Glycolipopeptide Biosurfactants
Kerr; Russell Greig ; et al.
U.S. patent application number 16/090888 was filed with the patent office on 2019-05-02 for glycolipopeptide biosurfactants. This patent application is currently assigned to Croda International PLC. The applicant listed for this patent is Croda International PLC. Invention is credited to Fabrice Berrue, Bradley Arnold Haltli, Russell Greig Kerr, Douglas Hubert Marchbank.
| Application Number | 20190127411 16/090888 |
| Document ID | / |
| Family ID | 58632356 |
| Filed Date | 2019-05-02 |






View All Diagrams
| United States Patent Application | 20190127411 |
| Kind Code | A1 |
| Kerr; Russell Greig ; et al. | May 2, 2019 |
GLYCOLIPOPEPTIDE BIOSURFACTANTS
Abstract
Surfactants based on a newly discovered class of compounds include a hydrophobic lipid oligomer covalently linked to a peptide or peptide-like chain and a carbohydrate moiety, and a serine-leucinol dipeptide linked to the lipid oligomer. Such surfactants can be used to create an oil-in-water or water-in-oil emulsion by mixing together a polar component; a non-polar component; and the surfactant. Biosurfactants of the newly discovered class can be made by isolating and culturing a microorganism which produces the biosurfactant, and then isolating the biosurfactant from the culture. A microorganism can be engineered to produce biosurfactant of this newly discovered class by expressing a set of heterologous genes involved in the biosynthesis of the biosurfactant in the microorganism.
| Inventors: | Kerr; Russell Greig; (Charlottetown, CA) ; Haltli; Bradley Arnold; (New Haven, CA) ; Marchbank; Douglas Hubert; (Stratford, CA) ; Berrue; Fabrice; (Halifax, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | Croda International PLC East Yorkshire GB |
||||||||||
| Family ID: | 58632356 | ||||||||||
| Appl. No.: | 16/090888 | ||||||||||
| Filed: | April 6, 2017 | ||||||||||
| PCT Filed: | April 6, 2017 | ||||||||||
| PCT NO: | PCT/EP2017/058296 | ||||||||||
| 371 Date: | October 3, 2018 |
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C11D 1/10 20130101; C12P 19/44 20130101; C11D 3/381 20130101; C07H 15/04 20130101; C11D 1/008 20130101; C11D 1/662 20130101; C02F 3/34 20130101; C12N 15/52 20130101; C12R 1/01 20130101; C07H 1/08 20130101 |
| International Class: | C07H 15/04 20060101 C07H015/04; C07H 1/08 20060101 C07H001/08; C11D 1/66 20060101 C11D001/66; C12N 15/52 20060101 C12N015/52; C12P 19/44 20060101 C12P019/44 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Apr 6, 2016 | GB | 1605875.2 |
Claims
1. A purified biosurfactant comprising a hydrophobic lipid component comprising a carboxyl end and a hydroxyl end, wherein the lipid component is covalently linked to (i) a peptide or peptide-like chain at the carboxyl end of the lipid component and (ii) a carbohydrate moiety at the hydroxyl end of the lipid component via a glycosidic linkage.
2. The purified biosurfactant according to claim 1, wherein the peptide chain comprises in the range of between 2 and 10 amino acids.
3. The purified biosurfactant according to claim 1, wherein the lipid component comprises in the range of between 1 and 6 alkanoic acid moieties
4. The purified biosurfactant according to claim 1, wherein the lipid component comprises an acyle chain, and wherein the length of each said acyl chain is in the range of between C.sub.4 to C.sub.20.
5. The purified biosurfactant according to claim 1, wherein the carbohydrate moiety may be selected from saccharides including glucose, fructose, galactose, mannose, ribose, or deoxy saccharide variants including deoxyribose, fucose, or rhamnose.
6. The purified biosurfactant according to claim 1, wherein the peptide or peptide-like chain comprises a serine-leucinol dipeptide.
7. The purified biosurfactant according to claim 1, wherein the lipid component comprises three 3-hydroxyalkanoic acid moieties.
8. The purified biosurfactant of claim 4, wherein the length of each acyl chain of the lipid component is C.sub.10.
9. The purified biosurfactant according to claim 1, wherein the carbohydrate moiety comprises a rhamnose moiety attached to the lipid component via a glycosidic linkage.
10. The purified biosurfactant according to claim 9, wherein the carbohydrate moiety comprises two rhamnose moieties.
11. The purified biosurfactant according to claim 1, wherein the lipid component comprises three .beta.-hydroxyalkanoic acid moieties, the length of each acyl chain of the lipid component is C.sub.10, and the carbohydrate moiety comprises a rhamnose moiety attached to the lipid component via a glycosidic linkage.
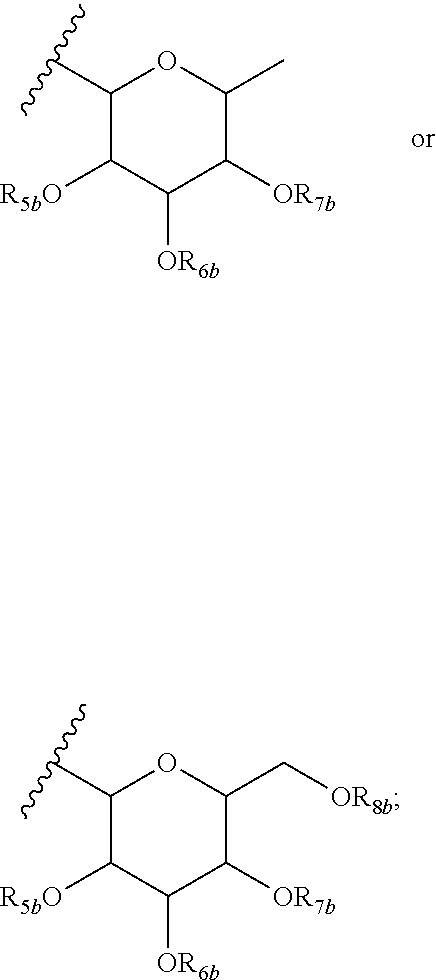
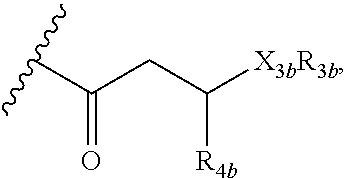
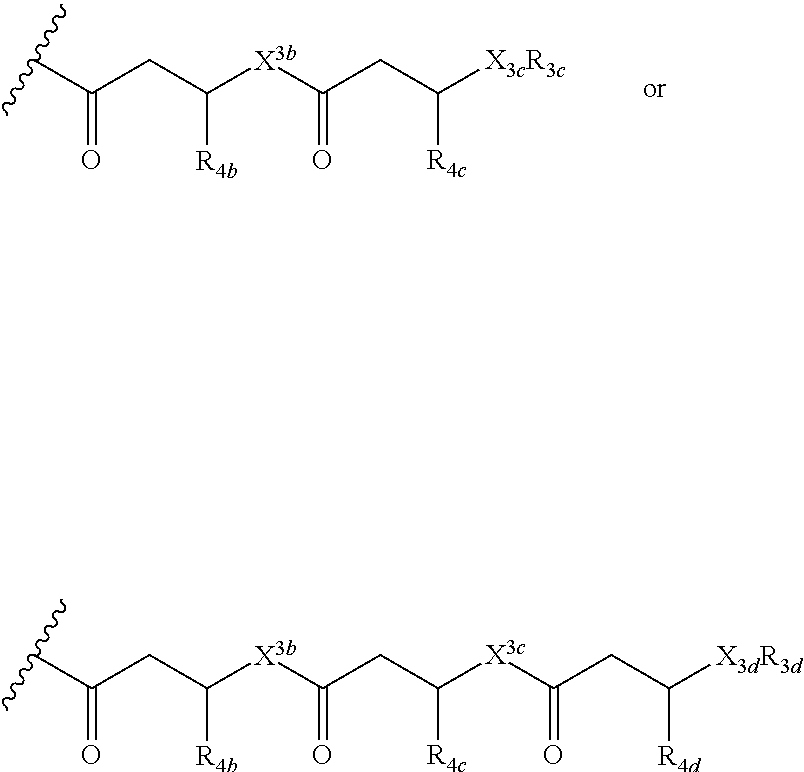
12. A purified biosurfactant comprising a peptide or peptide-like portion covalently bound to a lipid portion, wherein the biosurfactant comprises the structure: ##STR00049## wherein R.sub.1a is selected from the group consisting of H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), N(CH.sub.3).sub.2, and a peptide or peptide-like structure having the structure: ##STR00050## wherein R.sub.1b, R.sub.1c, and R.sub.1d, are selected from the group consisting of H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), and N(CH.sub.3).sub.2; R.sub.2a, R.sub.2b, R.sub.2c, and R.sub.2d are each independently an amino acid side chain; X.sub.1a, X.sub.1b, X.sub.1c, and X.sub.1d are each independently selected from the group consisting of one oxygen atom and two hydrogen atoms; X.sub.2a, X.sub.2b, X.sub.2c, and X.sub.2d are each independently selected from the group consisting of NH, N(CH.sub.3), and O; R.sub.3a is selected from the group consisting of a carbohydrate portion, and a lipid selected from the group consisting of a monomer having the structure: ##STR00051## and an oligomer selected from the group consisting of: ##STR00052## wherein X.sub.3a, X.sub.3b, X.sub.3c, and X.sub.3d are each independently selected from the group consisting of NH, N(CH.sub.3), and O; R.sub.3a, R.sub.3b, R.sub.3c, and R.sub.3d comprises a carbohydrate portion comprising a monomer selected from the group consisting of: ##STR00053## wherein R.sub.5a, R.sub.6a, R.sub.7a, and R.sub.8a are each independently selected from the group consisting of a hydrogen atom, methyl, acetyl, and a carbohydrate; and R.sub.4a, R.sub.4b, R.sub.4c, and R.sub.4d are each independently selected from the group consisting of a hydrogen atom, methyl, and a C.sub.2 to C.sub.19 saturated or unsaturated linear, branched-chain, cyclic, or aromatic hydrocarbon groups.
13. The purified biosurfactant of claim 12, wherein at least one of R.sub.6a, R.sub.7a, and R.sub.8a comprises a carbohydrate comprising a monomer selected from the group consisting of: ##STR00054## wherein R.sub.5b, R.sub.6b, R.sub.7b, and R.sub.8b are each independently selected from the group consisting of a hydrogen atom, methyl, acetyl, and a carbohydrate.
14. The purified biosurfactant of claim 12, wherein the peptide or peptide-like portion comprises at least one proline or proline-like monomer having the structure: ##STR00055## wherein X.sub.4 is selected from the group consisting of one oxygen atom and two hydrogen atoms.
15. The purified biosurfactant of claim 14, wherein the peptide or peptide-like portion comprises a single proline or proline-like monomer or a terminal proline or proline-like monomer having the structure: ##STR00056## wherein R.sub.9 is selected from the group consisting of H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), and N(CH.sub.3).sub.2; and X.sub.4 is selected from the group consisting of one oxygen atom and two hydrogen atoms.
16. A purified biosurfactant, wherein the biosurfactant has the structure: ##STR00057## wherein R.sub.5a, R.sub.6a, R.sub.7a, R.sub.10, and R.sub.11 are each independently selected from the group consisting of a hydrogen atom and acetyl; and n.sub.1, n.sub.2, and n.sub.3 are integers each independently selected from 1 to 7.
17. The purified biosurfactant of claim 16, wherein the biosurfactant has the structure: ##STR00058## wherein R.sub.5a, R.sub.5b, R.sub.6b, R.sub.7a, R.sub.7b, R.sub.10, and R.sub.11 are each independently selected from the group consisting of a hydrogen atom and acetyl; and n.sub.1, n.sub.2, and n.sub.3 are integers each independently selected from 1 to 7.
18. The purified biosurfactant of claim 16, wherein the biosurfactant has the structure: ##STR00059##
19. The purified biosurfactant of claim 17, wherein the biosurfactant has the structure: ##STR00060##
20. The purified biosurfactant of claim 17, wherein the biosurfactant has the structure: ##STR00061##
21. A method of making the biosurfactant of claim 1, the method comprising the steps of: (a) isolating a microorganism which comprises the biosurfactant; (b) placing the microorganism in a culture under conditions that promote the synthesis of the biosurfactant; and (c) isolating the biosurfactant from the culture.
22. The method according to claim 21, wherein the microorganism belongs to the genus and species Variovorax paradoxus and is strain RKNM-096 as deposited at the NRRL under accession number B-67038.
23. An organism consisting of Variovorax paradoxus, strain B-67038, Agricultural Research Service Culture Collection accession number B-67038.
24. An emulsified oil-in-water or water-in-oil composition comprising a polar component, a non-polar component, and the biosurfactant as claimed in claim 1.
25. An isolated microorganism engineered to produce the biosurfactant of claim 1, wherein a set of heterologous genes exhibiting at least 70% similarity to SEQ IDs 3, 5, 7, 9, 11 and 13 have been introduced into the microorganism.
26. A method of modifying natural glyclolipopeptide surfactants by adding additional rhamnose moieties using recombinantly expressed RIpE [SEQ IDs 11, 12, 23 and 24].
Description
FIELD OF THE INVENTION
[0001] This invention relates generally to the fields of surfactant chemistry, biochemistry, and microbiology. More specifically the invention relates to biosurfactants having a hydrophobic lipid oligomer covalently linked to a peptide or peptide-like (e.g. non-proteinogenic amino acid or single amino acid) chain and a carbohydrate moiety, various amino acid and nucleic acid sequences which encode components of biosynthetic pathways for these biosurfactants, and methods of making and using these biosurfactants.
BACKGROUND
[0002] Surfactants are amphiphilic chemicals that possess both hydrophobic and hydrophilic moieties which allow them to interact with polar and non-polar systems. Surfactants exert their activity at interfaces between different phases (gas, liquid, solid) and as a result exhibit a range of functions including, but not limited to the ability to act as detergents, emulsifiers, wetting agents and foaming agents. Most chemical surfactants are alkyl sulfates or sulfonates derived from petro- or oleo-chemical sources. The use of these products has been steadily growing with an estimated worldwide consumption of 13 million tonnes in 2008 and an estimated market value of $27 billion (USD) in 2012. In response to environmental and sustainability concerns, many companies utilizing chemical surfactants in their products have been exploring environmentally responsible alternatives as partial or full replacements for chemical surfactants. An alternative to chemical surfactants are biosurfactants, which are surface active molecules originating from microorganisms. These surfactants offer advantages over chemical surfactants such as production from sustainably produced feed stocks, biodegradability and lower toxicity.
SUMMARY
[0003] It was discovered that the bacterium Variovorax paradoxus RKNM-096, deposited on Apr. 10, 2015 as accession number NRRL B-67038 under the terms of the Budapest Treaty with the Agricultural Research Service Culture Collection (NRRL, 1818 North University Street, Peoria, Ill., 61064) produces a previously unknown class of biosurfactants termed "glycolipopeptides". Unlike known biosurfactants, glycolipopeptides typically contain a hydrophobic lipid oligomer covalently linked to a peptide chain and a carbohydrate moiety.
[0004] The deposit of NRRL B-67038 in support of this application was made by Nautilus Bioscience Canada Inc., 550 Unv. Ave., Charlottetown, PE, Canada, C1A4P3. Nautilus Bioscience Canada Inc. authorise the applicant to refer to the deposited biological material in this application and give their unreserved and irrevocable consent to the materials being made available to the public in accordance with appropriate national laws governing the deposit of these materials, such as Rule 31 and 33 EPC. The expert solution under Rule 32 EPC is also hereby requested.
[0005] Described herein are purified biosurfactants that include a hydrophobic lipid component including a carboxyl end and a hydroxyl end, wherein the lipid component is covalently linked to (i) a peptide or peptide-like chain at the carboxyl end of the lipid component and (ii) a carbohydrate moiety at the hydroxyl end of the lipid component via a glycosidic linkage. The peptide or peptide-like chain can include a serine-leucinol dipeptide, the lipid component can include three .beta.-hydroxyalkanoic acid moieties (e.g., wherein the length of each acyl chain of the lipid component is C.sub.6, C.sub.8, C.sub.10, or C.sub.12), and the carbohydrate moiety can include a rhamnose moiety attached to the lipid component via a glycosidic linkage. In certain embodiments, the carbohydrate moiety can include two rhamnose moieties and/or an acetyl group. Analogues and derivatives of these glycolipopeptides can be made by conventional methods.
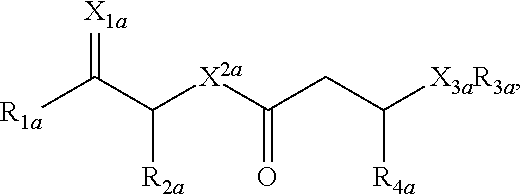
[0006] Glycolipopeptides can have the structure:
##STR00001##
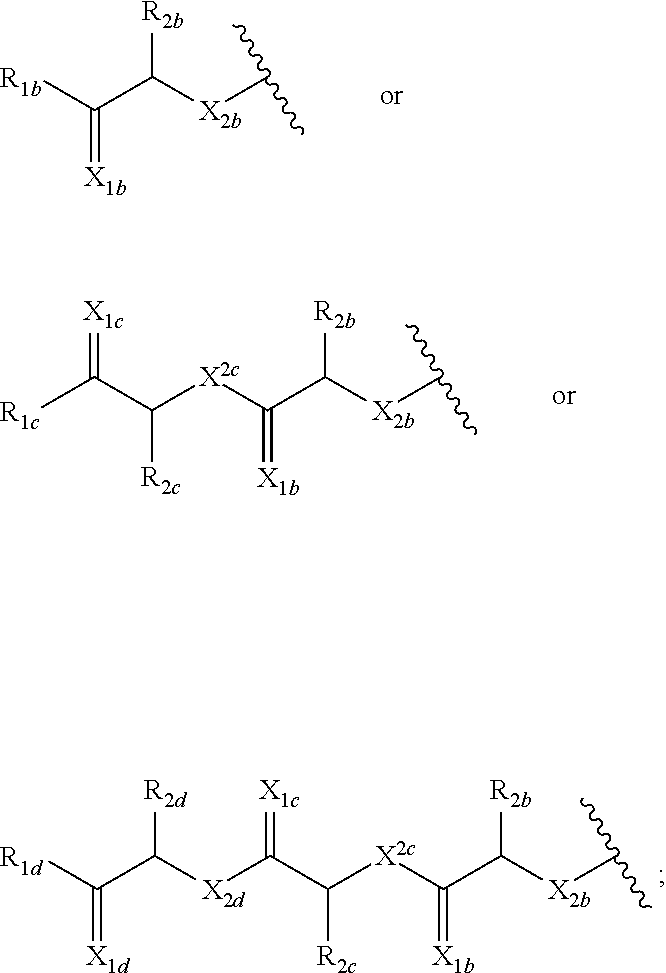
wherein R.sub.1a is H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), N(CH.sub.3).sub.2, or a peptide or peptide-like structure having the structure:
##STR00002##
wherein R.sub.1b, R.sub.1c, and R.sub.1d, are H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), or N(CH.sub.3).sub.2; R.sub.2a, R.sub.2b, R.sub.2c, and R.sub.2d are each independently an amino acid side chain; X.sub.1a, X.sub.1b, X.sub.1c, and X.sub.1d are each independently one oxygen atom or two hydrogen atoms; X.sub.2a, X.sub.2b, X.sub.2c, and X.sub.2d are each independently NH, N(CH.sub.3), or O; R.sub.3a is a carbohydrate portion or a lipid monomer having the structure:
##STR00003##
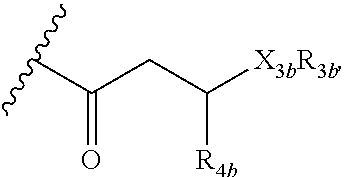
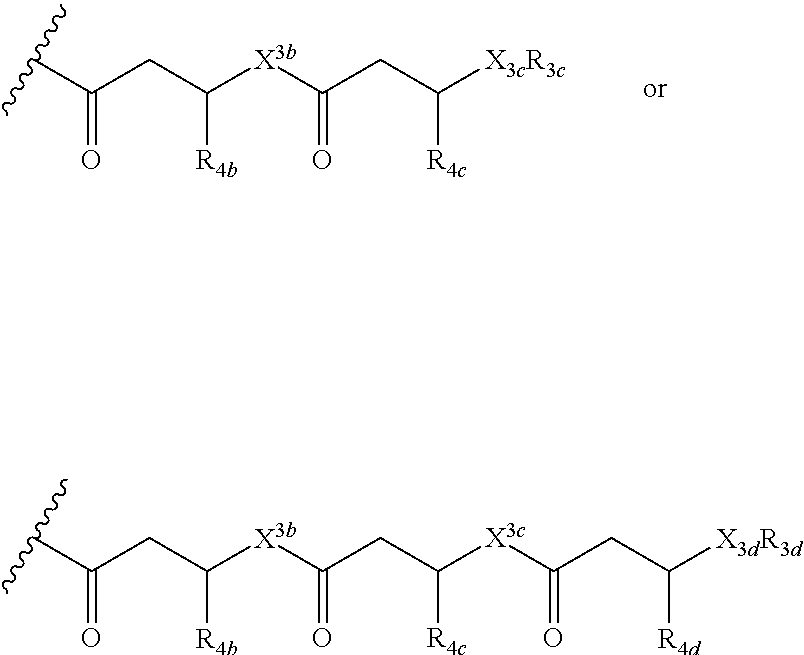
or a lipid oligomer having the structure of:
##STR00004##
wherein X.sub.3a, X.sub.3b, X.sub.3c, and X.sub.3d are each independently NH, N(CH.sub.3), or O; R.sub.3a, R.sub.3b, R.sub.3c, and R.sub.3d includes a carbohydrate portion including a monomer having the structure:
##STR00005##
wherein R.sub.5a, R.sub.6a, R.sub.7a, and R.sub.8a are each independently a hydrogen atom, methyl, acetyl, or a carbohydrate; and R.sub.4a, R.sub.4b, R.sub.4c, and R.sub.4d are each independently a hydrogen atom, methyl, or a C.sub.2 to C.sub.19 saturated or unsaturated linear, branched-chain, cyclic, or aromatic hydrocarbon groups. Naturally occurring glycolipopeptides include those having the following structures:
##STR00006##
[0007] Also described herein are emulsified compositions (e.g., oil-in-water or water-in-oil emulsions) including: a polar component, a non-polar component, and one or more of the above described biosurfactants; and a method of making an water-in-oil or oil-in-water emulsion by mixing together a polar component, a non-polar component, and one or more of the above described biosurfactants. Further described herein are a method of making one of the above described biosurfactants by [0008] (a) isolating a microorganism which includes the biosurfactant, [0009] (b) placing the microorganism in a culture under conditions that promote the synthesis of the biosurfactant, and [0010] (c) isolating the biosurfactant from the culture; and an isolated microorganism engineered to produce one of the above described biosurfactants, wherein a set of heterologous genes involved in the biosynthesis of the biosurfactant has been introduced into the microorganism.
[0011] Unless otherwise defined, all technical terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Commonly understood definitions of chemical and biological terms can be found in Rieger et al., Glossary of Genetics: Classical and Molecular, 5th edition, Springer-Verlag: New York, 1991; and A Dictionary of Chemistry, Ed. J. Daintith, 7.sup.th Ed., Oxford University Press, 2016.
[0012] As used herein, when referring to a chemical or molecule, the term "purified" means separated from components that occur with it in nature or in an artificially produced mixture. Typically, a molecule is purified when it is at least about 10% (e.g., at least 9%, 10%, 20%, 30% 40%, 50%, 60%, 70%, 80%, 90%, 95%, 98%, 99%, 99.9%, and 100%), by weight (excluding solvent), free from components that occur with it in nature or in an artificially produced mixture. Purity can be measured by any appropriate method, e.g., column chromatography, polyacrylamide gel electrophoresis, or HPLC analysis.
[0013] By "sequence identity" is meant the relatedness between two amino acid sequences or between two nucleotide sequences. Herein, the degree of identity between two amino acid sequences or two deoxyribonucleotide sequences is determined using the Needleman-Wunsch algorithm (Needleman and Wunsch, 1970, J. Mol. Biol. 48: 443-453) as implemented in the Needle program of the EMBOSS package (EMBOSS: The European Molecular Biology Open Software Suite, Rice et al., 2000, Trends in Genetics 16: 276-277; http://emboss.org), preferably version 3.0.0 or later. The optional parameters used are gap open penalty of 10, gap extension penalty of 0.5, and the EBLOSUM62 (EMBOSS version of BLOSUM62) substitution matrix for amino acid sequences or the EDNAFULL (EMBOSS version of NCBI NUC4.4) substitution matrix for nucleotide sequence. The output of Needle labeled "longest identity" (obtained using the -nobrief option) is used as the percent identity and is calculated as follows:
(Identical Amino Acid of Nucleotide Residues.times.100)/(Length of Alignment-Total Number of Gaps in Alignment).
[0014] Although methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, suitable methods and materials are described below. All patents, patent applications, and publications mentioned herein are incorporated by reference in their entirety. In the case of conflict, the present specification, including definitions will control.
[0015] In addition, the particular embodiments discussed below are illustrative only and not intended to be limiting.
BRIEF DESCRIPTION OF THE DRAWINGS
[0016] FIG. 1 is an illustration of selected HMBC (.sup.1H.fwdarw..sup.13C) and COSY correlations (bold bonds) of NB-RLP1006 and assigned fragment ions from MS/MS collision-induced dissociation of the glycolipopeptides.
[0017] FIG. 2 is a denaturing polyacrylamide gel showing purified His-tagged R1pE (A) and the UPLC-HRMS analysis of enzyme reactions in which the enzyme was incubated with NB-RLP860 and dTDP-L-rhamnose.
[0018] FIG. 3 is a schematic comparison of the V. paradoxus RKNM-096 glycolipopeptide gene cluster to homologous gene clusters identified in I. limosus DSM 16000 and J. agaricidamnosum DSM 9628. Genes encoding proteins homologous to proteins in the V. paradoxus gene cluster are indicated by arrow filling patterns. Identity and similarity to V. paradoxus proteins is indicated under arrows (identity %/similarity %). NRPS domain organization is indicated under arrows representing genes encoding non-ribosomal peptide synthetases (NRPSs). Domains: C--condensation, A--adenylation, T--thiolation/peptidyl-carrier protein, R--reductase. Subscript notation indicates putative A-domain substrate. Labels above arrows in the I. limosus and J. agaricidamnosum gene clusters indicate protein IDs.
[0019] FIG. 4 is the nucleic acid sequence SEQ ID NO:1.
[0020] FIG. 5 is the nucleic acid sequence SEQ ID NO:2.
[0021] FIG. 6 is the nucleic acid sequence SEQ ID NO:3.
[0022] FIG. 7 is the nucleic acid sequence SEQ ID NO:5.
[0023] FIG. 8 is the nucleic acid sequence SEQ ID NO:7.
[0024] FIG. 9 is the nucleic acid sequence SEQ ID NO:9.
[0025] FIG. 10 is the nucleic acid sequence SEQ ID NO:11.
[0026] FIG. 11 is the nucleic acid sequence SEQ ID NO:13.
[0027] FIG. 12 is the nucleic acid sequence SEQ ID NO:15.
[0028] FIG. 13 is the nucleic acid sequence SEQ ID NO:17.
[0029] FIG. 14 is the nucleic acid sequence SEQ ID NO:19.
[0030] FIG. 15 is the nucleic acid sequence SEQ ID NO:21.
[0031] FIG. 16 is the amino acid sequence SEQ ID NO:4.
[0032] FIG. 17 is the amino acid sequence SEQ ID NO:6.
[0033] FIG. 18 is the amino acid sequence SEQ ID NO:8.
[0034] FIG. 19 is the amino acid sequence SEQ ID NO:10.
[0035] FIG. 20 is the amino acid sequence SEQ ID NO:12.
[0036] FIG. 21 is the amino acid sequence SEQ ID NO:14.
[0037] FIG. 22 is the amino acid sequence SEQ ID NO:16.
[0038] FIG. 23 is the amino acid sequence SEQ ID NO:18.
[0039] FIG. 24 is the amino acid sequence SEQ ID NO:20.
[0040] FIG. 25 is the amino acid sequence SEQ ID NO:22.
[0041] FIG. 26 is the amino acid sequence SEQ ID NO:23.
[0042] FIG. 27 is the amino acid sequence SEQ ID NO:24.
DETAILED DESCRIPTION
[0043] The invention encompasses glycolipopeptide surfactant compositions, methods of making and using such biosurfactants, and bacteria and bacterial culture that produce glycolipopeptides. The below described preferred embodiments illustrate adaptation of these compositions and methods. Nonetheless, from the description of these embodiments, other aspects of the invention can be made and/or practiced based on the description provided below.
General Methodology
[0044] Methods involving conventional organic chemistry, biochemistry, microbiology, and molecular biology are described herein. Such methods are described in, e.g., Clayden et al., Organic Chemistry, Oxford University Press, 1st edition (2000); Molecular Cloning: A Laboratory Manual, 2nd ed., vol. 1-3, Sambrook et al., ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 2001; Current Protocols in Molecular Biology, Ausubel et al., ed., Greene Publishing and Wiley-Interscience, New York; and in the various volumes of Methods in Microbiology and Methods in Biochemistry and Molecular Biology both published by Elsevier.
Glycolipopeptides
[0045] Naturally occurring glycolipopeptides and synthetic analogues and derivatives thereof typically include a hydrophobic lipid component including a carboxyl end and a hydroxyl end, wherein the lipid component is covalently linked to (i) a peptide or peptide-like chain at the carboxyl end of the lipid component and (ii) a carbohydrate moiety at the hydroxyl end of the lipid component via a glycosidic linkage.
[0046] The peptide chain may comprise in the range of between 2 and 10 amino acids, preferably 2 to 8, more preferably 2 to 4 amino acids. The peptide chain may most preferably comprise 2 amino acids. The peptide or peptide-like chain can comprise and/or consist of a serine-leucinol dipeptide.
[0047] The lipid component may comprise in the range of between 1 and 6 alkanoic acid moieties, preferably 2 to 4, and more preferably 3. Most preferably the lipid component can include three .beta.-hydroxyalkanoic acid moieties. The length of each acyl chain of the lipid component may be in the range of between C.sub.4 to C.sub.20, preferably C.sub.6 to C.sub.16, more preferably C.sub.8 to C.sub.14. Most preferably the length of each acyl chain may be selected from C.sub.8, C.sub.10, or C.sub.12.
[0048] The carbohydrate moiety may be selected from saccharides including glucose, fructose, galactose, mannose, ribose, or deoxy saccharide variants including deoxyribose, fucose, or rhamnose. Preferably the carbohydrayte moiety is rhamnose. In particular, a rhamnose moiety attached to the lipid component via a glycosidic linkage. In certain embodiments, the carbohydrate moiety can include one, two, or three rhamnose moieties and/or an acetyl groups. Preferably the carbohydrate moiety includes two.
[0049] Glycolipopeptides can include the structure:
##STR00007##
wherein R.sub.1a is H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), N(CH.sub.3).sub.2, or a peptide or peptide-like structure having the structure:
##STR00008##
wherein R.sub.1b, R.sub.1c, and R.sub.1d, are H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), or N(CH.sub.3).sub.2; R.sub.2a, R.sub.2b, R.sub.2c, and R.sub.2d are each independently an amino acid side chain; X.sub.1a, X.sub.1b, X.sub.1c, and X.sub.1d are each independently one oxygen atom or two hydrogen atoms; X.sub.2a, X.sub.2b, X.sub.2c, and X.sub.2d are each independently NH, N(CH.sub.3), or O; R.sub.3a is a carbohydrate portion or a lipid monomer having the structure:
##STR00009##
or a lipid oligomer having the structure of:
##STR00010##
wherein X.sub.3a, X.sub.3b, X.sub.3c, and X.sub.3d are each independently NH, N(CH.sub.3), or O; R.sub.3a, R.sub.3b, R.sub.3c, and R.sub.3d includes a carbohydrate portion including a monomer having the structure:
##STR00011##
wherein R.sub.5a, R.sub.6a, R.sub.7a, and R.sub.4d are each independently a hydrogen atom, methyl, acetyl, or a carbohydrate; and R.sub.4a, R.sub.4b, R.sub.4c, and R.sub.4d are each independently a hydrogen atom, methyl, or a C.sub.2 to C.sub.19 saturated or unsaturated linear, branched-chain, cyclic, or aromatic hydrocarbon groups.
[0050] In the foregoing, at least one of R.sub.6a, R.sub.7a, and R.sub.8a can include a carbohydrate monomer having the structure:
##STR00012##
wherein R.sub.5b, R.sub.6b, R.sub.7b, and R.sub.8b are each independently a hydrogen atom, methyl, acetyl, or a carbohydrate.
[0051] In certain embodiments the peptide or peptide-like portion includes at least one proline or proline-like monomer having the structure:
##STR00013##
wherein X.sub.4 is one oxygen atom or two hydrogen atoms, or a single proline or proline-like monomer or a terminal proline or proline-like monomer having the structure:
##STR00014##
wherein R.sub.9 is of H, OH, OCH.sub.3, SH, S(CH.sub.3), NH.sub.2, NH(CH.sub.3), or N(CH.sub.3).sub.2; and X.sub.4 is one oxygen atom or two hydrogen atoms.
[0052] Glycolipopeptides can have the following structures:
##STR00015##
[0053] wherein R.sub.5a, R.sub.6a, R.sub.7a, R.sub.10, and R.sub.11 are each independently a hydrogen atom or acetyl; and n.sub.1, n.sub.2, and n.sub.3 are integers each independently ranging from 1 to 7;
##STR00016##
wherein R.sub.5a, R.sub.5b, R.sub.6b, R.sub.7a, R.sub.7b, R.sub.10, and R.sub.11 are each independently a hydrogen atom or acetyl; and n.sub.1, n.sub.2, and n.sub.3 are integers each independently ranging from 1 to 7;
##STR00017##
[0054] Derivatives, analogues, and other variants of the foregoing glycolipopeptides can be made by one of skill in the art. For instance, the amino acid composition and length of the peptide chain could be modified in a combinatorial fashion, introducing either proteinogenic or unnatural amino acids to modulate the solubility, hydrophilic-lipophilic balance (HLB), and other surfactant characteristics of the glycolipopeptides. The peptide portion may also contain amino acids with charged functional groups, which may result in cationic, anionic, or zwitterionic surfactants with unique surfactant applications. The carboxylic acid functionality at the C-terminus position of the peptide may also be reduced to a primary hydroxyl group. Similarly, the lipid portion may contain various numbers (e.g., 1, 2, 3, 4 or more) of .beta.-hydroxyalkanoate units, which themselves may be comprised of C.sub.2 to C.sub.19 saturated or unsaturated linear, branched-chain, cyclic, or aromatic hydrocarbon groups. The rhamnose moieties could be linked together via 1,2-, 1,3-, or 1,4-glycosidic linkages, which may possess either the .alpha.- or .beta.-configuration. In addition to rhamnose, the carbohydrate portion may also be composed of glucose or other monosaccharide units.
[0055] Variants of the Variovorax paradoxus RKNM-096 glycolipopeptide biosurfactants that have altered properties could be made. Altered properties of such variants may include, but are not limited to, alterations in emulsification, foaming and surface tension reducing properties exhibited under differing physiochemical conditions such as, but not limited to, temperature, pH, and salinity.
[0056] The variovaricins describe herein may be at least 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 95, 99, 99.5, 99.9, or 99.99 percent purified (by weight). They may be in crystalline or non-crystalline (amorphous) form, and in some cases also be obtained as salts derived from such organic and inorganic acids as: acetic, trifluoroacetic, lactic, citric, tartaric, formate, succinic maleic, malonic, gluconic, hydrochloric, hydrobromic, phosphoric, nitric, sulfuric, methane sulfonic and similarly known acids. The salts can be prepared by adapting commonly known procedures.
