Novel Guide Rna/cas Endonuclease Systems
CIGAN; ANDREW MARK ; et al.
U.S. patent application number 15/573953 was filed with the patent office on 2019-05-09 for novel guide rna/cas endonuclease systems. This patent application is currently assigned to PIONEER HI-BRED INTERNATIONAL, INC.. The applicant listed for this patent is PIONEER HI-BRED INTERNATIONAL, INC.. Invention is credited to ANDREW MARK CIGAN, GIEDRIUS GASIUNAS, TAUTVYDAS KARVELIS, VIRGINIJUS SIKSNYS, JOSHUA K YOUNG.
| Application Number | 20190136248 15/573953 |
| Document ID | / |
| Family ID | 56072462 |
| Filed Date | 2019-05-09 |











View All Diagrams
| United States Patent Application | 20190136248 |
| Kind Code | A1 |
| CIGAN; ANDREW MARK ; et al. | May 9, 2019 |
NOVEL GUIDE RNA/CAS ENDONUCLEASE SYSTEMS
Abstract
Compositions and methods are provided for novel guide RNA/Cas endonuclease systems. Type II Cas9 endonuclease systems originating from Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides are described herein. The present disclosure also describes methods for genome modification of a target sequence in the genome of a cell, for gene editing, and for inserting a polynucleotide of interest into the genome of a cell.
| Inventors: | CIGAN; ANDREW MARK; (MADISON, WI) ; GASIUNAS; GIEDRIUS; (VILNIUS, LT) ; KARVELIS; TAUTVYDAS; (VILNIUS, LT) ; SIKSNYS; VIRGINIJUS; (VILNIUS, LT) ; YOUNG; JOSHUA K; (JOHNSTON, IA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | PIONEER HI-BRED INTERNATIONAL,
INC. JOHNSTON IA |
||||||||||
| Family ID: | 56072462 | ||||||||||
| Appl. No.: | 15/573953 | ||||||||||
| Filed: | May 12, 2016 | ||||||||||
| PCT Filed: | May 12, 2016 | ||||||||||
| PCT NO: | PCT/US2016/032073 | ||||||||||
| 371 Date: | November 14, 2017 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 62162377 | May 15, 2015 | |||
| 62162353 | May 15, 2015 | |||
| 62196535 | Jul 24, 2015 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 15/102 20130101; C12Q 1/6811 20130101; C12N 9/22 20130101; C12N 15/902 20130101; C12N 2800/80 20130101; C12N 15/8213 20130101; C12N 2320/10 20130101; C12N 15/113 20130101; C12N 2310/20 20170501; C12N 15/11 20130101; C12N 15/1093 20130101; C40B 20/04 20130101; C40B 40/06 20130101; C12N 15/1051 20130101; C12Q 1/6811 20130101; C12Q 2521/301 20130101; C12Q 2525/179 20130101; C12Q 2525/191 20130101; C12Q 2535/122 20130101 |
| International Class: | C12N 15/82 20060101 C12N015/82; C12N 15/11 20060101 C12N015/11; C12N 9/22 20060101 C12N009/22 |
Claims
1. A single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139.
2. A single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA comprises a chimeric non-naturally occurring crRNA linked to a tracrRNA, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184.
3. A single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA comprises a chimeric non-naturally occurring crRNA linked to a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160.
4. A guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence.
5. A guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence.
6. A guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence.
7. A guide RNA/Cas9 endonuclease complex comprising a Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, and at least one guide RNA, wherein said guide RNA/Cas9 endonuclease complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a target sequence.
8. A guide RNA/Cas9 endonuclease complex comprising at least one guide RNA and a Cas9 endonuclease, wherein said Cas9 endonuclease is encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, and 80, wherein said guide RNA/Cas9 endonuclease complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a target sequence.
9. The guide RNA/Cas9 endonuclease complex of claim 7, wherein said guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139.
10. The guide RNA/Cas9 endonuclease complex of claim 7, wherein said target sequence is located in the genome of a cell.
11. A method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, and at least one guide RNA, wherein said guide RNA and Cas9 endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site.
12. The method of claim 10, further comprising identifying at least one cell that has a modification at said target, wherein the modification at said target site is selected from the group consisting of (i) a replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, and (iv) any combination of (i)-(iii).
13. A method for editing a nucleotide sequence in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, a polynucleotide modification template, and at least one guide RNA, wherein said polynucleotide modification template comprises at least one nucleotide modification of said nucleotide sequence, wherein said guide RNA and Cas9 endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site.
14. A method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one guide RNA, at least one donor DNA, and at least one Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, wherein said at least one guide RNA and at least one Cas9 endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site, wherein said donor DNA comprises a polynucleotide of interest.
15. The method of claim 11, 13 or 14, wherein said guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139.
16. The method of claim 13, further comprising identifying at least one cell that said polynucleotide of interest integrated in or near said target site.
17. The method of any one of claims 10-14, wherein the cell is selected from the group consisting of a human, non-human, animal, bacterial, fungal, insect, yeast, non-conventional yeast, and plant cell.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is the 371 national stage entry of International Application Number PCT/US2016/032073, filed on 12 May 2016, which claims the benefit of U.S. Provisional Application No. 62/162,377, filed May 15, 2015, U.S. Provisional Application No. 62/162,353, filed May 15, 2015 and U.S. Provisional Application No. 62/196,535, filed Jul. 24, 2015, each of which is incorporated herein in their entirety by reference.
FIELD
[0002] The disclosure relates to the field of plant molecular biology, in particular, to compositions for novel guide RNA/Cas endonuclease systems and compositions and methods for altering the genome of a cell.
REFERENCE TO SEQUENCE LISTING SUBMITTED ELECTRONICALLY
[0003] The official copy of the sequence listing is submitted electronically via EFS-Web as an ASCII formatted sequence listing with a file named 20160502_BB2539PCT_SequenceListing.txt created on May 2, 2016 and having a size 236 kilobytes and is filed concurrently with the specification. The sequence listing contained in this ASCII formatted document is part of the specification and is herein incorporated by reference in its entirety.
BACKGROUND
[0004] Recombinant DNA technology has made it possible to insert DNA sequences at targeted genomic locations and/or modify (edit) specific endogenous chromosomal sequences, thus altering the organism's phenotype. Site-specific integration techniques, which employ site-specific recombination systems, as well as other types of recombination technologies, have been used to generate targeted insertions of genes of interest in a variety of organism. Genome-editing techniques such as designer zinc finger nucleases (ZFNs) or transcription activator-like effector nucleases (TALENs), or homing meganucleases, are available for producing targeted genome perturbations, but these systems tends to have a low specificity and employ designed nucleases that need to be redesigned for each target site, which renders them costly and time-consuming to prepare.
[0005] Although several approaches have been developed to target a specific site for modification in the genome of an organism, there still remains a need for new genome engineering technologies that are affordable, easy to set up, scalable, and amenable to targeting multiple positions within the genome of an organism
BRIEF SUMMARY
[0006] Compositions and methods are provided for rapid characterization of novel Cas endonuclease systems and the elements comprising such a systems, including, but not limiting to, rapid characterization of PAM sequences, guide RNA elements and CAS endonucleases.
[0007] In one embodiment of the disclosure, the guide RNA is a guide RNAs capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence, wherein said tracrRNA is originated from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755.
[0008] In another embodiment of the disclosure, the guide RNA is a guide RNA capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a single molecule comprising a chimeric non-naturally occurring crRNA linked to a tracrRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence.
[0009] In another embodiment of the disclosure, the guide RNA is a guide RNA capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises at least a fragment of a crRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence.
[0010] In another embodiment of the disclosure, the guide RNA is a guide RNA capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a single molecule comprising a tracrRNA linked to a chimeric non-naturally occurring crRNA comprising at least a fragment of a crRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence.
[0011] Also provided are nucleic acid constructs, plants, plant cells, explants, seeds and grain having an altered target site or altered polynucleotide of interest produced by the methods described herein. Additional embodiments of the methods and compositions of the present disclosure are shown herein.
BRIEF DESCRIPTION OF THE DRAWINGS AND THE SEQUENCE LISTING
[0012] The disclosure can be more fully understood from the following detailed description and the accompanying drawings and Sequence Listing, which form a part of this application. The sequence descriptions and sequence listing attached hereto comply with the rules governing nucleotide and amino acid sequence disclosures in patent applications as set forth in 37 C.F.R. .sctn..sctn. 1.821-1.825. The sequence descriptions contain the three letter codes for amino acids as defined in 37 C.F.R. .sctn..sctn. 1.821-1.825, which are incorporated herein by reference.
FIGURES
[0013] FIG. 1 shows a diagram of the formation of a full Oligoduplex II comprising a restriction enzyme recognition site (RE1), a target sequence and a randomized Protospacer-Adjacent-Motif (PAM) sequence.
[0014] FIG. 2 show a diagram of the design and construction of a 5 nucleotide (5N) randomized Protospacer-Adjacent-Motif (PAM) plasmid library and host cell library. RE1=restriction endonuclease 1, RE2=restriction endonuclease 2.
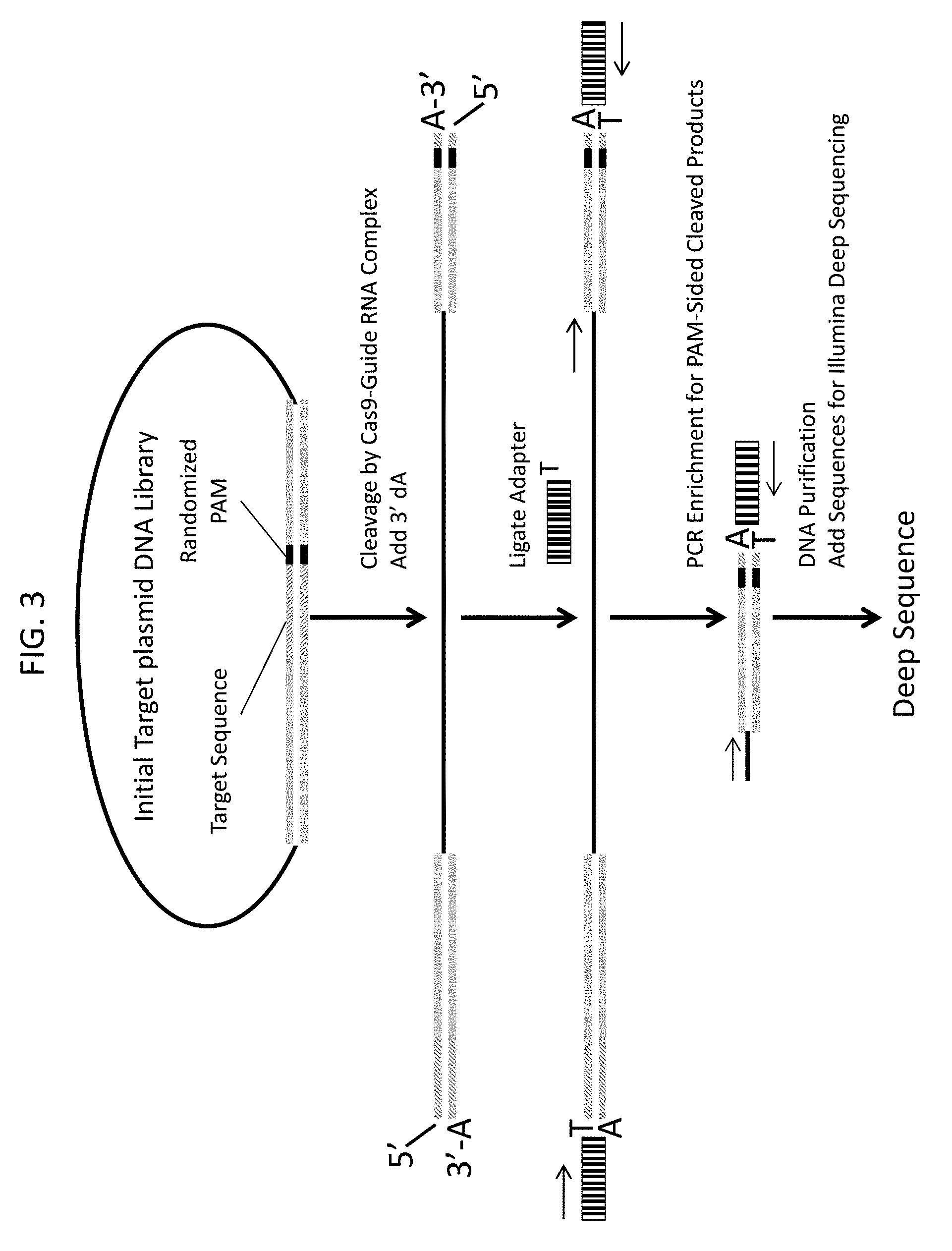
[0015] FIG. 3 shows a diagram of the production of enriched PAM sided products for deep sequencing and identification of PAM preferences.
[0016] FIG. 4 depicts the PAM sequence distribution from a 5 nucleotide (5N) randomized Protospacer-Adjacent-Motif (PAM) plasmid library.
[0017] FIG. 5 shows the PAM preferences (NGGNG) for Streptococcus thermophilus CRISPR3 (Sth3) Cas9 endonuclease in both 50 nM and 100 nM digests.
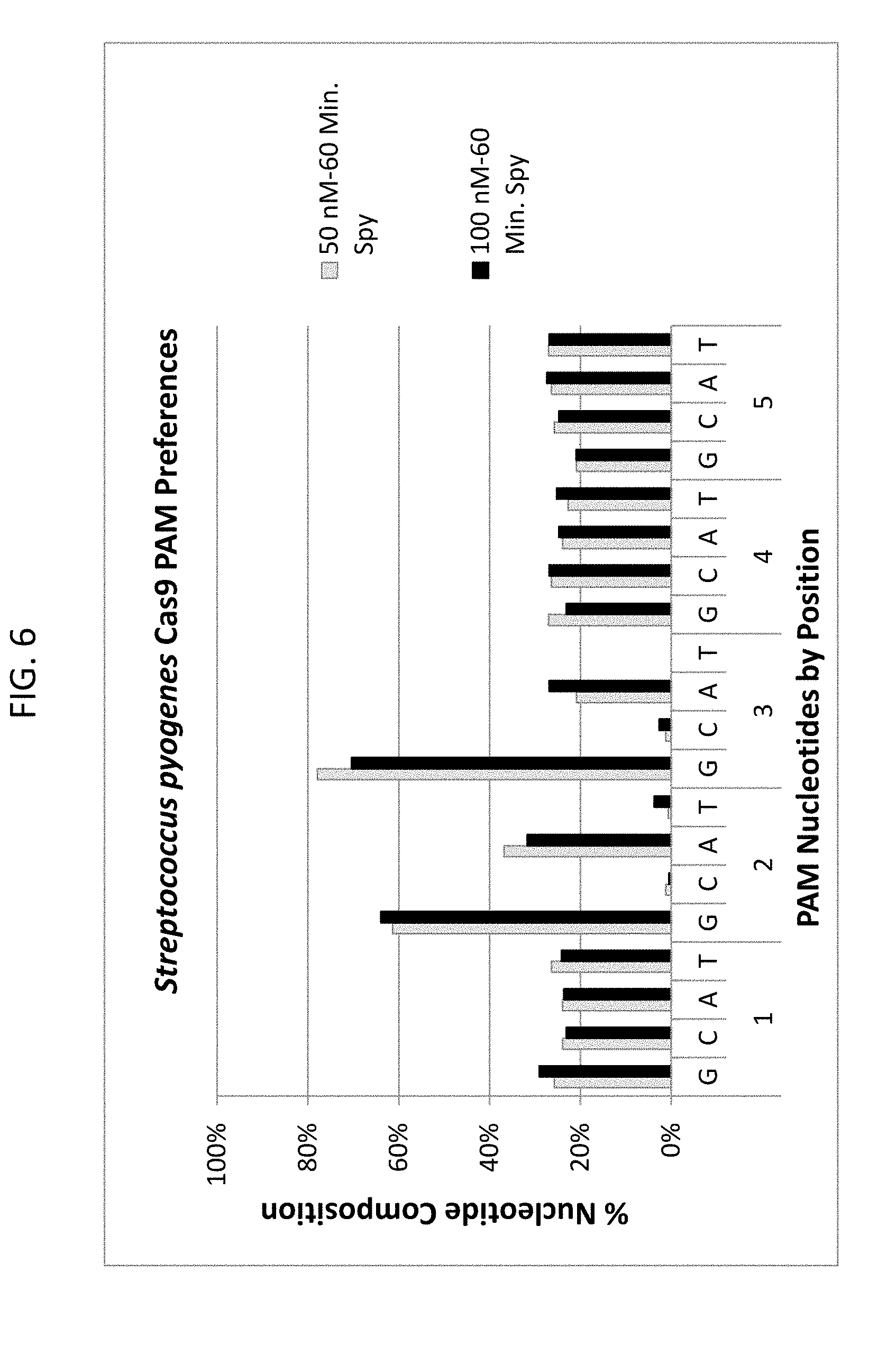
[0018] FIG. 6 shows the PAM preferences (NGG) for Streptococcus pyogenes (Spy) Cas9 endonuclease in both 50 nM and 100 nM digests.
[0019] FIG. 7 shows the effect of decreasing Sth3 and Spy Cas9-crRNA-tracrRNA complex concentration and digestion time to determine the minimal Sth3 and Spy Cas9 concentration and shortest digestion time where PCR amplified cleavage products may still be obtained from the randomized PAM plasmid library.
[0020] FIG. 8 shows the PAM preferences (NGGNG) for Streptococcus thermophilus CRISPR3 (Sth3) Cas9 endonuclease positive controls in both 50 nM and 100 nM digests.
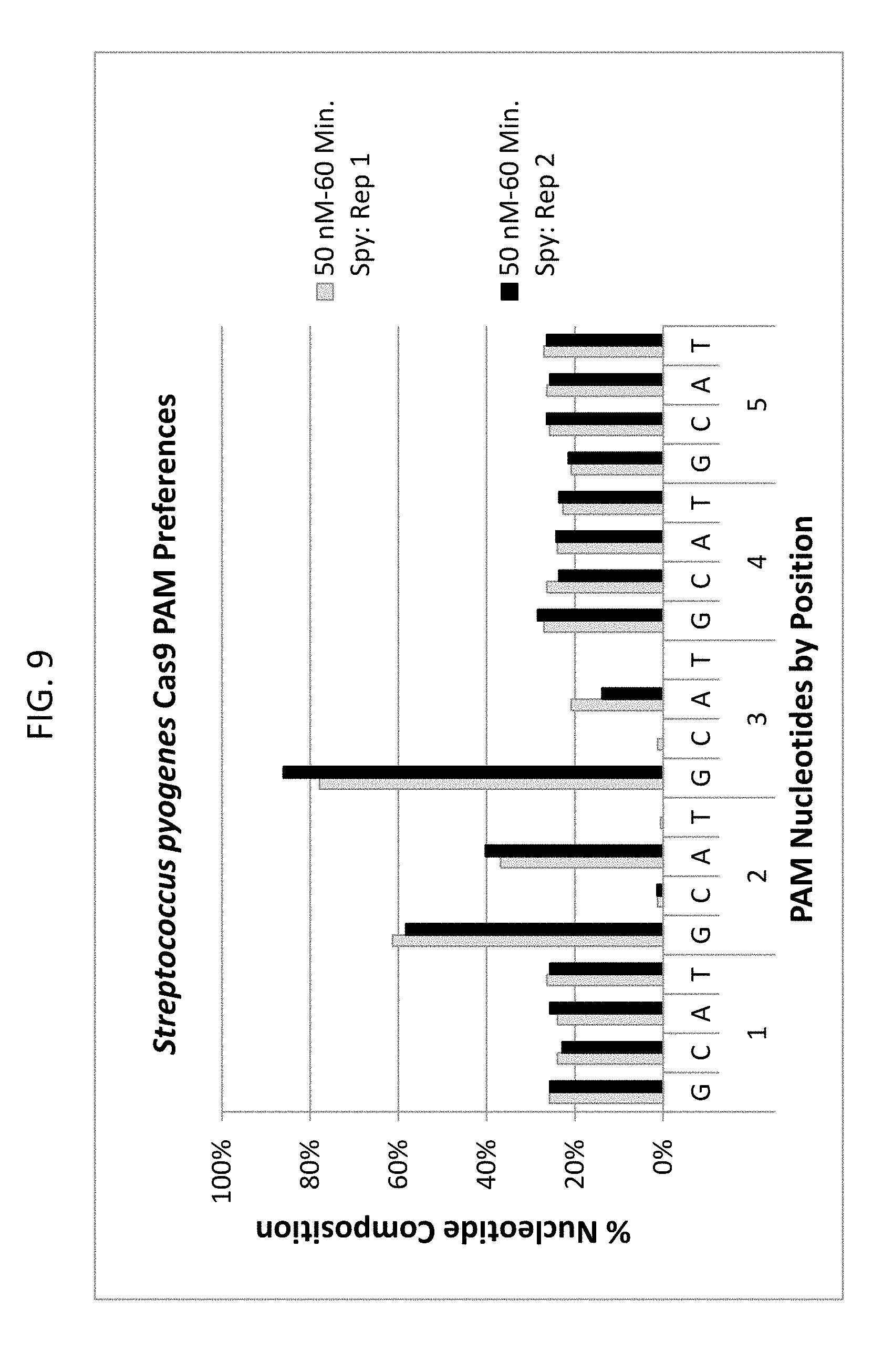
[0021] FIG. 9 shows the PAM preferences (NGG) for Streptococcus pyogenes (Spy) Cas9 endonuclease positive controls in both 50 nM and 100 nM digests.
[0022] FIG. 10 shown the PAM preferences (NGGNG) observed in the minimally Streptococcus thermophilus Sth3 digested libraries (0.5 nM-60 min and 50 nM-1 min) compared to that exhibited by the respective 50 nM-60 minute positive control.
[0023] FIG. 11 shown the PAM preferences (NGGNG) observed in the minimally Streptococcus pyogenes Spy digested libraries (0.5 nM-60 min and 50 nM-1 min) compared to that exhibited by the respective 50 nM-60 minute positive control.
[0024] FIG. 12 shows the PAM preferences for Streptococcus pyogenes (Spy) Cas9 endonuclease guided by a single guide RNA (sgRNA) or guided by a crRNA:tracrRNA duplex. The NGGNG PAM preference is nearly identical regardless of the type of guide RNA used
[0025] FIG. 13 shows the PAM preferences (NGG) for Streptococcus pyogenes (Spy) Cas9 endonuclease guided by a single guide RNA (sgRNA) or guided by a crRNA:tracrRNA duplex. The NGG PAM preference is nearly identical regardless of the type of guide RNA used
[0026] FIG. 14 shows the PAM preferences for Streptococcus thermophilus CRISPR3 (Sth3) Cas9 endonuclease positive controls for comparing of a 5N randomized PAM plasmid DNA library and a 7N randomized PAM plasmid DNA library.
[0027] FIG. 15 shows the PAM preferences (NGG) for Streptococcus pyogenes (Spy) Cas9 endonuclease positive controls for comparing of a 5N randomized PAM plasmid DNA library and a 7N randomized PAM plasmid DNA library.
[0028] FIG. 16 shows the PAM preferences (NNAGAAW) for Streptococcus thermophilus CRISPR1 (Sth1) Cas9 endonuclease in both 50 nM and 0.5 nM nM digests.
[0029] FIG. 17-A shows a genomic DNA region from, Brevibacillus laterosporus representing the Type II CRISPR-Cas system described herein. FIG. 17-B list 8 repeat sequences (SEQ ID NOs:37-44) of the genomic DNA region from the Brevibacillus laterosporus.
[0030] FIG. 18 shows a diagram of the "direct" scenario and the "reverse" scenario of the tracrRNA and CRISPR array to determine a guide RNA for the Cas9 protein identified from the Brevibacillus laterosporus (Blat).
[0031] FIG. 19 shows the secondary structure of the "direct" tracrRNA region downstream of the anti-repeat (SEQ ID NO: 68) from, Brevibacillus laterosporus.
[0032] FIG. 20 shows the secondary structure of the "reverse" tracrRNA region downstream of the anti-repeat (SEQ ID NO: 69) from, Brevibacillus laterosporus.
[0033] FIG. 21 shown an agarose gel with reaction products, indicating that only the "direct" sgRNA (dirsgRNA), but not the "reverse" sgRNA (revsgRNA) supported plasmid library cleavage in combination with a Cas9 endonuclease originating from Brevibacillus laterosporus. (BlatCas9).
[0034] FIG. 22 shows the effect of decreasing BlatCas9 concentration and digestion time to determine the minimal Blast Cas9 concentration and shortest digestion time where PCR amplified cleavage products may still be obtained from the randomized PAM plasmid library.
[0035] FIG. 23 shows the PAM preferences (NNNNCND) for Brevibacillus laterosporus (Blat) Cas9 endonuclease in both 50 nM and 0.5 nM digests.
[0036] FIG. 24 depict sequencing results indicating that plasmid DNA cleavage occurred in the protospacer 3 bp away from the PAM sequence.
[0037] FIG. 25 shows a genomic DNA region from Lactobacillus reuteri MIc3 representing an example of a Type II CRISPR-Cas system described herein.
[0038] FIG. 26 shows a genomic DNA region from Lactobacillus rossiae DSM 15814 representing an example of a Type II CRISPR-Cas system described herein.
[0039] FIG. 27 shows a genomic DNA region from Pediococcus pentosaceus SL4 representing an example of a Type II CRISPR-Cas system described herein.
[0040] FIG. 28 shows a genomic DNA region from Lactobacillus nodensis JCM 14932 representing an example of a Type II CRISPR-Cas system described herein.
[0041] FIG. 29 shows a genomic DNA region from Sulfurospirillum sp. SCADC representing an example of a Type II CRISPR-Cas system described herein.
[0042] FIG. 30 shows a genomic DNA region from Bifidobacterium thermophilum DSM 20210 representing an example of a Type II CRISPR-Cas system described herein.
[0043] FIG. 31 shows a genomic DNA region from Loktanella vestfoldensis representing an example of a Type II CRISPR-Cas system described herein.
[0044] FIG. 32 shows a genomic DNA region from Sphingomonas sanxanigenens NX02 representing an example of a Type II CRISPR-Cas system described herein.
[0045] FIG. 33 shows a genomic DNA region from Epilithonimonas tenax DSM 16811 representing an example of a Type II CRISPR-Cas system described herein.
[0046] FIG. 34 shows a genomic DNA region from Sporocytophaga myxococcoides representing an example of a Type II CRISPR-Cas system described herein.
[0047] FIG. 35 shows a genomic DNA region from Psychroflexus torquis ATCC 700755 representing an example of a Type II CRISPR-Cas system described herein.
[0048] FIG. 36 Bifidobacterium thermophilum (Bthe) Cas9 non-homologous end-joining (NHEJ) mutation frequencies with different single guide RNA (sgRNA) variable targeting domain (spacer) lengths (20 nt, 25 nt and 29 nt) at 2 maize target sites. NHEJ mutations were detected by deep sequencing 2 days after transformation.
SEQUENCES
TABLE-US-00001 [0049] TABLE 1 Summary of Nucleic Acid and Protein SEQ ID Numbers Nucleic acid Protein Description SEQ ID NO. SEQ ID NO. Target sequence T1 1 (80 bases) Single oligonucleotide GG-821N 2 (47 bases) Oligonucleotide GG-820 3 (44 bases) TK-119 primer 4 (22 bases) pUC-dir primer 5 (22 bases) JKYS800.1 forward primer 6 (59 bases) JKYS803 reverse primer 7 (53 bases) Universal Forward primer 8 (43 bases) Universal Reverse primer 9 (18 bases) Sth1-dir primer 10 (34 bases) Sth1-rev primer 11 (27 bases) Sth3-dir primer 12 (26 bases) Sth3-rev primer 13 (30 bases) Spy-dir primer 14 (38 bases) Spy-rev primer 15 (32 bases) Streptococcus thermophilus (Sth3) crRNA 16 (42 bases) Streptococcus thermophilus (Sth3) tracrRNA 17 (78 bases) Streptococcus pyogenes (Spy) crRNA 18 (42 bases) Streptococcus pyogenes (Spy) tracrRNA 19 (78 bases) TK-117 20 (31 bases) TK-111 21 (30 bases) JKYS807.1 primer 22 (56 bases) JKYS807.2 primer 23 (56 bases) JKYS807.3 primer 24 (56 bases) JKYS807.4 primer 25 (56 bases) Sth3 sgRNA 26 (123 bases) Spy sgRNA 27 (105 bases) GG-940-G oligonucleotide 28 (59 bases) GG-940-C oligonucleotide 29 (59 bases) GG-940-A oligonucleotide 30 (59 bases) GG-940-T oligonucleotide 31 (59 bases) JKYS812 32 (49 bases) Streptococcus thermophilus CRISPR1 (Sth1) crRNA 33 (42 bases) Streptococcus thermophilus CRISPR1 Sth1 tracrRNA 34 (80 bases) Streptococcus thermophilus CRISPR3 (Sth3) Cas9 35 (1388 aa) Cas9 single long open-reading-frame from the 36 (3279 bases) Brevibacillus laterosporus bacterial strain SSP360D4 Repeat 1, Brevibacillus laterosporus SSP360D4 37 (36 bases) Repeat 2, Brevibacillus laterosporus SSP360D4 38 (36 bases) Repeat 3, Brevibacillus laterosporus SSP360D4 39 (36 bases) Repeat 4, Brevibacillus laterosporus SSP360D4 40 (36 bases) Repeat 5, Brevibacillus laterosporus SSP360D4 41 (36 bases) Repeat 6, Brevibacillus laterosporus SSP360D4 42 (36 bases) Repeat 7, Brevibacillus laterosporus SSP360D4 43 (36 bases) Repeat 8, Brevibacillus laterosporus SSP360D4 44 (36 bases) Blat-Cas9-dir 45 (29 bases) Blat-Cas9-rev 46 35 bases) Blat sgRNA Direct 47 (177 bases) Blat sgRNA Reverse 48 (118 bases) GG-969 oligonucleotide 49 (68 bases) GG-839 oligonucleotide 50 (62 bases) TK-149 51 55 bases) TK-150 52 (62 bases) GG-840 53 (71 bases) GG-841 54 (75 bases) TK-124 55 (37 bases) TK-151 56 (26 bases) TK-126; 57 (32 bases) GG-935 58 (37 bases) GG-936 59 (45 bases) pUC-EheD primer 60 (21 bases) pUC-LguR primer 61 (22 bases) Sense DNA Strand of Cleaved Sequencing 62 (21 bases) Template Anti-Sense DNA Strand Sequencing Read 63 (11 bases) Anti-Sense DNA Strand of Cleaved Sequencing 64 (21 bases) Template Sense DNA Strand of DNA Sequencing Read 65 (11 bases) Sense DNA Strand of Target and PAM 66 (27 bases) Anti-Sense DNA Strand of Target and PAM 67 (27 bases) "Direct" tracrRNA region downstream of the 68 (118 bases) anti-repeat "Reverse" tracrRNA region downstream of the 69 (58 bases) anti-repeat Lactobacillus reuteri Mlc3 (Lreu) Cas9 Open 70 (4107 bases) Reading Frame Lactobacillus rossiae DSM 15814 (Lros) Cas9 71 (4110 bases) Open Reading Frame Pediococcus pentosaceus SL4 (Ppen) Cas9 72 (4041 bases) Open Reading Frame Lactobacillus nodensis JCM 14932 (Lnod) 73 (3393 bases) Cas9 Open Reading Frame Sulfurospirillum sp. SCADC (Sspe) Cas9 Open 74 (4086 bases) Reading Frame Bifidobacterium thermophilum DSM 20210 75 (3444 bases) (Bthe) Cas9 Open Reading Frame Loktanella vestfoldensis (Lves) Cas9 Open 76 (3216 bases) Reading Frame Sphingomonas sanxanigenens NX02 (Ssan) 77 (3318 bases) Cas9 Open Reading Frame Epilithonimonas tenax DSM 16811 (Eten) Cas9 78 (4200 bases) Open Reading Frame Sporocytophaga myxococcoides (Smyx) Cas9 79 (4362 bases) Open Reading Frame Psychroflexus torquis ATCC 700755 (Ptor) 80 (4530 bases) Cas9 Open Reading Frame Lreu Cas9 Endonuclease 81 (1368 aa) Lros Cas9 Endonuclease 82 (1369 aa) Ppen Cas9 Endonuclease 83 (1346 aa) Lnod Cas9 Endonuclease 84 (1130 aa) Sspe Cas9 Endonuclease 85 (1361 aa) Bthe Cas9 Endonuclease 86 (1147 aa) Lves Cas9 Endonuclease 87 (1071 aa) Ssan Cas9 Endonuclease 88 (1105 aa) Eten Cas9 Endonuclease 89 (1399 aa) Smyx Cas9 Endonuclease 90 (1453 aa) Ptor Cas9 Endonuclease 91 (1509 aa) Lreu CRISPR Repeat Consensus 92 (36 bases) Lros CRISPR Repeat Consensus 93 (36 bases) Ppen CRISPR Repeat Consensus 94 (36 bases) Lnod CRISPR Repeat Consensus 95 (36 bases) Sspe CRISPR Repeat Consensus 96 (36 bases) Bthe CRISPR Repeat Consensus 97 (36 bases) Lves CRISPR Repeat Consensus 98 (36 bases) Ssan CRISPR Repeat Consensus 99 (36 bases) Eten CRISPR Repeat Consensus 100 (47 bases) Smyx CRISPR Repeat Consensus 101 (47 bases) Ptor CRISPR Repeat Consensus 102 46 bases) Lreu Anti-Repeat 103 (36 bases) Lros Anti-Repeat 104 (37 bases) Ppen Anti-Repeat 105 (37 bases) Lnod Anti-Repeat 106 (38 bases) Sspe Anti-Repeat 107 (39 bases) Bthe Anti-Repeat 108 (36 bases) Lves Anti-Repeat 109 (36 bases) Ssan Anti-Repeat 110 (36 bases) Eten Anti-Repeat 111 (47 bases) Smyx Anti-Repeat 112 (47 bases) Ptor Anti-Repeat 113 (46 bases) Lreu Single guide RNA 114 (169 bases) Lros Single guide RNA 115 (166 bases) Ppen Single guide RNA 116 (168 bases) Lnod Single guide RNA 117 (114 bases) Sspe Single guide RNA 118 (180 bases) Sspe Single guide RNA 119 (117 bases) Bthe Single guide RNA 120 (254 bases) Lves Single guide RNA 121 (200 bases) Ssan Single guide RNA 122 (195 bases) Eten Single guide RNA 123 (155 bases) Smyx Single guide RNA 124 (149 bases) Ptor Single guide RNA 125 (155 bases) GG-939 126 (57 bases) Single guide RNA 127 (174 bases) Lreu Single guide RNA 128 (166 bases) Lros Single guide RNA 129 (163 bases) Ppen Single guide RNA 130 (165 bases) Lnod Single guide RNA 131 (111 bases) Sspe Single guide RNA 132 (177 bases) Sspe Single guide RNA 133 (114 bases) Bthe Single guide RNA 134 (251 bases) Lves Single guide RNA 135 (197 bases) Ssan Single guide RNA 136 (192 bases) Eten Single guide RNA 137 (152 bases) Smyx Single guide RNA 138 (146 bases) Ptor Single guide RNA 139 (152 bases) Cas9 endonuclease Brevibacillus laterosporus 140 (1092 aa) bacterial strain SSP360D4 Variable Targeting domain-direct 141 Variable Targeting domain-reverse 142 16 nt loop of the repeat-direct 143 16 nt loop of the repeat-reverse 144 anti-repeat region-direct 145 anti-repeat region-reverse 146 Putative 3' tracrRNA Sequence - direct 147 Putative 3' tracrRNA Sequence - reverse 148 Lactobacillus reuteri Mlc3 (Lreu) crRNA repeat 149 region Lactobacillus rossiae DSM 15814 (Lros) crRNA 150 repeat region Pediococcus pentosaceus SL4 (Ppen) crRNA 151 repeat region Lactobacillus nodensis JCM 14932 (Lnod) 152 crRNA repeat region Sulfurospirillum sp. SCADC (Sspe) crRNA 153-154 repeat region Bifidobacterium thermophilum DSM 20210 155 (Bthe) crRNA repeat region Loktanella vestfoldensis (Lves) crRNA repeat 156 region Sphingomonas sanxanigenens NX02 (Ssan) 157 crRNA repeat region Epilithonimonas tenax DSM 16811 (Eten) 158 crRNA repeat region Sporocytophaga myxococcoides (Smyx) crRNA 159 repeat region Psychroflexus torquis ATCC 700755 (Ptor) 160 crRNA repeat region Lactobacillus reuteri Mlc3 (Lreu) tracrRNA anti- 161 repeat Lactobacillus rossiae DSM 15814 (Lros) 162 tracrRNA anti-repeat Pediococcus pentosaceus SL4 (Ppen) 163 tracrRNA anti-repeat Lactobacillus nodensis JCM 14932 (Lnod) 164 tracrRNA anti-repeat Sulfurospirillum sp. SCADC (Sspe) tracrRNA 165-166 anti-repeat Bifidobacterium thermophilum DSM 20210 167 (Bthe) tracrRNA anti-repeat Loktanella vestfoldensis (Lves) tracrRNA anti- 168 repeat Sphingomonas sanxanigenens NX02 (Ssan) 169 tracrRNA anti-repeat Epilithonimonas tenax DSM 16811 (Eten) 170 tracrRNA anti-repeat Sporocytophaga myxococcoides (Smyx) 171 tracrRNA anti-repeat Psychroflexus torquis ATCC 700755 (Ptor) 172 tracrRNA anti-repeat Lactobacillus reuteri Mlc3 (Lreu) 3' tracrRNA 173 Lactobacillus rossiae DSM 15814 (Lros) 3' 174 tracrRNA Pediococcus pentosaceus SL4 (Ppen) 3' 175 tracrRNA Lactobacillus nodensis JCM 14932 (Lnod) 3' 176 tracrRNA Sulfurospirillum sp. SCADC (Sspe) 3' tracrRNA 177-178 Bifidobacterium thermophilum DSM 20210 179 (Bthe) 3' tracrRNA Loktanella vestfoldensis (Lves) 3' tracrRNA 180 Sphingomonas sanxanigenens NX02 (Ssan) 3' 181 tracrRNA Epilithonimonas tenax DSM 16811 (Eten) 3' 182 tracrRNA Sporocytophaga myxococcoides (Smyx) 3' 183 tracrRNA Psychroflexus torquis ATCC 700755 (Ptor) 3' 184 tracrRNA
DETAILED DESCRIPTION
[0050] Compositions and methods are provided for rapid characterization of Cas endonuclease systems and the elements comprising such a systems, including, but not limiting to, rapid characterization of PAM sequences, guide RNA elements and Cas endonucleases. Cas9 endonuclease systems originating from Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides are described herein.
[0051] The present disclosure also describes methods for genome modification of a target sequence in the genome of a cell, for gene editing, and for inserting a polynucleotide of interest into the genome of a cell.
[0052] CRISPR (clustered regularly interspaced short palindromic repeats) loci refers to certain genetic loci encoding factors of DNA cleavage systems, for example, used by bacterial and archaeal cells to destroy foreign DNA (Horvath and Barrangou, 2010, Science 327:167-170). A CRISPR locus can consist of a CRISPR array, comprising short direct repeats separated by short variable DNA sequences (called `spacers`), which can be flanked by diverse Cas (CRISPR-associated) genes. Multiple CRISPR-Cas systems have been described including Class 1 systems, with multisubunit effector complexes, and Class 2 systems, with single protein effectors (such as but not limiting to Cas9, Cpf1, C2c1, C2c2, C2c3). (Zetsche et al., 2015, Cell 163, 1-13; Shmakov et al., 2015, Molecular Cell 60, 1-13; Makarova et al. 2015, Nature Reviews Microbiology Vol. 13:1-15; WO 2013/176772 A1 published on Nov. 23, 2013 and incorporated by its entirety by reference herein).
[0053] The type II CRISPR/Cas system from bacteria employs a crRNA (CRISPR RNA) and tracrRNA (trans-activating CRISPR RNA) to guide a Cas9 endonuclease (encoded by a cas9 gene) to its DNA target. The crRNA contains a spacer region complementary to one strand of the double strand DNA target and a region that base pairs with the tracrRNA (trans-activating CRISPR RNA) forming a RNA duplex that directs the Cas9 endonuclease to cleave the DNA target. Spacers are acquired through a not fully understood process involving Cas1 and Cas2 proteins. All type II CRISPR-Cas loci contain cas1 and cas2 genes in addition to the cas9 gene (Makarova et al. 2015, Nature Reviews Microbiology Vol. 13:1-15). Type II CRISR-Cas loci can encode a tracrRNA, which is partially complementary to the repeats within the respective CRISPR array, and can comprise other proteins such as Csn1 and Csn2. The presence of cas9 in the vicinity of cas1 and cas2 genes is the hallmark of type II loci (Makarova et al. 2015, Nature Reviews Microbiology Vol. 13:1-15).
[0054] The number of CRISPR-associated genes at a given CRISPR locus can vary between species (Haft et al., 2005, Computational Biology, PLoS Comput Biol 1(6): e60. doi:10.1371/journal.pcbi.0010060; Makarova et al. 2015, Nature Reviews Microbiology Vol. 13:1-15; WO 2013/176772 A1 published on Nov. 23, 2013 and incorporated by its entirety by reference herein).
[0055] The term "Cas gene" herein refers to a gene that is generally coupled, associated or close to, or in the vicinity of flanking CRISPR loci. The terms "Cas gene", "CRISPR-associated (Cas) gene" are used interchangeably herein.
[0056] The term "Cas endonuclease" herein refers to a protein encoded by a Cas gene. A Cas endonuclease herein, when in complex with a suitable polynucleotide component, is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a specific DNA target sequence. A Cas endonuclease described herein comprises one or more nuclease domains. Cas endonucleases of the disclosure includes those having a HNH or HNH-like nuclease domain and/or a RuvC or RuvC-like nuclease domain. A Cas endonuclease of the disclosure includes a Cas9 protein, a Cpf1 protein, a C2c1 protein, a C2c2 protein, a C2c3 protein, Cas3, Cas 5, Cas7, Cas8, Cas10, or complexes of these.
[0057] As used herein, the terms "guide polynucleotide/Cas endonuclease complex", "guide polynucleotide/Cas endonuclease system", "guide polynucleotide/Cas complex", "guide polynucleotide/Cas system" are used interchangeably herein and refer to at least one guide polynucleotide and at least one Cas endonuclease that are capable of forming a complex, wherein said guide polynucleotide/Cas endonuclease complex can direct the Cas endonuclease to a DNA target site, enabling the Cas endonuclease to recognize, bind to, and optionally nick or cleave (introduce a single or double strand break) into the DNA target site. A guide polynucleotide/Cas endonuclease complex herein can comprise Cas protein(s) and suitable polynucleotide component(s) of any of the four known CRISPR systems (Horvath and Barrangou, Science 327:167-170) such as a type I, II, or III CRISPR system. A Cas endonuclease unwinds the DNA duplex at the target sequence and optionally cleaves at least one DNA strand, as mediated by recognition of the target sequence by a polynucleotide (such as, but not limited to, a crRNA or guide RNA) that is in complex with the Cas protein. Such recognition and cutting of a target sequence by a Cas endonuclease typically occurs if the correct protospacer-adjacent motif (PAM) is located at or adjacent to the 3' end of the DNA target sequence. Alternatively, a Cas protein herein may lack DNA cleavage or nicking activity, but can still specifically bind to a DNA target sequence when complexed with a suitable RNA component. (See also U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015 and US 2015-0059010 A1, published on Feb. 26, 2015, both are hereby incorporated in its entirety by reference).
[0058] A guide polynucleotide/Cas endonuclease complex can cleave one or both strands of a DNA target sequence. A guide polynucleotide/Cas endonuclease complex that can cleave both strands of a DNA target sequence typically comprises a Cas protein that has all of its endonuclease domains in a functional state (e.g., wild type endonuclease domains or variants thereof retaining some or all activity in each endonuclease domain). Thus, a wild type Cas protein (e.g., a Cas9 protein disclosed herein), or a variant thereof retaining some or all activity in each endonuclease domain of the Cas protein, is a suitable example of a Cas endonuclease that can cleave both strands of a DNA target sequence. A Cas9 protein comprising functional RuvC and HNH nuclease domains is an example of a Cas protein that can cleave both strands of a DNA target sequence. A guide polynucleotide/Cas endonuclease complex that can cleave one strand of a DNA target sequence can be characterized herein as having nickase activity (e.g., partial cleaving capability). A Cas nickase typically comprises one functional endonuclease domain that allows the Cas to cleave only one strand (i.e., make a nick) of a DNA target sequence. For example, a Cas9 nickase may comprise (i) a mutant, dysfunctional RuvC domain and (ii) a functional HNH domain (e.g., wild type HNH domain). As another example, a Cas9 nickase may comprise (i) a functional RuvC domain (e.g., wild type RuvC domain) and (ii) a mutant, dysfunctional HNH domain. Non-limiting examples of Cas9 nickases suitable for use herein are disclosed by Gasiunas et al. (Proc. Natl. Acad. Sci. U.S.A. 109:E2579-E2586), Jinek et al. (Science 337:816-821), Sapranauskas et al. (Nucleic Acids Res. 39:9275-9282) and in U.S. Patent Appl. Publ. No. 2014/0189896, which are incorporated herein by reference.
[0059] A pair of Cas9 nickases can be used to increase the specificity of DNA targeting. In general, this can be done by providing two Cas9 nickases that, by virtue of being associated with RNA components with different guide sequences, target and nick nearby DNA sequences on opposite strands in the region for desired targeting. Such nearby cleavage of each DNA strand creates a double strand break (i.e., a DSB with single-stranded overhangs), which is then recognized as a substrate for non-homologous-end-joining, NHEJ (leading to indel formation) or homologous recombination, HR. Each nick in these embodiments can be at least about 5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, or 100 (or any integer between 5 and 100) bases apart from each other, for example. One or two Cas9 nickase proteins herein can be used in a Cas9 nickase pair. For example, a Cas9 nickase with a mutant RuvC domain, but functioning HNH domain (i.e., Cas9 HNH+/RuvC-), could be used (e.g., Streptococcus pyogenes Cas9 HNH+/RuvC-). Each Cas9 nickase (e.g., Cas9 HNH+/RuvC-) would be directed to specific DNA sites nearby each other (up to 100 base pairs apart) by using suitable RNA components herein with guide RNA sequences targeting each nickase to each specific DNA site.
[0060] A Cas protein can be part of a fusion protein comprising one or more heterologous protein domains (e.g., 1, 2, 3, or more domains in addition to the Cas protein). Such a fusion protein may comprise any additional protein sequence, and optionally a linker sequence between any two domains, such as between Cas and a first heterologous domain. Examples of protein domains that may be fused to a Cas protein herein include, without limitation, epitope tags (e.g., histidine [His], V5, FLAG, influenza hemagglutinin [HA], myc, VSV-G, thioredoxin [Trx]), reporters (e.g., glutathione-5-transferase [GST], horseradish peroxidase [HRP], chloramphenicol acetyltransferase [CAT], beta-galactosidase, beta-glucuronidase [GUS], luciferase, green fluorescent protein [GFP], HcRed, DsRed, cyan fluorescent protein [CFP], yellow fluorescent protein [YFP], blue fluorescent protein [BFP]), and domains having one or more of the following activities: methylase activity, demethylase activity, transcription activation activity (e.g., VP16 or VP64), transcription repression activity, transcription release factor activity, histone modification activity, RNA cleavage activity and nucleic acid binding activity. A Cas protein can also be in fusion with a protein that binds DNA molecules or other molecules, such as maltose binding protein (MBP), S-tag, Lex A DNA binding domain (DBD), GAL4A DNA binding domain, and herpes simplex virus (HSV) VP16.
[0061] A guide polynucleotide/Cas endonuclease complex in certain embodiments can bind to a DNA target site sequence, but does not cleave any strand at the target site sequence. Such a complex may comprise a Cas protein in which all of its nuclease domains are mutant, dysfunctional. For example, a Cas9 protein herein that can bind to a DNA target site sequence, but does not cleave any strand at the target site sequence, may comprise both a mutant, dysfunctional RuvC domain and a mutant, dysfunctional HNH domain. A Cas protein herein that binds, but does not cleave, a target DNA sequence can be used to modulate gene expression, for example, in which case the Cas protein could be fused with a transcription factor (or portion thereof) (e.g., a repressor or activator, such as any of those disclosed herein).
[0062] In one embodiment, the Cas endonuclease gene is a Type II Cas9 endonuclease, such as but not limited to, Cas9 genes listed in SEQ ID NOs: 462, 474, 489, 494, 499, 505, and 518 of WO2007/025097 published Mar. 1, 2007, and incorporated herein by reference. In another embodiment, the Cas endonuclease gene is a plant, maize or soybean optimized Cas9 endonuclease gene. The Cas endonuclease gene can be operably linked to a SV40 nuclear targeting signal upstream of the Cas codon region and a bipartite VirD2 nuclear localization signal (Tinland et al. (1992) Proc. Natl. Acad. Sci. USA 89:7442-6) downstream of the Cas codon region.
[0063] "Cas9" (formerly referred to as Cas5, Csn1, or Csx12) herein refers to a Cas endonuclease of a type II CRISPR system that forms a complex with a crNucleotide and a tracrNucleotide, or with a single guide polynucleotide, for specifically recognizing and cleaving all or part of a DNA target sequence. Cas9 protein comprises a RuvC nuclease domain and an HNH (H--N--H) nuclease domain, each of which can cleave a single DNA strand at a target sequence (the concerted action of both domains leads to DNA double-strand cleavage, whereas activity of one domain leads to a nick). In general, the RuvC domain comprises subdomains I, II and III, where domain I is located near the N-terminus of Cas9 and subdomains II and III are located in the middle of the protein, flanking the HNH domain (Hsu et al, Cell 157:1262-1278).
[0064] Cas9 endonculeases are typically derived from a type II CRISPR system, which includes a DNA cleavage system utilizing a Cas9 endonuclease in complex with at least one polynucleotide component. For example, a Cas9 can be in complex with a CRISPR RNA (crRNA) and a trans-activating CRISPR RNA (tracrRNA). In another example, a Cas9 can be in complex with a single guide RNA
[0065] In one embodiment of the disclosure, the composition comprises at least one Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81-91, or a functional fragment thereof.
[0066] In one embodiment of the disclosure, the composition comprises at least one recombinant DNA vector encoding the Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81-91 (such as the DNA sequences form SEQ ID NO: 70-80), or mRNA encoding Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81-91. The Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81-91 can form a (Ribonucleotide Protein--RNP) complex with at least one guide RNA, wherein said complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a target site.
[0067] Recombinant DNA expressing the Cas9 endonucleases described herein (including functional fragments thereof, plant or microbe codon optimized Cas9 endonuclease) can be stably integrated into the genome of an organism. For example, plants can be produced that comprise a cas9 gene stably integrated in the plant's genome. Plants expressing a stably integrated Cas endonuclease can be exposed to at least one guide RNA and/or a polynucleotide modification templates and/or donor DNAs to enable genome modifications such as gene knockout, gene editing or DNA insertions.
[0068] A variant of a Cas9 protein sequence may be used, but should have specific binding activity, and optionally endonucleolytic activity, toward DNA when associated with an RNA component herein. Such a variant may comprise an amino acid sequence that is at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% A identical to the amino acid sequence of the reference Cas9. Alternatively, a Cas9 protein may comprise an amino acid sequence that is at least about 80%.sup., 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identical to any of the foregoing amino acid sequences, for example. Such a variant Cas9 protein should have specific binding activity, and optionally cleavage or nicking activity, toward DNA when associated with an RNA component herein.
[0069] The Cas endonuclease can comprise a modified form of the Cas9 polypeptide. The modified form of the Cas9 polypeptide can include an amino acid change (e.g., deletion, insertion, or substitution) that reduces the naturally-occurring nuclease activity of the Cas9 protein. For example, in some instances, the modified form of the Cas9 protein has less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5%, or less than 1% of the nuclease activity of the corresponding wild-type Cas9 polypeptide (US patent application US20140068797 A1 published on Mar. 6, 2014). In some cases, the modified form of the Cas9 polypeptide has no substantial nuclease activity and is referred to as catalytically "inactivated Cas9" or "deactivated cas9 (dCas9)." Catalytically inactivated Cas9 variants include Cas9 variants that contain mutations in the HNH and RuvC nuclease domains. These catalytically inactivated Cas9 variants are capable of interacting with sgRNA and binding to the target site in vivo but cannot cleave either strand of the target DNA.
[0070] A catalytically inactive Cas9 can be fused to a heterologous sequence (US patent application US20140068797 A1 published on Mar. 6, 2014). Suitable fusion partners include, but are not limited to, a polypeptide that provides an activity that indirectly increases transcription by acting directly on the target DNA or on a polypeptide (e.g., a histone or other DNA-binding protein) associated with the target DNA. Additional suitable fusion partners include, but are not limited to, a polypeptide that provides for methyltransferase activity, demethylase activity, acetyltransferase activity, deacetylase activity, kinase activity, phosphatase activity, ubiquitin ligase activity, deubiquitinating activity, adenylation activity, deadenylation activity, SUMOylating activity, deSUMOylating activity, ribosylation activity, deribosylation activity, myristoylation activity, or demyristoylation activity. Further suitable fusion partners include, but are not limited to, a polypeptide that directly provides for increased transcription of the target nucleic acid (e.g., a transcription activator or a fragment thereof, a protein or fragment thereof that recruits a transcription activator, a small molecule/drug-responsive transcription regulator, etc.). A catalytically inactive Cas9 can also be fused to a Fokl nuclease to generate double strand breaks (Guilinger et al. Nature biotechnology, volume 32, number 6, June 2014).
[0071] A Cas protein herein such as a Cas9 endonuclease protein can comprise a heterologous nuclear localization sequence (NLS). A heterologous NLS amino acid sequence herein may be of sufficient strength to drive accumulation of a Cas protein in a detectable amount in the nucleus of a yeast cell herein, for example. An NLS may comprise one (monopartite) or more (e.g., bipartite) short sequences (e.g., 2 to 20 residues) of basic, positively charged residues (e.g., lysine and/or arginine), and can be located anywhere in a Cas amino acid sequence but such that it is exposed on the protein surface. An NLS may be operably linked to the N-terminus or C-terminus of a Cas protein herein, for example. Two or more NLS sequences can be linked to a Cas protein, for example, such as on both the N- and C-termini of a Cas protein. The Cas endonuclease gene can be operably linked to a SV40 nuclear targeting signal upstream of the Cas codon region and a bipartite VirD2 nuclear localization signal (Tinland et al. (1992) Proc. Natl. Acad. Sci. USA 89:7442-6) downstream of the Cas codon region. Non-limiting examples of suitable NLS sequences herein include those disclosed in U.S. Pat. No. 7,309,576, which is incorporated herein by reference.
[0072] The terms "functional fragment", "fragment that is functionally equivalent" and "functionally equivalent fragment" of a Cas endonuclease are used interchangeably herein, and refer to a portion or subsequence of the Cas endonuclease sequence of the present disclosure in which the ability to recognize, bind to, and optionally nick or cleave (introduce a single or double strand break in) the target site is retained.
[0073] The terms "functional variant", "Variant that is functionally equivalent" and "functionally equivalent variant" of a Cas endonuclease are used interchangeably herein, and refer to a variant of the Cas endonuclease of the present disclosure in which the ability to recognize, bind to, and optionally nick or cleave (introduce a single or double strand break in) the target site is retained. Fragments and variants can be obtained via methods such as site-directed mutagenesis and synthetic construction.
[0074] In one embodiment, the Cas endonuclease gene is a plant codon optimized Streptococcus pyogenes Cas9 gene that can recognize any genomic sequence of the form N(12-30)NGG can in principle be targeted.
[0075] In one embodiment, the Cas endonuclease is a Cas9 endonuclease originated from organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said Cas9 endonuclease can form a guide RNA/Cas endonuclease complex capable of recognizing, binding to, and optionally nicking or cleaving all or part of a DNA target sequence.
[0076] The Cas endonuclease can be introduced directly into a cell by any method known in the art, for example, but not limited to transient introduction methods, transfection and/or topical application.
[0077] The guide polynucleotides and guide polynucleotide/Cas endonuclease systems described herein include guide polynucleotides comprising a crRNA (comprising a variable targeting (VT) domain linked to tracr-mate sequence that can hybridized to the tracr nucleotide) wherein said guide polynucleotide directs sequence-specific binding of the guide polynucleotide/Cas endonuclease complex to a target sequence in a eukaryotic cell. In an aspect, the guide polynucleotide targets a target sequence in a non-human eukaryotic organism preferably a multicellular eukaryotic organism, comprising a eukaryotic host cell. In one aspect, the guide polynucleotide is a non-naturally occurring guide polynucleotide or a guide polynucleotide targeting a target sequence that is not natural to bacteria. The disclosed guide polynucleotides can be reprogrammed to target nucleotide sequences in non-bacterial cells such as, but not limiting to changing the VT domain to target non-bacterial target sequences and sequences not naturally acquired by the system from which the crRNA was obtained. Alternatively, the VT domain can be programmed to guide the crRNA to a target sequence in a eukaryotic genome. Any sequence in a eukaryotic genome can be targeted using the disclosed guide polynucleotides, such as, mammalian (e.g. human, mouse, etc.), yeast, insect, animal, and plant sequences. In other embodiments, the VT domain can be programmed to guide the crRNA to a target sequence in a prokaryotic genome or bacterial plasmid sequence that is not naturally targeted by the native system.
[0078] In some embodiments, the guide polynucleotide/Cas endonuclease complex comprises one or more nuclear localization sequences of sufficient strength to drive accumulation of said complex in a detectable amount in the nucleus of a eukaryotic cell. For example, nuclear localization signals can be added to the N- or C- or both the N- and C-terminus of the Cas protein. In other embodiments, one or more cellular localization signals can be included in the complex to provide for accumulation of the complex in a detectable amount in cellular organelles in which a desired target sequence is contained. For example, chloroplast targeting sequences can be added to the Cas protein to provide accumulation in a chloroplast organelle in a plant cell where the desired target sequence is found in the plant chloroplast genome.
[0079] The guide polynucleotide/Cas endonuclease system described herein can be provided to eukaryotic cells and reprogrammed to facilitate cleavage of endogenous eukaryotic target polynucleotides.
[0080] Endonucleases are enzymes that cleave the phosphodiester bond within a polynucleotide chain, and include restriction endonucleases that cleave DNA at specific sites without damaging the bases. Restriction endonucleases include Type I, Type II, Type III, and Type IV endonucleases, which further include subtypes. In the Type I and Type III systems, both the methylase and restriction activities are contained in a single complex. Endonucleases also include meganucleases, also known as homing endonucleases (HEases), which like restriction endonucleases, bind and cut at a specific recognition site, however the recognition sites for meganucleases are typically longer, about 18 bp or more (patent application WO-PCT PCT/US12/30061 filed on Mar. 22, 2012). Meganucleases have been classified into four families based on conserved sequence motifs, the families are the LAGLIDADG, GIY-YIG, H-N-H, and His-Cys box families. These motifs participate in the coordination of metal ions and hydrolysis of phosphodiester bonds. HEases are notable for their long recognition sites, and for tolerating some sequence polymorphisms in their DNA substrates. The naming convention for meganuclease is similar to the convention for other restriction endonuclease. Meganucleases are also characterized by prefix F-, I-, or PI- for enzymes encoded by free-standing ORFs, introns, and inteins, respectively. One step in the recombination process involves polynucleotide cleavage at or near the recognition site. This cleaving activity can be used to produce a double-strand break. For reviews of site-specific recombinases and their recognition sites, see, Sauer (1994) Curr Op Biotechnol 5:521-7; and Sadowski (1993) FASEB 7:760-7. In some examples the recombinase is from the Integrase or Resolvase families.
[0081] TAL effector nucleases are a new class of sequence-specific nucleases that can be used to make double-strand breaks at specific target sequences in the genome of a plant or other organism. (Miller et al. (2011) Nature Biotechnology 29:143-148). Zinc finger nucleases (ZFNs) are engineered double-strand break inducing agents comprised of a zinc finger DNA binding domain and a double-strand-break-inducing agent domain. Recognition site specificity is conferred by the zinc finger domain, which typically comprising two, three, or four zinc fingers, for example having a C2H2 structure, however other zinc finger structures are known and have been engineered. Zinc finger domains are amenable for designing polypeptides which specifically bind a selected polynucleotide recognition sequence. ZFNs include an engineered DNA-binding zinc finger domain linked to a non-specific endonuclease domain, for example nuclease domain from a Type IIs endonuclease such as Fokl. Additional functionalities can be fused to the zinc-finger binding domain, including transcriptional activator domains, transcription repressor domains, and methylases. In some examples, dimerization of nuclease domain is required for cleavage activity. Each zinc finger recognizes three consecutive base pairs in the target DNA. For example, a 3 finger domain recognized a sequence of 9 contiguous nucleotides, with a dimerization requirement of the nuclease, two sets of zinc finger triplets are used to bind an 18 nucleotide recognition sequence.
[0082] As used herein, the term "guide polynucleotide", relates to a polynucleotide sequence that can form a complex with a Cas endonuclease and enables the Cas endonuclease to recognize, bind to, and optionally cleave a DNA target site. The guide polynucleotide can be a single molecule or a double molecule. The guide polynucleotide sequence can be a RNA sequence, a DNA sequence, or a combination thereof (a RNA-DNA combination sequence). Optionally, the guide polynucleotide can comprise at least one nucleotide, phosphodiester bond or linkage modification such as, but not limited, to Locked Nucleic Acid (LNA), 5-methyl dC, 2,6-Diaminopurine, 2'-Fluoro A, 2'-Fluoro U, 2'-O-Methyl RNA, phosphorothioate bond, linkage to a cholesterol molecule, linkage to a polyethylene glycol molecule, linkage to a spacer 18 (hexaethylene glycol chain) molecule, or 5' to 3' covalent linkage resulting in circularization. A guide polynucleotide that solely comprises ribonucleic acids is also referred to as a "guide RNA" or "gRNA" (See also U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015 and US 2015-0059010 A1, published on Feb. 26, 2015, both are hereby incorporated in its entirety by reference).
[0083] In one embodiment of the disclosure, the guide polynucleotide is a single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139.
[0084] In one embodiment of the disclosure, the guide polynucleotide is a single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA comprises a chimeric non-naturally occurring crRNA linked to a tracrRNA, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160.
[0085] In one embodiment of the disclosure, the guide polynucleotide is a guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence.
[0086] The guide polynucleotide can be a double molecule (also referred to as duplex guide polynucleotide) comprising a crNucleotide sequence and a tracrNucleotide sequence. The crNucleotide includes a first nucleotide sequence domain (referred to as Variable Targeting domain or VT domain) that can hybridize to a nucleotide sequence in a target DNA and a second nucleotide sequence (also referred to as a tracr mate sequence) that is part of a Cas endonuclease recognition (CER) domain. The tracr mate sequence can hybridized to a tracrNucleotide along a region of complementarity and together form the Cas endonuclease recognition domain or CER domain. The CER domain is capable of interacting with a Cas endonuclease polypeptide. The crNucleotide and the tracrNucleotide of the duplex guide polynucleotide can be RNA, DNA, and/or RNA-DNA-combination sequences. In some embodiments, the crNucleotide molecule of the duplex guide polynucleotide is referred to as "crDNA" (when composed of a contiguous stretch of DNA nucleotides) or "crRNA" (when composed of a contiguous stretch of RNA nucleotides), or "crDNA-RNA" (when composed of a combination of DNA and RNA nucleotides). The crNucleotide can comprise a fragment of the crRNA naturally occurring in Bacteria and Archaea. The size of the fragment of the crRNA naturally occurring in Bacteria and Archaea that can be present in a crNucleotide disclosed herein can range from, but is not limited to, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 or more nucleotides. In some embodiments the tracrNucleotide is referred to as "tracrRNA" (when composed of a contiguous stretch of RNA nucleotides) or "tracrDNA" (when composed of a contiguous stretch of DNA nucleotides) or "tracrDNA-RNA" (when composed of a combination of DNA and RNA nucleotides. In one embodiment, the RNA that guides the RNA/Cas9 endonuclease complex is a duplexed RNA comprising a duplex crRNA-tracrRNA. The tracrRNA (trans-activating CRISPR RNA) contains, in the 5'-to-3' direction, (i) a sequence that anneals with the repeat region of CRISPR type II crRNA and (ii) a stem loop-containing portion (Deltcheva et al., Nature 471:602-607). The duplex guide polynucleotide can form a complex with a Cas endonuclease, wherein said guide polynucleotide/Cas endonuclease complex (also referred to as a guide polynucleotide/Cas endonuclease system) can direct the Cas endonuclease to a genomic target site, enabling the Cas endonuclease to recognize, bind to, and optionally nick or cleave (introduce a single or double strand break) into the target site. (See also U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015 and US 2015-0059010 A1, published on Feb. 26, 2015, both are hereby incorporated in its entirety by reference.)
[0087] The guide polynucleotide can also be a single molecule (also referred to as single guide polynucleotide) comprising a crNucleotide sequence linked to a tracrNucleotide sequence. The single guide polynucleotide comprises a first nucleotide sequence domain (referred to as Variable Targeting domain or VT domain) that can hybridize to a nucleotide sequence in a target DNA and a Cas endonuclease recognition domain (CER domain), that interacts with a Cas endonuclease polypeptide. By "domain" it is meant a contiguous stretch of nucleotides that can be RNA, DNA, and/or RNA-DNA-combination sequence. The VT domain and/or the CER domain of a single guide polynucleotide can comprise a RNA sequence, a DNA sequence, or a RNA-DNA-combination sequence. The single guide polynucleotide being comprised of sequences from the crNucleotide and the tracrNucleotide may be referred to as "single guide RNA" (when composed of a contiguous stretch of RNA nucleotides) or "single guide DNA" (when composed of a contiguous stretch of DNA nucleotides) or "single guide RNA-DNA" (when composed of a combination of RNA and DNA nucleotides). The single guide polynucleotide can form a complex with a Cas endonuclease, wherein said guide polynucleotide/Cas endonuclease complex (also referred to as a guide polynucleotide/Cas endonuclease system) can direct the Cas endonuclease to a genomic target site, enabling the Cas endonuclease to recognize, bind to, and optionally nick or cleave (introduce a single or double strand break) the target site. (See also U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015 and US 2015-0059010 A1, published on Feb. 26, 2015, both are hereby incorporated in its entirety by reference.)
[0088] The term "variable targeting domain" or "VT domain" is used interchangeably herein and includes a nucleotide sequence that can hybridize (is complementary) to one strand (nucleotide sequence) of a double strand DNA target site. The % complementation between the first nucleotide sequence domain (VT domain) and the target sequence can be at least 50%, 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 63%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100%. The variable target domain can be at least 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29 or 30 nucleotides in length. In some embodiments, the variable targeting domain comprises a contiguous stretch of 12 to 30 nucleotides. The variable targeting domain can be composed of a DNA sequence, a RNA sequence, a modified DNA sequence, a modified RNA sequence, or any combination thereof.
[0089] The term "Cas endonuclease recognition domain" or "CER domain" (of a guide polynucleotide) is used interchangeably herein and includes a nucleotide sequence that interacts with a Cas endonuclease polypeptide. A CER domain comprises a tracrNucleotide mate sequence followed by a tracrNucleotide sequence. The CER domain can be composed of a DNA sequence, a RNA sequence, a modified DNA sequence, a modified RNA sequence (see for example US 2015-0059010 A1, published on Feb. 26, 2015, incorporated in its entirety by reference herein), or any combination thereof.
[0090] The nucleotide sequence linking the crNucleotide and the tracrNucleotide of a single guide polynucleotide can comprise a RNA sequence, a DNA sequence, or a RNA-DNA combination sequence. In one embodiment, the nucleotide sequence linking the crNucleotide and the tracrNucleotide of a single guide polynucleotide can be at least 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99 or 100 nucleotides in length. In another embodiment, the nucleotide sequence linking the crNucleotide and the tracrNucleotide of a single guide polynucleotide can comprise a tetraloop sequence, such as, but not limiting to a GAAA tetraloop sequence.
[0091] Nucleotide sequence modification of the guide polynucleotide, VT domain and/or CER domain can be selected from, but not limited to, the group consisting of a 5' cap, a 3' polyadenylated tail, a riboswitch sequence, a stability control sequence, a sequence that forms a dsRNA duplex, a modification or sequence that targets the guide poly nucleotide to a subcellular location, a modification or sequence that provides for tracking, a modification or sequence that provides a binding site for proteins, a Locked Nucleic Acid (LNA), a 5-methyl dC nucleotide, a 2,6-Diaminopurine nucleotide, a 2'-Fluoro A nucleotide, a 2'-Fluoro U nucleotide; a 2'-O-Methyl RNA nucleotide, a phosphorothioate bond, linkage to a cholesterol molecule, linkage to a polyethylene glycol molecule, linkage to a spacer 18 molecule, a 5' to 3' covalent linkage, or any combination thereof. These modifications can result in at least one additional beneficial feature, wherein the additional beneficial feature is selected from the group of a modified or regulated stability, a subcellular targeting, tracking, a fluorescent label, a binding site for a protein or protein complex, modified binding affinity to complementary target sequence, modified resistance to cellular degradation, and increased cellular permeability.
[0092] The terms "a polynucleotide originating from organism", "a polynucleotide derived from organism" are used interchangeably herein and refer to a polynucleotide (such as but not limited to crRNA and tracrRNA) that is naturally occurring in said organism (native to said organism) or is isolated from said organism, or is a synthetic oligonucleotide that is identical to the polynucleotide isolated from said organism). For example, a tracrRNA originating from Brevibacillus laterosporus refers to a tracrRNA that occurs in Brevibacillus laterosporus, or is isolated from Brevibacillus laterosporus, or is a synthetic oligonucleotide that is identical to the tracrRNA isolated from Brevibacillus laterosporus.
[0093] The terms "functional fragment", "fragment that is functionally equivalent" and "functionally equivalent fragment" of a guide RNA, crRNA or tracrRNA are used interchangeably herein, and refer to a portion or subsequence of the guide RNA, crRNA or tracrRNA, respectively, of the present disclosure in which the ability to function as a guide RNA, crRNA or tracrRNA, respectively, is retained.
[0094] The terms "functional variant", "Variant that is functionally equivalent" and "functionally equivalent variant" of a guide RNA, crRNA or tracrRNA (respectively) are used interchangeably herein, and refer to a variant of the guide RNA, crRNA or tracrRNA, respectively, of the present disclosure in which the ability to function as a guide RNA, crRNA or tracrRNA, respectively, is retained.
[0095] As used herein, the terms "single guide RNA" and "sgRNA" are used interchangeably herein and relate to a synthetic fusion of two RNA molecules, a crRNA (CRISPR RNA) comprising a variable targeting domain (linked to a tracr mate sequence that hybridizes to a tracrRNA), fused to a tracrRNA (trans-activating CRISPR RNA). The single guide RNA can comprise a crRNA or crRNA fragment and a tracrRNA or tracrRNA fragment of the type II CRISP R/Cas system that can form a complex with a type II Cas endonuclease, wherein said guide RNA/Cas endonuclease complex can direct the Cas endonuclease to a genomic target site, enabling the Cas endonuclease to t recognize, bind to, and optionally nick or cleave (introduce a single or double strand break) into a genomic target site.
[0096] The components of the single or dual guide polynucleotides described herein (such as but no limiting to the crRNA, tracrRNA, variable targeting domain, crRNA repeat, tracr-mate domain, loop, tracrRNA anti-repeat, 3'tracrRNA sequence) can be modified to create functional variants of these components such that these functional variants can be combined to create a functional single or dual guide polynucleotide. Examples of guide polynucleotide component modifications are described herein and include nucleotide extensions at the 3' end, 5' end, or both end of any of components of the guide polynucleotide, and/or nucleotide sequence modifications (substitutions, insertions, deletions), and/or chemical modifications, and/or linkage modifications, or any combinations thereof.
[0097] Extensions at 3' end, 5' end, or both ends of any of components of the guide polynucleotide can be can be at least 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99 or 100 nucleotides in length.
[0098] Nucleotide sequence modification of the guide polynucleotide components include a 5' cap, a 3' polyadenylated tail, a riboswitch sequence, a stability control sequence, a sequence that forms a dsRNA duplex, a modification or sequence that targets the guide polynucleotide to a subcellular location, a modification or sequence that provides for tracking, a modification or sequence that provides a binding site for proteins, a Locked Nucleic Acid (LNA), a 5-methyl dC nucleotide, a 2,6-Diaminopurine nucleotide, a 2'-Fluoro A nucleotide, a 2'-Fluoro U nucleotide; a 2'-O-Methyl RNA nucleotide, a phosphorothioate bond, linkage to a cholesterol molecule, linkage to a polyethylene glycol molecule, linkage to a spacer 18 molecule, a 5' to 3' covalent linkage, or any combination thereof.
[0099] In one aspect, the functional variant single or dual guide polynucleotide has a similar activity than the guide polynucleotides of SEQ ID NOs: 127-139. In another aspect, the functional variant single or dual guide polynucleotide has an increased activity when compared to the guide polynucleotides of SEQ ID NOs: 127-139. The guide activity includes guide polynucleotide/Cas endonuclease ability to recognize, bind to and cleave a double strand break and/or RGEN mutation frequency.
[0100] The terms "guide RNA/Cas endonuclease complex", "guide RNA/Cas endonuclease system", "guide RNA/Cas complex", "guide RNA/Cas system", "gRNA/Cas complex", "gRNA/Cas system", "RNA-guided endonuclease", "RGEN" are used interchangeably herein and refer to at least one RNA component and at least one Cas endonuclease that are capable of forming a complex, wherein said guide RNA/Cas endonuclease complex can direct the Cas endonuclease to a DNA target site, enabling the Cas endonuclease to recognize, bind to, and optionally nick or cleave (introduce a single or double strand break) the DNA target site. A guide RNA/Cas endonuclease complex herein can comprise Cas protein(s) and suitable RNA component(s) of any of the four known CRISPR systems (Horvath and Barrangou, Science 327:167-170) such as a type I, II, or III CRISPR system. A guide RNA/Cas endonuclease complex can comprise a Type II Cas9 endonuclease and at least one RNA component (e.g., a crRNA and tracrRNA, or a gRNA). (See also U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015 and US 2015-0059010 A1, published on Feb. 26, 2015, both are hereby incorporated in its entirety by reference).
[0101] The guide polynucleotide can be introduced into a cell transiently, as single stranded polynucleotide or a double stranded polynucleotide, using any method known in the art such as, but not limited to, particle bombardment, Agrobacterium transformation or topical applications. The guide polynucleotide can also be introduced indirectly into a cell by introducing a recombinant DNA molecule (via methods such as, but not limited to, particle bombardment or Agrobacterium transformation) comprising a heterologous nucleic acid fragment encoding a guide polynucleotide, operably linked to a specific promoter that is capable of transcribing the guide RNA in said cell. The specific promoter can be, but is not limited to, a RNA polymerase III promoter, which allow for transcription of RNA with precisely defined, unmodified, 5'- and 3'-ends (DiCarlo et al., Nucleic Acids Res. 41: 4336-4343; Ma et al., Mol. Ther. Nucleic Acids 3:e161).
