Methods Of Treating Cancer
Knutson; Sarah K. ; et al.
U.S. patent application number 15/979916 was filed with the patent office on 2019-04-04 for methods of treating cancer. The applicant listed for this patent is Epizyme, Inc.. Invention is credited to Heike Keilhack, Sarah K. Knutson, Natalie Warholic.
| Application Number | 20190100514 15/979916 |
| Document ID | / |
| Family ID | 49519104 |
| Filed Date | 2019-04-04 |










View All Diagrams
| United States Patent Application | 20190100514 |
| Kind Code | A1 |
| Knutson; Sarah K. ; et al. | April 4, 2019 |
METHODS OF TREATING CANCER
Abstract
The present invention relates to methods of treating cancer by administering the EZH2 inhibitor compounds and pharmaceutical compositions to subjects in need thereof. The present invention also relates to the use of such compounds for research or other non-therapeutic purposes.
| Inventors: | Knutson; Sarah K.; (Lincoln, MA) ; Warholic; Natalie; (Cambridge, MA) ; Keilhack; Heike; (Belmont, MA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 49519104 | ||||||||||
| Appl. No.: | 15/979916 | ||||||||||
| Filed: | May 15, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15598262 | May 17, 2017 | |||
| 15979916 | ||||
| 14054646 | Oct 15, 2013 | 9688665 | ||
| 15598262 | ||||
| 61786277 | Mar 14, 2013 | |||
| 61780703 | Mar 13, 2013 | |||
| 61758972 | Jan 31, 2013 | |||
| 61714140 | Oct 15, 2012 | |||
| 61714145 | Oct 15, 2012 | |||
| 61714045 | Oct 15, 2012 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07D 405/14 20130101; A61P 11/00 20180101; A61P 15/00 20180101; A61P 43/00 20180101; A61K 31/4545 20130101; A61P 7/00 20180101; A61P 35/00 20180101; A61P 25/00 20180101; C07D 405/12 20130101; A61P 13/08 20180101; A61P 35/02 20180101; C12Q 1/6886 20130101; A61P 13/12 20180101; A61K 31/4412 20130101; C07D 211/86 20130101; A61P 1/18 20180101; A61K 31/5377 20130101 |
| International Class: | C07D 405/14 20060101 C07D405/14; A61K 31/4545 20060101 A61K031/4545; A61K 31/4412 20060101 A61K031/4412; C12Q 1/6886 20060101 C12Q001/6886; C07D 405/12 20060101 C07D405/12; C07D 211/86 20060101 C07D211/86; A61K 31/5377 20060101 A61K031/5377 |
Claims
1. A method for treating or alleviating a symptom of a SWI/SNF-associated cancer in a subject comprising administering to a subject in need thereof a therapeutically effective amount of an EZH2 inhibitor, wherein the subject has a cancer selected from the group consisting of brain and central nervous system cancer, head and neck cancer, kidney cancer, ovarian cancer, pancreatic cancer, leukemia, lung cancer, lymphoma, myeloma, sarcoma, breast cancer, and prostate cancer.
2. The method of claim 1, wherein the SWI/SNF-associated cancer is characterized by reduced expression or loss of function of the SWI/SNF complex or one or more components of the SWI/SNF complex.
3. The method of claim 1, wherein the subject has a cancer selected from the group consisting of medulloblastoma, malignant rhabdoid tumor and atypical teratoid/rhabdoid tumor.
4. The method of claim 2, wherein the one or more components are selected from the group consisting of SNF5, ATRX and ARID1A.
5. The method of claim 2, wherein the loss of function is caused by a loss of function mutation resulting from a point mutation, a deletion and/or an insertion.
6. The method of claim 1, wherein the subject has a deletion of SNF5.
7. The method of claim 1, wherein the subject has a mutation of ATRX selected from the group consisting of a substitution of asparagine (N) for the wild type residue lysine (K) at amino acid position 688 of SEQ ID NO: 5 (K688N), and a substitution of isoleucine (I) for the wild type residue methionine (M) at amino acid position 366 of SEQ ID NO: 5 (M366I).
8. The method of claim 1, wherein said subject has a mutation of ARID1A selected from the group consisting of a nonsense mutation for the wild type residue cysteine (C) at amino acid position 884 of SEQ ID NO: 11 (C884*), a substitution of lysine (K) for the wild type residue glutamic acid (E) at amino acid position 966 (E966K), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1411 of SEQ ID NO: 11 (Q1411*), a frame shift mutation at the wild type residue phenylalanine (F) at amino acid position 1720 of SEQ ID NO: 11 (F1720fs), a frame shift mutation after the wild type residue glycine (G) at amino acid position 1847 of SEQ ID NO: 11 (G1847fs), a frame shift mutation at the wild type residue cysteine (C) at amino acid position 1874 of SEQ ID NO: 11 (C1874fs), a substitution of glutamic acid (E) for the wild type residue aspartic acid (D) at amino acid position 1957 (D1957E), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1430 of SEQ ID NO: 11 (Q1430*), a frame shift mutation at the wild type residue arginine (R) at amino acid position 1721 of SEQ ID NO: 11 (R1721fs), a substitution of glutamic acid (E) for the wild type residue glycine (G) at amino acid position 1255 (G1255E), a frame shift mutation at the wild type residue glycine (G) at amino acid position 284 of SEQ ID NO: 11 (G284fs), a nonsense mutation for the wild type residue arginine (R) at amino acid position 1722 of SEQ ID NO: 11 (R1722*), a frame shift mutation at the wild type residue methionine (M) at amino acid position 274 of SEQ ID NO: 11 (M274fs), a frame shift mutation at the wild type residue glycine (G) at amino acid position 1847 of SEQ ID NO: 11 (G1847fs), a frame shift mutation at the wild type residue P at amino acid position 559 of SEQ ID NO: 11 (P559fs), a nonsense mutation for the wild type residue arginine (R) at amino acid position 1276 of SEQ ID NO: 11 (R1276*), a frame shift mutation at the wild type residue glutamine (Q) at amino acid position 2176 of SEQ ID NO: 11 (Q2176fs), a frame shift mutation at the wild type residue histidine (H) at amino acid position 203 of SEQ ID NO: 11 (H203fs), a frame shift mutation at the wild type residue alanine (A) at amino acid position 591 of SEQ ID NO: 11 (A591fs), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1322 of SEQ ID NO: 11 (Q1322*), a nonsense mutation for the wild type residue serine (S) at amino acid position 2264 of SEQ ID NO: 11 (S2264*), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 586 of SEQ ID NO: 11 (Q586*), a frame shift mutation at the wild type residue glutamine (Q) at amino acid position 548 of SEQ ID NO: 11 (Q548fs), and a frame shift mutation at the wild type residue asparagine (N) at amino acid position 756 of SEQ ID NO: 11 (N756fs).
9-18. (canceled)
19. A method of inducing neuronal differentiation, cell cycle inhibition, gene expression or tumor suppression comprising contacting a cell with an EZH2 inhibitor.
20. The method of claim 19, wherein the EZH2 inhibitor is in an amount sufficient to increase expression of at least one gene selected from the group consisting of CD133, DOCK4, PTPRK, CKDN1A, CDKN2A, and BIN1.
21. A method of inhibiting hedgehog signaling or gene expression comprising contacting a cell with an EZH2 inhibitor.
22. The method of claim 21, wherein the EZH2 inhibitor is in an amount sufficient to reduce expression of GLI1 and/or PTCH1.
23. (canceled)
24. The method of claim 19, wherein the EZH2 inhibitor is in an amount sufficient to induce neuronal differentiation, cell cycle inhibition and/or tumor suppression.
25-26. (canceled)
27. The method of claim 21, wherein the EZH2 inhibitor is in an amount sufficient to inhibit hedgehog signaling.
28. (canceled)
29. The method of claim 19, wherein the cell comprises loss of function of SNF5, ARID1A, ATRX, and/or a component of the SWI/SNF complex.
30. The method of claim 29, wherein the loss of function is caused by a deletion of SNF5.
31. The method of claim 19, wherein the cell is a cancer cell.
32. The method of claim 31, wherein the cancer is selected from the group consisting of medulloblastoma, malignant rhabdoid tumor, and atypical teratoid rhabdoid tumor.
33. The method of claim 1, wherein the EZH2 inhibitor is: ##STR00051## or pharmaceutically acceptable salt thereof.
34. The method of claim 1, wherein the EZH2 inhibitor is: ##STR00052## or pharmaceutically acceptable salt thereof.
Description
RELATED APPLICATIONS
[0001] This application is a continuation application of U.S. application Ser. No. 15/598,262, filed May 17, 2017, which is a continuation application of U.S. application Ser. No. 14/054,646, filed Oct. 15, 2013 (now U.S. Pat. No. 9,688,665), which claims priority to, and the benefit of U.S. Provisional Application Nos. 61/714,045, filed Oct. 15, 2012, 61/758,972, filed Jan. 31, 2013, 61/714,140, filed Oct. 15, 2012, 61/714,145, filed Oct. 15, 2012, 61/780,703, filed Mar. 13, 2013, and 61/786,277, filed Mar. 14, 2013. The contents of each of these provisional applications are incorporated herein by reference in their entireties.
INCORPORATION-BY-REFERENCE OF SEQUENCE LISTING
[0002] The contents of the text file named "41478-513001US_ST25.txt", which was created on Jan. 10, 2014 and is 141 KB in size, are hereby incorporated by reference in their entireties.
FIELD OF INVENTION
[0003] The present invention relates generally to the field of cancer treatment, and in particular, the treatment of cancer associated with the SWI/SNF complex (i.e., SWI/SNF mediated cancer). More particularly, the present invention provides methods and compositions which treat, alleviate, prevent, diminish or otherwise ameliorate the symptoms of cancer associated with the SWI/SNF complex.
BACKGROUND OF THE INVENTION
[0004] Disease-associated chromatin-modifying enzymes (e.g., EZH2) play a role in diseases such as proliferative disorders, metabolic disorders, and blood disorders. Thus, there is a need for the development of small molecules that are capable of modulating the activity of EZH2.
SUMMARY OF THE INVENTION
[0005] The present invention provides a method for treating or alleviating a symptom of a SWI/SNF-associated cancer in a subject by administering to a subject in need thereof a therapeutically effective amount of an EZH2 inhibitor, where the subject has a cancer selected from the group consisting of brain and central nervous system cancer, head and neck cancer, kidney cancer, ovarian cancer, pancreatic cancer, leukemia, lung cancer, lymphoma, myeloma, sarcoma, breast cancer, and prostate cancer. For example, the SWI/SNF-associated cancer is characterized by reduced expression and/or loss of function of the SWI/SNF complex or one or more components of the SWI/SNF complex.
[0006] For example, the subject has a cancer selected from the group consisting of medulloblastoma, malignant rhabdoid tumor, and atypical teratoid/rhabdoid tumor.
[0007] For example, the one or more components are selected from the group consisting of SNF5, ATRX, and ARID1A.
[0008] For example, the loss of function is caused by a loss of function mutation resulting from a point mutation, a deletion, and/or an insertion.
[0009] For example, the subject has a deletion of SNF5.
[0010] For example, the subject has a mutation of ATRX selected from the group consisting of a substitution of asparagine (N) for the wild type residue lysine (K) at amino acid position 688 of SEQ ID NO: 5 (K688N), and a substitution of isoleucine (I) for the wild type residue methionine (M) at amino acid position 366 of SEQ ID NO: 5 (M366I).
[0011] For example, subject has a mutation of ARID1A selected from the group consisting of a nonsense mutation for the wild type residue cysteine (C) at amino acid position 884 of SEQ ID NO: 11 (C884*), a substitution of lysine (K) for the wild type residue glutamic acid (E) at amino acid position 966 (E966K), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1411 of SEQ ID NO: 11 (Q1411*), a frame shift mutation at the wild type residue phenylalanine (F) at amino acid position 1720 of SEQ ID NO: 11 (F1720fs), a frame shift mutation after the wild type residue glycine (G) at amino acid position 1847 of SEQ ID NO: 11 (G1847fs), a frame shift mutation at the wild type residue cysteine (C) at amino acid position 1874 of SEQ ID NO: 11 (C1874fs), a substitution of glutamic acid (E) for the wild type residue aspartic acid (D) at amino acid position 1957 (D1957E), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1430 of SEQ ID NO: 11 (Q1430*), a frame shift mutation at the wild type residue arginine (R) at amino acid position 1721 of SEQ ID NO: 11 (R1721fs), a substitution of glutamic acid (E) for the wild type residue glycine (G) at amino acid position 1255 (G1255E), a frame shift mutation at the wild type residue glycine (G) at amino acid position 284 of SEQ ID NO: 11 (G284fs), a nonsense mutation for the wild type residue arginine (R) at amino acid position 1722 of SEQ ID NO: 11 (R1722*), a frame shift mutation at the wild type residue methionine (M) at amino acid position 274 of SEQ ID NO: 11 (M274fs), a frame shift mutation at the wild type residue glycine (G) at amino acid position 1847 of SEQ ID NO: 11 (G1847fs), a frame shift mutation at the wild type residue P at amino acid position 559 of SEQ ID NO: 11 (P559fs), a nonsense mutation for the wild type residue arginine (R) at amino acid position 1276 of SEQ ID NO: 11 (R1276*), a frame shift mutation at the wild type residue glutamine (Q) at amino acid position 2176 of SEQ ID NO: 11 (Q2176fs), a frame shift mutation at the wild type residue histidine (H) at amino acid position 203 of SEQ ID NO: 11 (H203fs), a frame shift mutation at the wild type residue alanine (A) at amino acid position 591 of SEQ ID NO: 11 (A591fs), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1322 of SEQ ID NO: 11 (Q1322*), a nonsense mutation for the wild type residue serine (S) at amino acid position 2264 of SEQ ID NO: 11 (S2264*), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 586 of SEQ ID NO: 11 (Q586*), a frame shift mutation at the wild type residue glutamine (Q) at amino acid position 548 of SEQ ID NO: 11 (Q548fs), and a frame shift mutation at the wild type residue asparagine (N) at amino acid position 756 of SEQ ID NO: 11 (N756fs).
[0012] The present invention also provides a method of treating or alleviating a symptom of a SWI/SNF-associated cancer in a subject in need thereof by (a) determining the expression level of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes and tumor suppressor genes in a sample obtained from the subject; (b) selecting the subject having a decreased expression level of at least one gene in step a; and (c) administering to the subject selected in step b an effective amount of an EZH2 inhibitor, thereby treating or alleviating a symptom of cancer in the subject.
[0013] The present invention further provides a method of treating or alleviating a symptom of a SWI/SNF-associated cancer in a subject in need thereof by (a) determining the expression level of at least one gene selected from the group consisting of hedgehog pathway genes, myc pathway genes and histone methyltransferase genesin a sample obtained from the subject; (b) selecting the subject having an increased expression level of at least one gene in step a; and (c) administering to the subject selected in step b an effective amount of an EZH2 inhibitor, thereby treating or alleviating a symptom of cancer in the subject.
[0014] For example, the cancer can be medulloblastoma, malignant rhabdoid tumor or atypical teratoid rhabdoid tumor.
[0015] For example, the neuronal differentiation gene is CD133, DOCK4, or PTPRK.
[0016] For example, the cell cycle inhibition gene is CKDN1A or CDKN2A.
[0017] For example, the tumor suppressor gene is BIN1.
[0018] For example, the hedgehog pathway gene is GLI1 or PTCH1.
[0019] For example, the myc pathway gene is MYC.
[0020] For example, the histone methyltransferase gene is EZH2.
[0021] The present invention also provides a method of inducing neuronal differentiation, cell cycle inhibition or tumor suppression by contacting a cell with an EZH2 inhibitor. The EZH2 inhibitor may be in an amount sufficient to increase expression of at least one gene selected from the group consisting of CD133, DOCK4, PTPRK, CKDN1A, CDKN2A and BIN1.
[0022] The present invention also provides a method of inhibiting hedgehog signaling by contacting a cell with an EZH2 inhibitor. The EZH2 inhibitor can be in an amount sufficient to reduce expression of GLI1 and/or PTCH1.
[0023] The present invention also provides a method of inducing gene expression by contacting a cell with an EZH2 inhibitor. The EZH2 inhibitor can be in an amount sufficient to induce neuronal differentiation, cell cycle inhibition and/or tumor suppression. For example, the gene can be CD133, DOCK4, PTPRK, CKDN1A, CKDN2A or BIN1.
[0024] The present invention also provides a method of inhibiting gene expression by contacting a cell with an EZH2 inhibitor. The EZH2 inhibitor is in an amount sufficient to inhibit hedgehog signaling. For example, the gene can be GLI1 or PTCH1.
[0025] For example, the cell may have loss of function of SNF5, ARID1A, ATRX, and/or a component of the SWI/SNF complex.
[0026] For example, the loss of function is caused by a deletion of SNF5.
[0027] For example, the cell is a cancer cell. The cancer can be medulloblastoma, malignant rhabdoid tumor or atypical teratoid rhabdoid tumor.
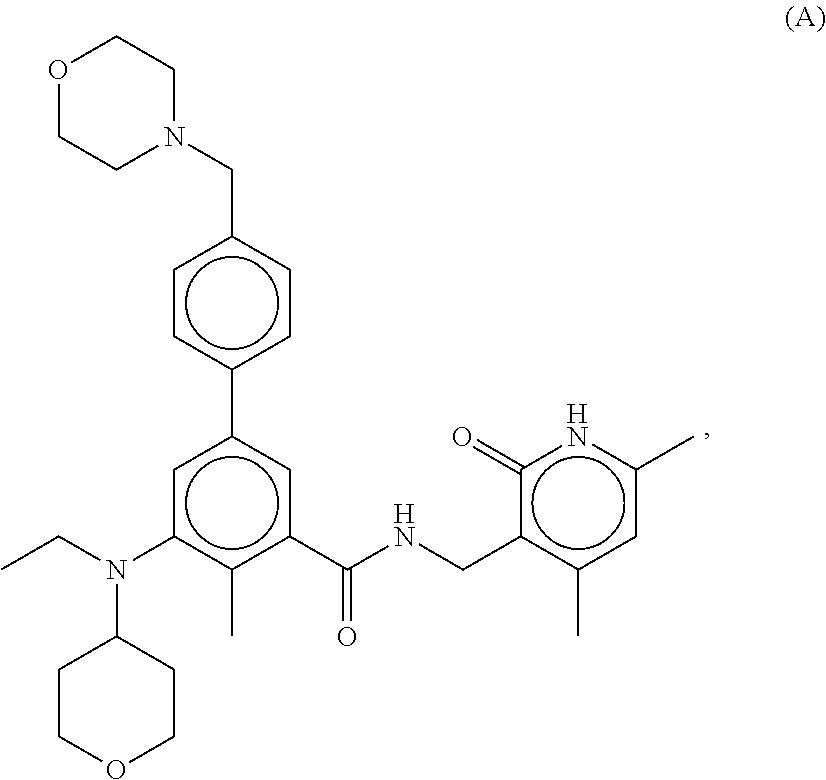
[0028] For example, the EZH2 inhibitor is Compound A having the following formula:
##STR00001##
stereoisomers thereof, or pharmaceutically acceptable salts or solvates thereof.
[0029] For example, the EZH2 inhibitor is Compound B having the following formula:
##STR00002##
stereoisomers thereof, or pharmaceutically acceptable salts or solvates thereof.
[0030] For example, the EZH2 inhibitor is Compound C having the following formula:
##STR00003##
stereoisomers thereof, or pharmaceutically acceptable salts or solvates thereof.
[0031] For example, the EZH2 inhibitor is Compound D having the following formula:
##STR00004##
stereoisomers thereof, or pharmaceutically acceptable salts or solvates thereof.
[0032] For example, the EZH2 inhibitor is Compound E having the following formula:
##STR00005##
stereoisomers thereof, or pharmaceutically acceptable salts or solvates thereof.
[0033] Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. In the specification, the singular forms also include the plural unless the context clearly dictates otherwise. Although methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, suitable methods and materials are described below. All publications, patent applications, patents and other references mentioned herein are incorporated by reference. The references cited herein are not admitted to be prior art to the claimed invention. In the case of conflict, the present specification, including definitions, will control. In addition, the materials, methods and examples are illustrative only and are not intended to be limiting.
[0034] Other features and advantages of the invention will be apparent from the following detailed description and claims.
BRIEF DESCRIPTIONS OF FIGURES
[0035] FIGS. 1A and 1B are a series of Western blot analyses of cell lines with wild type (RD and SJCRH30) and mutant SNF5.
[0036] FIGS. 2A-2E are a series of graphs establishing that SNF5 mutant cell lines A204 (FIG. 2C), G401 (FIG. 2D) and G402 (FIG. 2E) selectively respond to EZH2 compound (Compound E) compared to wild type cell lines RD (FIG. 2A) and SJCRH30 (FIG. 2B).
[0037] FIGS. 3A-3D are a series of bar graphs showing that G401 SNF mutant cell line is responding to Compound E after 7 days in soft agar compared to wild type cells RD. FIG. 3A shows cell line RD (5,000 cells/well). FIG. 3B shows G401 cells (5,000 cells/well). FIG. 3C shows G401 cells in 2D growth. FIG. 3D shows G401 cells (10,000 cells/well).
[0038] FIGS. 4A-4D are four graphs showing that G401 SNF5 mutant cell line is sensitive to Compound A in vitro. Wild type cell lines SJCRH30 (FIG. 4A) and RD (FIG. 4C) and SNF5 mutant cell lines G401 (FIG. 4B) and A204 (FIG. 4D) were pretreated for 7 days with indicated concentrations of Compound A and replated on day 0. Cell viability was determined by CellTiter-Glo.RTM. Luminescent Cell Viability Assay.
[0039] FIGS. 5A-5E are a series of graphs showing durable regressions in G401 xenografts (malignant rhabdoid tumor model) with Compound A treatment. FIG. 5A shows tumor regressions induced by Compound A at the indicated doses. FIG. 5B shows tumor regressions induced by twice daily administration of Compound A at the indicated doses. Data represent the mean values .+-.SEM (n=8). Compound administration was stopped on day 28. FIG. 5C shows the EZH2 target inhibition in G401 xenograft tumor tissue collected from a parallel cohort of mice on day 21. Each point shows the ratio of H3K27Me3 to total H3. Horizontal lines represent group mean values. BLLQ=below lower limit of quantification. FIGS. 5D and 5E show immunohistochemical staining of tumor histone methylation of tumor samples from the vehicle treated (FIG. 5D) and Compound A treated (FIG. 5E) (at 125 mg/kg) mice.
[0040] FIG. 6 is a graph showing the locations of ATRX mutations identified in SCLC cell lines.
[0041] FIG. 7A is a graph showing that LNCAP prostate cancer cells display dose-dependent cell growth inhibition with Compound E treatment in vitro.
[0042] FIG. 7B is a graph showing IC50 value of Compound E at day 11 and day 14 for WSU-DLCL2 and LNCAP cells.
[0043] FIGS. 8A-8C are three graphs establishing that ATRX mutant SCLC lines NCI-H446 (FIG. 8A), SW1271 (FIG. 8B) and NCI-H841 (FIG. 8C) are responding to Compound E.
[0044] FIGS. 9A-9C are three microscopy images showing that SCLC line NCI-H841 changes morphology after treatment with vehicle (FIG. 9A) or Compound E at concentration of 4.1E-02 uM (FIG. 9B) or 3.3 uM (FIG. 9C).
[0045] FIGS. 10A-10F are a series of graphs showing effects of Compound A on cellular global histone methylation and cell viability. FIG. 10A shows the chemical structure of Compound A. FIG. 10B illustrates the concentration-dependent inhibition of cellular H3K27Me3 levels in G401 and RD cells. FIGS. 10C through 10F illustrate the selective inhibition of proliferation of SMARCB1-deleted G401 cells by Compound A in vitro (measured by ATP content). G401 (FIGS. 10C and 10D) and RD cells (FIGS. 10E and 10F) were re-plated at the original seeding densities on day 7. Each point represents the mean for each concentration (n=3).
[0046] FIGS. 11A and 11B are a series of graphs showing biochemical mechanism of action studies. The IC.sub.50 value of Compound A increases with increasing SAM concentration (FIG. 11A) and is minimally affected by increasing oligonucleosome concentration (FIG. 11B), indicating SAM-competitive and nucleosome-noncompetitive mechanism of action.
[0047] FIGS. 12A and 12B are a series of immunoblots demonstrating verification of SMARCB1 and EZH2 expression in cell lines and specificity of Compound A for inhibition of cellular histone methylation. FIG. 12A shows cell lysates analyzed by immunoblot with antibodies specific to SMARCB1, EZH2 and Actin (loading control). FIG. 12B illustrates selective inhibition of cellular H3K27 methylation in G401 and RD cells. Cells were incubated with Compound A for 4 days, and acid-extracted histones were analyzed by immunoblot.
[0048] FIGS. 13A and 13B are a series of bar graphs demonstrating that Compound A induces G.sub.1 arrest and apoptosis in SMARCB1-deleted MRT cells. Cell cycle analysis (by flow cytometry) and determination of apoptosis (by TUNEL assay) in RD (FIG. 13A) or G401 cells (FIG. 13B) during incubation with either vehicle or 1 .mu.M Compound A for up to 14 days. G.sub.1 arrest was observed as of day 7 and apoptosis was induced as of day 11. Data are represented as mean values.+-.SEM (n=2). The DMSO control values shown are the average.+-.SEM from each time point. Cells were split and re-plated on days 4, 7 and 11 at the original seeding density.
[0049] FIGS. 14A-14L are a series of graphs showing that Compound A induces changes in expression of SMARCB1 regulated genes and cell morphology. FIG. 14A shows the basal expression of SMARCB1 regulated genes in G401 SMARCB1-deleted cells, relative to RD control cells (measured by qPCR, n=2). FIGS. 14B-14K show the expression of GLI1 (FIG. 14B), PTCh1 (FIG. 14C), DOCK4 (FIG. 14D), CD133 (FIG. 14E), PTPRK (FIG. 14F), BIN1 (FIG. 14G), CDKN1A (FIG. 14H), CDKN2A (FIG. 14I), EZH2 (FIG. 14J), and MYC (FIG. 14K) genes in G401 and RD cells. The cells were incubated with either DMSO or 1 .mu.M Compound A for 2, 4 and 7 days. Gene expression was determined by qPCR (n=2) and is expressed relative to the DMSO control of each time point. FIG. 14L illustrates the morphology of G402 cells that were incubated with either DMSO (left panel) or 1 .mu.M Compound A (right panel) for 14 days. Cells were split and re-plated to the original seeding density on day 7.
[0050] FIGS. 15A-15D are a series of graphs demonstrating body weights, tumor regressions and plasma levels in G401 xenograft bearing mice treated with Compound A. FIG. 15A shows body weights that were determined twice a week for animals treated with Compound A on a BID schedule for 28 days. Data are presented as mean values.+-.SEM (n=16 until day 21, n=8 from day 22 to 60). FIG. 15B shows tumor regressions induced by twice daily (BID) administration of Compound A for 21 days at the indicated doses (mean values.+-.SEM, n=16). *p<0.05, **p<0.01, repeated measures ANOVA, Dunnett's post-test vs. vehicle. FIG. 15C shows the tumor weights of 8 mice euthanized on day 21. ****p<0.0001, Fisher's exact test. FIG. 15D shows plasma levels of Compound A. Plasma was collected 5 min before and 3 h after dosing of Compound A on day 21, and compound levels were measured by LC-MS/MS. Animals were euthanized, and tumors were collected 3 h after dosing on day 21. Tumor homogenates were generated and subjected to LC-MS/MS analysis to determine Compound A concentrations. Note that tumor compound levels could not be determined from all animals especially in the higher dose groups because the xenografts were too small on day 21. Dots represent values for the individual animals; horizontal lines represent group mean values.
[0051] FIGS. 16A-16F are a series of graphs showing that Compound A eradicates SMARCB1-deleted MRT xenografts in SCID mice. FIG. 16A illustrates tumor regressions induced by twice daily (BID) administration of Compound A for 28 days at the indicated doses. Compound administration was stopped on day 28 and tumors were allowed to re-grow until they reached 2000 mm.sup.3 (data shown as mean values.+-.SEM, n=8). FIG. 16B shows the EZH2 target inhibition in G401 xenograft tumor tissue collected from mice euthanized on day 21. Each point shows the ratio of H3K27Me3 to total H3, measured by ELISA. Horizontal lines represent group mean values; grey symbols are values outside of the ELISA standard curve. FIGS. 16C-16F summarize the change in gene expression in G401 xenograft tumor tissue collected from mice treated with Compound A for 21 days. The graphs correspond to genes CD133 (FIG. 16C), PTPRK (FIG. 16D), DOCK4 (FIG. 16E), and GLI1 (FIG. 16F), respectively. Data are presented as fold change compared to vehicle.+-.SEM (n=6, n=4 for 500 mg/kg group). *p<0.05, **p<0.01, ****p<0.0001, vs. vehicle, Fisher's exact test.
DETAILED DESCRIPTION OF THE INVENTION
[0052] The present invention is based in part upon the discovery that EZH2 inhibitors can effectively treat SWI/SNF-associated cancers that are characterized by altered expressions and/or loss of function of certain biomarkers or genes. Specifically, tumors or tumor cells having altered expressions and/or loss of function of selected biomarkers or genes are sensitive to the EZH2 inhibitors of the present invention. Accordingly, the present invention provides methods of treating or alleviating a symptom of cancers in a subject by administering a therapeutically effective amount of an EZH2 inhibitor to the subject, particular treating cancers associated with altered expression and/or loss of function of certain biomarkers or genes. For example, the biomarker is one component of the SWI/SNF complex. For example, the gene is selected from the group consisting of neuronal differentiation genes, cell cycle gene inhibition genes, tumor suppressor genes, hedgehog pathway genes, myc pathway genes and histone methyltransferase genes.
[0053] The SWI/SNF complex in human includes at least evolutionarily conserved core subunits and variant subunits. Evolutionarily conserved core subunits include SNF5 (also called SMARCB1, INI1 or BAF47), SMARCA4 (also known as BRM/SWI2-related gene 1, BRG1), BAF155, and BAF170. Variant subunits include BAF53 (A or B), BAF60 (A, B or C), BAF 57, BAF45 (A, B, C, or D). Other subunits include ARIDI1A (also known as SMARCF1), ARID1B, SMARCA2 (also known as brahma homologue, BRM), ATRX, BAF200, BAF180 (also known as PBRM1), and bromodomain-containing 7 (BRD7). The at least one component of the SWI/SNF complex can by any component of the complex, for example, the component/subunit described herein or known in the art.
[0054] In any methods presented herein, neuronal differentiation gene may be, but is not limited to, CD133 (also called PROM1), DOCK4, PTPRK, PROM2, LHX1, LHX6, LHX9, PAX6, PAX7, VEFGA, FZD3B, FYN, HIF1A, HTRA2, EVX1, CCDC64, or GFAP.
[0055] In any methods presented herein, cell cycle inhibition gene may be, but is not limited to, CKDN1A, CDKN2A, MEN1, CHEK1, IRF6, ALOX15B, CYP27B1, DBC1, NME6, GMNN, HEXIM1, LATS1, MYC, HRAS, TGFB1, IFNG, WNT1, TP53, THBS1, INHBA, IL8, IRF1, TPR, BMP2, BMP4, ETS1, HPGD, BMP7, GATA3, NR2F2, APC, PTPN3, CALR, IL12A, IL12B, PML, CDKN2B, CDKN2C, CDKN1B, SOX2, TAF6, DNA2, PLK1, TERF1, GAS1, CDKN2D, MLF1, PTEN, TGFB2, SMAD3, FOXO4, CDK6, TFAP4, MAP2K1, NOTCH2, FOXC1, DLG1, MAD2L1, ATM, NAE1, DGKZ, FHL1, SCRIB, BTG3, PTPRK, RPS6KA2, STK11, CDKN3, TBRG1, CDC73, THAP5, CRLF3, DCUN1D3, MYOCD, PAF1, LILRB1, UHMK1, PNPT1, USP47, HEXIM2, CDK5RAP1, NKX3-1, TIPIN, PCBP4, USP44, RBM38, CDT1, RGCC, RNF167, CLSPN, CHMP1A, WDR6, TCF7L2, LATS2, RASSF1, MLTK, MAD2L2, FBXO5, ING4, or TRIM35.
[0056] In any methods presented herein, tumor suppressor gene may be, but is not limited to, BIN1. As used herein, the term "tumor suppressor gene" has its commonly understood meaning in the art, i.e. a gene whose expression and normal function act to suppress the neoplastic phenotype or induce apoptosis, or both. In some embodiments, tumor suppressor genes include cell cycle inhibition genes. Exemplary categories of tumor suppressors based on their functions include, but not limited to:
(1) genes that inhibit cell cycles; (2) genes that are coupling the cell cycle to DNA damage. When there is damaged DNA in the cell, the cell should not divide. If the damage can be repaired, the cell cycle can continue. If the damage cannot be repaired, the cell should initiate apoptosis (programmed cell death); (3) genes that prevent tumor cells from dispersing, block loss of contact inhibition, and inhibit metastasis. These genes and their encoded proteins are also known as metastasis suppressors; and (4) DNA repair proteins. Mutations in these genes increase the risk of cancer.
[0057] In any methods presented herein, hedgehog signaling pathway gene may be, but is not limited to, GLI1, PTCH1, SUFU, KIF7, GLI2, BMP4, MAP3K10, SHH, TCTN3, DYRK2, PTCHD1, or SMO.
[0058] In any methods presented herein, myc pathway gene may be, but is not limited to, MYC NMI, NFYC, NFYB, Cyclin T1, RuvB-like 1, GTF2I, BRCA1, T-cell lymphoma invasion and metastasis-inducing protein 1, ACTL6A, PCAF, MYCBP2, MAPK8, Bcl-2, Transcription initiation protein SPT3 homolog, SAP130, DNMT3A, mothers against decapentaplegic homolog 3, MAX, mothers against decapentaplegic homolog 2, MYCBP, HTATIP, ZBTB17, Transformation/transcription domain-associated protein, TADA2L, PFDN5, MAPK1, TFAP2A, P73, TAF9, YY1, SMARCB1, SMARCA4, MLH1, EP400 or let-7.
[0059] In any methods presented herein, histone methyltransferase gene may be, but is not limited to, EZH2.
[0060] Compounds of the present invention inhibit the histone methyltransferase activity of EZH2 or a mutant thereof and, accordingly, in one aspect of the invention, compounds disclosed herein are candidates for treating or preventing certain conditions and diseases. The present invention provides methods for treating, preventing or alleviating a symptom of cancer or a precancerous condition. The method includes administering to a subject in need thereof, a therapeutically effective amount of a compound of the present invention, or a pharmaceutically acceptable salt, polymorph, solvate, or stereoisomer thereof. Exemplary cancers that may be treated include medulloblastoma, oligodendroglioma, ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, ovarian serous adenocarcinoma, pancreatic ductal adenocarcinoma, pancreatic endocrine tumor, malignant rhabdoid tumor, astrocytoma, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, ependymoma, glioblastoma, meningioma, neuroglial tumor, oligoastrocytoma, oligodendroglioma, pineoblastoma, carcinosarcoma, chordoma, extragonadal germ cell tumor, extrarenal rhabdoid tumor, schwannoma, skin squamous cell carcinoma, chondrosarcoma, clear cell sarcoma of soft tissue, ewing sarcoma, gastrointestinal stromal tumor, osteosarcoma, rhabdomyosarcoma, epithelioid sarcoma, renal medullo carcinoma, diffuse large B-cell lymphoma, follicular lymphoma and not otherwise specified (NOS) sarcoma. Alternatively, cancers to be treated by the compounds of the present invention are non NHL cancers.
[0061] The present invention further provides the use of a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof in the treatment of cancer or precancer, or, for the preparation of a medicament useful for the treatment of such cancer or pre-cancer. Exemplary cancers that may be treated include medulloblastoma, oligodendroglioma, ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, ovarian serous adenocarcinoma, pancreatic ductal adenocarcinoma, pancreatic endocrine tumor, malignant rhabdoid tumor, astrocytoma, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, ependymoma, glioblastoma, meningioma, neuroglial tumor, oligoastrocytoma, oligodendroglioma, pineoblastoma, carcinosarcoma, chordoma, extragonadal germ cell tumor, extrarenal rhabdoid tumor, schwannoma, skin squamous cell carcinoma, chondrosarcoma, clear cell sarcoma of soft tissue, ewing sarcoma, gastrointestinal stromal tumor, osteosarcoma, rhabdomyosarcoma, epithelioid sarcoma, renal medullo carcinoma, diffuse large B-cell lymphoma, follicular lymphoma and not otherwise specified (NOS) sarcoma. Alternatively, the compound of the present invention can be used for the treatment of non NHL cancers, or, for the preparation of a medicament useful for the treatment of non NHL cancers.
[0062] The compounds of this invention can be used to modulate protein (e.g., histone) methylation, e.g., to modulate histone methyltransferase or histone demethylase enzyme activity. The compounds of the invention can be used in vivo or in vitro for modulating protein methylation. Based upon the surprising discovery that methylation regulation by EZH2 involves in tumor formation, particular tumors bearing altered expression and/or loss of function of selected biomarkers/genes, the compounds described herein are suitable candidates for treating these diseases, i.e., to decrease methylation or restore methylation to roughly its level in counterpart normal cells.
[0063] In some embodiments, compounds of the present invention can selectively inhibit proliferation of the SWI/SNF complex associated tumor or tumor cells (as shown in FIGS. 1-9). Accordingly, the present invention provides methods for treating, preventing or alleviating a symptom of the SWI/SNF complex associated cancer or a precancerous condition by a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof. The present invention further provides the use of a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof in the treatment of the SWI/SNF complex associated cancer or a precancer condition, or, for the preparation of a medicament useful for the treatment of such cancer or pre-cancer.
[0064] Also provided in the present invention are methods for determining responsiveness of a subject having a cancer to an EZH2 inhibitor. The method includes the steps of obtaining a sample (a nucleic acid sample or a protein sample) from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of at least one component of the SWI/SNF complex, detecting the expression and/or function of this component, and the presence of such reduced expression, haploinsufficiency, and/or loss of function indicates that the subject is responsive to the EZH2 inhibitor. The term "sample" means any biological sample derived from the subject, includes but is not limited to, cells, tissues samples, body fluids (including, but not limited to, mucus, blood, plasma, serum, urine, saliva, and semen), tumor cells, and tumor tissues. Samples can be provided by the subject under treatment or testing. Alternatively samples can be obtained by the physician according to routine practice in the art.
[0065] The present invention also provides methods for determining predisposition of a subject to a cancer or a precancerous condition by obtaining a sample from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of at least one component of the SWI/SNF complex, and the presence of such reduced expression, haploinsufficiency, and/or loss of function indicates that the subject is predisposed to (i.e., having higher risk of) developing the cancer or the precancerous condition compared to a subject without such loss of function of the at least one component of the SWI/SNF complex.
[0066] The term "predisposed" as used herein in relation to cancer or a precancerous condition is to be understood to mean the increased probability (e.g., at least 1%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, 150%, 200%, or more increase in probability) that a subject with reduced expression, haploinsufficiency, and/or loss of function of at least one component of the SWI/SNF complex, will suffer cancer or a precancerous condition, as compared to the probability that another subject not having reduced expression, haploinsufficiency, and/or loss of function of at least one component of the SWI/SNF complex, will suffer cancer or a precancerous condition, under circumstances where other risk factors (e.g., chemical/environment, food, and smoking history, etc.) for having cancer or a precancerous condition between the subjects are the same.
[0067] "Risk" in the context of the present invention, relates to the probability that an event will occur over a specific time period and can mean a subject's "absolute" risk or "relative" risk. Absolute risk can be measured with reference to either actual observation post-measurement for the relevant time cohort, or with reference to index values developed from statistically valid historical cohorts that have been followed for the relevant time period. Relative risk refers to the ratio of absolute risks of a subject compared either to the absolute risks of low risk cohorts or an average population risk, which can vary by how clinical risk factors are assessed. Odds ratios, the proportion of positive events to negative events for a given test result, are also commonly used (odds are according to the formula p/(1-p) where p is the probability of event and (1-p) is the probability of no event) to no-conversion.
[0068] Accordingly, the present invention provides personalized medicine, treatment and/or cancer management for a subject by genetic screening of reduced expression, haploinsufficiency, and/or loss of function of at least one component of the SWI/SNF complex in the subject. For example, the present invention provides methods for treating, preventing or alleviating a symptom of cancer or a precancerous condition by determining responsiveness of the subject to an EZH2 inhibitor and when the subject is responsive to the EZH2 inhibitor, administering to the subject a therapeutically effective amount of the EZH2 inhibitor, or a pharmaceutically acceptable salt, solvate, or stereoisomer thereof. The responsiveness is determined by obtaining a sample from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of at least one component of the SWI/SNF complex (such as SNF5, ARID1A or ATRX), and the presence of such loss of function indicates that the subject is responsive to the EZH2 inhibitor.
[0069] In other example, the present invention provides methods of cancer management in a subject by determining predisposition of the subject to a cancer or a precancerous condition periodically. The methods include steps of obtaining a sample from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of at least one component of the SWI/SNF complex, and the presence of such reduced expression, haploinsufficiency, and/or loss of function indicates that the subject is predisposed to developing the cancer or the precancerous condition compared to a subject without such reduced expression, haploinsufficiency, and/or loss of function of the at least one component of the SWI/SNF complex.
[0070] In merely illustrative embodiments, the methods of treatment presented herein include steps of (a) collecting a nucleic acid sample or a protein sample from a biological sample obtained from a subject, (b) measuring the expression level or function level of a component of the SWI/SNF complex in the sample, (c) measuring the expression level or function level of the component of the SWI/SNF in a control sample; (d) comparing the expression level or the function level of the component measured in step (b) in the tested sample to the expression level or the function level of the component measured in step (c) in the control sample (or a reference value); (e) identifying the subject as a candidate for treatment when the expression level or the function level of the component measured in step (b) is reduced or lost (e.g., haploinsufficiency or loss of function) compared to the expression level or the function level of the component measured in step (c); and (f) administering a therapeutically effective amount of an EZH2 inhibitor to the subject identified in step (e) or selecting a treatment regimen for the subject identified in step (e). The expression level or the function level of component in the subject sample is reduced, for example, 10%, 25%, 50% or 1-, 2-, 5- or more fold compared to the expression level or the function level of the component in the control sample. Any suitable methods known in the art can be utilized to measure the expression level or the function level of the component of the SWI/SNF complex. In some embodiments, the subject has malignant rhabdoid tumor, medulloblastoma or atypical teratoid rhabdoid tumor. In some embodiments, the component is SNF5, ARID1A or ATRX.
[0071] For example, the identified subject can be treated with the standard of care treatment as described in the most current National Comprehensive Cancer Network (NCCN) guidelines.
[0072] For example, a control sample is obtained from a healthy, normal subject. Alternatively, a control sample is obtained from a subject who is not suffering, has not been diagnosed, or is not at risk of developing cancer associated with the SWI/SNF complex.
[0073] In one preferred aspect, the present invention provides a method for treating or alleviating a symptom of cancer in a subject by determining responsiveness of the subject to an EZH2 inhibitor and administering to the subject a therapeutically effective amount of the EZH2 inhibitor if the subject is responsive to the EZH2 inhibitor and the subject has a cancer selected from the group consisting of brain and CNS cancer, kidney cancer, ovarian cancer, pancreatic cancer, leukemia, lymphoma, myeloma, and/or sarcoma. Such responsiveness is determined by obtaining a sample from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of SNF5, ARID1A, and/or ATRX, and the presence of the reduced expression, haploinsufficiency, and/or loss of function indicates the subject is responsive to the EZH2 inhibitor.
[0074] In another preferred aspect, the present invention provides a method for treating or alleviating a symptom of malignant rhabdoid tumor in a subject by determining responsiveness of the subject to an EZH2 inhibitor and administering to the subject a therapeutically effective amount of the EZH2 inhibitor if the subject is responsive to the EZH2 inhibitor. Such responsiveness is determined by obtaining a sample from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of SNF5, ARID1A, and/or ATRX, and the presence of the reduced expression, haploinsufficiency, and/or loss of function indicates the subject is responsive to the EZH2 inhibitor.
[0075] In another preferred aspect, the present invention provides a method for treating or alleviating a symptom of medulloblastoma in a subject by determining responsiveness of the subject to an EZH2 inhibitor and administering to the subject a therapeutically effective amount of the EZH2 inhibitor if the subject is responsive to the EZH2 inhibitor. Such responsiveness is determined by obtaining a sample from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of SNF5, ARID1A, and/or ATRX, and the presence of the reduced expression, haploinsufficiency, and/or loss of function indicates the subject is responsive to the EZH2 inhibitor.
[0076] In another preferred aspect, the present invention provides a method for treating or alleviating a symptom of atypical teratoid rhabdoid tumor in a subject by determining responsiveness of the subject to an EZH2 inhibitor and administering to the subject a therapeutically effective amount of the EZH2 inhibitor if the subject is responsive to the EZH2 inhibitor. Such responsiveness is determined by obtaining a sample from the subject and detecting reduced expression, haploinsufficiency, and/or loss of function of SNF5, ARID1A, and/or ATRX, and the presence of the reduced expression, haploinsufficiency, and/or loss of function indicates the subject is responsive to the EZH2 inhibitor.
[0077] Malignant rhabdoid tumors (MRTs) and atypical teratoid rhabdoid tumors (ATRTs) are extremely aggressive pediatric cancers of the brain, kidney, and soft tissues that are highly malignant, locally invasive, frequently metastatic, and particularly lethal. They are typically diploid and lack genomic aberrations; however, they are characterized by an almost complete penetrance of loss of SMARCB1 (also called SNF5, INI1 or BAF47), a core component of the SWI/SNF chromatin remodeling complex. The biallelic inactivation of SMARCB1 is in essence the sole genetic event in MRTs and ATRTs which suggests a driver role for this genetic aberration.
[0078] Without being bound by any theory, a compound of the present invention specifically inhibits cellular H3K27 methylation leading to selective apoptotic killing of SMARCB1 mutant MRT cells. For example, in vitro treatment of SMARCB1-deleted MRT cell lines with Compound A induced strong anti-proliferative effects with IC.sub.50 values in the nM range; while the control (wild-type) cell lines were minimally affected (FIG. 10C and table 6). Furthermore, the compound of the present invention induces genes of neuronal differentiation, cell cycle inhibition and tumor suppression while suppressing expression of hedgehog pathway genes, MYC and EZH2. For example, Compound A treatment of G401 SMARCB1-deleted cells for up to 7 days strongly induced expression of CD133, DOCK4 and PTPRK and up-regulated cell cycle inhibitors CDKAT1A and CDKN2A and tumor suppressor BIN1, all in a time-dependent manner (FIG. 14B). Simultaneously, the expression of hedgehog pathway genes, MYC and EZH2 were reduced. Notably, G402 SMARCB1-deleted cells exposed to Compound A for 14 days assumed a neuron-like morphology (FIG. 14C).
[0079] Accordingly, the present invention further provides methods of treating or alleviating a symptom of cancer in a subject in need thereof by (a) determining the expression level of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes and tumor suppressor genes in a sample obtained from the subject; (b) selecting a subject having a decreased expression level of at least one gene in step (a); and (c) administering to the subject selected in step (b) an effective amount of a compound of the invention, thus treating or alleviating a symptom of cancer in the subject.
[0080] The present invention also provides methods of treating or alleviating a symptom of cancer in a subject in need thereof by (a) determining the expression level of at least one gene selected from the group consisting of hedgehog pathway genes, myc pathway genes and histone methyltransferase genes in a sample obtained from the subject; (b) selecting a subject having an increased expression level of at least one gene in step (a); and (c) administering to the subject selected in step (b) an effective amount of a compound of the invention, thus treating or alleviating a symptom of cancer in the subject.
[0081] Also provided herein are methods of selecting a cancer therapy for a subject in need thereof by (a) determining the expression level of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes, and tumor suppressor genes in a sample obtained from the subject, and (b) selecting a cancer therapy when the subject has a decreased expression level of at least one gene in step (a), where the cancer therapy includes the administration of an effective amount of a compound of the invention to the subject.
[0082] The present invention further provides methods of selecting a cancer therapy for a subject in need thereof by (a) determining the expression level of at least one gene selected from the group consisting of hedgehog pathway genes, myc pathway genes and histone methyltransferase genes in a sample obtained from the subject, and (b) selecting a cancer therapy when the subject has an increased expression level of at least one gene in step (a), where the cancer therapy includes the administration of an effective amount of a compound of the invention to the subject.
[0083] In merely illustrative embodiments, the methods presented herein may include the steps of (a) collecting a nucleic acid or a protein sample from a biological sample obtained from a subject, (b) measuring the expression level of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes, and tumor suppressor genes in the sample, (c) measuring the expression level of the same gene(s) in a control sample; (d) comparing the expression level of the gene measured in step (b) in the tested sample to the expression level of the gene measured in step (c) in the control sample (or to a reference value); (e) identifying the subject as a candidate for treatment when the expression level of the component measured in step (b) is reduced compared to the expression level of the gene measured in step (c); and (f) administering a therapeutically effective amount of an EZH2 inhibitor to the subject identified in step (e) or selecting a treatment regimen for the subject identified in step (e). The expression level of the gene in the tested subject is reduced, for example, 10%, 25%, 50% or 1-, 2-, 5- or more fold compared to the expression level of the gene in the control sample.
[0084] In merely illustrative embodiments, the methods presented herein may include the steps of (a) collecting a nucleic acid or a protein sample from a biological sample obtained from a subject, (b) measuring the expression level of at least one gene selected from the group consisting of hedgehog pathway genes, myc pathway genes and histone methyltransferase genes in the sample, (c) measuring the expression level of the same gene(s) in a control sample; (d) comparing the expression level of the gene measured in step (b) in the tested sample to the expression level of the gene measured in step (c) in the control sample (or to a reference value); (e) identifying the subject as a candidate for treatment when the expression level of the component measured in step (b) is increased compared to the expression level of the gene measured in step (c); and (f) administering a therapeutically effective amount of an EZH2 inhibitor to the subject identified in step (e) or selecting a treatment regimen for the subject identified in step (e). The expression level of the gene in the tested subject is increased, for example, 10%, 25%, 50% or 1-, 2-, 5- or more fold compared to the expression level of the gene in the control sample.
[0085] The term "expression level" refers to protein, RNA, or mRNA level of a particular gene of interest. Any methods known in the art can be utilized to determine the expression level of a particular gene of interest. Examples include, but are not limited to, reverse transcription and amplification assays (such as PCR, ligation RT-PCR or quantitative RT-PCT), hybridization assays, Northern blotting, dot blotting, in situ hybridization, gel electrophoresis, capillary electrophoresis, column chromatography, Western blotting, immunohistochemistry, immunostaining, or mass spectrometry. Assays can be performed directly on biological samples or on protein/nucleic acids isolated from the samples. It is routine practice in the relevant art to carry out these assays. For example, the measuring step in any method described herein includes contacting the nucleic acid sample from the biological sample obtained from the subject with one or more primers that specifically hybridize to the gene of interest presented herein. Alternatively, the measuring step of any method described herein includes contacting the protein sample from the biological sample obtained from the subject with one or more antibodies that bind to the biomarker of the interest presented herein.
[0086] A decreased expression level of a particular gene means a decrease in its expression level by at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 300%, 400%, 500%, 1000%, 1500%, or more compared to a reference value or the expression level of this gene measured in a different (or previous) sample obtained from the same subject.
[0087] An increased expression level of a particular gene means an increase in its expression level by at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 300%, 400%, 500%, 1000%, 1500%, or more compared to a reference value or the expression level of this gene measured in a different (or previous) sample obtained from the same subject.
[0088] A "reference or baseline level/value" as used herein can be used interchangeably and is meant to be relative to a number or value derived from population studies, including without limitation, such subjects having similar age range, disease status (e.g., stage), subjects in the same or similar ethnic group, or relative to the starting sample of a subject undergoing treatment for cancer. Such reference values can be derived from statistical analyses and/or risk prediction data of populations obtained from mathematical algorithms and computed indices of cancer. Reference indices can also be constructed and used using algorithms and other methods of statistical and structural classification.
[0089] In some embodiments of the present invention, the reference or baseline value is the expression level of a particular gene of interest in a control sample derived from one or more healthy subjects or subjects who have not been diagnosed with any cancer.
[0090] In some embodiments of the present invention, the reference or baseline value is the expression level of a particular gene of interest in a sample obtained from the same subject prior to any cancer treatment. In other embodiments of the present invention, the reference or baseline value is the expression level of a particular gene of interest in a sample obtained from the same subject during a cancer treatment. Alternatively, the reference or baseline value is a prior measurement of the expression level of a particular gene of interest in a previously obtained sample from the same subject or from a subject having similar age range, disease status (e.g., stage) to the tested subject.
[0091] In some embodiments, an effective amount means an amount sufficient to increase the expression level of at least one gene which is decreased in the subject prior to the treatment or an amount sufficient to alleviate one or more symptoms of cancer. For example, an effective amount is an amount sufficient to increase the expression level of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes, and tumor suppressor genes by at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 300%, 400%, 500%, 1000%, 1500%, or more compared to a reference value or the expression level without the treatment of any compound.
[0092] In some embodiments, an effective amount means an amount sufficient to decrease the expression level of at least one gene which is increased in the subject prior to the treatment or an amount sufficient to alleviate one or more symptoms of cancer. For example, an effective amount is an amount sufficient to decrease the expression level of at least one gene selected from the group consisting of hedgehog pathway genes, MYC and EZH2 by at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 300%, 400%, 500%, 1000%, 1500%, or more compared to a reference value or the expression level without the treatment of any compound.
[0093] The precise effective amount for a subject will depend upon the subject's body weight, size, and health; the nature and extent of the condition; and the therapeutic selected for administration. An effective amount for a given situation can be determined by routine experimentation that is within the skill and judgment of the clinician.
[0094] The present invention further provides a method of determining efficacy of a cancer treatment in a subject in need thereof by (a) measuring the expression level of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes, and tumor suppressor genes in a sample obtained from the subject, (b) comparing the expression level of at least one gene in step (a) to a reference value or a prior measurement, and (c) determining the efficacy of the cancer treatment based on the comparison step. An exemplary cancer treatment is administering a compound of the invention to the tested subject.
[0095] The treatment is effective when the tested subject has an increased expression of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes and tumor suppressor genes 1) compared to a reference value or a prior measurement; or 2) over the period of time being monitored, such as 1, 2, 3, 4, 5, 6, or 7 days, or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 weeks, or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12 months or longer. When the existing treatment is not effective, a new treatment or an increased dosage of the existing treatment (for example, increasing the dosage of the compound administered to the subject) should be sought for the tested subject.
[0096] The present invention also provides a method of determining efficacy of a cancer treatment in a subject in need thereof by (a) measuring the expression level of at least one gene selected from the group consisting of hedgehog pathway genes, myc pathway genes and histone methyltransferase genes in a sample obtained from the subject, (b) comparing the expression level of at least one gene in step (a) to a reference value or a prior measurement, and (c) determining the efficacy of the cancer treatment based on the comparison step. An exemplary cancer treatment is administering an EZH2 inhibitor of the invention to the tested subject.
[0097] For example, the treatment is effective when the tested subject has a decreased expression of at least one gene selected from the group consisting of hedgehog pathway genes, myc pathway genes and histone methyltransferase genes 1) compared to a reference value or a prior measurement; or 2) over the period of time being monitored, such as 1, 2, 3, 4, 5, 6, or 7 days, or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 weeks, or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12 months or longer. When the existing treatment is not effective, a new treatment or an increased dosage of the existing treatment (for example, increasing the dosage of the compound administered to the subject) should be sought for the tested subject.
[0098] In any methods presented herein, cancer is selected from the group consisting of brain and central nervous system (CNS) cancer, head and neck cancer, kidney cancer, ovarian cancer, pancreatic cancer, leukemia, lung cancer, lymphoma, myeloma, sarcoma, breast cancer, and prostate cancer. Preferably, cancer is selected from the group consisting of medulloblastoma, oligodendroglioma, ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, ovarian serous adenocarcinoma, pancreatic ductal adenocarcinoma, pancreatic endocrine tumor, malignant rhabdoid tumor, astrocytoma, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, ependymoma, glioblastoma, meningioma, neuroglial tumor, oligoastrocytoma, oligodendroglioma, pineoblastoma, carcinosarcoma, chordoma, extragonadal germ cell tumor, extrarenal rhabdoid tumor, schwannoma, skin squamous cell carcinoma, chondrosarcoma, clear cell sarcoma of soft tissue, ewing sarcoma, gastrointestinal stromal tumor, osteosarcoma, rhabdomyosarcoma, epithelioid sarcoma, renal medullo carcinoma, diffuse large B-cell lymphoma, follicular lymphoma and not otherwise specified (NOS) sarcoma. More preferably, cancer is medulloblastoma, malignant rhabdoid tumor, or atypical teratoid rhabdoid tumor.
[0099] As used herein, the term "responsiveness" is interchangeable with terms "responsive", "sensitive", and "sensitivity", and it is meant that a subject is showing therapeutic responses when administered an EZH inhibitor, e.g., tumor cells or tumor tissues of the subject undergo apoptosis and/or necrosis, and/or display reduced growing, dividing, or proliferation. This term is also meant that a subject will or has a higher probability, relative to the population at large, of showing therapeutic responses when administered an EZH inhibitor, e.g., tumor cells or tumor tissues of the subject undergo apoptosis and/or necrosis, and/or display reduced growing, dividing, or proliferation.
[0100] As used herein, a "subject" is interchangeable with a "subject in need thereof", both of which refer to a subject having a disorder in which EZH2-mediated protein methylation plays a part, or a subject having an increased risk of developing such disorder relative to the population at large. A subject in need thereof may be a subject having a disorder associated with SWI/SNF complex. A subject in need thereof can have a precancerous condition. Preferably, a subject in need thereof has cancer. A subject in need thereof can have cancer associated with SWI/SNF complex. A subject in need thereof can have cancer associated with loss of function in at least one component of SWI/SNF complex. In a preferred aspect, a subject in need thereof has one or more cancers selected from the group consisting of brain and central nervous system (CNS) cancer, head and neck cancer, kidney cancer, ovarian cancer, pancreatic cancer, leukemia, lung cancer, lymphoma, myeloma, sarcoma, breast cancer, and prostate cancer. Preferably, a subject in need thereof has medulloblastoma, oligodendroglioma, ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, ovarian serous adenocarcinoma, pancreatic ductal adenocarcinoma, pancreatic endocrine tumor, malignant rhabdoid tumor, astrocytoma, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, ependymoma, glioblastoma, meningioma, neuroglial tumor, oligoastrocytoma, oligodendroglioma, pineoblastoma, carcinosarcoma, chordoma, extragonadal germ cell tumor, extrarenal rhabdoid tumor, schwannoma, skin squamous cell carcinoma, chondrosarcoma, clear cell sarcoma of soft tissue, ewing sarcoma, gastrointestinal stromal tumor, osteosarcoma, rhabdomyosarcoma, epithelioid sarcoma, renal medullo carcinoma, diffuse large B-cell lymphoma, follicular lymphoma and not otherwise specified (NOS) sarcoma. Alternatively, a subject in need thereof has a non NHL cancer.
[0101] As used herein, a "subject" includes a mammal. The mammal can be e.g., a human or appropriate non-human mammal, such as primate, mouse, rat, dog, cat, cow, horse, goat, camel, sheep or a pig. The subject can also be a bird or fowl. In one embodiment, the mammal is a human. A subject can be male or female.