[0057] In some embodiments, the composition includes additional compounds such as carriers, other surfactants (e.g., non-glycolipopeptide surfactants), or biologically active compounds (non-glycolipopeptide surfactants, such as pharmaceutical agents or other non-glycolipopeptide antimicrobial agents). The addition of the aforementioned agents to glycolipopeptide surfactants can be selected by one skilled in the art based on the chosen application.
[0058] The composition can include a carrier, such as conventional pharmaceutically acceptable carriers as described in Remington: The Science and Practice of Pharmacy, The University of the Sciences in Philadelphia, Editors, Lippincott, Williams, & Wilkins, Philadelphia, Pa., 21.sup.st Edition (2005). Pharmaceutically acceptable carriers vary depending on the mode of administration. Fluid formulations used for parenteral injection may include fluids such as water, physiological saline, aqueous dextrose or glycerol. Solid formulations may include highly purified solid carriers such as magnesium stearate, starch, or lactose. Pharmaceutical compositions may also contain minor quantities of non-toxic auxiliary substances, such as buffers and preservatives.
[0059] In some embodiments, the compositions include a non-glycolipopeptide surfactant. Examples include non-ionic, cationic, anionic and amphoteric surfactants. Representative examples of anionic surfactants include carboxylates, sulfonates, petroleum sulfonates, alkylbenzene sulfonates, naphthalene sulfonates, olefin sulfonates, alkyl sulfates, sulfates, sulfated natural oils and fats, sulfated esters, sulfated alkanolamides, alkylphenols, ethoxylated and sulfated aklylphenols and rhamnolipids. Examples of cationic surfactants include quaternary ammonium salts, N, N, N', N' tetrakis substituted ethylenediamines and 2-alkyl-1-hydroxethyl-2-imidazolines. Examples of non-ionic surfactants include ethyoxylated aliphatic alcohols, polyoxyethylene surfactants, carboxylic esters, polyethylene glycol esters, anhydrosorbitol ester and ethoxylated derivatives, glycol esters of fatty acids, carboxylic amides, monoalkanolamine condensates and polyoxyethylene fatty acid amides. Examples of amphoteric surfactants include sodium salts of N-coco-3-aminopropionic acid, N-tallow-3-iminodipropionate and N-cocoamidethyl-N-hydroxyethylglycine, as well as N-carboxymethyl-N-dimethyl-N-(9-octadecenyl) ammonium hydroxide. In further embodiments, the composition includes one or more food or food additive, cosmetic or pharmaceutical agents or antimicrobial agents (such as an antibacterial or antifungal agents).
Methods of Making Glycolipopeptides
[0060] The glycolipopeptides described herein may be made by isolation or purification from bacteria strains which produce them, such as Variovorax paradoxus RKNM-096. As described in the Examples section below, bacteria which produce one or more glycolipopeptides can be isolated from natural habitats or obtained from publicly accessible sources. Bacteria can be determined to produce glycolipopeptides by the methods described in the Examples. The glycolipopeptide-producing bacterium can be placed in a bioreactor (vessel) containing suitable culture medium, and then incubated under conditions that promote bacterial replication and production of one or more glycolipopeptides. The produced glycolipopeptide(s) can be purified or isolated from the culture mixture by conventional techniques such as extraction followed by chromatographic separation (e.g., using ultra high performance liquid chromatography). Chemical analyses (determination of molecular weight, melting point, NMR, IS spectroscopy, etc.) can be performed to confirm the structure and purity of the isolated glycolipopeptide(s). Alternatively, the glycolipopeptides described herein may be made by total synthesis or semi-synthesis, e.g. as described herein.
Glycolipopeptides Gene Clusters and Methods of Use
[0061] As described in Example 7 below, the glycolipopeptide and rhamnose biosynthetic gene clusters of V. paradoxus RKNM-096 were characterized. The polypeptides encoded in the gene cluster function in a coordinated fashion to synthesize the NB-RLP series of biosurfactants. The nucleotide sequence encoding these genes and the amino acid sequences of the corresponding polypeptides are shown in the sequence listing. Other amino acid sequences and the nucleic acid sequences that share at least 70% (e.g., at least 70, 80, 90, 95, 97, 98, or 99%) sequence identity with those shown in the sequence listing might also be used in the methods and compositions described herein particularly when such other sequences exhibit (or encode a molecule exhibiting) at least 50% (e.g., at least 50, 60, 70, 80, 90, or 100%) of the corresponding native polypeptide enzymatic activity. Nucleic acid sequences which encode the same polypeptides described herein but are not included in the sequence listing might also be used.
[0062] The foregoing polynucleotides might be used in a method for producing recombinant biosynthetic enzymes. As one example, such a method might include culturing a host cell (e.g., E. coli or another suitable prokaryotic or eukaryotic host cell) which contains an expression vector having a nucleic acid sequence of one or more of SEQ ID NO:1, SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:5, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:11, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:17, SEQ ID NO:19 and SEQ ID NO:21 in a culture medium under conditions suitable for expression of the recombinant protein in the host cell, and b) isolating the recombinant protein(s) from the host cell or the culture medium.
[0063] Also contemplated is method of producing a glycolipopeptide in a heterologous host cell by expressing the complete or partial biosynthetic gene cluster. This method might include the steps of a) culturing a host cell which contains an expression vector having nucleic acid sequences comprising SEQ ID NO:1, SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:5, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:11, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:17, SEQ ID NO:19 and SEQ ID NO:21 in a culture medium under conditions suitable for expression of the recombinant proteins in the host cell, and b) isolating produced glycolipopeptides from the culture medium.
[0064] Further contemplated are methods for using a nucleic acid molecule that hybridizes to or includes a portion of SEQ ID NO:1, SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:5, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:11, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:17, SEQ ID NO:19 or SEQ ID NO:21 as a probe or PCR primer to identify other organisms capable of producing glycolipopeptides or structurally similar biosurfactants.
Synthesis
[0065] The compounds of the present invention may be achieved using chemical methods as noted herein.
[0066] The total synthesis of the glycolipopeptides can be achieved using established synthetic methodology to assemble commercially available building blocks. A retrosynthetic analysis of NB-RLP1006 (1) demonstrates the feasibility of the total synthesis. As an example, one skilled in the art of organic synthesis may couple the dipeptide substituent (4) to the tridecanoic acid (5) and perform a chemical glycosylation of the lipopeptide intermediate (2) using glycosyl donor 3 as shown below. The dipeptide moiety can be prepared using standard amide coupling methods, while the tridecanoic acid can be generated from commercially available ethyl trans-2-decenoate. Meanwhile, the .alpha.-1,3-linked dirhamnose substituent (3) can be assembled using glycosyl donor 6 and glycosyl acceptor 7. It is understood that this general approach, or other similar approaches in which one assembles commercially available starting materials, could enable the synthesis of glycolipopeptide analogues. For instance, different amino acids can be incorporated into the peptide or peptide-like portion while the length of the peptide chain can be increased or decreased. Similarly, structural modifications could be made to the lipid and carbohydrate portions of the glycolipopeptides to produce analogues with potentially useful biosurfactant characteristics.
[0067] To generate the carbohydrate substituent, .alpha.-1,3 linked dirhamnose, a number of protecting group manipulations must be performed to enable regioselective glycosylation of the rhamnose sugar at the 3-OH position (Scheme 2). The p-methoxyphenyl .alpha.-L-rhamnopyranoside (8), which serves as a synthetic precursor to both rhamnose moieties, can be synthesized from commercially available L-rhamnose in three steps. The terminal rhamnose sugar can then be prepared by perbenzylation of 8 and removal of the p-methoxyphenyl substituent to allow synthesis of the rhamnosyl trichloroacetimidate (9). Meanwhile, a six-step sequence of protecting group manipulations can provide p-methoxyphenyl 2,4-di-O-benzyloxyrhamnopyranoside (10) (Cai, X.; et al. Carbohydr. Res. 2010, 345, 1230), which can be glycosylated at the 3-OH position to achieve the .alpha.-1,3 glycosidic linkage between the two rhamnose substituents. The anomeric effect is expected to direct the formation of an .alpha.-glycosidic linkage with high stereoselectivity in this chemical glycosylation (Takahashi, O.; et al. Carbohydr. Res. 2007, 342, 1202).
##STR00018##
[0068] To assemble the dirhamnose substituent, glycosyl donor 9 can be linked to glycosyl acceptor 10 through activation of the anomeric trichloroacetimidate using either BF.sub.3.Et.sub.2O or TMSOTf (Scheme 3). The anomeric p-methoxyphenyl protecting group must then be replaced with a good leaving group, such as a trichloroacetimidate, to enable glycosylation of the decanoic acid moiety. Alternatively, a thiophenyl group could be installed instead of the p-methoxyphenyl group during reaction A of Scheme 2. This approach would allow an orthogonal glycosylation to be pursued given the dual role of the anomeric thiophenyl group as a protecting and leaving group (Gampe, C. M.; et al. Tetrahedron 2011, 67, 9771; Wu, C.-Y.; Wong, C.-H. Top. Curr. Chem. 2011, 301, 223). It is known that the total synthesis of NB-RLP860 could be achieved using rhamnosyl donor 9 or other suitable rhamnosyl donors. Furthermore, it is recognized that a skilled chemist could modify the carbohydrate moiety of the glycolipopeptides by using glycosyl donors other than 9, 11, or 12.
##STR00019## ##STR00020##
[0069] To assemble the tridecanoic acid moiety, commercially available ethyl trans-2-decenoate (13) can serve as a precursor to the synthesis of 3-hydroxydecanoic acid (14) through an established five-step .beta.-oxidation (Schneekloth, J. S.; et al. Bioorg. Med. Chem. Lett. 2006, 16, 3855; Pandey, S. K.; Kumar, P. Eur. J. Org. Chem. 2007, 369). Overall yields reported in the literature for this synthesis vary between 50-85% (Scheme 4). Although (.+-.)-3-hydroxydecanoic acid is also commercially available, this racemic precursor is cost-prohibitive and does not likely represent an economically viable route. Upon generating the 3-(tert-butyldimethylsilyl) decanoic acid (15), this building block can be linked twice to 14 via Steglich esterification to generate the silylated di- and tri-decanoic acid (16 and 17, respectively) of NB-RLP1006 and other glycolipopeptides. In this approach, carboxylic acid 15 is activated as a N-hydroxysuccinimide ester prior to the addition of 14, obviating an additional protecting group for the carboxylic acid functionality of 14.
##STR00021##
[0070] An alternative approach is also available in which the carboxylic acid group of 14 is protected as a benzyl ester before esterification (Scheme 5). In this approach, building blocks 15 and 18 are linked together in a synthesis that requires additional steps for installing and removing silyl ether and benzyl ester protecting groups. It is known that a chemist skilled in the art of organic synthesis could utilize either approach to introduce C.sub.2 to C.sub.19 saturated or unsaturated linear, branched-chain, cyclic, or aromatic hydrocarbon moieties in order to modify the lipid portion of the glycolipopeptides. It is anticipated that analogues generated through this approach may also exhibit surfactant properties.
##STR00022##
[0071] The leucinol-serine dipeptide can be assembled from commercially available Boc-leucinol (19) and Fmoc-Ser(Bzl)-OH (20) using well-established amide coupling chemistry (Scheme 6) (Valeur, E.; Bradley, M. Chem. Soc. Rev. 2009, 38, 606). The five-step reaction sequence involves protecting the primary hydroxyl group of 19 as a benzyl ether and coupling the two amino acids before removing the 9-fluorenylmethyloxycarbonyl (Fmoc) protecting group to generate the leucinol-serine dipeptide (21) that is poised for amide coupling to the decanoic acid. It is conceivable that other commercially available amino acids, including but not limited to D-amino acids and .beta.-amino acids, could be assembled in a similar fashion to introduce structural modifications at the peptide portion of the glycolipopeptide. The C-terminus of the peptide or peptide-like portion could exist as a carboxylic acid functionality or be reduced to a primary hydroxyl group. Other modifications of the C-terminus position include, but are not limited to, alkylation, acylation, glycosylation, phosphorylation, and sulfation. The chain length of the peptide or peptide-like portion could be increased by coupling additional amino acid monomers to the dipeptide intermediate. Alternatively, a single amino acid monomer could be coupled to the tridecanoic acid intermediate (17) to decrease the chain length.
##STR00023##
[0072] The tridecanoic acid (17) can readily undergo amide coupling to the benzylated dipeptide (21), upon which the tert-butyldimethyl silyl ether protecting group can be removed using tetrabutylammonium fluoride to provide glycosyl acceptor 22 (Scheme 7). Glycosylation of 22 with either 11 or 12, followed by global deprotection via hydrogenolysis of the benzyl ethers, furnishes the deprotected glycolipopeptide NB-RLP1006.
##STR00024##
[0073] Although the total synthesis of NB-RLP1006 may require between 14-18 steps (longest linear sequence, 34-40 steps total), the synthesis could be expedited by utilizing solid-phase synthetic techniques in which the terminal leucinol residue is immobilized onto a solid support. For example, the Leucinol(Bzl) (19) can be tethered to a polystyrene-bound p-alkoxybenzyl hydroxyl group (Wang resin) through a silyl ether linkage (Scheme 8) (Scott, P. J. H. Linker Strategies in Solid-Phase Synthesis, John Wiley & Sons Ltd: Chichester, U.K., 2009; pp 50-51). Following previously described amide coupling and Steglich esterification methodologies (Coin, I.; et al. Nat. Protoc. 2007, 2, 3247.), the remaining serine and decanoic acid residues can then be attached in a step-wise approach using Fmoc-Ser(Bzl)-OH (20) and 3-(tert-butyldimethylsilyl)decanoic acid (23). After releasing the lipopeptide intermediate, the primary hydroxyl group can be selectively protected as a tert-butyldiphenylsilyl ether to provide glycosyl acceptor 24. The glycolipopeptide NB-RLP1006 can then be synthesized by chemical glycosylation and removal of the silyl and benzyl ether protecting groups. Analogues of the glycolipopeptides can also be produced using solid-phase synthetic techniques as described in the foregoing solution-phase synthesis of NB-RLP1006.
##STR00025## ##STR00026##
Semisynthesis of the Glycolipopeptides
[0074] Synthetic analogues of NB-RLP1006 and other glycolipopeptides may also be of interest for assessing the structure-activity relationships of this class of biosurfactants. Unlike the total synthesis, a semisynthesis could represent a rapid approach for developing a number of glycolipopeptide analogues. For instance, strategies may involve a semisynthesis of the tridecanoic acid (23) by acid hydrolysis of the glycolipopeptide mixture (Scheme 9). See Miao, S.; et al. J. Agric. Food Chem. 2015, 63, 3367. Tridecanoic acid (23) could then be coupled to the peptide portion and glycosylated with commercially available disaccharides, such as lactose or maltose, to generate novel glycolipopeptide analogues (e.g. 24). The aglycone of glycolipopeptides may also be produced by V. paradoxus RKNM-096 and undergo chemical glycosylation to produce similar analogues. It is also known that the rhamnolipids could be utilized as an advanced precursor and linked to various dipeptides (e.g. 21) through the carboxylic acid functional group to produce glycolipopeptides similar to NB-RLP1006 (Scheme 10). Given the commercial availability of the rhamnolipids, conceivably one skilled in the art of organic synthesis would also recognize that peptide chains other than leucinol-serine could be introduced to expand on the structural diversity of glycolipopeptide analogues accessible through this semisynthetic approach.
##STR00027## ##STR00028##
##STR00029##
[0075] Conceivably one skilled in the art of organic synthesis could isolate naturally occurring glycolipopeptides from a microbial fermentation and synthesize derivatives, analogues, and other structural variants. For instance, modifications that could occur at R.sub.5a, R.sub.5b, R.sub.6a, R.sub.6b, R.sub.7a, R.sub.7b, R.sub.10, and R.sub.11 include, but are not limited to, alkylation, acylation, glycosylation, phosphorylation, and sulfation. The glycolipopeptides could also undergo a base hydrolysis to produce rhamnolipid-like compounds with potentially useful surfactant properties. It is also known that a base hydrolysis reaction would provide NB-RLP374.
Methods of Use
[0076] The glycolipopeptides described herein might be used similarly to other surfactants. They may, for example, be used as detergents, emulsifiers, dispersants, wetting agents, foaming agents, or biofilm inhibitors/disruptors. A typical use would be for the preparation of emulsions for cosmetic or pharmaceutical formulations (eg., water-in-oil or oil-in-water emulsions), where one or more glycolipopeptides or derivatives or analogues thereof is mixed with a polar component and a non-polar component.
[0077] The properties of the surfactants of this invention also make them suitable as emulsifiers particularly in oil in water or water-in-oil emulsions e.g. in personal care applications. Personal care emulsion products can take the form of creams and milks desirably and typically include emulsifier to aid formation and stability of the emulsion. Typically, personal care emulsion products use emulsifiers (including emulsion stabilisers) in amounts of about 3 to about 5% by weight of the emulsion.
[0078] The oil phase of such emulsions are typically emollient oils of the type used in personal care or cosmetic products, which are oily materials which is liquid at ambient temperature or solid at ambient temperature, in bulk usually being a waxy solid, provided it is liquid at an elevated temperature, typically up to 100.degree. C. more usually about 80.degree. C., so such solid emollients desirably have melting temperatures less than 100.degree. C., and usually less than 70.degree. C., at which it can be included in and emulsified in the composition.
[0079] The concentration of the oil phase may vary widely and the amount of oil is typically from 1 to 90%, usually 3 to 60%, more usually 5 to 40%, particularly 8 to 20%, and especially 10 to 15% by weight of the total emulsion. The amount of water (or polyol, e.g. glycerin) present in the emulsion is typically greater than 5%, usually from 30 to 90%, more usually 50 to 90%, particularly 70 to 85%, and especially 75 to 80% by weight of the total composition. The amount of surfactant used in such emulsions may be in the range from 0.001 to 10% by weight of the emulsio, preferably 0.01 to 6% by weight, more preferably 0.1 to 5% by weight, further preferably 1 to 3% by weight. The amount of surfactant used on such emulsions is typically from 2 to 5.5%, by weight of the emulsion.
[0080] The end uses formulations of such emulsions include moisturizers, sunscreens, after sun products, body butters, gel creams, high perfume containing products, perfume creams, baby care products, hair conditioners, skin toning and skin whitening products, water-free products, anti-perspirant and deodorant products, tanning products, cleansers, 2-in-1 foaming emulsions, multiple emulsions, preservative free products, emulsifier free products, mild formulations, scrub formulations e.g. containing solid beads, silicone in water formulations, pigment containing products, sprayable emulsions, colour cosmetics, conditioners, shower products, foaming emulsions, make-up remover, eye make-up remover, and wipes. A preferred formulation type is a sunscreen containing one or more organic sunscreens and/or inorganic sunscreens such as metal oxides, but desirably includes at least one particulate titanium dioxide and/or zinc oxide.
[0081] All of the features described herein may be combined with any of the above aspects, in any combination. It is to be understood that the invention is not to be limited to the details of the above embodiments, which are described by way of example only. Many variations are possible.
[0082] In order that the present invention may be more readily understood, reference will now be made, by way of example, to the following description.
Examples
Example 1: Isolation of Variovorax paradoxus RKNM-096
[0083] Bacterial strain RKNM-096 was isolated from soil collected from the Battle Bluffs area west of Kamloops, British Columbia. RKNM-096 was isolated as a mucoid, yellow pigmented colony, and purified by serial subculturing. The bacterium was identified by 16S rRNA gene analysis, which indicated that RKNM-096 was a strain of V. paradoxus.
Example 2: Identifying Variovorax paradoxus RKNM-096 as a Biosurfactant Producer
[0084] V. paradoxus RKNM-096 was identified as a biosurfactant producer in a screen aimed at identifying bacterial producers of biosurfactants with emulsifying properties. The assay utilized to identify bacterial producers of biosurfactants was the emulsification activity assay. In this assay cultures were grown in 10 mL of liquid medium in 25 mm.times.150 mm glass tubes at 30.degree. C. with shaking at 200 rpm for 5 days. After 5 days, the cells were removed by centrifugation and 3.5 mL of cell free culture broth was mixed with 3.5 mL of kerosene in a 13 mm.times.100 mm test tube with a screw cap tube. The tubes were vortexed for two minutes and then allowed to stand overnight at room temperature after which the height of the emulsion (h.sub.emuls) and the total height (h.sub.total) of the liquid in the tube were measured. The emulsification index (E.sub.24) was calculated using the equation E.sub.24=h.sub.emuls/h.sub.total.times.100%. Fermentation broths of V. paradoxus RKNM-096 cultured in ISP2 broth (0.4% maltose, 0.4% yeast extract, 1.0% dextrose, pH 7.0) exhibited an E.sub.24 value of 50.7%.
Example 3: Identification of Glycolipopeptide Biosurfactants Produced by Variovorax paradoxus RKNM-096
[0085] To determine if a small molecule was responsible for the observed emulsification activity, V. paradoxus RKNM-096 was fermented in ISP2 broth as described above and the broth was extracted twice with 10 mL of ethyl acetate (EtOAc). The EtOAc extract was then washed twice with 10 mL of water to remove any remaining polar media components from the EtOAc extract. For comparison purposes an ISP2 media blank was extracted in an identical manner. The EtOAc extracts were evaporated in vacuo and reconstituted in CH.sub.3OH at a concentration of 0.5 mg/mL.
[0086] The extracts were separated by ultra high performance liquid chromatography (UPLC; Accela.TM. Thermo Fisher Scientific Mississauga, ON, Canada) and the eluates analyzed with a photodiode array detector (200-600 nm) (PDA; Accela.TM., Thermo Fisher Scientific Mississauga, ON, Canada), an evaporative light scattering detector (ELSD; Sedex, Sedere, Alfortville, France) and a high resolution mass spectrometer utilizing electrospray ionization (HRESIMS) (Orbitrap Exactive; Thermo Fisher Scientific, Mississauga, ON, Canada) (positive mode, monitoring m/z 200-2000). Chromatographic separation was achieved with a Kinetex 1.7 .mu.m C.sub.18 100 .ANG. 50.times.2.1 mm column (Phenomenex, Torrance, Calif., USA) and a linear gradient from 95% H.sub.2O/0.1% formic acid (FA) (solvent A) and 5% acetonitrile (CH.sub.3CN)/0.1% FA (solvent B) to 100% solvent B over 5 min followed by a hold of 100% solvent B for 3 min with a flow rate of 400 .mu.L/min. Examination of the ELSD chromatogram of the V. paradoxus RKNM-096 extract revealed five prominent peaks. The first peak eluted at 0.50 min and was present in the media blank indicating this peak was composed of media components. The following four peaks (1-4) eluted at 3.0 min, 5.04 min, 5.29 min and 5.39 min in the ELSD chromatogram, respectively. These peaks were not observed in the media blank extracts, indicating that these peaks were metabolic products of V. paradoxus RKNM-096. Peak 1 eluted at 3.00 min and examination of the mass spectrum of the corresponding peak in the total ion chromatogram (3.04 min) revealed the presence of two ions with mass to charge ratios (m/z) of 375.2855 and 397.2673, which is consistent with the anticipated [M+11].sup.+ and [M+Na].sup.+ for a compound with a molecular formula of C.sub.19H.sub.38N.sub.2O.sub.5 and mass of 374.2781. The mass spectra of peaks 2-4 were examined in an identical manner and the [M+H].sup.+ ions were identified as m/z 1007.6628, 1049.6778, and 1049.6734, respectively. The difference in mass between the [M+H].sup.+ ions associated with peaks 3 and 4 was 4.2 ppm, suggesting that these two compounds likely had an identical molecular formula, however the slight difference in retention time indicated that they were probably closely related structural analogues.
[0087] The compounds were also elucidated using NMR. The NMR data indicated the presence of four carbonyl groups in addition to two sugar residues with characteristic anomeric carbon chemical shifts at .delta..sub.C 101.4 and .delta..sub.C 103.9. Key COSY and HMBC correlations allowed the chemical characterization of the amino acid-derived leucinol, a serine residue, and three 3-hydroxydecanoic acids (FIG. 1). The connectivity between the different moieties was further confirmed by tandem mass spectrometry. The two deoxyhexose residues were identified by interpretation of .sup.1H-.sup.1H COSY correlations and coupling constant analysis. The small J-coupling exhibited by the anomeric proton H-1' (.delta..sub.H 4.79, d, J=1.4 Hz) and the methine proton H-2' (.delta..sub.H 3.86, dd, J=3.2, 1.4 Hz) placed protons H-1' and H-2' in the equatorial position, while the larger J-coupling for H-4' (.delta..sub.H 3.53-3.48, app. t) indicated the axial relationship with H-3' and H-5', and therefore suggested an .alpha.-rhamnopyranosyl residue. The HMBC cross peak between the anomeric proton H-1' and C-3C (.delta..sub.C 76.5) demonstrated the attachment of this sugar to the 3-hydroxydecanoic acid moiety. The second sugar residue was also identified as an .alpha.-rhamnopyranose on the basis of coupling constant values. The small J-coupling for H-1'' (6H 5.01, d, J=1.5 Hz) and H-2'' (.delta..sub.H 3.98, dd, J=3.3, 1.5 Hz) indicated the equatorial orientation of these protons, while the larger coupling constant for H-4'' OH 3.40, app. t, J=9.5 Hz) demonstrated its axial relationship with H-3'' and H-5''. A key HMBC correlation between H-3' and C-1'' established a 1,3-.alpha.-glycosidic linkage between the two rhamnopyranose moieties.
[0088] The structure of NB-RLP1006 is:
##STR00030##
[0089] Organic extracts from V. paradoxus RKNM-096 were also fractionated by automated normal-phase chromatography followed by reversed-phase HPLC, which provided NB-RLP1048A and NB-RLP1048B. On the basis of HRESIMS analysis (NB-RLP1048A: HRESIMS m/z 1049.6778 [M+H].sup.+; NB-RLP1048B: HRESIMS 1049.6734 [M+H].sup.+, calcd for C.sub.53H.sub.97N.sub.2O.sub.18, 1049.6731), these compounds were determined to be mono-acetylated analogues of NB-RLP1006. The apparent molecular formula of these compounds is C.sub.53H.sub.96N.sub.2O.sub.18. Based on NMR analysis, NB-RLP1048A consisted of an inseparable mixture of acetylated glycolipopeptides with the structure:
##STR00031##
where any single R-group is an acetyl group, while all other R-groups are hydrogen atoms.
[0090] The chemical structure of NB-RLP1048B was determined by 1D and 2D NMR spectroscopic techniques, confirming the location of the acetyl group at the C-3'' position.
[0091] The chemical structure of NB-RLP1048B is:
##STR00032##