[0102] The terms "target site", "target sequence", "target site sequence, "target DNA", "target locus", "genomic target site", "genomic target sequence", "genomic target locus" and "protospacer", are used interchangeably herein and refer to a polynucleotide sequence such as, but not limited to, a nucleotide sequence on a chromosome, episome, or any other DNA molecule in the genome (including chromosomal, choloroplastic, mitochondrial DNA, plasmid DNA) of a cell, at which a guide polynucleotide/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave. The target site can be an endogenous site in the genome of a cell, or alternatively, the target site can be heterologous to the cell and thereby not be naturally occurring in the genome of the cell, or the target site can be found in a heterologous genomic location compared to where it occurs in nature. As used herein, terms "endogenous target sequence" and "native target sequence" are used interchangeable herein to refer to a target sequence that is endogenous or native to the genome of a cell and is at the endogenous or native position of that target sequence in the genome of the cell. Cells include, but are not limited to, human, non-human, animal, bacterial, fungal, insect, yeast, non-conventional yeast, and plant cells as well as plants and seeds produced by the methods described herein. An "artificial target site" or "artificial target sequence" are used interchangeably herein and refer to a target sequence that has been introduced into the genome of a cell. Such an artificial target sequence can be identical in sequence to an endogenous or native target sequence in the genome of a cell but be located in a different position (i.e., a non-endogenous or non-native position) in the genome of a cell.
[0103] An "altered target site", "altered target sequence", "modified target site", "modified target sequence" are used interchangeably herein and refer to a target sequence as disclosed herein that comprises at least one alteration when compared to non-altered target sequence. Such "alterations" include, for example:
(i) replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, or (iv) any combination of (i)-(iii).
[0104] The length of the target DNA sequence (target site) can vary, and includes, for example, target sites that are at least 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 or more nucleotides in length. It is further possible that the target site can be palindromic, that is, the sequence on one strand reads the same in the opposite direction on the complementary strand. The nick/cleavage site can be within the target sequence or the nick/cleavage site could be outside of the target sequence. In another variation, the cleavage could occur at nucleotide positions immediately opposite each other to produce a blunt end cut or, in other Cases, the incisions could be staggered to produce single-stranded overhangs, also called "sticky ends", which can be either 5' overhangs, or 3' overhangs. Active variants of genomic target sites can also be used. Such active variants can comprise at least 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to the given target site, wherein the active variants retain biological activity and hence are capable of being recognized and cleaved by an Cas endonuclease. Assays to measure the single or double-strand break of a target site by an endonuclease are known in the art and generally measure the overall activity and specificity of the agent on DNA substrates containing recognition sites.
[0105] A "protospacer adjacent motif" (PAM) herein refers to a short nucleotide sequence adjacent to a target sequence (protospacer) that is recognized (targeted) by a guide polynucleotide/Cas endonuclease system described herein. The Cas endonuclease may not successfully recognize a target DNA sequence if the target DNA sequence is not followed by a PAM sequence. The sequence and length of a PAM herein can differ depending on the Cas protein or Cas protein complex used. The PAM sequence can be of any length but is typically 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 nucleotides long.
[0106] A "randomized PAM" and "randomized protospacer adjacent motif" are used interchangeably herein, and refer to a random DNA sequence adjacent to a target sequence (protospacer) that is recognized (targeted) by a guide polynucleotide/Cas endonuclease system described herein. The randomized PAM sequence can be of any length but is typically 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 nucleotides long. A randomized nucleotide includes anyone of the nucleotides A, C, G or T.
[0107] The PAM sequence plays a key role in target recognition by licensing crRNA-guided base pairing to the protospacer sequence (Szczelkun et al, 2014, Proc. Natl. Acad. Sci. U.S.A 111: 9798-803). A strict PAM requirement constrains DNA target selection and poses a limit to Cas9 genome editing applications. Target site selection may be further confined if unique genomic sites are required especially in large complex plant genomes like maize (Xie et al, 2014, Mol. Plant 7: 923-6). These constraints imposed by the PAM and the specificity of the Spy Cas9 can be overcome by systematically redesigning the PAM specificity of a single Cas9 protein (Kleinstiver et al, 2015, Nature 523, 481-485. Described herein is a different method to overcome constraints imposed by the PAM and the specificity of the Cas9, namely by exploring the natural diversity of Cas9 proteins. The method described herein can also be combined with the method of systematically redesigning the PAM specificity to overcome constraints imposed by the PAM and the specificity of the Cas endonucleases.
[0108] Cas9 proteins from different bacteria recognize different PAM sequences (Horvath et al, 2008, J. Bacteriol. 190: 1401-12; Jinek et al, 2012, Science 337: 816-21; Gasiunas et al, 2012, Cell 154: 442-451; Zhang et al, 2013, Cell 50: 488-503; Fonfara et al, 2014, Nucleic Acids Res. 42: 2577-2590). Typically, the PAM sequences of new Cas9 proteins are identified by computational analysis of sequences immediately flanking putative protospacers in bacteriophage genomes (Shah et al, 2013, RNA Biol. 10: 1-9). Currently, with >1000 Cas9 protein orthologues available (Chylinski et al, 2014 Nucleic Acids Res. 42: 6091-6105; Hsu et al, 2014, Cell 157: 1262-1278), most spacers in Type II CRISPR arrays show only a few if any matches to the phage sequences present in databases, indicating that the vast majority of the phage universe is still unexplored. This constrains computational PAM identification methods and hinders the exploration of Cas9 protein diversity for genome editing applications.
[0109] As described herein, to address this problem a method was developed to empirically examine the PAM sequence requirements for any Cas9 protein. The method is based on the analysis of the in vitro cleavage products of a plasmid DNA library which contains a fixed protospacer target sequence and a stretch of 5 or 7 randomized base pairs in the putative PAM region. Based on the methods described herein, the a stretch of randomized base pairs in the putative PAM region can be at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 base pairs.
[0110] Using the method described herein, the canonical PAM preferences for Cas9 proteins of S. pyogenes and S. thermophilus CRISPR1 and CRISPR3 systems were first confirmed. Next, the method described herein was applied to identify the PAM and guide RNA requirements for a novel Cas9 protein from the Type II CRISPR-Cas system of B. laterosporus SPP360D4. In the Type II system of B. laterosporus, the transcriptional direction of the tracrRNA and CRISPR region could not be reliably predicted by computational approaches. Therefore, two single guide RNA (sgRNA) variants for both possible sense and anti-sense expression scenarios of the tracrRNA and CRISPR array (Examples 5-8, 10-12 described herein) were synthesized and only one of the designed sgRNAs supported cleavage of the randomized PAM plasmid library by B. laterosporus Cas9. Deep sequencing analysis of the cleavage products revealed a novel PAM requirement for the B. laterosporus Cas9. One that requires a strong preference for a C residue at position 5 of the PAM sequence followed by moderate preferences for A residues at positions 7 and 8 with an overall PAM consensus of NNNNCNDD (N=G, C, A or T; D=A, G or T). With a strong preference for just a single nucleotide, B. laterosporus Cas9 provides a useful addition to the Cas9 genome editing toolbox.
[0111] To examine the genome editing potential of a novel Cas9 and sgRNA characterized with the method described herein, the B. laterosporus SPP360D4 Cas9 and sgRNA were tested in maize (Examples 5-8, 10-12, described herein). As a result of cleavage, imperfect DNA repair resulted in INDEL mutations at all 3 chromosomal sites tested with robust INDEL frequencies observed at 2 of the 3 sites. Interestingly, at one of the sites, a .about.30% enhancement in the recovery of INDEL mutations was observed for the B. laterosporus Cas9 over the S. pyogenes Cas9 (Example 12).
[0112] In one embodiment described herein it is shown that cleavage of permissive PAMs is dependent on Cas9 concentration. For all Cas9 proteins analyzed, PAM sequences licensing plasmid DNA cleavage at higher (50 nM) Cas9 concentrations were more relaxed than PAM sequences identified at low (0.5 nM) Cas9 concentrations. This finding corroborates previous studies which demonstrated that lowering Cas9 concentration and shortening cleavage time prevents off-target cleavage by S. pyogenes Cas9 (Pattanayak et al, 2013, Nat. Biotechnol.: 1-7; Lin et al, 2014, Elife 3: e04766. doi: 10.7554/eLife.04766.). Since most other PAM determination methods have been performed in cells or cell extracts by expressing Cas9 at undefined concentrations (Ran et al, 2015, 2015 Apr. 9; 520(7546):186-91. doi: 10.1038/nature14299; Jiang et al, 2013, Nat. Biotechnol. 31: 233-9; Esvelt et al, 2013, November; 10(11):1116-21. doi: 10.1038/nmeth.2681; Kleinstiver et al, 2015), our method further refines PAM specificity assessments by the dose-dependent control of recombinant Cas9 protein in vitro. This allows the careful detailed examination of Cas9 PAM specificity as a function of Cas9 guide RNA complex concentration.
[0113] In one embodiment, the method describes herein further refines Cas9 PAM discovery efforts by the use of recombinant Cas9 protein and reframes PAM specificity as being non-static and dependent on Cas9-guide RNA complex concentration.
[0114] Described herein are novel Cas endonucleases derived from diverse organisms capable or forming guide polynucleotide/Cas endonuclease complexes with guide polynucleotides comprising crRNA and tracrRNA sequences fragments derived from their respective organisms. In one example, a Cas endonuclease derived from Brevibacillus laterosporus (SEQ ID NO: 140) was able to from a RGEN complex with a guide polynucleotide comprising a crRNA and a tracrRNA fragment derived from Brevibacillus laterosporus (such as SEQ ID NO: 47 or 127).
[0115] The Cas endonucleases described herein can also be used in complexes with guide polynucleotides derived from other Cas systems. In one example, the crRNA and/or tracrRNA domains of a guide polynucleotide capable of forming a complex with a Cas endonuclease from organism 1 (such that said RGEN complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a specific DNA target sequence), can be exchanged with a crRNA and/or tracrRNA domain, or fragment thereof, derived from a different organism (organism 2), thereby forming a chimeric guide, and still be able to form a functional complex with the Cas endonuclease derived from organism 1.
[0116] Similarities in guide RNA s between different Cas systems can be determined based on sequence composition and secondary structures of the guide RNAs. In one example, the secondary structure and sequence similarity of the sgRNAs from Lactobacillus reuteri MIc3 (Lreu) (SEQ ID NO: 114), Lactobacillus rossiae DSM 15814 (Lros) SEQ ID NO: 115) and Pediococcus pentosaceus SL4 (Ppen) SEQ ID NO: 116) were determined and revealed that these three sgRNAs have very similar secondary structures. It is anticipated that fragments from Lreu, Lros and PPen guide RNAs, such as but not limited to repeat structures or anti-repeat structures or any-one guide RNA domain, can be exchanged and/or mixed with one another to create chimeric guides capable of forming a RGEN with any one of the Lrue, Lros or Ppen Cas endonuclease (SEQ ID NOs: 81. 82 and 93, respectively). In another example, the secondary structure and sequence similarity of the sgRNAs from Lactobacillus nodensis JCM 14932 (Lnod) (SEQ ID NO:117), Loktanella vestfoldensis (Lves) (SEQ ID NO:121) and Sphingomonas sanxanigenens NX02 (Ssan) (SEQ ID NO: 122) was determined to be very similar, indicating that fragments from Lnod, Lves and Ssan guide RNAs, such as but not limited to repeat structures or anti-repeat structures or any-one guide RNA domain, can be exchanged and/or mixed with one another to create chimeric guides capable of forming a RGEN with any one of the Lnod, Lves or Ssan Cas endonuclease (SEQ ID NOs: 84, 87 and 88, respectively).
[0117] In another example, the secondary structure and sequence similarity of the sgRNAs from Epilithonimonas tenax DSM 16811 (Eten) (SEQ ID NO:123), Sporocytophaga myxococcoides (Smyx) (SEQ ID NO:138) and Psychroflexus torquis ATCC 700755 (Ptor) (SEQ ID NO: 139) was determined to be very similar, indicating that fragments from Eten, Smyx and Ptor guide RNAs, such as but not limited to repeat structures or anti-repeat structures or any-one guide RNA domain, can be exchanged and/or mixed with one another to create chimeric guides capable of forming a RGEN with any one of the Eten, Smyx or Ptor Cas endonuclease (SEQ ID NOs: 89, 90 and 91, respectively).
[0118] In one aspect, the Cas endonuclease and the crRNA and/or tracrRNA (or sgRNA) capable of forming a functional complex are derived or obtained from phylogenetically related groups. (See, for example, Fonfara et al Nucleic acid research 2014 Vol 42, No 4 pg. 2577-2590). It is understood that, based on the components of the novel Cas endonuclease systems described herein (crRNAs, tracrRNAs, Cas endonucleases, PAM sequences) one skilled in the art can exchange and/or mix any one component derived from one organism with any one component derived from another organism to make a functional guide polynucleotide/Cas endonuclease complex.
[0119] Guide polynucleotides can be modified to contain different sequence or structure yet be functionally equivalent or possess superior activity (binding, cutting, specificity). In one aspect, the chimeric guide polynucleotide can comprise at least one nucleotide, phosphodiester bond or linkage modification, or chemical modification such as, but not limited, to Locked Nucleic Acid (LNA), 5-methyl dC, 2,6-Diaminopurine, 2'-Fluoro A, 2'-Fluoro U, 2'-O-Methyl RNA, 2'-O-Methyl (M) modification, 2'-O-Methyl 3'phosphorothioate (MS) modification, 2'-O-Methyl 3'thioPACE (MSP) modification, phosphorothioate bond, linkage to a cholesterol molecule, linkage to a polyethylene glycol molecule, linkage to a spacer 18 (hexaethylene glycol chain) molecule, or 5' to 3' covalent linkage resulting in circularization (Hendel et al. 2015 Nature Biotechnology Vol. 33 pg. 985-991). Chimeric guide polynucleotides can be generated chemically, with or without sugar or backbone modifications. Chimeric guide polynucleotides can also be generated by in vitro transcription or delivered by DNA molecules containing promoters for expression
[0120] The PAM interacting domain, HNH or HNH-like nuclease domain, and/or RuvC or RuvC-like nuclease domains from the Cas endonuclease proteins described herein find use for creating Cas scaffolds (US2016/0102324 entitled "New compact scaffold of Cas9 in the type II CRISPR system, published Apr. 14, 2016 and incorporated herein by reference). The boundaries of the PAM interacting domain, RuvC and HNH domains of the Cas endonuclease described herein can be determined and new shorter Cas endonucleases derived from the Cas endonucleases described herein (or any one functional combination/fusion protein thereof) can be designed,
[0121] The terms "targeting", "gene targeting" and "DNA targeting" are used interchangeably herein. DNA targeting herein may be the specific introduction of a knock-out, edit, or knock-in at a particular DNA sequence, such as in a chromosome or plasmid of a cell. In general, DNA targeting can be performed herein by cleaving one or both strands at a specific DNA sequence in a cell with a Cas protein associated with a suitable polynucleotide component. Such DNA cleavage, if a double-strand break (DSB), can prompt NHEJ or HDR processes which can lead to modifications at the target site.
[0122] The terms "knock-out", "gene knock-out" and "genetic knock-out" are used interchangeably herein. A knock-out represents a DNA sequence of a cell that has been rendered partially or completely inoperative by targeting with a Cas protein; such a DNA sequence prior to knock-out could have encoded an amino acid sequence, or could have had a regulatory function (e.g., promoter), for example. A knock-out may be produced by an indel (insertion or deletion of nucleotide bases in a target DNA sequence through NHEJ), or by specific removal of sequence that reduces or completely destroys the function of sequence at or near the targeting site.
[0123] In one embodiment of the disclosure, the method comprises a method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, and at least one guide RNA, wherein said guide RNA and Cas endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site. The embodiment can further comprise identifying at least one cell that has a modification at said target, wherein the modification at said target site is selected from the group consisting of (i) a replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, and (iv) any combination of (i)-(iii).
[0124] The guide polynucleotide/Cas endonuclease system can be used in combination with a co-delivered polynucleotide modification template to allow for editing (modification) of a genomic nucleotide sequence of interest. (See also U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015 and WO2015/026886 A1, published on Feb. 26, 2015, both are hereby incorporated in its entirety by reference.)
[0125] A "modified nucleotide" or "edited nucleotide" refers to a nucleotide sequence of interest that comprises at least one alteration when compared to its non-modified nucleotide sequence. Such "alterations" include, for example: (i) replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, or (iv) any combination of (i)-(iii).
[0126] The term "polynucleotide modification template" includes a polynucleotide that comprises at least one nucleotide modification when compared to the nucleotide sequence to be edited. A nucleotide modification can be at least one nucleotide substitution, addition or deletion. Optionally, the polynucleotide modification template can further comprise homologous nucleotide sequences flanking the at least one nucleotide modification, wherein the flanking homologous nucleotide sequences provide sufficient homology to the desired nucleotide sequence to be edited.
[0127] In one embodiment, the disclosure describes a method for editing a nucleotide sequence in the genome of a cell, the method comprising providing a guide polynucleotide, a polynucleotide modification template, and at least one Cas endonuclease to a cell, wherein the Cas endonuclease is capable of introducing a single or double-strand break at a target sequence in the genome of said cell, wherein said polynucleotide modification template includes at least one nucleotide modification of said nucleotide sequence. Cells include, but are not limited to, human, non-human, animal, bacterial, fungal, insect, yeast, and plant cells as well as plants and seeds produced by the methods described herein. Plant cells include cells selected from the group consisting of maize, rice, sorghum, rye, barley, wheat, millet, oats, sugarcane, turfgrass, or switchgrass, soybean, canola, alfalfa, sunflower, cotton, tobacco, peanut, potato, tobacco, Arabidopsis, and safflower cells. The nucleotide to be edited can be located within or outside a target site recognized and cleaved by a Cas endonuclease. In one embodiment, the at least one nucleotide modification is not a modification at a target site recognized and cleaved by a Cas endonuclease. In another embodiment, there are at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 30, 40, 50, 100, 200, 300, 400, 500, 600, 700, 900 or 1000 nucleotides between the at least one nucleotide to be edited and the genomic target site.
[0128] In one embodiment of the disclosure, the method comprises a method for editing a nucleotide sequence in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, a polynucleotide modification template, and at least one guide RNA, wherein said polynucleotide modification template comprises at least one nucleotide modification of said nucleotide sequence, wherein said guide RNA and Cas endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site. Cells include, but are not limited to, human, non-human, animal, bacterial, fungal, insect, yeast, and plant cells as well as plants and seeds produced by the methods described herein.
[0129] Genome editing can be accomplished using any method of gene editing available. For example, gene editing can be accomplished through the introduction into a host cell of a polynucleotide modification template (sometimes also referred to as a gene repair oligonucleotide) containing a targeted modification to a gene within the genome of the host cell. The polynucleotide modification template for use in such methods can be either single-stranded or double-stranded. Examples of such methods are generally described, for example, in US Publication No. 2013/0019349.
[0130] In some embodiments, gene editing may be facilitated through the induction of a double-stranded break (DSB) in a defined position in the genome near the desired alteration. DSBs can be induced using any DSB-inducing agent available, including, but not limited to, TALENs, meganucleases, zinc finger nucleases, Cas9-gRNA systems (based on bacterial CRISPR-Cas systems), and the like. In some embodiments, the introduction of a DSB can be combined with the introduction of a polynucleotide modification template.
[0131] The process for editing a genomic sequence combining DSB and modification templates generally comprises: providing to a host cell, a DSB-inducing agent, or a nucleic acid encoding a DSB-inducing agent, that recognizes a target sequence in the chromosomal sequence and is able to induce a DSB in the genomic sequence, and at least one polynucleotide modification template comprising at least one nucleotide alteration when compared to the nucleotide sequence to be edited. The polynucleotide modification template can further comprise nucleotide sequences flanking the at least one nucleotide alteration, in which the flanking sequences are substantially homologous to the chromosomal region flanking the DSB. Genome editing using DSB-inducing agents, such as Cas9-gRNA complexes, has been described, for example in U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015, WO2015/026886 A1, published on Feb. 26, 2015, U.S. application 62/023,246, filed on Jul. 7, 2014, and U.S. application 62/036,652, filed on Aug. 13, 2014, all of which are incorporated by reference herein.
[0132] The terms "knock-in", "gene knock-in, "gene insertion" and "genetic knock-in" are used interchangeably herein. A knock-in represents the replacement or insertion of a DNA sequence at a specific DNA sequence in cell by targeting with a Cas protein (by HR, wherein a suitable donor DNA polynucleotide is also used). Examples of knock-ins are a specific insertion of a heterologous amino acid coding sequence in a coding region of a gene, or a specific insertion of a transcriptional regulatory element in a genetic locus.
[0133] Various methods and compositions can be employed to obtain a cell or organism having a polynucleotide of interest inserted in a target site for a Cas endonuclease. Such methods can employ homologous recombination to provide integration of the polynucleotide of Interest at the target site. In one method provided, a polynucleotide of interest is provided to the organism cell in a donor DNA construct. As used herein, "donor DNA" is a DNA construct that comprises a polynucleotide of Interest to be inserted into the target site of a Cas endonuclease. The donor DNA construct further comprises a first and a second region of homology that flank the polynucleotide of Interest. The first and second regions of homology of the donor DNA share homology to a first and a second genomic region, respectively, present in or flanking the target site of the cell or organism genome. By "homology" is meant DNA sequences that are similar. For example, a "region of homology to a genomic region" that is found on the donor DNA is a region of DNA that has a similar sequence to a given "genomic region" in the cell or organism genome. A region of homology can be of any length that is sufficient to promote homologous recombination at the cleaved target site. For example, the region of homology can comprise at least 5-10, 5-15, 5-20, 5-25, 5-30, 5-35, 5-40, 5-45, 5-50, 5-55, 5-60, 5-65, 5-70, 5-75, 5-80, 5-85, 5-90, 5-95, 5-100, 5-200, 5-300, 5-400, 5-500, 5-600, 5-700, 5-800, 5-900, 5-1000, 5-1100, 5-1200, 5-1300, 5-1400, 5-1500, 5-1600, 5-1700, 5-1800, 5-1900, 5-2000, 5-2100, 5-2200, 5-2300, 5-2400, 5-2500, 5-2600, 5-2700, 5-2800, 5-2900, 5-3000, 5-3100 or more bases in length such that the region of homology has sufficient homology to undergo homologous recombination with the corresponding genomic region. "Sufficient homology" indicates that two polynucleotide sequences have sufficient structural similarity to act as substrates for a homologous recombination reaction. The structural similarity includes overall length of each polynucleotide fragment, as well as the sequence similarity of the polynucleotides. Sequence similarity can be described by the percent sequence identity over the whole length of the sequences, and/or by conserved regions comprising localized similarities such as contiguous nucleotides having 100% sequence identity, and percent sequence identity over a portion of the length of the sequences.
[0134] The amount of homology or sequence identity shared by a target and a donor polynucleotide can vary and includes total lengths and/or regions having unit integral values in the ranges of about 1-20 bp, 20-50 bp, 50-100 bp, 75-150 bp, 100-250 bp, 150-300 bp, 200-400 bp, 250-500 bp, 300-600 bp, 350-750 bp, 400-800 bp, 450-900 bp, 500-1000 bp, 600-1250 bp, 700-1500 bp, 800-1750 bp, 900-2000 bp, 1-2.5 kb, 1.5-3 kb, 2-4 kb, 2.5-5 kb, 3-6 kb, 3.5-7 kb, 4-8 kb, 5-10 kb, or up to and including the total length of the target site. These ranges include every integer within the range, for example, the range of 1-20 bp includes 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 and 20 bps. The amount of homology can also described by percent sequence identity over the full aligned length of the two polynucleotides which includes percent sequence identity of about at least 50%, 55%, 60%, 65%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100%. Sufficient homology includes any combination of polynucleotide length, global percent sequence identity, and optionally conserved regions of contiguous nucleotides or local percent sequence identity, for example sufficient homology can be described as a region of 75-150 bp having at least 80% sequence identity to a region of the target locus. Sufficient homology can also be described by the predicted ability of two polynucleotides to specifically hybridize under high stringency conditions, see, for example, Sambrook et al., (1989) Molecular Cloning: A Laboratory Manual, (Cold Spring Harbor Laboratory Press, NY); Current Protocols in Molecular Biology, Ausubel et al., Eds (1994) Current Protocols, (Greene Publishing Associates, Inc. and John Wiley & Sons, Inc.); and, Tijssen (1993) Laboratory Techniques in Biochemistry and Molecular Biology--Hybridization with Nucleic Acid Probes, (Elsevier, New York).
[0135] In one embodiment of the disclosure, the method comprises a method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one guide RNA, at least one donor DNA, and at least one Cas9 endonuclease originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said at least one guide RNA and at least one Cas endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site, wherein said donor DNA comprises a polynucleotide of interest. Cells include, but are not limited to, human, non-human, animal, bacterial, fungal, insect, yeast, and plant cells as well as plants and seeds produced by the methods described herein. The embodiment can further comprise, identifying at least one cell that said polynucleotide of interest integrated in or near said target site.
[0136] As used herein, a "genomic region" is a segment of a chromosome in the genome of a cell that is present on either side of the target site or, alternatively, also comprises a portion of the target site. The genomic region can comprise at least 5-10, 5-15, 5-20, 5-25, 5-30, 5-35, 5-40, 5-45, 5-50, 5-55, 5-60, 5-65, 5-70, 5-75, 5-80, 5-85, 5-90, 5-95, 5-100, 5-200, 5-300, 5-400, 5-500, 5-600, 5-700, 5-800, 5-900, 5-1000, 5-1100, 5-1200, 5-1300, 5-1400, 5-1500, 5-1600, 5-1700, 5-1800, 5-1900, 5-2000, 5-2100, 5-2200, 5-2300, 5-2400, 5-2500, 5-2600, 5-2700, 5-2800. 5-2900, 5-3000, 5-3100 or more bases such that the genomic region has sufficient homology to undergo homologous recombination with the corresponding region of homology.
[0137] Polynucleotides of interest and/or traits can be stacked together in a complex trait locus as described in US-2013-0263324-A1, published 3 Oct. 2013 and in PCT/US13/22891, published Jan. 24, 2013, both applications are hereby incorporated by reference. The guide polynucleotide/Cas9 endonuclease system described herein provides for an efficient system to generate double strand breaks and allows for traits to be stacked in a complex trait locus.
[0138] The guide polynucleotide/Cas endonuclease system can be used for introducing one or more polynucleotides of interest or one or more traits of interest into one or more target sites by providing one or more guide polynucleotides, one Cas endonuclease, and optionally one or more donor DNAs to a plant cell. ((as described in U.S. patent application Ser. No. 14/463,687, file Aug. 20, 2014, incorporated by reference herein). A fertile plant can be produced from that plant cell that comprises an alteration at said one or more target sites, wherein the alteration is selected from the group consisting of (i) replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, and (iv) any combination of (i)-(iii). Plants comprising these altered target sites can be crossed with plants comprising at least one gene or trait of interest in the same complex trait locus, thereby further stacking traits in said complex trait locus. (see also US-2013-0263324-A1, published 3 Oct. 2013 and in PCT/US13/22891, published Jan. 24, 2013).
[0139] The structural similarity between a given genomic region and the corresponding region of homology found on the donor DNA can be any degree of sequence identity that allows for homologous recombination to occur. For example, the amount of homology or sequence identity shared by the "region of homology" of the donor DNA and the "genomic region" of the organism genome can be at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity, such that the sequences undergo homologous recombination
[0140] The region of homology on the donor DNA can have homology to any sequence flanking the target site. While in some embodiments the regions of homology share significant sequence homology to the genomic sequence immediately flanking the target site, it is recognized that the regions of homology can be designed to have sufficient homology to regions that may be further 5' or 3' to the target site. In still other embodiments, the regions of homology can also have homology with a fragment of the target site along with downstream genomic regions. In one embodiment, the first region of homology further comprises a first fragment of the target site and the second region of homology comprises a second fragment of the target site, wherein the first and second fragments are dissimilar.
[0141] As used herein, "homologous recombination" includes the exchange of DNA fragments between two DNA molecules at the sites of homology. The frequency of homologous recombination is influenced by a number of factors. Different organisms vary with respect to the amount of homologous recombination and the relative proportion of homologous to non-homologous recombination. Generally, the length of the region of homology affects the frequency of homologous recombination events: the longer the region of homology, the greater the frequency. The length of the homology region needed to observe homologous recombination is also species-variable. In many cases, at least 5 kb of homology has been utilized, but homologous recombination has been observed with as little as 25-50 bp of homology. See, for example, Singer et al., (1982) Cell 31:25-33; Shen and Huang, (1986) Genetics 112:441-57; Watt et al., (1985) Proc. Natl. Acad. Sci. USA 82:4768-72, Sugawara and Haber, (1992) Mol Cell Biol 12:563-75, Rubnitz and Subramani, (1984) Mol Cell Biol 4:2253-8; Ayares et al., (1986) Proc. Natl. Acad. Sci. USA 83:5199-203; Liskay et al., (1987) Genetics 115:161-7.
[0142] Homology-directed repair (HDR) is a mechanism in cells to repair double-stranded and single stranded DNA breaks. Homology-directed repair includes homologous recombination (HR) and single-strand annealing (SSA) (Lieber. 2010 Annu. Rev. Biochem. 79:181-211). The most common form of HDR is called homologous recombination (HR), which has the longest sequence homology requirements between the donor and acceptor DNA. Other forms of HDR include single-stranded annealing (SSA) and breakage-induced replication, and these require shorter sequence homology relative to HR. Homology-directed repair at nicks (single-stranded breaks) can occur via a mechanism distinct from HDR at double-strand breaks (Davis and Maizels. PNAS (0027-8424), 111 (10), p. E924-E932.
[0143] Alteration of the genome of a plant cell, for example, through homologous recombination (HR), is a powerful tool for genetic engineering. Despite the low frequency of homologous recombination in higher plants, there are a few examples of successful homologous recombination of plant endogenous genes. The parameters for homologous recombination in plants have primarily been investigated by rescuing introduced truncated selectable marker genes. In these experiments, the homologous DNA fragments were typically between 0.3 kb to 2 kb. Observed frequencies for homologous recombination were on the order of 10.sup.-4 to 10.sup.-5. See, for example, Halfter et al., (1992) Mol Gen Genet 231:186-93; Offringa et al., (1990) EMBO J 9:3077-84; Offringa et al., (1993) Proc. Natl. Acad. Sci. USA 90:7346-50; Paszkowski et al., (1988) EMBO J 7:4021-6; Hourda and Paszkowski, (1994) Mol Gen Genet 243:106-11; and Risseeuw et al., (1995) Plant J 7:109-19.
[0144] Homologous recombination has been demonstrated in insects. In Drosophila, Dray and Gloor found that as little as 3 kb of total template:target homology sufficed to copy a large non-homologous segment of DNA into the target with reasonable efficiency (Dray and Gloor, (1997) Genetics 147:689-99). Using FLP-mediated DNA integration at a target FRT in Drosophila, Golic et al., showed integration was approximately 10-fold more efficient when the donor and target shared 4.1 kb of homology as compared to 1.1 kb of homology (Golic et al., (1997) Nucleic Acids Res 25:3665). Data from Drosophila indicates that 2-4 kb of homology is sufficient for efficient targeting, but there is some evidence that much less homology may suffice, on the order of about 30 bp to about 100 bp (Nassif and Engels, (1993) Proc. Natl. Acad. Sci. USA 90:1262-6; Keeler and Gloor, (1997) Mol Cell Biol 17:627-34).
[0145] Homologous recombination has also been accomplished in other organisms. For example, at least 150-200 bp of homology was required for homologous recombination in the parasitic protozoan Leishmania (Papadopoulou and Dumas, (1997) Nucleic Acids Res 25:4278-86). In the filamentous fungus Aspergillus nidulans, gene replacement has been accomplished with as little as 50 bp flanking homology (Chaveroche et al., (2000) Nucleic Acids Res 28:e97). Targeted gene replacement has also been demonstrated in the ciliate Tetrahymena thermophila (Gaertig et al., (1994) Nucleic Acids Res 22:5391-8). In mammals, homologous recombination has been most successful in the mouse using pluripotent embryonic stem cell lines (ES) that can be grown in culture, transformed, selected and introduced into a mouse embryo. Embryos bearing inserted transgenic ES cells develop as genetically offspring. By interbreeding siblings, homozygous mice carrying the selected genes can be obtained. An overview of the process is provided in Watson et al., (1992) Recombinant DNA, 2nd Ed., (Scientific American Books distributed by WH Freeman & Co.); Capecchi, (1989) Trends Genet 5:70-6; and Bronson, (1994) J Biol Chem 269:27155-8. Homologous recombination in mammals other than mouse has been limited by the lack of stem cells capable of being transplanted to oocytes or developing embryos. However, McCreath et al., Nature 405:1066-9 (2000) reported successful homologous recombination in sheep by transformation and selection in primary embryo fibroblast cells.
[0146] Error-prone DNA repair mechanisms can produce mutations at double-strand break sites. The Non-Homologous-End-Joining (NHEJ) pathways are the most common repair mechanism to bring the broken ends together (Bleuyard et al., (2006) DNA Repair 5:1-12). The structural integrity of chromosomes is typically preserved by the repair, but deletions, insertions, or other rearrangements are possible. The two ends of one double-strand break are the most prevalent substrates of NHEJ (Kirik et al., (2000) EMBO J 19:5562-6), however if two different double-strand breaks occur, the free ends from different breaks can be ligated and result in chromosomal deletions (Siebert and Puchta, (2002) Plant Cell 14:1121-31), or chromosomal translocations between different chromosomes (Pacher et al., (2007) Genetics 175:21-9).
[0147] Episomal DNA molecules can also be ligated into the double-strand break, for example, integration of T-DNAs into chromosomal double-strand breaks (Chilton and Que, (2003) Plant Physiol 133:956-65; Salomon and Puchta, (1998) EMBO J 17:6086-95). Once the sequence around the double-strand breaks is altered, for example, by exonuclease activities involved in the maturation of double-strand breaks, gene conversion pathways can restore the original structure if a homologous sequence is available, such as a homologous chromosome in non-dividing somatic cells, or a sister chromatid after DNA replication (Molinier et al., (2004) Plant Cell 16:342-52). Ectopic and/or epigenic DNA sequences may also serve as a DNA repair template for homologous recombination (Puchta, (1999) Genetics 152:1173-81).
[0148] Once a double-strand break is induced in the DNA, the cell's DNA repair mechanism is activated to repair the break. Error-prone DNA repair mechanisms can produce mutations at double-strand break sites. The most common repair mechanism to bring the broken ends together is the nonhomologous end-joining (NHEJ) pathway (Bleuyard et al., (2006) DNA Repair 5:1-12). The structural integrity of chromosomes is typically preserved by the repair, but deletions, insertions, or other rearrangements are possible (Siebert and Puchta, (2002) Plant Cell 14:1121-31; Pacher et al., (2007) Genetics 175:21-9).
[0149] Alternatively, the double-strand break can be repaired by homologous recombination between homologous DNA sequences. Once the sequence around the double-strand break is altered, for example, by exonuclease activities involved in the maturation of double-strand breaks, gene conversion pathways can restore the original structure if a homologous sequence is available, such as a homologous chromosome in non-dividing somatic cells, or a sister chromatid after DNA replication (Molinier et al., (2004) Plant Cell 16:342-52). Ectopic and/or epigenic DNA sequences may also serve as a DNA repair template for homologous recombination (Puchta, (1999) Genetics 152:1173-81).
[0150] DNA double-strand breaks appear to be an effective factor to stimulate homologous recombination pathways (Puchta et al., (1995) Plant Mol Biol 28:281-92; Tzfira and White, (2005) Trends Biotechnol 23:567-9; Puchta, (2005) J Exp Bot 56:1-14). Using DNA-breaking agents, a two- to nine-fold increase of homologous recombination was observed between artificially constructed homologous DNA repeats in plants (Puchta et al., (1995) Plant Mol Biol 28:281-92). In maize protoplasts, experiments with linear DNA molecules demonstrated enhanced homologous recombination between plasmids (Lyznik et al., (1991) Mol Gen Genet 230:209-18).
[0151] The donor DNA may be introduced by any means known in the art. For example, a plant having a target site is provided. The donor DNA may be provided by any transformation method known in the art including, for example, Agrobacterium-mediated transformation or biolistic particle bombardment. The donor DNA may be present transiently in the cell or it could be introduced via a viral replicon. In the presence of the Cas endonuclease and the target site, the donor DNA is inserted into the transformed plant's genome.
[0152] Further uses for guide RNA/Cas endonuclease systems have been described (See U.S. Patent Application US 2015-0082478 A1, published on Mar. 19, 2015, WO2015/026886 A1, published on Feb. 26, 2015, US 2015-0059010 A1, published on Feb. 26, 2015, U.S. application 62/023,246, filed on Jul. 7, 2014, and U.S. application 62/036,652, filed on Aug. 13, 2014, all of which are incorporated by reference herein) and include but are not limited to modifying or replacing nucleotide sequences of interest (such as a regulatory elements), insertion of polynucleotides of interest, gene knock-out, gene-knock in, modification of splicing sites and/or introducing alternate splicing sites, modifications of nucleotide sequences encoding a protein of interest, amino acid and/or protein fusions, and gene silencing by expressing an inverted repeat into a gene of interest.
[0153] Given the diversity of Type II CRISPR-Cas systems (Fonfara et al. (2014) Nucleic Acids Res. 42:2577-2590), it is plausible that many of the Cas9 endonucleases and cognate guide RNAs may have unique sequence recognition and enzymatic properties different from those previously described or characterized. For example, cleavage activity and specificity may be enhanced or proto-spacer adjacent motif (PAM) sequence may be different leading to increased genomic target site density. To tap into this vast unexplored diversity and expand the repertoire of Cas9 endonucleases and cognate guide RNAs available for genome targeting, two components of target site recognition need to be cooperatively characterized for each new system, the PAM sequence and the guide RNA (either duplexed CRISPR RNA (crRNA) and trans-activating CRISPR RNA (tracrRNA) or chimeric fusion of crRNA and tracrRNA (single guide RNA (sgRNA. Rapid in vitro methods described herein have been developed to concertedly characterize both the guide RNA and PAM sequence of Type II Cas9 proteins.
[0154] Methods for assaying Cas9 PAM preferences have been described herein (see Example 3, Example 4 and Example 7). In one embodiment, the Cas9 endonuclease PAM preferences was assayed in a dose dependent manner by subjecting the randomized PAM libraries described herein to in vitro digestion with different concentrations of recombinant Cas9 protein preloaded with guide RNA. After digestion with Cas9-guide RNA ribonucleoprotein (RNP) complexes, PAM sequence combinations from the randomized PAM library that supported cleavage were captured by ligating adapters to the free-ends of the plasmid DNA molecules cleaved by the Cas9-guide RNA complex (FIG. 3). To promote efficient ligation and capture of the cleaved ends, the typically blunt-ended double-stranded DNA cut generated by Cas9 endonucleases was modified to contain a 3' dA overhang and adapters were modified to contain a complementary 3'dT overhang. To generate sufficient quantities of DNA for sequencing, DNA fragments harboring the PAM sequence supporting cleavage were PCR amplified using a primer in the adapter and another directly adjacent to the PAM region. The resulting PCR amplified Cas9 PAM libraries were converted into ampli-seq templates and single-read deep sequenced from the adapter-side of the amplicon. To ensure adequate coverage, the Cas9 PAM libraries were sequenced to a depth at least 5 times greater than the diversity in the initial randomized PAM library (5,120 and 81,920 reads for the 5 and 7 bp PAM randomized libraries, respectively). PAM sequences were identified from the resulting sequence data by only selecting those reads containing a 12 nt sequence match flanking either side of the 5 or 7 nt PAM sequence (depending on the randomized PAM library used); capturing only those PAM sequences resulting from perfect Cas9-guide RNA target site recognition and cleavage. To compensate for the inherent bias in the initial randomized PAM libraries, the frequency of each PAM sequence was normalized to its frequency in the starting library. The composition of the resulting PAM sequences can then be examined using a position frequency matrix (PFM) (Stormo, 2013 Quant. Biol. 1: 115-130)
[0155] As described herein, to validate the randomness of the PAM library disclosed herein (PAM library validation), PCR fragments spanning the 5 bp and 7 bp randomized PAM regions were generated by Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific) amplification (15 cycles of a 2-step amplification protocol) using the primer pair combinations TK-119/pUC-dir and TK-113/pUC-dir (SEQ ID NO: 175/SEQ ID NO:5) for the 5 bp and 7 bp libraries, respectively. The resulting 145 bp PCR product was purified using GeneJET PCR Purification Kit (Thermo Fisher Scientific) and the sequences necessary for amplicon-specific barcodes and IIlumina sequencing were "tailed" on through two rounds of PCR each consisting of 10 cycles. In some examples, the primer pair combinations in the first round of PCR were JKYS800.1/JKYS803 and JKYS921.1 (SEQ ID NO:176)/JKYS812 (SEQ ID NO: 32) for the 5 bp and 7 bp libraries, respectively. A set of primers, JKYS557 (SEQ ID NO: 177)/JKYS558 (SEQ ID NO: 178), universal to all primary PCR reactions were utilized for the secondary PCR amplification. The resulting PCR amplifications were purified with a Qiagen PCR purification spin column, concentration measured with a Hoechst dye-based fluorometric assay, combined in an equimolar ratio and single read 60-100 nucleotide-length deep sequencing was performed on IIlumina's MiSeq Personal Sequencer with a 5-10% (v/v) spike of PhiX control v3 (Illumina, FC-110-3001) to off-set sequence bias. The PAM sequence for only those reads containing a perfect 12 nt sequence match flanking either side of the randomized PAM sequence were captured and used to examine the frequency and diversity of PAM sequences present in the library.
[0156] In one embodiment of the disclosure, the method comprises a method for producing a plasmid DNA library containing a randomized Protospacer-Adjacent-Motif (PAM) sequence, the method comprising: a) providing a first single stranded oligonucleotide comprising a target sequence that can be recognized by a guide RNA/Cas endonuclease complex; b) providing a second single stranded oligonucleotide comprising a randomized PAM sequence adjacent to a nucleotide sequence capable of hybridizing with the target sequence of (a); c) producing an oligoduplex comprising said randomized PAM sequence by combining the first single stranded oligonucleotide of (a) and the second single stranded oligonucleotide of (b); d) producing a ligation product by ligating the oligoduplex from (c) with a linearized plasmid; and, e) transforming host cells with the ligation product of (e) and recovering multiple host cell colonies representing the plasmid library.
[0157] Host cells include, but are not limited to, human, non-human, animal, bacterial, fungal, insect, yeast, non-conventional yeast, and plant cells. One skilled in the art can ligate the oligoduplex of (c) directly into a linearized vector without restriction enzyme digestion, or can use two restriction enzyme sites, one upstream (5') and one downstream (3') of the target site. The first single stranded oligonucleotide can comprise a restriction endonuclease recognition site located upstream of a target sequence and the ligation product of (d) is produced by first cleaving the oligoduplex with a restriction endonuclease that recognizes the restriction endonuclease recognition site of (a) followed by ligating the cleaved oligoduplex from (d) with a linearized plasmid.
[0158] In one embodiment of the disclosure, the method comprises a method for producing a plasmid DNA library containing a randomized Protospacer-Adjacent-Motif (PAM) sequence, the method comprising transforming at least one host cell with a ligation product and recovering multiple host cell colonies representing the plasmid library, wherein said ligation product was generated by contacting a library of linear oligoduplexes with a linearized plasmid, wherein each oligoduplex member of said library of oligoduplexes comprises a first single stranded oligonucleotide comprising a-target sequence, and a second single stranded oligonucleotide comprising a randomized PAM sequence adjacent to a nucleotide sequence capable of hybridizing with said target sequence. One skilled in the art can ligate the oligoduplex of (c) directly into a linearized vector without restriction enzyme digestion, or can use two restriction enzyme sites, one upstream (5') and one downstream (3') of the target site.
[0159] In one embodiment of the disclosure, the method comprises a method for producing a ligation product containing a randomized Protospacer-Adjacent-Motif (PAM) sequence, the method comprising: a) providing a first single stranded oligonucleotide comprising restriction endonuclease recognition site located upstream of a target sequence that can be recognized by a guide RNA/Cas endonuclease complex; b) providing a second single stranded oligonucleotide comprising a randomized PAM sequence adjacent a nucleotide sequence capable of hybridizing with the target sequence of (a); c) producing an oligoduplex comprising said randomized PAM sequence by combining the first single stranded oligonucleotide of (a) and the second single stranded oligonucleotide of (b); and, d) producing a ligation product by ligating the oligoduplex from (c) with a linearized plasmid.
[0160] In one embodiment of the disclosure, the method comprises a method for identification of a Protospacer-Adjacent-Motif (PAM) sequence, the method comprising: a) providing a library of plasmid DNAs, wherein each one of said plasmid DNAs comprises a randomized Protospacer-Adjacent-Motif sequence integrated adjacent to a target sequence that can be recognized by a guide RNA/Cas endonuclease complex; b) providing to said library of plasmids a guide RNA and a Cas endonuclease protein, wherein said guide RNA and Cas endonuclease protein can form a complex that is capable of introducing a double strand break into the said target sequence, thereby creating a library of cleaved targets; c) ligating adaptors to the library of cleaved targets of (b) allowing for the library of cleaved targets to be amplified; d) amplifying the library of cleaved targets such that cleaved products containing the randomized PAM sequence are enriched, thereby producing a library of enriched PAM-sided targets; e) sequencing the library of (a) and the library of enriched PAM-sided targets of (d) and identifying the nucleotide sequence adjacent to the cleaved targets of (b) on either strand of the plasmid DNA, wherein said nucleotide sequence represents a putative Protospacer-Adjacent-Motif sequences; and, f) determining the fold enrichment of each nucleotide within the putative Protospacer-Adjacent-Motif sequence relative to the plasmid DNA library of (a).
[0161] The randomized PAM libraries described herein can also be used in combination with immunoprecipitation then sequencing approach using dCAS9 for further PAM discovery. The randomized PAM libraries can also be put on a microchip followed by cleaving the chip-array library. The randomized PAM libraries described herein can also be used in combination with Phage-display as a method to identify PAMs. (Isalan, M., Klug, A. and Choo, Y. (2001) A rapid, generally applicable method to engineer zinc fingers illustrated by targeting the HIV-1 120 promoter. Nat. Biotechnol., 19, 656-660; Dreier, B., Fuller, R. P., Segal, D. J., Lund, C., Blancafort, P., Huber, A., Koksch, B. and Barbas, C. F., III (2005) Development of zinc finger domains for recognition of the 50-CNN-30 family DNA sequences and their use in the construction of artificial transcription factors. J. Biol. 125 Chem., 280, 35588-35597).
[0162] In one embodiment of the disclosure, the method comprises a method for identification of a tracrRNA of an organism, the method comprising: a) providing a first single guide RNA candidate comprising a chimeric non-naturally occurring crRNA comprising a variable targeting domain capable of hybridizing to a target sequence in the genome of a cell, linked to a first nucleotide sequence representing the sense expression of a candidate tracrRNA naturally occurring in said organism; b) providing a second single guide RNA candidate comprising a chimeric non-naturally occurring crRNA comprising a variable targeting domain capable of hybridizing to a target sequence in the genome of said cell, linked to a second nucleotide sequence representing the sense expression of a candidate tracrRNA naturally occurring in said organism; c) providing to the first and second single guide RNA candidates a Cas endonuclease protein, wherein said Cas endonuclease protein can form a complex with either the first single guide RNA candidate or the second single guide RNA candidate, wherein said complex is capable of introducing a double strand break into said target sequence; and d) identification of the first or second guide RNA candidate and its tracrRNA component that complexes to the Cas endonuclease of (c) and results in cleavage of the target sequence in the genome of said cell.
[0163] In one embodiment of the disclosure, the method comprises a method for identification of a tracrRNA of an organism, the method comprising: a) identifying a CRISPR array repeat sequence in a genomic locus of said organism; b) aligning the CRISPR array repeat sequence of (a) with the sequence of the genomic locus of (a) and identifying an antirepeat sequence that encodes a tracrRNA; and, c) determining the transcriptional direction of the tracrRNA.
[0164] In one embodiment of the disclosure, the method comprises a method for designing a single guide RNA, the method comprising: a) aligning a tracrRNA sequence with a CRISPR array repeat sequence from a genomic locus of an organism, wherein said CRISPR array repeat sequence comprises a crRNA sequence; b) deducing the transcriptional direction of the CRISPR array, thereby also deducing the crRNA sequence; and, c) designing a single guide RNA comprising said tracrRNA and crRNA sequences.
[0165] In one embodiment of the disclosure, the method comprises a method for producing target sequences, the method comprising: a) identifying a polynucleotides of interest; b) introducing a Protospacer-Adjacent-Motif (PAM) sequence adjacent to said polynucleotide of interest, wherein said PAM sequence comprises the nucleotide sequence NNNNCND, thereby creating a thereby creating a target site for a guide RNA/Cas9 endonuclease complex; and, c) identifying a polynucleotides of interest.
[0166] Polynucleotides of interest are further described herein and include polynucleotides reflective of the commercial markets and interests of those involved in the development of the crop. Crops and markets of interest change, and as developing nations open up world markets, new crops and technologies will emerge also. In addition, as our understanding of agronomic traits and characteristics such as yield and heterosis increase, the choice of genes for genetic engineering will change accordingly.
[0167] Further provided are methods for identifying at least one plant cell, comprising in its genome, a polynucleotide of interest integrated at the target site. A variety of methods are available for identifying those plant cells with insertion into the genome at or near to the target site without using a screenable marker phenotype. Such methods can be viewed as directly analyzing a target sequence to detect any change in the target sequence, including but not limited to PCR methods, sequencing methods, nuclease digestion, Southern blots, and any combination thereof. See, for example, U.S. patent application Ser. No. 12/147,834, herein incorporated by reference to the extent necessary for the methods described herein. The method also comprises recovering a plant from the plant cell comprising a polynucleotide of Interest integrated into its genome. The plant may be sterile or fertile. It is recognized that any polynucleotide of interest can be provided, integrated into the plant genome at the target site, and expressed in a plant.
[0168] Polynucleotides/polypeptides of interest include, but are not limited to, herbicide-resistance coding sequences, insecticidal coding sequences, nematicidal coding sequences, antimicrobial coding sequences, antifungal coding sequences, antiviral coding sequences, abiotic and biotic stress tolerance coding sequences, or sequences modifying plant traits such as yield, grain quality, nutrient content, starch quality and quantity, nitrogen fixation and/or utilization, fatty acids, and oil content and/or composition. More specific polynucleotides of interest include, but are not limited to, genes that improve crop yield, polypeptides that improve desirability of crops, genes encoding proteins conferring resistance to abiotic stress, such as drought, nitrogen, temperature, salinity, toxic metals or trace elements, or those conferring resistance to toxins such as pesticides and herbicides, or to biotic stress, such as attacks by fungi, viruses, bacteria, insects, and nematodes, and development of diseases associated with these organisms. General categories of genes of interest include, for example, those genes involved in information, such as zinc fingers, those involved in communication, such as kinases, and those involved in housekeeping, such as heat shock proteins. More specific categories of transgenes, for example, include genes encoding important traits for agronomics, insect resistance, disease resistance, herbicide resistance, fertility or sterility, grain characteristics, and commercial products. Genes of interest include, generally, those involved in oil, starch, carbohydrate, or nutrient metabolism as well as those affecting kernel size, sucrose loading, and the like that can be stacked or used in combination with other traits, such as but not limited to herbicide resistance, described herein.
[0169] Agronomically important traits such as oil, starch, and protein content can be genetically altered in addition to using traditional breeding methods. Modifications include increasing content of oleic acid, saturated and unsaturated oils, increasing levels of lysine and sulfur, providing essential amino acids, and also modification of starch. Hordothionin protein modifications are described in U.S. Pat. Nos. 5,703,049, 5,885,801, 5,885,802, and 5,990,389, herein incorporated by reference.
[0170] Polynucleotide sequences of interest may encode proteins involved in providing disease or pest resistance. By "disease resistance" or "pest resistance" is intended that the plants avoid the harmful symptoms that are the outcome of the plant-pathogen interactions. Pest resistance genes may encode resistance to pests that have great yield drag such as rootworm, cutworm, European Corn Borer, and the like. Disease resistance and insect resistance genes such as lysozymes or cecropins for antibacterial protection, or proteins such as defensins, glucanases or chitinases for antifungal protection, or Bacillus thuringiensis endotoxins, protease inhibitors, collagenases, lectins, or glycosidases for controlling nematodes or insects are all examples of useful gene products. Genes encoding disease resistance traits include detoxification genes, such as against fumonisin (U.S. Pat. No. 5,792,931); avirulence (avr) and disease resistance (R) genes (Jones et al. (1994) Science 266:789; Martin et al. (1993) Science 262:1432; and Mindrinos et al. (1994) Cell 78:1089); and the like. Insect resistance genes may encode resistance to pests that have great yield drag such as rootworm, cutworm, European Corn Borer, and the like. Such genes include, for example, Bacillus thuringiensis toxic protein genes (U.S. Pat. Nos. 5,366,892; 5,747,450; 5,736,514; 5,723,756; 5,593,881; and Geiser et al. (1986) Gene 48:109); and the like.
[0171] An "herbicide resistance protein" or a protein resulting from expression of an "herbicide resistance-encoding nucleic acid molecule" includes proteins that confer upon a cell the ability to tolerate a higher concentration of an herbicide than cells that do not express the protein, or to tolerate a certain concentration of an herbicide for a longer period of time than cells that do not express the protein. Herbicide resistance traits may be introduced into plants by genes coding for resistance to herbicides that act to inhibit the action of acetolactate synthase (ALS), in particular the sulfonylurea-type herbicides, genes coding for resistance to herbicides that act to inhibit the action of glutamine synthase, such as phosphinothricin or basta (e.g., the bar gene), glyphosate (e.g., the EPSP synthase gene and the GAT gene), HPPD inhibitors (e.g, the HPPD gene) or other such genes known in the art. See, for example, U.S. Pat. Nos. 7,626,077, 5,310,667, 5,866,775, 6,225,114, 6,248,876, 7,169,970, 6,867,293, and U.S. Provisional Application No. 61/401,456, each of which is herein incorporated by reference. The bar gene encodes resistance to the herbicide basta, the nptll gene encodes resistance to the antibiotics kanamycin and geneticin, and the ALS-gene mutants encode resistance to the herbicide chlorsulfuron.
[0172] Sterility genes can also be encoded in an expression cassette and provide an alternative to physical detasseling. Examples of genes used in such ways include male fertility genes such as MS26 (see for example U.S. Pat. Nos. 7,098,388, 7,517,975, 7,612,251), MS45 (see for example U.S. Pat. Nos. 5,478,369, 6,265,640) or MSCA1 (see for example U.S. Pat. No. 7,919,676). Maize plants (Zea mays L.) can be bred by both self-pollination and cross-pollination techniques. Maize has male flowers, located on the tassel, and female flowers, located on the ear, on the same plant. It can self-pollinate ("selfing") or cross pollinate. Natural pollination occurs in maize when wind blows pollen from the tassels to the silks that protrude from the tops of the incipient ears. Pollination may be readily controlled by techniques known to those of skill in the art. The development of maize hybrids requires the development of homozygous inbred lines, the crossing of these lines, and the evaluation of the crosses. Pedigree breeding and recurrent selections are two of the breeding methods used to develop inbred lines from populations. Breeding programs combine desirable traits from two or more inbred lines or various broad-based sources into breeding pools from which new inbred lines are developed by selfing and selection of desired phenotypes. A hybrid maize variety is the cross of two such inbred lines, each of which may have one or more desirable characteristics lacked by the other or which complement the other. The new inbreds are crossed with other inbred lines and the hybrids from these crosses are evaluated to determine which have commercial potential. The hybrid progeny of the first generation is designated F1. The F1 hybrid is more vigorous than its inbred parents. This hybrid vigor, or heterosis, can be manifested in many ways, including increased vegetative growth and increased yield.
[0173] Hybrid maize seed can be produced by a male sterility system incorporating manual detasseling. To produce hybrid seed, the male tassel is removed from the growing female inbred parent, which can be planted in various alternating row patterns with the male inbred parent. Consequently, providing that there is sufficient isolation from sources of foreign maize pollen, the ears of the female inbred will be fertilized only with pollen from the male inbred. The resulting seed is therefore hybrid (F1) and will form hybrid plants.
[0174] Field variation impacting plant development can result in plants tasseling after manual detasseling of the female parent is completed. Or, a female inbred plant tassel may not be completely removed during the detasseling process. In any event, the result is that the female plant will successfully shed pollen and some female plants will be self-pollinated. This will result in seed of the female inbred being harvested along with the hybrid seed which is normally produced. Female inbred seed does not exhibit heterosis and therefore is not as productive as F1 seed. In addition, the presence of female inbred seed can represent a germplasm security risk for the company producing the hybrid.
[0175] Alternatively, the female inbred can be mechanically detasseled by machine. Mechanical detasseling is approximately as reliable as hand detasseling, but is faster and less costly. However, most detasseling machines produce more damage to the plants than hand detasseling. Thus, no form of detasseling is presently entirely satisfactory, and a need continues to exist for alternatives which further reduce production costs and to eliminate self-pollination of the female parent in the production of hybrid seed.
[0176] Mutations that cause male sterility in plants have the potential to be useful in methods for hybrid seed production for crop plants such as maize and can lower production costs by eliminating the need for the labor-intensive removal of male flowers (also known as de-tasseling) from the maternal parent plants used as a hybrid parent. Mutations that cause male sterility in maize have been produced by a variety of methods such as X-rays or UV-irradiations, chemical treatments, or transposable element insertions (ms23, ms25, ms26, ms32) (Chaubal et al. (2000) Am J Bot 87:1193-1201). Conditional regulation of fertility genes through fertility/sterility "molecular switches" could enhance the options for designing new male-sterility systems for crop improvement (Unger et al. (2002) Transgenic Res 11:455-465).
[0177] Besides identification of novel genes impacting male fertility, there remains a need to provide a reliable system of producing genetic male sterility.
[0178] In U.S. Pat. No. 5,478,369, a method is described by which the Ms45 male fertility gene was tagged and cloned on maize chromosome 9. Previously, there had been described a male fertility gene on chromosome 9, ms2, which had never been cloned and sequenced. It is not allelic to the gene referred to in the '369 patent. See Albertsen, M. and Phillips, R. L., "Developmental Cytology of 13 Genetic Male Sterile Loci in Maize" Canadian Journal of Genetics & Cytology 23:195-208 (January 1981). The only fertility gene cloned before that had been the Arabidopsis gene described at Aarts, et al., supra.
