195 200 205Pro Thr Trp Gly Asp Thr
Val Glu Val Asn Thr Trp Phe Ser Gln Ser 210 215 220Gly Lys Ile Gly
Met Ala Ser Asp Trp Leu Ile Ser Asp Cys Asn Thr225 230 235 240Gly
Glu Ile Leu Ile Arg Ala Thr Ser Val Trp Ala Met Met Asn Gln 245 250
255Lys Thr Arg Arg Phe Ser Arg Leu Pro Tyr Glu Val Arg Gln Glu Leu
260 265 270Thr Pro His Tyr Val Asp Ser Pro His Val Ile Glu Asp Asn
Asp Arg 275 280 285Lys Leu His Lys Phe Asp Val Lys Thr Gly Asp Ser
Ile Arg Lys Gly 290 295 300Leu Thr Pro Arg Trp Asn Asp Leu Asp Val
Asn Gln His Val Ser Asn305 310 315 320Val Lys Tyr Ile Gly Trp Ile
Leu Glu Ser Met Pro Ile Glu Val Leu 325 330 335Glu Thr Gln Glu Leu
Cys Ser Leu Thr Val Glu Tyr Arg Arg Glu Cys 340 345 350Gly Met Asp
Ser Val Leu Glu Ser Val Thr Ala Met Asp Pro Ser Glu 355 360 365Asp
Gly Gly Val Ser Gln Tyr Lys His Leu Leu Arg Leu Glu Asp Gly 370 375
380Thr Asp Ile Val Lys Gly Arg Thr Glu Trp Arg Pro Lys Asn Ala
Gly385 390 395 400Thr Asn Gly Ala Thr Ser Lys Ala Lys Thr Ser Asn
Gly Asn Ser Val 405 410 415Ser5417PRTCuphea llavea 5Met Val Ala Ala
Ala Ala Ser Ser Ala Phe Phe Pro Ala Pro Ala Pro1 5 10 15Gly Ser Ser
Pro Lys Pro Gly Lys Pro Gly Asn Trp Pro Ser Ser Leu 20 25 30Ser Pro
Ser Phe Lys Pro Lys Ser Ile Pro Asn Gly Arg Phe Gln Val 35 40 45Lys
Ala Asn Ala Ser Ala His Pro Lys Ala Asn Gly Ser Ala Val Asn 50 55
60Leu Lys Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr Ser Ser Ser Pro65
70 75 80Pro Pro Arg Ala Phe Leu Asn Gln Leu Pro Asp Trp Ser Met Leu
Leu 85 90 95Ser Ala Ile Thr Thr Val Phe Val Ala Ala Glu Lys Gln Trp
Thr Met 100 105 110Leu Asp Arg Lys Ser Lys Arg Pro Asp Met Leu Val
Asp Ser Val Gly 115 120 125Leu Lys Asn Ile Val Arg Asp Gly Leu Val
Ser Arg Gln Ser Phe Ser 130 135 140Ile Arg Ser Tyr Glu Ile Gly Ala
Asp Arg Thr Ala Ser Ile Glu Thr145 150 155 160Leu Met Asn His Leu
Gln Glu Thr Ser Ile Asn His Cys Lys Ser Leu 165 170 175Gly Leu Leu
Asn Asp Gly Phe Gly Arg Thr Pro Gly Met Cys Lys Asn 180 185 190Asp
Leu Ile Trp Val Leu Thr Lys Met Gln Ile Met Val Asn Arg Tyr 195 200
205Pro Ala Trp Gly Asp Thr Val Glu Ile Asn Thr Trp Phe Ser Gln Ser
210 215 220Gly Lys Ile Gly Met Gly Ser Asp Trp Leu Ile Ser Asp Cys
Asn Thr225 230 235 240Gly Glu Ile Leu Ile Arg Ala Thr Ser Val Trp
Ala Met Met Asn Gln 245 250 255Lys Thr Arg Arg Phe Ser Arg Leu Pro
Tyr Glu Val Arg Gln Glu Leu 260 265 270Thr Pro His Phe Val Asp Ser
Pro His Val Ile Glu Asp Asn Asp Arg 275 280 285Lys Leu His Lys Phe
Asp Val Lys Thr Gly Asp Ser Ile Arg Lys Gly 290 295 300Leu Thr Pro
Arg Trp Asn Asp Leu Asp Val Asn Gln His Val Ser Asn305 310 315
320Val Lys Tyr Ile Gly Trp Ile Leu Glu Ser Met Pro Ile Glu Val Leu
325 330 335Glu Thr Gln Glu Leu Cys Ser Leu Thr Val Glu Tyr Arg Arg
Glu Cys 340 345 350Gly Met Asp Ser Val Leu Glu Ser Val Thr Ala Ile
Asp Pro Ser Glu 355 360 365Asp Gly Gly Arg Ser Gln Tyr Asn His Leu
Leu Arg Leu Asp Asp Gly 370 375 380Thr Asp Val Val Lys Gly Arg Thr
Glu Trp Arg Pro Lys Asn Ala Gly385 390 395 400Thr Asn Gly Ala Ile
Ser Thr Gly Lys Thr Ser Asn Gly Asn Ser Val 405 410
415Ser6417PRTCuphea lophostoma 6Met Val Ala Ala Ala Ala Ser Ser Ala
Phe Phe Pro Val Pro Ala Pro1 5 10 15Gly Thr Ser Leu Lys Pro Trp Lys
Ser Gly Asn Trp Pro Ser Ser Leu 20 25 30Ser Pro Ser Phe Lys Pro Lys
Thr Ile Pro Ser Gly Gly Phe Gln Val 35 40 45Lys Ala Asn Ala Ser Ala
Gln Pro Lys Ala Asn Gly Ser Ala Val Asn 50 55 60Leu Lys Ser Gly Ser
Leu Asn Thr Gln Glu Asp Thr Thr Ser Ser Pro65 70 75 80Pro Pro Arg
Ala Phe Leu Asn Gln Leu Pro Asp Trp Ser Met Leu Leu 85 90 95Thr Ala
Ile Thr Thr Val Phe Val Ala Ala Glu Lys Gln Trp Thr Met 100 105
110Leu Asp Arg Lys Ser Lys Arg Pro Glu Lys Leu Val Asp Ser Val Gly
115 120 125Leu Lys Ser Ser Val Arg Asp Gly Leu Val Ser Arg Gln Ser
Phe Ser 130 135 140Ile Arg Ser Tyr Glu Ile Gly Ala Asp Arg Thr Ala
Ser Ile Glu Thr145 150 155 160Leu Met Asn His Leu Gln Glu Thr Ser
Ile Asn His Cys Lys Ser Leu 165 170 175Gly Leu Leu Asn Asp Gly Phe
Gly Arg Thr Pro Gly Met Cys Lys Asn 180 185 190Asp Leu Ile Trp Val
Leu Thr Lys Met Gln Ile Met Val Asn Arg Tyr 195 200 205Pro Thr Trp
Gly Asp Thr Val Glu Ile Asn Thr Trp Phe Ser Gln Ser 210 215 220Gly
Lys Ile Gly Met Ala Ser Asp Trp Leu Ile Ser Asp Cys Asn Thr225 230
235 240Gly Glu Ile Leu Ile Arg Ala Thr Ser Val Trp Ala Met Met Asn
Gln 245 250 255Lys Thr Arg Arg Phe Ser Arg Leu Pro Tyr Glu Val Arg
Gln Glu Leu 260 265 270Thr Pro His Tyr Val Asp Ser Pro His Val Ile
Glu Asp Asn Asp Arg 275 280 285Lys Leu His Lys Phe Asp Val Lys Thr
Gly Asp Ser Ile Arg Lys Gly 290 295 300Leu Thr Pro Arg Trp Asn Asp
Leu Asp Val Asn Gln His Val Ser Asn305 310 315 320Val Lys Tyr Ile
Gly Trp Ile Leu Glu Ser Met Pro Ile Glu Val Leu 325 330 335Glu Thr
Gln Glu Leu Cys Ser Leu Thr Val Glu Tyr Arg Arg Glu Cys 340 345
350Gly Met Asp Ser Val Leu Glu Ser Val Thr Ala Met Asp Pro Ser Glu
355 360 365Asp Glu Gly Arg Ser Gln Tyr Lys His
Leu Leu Arg Leu Glu Asp Gly 370 375 380Thr Asp Ile Val Lys Gly Arg
Thr Glu Trp Arg Pro Lys Asn Ala Gly385 390 395 400Thr Asn Gly Ala
Ile Ser Thr Ala Lys Asn Ser Asn Gly Asn Ser Val 405 410
415Ser7384PRTSassafras albidum 7Met Ala Thr Thr Ser Leu Ala Ser Ala
Phe Cys Ser Met Lys Ala Val1 5 10 15Met Leu Ala Arg Asp Gly Arg Gly
Met Lys Pro Arg Ser Ser Asp Leu 20 25 30Gln Leu Arg Ala Gly Asn Ala
Gln Thr Pro Leu Lys Met Ile Asn Gly 35 40 45Thr Lys Phe Ser Tyr Thr
Glu Ser Leu Lys Arg Leu Pro Asp Trp Ser 50 55 60Met Leu Phe Ala Val
Ile Thr Thr Ile Phe Ser Val Ala Glu Lys Gln65 70 75 80Trp Thr Asn
Leu Glu Trp Lys Pro Lys Pro Lys Pro Arg Leu Pro Gln 85 90 95Leu Leu
Asp Asp His Phe Gly Leu His Gly Leu Val Phe Arg Arg Thr 100 105
110Phe Ala Ile Arg Ser Tyr Glu Val Gly Pro Asp Arg Ser Thr Ser Ile
115 120 125Val Ala Val Met Asn His Leu Gln Glu Ala Thr Leu Asn His
Ala Lys 130 135 140Ser Val Gly Ile Leu Gly Asp Gly Phe Gly Thr Thr
Leu Glu Met Ser145 150 155 160Lys Arg Asp Leu Ala Trp Val Val Arg
Arg Thr His Val Ala Val Glu 165 170 175Arg Tyr Pro Ala Trp Gly Asp
Thr Val Glu Val Glu Cys Trp Ile Gly 180 185 190Ala Ser Gly Asn Asn
Gly Met Arg Arg Asp Phe Leu Val Arg Asp Cys 195 200 205Lys Thr Gly
Glu Ile Leu Thr Arg Cys Thr Ser Leu Ser Val Met Met 210 215 220Asn
Thr Arg Thr Arg Arg Leu Ser Lys Ile Pro Glu Glu Val Arg Gly225 230
235 240Glu Ile Gly Pro Leu Phe Ile Asp Asn Val Ala Val Lys Asp Glu
Glu 245 250 255Ile Lys Lys Leu Gln Lys Leu Asn Asp Ser Ser Ala Asp
Tyr Ile Gln 260 265 270Gly Gly Leu Thr Pro Arg Trp Asn Asp Leu Asp
Val Asn Gln His Val 275 280 285Asn Asn Ile Lys Tyr Val Gly Trp Ile
Leu Glu Thr Val Pro Asp Ser 290 295 300Ile Phe Glu Ser His His Ile
Ser Ser Ile Thr Leu Glu Tyr Arg Arg305 310 315 320Glu Cys Thr Arg
Asp Ser Val Leu Gln Ser Leu Thr Thr Val Ser Gly 325 330 335Gly Ser
Leu Glu Ala Gly Leu Val Cys Asp His Leu Leu Gln Leu Glu 340 345
350Gly Gly Ser Glu Val Leu Arg Ala Arg Thr Glu Trp Arg Pro Lys Leu
355 360 365Thr Asp Ser Phe Arg Gly Ile Ile Val Ile Pro Ala Glu Pro
Ser Val 370 375 3808384PRTSassafras albidum 8Met Ala Thr Thr Ser
Leu Ala Ser Ala Phe Cys Ser Met Lys Ala Val1 5 10 15Met Leu Ala Arg
Asp Gly Arg Gly Met Lys Pro Arg Ser Ser Asp Leu 20 25 30Gln Leu Arg
Ala Gly Asn Ala Gln Thr Pro Leu Lys Met Ile Asn Gly 35 40 45Thr Lys
Phe Ser Tyr Thr Glu Ser Leu Lys Arg Leu Pro Asp Trp Ser 50 55 60Met
Leu Phe Ala Val Ile Thr Thr Ile Phe Ser Val Ala Glu Lys Gln65 70 75
80Trp Thr Asn Leu Glu Trp Lys Pro Lys Pro Lys Pro Arg Leu Pro Gln
85 90 95Leu Leu Asp Asp His Phe Gly Leu His Gly Leu Val Phe Arg Arg
Thr 100 105 110Phe Ala Ile Arg Ser Tyr Glu Val Gly Pro Asp Arg Ser
Thr Ser Ile 115 120 125Val Ala Val Met Asn His Leu Gln Glu Ala Thr
Leu Asn His Ala Lys 130 135 140Ser Val Gly Ile Leu Gly Asp Gly Phe
Gly Thr Thr Leu Glu Met Ser145 150 155 160Lys Arg Asp Leu Ala Trp
Val Val Arg Arg Thr His Val Ala Val Glu 165 170 175Arg Tyr Pro Ala
Trp Gly Asp Thr Val Glu Val Glu Ala Trp Val Gly 180 185 190Ala Ser
Gly Asn Ile Gly Met Arg Arg Asp Phe Leu Val Arg Asp Cys 195 200
205Lys Thr Gly His Ile Leu Ala Arg Cys Thr Ser Val Ser Val Met Met
210 215 220Asn Ala Arg Thr Arg Arg Leu Ser Lys Ile Pro Gln Glu Val
Arg Ala225 230 235 240Glu Ile Asp Pro Leu Phe Ile Glu Lys Val Ala
Val Lys Glu Gly Glu 245 250 255Ile Lys Lys Leu Gln Lys Phe Asn Asp
Ser Thr Ala Asp Tyr Ile Gln 260 265 270Gly Gly Trp Thr Pro Arg Trp
Asn Asp Leu Asp Val Asn Gln His Val 275 280 285Asn Asn Ile Lys Tyr
Ile Gly Trp Ile Phe Lys Ser Val Pro Asp Ser 290 295 300Ile Ser Glu
Asn His Tyr Leu Ser Ser Ile Thr Leu Glu Tyr Arg Arg305 310 315
320Glu Cys Thr Arg Gly Ser Ala Leu Gln Ser Leu Thr Thr Val Cys Gly
325 330 335Asp Ser Ser Glu Ala Gly Ile Ile Cys Glu His Leu Leu Gln
Leu Glu 340 345 350Asp Gly Pro Glu Val Leu Arg Ala Arg Thr Glu Trp
Arg Pro Lys Leu 355 360 365Thr Asp Ser Phe Arg Gly Ile Ile Val Ile
Pro Ala Glu Pro Ser Val 370 375 3809380PRTLindera benzoin 9Met Val
Ala Thr Ser Leu Ala Ser Ala Phe Cys Ser Met Lys Ala Val1 5 10 15Met
Leu Ala Asp Asp Gly Arg Gly Met Lys Pro Arg Ser Ser Asp Leu 20 25
30Gln Leu Arg Ala Gly Asn Ala Gln Thr Ser Leu Lys Met Ile Asp Gly
35 40 45Thr Lys Phe Ser Tyr Thr Glu Ser Leu Lys Arg Leu Pro Asp Trp
Ser 50 55 60Lys Leu Leu Thr Val Ile Thr Thr Ile Phe Ser Ala Ala Glu
Lys Gln65 70 75 80Trp Thr Asn Leu Glu Arg Lys Pro Lys Pro Pro His
Leu Leu Asp Asp 85 90 95Arg Phe Gly Leu His Gly Leu Val Phe Arg Arg
Thr Phe Ala Ile Arg 100 105 110Ser Tyr Glu Val Gly Pro Asp Arg Ser
Ala Ser Ile Leu Ala Val Leu 115 120 125Asn His Leu Gln Glu Ala Thr
Leu Asn His Ala Glu Ser Val Gly Ile 130 135 140Leu Gly Asp Arg Phe
Gly Glu Thr Leu Glu Met Ser Lys Arg Asp Leu145 150 155 160Met Trp
Val Val Arg Arg Thr Tyr Val Ala Val Glu Arg Tyr Pro Ala 165 170
175Trp Gly Asp Thr Val Glu Ile Glu Ser Trp Ile Gly Ala Ser Gly Asn
180 185 190Asn Gly Met Arg Arg Glu Phe Leu Val Arg Asp Phe Lys Thr
Gly Glu 195 200 205Ile Leu Thr Arg Cys Thr Ser Leu Ser Val Met Met
Asn Thr Arg Thr 210 215 220Arg Arg Leu Ser Lys Ile Pro Glu Glu Val
Arg Gly Glu Ile Gly Pro225 230 235 240Val Phe Ile Asp Asn Val Ala
Val Lys Asp Glu Glu Ile Lys Lys Leu 245 250 255Gln Lys Leu Asn Asp
Ser Thr Ala Asp Tyr Ile Gln Gly Gly Leu Ile 260 265 270Pro Arg Trp
Asn Asp Leu Asp Leu Asn Gln His Val Asn Asn Ile Lys 275 280 285Tyr
Val Ser Trp Ile Leu Glu Thr Val Pro Asp Ser Ile Leu Glu Ser 290 295
300Tyr His Met Ser Ser Ile Thr Leu Glu Tyr Arg Arg Glu Cys Thr
Arg305 310 315 320Asp Ser Val Leu Gln Ser Leu Thr Thr Val Ser Gly
Gly Ser Ser Glu 325 330 335Ala Gly Leu Val Cys Glu His Ser Leu Leu
Leu Glu Gly Gly Ser Glu 340 345 350Val Leu Arg Ala Arg Thr Glu Trp
Arg Pro Lys Leu Thr Asp Ser Phe 355 360 365Arg Gly Ile Ser Val Ile
Pro Ala Glu Gln Ser Val 370 375 38010418PRTCuphea crassiflora 10Met
Val Ala Ala Ala Ala Ser Ser Ala Phe Phe Pro Val Pro Ala Pro1 5 10
15Gly Thr Ser Thr Lys Pro Arg Lys Ser Gly Asn Trp Pro Ser Arg Leu
20 25 30Ser Pro Ser Ser Lys Pro Lys Ser Ile Pro Asn Gly Gly Phe Gln
Val 35 40 45Lys Ala Asn Ala Ser Ala His Pro Lys Ala Asn Gly Ser Ala
Val Asn 50 55 60Leu Lys Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr Ser
Ser Ser Pro65 70 75 80Pro Pro Arg Ala Phe Leu Asn Gln Leu Pro Asp
Trp Ser Met Leu Leu 85 90 95Thr Ala Ile Thr Thr Val Phe Val Ala Ala
Glu Lys Gln Trp Thr Met 100 105 110Leu Asp Arg Lys Ser Lys Arg Pro
Asp Met Leu Val Asp Ser Val Gly 115 120 125Leu Lys Ser Ile Val Arg
Asp Gly Leu Val Ser Arg Gln Ser Phe Ser 130 135 140Ile Arg Ser Tyr
Glu Ile Gly Ala Asp Arg Thr Ala Ser Ile Glu Thr145 150 155 160Leu
Met Asn His Leu Gln Glu Thr Ser Ile Asn His Cys Lys Ser Leu 165 170
175Gly Leu Leu Asn Asp Gly Phe Gly Arg Thr Pro Gly Met Cys Lys Asn
180 185 190Asp Leu Ile Trp Val Leu Thr Lys Met Gln Ile Met Val Asn
Arg Tyr 195 200 205Pro Thr Trp Gly Asp Thr Val Glu Ile Asn Thr Trp
Phe Ser Gln Ser 210 215 220Gly Lys Ile Gly Met Gly Ser Asp Trp Leu
Ile Ser Asp Cys Asn Thr225 230 235 240Gly Glu Ile Leu Ile Arg Ala
Thr Ser Val Trp Ala Met Met Asn Gln 245 250 255Lys Thr Arg Arg Phe
Ser Arg Leu Pro Tyr Glu Val Arg Gln Glu Leu 260 265 270Thr Pro His
Phe Val Asp Ser Pro His Val Ile Glu Asp Asn Asp Arg 275 280 285Lys
Leu His Lys Phe Asp Val Lys Thr Gly Asp Ser Ile Arg Lys Gly 290 295
300Leu Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln His Val Ser
Asn305 310 315 320Val Lys Tyr Ile Gly Trp Ile Leu Glu Ser Met Pro
Ile Glu Val Leu 325 330 335Glu Thr Gln Glu Leu Cys Ser Leu Thr Val
Glu Tyr Arg Arg Glu Cys 340 345 350Gly Met Asp Ser Lys Leu Glu Ser
Val Thr Ala Met Asp Pro Ser Glu 355 360 365Glu Asp Gly Val Arg Ser
Gln Tyr Asn His Leu Leu Arg Leu Glu Asp 370 375 380Gly Thr Asp Val
Val Lys Gly Arg Thr Glu Trp Arg Pro Lys Asn Ala385 390 395 400Gly
Thr Asn Gly Ala Ile Ser Thr Gly Lys Thr Ser Asn Gly Asn Ser 405 410
415Val Ser11415PRTCuphea koehneana 11Met Val Thr Ala Ala Ala Ser
Ser Ala Phe Phe Pro Val Pro Ala Pro1 5 10 15Gly Thr Ser Pro Lys Pro
Gly Lys Ser Trp Pro Ser Ser Leu Ser Pro 20 25 30Ser Phe Lys Pro Lys
