Multiple Exon Skipping Compositions For Dmd
SAZANI; Peter ; et al.
U.S. patent application number 16/243926 was filed with the patent office on 2019-05-02 for multiple exon skipping compositions for dmd. This patent application is currently assigned to Sarepta Therapeutics, Inc.. The applicant listed for this patent is Sarepta Therapeutics, Inc.. Invention is credited to Ryszard KOLE, Peter SAZANI.
| Application Number | 20190127738 16/243926 |
| Document ID | / |
| Family ID | 41510869 |
| Filed Date | 2019-05-02 |

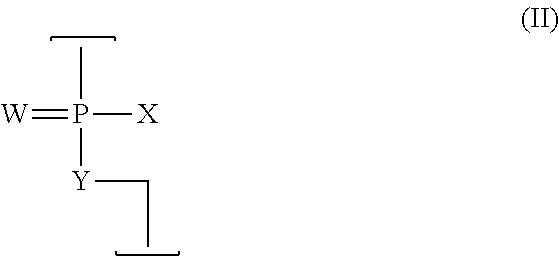










View All Diagrams
| United States Patent Application | 20190127738 |
| Kind Code | A1 |
| SAZANI; Peter ; et al. | May 2, 2019 |
MULTIPLE EXON SKIPPING COMPOSITIONS FOR DMD
Abstract
Provided are antisense molecules capable of binding to a selected target site in the human dystrophin gene to induce exon skipping, and methods of use thereof to treat muscular dystrophy.
| Inventors: | SAZANI; Peter; (Bothell, WA) ; KOLE; Ryszard; (Bellevue, WA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | Sarepta Therapeutics, Inc. Cambridge MA |
||||||||||
| Family ID: | 41510869 | ||||||||||
| Appl. No.: | 16/243926 | ||||||||||
| Filed: | January 9, 2019 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 16146328 | Sep 28, 2018 | |||
| 16243926 | ||||
| 15789862 | Oct 20, 2017 | |||
| 16146328 | ||||
| 15349778 | Nov 11, 2016 | |||
| 15789862 | ||||
| 14523610 | Oct 24, 2014 | |||
| 15349778 | ||||
| 12605276 | Oct 23, 2009 | 8871918 | ||
| 14523610 | ||||
| 61108416 | Oct 24, 2008 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12N 2310/3233 20130101; C12N 2310/3513 20130101; A61P 21/04 20180101; C12N 2310/11 20130101; C12N 2310/3341 20130101; C12N 15/111 20130101; C12N 15/113 20130101; A61P 21/00 20180101; C12N 2310/321 20130101; C12N 2310/331 20130101; C12N 2320/30 20130101; C12N 2320/33 20130101 |
| International Class: | C12N 15/113 20060101 C12N015/113; C12N 15/11 20060101 C12N015/11 |
Claims
1-65. (canceled)
66. An antisense oligonucleotide of 21 bases comprising a base sequence that is 100% complementary to 21 consecutive bases of exon 53 of the human dystrophin pre-mRNA, wherein the base sequence comprises 19 consecutive bases of TABLE-US-00009 (SEQ ID NO: 431) CTGTTGCCTCCGGTTCTGAAGGTGT,
wherein the antisense oligonucleotide is a morpholino oligomer, and wherein the antisense oligonucleotide induces exon 53 skipping; or a pharmaceutically acceptable salt thereof.
67. A pharmaceutical composition comprising an antisense oligonucleotide of 21 bases comprising a base sequence that is 100% complementary to 21 consecutive bases of exon 53 of the human dystrophin pre-mRNA, wherein the base sequence comprises 19 consecutive bases of TABLE-US-00010 (SEQ ID NO: 431) CTGTTGCCTCCGGTTCTGAAGGTGT,
wherein the antisense oligonucleotide is a morpholino oligomer, and wherein the antisense oligonucleotide induces exon 53 skipping; or a pharmaceutically acceptable salt thereof; and a pharmaceutically acceptable carrier.
Description
CROSS-REFERENCE TO RELATED APPLICATION
[0001] This application claims the benefit under 35 U.S.C. .sctn. 119(e) of U.S. Provisional Patent Application No. 61/108,416 filed Oct. 24, 2008; wherein this provisional application is incorporated herein by reference in its entirety.
STATEMENT REGARDING SEQUENCE LISTING
[0002] The Sequence Listing associated with this application is provided in text format in lieu of a paper copy, and is hereby incorporated by reference into the specification. The name of the text file containing the Sequence Listing is 120178_410_SEQUENCE_LISTING.txt. The text file is 157 KB, was created on Oct. 23, 2009 and is being submitted electronically via EFS-Web.
FIELD OF THE INVENTION
[0003] The present invention relates to novel antisense compounds and compositions suitable for facilitating exon skipping in the human dystrophin gene. It also provides methods for inducing exon skipping using the antisense compositions adapted for use in the methods of the invention.
BACKGROUND OF THE INVENTION
[0004] Antisense technologies are being developed using a range of chemistries to affect gene expression at a variety of different levels (transcription, splicing, stability, translation). Much of that research has focused on the use of antisense compounds to correct or compensate for abnormal or disease-associated genes in a wide range of indications. Antisense molecules are able to inhibit gene expression with specificity, and because of this, many research efforts concerning oligonucleotides as modulators of gene expression have focused on inhibiting the expression of targeted genes or the function of cis-acting elements. The antisense oligonucleotides are typically directed against RNA, either the sense strand (e.g., mRNA) or minus-strand in the case of some viral RNA targets. To achieve a desired effect of specific gene down-regulation, the oligonucleotides generally either promote the decay of the targeted mRNA, block translation of the mRNA or block the function of cis-acting RNA elements thereby effectively preventing either de novo synthesis of the target protein or replication of the viral RNA.
[0005] However, such techniques are not useful where the object is to up-regulate production of the native protein or compensate for mutations that induce premature termination of translation such as nonsense or frame-shifting mutations. In these cases, the defective gene transcript should not be subjected to targeted degradation or steric inhibition, so the antisense oligonucleotide chemistry should not promote target mRNA decay or block translation.
[0006] In a variety of genetic diseases, the effects of mutations on the eventual expression of a gene can be modulated through a process of targeted exon skipping during the splicing process. The splicing process is directed by complex multi-component machinery that brings adjacent exon-intron junctions in pre-mRNA into close proximity and performs cleavage of phosphodiester bonds at the ends of the introns with their subsequent reformation between exons that are to be spliced together. This complex and highly precise process is mediated by sequence motifs in the pre-mRNA that are relatively short semi-conserved RNA segments to which bind the various nuclear splicing factors that are then involved in the splicing reactions. By changing the way the splicing machinery reads or recognizes the motifs involved in pre-mRNA processing, it is possible to create differentially spliced mRNA molecules. It has now been recognized that the majority of human genes are alternatively spliced during normal gene expression, although the mechanisms involved have not been identified.
[0007] In cases where a normally functional protein is prematurely terminated because of mutations therein, a means for restoring some functional protein production through antisense technology has been shown to be possible through intervention during the splicing processes, and that if exons associated with disease-causing mutations can be specifically deleted from some genes, a shortened protein product can sometimes be produced that has similar biological properties of the native protein or has sufficient biological activity to ameliorate the disease caused by mutations associated with the exon (Sierakowska, Sambade et al. 1996; Wilton, Lloyd et al. 1999; van Deutekom, Bremmer-Bout et al. 2001; Lu, Mann et al. 2003; Aartsma-Rus, Janson et al. 2004). Kole et al. (U.S. Pat. Nos. 5,627,274; 5,916,808; 5,976,879; and 5,665,593) disclose methods of combating aberrant splicing using modified antisense oligonucleotide analogs that do not promote decay of the targeted pre-mRNA. Bennett et al (U.S. Pat. No. 6,210,892) describe antisense modulation of wild-type cellular mRNA processing also using antisense oligonucleotide analogs that do not induce RNAse H-mediated cleavage of the target RNA.
[0008] The process of targeted exon skipping is likely to be particularly useful in long genes where there are many exons and introns, where there is redundancy in the genetic constitution of the exons or where a protein is able to function without one or more particular exons. Efforts to redirect gene processing for the treatment of genetic diseases associated with truncations caused by mutations in various genes have focused on the use of antisense oligonucleotides that either: (1) fully or partially overlap with the elements involved in the splicing process; or (2) bind to the pre-mRNA at a position sufficiently close to the element to disrupt the binding and function of the splicing factors that would normally mediate a particular splicing reaction which occurs at that element.
[0009] Duchenne muscular dystrophy (DMD) is caused by a defect in the expression of the protein dystrophin. The gene encoding the protein contains 79 exons spread out over more than 2 million nucleotides of DNA. Any exonic mutation that changes the reading frame of the exon, or introduces a stop codon, or is characterized by removal of an entire out of frame exon or exons or duplications of one or more exons has the potential to disrupt production of functional dystrophin, resulting in DMD.
[0010] A less severe form of muscular dystrophy, Becker muscular dystrophy (BMD) has been found to arise where a mutation, typically a deletion of one or more exons, results in a correct reading frame along the entire dystrophin transcript, such that translation of mRNA into protein is not prematurely terminated. If the joining of the upstream and downstream exons in the processing of a mutated dystrophin pre-mRNA maintains the correct reading frame of the gene, the result is an mRNA coding for a protein with a short internal deletion that retains some activity resulting in a Becker phenotype.
[0011] Deletions of an exon or exons which do not alter the reading frame of a dystrophin protein give rise to a BMD phenotype, whereas an exon deletion that causes a frame-shift will give rise to DMD (Monaco, Bertelson et al. 1988). In general, dystrophin mutations including point mutations and exon deletions that change the reading frame and thus interrupt proper protein translation result in DMD. It should also be noted that some BMD and DMD patients have exon deletions covering multiple exons.
[0012] Although antisense molecules may provide a tool in the treatment of Duchenne Muscular Dystrophy (DMD), attempts to induce exon skipping using antisense molecules have had mixed success. Successful skipping of dystrophin exon 19 from the dystrophin pre-mRNA was achieved using a variety of antisense molecules directed at the flanking splice sites or motifs within the exon involved in exon definition as described by Errington et al., (Errington, Mann et al. 2003).
[0013] The first example of specific and reproducible exon skipping in the mdx mouse model was reported by Wilton et ael (Wilton, Lloyd et al. 1999). By directing an antisense molecule to the donor splice site, exon 23 skipping was induced in the dystrophin mRNA within 6 hours of treatment of the cultured cells. Wilton et al also describe targeting the acceptor region of the mouse dystrophin pre-mRNA with longer antisense oligonucleotides. While the first antisense oligonucleotide directed at the intron 23 donor splice site induced exon skipping in primary cultured myoblasts, this compound was found to be much less efficient in immortalized cell cultures expressing higher levels of dystrophin.
[0014] Despite these efforts, there remains a need for improved antisense oligomers targeted to multiple dystrophin exons and improved muscle delivery compositions and methods for DMD therapeutic applications.
BRIEF SUMMARY OF THE INVENTION
[0015] Embodiments of the present invention relate generally to antisense compounds capable of binding to a selected target to induce exon skipping, and methods of use thereof to induce exon skipping. In certain embodiments, it is possible to combine two or more antisense oligonucleotides of the present invention together to induce single or multiple exon skipping.
[0016] In certain embodiments, it is possible to improve exon skipping of a single or multiple exons by covalently linking together two or more antisense oligonucleotide molecules (see, e.g., Aartsma-Rus, Janson et al. 2004).
[0017] In certain embodiments, the antisense compounds of the present invention induce exon skipping in the human dystrophin gene, and thereby allow muscle cells to produce a functional dystrophin protein.
[0018] The antisense oligonucleotide compounds (also referred to herein as oligomers) of the present invention typically: (i) comprise morpholino subunits and phosphorus-containing intersubunit linkages joining a morpholino nitrogen of one subunit to a 5' exocyclic carbon of an adjacent subunit, (ii) contain between 10-40 nucleotide bases, preferably 20-35 bases (iii) comprise a base sequence effective to hybridize to at least 12 contiguous bases of a target sequence in dystrophin pre-mRNA and induce exon skipping.
[0019] In certain embodiments, the antisense compounds of the present invention may comprise phosphorus-containing intersubunit linkages joining a morpholino nitrogen of one subunit to a 5' exocyclic carbon of an adjacent subunit, in accordance with the following structure (I):
##STR00001##
wherein:
[0020] Y.sub.1 is --O--, --S--, --NH--, or --CH.sub.2--;
[0021] Z is O or S;
[0022] Pj is a purine or pyrimidine base-pairing moiety effective to bind, by base-specific hydrogen bonding, to a base in a polynucleotide; and
[0023] X is fluoro, optionally substituted alkyl, optionally substituted alkoxy, optionally substituted thioalkoxy, amino, optionally substituted alkylamino, or optionally substituted heterocyclyl.
[0024] In certain embodiments, the above intersubunit linkages, which are uncharged, may be interspersed with linkages that are positively charged at physiological pH, where the total number of positively charged linkages is between 2 and no more than half of the total number of linkages. For example, the positively charged linkages may have the above structure in which X is optionally substituted 1-piperazinyl. In other embodiments, the positively charged linkages may have the above structure in which X is substituted 1-piperazynyl, wherein the 1-piperazynyl is substituted at the 4-position with an optionally substituted alkyl guanidynyl moiety.
[0025] Where the antisense compound administered is effective to target a splice site of preprocessed human dystrophin, it may have a base sequence complementary to a target region containing at least 12 contiguous bases in a preprocessed messenger RNA (mRNA) human dystrophin transcript. Exemplary antisense sequences include those identified by SEQ ID NOS: 1 to 569 and 612 to 633.
[0026] In certain embodiments, an antisense sequence of the present invention is contained within:
[0027] (a) any of the sequences identified by SEQ ID NOS: 1-20, preferably SEQ ID NOS: 4, 8, 11 and 12, and more preferably SEQ ID NO:12 for use in producing skipping of exon 44 in the processing of human dystrophin pre-processed mRNA;
[0028] (b) any of the sequences identified by SEQ ID NOS: 21-76 and 612 to 624, preferably SEQ ID NOS: 27, 29, 34 and 39, and more preferably SEQ ID NO: 34 for use in producing skipping of exon 45 in the processing of human dystrophin pre-processed mRNA;
[0029] (c) any of the sequences identified by SEQ ID NOS: 77-125, preferably SEQ ID NOS: 21 to 53, and more preferably SEQ ID NOS: 82, 84-87, 90 96, 98, 99 and 101, for use in producing skipping of exon 46 in the processing of human dystrophin pre-processed mRNA;
[0030] (d) any of the sequences identified by SEQ ID NOS: 126-169, preferably SEQ ID NOS: 126-149, and more preferably SEQ ID NOS: 126, 128-130, 132, 144 and 146-149, for use in producing skipping of exon 47 in the processing of human dystrophin pre-processed mRNA;
[0031] (e) any of the sequences identified by SEQ ID NOS: 170-224 and 634, preferably SEQ ID NOS: 170-201 and 634, and more preferably SEQ ID NOS: 176, 178, 181-183, 194 and 198-201, for use in producing skipping of exon 48 in the processing of human dystrophin pre-processed mRNA;
[0032] (f) any of the sequences identified by SEQ ID NOS: 225-266, preferably SEQ ID NOS: 225-248, and more preferably SEQ ID NOS: 227, 229, 234, 236, 237 and 244-248, for use in producing skipping of exon 49 in the processing of human dystrophin pre-processed mRNA;
[0033] (g) any of the sequences identified by SEQ ID NOS: 267-308, preferably SEQ ID NOS: 277, 287 and 290, and more preferably SEQ ID NO: 287, for use in producing skipping of exon 50 in the processing of human dystrophin pre-processed mRNA; [0034] (h) any of the sequences identified by SEQ ID NOS: 309-371, preferably SEQ ID NOS: 324, 326 and 327, and more preferably SEQ ID NO: 327 for use in producing skipping of exon 51 in the processing of human dystrophin pre-processed mRNA; [0035] (i) any of the sequences identified by SEQ ID NOS: 372-415, preferably SEQ ID NOS: 372-397, and more preferably SEQ ID NOS: 379-382, 384, 390 and 392-395 for use in producing skipping of exon 52 in the processing of human dystrophin pre-processed mRNA;
[0036] (j) any of the sequences identified by SEQ ID NOS: 416-475 and 625-633, preferably SEQ ID NOS: 428, 429 and 431, and more preferably SEQ ID NO: 429, for use in producing skipping of exon 53 in the processing of human dystrophin pre-processed mRNA;
[0037] (k) any of the sequences identified by SEQ ID NOS: 476-519, preferably SEQ ID NOS: 476-499, and more preferably SEQ ID NOS: 479-482, 484, 489 and 491-493, for use in producing skipping of exon 54 in the processing of human dystrophin pre-processed mRNA; and
[0038] (l) any of the sequences identified by SEQ ID NOS: 520-569 and 635, preferably SEQ ID NOS: 520-546 and 635, and more preferably SEQ ID NOS: 524-528, 537, 539, 540, 542 and 544, for use in producing skipping of exon 55 in the processing of human dystrophin pre-processed mRNA;
[0039] In certain embodiments, the compound may be conjugated to an arginine-rich polypeptide effective to promote uptake of the compound into cells. Exemplary peptides include those identified by SEQ ID NOS: 570 to 578, among others described herein.
[0040] In one exemplary embodiment, the arginine-rich polypeptide is covalently coupled at its N-terminal or C-terminal residue to the 3' or 5' end of the antisense compound. Also in an exemplary embodiment, the antisense compound is composed of morpholino subunits and phosphorus-containing intersubunit linkages joining a morpholino nitrogen of one subunit to a 5' exocyclic carbon of an adjacent subunit.
[0041] In general, the peptide-oligomer conjugate may further comprise a homing peptide which is selective for a selected mammalian tissue, i.e., the same tissue being targeted by the cell-penetrating peptide. The conjugate may be of the form: cell penetrating peptide--homing peptide--antisense oligomer, or, more preferably, of the form: homing peptide--cell penetrating peptide--antisense oligomer. For example, a peptide conjugate compound for use in treating Duchenne muscular dystrophy, as described above, can further comprise a homing peptide which is selective for muscle tissue, such as the peptide having the sequence identified as SEQ ID NO: 579, conjugated to the cell-penetrating peptide. Exemplary conjugates of this type include those represented herein as CP06062-MSP-PMO (cell penetrating peptide--homing peptide--antisense oligomer) and as MSP-CP06062-PMO (homing peptide--cell penetrating peptide--antisense oligomer) (see SEQ ID NOs: 580-583).
[0042] In some embodiments, the peptide is conjugated to the oligomer via a linker moiety. In certain embodiments the linker moiety may comprise an optionally substituted piperazynyl moiety. In other embodiments, the linker moiety may further comprise a beta alanine and/or a 6-aminohexanoic acid subunit. In yet other embodiments, the peptide is conjugated directly to the oligomer without a linker moiety.
[0043] Conjugation of the peptide to the oligomer may be at any position suitable for forming a covalent bond between the peptide and the oligomer or between the linker moiety and the oligomer. For example, in some embodiments conjugation of the peptide may be at the 3' end of the oligomer. In other embodiments, conjugation of the peptide to the oligomer may be at the 5' end of the oligomer. In yet other embodiments, the peptide may be conjugated to the oligomer through any of the intersubunit linkages.
[0044] In some embodiments, the peptide is conjugated to the oligomer at the 5' end of the oligomer. In embodiments comprising phosphorus-containing intersubunit linkages, the peptide may be conjugated to the oligomer via a covalent bond to the phosphorous of the terminal linkage group. Conjugation in this manner may be with or without the linker moiety described above.
[0045] In yet other embodiments, the peptide may be conjugated to the oligomer at the 3' end of the oligomer. In some further embodiments, the peptide may be conjugated to the nitrogen atom of the 3' terminal morpolino group of the oligomer. In this respect, the peptide may be conjugated to the oligomer directly or via the linker moiety described above.
[0046] In some embodiments, the oligomer may be conjugated to a moiety that enhances the solubility of the oligomer in aqueous medium. In some embodiments, the moiety that enhances solubility of the oligomer in aqueous medium is a polyethyleneglycol. In yet further embodiments, the moiety that enhances solubility of the oligomer in aqueous medium is triethylene glycol. For example, in some embodiments the moiety that enhances solubility in aqueous medium may be conjugated to the oligomer at the 5' end of the oligomer. Conjugation of the moiety that enhances solubility of the oligomer in aqueous medium to the oligomer may be either directly or through the linker moiety described above.
[0047] Certain embodiments of the present invention provide antisense molecules selected and or adapted to aid in the prophylactic or therapeutic treatment of a genetic disorder comprising at least an antisense molecule in a form suitable for delivery to a patient.
[0048] Certain embodiments of the invention provide methods for treating a patient suffering from a genetic disease wherein there is a mutation in a gene encoding a particular protein and the affect of the mutation can be abrogated by exon skipping, comprising the steps of: (a) selecting an antisense molecule in accordance with the methods described herein; and (b) administering the molecule to a patient in need of such treatment. The present invention also includes the use of purified and isolated antisense oligonucleotides of the invention, for the manufacture of a medicament for treatment of a genetic disease.
[0049] Certain embodiments provide a method of treating muscular dystrophy, such as a condition characterized by Duchenne muscular dystrophy, which method comprises administering to a patient in need of treatment an effective amount of an appropriately designed antisense oligonucleotide, as described herein, relevant to the particular genetic lesion in that patient. Further, certain embodiments provide a method for prophylactically treating a patient to prevent or at least minimize muscular dystrophy, including Duchene muscular dystrophy, comprising the step of: administering to the patient an effective amount of an antisense oligonucleotide or a pharmaceutical composition comprising one or more of these biological molecules.
[0050] Certain embodiments relate to methods of treating muscular dystrophy in a subject, comprising administering to the subject an effective amount of a substantially uncharged antisense compound containing 20-35 morpholino subunits linked by phosphorus-containing intersubunit linkages joining a morpholino nitrogen of one subunit to a 5' exocyclic carbon of an adjacent subunit, comprising a sequence selected from the group consisting SEQ ID NOS:1 to 569 and 612 to 635, and capable of forming with the complementary mRNA sequence in a dystrophin-gene exon a heteroduplex structure between said compound and mRNA having a Tm of at least 45.degree. C., wherein the exon is selected from the group consisting of exons 44-55.
[0051] In certain embodiments, the muscular dystrophy is Duchenne's muscular dystrophy (DMD). In certain embodiments, the muscular dystrophy is Becker muscular dystrophy (BMD).
[0052] In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 1-20, and the exon is exon 44. In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 21-76 and 612 to 624, and the exon is exon 45.
[0053] In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 77-125, and the exon is exon 46. In certain embodiments, the sequence selected from the group consisting SEQ ID NOS: 126-169, and the exon is exon 47.
[0054] In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 170-224 and 634, and the exon is exon 48. In certain embodiments, the sequence selected from the group consisting SEQ ID NOS: 225-266, and the exon is exon 49.
[0055] In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 267-308, and the exon is exon 50. In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 309-371, and the exon is exon 51.
[0056] In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 372-415, and the exon is exon 52. In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 416-475 and 625-633, and the exon is exon 53. In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 476-519, and the exon is exon 54. In certain embodiments, the sequence is selected from the group consisting SEQ ID NOS: 520-569 and 635, and the exon is exon 55. In certain embodiments, the sequence comprises or consists essentially of SEQ ID NO:287.
[0057] Certain embodiments provide kits for treating a genetic disease, which kits comprise at least an antisense oligonudeotide of the present invention, packaged in a suitable container and instructions for its use.
[0058] These and other objects and features will be more fully understood when the following detailed description of the invention is read in conjunction with the figures.
BRIEF DESCRIPTION OF THE FIGURES
[0059] FIG. 1A shows an exemplary morpholino oligomer structure with a phosphorodiamidate linkage;
[0060] FIG. 1B shows a conjugate of an arginine-rich peptide and an antisense oligomer, in accordance with an embodiment of the invention;
[0061] FIG. 1C shows a conjugate as in FIG. 1B, wherein the backbone linkages contain one or more positively charged groups;
[0062] FIGS. 1D-G show the repeating subunit segment of exemplary morpholino oligonucleotides, designated D through G.
[0063] FIG. 2A shows the relative location and results of an antisense oligomer exon 51 scan designed to induce skipping of human dystrophin exon 51.
[0064] FIG. 2B-C shows the relative activity in cultured human rhabdomyosarcoma (RD) cells and human primary skeletal muscle cells of the three best oligomers selected from the exon 51 scan (SEQ ID NOs: 324, 326 and 327) relative to sequences (AVI-5658; SEQ ID NO: 588 and h51AON1; SEQ ID NO:594) that are effective at inducing exon 51 skipping. FIG. 2D shows the relative location within exon 51 of three selected oligomers compared to certain sequences.
[0065] FIG. 3A shows the relative location and results of an antisense oligomer exon 50 scan designed to induce skipping of human dystrophin exon 50 compared to other sequences that induce exon 50 skipping.
[0066] FIG. 3B shows the relative location and activity of antisense sequences selected from the exon 50 scan (SEQ ID NOS: 277, 287, 290 and 291) compared to other sequences (SEQ ID NOS: 584 and 585).
[0067] FIG. 4A shows the relative location and results of an antisense oligomer exon 53 scan designed to induce skipping of human dystrophin exon 53.
[0068] FIG. 4B shows the relative location of certain sequences used to compare the exon-skipping activity of those oligomers selected as being most active in the exon 53 scan.
[0069] FIGS. 4C-F show the results of dose-ranging studies, summarized in FIG. 4G, using the oligomers selected as being most efficacious in the exon 53 scan (SEQ ID NOS:422, 428, 429 and 431).
[0070] FIGS. 4H and 4I show the relative activity of certain sequences (SEQ ID NOS: 608-611) compared to the activity of the most active exon 53-skipping oligomer (SEQ ID NO:429) in both RD cells and human primary skeletal muscle cells.
[0071] FIG. 5A shows the relative location and results of an antisense oligomer exon 44 scan designed to induce skipping of human dystrophin exon 44.
[0072] FIG. 5B shows the relative location within exon 44 of certain sequences used to compare the exon-skipping activity to those oligomers selected as being most active in the exon 44 scan.
[0073] FIGS. 5C-G show the results of dose-ranging studies, summarized in FIG. 5H, using the oligomers selected as being most efficacious in the exon 44 scan (SEQ ID NOS: 4, 8, 11, 12 and 13).
[0074] FIGS. 5I and 5J show the relative activity of certain sequences (SEQ ID NOS: 600-603) compared to the activity of the most active exon 53-skipping oligomer (SEQ ID NO:12) in both RD cells and human primary skeletal muscle cells.
[0075] FIG. 6A shows the relative location and results of an antisense oligomer exon 45 scan designed to induce skipping of human dystrophin exon 45. FIG. 6B shows the relative location within exon 45 of certain sequences used to compare the exon-skipping activity to those oligomers selected as being most active in the exon 45 scan.
[0076] FIGS. 6C-F show the results of dose-ranging studies, summarized in FIG. 6H, using the oligomers selected as being most efficacious in the exon 45 scan (SEQ ID NOS: 27, 29, 34 and 39). FIG. 6G uses a relatively inactive oligomer (SEQ ID NO: 49) as a negative control.
[0077] FIGS. 6I and 6J show the relative activity of certain sequences (SEQ ID NOS: 604-607) compared to the activity of the most active exon 53-skipping oligomer (SEQ ID NO: 34) in both RD cells and human primary skeletal muscle cells.
DETAILED DESCRIPTION OF THE INVENTION
[0078] Embodiments of the present invention relate generally to improved antisense compounds, and methods of use thereof, which are specifically designed to induce exon skipping in the dystrophin gene. Dystrophin plays a vital role in muscle function, and various muscle-related diseases are characterized by mutated forms of this gene. Hence, in certain embodiments, the improved antisense compounds described herein induce exon skipping in mutated forms of the human dystrophin gene, such as the mutated dystrophin genes found in Duchenne's muscular dystrophy (DMD) and Becker's muscular dystrophy (BMD).
[0079] Due to aberrant mRNA splicing events caused by mutations, these mutated human dystrophin genes either express defective dystrophin protein or express no measurable dystrophin at all, a condition that leads to various forms of muscular dystrophy. To remedy this condition, the antisense compounds of the present invention typically hybridize to selected regions of a pre-processed RNA of a mutated human dystrophin gene, induce exon skipping and differential splicing in that otherwise aberrantly spliced dystrophin mRNA, and thereby allow muscle cells to produce an mRNA transcript that encodes a functional dystrophin protein. In certain embodiments, the resulting dystrophin protein is not necessarily the "wild-type" form of dystrophin, but is rather a truncated, yet functional or semi-functional, form of dystrophin.