[0179] Furthermore, it is recognized that the polynucleotide of interest may also comprise antisense sequences complementary to at least a portion of the messenger RNA (mRNA) for a targeted gene sequence of interest. Antisense nucleotides are constructed to hybridize with the corresponding mRNA. Modifications of the antisense sequences may be made as long as the sequences hybridize to and interfere with expression of the corresponding mRNA. In this manner, antisense constructions having 70%, 80%, or 85% sequence identity to the corresponding antisense sequences may be used. Furthermore, portions of the antisense nucleotides may be used to disrupt the expression of the target gene. Generally, sequences of at least 50 nucleotides, 100 nucleotides, 200 nucleotides, or greater may be used.
[0180] In addition, the polynucleotide of interest may also be used in the sense orientation to suppress the expression of endogenous genes in plants. Methods for suppressing gene expression in plants using polynucleotides in the sense orientation are known in the art. The methods generally involve transforming plants with a DNA construct comprising a promoter that drives expression in a plant operably linked to at least a portion of a nucleotide sequence that corresponds to the transcript of the endogenous gene. Typically, such a nucleotide sequence has substantial sequence identity to the sequence of the transcript of the endogenous gene, generally greater than about 65% sequence identity, about 85% sequence identity, or greater than about 95% sequence identity. See, U.S. Pat. Nos. 5,283,184 and 5,034,323; herein incorporated by reference.
[0181] The polynucleotide of interest can also be a phenotypic marker. A phenotypic marker is screenable or a selectable marker that includes visual markers and selectable markers whether it is a positive or negative selectable marker. Any phenotypic marker can be used. Specifically, a selectable or screenable marker comprises a DNA segment that allows one to identify, or select for or against a molecule or a cell that contains it, often under particular conditions. These markers can encode an activity, such as, but not limited to, production of RNA, peptide, or protein, or can provide a binding site for RNA, peptides, proteins, inorganic and organic compounds or compositions and the like.
[0182] Examples of selectable markers include, but are not limited to, DNA segments that comprise restriction enzyme sites; DNA segments that encode products which provide resistance against otherwise toxic compounds including antibiotics, such as, spectinomycin, ampicillin, kanamycin, tetracycline, Basta, neomycin phosphotransferase II (NEO) and hygromycin phosphotransferase (HPT)); DNA segments that encode products which are otherwise lacking in the recipient cell (e.g., tRNA genes, auxotrophic markers); DNA segments that encode products which can be readily identified (e.g., phenotypic markers such as .beta.-galactosidase, GUS; fluorescent proteins such as green fluorescent protein (GFP), cyan (CFP), yellow (YFP), red (RFP), and cell surface proteins); the generation of new primer sites for PCR (e.g., the juxtaposition of two DNA sequence not previously juxtaposed), the inclusion of DNA sequences not acted upon or acted upon by a restriction endonuclease or other DNA modifying enzyme, chemical, etc.; and, the inclusion of a DNA sequences required for a specific modification (e.g., methylation) that allows its identification.
[0183] Additional selectable markers include genes that confer resistance to herbicidal compounds, such as glufosinate ammonium, bromoxynil, imidazolinones, and 2,4-dichlorophenoxyacetate (2,4-D). See for example, Yarranton, (1992) Curr Opin Biotech 3:506-11; Christopherson et al., (1992) Proc. Natl. Acad. Sci. USA 89:6314-8; Yao et al., (1992) Cell 71:63-72; Reznikoff, (1992) Mol Microbiol 6:2419-22; Hu et al., (1987) Cell 48:555-66; Brown et al., (1987) Cell 49:603-12; Figge et al., (1988) Cell 52:713-22; Deuschle et al., (1989) Proc. Natl. Acad. Sci. USA 86:5400-4; Fuerst et al., (1989) Proc. Natl. Acad. Sci. USA 86:2549-53; Deuschle et al., (1990) Science 248:480-3; Gossen, (1993) Ph.D. Thesis, University of Heidelberg; Reines et al., (1993) Proc. Natl. Acad. Sci. USA 90:1917-21; Labow et al., (1990) Mol Cell Biol 10:3343-56; Zambretti et al., (1992) Proc. Natl. Acad. Sci. USA 89:3952-6; Baim et al., (1991) Proc. Natl. Acad. Sci. USA 88:5072-6; Wyborski et al., (1991) Nucleic Acids Res 19:4647-53; Hillen and Wissman, (1989) Topics Mol Struc Biol 10:143-62; Degenkolb et al., (1991) Antimicrob Agents Chemother 35:1591-5; Kleinschnidt et al., (1988) Biochemistry 27:1094-104; Bonin, (1993) Ph.D. Thesis, University of Heidelberg; Gossen et al., (1992) Proc. Natl. Acad. Sci. USA 89:5547-51; Oliva et al., (1992) Antimicrob Agents Chemother 36:913-9; Hlavka et al., (1985) Handbook of Experimental Pharmacology, Vol. 78 (Springer-Verlag, Berlin); Gill et al., (1988) Nature 334:721-4. Commercial traits can also be encoded on a gene or genes that could increase for example, starch for ethanol production, or provide expression of proteins. Another important commercial use of transformed plants is the production of polymers and bioplastics such as described in U.S. Pat. No. 5,602,321. Genes such as .beta.-Ketothiolase, PHBase (polyhydroxyburyrate synthase), and acetoacetyl-CoA reductase (see Schubert et al. (1988) J. Bacteriol. 170:5837-5847) facilitate expression of polyhyroxyalkanoates (PHAs).
[0184] Exogenous products include plant enzymes and products as well as those from other sources including prokaryotes and other eukaryotes. Such products include enzymes, cofactors, hormones, and the like. The level of proteins, particularly modified proteins having improved amino acid distribution to improve the nutrient value of the plant, can be increased. This is achieved by the expression of such proteins having enhanced amino acid content.
[0185] The transgenes, recombinant DNA molecules, DNA sequences of interest, and polynucleotides of interest can be comprise one or more DNA sequences for gene silencing. Methods for gene silencing involving the expression of DNA sequences in plant are known in the art include, but are not limited to, cosuppression, antisense suppression, double-stranded RNA (dsRNA) interference, hairpin RNA (hpRNA) interference, intron-containing hairpin RNA (ihpRNA) interference, transcriptional gene silencing, and micro RNA (miRNA) interference
[0186] As used herein, "nucleic acid" means a polynucleotide and includes a single or a double-stranded polymer of deoxyribonucleotide or ribonucleotide bases. Nucleic acids may also include fragments and modified nucleotides. Thus, the terms "polynucleotide", "nucleic acid sequence", "nucleotide sequence" and "nucleic acid fragment" are used interchangeably to denote a polymer of RNA and/or DNA that is single- or double-stranded, optionally containing synthetic, non-natural, or altered nucleotide bases. Nucleotides (usually found in their 5'-monophosphate form) are referred to by their single letter designation as follows: "A" for adenosine or deoxyadenosine (for RNA or DNA, respectively), "C" for cytosine or deoxycytosine, "G" for guanosine or deoxyguanosine, "U" for uridine, "T" for deoxythymidine, "R" for purines (A or G), "Y" for pyrimidines (C or T), "K" for G or T, "H" for A or C or T, "I" for inosine, and "N" for any nucleotide.
[0187] "Open reading frame" is abbreviated ORF.
[0188] The terms "subfragment that is functionally equivalent" and "functionally equivalent subfragment" are used interchangeably herein. These terms refer to a portion or subsequence of an isolated nucleic acid fragment in which the ability to alter gene expression or produce a certain phenotype is retained whether or not the fragment or subfragment encodes an active enzyme. For example, the fragment or subfragment can be used in the design of genes to produce the desired phenotype in a transformed plant. Genes can be designed for use in suppression by linking a nucleic acid fragment or subfragment thereof, whether or not it encodes an active enzyme, in the sense or antisense orientation relative to a plant promoter sequence.
[0189] The term "conserved domain" or "motif" means a set of amino acids conserved at specific positions along an aligned sequence of evolutionarily related proteins. While amino acids at other positions can vary between homologous proteins, amino acids that are highly conserved at specific positions indicate amino acids that are essential to the structure, the stability, or the activity of a protein. Because they are identified by their high degree of conservation in aligned sequences of a family of protein homologues, they can be used as identifiers, or "signatures", to determine if a protein with a newly determined sequence belongs to a previously identified protein family.
[0190] Polynucleotide and polypeptide sequences, variants thereof, and the structural relationships of these sequences can be described by the terms "homology", "homologous", "substantially identical", "substantially similar" and "corresponding substantially" which are used interchangeably herein. These refer to polypeptide or nucleic acid fragments wherein changes in one or more amino acids or nucleotide bases do not affect the function of the molecule, such as the ability to mediate gene expression or to produce a certain phenotype. These terms also refer to modification(s) of nucleic acid fragments that do not substantially alter the functional properties of the resulting nucleic acid fragment relative to the initial, unmodified fragment. These modifications include deletion, substitution, and/or insertion of one or more nucleotides in the nucleic acid fragment.
[0191] Substantially similar nucleic acid sequences encompassed may be defined by their ability to hybridize (under moderately stringent conditions, e.g., 0.5.times.SSC, 0.1% SDS, 60.degree. C.) with the sequences exemplified herein, or to any portion of the nucleotide sequences disclosed herein and which are functionally equivalent to any of the nucleic acid sequences disclosed herein. Stringency conditions can be adjusted to screen for moderately similar fragments, such as homologous sequences from distantly related organisms, to highly similar fragments, such as genes that duplicate functional enzymes from closely related organisms. Post-hybridization washes determine stringency conditions.
[0192] The term "selectively hybridizes" includes reference to hybridization, under stringent hybridization conditions, of a nucleic acid sequence to a specified nucleic acid target sequence to a detectably greater degree (e.g., at least 2-fold over background) than its hybridization to non-target nucleic acid sequences and to the substantial exclusion of non-target nucleic acids. Selectively hybridizing sequences typically have about at least 80% sequence identity, or 90% sequence identity, up to and including 100% sequence identity (i.e., fully complementary) with each other.
[0193] The term "stringent conditions" or "stringent hybridization conditions" includes reference to conditions under which a probe will selectively hybridize to its target sequence in an in vitro hybridization assay. Stringent conditions are sequence-dependent and will be different in different circumstances. By controlling the stringency of the hybridization and/or washing conditions, target sequences can be identified which are 100% complementary to the probe (homologous probing). Alternatively, stringency conditions can be adjusted to allow some mismatching in sequences so that lower degrees of similarity are detected (heterologous probing). Generally, a probe is less than about 1000 nucleotides in length, optionally less than 500 nucleotides in length.
[0194] Typically, stringent conditions will be those in which the salt concentration is less than about 1.5 M Na ion, typically about 0.01 to 1.0 M Na ion concentration (or other salt(s)) at pH 7.0 to 8.3, and at least about 30.degree. C. for short probes (e.g., 10 to 50 nucleotides) and at least about 60.degree. C. for long probes (e.g., greater than 50 nucleotides). Stringent conditions may also be achieved with the addition of destabilizing agents such as formamide. Exemplary low stringency conditions include hybridization with a buffer solution of 30 to 35% formamide, 1 M NaCl, 1% SDS (sodium dodecyl sulphate) at 37.degree. C., and a wash in 1.times. to 2.times.SSC (20.times.SSC=3.0 M NaCl/0.3 M trisodium citrate) at 50 to 55.degree. C. Exemplary moderate stringency conditions include hybridization in 40 to 45% formamide, 1 M NaCl, 1% SDS at 37.degree. C., and a wash in 0.5.times. to 1.times.SSC at 55 to 60.degree. C. Exemplary high stringency conditions include hybridization in 50% formamide, 1 M NaCl, 1% SDS at 37.degree. C., and a wash in 0.1.times.SSC at 60 to 65.degree. C.
[0195] "Sequence identity" or "identity" in the context of nucleic acid or polypeptide sequences refers to the nucleic acid bases or amino acid residues in two sequences that are the same when aligned for maximum correspondence over a specified comparison window.
[0196] The term "percentage of sequence identity" refers to the value determined by comparing two optimally aligned sequences over a comparison window, wherein the portion of the polynucleotide or polypeptide sequence in the comparison window may comprise additions or deletions (i.e., gaps) as compared to the reference sequence (which does not comprise additions or deletions) for optimal alignment of the two sequences. The percentage is calculated by determining the number of positions at which the identical nucleic acid base or amino acid residue occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison and multiplying the results by 100 to yield the percentage of sequence identity. Useful examples of percent sequence identities include, but are not limited to, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90% or 95%, or any integer percentage from 50% to 100%. These identities can be determined using any of the programs described herein.
[0197] Sequence alignments and percent identity or similarity calculations may be determined using a variety of comparison methods designed to detect homologous sequences including, but not limited to, the MegAlign.TM. program of the LASERGENE bioinformatics computing suite (DNASTAR Inc., Madison, Wis.). Within the context of this application it will be understood that where sequence analysis software is used for analysis, that the results of the analysis will be based on the "default values" of the program referenced, unless otherwise specified. As used herein "default values" will mean any set of values or parameters that originally load with the software when first initialized.
[0198] The "Clustal V method of alignment" corresponds to the alignment method labeled Clustal V (described by Higgins and Sharp, (1989) CABIOS 5:151-153; Higgins et al., (1992) Comput Appl Biosci 8:189-191) and found in the MegAlign.TM. program of the LASERGENE bioinformatics computing suite (DNASTAR Inc., Madison, Wis.). For multiple alignments, the default values correspond to GAP PENALTY=10 and GAP LENGTH PENALTY=10. Default parameters for pairwise alignments and calculation of percent identity of protein sequences using the Clustal method are KTUPLE=1, GAP PENALTY=3, WINDOW=5 and DIAGONALS SAVED=5. For nucleic acids these parameters are KTUPLE=2, GAP PENALTY=5, WINDOW=4 and DIAGONALS SAVED=4. After alignment of the sequences using the Clustal V program, it is possible to obtain a "percent identity" by viewing the "sequence distances" table in the same program.
[0199] The "Clustal W method of alignment" corresponds to the alignment method labeled Clustal W (described by Higgins and Sharp, (1989) CABIOS 5:151-153; Higgins et al., (1992) Comput Appl Biosci 8:189-191) and found in the MegAlign.TM. v6.1 program of the LASERGENE bioinformatics computing suite (DNASTAR Inc., Madison, Wis.). Default parameters for multiple alignment (GAP PENALTY=10, GAP LENGTH PENALTY=0.2, Delay Divergen Seqs (%)=30, DNA Transition Weight=0.5, Protein Weight Matrix=Gonnet Series, DNA Weight Matrix=IUB). After alignment of the sequences using the Clustal W program, it is possible to obtain a "percent identity" by viewing the "sequence distances" table in the same program.
[0200] Unless otherwise stated, sequence identity/similarity values provided herein refer to the value obtained using GAP Version 10 (GCG, Accelrys, San Diego, Calif.) using the following parameters: % identity and % similarity for a nucleotide sequence using a gap creation penalty weight of 50 and a gap length extension penalty weight of 3, and the nwsgapdna.cmp scoring matrix; % identity and % similarity for an amino acid sequence using a GAP creation penalty weight of 8 and a gap length extension penalty of 2, and the BLOSUM62 scoring matrix (Henikoff and Henikoff, (1989) Proc. Natl. Acad. Sci. USA 89:10915). GAP uses the algorithm of Needleman and Wunsch, (1970) J Mol Biol 48:443-53, to find an alignment of two complete sequences that maximizes the number of matches and minimizes the number of gaps. GAP considers all possible alignments and gap positions and creates the alignment with the largest number of matched bases and the fewest gaps, using a gap creation penalty and a gap extension penalty in units of matched bases.
[0201] "BLAST" is a searching algorithm provided by the National Center for Biotechnology Information (NCBI) used to find regions of similarity between biological sequences. The program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches to identify sequences having sufficient similarity to a query sequence such that the similarity would not be predicted to have occurred randomly. BLAST reports the identified sequences and their local alignment to the query sequence.
[0202] It is well understood by one skilled in the art that many levels of sequence identity are useful in identifying polypeptides from other species or modified naturally or synthetically wherein such polypeptides have the same or similar function or activity. Useful examples of percent identities include, but are not limited to, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90% or 95%, or any integer percentage from 50% to 100%. Indeed, any integer amino acid identity from 50% to 100% may be useful in describing the present disclosure, such as 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99%.
[0203] "Gene" includes a nucleic acid fragment that expresses a functional molecule such as, but not limited to, a specific protein, including regulatory sequences preceding (5' non-coding sequences) and following (3' non-coding sequences) the coding sequence. "Native gene" refers to a gene as found in nature with its own regulatory sequences.
[0204] A "mutated gene" is a gene that has been altered through human intervention. Such a "mutated gene" has a sequence that differs from the sequence of the corresponding non-mutated gene by at least one nucleotide addition, deletion, or substitution. In certain embodiments of the disclosure, the mutated gene comprises an alteration that results from a guide polynucleotide/Cas endonuclease system as disclosed herein. A mutated plant is a plant comprising a mutated gene.
[0205] As used herein, a "targeted mutation" is a mutation in a native gene that was made by altering a target sequence within the native gene using a method involving a double-strand-break-inducing agent that is capable of inducing a double-strand break in the DNA of the target sequence as disclosed herein or known in the art.
[0206] The guide RNA/Cas endonuclease induced targeted mutation can occur in a nucleotide sequence that is located within or outside a genomic target site that is recognized and cleaved by a Cas endonuclease.
[0207] The term "genome" as it applies to a plant cells encompasses not only chromosomal DNA found within the nucleus, but organelle DNA found within subcellular components (e.g., mitochondria, or plastid) of the cell.
[0208] A "codon-modified gene" or "codon-preferred gene" or "codon-optimized gene" is a gene having its frequency of codon usage designed to mimic the frequency of preferred codon usage of the host cell.
[0209] An "allele" is one of several alternative forms of a gene occupying a given locus on a chromosome. When all the alleles present at a given locus on a chromosome are the same, that plant is homozygous at that locus. If the alleles present at a given locus on a chromosome differ, that plant is heterozygous at that locus.
[0210] "Coding sequence" refers to a polynucleotide sequence which codes for a specific amino acid sequence. "Regulatory sequences" refer to nucleotide sequences located upstream (5' non-coding sequences), within, or downstream (3' non-coding sequences) of a coding sequence, and which influence the transcription, RNA processing or stability, or translation of the associated coding sequence. Regulatory sequences may include, but are not limited to: promoters, translation leader sequences, 5' untranslated sequences, 3' untranslated sequences, introns, polyadenylation target sequences, RNA processing sites, effector binding sites, and stem-loop structures.
[0211] "A plant-optimized nucleotide sequence" is nucleotide sequence that has been optimized for increased expression in plants, particularly for increased expression in plants or in one or more plants of interest. For example, a plant-optimized nucleotide sequence can be synthesized by modifying a nucleotide sequence encoding a protein such as, for example, double-strand-break-inducing agent (e.g., an endonuclease) as disclosed herein, using one or more plant-preferred codons for improved expression. See, for example, Campbell and Gowri (1990) Plant Physiol. 92:1-11 for a discussion of host-preferred codon usage.
[0212] Methods are available in the art for synthesizing plant-preferred genes. See, for example, U.S. Pat. Nos. 5,380,831, and 5,436,391, and Murray et al. (1989) Nucleic Acids Res. 17:477-498, herein incorporated by reference. Additional sequence modifications are known to enhance gene expression in a plant host. These include, for example, elimination of: one or more sequences encoding spurious polyadenylation signals, one or more exon-intron splice site signals, one or more transposon-like repeats, and other such well-characterized sequences that may be deleterious to gene expression. The G-C content of the sequence may be adjusted to levels average for a given plant host, as calculated by reference to known genes expressed in the host plant cell. When possible, the sequence is modified to avoid one or more predicted hairpin secondary mRNA structures. Thus, "a plant-optimized nucleotide sequence" of the present disclosure comprises one or more of such sequence modifications.
[0213] A promoter is a region of DNA involved in recognition and binding of RNA polymerase and other proteins to initiate transcription. The promoter sequence consists of proximal and more distal upstream elements, the latter elements often referred to as enhancers. An "enhancer" is a DNA sequence that can stimulate promoter activity, and may be an innate element of the promoter or a heterologous element inserted to enhance the level or tissue-specificity of a promoter. Promoters may be derived in their entirety from a native gene, or be composed of different elements derived from different promoters found in nature, and/or comprise synthetic DNA segments. It is understood by those skilled in the art that different promoters may direct the expression of a gene in different tissues or cell types, or at different stages of development, or in response to different environmental conditions. It is further recognized that since in most cases the exact boundaries of regulatory sequences have not been completely defined, DNA fragments of some variation may have identical promoter activity. Promoters that cause a gene to be expressed in most cell types at most times are commonly referred to as "constitutive promoters".
[0214] It has been shown that certain promoters are able to direct RNA synthesis at a higher rate than others. These are called "strong promoters". Certain other promoters have been shown to direct RNA synthesis at higher levels only in particular types of cells or tissues and are often referred to as "tissue specific promoters", or "tissue-preferred promoters" if the promoters direct RNA synthesis preferably in certain tissues but also in other tissues at reduced levels. Since patterns of expression of a chimeric gene (or genes) introduced into a plant are controlled using promoters, there is an ongoing interest in the isolation of novel promoters which are capable of controlling the expression of a chimeric gene or (genes) at certain levels in specific tissue types or at specific plant developmental stages.
[0215] A plant promoter can include a promoter capable of initiating transcription in a plant cell, for a review of plant promoters, see, Potenza et al., (2004) In Vitro Cell Dev Biol 40:1-22. Constitutive promoters include, for example, the core promoter of the Rsyn7 promoter and other constitutive promoters disclosed in WO99/43838 and U.S. Pat. No. 6,072,050; the core CaMV 35S promoter (Odell et al., (1985) Nature 313:810-2); rice actin (McElroy et al., (1990) Plant Cell 2:163-71); ubiquitin (Christensen et al., (1989) Plant Mol Biol 12:619-32; Christensen et al., (1992) Plant Mol Biol 18:675-89); pEMU (Last et al., (1991) Theor Appl Genet 81:581-8); MAS (Velten et al., (1984) EMBO J 3:2723-30); ALS promoter (U.S. Pat. No. 5,659,026), and the like. Other constitutive promoters are described in, for example, U.S. Pat. Nos. 5,608,149; 5,608,144; 5,604,121; 5,569,597; 5,466,785; 5,399,680; 5,268,463; 5,608,142 and 6,177,611. In some examples an inducible promoter may be used. Pathogen-inducible promoters induced following infection by a pathogen include, but are not limited to those regulating expression of PR proteins, SAR proteins, beta-1,3-glucanase, chitinase, etc.
[0216] Chemical-regulated promoters can be used to modulate the expression of a gene in a plant through the application of an exogenous chemical regulator. The promoter may be a chemical-inducible promoter, where application of the chemical induces gene expression, or a chemical-repressible promoter, where application of the chemical represses gene expression. Chemical-inducible promoters include, but are not limited to, the maize In2-2 promoter, activated by benzene sulfonamide herbicide safeners (De Veylder et al., (1997) Plant Cell Physiol 38:568-77), the maize GST promoter (GST-II-27, WO93/01294), activated by hydrophobic electrophilic compounds used as pre-emergent herbicides, and the tobacco PR-1a promoter (Ono et al., (2004) Biosci Biotechnol Biochem 68:803-7) activated by salicylic acid. Other chemical-regulated promoters include steroid-responsive promoters (see, for example, the glucocorticoid-inducible promoter (Schena et al., (1991) Proc. Natl. Acad. Sci. USA 88:10421-5; McNellis et al., (1998) Plant J 14:247-257); tetracycline-inducible and tetracycline-repressible promoters (Gatz et al., (1991) Mol Gen Genet 227:229-37; U.S. Pat. Nos. 5,814,618 and 5,789,156).
[0217] Tissue-preferred promoters can be utilized to target enhanced expression within a particular plant tissue. Tissue-preferred promoters include, for example, Kawamata et al., (1997) Plant Cell Physiol 38:792-803; Hansen et al., (1997) Mol Gen Genet 254:337-43; Russell et al., (1997) Transgenic Res 6:157-68; Rinehart et al., (1996) Plant Physiol 112:1331-41; Van Camp et al., (1996) Plant Physiol 112:525-35; Canevascini et al., (1996) Plant Physiol 112:513-524; Lam, (1994) Results Probl Cell Differ 20:181-96; and Guevara-Garcia et al., (1993) Plant J 4:495-505. Leaf-preferred promoters include, for example, Yamamoto et al., (1997) Plant J 12:255-65; Kwon et al., (1994) Plant Physiol 105:357-67; Yamamoto et al., (1994) Plant Cell Physiol 35:773-8; Gotor et al., (1993) Plant J 3:509-18; Orozco et al., (1993) Plant Mol Biol 23:1129-38; Matsuoka et al., (1993) Proc. Natl. Acad. Sci. USA 90:9586-90; Simpson et al., (1958) EMBO J 4:2723-9; Timko et al., (1988) Nature 318:57-8. Root-preferred promoters include, for example, Hire et al., (1992) Plant Mol Biol 20:207-18 (soybean root-specific glutamine synthase gene); Miao et al., (1991) Plant Cell 3:11-22 (cytosolic glutamine synthase (GS)); Keller and Baumgartner, (1991) Plant Cell 3:1051-61 (root-specific control element in the GRP 1.8 gene of French bean); Sanger et al., (1990) Plant Mol Biol 14:433-43 (root-specific promoter of A. tumefaciens mannopine synthase (MAS)); Bogusz et al., (1990) Plant Cell 2:633-41 (root-specific promoters isolated from Parasponia andersonii and Trema tomentosa); Leach and Aoyagi, (1991) Plant Sci 79:69-76 (A. rhizogenes roIC and roID root-inducing genes); Teeri et al., (1989) EMBO J 8:343-50 (Agrobacterium wound-induced TR1' and TR2' genes); VfENOD-GRP3 gene promoter (Kuster et al., (1995) Plant Mol Biol 29:759-72); and rolB promoter (Capana et al., (1994) Plant Mol Biol 25:681-91; phaseolin gene (Murai et al., (1983) Science 23:476-82; Sengopta-Gopalen et al., (1988) Proc. Natl. Acad. Sci. USA 82:3320-4). See also, U.S. Pat. Nos. 5,837,876; 5,750,386; 5,633,363; 5,459,252; 5,401,836; 5,110,732 and 5,023,179.
[0218] Seed-preferred promoters include both seed-specific promoters active during seed development, as well as seed-germinating promoters active during seed germination. See, Thompson et al., (1989) BioEssays 10:108. Seed-preferred promoters include, but are not limited to, Cim1 (cytokinin-induced message); cZ19B1 (maize 19 kDa zein); and milps (myo-inositol-1-phosphate synthase); (WO00/11177; and U.S. Pat. No. 6,225,529). For dicots, seed-preferred promoters include, but are not limited to, bean .beta.-phaseolin, napin, .beta.-conglycinin, soybean lectin, cruciferin, and the like. For monocots, seed-preferred promoters include, but are not limited to, maize 15 kDa zein, 22 kDa zein, 27 kDa gamma zein, waxy, shrunken 1, shrunken 2, globulin 1, oleosin, and nuc1. See also, WO00/12733, where seed-preferred promoters from END1 and END2 genes are disclosed.
[0219] The term "inducible promoter" refers to promoters that selectively express a coding sequence or functional RNA in response to the presence of an endogenous or exogenous stimulus, for example by chemical compounds (chemical inducers) or in response to environmental, hormonal, chemical, and/or developmental signals. Inducible or regulated promoters include, for example, promoters induced or regulated by light, heat, stress, flooding or drought, salt stress, osmotic stress, phytohormones, wounding, or chemicals such as ethanol, abscisic acid (ABA), jasmonate, salicylic acid, or safeners.
[0220] An example of a stress-inducible is RD29A promoter (Kasuga et al. (1999) Nature Biotechnol. 17:287-91). One of ordinary skill in the art is familiar with protocols for simulating drought conditions and for evaluating drought tolerance of plants that have been subjected to simulated or naturally-occurring drought conditions. For example, one can simulate drought conditions by giving plants less water than normally required or no water over a period of time, and one can evaluate drought tolerance by looking for differences in physiological and/or physical condition, including (but not limited to) vigor, growth, size, or root length, or in particular, leaf color or leaf area size. Other techniques for evaluating drought tolerance include measuring chlorophyll fluorescence, photosynthetic rates and gas exchange rates. Also, one of ordinary skill in the art is familiar with protocols for simulating stress conditions such as osmotic stress, salt stress and temperature stress and for evaluating stress tolerance of plants that have been subjected to simulated or naturally-occurring stress conditions.
[0221] Another example of an inducible promoter useful in plant cells has been described in US patent application, US 2013-0312137A1, published on Nov. 21, 2013, incorporated by reference herein. US patent application US 2013-0312137A1 describes a ZmCAS1 promoter from a CBSU-Anther_Subtraction library (CAS1) gene encoding a mannitol dehydrogenase from maize, and functional fragments thereof. The ZmCAS1 promoter (also refered to as "CAS1 promoter", "mannitol dehydrogenase promoter", "mdh promoter") can be induced by a chemical or stress treatment. The chemical can be a safener such as, but not limited to, N-(aminocarbonyl)-2-chlorobenzenesulfonamide (2-CBSU). The stress treatment can be a heat treatment such as, but not limited to, a heat shock treatment (see also U.S. provisional patent application, 62/120,421, filed on Feb. 25, 2015, incorporated by reference herein.
[0222] New promoters of various types useful in plant cells are constantly being discovered; numerous examples may be found in the compilation by Okamuro and Goldberg, (1989) In The Biochemistry of Plants, Vol. 115, Stumpf and Conn, eds (New York, N.Y.: Academic Press), pp. 1-82.
[0223] "Translation leader sequence" refers to a polynucleotide sequence located between the promoter sequence of a gene and the coding sequence. The translation leader sequence is present in the mRNA upstream of the translation start sequence. The translation leader sequence may affect processing of the primary transcript to mRNA, mRNA stability or translation efficiency. Examples of translation leader sequences have been described (e.g., Turner and Foster, (1995) Mol Biotechnol 3:225-236).
[0224] "3' non-coding sequences", "transcription terminator" or "termination sequences" refer to DNA sequences located downstream of a coding sequence and include polyadenylation recognition sequences and other sequences encoding regulatory signals capable of affecting mRNA processing or gene expression. The polyadenylation signal is usually characterized by affecting the addition of polyadenylic acid tracts to the 3' end of the mRNA precursor. The use of different 3' non-coding sequences is exemplified by Ingelbrecht et al., (1989) Plant Cell 1:671-680.
[0225] "RNA transcript" refers to the product resulting from RNA polymerase-catalyzed transcription of a DNA sequence. When the RNA transcript is a perfect complimentary copy of the DNA sequence, it is referred to as the primary transcript or pre-mRNA. A RNA transcript is referred to as the mature RNA or mRNA when it is a RNA sequence derived from post-transcriptional processing of the primary transcript pre mRNAt. "Messenger RNA" or "mRNA" refers to the RNA that is without introns and that can be translated into protein by the cell. "cDNA" refers to a DNA that is complementary to, and synthesized from, a mRNA template using the enzyme reverse transcriptase. The cDNA can be single-stranded or converted into double-stranded form using the Klenow fragment of DNA polymerase I. "Sense" RNA refers to RNA transcript that includes the mRNA and can be translated into protein within a cell or in vitro. "Antisense RNA" refers to an RNA transcript that is complementary to all or part of a target primary transcript or mRNA, and that blocks the expression of a target gene (see, e.g., U.S. Pat. No. 5,107,065). The complementarity of an antisense RNA may be with any part of the specific gene transcript, i.e., at the 5' non-coding sequence, 3' non-coding sequence, introns, or the coding sequence. "Functional RNA" refers to antisense RNA, ribozyme RNA, or other RNA that may not be translated but yet has an effect on cellular processes. The terms "complement" and "reverse complement" are used interchangeably herein with respect to mRNA transcripts, and are meant to define the antisense RNA of the message.
[0226] The term "operably linked" refers to the association of nucleic acid sequences on a single nucleic acid fragment so that the function of one is regulated by the other. For example, a promoter is operably linked with a coding sequence when it is capable of regulating the expression of that coding sequence (i.e., the coding sequence is under the transcriptional control of the promoter). Coding sequences can be operably linked to regulatory sequences in a sense or antisense orientation. In another example, the complementary RNA regions can be operably linked, either directly or indirectly, 5' to the target mRNA, or 3' to the target mRNA, or within the target mRNA, or a first complementary region is 5' and its complement is 3' to the target m RNA.
[0227] Standard recombinant DNA and molecular cloning techniques used herein are well known in the art and are described more fully in Sambrook et al., Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory: Cold Spring Harbor, N.Y. (1989). Transformation methods are well known to those skilled in the art and are described infra.
[0228] "PCR" or "polymerase chain reaction" is a technique for the synthesis of specific DNA segments and consists of a series of repetitive denaturation, annealing, and extension cycles. Typically, a double-stranded DNA is heat denatured, and two primers complementary to the 3' boundaries of the target segment are annealed to the DNA at low temperature, and then extended at an intermediate temperature. One set of these three consecutive steps is referred to as a "cycle".
[0229] The term "recombinant" refers to an artificial combination of two otherwise separated segments of sequence, e.g., by chemical synthesis, or manipulation of isolated segments of nucleic acids by genetic engineering techniques.
[0230] The terms "plasmid", "vector" and "cassette" refer to an extra chromosomal element often carrying genes that are not part of the central metabolism of the cell, and usually in the form of double-stranded DNA. Such elements may be autonomously replicating sequences, genome integrating sequences, phage, or nucleotide sequences, in linear or circular form, of a single- or double-stranded DNA or RNA, derived from any source, in which a number of nucleotide sequences have been joined or recombined into a unique construction which is capable of introducing a polynucleotide of interest into a cell. "Transformation cassette" refers to a specific vector containing a gene and having elements in addition to the gene that facilitates transformation of a particular host cell. "Expression cassette" refers to a specific vector containing a gene and having elements in addition to the gene that allow for expression of that gene in a host.
[0231] The terms "recombinant DNA molecule", "recombinant construct", "expression construct", "construct", "construct", and "recombinant DNA construct" are used interchangeably herein. A recombinant construct comprises an artificial combination of nucleic acid fragments, e.g., regulatory and coding sequences that are not all found together in nature. For example, a construct may comprise regulatory sequences and coding sequences that are derived from different sources, or regulatory sequences and coding sequences derived from the same source, but arranged in a manner different than that found in nature. Such a construct may be used by itself or may be used in conjunction with a vector. If a vector is used, then the choice of vector is dependent upon the method that will be used to transform host cells as is well known to those skilled in the art. For example, a plasmid vector can be used. The skilled artisan is well aware of the genetic elements that must be present on the vector in order to successfully transform, select and propagate host cells. The skilled artisan will also recognize that different independent transformation events may result in different levels and patterns of expression (Jones et al., (1985) EMBO J 4:2411-2418; De Almeida et al., (1989) Mol Gen Genetics 218:78-86), and thus that multiple events are typically screened in order to obtain lines displaying the desired expression level and pattern. Such screening may be accomplished standard molecular biological, biochemical, and other assays including Southern analysis of DNA, Northern analysis of mRNA expression, PCR, real time quantitative PCR (qPCR), reverse transcription PCR (RT-PCR), immunoblotting analysis of protein expression, enzyme or activity assays, and/or phenotypic analysis.
[0232] The term "expression", as used herein, refers to the production of a functional end-product (e.g., an mRNA, guide RNA, or a protein) in either precursor or mature form.
[0233] The term "providing" includes providing a nucleic acid (e.g., expression construct) or protein into a cell. Providing includes reference to the incorporation of a nucleic acid into a eukaryotic or prokaryotic cell where the nucleic acid may be incorporated into the genome of the cell, and includes reference to the transient provision of a nucleic acid or protein to the cell. Introduced includes reference to stable or transient transformation methods, as well as sexually crossing. Thus, "providing" in the context of inserting a nucleic acid fragment (e.g., a recombinant DNA construct/expression construct) into a cell, means "transfection" or "transformation" or "transduction" and includes reference to the incorporation of a nucleic acid fragment into a eukaryotic or prokaryotic cell where the nucleic acid fragment may be incorporated into the genome of the cell (e.g., chromosome, plasmid, plastid, or mitochondrial DNA), converted into an autonomous replicon, or transiently expressed (e.g., transfected mRNA).
[0234] "Mature" protein refers to a post-translationally processed polypeptide (i.e., one from which any pre- or propeptides present in the primary translation product have been removed). "Precursor" protein refers to the primary product of translation of mRNA (i.e., with pre- and propeptides still present). Pre- and propeptides may be but are not limited to intracellular localization signals.
[0235] "Stable transformation" refers to the transfer of a nucleic acid fragment into a genome of a host organism, including both nuclear and organellar genomes, resulting in genetically stable inheritance. In contrast, "transient transformation" refers to the transfer of a nucleic acid fragment into the nucleus, or other DNA-containing organelle, of a host organism resulting in gene expression without integration or stable inheritance. Host organisms containing the transformed nucleic acid fragments are referred to as "transgenic" organisms.
[0236] The commercial development of genetically improved germplasm has also advanced to the stage of introducing multiple traits into crop plants, often referred to as a gene stacking approach. In this approach, multiple genes conferring different characteristics of interest can be introduced into a plant. Gene stacking can be accomplished by many means including but not limited to co-transformation, retransformation, and crossing lines with different genes of interest.
[0237] The term "plant" refers to whole plants, plant organs, plant tissues, seeds, plant cells, seeds and progeny of the same. Plant cells include, without limitation, cells from seeds, suspension cultures, embryos, meristematic regions, callus tissue, leaves, roots, shoots, gametophytes, sporophytes, pollen and microspores. Plant parts include differentiated and undifferentiated tissues including, but not limited to roots, stems, shoots, leaves, pollens, seeds, tumor tissue and various forms of cells and culture (e.g., single cells, protoplasts, embryos, and callus tissue). The plant tissue may be in plant or in a plant organ, tissue or cell culture. The term "plant organ" refers to plant tissue or a group of tissues that constitute a morphologically and functionally distinct part of a plant. The term "genome" refers to the entire complement of genetic material (genes and non-coding sequences) that is present in each cell of an organism, or virus or organelle; and/or a complete set of chromosomes inherited as a (haploid) unit from one parent. "Progeny" comprises any subsequent generation of a plant.
[0238] A transgenic plant includes, for example, a plant which comprises within its genome a heterologous polynucleotide introduced by a transformation step. The heterologous polynucleotide can be stably integrated within the genome such that the polynucleotide is passed on to successive generations. The heterologous polynucleotide may be integrated into the genome alone or as part of a recombinant DNA construct. A transgenic plant can also comprise more than one heterologous polynucleotide within its genome. Each heterologous polynucleotide may confer a different trait to the transgenic plant. A heterologous polynucleotide can include a sequence that originates from a foreign species, or, if from the same species, can be substantially modified from its native form. Transgenic can include any cell, cell line, callus, tissue, plant part or plant, the genotype of which has been altered by the presence of heterologous nucleic acid including those transgenics initially so altered as well as those created by sexual crosses or asexual propagation from the initial transgenic. The alterations of the genome (chromosomal or extra-chromosomal) by conventional plant breeding methods, by the genome editing procedure described herein that does not result in an insertion of a foreign polynucleotide, or by naturally occurring events such as random cross-fertilization, non-recombinant viral infection, non-recombinant bacterial transformation, non-recombinant transposition, or spontaneous mutation are not intended to be regarded as transgenic.
[0239] In certain embodiments of the disclosure, a fertile plant is a plant that produces viable male and female gametes and is self-fertile. Such a self-fertile plant can produce a progeny plant without the contribution from any other plant of a gamete and the genetic material contained therein. Other embodiments of the disclosure can involve the use of a plant that is not self-fertile because the plant does not produce male gametes, or female gametes, or both, that are viable or otherwise capable of fertilization. As used herein, a "male sterile plant" is a plant that does not produce male gametes that are viable or otherwise capable of fertilization. As used herein, a "female sterile plant" is a plant that does not produce female gametes that are viable or otherwise capable of fertilization. It is recognized that male-sterile and female-sterile plants can be female-fertile and male-fertile, respectively. It is further recognized that a male fertile (but female sterile) plant can produce viable progeny when crossed with a female fertile plant and that a female fertile (but male sterile) plant can produce viable progeny when crossed with a male fertile plant.
[0240] The term "non-conventional yeast" herein refers to any yeast that is not a Saccharomyces (e.g., S. cerevisiae) or Schizosaccharomyces yeast species. Non-conventional yeast are described in Non-Conventional Yeasts in Genetics, Biochemistry and Biotechnology: Practical Protocols (K. Wolf, K. D. Breunig, G. Barth, Eds., Springer-Verlag, Berlin, Germany, 2003), which is incorporated herein by reference. Non-conventional yeast in certain embodiments may additionally (or alternatively) be yeast that favor non-homologous end-joining (NHEJ) DNA repair processes over repair processes mediated by homologous recombination (HR). Definition of a non-conventional yeast along these lines--preference of NHEJ over HR--is further disclosed by Chen et al. (PLoS ONE 8:e57952), which is incorporated herein by reference. Preferred non-conventional yeast herein are those of the genus Yarrowia (e.g., Yarrowia lipolytica). The term "yeast" herein refers to fungal species that predominantly exist in unicellular form. Yeast can alternative be referred to as "yeast cells" herein. (see also U.S. provisional application 62/036,652, filed on Aug. 13, 2014, which is incorporated by reference herein.
[0241] A "centimorgan" (cM) or "map unit" is the distance between two linked genes, markers, target sites, loci, or any pair thereof, wherein 1% of the products of meiosis are recombinant. Thus, a centimorgan is equivalent to a distance equal to a 1 average recombination frequency between the two linked genes, markers, target sites, loci, or any pair thereof.
[0242] The present disclosure finds use in the breeding of plants comprising one or more transgenic traits. Most commonly, transgenic traits are randomly inserted throughout the plant genome as a consequence of transformation systems based on Agrobacterium, biolistics, or other commonly used procedures. More recently, gene targeting protocols have been developed that enable directed transgene insertion. One important technology, site-specific integration (SSI) enables the targeting of a transgene to the same chromosomal location as a previously inserted transgene. Custom-designed meganucleases and custom-designed zinc finger meganucleases allow researchers to design nucleases to target specific chromosomal locations, and these reagents allow the targeting of transgenes at the chromosomal site cleaved by these nucleases.
[0243] The currently used systems for precision genetic engineering of eukaryotic genomes, e.g. plant genomes, rely upon homing endonucleases, meganucleases, zinc finger nucleases, and transcription activator-like effector nucleases (TALENs), which require de novo protein engineering for every new target locus. The highly specific, RNA-directed DNA nuclease, guide RNA/Cas9 endonuclease system described herein, is more easily customizable and therefore more useful when modification of many different target sequences is the goal. This disclosure takes further advantage of the two component nature of the guide RNA/Cas system, with its constant protein component, the Cas endonuclease, and its variable and easily reprogrammable targeting component, the guide RNA or the crRNA.
[0244] The guide RNA/Cas system described herein is especially useful for genome engineering, especially plant genome engineering, in circumstances where nuclease off-target cutting can be toxic to the targeted cells. In one embodiment of the guide RNA/Cas system described herein, the constant component, in the form of an expression-optimized Cas9 gene, is stably integrated into the target genome, e.g. plant genome. Expression of the Cas9 gene is under control of a promoter, e.g. plant promoter, which can be a constitutive promoter, tissue-specific promoter or inducible promoter, e.g. temperature-inducible, stress-inducible, developmental stage inducible, or chemically inducible promoter. In the absence of the variable component, i.e. the guide RNA or crRNA, the Cas9 protein is not able to cut DNA and therefore its presence in the plant cell should have little or no consequence. Hence a key advantage of the guide RNA/Cas system described herein is the ability to create and maintain a cell line or transgenic organism capable of efficient expression of the Cas9 protein with little or no consequence to cell viability. In order to induce cutting at desired genomic sites to achieve targeted genetic modifications, guide RNAs or crRNAs can be introduced by a variety of methods into cells containing the stably-integrated and expressed cas9 gene. For example, guide RNAs or crRNAs can be chemically or enzymatically synthesized, and introduced into the Cas9 expressing cells via direct delivery methods such a particle bombardment or electroporation.
[0245] Alternatively, genes capable of efficiently expressing guide RNAs or crRNAs in the target cells can be synthesized chemically, enzymatically or in a biological system, and these genes can be introduced into the Cas9 expressing cells via direct delivery methods such a particle bombardment, electroporation or biological delivery methods such as Agrobacterium mediated DNA delivery.
[0246] A guide RNA/Cas system mediating gene targeting can be used in methods for directing transgene insertion and/or for producing complex transgenic trait loci comprising multiple transgenes in a fashion similar as disclosed in WO2013/0198888 (published Aug. 1, 2013) where instead of using a double strand break inducing agent to introduce a gene of interest, a guide RNA/Cas system as disclosed herein is used. In one embodiment, a complex transgenic trait locus is a genomic locus that has multiple transgenes genetically linked to each other. By inserting independent transgenes within 0.1, 0.2, 0.3, 0.4, 0.5, 1.0, 2, or even 5 centimorgans (cM) from each other, the transgenes can be bred as a single genetic locus (see, for example, U.S. patent application Ser. No. 13/427,138) or PCT application PCT/US2012/030061. After selecting a plant comprising a transgene, plants containing (at least) one transgenes can be crossed to form an F1 that contains both transgenes. In progeny from these F1 (F2 or BC1) 1/500 progeny would have the two different transgenes recombined onto the same chromosome. The complex locus can then be bred as single genetic locus with both transgene traits. This process can be repeated to stack as many traits as desired.
[0247] Chromosomal intervals that correlate with a phenotype or trait of interest can be identified. A variety of methods well known in the art are available for identifying chromosomal intervals. The boundaries of such chromosomal intervals are drawn to encompass markers that will be linked to the gene controlling the trait of interest. In other words, the chromosomal interval is drawn such that any marker that lies within that interval (including the terminal markers that define the boundaries of the interval) can be used as a marker for northern leaf blight resistance. In one embodiment, the chromosomal interval comprises at least one QTL, and furthermore, may indeed comprise more than one QTL. Close proximity of multiple QTLs in the same interval may obfuscate the correlation of a particular marker with a particular QTL, as one marker may demonstrate linkage to more than one QTL. Conversely, e.g., if two markers in close proximity show co-segregation with the desired phenotypic trait, it is sometimes unclear if each of those markers identifies the same QTL or two different QTL. The term "quantitative trait locus" or "QTL" refers to a region of DNA that is associated with the differential expression of a quantitative phenotypic trait in at least one genetic background, e.g., in at least one breeding population. The region of the QTL encompasses or is closely linked to the gene or genes that affect the trait in question. An "allele of a QTL" can comprise multiple genes or other genetic factors within a contiguous genomic region or linkage group, such as a haplotype. An allele of a QTL can denote a haplotype within a specified window wherein said window is a contiguous genomic region that can be defined, and tracked, with a set of one or more polymorphic markers. A haplotype can be defined by the unique fingerprint of alleles at each marker within the specified window.
[0248] A variety of methods are available to identify those cells having an altered genome at or near a target site without using a screenable marker phenotype. Such methods can be viewed as directly analyzing a target sequence to detect any change in the target sequence, including but not limited to PCR methods, sequencing methods, nuclease digestion, Southern blots, and any combination thereof.
[0249] Proteins may be altered in various ways including amino acid substitutions, deletions, truncations, and insertions. Methods for such manipulations are generally known. For example, amino acid sequence variants of the protein(s) can be prepared by mutations in the DNA. Methods for mutagenesis and nucleotide sequence alterations include, for example, Kunkel, (1985) Proc. Natl. Acad. Sci. USA 82:488-92; Kunkel et al., (1987) Meth Enzymol 154:367-82; U.S. Pat. No. 4,873,192; Walker and Gaastra, eds. (1983) Techniques in Molecular Biology (MacMillan Publishing Company, New York) and the references cited therein. Guidance regarding amino acid substitutions not likely to affect biological activity of the protein is found, for example, in the model of Dayhoff et al., (1978) Atlas of Protein Sequence and Structure (Natl Biomed Res Found, Washington, D.C.). Conservative substitutions, such as exchanging one amino acid with another having similar properties, may be preferable. Conservative deletions, insertions, and amino acid substitutions are not expected to produce radical changes in the characteristics of the protein, and the effect of any substitution, deletion, insertion, or combination thereof can be evaluated by routine screening assays. Assays for double-strand-break-inducing activity are known and generally measure the overall activity and specificity of the agent on DNA substrates containing target sites.
[0250] A variety of methods are known for the introduction of nucleotide sequences and polypeptides into an organism, including, for example, transformation, sexual crossing, and the introduction of the polypeptide, DNA, or mRNA into the cell.
[0251] Methods for contacting, providing, and/or introducing a composition into various organisms are known and include but are not limited to, stable transformation methods, transient transformation methods, virus-mediated methods, and sexual breeding. Stable transformation indicates that the introduced polynucleotide integrates into the genome of the organism and is capable of being inherited by progeny thereof. Transient transformation indicates that the introduced composition is only temporarily expressed or present in the organism.
[0252] Protocols for introducing polynucleotides and polypeptides into plants may vary depending on the type of plant or plant cell targeted for transformation, such as monocot or dicot. Suitable methods of introducing polynucleotides and polypeptides into plant cells and subsequent insertion into the plant genome include microinjection (Crossway et al., (1986) Biotechniques 4:320-34 and U.S. Pat. No. 6,300,543), meristem transformation (U.S. Pat. No. 5,736,369), electroporation (Riggs et al., (1986) Proc. Natl. Acad. Sci. USA 83:5602-6, Agrobacterium-mediated transformation (U.S. Pat. Nos. 5,563,055 and 5,981,840), direct gene transfer (Paszkowski et al., (1984) EMBO J 3:2717-22), and ballistic particle acceleration (U.S. Pat. Nos. 4,945,050; 5,879,918; 5,886,244; 5,932,782; Tomes et al., (1995) "Direct DNA Transfer into Intact Plant Cells via Microprojectile Bombardment" in Plant Cell, Tissue, and Organ Culture: Fundamental Methods, ed. Gamborg & Phillips (Springer-Verlag, Berlin); McCabe et al., (1988) Biotechnology 6:923-6; Weissinger et al., (1988) Ann Rev Genet 22:421-77; Sanford et al., (1987) Particulate Science and Technology 5:27-37 (onion); Christou et al., (1988) Plant Physiol 87:671-4 (soybean); Finer and McMullen, (1991) In Vitro Cell Dev Biol 27P:175-82 (soybean); Singh et al., (1998) Theor Appl Genet 96:319-24 (soybean); Datta et al., (1990) Biotechnology 8:736-40 (rice); Klein et al., (1988) Proc. Natl. Acad. Sci. USA 85:4305-9 (maize); Klein et al., (1988) Biotechnology 6:559-63 (maize); U.S. Pat. Nos. 5,240,855; 5,322,783 and 5,324,646; Klein et al., (1988) Plant Physiol 91:440-4 (maize); Fromm et al., (1990) Biotechnology 8:833-9 (maize); Hooykaas-Van Slogteren et al., (1984) Nature 311:763-4; U.S. Pat. No. 5,736,369 (cereals); Bytebier et al., (1987) Proc. Natl. Acad. Sci. USA 84:5345-9 (Liliaceae); De Wet et al., (1985) in The Experimental Manipulation of Ovule Tissues, ed. Chapman et al., (Longman, New York), pp. 197-209 (pollen); Kaeppler et al., (1990) Plant Cell Rep 9:415-8) and Kaeppler et al., (1992) Theor Appl Genet 84:560-6 (whisker-mediated transformation); D'Halluin et al., (1992) Plant Cell 4:1495-505 (electroporation); Li et al., (1993) Plant Cell Rep 12:250-5; Christou and Ford (1995) Annals Botany 75:407-13 (rice) and Osjoda et al., (1996) Nat Biotechnol 14:745-50 (maize via Agrobacterium tumefaciens).
[0253] Alternatively, polynucleotides may be introduced into plants by contacting plants with a virus or viral nucleic acids. Generally, such methods involve incorporating a polynucleotide within a viral DNA or RNA molecule. In some examples a polypeptide of interest may be initially synthesized as part of a viral polyprotein, which is later processed by proteolysis in vivo or in vitro to produce the desired recombinant protein. Methods for introducing polynucleotides into plants and expressing a protein encoded therein, involving viral DNA or RNA molecules, are known, see, for example, U.S. Pat. Nos. 5,889,191, 5,889,190, 5,866,785, 5,589,367 and 5,316,931. Transient transformation methods include, but are not limited to, the introduction of polypeptides, such as a double-strand break inducing agent, directly into the organism, the introduction of polynucleotides such as DNA and/or RNA polynucleotides, and the introduction of the RNA transcript, such as an mRNA encoding a double-strand break inducing agent, into the organism. Such methods include, for example, microinjection or particle bombardment. See, for example Crossway et al., (1986) Mol Gen Genet 202:179-85; Nomura et al., (1986) Plant Sci 44:53-8; Hepler et al., (1994) Proc. Natl. Acad. Sci. USA 91:2176-80; and, Hush et al., (1994) J Cell Sci 107:775-84.
[0254] The term "dicot" refers to the subclass of angiosperm plants also knows as "dicotyledoneae" and includes reference to whole plants, plant organs (e.g., leaves, stems, roots, etc.), seeds, plant cells, and progeny of the same. Plant cell, as used herein includes, without limitation, seeds, suspension cultures, embryos, meristematic regions, callus tissue, leaves, roots, shoots, gametophytes, sporophytes, pollen, and microspores.
[0255] The term "crossed" or "cross" or "crossing" in the context of this disclosure means the fusion of gametes via pollination to produce progeny (i.e., cells, seeds, or plants). The term encompasses both sexual crosses (the pollination of one plant by another) and selfing (self-pollination, i.e., when the pollen and ovule (or microspores and megaspores) are from the same plant or genetically identical plants).
[0256] The term "introgression" refers to the transmission of a desired allele of a genetic locus from one genetic background to another. For example, introgression of a desired allele at a specified locus can be transmitted to at least one progeny plant via a sexual cross between two parent plants, where at least one of the parent plants has the desired allele within its genome. Alternatively, for example, transmission of an allele can occur by recombination between two donor genomes, e.g., in a fused protoplast, where at least one of the donor protoplasts has the desired allele in its genome. The desired allele can be, e.g., a transgene, a modified (mutated or edited) native allele, or a selected allele of a marker or QTL.
[0257] Standard DNA isolation, purification, molecular cloning, vector construction, and verification/characterization methods are well established, see, for example Sambrook et al., (1989) Molecular Cloning: A Laboratory Manual, (Cold Spring Harbor Laboratory Press, NY). Vectors and constructs include circular plasmids, and linear polynucleotides, comprising a polynucleotide of interest and optionally other components including linkers, adapters, regulatory or analysis. In some examples a recognition site and/or target site can be contained within an intron, coding sequence, 5' UTRs, 3' UTRs, and/or regulatory regions.
[0258] The present disclosure further provides expression constructs for expressing in a plant, plant cell, or plant part a guide RNA/Cas system that is capable of binding to and creating a double strand break in a target site. In one embodiment, the expression constructs of the disclosure comprise a promoter operably linked to a nucleotide sequence encoding a Cas gene and a promoter operably linked to a guide RNA of the present disclosure. The promoter is capable of driving expression of an operably linked nucleotide sequence in a plant cell.
[0259] A phenotypic marker is a screenable or selectable marker that includes visual markers and selectable markers whether it is a positive or negative selectable marker. Any phenotypic marker can be used. Specifically, a selectable or screenable marker comprises a DNA segment that allows one to identify, or select for or against a molecule or a cell that contains it, often under particular conditions. These markers can encode an activity, such as, but not limited to, production of RNA, peptide, or protein, or can provide a binding site for RNA, peptides, proteins, inorganic and organic compounds or compositions and the like.
[0260] Examples of selectable markers include, but are not limited to, DNA segments that comprise restriction enzyme sites; DNA segments that encode products which provide resistance against otherwise toxic compounds including antibiotics, such as, spectinomycin, ampicillin, kanamycin, tetracycline, Basta, neomycin phosphotransferase II (NEO) and hygromycin phosphotransferase (HPT)); DNA segments that encode products which are otherwise lacking in the recipient cell (e.g., tRNA genes, auxotrophic markers); DNA segments that encode products which can be readily identified (e.g., phenotypic markers such as .beta.-galactosidase, GUS; fluorescent proteins such as green fluorescent protein (GFP), cyan (CFP), yellow (YFP), red (RFP), and cell surface proteins); the generation of new primer sites for PCR (e.g., the juxtaposition of two DNA sequence not previously juxtaposed), the inclusion of DNA sequences not acted upon or acted upon by a restriction endonuclease or other DNA modifying enzyme, chemical, etc.; and, the inclusion of a DNA sequences required for a specific modification (e.g., methylation) that allows its identification.
[0261] Additional selectable markers include genes that confer resistance to herbicidal compounds, such as glufosinate ammonium, bromoxynil, imidazolinones, and 2,4-dichlorophenoxyacetate (2,4-D). See for example, Yarranton, (1992) Curr Opin Biotech 3:506-11; Christopherson et al., (1992) Proc. Natl. Acad. Sci. USA 89:6314-8; Yao et al., (1992) Cell 71:63-72; Reznikoff, (1992) Mol Microbiol 6:2419-22; Hu et al., (1987) Cell 48:555-66; Brown et al., (1987) Cell 49:603-12; Figge et al., (1988) Cell 52:713-22; Deuschle et al., (1989) Proc. Natl. Acad. Sci. USA 86:5400-4; Fuerst et al., (1989) Proc. Natl. Acad. Sci. USA 86:2549-53; Deuschle et al., (1990) Science 248:480-3; Gossen, (1993) Ph.D. Thesis, University of Heidelberg; Reines et al., (1993) Proc. Natl. Acad. Sci. USA 90:1917-21; Labow et al., (1990) Mol Cell Biol 10:3343-56; Zambretti et al., (1992) Proc. Natl. Acad. Sci. USA 89:3952-6; Baim et al., (1991) Proc. Natl. Acad. Sci. USA 88:5072-6; Wyborski et al., (1991) Nucleic Acids Res 19:4647-53; Hillen and Wissman, (1989) Topics Mol Struc Biol 10:143-62; Degenkolb et al., (1991) Antimicrob Agents Chemother 35:1591-5; Kleinschnidt et al., (1988) Biochemistry 27:1094-104; Bonin, (1993) Ph.D. Thesis, University of Heidelberg; Gossen et al., (1992) Proc. Natl. Acad. Sci. USA 89:5547-51; Oliva et al., (1992) Antimicrob Agents Chemother 36:913-9; Hlavka et al., (1985) Handbook of Experimental Pharmacology, Vol. 78 (Springer-Verlag, Berlin); Gill et al., (1988) Nature 334:721-4.
[0262] The cells having the introduced sequence may be grown or regenerated into plants using conventional conditions, see for example, McCormick et al., (1986) Plant Cell Rep 5:81-4. These plants may then be grown, and either pollinated with the same transformed strain or with a different transformed or untransformed strain, and the resulting progeny having the desired characteristic and/or comprising the introduced polynucleotide or polypeptide identified. Two or more generations may be grown to ensure that the polynucleotide is stably maintained and inherited, and seeds harvested.