Ser Ile Pro Asn Ala Gly Phe Gln Val Lys Ala 35 40 45Asn Ala Ser Ala
His Pro Lys Ala Asn Gly Ser Ala Val Asn Leu Lys 50 55 60Ser Gly Ser
Leu Asn Thr Gln Glu Asp Thr Ser Ser Ser Pro Pro Pro65 70 75 80Arg
Ala Phe Leu Asn Gln Leu Pro Asp Trp Ser Met Leu Leu Thr Ala 85 90
95Ile Thr Thr Val Phe Val Ala Ala Glu Lys Gln Trp Thr Met Arg Asp
100 105 110Arg Lys Ser Lys Arg Pro Asp Met Leu Val Asp Ser Val Gly
Ser Lys 115 120 125Ser Ile Val Leu Asp Gly Leu Val Ser Arg Gln Ile
Phe Ser Ile Arg 130 135 140Ser Tyr Glu Ile Gly Ala Asp Arg Thr Ala
Ser Ile Glu Thr Leu Met145 150 155 160Asn His Leu Gln Glu Thr Ser
Ile Asn His Cys Lys Ser Leu Gly Leu 165 170 175Leu Asn Asp Gly Phe
Gly Arg Thr Pro Gly Met Cys Lys Asn Asp Leu 180 185 190Ile Trp Val
Leu Thr Lys Met Gln Ile Met Val Asn Arg Tyr Pro Thr 195 200 205Trp
Gly Asp Thr Val Glu Ile Asn Thr Trp Phe Ser His Ser Gly Lys 210 215
220Ile Gly Met Ala Ser Asp Trp Leu Ile Thr Asp Cys Asn Thr Gly
Glu225 230 235 240Ile Leu Ile Arg Ala Thr Ser Val Trp Ala Met Met
Asn Gln Lys Thr 245 250 255Arg Arg Phe Ser Arg Leu Pro Tyr Glu Val
Arg Gln Glu Leu Thr Pro 260 265 270His Tyr Val Asp Ser Pro His Val
Ile Glu Asp Asn Asp Arg Lys Leu 275 280 285His Lys Phe Asp Val Lys
Thr Gly Asp Ser Ile Arg Lys Gly Leu Thr 290 295 300Pro Lys Trp Asn
Asp Leu Asp Val Asn Gln His Val Asn Asn Val Lys305 310 315 320Tyr
Ile Gly Trp Ile Leu Glu Ser Met Pro Ile Glu Val Leu Glu Thr 325 330
335Gln Glu Leu Cys Ser Leu Thr Val Glu Tyr Arg Arg Glu Cys Gly Met
340 345 350Asp Ser Val Leu Glu Ser Val Thr Ala Met Asp Pro Ser Glu
Asp Gly 355 360 365Gly Leu Ser Gln Tyr Lys His Leu Leu Arg Leu Glu
Asp Gly Thr Asp 370 375 380Ile Val Lys Gly Arg Thr Glu Trp Arg Pro
Lys Asn Ala Gly Thr Asn385 390 395 400Gly Ala Ile Ser Thr Ala Lys
Pro Ser Asn Gly Asn Ser Val Ser 405 410 41512417PRTCuphea leptopoda
12Met Val Gly Ala Ala Ala Ser Ser Ala Phe Phe Pro Ala Pro Ala Pro1
5 10 15Gly Thr Ser Pro Lys Pro Gly Lys Ser Gly Asn Trp Pro Ser Ser
Leu 20 25 30Ser Pro Ser Leu Lys Pro Lys Ser Ile Pro Asn Gly Gly Phe
Gln Val 35 40 45Lys Ala Asn Ala Ser Ala His Pro Lys Ala Asn Gly Ala
Ala Val Asn 50 55 60Leu Lys Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr
Ser Ser Ser Pro65 70 75 80Pro Pro Arg Ala Phe Leu Asn Gln Leu Pro
Asp Trp Ser Met Leu Leu 85 90 95Thr Ala Ile Thr Thr Val Phe Val Ala
Ala Glu Lys Gln Trp Thr Met 100 105 110Leu Asp Arg Lys Ser Lys Arg
Pro Asp Met Leu Val Asp Ser Val Gly 115 120 125Leu Lys Asn Ile Val
Arg Asp Gly Leu Val Ser Arg Gln Ser Phe Ser 130 135 140Ile Arg Ser
Tyr Glu Ile Gly Ala Asp Arg Thr Ala Ser Ile Glu Thr145 150 155
160Leu Met Asn His Leu Gln Glu Thr Ser Ile Asn His Cys Lys Ser Leu
165 170 175Gly Leu Leu Asn Asp Gly Phe Gly Arg Thr Pro Gly Met Cys
Lys Asn 180 185 190Asp Leu Ile Trp Val Leu Thr Lys Met Gln Ile Leu
Val Asn Arg Tyr 195 200 205Pro Ala Trp Gly Asp Thr Val Glu Ile Asn
Thr Trp Phe Ser Gln Ser 210 215 220Gly Lys Ile Gly Met Gly Ser Asp
Trp Leu Ile Ser Asp Cys Asn Thr225 230 235 240Gly Glu Ile Leu Ile
Arg Ala Thr Ser Val Trp Ala Met Met Asn Gln 245 250 255Lys Thr Arg
Arg Phe Ser Arg Leu Pro Tyr Glu Val Arg Gln Glu Leu 260 265 270Thr
Pro His Phe Val Asp Ser Pro His Val Ile Glu Asp Asn Asp Arg 275 280
285Lys Leu His Lys Phe Asp Val Lys Thr Gly Asp Ser Ile Arg Lys Gly
290 295 300Leu Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln His Val
Ser Asn305 310 315 320Val Lys Tyr Ile Gly Trp Ile Leu Glu Ser Met
Pro Ile Glu Val Leu 325 330 335Glu Thr Gln Glu Leu Cys Ser Leu Thr
Val Glu Tyr Arg Arg Glu Cys 340 345 350Gly Met Asp Ser Val Leu Glu
Ser Val Thr Ala Arg Asp Pro Ser Glu 355 360 365Asp Gly Gly Arg Ser
Gln Tyr Asn His Leu Leu Arg Leu Glu Asp Gly 370 375 380Thr Asp Val
Val Lys Gly Arg Thr Glu Trp Arg Ser Lys Asn Ala Gly385 390 395
400Thr Asn Gly Ala Thr Ser Thr Ala Lys Thr Ser Asn Gly Asn Ser Val
405 410 415Ser13417PRTCuphea angustifolia 13Met Val Ala Ala Ala Ala
Ser Ser Ala Phe Phe Pro Val Pro Ala Pro1 5 10
15Gly Thr Ser Leu Lys Pro Gly Lys Ser Gly Asn Trp Pro Ser Ser Leu
20 25 30Ser Pro Ser Phe Lys Pro Lys Thr Ile Pro Ser Gly Gly Leu Gln
Val 35 40 45Lys Ala Asn Ala Ser Ala His Pro Lys Ala Asn Gly Ser Ala
Val Asn 50 55 60Leu Lys Ser Gly Ser Leu Asp Thr Gln Glu Asp Thr Ser
Ser Ser Pro65 70 75 80Pro Pro Arg Ala Phe Leu Asn Gln Leu Pro Asp
Trp Ser Met Leu Leu 85 90 95Thr Ala Ile Thr Thr Val Phe Val Ala Ala
Glu Lys Gln Trp Thr Met 100 105 110Leu Asp Arg Lys Ser Lys Arg Pro
Glu Met Leu Val Asp Ser Val Gly 115 120 125Leu Lys Ser Ser Val Arg
Asp Gly Leu Val Ser Arg Gln Ser Phe Ser 130 135 140Ile Arg Ser Tyr
Glu Ile Gly Ala Asp Arg Thr Ala Ser Ile Glu Thr145 150 155 160Leu
Met Asn His Leu Gln Glu Thr Ser Ile Asn His Cys Lys Ser Leu 165 170
175Gly Leu Leu Asn Asp Gly Phe Gly Arg Thr Pro Gly Met Cys Lys Asn
180 185 190Asp Leu Ile Trp Val Leu Thr Lys Met Gln Ile Met Val Asn
Arg Tyr 195 200 205Pro Thr Trp Gly Asp Thr Val Glu Val Asn Thr Trp
Phe Ser Gln Ser 210 215 220Gly Lys Ile Gly Met Ala Ser Asp Trp Leu
Ile Ser Asp Cys Asn Thr225 230 235 240Gly Glu Ile Leu Ile Arg Ala
Thr Ser Val Trp Ala Met Met Asn Gln 245 250 255Lys Thr Arg Arg Phe
Ser Arg Leu Pro Tyr Glu Val Arg Gln Glu Leu 260 265 270Thr Pro His
Tyr Val Asp Ser Pro His Val Ile Glu Asp Asn Asp Arg 275 280 285Lys
Leu His Lys Phe Asp Val Lys Thr Gly Asp Ser Ile Arg Lys Gly 290 295
300Leu Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln His Val Ser
Asn305 310 315 320Val Lys Tyr Ile Gly Trp Ile Leu Glu Ser Met Pro
Ile Glu Val Leu 325 330 335Glu Thr Gln Glu Leu Cys Ser Leu Thr Val
Glu Tyr Arg Arg Glu Cys 340 345 350Gly Met Asp Ser Val Leu Glu Ser
Val Thr Ala Met Asp Pro Ser Glu 355 360 365Asp Gly Gly Val Ser Gln
Tyr Lys His Leu Leu Arg Leu Glu Asp Gly 370 375 380Thr Asp Ile Val
Lys Gly Arg Thr Glu Trp Arg Pro Lys Asn Ala Gly385 390 395 400Thr
Asn Gly Ala Thr Ser Lys Ala Lys Thr Ser Asn Gly Asn Ser Val 405 410
415Ser14417PRTCuphea llavea 14Met Val Ala Ala Ala Ala Ser Ser Ala
Phe Phe Pro Ala Pro Ala Pro1 5 10 15Gly Ser Ser Pro Lys Pro Gly Lys
Pro Gly Asn Trp Pro Ser Ser Leu 20 25 30Ser Pro Ser Phe Lys Pro Lys
Ser Ile Pro Asn Gly Arg Phe Gln Val 35 40 45Lys Ala Asn Ala Ser Ala
His Pro Lys Ala Asn Gly Ser Ala Val Asn 50 55 60Leu Lys Ser Gly Ser
Leu Asn Thr Gln Glu Asp Thr Ser Ser Ser Pro65 70 75 80Pro Pro Arg
Ala Phe Leu Asn Gln Leu Pro Asp Trp Ser Met Leu Leu 85 90 95Ser Ala
Ile Thr Thr Val Phe Val Ala Ala Glu Lys Gln Trp Thr Met 100 105
110Leu Asp Arg Lys Ser Lys Arg Pro Asp Met Leu Val Asp Ser Val Gly
115 120 125Leu Lys Asn Ile Val Arg Asp Gly Leu Val Ser Arg Gln Ser
Phe Ser 130 135 140Ile Arg Ser Tyr Glu Ile Gly Ala Asp Arg Thr Ala
Ser Ile Glu Thr145 150 155 160Leu Met Asn His Leu Gln Glu Thr Ser
Ile Asn His Cys Lys Ser Leu 165 170 175Gly Leu Leu Asn Asp Gly Phe
Gly Arg Thr Pro Gly Met Cys Lys Asn 180 185 190Asp Leu Ile Trp Val
Leu Thr Lys Met Gln Ile Met Val Asn Arg Tyr 195 200 205Pro Ala Trp
Gly Asp Thr Val Glu Ile Asn Thr Trp Phe Ser Gln Ser 210 215 220Gly
Lys Ile Gly Met Gly Ser Asp Trp Leu Ile Ser Asp Cys Asn Thr225 230
235 240Gly Glu Ile Leu Ile Arg Ala Thr Ser Val Trp Ala Met Met Asn
Gln 245 250 255Lys Thr Arg Arg Phe Ser Arg Leu Pro Tyr Glu Val Arg
Gln Glu Leu 260 265 270Thr Pro His Phe Val Asp Ser Pro His Val Ile
Glu Asp Asn Asp Arg 275 280 285Lys Leu His Lys Phe Asp Val Lys Thr
Gly Asp Ser Ile Arg Lys Gly 290 295 300Leu Thr Pro Arg Trp Asn Asp
Leu Asp Val Asn Gln His Val Ser Asn305 310 315 320Val Lys Tyr Ile
Gly Trp Ile Leu Glu Ser Met Pro Ile Glu Val Leu 325 330 335Glu Thr
Gln Glu Leu Cys Ser Leu Thr Val Glu Tyr Arg Arg Glu Cys 340 345
350Gly Met Asp Ser Val Leu Glu Ser Val Thr Ala Ile Asp Pro Ser Glu
355 360 365Asp Gly Gly Arg Ser Gln Tyr Asn His Leu Leu Arg Leu Asp
Asp Gly 370 375 380Thr Asp Val Val Lys Gly Arg Thr Glu Trp Arg Pro
Lys Asn Ala Gly385 390 395 400Thr Asn Gly Ala Ile Ser Thr Gly Lys
Thr Ser Asn Gly Asn Ser Val 405 410 415Ser15417PRTCuphea lophostoma
15Met Val Ala Ala Ala Ala Ser Ser Ala Phe Phe Pro Val Pro Ala Pro1
5 10 15Gly Thr Ser Leu Lys Pro Trp Lys Ser Gly Asn Trp Pro Ser Ser
Leu 20 25 30Ser Pro Ser Phe Lys Pro Lys Thr Ile Pro Ser Gly Gly Phe
Gln Val 35 40 45Lys Ala Asn Ala Ser Ala Gln Pro Lys Ala Asn Gly Ser
Ala Val Asn 50 55 60Leu Lys Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr
Thr Ser Ser Pro65 70 75 80Pro Pro Arg Ala Phe Leu Asn Gln Leu Pro
Asp Trp Ser Met Leu Leu 85 90 95Thr Ala Ile Thr Thr Val Phe Val Ala
Ala Glu Lys Gln Trp Thr Met 100 105 110Leu Asp Arg Lys Ser Lys Arg
Pro Glu Lys Leu Val Asp Ser Val Gly 115 120 125Leu Lys Ser Ser Val
Arg Asp Gly Leu Val Ser Arg Gln Ser Phe Ser 130 135 140Ile Arg Ser
Tyr Glu Ile Gly Ala Asp Arg Thr Ala Ser Ile Glu Thr145 150 155
160Leu Met Asn His Leu Gln Glu Thr Ser Ile Asn His Cys Lys Ser Leu
165 170 175Gly Leu Leu Asn Asp Gly Phe Gly Arg Thr Pro Gly Met Cys
Lys Asn 180 185 190Asp Leu Ile Trp Val Leu Thr Lys Met Gln Ile Met
Val Asn Arg Tyr 195 200 205Pro Thr Trp Gly Asp Thr Val Glu Ile Asn
Thr Trp Phe Ser Gln Ser 210 215 220Gly Lys Ile Gly Met Ala Ser Asp
Trp Leu Ile Ser Asp Cys Asn Thr225 230 235 240Gly Glu Ile Leu Ile
Arg Ala Thr Ser Val Trp Ala Met Met Asn Gln 245 250 255Lys Thr Arg
Arg Phe Ser Arg Leu Pro Tyr Glu Val Arg Gln Glu Leu 260 265 270Thr
Pro His Tyr Val Asp Ser Pro His Val Ile Glu Asp Asn Asp Arg 275 280
285Lys Leu His Lys Phe Asp Val Lys Thr Gly Asp Ser Ile Arg Lys Gly
290 295 300Leu Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln His Val
Ser Asn305 310 315 320Val Lys Tyr Ile Gly Trp Ile Leu Glu Ser Met
Pro Ile Glu Val Leu 325 330 335Glu Thr Gln Glu Leu Cys Ser Leu Thr
Val Glu Tyr Arg Arg Glu Cys 340 345 350Gly Met Asp Ser Val Leu Glu
Ser Val Thr Ala Met Asp Pro Ser Glu 355 360 365Asp Glu Gly Arg Ser
Gln Tyr Lys His Leu Leu Arg Leu Glu Asp Gly 370 375 380Thr Asp Ile
Val Lys Gly Arg Thr Glu Trp Arg Pro Lys Asn Ala Gly385 390 395
400Thr Asn Gly Ala Ile Ser Thr Ala Lys Asn Ser Asn Gly Asn Ser Val
405 410 415Ser16384PRTSassafras albidum 16Met Ala Thr Thr Ser Leu
Ala Ser Ala Phe Cys Ser Met Lys Ala Val1 5 10 15Met Leu Ala Arg Asp
Gly Arg Gly Met Lys Pro Arg Ser Ser Asp Leu 20 25 30Gln Leu Arg Ala
Gly Asn Ala Gln Thr Pro Leu Lys Met Ile Asn Gly 35 40 45Thr Lys Phe
Ser Tyr Thr Glu Ser Leu Lys Arg Leu Pro Asp Trp Ser 50 55 60Met Leu
Phe Ala Val Ile Thr Thr Ile Phe Ser Val Ala Glu Lys Gln65 70 75
80Trp Thr Asn Leu Glu Trp Lys Pro Lys Pro Lys Pro Arg Leu Pro Gln
85 90 95Leu Leu Asp Asp His Phe Gly Leu His Gly Leu Val Phe Arg Arg
Thr 100 105 110Phe Ala Ile Arg Ser Tyr Glu Val Gly Pro Asp Arg Ser
Thr Ser Ile 115 120 125Val Ala Val Met Asn His Leu Gln Glu Ala Thr
Leu Asn His Ala Lys 130 135 140Ser Val Gly Ile Leu Gly Asp Gly Phe
Gly Thr Thr Leu Glu Met Ser145 150 155 160Lys Arg Asp Leu Ala Trp
Val Val Arg Arg Thr His Val Ala Val Glu 165 170 175Arg Tyr Pro Ala
Trp Gly Asp Thr Val Glu Val Glu Cys Trp Ile Gly 180 185 190Ala Ser
Gly Asn Asn Gly Met Arg Arg Asp Phe Leu Val Arg Asp Cys 195 200
205Lys Thr Gly Glu Ile Leu Thr Arg Cys Thr Ser Leu Ser Val Met Met
210 215 220Asn Thr Arg Thr Arg Arg Leu Ser Lys Ile Pro Glu Glu Val
Arg Gly225 230 235 240Glu Ile Gly Pro Leu Phe Ile Asp Asn Val Ala
Val Lys Asp Glu Glu 245 250 255Ile Lys Lys Leu Gln Lys Leu Asn Asp
Ser Ser Ala Asp Tyr Ile Gln 260 265 270Gly Gly Leu Thr Pro Arg Trp
Asn Asp Leu Asp Val Asn Gln His Val 275 280 285Asn Asn Ile Lys Tyr
Val Gly Trp Ile Leu Glu Thr Val Pro Asp Ser 290 295 300Ile Phe Glu
Ser His His Ile Ser Ser Ile Thr Leu Glu Tyr Arg Arg305 310 315
320Glu Cys Thr Arg Asp Ser Val Leu Gln Ser Leu Thr Thr Val Ser Gly
325 330 335Gly Ser Leu Glu Ala Gly Leu Val Cys Asp His Leu Leu Gln
Leu Glu 340 345 350Gly Gly Ser Glu Val Leu Arg Ala Arg Thr Glu Trp
Arg Pro Lys Leu 355 360 365Thr Asp Ser Phe Arg Gly Ile Ile Val Ile
Pro Ala Glu Pro Ser Val 370 375 38017384PRTSassafras albidum 17Met
Ala Thr Thr Ser Leu Ala Ser Ala Phe Cys Ser Met Lys Ala Val1 5 10
15Met Leu Ala Arg Asp Gly Arg Gly Met Lys Pro Arg Ser Ser Asp Leu
20 25 30Gln Leu Arg Ala Gly Asn Ala Gln Thr Pro Leu Lys Met Ile Asn
Gly 35 40 45Thr Lys Phe Ser Tyr Thr Glu Ser Leu Lys Arg Leu Pro Asp
Trp Ser 50 55 60Met Leu Phe Ala Val Ile Thr Thr Ile Phe Ser Val Ala
Glu Lys Gln65 70 75 80Trp Thr Asn Leu Glu Trp Lys Pro Lys Pro Lys
Pro Arg Leu Pro Gln 85 90 95Leu Leu Asp Asp His Phe Gly Leu His Gly
Leu Val Phe Arg Arg Thr 100 105 110Phe Ala Ile Arg Ser Tyr Glu Val
Gly Pro Asp Arg Ser Thr Ser Ile 115 120 125Val Ala Val Met Asn His
Leu Gln Glu Ala Thr Leu Asn His Ala Lys 130 135 140Ser Val Gly Ile
Leu Gly Asp Gly Phe Gly Thr Thr Leu Glu Met Ser145 150 155 160Lys
Arg Asp Leu Ala Trp Val Val Arg Arg Thr His Val Ala Val Glu 165 170
175Arg Tyr Pro Ala Trp Gly Asp Thr Val Glu Val Glu Ala Trp Val Gly
180 185 190Ala Ser Gly Asn Ile Gly Met Arg Arg Asp Phe Leu Val Arg
Asp Cys 195 200 205Lys Thr Gly His Ile Leu Ala Arg Cys Thr Ser Val