[0080] By increasing the levels of functional dystrophin protein in muscle cells, these and related embodiments may be useful in the prophylaxis and treatment of muscular dystrophy, especially those forms of muscular dystrophy, such as DMD and BMD, that are characterized by the expression of defective dystrophin proteins due to aberrant mRNA splicing. The specific oligomers described herein further provide improved, dystrophin-exon-specific targeting over other oligomers in use, and thereby offer significant and practical advantages over alternate methods of treating relevant forms of muscular dystrophy.
[0081] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by those of ordinary skill in the art to which the invention belongs. Although any methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, preferred methods and materials are described. For the purposes of the present invention, the following terms are defined below.
Definitions
[0082] The articles "a" and "an" are used herein to refer to one or to more than one (i.e., to at least one) of the grammatical object of the article. By way of example, "an element" means one element or more than one element.
[0083] By "about" is meant a quantity, level, value, number, frequency, percentage, dimension, size, amount, weight or length that varies by as much as 30, 25, 20, 25, 10, 9, 8, 7, 6, 5, 4, 3, 2 or 1% to a reference quantity, level, value, number, frequency, percentage, dimension, size, amount, weight or length.
[0084] By "coding sequence" is meant any nucleic acid sequence that contributes to the code for the polypeptide product of a gene. By contrast, the term "non-coding sequence" refers to any nucleic acid sequence that does not contribute to the code for the polypeptide product of a gene.
[0085] Throughout this specification, unless the context requires otherwise, the words "comprise," "comprises," and "comprising" will be understood to imply the inclusion of a stated step or element or group of steps or elements but not the exclusion of any other step or element or group of steps or elements.
[0086] By "consisting of" is meant including, and limited to, whatever follows the phrase "consisting of." Thus, the phrase "consisting of" indicates that the listed elements are required or mandatory, and that no other elements may be present. By "consisting essentially of" is meant including any elements listed after the phrase, and limited to other elements that do not interfere with or contribute to the activity or action specified in the disclosure for the listed elements. Thus, the phrase "consisting essentially of" indicates that the listed elements are required or mandatory, but that other elements are optional and may or may not be present depending upon whether or not they materially affect the activity or action of the listed elements.
[0087] The terms "complementary" and "complementarity" refer to polynucleotides (i.e., a sequence of nucleotides) related by the base-pairing rules. For example, the sequence "A-G-T," is complementary to the sequence "T-C-A." Complementarity may be "partial," in which only some of the nucleic acids' bases are matched according to the base pairing rules. Or, there may be "complete" or "total" complementarity between the nucleic acids. The degree of complementarity between nucleic acid strands has significant effects on the efficiency and strength of hybridization between nucleic acid strands. While perfect complementarity is often desired, some embodiments can include one or more but preferably 6, 5, 4, 3, 2, or 1 mismatches with respect to the target RNA. Variations at any location within the oligomer are included. In certain embodiments, variations in sequence near the termini of an oligomer are generally preferable to variations in the interior, and if present are typically within about 6, 5, 4, 3, 2, or 1 nucleotides of the 5' and/or 3' terminus.
[0088] The terms "cell penetrating peptide" or "CPP" are used interchangeably and refer to cationic cell penetrating peptides, also called transport peptides, carrier peptides, or peptide transduction domains. The peptides, as shown herein, have the capability of inducing cell penetration within 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% of cells of a given cell culture population, including all integers in between, and allow macromolecular translocation within multiple tissues in vivo upon systemic administration.
[0089] The terms "antisense oligomer" or "antisense compound" are used interchangeably and refer to a sequence of cyclic subunits, each bearing a base-pairing moiety, linked by intersubunit linkages that allow the base-pairing moieties to hybridize to a target sequence in a nucleic acid (typically an RNA) by Watson-Crick base pairing, to form a nucleic acid:oligomer heteroduplex within the target sequence. The cyclic subunits are based on ribose or another pentose sugar or, in a preferred embodiment, a morpholino group (see description of morpholino oligomers below).
[0090] Such an antisense oligomer can be designed to block or inhibit translation of mRNA or to inhibit natural pre-mRNA splice processing, and may be said to be "directed to" or "targeted against" a target sequence with which it hybridizes. In certain embodiments, the target sequence includes a region including an AUG start codon of an mRNA, a 3' or 5' splice site of a pre-processed mRNA, or a branch point. The target sequence may be within an exon or within an intron. The target sequence for a splice site may include an mRNA sequence having its 5' end 1 to about 25 base pairs downstream of a normal splice acceptor junction in a preprocessed mRNA. A preferred target sequence for a splice is any region of a preprocessed mRNA that includes a splice site or is contained entirely within an exon coding sequence or spans a splice acceptor or donor site. An oligomer is more generally said to be "targeted against" a biologically relevant target, such as a protein, virus, or bacteria, when it is targeted against the nucleic acid of the target in the manner described above. Included are antisense oligomers that comprise, consist essentially of, or consist of one or more of SEQ ID NOS:1 to 569 and 612 to 635. Also included are variants of these antisense oligomers, including variant oligomers having 80%, 85%, 90%, 95%, 97%, 98%, or 99% (including all integers in between) sequence identity or sequence homology to any one of SEQ ID NOS:1 to 569 and 612 to 635, and/or variants that differ from these sequences by about 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 nucleotides, preferably those variants that induce exon skipping of one or more selected human dystrophin exons. Also included are oligomers of any on or more of SEQ ID NOS:584-611 and 634-635, which comprise a suitable number of charged linkages, as described herein, e.g. up to about 1 per every 2-5 uncharged linkages, such as about 4-5 per every 10 uncharged linkages, and/or which comprise an Arg-rich peptide attached thereto, as also described herein.
[0091] The terms "morpholino oligomer" or "PMO" (phosphoramidate- or phosphorodiamidate morpholino oligomer) refer to an oligonucleotide analog composed of morpholino subunit structures, where (i) the structures are linked together by phosphorus-containing linkages, one to three atoms long, preferably two atoms long, and preferably uncharged or cationic, joining the morpholino nitrogen of one subunit to a 5' exocyclic carbon of an adjacent subunit, and (ii) each morpholino ring bears a purine or pyrimidine base-pairing moiety effective to bind, by base specific hydrogen bonding, to a base in a polynucleotide. See, e.g., the structure in FIG. 1A, which shows a preferred phosphorodiamidate linkage type. Variations can be made to this linkage as long as they do not interfere with binding or activity. For example, the oxygen attached to phosphorus may be substituted with sulfur (thiophosphorodiamidate). The 5' oxygen may be substituted with amino or lower alkyl substituted amino. The pendant nitrogen attached to phosphorus may be unsubstituted, monosubstituted, or disubstituted with (optionally substituted) lower alkyl. See also the discussion of cationic linkages below. The synthesis, structures, and binding characteristics of morpholino oligomers are detailed in U.S. Pat. Nos. 5,698,685, 5,217,866, 5,142,047, 5,034,506, 5,166,315, 5,521,063, and 5,506,337, and PCT Appn. No. PCT/US07/11435 (cationic linkages), all of which are incorporated herein by reference.
[0092] The purine or pyrimidine base pairing moiety is typically adenine, cytosine, guanine, uracil, thymine or inosine. Also included are bases such as pyridin-4-one, pyridin-2-one, phenyl, pseudouracil, 2,4,6-trime115thoxy benzene, 3-methyl uracil, dihydrouridine, naphthyl, aminophenyl, 5-alkylcytidines (e.g., 5-methylcytidine), 5-alkyluridines (e.g., ribothymidine), 5-halouridine (e.g., 5-bromouridine) or 6-azapyrimidines or 6-alkylpyrimidines (e.g. 6-methyluridine), propyne, quesosine, 2-thiouridine, 4-thiouridine, wybutosine, wybutoxosine, 4-acetyltidine, 5-(carboxyhydroxymethyl)uridine, 5'-carboxymethylaminomethyl-2-thiouridine, 5-carboxymethylaminomethyluridine, .beta.-D-galactosylqueosine, 1-methyladenosine, 1-methylinosine, 2,2-dimethylguanosine, 3-methylcytidine, 2-methyladenosine, 2-methylguanosine, N6-methyladenosine, 7-methylguanosine, 5-methoxyaminomethyl-2-thiouridine, 5-methylaminomethyluridine, 5-methylcarbonyhnethyluridine, 5-methyloxyuridine, 5-methyl-2-thiouridine, 2-methylthio-N6-isopentenyladenosine, R-D-mannosylqueosine, uridine-5-oxyacetic acid, 2-thiocytidine, threonine derivatives and others (Burgin et al., 1996, Biochemistry, 35, 14090; Uhlman & Peyman, supra). By "modified bases" in this aspect is meant nucleotide bases other than adenine (A), guanine (G), cytosine (C), thymine (T), and uracil (U), as illustrated above; such bases can be used at any position in the antisense molecule. Persons skilled in the art will appreciate that depending on the uses of the oligomers, Ts and Us are interchangeable. For instance, with other antisense chemistries such as 2'-O-methyl antisense oligonucleotides that are more RNA-like, the T bases may be shown as U (see, e.g., Sequence ID Listing).
[0093] An "amino acid subunit" or "amino acid residue" can refer to an .alpha.-amino acid residue (e.g., --CO--CHR--NH--) or a .beta.- or other amino acid residue (e.g., --CO--(CH.sub.2).sub.nCHR--NH--), where R is a side chain (which may include hydrogen) and n is 1 to 6, preferably 1 to 4.
[0094] The term "naturally occurring amino acid" refers to an amino acid present in proteins found in nature, such as the 20 (L)-amino acids utilized during protein biosynthesis as well as others such as 4-hydroxyproline, hydroxylysine, desmosine, isodesmosine, homocysteine, citrulline and omithine. The term "non-natural amino acids" refers to those amino acids not present in proteins found in nature, examples include beta-alanine (.beta.-Ala; or B), 6-aminohexanoic acid (Ahx) and 6-aminopentanoic acid. Additional examples of "non-natural amino acids" include, without limitation, (D)-amino acids, norleucine, norvaline, p-fluorophenylalanine, ethionine and the like, which are known to a person skilled in the art.
[0095] An "effective amount" or "therapeutically effective amount" refers to an amount of therapeutic compound, such as an antisense oligomer, administered to a mammalian subject, either as a single dose or as part of a series of doses, which is effective to produce a desired physiological response or therapeutic effect in the subject. One example of a desired physiological response includes increased expression of a relatively functional or biologically active form of the dystrophin protein, mainly in muscle tissues or cells that contain a defective dystrophin protein or no dystrophin, as compared no antisense oligomer or a control oligomer. Examples of desired therapeutic effects include, without limitation, improvements in the symptoms or pathology of muscular dystrophy, reducing the progression of symptoms or pathology of muscular dystrophy, and slowing the onset of symptoms or pathology of muscular dystrophy, among others. Examples of such symptoms include fatigue, mental retardation, muscle weakness, difficulty with motor skills (e.g., running, hopping, jumping), frequent falls, and difficulty walking. The pathology of muscular dystrophy can be characterized, for example, by muscle fibre damage and membrane leakage. For an antisense oligomer, this effect is typically brought about by altering the splice-processing of a selected target sequence (e.g., dystrophin), such as to induce exon skipping.
[0096] An "exon" refers to a defined section of nucleic acid that encodes for a protein, or a nucleic acid sequence that is represented in the mature form of an RNA molecule after either portions of a pre-processed (or precursor) RNA have been removed by splicing. The mature RNA molecule can be a messenger RNA (mRNA) or a functional form of a non-coding RNA, such as rRNA or tRNA. The human dystrophin gene has about 75 exons.
[0097] An "intron" refers to a nucleic acid region (within a gene) that is not translated into a protein. An intron is a non-coding section that is transcribed into a precursor mRNA (pre-mRNA), and subsequently removed by splicing during formation of the mature RNA.
[0098] "Exon skipping" refers generally to the process by which an entire exon, or a portion thereof, is removed from a given pre-processed RNA, and is thereby excluded from being present in the mature RNA, such as the mature mRNA that is translated into a protein. Hence, the portion of the protein that is otherwise encoded by the skipped exon is not present in the expressed form of the protein, typically creating an altered, though still functional, form of the protein. In certain embodiments, the exon being skipped is an aberrant exon from the human dystrophin gene, which may contain a mutation or other alteration in its sequence that otherwise causes aberrant splicing. In certain embodiments, the exon being skipped is any one or more of exons 1-75 of the dystrophin gene, though any one or more of exons 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, and/or 55 of the human dystrophin gene are preferred.
[0099] "Dystrophin" is a rod-shaped cytoplasmic protein, and a vital part of the protein complex that connects the cytoskeleton of a muscle fiber to the surrounding extracellular matrix through the cell membrane. Dystrophin contains multiple functional domains. For instance, dystrophin contains an actin binding domain at about amino acids 14-240 and a central rod domain at about amino acids 253-3040. This large central domain is formed by 24 spectrin-like triple-helical elements of about 109 amino acids, which have homology to alpha-actinin and spectrin. The repeats are typically interrupted by four proline-rich non-repeat segments, also referred to as hinge regions. Repeats 15 and 16 are separated by an 18 amino acid stretch that appears to provide a major site for proteolytic cleavage of dystrophin. The sequence identity between most repeats ranges from 10-25%. One repeat contains three alpha-helices: 1, 2 and 3. Alpha-helices 1 and 3 are each formed by 7 helix turns, probably interacting as a coiled-coil through a hydrophobic interface. Alpha-helix 2 has a more complex structure and is formed by segments of four and three helix turns, separated by a Glycine or Proline residue. Each repeat is encoded by two exons, typically interrupted by an intron between amino acids 47 and 48 in the first part of alpha-helix 2. The other intron is found at different positions in the repeat, usually scattered over helix-3. Dystrophin also contains a cysteine-rich domain at about amino acids 3080-3360), including a cysteine-rich segment (i.e., 15 Cysteines in 280 amino acids) showing homology to the C-terminal domain of the slime mold (Dictyostelium discoideum) alpha-actinin. The carboxy-terminal domain is at about amino acids 3361-3685.
[0100] The amino-terminus of dystrophin binds to F-actin and the carboxy-terminus binds to the dystrophin-associated protein complex (DAPC) at the sarcolemma. The DAPC includes the dystroglycans, sarcoglycans, integrins and caveolin, and mutations in any of these components cause autosomally inherited muscular dystrophies. The DAPC is destabilized when dystrophin is absent, which results in diminished levels of the member proteins, and in turn leads to progressive fibre damage and membrane leakage. In various forms of muscular dystrophy, such as Duchenne's muscular dystrophy (DMD) and Becker's muscular dystrophy (BMD), muscle cells produce an altered and functionally defective form of dystrophin, or no dystrophin at all, mainly due to mutations in the gene sequence that lead to incorrect splicing. The predominant expression of the defective dystrophin protein, or the complete lack of dystrophin or a dystrophin-like protein, leads to rapid progression of muscle degeneration, as noted above. In this regard, a "defective" dystrophin protein may be characterized by the forms of dystrophin that are produced in certain subjects with DMD or BMD, as known in the art, or by the absence of detectable dystrophin.
[0101] Table A provides an illustration of the various dystrophin domains, the amino acid residues that encompass these domains, and the exons that encode them.
TABLE-US-00001 TABLE A Residue Domain Sub Domain Nos Exons actin binding 14-240 2-8 domain central rod 253-3040 8-61 domain hinge 1 253-327 (8)-9 repeat 1 337-447 10-11 repeat 2 448-556 12-14 repeat 3 557-667 14-16 hinge 2 668-717 17 repeat 4 718-828 (17)-20 repeat 5 829-934 20-21 repeat 6 935-1045 22-23 repeat 7 1046-1154 (23)-(26) repeat 8 1155-1263 26-27 repeat 9 1264-1367 28-(30) repeat 10 1368-1463 30-32 repeat 11 1464-1568 32-(34) repeat 12 1569-1676 34-35 repeat 13 1677-1778 36-37 repeat 14 1779-1874 38-(40) repeat 15 1875-1973 40-41 interruption 1974-1991 42 repeat 16 1992-2101 42-43 repeat 17 2102-2208 44-45 repeat 18 2209-2318 46-48 repeat 19 2319-2423 48-50 hinge 3 2424-2470 50-51 repeat 20 2471-2577 51-53 repeat 21 2578-2686 53-(55) repeat 22 2687-2802 55-(57) repeat 23 2803-2931 57-59 repeat 24 2932-3040 59-(61) hinge 4 3041-3112 61-64 Cysteine-rich 3080-3360 63-69 domain dystroglycan binding site 3080-3408 63-70 WW domain 3056-3092 62-63 EF-hand 1 3130-3157 65 EF-hand 2 3178-3206 65-66 ZZ domain 3307-3354 68-69 Carboxy-terminal 3361-3685 70-79 domain alpha1-syntrophin binding 3444-3494 73-74 site .beta.1-syntrophin binding site 3495-3535 74-75 (Leu)6-heptad repeat 3558-3593 75
[0102] As used herein, the terms "function" and "functional" and the like refer to a biological, enzymatic, or therapeutic function.
[0103] A "functional" dystrophin protein refers generally to a dystrophin protein having sufficient biological activity to reduce the progressive degradation of muscle tissue that is otherwise characteristic of muscular dystrophy, typically as compared to the altered or "defective" form of dystrophin protein that is present in certain subjects with DMD or BMD. In certain embodiments, a functional dystrophin protein may have about 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% (including all integers in between) of the in vitro or in vivo biological activity of wild-type dystrophin, as measured according to routine techniques in the art. As one example, dystrophin-related activity in muscle cultures in vitro can be measured according to myotube size, myofibril organization (or disorganization), contractile activity, and spontaneous clustering of acetylcholine receptors (see, e.g., Brown et al., Journal of Cell Science. 112:209-216, 1999). Animal models are also valuable resources for studying the pathogenesis of disease, and provide a means to test dystrophin-related activity. Two of the most widely used animal models for DMD research are the mdx mouse and the golden retriever muscular dystrophy (GRMD) dog, both of which are dystrophin negative (see, e.g., Collins & Morgan, Int J Exp Pathol 84: 165-172, 2003). These and other animal models can be used to measure the functional activity of various dystrophin proteins. Included are truncated forms of dystrophin, such as those forms that are produced by certain of the exon-skipping antisense compounds of the present invention.
[0104] By "gene" is meant a unit of inheritance that occupies a specific locus on a chromosome and consists of transcriptional and/or translational regulatory sequences and/or a coding region and/or non-translated sequences (i.e., introns, 5' and 3' untranslated sequences).
[0105] By "isolated" is meant material that is substantially or essentially free from components that normally accompany it in its native state. For example, an "isolated polynucleotide," as used herein, may refer to a polynucleotide that has been purified or removed from the sequences that flank it in a naturally-occurring state, e.g., a DNA fragment that has been removed from the sequences that are normally adjacent to the fragment.
[0106] By "enhance" or "enhancing," or "increase" or "increasing," or "stimulate" or "stimulating," refers generally to the ability of one or antisense compounds or compositions to produce or cause a greater physiological response (i.e., downstream effects) in a cell or a subject, as compared to the response caused by either no antisense compound or a control compound. A measurable physiological response may include increased expression of a functional form of a dystrophin protein, or increased dystrophin-related biological activity in muscle tissue, among other responses apparent from the understanding in the art and the description herein. Increased muscle function can also be measured, including increases or improvements in muscle function by about 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%. The percentage of muscle fibres that express a functional dystrophin can also be measured, including increased dystrophin expression in about 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% of muscle fibres. For instance, it has been shown that around 40% of muscle function improvement can occur if 25-30% of fibers express dystrophin (see, e.g., DelloRusso et al, Proc Natl Acad Sci USA 99: 12979-12984, 2002). An "increased" or "enhanced" amount is typically a "statistically significant" amount, and may include an increase that is 1.1, 1.2, 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, 30, 40, 50 or more times (e.g., 500, 1000 times) (including all integers and decimal points in between and above 1), e.g., 1.5, 1.6, 1.7. 1.8, etc.) the amount produced by no antisense compound (the absence of an agent) or a control compound.
[0107] The term "reduce" or "inhibit" may relate generally to the ability of one or more antisense compounds of the invention to "decrease" a relevant physiological or cellular response, such as a symptom of a disease or condition described herein, as measured according to routine techniques in the diagnostic art. Relevant physiological or cellular responses (in vivo or in vitro) will be apparent to persons skilled in the art, and may include reductions in the symptoms or pathology of muscular dystrophy, or reductions in the expression of defective forms of dystrophin, such as the altered forms of dystrophin that are expressed in individuals with DMD or BMD. A "decrease" in a response may be statistically significant as compared to the response produced by no antisense compound or a control composition, and may include a 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% decrease, including all integers in between.
[0108] "Homology" refers to the percentage number of amino acids that are identical or constitute conservative substitutions. Homology may be determined using sequence comparison programs such as GAP (Deveraux et al., 1984, Nucleic Acids Research 12, 387-395). In this way sequences of a similar or substantially different length to those cited herein could be compared by insertion of gaps into the alignment, such gaps being determined, for example, by the comparison algorithm used by GAP.
[0109] The recitations "sequence identity" or, for example, comprising a "sequence 50% identical to," as used herein, refer to the extent that sequences are identical on a nucleotide-by-nucleotide basis or an amino acid-by-amino acid basis over a window of comparison. Thus, a "percentage of sequence identity" may be calculated by comparing two optimally aligned sequences over the window of comparison, determining the number of positions at which the identical nucleic acid base (e.g., A, T, C, G, I) or the identical amino acid residue (e.g., Ala, Pro, Ser, Thr, Gly, Val, Leu, lie, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gin, Cys and Met) occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of sequence identity.
[0110] Terms used to describe sequence relationships between two or more polynucleotides or polypeptides include "reference sequence," "comparison window," "sequence identity," "percentage of sequence identity," and "substantial identity". A "reference sequence" is at least 8 or 10 but frequently 15 to 18 and often at least 25 monomer units, inclusive of nucleotides and amino acid residues, in length. Because two polynucleotides may each comprise (1) a sequence (i.e., only a portion of the complete polynucleotide sequence) that is similar between the two polynucleotides, and (2) a sequence that is divergent between the two polynucleotides, sequence comparisons between two (or more) polynucleotides are typically performed by comparing sequences of the two polynucleotides over a "comparison window" to identify and compare local regions of sequence similarity. A "comparison window" refers to a conceptual segment of at least 6 contiguous positions, usually about 50 to about 100, more usually about 100 to about 150 in which a sequence is compared to a reference sequence of the same number of contiguous positions after the two sequences are optimally aligned. The comparison window may comprise additions or deletions (i.e., gaps) of about 20% or less as compared to the reference sequence (which does not comprise additions or deletions) for optimal alignment of the two sequences. Optimal alignment of sequences for aligning a comparison window may be conducted by computerized implementations of algorithms (GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package Release 7.0, Genetics Computer Group, 575 Science Drive Madison, Wis., USA) or by inspection and the best alignment (i.e., resulting in the highest percentage homology over the comparison window) generated by any of the various methods selected. Reference also may be made to the BLAST family of programs as for example disclosed by Altschul et al., 1997, Nucl. Acids Res. 25:3389. A detailed discussion of sequence analysis can be found in Unit 19.3 of Ausubel et al., "Current Protocols in Molecular Biology," John Wiley & Sons Inc, 1994-1998, Chapter 15.
[0111] "Treatment" or "treating" of an individual (e.g., a mammal, such as a human) or a cell may include any type of intervention used in an attempt to alter the natural course of the individual or cell. Treatment includes, but is not limited to, administration of a pharmaceutical composition, and may be performed either prophylactically or subsequent to the initiation of a pathologic event or contact with an etiologic agent. Treatment includes any desirable effect on the symptoms or pathology of a disease or condition associated with the dystrophin protein, as in certain forms of muscular dystrophy, and may include, for example, minimal changes or improvements in one or more measurable markers of the disease or condition being treated. Also included are "prophylactic" treatments, which can be directed to reducing the rate of progression of the disease or condition being treated, delaying the onset of that disease or condition, or reducing the severity of its onset. "Treatment" or "prophylaxis" does not necessarily indicate complete eradication, cure, or prevention of the disease or condition, or associated symptoms thereof.
[0112] Hence, included are methods of treating muscular dystrophy, such as DMD and BMD, by administering one or more antisense oligomers of the present invention (e.g., SEQ ID NOS: 1 to 569 and 612 to 635, and variants thereof), optionally as part of a pharmaceutical formulation or dosage form, to a subject in need thereof. Also included are methods of inducing exon-skipping in a subject by administering one or more antisense oligomers, in which the exon is one of exons 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, and/or 55 from the dystrophin gene, preferably the human dystrophin gene. A "subject," as used herein, includes any animal that exhibits a symptom, or is at risk for exhibiting a symptom, which can be treated with an antisense compound of the invention, such as a subject that has or is at risk for having DMD or BMD, or any of the symptoms associated with these conditions (e.g., muscle fibre loss). Suitable subjects (patients) include laboratory animals (such as mouse, rat, rabbit, or guinea pig), farm animals, and domestic animals or pets (such as a cat or dog). Non-human primates and, preferably, human patients, are included.
[0113] Also included are vector delivery systems that are capable of expressing the oligomeric, dystrophin-targeting sequences of the present invention, such as vectors that express a polynucleotide sequence comprising any one or more of SEQ ID NOS: 1 to 569 and 612 to 635, or variants thereof, as described herein. By "vector" or "nucleic acid construct" is meant a polynucleotide molecule, preferably a DNA molecule derived, for example, from a plasmid, bacteriophage, yeast or virus, into which a polynucleotide can be inserted or cloned. A vector preferably contains one or more unique restriction sites and can be capable of autonomous replication in a defined host cell including a target cell or tissue or a progenitor cell or tissue thereof, or be integrable with the genome of the defined host such that the cloned sequence is reproducible. Accordingly, the vector can be an autonomously replicating vector, i.e., a vector that exists as an extra-chromosomal entity, the replication of which is independent of chromosomal replication, e.g., a linear or closed circular plasmid, an extra-chromosomal element, a mini-chromosome, or an artificial chromosome. The vector can contain any means for assuring self-replication. Alternatively, the vector can be one which, when introduced into the host cell, is integrated into the genome and replicated together with the chromosome(s) into which it has been integrated.
[0114] A vector or nucleic acid construct system can comprise a single vector or plasmid, two or more vectors or plasmids, which together contain the total DNA to be introduced into the genome of the host cell, or a transposon. The choice of the vector will typically depend on the compatibility of the vector with the host cell into which the vector is to be introduced. In the present case, the vector or nucleic acid construct is preferably one which is operably functional in a mammalian cell, such as a muscle cell. The vector can also include a selection marker such as an antibiotic or drug resistance gene, or a reporter gene (i.e., green fluorescent protein, luciferase), that can be used for selection or identification of suitable transformants or transfectants. Exemplary delivery systems may include viral vector systems (i.e., viral-mediated transduction) including, but not limited to, retroviral (e.g., lentiviral) vectors, adenoviral vectors, adeno-associated viral vectors, and herpes viral vectors, among others known in the art.