[0102] A subject in need thereof can be one who has not been previously diagnosed or identified as having cancer or a precancerous condition. A subject in need thereof can be one who has been previously diagnosed or identified as having cancer or a precancerous condition. A subject in need thereof can also be one who is having (suffering from) cancer or a precancerous condition. Alternatively, a subject in need thereof can be one who has a risk of developing such disorder relative to the population at large (i.e., a subject who is predisposed to developing such disorder relative to the population at large).
[0103] Optionally a subject in need thereof has already undergone, is undergoing or will undergo, at least one therapeutic intervention for the cancer or precancerous condition.
[0104] A subject in need thereof may have refractory cancer on most recent therapy. "Refractory cancer" means cancer that does not respond to treatment. The cancer may be resistant at the beginning of treatment or it may become resistant during treatment. Refractory cancer is also called resistant cancer. In some embodiments, the subject in need thereof has cancer recurrence following remission on most recent therapy. In some embodiments, the subject in need thereof received and failed all known effective therapies for cancer treatment. In some embodiments, the subject in need thereof received at least one prior therapy.
[0105] A subject in need thereof may be one who had, is having or is predisposed to developing a cancer or a precancerous condition associated with the SWI/SNF complex. A subject in need thereof may be one who had, is having or is predisposed to developing cancer or a precancerous condition associated with loss of function of at least one component of the SWI/SNF complex. In a preferred aspect, a subject in need thereof is one who had, is having or is predisposed to developing one or more cancers selected from the group consisting of brain and central nervous system (CNS) cancer, head and neck cancer, kidney cancer, ovarian cancer, pancreatic cancer, leukemia, lung cancer, lymphoma, myeloma, sarcoma, breast cancer, and prostate cancer. Preferably, a subject in need thereof is one who had, is having or is predisposed to developing brain and CNS cancer, kidney cancer, ovarian cancer, pancreatic cancer, leukemia, lymphoma, myeloma, and/or sarcoma. Exemplary brain and central CNS cancer includes medulloblastoma, oligodendroglioma, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, ependymoma, glioblastoma, meningioma, neuroglial tumor, oligoastrocytoma, oligodendroglioma, and pineoblastoma. Exemplary ovarian cancer includes ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, and ovarian serous adenocarcinoma. Exemplary pancreatic cancer includes pancreatic ductal adenocarcinoma and pancreatic endocrine tumor. Exemplary sarcoma includes chondrosarcoma, clear cell sarcoma of soft tissue, ewing sarcoma, gastrointestinal stromal tumor, osteosarcoma, rhabdomyosarcoma, and not otherwise specified (NOS) sarcoma. Alternatively, cancers to be treated by the compounds of the present invention are non NHL cancers.
[0106] Alternatively, a subject in need thereof is one who had, is having or is predisposed to developing one or more cancers selected from the group consisting of medulloblastoma, oligodendroglioma, ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, ovarian serous adenocarcinoma, pancreatic ductal adenocarcinoma, pancreatic endocrine tumor, malignant rhabdoid tumor, astrocytoma, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, ependymoma, glioblastoma, meningioma, neuroglial tumor, oligoastrocytoma, oligodendroglioma, pineoblastoma, carcinosarcoma, chordoma, extragonadal germ cell tumor, extrarenal rhabdoid tumor, schwannoma, skin squamous cell carcinoma, chondrosarcoma, clear cell sarcoma of soft tissue, ewing sarcoma, gastrointestinal stromal tumor, osteosarcoma, rhabdomyosarcoma, and not otherwise specified (NOS) sarcoma. Preferably, a subject is one who had, is having or is predisposed to developing medulloblastoma, ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, pancreatic ductal adenocarcinoma, malignant rhabdoid tumor, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, glioblastoma, meningioma, pineoblastoma, carcinosarcoma, extrarenal rhabdoid tumor, schwannoma, skin squamous cell carcinoma, chondrosarcoma, ewing sarcoma, epithelioid sarcoma, renal medullo carcinoma, diffuse large B-cell lymphoma, follicular lymphoma and/or NOS sarcoma. More preferably, a subject in need thereof is one who had, is having or is predisposed to developing malignant rhabdoid tumor, medulloblastoma and/or atypical teratoid rhabdoid tumor.
[0107] In some embodiments of the present invention, a subject in need thereof has a decreased expression level of at least one gene selected from the group consisting of neuronal differentiation genes, cell cycle inhibition genes, and tumor suppressor genes.
[0108] In some embodiments, a subject in need thereof has an increased expression level of at least one gene selected from the group consisting of hedgehog pathway genes, myc pathway genes and histone methyltransferase genes.
[0109] In some embodiments of the present invention, a subject in need thereof has loss of function of at least one component/subunit of the SWI/SNF complex. Alternatively, a subject in need thereof has reduced expression or haploinsufficiency of at least one component/subunit of the SWI/SNF complex. In certain embodiments, a subject in need thereof has loss of function of SNF5 subunit.
[0110] In any method of the present invention, a subject in need thereof may have reduced expression, haploinsufficiency or loss of function of at least one signaling component downstream of SWI/SNF complex. Such downstream component includes, but is not limited to, polycomb complex (PcG) and its targets.
[0111] As used herein, the term "loss of function" refers to less or no function of a gene product/protein compared to the wild type. Loss of function of a SWI/SNF complex component means the component/subunit or the entire SWI/SNF complex has less or no biological function compared to the wild type component/subunit or the entire SWI/SNF complex, respectively. Loss of function can be caused by transcriptional, post-transcription, or post translational mechanisms. In one aspect of the present invention, loss of function is caused by loss of function mutation resulted from a point mutation (e.g., a substitution, a missense mutation, or a nonsense mutation), an insertion, and/or a deletion in a polypeptide of a SWI/SNF complex component or a nucleic acid sequence encoding a polypeptide of a SWI/SNF complex component. The mutations referred herein are somatic mutations. The term "somatic mutation" refers to a deleterious alteration in at least one gene allele that is not found in every cell of the body, but is found only in isolated cells. A characteristic of the somatic mutations as used herein is, that they are restricted to particular tissues or even parts of tissues or cells within a tissue and are not present in the whole organism harboring the tissues or cells. The term "wild-type" refers to a gene or gene product that has the characteristics of that gene or gene product when isolated from a naturally occurring source. A wild-type gene is that which is most frequently observed in a population and is thus arbitrarily designed the "normal" or "wild-type" form of the gene.
[0112] Accordingly, a loss of function mutation or a reduced expression can be detected using any suitable method available in the art. For example, a loss of function mutation can be detected by measuring the biological function of a gene product, such as the ATP-dependent chromatin remodeling activity of the SWI/SNF complex. Alternatively, a loss of function mutation can be determined by detecting any alternation in a nucleic acid sequence encoding a component of the SWI/SNF complex. For example, a nucleic acid sequence encoding a component of the SWI/SNF complex having a loss of function mutation can be detected by whole-genome resequencing or target region resequencing (the latter also known as targeted resequencing) using suitably selected sources of DNA and polymerase chain reaction (PCR) primers in accordance with methods well known in the art. The method typically and generally entails the steps of genomic DNA purification, PCR amplification to amplify the region of interest, cycle sequencing, sequencing reaction cleanup, capillary electrophoresis, and/or data analysis. Alternatively or in addition, the method may include the use of microarray-based targeted region genomic DNA capture and/or sequencing. Kits, reagents, and methods for selecting appropriate PCR primers and performing resequencing are commercially available, for example, from Applied Biosystems, Agilent, and NimbleGen (Roche Diagnostics GmbH). Alternatively or in addition, a nucleic acid sequence encoding a SWI/SNF polypeptide having a loss of function mutation may be detected using a Southern blot in accordance with methods well known in the art. Optionally, a loss of function mutation can be detected by measuring the absence of the expression of a component polypeptide or by measuring the expression of the mutant component polypeptide. Detection of (mutant) polypeptide expression can be carried out with any suitable immunoassay in the art, such as Western blot analysis.
[0113] Human nucleic acid and amino acid sequence of components of the SWI/SNF complex have previously been described. See, e.g., GenBank Accession Nos NP_003064.2, NM_003073.3, NP_001007469.1, and NM_001007468.1 for SNF5, GenBank Accession Nos NM_000489.3, NP_000480.2, NM_138270.2, and NP_612114.1 for ATRX, GenBank Accession Nos NP_006006.3, NM_006015.4, NP_624361.1, and NM_139135.2 for ARID1A, each of which is incorporated herein by reference in its entirety.
[0114] Spectrum of hSNF5 somatic mutations in human has also been described in Sevenet et al., Human Molecular Genetics, 8: 2359-2368, 1999, which is incorporated herein by reference in its entirety.
[0115] A subject in need thereof may have reduced expression, haploinsufficiency, and/or loss of function of SNF5. For example, a subject can comprise a deletion of SNF5 in SNF5 polypeptide or a nucleic acid sequence encoding a SNF5 polypeptide.
TABLE-US-00001 SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily B member 1 isoform a (SMARCB1, also called SNF5)[Homo sapiens] (SEQ ID NO: 1) 1 mmmmalsktf gqkpvkfqle ddgefymigs evgnylrmfr gslykrypsl wrrlatveer 61 kkivasshgk ktkpntkdhg yttlatsvtl lkaseveeil dgndekykav sistepptyl 121 reqkakrnsq wvptlpnssh hldavpcstt inrnrmgrdk krtfplcfdd hdpavihena 181 sqpevlvpir ldmeidgqkl rdaftwnmne klmtpemfse ilcddldlnp ltfvpaiasa 241 irqqiesypt dsiledqsdq rviiklnihv gnislvdqfe wdmsekensp ekfalklcse 301 lglggefvtt iaysirgqls whqktyafse nplptveiai rntgdadqwc plletltdae 361 mekkirdqdr ntrrmrrlan tapaw Homo sapiens SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 (SMARCB1, also called SNF5), transcript variant 1, mRNA (SEQ ID NO: 2) 1 aacgccagcg cctgcgcact gagggcggcc tggtcgtcgt ctgcggcggc ggcggcggct 61 gaggagcccg gctgaggcgc cagtacccgg cccggtccgc atttcgcctt ccggcttcgg 121 tttccctcgg cccagcacgc cccggccccg ccccagccct cctgatccct cgcagcccgg 181 ctccggccgc ccgcctctgc cgccgcaatg atgatgatgg cgctgagcaa gaccttcggg 241 cagaagcccg tgaagttcca gctggaggac gacggcgagt tctacatgat cggctccgag 301 gtgggaaact acctccgtat gttccgaggt tctctgtaca agagataccc ctcactctgg 361 aggcgactag ccactgtgga agagaggaag aaaatagttg catcgtcaca tggtaaaaaa 421 acaaaaccta acactaagga tcacggatac acgactctag ccaccagtgt gaccctgtta 481 aaagcctcgg aagtggaaga gattctggat ggcaacgatg agaagtacaa ggctgtgtcc 541 atcagcacag agccccccac ctacctcagg gaacagaagg ccaagaggaa cagccagtgg 601 gtacccaccc tgcccaacag ctcccaccac ttagatgccg tgccatgctc cacaaccatc 661 aacaggaacc gcatgggccg agacaagaag agaaccttcc ccctttgctt tgatgaccat 721 gacccagctg tgatccatga gaacgcatct cagcccgagg tgctggtccc catccggctg 781 gacatggaga tcgatgggca gaagctgcga gacgccttca cctggaacat gaatgagaag 841 ttgatgacgc ctgagatgtt ttcagaaatc ctctgtgacg atctggattt gaacccgctg 901 acgtttgtgc cagccatcgc ctctgccatc agacagcaga tcgagtccta ccccacggac 961 agcatcctgg aggaccagtc agaccagcgc gtcatcatca agctgaacat ccatgtggga 1021 aacatttccc tggtggacca gtttgagtgg gacatgtcag agaaggagaa ctcaccagag 1081 aagtttgccc tgaagctgtg ctcggagctg gggttgggcg gggagtttgt caccaccatc 1141 gcatacagca tccggggaca gctgagctgg catcagaaga cctacgcctt cagcgagaac 1201 cctctgccca cagtggagat tgccatccgg aacacgggcg atgcggacca gtggtgccca 1261 ctgctggaga ctctgacaga cgctgagatg gagaagaaga tccgcgacca ggacaggaac 1321 acgaggcgga tgaggcgtct tgccaacacg gccccggcct ggtaaccagc ccatcagcac 1381 acggctccca cggagcatct cagaagattg ggccgcctct cctccatctt ctggcaagga 1441 cagaggcgag gggacagccc agcgccatcc tgaggatcgg gtgggggtgg agtgggggct 1501 tccaggtggc ccttcccggc acacattcca tttgttgagc cccagtcctg ccccccaccc 1561 caccctccct acccctcccc agtctctggg gtcaggaaga aaccttattt taggttgtgt 1621 tttgtttttg tataggagcc ccaggcaggg ctagtaacag tttttaaata aaaggcaaca 1681 ggtcatgttc aatttcttca acaaaaaaaa aaaaaaa SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily B member 1 isoform b [Homo sapiens](SMARCB1, also called SNF5) (SEQ ID NO: 3) 1 mmmmalsktf gqkpvkfqle ddgefymigs evgnylrmfr gslykrypsl wrrlatveer 61 kkivasshdh gyttlatsvt llkaseveei ldgndekyka vsisteppty lreqkakrns 121 qwvptlpnss hhldavpcst tinrnrmgrd kkrtfplcfd dhdpavihen asqpevlvpi 181 rldmeidgqk lrdaftwnmn eklmtpemfs eilcddldln pltfvpaias airqqiesyp 241 tdsiledqsd qrviiklnih vgnislvdqf ewdmsekens pekfalklcs elglggefvt 301 tiaysirgql swhqktyafs enplptveia irntgdadqw cplletltda emekkirdqd 361 rntrrmrrla ntapaw Homo sapiens SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 (SMARCB1, also called SNF5), transcript variant 2, mRNA (SEQ ID NO: 4) 1 aacgccagcg cctgcgcact gagggcggcc tggtcgtcgt ctgcggcggc ggcggcggct 61 gaggagcccg gctgaggcgc cagtacccgg cccggtccgc atttcgcctt ccggcttcgg 121 tttccctcgg cccagcacgc cccggccccg ccccagccct cctgatccct cgcagcccgg 181 ctccggccgc ccgcctctgc cgccgcaatg atgatgatgg cgctgagcaa gaccttcggg 241 cagaagcccg tgaagttcca gctggaggac gacggcgagt tctacatgat cggctccgag 301 gtgggaaact acctccgtat gttccgaggt tctctgtaca agagataccc ctcactctgg 361 aggcgactag ccactgtgga agagaggaag aaaatagttg catcgtcaca tgatcacgga 421 tacacgactc tagccaccag tgtgaccctg ttaaaagcct cggaagtgga agagattctg 481 gatggcaacg atgagaagta caaggctgtg tccatcagca cagagccccc cacctacctc 541 agggaacaga aggccaagag gaacagccag tgggtaccca ccctgcccaa cagctcccac 601 cacttagatg ccgtgccatg ctccacaacc atcaacagga accgcatggg ccgagacaag 661 aagagaacct tccccctttg ctttgatgac catgacccag ctgtgatcca tgagaacgca 721 tctcagcccg aggtgctggt ccccatccgg ctggacatgg agatcgatgg gcagaagctg 781 cgagacgcct tcacctggaa catgaatgag aagttgatga cgcctgagat gttttcagaa 841 atcctctgtg acgatctgga tttgaacccg ctgacgtttg tgccagccat cgcctctgcc 901 atcagacagc agatcgagtc ctaccccacg gacagcatcc tggaggacca gtcagaccag 961 cgcgtcatca tcaagctgaa catccatgtg ggaaacattt ccctggtgga ccagtttgag 1021 tgggacatgt cagagaagga gaactcacca gagaagtttg ccctgaagct gtgctcggag 1081 ctggggttgg gcggggagtt tgtcaccacc atcgcataca gcatccgggg acagctgagc 1141 tggcatcaga agacctacgc cttcagcgag aaccctctgc ccacagtgga gattgccatc 1201 cggaacacgg gcgatgcgga ccagtggtgc ccactgctgg agactctgac agacgctgag 1261 atggagaaga agatccgcga ccaggacagg aacacgaggc ggatgaggcg tcttgccaac 1321 acggccccgg cctggtaacc agcccatcag cacacggctc ccacggagca tctcagaaga 1381 ttgggccgcc tctcctccat cttctggcaa ggacagaggc gaggggacag cccagcgcca 1441 tcctgaggat cgggtggggg tggagtgggg gcttccaggt ggcccttccc ggcacacatt 1501 ccatttgttg agccccagtc ctgcccccca ccccaccctc cctacccctc cccagtctct 1561 ggggtcagga agaaacctta ttttaggttg tgttttgttt ttgtatagga gccccaggca 1621 gggctagtaa cagtttttaa ataaaaggca acaggtcatg ttcaatttct tcaacaaaaa 1681 aaaaaaaaaa
[0116] A subject in need thereof may have reduced expression, haploinsufficiency, and/or loss of function of ATRX. For example, a subject can comprise a mutation selected from the group consisting of a substitution of asparagine (N) for the wild type residue lysine (K) at amino acid position 688 of SEQ ID NO: 5 (K688N), and a substitution of isoleucine (I) for the wild type residue methionine (M) at amino acid position 366 of SEQ ID NO: 5 (M366I).
TABLE-US-00002 Homo sapiens alpha thalassemia/mental retardation syndrome X-linked (ATRX) isoform 1 (SEQ ID NO: 5) 1 mtaepmsesk lntlvqklhd flahsseese etsspprlam nqntdkisgs gsnsdmmens 61 keegtsssek skssgssrsk rkpsivtkyv esddekpldd etvnedasne nsenditmqs 121 lpkgtvivqp epvlnedkdd fkgpefrsrs kmktenlkkr gedglhgivs ctacgqqvnh 181 fqkdsiyrhp slqvlicknc fkyymsddis rdsdgmdeqc rwcaeggnli ccdfchnafc 241 kkcilrnlgr kelstimden nqwycyichp eplldlvtac nsvfenleql lqqnkkkikv 301 dseksnkvye htsrfspkkt ssncngeekk lddscsgsvt ysysalivpk emikkakkli 361 ettanmnssy vkflkqatdn seissatklr qlkafksvla dikkahlale edlnsefram 421 davnkekntk ehkvidakfe tkarkgekpc alekkdisks eaklsrkqvd sehmhqnvpt 481 eeqrtnkstg gehkksdrke epqyepants edldmdivsv pssvpedife nletamevqs 541 svdhqgdgss gteqevesss vklnisskdn rggiksktta kvtkelyvkl tpvslsnspi 601 kgadcqevpq dkdgykscgl npklekcglg qensdnehlv enevslllee sdlrrsprvk 661 ttplrrptet npvtsnsdee cnetvkekqk lsvpvrkkdk rnssdsaidn pkpnklpksk 721 qsetvdqnsd sdemlailke vsrmshssss dtdineihtn hktlydlktq agkddkgkrk 781 rksstsgsdf dtkkgksaks siiskkkrqt qsessnydse lekeiksmsk igaarttkkr 841 ipntkdfdss edekhskkgm dnqghknlkt sqegssddae rkqeretfss aegtvdkdtt 901 imelrdrlpk kqqasastdg vdklsgkeqs ftslevrkva etkekskhlk tktckkvgdg 961 lsdiaekflk kdqsdetsed dkkqskkgte ekkkpsdfkk kvikmeqqye sssdgteklp 1021 ereeichfpk gikqikngtt dgekkskkir dktskkkdel sdyaekstgk gdscdssedk 1081 kskngaygre kkrckllgks srkrqdcsss dtekysmked gcnssdkrlk rielrerrnl 1141 sskrntkeiq sgssssdaee ssednkkkkq rtsskkkavi vkekkrnslr tstkrkqadi 1201 tsssssdied ddqnsigegs sdeqkikpvt enlvlsshtg fcgssgdeal sksvpvtvdd 1261 ddddndpenr iakkmlleei kanlssdedg ssddepeegk krtgkqneen pgdeeaknqv 1321 nsesdsdsee skkpryrhrl lrhkltvsdg esgeekktkp kehkevkgrn rrkvssedse 1381 dsdfqesgvs eevsesedeq rprtrsakka eleenqrsyk qkkkrrrikv qedsssenks 1441 nseeeeeeke eeeeeeeeee eeeedendds kspgkgrkki rkilkddklr tetqnalkee 1501 eerrkriaer erereklrev ieiedasptk cpittklvld edeetkeplv qvhrnmvikl 1561 kphqvdgvqf mwdcccesvk ktkkspgsgc ilahcmglgk tlqvvsflht vllcdkldfs 1621 talvvcplnt alnwmnefek wqeglkddek levselatvk rpgersymlq rwqedggvmi 1681 igyemyrnla qgrnvksrkl keifnkalvd pgpdfvvcde ghilkneasa vskamnsirs 1741 rrriiltgtp lqnnlieyhc mvnfikenll gsikefrnrf inpiqngqca dstmvdvrvm 1801 kkrahilyem lagcvqrkdy taltkflppk heyvlavrmt siqcklyqyy ldhltgvgnn 1861 seggrgkaga klfqdfqmls riwthpwclq ldyiskenkg yfdedsmdef iasdsdetsm 1921 slssddytkk kkkgkkgkkd ssssgsgsdn dvevikvwns rsrgggegnv detgnnpsvs 1981 lkleeskats ssnpsspapd wykdfvtdad aevlehsgkm vllfeilrma eeigdkvlvf 2041 sqslisldli edflelasre ktedkdkpli ykgegkwlrn idyyrldgst taqsrkkwae 2101 efndetnvrg rlfiistkag slginlvaan rviifdaswn psydigsifr vyrfgqtkpv 2161 yvyrflaqgt medkiydrqv tkqslsfrvv dqqqverhft mneltelytf epdllddpns 2221 ekkkkrdtpm lpkdtilael lqihkehivg yhehdslldh keeeelteee rkaawaeyea 2281 ekkgltmrfn iptgtnlppv sfnsqtpyip fnlgalsams nqqledlinq grekvveatn 2341 svtavriqpl ediisavwke nmnlseaqvg alalsrqasq eldvkrreai yndvltkqqm 2401 liscvqrilm nrrlqqqynq qqqqqmtyqg atlghlmmpk ppnlimnpsn yqqidmrgmy 2461 qpvaggmqpp plqrapppmr sknpgpsqgk sm Homo sapiens alpha thalassemia/mental retardation syndrome X-linked (ATRX), transcript variant 1, mRNA (SEQ ID NO: 6) 1 aattctcctg cctgagcctc ggcccaacaa aatggcggcg gcagcggtgt cgctttgttt 61 ccgcggctcc tgcggcggtg gcagtggtag cggcctttga gctgtgggga ggttccagca 121 gcagctacag tgacgactaa gactccagtg catttctatc gtaaccgggc gcgggggagc 181 gcagatcggc gcccagcaat cacagaagcc gacaaggcgt tcaagcgaaa acatgaccgc 241 tgagcccatg agtgaaagca agttgaatac attggtgcag aagcttcatg acttccttgc 301 acactcatca gaagaatctg aagaaacaag ttctcctcca cgacttgcaa tgaatcaaaa 361 cacagataaa atcagtggtt ctggaagtaa ctctgatatg atggaaaaca gcaaggaaga 421 gggaactagc tcttcagaaa aatccaagtc ttcaggatcg tcacgatcaa agaggaaacc 481 ttcaattgta acaaagtatg tagaatcaga tgatgaaaaa cctttggatg atgaaactgt 541 aaatgaagat gcgtctaatg aaaattcaga aaatgatatt actatgcaga gcttgccaaa 601 aggtacagtg attgtacagc cagagccagt gctgaatgaa gacaaagatg attttaaagg 661 gcctgaattt agaagcagaa gtaaaatgaa aactgaaaat ctcaaaaaac gcggagaaga 721 tgggcttcat gggattgtga gctgcactgc ttgtggacaa caggtcaatc attttcaaaa 781 agattccatt tatagacacc cttcattgca agttcttatt tgtaagaatt gctttaagta 841 ttacatgagt gatgatatta gccgtgactc agatggaatg gatgaacaat gtaggtggtg 901 tgcggaaggt ggaaacttga tttgttgtga cttttgccat aatgctttct gcaagaaatg 961 cattctacgc aaccttggtc gaaaggagtt gtccacaata atggatgaaa acaaccaatg 1021 gtattgctac atttgtcacc cagagccttt gttggacttg gtcactgcat gtaacagcgt 1081 atttgagaat ttagaacagt tgttgcagca aaataagaag aagataaaag ttgacagtga 1141 aaagagtaat aaagtatatg aacatacatc cagattttct ccaaagaaga ctagttcaaa 1201 ttgtaatgga gaagaaaaga aattagatga ttcctgttct ggctctgtaa cctactctta 1261 ttccgcacta attgtgccca aagagatgat taagaaggca aaaaaactga ttgagaccac 1321 agccaacatg aactccagtt atgttaaatt tttaaagcag gcaacagata attcagaaat 1381 cagttctgct acaaaattac gtcagcttaa ggcttttaag tctgtgttgg ctgatattaa 1441 gaaggctcat cttgcattgg aagaagactt aaattccgag tttcgagcga tggatgctgt 1501 aaacaaagag aaaaatacca aagagcataa agtcatagat gctaagtttg aaacaaaagc 1561 acgaaaagga gaaaaacctt gtgctttgga aaagaaggat atttcaaagt cagaagctaa 1621 actttcaaga aaacaggtag atagtgagca catgcatcag aatgttccaa cagaggaaca 1681 aagaacaaat aaaagtaccg gtggtgaaca taagaaatct gatagaaaag aagaacctca 1741 atatgaacct gccaacactt ctgaagattt agacatggat attgtgtctg ttccttcctc 1801 agttccagaa gacatttttg agaatcttga gactgctatg gaagttcaga gttcagttga 1861 tcatcaaggg gatggcagca gtggaactga acaagaagtg gagagttcat ctgtaaaatt 1921 aaatatttct tcaaaagaca acagaggagg tattaaatca aaaactacag ctaaagtaac 1981 aaaagaatta tatgttaaac tcactcctgt ttccctttct aattccccaa ttaaaggtgc 2041 tgattgtcag gaagttccac aagataaaga tggctataaa agttgtggtc tgaaccccaa 2101 gttagagaaa tgtggacttg gacaggaaaa cagtgataat gagcatttgg ttgaaaatga 2161 agtttcatta cttttagagg aatctgatct tcgaagatcc ccacgtgtaa agactacacc 2221 cttgaggcga ccgacagaaa ctaaccctgt aacatctaat tcagatgaag aatgtaatga 2281 aacagttaag gagaaacaaa aactatcagt tccagtgaga aaaaaggata agcgtaattc 2341 ttctgacagt gctatagata atcctaagcc taataaattg ccaaaatcta agcaatcaga 2401 gactgtggat caaaattcag attctgatga aatgctagca atcctcaaag aggtgagcag 2461 gatgagtcac agttcttctt cagatactga tattaatgaa attcatacaa accataagac 2521 tttgtatgat ttaaagactc aggcggggaa agatgataaa ggaaaaagga aacgaaaaag 2581 ttctacatct ggctcagatt ttgatactaa aaagggcaaa tcagctaaga gctctataat 2641 ttctaaaaag aaacgacaaa cccagtctga gtcttctaat tatgactcag aattagaaaa 2701 agagataaag agcatgagta aaattggtgc tgccagaacc accaaaaaaa gaattccaaa 2761 tacaaaagat tttgactctt ctgaagatga gaaacacagc aaaaaaggaa tggataatca 2821 agggcacaaa aatttgaaga cctcacaaga aggatcatct gatgatgctg aaagaaaaca 2881 agagagagag actttctctt cagcagaagg cacagttgat aaagacacga ccatcatgga 2941 attaagagat cgacttccta agaagcagca agcaagtgct tccactgatg gtgtcgataa 3001 gctttctggg aaagagcaga gttttacttc tttggaagtt agaaaagttg ctgaaactaa 3061 agaaaagagc aagcatctca aaaccaaaac atgtaaaaaa gtacaggatg gcttatctga 3121 tattgcagag aaattcctaa agaaagacca gagcgatgaa acttctgaag atgataaaaa 3181 gcagagcaaa aagggaactg aagaaaaaaa gaaaccttca gactttaaga aaaaagtaat 3241 taaaatggaa caacagtatg aatcttcatc tgatggcact gaaaagttac ctgagcgaga 3301 agaaatttgt cattttccta agggcataaa acaaattaag aatggaacaa ctgatggaga 3361 aaagaaaagt aaaaaaataa gagataaaac ttctaaaaag aaggatgaat tatctgatta 3421 tgctgagaag tcaacaggga aaggagatag ttgtgactct tcagaggata aaaagagtaa 3481 gaatggagca tatggtagag agaagaaaag gtgcaagttg cttggaaaga gttcaaggaa 3541 gagacaagat tgttcatcat ctgatactga gaaatattcc atgaaagaag atggttgtaa 3601 ctcttctgat aagagactga aaagaataga attgagggaa agaagaaatt taagttcaaa 3661 gagaaatact aaggaaatac aaagtggctc atcatcatct gatgctgagg aaagttctga 3721 agataataaa aagaagaagc aaagaacttc atctaaaaag aaggcagtca ttgtcaagga 3781 gaaaaagaga aactccctaa gaacaagcac taaaaggaag caagctgaca ttacatcctc 3841 atcttcttct gatatagaag atgatgatca gaattctata ggtgagggaa gcagcgatga 3901 acagaaaatt aagcctgtga ctgaaaattt agtgctgtct tcacatactg gattttgcca 3961 atcttcagga gatgaagcct tatctaaatc agtgcctgtc acagtggatg atgatgatga 4021 cgacaatgat cctgagaata gaattgccaa gaagatgctt ttagaagaaa ttaaagccaa 4081 tctttcctct gatgaggatg gatcttcaga tgatgagcca gaagaaggga aaaaaagaac 4141 tggaaaacaa aatgaagaaa acccaggaga tgaggaagca aaaaatcaag tcaattctga 4201 atcagattca gattctgaag aatctaagaa gccaagatac agacataggc ttttgcggca 4261 caaattgact gtgagtgacg gagaatctgg agaagaaaaa aagacaaagc ctaaagagca 4321 taaagaagtc aaaggcagaa acagaagaaa ggtgagcagt gaagattcag aagattctga 4381 ttttcaggaa tcaggagtta gtgaagaagt tagtgaatcc gaagatgaac agcggcccag 4441 aacaaggtct gcaaagaaag cagagttgga agaaaatcag cggagctata aacagaaaaa 4501 gaaaaggcga cgtattaagg ttcaagaaga ttcatccagt gaaaacaaga gtaattctga 4561 ggaagaagag gaggaaaaag aagaggagga ggaagaggag gaggaggagg aagaggagga 4621 ggaagatgaa aatgatgatt ccaagtctcc tggaaaaggc agaaagaaaa ttcggaagat 4681 tcttaaagat gataaactga gaacagaaac acaaaatgct cttaaggaag aggaagagag
4741 acgaaaacgt attgctgaga gggagcgtga gcgagaaaaa ttgagagagg tgatagaaat 4801 tgaagatgct tcacccacca agtgtccaat aacaaccaag ttggttttag atgaagatga 4861 agaaaccaaa gaacctttag tgcaggttca tagaaatatg gttatcaaat tgaaacccca 4921 tcaagtagat ggtgttcagt ttatgtggga ttgctgctgt gagtctgtga aaaaaacaaa 4981 gaaatctcca ggttcaggat gcattcttgc ccactgtatg ggccttggta agactttaca 5041 ggtggtaagt tttcttcata cagttctttt gtgtgacaaa ctggatttca gcacggcgtt 5101 agtggtttgt cctcttaata ctgctttgaa ttggatgaat gaatttgaga agtggcaaga 5161 gggattaaaa gatgatgaga agcttgaggt ttctgaatta gcaactgtga aacgtcctca 5221 ggagagaagc tacatgctgc agaggtggca agaagatggt ggtgttatga tcataggcta 5281 tgagatgtat agaaatcttg ctcaaggaag gaatgtgaag agtcggaaac ttaaagaaat 5341 atttaacaaa gctttggttg atccaggccc tgattttgtt gtttgtgatg aaggccatat 5401 tctaaaaaat gaagcatctg ctgtttctaa agctatgaat tctatacgat caaggaggag 5461 gattatttta acaggaacac cacttcaaaa taacctaatt gagtatcatt gtatggttaa 5521 ttttatcaag gaaaatttac ttggatccat taaggagttc aggaatagat ttataaatcc 5581 aattcaaaat ggtcagtgtg cagattctac catggtagat gtcagagtga tgaaaaaacg 5641 tgctcacatt ctctatgaga tgttagctgg atgtgttcag aggaaagatt atacagcatt 5701 aacaaaattc ttgcctccaa aacacgaata tgtgttagct gtgagaatga cttctattca 5761 gtgcaagctc tatcagtact acttagatca cttaacaggt gtgggcaata atagtgaagg 5821 tggaagagga aaggcaggtg caaagctttt ccaagatttt cagatgttaa gtagaatatg 5881 gactcatcct tggtgtttgc agctagacta cattagcaaa gaaaataagg gttattttga 5941 tgaagacagt atggatgaat ttatagcctc agattctgat gaaacctcca tgagtttaag 6001 ctccgatgat tatacaaaaa agaagaaaaa agggaaaaag gggaaaaaag atagtagctc 6061 aagtggaagt ggcagtgaca atgatgttga agtgattaag gtctggaatt caagatctcg 6121 gggaggtggt gaaggaaatg tggatgaaac aggaaacaat ccttctgttt ctttaaaact 6181 ggaagaaagt aaagctactt cttcttctaa tccaagcagc ccagctccag actggtacaa 6241 agattttgtt acagatgctg atgctgaggt tttagagcat tctgggaaaa tggtacttct 6301 ctttgaaatt cttcgaatgg cagaggaaat tggggataaa gtccttgttt tcagccagtc 6361 cctcatatct ctggacttga ttgaagattt tcttgaatta gctagtaggg agaagacaga 6421 agataaagat aaacccctta tttataaagg tgaggggaag tggcttcgaa acattgacta 6481 ttaccgttta gatggttcca ctactgcaca gtcaaggaag aagtgggctg aagaatttaa 6541 tgatgaaact aatgtgagag gacgattatt tatcatttct actaaagcag gatctctagg 6601 aattaatctg gtagctgcta atcgagtaat tatattcgac gcttcttgga atccatctta 6661 tgacatccag agtatattca gagtttatcg ctttggacaa actaagcctg tttatgtata 6721 taggttctta gctcagggaa ccatggaaga taagatttat gatcggcaag taactaagca 6781 gtcactgtct tttcgagttg ttgatcagca gcaggtggag cgtcatttta ctatgaatga 6841 gcttactgaa ctttatactt ttgagccaga cttattagat gaccctaatt cagaaaagaa 6901 gaagaagagg gatactccca tgctgccaaa ggataccata cttgcagagc tccttcagat 6961 acataaagaa cacattgtag gataccatga acatgattct cttttggacc acaaagaaga 7021 agaagagttg actgaagaag aaagaaaagc agcttgggct gagtatgaag cagagaagaa 7081 gggactgacc atgcgtttca acataccaac tgggaccaat ttaccccctg tcagtttcaa 7141 ctctcaaact ccttatattc ctttcaattt gggagccctg tcagcaatga gtaatcaaca 7201 gctggaggac ctcattaatc aaggaagaga aaaagttgta gaagcaacaa acagtgtgac 7261 agcagtgagg attcaacctc ttgaggatat aatttcagct gtatggaagg agaacatgaa 7321 tctctcagag gcccaagtac aggcgttagc attaagtaga caagccagcc aggagcttga 7381 tgttaaacga agagaagcaa tctacaatga tgtattgaca aaacaacaga tgttaatcag 7441 ctgtgttcag cgaatactta tgaacagaag gctccagcag cagtacaatc agcagcaaca 7501 gcaacaaatg acttatcaac aagcaacact gggtcacctc atgatgccaa agcccccaaa 7561 tttgatcatg aatccttcta actaccagca gattgatatg agaggaatgt atcagccagt 7621 ggctggtggt atgcagccac caccattaca gcgtgcacca cccccaatga gaagcaaaaa 7681 tccaggacct tcccaaggga aatcaatgtg attttgcact aaaagcttaa tggattgtta 7741 aaatcataga aagatctttt atttttttag gaatcaatga cttaacagaa ctcaactgta 7801 taaatagttt ggtcccctta aatgccaatc ttccatatta gttttacttt tttttttttt 7861 aaatagggca taccatttct tcctgacatt tgtcagtgat gttgcctaga atcttcttac 7921 acacgctgag tacagaagat atttcaaatt gttttcagtg aaaacaagtc cttccataat 7981 agtaacaact ccacagattt cctctctaaa tttttatgcc tgcttttagc aaccataaaa 8041 ttgtcataaa attaataaat ttaggaaaga ataaagattt atatattcat tctttacata 8101 taaaaacaca cagctgagtt cttagagttg attcctcaag ttatgaaata cttttgtact 8161 taatccattt cttgattaaa gtgattgaaa tggttttaat gttcttttga ctgaagtctg 8221 aaactgggct cctgctttat tgtctctgtg actgaaagtt agaaactgag ggttatcttt 8281 gacacagaat tgtgtgcaat attcttaaat actactgctc taaaagttgg agaagtcttg 8341 cagttatctt agcattgtat aaacagcctt aagtatagcc taagaagaga attccttttt 8401 cttctttagt ccttctgcca ttttttattt tcagttatat gtgctgaaat aattactggt 8461 aaaatttcag ggttgtggat tatcttccac acatgaattt tctctctcct ggcacgaata 8521 taaagcacat ctcttaactg catggtgcca gtgctaatgc ttcatcctgt tgctggcagt 8581 gggatgtgga cttagaaaat caagttctag cattttagta ggttaacact gaagttgtgg 8641 ttgttaggtt cacaccctgt tttataaaca acatcaaaat ggcagaacca ttgctgactt 8701 taggttcaca tgaggaatgt acttttaaca attcccagta ctatcagtat tgtgaaataa 8761 ttcctctgaa agataagaat cactggcttc tatgcgcttc ttttctctca tcatcatgtt 8821 cttttacccc agtttcctta cattttttta aattgtttca gagtttgttt tttttttagt 8881 ttagattgtg aggcaattat taaatcaaaa ttaattcatc caatacccct ttactagaag 8941 ttttactaga aaatgtatta cattttattt tttcttaatc cagttctgca aaaatgacct 9001 ataaatttat tcatgtacaa ttttggttac ttgaattgtt aaagaaaaca ttgtttttga 9061 ctatgggagt caactcaaca tggcagaacc atttttgaga tgatgataca acaggtagtg 9121 aaacagctta agaattccaa aaaaaaaaaa aaaaaaaaaa aaaagaaaac tgggtttggg 9181 ctttgcttta ggtatcactg gattagaatg agtttaacat tagctaaaac tgctttgagt 9241 tgtttggatg attaagagat tgccattttt atcttggaag aactagtggt aaaacatcca 9301 agagcactag gattgtgata cagaatttgt gaggtttggt ggatccacgc ccctctcccc 9361 cactttccca tgatgaaata tcactaataa atcctgtata tttagatatt atgctagcca 9421 tgtaatcaga tttatttaat tgggtggggc aggtgtgtat ttactttaga aaaaatgaaa 9481 aagacaagat ttatgagaaa tatttgaagg cagtacactc tggccaactg ttaccagttg 9541 gtatttctac aagttcagaa tattttaaac ctgatttact agacctggga attttcaaca 9601 tggtctaatt atttactcaa agacatagat gtgaaaattt taggcaacct tctaaatctt 9661 tttcaccatg gatgaaacta taacttaaag aataatactt agaagggtta attggaaatc 9721 agagtttgaa ataaaacttg gaccactttg tatacactct tctcacttga cattttagct 9781 atataatatg tactttgagt ataacatcaa gctttaacaa atatttaaag acaaaaaaat 9841 cacgtcagta aaatactaaa aggctcattt ttatatttgt tttagatgtt ttaaatagtt 9901 gcaatggatt aaaaatgatg atttaaaatg ttgcttgtaa tacagttttg cctgctaaat 9961 tctccacatt ttgtaacctg ttttatttct ttgggtgtaa agcgtttttg cttagtattg 10021 tgatattgta tatgttttgt cccagttgta tagtaatgtt tcagtccatc atccagcttt 10081 ggctgctgaa atcatacagc tgtgaagact tgcctttgtt tctgttagac tgcttttcag 10141 ttctgtattg agtatcttaa gtactgtaga aaagatgtca cttcttcctt taaggctgtt 10201 ttgtaatata tataaggact ggaattgtgt ttttaaagaa aagcattcaa gtatgacaat 10261 atactatctg tgttttcacc attcaaagtg ctgtttagta gttgaaactt aaactattta 10321 atgtcattta ataaagtgac caaaatgtgt tgtgctcttt attgtatttt cacagctttg 10381 aaaatctgtg cacatactgt ttcatagaaa atgtatagct tttgttgtcc tatataatgg 10441 tggttctttt gcacatttag ttatttaata ttgagaggtc acgaagtttg gttattgaat 10501 ctgttatata ctaaattctg taaagggaga tctctcatct caaaaagaat ttacatacca 10561 ggaagtccat gtgtgtttgt gttagttttg gatgtctttg tgtaatccag ccccatttcc 10621 tgtttcccaa cagctgtaac actcatttta agtcaagcag ggctaccaac ccacacttga 10681 tagaaaagct gcttaccatt cagaagcttc cttattacct ggcctccaaa tgagctgaat 10741 attttgtagc cttcccttag ctatgttcat tttccctcca ttatcataaa atcagatcga 10801 tatttatgtg ccccaaacaa aactttaaga gcagttacat tctgtcccag tagcccttgt 10861 ttcctttgag agtagcatgt tgtgaggcta tagagactta ttctaccagt aaaacaggtc 10921 aatcctttta catgtttatt atactaaaaa ttatgttcag ggtatttact actttatttc 10981 accagactca gtctcaagtg acttggctat ctccaaatca gatctaccct tagagaataa 11041 acatttttct accgttattt tttttcaagt ctataatctg agccagtccc aaaggagtga 11101 tcaagtttca gaaatgcttt catcttcaca acattttata tatactatta tatggggtga 11161 ataaagtttt aaatccgaaa tataaaaaaa aaaaaaaaaa aa Homo sapiens alpha thalassemia/mental retardation syndrome X-linked (ATRX) isoform 2 (SEQ ID NO: 7) 1 mtaepmsesk lntlvqklhd flahsseese etsspprlam nqntdkisgs gsnsdmmens 61 keegtsssek skssgssrsk rkpsivtkyv esddekpldd etvnedasne nsenditmqs 121 lpkedglhgi vsctacgqqv nhfqkdsiyr hpslqvlick ncfkyymsdd isrdsdgmde 181 qcrwcaeggn liccdfchna fokkcilrnl grkelstimd ennqwycyic hpeplldlvt 241 acnsvfenle qllqqnkkki kvdseksnkv yehtsrfspk ktssncngee kklddscsgs 301 vtysysaliv pkemikkakk liettanmns syvkflkqat dnseissatk lrqlkafksv 361 ladikkahla leedlnsefr amdavnkekn tkehkvidak fetkarkgek pcalekkdis 421 kseaklsrkg vdsehmhqnv pteeqrtnks tggehkksdr keepqyepan tsedldmdiv 481 svpssvpedi fenletamev qssvdhqgdg ssgteqeves ssvklnissk dnrggikskt 541 takvtkelyv kltpvslsns pikgadcqev pqdkdgyksc glnpklekcg lgqensdneh 601 lvenevslll eesdlrrspr vkttplrrpt etnpvtsnsd eecnetvkek qklsvpvrkk 661 dkrnssdsai dnpkpnklpk skqsetvdqn sdsdemlail kevsrmshss ssdtdineih 721 tnhktlydlk tqagkddkgk rkrksstsgs dfdtkkgksa kssiiskkkr qtqsessnyd 781 selekeiksm skigaarttk kripntkdfd ssedekhskk gmdnqghknl ktsqegssdd 841 aerkqeretf ssaegtvdkd ttimelrdrl pkkqqasast dgvdklsgke qsftslevrk 901 vaetkekskh lktktckkvq dglsdiaekf lkkdqsdets eddkkqskkg teekkkpsdf
961 kkkvikmeqq yesssdgtek lpereeichf pkgikqikng ttdgekkskk irdktskkkd 1021 elsdyaekst gkgdscdsse dkkskngayg rekkrckllg kssrkrqdcs ssdtekysmk 1081 edgcnssdkr lkrielrerr nlsskrntke iqsgssssda eessednkkk kqrtsskkka 1141 vivkekkrns lrtstkrkqa ditsssssdi edddqnsige gssdeqkikp vtenlvlssh 1201 tgfcqssgde alsksvpvtv ddddddndpe nriakkmlle eikanlssde dgssddepee 1261 gkkrtgkqne enpgdeeakn qvnsesdsds eeskkpryrh rllrhkltvs dgesgeekkt 1321 kpkehkevkg rnrrkvssed sedsdfqesg vseevsesed eqrprtrsak kaeleenqrs 1381 ykqkkkrrri kvqedsssen ksnseeeeee keeeeeeeee eeeeeedend dskspgkgrk 1441 kirkilkddk lrtetqnalk eeeerrkria ererereklr evieiedasp tkcpittklv 1501 ldedeetkep lvqvhrnmvi klkphqvdgv qfmwdccces vkktkkspgs gcilahcmgl 1561 gktlqvvsfl htvllcdkld fstalvvcpl ntalnwmnef ekwqeglkdd eklevselat 1621 vkrpqersym lqrwqedggv miigyemyrn laqgrnvksr klkeifnkal vdpgpdfvvc 1681 deghilknea savskamnsi rsrrriiltg tplqnnliey hcmvnfiken llgsikefrn 1741 rfinpiqngq cadstmvdvr vmkkrahily emlagcvqrk dytaltkflp pkheyvlavr 1801 mtsiqcklyq yyldhltgvg nnseggrgka gaklfqdfqm lsriwthpwc lqldyisken 1861 kgyfdedsmd efiasdsdet smslssddyt kkkkkgkkgk kdssssgsgs dndvevikvw 1921 nsrsrgggeg nvdetgnnps vslkleeska tsssnpsspa pdwykdfvtd adaevlehsg 1981 kmvllfeilr maeeigdkvl vfsqslisld liedflelas rektedkdkp liykgegkwl 2041 rnidyyrldg sttaqsrkkw aeefndetnv rgrlfiistk agslginlva anrviifdas 2101 wnpsydiqsi frvyrfgqtk pvyvyrflaq gtmedkiydr qvtkqslsfr vvdqqqverh 2161 ftmneltely tfepdllddp nsekkkkrdt pmlpkdtila ellqihkehi vgyhehdsll 2221 dhkeeeelte eerkaawaey eaekkgltmr fniptgtnlp pvsfnsqtpy ipfnlgalsa 2281 msnqqledli nqgrekvvea tnsvtavriq plediisavw kenmnlseaq vqalalsrqa 2341 sqeldvkrre aiyndvltkq qmliscvqri lmnrrlqqqy nqqqqqqmty qqatlghlmm 2401 pkppnlimnp snyqqidmrg myqpvaggmq ppplqrappp mrsknpgpsq gksm Homo sapiens alpha thalassemia/mental retardation syndrome X-linked (ATRX), transcript variant 2, mRNA (SEQ ID NO: 8) 1 aattctcctg cctgagcctc ggcccaacaa aatggcggcg gcagcggtgt cgctttgttt 61 ccgcggctcc tgcggcggtg gcagtggtag cggcctttga gctgtgggga ggttccagca 121 gcagctacag tgacgactaa gactccagtg catttctatc gtaaccgggc gcgggggagc 181 gcagatcggc gcccagcaat cacagaagcc gacaaggcgt tcaagcgaaa acatgaccgc 241 tgagcccatg agtgaaagca agttgaatac attggtgcag aagcttcatg acttccttgc 301 acactcatca gaagaatctg aagaaacaag ttctcctcca cgacttgcaa tgaatcaaaa 361 cacagataaa atcagtggtt ctggaagtaa ctctgatatg atggaaaaca gcaaggaaga 421 gggaactagc tcttcagaaa aatccaagtc ttcaggatcg tcacgatcaa agaggaaacc 481 ttcaattgta acaaagtatg tagaatcaga tgatgaaaaa cctttggatg atgaaactgt 541 aaatgaagat gcgtctaatg aaaattcaga aaatgatatt actatgcaga gcttgccaaa 601 agaagatggg cttcatggga ttgtgagctg cactgcttgt ggacaacagg tcaatcattt 661 tcaaaaagat tccatttata gacacccttc attgcaagtt cttatttgta agaattgctt 721 taagtattac atgagtgatg atattagccg tgactcagat ggaatggatg aacaatgtag 781 gtggtgtgcg gaaggtggaa acttgatttg ttgtgacttt tgccataatg ctttctgcaa 841 gaaatgcatt ctacgcaacc ttggtcgaaa ggagttgtcc acaataatgg atgaaaacaa 901 ccaatggtat tgctacattt gtcacccaga gcctttgttg gacttggtca ctgcatgtaa 961 cagcgtattt gagaatttag aacagttgtt gcagcaaaat aagaagaaga taaaagttga 1021 cagtgaaaag agtaataaag tatatgaaca tacatccaga ttttctccaa agaagactag 1081 ttcaaattgt aatggagaag aaaagaaatt agatgattcc tgttctggct ctgtaaccta 1141 ctcttattcc gcactaattg tgcccaaaga gatgattaag aaggcaaaaa aactgattga 1201 gaccacagcc aacatgaact ccagttatgt taaattttta aagcaggcaa cagataattc 1261 agaaatcagt tctgctacaa aattacgtca gcttaaggct tttaagtctg tgttggctga 1321 tattaagaag gctcatcttg cattggaaga agacttaaat tccgagtttc gagcgatgga 1381 tgctgtaaac aaagagaaaa ataccaaaga gcataaagtc atagatgcta agtttgaaac 1441 aaaagcacga aaaggagaaa aaccttgtgc tttggaaaag aaggatattt caaagtcaga 1501 agctaaactt tcaagaaaac aggtagatag tgagcacatg catcagaatg ttccaacaga 1561 ggaacaaaga acaaataaaa gtaccggtgg tgaacataag aaatctgata gaaaagaaga 1621 acctcaatat gaacctgcca acacttctga agatttagac atggatattg tgtctgttcc 1681 ttcctcagtt ccagaagaca tttttgagaa tcttgagact gctatggaag ttcagagttc 1741 agttgatcat caaggggatg gcagcagtgg aactgaacaa gaagtggaga gttcatctgt 1801 aaaattaaat atttcttcaa aagacaacag aggaggtatt aaatcaaaaa ctacagctaa 1861 agtaacaaaa gaattatatg ttaaactcac tcctgtttcc ctttctaatt ccccaattaa 1921 aggtgctgat tgtcaggaag ttccacaaga taaagatggc tataaaagtt gtggtctgaa 1981 ccccaagtta gagaaatgtg gacttggaca ggaaaacagt gataatgagc atttggttga 2041 aaatgaagtt tcattacttt tagaggaatc tgatcttcga agatccccac gtgtaaagac 2101 tacacccttg aggcgaccga cagaaactaa ccctgtaaca tctaattcag atgaagaatg 2161 taatgaaaca gttaaggaga aacaaaaact atcagttcca gtgagaaaaa aggataagcg 2221 taattcttct gacagtgcta tagataatcc taagcctaat aaattgccaa aatctaagca 2281 atcagagact gtggatcaaa attcagattc tgatgaaatg ctagcaatcc tcaaagaggt 2341 gagcaggatg agtcacagtt cttcttcaga tactgatatt aatgaaattc atacaaacca 2401 taagactttg tatgatttaa agactcaggc ggggaaagat gataaaggaa aaaggaaacg 2461 aaaaagttct acatctggct cagattttga tactaaaaag ggcaaatcag ctaagagctc 2521 tataatttct aaaaagaaac gacaaaccca gtctgagtct tctaattatg actcagaatt 2581 agaaaaagag ataaagagca tgagtaaaat tggtgctgcc agaaccacca aaaaaagaat 2641 tccaaataca aaagattttg actcttctga agatgagaaa cacagcaaaa aaggaatgga 2701 taatcaaggg cacaaaaatt tgaagacctc acaagaagga tcatctgatg atgctgaaag 2761 aaaacaagag agagagactt tctcttcagc agaaggcaca gttgataaag acacgaccat 2821 catggaatta agagatcgac ttcctaagaa gcagcaagca agtgcttcca ctgatggtgt 2881 cgataagctt tctgggaaag agcagagttt tacttctttg gaagttagaa aagttgctga 2941 aactaaagaa aagagcaagc atctcaaaac caaaacatgt aaaaaagtac aggatggctt 3001 atctgatatt gcagagaaat tcctaaagaa agaccagagc gatgaaactt ctgaagatga 3061 taaaaagcag agcaaaaagg gaactgaaga aaaaaagaaa ccttcagact ttaagaaaaa 3121 agtaattaaa atggaacaac agtatgaatc ttcatctgat ggcactgaaa agttacctga 3181 gcgagaagaa atttgtcatt ttcctaaggg cataaaacaa attaagaatg gaacaactga 3241 tggagaaaag aaaagtaaaa aaataagaga taaaacttct aaaaagaagg atgaattatc 3301 tgattatgct gagaagtcaa cagggaaagg agatagttgt gactcttcag aggataaaaa 3361 gagtaagaat ggagcatatg gtagagagaa gaaaaggtgc aagttgcttg gaaagagttc 3421 aaggaagaga caagattgtt catcatctga tactgagaaa tattccatga aagaagatgg 3481 ttgtaactct tctgataaga gactgaaaag aatagaattg agggaaagaa gaaatttaag 3541 ttcaaagaga aatactaagg aaatacaaag tggctcatca tcatctgatg ctgaggaaag 3601 ttctgaagat aataaaaaga agaagcaaag aacttcatct aaaaagaagg cagtcattgt 3661 caaggagaaa aagagaaact ccctaagaac aagcactaaa aggaagcaag ctgacattac 3721 atcctcatct tcttctgata tagaagatga tgatcagaat tctataggtg agggaagcag 3781 cgatgaacag aaaattaagc ctgtgactga aaatttagtg ctgtcttcac atactggatt 3841 ttgccaatct tcaggagatg aagccttatc taaatcagtg cctgtcacag tggatgatga 3901 tgatgacgac aatgatcctg agaatagaat tgccaagaag atgcttttag aagaaattaa 3961 agccaatctt tcctctgatg aggatggatc ttcagatgat gagccagaag aagggaaaaa 4021 aagaactgga aaacaaaatg aagaaaaccc aggagatgag gaagcaaaaa atcaagtcaa 4081 ttctgaatca gattcagatt ctgaagaatc taagaagcca agatacagac ataggctttt 4141 gcggcacaaa ttgactgtga gtgacggaga atctggagaa gaaaaaaaga caaagcctaa 4201 agagcataaa gaagtcaaag gcagaaacag aagaaaggtg agcagtgaag attcagaaga 4261 ttctgatttt caggaatcag gagttagtga agaagttagt gaatccgaag atgaacagcg 4321 gcccagaaca aggtctgcaa agaaagcaga gttggaagaa aatcagcgga gctataaaca 4381 gaaaaagaaa aggcgacgta ttaaggttca agaagattca tccagtgaaa acaagagtaa 4441 ttctgaggaa gaagaggagg aaaaagaaga ggaggaggaa gaggaggagg aggaggaaga 4501 ggaggaggaa gatgaaaatg atgattccaa gtctcctgga aaaggcagaa agaaaattcg 4561 gaagattctt aaagatgata aactgagaac agaaacacaa aatgctctta aggaagagga 4621 agagagacga aaacgtattg ctgagaggga gcgtgagcga gaaaaattga gagaggtgat 4681 agaaattgaa gatgcttcac ccaccaagtg tccaataaca accaagttgg ttttagatga 4741 agatgaagaa accaaagaac ctttagtgca ggttcataga aatatggtta tcaaattgaa 4801 accccatcaa gtagatggtg ttcagtttat gtgggattgc tgctgtgagt ctgtgaaaaa 4861 aacaaagaaa tctccaggtt caggatgcat tcttgcccac tgtatgggcc ttggtaagac 4921 tttacaggtg gtaagttttc ttcatacagt tcttttgtgt gacaaactgg atttcagcac 4981 ggcgttagtg gtttgtcctc ttaatactgc tttgaattgg atgaatgaat ttgagaagtg 5041 gcaagaggga ttaaaagatg atgagaagct tgaggtttct gaattagcaa ctgtgaaacg 5101 tcctcaggag agaagctaca tgctgcagag gtggcaagaa gatggtggtg ttatgatcat 5161 aggctatgag atgtatagaa atcttgctca aggaaggaat gtgaagagtc ggaaacttaa 5221 agaaatattt aacaaagctt tggttgatcc aggccctgat tttgttgttt gtgatgaagg 5281 ccatattcta aaaaatgaag catctgctgt ttctaaagct atgaattcta tacgatcaag 5341 gaggaggatt attttaacag gaacaccact tcaaaataac ctaattgagt atcattgtat 5401 ggttaatttt atcaaggaaa atttacttgg atccattaag gagttcagga atagatttat 5461 aaatccaatt caaaatggtc agtgtgcaga ttctaccatg gtagatgtca gagtgatgaa 5521 aaaacgtgct cacattctct atgagatgtt agctggatgt gttcagagga aagattatac 5581 agcattaaca aaattcttgc ctccaaaaca cgaatatgtg ttagctgtga gaatgacttc 5641 tattcagtgc aagctctatc agtactactt agatcactta acaggtgtgg gcaataatag 5701 tgaaggtgga agaggaaagg caggtgcaaa gcttttccaa gattttcaga tgttaagtag 5761 aatatggact catccttggt gtttgcagct agactacatt agcaaagaaa ataagggtta 5821 ttttgatgaa gacagtatgg atgaatttat agcctcagat tctgatgaaa cctccatgag
5881 tttaagctcc gatgattata caaaaaagaa gaaaaaaggg aaaaagggga aaaaagatag 5941 tagctcaagt ggaagtggca gtgacaatga tgttgaagtg attaaggtct ggaattcaag 6001 atctcgggga ggtggtgaag gaaatgtgga tgaaacagga aacaatcctt ctgtttcttt 6061 aaaactggaa gaaagtaaag ctacttcttc ttctaatcca agcagcccag ctccagactg 6121 gtacaaagat tttgttacag atgctgatgc tgaggtttta gagcattctg ggaaaatggt 6181 acttctcttt gaaattcttc gaatggcaga ggaaattggg gataaagtcc ttgttttcag 6241 ccagtccctc atatctctgg acttgattga agattttctt gaattagcta gtagggagaa 6301 gacagaagat aaagataaac cccttattta taaaggtgag gggaagtggc ttcgaaacat 6361 tgactattac cgtttagatg gttccactac tgcacagtca aggaagaagt gggctgaaga 6421 atttaatgat gaaactaatg tgagaggacg attatttatc atttctacta aagcaggatc 6481 tctaggaatt aatctggtag ctgctaatcg agtaattata ttcgacgctt cttggaatcc 6541 atcttatgac atccagagta tattcagagt ttatcgcttt ggacaaacta agcctgttta 6601 tgtatatagg ttcttagctc agggaaccat ggaagataag atttatgatc ggcaagtaac 6661 taagcagtca ctgtcttttc gagttgttga tcagcagcag gtggagcgtc attttactat 6721 gaatgagctt actgaacttt atacttttga gccagactta ttagatgacc ctaattcaga 6781 aaagaagaag aagagggata ctcccatgct gccaaaggat accatacttg cagagctcct 6841 tcagatacat aaagaacaca ttgtaggata ccatgaacat gattctcttt tggaccacaa 6901 agaagaagaa gagttgactg aagaagaaag aaaagcagct tgggctgagt atgaagcaga 6961 gaagaaggga ctgaccatgc gtttcaacat accaactggg accaatttac cccctgtcag 7021 tttcaactct caaactcctt atattccttt caatttggga gccctgtcag caatgagtaa 7081 tcaacagctg gaggacctca ttaatcaagg aagagaaaaa gttgtagaag caacaaacag 7141 tgtgacagca gtgaggattc aacctcttga ggatataatt tcagctgtat ggaaggagaa 7201 catgaatctc tcagaggccc aagtacaggc gttagcatta agtagacaag ccagccagga 7261 gcttgatgtt aaacgaagag aagcaatcta caatgatgta ttgacaaaac aacagatgtt 7321 aatcagctgt gttcagcgaa tacttatgaa cagaaggctc cagcagcagt acaatcagca 7381 gcaacagcaa caaatgactt atcaacaagc aacactgggt cacctcatga tgccaaagcc 7441 cccaaatttg atcatgaatc cttctaacta ccagcagatt gatatgagag gaatgtatca 7501 gccagtggct ggtggtatgc agccaccacc attacagcgt gcaccacccc caatgagaag 7561 caaaaatcca ggaccttccc aagggaaatc aatgtgattt tgcactaaaa gcttaatgga 7621 ttgttaaaat catagaaaga tcttttattt ttttaggaat caatgactta acagaactca 7681 actgtataaa tagtttggtc cccttaaatg ccaatcttcc atattagttt tacttttttt 7741 ttttttaaat agggcatacc atttcttcct gacatttgtc agtgatgttg cctagaatct 7801 tcttacacac gctgagtaca gaagatattt caaattgttt tcagtgaaaa caagtccttc 7861 cataatagta acaactccac agatttcctc tctaaatttt tatgcctgct tttagcaacc 7921 ataaaattgt cataaaatta ataaatttag gaaagaataa agatttatat attcattctt 7981 tacatataaa aacacacagc tgagttctta gagttgattc ctcaagttat gaaatacttt 8041 tgtacttaat ccatttcttg attaaagtga ttgaaatggt tttaatgttc ttttgactga 8101 agtctgaaac tgggctcctg ctttattgtc tctgtgactg aaagttagaa actgagggtt 8161 atctttgaca cagaattgtg tgcaatattc ttaaatacta ctgctctaaa agttggagaa 8221 gtcttgcagt tatcttagca ttgtataaac agccttaagt atagcctaag aagagaattc 8281 ctttttcttc tttagtcctt ctgccatttt ttattttcag ttatatgtgc tgaaataatt 8341 actggtaaaa tttcagggtt gtggattatc ttccacacat gaattttctc tctcctggca 8401 cgaatataaa gcacatctct taactgcatg gtgccagtgc taatgcttca tcctgttgct 8461 ggcagtggga tgtggactta gaaaatcaag ttctagcatt ttagtaggtt aacactgaag 8521 ttgtggttgt taggttcaca ccctgtttta taaacaacat caaaatggca gaaccattgc 8581 tgactttagg ttcacatgag gaatgtactt ttaacaattc ccagtactat cagtattgtg 8641 aaataattcc tctgaaagat aagaatcact ggcttctatg cgcttctttt ctctcatcat 8701 catgttcttt taccccagtt tccttacatt tttttaaatt gtttcagagt ttgttttttt 8761 tttagtttag attgtgaggc aattattaaa tcaaaattaa ttcatccaat acccctttac 8821 tagaagtttt actagaaaat gtattacatt ttattttttc ttaatccagt tctgcaaaaa 8881 tgacctataa atttattcat gtacaatttt ggttacttga attgttaaag aaaacattgt 8941 ttttgactat gggagtcaac tcaacatggc agaaccattt ttgagatgat gatacaacag 9001 gtagtgaaac agcttaagaa ttccaaaaaa aaaaaaaaaa aaaaaaaaaa gaaaactggg 9061 tttgggcttt gctttaggta tcactggatt agaatgagtt taacattagc taaaactgct 9121 ttgagttgtt tggatgatta agagattgcc atttttatct tggaagaact agtggtaaaa 9181 catccaagag cactaggatt gtgatacaga atttgtgagg tttggtggat ccacgcccct 9241 ctcccccact ttcccatgat gaaatatcac taataaatcc tgtatattta gatattatgc 9301 tagccatgta atcagattta tttaattggg tggggcaggt gtgtatttac tttagaaaaa 9361 atgaaaaaga caagatttat gagaaatatt tgaaggcagt acactctggc caactgttac 9421 cagttggtat ttctacaagt tcagaatatt ttaaacctga tttactagac ctgggaattt 9481 tcaacatggt ctaattattt actcaaagac atagatgtga aaattttagg caaccttcta 9541 aatctttttc accatggatg aaactataac ttaaagaata atacttagaa gggttaattg 9601 gaaatcagag tttgaaataa aacttggacc actttgtata cactcttctc acttgacatt 9661 ttagctatat aatatgtact ttgagtataa catcaagctt taacaaatat ttaaagacaa 9721 aaaaatcacg tcagtaaaat actaaaaggc tcatttttat atttgtttta gatgttttaa 9781 atagttgcaa tggattaaaa atgatgattt aaaatgttgc ttgtaataca gttttgcctg 9841 ctaaattctc cacattttgt aacctgtttt atttctttgg gtgtaaagcg tttttgctta 9901 gtattgtgat attgtatatg ttttgtccca gttgtatagt aatgtttcag tccatcatcc 9961 agctttggct gctgaaatca tacagctgtg aagacttgcc tttgtttctg ttagactgct 10021 tttcagttct gtattgagta tcttaagtac tgtagaaaag atgtcacttc ttcctttaag 10081 gctgttttgt aatatatata aggactggaa ttgtgttttt aaagaaaagc attcaagtat 10141 gacaatatac tatctgtgtt ttcaccattc aaagtgctgt ttagtagttg aaacttaaac 10201 tatttaatgt catttaataa agtgaccaaa atgtgttgtg ctctttattg tattttcaca 10261 gctttgaaaa tctgtgcaca tactgtttca tagaaaatgt atagcttttg ttgtcctata 10321 taatggtggt tcttttgcac atttagttat ttaatattga gaggtcacga agtttggtta 10381 ttgaatctgt tatatactaa attctgtaaa gggagatctc tcatctcaaa aagaatttac 10441 ataccaggaa gtccatgtgt gtttgtgtta gttttggatg tctttgtgta atccagcccc 10501 atttcctgtt tcccaacagc tgtaacactc attttaagtc aagcagggct accaacccac 10561 acttgataga aaagctgctt accattcaga agcttcctta ttacctggcc tccaaatgag 10621 ctgaatattt tgtagccttc ccttagctat gttcattttc cctccattat cataaaatca 10681 gatcgatatt tatgtgcccc aaacaaaact ttaagagcag ttacattctg tcccagtagc 10741 ccttgtttcc tttgagagta gcatgttgtg aggctataga gacttattct accagtaaaa 10801 caggtcaatc cttttacatg tttattatac taaaaattat gttcagggta tttactactt 10861 tatttcacca gactcagtct caagtgactt ggctatctcc aaatcagatc tacccttaga 10921 gaataaacat ttttctaccg ttattttttt tcaagtctat aatctgagcc agtcccaaag 10981 gagtgatcaa gtttcagaaa tgctttcatc ttcacaacat tttatatata ctattatatg 11041 gggtgaataa agttttaaat ccgaaatata aaaaaaaaaa aaaaaaaa
[0117] A subject in need thereof may have reduced expression, haploinsufficiency, and/or loss of function of ARID1A. For example, a subject may comprise a mutation selected from the group consisting of a nonsense mutation for the wild type residue cysteine (C) at amino acid position 884 of SEQ ID NO: 11 (C884*), a substitution of lysine (K) for the wild type residue glutamic acid (E) at amino acid position 966 (E966K), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1411 of SEQ ID NO: 11 (Q1411*), a frame shift mutation at the wild type residue phenylalanine (F) at amino acid position 1720 of SEQ ID NO: 11 (F1720fs), a frame shift mutation after the wild type residue glycine (G) at amino acid position 1847 of SEQ ID NO: 11 (G1847fs), a frame shift mutation at the wild type residue cysteine (C) at amino acid position 1874 of SEQ ID NO: 11 (C1874fs), a substitution of glutamic acid (E) for the wild type residue aspartic acid (D) at amino acid position 1957 (D1957E), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1430 of SEQ ID NO: 11 (Q1430*), a frame shift mutation at the wild type residue arginine (R) at amino acid position 1721 of SEQ ID NO: 11 (R1721fs), a substitution of glutamic acid (E) for the wild type residue glycine (G) at amino acid position 1255 (G1255E), a frame shift mutation at the wild type residue glycine (G) at amino acid position 284 of SEQ ID NO: 11 (G284fs), a nonsense mutation for the wild type residue arginine (R) at amino acid position 1722 of SEQ ID NO: 11 (R1722*), a frame shift mutation at the wild type residue methionine (M) at amino acid position 274 of SEQ ID NO: 11 (M274fs), a frame shift mutation at the wild type residue glycine (G) at amino acid position 1847 of SEQ ID NO: 11 (G1847fs), a frame shift mutation at the wild type residue P at amino acid position 559 of SEQ ID NO: 11 (P559fs), a nonsense mutation for the wild type residue arginine (R) at amino acid position 1276 of SEQ ID NO: 11 (R1276*), a frame shift mutation at the wild type residue glutamine (Q) at amino acid position 2176 of SEQ ID NO: 11 (Q2176fs), a frame shift mutation at the wild type residue histidine (H) at amino acid position 203 of SEQ ID NO: 11 (H203fs), a frame shift mutation at the wild type residue alanine (A) at amino acid position 591 of SEQ ID NO: 11 (A591fs), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 1322 of SEQ ID NO: 11 (Q1322*), a nonsense mutation for the wild type residue serine (S) at amino acid position 2264 of SEQ ID NO: 11 (S2264*), a nonsense mutation for the wild type residue glutamine (Q) at amino acid position 586 of SEQ ID NO: 11 (Q586*), a frame shift mutation at the wild type residue glutamine (Q) at amino acid position 548 of SEQ ID NO: 11 (Q548fs), and a frame shift mutation at the wild type residue asparagine (N) at amino acid position 756 of SEQ ID NO: 11 (N756fs). "*" used herein refers to a stop codon. "fs" used herein refers to a frame shift.