[0263] Any plant can be used, including monocot and dicot plants. Examples of monocot plants that can be used include, but are not limited to, corn (Zea mays), rice (Oryza sativa), rye (Secale cereale), sorghum (Sorghum bicolor, Sorghum vulgare), millet (e.g., pearl millet (Pennisetum glaucum), proso millet (Panicum miliaceum), foxtail millet (Setaria italica), finger millet (Eleusine coracana)), wheat (Triticum aestivum), sugarcane (Saccharum spp.), oats (Avena), barley (Hordeum), switchgrass (Panicum virgatum), pineapple (Ananas comosus), banana (Musa spp.), palm, ornamentals, turfgrasses, and other grasses. Examples of dicot plants that can be used include, but are not limited to, soybean (Glycine max), canola (Brassica napus and B. campestris), alfalfa (Medicago sativa), tobacco (Nicotiana tabacum), Arabidopsis (Arabidopsis thaliana), sunflower (Helianthus annuus), cotton (Gossypium arboreum), and peanut (Arachis hypogaea), tomato (Solanum lycopersicum), potato (Solanum tuberosum) etc.
[0264] The transgenes, recombinant DNA molecules, DNA sequences of interest, and polynucleotides of interest can comprise one or more genes of interest. Such genes of interest can encode, for example, a protein that provides agronomic advantage to the plant.
[0265] Also provided are kits for performing any of the above methods described herein. The kits typically contain polynucleotides encoding one or more Cas endonuclease, or Cas endonuclease protein wherein the Cas endonuclease protein is provided as a purified protein, a cell lysate comprising said Cas endonuclease, a dilution of a cell lysate comprising said Cas endonuclease, an in-vitro translation mixture or an dilution of an in-vitro translation mixture, and/or single or dual guide polynucleotides, and/or template polynucleotides for gene editing and/or donor polynucleotides for inserting polynucleotides of interest into a genome of interest, as described herein. The kit can further contain instructions for administering all these components into the cells. The kits can also contain cells, buffers for transformation of cells, culture media for cells, and/or buffers for performing assays. The kits can further contain one or more inhibitors of proteins involved in NHEJ, or components which promote or increase homology-dependent repair (HDR) and instructions for introducing the Cas endonucleases and inhibitors into the cells such that Cas endonuclease-mediated gene disruption and/or targeted integration is enhanced. Optionally, cells containing the target site(s) of the Cas endonuclease may also be included in the kits described herein.
[0266] Inhibitors of non-homologous end joining (NHEJ) are known in the art and include molecules, such as but not limited to small molecules that inhibits (decrease) the binding or activity of a DNA-dependent-protein kinase catalytic subunit (DNA-PKcs), a Poly(ADP-ribose) polymerase 1/2 (PARPI/2), a PARPI, Ku70/80, a DNA-PKcs, a XRCC4/XLF, a Ligase IV, a Ligase III, a XRCCI, an Artemis Polynucleotide Kinase (PNK), SCR7, and any one combinations thereof (Sfeir et al. 2015, TIBS Vol 40 (11), pp 701-713; Srivastava, M. et al. An inhibitor of nonhomologous end-joining abrogates double-strand break repair and impedes cancer progression. Cell 151, 1474-1487 (2012); US patent application US2014/0242702, published on Aug. 28, 2014, herein incorporated in its entirety by reference). Other molecules that decrease the activity of the non-homologous end joining (NHEJ) DNA repair complex are known in the art and include RNAi-molecules, antisense nucleic acid molecules, ribozymes, compounds inhibiting the formation of a functional DNA Ligase IV (LIG4) complex and compounds enhancing proteolytic degradation of a functional DNA Ligase IV complex (US patent application 2014/0304847, published on Oct. 9, 2014, herein incorporated in its entirety by reference.
[0267] Activators of HDR are known in the art and include molecules, such as but not limited to RS1, RAD51 and RAD51B (Song et al. 2016 "RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency" Nature communications 7, Article number:10548; Takaku, M. et al 2009. Recombination activator function of the novel RAD51- and RAD51B-binding protein, human EVL. J. Biol. Chem. 284, 14326-14336 (2009).
[0268] In certain embodiments, the kits comprise at least one construct with a target gene and a Cas endonuclease described herein capable of cleaving within or in close proximity to the target gene. Such kits are useful for optimization of cleavage conditions in a variety of varying host cell types. In one aspect, the kit is a kit useful for increasing gene disruption, gene editing and/or targeted integration following Cas endonuclease mediated cleavage of a cell's genome.
[0269] In one embodiment, the kit includes a Cas endonuclease described herein capable of cleaving within a known target locus within a genome, and may additionally comprise a template DNA for gene editing and/or a donor nucleic acid for introducing a polynucleotide of interest into the cell's genome. Such kits are useful for optimization of conditions for template recognition, donor integration or for the construction of specifically modified cells, cell lines, and transgenic plants and animals containing gene disruptions, gene edits or targeted insertions. These and other aspects will be readily apparent to the skilled artisan in light of disclosure as a whole.
[0270] Also provided are kits containing any one or more of the elements disclosed in compositions described herein. In one aspect, the kits comprise a single guide polynucleotide comprising a crRNA, as described herein linked to a tracrRNA, wherein the crRNA comprises a variable targeting domain operably linked to a tracr mate sequence and/or one or more insertion sites for inserting or exchanging the variable targeting domain upstream of the tracr mate sequence, wherein when expressed, the single guide polynucleotide directs sequence-specific binding of a guide polynucleotide/Cas endonuclease complex to a target sequence in a eukaryotic cell. In another aspect, the kits comprise a dual guide polynucleotide comprising a crRNA molecule and a tracrRNA molecule, as described herein, wherein the crRNA molecule comprises a variable targeting domain operably linked to a tracr mate sequence and/or one or more insertion sites for inserting or exchanging the variable targeting domain upstream of the tracr mate sequence, wherein when expressed, the dual guide polynucleotide directs sequence-specific binding of a guide polynucleotide/Cas endonuclease complex to a target sequence in a eukaryotic cell.
[0271] The kits can contain one or more vectors encoding the guide polynucleotides, Cas endonucleases and/or template DNAs and/or donor DNAs described herein, and or the kits can contain the elements (guide polynucleotides, DNA templates, DNA donors and/or Cas endonucleases in purified or non-purified forms).
[0272] In one aspect, the kit comprises a Cas endonuclease as described herein, and/or a polynucleotide modification template and/or a donor DNA for inserting a polynucleotide of interest as described herein.
[0273] Components may be provide individually or in combinations, and may be provided in any suitable container, such as a vial, a bottle, or a tube. For example, a kit may provide one or more reaction or storage buffers. Reagents may be provided in a form that is usable in a particular assay, or in a form that requires addition of one or more other components before use (e.g. in concentrate or lyophilized form). A buffer can be any buffer, including but not limited to a sodium carbonate buffer, a sodium bicarbonate buffer, a borate buffer, a Tris buffer, a MOPS buffer, a HEPES buffer, and combinations thereof. In some embodiments, the buffer is alkaline. In some embodiments, the buffer has a pH from about 7 to about 10. In some aspects, the kit includes instructions in one or more languages, for example in more than one language.
[0274] The meaning of abbreviations is as follows: "sec" means second(s), "min" means minute(s), "h" means hour(s), "d" means day(s), ".mu.L" means microliter(s), "mL" means milliliter(s), "L" means liter(s), ".mu.M" means micromolar, "mM" means millimolar, "M" means molar, "mmol" means millimole(s), ".mu.mole" mean micromole(s), "g" means gram(s), ".mu.g" means microgram(s), "ng" means nanogram(s), "U" means unit(s), "bp" means base pair(s) and "kb" means kilobase(s).
[0275] Non-limiting examples of compositions and methods disclosed herein are as follows: [0276] 1. A method for producing a plasmid DNA library containing a randomized Protospacer-Adjacent-Motif (PAM) sequence, the method comprising: [0277] a) providing a first single stranded oligonucleotide comprising a target sequence that can be recognized by a guide RNA/Cas endonuclease complex; [0278] b) providing a second single stranded oligonucleotide comprising a randomized PAM sequence adjacent to a nucleotide sequence capable of hybridizing with the target sequence of (a); [0279] c) producing an oligoduplex comprising said randomized PAM sequence by combining the first single stranded oligonucleotide of (a) and the second single stranded oligonucleotide of (b); [0280] d) producing a ligation product by ligating the oligoduplex from (c) with a linearized plasmid; and, [0281] e) transforming host cells with the ligation product of (e) and recovering multiple host cell colonies representing the plasmid library. [0282] 2. A method for producing a ligation product containing a randomized Protospacer-Adjacent-Motif (PAM) sequence, the method comprising: [0283] a) providing a first single stranded oligonucleotide comprising restriction endonuclease recognition site located upstream of a target sequence that can be recognized by a guide RNA/Cas endonuclease complex; [0284] b) providing a second single stranded oligonucleotide comprising a randomized PAM sequence adjacent a nucleotide sequence capable of hybridizing with the target sequence of (a); [0285] c) producing an oligoduplex comprising said randomized PAM sequence by combining the first single stranded oligonucleotide of (a) and the second single stranded oligonucleotide of (b); and, [0286] d) producing a ligation product by ligating the oligoduplex from (c) with a linearized plasmid; [0287] 3. The method of embodiment 1, wherein the host cells of (e) are E. coli cells. [0288] 4. A ligation product produced by the method of anyone of embodiments 1-2. [0289] 5. A library of host cells produced by the method of embodiment 1. [0290] 6. The method of anyone of embodiments 1-2, wherein the first single stranded oligonucleotide comprises a restriction endonuclease recognition site located upstream of a target sequence and wherein the ligation product of (d) is produced by first cleaving the oligoduplex with a restriction endonuclease that recognizes the restriction endonuclease recognition site of (a) followed by ligating the cleaved oligoduplex from (d) with a linearized plasmid. [0291] 7. The method of anyone of embodiments 1-2, wherein the second single stranded oligonucleotide comprises a randomized PAM of at least 5 randomized nucleotides (5Ns). [0292] 8. The method of anyone of embodiments 1-2, wherein the second single stranded oligonucleotide comprises a randomized PAM of at least 7 randomized nucleotides (7Ns).
[0293] 9. A method for identification of a Protospacer-Adjacent-Motif (PAM) sequence, the method comprising: [0294] a) providing a library of plasmid DNAs, wherein each one of said plasmid DNAs comprises a randomized Protospacer-Adjacent-Motif sequence integrated adjacent to a target sequence that can be recognized by a guide RNA/Cas endonuclease complex; [0295] b) providing to said library of plasmids a guide RNA and a Cas endonuclease protein, wherein said guide RNA and Cas endonuclease protein can form a complex that is capable of introducing a double strand break into the said target sequence, thereby creating a library of cleaved targets; [0296] c) ligating adaptors to the library of cleaved targets of (b) allowing for the library of cleaved targets to be amplified; [0297] d) amplifying the library of cleaved targets such that cleaved products containing the randomized PAM sequence are enriched, thereby producing a library of enriched PAM-sided targets; [0298] e) sequencing the library of (a) and the library of enriched PAM-sided targets of (d) and identifying the nucleotide sequence adjacent to the cleaved targets of (b) on either strand of the plasmid DNA, wherein said nucleotide sequence represents a putative Protospacer-Adjacent-Motif sequences; and, [0299] f) determining the fold enrichment of each nucleotide within the putative Protospacer-Adjacent-Motif sequence relative to the plasmid DNA library of (a). [0300] 10. The method of anyone of embodiments 1-2 and 9, wherein the randomized PAM sequence comprises at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 randomized nucleotides. [0301] 11. The method of anyone of anyone of embodiments 1-2 and 9, wherein the target sequence is at least 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29 or 30 nucleotides in length. [0302] 12. The method of embodiment 9, wherein the Cas endonuclease is a Cas9 endonuclease from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755. [0303] 13. The method of embodiment 9, wherein the guide RNA comprises a single molecule of a chimeric non-naturally occurring crRNA linked to a tracrRNA. [0304] 14. The method of embodiment 9, wherein the guide RNA comprises a duplex molecule of a chimeric non-naturally occurring crRNA and a tracrRNA. [0305] 15. The method of embodiment 9, wherein the chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to a target sequence in the genome of an organism, wherein said crRNA is linked a tracrRNA originating from organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755. [0306] 16. The method of embodiment 9, wherein the chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to a target sequence in the genome of an organism, wherein said crRNA can form a duplex with a tracrRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755. [0307] 17. The method of embodiment 9, wherein the chimeric non-naturally occurring crRNA comprises at least a fragment of a crRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755. [0308] 18. A recombinant construct comprising at least one of the Protospacer-Adjacent-Motif (PAM) sequence identified by the method of embodiment 9. [0309] 19. A method for identification of a tracrRNA of an organism, the method comprising: [0310] a) providing a first single guide RNA candidate comprising a chimeric non-naturally occurring crRNA comprising a variable targeting domain capable of hybridizing to a target sequence in the genome of a cell, linked to a first nucleotide sequence representing the sense expression of a candidate tracrRNA naturally occurring in said organism; [0311] b) providing a second single guide RNA candidate comprising a chimeric non-naturally occurring crRNA comprising a variable targeting domain capable of hybridizing to a target sequence in the genome of said cell, linked to a second nucleotide sequence representing the sense expression of a candidate tracrRNA naturally occurring in said organism; [0312] c) providing to the first and second single guide RNA candidates a Cas endonuclease protein, wherein said Cas endonuclease protein can form a complex with either the first single guide RNA candidate or the second single guide RNA candidate, wherein said complex is capable of introducing a double strand break into said target sequence; and, [0313] d) identification of the first or second guide RNA candidate and its tracrRNA component that complexes to the Cas endonuclease of (c) and results in cleavage of the target sequence in the genome of said cell. [0314] 20. A method for identification of a tracrRNA of an organism, the method comprising: [0315] a) identifying a CRISPR array repeat sequence in a genomic locus of said organism; [0316] b) aligning the CRISPR array repeat sequence of (a) with the sequence of the genomic locus of (a) and identifying an antirepeat sequence that encodes a tracrRNA; and, [0317] c) determining the transcriptional direction of the tracrRNA. [0318] 21. A guide RNA capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence, wherein said tracrRNA is originated from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755. [0319] 22. A guide RNA capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a single molecule comprising a chimeric non-naturally occurring crRNA linked to a tracrRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence. [0320] 23. A guide RNA capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises at least a fragment of a crRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence. [0321] 24. A guide RNA capable of forming a guide RNA/Cas endonuclease complex, wherein said guide RNA/Cas endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a single molecule comprising a tracrRNA linked to a chimeric non-naturally occurring crRNA comprising at least a fragment of a crRNA originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence. [0322] 25. A guide RNA/Cas endonuclease complex comprising a Cas9 endonuclease originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, and at least one guide RNA, wherein said guide RNA/Cas9 endonuclease complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a target sequence. [0323] 26. The guide RNA/Cas endonuclease complex of embodiment 25 comprising at least one guide RNA of any one of embodiments 21-24. [0324] 27. The guide RNA/Cas endonuclease complex of embodiment 25, wherein said target sequence is located in the genome of a cell. [0325] 28. The guide RNA/Cas endonuclease complex of embodiment 25, wherein said Cas endonuclease is a Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 35 and 81-91, or a functional fragment thereof, wherein said guide RNA/Cas9 endonuclease capable of recognizing, binding to, and optionally nicking or cleaving all or part of a specific DNA target sequence. [0326] 29. A method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, and at least one guide RNA, wherein said guide RNA and Cas endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site. [0327] 30. The method of embodiment 29, further comprising identifying at least one cell that has a modification at said target, wherein the modification at said target site is selected from the group consisting of (i) a replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, and (iv) any combination of (i)-(iii). [0328] 31. A method for editing a nucleotide sequence in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, a polynucleotide modification template, and at least one guide RNA, wherein said polynucleotide modification template comprises at least one nucleotide modification of said nucleotide sequence, wherein said guide RNA and Cas endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site. [0329] 32. A method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one guide RNA, at least one donor DNA, and at least one Cas9 endonuclease originating from an organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said at least one guide RNA and at least one Cas endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site, wherein said donor DNA comprises a polynucleotide of interest. [0330] 33. The method of embodiment 32, further comprising identifying at least one cell that said polynucleotide of interest integrated in or near said target site. [0331] 34. The method of any one of embodiments 29-33, wherein the cell is selected from the group consisting of a human, non-human, animal, bacterial, fungal, insect, yeast, non-conventional yeast, and plant cell.
[0332] 35. The method of embodiment 34, wherein the plant cell is selected from the group consisting of a monocot and dicot cell. [0333] 36. The method of embodiment 35, wherein the plant cell is selected from the group consisting of maize, rice, sorghum, rye, barley, wheat, millet, oats, sugarcane, turfgrass, or switchgrass, soybean, canola, alfalfa, sunflower, cotton, tobacco, peanut, potato, tobacco, Arabidopsis, and safflower cell. [0334] 37. A plant comprising a modified target site, wherein said plant originates from a plant cell comprising a modified target site produced by the method of any of embodiments 29-36. [0335] 38. A plant comprising an edited nucleotide, wherein said plant originates from a plant cell comprising an edited nucleotide produced by the method of embodiment 31. [0336] 39. A method for designing a single guide RNA, the method comprising: [0337] a) aligning a tracrRNA sequence with a CRISPR array repeat sequence from a genomic locus of an organism, wherein said CRISPR array repeat sequence comprises a crRNA sequence; [0338] b) deducing the transcriptional direction of the CRISPR array, thereby also deducing the crRNA sequence; and, [0339] c) designing a single guide RNA comprising said tracrRNA and crRNA sequences [0340] 40. A method for producing target sequences, the method comprising: [0341] a) identifying a polynucleotides of interest; [0342] b) introducing a Protospacer-Adjacent-Motif (PAM) sequence adjacent to said polynucleotide of interest, wherein said PAM sequence comprises the nucleotide sequence NNNNCND, thereby creating a thereby creating a target site for a guide RNA/Cas9 endonuclease complex; and, [0343] c) identifying a polynucleotides of interest; [0344] 41. The method for embodiment 40, wherein the guide RNA/Cas9 endonuclease complex, comprises at least one Cas9 endonuclease originated from organism selected from the group consisting of Brevibacillus laterosporus, Lactobacillus reuteri MIc3, Lactobacillus rossiae DSM 15814, Pediococcus pentosaceus SL4, Lactobacillus nodensis JCM 14932, Sulfurospirillum sp. SCADC, Bifidobacterium thermophilum DSM 20210, Loktanella vestfoldensis, Sphingomonas sanxanigenens NX02, Epilithonimonas tenax DSM 16811, Sporocytophaga myxococcoides and Psychroflexus torquis ATCC 700755, wherein said guide RNA/Cas9 endonuclease complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a target sequence [0345] 42. A method for producing a plasmid DNA library containing a randomized Protospacer-Adjacent-Motif (PAM) sequence, the method comprising transforming at least one host cell with a ligation product and recovering multiple host cell colonies representing the plasmid library, wherein said ligation product was generated by contacting a library of linear oligoduplexes with a linearized plasmid, wherein each oligoduplex member of said library of oligoduplexes comprises a first single stranded oligonucleotide comprising a-target sequence, and a second single stranded oligonucleotide comprising a randomized PAM sequence adjacent to a nucleotide sequence capable of hybridizing with said target sequence. [0346] 43. A method for identification of a Protospacer-Adjacent-Motif (PAM), the method comprising: [0347] a) providing a library of plasmids, wherein each one of said plasmids comprise a randomized Protospacer-Adjacent-Motif sequence integrated adjacent to a target sequence that can be recognized by a guide RNA/Cas endonuclease complex; [0348] b) producing a 3 prime (3') or 5 prime (5') overhang into the target sequence of (a) by providing to the plasmids of (a) a 3 prime deoxy-adenine, a guide RNA and a Cas endonuclease protein, wherein said guide RNA and Cas endonuclease can form a complex that is capable of introducing a double strand break into said target sequence; [0349] c) ligating adapters to the 3 prime or 5 prime overhang of (c), thereby creating a library of cleaved targets that can be amplified; [0350] d) amplifying the library of cleaved targets such that cleaved products containing the randomized PAM sequence are enriched; [0351] e) sequencing the library of (a) and the library of enriched PAM-sided targets of (d) and identifying the nucleotide sequence adjacent to the cleaved targets of (b) on either strand of the plasmid DNA, wherein said nucleotide sequence represents a putative Protospacer-Adjacent-Motif sequences; and, [0352] f) determining the fold enrichment of each nucleotide within the putative Protospacer-Adjacent-Motif sequence relative to the plasmid DNA library of (a). [0353] 44. A single guide RNA selected from the group consisting of SEQ ID NOs: 47, 127, 114-125, and 128-139. [0354] 45. A single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139. [0355] 46. A single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA comprises a chimeric non-naturally occurring crRNA linked to a tracrRNA, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184. [0356] 47. A single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA comprises a chimeric non-naturally occurring crRNA linked to a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160. [0357] 48. A guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence. [0358] 49. A guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence. [0359] 50. A guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said guide RNA is a duplex molecule comprising a chimeric non-naturally occurring crRNA and a tracrRNA, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160, wherein said chimeric non-naturally occurring crRNA comprises a variable targeting domain capable of hybridizing to said target sequence. [0360] 51. A guide RNA/Cas9 endonuclease complex comprising a Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, and at least one guide RNA, wherein said guide RNA/Cas9 endonuclease complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a target sequence. [0361] 52. A guide RNA/Cas9 endonuclease complex comprising at least one guide RNA and a Cas9 endonuclease, wherein said Cas9 endonuclease is encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, and 80, wherein said guide RNA/Cas9 endonuclease complex is capable of recognizing, binding to, and optionally nicking or cleaving all or part of a target sequence. [0362] 53. The guide RNA/Cas9 endonuclease complex of embodiment 7, wherein said guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139. [0363] 54. The guide RNA/Cas9 endonuclease complex of embodiments 7, wherein said target sequence is located in the genome of a cell. [0364] 55. A method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, and at least one guide RNA, wherein said guide RNA and Cas9 endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site. [0365] 56. The method of embodiment 10, further comprising identifying at least one cell that has a modification at said target, wherein the modification at said target site is selected from the group consisting of (i) a replacement of at least one nucleotide, (ii) a deletion of at least one nucleotide, (iii) an insertion of at least one nucleotide, and (iv) any combination of (i)-(iii). [0366] 57. A method for editing a nucleotide sequence in the genome of a cell, the method comprising providing to said cell at least one Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, a polynucleotide modification template, and at least one guide RNA, wherein said polynucleotide modification template comprises at least one nucleotide modification of said nucleotide sequence, wherein said guide RNA and Cas9 endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site. [0367] 58. A method for modifying a target site in the genome of a cell, the method comprising providing to said cell at least one guide RNA, at least one donor DNA, and at least one Cas9 endonuclease selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91, or a functional fragment thereof, wherein said at least one guide RNA and at least one Cas9 endonuclease can form a complex that is capable of recognizing, binding to, and optionally nicking or cleaving all or part of said target site, wherein said donor DNA comprises a polynucleotide of interest. [0368] 59. The method of embodiments 11, 13 or 14, wherein said guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139. [0369] 60. The method of embodiment 13, further comprising identifying at least one cell that said polynucleotide of interest integrated in or near said target site. [0370] 61. The method of any one of embodiments 10-14, wherein the cell is selected from the group consisting of a human, non-human, animal, bacterial, fungal, insect, yeast, non-conventional yeast, and plant cell. [0371] 62. A single guide RNA capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave a target sequence, wherein said single guide RNA comprises a chimeric non-naturally occurring crRNA linked to a tracrRNA, wherein said tracrRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183 and 184, wherein said chimeric non-naturally occurring crRNA comprises a nucleotide sequence selected from the group consisting of SEQ ID NOs: 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159 and 160. [0372] 63. A kit for binding, cleaving or nicking a target sequence in eukaryotic cells or organisms comprising a guide RNA specific for said target DNA, and a Cas endonuclease protein selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91. [0373] 64. A kit for cleaving a target sequence in eukaryotic cells or organisms comprising a guide RNA specific for said target DNA, and a Cas endonuclease protein, wherein said guide RNA is capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave said target sequence, wherein said guide RNA is selected from the group consisting of 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139. [0374] 65. A kit for targeted mutagenesis in eukaryotic cells or organisms comprising a guide RNA specific for said target DNA, a polynucleotide modification template, and a Cas endonuclease protein, wherein said guide RNA is capable of forming a guide RNA/Cas9 endonuclease complex, wherein said guide RNA/Cas9 endonuclease complex can recognize, bind to, and optionally nick or cleave said target sequence, wherein said guide RNA is selected from the group consisting of SEQ ID NOs: 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138 and 139, wherein said Cas endonuclease protein is selected from the group consisting of SEQ ID NOs: 81, 82, 83, 84, 85, 86, 87, 88, 89, 90 and 91. [0375] 66. The kit of any one of embodiments 63-65, further comprising a molecule selected from the group consisting of an inhibitors of NHEJ, an activator of HDR or MMEJ repair pathways, an exogenous sequence, a homologous recombination DNA, a donor DNA, and any one combination thereof.
EXAMPLES
[0376] In the following Examples, unless otherwise stated, parts and percentages are by weight and degrees are Celsius. It should be understood that these Examples, while indicating embodiments of the disclosure, are given by way of illustration only. From the above discussion and these Examples, one skilled in the art can make various changes and modifications of the disclosure to adapt it to various usages and conditions. Such modifications are also intended to fall within the scope of the appended claims.
Example 1
Design and Construction of 5N Randomized Protospacer-Adjacent-Motif (PAM) Library for Assaying Cas9 PAM Preferences
[0377] To characterize the Protospacer-Adjacent-Motif (PAM) specificity of Cas9 proteins from Type II CRISPR (clustered, regularly interspaced, short palindromic repeats)-Cas (CRISPR-associated) nucleic acid-based adaptive immune systems found in most archaea and some bacteria, a plasmid DNA library containing a section of 5 random base pairs immediately adjacent to a 20 base pair target sequence, T1 (CGCTAAAGAGGAAGAGGACA (SEQ ID NO: 1), was developed. Randomization of the PAM sequence was generated through the synthesis of a single oligonucleotide, GG-821N (TGACCATGATTACGAATTCNNNNNTGTCCTCTTCCTCTTTAGCGAGC (SEQ ID NO: 2), with hand-mixing used to create a random incorporation of nucleotides across the 5 random residues (represented as N in the sequence of GG-821N). To convert the single stranded template of GG-821N into a double-stranded DNA template for cloning into the plasmid vector, a second oligonucleotide, GG-820 (AAGGATCCCCGGGTACCGAGCTGCTCGCTAAAGAGGAAGAGGAC (SEQ ID NO: 3), was synthesized with complementation to the 3' end of GG-821N to form a partial oligonucleotide duplex (oligoduplex I) as depicted in FIG. 1. The partial duplex was then extended by PCR using DreamTaq polymerase (Thermo Fisher Scientific) to generate a full duplex containing the target sequence, 5 NNNNN randomized base pairs downstream of the target sequence and cleavage site for the BamHI restriction enzyme (oligoduplex II in FIG. 1). To generate the plasmid library, the oligoduplex, purified using GeneJET PCR Purification Kit (Thermo Fisher Scientific), was digested with BamHI and ligated into pTZ57R/T vector (Thermo Fisher Scientific) pre-cleaved with BamHI. Linear pTZ57R/T vector contains protruding ddT nucleotide at the 3' ends, whereas PCR fragments generated with DreamTaq polymerase contains dA at the 3' ends. Therefore one end of the PCR fragment is ligated into the vector through BamHI sticky ends, while another through NT ends (FIG. 2). The E. coli DH5a strain was transformed (Ca.sup.2+ transformation) with the ligated plasmid library and plated onto Luria Broth (LB) containing agar. The transformation efficiency was estimated from plated dilutions. Overall, .about.12,000 colonies were recovered. The colonies were harvested from the plate by gently resuspending them in liquid LB media and plasmid DNA was purified using GeneJET Plasmid Miniprep kit (Thermo Fisher Scientific).
[0378] To validate the randomness of the resulting PAM library, PCR fragments spanning the 5 bp randomized PAM region were generated by Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific) amplification (15 cycles of a 2-step amplification protocol) using a TK-119 (GAGCTCGCTAAAGAGGAAGAGG (SEQ ID NO: 4) and pUC-dir (GCCAGGGTTTTCCCAGTCACGA (SEQ ID NO: 5) primer pair and 50 ng of plasmid DNA library as template. The resulting 122 bp PCR product was purified using GeneJET PCR Purification Kit (Thermo Fisher Scientific). 40 ng of the resulting PCR product was then amplified with Phusion.RTM. High Fidelity PCR Master Mix (New England Biolabs, M0531L) adding on the sequences necessary for amplicon-specific barcodes and Illumnia sequencing using "tailed" primers through two rounds of PCR each consisting of 10 cycles. The primers used in the primary PCR reaction are shown in Table 2 and a set of primers (AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACG (Universal Forward, SEQ ID NO: 8) and CAAGCAGAAGACGGCATA (Universal Reverse, SEQ ID NO: 9) universal to all primary PCR reactions were utilized for the secondary PCR amplification. The resulting PCR amplifications were purified with a Qiagen PCR purification spin column, concentration measured with a Hoechst dye-based fluorometric assay, combined in an equimolar ratio, and single read 60-100 nucleotide-length deep sequencing was performed on IIlumina's MiSeq Personal Sequencer with a 5-10% (v/v) spike of PhiX control v3 (Illumina, FC-110-3001) to off-set sequence bias. The PAM sequence for only those reads containing a perfect 12 nt sequence match flanking either side of the 5 nucleotide randomized PAM sequence were captured and used to examine the frequency and diversity of PAM sequences present in the library. The frequency of each PAM sequence was calculated by dividing the number of reads with a given PAM by the total number of reads. The PAM sequence distribution was visualized by ordering the frequency of each PAM from greatest to least and displaying them graphically and by calculating the standard deviation of the resulting PAM frequencies relative to the average. As shown in FIG. 4, all 1,024 possible PAM sequences were present at an average frequency of 0.10% with a coefficient of variation of 40.86%.
TABLE-US-00002 TABLE 2 Primary PCR primer sequences for tailing on the sequences needed for Illumina deep sequencing of initial uncut 5 bp randomized PAM pTZ57R/T library. Primer Primer SEQ Name Orientation Primary PCR Primer Sequence ID NO. JKYS800.1 Forward CTACACTCTTTCCCTACACGACGCTCTTCCGATCT 6 AAGTGAGCTCGCTAAAGAGGAAGA JKYS803 Reverse CAAGCAGAAGACGGCATACGAGCTCTTCCGATCTG 7 AATTCGAGCTCGGTACCT
Example 2
Protein Expression and Purification of Streptococcus pyogenes, Streptococcus thermophilus CRISPR1 and Streptococcus thermophilus CRISPR3 Cas9 Proteins
[0379] To examine the PAM specificity of the Cas9 proteins from the Streptococcus pyogenes (Spy) (Jinek et al. (2012) Science 337:816-21), Streptococcus thermophilus CRISPR1 (Sth1) (Horvath et al. (2008) Journal of Bacteriology 190:1401-12) and Streptococcus thermophilus CRISPR3 (Sth3) (Horvath et al. (2008) Journal of Bacteriology 190:1401-12) Type II CRISPR-Cas systems, Spy, Sth1 and Sth3 Cas9 proteins were E. coli expressed and purified. Briefly, the cas9 genes of the CRISPR1-Cas and CRISPR3-Cas systems of Streptococcus thermophilus (Sth1 and Sth3) were amplified from a genomic DNA sample, while the cas9 gene of Streptococcus pyogenes (Spy) was amplified from a plasmid, pMJ806 (Addgene plasmid #39312)). DNA fragments encoding Sth1, Sth3 and Spy Cas9 were PCR amplified using Sth1-dir/Sth1-rev (ACGTCTCACATGACTAAGCCATACTCAATTGGAC (SEQ ID NO: 10); ACTCGAGACCCTCTCCTAGTTTGGCAA (SEQ ID NO: 11), Sth3-dir/Sth3-rev (GGGGGGTCTCACATGAGTGACTTAGT (SEQ ID NO: 12); AATTACTCGAGAAAATCTAGCTTAGGCTTA (SEQ ID NO: 13) and Spy-dir/Spy-rev (AAGGTCTCCCATGGATAAGAAATACTCAATAGGCTTAG (SEQ ID NO: 14); TTCTCGAGGTCACCTCCTAGCTGACTCAAATC (SEQ ID NO: 15) primer pairs, accordingly, and ligated into a pBAD24-CHis expression vector digested over NcoI and XhoI sites.
[0380] Sth3 and Spy Cas9 proteins were expressed in E. coli DH10HB strain grown in LB broth supplemented with ampicillin (100 mg/ml). Cells were grown at 37.degree. C. to an OD 600 of 0.5 at which time the growth temperature was decreased to 16.degree. C. and expression induced with 0.2% (w/v) arabinose for 20 h. Cells were pelleted and resuspended in loading buffer (20 mM KH.sub.2PO.sub.4 pH7.0, 0.5 M NaCl, 10 mM imidazole, 5% glycerol) and disrupted by sonication. Cell debris was removed by centrifugation. The supernatant was loaded onto the Ni.sup.2+-charged 5 ml HiTrap chelating HP column (GE Healthcare) and eluted with a linear gradient of increasing imidazole concentration. The fractions containing Cas9 were pooled and subsequently loaded onto HiTrap heparin HP column (GE Healthcare) for elution using a linear gradient of increasing NaCl concentration (from 0.5 to 1 M NaCl). The fractions containing Cas9 were pooled and dialyzed against 10 mM Bis-Tris-HCl pH 7.0, 300 mM KCl, 1 mM EDTA, 1 mM DTT, 50% (v/v) glycerol and stored at -20.degree. C.
Example 3
Identification of PAM Preferences for Streptococcus pyogenes and Streptococcus thermophilus CRISPR3 Cas9 Proteins
[0381] To empirically examine the PAM preferences for Streptococcus pyogenes (Spy) and Streptococcus thermophilus CRISPR3 (Sth3) Cas9 proteins, the randomized PAM library described in Example 1 was subject to digestion with purified Sth3 and Spy Cas9 proteins and guide RNA containing a variable targeting domain that hybridizes with, i.e., is complementary to, a sequence in the target DNA molecule (referred herein as target sequence), T1 (SEQ ID NO: 1). Sth3 and Spy Cas9-crRNA-tracrRNA complexes were assembled by mixing Cas9 protein with pre-annealed crRNA and tracrRNA duplex (Table 3) at 1:1 molar ratio followed by incubation in a complex assembly buffer (10 mM Tris-HCl pH 7.5 at 37.degree. C., 100 mM NaCl, 1 mM EDTA, 1 mM DTT) at 37.degree. C. for 1 h. 1 .mu.g of plasmid DNA library with randomized 5 bp NNNNN PAM was cleaved with 50 nM and 100 nM of Cas9 complex in a reaction buffer (10 mM Tris-HCl pH 7.5 at 37.degree. C., 100 mM NaCl, 10 mM MgCl.sub.2, 1 mM DTT) for 60 min. at 37.degree. C. in a 100 .mu.l reaction volume (FIG. 3).
TABLE-US-00003 TABLE 3 RNA molecules used for Sth3 and Spy Cas9-crRNA-tracrRNA complex assembly. Name Sequence (5'-3') Origin SEQ ID NO. Sth3 crRNA CGCUAAAGAGGAAGAGGACAGUUUUAGAGC Synthetic 16 UGUGUUGUUUCG oligonucleotide Sth3 GGGCGAAACAACACAGCGAGUUAAAAUAAG In vitro 17 tracrRNA GCUUAGUCCGUACUCAACUUGAAAAGGUGG transcription CACCGAUUCGGUGUUUUU Spy crRNA CGCUAAAGAGGAAGAGGACAGUUUUAGAGC Synthetic 18 UAUGCUGUUUUG oligonucleotide Spy GGGAAACAGCAUAGCAAGUUAAAAUAAGGC In vitro 19 tracrRNA UAGUCCGUUAUCAACUUGAAAAAGUGGCAC transcription CGAGUCGGUGCUUUUUUU
[0382] To efficiently capture the blunt-ends of the plasmid library generated by Sth3 or Spy cleavage, a 3' dA was added by incubating the completed digestion reactions with 2.5 U of DreamTaq DNA Polymerase (Thermo Fisher Scientific) and 0.5 .mu.l of 10 mM dATP (or dNTP) for an additional 30 min. at 72.degree. C. (FIG. 3). Reaction products were purified using GeneJET PCR Purification Kit (Thermo Fisher Scientific). Next adapters with a 3' dT overhang were generated by annealing TK-117 (CGGCATTCCTGCTGAACCGCTCTTCCGATCT (SEQ ID NO: 20) and phosphorylated TK-111 (GATCGGAAGAGCGGTTCAGCAGGAATGCCG (SEQ ID NO: 21) oligonucleotides. 100 ng of the resulting adapter was ligated to an equal concentration of the purified 3' dA overhanging cleavage products for 1 hour at 22.degree. C. in a 25 .mu.l reaction volume in ligation buffer (40 mM Tris-HCl pH 7.8 at 25.degree. C., 10 mM MgCl.sub.2, 10 mM DTT, 0.5 mM ATP, 5% (w/v) PEG 4000, 0.5 U T4 Ligase; Thermo Fisher Scientific) (FIG. 3). Next, to selectively enrich for cleaved products containing the PAM sequence, PCR amplification was performed with a forward primer, pUC-dir (SEQ ID NO: 5), specific to the PAM-side of the cleaved pTZ57R/T plasmid vector and with a reverse primer, TK-117 (SEQ ID NO: 20), specific to the ligated TK-117/TK-111 adapter sequence (FIG. 3). PCR fragments were generated by Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific) amplification (15 cycles of a 2-step amplification protocol) with 10 .mu.l of ligation reaction mixtures as a template (in 100 .mu.l total volume). The resulting 131 bp PCR products amplified from the Cas9 pre-cleaved plasmid libraries were purified with GeneJET PCR Purification Kit (Thermo Fisher Scientific) and prepared for IIlumina deep sequencing as described in Example 1 except the barcode containing forward primers used in the primary reaction were specific to the TK-117/TK-111 adapter sequence and are shown in Table 4 (FIG. 3).
TABLE-US-00004 TABLE 4 Primary PCR primer sequences for tailing on the sequences needed for Illumina deep sequencing of cleaved and adapter ligated 5 bp randomized PAM pTZ57R/T library Primer Digestion Primer SEQ Name Experiment Orientation Primary PCR Primer Sequence ID NO. JKYS807.1 50 nM Sth3 Forward CTACACTCTTTCCCTACACGACGCTCTTCC 22 GATCTAAGGCGGCATTCCTGCTGAAC JKYS807.2 100 nM Sth3 Forward CTACACTCTTTCCCTACACGACGCTCTTCC 23 GATCTTTCCCGGCATTCCTGCTGAAC JKYS807.3 50 nM Spy Forward CTACACTCTTTCCCTACACGACGCTCTTCC 24 GATCTGGAACGGCATTCCTGCTGAAC JKYS807.4 100 nM Spy Forward CTACACTCTTTCCCTACACGACGCTCTTCC 25 GATCTCCTTCGGCATTCCTGCTGAAC
The resulting IIlumina compatible libraries were then sequenced as described in Example 1. The PAM sequence for only those reads containing a perfect 12 nt sequence match flanking either side of the 5 nucleotide randomized PAM sequence were captured and used to examine the frequency and diversity of PAM sequences present in the Sth3 and Spy Cas9-guide RNA cleaved libraries. Given the inherent bias in the uncut library observed in FIG. 4 and described in Example 1, PAM preferences were calculated relative to the uncut library by dividing the frequency of a given PAM from the Sth3 or Spy Cas9-guide RNA digested library by the frequency of the same PAM sequence in the uncut library with the resulting value being represented as a fold enrichment correlative to the uncut control. To examine the PAM preferences of Sth3 and Spy Cas9 proteins, the percent nucleotide composition of the PAM sequences with fold enrichment relative to the uncut control were examined. As shown in FIG. 5 and FIG. 6, the canonical PAM preferences for both Sth3 and Spy Cas9 proteins, NGGNG and NGG, respectively, are observed in both the 50 nM and 100 nM digests. For Sth3 Cas9 protein, a slight preference (not previously reported) for a C or T bp at position 1 is also evident. Next, the effect of decreasing Sth3 and Spy Cas9-crRNA-tracrRNA complex concentration and digestion time on PAM preferences was examined. To this end, the minimal Cas9 concentration and shortest time where PCR amplified cleavage products may still be obtained from the randomized PAM plasmid library were determined. First, the reaction time was held constant at 60 minutes while the Cas9-crRNA-tracrRNA complex concentration was varied between 0.5-100 nM. Next, the Cas9-crRNA-tracrRNA complex concentration was fixed at 50 nM and the reaction time was varied between 1-60 minutes. Optimization of the cleavage reaction conditions revealed that the concentration and cleavage time for Sth3 and Spy Cas9 complexes could be reduced to 0.5 nM (at a 60 min. incubation time) or 1 min. (at a 50 nM concentration of Cas9 complex), respectively (FIG. 7).
[0383] To examine the PAM sequences present in the minimally digested Sth3 and Spy Cas9-guide RNA libraries, 0.5 nM-60 minute and 50 nM-1 minute PCR amplified cleavage products were purified with the GeneJET PCR Purification Kit (Thermo Fisher Scientific) and subjected to IIlumina deep sequencing as described above for the 50 nM and 100 nM-60 minute Sth3 and Spy digests. As a positive control and to demonstrate the reproducibility of PAM preferences derived from our assay, the 50 nM-60 minute digests for Sth3 and Spy were repeated and IIlumina deep sequenced again. PAM preference analysis was carried-out as described above for the Sth3 and Spy (50 nM and 100 nM-60 minute digests) examining the percent nucleotide composition of the PAM sequences with fold enrichment relative to the uncut library. As shown in FIG. 8 and FIG. 9, the positive controls (Sth3 and Spy 50 nM-60 minute digests) demonstrated very similar trends in PAM preferences compared to that observed previously indicating a high degree of assay reproducibility. The PAM preferences observed in the minimally Sth3 and Spy digested libraries compared to that exhibited by the respective 50 nM-60 minute positive control are shown in FIG. 10 and FIG. 11. When the concentration of Sth3 Cas9-crRNA-tracrRNA complex is lowered to 0.5 nM, the percentage of uncanonical PAM residues cleaved by Sth3 decreases; resulting in a tightening of specificity (FIG. 10). This is most evident at positions 2 and 3 where on-nucleotide preferences for a G increase and off-nucleotide preferences decrease. A similar shift in PAM preference towards the reported PAM sequence for Spy (NGG) is observed when the Spy Cas9-crRNA-tracrRNA complex is lowered to 0.5 nM. Here the percentage of PAMs with an uncanonical A residue at position 2 declines from over 20% in the 50 nM-60 minute and 50 nM-1 minute digests to almost zero in the 0.5 nM-60 minute digest (FIG. 11).
[0384] Next, the effect of using a chimeric fusion of crRNA and tracrRNA (single guide RNA (sgRNA)) (Jinek et al. (2012) Science 337:816-21 and Gasiunas et al. (2012) Proc. Natl acad. Sci. USA 109: E2579-E2586) on Sth3 and Spy Cas9 PAM preferences was assayed. Digestion, enrichment, IIlumina deep sequencing and PAM preference analysis was carried-out as described above against the randomized 5 bp PAM plasmid DNA library except a sgRNA (Table 5) was used in place of the crRNA-tracrRNA duplex and digests were only performed with 0.5 nM of sgRNA-Cas9 complex for 60 min.
TABLE-US-00005 TABLE 5 RNA molecules used for Cas9-sgRNA complex assembly. Name Sequence (5'-3') Origin SEQ ID NO. Sth3 sgRNA GGGCGCUAAAGAGGAAGAGGACAGUUUUAGAGCU In vitro 26 GUGUUGUUUCGGUUAAAACAACACAGCGAGUUAA transcription AAUAAGGCUUAGUCCGUACUCAACUUGAAAAGGU GGCACCGAUUCGGUGUUUUUU Spy sgRNA GGGCGCUAAAGAGGAAGAGGACAGUUUUAGAGCU In vitro 27 AGAAAUAGCAAGUUAAAAUAAGGCUAGUCCGUUA transcription UCAACUUGAAAAAGUGGCACCGAGUCGGUGCUUU UUU
[0385] As shown in FIG. 12 and FIG. 13, the PAM preferences for Sth3 and Spy Cas9 proteins (NGGNG and NGG respectively) are nearly identical regardless of the type of guide RNA used; either a crRNA-tracrRNA duplex or sgRNA.
Example 4
Identification of PAM Preferences for Streptococcus thermophilus CRISPR1 Cas9 Protein
[0386] To empirically examine the PAM preferences for Streptococcus thermophilus CRISPR1 (Sth1) Cas9 protein with a reported PAM sequence of 7 nucleotides, NNAGAAW (Horvath et al. (2008) Journal of Bacteriology 190:1401-12), a randomized 7 bp PAM plasmid DNA library was generated as described for the 5 bp randomized PAM library in Example 1 with the following modifications. Randomization of the PAM sequence was generated through the synthesis of four oligonucleotides, GG-940-G (GTGCACGCCGGCGACGTTGGGTCAACTNNGNNNNTGTCCTCTTCCTCTTTAG CGTTTAG (SEQ ID NO: 28), GG-940-C (GTGCACGCCGGCGACGTTGGGTCAACTNNCNNNNTGTCCTCTTCCTCTTTAG CGTTTAG (SEQ ID NO: 29), GG-940-A (GTGCACGCCGGCGACGTTGGGTCAACTN NAN N N NTGTCCTCTTCCTCTTTAG CGTTTAG (SEQ ID NO: 30) and GG-940-T (GTGCACGCCGGCGACGTTGGGTCAACTN NTN N N NTGTCCTCTTCCTCTTTAG CGTTTAG (SEQ ID NO: 31), with hand-mixing used to create a random incorporation of nucleotides across the random residues (represented as N). The randomized single stranded oligonucleotides were each separately converted into double-stranded DNA templates for cloning into the plasmid vector using a second oligonucleotide, GG-939 (GACTAGACCTGCAGGGGATCCCGTCGACAAATTCTAAACGCTAAAGAGGAAG AGGAC (SEQ ID NO: 126), with complementation to the 3' end of GG-940-G, GG-940-C, GG-940-A and GG-940-T and by PCR extension with DreamTaq polymerase (Thermo Fisher Scientific) (oligoduplexes I & II FIG. 1). To avoid cleavage of some species of the randomized positions, the resulting double-stranded templates were each digested with an 8 bp cutting restriction endonuclease, SdaI, so that overhangs were present at each end; a PstI compatible overhang and a Taq added single 3' A overhang. The resulting overhangs were used to directionally ligate the 4 double-stranded templates into pTZ57R/T (Thermo Fisher Scientific) pre-cleaved with PstI. The ligations were Ca.sup.2+ transformed into DH5a E. coli cells, plasmid DNA was recovered and combined from each of the 4 transformants derived from GG-940-G, GG-940-C, GG-940-A and GG-940-T to generate the randomized 7 bp NNNNNNN PAM plasmid DNA library.
[0387] PAM preference experiments with Sth1 Cas9 protein on the resulting 7 bp randomized PAM plasmid DNA library were carried-out similarly to that described in Example 3 for the Streptococcus thermophilus CRISPR3 (Sth3) and Streptococcus pyogenes (Spy) Cas9 proteins (against the 5 bp randomized PAM library). Briefly, Sth1 Cas9-crRNA-tracrRNA complexes were assembled by mixing Cas9 protein with pre-annealed crRNA and tracrRNA duplex (Table 6) at 1:1 molar ratio followed by incubation in a complex assembly buffer (10 mM Tris-HCl pH 7.5 at 37.degree. C., 100 mM NaCl, 1 mM EDTA, 1 mM DTT) at 37.degree. C. for 1 h. Digests were performed using 1 .mu.g of randomized 7 bp PAM library with 50 nM Sth1 crRNA-tracrRNA-Cas9 complexes at 37.degree. C. for 60 min., 50 nM Sth1 crRNA-tracrRNA-Cas9 complexes at 37.degree. C. for 1 min. and 0.5 nM Sth1 crRNA-tracrRNA-Cas9 complexes at 37.degree. C. for 60 min. (FIG. 3). As a positive control, 1 .mu.g of the randomized 7 bp PAM library was also digested with Sth3 and Spy Cas9-sgRNA complexes (0.5 nM at 37.degree. C. for 60 min.). A 3' dA was added to the blunt-ends of the cleaved fragments (FIG. 3). Next, duplexed adapter TK-117/TK-111 with a 3' dT overhang was ligated to the A overhang (FIG. 3). Then, PCR was assembled using primers pUC-dir (SEQ ID NO: 5) and TK-117 (SEQ ID NO: 20) to enrich for PAM sequences that supported cleavage (FIG. 3). 40 ng of the resulting PCR product was then amplified with Phusion.RTM. High Fidelity PCR Master Mix (New England Biolabs, M0531L) adding on the sequences necessary for amplicon-specific barcodes and Illumina sequencing using "tailed" primers through two rounds of PCR each consisting of 10 cycles (FIG. 3). The sequences of the barcode specific forward primers used in the primary PCR reaction were similar to those listed in Table 3 and the reverse primer, JKYS812 (CAAGCAGAAGACGGCATACGAGCTCTTCCGATCTCGGCGACGTTGGGTC (SEQ ID NO: 32)), was paired with each of the forward primers. A set of primers, AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACG (Universal Forward, SEQ ID NO: 8) and CAAGCAGAAGACGGCATA (Universal Reverse, SEQ ID NO: 9), universal to all primary PCR reactions were utilized for the secondary PCR amplification.
TABLE-US-00006 TABLE 6 RNA molecules used for Sth1 Cas9-crRNA-tracrRNA complex assembly. Name Sequence (5'-3') Origin SEQ ID NO. Sth1 crRNA CGCUAAAGAGGAAGAGGACAGUUUUUGUACU Synthetic 33 CUCAAGAUUUA oligonucleotide Sth1 tracrRNA GGGUAAAUCUUGCAGAAGCUACAAAGAUAAG In vitro 34 GCUUCAUGCCGAAAUCAACACCCUGUCAUUU transcription UAUGGCAGGGUGUUUUCG
[0388] The resulting PCR amplifications were prepared and IIlumina deep sequenced as described in Example 1 and PAM preference analysis was carried-out as described in Example 3 for the Sth3 and Spy (50 nM and 100 nM-60 minute digests) examining the percent nucleotide composition of the PAM sequences with fold enrichment relative to the uncut library. As shown in FIG. 14 and FIG. 15, the PAM preferences for the positive controls, Sth3 and Spy Cas9 proteins, are nearly identical regardless of the length of the randomized PAM plasmid DNA library used; either 5 bp or 7 bp. The PAM preferences observed for the Sth1 Cas9 protein are shown in FIG. 16 and match those previously reported (NNAGAAW). Just as observed for Sth3 and Spy Cas9 proteins, the PAM specificity of Sth1 is more relaxed at higher concentrations of guide RNA-Cas9 complex. This is most evident at position 5 where an off-preference for a C nucleotide is less prevalent at the lower 0.5 nM complex concentration.
[0389] Since the canonical PAM sequence preferences for Spy, Sth1 and Sth3 Cas9 proteins may be recapitulated with our assay regardless of the type of guide RNA used (either crRNA-tracrRNA or sgRNA) or length of the randomized PAM sequence, suggests that the in vitro PAM library assay described herein or derivations of it may be used to directly interrogate PAM specificity from any Cas9 assuming the guide RNA sequences, either crRNA-tracrRNA or sgRNA, may be successfully deduced. Additionally, our assay grants precise control over the amount of Cas9 protein used in the in vitro digestion assays described herein allowing a detailed examination of Cas9 PAM specificity as a function of Cas9-guide RNA complex concentration as evident by the apparent broadening in PAM specificity as Cas9-guide RNA complex concentration was increased.
Example 5
Identification of Brevibacillus laterosporus crRNA, tracrRNA and Cas9 Endonuclease
[0390] To empirically examine the PAM preferences for a Cas9 protein whose PAM was undefined, a cas9 gene from an uncharacterized Type II CRISPR-Cas system was identified by searching internal Pioneer-DuPont databases consisting of microbial genomes with the amino acid sequence of S. thermophilus CRISPR3 (Sth3) Cas9 (SEQ ID NO: 35). Amino acid alignment of Sth3 revealed 12.9% identity and 24.4% similarity at the protein level with a protein derived from a single long open-reading-frame of 3279 nucleotides (SEQ ID NO: 36) from the Brevibacillus laterosporus bacterial strain SSP360D4. Translation of the open-reading-frame encodes a protein of 1092 amino acids (not including the stop codon). Based on PFAM database searches the protein contained HNH endonuclease and CRISPR-associated domains all hallmarks of a Cas9 protein. The cas9 gene of SSP360D4 was also located upstream of a CRISPR array comprised of 7 repeat-spacer units (FIG. 17A). The repeat and spacer length (36 and 30 bp, accordingly) is similar to other Type II CRISPR-Cas systems. However, 5 of 8 repeats contain 1 or 2 bp mutations (FIG. 17B). Sequences of Repeat1 (SEQ ID NO: 37), Repeat4 (SEQ ID NO: 40) and Repeat5 (SEQ ID NO: 41) are conserved; therefore this sequence was selected as a template for designing single guide RNAs (sgRNAs). A region upstream of the cas9 gene is partially complementary (anti-repeat) to the 5'-terminus of the repeat suggesting a putative tracrRNA (FIG. 17A). The possible transcriptional directions of the putative tracrRNA were considered by examining the secondary structures and possible termination signals present in a RNA version of the sense and anti-sense genomic DNA sequences surrounding the anti-repeat. However, the transcriptional direction of the tracrRNA and CRISPR region could not be reliably determined bioinformatically, so a method described in Example 7 was designed to empirically determine the appropriate directions of transcription. Other genes typically found in a Type II CRISPR-Cas locus were either truncated, as was the case for cas1, or missing (FIG. 17A).
Example 6
Protein Expression and Purification of Brevibacillus laterosporus Cas9 Protein
[0391] To examine the PAM specificity and guide RNA of Brevibacillus laterosporus (Blat) Cas9 protein with the in vitro cleavage assays described in Examples 7& 8, Blat Cas9 protein was E. coli expressed and purified. Briefly, a DNA fragment encoding the Brevibacillus laterosporus Cas9 protein was PCR amplified directly from the Pioneer-DuPont strain, SSP360D4, using Blat-Cas9-dir (TACCATGGCATACACAATGGGAATAGATG (SEQ ID NO: 45) and Blat-Cas9-rev (TTCTCGAGACGACTAGTTGATTTAATCGAATTGAC (SEQ ID NO: 46) primer pair and cloned into a pBAD24-CHis expression vector pre-cleaved over NcoI and XhoI sites. To establish optimal expression conditions three different E. coli strains, BL21 (DE3), DH10B and Rosetta (DE3), were analyzed. Highest expression yield of soluble Blat Cas9 protein was obtained in the BL21 (DE3) strain.
[0392] For purification, Blat Cas9 protein was expressed in E. coli BL21 (DE3) strain grown in LB broth supplemented with ampicillin (100 mg/ml). Cells were grown at 37.degree. C. to an OD 600 of 0.5 at which time the growth temperature was decreased to 16.degree. C. and expression induced with 0.2% (w/v) arabinose for 20 h. Cells were pelleted and resuspended in loading buffer (20 mM KH.sub.2PO.sub.4 pH7.0, 0.5 M NaCl, 10 mM imidazole, 5% glycerol) and disrupted by sonication. Cell debris was removed by centrifugation. The supernatant was loaded onto the Ni.sup.2+-charged 5 ml HiTrap chelating HP column (GE Healthcare) and eluted with a linear gradient of increasing imidazole concentration. The fractions containing Cas9 were pooled and subsequently loaded onto heparin column for elution using a linear gradient of increasing NaCl concentration (from 0.5 to 1 M NaCl). The fractions containing Cas9 were pooled and dialyzed against 10 mMBis-Tris-HCl pH 7.0, 300 mM KCl, 1 mM EDTA, 1 mM DTT, 50% (v/v) glycerol and stored at -20.degree. C.
Example 7
Determination of Guide RNAs for the Cas9 of Brevibacillus laterosporus
[0393] To determine a guide RNA for the Cas9 protein identified in the Brevibacillus laterosporus (Blat) Type II CRISPR-Cas system, we designed two single guide RNA (sgRNA) variants to account for both possible expression scenarios of the tracrRNA and CRISPR array (FIG. 18 & Table 7) and used them to probe which expression scenario supported cleavage activity of Blat Cas9 in the 7 bp randomized PAM plasmid DNA library from Example 4.
[0394] sgRNAs were designed by first identifying the boundaries of the putative tracrRNA molecules by analyzing regions which were partially complementary to the 22 nt 5' terminus of the repeat (anti-repeat). Next, to determine the 3' end of the tracrRNA, possible secondary structures and terminators were used to predict the region of termination in the downstream fragment (FIGS. 19 and 20) using Mini-fold (Markham et al. (2008) Methods in Molecular Biology 453: 3-31). The sgRNAs contained a T7 polymerase transcription initiation recognition signal at the 5' end followed by a 20 nt target recognition sequence, 16 nt of crRNA repeat, 4 nt self-folding hairpin loop and anti-repeat sequence complementary to the repeat region of the crRNA followed by the remaining 3' part of the putative tracrRNA (Table 7). The sgRNA variant which contains a putative tracrRNA transcribed in the same direction as the cas9 gene is termed "direct" sgRNA, while the sgRNA containing the tracrRNA transcribed in the opposite direction a "reverse" sgRNA (FIG. 18).
[0395] The "direct" sgRNA encoding gene was obtained in two PCR steps. First two fragments were generated by PCR using GG-969/GG-839 and TK-149/TK-150 oligonucleotide primer pairs (Table 8). The fragments were purified with the GeneJET PCR Purification Kit (Thermo Fisher Scientific) and the full length sgRNA gene was assembled from these fragments by overlapping PCR using GG-969/TK-150 primer pairs. The "reverse" sgRNA encoding gene was amplified by PCR using GG-840/GG-841 oligonucleotide primer pairs (Table 8). To generate the sgRNA encoding plasmids pUC-Blat-dir-sgRNA and pUC-Blat-rev-sgRNA, the PCR fragments were cloned into pUC18 vector digested with SacI.
TABLE-US-00007 TABLE 7 "Direct" and "reverse" Blat sgRNAs used to deduce transcriptional direction of crRNA and tracrRNA loci. Variable Remaining T7 Targeting 16 nt Putative 3' Blat Transcription domain of the Anti- tracrRNA SEQ sgRNA Initiation (SEQ ID NO:) repeat Loop Repeat Sequence ID NO: Direct GGG 193 195 GAAA 197 199 47 Reverse GGG 194 196 GAAA 198 200 48
TABLE-US-00008 TABLE 8 Oligonucleotides used for Blat sgRNA gene construction and sgRNA production. Name Sequence (5'-3') SEQ ID NO. GG-969 GGGCGCTAAAGAGGAAGAGGACAGCTATAGTTCCTTACTGAAAGGTAA 49 GTTGCTATAGTAAGGGCAAC GG-839 CTAAAAACGGGCTAGGCGATCCCCAACGCCTCGGGTCTGTTGCCCTTA 50 CTATAGCAACTTAC 149 GATCGCCTAGCCCGTTTTTACGGGCTCTCCCCATATTCAAAATAATGA CAGACGA TK-150 AAAAAAAAGCACCTCGGAAATAAATGCTCCAAGGTGCTCGTCTGTCAT TATTTTGAATATGG GG-840 GGGCGCTAAAGAGGAAGAGGACAATCATATCATATCGAGGAAACTTGA 53 TATGATATGATACTTTCATTTTA GG-841 CATAAAATAGACAGATAAATGAGATTGACTTCGATGATATATGGATAT 54 AAAATGAAAGTATCATATCATATCAAG TK-124 TAATACGACTCACTATAGGGCGCTAAAGAGGAAGAGG 55 TK-151 AAAAAAAAGCACCTCGGAAATAAATG 56 TK-126 ATAAAATAGACAGATAAATGAGATTGACTTCG 57 indicates data missing or illegible when filed
"Direct" and "reverse" Blat sgRNAs were obtained by in vitro transcription using TranscriptAid T7 High Yield Transcription Kit (Thermo Fisher Scientific) from the PCR fragments containing a T7 promoter at the proximal end of the RNA coding sequence. The "direct" sgRNA encoding fragment (177 nt) was generated using the TK-124/TK-151primer pair (Table 8) with pUC-Blat-dir-sgRNA plasmid DNA as template, whereas the "reverse" sgRNA encoding fragment (118 nt) was generated using the TK-124/TK-126 primer pair with pUC-Blat-rev-sgRNA plasmid as template (Table 7). The resulting sgRNAs were purified using GeneJET RNA Cleanup and Concentration Micro Kit (Thermo Fisher Scientific) and used for complex assembly. Blat Cas9-sgRNA complexes were assembled by mixing Cas9 protein with sgRNA at 1:1 molar ratio followed by incubation in a complex assembly buffer (10 mM Tris-HCl pH 7.5, 100 mM NaCl, 1 mM EDTA, 1 mM DTT) at 37.degree. C. for 1 h. Blat Cas9 cleavage of the 7 bp randomized PAM plasmid DNA library was performed similarly as described above for Spy and Sth3 Cas9 proteins (Example 3). Briefly, 50 nM of Blat Cas9 complexes, assembled using "direct" or "reverse" sgRNAs, respectively, were incubated with 1 .mu.g plasmid DNA for 60 min at 37.degree. C. After library digestion and addition of 3' dA overhangs, adapters were ligated and cleavage products were PCR amplified (FIG. 3). Analysis of reaction products by agarose gel electrophoresis revealed that the "direct" sgRNA, but not the "reverse" sgRNA supported plasmid library cleavage (FIG. 21). Single guide RNAs targeting a target site in the genome of an organism can be designed by changing the targeting sequence of SEQ ID NO: 47 with any random nucleotide that can hybridize to any desired target sequence (guide RNA as shown in SEQ ID NO: 127).
Example 8
Identification of PAM Preferences for Brevibacillus laterosporus Cas9 Protein
[0396] After determining a guide RNA for Brevibacillus laterosporus (Blat) Cas9, PAM identification was performed similarly to that described in Example 3 for the Spy and Sth3 Cas9 proteins. Briefly, 1 .mu.g of 7 bp randomized PAM plasmid library was digested with various concentrations of Blat Cas9-"direct" sgRNA complex, ranging between 0.5-50 nM, and at various reaction times, ranging from 1 to 60 minutes. Next, 3' dA overhangs were added to the cleavage products, adapters ligated and adapter-ligated cleavage products were PCR amplified. PCR reactions were then electrophoresed on a 1% agarose gel and visualized. As shown in FIG. 22 and similarly to that described for Sth3 and Spy Cas9 proteins, the minimal concentration and cleavage time needed to support visualization after PCR amplification were 0.5 nM (at a 60 min. incubation time) or 1 min. (at a 50 nM concentration of Cas9 complex). Next, the amplifications for the 50 nM-60 min., 50 nM-1 min. and 0.5 nM-60 min. digests were purified with the GeneJET PCR Purification Kit (Thermo Fisher Scientific) and Illumina sequencing anchors added by two rounds of PCR as described in Example 3 for the Sth3 and Spy Cas9 proteins when examining their PAM preferences with the 7 bp randomized PAM library. The resulting Illumina compatible libraries were then sequenced as described in Example 1 and PAM preference analysis was carried-out as described in Example 3 for the Sth3 and Spy (50 nM and 100 nM-60 minute digests) examining the percent nucleotide composition of the PAM sequences with fold enrichment relative to the uncut library. When the composition of the PAM sequences with .gtoreq.2 fold enrichment for the 50 nM-60 minute, 50 nM-1 minute and 0.5 nM-60 minute digests were analyzed, the consensus PAM sequence for the Blat Cas9 protein was NNNNCND (N=G, C, A or T; D=A, G or T) with a strong preference for a C at position 5 of the PAM sequence (FIG. 23). A moderate preference for an A was observed at position 7 and slight preferences for a C or T at position 4 and G, C or A over T at position 6 was also noted. Similarly to Sth1, Sth3 and Spy Cas9 proteins, the PAM specificity broadens as the Cas9-sgRNA complex concentration increases. This is most evident at position 5 where a larger proportion of PAM sequences containing an A residue support cleavage at 50 nM compared with 0.5 nM Cas9-sgRNA complexes.
[0397] To confirm the cleavage positions for the Blat Cas9 protein, we engineered the pUC18-T1-GTCCCGT-PAM plasmid containing a 20 base pair region matching the spacer T1 (SEQ ID NO: 1) followed by a PAM sequence, GTCCCGT, falling within the PAM consensus for Blat. To generate the plasmid, first the synthetic oligoduplex containing T1 and GTCCCGT PAM sequences was assembled by annealing complementary oligonucleotides GG-935 (CAAATTCTAAACGCTAAAGAGGAAGAGGACAGTCCCG (SEQ ID NO: 58) and GG-936 (AATTCGGGACTGTCCTCTTCCTCTTTAGCGTTTAGAATTTGAGCT (SEQ ID NO: 59) and ligated into pUC18 vector pre-cleaved with ScaI and EcoRI. 2.5 .mu.g of the resulting plasmid was then digested with 100 nM of the Blat Cas9-sgRNA complex in the 500 .mu.l of reaction buffer at 37.degree. C. for 60 min., purified using GeneJET PCR Purification Kit (Thermo Fisher Scientific) and electrophoresed on an agarose gel. Linear digestion products were then purified from the agarose gel using the GeneJET Gel Extraction Kit (Thermo Fisher Scientific). The cleaved region in Blat Cas9 linearized pUC18-T1-GTCCCGT-PAM plasmid was then directly sequenced with the pUC-EheD (CCGCATCAGGCGCCATTCGCC (SEQ ID NO: 60) and pUC-LguR (GCGAGGAAGCGGAAGAGCGCCC (SEQ ID NO: 61) primers. The sequence results confirmed that plasmid DNA cleavage occurred in the protospacer 3 bp away from the PAM sequence (FIG. 24) similar to that observed for Sth3 and Spy Cas9 proteins.
[0398] The NNNNCND PAM sequence identified herein, can be introduced adjacent to any polynucleotide of interest, thereby creating a target site that can be recognized by a guide RNA/Cas9 endonuclease complex described herein, wherein the guide RNA/Cas9 endonuclease system is capable of recognizing, binding to, and optionally nicking or cleaving all or part of the target sequence adjacent to the NNNNCND PAM sequence.
Example 9
Characterization of Cas9 Endonucleases and their PAM Preferences, and Cognate Guide RNAs from Diverse Organisms