[0115] The term "operably linked" as used herein means placing an oligomer-encoding sequence under the regulatory control of a promoter, which then controls the transcription of the oligomer.
[0116] A wild-type gene or gene product is that which is most frequently observed in a population and is thus arbitrarily designed the "normal" or "wild-type" form of the gene.
[0117] "Alkyl" or "alkylene" both refer to a saturated straight or branched chain hydrocarbon radical containing from 1 to 18 carbons. Examples include without limitation methyl, ethyl, propyl, iso-propyl, butyl, iso-butyl, tert-butyl, n-pentyl and n-hexyl. The term "lower alkyl" refers to an alkyl group, as defined herein, containing between 1 and 8 carbons.
[0118] "Alkenyl" refers to an unsaturated straight or branched chain hydrocarbon radical containing from 2 to 18 carbons and comprising at least one carbon to carbon double bond. Examples include without limitation ethenyl, propenyl, iso-propenyl, butenyl, iso-butenyl, tert-butenyl, n-pentenyl and n-hexenyl. The term "lower alkenyl" refers to an alkenyl group, as defined herein, containing between 2 and 8 carbons.
[0119] "Alkynyl" refers to an unsaturated straight or branched chain hydrocarbon radical containing from 2 to 18 carbons comprising at least one carbon to carbon triple bond. Examples include without limitation ethynyl, propynyl, iso-propynyl, butynyl, iso-butynyl, tert-butynyl, pentynyl and hexynyl. The term "lower alkynyl" refers to an alkynyl group, as defined herein, containing between 2 and 8 carbons.
[0120] "Cycloalkyl" refers to a mono- or poly-cyclic alkyl radical. Examples include without limitation cyclobutyl, cycopentyl, cyclohexyl, cycloheptyl and cyclooctyl.
[0121] "Aryl" refers to a cyclic aromatic hydrocarbon moiety containing from 5 to 18 carbons having one or more closed ring(s). Examples include without limitation phenyl, benzyl, naphthyl, anthracenyl, phenanthracenyl and biphenyl.
[0122] "Aralkyl" refers to a radical of the formula RaRb where Ra is an alkylene chain as defined above and Rb is one or more aryl radicals as defined above, for example, benzyl, diphenylmethyl and the like.
[0123] "Thioalkoxy" refers to a radical of the formula --SRc where Rc is an alkyl radical as defined herein. The term "lower thioalkoxy" refers to an alkoxy group, as defined herein, containing between 1 and 8 carbons.
[0124] "Alkoxy" refers to a radical of the formula --ORda where Rd is an alkyl radical as defined herein. The term "lower alkoxy" refers to an alkoxy group, as defined herein, containing between 1 and 8 carbons. Examples of alkoxy groups include, without limitation, methoxy and ethoxy.
[0125] "Alkoxyalkyl" refers to an alkyl group substituted with an alkoxy group.
[0126] "Carbonyl" refers to the --C(.dbd.O)-- radical.
[0127] "Guanidynyl" refers to the H.sub.2N(C.dbd.NH.sub.2)--NH-- radical.
[0128] "Amidinyl" refers to the H.sub.2N(C.dbd.NH.sub.2)CH-- radical.
[0129] "Amino" refers to the --NH.sub.2 radical.
[0130] "Alkylamino" refers to a radical of the formula --NHRd or --NRdRd where each Rd is, independently, an alkyl radical as defined herein. The term "lower alkylamino" refers to an alkylamino group, as defined herein, containing between 1 and 8 carbons.
[0131] "Heterocycle" means a 5- to 7-membered monocyclic, or 7- to 10-membered bicyclic, heterocyclic ring which is either saturated, unsaturated, or aromatic, and which contains from 1 to 4 heteroatoms independently selected from nitrogen, oxygen and sulfur, and wherein the nitrogen and sulfur heteroatoms may be optionally oxidized, and the nitrogen heteroatom may be optionally quaternized, including bicyclic rings in which any of the above heterocycles are fused to a benzene ring. The heterocycle may be attached via any heteroatom or carbon atom. Heterocycles include heteroaryls as defined below. Thus, in addition to the heteroaryls listed below, heterocycles also include morpholinyl, pyrrolidinonyl, pyrrolidinyl, piperidinyl, piperizynyl, hydantoinyl, valerolactamyl, oxiranyl, oxetanyl, tetrahydrofuranyl, tetrahydropyranyl, tetrahydropyridinyl, tetrahydrothiophenyl, tetrahydrothiopyranyl, tetrahydropyrimidinyl, tetrahydrothiopyranyl, and the like.
[0132] "Heteroaryl" means an aromatic heterocycle ring of 5- to 10 members and having at least one heteroatom selected from nitrogen, oxygen and sulfur, and containing at least 1 carbon atom, including both mono- and bicyclic ring systems. Representative heteroaryls are pyridyl, furyl, benzofuranyl, thiophenyl, benzothiophenyl, quinolinyl, pyrrolyl, indolyl, oxazolyl, benzoxazolyl, imidazolyl, benzimidazolyl, thiazolyl, benzothiazolyl, isoxazolyl, pyrazolyl, isothiazolyl, pyridazinyl, pyrimidinyl, pyrazinyl, triazinyl, cinnolinyl, phthalazinyl, and quinazolinyl.
[0133] The terms "optionally substituted alkyl", "optionally substituted alkenyl", "optionally substituted alkoxy", "optionally substituted thioalkoxy", "optionally substituted alkyl amino", "optionally substituted lower alkyl", "optionally substituted lower alkenyl", "optionally substituted lower alkoxy", "optionally substituted lower thioalkoxy", "optionally substituted lower alkyl amino" and "optionally substituted heterocyclyl" mean that, when substituted, at least one hydrogen atom is replaced with a substituent. In the case of an oxo substituent (.dbd.O) two hydrogen atoms are replaced. In this regard, substituents include: deuterium, optionally substituted alkyl, optionally substituted alkenyl, optionally substituted alkynyl, optionally substituted aryl, optionally substituted heterocycle, optionally substituted cycloalkyl, oxo, halogen, --CN, --ORx, NRxRy, NRxC(.dbd.O)Ry, NRxSO2Ry, --NRxC(.dbd.O)NRxRy, C(.dbd.O)Rx, C(.dbd.O)ORx, C(.dbd.O)NRxRy, --SOmRx and --SOmNRxRy, wherein m is 0, 1 or 2, Rx and Ry are the same or different and independently hydrogen, optionally substituted alkyl, optionally substituted alkenyl, optionally substituted alkynyl, optionally substituted aryl, optionally substituted heterocycle or optionally substituted cycloalkyl and each of said optionally substituted alkyl, optionally substituted alkenyl, optionally substituted alkynyl, optionally substituted aryl, optionally substituted heterocycle and optionally substituted cycloalkyl substituents may be further substituted with one or more of oxo, halogen, --CN, --ORx, NRxRy, NRxC(.dbd.O)Ry, NRxSO2Ry, --NRxC(.dbd.O)NRxRy, C(.dbd.O)Rx, C(.dbd.O)ORx, C(.dbd.O)NRxRy, --SOmRx and --SOmNRxRy.
Constructing Antisense Oligonucleotides
[0134] Examples of morpholino oligonucleotides having phosphorus-containing backbone linkages are illustrated in FIGS. 1A-1C. Especially preferred is a phosphorodiamidate-linked morpholino oligonucleotide such as shown in FIG. 1C, which is modified, in accordance with one aspect of the present invention, to contain positively charged groups at preferably 10%-50% of its backbone linkages. Morpholino oligonucleotides with uncharged backbone linkages and their preparation, including antisense oligonucleotides, are detailed, for example, in (Summerton and Weller 1997) and in co-owned U.S. Pat. Nos. 5,698,685, 5,217,866, 5,142,047, 5,034,506, 5,166,315, 5,185, 444, 5,521,063, and 5,506,337, all of which are expressly incorporated by reference herein.
[0135] Important properties of the morpholino-based subunits include: 1) the ability to be linked in a oligomeric form by stable, uncharged or positively charged backbone linkages; 2) the ability to support a nucleotide base (e.g. adenine, cytosine, guanine, thymidine, uracil and inosine) such that the polymer formed can hybridize with a complementary-base target nucleic acid, including target RNA, Tm values above about 45.degree. C. in relatively short oligonucleotides (e.g., 10-15 bases); 3) the ability of the oligonudeotide to be actively or passively transported into mammalian cells; and 4) the ability of the antisense oligonucleotide:RNA heteroduplex to resist RNAse and RNaseH degradation, respectively.
[0136] Exemplary backbone structures for antisense oligonucleotides of the claimed subject matter include the morpholino subunit types shown in FIGS. 1D-G, each linked by an uncharged or positively charged, phosphorus-containing subunit linkage. FIG. 1D shows a phosphorus-containing linkage which forms the five atom repeating-unit backbone, wherein the morpholino rings are linked by a 1-atom phosphoamide linkage. FIG. 1E shows a linkage which produces a 6-atom repeating-unit backbone. In this structure, the atom Y linking the 5' morpholino carbon to the phosphorus group may be sulfur, nitrogen, carbon or, preferably, oxygen. The X moiety pendant from the phosphorus may be fluorine, an alkyl or substituted alkyl, an alkoxy or substituted alkoxy, a thioalkoxy or substituted thioalkoxy, or unsubstituted, monosubstituted, or disubstituted nitrogen, including cyclic structures, such as morpholines or piperidines. Alkyl, alkoxy and thioalkoxy preferably include 1-6 carbon atoms. The Z moieties are sulfur or oxygen, and are preferably oxygen.
[0137] The linkages shown in FIGS. 1F and 1G are designed for 7-atom unit-length backbones. In structure 1F, the X moiety is as in Structure 1E, and the Y moiety may be methylene, sulfur, or, preferably, oxygen. In Structure 1G, the X and Y moieties are as in Structure 1E. Particularly preferred morpholino oligonucleotides include those composed of morpholino subunit structures of the form shown in FIG. 1E, where X=NH.sub.2, N(CH.sub.3).sub.2, optionally substituted 1-piperazinyl, or other charged group, Y.dbd.O, and Z.dbd.O.
[0138] As noted above, the uncharged or substantially uncharged oligonucleotide may be modified, in accordance with an aspect of the invention, to include charged linkages, e.g. up to about 1 per every 2-5 uncharged linkages, such as about 4-5 per every 10 uncharged linkages. Optimal improvement in antisense activity may be seen when about 25% of the backbone linkages are cationic, including about 20% to about 30%. Also included are oligomers in which about 35%, 40%, 45%, 50%, 55%, 60% (including all integers in between), or more of the backbone linkages are cationic. Enhancement is also seen with a small number, e.g., 5% or 10-20%, of cationic linkages.
[0139] A substantially uncharged, phosphorus containing backbone in an oligonucleotide analog is typically one in which a majority of the subunit linkages, e.g., between 50%-100%, typically at least 60% to 100% or 75% or 80% of its linkages, are uncharged at physiological pH and contain a single phosphorous atom.
[0140] Additional experiments conducted in support of the present invention indicate that the enhancement seen with added cationic backbone charges may, in some cases, be further enhanced by distributing the bulk of the charges close to the "center-region" backbone linkages of the antisense oligonucleotide, e.g., in a 20mer oligonucleotide with 8 cationic backbone linkages, having at least 70% of these charged linkages localized in the 10 centermost linkages.
[0141] The antisense compounds can be prepared by stepwise solid-phase synthesis, employing methods detailed in the references cited above, and below with respect to the synthesis of oligonucleotides having a mixture of uncharged and cationic backbone linkages. In some cases, it may be desirable to add additional chemical moieties to the antisense compound, e.g. to enhance pharmacokinetics or to facilitate capture or detection of the compound. Such a moiety may be covalently attached, typically to a terminus of the oligomer, according to standard synthetic methods. For example, addition of a polyethyleneglycol moiety or other hydrophilic polymer, e.g., one having 10-100 monomeric subunits, may be useful in enhancing solubility. One or more charged groups, e.g., anionic charged groups such as an organic acid, may enhance cell uptake. A reporter moiety, such as fluorescein or a radiolabeled group, may be attached for purposes of detection. Alternatively, the reporter label attached to the oligomer may be a ligand, such as an antigen or biotin, capable of binding a labeled antibody or streptavidin. In selecting a moiety for attachment or modification of an antisense compound, it is generally of course desirable to select chemical compounds of groups that are biocompatible and likely to be tolerated by a subject without undesirable side effects.
[0142] As noted above, the antisense compound can be constructed to contain a selected number of cationic linkages interspersed with uncharged linkages of the type described above. The intersubunit linkages, both uncharged and cationic, preferably are phosphorus-containing linkages, having the structure (II):
##STR00002##
wherein:
[0143] W is --S-- or --O--, and is preferably --O--,
[0144] X=--NR.sup.1R.sup.2 or --OR.sup.6,
[0145] Y=--O-- or --NR.sup.7, and
[0146] each said linkage in the oligomer is selected from:
[0147] (a) an uncharged linkage (a), wherein each of R.sup.1, R.sup.2, R.sup.6 and R.sup.7 is independently selected from hydrogen and lower alkyl;
[0148] (b1) a cationic linkage (b1), wherein X=--NR.sup.1R.sup.2 and Y=--O--, and --NR.sup.1R.sup.2 represents an optionally substituted piperazinyl moiety, such that R.sup.1R.sup.2=--CHRCHRN(R.sup.3)(R.sup.4)CHRCHR--, wherein:
[0149] each R is independently H or --CH.sub.3,
[0150] R.sup.4 is H, --CH.sub.3, or an electron pair, and
[0151] R.sup.3 is selected from H, optionally substituted lower alkyl, --C(.dbd.NH)NH.sub.2, --Z-L-NHC(.dbd.NH)NH.sub.2, and [--C(.dbd.O)CHR'NH].sub.mH, where: Z is --C(.dbd.O)-- or a direct bond, L is an optional linker up to 18 atoms in length, preferably up to 12 atoms, and more preferably up to 8 atoms in length, having bonds selected from optionally substituted alkyl, optionally substituted alkoxy, and optionally substituted alkylamino, R' is a side chain of a naturally occurring amino acid or a one- or two-carbon homolog thereof, and m is 1 to 6, preferably 1 to 4;
[0152] (b2) a cationic linkage (b2), wherein X=--NR.sup.1R.sup.2 and Y=--O--, R.sup.1=H or --CH.sub.3, and R.sup.2=LNR.sup.3R.sup.4R.sup.5, wherein L, R.sup.3, and R.sup.4 are as defined above, and R.sup.5 is H, optionally substituted lower alkyl, or optionally substituted lower (alkoxy)alkyl; and
[0153] (b3) a cationic linkage (b3), wherein Y=--NR.sup.7 and X=--OR.sup.6, and R.sup.7=-LNR.sup.3R.sup.4R.sup.5, wherein L, R.sup.3, R.sup.4 and R.sup.5 are as defined above, and R.sup.6 is H or optionally substituted lower alkyl; and
[0154] at least one said linkage is selected from cationic linkages (b1), (b2), and (b3).
[0155] Preferably, the oligomer includes at least two consecutive linkages of type (a) (i.e. uncharged linkages). In further embodiments, at least 5% of the linkages in the oligomer are cationic linkages (i.e. type (b1), (b2), or (b3)); for example, 10% to 60%, and preferably 20-50% linkages may be cationic linkages.
[0156] In one embodiment, at least one linkage is of type (b1), where, preferably, each R is H, R.sup.4 is H, --CH.sub.3, or an electron pair, and R.sup.3 is selected from H, optionally substituted lower alkyl, --C(.dbd.NH)NH.sub.2, and --C(.dbd.O)-L-NHC(.dbd.NH)NH.sub.2. The latter two embodiments of R.sup.3 provide a guanidino moiety, either attached directly to the piperazine ring, or pendant to a linker group L, respectively. For ease of synthesis, the variable Z in R.sup.3 is preferably --C(.dbd.O)--, as shown.
[0157] The linker group L, as noted above, contains bonds in its backbone selected from optionally substituted alkyl, optionally substituted alkoxy, and optionally substituted alkylamino, wherein the terminal atoms in L (e.g., those adjacent to carbonyl or nitrogen) are carbon atoms. Although branched linkages are possible, the linker is preferably unbranched. In one embodiment, the linker is a linear alkyl linker. Such a linker may have the structure --(CH.sub.2)--, where n is 1-12, preferably 2-8, and more preferably 2-6.
[0158] The morpholino subunits have the following structure (III):
##STR00003##
wherein Pi is a base-pairing moiety, and the linkages depicted above connect the nitrogen atom of (III) to the 5' carbon of an adjacent subunit. The base-pairing moieties Pi may be the same or different, and are generally designed to provide a sequence which binds to a target nucleic acid.
[0159] The use of embodiments of linkage types (b1), (b2) and (b3) above to link morpholino subunits (III) may be illustrated graphically as follows:
##STR00004##
[0160] Preferably, all cationic linkages in the oligomer are of the same type; i.e. all of type (b1), all of type (b2), or all of type (b3).
[0161] In further embodiments, the cationic linkages are selected from linkages (b1') and (b1'') as shown below, where (b1') is referred to herein as a "Pip" linkage and (b1'') is referred to herein as a "GuX" linkage:
##STR00005##
[0162] In the structures above, W is S or O, and is preferably O; each of R.sup.1 and R.sup.2 is independently selected from hydrogen and optionally substituted lower alkyl, and is preferably methyl; and A represents hydrogen or a non-interfering substituent (i.e. a substituent that does not adversely affect the ability of an oligomer to bind to its intended target) on one or more carbon atoms in (b1') and (b1''). Preferably, the ring carbons in the piperazine ring are unsubstituted; however, the ring carbons of the piperazine ring may include non-interfering substituents, such as methyl or fluorine. Preferably, at most one or two carbon atoms is so substituted.
[0163] In further embodiments, at least 10% of the linkages are of type (b1') or (b1''); for example, 10%-60% and preferably 20% to 50%, of the linkages may be of type (b1') or (b1'').
[0164] In other embodiments, the oligomer contains no linkages of the type (b1') above. Alternatively, the oligomer contains no linkages of type (b1) where each R is H, R.sup.3 is H or --CH.sub.3, and R.sup.4 is H, --CH.sub.3, or an electron pair.
[0165] The morpholino subunits may also be linked by non-phosphorus-based intersubunit linkages, as described further below, where at least one linkage is modified with a pendant cationic group as described above.
[0166] Other oligonucleotide analog linkages which are uncharged in their unmodified state but which could also bear a pendant amine substituent could be used. For example, a 5'nitrogen atom on a morpholino ring could be employed in a sulfamide linkage or a urea linkage (where phosphorus is replaced with carbon or sulfur, respectively) and modified in a manner analogous to the 5'-nitrogen atom in structure (b3) above.
[0167] Oligomers having any number of cationic linkages are provided, including fully cationic-linked oligomers. Preferably, however, the oligomers are partially charged, having, for example, 10%-80%. In preferred embodiments, about 10% to 60%, and preferably 20% to 50% of the linkages are cationic.
[0168] In one embodiment, the cationic linkages are interspersed along the backbone. The partially charged oligomers preferably contain at least two consecutive uncharged linkages; that is, the oligomer preferably does not have a strictly alternating pattern along its entire length.
[0169] Also considered are oligomers having blocks of cationic linkages and blocks of uncharged linkages; for example, a central block of uncharged linkages may be flanked by blocks of cationic linkages, or vice versa. In one embodiment, the oligomer has approximately equal-length 5', 3' and center regions, and the percentage of cationic linkages in the center region is greater than about 50%, preferably greater than about 70%.
[0170] Oligomers for use in antisense applications generally range in length from about 10 to about 40 subunits, more preferably about 10 to 30 subunits, and typically 15-25 bases, including those having 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, or 40 bases. In certain embodiments, an oligomer of the invention having 19-20 subunits, a useful length for an antisense compound, may ideally have two to ten, e.g., four to eight, cationic linkages, and the remainder uncharged linkages. An oligomer having 14-15 subunits may ideally have two to seven, e.g., 3 to 5, cationic linkages, and the remainder uncharged linkages.
[0171] Each morpholino ring structure supports a base pairing moiety, to form a sequence of base pairing moieties which is typically designed to hybridize to a selected antisense target in a cell or in a subject being treated. The base pairing moiety may be a purine or pyrimidine found in native DNA or RNA (e.g., A, G, C, T or U) or an analog, such as hypoxanthine (the base component of the nucleoside inosine) or 5-methyl cytosine.
Peptide Transporters
[0172] The antisense compounds of the invention may include an oligonucleotide moiety conjugated to an arginine-rich peptide transport moiety effective to enhance transport of the compound into cells. The transport moiety is preferably attached to a terminus of the oligomer, as shown, for example, in FIGS. 1B and 1C. The peptide transport moiety preferably comprises 6 to 16 subunits selected from X' subunits, Y subunits, and Z' subunits, wherein:
[0173] (a) each X' subunit independently represents lysine, arginine or an arginine analog, said analog being a cationic .alpha.-amino acid comprising a side chain of the structure R'N=C(NH.sub.2)R.sup.2, where R.sup.1 is H or R; R.sup.2 is R, --NH.sub.2, --NHR, or --NR.sub.2, where R is optionally substituted lower alkyl or optionally substituted lower alkenyl; R.sup.1 and R.sup.2 may join together to form a ring; and the side chain is linked to said amino acid via R.sup.1 or R.sup.2;
[0174] (b) each Y' subunit independently represents a neutral amino acid --C(.dbd.O)--(CHR).sub.n--NH--, where n is 2 to 7 and each R is independently H or methyl; and
[0175] (c) each Z' subunit independently represents an .alpha.-amino acid having a neutral aralkyl side chain;
wherein the peptide comprises a sequence represented by at least one of (X'Y'X').sub.p, (X'Y').sub.m, and/or (X'Z'Z'), where p is 2 to 5 and m is 2 to 8. Certain embodiments include various combinations selected independently from (X'Y'X'), (X'Y').sub.m. and/or (X'Z'Z').sub.p, including, for example, peptides having the sequence (X'Y'X')(X'Z'Z')(X'Y'X'X'Z'Z') (SEQ ID NO:637).
[0176] In selected embodiments, for each X', the side chain moiety is guanidyl, as in the amino acid subunit arginine (Arg). In certain embodiments, each Y' is independently --C(.dbd.O)--(CH.sub.2).sub.n--CHR--NH--, where n is 2 to 7 and R is H. For example, when n is 5 and R is H, Y' is a 6-aminohexanoic acid subunit, abbreviated herein as Ahx; when n is 2 and R is H, Y' is a .beta.-alanine subunit, abbreviated herein as B. Certain embodiments relate to carrier peptides having a combination of different neutral amino acids, including, for example, peptides comprising the sequence -RahxRRBRRAhxRRBRAhxB- (SEQ ID NO:578), which contains both .beta.-alanine and 6-aminohexanoic acid.
[0177] Preferred peptides of this type include those comprising arginine dimers alternating with single Y' subunits, where Y' is preferably Ahx or B or both. Examples include peptides having the formula (RY'R).sub.p and/or the formula (RRY').sub.p, where p is 1 to 2 to 5 and where Y' is preferably Ahx. In one embodiment, Y' is a 6-aminohexanoic acid subunit, R is arginine and p is 4. Certain embodiments include various linear combinations of at least two of (RY'R).sub.p and (RRY').sub.p, including, for example, illustrative peptides having the sequence (RY'R)(RRY')(RY'R)(RRY') (SEQ ID NO:638), or (RRY')(RY'R)(RRY') (SEQ ID NO:639). Other combinations are contemplated. In a further illustrative embodiment, each Z' is phenylalanine, and m is 3 or 4.
[0178] The conjugated peptide is preferably linked to a terminus of the oligomer via a linker Ahx-B, where Ahx is a 6-aminohexanoic acid subunit and B is a .beta.-alanine subunit, as shown, for example, in FIGS. 1B and 1C.
[0179] In selected embodiments, for each X', the side chain moiety is independently selected from the group consisting of guanidyl (HN.dbd.C(NH.sub.2)NH--), amidinyl (HN.dbd.C(NH.sub.2)CH--), 2-aminodihydropyrimidyl, 2-aminotetrahydropyrimidyl, 2-aminopyridinyl, and 2-aminopyrimidonyl, and it is preferably selected from guanidyl and amidinyl. In one embodiment, the side chain moiety is guanidyl, as in the amino acid subunit arginine (Arg).
[0180] In certain embodiments, the Y' subunits may be contiguous, in that no X' subunits intervene between Y' subunits, or interspersed singly between X' subunits. In certain embodiments, the linking subunit may be between Y' subunits. In one embodiment, the Y' subunits are at a terminus of the transporter; in other embodiments, they are flanked by X' subunits. In further preferred embodiments, each Y' is --C(.dbd.O)--(CH.sub.2).sub.nCHR--NH--, where n is 2 to 7 and R is H. For example, when n is 5 and R is H, Y is a 6-aminohexanoic acid subunit, abbreviated herein as Ahx. In selected embodiments of this group, each X' comprises a guanidyl side chain moiety, as in an arginine subunit. Preferred peptides of this type include those comprising arginine dimers alternating with single Y' subunits, where Y is preferably Ahx. Examples include peptides having the formula (RY'R).sub.4 or the formula (RRY').sub.4 where Y' is preferably Ahx. In the latter case, the nucleic acid analog is preferably linked to a terminal Y' subunit, preferably at the C-terminus, as shown, for example, in FIGS. 1B and 1C. The preferred linker is of the structure AhxB, where Ahx is a 6-aminohexanoic acid subunit and B is a .beta.-alanine subunit.
[0181] The transport moieties as described above have been shown to greatly enhance cell entry of attached oligomers, relative to uptake of the oligomer in the absence of the attached transport moiety, and relative to uptake by an attached transport moiety lacking the hydrophobic subunits Y'. Such enhanced uptake is preferably evidenced by at least a two-fold increase, and preferably a four-fold increase, in the uptake of the compound into mammalian cells relative to uptake of the agent by an attached transport moiety lacking the hydrophobic subunits Y'. Uptake is preferably enhanced at least twenty fold, and more preferably forty fold, relative to the unconjugated compound.
[0182] A further benefit of the transport moiety is its expected ability to stabilize a duplex between an antisense compound and its target nucleic acid sequence, presumably by virtue of electrostatic interaction between the positively charged transport moiety and the negatively charged nucleic acid. The number of charged subunits in the transporter is less than 14, as noted above, and preferably between 8 and 11, since too high a number of charged subunits may lead to a reduction in sequence specificity.
[0183] The use of arginine-rich peptide transporters (i.e., cell-penetrating peptides) is particularly useful in practicing the present invention. Certain peptide transporters have been shown to be highly effective at delivery of antisense compounds into primary cells including muscle cells (Marshall, Oda et al. 2007; Jearawiriyapaisam, Moulton et al. 2008; Wu, Moulton et al. 2008). Furthermore, compared to other peptide transporters such as Penetratin and the Tat peptide, the peptide transporters described herein, when conjugated to an antisense PMO, demonstrate an enhanced ability to alter splicing of several gene transcripts (Marshall, Oda et al. 2007). Especially preferred are the P007, CP06062 and CP04057 transport peptides listed below in Table 3 (SEQ ID NOS: 573, 578 and 577, respectively).
[0184] Exemplary peptide transporters, including linkers (B or AhxB) are given below in Table B below. Preferred sequences are those designated CP06062 (SEQ ID NO: 578), P007 (SEQ ID NO: 573) and CP04057 (SEQ ID NO: 577).
TABLE-US-00002 TABLE B Exemplary Peptide Transporters for intracellular Delivery of PMO SEQ Sequence (N-terminal to ID Peptide C-terminal) NO: rTAT RRRQRRKKRC 570 R.sub.9F.sub.2 RRRRRRRRRFFC 571 (RRAhx).sub.4B RRAhxRRAhxRRAhxRRAhxB 572 (RAhxR).sub.4AhxB; RAhxRRAhxRRAhxRRAhxRAhxB 573 (P007) (AhxRR).sub.4AhxB AhxRRAhxRRAhxRRAhxRRAhxB 574 (RAhx).sub.6B RAhxRAhxRAhxRAhxRAhxRAhxB 575 (RAhx).sub.8B RAhxRAhxRAhxRAhxRAhxRAhxRAhx 576 B (RAhxR).sub.5AhxB RAhxRRAhxRRAhxRRAhxRRAhxRAhx 577 (CP05057) B (RAhxRRBR).sub.2AhxB; RAhxRRBRRAhxRRBRAhxB 578 (CP06062) MSP ASSLNIA 579
Formulations
[0185] In certain embodiments, the present invention provides formulations or compositions suitable for the therapeutic delivery of antisense oligomers, as described herein. Hence, in certain embodiments, the present invention provides pharmaceutically acceptable compositions that comprise a therapeutically-effective amount of one or more of the oligomers described herein, formulated together with one or more pharmaceutically acceptable carriers (additives) and/or diluents. While it is possible for an oligomer of the present invention to be administered alone, it is preferable to administer the compound as a pharmaceutical formulation (composition).
[0186] Methods for the delivery of nucleic acid molecules are described, for example, in Akhtar et al., 1992, Trends Cell Bio., 2:139; and Delivery Strategies for Antisense Oligonucleotide Therapeutics, ed. Akhtar; Sullivan et al., PCT WO 94/02595. These and other protocols can be utilized for the delivery of virtually any nucleic acid molecule, including the isolated oligomers of the present invention.
[0187] As detailed below, the pharmaceutical compositions of the present invention may be specially formulated for administration in solid or liquid form, including those adapted for the following: (1) oral administration, for example, drenches (aqueous or non-aqueous solutions or suspensions), tablets, e.g., those targeted for buccal, sublingual, and systemic absorption, boluses, powders, granules, pastes for application to the tongue; (2) parenteral administration, for example, by subcutaneous, intramuscular, intravenous or epidural injection as, for example, a sterile solution or suspension, or sustained-release formulation; (3) topical application, for example, as a cream, ointment, or a controlled-release patch or spray applied to the skin; (4) intravaginally or intrarectally, for example, as a pessary, cream or foam; (5) sublingually; (6) ocularly; (7) transdermally; or (8) nasally.
[0188] The phrase "pharmaceutically acceptable" is employed herein to refer to those compounds, materials, compositions, and/or dosage forms which are, within the scope of sound medical judgment, suitable for use in contact with the tissues of human beings and animals without excessive toxicity, irritation, allergic response, or other problem or complication, commensurate with a reasonable benefit/risk ratio.
[0189] The phrase "pharmaceutically-acceptable carrier" as used herein means a pharmaceutically-acceptable material, composition or vehicle, such as a liquid or solid filler, diluent, excipient, manufacturing aid (e.g., lubricant, talc magnesium, calcium or zinc stearate, or steric acid), or solvent encapsulating material, involved in carrying or transporting the subject compound from one organ, or portion of the body, to another organ, or portion of the body. Each carrier must be "acceptable" in the sense of being compatible with the other ingredients of the formulation and not injurious to the patient.