TABLE-US-00003 AT-rich interactive domain-containing protein 1A (ARID1A) isoform a [Homo sapiens] (SEQ ID NO: 9) 1 maaqvapaaa sslgnppppp pselkkaeqq qreeaggeaa aaaaaergem kaaagqeseg 61 pavgppqplg kelqdgaesn gggggggags gggpgaepdl knsngnagpr palnnnltep 121 pggggggssd gvgapphsaa aalpppaygf gqpygrspsa vaaaaaavfh qqhggqqspg 181 laalqsgggg glepyagpqq nshdhgfpnh qynsyypnrs aypppapaya lssprggtpg 241 sgaaaaagsk pppsssasas sssssfaqqr fgamggggps aagggtpqpt atptlnqllt 301 spssargyqg ypggdysggp qdggagkgpa dmasqcwgaa aaaaaaaaas ggaqqrshha 361 pmspgssggg gqplartpqp sspmdqmgkm rpqpyggtnp ysqqqgppsg pqqghgypgq 421 pygsqtpqry pmtmqgraqs amgglsytqq ippygqqgps gygqqgqtpy ynqqsphpqq 481 qqppysqqpp sqtphaqpsy qqqpqsqppq lqssqppysq qpsqpphqqs papypsqqst 541 tqqhpqsqpp ysqpqaqspy qqqqpqqpap stlsqqaayp qpqsqqsqqt aysqqrfppp 601 qelsqdsfgs qassapsmts skggqedmnl slqsrpsslp dlsgsiddlp mgtegalspg 661 vstsgisssq gegsnpaqsp fsphtsphlp girgpspspv gspasvaqsr sgplspaavp 721 gnqmpprpps gqsdsimhps mnqssiaqdr gymqrnpqmp qysspqpgsa lsprqpsggq 781 ihtgmgsyqq nsmgsygpqg gqygpqggyp rqpnynalpn anypsagmag ginpmgaggq 841 mhgqpgippy gtlppgrmsh asmgnrpygp nmanmppqvg sgmcpppggm nrktqetava 901 mhvaansiqn rppgypnmnq ggmmgtgppy gqginsmagm inpqgppysm ggtmannsag 961 maaspemmgl gdvkltpatk mnnkadgtpk teskskksss stttnekitk lyelggeper 1021 kmwvdrylaf teekamgmtn lpavgrkpld lyrlyvsvke iggltqvnkn kkwrelatnl 1081 nvgtsssaas slkkgyiqcl yafeckierg edpppdifaa adskksqpki qppspagsgs 1141 mqgpqtpqst sssmaeggdl kpptpastph sqipplpgms rsnsvgiqda fndgsdstfq 1201 krnsmtpnpg yqpsmntsdm mgrmsyepnk dpygsmrkap gsdpfmssgq gpnggmgdpy 1261 sraagpglgn vamgprqhyp yggpydrvrt epgigpegnm stgapqpnlm psnpdsgmys 1321 psryppqqqq qqqqrhdsyg nqfstqgtps gspfpsqqtt myqqqqqnyk rpmdgtygpp 1381 akrhegemys vpystgqgqp qqqqlppaqp qpasqqqaaq pspqqdvynq ygnaypatat 1441 aaterrpagg pqnqfpfqfg rdrvsappgt naqqnmppqm mggpiqasae vaqqgtmwqg 1501 rndmtynyan rqstgsapqg payhgvnrtd emlhtdqran hegswpshgt rqppygpsap 1561 vppmtrppps nyqpppsmqn hipqvsspap lprpmenrts pskspflhsg mkmqkagppv 1621 pashiapapv qppmirrdit fppgsveatq pvlkgrrrlt mkdigtpeaw rvmmslksgl 1681 laestwaldt inillyddns imtfnlsqlp gllellveyf rrclieifgi lkeyevgdpg 1741 qrtlldpgrf skvsspapme ggeeeeellg pkleeeeeee vvendeeiaf sgkdkpasen 1801 seekliskfd klpvkivqkn dpfvvdcsdk lgrvqefdsg llhwrigggd ttehiqthfe 1861 sktellpsrp hapcppaprk hvttaegtpg ttdqegpppd gppekritat mddmlstrss 1921 tltedgakss eaikesskfp fgispaqshr nikiledeph skdetplctl ldwqdslakr 1981 cvcvsntirs lsfvpgndfe mskhpgllli lgklillhhk hperkqaplt yekeeeqdqg 2041 vscnkvewww dclemlrent lvtlanisgq ldlspypesi clpvldgllh wavcpsaeaq 2101 dpfstlgpna vlspqrlvle tlsklsiqdn nvdlilatpp fsrleklyst mvrflsdrkn 2161 pvcremavvl lanlaggdsl aaraiavqkg signllgfle dslaatqfqq sgasllhmqn 2221 ppfeptsvdm mrraaralla lakvdenhse ftlyesrlld isvsplmnsl vsqvicdvlf 2281 ligqs Homo sapiens AT rich interactive domain 1A (SWI-like)(ARID1A), transcript variant 1, mRNA (SEQ ID NO: 10) 1 cagaaagcgg agagtcacag cggggccagg ccctggggag cggagcctcc accgcccccc 61 tcattcccag gcaagggctt ggggggaatg agccgggaga gccgggtccc gagcctacag 121 agccgggagc agctgagccg ccggcgcctc ggccgccgcc gccgcctcct cctcctccgc 181 cgccgccagc ccggagcctg agccggcggg gcggggggga gaggagcgag cgcagcgcag 241 cagcggagcc ccgcgaggcc cgcccgggcg ggtggggagg gcagcccggg ggactgggcc 301 ccggggcggg gtgggagggg gggagaagac gaagacaggg ccgggtctct ccgcggacga 361 gacagcgggg atcatggccg cgcaggtcgc ccccgccgcc gccagcagcc tgggcaaccc 421 gccgccgccg ccgccctcgg agctgaagaa agccgagcag cagcagcggg aggaggcggg 481 gggcgaggcg gcggcggcgg cagcggccga gcgcggggaa atgaaggcag ccgccgggca 541 ggaaagcgag ggccccgccg tggggccgcc gcagccgctg ggaaaggagc tgcaggacgg 601 ggccgagagc aatgggggtg gcggcggcgg cggagccggc agcggcggcg ggcccggcgc 661 ggagccggac ctgaagaact cgaacgggaa cgcgggccct aggcccgccc tgaacaataa 721 cctcacggag ccgcccggcg gcggcggtgg cggcagcagc gatggggtgg gggcgcctcc 781 tcactcagcc gcggccgcct tgccgccccc agcctacggc ttcgggcaac cctacggccg 841 gagcccgtct gccgtcgccg ccgccgcggc cgccgtcttc caccaacaac atggcggaca 901 acaaagccct ggcctggcag cgctgcagag cggcggcggc gggggcctgg agccctacgc 961 ggggccccag cagaactctc acgaccacgg cttccccaac caccagtaca actcctacta 1021 ccccaaccgc agcgcctacc ccccgcccgc cccggcctac gcgctgagct ccccgagagg 1081 tggcactccg ggctccggcg cggcggcggc tgccggctcc aagccgcctc cctcctccag 1141 cgcctccgcc tcctcgtcgt cttcgtcctt cgctcagcag cgcttcgggg ccatgggggg 1201 aggcggcccc tccgcggccg gcgggggaac tccccagccc accgccaccc ccaccctcaa 1261 ccaactgctc acgtcgccca gctcggcccg gggctaccag ggctaccccg ggggcgacta 1321 cagtggcggg ccccaggacg ggggcgccgg caagggcccg gcggacatgg cctcgcagtg 1381 ttggggggct gcggcggcgg cagctgcggc ggcggccgcc tcgggagggg cccaacaaag 1441 gagccaccac gcgcccatga gccccgggag cagcggcggc ggggggcagc cgctcgcccg 1501 gacccctcag ccatccagtc caatggatca gatgggcaag atgagacctc agccatatgg 1561 cgggactaac ccatactcgc agcaacaggg acctccgtca ggaccgcagc aaggacatgg 1621 gtacccaggg cagccatacg ggtcccagac cccgcagcgg tacccgatga ccatgcaggg 1681 ccgggcgcag agtgccatgg gcggcctctc ttatacacag cagattcctc cttatggaca 1741 acaaggcccc agcgggtatg gtcaacaggg ccagactcca tattacaacc agcaaagtcc 1801 tcaccctcag cagcagcagc caccctactc ccagcaacca ccgtcccaga cccctcatgc 1861 ccaaccttcg tatcagcagc agccacagtc tcaaccacca cagctccagt cctctcagcc 1921 tccatactcc cagcagccat cccagcctcc acatcagcag tccccggctc catacccctc 1981 ccagcagtcg acgacacagc agcaccccca gagccagccc ccctactcac agccacaggc 2041 tcagtctcct taccagcagc agcaacctca gcagccagca ccctcgacgc tctcccagca 2101 ggctgcgtat cctcagcccc agtctcagca gtcccagcaa actgcctatt cccagcagcg 2161 cttccctcca ccgcaggagc tatctcaaga ttcatttggg tctcaggcat cctcagcccc 2221 ctcaatgacc tccagtaagg gagggcaaga agatatgaac ctgagccttc agtcaagacc 2281 ctccagcttg cctgatctat ctggttcaat agatgacctc cccatgggga cagaaggagc 2341 tctgagtcct ggagtgagca catcagggat ttccagcagc caaggagagc agagtaatcc 2401 agctcagtct cctttctctc ctcatacctc ccctcacctg cctggcatcc gaggcccttc 2461 cccgtcccct gttggctctc ccgccagtgt tgctcagtct cgctcaggac cactctcgcc 2521 tgctgcagtg ccaggcaacc agatgccacc tcggccaccc agtggccagt cggacagcat 2581 catgcatcct tccatgaacc aatcaagcat tgcccaagat cgaggttata tgcagaggaa 2641 cccccagatg ccccagtaca gttcccccca gcccggctca gccttatctc cgcgtcagcc 2701 ttccggagga cagatacaca caggcatggg ctcctaccag cagaactcca tggggagcta 2761 tggtccccag gggggtcagt atggcccaca aggtggctac cccaggcagc caaactataa 2821 tgccttgccc aatgccaact accccagtgc aggcatggct ggaggcataa accccatggg 2881 tgccggaggt caaatgcatg gacagcctgg catcccacct tatggcacac tccctccagg 2941 gaggatgagt cacgcctcca tgggcaaccg gccttatggc cctaacatgg ccaatatgcc 3001 acctcaggtt gggtcaggga tgtgtccccc accagggggc atgaaccgga aaacccaaga 3061 aactgctgtc gccatgcatg ttgctgccaa ctctatccaa aacaggccgc caggctaccc 3121 caatatgaat caagggggca tgatgggaac tggacctcct tatggacaag ggattaatag 3181 tatggctggc atgatcaacc ctcagggacc cccatattcc atgggtggaa ccatggccaa 3241 caattctgca gggatggcag ccagcccaga gatgatgggc cttggggatg taaagttaac 3301 tccagccacc aaaatgaaca acaaggcaga tgggacaccc aagacagaat ccaaatccaa 3361 gaaatccagt tcttctacta caaccaatga gaagatcacc aagttgtatg agctgggtgg 3421 tgagcctgag aggaagatgt gggtggaccg ttatctggcc ttcactgagg agaaggccat 3481 gggcatgaca aatctgcctg ctgtgggtag gaaacctctg gacctctatc gcctctatgt 3541 gtctgtgaag gagattggtg gattgactca ggtcaacaag aacaaaaaat ggcgggaact 3601 tgcaaccaac ctcaatgtgg gcacatcaag cagtgctgcc agctccttga aaaagcagta 3661 tatccagtgt ctctatgcct ttgaatgcaa gattgaacgg ggagaagacc ctcccccaga 3721 catctttgca gctgctgatt ccaagaagtc ccagcccaag atccagcctc cctctcctgc 3781 gggatcagga tctatgcagg ggccccagac tccccagtca accagcagtt ccatggcaga 3841 aggaggagac ttaaagccac caactccagc atccacacca cacagtcaga tccccccatt 3901 gccaggcatg agcaggagca attcagttgg gatccaggat gcctttaatg atggaagtga 3961 ctccacattc cagaagcgga attccatgac tccaaaccct gggtatcagc ccagtatgaa 4021 tacctctgac atgatggggc gcatgtccta tgagccaaat aaggatcctt atggcagcat 4081 gaggaaagct ccagggagtg atcccttcat gtcctcaggg cagggcccca acggcgggat 4141 gggtgacccc tacagtcgtg ctgccggccc tgggctagga aatgtggcga tgggaccacg 4201 acagcactat ccctatggag gtccttatga cagagtgagg acggagcctg gaatagggcc 4261 tgagggaaac atgagcactg gggccccaca gccgaatctc atgccttcca acccagactc 4321 ggggatgtat tctcctagcc gctacccccc gcagcagcag cagcagcagc agcaacgaca 4381 tgattcctat ggcaatcagt tctccaccca aggcacccct tctggcagcc ccttccccag 4441 ccagcagact acaatgtatc aacagcaaca gcagaattac aagcggccaa tggatggcac 4501 atatggccct cctgccaagc ggcacgaagg ggagatgtac agcgtgccat acagcactgg 4561 gcaggggcag cctcagcagc agcagttgcc cccagcccag ccccagcctg ccagccagca 4621 acaagctgcc cagccttccc ctcagcaaga tgtatacaac cagtatggca atgcctatcc 4681 tgccactgcc acagctgcta ctgagcgccg accagcaggc ggcccccaga accaatttcc 4741 attccagttt ggccgagacc gtgtctctgc accccctggc accaatgccc agcaaaacat 4801 gccaccacaa atgatgggcg gccccataca ggcatcagct gaggttgctc agcaaggcac 4861 catgtggcag gggcgtaatg acatgaccta taattatgcc aacaggcaga gcacgggctc 4921 tgccccccag ggccccgcct atcatggcgt gaaccgaaca gatgaaatgc tgcacacaga
4981 tcagagggcc aaccacgaag gctcgtggcc ttcccatggc acacgccagc ccccatatgg 5041 tccctctgcc cctgtgcccc ccatgacaag gccccctcca tctaactacc agcccccacc 5101 aagcatgcag aatcacattc ctcaggtatc cagccctgct cccctgcccc ggccaatgga 5161 gaaccgcacc tctcctagca agtctccatt cctgcactct gggatgaaaa tgcagaaggc 5221 aggtccccca gtacctgcct cgcacatagc acctgcccct gtgcagcccc ccatgattcg 5281 gcgggatatc accttcccac ctggctctgt tgaagccaca cagcctgtgt tgaagcagag 5341 gaggcggctc acaatgaaag acattggaac cccggaggca tggcgggtaa tgatgtccct 5401 caagtctggt ctcctggcag agagcacatg ggcattagat accatcaaca tcctgctgta 5461 tgatgacaac agcatcatga ccttcaacct cagtcagctc ccagggttgc tagagctcct 5521 tgtagaatat ttccgacgat gcctgattga gatctttggc attttaaagg agtatgaggt 5581 gggtgaccca ggacagagaa cgctactgga tcctgggagg ttcagcaagg tgtctagtcc 5641 agctcccatg gagggtgggg aagaagaaga agaacttcta ggtcctaaac tagaagagga 5701 agaagaagag gaagtagttg aaaatgatga ggagatagcc ttttcaggca aggacaagcc 5761 agcttcagag aatagtgagg agaagctgat cagtaagttt gacaagcttc cagtaaagat 5821 cgtacagaag aatgatccat ttgtggtgga ctgctcagat aagcttgggc gtgtgcagga 5881 gtttgacagt ggcctgctgc actggcggat tggtgggggg gacaccactg agcatatcca 5941 gacccacttc gagagcaaga cagagctgct gccttcccgg cctcacgcac cctgcccacc 6001 agcccctcgg aagcatgtga caacagcaga gggtacacca gggacaacag accaggaggg 6061 gcccccacct gatggacctc cagaaaaacg gatcacagcc actatggatg acatgttgtc 6121 tactcggtct agcaccttga ccgaggatgg agctaagagt tcagaggcca tcaaggagag 6181 cagcaagttt ccatttggca ttagcccagc acagagccac cggaacatca agatcctaga 6241 ggacgaaccc cacagtaagg atgagacccc actgtgtacc cttctggact ggcaggattc 6301 tcttgccaag cgctgcgtct gtgtgtccaa taccattcga agcctgtcat ttgtgccagg 6361 caatgacttt gagatgtcca aacacccagg gctgctgctc atcctgggca agctgatcct 6421 gctgcaccac aagcacccag aacggaagca ggcaccacta acttatgaaa aggaggagga 6481 acaggaccaa ggggtgagct gcaacaaagt ggagtggtgg tgggactgct tggagatgct 6541 ccgggaaaac accttggtta cactcgccaa catctcgggg cagttggacc tatctccata 6601 ccccgagagc atttgcctgc ctgtcctgga cggactccta cactgggcag tttgcccttc 6661 agctgaagcc caggacccct tttccaccct gggccccaat gccgtccttt ccccgcagag 6721 actggtcttg gaaaccctca gcaaactcag catccaggac aacaatgtgg acctgattct 6781 ggccacaccc cccttcagcc gcctggagaa gttgtatagc actatggtgc gcttcctcag 6841 tgaccgaaag aacccggtgt gccgggagat ggctgtggta ctgctggcca acctggctca 6901 gggggacagc ctggcagctc gtgccattgc agtgcagaag ggcagtatcg gcaacctcct 6961 gggcttccta gaggacagcc ttgccgccac acagttccag cagagccagg ccagcctcct 7021 ccacatgcag aacccaccct ttgagccaac tagtgtggac atgatgcggc gggctgcccg 7081 cgcgctgctt gccttggcca aggtggacga gaaccactca gagtttactc tgtacgaatc 7141 acggctgttg gacatctcgg tatcaccgtt gatgaactca ttggtttcac aagtcatttg 7201 tgatgtactg tttttgattg gccagtcatg acagccgtgg gacacctccc ccccccgtgt 7261 gtgtgtgcgt gtgtggagaa cttagaaact gactgttgcc ctttatttat gcaaaaccac 7321 ctcagaatcc agtttaccct gtgctgtcca gcttctccct tgggaaaaag tctctcctgt 7381 ttctctctcc tccttccacc tcccctccct ccatcacctc acgcctttct gttccttgtc 7441 ctcaccttac tcccctcagg accctacccc accctctttg aaaagacaaa gctctgccta 7501 catagaagac tttttttatt ttaaccaaag ttactgttgt ttacagtgag tttggggaaa 7561 aaaaataaaa taaaaatggc tttcccagtc cttgcatcaa cgggatgcca catttcataa 7621 ctgtttttaa tggtaaaaaa aaaaaaaaaa aatacaaaaa aaaattctga aggacaaaaa 7681 aggtgactgc tgaactgtgt gtggtttatt gttgtacatt cacaatcttg caggagccaa 7741 gaagttcgca gttgtgaaca gaccctgttc actggagagg cctgtgcagt agagtgtaga 7801 ccctttcatg tactgtactg tacacctgat actgtaaaca tactgtaata ataatgtctc 7861 acatggaaac agaaaacgct gggtcagcag caagctgtag tttttaaaaa tgtttttagt 7921 taaacgttga ggagaaaaaa aaaaaaggct tttcccccaa agtatcatgt gtgaacctac 7981 aacaccctga cctctttctc tcctccttga ttgtatgaat aaccctgaga tcacctctta 8041 gaactggttt taacctttag ctgcagcggc tacgctgcca cgtgtgtata tatatgacgt 8101 tgtacattgc acataccctt ggatccccac agtttggtcc tcctcccagc taccccttta 8161 tagtatgacg agttaacaag ttggtgacct gcacaaagcg agacacagct atttaatctc 8221 ttgccagata tcgcccctct tggtgcgatg ctgtacaggt ctctgtaaaa agtccttgct 8281 gtctcagcag ccaatcaact tatagtttat ttttttctgg gtttttgttt tgttttgttt 8341 tctttctaat cgaggtgtga aaaagttcta ggttcagttg aagttctgat gaagaaacac 8401 aattgagatt ttttcagtga taaaatctgc atatttgtat ttcaacaatg tagctaaaac 8461 ttgatgtaaa ttcctccttt ttttcctttt ttggcttaat gaatatcatt tattcagtat 8521 gaaatcttta tactatatgt tccacgtgtt aagaataaat gtacattaaa tcttggtaag 8581 acttt AT-rich interactive domain-containing protein 1A (ARID1A) isoform b (SEQ ID NO: 11) 1 maaqvapaaa sslgnppppp pselkkaegq qreeaggeaa aaaaaergem kaaagqeseg 61 pavgppqplg kelqdgaesn gggggggags gggpgaepdl knsngnagpr palnnnltep 121 pggggggssd gvgapphsaa aalpppaygf gqpygrspsa vaaaaaavfh qqhggqqspg 181 laalqsgggg glepyagpqq nshdhgfpnh qynsyypnrs aypppapaya lssprggtpg 241 sgaaaaagsk pppsssasas sssssfaqqr fgamggggps aagggtpqpt atptlnqllt 301 spssargyqg ypggdysggp qdggagkgpa dmasqcwgaa aaaaaaaaas ggaqqrshha 361 pmspgssggg gqplartpqp sspmdqmgkm rpqpyggtnp ysqqqgppsg pqqghgypgq 421 pygsqtpqry pmtmggraqs amgglsytqq ippygqqgps gygqqgqtpy ynggsphpqq 481 qqppysqqpp sqtphaqpsy qqqpqsqppq lqssqppysq qpsqpphqqs papypsqqst 541 tqqhpqsqpp ysqpqaqspy qqqqpqqpap stlsqqaayp qpqsqqsqqt aysqqrfppp 601 qelsqdsfgs qassapsmts skggqedmnl slqsrpsslp dlsgsiddlp mgtegalspg 661 vstsgisssq geqsnpaqsp fsphtsphlp girgpspspv gspasvaqsr sgplspaavp 721 gnqmpprpps gqsdsimhps mnqssiaqdr gymqrnpqmp qysspqpgsa lsprqpsggq 781 ihtgmgsyqq nsmgsygpqg gqygpqggyp rqpnynalpn anypsagmag ginpmgaggq 841 mhgqpgippy gtlppgrmsh asmgnrpygp nmanmppqvg sgmcpppggm nrktqetava 901 mhvaansiqn rppgypnmnq ggmmgtgppy gqginsmagm inpqgppysm ggtmannsag 961 maaspemmgl gdvkltpatk mnnkadgtpk teskskksss stttnekitk lyelggeper 1021 kmwvdrylaf teekamgmtn lpavgrkpld lyrlyvsvke iggltqvnkn kkwrelatnl 1081 nvgtsssaas slkkgyiqcl yafeckierg edpppdifaa adskksqpki qppspagsgs 1141 mqgpqtpqst sssmaeggdl kpptpastph sqipplpgms rsnsvgiqda fndgsdstfq 1201 krnsmtpnpg yqpsmntsdm mgrmsyepnk dpygsmrkap gsdpfmssgq gpnggmgdpy 1261 sraagpglgn vamgprqhyp yggpydrvrt epgigpegnm stgapqpnlm psnpdsgmys 1321 psryppqqqq qqqqrhdsyg nqfstqgtps gspfpsqqtt myqqqqqvss paplprpmen 1381 rtspskspfl hsgmkmqkag ppvpashiap apvqppmirr ditfppgsve atqpvlkqrr 1441 rltmkdigtp eawrvmmslk sgllaestwa ldtinillyd dnsimtfnls qlpgllellv 1501 eyfrrcliei fgilkeyevg dpgqrtlldp grfskvsspa pmeggeeeee llgpkleeee 1561 eeevvendee iafsgkdkpa senseeklis kfdklpvkiv qkndpfvvdc sdklgrvqef 1621 dsgllhwrig ggdttehiqt hfesktellp srphapcppa prkhvttaeg tpgttdqegp 1681 ppdgppekri tatmddmlst rsstltedga ksseaikess kfpfgispaq shrnikiled 1741 ephskdetpl ctlldwqdsl akrcvcvsnt irslsfvpgn dfemskhpgl llilgklill 1801 hhkhperkqa pltyekeeeq dqgvscnkve wwwdclemlr entlvtlani sgqldlspyp 1861 esiclpvldg llhwavcpsa eaqdpfstlg pnavlspqrl vletlsklsi qdnnvdlila 1921 tppfsrlekl ystmvrflsd rknpvcrema vvllanlaqg dslaaraiav qkgsignllg 1981 fledslaatq fqqsqasllh mqnppfepts vdmmrraara llalakvden hseftlyesr 2041 lldisvsplm nslvsqvicd vlfligqs Homo sapiens AT rich interactive domain 1A (SWI-like)(ARID1A), transcript variant 2, mRNA (SEQ ID NO: 12) 1 cagaaagcgg agagtcacag cggggccagg ccctggggag cggagcctcc accgcccccc 61 tcattcccag gcaagggctt ggggggaatg agccgggaga gccgggtccc gagcctacag 121 agccgggagc agctgagccg ccggcgcctc ggccgccgcc gccgcctcct cctcctccgc 181 cgccgccagc ccggagcctg agccggcggg gcggggggga gaggagcgag cgcagcgcag 241 cagcggagcc ccgcgaggcc cgcccgggcg ggtggggagg gcagcccggg ggactgggcc 301 ccggggcggg gtgggagggg gggagaagac gaagacaggg ccgggtctct ccgcggacga 361 gacagcgggg atcatggccg cgcaggtcgc ccccgccgcc gccagcagcc tgggcaaccc 421 gccgccgccg ccgccctcgg agctgaagaa agccgagcag cagcagcggg aggaggcggg 481 gggcgaggcg gcggcggcgg cagcggccga gcgcggggaa atgaaggcag ccgccgggca 541 ggaaagcgag ggccccgccg tggggccgcc gcagccgctg ggaaaggagc tgcaggacgg 601 ggccgagagc aatgggggtg gcggcggcgg cggagccggc agcggcggcg ggcccggcgc 661 ggagccggac ctgaagaact cgaacgggaa cgcgggccct aggcccgccc tgaacaataa 721 cctcacggag ccgcccggcg gcggcggtgg cggcagcagc gatggggtgg gggcgcctcc 781 tcactcagcc gcggccgcct tgccgccccc agcctacggc ttcgggcaac cctacggccg 841 gagcccgtct gccgtcgccg ccgccgcggc cgccgtcttc caccaacaac atggcggaca 901 acaaagccct ggcctggcag cgctgcagag cggcggcggc gggggcctgg agccctacgc 961 ggggccccag cagaactctc acgaccacgg cttccccaac caccagtaca actcctacta 1021 ccccaaccgc agcgcctacc ccccgcccgc cccggcctac gcgctgagct ccccgagagg 1081 tggcactccg ggctccggcg cggcggcggc tgccggctcc aagccgcctc cctcctccag 1141 cgcctccgcc tcctcgtcgt cttcgtcctt cgctcagcag cgcttcgggg ccatgggggg 1201 aggcggcccc tccgcggccg gcgggggaac tccccagccc accgccaccc ccaccctcaa 1261 ccaactgctc acgtcgccca gctcggcccg gggctaccag ggctaccccg ggggcgacta 1321 cagtggcggg ccccaggacg ggggcgccgg caagggcccg gcggacatgg cctcgcagtg 1381 ttggggggct gcggcggcgg cagctgcggc ggcggccgcc tcgggagggg cccaacaaag 1441 gagccaccac gcgcccatga gccccgggag cagcggcggc ggggggcagc cgctcgcccg 1501 gacccctcag ccatccagtc caatggatca gatgggcaag atgagacctc agccatatgg
1561 cgggactaac ccatactcgc agcaacaggg acctccgtca ggaccgcagc aaggacatgg 1621 gtacccaggg cagccatacg ggtcccagac cccgcagcgg tacccgatga ccatgcaggg 1681 ccgggcgcag agtgccatgg gcggcctctc ttatacacag cagattcctc cttatggaca 1741 acaaggcccc agcgggtatg gtcaacaggg ccagactcca tattacaacc agcaaagtcc 1801 tcaccctcag cagcagcagc caccctactc ccagcaacca ccgtcccaga cccctcatgc 1861 ccaaccttcg tatcagcagc agccacagtc tcaaccacca cagctccagt cctctcagcc 1921 tccatactcc cagcagccat cccagcctcc acatcagcag tccccggctc catacccctc 1981 ccagcagtcg acgacacagc agcaccccca gagccagccc ccctactcac agccacaggc 2041 tcagtctcct taccagcagc agcaacctca gcagccagca ccctcgacgc tctcccagca 2101 ggctgcgtat cctcagcccc agtctcagca gtcccagcaa actgcctatt cccagcagcg 2161 cttccctcca ccgcaggagc tatctcaaga ttcatttggg tctcaggcat cctcagcccc 2221 ctcaatgacc tccagtaagg gagggcaaga agatatgaac ctgagccttc agtcaagacc 2281 ctccagcttg cctgatctat ctggttcaat agatgacctc cccatgggga cagaaggagc 2341 tctgagtcct ggagtgagca catcagggat ttccagcagc caaggagagc agagtaatcc 2401 agctcagtct cctttctctc ctcatacctc ccctcacctg cctggcatcc gaggcccttc 2461 cccgtcccct gttggctctc ccgccagtgt tgctcagtct cgctcaggac cactctcgcc 2521 tgctgcagtg ccaggcaacc agatgccacc tcggccaccc agtggccagt cggacagcat 2581 catgcatcct tccatgaacc aatcaagcat tgcccaagat cgaggttata tgcagaggaa 2641 cccccagatg ccccagtaca gttcccccca gcccggctca gccttatctc cgcgtcagcc 2701 ttccggagga cagatacaca caggcatggg ctcctaccag cagaactcca tggggagcta 2761 tggtccccag gggggtcagt atggcccaca aggtggctac cccaggcagc caaactataa 2821 tgccttgccc aatgccaact accccagtgc aggcatggct ggaggcataa accccatggg 2881 tgccggaggt caaatgcatg gacagcctgg catcccacct tatggcacac tccctccagg 2941 gaggatgagt cacgcctcca tgggcaaccg gccttatggc cctaacatgg ccaatatgcc 3001 acctcaggtt gggtcaggga tgtgtccccc accagggggc atgaaccgga aaacccaaga 3061 aactgctgtc gccatgcatg ttgctgccaa ctctatccaa aacaggccgc caggctaccc 3121 caatatgaat caagggggca tgatgggaac tggacctcct tatggacaag ggattaatag 3181 tatggctggc atgatcaacc ctcagggacc cccatattcc atgggtggaa ccatggccaa 3241 caattctgca gggatggcag ccagcccaga gatgatgggc cttggggatg taaagttaac 3301 tccagccacc aaaatgaaca acaaggcaga tgggacaccc aagacagaat ccaaatccaa 3361 gaaatccagt tcttctacta caaccaatga gaagatcacc aagttgtatg agctgggtgg 3421 tgagcctgag aggaagatgt gggtggaccg ttatctggcc ttcactgagg agaaggccat 3481 gggcatgaca aatctgcctg ctgtgggtag gaaacctctg gacctctatc gcctctatgt 3541 gtctgtgaag gagattggtg gattgactca ggtcaacaag aacaaaaaat ggcgggaact 3601 tgcaaccaac ctcaatgtgg gcacatcaag cagtgctgcc agctccttga aaaagcagta 3661 tatccagtgt ctctatgcct ttgaatgcaa gattgaacgg ggagaagacc ctcccccaga 3721 catctttgca gctgctgatt ccaagaagtc ccagcccaag atccagcctc cctctcctgc 3781 gggatcagga tctatgcagg ggccccagac tccccagtca accagcagtt ccatggcaga 3841 aggaggagac ttaaagccac caactccagc atccacacca cacagtcaga tccccccatt 3901 gccaggcatg agcaggagca attcagttgg gatccaggat gcctttaatg atggaagtga 3961 ctccacattc cagaagcgga attccatgac tccaaaccct gggtatcagc ccagtatgaa 4021 tacctctgac atgatggggc gcatgtccta tgagccaaat aaggatcctt atggcagcat 4081 gaggaaagct ccagggagtg atcccttcat gtcctcaggg cagggcccca acggcgggat 4141 gggtgacccc tacagtcgtg ctgccggccc tgggctagga aatgtggcga tgggaccacg 4201 acagcactat ccctatggag gtccttatga cagagtgagg acggagcctg gaatagggcc 4261 tgagggaaac atgagcactg gggccccaca gccgaatctc atgccttcca acccagactc 4321 ggggatgtat tctcctagcc gctacccccc gcagcagcag cagcagcagc agcaacgaca 4381 tgattcctat ggcaatcagt tctccaccca aggcacccct tctggcagcc ccttccccag 4441 ccagcagact acaatgtatc aacagcaaca gcaggtatcc agccctgctc ccctgccccg 4501 gccaatggag aaccgcacct ctcctagcaa gtctccattc ctgcactctg ggatgaaaat 4561 gcagaaggca ggtcccccag tacctgcctc gcacatagca cctgcccctg tgcagccccc 4621 catgattcgg cgggatatca ccttcccacc tggctctgtt gaagccacac agcctgtgtt 4681 gaagcagagg aggcggctca caatgaaaga cattggaacc ccggaggcat ggcgggtaat 4741 gatgtccctc aagtctggtc tcctggcaga gagcacatgg gcattagata ccatcaacat 4801 cctgctgtat gatgacaaca gcatcatgac cttcaacctc agtcagctcc cagggttgct 4861 agagctcctt gtagaatatt tccgacgatg cctgattgag atctttggca ttttaaagga 4921 gtatgaggtg ggtgacccag gacagagaac gctactggat cctgggaggt tcagcaaggt 4981 gtctagtcca gctcccatgg agggtgggga agaagaagaa gaacttctag gtcctaaact 5041 agaagaggaa gaagaagagg aagtagttga aaatgatgag gagatagcct tttcaggcaa 5101 ggacaagcca gcttcagaga atagtgagga gaagctgatc agtaagtttg acaagcttcc 5161 agtaaagatc gtacagaaga atgatccatt tgtggtggac tgctcagata agcttgggcg 5221 tgtgcaggag tttgacagtg gcctgctgca ctggcggatt ggtggggggg acaccactga 5281 gcatatccag acccacttcg agagcaagac agagctgctg ccttcccggc ctcacgcacc 5341 ctgcccacca gcccctcgga agcatgtgac aacagcagag ggtacaccag ggacaacaga 5401 ccaggagggg cccccacctg atggacctcc agaaaaacgg atcacagcca ctatggatga 5461 catgttgtct actcggtcta gcaccttgac cgaggatgga gctaagagtt cagaggccat 5521 caaggagagc agcaagtttc catttggcat tagcccagca cagagccacc ggaacatcaa 5581 gatcctagag gacgaacccc acagtaagga tgagacccca ctgtgtaccc ttctggactg 5641 gcaggattct cttgccaagc gctgcgtctg tgtgtccaat accattcgaa gcctgtcatt 5701 tgtgccaggc aatgactttg agatgtccaa acacccaggg ctgctgctca tcctgggcaa 5761 gctgatcctg ctgcaccaca agcacccaga acggaagcag gcaccactaa cttatgaaaa 5821 ggaggaggaa caggaccaag gggtgagctg caacaaagtg gagtggtggt gggactgctt 5881 ggagatgctc cgggaaaaca ccttggttac actcgccaac atctcggggc agttggacct 5941 atctccatac cccgagagca tttgcctgcc tgtcctggac ggactcctac actgggcagt 6001 ttgcccttca gctgaagccc aggacccctt ttccaccctg ggccccaatg ccgtcctttc 6061 cccgcagaga ctggtcttgg aaaccctcag caaactcagc atccaggaca acaatgtgga 6121 cctgattctg gccacacccc ccttcagccg cctggagaag ttgtatagca ctatggtgcg 6181 cttcctcagt gaccgaaaga acccggtgtg ccgggagatg gctgtggtac tgctggccaa 6241 cctggctcag ggggacagcc tggcagctcg tgccattgca gtgcagaagg gcagtatcgg 6301 caacctcctg ggcttcctag aggacagcct tgccgccaca cagttccagc agagccaggc 6361 cagcctcctc cacatgcaga acccaccctt tgagccaact agtgtggaca tgatgcggcg 6421 ggctgcccgc gcgctgcttg ccttggccaa ggtggacgag aaccactcag agtttactct 6481 gtacgaatca cggctgttgg acatctcggt atcaccgttg atgaactcat tggtttcaca 6541 agtcatttgt gatgtactgt ttttgattgg ccagtcatga cagccgtggg acacctcccc 6601 cccccgtgtg tgtgtgcgtg tgtggagaac ttagaaactg actgttgccc tttatttatg 6661 caaaaccacc tcagaatcca gtttaccctg tgctgtccag cttctccctt gggaaaaagt 6721 ctctcctgtt tctctctcct ccttccacct cccctccctc catcacctca cgcctttctg 6781 ttccttgtcc tcaccttact cccctcagga ccctacccca ccctctttga aaagacaaag 6841 ctctgcctac atagaagact ttttttattt taaccaaagt tactgttgtt tacagtgagt 6901 ttggggaaaa aaaataaaat aaaaatggct ttcccagtcc ttgcatcaac gggatgccac 6961 atttcataac tgtttttaat ggtaaaaaaa aaaaaaaaaa atacaaaaaa aaattctgaa 7021 ggacaaaaaa ggtgactgct gaactgtgtg tggtttattg ttgtacattc acaatcttgc 7081 aggagccaag aagttcgcag ttgtgaacag accctgttca ctggagaggc ctgtgcagta 7141 gagtgtagac cctttcatgt actgtactgt acacctgata ctgtaaacat actgtaataa 7201 taatgtctca catggaaaca gaaaacgctg ggtcagcagc aagctgtagt ttttaaaaat 7261 gtttttagtt aaacgttgag gagaaaaaaa aaaaaggctt ttcccccaaa gtatcatgtg 7321 tgaacctaca acaccctgac ctctttctct cctccttgat tgtatgaata accctgagat 7381 cacctcttag aactggtttt aacctttagc tgcagcggct acgctgccac gtgtgtatat 7441 atatgacgtt gtacattgca catacccttg gatccccaca gtttggtcct cctcccagct 7501 acccctttat agtatgacga gttaacaagt tggtgacctg cacaaagcga gacacagcta 7561 tttaatctct tgccagatat cgcccctctt ggtgcgatgc tgtacaggtc tctgtaaaaa 7621 gtccttgctg tctcagcagc caatcaactt atagtttatt tttttctggg tttttgtttt 7681 gttttgtttt ctttctaatc gaggtgtgaa aaagttctag gttcagttga agttctgatg 7741 aagaaacaca attgagattt tttcagtgat aaaatctgca tatttgtatt tcaacaatgt 7801 agctaaaact tgatgtaaat tcctcctttt tttccttttt tggcttaatg aatatcattt 7861 attcagtatg aaatctttat actatatgtt ccacgtgtta agaataaatg tacattaaat 7921 cttggtaaga cttt
[0118] The present invention also provides methods of inducing neuronal differentiation by contacting a cell with a compound (i.e., an EZH2 inhibitor) of the invention. Preferably, the compound is in an amount sufficient to increase expression of at least one gene selected from the group consisting of CD133 (also called PROM1), DOCK4, PTPRK, PROM2, LHX1, LHX6, LHX9, PAX6, PAX7, VEFGA, FZD3B, FYN, HIF1A, HTRA2, EVX1, CCDC64, and GFAP.
[0119] The term "inducing neuronal differentiation" used herein refers to causing a cell to develop into a cell of the neuronal lineage as a result of a direct or intentional effect on the cell.
[0120] The present invention also provides methods of inducing cell cycle inhibition by contacting a cell with a compound of the invention. Preferably, the compound is in an amount sufficient to increase expression of at least one gene selected from the group consisting of CKDN1A, CDKN2A, MEN1, CHEK1, IRF6, ALOX15B, CYP27B1, DBC1, NME6, GMNN, HEXIM1, LATS1, MYC, HRAS, TGFB1, IFNG, WNT1, TP53, THBS1, INHBA, IL8, IRF1, TPR, BMP2, BMP4, ETS1, HPGD, BMP7, GATA3, NR2F2, APC, PTPN3, CALR, IL12A, IL12B, PML, CDKN2B, CDKN2C, CDKN1B, SOX2, TAF6, DNA2, PLK1, TERF1, GAS1, CDKN2D, MLF1, PTEN, TGFB2, SMAD3, FOXO4, CDK6, TFAP4, MAP2K1, NOTCH2, FOXC1, DLG1, MAD2L1, ATM, NAE1, DGKZ, FHL1, SCRIB, BTG3, PTPRK, RPS6KA2, STK11, CDKN3, TBRG1, CDC73, THAP5, CRLF3, DCUN1D3, MYOCD, PAF1, LILRB1, UHMK1, PNPT1, USP47, HEXIM2, CDK5RAP1, NKX3-1, TIPIN, PCBP4, USP44, RBM38, CDT1, RGCC, RNF167, CLSPN, CHMP1A, WDR6, TCF7L2, LATS2, RASSF1, MLTK, MAD2L2, FBXO5, ING4, and TRIM35.
[0121] The term "inducing cell cycle inhibition" used herein refers to causing an accumulation or an arrest at any phase during cell division and/or duplication.
[0122] The present invention also provides methods of inducing tumor suppression by contacting a cell with a compound of the invention. Preferably, the compound is in an amount sufficient to increase expression of BIN1 or any tumor suppressors.
[0123] The term "inducing tumor suppression" may include, but is not limited to, a reduction in size of a tumor, a reduction in tumor volume, a decrease in number of tumors, a decrease in number of metastatic lesions in other tissues or organs distant from the primary tumor site, an increase in average survival time of a population of treated subjects in comparison to a population receiving carrier alone, an increase in average survival time of a population of treated subjects in comparison to a population of untreated subjects, an increase in average survival time of a population of treated subjects in comparison to a population receiving monotherapy with a drug that is not a compound of the present invention, a decrease in the mortality rate of a population of treated subjects in comparison to a population receiving carrier alone, a decrease in tumor growth rate, or a decrease in tumor regrowth rate.
[0124] The present invention also provides methods of inhibiting hedgehog signaling by contacting a cell with a compound of the invention. Preferably, the compound is in an amount sufficient to reduce expression of at least one gene selected from the group consisting of GLI1, PTCH1, SUFU, KIF7, GLI2, BMP4, MAP3K10, SHH, TCTN3, DYRK2, PTCHD1, and SMO.
[0125] The phrase "inhibiting hedgehog signaling" means the hedgehog signaling strength (intensity) with a compound treatment is reduced by at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 300%, 400%, 500%, 1000%, 1500%, or more compared to the hedgehog signaling strength (intensity) without any compound treatment.
[0126] The present invention also provides methods of inducing a gene expression by contacting a cell with a compound of the invention. Preferably, the compound is in an amount sufficient to induce neuronal differentiation, cell cycle inhibition and/or tumor suppression. Such gene is selected from the group consisting of CD133 (also called PROM1), DOCK4, PTPRK, PROM2, LHX1, LHX6, LHX9, PAX6, PAX7, VEFGA, FZD3B, FYN, HIF1A, HTRA2, EVX1, CCDC64, GFAP, CKDN1A, CDKN2A, MEN1, CHEK1, IRF6, ALOX15B, CYP27B1, DBC1, NME6, GMNN, HEXIM1, LATS1, MYC, HRAS, TGFB1, IFNG, WNT1, TP53, THBS1, INHBA, IL8, IRF1, TPR, BMP2, BMP4, ETS1, HPGD, BMP7, GATA3, NR2F2, APC, PTPN3, CALR, IL12A, IL12B, PML, CDKN2B, CDKN2C, CDKN1B, SOX2, TAF6, DNA2, PLK1, TERF1, GAS1, CDKN2D, MLF1, PTEN, TGFB2, SMAD3, FOXO4, CDK6, TFAP4, MAP2K1, NOTCH2, FOXC1, DLG1, MAD2L1, ATM, NAE1, DGKZ, FHL1, SCRIB, BTG3, PTPRK, RPS6KA2, STK11, CDKN3, TBRG1, CDC73, THAP5, CRLF3, DCUN1D3, MYOCD, PAF1, LILRB1, UHMK1, PNPT1, USP47, HEXIM2, CDK5RAP1, NKX3-1, TIPIN, PCBP4, USP44, RBM38, CDT1, RGCC, RNF167, CLSPN, CHMP1A, WDR6, TCF7L2, LATS2, RASSF1, MLTK, MAD2L2, FBXO5, ING4, TRIM35, BIN1 and any tumor suppressors.
[0127] The phrase "inducing a gene expression" means the expression level of a particular gene of interest is increased by at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 300%, 400%, 500%, 1000%, 1500%, or more compared to the expression level of this gene without any compound treatment.
[0128] The present invention also provides methods of inhibiting a gene expression comprising contacting a cell with a compound of the invention. Preferably, the compound is in an amount sufficient to inhibit hedgehog signaling. Such gene is GLI1, PTCH1, SUFU, KIF7, GLI2, BMP4, MAP3K10, SHH, TCTN3, DYRK2, PTCHD1, or SMO.
[0129] The phrase "inhibiting a gene expression" means the expression level of a particular gene of interest is reduced by at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 200%, 300%, 400%, 500%, 1000%, 1500%, or more compared to the expression level of this gene without any compound treatment.
[0130] Neuronal differentiation, cell cycle inhibition, tumor suppression and hedgehog signaling inhibition can be determined by any methods known in the art.
[0131] As used herein, a cell refers to any cell that can be obtained and used by a method described herein. For example, a cell may be obtained from a cell culture. Alternatively, a cell may be isolated from a subject. A cell may also refer to a cell of a subject.
[0132] A cell may comprise loss of function of SNF5, ARID1A, ATRX, and/or a component of the SWI/SNF complex. Preferably, a cell may comprise a deletion of SNF5.
[0133] A cell may be a cancer cell, where the cancer is selected from the group consisting of medulloblastoma, oligodendroglioma, ovarian clear cell adenocarcinoma, ovarian endomethrioid adenocarcinoma, ovarian serous adenocarcinoma, pancreatic ductal adenocarcinoma, pancreatic endocrine tumor, malignant rhabdoid tumor, astrocytoma, atypical teratoid rhabdoid tumor, choroid plexus carcinoma, choroid plexus papilloma, ependymoma, glioblastoma, meningioma, neuroglial tumor, oligoastrocytoma, oligodendroglioma, pineoblastoma, carcinosarcoma, chordoma, extragonadal germ cell tumor, extrarenal rhabdoid tumor, schwannoma, skin squamous cell carcinoma, chondrosarcoma, clear cell sarcoma of soft tissue, ewing sarcoma, gastrointestinal stromal tumor, osteosarcoma, rhabdomyosarcoma, epithelioid sarcoma, renal medullo carcinoma, diffuse large B-cell lymphoma, follicular lymphoma and not otherwise specified (NOS) sarcoma. More preferably a cell is a cancer cell of medulloblastoma, malignant rhabdoid tumor, or atypical teratoid rhabdoid tumor.
[0134] A cancer that is to be treated can be staged according to the American Joint Committee on Cancer (AJCC) TNM classification system, where the tumor (T) has been assigned a stage of TX, T1, T1mic, T1a, T1b, T1c, T2, T3, T4, T4a, T4b, T4c, or T4d; and where the regional lymph nodes (N) have been assigned a stage of NX, NO, N1, N2, N2a, N2b, N3, N3a, N3b, or N3c; and where distant metastasis (M) can be assigned a stage of MX, M0, or M1. A cancer that is to be treated can be staged according to an American Joint Committee on Cancer (AJCC) classification as Stage I, Stage IIA, Stage IIB, Stage IIIA, Stage IIIB, Stage IIIC, or Stage IV. A cancer that is to be treated can be assigned a grade according to an AJCC classification as Grade GX (e.g., grade cannot be assessed), Grade 1, Grade 2, Grade 3 or Grade 4. A cancer that is to be treated can be staged according to an AJCC pathologic classification (pN) of pNX, pN0, PN0 (I-), PN0 (I+), PN0 (mol-), PN0 (mol+), PN1, PN1(mi), PN1a, PN1b, PN1c, pN2, pN2a, pN2b, pN3, pN3a, pN3b, or pN3c.
[0135] A cancer that is to be treated can be evaluated by DNA cytometry, flow cytometry, or image cytometry. A cancer that is to be treated can be typed as having 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, or 90% of cells in the synthesis stage of cell division (e.g., in S phase of cell division). A cancer that is to be treated can be typed as having a low S-phase fraction or a high S-phase fraction.
[0136] As used herein, a "normal cell" is a cell that cannot be classified as part of a "cell proliferative disorder". A normal cell lacks unregulated or abnormal growth, or both, that can lead to the development of an unwanted condition or disease. Preferably, a normal cell possesses normally functioning cell cycle checkpoint control mechanisms.
[0137] As used herein, "contacting a cell" refers to a condition in which a compound or other composition of matter is in direct contact with a cell, or is close enough to induce a desired biological effect in a cell.