[0092] The described fractionation scheme also yielded several other glycolipopeptide analogues produced by V. paradoxus RKNM-096 in smaller quantities, including 10.9 mg of NB-RLP978. The .sup.1H and .sup.13C NMR data were nearly identical to that of NB-RLP1006. The apparent molecular formula of NB-RLP978 is C.sub.49H.sub.90N.sub.2O.sub.17 (HRESIMS m/z 979.6307 [M+H].sup.+, calcd for C.sub.49H.sub.91N.sub.2O.sub.17, 979.6312). On the basis of tandem mass spectrometry, NB-RLP978 was determined to be an inseparable mixture of three closely related analogues, NB-RLP978A, NB-RLP978B, and NB-RLP978C, containing a C.sub.8 acyl chain at one of the 3-hydroxyalkanoic acid positions.
[0093] The chemical structure of NB-RLP978A-C is:
##STR00033##
where any single acyl chain is C.sub.8 (i.e. n.sub.1, n.sub.2, or n.sub.3=3) while the remaining acyl chains are C.sub.10 (i.e. n=5). NB-RLP978A: n.sub.1=n.sub.2=5, n.sub.3=3; NB-RLP978B: n.sub.1=n.sub.3=5, n.sub.2=3; NB-RLP978C: n.sub.1=3, n.sub.2=n.sub.3=5.
[0094] The reversed-phase HPLC purification of NB-RLP1006 and NB-RLP978A-C also yielded NB-RLP950, an inseparable mixture of compounds with an apparent molecular formula of C.sub.47H.sub.86N.sub.2O.sub.18 (HRESIMS m/z 951.5982 [M+H].sup.+, calcd for C.sub.47H.sub.87N.sub.2O.sub.18, 951.5999), which is consistent with an analogue of NB-RLP1006 lacking four methylene groups. The .sup.1H NMR spectrum of NB-RLP950 was nearly identical to that of NB-RLP1006 and NB-RLP978. A .sup.13C spectrum was not obtained due to insufficient material. On the basis of tandem mass spectrometry, NB-RLP950 was determined to be a mixture of six closely related analogues: NB-RLP950A, NB-RLP950B, NB-RLP950C, NB-RLP950D, NB-RLP950E, and NB-RLP950F. These glycolipopeptide analogues either contain two C.sub.8 acyl chains or one C.sub.6 acyl chain at the 3-hydroxyalkanoic acid positions.
[0095] The chemical structure of NB-RLP950A-F is:
##STR00034##
where any two acyl chains are C.sub.8 (e.g. n.sub.1=n.sub.2=3 and n.sub.3=5) while the remaining acyl chain is C.sub.6 (i.e. n=1). NB-RLP950A: n.sub.1=5, n.sub.2=n.sub.3=3; NB-RLP950B: n.sub.2=5, n.sub.1=n.sub.3=3; NB-RLP950C: n.sub.3=5, n.sub.1=n.sub.2=3; NB-RLP950D: n.sub.1=n.sub.2=5, n.sub.3=1; NB-RLP950E: n.sub.1=n.sub.3=5, n.sub.2=1; NB-RLP950F: n.sub.2=n.sub.3=5, n.sub.2=1.
[0096] The reversed-phase HPLC purification of NB-RLP1048B also yielded NB-RLP1020. The .sup.1H and .sup.13C NMR data of NB-RLP1020 were nearly identical to that of NB-RLP1048B. The apparent molecular formula of NB-RLP1020 is C.sub.51H.sub.92N.sub.2O.sub.18 (HRESIMS m/z 1021.6415 [M+H].sup.+, calcd for C.sub.51H.sub.93N.sub.2O.sub.18, 1021.6418). On the basis of tandem mass spectrometry, NB-RLP1020 was determined to be an inseparable mixture of three closely related analogues, NB-RLP1020A, NB-RLP1020B, and NB-RLP1020C, comprising a C.sub.8 acyl chain at one of the 3-hydroxyalkanoic acid positions.
[0097] The chemical structure of NB-RLP 1020A-C is:
##STR00035##
[0098] where any single acyl chain is C.sub.8 (i.e. n.sub.1, n.sub.2, or n.sub.3=3) while the remaining acyl chains are C.sub.10 (i.e. n=5). NB-RLP1020A: n.sub.1=n.sub.2=5, n.sub.3=3; NB-RLP1020B: n.sub.1=n.sub.3=5, n.sub.2=3; NB-RLP1020C: n.sub.1=3, n.sub.2=n.sub.3=5.
[0099] The reversed-phase HPLC fractionation also yielded an inseparable mixture of compounds with an apparent molecular formula of C.sub.51H.sub.92N.sub.2O.sub.18 (HRESIMS m/z 1021.6477 [M+H].sup.+, calcd for C.sub.51H.sub.93N.sub.2O.sub.18, 1021.6418). Similar to NB-RLP1020A-C, the structure of these compounds is:
##STR00036##
where any single R-group is an acetyl group, while all other R-groups are hydrogen atoms and where any single acyl chain is C.sub.8 (i.e. n.sub.1, n.sub.2, or n.sub.3=3) while the remaining acyl chains are C.sub.10 (i.e. n=5).
[0100] The reversed-phase HPLC fractionation also yielded NB-RLP1076. The .sup.1H and .sup.13C NMR data were nearly identical to that of NB-RLP1020A-C and NB-RLP1048B. The apparent molecular formula of NB-RLP1076 is C.sub.55H.sub.100N.sub.2O.sub.18 (HRESIMS m/z 1077.7046 [M+H].sup.+, calcd for C.sub.55H.sub.101N.sub.2O.sub.18, 1077.7044). On the basis of tandem mass spectrometry, NB-RLP1076 was determined to be an inseparable mixture of three closely related analogues, NB-RLP1076A, NB-RLP1076B, and NB-RLP1076C, comprising a C.sub.12 acyl chain at one of the 3-hydroxyalkanoic acid positions.
[0101] The chemical structure of NB-RLP1076A-C is:
##STR00037##
where any single acyl chain is C.sub.12 (i.e. n.sub.1, n.sub.2, or n.sub.3=7) while the remaining acyl chains are C.sub.10 (i.e. n=5). NB-RLP1076A: n.sub.1=n.sub.2=5, n.sub.3=7; NB-RLP1076B: n.sub.1=n.sub.3=5, n.sub.2=7; NB-RLP1076C: n.sub.1=7, n.sub.2=n.sub.3=5.
[0102] The reversed-phase HPLC fractionation also yielded an inseparable mixture of compounds with an apparent molecular formula of C.sub.55H.sub.100N.sub.2O.sub.18 (HRESIMS m/z 1077.7098 [M+H].sup.+, calcd for C.sub.55H.sub.101N.sub.2O.sub.18, 1077.7044). Similar to NB-RLP1076A-C, the structure of these compounds is:
##STR00038##
where any single R-group is an acetyl group, while all other R-groups are hydrogen atoms and where any single acyl chain is C.sub.12 (i.e. n.sub.1, n.sub.2, or n.sub.3=7) while the remaining acyl chains are Cu) (i.e. n=5).
[0103] Using portions of the V. paradoxus RKNM-096 glycolipopeptide biosurfactant biosynthetic gene cluster as in silico probes against published bacteria genomes (described below), we identified Janthinobacterium agaricidamnosum DSM 9628 as a potential producer of glycolipopeptide biosurfactants similar to those isolated from V. paradoxus RKNM-096. J. agaricidamnosum was cultured and extracted as described above for V. paradoxus RKNM-096 and the resulting organic extract (110.4 mg) of was subjected to automated reversed-phase chromatography with a RediSep C.sub.18 column using a H.sub.2O/CH.sub.3OH gradient. Fractions containing the glycolipopeptide (77.6 mg) were combined and a portion of this material was subjected to further separation by reversed-phase HPLC, which yielded 17.1 mg of NB-RLP860 and 6.4 mg of NB-RLP832. Analysis of NB-RLP860 by HRESIMS (HRESIMS m/z 861.6033 [M+H].sup.+, calcd for C.sub.45H.sub.85N.sub.2O.sub.13, 861.6046) indicated an apparent molecular formula of C.sub.45H.sub.84N.sub.2O.sub.13 and five degrees of unsaturation. The .sup.1H and .sup.13C NMR data of NB-RLP860 were similar to NB-RLP1006, except the NMR spectra lacked resonances belonging to the second .alpha.-rhamnopyranose moiety.
[0104] The chemical structure of NB-RLP860 was determined by 1D and 2D NMR spectroscopy. The structure of NB-RLP860 is:
##STR00039##
[0105] Analysis of NB-RLP832 by HRESIMS (HRESIMS m/z 833.5734 [M+H].sup.+, calcd for C.sub.45H.sub.81N.sub.2O.sub.13, 833.5733) indicated an apparent molecular formula of C.sub.43H.sub.80N.sub.2O.sub.13. The .sup.1H and .sup.13C NMR data were nearly identical to that of NB-RLP860. On the basis of tandem mass spectrometry, NB-RLP832 was determined to be an inseparable mixture of three closely related analogues, NB-RLP832A, NB-RLP832B, and NB-RLP832C, comprising a C.sub.8 acyl chain at one of the 3-hydroxyalkanoic acid positions.
[0106] The chemical structure of NB-RLP832A-C is:
##STR00040##
where any single acyl chain is C.sub.8 (i.e. n.sub.1, n.sub.2, or n.sub.3=3) while the remaining acyl chains are C.sub.10 (i.e. n=5). NB-RLP832A: n.sub.1=n.sub.2=5, n.sub.3=3; NB-RLP832B: n.sub.1=n.sub.3=5, n.sub.2=3; NB-RLP832C: n.sub.1=3, n.sub.2=n.sub.3=5.
[0107] Glycolipopeptides NB-RLP860 and NB-RLP832A-C were also detected in small quantities in organic extracts of V. paradoxus RKNM-096 by LC-MS analysis. Analysis of HRESIMS chromatograms revealed [M+H].sup.+ ions of m/z 861.6073 and m/z 833.5749, which are consistent with the predicted m/z of [M+H].sup.+ ions for NB-RLP860 (calcd for C.sub.45H.sub.85N.sub.2O.sub.13, m/z 861.6046 [M+H].sup.+) and NB-RLP832A-C (calcd for C.sub.45H.sub.81N.sub.2O.sub.13, m/z 833.5733 [M+H].sup.+).
[0108] Analysis of organic extracts of V. paradoxus RKNM-096 also revealed three peaks in the HRESIMS chromatogram exhibiting [M+H].sup.+ ions of m/z 903.6213, which is consistent with the predicted [M+H].sup.+ ions for an acetylated analogue of NB-RLP860 (m/z 903.6152 [M+H].sup.+). As these compounds were produced in small quantities, attempts to determine their structures unambiguously by NMR spectroscopy were prohibited. These compounds were not detected in organic extracts from J. agaricidamnosum DSM 9628. Given the observed fragment ions of m/z 715.5480 (b) and 598.4310 (bf), these compounds were identified as acetylated glycolipopeptides NB-RLP902 with the structure:
##STR00041##
where any single R-group is an acetyl group, while all other R-groups are hydrogen atoms.
[0109] Fractions generated by automated reversed-phase chromatography of organic extracts from V. paradoxus RKNM-096 were enriched with NB-RLP902. Also detected in the HRESIMS chromatograms of these fractions was a small peak exhibiting a [M+H].sup.+ ion of m/z 875.5888, which is consistent with an analogue of NB-RLP902 lacking two methylene groups. This [M+H].sup.+ ion was not observed in organic extracts from J. agaricidamnosum DSM 9628. The observed fragment ion of 687.5164 (b) indicates that this compound is also a glycolipopeptide. Similar to NB-RLP978A-C, NB-RLP1020A-C, and NB-RLP832A-C, it is proposed that this peak is comprised of three compounds NB-RLP874A-C with the structure:
##STR00042##
where any single R-group is an acetyl group, while all other R-groups are hydrogen atoms and where any single acyl chain is C.sub.8 (i.e. n.sub.1, n.sub.2, or n.sub.3=3) while the remaining acyl chains are C.sub.10 (i.e. n=5).
Example 4: Deacetylation of NB-RLP1048A and Other Acetylated Glycolipopeptide Biosurfactants Produced by Variovorax paradoxus RKNM-096
[0110] It is known that the relative amount of NB-RLP1006 and acetylated glycolipopeptides (e.g. NB-RLP1048A) produced by V. paradoxus RKNM-096 may vary between batches using different culture media and fermentation conditions. As a result, the surfactant properties of the extracted glycolipopeptide product may also vary. As product consistency is important to be competitive in the biosurfactant industry, a method to selectively remove the acetate from R.sub.5a, R.sub.5b, R.sub.6a, R.sub.6b, R.sub.7a, R.sub.1b, R.sub.10, and R.sub.11 was developed to generate a consistent glycolipopeptide product comprised of NB-RLP1006 with >95% purity by weight (Scheme 1). The method utilizes NaOH within a narrow concentration range to selectively remove acetate moieties without inducing further hydrolysis of the amide, ester, or glycosidic linkages of the glycolipopeptide. The NaOH concentration and reaction solvent both have a demonstrated role in controlling the extent of hydrolysis and achieving selectively. Optimal NaOH concentrations are directly proportional to the concentration and composition of the glycolipopeptides in the reaction medium. Reaction solvents with higher water composition, such as H.sub.2O:acetone (9:1), showed better selectivity and minimized the hydrolysis of the ester linkages between the .beta.-hydroxyalkanoic acid moieties.
##STR00043##
[0111] It is known that deacetylation of the glycolipopeptide mixture may be achieved with variations to the method described herein. It is possible that inorganic bases other than NaOH, including but not limited to LiOH, KOH, Na.sub.2CO.sub.3, NH.sub.3, and NH.sub.4OH, or organic bases, including but not limited to tetrabutylammonium hydroxide or alkylamines, may be utilized. The selective deacetylation may also be achieved enzymatically using esterases, including but not limited to acetylesterases and lipases.
[0112] Hydrolysis of the glycolipopeptide mixture is known to produce several products, including but not limited to the lipopeptides NB-RLP356 (HRESIMS m/z 357.2745 [M+H].sup.+, calcd for C.sub.19H.sub.37N.sub.2O.sub.4, 357.2748; m/z 379.2567 [M+H].sup.+, calcd for C.sub.19H.sub.36N.sub.2O.sub.4Na, 379.2565), NB-RLP374 (HRESIMS m/z 375.2851 [M+H].sup.+, calcd for C.sub.19H.sub.39N.sub.2O.sub.5, 375.2854), and NB-RLP526 (HRESIMS m/z 527.4054 [M+H].sup.+, calcd for C.sub.29H.sub.55N.sub.2O.sub.6, 527.4055), and the glycolipids NB-RLP480 (HRESIMS m/z 481.2599 [M+H].sup.+, calcd for C.sub.22H.sub.41O.sub.11, 481.2643; m/z 503.2465 [M+Na].sup.+, calcd for C.sub.22H.sub.40O.sub.11Na, 503.2463) and NB-RLP650 (HRESIMS m/z 651.3962 [M+H.sub.]+, calcd for C.sub.32H.sub.59O.sub.13, 651.3950). Given their amphiphilic structures, these compounds are also expected to behave as surface active agents and may exhibit surfactant properties that may be unique or complementary to the glycolipopeptides. These compounds are known to be formed during the deacetylation process described herein and are thus present in the glycolipopeptide final product. Although normally present in small quantities (<5% by weight), these compounds may contribute to the surfactant characteristics of the glycolipopeptide product. Hydrolysis of the glycolipopeptides may also occur spontaneously, for instance during the extraction and purification, to generate these compounds. For instance, the lipopeptide NB-RLP374 is detected in the organic extract of V. paradoxus RKNM-096 before the glycolipopeptide material is subjected to any downstream modification.
[0113] The chemical structure of NB-RLP356 is:
##STR00044##
[0114] The chemical structure of NB-RLP374 is:
##STR00045##
[0115] The chemical structure of NB-RLP526 is:
##STR00046##
[0116] The chemical structure of NB-RLP480 is:
##STR00047##
[0117] The chemical structure of NB-RLP650 is:
##STR00048##
Example 5: Surface Activity
[0118] As summarized in Table 1, the critical micelle concentrations (CMCs) of NB-RLP1006, NB-RLP978, NB-RLP860, and NB-RLP-1048B were determined by the Du Nouy method utilizing a Kibron Delta-8 multichannel microtensiometer (Kibron Inc., Helsinki, Finland). All samples were prepared in degassed deionized water (Millipore, Etobicoke, ON, CA) at concentrations ranging from 0 to 2.0 mM. All measurements were recorded between 24 and 25.degree. C. and performed in duplicate. The critical micelle concentration of both NB-RLP1006 and NB-RLP978 was 0.20 mM (0.02 wt %). Surface tension measurements indicated that NB-RLP1006 and NB-RLP978 were capable of reducing the surface tension of water from 72 to 35.5 mN/m at their CMC. Meanwhile, NB-RLP860 and NB-RLP1048B exhibited CMC values of 0.85 mM (0.07 and 0.09 wt %, respectively), reducing the surface tension of water to 36.2 and 36.9 mN/m, respectively. The surface activity of NB-RLP1006 was compared to rhamnolipids A and B, which were purified from a commercially available rhamnolipid mixture (R90; AGAE Technologies, Corvallis, Oreg., USA) by reversed-phase HPLC. Rhamnolipids A and B both exhibited a CMC of 0.06 mM (0.003 and 0.004 wt %, respectively) in which the surface tension of water was reduced to 28.2 and 39.0 mN/m, respectively. The higher CMC values for NB-RLP860 and NB-RLP1048B may be due to their poor aqueous solubility.
TABLE-US-00001 TABLE 1 Surfactant properties of isolated glycolipopeptides compared to rhamnolipids. Critical micelle concentration (CMC) and surface tension reduction of water are shown. Minimum CMC Surface Tension Compound (mM) (mN/m) NB-RLP1006 0.20 35.5 NB-RLP1048B 0.85 36.9 NB-RLP860 0.85 36.2 NB-RLP978 0.20 35.5 Rhamnolipid A 0.06 28.2 Rhamnolipid B 0.06 39.0
[0119] The characteristic curvature (Cc) of NB-RLP1006 was determined using the hydrophilic-lipophilic difference-net average curvature (HLD-NAC) model to calculate the shift in chemical potential when NB-RLP1006 is transferred from the oil to the aqueous phase as a function of salinity by the following general equation:
HLD=F(S)-k.times.EACN+F(A)-.varies..times..DELTA.T+Cc
where F(S) is a function of salinity, k is a coefficient equal to 0.17, EACN (effective alkane carbon number) is the number of carbons in the alkane oil phase, .alpha. is a coefficient dependent on the type of surfactant (ionic, ethoxylates, etc), and .DELTA.T is the effect of temperature. Four mixtures of NB-RLP1006 and sodium dihexyl sulfosuccinate (SDHS) were prepared with a total surfactant concentration of 1.8 mg/mL using the following NB-RLP1006/SDHS ratios: 0, 12, 24, and 40 wt % NB-RLP1006. An electrolyte scan was performed for each mixture by varying the NaCl concentration from 0 to 6.0% (w/v). Each mixture was added to an equal volume of toluene, which constituted the oil phase, and shaken vigorously. The optimal salinity (S*) was identified as the concentration of NaCl in which a Winsor Type III microemulsion was formed, wherein the separate middle phase was composed of an equal volume of oil and water. A plot of the NB-RLP1006/SDHS molar ratios versus S* was generated and Cc was calculated from the line of best fit. The Cc value for NB-RLP1006 was determined to be +5.2, a value that reflects the hydrophobic nature of this biosurfactant.
[0120] The emulsifying properties of NB-RLP1006 were determined using the emulsification index as described above. Pure NB-RLP1006 exhibited strong emulsification activity with an E.sub.24 value of 53% at 1 mg/mL in deionized water. The emulsification of NB-RLP1006 is pH-dependent with E.sub.24 values of 8, 38, and 31% at pH 3, 6, and 8, respectively. The type of emulsion formed by NB-RLP1006 (e.g. oil-in-water or water-in-oil) was determined using the drop dilution test. An emulsion was formed by vigorously mixing a 1 mg/mL solution of RLP1006 in deionized water with an equal volume of kerosene for 1 min. A portion (20 .mu.L) of the emulsion was transferred to 0.5 mL of deionized water and 0.5 mL of kerosene and dilution of the emulsion in each liquid was monitored. The emulsion formed by NB-RLP1006 was readily dispersed in the aqueous phase, indicating that the continuous phase of the emulsion was water and that an oil-in-water (o/w) emulsion was formed by NB-RLP1006 under these conditions.
[0121] These results established that NB-RLP1006 is a potent biosurfactant capable of lowering the surface tension of water to 35.5 mN/m with a CMC comparable to that of two other well-characterized biosurfactants, rhamnolipids A and B. NB-RLP1006 also exhibits strong emulsification activity forming o/w emulsions under the conditions described herein.
Example 6: Cytotoxicity Testing of the Glycolipopeptides
[0122] To evaluate the safety profile of the glycolipopeptides, cytotoxicity testing was conducted against two normal human cell lines, BJ fibroblast cells ATCC CRL-2522 and adult epidermal keratinocytes (HEKa; Life Technologies, Carlsbad, Calif., USA). BJ fibroblasts were grown and maintained in 15 mL Eagle's minimal essential medium supplemented with fetal bovine serum (10% v/v), penicillin (100 .mu.U) and streptomycin (100 .mu.g/mL). HEKa cells were grown and maintained in 15 mL of EPI life medium (Life Technologies, Carlsbad, Calif., USA) supplemented with HKGS growth supplement (10% v/v; Life Technologies, Carlsbad, Calif., USA) and 50 .mu.g/mL gentamicin (Sigma-Aldrich, St. Louis, Mo., USA). Cells were cultured in T75 cm.sup.2 cell culture flasks and incubated at 37.degree. C. in a humidified atmosphere of 5% CO.sub.2. For BJ fibroblasts culture media was refreshed every two to three days and cells were not allowed to exceed 80% confluence. For HEKa cells growth medium was refreshed every 2 d until the cells reached 50% confluence and then the medium was refreshed every 24 h until 80% confluence was obtained. At 80% confluence, the cells were counted, diluted to 10,000 cells/well in growth medium lacking antibiotics and 90 .mu.L of cell suspension was transferred into the wells of 96-well treated cell culture plates. The plates were incubated as before to allow cells to adhere to the plates for 24 h before treatment. DMSO was used as the vehicle at a final concentration of 1%. All compounds tested were re-solubilized in DMSO and a dilution series was prepared for each cell line using the respective cell culture growth medium, 10 .mu.L of which were added to the assay wells yielding eight final concentrations ranging from 512 .mu.g/mL to 8 .mu.g/mL per well (final well volume of 100 .mu.L). The fibroblasts and HEKa cells were incubated as previously described for 24 h. All samples were tested in triplicate. Each plate contained four un-inoculated media blanks (media+1% DMSO), four untreated growth controls (media+1% DMSO+cells), and one column containing a serially diluted zinc pyrithione positive control. AlamarBlue (Life Technologies, Carlsbad, Calif., USA) was added to each well 24 h after treatment (10% v/v). Fluorescence (560/12 excitation, 590 nm emission) was monitored using a Varioskan Flash Multimode plate reader both at time zero and 4 h after the addition of alamarBlue. After subtraction of fluorescence at time zero from 4 h readings the percentage of cell viability relative to vehicle control wells was calculated. Low cytotoxic activity was displayed against the HEKa and BJ fibroblast cell lines. The observed IC.sub.50 and MIC.sub.90 values for the glycolipopeptides were significantly higher than the positive control zinc pyrithione, which served as an industry benchmark for topical antimicrobial agents (Table 2). These results indicate that the glycolipopeptides exhibit low cytotoxicity towards human skin cells and thus may be safe for use in applications which result in dermal contact such as cosmetic products.
TABLE-US-00002 TABLE 2 Cytotoxicity testing results for the glycolipopeptides. Values indicate the half maximal inhibitory concentrations (IC.sub.50) and minimum inhibitory concentration that results in 90% of growth inhibition (MIC.sub.90) in .mu.g/mL. Error is reported as standard deviation. Eukaryotic Cells HEKa HEKa BJ BJ Compound (IC.sub.50) (MIC.sub.90) (IC.sub.50) (MIC.sub.90) NB-RLP1006 15.5 .+-. 1.7 64-128 19.5 .+-. 2.4 32 NB-RLP1048B 19.3 .+-. 4.0 64-128 18.7 .+-. 1.6 32 NB-RLP860 15.5 .+-. 1.6 128 16.3 .+-. 0.3 32 Zinc pyrithione 0.20 .+-. 0.001 1 2.2 .+-. 0.3 4
Example 7: Sequencing of the V. paradoxus RKNM-096 Glycolipopeptide and Rhamnose Biosynthetic Gene Clusters
[0123] To establish the genetic basis for the biosynthesis of the novel glycolipopeptide biosurfactants described here, the genome of V. paradoxus RKNM-096 was sequenced. V. paradoxus RKNM-096 was cultured in ISP2 broth and genomic DNA was isolated using the UltraClean.RTM. Microbial DNA Isolation Kit according to the manufacturer's recommendations (Mo Bio, Carlsbad, Calif., USA). The genome was sequenced at the McGill University and Genome Quebec Innovation Centre (Montreal, QC, CA) using 2 SMRT Cells in a PacBio RSII sequencer (Pacific Biosciences, Menlo Park, Calif., USA). A total of 140, 476 raw subreads with an average length of 11,269 bp were generated and genome assembly was achieved using a HGAP workflow (Chin et al. [2013] Nature Methods 10, 563). Briefly, raw subreads were generated from raw .bas.h5 PacBio data files. A subread length cutoff value (30.times.) was extracted from subreads and used in the preassembly (BLASR) step, which consists of aligning short subreads on long subreads (Chaisson and Tesler [2012] BMC Bioinformatics 13, 238). Since errors in PacBio reads are random, the alignment of multiple short reads on longer reads enables correction of sequencing errors on long reads. These long corrected reads were then used as seeds in a subsequent assembly prepared using the Celera assembler (Myers et al. [2000] Science 287, 2196), which generates contigs. These contigs were then `polished` by aligning raw reads on contigs (BLASR) which were then processed through a variant calling algorithm (Quiver) that generates high quality consensus sequences using local realignments and PacBio quality scores (Chin et al. [2013] Nature Methods 10, 563). Over 161,717,463 bp of corrected long subreads were obtained and resulted in the assembly of two contigs. One contig contained 7,193,071 bp while the other contained 1,767 bp. The genome was annotated using the RAST server (Aziz et al. [2008] BMC Genomics 9, 75; Overbeek et al. [2014] Nucleic Acid Res. 42, D206; Brettin et al. [2015] Sci Rep. 5, 8265). The function of open reading frames (ORFS) identified by the RAST annotation were further explored by BLASTP (Altscul et al. [1997] Nucleic Acids Res. 25, 3389) and conserved domain (Marchler-Bauer and Bryant [2004] Nucleic Acids Res. 32, W327) analysis of deduced amino acid sequences.
[0124] Based on the structure of NB-RLP1006 it was hypothesized that its biosynthesis would require a NRPS to synthesize the dipeptide, one or more acyltransferases to acylate the peptide and generate the 3-(3-(3-hydroxydecanoyloxy) decanoyloxy) decanoyl moiety and one or more glycosyltransferases. Scanning the genome for genes encoding NRPSs identified two loci. One locus contained a single NRPS-encoding gene followed by two glycosyltransferases, thus this locus (12,721 bp) was analyzed further. Six ORFs were identified in this locus, which were predicted to play an integral role in glycolipopeptide biosynthesis (Table 3). The six genes, designated rlpA to rlpE, are oriented in the same direction and form a contiguous region in the V. paradoxus RKNM-096 genome.
TABLE-US-00003 TABLE 3 Deduced functions of Orfs identified in the V. paradoxus RKNM-096 glycolipopeptide (Seq. ID: 1) and dTDP-L-rhamnose biosynthetic gene clusters (Seq. ID: 2). Seq. ID. Seq. ID. Size (DNA) Source Name Start Stop (Protein) (aa) Proposed Function 3 Seq. ID: 1 rlpA 121 1035 4 304 LysR transcriptional regulator 5 Seq. ID: 1 rlpB 1437 8912 6 2491 Nonribosomal peptide synthetase 7 Seq. ID: 1 rlpC 8924 10243 8 439 dTDP-rhamnosyl transferase 9 Seq. ID: 1 rlpD 10276 10488 10 70 MbtH protein 11 Seq. ID: 1 rlpE 10497 11465 12 322 dTDP-rhamnosyl transferase 13 Seq. ID: 1 rlpF 11462 12721 14 419 MFS transporter 15 Seq ID: 2 rmlB 299 1378 16 359 dTDP-glucose 4,6-dehydratase 17 Seq ID: 2 rmlD 1375 2265 18 296 dTDP-4-dehydrorhamnose reductase 19 Seq ID: 2 rmlA 2298 3194 20 298 Glucose-1-phosphate thymidylyltransferase 21 Seq ID: 2 rmlC 3191 3736 22 181 dTDP-4-dehydrorhamnose 3,5-epimerase
[0125] Genes involved in regulation. The first gene, rlpA, encodes a protein that exhibits similarity to transcriptional regulators belonging to the LysR family. Conserved domain analysis indicated that R1pA contained an amino-terminal helix-turn-helix domain and a carboxy-terminal LysR substrate binding domain, which is consistent with the domain architecture of LysR transcriptional regulators. This family of regulators can function as transcriptional activators or repressors (Maddocks and Oyston [2009] Microbiology 154, 3609), thus it is likely that R1pA plays a role in the regulation of glycolipopeptide biosynthesis.
[0126] Genes involved in peptide biosynthesis. Following rlpA is large gene, rlpB, (7,476 bp), which encodes a NRPS. Domain analysis (Bachmann and Ravel [2009] Meth. Enzymol. 458, 181) indicated that that the NRPS consists of two modules (M1 and M2) with the following domain organization (C-A-PCP).sub.M1-(C-A-PCP-R).sub.M2. The dimodular structure and domain organization suggests that R1pB generates a dipeptide, which is consistent with structure of the V. paradoxus RKNM-096 glycolipopeptides. The first domain of the first module of R1pB is a condensation domain. The presence of a C-domain at the beginning of a NRPS initiation module is characteristic of acylated peptides. Amino-terminal C-domains can catalyze amide bond formation between the first amino acid of a peptide and a fatty acid. The fatty acid can be presented to the C-domain as an acyl-ACP intermediate, as in the case of CDA biosynthesis (Kopp et al. [2008] J. Am. Chem. Soc. 130, 2656), or an acyl-CoA intermediate, as in the case of surfactin biosynthesis (Krass et al. [2010] Chem. Biol. 17, 872). A phylogenetic analysis of the R1pB initiation module C-domain (residues 12-437) was conducted using the NaPDoS program (Ziemert et al. [2012] PLoS One 7, e34064). The R1pB domain clustered closely with C-domains from initiation modules that catalyze the condensation of a fatty acid precursor with an amino acid. The most closely related C-domain in the NaPDoS reference database was the initiation module of the bacillibactin NRPS (38% identity), which catalyzes the condensation of 2,3-dihydroxybenzoyl-ACP with glycine (May et al. [2001] J. Biol. Chem. 278, 7209). This suggests that glycolipopeptide biosynthesis starts with the condensation of a fatty acid with the first amino acid of the peptide (serine). Similar analysis of the second C-domain indicated it was most closely related to the second C-domain of the bacillibactin dimodular NRPS, DhbF (54% identity). Phylogenetic analysis revealed that the M2 C-domain of R1pB clustered with C-domains catalyzing the condensation of two L-amino acids (Ziemert et al. [2012] PLoS One 7, e34064), which is consistent with the glycolipopeptide structure.