[0399] The rapid in vitro methods described herein (Examples 1-8) can be used to identify and characterize Cas endonucleases from any organism and their related PAM preferences and guide RNAs elements.
[0400] Cas9 proteins of Type II-A, II-B and II-C subtypes were identified from the NCBI NR database using the PSI-BLAST program (Altschul S F, et al. (1997) Nucleic Acids Res. 25:3389-3402). A phylogenetic relationship of each Cas9 protein was visualized with CLANs software (Frickey T, Lupas A. (2004) Bioinformatics 20:3702-3704) and putative Cas9 endonucleases from different groupings were selected. Genomic DNA regions derived from non-pathogenic sources and those containing a clustered-regularly-interspace-short-palindromic repeat (CRISPR) array and a putative trans-activating CRISPR RNA (tracrRNA) coding region (defined by homology to the CRISPR repeat and termed the anti-repeat) in the vicinity of the Cas9 were chosen. In total, 11 diverse genomic DNA regions were selected for further analysis (Table 9)
[0401] A schematic of the genomic locus for each system is depicted in FIGS. 25-35. The cas9 gene open-reading-frame (ORF), CRISPR array, anti-repeat (the genomic DNA region demonstrating partial homology to the repeat consensus that indicates the location of the encoded tracrRNA) and other CRISPR-Cas genes are indicated for each system. The genomic DNA sequence and length of each cas9 gene ORF and cas9 gene translation (not including the stop codon) are referenced in Table 10 for each system. Table 10 lists the consensus sequence of the CRISPR array repeats from the genomic DNA locus of each system and the sequences of the anti-repeat for each system (as genomic DNA sequence on the same strand as the cas9 gene ORF).
[0402] As was done for the Brevibacillus laterosporus (BLAT) Type II CRISP R/Cas system (described in Example 6), the possible transcriptional directions of the putative tracrRNAs for each new system were considered by examining the secondary structures and possible termination signals present in a RNA version of the sense and anti-sense genomic DNA sequences surrounding the anti-repeat. Based on the hairpin-like secondary structures present for each system, the transcriptional direction of the tracrRNA was deduced for 10 of the 11 diverse Type II CRISPR-Cas systems. Because the anti-repeat in the tracrRNA can hybridize to the crRNA derived from the CRISPR array to form a duplexed RNA capable of guiding the Cas9 endonuclease to cleave invading DNA the transcriptional direction of the CRISPR array may also be determined based of the direction of tracrRNA transcription (since double-stranded RNA hybridizes with 5' to 3' directionality). The deduced transcriptional directions of both the tracrRNA and CRISPR array for each system are listed in Table 10 and are depicted in FIGS. 25-35. Based on the likely transcriptional direction of the tracrRNA and CRISPR array, single guide RNAs (sgRNAs) were also designed and are shown in Table 12. For the system, Sulfurospirillum sp. SCADC, where the transcriptional direction of the tracrRNA and CRISPR array could not be deduced two sgRNAs were designed (as described in Example 7 for the Blat Type II CRISPR-Cas system); one for each possible direction of tracrRNA transcription (Table 12).
[0403] Next the sgRNAs, will be complexed with the respective purified Cas9 protein and assayed for their ability to support cleavage of the 7 bp randomized PAM plasmid DNA library (as described in Example 7 for the Blat Type II CRISPR-Cas system). If the sgRNA does not support cleavage activity, new guide RNA designs (either sgRNA or duplexed crRNA and tracrRNA; in both possible transcriptional directions of the CRISPR array and anti-repeat region) will be tested for their ability to support cleavage.
[0404] Once a guide RNA that supports Cas9 cleavage has been established, the PAM specificity of each Cas9 endonuclease can be assayed (as described in Example 7 for the Blat Type II CRISPR-Cas system). After PAM preferences have been determined, the sgRNAs may be further refined for maximal activity or cellular transcription by either increasing or decreasing the tracrRNA 3' end tail length, increasing or decreasing crRNA repeat and tracrRNA anti-repeat length, modifying the 4 nt self-folding loop or altering the sequence composition.
TABLE-US-00009 TABLE 9 List of 11 organisms selected for the identification of diverse Type II CRISPR-Cas systems described herein. CRISPR-Cas Bacterial Origin Abbreviation System Subtype Isolated from Lactobacillus reuteri Mlc3 Lreu II-A Sourdough Lactobacillus rossiae DSM 15814 Lros II-A Sourdough Pediococcus pentosaceus SL4 Ppen II-A Meat Lactobacillus nodensis JCM 14932 Lnod II-A Dairy Sulfurospirillum sp. SCADC Sspe II-B Oil sands tailings pond Bifidobacterium thermophilum DSM 20210 Bthe II-C Dairy Loktanella vestfoldensis Lves II-C Lakes Ace and Pendant, Vestfold Hills, Antarctica Sphingomonas sanxanigenens NX02 Ssan II-C Isolated from soil Epilithonimonas tenax DSM 16811 Eten II-C River epilthon Sporocytophaga myxococcoides Smyx II-C From soil, cellulose decomposing organism Psychroflexus torquis ATCC 700755 Ptor II-C Prydz Bay, Antarctica
TABLE-US-00010 TABLE 10 Sequence and length of the cas9 gene ORF and cas9 gene translation from each Type II CRISPR-Cas system identified by the methods described herein. Translation of cas9 Gene Length of ORF (not cas9 Gene cas9 Gene Length of including the Translation Bacterial ORF (SEQ cas9 Gene stop codon) (No. of Amino Origin ID NO) ORF (bp) (SEQ ID NO) Acids) Lreu 70 4107 81 1368 Lros 71 4110 82 1369 Ppen 72 4041 83 1346 Lnod 73 3393 84 1130 Sspe 74 4086 85 1361 Bthe 75 3444 86 1147 Lves 76 3216 87 1071 Ssan 77 3318 88 1105 Eten 78 4200 89 1399 Smyx 79 4362 90 1453 Ptor 80 4530 91 1509
TABLE-US-00011 TABLE 11 CRISPR repeat consensus, anti-repeat (putative tracrRNA coding region) and deduced transcriptional directions of tracrRNA and CRISPR array relative to the cas9 gene ORF for 11 diverse Type II CRISPR-Cas systems. tracrRNA CRISPR Array CRISPR Transcriptional Transcriptional Repeat Anti- Direction Direction Consensus Repeat (Relative to (Relative to Bacterial (SEQ ID (SEQ ID the cas9 Gene the cas9 Gene Origin NO) NO) ORF) ORF) Lreu 92 103 Antisense Sense Lros 93 104 Antisense Sense Ppen 94 105 Antisense Sense Lnod 95 106 Sense Sense Sspe 96 107 Sense/ Sense/ Antisense Antisense Bthe 97 108 Sense Antisense Lves 98 109 Antisense Antisense Ssan 99 110 Antisense Antisense Eten 100 111 Antisense Antisense Smyx 101 112 Antisense Sense Ptor 102 113 Antisense Antisense
TABLE-US-00012 TABLE 12 Examples of sgRNAs components for each new diverse Type II CRISPR-Cas system described herein. Variable Remaining T7 Targeting Putative 3' SEQ Bacterial Transcription domain crRNA tracrRNA tracrRNA ID Origin Initiation (VT) Repeat Loop Anti-Repeat Sequence NO: Lreu GGG N.sub.20 .sub.(*.sub.) 149 N.sub.4 .sub.(**.sub.) 161 173 128 Lros GGG N.sub.20 .sub.(*.sub.) 150 N.sub.4 .sub.(**.sub.) 162 174 129 Ppen GGG N.sub.20 .sub.(*.sub.) 151 N.sub.4 .sub.(**.sub.) 163 175 130 Lnod GGG N.sub.20 .sub.(*.sub.) 152 N.sub.4 .sub.(**.sub.) 164 176 131 Sspe GGG N.sub.20 .sub.(*.sub.) 153 N.sub.4 .sub.(**.sub.) 165 177 132 (tracrRNA Sense- crRNA Sense) Sspe GGG N.sub.20 .sub.(*.sub.) 154 N.sub.4 .sub.(**.sub.) 166 178 133 (tracrRNA Antisense- crRNA Antisense) Bthe GGG N.sub.20 .sub.(*.sub.) 155 N.sub.4 .sub.(**.sub.) 167 179 134 Lves GGG N.sub.20 .sub.(*.sub.) 156 N.sub.4 .sub.(**.sub.) 168 180 135 Ssan GGG N.sub.20 .sub.(*.sub.) 157 N.sub.4 .sub.(**.sub.) 169 181 136 Eten GGG N.sub.20 .sub.(*.sub.) 158 N.sub.4 .sub.(**.sub.) 170 182 137 Smyx GGG N.sub.20 .sub.(*.sub.) 159 N.sub.4 .sub.(**.sub.) 171 183 138 Ptor GGG N.sub.20 .sub.(*.sub.) 160 N.sub.4 .sub.(**.sub.) 172 184 139
[0405] N.sub.20(*) indicates a series of 20 nucleotides as one example of a sgRNA variable targeting domain. As described herein, the variable targeting domain of a sgRNA can vary for example, but not limiting from at least 12 to 30 nucleotides. N.sub.4(**) indicates a loop of 4 nucleotides such as but not limiting to GAAA. As described herein, the length of the loop can vary from at least 3 nucleotides to 100 nucleotides.
[0406] Single guide RNAs targeting a target site in the genome of an organism can be designed by changing the targeting sequence of any one of SEQ ID NOs: 114-125 with any random nucleotide that can hybridize to any desired target sequence (such as, but not limiting to, guide RNAs as shown in SEQ ID NO: 128-139).
Example 10
PAM Specificity is not Greatly Influenced by the Type or Composition of the Guide RNA
[0407] As described in Example 3 and 4, to empirically examine the PAM preferences for Streptococcus pyogenes (Spy), Streptococcus thermophilus CRISPR3 (Sth3) and Streptococcus thermophilus CRISPR1 Cas9 proteins, two randomized PAM libraries (described in Example 1 and 4) were generated. The two libraries increased in size and complexity from 5 randomized base pairs (1,024 potential PAM combinations) to 7 randomized base pairs (16,384 potential PAM combinations). These randomized libraries were subject to digestion with purified Sth3 and Spy Cas9 proteins (5N library, Example 3) and Sth1 (Example 4, 7N library) and guide RNA containing a variable targeting domain T1 that hybridizes with, i.e., is complementary to, a sequence in the target DNA molecule (referred herein as target sequence), T1 (SEQ ID NO: 1).
[0408] To confirm that PAM specificity is independent of the type of guide RNA, duplexed crRNA: tracrRNA or single guide RNA (sgRNA), Spy, Sth3 and Sth1 Cas9 PAM preferences were examined using Cas9 sgRNA RNP complexes instead of Cas9 and crRNA:tracrRNA RNP complexes. Digestion was carried-out at a single RNP complex concentration of 0.5 nM and PAM preference analysis was performed as described herein. PAM preferences were nearly identical regardless of the type of guide RNA used; either a crRNA:tracrRNA duplex or sgRNA (U.S. patent application 62/196,535, filed Jul. 24, 2015, which is incorporated herein in its entirety by reference).
[0409] To confirm that PAM specificity is not greatly influenced by the composition of the target DNA or spacer sequence, the sequence on the opposite side of the 5 or 7 bp randomized library was targeted for cleavage with a different variable targeting domain, T2-5 for the 5 bp library or T2-7 for the 7 bp library. Spy and Sth3 Cas9 proteins preloaded with sgRNAs targeting the T2 sequence were used to interrogate the 5 bp randomized PAM library while the Sth1 Cas9-T2 sgRNA complexes were used to digest the 7 bp randomized PAM library. The library was digested with Spy, Sth3 and Sth1 Cas9 proteins preloaded with sgRNAs targeting the T2 sequence and PAM preferences were assayed as described above. The PAM preferences for all 3 Cas9 proteins were nearly identical regardless of spacer and target DNA sequence (U.S. patent application 62/196,535, filed Jul. 24, 2015).
Example 11
Identification of Extended PAM Sequences
[0410] As shown in FIG. 23 (Example 8), the PAM consensus for the Blat Cas9 protein under the 0.5 nM digest conditions was NNNNCND (N=G,C, A or T; D=A, G or T) with a strong preference for a C at position 5 of the PAM sequence. A moderate preference for an A was observed at position 7 and slight preferences for a C or T at position 4 and G, C or A over T at position 6 were also noted when closely examining FIG. 23. Similarly to Spy, Sth3 and Sth1 Cas9 proteins, the PAM specificity broadens as the Cas9-sgRNA complex concentration increases. This was most evident at position 5 where a larger proportion of PAM sequences containing an A residue support cleavage at 50 nM compared with 0.5 nM digest conditions (FIG. 23).
[0411] Since Blat Cas9 may accept any base in the first 3 positions of its PAM sequence (FIG. 23), the spacer domain T1 (and corresponding variable targeting domain in the guide RNA) was shifted by 3 nucleotides to allow PAM identification to be extended from 7 to 10 bp. The shifted T1 variable targeting domain, T1-3, was incorporated into the Blat "direct" sgRNA resulting in a sgRNA referred to as Blat sgRNA (T1-3) and PAM identification was performed as described previously for Spy, Sth3, Sth1 and Blat Cas9 proteins. PAM preference analysis revealed the PAM specificity for Blat Cas9 can be extended out to position 8 where there is a moderate preference for an additional A (U.S. patent application 62/196,535, filed Jul. 24, 2015).
[0412] To validate the PAM specificity for Blat Cas9, plasmids were engineered to contain mutations (GTCCCGAA (reference), GTCACGAA, GTCCTGAA, GTCCCGCA, GTCCCGAC, GTCCCGCC with mutations shown in bold and underlined, U.S. patent application 62/196,535, filed Jul. 24, 2015) in the most conserved residues of the PAM immediately downstream of a 20 base pair region matching the variable targeting domain T1. In vitro cleavage reactions with the various PAM sequences were initiated by mixing supercoiled plasmid DNA with pre-assembled Blat Cas9-sgRNA complex (1:1 v/v ratio) at 15.degree. C. The final reaction mixture contained 3 nM plasmid, 50 nM Cas9, 10 mMTris-HCl (pH 7.5 at 37.degree. C.), 100 mM NaCl, 1 mM DTT and 10 mM MgCl2 in a 100 .mu.l reaction volume. Aliquots were removed at timed intervals and quenched with phenol/chloroform. The aqueous phase was mixed with 3.times. loading dye solution (0.01% (w/v) bromophenol blue and 75 mM EDTA in 50% (v/v) glycerol) and reaction products analyzed by agarose gel electrophoresis. The amount of supercoiled (SC) form was evaluated by densitometric analysis of ethidium bromide stained gels using the software ImageJ. Values of reaction rate constants were obtained as described by Szczelkun et al, 2014, Proc. Natl. Acad. Sci. U.S.A 111: 9798-803). Replacement of the C nucleotide at the 5th position abolished plasmid DNA cleavage confirming its key role in Blat Cas9 PAM recognition. Replacement of A nucleotides at the 7th and 8th positions significantly reduced (43.times. and 12.times., respectively) the cleavage rate of supercoiled plasmid also indicating the importance of these nucleotides in Blat Cas9 PAM recognition.
[0413] To confirm the cleavage positions for the Blat Cas9 protein with an optimal PAM sequence, a plasmid was engineered that contained a 20 base pair region matching the variable targeting domain T1 followed by a PAM sequence, GTCCCGAA, falling within the PAM consensus for Blat Cas9, NNNNCNDD. We used direct sequencing to determine the ends of the linear DNA molecule generated by the Blat Cas9 RNP complex. The sequence results confirmed that plasmid DNA cleavage occurred in the protospacer 3 nucleotides away from the PAM sequence (similar to that observed for Spy, Sth3 and Sth1 Cas9 proteins (Garneau et al, 2010, Nature 468: 67-71; Gasiunas et al, 2012, Proc. Natl. Acad. Sci. U.S.A 109: E2579-2586; Jinek et al, 2012, Science 337: 816-21).
Example 12
In Planta Genome Editing Using Blat Cas9 and sgRNA
[0414] Following elucidation of the sgRNA and PAM preferences for Blat Cas9, maize optimized Cas9 and sgRNA expression cassettes were generated for in planta testing. The Blat cas9 gene was maize codon optimized and intron 2 of the potato ST-LSI gene was inserted to disrupt expression in E. coli and facilitate optimal splicing (Libiakova et al, 2001. Plant Cell Rep. 20: 610-615). To facilitate nuclear localization of the Blat Cas9 protein in maize cells, Simian virus 40 (SV40) monopartite and Agrobacterium tumefaciens bipartite VirD2 T-DNA border endonuclease nuclear localization signals were incorporated at the amino and carboxyl-termini of the Cas9 open reading frame, respectively (U.S. patent application 62/196,535, filed Jul. 24, 2015). To express the resulting maize optimized Blat cas9 gene in a robust constitutive manner, it was operably linked to a maize Ubiquitin promoter, 5' UTR and intron (Christensen et al, 1992, Plant Mol. Biol. 18: 675-689) and pinII terminator (An et al, 1989, Plant Cell 1: 115-122) in a plasmid DNA vector. To confer efficient sgRNA expression in maize cells, a maize U6 polymerase III promoter region isolated from Zea mays cultivar B73 residing on chromosome 8 at position 165,535,024-165,536,023 (B73 RefGen_v3) and terminator (TTTTTTTT) were isolated and operably fused to the 5' and 3' ends of a modified Blat sgRNA encoding DNA sequence. The modified Blat sgRNA contained two modifications from the sgRNA that was used in the in vitro studies (see Blat sgRNA (T1) direct; SEQ ID NO: 151), a T to G alteration at position 101 and a T to C modification at 159. The changes were introduced to remove potential premature U6 polymerase III signals in the Blat sgRNA. Alterations were introduced to have minimal impact on the secondary structure of the sgRNA compared to the version used in the in vitro studies. For a direct comparison with the Blat Cas9 sgRNA system, equivalent Cas9 and sgRNA DNA expression vectors were also prepared for the Spy Cas9 sgRNA system.
[0415] To carefully compare the mutational efficiency resulting from the imperfect non-homologous end-joing (NHEJ) repair of DNA double-strand breaks (DSBs) resulting from Spy and Blat Cas9 cleavage, protospacer identical genomic target sites were selected by identifying targets with Spy and Blat Cas9 compatible PAMs, NGGYCVAA. Since Blat and Spy Cas9 both cleave between the 3 and 4 bp upstream of their respective PAM, genomic targets will be cleaved at the exact same position allowing a tighter correlation between NHEJ mutation frequency and cleavage activity. Identical variable targeting domain sequences were selected for Blat and Spy Cas9 by capturing the 18 to 21 nt sequence immediately upstream of the PAM. To ensure optimal U6 polymerase III expression and not introduce a mismatch within the sgRNA variable targeting domain (spacer), all target sequences were selected to naturally terminate in a G at their 5' end. Targets were selected in exon 1 and 4 of the maize fertility gene Ms45 (referred to as MS45 Exon1 and MS45 Exon 4; see also U.S. Pat. No. 5,478,369 incorporated herein by reference) and within the promoter region of the maize liguleless-1 gene (refered to as LIG34 Promoter target herein; Moreno et al. 1997. Genes and Development 11:616-628).
[0416] To rapidly examine the mutational activity of Blat Cas9 with the PAM and sgRNA identified herein, Blat and the equivalent Spy Cas9 and sgRNA DNA expression vectors were independently introduced into maize Hi-II (Armstrong & Green, 1985, Planta 164: 207-214) immature embryos (IEs) by particle gun transformation similar to that described in (Ananiev et al, 2009, Chromosoma 118: 157-177). Since particle gun transformation can be highly variable, a visual marker DNA expression cassette, Ds-Red, was also co-delivered with the Cas9 and sgRNA expression vectors to aid in the selection of evenly transformed IEs. In total, 3 transformation replicates were performed on 60-90 IEs and 20-30 of the most evenly transformed IEs from each replicate were harvested 3 days after transformation. Total genomic DNA was extracted and the region surrounding the target site was PCR amplified and deep sequenced to a read depth in excess of 300,000. The resulting reads were examined for the presence of mutations at the expected site of cleavage by comparison to control experiments where only the Cas9 DNA expression cassette was transformed. Mutations arising at the expected site of cleavage for Blat Cas9 were detected with the most prevalent types of mutations being single base pair insertions or deletions. This pattern of imprecise mutagenic repair of the double-stranded DNA cut introduced by Blat Cas9 was also observed for the Spy Cas9 (U.S. patent application 62/196,535, filed Jul. 24, 2015) and at other Cas9 sites in maize (data not shown). The mutational activity for Blat Cas9 was robust at 2 of the 3 sites tested and exceeded that of the Spy Cas9 at the Ms45 Exon 4 target site by .about.30%.
[0417] In Planta Mutation Detection
[0418] The DNA region surrounding the expected site of cleavage for each Cas9-guide RNA was amplified by PCR using Phusion.RTM. High Fidelity PCR Master Mix (NEB,USA) "tailing" on the sequences necessary for amplicon-specific barcodes and IIlumina sequences through two rounds of PCR each consisting of 20 cycles. The primer pairs used in the primary PCR were primer pairs corresponding to the Ms45 exon 1, Ms45 exon 4 and Lig34 promoter regions, respectively. A set of primers universal to the products from the primary reactions, were used in the secondary PCR reaction (U.S. patent application 62/196,535, filed Jul. 24, 2015). The resulting PCR amplifications were purified with a Qiagen PCR purification spin column (Qiagen, Germany), concentration measured with a Hoechst dye-based fluorometric assay, combined in an equimolar ratio, and single read 100 nucleotide-length amplicon sequencing was performed on Illumina's MiSeq Personal Sequencer with a 5-10% (v/v) spike of PhiX control v3 (Illumina, FC-110-3001) to off-set sequence bias. Only those reads with a .gtoreq.1 nucleotide INDEL arising within the 10 nt window centered over the expected site of cleavage and not found in the negative controls were classified as mutations. Mutant reads with an identical mutation were counted and collapsed into a single read and the top 10 most prevalent mutations were visually confirmed as arising within the expected site of cleavage. The total numbers of visually confirmed mutations were then used to calculate the percentage of mutant reads based on the total number of reads of an appropriate length containing a perfect match to the barcode and forward primer.
Example 13
Simplified Construction of Randomized Protospacer-Adjacent-Motif (PAM) Libraries for Assaying Cas Endonuclease PAM Preferences
[0419] To simplify construction for randomized PAM libraries, a fully double-stranded DNA oligoduplex as described in Example 1 (oligoduplex II) containing a region of randomization immediately adjacent to a DNA target sequence may be used directly as template for Cas endonuclease digestion. This would eliminate the cloning of the oligoduplex II fragment into a plasmid DNA vector allowing randomized PAM libraries to be constructed without the downstream E. coli transformation and plasmid DNA isolation steps. PAM sequences supporting Cas endonuclease cleavage in these linearized double-stranded DNA libraries would be captured and deep sequenced as described in Examples 3, 4 and 8 for Spy, Sth3, Sth1 and Blat Cas9 proteins. To identify those sequences that have truly been cleaved by a Cas endonuclease and not just the result of adaptor ligation to the end of an un-cleaved oligoduplex, an in silico enrichment step may be applied to the resulting deep sequencing reads by selecting for only those reads that contain an appropriate sequence junction resulting for Cas endonuclease cleavage and adapter ligation. Once reads harboring a PAM sequence that supported cleavage have been identified, their nucleotide composition may be analyzed similar to that described for Spy, Sth3, Sth1 and Blat Cas9 proteins in Examples 3, 4 and 8.
Example 14
[0420] Cas Endonuclease Proto-Spacer Adjacent Motifs (PAMs) May be Assayed Directly in E. coli Cell Lysate
[0421] Cas endonuclease protein produced in E. coli may be directly (without subsequent purification steps) used to assay proto-spacer adjacent motif (PAM) recognition and single guide RNA (sgRNA) requirements upon cell lysis.
[0422] Streptococcus thermophilus CRISPR1 (Sth1) and Streptococcus thermophilus CRISPR3 (Sth3) Cas9 protein was produced in E. coli cells as described in Example 2 but without the purification steps. In brief, after cultures were grown, induced and allowed to express Cas9 protein, cell lysis was performed via sonication and cell debri was pelleted by centrifugation resulting in a cell lysate containing soluble Cas9 protein. Cas9-guide RNA complexes were assembled by combining 20 .mu.l of resulting cell lysate with RiboLock RNase Inhibitor (40 U; Thermo Fisher Scientific) and 2 .mu.g of T7 in vitro transcribed sgRNA (generated as described in Example 7) and incubated at room temperature for 15 min. To examine PAM preferences at different Cas9 concentrations, 1 .mu.g of the 7 bp randomized PAM library (Example 4) was incubated with 10 .mu.l of various dilutions (1-fold (undiluted), 10-fold and 100-fold) of cell lysate containing assembled Cas9 complexes in a 100 .mu.l reaction buffer (10 mM Tris-HCl pH 7.5 at 37.degree. C., 100 mM NaCl, 10 mM MgCl.sub.2, 1 mM DTT) so that E. coli lysate was diluted to a final concentration of either 10-fold, 100-fold or 1000-fold, respectively. Reactions mixtures were incubated for 60 min. at 37.degree., DNA end repaired with 2.5 U T4 DNA polymerase (Thermo Fisher Scientific), RNA digested with 1 .mu.l RNase A/T1 Mix (Thermo Fisher Scientific) and 3' dA added with 2.5 U of DreamTaq DNA Polymerase (Thermo Fisher Scientific). Finally, DNA was recovered using a GeneJET PCR Purification Kit (Thermo Fisher Scientific). DNA fragments resulting from cleavage by Cas9 were tagged with adapters, captured and prepared for IIlumina deep sequencing as described in Example 3 (FIG. 3). The resulting libraries were deep sequenced as described in Example 1. PAM sequences were identified from the resulting sequence data as described in Example 3 by only selecting those reads containing a perfect 12 nt sequence match flanking either side of the 7 nt PAM sequence capturing only those PAM sequences resulting from perfect Cas9-guide RNA target site recognition, cleavage and adapter ligation. The collection of resulting PAM sequences were then collapsed into like sequences, counted, and frequency of each PAM supporting cleavage calculated. To compensate for inherent bias in the initial randomized PAM libraries, the frequency of each PAM sequence was next normalized to its frequency in the starting library. Next, a PAM consensus was calculated using a position frequency matrix (PFM). This was accomplished by first aligning the collapsed PAM sequences. Then, each nucleotide (G, C, A, or T) at each position of the PAM was weighted based on the frequency of the PAM sequence with which it was associated. Finally, the total contribution of each nucleotide (G, C, A, or T) at each PAM position was summed to generate the overall probability of identifying a given nucleotide at each PAM position within the dataset.
[0423] Tables 13-18 represent the position frequency matrix (PFM) and resulting PAM consensus at each position of the 7 bp randomized PAM library for the Streptococcus thermophilus CRISPR1 (Sth1) and Streptococcus thermophilus CRISPR3 (Sth3) Cas9 proteins when assayed at different concentrations of E. coli cell lysate. The nucleotide positions of the 7 bp randomized PAM library are indicated by 1, 2, 3, 4, 5, 6, and 7 in a 5' to 3' direction with 1 being the closest to the DNA sequence involved in spacer target site recognition. The frequency of each nucleotide (G, C, A, T) at a respective position is indicated as a %. The consensus PAM preference is listed at the bottom of the table (consensus). The numbers marked with an asterisk (*) indicate the nucleotide preference(s) at each position of the protospacer adjacent motif (PAM). The percentages in the position frequency matrix (PFM) tables represent the probability of finding the corresponding nucleotide at each position of the PAM sequence and can be used to infer the strength of PAM recognition at each position.
TABLE-US-00013 TABLE 13 Position frequency matrix (PFM) and PAM consensus for Streptococcus thermophilus CRISPR1 Cas9 with Cas9 protein provided via 10 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 17.69% 14.97% 22.16% 41.47%* 9.34% 8.56% 21.79% C 27.64% 29.63% 5.67% 17.96% 28.97% 10.45% 13.89% A 26.54% 25.79% 70.38%* 16.85% 55.56%* 64.09%* 26.22%* T 28.13% 29.61% 1.79% 23.72% 6.13% 16.90% 38.10%* Consensus N N A G A A W
TABLE-US-00014 TABLE 14 Position frequency matrix (PFM) and PAM consensus for Streptococcus thermophilus CRISPR1 Cas9 with Cas9 protein provided via 100 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 19.80% 16.70% 27.37% 43.52%* 11.01% 7.87% 20.20% C 25.74% 27.47% 6.01% 16.02% 24.04% 8.77% 12.49% A 29.40% 25.80% 64.19%* 18.73% 59.60%* 69.09%* 27.66%* T 25.06% 30.03% 2.43% 21.73% 5.36% 14.27% 39.65%* Consensus N N A G A A W
TABLE-US-00015 TABLE 15 Position frequency matrix (PFM) and PAM consensus for Streptococcus thermophilus CRISPR1 Cas9 with Cas9 protein provided via 1000 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 19.72% 16.25% 24.92% 53.70%* 10.39% 3.79% 18.40% C 26.89% 30.09% 4.08% 13.55% 22.65% 3.32% 10.18% A 27.92% 26.35% 70.37%* 15.20% 64.60%* 86.15%* 33.19%* T 25.46% 27.30% 0.64% 17.55% 2.37% 6.73% 38.23%* Consensus N N A G A A W
TABLE-US-00016 TABLE 16 Position frequency matrix (PFM) and PAM consensus for Streptococcus thermophilus CRISPR3 Cas9 with Cas9 protein provided via 10 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 12.46% 49.67%* 80.76%* 21.03% 49.94%* 23.46% 21.96% C 26.60% 9.72% 5.67% 15.73% 10.22% 20.97% 24.97% A 16.71% 22.42% 8.85% 35.35% 19.75% 27.10% 25.69% T 44.23% 18.18% 4.72% 27.89% 20.10% 28.46% 27.39% Consensus N G G N G N N
TABLE-US-00017 TABLE 17 Position frequency matrix (PFM) and PAM consensus for Streptococcus thermophilus CRISPR3 Cas9 with Cas9 protein provided via 100 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 12.06% 55.16%* 82.16%* 23.38% 53.61%* 23.02% 22.39% C 28.81% 11.09% 5.10% 17.36% 10.19% 21.26% 24.06% A 22.84% 17.33% 9.02% 31.55% 18.80% 25.87% 25.64% T 36.28% 16.42% 3.72% 27.71% 17.40% 29.84% 27.91% Consensus N G G N G N N
TABLE-US-00018 TABLE 18 Position frequency matrix (PFM) and PAM consensus for Streptococcus thermophilus CRISPR3 Cas9 with Cas9 protein provided via 1000 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 12.26% 63.66%* 89.19%* 27.07% 54.77%* 26.19% 23.09% C 30.31% 7.86% 2.78% 17.23% 9.70% 19.39% 22.85% A 21.26% 15.31% 6.18% 29.16% 17.45% 26.56% 26.21% T 36.17% 13.17% 1.86% 26.55% 18.08% 27.87% 27.86% Consensus N G G N G N N
[0424] As shown in Tables 13-18, all lysate dilutions yielded the canonical PAM preferences for Sth1 and Sth3 Cas9 proteins, NNAGAAW and NGGNG, respectively. Similar to the results with purified protein in Examples 3, 4 and 8, higher concentrations of lysate and consequentially Cas9 protein resulted in a relaxation of PAM specificity. This was most notable for the Sth3 Cas9 protein at PAM position 2 where the preference for a G residue is reduced from approximately 64% in the PFM in the 1000-fold dilution (final concentration) reaction to around 50% in the 10-fold dilution (final concentration) experiment (Tables 16-18). For Sth1 Cas9 protein, PAM positions 4, 5 and 6 were most particularly affected by different concentrations of Cas9 protein in the lysate dilution experiments.
[0425] This data indicates that the in vitro PAM library assay described herein obtained the same results for the PAM preferences for Sth1 and Sth3 Cas9 proteins when compared to assays where the Sth1 and Sth3 Cas9 proteins are stably expressed (in-vivo expressed). Hence, the in vitro PAM library assay described herein, or derivations of it, may be used to assay PAM specificity from any Cas endonuclease using unpurified Cas protein coming directly from E. coli lysate. Additionally by diluting E. coli lysate containing Cas9 protein, the in vitro PAM library assay permits the measurement of PAM specificity to be examined as a function of Cas endonuclease concentration as is evident by the apparent broadening in PAM specificity as E. coli lysate containing Cas9 protein was increased.
Example 15
Cas Endonuclease Proto-Spacer Adjacent Motifs (PAMs) May be Assayed Directly with In Vitro Translated Protein
[0426] Cas endonuclease protein produced by in vitro translation may be used to directly (without subsequent purification steps) assay proto-spacer adjacent motifs (PAM) and single guide RNA (sgRNA) requirements.
[0427] The Streptococcus pyogenes (Spy) cas9 gene was codon optimized for expression in eukaryotes (maize) with standard methods known in the art and operably linked to the in vitro translation (IVT) vector pT7CFE1-NHIS-GST-CHA (Thermo Fisher Scientific). To eliminate expression of the HA tag, a stop codon was included between the Spy cas9 gene and C-terminal tag. The resulting plasmid was purified by phenol:chloroform extraction to remove residual RNases and further purified by precipitation with 2 volumes of ethanol in the presence of sodium acetate. Next, Spy protein was produced in vitro using a 1-Step Human Coupled IVT Kit (Thermo Fisher Scientific) per the manufacturer's instruction allowing the reaction to proceed overnight at 30.degree. C. Following the incubation, the reactions were centrifuged at 10,000 rpm for 5 min. 20 .mu.l of supernatant containing soluble Cas9 protein was mixed with 2 .mu.g of T7 in vitro transcribed sgRNA (generated as described in Example 7) and incubated for 15 min. at room temperature. To examine PAM preferences at different Cas9 concentrations, 1 .mu.g of the 7 bp randomized PAM library (Example 4) was incubated with 10 .mu.l of various dilutions (1-fold (undiluted), 10-fold and 100-fold) of in vitro translation mixtures containing assembled Cas9 complexes in a 100 .mu.l reaction buffer (10 mM Tris-HCl pH 7.5 at 37.degree. C., 100 mM NaCl, 10 mM MgCl.sub.2, 1 mM DTT) so that IVT supernatant was diluted to a final concentration of either 10-fold, 100-fold or 1000-fold. Reactions mixtures were incubated for 60 min at 37.degree., DNA end repaired with 2.5 U T4 DNA polymerase (Thermo Fisher Scientific), RNA digested with 1 .mu.l RNase A/T1 Mix (Thermo Fisher Scientific) and 3' dA added with 2.5 U of DreamTaq DNA Polymerase (Thermo Fisher Scientific). Finally, DNA was recovered using a GeneJET PCR Purification Kit (Thermo Fisher Scientific). PAM sequences supporting cleavage were captured by adapter ligation and enriched for as described in Example 3 (FIG. 3). The resulting libraries were deep sequenced as described in Example 1. PAM sequences were identified from the resulting sequence data as described in Example 3 by only selecting those reads containing a perfect 12 nt sequence match flanking either side of the 5 or 7 nt PAM sequence capturing only those PAM sequences resulting from perfect Cas9-guide RNA target site recognition, cleavage and adapter ligation. To compensate for inherent bias in the initial randomized PAM library, the frequency of each PAM sequence was normalized to its frequency in the starting library and a PAM consensus was then calculated with a position frequency matrix (PFM) as described in Example 14.
[0428] Tables 19-21 represent the position frequency matrix (PFM) and resulting PAM consensus at each position of the 7 bp randomized PAM library for the Streptococcus pyogenes Cas9 protein when assayed at different concentrations of in vitro translated (IVT) supernatant. The nucleotide positions of the 7 bp randomized PAM library are indicated by 1, 2, 3, 4, 5, 6, and 7 in a 5' to 3' direction with 1 being the closest to the DNA sequence involved in spacer target site recognition. The frequency of each nucleotide (G, C, A, T) at a respective position is indicated as a %. The consensus PAM preference is listed at the bottom of the table (consensus). The numbers marked with an asterisk (*) indicate the nucleotide preference(s) at each position of the protospacer adjacent motif (PAM). The percentages in the position frequency matrix (PFM) tables represent the probability of finding the corresponding nucleotide at each position of the PAM sequence and can be used to infer the strength of PAM recognition at each position.
TABLE-US-00019 TABLE 19 Position frequency matrix (PFM) and PAM consensus for Streptococcus pyogenes Cas9 with Cas9 protein provided via 10 fold dilution of in-vitro translated solution (IVT). 1 2 3 4 5 6 7 G 24.18% 53.04%* 72.63%* 19.30% 14.19% 19.97% 23.65% C 25.97% 7.16% 8.52% 24.26% 25.67% 28.52% 27.44% A 25.21% 28.71% 14.69% 22.57% 23.80% 19.66% 20.39% T 24.64% 11.09% 4.16% 33.87% 36.34% 31.85% 28.52% Consensus N G G N N N N
TABLE-US-00020 TABLE 20 Position frequency matrix (PFM) and PAM consensus for Streptococcus pyogenes Cas9 with Cas9 protein provided via 100 fold dilution of in-vitro translated solution (IVT). 1 2 3 4 5 6 7 G 23.84% 52.07%* 78.60%* 21.17% 14.72% 19.66% 22.39% C 24.16% 6.26% 4.34% 21.69% 23.72% 28.60% 27.09% A 26.64% 34.55% 14.85% 25.33% 25.90% 20.48% 21.29% T 25.36% 7.12% 2.21% 31.82% 35.66% 31.26% 29.23% Consensus N G G N N N N
TABLE-US-00021 TABLE 21 Position frequency matrix (PFM) and PAM consensus for Streptococcus pyogenes Cas9 with Cas9 protein provided via 1000 fold dilution of in-vitro translated solution (IVT). 1 2 3 4 5 6 7 G 23.39% 81.14%* 95.35%* 27.51% 15.79% 19.98% 22.92% C 22.34% 2.54% 0.80% 14.69% 23.08% 26.85% 25.30% A 29.08% 12.52% 3.07% 26.65% 25.51% 22.87% 22.57% T 25.19% 3.80% 0.78% 31.15% 35.63% 30.29% 29.22% Consensus N G G N N N N
[0429] As illustrated in Tables 19-21, the PAM requirement preferences reported for the Spy Cas9 protein (NGG) may be recapitulated under all IVT dilutions. Similar to the results with purified protein in Examples 3, 4 and 8, higher concentrations of IVT supernatant and consequentially Cas9 protein resulted in a broadening of PAM specificity. This was most notable for Spy Cas9 at PAM position 2 where the frequency for an uncanonical A residue increases from approximately 13% in the PFM with the 1000-fold dilution (final concentration) reaction to around 29% in the 10-fold dilution (final concentration) experiment.
[0430] This data indicates that the in vitro translation (IVT) assay described herein obtained the same results for the PAM preferences for Spy Cas9 protein when compared to assays where the Spy Cas9 protein is stably expressed (in-vivo expressed). Hence, the in vitro translation (IVT) assay described herein, or derivations of it, may be used to assay PAM specificity from any Cas endonuclease. Additionally by diluting IVT products containing Cas9 protein, our assay permits the measurement of PAM specificity to be examined as a function of Cas endonuclease concentration as evident by the apparent broadening in PAM specificity as IVT supernatant containing Cas9 protein was increased.
Example 16
Guide RNA and PAM Requirements for Novel Cas Endonucleases
[0431] The single guide RNA (sgRNA) and PAM requirements of the Cas9 endonucleases from Lactobacillus reuteri MIc3 (Lreu), Lactobacillus nodensis JCM 14932 (Lnod), Sulfurospirillum sp. SCADC (Sspe), Bifidobacterium thermophilum DSM 20210 (Bthe), Loktanella vestfoldensis (Lves), Epilithonimonas tenax DSM 16811 (Eten) and Sporocytophaga myxococcoides (Smyx) (Example 9) were determined with the methods described herein.
[0432] If purified protein could not be easily obtained as described in Example 2, Cas9 protein from E. coli cell lysate as described in Example 14 or in vitro translated (IVT) Cas9 protein as described in Example 15 was utilized. Once a source of Cas9 protein was established, 1 .mu.g of the 7 bp randomized PAM plasmid DNA library (Example 4) was subject to Cas9-guide RNA digestion at various concentrations of either purified protein, lysate, or IVT protein. DNA fragments resulting from cleavage by Cas9 were ligated to adapters, captured and prepared for IIlumina deep sequencing as described in Example 3 (FIG. 3). The resulting libraries were deep sequenced as described in Example 1. Since the position of cleavage within target sites for novel Cas9 proteins is unknown, reads were 1.sup.st examined for the most predominant cleavage location by examining the junction resulting from cleavage and adapter ligation. After properly defining the position of cleavage, PAM sequences were identified from the resulting sequence data as described in Example 3 by only selecting those reads containing a perfect 12 nt sequence match flanking either side of the 5 or 7 nt PAM sequence. To compensate for inherent bias in the initial randomized PAM library, the frequency of each PAM sequence was normalized to its frequency in the starting library and a PAM consensus was then calculated with a position frequency matrix (PFM) as described in Example 14. To obtain the most accurate read-out on PAM specificity and avoid conditions that are conducive to promiscuous PAM recognition (Examples 3, 4, 8, 14 and 15), the lowest concentration of Cas9 (purified, E. coli lysate or IVT supernatant) that supported cleavage was used to ascertain the PAM recognition of each Cas9 protein.
[0433] Tables 22-28 represent the position frequency matrix (PFM) and resulting PAM consensus at each position of the 7 bp randomized PAM library for several previously uncharacterized Cas9 proteins. Results derived from the lowest concentration of Cas9 coming from either purified, E. coli lysate or in vitro translation (IVT) supernatant that supported cleavage are shown. The nucleotide positions of the 7 bp randomized PAM library are indicated by 1, 2, 3, 4, 5, 6, and 7 in a 5' to 3' direction with 1 being the closest to the DNA sequence involved in spacer target site recognition. The frequency of each nucleotide (G, C, A, T) at a respective position is indicated as a %. The consensus PAM preference is listed at the bottom of the table (consensus). The numbers marked with an asterisk (*) indicate the nucleotide preference(s) at each position of the protospacer adjacent motif (PAM). The percentages in the position frequency matrix (PFM) tables represent the probability of finding the corresponding nucleotide at each position of the PAM sequence and can be used to infer the strength of PAM recognition at each position.
TABLE-US-00022 TABLE 22 Position frequency matrix (PFM) and PAM consensus for Lactobacillus reuteri Cas9 when purified Cas9 protein was used (0.5 nM Cas9-guide RNA complex and 60 minute digestion time). 1 2 3 4 5 6 7 G 15.57% 83.27%* 98.90%* 31.64% 39.04%* 25.51% 15.86% C 15.96% 2.44% 0.12% 17.94% 24.13% 26.77% 34.32% A 17.74% 11.81% 0.66% 14.84% 11.30% 22.13% 18.37% T 50.73% 2.48% 0.32% 35.58% 25.53% 25.58% 31.44% Consensus N (T > V) G G N N (G > H) N N
TABLE-US-00023 TABLE 23 Position frequency matrix (PFM) and PAM consensus for Lactobacillus nodensis Cas9 when purified Cas9 protein was used (50 nM Cas9-guide RNA complex and 60 minute digestion time). 1 2 3 4 5 6 7 G 21.47% 13.95% 2.62% 7.92% 4.07% 5.67% 24.14% C 25.74% 23.76% 2.07% 1.53% 1.68% 1.29% 16.67% A 22.41% 19.73% 94.31%* 89.34%* 93.77%* 91.48%* 33.13% T 30.38% 42.56%* 0.99% 1.22% 0.48% 1.55% 26.07% Consensus N N (T > V) A A A A N
TABLE-US-00024 TABLE 24 Position frequency matrix (PFM) and PAM consensus for Sulfurospirillum sp. SCADC Cas9 with Cas9 protein provided via 1000 fold dilution of in vitro translated solution (IVT). 1 2 3 4 5 6 7 G 16.26% 97.32%* 97.67%* 18.52% 22.18% 18.86% 23.20% C 24.43% 0.95% 0.85% 20.37% 20.90% 25.19% 22.14% A 35.19% 1.11% 0.74% 31.97% 22.61% 26.12% 23.94% T 24.13% 0.61% 0.74% 29.13% 34.31% 29.82% 30.72% Consensus N G G N N N N
TABLE-US-00025 TABLE 25 Position frequency matrix (PFM) and PAM consensus for Bifidobacterium thermophilum Cas9 when purified Cas9 protein was used (0.5 nM Cas9-guide RNA complex and 60 minute digestion time). 1 2 3 4 5 6 7 G 18.93% 16.16% 20.28% 0.10% 0.03% 2.53% 3.19% C 34.69% 31.11% 27.80% 99.55%* 99.05%* 5.34% 47.56%* A 23.13% 28.52% 28.76% 0.13% 0.40% 91.44%* 1.17% T 23.24% 24.20% 23.17% 0.21% 0.52% 0.69% 48.08%* Consensus N N N C C A Y
TABLE-US-00026 TABLE 26 Position frequency matrix (PFM) and PAM consensus for Loktanella vestfoldensis Cas9 with Cas9 protein provided via 1000 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 21.74% 62.30%* 51.21%* 13.71% 17.79% 32.03%* 23.70% C 29.99% 8.00% 5.94% 10.17% 5.73% 14.72% 23.82% A 16.37% 21.66% 37.33%* 63.65%* 64.49%* 13.01% 25.97% T 31.89% 8.03% 5.51% 12.47% 11.99% 40.24%* 26.51% Consensus N G R (G > A) A A K N
TABLE-US-00027 TABLE 27 Position frequency matrix (PFM) and PAM consensus for Epilithonimonas tenax Cas9 with Cas9 protein provided via 10 fold dilution of E. coli cell lysate. 1 2 3 4 5 6 7 G 30.87%* 25.83% 39.60%* 18.03% 14.19% 87.26%* 91.40%* C 30.34%* 7.27% 3.14% 7.13% 11.68% 2.31% 2.27% A 15.84% 63.47%* 54.64%* 71.53%* 31.83%* 3.18% 2.88% T 22.95% 3.43% 2.61% 3.30% 42.29%* 7.25% 3.45% Consensus N (S > W) A R A N (W > S) G G
TABLE-US-00028 TABLE 28 Position frequency matrix (PFM) and PAM consensus for Sporocytophaga myxococcoides Cas9 when purified Cas9 protein was used (50 nM Cas9-guide RNA complex and 60 minute digestion time). 1 2 3 4 5 6 7 G 10.48% 19.15% 2.54% 4.72% 1.02% 7.00% 23.48% C 26.01% 14.45% 0.56% 23.74% 0.80% 3.23% 17.02% A 19.94% 59.05%* 96.61%* 7.74% 97.97%* 79.56%* 28.21% T 43.58% 7.35% 0.29% 63.80%* 0.21% 10.21% 31.28% Consensus N (T > V) A A T A A N
TABLE-US-00029 TABLE 29 Summary of sgRNA and PAM requirement for novel Cas endonucleases. PAM sgRNA Bacterial Origin Abbreviation consensus SEQ ID NO: Lactobacillus reuteri Mlc3 Lreu Table 22 114 Lactobacillus nodensis JCM 14932 Lnod Table 23 117 Sulfurospirillum sp. SCADC Sspe Table 24 119 Bifidobacterium thermophilum DSM 20210 Bthe Table 25 120 Loktanella vestfoldensis Lves Table 26 121 Epilithonimonas tenax DSM 16811 Eten Table 27 123 Sporocytophaga myxococcoides Smyx Table 28 124
[0434] Among the Cas9 proteins examined, both the length and composition of PAM recognition was diverse. Two of the Cas9 proteins, Lreu and Sspe (Tables 22-23), exhibited PAM recognition similar to the Streptococcus pyogenes (Spy) Cas9 protein which predominantly recognizes a NGG PAM while others exhibited very C-rich (Bthe, Table 25) or A-rich (Lnod and Smyx; Tables 23 and 28) PAM recognition. Additionally, a couple of the Cas9 proteins, Eten and Lves (Tables 26 and 27), yielded characteristics of both G-rich and A-rich PAM recognition.
[0435] Unlike the diversity observed for PAM recognition, the position of target site cleavage did not differ greatly and was determined to be between the 3.sup.rd and 4th bp upstream (5 prime) of the PAM for all Cas9 proteins except for one, the Cas9 protein from Sulfurospirillum sp. SCADC. Interestingly, the predominant cleavage location by examining the junction resulting from cleavage and adapter ligation was around the 7.sup.th bp upstream (5 prime) of the PAM sequence.
[0436] Taken together, these data further suggest that the methods described herein can be used to characterize novel Cas endonuclease PAM and guide RNA requirements.
Example 17
In Planta Genome Editing with Novel Cas9 Endonucleases
[0437] After determining the proto-spacer adjacent motif (PAM) and guide RNA requirement as described herein, Cas9 proteins with novel PAM recognition were selected and tested for their ability to cleave and mutagenize maize chromosomal DNA as described in Example 12.
[0438] To expand the number and diversity of sites available for genome editing, Cas9 proteins with diverse PAM recognition were selected for evaluation in corn by preferentially choosing systems with either A, T or C-rich PAM recognition to best complement the G-rich PAM of the Streptococcus pyogenes (Spy) Cas9 protein. Once systems were selected, DNA target sites adjacent to the appropriate PAM sequence were chosen and maize optimized cas9 gene and single guide RNA (sgRNA) expression vectors were constructed and delivered into maize immature embryos as described in Example 12. Embryos were harvested two days after transformation and chromosomal DNA was analyzed for the presence of mutations resulting from DNA target site cleavage and repair as described in Example 12. The frequency of mutations identified at each target site for each Cas9 is listed in Table 30.
[0439] Interestingly, the Bifidobacterium thermophilum (Bthe) Cas9 protein failed to effectively mutagenize its target sites. However when different spacer lengths were tested for Bthe, the frequency of mutagenesis improved dramatically with a spacer length around 25 nt being the most optimal (FIG. 36). Since the minimal spacer length for the Streptococcus pyogenes (Spy) Cas9 sgRNA is approximately 17 nt in length, it seems that the sgRNA spacer DNA target interactions for Bthe Cas9 may provide enhanced specificity relative to the Spy Cas9 protein.
TABLE-US-00030 TABLE 30 Maize chromosomal target DNA mutation frequencies two days after transformation by particle gun. sgRNA DNA Target Spacer Mutation Origin of cas9 gene Location Length Frequency Bifidobacterium thermophilum Chr1: 51.81 cM 25 0.29% DSM 20210 Chr9: 119.15 cM 25 0.05% Lactobacillus nodensis JCM Chr1: 51.81 cM 21 0.06% 14932 Chr9: 119.15 cM 22 0.28%
[0440] Taken together, these results indicate that the methods described herein to characterize Cas endonuclease PAM recognition and guide RNA requirements are robust. Ultimately, allowing new Cas endonuclease systems to be characterized for genome editing applications.
Sequence CWU 1
1
184120DNAArtificial Sequencesynthesized T1 sequence 1cgctaaagag
gaagaggaca 20247DNAArtificial Sequencesynthesized
GG-821Nmisc_feature(20)..(24)n is a, c, g, or t 2tgaccatgat
tacgaattcn nnnntgtcct cttcctcttt agcgagc 47344DNAArtificial
Sequencesynthesized GG-820 3aaggatcccc gggtaccgag ctgctcgcta
aagaggaaga ggac 44422DNAArtificial Sequencesynthesized sequence
TK-119 4gagctcgcta aagaggaaga gg 22522DNAArtificial
Sequencesynthesized sequence pUC-dir 5gccagggttt tcccagtcac ga
22659DNAArtificial Sequencesynthesized sequence JKYS800.1
6ctacactctt tccctacacg acgctcttcc gatctaagtg agctcgctaa agaggaaga
59753DNAArtificial Sequencesynthesized sequence JKYS803 7caagcagaag
acggcatacg agctcttccg atctgaattc gagctcggta cct 53843DNAArtificial
Sequencesynthesized sequence Universal Forward 8aatgatacgg
cgaccaccga gatctacact ctttccctac acg 43918DNAArtificial
Sequencesynthesized sequence Universal Reverse 9caagcagaag acggcata
181034DNAArtificial Sequencesynthesized sequence Sth1-dir
10acgtctcaca tgactaagcc atactcaatt ggac 341127DNAArtificial
Sequencesynthesized sequence Sth1-rev 11actcgagacc ctctcctagt
ttggcaa 271226DNAArtificial Sequencesynthesized sequence Sth3-dir
12ggggggtctc acatgagtga cttagt 261330DNAArtificial
Sequencesynthesized sequence Sth3-rev 13aattactcga gaaaatctag
cttaggctta 301438DNAArtificial Sequencesynthesized sequence Spy-dir
14aaggtctccc atggataaga aatactcaat aggcttag 381532DNAArtificial
Sequencesynthesized sequence Spy-rev 15ttctcgaggt cacctcctag
ctgactcaaa tc 321642RNAStreptococcus thermophilus 16cgcuaaagag
gaagaggaca guuuuagagc uguguuguuu cg 421778RNAStreptococcus
thermophilus 17gggcgaaaca acacagcgag uuaaaauaag gcuuaguccg
uacucaacuu gaaaaggugg 60caccgauucg guguuuuu 781842RNAStreptococcus
pyogenes 18cgcuaaagag gaagaggaca guuuuagagc uaugcuguuu ug
421978RNAStreptococcus pyogenes 19gggaaacagc auagcaaguu aaaauaaggc
uaguccguua ucaacuugaa aaaguggcac 60cgagucggug cuuuuuuu
782031DNAArtificial Sequencesynthesized sequence TK-117
20cggcattcct gctgaaccgc tcttccgatc t 312130DNAArtificial
Sequencesynthesized sequence TK-111 21gatcggaaga gcggttcagc
aggaatgccg 302256DNAArtificial Sequencesynthesized sequence
JKYS807.1 22ctacactctt tccctacacg acgctcttcc gatctaaggc ggcattcctg
ctgaac 562356DNAArtificial Sequencesynthesized sequence JKYS807.2
23ctacactctt tccctacacg acgctcttcc gatctttccc ggcattcctg ctgaac
562456DNAArtificial Sequencesynthesized sequence JKYS807.3
24ctacactctt tccctacacg acgctcttcc gatctggaac ggcattcctg ctgaac
562556DNAArtificial Sequencesynthesized sequence JKYS807.4
25ctacactctt tccctacacg acgctcttcc gatctccttc ggcattcctg ctgaac
5626123RNAArtificial Sequencesynthesized sequence Sth3 sgRNA
26gggcgcuaaa gaggaagagg acaguuuuag agcuguguug uuucgguuaa aacaacacag
60cgaguuaaaa uaaggcuuag uccguacuca acuugaaaag guggcaccga uucgguguuu
120uuu 12327105RNAArtificial Sequencesynthesized sequence Spy sgRNA
27gggcgcuaaa gaggaagagg acaguuuuag agcuagaaau agcaaguuaa aauaaggcua
60guccguuauc aacuugaaaa aguggcaccg agucggugcu uuuuu
1052859DNAArtificial Sequencesynthesized sequence
GG-940-Gmisc_feature(28)..(29)n is a, c, g, or
tmisc_feature(31)..(34)n is a, c, g, or t 28gtgcacgccg gcgacgttgg
gtcaactnng nnnntgtcct cttcctcttt agcgtttag 592959DNAArtificial
Sequencesynthesized sequence GG-940-Cmisc_feature(28)..(29)n is a,
c, g, or tmisc_feature(31)..(34)n is a, c, g, or t 29gtgcacgccg
gcgacgttgg gtcaactnnc nnnntgtcct cttcctcttt agcgtttag
593059DNAArtificial Sequencesynthesized sequence
GG-940-Amisc_feature(28)..(29)n is a, c, g, or
tmisc_feature(31)..(34)n is a, c, g, or t 30gtgcacgccg gcgacgttgg
gtcaactnna nnnntgtcct cttcctcttt agcgtttag 593159DNAArtificial
Sequencesynthesized sequence GG-940-Tmisc_feature(28)..(29)n is a,
c, g, or tmisc_feature(31)..(34)n is a, c, g, or t 31gtgcacgccg
gcgacgttgg gtcaactnnt nnnntgtcct cttcctcttt agcgtttag
593249DNAArtificial Sequencesynthesized sequence JKYS812
32caagcagaag acggcatacg agctcttccg atctcggcga cgttgggtc
493342RNAStreptococcus thermophilus 33cgcuaaagag gaagaggaca
guuuuuguac ucucaagauu ua 423480RNAStreptococcus thermophilus
34ggguaaaucu ugcagaagcu acaaagauaa ggcuucaugc cgaaaucaac acccugucau
60uuuauggcag gguguuuucg 80351388PRTStreptococcus thermophilus 35Met
Thr Lys Pro Tyr Ser Ile Gly Leu Asp Ile Gly Thr Asn Ser Val1 5 10
15Gly Trp Ala Val Ile Thr Asp Asn Tyr Lys Val Pro Ser Lys Lys Met
20 25 30Lys Val Leu Gly Asn Thr Ser Lys Lys Tyr Ile Lys Lys Asn Leu
Leu 35 40 45Gly Val Leu Leu Phe Asp Ser Gly Ile Thr Ala Glu Gly Arg
Arg Leu 50 55 60Lys Arg Thr Ala Arg Arg Arg Tyr Thr Arg Arg Arg Asn
Arg Ile Leu65 70 75 80Tyr Leu Gln Glu Ile Phe Ser Thr Glu Met Ala
Thr Leu Asp Asp Ala 85 90 95Phe Phe Gln Arg Leu Asp Asp Ser Phe Leu
Val Pro Asp Asp Lys Arg 100 105 110Asp Ser Lys Tyr Pro Ile Phe Gly
Asn Leu Val Glu Glu Lys Val Tyr 115 120 125His Asp Glu Phe Pro Thr
Ile Tyr His Leu Arg Lys Tyr Leu Ala Asp 130 135 140Ser Thr Lys Lys
Ala Asp Leu Arg Leu Val Tyr Leu Ala Leu Ala His145 150 155 160Met
Ile Lys Tyr Arg Gly His Phe Leu Ile Glu Gly Glu Phe Asn Ser 165 170
175Lys Asn Asn Asp Ile Gln Lys Asn Phe Gln Asp Phe Leu Asp Thr Tyr
180 185 190Asn Ala Ile Phe Glu Ser Asp Leu Ser Leu Glu Asn Ser Lys
Gln Leu 195 200 205Glu Glu Ile Val Lys Asp Lys Ile Ser Lys Leu Glu
Lys Lys Asp Arg 210 215 220Ile Leu Lys Leu Phe Pro Gly Glu Lys Asn
Ser Gly Ile Phe Ser Glu225 230 235 240Phe Leu Lys Leu Ile Val Gly
Asn Gln Ala Asp Phe Arg Lys Cys Phe 245 250 255Asn Leu Asp Glu Lys
Ala Ser Leu His Phe Ser Lys Glu Ser Tyr Asp 260 265 270Glu Asp Leu
Glu Thr Leu Leu Gly Tyr Ile Gly Asp Asp Tyr Ser Asp 275 280 285Val
Phe Leu Lys Ala Lys Lys Leu Tyr Asp Ala Ile Leu Leu Ser Gly 290 295
300Phe Leu Thr Val Thr Asp Asn Glu Thr Glu Ala Pro Leu Ser Ser
Ala305 310 315 320Met Ile Lys Arg Tyr Asn Glu His Lys Glu Asp Leu
Ala Leu Leu Lys 325 330 335Glu Tyr Ile Arg Asn Ile Ser Leu Lys Thr
Tyr Asn Glu Val Phe Lys 340 345 350Asp Asp Thr Lys Asn Gly Tyr Ala
Gly Tyr Ile Asp Gly Lys Thr Asn 355 360 365Gln Glu Asp Phe Tyr Val
Tyr Leu Lys Asn Leu Leu Ala Glu Phe Glu 370 375 380Gly Ala Asp Tyr
Phe Leu Glu Lys Ile Asp Arg Glu Asp Phe Leu Arg385 390 395 400Lys
Gln Arg Thr Phe Asp Asn Gly Ser Ile Pro Tyr Gln Ile His Leu 405 410
415Gln Glu Met Arg Ala Ile Leu Asp Lys Gln Ala Lys Phe Tyr Pro Phe
420 425 430Leu Ala Lys Asn Lys Glu Arg Ile Glu Lys Ile Leu Thr Phe
Arg Ile 435 440 445Pro Tyr Tyr Val Gly Pro Leu Ala Arg Gly Asn Ser
Asp Phe Ala Trp 450 455 460Ser Ile Arg Lys Arg Asn Glu Lys Ile Thr
Pro Trp Asn Phe Glu Asp465 470 475 480Val Ile Asp Lys Glu Ser Ser
Ala Glu Ala Phe Ile Asn Arg Met Thr 485 490 495Ser Phe Asp Leu Tyr
Leu Pro Glu Glu Lys Val Leu Pro Lys His Ser 500 505 510Leu Leu Tyr
Glu Thr Phe Asn Val Tyr Asn Glu Leu Thr Lys Val Arg 515 520 525Phe
Ile Ala Glu Ser Met Arg Asp Tyr Gln Phe Leu Asp Ser Lys Gln 530 535
540Lys Lys Asp Ile Val Arg Leu Tyr Phe Lys Asp Lys Arg Lys Val
Thr545 550 555 560Asp Lys Asp Ile Ile Glu Tyr Leu His Ala Ile Tyr
Gly Tyr Asp Gly 565 570 575Ile Glu Leu Lys Gly Ile Glu Lys Gln Phe
Asn Ser Ser Leu Ser Thr 580 585 590Tyr His Asp Leu Leu Asn Ile Ile
Asn Asp Lys Glu Phe Leu Asp Asp 595 600 605Ser Ser Asn Glu Ala Ile
Ile Glu Glu Ile Ile His Thr Leu Thr Ile 610 615 620Phe Glu Asp Arg
Glu Met Ile Lys Gln Arg Leu Ser Lys Phe Glu Asn625 630 635 640Ile
Phe Asp Lys Ser Val Leu Lys Lys Leu Ser Arg Arg His Tyr Thr 645 650
655Gly Trp Gly Lys Leu Ser Ala Lys Leu Ile Asn Gly Ile Arg Asp Glu
660 665 670Lys Ser Gly Asn Thr Ile Leu Asp Tyr Leu Ile Asp Asp Gly
Ile Ser 675 680 685Asn Arg Asn Phe Met Gln Leu Ile His Asp Asp Ala
Leu Ser Phe Lys 690 695 700Lys Lys Ile Gln Lys Ala Gln Ile Ile Gly
Asp Glu Asp Lys Gly Asn705 710 715 720Ile Lys Glu Val Val Lys Ser
Leu Pro Gly Ser Pro Ala Ile Lys Lys 725 730 735Gly Ile Leu Gln Ser
Ile Lys Ile Val Asp Glu Leu Val Lys Val Met 740 745 750Gly Gly Arg
Lys Pro Glu Ser Ile Val Val Glu Met Ala Arg Glu Asn 755 760 765Gln
Tyr Thr Asn Gln Gly Lys Ser Asn Ser Gln Gln Arg Leu Lys Arg 770 775
780Leu Glu Lys Ser Leu Lys Glu Leu Gly Ser Lys Ile Leu Lys Glu
Asn785 790 795 800Ile Pro Ala Lys Leu Ser Lys Ile Asp Asn Asn Ala
Leu Gln Asn Asp 805 810 815Arg Leu Tyr Leu Tyr Tyr Leu Gln Asn Gly
Lys Asp Met Tyr Thr Gly 820 825 830Asp Asp Leu Asp Ile Asp Arg Leu
Ser Asn Tyr Asp Ile Asp His Ile 835 840 845Ile Pro Gln Ala Phe Leu
Lys Asp Asn Ser Ile Asp Asn Lys Val Leu 850 855 860Val Ser Ser Ala
Ser Asn Arg Gly Lys Ser Asp Asp Phe Pro Ser Leu865 870 875 880Glu
Val Val Lys Lys Arg Lys Thr Phe Trp Tyr Gln Leu Leu Lys Ser 885 890
895Lys Leu Ile Ser Gln Arg Lys Phe Asp Asn Leu Thr Lys Ala Glu Arg
900 905 910Gly Gly Leu Leu Pro Glu Asp Lys Ala Gly Phe Ile Gln Arg
Gln Leu 915 920 925Val Glu Thr Arg Gln Ile Thr Lys His Val Ala Arg
Leu Leu Asp Glu 930 935 940Lys Phe Asn Asn Lys Lys Asp Glu Asn Asn
Arg Ala Val Arg Thr Val945 950 955 960Lys Ile Ile Thr Leu Lys Ser
Thr Leu Val Ser Gln Phe Arg Lys Asp 965 970 975Phe Glu Leu Tyr Lys
Val Arg Glu Ile Asn Asp Phe His His Ala His 980 985 990Asp Ala Tyr
Leu Asn Ala Val Ile Ala Ser Ala Leu Leu Lys Lys Tyr 995 1000
1005Pro Lys Leu Glu Pro Glu Phe Val Tyr Gly Asp Tyr Pro Lys Tyr
1010 1015 1020Asn Ser Phe Arg Glu Arg Lys Ser Ala Thr Glu Lys Val
Tyr Phe 1025 1030 1035Tyr Ser Asn Ile Met Asn Ile Phe Lys Lys Ser
Ile Ser Leu Ala 1040 1045 1050Asp Gly Arg Val Ile Glu Arg Pro Leu
Ile Glu Val Asn Glu Glu 1055 1060 1065Thr Gly Glu Ser Val Trp Asn
Lys Glu Ser Asp Leu Ala Thr Val 1070 1075 1080Arg Arg Val Leu Ser
Tyr Pro Gln Val Asn Val Val Lys Lys Val 1085 1090 1095Glu Glu Gln
Asn His Gly Leu Asp Arg Gly Lys Pro Lys Gly Leu 1100 1105 1110Phe
Asn Ala Asn Leu Ser Ser Lys Pro Lys Pro Asn Ser Asn Glu 1115 1120
1125Asn Leu Val Gly Ala Lys Glu Tyr Leu Asp Pro Lys Lys Tyr Gly
1130 1135 1140Gly Tyr Ala Gly Ile Ser Asn Ser Phe Ala Val Leu Val
Lys Gly 1145 1150 1155Thr Ile Glu Lys Gly Ala Lys Lys Lys Ile Thr
Asn Val Leu Glu 1160 1165 1170Phe Gln Gly Ile Ser Ile Leu Asp Arg
Ile Asn Tyr Arg Lys Asp 1175 1180 1185Lys Leu Asn Phe Leu Leu Glu
Lys Gly Tyr Lys Asp Ile Glu Leu 1190 1195 1200Ile Ile Glu Leu Pro
Lys Tyr Ser Leu Phe Glu Leu Ser Asp Gly 1205 1210 1215Ser Arg Arg
Met Leu Ala Ser Ile Leu Ser Thr Asn Asn Lys Arg 1220 1225 1230Gly
Glu Ile His Lys Gly Asn Gln Ile Phe Leu Ser Gln Lys Phe 1235 1240
1245Val Lys Leu Leu Tyr His Ala Lys Arg Ile Ser Asn Thr Ile Asn
1250 1255 1260Glu Asn His Arg Lys Tyr Val Glu Asn His Lys Lys Glu
Phe Glu 1265 1270 1275Glu Leu Phe Tyr Tyr Ile Leu Glu Phe Asn Glu
Asn Tyr Val Gly 1280 1285 1290Ala Lys Lys Asn Gly Lys Leu Leu Asn
Ser Ala Phe Gln Ser Trp 1295 1300 1305Gln Asn His Ser Ile Asp Glu
Leu Cys Ser Ser Phe Ile Gly Pro 1310 1315 1320Thr Gly Ser Glu Arg
Lys Gly Leu Phe Glu Leu Thr Ser Arg Gly 1325 1330 1335Ser Ala Ala
Asp Phe Glu Phe Leu Gly Val Lys Ile Pro Arg Tyr 1340 1345 1350Arg
Asp Tyr Thr Pro Ser Ser Leu Leu Lys Asp Ala Thr Leu Ile 1355 1360
1365His Gln Ser Val Thr Gly Leu Tyr Glu Thr Arg Ile Asp Leu Ala
1370 1375 1380Lys Leu Gly Glu Gly 1385363279DNABrevibacillus
laterosporusmisc_feature(1)..(3279)Cas9 single long
open-reading-frame 36atggcataca caatgggaat agatgtgggg attgcttcgt
gtggatgggc cattgtagat 60cttgaaagac aacgaataat agatataggt gttcgaactt
ttgagaaagc agagaatccg 120aaaaatggag aggctcttgc tgttccaagg
agagaagcta gatcaagccg taggagatta 180cggagaaaaa aacatcgcat
tgaaagatta aagcatatgt ttgttcggaa tggactggcg 240gttgatattc
aacatcttga gcagacgtta cgtagtcaaa atgaaataga tgtatggcaa
300ttacgagtag atggtttgga tcgaatgtta actcaaaaag agtggcttcg
tgtattaatt 360catcttgcac aacgtcgtgg ttttcaatca aatcgaaaaa
cagatggttc aagtgaagat 420ggacaggttc ttgtaaatgt aacggagaat
gacagattaa tggaagagaa agattataga 480accgtagcgg aaatgatggt
aaaagatgaa aaattttctg accataagcg aaataaaaat 540ggaaattatc
atggagtagt gagcagatct tctttactag ttgaaataca tacattattt
600gaaacccaaa gacagcacca taattctttg gcatcgaaag atttcgagct
ggaatatgtt 660aatatttggt ctgcacaacg gcctgttgca acaaaagatc
aaatagaaaa aatgattggt 720acatgtactt tcttaccgaa agaaaagaga
gctccaaagg catcttggca ttttcaatat 780tttatgcttt tgcaaacgat
caatcatatc cgtataacaa atgtacaagg cacgagatca 840ttgaataaag
aagaaattga acaagtggtc aatatggcac ttaccaaatc aaaggtatct
900taccatgata ctagaaagat ccttgattta tcagaagaat atcaatttgt
tggcttggat 960tatggaaaag aggatgaaaa aaagaaagtt gaaagtaagg
aaacgatcat caaattagat 1020gattaccata agttaaataa gatttttaat
gaagtggaat tagctaaagg agaaacgtgg 1080gaagctgatg attatgatac
agtagcgtat gccctcactt tctttaaaga tgatgaagat 1140attagagatt
atttgcagaa taaatataaa gatagtaaaa atcgtcttgt taagaacttg
1200gctaataaag aatacacaaa tgaattgata ggaaaagtga gtacgctatc
ttttcgtaaa 1260gtggggcatt tatcattgaa agccttacga aagatcatcc
cgtttttgga acaaggtatg 1320acgtatgata aggcgtgcca agcggctggt
tttgactttc aaggtatatc aaaaaagaag 1380agatctgtcg tcttgccagt
cattgaccaa atttccaacc cagtagtaaa tcgtgctctt 1440actcaaacac
gtaaagtaat taatgctcta atcaaaaaat atggttcacc tgaaaccata
1500cacattgaaa cagcaagaga gttatcaaag acatttgatg aacgaaagaa
tattaccaag 1560gattataagg agaatagaga caagaacgaa catgcaaaaa
aacacttatc tgaattgggg 1620attattaatc ctactggtct tgatatagta
aagtacaagt tgtggtgtga gcaacaaggg 1680cgctgtatgt acagcaatca
acccatctcc tttgaaagac tgaaagaatc aggttatacc 1740gaggtagatc
acattattcc atacagccga agtatgaatg atagttataa caatcgcgta
1800ttggttatga ctagagaaaa tcgtgaaaag ggtaatcaaa ctcctttcga
atatatgggg 1860aatgatacgc aaagatggta tgagtttgaa cagagagtca
caacaaatcc acaaataaaa 1920aaggagaaac gacagaacct cttgctaaaa
ggattcacta atcgtcggga actagaaatg 1980ttagaaagaa acttaaatga
tacacgttat ataactaagt
atctttcaca ttttattagt 2040acgaatttgg aattctctcc tagtgataaa
aagaaaaagg tagttaatac aagcggtcgt 2100atcacttccc atttaagaag
tagatgggga ttagaaaaaa atcgtgggca aaatgaccta 2160caccatgcaa
tggatgcgat cgtcattgct gtcacctcag actcgtttat tcaacaggta
2220acgaattact ataaacggaa agaaagaaga gagttgaatg gggatgataa
gttccctcta 2280ccttggaagt tttttagaga agaagtcata gcaagattaa
gccctaaccc aaaagaacaa 2340attgaagcat tgcccaacca tttttatagc
gaagatgaac ttgctgatct tcaaccgata 2400tttgtgtcaa gaatgcccaa
gcgtagtata actggagagg cgcatcaggc acagtttcgt 2460cgagttgtgg
gtaaaactaa agagggtaaa aatattactg caaagaaaac tgctttagtt
2520gatattagct atgataaaaa tggtgatttt aatatgtatg gaagagaaac
agatcctgct 2580acttatgaag caattaaaga aagatatctc gaatttggag
gaaatgttaa aaaagcattt 2640tcgacggatt tacataaacc gaaaaaagat
ggcaccaaag gtccactgat aaaatctgtg 2700agaataatgg aaaataaaac
attggtacat cccgtaaata aaggaaaagg cgttgtgtat 2760aacagctcta
ttgtaaggac agatgtattt caaagaaaag agaaatatta tttactacct
2820gtgtacgtaa cagatgtaac caaggggaaa ctaccaaata aggtgatcgt
tgccaaaaag 2880ggatatcatg attggattga ggttgatgat agcttcacat
ttttattcag tctatatccg 2940aatgatttaa tattcatcag acaaaatcca
aaaaagaaaa tatcattaaa aaaacgtatt 3000gagagtcatt ctatttctga
tagtaaagag gttcaggaaa ttcacgccta ttataaaggg 3060gttgatagtt
caactgctgc tatagagttt attattcatg atggtagtta ctatgcaaaa
3120ggtgttggtg tccaaaatct agattgtttt gaaaaatatc aagttgatat
tctaggtaat 3180tatttcaaag tgaaaggaga aaaacgactt gagttggaga
catctgatag taaccacaaa 3240ggcaaagatg tcaattcgat taaatcaact
agtcgttaa 32793736DNABrevibacillus
laterosporusmisc_feature(1)..(36)Repeat1 37atcatatcat atcgagtttt
agtaaggaac tatagc 363836DNABrevibacillus
laterosporusmisc_feature(1)..(36)Repeat2 38atcatatcat atcgagcttt
agtaaggaac tatagc 363936DNABrevibacillus
laterosporusmisc_feature(1)..(36)Repeat3 39atcatatcat atcgagtttt
agtaaggaac catagc 364036DNABrevibacillus
laterosporusmisc_feature(1)..(36)Repeat4 40atcatatcat atcgagtttt
agtaaggaac tatagc 364136DNABrevibacillus laterosporus 41atcatatcat
atcgagtttt agtaaggaac tatagc 364236DNABrevibacillus
laterosporusmisc_feature(1)..(36)Repeat6 42atcatatcat atcgagcttc
agtaaggaac tatagc 364336DNABrevibacillus
laterosporusmisc_feature(1)..(36)Repeat7 43atcatatcat atcaagcttt
agtaaggaac tatagc 364436DNABrevibacillus
laterosporusmisc_feature(1)..(36)Repeat8 44atcatatcat atcgagtttt
agtaaggaac tatagt 364529DNAArtificial Sequencesynthesized sequence
Blat-Cas9-dir 45taccatggca tacacaatgg gaatagatg 294635DNAArtificial
Sequencesynthesized sequence Blat-Cas9-rev 46ttctcgagac gactagttga
tttaatcgaa ttgac 3547177RNAArtificial Sequencesynthesized sequence
Blat sgRNA Direct 47gggcgcuaaa gaggaagagg acagcuauag uuccuuacug
aaagguaagu ugcuauagua 60agggcaacag acccgaggcg uuggggaucg ccuagcccgu
uuuuacgggc ucuccccaua 120uucaaaauaa ugacagacga gcaccuugga
gcauuuauuu ccgaggugcu uuuuuuu 17748118RNAArtificial
Sequencesynthesized sequence Blat sgRNA Reverse 48gggcgcuaaa
gaggaagagg acaaucauau cauaucgagg aaacuugaua ugauaugaua 60cuuucauuuu
auauccauau aucaucgaag ucaaucucau uuaucugucu auuuuaug
1184968DNAArtificial Sequencesynthesized sequence GG-969
49gggcgctaaa gaggaagagg acagctatag ttccttactg aaaggtaagt tgctatagta
60agggcaac 685062DNAArtificial Sequencesynthesized sequence GG-839
50ctaaaaacgg gctaggcgat ccccaacgcc tcgggtctgt tgcccttact atagcaactt
60ac 625155DNAArtificial Sequencesynthesized sequence TK-149
51gatcgcctag cccgttttta cgggctctcc ccatattcaa aataatgaca gacga
555262DNAArtificial Sequencesynthesized sequence TK-150
52aaaaaaaagc acctcggaaa taaatgctcc aaggtgctcg tctgtcatta ttttgaatat
60gg 625371DNAArtificial Sequencesynthesized sequence GG-840
53gggcgctaaa gaggaagagg acaatcatat catatcgagg aaacttgata tgatatgata
60ctttcatttt a 715475DNAArtificial Sequencesynthesized sequence
GG-841 54cataaaatag acagataaat gagattgact tcgatgatat atggatataa
aatgaaagta 60tcatatcata tcaag 755537DNAArtificial
Sequencesynthesized sequence TK-124 55taatacgact cactataggg
cgctaaagag gaagagg 375626DNAArtificial Sequencesynthesized sequence
TK-151 56aaaaaaaagc acctcggaaa taaatg 265732DNAArtificial
Sequencesynthesized sequence TK-126 57ataaaataga cagataaatg
agattgactt cg 325837DNAArtificial Sequencesynthesized sequence
GG-935 58caaattctaa acgctaaaga ggaagaggac agtcccg
375945DNAArtificial Sequencesynthesized sequence GG-936
59aattcgggac tgtcctcttc ctctttagcg tttagaattt gagct
456021DNAArtificial Sequencesynthesized sequence pUC-EheD
60ccgcatcagg cgccattcgc c 216122DNAArtificial Sequencesynthesized
sequence pUC-LguR 61gcgaggaagc ggaagagcgc cc 226221DNAArtificial
Sequencesynthesized sequence Sense DNA Strand of Cleaved Sequencing
Template 62gaggaagagg accagtcccg t 216311DNAArtificial
Sequencesynthesized Anti-Sense DNA Strand Sequencing
Readmisc_feature(11)..(11)n is a, c, g, or t 63acgggactgt n
116421DNAArtificial Sequencesynthesized Anti-Sense DNA Strand of
Cleaved Sequencing Template 64acgggacctg tcctcttcct c
216511DNAArtificial Sequencesynthesized Sense DNA Strand of DNA
Sequencing Readmisc_feature(11)..(11)n is a, c, g, or t
65gaggaagagg n 116627DNAArtificial Sequencesynthesized Sense DNA
Strand of Target and PAM 66cgctaaagag gaagaggaca gtcccgt
276727DNAArtificial Sequencesynthesized Anti-Sense DNA Strand of
Target and PAM 67acgggactgt cctcttcctc tttagcg
2768118RNABrevibacillus laterosporusmisc_feature(1)..(118)"Direct"
tracrRNA region downstream of the anti-repeat 68aagggcaaca
gacccgaggc guuggggauc gccuagcccg uuuuuacggg cucuccccau 60auucaaaaua
augacagacg agcaccuugg agcauuuauu uccgaggugc uuuuuuuu
1186958RNABrevibacillus laterosporusmisc_feature(1)..(58)"Reverse"
tracrRNA region downstream of the anti-repeat 69acuuucauuu
uauauccaua uaucaucgaa gucaaucuca uuuaucuguc uauuuuau
58704107DNALactobacillus reuterimisc_feature(1)..(4107)Lreu Cas9
Open Reading Frame 70atgataaaga aagactataa tattggactt gatattgggg
caacttcagt cggctttgct 60ggtattgatg aacagtatga cccaattaag ttaaagggaa
aaacagtagt gggggttaac 120ctatttgaag aagggcaaac agcggcagat
cggcgttcct ttcgcactac ccgtcggcga 180ttgaaccggc gtaaatggcg
cttatcatta ttagaagagt tttttgaccc atatattacg 240cctgttgatc
ccgcattctt tgcacgtttg aaggaatcaa atctttcacc aaaggataat
300aacaaaaatt ttagtagatc attacttttt cctgatatta cagatcagaa
attttatgaa 360gaatatccga caatttatca tctgcgatat gcattgatga
ctgaaaataa aaaatttgat 420cttcgggcta tcttcttagc aattcaccat