[0138] As used herein, "monotherapy" refers to the administration of a single active or therapeutic compound to a subject in need thereof. Preferably, monotherapy will involve administration of a therapeutically effective amount of an active compound. For example, cancer monotherapy with one of the compound of the present invention, or a pharmaceutically acceptable salt, polymorph, solvate, analog or derivative thereof, to a subject in need of treatment of cancer. Monotherapy may be contrasted with combination therapy, in which a combination of multiple active compounds is administered, preferably with each component of the combination present in a therapeutically effective amount. In one aspect, monotherapy with a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, is more effective than combination therapy in inducing a desired biological effect.
[0139] As used herein, "treating" or "treat" describes the management and care of a patient for the purpose of combating a disease, condition, or disorder and includes the administration of a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, to alleviate one or more symptoms or complications of a disease, condition or disorder, or to eliminate the disease, condition or disorder. The term "treat" can also include treatment of a cell in vitro or an animal model.
[0140] A compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, can also be used to prevent a disease, condition or disorder, or used to identify suitable candidates for such purposes. As used herein, "preventing" or "prevent" describes reducing or eliminating the onset of the symptoms or complications of the disease, condition or disorder.
[0141] As used herein, the term "alleviate" is meant to describe a process by which the severity of a sign or symptom of a disorder is decreased. Importantly, a sign or symptom can be alleviated without being eliminated. In a preferred embodiment, the administration of pharmaceutical compositions of the invention leads to the elimination of a sign or symptom, however, elimination is not required. Effective dosages are expected to decrease the severity of a sign or symptom. For instance, a sign or symptom of a disorder such as cancer, which can occur in multiple locations, is alleviated if the severity of the cancer is decreased within at least one of multiple locations.
[0142] As used herein, the term "severity" is meant to describe the potential of cancer to transform from a precancerous, or benign, state into a malignant state. Alternatively, or in addition, severity is meant to describe a cancer stage, for example, according to the TNM system (accepted by the International Union Against Cancer (UICC) and the American Joint Committee on Cancer (AJCC)) or by other art-recognized methods. Cancer stage refers to the extent or severity of the cancer, based on factors such as the location of the primary tumor, tumor size, number of tumors, and lymph node involvement (spread of cancer into lymph nodes). Alternatively, or in addition, severity is meant to describe the tumor grade by art-recognized methods (see, National Cancer Institute, www.cancer.gov). Tumor grade is a system used to classify cancer cells in terms of how abnormal they look under a microscope and how quickly the tumor is likely to grow and spread. Many factors are considered when determining tumor grade, including the structure and growth pattern of the cells. The specific factors used to determine tumor grade vary with each type of cancer. Severity also describes a histologic grade, also called differentiation, which refers to how much the tumor cells resemble normal cells of the same tissue type (see, National Cancer Institute, www.cancer.gov). Furthermore, severity describes a nuclear grade, which refers to the size and shape of the nucleus in tumor cells and the percentage of tumor cells that are dividing (see, National Cancer Institute, www.cancer.gov).
[0143] In another aspect of the invention, severity describes the degree to which a tumor has secreted growth factors, degraded the extracellular matrix, become vascularized, lost adhesion to juxtaposed tissues, or metastasized. Moreover, severity describes the number of locations to which a primary tumor has metastasized. Finally, severity includes the difficulty of treating tumors of varying types and locations. For example, inoperable tumors, those cancers which have greater access to multiple body systems (hematological and immunological tumors), and those which are the most resistant to traditional treatments are considered most severe. In these situations, prolonging the life expectancy of the subject and/or reducing pain, decreasing the proportion of cancerous cells or restricting cells to one system, and improving cancer stage/tumor grade/histological grade/nuclear grade are considered alleviating a sign or symptom of the cancer.
[0144] As used herein the term "symptom" is defined as an indication of disease, illness, injury, or that something is not right in the body. Symptoms are felt or noticed by the individual experiencing the symptom, but may not easily be noticed by others. Others are defined as non-health-care professionals.
[0145] As used herein the term "sign" is also defined as an indication that something is not right in the body. But signs are defined as things that can be seen by a doctor, nurse, or other health care professional.
[0146] Cancer is a group of diseases that may cause almost any sign or symptom. The signs and symptoms will depend on where the cancer is, the size of the cancer, and how much it affects the nearby organs or structures. If a cancer spreads (metastasizes), then symptoms may appear in different parts of the body.
[0147] Treating cancer can result in a reduction in size of a tumor. A reduction in size of a tumor may also be referred to as "tumor regression". Preferably, after treatment, tumor size is reduced by 5% or greater relative to its size prior to treatment; more preferably, tumor size is reduced by 10% or greater; more preferably, reduced by 20% or greater; more preferably, reduced by 30% or greater; more preferably, reduced by 40% or greater; even more preferably, reduced by 50% or greater; and most preferably, reduced by greater than 75% or greater. Size of a tumor may be measured by any reproducible means of measurement. The size of a tumor may be measured as a diameter of the tumor.
[0148] Treating cancer can result in a reduction in tumor volume. Preferably, after treatment, tumor volume is reduced by 5% or greater relative to its size prior to treatment; more preferably, tumor volume is reduced by 10% or greater; more preferably, reduced by 20% or greater; more preferably, reduced by 30% or greater; more preferably, reduced by 40% or greater; even more preferably, reduced by 50% or greater; and most preferably, reduced by greater than 75% or greater. Tumor volume may be measured by any reproducible means of measurement.
[0149] Treating cancer results in a decrease in number of tumors. Preferably, after treatment, tumor number is reduced by 5% or greater relative to number prior to treatment; more preferably, tumor number is reduced by 10% or greater; more preferably, reduced by 20% or greater; more preferably, reduced by 30% or greater; more preferably, reduced by 40% or greater; even more preferably, reduced by 50% or greater; and most preferably, reduced by greater than 75%. Number of tumors may be measured by any reproducible means of measurement. The number of tumors may be measured by counting tumors visible to the naked eye or at a specified magnification. Preferably, the specified magnification is 2.times., 3.times., 4.times., 5.times., 10.times., or 50.times..
[0150] Treating cancer can result in a decrease in number of metastatic lesions in other tissues or organs distant from the primary tumor site. Preferably, after treatment, the number of metastatic lesions is reduced by 5% or greater relative to number prior to treatment; more preferably, the number of metastatic lesions is reduced by 10% or greater; more preferably, reduced by 20% or greater; more preferably, reduced by 30% or greater; more preferably, reduced by 40% or greater; even more preferably, reduced by 50% or greater; and most preferably, reduced by greater than 75%. The number of metastatic lesions may be measured by any reproducible means of measurement. The number of metastatic lesions may be measured by counting metastatic lesions visible to the naked eye or at a specified magnification. Preferably, the specified magnification is 2.times., 3.times., 4.times., 5.times., 10.times., or 50.times..
[0151] Treating cancer can result in an increase in average survival time of a population of treated subjects in comparison to a population receiving carrier alone. Preferably, the average survival time is increased by more than 30 days; more preferably, by more than 60 days; more preferably, by more than 90 days; and most preferably, by more than 120 days. An increase in average survival time of a population may be measured by any reproducible means. An increase in average survival time of a population may be measured, for example, by calculating for a population the average length of survival following initiation of treatment with an active compound. An increase in average survival time of a population may also be measured, for example, by calculating for a population the average length of survival following completion of a first round of treatment with an active compound.
[0152] Treating cancer can result in an increase in average survival time of a population of treated subjects in comparison to a population of untreated subjects. Preferably, the average survival time is increased by more than 30 days; more preferably, by more than 60 days; more preferably, by more than 90 days; and most preferably, by more than 120 days. An increase in average survival time of a population may be measured by any reproducible means. An increase in average survival time of a population may be measured, for example, by calculating for a population the average length of survival following initiation of treatment with an active compound. An increase in average survival time of a population may also be measured, for example, by calculating for a population the average length of survival following completion of a first round of treatment with an active compound.
[0153] Treating cancer can result in increase in average survival time of a population of treated subjects in comparison to a population receiving monotherapy with a drug that is not a compound of the present invention, or a pharmaceutically acceptable salt, polymorph, solvate, analog or derivative thereof. Preferably, the average survival time is increased by more than 30 days; more preferably, by more than 60 days; more preferably, by more than 90 days; and most preferably, by more than 120 days. An increase in average survival time of a population may be measured by any reproducible means. An increase in average survival time of a population may be measured, for example, by calculating for a population the average length of survival following initiation of treatment with an active compound. An increase in average survival time of a population may also be measured, for example, by calculating for a population the average length of survival following completion of a first round of treatment with an active compound.
[0154] Treating cancer can result in a decrease in the mortality rate of a population of treated subjects in comparison to a population receiving carrier alone. Treating cancer can result in a decrease in the mortality rate of a population of treated subjects in comparison to an untreated population. Treating cancer can result in a decrease in the mortality rate of a population of treated subjects in comparison to a population receiving monotherapy with a drug that is not a compound of the present invention, or a pharmaceutically acceptable salt, polymorph, solvate, analog or derivative thereof. Preferably, the mortality rate is decreased by more than 2%; more preferably, by more than 5%; more preferably, by more than 10%; and most preferably, by more than 25%. A decrease in the mortality rate of a population of treated subjects may be measured by any reproducible means. A decrease in the mortality rate of a population may be measured, for example, by calculating for a population the average number of disease-related deaths per unit time following initiation of treatment with an active compound. A decrease in the mortality rate of a population may also be measured, for example, by calculating for a population the average number of disease-related deaths per unit time following completion of a first round of treatment with an active compound.
[0155] Treating cancer can result in a decrease in tumor growth rate. Preferably, after treatment, tumor growth rate is reduced by at least 5% relative to number prior to treatment; more preferably, tumor growth rate is reduced by at least 10%; more preferably, reduced by at least 20%; more preferably, reduced by at least 30%; more preferably, reduced by at least 40%; more preferably, reduced by at least 50%; even more preferably, reduced by at least 50%; and most preferably, reduced by at least 75%. Tumor growth rate may be measured by any reproducible means of measurement. Tumor growth rate can be measured according to a change in tumor diameter per unit time.
[0156] Treating cancer can result in a decrease in tumor regrowth. Preferably, after treatment, tumor regrowth is less than 5%; more preferably, tumor regrowth is less than 10%; more preferably, less than 20%; more preferably, less than 30%; more preferably, less than 40%; more preferably, less than 50%; even more preferably, less than 50%; and most preferably, less than 75%. Tumor regrowth may be measured by any reproducible means of measurement. Tumor regrowth is measured, for example, by measuring an increase in the diameter of a tumor after a prior tumor shrinkage that followed treatment. A decrease in tumor regrowth is indicated by failure of tumors to reoccur after treatment has stopped.
[0157] Treating cancer can result in a reduction in the rate of cellular proliferation. Preferably, after treatment, the rate of cellular proliferation is reduced by at least 5%; more preferably, by at least 10%; more preferably, by at least 20%; more preferably, by at least 30%; more preferably, by at least 40%; more preferably, by at least 50%; even more preferably, by at least 50%; and most preferably, by at least 75%. The rate of cellular proliferation may be measured by any reproducible means of measurement. The rate of cellular proliferation is measured, for example, by measuring the number of dividing cells in a tissue sample per unit time.
[0158] Treating cancer can result in a reduction in the proportion of proliferating cells. Preferably, after treatment, the proportion of proliferating cells is reduced by at least 5%; more preferably, by at least 10%; more preferably, by at least 20%; more preferably, by at least 30%; more preferably, by at least 40%; more preferably, by at least 50%; even more preferably, by at least 50%; and most preferably, by at least 75%. The proportion of proliferating cells may be measured by any reproducible means of measurement. Preferably, the proportion of proliferating cells is measured, for example, by quantifying the number of dividing cells relative to the number of nondividing cells in a tissue sample. The proportion of proliferating cells can be equivalent to the mitotic index.
[0159] Treating cancer can result in a decrease in size of an area or zone of cellular proliferation. Preferably, after treatment, size of an area or zone of cellular proliferation is reduced by at least 5% relative to its size prior to treatment; more preferably, reduced by at least 10%; more preferably, reduced by at least 20%; more preferably, reduced by at least 30%; more preferably, reduced by at least 40%; more preferably, reduced by at least 50%; even more preferably, reduced by at least 50%; and most preferably, reduced by at least 75%. Size of an area or zone of cellular proliferation may be measured by any reproducible means of measurement. The size of an area or zone of cellular proliferation may be measured as a diameter or width of an area or zone of cellular proliferation.
[0160] Treating cancer can result in a decrease in the number or proportion of cells having an abnormal appearance or morphology. Preferably, after treatment, the number of cells having an abnormal morphology is reduced by at least 5% relative to its size prior to treatment; more preferably, reduced by at least 10%; more preferably, reduced by at least 20%; more preferably, reduced by at least 30%; more preferably, reduced by at least 40%; more preferably, reduced by at least 50%; even more preferably, reduced by at least 50%; and most preferably, reduced by at least 75%. An abnormal cellular appearance or morphology may be measured by any reproducible means of measurement. An abnormal cellular morphology can be measured by microscopy, e.g., using an inverted tissue culture microscope. An abnormal cellular morphology can take the form of nuclear pleiomorphism.
[0161] Treating cancer can result in cell death, and preferably, cell death results in a decrease of at least 10% in number of cells in a population. More preferably, cell death means a decrease of at least 20%; more preferably, a decrease of at least 30%; more preferably, a decrease of at least 40%; more preferably, a decrease of at least 50%; most preferably, a decrease of at least 75%. Number of cells in a population may be measured by any reproducible means. A number of cells in a population can be measured by fluorescence activated cell sorting (FACS), immunofluorescence microscopy and light microscopy. Methods of measuring cell death are as shown in Li et al., Proc Natl Acad Sci USA. 100(5): 2674-8, 2003. In an aspect, cell death occurs by apoptosis.
[0162] As used herein, the term "selectively" means tending to occur at a higher frequency in one population than in another population. The compared populations can be cell populations. Preferably, a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, acts selectively on a cancer or precancerous cell but not on a normal cell. Preferably, a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, acts selectively to modulate one molecular target (e.g., a target protein methyltransferase) but does not significantly modulate another molecular target (e.g., a non-target protein methyltransferase). The invention also provides a method for selectively inhibiting the activity of an enzyme, such as a protein methyltransferase. Preferably, an event occurs selectively in population A relative to population B if it occurs greater than two times more frequently in population A as compared to population B. An event occurs selectively if it occurs greater than five times more frequently in population A. An event occurs selectively if it occurs greater than ten times more frequently in population A; more preferably, greater than fifty times; even more preferably, greater than 100 times; and most preferably, greater than 1000 times more frequently in population A as compared to population B. For example, cell death would be said to occur selectively in cancer cells if it occurred greater than twice as frequently in cancer cells as compared to normal cells.
[0163] A compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, can modulate the activity of a molecular target (e.g., a target protein methyltransferase). Modulating refers to stimulating or inhibiting an activity of a molecular target. Preferably, a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, modulates the activity of a molecular target if it stimulates or inhibits the activity of the molecular target by at least 2-fold relative to the activity of the molecular target under the same conditions but lacking only the presence of said compound. More preferably, a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, modulates the activity of a molecular target if it stimulates or inhibits the activity of the molecular target by at least 5-fold, at least 10-fold, at least 20-fold, at least 50-fold, at least 100-fold relative to the activity of the molecular target under the same conditions but lacking only the presence of said compound. The activity of a molecular target may be measured by any reproducible means. The activity of a molecular target may be measured in vitro or in vivo. For example, the activity of a molecular target may be measured in vitro by an enzymatic activity assay or a DNA binding assay, or the activity of a molecular target may be measured in vivo by assaying for expression of a reporter gene.
[0164] A compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, does not significantly modulate the activity of a molecular target if the addition of the compound does not stimulate or inhibit the activity of the molecular target by greater than 10% relative to the activity of the molecular target under the same conditions but lacking only the presence of said compound.
[0165] As used herein, the term "isozyme selective" means preferential inhibition or stimulation of a first isoform of an enzyme in comparison to a second isoform of an enzyme (e.g., preferential inhibition or stimulation of a protein methyltransferase isozyme alpha in comparison to a protein methyltransferase isozyme beta). Preferably, a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, demonstrates a minimum of a fourfold differential, preferably a tenfold differential, more preferably a fifty fold differential, in the dosage required to achieve a biological effect. Preferably, a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, demonstrates this differential across the range of inhibition, and the differential is exemplified at the IC.sub.50, i.e., a 50% inhibition, for a molecular target of interest.
[0166] Administering a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, to a cell or a subject in need thereof can result in modulation (i.e., stimulation or inhibition) of an activity of a protein methyltransferase of interest.
[0167] Detection of methylation of H3-K27, formation of trimethylated H3-K27, conversion of monomethylated H3-K27 to dimethylated H3-K27, or conversion of dimethylated H3-K27 to trimethylated H3-K27 can be accomplished using any suitable method. Exemplary methods can be found in US20120071418, the contents of which are incorporated herein by reference.
[0168] Administering a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, to a cell or a subject in need thereof results in modulation (i.e., stimulation or inhibition) of an activity of an intracellular target (e.g., substrate). Several intracellular targets can be modulated with the compounds of the present invention, including, but not limited to, protein methyltrasferase.
[0169] Preferably, an effective amount of a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, is not significantly cytotoxic to normal cells. A therapeutically effective amount of a compound is not significantly cytotoxic to normal cells if administration of the compound in a therapeutically effective amount does not induce cell death in greater than 10% of normal cells. A therapeutically effective amount of a compound does not significantly affect the viability of normal cells if administration of the compound in a therapeutically effective amount does not induce cell death in greater than 10% of normal cells. In an aspect, cell death occurs by apoptosis.
[0170] Contacting a cell with a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, can induce or activate cell death selectively in cancer cells. Administering to a subject in need thereof a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, can induce or activate cell death selectively in cancer cells. Contacting a cell with a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, can induce cell death selectively in one or more cells affected by a cell proliferative disorder. Preferably, administering to a subject in need thereof a compound of the present invention, or a pharmaceutically acceptable salt, polymorph or solvate thereof, induces cell death selectively in one or more cells affected by a cell proliferative disorder.
[0171] One skilled in the art may refer to general reference texts for detailed descriptions of known techniques discussed herein or equivalent techniques. These texts include Ausubel et al., Current Protocols in Molecular Biology, John Wiley and Sons, Inc. (2005); Sambrook et al., Molecular Cloning, A Laboratory Manual (3.sup.rd edition), Cold Spring Harbor Press, Cold Spring Harbor, N.Y. (2000); Coligan et al., Current Protocols in Immunology, John Wiley & Sons, N.Y.; Enna et al., Current Protocols in Pharmacology, John Wiley & Sons, N.Y.; Fingl et al., The Pharmacological Basis of Therapeutics (1975), Remington's Pharmaceutical Sciences, Mack Publishing Co., Easton, Pa., 18.sup.th edition (1990). These texts can, of course, also be referred to in making or using an aspect of the invention.
[0172] A compound (i.e., an EZH2 inhibitor) that can be used in any methods described herein may have the following Formula I:
##STR00006##
or a pharmaceutically acceptable salt thereof; wherein
[0173] R.sup.701 is H, F, OR.sup.707, NHR.sup.707, --(C.ident.C)--(CH.sub.2).sub.n7--R.sup.708, phenyl, 5- or 6-membered heteroaryl, C.sub.3-8 cycloalkyl, or 4-7 membered heterocycloalkyl containing 1-3 heteroatoms, wherein the phenyl, 5- or 6-membered heteroaryl, C.sub.3-8 cycloalkyl or 4-7 membered heterocycloalkyl each independently is optionally substituted with one or more groups selected from halo, C.sub.1-3 alkyl, OH, O--C.sub.1-6 alkyl, NH--C.sub.1-6 alkyl, and, C.sub.1-3 alkyl substituted with C.sub.3-8 cycloalkyl or 4-7 membered heterocycloalkyl containing 1-3 heteroatoms, wherein each of the O--C.sub.1-6 alkyl and NH--C.sub.1-6 alkyl is optionally substituted with hydroxyl, O--C.sub.1-3 alkyl or NH--C.sub.1-3 alkyl, each of the O--C.sub.1-3 alkyl and NH--C.sub.1-3 alkyl being optionally further substituted with O--C.sub.1-3 alkyl or NH--C.sub.1-3 alkyl;
[0174] each of R.sup.702 and R.sup.703, independently is H, halo, C.sub.1-4 alkyl, C.sub.1-6 alkoxyl or C.sub.6-C.sub.10 aryloxy, each optionally substituted with one or more halo;
[0175] each of R.sup.704 and R.sup.705, independently is C.sub.1-4 alkyl;
[0176] R.sup.706 is cyclohexyl substituted by N(C.sub.1-4 alkyl).sub.2 wherein one or both of the C.sub.1-4 alkyl is substituted with C.sub.1-6 alkoxy; or R.sup.706 is tetrahydropyranyl;
[0177] R.sup.707 is C.sub.1-4 alkyl optionally substituted with one or more groups selected from hydroxyl, C.sub.1-4 alkoxy, amino, mono- or di-C.sub.1-4 alkylamino, C.sub.3-8 cycloalkyl, and 4-7 membered heterocycloalkyl containing 1-3 heteroatoms, wherein the C.sub.3-8 cycloalkyl or 4-7 membered heterocycloalkyl each independently is further optionally substituted with C.sub.1-3 alkyl;
[0178] R.sup.708 is C.sub.1-4 alkyl optionally substituted with one or more groups selected from OH, halo, and C.sub.1-4 alkoxy, 4-7 membered heterocycloalkyl containing 1-3 heteroatoms, or O--C.sub.1-6 alkyl, wherein the 4-7 membered heterocycloalkyl can be optionally further substituted with OH or C.sub.1-6 alkyl; and
[0179] n 7 is 0, 1 or 2.
[0180] For example, R.sup.706 is cyclohexyl substituted by N(C.sub.1-4 alkyl).sub.2 wherein one of the C.sub.1-4 alkyl is unsubstituted and the other is substituted with methoxy.
[0181] For example, R.sup.706 is
##STR00007##
[0182] For example, the compound is of Formula II:
##STR00008##
[0183] For example, R.sup.702 is methyl or isopropyl and R.sup.703 is methyl or methoxyl.
[0184] For example, R.sup.704 is methyl.
[0185] For example, R.sup.701 is OR.sup.707 and R.sup.707 is C.sub.1-3 alkyl optionally substituted with OCH.sub.3 or morpholine.
[0186] For example, R.sup.701 is H or F.
[0187] For example, R.sup.701 is tetrahydropyranyl, phenyl, pyridyl, pyrimidyl, pyrazinyl, imidazolyl, or pyrazolyl, each of which is optionally substituted with methyl, methoxy, ethyl substituted with morpholine, or --OCH.sub.2CH.sub.2OCH.sub.3.
[0188] For example, R.sup.708 is morpholine, piperidine, piperazine, pyrrolidine, diazepane, or azetidine, each of which is optionally substituted with OH or C.sub.1-6 alkyl.
[0189] For example, R.sup.708 is morpholine
[0190] For example, R.sup.708 is piperazine substituted with C.sub.1-6 alkyl.
[0191] For example, R.sup.708 is methyl, t-butyl or C(CH.sub.3).sub.2OH.
[0192] A compound (i.e., an EZH2 inhibitor) that can be used in any methods described herein may have the following Formula III:
##STR00009##
or a pharmaceutically acceptable salt thereof.
[0193] In this formula:
[0194] R.sup.801 is C.sub.1-6 alkyl, C.sub.2-6 alkenyl, C.sub.2-6 alkynyl, C.sub.3-8 cycloalkyl, 4-7 membered heterocycloalkyl containing 1-3 heteroatoms, phenyl or 5- or 6-membered heteroaryl, each of which is substituted with O--C.sub.1-6 alkyl-R.sub.x or NH--C.sub.1-6 alkyl-R.sub.x, wherein R.sub.x is hydroxyl, O--C.sub.1-3 alkyl or NH--C.sub.1-3 alkyl, and R.sub.x is optionally further substituted with O--C.sub.1-3 alkyl or NH--C.sub.1-3 alkyl except when R.sub.x is hydroxyl; or R.sup.801 is phenyl substituted with -Q.sub.2-T.sub.2, wherein Q.sub.2 is a bond or C.sub.1-C.sub.3 alkyl linker optionally substituted with halo, cyano, hydroxyl or C.sub.1-C.sub.6 alkoxy, and T.sub.2 is optionally substituted 4- to 12-membered heterocycloalkyl; and R.sup.801 is optionally further substituted;
[0195] each of R.sup.802 and R.sup.803, independently is H, halo, C.sub.1-4 alkyl, C.sub.1-6 alkoxyl or C.sub.6-C.sub.10 aryloxy, each optionally substituted with one or more halo;
[0196] each of R.sup.804 and R.sup.805, independently is C.sub.1-4 alkyl; and
[0197] R.sup.806 is -Q.sub.x-T.sub.x, wherein Q.sub.x is a bond or C.sub.1-4 alkyl linker, T.sub.x is H, optionally substituted C.sub.1-4 alkyl, optionally substituted C.sub.3-C.sub.8 cycloalkyl or optionally substituted 4- to 14-membered heterocycloalkyl.
[0198] For example, each of Q.sub.x and Q.sub.2 independently is a bond or methyl linker, and each of T.sub.x and T.sub.2 independently is tetrahydropyranyl, piperidinyl substituted by 1, 2, or 3 C.sub.1-4 alkyl groups, or cyclohexyl substituted by N(C.sub.1-4 alkyl).sub.2 wherein one or both of the C.sub.1-4 alkyl is optionally substituted with C.sub.1-6 alkoxy;
[0199] For example, R.sup.806 is cyclohexyl substituted by N(C.sub.1-4 alkyl).sub.2 or R.sup.806 is tetrahydropyranyl.
##STR00010##
[0200] For example, R.sup.806 is
[0201] For example, R.sup.801 is phenyl or 5- or 6-membered heteroaryl substituted with O--C.sub.1-6 alkyl-R.sub.x, or R.sup.801 is phenyl substituted with CH.sub.2-tetrahydropyranyl.
[0202] For example, a compound of the present invention is of Formula IVa or IVb:
##STR00011##
wherein Z' is CH or N, and R.sup.807 is C.sub.2-3 alkyl-R.sub.x.
[0203] For example, R.sup.807 is --CH.sub.2CH.sub.2OH, --CH.sub.2CH.sub.2OCH.sub.3, or --CH.sub.2CH.sub.2OCH.sub.2CH.sub.2OCH.sub.3.
[0204] For example, R.sup.802 is methyl or isopropyl and R.sup.803 is methyl or methoxyl.
[0205] For example, R.sup.804 is methyl.
[0206] A compound of the present invention may have the following Formula (V):
##STR00012##
or a pharmaceutically acceptable salt or ester thereof.
[0207] In this formula:
[0208] R.sub.2, R.sub.4 and R.sub.12 are each independently C.sub.1-6 alkyl;
[0209] R.sub.6 is C.sub.6-C.sub.10 aryl or 5- or 6-membered heteroaryl, each of which is optionally substituted with one or more -Q.sub.2-T.sub.2, wherein Q.sub.2 is a bond or C.sub.1-C.sub.3 alkyl linker optionally substituted with halo, cyano, hydroxyl or C.sub.1-C.sub.6 alkoxy, and T2 is H, halo, cyano, --OR.sub.a, --NR.sub.aR.sub.b, --(NR.sub.aR.sub.bR.sub.c).sup.+A.sup.-, --C(O)R.sub.a, --C(O)OR.sub.a, --C(O)NR.sub.aR.sub.b, --NR.sub.bC(O)R.sub.a, --NR.sub.bC(O)OR.sub.a, --S(O).sub.2R.sub.a, --S(O).sub.2NR.sub.aR.sub.b, or R.sub.S2, in which each of R.sub.a, R.sub.b, and R.sub.c, independently is H or R.sub.S3, A.sup.- is a pharmaceutically acceptable anion, each of R.sub.S2 and R.sub.S3, independently, is C.sub.1-C.sub.6 alkyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, or R.sub.a and R.sub.b, together with the N atom to which they are attached, form a 4 to 12-membered heterocycloalkyl ring having 0 or 1 additional heteroatom, and each of R.sub.S2, R.sub.S3, and the 4 to 12-membered heterocycloalkyl ring formed by R.sub.a and R.sub.b, is optionally substituted with one or more -Q.sub.3-T.sub.3, wherein Q.sub.3 is a bond or C.sub.1-C.sub.3 alkyl linker each optionally substituted with halo, cyano, hydroxyl or C.sub.1-C.sub.6 alkoxy, and T.sub.3 is selected from the group consisting of halo, cyano, C.sub.1-C.sub.6 alkyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, 5- or 6-membered heteroaryl, OR.sub.d, COOR.sub.d, --S(O).sub.2R.sub.d, --NR.sub.dR.sub.e, and --C(O)NR.sub.dR.sub.e, each of R.sub.d and R.sub.e independently being H or C.sub.1-C.sub.6 alkyl, or -Q.sub.3-T.sub.3 is oxo; or any two neighboring -Q.sub.2-T.sub.2, together with the atoms to which they are attached form a 5- or 6-membered ring optionally containing 1-4 heteroatoms selected from N, O and S and optionally substituted with one or more substituents selected from the group consisting of halo, hydroxyl, COOH, C(O)O--C.sub.1-C.sub.6 alkyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, and 5- or 6-membered heteroaryl;
[0210] R.sub.7 is -Q.sub.4-T.sub.4, in which Q.sub.4 is a bond, C.sub.1-C.sub.4 alkyl linker, or C.sub.2-C.sub.4 alkenyl linker, each linker optionally substituted with halo, cyano, hydroxyl or C.sub.1-C.sub.6 alkoxy, and T.sub.4 is H, halo, cyano, NR.sub.fR.sub.g, --OR.sub.f, --C(O)R.sub.f, --C(O)OR.sub.f, --C(O)NR.sub.fR.sub.g, --C(O)NR.sub.fOR.sub.g, --NR.sub.fC(O)R.sub.g, --S(O).sub.2R.sub.f, or R.sub.S4, in which each of R.sub.f and R.sub.g, independently is H or R.sub.S5, each of R.sub.S4 and R.sub.S5, independently is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, and each of R.sub.S4 and R.sub.S5 is optionally substituted with one or more -Q.sub.5-T.sub.5, wherein Q.sub.5 is a bond, C(O), C(O)NR.sub.k, NR.sub.kC(O), S(O).sub.2, or C.sub.1-C.sub.3 alkyl linker, R.sub.k being H or C.sub.1-C.sub.6 alkyl, and T.sub.5 is H, halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, 5- or 6-membered heteroaryl, or S(O).sub.qR.sub.q in which q is 0, 1, or 2 and R.sub.q is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, and T.sub.5 is optionally substituted with one or more substituents selected from the group consisting of halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, and 5- or 6-membered heteroaryl except when T.sub.5 is H, halo, hydroxyl, or cyano; or -Q.sub.5-T.sub.5 is oxo; and
[0211] R.sub.8 is H, halo, hydroxyl, COOH, cyano, R.sub.S6, OR.sub.S6, or COOR.sub.S6, in which R.sub.S6 is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, C.sub.3-C.sub.8 cycloalkyl, 4 to 12-membered heterocycloalkyl, amino, mono-C.sub.1-C.sub.6 alkylamino, or di-C.sub.1-C.sub.6 alkylamino, and R.sub.S6 is optionally substituted with one or more substituents selected from the group consisting of halo, hydroxyl, COOH, C(O)O--C.sub.1-C.sub.6 alkyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, and di-C.sub.1-C.sub.6 alkylamino; or R.sub.7 and R.sub.8, together with the N atom to which they are attached, form a 4 to 11-membered heterocycloalkyl ring having 0 to 2 additional heteroatoms, and the 4 to 11-membered heterocycloalkyl ring formed by R.sub.7 and R.sub.8 is optionally substituted with one or more -Q.sub.6-T.sub.6, wherein Q.sub.6 is a bond, C(O), C(O)NR.sub.m, NR.sub.mC(O), S(O).sub.2, or C.sub.1-C.sub.3 alkyl linker, R.sub.m being H or C.sub.1-C.sub.6 alkyl, and T6 is H, halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, 5- or 6-membered heteroaryl, or S(O).sub.pR.sub.p in which p is 0, 1, or 2 and R.sub.p is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, and T6 is optionally substituted with one or more substituents selected from the group consisting of halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, and 5- or 6-membered heteroaryl except when T6 is H, halo, hydroxyl, or cyano; or -Q.sub.6-T.sub.6 is oxo.
[0212] For example, R6 is C.sub.6-C.sub.10 aryl or 5- or 6-membered heteroaryl, each of which is optionally, independently substituted with one or more -Q.sub.2-T.sub.2, wherein Q.sub.2 is a bond or C.sub.1-C.sub.3 alkyl linker, and T.sub.2 is H, halo, cyano, --OR.sub.a, --NR.sub.aR.sub.b,
--(NR.sub.aR.sub.bR.sub.c).sup.+A.sup.-, --C(O)NR.sub.aR.sub.b, --NR.sub.bC(O)R.sub.a, --S(O).sub.2R.sub.a, or R.sub.S2, in which each of R.sub.a and R.sub.b, independently is H or R.sub.S3, each of R.sub.S2 and R.sub.S3, independently, is C.sub.1-C.sub.6 alkyl, or R.sub.a and R.sub.b, together with the N atom to which they are attached, form a 4 to 7-membered heterocycloalkyl ring having 0 or 1 additional heteroatom, and each of R.sub.S2, R.sub.S3, and the 4 to 7-membered heterocycloalkyl ring formed by R.sub.a and R.sub.b, is optionally, independently substituted with one or more -Q.sub.3-T.sub.3, wherein Q.sub.3 is a bond or C.sub.1-C.sub.3 alkyl linker and T.sub.3 is selected from the group consisting of halo, C.sub.1-C.sub.6 alkyl, 4 to 7-membered heterocycloalkyl, OR.sub.d, --S(O).sub.2R.sub.d, and --NR.sub.dR.sub.e, each of R.sub.d and R.sub.e independently being H or C.sub.1-C.sub.6 alkyl, or -Q.sub.3-T.sub.3 is oxo; or any two neighboring -Q.sub.2-T.sub.2, together with the atoms to which they are attached form a 5- or 6-membered ring optionally containing 1-4 heteroatoms selected from N, O and S.
[0213] For example, the compound of the present invention is of Formula (VI):
##STR00013##
or a pharmaceutically acceptable salt thereof, wherein Q.sub.2 is a bond or methyl linker, T.sub.2 is H, halo, --OR.sub.a, --NR.sub.aR.sub.b, --(NR.sub.aR.sub.bR.sub.c).sup.+A.sup.-, or --S(O).sub.2NR.sub.aR.sub.b, R.sub.7 is piperidinyl, tetrahydropyran, cyclopentyl, or cyclohexyl, each optionally substituted with one -Q.sub.5-T.sub.5 and R.sub.8 is ethyl.
[0214] A compound of the present invention may have the following Formula (VIa):
##STR00014##
wherein
[0215] each of R.sub.a and R.sub.b, independently is H or R.sub.S3, R.sub.S3 being C.sub.1-C.sub.6 alkyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, or R.sub.a and R.sub.b, together with the N atom to which they are attached, form a 4 to 12-membered heterocycloalkyl ring having 0 or 1 additional heteroatom, and each of R.sub.S3 and the 4 to 12-membered heterocycloalkyl ring formed by R.sub.a and R.sub.b, is optionally substituted with one or more -Q.sub.3-T.sub.3, wherein Q.sub.3 is a bond or C.sub.1-C.sub.3 alkyl linker each optionally substituted with halo, cyano, hydroxyl or C.sub.1-C.sub.6 alkoxy, and T.sub.3 is selected from the group consisting of halo, cyano, C.sub.1-C.sub.6 alkyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 12-membered heterocycloalkyl, 5- or 6-membered heteroaryl, OR.sub.d, COOR.sub.d, --S(O).sub.2R.sub.d, --NR.sub.dR.sub.e, and --C(O)NR.sub.dR.sub.e, each of R.sub.d and R.sub.e independently being H or C.sub.1-C.sub.6 alkyl, or -Q.sub.3-T.sub.3 is oxo;
[0216] R.sub.7 is -Q.sub.4-T.sub.4, in which Q.sub.4 is a bond, C.sub.1-C.sub.4 alkyl linker, or C.sub.2-C.sub.4 alkenyl linker, each linker optionally substituted with halo, cyano, hydroxyl or C.sub.1-C.sub.6 alkoxy, and T.sub.4 is H, halo, cyano, NR.sub.fR.sub.g, --OR.sub.f, --C(O)R.sub.f, --C(O)OR.sub.f, --C(O)NR.sub.fR.sub.g, --C(O)NR.sub.fOR.sub.g, --NR.sub.fC(O)R.sub.g, --S(O).sub.2R.sub.f, or R.sub.S4, in which each of R.sub.f and R.sub.g, independently is H or R.sub.S5, each of R.sub.S4 and R.sub.S5, independently is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 7-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, and each of R.sub.S4 and R.sub.S5 is optionally substituted with one or more -Q.sub.5-T.sub.5, wherein Q.sub.5 is a bond, C(O), C(O)NR.sub.k, NR.sub.kC(O), S(O).sub.2, or C.sub.1-C.sub.3 alkyl linker, R.sub.k being H or C.sub.1-C.sub.6 alkyl, and T.sub.5 is H, halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 7-membered heterocycloalkyl, 5- or 6-membered heteroaryl, or S(O).sub.qR.sub.q in which q is 0, 1, or 2 and R.sub.q is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, C.sub.3-C.sub.8cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 7-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, and T.sub.5 is optionally substituted with one or more substituents selected from the group consisting of halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 7-membered heterocycloalkyl, and 5- or 6-membered heteroaryl except when T.sub.5 is H, halo, hydroxyl, or cyano; or -Q.sub.5-T.sub.5 is oxo; provided that R.sub.7 is not H; and
[0217] R.sub.8 is H, halo, hydroxyl, COOH, cyano, R.sub.S6, OR.sub.S6, or COOR.sub.S6, in which R.sub.S6 is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, amino, mono-C.sub.1-C.sub.6 alkylamino, or di-C.sub.1-C.sub.6 alkylamino, and R.sub.S6 is optionally substituted with one or more substituents selected from the group consisting of halo, hydroxyl, COOH, C(O)O--C.sub.1-C.sub.6 alkyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, and di-C.sub.1-C.sub.6 alkylamino; or R.sub.7 and R.sub.8, together with the N atom to which they are attached, form a 4 to 11-membered heterocycloalkyl ring which has 0 to 2 additional heteroatoms and is optionally substituted with one or more -Q.sub.6-T.sub.6, wherein Q.sub.6 is a bond, C(O), C(O)NR.sub.m, NR.sub.mC(O), S(O).sub.2, or C.sub.1-C.sub.3 alkyl linker, R.sub.m being H or C.sub.1-C.sub.6 alkyl, and T.sub.6 is H, halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 7-membered heterocycloalkyl, 5- or 6-membered heteroaryl, or S(O).sub.pR.sub.p in which p is 0, 1, or 2 and R.sub.p is C.sub.1-C.sub.6 alkyl, C.sub.2-C.sub.6 alkenyl, C.sub.2-C.sub.6 alkynyl, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 7-membered heterocycloalkyl, or 5- or 6-membered heteroaryl, and T.sub.6 is optionally substituted with one or more substituents selected from the group consisting of halo, C.sub.1-C.sub.6 alkyl, hydroxyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, di-C.sub.1-C.sub.6 alkylamino, C.sub.3-C.sub.8 cycloalkyl, C.sub.6-C.sub.10 aryl, 4 to 7-membered heterocycloalkyl, and 5- or 6-membered heteroaryl except when T6 is H, halo, hydroxyl, or cyano; or -Q6-T6 is oxo.
[0218] For example, R.sub.a and R.sub.b, together with the N atom to which they are attached, form a 4 to 7-membered heterocycloalkyl ring having 0 or 1 additional heteroatoms to the N atom and the ring is optionally substituted with one or more -Q.sub.3-T.sub.3, wherein the heterocycloalkyl is azetidinyl, pyrrolidinyl, imidazolidinyl, pyrazolidinyl, oxazolidinyl, isoxazolidinyl, triazolidinyl, piperidinyl, 1,2,3,6-tetrahydropyridinyl, piperazinyl, or morpholinyl.
[0219] For example, R.sub.7 is C.sub.3-C.sub.8 cycloalkyl or 4 to 7-membered heterocycloalkyl, each optionally substituted with one or more -Q.sub.5-T.sub.5.
[0220] For example, R.sub.7 is piperidinyl, tetrahydropyran, tetrahydro-2H-thiopyranyl, cyclopentyl, cyclohexyl, pyrrolidinyl, or cycloheptyl, each optionally substituted with one or more -Q.sub.5-T.sub.5.
[0221] For example, R.sub.8 is H or C.sub.1-C.sub.6 alkyl which is optionally substituted with one or more substituents selected from the group consisting of halo, hydroxyl, COOH, C(O)O--C.sub.1-C.sub.6 alkyl, cyano, C.sub.1-C.sub.6 alkoxyl, amino, mono-C.sub.1-C.sub.6 alkylamino, and di-C.sub.1-C.sub.6 alkylamino.
[0222] In some embodiments, a compound that can be used in any methods presented here is:
##STR00015##
stereoisomers thereof or pharmaceutically acceptable salt or solvate thereof.
[0223] In some embodiments, a compound that can be used in any methods presented here is:
##STR00016## ##STR00017##
stereoisomers thereof or pharmaceutically acceptable salts and solvates thereof.
[0224] In some embodiments, a compound that can be used in any methods presented here is:
##STR00018##
stereoisomers thereof or pharmaceutically acceptable salts and solvates thereof.
[0225] In some embodiments, the compounds suitable for use in the method of this invention include compounds of Formula (VII):
##STR00019##
wherein,
[0226] V.sup.1 is N or CR.sup.7,
[0227] V.sup.2 is N or CR.sup.2, provided when V.sup.1 is N, V.sup.2 is N,
[0228] X and Z are selected independently from the group consisting of hydrogen, (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, unsubstituted or substituted (C.sub.3-C.sub.8)cycloalkyl, unsubstituted or substituted (C.sub.3-C.sub.8)cycloalkyl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, unsubstituted or substituted (C.sub.5-C.sub.8)cycloalkenyl, unsubstituted or substituted (C.sub.5-C.sub.8)cycloalkenyl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, (C.sub.6-C.sub.10)bicycloalkyl, unsubstituted or substituted heterocycloalkyl, unsubstituted or substituted heterocycloalkyl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, unsubstituted or substituted aryl, unsubstituted or substituted aryl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, unsubstituted or substituted heteroaryl, unsubstituted or substituted heteroaryl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, halo, cyano,
--COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --CONR.sup.aNR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --NR.sup.aNR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)OR.sup.a, --OR.sup.a, --OC(O)R.sup.a, and --OC(O)NR.sup.aR.sup.b;
[0229] Y is H or halo;
[0230] R.sup.1 is (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, unsubstituted or substituted (C.sub.3-C.sub.8)cycloalkyl, unsubstituted or substituted (C.sub.3-C.sub.8)cycloalkyl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, unsubstituted or substituted (C.sub.5-C.sub.8)cycloalkenyl, unsubstituted or substituted (C.sub.5-C.sub.8)cycloalkenyl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, unsubstituted or substituted (C.sub.6-C.sub.10)bicycloalkyl, unsubstituted or substituted heterocycloalkyl or --(C.sub.2-C.sub.8)alkenyl, unsubstituted or substituted heterocycloalkyl-(C.sub.1-C.sub.8)alkyl, unsubstituted or substituted aryl, unsubstituted or substituted aryl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, unsubstituted or substituted heteroaryl, unsubstituted or substituted heteroaryl-(C.sub.1-C.sub.8)alkyl or --(C.sub.2-C.sub.8)alkenyl, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --CONR.sup.aNR.sup.aR.sup.b;
[0231] R.sup.2 is hydrogen, (C.sub.1-C.sub.8)alkyl, trifluoromethyl, alkoxy, or halo, in which said (C.sub.1-C.sub.8)alkyl is optionally substituted with one to two groups selected from amino and (C.sub.1-C.sub.3)alkylamino;
[0232] R.sup.7 is hydrogen, (C.sub.1-C.sub.3)alkyl, or alkoxy;
[0233] R.sup.3 is hydrogen, (C.sub.1-C.sub.8)alkyl, cyano, trifluoromethyl, --NR.sup.aR.sup.b, or halo;
[0234] R.sup.6 is selected from the group consisting of hydrogen, halo, (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, unsubstituted or substituted (C.sub.3-C.sub.8)cycloalkyl, unsubstituted or substituted (C.sub.3-C.sub.8)cycloalkyl-(C.sub.1-C.sub.8)alkyl, unsubstituted or substituted (C.sub.5-C.sub.8)cycloalkenyl, unsubstituted or substituted (C.sub.5-C.sub.8)cycloalkenyl(C.sub.1-C.sub.8)alkyl, (C.sub.6-C.sub.10)bicycloalkyl, unsubstituted or substituted heterocycloalkyl, unsubstituted or substituted heterocycloalkyl-(C.sub.1-C.sub.8)alkyl, unsubstituted or substituted aryl, unsubstituted or substituted aryl-(C.sub.1-C.sub.8)alkyl, unsubstituted or substituted heteroaryl, unsubstituted or substituted heteroaryl-(C.sub.1-C.sub.8)alkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a,
--CONR.sup.aR.sup.b, --CONR.sup.aNR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --NR.sup.aNR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)OR.sup.a, --OR.sup.a, --OC(O)R.sup.a, --OC(O)NR.sup.aR.sup.b;
[0235] wherein any (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, cycloalkyl, cycloalkenyl, bicycloalkyl, heterocycloalkyl, aryl, or heteroaryl group is optionally substituted by 1, 2 or 3 groups independently selected from the group consisting of --O(C.sub.1-C.sub.6)alkyl(R.sup.c).sub.1-2, --S(C.sub.1-C.sub.6)alkyl(R.sup.c).sub.1-2, --(C.sub.1-C.sub.6)alkyl(R.sup.c).sub.1-2, --(C.sub.1-C.sub.8)alkyl-heterocycloalkyl, (C.sub.3-C.sub.8)cycloalkyl-heterocycloalkyl, halo, (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, OC(O)NR.sup.aR.sup.b, heterocycloalkyl, aryl, heteroaryl, aryl(C.sub.1-C.sub.4)alkyl, and heteroaryl(C.sub.1-C.sub.4)alkyl; [0236] wherein any aryl or heteroaryl moiety of said aryl, heteroaryl, aryl(C.sub.1-C.sub.4)alkyl, or heteroaryl(C.sub.1-C.sub.4)alkyl is optionally substituted by 1, 2 or 3 groups independently selected from the group consisting of halo, (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, [0237] SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, and --OC(O)NR.sup.aR.sup.b;
[0238] R.sup.a and R.sup.b are each independently hydrogen, (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.6-C.sub.10)bicycloalkyl, heterocycloalkyl, aryl, or heteroaryl, wherein said (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, cycloalkyl, cycloalkenyl, bicycloalkyl, heterocycloalkyl, aryl or heteroaryl group is optionally substituted by 1, 2 or 3 groups independently selected from halo, hydroxyl, (C.sub.1-C.sub.4)alkoxy, amino, (C.sub.1-C.sub.4)alkylamino, ((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl)amino, --CO.sub.2H, --CO.sub.2(C.sub.1-C.sub.4)alkyl, --CONH.sub.2, --CONH(C.sub.1-C.sub.4)alkyl,
--CON((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl), --SO.sub.2(C.sub.1-C.sub.4)alkyl, --SO.sub.2NH.sub.2, --SO.sub.2NH(C.sub.1-C.sub.4)alkyl, and SO.sub.2N((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl);
[0239] or R.sup.a and R.sup.b taken together with the nitrogen to which they are attached represent a 5-8 membered saturated or unsaturated ring, optionally containing an additional heteroatom selected from oxygen, nitrogen, and sulfur, wherein said ring is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.4)alkyl, (C.sub.1-C.sub.4)haloalkyl, amino, (C.sub.1-C.sub.4)alkylamino, ((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl)amino, hydroxyl, oxo, (C.sub.1-C.sub.4)alkoxy, and (C.sub.1-C.sub.4)alkoxy(C.sub.1-C.sub.4)alkyl, wherein said ring is optionally fused to a (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, or heteroaryl ring;
[0240] or R.sup.a and R.sup.b taken together with the nitrogen to which they are attached represent a 6- to 10-membered bridged bicyclic ring system optionally fused to a (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, or heteroaryl ring;
[0241] each R.sup.c is independently (C.sub.1-C.sub.4)alkylamino, --NR.sup.aSO.sub.2R.sup.b, --SOR.sup.a, --SO.sub.2R.sup.a, --NR.sup.aC(O)OR.sup.a, --NR.sup.aR.sup.b, or --CO.sub.2R.sup.a;
[0242] or a salt thereof.
[0243] Subgroups of the compounds encompassed by the general structure of Formula (I) are represented as follows:
[0244] Subgroup A of Formula (VII)
[0245] X and Z are selected from the group consisting of (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, heteroaryl, --NR.sup.aR.sup.b, and --OR.sup.a;
[0246] Y is H or F;
[0247] R.sup.1 is selected from the group consisting of (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, and heteroaryl;
[0248] R.sup.2 is hydrogen, (C.sub.1-C.sub.8)alkyl, trifluoromethyl, alkoxy, or halo, in which said (C.sub.1-C.sub.8)alkyl is optionally substituted with one to two groups selected from amino and (C.sub.1-C.sub.3)alkylamino;
[0249] R.sup.7 is hydrogen, (C.sub.1-C.sub.3)alkyl, or alkoxy;
[0250] R.sup.3 is selected from the group consisting of hydrogen, (C.sub.1-C.sub.8)alkyl, cyano, trifluoromethyl, --NR.sup.aR.sup.b, and halo;
[0251] R.sup.6 is selected from the group consisting of hydrogen, halo, cyano, trifluoromethyl, amino, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl; aryl, heteroaryl, acylamino; (C.sub.2-C.sub.8)alkynyl, arylalkynyl, heteroarylalkynyl; --SO.sub.2R.sup.a; --SO.sub.2NR.sup.aR.sup.b and --NR.sup.aSO.sub.2R.sup.b; [0252] wherein any (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.2-C.sub.8)alkynyl, arylalkynyl, heteroarylalkynyl group is optionally substituted by 1, 2 or 3 groups independently selected from --O(C.sub.1-C.sub.6)alkyl(R.sup.c).sub.1-2, --S(C.sub.1-C.sub.6)alkyl(R.sup.c).sub.1-2, --(C.sub.1-C.sub.6)alkyl(R.sup.c).sub.1-2, --(C.sub.1-C.sub.8)alkyl-heterocycloalkyl, (C.sub.3-C.sub.8)cycloalkyl-heterocycloalkyl, halo, (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, --OC(O)NR.sup.aR.sup.b, heterocycloalkyl, aryl, heteroaryl, aryl(C.sub.1-C.sub.4)alkyl, and heteroaryl(C.sub.1-C.sub.4)alkyl;
[0253] R.sup.a and R.sup.b are each independently hydrogen, (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.6-C.sub.10)bicycloalkyl, heterocycloalkyl, aryl, or heteroaryl, wherein said (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, cycloalkyl, cycloalkenyl, bicycloalkyl, heterocycloalkyl, aryl or heteroaryl group is optionally substituted by 1, 2 or 3 groups independently selected from halo, hydroxyl, (C.sub.1-C.sub.4)alkoxy, amino, (C.sub.1-C.sub.4)alkylamino, ((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl)amino, --CO.sub.2H, --CO.sub.2(C.sub.1-C.sub.4)alkyl, --CONH.sub.2, --CONH(C.sub.1-C.sub.4)alkyl, --CON((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl), --SO.sub.2(C.sub.1-C.sub.4)alkyl, --SO.sub.2NH.sub.2, --SO.sub.2NH(C.sub.1-C.sub.4)alkyl, and
--SO.sub.2N((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl);
[0254] or R.sup.a and R.sup.b taken together with the nitrogen to which they are attached represent a 5-8 membered saturated or unsaturated ring, optionally containing an additional heteroatom selected from oxygen, nitrogen, and sulfur, wherein said ring is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.4)alkyl, (C.sub.1-C.sub.4)haloalkyl, amino, (C.sub.1-C.sub.4)alkylamino, ((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl)amino, hydroxyl, oxo, (C.sub.1-C.sub.4)alkoxy, and (C.sub.1-C.sub.4)alkoxy(C.sub.1-C.sub.4)alkyl, wherein said ring is optionally fused to a (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, or heteroaryl ring;
[0255] or R.sup.a and R.sup.b taken together with the nitrogen to which they are attached represent a 6- to 10-membered bridged bicyclic ring system optionally fused to a (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, or heteroaryl ring. An aryl or heteroaryl group in this particular subgroup A is selected independently from the group consisting of furan, thiophene, pyrrole, oxazole, thiazole, imidazole, pyrazole, oxadiazole, thiadiazole, triazole, tetrazole, benzofuran, benzothiophene, benzoxazole, benzothiazole, phenyl, pyridine, pyridazine, pyrimidine, pyrazine, triazine, tetrazine, quinoline, cinnoline, quinazoline, quinoxaline, and naphthyridine or another aryl or heteroaryl group as follows:
##STR00020##
wherein in (1),
[0256] A is O, NH, or S; B is CH or N, and C is hydrogen or C.sub.1-C.sub.8 alkyl; or
##STR00021##
wherein in (2),
[0257] D is N or C optionally substituted by hydrogen or C.sub.1-C.sub.8 alkyl; or
##STR00022##
wherein in (3),
[0258] E is NH or CH.sub.2; F is O or CO; and G is NH or CH.sub.2; or
##STR00023##
wherein in (4), [0259] J is O, S or CO; or
##STR00024##
[0259] wherein in (5),
[0260] Q is CH or N;
[0261] M is CH or N; and
[0262] L/(5) is hydrogen, halo, amino, cyano, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --CONR.sup.aNR.sup.aR.sup.b, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --NR.sup.aNR.sup.aC(O)NR.sup.aR.sup.b, or --OR.sup.a, [0263] wherein any (C.sub.1-C.sub.8)alkyl or (C.sub.3-C.sub.8)cycloalkyl group is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, and --OC(O)NR.sup.aR.sup.b; wherein R.sup.a and R.sup.b are defined as above; or
##STR00025##
[0263] wherein in (6),
[0264] L/(6) is NH or CH.sub.2; or
##STR00026##
wherein in 7, [0265] M/(7) is hydrogen, halo, amino, cyano, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --CONR.sup.aNR.sup.aR.sup.b, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --NR.sup.aNR.sup.aC(O)NR.sup.aR.sup.b, or --OR.sup.a, [0266] wherein any (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, or heterocycloalkyl group is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, and --OC(O)NR.sup.aR.sup.b; wherein R.sup.a and R.sup.b are defined as above; or
##STR00027##
[0266] wherein in (8),
[0267] P is CH.sub.2, NH, O, or S; Q/(8) is CH or N; and n is 0-2; or
##STR00028##
wherein in (9),
[0268] S/(9) and T/(9) is C, or S/(9) is C and T/(9) is N, or S/(9) is N and T/(9) is C;
[0269] R is hydrogen, amino, methyl, trifluoromethyl, or halo;
[0270] U is hydrogen, halo, amino, cyano, nitro, trifluoromethyl, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aSO.sub.2R.sup.b,
--NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --OR.sup.a, or 4-(1H-pyrazol-4-yl), [0271] wherein any (C.sub.1-C.sub.8)alkyl or (C.sub.3-C.sub.8)cycloalkyl group is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, and --OC(O)NR.sup.aR.sup.b; wherein R.sup.a and R.sup.b are defined as above.
[0272] Subgroup B of Formula (VII)
[0273] X and Z are selected independently from the group consisting of (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, heteroaryl, --NR.sup.aR.sup.b, and --OR.sup.a;
[0274] Y is H;
[0275] R.sup.1 is (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, or heterocycloalkyl;
[0276] R.sup.2 is hydrogen, (C.sub.1-C.sub.3)alkyl, or halo, in which said (C.sub.1-C.sub.3)alkyl is optionally substituted with one to two groups selected from amino and (C.sub.1-C.sub.3)alkylamino;
[0277] R.sup.7 is hydrogen, (C.sub.1-C.sub.3)alkyl, or alkoxy;
[0278] R.sup.3 is hydrogen, (C.sub.1-C.sub.8)alkyl or halo;
[0279] R.sup.6 is hydrogen, halo, cyano, trifluoromethyl, amino, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, aryl, heteroaryl, acylamino; (C.sub.2-C.sub.8)alkynyl, arylalkynyl, heteroarylalkynyl, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, or
--NR.sup.aSO.sub.2R.sup.b; [0280] wherein any (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.2-C.sub.8)alkynyl, arylalkynyl, or heteroarylalkynyl group is optionally substituted by 1, 2 or 3 groups independently selected from halo, (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, --OC(O)NR.sup.aR.sup.b, heterocycloalkyl, aryl, heteroaryl, aryl(C.sub.1-C.sub.4)alkyl, and heteroaryl(C.sub.1-C.sub.4)alkyl;
[0281] R.sup.a and R.sup.b are each independently hydrogen, (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.6-C.sub.10)bicycloalkyl, heterocycloalkyl, aryl, or heteroaryl, wherein said (C.sub.1-C.sub.8)alkyl, (C.sub.2-C.sub.8)alkenyl, (C.sub.2-C.sub.8)alkynyl, cycloalkyl, cycloalkenyl, bicycloalkyl, heterocycloalkyl, aryl or heteroaryl group is optionally substituted by 1, 2 or 3 groups independently selected from halo, hydroxyl, (C.sub.1-C.sub.4)alkoxy, amino, (C.sub.1-C.sub.4)alkylamino, ((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl)amino, --CO.sub.2H, --CO.sub.2(C.sub.1-C.sub.4)alkyl, --CONH.sub.2, --CONH(C.sub.1-C.sub.4)alkyl,
--CON((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl), --SO.sub.2(C.sub.1-C.sub.4)alkyl, --SO.sub.2NH.sub.2, --SO.sub.2NH(C.sub.1-C.sub.4)alkyl, and --SO.sub.2N((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl);
[0282] or R.sup.a and R.sup.b taken together with the nitrogen to which they are attached represent a 5-8 membered saturated or unsaturated ring, optionally containing an additional heteroatom selected from oxygen, nitrogen, and sulfur, wherein said ring is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.4)alkyl, (C.sub.1-C.sub.4)haloalkyl, amino, (C.sub.1-C.sub.4)alkylamino, ((C.sub.1-C.sub.4)alkyl)((C.sub.1-C.sub.4)alkyl)amino, hydroxyl, oxo, (C.sub.1-C.sub.4)alkoxy, and (C.sub.1-C.sub.4)alkoxy(C.sub.1-C.sub.4)alkyl, wherein said ring is optionally fused to a (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, or heteroaryl ring;
[0283] or R.sup.a and R.sup.b taken together with the nitrogen to which they are attached represent a 6- to 10-membered bridged bicyclic ring system optionally fused to a (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, aryl, or heteroaryl ring. Aryl and heteroaryl in this definition are selected from the group consisting of furan, thiophene, pyrrole, oxazole, thiazole, imidazole, pyrazole, oxadiazole, thiadiazole, triazole, tetrazole, benzofuran, benzothiophene, benzoxazole, benzothiazole, phenyl, pyridine, pyridazine, pyrimidine, pyrazine, triazine, tetrazine, quinoline, cinnoline, quinazoline, quinoxaline, and naphthyridine or a compound of another aryl or heteroaryl group as follows:
##STR00029##
wherein in (1),
[0284] A is O, NH, or S; B is CH or N, and C is hydrogen or C.sub.1-C.sub.8 alkyl; or
##STR00030##
wherein in (2),
[0285] D is N or C optionally substituted by hydrogen or C.sub.1-C.sub.8 alkyl; or
##STR00031##
wherein in (3),
[0286] E is NH or CH.sub.2; F is O or CO; and G is NH or CH.sub.2; or
##STR00032##
wherein in (4), [0287] J is O, S or CO; or
##STR00033##
[0287] wherein in (5),
[0288] Q is CH or N;
[0289] M is CH or N; and
[0290] L/(5) is hydrogen, halo, amino, cyano, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --CONR.sup.aNR.sup.aR.sup.b, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --NR.sup.aNR.sup.aC(O)NR.sup.aR.sup.b, or --OR.sup.a, [0291] wherein any (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, group is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, and --OC(O)NR.sup.aR.sup.b, wherein R.sup.a and R.sup.b are defined as above; or
##STR00034##
[0291] wherein in (6),
[0292] L/(6) is NH or CH.sub.2; or
##STR00035##
wherein in (7), [0293] M/(7) is hydrogen, halo, amino, cyano, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --CONR.sup.aNR.sup.aR.sup.b, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --NR.sup.aNR.sup.aC(O)NR.sup.aR.sup.b, or --OR.sup.a, [0294] wherein any (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, heterocycloalkyl group is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SR.sup.a, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, --OC(O)NR.sup.aR.sup.b; wherein R.sup.a and R.sup.b are defined as above; or
##STR00036##
[0294] wherein in (8),
[0295] P is CH.sub.2, NH, O, or S; Q/(8) is CH or N; and n is 0-2; or
##STR00037##
wherein in (9),
[0296] S/(9) and T/(9) is C, or S/(9) is C and T/(9) is N, or S/(9) is N and T/(9) is C;
[0297] R is hydrogen, amino, methyl, trifluoromethyl, halo;
[0298] U is hydrogen, halo, amino, cyano, nitro, trifluoromethyl, (C.sub.1-C.sub.8)alkyl, (C.sub.3-C.sub.8)cycloalkyl, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aSO.sub.2R.sup.b,
--NR.sup.aSO.sub.2NR.sup.aR.sup.b, --NR.sup.aNR.sup.aR.sup.b, --NR.sup.aNR.sup.aC(O)R.sup.b, --OR.sup.a, or 4-(1H-pyrazol-4-yl), [0299] wherein any (C.sub.1-C.sub.8)alkyl, or (C.sub.3-C.sub.8)cycloalkyl group is optionally substituted by 1, 2 or 3 groups independently selected from (C.sub.1-C.sub.6)alkyl, (C.sub.3-C.sub.8)cycloalkyl, (C.sub.5-C.sub.8)cycloalkenyl, (C.sub.1-C.sub.6)haloalkyl, cyano, --COR.sup.a, --CO.sub.2R.sup.a, --CONR.sup.aR.sup.b, --SOR.sup.a, --SO.sub.2R.sup.a, --SO.sub.2NR.sup.aR.sup.b, nitro, --NR.sup.aR.sup.b, --NR.sup.aC(O)R.sup.b, --NR.sup.aC(O)NR.sup.aR.sup.b, --NR.sup.aC(O)OR.sup.a, --NR.sup.aSO.sub.2R.sup.b, --NR.sup.aSO.sub.2NR.sup.aR.sup.b, --OR.sup.a, --OC(O)R.sup.a, and --OC(O)NR.sup.aR.sup.b, wherein Ra and Rb are defined as above.