[0127] To predict the substrate specificity of the R1pB A-domains, the substrate specificity codes were extracted from the A-domain active sites (8 residues between motifs A3 and A6) and compared to known A-domain specificity codes using the NRPS Predictive Blast tool (Bachmann and Ravel [2009] Meth. Enzymol. 458, 181). The specificity code of the M1 A-domain was most similar to A-domains from the nostopeptolide, pyoverdin, CDA and enterobactin NRPSs that activate L-serine (75-87% identity, 87-100% similarity, E-value 0.023-0.039), suggesting that L-serine is incorporated by M1. This observation is consistent with the structure of the glycolipopeptides. The M2 A-domain specificity code showed low homology (50% identity, 100% similarity, E-value 0.98) to an A-domain of the tyrocidine NRPS (TycB), which activates L-phenylalanine or L-tryptophan (Mootz and Marahiel [1997] J. Bacteriol. 179, 6843). This low level of similarity precludes prediction of the substrate specificity of this A-domain. Based on the structure of the V. paradoxus RKNM-096 glycolipopeptides the second A-domain would be expected to activate L-leucine. Comparison of the A-domain specificity code of R1pB module 2 to leucine specificity codes (Stachelhaus et al. [1999] Chem. Biol. 6, 493) also revealed low similarity, thus the R1pB M2 A-domain specificity code may represent a novel variant for leucine, although biochemical evidence would be need to establish the substrate specificity of this domain. The PCP domains of R1pB were also analyzed and both were found to contain the core PCP domain motif with an invariant serine which represents the 4'-phosphopantetheine attachment site (Konz and Marahiel [1999] Chem. Biol. 6, R39).
[0128] The final domain of R1pB is an R-domain. R-domains utilize NAD(P)H as a co-factor to reductively release PCP-bound final products as an aldehyde or alcohol (Du and Lou [2010] Nat. Prod. Rep. 27, 255). The presence of a leucinol residue at the carboxy-terminus of the glycolipopeptide dipeptide moiety is consistent with release of an acylated dipeptide intermediate by an R-domain. Collectively, the domain structure and organization of R1pB, as well as the predicted substrate specificity of the individual domains are consistent with the structure of the glycolipopeptides produced by V. paradoxus RKNM-096.
[0129] A small gene (rlpD) encoding a 70 amino acid protein that shows similarity to MbtH-like proteins was found downstream of rlpB. These proteins are often found in association with NRPSs and have been demonstrated to be essential for non-ribosomal peptide the production. (Baltz [2014] J. Ind. Microbiol. Biotechnol. 41, 357). Recently these proteins have been shown to facilitate adenylation reactions via direct interaction with A-domains (Herbst et al. [2013] J. Biol. Chem. 288, 1991). Thus we predict that R1pD interacts with one or both A-domains of R1pB to facilitate dipeptide formation.
[0130] Genes involved in glycosylation. Glycosylation of the acylated dipeptide generated by R1pB is likely catalyzed by two ORFs (rlpC and E) downstream of rlpB. The deduced amino acid sequence of rlpC (439 aa) shows similarity to the GT1 family of glycosyltransferases, which utilize activated sugars as substrates to transfer sugar moieties to a diverse array of acceptor molecules (Breton et al. [2006] Glycobiology 16, 29R). The deduced amino acid sequence of rlpE (322 aa) shows similarity to dTDP-rhamnosyltransferases. In rhamnolipid biosynthesis two glycosyltransferases are utilized to sequentially transfer two rhamnosyl units to the lipid component of rhamnolipid (Deziel et al. [2003] Microbiology 149, 2005). Rh1B transfers rhamnose from dTDP-L-rhamnose to the free .beta.-hydroxyl group of 3-(3'-hydroxydecanoyloxy)decanoic acid (HDD) to generate mono-RL, while di-RL is formed by the transfer of an additional rhamnose from dTDP-L-rhamnose to mono-RL by Rh1C (Abdel-Mawgoud et al. [2011] in Biosurfactants, Springer-Verlag, Berlin Heidelberg). The relationship between R1pC and R1pE and the Rh1B and Rh1C homologs from P. aeruginosa PAO1, B. thialandensis E264 and B. psuedomallei 1710B was investigated via the generation of a phylogenetic tree (unweighted pair group method with arithmetic mean method). In this analysis R1pC clustered with the Rh1B orthologs while R1pE clustered with the Rh1C orthologs. While R1pC clustered with the Rh1B orthologs, it did not cluster tightly as it showed limited sequence identity with these enzymes (18.6-23.1%). In contrast, R1pE shared between 39.6-40.7% identity with the Rh1C orthologs. This data suggests that R1pC and R1pE perform similar functions as Rh1B and Rh1C, respectively. We hypothesize that R1pC catalyzes the rhamnosylation of an acylated dipeptide intermediate utilizing dTDP-L-rhamnose as the carbohydrate donor. The limited sequence homology between R1pC and the Rh1B orthologs may reflect the significant difference in glycosylation substrates utilized by the enzymes. R1pE is predicted to catalyze the second glycosylation reaction, transferring rhamnose from dTDP-L-rhamnose to the R1pC reaction product.
[0131] Genes encoding dTDP-L-rhamnose biosynthesis were not found in close proximity to the glycolipopeptide gene cluster. Scanning the genome for homologs of P. aeruginosa PAO1 rhamnose biosynthetic genes (rmlBDAC) identified four genes that exhibited strong sequence similarity to those from P. aeruginosa (identity/similarity: Rm1B--79%/89%, Rm1D--60%/71%, Rm1A--78%/89%, Rm1C--66%/80%). In the V. paradoxus RKNM-096 genome the four genes are clustered and are found in the same order as in P. aeruginosa (rmlBDAC) (Rahim et al. [2000] Microbiology 146, 2803). This locus likely provides the dTDP-L-rhamnose substrates utilized by R1pC and R1pE. Modulation of expression of one or more components of the dTDP-L-rhamnose biosynthetic pathway by one skilled in the art may be an effective approach to increase glycolipopeptide yields.
[0132] Genes involved in transport. Directly downstream of rlpF, is an ORF (rlpF) encoding a protein, which is similar to major facilitator superfamily transporters from a variety of bacteria. R1pF exhibits 38% identity and 54% similarity to PA1131 from P. aeruginosa PAO1, which is immediately upstream of rhlC (Dubeau [2009] BMC Microbiol 9, 263). R1pF is likely involved in glycolipopeptide efflux.
[0133] Genes involved in the biosynthesis of the lipid moiety. In rhamnolipid biosynthesis the HDD moiety is produced by Rh1A, which condenses two .beta.-hydroxydecanoyl-ACP molecules from fatty acid biosynthesis to yield 3-(3'-hydroxydecanoyloxy)decanoic acid. Scanning of the V. paradoxus RKNM-096 genome for Rh1A homologs did not identify any proteins with significant similarity to Rh1A. Thus generation of the lipid moiety of the RKNM-096 glycolipopeptides is likely directed by a novel, yet to be identified mechanism.
[0134] Genes involved in glycolipopeptide acetylation. Acetylated analogues of NB-RLP1006 are abundant in V. paradoxus RKNM-096 fermentation broths. No genes encoding acetyltransferases were identified in the gene cluster. Thus it is likely that acetylation is catalyzed by an enzyme encoded elsewhere in the V. paradoxus RKNM-096 genome.
[0135] Proposed biosynthesis. Glycolipopeptide biosynthesis presumably starts with the formation of the 3-(3-(3-hydroxydecanoyloxy)decanoyloxy)decanoyl moiety via a yet to be identified mechanism. After formation of the lipid moiety it is likely presented to the C-domain of R1pB M1 which condenses the lipid moiety with L-serine. R1pB M2 then incorporates L-leucine to form a PCP-bound acylated dipeptide intermediate which is released from the enzyme by the C-terminal R-domain of R1pB, resulting in the formation of a terminal L-leucinol residue. dTDP-L-Rhamnose, produced by the rmlBDAC operon, is then utilized by the rhamnosyltransferases R1pC and R1pE to sequentially glycosylate the aglycone resulting in the production of the final glycosylated glycolipopeptide NB-RLP1006. NB-RLP1006 would serve as a substrate for acetylation to form NB-RLP1048A and NB-RLP1048B.
[0136] To prove the involvement of the rlpA-rplF gene cluster in the biosynthesis of glycolipopeptides in V. paradoxus RKNM-096 rlpE was expressed in E. coli and the activity of the enzyme demonstrated using NB-RPL860 as a substrate. Bioinformatics analysis indicated R1pE catalyzes the second rhamnosylation in glycolipopeptide biosynthesis, converting mono-rhamnosylated glyclipopeptides (e.g. NB-RLP832 and NB-RLP860) to di-rhamnosylated glycolipopeptides (e.g. NB-RLP978 and NB-RLP1006). The rlpE gene was cloned in pET28a (EMD Millipore, Darmstadt, DE) with an amino-terminal hexa-histidine tag using standard cloning techniques and mutation-free cloning was verified by sequencing. Due to the high GC content of rlpE, E. coli Rossetta DE3 pLysS (EMD Millipore) was chosen as the expression host as this strain expresses tRNAs for rare GC-rich codons (AGG, CCA, GGA). A single colony was used to inoculate 50 mL of LB Miller (EMD Millipore) supplemented with 50 .mu.g/mL of kanamycin (Sigma-Aldrich) and 34 .mu.g/mL of chloramphenicol (Sigma-Aldrich) and the flask was incubated at 37.degree. C. with shaking at 250 rpm overnight. Expression cultures (50 mL) were performed in LB Miller supplemented with kanamycin and chloramphenicol. These cultures were inoculated with 0.5 mL of the overnight culture and cultured at 37.degree. C. and 250 rpm until the optical density (600 nm) reached 0.5, following which IPTG was added to a final concentration of 1.0 mM to induce protein expression and the cultures were incubated at 15.degree. C. for 24 h. Cells were harvested by centrifugation (6 000.times.g for 5 min) and washed once with 20 mM Tris-HCl (pH 8.0). The cell pellet was frozen at -80.degree. C. until purification could be performed. To purify His-tagged R1pE, the cells were thawed, suspended in lysis buffer (500 mM NaCl, 5% glycerol, 1% Triton X-100, 25 mM Tris-HCl, pH 8.0) and then lysed via sonication. Cell debris and insoluble protein was removed by centrifugation at 15 000.times.g for 30 min. The supernatant was mixed with 0.5 mL of HisPur Ni-NTA resin (Thermo Fisher Scientific). The resin was washed six times with 1.0 mL of 75 mM imidazole. His-tagged R1pE was eluted with 1.0 mL of 250 mM imidazole. Four batch elutions were performed and pooled. The imidazole elution buffer was exchanged with enzyme buffer (25 mM Tris-HCl, 10% glycerol) and concentrated by centrifugal filtration using a Macrosep 3 kDa spin filter (Pall). Following concentration the enzyme was aliquoted and stored at -80.degree. C. The purity of the enzyme was analyzed by denaturing polyacrylamide gel electrophoresis (4-15% Mini-PROTEAN precast gel, 160 V, 30 min; Bio-Rad). The calculated molecular weight of His-tagged R1pE was 38.2 KDa. The apparent molecular weight of the purified protein was 33.05 kDa, which was in good agreement with the expected molecular weight (FIG. 2A).
[0137] The activity of R1pE was established by incubating the enzyme (0.1 .mu.M) in reaction buffer (25 mM Tris-HCl pH 8.0, 2.5 mM MgCl.sub.2) with 1 mM of TDP-L-rhamnose and 0.5 mM NB-RLP860. Reactions (200 .mu.L) were incubated at 30.degree. C. for 4 h. A portion (25 .mu.L) of the reaction was removed at 15 s, 1 min, 5 min, 20 min, 1 h and 4 h. The reaction was stopped by the addition of two volumes of methanol followed by flash freezing. Quenched reactions were separated by UPLC (Accela.TM., Thermo Fisher Scientific Mississauga, ON, Canada) and the eluates analyzed by HRESIMS (LTQ Orbitrap Velos; Thermo Fisher Scientific) (positive mode, monitoring m/z 200-2000). Chromatographic separation was achieved with a Hypersil Gold 1.9 .mu.m C.sub.18 175 50.times.2.1 mm column (thermo Fisher Scientific) and a linear gradient from 50% H.sub.2O/0.1% FA (solvent A) and 50% acetonitrile (CH.sub.3CN)/0.1% FA (solvent B) to 100% solvent B over 5 min followed by a hold of 100% solvent B for 3 min with a flow rate of 300 .mu.L/min. Reactions conducted with boiled enzyme showed no conversion of NB-RLP860 to NB-RLP1006. In contrast, enzyme reactions containing intact 6.times.His-R1PE resulted in the complete conversion of NB-RLP860 to NB-RLP1006 after 4 h (FIG. 2B). This data indicates that R1pE catalyzes the second rhamnosylation step in glycolipopeptide biosynthesis in V. paradoxus RKN-096. As genes for the biosynthesis of natural products in bacteria are typically clustered, this finding also provides strong evidence confirming the proposed gene cluster as the locus responsible for glycolipopeptide biosynthesis.
[0138] We also explored the ability of purified 6His-R1pE to iteratively add rhamnose units to NB-RLP860 by scanning the HRMS data for masses consistent with glycolipopeptide surfactants containing three rhamnose residues (calc'd [M+H].sup.+ 1153.7204), four rhamnose residues (calc'd [M+H].sup.+ 1299.7783) and five rhamnose residues (calc'd [M+H].sup.+ 1153.7204). Masses consistent with trirhamnosylated and tetrarhamnosylated reaction products were obtained and differed from the expected molecular weights by <1.03 parts per million (ppm) and <0.6 ppm, respectively. Interestingly, two peaks were observed for each mass, suggesting additional rhamnose units are attached at two different positions of the NB-RLP1006 structure. Relative to the production of NB-RLP1006 the tri-rhamnosylated and tetra-rhamnosylated glycolipopeptides constituted 2.99% and 0.14% of the reaction products. No penta-rhamnosylated glycolipopeptides were detected. This data indicates that recombinantly expressed R1pE can be used to generate glycolipopeptide analogs with up to four rhamnose residues. Such modifications may alter the functional properties of the glycolipopeptide. The properties can include but are not limited to wetting, foaming, surfactancy and emulsification.
[0139] Elucidation of the biosynthetic pathway for the glycolipopeptide biosurfactants produced by V. paradoxus RKNM-096 sets the stage for rational modification of the biosynthetic pathway to generate novel analogues or to increase yields. Analogues may be generated by those skilled in the art via modification of the enzymes responsible for the biosynthesis and incorporation of the lipid, peptide and carbohydrate portions of the molecule. Yields can be increased by those skilled in the art by modification of regulatory genes and or promoters, by overexpressing enzymes that represent rate limiting steps in the biosynthetic pathway or by inactivating enzymes which perform undesirable reactions. Knowledge of the biosynthetic pathway also enables expression in a heterologous host, which may enable yield improvements or the generation of glycolipopeptide analogues.
Example 8: Identification of Related Biosurfactants in Other Bacteria
[0140] Sequencing of the V. paradoxus RKNM-096 glycolipopeptide and rhamnose biosynthetic gene clusters was performed. Prior to the discovery of the glycolipopeptide series of biosurfactants and the associated biosynthetic gene cluster described herein, it would not have been possible to accurately predict the production of related glycolipopeptide biosurfactants based solely on DNA sequence analysis. Identification of the glycolipopeptide biosynthetic gene cluster now allows for targeted interrogation of microbial genomes for related gene clusters, which may have the potential to produce novel glycolipopeptide biosurfactants. As rlpC encodes a novel rhamnosyltransferase, which glycosylates an acylated dipeptide intermediate characteristic of the glycolipopeptide class of biosurfactants, we used the deduced amino acid sequence of this gene to search available bacterial genomes for homologs. This search identified homologs exhibiting to R1pC from a wide variety of bacteria. We then investigated genomic regions flanking the genes encoding the R1pC homologs for the presence of homologs of the other glycolipopeptide biosynthetic genes. Two examples will be presented to demonstrate the utility of using sequences from the glycolipopeptide gene cluster as probes to discover producers of putatively novel biosurfactants.
[0141] A homologous gene cluster was identified in the Janthinobacterium agaricidamnosum DSM 9628 genome (GenBank accession no. NZ_HG322949.1) (FIG. 3). J. agaricidamnosum is a beta-proteobacterium like V. paradoxus, but belongs to a different family. The R1pC homolog in this strain (WP_038493268.1) exhibited 68% identity to R1pC. Scanning the genome around the R1pC homolog identified other homologs of genes present in the glycolipopeptide gene cluster. Directly downstream of the R1pC homolog was an MtbH-like protein (WP_038493269.1) which shared 69% identity with R1pD. Upstream a dimodular NRPS was identified (WP)038499875.1), which showed 68% identity to R1pB and contained an identical domain organization ([C-A-PCP].sub.M1-[C-A-PCP-R].sub.M2). Active site analysis (Bachmann and Ravel [2009] Meth. Enzymol. 458, 181) indicated that the predicted substrate specificity also matched that of R1pB, with the M1 A-domain specificity code matching that for L-serine and the M2 A-domain specificity code matching that of the M2 A-domain of R1pB, indicating L-leucine is incorporated by M2 (FIG. 3). A C-domain and R-domain were also found at the amino and carboxy-termini of the J. agaricidamnosum NRPS, respectively. This suggests that biosynthesis is initiated by condensation of an acyl intermediate with serine, and terminated by reductive release of an acylated dipeptide, similar to what is predicted for glycolipopeptide biosynthesis in V. paradoxus RKNM-096. No homolog to R1pE was found in the J. agaricidamnosum DSM 9628 gene cluster, indicating that the product of the cluster likely contains a single rhamnose residue. A gene cluster with a highly similar organization to that in J. agaricidamnosum DSM 9628 was also detected in the genome of V. paradoxus DSM 21786 (GenBank accession no. NC_022247.1). Collectively, this data suggests that J. agaricidamnosum DSM9628 and V. paradoxus DSM 21786 possess the ability to produce novel biosurfactants with structures related to those produced by V. paradoxus RKNM-096. Based on the bioinformatics analysis presented here, we predict the compound(s) produced by these bacteria would be a N-acylated L-serinyl-L-leucinol dipeptide bearing a single rhamnose residue.
[0142] Genome scanning using the R1pC sequence also identified a putative biosurfactant gene cluster in the more distantly related alpha-proteobacterium Inquilinus limosus DSM 16000 (Genbank accession no. NZ_AUHM01000002.1) (FIG. 3). The R1pC homolog (WP_026869107.1) in I. limosus shared 61% identity with the V. paradoxus RKNM-096 protein. Genes encoding a MtbH-like protein (WP_026869104.1) and a NRPS (WP_026869105.1) were identified immediately upstream of the R1pC homolog. The MbtH-like protein shared 56% identity with R1pD. The NRPS was a monomodular enzyme with the following domain organization: C-A-T-R (FIG. 3). Active site analysis of the A-domain (Bachmann and Ravel [2009] Meth. Enzymol. 458, 181) indicated that L-serine is the likely substrate of this enzyme. The presence of a C-domain at the N-terminus and an R-domain at the C-terminus suggests that the product of the NRPS is an acylated serinol. An R1pE homolog (WP_034850803.1) was also detected in the I. limosus gene cluster (41% identity) suggesting that the acylated serinol intermediate may be sequentially glycosylated to yield a product bearing a dirhamnosyl moiety similar to NB-RLP1006. The final product may be exported out of the cell via the action of a MFS exporter (WP_034850806.1) which shares 52% identity with R1pF.
[0143] To validate our in silico approach to identifying producers of glycolipopeptide biosurfactants we obtained J. agaricidamnosum DSM 9628 and V. paradoxus DSM 21786 from the Deutsche Sammlung von Mikroorganismen and Zellkulturen (DSMZ) culture collection. Each strain was fermented in a variety of culture media to promote production of predicted biosurfactants. Fermentations were extracted twice with an equal volume of EtOAc. The organic layer was evaporated and the resulting concentrated extracts were analyzed by UPLC-PDA-ELSD-HRESIMS as described above for NB-RLP1006 (Example 3). Three prominent peaks eluting at 3.07, 5.05 and 5.51 min were observed in the ELSD and HRESIMS chromatograms of J. agaricidamnosum DSM 9628. The peak at 3.07 min (HRESIMS m/z 1182.6217 [M+H].sup.+, calcd for C.sub.56H.sub.85N.sub.12016, 1181.6201) could be attributed to the known compound jagaracin previously reported from this strain (Graupner et al. [2012] Angew Chem. Int. Ed. Engl. 51:13173). Extraction of the mass spectra for peaks eluting at 5.05 and 5.51 min revealed [M+H].sup.+ ions of m/z 833.5741 and m/z 861.6033, respectively. The observed [M+H].sup.+ ions showed a -1.5 and 1.0 ppm mass difference from predicted m/z [M+H].sup.+ ions for the monorhamnosyl glycolipopeptides NB-RLP832 (m/z 833.5741 [M+H].sup.+) and NB-RLP860 (m/z 861.6033 [M+H].sup.+), respectively, indicating the expected compounds had been produced by J. agaricidamnosum DSM 9628. Similar to NB-RLP978A-C and NB-RLP1020A-C, the mass of NB-RLP832 closely matched that predicted for an analogue of NB-RLP860 lacking two methylene groups. These compounds were purified and their structures elucidated using a combination of 1D and 2D NMR experiments. This analysis unambiguously confirmed that the expected monorhamnosylated biosurfactant had been produced by J. agaricidamnosum DSM 9628 (see Example 3).
[0144] Identical analysis of V. paradoxus DSM 21786 fermentation extracts also revealed the presence of a peak eluting at 5.51 min in the HRESIMS chromatogram. Inspection of the mass spectrum associated with this peak revealed the presence of a [M+H].sup.+ ion with a m/z of 861.6104, which differed from the expected mass ([M+H].sup.+ m/z 861.6046) by 5.8 ppm. The identical retention time and monoisotopic mass indicated that both J. agaricidamnosum DSM 9628 and V. paradoxus DSM 21786 produce NB-RLP860.
Sequence CWU 1
1
24112721DNAVariovorax paradoxus 1gtcgtgtctc cttcttttcg tggggtgttc
caacgggccg actgggaggt cggctgaaaa 60ccgctcgcca gtgtgcgtgc cgcaaggttt
gccttcaata aaataatcaa gctaagtaat 120atgaatggca tgcatatcga
ctcggtcgac ctcaatctgc tgcgcctgtt cgatgcggtc 180taccgcgagc
gcagcgtgag ccgcgccgcg gagtcgctgg gcctcacgca gcctgcggca
240agccatgggc tgggacggct gcggctgctt ttgaaagacg cgctcttcac
gcgtgccccc 300ggcggcgtgg cgcccacgcc gcgcgccgac cggctcgcgg
tggcggtgca ggcggcgctc 360ggcacgatcg aagcggcgct gcacgagccc
gatcgcttcg agccccaggt gtcgcgcaag 420agctttcgta ttcacatgag
cgacatcggc gaggggcgct tcctgcccgc gctgatggcg 480cggctcggcg
agctggcgcc cggcgtgcgg ctggagaccc tgccgctctt gcctgcggag
540gttgcgcccg cactcgacag cggccgcatc gatttcgcct tcggctttct
ctcgaccgtg 600cgcgacacgc agcgcacgca tcttctgaaa gaccgctaca
tcgtgctgct gcgcaagggc 660catccctttg tgaagcgccg gcgcaagggg
caggcgctgc tcgaggcgct gcaggagctc 720gactacgtgg cggtgcgcac
gcacgccgac acgctgcgca tcttgcagtt gctcaacctc 780gaagaccgcc
tgcgcctcac gaccgagcac ttcatggtgc taccggccat cgtgcgcgcc
840accgatctcg cggtggtgat gccgcgcaac atcgcgcgag ggtttgcgga
ggagggcggc 900tacgcgatcg tcgagccgcc gtttccgctg cgcgatttca
gcgtgtcgct gcactggagc 960aagcgcttcg agggcgaccc ggccaaccgt
tggttgcggc aggtgatcac ggcgctgttc 1020tccgagcgcg gctgaagttc
gaccaccaaa gtacgcgccg cgcggtgcaa gcgcgcgcga 1080ctgcgcgagt
aacacgccga gagattcccc tacagctttc tcgcccagtt gctgcatcgc
1140aacattcttt tggggtgcat gacgcgcgaa atacgatgaa agccttcgat
tccgaaagcc 1200gcgattcagg tcgcaacttc gggatgaaat ctttcgcgct
caaagacgtt cgtgaaatgt 1260tttcttccct aaaaccgtca ctgaaagtgt
tgaaaccact tgtacagtgg actggcaatg 1320tgaacggatt gttaccgcgg
agcaccggca tttctccttg agcggccgat gcacgacgcg 1380tccatttcac
gcgcacatgc atcgttgcca atttcactca agacctggag aagtgcatga
1440gtaccgtcga tcagctgggc cgcaccgccc cccttacctc ggggcagatg
gcgatgtggc 1500tcggcgcaaa gttcgcgtcg cccgacacca atttcaatct
cgccgaagcc atcgacatcg 1560caggcgagat cgaccccgcg atcttcctgg
cggccatgcg acaggtggcc gatgaagtcg 1620aggccacgcg cctgagcttc
atcgataccc cgcaagggcc acgacaggtc gtcgcgcccg 1680ttttcaccgg
cgagatcccc tacctcgacc tcagcggcga gagcgatccg caggccgagg
1740ccgagcgctg gatgcatgcg gactacaccc gcagcatcga cctcgcgcac
gggcagctgt 1800ggctgtccgc gctgatccgc ctcgcgcccg atcgccacat
ctggtaccac cgcagccatc 1860acatcgcgct cgacggcttc agcggcggcc
tcatcgcacg ccgcttcgcc gacatctaca 1920ccgcgatggt cgacaacaac
gcagcggtgc ccgaagactc gcgccttgca ccgatctcgc 1980agctggccga
cgaagaacat gcctatcgcg agtccggccg cttcccgcgc gaccgccagt
2040actggaccga gcgcttcgcc gatgcacccg atccgttgag cctcgcctcg
caccgctcgg 2100tcaacgtcgg tggcctcttg cgccagacgg tgcacctgcc
ggcggccagc gtgcaagccc 2160tgcagaccat cgcgcaagag ctcggcacca
cgctgccgca aatcctcatc gccaccaccg 2220cggcctacct gtaccgcgca
acgggcatcg aggacatggc aatcggcatc cccgtcaccg 2280cgcgccacaa
cgaccgcatg cgccgcgtgc ccgcgatggt ggccaacgcg ctgccgctgc
2340gcctggcgat gcgcgcggac ctgccgattc cggaactgat ccgcgaagtc
ggccggcaga 2400tgcggcagat cctgcggcac cagtcgtatc gctacgagca
tttgcgcagc gacctcaaca 2460tgctggtgaa caaccggcag ctcttcacca
ccgtggtcaa cgtcgagccc ttcgactacg 2520acttccgctt tgcgggccat
gccgcgaagc cgcgcaacct ctcgaacggc acggccgagg 2580acctcggcat
cttcctgtac gagcgcggca acgggcagga cctgcagatc gacttcgacg
2640ccaaccccgc ggtgcacacc gcagaggaac tggccgatca ccagcgccgg
ctgcttgcct 2700tcatcgacgc cgtgatccgc ctgccgttgc aggccgtcgg
ccagatcgac ctgctcggtg 2760ccgaagagcg gcagcaattg ctggtcgagt
ggaacgacac ggcccacgcc gtgcccgaca 2820cccatctcac cgcgttgatc
gaagcgcagc tcgcagccga tccgcaagcc atcgcattgc 2880gcttcgacgg
cgaggcgatg aacaacgaag aactgaaccg ccgcgccaac cgtctcgccc
2940acctgctgcg cgcacgcggc gctggcccgg agcgcaccgt ggcgctcgcg
atcccgcgtt 3000cgatggacct gatgattgcc ttgctcgcca cgttgaagac
cggcgcggcc tacctgccgg 3060tcgatccgga tttcccggcg gaccgcatcg
ccttcatgct cggcgatgcg cagcccgtgt 3120gcctcgtcac gaccgaagcc
ctcgcggagt cgctgccggc agccgccccc acattgctgc 3180tcgatgtagc
gcaaacgatt gcggatctgg agagttgcaa cgacaccaac ccgggcatcg
3240cgatcgaccc ttcgcatccg gcctatgtga tctacacctc gggctcgacc
ggcatgccca 3300agggtgcggt cgtgtcgcac cgcgccatcg tcaaccgcct
gcgctggatg caggaccgct 3360acggccttca ggccgacgac cgcgtgctgc
agaagacgcc ttccagcttc gacgtgtcgg 3420tgtgggagtt cttctggccg
ctgatcgacg gtgccacgct ggtgcttgcg aaaccgggcg 3480gccacaagga
tgcggcctac ctcgcggggc tgatcgcgga ggagggcatc accacgatcc
3540acttcgtgcc gtcgatgctc gaggtcttcc tgctcgagcc cacggcgggc
gcatgcacca 3600cgctgcgccg cgtgatctgc agcggcgaag ccttgtcgcc
cgcgctgcaa tcgcagttcc 3660agcagcacct ctcgtgcgag ctgcacaacc
tctacggtcc gaccgaggcc gcggtcgacg 3720tcacctcgtg ggagtgcgaa
cgcacggacg acgcagaagc ctcgagcgtt cccatcggcc 3780gcccgatctg
gaacacccag atgcacgtgc tcgacagcgg cctgcagccc gtgccggccg
3840gcgtgactgg cgagctgtac atcgcgggcg tcggcctcgc acgcggctac
ctcaagcgcc 3900cgttgctgag cgccgagcgt ttcatcgcca acccctacgg
cacacccggc agccgcatgt 3960accgcaccgg cgacctcgcg cgctggcgca
aggacggcag ccttgacttc ctcggccgcg 4020ccgaccagca ggtgaagatc
cggggcctgc gcatcgagcc gggagagatc gaatccgtgc 4080tgctgcagca
tccgcaagtc gcgcaggccg ccgtggtggc gcgcgaagac gtaccgggcg
4140aaaagcgtct cgtggcctac gtcgttgcga cggacgctgc cgatccgcaa
gcggccgaac 4200tgcgcacgcg cctcgcgcaa tcgctgcccg agtacatggt
gccttcggcc ttcgtcagcc 4260tcccgtcgct gccgctcgga cccagcggca
agctcgaccg caaggcgctg ccgccccccg 4320aagtgcaggc cgccacgccg
tacgccgcgc cgcgcacgcc gaccgaaaag atcctggccg 4380gcctctgggc
cgagacgctg catttgccgc gcgtcggtgt caacgacaac ttcttcgaac
4440tcggcggcca ctcgctgatg atcgtgcagc tcatgtcgat gatccggcag
caattcatga 4500tcgacctgcc ggtcgacacg ctgttccagg tctccaccat
cgcgggcctt gccgagctgc 4560tcgaccagga atcggtcgcc cgtccgagcc
tgactccgat gccgcgcccc gcgcgcattc 4620cgctgtcctt cgcgcagcgc
cgcctgtggc tgatgaacca gctcgaaggc gcgaacccgg 4680cctacaacat
gccgctcgcg ctgcgcctgt cgggtgtgct cgatcgcacc gcattgcatg
4740cggcgctcgg cgacctggtg cagcgccacg agagcctgcg cacggtctac
ccgaacgaag 4800acgggctgcc gtaccagcac atcctcgacg gcgcggatgc
gcgtccggcg gtgatcgagg 4860ccgacagcag cgaagaagaa atcgcggcgc
agcttcacgc cgctgcgggc catgccttcg 4920atctcggcag cgcggcgccc
ttgcgcgtct acctgttcaa gctcgccggc gacgaacacg 4980tgctgctgct
gctcacgcac cacattgccg gcgatggcgc ctcgctgctg ccgctagcgc
5040gcgacatcag cgtggcctat gccgcgcgct gcgaaggcaa ggcgccgggc
tgggagccgc 5100tgccgctgca atacgccgac tacgcgctgt ggcagcagga
gctgctcggc agcgaagacg 5160atgccgagag catggccggc cgccagcgtg
agttctggcg ttcctcgctg agcgacctgc 5220ccgagcaact ggcgctgccc
gtcgaccacg cacggccgct cgtgccgacc taccgcggcg 5280atgtggtccc
gctgcagatt ccgtcgcatg tgcatgaacg catcctgcaa ctggcgcgcg
5340acgggcaggc cagcgtcttc atggtgctgc aggccgcact cgcgggcctc
ctgagccgcc 5400tcggcgcggg cgacgacatc gtcatcggca gcccggtcgc
ggggcgcagc gaccatgcgc 5460tggacgaact catcggctgc ttcgtcaaca
cgctggtgct gcgcactgac acctcgggcc 5520agccgagcct gcgcgagctg
gtctcgcgcg tgcgcgccac caacctcgcg gcctatgcga 5580accaggagtt
tccgtacgac cgcctcgtgg agctgctgcg tccgggccgc tcgcgcgcca
5640acctgccgct gttccaggtc atgctgggct tccagggcac gagccgcctg
tcgttcagcc 5700tgccgggcct gtcgatcgcg ccgcagccgg tggccatcga
caccgcgaag ttcgacctgt 5760cgttcatcct cggcgagcaa cgcggtgccg
atggcctgcc gggcggcatc tccggcggca 5820tccagtacag caccgacctg
ttcgagcgca gcacggtcga ggccatgggc gcgcggctgg 5880tgcgtttgct
ggaagaggcc tgcgaggcgc ccgacgatgc ggtgagtggc ctcgccatcc
5940tgagcgcgga agaaaccgac cgcctgctgt ccgactggag cggccgcacg
cgcgaccttg 6000cgccgctctc gttcgccgac atggtggcct cgcatgccgc
ggagcgcccg cttgcagatg 6060cagtggtgct cgacgacgcg accgtcagct
acgccgaact cgatgcacgc gccaaccggc 6120tctcgcacct gctgcgtgcg
caaggcatcg gggttggcgc catcgtcgcg acagtgctgc 6180cgcgttcgct
cgacctcatc gtggcgcact tggccatcgt gaaggccggc gcggcctacc
6240tgcccatcga ccccaaccac atggccgcgc gcagcgcctt cgtgttcgag