Ser Val Met Met 210 215 220Asn Ala Arg Thr Arg Arg Leu Ser Lys Ile
Pro Gln Glu Val Arg Ala225 230 235 240Glu Ile Asp Pro Leu Phe Ile
Glu Lys Val Ala Val Lys Glu Gly Glu 245 250 255Ile Lys Lys Leu Gln
Lys Phe Asn Asp Ser Thr Ala Asp Tyr Ile Gln 260 265 270Gly Gly Trp
Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln His Val 275 280 285Asn
Asn Ile Lys Tyr Ile Gly Trp Ile Phe Lys Ser Val Pro Asp Ser 290 295
300Ile Ser Glu Asn His Tyr Leu Ser Ser Ile Thr Leu Glu Tyr Arg
Arg305 310 315 320Glu Cys Thr Arg Gly Ser Ala Leu Gln Ser Leu Thr
Thr Val Cys Gly 325 330 335Asp Ser Ser Glu Ala Gly Ile Ile Cys Glu
His Leu Leu Gln Leu Glu 340 345 350Asp Gly Pro Glu Val Leu Arg Ala
Arg Thr Glu Trp Arg Pro Lys Leu 355 360 365Thr Asp Ser Phe Arg Gly
Ile Ile Val Ile Pro Ala Glu Pro Ser Val 370 375 38018380PRTLindera
benzoin 18Met Val Ala Thr Ser Leu Ala Ser Ala Phe Cys Ser Met Lys
Ala Val1 5 10 15Met Leu Ala Asp Asp Gly Arg Gly Met Lys Pro Arg Ser
Ser Asp Leu 20 25 30Gln Leu Arg Ala Gly Asn Ala Gln Thr Ser Leu Lys
Met Ile Asp Gly 35 40 45Thr Lys Phe Ser Tyr Thr Glu Ser Leu Lys Arg
Leu Pro Asp Trp Ser 50 55 60Lys Leu Leu Thr Val Ile Thr Thr Ile Phe
Ser Ala Ala Glu Lys Gln65 70 75 80Trp Thr Asn Leu Glu Arg Lys Pro
Lys Pro Pro His Leu Leu Asp Asp 85 90 95Arg Phe Gly Leu His Gly Leu
Val Phe Arg Arg Thr Phe Ala Ile Arg 100 105 110Ser Tyr Glu Val Gly
Pro Asp Arg Ser Ala Ser Ile Leu Ala Val Leu 115 120 125Asn His Leu
Gln Glu Ala Thr Leu Asn His Ala Glu Ser Val Gly Ile 130 135 140Leu
Gly Asp Arg Phe Gly Glu Thr Leu Glu Met Ser Lys Arg Asp Leu145 150
155 160Met Trp Val Val Arg Arg Thr Tyr Val Ala Val Glu Arg Tyr Pro
Ala 165 170 175Trp Gly Asp Thr Val Glu Ile Glu Ser Trp Ile Gly Ala
Ser Gly Asn 180 185 190Asn Gly Met Arg Arg Glu Phe Leu Val Arg Asp
Phe Lys Thr Gly Glu 195 200 205Ile Leu Thr Arg Cys Thr Ser Leu Ser
Val Met Met Asn Thr Arg Thr 210 215 220Arg Arg Leu Ser Lys Ile Pro
Glu Glu Val Arg Gly Glu Ile Gly Pro225 230 235 240Val Phe Ile Asp
Asn Val Ala Val Lys Asp Glu Glu Ile Lys Lys Leu 245 250 255Gln Lys
Leu Asn Asp Ser Thr Ala Asp Tyr Ile Gln Gly Gly Leu Ile 260 265
270Pro Arg Trp Asn Asp Leu Asp Leu Asn Gln His Val Asn Asn Ile Lys
275 280 285Tyr Val Ser Trp Ile Leu Glu Thr Val Pro Asp Ser Ile Leu
Glu Ser 290 295 300Tyr His Met Ser Ser Ile Thr Leu Glu Tyr Arg Arg
Glu Cys Thr Arg305 310 315 320Asp Ser Val Leu Gln Ser Leu Thr Thr
Val Ser Gly Gly Ser Ser Glu 325 330 335Ala Gly Leu Val Cys Glu His
Ser Leu Leu Leu Glu Gly Gly Ser Glu 340 345 350Val Leu Arg Ala Arg
Thr Glu Trp Arg Pro Lys Leu Thr Asp Ser Phe 355 360 365Arg Gly Ile
Ser Val Ile Pro Ala Glu Gln Ser Val 370 375 380191257DNACuphea
crassiflora 19atggtggctg ctgcagcaag ttctgcattc ttccctgttc
ctgccccagg aacctccact 60aaacccagga agtccggcaa ttggccatcg agattgagcc
cttcctccaa gcccaagtca 120atccccaatg gcggatttca ggttaaggca
aatgccagtg cccatcctaa ggctaacggt 180tctgcagtaa atctaaagtc
tggcagcctc aacactcagg aggacacttc gtcgtcccct 240cctcctcggg
ctttccttaa ccagttgcct gattggagta tgcttctgac tgcaatcacg
300accgttttcg tggcggcaga gaagcagtgg acaatgcttg atcggaaatc
taagaggcct 360gacatgctcg tggactcggt tgggttgaag agtattgttc
gggatgggct cgtgtccaga 420caaagttttt cgatcaggtc
ttatgaaata ggcgctgatc gaacagcctc tatagagacg 480ctgatgaacc
acttgcagga aacatctatt aatcattgta agagtttggg ccttctcaat
540gacggctttg gtcggactcc tgggatgtgt aaaaacgacc tcatttgggt
gcttacaaaa 600atgcagatca tggtgaatcg ctacccaact tggggcgata
ctgttgagat caatacctgg 660ttctcccagt cggggaaaat cggtatgggt
agcgattggc taataagtga ttgcaataca 720ggagaaattc ttataagggc
aacgagcgtg tgggccatga tgaatcaaaa gacgagaaga 780ttctcaagac
ttccatacga ggttcgccag gagttaacgc ctcattttgt ggactctcct
840catgtcattg aagacaatga tcggaaattg cataagtttg atgtgaagac
tggcgattct 900attcgcaagg gtctaactcc gaggtggaat gatttggatg
tcaatcagca cgtaagcaac 960gtgaagtaca ttgggtggat tctcgagagt
atgccaatag aagttctgga gacccaggag 1020ctatgctctc tgacagttga
atataggcgg gaatgcggaa tggacagtaa gctggagtcc 1080gtgactgcta
tggatccctc agaagaagat ggagtccggt ctcagtacaa tcaccttctg
1140cggcttgagg atgggactga tgtcgtgaag ggcagaactg agtggcgacc
gaagaatgca 1200ggaactaacg gggcgatatc aacaggaaag acttcaaatg
gaaactcggt ttcttag 1257201248DNACuphea koehneana 20atggtcactg
ctgcagcaag ttctgcattc ttccctgttc cagccccggg aacctcccct 60aaacccggga
agtcctggcc atcgagcttg agcccttcct tcaagcccaa gtcaatcccc
120aatgccggat ttcaggttaa ggcaaatgcc agtgcccatc ctaaggctaa
cggttctgca 180gtaaatctaa agtctggcag cctcaacact caggaggaca
cttcgtcgtc ccctcctcct 240cgggctttcc ttaaccagtt gcctgattgg
agtatgcttc tgactgcaat cacgaccgtc 300ttcgtggcgg cagagaagca
gtggactatg cgtgatcgga aatctaagag gcctgacatg 360ctcgtggact
cggttggatc gaagagtatt gttctggatg ggctcgtgtc cagacagatt
420ttttcgatta gatcttatga aataggcgct gatcgaacag cctctataga
gacgctgatg 480aaccacttgc aggaaacatc tatcaatcat tgtaagagtt
tgggtcttct caatgacggc 540tttggtcgta ctcctgggat gtgtaaaaac
gacctcattt gggtgcttac aaaaatgcag 600atcatggtga atcgctaccc
aacttggggc gatactgttg agatcaatac ctggttctcc 660cattcgggga
aaatcggtat ggctagcgat tggctaataa ctgattgcaa cacaggagaa
720attcttataa gagcaacgag cgtgtgggcc atgatgaatc aaaagacgag
aagattctca 780agacttccat acgaggttcg ccaggagtta acgcctcatt
atgtggactc tcctcatgtc 840attgaagata atgatcggaa attgcataag
tttgatgtga agactggtga ttccattcgt 900aagggtctaa ctccgaagtg
gaatgacttg gatgtcaatc agcacgtcaa caacgtgaag 960tacatcgggt
ggattctcga gagtatgcca atagaagttt tggagactca ggagctatgc
1020tctctcaccg ttgaatatag gcgggaatgc ggaatggaca gtgtgctgga
gtccgtgact 1080gctatggatc cctcagaaga tggaggccta tctcagtaca
agcaccttct gcggcttgag 1140gatgggactg acatcgtgaa gggcagaact
gagtggcgac cgaagaatgc aggaactaac 1200ggggcgatat caacagcaaa
gccttcaaat ggaaactcgg tctcttag 1248211254DNACuphea leptopoda
21atggtgggtg ctgcagcaag ttctgcattc ttccctgctc cagccccggg aacctcccct
60aaacccggga agtccggcaa ttggccatca agcttgagcc cttccttaaa gcccaagtca
120atccccaatg gcggatttca ggttaaggca aatgccagtg cccatcctaa
ggctaacggt 180gctgcagtaa atctaaagtc tggcagcctc aacactcagg
aggacacttc gtcgtcccct 240cctcctcggg ctttccttaa ccagttgcct
gattggagta tgcttctgac tgcaatcacg 300accgtcttcg tggcggcaga
gaagcagtgg actatgcttg atcggaaatc taagaggcct 360gacatgctcg
tggactcggt tgggttgaag aatattgttc gggatgggct cgtgtccaga
420cagagttttt cgatcaggtc ttatgaaata ggcgctgatc gaacagcctc
tatagagacg 480ctgatgaacc acttgcagga aacatctatc aatcattgta
agagtttggg tcttctcaat 540gacggctttg gtcgtactcc tgggatgtgt
aaaaacgacc tcatttgggt gcttacaaaa 600atgcagatcc tggtgaatcg
ctacccagct tggggagata ctgttgagat caatacctgg 660ttctctcagt
cggggaaaat cggcatgggt agtgattggc taataagtga ttgcaacaca
720ggagaaattc ttataagagc aacgagcgtg tgggcaatga tgaatcaaaa
gacgagaaga 780ttctcaagac ttccatacga ggttcgccag gagttaacgc
ctcattttgt agactcacct 840catgtcattg aagacaatga tcggaaattg
cataagtttg atgtgaagac tggtgattct 900attcgcaagg gtctaactcc
gaggtggaat gacttggatg tcaatcaaca cgtaagcaac 960gtgaagtaca
ttgggtggat tctcgagagt atgccaatag aagttttgga gactcaggag
1020ctatgctctc tcaccgttga atataggcgg gaatgcggaa tggacagtgt
gctggagtcc 1080gtgactgcta gggatccctc agaagatgga ggccggtctc
agtacaatca ccttctgcgg 1140cttgaggatg ggactgatgt cgtgaagggc
agaactgagt ggcgatcgaa gaatgcagga 1200actaacgggg cgacatcaac
agcaaagact tcaaatggaa actcggtctc ttag 1254221254DNACuphea
angustifolia 22atggtggctg ctgcagcaag ttctgcattc ttccctgttc
cagccccggg aacatccctt 60aaacccggga agtccggcaa ttggccatcg agcttgagcc
cttccttcaa gcccaagaca 120atccccagtg gcggacttca ggttaaggca
aatgccagtg cccatcctaa ggctaacggt 180tctgcagtaa atctaaagtc
tggcagcctc gacactcagg aggacacttc gtcgtcccct 240cctcctcggg
ctttccttaa ccagttgcct gattggagta tgcttctgac tgcaatcacg
300accgtcttcg tggcggcaga gaagcagtgg actatgcttg ataggaaatc
taagaggcct 360gaaatgctcg tggactcggt tgggttgaag agtagtgttc
gggatgggct cgtgtccaga 420cagagttttt cgattaggtc ttatgaaata
ggcgctgatc gaacagcctc tatagagacg 480ctgatgaacc acttgcagga
aacatctatc aatcattgta agagtttggg tcttctcaac 540gatggctttg
gtcgtactcc tgggatgtgt aaaaacgacc tcatttgggt gcttacaaaa
600atgcagatca tggtgaatcg ctacccaact tggggcgata ctgttgaggt
caatacctgg 660ttctcccagt cggggaaaat cggtatggct agcgattggc
taatcagtga ttgcaacaca 720ggagaaattc ttataagagc aacaagcgtg
tgggccatga tgaatcaaaa gacgagaaga 780ttctcaagac ttccatacga
ggttcgccag gagctaacac ctcattatgt ggactctcct 840catgtcattg
aagataatga tcggaaattg cataagtttg atgtgaagac tggtgattcc
900attcgcaagg gtctaactcc gaggtggaat gacttggatg tcaatcagca
cgtaagcaac 960gtgaagtaca ttgggtggat tcttgagagt atgccaatag
aagttttgga gacccaggag 1020ctatgctctc tcaccgttga atataggcgg
gaatgcggaa tggacagtgt gctggagtcc 1080gtgactgcta tggatccctc
agaagatgga ggcgtgtctc agtacaagca ccttctgcgg 1140cttgaggatg
ggactgatat cgtgaagggc agaactgaat ggcgaccgaa gaatgcagga
1200actaatgggg cgacatcaaa agcaaagact tcaaatggaa actcggtctc ttag
1254231254DNACuphea llavea 23atggtggctg ctgcagcaag ttctgcattc
ttccctgctc cagccccggg atcctcacct 60aaacccggga agcccggtaa ttggccatcg
agcttgagcc cttccttcaa gcccaagtca 120atccccaatg gccgatttca
ggttaaggca aatgcgagtg cccatcctaa ggctaacggt 180tctgcagtaa
atctaaagtc tggcagcctc aacactcagg aggacacttc gtcgtcccct
240cctcctcggg ctttccttaa ccagttgcct gattggagta tgcttctgtc
tgcaatcacg 300actgtattcg tggcggcaga gaagcagtgg actatgcttg
atcggaaatc taagaggcct 360gacatgcttg tggactcggt tgggttgaag
aatattgttc gggatgggct cgtgtccaga 420cagagttttt cgattagatc
ttatgaaata ggcgctgatc gaacagcttc tatagagaca 480ctgatgaacc
acttgcagga aacatctatc aatcattgta agagtttggg tcttctcaat
540gacggctttg gtcgtactcc tgggatgtgt aaaaacgacc tcatttgggt
gcttacaaaa 600atgcagatca tggtgaatcg ctacccagct tggggcgata
ctgttgagat caatacatgg 660ttctcccagt cggggaaaat cggtatgggt
agcgattggc taataagtga ttgcaacaca 720ggagaaattc ttataagagc
aacgagcgtg tgggccatga tgaatcaaaa gacgagaaga 780ttctcaagac
ttccatatga ggttcgccag gagttaacgc ctcattttgt ggactctcct
840catgtcattg aagacaatga tcggaaattg cataagttcg atgtgaagac
tggtgattct 900attcgcaagg gtctaactcc gaggtggaat gacttggatg
tcaatcaaca cgtaagcaac 960gtgaagtaca ttgggtggat tctcgagagt
atgccaatag aagttttgga gacccaggaa 1020ctatgctctc tcacagttga
atataggcgg gaatgcggaa tggacagtgt gctggagtcc 1080gtgactgcta
tagatccctc agaagatgga gggcggtctc agtacaatca ccttctgcgg
1140cttgatgatg ggactgatgt cgtgaagggc agaacagagt ggcgaccgaa
gaatgcagga 1200actaacgggg cgatatcaac aggaaagact tcaaatggga
actcggtctc ctag 1254241254DNACuphea lophostoma 24atggtggctg
ctgcagcaag ttctgcattc ttccctgttc cagccccggg aacctccctt 60aaaccctgga
agtccggaaa ttggccatcg agcttgagcc cttccttcaa gcccaagaca
120atccccagtg gcggatttca ggttaaggca aatgccagtg cccagcctaa
ggctaacggt 180tctgcagtaa atctaaagtc tggcagcctc aacactcagg
aggacacaac gtcgtcgcct 240cctcctcggg ctttccttaa ccagttgcct
gattggagta tgcttctgac tgcaatcacg 300accgtcttcg tggcggcgga
gaagcagtgg acaatgcttg ataggaaatc taagaggcct 360gaaaagctcg
tggactcggt tgggttgaag agtagtgttc gggatgggct cgtgtccaga
420cagagttttt cgattaggtc ttatgaaata ggcgctgatc gaacagcctc
tatagagacg 480ttgatgaacc acttgcagga aacatctatc aatcattgta
agagtttggg tcttctcaac 540gacggctttg gtcgtactcc tgggatgtgt
aaaaacgacc tcatttgggt gcttacgaaa 600atgcagatca tggtgaatcg
ctacccaact tggggcgata ctgttgagat caatacctgg 660ttctcccagt
cggggaaaat cggtatggct agcgattggc taataagtga ttgcaacaca
720ggagaaattc ttataagagc aacgagcgtg tgggccatga tgaatcaaaa
gacgagaagg 780ttctcaagac ttccatacga ggttcgccag gagttaacgc
ctcattatgt ggactctcct 840catgtcattg aagacaatga tcggaaattg
cataagtttg atgtgaagac tggtgattcc 900attcgcaagg gtctgactcc
gaggtggaat gacttggatg tcaatcagca cgtaagcaac 960gtgaagtaca
ttgggtggat tctggagagt atgccaatag aagttttgga gacccaggag
1020ctatgctctc tcaccgttga atataggcgg gaatgcggga tggacagtgt
gctggagtcc 1080gtgactgcta tggatccctc agaagatgaa ggccggtctc
agtacaagca ccttctgcgg 1140cttgaggatg ggactgatat cgtgaagggc
agaactgagt ggcgaccgaa gaatgcagga 1200actaacgggg cgatatcaac
agcaaagaat tcaaatggaa actcggtctc ttag 1254251155DNASassafras
albidum 25atggccacca cctctttagc ttctgctttc tgctcgatga aagctgtaat
gttggctcgt 60gatggcaggg gcatgaaacc caggagcagt gatttgcagc tgagggcggg
aaatgcacaa 120acccctttga agatgatcaa tgggaccaag ttcagttaca
cggagagctt gaaaaggttg 180cctgactgga gcatgctctt tgcagtgatc
acaaccatct tttcggttgc tgagaagcag 240tggaccaatc tagagtggaa
gccgaagccg aagccgaggc taccccagtt gcttgatgac 300cattttggac
tgcatgggtt agttttcagg cgcacctttg ccatcagatc ttatgaggtc
360ggacctgacc gctccacatc tatagtggct gttatgaatc acttgcagga
ggctacactt 420aatcatgcga agagtgtggg aattctagga gatggattcg
gtacgacgct agagatgagt 480aagagagatc tggcgtgggt tgtgagacgc
acgcatgttg ctgtggaacg gtaccctgct 540tggggtgata ctgttgaagt
agagtgctgg attggtgcat ctggaaataa tggcatgcgc 600cgtgatttcc
ttgtccggga ctgcaaaaca ggcgaaattc ttacaagatg taccagtctt
660tcggtgatga tgaatacaag gacaaggagg ttgtccaaaa tccctgaaga
agttagaggg 720gagatagggc ctctattcat tgataatgtg gctgtcaagg
acgaggaaat taagaaacta 780cagaagctca atgacagctc tgcagattac
atccaaggag gtttgactcc tcgatggaat 840gatttggatg tcaatcagca
tgttaacaac atcaaatacg ttggctggat tcttgagact 900gtcccagact
ccatctttga gagtcatcat atttccagca tcactcttga atacaggaga