atgatcaagt accggggaaa tttcttaaat 480tccacccctg tagcacattt
tgatacgagt aagattgatt ttgcgaatga ttttagtaaa 540ctaaatcgtc
tttacttgaa tgaagatcct aataatattt ttgaaattaa tttacaaaat
600gtaaaagaaa taagtgatat tttactggat catagtatta agaagtttga
taagcaaaag 660caagttgcta agcttttact tacatctcaa aatgataagg
aactggataa aagaaataag 720cagattgcta ctcagatcag taaagcaatt
cttggctata atttttcact aaatgaaatt 780ttgaaacttg aagcagtaaa
taaaagcaaa tggaaactaa actttagtag cgccgacatt 840gatgatacgt
tacccgactt aatttccgaa cttgatgaaa gtcaagaatc aattttaaat
900attattttaa gtttgtactc acggttgact ttaaatggaa ttgtcccaag
tggaatgagt 960ctttccgaat caatgattga taaatatggt actcataagg
aacaccttga tttattaaag 1020aaatatttaa aaacccttcc tataaaaaat
cggaaagaga ttgctgaagc ctatgccgaa 1080tatgttggaa attctttgaa
gaaaagcgga catatttctc aagaagaatt ttataaagca 1140gtaaagaaga
atcttgataa atccgaaaca gctcaaaaaa ttcttagctt aattagtgaa
1200gagaagttta tgcctaagca acggaccaat caaaatgggg tcattccata
tcaacttcac 1260caaaaggaac ttgatcagat tattgtaaac caatctcaat
attatccatg gttagcagaa 1320ttaaatccag ttacggagca taaagatgca
aagtataaac tagatgaatt gattgctttc 1380cgggtaccgt attatgttgg
accgttaatt gatcctaaaa caatccctca aacagaacaa 1440ggaaataaaa
atgcttcatt tgcgtggatg gttcgtaaag aaaatggtca aataacgcca
1500tggaattttg ataaaaaagt agatagaatt tcgtctgcaa ataactttat
taagcggatg 1560actactaagg atacgtattt aatcggcgaa gatgtattac
ctgctcatag ccttatttat 1620gaacgtttta aggtattaaa tgaactaaat
atgattcggg taaatggtaa aaaactctct 1680gtttcggtta aacaaaatct
ttataatgac ctatttaaac aacaaaagac aattaatagg 1740aagaagttag
ctaattatct ccaagcaaat cttggtatcc ctgaacgacc tcaaattaca
1800ggtctctcag atcctgaaaa atttaattca caattaagct catatattga
tttacaaaaa 1860atactaggaa gtgaaattgt tgatgatcct aataagcagg
atgatttaga aaagattatt 1920gaatggtcaa ctgtttttga agattcacgg
atctataagg ttaagttaca agaaattgga 1980tggttcactg aaaaacaaaa
gaatgagtta gtaagtcatc gctatcaagg ctggggacga 2040ttatcaaaga
aattactagt tgagctaaaa gataagaatg ggagatcaat tattgatttg
2100ctttggaatt cacagcgaac atttatggag attcaaagtc gtccagagtt
tgctgaacag 2160attactaatg agaaccaaga taaattaact gaggataatt
acgaagacgt actggccgat 2220gcctatactt caccgcaaaa caaaaaggca
attcgtcagg taattaaagt tgtcgatgat 2280attgttaagg caactggaaa
agcaccgaag tttatttcgt tagagtttgc tcgctcggat 2340gaacggtcag
atcgtgtaaa atcaaggaaa acacatatcc aaaaaattta tgaaactacc
2400gctaaagaat tactgaaaga tgatcagtta attaaagagt taggtagcgt
ttcagattta 2460tcagatcggt tatacttata ctttacccag cttggtcgtg
atatgtatac aggtaagcca 2520attaatattg atgaaatttc gactatgtat
gacattgacc atatccttcc gcaggcgttt 2580cttaaggacg attcattaga
taaccgggta cttgtaagac gacaggataa taacgctaaa 2640tctgacactg
ttccggcttt gaaatttgga aagatgaaac cattttggaa caaattacaa
2700aagcacggct tgattagtaa acgtaaactt aataacttgc aaactaatcc
tgaaagtatc 2760gataagttta aagcagttgg ttttgtgaac cgtcagttag
ttgaaacacg ccaagttata 2820aaattagctg caaatatttt agcaagccgt
tatcctgatt ctaagattat tgaagttaaa 2880gcaagtttaa ctcatcaaat
gcgtgaatca tttaacctaa ttaagaatcg ggatgttaat 2940gactatcatc
atgctgttga tgcgtattta agtgcatttg tagggcaata cctatataat
3000cgttatccta aattgcagcc atattttgtt tatggtcagt ttaaaaaatt
tgataaacaa 3060agtactcgga ttgggatgaa aactaaccac tttaattttt
tgtatgatct tgagcccgaa 3120ggtaaaaatg taaaaataaa gaaaccaact
aagattataa acaaggaaac tggtgaaatt 3180attggtgatc gtgatgaatt
agttgccaaa ttgaaccgtg tttataattt taagtacatg 3240ttagtttcac
aagaagtata tacgcgaagt ggtgcattgt ttgatcagac aatttaccca
3300gctaattcag gtaagaaatt aatcccgctg aagcaaaata aaactacagc
aatttatggt 3360ggatacagtg gaagtaaagc agcgtatatg tctatcatta
ggttacgaga taaaaagggt 3420ggaacttacc gtatcgttgg aatcccagta
agagcagtta ataaattaaa tcaagcgaag 3480aaaaagagta atgaaaaata
tcttgcagaa ttaaaggcag tcattgagcc acaaattgct 3540aagacaaaga
aggatcgaaa gactggtcaa agggttcttg taccacaaga atttgatgtg
3600attatccctg aagttatgta tcgtcagtta atagttgatg gtgaccagaa
atttacgttg 3660ggtggaacta ttgatagata taatgctgtt caattagttt
tgaatcaaga aatattgaca 3720ttccttgagc aacctactaa atataaggac
gccgatacaa aattgttgga tatatatgat 3780caaatagtaa atttagtaga
gaaatatttt atgctgtttg atagcaaacg acttgctgca 3840ggtcgagttg
cgtttgaaaa attacctact ttacaacctg tcgataaaat gccttctaaa
3900ttaataataa ttagaagaat tattcaggga cttcatgaca atgctgctcg
aacagattta 3960aaggcgatta atggaagttc gtcatttggt aggttacaaa
aaagaaatgg aattatacta 4020tcacctaatg catgcctgat ttaccagtca
ccaactggct tatttgagag aaaagtttat 4080ctaaatacta tttcaccatt aaaatag
4107714110DNALactobacillus rossiaemisc_feature(1)..(4110)Lros Cas9
Open Reading Frame 71atggataaat caaaaccata tggaattggc ctagatattg
gtacaaattc tgtcgggttc 60gtggcgactg atgctgaggg gcaccttatc cgattgaaag
gaaagacggt tattggagcg 120tacctgttta atgctggaat atcggcagcg
gaaagacggg ggtttcggac gacaagacgt 180cgattatcgc gggttaaatg
gcgcctagga cttcttagag aaatatttga gactcatttt 240caggaaagta
tgggagaaaa tgaggataat gatttctttt tacgtttcaa atactcgaat
300atttcaccta aagacccaca attttcgacg gctaagggtt tatttaatga
tcgaacggat 360aaggaatttt atgatcagta tccgactatt tatcatttgc
gtcgggcttt gatgactgaa 420gaccatcagt ttgatattcg ggaaatatac
atcgcaatgc atcacattgt gaaatatcgt 480ggtcattttt taaaagaagg
acgcgccaaa gactttaaag ttggggattt aagactgctg 540gataatttca
aaatgatgaa tgaacaaatt gaagaaatta atcctttgtg gcagttgaaa
600ttaccaactg atgacgcctc tattaagtcg ataaccgcta ttttactaga
caatactcag 660agccaaaatg atcgtcaaaa agcagtgacg aaagtaattc
tggcaacact tgttaaggcg 720agtgacaaag acattaacgc tgcacgtaaa
cggtttgttg gcgaattgag taaggctatg 780gttggcctta aaacaaaact
ttgggttctt gcagatgttt cgcagaatgg tgattgggaa 840attaagtatg
aaaactacgc tgactttgcg gaaacgatcg gttccggtga aagtgacacc
900attcaaagtc tttttaacga gattaatgat ctatatgggg ttattacgtt
ggctggcatt 960attcccaaag aagctgagtc attttctgac ggaatggtcc
gcaagtatga acatcatcgt 1020aaaaatctag aactgttaaa agtttattgt
gcggaacaat cagatggtaa acggggacga 1080cagattcgtc agacttatga
taaatatatt gatggggtag atagcaaaca gtttacgcag 1140gaagactttt
ataaggcatt aagcaagttc actgcgaagg atgaagcgac tagtgaaaat
1200gccaaattaa ttgctcagga aattgcagtt ggaactttca tgcctaagct
gcgaacaaag 1260gctaatggca ccattccaca tcagttgcac caaaaagaat
tggatgcaat tattgaaaat 1320caaaaaaagt attacccatg gcttggtgaa
gtcaatcccg ttgagagtca tcgtcgcgca 1380ttgccataca aattggatga
attagtcagt tttaggattc catattatgt tggaccaatg 1440gttacgccaa
caaagggaga tccagaaaaa agtaaatttg cctggatggt tcgaaaggaa
1500ccgggtacca ttacgccatg gaatttcgat caaaaagtag accgatcggc
gtctggtgaa 1560gcgtttattc aacgaatgaa aacgactgat acatttttaa
ttggcgaaga tgtattgcca 1620caacaaagtc tgctgtatca gaaatttgaa
gtactaaatg aattaaacaa aattatgata 1680aacggtaagc caatttgcag
agaacagaaa caacggcttt tcaaacaact gtttatgcaa 1740tataagactg
tgacagtgaa gaaagttcaa caaaatttaa ttgcgaatgg tgaagagtct
1800gaaaatgtgc caattactgg cttgtcagat cctttacggt ttaatagttc
attcagcact 1860tatatcgatt acaaagatat tttgggtaca gctgctgtta
acgataatgc aaagcaaagc 1920gatattgagc agataattgc atggtccaca
atttttgaag atgcagccat tttccgagag 1980aagttaaatg atattacttg
gcttaatgac gatcaacgca ataagctcag tcataaacgt 2040tatcgcggtt
ggggtcgtca ttctcgcaaa ttgttagctg gtcttcgtga cggagaaggc
2100cagactatca tcgaacggct atggaacaca aatgacaact tcatgcagat
tcaaaacgat 2160agtgaaatag cgcgtcaaat taccgaagca aactcaagca
agatggcaac cgctgaggga 2220acagacgaaa ttatcgatgg cttctacact
tcacccgaaa acaaaaaggc cctgcgtgaa 2280gtaatgaaag ttgtgaagga
tattcaacgt gcgcatcatg gtcaggcacc tgcttgggtt 2340tatattgaaa
gtccgcgaga gacaccaaga cccggtcagc ggacagttag tcgagaacaa
2400cagttaacag atttgtatga gggtgcagca aaagaaatcg ttgatgatgc
cgttttaaat 2460gaactaaagg acaaggttaa atccaaggaa aactttacgg
ataaattagt tttgtatttc 2520ttgcaaaatg gacatgatat ttatgcgaat
gacagtatca acattgataa ccttaatgca 2580tatgatatcg atcatgtttt
accgcaaagt ttgataaaag atgatagcct agacaatcgg 2640gttctcacaa
cacatgaaag aaatcttaaa aaatctaatc ggtttgctac agaactgttt
2700gccgatcaaa ggaaaaaatg ggaaaaatgg catcgattag gattgatttc
gtcacgtaaa 2760ttaaaacatt taactatgca gcccaattca gtagaaaaat
ttgcacatgg atttattgca 2820cgtcaactca ctgaaacgag gcaaataatt
catttaacgg cgaatgtact ttcaaattta 2880tatcaagaaa atgacactaa
aatcgttatg attaaggcgg gattaaactc ggagtttagg 2940cgtacatttg
attttccaaa gaatcgtagt gtaaatgatt accaccatgc ctttgatgca
3000tttttaacgg ccaaaattgg acgttactta ttagcgagat atccaaaatt
agaacccttt 3060tttgtttatg gaaacttcgt taagaatcct aaagcaatga
agcgattgag tagttttgat 3120ttcattgctc agttagctgc taaaactgat
gatactagtc atatagatca acgcagttta 3180aaacaagttc ccgttgttaa
tgaagaaaca ggggaaattg tttgggataa ggatattgag 3240cttgctgaac
tggacaagac gtataactac aaaactatgt tagtgaaacg tgctcagact
3300gaaaataatg ctcaaatgtt caaacaaaca gtttttaagg ctcgtgacaa
tcaaaataaa 3360acgctaattc ctgtcaaaaa tggattatct actgacgttt
atggtggtca ttcacaacag 3420gcgatttctt acttatgtat cgtttgggtt
ggacaaaaga aaaaatatcg cgttttggga 3480attagtaccg cccatgcagg
catattgaac aatttcgaaa aaaactacgg ccggtttgaa 3540gcgaaaaaga
aactacaaga aattgtctcc aatacactag ataatgcgga cagaaacgat
3600tttaagatag ttgctcctaa agttttgttt gaacaagtgg ttgaggacga
taacatgaag 3660tttgggttgg gtagtgctag tgactataga aatgttcaac
agttgttttt atcccggaaa 3720aatcagttgc tattggctaa tatgatgact
gaccaaatac acgaccaaga tttagtccac 3780ctttttgacg aaatagttgg
tcaaatgaat gcgcattttc ccatttttga tcgcggaggc 3840tatcgaagtt
cattgaccca atctcgtgat aaatttttga aattaccttt taaaaagaat
3900gaggatttga tcacgaaaca agaggttatt cgaagaattt tagatggatt
acatgctaat 3960gcaaatcgta aagatcttaa aatcattggt agtaaaggtg
attttgggag attgggaaca 4020aagaaaatct acttatcgaa agatgcgaag
cttatttaca cctcaccaac atgtcttttt 4080actcgtactg ttccactcag
ttctttgtaa 4110724041DNAPediococcus
pentosaceusmisc_feature(1)..(4041)Ppen Cas9 Open Reading Frame
72atggaaaaag taccatataa tattggtctc gatattggga cgtcatcaat tggttttgca
60gcaaccgata atcttaacaa gcctattcgg gcaaaaggta aaaccgttat tggagtacga
120ttatttgaag agggtaaaac tgcagctgat cgacgtggat ttagaacaac
acgtcggcga 180ttatcaagaa gaaagtggcg tttaaggtta cttgatgaga
ttttcgataa agaaatggct 240aaggtagata ataccttttt tgctcgtttg
aaggagtcga atctttcacc aaaagatgct 300aataagaaat atttaggatc
attattattt ccagaaaaga aggattttaa gttttatgaa 360gattatccaa
cgatatatca tctacgttat gctctaatgc atgaaaaacg tcaatttgat
420attcgtgaag tttatttagc aatgcatcat attatcaaat atcgtggtaa
ttttttaaat
480tcagccccaa tgaattcttt taaaacacaa gattttgatt ttgtggctaa
atttgaaaaa 540ttaaacgaac tctttgaaag tattgatgcg gaacatgaaa
cgaaatttga tattgaaaat 600atttctaagt ttagagatat catgctaaat
caagatattc gtaagttaga ccgaaaaaaa 660caagccgcta aaattttaat
cctagatagt acagataaga cggctaaaaa gattaataat 720aaaattgcaa
ctgcagtagc caattcagct ttaggatata agtttgcgtt ggatgctatt
780ttgaaattag atgtagaaga gtctaaagat tggtcgatta gtttaaacga
tgaagaaatc 840gatagtattt tagataactt aactagtgat ctggatgcgg
aacgcatcga aatcattgag 900atattgagag atttatacag tcatattgct
ttaaatgaaa ttgtacccaa cggacaatcc 960ttgtctaaat ccatgatgga
taaatacgat aagcatcatg cggatttaga tgtattgaaa 1020aaagttattt
ctaatatgga cgatcgaaaa aaagctaaat ctttaaaaaa tatttacaat
1080cagtatgtgg gtaaaactaa tgataaagtt ttggataaag atgaattcta
taaacaaatc 1140caaaagaatt tagatgagtc tgaagatgct atgaagattg
tcaatgaaat tgaacttgat 1200caatttatgc caaaacagcg tacatctcaa
aatggagtga ttccgcatca acttcaccaa 1260aaagaattag atgaaatcat
tgaaaatcaa aagcaatact atccattttt agcagagcct 1320aaccctaacg
agaaaagaaa acctcatgct caatttaaat tagatgagtt gatagctttc
1380aaaattcctt actatgttgg tcctttgatt acaaaagaag aacaacaagc
ccaatctggt 1440gctaaatttg cctggatgaa gcgtaagcaa gatggggtga
taactccttg gaatttcgat 1500gaaaaagtgg atcgaatggc gtcagctaat
gaatttattc gccgaatgac gactaaagac 1560acgtatttgt taggtgaaga
tgtacttcca gacgaaagct taatctatca aaaatttaaa 1620gttttgaatg
aattgaataa tgtcaaagtt aacgataaaa aactgacagt ttctgataag
1680caggatattt tcaatgacct gtttaagaaa cagaaaactg tttccgtaag
taaacttcaa 1740aagtactttg ttacagaaaa gcattacttg acagagccaa
cgatcaaggg gttatcagat 1800acgaagaaat tttctaatag cttatcaact
tatattgatt ttgaaaaaat atttggaaat 1860gaaattcttg ctgatcaaaa
taagcaaaat gatttagaaa aaattattga gtggtcaact 1920atttttgaag
atcgtaaaat ttttgaagat aaattgcatg agattgaatg gttaactgaa
1980aagcagatta aagctgttcg gagatatcgt ggatggggac ggctttctaa
aaagctacta 2040gtcgatttaa gaaataatga aggtaaaagc attttagatg
aactttggcg taccaatgat 2100aactttatgc agattcaagc acgtgaggaa
tttgctaagt cgattgtcga agctaaccaa 2160aagttaatga atcttggtgg
agctgccagt gttcaaaata ctgtggaaag taccttagag 2220gatgcctata
cctctcccca aaataaaaag gctattcgtc aagttattaa agttgttgaa
2280gacatagtca aagcggttgg ctacgctcca gagaaaatca caattgaatt
tactcgaggg 2340gcagataaaa atcctagacg gactcaaaat cgtcaaaagc
aaatcatgga agtatataaa 2400accgcggcta aggaaatcgt agatgcaact
ttgaagggac aacttgaaaa tgaggaaaag 2460ttaactgata agttatattt
gtatttcacc cagttaggta aagatattta tagtggcgaa 2520acaattaata
tcgatcaact taataattat gatattgacc atattttgcc acaagctttt
2580attaaagatg attctctgga taatcgagtg ttaacaagtc gggatttaaa
caacggtaaa 2640agtgatagtg tgcctgtgaa gcagtttggt gcaaacatga
aatcattttg gatgagactc 2700caagcacatg gattaatctc aaaaagaaag
cttaataatt taatgacaga tccagacagt 2760attggaaaat ataccatgca
aggatttgtt agaagacagc tagttgaaac tagccaagtt 2820attaagttga
ctgcaaatat actaggtgcc atctatgggg aaaatactga tattgtggaa
2880atcccagcga agctgactca tcaaatgcgg gagaagttca atctgtataa
ggtgcgtgag 2940gtaaatgatt atcatcacgc ctttgatgcg tatttaacga
cctttgttgg taactattta 3000ttcaaacgct atccaaaact acgtccttat
tttgtatatg gcgatttcaa aatgactgat 3060aatgccttga aagggatgcg
tagattcaat ttcttacatg acttaaaaga tgatgaagta 3120ttagtagata
acgaaactgg tgaagtttta tgggaaggtc aaaagtctat tgaagagttg
3180aagaagattt atggttataa gtttatgtta acgactcatg aagcatatac
gcaacatggt 3240ccaatgttta agcaaactgt atatagttct gatacgcctg
gcaaactaat taagataaag 3300aacaataagc ctactgaaat atatggtgga
tatacttcga atacagatgc ttatatggcg 3360attgtaagaa taaaagctag
aaagggggat acttataagg ttgtcggagt acctagaaaa 3420gaaggagatg
ctttaagtcg aatcaagtta ttagacgaaa gtaaatacca tgagaacttg
3480aagaatattt tagcaattaa tttgaataaa tctttgaaca agtttgatgt
ggttctcgat 3540aaagttaaat atcggcaggt tatttatgac ggtacagatc
atttaatgtt aggtagttca 3600aaatataaat atagtactaa acaattagta
ttatcagacc aatctatgaa aattttaagc 3660tctactggtg aattctcgga
cgaggaactt atcaaggtgt tcgacgagat tatgtacatt 3720gttaataaga
actttagttt gtatgatacc cgaggatttc gtgataagtt aaataatgct
3780agggaaaatt tcataaaatt gcctaataaa acattgttcg aaaagggaaa
attaaaacaa 3840catagcaagt tagaaatatt aaaacaaatt ttaattggac
tgcatgcgaa tgctggtcga 3900ggtgatttaa aagatattgg agttaataat
tttggagcta tggttgttac tgctggggta 3960acgttatcac ctgacgctac
aattgtttat caatccccaa ccgggctatt tgaacgcaaa 4020gttaaattga
gcgaccttta a 4041733393DNALactobacillus
nodensismisc_feature(1)..(3393)Lnod Cas9 Open Reading Frame
73atggaaaaca aaattagctt aggattagat attggagttg cttctgttgg ttttagtgta
60attgatgcac aagaaggtaa agtactggag ttaggggcca gactattcaa tagttcagtt
120tccgcggaga atcaaactcg tcgagatatg cgagggtcca gaagactaag
taatcgcaag 180aaacaacgtc gtaaagatgt agctcaacta tttaagactt
ttggattaat aaattatttt 240gataaagaaa attattttga taactttcaa
aataatttaa attcttatga attgagagtt 300aaaggattat cagagaaatt
atcaaaagaa gaactggtga atagtttgta tcatattgtc 360aaaaggcgtg
gtatcagtta tgacttggca gatgctgaca ctgatttcga tggttctgat
420tactcatcta gtttgaatca gaatcaattg gaattacaga cgaagactcc
agctgaaata 480caacttactc gcttgaatgt acatggagct gtacgtggaa
aagttacgat caatggcgag 540gatgaggaca ctatgcaagt attgttgaat
gtgttcccca ccaaatcatt tgtatctgaa 600gcaaaaaaga ttattcaaac
acaacaacaa tattatcctg atattttgac ggatatattt 660gaaactaagt
atttagaaat tctagaacgt aaaagagaat attttgtcgg acctggtagc
720gaaaaatcta gaactgattt tggtatctat aagaaagatg gtagaacttt
ggataatttg 780ttcgaagagt taattggtca tgacaaaata tatcctgatg
agttacgtgc gtctggtgca 840tcttatactg cacaggtttt taatgtctta
aatgatttaa ataatttacg tatttctagt 900tatgaaaacg ggaaaatttc
tacggaagat aaaataaaga ttgttgaaga tttaagaaac 960aatatcggca
atgtaattat aatgaaaatc attaaaaagg ttgccggttg tgaagatgat
1020gatattaaag gctatagatt ggatagcaag gacaaaccgg atattcattc
aatggccgtt 1080tatcgaaaag ttcatcgtga tttattgaaa tatgatgtgg
atattattaa gtggccaacg 1140gaatttattg atgaattaag tccaatatta
actttgaata cggagaacgg tgaaattcgg 1200aaacaaatgg tcaataagtt
gcaaccaaag tattcatttt taacggatga acttattcaa 1260gttattatta
ataacaaatc tagttttgac gttacttcca ataataaatg gcatcgtttc
1320tcattgaaaa caatgaatgt tttgatcgaa gaaatgttca gacgaccagt
tgaacaaatg 1380actcttattc aagaactggg attgattaaa gactcaggta
agagatttga aaattgtaaa 1440cttttgccat atagggaaat ctcaaaggat
atttttaatc cggtagcgtc caagtcagtt 1500cgtgaggctc ttaaaattgt
taatgccgta atgaagaagt atggtcaaat tgattatctg 1560gttatcgaga
tgcctcgtga taagaatgaa gaagaaatga agaaacaaat tgaaaaattc
1620caaaaggaaa ataataagca aaaagatgaa gcaataaatg agtttgttaa
aaaaattggg 1680aataagaatg ctgttgatga tggactaaga cgatatggtg
gtaaattata tttcaagatt 1740agattgtggt atcaacaaga tggaattgat
ttatataatg gcaaagtaat tgaaccgttt 1800gacttattga ataatattaa
taaatttgaa gtggatcata ttattccaga atcaatttca 1860tttgacgaca
gtattaataa caagacactt tgctatgctg atatgaatca aatcaaagga
1920caaaagacac cgtttgaatt tatgaacgag ggacatggcc agggctttgc
aaaaatgaag 1980gctatggtaa ataagaattc aaagttaaaa ggtaaacgaa
aaaattattt atttgatgaa 2040aatattagtg atattgaaac cagaaaaagg
tttatatctc gtaaccttgt tgatactcag 2100tattcgtcaa gagtggtttt
gaatagtttg caagagttct ttaaggaaaa agaaaccgga 2160actaaggtga
ctgtcgtccg aggtaagttt acttctaatt tacgcaagca ttggcatatc
2220aataagacac gtgatacatt tcatcatcac gccatcgatg catcaataat
tgccgcgaca 2280ccatttttaa gaatttggaa aaaagatgcc agtttattcc
ccatgcatgt ctctgaaaat 2340actgttgata ttgaaactgg tgagattctt
aatgatactg aatttaaaaa agacttttac 2400ggattaccgt atagcagttt
tattgaagaa ctgaacggtg ctgatgaccg gattaagttt 2460tcgcatcagg
ttgataggaa gatgaatcgt aaggtaagtg atgcgacaat ttattcgaca
2520cgtaaaggaa tgttgaataa agataaagaa gaaactgatt atgtcttggg
aaaaatcaaa 2580aatatttatg atgttagtga gtataacaaa ttcaaacgaa
tttacgataa agatccaaat 2640aaatttctgt tggctcatca tgatccaaaa
agctttgatg aacttagaaa aatcatgcag 2700gaatatccta gtaaaattga
taaagtaatg actaatggta aagtaaaatc tgtggatatt 2760tctccatttg
aattgtatcg tcgcaaacat ggaatggtac agaaatactc taataaaaat
2820aaaggccctg ttattaaaca acttaagtat ttagacaaga agttgggatc
tcacattgat 2880attactccga aggatacatt gaatgatagg catgttgtat
tacagtcact gaagccgtgg 2940agaactgatg tgtattacaa tagtgtgacc
ggtgaatatg aaattatggg tattaagtat 3000tctgatttga agttcaatgg
tggtgagtat ggaatcaaga tgagcaaata tttggaaata 3060aaagagcgtg
aacaggtttc agatgagtct gagtttctgt ttactcttta taagagagat
3120agaatccaaa ttataaattg tgaaaatgac gaaattgtag aaatgctttt
ctggtctaga 3180aataattcta atattgggta tgccgaactt aaaccggtgt
ataaatataa aactgaggat 3240gaaagttggc cggtatacgg atatggaaaa
aatcaaatcc taaaaagatt ggtaccaaag 3300aattgtaaaa tattgaaagt
aaataccgat atactgggca atccatacta tattaaaaaa 3360gaatccaaaa
acccaaaaaa tattcttgat taa 3393744086DNAArtificial Sequencesequence
from Sulfurospirillum sp. SCADC (Sspe)misc_feature(1)..(4086)Sspe
Cas9 Open Reading Frame 74atgacatcgt tgatttcact ggatttgggt
gggaaaaata cgggattttt tagttttaca 60gctaaagacg tttcaataat agatactttt
caatcaggaa cgattatcta tgatgaaagc 120tttgtgctct cacaagtcgc
acgacgaggc aaacgacacg gtaaacgtaa caacctacgc 180aacacgctag
tcaagcgact ttttttgctt cttttacaaa agcattatgg gcttagtctt
240gattttttac ccgacgaaat tttgggactt tttaataaac gaggttacac
ttatgcaagt 300tttgaaatca aagaagatga aaaagaaaat cttgaaagcg
atattttaaa agagtttttg 360aatgacaaac tcaattacac tatacaaaat
gatgacgaag tagaagagtt tttaaatcag 420atagcttcaa atgaagagac
gttcaagaac tataaaaagg attttgaaaa tttatttgga 480gcttcgacgc
atcagccaaa aaaacaaatt gaacttattg atgagataaa aaaagacctt
540gaaaaagaag atgccaaaga acttttagat gggttaaaag tcataaaaaa
aattatagac 600gaatttcaca aacaacaaaa ccaaggcaat ctcccacggg
ctaagtattt tgaagaactt 660tatcttgaaa tagaatacaa tctaaaaatc
caaaaattct ttacatgtaa ccatcttcac 720atcaatgaca tgcaatatct
catcggcaat ctctcaaact atcagctcaa agagctaaga 780cgctacttta
acgatgaagc tatggcaaaa gaagactttt ggtcttgcga aaaactgcat
840cgcatcacat ggcgatttat ccaaagttgg cacccaaaaa gtcctgaaga
caaacaaagg 900caaaaagaga atcttgcaaa ccttaaaacc aaaagtatta
ttgagttttt aaccaccacc 960aacccactga tgaccattcc gccgtatgac
gatatgaaca accgtggagc ggtcaaatgc 1020caaacattgc gactcaatga
ggagtatttg gataagcatc taccacattg gagagccata 1080gcacatacgc
ttgcaagtga gactcaaaaa gaaaatcttg agggtgtgac cgtaaaaggc
1140tacagtgaag atagcacgct tttacaccgc attttggaca cctctagcat
cattgaccct 1200tatagactta gaagtgatga gatagatagt tattgtgatg
tattgaccaa agacaatgct 1260tttgcattga aaaaatttgc aaaagagtac
tatcaacttg tcaaagaaaa agtacgaaca 1320ggtatttgga caaaagacga
tgatatgttc aaaaagtgtg accacaatcc accacacaaa 1380aacaatcaaa
tccacaattt agtcgcagga atcttaggca aacccatcgc aaaagagcgg
1440tttgaagcgt ttgaaaatga actttggaat gtaaaatttg gcaacaaaaa
actctcaagc 1500tattgtaaaa acattgaaga gtttagaaaa tcaaacggca
acctttttaa gcagattgta 1560gaattgggtg aagataaaga ggtacaaaaa
tatcaaaaag agttaaacga atgggtacga 1620aaaatcggtg agttttttaa
catcgaaacg ccttatagag cacgctttaa caaccttttt 1680tccatggcgc
aacttcatac catcatcgat accacacgca gtggttttaa cgctacatgt
1740aaatggtgta gctgcgaaaa tcaataccga gctagcacaa gaatagagat
agatgagcaa 1800acaggggaaa ttactaccaa tgccaactgc caacggctcc
cagccgatac acaaagacct 1860tttagtggca agattgagcg gtacatcgac
aaactaggtt acgagatagc caaagtcaaa 1920gccaaagagt tggagggcat
caaagaagat acgatagatt taaaaatcat cttagagcaa 1980aacgcttttg
cgtatgaaga gtctattcga aatgccaaaa tcaaaaatgc caatgcaaaa
2040gctaaaaaag ctcttgaaga agcacaaaaa agagggctta aaaatattga
agacaaaacc 2100aaaagaatta aagattttag caacagcatt tgcccctatt
gtgggcaaag tttaggcgaa 2160gatggcgaga tagaccatat cttatcacga
agttacaccc ttaaaaagta tgataccgtt 2220ttcaacagcg aaggcaatct
tttgtatgtt caccaaaaat gcaatcaagc aaaacttgca 2280aaaacagatt
attctttaca agatttaaaa atagatattt ctcaaaaatg gatagaagaa
2340caaatcgcta ctatcaaaac ctacaaaact ttttcagttc ttacgcaaga
acaacaaaaa 2400gcttttaaat atgccctatt tttagataac tcaaatgaag
cttaccaaaa agtaataagt 2460tggcttagaa ccgaccaaag ctcacgagta
aacggtacgc aaaaatattt ggctaaaaaa 2520atccaagaaa agctcaaagc
gatgtttcca gctaaaacat ttaactttga gtttatctta 2580gccgatgcca
atgatgtgca tgatttgaga attaaagcgt atcaattgcc agaaaagcca
2640aaagactcta aacaagaaac ctacagtcat actatagatg cggttatgag
tttggtcagt 2700gtatgggata aggtattgcc aaaaacagaa aaacctacaa
aagaagatat tttaaaattt 2760gccaatgtcg aaaattggag tgccctcaat
aatgaatttt taaccaaagg caaatcagca 2820aatcaaaaaa tagaagagat
gatacaagcc aatgattttg gtcaaaaaaa tatgaggcaa 2880gtttttagta
agcccatttt taaagatgaa tcaataggag aaaggtataa gccatttgtt
2940cgctatcaca atcaatttta cataggctat ccgataagca taaaagacgg
atacgatatg 3000cagcattgtc aagcaatgat ttcaaagaac gatatttcaa
gggtagagga gattcttaaa 3060gacacctctt tatgcactct tttaaaagaa
aagaatggca ttaaacttta ttctattaac 3120aagcaaagta tcaatgaact
gtcaaatcag ttctttaatt taaactatca aaacttaaac 3180gatgcacaga
agaaaaaatc tgaactggca gaatttgtca tcaatcattg taaatattat
3240gtcaaaaaaa catcggttat caatgcgcca caatttattg ataaagattc
gatgaagccg 3300tacccgtttt ataaagactg gcaaaaattt cacgaagctt
acaaaaaaga gcttgatgct 3360gaaccaaaaa ccaaaaaaga taacgggaaa
ttagtttatg atatatcagg catagatgac 3420ttctggactg aattttgcaa
aaaatatttt ggtataaaaa caaaagacaa tcgaaacaaa 3480gcgagaaagg
ttttttctat tgtggcgctt acttcagccc caggaacagt ctttagaatc
3540aaacgaaaaa cgcctaaagg acacatatac caagcaacgg cgattgacaa
tcaacaaatc 3600agcggagatt atgcaaatgt tctgttagct ggtaactcaa
aaacacttgc gttggcaggt 3660caaaaacctt catctgactt gaaaaaagag
ttgagtgtca aagagtcaaa agatattcga 3720gatataaagc ttgagccatc
aagatttttc aaagaggggt ttgattgtcg tggcattgaa 3780gttattgtca
ataaaacaag tgcgaccatt aaaaatttcc cacttacaaa aattgataaa
3840aaaatcaaaa aacttatttt taaaactctc tttgaaaaaa aagatggtaa
acgacaaaag 3900caaaaaacat caatttcatt gaaagaaaaa aatacaatgc
aagagacact aaaaaaatta 3960ttaaaagata gtataaaagt tactattaga
gatggtagca tttcgggtat tgaaattagc 4020aaaaaaactg ttaattttac
tttgccgttc aaaagtgaaa acttggcaaa actcttagat 4080gactaa
4086753444DNABifidobacterium
thermophilummisc_feature(1)..(3444)Bthe Cas9 Open Reading Frame
75atgtctgata agacatatcg tatcggtatc gacgttggtt tgtactctgt tggcttgtct
60gccattcagg taaatgatga tgacgatcca gtaagaatac tgaacgcaca gagtgtcatc
120catgatggcg gtgtcgatcc gaatgctcag aaaagcgctg atagccgacg
tgctcagtct 180gggattgccc gtcgcacgcg tcgtatgcgc cgtaatagga
agaaacgtct gaagcgtctt 240gatcaaattc ttgtggaatc tggtttccct
gtgagcagtg agaatgatct tgaaggattt 300gaaccgtggc ttcttagggc
gcaagcggct gatgccttca ttgaagatga ggatattcgc 360aagcgtgcaa
tttcggtttc atgtcgccat atcgcgcgcc atcgtggatg gagaaatcct
420tatttggatg ttcgaacgct gctggcggtt gactcgcctt cctcggcatt
ctatgacaag 480ctcgtggaga atgctgctct tgaaatggac ggccagatgc
ctgacagcga tgccacgccg 540gcacagattg tgcgcgatgt gcttgagtac
aagcgtggtg aagctgctat ccgtcttcga 600aagtcaacgg ctgagaataa
gaaaaatcgc ctcgctttgt tccctgagaa gatgatgcag 660gatgactatg
cttatgaatt gcgtttgatt ctggcgaagc aagcggttcc caaagatatc
720gctcgcaagc tcattttggc ggtgttccaa tcgcagtcgc caaaaggttc
tgcggagaag 780cgtgttggga aagatcccct tgatccgtct cagcctcggg
ctttgaaggc ttctctggca 840ttccaagaat atcgaatatt gaatattctt
accaatctgc gtttgcagga tggtggcgcg 900gagcgtcgtc tgtctattga
ggaaaagcag aagctctaca agatgctcgt ggaagatacg 960ggtcgcgaaa
agaagtacga gacatggaca gatatcgcat cggccatgga gtggaagcga
1020aattggctca agggagttgg gagtttgacg gccgatggcg atgaccgagt
cactagtcgc 1080cctcctcata tcgatattgt agaaaagctc aacggcatta
aggatacgaa attcagaaag 1140agtattttgt cgtggtggaa atctgcgact
gatgtcaacc gtgaagcgat gattgccctg 1200ctttccaata cagtcgatat
cgccaagaaa caggatgacc cggatttctc atccgccgtg 1260gatttcattg
attcgatgga tgatagcgat ttgcagattc tcgataccat cagcattcag
1320cctggtcgtg ctgcgtattc gtcgaagacg ttgcgcgcat tatcaaaacg
catgtattcc 1380acggatgacg atttacacga tgcaagaaag catgtttttg
gtgtagatga ttcatggcgt 1440cctccacagc ctgcaattgg ggcgccattg
ggtaatccgt cagttgatcg agtcgcgaaa 1500attgtcaatc gttggcttct
tgcatgccag tcccggtggg gaaacccgtt gtctatccaa 1560atcgaacatg
tacgagatgc gttgtcttcg gcagcgactg caaccgctga caagcgtgca
1620tatgaacggg cgttaggcaa gcggaacgcc gagaaaatga aagtcaagaa
tgagcttcgg 1680ttgcaaggac tcaatgaacc gcatgaatct gatgtccggc
gtcaagaagc tatcactcgt 1740caacagggca aatgcttgta ctgtggcgat
gatatcacat tcagtacctg tgagatggat 1800cacatcgttc ctcgaaaggg
gcatggctct acaaatacgc gcgataatct cgctgctgta 1860tgtattcagt
gtaatcggca gaagagtaat acgccgtttg ctctatggtg ccagactcca
1920gaggcaaaaa gccgcggcgt gagtcttgaa gctgcaattc atcgcgtgaa
aggtttcttc 1980accgaatcga aggagctaac agggcggcag gcgaaggttt
tcaccagctc gatgataatg 2040cgtttgaagc agacgacagc tgacgatcct
attgatagtc gctcgataga atcggtggca 2100tggatggctg atgagttgca
tcgtcgcatc gattggcatt tcaatggaga tgcctctgaa 2160agcgaccatg
gtcgtagagt gctggttgct gtataccaag ggcgtattac ttctgaagcg
2220cgcaatgtca tgcgtttcca agcgggtggt gatttccact ttgttggcgg
acatggcaaa 2280acacgcctcg atagaagaca ccacgcagtc gatgcgtcgg
tgattgccat gatgactcct 2340gcggcagcac ttacgctggc ggaacgtatc
aatctccgag atagccaacg gtgcattgga 2400cgtataagag agggggagat
tgactggaaa cagtggccca acgagcctac agaaaaatat 2460cagcattggc
ttgataatgg gaaacgcctc ttcgcattga tcaatgatgc gctggataat
2520gaccgtattc caataaccca ttggcagcgc tatgcgttgg gcaattctat
tgcacacgaa 2580gcaacgattc accctctgcg gaagattcca ctgggcagtg
ccattgatta tgagacaatt 2640agtcgagctg caactcctgc cctgtattgc
gccttgacgc gttgcccgga ctattcggtg 2700aatgatggtt tgccggagaa
caaacagagg catattacag tgaatggcaa agtttatggg 2760ccggaagatg
aggtagcgtt ctttgcttct gaccatgtgg agcttgctgt gcaaggtggt
2820tcggcagata ttgggaaaac tatccatcat gctcgggttt accgttgtta
ctttgttgat 2880cgtaggggac agaaaaagtg gttttacggg atgattcgtg
tcttccgagt tgatttgata 2940catgcaaggc atgagaacct attcacttat
cctttgccct ctcaatcaat atcaatgaga 3000tatgcggaaa caagaacggc
tgaggcggtg ttatgcggac atgccgaaca tgtcggtaac 3060ctggtagctg
gagatgaaat cgaagttcca atgattggaa agctgacagg caaaatcgat
3120acgtttgcaa aattctttaa tgaatctttg aataatgaat atgttgcaga
acggtggagc 3180attgatggtt tcgatagtga atcgaagttg ttgcttcgcc
cgctaatgct ggctgaagag 3240ggcatagcga attgggaaga taactctcac
ttatctattc ccgatgatgt caagggactc 3300attgctagag gctggcggcc
atcggtggac acggtatttg cgaagaaacc gcggattgtc 3360cgtcgaaatg
tcctaggaga acccagatgg aagtcgcgtt caggcatgcc cgtttcttgg
3420cgagtcacag gctctgaggc atag 3444763216DNALoktanella
vestfoldensismisc_feature(1)..(3216)Lves Cas9 Open Reading Frame
76atgcgtcttg gttttgatat tggaacgaat tcgattggct ggtggttgta
tgcgaccgac
60ggcaacgaga tcaccggggt gatcgacggg ggcgtgcgga ttttctcgga cgggcgcgat
120cctaaatcca aggcctcttt ggccgttgat cgccgcggcg cccgcgctca
acgccgccgc 180cgtgaccgtt acctgcggcg caaggcggcg ttgatgaaac
gaatggccgc cgccggtttg 240atgcccgccg atcctgtcgc ggcaaaggcg
ttggaacagc ttgatcccta tgctttgcgc 300gcttcgggat tggatcagga
attgcccttg acccatctcg gccgtgcctt gttccacctg 360aaccagcggc
gcggctttaa gtccaaccgc aagacggaca agggcgacaa cgaaagcggc
420aagatcaagg atgcgacagc acgtcttgat caggcgatga tagccaaggg
tgcgcgtact 480tacggcgaat ttctacatat gcggcgcgct tcagccccgg
acccaaaatg cgtgccgact 540gtacgtaccc gactgtccat cgcaccgcgt
gataacgccg aaaaggcaga ggcgggttat 600gacttctacc cggaccggcg
gcatttgttc gaagaattta ccaaactctg ggcggcgcag 660gccgcaaatt
ccccggatat tcttaccgat gagctacgcg atgaaatctc ggtgatcatc
720tttcaccagc gcccgctgaa aacacccgag gtcggcctgt gcctgttctc
aggatctcat 780ggcgttccgc agaatgataa gcgcatcccg agcgcgcatc
cgcagaacca gcgccgtatt 840ctttttgaga cagtgaacaa cctgaaggtg
gctgcacggg gcgaactggc acgcgggctg 900acgcgcgatg agcgggacac
aattgctcat gcgttagata acaaggcgca taccaagtcg 960ctgtccggca
tgtcgatgaa gctcaaggcg cttggcaagc tgatcaagct gcgtcccgaa
1020caatcattca cactggaaac tgccaaccgc gattccatcg tctgcgatcc
ggtgcgggcc 1080agcctgtcgc atcccgaccg gttcgggtcg cgttggtcca
cactggacgc cgaagcacaa 1140tgggatttgg tgcagcgcat ccgcgccgtg
cagagcgatg cagaacataa cgcgttggtc 1200gcatggctga tcgcgacgca
cggacttgac cgcgtgcacg ccgagaacgt ggcaaatgcc 1260ccactgcccg
aaggccacgg ccgtctggga atgacggcga caaagcggat attggcggcg
1320ctggaagccg aagtgatccc ctatagcgcg gcggtggcgg cttgtgggtg
gcaccattcg 1380gatgggcgta ccggcgaggt tctgactgaa ctgccctatt
acggccagat cctcgaccgt 1440cacgtgatcc ccggcacata cgacgagaac
gacgacgaag tgacgcgcta tggccgcatc 1500accaacccga cggttcatat
cggcctgaac cagctgcggc gtttggtgaa caaaattgtc 1560accgtttacg
gcaagcctga tgaaatcgtc gttgagctag cgcgcgatct gaagctgtcc
1620gaagatcaga agcgcgacgt gcagcgcgat atcaagaaga acaccgaagc
tgcgattacc 1680agaggtcaaa agatcgaaga attgggctat gcaaacacag
gtgccaatcg ggtgatgtat 1740cgcctttggg aggaactcgg cccggcaatc
ggcccgcgct gttgcccata ttccggcaag 1800ccgatcagtg cctcgatgat
ctttgacggc tcctgtgatg tcgatcatat ccttccctat 1860tcgcgcacgc
tggaggacgg attttccaac cgtactctgt gcctgaagga gttcaatcgc
1920cagaaaacca acaagactcc atgggaggca tggggcaata ccccggcctg
ggatgccatc 1980gaagccaacc tcaaaaatct gcccgcgaac aaggcttggc
gttttgcccc cgatgcgatg 2040gagcgtttcg agggcgagaa cgatttctct
gcccgcgctc tgaaagacac tcaatacctg 2100tcgcgcatcg cccgcagcta
tctcgatgcg ctttacgatg gggcggacgg caagagccat 2160gtctgggttg
tgcctggccg cctgaccgaa atgctgcgcc gccattgggg gctgaacagc
2220ctgcttcctg ataaggatgg cgcggtcaaa gccaaaaacc gcagcgacca
ccgtcaccac 2280gcgatagatg ctgccgttgt cgcggctacc gaccggtcct
tggtccagcg gatcagtaaa 2340atggcgcaaa gggacgaagt gaatggggcc
gaagaggttg cccgctccgt ccctcctcct 2400tgggacgact ttcgaaccga
tatcaaatcg caactagacc gtatcatcgt cagccaccgc 2460gccgatcacg
gccgaattga ctttgccgca cggcaaacgg gcaacgacag caccagcggc
2520gcgttgcacg aggcgacggc actctcaatt attgacgacc agaacgtcgc
agtccgcatt 2580ccgctcttgt ctttgtccgc agcacagttt gaggaaggcg
ggcggtcagg ctgggttcgt 2640gacccgcagc tacgcggcgc tttgcatttg
gcaaccaagg gcaaggacaa aaaagacttc 2700gaggcggcgc tcctatcatt
cgctgccaag cccgggccat atcacggaat atcaagggta 2760cgtatcgaaa
aacctttgca agatactgcc cgggtctacg tgcccgccga tgcgccgatt
2820aaggcttacc aaggcggtag caatcatcgc tacgaagttt ggaagcttcc
cgatggtaaa 2880gtcctccatc acgttgtttc aatgtttgtt gcccatcagg
gaaacctgac acgcccccat 2940cctgccgcaa aacgaattta tcaattcatg
aagggtgatc tggtgagact tgaggacagc 3000aagtttggac cagtgatcgc
aacggtggag aagttcaacg gaaaggggat gattgagctg 3060gttccgcata
acgaagctaa tgcctcggac cgatatcgca aaaccaagga agatctctat
3120atccgcctcg gcgcaacaac ccttctcagg gccaaagccc gccgtgtcca
tgtcgatgaa 3180atggggcgtc tgcgcgatcc tggcccaccg caatag
3216773318DNASphingomonas sanxanigenensmisc_feature(1)..(3318)Ssan
Cas9 Open Reading Frame 77atggatatgg cgtggcggct cgggctcgat
cttggcacca attcgcttgg ctgggcggcg 60ctgtcgctcg atgcggcggg ggcgccggat
gccattctgg cagccggatc gcgcatcttc 120ggtgatggcc gcgatccgca
gtcgggcacg tcgctggcgg tggatcgccg cgcggcgcgg 180gcggcgcggc
ggcggcggga ccggttcaag caaaggcagc gcgcgctgct gaagcatctc
240gaggcggacg ggctgttccc cgccgatccc gaggtgcagc aggcgctggc
cgcgctcgat 300ccctatgcgt tgcgcgcgcg ggcgctggac gaggcgctga
gcctgcatga gatcggccgg 360gcgctgttcc acctcaacca gcggcggggc
ttccagtcca accgcaaggc ggatcgcggc 420aaggatgagg atgcgggcaa
gatcgcgatc ggcgtcgatc ggctgaagga tgccatcgcg 480gcggcgggcg
cgcggacgtt cggcgaattc ctccgccagc gccgtgcggg cgccaccggc
540gagaatcaga tacccagcgt gcggacgcgg ctgcgcgccg agacgggcga
gggggcgaag 600ggcagcggct atgatttcta tcccagccgc gcgctgctga
aagacgagtt cgacgcgatc 660tggcatgcgc aggcggagca tcatcccaag
gtgctgaccg acgaggccta tcaccgcctc 720cacgagatcg tcttccgcca
gcgcccgttg agggcgccga aggtgggcgc ctgcaccctg 780gtgccgggcg
aagcgcggct gcccaaggcg cacccgctgt tccagcgccg ccggctgctt
840gaggaactga acgcgctgat gatcgtccgc gccggcgcgg tcgcggaacg
gctgacgccg 900gagcagcgcg acctgctgct gctcaagctg aaggacaagg
gtaaggtcac gttcaagagc 960ctgcgcgaga aggtgctgaa gctggacggc
gatgcgcgtt ttaacaagga aagcgagcac 1020cgcaccgagt tgaagggcga
tgaggtcgcc gcggaaatgg gcggcaagac gcgcttcggc 1080tccttgtggc
cgcacctttc cacggagcag caatggacgg tgatcgagcg gcagcaggcg
1140ctggagagcg atgccgacga ggcggcgttc cgcacctggc tggtcgagac
gcacaagctg 1200acagccgagc aggcgagggc ggtggcgggc gcgcggctgc
ccgctgggca tggccgcttc 1260gggctcaccg ccaccgcggc cctgctcgcc
gcgttgcgcg atggccgcac gcccgagggg 1320cgcggcccga atgcgcccgc
gaccgaagcg ggccgggtgg tggtctatag cgaggcggcg 1380gcgatcgccg
gctatcatca cagcgatcac cgcagtggcg aggtcttcac ggacgccaag
1440ggccgcccgg ccttgcccta ttatggcgtg ccgctggatc ggcatatcgt
gccgggcacc 1500gcagaccccg acgagccgga tgaggcggcg cgcatcggcc
ggctgaccaa cccgaccgtg 1560catatcgcgc tcaaccaact ccgtcgcgtc
gtcaaccggc tgatccgcgt ctatggcccg 1620cccgccgagg tcgcgctgga
actggcgcgc gaactcaagc tgtccgagga tgagaagaag 1680gagcgcaacc
gggagaacag ccgcaaccgc ctcgatgccg agaagcggtc ctccaagctc
1740gccgaactcg gccagcgcga caatggcggc aaccgcgcgc tgctcaagct
gtgggaggaa 1800ctgaaccccg agaatatcct cgatcggcgc tgcatctatt
cggggcggca gatctcgatc 1860ggcatgatct tctcgggcgc ggcggaggtg
gaccatatcc tgcctttcga cgccacgctg 1920gatgattcca acgccaacaa
gatcctctgc ctgcgcgagg ccaatcgcga caagcgcaag 1980cgatccccgt
tcgaggcgtg gggcggcacc ccgcaatggg aagagatcgc cgagcgcgct
2040tcccgcctgc cgcgcaacaa gcgctggcgg ttcgagcccg acgcgatgga
gcgcttcgcc 2100gaggagggcg gcttcctcgc ccgccatctc gtcgacacgc
aatatctcgg ccggatcgct 2160catgactatc tgcgctgcct ctatcccgac
aagggagacg gcagcagcca tgtgtgggtt 2220tcgccaggcc ggctgaccga
gatggtgcgc cgcaagctgg ggctgaacgg tctgctcggc 2280gaccataatc
tcggcgccga acagcccaag aaccgcaagg atcatcgcca ccacgcgatc
2340gacgcggtcg tcactgcgat cctcgatcgc tcgatgctcc agcgtatcca
gcgggcttcg 2400ggcgaggggg cgagcgatgc cgagctgctg cgcatcatcg
tgcccgaacc ctggacgggg 2460tttcgggacg acctgcaacg cgcggttgac
cgcatcgtcg tggcgcaccg tgcggaccat 2520ggcacggtgg ccaaggccgc
gacgcgggga cgggaccaga ccgcggcccg gctgcacaac 2580gatactgcct
acggcttcac cggcgaggcg gatgccaagg gcacgccgat tgtggtccat
2640cgggcgccgc tgggggcgct caaaaagccc gagcatatcg atcaggtgcg
cgatccgctg 2700ctgcgcgcgg cattgcacga tttcacggcg gggctgagcg
gcaagccttt cgaggatcgc 2760atcgccgcct tcccccggct tgggccgctc
gactatcgcg gcatccgccg ggtgcgcgtg 2820gtggagccgc tgcaggtgat
cccgatccgc gacgccgccg gccgcgccta caagggctat 2880aaaggcgatt
ccaactatcg ctacgatgtg tgggaactgc ccgacggtaa atgggagcag
2940cgtgtgctgc aaatgtacta cgcgcatcag tccggcgagc caccgcgccc
gcatccagcg 3000gcgcgcaagg ttctgagcct gcaccgggac gatgtgctcg
cgatcgagcg aggggagggg 3060gggcgagaac tcgtccgcgt cgtgaaattc
tcgaccaatg agttcacgct agcgccgctg 3120aatgagggcg gggcgctgaa
gtcgcgtcat gcggacaagg gagatccgtt cagatatctt 3180tatccctcac
cctcgacttt gaaagcgtgg cgcgcacggg aggtgcgcgt ggacgaactc
3240ggccgggtgc tcgatcccgg cttccccgcg cgcaagcggc ggcgggtcac
gcggcccggc 3300cgcgcggacg cggattga 3318784200DNAEpilithonimonas
tenaxmisc_feature(1)..(4200)Eten Cas9 Open Reading Frame
78atgacaaaaa atattcttgg attagacttg ggagtttcat caatcggttg ggcttatgtt
60caggaagatg ataaaaattc tgcgaataat aaaatcatca agttaggagt tcgtgtaaat
120cctttgactg ttgatgaaca aataaatttc gaaaaaggaa aaccgattac
aaccaatgcg 180ggaagaactt tagcaagagg agcaagaaga aatctgcaaa
ggtttaagtt gaggagagct 240aatgttattg atgttttgac aaagggaaat
atcttgaaag atggtgattt gcttacagaa 300gttgggaaaa actctacgtt
tcaaactcag gaattaagag caaaatctgc gaaagaaaaa 360atcgaacttt
cagattttgt gagggtttta cttttaatta ataaaaaaag aggctataaa
420agcagtagaa aagcaaagaa cgaagatgaa ggacaaatca ttgacggaat
ggcggttgct 480aaaaaattat atgaagaaag cctgactcca ggagaatatt
cttaccaact tttaatagaa 540gggaagaagc aattgccaga tttttatcgt
tctgatttac aatctgaatt tgataaagta 600tggaaatttc aaaagcaatt
ttattctgaa atcttaattg atgaacttta taaagaacta 660caagcaaaaa
ataaaaatgc gacgtgggca attttgaaag aaccgttttc tcttgttgga
720atcaaacaaa tgggaactat gcaggagaaa aagattgaaa aatatctttg
gagaagtgaa 780gctgcaaaaa aacaattgga ttttgaaagc ttagctgtcg
tgtttcaaga gattaacagt 840aacctaaata attctagcgg ctatctcggc
gcaataagtg atcgaagtaa agaattgtat 900tttaatcata tgactgttgg
tgaatatctt tatcaacaac ttaaagcaaa ccctcatacc 960aaacttaaaa
atcaagtttt ttatcgacag gattatttgg atgaatttga aaaaatatgg
1020gagacacaat ctcaatatca ttcagaatta accaaggaat taaaagaaca
agttcgtgat 1080gtcgtgattt tctatcagag aaaactcaaa tcccaaaaag
gtttaattag tatttgcgaa 1140tttgagaaca gagaaattga aatcatagaa
aacggaaaaa caaagaagaa aacagtagga 1200ttaaaggttg ctcctaaatc
ttcgccattg ttccaagagt ttaaaatttg gcaagtttta 1260aataatctgc
aatttcaaaa tttagaaaca aaagaaattt tcccaatcga tttagatttc
1320aaacaatcaa tttttgatga agttaatgtc aaaggaagac tttctgcaaa
agaggtttta 1380gatattgtag gttattccgg taaagaatgg aaaactaatt
ttaaagatat tgaaggcaat 1440aataccaatg aaaatctata cactgctttt
ctgaaaatta ttgctaatga aggaaaagaa 1500ttcccaaaag agtttaagtt
aacgattgaa gacgatatta aagttactaa gattcattct 1560tctgcaagta
aaataaaaga atttgtaaaa gagaattttt cgtcattagg aataaataca
1620tcaatattag attttaatcc agaatttgac ggaaaagatt ttgagaagca
aagttcctat 1680cagctttggc atttattgta ttcttacgaa ggcgatgatt
ctgcttctgg taatgaaaaa 1740ttatatgagc ttctggagaa gaaatttagc
tttaaaaaag aacattccaa aatattggca 1800gaaattggtt tctctccaga
ttatggaagt ttgagttcaa aagcgatgcg aaagatttgc 1860acttatatca
aagagcataa atacagcgac gcttgtaatt tagctggtta taatcactct
1920aaaaattcat tgactaaaga acaattagca aatagaattt taaaagaaaa
actagaaatt 1980cttccaaaaa attctttgag aaatcctgtg gttgaaaaga
ttttgaatca aatgataaat 2040gtggtcaatg aagtatccaa agaatacgga
agaccagacg aaataagaat tgagctagca 2100agagaactga aaaaaaatgc
agaggagcgt gcaaatatga cttctgaaat tggcaaagca 2160accttgcttc
atcaaaaata tgcagaaata ttacagaaag aatatgggat aaaagtacca
2220tcaagaaacg atattattcg gtacaaactg tatttggagc tggcaaataa
cggttttaaa 2280gatttataca caggtcaaaa aatagaaaag gaaaacattt
ttaccgacaa atatgatatt 2340gaccatatta tcccgcaatc tcgatttttt
gatgacagtt tttcaaataa agttttggta 2400ccgagaggtg caaatcttaa
aaaaggaaac gcaactgcat ttgattattt ggaaatggaa 2460gggaaagatc
aactggaaaa attcctcaac accatcaaag atttgtttga taaaagtttg
2520atttctaaag ctaaatttga aaaacttcag aaaaaaggaa gtgaaattgg
agatggtttt 2580atacaaagag atttacgaga tacacagtat atagcgaaaa
aagcaaaaga gattcttttc 2640gaaattacca attctgtggt ttctacatca
ggaagaatca cagataaatt acgtgaggat 2700tggaatctcg tcaacacaat
gaaagaactt aatctcgata aatacaggaa attaggttta 2760acggaaacgg
taattaattc taaaggagaa gaaaaagaaa gaattacgga ttggagtaaa
2820agaaatgatc atcgccatca cgcaatggac gctttgacag tagcttttac
gacgcataat 2880catatacaat atttgaatca tttgaatgct agaaaagatg
aaaagcataa tcagcatatc 2940attatttcaa acattgaaaa tcttattaca
aaagtctatg aaaagaaaaa tggttctact 3000aaaagaaagt ttgtagaacc
gattcataat tttagaattg aagccaaaaa gcatttagat 3060gaaatattga
tttctcacaa aacgaaaaat aaggttgtta ccaaaaacat caataaaacc
3120aagaaaaagg gtggagtagt tgctaaagta gtgttgacac cgagaggtca
acttcataaa 3180gaaactattt atggaagctc gaaatttctt aaaactaaag
aagaaaaagt atcggggaaa 3240tttgatttgg aaaccattaa caaagttcag
aatgagaaat ttagaattgc tttattagaa 3300agattaaaag aatttaatgg
agattctaaa aaagcattta caggaaaaaa tgtgttggca 3360aaaaatccga
tttatttgaa tgaagaaaag actgaacaag tttctgaaag tgtaattctg
3420gcttggtacg aaaaggctta taccattagg aaagcagtaa attctgataa
ttttaaagat 3480tataaaaacc ttgaaaaagt aattgataat ggcgtaaaag
aaatcttaaa aaatcgtctg 3540gatgcattta agggtaatgc aaaagaagca
ttttctgatt tagaaaaaaa tccgatttgg 3600ttgaatgagt ccaaaggaat
agccataaaa acagttacca taacgggaat caataatgca 3660gaaactttgc
attacaaaaa agaccatttt ggaaaagaaa ttctagatga aaatggtaaa
3720agaatagccg ttgattttgt aagtactgga aataatcatc acgttgctat
ttatgaagat 3780gcagatggaa atttgcagga aagagtagtg agtttctacg
aagctgttga aagagtgaat 3840caggggcttt caattatcga taaagaatat
aattctggat taggatggaa gtttcttttt 3900acaatgaagc agaatgaaat
gtttttgttt ccatcagaag attttaatcc aaaagaaaat 3960gatttatttg
atgaaaagaa tctgagtttg atttctaaga atatgtttag agtacaaaag
4020tttggggaat tatctaagtc aggtttttgg tttagacatc atttggaaac
atcagtggaa 4080cttaaaaaag aattgagaag tacttcttat ttagattttt
acagtaaaga ttttatgaaa 4140actattgtta aagtaagatt gaatcatctt
ggaaaaatag ttcaaatagg cgaatattaa 4200794362DNASporocytophaga
myxococcoidesmisc_feature(1)..(4362)Smyx Cas9 Open Reading Frame
79atgaaaacag ttcttggtct tgatttaggt acaaactcta ttggttgggc attgattcaa
60catgattttg atagtaaaaa aggtgaaata cttggaatgg gtagccggat aattccaatg
120agtcaggata ttttaggtga atttggaaag ggaaattctg tttctcaaac
agctgacagg 180actaaattca ggagtgcaag aaggctccgg gaacgtcatc
ttttaagacg agagcgttta 240catcgcgttt taaatatcct tggttttctt
cctcgtcatt atgccgctga tattgatttt 300gaaaaacgat tagggcaatt
ttttgaagga aaagaaccta aactagctta tgacaacaat 360cagttcattt
ttacaaaatc ctttggaaaa atgctggctg attttcgtca gcatcagcct
420gattttttaa aggatgaaaa aagtaatgat ttattaatac cttatgattg
gtctatttat 480tatttgcgta aagaagcttt aacaaaaaaa atagagaagg
aagagcttgc atggataatt 540ctcaatttta atcagaagcg tggatactac
caattacggg gagaggagga agaggagaat 600ttaaacaagt tggtagagtt
tcattcttta aagattatag atgtagttgc ggatgaaaaa 660cttaataata
aaggtgagac atggtattcg cttcttcttg aaaatggatg gacatacaga
720cgttcaagta aggttccatt atttgattgg aaagagaaag tcagagattt
tattgttact 780actgatataa atgatgacgg gtctgtaaaa acagataaag
agggaaatga aaaaagaagc 840ttcagagcgc ccagtgatga tgattggact
ttactgaaaa agaaaacgga acaggatatc 900gacaagtcaa ggaaaccaat
tggggcttat atttatgatg caatacttaa aaaccccaag 960caaaaaataa
atggtaagct tgtacgtaca gttgaacgta aattctataa ggaagagctt
1020aaactgattc tacagaagca aaaggaattt catcctgaat tgcagagtac
agatttgtat 1080aatgaatgta ttcgggagtt gtataaaaat aatgatgcac
atcaattaca gttaaataaa 1140aaagattttg ttcatctatt tttggaagat
attatttttt atcagcgtcc gcttcgaagt 1200cagaaatctt ctgttggtaa
ttgtccttta gaattcagaa aatttaaaga tagtgagggg 1260attgagaagg
tagaatattt gaagacaatt cctaagtcca atccatattt tcaggagttc
1320cgcatttggc agtggatgta taatctttca atatacaaga aggatgatga
tgaaaatgtt 1380acccgtgatt ttctgaaaac tatagaagat tgggaggatc
tattcgaatt tcttaatcat 1440cgaaaagata ttgagcaaga gacattgttg
aagtttttgt taggaaagaa tggggtaaaa 1500ggaaaggctc ttaaagtaga
agcagggaaa tttcgttgga attatgtatc agataagatt 1560tatccatgta
atgagaccaa agctcttatt atttctaagc ttgaaaaagt aaaaggtgta
1620gaggagaaat ttttgacaga agagatagag tataagcttt ggcatcttat
ttattctgta 1680acagataaag ttgagtatga aaaggctcta aaggcttttg
ctggtaaaca aaagttggat 1740ttggtttcct ttgtagattc atttaaaagg
tttccaccat ttaaaaatga gtatggtgct 1800tattctgaga aagcaattaa
aaagattctg cccttattac ggacgggaaa atcttggaat 1860tggatggcta
ttgatagcaa agtaagggat agaatcaata aaataattac tggagaattt
1920gatgaagaaa taaagaataa agttcgtgag aaggcggaaa agcatagttt
aaagaaggaa 1980aacgattttc aaggcttacc attatggttg gcgcaatatg
tagtatatgg aagacattca 2040gaagcaagtt tttcaggtaa atggaactcc
gttgatgatt taaaaaagta tctcgaagaa 2100tttaagcaac attcattgag
aaacccgata gtagaacaag tattgacgga gactttacgg 2160gttgttgctg
atatttggca attttatgga aagggagaaa aagatttctt tagtgagatt
2220catattgaac tcggaagaga aatgaaaaat acagctgaag accgtaaaga
aatgagtgcc 2280ataatacagg cgaacgaaac aactaacctt agaataaaag
ctctattaaa tgaactgttg 2340caagacaaga aggtggaaaa tgtaagacct
tattctcctt ctcaacagga aattttaaag 2400atatatgaag atggcgtttt
aagttctgat attgaaattc cagatgatat tcagaaaatc 2460agtaaagcag
cccaaccaac gaaatcagaa cttcagagat ataagttatg gcttgagcaa
2520aaatatcgat ctccatacac tggagcaatg attccattgg gcaagttgtt
tacatctgaa 2580tatgaaattg aacatatcat tcctcagagt ctttattttg
acgatagttt tagtaataaa 2640gtaatttgtg agtcggccgt caataaactt
aaagatagta ggttgggaat ggaatttata 2700aaagaatgcc atgggatggt
tgtggaaaca ggctttggta agtcggttac tgtttttgag 2760gaagaaactt
atagagattt tgtggttcaa aattatagta agaatcattc caaaaaaagt
2820aagcttcttt tggaagaaat tcctgagaag atgatcgaaa ggcagatgaa
tgacactcgt 2880tatataagta agtttatatc atctgtatta tctaatatag
tgcgtgagga ggtcaatgat 2940gatggagtta actcaaaaaa tattgttccg
ggtaatggta aaattacaac acagttaaaa 3000caggactggg gtttaaatga
tatttggaat gaattgattt taccacgctt tgaacgtttg 3060aatgttttga
ctgattcaaa acattttaca gcttggagtg aaaatcatca gaggctattg
3120ccaactgtac cgatagggct ttctaaagga ttttccaaga aaagaattga
tcatagacat 3180catgctttgg atgccttggt tatagcctgt gcctccagaa
atcatatcaa ttttttgaac 3240aatgcacatg caatagataa aaagaaaaat
tcggaggaaa aacaaaaatt tagacatgat 3300ttaaaagcga ttctttgtga
taaaaagtat agtgataaat ccgaaaagaa ttacaggtgg 3360atattcaaga
aaccttggga taattttact attgattcaa aaaacgcttt ggataaaatt
3420attgtaagct ttaaacaaaa tcttcgaatt atcaataagg caacaaaccg
ttatgaaaaa 3480tgggtagata gagatgggat aaaagtaaaa gaatggcata
agcaagaagg aataaactgg 3540gctactagaa agcctttgca taaggataca
gtttcaggta aagttgattt aaaaagggta 3600acagttccag aaggaaagat
attgaccgct accaggaaaa gcttagatac atcgtttgac 3660ttgaaggtca
ttgaatctat tacagatacc ggtattcaaa aaattctaaa aaattatctt
3720actagtaaat caaataatcc ggagctcgca ttttcttcgg agggtgttga
ggatatgaat 3780aagaatatca gaaaatataa tgatggaaaa ttgcatcaac
ctatatataa agttagaata 3840tttgagttgg ggagtaaatt tccattggga
caatttggta ataaaaagtt taagtatgta 3900gaaacagcaa aagggactaa
tcttttcttt gctgtatatg aagatgaaaa taaaaatagg 3960aattatgaaa
cgattccttt aaatgtagtt attgaaaggc aaaaacaagg actctcttct
4020gtgccaataa aaagtgaaaa agggcataaa ttgttattct
atttgtcccc aaatgatata 4080gtctatgttc ctggtagtaa tgaagaattc
agtatagata ggctttatag atttacagat 4140tccagcgata agactgctaa
ttttatacca cttagtgttt caagtttaat ttttagctca 4200aataaaaatg
agcaaaaaaa aatagggatt agctatccaa ttcaagatga gtttggtcta
4260ggaagtccac agtcaaaaaa tcaaaagtcg atcgacggaa taatgattaa
agagaaatgt 4320gtaaaagtga atatagacag gcttggtagg gtgtcatttt ga
4362804530DNAPsychroflexus torquismisc_feature(1)..(4530)Ptor Cas9
Open Reading Frame 80atgaaaagaa ttttagggtt agatttagga acaaactcga
ttgggtggag tctaattgaa 60catgatttta aaaataagca aggacaaatt gaaggattag
gtgttcgtat aattcctatg 120agtcaagaga tactaggtaa atttgatgct
gggcaatcaa tctctcaaac cgcagataga 180acaaaatata gaggcgttag
acgattgtat caaagagaca atcttcgaag agaacgtcta 240catagagtgt
taaaaatttt agactttctt ccaaagcatt atagcgaaag cattgacttt
300caggataaag taggtcaatt caaacctaaa caagaggtta aactcaatta
tcgcaaaaat 360gagaaaaata agcacgaatt tgtttttatg aattcattta
ttgaaatggt aagtgaattc 420aaaaatgccc aaccagagct tttttacaat
aaaggtaatg gagaagaaac aaaaatacca 480tatgactgga cattatacta
tctccgtaaa aaagccttaa ctcaacaaat tactaaagaa 540gaactagcct
ggttaatttt aaatttcaat caaaaacgag gctattatca attacgtgga
600gaagacatag atgaggacaa gaacaagaaa tacatgcaac ttaaagtcaa
caatttgatt 660gattctggtg caaaagtgaa aggaaaagta ttgtataatg
taatttttga taatgggtgg 720aaatatgaaa aacaaattgt caataaagat
gaatgggaag gaaggacaaa ggaatttatt 780attaccacta aaacactgaa
aaatggtaat attaaaagaa catataaagc tgtagactcg 840gaaatagatt
gggctgcgat caaagctaaa actgaacaag acattaataa agcaaataaa
900acagtaggcg aatatattta cgaatctcta ttagacaatc cttctcaaaa
aataagaggg 960aaattggtta aaactataga acgaaaattt tataaagagg
aatttgaaaa gctactttct 1020aaacaaattg aattacaacc tgaactattt
aatgagtcac tgtacaaagc ttgtattaaa 1080gaattatacc ctagaaatga
aaatcatcaa agtaataata aaaaacaagg gtttgagtat 1140ttatttacgg
aggatattat cttttatcaa agaccactta aaagtcaaaa atctaatatt
1200tctggttgtc aatttgagca taaaatttat aagcaaaaaa ataaaaaaac
aggtaagctt 1260gaattaataa aagaacctat aaaaaccatt tccagatcac
atcccttatt tcaggaattt 1320agaatttggc aatggttaca aaatcttaaa
atctataata aagaaaaaat agaaaacgga 1380aagttagaag atgtaacaac
tcaactatta cctaataatg aggcctatgt taccttattt 1440gattttttga
acactaaaaa agaacttgag caaaagcaat ttatcgagta ttttgtaaaa
1500aagaaattaa tagataaaaa agaaaaagag cattttcgtt ggaactttgt
agaagataaa 1560aaatatcctt tttctgaaac cagagcccaa tttttatcac
gcttagccaa agttaaagga 1620ataaaaaata ctgaagattt tttaaataaa
aatacgcaag taggaagcaa agaaaatagt 1680ccatttatta aacgaataga
gcaattatgg cacatcatat attcggtttc tgatctaaaa 1740gagtatgaaa
aagcactcga aaaatttgca gaaaaacaca acttagaaaa ggactccttt
1800ttaaaaaact tcaaaaaatt tcctccattt gttagcgatt atgccagcta
ctctaagaaa 1860gctatatcaa aactattacc aattatgcgc atgggtaaat
attggagtga aagcgccgtt 1920ccaacccaag tgaaagagcg ttccttatca
attatggaaa gagtaaaagt cttaccctta 1980aaagaaggtt actctgacaa
agatttagct gatttgcttt caagagtaag tgacgatgat 2040ataccgaagc
aattaataaa aagttttatt tcttttaaag acaaaaaccc attaaaaggc
2100ttaaatacat accaagccaa ttatttagta tatggtagac actcagaaac
tggagatata 2160caacattgga aaacacctga ggatattgac aggtatttaa
ataatttcaa acagcattcc 2220ctacgcaatc ctattgtaga gcaagtggtt
atggaaacct tgcgtgtagt gagggacatt 2280tgggagcatt atggtaataa
tgaaaaagac ttctttaagg aaatccatgt ggaattaggc 2340agagagatga
aaagccctgc tggtaaacga gaaaaattat cacaaagaaa taccgaaaat
2400gaaaacacta atcatcgtat ccgggaagtt ttaaaagaat tgatgaatga
tgcctctgtt 2460gaaggtggtg tacgcgatta ttctccgagc cagcaggaga
ttcttaaact ttatgaggag 2520ggcatttacc aaaatcccaa caccaattat
ttaaaagttg atgaagatga aatactaaaa 2580atacgcaaaa agaataatcc
tactcaaaaa gagatacaac gctacaaact atggttagaa 2640caaggatata
tctcacctta tacaggaaag attattccct taacaaaact gtttactcac
2700gaatatcaaa ttgaacatat cataccacaa tctaggtatt atgataattc
tctaggaaat 2760aaaatcatct gcgaaagcga agtcaatgaa gataaggata
ataaaacagc ttatgaatat 2820ttaaaggttg aaaaaggaag tattgttttt
gggcataagc ttctcaattt ggacgaatat 2880gaagctcacg tgaacaaata
tttcaaaaaa aataaaacaa aactaaaaaa tttattaagc 2940gaagatatcc
ctgaaggttt tattaaccga caacttaatg atagtagata catcagtaag
3000ctggtaaaag gactattgag taatattgta cgagaaaatg gggaacaaga
agcaacatca 3060aaaaacctta ttccagtaac tggtgtagtc acctcaaaac
taaaacaaga ttgggggctt 3120aatgataaat ggaatgaaat cattgctcct
cgttttaagc gattaaataa actcacaaat 3180tcaaatgatt ttggtttttg
ggataatgac attaacgcct ttagaattca ggttccagat 3240agtcttatca
aaggttttag caaaaaaaga atagaccatc gccaccatgc cttagatgcc
3300ttggtggtag cttgtacttc aagaaatcat acacattact tgagtgcatt
aaatgccgaa 3360aataaaaatt acagcttgcg tgacaaactt gtaatcaaaa
atgagaacgg tgactatact 3420aagacctttc aaataccgtg gcaaggtttt
acaatagaag ctaaaaacaa cttagaaaaa 3480acggttgtaa gctttaaaaa
gaacctgcgt gttattaaca aaaccaataa taaattttgg 3540tcttacaaag
atgaaaacgg aaatctaaac cttggaaaag atggaaaacc taagaagaaa
3600cttcgcaaac agaccaaagg ttataactgg gcgattcgca aacctttgca
taaagaaact 3660gtatcgggaa tttataatat taatgcgcca aaaaataaaa
ttgcaacttc agtaaggact 3720ttattaacag aaatcaaaaa tgaaaaacat
ttagcaaaaa taaccgattt acgtattaga 3780gaaacgattc tacctaatca
tctaaagcat tacctaaaca ataaagggga agcaaatttt 3840agtgaagcat
ttagccaagg gggtattgaa gatttaaaca aaaaaattac gactttaaac
3900gaaggaaaaa aacatcaacc tatttataga gtgaaaatat ttgaagtagg
cagtaagttt 3960tcaatttcag aggatgaaaa ctctgcaaaa agtaagaaat
atgtagaagc tgcaaaaggc 4020accaacttgt tctttgctat ttatttggat
gaggagaata aaaaacggaa ctatgagacc 4080attccgctta atgaagtaat
cacccatcaa aaacaagtag caggttttcc taaatctgaa 4140agattatctg
tacaacctga ttctcaaaaa ggcacattct tattcaccct ttctcctaat
4200gatttagttt atgtaccaaa taatgaggaa cttgaaaatc gtgatttatt
taatttgggg 4260aatttgaatg tcgaacaaat aagtagaatt tacaaattca
ctgactcaag tgataaaact 4320tgtaatttta taccatttca agtatcaaaa
ttgatattta atttaaaaaa aaaggagcaa 4380aaaaaattag atgttgattt
tattattcaa aatgagtttg gcttaggaag tcctcaatct 4440aaaaatcaaa
aatcaattga tgatgttatg ataaaagaaa aatgcattaa actcaaaata
4500gacagattag ggaacatttc aaaagcctag 4530811368PRTLactobacillus
reuteri 81Met Ile Lys Lys Asp Tyr Asn Ile Gly Leu Asp Ile Gly Ala
Thr Ser1 5 10 15Val Gly Phe Ala Gly Ile Asp Glu Gln Tyr Asp Pro Ile
Lys Leu Lys 20 25 30Gly Lys Thr Val Val Gly Val Asn Leu Phe Glu Glu
Gly Gln Thr Ala 35 40 45Ala Asp Arg Arg Ser Phe Arg Thr Thr Arg Arg
Arg Leu Asn Arg Arg 50 55 60Lys Trp Arg Leu Ser Leu Leu Glu Glu Phe
Phe Asp Pro Tyr Ile Thr65 70 75 80Pro Val Asp Pro Ala Phe Phe Ala
Arg Leu Lys Glu Ser Asn Leu Ser 85 90 95Pro Lys Asp Asn Asn Lys Asn
Phe Ser Arg Ser Leu Leu Phe Pro Asp 100 105 110Ile Thr Asp Gln Lys
Phe Tyr Glu Glu Tyr Pro Thr Ile Tyr His Leu 115 120 125Arg Tyr Ala
Leu Met Thr Glu Asn Lys Lys Phe Asp Leu Arg Ala Ile 130 135 140Phe
Leu Ala Ile His His Met Ile Lys Tyr Arg Gly Asn Phe Leu Asn145 150
155 160Ser Thr Pro Val Ala His Phe Asp Thr Ser Lys Ile Asp Phe Ala
Asn 165 170 175Asp Phe Ser Lys Leu Asn Arg Leu Tyr Leu Asn Glu Asp
Pro Asn Asn 180 185 190Ile Phe Glu Ile Asn Leu Gln Asn Val Lys Glu
Ile Ser Asp Ile Leu 195 200 205Leu Asp His Ser Ile Lys Lys Phe Asp
Lys Gln Lys Gln Val Ala Lys 210 215 220Leu Leu Leu Thr Ser Gln Asn
Asp Lys Glu Leu Asp Lys Arg Asn Lys225 230 235 240Gln Ile Ala Thr
Gln Ile Ser Lys Ala Ile Leu Gly Tyr Asn Phe Ser 245 250 255Leu Asn
Glu Ile Leu Lys Leu Glu Ala Val Asn Lys Ser Lys Trp Lys 260 265
270Leu Asn Phe Ser Ser Ala Asp Ile Asp Asp Thr Leu Pro Asp Leu Ile
275 280 285Ser Glu Leu Asp Glu Ser Gln Glu Ser Ile Leu Asn Ile Ile
Leu Ser 290 295 300Leu Tyr Ser Arg Leu Thr Leu Asn Gly Ile Val Pro
Ser Gly Met Ser305 310 315 320Leu Ser Glu Ser Met Ile Asp Lys Tyr
Gly Thr His Lys Glu His Leu 325 330 335Asp Leu Leu Lys Lys Tyr Leu
Lys Thr Leu Pro Ile Lys Asn Arg Lys 340 345 350Glu Ile Ala Glu Ala
Tyr Ala Glu Tyr Val Gly Asn Ser Leu Lys Lys 355 360 365Ser Gly His
Ile Ser Gln Glu Glu Phe Tyr Lys Ala Val Lys Lys Asn 370 375 380Leu
Asp Lys Ser Glu Thr Ala Gln Lys Ile Leu Ser Leu Ile Ser Glu385 390
395 400Glu Lys Phe Met Pro Lys Gln Arg Thr Asn Gln Asn Gly Val Ile
Pro 405 410 415Tyr Gln Leu His Gln Lys Glu Leu Asp Gln Ile Ile Val
Asn Gln Ser 420 425 430Gln Tyr Tyr Pro Trp Leu Ala Glu Leu Asn Pro
Val Thr Glu His Lys 435 440 445Asp Ala Lys Tyr Lys Leu Asp Glu Leu
Ile Ala Phe Arg Val Pro Tyr 450 455 460Tyr Val Gly Pro Leu Ile Asp
Pro Lys Thr Ile Pro Gln Thr Glu Gln465 470 475 480Gly Asn Lys Asn
Ala Ser Phe Ala Trp Met Val Arg Lys Glu Asn Gly 485 490 495Gln Ile
Thr Pro Trp Asn Phe Asp Lys Lys Val Asp Arg Ile Ser Ser 500 505
510Ala Asn Asn Phe Ile Lys Arg Met Thr Thr Lys Asp Thr Tyr Leu Ile
515 520 525Gly Glu Asp Val Leu Pro Ala His Ser Leu Ile Tyr Glu Arg
Phe Lys 530 535 540Val Leu Asn Glu Leu Asn Met Ile Arg Val Asn Gly
Lys Lys Leu Ser545 550 555 560Val Ser Val Lys Gln Asn Leu Tyr Asn
Asp Leu Phe Lys Gln Gln Lys 565 570 575Thr Ile Asn Arg Lys Lys Leu
Ala Asn Tyr Leu Gln Ala Asn Leu Gly 580 585 590Ile Pro Glu Arg Pro
Gln Ile Thr Gly Leu Ser Asp Pro Glu Lys Phe 595 600 605Asn Ser Gln
Leu Ser Ser Tyr Ile Asp Leu Gln Lys Ile Leu Gly Ser 610 615 620Glu
Ile Val Asp Asp Pro Asn Lys Gln Asp Asp Leu Glu Lys Ile Ile625 630
635 640Glu Trp Ser Thr Val Phe Glu Asp Ser Arg Ile Tyr Lys Val Lys
Leu 645 650 655Gln Glu Ile Gly Trp Phe Thr Glu Lys Gln Lys Asn Glu
Leu Val Ser 660 665 670His Arg Tyr Gln Gly Trp Gly Arg Leu Ser Lys
Lys Leu Leu Val Glu 675 680 685Leu Lys Asp Lys Asn Gly Arg Ser Ile
Ile Asp Leu Leu Trp Asn Ser 690 695 700Gln Arg Thr Phe Met Glu Ile
Gln Ser Arg Pro Glu Phe Ala Glu Gln705 710 715 720Ile Thr Asn Glu
Asn Gln Asp Lys Leu Thr Glu Asp Asn Tyr Glu Asp 725 730 735Val Leu
Ala Asp Ala Tyr Thr Ser Pro Gln Asn Lys Lys Ala Ile Arg 740 745
750Gln Val Ile Lys Val Val Asp Asp Ile Val Lys Ala Thr Gly Lys Ala
755 760 765Pro Lys Phe Ile Ser Leu Glu Phe Ala Arg Ser Asp Glu Arg
Ser Asp 770 775 780Arg Val Lys Ser Arg Lys Thr His Ile Gln Lys Ile
Tyr Glu Thr Thr785 790 795 800Ala Lys Glu Leu Leu Lys Asp Asp Gln
Leu Ile Lys Glu Leu Gly Ser 805 810 815Val Ser Asp Leu Ser Asp Arg
Leu Tyr Leu Tyr Phe Thr Gln Leu Gly 820 825 830Arg Asp Met Tyr Thr
Gly Lys Pro Ile Asn Ile Asp Glu Ile Ser Thr 835 840 845Met Tyr Asp
Ile Asp His Ile Leu Pro Gln Ala Phe Leu Lys Asp Asp 850 855 860Ser
Leu Asp Asn Arg Val Leu Val Arg Arg Gln Asp Asn Asn Ala Lys865 870
875 880Ser Asp Thr Val Pro Ala Leu Lys Phe Gly Lys Met Lys Pro Phe
Trp 885 890 895Asn Lys Leu Gln Lys His Gly Leu Ile Ser Lys Arg Lys
Leu Asn Asn 900 905 910Leu Gln Thr Asn Pro Glu Ser Ile Asp Lys Phe
Lys Ala Val Gly Phe 915 920 925Val Asn Arg Gln Leu Val Glu Thr Arg
Gln Val Ile Lys Leu Ala Ala 930 935 940Asn Ile Leu Ala Ser Arg Tyr
Pro Asp Ser Lys Ile Ile Glu Val Lys945 950 955 960Ala Ser Leu Thr
His Gln Met Arg Glu Ser Phe Asn Leu Ile Lys Asn 965 970 975Arg Asp
Val Asn Asp Tyr His His Ala Val Asp Ala Tyr Leu Ser Ala 980 985
990Phe Val Gly Gln Tyr Leu Tyr Asn Arg Tyr Pro Lys Leu Gln Pro Tyr
995 1000 1005Phe Val Tyr Gly Gln Phe Lys Lys Phe Asp Lys Gln Ser
Thr Arg 1010 1015 1020Ile Gly Met Lys Thr Asn His Phe Asn Phe Leu
Tyr Asp Leu Glu 1025 1030 1035Pro Glu Gly Lys Asn Val Lys Ile Lys
Lys Pro Thr Lys Ile Ile 1040 1045 1050Asn Lys Glu Thr Gly Glu Ile
Ile Gly Asp Arg Asp Glu Leu Val 1055 1060 1065Ala Lys Leu Asn Arg
Val Tyr Asn Phe Lys Tyr Met Leu Val Ser 1070 1075 1080Gln Glu Val
Tyr Thr Arg Ser Gly Ala Leu Phe Asp Gln Thr Ile 1085 1090 1095Tyr
Pro Ala Asn Ser Gly Lys Lys Leu Ile Pro Leu Lys Gln Asn 1100 1105
1110Lys Thr Thr Ala Ile Tyr Gly Gly Tyr Ser Gly Ser Lys Ala Ala
1115 1120 1125Tyr Met Ser Ile Ile Arg Leu Arg Asp Lys Lys Gly Gly
Thr Tyr 1130 1135 1140Arg Ile Val Gly Ile Pro Val Arg Ala Val Asn
Lys Leu Asn Gln 1145 1150 1155Ala Lys Lys Lys Ser Asn Glu Lys Tyr
Leu Ala Glu Leu Lys Ala 1160 1165 1170Val Ile Glu Pro Gln Ile Ala
Lys Thr Lys Lys Asp Arg Lys Thr 1175 1180 1185Gly Gln Arg Val Leu
Val Pro Gln Glu Phe Asp Val Ile Ile Pro 1190 1195 1200Glu Val Met
Tyr Arg Gln Leu Ile Val Asp Gly Asp Gln Lys Phe 1205 1210 1215Thr
Leu Gly Gly Thr Ile Asp Arg Tyr Asn Ala Val Gln Leu Val 1220 1225
1230Leu Asn Gln Glu Ile Leu Thr Phe Leu Glu Gln Pro Thr Lys Tyr
1235 1240 1245Lys Asp Ala Asp Thr Lys Leu Leu Asp Ile Tyr Asp Gln
Ile Val 1250 1255 1260Asn Leu Val Glu Lys Tyr Phe Met Leu Phe Asp
Ser Lys Arg Leu 1265 1270 1275Ala Ala Gly Arg Val Ala Phe Glu Lys
Leu Pro Thr Leu Gln Pro 1280 1285 1290Val Asp Lys Met Pro Ser Lys
Leu Ile Ile Ile Arg Arg Ile Ile 1295 1300 1305Gln Gly Leu His Asp
Asn Ala Ala Arg Thr Asp Leu Lys Ala Ile 1310 1315 1320Asn Gly Ser
Ser Ser Phe Gly Arg Leu Gln Lys Arg Asn Gly Ile 1325 1330 1335Ile
Leu Ser Pro Asn Ala Cys Leu Ile Tyr Gln Ser Pro Thr Gly 1340 1345
1350Leu Phe Glu Arg Lys Val Tyr Leu Asn Thr Ile Ser Pro Leu Lys
1355 1360 1365821369PRTLactobacillus rossiae 82Met Asp Lys Ser Lys
Pro Tyr Gly Ile Gly Leu Asp Ile Gly Thr Asn1 5 10 15Ser Val Gly Phe
Val Ala Thr Asp Ala Glu Gly His Leu Ile Arg Leu 20 25 30Lys Gly Lys
Thr Val Ile Gly Ala Tyr Leu Phe Asn Ala Gly Ile Ser 35 40 45Ala Ala
Glu Arg Arg Gly Phe Arg Thr Thr Arg Arg Arg Leu Ser Arg 50 55 60Val
Lys Trp Arg Leu Gly Leu Leu Arg Glu Ile Phe Glu Thr His Phe65 70 75
80Gln Glu Ser Met Gly Glu Asn Glu Asp Asn Asp Phe Phe Leu Arg Phe
85 90 95Lys Tyr Ser Asn Ile Ser Pro Lys Asp Pro Gln Phe Ser Thr Ala
Lys 100 105 110Gly Leu Phe Asn Asp Arg Thr Asp Lys Glu Phe Tyr Asp
Gln Tyr Pro 115 120 125Thr Ile Tyr His Leu Arg Arg Ala Leu Met Thr
Glu Asp His Gln Phe 130 135 140Asp Ile Arg Glu Ile Tyr Ile Ala Met
His His Ile Val Lys Tyr Arg145 150 155 160Gly His Phe Leu Lys Glu
Gly Arg Ala Lys Asp Phe Lys Val Gly Asp 165 170 175Leu Arg Leu Leu
Asp Asn Phe Lys Met Met Asn Glu Gln Ile Glu Glu 180 185 190Ile Asn
Pro Leu Trp Gln Leu Lys Leu Pro Thr Asp Asp Ala Ser Ile 195 200
205Lys Ser Ile Thr Ala Ile Leu Leu Asp Asn Thr Gln Ser Gln Asn Asp
210 215 220Arg Gln Lys Ala Val Thr Lys Val Ile Leu Ala Thr Leu Val
Lys Ala225 230 235 240Ser Asp Lys Asp Ile Asn Ala Ala Arg Lys Arg
Phe Val Gly Glu Leu 245 250 255Ser Lys Ala Met Val Gly Leu Lys Thr
Lys Leu Trp Val Leu Ala Asp 260 265
270Val Ser Gln Asn Gly Asp Trp Glu Ile Lys Tyr Glu Asn Tyr Ala Asp
275 280 285Phe Ala Glu Thr Ile Gly Ser Gly Glu Ser Asp Thr Ile Gln
Ser Leu 290 295 300Phe Asn Glu Ile Asn Asp Leu Tyr Gly Val Ile Thr
Leu Ala Gly Ile305 310 315 320Ile Pro Lys Glu Ala Glu Ser Phe Ser
Asp Gly Met Val Arg Lys Tyr 325 330 335Glu His His Arg Lys Asn Leu
Glu Leu Leu Lys Val Tyr Cys Ala Glu 340 345 350Gln Ser Asp Gly Lys
Arg Gly Arg Gln Ile Arg Gln Thr Tyr Asp Lys 355 360 365Tyr Ile Asp
Gly Val Asp Ser Lys Gln Phe Thr Gln Glu Asp Phe Tyr 370 375 380Lys
Ala Leu Ser Lys Phe Thr Ala Lys Asp Glu Ala Thr Ser Glu Asn385 390
395 400Ala Lys Leu Ile Ala Gln Glu Ile Ala Val Gly Thr Phe Met Pro
Lys 405 410 415Leu Arg Thr Lys Ala Asn Gly Thr Ile Pro His Gln Leu
His Gln Lys 420 425 430Glu Leu Asp Ala Ile Ile Glu Asn Gln Lys Lys
Tyr Tyr Pro Trp Leu 435 440 445Gly Glu Val Asn Pro Val Glu Ser His
Arg Arg Ala Leu Pro Tyr Lys 450 455 460Leu Asp Glu Leu Val Ser Phe
Arg Ile Pro Tyr Tyr Val Gly Pro Met465 470 475 480Val Thr Pro Thr
Lys Gly Asp Pro Glu Lys Ser Lys Phe Ala Trp Met 485 490 495Val Arg
Lys Glu Pro Gly Thr Ile Thr Pro Trp Asn Phe Asp Gln Lys 500 505
510Val Asp Arg Ser Ala Ser Gly Glu Ala Phe Ile Gln Arg Met Lys Thr
515 520 525Thr Asp Thr Phe Leu Ile Gly Glu Asp Val Leu Pro Gln Gln
Ser Leu 530 535 540Leu Tyr Gln Lys Phe Glu Val Leu Asn Glu Leu Asn
Lys Ile Met Ile545 550 555 560Asn Gly Lys Pro Ile Cys Arg Glu Gln
Lys Gln Arg Leu Phe Lys Gln 565 570 575Leu Phe Met Gln Tyr Lys Thr
Val Thr Val Lys Lys Val Gln Gln Asn 580 585 590Leu Ile Ala Asn Gly
Glu Glu Ser Glu Asn Val Pro Ile Thr Gly Leu 595 600 605Ser Asp Pro
Leu Arg Phe Asn Ser Ser Phe Ser Thr Tyr Ile Asp Tyr 610 615 620Lys
Asp Ile Leu Gly Thr Ala Ala Val Asn Asp Asn Ala Lys Gln Ser625 630
635 640Asp Ile Glu Gln Ile Ile Ala Trp Ser Thr Ile Phe Glu Asp Ala
Ala 645 650 655Ile Phe Arg Glu Lys Leu Asn Asp Ile Thr Trp Leu Asn
Asp Asp Gln 660 665 670Arg Asn Lys Leu Ser His Lys Arg Tyr Arg Gly
Trp Gly Arg His Ser 675 680 685Arg Lys Leu Leu Ala Gly Leu Arg Asp
Gly Glu Gly Gln Thr Ile Ile 690 695 700Glu Arg Leu Trp Asn Thr Asn
Asp Asn Phe Met Gln Ile Gln Asn Asp705 710 715 720Ser Glu Ile Ala
Arg Gln Ile Thr Glu Ala Asn Ser Ser Lys Met Ala 725 730 735Thr Ala
Glu Gly Thr Asp Glu Ile Ile Asp Gly Phe Tyr Thr Ser Pro 740 745
750Glu Asn Lys Lys Ala Leu Arg Glu Val Met Lys Val Val Lys Asp Ile
755 760 765Gln Arg Ala His His Gly Gln Ala Pro Ala Trp Val Tyr Ile
Glu Ser 770 775 780Pro Arg Glu Thr Pro Arg Pro Gly Gln Arg Thr Val
Ser Arg Glu Gln785 790 795 800Gln Leu Thr Asp Leu Tyr Glu Gly Ala
Ala Lys Glu Ile Val Asp Asp 805 810 815Ala Val Leu Asn Glu Leu Lys
Asp Lys Val Lys Ser Lys Glu Asn Phe 820 825 830Thr Asp Lys Leu Val
Leu Tyr Phe Leu Gln Asn Gly His Asp Ile Tyr 835 840 845Ala Asn Asp
Ser Ile Asn Ile Asp Asn Leu Asn Ala Tyr Asp Ile Asp 850 855 860His
Val Leu Pro Gln Ser Leu Ile Lys Asp Asp Ser Leu Asp Asn Arg865 870
875 880Val Leu Thr Thr His Glu Arg Asn Leu Lys Lys Ser Asn Arg Phe
Ala 885 890 895Thr Glu Leu Phe Ala Asp Gln Arg Lys Lys Trp Glu Lys
Trp His Arg 900 905 910Leu Gly Leu Ile Ser Ser Arg Lys Leu Lys His
Leu Thr Met Gln Pro 915 920 925Asn Ser Val Glu Lys Phe Ala His Gly
Phe Ile Ala Arg Gln Leu Thr 930 935 940Glu Thr Arg Gln Ile Ile His
Leu Thr Ala Asn Val Leu Ser Asn Leu945 950 955 960Tyr Gln Glu Asn
Asp Thr Lys Ile Val Met Ile Lys Ala Gly Leu Asn 965 970 975Ser Glu
Phe Arg Arg Thr Phe Asp Phe Pro Lys Asn Arg Ser Val Asn 980 985
990Asp Tyr His His Ala Phe Asp Ala Phe Leu Thr Ala Lys Ile Gly Arg
995 1000 1005Tyr Leu Leu Ala Arg Tyr Pro Lys Leu Glu Pro Phe Phe
Val Tyr 1010 1015 1020Gly Asn Phe Val Lys Asn Pro Lys Ala Met Lys
Arg Leu Ser Ser 1025 1030 1035Phe Asp Phe Ile Ala Gln Leu Ala Ala
Lys Thr Asp Asp Thr Ser 1040 1045 1050His Ile Asp Gln Arg Ser Leu
Lys Gln Val Pro Val Val Asn Glu 1055 1060 1065Glu Thr Gly Glu Ile
Val Trp Asp Lys Asp Ile Glu Leu Ala Glu 1070 1075 1080Leu Asp Lys
Thr Tyr Asn Tyr Lys Thr Met Leu Val Lys Arg Ala 1085 1090 1095Gln
Thr Glu Asn Asn Ala Gln Met Phe Lys Gln Thr Val Phe Lys 1100 1105
1110Ala Arg Asp Asn Gln Asn Lys Thr Leu Ile Pro Val Lys Asn Gly
1115 1120 1125Leu Ser Thr Asp Val Tyr Gly Gly His Ser Gln Gln Ala
Ile Ser 1130 1135 1140Tyr Leu Cys Ile Val Trp Val Gly Gln Lys Lys
Lys Tyr Arg Val 1145 1150 1155Leu Gly Ile Ser Thr Ala His Ala Gly
Ile Leu Asn Asn Phe Glu 1160 1165 1170Lys Asn Tyr Gly Arg Phe Glu
Ala Lys Lys Lys Leu Gln Glu Ile 1175 1180 1185Val Ser Asn Thr Leu
Asp Asn Ala Asp Arg Asn Asp Phe Lys Ile 1190 1195 1200Val Ala Pro
Lys Val Leu Phe Glu Gln Val Val Glu Asp Asp Asn 1205 1210 1215Met
Lys Phe Gly Leu Gly Ser Ala Ser Asp Tyr Arg Asn Val Gln 1220 1225
1230Gln Leu Phe Leu Ser Arg Lys Asn Gln Leu Leu Leu Ala Asn Met
1235 1240 1245Met Thr Asp Gln Ile His Asp Gln Asp Leu Val His Leu
Phe Asp 1250 1255 1260Glu Ile Val Gly Gln Met Asn Ala His Phe Pro
Ile Phe Asp Arg 1265 1270 1275Gly Gly Tyr Arg Ser Ser Leu Thr Gln
Ser Arg Asp Lys Phe Leu 1280 1285 1290Lys Leu Pro Phe Lys Lys Asn
Glu Asp Leu Ile Thr Lys Gln Glu 1295 1300 1305Val Ile Arg Arg Ile
Leu Asp Gly Leu His Ala Asn Ala Asn Arg 1310 1315 1320Lys Asp Leu
Lys Ile Ile Gly Ser Lys Gly Asp Phe Gly Arg Leu 1325 1330 1335Gly
Thr Lys Lys Ile Tyr Leu Ser Lys Asp Ala Lys Leu Ile Tyr 1340 1345
1350Thr Ser Pro Thr Cys Leu Phe Thr Arg Thr Val Pro Leu Ser Ser
1355 1360 1365Leu831346PRTPediococcus pentosaceus 83Met Glu Lys Val
Pro Tyr Asn Ile Gly Leu Asp Ile Gly Thr Ser Ser1 5 10 15Ile Gly Phe
Ala Ala Thr Asp Asn Leu Asn Lys Pro Ile Arg Ala Lys 20 25 30Gly Lys
Thr Val Ile Gly Val Arg Leu Phe Glu Glu Gly Lys Thr Ala 35 40 45Ala
Asp Arg Arg Gly Phe Arg Thr Thr Arg Arg Arg Leu Ser Arg Arg 50 55
60Lys Trp Arg Leu Arg Leu Leu Asp Glu Ile Phe Asp Lys Glu Met Ala65
70 75 80Lys Val Asp Asn Thr Phe Phe Ala Arg Leu Lys Glu Ser Asn Leu
Ser 85 90 95Pro Lys Asp Ala Asn Lys Lys Tyr Leu Gly Ser Leu Leu Phe
Pro Glu 100 105 110Lys Lys Asp Phe Lys Phe Tyr Glu Asp Tyr Pro Thr
Ile Tyr His Leu 115 120 125Arg Tyr Ala Leu Met His Glu Lys Arg Gln
Phe Asp Ile Arg Glu Val 130 135 140Tyr Leu Ala Met His His Ile Ile
Lys Tyr Arg Gly Asn Phe Leu Asn145 150 155 160Ser Ala Pro Met Asn
Ser Phe Lys Thr Gln Asp Phe Asp Phe Val Ala 165 170 175Lys Phe Glu
Lys Leu Asn Glu Leu Phe Glu Ser Ile Asp Ala Glu His 180 185 190Glu
Thr Lys Phe Asp Ile Glu Asn Ile Ser Lys Phe Arg Asp Ile Met 195 200
205Leu Asn Gln Asp Ile Arg Lys Leu Asp Arg Lys Lys Gln Ala Ala Lys
210 215 220Ile Leu Ile Leu Asp Ser Thr Asp Lys Thr Ala Lys Lys Ile
Asn Asn225 230 235 240Lys Ile Ala Thr Ala Val Ala Asn Ser Ala Leu
Gly Tyr Lys Phe Ala 245 250 255Leu Asp Ala Ile Leu Lys Leu Asp Val
Glu Glu Ser Lys Asp Trp Ser 260 265 270Ile Ser Leu Asn Asp Glu Glu
Ile Asp Ser Ile Leu Asp Asn Leu Thr 275 280 285Ser Asp Leu Asp Ala
Glu Arg Ile Glu Ile Ile Glu Ile Leu Arg Asp 290 295 300Leu Tyr Ser
His Ile Ala Leu Asn Glu Ile Val Pro Asn Gly Gln Ser305 310 315
320Leu Ser Lys Ser Met Met Asp Lys Tyr Asp Lys His His Ala Asp Leu
325 330 335Asp Val Leu Lys Lys Val Ile Ser Asn Met Asp Asp Arg Lys
Lys Ala 340 345 350Lys Ser Leu Lys Asn Ile Tyr Asn Gln Tyr Val Gly
Lys Thr Asn Asp 355 360 365Lys Val Leu Asp Lys Asp Glu Phe Tyr Lys
Gln Ile Gln Lys Asn Leu 370 375 380Asp Glu Ser Glu Asp Ala Met Lys
Ile Val Asn Glu Ile Glu Leu Asp385 390 395 400Gln Phe Met Pro Lys
Gln Arg Thr Ser Gln Asn Gly Val Ile Pro His 405 410 415Gln Leu His
Gln Lys Glu Leu Asp Glu Ile Ile Glu Asn Gln Lys Gln 420 425 430Tyr
Tyr Pro Phe Leu Ala Glu Pro Asn Pro Asn Glu Lys Arg Lys Pro 435 440
445His Ala Gln Phe Lys Leu Asp Glu Leu Ile Ala Phe Lys Ile Pro Tyr
450 455 460Tyr Val Gly Pro Leu Ile Thr Lys Glu Glu Gln Gln Ala Gln
Ser Gly465 470 475 480Ala Lys Phe Ala Trp Met Lys Arg Lys Gln Asp
Gly Val Ile Thr Pro 485 490 495Trp Asn Phe Asp Glu Lys Val Asp Arg
Met Ala Ser Ala Asn Glu Phe 500 505 510Ile Arg Arg Met Thr Thr Lys
Asp Thr Tyr Leu Leu Gly Glu Asp Val 515 520 525Leu Pro Asp Glu Ser
Leu Ile Tyr Gln Lys Phe Lys Val Leu Asn Glu 530 535 540Leu Asn Asn
Val Lys Val Asn Asp Lys Lys Leu Thr Val Ser Asp Lys545 550 555
560Gln Asp Ile Phe Asn Asp Leu Phe Lys Lys Gln Lys Thr Val Ser Val
565 570 575Ser Lys Leu Gln Lys Tyr Phe Val Thr Glu Lys His Tyr Leu
Thr Glu 580 585 590Pro Thr Ile Lys Gly Leu Ser Asp Thr Lys Lys Phe
Ser Asn Ser Leu 595 600 605Ser Thr Tyr Ile Asp Phe Glu Lys Ile Phe
Gly Asn Glu Ile Leu Ala 610 615 620Asp Gln Asn Lys Gln Asn Asp Leu
Glu Lys Ile Ile Glu Trp Ser Thr625 630 635 640Ile Phe Glu Asp Arg
Lys Ile Phe Glu Asp Lys Leu His Glu Ile Glu 645 650 655Trp Leu Thr
Glu Lys Gln Ile Lys Ala Val Arg Arg Tyr Arg Gly Trp 660 665 670Gly
Arg Leu Ser Lys Lys Leu Leu Val Asp Leu Arg Asn Asn Glu Gly 675 680
685Lys Ser Ile Leu Asp Glu Leu Trp Arg Thr Asn Asp Asn Phe Met Gln
690 695 700Ile Gln Ala Arg Glu Glu Phe Ala Lys Ser Ile Val Glu Ala
Asn Gln705 710 715 720Lys Leu Met Asn Leu Gly Gly Ala Ala Ser Val
Gln Asn Thr Val Glu 725 730 735Ser Thr Leu Glu Asp Ala Tyr Thr Ser
Pro Gln Asn Lys Lys Ala Ile 740 745 750Arg Gln Val Ile Lys Val Val
Glu Asp Ile Val Lys Ala Val Gly Tyr 755 760 765Ala Pro Glu Lys Ile
Thr Ile Glu Phe Thr Arg Gly Ala Asp Lys Asn 770 775 780Pro Arg Arg
Thr Gln Asn Arg Gln Lys Gln Ile Met Glu Val Tyr Lys785 790 795
800Thr Ala Ala Lys Glu Ile Val Asp Ala Thr Leu Lys Gly Gln Leu Glu
805 810 815Asn Glu Glu Lys Leu Thr Asp Lys Leu Tyr Leu Tyr Phe Thr
Gln Leu 820 825 830Gly Lys Asp Ile Tyr Ser Gly Glu Thr Ile Asn Ile
Asp Gln Leu Asn 835 840 845Asn Tyr Asp Ile Asp His Ile Leu Pro Gln
Ala Phe Ile Lys Asp Asp 850 855 860Ser Leu Asp Asn Arg Val Leu Thr
Ser Arg Asp Leu Asn Asn Gly Lys865 870 875 880Ser Asp Ser Val Pro
Val Lys Gln Phe Gly Ala Asn Met Lys Ser Phe 885 890 895Trp Met Arg
Leu Gln Ala His Gly Leu Ile Ser Lys Arg Lys Leu Asn 900 905 910Asn
Leu Met Thr Asp Pro Asp Ser Ile Gly Lys Tyr Thr Met Gln Gly 915 920
925Phe Val Arg Arg Gln Leu Val Glu Thr Ser Gln Val Ile Lys Leu Thr
930 935 940Ala Asn Ile Leu Gly Ala Ile Tyr Gly Glu Asn Thr Asp Ile
Val Glu945 950 955 960Ile Pro Ala Lys Leu Thr His Gln Met Arg Glu
Lys Phe Asn Leu Tyr 965 970 975Lys Val Arg Glu Val Asn Asp Tyr His
His Ala Phe Asp Ala Tyr Leu 980 985 990Thr Thr Phe Val Gly Asn Tyr
Leu Phe Lys Arg Tyr Pro Lys Leu Arg 995 1000 1005Pro Tyr Phe Val
Tyr Gly Asp Phe Lys Met Thr Asp Asn Ala Leu 1010 1015 1020Lys Gly
Met Arg Arg Phe Asn Phe Leu His Asp Leu Lys Asp Asp 1025 1030
1035Glu Val Leu Val Asp Asn Glu Thr Gly Glu Val Leu Trp Glu Gly
1040 1045 1050Gln Lys Ser Ile Glu Glu Leu Lys Lys Ile Tyr Gly Tyr
Lys Phe 1055 1060 1065Met Leu Thr Thr His Glu Ala Tyr Thr Gln His
Gly Pro Met Phe 1070 1075 1080Lys Gln Thr Val Tyr Ser Ser Asp Thr