gaggccgcgc 6300ccgccgcggt gctgacgcac gatgcgctgt tgcccgagct
ggtcggcgtt ccccgctgca 6360tcgcgctcga cagcgacagc atggttgccg
cgctggccat ccagtcggat acgccgctgg 6420tgcatgcggc caatccacag
gatgccgcct acctcatcta cacctccggc tccaccggca 6480tgcccaaggg
cgtggtggtg ccgcatgcgg gcctgggcag cctcggcacc gcgatggcgg
6540agcggctcgt catcggccac ggctcgcgcg tgctgcagtt ctcctccagc
ggcttcgacg 6600cgtcggtgat ggaccagctg atggcctttg gcgccggtgc
cgcgctggtg gtgccggggc 6660cggagcaact gctcggcacg gagctggccg
atctgctcga gaagcaggcc gtgagccacg 6720cgctgattcc gcccgccgcg
ctcgcgaccc tgccgcacgg cgagttcccg cacctgcaga 6780cgctggtggt
cggcggcgat gcctgcaccg ccgcgctggc ggcgaagtgg tcgcaaggcc
6840gccgcatgat caacgcctac ggcccgaccg agatcaccat ctgcgcgagc
atgagcgcgc 6900cgatgacggc cgaggagttg ccctccatcg gccagccgat
ctggaacacg cggatgtatg 6960tgctcgacag cgccctgcaa ccggtgccgc
cgggtgtcgc gggcgagctc tacatcgccg 7020gcagcggcgt ggcgcgcggc
tatctcaacc ggccggcatt gagtgcggaa cgcttcatcg 7080ccgacccgca
tggcgcgccc ggcagccgca tgtaccgcag cggcgacctc gcacgctggc
7140gcgccgacgg cacgctcgac ttcctcggcc gcgccgacca gcaggtgaag
atccggggct 7200tccgcatcga gccgggcgag atcgaatccg tgctgctcaa
gcacccgttg atcacgcagg 7260ccgccgtgat cgcccgcgag gacgtgcccg
gcgagaagcg cctggtcgcc tacttcgtcg 7320ccggttccga gccgcagccc
accgagctgc gcgcccacat ggcgcaggcc ttgcccgact 7380acatggtgcc
ttcggccttc gtgcgcctgc cgtcgctgcc gctcacgcaa agcggcaagc
7440tcgacaagaa ggcgctgccg gtgcccgacc agcagcccgc cgcgctgtac
gtggagcccc 7500gcacgccgac cgagaaactg ctcgcgggcc tctggtccga
gacgctgcac ctggagcgtg 7560tcggcatcca cgacaacttc ttcgagatcg
gcgggcattc gctcatggcg atccagctgg 7620gcatgcgcat ccgccagcag
gtgcgcgcgg acttcccgca cgccgaggtc tacaaccgcc 7680cgacgattgc
cgacctggcc gcctggctcg acaacgaagg cggcacggtc gaggcgctgg
7740acctgtcgcg cgagctcgac ctgcccgcgc acatccgccc gcaggccact
gcaccgaagc 7800tcgcaccgcg ccgcgtgttc ctcaccggcg cgagcggctt
cgtcggcagt cacctgctgg 7860ccgcgctgtt gcgcgacacc gcggcctgcg
tggtctgcca cgtgcgcgcg cccgacgagc 7920aggccggcga gcagcgcctc
aagcgcacgc tggcccagcg ccagctcggt gcgatctggg 7980acaacgcgcg
catcaaggtc gtgaccggcg acctcggcaa gccgcgcctg ggcctcgatg
8040acgctgccgt gcaactggtg cgcgacggct gcgacgccat ctaccactgc
gccgcgcagg 8100tcgacttcct gcatccctac gcgagcctca agcccgcgaa
cgtcgacagc gtggtcacgc 8160tgctcgaatg gacggcgcag gggcgcgcga
agagcatgca ctacgtctcc acgctggctg 8220tgatcgacca gaacaacaag
gaagacacca tcaccgagca atcggcgctg gcctcatgga 8280gcgggctggt
cgacggctac agccagagca agtgggtcgg cgatgcgctg gcccgcgagg
8340cgcaggcgcg cggcatgccg gtggcgatct accggctggg ggcagtcacc
ggcgaccaca 8400cgcacgcgat ctgcaatgcc gacgacctga tctggcgcgt
ggcgcatctc tatgccgacc 8460tggaagcgat tcccgatatg gacctgccgc
tcaacctcac accggtggac gacgtggcgc 8520gcgccatcct cggccttgcg
gcgcaggagg cctcgtgggg ccaggtgttc cacctgatga 8580gccaggcggc
gctgcgggtg cgcgacattc cgcacgtctt cgagcgcatg ggcatgcggc
8640tggagccggt cgggctggag ccctggctgc agcgcgcgca tgcacggctg
gccgtcgcgc 8700atgaccgcga cctggccgcg gtgctcgcca tcctcgaccg
ctacgacacc acggccacgc 8760cgccgcaggt gagcggcgcg gccacgcatg
cgcagctcga ggccatcggc gcgccgatcc 8820gcccggtgga ccgcgacctg
ctgcagcgct acttcgtcga cctgggcatc gacaccaagg 8880cgcgccgcgc
cctggaaacc accacttcat aggagcacac ggaatggcac gctatctcat
8940cgcagcaacc gccttgccgg gacacgtcct gccgatgctg gccatcgcgc
agcatctggt 9000gaaccagggg cacgaggtgc gggtgcacac cgcgagccag
ttcagggcgc aggccgaggc 9060gaccggtgcg ggcttcacgc ccttcgagcg
cacgatcgac ttcgactacc gcgacctgga 9120caagcgcttt cccgagcgcc
agcgcatcgc ctcggcgcat gcgcagctgt gcttcggcct 9180gaagcacttc
tttgccgatg cgatggccgc gcagcatgcg ggcctgcaat cgatcctcga
9240agacttcgag gccgatgcca tcgtggtcga cacgatgttc tgcggcactt
tcccgctgct 9300gctaggcaag gagcgcgaag accgcccggc catcgtcggc
atcggcatct cggcgctgcc 9360gctctcgagc tgcgacaccg ccttcttcgg
caccgcgctg ccgccgtcgt ccacgccgga 9420agggcgggtg cgcaacaagg
cgatgaacgc caacctcaaa caggcgatgt tcggcgaggt 9480gcaacgctac
ttcgacacgc tgctcgcgcg ttcgggcctg gccgcgctgc ccgatttctt
9540cgtcgatgcg atggtgaagc tgcccgatct ttacctgcag ctcaccgcgc
cttcgttcga 9600atacccgcgc agcgacctgc ccgcgtcggt gcatttcgtc
ggcccgctgc tctcgcccgc 9660gagccgcgac ttcacgccgc ccgagtggtg
gcacgagctg gacgacggcc gctcggtcgt 9720gctggtcacg cagggcacgc
tggccaacca gaatccgtcg cagctgatcg gcccgacgct 9780gcaggcgctg
gccggcgaca agaacatcct cgtcatcgcc accaccggcg gcccggtgcc
9840gcccgccctg acggtgaacc tgcccgccaa cgcccgcgtg gtgccgttcc
tgccctacga 9900ccggctgctg cccaagctgc acgcgatggt caccaacggc
ggctacggct cggtcaacca 9960tgcattgagc ctcggtgtgc cgctggtggt
ggccggcacc tccgaagaga agcccgagat 10020cgccgcgcgc gtggcctggt
cgggcgcggg catcaacctc gccaccggcc agccgaccgc 10080gcgccaggtc
ggcgacgcgg tgcgcaaggt actgggcaac tcgacctatc gccagcgtgc
10140ggcggtgctg cgtgaggact tcgcttgcca tcgcgcgctg accggcatcg
ccggcgccct 10200cgaggcactt ctgcaaacct tcgcatccgc ggaaatggct
tgaacctgaa ccccatacga 10260caaaggaaat cccagatgag caacccgttc
gacgacaaga acgccagctt ccaggtgctg 10320gtgaacgacg agggccagca
ctcgctgtgg cccgccttca tcgccgtgcc cgccggctgg 10380caggtggcgc
tggcgccgac cgaccgcgac gcctgcagcg cctacatcgc ggcgaactgg
10440caggacatgc gcccgcgttc gctggtggtg gccacggcgg ccggctgacg
ccgaggatgt 10500ccttcccgtt cggtgccgtc gtcgtcacct atttcccgac
cggcgagcaa gtggcgaacc 10560tccattcgct ggcggcctcg tgtccgcacc
tctgcgtggt cgacaacacg ccgcaggtgg 10620gcgattggca tgcggcgctc
gtcgatgcgg gcgtttcggt gctgcacaac ggcaaccgcg 10680gcggcatcgc
gggcgccttc aaccgcggca tcatcgacct cgaagcgcgg ggcgccgaac
10740tcttcttcct gctcgaccag gattcgaagc tgccacccgg ctacttcgat
gccatgtgcg 10800aggctgcgat ggtggcccgg gagcggaagg gcgagggcaa
tggtgaggaa gacgcggcct 10860tcctgatcgg cccgctcgtc cacgacacga
acctggacgc gctgatcccg caattcggcc 10920tccagggcaa acgcgtctac
cagttcgacc tgcggcagcc cttcaccgag ccgctgatgc 10980gctgcgcctt
catgatttcc tcgggctccc tgatttcgcg cggcgcctgg gcccggatcg
11040gccggttcga cgagcgctat gtgatcgacc acgtggacac cgactactgc
atgcgtgccc 11100tgggtcgcgg cgtgccgctc tacctgaatc cgcacgtcgt
gctgcggcac cagattggcg 11160acatccgtgc ccggtcgctg ttcggctgga
agatccactt catcaactac ccggccgcgc 11220ggcgctacta catcgcgcgc
aatgccatcg atctctcgcg ggcgcatgtg cgcgcctttc 11280ccgcgatcct
gttcatcaac gtttacacgc tcaagcagat cctgccgatg ctgatgttcg
11340agcgcgaccg cttcaagaag accatcgcgc tgatgctcgg ctgcttcgat
ggcctgttcg 11400ggcggctcgg gggcctcggc gaggtgcatc cgcggatggg
caaatacctg ggccgcagcg 11460attgaccgcc acccttccag cgccgcgcgt
acgccgcgcc gcgctcgcct tcatcttcgt 11520cacggtgctg atcgacttca
tggcgttcgg cctgatcctg cccggcctgc cgcacctggt 11580ggagcggctg
gccggcggca gcacggtaac ggcggcgtac tggatcgctg tgttcggcac
11640cgcgttcgcg gcgatccagt tcgtgagctc gccgatccag ggcgcgctgt
ccgaccgctt 11700cgggcggcgg ccggtgatcc tgctgtcgtg cttcggcctc
ggcgtggatt tcgtgttcat 11760ggccctggcc gacagcctgc cgtggctgtt
cgtcggccgg gtggtctccg gcgtgttctc 11820ggccagcttc accatcgcca
atgcctacat cgccgatgtg acgctgccgg aggagcgcgc 11880ccgcagctac
ggcatcgtgg gggccgcgtt cggcatgggc ctggtgttcg ggccggtgct
11940cggcgggcaa ctgagccaca tcgatccgcg cctgccgttc tggttcgcgg
ccggcttgac 12000gctgctcagc ttctgctacg gatggttcgt gttgcccgaa
tcgctgccgc ccgagcggcg 12060tgcccgcaag ttcgactggt cgcatgccaa
tccggttggg acgctggtgc tgctcaagcg 12120ctatccgcag gtgttcggac
tggcggcggt gatcttcctc gtgaacctgg ctcagtacgt 12180ctatcccagc
gtgttcgtgc tgttcgccga ctaccggtat cactggaagg aagacgccgt
12240gggctgggtg ctcggcgcgg tgggcgtgct cagcgtgctg gtcaatgcgc
tgttgatcgg 12300gccgggcgtg aagcgcttcg gcgagcgccg cgccctgttg
ctcggcatgg gcttcggcgt 12360gctcggcttc gtcatcatcg ggtttgccga
cgctggatgg atcctcctgg ccggggtgcc 12420gttcggcatt ctgctggcgt
tcgccggacc ggcggcgcag gcgctggtca cgctgcaggt 12480cggcaccgcc
gagcagggcc gcatccaggg ggcgctcacc agcctggtgt cggtggcggg
12540catcgtcggg ccggcgatgt tcgccggcag cttcggttac ttcatcggcg
cggacgcgcc 12600ggtgcacttg ccgggcgcgc cgtttttcct cgctgcggcg
ttcctctgca tcggcacgct 12660gatcgcgtgg cgctacgcac agccgaagcc
cgcgacggca gcggtgcccg agccgacctg 12720a 1272123959DNAVariovorax
paradoxus 2ccgctgcgcc tcgcaacggg tttgctcctt cggtgcatcg cgatccctgc
gggtgcgatg 60gctctccaga cggcgtttga tgtgatgcag tactgacccc ctgttcgggc
cgacctgagc 120gtttatggga gtttgcgcct tcggtagggc caccggggtg
gcccgctctc ctgcagtggg 180gcgattgtag gtgggcactg ccaatgcgcc
aaccccggga gtttcggccc ttgggccgat 240gggataatca tccgttcatt
cgccggaggg cgatcgttcg acaacaacag gggaccccat 300gatcctggta
accggcggcg caggcttcat tggcgccaat ttcgtactcg actggctcgc
360acagagcgat gaaccggtcg tgaacctaga caagctgacc tacgcgggca
acctcgagac 420gctcgcatcg ctcaaggaca acccgaagca catcttcgtg
cagggcgaca tcggcgacag 480cgcgctgctc gaccgcctgc tggccgagca
caagccgcgt gccgtggtca acttcgcggc 540cgaatcgcac gtcgaccgct
cgatccacgg ccccgaagac ttcgtgcaga ccaacgtgct 600gggcaccttc
cgcctgctcg aatccgtgcg cggtttctgg aatgccctgc cggccgacca
660gaaggccgcc ttccgcttcc tgcatgtgtc gaccgacgag gtctacggct
cgctctccaa 720gaccgacccg gccttcaccg aagagaacaa gtacgagccc
aacagcccgt actcggccag 780caaggccgcc agcgaccacc tcgtgcgcgc
ctggcaccac acctacggcc tgccggtggt 840caccaccaac tgctcgaaca
actacgggcc gttccacttc cccgagaagc tcattcccct 900gatgatcgtc
aacgcgctgg cgggcaagcc gctgcccgtg tacggcgacg gcatgcaggt
960gcgcgactgg ctctacgtga aggaccactg cagcgccatc cgccgcgtgc
tcgaagccgg 1020caagctcggc gagacctaca acgtgggcgg ctggaacgag
aagcccaaca tcgagatcgt 1080caacaccgtc tgcgcgctgc tcgacgagct
gagccccaag gccggcggca agccgtacaa 1140ggaacagatc acctatgtga
ccgaccgccc cggccacgac cgccgctacg cgatcgacgc 1200acgcaagctc
gagcgcgaac tcggctggaa acctgccgag accttcgaca gcggcatccg
1260caagacggtc gagtggtacc tcgcgaacgg cgagtgggtg cgcaacgtgc
aaagcggcgc 1320gtaccgcgag tgggtcgaga agcaatacga cgccgcaccg
gcgaaggcca ccgcatgaag 1380ctgctgctgc tgggcaaggg cggacaggtc
ggctgggagc tgcaacgcag cctcgcgccc 1440ctgggcgaac tggtggcgct
cgatttcgac agcaccgact tcaacgccga cttcagtcgc 1500cccgagcagc
tggccgagac agtgctgaag gtgcgccccg acgtcatcgt caatgccgca
1560gcgcacaccg cggtcgacaa ggccgagagc gagcccgagt tcgcgcgcaa
gctcaacgcc 1620acctcgcccg gcgtggtggc cgaagccgcg cagcagatcg
gcgcgctgat ggttcactac 1680tcgaccgact acgtcttcga cggcagcggc
agcaagccgt ggaaagaaga cgatgcgacc 1740ggcccgctca gcgtctacgg
cagcaccaag ctcgaaggcg agcaactggt ggcaaagcac 1800tgtgcgaagc
acctgatctt tcgcaccagc tgggtctatg ccgcgcgcgg cggcaacttc
1860gccaagacca tgctgcgcat cgccaaggag cgcgacaagc tgaccgtcat
cgacgaccag 1920ttcggcgcgc ccaccggcgc ggaactgctg gccgacatca
ccgcgcacgc gattcgcgcg 1980acgctgcagg acccgtccaa ggccgggctc
tatcacgcgg tggccggtgg cgtgaccacg 2040tggcacggct atgcgcgctt
cgtgatcgag caggccaagg cggcgggcgt ggaactgaag 2100gccggccccg
aagcggtcga gcccgtgccc accacggcat tcccgacgcc ggccaggcgg
2160ccgcacaact cgcgcctgga caccaccaag ctgcaatcga ccttcggcct
cgtgctgccc
2220gagtggcagt ccggcgtcgc ccgcatgttg cgcgaaacct tctgatattc
gcagagcaag 2280agagacacga acaccccatg accaagacga cgcaacgcaa
aggcatcatc ctcgccggtg 2340gctcgggcac ccgcctgcac cccgcgacgc
ttgccatgag caaacaactg ctgccggtgt 2400acgacaagcc gatgatctat
tacccgctga gcacgctgat gctgggcggc atgcgcgaca 2460tcctgatcat
cagcacgccg caggacacgc cgcgtttcca gcaactgctg ggggatggca
2520gccaatgggg catcaacctg cagtacgcgg tgcagccgag cccggatggt
ctggcgcagg 2580cgttcatcat cggtgacaag ttcgtgggca acgacccgag
tgcgctggtg ctgggggaca 2640acatcttcta tggccacgac ttcgcccatc
tgctggccga tgccgacgcc aagacctcgg 2700gtgcgacggt gttcgcctac
cacgtgcacg accccgagcg ctacggcgtg gtggccttcg 2760atgccaaggg
cagggcgagc agcatcgaag aaaagccgct caagcccaag agcagctatg
2820cggtcacggg cctctacttc tacgacaacc aggtcgtcga catcgccaag
gccgtgaagc 2880cgagcgcgcg cggcgaactc gagatcaccg cggtcaacca
ggcgtatctc gacctcgacc 2940agctgaacgt gcagatcatg cagcgcggct
atgcgtggct cgataccggt acgcacgaca 3000gcctgctgga agccgggcag
ttcattgcca cgctcgagca ccgccagggg ctgaagatcg 3060catgccccga
agagatcgca tggcgcaatg gcttcatctc aaccgagcaa ctcgaaaagc
3120tcgcggcgcc gctggaaaag agcggctacg gcaagtacct caagcacctg
ctgaacgacg 3180aggtgcgctc gtgaaggcca cgcccacctc gattcctgac
gtgctcgtga tcgagccgaa 3240ggtgtttggc gatgcacggg gcttcttctt
cgaaagcttc aaccagaagg ccttcgacga 3300agcgatcggc aagcatgtcg
acttcgtgca ggacaaccat tcgcgatcgg ccaagggtgt 3360gctgcggggg
ctgcattacc aggtccagca gccgcaaggc aagctcgtgc gggtggtgcg
3420tggtgcggtg ttcgacgtgg ccgtcgacat ccgcaagtcg tcgccgactt
ttggcaaatg 3480ggtgggtgtc gagttgaacg aagacaacca caagcagctc
tgggtgccgg caggattcgc 3540gcacggtttc ctggtgttga gcgagaccgc
ggaattcctc tacaagacca ccgactacta 3600cgcgcccgcc cacgagcgcg
cgattgtctg gaacgacccc gctgtcggta ttcgatggcc 3660ggatgtggga
ggggcaccgg tcctgtcgaa gaaggacgaa gacgggtgtc ttctgcaagc
3720ggcagaggtt ttctagtgtc ctttcgtcag atagcggggc ggcttcgcgt
atcgggatcc 3780cgcgttgagc ccgcaagagt gccctgagag ggggggcgaa
aaactcacaa cgccactgcc 3840tcgagcaaac gtgcgtctcg cagctttctg
aagttgttgc accttctttt tttttctctt 3900acatctttga aatgattttg
aaaatccgcg gcgatcgcat gcatgctgct ggaatcacc 39593915DNAVariovorax
paradoxus 3atgaatggca tgcatatcga ctcggtcgac ctcaatctgc tgcgcctgtt
cgatgcggtc 60taccgcgagc gcagcgtgag ccgcgccgcg gagtcgctgg gcctcacgca
gcctgcggca 120agccatgggc tgggacggct gcggctgctt ttgaaagacg
cgctcttcac gcgtgccccc 180ggcggcgtgg cgcccacgcc gcgcgccgac
cggctcgcgg tggcggtgca ggcggcgctc 240ggcacgatcg aagcggcgct
gcacgagccc gatcgcttcg agccccaggt gtcgcgcaag 300agctttcgta
ttcacatgag cgacatcggc gaggggcgct tcctgcccgc gctgatggcg
360cggctcggcg agctggcgcc cggcgtgcgg ctggagaccc tgccgctctt
gcctgcggag 420gttgcgcccg cactcgacag cggccgcatc gatttcgcct
tcggctttct ctcgaccgtg 480cgcgacacgc agcgcacgca tcttctgaaa
gaccgctaca tcgtgctgct gcgcaagggc 540catccctttg tgaagcgccg
gcgcaagggg caggcgctgc tcgaggcgct gcaggagctc 600gactacgtgg
cggtgcgcac gcacgccgac acgctgcgca tcttgcagtt gctcaacctc
660gaagaccgcc tgcgcctcac gaccgagcac ttcatggtgc taccggccat
cgtgcgcgcc 720accgatctcg cggtggtgat gccgcgcaac atcgcgcgag
ggtttgcgga ggagggcggc 780tacgcgatcg tcgagccgcc gtttccgctg
cgcgatttca gcgtgtcgct gcactggagc 840aagcgcttcg agggcgaccc
ggccaaccgt tggttgcggc aggtgatcac ggcgctgttc 900tccgagcgcg gctga
9154304PRTVariovorax paradoxus 4Met Asn Gly Met His Ile Asp Ser Val
Asp Leu Asn Leu Leu Arg Leu1 5 10 15Phe Asp Ala Val Tyr Arg Glu Arg
Ser Val Ser Arg Ala Ala Glu Ser 20 25 30Leu Gly Leu Thr Gln Pro Ala
Ala Ser His Gly Leu Gly Arg Leu Arg 35 40 45Leu Leu Leu Lys Asp Ala
Leu Phe Thr Arg Ala Pro Gly Gly Val Ala 50 55 60Pro Thr Pro Arg Ala
Asp Arg Leu Ala Val Ala Val Gln Ala Ala Leu65 70 75 80Gly Thr Ile
Glu Ala Ala Leu His Glu Pro Asp Arg Phe Glu Pro Gln 85 90 95Val Ser
Arg Lys Ser Phe Arg Ile His Met Ser Asp Ile Gly Glu Gly 100 105
110Arg Phe Leu Pro Ala Leu Met Ala Arg Leu Gly Glu Leu Ala Pro Gly
115 120 125Val Arg Leu Glu Thr Leu Pro Leu Leu Pro Ala Glu Val Ala
Pro Ala 130 135 140Leu Asp Ser Gly Arg Ile Asp Phe Ala Phe Gly Phe
Leu Ser Thr Val145 150 155 160Arg Asp Thr Gln Arg Thr His Leu Leu
Lys Asp Arg Tyr Ile Val Leu 165 170 175Leu Arg Lys Gly His Pro Phe
Val Lys Arg Arg Arg Lys Gly Gln Ala 180 185 190Leu Leu Glu Ala Leu
Gln Glu Leu Asp Tyr Val Ala Val Arg Thr His 195 200 205Ala Asp Thr
Leu Arg Ile Leu Gln Leu Leu Asn Leu Glu Asp Arg Leu 210 215 220Arg
Leu Thr Thr Glu His Phe Met Val Leu Pro Ala Ile Val Arg Ala225 230
235 240Thr Asp Leu Ala Val Val Met Pro Arg Asn Ile Ala Arg Gly Phe
Ala 245 250 255Glu Glu Gly Gly Tyr Ala Ile Val Glu Pro Pro Phe Pro
Leu Arg Asp 260 265 270Phe Ser Val Ser Leu His Trp Ser Lys Arg Phe
Glu Gly Asp Pro Ala 275 280 285Asn Arg Trp Leu Arg Gln Val Ile Thr
Ala Leu Phe Ser Glu Arg Gly 290 295 30057476DNAVariovorax paradoxus
5atgagtaccg tcgatcagct gggccgcacc gcccccctta cctcggggca gatggcgatg
60tggctcggcg caaagttcgc gtcgcccgac accaatttca atctcgccga agccatcgac
120atcgcaggcg agatcgaccc cgcgatcttc ctggcggcca tgcgacaggt
ggccgatgaa 180gtcgaggcca cgcgcctgag cttcatcgat accccgcaag
ggccacgaca ggtcgtcgcg 240cccgttttca ccggcgagat cccctacctc
gacctcagcg gcgagagcga tccgcaggcc 300gaggccgagc gctggatgca
tgcggactac acccgcagca tcgacctcgc gcacgggcag 360ctgtggctgt
ccgcgctgat ccgcctcgcg cccgatcgcc acatctggta ccaccgcagc
420catcacatcg cgctcgacgg cttcagcggc ggcctcatcg cacgccgctt
cgccgacatc 480tacaccgcga tggtcgacaa caacgcagcg gtgcccgaag
actcgcgcct tgcaccgatc 540tcgcagctgg ccgacgaaga acatgcctat
cgcgagtccg gccgcttccc gcgcgaccgc 600cagtactgga ccgagcgctt
cgccgatgca cccgatccgt tgagcctcgc ctcgcaccgc 660tcggtcaacg
tcggtggcct cttgcgccag acggtgcacc tgccggcggc cagcgtgcaa
720gccctgcaga ccatcgcgca agagctcggc accacgctgc cgcaaatcct
catcgccacc 780accgcggcct acctgtaccg cgcaacgggc atcgaggaca
tggcaatcgg catccccgtc 840accgcgcgcc acaacgaccg catgcgccgc
gtgcccgcga tggtggccaa cgcgctgccg 900ctgcgcctgg cgatgcgcgc
ggacctgccg attccggaac tgatccgcga agtcggccgg 960cagatgcggc
agatcctgcg gcaccagtcg tatcgctacg agcatttgcg cagcgacctc
1020aacatgctgg tgaacaaccg gcagctcttc accaccgtgg tcaacgtcga
gcccttcgac 1080tacgacttcc gctttgcggg ccatgccgcg aagccgcgca
acctctcgaa cggcacggcc 1140gaggacctcg gcatcttcct gtacgagcgc
ggcaacgggc aggacctgca gatcgacttc 1200gacgccaacc ccgcggtgca
caccgcagag gaactggccg atcaccagcg ccggctgctt 1260gccttcatcg
acgccgtgat ccgcctgccg ttgcaggccg tcggccagat cgacctgctc
1320ggtgccgaag agcggcagca attgctggtc gagtggaacg acacggccca
cgccgtgccc 1380gacacccatc tcaccgcgtt gatcgaagcg cagctcgcag
ccgatccgca agccatcgca 1440ttgcgcttcg acggcgaggc gatgaacaac
gaagaactga accgccgcgc caaccgtctc 1500gcccacctgc tgcgcgcacg
cggcgctggc ccggagcgca ccgtggcgct cgcgatcccg 1560cgttcgatgg
acctgatgat tgccttgctc gccacgttga agaccggcgc ggcctacctg
1620ccggtcgatc cggatttccc ggcggaccgc atcgccttca tgctcggcga
tgcgcagccc 1680gtgtgcctcg tcacgaccga agccctcgcg gagtcgctgc
cggcagccgc ccccacattg 1740ctgctcgatg tagcgcaaac gattgcggat
ctggagagtt gcaacgacac caacccgggc 1800atcgcgatcg acccttcgca
tccggcctat gtgatctaca cctcgggctc gaccggcatg 1860cccaagggtg
cggtcgtgtc gcaccgcgcc atcgtcaacc gcctgcgctg gatgcaggac
1920cgctacggcc ttcaggccga cgaccgcgtg ctgcagaaga cgccttccag
cttcgacgtg 1980tcggtgtggg agttcttctg gccgctgatc gacggtgcca
cgctggtgct tgcgaaaccg 2040ggcggccaca aggatgcggc ctacctcgcg
gggctgatcg cggaggaggg catcaccacg 2100atccacttcg tgccgtcgat
gctcgaggtc ttcctgctcg agcccacggc gggcgcatgc 2160accacgctgc
gccgcgtgat ctgcagcggc gaagccttgt cgcccgcgct gcaatcgcag
2220ttccagcagc acctctcgtg cgagctgcac aacctctacg gtccgaccga
ggccgcggtc 2280gacgtcacct cgtgggagtg cgaacgcacg gacgacgcag
aagcctcgag cgttcccatc 2340ggccgcccga tctggaacac ccagatgcac
gtgctcgaca gcggcctgca gcccgtgccg 2400gccggcgtga ctggcgagct
gtacatcgcg ggcgtcggcc tcgcacgcgg ctacctcaag 2460cgcccgttgc
tgagcgccga gcgtttcatc gccaacccct acggcacacc cggcagccgc
2520atgtaccgca ccggcgacct cgcgcgctgg cgcaaggacg gcagccttga
cttcctcggc 2580cgcgccgacc agcaggtgaa gatccggggc ctgcgcatcg
agccgggaga gatcgaatcc 2640gtgctgctgc agcatccgca agtcgcgcag
gccgccgtgg tggcgcgcga agacgtaccg 2700ggcgaaaagc gtctcgtggc
ctacgtcgtt gcgacggacg ctgccgatcc gcaagcggcc 2760gaactgcgca
cgcgcctcgc gcaatcgctg cccgagtaca tggtgccttc ggccttcgtc
2820agcctcccgt cgctgccgct cggacccagc ggcaagctcg accgcaaggc
gctgccgccc 2880cccgaagtgc aggccgccac gccgtacgcc gcgccgcgca
cgccgaccga aaagatcctg 2940gccggcctct gggccgagac gctgcatttg
ccgcgcgtcg gtgtcaacga caacttcttc 3000gaactcggcg gccactcgct
gatgatcgtg cagctcatgt cgatgatccg gcagcaattc 3060atgatcgacc
tgccggtcga cacgctgttc caggtctcca ccatcgcggg ccttgccgag
3120ctgctcgacc aggaatcggt cgcccgtccg agcctgactc cgatgccgcg
ccccgcgcgc 3180attccgctgt ccttcgcgca gcgccgcctg tggctgatga
accagctcga aggcgcgaac 3240ccggcctaca acatgccgct cgcgctgcgc
ctgtcgggtg tgctcgatcg caccgcattg 3300catgcggcgc tcggcgacct
ggtgcagcgc cacgagagcc tgcgcacggt ctacccgaac 3360gaagacgggc
tgccgtacca gcacatcctc gacggcgcgg atgcgcgtcc ggcggtgatc
3420gaggccgaca gcagcgaaga agaaatcgcg gcgcagcttc acgccgctgc
gggccatgcc 3480ttcgatctcg gcagcgcggc gcccttgcgc gtctacctgt
tcaagctcgc cggcgacgaa 3540cacgtgctgc tgctgctcac gcaccacatt
gccggcgatg gcgcctcgct gctgccgcta 3600gcgcgcgaca tcagcgtggc
ctatgccgcg cgctgcgaag gcaaggcgcc gggctgggag 3660ccgctgccgc
tgcaatacgc cgactacgcg ctgtggcagc aggagctgct cggcagcgaa
3720gacgatgccg agagcatggc cggccgccag cgtgagttct ggcgttcctc
gctgagcgac 3780ctgcccgagc aactggcgct gcccgtcgac cacgcacggc
cgctcgtgcc gacctaccgc 3840ggcgatgtgg tcccgctgca gattccgtcg
catgtgcatg aacgcatcct gcaactggcg 3900cgcgacgggc aggccagcgt
cttcatggtg ctgcaggccg cactcgcggg cctcctgagc 3960cgcctcggcg
cgggcgacga catcgtcatc ggcagcccgg tcgcggggcg cagcgaccat
4020gcgctggacg aactcatcgg ctgcttcgtc aacacgctgg tgctgcgcac
tgacacctcg 4080ggccagccga gcctgcgcga gctggtctcg cgcgtgcgcg
ccaccaacct cgcggcctat 4140gcgaaccagg agtttccgta cgaccgcctc
gtggagctgc tgcgtccggg ccgctcgcgc 4200gccaacctgc cgctgttcca
ggtcatgctg ggcttccagg gcacgagccg cctgtcgttc 4260agcctgccgg
gcctgtcgat cgcgccgcag ccggtggcca tcgacaccgc gaagttcgac
4320ctgtcgttca tcctcggcga gcaacgcggt gccgatggcc tgccgggcgg
catctccggc 4380ggcatccagt acagcaccga cctgttcgag cgcagcacgg
tcgaggccat gggcgcgcgg 4440ctggtgcgtt tgctggaaga ggcctgcgag
gcgcccgacg atgcggtgag tggcctcgcc 4500atcctgagcg cggaagaaac
cgaccgcctg ctgtccgact ggagcggccg cacgcgcgac 4560cttgcgccgc
tctcgttcgc cgacatggtg gcctcgcatg ccgcggagcg cccgcttgca
4620gatgcagtgg tgctcgacga cgcgaccgtc agctacgccg aactcgatgc
acgcgccaac 4680cggctctcgc acctgctgcg tgcgcaaggc atcggggttg
gcgccatcgt cgcgacagtg 4740ctgccgcgtt cgctcgacct catcgtggcg
cacttggcca tcgtgaaggc cggcgcggcc 4800tacctgccca tcgaccccaa
ccacatggcc gcgcgcagcg ccttcgtgtt cgaggaggcc 4860gcgcccgccg
cggtgctgac gcacgatgcg ctgttgcccg agctggtcgg cgttccccgc
4920tgcatcgcgc tcgacagcga cagcatggtt gccgcgctgg ccatccagtc
ggatacgccg 4980ctggtgcatg cggccaatcc acaggatgcc gcctacctca
tctacacctc cggctccacc 5040ggcatgccca agggcgtggt ggtgccgcat
gcgggcctgg gcagcctcgg caccgcgatg 5100gcggagcggc tcgtcatcgg
ccacggctcg cgcgtgctgc agttctcctc cagcggcttc 5160gacgcgtcgg
tgatggacca gctgatggcc tttggcgccg gtgccgcgct ggtggtgccg
5220gggccggagc aactgctcgg cacggagctg gccgatctgc tcgagaagca
ggccgtgagc 5280cacgcgctga ttccgcccgc cgcgctcgcg accctgccgc
acggcgagtt cccgcacctg 5340cagacgctgg tggtcggcgg cgatgcctgc
accgccgcgc tggcggcgaa gtggtcgcaa 5400ggccgccgca tgatcaacgc
ctacggcccg accgagatca ccatctgcgc gagcatgagc 5460gcgccgatga
cggccgagga gttgccctcc atcggccagc cgatctggaa cacgcggatg
5520tatgtgctcg acagcgccct gcaaccggtg ccgccgggtg tcgcgggcga
gctctacatc 5580gccggcagcg gcgtggcgcg cggctatctc aaccggccgg
cattgagtgc ggaacgcttc 5640atcgccgacc cgcatggcgc gcccggcagc
cgcatgtacc gcagcggcga cctcgcacgc 5700tggcgcgccg acggcacgct
cgacttcctc ggccgcgccg accagcaggt gaagatccgg 5760ggcttccgca
tcgagccggg cgagatcgaa tccgtgctgc tcaagcaccc gttgatcacg
5820caggccgccg tgatcgcccg cgaggacgtg cccggcgaga agcgcctggt
cgcctacttc 5880gtcgccggtt ccgagccgca gcccaccgag ctgcgcgccc
acatggcgca ggccttgccc 5940gactacatgg tgccttcggc cttcgtgcgc
ctgccgtcgc tgccgctcac gcaaagcggc 6000aagctcgaca agaaggcgct
gccggtgccc gaccagcagc ccgccgcgct gtacgtggag 6060ccccgcacgc
cgaccgagaa actgctcgcg ggcctctggt ccgagacgct gcacctggag
6120cgtgtcggca tccacgacaa cttcttcgag atcggcgggc attcgctcat
ggcgatccag 6180ctgggcatgc gcatccgcca gcaggtgcgc gcggacttcc
cgcacgccga ggtctacaac 6240cgcccgacga ttgccgacct ggccgcctgg
ctcgacaacg aaggcggcac ggtcgaggcg 6300ctggacctgt cgcgcgagct
cgacctgccc gcgcacatcc gcccgcaggc cactgcaccg 6360aagctcgcac
cgcgccgcgt gttcctcacc ggcgcgagcg gcttcgtcgg cagtcacctg
6420ctggccgcgc tgttgcgcga caccgcggcc tgcgtggtct gccacgtgcg
cgcgcccgac 6480gagcaggccg gcgagcagcg cctcaagcgc acgctggccc
agcgccagct cggtgcgatc 6540tgggacaacg cgcgcatcaa ggtcgtgacc
ggcgacctcg gcaagccgcg cctgggcctc 6600gatgacgctg ccgtgcaact
ggtgcgcgac ggctgcgacg ccatctacca ctgcgccgcg 6660caggtcgact
tcctgcatcc ctacgcgagc ctcaagcccg cgaacgtcga cagcgtggtc
6720acgctgctcg aatggacggc gcaggggcgc gcgaagagca tgcactacgt
ctccacgctg 6780gctgtgatcg accagaacaa caaggaagac accatcaccg
agcaatcggc gctggcctca 6840tggagcgggc tggtcgacgg ctacagccag
agcaagtggg tcggcgatgc gctggcccgc 6900gaggcgcagg cgcgcggcat
gccggtggcg atctaccggc tgggggcagt caccggcgac 6960cacacgcacg
cgatctgcaa tgccgacgac ctgatctggc gcgtggcgca tctctatgcc
7020gacctggaag cgattcccga tatggacctg ccgctcaacc tcacaccggt
ggacgacgtg 7080gcgcgcgcca tcctcggcct tgcggcgcag gaggcctcgt
ggggccaggt gttccacctg 7140atgagccagg cggcgctgcg ggtgcgcgac
attccgcacg tcttcgagcg catgggcatg 7200cggctggagc cggtcgggct
ggagccctgg ctgcagcgcg cgcatgcacg gctggccgtc 7260gcgcatgacc
gcgacctggc cgcggtgctc gccatcctcg accgctacga caccacggcc
7320acgccgccgc aggtgagcgg cgcggccacg catgcgcagc tcgaggccat
cggcgcgccg 7380atccgcccgg tggaccgcga cctgctgcag cgctacttcg
tcgacctggg catcgacacc 7440aaggcgcgcc gcgccctgga aaccaccact tcatag
747662491PRTVariovorax paradoxus 6Met Ser Thr Val Asp Gln Leu Gly
Arg Thr Ala Pro Leu Thr Ser Gly1 5 10 15Gln Met Ala Met Trp Leu Gly
Ala Lys Phe Ala Ser Pro Asp Thr Asn 20 25 30Phe Asn Leu Ala Glu Ala
Ile Asp Ile Ala Gly Glu Ile Asp Pro Ala 35 40 45Ile Phe Leu Ala Ala
Met Arg Gln Val Ala Asp Glu Val Glu Ala Thr 50 55 60Arg Leu Ser Phe
Ile Asp Thr Pro Gln Gly Pro Arg Gln Val Val Ala65 70 75 80Pro Val
Phe Thr Gly Glu Ile Pro Tyr Leu Asp Leu Ser Gly Glu Ser 85 90 95Asp
Pro Gln Ala Glu Ala Glu Arg Trp Met His Ala Asp Tyr Thr Arg 100 105
110Ser Ile Asp Leu Ala His Gly Gln Leu Trp Leu Ser Ala Leu Ile Arg
115 120 125Leu Ala Pro Asp Arg His Ile Trp Tyr His Arg Ser His His
Ile Ala 130 135 140Leu Asp Gly Phe Ser Gly Gly Leu Ile Ala Arg Arg
Phe Ala Asp Ile145 150 155 160Tyr Thr Ala Met Val Asp Asn Asn Ala
Ala Val Pro Glu Asp Ser Arg 165 170 175Leu Ala Pro Ile Ser Gln Leu
Ala Asp Glu Glu His Ala Tyr Arg Glu 180 185 190Ser Gly Arg Phe Pro
Arg Asp Arg Gln Tyr Trp Thr Glu Arg Phe Ala 195 200 205Asp Ala Pro
Asp Pro Leu Ser Leu Ala Ser His Arg Ser Val Asn Val 210 215 220Gly
Gly Leu Leu Arg Gln Thr Val His Leu Pro Ala Ala Ser Val Gln225 230
235 240Ala Leu Gln Thr Ile Ala Gln Glu Leu Gly Thr Thr Leu Pro Gln
Ile 245 250 255Leu Ile Ala Thr Thr Ala Ala Tyr Leu Tyr Arg Ala Thr
Gly Ile Glu 260 265 270Asp Met Ala Ile Gly Ile Pro Val Thr Ala Arg
His Asn Asp Arg Met 275 280 285Arg Arg Val Pro Ala Met Val Ala Asn
Ala Leu Pro Leu Arg Leu Ala 290 295 300Met Arg Ala Asp Leu Pro Ile
Pro Glu Leu Ile Arg Glu Val Gly Arg305 310 315 320Gln Met Arg Gln
Ile Leu Arg His Gln Ser Tyr Arg Tyr Glu His Leu 325 330 335Arg Ser
Asp Leu Asn Met Leu Val Asn Asn Arg Gln Leu Phe Thr Thr 340 345
350Val Val Asn Val Glu Pro Phe Asp Tyr Asp Phe Arg Phe Ala Gly His
355 360 365Ala Ala Lys Pro Arg Asn Leu Ser Asn Gly Thr Ala Glu Asp
Leu Gly 370 375 380Ile Phe Leu Tyr Glu Arg Gly Asn Gly Gln Asp Leu
Gln Ile Asp Phe385 390 395 400Asp Ala Asn Pro Ala Val His Thr Ala
Glu Glu Leu Ala Asp His Gln 405 410 415Arg Arg Leu Leu Ala Phe Ile
Asp Ala Val Ile Arg Leu Pro Leu Gln 420 425 430Ala Val Gly Gln Ile
Asp Leu Leu Gly Ala Glu Glu Arg Gln Gln Leu 435 440 445Leu Val Glu
Trp Asn Asp Thr Ala His Ala Val Pro Asp Thr His Leu 450 455 460Thr
Ala Leu Ile Glu Ala Gln Leu Ala Ala Asp Pro Gln Ala Ile Ala465 470
475
480Leu Arg Phe Asp Gly Glu Ala Met Asn Asn Glu Glu Leu Asn Arg Arg
485 490 495Ala Asn Arg Leu Ala His Leu Leu Arg Ala Arg Gly Ala Gly
Pro Glu 500 505 510Arg Thr Val Ala Leu Ala Ile Pro Arg Ser Met Asp
Leu Met Ile Ala 515 520 525Leu Leu Ala Thr Leu Lys Thr Gly Ala Ala
Tyr Leu Pro Val Asp Pro 530 535 540Asp Phe Pro Ala Asp Arg Ile Ala
Phe Met Leu Gly Asp Ala Gln Pro545 550 555 560Val Cys Leu Val Thr
Thr Glu Ala Leu Ala Glu Ser Leu Pro Ala Ala 565 570 575Ala Pro Thr
Leu Leu Leu Asp Val Ala Gln Thr Ile Ala Asp Leu Glu 580 585 590Ser
Cys Asn Asp Thr Asn Pro Gly Ile Ala Ile Asp Pro Ser His Pro 595 600
605Ala Tyr Val Ile Tyr Thr Ser Gly Ser Thr Gly Met Pro Lys Gly Ala
610 615 620Val Val Ser His Arg Ala Ile Val Asn Arg Leu Arg Trp Met
Gln Asp625 630 635 640Arg Tyr Gly Leu Gln Ala Asp Asp Arg Val Leu
Gln Lys Thr Pro Ser 645 650 655Ser Phe Asp Val Ser Val Trp Glu Phe
Phe Trp Pro Leu Ile Asp Gly 660 665 670Ala Thr Leu Val Leu Ala Lys
Pro Gly Gly His Lys Asp Ala Ala Tyr 675 680 685Leu Ala Gly Leu Ile
Ala Glu Glu Gly Ile Thr Thr Ile His Phe Val 690 695 700Pro Ser Met
Leu Glu Val Phe Leu Leu Glu Pro Thr Ala Gly Ala Cys705 710 715
720Thr Thr Leu Arg Arg Val Ile Cys Ser Gly Glu Ala Leu Ser Pro Ala
725 730 735Leu Gln Ser Gln Phe Gln Gln His Leu Ser Cys Glu Leu His
Asn Leu 740 745 750Tyr Gly Pro Thr Glu Ala Ala Val Asp Val Thr Ser
Trp Glu Cys Glu 755 760 765Arg Thr Asp Asp Ala Glu Ala Ser Ser Val
Pro Ile Gly Arg Pro Ile 770 775 780Trp Asn Thr Gln Met His Val Leu
Asp Ser Gly Leu Gln Pro Val Pro785 790 795 800Ala Gly Val Thr Gly
Glu Leu Tyr Ile Ala Gly Val Gly Leu Ala Arg 805 810 815Gly Tyr Leu
Lys Arg Pro Leu Leu Ser Ala Glu Arg Phe Ile Ala Asn 820 825 830Pro
Tyr Gly Thr Pro Gly Ser Arg Met Tyr Arg Thr Gly Asp Leu Ala 835 840
845Arg Trp Arg Lys Asp Gly Ser Leu Asp Phe Leu Gly Arg Ala Asp Gln
850 855 860Gln Val Lys Ile Arg Gly Leu Arg Ile Glu Pro Gly Glu Ile
Glu Ser865 870 875 880Val Leu Leu Gln His Pro Gln Val Ala Gln Ala
Ala Val Val Ala Arg 885 890 895Glu Asp Val Pro Gly Glu Lys Arg Leu
Val Ala Tyr Val Val Ala Thr 900 905 910Asp Ala Ala Asp Pro Gln Ala
Ala Glu Leu Arg Thr Arg Leu Ala Gln 915 920 925Ser Leu Pro Glu Tyr
Met Val Pro Ser Ala Phe Val Ser Leu Pro Ser 930 935 940Leu Pro Leu
Gly Pro Ser Gly Lys Leu Asp Arg Lys Ala Leu Pro Pro945 950 955
960Pro Glu Val Gln Ala Ala Thr Pro Tyr Ala Ala Pro Arg Thr Pro Thr
965 970 975Glu Lys Ile Leu Ala Gly Leu Trp Ala Glu Thr Leu His Leu
Pro Arg 980 985 990Val Gly Val Asn Asp Asn Phe Phe Glu Leu Gly Gly
His Ser Leu Met 995 1000 1005Ile Val Gln Leu Met Ser Met Ile Arg
Gln Gln Phe Met Ile Asp 1010 1015 1020Leu Pro Val Asp Thr Leu Phe
Gln Val Ser Thr Ile Ala Gly Leu 1025 1030 1035Ala Glu Leu Leu Asp
Gln Glu Ser Val Ala Arg Pro Ser Leu Thr 1040 1045 1050Pro Met Pro
Arg Pro Ala Arg Ile Pro Leu Ser Phe Ala Gln Arg 1055 1060 1065Arg
Leu Trp Leu Met Asn Gln Leu Glu Gly Ala Asn Pro Ala Tyr 1070 1075
1080Asn Met Pro Leu Ala Leu Arg Leu Ser Gly Val Leu Asp Arg Thr
1085 1090 1095Ala Leu His Ala Ala Leu Gly Asp Leu Val Gln Arg His
Glu Ser 1100 1105 1110Leu Arg Thr Val Tyr Pro Asn Glu Asp Gly Leu
Pro Tyr Gln His 1115 1120 1125Ile Leu Asp Gly Ala Asp Ala Arg Pro
Ala Val Ile Glu Ala Asp 1130 1135 1140Ser Ser Glu Glu Glu Ile Ala
Ala Gln Leu His Ala Ala Ala Gly 1145 1150 1155His Ala Phe Asp Leu
Gly Ser Ala Ala Pro Leu Arg Val Tyr Leu 1160 1165 1170Phe Lys Leu
Ala Gly Asp Glu His Val Leu Leu Leu Leu Thr His 1175 1180 1185His