960gagtgcacca gggatagcgt gctgcagtcc ctgaccactg tctccggtgg
ctcgttggag 1020gctgggttag tgtgcgatca cttgctccag cttgaaggtg
ggtctgaggt attgagggca 1080agaacagagt ggaggcctaa gcttaccgat
agtttcagag ggattattgt gatacccgca 1140gaaccgagtg tgtaa
1155261155DNASassafras albidum 26atggccacca cctctttagc ttctgctttc
tgctcgatga aagctgtaat gttggctcgt 60gatggcaggg gcatgaaacc caggagcagt
gatttgcagc tgagggcggg aaatgcacaa 120acccctttga agatgatcaa
tgggaccaag ttcagttaca cggagagctt gaaaaggttg 180cctgactgga
gcatgctctt tgcagtgatc acaaccatct tttcggttgc tgagaagcag
240tggaccaatc tagagtggaa gccgaagccg aagccgaggc taccccagtt
gcttgatgac 300cattttggac tgcatgggtt agttttcagg cgcacctttg
ccatcagatc ttatgaggtc 360ggacctgacc gctccacatc tatagtggct
gttatgaatc acttgcagga ggctacactt 420aatcatgcga agagtgtggg
aattctagga gatggattcg gtacgacgct agagatgagt 480aagagagatc
tggcgtgggt tgtgagacgc acgcatgttg ctgtggaacg gtaccccgct
540tggggcgata ctgttgaagt cgaggcctgg gtcggtgcat ctggaaacat
tggcatgcgc 600cgcgattttc ttgtccgcga ctgcaaaact ggccacattc
ttgcaagatg taccagtgtt 660tcagtgatga tgaatgcgag gacacggaga
ttgtccaaaa ttccccaaga agttagagcc 720gagattgacc ctcttttcat
tgaaaaggtt gcggtcaagg aaggggaaat taagaaatta 780cagaagttca
atgatagcac tgcagattac attcaagggg gttggactcc tcgatggaat
840gatttggatg tcaatcagca cgtgaacaat atcaaataca ttggctggat
ttttaagagc 900gtcccagact ctatctctga gaatcattat ctttctagca
tcactctcga atacaggaga 960gagtgcacaa ggggcagcgc gctgcagtcc
ctgaccactg tttgtggtga ctcgtcggaa 1020gctgggatca tatgtgagca
cctactccag cttgaggatg ggcctgaggt tttgagggca 1080agaacagagt
ggaggcctaa gcttaccgat agtttcagag ggattattgt gatacccgca
1140gaaccgagtg tgtaa 1155271248DNALindera benzoin 27atggtcactg
ctgcagcaag ttctgcattc ttccctgttc cagccccggg aacctcccct 60aaacccggga
agtcctggcc atcgagcttg agcccttcct tcaagcccaa gtcaatcccc
120aatgccggat ttcaggttaa ggcaaatgcc agtgcccatc ctaaggctaa
cggttctgca 180gtaaatctaa agtctggcag cctcaacact caggaggaca
cttcgtcgtc ccctcctcct 240cgggctttcc ttaaccagtt gcctgattgg
agtatgcttc tgactgcaat cacgaccgtc 300ttcgtggcgg cagagaagca
gtggactatg cgtgatcgga aatctaagag gcctgacatg 360ctcgtggact
cggttggatc gaagagtatt gttctggatg ggctcgtgtc cagacagatt
420ttttcgatta gatcttatga aataggcgct gatcgaacag cctctataga
gacgctgatg 480aaccacttgc aggaaacatc tatcaatcat tgtaagagtt
tgggtcttct caatgacggc 540tttggtcgta ctcctgggat gtgtaaaaac
gacctcattt gggtgcttac aaaaatgcag 600atcatggtga atcgctaccc
aacttggggc gatactgttg agatcaatac ctggttctcc 660cattcgggga
aaatcggtat ggctagcgat tggctaataa ctgattgcaa cacaggagaa
720attcttataa gagcaacgag cgtgtgggcc atgatgaatc aaaagacgag
aagattctca 780agacttccat acgaggttcg ccaggagtta acgcctcatt
atgtggactc tcctcatgtc 840attgaagata atgatcggaa attgcataag
tttgatgtga agactggtga ttccattcgt 900aagggtctaa ctccgaagtg
gaatgacttg gatgtcaatc agcacgtcaa caacgtgaag 960tacatcgggt
ggattctcga gagtatgcca atagaagttt tggagactca ggagctatgc
1020tctctcaccg ttgaatatag gcgggaatgc ggaatggaca gtgtgctgga
gtccgtgact 1080gctatggatc cctcagaaga tggaggccta tctcagtaca
agcaccttct gcggcttgag 1140gatgggactg acatcgtgaa gggcagaact
gagtggcgac cgaagaatgc aggaactaac 1200ggggcgatat caacagcaaa
gccttcaaat ggaaactcgg tctcttag 1248281257DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
28atggtggccg ccgccgcctc ctccgccttc ttccccgtgc ccgcccccgg cacctccacc
60aagccccgca agtccggcaa ctggccctcc cgcctgtccc cctcctccaa gcccaagtcc
120atccccaacg gcggcttcca ggtgaaggcc aacgcctccg cccaccccaa
ggccaacggc 180tccgccgtga acctgaagtc cggctccctg aacacccagg
aggacacctc ctcctccccc 240cccccccgcg ccttcctgaa ccagctgccc
gactggtcca tgctgctgac cgccatcacc 300accgtgttcg tggccgccga
gaagcagtgg accatgctgg accgcaagtc caagcgcccc 360gacatgctgg
tggactccgt gggcctgaag tccatcgtgc gcgacggcct ggtgtcccgc
420cagtccttct ccatccgctc ctacgagatc ggcgccgacc gcaccgcctc
catcgagacc 480ctgatgaacc acctgcagga gacctccatc aaccactgca
agtccctggg cctgctgaac 540gacggcttcg gccgcacccc cggcatgtgc
aagaacgacc tgatctgggt gctgaccaag 600atgcagatca tggtgaaccg
ctaccccacc tggggcgaca ccgtggagat caacacctgg 660ttctcccagt
ccggcaagat cggcatgggc tccgactggc tgatctccga ctgcaacacc
720ggcgagatcc tgatccgcgc cacctccgtg tgggccatga tgaaccagaa
gacccgccgc 780ttctcccgcc tgccctacga ggtgcgccag gagctgaccc
cccacttcgt ggactccccc 840cacgtgatcg aggacaacga ccgcaagctg
cacaagttcg acgtgaagac cggcgactcc 900atccgcaagg gcctgacccc
ccgctggaac gacctggacg tgaaccagca cgtgtccaac 960gtgaagtaca
tcggctggat cctggagtcc atgcccatcg aggtgctgga gacccaggag
1020ctgtgctccc tgaccgtgga gtaccgccgc gagtgcggca tggactccaa
gctggagtcc 1080gtgaccgcca tggacccctc cgaggaggac ggcgtgcgct
cccagtacaa ccacctgctg 1140cgcctggagg acggcaccga cgtggtgaag
ggccgcaccg agtggcgccc caagaacgcc 1200ggcaccaacg gcgccatctc
caccggcaag acctccaacg gcaactccgt gtcctga 1257291248DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
29atggtgaccg ccgccgcctc ctccgccttc ttccccgtgc ccgcccccgg cacctccccc
60aagcccggca agtcctggcc ctcctccctg tccccctcct tcaagcccaa gtccatcccc
120aacgccggct tccaggtgaa ggccaacgcc tccgcccacc ccaaggccaa
cggctccgcc 180gtgaacctga agtccggctc cctgaacacc caggaggaca
cctcctcctc cccccccccc 240cgcgccttcc tgaaccagct gcccgactgg
tccatgctgc tgaccgccat caccaccgtg 300ttcgtggccg ccgagaagca
gtggaccatg cgcgaccgca agtccaagcg ccccgacatg 360ctggtggact
ccgtgggctc caagtccatc gtgctggacg gcctggtgtc ccgccagatc
420ttctccatcc gctcctacga gatcggcgcc gaccgcaccg cctccatcga
gaccctgatg 480aaccacctgc aggagacctc catcaaccac tgcaagtccc
tgggcctgct gaacgacggc 540ttcggccgca cccccggcat gtgcaagaac
gacctgatct gggtgctgac caagatgcag 600atcatggtga accgctaccc
cacctggggc gacaccgtgg agatcaacac ctggttctcc 660cactccggca
agatcggcat ggcctccgac tggctgatca ccgactgcaa caccggcgag
720atcctgatcc gcgccacctc cgtgtgggcc atgatgaacc agaagacccg
ccgcttctcc 780cgcctgccct acgaggtgcg ccaggagctg accccccact
acgtggactc cccccacgtg 840atcgaggaca acgaccgcaa gctgcacaag
ttcgacgtga agaccggcga ctccatccgc 900aagggcctga cccccaagtg
gaacgacctg gacgtgaacc agcacgtgaa caacgtgaag 960tacatcggct
ggatcctgga gtccatgccc atcgaggtgc tggagaccca ggagctgtgc
1020tccctgaccg tggagtaccg ccgcgagtgc ggcatggact ccgtgctgga
gtccgtgacc 1080gccatggacc cctccgagga cggcggcctg tcccagtaca
agcacctgct gcgcctggag 1140gacggcaccg acatcgtgaa gggccgcacc
gagtggcgcc ccaagaacgc cggcaccaac 1200ggcgccatct ccaccgccaa
gccctccaac ggcaactccg tgtcctga 1248301254DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
30atggtgggcg ccgccgcctc ctccgccttc ttccccgccc ccgcccccgg cacctccccc
60aagcccggca agtccggcaa ctggccctcc tccctgtccc cctccctgaa gcccaagtcc
120atccccaacg gcggcttcca ggtgaaggcc aacgcctccg cccaccccaa
ggccaacggc 180gccgccgtga acctgaagtc cggctccctg aacacccagg
aggacacctc ctcctccccc 240cccccccgcg ccttcctgaa ccagctgccc
gactggtcca tgctgctgac cgccatcacc 300accgtgttcg tggccgccga
gaagcagtgg accatgctgg accgcaagtc caagcgcccc 360gacatgctgg
tggactccgt gggcctgaag aacatcgtgc gcgacggcct ggtgtcccgc
420cagtccttct ccatccgctc ctacgagatc ggcgccgacc gcaccgcctc
catcgagacc 480ctgatgaacc acctgcagga gacctccatc aaccactgca
agtccctggg cctgctgaac 540gacggcttcg gccgcacccc cggcatgtgc
aagaacgacc tgatctgggt gctgaccaag 600atgcagatcc tggtgaaccg
ctaccccgcc tggggcgaca ccgtggagat caacacctgg 660ttctcccagt
ccggcaagat cggcatgggc tccgactggc tgatctccga ctgcaacacc
720ggcgagatcc tgatccgcgc cacctccgtg tgggccatga tgaaccagaa
gacccgccgc 780ttctcccgcc tgccctacga ggtgcgccag gagctgaccc
cccacttcgt ggactccccc 840cacgtgatcg aggacaacga ccgcaagctg
cacaagttcg acgtgaagac cggcgactcc 900atccgcaagg gcctgacccc
ccgctggaac gacctggacg tgaaccagca cgtgtccaac 960gtgaagtaca
tcggctggat cctggagtcc atgcccatcg aggtgctgga gacccaggag
1020ctgtgctccc tgaccgtgga gtaccgccgc gagtgcggca tggactccgt
gctggagtcc 1080gtgaccgccc gcgacccctc cgaggacggc ggccgctccc
agtacaacca cctgctgcgc 1140ctggaggacg gcaccgacgt ggtgaagggc
cgcaccgagt ggcgctccaa gaacgccggc 1200accaacggcg ccacctccac
cgccaagacc tccaacggca actccgtgtc ctga
1254311254DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 31atggtggccg ccgccgcctc ctccgccttc
ttccccgtgc ccgcccccgg cacctccctg 60aagcccggca agtccggcaa ctggccctcc
tccctgtccc cctccttcaa gcccaagacc 120atcccctccg gcggcctgca
ggtgaaggcc aacgcctccg cccaccccaa ggccaacggc 180tccgccgtga
acctgaagtc cggctccctg gacacccagg aggacacctc ctcctccccc
240cccccccgcg ccttcctgaa ccagctgccc gactggtcca tgctgctgac
cgccatcacc 300accgtgttcg tggccgccga gaagcagtgg accatgctgg
accgcaagtc caagcgcccc 360gagatgctgg tggactccgt gggcctgaag
tcctccgtgc gcgacggcct ggtgtcccgc 420cagtccttct ccatccgctc
ctacgagatc ggcgccgacc gcaccgcctc catcgagacc 480ctgatgaacc
acctgcagga gacctccatc aaccactgca agtccctggg cctgctgaac
540gacggcttcg gccgcacccc cggcatgtgc aagaacgacc tgatctgggt
gctgaccaag 600atgcagatca tggtgaaccg ctaccccacc tggggcgaca
ccgtggaggt gaacacctgg 660ttctcccagt ccggcaagat cggcatggcc
tccgactggc tgatctccga ctgcaacacc 720ggcgagatcc tgatccgcgc
cacctccgtg tgggccatga tgaaccagaa gacccgccgc 780ttctcccgcc
tgccctacga ggtgcgccag gagctgaccc cccactacgt ggactccccc
840cacgtgatcg aggacaacga ccgcaagctg cacaagttcg acgtgaagac
cggcgactcc 900atccgcaagg gcctgacccc ccgctggaac gacctggacg
tgaaccagca cgtgtccaac 960gtgaagtaca tcggctggat cctggagtcc
atgcccatcg aggtgctgga gacccaggag 1020ctgtgctccc tgaccgtgga
gtaccgccgc gagtgcggca tggactccgt gctggagtcc 1080gtgaccgcca
tggacccctc cgaggacggc ggcgtgtccc agtacaagca cctgctgcgc
1140ctggaggacg gcaccgacat cgtgaagggc cgcaccgagt ggcgccccaa
gaacgccggc 1200accaacggcg ccacctccaa ggccaagacc tccaacggca
actccgtgtc ctga 1254321254DNAArtificial SequenceDescription of
Artificial Sequence Synthetic polynucleotide 32atggtggccg
ccgccgcctc ctccgccttc ttccccgccc ccgcccccgg ctcctccccc 60aagcccggca
agcccggcaa ctggccctcc tccctgtccc cctccttcaa gcccaagtcc
120atccccaacg gccgcttcca ggtgaaggcc aacgcctccg cccaccccaa
ggccaacggc 180tccgccgtga acctgaagtc cggctccctg aacacccagg
aggacacctc ctcctccccc 240cccccccgcg ccttcctgaa ccagctgccc
gactggtcca tgctgctgtc cgccatcacc 300accgtgttcg tggccgccga
gaagcagtgg accatgctgg accgcaagtc caagcgcccc 360gacatgctgg
tggactccgt gggcctgaag aacatcgtgc gcgacggcct ggtgtcccgc
420cagtccttct ccatccgctc ctacgagatc ggcgccgacc gcaccgcctc
catcgagacc 480ctgatgaacc acctgcagga gacctccatc aaccactgca
agtccctggg cctgctgaac 540gacggcttcg gccgcacccc cggcatgtgc
aagaacgacc tgatctgggt gctgaccaag 600atgcagatca tggtgaaccg
ctaccccgcc tggggcgaca ccgtggagat caacacctgg 660ttctcccagt
ccggcaagat cggcatgggc tccgactggc tgatctccga ctgcaacacc
720ggcgagatcc tgatccgcgc cacctccgtg tgggccatga tgaaccagaa
gacccgccgc 780ttctcccgcc tgccctacga ggtgcgccag gagctgaccc
cccacttcgt ggactccccc 840cacgtgatcg aggacaacga ccgcaagctg
cacaagttcg acgtgaagac cggcgactcc 900atccgcaagg gcctgacccc
ccgctggaac gacctggacg tgaaccagca cgtgtccaac 960gtgaagtaca
tcggctggat cctggagtcc atgcccatcg aggtgctgga gacccaggag
1020ctgtgctccc tgaccgtgga gtaccgccgc gagtgcggca tggactccgt
gctggagtcc 1080gtgaccgcca tcgacccctc cgaggacggc ggccgctccc
agtacaacca cctgctgcgc 1140ctggacgacg gcaccgacgt ggtgaagggc
cgcaccgagt ggcgccccaa gaacgccggc 1200accaacggcg ccatctccac
cggcaagacc tccaacggca actccgtgtc ctga 1254331254DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
33atggtggccg ccgccgcctc ctccgccttc ttccccgtgc ccgcccccgg cacctccctg
60aagccctgga agtccggcaa ctggccctcc tccctgtccc cctccttcaa gcccaagacc
120atcccctccg gcggcttcca ggtgaaggcc aacgcctccg cccagcccaa
ggccaacggc 180tccgccgtga acctgaagtc cggctccctg aacacccagg
aggacaccac ctcctccccc 240cccccccgcg ccttcctgaa ccagctgccc
gactggtcca tgctgctgac cgccatcacc 300accgtgttcg tggccgccga
gaagcagtgg accatgctgg accgcaagtc caagcgcccc 360gagaagctgg
tggactccgt gggcctgaag tcctccgtgc gcgacggcct ggtgtcccgc
420cagtccttct ccatccgctc ctacgagatc ggcgccgacc gcaccgcctc
catcgagacc 480ctgatgaacc acctgcagga gacctccatc aaccactgca
agtccctggg cctgctgaac 540gacggcttcg gccgcacccc cggcatgtgc
aagaacgacc tgatctgggt gctgaccaag 600atgcagatca tggtgaaccg
ctaccccacc tggggcgaca ccgtggagat caacacctgg 660ttctcccagt
ccggcaagat cggcatggcc tccgactggc tgatctccga ctgcaacacc
720ggcgagatcc tgatccgcgc cacctccgtg tgggccatga tgaaccagaa
gacccgccgc 780ttctcccgcc tgccctacga ggtgcgccag gagctgaccc
cccactacgt ggactccccc 840cacgtgatcg aggacaacga ccgcaagctg
cacaagttcg acgtgaagac cggcgactcc 900atccgcaagg gcctgacccc
ccgctggaac gacctggacg tgaaccagca cgtgtccaac 960gtgaagtaca
tcggctggat cctggagtcc atgcccatcg aggtgctgga gacccaggag
1020ctgtgctccc tgaccgtgga gtaccgccgc gagtgcggca tggactccgt
gctggagtcc 1080gtgaccgcca tggacccctc cgaggacgag ggccgctccc
agtacaagca cctgctgcgc 1140ctggaggacg gcaccgacat cgtgaagggc
cgcaccgagt ggcgccccaa gaacgccggc 1200accaacggcg ccatctccac
cgccaagaac tccaacggca actccgtgtc ctga 1254341155DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
34atggccacca cctccctggc ctccgccttc tgctccatga aggccgtgat gctggcccgc
60gacggccgcg gcatgaagcc ccgctcctcc gacctgcagc tgcgcgccgg caacgcccag
120acccccctga agatgatcaa cggcaccaag ttctcctaca ccgagtccct
gaagcgcctg 180cccgactggt ccatgctgtt cgccgtgatc accaccatct
tctccgtggc cgagaagcag 240tggaccaacc tggagtggaa gcccaagccc
aagccccgcc tgccccagct gctggacgac 300cacttcggcc tgcacggcct