[0300] In some embodiments, the EZH2 inhibitor is:
##STR00038##
stereoisomers thereof or pharmaceutically acceptable salt or solvate thereof.
[0301] In some embodiments, the EZH2 inhibitor is
##STR00039##
stereoisomers thereof or pharmaceutically acceptable salt or solvate thereof.
[0302] The compounds described herein can be synthesized according to any method known in the art. For example, the compounds having the Formula (VII) can be synthesized according to the method described in WO 2011/140325; WO 2011/140324; and WO 2012/005805, each of which is incorporated by reference in its entirety.
[0303] As used herein, "alkyl", "C.sub.1, C.sub.2, C.sub.3, C.sub.4, C.sub.5 or C.sub.6 alkyl" or "C.sub.1-C.sub.6 alkyl" is intended to include C.sub.1, C.sub.2, C.sub.3, C.sub.4, C.sub.5 or C.sub.6 straight chain (linear) saturated aliphatic hydrocarbon groups and C.sub.3, C.sub.4, C.sub.5 or C.sub.6 branched saturated aliphatic hydrocarbon groups. For example, C.sub.1-C.sub.6 alkyl is intended to include C.sub.1, C.sub.2, C.sub.3, C.sub.4, C.sub.5 and C.sub.6 alkyl groups. Examples of alkyl include, moieties having from one to six carbon atoms, such as, but not limited to, methyl, ethyl, n-propyl, i-propyl, n-butyl, s-butyl, t-butyl, n-pentyl, s-pentyl or n-hexyl.
[0304] In certain embodiments, a straight chain or branched alkyl has six or fewer carbon atoms (e.g., C.sub.1-C.sub.6 for straight chain, C.sub.3-C.sub.6 for branched chain), and in another embodiment, a straight chain or branched alkyl has four or fewer carbon atoms.
[0305] As used herein, the term "cycloalkyl" refers to a saturated or unsaturated nonaromatic hydrocarbon mono- or multi-ring (e.g., fused, bridged, or spiro rings) system having 3 to 30 carbon atoms (e.g., C.sub.3-C.sub.10). Examples of cycloalkyl include, but are not limited to, cyclopropyl, cyclobutyl, cyclopentyl, cyclohexyl, cycloheptyl, cyclooctyl, cyclopentenyl, cyclohexenyl, cycloheptenyl, and adamantyl. The term "heterocycloalkyl" refers to a saturated or unsaturated nonaromatic 3-8 membered monocyclic, 7-12 membered bicyclic (fused, bridged, or spiro rings), or 11-14 membered tricyclic ring system (fused, bridged, or spiro rings) having one or more heteroatoms (such as O, N, S, or Se), unless specified otherwise. Examples of heterocycloalkyl groups include, but are not limited to, piperidinyl, piperazinyl, pyrrolidinyl, dioxanyl, tetrahydrofuranyl, isoindolinyl, indolinyl, imidazolidinyl, pyrazolidinyl, oxazolidinyl, isoxazolidinyl, triazolidinyl, tetrahyrofuranyl, oxiranyl, azetidinyl, oxetanyl, thietanyl, 1,2,3,6-tetrahydropyridinyl, tetrahydropyranyl, dihydropyranyl, pyranyl, morpholinyl, 1,4-diazepanyl, 1,4-oxazepanyl, 2-oxa-5-azabicyclo[2.2.1]heptanyl, diazabicyclo[2.2.1]heptanyl, 2-oxa-6-azaspiro[3.3]heptanyl, 2,6-diazaspiro[3.3]heptanyl, 1,4-dioxa-8-azaspiro[4.5]decanyl and the like.
[0306] The term "optionally substituted alkyl" refers to unsubstituted alkyl or alkyl having designated substituents replacing one or more hydrogen atoms on one or more carbons of the hydrocarbon backbone. Such substituents can include, for example, alkyl, alkenyl, alkynyl, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, amino (including alkylamino, dialkylamino, arylamino, diarylamino and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety.
[0307] An "arylalkyl" or an "aralkyl" moiety is an alkyl substituted with an aryl (e.g., phenylmethyl (benzyl)). An "alkylaryl" moiety is an aryl substituted with an alkyl (e.g., methylphenyl).
[0308] As used herein, "alkyl linker" is intended to include C.sub.1, C.sub.2, C.sub.3, C.sub.4, C.sub.5 or C.sub.6 straight chain (linear) saturated divalent aliphatic hydrocarbon groups and C.sub.3, C.sub.4, C.sub.5 or C.sub.6 branched saturated aliphatic hydrocarbon groups. For example, C.sub.1-C.sub.6 alkyl linker is intended to include C.sub.1, C.sub.2, C.sub.3, C.sub.4, C.sub.5 and C.sub.6 alkyl linker groups. Examples of alkyl linker include, moieties having from one to six carbon atoms, such as, but not limited to, methyl (--CH.sub.2--), ethyl (--CH.sub.2CH.sub.2--), n-propyl (--CH.sub.2CH.sub.2CH.sub.2--), i-propyl (--CHCH.sub.3CH.sub.2--), n-butyl (--CH.sub.2CH.sub.2CH.sub.2CH.sub.2--), s-butyl (--CHCH.sub.3CH.sub.2CH.sub.2--), i-butyl (--C(CH.sub.3).sub.2CH.sub.2--), n-pentyl (--CH.sub.2CH.sub.2CH.sub.2CH.sub.2CH.sub.2--), s-pentyl (--CHCH.sub.3CH.sub.2CH.sub.2CH.sub.2--) or n-hexyl (--CH.sub.2CH.sub.2CH.sub.2CH.sub.2CH.sub.2CH.sub.2--).
[0309] "Alkenyl" includes unsaturated aliphatic groups analogous in length and possible substitution to the alkyls described above, but that contain at least one double bond. For example, the term "alkenyl" includes straight chain alkenyl groups (e.g., ethenyl, propenyl, butenyl, pentenyl, hexenyl, heptenyl, octenyl, nonenyl, decenyl), and branched alkenyl groups. In certain embodiments, a straight chain or branched alkenyl group has six or fewer carbon atoms in its backbone (e.g., C.sub.2-C.sub.6 for straight chain, C.sub.3-C.sub.6 for branched chain). The term "C.sub.2-C.sub.6" includes alkenyl groups containing two to six carbon atoms. The term "C.sub.3-C.sub.6" includes alkenyl groups containing three to six carbon atoms.
[0310] The term "optionally substituted alkenyl" refers to unsubstituted alkenyl or alkenyl having designated substituents replacing one or more hydrogen atoms on one or more hydrocarbon backbone carbon atoms. Such substituents can include, for example, alkyl, alkenyl, alkynyl, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, amino (including alkylamino, dialkylamino, arylamino, diarylamino and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety.
[0311] "Alkynyl" includes unsaturated aliphatic groups analogous in length and possible substitution to the alkyls described above, but which contain at least one triple bond. For example, "alkynyl" includes straight chain alkynyl groups (e.g., ethynyl, propynyl, butynyl, pentynyl, hexynyl, heptynyl, octynyl, nonynyl, decynyl), and branched alkynyl groups. In certain embodiments, a straight chain or branched alkynyl group has six or fewer carbon atoms in its backbone (e.g., C.sub.2-C.sub.6 for straight chain, C.sub.3-C.sub.6 for branched chain). The term "C.sub.2-C.sub.6" includes alkynyl groups containing two to six carbon atoms. The term "C.sub.3-C.sub.6" includes alkynyl groups containing three to six carbon atoms.
[0312] The term "optionally substituted alkynyl" refers to unsubstituted alkynyl or alkynyl having designated substituents replacing one or more hydrogen atoms on one or more hydrocarbon backbone carbon atoms. Such substituents can include, for example, alkyl, alkenyl, alkynyl, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, amino (including alkylamino, dialkylamino, arylamino, diarylamino and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety.
[0313] Other optionally substituted moieties (such as optionally substituted cycloalkyl, heterocycloalkyl, aryl, or heteroaryl) include both the unsubstituted moieties and the moieties having one or more of the designated substituents. For example, substituted heterocycloalkyl includes those substituted with one or more alkyl groups, such as 2,2,6,6-tetramethyl-piperidinyl and 2,2,6,6-tetramethyl-1,2,3,6-tetrahydropyridinyl.
[0314] "Aryl" includes groups with aromaticity, including "conjugated," or multicyclic systems with at least one aromatic ring and do not contain any heteroatom in the ring structure. Examples include phenyl, benzyl, 1,2,3,4-tetrahydronaphthalenyl, etc.
[0315] "Heteroaryl" groups are aryl groups, as defined above, except having from one to four heteroatoms in the ring structure, and may also be referred to as "aryl heterocycles" or "heteroaromatics." As used herein, the term "heteroaryl" is intended to include a stable 5-, 6-, or 7-membered monocyclic or 7-, 8-, 9-, 10-, 11- or 12-membered bicyclic aromatic heterocyclic ring which consists of carbon atoms and one or more heteroatoms, e.g., 1 or 1-2 or 1-3 or 1-4 or 1-5 or 1-6 heteroatoms, or e.g. 2, 3, 4, 5, or 6 heteroatoms, independently selected from the group consisting of nitrogen, oxygen and sulfur. The nitrogen atom may be substituted or unsubstituted (i.e., N or NR wherein R is H or other substituents, as defined). The nitrogen and sulfur heteroatoms may optionally be oxidized (i.e., N.fwdarw.O and S(O).sub.p, where p=1 or 2). It is to be noted that total number of S and O atoms in the aromatic heterocycle is not more than 1.
[0316] Examples of heteroaryl groups include pyrrole, furan, thiophene, thiazole, isothiazole, imidazole, triazole, tetrazole, pyrazole, oxazole, isoxazole, pyridine, pyrazine, pyridazine, pyrimidine, and the like.
[0317] Furthermore, the terms "aryl" and "heteroaryl" include multicyclic aryl and heteroaryl groups, e.g., tricyclic, bicyclic, e.g., naphthalene, benzoxazole, benzodioxazole, benzothiazole, benzoimidazole, benzothiophene, methylenedioxyphenyl, quinoline, isoquinoline, naphthrydine, indole, benzofuran, purine, benzofuran, deazapurine, indolizine.
[0318] In the case of multicyclic aromatic rings, only one of the rings needs to be aromatic (e.g., 2,3-dihydroindole), although all of the rings may be aromatic (e.g., quinoline). The second ring can also be fused or bridged.
[0319] The cycloalkyl, heterocycloalkyl, aryl, or heteroaryl ring can be substituted at one or more ring positions (e.g., the ring-forming carbon or heteroatom such as N) with such substituents as described above, for example, alkyl, alkenyl, alkynyl, halogen, hydroxyl, alkoxy, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, alkylaminocarbonyl, aralkylaminocarbonyl, alkenylaminocarbonyl, alkylcarbonyl, arylcarbonyl, aralkylcarbonyl, alkenylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylthiocarbonyl, phosphate, phosphonato, phosphinato, amino (including alkylamino, dialkylamino, arylamino, diarylamino and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety. Aryl and heteroaryl groups can also be fused or bridged with alicyclic or heterocyclic rings, which are not aromatic so as to form a multicyclic system (e.g., tetralin, methylenedioxyphenyl).
[0320] As used herein, "carbocycle" or "carbocyclic ring" is intended to include any stable monocyclic, bicyclic or tricyclic ring having the specified number of carbons, any of which may be saturated, unsaturated, or aromatic. Carbocycle includes cycloalkyl and aryl. For example, a C.sub.3-C.sub.14 carbocycle is intended to include a monocyclic, bicyclic or tricyclic ring having 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13 or 14 carbon atoms. Examples of carbocycles include, but are not limited to, cyclopropyl, cyclobutyl, cyclobutenyl, cyclopentyl, cyclopentenyl, cyclohexyl, cycloheptenyl, cycloheptyl, cycloheptenyl, adamantyl, cyclooctyl, cyclooctenyl, cyclooctadienyl, fluorenyl, phenyl, naphthyl, indanyl, adamantyl and tetrahydronaphthyl. Bridged rings are also included in the definition of carbocycle, including, for example, [3.3.0]bicyclooctane, [4.3.0]bicyclononane, [4.4.0]bicyclodecane and [2.2.2]bicyclooctane. A bridged ring occurs when one or more carbon atoms link two non-adjacent carbon atoms. In one embodiment, bridge rings are one or two carbon atoms. It is noted that a bridge always converts a monocyclic ring into a tricyclic ring. When a ring is bridged, the substituents recited for the ring may also be present on the bridge. Fused (e.g., naphthyl, tetrahydronaphthyl) and spiro rings are also included.
[0321] As used herein, "heterocycle" or "heterocyclic group" includes any ring structure (saturated, unsaturated, or aromatic) which contains at least one ring heteroatom (e.g., N, O or S). Heterocycle includes heterocycloalkyl and heteroaryl. Examples of heterocycles include, but are not limited to, morpholine, pyrrolidine, tetrahydrothiophene, piperidine, piperazine, oxetane, pyran, tetrahydropyran, azetidine, and tetrahydrofuran.
[0322] Examples of heterocyclic groups include, but are not limited to, acridinyl, azocinyl, benzimidazolyl, benzofuranyl, benzothiofuranyl, benzothiophenyl, benzoxazolyl, benzoxazolinyl, benzthiazolyl, benztriazolyl, benztetrazolyl, benzisoxazolyl, benzisothiazolyl, benzimidazolinyl, carbazolyl, 4aH-carbazolyl, carbolinyl, chromanyl, chromenyl, cinnolinyl, decahydroquinolinyl, 2H,6H-1,5,2-dithiazinyl, dihydrofuro[2,3-b]tetrahydrofuran, furanyl, furazanyl, imidazolidinyl, imidazolinyl, imidazolyl, 1H-indazolyl, indolenyl, indolinyl, indolizinyl, indolyl, 3H-indolyl, isatinoyl, isobenzofuranyl, isochromanyl, isoindazolyl, isoindolinyl, isoindolyl, isoquinolinyl, isothiazolyl, isoxazolyl, methylenedioxyphenyl, morpholinyl, naphthyridinyl, octahydroisoquinolinyl, oxadiazolyl, 1,2,3-oxadiazolyl, 1,2,4-oxadiazolyl, 1,2,5-oxadiazolyl, 1,3,4-oxadiazolyl, 1,2,4-oxadiazol5(4H)-one, oxazolidinyl, oxazolyl, oxindolyl, pyrimidinyl, phenanthridinyl, phenanthrolinyl, phenazinyl, phenothiazinyl, phenoxathinyl, phenoxazinyl, phthalazinyl, piperazinyl, piperidinyl, piperidonyl, 4-piperidonyl, piperonyl, pteridinyl, purinyl, pyranyl, pyrazinyl, pyrazolidinyl, pyrazolinyl, pyrazolyl, pyridazinyl, pyridooxazole, pyridoimidazole, pyridothiazole, pyridinyl, pyridyl, pyrimidinyl, pyrrolidinyl, pyrrolinyl, 2H-pyrrolyl, pyrrolyl, quinazolinyl, quinolinyl, 4H-quinolizinyl, quinoxalinyl, quinuclidinyl, tetrahydrofuranyl, tetrahydroisoquinolinyl, tetrahydroquinolinyl, tetrazolyl, 6H-1,2,5-thiadiazinyl, 1,2,3-thiadiazolyl, 1,2,4-thiadiazolyl, 1,2,5-thiadiazolyl, 1,3,4-thiadiazolyl, thianthrenyl, thiazolyl, thienyl, thienothiazolyl, thienooxazolyl, thienoimidazolyl, thiophenyl, triazinyl, 1,2,3-triazolyl, 1,2,4-triazolyl, 1,2,5-triazolyl, 1,3,4-triazolyl and xanthenyl.
[0323] The term "substituted," as used herein, means that any one or more hydrogen atoms on the designated atom is replaced with a selection from the indicated groups, provided that the designated atom's normal valency is not exceeded, and that the substitution results in a stable compound. When a substituent is oxo or keto (i.e., .dbd.O), then 2 hydrogen atoms on the atom are replaced. Keto substituents are not present on aromatic moieties. Ring double bonds, as used herein, are double bonds that are formed between two adjacent ring atoms (e.g., C.dbd.C, C.dbd.N or N.dbd.N). "Stable compound" and "stable structure" are meant to indicate a compound that is sufficiently robust to survive isolation to a useful degree of purity from a reaction mixture, and formulation into an efficacious therapeutic agent.
[0324] When a bond to a substituent is shown to cross a bond connecting two atoms in a ring, then such substituent may be bonded to any atom in the ring. When a substituent is listed without indicating the atom via which such substituent is bonded to the rest of the compound of a given formula, then such substituent may be bonded via any atom in such formula. Combinations of substituents and/or variables are permissible, but only if such combinations result in stable compounds.
[0325] When any variable (e.g., R.sub.1) occurs more than one time in any constituent or formula for a compound, its definition at each occurrence is independent of its definition at every other occurrence. Thus, for example, if a group is shown to be substituted with 0-2 R.sub.1 moieties, then the group may optionally be substituted with up to two R.sub.1 moieties and R.sub.1 at each occurrence is selected independently from the definition of R.sub.1. Also, combinations of substituents and/or variables are permissible, but only if such combinations result in stable compounds.
[0326] The term "hydroxy" or "hydroxyl" includes groups with an --OH or --O.sup.-.
[0327] As used herein, "halo" or "halogen" refers to fluoro, chloro, bromo and iodo. The term "perhalogenated" generally refers to a moiety wherein all hydrogen atoms are replaced by halogen atoms. The term "haloalkyl" or "haloalkoxyl" refers to an alkyl or alkoxyl substituted with one or more halogen atoms.
[0328] The term "carbonyl" includes compounds and moieties which contain a carbon connected with a double bond to an oxygen atom. Examples of moieties containing a carbonyl include, but are not limited to, aldehydes, ketones, carboxylic acids, amides, esters, anhydrides, etc.
[0329] The term "carboxyl" refers to --COOH or its C.sub.1-C.sub.6 alkyl ester.
[0330] "Acyl" includes moieties that contain the acyl radical (R--C(O)--) or a carbonyl group. "Substituted acyl" includes acyl groups where one or more of the hydrogen atoms are replaced by, for example, alkyl groups, alkynyl groups, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, amino (including alkylamino, dialkylamino, arylamino, diarylamino and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moiety.
[0331] "Aroyl" includes moieties with an aryl or heteroaromatic moiety bound to a carbonyl group. Examples of aroyl groups include phenylcarboxy, naphthyl carboxy, etc.
[0332] "Alkoxyalkyl," "alkylaminoalkyl," and "thioalkoxyalkyl" include alkyl groups, as described above, wherein oxygen, nitrogen, or sulfur atoms replace one or more hydrocarbon backbone carbon atoms.
[0333] The term "alkoxy" or "alkoxyl" includes substituted and unsubstituted alkyl, alkenyl and alkynyl groups covalently linked to an oxygen atom. Examples of alkoxy groups or alkoxyl radicals include, but are not limited to, methoxy, ethoxy, isopropyloxy, propoxy, butoxy and pentoxy groups. Examples of substituted alkoxy groups include halogenated alkoxy groups. The alkoxy groups can be substituted with groups such as alkenyl, alkynyl, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, phosphate, phosphonato, phosphinato, amino (including alkylamino, dialkylamino, arylamino, diarylamino, and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moieties. Examples of halogen substituted alkoxy groups include, but are not limited to, fluoromethoxy, difluoromethoxy, trifluoromethoxy, chloromethoxy, dichloromethoxy and trichloromethoxy.
[0334] The term "ether" or "alkoxy" includes compounds or moieties which contain an oxygen bonded to two carbon atoms or heteroatoms. For example, the term includes "alkoxyalkyl," which refers to an alkyl, alkenyl, or alkynyl group covalently bonded to an oxygen atom which is covalently bonded to an alkyl group.
[0335] The term "ester" includes compounds or moieties which contain a carbon or a heteroatom bound to an oxygen atom which is bonded to the carbon of a carbonyl group. The term "ester" includes alkoxycarboxy groups such as methoxycarbonyl, ethoxycarbonyl, propoxycarbonyl, butoxycarbonyl, pentoxycarbonyl, etc.
[0336] The term "thioalkyl" includes compounds or moieties which contain an alkyl group connected with a sulfur atom. The thioalkyl groups can be substituted with groups such as alkyl, alkenyl, alkynyl, halogen, hydroxyl, alkylcarbonyloxy, arylcarbonyloxy, alkoxycarbonyloxy, aryloxycarbonyloxy, carboxylate, carboxyacid, alkylcarbonyl, arylcarbonyl, alkoxycarbonyl, aminocarbonyl, alkylaminocarbonyl, dialkylaminocarbonyl, alkylthiocarbonyl, alkoxyl, amino (including alkylamino, dialkylamino, arylamino, diarylamino and alkylarylamino), acylamino (including alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido), amidino, imino, sulfhydryl, alkylthio, arylthio, thiocarboxylate, sulfates, alkylsulfinyl, sulfonato, sulfamoyl, sulfonamido, nitro, trifluoromethyl, cyano, azido, heterocyclyl, alkylaryl, or an aromatic or heteroaromatic moieties.
[0337] The term "thiocarbonyl" or "thiocarboxy" includes compounds and moieties which contain a carbon connected with a double bond to a sulfur atom.
[0338] The term "thioether" includes moieties which contain a sulfur atom bonded to two carbon atoms or heteroatoms. Examples of thioethers include, but are not limited to alkthioalkyls, alkthioalkenyls, and alkthioalkynyls. The term "alkthioalkyls" include moieties with an alkyl, alkenyl, or alkynyl group bonded to a sulfur atom which is bonded to an alkyl group. Similarly, the term "alkthioalkenyls" refers to moieties wherein an alkyl, alkenyl or alkynyl group is bonded to a sulfur atom which is covalently bonded to an alkenyl group; and alkthioalkynyls" refers to moieties wherein an alkyl, alkenyl or alkynyl group is bonded to a sulfur atom which is covalently bonded to an alkynyl group.
[0339] As used herein, "amine" or "amino" refers to unsubstituted or substituted --NH.sub.2. "Alkylamino" includes groups of compounds wherein nitrogen of --NH.sub.2 is bound to at least one alkyl group. Examples of alkylamino groups include benzylamino, methylamino, ethylamino, phenethylamino, etc. "Dialkylamino" includes groups wherein the nitrogen of --NH.sub.2 is bound to at least two additional alkyl groups. Examples of dialkylamino groups include, but are not limited to, dimethylamino and diethylamino. "Arylamino" and "diarylamino" include groups wherein the nitrogen is bound to at least one or two aryl groups, respectively. "Aminoaryl" and "aminoaryloxy" refer to aryl and aryloxy substituted with amino. "Alkylarylamino," "alkylaminoaryl" or "arylaminoalkyl" refers to an amino group which is bound to at least one alkyl group and at least one aryl group. "Alkaminoalkyl" refers to an alkyl, alkenyl, or alkynyl group bound to a nitrogen atom which is also bound to an alkyl group. "Acylamino" includes groups wherein nitrogen is bound to an acyl group. Examples of acylamino include, but are not limited to, alkylcarbonylamino, arylcarbonylamino, carbamoyl and ureido groups.
[0340] The term "amide" or "aminocarboxy" includes compounds or moieties that contain a nitrogen atom that is bound to the carbon of a carbonyl or a thiocarbonyl group. The term includes "alkaminocarboxy" groups that include alkyl, alkenyl or alkynyl groups bound to an amino group which is bound to the carbon of a carbonyl or thiocarbonyl group. It also includes "arylaminocarboxy" groups that include aryl or heteroaryl moieties bound to an amino group that is bound to the carbon of a carbonyl or thiocarbonyl group. The terms "alkylaminocarboxy", "alkenylaminocarboxy", "alkynylaminocarboxy" and "arylaminocarboxy" include moieties wherein alkyl, alkenyl, alkynyl and aryl moieties, respectively, are bound to a nitrogen atom which is in turn bound to the carbon of a carbonyl group. Amides can be substituted with substituents such as straight chain alkyl, branched alkyl, cycloalkyl, aryl, heteroaryl or heterocycle. Substituents on amide groups may be further substituted.
[0341] In the present specification, the structural formula of the compound represents a certain isomer for convenience in some cases, but the present invention includes all isomers, such as geometrical isomers, optical isomers based on an asymmetrical carbon, stereoisomers, tautomers, and the like, it being understood that not all isomers may have the same level of activity. In addition, a crystal polymorphism may be present for the compounds represented by the formula. It is noted that any crystal form, crystal form mixture, or anhydride or hydrate thereof is included in the scope of the present invention. Furthermore, so-called metabolite which is produced by degradation of the present compound in vivo is included in the scope of the present invention.
[0342] "Isomerism" means compounds that have identical molecular formulae but differ in the sequence of bonding of their atoms or in the arrangement of their atoms in space. Isomers that differ in the arrangement of their atoms in space are termed "stereoisomers." Stereoisomers that are not mirror images of one another are termed "diastereoisomers," and stereoisomers that are non-superimposable mirror images of each other are termed "enantiomers" or sometimes optical isomers. A mixture containing equal amounts of individual enantiomeric forms of opposite chirality is termed a "racemic mixture."
[0343] A carbon atom bonded to four nonidentical substituents is termed a "chiral center."
[0344] "Chiral isomer" means a compound with at least one chiral center. Compounds with more than one chiral center may exist either as an individual diastereomer or as a mixture of diastereomers, termed "diastereomeric mixture." When one chiral center is present, a stereoisomer may be characterized by the absolute configuration (R or S) of that chiral center. Absolute configuration refers to the arrangement in space of the substituents attached to the chiral center. The substituents attached to the chiral center under consideration are ranked in accordance with the Sequence Rule of Cahn, Ingold and Prelog. (Cahn et al., Angew. Chem. Inter. Edit. 1966, 5, 385; errata 511; Cahn et al., Angew. Chem. 1966, 78, 413; Cahn and Ingold, J. Chem. Soc. 1951 (London), 612; Cahn et al., Experientia 1956, 12, 81; Cahn, J. Chem. Educ. 1964, 41, 116).
[0345] "Geometric isomer" means the diastereomers that owe their existence to hindered rotation about double bonds or a cycloalkyl linker (e.g., 1,3-cylcobutyl). These configurations are differentiated in their names by the prefixes cis and trans, or Z and E, which indicate that the groups are on the same or opposite side of the double bond in the molecule according to the Cahn-Ingold-Prelog rules.
[0346] It is to be understood that the compounds of the present invention may be depicted as different chiral isomers or geometric isomers. It should also be understood that when compounds have chiral isomeric or geometric isomeric forms, all isomeric forms are intended to be included in the scope of the present invention, and the naming of the compounds does not exclude any isomeric forms, it being understood that not all isomers may have the same level of activity.
[0347] Furthermore, the structures and other compounds discussed in this invention include all atropic isomers thereof, it being understood that not all atropic isomers may have the same level of activity. "Atropic isomers" are a type of stereoisomer in which the atoms of two isomers are arranged differently in space. Atropic isomers owe their existence to a restricted rotation caused by hindrance of rotation of large groups about a central bond. Such atropic isomers typically exist as a mixture, however as a result of recent advances in chromatography techniques, it has been possible to separate mixtures of two atropic isomers in select cases.
[0348] "Tautomer" is one of two or more structural isomers that exist in equilibrium and is readily converted from one isomeric form to another. This conversion results in the formal migration of a hydrogen atom accompanied by a switch of adjacent conjugated double bonds. Tautomers exist as a mixture of a tautomeric set in solution. In solutions where tautomerization is possible, a chemical equilibrium of the tautomers will be reached. The exact ratio of the tautomers depends on several factors, including temperature, solvent and pH. The concept of tautomers that are interconvertable by tautomerizations is called tautomerism.
[0349] Of the various types of tautomerism that are possible, two are commonly observed. In keto-enol tautomerism a simultaneous shift of electrons and a hydrogen atom occurs. Ring-chain tautomerism arises as a result of the aldehyde group (--CHO) in a sugar chain molecule reacting with one of the hydroxy groups (--OH) in the same molecule to give it a cyclic (ring-shaped) form as exhibited by glucose.
[0350] Common tautomeric pairs are: ketone-enol, amide-nitrile, lactam-lactim, amide-imidic acid tautomerism in heterocyclic rings (e.g., in nucleobases such as guanine, thymine and cytosine), imine-enamine and enamine-enamine. An example of keto-enol equilibria is between pyridin-2(1H)-ones and the corresponding pyridin-2-ols, as shown below.
##STR00040##
[0351] It is to be understood that the compounds of the present invention may be depicted as different tautomers. It should also be understood that when compounds have tautomeric forms, all tautomeric forms are intended to be included in the scope of the present invention, and the naming of the compounds does not exclude any tautomer form. It will be understood that certain tautomers may have a higher level of activity than others.
[0352] The term "crystal polymorphs", "polymorphs" or "crystal forms" means crystal structures in which a compound (or a salt or solvate thereof) can crystallize in different crystal packing arrangements, all of which have the same elemental composition. Different crystal forms usually have different X-ray diffraction patterns, infrared spectral, melting points, density hardness, crystal shape, optical and electrical properties, stability and solubility. Recrystallization solvent, rate of crystallization, storage temperature, and other factors may cause one crystal form to dominate. Crystal polymorphs of the compounds can be prepared by crystallization under different conditions.
[0353] The compounds of any of Formulae disclosed herein include the compounds themselves, as well as their salts or their solvates, if applicable. A salt, for example, can be formed between an anion and a positively charged group (e.g., amino) on an aryl- or heteroaryl-substituted benzene compound. Suitable anions include chloride, bromide, iodide, sulfate, bisulfate, sulfamate, nitrate, phosphate, citrate, methanesulfonate, trifluoroacetate, glutamate, glucuronate, glutarate, malate, maleate, succinate, fumarate, tartrate, tosylate, salicylate, lactate, naphthalenesulfonate, and acetate (e.g., trifluoroacetate). The term "pharmaceutically acceptable anion" refers to an anion suitable for forming a pharmaceutically acceptable salt. Likewise, a salt can also be formed between a cation and a negatively charged group (e.g., carboxylate) on an aryl- or heteroaryl-substituted benzene compound. Suitable cations include sodium ion, potassium ion, magnesium ion, calcium ion, and an ammonium cation such as tetramethylammonium ion. The aryl- or heteroaryl-substituted benzene compounds also include those salts containing quaternary nitrogen atoms.
[0354] Additionally, the compounds of the present invention, for example, the salts of the compounds, can exist in either hydrated or unhydrated (the anhydrous) form or as solvates with other solvent molecules. Nonlimiting examples of hydrates include monohydrates, dihydrates, etc. Nonlimiting examples of solvates include ethanol solvates, acetone solvates, etc.
[0355] "Solvate" means solvent addition forms that contain either stoichiometric or non stoichiometric amounts of solvent. Some compounds have a tendency to trap a fixed molar ratio of solvent molecules in the crystalline solid state, thus forming a solvate. If the solvent is water the solvate formed is a hydrate; and if the solvent is alcohol, the solvate formed is an alcoholate. Hydrates are formed by the combination of one or more molecules of water with one molecule of the substance in which the water retains its molecular state as H.sub.2O.
[0356] As used herein, the term "analog" refers to a chemical compound that is structurally similar to another but differs slightly in composition (as in the replacement of one atom by an atom of a different element or in the presence of a particular functional group, or the replacement of one functional group by another functional group). Thus, an analog is a compound that is similar or comparable in function and appearance, but not in structure or origin to the reference compound.
[0357] As defined herein, the term "derivative" refers to compounds that have a common core structure, and are substituted with various groups as described herein. For example, all of the compounds represented by Formula (I) are aryl- or heteroaryl-substituted benzene compounds, and have Formula (I) as a common core.
[0358] The term "bioisostere" refers to a compound resulting from the exchange of an atom or of a group of atoms with another, broadly similar, atom or group of atoms. The objective of a bioisosteric replacement is to create a new compound with similar biological properties to the parent compound. The bioisosteric replacement may be physicochemically or topologically based. Examples of carboxylic acid bioisosteres include, but are not limited to, acyl sulfonimides, tetrazoles, sulfonates and phosphonates. See, e.g., Patani and LaVoie, Chem. Rev. 96, 3147-3176, 1996.
[0359] The present invention is intended to include all isotopes of atoms occurring in the present compounds. Isotopes include those atoms having the same atomic number but different mass numbers. By way of general example and without limitation, isotopes of hydrogen include tritium and deuterium, and isotopes of carbon include C-13 and C-14.
[0360] The present invention provides methods for the synthesis of the compounds of any Formula disclosed herein. The present invention also provides detailed methods for the synthesis of various disclosed compounds of the present invention according to the following schemes as shown in the Examples.
[0361] Throughout the description, where compositions are described as having, including, or comprising specific components, it is contemplated that compositions also consist essentially of, or consist of, the recited components. Similarly, where methods or processes are described as having, including, or comprising specific process steps, the processes also consist essentially of, or consist of, the recited processing steps. Further, it should be understood that the order of steps or order for performing certain actions is immaterial so long as the invention remains operable. Moreover, two or more steps or actions can be conducted simultaneously.
[0362] The synthetic processes of the invention can tolerate a wide variety of functional groups, therefore various substituted starting materials can be used. The processes generally provide the desired final compound at or near the end of the overall process, although it may be desirable in certain instances to further convert the compound to a pharmaceutically acceptable salt, polymorph or solvate thereof.
[0363] Compounds of the present invention can be prepared in a variety of ways using commercially available starting materials, compounds known in the literature, or from readily prepared intermediates, by employing standard synthetic methods and procedures either known to those skilled in the art, or which will be apparent to the skilled artisan in light of the teachings herein. Standard synthetic methods and procedures for the preparation of organic molecules and functional group transformations and manipulations can be obtained from the relevant scientific literature or from standard textbooks in the field. Although not limited to any one or several sources, classic texts such as Smith, M. B., March, J., March's Advanced Organic Chemistry: Reactions, Mechanisms, and Structure, 5.sup.th edition, John Wiley & Sons: New York, 2001; Greene, T. W., Wuts, P. G. M., Protective Groups in Organic Synthesis, 3.sup.rd edition, John Wiley & Sons: New York, 1999; R. Larock, Comprehensive Organic Transformations, VCH Publishers (1989); L. Fieser and M. Fieser, Fieser and Fieser's Reagents for Organic Synthesis, John Wiley and Sons (1994); and L. Paquette, ed., Encyclopedia of Reagents for Organic Synthesis, John Wiley and Sons (1995), incorporated by reference herein, are useful and recognized reference textbooks of organic synthesis known to those in the art. The following descriptions of synthetic methods are designed to illustrate, but not to limit, general procedures for the preparation of compounds of the present invention.
[0364] Compounds of the present invention can be conveniently prepared by a variety of methods familiar to those skilled in the art. The compounds of this invention with any Formula disclosed herein may be prepared according to the procedures illustrated in Schemes 1-10 below, from commercially available starting materials or starting materials which can be prepared using literature procedures. The Z and R groups (such as R.sub.2, R.sub.3, R.sub.4, R.sub.6, R.sub.7, R.sub.8, and R.sub.12) in Schemes 1-10 are as defined in any of Formulae disclosed herein, unless otherwise specified.
[0365] One of ordinary skill in the art will note that, during the reaction sequences and synthetic schemes described herein, the order of certain steps may be changed, such as the introduction and removal of protecting groups.
[0366] One of ordinary skill in the art will recognize that certain groups may require protection from the reaction conditions via the use of protecting groups. Protecting groups may also be used to differentiate similar functional groups in molecules. A list of protecting groups and how to introduce and remove these groups can be found in Greene, T. W., Wuts, P. G. M., Protective Groups in Organic Synthesis, 3.sup.rd edition, John Wiley & Sons: New York, 1999.
[0367] Preferred protecting groups include, but are not limited to:
[0368] For a hydroxyl moiety: TBS, benzyl, THP, Ac
[0369] For carboxylic acids: benzyl ester, methyl ester, ethyl ester, allyl ester
[0370] For amines: Cbz, BOC, DMB
[0371] For diols: Ac (.times.2) TBS (.times.2), or when taken together acetonides
[0372] For thiols: Ac
[0373] For benzimidazoles: SEM, benzyl, PMB, DMB
[0374] For aldehydes: di-alkyl acetals such as dimethoxy acetal or diethyl acetyl.
[0375] In the reaction schemes described herein, multiple stereoisomers may be produced. When no particular stereoisomer is indicated, it is understood to mean all possible stereoisomers that could be produced from the reaction. A person of ordinary skill in the art will recognize that the reactions can be optimized to give one isomer preferentially, or new schemes may be devised to produce a single isomer. If mixtures are produced, techniques such as preparative thin layer chromatography, preparative HPLC, preparative chiral HPLC, or preparative SFC may be used to separate the isomers.
[0376] The following abbreviations are used throughout the specification and are defined below: [0377] Ac acetyl [0378] AcOH acetic acid [0379] aq. aqueous [0380] BID or b.i.d. bis in die (twice a day) [0381] BOC tert-butoxy carbonyl [0382] Cbz benzyloxy carbonyl [0383] CDCl.sub.3 deuterated chloroform [0384] CH.sub.2Cl.sub.2 dichloromethane [0385] DCM dichloromethane [0386] DMB 2,4 dimethoxy benzyl [0387] DMF N,N-Dimethylformamide [0388] DMSO Dimethyl sulfoxide [0389] EA or EtOAc Ethyl acetate [0390] EDC or EDCI N-(3-Dimethylaminopropyl)-N'-ethylcarbodiimide [0391] ESI- Electrospray negative mode [0392] ESI+ Electrospray positive mode [0393] EtOH ethanol [0394] h hours [0395] H.sub.2O water [0396] HOBt 1-Hydroxybenzotriazole [0397] HCl hydrogen chloride or hydrochloric acid [0398] HPLC High performance liquid chromatography [0399] K.sub.2CO.sub.3 potassium carbonate [0400] LC/MS or LC-MS Liquid chromatography mass spectrum [0401] M Molar [0402] MeCN Acetonitrile [0403] min minutes [0404] Na.sub.2CO.sub.3 sodium carbonate [0405] Na.sub.2SO.sub.4 sodium sulfate [0406] NaHCO.sub.3 sodium bicarbonate [0407] NaHMDs Sodium hexamethyldisilazide [0408] NaOH sodium hydroxide [0409] NaHCO.sub.3 sodium bicarbonate [0410] Na.sub.2SO.sub.4 sodium sulfate [0411] NMR Nuclear Magnetic Resonance [0412] Pd(OH).sub.2 Palladium dihydroxide [0413] PMB para methoxybenzyl [0414] p.o. per os (oral administration) [0415] ppm parts per million [0416] prep HPLC preparative High Performance Liquid Chromatography [0417] PYBOP (Benzotriazol-1-yloxy)tripyrrolidinophosphonium [0418] hexafluorophosphate [0419] Rt or RT Room temperature [0420] TBME tert-Butyl methyl ether [0421] TFA trifluoroacetic acid [0422] THF tetrahydrofuran [0423] THP tetrahydropyran
[0424] The present invention also provides pharmaceutical compositions comprising a compound of any Formula disclosed herein in combination with at least one pharmaceutically acceptable excipient or carrier.
[0425] A "pharmaceutical composition" is a formulation containing the compounds of the present invention in a form suitable for administration to a subject. In one embodiment, the pharmaceutical composition is in bulk or in unit dosage form. The unit dosage form is any of a variety of forms, including, for example, a capsule, an IV bag, a tablet, a single pump on an aerosol inhaler or a vial. The quantity of active ingredient (e.g., a formulation of the disclosed compound or salt, hydrate, solvate or isomer thereof) in a unit dose of composition is an effective amount and is varied according to the particular treatment involved. One skilled in the art will appreciate that it is sometimes necessary to make routine variations to the dosage depending on the age and condition of the patient. The dosage will also depend on the route of administration. A variety of routes are contemplated, including oral, pulmonary, rectal, parenteral, transdermal, subcutaneous, intravenous, intramuscular, intraperitoneal, inhalational, buccal, sublingual, intrapleural, intrathecal, intranasal, and the like. Dosage forms for the topical or transdermal administration of a compound of this invention include powders, sprays, ointments, pastes, creams, lotions, gels, solutions, patches and inhalants. In one embodiment, the active compound is mixed under sterile conditions with a pharmaceutically acceptable carrier, and with any preservatives, buffers, or propellants that are required.
[0426] As used herein, the phrase "pharmaceutically acceptable" refers to those compounds, anions, cations, materials, compositions, carriers, and/or dosage forms which are, within the scope of sound medical judgment, suitable for use in contact with the tissues of human beings and animals without excessive toxicity, irritation, allergic response, or other problem or complication, commensurate with a reasonable benefit/risk ratio.
[0427] "Pharmaceutically acceptable excipient" means an excipient that is useful in preparing a pharmaceutical composition that is generally safe, non-toxic and neither biologically nor otherwise undesirable, and includes excipient that is acceptable for veterinary use as well as human pharmaceutical use. A "pharmaceutically acceptable excipient" as used in the specification and claims includes both one and more than one such excipient.
[0428] A pharmaceutical composition of the invention is formulated to be compatible with its intended route of administration. Examples of routes of administration include parenteral, e.g., intravenous, intradermal, subcutaneous, oral (e.g., inhalation), transdermal (topical), and transmucosal administration. Solutions or suspensions used for parenteral, intradermal, or subcutaneous application can include the following components: a sterile diluent such as water for injection, saline solution, fixed oils, polyethylene glycols, glycerine, propylene glycol or other synthetic solvents; antibacterial agents such as benzyl alcohol or methyl parabens; antioxidants such as ascorbic acid or sodium bisulfite; chelating agents such as ethylenediaminetetraacetic acid; buffers such as acetates, citrates or phosphates, and agents for the adjustment of tonicity such as sodium chloride or dextrose. The pH can be adjusted with acids or bases, such as hydrochloric acid or sodium hydroxide. The parenteral preparation can be enclosed in ampoules, disposable syringes or multiple dose vials made of glass or plastic.
[0429] A compound or pharmaceutical composition of the invention can be administered to a subject in many of the well-known methods currently used for chemotherapeutic treatment. For example, for treatment of cancers, a compound of the invention may be injected directly into tumors, injected into the blood stream or body cavities or taken orally or applied through the skin with patches. The dose chosen should be sufficient to constitute effective treatment but not so high as to cause unacceptable side effects. The state of the disease condition (e.g., cancer, precancer, and the like) and the health of the patient should preferably be closely monitored during and for a reasonable period after treatment.
[0430] The term "therapeutically effective amount", as used herein, refers to an amount of a pharmaceutical agent to treat, ameliorate, or prevent an identified disease or condition, or to exhibit a detectable therapeutic or inhibitory effect. The effect can be detected by any assay method known in the art. The precise effective amount for a subject will depend upon the subject's body weight, size, and health; the nature and extent of the condition; and the therapeutic or combination of therapeutics selected for administration. Therapeutically effective amounts for a given situation can be determined by routine experimentation that is within the skill and judgment of the clinician. In a preferred aspect, the disease or condition to be treated is cancer. In another aspect, the disease or condition to be treated is a cell proliferative disorder.
[0431] For any compound, the therapeutically effective amount can be estimated initially either in cell culture assays, e.g., of neoplastic cells, or in animal models, usually rats, mice, rabbits, dogs, or pigs. The animal model may also be used to determine the appropriate concentration range and route of administration. Such information can then be used to determine useful doses and routes for administration in humans. Therapeutic/prophylactic efficacy and toxicity may be determined by standard pharmaceutical procedures in cell cultures or experimental animals, e.g., ED.sub.50 (the dose therapeutically effective in 50% of the population) and LD.sub.50 (the dose lethal to 50% of the population). The dose ratio between toxic and therapeutic effects is the therapeutic index, and it can be expressed as the ratio, LD.sub.50/ED.sub.50. Pharmaceutical compositions that exhibit large therapeutic indices are preferred. The dosage may vary within this range depending upon the dosage form employed, sensitivity of the patient, and the route of administration.
[0432] Dosage and administration are adjusted to provide sufficient levels of the active agent(s) or to maintain the desired effect. Factors which may be taken into account include the severity of the disease state, general health of the subject, age, weight, and gender of the subject, diet, time and frequency of administration, drug combination(s), reaction sensitivities, and tolerance/response to therapy. Long-acting pharmaceutical compositions may be administered every 3 to 4 days, every week, or once every two weeks depending on half-life and clearance rate of the particular formulation.
[0433] The pharmaceutical compositions containing active compounds of the present invention may be manufactured in a manner that is generally known, e.g., by means of conventional mixing, dissolving, granulating, dragee-making, levigating, emulsifying, encapsulating, entrapping, or lyophilizing processes. Pharmaceutical compositions may be formulated in a conventional manner using one or more pharmaceutically acceptable carriers comprising excipients and/or auxiliaries that facilitate processing of the active compounds into preparations that can be used pharmaceutically. Of course, the appropriate formulation is dependent upon the route of administration chosen.
[0434] Pharmaceutical compositions suitable for injectable use include sterile aqueous solutions (where water soluble) or dispersions and sterile powders for the extemporaneous preparation of sterile injectable solutions or dispersion. For intravenous administration, suitable carriers include physiological saline, bacteriostatic water, Cremophor EL.TM. (BASF, Parsippany, N.J.) or phosphate buffered saline (PBS). In all cases, the composition must be sterile and should be fluid to the extent that easy syringeability exists. It must be stable under the conditions of manufacture and storage and must be preserved against the contaminating action of microorganisms such as bacteria and fungi. The carrier can be a solvent or dispersion medium containing, for example, water, ethanol, polyol (for example, glycerol, propylene glycol, and liquid polyethylene glycol, and the like), and suitable mixtures thereof. The proper fluidity can be maintained, for example, by the use of a coating such as lecithin, by the maintenance of the required particle size in the case of dispersion and by the use of surfactants. Prevention of the action of microorganisms can be achieved by various antibacterial and antifungal agents, for example, parabens, chlorobutanol, phenol, ascorbic acid, thimerosal, and the like. In many cases, it will be preferable to include isotonic agents, for example, sugars, polyalcohols such as manitol and sorbitol, and sodium chloride in the composition. Prolonged absorption of the injectable compositions can be brought about by including in the composition an agent which delays absorption, for example, aluminum monostearate and gelatin.
[0435] Sterile injectable solutions can be prepared by incorporating the active compound in the required amount in an appropriate solvent with one or a combination of ingredients enumerated above, as required, followed by filtered sterilization. Generally, dispersions are prepared by incorporating the active compound into a sterile vehicle that contains a basic dispersion medium and the required other ingredients from those enumerated above. In the case of sterile powders for the preparation of sterile injectable solutions, methods of preparation are vacuum drying and freeze-drying that yields a powder of the active ingredient plus any additional desired ingredient from a previously sterile-filtered solution thereof.
[0436] Oral compositions generally include an inert diluent or an edible pharmaceutically acceptable carrier. They can be enclosed in gelatin capsules or compressed into tablets. For the purpose of oral therapeutic administration, the active compound can be incorporated with excipients and used in the form of tablets, troches, or capsules. Oral compositions can also be prepared using a fluid carrier for use as a mouthwash, wherein the compound in the fluid carrier is applied orally and swished and expectorated or swallowed. Pharmaceutically compatible binding agents, and/or adjuvant materials can be included as part of the composition. The tablets, pills, capsules, troches and the like can contain any of the following ingredients, or compounds of a similar nature: a binder such as microcrystalline cellulose, gum tragacanth or gelatin; an excipient such as starch or lactose, a disintegrating agent such as alginic acid, Primogel, or corn starch; a lubricant such as magnesium stearate or Sterotes; a glidant such as colloidal silicon dioxide; a sweetening agent such as sucrose or saccharin; or a flavoring agent such as peppermint, methyl salicylate, or orange flavoring.
[0437] For administration by inhalation, the compounds are delivered in the form of an aerosol spray from pressured container or dispenser, which contains a suitable propellant, e.g., a gas such as carbon dioxide, or a nebulizer.
[0438] Systemic administration can also be by transmucosal or transdermal means. For transmucosal or transdermal administration, penetrants appropriate to the barrier to be permeated are used in the formulation. Such penetrants are generally known in the art, and include, for example, for transmucosal administration, detergents, bile salts, and fusidic acid derivatives. Transmucosal administration can be accomplished through the use of nasal sprays or suppositories. For transdermal administration, the active compounds are formulated into ointments, salves, gels, or creams as generally known in the art.
[0439] The active compounds can be prepared with pharmaceutically acceptable carriers that will protect the compound against rapid elimination from the body, such as a controlled release formulation, including implants and microencapsulated delivery systems. Biodegradable, biocompatible polymers can be used, such as ethylene vinyl acetate, polyanhydrides, polyglycolic acid, collagen, polyorthoesters, and polylactic acid. Methods for preparation of such formulations will be apparent to those skilled in the art. The materials can also be obtained commercially from Alza Corporation and Nova Pharmaceuticals, Inc. Liposomal suspensions (including liposomes targeted to infected cells with monoclonal antibodies to viral antigens) can also be used as pharmaceutically acceptable carriers. These can be prepared according to methods known to those skilled in the art, for example, as described in U.S. Pat. No. 4,522,811.
[0440] It is especially advantageous to formulate oral or parenteral compositions in dosage unit form for ease of administration and uniformity of dosage. Dosage unit form as used herein refers to physically discrete units suited as unitary dosages for the subject to be treated; each unit containing a predetermined quantity of active compound calculated to produce the desired therapeutic effect in association with the required pharmaceutical carrier. The specification for the dosage unit forms of the invention are dictated by and directly dependent on the unique characteristics of the active compound and the particular therapeutic effect to be achieved.
[0441] In therapeutic applications, the dosages of the pharmaceutical compositions used in accordance with the invention vary depending on the agent, the age, weight, and clinical condition of the recipient patient, and the experience and judgment of the clinician or practitioner administering the therapy, among other factors affecting the selected dosage. Generally, the dose should be sufficient to result in slowing, and preferably regressing, the growth of the tumors and also preferably causing complete regression of the cancer. Dosages can range from about 0.01 mg/kg per day to about 5000 mg/kg per day. In preferred aspects, dosages can range from about 1 mg/kg per day to about 1000 mg/kg per day. In an aspect, the dose will be in the range of about 0.1 mg/day to about 50 g/day; about 0.1 mg/day to about 25 g/day; about 0.1 mg/day to about 10 g/day; about 0.1 mg to about 3 g/day; or about 0.1 mg to about 1 g/day, in single, divided, or continuous doses (which dose may be adjusted for the patient's weight in kg, body surface area in m.sup.2, and age in years). An effective amount of a pharmaceutical agent is that which provides an objectively identifiable improvement as noted by the clinician or other qualified observer. For example, regression of a tumor in a patient may be measured with reference to the diameter of a tumor. Decrease in the diameter of a tumor indicates regression. Regression is also indicated by failure of tumors to reoccur after treatment has stopped. As used herein, the term "dosage effective manner" refers to amount of an active compound to produce the desired biological effect in a subject or cell.
[0442] The pharmaceutical compositions can be included in a container, pack, or dispenser together with instructions for administration.
[0443] The compounds of the present invention are capable of further forming salts. All of these forms are also contemplated within the scope of the claimed invention.
[0444] As used herein, "pharmaceutically acceptable salts" refer to derivatives of the compounds of the present invention wherein the parent compound is modified by making acid or base salts thereof. Examples of pharmaceutically acceptable salts include, but are not limited to, mineral or organic acid salts of basic residues such as amines, alkali or organic salts of acidic residues such as carboxylic acids, and the like. The pharmaceutically acceptable salts include the conventional non-toxic salts or the quaternary ammonium salts of the parent compound formed, for example, from non-toxic inorganic or organic acids. For example, such conventional non-toxic salts include, but are not limited to, those derived from inorganic and organic acids selected from 2-acetoxybenzoic, 2-hydroxyethane sulfonic, acetic, ascorbic, benzene sulfonic, benzoic, bicarbonic, carbonic, citric, edetic, ethane disulfonic, 1,2-ethane sulfonic, fumaric, glucoheptonic, gluconic, glutamic, glycolic, glycollyarsanilic, hexylresorcinic, hydrabamic, hydrobromic, hydrochloric, hydroiodic, hydroxymaleic, hydroxynaphthoic, isethionic, lactic, lactobionic, lauryl sulfonic, maleic, malic, mandelic, methane sulfonic, napsylic, nitric, oxalic, pamoic, pantothenic, phenylacetic, phosphoric, polygalacturonic, propionic, salicyclic, stearic, subacetic, succinic, sulfamic, sulfanilic, sulfuric, tannic, tartaric, toluene sulfonic, and the commonly occurring amine acids, e.g., glycine, alanine, phenylalanine, arginine, etc.
[0445] Other examples of pharmaceutically acceptable salts include hexanoic acid, cyclopentane propionic acid, pyruvic acid, malonic acid, 3-(4-hydroxybenzoyl)benzoic acid, cinnamic acid, 4-chlorobenzenesulfonic acid, 2-naphthalenesulfonic acid, 4-toluenesulfonic acid, camphorsulfonic acid, 4-methylbicyclo-[2.2.2]-oct-2-ene-1-carboxylic acid, 3-phenylpropionic acid, trimethylacetic acid, tertiary butylacetic acid, muconic acid, and the like. The present invention also encompasses salts formed when an acidic proton present in the parent compound either is replaced by a metal ion, e.g., an alkali metal ion, an alkaline earth ion, or an aluminum ion; or coordinates with an organic base such as ethanolamine, diethanolamine, triethanolamine, tromethamine, N-methylglucamine, and the like. In the salt form, it is understood that the ratio of the compound to the cation or anion of the salt can be 1:1, or any ration other than 1:1, e.g., 3:1, 2:1, 1:2, or 1:3.
[0446] It should be understood that all references to pharmaceutically acceptable salts include solvent addition forms (solvates) or crystal forms (polymorphs) as defined herein, of the same salt.
[0447] The compounds of the present invention can also be prepared as esters, for example, pharmaceutically acceptable esters. For example, a carboxylic acid function group in a compound can be converted to its corresponding ester, e.g., a methyl, ethyl or other ester. Also, an alcohol group in a compound can be converted to its corresponding ester, e.g., acetate, propionate or other ester.
[0448] The compounds, or pharmaceutically acceptable salts or solvates thereof, are administered orally, nasally, transdermally, pulmonary, inhalationally, buccally, sublingually, intraperintoneally, subcutaneously, intramuscularly, intravenously, rectally, intrapleurally, intrathecally and parenterally. In one embodiment, the compound is administered orally. One skilled in the art will recognize the advantages of certain routes of administration.
[0449] The dosage regimen utilizing the compounds is selected in accordance with a variety of factors including type, species, age, weight, sex and medical condition of the patient; the severity of the condition to be treated; the route of administration; the renal and hepatic function of the patient; and the particular compound or salt thereof employed. An ordinarily skilled physician or veterinarian can readily determine and prescribe the effective amount of the drug required to prevent, counter, or arrest the progress of the condition.
[0450] Techniques for formulation and administration of the disclosed compounds of the invention can be found in Remington: the Science and Practice of Pharmacy, 19.sup.th edition, Mack Publishing Co., Easton, Pa. (1995). In an embodiment, the compounds described herein, and the pharmaceutically acceptable salts thereof, are used in pharmaceutical preparations in combination with a pharmaceutically acceptable carrier or diluent. Suitable pharmaceutically acceptable carriers include inert solid fillers or diluents and sterile aqueous or organic solutions. The compounds will be present in such pharmaceutical compositions in amounts sufficient to provide the desired dosage amount in the range described herein.
[0451] All percentages and ratios used herein, unless otherwise indicated, are by weight. Other features and advantages of the present invention are apparent from the different examples. The provided examples illustrate different components and methodology useful in practicing the present invention. The examples do not limit the claimed invention. Based on the present disclosure the skilled artisan can identify and employ other components and methodology useful for practicing the present invention.
[0452] In the synthetic schemes described herein, compounds may be drawn with one particular configuration for simplicity. Such particular configurations are not to be construed as limiting the invention to one or another isomer, tautomer, regioisomer or stereoisomer, nor does it exclude mixtures of isomers, tautomers, regioisomers or stereoisomers; however, it will be understood that a given isomer, tautomer, regioisomer or stereoisomer may have a higher level of activity than another isomer, tautomer, regioisomer or stereoisomer.