Ile Ala Gly Asp Gly Ala Ser Leu Leu Pro Leu Ala Arg Asp 1190 1195
1200Ile Ser Val Ala Tyr Ala Ala Arg Cys Glu Gly Lys Ala Pro Gly
1205 1210 1215Trp Glu Pro Leu Pro Leu Gln Tyr Ala Asp Tyr Ala Leu
Trp Gln 1220 1225 1230Gln Glu Leu Leu Gly Ser Glu Asp Asp Ala Glu
Ser Met Ala Gly 1235 1240 1245Arg Gln Arg Glu Phe Trp Arg Ser Ser
Leu Ser Asp Leu Pro Glu 1250 1255 1260Gln Leu Ala Leu Pro Val Asp
His Ala Arg Pro Leu Val Pro Thr 1265 1270 1275Tyr Arg Gly Asp Val
Val Pro Leu Gln Ile Pro Ser His Val His 1280 1285 1290Glu Arg Ile
Leu Gln Leu Ala Arg Asp Gly Gln Ala Ser Val Phe 1295 1300 1305Met
Val Leu Gln Ala Ala Leu Ala Gly Leu Leu Ser Arg Leu Gly 1310 1315
1320Ala Gly Asp Asp Ile Val Ile Gly Ser Pro Val Ala Gly Arg Ser
1325 1330 1335Asp His Ala Leu Asp Glu Leu Ile Gly Cys Phe Val Asn
Thr Leu 1340 1345 1350Val Leu Arg Thr Asp Thr Ser Gly Gln Pro Ser
Leu Arg Glu Leu 1355 1360 1365Val Ser Arg Val Arg Ala Thr Asn Leu
Ala Ala Tyr Ala Asn Gln 1370 1375 1380Glu Phe Pro Tyr Asp Arg Leu
Val Glu Leu Leu Arg Pro Gly Arg 1385 1390 1395Ser Arg Ala Asn Leu
Pro Leu Phe Gln Val Met Leu Gly Phe Gln 1400 1405 1410Gly Thr Ser
Arg Leu Ser Phe Ser Leu Pro Gly Leu Ser Ile Ala 1415 1420 1425Pro
Gln Pro Val Ala Ile Asp Thr Ala Lys Phe Asp Leu Ser Phe 1430 1435
1440Ile Leu Gly Glu Gln Arg Gly Ala Asp Gly Leu Pro Gly Gly Ile
1445 1450 1455Ser Gly Gly Ile Gln Tyr Ser Thr Asp Leu Phe Glu Arg
Ser Thr 1460 1465 1470Val Glu Ala Met Gly Ala Arg Leu Val Arg Leu
Leu Glu Glu Ala 1475 1480 1485Cys Glu Ala Pro Asp Asp Ala Val Ser
Gly Leu Ala Ile Leu Ser 1490 1495 1500Ala Glu Glu Thr Asp Arg Leu
Leu Ser Asp Trp Ser Gly Arg Thr 1505 1510 1515Arg Asp Leu Ala Pro
Leu Ser Phe Ala Asp Met Val Ala Ser His 1520 1525 1530Ala Ala Glu
Arg Pro Leu Ala Asp Ala Val Val Leu Asp Asp Ala 1535 1540 1545Thr
Val Ser Tyr Ala Glu Leu Asp Ala Arg Ala Asn Arg Leu Ser 1550 1555
1560His Leu Leu Arg Ala Gln Gly Ile Gly Val Gly Ala Ile Val Ala
1565 1570 1575Thr Val Leu Pro Arg Ser Leu Asp Leu Ile Val Ala His
Leu Ala 1580 1585 1590Ile Val Lys Ala Gly Ala Ala Tyr Leu Pro Ile
Asp Pro Asn His 1595 1600 1605Met Ala Ala Arg Ser Ala Phe Val Phe
Glu Glu Ala Ala Pro Ala 1610 1615 1620Ala Val Leu Thr His Asp Ala
Leu Leu Pro Glu Leu Val Gly Val 1625 1630 1635Pro Arg Cys Ile Ala
Leu Asp Ser Asp Ser Met Val Ala Ala Leu 1640 1645 1650Ala Ile Gln
Ser Asp Thr Pro Leu Val His Ala Ala Asn Pro Gln 1655 1660 1665Asp
Ala Ala Tyr Leu Ile Tyr Thr Ser Gly Ser Thr Gly Met Pro 1670 1675
1680Lys Gly Val Val Val Pro His Ala Gly Leu Gly Ser Leu Gly Thr
1685 1690 1695Ala Met Ala Glu Arg Leu Val Ile Gly His Gly Ser Arg
Val Leu 1700 1705 1710Gln Phe Ser Ser Ser Gly Phe Asp Ala Ser Val
Met Asp Gln Leu 1715 1720 1725Met Ala Phe Gly Ala Gly Ala Ala Leu
Val Val Pro Gly Pro Glu 1730 1735 1740Gln Leu Leu Gly Thr Glu Leu
Ala Asp Leu Leu Glu Lys Gln Ala 1745 1750 1755Val Ser His Ala Leu
Ile Pro Pro Ala Ala Leu Ala Thr Leu Pro 1760 1765 1770His Gly Glu
Phe Pro His Leu Gln Thr Leu Val Val Gly Gly Asp 1775 1780 1785Ala
Cys Thr Ala Ala Leu Ala Ala Lys Trp Ser Gln Gly Arg Arg 1790 1795
1800Met Ile Asn Ala Tyr Gly Pro Thr Glu Ile Thr Ile Cys Ala Ser
1805 1810 1815Met Ser Ala Pro Met Thr Ala Glu Glu Leu Pro Ser Ile
Gly Gln 1820 1825 1830Pro Ile Trp Asn Thr Arg Met Tyr Val Leu Asp
Ser Ala Leu Gln 1835 1840 1845Pro Val Pro Pro Gly Val Ala Gly Glu
Leu Tyr Ile Ala Gly Ser 1850 1855 1860Gly Val Ala Arg Gly Tyr Leu
Asn Arg Pro Ala Leu Ser Ala Glu 1865 1870 1875Arg Phe Ile Ala Asp
Pro His Gly Ala Pro Gly Ser Arg Met Tyr 1880 1885 1890Arg Ser Gly
Asp Leu Ala Arg Trp Arg Ala Asp Gly Thr Leu Asp 1895 1900 1905Phe
Leu Gly Arg Ala Asp Gln Gln Val Lys Ile Arg Gly Phe Arg 1910 1915
1920Ile Glu Pro Gly Glu Ile Glu Ser Val Leu Leu Lys His Pro Leu
1925 1930 1935Ile Thr Gln Ala Ala Val Ile Ala Arg Glu Asp Val Pro
Gly Glu 1940 1945 1950Lys Arg Leu Val Ala Tyr Phe Val Ala Gly Ser
Glu Pro Gln Pro 1955 1960 1965Thr Glu Leu Arg Ala His Met Ala Gln
Ala Leu Pro Asp Tyr Met 1970 1975 1980Val Pro Ser Ala Phe Val Arg
Leu Pro Ser Leu Pro Leu Thr Gln 1985 1990 1995Ser Gly Lys Leu Asp
Lys Lys Ala Leu Pro Val Pro Asp Gln Gln 2000 2005 2010Pro Ala Ala
Leu Tyr Val Glu Pro Arg Thr Pro Thr Glu Lys Leu 2015 2020 2025Leu
Ala Gly Leu Trp Ser Glu Thr Leu His Leu Glu Arg Val Gly 2030 2035
2040Ile His Asp Asn Phe Phe Glu Ile Gly Gly His Ser Leu Met Ala
2045 2050 2055Ile Gln Leu Gly Met Arg Ile Arg Gln Gln Val Arg Ala
Asp Phe 2060 2065 2070Pro His Ala Glu Val Tyr Asn Arg Pro Thr Ile
Ala Asp Leu Ala 2075 2080 2085Ala Trp Leu Asp Asn Glu Gly Gly Thr
Val Glu Ala Leu Asp Leu 2090 2095 2100Ser Arg Glu Leu Asp Leu Pro
Ala His Ile Arg Pro Gln Ala Thr 2105 2110 2115Ala Pro Lys Leu Ala
Pro Arg Arg Val Phe Leu Thr Gly Ala Ser 2120 2125 2130Gly Phe Val
Gly Ser His Leu Leu Ala Ala Leu Leu Arg Asp Thr 2135 2140 2145Ala
Ala Cys Val Val Cys His Val Arg Ala Pro Asp Glu Gln Ala 2150 2155
2160Gly Glu Gln Arg Leu Lys Arg Thr Leu Ala Gln Arg Gln Leu Gly
2165 2170 2175Ala Ile Trp Asp Asn Ala Arg Ile Lys Val Val Thr Gly
Asp Leu 2180 2185 2190Gly Lys Pro Arg Leu Gly Leu Asp Asp Ala Ala
Val Gln Leu Val 2195 2200 2205Arg Asp Gly Cys Asp Ala Ile Tyr His
Cys Ala Ala Gln Val Asp 2210 2215 2220Phe Leu His Pro Tyr Ala Ser
Leu Lys Pro Ala Asn Val Asp Ser 2225 2230 2235Val Val Thr Leu Leu
Glu Trp Thr Ala Gln Gly Arg Ala Lys Ser 2240 2245 2250Met His Tyr
Val Ser Thr Leu Ala Val Ile Asp Gln Asn Asn Lys 2255 2260 2265Glu
Asp Thr Ile Thr Glu Gln Ser Ala Leu Ala Ser Trp Ser Gly 2270 2275
2280Leu Val Asp Gly Tyr Ser Gln Ser Lys Trp Val Gly Asp Ala Leu
2285 2290 2295Ala Arg Glu Ala Gln Ala Arg Gly Met Pro Val Ala Ile
Tyr Arg 2300 2305 2310Leu Gly Ala Val Thr Gly Asp His Thr His Ala
Ile Cys Asn Ala 2315 2320 2325Asp Asp Leu Ile Trp Arg Val Ala His
Leu Tyr Ala Asp Leu Glu 2330 2335 2340Ala Ile Pro Asp Met Asp Leu
Pro Leu Asn Leu Thr Pro Val Asp 2345 2350 2355Asp Val Ala Arg Ala
Ile Leu Gly Leu Ala Ala Gln Glu Ala Ser 2360 2365 2370Trp Gly Gln
Val Phe His Leu Met Ser Gln Ala Ala Leu Arg Val 2375 2380 2385Arg
Asp Ile Pro His Val Phe Glu Arg Met Gly Met Arg Leu Glu 2390 2395
2400Pro Val Gly Leu Glu Pro Trp Leu Gln Arg Ala His Ala Arg Leu
2405 2410 2415Ala Val Ala His Asp Arg Asp Leu Ala Ala Val Leu Ala
Ile Leu 2420 2425 2430Asp Arg Tyr Asp Thr Thr Ala Thr Pro Pro Gln
Val Ser Gly Ala 2435 2440 2445Ala Thr His Ala Gln Leu Glu Ala Ile
Gly Ala Pro Ile Arg Pro 2450 2455 2460Val Asp Arg Asp Leu Leu Gln
Arg Tyr Phe Val Asp Leu Gly Ile 2465 2470 2475Asp Thr Lys Ala Arg
Arg Ala Leu Glu Thr Thr Thr Ser 2480 2485 249071320DNAVariovorax
paradoxus 7atggcacgct atctcatcgc agcaaccgcc ttgccgggac acgtcctgcc
gatgctggcc 60atcgcgcagc atctggtgaa ccaggggcac gaggtgcggg tgcacaccgc
gagccagttc 120agggcgcagg ccgaggcgac cggtgcgggc ttcacgccct
tcgagcgcac gatcgacttc 180gactaccgcg acctggacaa gcgctttccc
gagcgccagc gcatcgcctc ggcgcatgcg 240cagctgtgct tcggcctgaa
gcacttcttt gccgatgcga tggccgcgca gcatgcgggc 300ctgcaatcga
tcctcgaaga cttcgaggcc gatgccatcg tggtcgacac gatgttctgc
360ggcactttcc cgctgctgct aggcaaggag cgcgaagacc gcccggccat
cgtcggcatc 420ggcatctcgg cgctgccgct ctcgagctgc gacaccgcct
tcttcggcac cgcgctgccg 480ccgtcgtcca cgccggaagg gcgggtgcgc
aacaaggcga tgaacgccaa cctcaaacag 540gcgatgttcg gcgaggtgca
acgctacttc gacacgctgc tcgcgcgttc gggcctggcc 600gcgctgcccg
atttcttcgt cgatgcgatg gtgaagctgc ccgatcttta cctgcagctc
660accgcgcctt cgttcgaata cccgcgcagc gacctgcccg cgtcggtgca
tttcgtcggc 720ccgctgctct cgcccgcgag ccgcgacttc acgccgcccg
agtggtggca cgagctggac 780gacggccgct cggtcgtgct ggtcacgcag
ggcacgctgg ccaaccagaa tccgtcgcag 840ctgatcggcc cgacgctgca
ggcgctggcc ggcgacaaga acatcctcgt catcgccacc 900accggcggcc
cggtgccgcc cgccctgacg gtgaacctgc ccgccaacgc ccgcgtggtg
960ccgttcctgc cctacgaccg gctgctgccc aagctgcacg cgatggtcac
caacggcggc 1020tacggctcgg tcaaccatgc attgagcctc ggtgtgccgc
tggtggtggc cggcacctcc 1080gaagagaagc ccgagatcgc cgcgcgcgtg
gcctggtcgg gcgcgggcat caacctcgcc 1140accggccagc cgaccgcgcg
ccaggtcggc gacgcggtgc gcaaggtact gggcaactcg 1200acctatcgcc
agcgtgcggc ggtgctgcgt gaggacttcg cttgccatcg cgcgctgacc
1260ggcatcgccg gcgccctcga ggcacttctg caaaccttcg catccgcgga
aatggcttga 13208439PRTVariovorax paradoxus 8Met Ala Arg Tyr Leu Ile
Ala Ala Thr Ala Leu Pro Gly His Val Leu1 5 10 15Pro Met Leu Ala Ile
Ala Gln His Leu Val Asn Gln Gly His Glu Val 20 25 30Arg Val His Thr
Ala Ser Gln Phe Arg Ala Gln Ala Glu Ala Thr Gly 35 40 45Ala Gly Phe
Thr Pro Phe Glu Arg Thr Ile Asp Phe Asp Tyr Arg Asp 50 55 60Leu Asp
Lys Arg Phe Pro Glu Arg Gln Arg Ile Ala Ser Ala His Ala65 70 75
80Gln Leu Cys Phe Gly Leu Lys His Phe Phe Ala Asp Ala Met Ala Ala
85 90 95Gln His Ala Gly Leu Gln Ser Ile Leu Glu Asp Phe Glu Ala Asp
Ala 100 105 110Ile Val Val Asp Thr Met Phe Cys Gly Thr Phe Pro Leu
Leu Leu Gly 115 120 125Lys Glu Arg Glu Asp Arg Pro Ala Ile Val Gly
Ile Gly Ile Ser Ala 130 135 140Leu Pro Leu Ser Ser Cys Asp Thr Ala
Phe Phe Gly Thr Ala Leu Pro145 150 155 160Pro Ser Ser Thr Pro Glu
Gly Arg Val Arg Asn Lys Ala Met Asn Ala 165 170 175Asn Leu Lys Gln
Ala Met Phe Gly Glu Val Gln Arg Tyr Phe Asp Thr 180 185 190Leu Leu
Ala Arg Ser Gly Leu Ala Ala Leu Pro Asp Phe Phe Val Asp 195 200
205Ala Met Val Lys Leu Pro Asp
Leu Tyr Leu Gln Leu Thr Ala Pro Ser 210 215 220Phe Glu Tyr Pro Arg
Ser Asp Leu Pro Ala Ser Val His Phe Val Gly225 230 235 240Pro Leu
Leu Ser Pro Ala Ser Arg Asp Phe Thr Pro Pro Glu Trp Trp 245 250
255His Glu Leu Asp Asp Gly Arg Ser Val Val Leu Val Thr Gln Gly Thr
260 265 270Leu Ala Asn Gln Asn Pro Ser Gln Leu Ile Gly Pro Thr Leu
Gln Ala 275 280 285Leu Ala Gly Asp Lys Asn Ile Leu Val Ile Ala Thr
Thr Gly Gly Pro 290 295 300Val Pro Pro Ala Leu Thr Val Asn Leu Pro
Ala Asn Ala Arg Val Val305 310 315 320Pro Phe Leu Pro Tyr Asp Arg
Leu Leu Pro Lys Leu His Ala Met Val 325 330 335Thr Asn Gly Gly Tyr
Gly Ser Val Asn His Ala Leu Ser Leu Gly Val 340 345 350Pro Leu Val
Val Ala Gly Thr Ser Glu Glu Lys Pro Glu Ile Ala Ala 355 360 365Arg
Val Ala Trp Ser Gly Ala Gly Ile Asn Leu Ala Thr Gly Gln Pro 370 375
380Thr Ala Arg Gln Val Gly Asp Ala Val Arg Lys Val Leu Gly Asn
Ser385 390 395 400Thr Tyr Arg Gln Arg Ala Ala Val Leu Arg Glu Asp
Phe Ala Cys His 405 410 415Arg Ala Leu Thr Gly Ile Ala Gly Ala Leu
Glu Ala Leu Leu Gln Thr 420 425 430Phe Ala Ser Ala Glu Met Ala
4359213DNAVariovorax paradoxus 9atgagcaacc cgttcgacga caagaacgcc
agcttccagg tgctggtgaa cgacgagggc 60cagcactcgc tgtggcccgc cttcatcgcc
gtgcccgccg gctggcaggt ggcgctggcg 120ccgaccgacc gcgacgcctg
cagcgcctac atcgcggcga actggcagga catgcgcccg 180cgttcgctgg
tggtggccac ggcggccggc tga 2131070PRTVariovorax paradoxus 10Met Ser
Asn Pro Phe Asp Asp Lys Asn Ala Ser Phe Gln Val Leu Val1 5 10 15Asn
Asp Glu Gly Gln His Ser Leu Trp Pro Ala Phe Ile Ala Val Pro 20 25
30Ala Gly Trp Gln Val Ala Leu Ala Pro Thr Asp Arg Asp Ala Cys Ser
35 40 45Ala Tyr Ile Ala Ala Asn Trp Gln Asp Met Arg Pro Arg Ser Leu
Val 50 55 60Val Ala Thr Ala Ala Gly65 7011969DNAVariovorax
paradoxus 11atgtccttcc cgttcggtgc cgtcgtcgtc acctatttcc cgaccggcga
gcaagtggcg 60aacctccatt cgctggcggc ctcgtgtccg cacctctgcg tggtcgacaa
cacgccgcag 120gtgggcgatt ggcatgcggc gctcgtcgat gcgggcgttt
cggtgctgca caacggcaac 180cgcggcggca tcgcgggcgc cttcaaccgc
ggcatcatcg acctcgaagc gcggggcgcc 240gaactcttct tcctgctcga
ccaggattcg aagctgccac ccggctactt cgatgccatg 300tgcgaggctg
cgatggtggc ccgggagcgg aagggcgagg gcaatggtga ggaagacgcg
360gccttcctga tcggcccgct cgtccacgac acgaacctgg acgcgctgat
cccgcaattc 420ggcctccagg gcaaacgcgt ctaccagttc gacctgcggc
agcccttcac cgagccgctg 480atgcgctgcg ccttcatgat ttcctcgggc
tccctgattt cgcgcggcgc ctgggcccgg 540atcggccggt tcgacgagcg
ctatgtgatc gaccacgtgg acaccgacta ctgcatgcgt 600gccctgggtc
gcggcgtgcc gctctacctg aatccgcacg tcgtgctgcg gcaccagatt
660ggcgacatcc gtgcccggtc gctgttcggc tggaagatcc acttcatcaa
ctacccggcc 720gcgcggcgct actacatcgc gcgcaatgcc atcgatctct
cgcgggcgca tgtgcgcgcc 780tttcccgcga tcctgttcat caacgtttac
acgctcaagc agatcctgcc gatgctgatg 840ttcgagcgcg accgcttcaa
gaagaccatc gcgctgatgc tcggctgctt cgatggcctg 900ttcgggcggc
tcgggggcct cggcgaggtg catccgcgga tgggcaaata cctgggccgc 960agcgattga
96912322PRTVariovorax paradoxus 12Met Ser Phe Pro Phe Gly Ala Val
Val Val Thr Tyr Phe Pro Thr Gly1 5 10 15Glu Gln Val Ala Asn Leu His
Ser Leu Ala Ala Ser Cys Pro His Leu 20 25 30Cys Val Val Asp Asn Thr
Pro Gln Val Gly Asp Trp His Ala Ala Leu 35 40 45Val Asp Ala Gly Val
Ser Val Leu His Asn Gly Asn Arg Gly Gly Ile 50 55 60Ala Gly Ala Phe
Asn Arg Gly Ile Ile Asp Leu Glu Ala Arg Gly Ala65 70 75 80Glu Leu
Phe Phe Leu Leu Asp Gln Asp Ser Lys Leu Pro Pro Gly Tyr 85 90 95Phe
Asp Ala Met Cys Glu Ala Ala Met Val Ala Arg Glu Arg Lys Gly 100 105
110Glu Gly Asn Gly Glu Glu Asp Ala Ala Phe Leu Ile Gly Pro Leu Val
115 120 125His Asp Thr Asn Leu Asp Ala Leu Ile Pro Gln Phe Gly Leu
Gln Gly 130 135 140Lys Arg Val Tyr Gln Phe Asp Leu Arg Gln Pro Phe
Thr Glu Pro Leu145 150 155 160Met Arg Cys Ala Phe Met Ile Ser Ser
Gly Ser Leu Ile Ser Arg Gly 165 170 175Ala Trp Ala Arg Ile Gly Arg
Phe Asp Glu Arg Tyr Val Ile Asp His 180 185 190Val Asp Thr Asp Tyr
Cys Met Arg Ala Leu Gly Arg Gly Val Pro Leu 195 200 205Tyr Leu Asn
Pro His Val Val Leu Arg His Gln Ile Gly Asp Ile Arg 210 215 220Ala
Arg Ser Leu Phe Gly Trp Lys Ile His Phe Ile Asn Tyr Pro Ala225 230
235 240Ala Arg Arg Tyr Tyr Ile Ala Arg Asn Ala Ile Asp Leu Ser Arg
Ala 245 250 255His Val Arg Ala Phe Pro Ala Ile Leu Phe Ile Asn Val
Tyr Thr Leu 260 265 270Lys Gln Ile Leu Pro Met Leu Met Phe Glu Arg
Asp Arg Phe Lys Lys 275 280 285Thr Ile Ala Leu Met Leu Gly Cys Phe
Asp Gly Leu Phe Gly Arg Leu 290 295 300Gly Gly Leu Gly Glu Val His
Pro Arg Met Gly Lys Tyr Leu Gly Arg305 310 315 320Ser
Asp131260DNAVariovorax paradoxus 13ttgaccgcca cccttccagc gccgcgcgta
cgccgcgccg cgctcgcctt catcttcgtc 60acggtgctga tcgacttcat ggcgttcggc
ctgatcctgc ccggcctgcc gcacctggtg 120gagcggctgg ccggcggcag
cacggtaacg gcggcgtact ggatcgctgt gttcggcacc 180gcgttcgcgg
cgatccagtt cgtgagctcg ccgatccagg gcgcgctgtc cgaccgcttc
240gggcggcggc cggtgatcct gctgtcgtgc ttcggcctcg gcgtggattt
cgtgttcatg 300gccctggccg acagcctgcc gtggctgttc gtcggccggg
tggtctccgg cgtgttctcg 360gccagcttca ccatcgccaa tgcctacatc
gccgatgtga cgctgccgga ggagcgcgcc 420cgcagctacg gcatcgtggg
ggccgcgttc ggcatgggcc tggtgttcgg gccggtgctc 480ggcgggcaac
tgagccacat cgatccgcgc ctgccgttct ggttcgcggc cggcttgacg
540ctgctcagct tctgctacgg atggttcgtg ttgcccgaat cgctgccgcc
cgagcggcgt 600gcccgcaagt tcgactggtc gcatgccaat ccggttggga
cgctggtgct gctcaagcgc 660tatccgcagg tgttcggact ggcggcggtg
atcttcctcg tgaacctggc tcagtacgtc 720tatcccagcg tgttcgtgct
gttcgccgac taccggtatc actggaagga agacgccgtg 780ggctgggtgc
tcggcgcggt gggcgtgctc agcgtgctgg tcaatgcgct gttgatcggg
840ccgggcgtga agcgcttcgg cgagcgccgc gccctgttgc tcggcatggg
cttcggcgtg 900ctcggcttcg tcatcatcgg gtttgccgac gctggatgga
tcctcctggc cggggtgccg 960ttcggcattc tgctggcgtt cgccggaccg
gcggcgcagg cgctggtcac gctgcaggtc 1020ggcaccgccg agcagggccg
catccagggg gcgctcacca gcctggtgtc ggtggcgggc 1080atcgtcgggc
cggcgatgtt cgccggcagc ttcggttact tcatcggcgc ggacgcgccg
1140gtgcacttgc cgggcgcgcc gtttttcctc gctgcggcgt tcctctgcat
cggcacgctg 1200atcgcgtggc gctacgcaca gccgaagccc gcgacggcag
cggtgcccga gccgacctga 126014419PRTVariovorax paradoxus 14Met Thr
Ala Thr Leu Pro Ala Pro Arg Val Arg Arg Ala Ala Leu Ala1 5 10 15Phe
Ile Phe Val Thr Val Leu Ile Asp Phe Met Ala Phe Gly Leu Ile 20 25
30Leu Pro Gly Leu Pro His Leu Val Glu Arg Leu Ala Gly Gly Ser Thr
35 40 45Val Thr Ala Ala Tyr Trp Ile Ala Val Phe Gly Thr Ala Phe Ala
Ala 50 55 60Ile Gln Phe Val Ser Ser Pro Ile Gln Gly Ala Leu Ser Asp
Arg Phe65 70 75 80Gly Arg Arg Pro Val Ile Leu Leu Ser Cys Phe Gly
Leu Gly Val Asp 85 90 95Phe Val Phe Met Ala Leu Ala Asp Ser Leu Pro
Trp Leu Phe Val Gly 100 105 110Arg Val Val Ser Gly Val Phe Ser Ala
Ser Phe Thr Ile Ala Asn Ala 115 120 125Tyr Ile Ala Asp Val Thr Leu
Pro Glu Glu Arg Ala Arg Ser Tyr Gly 130 135 140Ile Val Gly Ala Ala
Phe Gly Met Gly Leu Val Phe Gly Pro Val Leu145 150 155 160Gly Gly
Gln Leu Ser His Ile Asp Pro Arg Leu Pro Phe Trp Phe Ala 165 170
175Ala Gly Leu Thr Leu Leu Ser Phe Cys Tyr Gly Trp Phe Val Leu Pro
180 185 190Glu Ser Leu Pro Pro Glu Arg Arg Ala Arg Lys Phe Asp Trp
Ser His 195 200 205Ala Asn Pro Val Gly Thr Leu Val Leu Leu Lys Arg
Tyr Pro Gln Val 210 215 220Phe Gly Leu Ala Ala Val Ile Phe Leu Val
Asn Leu Ala Gln Tyr Val225 230 235 240Tyr Pro Ser Val Phe Val Leu
Phe Ala Asp Tyr Arg Tyr His Trp Lys 245 250 255Glu Asp Ala Val Gly
Trp Val Leu Gly Ala Val Gly Val Leu Ser Val 260 265 270Leu Val Asn
Ala Leu Leu Ile Gly Pro Gly Val Lys Arg Phe Gly Glu 275 280 285Arg
Arg Ala Leu Leu Leu Gly Met Gly Phe Gly Val Leu Gly Phe Val 290 295
300Ile Ile Gly Phe Ala Asp Ala Gly Trp Ile Leu Leu Ala Gly Val
Pro305 310 315 320Phe Gly Ile Leu Leu Ala Phe Ala Gly Pro Ala Ala
Gln Ala Leu Val 325 330 335Thr Leu Gln Val Gly Thr Ala Glu Gln Gly
Arg Ile Gln Gly Ala Leu 340 345 350Thr Ser Leu Val Ser Val Ala Gly
Ile Val Gly Pro Ala Met Phe Ala 355 360 365Gly Ser Phe Gly Tyr Phe
Ile Gly Ala Asp Ala Pro Val His Leu Pro 370 375 380Gly Ala Pro Phe
Phe Leu Ala Ala Ala Phe Leu Cys Ile Gly Thr Leu385 390 395 400Ile
Ala Trp Arg Tyr Ala Gln Pro Lys Pro Ala Thr Ala Ala Val Pro 405 410
415Glu Pro Thr151080DNAVariovorax paradoxus 15atgatcctgg taaccggcgg
cgcaggcttc attggcgcca atttcgtact cgactggctc 60gcacagagcg atgaaccggt
cgtgaaccta gacaagctga cctacgcggg caacctcgag 120acgctcgcat
cgctcaagga caacccgaag cacatcttcg tgcagggcga catcggcgac
180agcgcgctgc tcgaccgcct gctggccgag cacaagccgc gtgccgtggt
caacttcgcg 240gccgaatcgc acgtcgaccg ctcgatccac ggccccgaag
acttcgtgca gaccaacgtg 300ctgggcacct tccgcctgct cgaatccgtg
cgcggtttct ggaatgccct gccggccgac 360cagaaggccg ccttccgctt
cctgcatgtg tcgaccgacg aggtctacgg ctcgctctcc 420aagaccgacc
cggccttcac cgaagagaac aagtacgagc ccaacagccc gtactcggcc
480agcaaggccg ccagcgacca cctcgtgcgc gcctggcacc acacctacgg
cctgccggtg 540gtcaccacca actgctcgaa caactacggg ccgttccact
tccccgagaa gctcattccc 600ctgatgatcg tcaacgcgct ggcgggcaag
ccgctgcccg tgtacggcga cggcatgcag 660gtgcgcgact ggctctacgt
gaaggaccac tgcagcgcca tccgccgcgt gctcgaagcc 720ggcaagctcg
gcgagaccta caacgtgggc ggctggaacg agaagcccaa catcgagatc
780gtcaacaccg tctgcgcgct gctcgacgag ctgagcccca aggccggcgg
caagccgtac 840aaggaacaga tcacctatgt gaccgaccgc cccggccacg
accgccgcta cgcgatcgac 900gcacgcaagc tcgagcgcga actcggctgg
aaacctgccg agaccttcga cagcggcatc 960cgcaagacgg tcgagtggta
cctcgcgaac ggcgagtggg tgcgcaacgt gcaaagcggc 1020gcgtaccgcg
agtgggtcga gaagcaatac gacgccgcac cggcgaaggc caccgcatga
108016359PRTVariovorax paradoxus 16Met Ile Leu Val Thr Gly Gly Ala
Gly Phe Ile Gly Ala Asn Phe Val1 5 10 15Leu Asp Trp Leu Ala Gln Ser
Asp Glu Pro Val Val Asn Leu Asp Lys 20 25 30Leu Thr Tyr Ala Gly Asn
Leu Glu Thr Leu Ala Ser Leu Lys Asp Asn 35 40 45Pro Lys His Ile Phe
Val Gln Gly Asp Ile Gly Asp Ser Ala Leu Leu 50 55 60Asp Arg Leu Leu
Ala Glu His Lys Pro Arg Ala Val Val Asn Phe Ala65 70 75 80Ala Glu
Ser His Val Asp Arg Ser Ile His Gly Pro Glu Asp Phe Val 85 90 95Gln
Thr Asn Val Leu Gly Thr Phe Arg Leu Leu Glu Ser Val Arg Gly 100 105
110Phe Trp Asn Ala Leu Pro Ala Asp Gln Lys Ala Ala Phe Arg Phe Leu
115 120 125His Val Ser Thr Asp Glu Val Tyr Gly Ser Leu Ser Lys Thr
Asp Pro 130 135 140Ala Phe Thr Glu Glu Asn Lys Tyr Glu Pro Asn Ser
Pro Tyr Ser Ala145 150 155 160Ser Lys Ala Ala Ser Asp His Leu Val
Arg Ala Trp His His Thr Tyr 165 170 175Gly Leu Pro Val Val Thr Thr
Asn Cys Ser Asn Asn Tyr Gly Pro Phe 180 185 190His Phe Pro Glu Lys
Leu Ile Pro Leu Met Ile Val Asn Ala Leu Ala 195 200 205Gly Lys Pro
Leu Pro Val Tyr Gly Asp Gly Met Gln Val Arg Asp Trp 210 215 220Leu
Tyr Val Lys Asp His Cys Ser Ala Ile Arg Arg Val Leu Glu Ala225 230
235 240Gly Lys Leu Gly Glu Thr Tyr Asn Val Gly Gly Trp Asn Glu Lys
Pro 245 250 255Asn Ile Glu Ile Val Asn Thr Val Cys Ala Leu Leu Asp
Glu Leu Ser 260 265 270Pro Lys Ala Gly Gly Lys Pro Tyr Lys Glu Gln
Ile Thr Tyr Val Thr 275 280 285Asp Arg Pro Gly His Asp Arg Arg Tyr
Ala Ile Asp Ala Arg Lys Leu 290 295 300Glu Arg Glu Leu Gly Trp Lys
Pro Ala Glu Thr Phe Asp Ser Gly Ile305 310 315 320Arg Lys Thr Val
Glu Trp Tyr Leu Ala Asn Gly Glu Trp Val Arg Asn 325 330 335Val Gln
Ser Gly Ala Tyr Arg Glu Trp Val Glu Lys Gln Tyr Asp Ala 340 345
350Ala Pro Ala Lys Ala Thr Ala 35517891DNAVariovorax paradoxus
17atgaagctgc tgctgctggg caagggcgga caggtcggct gggagctgca acgcagcctc
60gcgcccctgg gcgaactggt ggcgctcgat ttcgacagca ccgacttcaa cgccgacttc
120agtcgccccg agcagctggc cgagacagtg ctgaaggtgc gccccgacgt
catcgtcaat 180gccgcagcgc acaccgcggt cgacaaggcc gagagcgagc
ccgagttcgc gcgcaagctc 240aacgccacct cgcccggcgt ggtggccgaa
gccgcgcagc agatcggcgc gctgatggtt 300cactactcga ccgactacgt
cttcgacggc agcggcagca agccgtggaa agaagacgat 360gcgaccggcc
cgctcagcgt ctacggcagc accaagctcg aaggcgagca actggtggca
420aagcactgtg cgaagcacct gatctttcgc accagctggg tctatgccgc
gcgcggcggc 480aacttcgcca agaccatgct gcgcatcgcc aaggagcgcg
acaagctgac cgtcatcgac 540gaccagttcg gcgcgcccac cggcgcggaa
ctgctggccg acatcaccgc gcacgcgatt 600cgcgcgacgc tgcaggaccc
gtccaaggcc gggctctatc acgcggtggc cggtggcgtg 660accacgtggc
acggctatgc gcgcttcgtg atcgagcagg ccaaggcggc gggcgtggaa
720ctgaaggccg gccccgaagc ggtcgagccc gtgcccacca cggcattccc
gacgccggcc 780aggcggccgc acaactcgcg cctggacacc accaagctgc
aatcgacctt cggcctcgtg 840ctgcccgagt ggcagtccgg cgtcgcccgc
atgttgcgcg aaaccttctg a 89118296PRTVariovorax paradoxus 18Met Lys
Leu Leu Leu Leu Gly Lys Gly Gly Gln Val Gly Trp Glu Leu1 5 10 15Gln
Arg Ser Leu Ala Pro Leu Gly Glu Leu Val Ala Leu Asp Phe Asp 20 25
30Ser Thr Asp Phe Asn Ala Asp Phe Ser Arg Pro Glu Gln Leu Ala Glu
35 40 45Thr Val Leu Lys Val Arg Pro Asp Val Ile Val Asn Ala Ala Ala
His 50 55 60Thr Ala Val Asp Lys Ala Glu Ser Glu Pro Glu Phe Ala Arg
Lys Leu65 70 75 80Asn Ala Thr Ser Pro Gly Val Val Ala Glu Ala Ala
Gln Gln Ile Gly 85 90 95Ala Leu Met Val His Tyr Ser Thr Asp Tyr Val
Phe Asp Gly Ser Gly 100 105 110Ser Lys Pro Trp Lys Glu Asp Asp Ala
Thr Gly Pro Leu Ser Val Tyr 115 120 125Gly Ser Thr Lys Leu Glu Gly
Glu Gln Leu Val Ala Lys His Cys Ala 130 135 140Lys His Leu Ile Phe
Arg Thr Ser Trp Val Tyr Ala Ala Arg Gly Gly145 150 155 160Asn Phe
Ala Lys Thr Met Leu Arg Ile Ala Lys Glu Arg Asp Lys Leu 165 170
175Thr Val Ile Asp Asp Gln Phe Gly Ala Pro Thr Gly Ala Glu Leu Leu
180 185 190Ala Asp Ile Thr Ala His Ala Ile Arg Ala Thr Leu Gln Asp
Pro Ser 195 200 205Lys Ala Gly Leu Tyr His Ala Val Ala Gly Gly Val
Thr Thr Trp His 210 215 220Gly Tyr Ala Arg Phe Val Ile Glu Gln Ala
Lys Ala Ala Gly Val Glu225 230 235 240Leu Lys Ala Gly Pro Glu Ala
Val Glu Pro Val Pro Thr Thr Ala Phe 245 250 255Pro Thr Pro Ala Arg
Arg Pro His Asn Ser Arg Leu Asp Thr Thr Lys 260 265 270Leu Gln Ser
Thr Phe Gly Leu Val Leu Pro Glu Trp Gln Ser Gly Val 275 280 285Ala
Arg Met Leu Arg Glu Thr Phe 290
29519897DNAVariovorax paradoxus 19atgaccaaga cgacgcaacg caaaggcatc
atcctcgccg gtggctcggg cacccgcctg 60caccccgcga cgcttgccat gagcaaacaa
ctgctgccgg tgtacgacaa gccgatgatc 120tattacccgc tgagcacgct
gatgctgggc ggcatgcgcg acatcctgat catcagcacg 180ccgcaggaca
cgccgcgttt ccagcaactg ctgggggatg gcagccaatg gggcatcaac
240ctgcagtacg cggtgcagcc gagcccggat ggtctggcgc aggcgttcat
catcggtgac 300aagttcgtgg gcaacgaccc gagtgcgctg gtgctggggg
acaacatctt ctatggccac 360gacttcgccc atctgctggc cgatgccgac
gccaagacct cgggtgcgac ggtgttcgcc 420taccacgtgc acgaccccga
gcgctacggc gtggtggcct tcgatgccaa gggcagggcg 480agcagcatcg
aagaaaagcc gctcaagccc aagagcagct atgcggtcac gggcctctac
540ttctacgaca accaggtcgt cgacatcgcc aaggccgtga agccgagcgc
gcgcggcgaa 600ctcgagatca ccgcggtcaa ccaggcgtat ctcgacctcg
accagctgaa cgtgcagatc 660atgcagcgcg gctatgcgtg gctcgatacc
ggtacgcacg acagcctgct ggaagccggg 720cagttcattg ccacgctcga
gcaccgccag gggctgaaga tcgcatgccc cgaagagatc 780gcatggcgca
atggcttcat ctcaaccgag caactcgaaa agctcgcggc gccgctggaa
840aagagcggct acggcaagta cctcaagcac ctgctgaacg acgaggtgcg ctcgtga
89720298PRTVariovorax paradoxus 20Met Thr Lys Thr Thr Gln Arg Lys
Gly Ile Ile Leu Ala Gly Gly Ser1 5 10 15Gly Thr Arg Leu His Pro Ala
Thr Leu Ala Met Ser Lys Gln Leu Leu 20 25 30Pro Val Tyr Asp Lys Pro
Met Ile Tyr Tyr Pro Leu Ser Thr Leu Met 35 40 45Leu Gly Gly Met Arg
Asp Ile Leu Ile Ile Ser Thr Pro Gln Asp Thr 50 55 60Pro Arg Phe Gln
Gln Leu Leu Gly Asp Gly Ser Gln Trp Gly Ile Asn65 70 75 80Leu Gln
Tyr Ala Val Gln Pro Ser Pro Asp Gly Leu Ala Gln Ala Phe 85 90 95Ile
Ile Gly Asp Lys Phe Val Gly Asn Asp Pro Ser Ala Leu Val Leu 100 105
110Gly Asp Asn Ile Phe Tyr Gly His Asp Phe Ala His Leu Leu Ala Asp
115 120 125Ala Asp Ala Lys Thr Ser Gly Ala Thr Val Phe Ala Tyr His
Val His 130 135 140Asp Pro Glu Arg Tyr Gly Val Val Ala Phe Asp Ala
Lys Gly Arg Ala145 150 155 160Ser Ser Ile Glu Glu Lys Pro Leu Lys
Pro Lys Ser Ser Tyr Ala Val 165 170 175Thr Gly Leu Tyr Phe Tyr Asp
Asn Gln Val Val Asp Ile Ala Lys Ala 180 185 190Val Lys Pro Ser Ala
Arg Gly Glu Leu Glu Ile Thr Ala Val Asn Gln 195 200 205Ala Tyr Leu
Asp Leu Asp Gln Leu Asn Val Gln Ile Met Gln Arg Gly 210 215 220Tyr
Ala Trp Leu Asp Thr Gly Thr His Asp Ser Leu Leu Glu Ala Gly225 230
235 240Gln Phe Ile Ala Thr Leu Glu His Arg Gln Gly Leu Lys Ile Ala
Cys 245 250 255Pro Glu Glu Ile Ala Trp Arg Asn Gly Phe Ile Ser Thr
Glu Gln Leu 260 265 270Glu Lys Leu Ala Ala Pro Leu Glu Lys Ser Gly
Tyr Gly Lys Tyr Leu 275 280 285Lys His Leu Leu Asn Asp Glu Val Arg
Ser 290 29521546DNAVariovorax paradoxus 21gtgaaggcca cgcccacctc
gattcctgac gtgctcgtga tcgagccgaa ggtgtttggc 60gatgcacggg gcttcttctt
cgaaagcttc aaccagaagg ccttcgacga agcgatcggc 120aagcatgtcg
acttcgtgca ggacaaccat tcgcgatcgg ccaagggtgt gctgcggggg
180ctgcattacc aggtccagca gccgcaaggc aagctcgtgc gggtggtgcg
tggtgcggtg 240ttcgacgtgg ccgtcgacat ccgcaagtcg tcgccgactt
ttggcaaatg ggtgggtgtc 300gagttgaacg aagacaacca caagcagctc
tgggtgccgg caggattcgc gcacggtttc 360ctggtgttga gcgagaccgc
ggaattcctc tacaagacca ccgactacta cgcgcccgcc 420cacgagcgcg
cgattgtctg gaacgacccc gctgtcggta ttcgatggcc ggatgtggga
480ggggcaccgg tcctgtcgaa gaaggacgaa gacgggtgtc ttctgcaagc
ggcagaggtt 540ttctag 54622181PRTVariovorax paradoxus 22Met Lys Ala
Thr Pro Thr Ser Ile Pro Asp Val Leu Val Ile Glu Pro1 5 10 15Lys Val
Phe Gly Asp Ala Arg Gly Phe Phe Phe Glu Ser Phe Asn Gln 20 25 30Lys
Ala Phe Asp Glu Ala Ile Gly Lys His Val Asp Phe Val Gln Asp 35 40
45Asn His Ser Arg Ser Ala Lys Gly Val Leu Arg Gly Leu His Tyr Gln
50 55 60Val Gln Gln Pro Gln Gly Lys Leu Val Arg Val Val Arg Gly Ala
Val65 70 75 80Phe Asp Val Ala Val Asp Ile Arg Lys Ser Ser Pro Thr
Phe Gly Lys 85 90 95Trp Val Gly Val Glu Leu Asn Glu Asp Asn His Lys
Gln Leu Trp Val 100 105 110Pro Ala Gly Phe Ala His Gly Phe Leu Val
Leu Ser Glu Thr Ala Glu 115 120 125Phe Leu Tyr Lys Thr Thr Asp Tyr
Tyr Ala Pro Ala His Glu Arg Ala 130 135 140Ile Val Trp Asn Asp Pro
Ala Val Gly Ile Arg Trp Pro Asp Val Gly145 150 155 160Gly Ala Pro
Val Leu Ser Lys Lys Asp Glu Asp Gly Cys Leu Leu Gln 165 170 175Ala
Ala Glu Val Phe 180231029DNAVariovorax paradoxus 23atgggcagca
gccatcatca tcatcatcac agcagcggcc tggtgccgcg cggcagccat 60atgtccttcc
cgttcggtgc cgtcgtcgtc acctatttcc cgaccggcga gcaagtggcg
120aacctccatt cgctggcggc ctcgtgtccg cacctctgcg tggtcgacaa
cacgccgcag 180gtgggcgatt ggcatgcggc gctcgtcgat gcgggcgttt
cggtgctgca caacggcaac 240cgcggcggca tcgcgggcgc cttcaaccgc
ggcatcatcg acctcgaagc gcggggcgcc 300gaactcttct tcctgctcga
ccaggattcg aagctgccac ccggctactt cgatgccatg 360tgcgaggctg
cgatggtggc ccgggagcgg aagggcgagg gcaatggtga ggaagacgcg
420gccttcctga tcggcccgct cgtccacgac acgaacctgg acgcgctgat
cccgcaattc 480ggcctccagg gcaaacgcgt ctaccagttc gacctgcggc
agcccttcac cgagccgctg 540atgcgctgcg ccttcatgat ttcctcgggc
tccctgattt cgcgcggcgc ctgggcccgg 600atcggccggt tcgacgagcg
ctatgtgatc gaccacgtgg acaccgacta ctgcatgcgt 660gccctgggtc
gcggcgtgcc gctctacctg aatccgcacg tcgtgctgcg gcaccagatt
720ggcgacatcc gtgcccggtc gctgttcggc tggaagatcc acttcatcaa
ctacccggcc 780gcgcggcgct actacatcgc gcgcaatgcc atcgatctct
cgcgggcgca tgtgcgcgcc 840tttcccgcga tcctgttcat caacgtttac
acgctcaagc agatcctgcc gatgctgatg 900ttcgagcgcg accgcttcaa
gaagaccatc gcgctgatgc tcggctgctt cgatggcctg 960ttcgggcggc
tcgggggcct cggcgaggtg catccgcgga tgggcaaata cctgggccgc
1020agcgattga 102924342PRTVariovorax paradoxus 24Met Gly Ser Ser
His His His His His His Ser Ser Gly Leu Val Pro1 5 10 15Arg Gly Ser
His Met Ser Phe Pro Phe Gly Ala Val Val Val Thr Tyr 20 25 30Phe Pro
Thr Gly Glu Gln Val Ala Asn Leu His Ser Leu Ala Ala Ser 35 40 45Cys
Pro His Leu Cys Val Val Asp Asn Thr Pro Gln Val Gly Asp Trp 50 55
60His Ala Ala Leu Val Asp Ala Gly Val Ser Val Leu His Asn Gly Asn65
70 75 80Arg Gly Gly Ile Ala Gly Ala Phe Asn Arg Gly Ile Ile Asp Leu
Glu 85 90 95Ala Arg Gly Ala Glu Leu Phe Phe Leu Leu Asp Gln Asp Ser
Lys Leu 100 105 110Pro Pro Gly Tyr Phe Asp Ala Met Cys Glu Ala Ala
Met Val Ala Arg 115 120 125Glu Arg Lys Gly Glu Gly Asn Gly Glu Glu
Asp Ala Ala Phe Leu Ile 130 135 140Gly Pro Leu Val His Asp Thr Asn
Leu Asp Ala Leu Ile Pro Gln Phe145 150 155 160Gly Leu Gln Gly Lys
Arg Val Tyr Gln Phe Asp Leu Arg Gln Pro Phe 165 170 175Thr Glu Pro
Leu Met Arg Cys Ala Phe Met Ile Ser Ser Gly Ser Leu 180 185 190Ile
Ser Arg Gly Ala Trp Ala Arg Ile Gly Arg Phe Asp Glu Arg Tyr 195 200
205Val Ile Asp His Val Asp Thr Asp Tyr Cys Met Arg Ala Leu Gly Arg
210 215 220Gly Val Pro Leu Tyr Leu Asn Pro His Val Val Leu Arg His
Gln Ile225 230 235 240Gly Asp Ile Arg Ala Arg Ser Leu Phe Gly Trp
Lys Ile His Phe Ile 245 250 255Asn Tyr Pro Ala Ala Arg Arg Tyr Tyr
Ile Ala Arg Asn Ala Ile Asp 260 265 270Leu Ser Arg Ala His Val Arg
Ala Phe Pro Ala Ile Leu Phe Ile Asn 275 280 285Val Tyr Thr Leu Lys
Gln Ile Leu Pro Met Leu Met Phe Glu Arg Asp 290 295 300Arg Phe Lys
Lys Thr Ile Ala Leu Met Leu Gly Cys Phe Asp Gly Leu305 310 315
320Phe Gly Arg Leu Gly Gly Leu Gly Glu Val His Pro Arg Met Gly Lys
325 330 335Tyr Leu Gly Arg Ser Asp 340
References