ggtgttccgc cgcaccttcg ccatccgctc ctacgaggtg 360ggccccgacc
gctccacctc catcgtggcc gtgatgaacc acctgcagga ggccaccctg
420aaccacgcca agtccgtggg catcctgggc gacggcttcg gcaccaccct
ggagatgtcc 480aagcgcgacc tggcctgggt ggtgcgccgc acccacgtgg
ccgtggagcg ctaccccgcc 540tggggcgaca ccgtggaggt ggagtgctgg
atcggcgcct ccggcaacaa cggcatgcgc 600cgcgacttcc tggtgcgcga
ctgcaagacc ggcgagatcc tgacccgctg cacctccctg 660tccgtgatga
tgaacacccg cacccgccgc ctgtccaaga tccccgagga ggtgcgcggc
720gagatcggcc ccctgttcat cgacaacgtg gccgtgaagg acgaggagat
caagaagctg 780cagaagctga acgactcctc cgccgactac atccagggcg
gcctgacccc ccgctggaac 840gacctggacg tgaaccagca cgtgaacaac
atcaagtacg tgggctggat cctggagacc 900gtgcccgact ccatcttcga
gtcccaccac atctcctcca tcaccctgga gtaccgccgc 960gagtgcaccc
gcgactccgt gctgcagtcc ctgaccaccg tgtccggcgg ctccctggag
1020gccggcctgg tgtgcgacca cctgctgcag ctggagggcg gctccgaggt
gctgcgcgcc 1080cgcaccgagt ggcgccccaa gctgaccgac tccttccgcg
gcatcatcgt gatccccgcc 1140gagccctccg tgtga 1155351155DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
35atggccacca cctccctggc ctccgccttc tgctccatga aggccgtgat gctggcccgc
60gacggccgcg gcatgaagcc ccgctcctcc gacctgcagc tgcgcgccgg caacgcccag
120acccccctga agatgatcaa cggcaccaag ttctcctaca ccgagtccct
gaagcgcctg 180cccgactggt ccatgctgtt cgccgtgatc accaccatct
tctccgtggc cgagaagcag 240tggaccaacc tggagtggaa gcccaagccc
aagccccgcc tgccccagct gctggacgac 300cacttcggcc tgcacggcct
ggtgttccgc cgcaccttcg ccatccgctc ctacgaggtg 360ggccccgacc
gctccacctc catcgtggcc gtgatgaacc acctgcagga ggccaccctg
420aaccacgcca agtccgtggg catcctgggc gacggcttcg gcaccaccct
ggagatgtcc 480aagcgcgacc tggcctgggt ggtgcgccgc acccacgtgg
ccgtggagcg ctaccccgcc 540tggggcgaca ccgtggaggt ggaggcctgg
gtgggcgcct ccggcaacat cggcatgcgc 600cgcgacttcc tggtgcgcga
ctgcaagacc ggccacatcc tggcccgctg cacctccgtg 660tccgtgatga
tgaacgcccg cacccgccgc ctgtccaaga tcccccagga ggtgcgcgcc
720gagatcgacc ccctgttcat cgagaaggtg gccgtgaagg agggcgagat
caagaagctg 780cagaagttca acgactccac cgccgactac atccagggcg
gctggacccc ccgctggaac 840gacctggacg tgaaccagca cgtgaacaac
atcaagtaca tcggctggat cttcaagtcc 900gtgcccgact ccatctccga
gaaccactac ctgtcctcca tcaccctgga gtaccgccgc 960gagtgcaccc
gcggctccgc cctgcagtcc ctgaccaccg tgtgcggcga ctcctccgag
1020gccggcatca tctgcgagca cctgctgcag ctggaggacg gccccgaggt
gctgcgcgcc 1080cgcaccgagt ggcgccccaa gctgaccgac tccttccgcg
gcatcatcgt gatccccgcc 1140gagccctccg tgtga 1155361248DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
36atggtgaccg ccgccgcctc ctccgccttc ttccccgtgc ccgcccccgg cacctccccc
60aagcccggca agtcctggcc ctcctccctg tccccctcct tcaagcccaa gtccatcccc
120aacgccggct tccaggtgaa ggccaacgcc tccgcccacc ccaaggccaa
cggctccgcc 180gtgaacctga agtccggctc cctgaacacc caggaggaca
cctcctcctc cccccccccc 240cgcgccttcc tgaaccagct gcccgactgg
tccatgctgc tgaccgccat caccaccgtg 300ttcgtggccg ccgagaagca
gtggaccatg cgcgaccgca agtccaagcg ccccgacatg 360ctggtggact
ccgtgggctc caagtccatc gtgctggacg gcctggtgtc ccgccagatc
420ttctccatcc gctcctacga gatcggcgcc gaccgcaccg cctccatcga
gaccctgatg 480aaccacctgc aggagacctc catcaaccac tgcaagtccc
tgggcctgct gaacgacggc 540ttcggccgca cccccggcat gtgcaagaac
gacctgatct gggtgctgac caagatgcag 600atcatggtga accgctaccc
cacctggggc gacaccgtgg agatcaacac ctggttctcc 660cactccggca
agatcggcat ggcctccgac tggctgatca ccgactgcaa caccggcgag
720atcctgatcc gcgccacctc cgtgtgggcc atgatgaacc agaagacccg
ccgcttctcc 780cgcctgccct acgaggtgcg ccaggagctg accccccact
acgtggactc cccccacgtg 840atcgaggaca acgaccgcaa gctgcacaag
ttcgacgtga agaccggcga ctccatccgc 900aagggcctga cccccaagtg
gaacgacctg gacgtgaacc agcacgtgaa caacgtgaag 960tacatcggct
ggatcctgga gtccatgccc atcgaggtgc tggagaccca ggagctgtgc
1020tccctgaccg tggagtaccg ccgcgagtgc ggcatggact ccgtgctgga
gtccgtgacc 1080gccatggacc cctccgagga cggcggcctg tcccagtaca
agcacctgct gcgcctggag 1140gacggcaccg acatcgtgaa gggccgcacc
gagtggcgcc ccaagaacgc cggcaccaac 1200ggcgccatct ccaccgccaa
gccctccaac ggcaactccg tgtcctga 12483738PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
37Met Ala Thr Ala Ser Thr Phe Ser Ala Phe Asn Ala Arg Cys Gly Asp1
5 10 15Leu Arg Arg Ser Ala Gly Ser Gly Pro Arg Arg Pro Ala Arg Pro
Leu 20 25 30Pro Val Arg Ala Ala Ile 35386398DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
38gaagagcgcc caatgtttaa acccctcaac tgcgacgctg ggaaccttct ccgggcaggc
60gatgtgcgtg ggtttgcctc cttggcacgg ctctacaccg tcgagtacgc catgaggcgg
120tgatggctgt gtcggttgcc acttcgtcca gagacggcaa gtcgtccatc
ctctgcgtgt 180gtggcgcgac gctgcagcag tccctctgca gcagatgagc
gtgactttgg ccatttcacg 240cactcgagtg tacacaatcc atttttctta
aagcaaatga ctgctgattg accagatact 300gtaacgctga tttcgctcca
gatcgcacag atagcgacca tgttgctgcg tctgaaaatc 360tggattccga
attcgaccct ggcgctccat ccatgcaaca gatggcgaca cttgttacaa
420ttcctgtcac ccatcggcat ggagcaggtc cacttagatt cccgatcacc
cacgcacatc 480tcgctaatag tcattcgttc gtgtcttcga tcaatctcaa
gtgagtgtgc atggatcttg 540gttgacgatg cggtatgggt ttgcgccgct
ggctgcaggg tctgcccaag gcaagctaac 600ccagctcctc tccccgacaa
tactctcgca ggcaaagccg gtcacttgcc ttccagattg 660ccaataaact
caattatggc ctctgtcatg ccatccatgg gtctgatgaa tggtcacgct
720cgtgtcctga ccgttcccca gcctctggcg tcccctgccc cgcccaccag
cccacgccgc 780gcggcagtcg ctgccaaggc tgtctcggag gtaccctttc
ttgcgctatg acacttccag 840caaaaggtag ggcgggctgc gagacggctt
cccggcgctg catgcaacac cgatgatgct 900tcgacccccc gaagctcctt
cggggctgca tgggcgctcc gatgccgctc cagggcgagc 960gctgtttaaa
tagccaggcc cccgattgca aagacattat agcgagctac caaagccata
1020ttcaaacacc tagatcacta ccacttctac acaggccact cgagcttgtg
atcgcactcc 1080gctaaggggg cgcctcttcc tcttcgtttc agtcacaacc
cgcaaactct agaatatcaa 1140tgctgctgca ggccttcctg ttcctgctgg
ccggcttcgc cgccaagatc agcgcctcca 1200tgacgaacga gacgtccgac
cgccccctgg tgcacttcac ccccaacaag ggctggatga 1260acgaccccaa
cggcctgtgg tacgacgaga aggacgccaa gtggcacctg tacttccagt
1320acaacccgaa cgacaccgtc tgggggacgc ccttgttctg gggccacgcc
acgtccgacg 1380acctgaccaa ctgggaggac cagcccatcg ccatcgcccc
gaagcgcaac gactccggcg 1440ccttctccgg ctccatggtg gtggactaca
acaacacctc cggcttcttc aacgacacca 1500tcgacccgcg ccagcgctgc
gtggccatct ggacctacaa caccccggag tccgaggagc 1560agtacatctc
ctacagcctg gacggcggct acaccttcac cgagtaccag aagaaccccg
1620tgctggccgc caactccacc cagttccgcg acccgaaggt cttctggtac
gagccctccc 1680agaagtggat catgaccgcg gccaagtccc aggactacaa
gatcgagatc tactcctccg 1740acgacctgaa gtcctggaag ctggagtccg
cgttcgccaa cgagggcttc ctcggctacc 1800agtacgagtg ccccggcctg
atcgaggtcc ccaccgagca ggaccccagc aagtcctact 1860gggtgatgtt
catctccatc aaccccggcg ccccggccgg cggctccttc aaccagtact
1920tcgtcggcag cttcaacggc acccacttcg aggccttcga caaccagtcc
cgcgtggtgg 1980acttcggcaa ggactactac gccctgcaga ccttcttcaa
caccgacccg acctacggga 2040gcgccctggg catcgcgtgg gcctccaact
gggagtactc cgccttcgtg cccaccaacc 2100cctggcgctc ctccatgtcc
ctcgtgcgca agttctccct caacaccgag taccaggcca 2160acccggagac
ggagctgatc aacctgaagg ccgagccgat cctgaacatc agcaacgccg
2220gcccctggag ccggttcgcc accaacacca cgttgacgaa ggccaacagc
tacaacgtcg 2280acctgtccaa cagcaccggc accctggagt tcgagctggt
gtacgccgtc aacaccaccc 2340agacgatctc caagtccgtg ttcgcggacc
tctccctctg gttcaagggc ctggaggacc 2400ccgaggagta cctccgcatg
ggcttcgagg tgtccgcgtc ctccttcttc ctggaccgcg 2460ggaacagcaa
ggtgaagttc gtgaaggaga acccctactt caccaaccgc atgagcgtga
2520acaaccagcc cttcaagagc gagaacgacc tgtcctacta caaggtgtac
ggcttgctgg 2580accagaacat cctggagctg tacttcaacg acggcgacgt
cgtgtccacc aacacctact 2640tcatgaccac cgggaacgcc ctgggctccg
tgaacatgac gacgggggtg gacaacctgt 2700tctacatcga caagttccag
gtgcgcgagg tcaagtgaca attgacgccc gcgcggcgca 2760cctgacctgt
tctctcgagg gcgcctgttc tgccttgcga aacaagcccc tggagcatgc
2820gtgcatgatc gtctctggcg ccccgccgcg cggtttgtcg ccctcgcggg
cgccgcggcc 2880gcgggggcgc attgaaattg ttgcaaaccc cacctgacag
attgagggcc caggcaggaa 2940ggcgttgaga tggaggtaca ggagtcaagt
aactgaaagt ttttatgata actaacaaca 3000aagggtcgtt tctggccagc
gaatgacaag aacaagattc cacatttccg tgtagaggct 3060tgccatcgaa
tgtgagcggg cgggccgcgg acccgacaaa acccttacga cgtggtaaga
3120aaaacgtggc gggcactgtc cctgtagcct gaagaccagc aggagacgat
cggaagcatc 3180acagcacagg atcccgcgtc tcgaacagag cgcgcagagg
aacgctgaag gtctcgcctc 3240tgtcgcacct cagcgcggca tacaccacaa
taaccacctg acgaatgcgc ttggttcttc 3300gtccattagc gaagcgtccg
gttcacacac gtgccacgtt ggcgaggtgg caggtgacaa 3360tgatcggtgg
agctgatggt cgaaacgttc acagcctagg gatatcgtga aaactcgctc
3420gaccgcccgc gtcccgcagg cagcgatgac gtgtgcgtga cctgggtgtt
tcgtcgaaag 3480gccagcaacc ccaaatcgca ggcgatccgg agattgggat
ctgatccgag cttggaccag 3540atcccccacg atgcggcacg ggaactgcat
cgactcggcg cggaacccag ctttcgtaaa 3600tgccagattg gtgtccgata
ccttgatttg ccatcagcga aacaagactt cagcagcgag 3660cgtatttggc
gggcgtgcta ccagggttgc atacattgcc catttctgtc tggaccgctt
3720taccggcgca gagggtgagt tgatggggtt ggcaggcatc gaaacgcgcg
tgcatggtgt 3780gtgtgtctgt tttcggctgc acaatttcaa tagtcggatg
ggcgacggta gaattgggtg 3840ttgcgctcgc gtgcatgcct cgccccgtcg
ggtgtcatga ccgggactgg aatcccccct 3900cgcgaccctc ctgctaacgc
tcccgactct cccgcccgcg cgcaggatag actctagttc 3960aaccaatcga
caactagtat ggccaccgcc tccaccttct ccgccttcaa cgcccgctgc
4020ggcgacctgc gccgctccgc cggctccggc ccccgccgcc ccgcccgccc
cctgcccgtg 4080cgcgccgcca tcaacgcctc cgcccacccc aaggccaacg
gctccgccgt gaacctgaag 4140tccggctccc tgaacaccca ggaggacacc
tcctcctccc cccccccccg cgccttcctg 4200aaccagctgc ccgactggtc
catgctgctg accgccatca ccaccgtgtt cgtggccgcc 4260gagaagcagt
ggaccatgct ggaccgcaag tccaagcgcc ccgacatgct ggtggactcc
4320gtgggcctga agtccatcgt gcgcgacggc ctggtgtccc gccagtcctt
ctccatccgc 4380tcctacgaga tcggcgccga ccgcaccgcc tccatcgaga
ccctgatgaa ccacctgcag 4440gagacctcca tcaaccactg caagtccctg
ggcctgctga acgacggctt cggccgcacc 4500cccggcatgt gcaagaacga
cctgatctgg gtgctgacca agatgcagat catggtgaac 4560cgctacccca
cctggggcga caccgtggag atcaacacct ggttctccca gtccggcaag
4620atcggcatgg gctccgactg gctgatctcc gactgcaaca ccggcgagat
cctgatccgc 4680gccacctccg tgtgggccat gatgaaccag aagacccgcc
gcttctcccg cctgccctac 4740gaggtgcgcc aggagctgac cccccacttc
gtggactccc cccacgtgat cgaggacaac 4800gaccgcaagc tgcacaagtt
cgacgtgaag accggcgact ccatccgcaa gggcctgacc 4860ccccgctgga
acgacctgga cgtgaaccag cacgtgtcca acgtgaagta catcggctgg
4920atcctggagt ccatgcccat cgaggtgctg gagacccagg agctgtgctc
cctgaccgtg 4980gagtaccgcc gcgagtgcgg catggactcc aagctggagt
ccgtgaccgc catggacccc 5040tccgaggagg acggcgtgcg ctcccagtac
aaccacctgc tgcgcctgga ggacggcacc 5100gacgtggtga agggccgcac
cgagtggcgc cccaagaacg ccggcaccaa cggcgccatc 5160tccaccggca
agacctccaa cggcaactcc gtgtccatgg actacaagga ccacgacggc
5220gactacaagg accacgacat cgactacaag gacgacgacg acaagtgact
cgaggcagca 5280gcagctcaga tagtatcgac acactctgga cgctggtcgt
gtgatggact gttgccgcca 5340cacttgctgc cttgacctgt gaatatccct
gccgctttta tcaaacagcc tcagtgtgtt 5400tgatcttgtg tgtacgcgct
tttgcgagtt gctagctgct tgtgctattt gcgaatacca 5460cccccagcat
ccccttccct cgtttcatat cgcttgcatc ccaaccgcaa cttatctacg
5520ctgtcctgct atccctcagc gctgctcctg ctcctgctca ctgcccctcg
cacagccttg 5580gtttgggctc cgcctgtatt ctcctggtac tgcaacctgt
aaaccagcac tgcaatgctg 5640atgcacggga agtagtggga tgggaacaca
aatggaaagc ttgagctcca gcgccatgcc 5700acgccctttg atggcttcaa
gtacgattac ggtgttggat tgtgtgtttg ttgcgtagtg 5760tgcatggttt
agaataatac acttgatttc ttgctcacgg caatctcggc ttgtccgcag
5820gttcaacccc atttcggagt ctcaggtcag ccgcgcaatg accagccgct
acttcaagga 5880cttgcacgac aacgccgagg tgagctatgt ttaggacttg
attggaaatt gtcgtcgacg 5940catattcgcg ctccgcgaca gcacccaagc
aaaatgtcaa gtgcgttccg atttgcgtcc 6000gcaggtcgat gttgtgatcg
tcggcgccgg atccgccggt ctgtcctgcg cttacgagct 6060gaccaagcac
cctgacgtcc gggtacgcga gctgagattc gattagacat aaattgaaga
6120ttaaacccgt agaaaaattt gatggtcgcg aaactgtgct cgattgcaag
aaattgatcg 6180tcctccactc cgcaggtcgc catcatcgag cagggcgttg
ctcccggcgg cggcgcctgg 6240ctggggggac agctgttctc ggccatgtgt
gtacgtagaa ggatgaattt cagctggttt 6300tcgttgcaca gctgtttgtg
catgatttgt ttcagactat tgttgaatgt ttttagattt 6360cttaggatgc
atgatttgtc tgcatgcgac tgaagagc 63983932PRTSassafras albidum 39Leu
Phe Ala Val Ile Thr Thr Ile Phe Ser Val Ala Glu Lys Gln Trp1 5 10
15Thr Asn Leu Glu Trp Lys Pro Lys Pro Lys Pro Arg Leu Pro Gln Leu
20 25 3040352PRTArtificial SequenceDescription of Artificial
Sequence
Synthetic polypeptide 40Met Ala Thr Thr Ser Leu Ala Ser Ala Phe Cys
Ser Met Lys Ala Val1 5 10 15Met Leu Ala Arg Asp Gly Arg Gly Met Lys
Pro Arg Ser Ser Asp Leu 20 25 30Gln Leu Arg Ala Gly Asn Ala Gln Thr
Pro Leu Lys Met Ile Asn Gly 35 40 45Thr Lys Phe Ser Tyr Thr Glu Ser
Leu Lys Arg Leu Pro Asp Trp Ser 50 55 60Met Leu Asp Asp His Phe Gly
Leu His Gly Leu Val Phe Arg Arg Thr65 70 75 80Phe Ala Ile Arg Ser
Tyr Glu Val Gly Pro Asp Arg Ser Thr Ser Ile 85 90 95Val Ala Val Met
Asn His Leu Gln Glu Ala Thr Leu Asn His Ala Lys 100 105 110Ser Val
Gly Ile Leu Gly Asp Gly Phe Gly Thr Thr Leu Glu Met Ser 115 120
125Lys Arg Asp Leu Ala Trp Val Val Arg Arg Thr His Val Ala Val Glu
130 135 140Arg Tyr Pro Ala Trp Gly Asp Thr Val Glu Val Glu Cys Trp