Pro Gly Lys Leu Ile Lys 1085 1090 1095Ile Lys Asn Asn Lys Pro Thr
Glu Ile Tyr Gly Gly Tyr Thr Ser 1100 1105 1110Asn Thr Asp Ala Tyr
Met Ala Ile Val Arg Ile Lys Ala Arg Lys 1115 1120 1125Gly Asp Thr
Tyr Lys Val Val Gly Val Pro Arg Lys Glu Gly Asp 1130 1135 1140Ala
Leu Ser Arg Ile Lys Leu Leu Asp Glu Ser Lys Tyr His Glu 1145 1150
1155Asn Leu Lys Asn Ile Leu Ala Ile Asn Leu Asn Lys Ser Leu Asn
1160 1165 1170Lys Phe Asp Val Val Leu Asp Lys Val Lys Tyr Arg Gln
Val Ile 1175 1180 1185Tyr Asp Gly Thr Asp His Leu Met Leu Gly Ser
Ser Lys Tyr Lys 1190 1195 1200Tyr Ser Thr Lys Gln Leu Val Leu Ser
Asp Gln Ser Met Lys Ile 1205 1210 1215Leu Ser Ser Thr Gly Glu Phe
Ser Asp Glu Glu Leu Ile Lys Val 1220 1225 1230Phe Asp Glu Ile Met
Tyr Ile Val Asn Lys Asn Phe Ser Leu Tyr 1235 1240 1245Asp Thr Arg
Gly Phe Arg Asp Lys Leu Asn Asn Ala Arg Glu Asn 1250 1255 1260Phe
Ile Lys Leu Pro Asn Lys Thr Leu Phe Glu Lys Gly Lys Leu 1265 1270
1275Lys Gln His Ser Lys Leu Glu Ile Leu Lys Gln Ile Leu Ile Gly
1280 1285 1290Leu His Ala Asn Ala Gly Arg Gly Asp Leu Lys Asp Ile
Gly Val 1295 1300 1305Asn Asn Phe Gly Ala Met Val Val Thr Ala Gly
Val Thr Leu Ser 1310 1315 1320Pro Asp Ala Thr Ile Val Tyr Gln Ser
Pro Thr Gly Leu Phe Glu 1325 1330 1335Arg Lys Val Lys Leu Ser Asp
Leu 1340 1345841130PRTLactobacillus nodensis 84Met Glu Asn Lys Ile
Ser Leu Gly Leu Asp Ile Gly Val Ala Ser
Val1 5 10 15Gly Phe Ser Val Ile Asp Ala Gln Glu Gly Lys Val Leu Glu
Leu Gly 20 25 30Ala Arg Leu Phe Asn Ser Ser Val Ser Ala Glu Asn Gln
Thr Arg Arg 35 40 45Asp Met Arg Gly Ser Arg Arg Leu Ser Asn Arg Lys
Lys Gln Arg Arg 50 55 60Lys Asp Val Ala Gln Leu Phe Lys Thr Phe Gly
Leu Ile Asn Tyr Phe65 70 75 80Asp Lys Glu Asn Tyr Phe Asp Asn Phe
Gln Asn Asn Leu Asn Ser Tyr 85 90 95Glu Leu Arg Val Lys Gly Leu Ser
Glu Lys Leu Ser Lys Glu Glu Leu 100 105 110Val Asn Ser Leu Tyr His
Ile Val Lys Arg Arg Gly Ile Ser Tyr Asp 115 120 125Leu Ala Asp Ala
Asp Thr Asp Phe Asp Gly Ser Asp Tyr Ser Ser Ser 130 135 140Leu Asn
Gln Asn Gln Leu Glu Leu Gln Thr Lys Thr Pro Ala Glu Ile145 150 155
160Gln Leu Thr Arg Leu Asn Val His Gly Ala Val Arg Gly Lys Val Thr
165 170 175Ile Asn Gly Glu Asp Glu Asp Thr Met Gln Val Leu Leu Asn
Val Phe 180 185 190Pro Thr Lys Ser Phe Val Ser Glu Ala Lys Lys Ile
Ile Gln Thr Gln 195 200 205Gln Gln Tyr Tyr Pro Asp Ile Leu Thr Asp
Ile Phe Glu Thr Lys Tyr 210 215 220Leu Glu Ile Leu Glu Arg Lys Arg
Glu Tyr Phe Val Gly Pro Gly Ser225 230 235 240Glu Lys Ser Arg Thr
Asp Phe Gly Ile Tyr Lys Lys Asp Gly Arg Thr 245 250 255Leu Asp Asn
Leu Phe Glu Glu Leu Ile Gly His Asp Lys Ile Tyr Pro 260 265 270Asp
Glu Leu Arg Ala Ser Gly Ala Ser Tyr Thr Ala Gln Val Phe Asn 275 280
285Val Leu Asn Asp Leu Asn Asn Leu Arg Ile Ser Ser Tyr Glu Asn Gly
290 295 300Lys Ile Ser Thr Glu Asp Lys Ile Lys Ile Val Glu Asp Leu
Arg Asn305 310 315 320Asn Ile Gly Asn Val Ile Ile Met Lys Ile Ile
Lys Lys Val Ala Gly 325 330 335Cys Glu Asp Asp Asp Ile Lys Gly Tyr
Arg Leu Asp Ser Lys Asp Lys 340 345 350Pro Asp Ile His Ser Met Ala
Val Tyr Arg Lys Val His Arg Asp Leu 355 360 365Leu Lys Tyr Asp Val
Asp Ile Ile Lys Trp Pro Thr Glu Phe Ile Asp 370 375 380Glu Leu Ser
Pro Ile Leu Thr Leu Asn Thr Glu Asn Gly Glu Ile Arg385 390 395
400Lys Gln Met Val Asn Lys Leu Gln Pro Lys Tyr Ser Phe Leu Thr Asp
405 410 415Glu Leu Ile Gln Val Ile Ile Asn Asn Lys Ser Ser Phe Asp
Val Thr 420 425 430Ser Asn Asn Lys Trp His Arg Phe Ser Leu Lys Thr
Met Asn Val Leu 435 440 445Ile Glu Glu Met Phe Arg Arg Pro Val Glu
Gln Met Thr Leu Ile Gln 450 455 460Glu Leu Gly Leu Ile Lys Asp Ser
Gly Lys Arg Phe Glu Asn Cys Lys465 470 475 480Leu Leu Pro Tyr Arg
Glu Ile Ser Lys Asp Ile Phe Asn Pro Val Ala 485 490 495Ser Lys Ser
Val Arg Glu Ala Leu Lys Ile Val Asn Ala Val Met Lys 500 505 510Lys
Tyr Gly Gln Ile Asp Tyr Leu Val Ile Glu Met Pro Arg Asp Lys 515 520
525Asn Glu Glu Glu Met Lys Lys Gln Ile Glu Lys Phe Gln Lys Glu Asn
530 535 540Asn Lys Gln Lys Asp Glu Ala Ile Asn Glu Phe Val Lys Lys
Ile Gly545 550 555 560Asn Lys Asn Ala Val Asp Asp Gly Leu Arg Arg
Tyr Gly Gly Lys Leu 565 570 575Tyr Phe Lys Ile Arg Leu Trp Tyr Gln
Gln Asp Gly Ile Asp Leu Tyr 580 585 590Asn Gly Lys Val Ile Glu Pro
Phe Asp Leu Leu Asn Asn Ile Asn Lys 595 600 605Phe Glu Val Asp His
Ile Ile Pro Glu Ser Ile Ser Phe Asp Asp Ser 610 615 620Ile Asn Asn
Lys Thr Leu Cys Tyr Ala Asp Met Asn Gln Ile Lys Gly625 630 635
640Gln Lys Thr Pro Phe Glu Phe Met Asn Glu Gly His Gly Gln Gly Phe
645 650 655Ala Lys Met Lys Ala Met Val Asn Lys Asn Ser Lys Leu Lys
Gly Lys 660 665 670Arg Lys Asn Tyr Leu Phe Asp Glu Asn Ile Ser Asp
Ile Glu Thr Arg 675 680 685Lys Arg Phe Ile Ser Arg Asn Leu Val Asp
Thr Gln Tyr Ser Ser Arg 690 695 700Val Val Leu Asn Ser Leu Gln Glu
Phe Phe Lys Glu Lys Glu Thr Gly705 710 715 720Thr Lys Val Thr Val
Val Arg Gly Lys Phe Thr Ser Asn Leu Arg Lys 725 730 735His Trp His
Ile Asn Lys Thr Arg Asp Thr Phe His His His Ala Ile 740 745 750Asp
Ala Ser Ile Ile Ala Ala Thr Pro Phe Leu Arg Ile Trp Lys Lys 755 760
765Asp Ala Ser Leu Phe Pro Met His Val Ser Glu Asn Thr Val Asp Ile
770 775 780Glu Thr Gly Glu Ile Leu Asn Asp Thr Glu Phe Lys Lys Asp
Phe Tyr785 790 795 800Gly Leu Pro Tyr Ser Ser Phe Ile Glu Glu Leu
Asn Gly Ala Asp Asp 805 810 815Arg Ile Lys Phe Ser His Gln Val Asp
Arg Lys Met Asn Arg Lys Val 820 825 830Ser Asp Ala Thr Ile Tyr Ser
Thr Arg Lys Gly Met Leu Asn Lys Asp 835 840 845Lys Glu Glu Thr Asp
Tyr Val Leu Gly Lys Ile Lys Asn Ile Tyr Asp 850 855 860Val Ser Glu
Tyr Asn Lys Phe Lys Arg Ile Tyr Asp Lys Asp Pro Asn865 870 875
880Lys Phe Leu Leu Ala His His Asp Pro Lys Ser Phe Asp Glu Leu Arg
885 890 895Lys Ile Met Gln Glu Tyr Pro Ser Lys Ile Asp Lys Val Met
Thr Asn 900 905 910Gly Lys Val Lys Ser Val Asp Ile Ser Pro Phe Glu
Leu Tyr Arg Arg 915 920 925Lys His Gly Met Val Gln Lys Tyr Ser Asn
Lys Asn Lys Gly Pro Val 930 935 940Ile Lys Gln Leu Lys Tyr Leu Asp
Lys Lys Leu Gly Ser His Ile Asp945 950 955 960Ile Thr Pro Lys Asp
Thr Leu Asn Asp Arg His Val Val Leu Gln Ser 965 970 975Leu Lys Pro
Trp Arg Thr Asp Val Tyr Tyr Asn Ser Val Thr Gly Glu 980 985 990Tyr
Glu Ile Met Gly Ile Lys Tyr Ser Asp Leu Lys Phe Asn Gly Gly 995
1000 1005Glu Tyr Gly Ile Lys Met Ser Lys Tyr Leu Glu Ile Lys Glu
Arg 1010 1015 1020Glu Gln Val Ser Asp Glu Ser Glu Phe Leu Phe Thr
Leu Tyr Lys 1025 1030 1035Arg Asp Arg Ile Gln Ile Ile Asn Cys Glu
Asn Asp Glu Ile Val 1040 1045 1050Glu Met Leu Phe Trp Ser Arg Asn
Asn Ser Asn Ile Gly Tyr Ala 1055 1060 1065Glu Leu Lys Pro Val Tyr
Lys Tyr Lys Thr Glu Asp Glu Ser Trp 1070 1075 1080Pro Val Tyr Gly
Tyr Gly Lys Asn Gln Ile Leu Lys Arg Leu Val 1085 1090 1095Pro Lys
Asn Cys Lys Ile Leu Lys Val Asn Thr Asp Ile Leu Gly 1100 1105
1110Asn Pro Tyr Tyr Ile Lys Lys Glu Ser Lys Asn Pro Lys Asn Ile
1115 1120 1125Leu Asp 1130851361PRTSulfurospirillum sp. 85Met Thr
Ser Leu Ile Ser Leu Asp Leu Gly Gly Lys Asn Thr Gly Phe1 5 10 15Phe
Ser Phe Thr Ala Lys Asp Val Ser Ile Ile Asp Thr Phe Gln Ser 20 25
30Gly Thr Ile Ile Tyr Asp Glu Ser Phe Val Leu Ser Gln Val Ala Arg
35 40 45Arg Gly Lys Arg His Gly Lys Arg Asn Asn Leu Arg Asn Thr Leu
Val 50 55 60Lys Arg Leu Phe Leu Leu Leu Leu Gln Lys His Tyr Gly Leu
Ser Leu65 70 75 80Asp Phe Leu Pro Asp Glu Ile Leu Gly Leu Phe Asn
Lys Arg Gly Tyr 85 90 95Thr Tyr Ala Ser Phe Glu Ile Lys Glu Asp Glu
Lys Glu Asn Leu Glu 100 105 110Ser Asp Ile Leu Lys Glu Phe Leu Asn
Asp Lys Leu Asn Tyr Thr Ile 115 120 125Gln Asn Asp Asp Glu Val Glu
Glu Phe Leu Asn Gln Ile Ala Ser Asn 130 135 140Glu Glu Thr Phe Lys
Asn Tyr Lys Lys Asp Phe Glu Asn Leu Phe Gly145 150 155 160Ala Ser
Thr His Gln Pro Lys Lys Gln Ile Glu Leu Ile Asp Glu Ile 165 170
175Lys Lys Asp Leu Glu Lys Glu Asp Ala Lys Glu Leu Leu Asp Gly Leu
180 185 190Lys Val Ile Lys Lys Ile Ile Asp Glu Phe His Lys Gln Gln
Asn Gln 195 200 205Gly Asn Leu Pro Arg Ala Lys Tyr Phe Glu Glu Leu
Tyr Leu Glu Ile 210 215 220Glu Tyr Asn Leu Lys Ile Gln Lys Phe Phe
Thr Cys Asn His Leu His225 230 235 240Ile Asn Asp Met Gln Tyr Leu
Ile Gly Asn Leu Ser Asn Tyr Gln Leu 245 250 255Lys Glu Leu Arg Arg
Tyr Phe Asn Asp Glu Ala Met Ala Lys Glu Asp 260 265 270Phe Trp Ser
Cys Glu Lys Leu His Arg Ile Thr Trp Arg Phe Ile Gln 275 280 285Ser
Trp His Pro Lys Ser Pro Glu Asp Lys Gln Arg Gln Lys Glu Asn 290 295
300Leu Ala Asn Leu Lys Thr Lys Ser Ile Ile Glu Phe Leu Thr Thr
Thr305 310 315 320Asn Pro Leu Met Thr Ile Pro Pro Tyr Asp Asp Met
Asn Asn Arg Gly 325 330 335Ala Val Lys Cys Gln Thr Leu Arg Leu Asn
Glu Glu Tyr Leu Asp Lys 340 345 350His Leu Pro His Trp Arg Ala Ile
Ala His Thr Leu Ala Ser Glu Thr 355 360 365Gln Lys Glu Asn Leu Glu
Gly Val Thr Val Lys Gly Tyr Ser Glu Asp 370 375 380Ser Thr Leu Leu
His Arg Ile Leu Asp Thr Ser Ser Ile Ile Asp Pro385 390 395 400Tyr
Arg Leu Arg Ser Asp Glu Ile Asp Ser Tyr Cys Asp Val Leu Thr 405 410
415Lys Asp Asn Ala Phe Ala Leu Lys Lys Phe Ala Lys Glu Tyr Tyr Gln
420 425 430Leu Val Lys Glu Lys Val Arg Thr Gly Ile Trp Thr Lys Asp
Asp Asp 435 440 445Met Phe Lys Lys Cys Asp His Asn Pro Pro His Lys
Asn Asn Gln Ile 450 455 460His Asn Leu Val Ala Gly Ile Leu Gly Lys
Pro Ile Ala Lys Glu Arg465 470 475 480Phe Glu Ala Phe Glu Asn Glu
Leu Trp Asn Val Lys Phe Gly Asn Lys 485 490 495Lys Leu Ser Ser Tyr
Cys Lys Asn Ile Glu Glu Phe Arg Lys Ser Asn 500 505 510Gly Asn Leu
Phe Lys Gln Ile Val Glu Leu Gly Glu Asp Lys Glu Val 515 520 525Gln
Lys Tyr Gln Lys Glu Leu Asn Glu Trp Val Arg Lys Ile Gly Glu 530 535
540Phe Phe Asn Ile Glu Thr Pro Tyr Arg Ala Arg Phe Asn Asn Leu
Phe545 550 555 560Ser Met Ala Gln Leu His Thr Ile Ile Asp Thr Thr
Arg Ser Gly Phe 565 570 575Asn Ala Thr Cys Lys Trp Cys Ser Cys Glu
Asn Gln Tyr Arg Ala Ser 580 585 590Thr Arg Ile Glu Ile Asp Glu Gln
Thr Gly Glu Ile Thr Thr Asn Ala 595 600 605Asn Cys Gln Arg Leu Pro
Ala Asp Thr Gln Arg Pro Phe Ser Gly Lys 610 615 620Ile Glu Arg Tyr
Ile Asp Lys Leu Gly Tyr Glu Ile Ala Lys Val Lys625 630 635 640Ala
Lys Glu Leu Glu Gly Ile Lys Glu Asp Thr Ile Asp Leu Lys Ile 645 650
655Ile Leu Glu Gln Asn Ala Phe Ala Tyr Glu Glu Ser Ile Arg Asn Ala
660 665 670Lys Ile Lys Asn Ala Asn Ala Lys Ala Lys Lys Ala Leu Glu
Glu Ala 675 680 685Gln Lys Arg Gly Leu Lys Asn Ile Glu Asp Lys Thr
Lys Arg Ile Lys 690 695 700Asp Phe Ser Asn Ser Ile Cys Pro Tyr Cys
Gly Gln Ser Leu Gly Glu705 710 715 720Asp Gly Glu Ile Asp His Ile
Leu Ser Arg Ser Tyr Thr Leu Lys Lys 725 730 735Tyr Asp Thr Val Phe
Asn Ser Glu Gly Asn Leu Leu Tyr Val His Gln 740 745 750Lys Cys Asn
Gln Ala Lys Leu Ala Lys Thr Asp Tyr Ser Leu Gln Asp 755 760 765Leu
Lys Ile Asp Ile Ser Gln Lys Trp Ile Glu Glu Gln Ile Ala Thr 770 775
780Ile Lys Thr Tyr Lys Thr Phe Ser Val Leu Thr Gln Glu Gln Gln
Lys785 790 795 800Ala Phe Lys Tyr Ala Leu Phe Leu Asp Asn Ser Asn
Glu Ala Tyr Gln 805 810 815Lys Val Ile Ser Trp Leu Arg Thr Asp Gln
Ser Ser Arg Val Asn Gly 820 825 830Thr Gln Lys Tyr Leu Ala Lys Lys
Ile Gln Glu Lys Leu Lys Ala Met 835 840 845Phe Pro Ala Lys Thr Phe
Asn Phe Glu Phe Ile Leu Ala Asp Ala Asn 850 855 860Asp Val His Asp
Leu Arg Ile Lys Ala Tyr Gln Leu Pro Glu Lys Pro865 870 875 880Lys
Asp Ser Lys Gln Glu Thr Tyr Ser His Thr Ile Asp Ala Val Met 885 890
895Ser Leu Val Ser Val Trp Asp Lys Val Leu Pro Lys Thr Glu Lys Pro
900 905 910Thr Lys Glu Asp Ile Leu Lys Phe Ala Asn Val Glu Asn Trp
Ser Ala 915 920 925Leu Asn Asn Glu Phe Leu Thr Lys Gly Lys Ser Ala
Asn Gln Lys Ile 930 935 940Glu Glu Met Ile Gln Ala Asn Asp Phe Gly
Gln Lys Asn Met Arg Gln945 950 955 960Val Phe Ser Lys Pro Ile Phe
Lys Asp Glu Ser Ile Gly Glu Arg Tyr 965 970 975Lys Pro Phe Val Arg
Tyr His Asn Gln Phe Tyr Ile Gly Tyr Pro Ile 980 985 990Ser Ile Lys
Asp Gly Tyr Asp Met Gln His Cys Gln Ala Met Ile Ser 995 1000
1005Lys Asn Asp Ile Ser Arg Val Glu Glu Ile Leu Lys Asp Thr Ser
1010 1015 1020Leu Cys Thr Leu Leu Lys Glu Lys Asn Gly Ile Lys Leu
Tyr Ser 1025 1030 1035Ile Asn Lys Gln Ser Ile Asn Glu Leu Ser Asn
Gln Phe Phe Asn 1040 1045 1050Leu Asn Tyr Gln Asn Leu Asn Asp Ala
Gln Lys Lys Lys Ser Glu 1055 1060 1065Leu Ala Glu Phe Val Ile Asn
His Cys Lys Tyr Tyr Val Lys Lys 1070 1075 1080Thr Ser Val Ile Asn
Ala Pro Gln Phe Ile Asp Lys Asp Ser Met 1085 1090 1095Lys Pro Tyr
Pro Phe Tyr Lys Asp Trp Gln Lys Phe His Glu Ala 1100 1105 1110Tyr
Lys Lys Glu Leu Asp Ala Glu Pro Lys Thr Lys Lys Asp Asn 1115 1120
1125Gly Lys Leu Val Tyr Asp Ile Ser Gly Ile Asp Asp Phe Trp Thr
1130 1135 1140Glu Phe Cys Lys Lys Tyr Phe Gly Ile Lys Thr Lys Asp
Asn Arg 1145 1150 1155Asn Lys Ala Arg Lys Val Phe Ser Ile Val Ala
Leu Thr Ser Ala 1160 1165 1170Pro Gly Thr Val Phe Arg Ile Lys Arg
Lys Thr Pro Lys Gly His 1175 1180 1185Ile Tyr Gln Ala Thr Ala Ile
Asp Asn Gln Gln Ile Ser Gly Asp 1190 1195 1200Tyr Ala Asn Val Leu
Leu Ala Gly Asn Ser Lys Thr Leu Ala Leu 1205 1210 1215Ala Gly Gln
Lys Pro Ser Ser Asp Leu Lys Lys Glu Leu Ser Val 1220 1225 1230Lys
Glu Ser Lys Asp Ile Arg Asp Ile Lys Leu Glu Pro Ser Arg 1235 1240
1245Phe Phe Lys Glu Gly Phe Asp Cys Arg Gly Ile Glu Val Ile Val
1250 1255 1260Asn Lys Thr Ser Ala Thr Ile Lys Asn Phe Pro Leu Thr
Lys Ile 1265 1270 1275Asp Lys Lys Ile Lys Lys Leu Ile Phe Lys Thr
Leu Phe Glu Lys 1280 1285 1290Lys Asp Gly Lys Arg Gln Lys Gln Lys
Thr Ser Ile Ser Leu Lys 1295 1300 1305Glu Lys Asn Thr Met Gln Glu
Thr Leu Lys Lys Leu Leu Lys Asp 1310 1315 1320Ser Ile Lys Val Thr
Ile Arg Asp Gly Ser Ile Ser Gly Ile Glu 1325 1330 1335Ile Ser Lys
Lys
Thr Val Asn Phe Thr Leu Pro Phe Lys Ser Glu 1340 1345 1350Asn Leu
Ala Lys Leu Leu Asp Asp 1355 1360861147PRTBifidobacterium
thermophilum 86Met Ser Asp Lys Thr Tyr Arg Ile Gly Ile Asp Val Gly
Leu Tyr Ser1 5 10 15Val Gly Leu Ser Ala Ile Gln Val Asn Asp Asp Asp
Asp Pro Val Arg 20 25 30Ile Leu Asn Ala Gln Ser Val Ile His Asp Gly
Gly Val Asp Pro Asn 35 40 45Ala Gln Lys Ser Ala Asp Ser Arg Arg Ala
Gln Ser Gly Ile Ala Arg 50 55 60Arg Thr Arg Arg Met Arg Arg Asn Arg
Lys Lys Arg Leu Lys Arg Leu65 70 75 80Asp Gln Ile Leu Val Glu Ser
Gly Phe Pro Val Ser Ser Glu Asn Asp 85 90 95Leu Glu Gly Phe Glu Pro
Trp Leu Leu Arg Ala Gln Ala Ala Asp Ala 100 105 110Phe Ile Glu Asp
Glu Asp Ile Arg Lys Arg Ala Ile Ser Val Ser Cys 115 120 125Arg His
Ile Ala Arg His Arg Gly Trp Arg Asn Pro Tyr Leu Asp Val 130 135
140Arg Thr Leu Leu Ala Val Asp Ser Pro Ser Ser Ala Phe Tyr Asp
Lys145 150 155 160Leu Val Glu Asn Ala Ala Leu Glu Met Asp Gly Gln
Met Pro Asp Ser 165 170 175Asp Ala Thr Pro Ala Gln Ile Val Arg Asp
Val Leu Glu Tyr Lys Arg 180 185 190Gly Glu Ala Ala Ile Arg Leu Arg
Lys Ser Thr Ala Glu Asn Lys Lys 195 200 205Asn Arg Leu Ala Leu Phe
Pro Glu Lys Met Met Gln Asp Asp Tyr Ala 210 215 220Tyr Glu Leu Arg
Leu Ile Leu Ala Lys Gln Ala Val Pro Lys Asp Ile225 230 235 240Ala
Arg Lys Leu Ile Leu Ala Val Phe Gln Ser Gln Ser Pro Lys Gly 245 250
255Ser Ala Glu Lys Arg Val Gly Lys Asp Pro Leu Asp Pro Ser Gln Pro
260 265 270Arg Ala Leu Lys Ala Ser Leu Ala Phe Gln Glu Tyr Arg Ile
Leu Asn 275 280 285Ile Leu Thr Asn Leu Arg Leu Gln Asp Gly Gly Ala
Glu Arg Arg Leu 290 295 300Ser Ile Glu Glu Lys Gln Lys Leu Tyr Lys
Met Leu Val Glu Asp Thr305 310 315 320Gly Arg Glu Lys Lys Tyr Glu
Thr Trp Thr Asp Ile Ala Ser Ala Met 325 330 335Glu Trp Lys Arg Asn
Trp Leu Lys Gly Val Gly Ser Leu Thr Ala Asp 340 345 350Gly Asp Asp
Arg Val Thr Ser Arg Pro Pro His Ile Asp Ile Val Glu 355 360 365Lys
Leu Asn Gly Ile Lys Asp Thr Lys Phe Arg Lys Ser Ile Leu Ser 370 375
380Trp Trp Lys Ser Ala Thr Asp Val Asn Arg Glu Ala Met Ile Ala
Leu385 390 395 400Leu Ser Asn Thr Val Asp Ile Ala Lys Lys Gln Asp
Asp Pro Asp Phe 405 410 415Ser Ser Ala Val Asp Phe Ile Asp Ser Met
Asp Asp Ser Asp Leu Gln 420 425 430Ile Leu Asp Thr Ile Ser Ile Gln
Pro Gly Arg Ala Ala Tyr Ser Ser 435 440 445Lys Thr Leu Arg Ala Leu
Ser Lys Arg Met Tyr Ser Thr Asp Asp Asp 450 455 460Leu His Asp Ala
Arg Lys His Val Phe Gly Val Asp Asp Ser Trp Arg465 470 475 480Pro
Pro Gln Pro Ala Ile Gly Ala Pro Leu Gly Asn Pro Ser Val Asp 485 490
495Arg Val Ala Lys Ile Val Asn Arg Trp Leu Leu Ala Cys Gln Ser Arg
500 505 510Trp Gly Asn Pro Leu Ser Ile Gln Ile Glu His Val Arg Asp
Ala Leu 515 520 525Ser Ser Ala Ala Thr Ala Thr Ala Asp Lys Arg Ala
Tyr Glu Arg Ala 530 535 540Leu Gly Lys Arg Asn Ala Glu Lys Met Lys
Val Lys Asn Glu Leu Arg545 550 555 560Leu Gln Gly Leu Asn Glu Pro
His Glu Ser Asp Val Arg Arg Gln Glu 565 570 575Ala Ile Thr Arg Gln
Gln Gly Lys Cys Leu Tyr Cys Gly Asp Asp Ile 580 585 590Thr Phe Ser
Thr Cys Glu Met Asp His Ile Val Pro Arg Lys Gly His 595 600 605Gly
Ser Thr Asn Thr Arg Asp Asn Leu Ala Ala Val Cys Ile Gln Cys 610 615
620Asn Arg Gln Lys Ser Asn Thr Pro Phe Ala Leu Trp Cys Gln Thr
Pro625 630 635 640Glu Ala Lys Ser Arg Gly Val Ser Leu Glu Ala Ala
Ile His Arg Val 645 650 655Lys Gly Phe Phe Thr Glu Ser Lys Glu Leu
Thr Gly Arg Gln Ala Lys 660 665 670Val Phe Thr Ser Ser Met Ile Met
Arg Leu Lys Gln Thr Thr Ala Asp 675 680 685Asp Pro Ile Asp Ser Arg
Ser Ile Glu Ser Val Ala Trp Met Ala Asp 690 695 700Glu Leu His Arg
Arg Ile Asp Trp His Phe Asn Gly Asp Ala Ser Glu705 710 715 720Ser
Asp His Gly Arg Arg Val Leu Val Ala Val Tyr Gln Gly Arg Ile 725 730
735Thr Ser Glu Ala Arg Asn Val Met Arg Phe Gln Ala Gly Gly Asp Phe
740 745 750His Phe Val Gly Gly His Gly Lys Thr Arg Leu Asp Arg Arg
His His 755 760 765Ala Val Asp Ala Ser Val Ile Ala Met Met Thr Pro
Ala Ala Ala Leu 770 775 780Thr Leu Ala Glu Arg Ile Asn Leu Arg Asp
Ser Gln Arg Cys Ile Gly785 790 795 800Arg Ile Arg Glu Gly Glu Ile
Asp Trp Lys Gln Trp Pro Asn Glu Pro 805 810 815Thr Glu Lys Tyr Gln
His Trp Leu Asp Asn Gly Lys Arg Leu Phe Ala 820 825 830Leu Ile Asn
Asp Ala Leu Asp Asn Asp Arg Ile Pro Ile Thr His Trp 835 840 845Gln
Arg Tyr Ala Leu Gly Asn Ser Ile Ala His Glu Ala Thr Ile His 850 855
860Pro Leu Arg Lys Ile Pro Leu Gly Ser Ala Ile Asp Tyr Glu Thr
Ile865 870 875 880Ser Arg Ala Ala Thr Pro Ala Leu Tyr Cys Ala Leu
Thr Arg Cys Pro 885 890 895Asp Tyr Ser Val Asn Asp Gly Leu Pro Glu
Asn Lys Gln Arg His Ile 900 905 910Thr Val Asn Gly Lys Val Tyr Gly
Pro Glu Asp Glu Val Ala Phe Phe 915 920 925Ala Ser Asp His Val Glu
Leu Ala Val Gln Gly Gly Ser Ala Asp Ile 930 935 940Gly Lys Thr Ile
His His Ala Arg Val Tyr Arg Cys Tyr Phe Val Asp945 950 955 960Arg
Arg Gly Gln Lys Lys Trp Phe Tyr Gly Met Ile Arg Val Phe Arg 965 970
975Val Asp Leu Ile His Ala Arg His Glu Asn Leu Phe Thr Tyr Pro Leu
980 985 990Pro Ser Gln Ser Ile Ser Met Arg Tyr Ala Glu Thr Arg Thr
Ala Glu 995 1000 1005Ala Val Leu Cys Gly His Ala Glu His Val Gly
Asn Leu Val Ala 1010 1015 1020Gly Asp Glu Ile Glu Val Pro Met Ile
Gly Lys Leu Thr Gly Lys 1025 1030 1035Ile Asp Thr Phe Ala Lys Phe
Phe Asn Glu Ser Leu Asn Asn Glu 1040 1045 1050Tyr Val Ala Glu Arg
Trp Ser Ile Asp Gly Phe Asp Ser Glu Ser 1055 1060 1065Lys Leu Leu
Leu Arg Pro Leu Met Leu Ala Glu Glu Gly Ile Ala 1070 1075 1080Asn
Trp Glu Asp Asn Ser His Leu Ser Ile Pro Asp Asp Val Lys 1085 1090
1095Gly Leu Ile Ala Arg Gly Trp Arg Pro Ser Val Asp Thr Val Phe
1100 1105 1110Ala Lys Lys Pro Arg Ile Val Arg Arg Asn Val Leu Gly
Glu Pro 1115 1120 1125Arg Trp Lys Ser Arg Ser Gly Met Pro Val Ser
Trp Arg Val Thr 1130 1135 1140Gly Ser Glu Ala
1145871071PRTLoktanella vestfoldensis 87Met Arg Leu Gly Phe Asp Ile
Gly Thr Asn Ser Ile Gly Trp Trp Leu1 5 10 15Tyr Ala Thr Asp Gly Asn
Glu Ile Thr Gly Val Ile Asp Gly Gly Val 20 25 30Arg Ile Phe Ser Asp
Gly Arg Asp Pro Lys Ser Lys Ala Ser Leu Ala 35 40 45Val Asp Arg Arg
Gly Ala Arg Ala Gln Arg Arg Arg Arg Asp Arg Tyr 50 55 60Leu Arg Arg
Lys Ala Ala Leu Met Lys Arg Met Ala Ala Ala Gly Leu65 70 75 80Met
Pro Ala Asp Pro Val Ala Ala Lys Ala Leu Glu Gln Leu Asp Pro 85 90
95Tyr Ala Leu Arg Ala Ser Gly Leu Asp Gln Glu Leu Pro Leu Thr His
100 105 110Leu Gly Arg Ala Leu Phe His Leu Asn Gln Arg Arg Gly Phe
Lys Ser 115 120 125Asn Arg Lys Thr Asp Lys Gly Asp Asn Glu Ser Gly
Lys Ile Lys Asp 130 135 140Ala Thr Ala Arg Leu Asp Gln Ala Met Ile
Ala Lys Gly Ala Arg Thr145 150 155 160Tyr Gly Glu Phe Leu His Met
Arg Arg Ala Ser Ala Pro Asp Pro Lys 165 170 175Cys Val Pro Thr Val
Arg Thr Arg Leu Ser Ile Ala Pro Arg Asp Asn 180 185 190Ala Glu Lys
Ala Glu Ala Gly Tyr Asp Phe Tyr Pro Asp Arg Arg His 195 200 205Leu
Phe Glu Glu Phe Thr Lys Leu Trp Ala Ala Gln Ala Ala Asn Ser 210 215
220Pro Asp Ile Leu Thr Asp Glu Leu Arg Asp Glu Ile Ser Val Ile
Ile225 230 235 240Phe His Gln Arg Pro Leu Lys Thr Pro Glu Val Gly
Leu Cys Leu Phe 245 250 255Ser Gly Ser His Gly Val Pro Gln Asn Asp
Lys Arg Ile Pro Ser Ala 260 265 270His Pro Gln Asn Gln Arg Arg Ile
Leu Phe Glu Thr Val Asn Asn Leu 275 280 285Lys Val Ala Ala Arg Gly
Glu Leu Ala Arg Gly Leu Thr Arg Asp Glu 290 295 300Arg Asp Thr Ile
Ala His Ala Leu Asp Asn Lys Ala His Thr Lys Ser305 310 315 320Leu
Ser Gly Met Ser Met Lys Leu Lys Ala Leu Gly Lys Leu Ile Lys 325 330
335Leu Arg Pro Glu Gln Ser Phe Thr Leu Glu Thr Ala Asn Arg Asp Ser
340 345 350Ile Val Cys Asp Pro Val Arg Ala Ser Leu Ser His Pro Asp
Arg Phe 355 360 365Gly Ser Arg Trp Ser Thr Leu Asp Ala Glu Ala Gln
Trp Asp Leu Val 370 375 380Gln Arg Ile Arg Ala Val Gln Ser Asp Ala
Glu His Asn Ala Leu Val385 390 395 400Ala Trp Leu Ile Ala Thr His
Gly Leu Asp Arg Val His Ala Glu Asn 405 410 415Val Ala Asn Ala Pro
Leu Pro Glu Gly His Gly Arg Leu Gly Met Thr 420 425 430Ala Thr Lys
Arg Ile Leu Ala Ala Leu Glu Ala Glu Val Ile Pro Tyr 435 440 445Ser
Ala Ala Val Ala Ala Cys Gly Trp His His Ser Asp Gly Arg Thr 450 455
460Gly Glu Val Leu Thr Glu Leu Pro Tyr Tyr Gly Gln Ile Leu Asp
Arg465 470 475 480His Val Ile Pro Gly Thr Tyr Asp Glu Asn Asp Asp
Glu Val Thr Arg 485 490 495Tyr Gly Arg Ile Thr Asn Pro Thr Val His
Ile Gly Leu Asn Gln Leu 500 505 510Arg Arg Leu Val Asn Lys Ile Val
Thr Val Tyr Gly Lys Pro Asp Glu 515 520 525Ile Val Val Glu Leu Ala
Arg Asp Leu Lys Leu Ser Glu Asp Gln Lys 530 535 540Arg Asp Val Gln
Arg Asp Ile Lys Lys Asn Thr Glu Ala Ala Ile Thr545 550 555 560Arg
Gly Gln Lys Ile Glu Glu Leu Gly Tyr Ala Asn Thr Gly Ala Asn 565 570
575Arg Val Met Tyr Arg Leu Trp Glu Glu Leu Gly Pro Ala Ile Gly Pro
580 585 590Arg Cys Cys Pro Tyr Ser Gly Lys Pro Ile Ser Ala Ser Met
Ile Phe 595 600 605Asp Gly Ser Cys Asp Val Asp His Ile Leu Pro Tyr
Ser Arg Thr Leu 610 615 620Glu Asp Gly Phe Ser Asn Arg Thr Leu Cys
Leu Lys Glu Phe Asn Arg625 630 635 640Gln Lys Thr Asn Lys Thr Pro
Trp Glu Ala Trp Gly Asn Thr Pro Ala 645 650 655Trp Asp Ala Ile Glu
Ala Asn Leu Lys Asn Leu Pro Ala Asn Lys Ala 660 665 670Trp Arg Phe
Ala Pro Asp Ala Met Glu Arg Phe Glu Gly Glu Asn Asp 675 680 685Phe
Ser Ala Arg Ala Leu Lys Asp Thr Gln Tyr Leu Ser Arg Ile Ala 690 695
700Arg Ser Tyr Leu Asp Ala Leu Tyr Asp Gly Ala Asp Gly Lys Ser
His705 710 715 720Val Trp Val Val Pro Gly Arg Leu Thr Glu Met Leu
Arg Arg His Trp 725 730 735Gly Leu Asn Ser Leu Leu Pro Asp Lys Asp
Gly Ala Val Lys Ala Lys 740 745 750Asn Arg Ser Asp His Arg His His
Ala Ile Asp Ala Ala Val Val Ala 755 760 765Ala Thr Asp Arg Ser Leu
Val Gln Arg Ile Ser Lys Met Ala Gln Arg 770 775 780Asp Glu Val Asn
Gly Ala Glu Glu Val Ala Arg Ser Val Pro Pro Pro785 790 795 800Trp
Asp Asp Phe Arg Thr Asp Ile Lys Ser Gln Leu Asp Arg Ile Ile 805 810
815Val Ser His Arg Ala Asp His Gly Arg Ile Asp Phe Ala Ala Arg Gln
820 825 830Thr Gly Asn Asp Ser Thr Ser Gly Ala Leu His Glu Ala Thr
Ala Leu 835 840 845Ser Ile Ile Asp Asp Gln Asn Val Ala Val Arg Ile
Pro Leu Leu Ser 850 855 860Leu Ser Ala Ala Gln Phe Glu Glu Gly Gly
Arg Ser Gly Trp Val Arg865 870 875 880Asp Pro Gln Leu Arg Gly Ala
Leu His Leu Ala Thr Lys Gly Lys Asp 885 890 895Lys Lys Asp Phe Glu
Ala Ala Leu Leu Ser Phe Ala Ala Lys Pro Gly 900 905 910Pro Tyr His
Gly Ile Ser Arg Val Arg Ile Glu Lys Pro Leu Gln Asp 915 920 925Thr
Ala Arg Val Tyr Val Pro Ala Asp Ala Pro Ile Lys Ala Tyr Gln 930 935
940Gly Gly Ser Asn His Arg Tyr Glu Val Trp Lys Leu Pro Asp Gly
Lys945 950 955 960Val Leu His His Val Val Ser Met Phe Val Ala His
Gln Gly Asn Leu 965 970 975Thr Arg Pro His Pro Ala Ala Lys Arg Ile
Tyr Gln Phe Met Lys Gly 980 985 990Asp Leu Val Arg Leu Glu Asp Ser
Lys Phe Gly Pro Val Ile Ala Thr 995 1000 1005Val Glu Lys Phe Asn
Gly Lys Gly Met Ile Glu Leu Val Pro His 1010 1015 1020Asn Glu Ala
Asn Ala Ser Asp Arg Tyr Arg Lys Thr Lys Glu Asp 1025 1030 1035Leu
Tyr Ile Arg Leu Gly Ala Thr Thr Leu Leu Arg Ala Lys Ala 1040 1045
1050Arg Arg Val His Val Asp Glu Met Gly Arg Leu Arg Asp Pro Gly
1055 1060 1065Pro Pro Gln 1070881105PRTSphingomonas sanxanigenens
88Met Asp Met Ala Trp Arg Leu Gly Leu Asp Leu Gly Thr Asn Ser Leu1
5 10 15Gly Trp Ala Ala Leu Ser Leu Asp Ala Ala Gly Ala Pro Asp Ala
Ile 20 25 30Leu Ala Ala Gly Ser Arg Ile Phe Gly Asp Gly Arg Asp Pro
Gln Ser 35 40 45Gly Thr Ser Leu Ala Val Asp Arg Arg Ala Ala Arg Ala
Ala Arg Arg 50 55 60Arg Arg Asp Arg Phe Lys Gln Arg Gln Arg Ala Leu
Leu Lys His Leu65 70 75 80Glu Ala Asp Gly Leu Phe Pro Ala Asp Pro
Glu Val Gln Gln Ala Leu 85 90 95Ala Ala Leu Asp Pro Tyr Ala Leu Arg
Ala Arg Ala Leu Asp Glu Ala 100 105 110Leu Ser Leu His Glu Ile Gly
Arg Ala Leu Phe His Leu Asn Gln Arg 115 120 125Arg Gly Phe Gln Ser
Asn Arg Lys Ala Asp Arg Gly Lys Asp Glu Asp 130 135 140Ala Gly Lys
Ile Ala Ile Gly Val Asp Arg Leu Lys Asp Ala Ile Ala145 150 155
160Ala Ala Gly Ala Arg Thr Phe Gly Glu Phe Leu Arg Gln Arg Arg Ala
165 170 175Gly Ala Thr Gly Glu Asn Gln Ile Pro Ser Val Arg Thr Arg
Leu Arg 180 185 190Ala Glu Thr Gly Glu Gly Ala Lys Gly Ser Gly Tyr
Asp Phe Tyr Pro 195 200 205Ser Arg Ala Leu Leu Lys Asp Glu Phe Asp
Ala Ile Trp His Ala Gln 210
215 220Ala Glu His His Pro Lys Val Leu Thr Asp Glu Ala Tyr His Arg
Leu225 230 235 240His Glu Ile Val Phe Arg Gln Arg Pro Leu Arg Ala
Pro Lys Val Gly 245 250 255Ala Cys Thr Leu Val Pro Gly Glu Ala Arg
Leu Pro Lys Ala His Pro 260 265 270Leu Phe Gln Arg Arg Arg Leu Leu
Glu Glu Leu Asn Ala Leu Met Ile 275 280 285Val Arg Ala Gly Ala Val
Ala Glu Arg Leu Thr Pro Glu Gln Arg Asp 290 295 300Leu Leu Leu Leu
Lys Leu Lys Asp Lys Gly Lys Val Thr Phe Lys Ser305 310 315 320Leu
Arg Glu Lys Val Leu Lys Leu Asp Gly Asp Ala Arg Phe Asn Lys 325 330
335Glu Ser Glu His Arg Thr Glu Leu Lys Gly Asp Glu Val Ala Ala Glu
340 345 350Met Gly Gly Lys Thr Arg Phe Gly Ser Leu Trp Pro His Leu
Ser Thr 355 360 365Glu Gln Gln Trp Thr Val Ile Glu Arg Gln Gln Ala
Leu Glu Ser Asp 370 375 380Ala Asp Glu Ala Ala Phe Arg Thr Trp Leu
Val Glu Thr His Lys Leu385 390 395 400Thr Ala Glu Gln Ala Arg Ala
Val Ala Gly Ala Arg Leu Pro Ala Gly 405 410 415His Gly Arg Phe Gly
Leu Thr Ala Thr Ala Ala Leu Leu Ala Ala Leu 420 425 430Arg Asp Gly
Arg Thr Pro Glu Gly Arg Gly Pro Asn Ala Pro Ala Thr 435 440 445Glu
Ala Gly Arg Val Val Val Tyr Ser Glu Ala Ala Ala Ile Ala Gly 450 455
460Tyr His His Ser Asp His Arg Ser Gly Glu Val Phe Thr Asp Ala
Lys465 470 475 480Gly Arg Pro Ala Leu Pro Tyr Tyr Gly Val Pro Leu
Asp Arg His Ile 485 490 495Val Pro Gly Thr Ala Asp Pro Asp Glu Pro
Asp Glu Ala Ala Arg Ile 500 505 510Gly Arg Leu Thr Asn Pro Thr Val
His Ile Ala Leu Asn Gln Leu Arg 515 520 525Arg Val Val Asn Arg Leu
Ile Arg Val Tyr Gly Pro Pro Ala Glu Val 530 535 540Ala Leu Glu Leu
Ala Arg Glu Leu Lys Leu Ser Glu Asp Glu Lys Lys545 550 555 560Glu
Arg Asn Arg Glu Asn Ser Arg Asn Arg Leu Asp Ala Glu Lys Arg 565 570
575Ser Ser Lys Leu Ala Glu Leu Gly Gln Arg Asp Asn Gly Gly Asn Arg
580 585 590Ala Leu Leu Lys Leu Trp Glu Glu Leu Asn Pro Glu Asn Ile
Leu Asp 595 600 605Arg Arg Cys Ile Tyr Ser Gly Arg Gln Ile Ser Ile
Gly Met Ile Phe 610 615 620Ser Gly Ala Ala Glu Val Asp His Ile Leu
Pro Phe Asp Ala Thr Leu625 630 635 640Asp Asp Ser Asn Ala Asn Lys
Ile Leu Cys Leu Arg Glu Ala Asn Arg 645 650 655Asp Lys Arg Lys Arg
Ser Pro Phe Glu Ala Trp Gly Gly Thr Pro Gln 660 665 670Trp Glu Glu
Ile Ala Glu Arg Ala Ser Arg Leu Pro Arg Asn Lys Arg 675 680 685Trp
Arg Phe Glu Pro Asp Ala Met Glu Arg Phe Ala Glu Glu Gly Gly 690 695
700Phe Leu Ala Arg His Leu Val Asp Thr Gln Tyr Leu Gly Arg Ile
Ala705 710 715 720His Asp Tyr Leu Arg Cys Leu Tyr Pro Asp Lys Gly
Asp Gly Ser Ser 725 730 735His Val Trp Val Ser Pro Gly Arg Leu Thr
Glu Met Val Arg Arg Lys 740 745 750Leu Gly Leu Asn Gly Leu Leu Gly
Asp His Asn Leu Gly Ala Glu Gln 755 760 765Pro Lys Asn Arg Lys Asp
His Arg His His Ala Ile Asp Ala Val Val 770 775 780Thr Ala Ile Leu
Asp Arg Ser Met Leu Gln Arg Ile Gln Arg Ala Ser785 790 795 800Gly
Glu Gly Ala Ser Asp Ala Glu Leu Leu Arg Ile Ile Val Pro Glu 805 810
815Pro Trp Thr Gly Phe Arg Asp Asp Leu Gln Arg Ala Val Asp Arg Ile
820 825 830Val Val Ala His Arg Ala Asp His Gly Thr Val Ala Lys Ala
Ala Thr 835 840 845Arg Gly Arg Asp Gln Thr Ala Ala Arg Leu His Asn
Asp Thr Ala Tyr 850 855 860Gly Phe Thr Gly Glu Ala Asp Ala Lys Gly
Thr Pro Ile Val Val His865 870 875 880Arg Ala Pro Leu Gly Ala Leu
Lys Lys Pro Glu His Ile Asp Gln Val 885 890 895Arg Asp Pro Leu Leu
Arg Ala Ala Leu His Asp Phe Thr Ala Gly Leu 900 905 910Ser Gly Lys
Pro Phe Glu Asp Arg Ile Ala Ala Phe Pro Arg Leu Gly 915 920 925Pro
Leu Asp Tyr Arg Gly Ile Arg Arg Val Arg Val Val Glu Pro Leu 930 935
940Gln Val Ile Pro Ile Arg Asp Ala Ala Gly Arg Ala Tyr Lys Gly
Tyr945 950 955 960Lys Gly Asp Ser Asn Tyr Arg Tyr Asp Val Trp Glu
Leu Pro Asp Gly 965 970 975Lys Trp Glu Gln Arg Val Leu Gln Met Tyr
Tyr Ala His Gln Ser Gly 980 985 990Glu Pro Pro Arg Pro His Pro Ala
Ala Arg Lys Val Leu Ser Leu His 995 1000 1005Arg Asp Asp Val Leu
Ala Ile Glu Arg Gly Glu Gly Gly Arg Glu 1010 1015 1020Leu Val Arg
Val Val Lys Phe Ser Thr Asn Glu Phe Thr Leu Ala 1025 1030 1035Pro
Leu Asn Glu Gly Gly Ala Leu Lys Ser Arg His Ala Asp Lys 1040 1045
1050Gly Asp Pro Phe Arg Tyr Leu Tyr Pro Ser Pro Ser Thr Leu Lys
1055 1060 1065Ala Trp Arg Ala Arg Glu Val Arg Val Asp Glu Leu Gly
Arg Val 1070 1075 1080Leu Asp Pro Gly Phe Pro Ala Arg Lys Arg Arg
Arg Val Thr Arg 1085 1090 1095Pro Gly Arg Ala Asp Ala Asp 1100
1105891399PRTEpilithonimonas tenax 89Met Thr Lys Asn Ile Leu Gly
Leu Asp Leu Gly Val Ser Ser Ile Gly1 5 10 15Trp Ala Tyr Val Gln Glu
Asp Asp Lys Asn Ser Ala Asn Asn Lys Ile 20 25 30Ile Lys Leu Gly Val
Arg Val Asn Pro Leu Thr Val Asp Glu Gln Ile 35 40 45Asn Phe Glu Lys
Gly Lys Pro Ile Thr Thr Asn Ala Gly Arg Thr Leu 50 55 60Ala Arg Gly
Ala Arg Arg Asn Leu Gln Arg Phe Lys Leu Arg Arg Ala65 70 75 80Asn
Val Ile Asp Val Leu Thr Lys Gly Asn Ile Leu Lys Asp Gly Asp 85 90
95Leu Leu Thr Glu Val Gly Lys Asn Ser Thr Phe Gln Thr Gln Glu Leu
100 105 110Arg Ala Lys Ser Ala Lys Glu Lys Ile Glu Leu Ser Asp Phe
Val Arg 115 120 125Val Leu Leu Leu Ile Asn Lys Lys Arg Gly Tyr Lys
Ser Ser Arg Lys 130 135 140Ala Lys Asn Glu Asp Glu Gly Gln Ile Ile
Asp Gly Met Ala Val Ala145 150 155 160Lys Lys Leu Tyr Glu Glu Ser
Leu Thr Pro Gly Glu Tyr Ser Tyr Gln 165 170 175Leu Leu Ile Glu Gly
Lys Lys Gln Leu Pro Asp Phe Tyr Arg Ser Asp 180 185 190Leu Gln Ser
Glu Phe Asp Lys Val Trp Lys Phe Gln Lys Gln Phe Tyr 195 200 205Ser
Glu Ile Leu Ile Asp Glu Leu Tyr Lys Glu Leu Gln Ala Lys Asn 210 215
220Lys Asn Ala Thr Trp Ala Ile Leu Lys Glu Pro Phe Ser Leu Val
Gly225 230 235 240Ile Lys Gln Met Gly Thr Met Gln Glu Lys Lys Ile
Glu Lys Tyr Leu 245 250 255Trp Arg Ser Glu Ala Ala Lys Lys Gln Leu
Asp Phe Glu Ser Leu Ala 260 265 270Val Val Phe Gln Glu Ile Asn Ser
Asn Leu Asn Asn Ser Ser Gly Tyr 275 280 285Leu Gly Ala Ile Ser Asp
Arg Ser Lys Glu Leu Tyr Phe Asn His Met 290 295 300Thr Val Gly Glu
Tyr Leu Tyr Gln Gln Leu Lys Ala Asn Pro His Thr305 310 315 320Lys
Leu Lys Asn Gln Val Phe Tyr Arg Gln Asp Tyr Leu Asp Glu Phe 325 330
335Glu Lys Ile Trp Glu Thr Gln Ser Gln Tyr His Ser Glu Leu Thr Lys
340 345 350Glu Leu Lys Glu Gln Val Arg Asp Val Val Ile Phe Tyr Gln
Arg Lys 355 360 365Leu Lys Ser Gln Lys Gly Leu Ile Ser Ile Cys Glu
Phe Glu Asn Arg 370 375 380Glu Ile Glu Ile Ile Glu Asn Gly Lys Thr
Lys Lys Lys Thr Val Gly385 390 395 400Leu Lys Val Ala Pro Lys Ser
Ser Pro Leu Phe Gln Glu Phe Lys Ile 405 410 415Trp Gln Val Leu Asn
Asn Leu Gln Phe Gln Asn Leu Glu Thr Lys Glu 420 425 430Ile Phe Pro
Ile Asp Leu Asp Phe Lys Gln Ser Ile Phe Asp Glu Val 435 440 445Asn
Val Lys Gly Arg Leu Ser Ala Lys Glu Val Leu Asp Ile Val Gly 450 455
460Tyr Ser Gly Lys Glu Trp Lys Thr Asn Phe Lys Asp Ile Glu Gly
Asn465 470 475 480Asn Thr Asn Glu Asn Leu Tyr Thr Ala Phe Leu Lys
Ile Ile Ala Asn 485 490 495Glu Gly Lys Glu Phe Pro Lys Glu Phe Lys
Leu Thr Ile Glu Asp Asp 500 505 510Ile Lys Val Thr Lys Ile His Ser
Ser Ala Ser Lys Ile Lys Glu Phe 515 520 525Val Lys Glu Asn Phe Ser
Ser Leu Gly Ile Asn Thr Ser Ile Leu Asp 530 535 540Phe Asn Pro Glu
Phe Asp Gly Lys Asp Phe Glu Lys Gln Ser Ser Tyr545 550 555 560Gln
Leu Trp His Leu Leu Tyr Ser Tyr Glu Gly Asp Asp Ser Ala Ser 565 570
575Gly Asn Glu Lys Leu Tyr Glu Leu Leu Glu Lys Lys Phe Ser Phe Lys
580 585 590Lys Glu His Ser Lys Ile Leu Ala Glu Ile Gly Phe Ser Pro
Asp Tyr 595 600 605Gly Ser Leu Ser Ser Lys Ala Met Arg Lys Ile Cys
Thr Tyr Ile Lys 610 615 620Glu His Lys Tyr Ser Asp Ala Cys Asn Leu
Ala Gly Tyr Asn His Ser625 630 635 640Lys Asn Ser Leu Thr Lys Glu
Gln Leu Ala Asn Arg Ile Leu Lys Glu 645 650 655Lys Leu Glu Ile Leu
Pro Lys Asn Ser Leu Arg Asn Pro Val Val Glu 660 665 670Lys Ile Leu
Asn Gln Met Ile Asn Val Val Asn Glu Val Ser Lys Glu 675 680 685Tyr
Gly Arg Pro Asp Glu Ile Arg Ile Glu Leu Ala Arg Glu Leu Lys 690 695
700Lys Asn Ala Glu Glu Arg Ala Asn Met Thr Ser Glu Ile Gly Lys
Ala705 710 715 720Thr Leu Leu His Gln Lys Tyr Ala Glu Ile Leu Gln
Lys Glu Tyr Gly 725 730 735Ile Lys Val Pro Ser Arg Asn Asp Ile Ile
Arg Tyr Lys Leu Tyr Leu 740 745 750Glu Leu Ala Asn Asn Gly Phe Lys
Asp Leu Tyr Thr Gly Gln Lys Ile 755 760 765Glu Lys Glu Asn Ile Phe
Thr Asp Lys Tyr Asp Ile Asp His Ile Ile 770 775 780Pro Gln Ser Arg
Phe Phe Asp Asp Ser Phe Ser Asn Lys Val Leu Val785 790 795 800Pro
Arg Gly Ala Asn Leu Lys Lys Gly Asn Ala Thr Ala Phe Asp Tyr 805 810
815Leu Glu Met Glu Gly Lys Asp Gln Leu Glu Lys Phe Leu Asn Thr Ile
820 825 830Lys Asp Leu Phe Asp Lys Ser Leu Ile Ser Lys Ala Lys Phe
Glu Lys 835 840 845Leu Gln Lys Lys Gly Ser Glu Ile Gly Asp Gly Phe
Ile Gln Arg Asp 850 855 860Leu Arg Asp Thr Gln Tyr Ile Ala Lys Lys
Ala Lys Glu Ile Leu Phe865 870 875 880Glu Ile Thr Asn Ser Val Val
Ser Thr Ser Gly Arg Ile Thr Asp Lys 885 890 895Leu Arg Glu Asp Trp
Asn Leu Val Asn Thr Met Lys Glu Leu Asn Leu 900 905 910Asp Lys Tyr
Arg Lys Leu Gly Leu Thr Glu Thr Val Ile Asn Ser Lys 915 920 925Gly
Glu Glu Lys Glu Arg Ile Thr Asp Trp Ser Lys Arg Asn Asp His 930 935
940Arg His His Ala Met Asp Ala Leu Thr Val Ala Phe Thr Thr His
Asn945 950 955 960His Ile Gln Tyr Leu Asn His Leu Asn Ala Arg Lys
Asp Glu Lys His 965 970 975Asn Gln His Ile Ile Ile Ser Asn Ile Glu
Asn Leu Ile Thr Lys Val 980 985 990Tyr Glu Lys Lys Asn Gly Ser Thr
Lys Arg Lys Phe Val Glu Pro Ile 995 1000 1005His Asn Phe Arg Ile
Glu Ala Lys Lys His Leu Asp Glu Ile Leu 1010 1015 1020Ile Ser His
Lys Thr Lys Asn Lys Val Val Thr Lys Asn Ile Asn 1025 1030 1035Lys
Thr Lys Lys Lys Gly Gly Val Val Ala Lys Val Val Leu Thr 1040 1045
1050Pro Arg Gly Gln Leu His Lys Glu Thr Ile Tyr Gly Ser Ser Lys
1055 1060 1065Phe Leu Lys Thr Lys Glu Glu Lys Val Ser Gly Lys Phe
Asp Leu 1070 1075 1080Glu Thr Ile Asn Lys Val Gln Asn Glu Lys Phe
Arg Ile Ala Leu 1085 1090 1095Leu Glu Arg Leu Lys Glu Phe Asn Gly
Asp Ser Lys Lys Ala Phe 1100 1105 1110Thr Gly Lys Asn Val Leu Ala
Lys Asn Pro Ile Tyr Leu Asn Glu 1115 1120 1125Glu Lys Thr Glu Gln
Val Ser Glu Ser Val Ile Leu Ala Trp Tyr 1130 1135 1140Glu Lys Ala
Tyr Thr Ile Arg Lys Ala Val Asn Ser Asp Asn Phe 1145 1150 1155Lys
Asp Tyr Lys Asn Leu Glu Lys Val Ile Asp Asn Gly Val Lys 1160 1165
1170Glu Ile Leu Lys Asn Arg Leu Asp Ala Phe Lys Gly Asn Ala Lys
1175 1180 1185Glu Ala Phe Ser Asp Leu Glu Lys Asn Pro Ile Trp Leu
Asn Glu 1190 1195 1200Ser Lys Gly Ile Ala Ile Lys Thr Val Thr Ile
Thr Gly Ile Asn 1205 1210 1215Asn Ala Glu Thr Leu His Tyr Lys Lys
Asp His Phe Gly Lys Glu 1220 1225 1230Ile Leu Asp Glu Asn Gly Lys
Arg Ile Ala Val Asp Phe Val Ser 1235 1240 1245Thr Gly Asn Asn His
His Val Ala Ile Tyr Glu Asp Ala Asp Gly 1250 1255 1260Asn Leu Gln
Glu Arg Val Val Ser Phe Tyr Glu Ala Val Glu Arg 1265 1270 1275Val
Asn Gln Gly Leu Ser Ile Ile Asp Lys Glu Tyr Asn Ser Gly 1280 1285
1290Leu Gly Trp Lys Phe Leu Phe Thr Met Lys Gln Asn Glu Met Phe
1295 1300 1305Leu Phe Pro Ser Glu Asp Phe Asn Pro Lys Glu Asn Asp
Leu Phe 1310 1315 1320Asp Glu Lys Asn Leu Ser Leu Ile Ser Lys Asn
Met Phe Arg Val 1325 1330 1335Gln Lys Phe Gly Glu Leu Ser Lys Ser
Gly Phe Trp Phe Arg His 1340 1345 1350His Leu Glu Thr Ser Val Glu
Leu Lys Lys Glu Leu Arg Ser Thr 1355 1360 1365Ser Tyr Leu Asp Phe
Tyr Ser Lys Asp Phe Met Lys Thr Ile Val 1370 1375 1380Lys Val Arg
Leu Asn His Leu Gly Lys Ile Val Gln Ile Gly Glu 1385 1390
1395Tyr901453PRTSporocytophaga myxococcoides 90Met Lys Thr Val Leu
Gly Leu Asp Leu Gly Thr Asn Ser Ile Gly Trp1 5 10 15Ala Leu Ile Gln
His Asp Phe Asp Ser Lys Lys Gly Glu Ile Leu Gly 20 25 30Met Gly Ser
Arg Ile Ile Pro Met Ser Gln Asp Ile Leu Gly Glu Phe 35 40 45Gly Lys
Gly Asn Ser Val Ser Gln Thr Ala Asp Arg Thr Lys Phe Arg 50 55 60Ser
Ala Arg Arg Leu Arg Glu Arg His Leu Leu Arg Arg Glu Arg Leu65 70 75
80His Arg Val Leu Asn Ile Leu Gly Phe Leu Pro Arg His Tyr Ala Ala
85 90 95Asp Ile Asp Phe Glu Lys Arg Leu Gly Gln Phe Phe Glu Gly Lys
Glu 100 105 110Pro Lys Leu Ala Tyr Asp Asn Asn Gln Phe Ile Phe Thr
Lys Ser Phe 115 120 125Gly Lys Met Leu Ala Asp Phe Arg Gln His Gln
Pro Asp Phe Leu Lys 130 135 140Asp Glu Lys Ser Asn Asp Leu Leu Ile
Pro Tyr Asp Trp Ser Ile Tyr145 150 155 160Tyr Leu Arg Lys Glu Ala
Leu Thr Lys Lys Ile Glu Lys Glu Glu Leu
165 170 175Ala Trp Ile Ile Leu Asn Phe Asn Gln Lys Arg Gly Tyr Tyr
Gln Leu 180 185 190Arg Gly Glu Glu Glu Glu Glu Asn Leu Asn Lys Leu
Val Glu Phe His 195 200 205Ser Leu Lys Ile Ile Asp Val Val Ala Asp
Glu Lys Leu Asn Asn Lys 210 215 220Gly Glu Thr Trp Tyr Ser Leu Leu
Leu Glu Asn Gly Trp Thr Tyr Arg225 230 235 240Arg Ser Ser Lys Val
Pro Leu Phe Asp Trp Lys Glu Lys Val Arg Asp 245 250 255Phe Ile Val
Thr Thr Asp Ile Asn Asp Asp Gly Ser Val Lys Thr Asp 260 265 270Lys
Glu Gly Asn Glu Lys Arg Ser Phe Arg Ala Pro Ser Asp Asp Asp 275 280
285Trp Thr Leu Leu Lys Lys Lys Thr Glu Gln Asp Ile Asp Lys Ser Arg
290 295 300Lys Pro Ile Gly Ala Tyr Ile Tyr Asp Ala Ile Leu Lys Asn
Pro Lys305 310 315 320Gln Lys Ile Asn Gly Lys Leu Val Arg Thr Val
Glu Arg Lys Phe Tyr 325 330 335Lys Glu Glu Leu Lys Leu Ile Leu Gln
Lys Gln Lys Glu Phe His Pro 340 345 350Glu Leu Gln Ser Thr Asp Leu
Tyr Asn Glu Cys Ile Arg Glu Leu Tyr 355 360 365Lys Asn Asn Asp Ala
His Gln Leu Gln Leu Asn Lys Lys Asp Phe Val 370 375 380His Leu Phe
Leu Glu Asp Ile Ile Phe Tyr Gln Arg Pro Leu Arg Ser385 390 395
400Gln Lys Ser Ser Val Gly Asn Cys Pro Leu Glu Phe Arg Lys Phe Lys
405 410 415Asp Ser Glu Gly Ile Glu Lys Val Glu Tyr Leu Lys Thr Ile
Pro Lys 420 425 430Ser Asn Pro Tyr Phe Gln Glu Phe Arg Ile Trp Gln
Trp Met Tyr Asn 435 440 445Leu Ser Ile Tyr Lys Lys Asp Asp Asp Glu
Asn Val Thr Arg Asp Phe 450 455 460Leu Lys Thr Ile Glu Asp Trp Glu
Asp Leu Phe Glu Phe Leu Asn His465 470 475 480Arg Lys Asp Ile Glu
Gln Glu Thr Leu Leu Lys Phe Leu Leu Gly Lys 485 490 495Asn Gly Val
Lys Gly Lys Ala Leu Lys Val Glu Ala Gly Lys Phe Arg 500 505 510Trp
Asn Tyr Val Ser Asp Lys Ile Tyr Pro Cys Asn Glu Thr Lys Ala 515 520
525Leu Ile Ile Ser Lys Leu Glu Lys Val Lys Gly Val Glu Glu Lys Phe
530 535 540Leu Thr Glu Glu Ile Glu Tyr Lys Leu Trp His Leu Ile Tyr
Ser Val545 550 555 560Thr Asp Lys Val Glu Tyr Glu Lys Ala Leu Lys
Ala Phe Ala Gly Lys 565 570 575Gln Lys Leu Asp Leu Val Ser Phe Val
Asp Ser Phe Lys Arg Phe Pro 580 585 590Pro Phe Lys Asn Glu Tyr Gly
Ala Tyr Ser Glu Lys Ala Ile Lys Lys 595 600 605Ile Leu Pro Leu Leu
Arg Thr Gly Lys Ser Trp Asn Trp Met Ala Ile 610 615 620Asp Ser Lys
Val Arg Asp Arg Ile Asn Lys Ile Ile Thr Gly Glu Phe625 630 635
640Asp Glu Glu Ile Lys Asn Lys Val Arg Glu Lys Ala Glu Lys His Ser
645 650 655Leu Lys Lys Glu Asn Asp Phe Gln Gly Leu Pro Leu Trp Leu
Ala Gln 660 665 670Tyr Val Val Tyr Gly Arg His Ser Glu Ala Ser Phe
Ser Gly Lys Trp 675 680 685Asn Ser Val Asp Asp Leu Lys Lys Tyr Leu
Glu Glu Phe Lys Gln His 690 695 700Ser Leu Arg Asn Pro Ile Val Glu
Gln Val Leu Thr Glu Thr Leu Arg705 710 715 720Val Val Ala Asp Ile
Trp Gln Phe Tyr Gly Lys Gly Glu Lys Asp Phe 725 730 735Phe Ser Glu
Ile His Ile Glu Leu Gly Arg Glu Met Lys Asn Thr Ala 740 745 750Glu
Asp Arg Lys Glu Met Ser Ala Ile Ile Gln Ala Asn Glu Thr Thr 755 760
765Asn Leu Arg Ile Lys Ala Leu Leu Asn Glu Leu Leu Gln Asp Lys Lys
770 775 780Val Glu Asn Val Arg Pro Tyr Ser Pro Ser Gln Gln Glu Ile
Leu Lys785 790 795 800Ile Tyr Glu Asp Gly Val Leu Ser Ser Asp Ile
Glu Ile Pro Asp Asp 805 810 815Ile Gln Lys Ile Ser Lys Ala Ala Gln
Pro Thr Lys Ser Glu Leu Gln 820 825 830Arg Tyr Lys Leu Trp Leu Glu
Gln Lys Tyr Arg Ser Pro Tyr Thr Gly 835 840 845Ala Met Ile Pro Leu
Gly Lys Leu Phe Thr Ser Glu Tyr Glu Ile Glu 850 855 860His Ile Ile
Pro Gln Ser Leu Tyr Phe Asp Asp Ser Phe Ser Asn Lys865 870 875
880Val Ile Cys Glu Ser Ala Val Asn Lys Leu Lys Asp Ser Arg Leu Gly
885 890 895Met Glu Phe Ile Lys Glu Cys His Gly Met Val Val Glu Thr
Gly Phe 900 905 910Gly Lys Ser Val Thr Val Phe Glu Glu Glu Thr Tyr
Arg Asp Phe Val 915 920 925Val Gln Asn Tyr Ser Lys Asn His Ser Lys
Lys Ser Lys Leu Leu Leu 930 935 940Glu Glu Ile Pro Glu Lys Met Ile
Glu Arg Gln Met Asn Asp Thr Arg945 950 955 960Tyr Ile Ser Lys Phe
Ile Ser Ser Val Leu Ser Asn Ile Val Arg Glu 965 970 975Glu Val Asn
Asp Asp Gly Val Asn Ser Lys Asn Ile Val Pro Gly Asn 980 985 990Gly
Lys Ile Thr Thr Gln Leu Lys Gln Asp Trp Gly Leu Asn Asp Ile 995
1000 1005Trp Asn Glu Leu Ile Leu Pro Arg Phe Glu Arg Leu Asn Val
Leu 1010 1015 1020Thr Asp Ser Lys His Phe Thr Ala Trp Ser Glu Asn
His Gln Arg 1025 1030 1035Leu Leu Pro Thr Val Pro Ile Gly Leu Ser
Lys Gly Phe Ser Lys 1040 1045 1050Lys Arg Ile Asp His Arg His His
Ala Leu Asp Ala Leu Val Ile 1055 1060 1065Ala Cys Ala Ser Arg Asn
His Ile Asn Phe Leu Asn Asn Ala His 1070 1075 1080Ala Ile Asp Lys
Lys Lys Asn Ser Glu Glu Lys Gln Lys Phe Arg 1085 1090 1095His Asp
Leu Lys Ala Ile Leu Cys Asp Lys Lys Tyr Ser Asp Lys 1100 1105
1110Ser Glu Lys Asn Tyr Arg Trp Ile Phe Lys Lys Pro Trp Asp Asn
1115 1120 1125Phe Thr Ile Asp Ser Lys Asn Ala Leu Asp Lys Ile Ile
Val Ser 1130 1135 1140Phe Lys Gln Asn Leu Arg Ile Ile Asn Lys Ala
Thr Asn Arg Tyr 1145 1150 1155Glu Lys Trp Val Asp Arg Asp Gly Ile
Lys Val Lys Glu Trp His 1160 1165 1170Lys Gln Glu Gly Ile Asn Trp
Ala Thr Arg Lys Pro Leu His Lys 1175 1180 1185Asp Thr Val Ser Gly
Lys Val Asp Leu Lys Arg Val Thr Val Pro 1190 1195 1200Glu Gly Lys
Ile Leu Thr Ala Thr Arg Lys Ser Leu Asp Thr Ser 1205 1210 1215Phe
Asp Leu Lys Val Ile Glu Ser Ile Thr Asp Thr Gly Ile Gln 1220 1225
1230Lys Ile Leu Lys Asn Tyr Leu Thr Ser Lys Ser Asn Asn Pro Glu
1235 1240 1245Leu Ala Phe Ser Ser Glu Gly Val Glu Asp Met Asn Lys
Asn Ile 1250 1255 1260Arg Lys Tyr Asn Asp Gly Lys Leu His Gln Pro
Ile Tyr Lys Val 1265 1270 1275Arg Ile Phe Glu Leu Gly Ser Lys Phe
Pro Leu Gly Gln Phe Gly 1280 1285 1290Asn Lys Lys Phe Lys Tyr Val
Glu Thr Ala Lys Gly Thr Asn Leu 1295 1300 1305Phe Phe Ala Val Tyr
Glu Asp Glu Asn Lys Asn Arg Asn Tyr Glu 1310 1315 1320Thr Ile Pro
Leu Asn Val Val Ile Glu Arg Gln Lys Gln Gly Leu 1325 1330 1335Ser
Ser Val Pro Ile Lys Ser Glu Lys Gly His Lys Leu Leu Phe 1340 1345
1350Tyr Leu Ser Pro Asn Asp Ile Val Tyr Val Pro Gly Ser Asn Glu
1355 1360 1365Glu Phe Ser Ile Asp Arg Leu Tyr Arg Phe Thr Asp Ser
Ser Asp 1370 1375 1380Lys Thr Ala Asn Phe Ile Pro Leu Ser Val Ser
Ser Leu Ile Phe 1385 1390 1395Ser Ser Asn Lys Asn Glu Gln Lys Lys
Ile Gly Ile Ser Tyr Pro 1400 1405 1410Ile Gln Asp Glu Phe Gly Leu
Gly Ser Pro Gln Ser Lys Asn Gln 1415 1420 1425Lys Ser Ile Asp Gly
Ile Met Ile Lys Glu Lys Cys Val Lys Val 1430 1435 1440Asn Ile Asp
Arg Leu Gly Arg Val Ser Phe 1445 1450911509PRTPsychroflexus torquis
91Met Lys Arg Ile Leu Gly Leu Asp Leu Gly Thr Asn Ser Ile Gly Trp1
5 10 15Ser Leu Ile Glu His Asp Phe Lys Asn Lys Gln Gly Gln Ile Glu
Gly 20 25 30Leu Gly Val Arg Ile Ile Pro Met Ser Gln Glu Ile Leu Gly
Lys Phe 35 40 45Asp Ala Gly Gln Ser Ile Ser Gln Thr Ala Asp Arg Thr
Lys Tyr Arg 50 55 60Gly Val Arg Arg Leu Tyr Gln Arg Asp Asn Leu Arg
Arg Glu Arg Leu65 70 75 80His Arg Val Leu Lys Ile Leu Asp Phe Leu
Pro Lys His Tyr Ser Glu 85 90 95Ser Ile Asp Phe Gln Asp Lys Val Gly
Gln Phe Lys Pro Lys Gln Glu 100 105 110Val Lys Leu Asn Tyr Arg Lys
Asn Glu Lys Asn Lys His Glu Phe Val 115 120 125Phe Met Asn Ser Phe
Ile Glu Met Val Ser Glu Phe Lys Asn Ala Gln 130 135 140Pro Glu Leu
Phe Tyr Asn Lys Gly Asn Gly Glu Glu Thr Lys Ile Pro145 150 155
160Tyr Asp Trp Thr Leu Tyr Tyr Leu Arg Lys Lys Ala Leu Thr Gln Gln
165 170 175Ile Thr Lys Glu Glu Leu Ala Trp Leu Ile Leu Asn Phe Asn
Gln Lys 180 185 190Arg Gly Tyr Tyr Gln Leu Arg Gly Glu Asp Ile Asp
Glu Asp Lys Asn 195 200 205Lys Lys Tyr Met Gln Leu Lys Val Asn Asn
Leu Ile Asp Ser Gly Ala 210 215 220Lys Val Lys Gly Lys Val Leu Tyr
Asn Val Ile Phe Asp Asn Gly Trp225 230 235 240Lys Tyr Glu Lys Gln
Ile Val Asn Lys Asp Glu Trp Glu Gly Arg Thr 245 250 255Lys Glu Phe
Ile Ile Thr Thr Lys Thr Leu Lys Asn Gly Asn Ile Lys 260 265 270Arg
Thr Tyr Lys Ala Val Asp Ser Glu Ile Asp Trp Ala Ala Ile Lys 275 280
285Ala Lys Thr Glu Gln Asp Ile Asn Lys Ala Asn Lys Thr Val Gly Glu
290 295 300Tyr Ile Tyr Glu Ser Leu Leu Asp Asn Pro Ser Gln Lys Ile
Arg Gly305 310 315 320Lys Leu Val Lys Thr Ile Glu Arg Lys Phe Tyr
Lys Glu Glu Phe Glu 325 330 335Lys Leu Leu Ser Lys Gln Ile Glu Leu
Gln Pro Glu Leu Phe Asn Glu 340 345 350Ser Leu Tyr Lys Ala Cys Ile
Lys Glu Leu Tyr Pro Arg Asn Glu Asn 355 360 365His Gln Ser Asn Asn
Lys Lys Gln Gly Phe Glu Tyr Leu Phe Thr Glu 370 375 380Asp Ile Ile
Phe Tyr Gln Arg Pro Leu Lys Ser Gln Lys Ser Asn Ile385 390 395
400Ser Gly Cys Gln Phe Glu His Lys Ile Tyr Lys Gln Lys Asn Lys Lys
405 410 415Thr Gly Lys Leu Glu Leu Ile Lys Glu Pro Ile Lys Thr Ile
Ser Arg 420 425 430Ser His Pro Leu Phe Gln Glu Phe Arg Ile Trp Gln
Trp Leu Gln Asn 435 440 445Leu Lys Ile Tyr Asn Lys Glu Lys Ile Glu
Asn Gly Lys Leu Glu Asp 450 455 460Val Thr Thr Gln Leu Leu Pro Asn
Asn Glu Ala Tyr Val Thr Leu Phe465 470 475 480Asp Phe Leu Asn Thr
Lys Lys Glu Leu Glu Gln Lys Gln Phe Ile Glu 485 490 495Tyr Phe Val
Lys Lys Lys Leu Ile Asp Lys Lys Glu Lys Glu His Phe 500 505 510Arg
Trp Asn Phe Val Glu Asp Lys Lys Tyr Pro Phe Ser Glu Thr Arg 515 520
525Ala Gln Phe Leu Ser Arg Leu Ala Lys Val Lys Gly Ile Lys Asn Thr
530 535 540Glu Asp Phe Leu Asn Lys Asn Thr Gln Val Gly Ser Lys Glu
Asn Ser545 550 555 560Pro Phe Ile Lys Arg Ile Glu Gln Leu Trp His
Ile Ile Tyr Ser Val 565 570 575Ser Asp Leu Lys Glu Tyr Glu Lys Ala
Leu Glu Lys Phe Ala Glu Lys 580 585 590His Asn Leu Glu Lys Asp Ser
Phe Leu Lys Asn Phe Lys Lys Phe Pro 595 600 605Pro Phe Val Ser Asp
Tyr Ala Ser Tyr Ser Lys Lys Ala Ile Ser Lys 610 615 620Leu Leu Pro
Ile Met Arg Met Gly Lys Tyr Trp Ser Glu Ser Ala Val625 630 635
640Pro Thr Gln Val Lys Glu Arg Ser Leu Ser Ile Met Glu Arg Val Lys
645 650 655Val Leu Pro Leu Lys Glu Gly Tyr Ser Asp Lys Asp Leu Ala
Asp Leu 660 665 670Leu Ser Arg Val Ser Asp Asp Asp Ile Pro Lys Gln
Leu Ile Lys Ser 675 680 685Phe Ile Ser Phe Lys Asp Lys Asn Pro Leu
Lys Gly Leu Asn Thr Tyr 690 695 700Gln Ala Asn Tyr Leu Val Tyr Gly
Arg His Ser Glu Thr Gly Asp Ile705 710 715 720Gln His Trp Lys Thr
Pro Glu Asp Ile Asp Arg Tyr Leu Asn Asn Phe 725 730 735Lys Gln His
Ser Leu Arg Asn Pro Ile Val Glu Gln Val Val Met Glu 740 745 750Thr
Leu Arg Val Val Arg Asp Ile Trp Glu His Tyr Gly Asn Asn Glu 755 760
765Lys Asp Phe Phe Lys Glu Ile His Val Glu Leu Gly Arg Glu Met Lys
770 775 780Ser Pro Ala Gly Lys Arg Glu Lys Leu Ser Gln Arg Asn Thr
Glu Asn785 790 795 800Glu Asn Thr Asn His Arg Ile Arg Glu Val Leu
Lys Glu Leu Met Asn 805 810 815Asp Ala Ser Val Glu Gly Gly Val Arg
Asp Tyr Ser Pro Ser Gln Gln 820 825 830Glu Ile Leu Lys Leu Tyr Glu
Glu Gly Ile Tyr Gln Asn Pro Asn Thr 835 840 845Asn Tyr Leu Lys Val
Asp Glu Asp Glu Ile Leu Lys Ile Arg Lys Lys 850 855 860Asn Asn Pro
Thr Gln Lys Glu Ile Gln Arg Tyr Lys Leu Trp Leu Glu865 870 875
880Gln Gly Tyr Ile Ser Pro Tyr Thr Gly Lys Ile Ile Pro Leu Thr Lys
885 890 895Leu Phe Thr His Glu Tyr Gln Ile Glu His Ile Ile Pro Gln
Ser Arg 900 905 910Tyr Tyr Asp Asn Ser Leu Gly Asn Lys Ile Ile Cys
Glu Ser Glu Val 915 920 925Asn Glu Asp Lys Asp Asn Lys Thr Ala Tyr
Glu Tyr Leu Lys Val Glu 930 935 940Lys Gly Ser Ile Val Phe Gly His
Lys Leu Leu Asn Leu Asp Glu Tyr945 950 955 960Glu Ala His Val Asn
Lys Tyr Phe Lys Lys Asn Lys Thr Lys Leu Lys 965 970 975Asn Leu Leu
Ser Glu Asp Ile Pro Glu Gly Phe Ile Asn Arg Gln Leu 980 985 990Asn
Asp Ser Arg Tyr Ile Ser Lys Leu Val Lys Gly Leu Leu Ser Asn 995
1000 1005Ile Val Arg Glu Asn Gly Glu Gln Glu Ala Thr Ser Lys Asn
Leu 1010 1015 1020Ile Pro Val Thr Gly Val Val Thr Ser Lys Leu Lys
Gln Asp Trp 1025 1030 1035Gly Leu Asn Asp Lys Trp Asn Glu Ile Ile
Ala Pro Arg Phe Lys 1040 1045 1050Arg Leu Asn Lys Leu Thr Asn Ser
Asn Asp Phe Gly Phe Trp Asp 1055 1060 1065Asn Asp Ile Asn Ala Phe
Arg Ile Gln Val Pro Asp Ser Leu Ile 1070 1075 1080Lys Gly Phe Ser
Lys Lys Arg Ile Asp His Arg His His Ala Leu 1085 1090 1095Asp Ala
Leu Val Val Ala Cys Thr Ser Arg Asn His Thr His Tyr 1100 1105
1110Leu Ser Ala Leu Asn Ala Glu Asn Lys Asn Tyr Ser Leu Arg Asp
1115 1120 1125Lys Leu Val Ile Lys Asn Glu Asn Gly Asp Tyr Thr Lys
Thr Phe 1130 1135 1140Gln Ile Pro Trp Gln Gly Phe Thr Ile Glu Ala
Lys Asn Asn Leu 1145 1150 1155Glu Lys Thr Val Val Ser Phe Lys Lys
Asn Leu Arg Val Ile Asn 1160 1165 1170Lys Thr Asn Asn
Lys Phe Trp Ser Tyr Lys Asp Glu Asn Gly Asn 1175 1180 1185Leu Asn
Leu Gly Lys Asp Gly Lys Pro Lys Lys Lys Leu Arg Lys 1190 1195
1200Gln Thr Lys Gly Tyr Asn Trp Ala Ile Arg Lys Pro Leu His Lys
1205 1210 1215Glu Thr Val Ser Gly Ile Tyr Asn Ile Asn Ala Pro Lys
Asn Lys 1220 1225 1230Ile Ala Thr Ser Val Arg Thr Leu Leu Thr Glu
Ile Lys Asn Glu 1235 1240 1245Lys His Leu Ala Lys Ile Thr Asp Leu
Arg Ile Arg Glu Thr Ile 1250 1255 1260Leu Pro Asn His Leu Lys His
Tyr Leu Asn Asn Lys Gly Glu Ala 1265 1270 1275Asn Phe Ser Glu Ala
Phe Ser Gln Gly Gly Ile Glu Asp Leu Asn 1280 1285 1290Lys Lys Ile
Thr Thr Leu Asn Glu Gly Lys Lys His Gln Pro Ile 1295 1300 1305Tyr
Arg Val Lys Ile Phe Glu Val Gly Ser Lys Phe Ser Ile Ser 1310 1315
1320Glu Asp Glu Asn Ser Ala Lys Ser Lys Lys Tyr Val Glu Ala Ala
1325 1330 1335Lys Gly Thr Asn Leu Phe Phe Ala Ile Tyr Leu Asp Glu
Glu Asn 1340 1345 1350Lys Lys Arg Asn Tyr Glu Thr Ile Pro Leu Asn
Glu Val Ile Thr 1355 1360 1365His Gln Lys Gln Val Ala Gly Phe Pro
Lys Ser Glu Arg Leu Ser 1370 1375 1380Val Gln Pro Asp Ser Gln Lys
Gly Thr Phe Leu Phe Thr Leu Ser 1385 1390 1395Pro Asn Asp Leu Val
Tyr Val Pro Asn Asn Glu Glu Leu Glu Asn 1400 1405 1410Arg Asp Leu
Phe Asn Leu Gly Asn Leu Asn Val Glu Gln Ile Ser 1415 1420 1425Arg
Ile Tyr Lys Phe Thr Asp Ser Ser Asp Lys Thr Cys Asn Phe 1430 1435
1440Ile Pro Phe Gln Val Ser Lys Leu Ile Phe Asn Leu Lys Lys Lys
1445 1450 1455Glu Gln Lys Lys Leu Asp Val Asp Phe Ile Ile Gln Asn
Glu Phe 1460 1465 1470Gly Leu Gly Ser Pro Gln Ser Lys Asn Gln Lys
Ser Ile Asp Asp 1475 1480 1485Val Met Ile Lys Glu Lys Cys Ile Lys
Leu Lys Ile Asp Arg Leu 1490 1495 1500Gly Asn Ile Ser Lys Ala
15059236DNALactobacillus reuteri 92gttttagatg tacttcaaat caataatgtt
tagaac 369336DNALactobacillus rossiae 93gttttagatg tatgtcagat
caatagggtt aagaac 369436DNAPediococcus pentosaceus 94gtttcagaag
agtgttaaat caataagttc aagtac 369536DNALactobacillus nodensis
95gttttagtac tctcaagaat ttagtaacag taaaac 369637DNAArtificial
sequencesequence from Sulfurospirillum sp. SCADC (Sspe)
96acgacacttt acaacaccat cgcttagcaa ctgraac 379736DNABifidobacterium
thermophilum 97caagtctatc aagaaggaag gatgctaatt ctcagc
369836DNALoktanella vestfoldensis 98attatagctg ttcaaaattc
gcggtccagc cgcaac 369936DNASphingomonas sanxanigenens 99accatagctg
ttcagagatc gcggtccagc ggcaac 3610047DNAEpilithonimonas tenax DSM
100gttgttgtgt atcccaaaga tacttaaaaa tgaaagcaat tcacaac
4710147DNASporocytophaga myxococcoides 101gttgtgattt gctttcaaac
aacgatcttt gaattatcgg aaacaac 4710246DNAPsychroflexus torquis
102cctgtgaatt atcactaaaa atacaatttt gaaagcaatt cacaac
4610336DNALactobacillus reuteri 103tttaactctg tacttcaaat caataatatt
tagtag 3610437DNALactobacillus rossiae 104tttaacgctg tatgtcagat
caatagggtc aaaagtt 3710537DNAPediococcus pentosaceus 105tttcaaaaca
gaatgttaaa tcaataagtt taagtac 3710638DNALactobacillus nodensis
106gttttactgt tactaaattc ttgagaacct actaaaat 3810739DNAartificial
sequencesequence from Sulfurospirillum sp. SCADC (Sspe)
107atttcagtgc atcgaacgat acgtgttgta aagtgtcgt
3910836DNABifidobacterium thermophilum 108gtgtagcgcg atatactaaa
aaagctaatt ctcagt 3610936DNALoktanella vestfoldensis 109ttaacggctc
gaccgcgaat tttgaacatt tggata 3611036DNASphingomonas sanxanigenens
110ttaaccgcca gaccacgatc tctgaagtgc tcaggc
3611147DNAEpilithonimonas tenax 111attgtgaatt gctttcggat acaaatgtag
aaattaatta tttatat 4711247DNASporocytophaga myxococcoides
112attgtgattt gctttcacta gcaaatatat ataaatttaa ttttatt
4711346DNAPsychroflexus torquis 113attgtgaatt gcttcaaaat tgtgtaccaa
atatattaat attata 46114169RNAArtificial Sequencesynthesized Lreu
Single Guide RNA 114gggcgcuaaa gaggaagagg acaguuuuag auguacuucg
aaagaaguac agaguuaaaa 60ucaaacaagu gcuucagcac aaguuucuac uuuugaguca
guuaucugac caauacauaa 120aagggaauua aaucguucac augagcguuu
uaauucccuu ucucucuuu 169115166RNAArtificial Sequencesynthesized
Lros Single Guide RNA 115gggcgcuaaa gaggaagagg acaguuuuag
auguaugucg aaagacauac agcguuaaaa 60ucaagcaagg cuuucgagcc aaguuuaaau
cuuuggguuc gcuauucgga ccguacauag 120uaaaagggcg ucaucgaguu
caaauucgau ggcgcccuuu aguuuu 166116168RNAArtificial
Sequencesynthesized Ppen Single Guide RN 116gggcgcuaaa gaggaagagg
acaguuucag aagaguguug aaaaacauuc uguuuugaaa 60ucaaacaacg cuuuacgcgg
aguuuacaca ucugucccau uauaugggca uuacauaaua 120aaagagaaau
caccuuuaag guugauuccu cuuuuucuug cguuccuu 168117114RNAArtificial
Sequence2synthesized Lnod Single Guide RNA 117gggcgcuaaa gaggaagagg
acaguuuuag uacucucaag gaaacuugag aaccuacuaa 60aauaaggauu uauuccgaau
uuaccaccua uuuuuuaaua ggugguuuuu uuuu 114118180RNAArtificial
Sequencesynthesized Sspe Single Guide RNA 118gggcgcuaaa gaggaagagg
acaacgacac uuuacaacac gaaaguguug uaaagugucg 60ugagugaaag uauagcauuu
uucucaaaau aucgcacuuu uuuugcaagg aaauaauuug 120cgaaaaaaag
agacaauuuc gauacaagaa agccccauca agucgaugau uuguaaggcu
180119117RNAArtificial Sequencesynthesized Sspe Single Guide RNA
(tracrRNA Antisense-crRNA Antisens 119gggcgcuaaa gaggaagagg
acaguuccag uugcuaagcg aaaguucgau gcacugaaau 60caucgaaaag auauauagac
ccgcccaacu gccuuaggca uggcgggcuu cuuucuu 117120254RNAArtificial
Sequencesynthesized Bthe Single Guide RNA 120gggcgcuaaa gaggaagagg
acagcugaga auuagcaucg aaaaaagcua auucucagua 60aguacagcaa uuuauagcug
uaucugaaug cuaagcgguu agccgcaggg gagagcuucg 120gcucuccccg
uucuuuucua gucgcacgug acacgccaua gguagauugc cugugagcaa
180cucgagaaua guaguauucu ucucuguaaa acgagacauc guugcugaug
acuccauuga 240caaagaaggg aauu 254121200RNAArtificial
Sequencesynthesized Lves Single Guide RNA 121gggcgcuaaa gaggaagagg
acaguugcgg cuggaccgcg aaagcggucg agccguuaac 60aagcauucga uugcaccaca
uugaaacgca ggcuacggcc ugcguuuucu uauuuccggu 120augguucguc
uugucaaagu gucucagaga ugcccgcuuu cuaaaugugg auugucgagg
180uucgcacccg uagcguuuuu 200122195RNAArtificial Sequencesynthesized
Ssan Single Guide RNA 122gggcgcuaaa gaggaagagg acaguugccg
cuggaccgcg aaaguggucu ggcgguuaac 60aagcagccag ucugcaccag auaagggcgg
cgcuccggcg ccgccuuuuu uguuugcccg 120gaauguugag acaggcgcca
aaccaucccc ccuccccucg gggcgugcgg caaucgcugc 180uuucagcuca cucca
195123155RNAArtificial Sequencesynthesized Eten Single Guide RNA
123gggcgcuaaa gaggaagagg acaguuguga auugcuuucg aaagaaagca
auucacaaua 60aggauuauuc cguugugaaa acauuuagca ccucgccuau cugcggggua
uuuuuuauuu 120aauacauuca aacaaugaag ucgauugcau uuauu
155124149RNAArtificial Sequencessynthesized Smyx Single Guide RNA
124gggcgcuaaa gaggaagagg acaguuguga uuugcuuucg aaaaaagcaa
aucacaauaa 60ggauuauucc guugugaaaa cauauaagcc gccucgucuu acaauacggg
gcuuuuuuua 120uuccccauuc acauauggcc auccauuuu
149125155RNAArtificial Sequencesynthesized Ptor Single Guide RNA
125gggcgcuaaa gaggaagagg acaguuguga auugcuuuca gaaaugaagc
aauucacaau 60aaggauuauu ccguugugaa aacauuuaaa gcggccuuua cgggucgcuu
ucuuuuuuua 120aagaacuuaa acuuaauaaa uucuuuacaa acuca
15512657DNAArtificial sequencesynthesized GG-939 126gactagacct
gcaggggatc ccgtcgacaa attctaaacg ctaaagagga agaggac
57127174RNAArtificial sequencesynthesized single guide
RNAmisc_feature(1)..(20)n is a, c, g, or u 127nnnnnnnnnn nnnnnnnnnn
gcuauaguuc cuuacugaaa gguaaguugc uauaguaagg 60gcaacagacc cgaggcguug
gggaucgccu agcccguuuu uacgggcucu ccccauauuc 120aaaauaauga
cagacgagca ccuuggagca uuuauuuccg aggugcuuuu uuuu
174128166RNAArtificial Sequencesynthesized Lreu Single Guide
RNAmisc_feature(1)..(20)n is a, c, g, or u 128nnnnnnnnnn nnnnnnnnnn
guuuuagaug uacuucgaaa gaaguacaga guuaaaauca 60aacaagugcu ucagcacaag
uuucuacuuu ugagucaguu aucugaccaa uacauaaaag 120ggaauuaaau
cguucacaug agcguuuuaa uucccuuucu cucuuu 166129163RNAArtificial
Sequencesynthesized Lros Single Guide RNAmisc_feature(1)..(20)n is
a, c, g, or u 129nnnnnnnnnn nnnnnnnnnn guuuuagaug uaugucgaaa
gacauacagc guuaaaauca 60agcaaggcuu ucgagccaag uuuaaaucuu uggguucgcu
auucggaccg uacauaguaa 120aagggcguca ucgaguucaa auucgauggc
gcccuuuagu uuu 163130165RNAArtificial Sequencesynthesized Ppen
Single Guide RNAmisc_feature(1)..(20)n is a, c, g, or u
130nnnnnnnnnn nnnnnnnnnn guuucagaag aguguugaaa aacauucugu
uuugaaauca 60aacaacgcuu uacgcggagu uuacacaucu gucccauuau augggcauua
cauaauaaaa 120gagaaaucac cuuuaagguu gauuccucuu uuucuugcgu uccuu
165131111RNAArtificial Sequencesynthesized Lnod Single Guide
RNAmisc_feature(1)..(20)n is a, c, g, or u 131nnnnnnnnnn nnnnnnnnnn
guuuuaguac ucucaaggaa acuugagaac cuacuaaaau 60aaggauuuau uccgaauuua
ccaccuauuu uuuaauaggu gguuuuuuuu u 111132177RNAArtificial
Sequencesynthesized Sspe Single Guide RNAmisc_feature(1)..(20)n is
a, c, g, or u 132nnnnnnnnnn nnnnnnnnnn acgacacuuu acaacacgaa
aguguuguaa agugucguga 60gugaaaguau agcauuuuuc ucaaaauauc gcacuuuuuu
ugcaaggaaa uaauuugcga 120aaaaaagaga caauuucgau acaagaaagc
cccaucaagu cgaugauuug uaaggcu 177133114RNAArtificial
Sequencesynthesized Sspe Single Guide RNA (tracrRNA Antisense-crRNA
Antisensemisc_feature(1)..(20)n is a, c, g, or u 133nnnnnnnnnn
nnnnnnnnnn guuccaguug cuaagcgaaa guucgaugca cugaaaucau 60cgaaaagaua
uauagacccg cccaacugcc uuaggcaugg cgggcuucuu ucuu
114134251RNAArtificial Sequencesynthesized Bthe Single Guide
RNAmisc_feature(1)..(20)n is a, c, g, or u 134nnnnnnnnnn nnnnnnnnnn
gcugagaauu agcaucgaaa aaagcuaauu cucaguaagu 60acagcaauuu auagcuguau
cugaaugcua agcgguuagc cgcaggggag agcuucggcu 120cuccccguuc
uuuucuaguc gcacgugaca cgccauaggu agauugccug ugagcaacuc
180gagaauagua guauucuucu cuguaaaacg agacaucguu gcugaugacu
ccauugacaa 240agaagggaau u 251135197RNAArtificial
Sequencesynthesized Lves Single Guide RNAmisc_feature(1)..(20)n is
a, c, g, or u 135nnnnnnnnnn nnnnnnnnnn guugcggcug gaccgcgaaa
gcggucgagc cguuaacaag 60cauucgauug caccacauug aaacgcaggc uacggccugc
guuuucuuau uuccgguaug 120guucgucuug ucaaaguguc ucagagaugc
ccgcuuucua aauguggauu gucgagguuc 180gcacccguag cguuuuu
197136192RNAArtificial Sequencesynthesized Ssan Single Guide
RNAmisc_feature(1)..(20)n is a, c, g, or u 136nnnnnnnnnn nnnnnnnnnn
guugccgcug gaccgcgaaa guggucuggc gguuaacaag 60cagccagucu gcaccagaua
agggcggcgc uccggcgccg ccuuuuuugu uugcccggaa 120uguugagaca
ggcgccaaac cauccccccu ccccucgggg cgugcggcaa ucgcugcuuu
180cagcucacuc ca 192137152RNAArtificial Sequencesynthesized Eten
Single Guide RNAmisc_feature(1)..(20)n is a, c, g, or u
137nnnnnnnnnn nnnnnnnnnn guugugaauu gcuuucgaaa gaaagcaauu
cacaauaagg 60auuauuccgu ugugaaaaca uuuagcaccu cgccuaucug cgggguauuu
uuuauuuaau 120acauucaaac aaugaagucg auugcauuua uu
152138146RNAArtificial Sequencesynthesized Smyx Single Guide
RNAmisc_feature(1)..(20)n is a, c, g, or u 138nnnnnnnnnn nnnnnnnnnn
guugugauuu gcuuucgaaa aaagcaaauc acaauaagga 60uuauuccguu gugaaaacau
auaagccgcc ucgucuuaca auacggggcu uuuuuuauuc 120cccauucaca
uauggccauc cauuuu 146139152RNAArtificial Sequencesynthesized Ptor
Single Guide RNAmisc_feature(1)..(20)n is a, c, g, or u
139nnnnnnnnnn nnnnnnnnnn guugugaauu gcuuucagaa augaagcaau
ucacaauaag 60gauuauuccg uugugaaaac auuuaaagcg gccuuuacgg gucgcuuucu
uuuuuuaaag 120aacuuaaacu uaauaaauuc uuuacaaacu ca
1521401092PRTBrevibacillus laterosporus 140Met Ala Tyr Thr Met Gly
Ile Asp Val Gly Ile Ala Ser Cys Gly Trp1 5 10 15Ala Ile Val Asp Leu
Glu Arg Gln Arg Ile Ile Asp Ile Gly Val Arg 20 25 30Thr Phe Glu Lys
Ala Glu Asn Pro Lys Asn Gly Glu Ala Leu Ala Val 35 40 45Pro Arg Arg
Glu Ala Arg Ser Ser Arg Arg Arg Leu Arg Arg Lys Lys 50 55 60His Arg
Ile Glu Arg Leu Lys His Met Phe Val Arg Asn Gly Leu Ala65 70 75
80Val Asp Ile Gln His Leu Glu Gln Thr Leu Arg Ser Gln Asn Glu Ile
85 90 95Asp Val Trp Gln Leu Arg Val Asp Gly Leu Asp Arg Met Leu Thr
Gln 100 105 110Lys Glu Trp Leu Arg Val Leu Ile His Leu Ala Gln Arg
Arg Gly Phe 115 120 125Gln Ser Asn Arg Lys Thr Asp Gly Ser Ser Glu
Asp Gly Gln Val Leu 130 135 140Val Asn Val Thr Glu Asn Asp Arg Leu
Met Glu Glu Lys Asp Tyr Arg145 150 155 160Thr Val Ala Glu Met Met
Val Lys Asp Glu Lys Phe Ser Asp His Lys 165 170 175Arg Asn Lys Asn
Gly Asn Tyr His Gly Val Val Ser Arg Ser Ser Leu 180 185 190Leu Val
Glu Ile His Thr Leu Phe Glu Thr Gln Arg Gln His His Asn 195 200
205Ser Leu Ala Ser Lys Asp Phe Glu Leu Glu Tyr Val Asn Ile Trp Ser
210 215 220Ala Gln Arg Pro Val Ala Thr Lys Asp Gln Ile Glu Lys Met
Ile Gly225 230 235 240Thr Cys Thr Phe Leu Pro Lys Glu Lys Arg Ala
Pro Lys Ala Ser Trp 245 250 255His Phe Gln Tyr Phe Met Leu Leu Gln
Thr Ile Asn His Ile Arg Ile 260 265 270Thr Asn Val Gln Gly Thr Arg
Ser Leu Asn Lys Glu Glu Ile Glu Gln 275 280 285Val Val Asn Met Ala
Leu Thr Lys Ser Lys Val Ser Tyr His Asp Thr 290 295 300Arg Lys Ile
Leu Asp Leu Ser Glu Glu Tyr Gln Phe Val Gly Leu Asp305 310 315
320Tyr Gly Lys Glu Asp Glu Lys Lys Lys Val Glu Ser Lys Glu Thr Ile
325 330 335Ile Lys Leu Asp Asp Tyr His Lys Leu Asn Lys Ile Phe Asn
Glu Val 340 345 350Glu Leu Ala Lys Gly Glu Thr Trp Glu Ala Asp Asp
Tyr Asp Thr Val 355 360 365Ala Tyr Ala Leu Thr Phe Phe Lys Asp Asp
Glu Asp Ile Arg Asp Tyr 370 375 380Leu Gln Asn Lys Tyr Lys Asp Ser
Lys Asn Arg Leu Val Lys Asn Leu385 390 395 400Ala Asn Lys Glu Tyr
Thr Asn Glu Leu Ile Gly Lys Val Ser Thr Leu 405 410 415Ser Phe Arg
Lys Val Gly His Leu Ser Leu Lys Ala Leu Arg Lys Ile 420 425 430Ile
Pro Phe Leu Glu Gln Gly Met Thr Tyr Asp Lys Ala Cys Gln Ala 435 440
445Ala Gly Phe Asp Phe Gln Gly Ile Ser Lys Lys Lys Arg Ser Val Val
450 455 460Leu Pro Val Ile Asp Gln Ile Ser Asn Pro Val Val Asn Arg
Ala Leu465 470 475 480Thr Gln Thr Arg Lys Val Ile Asn Ala Leu Ile
Lys Lys Tyr Gly Ser 485 490 495Pro Glu Thr Ile His Ile Glu Thr Ala
Arg Glu Leu Ser Lys Thr Phe 500 505 510Asp Glu Arg Lys Asn Ile Thr
Lys Asp Tyr Lys Glu Asn Arg Asp Lys 515 520 525Asn Glu His Ala Lys
Lys His Leu Ser Glu Leu Gly Ile Ile Asn Pro 530 535 540Thr Gly Leu
Asp Ile Val Lys Tyr Lys Leu Trp Cys Glu Gln Gln Gly545 550 555
560Arg Cys Met Tyr Ser Asn Gln Pro Ile Ser Phe Glu Arg Leu Lys Glu
565 570 575Ser Gly Tyr Thr Glu Val Asp His Ile Ile Pro Tyr Ser Arg
Ser Met 580 585 590Asn Asp Ser Tyr Asn Asn Arg Val Leu Val Met Thr
Arg Glu Asn Arg 595 600 605Glu Lys Gly Asn Gln Thr Pro Phe Glu Tyr
Met Gly Asn Asp Thr Gln 610 615 620Arg Trp Tyr Glu Phe Glu Gln Arg
Val Thr Thr Asn Pro Gln Ile Lys625 630 635 640Lys Glu Lys Arg Gln
Asn Leu Leu Leu Lys Gly Phe Thr Asn Arg Arg 645 650 655Glu Leu Glu
Met Leu Glu Arg Asn Leu Asn Asp Thr Arg Tyr Ile Thr 660
665 670Lys Tyr Leu Ser His Phe Ile Ser Thr Asn Leu Glu Phe Ser Pro
Ser 675 680 685Asp Lys Lys Lys Lys Val Val Asn Thr Ser Gly Arg Ile
Thr Ser His 690 695 700Leu Arg Ser Arg Trp Gly Leu Glu Lys Asn Arg
Gly Gln Asn Asp Leu705 710 715 720His His Ala Met Asp Ala Ile Val
Ile Ala Val Thr Ser Asp Ser Phe 725 730 735Ile Gln Gln Val Thr Asn
Tyr Tyr Lys Arg Lys Glu Arg Arg Glu Leu 740 745 750Asn Gly Asp Asp
Lys Phe Pro Leu Pro Trp Lys Phe Phe Arg Glu Glu 755 760 765Val Ile
Ala Arg Leu Ser Pro Asn Pro Lys Glu Gln Ile Glu Ala Leu 770 775
780Pro Asn His Phe Tyr Ser Glu Asp Glu Leu Ala Asp Leu Gln Pro
Ile785 790 795 800Phe Val Ser Arg Met Pro Lys Arg Ser Ile Thr Gly
Glu Ala His Gln 805 810 815Ala Gln Phe Arg Arg Val Val Gly Lys Thr
Lys Glu Gly Lys Asn Ile 820 825 830Thr Ala Lys Lys Thr Ala Leu Val
Asp Ile Ser Tyr Asp Lys Asn Gly 835 840 845Asp Phe Asn Met Tyr Gly
Arg Glu Thr Asp Pro Ala Thr Tyr Glu Ala 850 855 860Ile Lys Glu Arg
Tyr Leu Glu Phe Gly Gly Asn Val Lys Lys Ala Phe865 870 875 880Ser
Thr Asp Leu His Lys Pro Lys Lys Asp Gly Thr Lys Gly Pro Leu 885 890
895Ile Lys Ser Val Arg Ile Met Glu Asn Lys Thr Leu Val His Pro Val
900 905 910Asn Lys Gly Lys Gly Val Val Tyr Asn Ser Ser Ile Val Arg
Thr Asp 915 920 925Val Phe Gln Arg Lys Glu Lys Tyr Tyr Leu Leu Pro
Val Tyr Val Thr 930 935 940Asp Val Thr Lys Gly Lys Leu Pro Asn Lys
Val Ile Val Ala Lys Lys945 950 955 960Gly Tyr His Asp Trp Ile Glu
Val Asp Asp Ser Phe Thr Phe Leu Phe 965 970 975Ser Leu Tyr Pro Asn
Asp Leu Ile Phe Ile Arg Gln Asn Pro Lys Lys 980 985 990Lys Ile Ser
Leu Lys Lys Arg Ile Glu Ser His Ser Ile Ser Asp Ser 995 1000
1005Lys Glu Val Gln Glu Ile His Ala Tyr Tyr Lys Gly Val Asp Ser
1010 1015 1020Ser Thr Ala Ala Ile Glu Phe Ile Ile His Asp Gly Ser
Tyr Tyr 1025 1030 1035Ala Lys Gly Val Gly Val Gln Asn Leu Asp Cys
Phe Glu Lys Tyr 1040 1045 1050Gln Val Asp Ile Leu Gly Asn Tyr Phe
Lys Val Lys Gly Glu Lys 1055 1060 1065Arg Leu Glu Leu Glu Thr Ser
Asp Ser Asn His Lys Gly Lys Asp 1070 1075 1080Val Asn Ser Ile Lys
Ser Thr Ser Arg 1085 109014120RNAArtificial sequencesynthesized
Variable Targeting domain-direct 141cgcuaaagag gaagaggaca
2014220RNAArtificial sequenceynthesized Variable Targeting
domain-reverse 142cgcuaaagag gaagaggaca 2014316RNABrevibacillus
laterosporus 143gcuauaguuc cuuacu 1614416RNABrevibacillus
laterosporus 144aucauaucau aucgag 1614516RNABrevibacillus
laterosporus 145gguaaguugc uauagu 1614616RNABrevibacillus
laterosporus 146cuugauauga uaugau 16147118RNABrevibacillus
laterosporus 147aagggcaaca gacccgaggc guuggggauc gccuagcccg
uuuuuacggg cucuccccau 60auucaaaaua augacagacg agcaccuugg agcauuuauu
uccgaggugc uuuuuuuu 11814859RNABrevibacillus laterosporus
148acuuucauuu uauauccaua uaucaucgaa gucaaucuca uuuaucuguc uauuuuaug
5914916RNALactobacillus reuteri 149guuuuagaug uacuuc
1615016RNALactobacillus rossiae 150guuuuagaug uauguc
1615116RNAPediococcus pentosaceus 151guuucagaag aguguu
1615217RNALactobacillus nodensis 152guuuuaguac ucucaag
1715317RNAArtificial Sequencesequence of Sulfurospirillum species
SCADC (Sspe) 153acgacacuuu acaacac 1715416RNAArtificial
Sequencesequence of Sulfurospirillum species SCADC (Sspe)
154guuccaguug cuaagc 1615516RNABifidobacterium thermophilum
155gcugagaauu agcauc 1615616RNALoktanella vestfoldensis
156guugcggcug gaccgc 1615716RNASphingomonas sanxanigenens
157guugccgcug gaccgc 1615816RNAEpilithonimonas tenax 158guugugaauu
gcuuuc 1615916RNASporocytophaga myxococcoides 159guugugauuu gcuuuc
1616017RNAPsychroflexus torquis 160guugugaauu gcuuuca
1716118RNALactobacillus reuteri 161gaaguacaga guuaaaau
1816218RNALactobacillus rossiae 162gacauacagc guuaaaau
1816318RNAPediococcus pentosaceus 163aacauucugu uuugaaau
1816419RNALactobacillus nodensis 164cuugagaacc uacuaaaau
1916517RNAArtificial Sequencesequence of Sulfurospirillum species
SCADC (Sspe) 165guguuguaaa gugucgu 1716613RNAArtificial
Sequencesequence of Sulfurospirillum species SCADC (Sspe)
166guucgaugca cug 1316716RNABifidobacterium thermophilum
167aaagcuaauu cucagu 1616817RNALoktanella vestfoldensis
168gcggucgagc cguuaac 1716917RNASphingomonas sanxanigenens
169guggucuggc gguuaac 1717016RNAEpilithonimonas tenax 170gaaagcaauu
cacaau 1617115RNASporocytophaga myxococcoides 171aaagcaaauc acaau
1517216RNAPsychroflexus torquis 172ugaagcaauu cacaau
16173108RNALactobacillus reuteri 173caaacaagug cuucagcaca
aguuucuacu uuugagucag uuaucugacc aauacauaaa 60agggaauuaa aucguucaca
ugagcguuuu aauucccuuu cucucuuu 108174105RNALactobacillus rossiae
174caagcaaggc uuucgagcca aguuuaaauc uuuggguucg cuauucggac
cguacauagu 60aaaagggcgu caucgaguuc aaauucgaug gcgcccuuua guuuu
105175107RNAPediococcus pentosaceus 175caaacaacgc uuuacgcgga
guuuacacau cugucccauu auaugggcau uacauaauaa 60aagagaaauc accuuuaagg
uugauuccuc uuuuucuugc guuccuu 10717651RNALactobacillus nodensis
176aaggauuuau uccgaauuua ccaccuauuu uuuaauaggu gguuuuuuuu u
51177119RNAArtificial Sequencesequence of Sulfurospirillum species
SCADC (Sspe) 177gagugaaagu auagcauuuu ucucaaaaua ucgcacuuuu
uuugcaagga aauaauuugc 60gaaaaaaaga gacaauuucg auacaagaaa gccccaucaa
gucgaugauu uguaaggcu 11917861RNAArtificial Sequencesequence of
Sulfurospirillum species SCADC (Sspe) 178aaaucaucga aaagauauau
agacccgccc aacugccuua ggcauggcgg gcuucuuucu 60u
61179195RNABifidobacterium thermophilum 179aaguacagca auuuauagcu
guaucugaau gcuaagcggu uagccgcagg ggagagcuuc 60ggcucucccc guucuuuucu
agucgcacgu gacacgccau agguagauug ccugugagca 120acucgagaau
aguaguauuc uucucuguaa aacgagacau cguugcugau gacuccauug
180acaaagaagg gaauu 195180140RNALoktanella vestfoldensis
180aagcauucga uugcaccaca uugaaacgca ggcuacggcc ugcguuuucu
uauuuccggu 60augguucguc uugucaaagu gucucagaga ugcccgcuuu cuaaaugugg
auugucgagg 120uucgcacccg uagcguuuuu 140181135RNASphingomonas
sanxanigenens 181aagcagccag ucugcaccag auaagggcgg cgcuccggcg
ccgccuuuuu uguuugcccg 60gaauguugag acaggcgcca aaccaucccc ccuccccucg
gggcgugcgg caaucgcugc 120uuucagcuca cucca
13518296RNAEpilithonimonas tenax 182aaggauuauu ccguugugaa
aacauuuagc accucgccua ucugcggggu auuuuuuauu 60uaauacauuc aaacaaugaa
gucgauugca uuuauu 9618391RNASporocytophaga myxococcoides
183aaggauuauu ccguugugaa aacauauaag ccgccucguc uuacaauacg
gggcuuuuuu 60uauuccccau ucacauaugg ccauccauuu u
9118495RNAPsychroflexus torquis 184aaggauuauu ccguugugaa aacauuuaaa
gcggccuuua cgggucgcuu ucuuuuuuua 60aagaacuuaa acuuaauaaa uucuuuacaa
acuca 95
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015
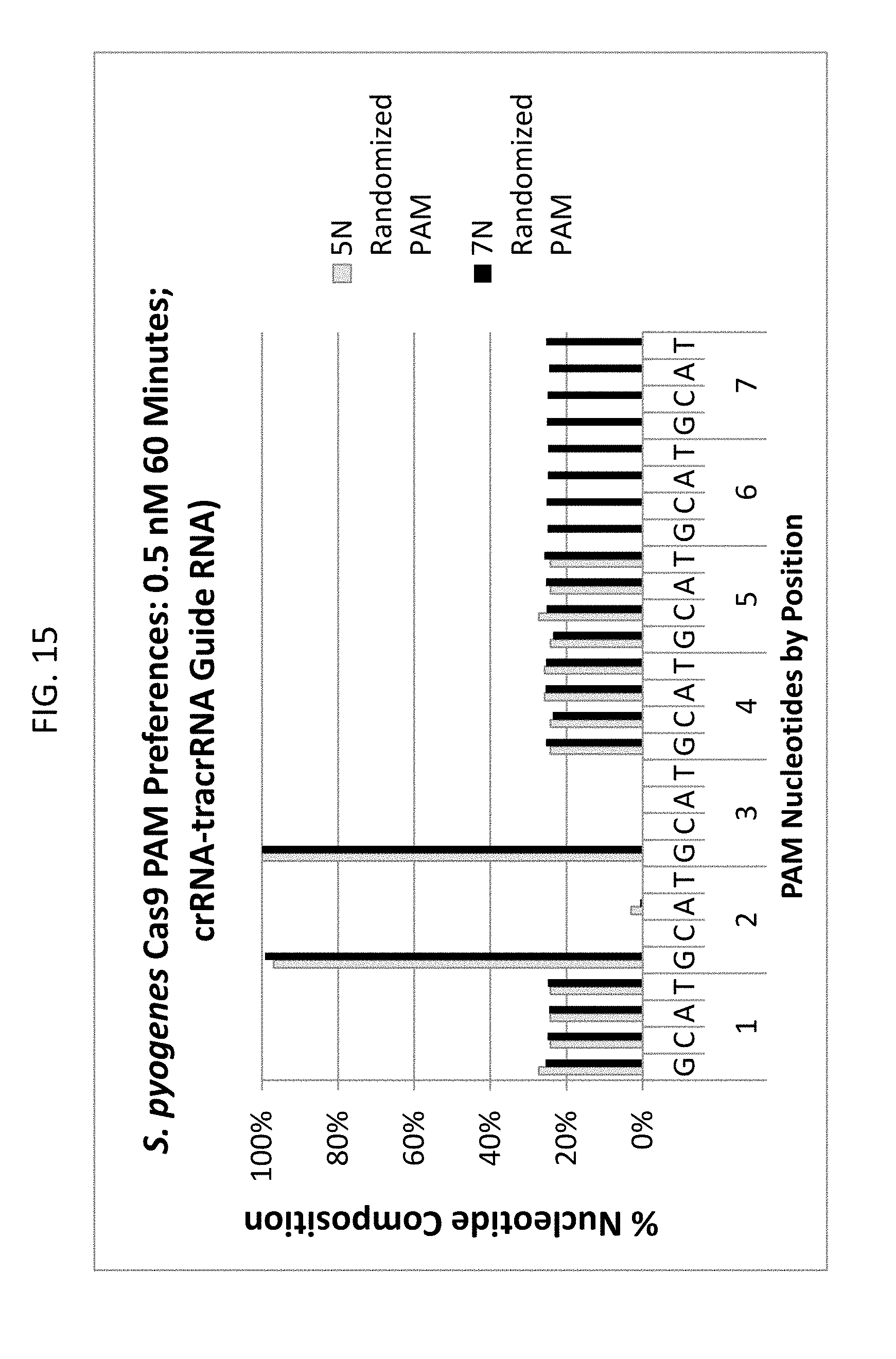
D00016