[0190] Some examples of materials that can serve as pharmaceutically-acceptable carriers include, without limitation: (1) sugars, such as lactose, glucose and sucrose; (2) starches, such as corn starch and potato starch; (3) cellulose, and its derivatives, such as sodium carboxymethyl cellulose, ethyl cellulose and cellulose acetate; (4) powdered tragacanth; (5) malt; (6) gelatin; (7) talc; (8) excipients, such as cocoa butter and suppository waxes; (9) oils, such as peanut oil, cottonseed oil, safflower oil, sesame oil, olive oil, corn oil and soybean oil; (10) glycols, such as propylene glycol; (11) polyols, such as glycerin, sorbitol, mannitol and polyethylene glycol; (12) esters, such as ethyl oleate and ethyl laurate; (13) agar; (14) buffering agents, such as magnesium hydroxide and aluminum hydroxide; (15) alginic acid; (16) pyrogen-free water; (17) isotonic saline; (18) Ringer's solution; (19) ethyl alcohol; (20) pH buffered solutions; (21) polyesters, polycarbonates and/or polyanhydrides; and (22) other non-toxic compatible substances employed in pharmaceutical formulations.
[0191] Additional non-limiting examples of agents suitable for formulation with the antisense oligomers of the instant invention include: PEG conjugated nucleic acids, phospholipid conjugated nucleic acids, nucleic acids containing lipophilic moieties, phosphorothioates, P-glycoprotein inhibitors (such as Pluronic P85) which can enhance entry of drugs into various tissues; biodegradable polymers, such as poly (DL-lactide-coglycolide) microspheres for sustained release delivery after implantation (Emerich, D F et al., 1999, Cell Transplant, 8, 47-58) Alkermes, Inc. Cambridge, Mass.; and loaded nanoparticles, such as those made of polybutylcyanoacrylate, which can deliver drugs across the blood brain barrier and can alter neuronal uptake mechanisms (Prog Neuropsychopharmacol Biol Psychiatry, 23, 941-949, 1999).
[0192] The invention also features the use of the composition comprising surface-modified liposomes containing poly (ethylene glycol) lipids (PEG-modified, branched and unbranched or combinations thereof, or long-circulating liposomes or stealth liposomes). Oligomers of the invention can also comprise covalently attached PEG molecules of various molecular weights. These formulations offer a method for increasing the accumulation of drugs in target tissues. This class of drug carriers resists opsonization and elimination by the mononuclear phagocytic system (MPS or RES), thereby enabling longer blood circulation times and enhanced tissue exposure for the encapsulated drug (Lasic et al. Chem. Rev. 1995, 95, 2601-2627; Ishiwata et al., Chem. Pharm. Bull. 1995, 43, 1005-1011). Such liposomes have been shown to accumulate selectively in tumors, presumably by extravasation and capture in the neovascularized target tissues (Lasic et al., Science 1995, 267, 1275-1276; Oku et al., 1995, Biochim. Biophys. Acta, 1238, 86-90). The long-circulating liposomes enhance the pharmacokinetics and pharmacodynamics of DNA and RNA, particularly compared to conventional cationic liposomes which are known to accumulate in tissues of the MPS (Liu et al., J. Biol. Chem. 1995, 42, 24864-24870; Choi et al., International PCT Publication No. WO 96/10391; Ansell et al., International PCT Publication No. WO 96/10390; Holland et al., International PCT Publication No. WO 96/10392). Long-circulating liposomes are also likely to protect drugs from nuclease degradation to a greater extent compared to cationic liposomes, based on their ability to avoid accumulation in metabolically aggressive MPS tissues such as the liver and spleen.
[0193] In a further embodiment, the present invention includes oligomer compositions prepared for delivery as described in U.S. Pat. Nos. 6,692,911, 7,163,695 and 7,070,807. In this regard, in one embodiment, the present invention provides an oligomer of the present invention in a composition comprising copolymers of lysine and histidine (HK) as described in U.S. Pat. Nos. 7,163,695, 7,070,807, and 6,692,911 either alone or in combination with PEG (e.g., branched or unbranched PEG or a mixture of both), in combination with PEG and a targeting moiety or any of the foregoing in combination with a crosslinking agent. In certain embodiments, the present invention provides antisense oligomers in compositions comprising gluconic-acid-modified polyhistidine or gluconylated-polyhistidine/transferrin-polylysine. One skilled in the art will also recognize that amino acids with properties similar to His and Lys may be substituted within the composition.
[0194] Certain embodiments of the oligomers described herein may contain a basic functional group, such as amino or alkylamino, and are, thus, capable of forming pharmaceutically-acceptable salts with pharmaceutically-acceptable acids. The term "pharmaceutically-acceptable salts" in this respect, refers to the relatively non-toxic, inorganic and organic acid addition salts of compounds of the present invention. These salts can be prepared in situ in the administration vehicle or the dosage form manufacturing process, or by separately reacting a purified compound of the invention in its free base form with a suitable organic or inorganic acid, and isolating the salt thus formed during subsequent purification. Representative salts include the hydrobromide, hydrochloride, sulfate, bisulfate, phosphate, nitrate, acetate, valerate, oleate, palmitate, stearate, laurate, benzoate, lactate, phosphate, tosylate, citrate, maleate, fumarate, succinate, tartrate, napthylate, mesylate, glucoheptonate, lactobionate, and laurylsulphonate salts and the like. (See, e.g., Berge et al. (1977) "Pharmaceutical Salts", J. Pharm. Sci. 66:1-19).
[0195] The pharmaceutically acceptable salts of the subject oligomers include the conventional nontoxic salts or quaternary ammonium salts of the compounds, e.g., from non-toxic organic or inorganic acids. For example, such conventional nontoxic salts include those derived from inorganic acids such as hydrochloride, hydrobromic, sulfuric, sulfamic, phosphoric, nitric, and the like; and the salts prepared from organic acids such as acetic, propionic, succinic, glycolic, stearic, lactic, malic, tartaric, citric, ascorbic, palmitic, maleic, hydroxymaleic, phenylacetic, glutamic, benzoic, salicyclic, sulfanilic, 2-acetoxybenzoic, fumaric, toluenesulfonic, methanesulfonic, ethane disulfonic, oxalic, isothionic, and the like.
[0196] In certain embodiments, the oligomers of the present invention may contain one or more acidic functional groups and, thus, are capable of forming pharmaceutically-acceptable salts with pharmaceutically-acceptable bases. The term "pharmaceutically-acceptable salts" in these instances refers to the relatively non-toxic, inorganic and organic base addition salts of compounds of the present invention. These salts can likewise be prepared in situ in the administration vehicle or the dosage form manufacturing process, or by separately reacting the purified compound in its free acid form with a suitable base, such as the hydroxide, carbonate or bicarbonate of a pharmaceutically-acceptable metal cation, with ammonia, or with a pharmaceutically-acceptable organic primary, secondary or tertiary amine. Representative alkali or alkaline earth salts include the lithium, sodium, potassium, calcium, magnesium, and aluminum salts and the like. Representative organic amines useful for the formation of base addition salts include ethylamine, diethylamine, ethylenediamine, ethanolamine, diethanolamine, piperazine and the like. (See, e.g., Berge et al., supra).
[0197] Wetting agents, emulsifiers and lubricants, such as sodium lauryl sulfate and magnesium stearate, as well as coloring agents, release agents, coating agents, sweetening, flavoring and perfuming agents, preservatives and antioxidants can also be present in the compositions.
[0198] Examples of pharmaceutically-acceptable antioxidants include: (1) water soluble antioxidants, such as ascorbic acid, cysteine hydrochloride, sodium bisulfate, sodium metabisulfite, sodium sulfite and the like; (2) oil-soluble antioxidants, such as ascorbyl palmitate, butylated hydroxyanisole (BHA), butylated hydroxytoluene (BHT), lecithin, propyl gallate, alpha-tocopherol, and the like; and (3) metal chelating agents, such as citric acid, ethylenediamine tetraacetic acid (EDTA), sorbitol, tartaric acid, phosphoric acid, and the like.
[0199] Formulations of the present invention include those suitable for oral, nasal, topical (including buccal and sublingual), rectal, vaginal and/or parenteral administration. The formulations may conveniently be presented in unit dosage form and may be prepared by any methods well known in the art of pharmacy. The amount of active ingredient that can be combined with a carrier material to produce a single dosage form will vary depending upon the host being treated, the particular mode of administration. The amount of active ingredient which can be combined with a carrier material to produce a single dosage form will generally be that amount of the compound which produces a therapeutic effect. Generally, out of one hundred percent, this amount will range from about 0.1 percent to about ninety-nine percent of active ingredient, preferably from about 5 percent to about 70 percent, most preferably from about 10 percent to about 30 percent.
[0200] In certain embodiments, a formulation of the present invention comprises an excipient selected from cyclodextrins, celluloses, liposomes, micelle forming agents, e.g., bile acids, and polymeric carriers, e.g., polyesters and polyanhydrides; and an oligomer of the present invention. In certain embodiments, an aforementioned formulation renders orally bioavailable an oligomer of the present invention.
[0201] Methods of preparing these formulations or compositions include the step of bringing into association an oligomer of the present invention with the carrier and, optionally, one or more accessory ingredients. In general, the formulations are prepared by uniformly and intimately bringing into association a compound of the present invention with liquid carriers, or finely divided solid carriers, or both, and then, if necessary, shaping the product.
[0202] Formulations of the invention suitable for oral administration may be in the form of capsules, cachets, pills, tablets, lozenges (using a flavored basis, usually sucrose and acacia or tragacanth), powders, granules, or as a solution or a suspension in an aqueous or non-aqueous liquid, or as an oil-in-water or water-in-oil liquid emulsion, or as an elixir or syrup, or as pastilles (using an inert base, such as gelatin and glycerin, or sucrose and acacia) and/or as mouth washes and the like, each containing a predetermined amount of a compound of the present invention as an active ingredient. An oligomer of the present invention may also be administered as a bolus, electuary or paste.
[0203] In solid dosage forms of the invention for oral administration (capsules, tablets, pills, dragees, powders, granules, trouches and the like), the active ingredient may be mixed with one or more pharmaceutically-acceptable carriers, such as sodium citrate or dicalcium phosphate, and/or any of the following: (1) fillers or extenders, such as starches, lactose, sucrose, glucose, mannitol, and/or silicic acid; (2) binders, such as, for example, carboxymethylcellulose, alginates, gelatin, polyvinyl pyrrolidone, sucrose and/or acacia; (3) humectants, such as glycerol; (4) disintegrating agents, such as agar-agar, calcium carbonate, potato or tapioca starch, alginic acid, certain silicates, and sodium carbonate; (5) solution retarding agents, such as paraffin; (6) absorption accelerators, such as quaternary ammonium compounds and surfactants, such as poloxamer and sodium lauryl sulfate; (7) wetting agents, such as, for example, cetyl alcohol, glycerol monostearate, and non-ionic surfactants; (8) absorbents, such as kaolin and bentonite clay; (9) lubricants, such as talc, calcium stearate, magnesium stearate, solid polyethylene glycols, sodium lauryl sulfate, zinc stearate, sodium stearate, stearic acid, and mixtures thereof; (10) coloring agents; and (11) controlled release agents such as crospovidone or ethyl cellulose. In the case of capsules, tablets and pills, the pharmaceutical compositions may also comprise buffering agents. Solid compositions of a similar type may also be employed as fillers in soft and hard-shelled gelatin capsules using such excipients as lactose or milk sugars, as well as high molecular weight polyethylene glycols and the like.
[0204] A tablet may be made by compression or molding, optionally with one or more accessory ingredients. Compressed tablets may be prepared using binder (e.g., gelatin or hydroxypropylmethyl cellulose), lubricant, inert diluent, preservative, disintegrant (for example, sodium starch glycolate or cross-linked sodium carboxymethyl cellulose), surface-active or dispersing agent. Molded tablets may be made by molding in a suitable machine a mixture of the powdered compound moistened with an inert liquid diluent.
[0205] The tablets, and other solid dosage forms of the pharmaceutical compositions of the present invention, such as dragees, capsules, pills and granules, may optionally be scored or prepared with coatings and shells, such as enteric coatings and other coatings well known in the pharmaceutical-formulating art. They may also be formulated so as to provide slow or controlled release of the active ingredient therein using, for example, hydroxypropylmethyl cellulose in varying proportions to provide the desired release profile, other polymer matrices, liposomes and/or microspheres. They may be formulated for rapid release, e.g., freeze-dried. They may be sterilized by, for example, filtration through a bacteria-retaining filter, or by incorporating sterilizing agents in the form of sterile solid compositions which can be dissolved in sterile water, or some other sterile injectable medium immediately before use. These compositions may also optionally contain opacifying agents and may be of a composition that they release the active ingredient(s) only, or preferentially, in a certain portion of the gastrointestinal tract, optionally, in a delayed manner. Examples of embedding compositions which can be used include polymeric substances and waxes. The active ingredient can also be in micro-encapsulated form, if appropriate, with one or more of the above-described excipients.
[0206] Liquid dosage forms for oral administration of the compounds of the invention include pharmaceutically acceptable emulsions, microemulsions, solutions, suspensions, syrups and elixirs. In addition to the active ingredient, the liquid dosage forms may contain inert diluents commonly used in the art, such as, for example, water or other solvents, solubilizing agents and emulsifiers, such as ethyl alcohol, isopropyl alcohol, ethyl carbonate, ethyl acetate, benzyl alcohol, benzyl benzoate, propylene glycol, 1,3-butylene glycol, oils (in particular, cottonseed, groundnut, corn, germ, olive, castor and sesame oils), glycerol, tetrahydrofuryl alcohol, polyethylene glycols and fatty acid esters of sorbitan, and mixtures thereof.
[0207] Besides inert diluents, the oral compositions can also include adjuvants such as wetting agents, emulsifying and suspending agents, sweetening, flavoring, coloring, perfuming and preservative agents.
[0208] Suspensions, in addition to the active compounds, may contain suspending agents as, for example, ethoxylated isostearyl alcohols, polyoxyethylene sorbitol and sorbitan esters, microcrystalline cellulose, aluminum metahydroxide, bentonite, agar-agar and tragacanth, and mixtures thereof.
[0209] Formulations for rectal or vaginal administration may be presented as a suppository, which may be prepared by mixing one or more compounds of the invention with one or more suitable nonirritating excipients or carriers comprising, for example, cocoa butter, polyethylene glycol, a suppository wax or a salicylate, and which is solid at room temperature, but liquid at body temperature and, therefore, will melt in the rectum or vaginal cavity and release the active compound.
[0210] Formulations or dosage forms for the topical or transdermal administration of an oligomer as provided herein include powders, sprays, ointments, pastes, creams, lotions, gels, solutions, patches and inhalants. The active oligomers may be mixed under sterile conditions with a pharmaceutically-acceptable carrier, and with any preservatives, buffers, or propellants which may be required. The ointments, pastes, creams and gels may contain, in addition to an active compound of this invention, excipients, such as animal and vegetable fats, oils, waxes, paraffins, starch, tragacanth, cellulose derivatives, polyethylene glycols, silicones, bentonites, silicic acid, talc and zinc oxide, or mixtures thereof.
[0211] Powders and sprays can contain, in addition to an oligomer of the present invention, excipients such as lactose, talc, silicic acid, aluminum hydroxide, calcium silicates and polyamide powder, or mixtures of these substances. Sprays can additionally contain customary propellants, such as chlorofluorohydrocarbons and volatile unsubstituted hydrocarbons, such as butane and propane.
[0212] Transdermal patches have the added advantage of providing controlled delivery of an oligomer of the present invention to the body. Such dosage forms can be made by dissolving or dispersing the oligomer in the proper medium. Absorption enhancers can also be used to increase the flux of the agent across the skin. The rate of such flux can be controlled by either providing a rate controlling membrane or dispersing the agent in a polymer matrix or gel, among other methods known in the art.
[0213] Pharmaceutical compositions suitable for parenteral administration may comprise one or more oligomers of the invention in combination with one or more pharmaceutically-acceptable sterile isotonic aqueous or nonaqueous solutions, dispersions, suspensions or emulsions, or sterile powders which may be reconstituted into sterile injectable solutions or dispersions just prior to use, which may contain sugars, alcohols, antioxidants, buffers, bacteriostats, solutes which render the formulation isotonic with the blood of the intended recipient or suspending or thickening agents. Examples of suitable aqueous and nonaqueous carriers which may be employed in the pharmaceutical compositions of the invention include water, ethanol, polyols (such as glycerol, propylene glycol, polyethylene glycol, and the like), and suitable mixtures thereof, vegetable oils, such as olive oil, and injectable organic esters, such as ethyl oleate. Proper fluidity can be maintained, for example, by the use of coating materials, such as lecithin, by the maintenance of the required particle size in the case of dispersions, and by the use of surfactants.
[0214] These compositions may also contain adjuvants such as preservatives, wetting agents, emulsifying agents and dispersing agents. Prevention of the action of microorganisms upon the subject oligomers may be ensured by the inclusion of various antibacterial and antifungal agents, for example, paraben, chlorobutanol, phenol sorbic acid, and the like. It may also be desirable to include isotonic agents, such as sugars, sodium chloride, and the like into the compositions. In addition, prolonged absorption of the injectable pharmaceutical form may be brought about by the inclusion of agents which delay absorption such as aluminum monostearate and gelatin.
[0215] In some cases, in order to prolong the effect of a drug, it is desirable to slow the absorption of the drug from subcutaneous or intramuscular injection. This may be accomplished by the use of a liquid suspension of crystalline or amorphous material having poor water solubility, among other methods known in the art. The rate of absorption of the drug then depends upon its rate of dissolution which, in turn, may depend upon crystal size and crystalline form. Alternatively, delayed absorption of a parenterally-administered drug form is accomplished by dissolving or suspending the drug in an oil vehicle.
[0216] Injectable depot forms may be made by forming microencapsule matrices of the subject oligomers in biodegradable polymers such as polylactide-polyglycolide. Depending on the ratio of oligomer to polymer, and the nature of the particular polymer employed, the rate of oligomer release can be controlled. Examples of other biodegradable polymers include poly(orthoesters) and poly(anhydrides). Depot injectable formulations may also prepared by entrapping the drug in liposomes or microemulsions that are compatible with body tissues.
[0217] When the oligomers of the present invention are administered as pharmaceuticals, to humans and animals, they can be given per se or as a pharmaceutical composition containing, for example, 0.1 to 99% (more preferably, 10 to 30%) of active ingredient in combination with a pharmaceutically acceptable carrier.
[0218] As noted above, the formulations or preparations of the present invention may be given orally, parenterally, topically, or rectally. They are typically given in forms suitable for each administration route. For example, they are administered in tablets or capsule form, by injection, inhalation, eye lotion, ointment, suppository, etc. administration by injection, infusion or inhalation; topical by lotion or ointment; and rectal by suppositories.
[0219] The phrases "parenteral administration" and "administered parenterally" as used herein means modes of administration other than enteral and topical administration, usually by injection, and includes, without limitation, intravenous, intramuscular, intraarterial, intrathecal, intracapsular, intraorbital, intracardiac, intradermal, intraperitoneal, transtracheal, subcutaneous, subcuticular, intraarticulare, subcapsular, subarachnoid, intraspinal and intrastemal injection and infusion.
[0220] The phrases "systemic administration," "administered systemically," "peripheral administration" and "administered peripherally" as used herein mean the administration of a compound, drug or other material other than directly into the central nervous system, such that it enters the patient's system and, thus, is subject to metabolism and other like processes, for example, subcutaneous administration.
[0221] Regardless of the route of administration selected, the oligomers of the present invention, which may be used in a suitable hydrated form, and/or the pharmaceutical compositions of the present invention, may be formulated into pharmaceutically-acceptable dosage forms by conventional methods known to those of skill in the art. Actual dosage levels of the active ingredients in the pharmaceutical compositions of this invention may be varied so as to obtain an amount of the active ingredient which is effective to achieve the desired therapeutic response for a particular patient, composition, and mode of administration, without being unacceptably toxic to the patient.
[0222] The selected dosage level will depend upon a variety of factors including the activity of the particular oligomer of the present invention employed, or the ester, salt or amide thereof, the route of administration, the time of administration, the rate of excretion or metabolism of the particular oligomer being employed, the rate and extent of absorption, the duration of the treatment, other drugs, compounds and/or materials used in combination with the particular oligomer employed, the age, sex, weight, condition, general health and prior medical history of the patient being treated, and like factors well known in the medical arts.
[0223] A physician or veterinarian having ordinary skill in the art can readily determine and prescribe the effective amount of the pharmaceutical composition required. For example, the physician or veterinarian could start doses of the compounds of the invention employed in the pharmaceutical composition at levels lower than that required in order to achieve the desired therapeutic effect and gradually increase the dosage until the desired effect is achieved. In general, a suitable daily dose of a compound of the invention will be that amount of the compound which is the lowest dose effective to produce a therapeutic effect. Such an effective dose will generally depend upon the factors described above. Generally, oral, intravenous, intracerebroventricular and subcutaneous doses of the compounds of this invention for a patient, when used for the indicated effects, will range from about 0.0001 to about 100 mg per kilogram of body weight per day.
[0224] If desired, the effective daily dose of the active compound may be administered as two, three, four, five, six or more sub-doses administered separately at appropriate intervals throughout the day, optionally, in unit dosage forms. In certain situations, dosing is one administration per day. In certain embodiments, dosing is one or more administration per every 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 days, or every 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12 weeks, or every 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12 months, as needed, to maintain the desired expression of a functional dystrophin protein.
[0225] Nucleic acid molecules can be administered to cells by a variety of methods known to those familiar to the art, including, but not restricted to, encapsulation in liposomes, by iontophoresis, or by incorporation into other vehicles, such as hydrogels, cyclodextrins, biodegradable nanocapsules, and bioadhesive microspheres, as described herein and known in the art. In certain embodiments, microemulsification technology may be utilized to improve bioavailability of lipophilic (water insoluble) pharmaceutical agents. Examples include Trimetrine (Dordunoo, S. K., et al., Drug Development and Industrial Pharmacy, 17(12), 1685-1713, 1991 and REV 5901 (Sheen, P. C., et al., J Pharm Sci 80(7), 712-714, 1991). Among other benefits, microemulsification provides enhanced bioavailability by preferentially directing absorption to the lymphatic system instead of the circulatory system, which thereby bypasses the liver, and prevents destruction of the compounds in the hepatobiliary circulation.
[0226] In one aspect of invention, the formulations contain micelles formed from an oligomer as provided herein and at least one amphiphilic carrier, in which the micelles have an average diameter of less than about 100 nm. More preferred embodiments provide micelles having an average diameter less than about 50 nm, and even more preferred embodiments provide micelles having an average diameter less than about 30 nm, or even less than about 20 nm.
[0227] While all suitable amphiphilic carriers are contemplated, the presently preferred carriers are generally those that have Generally-Recognized-as-Safe (GRAS) status, and that can both solubilize the compound of the present invention and microemulsify it at a later stage when the solution comes into a contact with a complex water phase (such as one found in human gastro-intestinal tract). Usually, amphiphilic ingredients that satisfy these requirements have HLB (hydrophilic to lipophilic balance) values of 2-20, and their structures contain straight chain aliphatic radicals in the range of C-6 to C-20. Examples are polyethylene-glycolized fatty glycerides and polyethylene glycols.
[0228] Examples of amphiphilic carriers include saturated and monounsaturated polyethyleneglycolyzed fatty acid glycerides, such as those obtained from fully or partially hydrogenated various vegetable oils. Such oils may advantageously consist of tri-, di-, and mono-fatty acid glycerides and di- and mono-polyethyleneglycol esters of the corresponding fatty acids, with a particularly preferred fatty acid composition including capric acid 4-10, capric acid 3-9, lauric acid 40-50, myristic acid 14-24, palmitic acid 4-14 and stearic acid 5-15%. Another useful class of amphiphilic carriers includes partially esterified sorbitan and/or sorbitol, with saturated or mono-unsaturated fatty acids (SPAN-series) or corresponding ethoxylated analogs (TWEEN-series).
[0229] Commercially available amphiphilic carriers may be particularly useful, including Gelucire-series, Labrafil, Labrasol, or Lauroglycol (all manufactured and distributed by Gattefosse Corporation, Saint Priest, France), PEG-mono-oleate, PEG-di-oleate, PEG-mono-laurate and di-laurate, Lecithin, Polysorbate 80, etc (produced and distributed by a number of companies in USA and worldwide).
[0230] In certain embodiments, the delivery may occur by use of liposomes, nanocapsules, microparticles, microspheres, lipid particles, vesicles, and the like, for the introduction of the compositions of the present invention into suitable host cells. In particular, the compositions of the present invention may be formulated for delivery either encapsulated in a lipid particle, a liposome, a vesicle, a nanosphere, a nanoparticle or the like. The formulation and use of such delivery vehicles can be carried out using known and conventional techniques.
[0231] Hydrophilic polymers suitable for use in the present invention are those which are readily water-soluble, can be covalently attached to a vesicle-forming lipid, and which are tolerated in vivo without toxic effects (i.e., are biocompatible). Suitable polymers include polyethylene glycol (PEG), polylactic (also termed polylactide), polyglycolic acid (also termed polyglycolide), a polylactic-polyglycolic acid copolymer, and polyvinyl alcohol. In certain embodiments, polymers have a molecular weight of from about 100 or 120 daltons up to about 5,000 or 10,000 daltons, or from about 300 daltons to about 5,000 daltons. In other embodiments, the polymer is polyethyleneglycol having a molecular weight of from about 100 to about 5,000 daltons, or having a molecular weight of from about 300 to about 5,000 daltons. In certain embodiments, the polymer is polyethyleneglycol of 750 daltons (PEG(750)). Polymers may also be defined by the number of monomers therein; a preferred embodiment of the present invention utilizes polymers of at least about three monomers, such PEG polymers consisting of three monomers (approximately 150 daltons).
[0232] Other hydrophilic polymers which may be suitable for use in the present invention include polyvinylpyrrolidone, polymethoxazoline, polyethyloxazoline, polyhydroxypropyl methacrylamide, polymethacrylamide, polydimethylacrylamide, and derivatized celluloses such as hydroxymethylcellulose or hydroxyethylcellulose.
[0233] In certain embodiments, a formulation of the present invention comprises a biocompatible polymer selected from the group consisting of polyamides, polycarbonates, polyalkylenes, polymers of acrylic and methacrylic esters, polyvinyl polymers, polyglycolides, polysiloxanes, polyurethanes and copolymers thereof, celluloses, polypropylene, polyethylenes, polystyrene, polymers of lactic acid and glycolic acid, polyanhydrides, poly(ortho)esters, poly(butic acid), poly(valeric acid), poly(lactide-co-caprolactone), polysaccharides, proteins, polyhyaluronic acids, polycyanoacrylates, and blends, mixtures, or copolymers thereof.