[0453] Compounds designed, selected and/or optimized by methods described above, once produced, can be characterized using a variety of assays known to those skilled in the art to determine whether the compounds have biological activity. For example, the molecules can be characterized by conventional assays, including but not limited to those assays described below, to determine whether they have a predicted activity, binding activity and/or binding specificity.
[0454] Furthermore, high-throughput screening can be used to speed up analysis using such assays. As a result, it can be possible to rapidly screen the molecules described herein for activity, using techniques known in the art. General methodologies for performing high-throughput screening are described, for example, in Devlin (1998) High Throughput Screening, Marcel Dekker; and U.S. Pat. No. 5,763,263. High-throughput assays can use one or more different assay techniques including, but not limited to, those described below.
[0455] An EZH2 inhibitor of the present invention may, if desired, be presented in a kit (e.g., a pack or dispenser device) which may contain one or more unit dosage forms containing the EZH2 inhibitor. The pack may for example comprise metal or plastic foil, such as a blister pack. The pack or dispenser device may be accompanied by instructions for administration. Compositions comprising an EZH2 inhibitor of the invention formulated in a compatible pharmaceutical carrier may also be prepared, placed in an appropriate container, and labeled for treatment of an indicated condition. Instructions for use may also be provided.
[0456] Also provided herein are kits comprising a plurality of methylation detection reagents that detect the methylated H3-K27. For example, the kit includes mono-methylated H3-K27, di-methylated H3-K27 and tri-methylated H3-K27 detection reagents. The detection reagent is for example antibodies or fragments thereof, polypeptide or aptamers.
[0457] A kit may also include reagents for detecting loss of function of at least one component of the SWI/SNF complex, e.g., nucleic acids that specifically identify a mutant component nucleic acid sequence by having homologous nucleic acid sequences, such as oligonucleotide sequences, complementary to a portion of the mutant component nucleic acid sequence or antibodies to proteins encoded by the wild type and/or mutant component nucleic acids packaged together in the form of a kit. The oligonucleotides can be fragments of the component gene. For example the oligonucleotides can be 200, 150, 100, 50, 25, 10 or less nucleotides in length. The kit may contain in separate containers an aptamer or an antibody, control formulations (positive and/or negative), and/or a detectable label such as fluorescein, green fluorescent protein, rhodamine, cyanine dyes, Alexa dyes, luciferase, radiolabels, among others. In addition, reagents for detecting the biological activity of the SWI/SNF complex (such as its chromatin remodeling activity) may be included in the kit.
[0458] Instructions (e.g., written, tape, VCR, CD-ROM, etc.) for carrying out the assay may be included in the kit. The assay may for example be in the form of a Western Blot analysis, Immunohistochemistry (IHC), immunofluorescence (IF), sequencing and Mass spectrometry (MS) as known in the art.
Example 1: Durable Tumor Regression in Genetically Altered Lymphomas and Malignant Rhabdoid Tumors by Inhibition of EZH2
[0459] Compound A is a Potent and Selective Inhibitor of EZH2:
[0460] Cell free biochemical assays that included radiolabeled SAM and either chicken erythrocyte oligonucleosomes or peptides corresponding to H3K27 as substrates showed that Compound A selectively inhibited the activity of human PRC2 containing wild-type EZH2 with an inhibition constant (Ki) value of 2.5.+-.0.5 nmol/L and IC.sub.50 values of 11.+-.5 nM (nucleosome assay) or 16.+-.12 nM (peptide assay). The IC.sub.50 values were similar for human and rat EZH2 enzymes as well as for EZH2 proteins bearing all known lymphoma change-of-function mutations. The IC.sub.50 value of Compound A increased with increasing concentration of SAM, but was minimally affected by increasing the amount of oligonucleosome which is consistent with a SAM-competitive and nucleosome-noncompetitive modality of inhibition. In order to demonstrate HMT selectivity, inhibition by Compound A against a panel of HMTs other than EZH2 encompassing both lysine and arginine HMTs was assessed. Compound A displayed a 35-fold selectivity versus EZH1 and greater than 4500-fold selectivity relative to the 14 other HMTs tested.
[0461] Compound A Specifically Inhibits Cellular H3K27 Methylation in Cells:
[0462] When WSU-DLCL2 EZH2 Y641F mutant lymphoma cells were incubated with Compound A for 4 days, a concentration-dependent reduction in global H3K27Me3 levels was observed with an average IC.sub.50 value of 0.26 .mu.M (H3K27Me3 levels determined by ELISA). When studying the kinetics of methylation inhibition, the half-life of H3K27Me3 was approximately 1 day as 90% inhibition was only achieved after 3 to 4 days of incubation. When OCI-LY19 EZH2 wild-type lymphoma cells were incubated with 2.7 .mu.M Compound A for 4 days, the only methyl marks affected were the H3K27Me1, H3K27Me2 and H3K27Me3, the three known products of PRC2 catalysis. Incubation with Compound A also resulted in an increase in H3K27 acetylation. The ability of Compound A to reduce global H3K27 trimethylation levels was further tested in several other human lymphoma cell lines including lines expressing either wild-type or mutant EZH2. Compound A reduced H3K27Me3 with similar potency in all cell lines independent of the EZH2 status (Table 1).
[0463] Compound A Leads to Selective Killing of Lymphoma Cell Lines Bearing EZH2 Point Mutations:
[0464] Incubation of WSU-DLCL2 EZH2 Y641F mutant cells with Compound A lead to anti-proliferative effects with an average IC.sub.50 value of 0.28.+-.0.14 .mu.M in a 6 day proliferation assay. The kinetics of the effect of Compound A on viable cell number was further tested over an extended period of 11 days. The antiproliferative effect of Compound A was apparent after WSU-DLCL2 cells had been exposed to compound for longer than 4 days, consistent with the kinetics of Compound A-mediated cellular H3K27 methylation inhibition. The IC.sub.50 value for Compound A inhibition of proliferation of WSU-DLCL2 cells in the 11-day assay (0.0086 Table 1) was lower when compared with results obtained with a 6-day proliferation assay, suggesting increased sensitivity with longer incubation periods. In contrast to the WSU-DLCL2 cells, the growth of OCI-LY19 human lymphoma cells (EZH2 wild type for residue Y641) over 11 days was not significantly affected, despite comparable IC.sub.50 values for H3K27Me3 inhibition for both cell lines (Table 1). In order to identify a concentration at which cells stop proliferating considering the entire incubation period of 11 days, the lowest cytotoxic concentration (LCC) for a particular cell line was calculated. The LCC value for WSU-DLCL2 EZH2 Y641F mutant human lymphoma cells was significantly lower when compared with OCI-LY19 cells that are wild type for EZH2 (Table 1). This context specific cell killing was further supported by results from 11-day proliferation assays with an extended lymphoma cell line panel. All cell lines harboring an EZH2 mutation, with the exception of the RL cell line (EZH2 Y641N), were more sensitive to the antiproliferative effects of Compound A when compared with cell lines with wild-type EZH2 (Table 1). The Pfeiffer cell line (EZH2 A677G) showed a 20 to 300 fold increase in sensitivity to Compound A, as measured by IC.sub.50 value and LCC, respectively, over the Y641 mutant cell lines. Next the minimum time of compound exposure necessary for sustained cell killing was investigated by washout experiments. The LCC values on day 11 or 14 for WSU-DLCL2 cells that were either incubated with Compound A for 7 days (followed by 7 days of compound washout) or continuously for 14 days were similar (Table 2). Drug exposure for only 4 days, however, was not sufficient to induce LCC values similar to continuous incubation.
[0465] Compound A Induces G.sub.1 Arrest and Apoptosis in EZH2 Mutant Lymphoma Cells:
[0466] Next, the effects of incubation with Compound A (1 .mu.M) for 7 days on cell cycle progression and apoptosis in WSU-DLCL2 cells were assessed. An increase in the percentage of cells in G.sub.1 phase, and a decrease in the percentage of cells in S phase and G.sub.2/M phase was apparent after 2 days of Compound A incubation. The maximum effect was achieved after 4 days. There was no apparent increase in the sub-G.sub.1 fraction suggesting that apoptosis was not induced by Compound A incubation for 7 days. This is in agreement with the growth curves of WSU-DLCL2 cells in the presence of Compound A indicating that cytotoxic effects were observed only after 7 days of incubation. Following incubation of WSU-DLCL2 cells with Compound A for up to 14 days, the fraction of apoptotic cells determined by TUNEL assay was significantly increased on day 14 compared to vehicle, indicating that Compound A-mediated cell death occurred through the induction of apoptosis.
[0467] Oral Administration of Compound A Leads to EZH2 Target Inhibition in EZH2 Mutant Xenograft Models in Mice:
[0468] The effect of oral dosing of Compound A on systemic compound exposure and in vivo target inhibition in mice bearing EZH2 mutant lymphoma xenografts was investigated. First, SCID mice implanted subcutaneously with WSU-DLCL2 xenografts were orally dosed with Compound A for 4 or 7 days. Measuring Compound A plasma levels either 5 minutes before or 3 hours after the last dose revealed a clear dose dependent increase in exposure. Only animals dosed at 160 mg/kg TID or 213 mg/kg BID maintained mean compound levels in plasma above the LCC for WSU-DLCL2 cells throughout a dosing cycle (1652 ng/mL, with mouse plasma protein binding considered). Compound determination in homogenates from tumors collected 3 hours after the last dose revealed that only for the highest dose groups compound levels in the 2 compartments were similar. When H3K27Me3 levels in tumors were analyzed, dose dependent EZH2 target inhibition was observed. H3K27Me3 inhibition was less in tumors from mice dosed at 213 mg/kg QD, suggesting that maintaining a plasma concentration above LCC throughout a dosing cycle is required for optimal target inhibition. Dosing for 4 days at 160 mg/kg TID resulted in slightly lower target inhibition than dosing for 7 days at the same dose and schedule, indicating that prolonged dosing increased the degree of target inhibition in WSU-DLCL2 tumors. A similar 7-day study in nude mice implanted subcutaneously with KARAPS-422 xenografts assessing both BID and QD schedules was performed. Compound A induced a dose-dependent reduction of tumor H3K27Me3 levels at both regimens.
[0469] Compound A Induces Significant Antitumor Effects in Several EZH2 Mutant Lymphoma Xenografts:
[0470] When WSU-DLCL2 EZH2 Y641F mutant xenograft tumor bearing SCID mice were treated with Compound A for 28 days, dose-dependent tumor growth inhibition, 58% at the highest dose of 150 mg/kg TID, was observed. Only animals administered the highest dose maintained mean Compound A plasma levels above LCC for WSU-DLCL2 cells throughout the dosing cycle. Dosing of Compound A for 28 days led to a relative compound accumulation in tumor tissue compared with plasma, in contrast to what was observed with 7-day dosing. ELISA analysis of histones from tumors collected on day 28 indicated dose-dependent target inhibition. H3K27Me3 levels in WSU-DLCL2 xenografts were lower in mice dosed for 28 days compared with 7 days indicating that prolonged administration of Compound A increased the degree of target inhibition. In KARPAS-422 EZH2 Y461N mutant xenografts, 28-day dosing of Compound A on a BID schedule had much more dramatic effects. Tumor growth inhibition was observed at doses as low as 80.5 mg/kg BID, but higher doses eradicated the xenografts, and no re-growth was observed for up to 90 days after cessation of dosing. When intermittent dosing schedules were investigated in KARPAS-422 xenograft bearing mice, Compound A again showed significant dose-dependent antitumor effects with two cycles of 7-day on/7-day off and 21 day on/7 day off schedules. For all dosing schedules, tumor growth inhibition and complete regressions were observed at 90 and 361 mg/kg BID, respectively. The Pfeiffer EZH2 A677G mutant xenograft model was the most sensitive tumor model, as suggested by the potent anti-proliferative effects of Compound A on this cell line in vitro. All Compound A dose groups (QD schedule) except the lowest one (30 mg/kg QD) showed complete tumor regressions in all animals. Again, tumor re-growth was not observed until the end of the study (36 days after stopping Compound A administration). Although tumor re-growth was observed at 30 mg/kg QD, this very low dose induced tumor stasis during the administration period. Due to tolerability issues dosing was stopped on day 12 for mice administered 1140 mg/kg QD; still, durable complete regressions were observed in this group that were only exposed to Compound A for 12 days.
[0471] Compound A Selectively Kills SMARCB1 Mutant MRT Cells In Vitro and In Vivo:
[0472] Whether EZH2 inhibition had any effects on the growth and survival of SMARCB1-deleted MRT cells was tested. Incubating SMARCB1-deleted MRT cell lines G401 and A204 with Compound A in a 14-day proliferation assay in vitro induced strong anti-proliferative effects with IC.sub.50 values in the nM range while the control cell lines RD and SJCRH30 which expressed SMARCB1 were minimally affected (Table 3). Dosing of SCID mice bearing subcutaneous G401 xenografts with Compound A at 266 or 532 mg/kg BID for 28 days eliminated those extremely fast growing tumors. Similar to the KARPAS-422 and Pfeiffer EZH2 mutant NHL xenograft models re-growth was not observed at study end, 32 days after dosing stop. Compound A dosed at 133 mg/kg induced stasis during the administration period, and produced a significant tumor growth delay compared to vehicle after dosing stop. Tumors that were harvested from subsets of mice from each group on day 21 showed strong EZH2 target inhibition at all doses.
[0473] Compound A Inhibits 113K27 Methylation in Nontumor Tissues in a Dose Dependent Manner:
[0474] The data described above demonstrate that Compound A represents a new treatment modality for SWI/SNF driven cancers and MRTs. Measuring pharmacodynamic biomarker modulation post-dose is often performed in early clinical trials to assess the degree of target inhibition that is predicted to produce a response based on data from preclinical models. Since the collection of post-dose tumor biopsies is often not possible, easier accessible surrogate tissues such as peripheral blood mononuclear cells (PBMCs), skin or bone marrow are often collected instead. To test EZH2 target inhibition in surrogate tissues male and female Sprague Dawley rats were orally administered 100, 300, or 1000 mg/kg Compound A for 28 days, and PBMCs, bone marrow and skin samples were collected at study end. Plasma levels of Compound A increased dose-dependently in both male and female rats, and the plasma levels were generally higher in females compared with those in males. Due to tolerability issues, females in the 1000 mg/kg group had to be euthanized on day 23. Dose-dependent target inhibition was observed in PBMCs and bone marrow from rats dosed with Compound A, as measured by ELISA. The degree of target inhibition was less pronounced for PBMCs from females that were dosed for 22 days compared with males that were dosed for 28 days (same dose of 1000 mg/kg). A dose dependent reduction in H3K27Me3 positive cells was observed in the epidermis of skin of Compound A-dosed rats, as assessed by an IHC assay. The maximum effect was observed at the highest dose, and was already evident after 22 days of Compound A administration.
[0475] Compound A displayed similar properties as other EZH2 inhibitors in vitro, such as very high specificity for EZH2 in biochemical assays when compared with other HMTs and specific inhibition of cellular H3K27 methylation leading to context specific killing of EZH2 mutated NHL cell lines. However, this compound achieved an approximately 10-fold increase in potency, reflected by decreased K.sub.i and IC.sub.50 values determined in biochemical and cell-functional assays. In addition, Compound A showed excellent oral bioavailability when administered to rodents which lead to dose dependent EZH2 target inhibition in xenograft tumor and nontumor tissues. Importantly, dosing of Compound A induced significant antitumor effects in mice bearing EZH2 mutant lymphoma xenografts. The responses ranged from tumor eradication (no regrowth after dosing cessation) to dose-dependent tumor growth inhibition. The delayed onset of antitumor activity (after 4 to 7 days) was consistent with the kinetics of methylation inhibition and antiproliferative activity induced by incubation of cells with Compound A in vitro. Keeping Compound A plasma levels above LCC throughout a dosing cycle was necessary for the WSU-DLCL2 xenograft model to induce maximal target inhibition and antitumor response. The other two lymphoma xenograft models (KARPAS-422 and Pfeiffer), however, were extremely sensitive to Compound A administration, and keeping plasma levels above LCC was not necessary. Pfeiffer EZH2 A677G mutant xenograft tumors disappeared permanently with very low doses or short dosing periods, suggesting that patients with this type of genetically defined NHL would have a significant treatment effect with Compound A.
[0476] MRTs are extremely aggressive pediatric cancers of the brain, kidney, and soft tissues that are highly malignant, locally invasive, frequently metastatic, and particularly lethal, but they are typically diploid and lack genomic aberrations. They are, however, characterized by an almost complete penetrance of loss of expression of the SMARCB1, a core component of the SWI/SNF chromatin remodeling complex. The biallelic inactivation of SMARCB1, for instance induced by mutations, is in essence the sole genetic event in MRTs which suggests a driver role for this genetic aberration. Through genetic studies it has been suggested that PRC2 and SWI/SNF antagonistically regulate gene expression around the RB, Cyclin D1 and MYC pathways. Here, it has been demonstrated pharmacological EZH2 inhibition induced antiproliferative effects in SMARCB1 deleted MRT cell lines and permanently eradicated MRT xenografts in mice. This confirms the dependency of such cancers, in which EZH2 itself is not genetically altered, on PRC2 activity.
[0477] Compound A represents a new treatment modality for genetically defined subsets of NHL and for MRTs. The ability to measure dose-dependent changes in H3K27Me3 levels in skin, PBMCs and bone marrow portends the use of signal from these surrogate tissues as a non-invasive pharmacodynamics biomarker in human clinical trials.
TABLE-US-00004 TABLE 1 IC.sub.50 Values for Methylation and Proliferation as well as LCC Values for Compound A in Human Lymphoma Cell Lines Methylation EZH2 IC.sub.50 Proliferation IC.sub.50 LCC Cell Line Status (nmol/L).sup.a (.mu.mol/L).sup.b (.mu.mol/L).sup.b DOHH-2 Wild Type ND 1.7 >10 Farage Wild Type ND 0.099 >10 OCI-LY19 Wild Type 8 6.2 10-25 Toledo Wild Type ND 7.6 >10 Karpas-422 Y641N 90 0.0018 0.12 Pfeiffer A677G 2 0.00049 0.0005 RL Y641N 22 5.8 >25 SU-DHL-10 Y641F ND 0.0058 0.14 SU-DHL-6 Y641N 20 0.0047 0.21 WSU-DLCL2 Y641F 9 0.0086 0.17 .sup.aDerived after incubation for 4 days by immunoblot. Values represent the result from one experiment. .sup.bDerived after incubation for 11 days. Compound incubations for each experiment were performed in triplicate, and values represent one experiment for all cell lines except OCI-LY19, Pfeiffer, and WSU-DLCL2. For the remaining three cell lines, values represent the mean from the following number of experiments: OCI-LY19 n = 9; Pfeiffer n = 2 and WSU-DLCL2 n = 15.
TABLE-US-00005 TABLE 2 LCC Values for Compound A for WSU-DLCL2 Human Lymphoma Cells Dosed Either Continuously or After Compound Washout Day 11 Day 14 WSU-DLCL2 Washout LCC (.mu.M) LCC (.mu.M) No Washout 0.17 0.11 4-day Compound A; 11-day Washout 0.36 0.42 7-day Compound A; 7-day Washout 0.19 0.075 Values represent the mean of duplicate experiments with three replicates per incubation concentration within the experiments.
TABLE-US-00006 TABLE 3 IC.sub.50 Values for Compound A for SMARCB1 Negative MRT Cell Lines and SMARCB1 Positive Control Cell Lines Proliferation Proliferation Cell Line SMARCB1 Status IC.sub.50 (.mu.M), day 7 IC.sub.50 (.mu.M), day 14 RD Wild Type 9.2 5.2 SJCRH30 Wild Type 6.1 8.8 G401 Mutant 0.087 0.042 A204 Mutant 3.2 0.14 Values represent the mean of duplicate experiments with three replicates per incubation concentration within the experiments.
Example 2: Durable Tumor Regression in Genetically Altered Malignant Rhabdoid Tumors by Inhibition of EZH2
[0478] Compound A is a Potent and Selective Inhibitor of EZH2:
[0479] Compound A was developed through iterative medicinal chemistry (FIG. 10A). Compound A inhibited the activity of human PRC2 containing wild-type EZH2 with an inhibition constant (Ki) value of 2.5.+-.0.5 nM, and similar potency was observed for EZH2 proteins bearing all known lymphoma change-of-function mutations (Table 5). The compound was found to be SAM-competitive and nucleosome-noncompetitive by steady state kinetic studies (FIG. 11). Inhibition by Compound A against a panel of HMTs other than EZH2 encompassing both lysine and arginine HMTs was also assessed. Compound A displayed a 35-fold selectivity versus EZH1 and >4500-fold selectivity relative to 14 other HMTs tested (Table 5).
TABLE-US-00007 TABLE 4 Histone Methyltransferase Inhibition by Compound A % Inhibition at 1 .mu.M Enzyme Assay IC.sub.50 (nM) Compound A.sup.a CARM1 >50,000.sup.b 5 .+-. 3 DOT1L >50,000.sup.c 2 .+-. 8 EHMT1 >50,000.sup.c 6 .+-. 6 EHMT2 >50,000.sup.c 7 .+-. 3 EZH1.sup.d,e 392 .+-. 72.sup.f 98 .+-. 1 EZH2 Peptide Assay.sup.d,e 11 .+-. 5.sup.f ND EZH2 Nucleosome Assay.sup.d 16 .+-. 12.sup.f 100 .+-. 1 A677G EZH2.sup.d,e 2.sup.b ND A687V EZH2.sup.d,e 2.sup.b ND Y641F EZH2.sup.d,e 14 .+-. 5.sup.f ND Y641C EZH2.sup.d,e 16.sup.c ND Y641H EZH2.sup.d,e 6.sup.c ND Y641N EZH2.sup.d,e 38.sup.b ND Y641S EZH2.sup.d,e 6.sup.c ND rat EZH2.sup.d,e 4.sup.c ND PRMT1 >50,000.sup.c 5 .+-. 4 PRMT3 ND 2 .+-. 2 PRMT5/MEP50 >50,000.sup.c 2 .+-. 6 PRMT6 ND 3 .+-. 3 PRMT8 >50,000.sup.c 7 .+-. 3 SETD7 ND 4 .+-. 3 SMYD2 >50,000.sup.c 1 .+-. 2 SMYD3 ND 0 .+-. 5 WHSC1 >100,000.sup.c 8 .+-. 3 WHSC1L1 >100,000.sup.c 9 .+-. 8 .sup.aValues represent the mean and standard deviation of duplicate experiments determined at 10 .mu.mol/L Compound A. .sup.bValues represent the mean of duplicate experiments with two replicates per experiment. .sup.cValues represent one experiment with two replicates per experiment. .sup.dAll EZH1 and EZH2 proteins were assayed in the context of 4 PRC2 components (EZH1/2, SUZ12, RBAP48, EED). .sup.eAssayed with H3K27 peptides as substrates.
[0480] Compound A Specifically Inhibits Cellular 113K27 Methylation Leading to Selective Apoptotic Killing of SMARCB1 Mutant MRT Cells:
[0481] A panel of SMARCB1 deficient MRT cells and SMARCB1 wild-type control cells (confirmed by immunoblot, FIG. 12A) were treated with Compound A for 4 days, resulting in concentration-dependent reductions in global H3K27Me3 levels (FIG. 10B and table 6). Treatment of either wild-type or mutant cells resulted in diminution only of methyl marks on H3K27, with no other histone methyl marks being affected (FIG. 12B). In vitro treatment of SMARCB1-deleted MRT cell lines with Compound A induced strong anti-proliferative effects with IC.sub.50 values in the nM range; while the control (wild-type) cell lines were minimally affected (FIG. 10C and table 6). Antiproliferative effects were apparent in SMARCB1-deleted MRT cells after 7 days of compound exposure, but required 14 days of exposure for maximal activity. The effects of incubation with Compound A (1 .mu.M) for 14 days on cell cycle progression and apoptosis in G401 and RD cells were also assessed. Compound A incubation of RD SMARCB1 wild-type cells showed no changes in cell cycle or apoptosis compared to the DMSO control (FIG. 13A). In contrast, G401 SMARCB1-deleted cells showed an increase in the percentage of cells in G.sub.1 phase, and a concomitant decrease in S phase and G.sub.2/M phase after 7 days (FIG. 13B). There was no apparent increase in the sub-G.sub.1 fraction through day 7, suggesting that apoptosis was not yet induced by that time. This coincides with the growth curves of G401 cells in the presence of Compound A that display cytotoxicity only after 7 days of incubation (FIG. 10C). Following Compound A treatment of G401 cells for up to 14 days, the fraction of cells in sub-G.sub.1 as well as apoptotic cells determined by TUNEL assay increased in a time dependent manner through days 11 and 14, indicating that Compound A-mediated cell death occurred through the induction of apoptosis (FIG. 13B).
TABLE-US-00008 TABLE 6 Methylation Proliferation IC.sub.50 on Day Cell Line SMARCB1 Status IC.sub.50 (nM).sup.a 14 (nM).sup.b G401 mutant 2.7 135 A204 mutant 1.4 590 G402 mutant 1.7 144 KYM-1 mutant 4.3 32 RD wild-type 5.6 6100, >10000.sup.c 293 wild-type 2.4 >10000 SJCRH30 wild-type 4.9 5100, >10000.sup.c .sup.aDerived after incubation for 4 days, extraction of histones, immunoblot and densitometry. Values represent the mean from two experiments. .sup.bCompound incubations for each experiment were performed in triplicate, and values represent the mean of 2 experiments for all cell lines. .sup.cMean calculation of duplicate experiment not possible.
[0482] Compound A Induces Genes of Neuronal Differentiation and Cell Cycle Inhibition while Suppressing Expression of Hedgehog Pathway Genes, MYC and EZH2:
[0483] It has been suggested that SMARCB1 loss drives cancer formation through simultaneous epigenetic perturbation of key cancer pathways. The present data confirmed the previously described reduced expression of genes important for neuronal differentiation (CD133, DOCK4, PTPRK), cell cycle inhibition (CDKN2A) and tumor suppression (BIN1), as well as increased expression of the hedgehog pathway gene GLI1 in SMARCB1-deleted G401 cells compared to control cells (FIG. 14A). Compound A treatment of G401 cells for up to 7 days strongly induced expression of CD133, DOCK4 and PTPRK and up-regulated cell cycle inhibitors CDKN1A and CDKN2A and tumor suppressor BIN1, all in a time-dependent manner (FIG. 14B). Simultaneously, the expression of hedgehog pathway genes, MYC and EZH2 were reduced. Notably, G402 SMARCB1-deleted cells exposed to Compound A for 14 days assumed a neuron-like morphology (FIG. 14C). In contrast, Compound A incubation of RD control cells had minimal effect on expression of the above-mentioned genes.
[0484] Compound A Eradicates SMARCB1 Mutant MRT Xenografts:
[0485] Oral dosing of Compound A led to systemic compound exposure, in vivo target inhibition and antitumor activity in mice bearing SMARCB1-deleted MRT xenografts. A study in SCID mice bearing subcutaneous G401 xenografts was performed where animals were dosed for 21 days with Compound A. Half of the mice per group were euthanized on day 21 to collect blood and tissues, while the remaining animals were treated for an additional 7 days and then left without dosing for another 32 days. Compound A was well tolerated at all doses with minimal effect on body weight (FIG. 15A). Dosing at 250 or 500 mg/kg twice daily (BID) for 21 to 28 days practically eliminated the fast-growing G401 tumors (FIGS. 15B, 14C and 16A). Re-growth was not observed for 32 days after dose cessation. Compound A dosed at 125 mg/kg induced tumor stasis during the administration period, and produced a significant tumor growth delay compared to vehicle after the dosing period. Measuring Compound A plasma levels either 5 min before or 3 h after dosing on day 21 revealed a clear dose-dependent increase in systemic exposure (FIG. 15D). Tumors that were harvested from subsets of mice from each group on day 21 showed strong inhibition of H3K27me3, correlating with the antitumor activity (maximum effect achieved at 250 mg/kg, FIG. 16B). In addition, dose-dependent changes in the expression of CD133, PTPRK, DOCK4 and GLI1 were detected in the G401 xenograft tumors (FIG. 16C).
[0486] The present data demonstrate that pharmacological inhibition of EZH2 induced antiproliferative effects specifically in SMARCB1-deleted MRT cell lines and permanently eradicated MRT xenografts in mice. This confirms the dependency of such cancers on PRC2 activity, despite the fact that EZH2 itself is not genetically altered in this context. Data presented herein show that in the context of SMARCB1-deleted MRT, inhibition of EZH2 functions as a SMARCB1 surrogate and de-represses neuronal differentiation genes, cell cycle inhibitors and tumor suppressors while reducing GLI1, PTCH1, MYC and EZH2. The sum of the effects of Compound A mediated EZH2 inhibition on several cancer pathways is the cause for the dramatic and permanent anti-tumor activity seen in MRT models. Thus, Compound A represents a new treatment modality for these lethal childhood tumors.
[0487] Furthermore, since several members of the SWI/SNF complex are genetically altered in other cancer types besides MRT, it is conceivable that EZH2 also plays a role in tumor maintenance and survival in a spectrum of cancer types. Combined with recent reports demonstrating the effectiveness of EZH2 inhibitors in selective killing of EZH2 mutant bearing non-Hodgkin lymphomas, the present data demonstrate that small molecule-based inhibition of EZH2 is an effective mechanism of therapeutic intervention in a variety of hematologic and solid tumors for which genetic alterations--either target-directed or indirect--confer a proliferative dependency on EZH2 enzymatic activity.
Example 3: Material and Methods
[0488] Cell Culture:
[0489] Cell lines 293T, RD, SJCRH30, A204, G401, G402, and KYM-1. 293T (CRL-11268), RD (CRL-136), SJCRH30 (CRL-2061), A204 (HTB-82), G401 (CRL-1441), and G402 (CRL-1440) were obtained from ATCC. KYM-1 (JCRB0627) was obtained from JCRB. 293T and RD cells were cultured in DMEM+10% FBS. SJCRH30 cells were cultured in RPMI+10% FBS. A204, G401, and G402 cells were cultured in McCoys 5a+10% FBS. KYM-1 cells were cultured in DMEM/Ham's F12+10% FBS.
[0490] Western Blots Analysis:
[0491] Histones were acid extracted as previously described (Daigle et al., Blood. 2013 Aug. 8; 122(6):1017-25). Western blots for acid extracted histones were performed as previously described (Knutson et al., Proc Natl Acad Sci USA. 2013 May 7; 110(19):7922-7). Whole cell lysates (WCL) were prepared using a modified RIPA buffer (10.times.RIPA Lysis Buffer (Millipore #20-188), 0.1% SDS (Invitrogen AM9823), protease mini-tablet (Roche #1836153)). Cells were pelleted, washed with ice cold PBS, resuspended in ice cold RIPA buffer, and incubated on ice for 5 minutes. Lysates were sonicated 3.times. for 10 sec at 50% power, then incubated on ice for 10 minutes. Lysates were then centrifuged at max speed for 15 minutes at 4 degrees in a table top centrifuge. Clarified lysates were aliquoted to a fresh tube, and protein concentrations for WCL were determined by BCA assay (Pierce). Ten micrograms of each lysate was fractionated on 10-20% Tris-Glycine gel (Biorad), transferred using iBlot (7 minutes on program 3, using Nitrocellulose transfer stacks), and probed with the following antibodies in Odyssey blocking buffer: SNF5 (CST #8745), EZH2 (CST #5246), and Beta-actin (CST #3700).
[0492] In Vitro Cell Assays:
[0493] For the adherent cell line proliferation assays (all cell lines except KYM-1, which was analyzed as previously described for suspension cell lines (Daigle et al., Blood. 2013 Aug. 8; 122(6):1017-25), plating densities for each cell line were determined based on growth curves (measured by ATP viability) and density over a 7 day timecourse. On the day before compound treatment, cells were plated in either 96-well plates in triplicate (for the day 0-7 timecourse) or 6-well plates (for replating on day 7 for the remainder of the timecourse). On Day 0, cells were either untreated, DMSO-treated, or treated with Compound A starting at 10 uM and decreasing in either 3- or 4-fold dilutions. Plates were read on Day 0, Day 4, and Day 7 using CellTiter-Glo.RTM. (Promega), with compound/media being replenished on Day 4. On Day 7, the 6-well plates were trypsinized, centrifuged, and resuspended in fresh media for counting by Vi-Cell. Cells from each treatment were replated at the original density in 96-well plates in triplicate. Cells were allowed to adhere to the plate overnight, and cells were treated as on Day 0. On Day 7, 11 and 14, plates were read using CellTiter-Glo.RTM., with compound/media being replenished on Day 11. Averages of triplicates were used to plot proliferation over the timecourse, and calculate IC50 values. For cell cycle and apoptosis, G401 and RD cells were plated in 15 cm dishes in duplicate at a density of 1.times.10.sup.6 cells per plate. Cells were incubated with Compound A at 1 uM, in a total of 25 mL, over a course of 14 days, with cells being split back to original plating density on day 4, 7, and 11. Cell cycle analysis and TUNEL assay were performed using a Guava.RTM. flow cytometer, following the manufacturer's protocol.
[0494] Gene Expression Analysis:
[0495] G401 and RD cells were plated in T-75 flasks at 175,000 cells/flask and 117,000 cells/flask respectively and allowed to adhere overnight. On Day 0, cells were treated in duplicates with DMSO or 1 uM Compound A. Cells were harvested and pelleted on Day 2, 4, and 7 with media and compound being replenished on Day 4. Tumor tissue from the G401 xenograft animals dosed for 21 days (vehicle, 125 mg/kg, and 250 mg/kg (6 animals each) and 500 mg/kg (4 animals) Compound A dose groups) were used for gene expression analysis. Total mRNA was extracted from cell pellets and tumor tissue using the RNeasy Mini Kit (Qiagen #74106) and reverse transcribed by the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems (AB) #4368813). RT-PCR was performed by ViiA.TM. 7 Real-Time PCR Systems (AB) using TaqMan Fast Advanced Master Mix (AB #4444964) and TaqMan primer/probe sets in table below. Gene expression was normalized to 18S (AB #Hs99999901_s1) and fold change was calculated using the .DELTA..DELTA.Ct method. For the in vivo samples, the average Ct value+/-SD was determined for each dose group and fold change compared to vehicle dose group was calculated using the .DELTA..DELTA.Ct method.
TABLE-US-00009 Gene AB# MYC Hs00153408_m1 EZH2 Hs00172783_m1 PTCH1 Hs00181117_m1 PROM1 Hs01009250_m1 (CD133) GLI1 Hs01110766_m1 DOCK4 Hs00206807_m1 PTPRK Hs00267788_m1 BIN1 Hs00184913_m1
[0496] ELISA:
[0497] Histones were isolated from tumors as previously described (Daigle et al) and were prepared in equivalent concentrations (0.5 ng/ul for H3 and 4 ng/ul for H3K27Me3) in coating buffer (PBS with 0.05% BSA). Sample or standard (100 .mu.L) was added in duplicate to two 96-well ELISA plates (Thermo Labsystems, Immulon 4HBX #3885). Histones isolated from G401 cells that were treated with DMSO or 10 .mu.mol/L Compound A for 4 days were added to control wells at the same histone concentration as the tumor histone samples. The plates were sealed and incubated overnight at 4.degree. C. The following day, plates were washed 3 times with 300 .mu.L/well PBST (PBS with 0.05% Tween 20; 10.times.PBST, KPL #51-14-02) on a Bio Tek plate washer. Plates were blocked with 300 .mu.L/well of diluent (PBS+2% BSA+0.05% Tween 20), incubated at room temperature for 2 hours, and washed 3 times with PBST. All antibodies were diluted in diluent. 100 uL/well of anti-H3K27Me3 (CST #9733, 50% glycerol stock 1:1000) or anti-total H3 (Abcam #ab1791, 50% glycerol stock 1:10,000) was added to each plate. Plates were incubated for 90 minutes at room temperature and washed 3 times with PBST. 100 .mu.L/well of anti-Rb-IgG-HRP (Cell Signaling Technology, 7074) was added 1:2000 to the H3K27Me3 plate and 1:6000 to the H3 plate and incubated for 90 minutes at room temperature. Plates were washed 4 times with PB ST. For detection, 100 .mu.L/well of TMB substrate (BioFx Laboratories, #TMBS) was added and plates incubated in the dark at room temperature for 5 minutes. Reaction was stopped with 100 .mu.L/well 1N H.sub.2SO.sub.4. Absorbance at 450 nm was read on SpectraMax M5 Microplate reader.
[0498] Xenograft Study:
[0499] All the procedures related to animal handling, care and the treatment in this study were performed according to the guidelines approved by the Institutional Animal Care and Use Committee (IACUC) of Shanghai Chemparner following the guidance of the Association for Assessment and Accreditation of Laboratory Animal Care (AAALAC). For the in vivo study, mice were inoculated subcutaneously at the right flank with G-401 tumor cells (5.times.10.sup.6/mouse) in 0.2 ml mixture of base media and Matrigel (McCoy's 5A:Matrigel=1:1) for tumor development. The treatments were started when the tumor size reached approximately 157 mm3 for the tumor efficacy study (n=16 mice per group). Compound A or vehicle (0.5% NaCMC+0.1% Tween-80 in water) was administered orally BID at a dose volume of 10 .mu.L/g for either 21 or 28 days. Animal body weights were measured every day during the first week, then twice weekly for the remainder of the study. Tumor size was measured twice weekly in two dimensions using a caliper, and the volume was expressed in mm.sup.3. For PK/PD analysis, 8 mice with the largest tumor burden were euthanized for tumor and blood collection after 21 days of dosing. The remaining mice continued dosing for one more week, and from day 29, treatment was stopped and the mice were enrolled in a tumor growth delay study. Mice were observed as individuals until they reached the tumor weight endpoint (2000 mm.sup.3) or until day 60 (whichever came first).
[0500] Pharmacokinetic Analyses:
[0501] Dexamethasone was used as internal standard. An aliquot of 30 .mu.L plasma sample was added with 30 .mu.L IS (Dexamethasone, 1000 ng/mL) and 150 .mu.L ACN. The mixture was vortexed for 5 min and centrifuged at 14000 rpm for 5 min. An aliquot of 2 .mu.L supernatant was injected for LC-MS/MS analysis (Q-trap 3200). For 10-fold diluted plasma samples an aliquot of 3 .mu.L plasma sample was added with 27 .mu.L blank plasma, the dilution factor was 10, then added with 30 .mu.L IS (Dexamethasone, 1000 ng/mL) and 150 .mu.L ACN. The mixture was vortexed for 5 min and centrifuged at 14000 rpm for 5 min. An aliquot of 2 .mu.L supernatant was injected for LC-MS/MS analysis. Tumor samples were homogenized on Beadbeater.RTM. for 30 seconds with 3.times.PBS (w/v) to obtain a tumor homogenate. An aliquot of 30 .mu.L tumor homogenate sample was added with 30 .mu.L IS (Dexamethasone, 1000 ng/mL) and 150 .mu.L ACN. The mixture was vortexed for 5 min and centrifuged at 14000 rpm for 5 min. An aliquot of 2 .mu.L supernatant was injected for LC-MS/MS analysis.
Example 4: General Experimental Procedures
NMR
[0502] .sup.1H-NMR spectra were taken using CDCl.sub.3 unless otherwise stated and were recorded at 400 or 500 MHz using a Varian or Oxford instruments magnet (500 MHz) instruments. Multiplicities indicated are s=singlet, d=doublet, t=triplet, q=quartet, quint=quintet, sxt=sextet, m=multiplet, dd=doublet of doublets, dt=doublet of triplets; br indicates a broad signal.
LCMS and HPLC
[0503] Shimadzu LC-Q, Shimadzu LCMS-2010EV or Waters Acquity Ultra Performance LC. HPLC: Products were analyzed by Shimadzu SPD-20A with 150.times.4.5 mm YMC ODS-M80 column or 150.times.4.6 mm YMC-Pack Pro C18 column at 1.0 ml/min.
[0504] Mobile phase was MeCN:H2O=3:2 (containing 0.3% SDS and 0.05% H.sub.3PO.sub.4),
[0505] 0.05% TFA in water, 0.05% TFA in acetonitrile (gradient Initial 20%, then 0.05% TFA/MeCN to conc. to 95% in 3 min. holds for 0.5 min. at 3.51 to 4.50 min then 0.05% TFA/MeCN conc. 20%).
[0506] Alternatively the LCMS, 2 different methods were used; the one we use the most is the high pH (METCR1600) and the other one for more standard compounds (METCR1416).
[0507] 0.1% Formic acid in water--Mobile phase "A" 0.1% Formic acid in acetonitrile--Mobile phase "B" utilizing Waters Atlantis dC18, 2.1 mm.times.100 mm, 3 .mu.m column, with a flow rate=0.6 ml/min Column temperature=40.degree. C.; Time (mins) % B 0.00 min 5% B. 5.0 mins 100% B, 5.4 mins 100% B and 0.42 mins 5% B
[0508] 3.5 minute method refers to Atlantis dC18, 2.1 mm.times.50 mm, 3 .mu.m column, flow rate of 1 ml/min at 40 C. Mobile phase A Formic acid (aq.) 0.1% mobile phase B formic acid (MeCN) 0.1%, injection 3 .mu.L, gradient 0 mins (5% organic), 2.5 min (100% organic), 2.7 mins (100% organic), 2.71 min (5% organic), 3.5 min (5% organic)
[0509] 7.0 minute method refers to Atlantis dC18, 2.1 mm.times.100 mm, 3 .mu.m column, flow rate of 0.6 ml/min at 40 C. Mobile phase A Formic acid (aq.) 0.1% mobile phase B formic acid (MeCN) 0.1%, injection 3 .mu.L, gradient 0 mins (5% organic), 5 min (100% organic), 5.4 mins (100% organic), 5.42 min (5% organic), 7 min (5% organic)
[0510] Both the 3. 5 and 7 minute methods were performed on a MS18 Shimadzu LCMS-2010EV or a MS19 Shimadzu LCMS-2010EV system utilizing LC-20AB pumps and SPD-M20A PDA detectors.
[0511] Products were purified by HPLC/MS using Waters AutoPurification System with 3100 Mass Detector.
[0512] HPLC analyses may also be performed on a Shimdazu LC-2010CHT using an YMC ODS-A, C18, (150.times.4.6.times.5 .mu.m) column at ambient temperature with a flow Rate of 1.4 ml/min. An injection volume of 10 .mu.l is utilized and detection occurs via UV/PDA. Mobile Phase A is 0.05 TFA in water and Mobile Phase B is 0.05 TFA in acetonitrile with a gradient program of Initial 5% B to 95% B in 8 min, hold for 1.5 min, at 9.51 to 12 min B. conc. 0.5%. The diluent is the mobile phase
Other
[0513] Automated flash column chromatography was performed on a Biotage Isolera version 4. 10 g SNAP cartridge running at 12 ml/min or a 25 g SNAP cartridge running at 25 ml/min and detecting at 254 nm and 280 nm.
[0514] Select Nitrile reductions may be performed on a ThalesNano H-Cube.RTM. according to the conditions described in the experimental procedure.
[0515] Other related general procedures can also be found in PCT publication No. WO12/118812, PCT application No. PCT/US2012/033648 and PCT application No. PCT/US2012/033662, each of which is incorporated herein by reference in its entirety.
Example 5: Synthesis of N-((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)-5-(ethyl (tetrahydro-2H-pyran-4-yl)amino)-4-methyl-4'-(morpholinomethyl)-[1,1'-bip- henyl]-3-carboxamide
##STR00041##
[0516] Step 1: Synthesis of 5-bromo-2-methyl-3-nitrobenzoic Acid
##STR00042##
[0518] To stirred solution of 2-methyl-3-nitrobenzoic acid (100 g, 552 mmol) in conc. H.sub.2SO.sub.4 (400 mL), 1,3-dibromo-5,5-dimethyl-2,4-imidazolidinedione (88 g, 308 mmol) was added in a portion wise manner at room temperature and the reaction mixture was then stirred at room temperature for 5 h. The reaction mixture was poured onto ice cold water, the precipitated solid was filtered off, washed with water and dried under vacuum to afford the desired compound as a solid (140 g, 98%). The isolated compound was taken directly into the next step. .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 8.31 (s, 1H), 8.17 (s, 1H), 2.43 (s, 3H).
Step 2: Synthesis of Methyl 5-bromo-2-methyl-3-nitrobenzoate
##STR00043##
[0520] To a stirred solution of 5-bromo-2-methyl-3-nitrobenzoic acid (285 g, 1105 mmol) in DMF (2.8 L) at room temperature was added sodium carbonate (468 g, 4415 mmol) followed by addition of methyl iodide (626.6 g, 4415 mmol). The resulting reaction mixture was heated at 60.degree. C. for 8 h. After completion (monitored by TLC), the reaction mixture was filtered (to remove sodium carbonate) and washed with ethyl acetate (1 L.times.3). The combined filtrate was washed with water (3 L.times.5) and the aqueous phase was back extracted with ethyl acetate (1 L.times.3). The combined organic layers were dried over anhydrous sodium sulfate, filtered and concentrated under reduced pressure to afford the title compound as a solid (290 g, 97% yield). The isolated compound was taken directly into the next step. .sup.1H NMR (CDCl.sub.3, 400 MHz) .delta. 8.17 (s, 1H), 7.91 (s, 1H), 3.96 (s, 3H), 2.59 (s, 3H).
Step 3: Synthesis of Methyl 3-amino-5-bromo-2-methylbenzoate
##STR00044##
[0522] To a stirred solution of methyl 5-bromo-2-methyl-3-nitrobenzoate (290 g, 1058 mmol) in ethanol (1.5 L) was added aqueous ammonium chloride (283 g, 5290 mmol dissolved in 1.5 L water). The resulting mixture was stirred at 80.degree. C. to which iron powder (472 g, 8451 mmol) was added in a portion wise manner. The resulting reaction mixture was heated at 80.degree. C. for 12 h. Upon completion as determined by TLC, the reaction mixture was hot filtered over Celite.RTM. and the celite bed was washed with methanol (5 L) followed by washing with 30% MeOH in DCM (5 L). The combined filtrate was concentrated in-vacuo, the residue obtained was diluted with aqueous sodium bicarbonate solution (2 L) and extracted with ethyl acetate (5 L.times.3). The combined organic layers were dried over anhydrous sodium sulfate, filtered and concentrated under reduced pressure to afford the title compound as a solid (220 g, 85%). The compound was taken directly into the next step. .sup.1H NMR (CDCl.sub.3, 400 MHz) .delta. 7.37 (s, 1H), 6.92 (s, 1H), 3.94 (s, 3H), 3.80 (bs, 2H), 2.31 (s, 3H).
Step 4: Synthesis of Methyl 5-bromo-2-methyl-3-((tetrahydro-2H-pyran-4-yl) amino) Benzoate
##STR00045##
[0524] To a stirred solution of methyl 3-amino-5-bromo-2-methylbenzoate (15 g, 61.5 mmol) and dihydro-2H-pyran-4(3)-one (9.2 g, 92 mmol) in dichloroethane (300 mL) was added acetic acid (22 g, 369 mmol) and the reaction mixture stirred at room temperature for 15 minutes, then the reaction mixture was cooled to 0.degree. C. and sodium triacetoxyborohydride (39 g, 184 mmol) was added. The reaction mixture was stirred overnight at room temperature. Upon completion of the reaction as determined by TLC, aqueous sodium bicarbonate solution was added to the reaction mixture until a pH of 7-8 was obtained. The organic phase was separated and the aqueous phase was extracted with ethyl acetate. The combined organic layers were dried over anhydrous sodium sulfate, filtered and concentrated under reduced pressure. The crude compound was purified by column chromatography (100-200 mesh silica gel) eluting with ethyl acetate:hexane to afford the desired compound as a solid (14 g, 69%). .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 7.01 (s, 1H), 6.98 (s, 1H), 5.00 (d, 1H, J=7.6 Hz), 3.84-3.87 (m, 2H), 3.79 (s, 3H), 3.54-3.56 (m, 1H), 3.43 (t, 2H, J=12 Hz), 2.14 (s, 3H), 1.81-1.84 (m, 2H), 1.47-1.55 (m, 2H).
Step 5: Synthesis of Methyl 5-bromo-3-(ethyl (tetrahydro-2H-pyran-4-yl) amino)-2-methylbenzoate
##STR00046##
[0526] To a stirred solution of methyl 5-bromo-2-methyl-3-((tetrahydro-2H-pyran-4-yl) amino) benzoate (14 g, 42.7 mmol) in dichloroethane (150 mL) was added acetaldehyde (3.75 g, 85.2 mmol) and acetic acid (15.3 g, 256 mmol). The resulting reaction mixture was stirred at room temperature for 15 minutes. The mixture was cooled to 0.degree. C. and sodium triacetoxyborohydride (27 g, 128 mmol) was added. The reaction mixture was stirred at room temperature for 3 hours. Upon completion of the reaction as determined by TLC, aqueous sodium bicarbonate solution was added to the reaction mixture until a pH 7-8 was obtained, the organic phase was separated and the aqueous phase was extracted with ethyl acetate. The combined organic layers were dried over anhydrous sodium sulfate, filtered and concentrated under reduced pressure. The crude compound was purified by column chromatography (100-200 mesh silica gel) eluting with ethyl acetate:hexane to afford the desired compound as a viscous liquid (14 g, 93%). .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 7.62 (s, 1H), 7.52 (s, 1H), 3.80 (bs, 5H), 3.31 (t, 2H), 2.97-3.05 (m, 2H), 2.87-2.96 (m, 1H), 2.38 (s, 3H), 1.52-1.61 (m, 2H), 1.37-1.50 (m, 2H), 0.87 (t, 3H, J=6.8 Hz).
Step 6: Synthesis of 5-bromo-N-((4, 6-dimethyl-2-oxo-1, 2-dihydropyridin-3-yl) methyl)-3-(ethyl (tetrahydro-2H-pyran-4-yl) amino)-2-methylbenzamide
##STR00047##
[0528] To a stirred solution of 5-bromo-3-(ethyl (tetrahydro-2H-pyran-4-yl) amino)-2-methylbenzoate (14 g, 39.4 mmol) in ethanol (100 mL) was added aqueous NaOH (2.36 g, 59.2 mmol in 25 mL water) and the resulting mixture was stirred at 60.degree. C. for 1 h. Upon completion of the reaction as determined by TLC, the solvent was removed under reduced pressure and the residue obtained was acidified with 1N HCl until a pH 7 was obtained and then aqueous citric acid solution was added until a pH 5-6 was obtained. The aqueous layer was extracted with 10% MeOH in DCM (200 mL.times.3), the combined organic layers were dried over anhydrous sodium sulfate, filtered and concentrated under reduced pressure to give the respective acid (14 g, 100%).
[0529] The above acid (14 g, 40.9 mmol) was then dissolved in DMSO (70 mL) and 3-(amino methyl)-4, 6-dimethylpyridin-2(1H)-one (12.4 g, 81.9 mmol) was added to it. The reaction mixture was stirred at room temperature for 15 minutes, then PYBOP (31.9 g, 61.4 mmol) was added and stirring was continued for overnight at room temperature. Upon completion of the reaction as determined by TLC, the reaction mixture was poured onto ice-cold water (700 mL), stirred for 30 minutes and the precipitated solid was collected by filtration, washed with water (500 mL) and air dried. The solid obtained was stirred with acetonitrile (75 mL.times.2), filtered and air dried. The solid obtained was again stirred with 5% MeOH in DCM (100 mL), filtered and dried completely under vacuum to afford the title compound as a solid (14 g, 74%). .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 11.47 (s, 1H), 8.23 (t, 1H), 7.30 (s, 1H), 7.08 (s, 1H), 5.85 (s, 1H), 4.23 (d, 2H, J=4.4 Hz), 3.81 (d, 2H, J=10.4 Hz), 3.20-3.26 (m, 2H), 3.00-3.07 (m, 1H), 2.91-2.96 (m, 2H), 2.18 (s, 3H), 2.14 (s, 3H), 2.10 (s, 3H), 1.58-1.60 (m, 2H), 1.45-1.50 (m, 2H), 0.78 (t, 3H, J=6.8 Hz).
Step 7: Synthesis of N-((4, 6-dimethyl-2-oxo-1, 2-dihydropyridin-3-yl) methyl)-5-(ethyl (tetrahydro-2H-pyran-4-yl) amino)-4-methyl-4'-(morpholinomethyl)-[1, 1'-biphenyl]-3-carboxamide
##STR00048##
[0531] To a stirred solution of 5-bromo-N-((4, 6-dimethyl-2-oxo-1, 2-dihydropyridin-3-yl) methyl)-3-(ethyl (tetrahydro-2H-pyran-4-yl) amino)-2-methylbenzamide (14 g, 29.5 mmol) in dioxane/water mixture (70 mL/14 mL) was added 4-(4-(4, 4, 5, 5-tetramethyl-1, 3, 2-dioxaborolan-2-yl) benzyl) morpholine (13.4 g, 44.2 mmol) followed by addition of Na.sub.2CO.sub.3 (11.2 g, 106.1 mmol). The solution was purged with argon for 15 minutes and then Pd (PPh.sub.3).sub.4 (3.40 g, 2.94 mmol) was added and the solution was again purged with argon for a further 10 min. The reaction mixture was heated at 100.degree. C. for 4 h. After completion (monitored by TLC), the reaction mixture was diluted with water and extracted with 10% MeOH/DCM. The combined organic layers were dried over anhydrous sodium sulphate, filtered and concentrated under reduced pressure. The crude compound was purified by column chromatography (100-200 mesh silica gel) eluting with methanol: DCM to the title compound as a solid (12 g, 71%). Analytical Data: LCMS: 573.35 (M+1).sup.+; HPLC: 99.5% (@ 254 nm) (R.sub.t; 3.999; Method: Column: YMC ODS-A 150 mm.times.4.6 mm.times.5.mu.; Mobile Phase: A; 0.05% TFA in water/B; 0.05% TFA in acetonitrile; Inj. Vol: 10 .mu.L, Col. Temp.: 30.degree. C.; Flow rate: 1.4 mL/min.; Gradient: 5% B to 95% B in 8 min, Hold for 1.5 min, 9.51-12 min 5% B); .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 11.46 (s, 1H), 8.19 (t, 1H), 7.57 (d, 2H, J=7.2 Hz), 7.36-7.39 (m, 3H), 7.21 (s, 1H), 5.85 (s, 1H), 4.28 (d, 2H, J=2.8 Hz), 3.82 (d, 2H, J=9.6 Hz), 3.57 (bs, 4H), 3.48 (s, 2H), 3.24 (t, 2H, J=10.8 Hz), 3.07-3.09 (m, 2H), 3.01 (m, 1H), 2.36 (m, 4H), 2.24 (s, 3H), 2.20 (s, 3H), 2.10 (s, 3H), 1.64-1.67 (m, 2H), 1.51-1.53 (m, 2H), 0.83 (t, 3H, J=6.4 Hz).
Step 8: Synthesis of N-((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)-5-(ethyl (tetrahydro-2H-pyran-4-yl)amino)-4-methyl-4'-(morpholinomethyl)-[1,1'-bip- henyl]-3-carboxamide Trihydrochloride
##STR00049##
[0533] N-((4, 6-dimethyl-2-oxo-1, 2-dihydropyridin-3-yl) methyl)-5-(ethyl (tetrahydro-2H-pyran-4-yl) amino)-4-methyl-4'-(morpholinomethyl)-[1, 1'-biphenyl]-3-carboxamide (12 g, 21.0 mmol) was dissolved in methanolic HCl (200 mL) and stirred at room temperature for 3 h. After three hours of stirring, the reaction mixture was concentrated under reduced pressure. The solid obtained was stirred with ether (100 mL.times.2) to afford the desired salt as a solid (11 g, 77%). Analytical Data of the tri-HCl salt: LCMS: 573.40 (M+1).sup.+; HPLC: 99.1% (@ 254 nm) (R.sub.t; 3.961; Method: Column: YMC ODS-A 150 mm.times.4.6 mm.times.5.mu.; Mobile Phase: A; 0.05% TFA in water/B; 0.05% TFA in acetonitrile; Inj. Vol: 10 .mu.L, Col. Temp.: 30.degree. C.; Flow rate: 1.4 mL/min.; Gradient: 5% B to 95% B in 8 min, Hold for 1.5 min, 9.51-12 min 5% B); .sup.1H NMR (D.sub.2O 400 MHz) .delta. 7.92 (bs, 1H,) 7.80 (s, 1H), 7.77 (d, 2H, J=8 Hz), 7.63 (s, 1H), 7.61 (s, 1H), 6.30 (s, 1H), 4.48 (s, 2H), 4.42 (s, 2H), 4.09-4.11 (m, 4H), 3.95-3.97 (m, 2H), 3.77 (t, 3H, J=10.4 Hz), 3.44-3.47 (m, 3H), 3.24-3.32 (m, 3H), 2.42 (s, 3H), 2.35 (s, 3H), 2.26 (s, 3H), 2.01 (m, 2H), 1.76 (m, 2H), 1.04 (t, 3H, J=6.8 Hz).
Example 6: N-((4, 6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)-5-(((1r,4r)-4-(dimethyla- mino)cyclohexyl)(ethyl)amino)-4-methyl-4'-(morpholinomethyl)-[1,1'-bipheny- l]-3-carboxamide
##STR00050##
[0534] Step 1: 5-bromo-2-methyl-3-nitrobenzoic Acid
[0535] To stirred solution of 2-methyl-3-nitrobenzoic acid (100 g, 552.48 mmol) in conc. H.sub.2SO.sub.4 (400 mL), 1,3-dibromo-5,5-dimethyl-2,4-imidazolidinedione (87.98 g, 307.70 mmol) was added in a portion-wise manner at room temperature. The reaction mixture was then stirred at room temperature for 5 h. The reaction mixture was poured into ice cold water, the precipitated solid collected by filtration, washed with water and dried under vacuum to afford desired 5-bromo-2-methyl-3-nitrobenzoic acid as off-white solid (140 g, 97.90% yield). .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 8.31 (s, 1H), 8.17 (s, 1H), 2.43 (s, 3H).
Step 2: methyl 5-bromo-2-methyl-3-nitrobenzoate
[0536] To a stirred solution of 5-bromo-2-methyl-3-nitrobenzoic acid (285 g, 1104.65 mmol) in DMF (2.8 L) was added sodium carbonate (468 g, 4415.09 mmol) followed by addition of methyl iodide (626.63 g, 4415 mmol) at room temperature. The resulting reaction mixture was stirred at 60.degree. C. for 8 h. The reaction mixture was then filtered to remove suspended solids which were washed well with ethyl acetate (3.times.1 L). The combined filtrates were washed well with water (5.times.3 L) and the aqueous phase back extracted with ethyl acetate (3.times.1 L). The combined organic extracts dried over anhydrous sodium sulfate, filtered and concentrated under reduced pressure to afford methyl 5-bromo-2-methyl-3-nitrobenzoate as an off-white solid (290 g, 97% yield). .sup.1H NMR (CDCl.sub.3, 400 MHz) .delta. 8.17 (s, 1H), 7.91 (s, 1H), 3.96 (s, 3H), 2.59 (s, 3H).