Ile Gly145 150 155 160Ala Ser Gly Asn Asn Gly Met Arg Arg Asp Phe
Leu Val Arg Asp Cys 165 170 175Lys Thr Gly Glu Ile Leu Thr Arg Cys
Thr Ser Leu Ser Val Met Met 180 185 190Asn Thr Arg Thr Arg Arg Leu
Ser Lys Ile Pro Glu Glu Val Arg Gly 195 200 205Glu Ile Gly Pro Leu
Phe Ile Asp Asn Val Ala Val Lys Asp Glu Glu 210 215 220Ile Lys Lys
Leu Gln Lys Leu Asn Asp Ser Ser Ala Asp Tyr Ile Gln225 230 235
240Gly Gly Leu Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln His Val
245 250 255Asn Asn Ile Lys Tyr Val Gly Trp Ile Leu Glu Thr Val Pro
Asp Ser 260 265 270Ile Phe Glu Ser His His Ile Ser Ser Ile Thr Leu
Glu Tyr Arg Arg 275 280 285Glu Cys Thr Arg Asp Ser Val Leu Gln Ser
Leu Thr Thr Val Ser Gly 290 295 300Gly Ser Leu Glu Ala Gly Leu Val
Cys Asp His Leu Leu Gln Leu Glu305 310 315 320Gly Gly Ser Glu Val
Leu Arg Ala Arg Thr Glu Trp Arg Pro Lys Leu 325 330 335Thr Asp Ser
Phe Arg Gly Ile Ile Val Ile Pro Ala Glu Pro Ser Val 340 345
35041352PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 41Met Val Ala Thr Ser Leu Ala Ser Ala Phe Cys
Ser Met Lys Ala Val1 5 10 15Met Leu Ala Asp Asp Gly Arg Gly Met Lys
Pro Arg Ser Ser Asp Leu 20 25 30Gln Leu Arg Ala Gly Asn Ala Gln Thr
Ser Leu Lys Met Ile Asp Gly 35 40 45Thr Lys Phe Ser Tyr Thr Glu Ser
Leu Lys Arg Leu Pro Asp Trp Ser 50 55 60Lys Leu Asp Asp Arg Phe Gly
Leu His Gly Leu Val Phe Arg Arg Thr65 70 75 80Phe Ala Ile Arg Ser
Tyr Glu Val Gly Pro Asp Arg Ser Ala Ser Ile 85 90 95Leu Ala Val Leu
Asn His Leu Gln Glu Ala Thr Leu Asn His Ala Glu 100 105 110Ser Val
Gly Ile Leu Gly Asp Arg Phe Gly Glu Thr Leu Glu Met Ser 115 120
125Lys Arg Asp Leu Met Trp Val Val Arg Arg Thr Tyr Val Ala Val Glu
130 135 140Arg Tyr Pro Ala Trp Gly Asp Thr Val Glu Ile Glu Ser Trp
Ile Gly145 150 155 160Ala Ser Gly Asn Asn Gly Met Arg Arg Glu Phe
Leu Val Arg Asp Phe 165 170 175Lys Thr Gly Glu Ile Leu Thr Arg Cys
Thr Ser Leu Ser Val Met Met 180 185 190Asn Thr Arg Thr Arg Arg Leu
Ser Lys Ile Pro Glu Glu Val Arg Gly 195 200 205Glu Ile Gly Pro Val
Phe Ile Asp Asn Val Ala Val Lys Asp Glu Glu 210 215 220Ile Lys Lys
Leu Gln Lys Leu Asn Asp Ser Thr Ala Asp Tyr Ile Gln225 230 235
240Gly Gly Leu Ile Pro Arg Trp Asn Asp Leu Asp Leu Asn Gln His Val
245 250 255Asn Asn Ile Lys Tyr Val Ser Trp Ile Leu Glu Thr Val Pro
Asp Ser 260 265 270Ile Leu Glu Ser Tyr His Met Ser Ser Ile Thr Leu
Glu Tyr Arg Arg 275 280 285Glu Cys Thr Arg Asp Ser Val Leu Gln Ser
Leu Thr Thr Val Ser Gly 290 295 300Gly Ser Ser Glu Ala Gly Leu Val
Cys Glu His Ser Leu Leu Leu Glu305 310 315 320Gly Gly Ser Glu Val
Leu Arg Ala Arg Thr Glu Trp Arg Pro Lys Leu 325 330 335Thr Asp Ser
Phe Arg Gly Ile Ser Val Ile Pro Ala Glu Gln Ser Val 340 345
35042316PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 42Gly Asn Ala Gln Thr Pro Leu Lys Met Ile Asn
Gly Thr Lys Phe Ser1 5 10 15Tyr Thr Glu Ser Leu Lys Arg Leu Pro Asp
Trp Ser Met Leu Asp Asp 20 25 30His Phe Gly Leu His Gly Leu Val Phe
Arg Arg Thr Phe Ala Ile Arg 35 40 45Ser Tyr Glu Val Gly Pro Asp Arg
Ser Thr Ser Ile Val Ala Val Met 50 55 60Asn His Leu Gln Glu Ala Thr
Leu Asn His Ala Lys Ser Val Gly Ile65 70 75 80Leu Gly Asp Gly Phe
Gly Thr Thr Leu Glu Met Ser Lys Arg Asp Leu 85 90 95Ala Trp Val Val
Arg Arg Thr His Val Ala Val Glu Arg Tyr Pro Ala 100 105 110Trp Gly
Asp Thr Val Glu Val Glu Cys Trp Ile Gly Ala Ser Gly Asn 115 120
125Asn Gly Met Arg Arg Asp Phe Leu Val Arg Asp Cys Lys Thr Gly Glu
130 135 140Ile Leu Thr Arg Cys Thr Ser Leu Ser Val Met Met Asn Thr
Arg Thr145 150 155 160Arg Arg Leu Ser Lys Ile Pro Glu Glu Val Arg
Gly Glu Ile Gly Pro 165 170 175Leu Phe Ile Asp Asn Val Ala Val Lys
Asp Glu Glu Ile Lys Lys Leu 180 185 190Gln Lys Leu Asn Asp Ser Ser
Ala Asp Tyr Ile Gln Gly Gly Leu Thr 195 200 205Pro Arg Trp Asn Asp
Leu Asp Val Asn Gln His Val Asn Asn Ile Lys 210 215 220Tyr Val Gly
Trp Ile Leu Glu Thr Val Pro Asp Ser Ile Phe Glu Ser225 230 235
240His His Ile Ser Ser Ile Thr Leu Glu Tyr Arg Arg Glu Cys Thr Arg
245 250 255Asp Ser Val Leu Gln Ser Leu Thr Thr Val Ser Gly Gly Ser
Leu Glu 260 265 270Ala Gly Leu Val Cys Asp His Leu Leu Gln Leu Glu
Gly Gly Ser Glu 275 280 285Val Leu Arg Ala Arg Thr Glu Trp Arg Pro
Lys Leu Thr Asp Ser Phe 290 295 300Arg Gly Ile Ile Val Ile Pro Ala
Glu Pro Ser Val305 310 31543316PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 43Gly Asn Ala Gln Thr Ser
Leu Lys Met Ile Asp Gly Thr Lys Phe Ser1 5 10 15Tyr Thr Glu Ser Leu
Lys Arg Leu Pro Asp Trp Ser Lys Leu Asp Asp 20 25 30Arg Phe Gly Leu
His Gly Leu Val Phe Arg Arg Thr Phe Ala Ile Arg 35 40 45Ser Tyr Glu
Val Gly Pro Asp Arg Ser Ala Ser Ile Leu Ala Val Leu 50 55 60Asn His
Leu Gln Glu Ala Thr Leu Asn His Ala Glu Ser Val Gly Ile65 70 75
80Leu Gly Asp Arg Phe Gly Glu Thr Leu Glu Met Ser Lys Arg Asp Leu
85 90 95Met Trp Val Val Arg Arg Thr Tyr Val Ala Val Glu Arg Tyr Pro
Ala 100 105 110Trp Gly Asp Thr Val Glu Ile Glu Ser Trp Ile Gly Ala
Ser Gly Asn 115 120 125Asn Gly Met Arg Arg Glu Phe Leu Val Arg Asp
Phe Lys Thr Gly Glu 130 135 140Ile Leu Thr Arg Cys Thr Ser Leu Ser
Val Met Met Asn Thr Arg Thr145 150 155 160Arg Arg Leu Ser Lys Ile
Pro Glu Glu Val Arg Gly Glu Ile Gly Pro 165 170 175Val Phe Ile Asp
Asn Val Ala Val Lys Asp Glu Glu Ile Lys Lys Leu 180 185 190Gln Lys
Leu Asn Asp Ser Thr Ala Asp Tyr Ile Gln Gly Gly Leu Ile 195 200
205Pro Arg Trp Asn Asp Leu Asp Leu Asn Gln His Val Asn Asn Ile Lys
210 215 220Tyr Val Ser Trp Ile Leu Glu Thr Val Pro Asp Ser Ile Leu
Glu Ser225 230 235 240Tyr His Met Ser Ser Ile Thr Leu Glu Tyr Arg
Arg Glu Cys Thr Arg 245 250 255Asp Ser Val Leu Gln Ser Leu Thr Thr
Val Ser Gly Gly Ser Ser Glu 260 265 270Ala Gly Leu Val Cys Glu His
Ser Leu Leu Leu Glu Gly Gly Ser Glu 275 280 285Val Leu Arg Ala Arg
Thr Glu Trp Arg Pro Lys Leu Thr Asp Ser Phe 290 295 300Arg Gly Ile
Ser Val Ile Pro Ala Glu Gln Ser Val305 310 315441134DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
44atggccaccg cctccacctt ctccgccttc aacgcccgct gcggcgacct gcgccgctcc
60gccggctccg gcccccgccg ccccgcccgc cccctgcccg tgcgcgccgc catcggcaac
120gcccagaccc ccctgaagat gatcaacggc accaagttct cctacaccga
gtccctgaag 180cgcctgcccg actggtccat gctggacgac cacttcggcc
tgcacggcct ggtgttccgc 240cgcaccttcg ccatccgctc ctacgaggtg
ggccccgacc gctccacctc catcgtggcc 300gtgatgaacc acctgcagga
ggccaccctg aaccacgcca agtccgtggg catcctgggc 360gacggcttcg
gcaccaccct ggagatgtcc aagcgcgacc tggcctgggt ggtgcgccgc
420acccacgtgg ccgtggagcg ctaccccgcc tggggcgaca ccgtggaggt
ggagtgctgg 480atcggcgcct ccggcaacaa cggcatgcgc cgcgacttcc
tggtgcgcga ctgcaagacc 540ggcgagatcc tgacccgctg cacctccctg
tccgtgatga tgaacacccg cacccgccgc 600ctgtccaaga tccccgagga
ggtgcgcggc gagatcggcc ccctgttcat cgacaacgtg 660gccgtgaagg
acgaggagat caagaagctg cagaagctga acgactcctc cgccgactac
720atccagggcg gcctgacccc ccgctggaac gacctggacg tgaaccagca
cgtgaacaac 780atcaagtacg tgggctggat cctggagacc gtgcccgact
ccatcttcga gtcccaccac 840atctcctcca tcaccctgga gtaccgccgc
gagtgcaccc gcgactccgt gctgcagtcc 900ctgaccaccg tgtccggcgg
ctccctggag gccggcctgg tgtgcgacca cctgctgcag 960ctggagggcg
gctccgaggt gctgcgcgcc cgcaccgagt ggcgccccaa gctgaccgac
1020tccttccgcg gcatcatcgt gatccccgcc gagccctccg tgatggacta
caaggaccac 1080gacggcgact acaaggacca cgacatcgac tacaaggacg
acgacgacaa gtga 1134451134DNAArtificial SequenceDescription of
Artificial Sequence Synthetic polynucleotide 45atggccaccg
cctccacctt ctccgccttc aacgcccgct gcggcgacct gcgccgctcc 60gccggctccg
gcccccgccg ccccgcccgc cccctgcccg tgcgcgccgc catcggcaac
120gcccagacct ccctgaagat gatcgacggc accaagttct cctacaccga
gtccctgaag 180cgcctgcccg actggtccaa gctggacgac cgcttcggcc
tgcacggcct ggtgttccgc 240cgcaccttcg ccatccgctc ctacgaggtg
ggccccgacc gctccgcctc catcctggcc 300gtgctgaacc acctgcagga
ggccaccctg aaccacgccg agtccgtggg catcctgggc 360gaccgcttcg
gcgagaccct ggagatgtcc aagcgcgacc tgatgtgggt ggtgcgccgc
420acctacgtgg ccgtggagcg ctaccccgcc tggggcgaca ccgtggagat
cgagtcctgg 480atcggcgcct ccggcaacaa cggcatgcgc cgcgagttcc
tggtgcgcga cttcaagacc 540ggcgagatcc tgacccgctg cacctccctg
tccgtgatga tgaacacccg cacccgccgc 600ctgtccaaga tccccgagga
ggtgcgcggc gagatcggcc ccgtgttcat cgacaacgtg 660gccgtgaagg
acgaggagat caagaagctg cagaagctga acgactccac cgccgactac
720atccagggcg gcctgatccc ccgctggaac gacctggacc tgaaccagca
cgtgaacaac 780atcaagtacg tgtcctggat cctggagacc gtgcccgact
ccatcctgga gtcctaccac 840atgtcctcca tcaccctgga gtaccgccgc
gagtgcaccc gcgactccgt gctgcagtcc 900ctgaccaccg tgtccggcgg
ctcctccgag gccggcctgg tgtgcgagca ctccctgctg 960ctggagggcg
gctccgaggt gctgcgcgcc cgcaccgagt ggcgccccaa gctgaccgac
1020tccttccgcg gcatctccgt gatccccgcc gagcagtccg tgatggacta
caaggaccac 1080gacggcgact acaaggacca cgacatcgac tacaaggacg
acgacgacaa gtga 113446357PRTCinnamomum camphora 46Met Val Thr Thr
Ser Leu Ala Ser Ala Tyr Phe Ser Met Lys Ala Val1 5 10 15Met Leu Ala
Pro Asp Gly Arg Gly Ile Lys Pro Arg Ser Ser Gly Leu 20 25 30Gln Val
Arg Ala Gly Asn Glu Arg Asn Ser Cys Lys Val Ile Asn Gly 35 40 45Thr
Lys Val Lys Asp Thr Glu Gly Leu Lys Gly Cys Ser Thr Leu Gln 50 55
60Gly Gln Ser Met Leu Asp Asp His Phe Gly Leu His Gly Leu Val Phe65
70 75 80Arg Arg Thr Phe Ala Ile Arg Cys Tyr Glu Val Gly Pro Asp Arg
Ser 85 90 95Thr Ser Ile Met Ala Val Met Asn His Leu Gln Glu Ala Ala
Arg Asn 100 105 110His Ala Glu Ser Leu Gly Leu Leu Gly Asp Gly Phe
Gly Glu Thr Leu 115 120 125Glu Met Ser Lys Arg Asp Leu Ile Trp Val
Val Arg Arg Thr His Val 130 135 140Ala Val Glu Arg Tyr Pro Ala Trp
Gly Asp Thr Val Glu Val Glu Ala145 150 155 160Trp Val Gly Ala Ser
Gly Asn Thr Gly Met Arg Arg Asp Phe Leu Val 165 170 175Arg Asp Cys
Lys Thr Gly His Ile Leu Thr Arg Cys Thr Ser Val Ser 180 185 190Val
Met Met Asn Met Arg Thr Arg Arg Leu Ser Lys Ile Pro Gln Glu 195 200
205Val Arg Ala Glu Ile Asp Pro Leu Phe Ile Glu Lys Val Ala Val Lys
210 215 220Glu Gly Glu Ile Lys Lys Leu Gln Lys Leu Asn Asp Ser Thr
Ala Asp225 230 235 240Tyr Ile Gln Gly Gly Trp Thr Pro Arg Trp Asn
Asp Leu Asp Val Asn 245 250 255Gln His Val Asn Asn Ile Ile Tyr Val
Gly Trp Ile Phe Lys Ser Val 260 265 270Pro Asp Ser Ile Ser Glu Asn
His His Leu Ser Ser Ile Thr Leu Glu 275 280 285Tyr Arg Arg Glu Cys
Thr Arg Gly Asn Lys Leu Gln Ser Leu Thr Thr 290 295 300Val Cys Gly
Gly Ser Ser Glu Ala Gly Ile Ile Cys Glu His Leu Leu305 310 315
320Gln Leu Glu Asp Gly Ser Glu Val Leu Arg Ala Arg Thr Glu Trp Arg
325 330 335Pro Lys His Thr Asp Ser Phe Gln Gly Ile Ser Glu Arg Phe
Pro Gln 340 345 350Gln Glu Pro His Lys 3554732PRTSassafras albidum
47Leu Phe Ala Val Ile Thr Thr Ile Phe Ser Val Ala Glu Lys Gln Trp1
5 10 15Thr Asn Leu Glu Trp Lys Pro Lys Pro Lys Pro Arg Leu Pro Gln
Leu 20 25 304828PRTLindera benzoin 48Leu Leu Thr Val Ile Thr Thr
Ile Phe Ser Ala Ala Glu Lys Gln Trp1 5 10 15Thr Asn Leu Glu Arg Lys
Pro Lys Pro Pro His Leu 20 2549405PRTArtificial SequenceDescription
of Artificial Sequence Synthetic polypeptide 49Met Ala Thr Ala Ser
Thr Phe Ser Ala Phe Asn Ala Arg Cys Gly Asp1 5 10 15Leu Arg Arg Ser
Ala Gly Ser Gly Pro Arg Arg Pro Ala Arg Pro Leu 20 25 30Pro Val Arg
Ala Ala Ile Asn Ala Ser Ala His Pro Lys Ala Asn Gly 35 40 45Ser Ala
Val Asn Leu Lys Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr 50 55 60Ser
Ser Ser Pro Pro Pro Arg Ala Phe Leu Asn Gln Leu Pro Asp Trp65 70 75
80Ser Met Leu Leu Thr Ala Ile Thr Thr Val Phe Val Ala Ala Glu Lys
85 90 95Gln Trp Thr Met Arg Asp Arg Lys Ser Lys Arg Pro Asp Met Leu
Val 100 105 110Asp Ser Val Gly Leu Lys Ser Val Val Leu Asp Gly Leu
Val Ser Arg 115 120 125Gln Ile Phe Ser Ile Arg Ser Tyr Glu Ile Gly
Ala Asp Arg Thr Ala 130 135 140Ser Ile Glu Thr Leu Met Asn His Leu
Gln Glu Thr Ser Ile Asn His145 150 155 160Cys Lys Ser Leu Gly Leu
Leu Asn Asp Gly Phe Gly Arg Thr Pro Gly 165 170 175Met Cys Lys Asn
Asp Leu Ile Trp Val Leu Thr Lys Met Gln Ile Met 180 185 190Val Asn
Arg Tyr Pro Thr Trp Gly Asp Thr Val Glu Ile Asn Thr Trp 195 200
205Phe Ser His Ser Gly Lys Ile Gly Met Ala Ser Asp Trp Leu Ile Thr
210 215 220Asp Cys Asn Thr Gly Glu Ile Leu Ile Arg Ala Thr Ser Val
Trp Ala225 230 235 240Met Met Asn Gln Lys Thr Arg Arg Phe Ser Arg
Leu Pro Tyr Glu Val 245 250 255Arg Gln Glu Leu Thr
Pro His Tyr Val Asp Ser Pro His Val Ile Glu 260 265 270Asp Asn Asp
Arg Lys Leu His Lys Phe Asp Val Lys Thr Gly Asp Ser 275 280 285Ile
Arg Lys Gly Leu Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln 290 295
300His Val Ser Asn Val Lys Tyr Ile Gly Trp Ile Leu Glu Ser Met
Pro305 310 315 320Ile Glu Val Leu Glu Thr Gln Glu Leu Cys Ser Leu
Thr Val Glu Tyr 325 330 335Arg Arg Glu Cys Gly Met Asp Ser Val Leu
Glu Ser Val Thr Ala Met 340 345 350Asp Pro Ser Glu Asp Glu Gly Arg
Ser Gln Tyr Lys His Leu Leu Arg 355 360 365Leu Glu Asp Gly Thr Asp
Ile Val Lys Gly Arg Thr Glu Trp Arg Pro 370 375 380Lys Asn Ala Gly
Thr Asn Gly Ala Ile Ser Thr Ala Lys Pro Ser Asn385 390 395 400Gly
Asn Ser Val Ser 40550377PRTArtificial SequenceDescription of
Artificial Sequence Synthetic polypeptide 50Met Ala Thr Ala Ser Thr