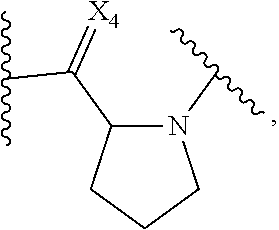

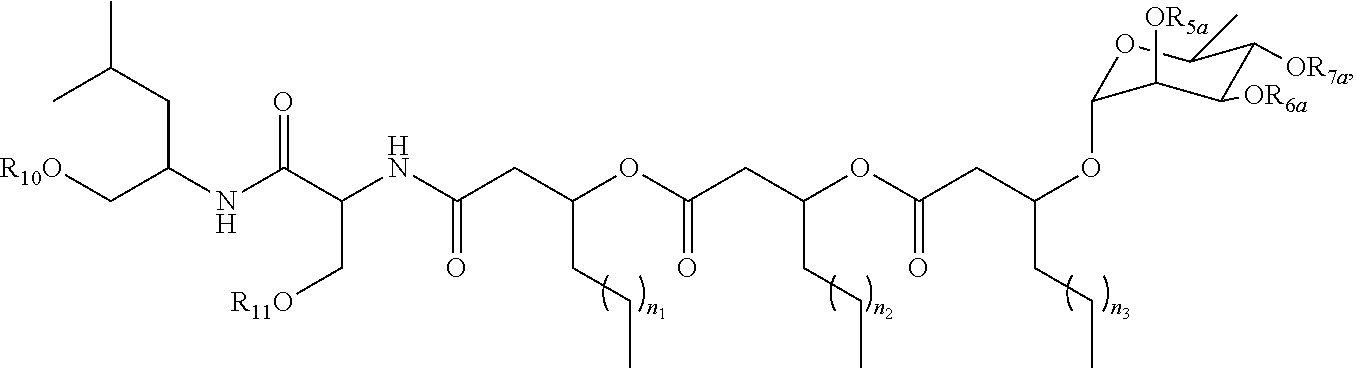
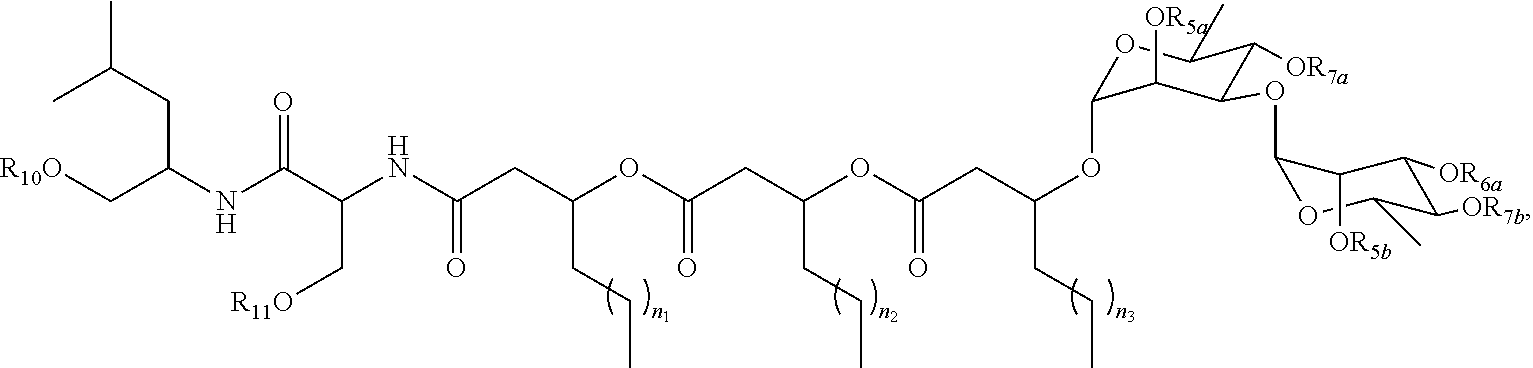
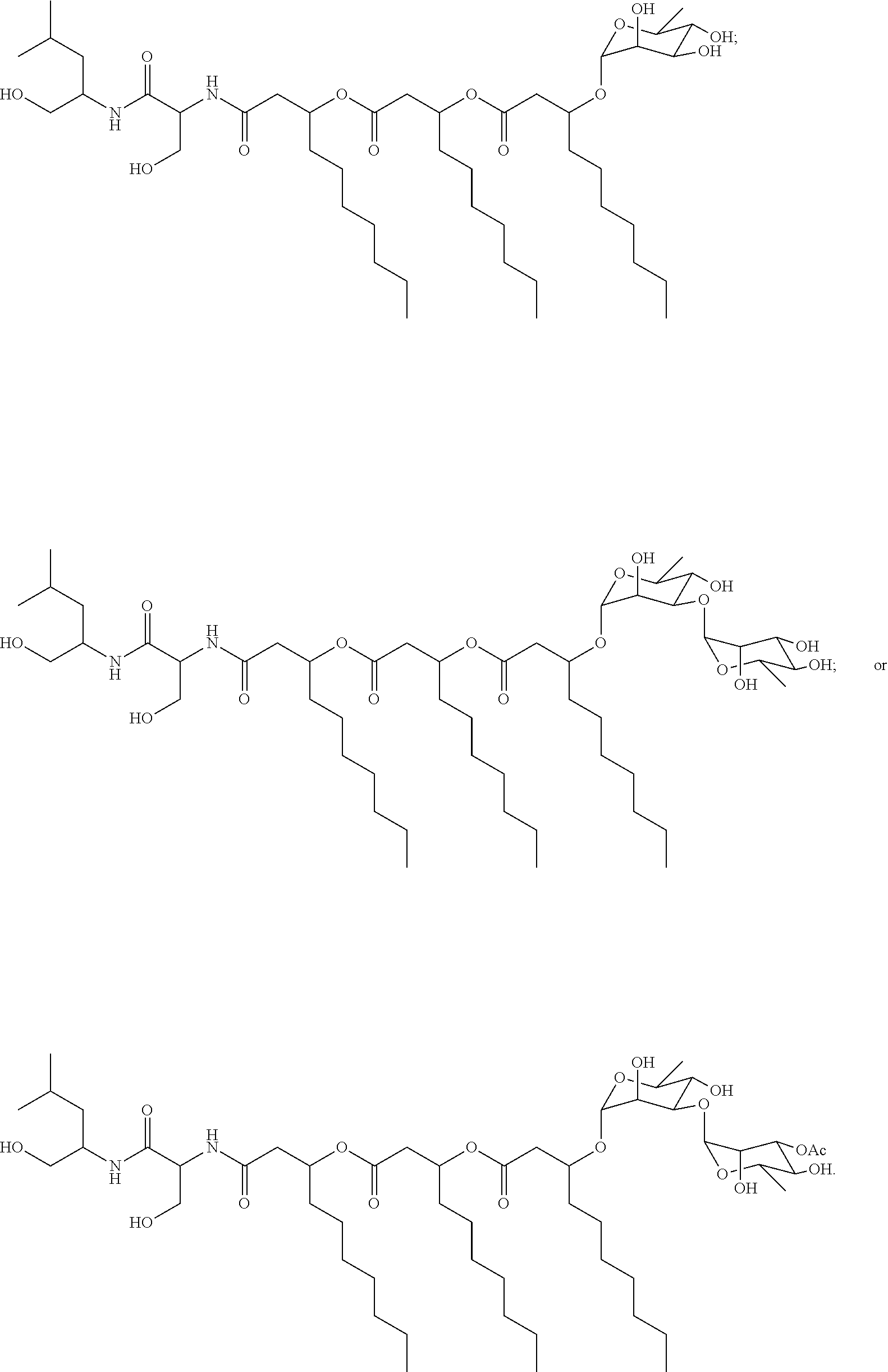
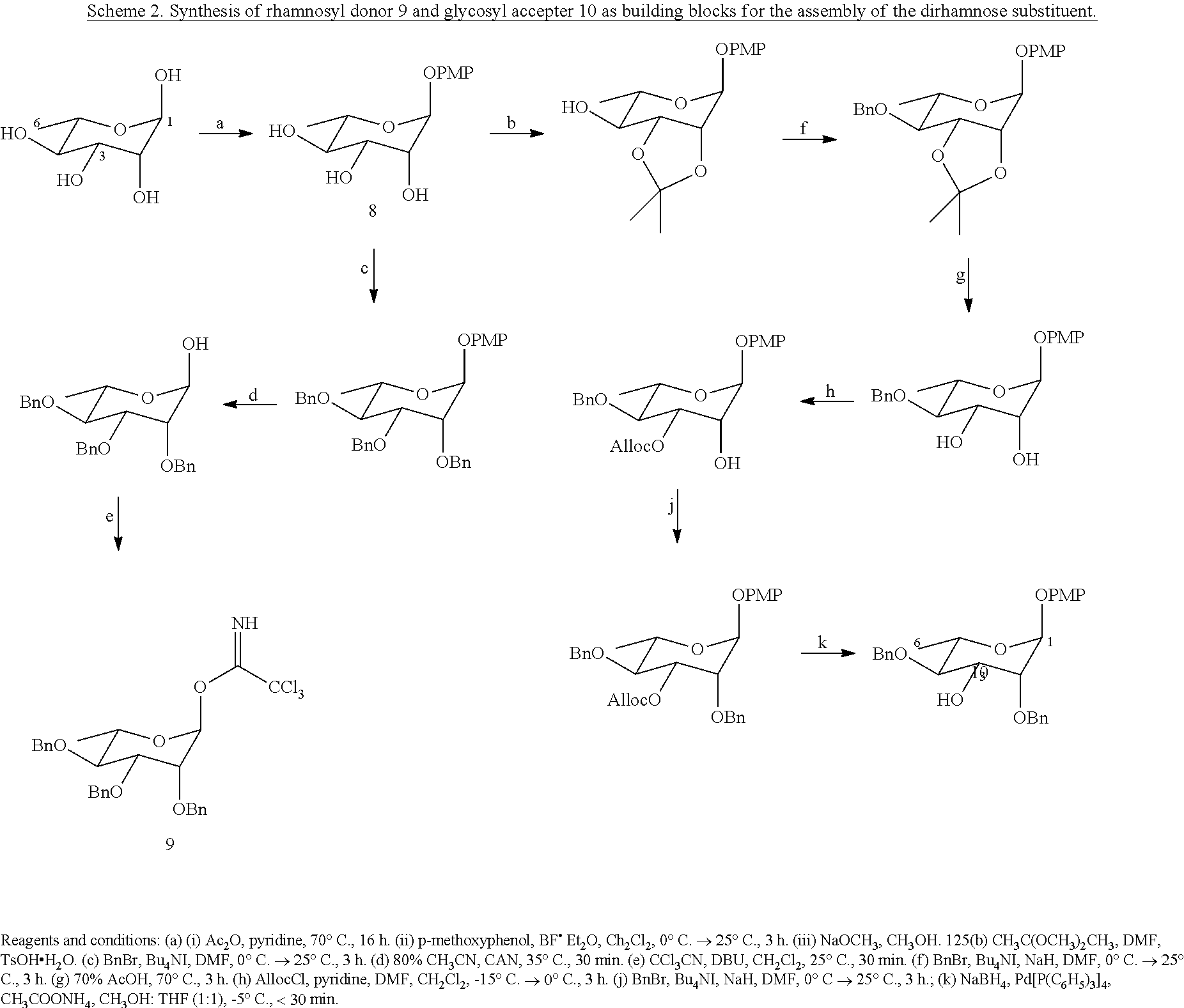
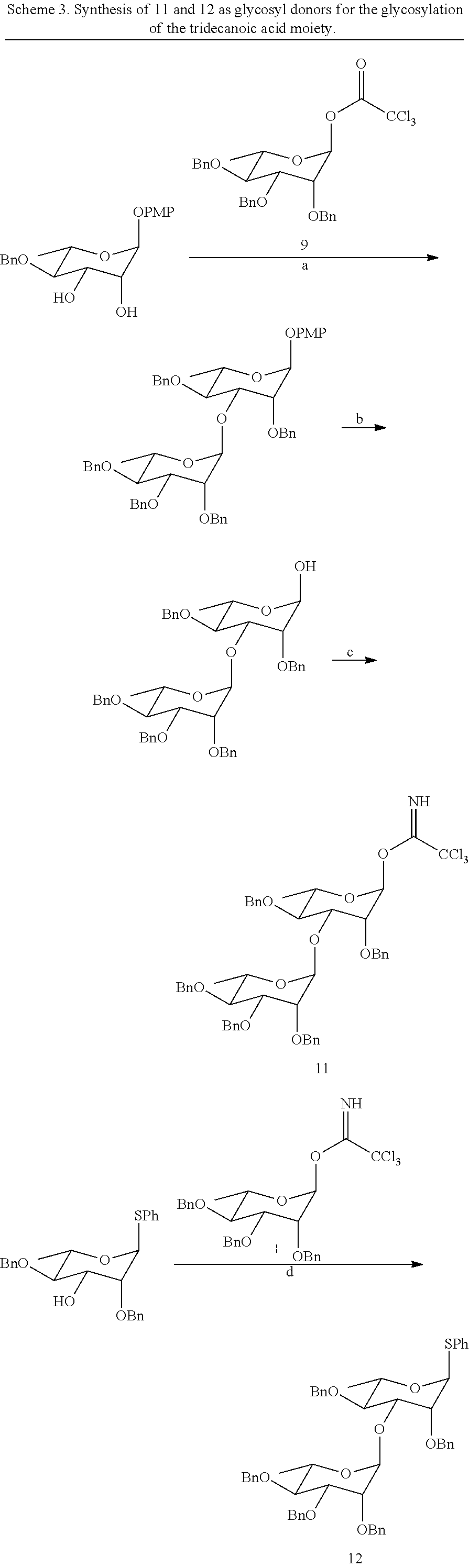

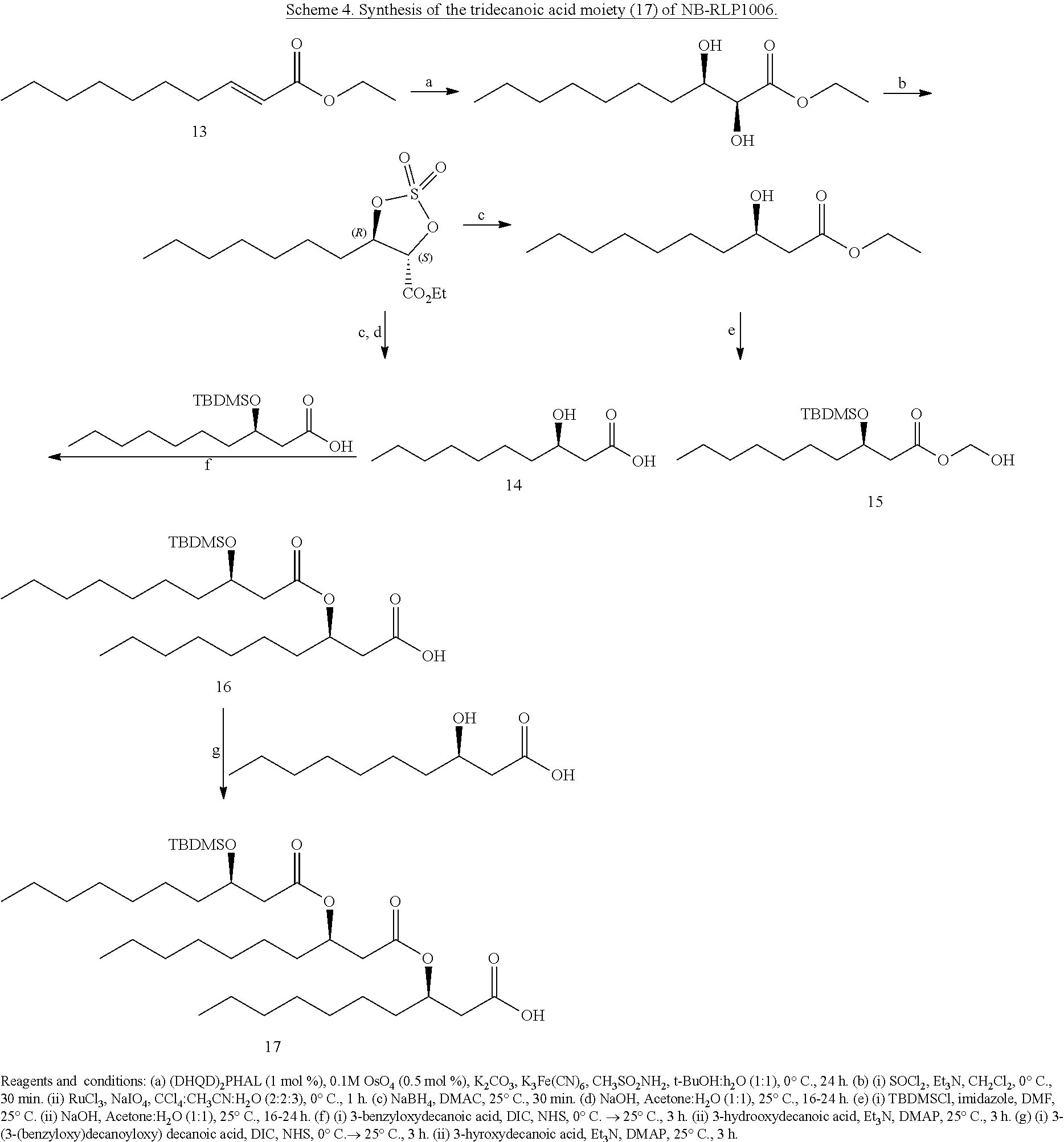


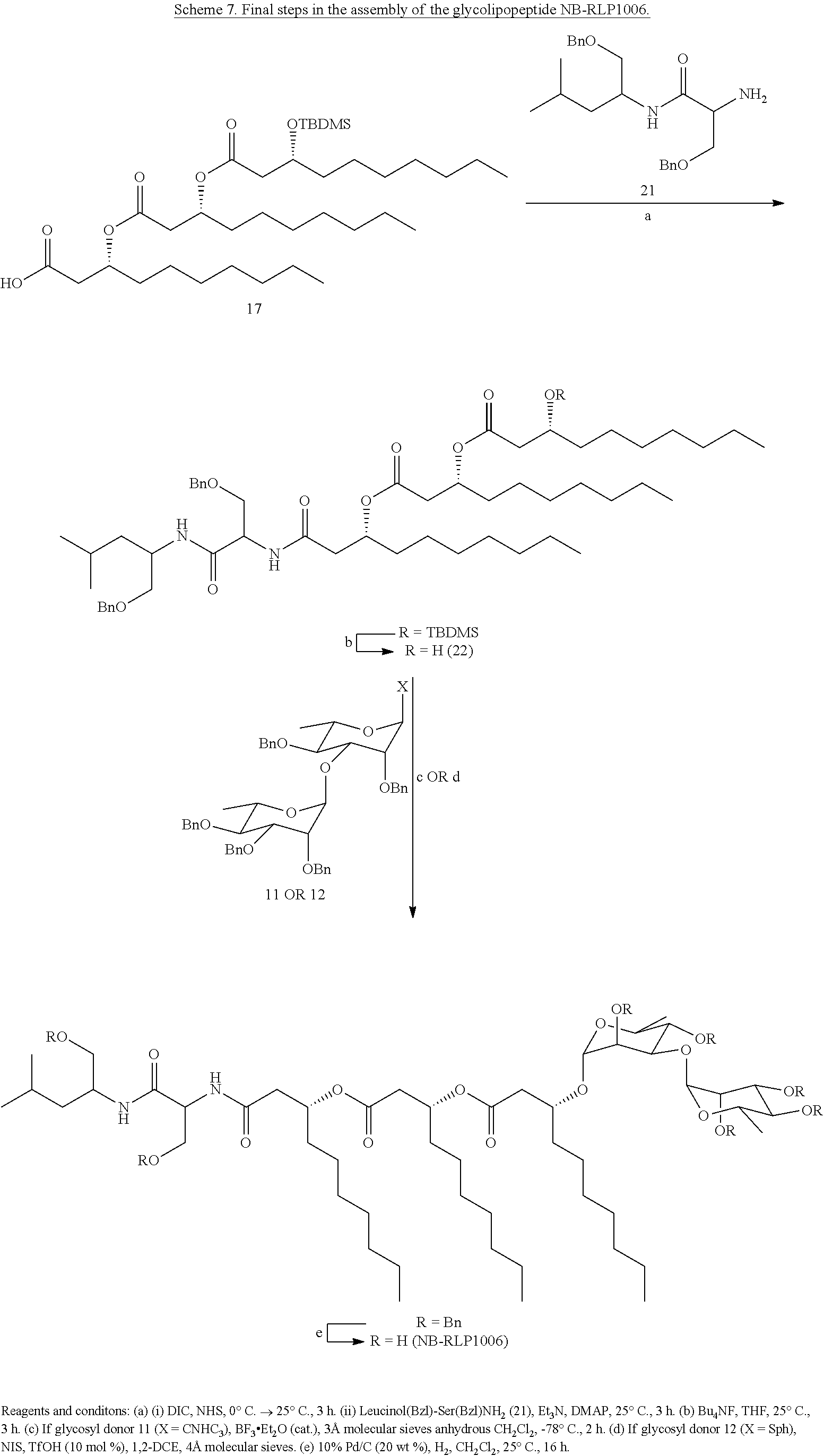
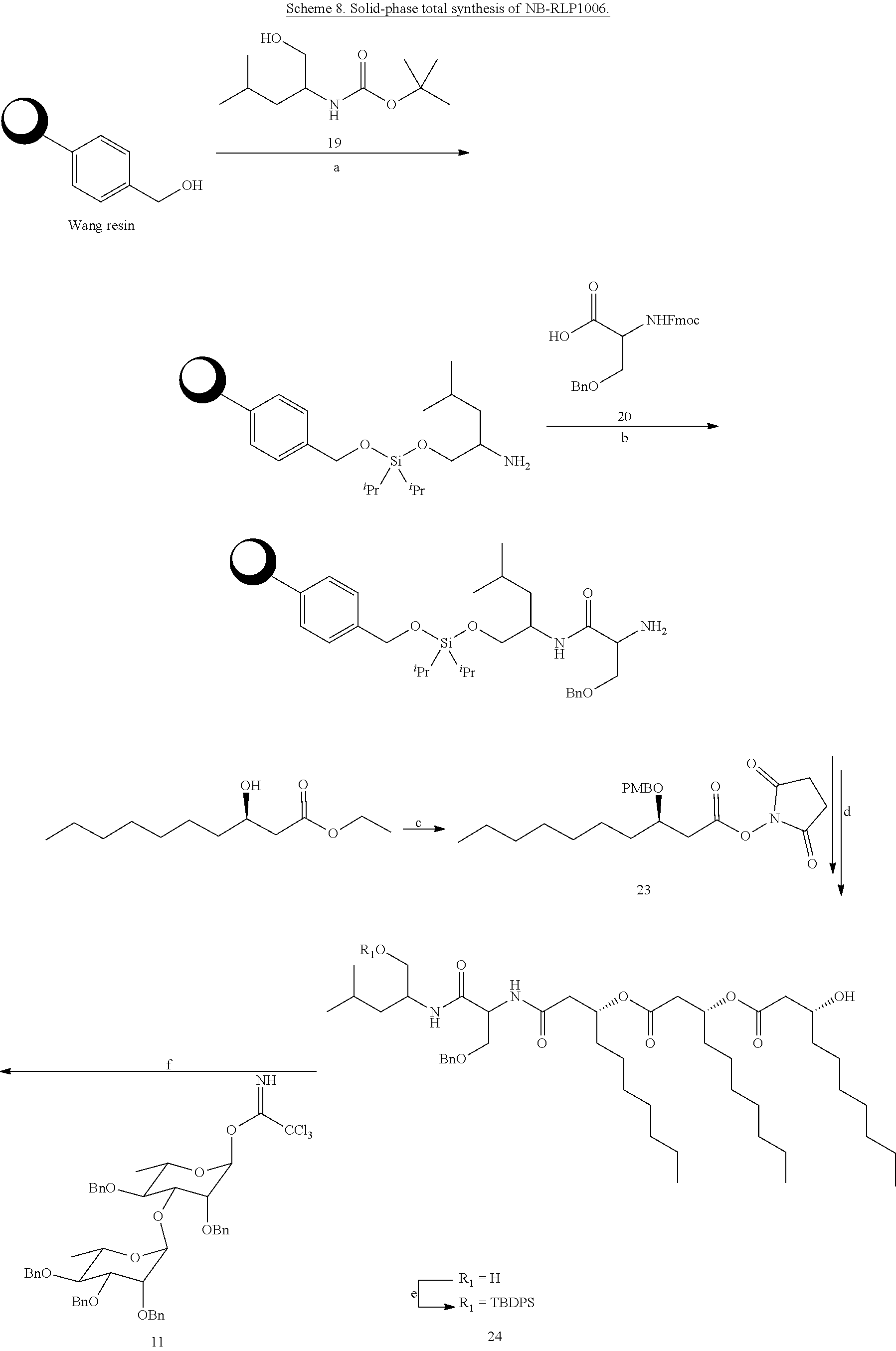