Step 3: Methyl 3-amino-5-bromo-2-methylbenzoate
[0537] To a stirred solution of methyl 5-bromo-2-methyl-3-nitrobenzoate (290 g, 1058.39 mmol) in ethanol (1.5 L) was added aqueous ammonium chloride (283 g, 5290 mmol dissolved in 1.5 L water). The resulting mixture was stirred and heated at 80.degree. C. followed by addition of iron powder (472 g, 8451 mmol) in portions at 80.degree. C. The resulting reaction mixture was heated at 80.degree. C. for 12 h. The reaction mixture was then hot filtered through Celite.RTM. and the Celite.RTM. bed washed well methanol (5 L) and then with 30% MeOH in DCM (5 L). The combined filtrates were concentrated in vacuo and the residue obtained was diluted with aqueous bicarbonate (2 L) and extracted with ethyl acetate (3.times.5 L). The combined organic layers were dried over anhydrous sodium sulfate, filtered and concentrated under reduced pressure to afford methyl 3-amino-5-bromo-2-methylbenzoate as a brown solid (220 g, 89.41% yield).
[0538] A portion of the product (5 g) was dissolved in hot ethanol (20 mL), insoluble residue filtered off and mother liquor concentrated to obtain methyl 3-amino-5-bromo-2-methylbenzoate (3.5 g, 70% yield) with HPLC purity 93.81% as light brown solid. .sup.1H NMR (CDCl.sub.3, 400 MHz) .delta. 7.37 (s, 1H), 6.92 (s, 1H), 3.94 (s, 3H), 3.80 (bs, 2H), 2.31 (s, 3H).
Step 4: Methyl 5-bromo-3-(((1r,4r)-4-((tert-butoxycarbonyl)amino)cyclohexyl)amino)-2-met- hylbenzoate
[0539] To a stirred solution of methyl 3-amino-5-bromo-2-methylbenzoate (5 g, 20.5 mmol) and tert-butyl (4-oxocyclohexyl)carbamate (5.69 g, 26.7 mmol) in dichloroethane (50 mL), acetic acid (7.4 g, 123 mmol) was added and the reaction was stirred at room temperature for 10 minutes. Sodium triacetoxyborohydride (13.1 g, 61.7 mmol) was then added at 0.degree. C. and reaction was stirred at room temperature for 16 hours. The reaction was quenched with aqueous sodium bicarbonate, the organic phase separated and the aqueous phase extracted with dichloromethane. The combined organic layers were dried over anhydrous sodium sulfate and concentrated in vacuo. The crude product was purified by silica gel column chromatography (100-200 mesh size) eluting with 10% ethyl acetate in hexane to afford 3.5 g of the more polar (trans) isomer, methyl 5-bromo-3-(((1r,4r)-4-((tert-butoxycarbonyl)amino)cyclohexyl)amino)-2-met- hylbenzoate, as solid (38.46%). .sup.1H NMR (CDCl.sub.3, 400 MHz) .delta. 7.21 (s, 1H), 6.80 (s, 1H), 4.41 (bs, 1H), 3.85 (s, 3H), 3.60 (m, 1H), 3.45 (m, 1H), 3.20 (m, 1H), 2.22 (s, 3H), 2.15 (bs, 2H), 2.05 (bs, 2H), 1.45 (s, 9H), 1.30 (m, 4H).
Step 5: methyl 5-bromo-3-(((1r,4r)-4-((tert-butoxycarbonyl)amino)cyclohexyl)-(ethyl)amin- o)-2-methylbenzoate
[0540] To a stirred solution of methyl 5-bromo-3-(((1r,4r)-4-((tert-butoxycarbonyl)amino)-cyclohexyl)(ethyl)amin- o)-2-methylbenzoate (55 g, 0.124 mol) and acetaldehyde (11 g, 0.25 mol) in dichloroethane (550 mL), acetic acid (44.64 g, 0.744 mol) was added and the reaction mixture stirred at room temperature for 10 minutes. Sodium triacetoxyborohydride (79 g, 0.372 mol) was then added at 0.degree. C. and the reaction mixture was stirred at room temperature for 16 hours. The reaction was quenched with aqueous sodium bicarbonate, the organic phase separated and the aqueous phase extracted with dichloromethane. The combined extracts were dried over anhydrous sodium sulfate and concentrated in-vacuo. The crude compound was purified by silica gel column chromatography (100-200 mesh size) eluting with 10% ethyl acetate in hexane to afford 44 g of methyl 5-bromo-3-(((1r,4r)-4-((tert-butoxycarbonyl)amino)cyclohexyl)-(ethyl)amin- o)-2-methylbenzoate (75.2%) as solid. .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 7.55 (s, 1H), 7.45 (s, 1H), 6.65 (d, 1H), 3.80 (s, 3H), 3.15 (bs, 1H), 3.05 (q, 2H), 2.60 (m, 1H), 2.30 (s, 3H), 1.75 (m, 4H), 1.40 (m, 2H), 1.35 (s, 9H), 1.10 (m, 2H), 0.80 (t, 3H).
Step 6: Tert-butyl ((1r,4r)-4-((5-bromo-3-(((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)meth- yl)carbamoyl)-2-methylphenyl)(ethyl)amino)cyclohexyl)carbamate
[0541] Aqueous NaOH (3.5 g, 0.08 mol in 10 mL H.sub.2O) was added to a solution of methyl 5-bromo-3-(((1r,4r)-4-((tert-butoxycarbonyl)amino)cyclohexyl)-(ethyl)amin- o)-2-methylbenzoate (25 g, 0.053 mol) in EtOH (100 mL) and stirred at 60.degree. C. for 1 h. The ethanol was then removed under reduced pressure and acidified to pH 8 with dilute HCl and to pH 6 with citric acid. The mixture was extracted with 10% methanol in DCM (3.times.200 mL). The combined organic layers were dried and concentrated giving the respective acid (24.2 g, 99.0%). .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 13.13 (s, 1H), 7.54 (s, 1H), 7.43 (s, 1H), 6.68 (d, 1H), 3.14 (bs, 1H), 3.03 (q, 2H), 2.56 (m, 1H), 2.33 (s, 3H), 1.80-1.65 (m, 4H), 1.40 (m, 2H), 1.35 (s, 9H), 1.10 (m, 2H), 0.77 (t, 3H).
[0542] The acid (24 g, 0.053 mol) was dissolved in DMSO (100 mL) and 3-(aminomethyl)-4,6-dimethylpyridin-2(1H)-one (16 g, 0.106 mol) and triethylamine (5.3 g, 0.053 mol) was added. The reaction mixture was stirred at room temperature for 15 min before PyBop (41 g, 0.079 mmol) was added and stirring was then continued for overnight at room temperature. The reaction mixture was poured into ice water (1 L). The resulting precipitate was collected by filtration, washed well with water (2.times.1 L) and dried. The product obtained was further purified by washings with acetonitrile (3.times.200 mL) and DCM (100 mL) to afford tert-butyl ((1r,4r)-4-((5-bromo-3-(((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)meth- yl)carbamoyl)-2-methylphenyl)(ethyl)amino)cyclohexyl)-carbamate (24 g, 77%). .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 11.47 (s, 1H), 8.24 (t, 1H), 7.25 (s, 1H), 7.04 (s, 1H), 6.67 (d, 1H), 5.85 (s, 1H), 4.24 (d, 2H), 3.13 (bs, 1H), 3.01 (q, 2H), 2.53 (m, 1H), 2.18 (s, 3H), 2.10 (s, 6H), 1.80-1.65 (m, 4H), 1.40 (m, 2H), 1.35 (s, 9H), 1.10 (m, 2H), 0.77 (t, 3H).
Step 7: Tert-butyl ((1r,4r)-4-((5-(((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)carba- moyl)-4-methyl-4'-(morpholinomethyl)-[1,1'-biphenyl]-3-yl)(ethyl)amino)cyc- lohexyl)carbamate
[0543] To a stirred solution of tert-butyl ((1r,4r)-4-((5-bromo-3-(((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)meth- yl)carbamoyl)-2-methylphenyl)(ethyl)amino)cyclohexyl)-carbamate (24 g, 0.041 mol) and 4-(4-(4,4,5,5-tetramethyl-1,3,2-dioxaborolan-2-yl)benzyl)morpholine (18 g, 0.061 mol) in dioxane/water mixture (160 mL+40 mL), Na.sub.2CO.sub.3 (15 g, 0.15 mol) was added and solution purged with argon for 15 min. Pd(PPh.sub.3).sub.4 (4.7 g, 0.041 mol) was then added and the reaction mixture again purged with argon for 10 min. The reaction mixture was heated at 100.degree. C. for 4 h. The reaction mixture was then diluted with 10% MeOH/DCM (500 mL) and filtered. The filtrate was concentrated, diluted with water (500 mL) and extracted with 10% MeOH in DCM (3.times.500 mL). The combined organic layers were dried over Na.sub.2SO.sub.4 and solvent removed under reduced pressure. The crude product was purified by silica gel column chromatography (100-200 mesh) eluting with 7% MeOH in DCM to afford tert-butyl ((1r,4r)-4-((5-(((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)carba- moyl)-4-methyl-4'-(morpholinomethyl)-[1,1'-biphenyl]-3-yl)(ethyl)amino)cyc- lohexyl)carbamate (20 g, 71.43%). .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 11.46 (s, 1H), 8.20 (t, 1H), 7.56 (d, 2H), 7.36 (m, 3H), 7.17 (s, 1H), 6.66 (d, 1H), 5.85 (s, 1H), 4.28 (d, 2H), 3.57 (bs, 4H), 3.48 (s, 2H), 3.20-3.05 (m, 3H), 2.62 (m, 1H), 2.36 (bs, 4H), 2.20 (s, 6H), 2.10 (s, 3H), 1.75 (m, 4H), 1.42 (m, 2H), 1.35 (s, 9H), 1.10 (m, 2H), 0.82 (t, 3H).
Step 8: 5-(((1r,4r)-4-aminocyclohexyl)(ethyl)amino)-N-((4,6-dimethyl-2-oxo- -1,2-dihydropyridin-3-yl)methyl)-4-methyl-4'-(morpholinomethyl)-[1,1'-biph- enyl]-3-carboxamide
[0544] To a stirred solution of tert-butyl ((1r,4r)-4-((5-(((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)carba- moyl)-4-methyl-4'-(morpholinomethyl)-[1,1'-biphenyl]-3-yl)(ethyl)amino)cyc- lohexyl)carbamate (20 g, 0.03 mol) in DCM (200 mL) at 0.degree. C., TFA (75 mL) was added and reaction was stirred for 2 h at room temperature. The reaction mixture was then concentrated to dryness and the residue basified with aqueous saturated bicarbonate solution (300 mL) to pH 8. The mixture was extracted with 20% methanol in DCM (4.times.200 m). The combined extracts were dried over Na.sub.2SO.sub.4 and the solvent removed under reduced pressure to afford 5-(((1r,4r)-4-aminocyclohexyl)(ethyl)amino)-N-((4,6-dimethyl-2-oxo-1,2-di- hydropyridin-3-yl)methyl)-4-methyl-4'-(morpholinomethyl)-[1,1'-biphenyl]-3- -carboxamide (15.5 g, 91%) which was used as is in the next reaction. .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 8.18 (bs, 1H), 7.57 (d, 2H), 7.38 (m, 3H), 7.20 (s, 1H), 5.85 (s, 1H), 4.29 (d, 2H), 3.57 (bs, 4H), 3.48 (s, 2H), 3.31 (bs, 2H), 3.10 (m, 2H), 2.91 (m, 1H), 2.67 (m, 1H), 2.36 (bs, 4H), 2.21 (s, 3H), 2.20 (s, 3H), 2.10 (s, 3H), 1.90 (m, 2H), 1.83 (m, 2H), 1.45 (m, 2H), 1.23 (m, 2H), 0.83 (t, 3H).
Step 9: N-((4,6-dimethyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)-5-(((1r,4r)- -4-(dimethylamino)cyclohexyl)(ethyl)amino)-4-methyl-4'-(morpholinomethyl)-- [1,1'-biphenyl]-3-carboxamide
[0545] To a stirred solution of 5-(((1r,4r)-4-aminocyclohexyl)(ethyl)amino)-N-((4,6-dimethyl-2-oxo-1,2-di- hydropyridin-3-yl)methyl)-4-methyl-4'-(morpholinomethyl)-[1,1'-biphenyl]-3- -carboxamide (14 g, 0.023 mol) in dichloromethane (150 mL) was added aqueous 35% formaldehyde solution (2.4 g, 0.080 mol) at 0.degree. C. After stirring for 20 min, Na(OAc).sub.3BH (12.2 g, 0.057 mol) was added and stirring continued for 2 h at 0.degree. C. Water (100 mL) was then added to the reaction mixture and the mixture extracted with 20% methanol in DCM (3.times.200 mL). The combined extracts were dried over Na.sub.2SO.sub.4 and the solvent removed under reduced pressure. The crude product was purified by basic alumina column chromatography eluting with 6-7% MeOH in DCM to afford the title compound (10 g, 63.6%). LCMS: 614.65 (M+1).sup.+; HPLC: 98.88% (@ 210-370 nm) (R.sub.t; 3.724; Method: Column: YMC ODS-A 150 mm.times.4.6 mm.times.5.mu.; Mobile Phase: A; 0.05% TFA in water/B; 0.05% TFA in acetonitrile; Inj. Vol: 10 .mu.L, Col. Temp.: 30.degree. C.; Flow rate: 1.4 mL/min.; Gradient: 5% B to 95% B in 8 min, Hold for 1.5 min, 9.51-12 min 5% B); .sup.1H NMR (DMSO-d.sub.6, 400 MHz) .delta. 11.45 (s, 1H), 8.17 (t, 1H), 7.56 (d, 2H, J=8 Hz), 7.36 (m, 3H), 7.17 (s, 1H), 5.85 (s, 1H), 4.29 (d, 2H, J=4.4 Hz), 3.57 (bs, 4H), 3.48 (s, 2H), 3.09 (q, 2H), 2.66 (m, 1H), 2.36 (bs, 4H), 2.21 (s, 3H), 2.20 (s, 3H), 2.11 (s, 9H), 1.79 (m, 4H), 1.36 (m, 2H), 1.11 (m, 2H), 0.82 (t, 3H, J=6.4&6.8 Hz).
Example 7: Bioassay Protocol and General Methods
Protocol for Wild-Type and Mutant PRC2 Enzyme Assays
[0546] General Materials.
[0547] S-adenosylmethionine (SAM), S-adenosylhomocyteine (SAH), bicine, KCl, Tween20, dimethylsulfoxide (DMSO) and bovine skin gelatin (BSG) were purchased from Sigma-Aldrich at the highest level of purity possible. Dithiothreitol (DTT) was purchased from EMD. .sup.3H-SAM was purchased from American Radiolabeled Chemicals with a specific activity of 80 Ci/mmol. 384-well streptavidin Flashplates were purchased from PerkinElmer.
[0548] Substrates.
[0549] Peptides representative of human histone H3 residues 21-44 containing either an unmodified lysine 27 (H3K27me0) or dimethylated lysine 27 (H3K27me2) were synthesized with a C-terminal G(K-biotin) linker-affinity tag motif and a C-terminal amide cap by 21.sup.st Century Biochemicals. The peptides were high-performance liquid chromatography (HPLC) purified to greater than 95% purity and confirmed by liquid chromatography mass spectrometry (LC-MS). The sequences are listed below.
TABLE-US-00010 H3K27me0: (SEQ ID NO: 13) ATKAARKSAPATGGVKKPHRYRPGGK(biotin)-amide H3K27me2: (SEQ ID NO: 14) ATKAARK(me2)SAPATGGVKKPHRYRPGGK(biotin)-amide
[0550] Chicken erythrocyte oligonucleosomes were purified from chicken blood according to established procedures.
[0551] Recombinant PRC2 Complexes.
[0552] Human PRC2 complexes were purified as 4-component enzyme complexes co-expressed in Spodoptera frugiperda (sf9) cells using a baculovirus expression system. The subunits expressed were wild-type EZH2 (NM_004456) or EZH2 Y641F, N, H, S or C mutants generated from the wild-type EZH2 construct, EED (NM_003797), Suz12 (NM_015355) and RbAp48 (NM_005610). The EED subunit contained an N-terminal FLAG tag that was used to purify the entire 4-component complex from sf9 cell lysates. The purity of the complexes met or exceeded 95% as determined by SDS-PAGE and Agilent Bioanalyzer analysis. Concentrations of enzyme stock concentrations (generally 0.3-1.0 mg/mL) was determined using a Bradford assay against a bovine serum albumin (BSA) standard.
[0553] General Procedure for PRC2 Enzyme Assays on Peptide Substrates.
[0554] The assays were all performed in a buffer consisting of 20 mM bicine (pH=7.6), 0.5 mM DTT, 0.005% BSG and 0.002% Tween20, prepared on the day of use. Compounds in 100% DMSO (1 .mu.L) were spotted into polypropylene 384-well V-bottom plates (Greiner) using a Platemate 2.times.3 outfitted with a 384-channel pipet head (Thermo). DMSO (1 .mu.L) was added to columns 11, 12, 23, 24, rows A-H for the maximum signal control, and SAH, a known product and inhibitor of PRC2 (1 .mu.L) was added to columns 11, 12, 23, 24, rows I-P for the minimum signal control. A cocktail (40 .mu.L) containing the wild-type PRC2 enzyme and H3K27me0 peptide or any of the Y641 mutant enzymes and H3K27me2 peptide was added by Multidrop Combi (Thermo). The compounds were allowed to incubate with PRC2 for 30 min at 25.degree. C., then a cocktail (10 .mu.L) containing a mixture of non-radioactive and .sup.3H-SAM was added to initiate the reaction (final volume=51 .mu.L). In all cases, the final concentrations were as follows: wild-type or mutant PRC2 enzyme was 4 nM, SAH in the minimum signal control wells was 1 mM and the DMSO concentration was 1%. The final concentrations of the rest of the components are indicated in Table 7, below. The assays were stopped by the addition of non-radioactive SAM (10 .mu.L) to a final concentration of 600 .mu.M, which dilutes the .sup.3H-SAM to a level where its incorporation into the peptide substrate is no longer detectable. 50 .mu.L of the reaction in the 384-well polypropylene plate was then transferred to a 384-well Flashplate and the biotinylated peptides were allowed to bind to the streptavidin surface for at least 1 h before being washed three times with 0.1% Tween20 in a Biotek ELx405 plate washer. The plates were then read in a PerkinElmer TopCount platereader to measure the quantity of .sup.3H-labeled peptide bound to the Flashplate surface, measured as disintegrations per minute (dpm) or alternatively, referred to as counts per minute (cpm).
TABLE-US-00011 TABLE 7 Final concentrations of components for each assay variation based upon EZH2 identity (wild-type or Y641 mutant EZH2) PRC2 Enzyme (denoted by EZH2 Non-radioactive identity) Peptide (nM) SAM (nM) .sup.3H-SAM (nM) Wild-type 185 1800 150 Y641F 200 850 150 Y641N 200 850 150 Y641H 200 1750 250 Y641S 200 1300 200 Y641C 200 3750 250
[0555] General Procedure for Wild-Type PRC2 Enzyme Assay on Oligonucleosome Substrate.
[0556] The assays was performed in a buffer consisting of 20 mM bicine (pH=7.6), 0.5 mM DTT, 0.005% BSG, 100 mM KCl and 0.002% Tween20, prepared on the day of use. Compounds in 100% DMSO (1 .mu.L) were spotted into polypropylene 384-well V-bottom plates (Greiner) using a Platemate 2.times.3 outfitted with a 384-channel pipet head (Thermo). DMSO (1 .mu.L) was added to columns 11, 12, 23, 24, rows A-H for the maximum signal control, and SAH, a known product and inhibitor of PRC2 (1 .mu.L) was added to columns 11, 12, 23, 24, rows I-P for the minimum signal control. A cocktail (40 .mu.L) containing the wild-type PRC2 enzyme and chicken erythrocyte oligonucleosome was added by Multidrop Combi (Thermo). The compounds were allowed to incubate with PRC2 for 30 min at 25.degree. C., then a cocktail (10 .mu.L) containing a mixture of non-radioactive and .sup.3H-SAM was added to initiate the reaction (final volume=51 .mu.L). The final concentrations were as follows: wild-type PRC2 enzyme was 4 nM, non-radioactive SAM was 430 nM, .sup.3H-SAM was 120 nM, chicken erythrocyte olignonucleosome was 120 nM, SAH in the minimum signal control wells was 1 mM and the DMSO concentration was 1%. The assay was stopped by the addition of non-radioactive SAM (10 .mu.L) to a final concentration of 600 .mu.M, which dilutes the .sup.3H-SAM to a level where its incorporation into the chicken erythrocyte olignonucleosome substrate is no longer detectable. 50 .mu.L of the reaction in the 384-well polypropylene plate was then transferred to a 384-well Flashplate and the chicken erythrocyte nucleosomes were immobilized to the surface of the plate, which was then washed three times with 0.1% Tween20 in a Biotek ELx405 plate washer. The plates were then read in a PerkinElmer TopCount platereader to measure the quantity of .sup.3H-labeled chicken erythrocyte oligonucleosome bound to the Flashplate surface, measured as disintegrations per minute (dpm) or alternatively, referred to as counts per minute (cpm).
% Inhibition Calculation ##EQU00001## % inh = 100 - ( dpm cmpd - dpm m i n dpm m ax - dpm m i n ) .times. 100 ##EQU00001.2##
[0557] Where dpm=disintegrations per minute, cmpd=signal in assay well, and min and max are the respective minimum and maximum signal controls.
Four - parameter IC 50 fit ##EQU00002## Y = Bottom + ( Top - Bottom ) 1 + ( X IC 50 ) Hill Coefficient ##EQU00002.2##
[0558] Where top and bottom are the normally allowed to float, but may be fixed at 100 or 0 respectively in a 3-parameter fit. The Hill Coefficient normally allowed to float but may also be fixed at 1 in a 3-parameter fit. Y is the % inhibition and X is the compound concentration.
[0559] IC.sub.50 values for the PRC2 enzyme assays on peptide substrates (e.g., EZH2 wild type and Y641F) are presented in Table 8 below.
[0560] WSU-DLCL2 Methylation Assay
[0561] WSU-DLCL2 suspension cells were purchased from DSMZ (German Collection of Microorganisms and Cell Cultures, Braunschweig, Germany). RPMI/Glutamax Medium, Penicillin-Streptomycin, Heat Inactivated Fetal Bovine Serum, and D-PBS were purchased from Life Technologies, Grand Island, N.Y., USA. Extraction Buffer and Neutralization Buffer (5.times.) were purchased from Active Motif, Carlsbad, Calif., USA. Rabbit anti-Histone H3 antibody was purchased from Abcam, Cambridge, Mass., USA. Rabbit anti-H3K27me3 and HRP-conjugated anti-rabbit-IgG were purchased from Cell Signaling Technology, Danvers, Mass., USA. TMB "Super Sensitive" substrate was sourced from BioFX Laboratories, Owings Mills, Md., USA. IgG-free Bovine Serum Albumin was purchased from Jackson ImmunoResearch, West Grove, Pa., USA. PBS with Tween (10.times.PBST) was purchased from KPL, Gaithersburg, Md., USA. Sulfuric Acid was purchased from Ricca Chemical, Arlington, Tex., USA. Immulon ELISA plates were purchased from Thermo, Rochester, N.Y., USA. V-bottom cell culture plates were purchased from Corning Inc., Corning, N.Y., USA. V-bottom polypropylene plates were purchased from Greiner Bio-One, Monroe, N.C., USA.
[0562] WSU-DLCL2 suspension cells were maintained in growth medium (RPMI 1640 supplemented with 10% v/v heat inactivated fetal bovine serum and 100 units/mL penicillin-streptomycin) and cultured at 37.degree. C. under 5% CO.sub.2. Under assay conditions, cells were incubated in Assay Medium (RPMI 1640 supplemented with 20% v/v heat inactivated fetal bovine serum and 100 units/mL penicillin-streptomycin) at 37.degree. C. under 5% CO.sub.2 on a plate shaker.
[0563] WSU-DLCL2 cells were seeded in assay medium at a concentration of 50,000 cells per mL to a 96-well V-bottom cell culture plate with 200 .mu.L per well. Compound (1 .mu.L) from 96 well source plates was added directly to V-bottom cell plate. Plates were incubated on a titer-plate shaker at 37.degree. C., 5% CO2 for 96 hours. After four days of incubation, plates were spun at 241.times.g for five minutes and medium was aspirated gently from each well of cell plate without disturbing cell pellet. Pellet was resuspended in 200 .mu.L DPBS and plates were spun again at 241.times.g for five minutes. The supernatant was aspirated and cold (4.degree. C.) Extraction buffer (100 .mu.L) was added per well. Plates were incubated at 4.degree. C. on orbital shaker for two hours. Plates were spun at 3427.times.g.times.10 minutes. Supernatant (80 .mu.L per well) was transferred to its respective well in 96 well V-bottom polypropylene plate. Neutralization Buffer 5.times. (20 per well) was added to V-bottom polypropylene plate containing supernatant. V-bottom polypropylene plates containing crude histone preparation (CHP) were incubated on orbital shaker x five minutes. Crude Histone Preparations were added (24, per well) to each respective well into duplicate 96 well ELISA plates containing 100 .mu.L Coating Buffer (1.times.PBS+BSA 0.05% w/v). Plates were sealed and incubated overnight at 4.degree. C. The following day, plates were washed three times with 300 .mu.L per well 1.times.PBST. Wells were blocked for two hours with 300 .mu.L per well ELISA Diluent ((PBS (1.times.) BSA (2% w/v) and Tween20 (0.05% v/v)). Plates were washed three times with 1.times.PBST. For the Histone H3 detection plate, 100 per well were added of anti-Histone-H3 antibody (Abcam, ab1791) diluted 1:10,000 in ELISA Diluent. For H3K27 trimethylation detection plate, 100 .mu.L per well were added of anti-H3K27me3 diluted 1:2000 in ELISA diluent. Plates were incubated for 90 minutes at room temperature. Plates were washed three times with 300 .mu.L 1.times.PBST per well. For Histone H3 detection, 100 .mu.L of HRP-conjugated anti-rabbit IgG antibody diluted to 1:6000 in ELISA diluent was added per well. For H3K27me3 detection, 100 .mu.L of HRP conjugated anti-rabbit IgG antibody diluted to 1:4000 in ELISA diluent was added per well. Plates were incubated at room temperature for 90 minutes. Plates were washed four times with 1.times.PBST 300 .mu.L per well. TMB substrate 100 .mu.L was added per well. Histone H3 plates were incubated for five minutes at room temperature. H3K27me3 plates were incubated for 10 minutes at room temperature. The reaction was stopped with sulfuric acid 1N (100 .mu.L per well). Absorbance for each plate was read at 450 nm.
[0564] First, the ratio for each well was determined by:
( H 3 K 27 me 3 OD 450 value Histone H 3 OD 450 value ) ##EQU00003##
[0565] Each plate included eight control wells of DMSO only treatment (Minimum Inhibition) as well as eight control wells for maximum inhibition (Background wells).
[0566] The average of the ratio values for each control type was calculated and used to determine the percent inhibition for each test well in the plate. Test compound was serially diluted three-fold in DMSO for a total of ten test concentrations, beginning at 25 .mu.M. Percent inhibition was determined and IC.sub.50 curves were generated using duplicate wells per concentration of compound. IC.sub.50 values for this assay are presented in Table 8 below.
Percent Inhibition = 100 - ( ( ( Individual Test Sample Ratio ) - ( Background Avg Ratio ) ( Minimum Inhibition Ratio ) - ( Background Average Ratio ) ) * 100 ) ##EQU00004##
[0567] Cell Proliferation Analysis
[0568] WSU-DLCL2 suspension cells were purchased from DSMZ (German Collection of Microorganisms and Cell Cultures, Braunschweig, Germany). RPMI/Glutamax Medium, Penicillin-Streptomycin, Heat Inactivated Fetal Bovine Serum were purchased from Life Technologies, Grand Island, N.Y., USA. V-bottom polypropylene 384-well plates were purchased from Greiner Bio-One, Monroe, N.C., USA. Cell culture 384-well white opaque plates were purchased from Perkin Elmer, Waltham, Mass., USA. Cell-Titer Glo.RTM. was purchased from Promega Corporation, Madison, Wis., USA. SpectraMax M5 plate reader was purchased from Molecular Devices LLC, Sunnyvale, Calif., USA.
[0569] WSU-DLCL2 suspension cells were maintained in growth medium (RPMI 1640 supplemented with 10% v/v heat inactivated fetal bovine serum and cultured at 37.degree. C. under 5% CO.sub.2. Under assay conditions, cells were incubated in Assay Medium (RPMI 1640 supplemented with 20% v/v heat inactivated fetal bovine serum and 100 units/mL penicillin-streptomycin) at 37.degree. C. under 5% CO.sub.2.
[0570] For the assessment of the effect of compounds on the proliferation of the WSU-DLCL2 cell line, exponentially growing cells were plated in 384-well white opaque plates at a density of 1250 cell/ml in a final volume of 50 .mu.l of assay medium. A compound source plate was prepared by performing triplicate nine-point 3-fold serial dilutions in DMSO, beginning at 10 mM (final top concentration of compound in the assay was 20 .mu.M and the DMSO was 0.2%). A 100 nL aliquot from the compound stock plate was added to its respective well in the cell plate. The 100% inhibition control consisted of cells treated with 200 nM final concentration of staurosporine and the 0% inhibition control consisted of DMSO treated cells. After addition of compounds, assay plates were incubated for 6 days at 37.degree. C., 5% CO.sub.2, relative humidity >90% for 6 days. Cell viability was measured by quantization of ATP present in the cell cultures, adding 35 .mu.l of CellTiter-Glo.RTM..RTM. reagent to the cell plates. Luminescence was read in the SpectraMax M5. The concentration inhibiting cell viability by 50% was determined using a 4-parametric fit of the normalized dose response curves.
INCORPORATION BY REFERENCE
[0571] The entire disclosure of each of the patent documents and scientific articles referred to herein is incorporated by reference for all purposes.
[0572] All publications and patent documents cited herein are incorporated herein by reference as if each such publication or document was specifically and individually indicated to be incorporated herein by reference. Citation of publications and patent documents is not intended as an admission that any is pertinent prior art, nor does it constitute any admission as to the contents or date of the same. The invention having now been described by way of written description, those of skill in the art will recognize that the invention can be practiced in a variety of embodiments and that the foregoing description and examples above are for purposes of illustration and not limitation of the claims that follow.
EQUIVALENTS
[0573] The invention can be embodied in other specific forms without departing from the spirit or essential characteristics thereof. The foregoing embodiments are therefore to be considered in all respects illustrative rather than limiting on the invention described herein. Scope of the invention is thus indicated by the appended claims rather than by the foregoing description, and all changes that come within the meaning and range of equivalency of the claims are intended to be embraced therein.
Sequence CWU 1
1
141385PRTHomo sapiens 1Met Met Met Met Ala Leu Ser Lys Thr Phe Gly
Gln Lys Pro Val Lys1 5 10 15Phe Gln Leu Glu Asp Asp Gly Glu Phe Tyr
Met Ile Gly Ser Glu Val 20 25 30Gly Asn Tyr Leu Arg Met Phe Arg Gly
Ser Leu Tyr Lys Arg Tyr Pro 35 40 45Ser Leu Trp Arg Arg Leu Ala Thr
Val Glu Glu Arg Lys Lys Ile Val 50 55 60Ala Ser Ser His Gly Lys Lys
Thr Lys Pro Asn Thr Lys Asp His Gly65 70 75 80Tyr Thr Thr Leu Ala
Thr Ser Val Thr Leu Leu Lys Ala Ser Glu Val 85 90 95Glu Glu Ile Leu
Asp Gly Asn Asp Glu Lys Tyr Lys Ala Val Ser Ile 100 105 110Ser Thr
Glu Pro Pro Thr Tyr Leu Arg Glu Gln Lys Ala Lys Arg Asn 115 120
125Ser Gln Trp Val Pro Thr Leu Pro Asn Ser Ser His His Leu Asp Ala
130 135 140Val Pro Cys Ser Thr Thr Ile Asn Arg Asn Arg Met Gly Arg
Asp Lys145 150 155 160Lys Arg Thr Phe Pro Leu Cys Phe Asp Asp His
Asp Pro Ala Val Ile 165 170 175His Glu Asn Ala Ser Gln Pro Glu Val
Leu Val Pro Ile Arg Leu Asp 180 185 190Met Glu Ile Asp Gly Gln Lys
Leu Arg Asp Ala Phe Thr Trp Asn Met 195 200 205Asn Glu Lys Leu Met
Thr Pro Glu Met Phe Ser Glu Ile Leu Cys Asp 210 215 220Asp Leu Asp
Leu Asn Pro Leu Thr Phe Val Pro Ala Ile Ala Ser Ala225 230 235
240Ile Arg Gln Gln Ile Glu Ser Tyr Pro Thr Asp Ser Ile Leu Glu Asp
245 250 255Gln Ser Asp Gln Arg Val Ile Ile Lys Leu Asn Ile His Val
Gly Asn 260 265 270Ile Ser Leu Val Asp Gln Phe Glu Trp Asp Met Ser
Glu Lys Glu Asn 275 280 285Ser Pro Glu Lys Phe Ala Leu Lys Leu Cys
Ser Glu Leu Gly Leu Gly 290 295 300Gly Glu Phe Val Thr Thr Ile Ala
Tyr Ser Ile Arg Gly Gln Leu Ser305 310 315 320Trp His Gln Lys Thr
Tyr Ala Phe Ser Glu Asn Pro Leu Pro Thr Val 325 330 335Glu Ile Ala
Ile Arg Asn Thr Gly Asp Ala Asp Gln Trp Cys Pro Leu 340 345 350Leu
Glu Thr Leu Thr Asp Ala Glu Met Glu Lys Lys Ile Arg Asp Gln 355 360
365Asp Arg Asn Thr Arg Arg Met Arg Arg Leu Ala Asn Thr Ala Pro Ala
370 375 380Trp38521717DNAHomo sapiens 2aacgccagcg cctgcgcact
gagggcggcc tggtcgtcgt ctgcggcggc ggcggcggct 60gaggagcccg gctgaggcgc
cagtacccgg cccggtccgc atttcgcctt ccggcttcgg 120tttccctcgg
cccagcacgc cccggccccg ccccagccct cctgatccct cgcagcccgg
180ctccggccgc ccgcctctgc cgccgcaatg atgatgatgg cgctgagcaa
gaccttcggg 240cagaagcccg tgaagttcca gctggaggac gacggcgagt
tctacatgat cggctccgag 300gtgggaaact acctccgtat gttccgaggt
tctctgtaca agagataccc ctcactctgg 360aggcgactag ccactgtgga
agagaggaag aaaatagttg catcgtcaca tggtaaaaaa 420acaaaaccta
acactaagga tcacggatac acgactctag ccaccagtgt gaccctgtta
480aaagcctcgg aagtggaaga gattctggat ggcaacgatg agaagtacaa
ggctgtgtcc 540atcagcacag agccccccac ctacctcagg gaacagaagg
ccaagaggaa cagccagtgg 600gtacccaccc tgcccaacag ctcccaccac
ttagatgccg tgccatgctc cacaaccatc 660aacaggaacc gcatgggccg
agacaagaag agaaccttcc ccctttgctt tgatgaccat 720gacccagctg
tgatccatga gaacgcatct cagcccgagg tgctggtccc catccggctg
780gacatggaga tcgatgggca gaagctgcga gacgccttca cctggaacat
gaatgagaag 840ttgatgacgc ctgagatgtt ttcagaaatc ctctgtgacg
atctggattt gaacccgctg 900acgtttgtgc cagccatcgc ctctgccatc
agacagcaga tcgagtccta ccccacggac 960agcatcctgg aggaccagtc
agaccagcgc gtcatcatca agctgaacat ccatgtggga 1020aacatttccc
tggtggacca gtttgagtgg gacatgtcag agaaggagaa ctcaccagag
1080aagtttgccc tgaagctgtg ctcggagctg gggttgggcg gggagtttgt
caccaccatc 1140gcatacagca tccggggaca gctgagctgg catcagaaga
cctacgcctt cagcgagaac 1200cctctgccca cagtggagat tgccatccgg
aacacgggcg atgcggacca gtggtgccca 1260ctgctggaga ctctgacaga
cgctgagatg gagaagaaga tccgcgacca ggacaggaac 1320acgaggcgga
tgaggcgtct tgccaacacg gccccggcct ggtaaccagc ccatcagcac
1380acggctccca cggagcatct cagaagattg ggccgcctct cctccatctt
ctggcaagga 1440cagaggcgag gggacagccc agcgccatcc tgaggatcgg
gtgggggtgg agtgggggct 1500tccaggtggc ccttcccggc acacattcca
tttgttgagc cccagtcctg ccccccaccc 1560caccctccct acccctcccc
agtctctggg gtcaggaaga aaccttattt taggttgtgt 1620tttgtttttg
tataggagcc ccaggcaggg ctagtaacag tttttaaata aaaggcaaca
1680ggtcatgttc aatttcttca acaaaaaaaa aaaaaaa 17173376PRTHomo
sapiens 3Met Met Met Met Ala Leu Ser Lys Thr Phe Gly Gln Lys Pro
Val Lys1 5 10 15Phe Gln Leu Glu Asp Asp Gly Glu Phe Tyr Met Ile Gly
Ser Glu Val 20 25 30Gly Asn Tyr Leu Arg Met Phe Arg Gly Ser Leu Tyr
Lys Arg Tyr Pro 35 40 45Ser Leu Trp Arg Arg Leu Ala Thr Val Glu Glu
Arg Lys Lys Ile Val 50 55 60Ala Ser Ser His Asp His Gly Tyr Thr Thr
Leu Ala Thr Ser Val Thr65 70 75 80Leu Leu Lys Ala Ser Glu Val Glu
Glu Ile Leu Asp Gly Asn Asp Glu 85 90 95Lys Tyr Lys Ala Val Ser Ile
Ser Thr Glu Pro Pro Thr Tyr Leu Arg 100 105 110Glu Gln Lys Ala Lys
Arg Asn Ser Gln Trp Val Pro Thr Leu Pro Asn 115 120 125Ser Ser His
His Leu Asp Ala Val Pro Cys Ser Thr Thr Ile Asn Arg 130 135 140Asn
Arg Met Gly Arg Asp Lys Lys Arg Thr Phe Pro Leu Cys Phe Asp145 150
155 160Asp His Asp Pro Ala Val Ile His Glu Asn Ala Ser Gln Pro Glu
Val 165 170 175Leu Val Pro Ile Arg Leu Asp Met Glu Ile Asp Gly Gln
Lys Leu Arg 180 185 190Asp Ala Phe Thr Trp Asn Met Asn Glu Lys Leu
Met Thr Pro Glu Met 195 200 205Phe Ser Glu Ile Leu Cys Asp Asp Leu
Asp Leu Asn Pro Leu Thr Phe 210 215 220Val Pro Ala Ile Ala Ser Ala
Ile Arg Gln Gln Ile Glu Ser Tyr Pro225 230 235 240Thr Asp Ser Ile
Leu Glu Asp Gln Ser Asp Gln Arg Val Ile Ile Lys 245 250 255Leu Asn
Ile His Val Gly Asn Ile Ser Leu Val Asp Gln Phe Glu Trp 260 265
270Asp Met Ser Glu Lys Glu Asn Ser Pro Glu Lys Phe Ala Leu Lys Leu
275 280 285Cys Ser Glu Leu Gly Leu Gly Gly Glu Phe Val Thr Thr Ile
Ala Tyr 290 295 300Ser Ile Arg Gly Gln Leu Ser Trp His Gln Lys Thr
Tyr Ala Phe Ser305 310 315 320Glu Asn Pro Leu Pro Thr Val Glu Ile
Ala Ile Arg Asn Thr Gly Asp 325 330 335Ala Asp Gln Trp Cys Pro Leu
Leu Glu Thr Leu Thr Asp Ala Glu Met 340 345 350Glu Lys Lys Ile Arg
Asp Gln Asp Arg Asn Thr Arg Arg Met Arg Arg 355 360 365Leu Ala Asn
Thr Ala Pro Ala Trp 370 37541690DNAHomo sapiens 4aacgccagcg
cctgcgcact gagggcggcc tggtcgtcgt ctgcggcggc ggcggcggct 60gaggagcccg
gctgaggcgc cagtacccgg cccggtccgc atttcgcctt ccggcttcgg
120tttccctcgg cccagcacgc cccggccccg ccccagccct cctgatccct
cgcagcccgg 180ctccggccgc ccgcctctgc cgccgcaatg atgatgatgg
cgctgagcaa gaccttcggg 240cagaagcccg tgaagttcca gctggaggac
gacggcgagt tctacatgat cggctccgag 300gtgggaaact acctccgtat
gttccgaggt tctctgtaca agagataccc ctcactctgg 360aggcgactag
ccactgtgga agagaggaag aaaatagttg catcgtcaca tgatcacgga
420tacacgactc tagccaccag tgtgaccctg ttaaaagcct cggaagtgga
agagattctg 480gatggcaacg atgagaagta caaggctgtg tccatcagca
cagagccccc cacctacctc 540agggaacaga aggccaagag gaacagccag
tgggtaccca ccctgcccaa cagctcccac 600cacttagatg ccgtgccatg
ctccacaacc atcaacagga accgcatggg ccgagacaag 660aagagaacct
tccccctttg ctttgatgac catgacccag ctgtgatcca tgagaacgca
720tctcagcccg aggtgctggt ccccatccgg ctggacatgg agatcgatgg
gcagaagctg 780cgagacgcct tcacctggaa catgaatgag aagttgatga
cgcctgagat gttttcagaa 840atcctctgtg acgatctgga tttgaacccg
ctgacgtttg tgccagccat cgcctctgcc 900atcagacagc agatcgagtc
ctaccccacg gacagcatcc tggaggacca gtcagaccag 960cgcgtcatca
tcaagctgaa catccatgtg ggaaacattt ccctggtgga ccagtttgag
1020tgggacatgt cagagaagga gaactcacca gagaagtttg ccctgaagct
gtgctcggag 1080ctggggttgg gcggggagtt tgtcaccacc atcgcataca
gcatccgggg acagctgagc 1140tggcatcaga agacctacgc cttcagcgag
aaccctctgc ccacagtgga gattgccatc 1200cggaacacgg gcgatgcgga
ccagtggtgc ccactgctgg agactctgac agacgctgag 1260atggagaaga
agatccgcga ccaggacagg aacacgaggc ggatgaggcg tcttgccaac
1320acggccccgg cctggtaacc agcccatcag cacacggctc ccacggagca
tctcagaaga 1380ttgggccgcc tctcctccat cttctggcaa ggacagaggc
gaggggacag cccagcgcca 1440tcctgaggat cgggtggggg tggagtgggg
gcttccaggt ggcccttccc ggcacacatt 1500ccatttgttg agccccagtc
ctgcccccca ccccaccctc cctacccctc cccagtctct 1560ggggtcagga
agaaacctta ttttaggttg tgttttgttt ttgtatagga gccccaggca
1620gggctagtaa cagtttttaa ataaaaggca acaggtcatg ttcaatttct
tcaacaaaaa 1680aaaaaaaaaa 169052492PRTHomo sapiens 5Met Thr Ala Glu
Pro Met Ser Glu Ser Lys Leu Asn Thr Leu Val Gln1 5 10 15Lys Leu His
Asp Phe Leu Ala His Ser Ser Glu Glu Ser Glu Glu Thr 20 25 30Ser Ser
Pro Pro Arg Leu Ala Met Asn Gln Asn Thr Asp Lys Ile Ser 35 40 45Gly
Ser Gly Ser Asn Ser Asp Met Met Glu Asn Ser Lys Glu Glu Gly 50 55
60Thr Ser Ser Ser Glu Lys Ser Lys Ser Ser Gly Ser Ser Arg Ser Lys65
70 75 80Arg Lys Pro Ser Ile Val Thr Lys Tyr Val Glu Ser Asp Asp Glu
Lys 85 90 95Pro Leu Asp Asp Glu Thr Val Asn Glu Asp Ala Ser Asn Glu
Asn Ser 100 105 110Glu Asn Asp Ile Thr Met Gln Ser Leu Pro Lys Gly
Thr Val Ile Val 115 120 125Gln Pro Glu Pro Val Leu Asn Glu Asp Lys
Asp Asp Phe Lys Gly Pro 130 135 140Glu Phe Arg Ser Arg Ser Lys Met
Lys Thr Glu Asn Leu Lys Lys Arg145 150 155 160Gly Glu Asp Gly Leu
His Gly Ile Val Ser Cys Thr Ala Cys Gly Gln 165 170 175Gln Val Asn
His Phe Gln Lys Asp Ser Ile Tyr Arg His Pro Ser Leu 180 185 190Gln
Val Leu Ile Cys Lys Asn Cys Phe Lys Tyr Tyr Met Ser Asp Asp 195 200
205Ile Ser Arg Asp Ser Asp Gly Met Asp Glu Gln Cys Arg Trp Cys Ala
210 215 220Glu Gly Gly Asn Leu Ile Cys Cys Asp Phe Cys His Asn Ala
Phe Cys225 230 235 240Lys Lys Cys Ile Leu Arg Asn Leu Gly Arg Lys
Glu Leu Ser Thr Ile 245 250 255Met Asp Glu Asn Asn Gln Trp Tyr Cys
Tyr Ile Cys His Pro Glu Pro 260 265 270Leu Leu Asp Leu Val Thr Ala
Cys Asn Ser Val Phe Glu Asn Leu Glu 275 280 285Gln Leu Leu Gln Gln
Asn Lys Lys Lys Ile Lys Val Asp Ser Glu Lys 290 295 300Ser Asn Lys
Val Tyr Glu His Thr Ser Arg Phe Ser Pro Lys Lys Thr305 310 315
320Ser Ser Asn Cys Asn Gly Glu Glu Lys Lys Leu Asp Asp Ser Cys Ser
325 330 335Gly Ser Val Thr Tyr Ser Tyr Ser Ala Leu Ile Val Pro Lys
Glu Met 340 345 350Ile Lys Lys Ala Lys Lys Leu Ile Glu Thr Thr Ala
Asn Met Asn Ser 355 360 365Ser Tyr Val Lys Phe Leu Lys Gln Ala Thr
Asp Asn Ser Glu Ile Ser 370 375 380Ser Ala Thr Lys Leu Arg Gln Leu
Lys Ala Phe Lys Ser Val Leu Ala385 390 395 400Asp Ile Lys Lys Ala
His Leu Ala Leu Glu Glu Asp Leu Asn Ser Glu 405 410 415Phe Arg Ala
Met Asp Ala Val Asn Lys Glu Lys Asn Thr Lys Glu His 420 425 430Lys
Val Ile Asp Ala Lys Phe Glu Thr Lys Ala Arg Lys Gly Glu Lys 435 440
445Pro Cys Ala Leu Glu Lys Lys Asp Ile Ser Lys Ser Glu Ala Lys Leu
450 455 460Ser Arg Lys Gln Val Asp Ser Glu His Met His Gln Asn Val
Pro Thr465 470 475 480Glu Glu Gln Arg Thr Asn Lys Ser Thr Gly Gly
Glu His Lys Lys Ser 485 490 495Asp Arg Lys Glu Glu Pro Gln Tyr Glu
Pro Ala Asn Thr Ser Glu Asp 500 505 510Leu Asp Met Asp Ile Val Ser
Val Pro Ser Ser Val Pro Glu Asp Ile 515 520 525Phe Glu Asn Leu Glu
Thr Ala Met Glu Val Gln Ser Ser Val Asp His 530 535 540Gln Gly Asp
Gly Ser Ser Gly Thr Glu Gln Glu Val Glu Ser Ser Ser545 550 555
560Val Lys Leu Asn Ile Ser Ser Lys Asp Asn Arg Gly Gly Ile Lys Ser
565 570 575Lys Thr Thr Ala Lys Val Thr Lys Glu Leu Tyr Val Lys Leu
Thr Pro 580 585 590Val Ser Leu Ser Asn Ser Pro Ile Lys Gly Ala Asp
Cys Gln Glu Val 595 600 605Pro Gln Asp Lys Asp Gly Tyr Lys Ser Cys
Gly Leu Asn Pro Lys Leu 610 615 620Glu Lys Cys Gly Leu Gly Gln Glu
Asn Ser Asp Asn Glu His Leu Val625 630 635 640Glu Asn Glu Val Ser
Leu Leu Leu Glu Glu Ser Asp Leu Arg Arg Ser 645 650 655Pro Arg Val
Lys Thr Thr Pro Leu Arg Arg Pro Thr Glu Thr Asn Pro 660 665 670Val
Thr Ser Asn Ser Asp Glu Glu Cys Asn Glu Thr Val Lys Glu Lys 675 680
685Gln Lys Leu Ser Val Pro Val Arg Lys Lys Asp Lys Arg Asn Ser Ser
690 695 700Asp Ser Ala Ile Asp Asn Pro Lys Pro Asn Lys Leu Pro Lys
Ser Lys705 710 715 720Gln Ser Glu Thr Val Asp Gln Asn Ser Asp Ser
Asp Glu Met Leu Ala 725 730 735Ile Leu Lys Glu Val Ser Arg Met Ser
His Ser Ser Ser Ser Asp Thr 740 745 750Asp Ile Asn Glu Ile His Thr
Asn His Lys Thr Leu Tyr Asp Leu Lys 755 760 765Thr Gln Ala Gly Lys
Asp Asp Lys Gly Lys Arg Lys Arg Lys Ser Ser 770 775 780Thr Ser Gly
Ser Asp Phe Asp Thr Lys Lys Gly Lys Ser Ala Lys Ser785 790 795
800Ser Ile Ile Ser Lys Lys Lys Arg Gln Thr Gln Ser Glu Ser Ser Asn
805 810 815Tyr Asp Ser Glu Leu Glu Lys Glu Ile Lys Ser Met Ser Lys
Ile Gly 820 825 830Ala Ala Arg Thr Thr Lys Lys Arg Ile Pro Asn Thr
Lys Asp Phe Asp 835 840 845Ser Ser Glu Asp Glu Lys His Ser Lys Lys
Gly Met Asp Asn Gln Gly 850 855 860His Lys Asn Leu Lys Thr Ser Gln
Glu Gly Ser Ser Asp Asp Ala Glu865 870 875 880Arg Lys Gln Glu Arg
Glu Thr Phe Ser Ser Ala Glu Gly Thr Val Asp 885 890 895Lys Asp Thr
Thr Ile Met Glu Leu Arg Asp Arg Leu Pro Lys Lys Gln 900 905 910Gln
Ala Ser Ala Ser Thr Asp Gly Val Asp Lys Leu Ser Gly Lys Glu 915 920
925Gln Ser Phe Thr Ser Leu Glu Val Arg Lys Val Ala Glu Thr Lys Glu
930 935 940Lys Ser Lys His Leu Lys Thr Lys Thr Cys Lys Lys Val Gln
Asp Gly945 950 955 960Leu Ser Asp Ile Ala Glu Lys Phe Leu Lys Lys
Asp Gln Ser Asp Glu 965 970 975Thr Ser Glu Asp Asp Lys Lys Gln Ser
Lys Lys Gly Thr Glu Glu Lys 980 985 990Lys Lys Pro Ser Asp Phe Lys
Lys Lys Val Ile Lys Met Glu Gln Gln 995 1000 1005Tyr Glu Ser Ser
Ser Asp Gly Thr Glu Lys Leu Pro Glu Arg Glu 1010 1015 1020Glu Ile
Cys His Phe Pro Lys Gly Ile Lys Gln Ile Lys Asn Gly 1025 1030
1035Thr Thr Asp Gly Glu Lys Lys Ser Lys Lys Ile Arg Asp Lys Thr
1040 1045 1050Ser Lys Lys Lys Asp Glu Leu Ser Asp Tyr Ala Glu Lys
Ser Thr 1055 1060 1065Gly Lys Gly Asp Ser Cys Asp Ser Ser Glu Asp
Lys Lys Ser Lys 1070 1075 1080Asn Gly Ala Tyr Gly Arg Glu Lys Lys
Arg Cys Lys Leu Leu Gly 1085 1090 1095Lys Ser Ser Arg Lys Arg Gln
Asp Cys Ser Ser Ser Asp Thr Glu 1100 1105 1110Lys Tyr Ser Met Lys
Glu Asp Gly Cys Asn Ser Ser Asp Lys Arg 1115 1120
1125Leu Lys Arg Ile Glu Leu Arg Glu Arg Arg Asn Leu Ser Ser Lys
1130 1135 1140Arg Asn Thr Lys Glu Ile Gln Ser Gly Ser Ser Ser Ser
Asp Ala 1145 1150 1155Glu Glu Ser Ser Glu Asp Asn Lys Lys Lys Lys
Gln Arg Thr Ser 1160 1165 1170Ser Lys Lys Lys Ala Val Ile Val Lys
Glu Lys Lys Arg Asn Ser 1175 1180 1185Leu Arg Thr Ser Thr Lys Arg
Lys Gln Ala Asp Ile Thr Ser Ser 1190 1195 1200Ser Ser Ser Asp Ile
Glu Asp Asp Asp Gln Asn Ser Ile Gly Glu 1205 1210 1215Gly Ser Ser
Asp Glu Gln Lys Ile Lys Pro Val Thr Glu Asn Leu 1220 1225 1230Val