Phe Ser Ala Phe Asn Ala Arg Cys Gly Asp1 5 10 15Leu Arg Arg Ser Ala
Gly Ser Gly Pro Arg Arg Pro Ala Arg Pro Leu 20 25 30Pro Val Arg Ala
Ala Ile Asn Ala Ser Ala His Pro Lys Ala Asn Gly 35 40 45Ser Ala Val
Asn Leu Lys Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr 50 55 60Ser Ser
Ser Pro Pro Pro Arg Ala Phe Leu Asn Gln Leu Pro Asp Trp65 70 75
80Ser Met Leu Val Asp Ser Val Gly Leu Lys Ser Val Val Leu Asp Gly
85 90 95Leu Val Ser Arg Gln Ile Phe Ser Ile Arg Ser Tyr Glu Ile Gly
Ala 100 105 110Asp Arg Thr Ala Ser Ile Glu Thr Leu Met Asn His Leu
Gln Glu Thr 115 120 125Ser Ile Asn His Cys Lys Ser Leu Gly Leu Leu
Asn Asp Gly Phe Gly 130 135 140Arg Thr Pro Gly Met Cys Lys Asn Asp
Leu Ile Trp Val Leu Thr Lys145 150 155 160Met Gln Ile Met Val Asn
Arg Tyr Pro Thr Trp Gly Asp Thr Val Glu 165 170 175Ile Asn Thr Trp
Phe Ser His Ser Gly Lys Ile Gly Met Ala Ser Asp 180 185 190Trp Leu
Ile Thr Asp Cys Asn Thr Gly Glu Ile Leu Ile Arg Ala Thr 195 200
205Ser Val Trp Ala Met Met Asn Gln Lys Thr Arg Arg Phe Ser Arg Leu
210 215 220Pro Tyr Glu Val Arg Gln Glu Leu Thr Pro His Tyr Val Asp
Ser Pro225 230 235 240His Val Ile Glu Asp Asn Asp Arg Lys Leu His
Lys Phe Asp Val Lys 245 250 255Thr Gly Asp Ser Ile Arg Lys Gly Leu
Thr Pro Arg Trp Asn Asp Leu 260 265 270Asp Val Asn Gln His Val Ser
Asn Val Lys Tyr Ile Gly Trp Ile Leu 275 280 285Glu Ser Met Pro Ile
Glu Val Leu Glu Thr Gln Glu Leu Cys Ser Leu 290 295 300Thr Val Glu
Tyr Arg Arg Glu Cys Gly Met Asp Ser Val Leu Glu Ser305 310 315
320Val Thr Ala Met Asp Pro Ser Glu Asp Glu Gly Arg Ser Gln Tyr Lys
325 330 335His Leu Leu Arg Leu Glu Asp Gly Thr Asp Ile Val Lys Gly
Arg Thr 340 345 350Glu Trp Arg Pro Lys Asn Ala Gly Thr Asn Gly Ala
Ile Ser Thr Ala 355 360 365Lys Pro Ser Asn Gly Asn Ser Val Ser 370
37551339PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 51Asn Ala Ser Ala His Pro Lys Ala Asn Gly Ser
Ala Val Asn Leu Lys1 5 10 15Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr
Ser Ser Ser Pro Pro Pro 20 25 30Arg Ala Phe Leu Asn Gln Leu Pro Asp
Trp Ser Met Leu Val Asp Ser 35 40 45Val Gly Leu Lys Ser Val Val Leu
Asp Gly Leu Val Ser Arg Gln Ile 50 55 60Phe Ser Ile Arg Ser Tyr Glu
Ile Gly Ala Asp Arg Thr Ala Ser Ile65 70 75 80Glu Thr Leu Met Asn
His Leu Gln Glu Thr Ser Ile Asn His Cys Lys 85 90 95Ser Leu Gly Leu
Leu Asn Asp Gly Phe Gly Arg Thr Pro Gly Met Cys 100 105 110Lys Asn
Asp Leu Ile Trp Val Leu Thr Lys Met Gln Ile Met Val Asn 115 120
125Arg Tyr Pro Thr Trp Gly Asp Thr Val Glu Ile Asn Thr Trp Phe Ser
130 135 140His Ser Gly Lys Ile Gly Met Ala Ser Asp Trp Leu Ile Thr
Asp Cys145 150 155 160Asn Thr Gly Glu Ile Leu Ile Arg Ala Thr Ser
Val Trp Ala Met Met 165 170 175Asn Gln Lys Thr Arg Arg Phe Ser Arg
Leu Pro Tyr Glu Val Arg Gln 180 185 190Glu Leu Thr Pro His Tyr Val
Asp Ser Pro His Val Ile Glu Asp Asn 195 200 205Asp Arg Lys Leu His
Lys Phe Asp Val Lys Thr Gly Asp Ser Ile Arg 210 215 220Lys Gly Leu
Thr Pro Arg Trp Asn Asp Leu Asp Val Asn Gln His Val225 230 235
240Ser Asn Val Lys Tyr Ile Gly Trp Ile Leu Glu Ser Met Pro Ile Glu
245 250 255Val Leu Glu Thr Gln Glu Leu Cys Ser Leu Thr Val Glu Tyr
Arg Arg 260 265 270Glu Cys Gly Met Asp Ser Val Leu Glu Ser Val Thr
Ala Met Asp Pro 275 280 285Ser Glu Asp Glu Gly Arg Ser Gln Tyr Lys
His Leu Leu Arg Leu Glu 290 295 300Asp Gly Thr Asp Ile Val Lys Gly
Arg Thr Glu Trp Arg Pro Lys Asn305 310 315 320Ala Gly Thr Asn Gly
Ala Ile Ser Thr Ala Lys Pro Ser Asn Gly Asn 325 330 335Ser Val
Ser521203DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 52atggccaccg cctccacctt ctccgccttc
aacgcccgct gcggcgacct gcgccgctcc 60gccggctccg gcccccgccg ccccgcccgc
cccctgcccg tgcgcgccgc catcaacgcc 120tccgcccacc ccaaggccaa
cggctccgcc gtgaacctga agtccggctc cctgaacacc 180caggaggaca
cctcctcctc cccccccccc cgcgccttcc tgaaccagct gcccgactgg
240tccatgctgg tggactccgt gggcctgaag tccgtggtgc tggacggcct
ggtgtcccgc 300cagatcttct ccatccgctc ctacgagatc ggcgccgacc
gcaccgcctc catcgagacc 360ctgatgaacc acctgcagga gacctccatc
aaccactgca agtccctggg cctgctgaac 420gacggcttcg gccgcacccc
cggcatgtgc aagaacgacc tgatctgggt gctgaccaag 480atgcagatca
tggtgaaccg ctaccccacc tggggcgaca ccgtggagat caacacctgg
540ttctcccact ccggcaagat cggcatggcc tccgactggc tgatcaccga
ctgcaacacc 600ggcgagatcc tgatccgcgc cacctccgtg tgggccatga
tgaaccagaa gacccgccgc 660ttctcccgcc tgccctacga ggtgcgccag
gagctgaccc cccactacgt ggactccccc 720cacgtgatcg aggacaacga
ccgcaagctg cacaagttcg acgtgaagac cggcgactcc 780atccgcaagg
gcctgacccc ccgctggaac gacctggacg tgaaccagca cgtgtccaac
840gtgaagtaca tcggctggat cctggagtcc atgcccatcg aggtgctgga
gacccaggag 900ctgtgctccc tgaccgtgga gtaccgccgc gagtgcggca
tggactccgt gctggagtcc 960gtgaccgcca tggacccctc cgaggacgag
ggccgctccc agtacaagca cctgctgcgc 1020ctggaggacg gcaccgacat
cgtgaagggc cgcaccgagt ggcgccccaa gaacgccggc 1080accaacggcg
ccatctccac cgccaagccc tccaacggca actccgtgtc catggactac
1140aaggaccacg acggcgacta caaggaccac gacatcgact acaaggacga
cgacgacaag 1200tga 120353415PRTCuphea hookeriana 53Met Val Ala Ala
Ala Ala Ser Ser Ala Phe Phe Pro Val Pro Ala Pro1 5 10 15Gly Ala Ser
Pro Lys Pro Gly Lys Phe Gly Asn Trp Pro Ser Ser Leu 20 25 30Ser Pro
Ser Phe Lys Pro Lys Ser Ile Pro Asn Gly Gly Phe Gln Val 35 40 45Lys
Ala Asn Asp Ser Ala His Pro Lys Ala Asn Gly Ser Ala Val Ser 50 55
60Leu Lys Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr Ser Ser Ser Pro65
70 75 80Pro Pro Arg Thr Phe Leu His Gln Leu Pro Asp Trp Ser Arg Leu
Leu 85 90 95Thr Ala Ile Thr Thr Val Phe Val Lys Ser Lys Arg Pro Asp
Met His 100 105 110Asp Arg Lys Ser Lys Arg Pro Asp Met Leu Val Asp
Ser Phe Gly Leu 115 120 125Glu Ser Thr Val Gln Asp Gly Leu Val Phe
Arg Gln Ser Phe Ser Ile 130 135 140Arg Ser Tyr Glu Ile Gly Thr Asp
Arg Thr Ala Ser Ile Glu Thr Leu145 150 155 160Met Asn His Leu Gln
Glu Thr Ser Leu Asn His Cys Lys Ser Thr Gly 165 170 175Ile Leu Leu
Asp Gly Phe Gly Arg Thr Leu Glu Met Cys Lys Arg Asp 180 185 190Leu
Ile Trp Val Val Ile Lys Met Gln Ile Lys Val Asn Arg Tyr Pro 195 200
205Ala Trp Gly Asp Thr Val Glu Ile Asn Thr Arg Phe Ser Arg Leu Gly
210 215 220Lys Ile Gly Met Gly Arg Asp Trp Leu Ile Ser Asp Cys Asn
Thr Gly225 230 235 240Glu Ile Leu Val Arg Ala Thr Ser Ala Tyr Ala
Met Met Asn Gln Lys 245 250 255Thr Arg Arg Leu Ser Lys Leu Pro Tyr
Glu Val His Gln Glu Ile Val 260 265 270Pro Leu Phe Val Asp Ser Pro
Val Ile Glu Asp Ser Asp Leu Lys Val 275 280 285His Lys Phe Lys Val
Lys Thr Gly Asp Ser Ile Gln Lys Gly Leu Thr 290 295 300Pro Gly Trp
Asn Asp Leu Asp Val Asn Gln His Val Ser Asn Val Lys305 310 315
320Tyr Ile Gly Trp Ile Leu Glu Ser Met Pro Thr Glu Val Leu Glu Thr
325 330 335Gln Glu Leu Cys Ser Leu Ala Leu Glu Tyr Arg Arg Glu Cys
Gly Arg 340 345 350Asp Ser Val Leu Glu Ser Val Thr Ala Met Asp Pro
Ser Lys Val Gly 355 360 365Val Arg Ser Gln Tyr Gln His Leu Leu Arg
Leu Glu Asp Gly Thr Ala 370 375 380Ile Val Asn Gly Ala Thr Glu Trp
Arg Pro Lys Asn Ala Gly Ala Asn385 390 395 400Gly Ala Ile Ser Thr
Gly Lys Thr Ser Asn Gly Asn Ser Val Ser 405 410
41554376PRTArtificial SequenceDescription of Artificial Sequence
Synthetic polypeptide 54Met Ala Thr Ala Ser Thr Phe Ser Ala Phe Asn
Ala Arg Cys Gly Asp1 5 10 15Leu Arg Arg Ser Ala Gly Ser Gly Pro Arg
Arg Pro Ala Arg Pro Leu 20 25 30Pro Val Arg Ala Ala Ile Asn Asp Ser
Ala His Pro Lys Ala Asn Gly 35 40 45Ser Ala Val Ser Leu Lys Ser Gly
Ser Leu Asn Thr Gln Glu Asp Thr 50 55 60Ser Ser Ser Pro Pro Pro Arg
Thr Phe Leu His Gln Leu Pro Asp Trp65 70 75 80Ser Arg Leu Val Asp
Ser Phe Gly Leu Glu Ser Thr Val Gln Asp Gly 85 90 95Leu Val Phe Arg
Gln Ser Phe Ser Ile Arg Ser Tyr Glu Ile Gly Thr 100 105 110Asp Arg
Thr Ala Ser Ile Glu Thr Leu Met Asn His Leu Gln Glu Thr 115 120
125Ser Leu Asn His Cys Lys Ser Thr Gly Ile Leu Leu Asp Gly Phe Gly
130 135 140Arg Thr Leu Glu Met Cys Lys Arg Asp Leu Ile Trp Val Val
Ile Lys145 150 155 160Met Gln Ile Lys Val Asn Arg Tyr Pro Ala Trp
Gly Asp Thr Val Glu 165 170 175Ile Asn Thr Arg Phe Ser Arg Leu Gly
Lys Ile Gly Met Gly Arg Asp 180 185 190Trp Leu Ile Ser Asp Cys Asn
Thr Gly Glu Ile Leu Val Arg Ala Thr 195 200 205Ser Ala Tyr Ala Met
Met Asn Gln Lys Thr Arg Arg Leu Ser Lys Leu 210 215 220Pro Tyr Glu
Val His Gln Glu Ile Val Pro Leu Phe Val Asp Ser Pro225 230 235
240Val Ile Glu Asp Ser Asp Leu Lys Val His Lys Phe Lys Val Lys Thr
245 250 255Gly Asp Ser Ile Gln Lys Gly Leu Thr Pro Gly Trp Asn Asp
Leu Asp 260 265 270Val Asn Gln His Val Ser Asn Val Lys Tyr Ile Gly
Trp Ile Leu Glu 275 280 285Ser Met Pro Thr Glu Val Leu Glu Thr Gln
Glu Leu Cys Ser Leu Ala 290 295 300Leu Glu Tyr Arg Arg Glu Cys Gly
Arg Asp Ser Val Leu Glu Ser Val305 310 315 320Thr Ala Met Asp Pro
Ser Lys Val Gly Val Arg Ser Gln Tyr Gln His 325 330 335Leu Leu Arg
Leu Glu Asp Gly Thr Ala Ile Val Asn Gly Ala Thr Glu 340 345 350Trp
Arg Pro Lys Asn Ala Gly Ala Asn Gly Ala Ile Ser Thr Gly Lys 355 360
365Thr Ser Asn Gly Asn Ser Val Ser 370 37555338PRTArtificial
SequenceDescription of Artificial Sequence Synthetic polypeptide
55Asn Asp Ser Ala His Pro Lys Ala Asn Gly Ser Ala Val Ser Leu Lys1
5 10 15Ser Gly Ser Leu Asn Thr Gln Glu Asp Thr Ser Ser Ser Pro Pro
Pro 20 25 30Arg Thr Phe Leu His Gln Leu Pro Asp Trp Ser Arg Leu Val
Asp Ser 35 40 45Phe Gly Leu Glu Ser Thr Val Gln Asp Gly Leu Val Phe
Arg Gln Ser 50 55 60Phe Ser Ile Arg Ser Tyr Glu Ile Gly Thr Asp Arg
Thr Ala Ser Ile65 70 75 80Glu Thr Leu Met Asn His Leu Gln Glu Thr
Ser Leu Asn His Cys Lys 85 90 95Ser Thr Gly Ile Leu Leu Asp Gly Phe
Gly Arg Thr Leu Glu Met Cys 100 105 110Lys Arg Asp Leu Ile Trp Val
Val Ile Lys Met Gln Ile Lys Val Asn 115 120 125Arg Tyr Pro Ala Trp
Gly Asp Thr Val Glu Ile Asn Thr Arg Phe Ser 130 135 140Arg Leu Gly
Lys Ile Gly Met Gly Arg Asp Trp Leu Ile Ser Asp Cys145 150 155
160Asn Thr Gly Glu Ile Leu Val Arg Ala Thr Ser Ala Tyr Ala Met Met
165 170 175Asn Gln Lys Thr Arg Arg Leu Ser Lys Leu Pro Tyr Glu Val
His Gln 180 185 190Glu Ile Val Pro Leu Phe Val Asp Ser Pro Val Ile
Glu Asp Ser Asp 195 200 205Leu Lys Val His Lys Phe Lys Val Lys Thr
Gly Asp Ser Ile Gln Lys 210 215 220Gly Leu Thr Pro Gly Trp Asn Asp
Leu Asp Val Asn Gln His Val Ser225 230 235 240Asn Val Lys Tyr Ile
Gly Trp Ile Leu Glu Ser Met Pro Thr Glu Val 245 250 255Leu Glu Thr
Gln Glu Leu Cys Ser Leu Ala Leu Glu Tyr Arg Arg Glu 260 265 270Cys
Gly Arg Asp Ser Val Leu Glu Ser Val Thr Ala Met Asp Pro Ser 275 280
285Lys Val Gly Val Arg Ser Gln Tyr Gln His Leu Leu Arg Leu Glu Asp
290 295 300Gly Thr Ala Ile Val Asn Gly Ala Thr Glu Trp Arg Pro Lys
Asn Ala305 310 315 320Gly Ala Asn Gly Ala Ile Ser Thr Gly Lys Thr
Ser Asn Gly Asn Ser 325 330 335Val Ser561200DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
56atggccaccg cctccacctt ctccgccttc aacgcccgct gcggcgacct gcgccgctcc
60gccggctccg gcccccgccg ccccgcccgc cccctgcccg tgcgcgccgc catcaacgac
120tccgcccacc ccaaggccaa cggctccgcc gtgagcctga agtccggcag
cctgaacacc 180caggaggaca cctcctccag cccccccccc cgcaccttcc
tgcaccagct gcccgactgg 240agccgcctgg tggacagctt cggcctggag
tccaccgtgc aggacggcct ggtgttccgc 300cagtccttct ccatccgctc
ctacgagatc ggcaccgacc gcaccgccag catcgagacc 360ctgatgaacc
acctgcagga gacctccctg aaccactgca agagcaccgg catcctgctg
420gacggcttcg gccgcaccct ggagatgtgc aagcgcgacc tgatctgggt
ggtgatcaag 480atgcagatca aggtgaaccg ctaccccgcc tggggcgaca
ccgtggagat caacacccgc 540ttcagccgcc tgggcaagat cggcatgggc
cgcgactggc tgatctccga ctgcaacacc 600ggcgagatcc tggtgcgcgc
caccagcgcc tacgccatga tgaaccagaa gacccgccgc 660ctgtccaagc
tgccctacga ggtgcaccag gagatcgtgc ccctgttcgt ggacagcccc
720gtgatcgagg actccgacct gaaggtgcac aagttcaagg tgaagaccgg
cgacagcatc 780cagaagggcc tgacccccgg ctggaacgac ctggacgtga
accagcacgt gtccaacgtg 840aagtacatcg gctggatcct ggagagcatg
cccaccgagg tgctggagac ccaggagctg 900tgctccctgg ccctggagta
ccgccgcgag tgcggccgcg actccgtgct ggagagcgtg 960accgccatgg
accccagcaa ggtgggcgtg cgctcccagt accagcacct gctgcgcctg
1020gaggacggca ccgccatcgt gaacggcgcc accgagtggc gccccaagaa
cgccggcgcc 1080aacggcgcca tctccaccgg caagaccagc aacggcaact
ccgtgtccat ggactacaag 1140gaccacgacg gcgactacaa ggaccacgac
atcgactaca aggacgacga cgacaagtga
1200576046DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 57caccggcgcg ctgcttcgcg tgccgggtgc
agcaatcaga tccaagtctg acgacttgcg 60cgcacgcgcc ggatccttca attccaaagt
gtcgtccgcg tgcgcttctt cgccttcgtc 120ctcttgaaca tccagcgacg
caagcgcagg gcgctgggcg gctggcgtcc cgaaccggcc 180tcggcgcacg
cggctgaaat tgccgatgtc ggcaatgtag tgccgctccg cccacctctc
240aattaagttt ttcagcgcgt ggttgggaat gatctgcgct catggggcga
aagaaggggt 300tcagaggtgc tttattgtta ctcgactggg cgtaccagca
ttcgtgcatg actgattata 360catacaaaag tacagctcgc ttcaatgccc
tgcgattcct actcccgagc gagcactcct 420ctcaccgtcg ggttgcttcc
cacgaccacg ccggtaagag ggtctgtggc ctcgcgcccc 480tcgcgagcgc
atctttccag ccacgtctgt atgattttgc gctcatacgt ctggcccgtc
540gaccccaaaa tgacgggatc ctgcataata tcgcccgaaa tgggatccag
gcattcgtca 600ggaggcgtca gccccgcggg agatgccggt cccgccgcat