[0234] Cyclodextrins are cyclic oligosaccharides, consisting of 6, 7 or 8 glucose units, designated by the Greek letter .alpha., .beta.. or .gamma., respectively. The glucose units are linked by .alpha.-1,4-glucosidic bonds. As a consequence of the chair conformation of the sugar units, all secondary hydroxyl groups (at C-2, C-3) are located on one side of the ring, while all the primary hydroxyl groups at C-6 are situated on the other side. As a result, the external faces are hydrophilic, making the cyclodextrins water-soluble. In contrast, the cavities of the cyclodextrins are hydrophobic, since they are lined by the hydrogen of atoms C-3 and C-5, and by ether-like oxygens. These matrices allow complexation with a variety of relatively hydrophobic compounds, including, for instance, steroid compounds such as 17.alpha.-estradiol (see, e.g., van Uden et al. Plant Cell Tiss. Org. Cult. 38:1-3-113 (1994)). The complexation takes place by Van der Waals interactions and by hydrogen bond formation. For a general review of the chemistry of cyclodextrins, see, Wenz, Agnew. Chem. Int. Ed. Engl., 33:803-822 (1994).
[0235] The physico-chemical properties of the cyclodextrin derivatives depend strongly on the kind and the degree of substitution. For example, their solubility in water ranges from insoluble (e.g., triacetyl-beta-cydodextrin) to 147% soluble (w/v) (G-2-beta-cydodextrin). In addition, they are soluble in many organic solvents. The properties of the cyclodextrins enable the control over solubility of various formulation components by increasing or decreasing their solubility.
[0236] Numerous cyclodextrins and methods for their preparation have been described. For example, Parmeter (I), et al. (U.S. Pat. No. 3,453,259) and Gramera, et al. (U.S. Pat. No. 3,459,731) described electroneutral cyclodextrins. Other derivatives include cyclodextrins with cationic properties [Parmeter (II), U.S. Pat. No. 3,453,257], insoluble crosslinked cyclodextrins (Solms, U.S. Pat. No. 3,420,788), and cyclodextrins with anionic properties [Parmeter (III), U.S. Pat. No. 3,426,011]. Among the cyclodextrin derivatives with anionic properties, carboxylic acids, phosphorous acids, phosphinous acids, phosphonic acids, phosphoric acids, thiophosphonic acids, thiosulphinic acids, and sulfonic acids have been appended to the parent cyclodextrin [see, Parmeter (III), supra]. Furthermore, sulfoalkyl ether cyclodextrin derivatives have been described by Stella, et al. (U.S. Pat. No. 5,134,127).
[0237] Liposomes consist of at least one lipid bilayer membrane enclosing an aqueous internal compartment. Liposomes may be characterized by membrane type and by size. Small unilamellar vesicles (SUVs) have a single membrane and typically range between 0.02 and 0.05 .mu.m in diameter; large unilamellar vesicles (LUVS) are typically larger than 0.05 .mu.m. Oligolamellar large vesicles and multilamellar vesicles have multiple, usually concentric, membrane layers and are typically larger than 0.1 .mu.m. Liposomes with several nonconcentric membranes, i.e., several smaller vesicles contained within a larger vesicle, are termed multivesicular vesicles.
[0238] One aspect of the present invention relates to formulations comprising liposomes containing an oligomer of the present invention, where the liposome membrane is formulated to provide a liposome with increased carrying capacity. Alternatively or in addition, the compound of the present invention may be contained within, or adsorbed onto, the liposome bilayer of the liposome. An oligomer of the present invention may be aggregated with a lipid surfactant and carried within the liposome's internal space; in these cases, the liposome membrane is formulated to resist the disruptive effects of the active agent-surfactant aggregate.
[0239] According to one embodiment of the present invention, the lipid bilayer of a liposome contains lipids derivatized with polyethylene glycol (PEG), such that the PEG chains extend from the inner surface of the lipid bilayer into the interior space encapsulated by the liposome, and extend from the exterior of the lipid bilayer into the surrounding environment.
[0240] Active agents contained within liposomes of the present invention are in solubilized form. Aggregates of surfactant and active agent (such as emulsions or micelles containing the active agent of interest) may be entrapped within the interior space of liposomes according to the present invention. A surfactant acts to disperse and solubilize the active agent, and may be selected from any suitable aliphatic, cycloaliphatic or aromatic surfactant, including but not limited to biocompatible lysophosphatidylcholines (LPCs) of varying chain lengths (for example, from about C14 to about C20). Polymer-derivatized lipids such as PEG-lipids may also be utilized for micelle formation as they will act to inhibit micelle/membrane fusion, and as the addition of a polymer to surfactant molecules decreases the CMC of the surfactant and aids in micelle formation. Preferred are surfactants with CMCs in the micromolar range; higher CMC surfactants may be utilized to prepare micelles entrapped within liposomes of the present invention.
[0241] Liposomes according to the present invention may be prepared by any of a variety of techniques that are known in the art. See, e.g., U.S. Pat. No. 4,235,871; Published PCT applications WO 96/14057; New RRC, Liposomes: A practical approach, IRL Press, Oxford (1990), pages 33-104; Lasic D D, Liposomes from physics to applications, Elsevier Science Publishers BV, Amsterdam, 1993. For example, liposomes of the present invention may be prepared by diffusing a lipid derivatized with a hydrophilic polymer into preformed liposomes, such as by exposing preformed liposomes to micelles composed of lipid-grafted polymers, at lipid concentrations corresponding to the final mole percent of derivatized lipid which is desired in the liposome. Liposomes containing a hydrophilic polymer can also be formed by homogenization, lipid-field hydration, or extrusion techniques, as are known in the art.
[0242] In another exemplary formulation procedure, the active agent is first dispersed by sonication in a lysophosphatidylcholine or other low CMC surfactant (including polymer grafted lipids) that readily solubilizes hydrophobic molecules. The resulting micellar suspension of active agent is then used to rehydrate a dried lipid sample that contains a suitable mole percent of polymer-grafted lipid, or cholesterol. The lipid and active agent suspension is then formed into liposomes using extrusion techniques as are known in the art, and the resulting liposomes separated from the unencapsulated solution by standard column separation.
[0243] In one aspect of the present invention, the liposomes are prepared to have substantially homogeneous sizes in a selected size range. One effective sizing method involves extruding an aqueous suspension of the liposomes through a series of polycarbonate membranes having a selected uniform pore size; the pore size of the membrane will correspond roughly with the largest sizes of liposomes produced by extrusion through that membrane. See e.g., U.S. Pat. No. 4,737,323 (Apr. 12, 1988). In certain embodiments, reagents such as DharmaFECT.RTM. and Lipofectamine.RTM. may be utilized to introduce polynucleotides or proteins into cells.
[0244] The release characteristics of a formulation of the present invention depend on the encapsulating material, the concentration of encapsulated drug, and the presence of release modifiers. For example, release can be manipulated to be pH dependent, for example, using a pH sensitive coating that releases only at a low pH, as in the stomach, or a higher pH, as in the intestine. An enteric coating can be used to prevent release from occurring until after passage through the stomach. Multiple coatings or mixtures of cyanamide encapsulated in different materials can be used to obtain an initial release in the stomach, followed by later release in the intestine. Release can also be manipulated by inclusion of salts or pore forming agents, which can increase water uptake or release of drug by diffusion from the capsule. Excipients which modify the solubility of the drug can also be used to control the release rate. Agents which enhance degradation of the matrix or release from the matrix can also be incorporated. They can be added to the drug, added as a separate phase (i.e., as particulates), or can be co-dissolved in the polymer phase depending on the compound. In most cases the amount should be between 0.1 and thirty percent (w/w polymer). Types of degradation enhancers include inorganic salts such as ammonium sulfate and ammonium chloride, organic acids such as citric acid, benzoic acid, and ascorbic acid, inorganic bases such as sodium carbonate, potassium carbonate, calcium carbonate, zinc carbonate, and zinc hydroxide, and organic bases such as protamine sulfate, spermine, choline, ethanolamine, diethanolamine, and triethanolamine and surfactants such as Tween.RTM. and Pluronic.RTM.. Pore forming agents which add microstructure to the matrices (i.e., water soluble compounds such as inorganic salts and sugars) are added as particulates. The range is typically between one and thirty percent (w/w polymer).
[0245] Uptake can also be manipulated by altering residence time of the particles in the gut. This can be achieved, for example, by coating the particle with, or selecting as the encapsulating material, a mucosal adhesive polymer. Examples include most polymers with free carboxyl groups, such as chitosan, celluloses, and especially polyacrylates (as used herein, polyacrylates refers to polymers including acrylate groups and modified acrylate groups such as cyanoacrylates and methacrylates).
[0246] An oligomer may be formulated to be contained within, or, adapted to release by a surgical or medical device or implant. In certain aspects, an implant may be coated or otherwise treated with an oligomer. For example, hydrogels, or other polymers, such as biocompatible and/or biodegradable polymers, may be used to coat an implant with the compositions of the present invention (i.e., the composition may be adapted for use with a medical device by using a hydrogel or other polymer). Polymers and copolymers for coating medical devices with an agent are well-known in the art. Examples of implants include, but are not limited to, stents, drug-eluting stents, sutures, prosthesis, vascular catheters, dialysis catheters, vascular grafts, prosthetic heart valves, cardiac pacemakers, implantable cardioverter defibrillators, IV needles, devices for bone setting and formation, such as pins, screws, plates, and other devices, and artificial tissue matrices for wound healing.
[0247] In addition to the methods provided herein, the oligomers for use according to the invention may be formulated for administration in any convenient way for use in human or veterinary medicine, by analogy with other pharmaceuticals. The antisense oligomers and their corresponding formulations may be administered alone or in combination with other therapeutic strategies in the treatment of muscular dystrophy, such as myoblast transplantation, stem cell therapies, administration of aminoglycoside antibiotics, proteasome inhibitors, and up-regulation therapies (e.g., upregulation of utrophin, an autosomal paralogue of dystrophin).
[0248] All publications and patent applications cited in this specification are herein incorporated by reference as if each individual publication or patent application were specifically and individually indicated to be incorporated by reference.
[0249] Although the foregoing invention has been described in some detail by way of illustration and example for purposes of clarity of understanding, it will be readily apparent to one of ordinary skill in the art in light of the teachings of this invention that certain changes and modifications may be made thereto without departing from the spirit or scope of the appended claims. The following examples are provided by way of illustration only and not by way of limitation. Those of skill in the art will readily recognize a variety of noncritical parameters that could be changed or modified to yield essentially similar results.
REFERENCES
[0250] Aartsma-Rus, A., A. A. Janson, et al. (2004). "Antisense-induced multiexon skipping for Duchenne muscular dystrophy makes more sense." Am J Hum Genet 74(1): 83-92. [0251] Dunckley, M. G., I. C. Eperon, et al. (1997). "Modulation of splicing in the DMD gene by antisense oligoribonucleotides." Nucleosides & Nucleotides 16(7-9): 1665-1668. [0252] Dunckley, M. G., M. Manoharan, et al. (1998). "Modification of splicing in the dystrophin gene in cultured Mdx muscle cells by antisense oligoribonudeotides." Hum Mol Genet 7(7): 1083-90. [0253] Errington, S. J., C. J. Mann, et al. (2003). "Target selection for antisense oligonucleotide induced exon skipping in the dystrophin gene." J Gene Med 5(6): 518-27. [0254] Jearawiriyapaisam, N., H. M. Moulton, et al. (2008). "Sustained Dystrophin Expression Induced by Peptide-conjugated Morpholino Oligomers in the Muscles of mdx Mice." Mol Ther. [0255] Lu, Q. L., C. J. Mann, et al. (2003). "Functional amounts of dystrophin produced by skipping the mutated exon in the mdx dystrophic mouse." Nat Med 9(8): 1009-14. [0256] Mann, C. J., K. Honeyman, et al. (2002). "Improved antisense oligonucleotide induced exon skipping in the mdx mouse model of muscular dystrophy." J Gene Med 4(6): 644-54. [0257] Marshall, N. B., S. K. Oda, et al. (2007). "Arginine-rich cell-penetrating peptides facilitate delivery of antisense oligomers into murine leukocytes and alter pre-mRNA splicing." Journal of Immunological Methods 325(1-2): 114-126. [0258] Matsuo, M., T. Masumura, et al. (1991). "Exon skipping during splicing of dystrophin mRNA precursor due to an intraexon deletion in the dystrophin gene of Duchenne muscular dystrophy kobe." J Clin Invest 87(6): 2127-31. [0259] Monaco, A. P., C. J. Bertelson, et al. (1988). "An explanation for the phenotypic differences between patients bearing partial deletions of the DMD locus." Genomics 2(1): 90-5. [0260] Pramono, Z. A., Y. Takeshima, et al. (1996). "Induction of exon skipping of the dystrophin transcript in lymphoblastoid cells by transfecting an antisense oligodeoxynucleotide complementary to an exon recognition sequence." Biochem Biophys Res Commun 226(2): 445-9. [0261] Sazani, P., R. Kole, et al. (2007). Splice switching oligomers for the TNF superfamily receptors and their use in treatment of disease. PCT WO2007058894, University of North Carolina [0262] Sierakowska, H., M. J. Sambade, et al. (1996). "Repair of thalassemic human beta-globin mRNA in mammalian cells by antisense oligonucleotides." Proc Natl Acad Sci USA 93(23): 12840-4. [0263] Summerton, J. and D. Weller (1997). "Morpholino antisense oligomers: design, preparation, and properties." Antisense Nucleic Acid Drug Dev 7(3): 187-95. [0264] Takeshima, Y., H. Nishio, et al. (1995). "Modulation of in vitro splicing of the upstream intron by modifying an intra-exon sequence which is deleted from the dystrophin gene in dystrophin Kobe." J Clin Invest 95(2): 515-20. [0265] van Deutekom, J. C., M. Bremmer-Bout, et al. (2001). "Antisense-induced exon skipping restores dystrophin expression in DMD patient derived muscle cells." Hum Mol Genet 10(15): 1547-54. [0266] van Deutekom, J. C., A. A. Janson, et al. (2007). "Local dystrophin restoration with antisense oligonucleotide PRO051." N Enal J Med 357(26): 2677-86. [0267] Wilton, S. D., A. M. Fall, et al. (2007). "Antisense oligonucleotide-induced exon skipping across the human dystrophin gene transcript." Mol Ther 15(7): 1288-96. [0268] Wilton, S. D., F. Lloyd, et al. (1999). "Specific removal of the nonsense mutation from the mdx dystrophin mRNA using antisense oligonucleotides." Neuromuscul Disord 9(5): 330-8. [0269] Wu, B., H. M. Moulton, et al. (2008). "Effective rescue of dystrophin improves cardiac function in dystrophin-deficient mice by a modified morpholino oligomer." Proc Natl Acad Sci USA 105(39): 14814-9. [0270] Yin, H., H. M. Moulton, et al. (2008). "Cell-penetrating peptide-conjugated antisense oligonucleotides restore systemic muscle and cardiac dystrophin expression and function." Hum Mol Genet 17(24): 3909-18.
EXAMPLES
Materials and Methods
[0271] Cells and Tissue Culture Treatment Conditions
[0272] Human Rhabdomyosarcoma cells (ATCC, CCL-136; RD cells) preserved in a 5% DMSO solution (Sigma) at a low passage number were thawed in a 37.degree. C. water bath until the ice sliver was no longer visible. Cells were seeded into tissue culture-treated T75 flasks (Nunc) at 1.5.times.10.sup.6 cells/flask in 24 mL of warmed DMEM with L-Glutamine (HyClone), 10% fetal bovine serum, and 1% Penicillin-Streptomycin antibiotic solution (CelGro); after 24 hours, media was aspirated, cells were washed once in warmed PBS, and fresh media was added. Cells were grown to 80% confluence in a 37.degree. C. incubator at 5.0% CO2.
[0273] Media was aspirated from T75 flasks; cells were washed once in warmed PBS and aspirated. 3 mL of Trypsin/EDTA, warmed in a 37.degree. C. water bath, was added to each T75. Cells were incubated at 37.degree. C. 5 2-5 minutes until, with gentle agitation, they released from the flask. Cell suspension was transferred to a 15.0 mL conical tube; flasks were rinsed with 1.0 mL of Trypsin/EDTA solution to gather remaining cells. Cells were counted with a Vi-Cell XR cell counter (Beckman Coulter). Cells were seeded into tissue culture-treated 12-well plates (Falcon) at 2.0.times.10.sup.5 viable cells per well in 1.0 mL media. Cells were incubated overnight in a 37.degree. C. incubator at 5.0% CO.sub.2.
[0274] Twelve-well seeded plates were examined for even cellular distribution and plate adherence. Lyophilized peptide conjugated phosphorodiamidate morpholino oligomers (PPMOs) were re-suspended at 2.0 mM in nuclease-free water (Ambion), and kept on ice during cell treatment; to verify molarity, PPMOs were measured using a NanoDrop 2000 spectrophotometer (Thermo Scientific). Immediately prior to PPMO treatment, media was aspirated, and cells were rinsed in warmed PBS. PPMOs were diluted in warmed media to the desired molarity; cells were treated in a total of 1.0 mL PPMO per well. PPMOs were tested in triplicate. For no-treatment controls, fresh, warmed media was added in 1.0 mL total volume. Cells were incubated for 48 hours in a 37.degree. C. incubator at 5.0% C02.
[0275] RNA Extraction
[0276] Media was aspirated, and cells were rinsed in warmed PBS. RNA was extracted with the QuickGene-Mini80 system, QuickGene RNA cultured cell HC kit S, and MagNAlyser with ceramic bead homogenization using the manufacturers' recommended protocols. Briefly, cells were lysed in treatment plates with 350 uL LRP (10 uL .beta.-Mercaptoethanol added per 100 uL LRP) lysis buffer; homogenate was gently triturated to ensure full lysis, and transferred to MagNAlyser tubes. Tubes were spun at 2800 rpm for 30 seconds in the MagNAlyser to ensure full homogenization, and iced briefly. 50 uL SRP solubilization buffer was added and homogenate was vortexed for 15 seconds. 170 uL >99% ethanol was added to each tube, and homogenate was vortexed for 60 seconds. Homogenate was flash-spun and transferred to Mini80 RNA cartridges, samples were pressurized and flow-through was discarded. Cartridges were washed in 750 uL WRP wash buffer and pressurized. 40 uL of DNase solution (1.25 uL Qiagen DNasel, 35 uL RDD Buffer, 3.75 uL nuclease-free water) was added directly to the cartridge membrane; cartridges were incubated four minutes at room temperature. Cartridges were washed twice with 750 uL WRP, pressurizing after each wash. Cartridges were placed over nuclease-free tubes. 50 uL CRP elution buffer was added to each membrane; membranes were incubated for five minutes at room-temperature. Cartridges were pressurized and eluate was collected. RNA was stored at -80.degree. C. pending quantification. RNA was quantified using the NanoDrop.TM. 2000 spectrophotometer.
[0277] Nested RT-PCR
[0278] Primer-specific, exon-specific, optimized nested RT-PCR amplification was performed using the primer pair sets for each dystrophin exon as shown below in Table 1.
TABLE-US-00003 TABLE 1 Primer pair sets used to PCR amplify human dystrophin mRNA to detect exon-skipping SEQ Name F/R I/O Sequence (5'-3') Exon Purpose ID NO: PS170 F O CCAGAGCTTTACCTGAGAAACAAG 48 Detection 640 PS172 F I CCAGCCACTCAGCCAGTGAAG 49 of Exon 50 641 PS174 R I CGATCCGTAATGATTGTTCTAGCC 52 and 51 642 PS176 R O CATTTCATTCAACTGTTGCCTCCG 53 Skipping in 643 Human Dystrophin PS186 F O CAATGCTCCTGACCTCTGTGC 42 Detection 644 PS187 F I GTCTACAACAAAGCTCAGGTCG 43 of Exon 44 645 PS189 F I GCAATGTTATCTGCTTCCTCCAACC 46 and 45 646 PS190 R O GCTCTTTTCCAGGTTCAAGTGG 46 Skipping in 647 Human Dystrophin PS192 F O CTTGGACAGAACTTACCGACTGG 51 Detection 648 PS193 F I GCAGGATTTGGAACAGAGGCG 52 of Exon 53 649 PS195 R I CATCTACATTTGTCTGCCACTGG 54 Skipping in 650 PS197 R O GTTTCTTCCAAAGCAGCCTCTCG 55 Human 651 Dystrophin
[0279] The indicated primer pairs are shown as either forward or reverse (F/R) and either outer or inner primer pairs (I/O) corresponding to primary or secondary amplifications, respectively. The location of the primer target is indicated in the Exon column and the Purpose indicates the exon-skipping events can be detected. For example, PS170 and PS176 primers amplify a region from exon 48 to 53 in the primary amplification. Primers PS172 and PS174 then amplify a region from exon 49 to 52 in the secondary amplification. This nested PCR reaction will detect exon skipping of both exons 50 and/or exon 51. The specific nested RT-PCR reaction conditions are provided below.
[0280] RNA extracted from treated cells (described above) was diluted to 20 ng/ul for all samples.
TABLE-US-00004 TABLE 2 Reaction setup for RT-PCR and primary amplification (50 .mu.l reaction): 2x Reaction mix 25 .mu.l PS XXX Forward Primer (30 .mu.M) 0.5 .mu.l (see Table 1) PS XXX Reverse Primer (30 .mu.M) 0.5 .mu.l (see Table 1) Superscript III Platinum Taq mix 2 .mu.l Template RNA (20 ng/.mu.l) 10 .mu.l Nuclease-Free Water (50 .mu.l total 12 .mu.l volume)
TABLE-US-00005 TABLE 3 RT-PCR and primary amplification program: Temperature Time Reverse 55.degree. C. 30 minutes Transcription RT Inactivation 94.degree. C. 2 minutes Denaturing 94.degree. C. 1 minute 8 Cycles Annealing 59.degree. C. 1 minute Extension 68.degree. C. 1 minute 4.degree. C. .infin.
TABLE-US-00006 TABLE 4 Reaction setup for nested secondary amplification (50 .mu.l reaction): 10x PCR Buffer 5 .mu.l dNTP solution (10 mM) 0.5 .mu.l 50 mM MgCl 1.5 .mu.l PS XXX Forward Primer (30 .mu.M) 0.33 .mu.l (see Table 1) PS XXX Reverse Primer (30 .mu.M) 0.33 .mu.l (see Table 1) Platinum Taq DNA polymerase 0.2 .mu.l 0.1 mM Cy5-dCTP 1 .mu.l RT-PCR product (from Step 1) 1 .mu.l Nuclease-Free Water (50 .mu.l total 40.15 .mu.l volume)
TABLE-US-00007 TABLE 5 Nested secondary amplification program: Temperature Time Primary 94.degree. C. 3 minutes Denature Denaturing 94.degree. C. 45 seconds 28-30 Annealing 59.degree. C. 30 seconds Cycles Extension 68.degree. C. 1 minute 4.degree. C. .infin.
[0281] Gel Electrophoresis Analysis
[0282] Ten microliters of 5.times. Ficoll loading dye was added to each 50 microliter nested RT-PCR reaction. Fifteen microliters of PCR/dye mixture was run on a 10% TBE gel at 300 volts for 30 minutes. After electrophoresis, the gel was washed in diH2O for at least one hour, changing the water every 30 minutes. The gel was then scanned on a Typhoon Trio Variable Mode Imager (GE Healthcare). For exon 44 skipping, the nested RT-PCR product from full-length dystrophin transcript is 571 bp, and 423 bp from Exon 44-skipped mRNA (exon 44 is 148 bp). For exon 45, the nested RT-PCR product from full-length dystrophin transcript is 571 bp, and 395 bp from Exon 45-skipped mRNA (exon 45 is 176 bp). For exon 53, the PCR product from full-length dystrophin transcript is 365 bp, and 153 bp from exon 53-skipped mRNA (exon 53 is 212 bp).
[0283] The gel images were subjected to quantitative analysis by measuring the band intensities of the full-length PCR product compared to the exon-skipped product. In some cases, the percent skipping at a fixed PPMO concentration (e.g., 3 micromolar) was used to determine the relative activity of a series of PPMO to induce exon skipping of a given exon. In other situations, a PPMO dose-range was used to treat cells (e.g., 0.1, 0.3, 1.0, 3.0 and 10 micromolar) and an EC.sub.50 was calculated based on the percent skipping induced at each concentration.
Example 1
Exon 51 Scan
[0284] A series of overlapping antisense PPMOs that target human dystrophin exon 51 were designed, synthesized and used to treat either human rhabdomyosarcoma cells (RD cells) or primary human skeletal muscle cells. This strategy is termed an "exon scan" and was used similarly for several other dystrophin exons as described below. All the PPMOs were synthesized as peptide-conjugated PMO (PPMO) using the CP06062 peptide (SEQ ID NO: 578) and a 3' terminal PMO linkage. For exon 51, a series of 26 PPMOs, each 26 bases in length, were made (SEQ ID NOS: 309-311, 314, 316, 317, 319, 321, 323, 324, 326, 327, 329-331, 333, 335, 336, 338-345) as shown in FIG. 2A. The PPMOs were evaluated for exon skipping efficacy by treating RD cells at various concentrations as described above in the Materials and Methods. Three PPMOs (SEQ ID NOS: 324, 326 and 327) were identified as effective in inducing exon-skipping and selected for additional evaluation. Dose-ranging experiments in RD cells and primary human skeletal muscle cells were used to confirm the relative efficacy of these three PPMO sequences. SEQ ID NO: 327 was shown to be most effective at inducing exon 51 skipping as shown in FIGS. 2B and 2C.
[0285] A comparison of the relative effectiveness of SEQ ID NO: 327 to other exon 51-targeted antisense sequences was performed in RD cells and primary human skeletal muscle cells, as described above. All the evaluated sequences were made as peptide-conjugated PMOs using the CP06062 peptide (SEQ ID NO: 578). This allowed direct comparison of the relative effectiveness of the antisense sequences without regard to antisense chemistry or cell delivery. The relative location of the certain exon 51-targeted oligos compared to SEQ ID NO: 327 is shown in FIG. 2D. As shown in FIG. 2C, there is a ranked hierarchy of exon-skipping effectiveness, with SEQ ID NO: 327 being the most effective by at least a factor of several-fold compared to other sequences.
Example 2
Exon 50 Scan
[0286] A series of overlapping antisense PPMOs that target human dystrophin exon 50 were designed and synthesized. For exon 50, a series of 17 PPMOs, each 25 bases in length, were made (SEQ ID NOS:267, 269, 271, 273, 275, 277, 279, 280, 282 and 284-291) as shown in FIG. 3A. The PPMOs were evaluated for exon skipping efficacy by treating RD cells at various concentrations as described above in the Materials and Methods. Four PPMOs (SEQ ID NOS: 277, 287, 290 and 291) were identified as effective in inducing exon-skipping and selected for additional evaluation. Dose-ranging experiments in RD cells were used to confirm the relative efficacy of these four PMO sequences. SEQ ID NOs: 584 (AVI-5656) and 287 (AVI-5038) were shown to be most effective at inducing exon 50 skipping as shown in FIG. 3B. The EC.sub.50 values were derived from the dose-ranging experiments and represent the calculated concentration where 50% of the PCR product is from the mRNA lacking exon 50 relative to the PCR product produced from the mRNA containing exon 50. Compared to other sequences (see, e.g., SEQ ID NOs: 584 and 585 correspond to SEQ ID NOs: 173 and 175 in WO2006/000057, respectively) AVI-5038 (SEQ ID NO: 287) is equivalent or better at inducing exon-skipping activity in the RD cell assay as shown in FIG. 3B.
Example 3
Exon 53 Scan
[0287] A series of overlapping antisense PPMOs that target human dystrophin exon 53 were designed and synthesized. For exon 53, a series of 24 PPMOs, each 25 bases in length, were made (SEQ ID NOS:416, 418, 420, 422, 424, 426, 428, 429, 431, 433, 434, 436, 438-440 and 443-451) as shown in FIG. 4A. The PPMOs were evaluated for exon skipping efficacy by treating RD cells and primary human skeletal muscle cells at various concentrations as described above in the Materials and Methods. Three PPMOs (SEQ ID NOS: 428, 429 and 431) were identified as effective in inducing exon-skipping and selected for additional evaluation. Dose-ranging experiments in RD cells were used to confirm the relative efficacy of these three PMO sequences. SEQ ID NO: 429 was shown to be most effective at inducing exon 53 skipping as shown in FIGS. 4B-F. However, when compared to other exon 53 antisense sequences, SEQ ID NO: 429 proved identical to H53A(+23+47) which is listed as SEQ ID NO: 195 in WO2006/000057 and SEQ ID NO: 609 in the present application. Other sequences were compared to SEQ ID NO: 429 including H53A(+39+69) and H53A(-12+10) (listed as SEQ ID NOs:193 and 199 in WO2006/000057, respectively) and h53AON1 (listed as SEQ ID NO:39 in U.S. application Ser. No. 11/233,507) and listed as SEQ ID NOs: 608, 611 and 610, respectively, in the present application. All the evaluated sequences were made as peptide-conjugated PMOs using the CP06062 peptide (SEQ ID NO: 578). This allowed direct comparison of the relative effectiveness of the antisense sequences without regard to antisense chemistry or cell delivery. As shown in FIGS. 4I and 4G-H, SEQ ID NO: 429 was shown to be superior to each of these four sequences.
Example 4
Exon 44 Scan
[0288] A series of overlapping antisense PPMOs that target human dystrophin exon 44 were designed and synthesized. For exon 44, a series of PPMOs, each 25 bases in length, were made (SEQ ID NOS:1-20) as shown in FIG. 5A. The PPMOs were evaluated for exon skipping efficacy by treating RD cells at various concentrations as described above in the Materials and Methods. Five PPMOs (SEQ ID NOS:4, 8, 11, 12 and 13) were identified as effective in inducing exon-skipping and selected for additional evaluation. Dose-ranging experiments in RD cells were used to confirm the relative efficacy of these five PPMO sequences as shown in FIGS. 5C to 5H. SEQ ID NOs: 8, 11 and 12 were shown to be most effective at inducing exon 44 skipping as shown in FIG. 5H with SEQ ID NO:12 proving the most efficacious.
[0289] Comparison of SEQ ID NO: 12 to other exon 44 antisense sequences was done in both RD cells and human primary skeletal muscle cells. All the evaluated sequences were made as peptide-conjugated PMOs using the CP06062 peptide (SEQ ID NO: 578). This allowed direct comparison of the relative effectiveness of the antisense sequences without regard to antisense chemistry or cell delivery.
[0290] The alignment of the sequences (SEQ ID NOS: 600, 601, 602 and 603) with SEQ ID NOS: 4, 8, 11 and 12 is shown in FIG. 5B. SEQ ID NOS: 601 and 603 are listed as SEQ ID NOS: 165 and 167 in WO2006/000057. SEQ ID NO:602 is listed in WO2004/083446 and as SEQ ID NO: 21 in U.S. application Ser. No. 11/233,507. SEQ ID NO:600 was published in 2007 (Wilton, Fall et al. 2007). The comparison in RD cells showed that both SEQ ID NOS: 602 and 603 were superior to SEQ ID NO:12 (FIG. 5I). However, as shown in FIG. 5J, in human primary skeletal muscle cells SEQ ID NO:12 was superior (8.86% exon skipping) to SEQ ID NO:602 (6.42%). Similar experiments are performed with SEQ ID NO:603.
Example 5
Exon 45 Scan