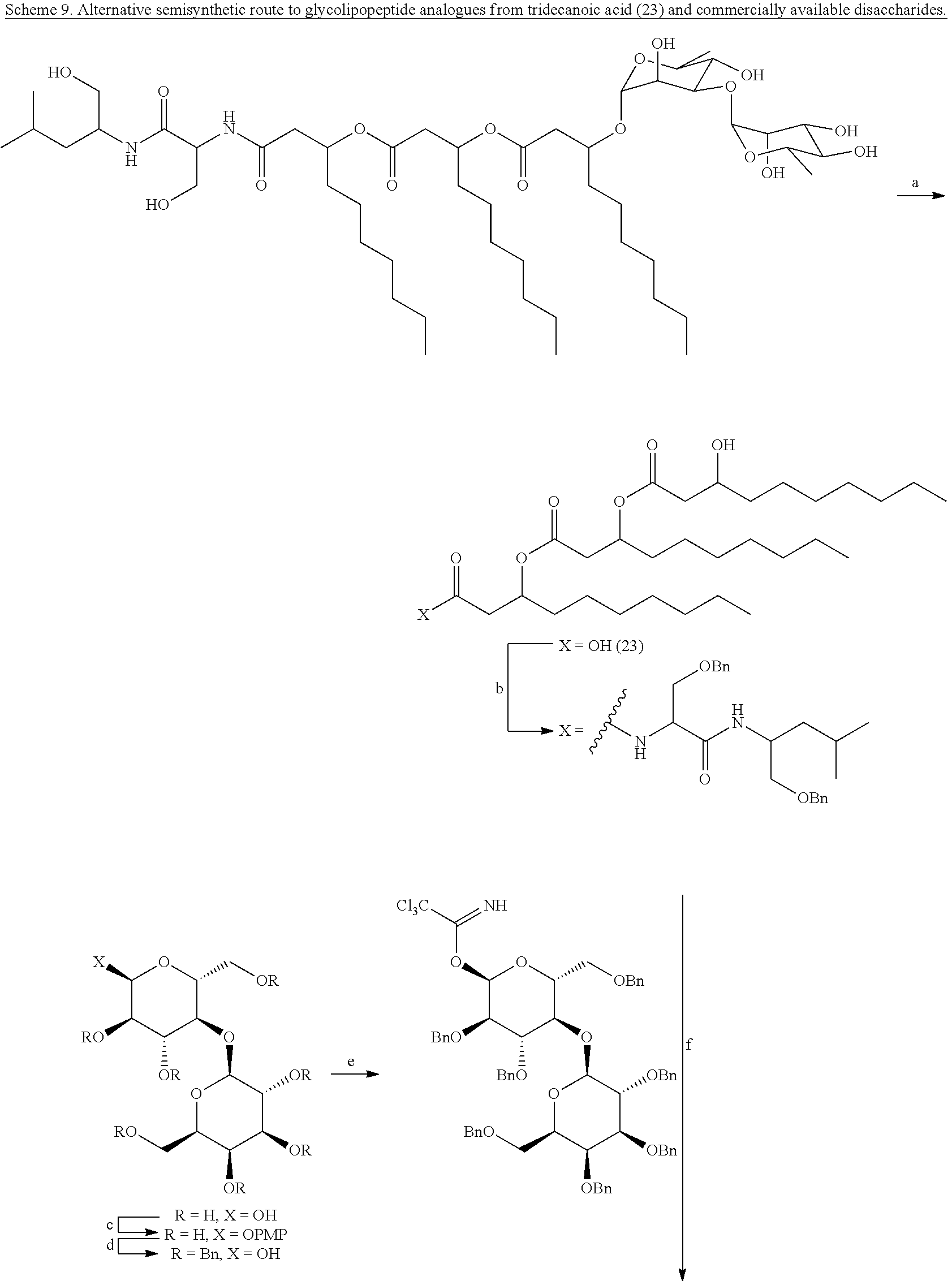
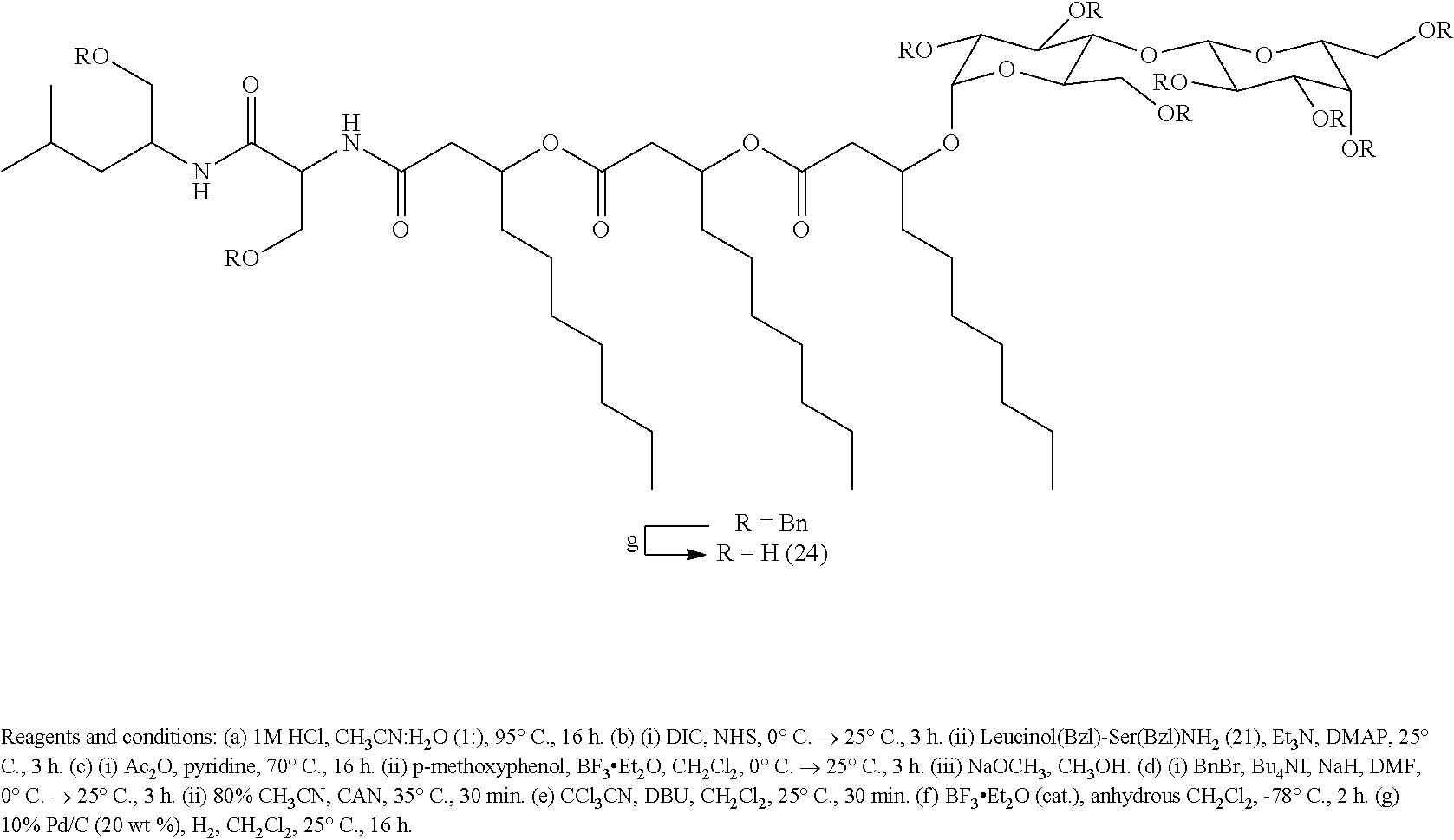
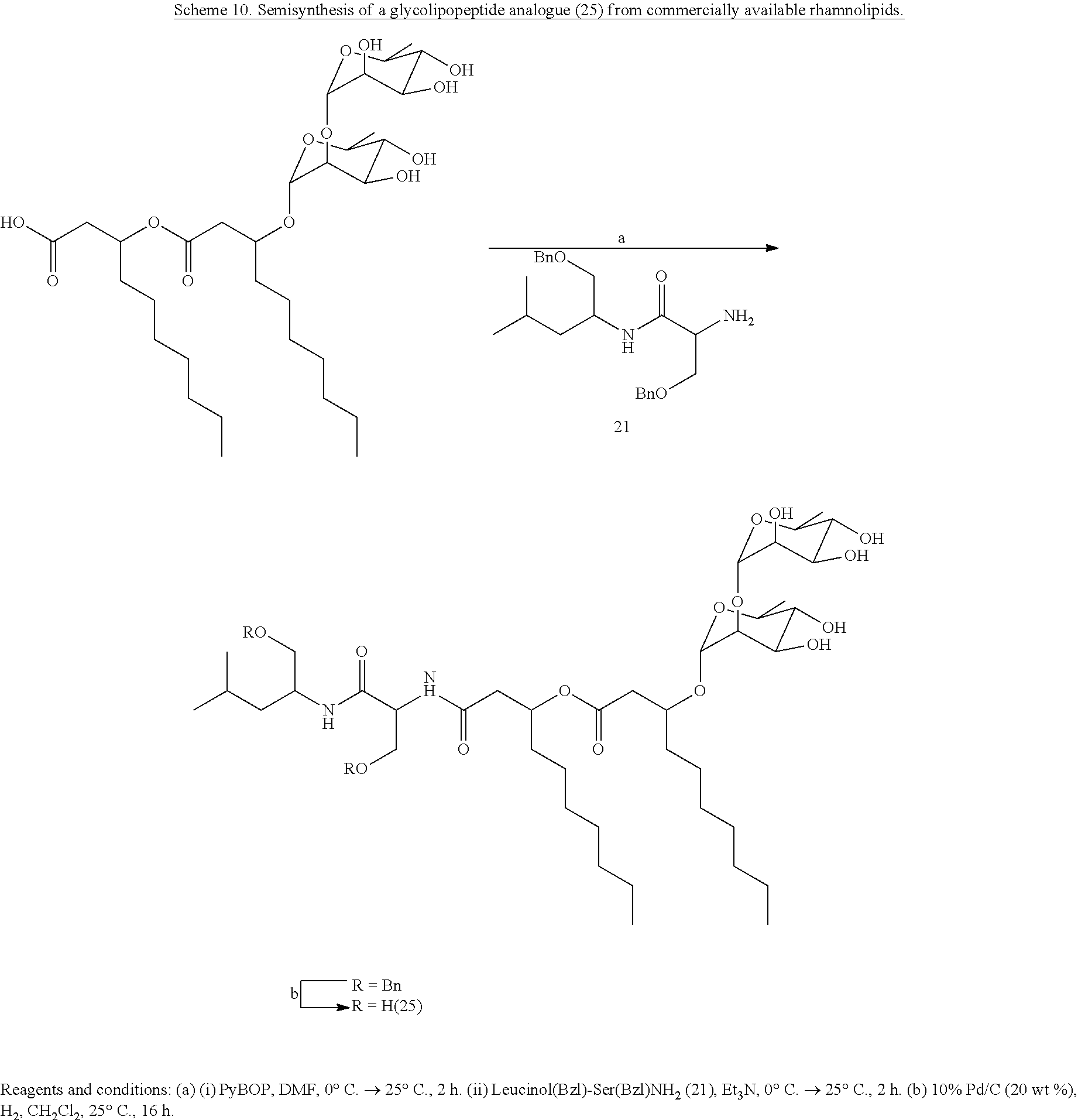



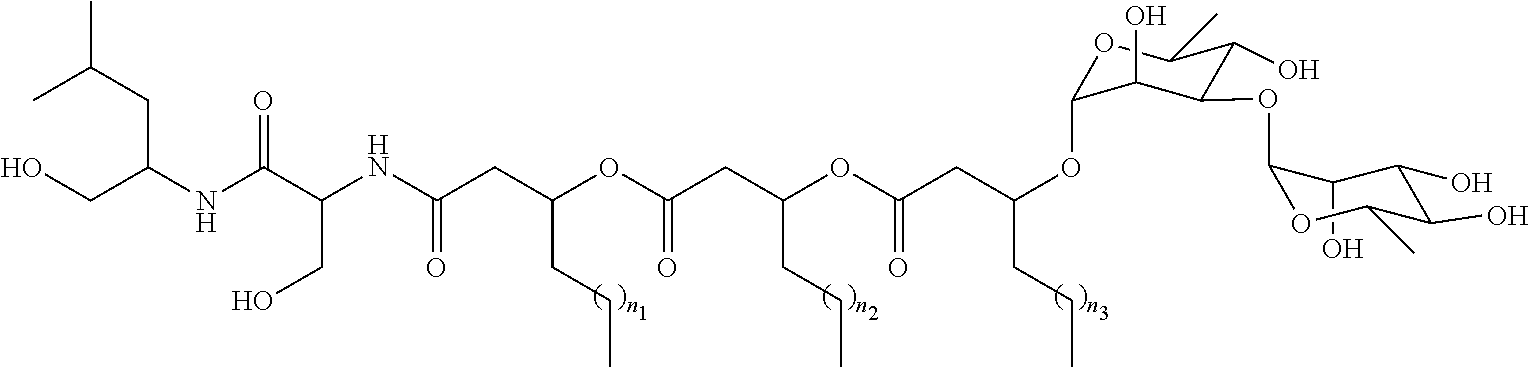





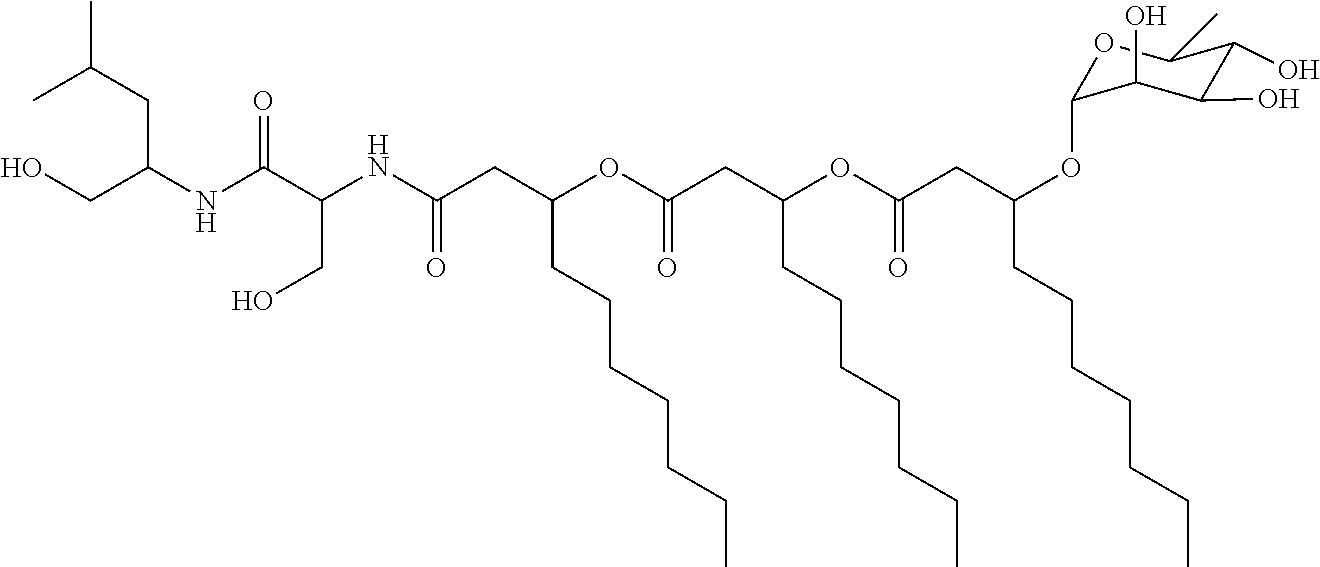
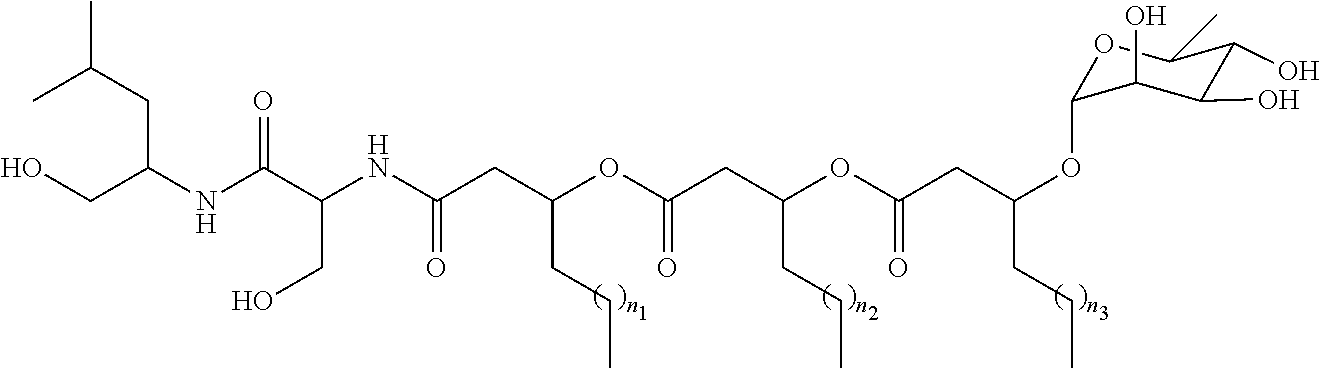

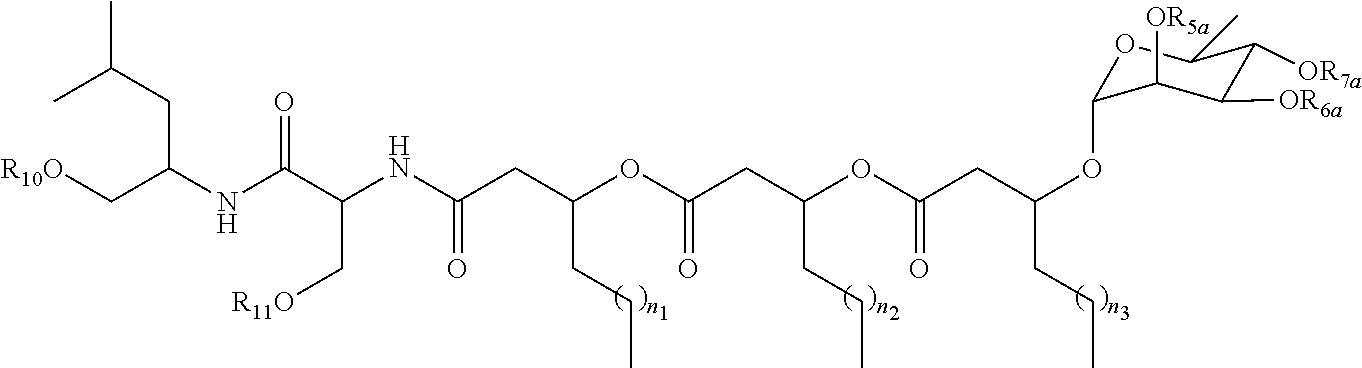

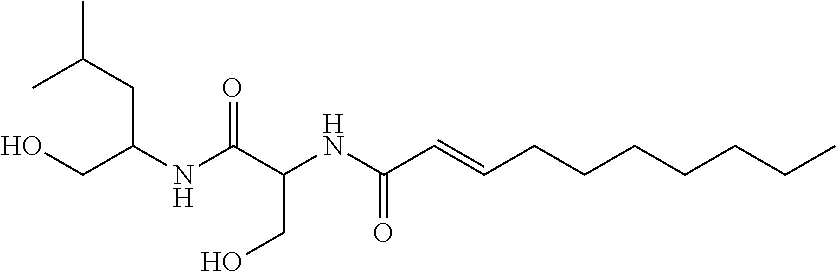

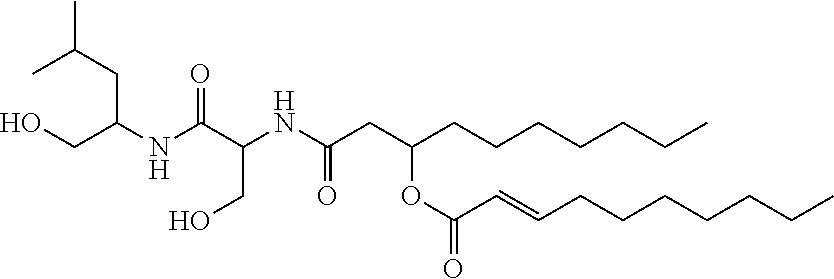
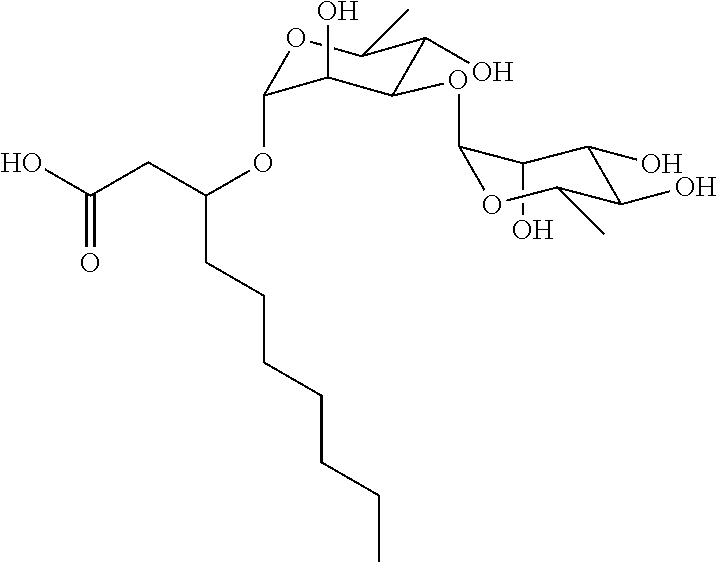
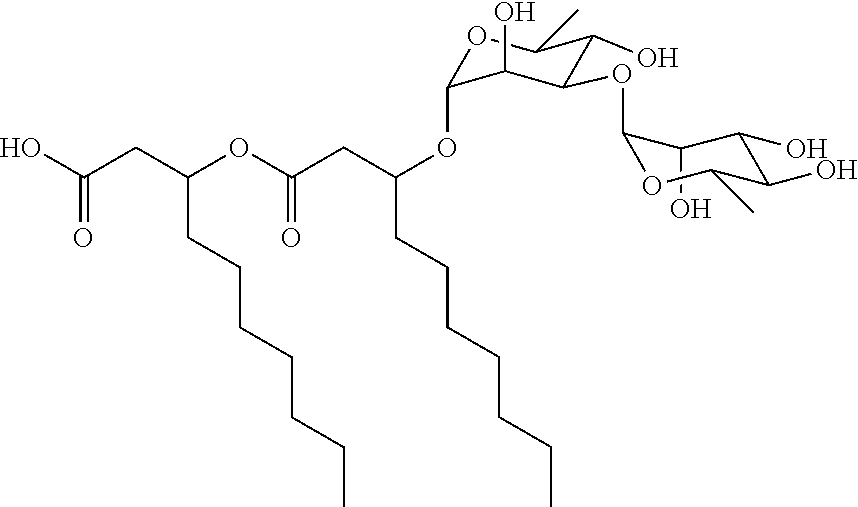
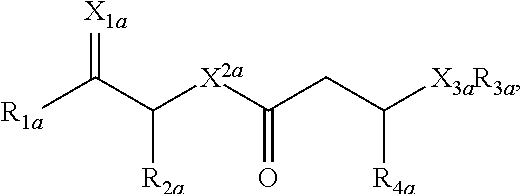

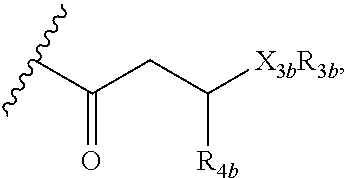

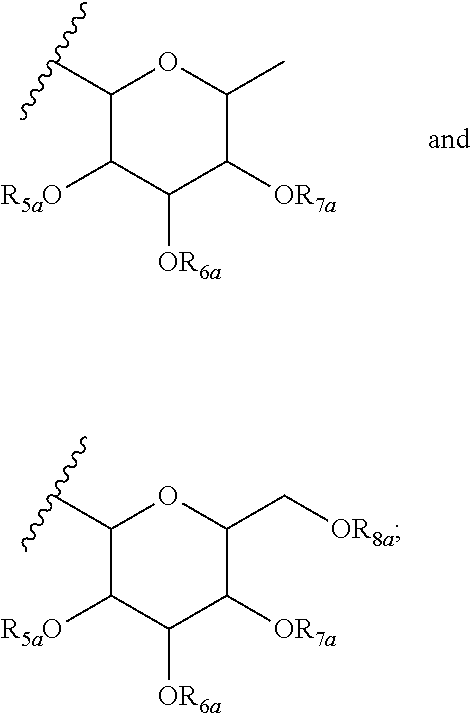
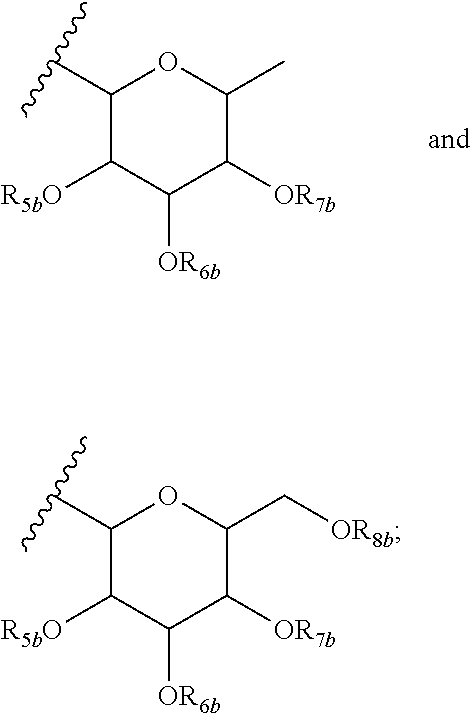







D00000
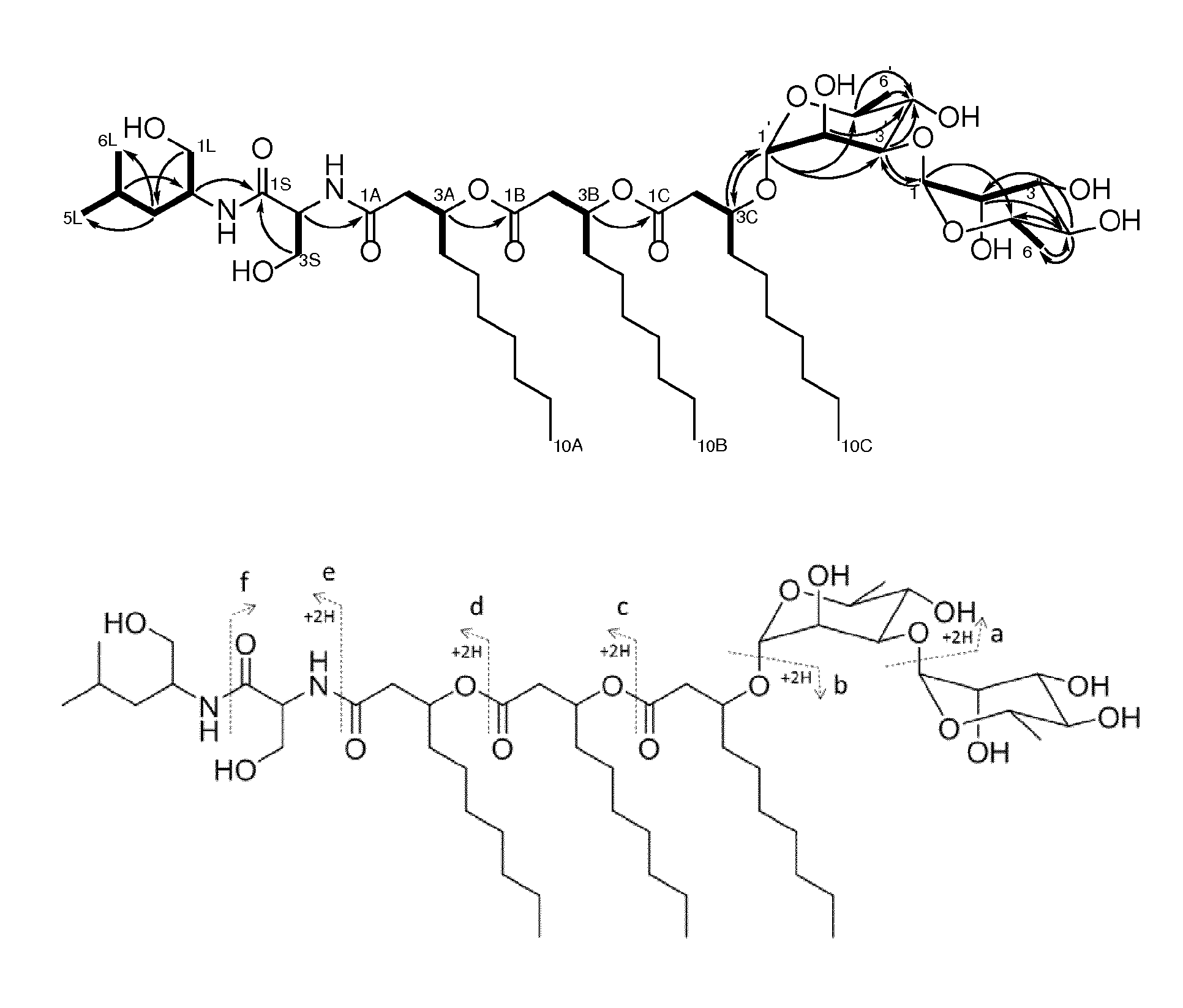
D00001

D00002
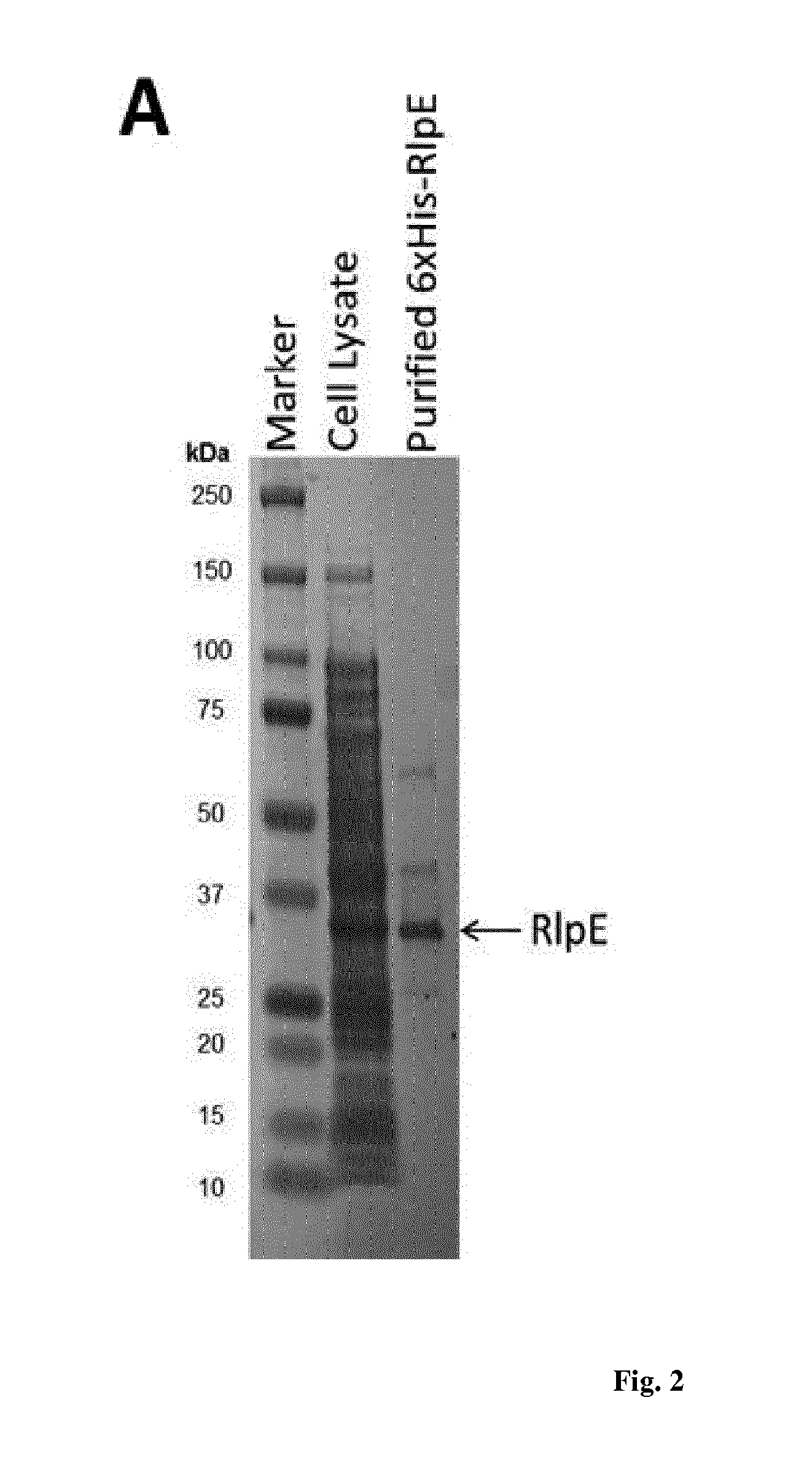
D00003
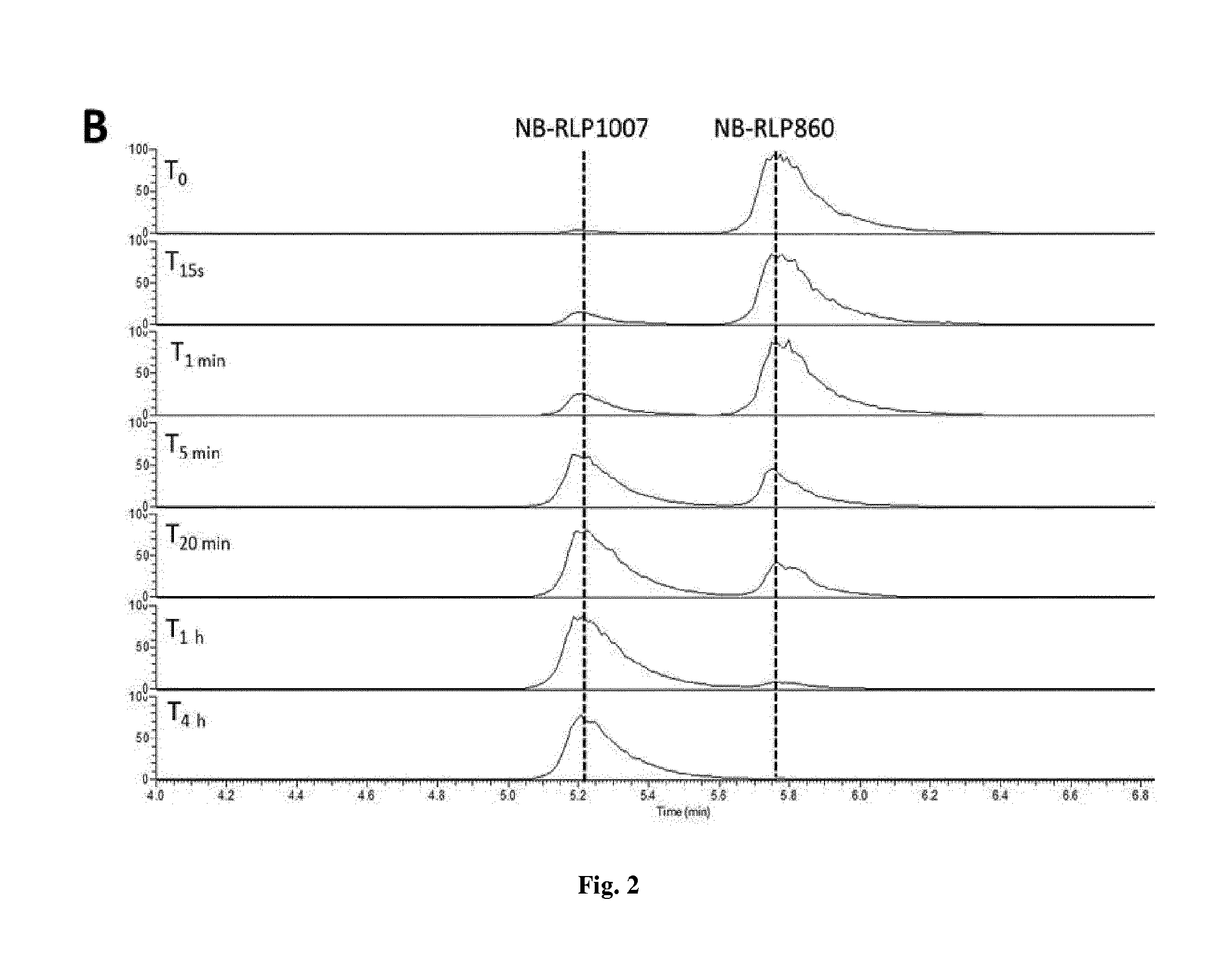
D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.