tggaaaggtg tagagggggt 660gaatccccca tttcatgaaa tgggtacccc
gctcccgtct ggtcctcacg ttcgtgtacg 720gcctggatcc cggaaagggc
ggatgcacgt ggtgttgccc cgccattggc gcccacgttt 780caaagtcccc
ggccagaaat gcacaggacc ggcccggctc gcacaggcca tgacgaatgc
840ccagatttcg acagcaaaac aatctggaat aatcgcaacc attcgcgttt
tgaacgaaac 900gaaaagacgc tgtttagcac gtttccgata tcgtgggggc
cgaagcatga ttggggggag 960gaaagcgtgg ccccaaggta gcccattctg
tgccacacgc cgacgaggac caatccccgg 1020catcagcctt catcgacggc
tgcgccgcac atataaagcc ggacgccttc ccgacacgtt 1080caaacagttt
tatttcctcc acttcctgaa tcaaacaaat cttcaaggaa gatcctgctc
1140ttgagcaact cgtatgttcg cgttctactt cctgacggcc tgcatctccc
tgaagggcgt 1200gttcggcgtc tccccctcct acaacggcct gggcctgacg
ccccagatgg gctgggacaa 1260ctggaacacg ttcgcctgcg acgtctccga
gcagctgctg ctggacacgg ccgaccgcat 1320ctccgacctg ggcctgaagg
acatgggcta caagtacatc atcctggacg actgctggtc 1380ctccggccgc
gactccgacg gcttcctggt cgccgacgag cagaagttcc ccaacggcat
1440gggccacgtc gccgaccacc tgcacaacaa ctccttcctg ttcggcatgt
actcctccgc 1500gggcgagtac acgtgcgccg gctaccccgg ctccctgggc
cgcgaggagg aggacgccca 1560gttcttcgcg aacaaccgcg tggactacct
gaagtacgac aactgctaca acaagggcca 1620gttcggcacg cccgagatct
cctaccaccg ctacaaggcc atgtccgacg ccctgaacaa 1680gacgggccgc
cccatcttct actccctgtg caactggggc caggacctga ccttctactg
1740gggctccggc atcgcgaact cctggcgcat gtccggcgac gtcacggcgg
agttcacgcg 1800ccccgactcc cgctgcccct gcgacggcga cgagtacgac
tgcaagtacg ccggcttcca 1860ctgctccatc atgaacatcc tgaacaaggc
cgcccccatg ggccagaacg cgggcgtcgg 1920cggctggaac gacctggaca
acctggaggt cggcgtcggc aacctgacgg acgacgagga 1980gaaggcgcac
ttctccatgt gggccatggt gaagtccccc ctgatcatcg gcgcgaacgt
2040gaacaacctg aaggcctcct cctactccat ctactcccag gcgtccgtca
tcgccatcaa 2100ccaggactcc aacggcatcc ccgccacgcg cgtctggcgc
tactacgtgt ccgacacgga 2160cgagtacggc cagggcgaga tccagatgtg
gtccggcccc ctggacaacg gcgaccaggt 2220cgtggcgctg ctgaacggcg
gctccgtgtc ccgccccatg aacacgaccc tggaggagat 2280cttcttcgac
tccaacctgg gctccaagaa gctgacctcc acctgggaca tctacgacct
2340gtgggcgaac cgcgtcgaca actccacggc gtccgccatc ctgggccgca
acaagaccgc 2400caccggcatc ctgtacaacg ccaccgagca gtcctacaag
gacggcctgt ccaagaacga 2460cacccgcctg ttcggccaga agatcggctc
cctgtccccc aacgcgatcc tgaacacgac 2520cgtccccgcc cacggcatcg
cgttctaccg cctgcgcccc tcctcctgat acaacttatt 2580acgtattctg
accggcgctg atgtggcgcg gacgccgtcg tactctttca gactttactc
2640ttgaggaatt gaacctttct cgcttgctgg catgtaaaca ttggcgcaat
taattgtgtg 2700atgaagaaag ggtggcacaa gatggatcgc gaatgtacga
gatcgacaac gatggtgatt 2760gttatgaggg gccaaacctg gctcaatctt
gtcgcatgtc cggcgcaatg tgatccagcg 2820gcgtgactct cgcaacctgg
tagtgtgtgc gcaccgggtc gctttgatta aaactgatcg 2880cattgccatc
ccgtcaactc acaagcctac tctagctccc attgcgcact cgggcgcccg
2940gctcgatcaa tgttctgagc ggagggcgaa gcgtcaggaa atcgtctcgg
cagctggaag 3000cgcatggaat gcggagcgga gatcgaatca ggatcccgcg
tctcgaacag agcgcgcaga 3060ggaacgctga aggtctcgcc tctgtcgcac
ctcagcgcgg catacaccac aataaccacc 3120tgacgaatgc gcttggttct
tcgtccatta gcgaagcgtc cggttcacac acgtgccacg 3180ttggcgaggt
ggcaggtgac aatgatcggt ggagctgatg gtcgaaacgt tcacagccta
3240gggatatcgt gaaaactcgc tcgaccgccc gcgtcccgca ggcagcgatg
acgtgtgcgt 3300gacctgggtg tttcgtcgaa aggccagcaa ccccaaatcg
caggcgatcc ggagattggg 3360atctgatccg agcttggacc agatccccca
cgatgcggca cgggaactgc atcgactcgg 3420cgcggaaccc agctttcgta
aatgccagat tggtgtccga taccttgatt tgccatcagc 3480gaaacaagac
ttcagcagcg agcgtatttg gcgggcgtgc taccagggtt gcatacattg
3540cccatttctg tctggaccgc tttaccggcg cagagggtga gttgatgggg
ttggcaggca 3600tcgaaacgcg cgtgcatggt gtgtgtgtct gttttcggct
gcacaatttc aatagtcgga 3660tgggcgacgg tagaattggg tgttgcgctc
gcgtgcatgc ctcgccccgt cgggtgtcat 3720gaccgggact ggaatccccc
ctcgcgaccc tcctgctaac gctcccgact ctcccgcccg 3780cgcgcaggat
agactctagt tcaaccaatc gacaactagt atggccaccg catccacttt
3840ctcggcgttc aatgcccgct gcggcgacct gcgtcgctcg gcgggctccg
ggccccggcg 3900cccagcgagg cccctccccg tgcgcgggcg cgcctcccag
ctgcgcaagc ccgccctgga 3960ccccctgcgc gccgtgatct ccgccgacca
gggctccatc tcccccgtga actcctgcac 4020ccccgccgac cgcctgcgcg
ccggccgcct gatggaggac ggctactcct acaaggagaa 4080gttcatcgtg
cgctcctacg aggtgggcat caacaagacc gccaccgtgg agaccatcgc
4140caacctgctg caggaggtgg cctgcaacca cgtgcagaag tgcggcttct
ccaccgacgg 4200cttcgccacc accctgacca tgcgcaagct gcacctgatc
tgggtgaccg cccgcatgca 4260catcgagatc tacaagtacc ccgcctggtc
cgacgtggtg gagatcgaga cctggtgcca 4320gtccgagggc cgcatcggca
cccgccgcga ctggatcctg cgcgactccg ccaccaacga 4380ggtgatcggc
cgcgccacct ccaagtgggt gatgatgaac caggacaccc gccgcctgca
4440gcgcgtgacc gacgaggtgc gcgacgagta cctggtgttc tgcccccgcg
agccccgcct 4500ggccttcccc gaggagaaca actcctccct gaagaagatc
cccaagctgg aggaccccgc 4560ccagtactcc atgctggagc tgaagccccg
ccgcgccgac ctggacatga accagcacgt 4620gaacaacgtg acctacatcg
gctgggtgct ggagtccatc ccccaggaga tcatcgacac 4680ccacgagctg
caggtgatca ccctggacta ccgccgcgag tgccagcagg acgacatcgt
4740ggactccctg accacctccg agatccccga cgaccccatc tccaagttca
ccggcaccaa 4800cggctccgcc atgtcctcca tccagggcca caacgagtcc
cagttcctgc acatgctgcg 4860cctgtccgag aacggccagg agatcaaccg
cggccgcacc cagtggcgca agaagtcctc 4920ccgcatggac tacaaggacc
acgacggcga ctacaaggac cacgacatcg actacaagga 4980cgacgacgac
aagtgaatcg atggagcgac gagtgtgcgt gcggggctgg cgggagtggg
5040acgccctcct cgctcctctc tgttctgaac ggaacaatcg gccaccccgc
gctacgcgcc 5100acgcatcgag caacgaagaa aaccccccga tgataggttg
cggtggctgc cgggatatag 5160atccggccgc acatcaaagg gcccctccgc
cagagaagaa gctcctttcc cagcagactc 5220cttctgctgc caaaacactt
ctctgtccac agcaacacca aaggatgaac agatcaactt 5280gcgtctccgc
gtagcttcct cggctagcgt gcttgcaaca ggtccctgca ctattatctt
5340cctgctttcc tctgaattat gcggcaggcg agcgctcgct ctggcgagcg
ctccttcgcg 5400ccgccctcgc tgatcgagtg tacagtcaat gaatggtgag
ctccgcgcct gcgcgaggac 5460gcagaacaac gctgccgccg tgtcttttgc
acgcgcgact ccggcgcttc gctggtggca 5520cccccataaa gaaaccctca
attctgtttg tggaagacac ggtgtacccc cacccaccca 5580cctgcacctc
tattattggt attattgacg cgggagtggg cgttgtaccc tacaacgtag
5640cttctctagt tttcagctgg ctcccaccat tgtaaattca tgctagaata
gtgcgtggtt 5700atgtgagagg tatagtgtgt ctgagcagac ggggcgggat
gcatgtcgtg gtggtgatct 5760ttggctcaag gcgtcgtcga cgtgacgtgc
ccgatcatga gagcaatacc gcgctcaaag 5820ccgacgcata gcctttactc
cgcaatccaa acgactgtcg ctcgtatttt ttggatatct 5880attttaaaga
gcgagcacag cgccgggcat gggcctgaaa ggcctcgcgg ccgtgctcgt
5940ggtgggggcc gcgagcgcgt ggggcatcgc ggcagtgcac caggcgcaga
cggaggaacg 6000catggtgcgt gcgcaatata agatacatgt attgttgtcc tgcagg
6046581176DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 58atggccaccg catccacttt ctcggcgttc
aatgcccgct gcggcgacct gcgtcgctcg 60gcgggctccg ggccccggcg cccagcgagg
cccctccccg tgcgcgggcg cgcctcccag 120ctgcgcaagc ccgccctgga
ccccctgcgc gccgtgatct ccgccgacca gggctccatc 180tcccccgtga
actcctgcac ccccgccgac cgcctgcgcg ccggccgcct gatggaggac
240ggctactcct acaaggagaa gttcatcgtg cgctcctacg aggtgggcat
caacaagacc 300gccaccgtgg agaccatcgc caacctgctg caggaggtgg
cctgcaacca cgtgcagaag 360tgcggcttct ccaccgccgg cttcgccacc
accctgacca tgcgcaagct gcacctgatc 420tgggtgaccg cccgcatgca
catcgagatc tacaagtacc ccgcctggtc cgacgtggtg 480gagatcgaga
cctggtgcca gtccgagggc cgcatcggca cccgccgcga ctggatcctg
540cgcgactccg ccaccaacga ggtgatcggc cgcgccacct ccaagtgggt
gatgatgaac 600caggacaccc gccgcctgca gcgcgtgacc gacgaggtgc
gcgacgagta cctggtgttc 660tgcccccgcg agccccgcct ggccttcccc
gaggagaaca actcctccct gaagaagatc 720cccaagctgg aggaccccgc
ccagtactcc atgctggagc tgaagccccg ccgcgccgac 780ctggacatga
accagcacgt gaacaacgtg acctacatcg gctgggtgct ggagtccatc
840ccccaggaga tcatcgacac ccacgagctg caggtgatca ccctggacta
ccgccgcgag 900tgccagcagg acgacatcgt ggactccctg accacctccg
agatccccga cgaccccatc 960tccaagttca ccggcaccaa cggctccgcc
atgtcctcca tccagggcca caacgagtcc 1020cagttcctgc acatgctgcg
cctgtccgag aacggccagg agatcaaccg cggccgcacc 1080cagtggcgca
agaagtcctc ccgcatggac tacaaggacc acgacggcga ctacaaggac
1140cacgacatcg actacaagga cgacgacgac aagtga 1176591176DNAArtificial
SequenceDescription of Artificial Sequence Synthetic polynucleotide
59atggccaccg catccacttt ctcggcgttc aatgcccgct gcggcgacct gcgtcgctcg
60gcgggctccg ggccccggcg cccagcgagg cccctccccg tgcgcgggcg cgcctcccag
120ctgcgcaagc ccgccctgga ccccctgcgc gccgtgatct ccgccgacca
gggctccatc 180tcccccgtga actcctgcac ccccgccgac cgcctgcgcg
ccggccgcct gatggaggac 240ggctactcct acaaggagaa gttcatcgtg
cgctcctacg aggtgggcat caacaagacc 300gccaccgtgg agaccatcgc
caacctgctg caggaggtgg cctgcaacca cgtgcagaag 360tgcggcttct
ccaccgacgg cttcgccacc accctgacca tgcgcaagct gcacctgatc
420tgggtgaccg cccgcatgca catcgagatc tacaagtacc ccgcctggtc
cgacgtggtg 480gagatcgaga cctggtgcca gtccgagggc cgcatcggca
cccgccgcga ctggatcctg 540cgcgactccg ccaccaacga ggtgatcggc
cgcgccacct ccaagtgggt gatgatgaac 600caggacaccc gccgcctgca
gcgcgtgacc gccgaggtgc gcgacgagta cctggtgttc 660tgcccccgcg
agccccgcct ggccttcccc gaggagaaca actcctccct gaagaagatc
720cccaagctgg aggaccccgc ccagtactcc atgctggagc tgaagccccg
ccgcgccgac 780ctggacatga accagcacgt gaacaacgtg acctacatcg
gctgggtgct ggagtccatc 840ccccaggaga tcatcgacac ccacgagctg
caggtgatca ccctggacta ccgccgcgag 900tgccagcagg acgacatcgt
ggactccctg accacctccg agatccccga cgaccccatc 960tccaagttca
ccggcaccaa cggctccgcc atgtcctcca tccagggcca caacgagtcc
1020cagttcctgc acatgctgcg cctgtccgag aacggccagg agatcaaccg
cggccgcacc 1080cagtggcgca agaagtcctc ccgcatggac tacaaggacc
acgacggcga ctacaaggac 1140cacgacatcg actacaagga cgacgacgac aagtga
1176601176DNAArtificial SequenceDescription of Artificial Sequence
Synthetic polynucleotide 60atggccaccg catccacttt ctcggcgttc
aatgcccgct gcggcgacct gcgtcgctcg 60gcgggctccg ggccccggcg cccagcgagg
cccctccccg tgcgcgggcg cgcctcccag 120ctgcgcaagc ccgccctgga
ccccctgcgc gccgtgatct ccgccgacca gggctccatc 180tcccccgtga
actcctgcac ccccgccgac cgcctgcgcg ccggccgcct gatggaggac
240ggctactcct acaaggagaa gttcatcgtg cgctcctacg aggtgggcat
caacaagacc 300gccaccgtgg agaccatcgc caacctgctg caggaggtgg
cctgcaacca cgtgcagaag 360tgcggcttct ccaccgccgg cttcgccacc
accctgacca tgcgcaagct gcacctgatc 420tgggtgaccg cccgcatgca
catcgagatc tacaagtacc ccgcctggtc cgacgtggtg 480gagatcgaga
cctggtgcca gtccgagggc cgcatcggca cccgccgcga ctggatcctg
540cgcgactccg ccaccaacga ggtgatcggc cgcgccacct ccaagtgggt
gatgatgaac 600caggacaccc gccgcctgca gcgcgtgacc gccgaggtgc
gcgacgagta cctggtgttc 660tgcccccgcg agccccgcct ggccttcccc
gaggagaaca actcctccct gaagaagatc 720cccaagctgg aggaccccgc
ccagtactcc atgctggagc tgaagccccg ccgcgccgac 780ctggacatga
accagcacgt gaacaacgtg acctacatcg gctgggtgct ggagtccatc
840ccccaggaga tcatcgacac ccacgagctg caggtgatca ccctggacta
ccgccgcgag 900tgccagcagg acgacatcgt ggactccctg accacctccg
agatccccga cgaccccatc 960tccaagttca ccggcaccaa cggctccgcc
atgtcctcca tccagggcca caacgagtcc 1020cagttcctgc acatgctgcg
cctgtccgag aacggccagg agatcaaccg cggccgcacc 1080cagtggcgca
agaagtcctc ccgcatggac tacaaggacc acgacggcga ctacaaggac
1140cacgacatcg actacaagga cgacgacgac aagtga 117661368PRTBrassica
napus 61Met Ala Thr Ala Ser Thr Phe Ser Ala Phe Asn Ala Arg Cys Gly
Asp1 5 10 15Leu Arg Arg Ser Ala Gly Ser Gly Pro Arg Arg Pro Ala Arg
Pro Leu 20 25 30Pro Val Arg Gly Arg Ala Ser Gln Leu Arg Lys Pro Ala
Leu Asp Pro 35 40 45Leu Arg Ala Val Ile Ser Ala Asp Gln Gly Ser Ile
Ser Pro Val Asn 50 55 60Ser Cys Thr Pro Ala Asp Arg Leu Arg Ala Gly
Arg Leu Met Glu Asp65 70 75 80Gly Tyr Ser Tyr Lys Glu Lys Phe Ile
Val Arg Ser Tyr Glu Val Gly 85 90 95Ile Asn Lys Thr Ala Thr Val Glu
Thr Ile Ala Asn Leu Leu Gln Glu 100 105 110Val Ala Cys Asn His Val
Gln Lys Cys Gly Phe Ser Thr Asp Gly Phe 115 120 125Ala Thr Thr Leu
Thr Met Arg Lys Leu His Leu Ile Trp Val Thr Ala 130 135 140Arg Met
His Ile Glu Ile Tyr Lys Tyr Pro Ala Trp Ser Asp Val Val145 150 155
160Glu Ile Glu Thr Trp Cys Gln Ser Glu Gly Arg Ile Gly Thr Arg Arg
165 170 175Asp Trp Ile Leu Arg Asp Ser Ala Thr Asn Glu Val Ile Gly
Arg Ala 180 185 190Thr Ser Lys Trp Val Met Met Asn Gln Asp Thr Arg
Arg Leu Gln Arg 195 200 205Val Thr Asp Glu Val Arg Asp Glu Tyr Leu
Val Phe Cys Pro Arg Glu 210 215 220Pro Arg Leu Ala Phe Pro Glu Glu
Asn Asn Ser Ser Leu Lys Lys Ile225 230 235 240Pro Lys Leu Glu Asp
Pro Ala Gln Tyr Ser Met Leu Glu Leu Lys Pro 245 250 255Arg Arg Ala
Asp Leu Asp Met Asn Gln His Val Asn Asn Val Thr Tyr 260 265 270Ile
Gly Trp Val Leu Glu Ser Ile Pro Gln Glu Ile Ile Asp Thr His 275 280
285Glu Leu Gln Val Ile Thr Leu Asp Tyr Arg Arg Glu Cys Gln Gln Asp
290 295 300Asp Ile Val Asp Ser Leu Thr Thr Ser Glu Ile Pro Asp Asp
Pro Ile305 310 315 320Ser Lys Phe Thr Gly Thr Asn Gly Ser Ala Met
Ser Ser Ile Gln Gly 325 330 335His Asn Glu Ser Gln Phe Leu His Met
Leu Arg Leu Ser Glu Asn Gly 340 345 350Gln Glu Ile Asn Arg Gly Arg
Thr Gln Trp Arg Lys Lys Ser Ser Arg 355 360
365624PRTUnknownDescription of Unknown hydrophobic domain motif
62Leu Pro Asp Trp1

















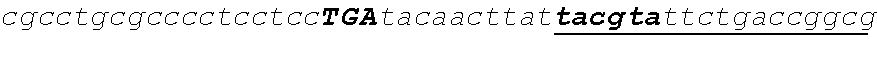




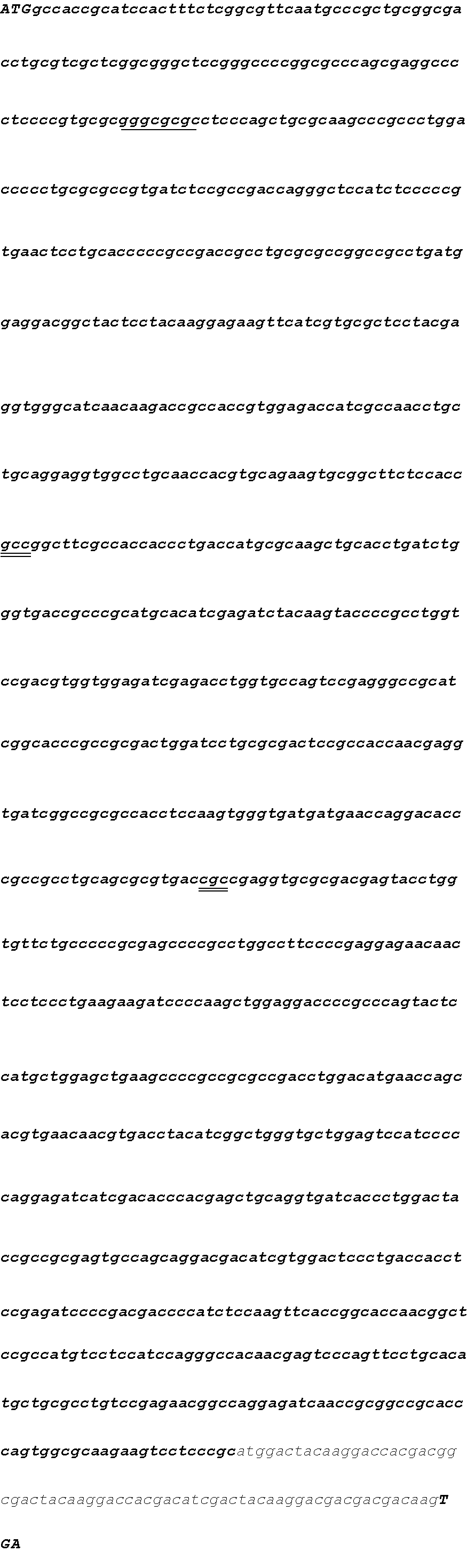
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.