[0291] A series of overlapping antisense PPMOs that target human dystrophin exon 45 were designed and synthesized. For exon 45, a series of 22 PPMOs, each 25 bases in length, were made (SEQ ID NOS: 21, 23, 25, 27, 29, 31, 32, 34, 35, 37, 39, 41, 43 and 45-53) as shown in FIG. 6A. The PPMOs were evaluated for exon skipping efficacy by treating RD cells and human primary skeletal muscle cells at various concentrations as described above in the Materials and Methods. Five PPMOs (SEQ ID NOS:27, 29, 34, and 39) were identified as effective in inducing exon-skipping and selected for additional evaluation. Dose-ranging experiments in RD cells were used to confirm the relative efficacy of these four PMO sequences as shown in FIGS. 6C-G and summarized in FIG. 6H. SEQ ID NO: 49 was used as a negative control in these experiments. SEQ ID NOs: 29 and 34 were shown to be most effective at inducing exon 45 skipping as shown in FIG. 6H.
[0292] Comparison of SEQ ID NO: 34 to other exon 45 antisense sequences was done in both RD cells and human primary skeletal muscle cells. All the evaluated sequences were made as peptide-conjugated PMOs using the CP06062 peptide (SEQ ID NO: 578). This allowed direct comparison of the relative effectiveness of the antisense sequences without regard to antisense chemistry or cell delivery. The alignment of the sequences (SEQ ID NOS: 604, 605, 606 and 607) with SEQ ID NOS: 27, 29, 34 and 39 is shown in FIG. 6B. SEQ ID NOS: 604 and 607 are listed as SEQ ID NOS: 211 and 207 in WO2006/000057, respectively. SEQ ID NOS:605 and 606 are listed in U.S. application Ser. No. 11/233,507 as SEQ ID NOS: 23 and 1, respectively. The comparison in RD cells showed that SEQ ID NO: 34 was superior to all four sequences evaluated as shown in FIG. 6I. Testing of these compounds in different populations of human primary skeletal muscle cells is performed as described above.
SEQUENCE ID LISTING
[0293] Sequences are shown using the nucleotide base symbols common for DNA: A, G, C and T. Other antisense chemistries such as 2'-O-methyl use U in place of T. Any of the bases may be substituted with inosine (I) especially in stretches of three or more G residues.
TABLE-US-00008 Name Sequences SEQ ID NO. Oligomer Targeting Sequences (5' to 3'); Hu.DMD.Exon44.25.001 CTGCAGGTAAAAGCATATGGATCAA 1 Hu.DMD.Exon44.25.002 ATCGCCTGCAGGTAAAAGCATATGG 2 Hu.DMD.Exon44.25.003 GTCAAATCGCCTGCAGGTAAAAGCA 3 Hu.DMD.Exon44.25.004 GATCTGTCAAATCGCCTGCAGGTAA 4 Hu.DMD.Exon44.25.005 CAACAGATCTGTCAAATCGCCTGCA 5 Hu.DMD.Exon44.25.006 TTTCTCAACAGATCTGTCAAATCGC 6 Hu.DMD.Exon44.25.007 CCATTTCTCAACAGATCTGTCAAAT 7 Hu.DMD.Exon44.25.008 ATAATGAAAACGCCGCCATTTCTCA 8 Hu.DMD.Exon44.25.009 AAATATCTTTATATCATAATGAAAA 9 Hu.DMD.Exon44.25.010 TGTTAGCCACTGATTAAATATCTTT 10 Hu.DMD.Exon44.25.011 AAACTGTTCAGCTTCTGTTAGCCAC 11 Hu.DMD.Exon44.25.012 TTGTGTCTTTCTGAGAAACTGTTCA 12 Hu.DMD.Exon44.25.013 CCAATTCTCAGGAATTTGTGTCTTT 13 Hu.DMD.Exon44.25.014 GTATTTAGCATGTTCCCAATTCTCA 14 Hu.DMD.Exon44.25.015 CTTAAGATACCATTTGTATTTAGCA 15 Hu.DMD.Exon44.25.016 CTTACCTTAAGATACCATTTGTATT 16 Hu.DMD.Exon44.25.017 AAAGACTTACCTTAAGATACCATTT 17 Hu.DMD.Exon44.25.018 AAATCAAAGACTTACCTTAAGATAC 18 Hu.DMD.Exon44.25.019 AAAACAAATCAAAGACTTACCTTAA 19 Hu.DMD.Exon44.25.020 TCGAAAAAACAAATCAAAGACTTAC 20 Hu.DMD.Exon45.25.001 CTGTAAGATACCAAAAAGGCAAAAC 21 Hu.DMD.Exon45.25.002 CCTGTAAGATACCAAAAAGGCAAAA 22 Hu.DMD.Exon45.25.002. AGTTCCTGTAAGATACCAAAAAGGC 23 2 Hu.DMD.Exon45.25.003 GAGTTCCTGTAAGATACCAAAAAGG 24 Hu.DMD.Exon45.25.003. CCTGGAGTTCCTGTAAGATACCAAA 25 2 Hu.DMD.Exon45.25.004 TCCTGGAGTTCCTGTAAGATACCAA 26 Hu.DMD.Exon45.25.004. GCCATCCTGGAGTTCCTGTAAGATA 27 2 Hu.DMD.Exon45.25.005 TGCCATCCTGGAGTTCCTGTAAGAT 28 Hu.DMD.Exon45.25.005. CCAATGCCATCCTGGAGTTCCTGTA 29 2 Hu.DMD.Exon45.25.006 CCCAATGCCATCCTGGAGTTCCTGT 30 Hu.DMD.Exon45.25.006. GCTGCCCAATGCCATCCTGGAGTTC 31 2 Hu.DMD.Exon45.25.007 CGCTGCCCAATGCCATCCTGGAGTT 32 Hu.DMD.Exon45.25.008 AACAGTTTGCCGCTGCCCAATGCCA 33 Hu.DMD.Exon45.25.008. CTGACAACAGTTTGCCGCTGCCCAA 34 2 Hu.DMD.Exon45.25.009 GTTGCATTCAATGTTCTGACAACAG 35 Hu.DMD.Exon45.25.010 GCTGAATTATTTCTTCCCCAGTTGC 36 Hu.DMD.Exon45.25.010. ATTATTTCTTCCCCAGTTGCATTCA 37 2 Hu.DMD.Exon45.25.011 GGCATCTGTTTTTGAGGATTGCTGA 38 Hu.DMD.Exon45.25.011. TTTGAGGATTGCTGAATTATTTCTT 39 2 Hu.DMD.Exon45.25.012 AATTTTTCCTGTAGAATACTGGCAT 40 Hu.DMD.Exon45.25.012. ATACTGGCATCTGTTTTTGAGGATT 41 2 Hu.DMD.Exon45.25.013 ACCGCAGATTCAGGCTTCCCAATTT 42 Hu.DMD.Exon45.25.013. AATTTTTCCTGTAGAATACTGGCAT 43 2 Hu.DMD.Exon45.25.014 CTGTTTGCAGACCTCCTGCCACCGC 44 Hu.DMD.Exon45.25.014. AGATTCAGGCTTCCCAATTTTTCCT 45 2 Hu.DMD.Exon45.25.015 CTCTTTTTTCTGTCTGACAGCTGTT 46 Hu.DMD.Exon45.25.015. ACCTCCTGCCACCGCAGATTCAGGC 47 2 Hu.DMD.Exon45.25.016 CCTACCTCTTTTTTCTGTCTGACAG 48 Hu.DMD.Exon45.25.016. GACAGCTGTTTGCAGACCTCCTGCC 49 2 Hu.DMD.Exon45.25.017 GTCGCCCTACCTCTTTTTTCTGTCT 50 Hu.DMD.Exon45.25.018 GATCTGTCGCCCTACCTCTTTTTTC 51 Hu.DMD.Exon45.25.019 TATTAGATCTGTCGCCCTACCTCTT 52 Hu.DMD.Exon45.25.020 ATTCCTATTAGATCTGTCGCCCTAC 53 Hu.DMD.Exon45.20.001 AGATACCAAAAAGGCAAAAC 54 Hu.DMD.Exon45.20.002 AAGATACCAAAAAGGCAAAA 55 Hu.DMD.Exon45.20.003 CCTGTAAGATACCAAAAAGG 56 Hu.DMD.Exon45.20.004 GAGTTCCTGTAAGATACCAA 57 Hu.DMD.Exon45.20.005 TCCTGGAGTTCCTGTAAGAT 58 Hu.DMD.Exon45.20.006 TGCCATCCTGGAGTTCCTGT 59 Hu.DMD.Exon45.20.007 CCCAATGCCATCCTGGAGTT 60 Hu.DMD.Exon45.20.008 CGCTGCCCAATGCCATCCTG 61 Hu.DMD.Exon45.20.009 CTGACAACAGTTTGCCGCTG 62 Hu.DMD.Exon45.20.010 GTTGCATTCAATGTTCTGAC 63 Hu.DMD.Exon45.20.011 ATTATTTCTTCCCCAGTTGC 64 Hu.DMD.Exon45.20.012 TTTGAGGATTGCTGAATTAT 65 Hu.DMD.Exon45.20.013 ATACTGGCATCTGTTTTTGA 66 Hu.DMD.Exon45.20.014 AATTTTTCCTGTAGAATACT 67 Hu.DMD.Exon45.20.015 AGATTCAGGCTTCCCAATTT 68 Hu.DMD.Exon45.20.016 ACCTCCTGCCACCGCAGATT 69 Hu.DMD.Exon45.20.017 GACAGCTGTTTGCAGACCTC 70 Hu.DMD.Exon45.20.018 CTCTTTTTTCTGTCTGACAG 71 Hu.DMD.Exon45.20.019 CCTACCTCTTTTTTCTGTCT 72 Hu.DMD.Exon45.20.020 GTCGCCCTACCTCTTTTTTC 73 Hu.DMD.Exon45.20.021 GATCTGTCGCCCTACCTCTT 74 Hu.DMD.Exon45.20.022 TATTAGATCTGTCGCCCTAC 75 Hu.DMD.Exon45.20.023 ATTCCTATTAGATCTGTCGC 76 Hu.DMD.Exon46.25.001 GGGGGATTTGAGAAAATAAAATTAC 77 Hu.DMD.Exon46.25.002 ATTTGAGAAAATAAAATTACCTTGA 78 Hu.DMD.Exon46.25.002. CTAGCCTGGAGAAAGAAGAATAAAA 79 2 Hu.DMD.Exon46.25.003 AGAAAATAAAATTACCTTGACTTGC 80 Hu.DMD.Exon46.25.003. TTCTTCTAGCCTGGAGAAAGAAGAA 81 2 Hu.DMD.Exon46.25.004 ATAAAATTACCTTGACTTGCTCAAG 82 Hu.DMD.Exon46.25.004. TTTTGTTCTTCTAGCCTGGAGAAAG 83 2 Hu.DMD.Exon46.25.005 ATTACCTTGACTTGCTCAAGCTTTT 84 Hu.DMD.Exon46.25.005. TATTCTTTTGTTCTTCTAGCCTGGA 85 2 Hu.DMD.Exon46.25.006 CTTGACTTGCTCAAGCTTTTCTTTT 86 Hu.DMD.Exon46.25.006. CAAGATATTCTTTTGTTCTTCTAGC 87 2 Hu.DMD.Exon46.25.007 CTTTTAGTTGCTGCTCTTTTCCAGG 88 Hu.DMD.Exon46 25.008 CCAGGTTCAAGTGGGATACTAGCAA 89 Hu.DMD.Exon46.25.008. ATCTCTTTGAAATTCTGACAAGATA 90 2 Hu.DMD.Exon46.25.009 AGCAATGTTATCTGCTTCCTCCAAC 91 Hu.DMD.Exon46.25.009. AACAAATTCATTTAAATCTCTTTGA 92 2 Hu.DMD.Exon46.25.010 CCAACCATAAAACAAATTCATTTAA 93 Hu.DMD.Exon46.25.010. TTCCTCCAACCATAAAACAAATTCA 94 2 Hu.DMD.Exon46.25.011 TTTAAATCTCTTTGAAATTCTGACA 95 Hu.DMD.Exon46.25.012 TGACAAGATATTCTTTTGTTCTTCT 96 Hu.DMD.Exon46.25.012. TTCAAGTGGGATACTAGCAATGTTA 97 2 Hu.DMD.Exon46.25.013 AGATATTCTTTTGTTCTTCTAGCCT 98 Hu.DMD.Exon46.25.013. CTGCTCTTTTCCAGGTTCAAGTGGG 99 2 Hu.DMD.Exon46.25.014 TTCTTTTGTTCTTCTAGCCTGGAGA 100 Hu.DMD.Exon46.25.014. CTTTTCTTTTAGTTGCTGCTCTTTT 101 2 Hu.DMD.Exon46.25.015 TTGTTCTTCTAGCCTGGAGAAAGAA 102 Hu.DMD.Exon46.25.016 CTTCTAGCCTGGAGAAAGAAGAATA 103 Hu.DMD.Exon46.25.017 AGCCTGGAGAAAGAAGAATAAAATT 104 Hu.DMD.Exon46.25.018 CTGGAGAAAGAAGAATAAAATTGTT 105 Hu.DMD.Exon46.20.001 GAAAGAAGAATAAAATTGTT 106 Hu.DMD.Exon46.20.002 GGAGAAAGAAGAATAAAATT 107 Hu.DMD.Exon46.20.003 AGCCTGGAGAAAGAAGAATA 108 Hu.DMD.Exon46.20.004 CTTCTAGCCTGGAGAAAGAA 109 Hu.DMD.Exon46.20.005 TTGTTCTTCTAGCCTGGAGA 110 Hu.DMD.Exon46.20.006 TTCTTTTGTTCTTCTAGCCT 111
Hu.DMD.Exon46.20.007 TGACAAGATATTCTTTTGTT 112 Hu.DMD.Exon46.20.008 ATCTCTTTGAAATTCTGACA 113 Hu.DMD.Exon46.20.009 AACAAATTCATTTAAATCTC 114 Hu.DMD.Exon46.20.010 TTCCTCCAACCATAAAACAA 115 Hu.DMD.Exon46.20.011 AGCAATGTTATCTGCTTCCT 116 Hu.DMD.Exon46.20.012 TTCAAGTGGGATACTAGCAA 117 Hu.DMD.Exon46.20.013 CTGCTCTTTTCCAGGTTCAA 118 Hu.DMD.Exon46.20.014 CTTTTCTTTTAGTTGCTGCT 119 Hu.DMD.Exon46.20.015 CTTGACTTGCTCAAGCTTTT 120 Hu.DMD.Exon46.20.016 ATTACCTTGACTTGCTCAAG 121 Hu.DMD.Exon46.20.017 ATAAAATTACCTTGACTTGC 122 Hu.DMD.Exon46.20.018 AGAAAATAAAATTACCTTGA 123 Hu.DMD.Exon46.20.019 ATTTGAGAAAATAAAATTAC 124 Hu.DMD.Exon46.20.020 GGGGGATTTGAGAAAATAAA 125 Hu.DMD.Exon47.25.001 CTGAAACAGACAAATGCAACAACGT 126 Hu.DMD.Exon47.25.002 AGTAACTGAAACAGACAAATGCAAC 127 Hu.DMD.Exon47.25.003 CCACCAGTAACTGAAACAGACAAAT 128 Hu.DMD.Exon47.25.004 CTCTTCCACCAGTAACTGAAACAGA 129 Hu.DMD.Exon47.25.005 GGCAACTCTTCCACCAGTAACTGAA 130 Hu.DMD.Exon47.25.006 GCAGGGGCAACTCTTCCACCAGTAA 131 Hu.DMD.Exon47.25.007 CTGGCGCAGGGGCAACTCTTCCACC 132 Hu.DMD.Exon47.25.008 TTTAATTGTTTGAGAATTCCCTGGC 133 Hu.DMD.Exon47.25.008. TTGTTTGAGAATTCCCTGGCGCAGG 134 2 Hu.DMD.Exon47.25.009 GCACGGGTCCTCCAGTTTCATTTAA 135 Hu.DMD.Exon47.25.009. TCCAGTTTCATTTAATTGTTTGAGA 136 2 Hu.DMD.Exon47.25.010 GCTTATGGGAGCACTTACAAGCACG 137 Hu.DMD.Exon47.25.010. TACAAGCACGGGTCCTCCAGTTTCA 138 2 Hu.DMD.Exon47.25.011 AGTTTATCTTGCTCTTCTGGGCTTA 139 Hu.DMD.Exon47.25.012 TCTGCTTGAGCTTATTTTCAAGTTT 140 Hu.DMD.Exon47.25.012. ATCTTGCTCTTCTGGGCTTATGGGA 141 2 Hu.DMD.Exon47.25.013 CTTTATCCACTGGAGATTTGTCTGC 142 Hu.DMD.Exon47.25.013. CTTATTTTCAAGTTTATCTTGCTCT 143 2 Hu.DMD.Exon47.25.014 CTAACCTTTATCCACTGGAGATTTG 144 Hu.DMD.Exon47.25.014. ATTTGTCTGCTTGAGCTTATTTTCA 145 2 Hu.DMD.Exon47.25.015 AATGTCTAACCTTTATCCACTGGAG 146 Hu.DMD.Exon47.25.016 TGGTTAATGTCTAACCTTTATCCAC 147 Hu.DMD.Exon47.25.017 AGAGATGGTTAATGTCTAACCTTTA 148 Hu.DMD.Exon47.25.018 ACGGAAGAGATGGTTAATGTCTAAC 149 Hu.DMD.Exon47.20.001 ACAGACAAATGCAACAACGT 150 Hu.DMD.Exon47.20.002 CTGAAACAGACAAATGCAAC 151 Hu.DMD.Exon47.20.003 AGTAACTGAAACAGACAAAT 152 Hu.DMD.Exon47.20.004 CCACCAGTAACTGAAACAGA 153 Hu.DMD.Exon47.20.005 CTCTTCCACCAGTAACTGAA 154 Hu.DMD.Exon47.20.006 GGCAACTCTTCCACCAGTAA 155 Hu.DMD.Exon47.20.007 CTGGCGCAGGGGCAACTCTT 156 Hu.DMD.Exon47.20.008 TTGTTTGAGAATTCCCTGGC 157 Hu.DMD.Exon47.20.009 TCCAGTTTCATTTAATTGTT 158 Hu.DMD.Exon47.20.010 TACAAGCACGGGTCCTCCAG 159 Hu.DMD.Exon47.20.011 GCTTATGGGAGCACTTACAA 160 Hu.DMD.Exon47.20.012 ATCTTGCTCTTCTGGGCTTA 161 Hu.DMD.Exon47.20.013 CTTATTTTCAAGTTTATCTT 162 Hu.DMD.Exon47.20.014 ATTTGTCTGCTTGAGCTTAT 163 Hu.DMD.Exon47.20.015 CTTTATCCACTGGAGATTTG 164 Hu.DMD.Exon47.20.016 CTAACCTTTATCCACTGGAG 165 Hu.DMD.Exon47.20.017 AATGTCTAACCTTTATCCAC 166 Hu.DMD.Exon47.20.018 TGGTTAATGTCTAACCTTTA 167 Hu.DMD.Exon47.20.019 AGAGATGGTTAATGTCTAAC 168 Hu.DMD.Exon47.20.020 ACGGAAGAGATGGTTAATGT 169 Hu.DMD.Exon48.25.001 CTGAAAGGAAAATACATTTTAAAAA 170 Hu.DMD.Exon48.25.002 CCTGAAAGGAAAATACATTTTAAAA 171 Hu.DMD.Exon48.25.002. GAAACCTGAAAGGAAAATACATTTT 172 2 Hu.DMD.Exon48.25.003 GGAAACCTGAAAGGAAAATACATTT 173 Hu.DMD.Exon48.25.003. CTCTGGAAACCTGAAAGGAAAATAC 174 2 Hu.DMD.Exon48.25.004 GCTCTGGAAACCTGAAAGGAAAATA 175 Hu.DMD.Exon48.25.004. TAAAGCTCTGGAAACCTGAAAGGAA 634 2 Hu.DMD.Exon48.25.005 GTAAAGCTCTGGAAACCTGAAAGGA 176 Hu.DMD.Exon48 25.005. TCAGGTAAAGCTCTGGAAACCTGAA 177 2 Hu.DMD.Exon48.25.006 CTCAGGTAAAGCTCTGGAAACCTGA 178 Hu.DMD.Exon48.25.006. GTTTCTCAGGTAAAGCTCTGGAAAC 179 2 Hu.DMD.Exon48.25.007 TGTTTCTCAGGTAAAGCTCTGGAAA 180 Hu.DMD.Exon48.25.007. AATTTCTCCTTGTTTCTCAGGTAAA 181 2 Hu.DMD.Exon48.25.008 TTTGAGCTTCAATTTCTCCTTGTTT 182 Hu.DMD.Exon48.25.008 TTTTATTTGAGCTTCAATTTCTCCT 183 Hu.DMD.Exon48.25.009 AAGCTGCCCAAGGTCTTTTATTTGA 184 Hu.DMD.Exon48.25.010 AGGTCTTCAAGCTTTTTTTCAAGCT 185 Hu.DMD.Exon48.25.010. TTCAAGCTTTTTTTCAAGCTGCCCA 186 2 Hu.DMD.Exon48.25.011 GATGATTTAACTGCTCTTCAAGGTG 187 Hu.DMD.Exon48.25.011. CTGCTCTTCAAGGTCTTCAAGCTTT 188 2 Hu.DMD.Exon48.25.012 AGGAGATAACCACAGCAGCAGATGA 189 Hu.DMD.Exon48.25.012. CAGCAGATGATTTAACTGCTCTTCA 190 2 Hu.DMD.Exon48.25.013 ATTTCCAACTGATTCCTAATAGGAG 191 Hu.DMD.Exon48.25.014 CTTGGTTTGGTTGGTTATAAATTTC 192 Hu.DMD.Exon48.25.014. CAACTGATTCCTAATAGGAGATAAC 193 2 Hu.DMD.Exon48.25.015 CTTAACGTCAAATGGTCCTTCTTGG 194 Hu.DMD.Exon48.25.015. TTGGTTATAAATTTCCAACTGATTC 195 2 Hu.DMD.Exon48.25.016 CCTACCTTAACGTCAAATGGTCCTT 196 Hu.DMD.Exon48.25.016. TCCTTCTTGGTTTGGTTGGTTATAA 197 2 Hu.DMD.Exon48.25.017 AGTTCCCTACCTTAACGTCAAATGG 198 Hu.DMD.Exon48.25.018 CAAAAAGTTCCCTACCTTAACGTCA 199 Hu.DMD.Exon48.25.019 TAAAGCAAAAAGTTCCCTACCTTAA 200 Hu.DMD.Exon48.25.020 ATATTTAAAGCAAAAAGTTCCCTAC 201 Hu.DMD.Exon48.20.001 AGGAAAATACATTTTAAAAA 202 Hu.DMD.Exon48.20.002 AAGGAAAATACATTTTAAAA 203 Hu.DMD.Exon48.20.003 CCTGAAAGGAAAATACATTT 204 Hu.DMD.Exon48.20.004 GGAAACCTGAAAGGAAAATA 205 Hu.DMD.Exon48.20.005 GCTCTGGAAACCTGAAAGGA 206 Hu.DMD.Exon48.20.006 GTAAAGCTCTGGAAACCTGA 207 Hu.DMD.Exon48.20.007 CTCAGGTAAAGCTCTGGAAA 208 Hu.DMD.Exon48.20.008 AATTTCTCCTTGTTTCTCAG 209 Hu.DMD.Exon48.20.009 TTTTATTTGAGCTTCAATTT 210 Hu.DMD.Exon48.20.010 AAGCTGCCCAAGGTCTTTTA 211 Hu.DMD.Exon48.20.011 TTCAAGCTTTTTTTCAAGCT 212 Hu.DMD.Exon48.20.012 CTGCTCTTCAAGGTCTTCAA 213 Hu.DMD.Exon48.20.013 CAGCAGATGATTTAACTGCT 214 Hu.DMD.Exon48.20.014 AGGAGATAACCACAGCAGCA 215 Hu.DMD.Exon48.20.015 CAACTGATTCCTAATAGGAG 216 Hu.DMD.Exon48.20.016 TTGGTTATAAATTTCCAACT 217 Hu.DMD.Exon48.20.017 TCCTTCTTGGTTTGGTTGGT 218 Hu.DMD.Exon48.20.018 CTTAACGTCAAATGGTCCTT 219 Hu.DMD.Exon48.20.019 CCTACCTTAACGTCAAATGG 220 Hu.DMD.Exon48.20.020 AGTTCCCTACCTTAACGTCA 221 Hu.DMD.Exon48.20.021 CAAAAAGTTCCCTACCTTAA 222 Hu.DMD.Exon48.20.022 TAAAGCAAAAAGTTCCCTAC 223 Hu.DMD.Exon48.20.023 ATATTTAAAGCAAAAAGTTC 224 Hu.DMD.Exon49.25.001 CTGGGGAAAAGAACCCATATAGTGC 225 Hu.DMD.Exon49.25.002 TCCTGGGGAAAAGAACCCATATAGT 226
Hu.DMD.Exon49.25.002. GTTTCCTGGGGAAAAGAACCCATAT 227 2 Hu.DMD.Exon49.25.003 CAGTTTCCTGGGGAAAAGAACCCAT 228 Hu.DMD.Exon49.25.003. TTTCAGTTTCCTGGGGAAAAGAACC 229 2 Hu.DMD.Exon49.25.004 TATTTCAGTTTCCTGGGGAAAAGAA 230 Hu.DMD.Exon49.25.004. TGCTATTTCAGTTTCCTGGGGAAAA 231 2 Hu.DMD.Exon49.25.005 ACTGCTATTTCAGTTTCCTGGGGAA 232 Hu.DMD.Exon49.25.005. TGAACTGCTATTTCAGTTTCCTGGG 233 2 Hu.DMD.Exon49.25.006 CTTGAACTGCTATTTCAGTTTCCTG 234 Hu.DMD.Exon49 25.006. TAGCTTGAACTGCTATTTCAGTTTC 235 2 Hu.DMD.Exon49.25.007 TTTAGCTTGAACTGCTATTTCAGTT 236 Hu.DMD.Exon49.25.008 TTCCACATCCGGTTGTTTAGCTTGA 237 Hu.DMD.Exon49.25.009 TGCCCTTTAGACAAAATCTCTTCCA 238 Hu.DMD.Exon49.25.009. TTTAGACAAAATCTCTTCCACATCC 239 2 Hu.DMD.Exon49.25.010 GTTTTTCCTTGTACAAATGCTGCCC 240 Hu.DMD.Exon49.25.010 GTACAAATGCTGCCCTTTAGACAAA 241 2 Hu.DMD.Exon49.25.011 CTTCACTGGCTGAGTGGCTGGTTTT 242 Hu.DMD.Exon49.25.011. GGCTGGTTTTTCCTTGTACAAATGC 243 2 Hu.DMD.Exon49.25.012 ATTACCTTCACTGGCTGAGTGGCTG 244 Hu.DMD.Exon49.25.013 GCTTCATTACCTTCACTGGCTGAGT 245 Hu.DMD.Exon49.25.014 AGGTTGCTTCATTACCTTCACTGGC 246 Hu.DMD.Exon49.25.015 GCTAGAGGTTGCTTCATTACCTTCA 247 Hu.DMD.Exon49.25.016 ATATTGCTAGAGGTTGCTTCATTAC 248 Hu.DMD.Exon49.20.001 GAAAAGAACCCATATAGTGC 249 Hu.DMD.Exon49.20.002 GGGAAAAGAACCCATATAGT 250 Hu.DMD.Exon49.20.003 TCCTGGGGAAAAGAACCCAT 251 Hu.DMD.Exon49.20.004 CAGTTTCCTGGGGAAAAGAA 252 Hu.DMD.Exon49.20.005 TATTTCAGTTTCCTGGGGAA 253 Hu.DMD.Exon49.20.006 ACTGCTATTTCAGTTTCCTG 254 Hu.DMD.Exon49.20.007 CTTGAACTGCTATTTCAGTT 255 Hu.DMD.Exon49.20.008 TTTAGCTTGAACTGCTATTT 256 Hu.DMD.Exon49.20.009 TTCCACATCCGGTTGTTTAG 257 Hu.DMD.Exon49.20.010 TTTAGACAAAATCTCTTCCA 258 Hu.DMD.Exon49.20.011 GTACAAATGCTGCCCTTTAG 259 Hu.DMD.Exon49.20.012 GGCTGGTTTTTCCTTGTACA 260 Hu.DMD.Exon49.20.013 CTTCACTGGCTGAGTGGCTG 261 Hu.DMD.Exon49.20.014 ATTACCTTCACTGGCTGAGT 262 Hu.DMD.Exon49.20.015 GCTTCATTACCTTCACTGGC 263 Hu.DMD.Exon49.20.016 AGGTTGCTTCATTACCTTCA 264 Hu.DMD.Exon49.20.017 GCTAGAGGTTGCTTCATTAC 265 Hu.DMD.Exon49.20.018 ATATTGCTAGAGGTTGCTTC 266 Hu.DMD.Exon50.25.001 CTTTAACAGAAAAGCATACACATTA 267 Hu.DMD.Exon50.25.002 TCCTCTTTAACAGAAAAGCATACAC 268 Hu.DMD.Exon50.25.002. TTCCTCTTTAACAGAAAAGCATACA 269 2 Hu.DMD.Exon50.25.003 TAACTTCCTCTTTAACAGAAAAGCA 270 Hu.DMD.Exon50.25.003. CTAACTTCCTCTTTAACAGAAAAGC 271 2 Hu.DMD.Exon50.25.004 TCTTCTAACTTCCTCTTTAACAGAA 272 Hu.DMD.Exon50.25.004. ATCTTCTAACTTCCTCTTTAACAGA 273 2 Hu.DMD.Exon50.25.005 TCAGATCTTCTAACTTCCTCTTTAA 274 Hu.DMD.Exon50.25.005. CTCAGATCTTCTAACTTCCTCTTTA 275 2 Hu.DMD.Exon50.25.006 AGAGCTCAGATCTTCTAACTTCCTC 276 Hu.DMD.Exon50.25.006. CAGAGCTCAGATCTTCTAACTTCCT 277 2 NG-08-0731 Hu.DMD.Exon50.25.007 CACTCAGAGCTCAGATCTTCTACT 278 Hu.DMD.Exon50.25.007. CCTTCCACTCAGAGCTCAGATCTTC 279 2 Hu.DMD.Exon50.25.008 GTAAACGGTTTACCGCCTTCCACTC 280 Hu.DMD.Exon50.25.009 CTTTGCCCTCAGCTCTTGAAGTAAA 281 Hu.DMD.Exon50.25.009. CCCTCAGCTCTTGAAGTAAACGGTT 282 2 Hu.DMD.Exon50.25.010 CCAGGAGCTAGGTCAGGCTGCTTTG 283 Hu.DMD.Exon50.25.010. GGTCAGGCTGCTTTGCCCTCAGCTC 284 2 Hu.DMD.Exon50.25.011 AGGCTCCAATAGTGGTCAGTCCAGG 285 Hu.DMD.Exon50.25.011. TCAGTCCAGGAGCTAGGTCAGGCTG 286 2 Hu.DMD.Exon50.25.012 CTTACAGGCTCCAATAGTGGTCAGT 287 AVI-5038 Hu.DMD.Exon50.25.013 GTATACTTACAGGCTCCAATAGTGG 288 Hu.DMD.Exon50.25.014 ATCCAGTATACTTACAGGCTCCAAT 289 Hu.DMD.Exon50.25.015 ATGGGATCCAGTATACTTACAGGCT 290 NG-08-0741 Hu.DMD.Exon50.25.016 AGAGAATGGGATCCAGTATACTTAC 291 NG-08-0742 Hu.DMD.Exon50.20.001 ACAGAAAAGCATACACATTA 292 Hu.DMD.Exon50.20.002 TTTAACAGAAAAGCATACAC 293 Hu.DMD.Exon50.20.003 TCCTCTTTAACAGAAAAGCA 294 Hu.DMD.Exon50.20.004 TAACTTCCTCTTTAACAGAA 295 Hu.DMD.Exon50.20.005 TCTTCTAACTTCCTCTTTAA 296 Hu.DMD.Exon50.20.006 TCAGATCTTCTAACTTCCTC 297 Hu.DMD.Exon50.20.007 CCTTCCACTCAGAGCTCAGA 298 Hu.DMD.Exon50.20.008 GTAAACGGTTTACCGCCTTC 299 Hu.DMD.Exon50.20.009 CCCTCAGCTCTTGAAGTAAA 300 Hu.DMD.Exon50.20.010 GGTCAGGCTGCTTTGCCCTC 301 Hu.DMD.Exon50.20.011 TCAGTCCAGGAGCTAGGTCA 302 Hu.DMD.Exon50.20.012 AGGCTCCAATAGTGGTCAGT 303 Hu.DMD.Exon50.20.013 CTTACAGGCTCCAATAGTGG 304 Hu.DMD.Exon50.20.014 GTATACTTACAGGCTCCAAT 305 Hu.DMD.Exon50.20.015 ATCCAGTATACTTACAGGCT 306 Hu.DMD.Exon50.20.016 ATGGGATCCAGTATACTTAC 307 Hu.DMD.Exon50.20.017 AGAGAATGGGATCCAGTATA 308 Hu.DMD.Exon51.25.001- CTAAAATATTTTGGGTTTTTGCAAAA 309 44 Hu.DmD.Exon51.25.002- GCTAAAATATTTTGGGTTTTTGCAAA 310 45 Hu.DMD.Exon51.25.002. TAGGAGCTAAAATATTTTGGGTTTTT 311 2-46 Hu.DMD.Exon51.25.003 AGTAGGAGCTAAAATATTTTGGGTT 312 Hu.DMD.Exon51.25.003. TGAGTAGGAGCTAAAATATTTTGGG 313 2 Hu.DMD.Exon51.25.004 CTGAGTAGGAGCTAAAATATTTTGGG 314 Hu.DMD.Exon51.25.004. CAGTCTGAGTAGGAGCTAAAATATT 315 2 Hu.DMD.Exon51.25.005 ACAGTCTGAGTAGGAGCTAAAATATT 316 Hu.DMD.Exon51.25.005. GAGTAACAGTCTGAGTAGGAGCTAAA 317 2 Hu.DMD.Exon51.25.006 CAGAGTAACAGTCTGAGTAGGAGCT 318 Hu.DMD.Exon51.25.006. CACCAGAGTAACAGTCTGAGTAGGAG 319 2 Hu.DMD.Exon51.25.007 GTCACCAGAGTAACAGTCTGAGTAG 320 Hu.DMD.Exon51.25.007. AACCACAGGTTGTGTCACCAGAGTAA 321 2 Hu.DMD.Exon51.25.008 GTTGTGTCACCAGAGTAACAGTCTG 322 Hu.DMD.Exon51.25.009 TGGCAGTTTCCTTAGTAACCACAGGT 323 Hu.DMD.Exon51.25.010 ATTTCTAGTTTGGAGATGGCAGTTTC 324 Hu.DMD.Exon51.25.010. GGAAGATGGCATTTCTAGTTTGGAG 325 2 Hu.DMD.Exon51.25.011 CATCAAGGAAGATGGCATTTCTAGTT 326 Hu.DMD.Exon51.25.011. GAGCAGGTACCTCCAACATCAAGGAA 327 2 Hu.DMD.Exon51.25.012 ATCTGCCAGAGCAGGTACCTCCAAC 328 Hu.DMD.Exon51.25.013 AAGTTCTGTCCAAGCCCGGTTGAAAT 329 Hu.DMD.Exon51.25.013. CGGTTGAAATCTGCCAGAGCAGGTAC 330 2 Hu.DMD.Exon51.25.014 GAGAAAGCCAGTCGGTAAGTTCTGTC 331 Hu.DMD.Exon51.25.014. GTCGGTAAGTTCTGTCCAAGCCCGG 332 2 Hu.DMD.Exon51.25.015 ATAACTTGATCAAGCAGAGAAAGCCA 333 Hu.DMD.Exon51.25.015. AAGCAGAGAAAGCCAGTCGGTAAGT 334 2 Hu.DMD.Exon51.25.016 CACCCTCTGTGATTTTATAACTTGAT 335
Hu.DMD.Exon51.25.017 CAAGGTCACCCACCATCACCCTCTGT 336 Hu.DMD.Exon51.25.017. CATCACCCTCTGTGATTTTATAACT 337 2 Hu.DMD.Exon51.25.018 CTTCTGCTTGATGATCATCTCGTTGA 338 Hu.DMD.Exon51.25.019 CCTTCTGCTTGATGATCATCTCGTTG 339 Hu.DMD.Exon51.25.019. ATCTCGTTGATATCCTCAAGGTCACC 340 2 Hu.DMD.Exon51.25.020 TCATACCTTCTGCTTGATGATCATCT 341 Hu.DMD.Exon51.25.020. TCATTTTTTCTCATACCTTCTGCTTG 342 2 Hu.DMD.Exon51.25.021 TTTTCTCATACCTTCTGCTTGATGAT 343 Hu.DMD.Exon51.25.022 TTTTATCATTTTTTCTCATACCTTCT 344 Hu.DMD.Exon51.25.023 CCAACTTTTATCATTTTTTCTCATAC 345 Hu.DMD.Exon51.20.001 ATATTTTGGGTTTTTGCAAA 346 Hu.DMD.Exon51 20.002 AAAATATTTTGGGTTTTTGC 347 Hu.DMD.Exon51.20.003 GAGCTAAAATATTTTGGGTT 348 Hu.DMD.Exon51.20.004 AGTAGGAGCTAAAATATTTT 349 Hu.DMD.Exon51.20.005 GTCTGAGTAGGAGCTAAAAT 350 Hu.DMD.Exon51.20.006 TAACAGTCTGAGTAGGAGCT 351 Hu.DMD.Exon51.20.007 CAGAGTAACAGTCTGAGTAG 352 Hu.DMD.Exon51.20.008 CACAGGTTGTGTCACCAGAG 353 Hu.DMD.Exon51.20.009 AGTTTCCTTAGTAACCACAG 354 Hu.DMD.Exon51.20.010 TAGTTTGGAGATGGCAGTTT 355 Hu.DMD.Exon51.20.011 GGAAGATGGCATTTCTAGTT 356 Hu.DMD.Exon51.20.012 TACCTCCAACATCAAGGAAG 357 Hu.DMD.Exon51.20.013 ATCTGCCAGAGCAGGTACCT 358 Hu.DMD.Exon51.20.014 CCAAGCCCGGTTGAAATCTG 359 Hu.DMD.Exon51.20.015 GTCGGTAAGTTCTGTCCAAG 360 Hu.DMD.Exon51.20.016 AAGCAGAGAAAGCCAGTCGG 361 Hu.DMD.Exon51.20.017 TTTTATAACTTGATCAAGCA 362 Hu.DMD.Exon51.20.018 CATCACCCTCTGTGATTTTA 363 Hu DMD.Exon51.20.019 CTCAAGGTCACCCACCATCA 364 Hu.DMD.Exon51.20.020 CATCTCGTTGATATCCTCAA 365 Hu.DMD.Exon51.20.021 CTTCTGCTTGATGATCATCT 366 Hu.DMD.Exon51.20.022 CATACCTTCTGCTTGATGAT 367 Hu.DMD.Exon51.20.023 TTTCTCATACCTTCTGCTTG 368 Hu.DMD.Exon51.20.024 CATTTTTTCTCATACCTTCT 369 Hu.DMD.Exon51.20.025 TTTATCATTTTTTCTCATAC 370 Hu.DMD.Exon51.20.026 CAACTTTTATCATTTTTTCT 371 Hu.DMD.Exon52.25.001 CTGTAAGAACAAATATCCCTTAGTA 372 Hu.DMD.Exon52.25.002 TGCCTGTAAGAACAAATATCCCTTA 373 Hu.DMD.Exon52 25.002. GTTGCCTGTAAGAACAAATATCCCT 374 2 Hu.DMD.Exon52.25.003 ATTGTTGCCTGTAAGAACAAATATC 375 Hu.DMD.Exon52.25.003. GCATTGTTGCCTGTAAGAACAAATA 376 2 Hu.DMD.Exon52.25.004 CCTGCATTGTTGCCTGTAAGAACAA 377 Hu.DMD.Exon52.25.004. ATCCTGCATTGTTGCCTGTAAGAAC 378 2 Hu.DMD.Exon52.25.005 CAAATCCTGCATTGTTGCCTGTAAG 379 Hu.DMD.Exon52.25.005. TCCAAATCCTGCATTGTTGCCTGTA 380 2 Hu.DMD.Exon52.25.006 TGTTCCAAATCCTGCATTGTTGCCT 381 Hu.DMD.Exon52.25.006. TCTGTTCCAAATCCTGCATTGTTGC 382 2 Hu.DMD.Exon52.25.007 AACTGGGGACGCCTCTGTTCCAAAT 383 Hu.DMD.Exon52.25.007. GCCTCTGTTCCAAATCCTGCATTGT 384 2 Hu.DMD.Exon52.25.008 CAGCGGTAATGAGTTCTTCCAACTG 385 Hu.DMD.Exon52.25.008. CTTCCAACTGGGGACGCCTCTGTTC 386 2 Hu.DMD.Exon52.25.009 CTTGTTTTTCAAATTTTGGGCAGCG 387 Hu.DMD.Exon52.25.010 CTAGCCTCTTGATTGCTGGTCTTGT 388 Hu.DMD.Exon52.25.010. TTTTCAAATTTTGGGCAGCGGTAAT 389 2 Hu.DMD.Exon52.25.011 TTCGATCCGTAATGATTGTTCTAGC 390 Hu.DMD.Exon52.25.011. GATTGCTGGTCTTGTTTTTCAAATT 391 2 Hu.DMD.Exon52.25.012 CTTACTTCGATCCGTAATGATTGTT 392 Hu.DMD.Exon52.25.012. TTGTTCTAGCCTCTTGATTGCTGGT 393 2 Hu.DMD.Exon52.25.013 AAAAACTTACTTCGATCCGTAATGA 394 Hu.DMD.Exon52.25.014 TGTAAAAAACTTACTTCGATCCGT 395 Hu.DMD.Exon52.25.015 ATGCTTGTTAAAAAACTTACTTCGA 396 Hu.DMD.Exon52.25.016 GTCCCATGCTTGTTAAAAAACTTAC 397 Hu.DMD.Exon52.20.001 AGAACAAATATCCCTTAGTA 398 Hu.DMD.Exon52.20.002 GTAAGAACAAATATCCCTTA 399 Hu.DMD.Exon52.20.003 TGCCTGTAAGAACAAATATC 400 Hu.DMD.Exon52.20.004 ATTGTTGCCTGTAAGAACAA 401 Hu.DMD.Exon52.20.005 CCTGCATTGTTGCCTGTAAG 402 Hu.DMD.Exon52.20.006 CAAATCCTGCATTGTTGCCT 403 Hu.DMD.Exon52.20.007 GCCTCTGTTCCAAATCCTGC 404 Hu.DMD.Exon52.20.008 CTTCCAACTGGGGACGCCTC 405 Hu.DMD.Exon52.20.009 CAGCGGTAATGAGTTCTTCC 406 Hu.DMD.Exon52.20.010 TTTTCAAATTTTGGGCAGCG 407 Hu.DMD.Exon52.20.011 GATTGCTGGTCTTGTTTTTC 408 Hu.DMD.Exon52.20.012 TTGTTCTAGCCTCTTGATTG 409 Hu.DMD.Exon52.20.013 TTCGATCCGTAATGATTGTT 410 Hu.DMD.Exon52.20.014 CTTACTTCGATCCGTAATGA 411 Hu.DMD.Exon52.20.015 AAAAACTTACTTCGATCCGT 412 Hu.DMD.Exon52.20.016 TGTTAAAAAACTTACTTCGA 413 Hu.DMD.Exon52.20.017 ATGCTTGTTAAAAAACTTAC 414 Hu.DMD.Exon52.20.018 GTCCCATGCTTGTTAAAAAA 415 Hu.DMD.Exon53.25.001 CTAGAATAAAAGGAAAAATAAATAT 416 Hu.DMD.Exon53.25.002 AACTAGAATAAAAGGAAAAATAAAT 417 Hu.DMD.Exon53.25.002. TTCAACTAGAATAAAAGGAAAAATA 418 2 Hu.DMD.Exon53.25.003 CTTTCAACTAGAATAAAAGGAAAAA 419 Hu.DMD.Exon53.25.003. ATTCTTTCAACTAGAATAAAAGGAA 420 2 Hu.DMD.Exon53.25.004 GAATTCTTTCAACTAGAATAAAAGG 421 Hu.DMD.Exon53.25.004. TCTGAATTCTTTCAACTAGAATAAA 422 2 Hu.DMD.Exon53.25.005 ATTCTGAATTCTTTCAACTAGAATA 423 Hu.DMD.Exon53.25.005. CTGATTCTGAATTCTTTCAACTAGA 424 2 Hu.DMD.Exon53.25.006 CACTGATTCTGAATTCTTTCAACTA 425 Hu.DMD.Exon53.25.006. TCCCACTGATTCTGAAfTCTTTCAA 426 2 Hu.DMD.Exon53.25.007 CATCCCACTGATTCTGAATTCTTTC 427 Hu.DMD.Exon53.25.008 TACTTCATCCCACTGATTCTGAATT 428 Hu.DMD.Exon53.25.008. CTGAAGGTGTTCTTGTACTTCATCC 429 2 Hu.DMD.Exon53.25.009 CGGTTCTGAAGGTGTTCTTGTACT 430 Hu.DMD.Exon53.25.009. CTGTTGCCTCCGGTTCTGAAGGTGT 431 2 Hu.DMD.Exon53.25.010 TTTCATTCAACTGTTGCCTCCGGTT 432 Hu.DMD.Exon53.25.010. TAACATTTCATTCAACTGTTGCCTC 433 2 Hu.DMD.Exon53.25.011 TTGTGTTGAATCCTTTAACATTTCA 434 Hu.DMD.Exon53.25.012 TCTTCCTTAGCTTCCAGCCATTGTG 435 Hu.DMD.Exon53.25.012. CTTAGCTTCCAGCCATTGTGTTGAA 436 2 Hu.DMD.Exon53.25.013 GTCCTAAGACCTGCTCAGCTTCTTC 437 Hu.DMD.Exon53.25.013. CTGCTCAGCTTCTTCCTTAGCTTCC 438 2 Hu.DMD.Exon53.25.014 CTCAAGCTTGGCTCTGGCCTGTCCT 439 Hu.DMD.Exon53.25.014. GGCCTGTCCTAAGACCTGCTCAGCT 440 2 Hu.DMD.Exon53.25.015 TAGGGACCCTCCTTCCATGACTCAA 441 Hu.DMD.Exon53.25.016 TTTGGATTGCATCTACTGTATAGGG 442 Hu.DMD.Exon53.25.016. ACCCTCCTTCCATGACTCAAGCTTG 443 2 Hu.DMD.Exon53.25.017 CTTGGTTTCTGTGATTTTCTTTTGG 444 Hu.DMD.Exon53.25.017. ATCTACTGTATAGGGACCCTCCTTC 445 2 Hu.DMD.Exon53.25.018 CTAACCTTGGTTTCTGTGATTTTCT 446 Hu.DMD.Exon53.25.018. TTTCTTTTGGATTGCATCTACTGTA 447 2
Hu.DMD.Exon53.25.019 TGATACTAACCTTGGTTTCTGTGAT 448 Hu.DMD.Exon53.25.020 ATCTTTGATACTAACCTTGGTTTCT 449 Hu.DMD.Exon53.25.021 AAGGTATCTTTGATACTAACCTTGG 450 Hu.DMD.Exon53.25.022 TTAAAAAGGTATCTTTGATACTAAC 451 Hu.DMD.Exon53.20.001 ATAAAAGGAAAAATAAATAT 452 Hu.DMD.Exon53.20.002 GAATAAAAGGAAAAATAAAT 453 Hu.DMD.Exon53.20.003 AACTAGAATAAAAGGAAAAA 454 Hu.DMD.Exon53.20.004 CTTTCAACTAGAATAAAAGG 455 Hu.DMD.Exon53.20.005 GAATTCTTTCAACTAGAATA 456 Hu.DMD.Exon53.20.006 ATTCTGAATTCTTTCAACTA 457 Hu.DMD.Exon53.20.007 TACTTCATCCCACTGATTCT 458 Hu.DMD.Exon53.20.008 CTGAAGGTGTTCTTGTACT 459 Hu.DMD.Exon53.20.009 CTGTTGCCTCCGGTTCTGAA 460 Hu.DMD.Exon53.20.010 TAACATTTCATTCAACTGTT 461 Hu.DMD.Exon53.20.011 TTGTGTTGAATCCTTTAACA 462 Hu.DMD.Exon53.20.012 CTTAGCTTCCAGCCATTGTG 463 Hu.DMD.Exon53.20.013 CTGCTCAGCTTCTTCCTTAG 464 Hu.DMD.Exon53.20.014 GGCCTGTCCTAAGACCTGCT 465 Hu.DMD.Exon53.20.015 CTCAAGCTTGGCTCTGGCCT 466 Hu.DMD.Exon53.20.016 ACCCTCCTTCCATGACTCAA 467 Hu.DMD.Exon53.20.017 ATCTACTGTATAGGGACCCT 468 Hu.DMD.Exon53.20.018 TTTCTTTTGGATTGCATCTA 469 Hu.DMD.Exon53.20.019 CTTGGTTTCTGTGATTTTCT 470 Hu.DMD.Exon53.20.020 CTAACCTTGGTTTCTGTGAT 471 Hu.DMD.Exon53.20.021 TGATACTAACCTTGGTTTCT 472 Hu.DMD.Exon53.20.022 ATCTTTGATACTAACCTTGG 473 Hu.DMD.Exon53.20.023 AAGGTATCTTTGATACTAAC 474 Hu.DMD.Exon53.20.024 TTAAAAAGGTATCTTTGATA 475 Hu.DMD.Exon54.25.001 CTATAGATTTTTATGAGAAAGAGA 476 Hu.DMD.Exon54.25.002 AACTGCTATAGATTTTATGAGAAA 477 Hu.DMD.Exon54.25.003 TGGCCAACTGCTATAGATTTTTATG 478 Hu.DMD.Exon54.25.004 GTCTTTGGCCAACTGCTATAGATTT 479 Hu.DMD.Exon54.25.005 CGGAGGTCTTTGGCCAACTGCTATA 480 Hu.DMD.Exon54.25.006 ACTGGCGGAGGTCTTTGGCCAACTG 481 Hu.DMD.Exon54.25.007 TTTGTCTGCCACTGGCGGAGGTCTT 482 Hu.DMD.Exon54.25.008 AGTCATTTGCCACATCTACATTTGT 483 Hu.DMD.Exon54.25.008. TTTGCCACATCTACATTTGTCTGCC 484 2 Hu.DMD.Exon54.25.009 CCGGAGAAGTTTCAGGGCCAAGTCA 485 Hu.DMD.Exon54.25.010 GTATCATCTGCAGAATAATCCCGGA 486 Hu.DMD.Exon54.25.010. TAATCCCGGAGAAGTTTCAGGGCCA 487 2 Hu.DMD.Exon54.25.011 TTATCATGTGGACTTTTCTGGTATC 488 Hu.DMD.Exon54.25.012 AGAGGCATTGATATTCTCTGTTATC 489 Hu.DMD.Exon54.25.012. ATGTGGACTTTTCTGGTATCATCTG 490 2 Hu.DMD.Exon54.25.013 CTTTTATGAATGCTTCTCCAAGAGG 491 Hu.DMD.Exon54.25.013. ATATTCTCTGTTATCATGTGGACTT 492 2 Hu.DMD.Exon54.25.014 CATACCTTTTATGAATGCTTCTCCA 493 Hu.DMD.Exon54.25.014. CTCCAAGAGGCATTGATATTCTCTG 494 2 Hu.DMD.Exon54.25.015 TAATTCATACCTTTTATGAATGCTT 495 Hu.DMD.Exon54.25.015. CTTTTATGAATGCTTCTCCAAGAGG 496 2 Hu.DMD.Exon54.25.016 TAATGTAATTCATACCTTTTATGAA 497 Hu.DMD.Exon54.25.017 AGAAATAATGTAATTCATACCTTTT 498 Hu.DMD.Exon54.25.018 GTTTTAGAAATAATGTAATTCATAC 499 Hu.DMD.Exon54.20.001 GATTTTTATGAGAAAGAGA 500 Hu.DMD.Exon54.20.002 CTATAGATTTTTATGAGAAA 501 Hu.DMD.Exon54.20.003 AACTGCTATAGATTTTTATG 502 Hu.DMD.Exon54.20.004 TGGCCAACTGCTATAGATTT 503 Hu.DMD.Exon54.20.005 GTCTTTGGCCAACTGCTATA 504 Hu.DMD.Exon54.20.006 CGGAGGTCTTTGGCCAACTG 505 Hu.DMD.Exon54.20.007 TTTGTCTGCCACTGGCGGAG 506 Hu.DMD.Exon54.20.008 TTTGCCACATCTACATTTGT 507 Hu.DMD.Exon54.20.009 TTCAGGGCCAAGTCATTTGC 508 Hu.DMD.Exon54.20.010 TAATCCCGGAGAAGTTTCAG 509 Hu.DMD.Exon54.20.011 GTATCATCTGCAGAATAATC 510 Hu.DMD.Exon54.20.012 ATGTGGACTTTTCTGGTATC 511 Hu.DMD.Exon54.20.013 ATATTCTCTGTTATCATGTG 512 Hu.DMD.Exon54.20.014 CTCCAAGAGGCATTGATATT 513 Hu.DMD.Exon54.20.015 CTTTTATGAATGCTTCTCCA 514 Hu.DMD.Exon54.20.016 CATACCTTTTATGAATGCTT 515 Hu.DMD.Exon54.20.017 TAATTCATACCTTTTATGAA 516 Hu.DMD.Exon54.20.018 TAATGTAATTCATACCTTTT 517 Hu.DMD.Exon54.20.019 AGAAATAATGTAATTCATAC 518 Hu.DMD.Exon54.20.020 GTTTTAGAAATAATGTAATT 519 Hu.DMD.Exon55.25.001 CTGCAAAGGACCAAATGTTCAGATG 520 Hu.DMD.Exon55.25.002 TCACCCTGCAAAGGACCAAATGTTC 521 Hu.DMD.Exon55.25.003 CTCACTCACCCTGCAAAGGACCAAA 522 Hu.DMD.Exon55.25.004 TCTCGCTCACTCACCCTGCAAAGGA 523 Hu.DMD.Exon55.25.005 CAGCCTCTCGCTCACTCACCCTGCA 524 Hu.DMD.Exon55.25.006 CAAAGCAGCCTCTCGCTCACTCACC 525 Hu.DMD.Exon55.25.007 TCTTCCAAAGCAGCCTCTCGCTCAC 526 Hu.DMD.Exon55.25.007. TCTATGAGTTTCTTCCAAAGCAGCC 527 2 Hu.DMD.Exon55.25.008 GTTGCAGTAATCTATGAGTTTCTTC 528 Hu.DMD.Exon55.25.008. GAACTGTTGCAGTAATCTATGAGTT 529 2 Hu.DMD.Exon55.25.009 TTCCAGGTCCAGGGGGAACTGTTGC 530 Hu.DMD.Exon55.25.010 GTAAGCCAGGCAAGAAACTTTTCCA 531 Hu.DMD.Exon55.25.010. CCAGGCAAGAAACTTTTCCAGGTCC 532 2 Hu.DMD.Exon55.25.011 TGGCAGTTGTTTCAGCTTCTGTAAG 533 Hu.DMD.Exon55.25.011. TTCAGCTTCTGTAAGCCAGGCAAGA 635 2 Hu.DMD.Exon55.25.012 GGTAGCATCCTGTAGGACATTGGCA 534 Hu.DMD.Exon55.25.012. GACATTGGCAGTTGTTTCAGCTTCT 535 2 Hu.DMD.Exon55.25.013 TCTAGGAGCCTTTCCTTACGGGTAG 536 Hu.DMD.Exon55.25.014 CTTTTACTCCCTTGGAGTCTTCTAG 537 Hu.DMD.Exon55.25.014. GAGCCTTTCCTTACGGGTAGCATCC 538 2 Hu.DMD.Exon55.25.015 TTGCCATTGTTTCATCAGCTCTTTT 539 Hu.DMD.Exon55.25.015. CTTGGAGTCTTCTAGGAGCCTTTCC 540 2 Hu.DMD.Exon55.25 016 CTTACTTGCCATTGTTTCATCAGCT 541 Hu.DMD.Exon55.25.016. CAGCTCTTTTACTCCCTTGGAGTCT 542 2 Hu.DMD.Exon55.25.017 CCTGACTTACTTGCCATTGTTTCAT 543 Hu.DMD.Exon55.25.018 AAATGCCTGACTTACTTGCCATTGT 544 Hu.DMD.Exon55.25.019 AGCGGAAATGCCTGACTTACTTGCC 545 Hu.DMD.Exon55.25.020 GCTAAAGCGGAAATGCCTGACTTAC 546 Hu.DMD.Exon55.20.001 AAGGACCAAATGTTCAGATG 547 Hu.DMD.Exon55.20.002 CTGCAAAGGACCAAATGTTC 548 Hu.DMD.Exon55.20.003 TCACCCTGCAAAGGACCAAA 549 Hu.DMD.Exon55.20.004 CTCACTCACCCTGCAAAGGA 550 Hu.DMD.Exon55.20.005 TCTCGCTCACTCACCCTGCA 551 Hu.DMD.Exon55.20.006 CAGCCTCTCGCTCACTCACC 552 Hu.DMD.Exon55.20.007 CAAAGCAGCCTCTCGCTCAC 553 Hu.DMD.Exon55.20.008 TCTATGAGTTTCTTCCAAAG 554 Hu.DMD.Exon55.20.009 GAACTGTTGCAGTAATCTAT 555 Hu.DMD.Exon55.20.010 TTCCAGGTCCAGGGGGAACT 556 Hu.DMD.Exon55.20.011 CCAGGCAAGAAACTTTTCCA 557 Hu.DMD.Exon55.20.012 TTCAGCTTCTGTAAGCCAGG 558 Hu.DMD.Exon55.20.013 GACATTGGCAGTTGTTTCAG 559 Hu.DMD.Exon55.20.014 GGTAGCATCCTGTAGGACAT 560 Hu.DMD.Exon55.20.015 GAGCCTTTCCTTACGGGTAG 561 Hu.DMD.Exon55.20.016 CTTGGAGTCTTCTAGGAGCC 562 Hu.DMD.Exon55.20.017 CAGCTCTTTTACTCCCTTGG 563 Hu.DMD.Exon55.20.018 TTGCCATTGTTTCATCAGCT 564