D00017

D00018

D00019

D00020

D00021
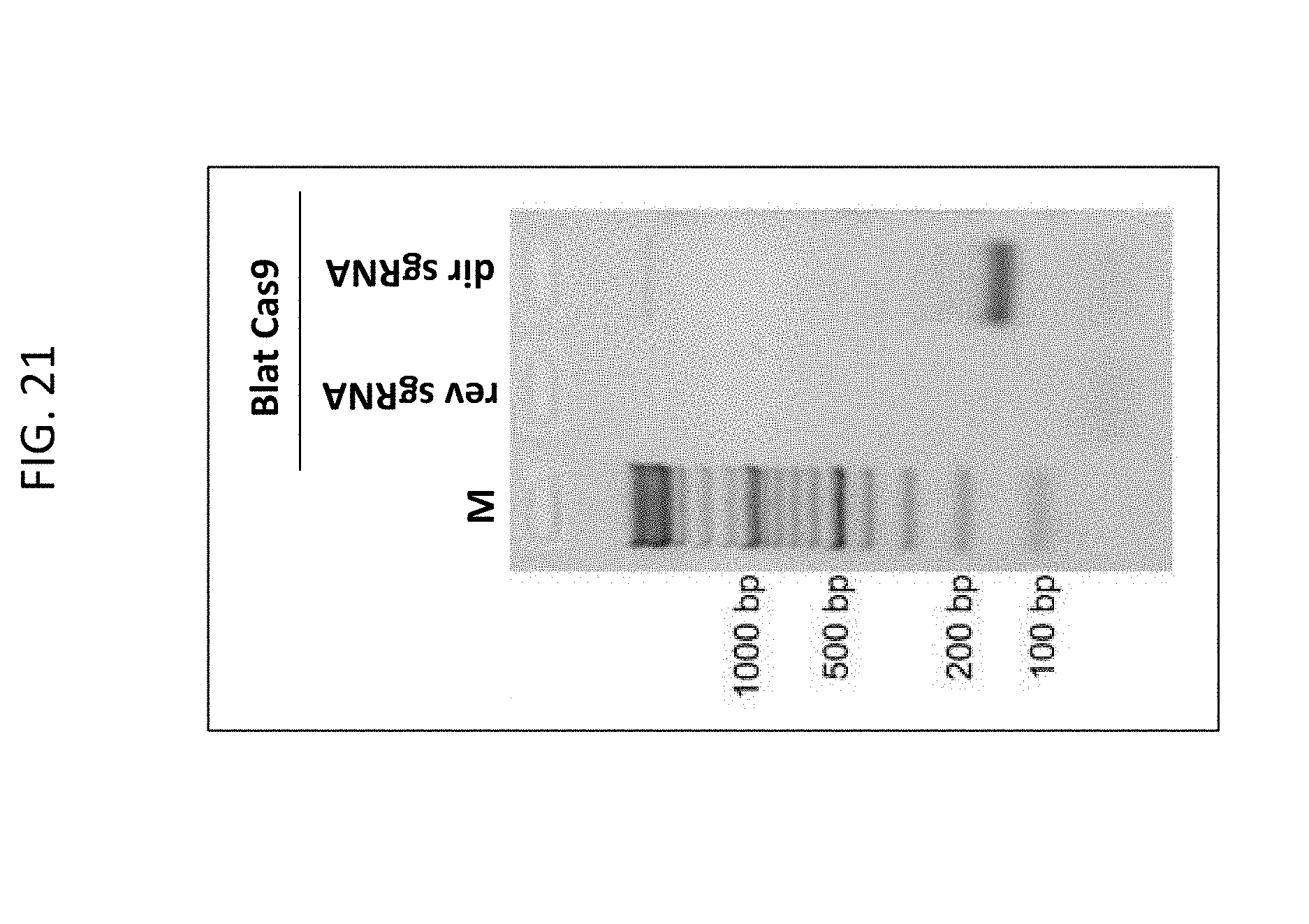
D00022

D00023

D00024
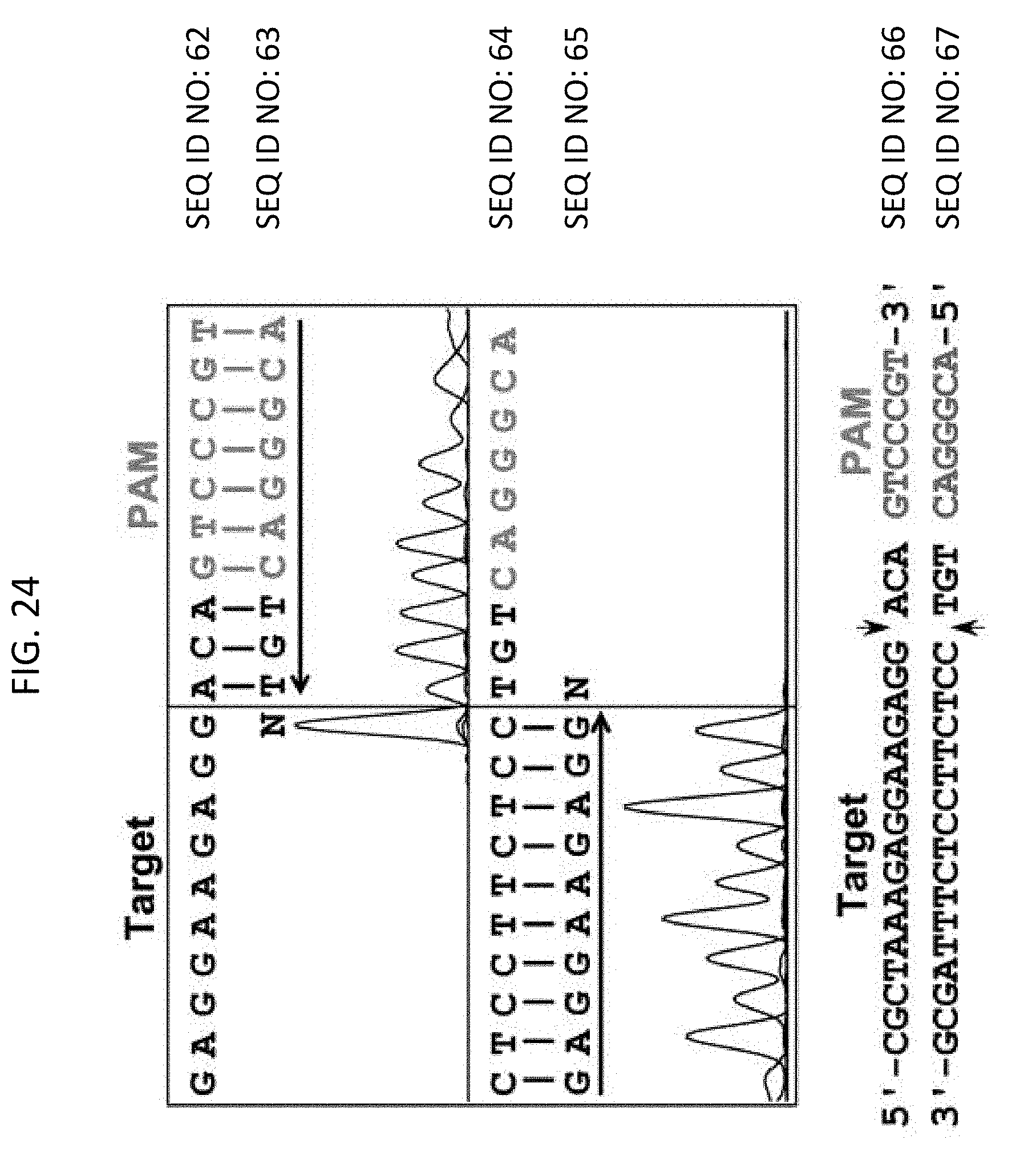
D00025
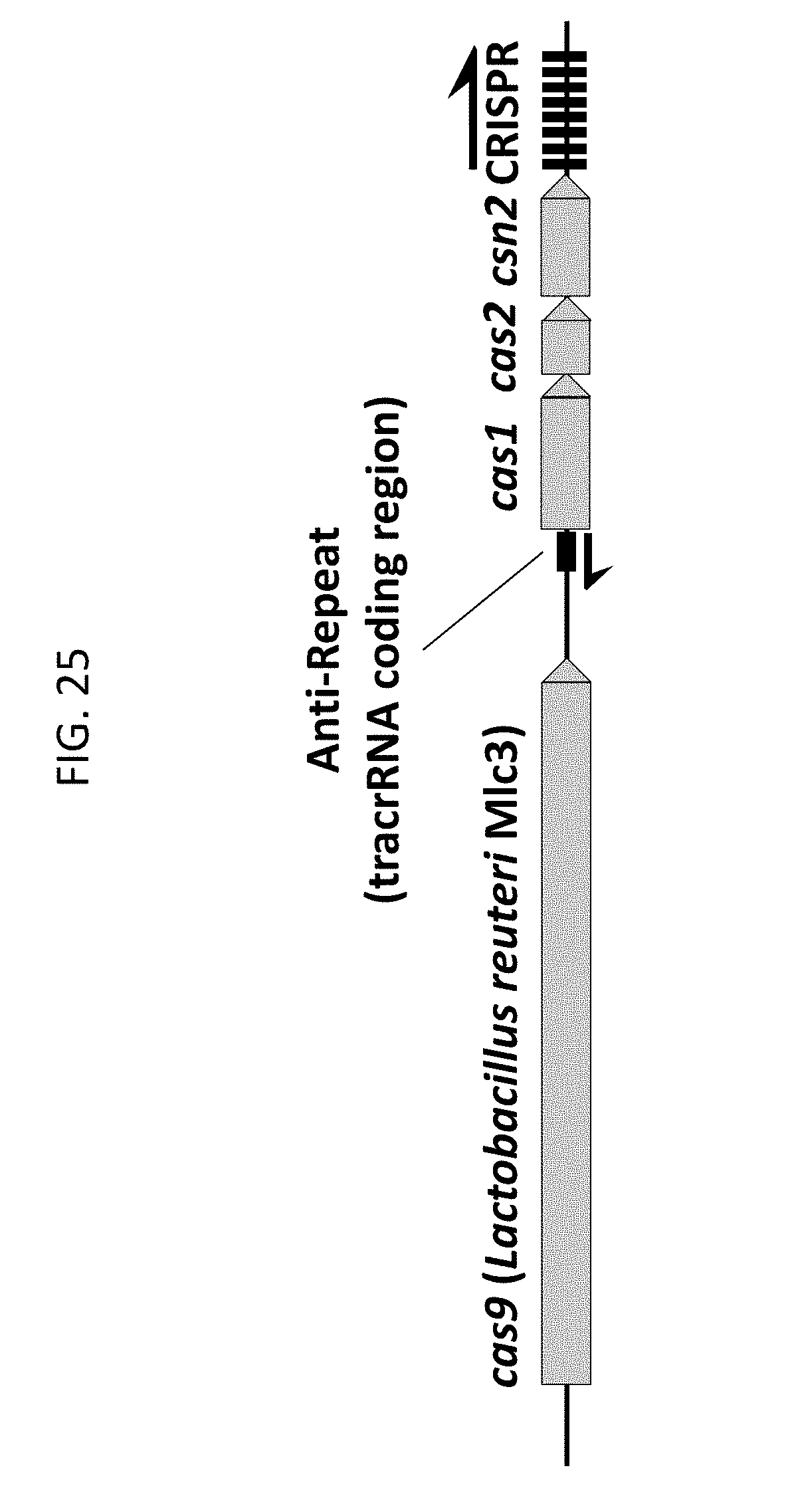
D00026

D00027

D00028
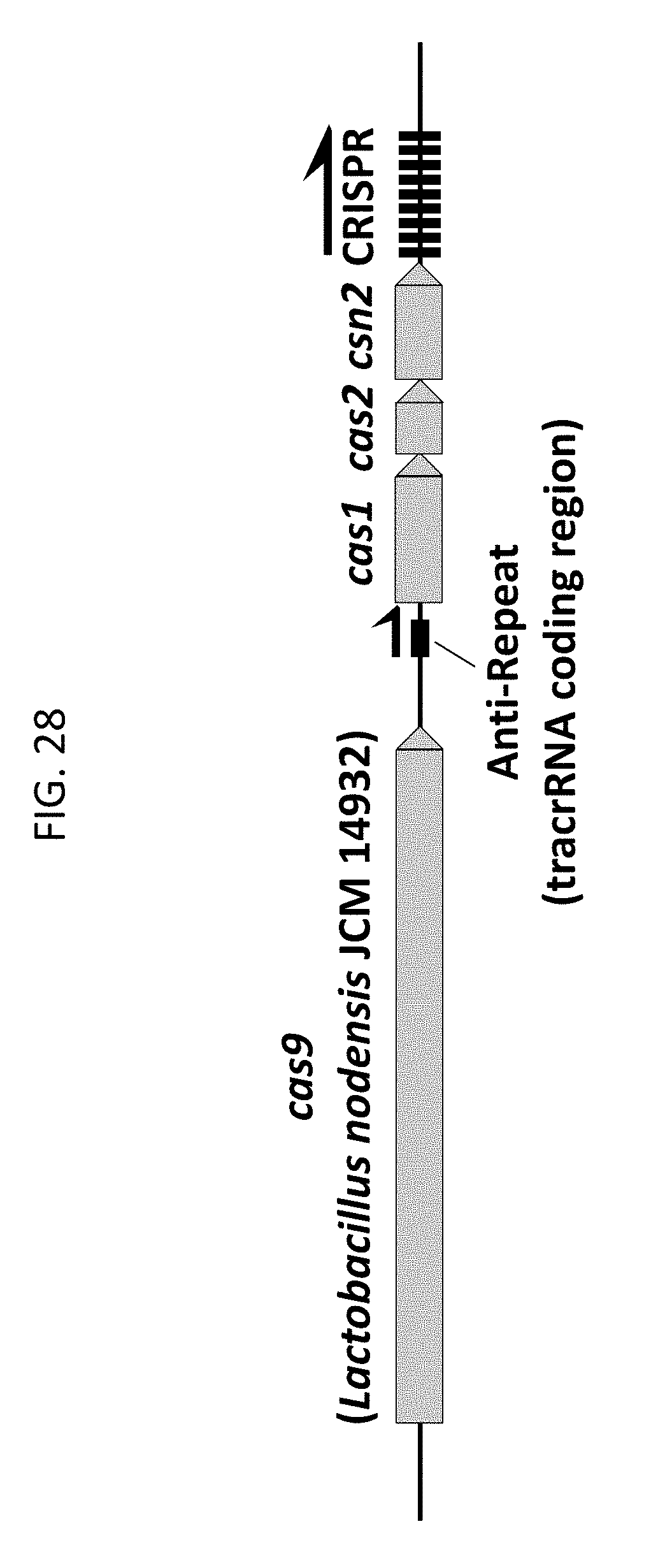
D00029

D00030
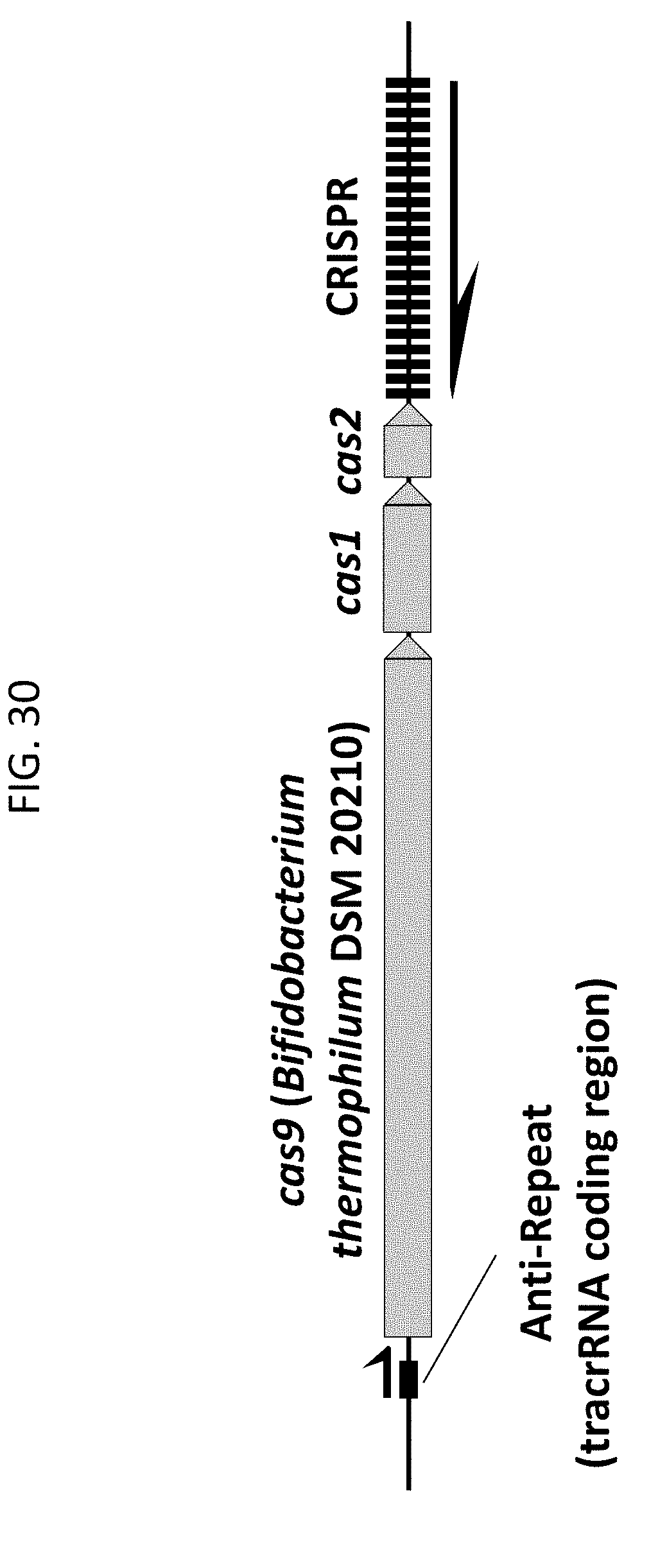
D00031

D00032
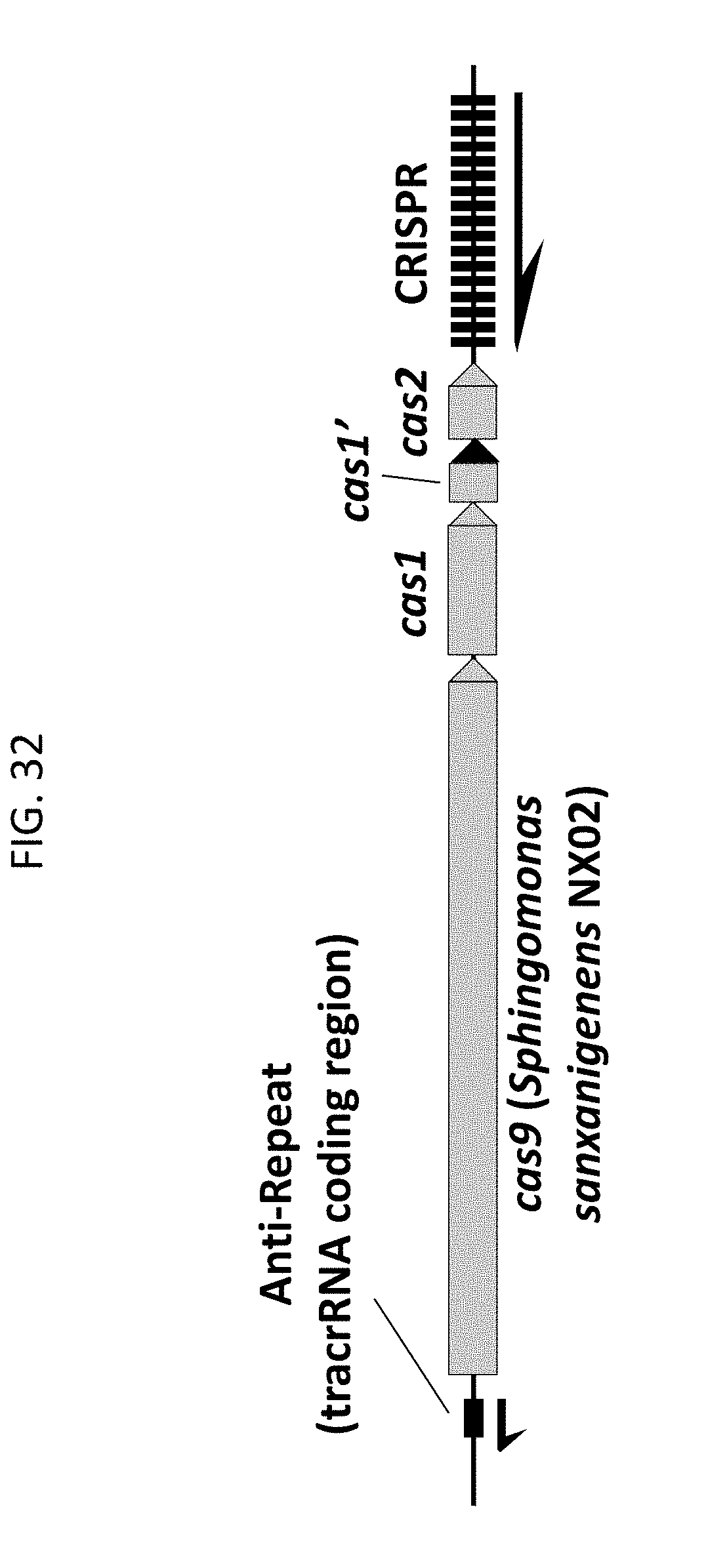
D00033

D00034

D00035

D00036
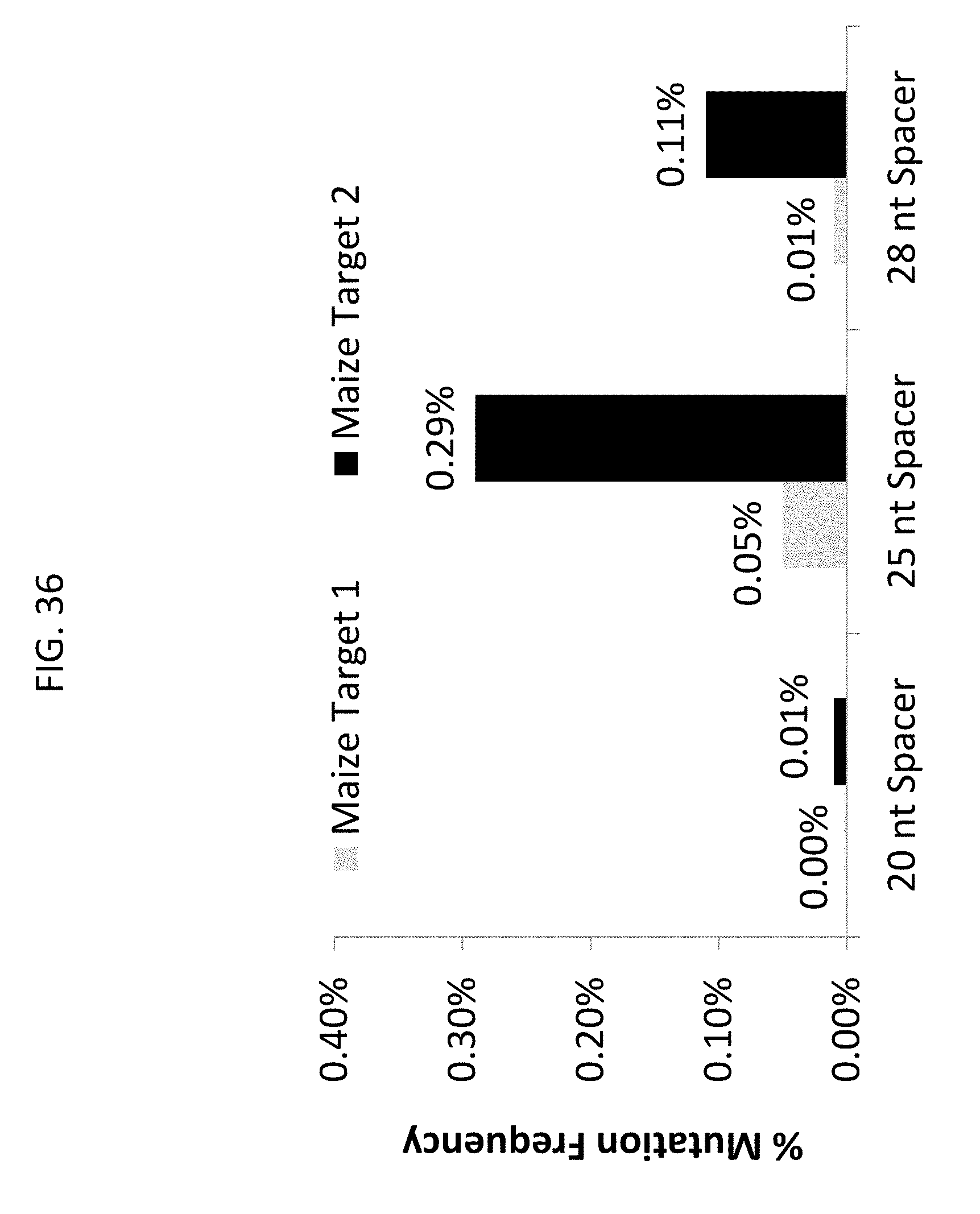
P00899
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.