Hu.DMD.Exon55.20.019 CTTACTTGCCATTGTTTCAT 565 Hu.DMD.Exon55.20.020 CCTGACTTACTTGCCATTGT 566 Hu.DMD.Exon55.20.021 AAATGCCTGACTTACTTGCC 567 Hu.DMD.Exon55.20.022 AGCGGAAATGCCTGACTTAC 568 Hu.DMD.Exon55.20.023 GCTAAAGCGGAAATGCCTGA 569 H50A(+02+30)-AVI-5656 CCACTCAGAGCTCAGATCTTCTAACTTC 584 C H50D(+07-18)-AVI-5915 GGGATCCAGTATACTTACAGGCTCC 585 H50A(+07+33) CTTCCACTCAGAGCTCAGATCTTCTAA 586 H51A(+61+90)-AVI-4657 ACATCAAGGAAGATGGCATTTCTAGTTT 587 GG H51A(+66+95)-AVI-4658 CTCCAACATCAAGGAAGATGGCATTTCT 588 AG H51A(+111+134) TTCTGTCCAAGCCCGGTTGAAATC 589 H51A(+175+195) CACCCACCATCACCCTCYGTG 590 H51A(+199+220) ATCATCTCGTTGATATCCTCAA 591 H51A(+66+90) ACATCAAGGAAGATGGCATTTCTAG 592 H51A(-01+25) ACCAGAGTAACAGTCTGAGTAGGAGC 593 h51AON1 TCAAGGAAGATGGCATTTCT 594 h51AON2 CCTCTGTGATTTTATAACTTGAT 595 H51D(+08-17) ATCATTTTTTCTCATACCTTCTGCT 596 H51D(+16-07) CTCATACCTTCTGCTTGATGATC 597 hAON#23 TGGCATTTCTAGTTTGG 598 hAON#24 CCAGAGCAGGTACCTCCAACATC 599 H44A(+61+84) TGTTCAGCTTCTGTTAGCCACTGA 600 H44A(+85+104 ) TTTGTGTCTTTCTGAGAAAC 601 h44AON1 CGCCGCCATTTCTCAACAG 602 H44A(-06+14) ATCTGTCAAATCGCCTGCAG 603 H45A(+71+90) TGTTTTTGAGGATTGCTGAA 604 h45AON1 GCTGAATTATTTCTTCCCC 605 h45AON5 GCCCAATGCCATCCTGG 606 H45A(-06+20) CCAATGCCATCCTGGAGTTCCTGTAA 607 H53A(+39+69) CATTCAACTGTTGCCTCCGGTTCTGAAG 608 GTG H53A(+23+47) CTGAAGGTGTTCTTGTACTTCATCC 609 h53AON1 CTGTTGCCTCCGGTTCTG 610 F153A(-12+10) ATTCTTTCAACTAGAATAAAAG 611 huEx45.30.66 GCCATCCTGGAGTTCCTGTAAGATACC 612 AAA huEx45.30.71 CCAATGCCATCCTGGAGTTCCTGTAAG 613 ATA huEx45.30.79 GCCGCTGCCCAATGCCATCCTGGAGTT 614 CCT huEx45.30.83 GTTTGCCGCTGCCCAATGCCATCCTGG 615 AGT huEx45.30.88 CAACAGTTTGCCGCTGCCCAATGCCAT 616 CCT huEx45.30.92 CTGACAACAGTTTGCCGCTGCCCAATG 617 CCA huEx45.30.96 TGTTCTGACAACAGTTTGCCGCTGCCC 618 AAT huEx45.30.99 CAATGTTCTGACAACAGTTTGCCGCTG 619 CCC huEx45.30.103 CATTCAATGTTCTGACAACAGTTTGCCG 620 CT huEx45.30.120 TATTTCTTCCCCAGTTGCATTCAATGTT 621 CT huEx45.30.127 GCTGAATTATTTCTTCCCCAGTTGCATT 622 CA huEx45.30.132 GGATTGCTGAATTATTTCTTCCCCAGTT 623 GC huEx45.30.137 TTTGAGGATTGCTGAATTATTTCTTCCC 624 CA huEx53.30.84 GTACTTCATCCCACTGATTCTGAATTCT 625 TT huEx53.30.88 TCTTGTACTTCATCCCACTGATTCTGAA 626 TT huEx53.30.91 TGTTCTTGTACTTCATCCCACTGATTCT 627 GA huEx53.30.103 CGGTTCTGAAGGTGTTCTTGTACTTCAT 628 CC huEx53.30.106 CTCCGGTTCTGAAGGTGTTCTTGTACTT 629 CA huEx53.30.109 TGCCTCCGGTTCTGAAGGTGTTCTTGTA 630 CT huEx53.30.112 TGTTGCCTCCGGTTCTGAAGGTGTTCTT 631 GT huEx53.30.115 AACTGTTGCCTCCGGTTCTGAAGGTGT 632 TCT huEx53.30.118 TTCAACTGTTGCCTCCGGTTCTGAAGGT 633 GT h50AON1 h50AON2 Peptide Transporters (NH.sub.2 to COOH)*: rTAT RRRQRRKKRC 570 R.sub.9F.sub.2 RRRRRRRRRFFC 571 (RRAnx).sub.4B RRAhxRRAhxRRAhxRRAhxB 572 (RAhxR).sub.4AhxB; (P007) RAhxRRAhxRRAhxRRAhxRAhxB 573 (AhxRR).sub.4AhxB AhxRRAhxRRAhxRRAhxRRAhxB 574 (RAhx).sub.6B RAhxRAhxRAhxRAhxRAhxRAhxB 575 (RAhx).sub.8B RAhxRAhxRAhxRAhxRAhxRAhxRAhxRAhx 576 B (RAhxR).sub.5AhxB RAhxRRAhxRRAhxRRAhxRRAhxRAhxB 577 (RAhxRRBR).sub.2AhxB; RAhxRRBRRAhxRRBRAhxB 578 (CP06062) MSP ASSLNIA 579 Cell Penetrating Peptide / Homing Peptide / PMO Conjugates (NH.sub.2 to COOH and 5' to 3') MSP-PMO ASSLNIA-XB- 580 GGCCAAACCTCGGCTTACCTGAAAT 636 CP06062-MSP-PMO RXRRBRRXRRBR-XB-ASSLNIA-X- 581 GGCCAAACCTCGGCTTACCTGAAAT 636 MSP-CP06062-PMO ASSLNIA-X-RXRRBRRXRRBR-B- 582 GGCCAAACCTCGGCTTACCTGAAAT 636 CP06062-PMO RXRRBRRXRRBR-XB- 583 GGCCAAACCTCGGCTTACCTGAAAT 636 *Ahx is 6-aminohexanoic acid and B is beta-aianine.
Sequence CWU 1
1
651125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 1ctgcaggtaa aagcatatgg atcaa
25225DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 2atcgcctgca ggtaaaagca tatgg
25325DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 3gtcaaatcgc ctgcaggtaa aagca
25425DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 4gatctgtcaa atcgcctgca ggtaa
25525DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 5caacagatct gtcaaatcgc ctgca
25625DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 6tttctcaaca gatctgtcaa atcgc
25725DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 7ccatttctca acagatctgt caaat
25825DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 8ataatgaaaa cgccgccatt tctca
25925DNAArtificial SequenceAntisense sequence to target splice site
of proprocessed human dystrophin 9aaatatcttt atatcataat gaaaa
251025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 10tgttagccac tgattaaata tcttt
251125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 11aaactgttca gcttctgtta gccac
251225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 12ttgtgtcttt ctgagaaact gttca
251325DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 13ccaattctca ggaatttgtg tcttt
251425DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 14gtatttagca tgttcccaat tctca
251525DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 15cttaagatac catttgtatt tagca
251625DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 16cttaccttaa gataccattt gtatt
251725DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 17aaagacttac cttaagatac cattt
251825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 18aaatcaaaga cttaccttaa gatac
251925DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 19aaaacaaatc aaagacttac cttaa
252025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 20tcgaaaaaac aaatcaaaga cttac
252125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 21ctgtaagata ccaaaaaggc aaaac
252225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 22cctgtaagat accaaaaagg caaaa
252325DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 23agttcctgta agataccaaa aaggc
252425DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 24gagttcctgt aagataccaa aaagg
252525DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 25cctggagttc ctgtaagata ccaaa
252625DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 26tcctggagtt cctgtaagat accaa
252725DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 27gccatcctgg agttcctgta agata
252825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 28tgccatcctg gagttcctgt aagat
252925DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 29ccaatgccat cctggagttc ctgta
253025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 30cccaatgcca tcctggagtt cctgt
253125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 31gctgcccaat gccatcctgg agttc
253225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 32cgctgcccaa tgccatcctg gagtt
253325DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 33aacagtttgc cgctgcccaa tgcca
253425DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 34ctgacaacag tttgccgctg cccaa
253525DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 35gttgcattca atgttctgac aacag
253625DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 36gctgaattat ttcttcccca gttgc
253725DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 37attatttctt ccccagttgc attca
253825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 38ggcatctgtt tttgaggatt gctga
253925DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 39tttgaggatt gctgaattat ttctt
254025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 40aatttttcct gtagaatact ggcat
254125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 41atactggcat ctgtttttga ggatt
254225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 42accgcagatt caggcttccc aattt
254325DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 43aatttttcct gtagaatact ggcat
254425DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 44ctgtttgcag acctcctgcc accgc
254525DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 45agattcaggc ttcccaattt ttcct
254625DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 46ctcttttttc tgtctgacag ctgtt
254725DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 47acctcctgcc accgcagatt caggc
254825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 48cctacctctt ttttctgtct gacag
254925DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 49gacagctgtt tgcagacctc ctgcc
255025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 50gtcgccctac ctcttttttc tgtct
255125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 51gatctgtcgc cctacctctt ttttc
255225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 52tattagatct gtcgccctac ctctt
255325DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 53attcctatta gatctgtcgc cctac
255420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 54agataccaaa aaggcaaaac
205520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 55aagataccaa aaaggcaaaa
205620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 56cctgtaagat accaaaaagg
205720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 57gagttcctgt aagataccaa
205820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 58tcctggagtt cctgtaagat
205920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 59tgccatcctg gagttcctgt
206020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 60cccaatgcca tcctggagtt
206120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 61cgctgcccaa tgccatcctg
206220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 62ctgacaacag tttgccgctg
206320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 63gttgcattca atgttctgac
206420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 64attatttctt ccccagttgc
206520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 65tttgaggatt gctgaattat
206620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 66atactggcat ctgtttttga
206720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 67aatttttcct gtagaatact
206820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 68agattcaggc ttcccaattt
206920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 69acctcctgcc accgcagatt
207020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 70gacagctgtt tgcagacctc
207120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 71ctcttttttc tgtctgacag
207220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 72cctacctctt ttttctgtct
207320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 73gtcgccctac ctcttttttc
207420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 74gatctgtcgc cctacctctt
207520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 75tattagatct gtcgccctac
207620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 76attcctatta gatctgtcgc
207725DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 77gggggatttg agaaaataaa attac
257825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 78atttgagaaa ataaaattac cttga
257925DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 79ctagcctgga gaaagaagaa taaaa
258025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 80agaaaataaa attaccttga cttgc
258125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 81ttcttctagc ctggagaaag aagaa
258225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 82ataaaattac cttgacttgc tcaag
258325DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 83ttttgttctt ctagcctgga gaaag
258425DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 84attaccttga cttgctcaag ctttt
258525DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 85tattcttttg ttcttctagc ctgga
258625DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 86cttgacttgc tcaagctttt ctttt
258725DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 87caagatattc ttttgttctt ctagc
258825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 88cttttagttg ctgctctttt ccagg
258925DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 89ccaggttcaa gtgggatact agcaa
259025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 90atctctttga aattctgaca agata
259125DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 91agcaatgtta tctgcttcct ccaac
259225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 92aacaaattca tttaaatctc tttga
259325DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 93ccaaccataa aacaaattca tttaa
259425DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 94ttcctccaac cataaaacaa attca
259525DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 95tttaaatctc tttgaaattc tgaca
259625DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 96tgacaagata ttcttttgtt cttct
259725DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 97ttcaagtggg atactagcaa tgtta
259825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 98agatattctt ttgttcttct agcct
259925DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 99ctgctctttt ccaggttcaa gtggg
2510025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 100ttcttttgtt cttctagcct
ggaga 2510125DNAArtificial SequenceAntisense sequence to target
splice site of
proprocessed human dystrophin 101cttttctttt agttgctgct ctttt
2510225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 102ttgttcttct agcctggaga
aagaa 2510325DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 103cttctagcct
ggagaaagaa gaata 2510425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 104agcctggaga
aagaagaata aaatt 2510525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 105ctggagaaag
aagaataaaa ttgtt 2510620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 106gaaagaagaa
taaaattgtt 2010720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 107ggagaaagaa
gaataaaatt 2010820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 108agcctggaga
aagaagaata 2010920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 109cttctagcct
ggagaaagaa 2011020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 110ttgttcttct
agcctggaga 2011120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 111ttcttttgtt
cttctagcct 2011220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 112tgacaagata
ttcttttgtt 2011320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 113atctctttga
aattctgaca 2011420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 114aacaaattca
tttaaatctc 2011520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 115ttcctccaac
cataaaacaa 2011620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 116agcaatgtta
tctgcttcct 2011720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 117ttcaagtggg
atactagcaa 2011820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 118ctgctctttt
ccaggttcaa 2011920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 119cttttctttt
agttgctgct 2012020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 120cttgacttgc
tcaagctttt 2012120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 121attaccttga
cttgctcaag 2012220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 122ataaaattac
cttgacttgc 2012320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 123agaaaataaa
attaccttga 2012420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 124atttgagaaa
ataaaattac 2012520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 125gggggatttg
agaaaataaa 2012625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 126ctgaaacaga
caaatgcaac aacgt 2512725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 127agtaactgaa
acagacaaat gcaac 2512825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 128ccaccagtaa
ctgaaacaga caaat 2512925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 129ctcttccacc
agtaactgaa acaga 2513025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 130ggcaactctt
ccaccagtaa ctgaa 2513125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 131gcaggggcaa
ctcttccacc agtaa 2513225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 132ctggcgcagg
ggcaactctt ccacc 2513325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 133tttaattgtt
tgagaattcc ctggc 2513425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 134ttgtttgaga
attccctggc gcagg 2513525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 135gcacgggtcc
tccagtttca tttaa 2513625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 136tccagtttca
tttaattgtt tgaga 2513725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 137gcttatggga
gcacttacaa gcacg 2513825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 138tacaagcacg
ggtcctccag tttca 2513925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 139agtttatctt
gctcttctgg gctta 2514025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 140tctgcttgag
cttattttca agttt 2514125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 141atcttgctct
tctgggctta tggga 2514225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 142ctttatccac
tggagatttg tctgc 2514325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 143cttattttca
agtttatctt gctct 2514425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 144ctaaccttta
tccactggag atttg 2514525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 145atttgtctgc
ttgagcttat tttca 2514625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 146aatgtctaac
ctttatccac tggag 2514725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 147tggttaatgt
ctaaccttta tccac 2514825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 148agagatggtt
aatgtctaac cttta 2514925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 149acggaagaga
tggttaatgt ctaac 2515020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 150acagacaaat
gcaacaacgt 2015120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 151ctgaaacaga
caaatgcaac 2015220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 152agtaactgaa
acagacaaat 2015320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 153ccaccagtaa
ctgaaacaga 2015420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 154ctcttccacc
agtaactgaa 2015520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 155ggcaactctt
ccaccagtaa 2015620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 156ctggcgcagg
ggcaactctt 2015720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 157ttgtttgaga
attccctggc 2015820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 158tccagtttca
tttaattgtt 2015920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 159tacaagcacg
ggtcctccag 2016020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 160gcttatggga
gcacttacaa 2016120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 161atcttgctct
tctgggctta 2016220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 162cttattttca
agtttatctt 2016320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 163atttgtctgc
ttgagcttat 2016420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 164ctttatccac
tggagatttg 2016520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 165ctaaccttta
tccactggag 2016620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 166aatgtctaac
ctttatccac 2016720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 167tggttaatgt
ctaaccttta 2016820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 168agagatggtt
aatgtctaac 2016920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 169acggaagaga
tggttaatgt 2017025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 170ctgaaaggaa
aatacatttt aaaaa 2517125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 171cctgaaagga
aaatacattt taaaa 2517225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 172gaaacctgaa
aggaaaatac atttt 2517325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 173ggaaacctga
aaggaaaata cattt 2517425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 174ctctggaaac
ctgaaaggaa aatac 2517525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 175gctctggaaa
cctgaaagga aaata 2517625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 176gtaaagctct
ggaaacctga aagga 2517725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 177tcaggtaaag
ctctggaaac ctgaa 2517825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 178ctcaggtaaa
gctctggaaa cctga 2517925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 179gtttctcagg
taaagctctg gaaac 2518025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 180tgtttctcag
gtaaagctct ggaaa 2518125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 181aatttctcct
tgtttctcag gtaaa 2518225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 182tttgagcttc
aatttctcct tgttt 2518325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 183ttttatttga
gcttcaattt ctcct 2518425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 184aagctgccca
aggtctttta tttga 2518525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 185aggtcttcaa
gctttttttc aagct 2518625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 186ttcaagcttt
ttttcaagct gccca 2518725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 187gatgatttaa
ctgctcttca aggtc 2518825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 188ctgctcttca
aggtcttcaa gcttt 2518925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 189aggagataac
cacagcagca gatga 2519025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 190cagcagatga
tttaactgct cttca 2519125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 191atttccaact
gattcctaat aggag 2519225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 192cttggtttgg
ttggttataa atttc 2519325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 193caactgattc
ctaataggag ataac 2519425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 194cttaacgtca
aatggtcctt cttgg 2519525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 195ttggttataa
atttccaact gattc 2519625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 196cctaccttaa
cgtcaaatgg tcctt 2519725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 197tccttcttgg
tttggttggt tataa 2519825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 198agttccctac
cttaacgtca aatgg 2519925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 199caaaaagttc
cctaccttaa cgtca 2520025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 200taaagcaaaa
agttccctac cttaa 2520125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 201atatttaaag
caaaaagttc cctac
2520220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 202aggaaaatac attttaaaaa
2020320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 203aaggaaaata cattttaaaa
2020420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 204cctgaaagga aaatacattt
2020520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 205ggaaacctga aaggaaaata
2020620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 206gctctggaaa cctgaaagga
2020720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 207gtaaagctct ggaaacctga
2020820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 208ctcaggtaaa gctctggaaa
2020920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 209aatttctcct tgtttctcag
2021020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 210ttttatttga gcttcaattt
2021120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 211aagctgccca aggtctttta
2021220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 212ttcaagcttt ttttcaagct
2021320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 213ctgctcttca aggtcttcaa
2021420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 214cagcagatga tttaactgct
2021520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 215aggagataac cacagcagca
2021620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 216caactgattc ctaataggag
2021720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 217ttggttataa atttccaact
2021820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 218tccttcttgg tttggttggt
2021920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 219cttaacgtca aatggtcctt
2022020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 220cctaccttaa cgtcaaatgg
2022120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 221agttccctac cttaacgtca
2022220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 222caaaaagttc cctaccttaa
2022320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 223taaagcaaaa agttccctac
2022420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 224atatttaaag caaaaagttc
2022525DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 225ctggggaaaa gaacccatat
agtgc 2522625DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 226tcctggggaa
aagaacccat atagt 2522725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 227gtttcctggg
gaaaagaacc catat 2522825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 228cagtttcctg
gggaaaagaa cccat 2522925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 229tttcagtttc
ctggggaaaa gaacc 2523025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 230tatttcagtt
tcctggggaa aagaa 2523125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 231tgctatttca
gtttcctggg gaaaa 2523225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 232actgctattt
cagtttcctg gggaa 2523325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 233tgaactgcta
tttcagtttc ctggg 2523425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 234cttgaactgc
tatttcagtt tcctg 2523525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 235tagcttgaac
tgctatttca gtttc 2523625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 236tttagcttga
actgctattt cagtt 2523725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 237ttccacatcc
ggttgtttag cttga 2523825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 238tgccctttag
acaaaatctc ttcca 2523925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 239tttagacaaa
atctcttcca catcc 2524025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 240gtttttcctt
gtacaaatgc tgccc 2524125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 241gtacaaatgc
tgccctttag acaaa 2524225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 242cttcactggc
tgagtggctg gtttt 2524325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 243ggctggtttt
tccttgtaca aatgc 2524425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 244attaccttca
ctggctgagt ggctg 2524525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 245gcttcattac
cttcactggc tgagt 2524625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 246aggttgcttc
attaccttca ctggc 2524725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 247gctagaggtt
gcttcattac cttca 2524825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 248atattgctag
aggttgcttc attac 2524920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 249gaaaagaacc
catatagtgc 2025020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 250gggaaaagaa
cccatatagt 2025120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 251tcctggggaa
aagaacccat 2025220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 252cagtttcctg
gggaaaagaa 2025320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 253tatttcagtt
tcctggggaa 2025420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 254actgctattt
cagtttcctg 2025520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 255cttgaactgc
tatttcagtt 2025620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 256tttagcttga
actgctattt 2025720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 257ttccacatcc
ggttgtttag 2025820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 258tttagacaaa
atctcttcca 2025920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 259gtacaaatgc
tgccctttag 2026020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 260ggctggtttt
tccttgtaca 2026120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 261cttcactggc
tgagtggctg 2026220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 262attaccttca
ctggctgagt 2026320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 263gcttcattac
cttcactggc 2026420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 264aggttgcttc
attaccttca 2026520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 265gctagaggtt
gcttcattac 2026620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 266atattgctag
aggttgcttc 2026725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 267ctttaacaga
aaagcataca catta 2526825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 268tcctctttaa
cagaaaagca tacac 2526925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 269ttcctcttta
acagaaaagc ataca 2527025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 270taacttcctc
tttaacagaa aagca 2527125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 271ctaacttcct
ctttaacaga aaagc 2527225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 272tcttctaact
tcctctttaa cagaa 2527325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 273atcttctaac
ttcctcttta acaga 2527425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 274tcagatcttc
taacttcctc tttaa 2527525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 275ctcagatctt
ctaacttcct cttta 2527625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 276agagctcaga
tcttctaact tcctc 2527725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 277cagagctcag
atcttctaac ttcct 2527824DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 278cactcagagc
tcagatcttc tact 2427925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 279ccttccactc
agagctcaga tcttc 2528025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 280gtaaacggtt
taccgccttc cactc 2528125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 281ctttgccctc
agctcttgaa gtaaa 2528225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 282ccctcagctc
ttgaagtaaa cggtt 2528325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 283ccaggagcta
ggtcaggctg ctttg 2528425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 284ggtcaggctg
ctttgccctc agctc 2528525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 285aggctccaat
agtggtcagt ccagg 2528625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 286tcagtccagg
agctaggtca ggctg 2528725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 287cttacaggct
ccaatagtgg tcagt 2528825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 288gtatacttac
aggctccaat agtgg 2528925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 289atccagtata
cttacaggct ccaat 2529025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 290atgggatcca
gtatacttac aggct 2529125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 291agagaatggg
atccagtata cttac 2529220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 292acagaaaagc
atacacatta 2029320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 293tttaacagaa
aagcatacac 2029420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 294tcctctttaa
cagaaaagca 2029520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 295taacttcctc
tttaacagaa 2029620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 296tcttctaact
tcctctttaa 2029720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 297tcagatcttc
taacttcctc 2029820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 298ccttccactc
agagctcaga 2029920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 299gtaaacggtt
taccgccttc 2030020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 300ccctcagctc
ttgaagtaaa 2030120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 301ggtcaggctg
ctttgccctc 2030220DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
302tcagtccagg agctaggtca 2030320DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
303aggctccaat agtggtcagt 2030420DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
304cttacaggct ccaatagtgg 2030520DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
305gtatacttac aggctccaat 2030620DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
306atccagtata cttacaggct 2030720DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
307atgggatcca gtatacttac 2030820DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
308agagaatggg atccagtata 2030926DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
309ctaaaatatt ttgggttttt gcaaaa 2631026DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 310gctaaaatat tttgggtttt tgcaaa
2631126DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 311taggagctaa aatattttgg
gttttt 2631225DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 312agtaggagct
aaaatatttt gggtt 2531325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 313tgagtaggag
ctaaaatatt ttggg 2531426DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 314ctgagtagga
gctaaaatat tttggg 2631525DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
315cagtctgagt aggagctaaa atatt 2531626DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 316acagtctgag taggagctaa aatatt
2631726DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 317gagtaacagt ctgagtagga
gctaaa 2631825DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 318cagagtaaca
gtctgagtag gagct 2531926DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 319caccagagta
acagtctgag taggag 2632025DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
320gtcaccagag taacagtctg agtag 2532126DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 321aaccacaggt tgtgtcacca gagtaa
2632225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 322gttgtgtcac cagagtaaca
gtctg 2532326DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 323tggcagtttc
cttagtaacc acaggt 2632426DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
324atttctagtt tggagatggc agtttc 2632525DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 325ggaagatggc atttctagtt tggag
2532626DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 326catcaaggaa gatggcattt
ctagtt 2632726DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 327gagcaggtac
ctccaacatc aaggaa 2632825DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
328atctgccaga gcaggtacct ccaac 2532926DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 329aagttctgtc caagcccggt tgaaat
2633026DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 330cggttgaaat ctgccagagc
aggtac 2633126DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 331gagaaagcca
gtcggtaagt tctgtc 2633225DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
332gtcggtaagt tctgtccaag cccgg 2533326DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 333ataacttgat caagcagaga aagcca
2633425DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 334aagcagagaa agccagtcgg
taagt 2533526DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 335caccctctgt
gattttataa cttgat 2633626DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
336caaggtcacc caccatcacc ctctgt 2633725DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 337catcaccctc tgtgatttta taact
2533826DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 338cttctgcttg atgatcatct
cgttga 2633926DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 339ccttctgctt
gatgatcatc tcgttg 2634026DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
340atctcgttga tatcctcaag gtcacc 2634126DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 341tcataccttc tgcttgatga tcatct
2634226DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 342tcattttttc tcataccttc
tgcttg 2634326DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 343ttttctcata
ccttctgctt gatgat 2634426DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
344ttttatcatt ttttctcata ccttct 2634526DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 345ccaactttta tcattttttc tcatac
2634620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 346atattttggg tttttgcaaa
2034720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 347aaaatatttt gggtttttgc
2034820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 348gagctaaaat attttgggtt
2034920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 349agtaggagct aaaatatttt
2035020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 350gtctgagtag gagctaaaat
2035120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 351taacagtctg agtaggagct
2035220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 352cagagtaaca gtctgagtag
2035320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 353cacaggttgt gtcaccagag
2035420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 354agtttcctta gtaaccacag
2035520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 355tagtttggag atggcagttt
2035620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 356ggaagatggc atttctagtt
2035720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 357tacctccaac atcaaggaag
2035820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 358atctgccaga gcaggtacct
2035920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 359ccaagcccgg ttgaaatctg
2036020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 360gtcggtaagt tctgtccaag
2036120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 361aagcagagaa agccagtcgg
2036220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 362ttttataact tgatcaagca
2036320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 363catcaccctc tgtgatttta
2036420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 364ctcaaggtca cccaccatca
2036520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 365catctcgttg atatcctcaa
2036620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 366cttctgcttg atgatcatct
2036720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 367cataccttct gcttgatgat
2036820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 368tttctcatac cttctgcttg
2036920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 369cattttttct cataccttct
2037020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 370tttatcattt tttctcatac
2037120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 371caacttttat cattttttct
2037225DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 372ctgtaagaac aaatatccct
tagta 2537325DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 373tgcctgtaag
aacaaatatc cctta 2537425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 374gttgcctgta
agaacaaata tccct 2537525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 375attgttgcct
gtaagaacaa atatc 2537625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 376gcattgttgc
ctgtaagaac aaata 2537725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 377cctgcattgt
tgcctgtaag aacaa 2537825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 378atcctgcatt
gttgcctgta agaac 2537925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 379caaatcctgc
attgttgcct gtaag 2538025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 380tccaaatcct
gcattgttgc ctgta 2538125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 381tgttccaaat
cctgcattgt tgcct 2538225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 382tctgttccaa
atcctgcatt gttgc 2538325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 383aactggggac
gcctctgttc caaat 2538425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 384gcctctgttc
caaatcctgc attgt 2538525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 385cagcggtaat
gagttcttcc aactg 2538625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 386cttccaactg
gggacgcctc tgttc 2538725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 387cttgtttttc
aaattttggg cagcg 2538825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 388ctagcctctt
gattgctggt cttgt 2538925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 389ttttcaaatt
ttgggcagcg gtaat 2539025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 390ttcgatccgt
aatgattgtt ctagc 2539125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 391gattgctggt
cttgtttttc aaatt 2539225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 392cttacttcga
tccgtaatga ttgtt 2539325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 393ttgttctagc
ctcttgattg ctggt 2539425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 394aaaaacttac
ttcgatccgt aatga 2539525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 395tgttaaaaaa
cttacttcga tccgt 2539625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 396atgcttgtta
aaaaacttac ttcga 2539725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 397gtcccatgct
tgttaaaaaa cttac 2539820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 398agaacaaata
tcccttagta 2039920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 399gtaagaacaa
atatccctta 2040020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 400tgcctgtaag
aacaaatatc 2040120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 401attgttgcct
gtaagaacaa 2040220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin
402cctgcattgt tgcctgtaag 2040320DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
403caaatcctgc attgttgcct 2040420DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
404gcctctgttc caaatcctgc 2040520DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
405cttccaactg gggacgcctc 2040620DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
406cagcggtaat gagttcttcc 2040720DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
407ttttcaaatt ttgggcagcg 2040820DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
408gattgctggt cttgtttttc 2040920DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
409ttgttctagc ctcttgattg 2041020DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
410ttcgatccgt aatgattgtt 2041120DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
411cttacttcga tccgtaatga 2041220DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
412aaaaacttac ttcgatccgt 2041320DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
413tgttaaaaaa cttacttcga 2041420DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
414atgcttgtta aaaaacttac 2041520DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
415gtcccatgct tgttaaaaaa 2041625DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
416ctagaataaa aggaaaaata aatat 2541725DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 417aactagaata aaaggaaaaa taaat
2541825DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 418ttcaactaga ataaaaggaa
aaata 2541925DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 419ctttcaacta
gaataaaagg aaaaa 2542025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 420attctttcaa
ctagaataaa aggaa 2542125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 421gaattctttc
aactagaata aaagg 2542225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 422tctgaattct
ttcaactaga ataaa 2542325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 423attctgaatt
ctttcaacta gaata 2542425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 424ctgattctga
attctttcaa ctaga 2542525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 425cactgattct
gaattctttc aacta 2542625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 426tcccactgat
tctgaattct ttcaa 2542725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 427catcccactg
attctgaatt ctttc 2542825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 428tacttcatcc
cactgattct gaatt 2542925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 429ctgaaggtgt
tcttgtactt catcc 2543024DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 430cggttctgaa
ggtgttcttg tact 2443125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 431ctgttgcctc
cggttctgaa ggtgt 2543225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 432tttcattcaa
ctgttgcctc cggtt 2543325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 433taacatttca
ttcaactgtt gcctc 2543425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 434ttgtgttgaa
tcctttaaca tttca 2543525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 435tcttccttag
cttccagcca ttgtg 2543625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 436cttagcttcc
agccattgtg ttgaa 2543725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 437gtcctaagac
ctgctcagct tcttc 2543825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 438ctgctcagct
tcttccttag cttcc 2543925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 439ctcaagcttg
gctctggcct gtcct 2544025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 440ggcctgtcct
aagacctgct cagct 2544125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 441tagggaccct
ccttccatga ctcaa 2544225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 442tttggattgc
atctactgta taggg 2544325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 443accctccttc
catgactcaa gcttg 2544425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 444cttggtttct
gtgattttct tttgg 2544525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 445atctactgta
tagggaccct ccttc 2544625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 446ctaaccttgg
tttctgtgat tttct 2544725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 447tttcttttgg
attgcatcta ctgta 2544825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 448tgatactaac
cttggtttct gtgat 2544925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 449atctttgata
ctaaccttgg tttct 2545025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 450aaggtatctt
tgatactaac cttgg 2545125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 451ttaaaaaggt
atctttgata ctaac 2545220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 452ataaaaggaa
aaataaatat 2045320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 453gaataaaagg
aaaaataaat 2045420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 454aactagaata
aaaggaaaaa 2045520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 455ctttcaacta
gaataaaagg 2045620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 456gaattctttc
aactagaata 2045720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 457attctgaatt
ctttcaacta 2045820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 458tacttcatcc
cactgattct 2045919DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 459ctgaaggtgt
tcttgtact 1946020DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 460ctgttgcctc
cggttctgaa 2046120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 461taacatttca
ttcaactgtt 2046220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 462ttgtgttgaa
tcctttaaca 2046320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 463cttagcttcc
agccattgtg 2046420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 464ctgctcagct
tcttccttag 2046520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 465ggcctgtcct
aagacctgct 2046620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 466ctcaagcttg
gctctggcct 2046720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 467accctccttc
catgactcaa 2046820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 468atctactgta
tagggaccct 2046920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 469tttcttttgg
attgcatcta 2047020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 470cttggtttct
gtgattttct 2047120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 471ctaaccttgg
tttctgtgat 2047220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 472tgatactaac
cttggtttct 2047320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 473atctttgata
ctaaccttgg 2047420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 474aaggtatctt
tgatactaac 2047520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 475ttaaaaaggt
atctttgata 2047624DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 476ctatagattt
ttatgagaaa gaga 2447725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 477aactgctata
gatttttatg agaaa 2547825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 478tggccaactg
ctatagattt ttatg 2547925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 479gtctttggcc
aactgctata gattt 2548025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 480cggaggtctt
tggccaactg ctata 2548125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 481actggcggag
gtctttggcc aactg 2548225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 482tttgtctgcc
actggcggag gtctt 2548325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 483agtcatttgc
cacatctaca tttgt 2548425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 484tttgccacat
ctacatttgt ctgcc 2548525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 485ccggagaagt
ttcagggcca agtca 2548625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 486gtatcatctg
cagaataatc ccgga 2548725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 487taatcccgga
gaagtttcag ggcca 2548825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 488ttatcatgtg
gacttttctg gtatc 2548925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 489agaggcattg
atattctctg ttatc 2549025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 490atgtggactt
ttctggtatc atctg 2549125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 491cttttatgaa
tgcttctcca agagg 2549225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 492atattctctg
ttatcatgtg gactt 2549325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 493catacctttt
atgaatgctt ctcca 2549425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 494ctccaagagg
cattgatatt ctctg 2549525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 495taattcatac
cttttatgaa tgctt 2549625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 496cttttatgaa
tgcttctcca agagg 2549725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 497taatgtaatt
catacctttt atgaa 2549825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 498agaaataatg
taattcatac ctttt 2549925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 499gttttagaaa
taatgtaatt catac 2550019DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 500gatttttatg
agaaagaga 1950120DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 501ctatagattt
ttatgagaaa 2050220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 502aactgctata
gatttttatg
2050320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 503tggccaactg ctatagattt
2050420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 504gtctttggcc aactgctata
2050520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 505cggaggtctt tggccaactg
2050620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 506tttgtctgcc actggcggag
2050720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 507tttgccacat ctacatttgt
2050820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 508ttcagggcca agtcatttgc
2050920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 509taatcccgga gaagtttcag
2051020DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 510gtatcatctg cagaataatc
2051120DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 511atgtggactt ttctggtatc
2051220DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 512atattctctg ttatcatgtg
2051320DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 513ctccaagagg cattgatatt
2051420DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 514cttttatgaa tgcttctcca
2051520DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 515catacctttt atgaatgctt
2051620DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 516taattcatac cttttatgaa
2051720DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 517taatgtaatt catacctttt
2051820DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 518agaaataatg taattcatac
2051920DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 519gttttagaaa taatgtaatt
2052025DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 520ctgcaaagga ccaaatgttc
agatg 2552125DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 521tcaccctgca
aaggaccaaa tgttc 2552225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 522ctcactcacc
ctgcaaagga ccaaa 2552325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 523tctcgctcac
tcaccctgca aagga 2552425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 524cagcctctcg
ctcactcacc ctgca 2552525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 525caaagcagcc
tctcgctcac tcacc 2552625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 526tcttccaaag
cagcctctcg ctcac 2552725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 527tctatgagtt
tcttccaaag cagcc 2552825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 528gttgcagtaa
tctatgagtt tcttc 2552925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 529gaactgttgc
agtaatctat gagtt 2553025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 530ttccaggtcc
agggggaact gttgc 2553125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 531gtaagccagg
caagaaactt ttcca 2553225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 532ccaggcaaga
aacttttcca ggtcc 2553325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 533tggcagttgt
ttcagcttct gtaag 2553425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 534ggtagcatcc
tgtaggacat tggca 2553525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 535gacattggca
gttgtttcag cttct 2553625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 536tctaggagcc
tttccttacg ggtag 2553725DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 537cttttactcc
cttggagtct tctag 2553825DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 538gagcctttcc
ttacgggtag catcc 2553925DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 539ttgccattgt
ttcatcagct ctttt 2554025DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 540cttggagtct
tctaggagcc tttcc 2554125DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 541cttacttgcc
attgtttcat cagct 2554225DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 542cagctctttt
actcccttgg agtct 2554325DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 543cctgacttac
ttgccattgt ttcat 2554425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 544aaatgcctga
cttacttgcc attgt 2554525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 545agcggaaatg
cctgacttac ttgcc 2554625DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 546gctaaagcgg
aaatgcctga cttac 2554720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 547aaggaccaaa
tgttcagatg 2054820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 548ctgcaaagga
ccaaatgttc 2054920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 549tcaccctgca
aaggaccaaa 2055020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 550ctcactcacc
ctgcaaagga 2055120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 551tctcgctcac
tcaccctgca 2055220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 552cagcctctcg
ctcactcacc 2055320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 553caaagcagcc
tctcgctcac 2055420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 554tctatgagtt
tcttccaaag 2055520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 555gaactgttgc
agtaatctat 2055620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 556ttccaggtcc
agggggaact 2055720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 557ccaggcaaga
aacttttcca 2055820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 558ttcagcttct
gtaagccagg 2055920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 559gacattggca
gttgtttcag 2056020DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 560ggtagcatcc
tgtaggacat 2056120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 561gagcctttcc
ttacgggtag 2056220DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 562cttggagtct
tctaggagcc 2056320DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 563cagctctttt
actcccttgg 2056420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 564ttgccattgt
ttcatcagct 2056520DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 565cttacttgcc
attgtttcat 2056620DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 566cctgacttac
ttgccattgt 2056720DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 567aaatgcctga
cttacttgcc 2056820DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 568agcggaaatg
cctgacttac 2056920DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 569gctaaagcgg
aaatgcctga 2057010PRTArtificial SequencePeptide Transporter for
Intracellular Delivery of PMO 570Arg Arg Arg Gln Arg Arg Lys Lys
Arg Cys1 5 1057112PRTArtificial SequencePeptide Transporter for
Intracellular Delivery of PMO 571Arg Arg Arg Arg Arg Arg Arg Arg
Arg Phe Phe Cys1 5 1057213PRTArtificial SequencePeptide Transporter
for Intracellular Delivery of PMOMOD_RES3, 6, 9, 12X =
6-aminohexanoic acidMOD_RES13X = beta alanine 572Arg Arg Xaa Arg
Arg Xaa Arg Arg Xaa Arg Arg Xaa Xaa1 5 1057314PRTArtificial
SequencePeptide Transporter for Intracellular Delivery of
PMOMOD_RES2, 5, 8, 11, 13X = 6-aminohexanoic acidMOD_RES14X = beta
alanine 573Arg Xaa Arg Arg Xaa Arg Arg Xaa Arg Arg Xaa Arg Xaa Xaa1
5 1057414PRTArtificial SequencePeptide Transporter for
Intracellular Delivery of PMOMOD_RES1, 4, 7, 10, 13X =
6-aminohexanoic acidMOD_RES14X = beta alanine 574Xaa Arg Arg Xaa
Arg Arg Xaa Arg Arg Xaa Arg Arg Xaa Xaa1 5 1057513PRTArtificial
SequencePeptide Transporter for Intracellular Delivery of
PMOMOD_RES2, 4, 6, 8, 10, 12X = 6-aminohexanoic acidMOD_RES13X =
beta alanine 575Arg Xaa Arg Xaa Arg Xaa Arg Xaa Arg Xaa Arg Xaa
Xaa1 5 1057617PRTArtificial SequencePeptide Transporter for
Intracellular Delivery of PMOMOD_RES2, 4, 6, 8, 10, 12, 14, 16X =
6-aminohexanoic acidMOD_RES17X = beta alanine 576Arg Xaa Arg Xaa
Arg Xaa Arg Xaa Arg Xaa Arg Xaa Arg Xaa Arg Xaa1 5 10
15Xaa57717PRTArtificial SequencePeptide Transporter for
Intracellular Delivery of PMOMOD_RES2, 5, 8, 11, 14, 16X =
6-aminohexanoic acidMOD_RES17X = beta alanine 577Arg Xaa Arg Arg
Xaa Arg Arg Xaa Arg Arg Xaa Arg Arg Xaa Arg Xaa1 5 10
15Xaa57814PRTArtificial SequencePeptide Transporter for
Intracellular Delivery of PMOMOD_RES2, 8, 13X = 6-aminohexanoic
acidMOD_RES5, 11, 14X = beta alanine 578Arg Xaa Arg Arg Xaa Arg Arg
Xaa Arg Arg Xaa Arg Xaa Xaa1 5 105797PRTArtificial SequencePeptide
Transporter for Intracellular Delivery of PMO 579Ala Ser Ser Leu
Asn Ile Ala1 55809PRTArtificial SequenceCell Penetrating Peptide /
Homing Peptide / PMO ConjugatesMOD_RES8X = a neutral amino acid
C(=O) (CHR)n NH-, where n is 2 to 7 and each R is independently H
or methylMOD_RES9X = Beta alanine 580Ala Ser Ser Leu Asn Ile Ala
Xaa Xaa1 558122PRTArtificial SequenceCell Penetrating Peptide /
Homing Peptide / PMO ConjugatesMOD_RES2, 8, 13, 22X = a neutral
amino acid C(=O) (CHR)n NH-, where n is 2 to 7 and each R is
independently H or methylMOD_RES5, 11, 14X = beta alanine 581Arg
Xaa Arg Arg Xaa Arg Arg Xaa Arg Arg Xaa Arg Xaa Xaa Ala Ser1 5 10
15Ser Leu Asn Ile Ala Xaa 2058221PRTArtificial SequenceCell
Penetrating Peptide / Homing Peptide / PMO ConjugatesMOD_RES8, 10,
16X = a neutral amino acid C(=O) (CHR)n NH-, where n is 2 to 7 and
each R is independently H or methylMOD_RES13, 19, 21Xaa = beta
alanine 582Ala Ser Ser Leu Asn Ile Ala Xaa Arg Xaa Arg Arg Xaa Arg
Arg Xaa1 5 10 15Arg Arg Xaa Arg Xaa 2058314PRTArtificial
SequenceCell Penetrating Peptide / Homing Peptide / PMO
ConjugatesMOD_RES2, 8, 13X = a neutral amino acid C(=O) (CHR)n NH-,
where n is 2 to 7 and each R is independently H or methylMOD_RES5,
11, 14X = beta alanine 583Arg Xaa Arg Arg Xaa Arg Arg Xaa Arg Arg
Xaa Arg Xaa Xaa1 5 1058429DNAArtificial SequenceAntisense sequence
to target splice site of proprocessed human dystrophin
584ccactcagag ctcagatctt ctaacttcc 2958525DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 585gggatccagt atacttacag gctcc
2558627DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 586cttccactca gagctcagat
cttctaa 2758730DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 587acatcaagga
agatggcatt tctagtttgg 3058830DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
588ctccaacatc aaggaagatg gcatttctag 3058924DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 589ttctgtccaa gcccggttga aatc 2459021DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 590cacccaccat caccctcygt g 2159122DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 591atcatctcgt tgatatcctc aa 2259225DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 592acatcaagga agatggcatt tctag
2559326DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 593accagagtaa cagtctgagt
aggagc 2659420DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 594tcaaggaaga
tggcatttct 2059523DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 595cctctgtgat
tttataactt gat
2359625DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 596atcatttttt ctcatacctt
ctgct 2559723DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 597ctcatacctt
ctgcttgatg atc 2359817DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 598tggcatttct
agtttgg 1759923DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 599ccagagcagg
tacctccaac atc 2360024DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 600tgttcagctt
ctgttagcca ctga 2460120DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 601tttgtgtctt
tctgagaaac 2060219DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 602cgccgccatt
tctcaacag 1960320DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 603atctgtcaaa
tcgcctgcag 2060420DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 604tgtttttgag
gattgctgaa 2060519DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 605gctgaattat
ttcttcccc 1960617DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 606gcccaatgcc atcctgg
1760726DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 607ccaatgccat cctggagttc
ctgtaa 2660831DNAArtificial SequenceAntisense sequence to target
splice site of proprocessed human dystrophin 608cattcaactg
ttgcctccgg ttctgaaggt g 3160925DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
609ctgaaggtgt tcttgtactt catcc 2561018DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 610ctgttgcctc cggttctg 1861122DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 611attctttcaa ctagaataaa ag 2261230DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 612gccatcctgg agttcctgta agataccaaa
3061330DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 613ccaatgccat cctggagttc
ctgtaagata 3061430DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 614gccgctgccc
aatgccatcc tggagttcct 3061530DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
615gtttgccgct gcccaatgcc atcctggagt 3061630DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 616caacagtttg ccgctgccca atgccatcct
3061730DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 617ctgacaacag tttgccgctg
cccaatgcca 3061830DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 618tgttctgaca
acagtttgcc gctgcccaat 3061930DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
619caatgttctg acaacagttt gccgctgccc 3062030DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 620cattcaatgt tctgacaaca gtttgccgct
3062130DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 621tatttcttcc ccagttgcat
tcaatgttct 3062230DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 622gctgaattat
ttcttcccca gttgcattca 3062330DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
623ggattgctga attatttctt ccccagttgc 3062430DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 624tttgaggatt gctgaattat ttcttcccca
3062530DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 625gtacttcatc ccactgattc
tgaattcttt 3062630DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 626tcttgtactt
catcccactg attctgaatt 3062730DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
627tgttcttgta cttcatccca ctgattctga 3062830DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 628cggttctgaa ggtgttcttg tacttcatcc
3062930DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 629ctccggttct gaaggtgttc
ttgtacttca 3063030DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 630tgcctccggt
tctgaaggtg ttcttgtact 3063130DNAArtificial SequenceAntisense
sequence to target splice site of proprocessed human dystrophin
631tgttgcctcc ggttctgaag gtgttcttgt 3063230DNAArtificial
SequenceAntisense sequence to target splice site of proprocessed
human dystrophin 632aactgttgcc tccggttctg aaggtgttct
3063330DNAArtificial SequenceAntisense sequence to target splice
site of proprocessed human dystrophin 633ttcaactgtt gcctccggtt
ctgaaggtgt 3063425DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 634taaagctctg
gaaacctgaa aggaa 2563525DNAArtificial SequenceAntisense sequence to
target splice site of proprocessed human dystrophin 635ttcagcttct
gtaagccagg caaga 2563625DNAArtificial SequenceAntisense oligomer
636ggccaaacct cggcttacct gaaat 2563712PRTArtificial
Sequencearginine-rich peptide transportVARIANT1, 3, 4, 7, 9, 10Xaa
= K, R or arginine analogMOD_RES2, 8Xaa = a neutral amino acid
C(=O) (CHR)n NH-, where n is 2 to 7 and each R is independently H
or methylVARIANT5, 6, 11, 12Xaa = an alpha-amino acid having a
neutral aralkyl side chain 637Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa
Xaa Xaa Xaa1 5 1063812PRTArtificial Sequencearginine-rich peptide
transportMOD_RES2, 6, 8, 12Xaa = a neutral amino acid C(=O) (CHR)n
NH-, where n is 2 to 7 and each R is independently H or methyl
638Arg Xaa Arg Arg Arg Xaa Arg Xaa Arg Arg Arg Xaa1 5
106399PRTArtificial Sequencearginine-rich peptide
transportMOD_RES3, 5, 9Xaa = a neutral amino acid C(=O) (CHR)n NH-,
where n is 2 to 7 and each R is independently H or methyl 639Arg
Arg Xaa Arg Xaa Arg Arg Arg Xaa1 564024DNAArtificial SequencePCR
Primer 640ccagagcttt acctgagaaa caag 2464121DNAArtificial
SequencePCR Primer 641ccagccactc agccagtgaa g 2164224DNAArtificial
SequencePCR Primer 642cgatccgtaa tgattgttct agcc
2464324DNAArtificial SequencePCR Primer 643catttcattc aactgttgcc
tccg 2464421DNAArtificial SequencePCR Primer 644caatgctcct
gacctctgtg c 2164522DNAArtificial SequencePCR Primer 645gtctacaaca
aagctcaggt cg 2264625DNAArtificial SequencePCR Primer 646gcaatgttat
ctgcttcctc caacc 2564722DNAArtificial SequencePCR Primer
647gctcttttcc aggttcaagt gg 2264823DNAArtificial SequencePCR Primer
648cttggacaga acttaccgac tgg 2364921DNAArtificial SequencePCR
Primer 649gcaggatttg gaacagaggc g 2165023DNAArtificial SequencePCR
Primer 650catctacatt tgtctgccac tgg 2365123DNAArtificial
SequencePCR Primer 651gtttcttcca aagcagcctc tcg 23

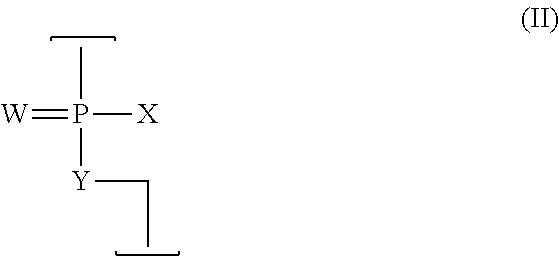



D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025

D00026

D00027

D00028

D00029

D00030

D00031

D00032

D00033

D00034

D00035

D00036

D00037

D00038

D00039

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.