Leu Ser Ser His Thr Gly Phe Cys Gln Ser Ser Gly Asp Glu 1235 1240
1245Ala Leu Ser Lys Ser Val Pro Val Thr Val Asp Asp Asp Asp Asp
1250 1255 1260Asp Asn Asp Pro Glu Asn Arg Ile Ala Lys Lys Met Leu
Leu Glu 1265 1270 1275Glu Ile Lys Ala Asn Leu Ser Ser Asp Glu Asp
Gly Ser Ser Asp 1280 1285 1290Asp Glu Pro Glu Glu Gly Lys Lys Arg
Thr Gly Lys Gln Asn Glu 1295 1300 1305Glu Asn Pro Gly Asp Glu Glu
Ala Lys Asn Gln Val Asn Ser Glu 1310 1315 1320Ser Asp Ser Asp Ser
Glu Glu Ser Lys Lys Pro Arg Tyr Arg His 1325 1330 1335Arg Leu Leu
Arg His Lys Leu Thr Val Ser Asp Gly Glu Ser Gly 1340 1345 1350Glu
Glu Lys Lys Thr Lys Pro Lys Glu His Lys Glu Val Lys Gly 1355 1360
1365Arg Asn Arg Arg Lys Val Ser Ser Glu Asp Ser Glu Asp Ser Asp
1370 1375 1380Phe Gln Glu Ser Gly Val Ser Glu Glu Val Ser Glu Ser
Glu Asp 1385 1390 1395Glu Gln Arg Pro Arg Thr Arg Ser Ala Lys Lys
Ala Glu Leu Glu 1400 1405 1410Glu Asn Gln Arg Ser Tyr Lys Gln Lys
Lys Lys Arg Arg Arg Ile 1415 1420 1425Lys Val Gln Glu Asp Ser Ser
Ser Glu Asn Lys Ser Asn Ser Glu 1430 1435 1440Glu Glu Glu Glu Glu
Lys Glu Glu Glu Glu Glu Glu Glu Glu Glu 1445 1450 1455Glu Glu Glu
Glu Glu Glu Asp Glu Asn Asp Asp Ser Lys Ser Pro 1460 1465 1470Gly
Lys Gly Arg Lys Lys Ile Arg Lys Ile Leu Lys Asp Asp Lys 1475 1480
1485Leu Arg Thr Glu Thr Gln Asn Ala Leu Lys Glu Glu Glu Glu Arg
1490 1495 1500Arg Lys Arg Ile Ala Glu Arg Glu Arg Glu Arg Glu Lys
Leu Arg 1505 1510 1515Glu Val Ile Glu Ile Glu Asp Ala Ser Pro Thr
Lys Cys Pro Ile 1520 1525 1530Thr Thr Lys Leu Val Leu Asp Glu Asp
Glu Glu Thr Lys Glu Pro 1535 1540 1545Leu Val Gln Val His Arg Asn
Met Val Ile Lys Leu Lys Pro His 1550 1555 1560Gln Val Asp Gly Val
Gln Phe Met Trp Asp Cys Cys Cys Glu Ser 1565 1570 1575Val Lys Lys
Thr Lys Lys Ser Pro Gly Ser Gly Cys Ile Leu Ala 1580 1585 1590His
Cys Met Gly Leu Gly Lys Thr Leu Gln Val Val Ser Phe Leu 1595 1600
1605His Thr Val Leu Leu Cys Asp Lys Leu Asp Phe Ser Thr Ala Leu
1610 1615 1620Val Val Cys Pro Leu Asn Thr Ala Leu Asn Trp Met Asn
Glu Phe 1625 1630 1635Glu Lys Trp Gln Glu Gly Leu Lys Asp Asp Glu
Lys Leu Glu Val 1640 1645 1650Ser Glu Leu Ala Thr Val Lys Arg Pro
Gln Glu Arg Ser Tyr Met 1655 1660 1665Leu Gln Arg Trp Gln Glu Asp
Gly Gly Val Met Ile Ile Gly Tyr 1670 1675 1680Glu Met Tyr Arg Asn
Leu Ala Gln Gly Arg Asn Val Lys Ser Arg 1685 1690 1695Lys Leu Lys
Glu Ile Phe Asn Lys Ala Leu Val Asp Pro Gly Pro 1700 1705 1710Asp
Phe Val Val Cys Asp Glu Gly His Ile Leu Lys Asn Glu Ala 1715 1720
1725Ser Ala Val Ser Lys Ala Met Asn Ser Ile Arg Ser Arg Arg Arg
1730 1735 1740Ile Ile Leu Thr Gly Thr Pro Leu Gln Asn Asn Leu Ile
Glu Tyr 1745 1750 1755His Cys Met Val Asn Phe Ile Lys Glu Asn Leu
Leu Gly Ser Ile 1760 1765 1770Lys Glu Phe Arg Asn Arg Phe Ile Asn
Pro Ile Gln Asn Gly Gln 1775 1780 1785Cys Ala Asp Ser Thr Met Val
Asp Val Arg Val Met Lys Lys Arg 1790 1795 1800Ala His Ile Leu Tyr
Glu Met Leu Ala Gly Cys Val Gln Arg Lys 1805 1810 1815Asp Tyr Thr
Ala Leu Thr Lys Phe Leu Pro Pro Lys His Glu Tyr 1820 1825 1830Val
Leu Ala Val Arg Met Thr Ser Ile Gln Cys Lys Leu Tyr Gln 1835 1840
1845Tyr Tyr Leu Asp His Leu Thr Gly Val Gly Asn Asn Ser Glu Gly
1850 1855 1860Gly Arg Gly Lys Ala Gly Ala Lys Leu Phe Gln Asp Phe
Gln Met 1865 1870 1875Leu Ser Arg Ile Trp Thr His Pro Trp Cys Leu
Gln Leu Asp Tyr 1880 1885 1890Ile Ser Lys Glu Asn Lys Gly Tyr Phe
Asp Glu Asp Ser Met Asp 1895 1900 1905Glu Phe Ile Ala Ser Asp Ser
Asp Glu Thr Ser Met Ser Leu Ser 1910 1915 1920Ser Asp Asp Tyr Thr
Lys Lys Lys Lys Lys Gly Lys Lys Gly Lys 1925 1930 1935Lys Asp Ser
Ser Ser Ser Gly Ser Gly Ser Asp Asn Asp Val Glu 1940 1945 1950Val
Ile Lys Val Trp Asn Ser Arg Ser Arg Gly Gly Gly Glu Gly 1955 1960
1965Asn Val Asp Glu Thr Gly Asn Asn Pro Ser Val Ser Leu Lys Leu
1970 1975 1980Glu Glu Ser Lys Ala Thr Ser Ser Ser Asn Pro Ser Ser
Pro Ala 1985 1990 1995Pro Asp Trp Tyr Lys Asp Phe Val Thr Asp Ala
Asp Ala Glu Val 2000 2005 2010Leu Glu His Ser Gly Lys Met Val Leu
Leu Phe Glu Ile Leu Arg 2015 2020 2025Met Ala Glu Glu Ile Gly Asp
Lys Val Leu Val Phe Ser Gln Ser 2030 2035 2040Leu Ile Ser Leu Asp
Leu Ile Glu Asp Phe Leu Glu Leu Ala Ser 2045 2050 2055Arg Glu Lys
Thr Glu Asp Lys Asp Lys Pro Leu Ile Tyr Lys Gly 2060 2065 2070Glu
Gly Lys Trp Leu Arg Asn Ile Asp Tyr Tyr Arg Leu Asp Gly 2075 2080
2085Ser Thr Thr Ala Gln Ser Arg Lys Lys Trp Ala Glu Glu Phe Asn
2090 2095 2100Asp Glu Thr Asn Val Arg Gly Arg Leu Phe Ile Ile Ser
Thr Lys 2105 2110 2115Ala Gly Ser Leu Gly Ile Asn Leu Val Ala Ala
Asn Arg Val Ile 2120 2125 2130Ile Phe Asp Ala Ser Trp Asn Pro Ser
Tyr Asp Ile Gln Ser Ile 2135 2140 2145Phe Arg Val Tyr Arg Phe Gly
Gln Thr Lys Pro Val Tyr Val Tyr 2150 2155 2160Arg Phe Leu Ala Gln
Gly Thr Met Glu Asp Lys Ile Tyr Asp Arg 2165 2170 2175Gln Val Thr
Lys Gln Ser Leu Ser Phe Arg Val Val Asp Gln Gln 2180 2185 2190Gln
Val Glu Arg His Phe Thr Met Asn Glu Leu Thr Glu Leu Tyr 2195 2200
2205Thr Phe Glu Pro Asp Leu Leu Asp Asp Pro Asn Ser Glu Lys Lys
2210 2215 2220Lys Lys Arg Asp Thr Pro Met Leu Pro Lys Asp Thr Ile
Leu Ala 2225 2230 2235Glu Leu Leu Gln Ile His Lys Glu His Ile Val
Gly Tyr His Glu 2240 2245 2250His Asp Ser Leu Leu Asp His Lys Glu
Glu Glu Glu Leu Thr Glu 2255 2260 2265Glu Glu Arg Lys Ala Ala Trp
Ala Glu Tyr Glu Ala Glu Lys Lys 2270 2275 2280Gly Leu Thr Met Arg
Phe Asn Ile Pro Thr Gly Thr Asn Leu Pro 2285 2290 2295Pro Val Ser
Phe Asn Ser Gln Thr Pro Tyr Ile Pro Phe Asn Leu 2300 2305 2310Gly
Ala Leu Ser Ala Met Ser Asn Gln Gln Leu Glu Asp Leu Ile 2315 2320
2325Asn Gln Gly Arg Glu Lys Val Val Glu Ala Thr Asn Ser Val Thr
2330 2335 2340Ala Val Arg Ile Gln Pro Leu Glu Asp Ile Ile Ser Ala
Val Trp 2345 2350 2355Lys Glu Asn Met Asn Leu Ser Glu Ala Gln Val
Gln Ala Leu Ala 2360 2365 2370Leu Ser Arg Gln Ala Ser Gln Glu Leu
Asp Val Lys Arg Arg Glu 2375 2380 2385Ala Ile Tyr Asn Asp Val Leu
Thr Lys Gln Gln Met Leu Ile Ser 2390 2395 2400Cys Val Gln Arg Ile
Leu Met Asn Arg Arg Leu Gln Gln Gln Tyr 2405 2410 2415Asn Gln Gln
Gln Gln Gln Gln Met Thr Tyr Gln Gln Ala Thr Leu 2420 2425 2430Gly
His Leu Met Met Pro Lys Pro Pro Asn Leu Ile Met Asn Pro 2435 2440
2445Ser Asn Tyr Gln Gln Ile Asp Met Arg Gly Met Tyr Gln Pro Val
2450 2455 2460Ala Gly Gly Met Gln Pro Pro Pro Leu Gln Arg Ala Pro
Pro Pro 2465 2470 2475Met Arg Ser Lys Asn Pro Gly Pro Ser Gln Gly
Lys Ser Met 2480 2485 2490611202DNAHomo sapiens 6aattctcctg
cctgagcctc ggcccaacaa aatggcggcg gcagcggtgt cgctttgttt 60ccgcggctcc
tgcggcggtg gcagtggtag cggcctttga gctgtgggga ggttccagca
120gcagctacag tgacgactaa gactccagtg catttctatc gtaaccgggc
gcgggggagc 180gcagatcggc gcccagcaat cacagaagcc gacaaggcgt
tcaagcgaaa acatgaccgc 240tgagcccatg agtgaaagca agttgaatac
attggtgcag aagcttcatg acttccttgc 300acactcatca gaagaatctg
aagaaacaag ttctcctcca cgacttgcaa tgaatcaaaa 360cacagataaa
atcagtggtt ctggaagtaa ctctgatatg atggaaaaca gcaaggaaga
420gggaactagc tcttcagaaa aatccaagtc ttcaggatcg tcacgatcaa
agaggaaacc 480ttcaattgta acaaagtatg tagaatcaga tgatgaaaaa
cctttggatg atgaaactgt 540aaatgaagat gcgtctaatg aaaattcaga
aaatgatatt actatgcaga gcttgccaaa 600aggtacagtg attgtacagc
cagagccagt gctgaatgaa gacaaagatg attttaaagg 660gcctgaattt
agaagcagaa gtaaaatgaa aactgaaaat ctcaaaaaac gcggagaaga
720tgggcttcat gggattgtga gctgcactgc ttgtggacaa caggtcaatc
attttcaaaa 780agattccatt tatagacacc cttcattgca agttcttatt
tgtaagaatt gctttaagta 840ttacatgagt gatgatatta gccgtgactc
agatggaatg gatgaacaat gtaggtggtg 900tgcggaaggt ggaaacttga
tttgttgtga cttttgccat aatgctttct gcaagaaatg 960cattctacgc
aaccttggtc gaaaggagtt gtccacaata atggatgaaa acaaccaatg
1020gtattgctac atttgtcacc cagagccttt gttggacttg gtcactgcat
gtaacagcgt 1080atttgagaat ttagaacagt tgttgcagca aaataagaag
aagataaaag ttgacagtga 1140aaagagtaat aaagtatatg aacatacatc
cagattttct ccaaagaaga ctagttcaaa 1200ttgtaatgga gaagaaaaga
aattagatga ttcctgttct ggctctgtaa cctactctta 1260ttccgcacta
attgtgccca aagagatgat taagaaggca aaaaaactga ttgagaccac
1320agccaacatg aactccagtt atgttaaatt tttaaagcag gcaacagata
attcagaaat 1380cagttctgct acaaaattac gtcagcttaa ggcttttaag
tctgtgttgg ctgatattaa 1440gaaggctcat cttgcattgg aagaagactt
aaattccgag tttcgagcga tggatgctgt 1500aaacaaagag aaaaatacca
aagagcataa agtcatagat gctaagtttg aaacaaaagc 1560acgaaaagga
gaaaaacctt gtgctttgga aaagaaggat atttcaaagt cagaagctaa
1620actttcaaga aaacaggtag atagtgagca catgcatcag aatgttccaa
cagaggaaca 1680aagaacaaat aaaagtaccg gtggtgaaca taagaaatct
gatagaaaag aagaacctca 1740atatgaacct gccaacactt ctgaagattt
agacatggat attgtgtctg ttccttcctc 1800agttccagaa gacatttttg
agaatcttga gactgctatg gaagttcaga gttcagttga 1860tcatcaaggg
gatggcagca gtggaactga acaagaagtg gagagttcat ctgtaaaatt
1920aaatatttct tcaaaagaca acagaggagg tattaaatca aaaactacag
ctaaagtaac 1980aaaagaatta tatgttaaac tcactcctgt ttccctttct
aattccccaa ttaaaggtgc 2040tgattgtcag gaagttccac aagataaaga
tggctataaa agttgtggtc tgaaccccaa 2100gttagagaaa tgtggacttg
gacaggaaaa cagtgataat gagcatttgg ttgaaaatga 2160agtttcatta
cttttagagg aatctgatct tcgaagatcc ccacgtgtaa agactacacc
2220cttgaggcga ccgacagaaa ctaaccctgt aacatctaat tcagatgaag
aatgtaatga 2280aacagttaag gagaaacaaa aactatcagt tccagtgaga
aaaaaggata agcgtaattc 2340ttctgacagt gctatagata atcctaagcc
taataaattg ccaaaatcta agcaatcaga 2400gactgtggat caaaattcag
attctgatga aatgctagca atcctcaaag aggtgagcag 2460gatgagtcac
agttcttctt cagatactga tattaatgaa attcatacaa accataagac
2520tttgtatgat ttaaagactc aggcggggaa agatgataaa ggaaaaagga
aacgaaaaag 2580ttctacatct ggctcagatt ttgatactaa aaagggcaaa
tcagctaaga gctctataat 2640ttctaaaaag aaacgacaaa cccagtctga
gtcttctaat tatgactcag aattagaaaa 2700agagataaag agcatgagta
aaattggtgc tgccagaacc accaaaaaaa gaattccaaa 2760tacaaaagat
tttgactctt ctgaagatga gaaacacagc aaaaaaggaa tggataatca
2820agggcacaaa aatttgaaga cctcacaaga aggatcatct gatgatgctg
aaagaaaaca 2880agagagagag actttctctt cagcagaagg cacagttgat
aaagacacga ccatcatgga 2940attaagagat cgacttccta agaagcagca
agcaagtgct tccactgatg gtgtcgataa 3000gctttctggg aaagagcaga
gttttacttc tttggaagtt agaaaagttg ctgaaactaa 3060agaaaagagc
aagcatctca aaaccaaaac atgtaaaaaa gtacaggatg gcttatctga
3120tattgcagag aaattcctaa agaaagacca gagcgatgaa acttctgaag
atgataaaaa 3180gcagagcaaa aagggaactg aagaaaaaaa gaaaccttca
gactttaaga aaaaagtaat 3240taaaatggaa caacagtatg aatcttcatc
tgatggcact gaaaagttac ctgagcgaga 3300agaaatttgt cattttccta
agggcataaa acaaattaag aatggaacaa ctgatggaga 3360aaagaaaagt
aaaaaaataa gagataaaac ttctaaaaag aaggatgaat tatctgatta
3420tgctgagaag tcaacaggga aaggagatag ttgtgactct tcagaggata
aaaagagtaa 3480gaatggagca tatggtagag agaagaaaag gtgcaagttg
cttggaaaga gttcaaggaa 3540gagacaagat tgttcatcat ctgatactga
gaaatattcc atgaaagaag atggttgtaa 3600ctcttctgat aagagactga
aaagaataga attgagggaa agaagaaatt taagttcaaa 3660gagaaatact
aaggaaatac aaagtggctc atcatcatct gatgctgagg aaagttctga
3720agataataaa aagaagaagc aaagaacttc atctaaaaag aaggcagtca
ttgtcaagga 3780gaaaaagaga aactccctaa gaacaagcac taaaaggaag
caagctgaca ttacatcctc 3840atcttcttct gatatagaag atgatgatca
gaattctata ggtgagggaa gcagcgatga 3900acagaaaatt aagcctgtga
ctgaaaattt agtgctgtct tcacatactg gattttgcca 3960atcttcagga
gatgaagcct tatctaaatc agtgcctgtc acagtggatg atgatgatga
4020cgacaatgat cctgagaata gaattgccaa gaagatgctt ttagaagaaa
ttaaagccaa 4080tctttcctct gatgaggatg gatcttcaga tgatgagcca
gaagaaggga aaaaaagaac 4140tggaaaacaa aatgaagaaa acccaggaga
tgaggaagca aaaaatcaag tcaattctga 4200atcagattca gattctgaag
aatctaagaa gccaagatac agacataggc ttttgcggca 4260caaattgact
gtgagtgacg gagaatctgg agaagaaaaa aagacaaagc ctaaagagca
4320taaagaagtc aaaggcagaa acagaagaaa ggtgagcagt gaagattcag
aagattctga 4380ttttcaggaa tcaggagtta gtgaagaagt tagtgaatcc
gaagatgaac agcggcccag 4440aacaaggtct gcaaagaaag cagagttgga
agaaaatcag cggagctata aacagaaaaa 4500gaaaaggcga cgtattaagg
ttcaagaaga ttcatccagt gaaaacaaga gtaattctga 4560ggaagaagag
gaggaaaaag aagaggagga ggaagaggag gaggaggagg aagaggagga
4620ggaagatgaa aatgatgatt ccaagtctcc tggaaaaggc agaaagaaaa
ttcggaagat 4680tcttaaagat gataaactga gaacagaaac acaaaatgct
cttaaggaag aggaagagag 4740acgaaaacgt attgctgaga gggagcgtga
gcgagaaaaa ttgagagagg tgatagaaat 4800tgaagatgct tcacccacca
agtgtccaat aacaaccaag ttggttttag atgaagatga 4860agaaaccaaa
gaacctttag tgcaggttca tagaaatatg gttatcaaat tgaaacccca
4920tcaagtagat ggtgttcagt ttatgtggga ttgctgctgt gagtctgtga
aaaaaacaaa 4980gaaatctcca ggttcaggat gcattcttgc ccactgtatg
ggccttggta agactttaca 5040ggtggtaagt tttcttcata cagttctttt
gtgtgacaaa ctggatttca gcacggcgtt 5100agtggtttgt cctcttaata
ctgctttgaa ttggatgaat gaatttgaga agtggcaaga 5160gggattaaaa
gatgatgaga agcttgaggt ttctgaatta gcaactgtga aacgtcctca
5220ggagagaagc tacatgctgc agaggtggca agaagatggt ggtgttatga
tcataggcta 5280tgagatgtat agaaatcttg ctcaaggaag gaatgtgaag
agtcggaaac ttaaagaaat 5340atttaacaaa gctttggttg atccaggccc
tgattttgtt gtttgtgatg aaggccatat 5400tctaaaaaat gaagcatctg
ctgtttctaa agctatgaat tctatacgat caaggaggag 5460gattatttta
acaggaacac cacttcaaaa taacctaatt gagtatcatt gtatggttaa
5520ttttatcaag gaaaatttac ttggatccat taaggagttc aggaatagat
ttataaatcc 5580aattcaaaat ggtcagtgtg cagattctac catggtagat
gtcagagtga tgaaaaaacg 5640tgctcacatt ctctatgaga tgttagctgg
atgtgttcag aggaaagatt atacagcatt 5700aacaaaattc ttgcctccaa
aacacgaata tgtgttagct gtgagaatga cttctattca 5760gtgcaagctc
tatcagtact acttagatca cttaacaggt gtgggcaata atagtgaagg
5820tggaagagga aaggcaggtg caaagctttt ccaagatttt cagatgttaa
gtagaatatg 5880gactcatcct tggtgtttgc agctagacta cattagcaaa
gaaaataagg gttattttga 5940tgaagacagt atggatgaat ttatagcctc
agattctgat gaaacctcca tgagtttaag 6000ctccgatgat tatacaaaaa
agaagaaaaa agggaaaaag gggaaaaaag atagtagctc 6060aagtggaagt
ggcagtgaca atgatgttga agtgattaag gtctggaatt caagatctcg
6120gggaggtggt gaaggaaatg tggatgaaac aggaaacaat ccttctgttt
ctttaaaact 6180ggaagaaagt aaagctactt cttcttctaa tccaagcagc
ccagctccag actggtacaa 6240agattttgtt acagatgctg atgctgaggt
tttagagcat tctgggaaaa tggtacttct 6300ctttgaaatt cttcgaatgg
cagaggaaat tggggataaa gtccttgttt tcagccagtc 6360cctcatatct
ctggacttga ttgaagattt tcttgaatta gctagtaggg agaagacaga
6420agataaagat aaacccctta tttataaagg tgaggggaag tggcttcgaa
acattgacta 6480ttaccgttta gatggttcca ctactgcaca gtcaaggaag
aagtgggctg aagaatttaa 6540tgatgaaact aatgtgagag gacgattatt
tatcatttct actaaagcag
gatctctagg 6600aattaatctg gtagctgcta atcgagtaat tatattcgac
gcttcttgga atccatctta 6660tgacatccag agtatattca gagtttatcg
ctttggacaa actaagcctg tttatgtata 6720taggttctta gctcagggaa
ccatggaaga taagatttat gatcggcaag taactaagca 6780gtcactgtct
tttcgagttg ttgatcagca gcaggtggag cgtcatttta ctatgaatga
6840gcttactgaa ctttatactt ttgagccaga cttattagat gaccctaatt
cagaaaagaa 6900gaagaagagg gatactccca tgctgccaaa ggataccata
cttgcagagc tccttcagat 6960acataaagaa cacattgtag gataccatga
acatgattct cttttggacc acaaagaaga 7020agaagagttg actgaagaag
aaagaaaagc agcttgggct gagtatgaag cagagaagaa 7080gggactgacc
atgcgtttca acataccaac tgggaccaat ttaccccctg tcagtttcaa
7140ctctcaaact ccttatattc ctttcaattt gggagccctg tcagcaatga
gtaatcaaca 7200gctggaggac ctcattaatc aaggaagaga aaaagttgta
gaagcaacaa acagtgtgac 7260agcagtgagg attcaacctc ttgaggatat
aatttcagct gtatggaagg agaacatgaa 7320tctctcagag gcccaagtac
aggcgttagc attaagtaga caagccagcc aggagcttga 7380tgttaaacga
agagaagcaa tctacaatga tgtattgaca aaacaacaga tgttaatcag
7440ctgtgttcag cgaatactta tgaacagaag gctccagcag cagtacaatc
agcagcaaca 7500gcaacaaatg acttatcaac aagcaacact gggtcacctc
atgatgccaa agcccccaaa 7560tttgatcatg aatccttcta actaccagca
gattgatatg agaggaatgt atcagccagt 7620ggctggtggt atgcagccac
caccattaca gcgtgcacca cccccaatga gaagcaaaaa 7680tccaggacct
tcccaaggga aatcaatgtg attttgcact aaaagcttaa tggattgtta
7740aaatcataga aagatctttt atttttttag gaatcaatga cttaacagaa
ctcaactgta 7800taaatagttt ggtcccctta aatgccaatc ttccatatta
gttttacttt tttttttttt 7860aaatagggca taccatttct tcctgacatt
tgtcagtgat gttgcctaga atcttcttac 7920acacgctgag tacagaagat
atttcaaatt gttttcagtg aaaacaagtc cttccataat 7980agtaacaact
ccacagattt cctctctaaa tttttatgcc tgcttttagc aaccataaaa
8040ttgtcataaa attaataaat ttaggaaaga ataaagattt atatattcat
tctttacata 8100taaaaacaca cagctgagtt cttagagttg attcctcaag
ttatgaaata cttttgtact 8160taatccattt cttgattaaa gtgattgaaa
tggttttaat gttcttttga ctgaagtctg 8220aaactgggct cctgctttat
tgtctctgtg actgaaagtt agaaactgag ggttatcttt 8280gacacagaat
tgtgtgcaat attcttaaat actactgctc taaaagttgg agaagtcttg
8340cagttatctt agcattgtat aaacagcctt aagtatagcc taagaagaga
attccttttt 8400cttctttagt ccttctgcca ttttttattt tcagttatat
gtgctgaaat aattactggt 8460aaaatttcag ggttgtggat tatcttccac
acatgaattt tctctctcct ggcacgaata 8520taaagcacat ctcttaactg
catggtgcca gtgctaatgc ttcatcctgt tgctggcagt 8580gggatgtgga
cttagaaaat caagttctag cattttagta ggttaacact gaagttgtgg
8640ttgttaggtt cacaccctgt tttataaaca acatcaaaat ggcagaacca
ttgctgactt 8700taggttcaca tgaggaatgt acttttaaca attcccagta
ctatcagtat tgtgaaataa 8760ttcctctgaa agataagaat cactggcttc
tatgcgcttc ttttctctca tcatcatgtt 8820cttttacccc agtttcctta
cattttttta aattgtttca gagtttgttt tttttttagt 8880ttagattgtg
aggcaattat taaatcaaaa ttaattcatc caatacccct ttactagaag
8940ttttactaga aaatgtatta cattttattt tttcttaatc cagttctgca
aaaatgacct 9000ataaatttat tcatgtacaa ttttggttac ttgaattgtt
aaagaaaaca ttgtttttga 9060ctatgggagt caactcaaca tggcagaacc
atttttgaga tgatgataca acaggtagtg 9120aaacagctta agaattccaa
aaaaaaaaaa aaaaaaaaaa aaaagaaaac tgggtttggg 9180ctttgcttta
ggtatcactg gattagaatg agtttaacat tagctaaaac tgctttgagt
9240tgtttggatg attaagagat tgccattttt atcttggaag aactagtggt
aaaacatcca 9300agagcactag gattgtgata cagaatttgt gaggtttggt
ggatccacgc ccctctcccc 9360cactttccca tgatgaaata tcactaataa
atcctgtata tttagatatt atgctagcca 9420tgtaatcaga tttatttaat
tgggtggggc aggtgtgtat ttactttaga aaaaatgaaa 9480aagacaagat
ttatgagaaa tatttgaagg cagtacactc tggccaactg ttaccagttg
9540gtatttctac aagttcagaa tattttaaac ctgatttact agacctggga
attttcaaca 9600tggtctaatt atttactcaa agacatagat gtgaaaattt
taggcaacct tctaaatctt 9660tttcaccatg gatgaaacta taacttaaag
aataatactt agaagggtta attggaaatc 9720agagtttgaa ataaaacttg
gaccactttg tatacactct tctcacttga cattttagct 9780atataatatg
tactttgagt ataacatcaa gctttaacaa atatttaaag acaaaaaaat
9840cacgtcagta aaatactaaa aggctcattt ttatatttgt tttagatgtt
ttaaatagtt 9900gcaatggatt aaaaatgatg atttaaaatg ttgcttgtaa
tacagttttg cctgctaaat 9960tctccacatt ttgtaacctg ttttatttct
ttgggtgtaa agcgtttttg cttagtattg 10020tgatattgta tatgttttgt
cccagttgta tagtaatgtt tcagtccatc atccagcttt 10080ggctgctgaa
atcatacagc tgtgaagact tgcctttgtt tctgttagac tgcttttcag
10140ttctgtattg agtatcttaa gtactgtaga aaagatgtca cttcttcctt
taaggctgtt 10200ttgtaatata tataaggact ggaattgtgt ttttaaagaa
aagcattcaa gtatgacaat 10260atactatctg tgttttcacc attcaaagtg
ctgtttagta gttgaaactt aaactattta 10320atgtcattta ataaagtgac
caaaatgtgt tgtgctcttt attgtatttt cacagctttg 10380aaaatctgtg
cacatactgt ttcatagaaa atgtatagct tttgttgtcc tatataatgg
10440tggttctttt gcacatttag ttatttaata ttgagaggtc acgaagtttg
gttattgaat 10500ctgttatata ctaaattctg taaagggaga tctctcatct
caaaaagaat ttacatacca 10560ggaagtccat gtgtgtttgt gttagttttg
gatgtctttg tgtaatccag ccccatttcc 10620tgtttcccaa cagctgtaac
actcatttta agtcaagcag ggctaccaac ccacacttga 10680tagaaaagct
gcttaccatt cagaagcttc cttattacct ggcctccaaa tgagctgaat
10740attttgtagc cttcccttag ctatgttcat tttccctcca ttatcataaa
atcagatcga 10800tatttatgtg ccccaaacaa aactttaaga gcagttacat
tctgtcccag tagcccttgt 10860ttcctttgag agtagcatgt tgtgaggcta
tagagactta ttctaccagt aaaacaggtc 10920aatcctttta catgtttatt
atactaaaaa ttatgttcag ggtatttact actttatttc 10980accagactca
gtctcaagtg acttggctat ctccaaatca gatctaccct tagagaataa
11040acatttttct accgttattt tttttcaagt ctataatctg agccagtccc
aaaggagtga 11100tcaagtttca gaaatgcttt catcttcaca acattttata
tatactatta tatggggtga 11160ataaagtttt aaatccgaaa tataaaaaaa
aaaaaaaaaa aa 1120272454PRTHomo sapiens 7Met Thr Ala Glu Pro Met
Ser Glu Ser Lys Leu Asn Thr Leu Val Gln1 5 10 15Lys Leu His Asp Phe
Leu Ala His Ser Ser Glu Glu Ser Glu Glu Thr 20 25 30Ser Ser Pro Pro
Arg Leu Ala Met Asn Gln Asn Thr Asp Lys Ile Ser 35 40 45Gly Ser Gly
Ser Asn Ser Asp Met Met Glu Asn Ser Lys Glu Glu Gly 50 55 60Thr Ser
Ser Ser Glu Lys Ser Lys Ser Ser Gly Ser Ser Arg Ser Lys65 70 75
80Arg Lys Pro Ser Ile Val Thr Lys Tyr Val Glu Ser Asp Asp Glu Lys
85 90 95Pro Leu Asp Asp Glu Thr Val Asn Glu Asp Ala Ser Asn Glu Asn
Ser 100 105 110Glu Asn Asp Ile Thr Met Gln Ser Leu Pro Lys Glu Asp
Gly Leu His 115 120 125Gly Ile Val Ser Cys Thr Ala Cys Gly Gln Gln
Val Asn His Phe Gln 130 135 140Lys Asp Ser Ile Tyr Arg His Pro Ser
Leu Gln Val Leu Ile Cys Lys145 150 155 160Asn Cys Phe Lys Tyr Tyr
Met Ser Asp Asp Ile Ser Arg Asp Ser Asp 165 170 175Gly Met Asp Glu
Gln Cys Arg Trp Cys Ala Glu Gly Gly Asn Leu Ile 180 185 190Cys Cys
Asp Phe Cys His Asn Ala Phe Cys Lys Lys Cys Ile Leu Arg 195 200
205Asn Leu Gly Arg Lys Glu Leu Ser Thr Ile Met Asp Glu Asn Asn Gln
210 215 220Trp Tyr Cys Tyr Ile Cys His Pro Glu Pro Leu Leu Asp Leu
Val Thr225 230 235 240Ala Cys Asn Ser Val Phe Glu Asn Leu Glu Gln
Leu Leu Gln Gln Asn 245 250 255Lys Lys Lys Ile Lys Val Asp Ser Glu
Lys Ser Asn Lys Val Tyr Glu 260 265 270His Thr Ser Arg Phe Ser Pro
Lys Lys Thr Ser Ser Asn Cys Asn Gly 275 280 285Glu Glu Lys Lys Leu
Asp Asp Ser Cys Ser Gly Ser Val Thr Tyr Ser 290 295 300Tyr Ser Ala
Leu Ile Val Pro Lys Glu Met Ile Lys Lys Ala Lys Lys305 310 315
320Leu Ile Glu Thr Thr Ala Asn Met Asn Ser Ser Tyr Val Lys Phe Leu
325 330 335Lys Gln Ala Thr Asp Asn Ser Glu Ile Ser Ser Ala Thr Lys
Leu Arg 340 345 350Gln Leu Lys Ala Phe Lys Ser Val Leu Ala Asp Ile
Lys Lys Ala His 355 360 365Leu Ala Leu Glu Glu Asp Leu Asn Ser Glu
Phe Arg Ala Met Asp Ala 370 375 380Val Asn Lys Glu Lys Asn Thr Lys
Glu His Lys Val Ile Asp Ala Lys385 390 395 400Phe Glu Thr Lys Ala
Arg Lys Gly Glu Lys Pro Cys Ala Leu Glu Lys 405 410 415Lys Asp Ile
Ser Lys Ser Glu Ala Lys Leu Ser Arg Lys Gln Val Asp 420 425 430Ser
Glu His Met His Gln Asn Val Pro Thr Glu Glu Gln Arg Thr Asn 435 440
445Lys Ser Thr Gly Gly Glu His Lys Lys Ser Asp Arg Lys Glu Glu Pro
450 455 460Gln Tyr Glu Pro Ala Asn Thr Ser Glu Asp Leu Asp Met Asp
Ile Val465 470 475 480Ser Val Pro Ser Ser Val Pro Glu Asp Ile Phe
Glu Asn Leu Glu Thr 485 490 495Ala Met Glu Val Gln Ser Ser Val Asp
His Gln Gly Asp Gly Ser Ser 500 505 510Gly Thr Glu Gln Glu Val Glu
Ser Ser Ser Val Lys Leu Asn Ile Ser 515 520 525Ser Lys Asp Asn Arg
Gly Gly Ile Lys Ser Lys Thr Thr Ala Lys Val 530 535 540Thr Lys Glu
Leu Tyr Val Lys Leu Thr Pro Val Ser Leu Ser Asn Ser545 550 555
560Pro Ile Lys Gly Ala Asp Cys Gln Glu Val Pro Gln Asp Lys Asp Gly
565 570 575Tyr Lys Ser Cys Gly Leu Asn Pro Lys Leu Glu Lys Cys Gly
Leu Gly 580 585 590Gln Glu Asn Ser Asp Asn Glu His Leu Val Glu Asn
Glu Val Ser Leu 595 600 605Leu Leu Glu Glu Ser Asp Leu Arg Arg Ser
Pro Arg Val Lys Thr Thr 610 615 620Pro Leu Arg Arg Pro Thr Glu Thr
Asn Pro Val Thr Ser Asn Ser Asp625 630 635 640Glu Glu Cys Asn Glu
Thr Val Lys Glu Lys Gln Lys Leu Ser Val Pro 645 650 655Val Arg Lys
Lys Asp Lys Arg Asn Ser Ser Asp Ser Ala Ile Asp Asn 660 665 670Pro
Lys Pro Asn Lys Leu Pro Lys Ser Lys Gln Ser Glu Thr Val Asp 675 680
685Gln Asn Ser Asp Ser Asp Glu Met Leu Ala Ile Leu Lys Glu Val Ser
690 695 700Arg Met Ser His Ser Ser Ser Ser Asp Thr Asp Ile Asn Glu
Ile His705 710 715 720Thr Asn His Lys Thr Leu Tyr Asp Leu Lys Thr
Gln Ala Gly Lys Asp 725 730 735Asp Lys Gly Lys Arg Lys Arg Lys Ser
Ser Thr Ser Gly Ser Asp Phe 740 745 750Asp Thr Lys Lys Gly Lys Ser
Ala Lys Ser Ser Ile Ile Ser Lys Lys 755 760 765Lys Arg Gln Thr Gln
Ser Glu Ser Ser Asn Tyr Asp Ser Glu Leu Glu 770 775 780Lys Glu Ile
Lys Ser Met Ser Lys Ile Gly Ala Ala Arg Thr Thr Lys785 790 795
800Lys Arg Ile Pro Asn Thr Lys Asp Phe Asp Ser Ser Glu Asp Glu Lys
805 810 815His Ser Lys Lys Gly Met Asp Asn Gln Gly His Lys Asn Leu
Lys Thr 820 825 830Ser Gln Glu Gly Ser Ser Asp Asp Ala Glu Arg Lys
Gln Glu Arg Glu 835 840 845Thr Phe Ser Ser Ala Glu Gly Thr Val Asp
Lys Asp Thr Thr Ile Met 850 855 860Glu Leu Arg Asp Arg Leu Pro Lys
Lys Gln Gln Ala Ser Ala Ser Thr865 870 875 880Asp Gly Val Asp Lys
Leu Ser Gly Lys Glu Gln Ser Phe Thr Ser Leu 885 890 895Glu Val Arg
Lys Val Ala Glu Thr Lys Glu Lys Ser Lys His Leu Lys 900 905 910Thr
Lys Thr Cys Lys Lys Val Gln Asp Gly Leu Ser Asp Ile Ala Glu 915 920
925Lys Phe Leu Lys Lys Asp Gln Ser Asp Glu Thr Ser Glu Asp Asp Lys
930 935 940Lys Gln Ser Lys Lys Gly Thr Glu Glu Lys Lys Lys Pro Ser
Asp Phe945 950 955 960Lys Lys Lys Val Ile Lys Met Glu Gln Gln Tyr
Glu Ser Ser Ser Asp 965 970 975Gly Thr Glu Lys Leu Pro Glu Arg Glu
Glu Ile Cys His Phe Pro Lys 980 985 990Gly Ile Lys Gln Ile Lys Asn
Gly Thr Thr Asp Gly Glu Lys Lys Ser 995 1000 1005Lys Lys Ile Arg
Asp Lys Thr Ser Lys Lys Lys Asp Glu Leu Ser 1010 1015 1020Asp Tyr
Ala Glu Lys Ser Thr Gly Lys Gly Asp Ser Cys Asp Ser 1025 1030
1035Ser Glu Asp Lys Lys Ser Lys Asn Gly Ala Tyr Gly Arg Glu Lys
1040 1045 1050Lys Arg Cys Lys Leu Leu Gly Lys Ser Ser Arg Lys Arg
Gln Asp 1055 1060 1065Cys Ser Ser Ser Asp Thr Glu Lys Tyr Ser Met
Lys Glu Asp Gly 1070 1075 1080Cys Asn Ser Ser Asp Lys Arg Leu Lys
Arg Ile Glu Leu Arg Glu 1085 1090 1095Arg Arg Asn Leu Ser Ser Lys
Arg Asn Thr Lys Glu Ile Gln Ser 1100 1105 1110Gly Ser Ser Ser Ser
Asp Ala Glu Glu Ser Ser Glu Asp Asn Lys 1115 1120 1125Lys Lys Lys
Gln Arg Thr Ser Ser Lys Lys Lys Ala Val Ile Val 1130 1135 1140Lys
Glu Lys Lys Arg Asn Ser Leu Arg Thr Ser Thr Lys Arg Lys 1145 1150
1155Gln Ala Asp Ile Thr Ser Ser Ser Ser Ser Asp Ile Glu Asp Asp
1160 1165 1170Asp Gln Asn Ser Ile Gly Glu Gly Ser Ser Asp Glu Gln
Lys Ile 1175 1180 1185Lys Pro Val Thr Glu Asn Leu Val Leu Ser Ser
His Thr Gly Phe 1190 1195 1200Cys Gln Ser Ser Gly Asp Glu Ala Leu
Ser Lys Ser Val Pro Val 1205 1210 1215Thr Val Asp Asp Asp Asp Asp
Asp Asn Asp Pro Glu Asn Arg Ile 1220 1225 1230Ala Lys Lys Met Leu
Leu Glu Glu Ile Lys Ala Asn Leu Ser Ser 1235 1240 1245Asp Glu Asp
Gly Ser Ser Asp Asp Glu Pro Glu Glu Gly Lys Lys 1250 1255 1260Arg
Thr Gly Lys Gln Asn Glu Glu Asn Pro Gly Asp Glu Glu Ala 1265 1270
1275Lys Asn Gln Val Asn Ser Glu Ser Asp Ser Asp Ser Glu Glu Ser
1280 1285 1290Lys Lys Pro Arg Tyr Arg His Arg Leu Leu Arg His Lys
Leu Thr 1295 1300 1305Val Ser Asp Gly Glu Ser Gly Glu Glu Lys Lys
Thr Lys Pro Lys 1310 1315 1320Glu His Lys Glu Val Lys Gly Arg Asn
Arg Arg Lys Val Ser Ser 1325 1330 1335Glu Asp Ser Glu Asp Ser Asp
Phe Gln Glu Ser Gly Val Ser Glu 1340 1345 1350Glu Val Ser Glu Ser
Glu Asp Glu Gln Arg Pro Arg Thr Arg Ser 1355 1360 1365Ala Lys Lys
Ala Glu Leu Glu Glu Asn Gln Arg Ser Tyr Lys Gln 1370 1375 1380Lys
Lys Lys Arg Arg Arg Ile Lys Val Gln Glu Asp Ser Ser Ser 1385 1390
1395Glu Asn Lys Ser Asn Ser Glu Glu Glu Glu Glu Glu Lys Glu Glu
1400 1405 1410Glu Glu Glu Glu Glu Glu Glu Glu Glu Glu Glu Glu Glu
Asp Glu 1415 1420 1425Asn Asp Asp Ser Lys Ser Pro Gly Lys Gly Arg
Lys Lys Ile Arg 1430 1435 1440Lys Ile Leu Lys Asp Asp Lys Leu Arg
Thr Glu Thr Gln Asn Ala 1445 1450 1455Leu Lys Glu Glu Glu Glu Arg
Arg Lys Arg Ile Ala Glu Arg Glu 1460 1465 1470Arg Glu Arg Glu Lys
Leu Arg Glu Val Ile Glu Ile Glu Asp Ala 1475 1480 1485Ser Pro Thr
Lys Cys Pro Ile Thr Thr Lys Leu Val Leu Asp Glu 1490 1495 1500Asp
Glu Glu Thr Lys Glu Pro Leu Val Gln Val His Arg Asn Met 1505 1510
1515Val Ile Lys Leu Lys Pro His Gln Val Asp Gly Val Gln Phe Met
1520 1525 1530Trp Asp Cys Cys Cys Glu Ser Val Lys Lys Thr Lys Lys
Ser Pro 1535 1540 1545Gly Ser Gly Cys Ile Leu Ala His Cys Met Gly
Leu Gly Lys Thr 1550 1555 1560Leu Gln Val Val Ser Phe Leu His Thr
Val Leu Leu Cys Asp Lys 1565 1570 1575Leu Asp Phe Ser Thr Ala Leu
Val Val Cys Pro Leu Asn Thr Ala 1580 1585 1590Leu Asn Trp Met Asn
Glu Phe Glu Lys Trp Gln Glu Gly Leu Lys 1595 1600 1605Asp Asp Glu
Lys Leu Glu Val Ser Glu Leu Ala Thr Val Lys Arg 1610 1615 1620Pro
Gln Glu Arg Ser Tyr Met Leu Gln Arg Trp Gln Glu Asp Gly 1625 1630
1635Gly Val Met Ile Ile Gly Tyr Glu Met Tyr Arg Asn Leu Ala Gln
1640 1645 1650Gly Arg Asn Val Lys Ser Arg Lys Leu Lys Glu Ile Phe
Asn Lys 1655 1660 1665Ala Leu Val Asp Pro Gly Pro Asp Phe Val Val
Cys Asp Glu Gly 1670 1675 1680His Ile Leu Lys Asn Glu Ala Ser Ala
Val Ser Lys Ala Met Asn 1685 1690 1695Ser Ile
Arg Ser Arg Arg Arg Ile Ile Leu Thr Gly Thr Pro Leu 1700 1705
1710Gln Asn Asn Leu Ile Glu Tyr His Cys Met Val Asn Phe Ile Lys
1715 1720 1725Glu Asn Leu Leu Gly Ser Ile Lys Glu Phe Arg Asn Arg
Phe Ile 1730 1735 1740Asn Pro Ile Gln Asn Gly Gln Cys Ala Asp Ser
Thr Met Val Asp 1745 1750 1755Val Arg Val Met Lys Lys Arg Ala His
Ile Leu Tyr Glu Met Leu 1760 1765 1770Ala Gly Cys Val Gln Arg Lys
Asp Tyr Thr Ala Leu Thr Lys Phe 1775 1780 1785Leu Pro Pro Lys His
Glu Tyr Val Leu Ala Val Arg Met Thr Ser 1790 1795 1800Ile Gln Cys
Lys Leu Tyr Gln Tyr Tyr Leu Asp His Leu Thr Gly 1805 1810 1815Val
Gly Asn Asn Ser Glu Gly Gly Arg Gly Lys Ala Gly Ala Lys 1820 1825
1830Leu Phe Gln Asp Phe Gln Met Leu Ser Arg Ile Trp Thr His Pro
1835 1840 1845Trp Cys Leu Gln Leu Asp Tyr Ile Ser Lys Glu Asn Lys
Gly Tyr 1850 1855 1860Phe Asp Glu Asp Ser Met Asp Glu Phe Ile Ala
Ser Asp Ser Asp 1865 1870 1875Glu Thr Ser Met Ser Leu Ser Ser Asp
Asp Tyr Thr Lys Lys Lys 1880 1885 1890Lys Lys Gly Lys Lys Gly Lys
Lys Asp Ser Ser Ser Ser Gly Ser 1895 1900 1905Gly Ser Asp Asn Asp
Val Glu Val Ile Lys Val Trp Asn Ser Arg 1910 1915 1920Ser Arg Gly
Gly Gly Glu Gly Asn Val Asp Glu Thr Gly Asn Asn 1925 1930 1935Pro
Ser Val Ser Leu Lys Leu Glu Glu Ser Lys Ala Thr Ser Ser 1940 1945
1950Ser Asn Pro Ser Ser Pro Ala Pro Asp Trp Tyr Lys Asp Phe Val
1955 1960 1965Thr Asp Ala Asp Ala Glu Val Leu Glu His Ser Gly Lys
Met Val 1970 1975 1980Leu Leu Phe Glu Ile Leu Arg Met Ala Glu Glu
Ile Gly Asp Lys 1985 1990 1995Val Leu Val Phe Ser Gln Ser Leu Ile
Ser Leu Asp Leu Ile Glu 2000 2005 2010Asp Phe Leu Glu Leu Ala Ser
Arg Glu Lys Thr Glu Asp Lys Asp 2015 2020 2025Lys Pro Leu Ile Tyr
Lys Gly Glu Gly Lys Trp Leu Arg Asn Ile 2030 2035 2040Asp Tyr Tyr
Arg Leu Asp Gly Ser Thr Thr Ala Gln Ser Arg Lys 2045 2050 2055Lys
Trp Ala Glu Glu Phe Asn Asp Glu Thr Asn Val Arg Gly Arg 2060 2065
2070Leu Phe Ile Ile Ser Thr Lys Ala Gly Ser Leu Gly Ile Asn Leu
2075 2080 2085Val Ala Ala Asn Arg Val Ile Ile Phe Asp Ala Ser Trp
Asn Pro 2090 2095 2100Ser Tyr Asp Ile Gln Ser Ile Phe Arg Val Tyr
Arg Phe Gly Gln 2105 2110 2115Thr Lys Pro Val Tyr Val Tyr Arg Phe
Leu Ala Gln Gly Thr Met 2120 2125 2130Glu Asp Lys Ile Tyr Asp Arg
Gln Val Thr Lys Gln Ser Leu Ser 2135 2140 2145Phe Arg Val Val Asp
Gln Gln Gln Val Glu Arg His Phe Thr Met 2150 2155 2160Asn Glu Leu
Thr Glu Leu Tyr Thr Phe Glu Pro Asp Leu Leu Asp 2165 2170 2175Asp
Pro Asn Ser Glu Lys Lys Lys Lys Arg Asp Thr Pro Met Leu 2180 2185
2190Pro Lys Asp Thr Ile Leu Ala Glu Leu Leu Gln Ile His Lys Glu
2195 2200 2205His Ile Val Gly Tyr His Glu His Asp Ser Leu Leu Asp
His Lys 2210 2215 2220Glu Glu Glu Glu Leu Thr Glu Glu Glu Arg Lys
Ala Ala Trp Ala 2225 2230 2235Glu Tyr Glu Ala Glu Lys Lys Gly Leu
Thr Met Arg Phe Asn Ile 2240 2245 2250Pro Thr Gly Thr Asn Leu Pro
Pro Val Ser Phe Asn Ser Gln Thr 2255 2260 2265Pro Tyr Ile Pro Phe
Asn Leu Gly Ala Leu Ser Ala Met Ser Asn 2270 2275 2280Gln Gln Leu
Glu Asp Leu Ile Asn Gln Gly Arg Glu Lys Val Val 2285 2290 2295Glu
Ala Thr Asn Ser Val Thr Ala Val Arg Ile Gln Pro Leu Glu 2300 2305
2310Asp Ile Ile Ser Ala Val Trp Lys Glu Asn Met Asn Leu Ser Glu
2315 2320 2325Ala Gln Val Gln Ala Leu Ala Leu Ser Arg Gln Ala Ser
Gln Glu 2330 2335 2340Leu Asp Val Lys Arg Arg Glu Ala Ile Tyr Asn
Asp Val Leu Thr 2345 2350 2355Lys Gln Gln Met Leu Ile Ser Cys Val
Gln Arg Ile Leu Met Asn 2360 2365 2370Arg Arg Leu Gln Gln Gln Tyr
Asn Gln Gln Gln Gln Gln Gln Met 2375 2380 2385Thr Tyr Gln Gln Ala
Thr Leu Gly His Leu Met Met Pro Lys Pro 2390 2395 2400Pro Asn Leu
Ile Met Asn Pro Ser Asn Tyr Gln Gln Ile Asp Met 2405 2410 2415Arg
Gly Met Tyr Gln Pro Val Ala Gly Gly Met Gln Pro Pro Pro 2420 2425
2430Leu Gln Arg Ala Pro Pro Pro Met Arg Ser Lys Asn Pro Gly Pro
2435 2440 2445Ser Gln Gly Lys Ser Met 2450811088DNAHomo sapiens
8aattctcctg cctgagcctc ggcccaacaa aatggcggcg gcagcggtgt cgctttgttt
60ccgcggctcc tgcggcggtg gcagtggtag cggcctttga gctgtgggga ggttccagca
120gcagctacag tgacgactaa gactccagtg catttctatc gtaaccgggc
gcgggggagc 180gcagatcggc gcccagcaat cacagaagcc gacaaggcgt
tcaagcgaaa acatgaccgc 240tgagcccatg agtgaaagca agttgaatac
attggtgcag aagcttcatg acttccttgc 300acactcatca gaagaatctg
aagaaacaag ttctcctcca cgacttgcaa tgaatcaaaa 360cacagataaa
atcagtggtt ctggaagtaa ctctgatatg atggaaaaca gcaaggaaga
420gggaactagc tcttcagaaa aatccaagtc ttcaggatcg tcacgatcaa
agaggaaacc 480ttcaattgta acaaagtatg tagaatcaga tgatgaaaaa
cctttggatg atgaaactgt 540aaatgaagat gcgtctaatg aaaattcaga
aaatgatatt actatgcaga gcttgccaaa 600agaagatggg cttcatggga
ttgtgagctg cactgcttgt ggacaacagg tcaatcattt 660tcaaaaagat
tccatttata gacacccttc attgcaagtt cttatttgta agaattgctt
720taagtattac atgagtgatg atattagccg tgactcagat ggaatggatg
aacaatgtag 780gtggtgtgcg gaaggtggaa acttgatttg ttgtgacttt
tgccataatg ctttctgcaa 840gaaatgcatt ctacgcaacc ttggtcgaaa
ggagttgtcc acaataatgg atgaaaacaa 900ccaatggtat tgctacattt
gtcacccaga gcctttgttg gacttggtca ctgcatgtaa 960cagcgtattt
gagaatttag aacagttgtt gcagcaaaat aagaagaaga taaaagttga
1020cagtgaaaag agtaataaag tatatgaaca tacatccaga ttttctccaa
agaagactag 1080ttcaaattgt aatggagaag aaaagaaatt agatgattcc
tgttctggct ctgtaaccta 1140ctcttattcc gcactaattg tgcccaaaga
gatgattaag aaggcaaaaa aactgattga 1200gaccacagcc aacatgaact
ccagttatgt taaattttta aagcaggcaa cagataattc 1260agaaatcagt
tctgctacaa aattacgtca gcttaaggct tttaagtctg tgttggctga
1320tattaagaag gctcatcttg cattggaaga agacttaaat tccgagtttc
gagcgatgga 1380tgctgtaaac aaagagaaaa ataccaaaga gcataaagtc
atagatgcta agtttgaaac 1440aaaagcacga aaaggagaaa aaccttgtgc
tttggaaaag aaggatattt caaagtcaga 1500agctaaactt tcaagaaaac
aggtagatag tgagcacatg catcagaatg ttccaacaga 1560ggaacaaaga
acaaataaaa gtaccggtgg tgaacataag aaatctgata gaaaagaaga
1620acctcaatat gaacctgcca acacttctga agatttagac atggatattg
tgtctgttcc 1680ttcctcagtt ccagaagaca tttttgagaa tcttgagact
gctatggaag ttcagagttc 1740agttgatcat caaggggatg gcagcagtgg
aactgaacaa gaagtggaga gttcatctgt 1800aaaattaaat atttcttcaa
aagacaacag aggaggtatt aaatcaaaaa ctacagctaa 1860agtaacaaaa
gaattatatg ttaaactcac tcctgtttcc ctttctaatt ccccaattaa
1920aggtgctgat tgtcaggaag ttccacaaga taaagatggc tataaaagtt
gtggtctgaa 1980ccccaagtta gagaaatgtg gacttggaca ggaaaacagt
gataatgagc atttggttga 2040aaatgaagtt tcattacttt tagaggaatc
tgatcttcga agatccccac gtgtaaagac 2100tacacccttg aggcgaccga
cagaaactaa ccctgtaaca tctaattcag atgaagaatg 2160taatgaaaca
gttaaggaga aacaaaaact atcagttcca gtgagaaaaa aggataagcg
2220taattcttct gacagtgcta tagataatcc taagcctaat aaattgccaa
aatctaagca 2280atcagagact gtggatcaaa attcagattc tgatgaaatg
ctagcaatcc tcaaagaggt 2340gagcaggatg agtcacagtt cttcttcaga
tactgatatt aatgaaattc atacaaacca 2400taagactttg tatgatttaa
agactcaggc ggggaaagat gataaaggaa aaaggaaacg 2460aaaaagttct
acatctggct cagattttga tactaaaaag ggcaaatcag ctaagagctc
2520tataatttct aaaaagaaac gacaaaccca gtctgagtct tctaattatg
actcagaatt 2580agaaaaagag ataaagagca tgagtaaaat tggtgctgcc
agaaccacca aaaaaagaat 2640tccaaataca aaagattttg actcttctga
agatgagaaa cacagcaaaa aaggaatgga 2700taatcaaggg cacaaaaatt
tgaagacctc acaagaagga tcatctgatg atgctgaaag 2760aaaacaagag
agagagactt tctcttcagc agaaggcaca gttgataaag acacgaccat
2820catggaatta agagatcgac ttcctaagaa gcagcaagca agtgcttcca
ctgatggtgt 2880cgataagctt tctgggaaag agcagagttt tacttctttg
gaagttagaa aagttgctga 2940aactaaagaa aagagcaagc atctcaaaac
caaaacatgt aaaaaagtac aggatggctt 3000atctgatatt gcagagaaat
tcctaaagaa agaccagagc gatgaaactt ctgaagatga 3060taaaaagcag
agcaaaaagg gaactgaaga aaaaaagaaa ccttcagact ttaagaaaaa
3120agtaattaaa atggaacaac agtatgaatc ttcatctgat ggcactgaaa
agttacctga 3180gcgagaagaa atttgtcatt ttcctaaggg cataaaacaa
attaagaatg gaacaactga 3240tggagaaaag aaaagtaaaa aaataagaga
taaaacttct aaaaagaagg atgaattatc 3300tgattatgct gagaagtcaa
cagggaaagg agatagttgt gactcttcag aggataaaaa 3360gagtaagaat
ggagcatatg gtagagagaa gaaaaggtgc aagttgcttg gaaagagttc
3420aaggaagaga caagattgtt catcatctga tactgagaaa tattccatga
aagaagatgg 3480ttgtaactct tctgataaga gactgaaaag aatagaattg
agggaaagaa gaaatttaag 3540ttcaaagaga aatactaagg aaatacaaag
tggctcatca tcatctgatg ctgaggaaag 3600ttctgaagat aataaaaaga
agaagcaaag aacttcatct aaaaagaagg cagtcattgt 3660caaggagaaa
aagagaaact ccctaagaac aagcactaaa aggaagcaag ctgacattac
3720atcctcatct tcttctgata tagaagatga tgatcagaat tctataggtg
agggaagcag 3780cgatgaacag aaaattaagc ctgtgactga aaatttagtg
ctgtcttcac atactggatt 3840ttgccaatct tcaggagatg aagccttatc
taaatcagtg cctgtcacag tggatgatga 3900tgatgacgac aatgatcctg
agaatagaat tgccaagaag atgcttttag aagaaattaa 3960agccaatctt
tcctctgatg aggatggatc ttcagatgat gagccagaag aagggaaaaa
4020aagaactgga aaacaaaatg aagaaaaccc aggagatgag gaagcaaaaa
atcaagtcaa 4080ttctgaatca gattcagatt ctgaagaatc taagaagcca
agatacagac ataggctttt 4140gcggcacaaa ttgactgtga gtgacggaga
atctggagaa gaaaaaaaga caaagcctaa 4200agagcataaa gaagtcaaag
gcagaaacag aagaaaggtg agcagtgaag attcagaaga 4260ttctgatttt
caggaatcag gagttagtga agaagttagt gaatccgaag atgaacagcg
4320gcccagaaca aggtctgcaa agaaagcaga gttggaagaa aatcagcgga
gctataaaca 4380gaaaaagaaa aggcgacgta ttaaggttca agaagattca
tccagtgaaa acaagagtaa 4440ttctgaggaa gaagaggagg aaaaagaaga
ggaggaggaa gaggaggagg aggaggaaga 4500ggaggaggaa gatgaaaatg
atgattccaa gtctcctgga aaaggcagaa agaaaattcg 4560gaagattctt
aaagatgata aactgagaac agaaacacaa aatgctctta aggaagagga
4620agagagacga aaacgtattg ctgagaggga gcgtgagcga gaaaaattga
gagaggtgat 4680agaaattgaa gatgcttcac ccaccaagtg tccaataaca
accaagttgg ttttagatga 4740agatgaagaa accaaagaac ctttagtgca
ggttcataga aatatggtta tcaaattgaa 4800accccatcaa gtagatggtg
ttcagtttat gtgggattgc tgctgtgagt ctgtgaaaaa 4860aacaaagaaa
tctccaggtt caggatgcat tcttgcccac tgtatgggcc ttggtaagac
4920tttacaggtg gtaagttttc ttcatacagt tcttttgtgt gacaaactgg
atttcagcac 4980ggcgttagtg gtttgtcctc ttaatactgc tttgaattgg
atgaatgaat ttgagaagtg 5040gcaagaggga ttaaaagatg atgagaagct
tgaggtttct gaattagcaa ctgtgaaacg 5100tcctcaggag agaagctaca
tgctgcagag gtggcaagaa gatggtggtg ttatgatcat 5160aggctatgag
atgtatagaa atcttgctca aggaaggaat gtgaagagtc ggaaacttaa
5220agaaatattt aacaaagctt tggttgatcc aggccctgat tttgttgttt
gtgatgaagg 5280ccatattcta aaaaatgaag catctgctgt ttctaaagct
atgaattcta tacgatcaag 5340gaggaggatt attttaacag gaacaccact
tcaaaataac ctaattgagt atcattgtat 5400ggttaatttt atcaaggaaa
atttacttgg atccattaag gagttcagga atagatttat 5460aaatccaatt
caaaatggtc agtgtgcaga ttctaccatg gtagatgtca gagtgatgaa
5520aaaacgtgct cacattctct atgagatgtt agctggatgt gttcagagga
aagattatac 5580agcattaaca aaattcttgc ctccaaaaca cgaatatgtg
ttagctgtga gaatgacttc 5640tattcagtgc aagctctatc agtactactt
agatcactta acaggtgtgg gcaataatag 5700tgaaggtgga agaggaaagg
caggtgcaaa gcttttccaa gattttcaga tgttaagtag 5760aatatggact
catccttggt gtttgcagct agactacatt agcaaagaaa ataagggtta
5820ttttgatgaa gacagtatgg atgaatttat agcctcagat tctgatgaaa
cctccatgag 5880tttaagctcc gatgattata caaaaaagaa gaaaaaaggg
aaaaagggga aaaaagatag 5940tagctcaagt ggaagtggca gtgacaatga
tgttgaagtg attaaggtct ggaattcaag 6000atctcgggga ggtggtgaag
gaaatgtgga tgaaacagga aacaatcctt ctgtttcttt 6060aaaactggaa
gaaagtaaag ctacttcttc ttctaatcca agcagcccag ctccagactg
6120gtacaaagat tttgttacag atgctgatgc tgaggtttta gagcattctg
ggaaaatggt 6180acttctcttt gaaattcttc gaatggcaga ggaaattggg
gataaagtcc ttgttttcag 6240ccagtccctc atatctctgg acttgattga
agattttctt gaattagcta gtagggagaa 6300gacagaagat aaagataaac
cccttattta taaaggtgag gggaagtggc ttcgaaacat 6360tgactattac
cgtttagatg gttccactac tgcacagtca aggaagaagt gggctgaaga
6420atttaatgat gaaactaatg tgagaggacg attatttatc atttctacta
aagcaggatc 6480tctaggaatt aatctggtag ctgctaatcg agtaattata
ttcgacgctt cttggaatcc 6540atcttatgac atccagagta tattcagagt
ttatcgcttt ggacaaacta agcctgttta 6600tgtatatagg ttcttagctc
agggaaccat ggaagataag atttatgatc ggcaagtaac 6660taagcagtca
ctgtcttttc gagttgttga tcagcagcag gtggagcgtc attttactat
6720gaatgagctt actgaacttt atacttttga gccagactta ttagatgacc
ctaattcaga 6780aaagaagaag aagagggata ctcccatgct gccaaaggat
accatacttg cagagctcct 6840tcagatacat aaagaacaca ttgtaggata
ccatgaacat gattctcttt tggaccacaa 6900agaagaagaa gagttgactg
aagaagaaag aaaagcagct tgggctgagt atgaagcaga 6960gaagaaggga
ctgaccatgc gtttcaacat accaactggg accaatttac cccctgtcag
7020tttcaactct caaactcctt atattccttt caatttggga gccctgtcag
caatgagtaa 7080tcaacagctg gaggacctca ttaatcaagg aagagaaaaa
gttgtagaag caacaaacag 7140tgtgacagca gtgaggattc aacctcttga
ggatataatt tcagctgtat ggaaggagaa 7200catgaatctc tcagaggccc
aagtacaggc gttagcatta agtagacaag ccagccagga 7260gcttgatgtt
aaacgaagag aagcaatcta caatgatgta ttgacaaaac aacagatgtt
7320aatcagctgt gttcagcgaa tacttatgaa cagaaggctc cagcagcagt
acaatcagca 7380gcaacagcaa caaatgactt atcaacaagc aacactgggt
cacctcatga tgccaaagcc 7440cccaaatttg atcatgaatc cttctaacta
ccagcagatt gatatgagag gaatgtatca 7500gccagtggct ggtggtatgc
agccaccacc attacagcgt gcaccacccc caatgagaag 7560caaaaatcca
ggaccttccc aagggaaatc aatgtgattt tgcactaaaa gcttaatgga
7620ttgttaaaat catagaaaga tcttttattt ttttaggaat caatgactta
acagaactca 7680actgtataaa tagtttggtc cccttaaatg ccaatcttcc
atattagttt tacttttttt 7740ttttttaaat agggcatacc atttcttcct
gacatttgtc agtgatgttg cctagaatct 7800tcttacacac gctgagtaca
gaagatattt caaattgttt tcagtgaaaa caagtccttc 7860cataatagta
acaactccac agatttcctc tctaaatttt tatgcctgct tttagcaacc
7920ataaaattgt cataaaatta ataaatttag gaaagaataa agatttatat
attcattctt 7980tacatataaa aacacacagc tgagttctta gagttgattc
ctcaagttat gaaatacttt 8040tgtacttaat ccatttcttg attaaagtga
ttgaaatggt tttaatgttc ttttgactga 8100agtctgaaac tgggctcctg
ctttattgtc tctgtgactg aaagttagaa actgagggtt 8160atctttgaca
cagaattgtg tgcaatattc ttaaatacta ctgctctaaa agttggagaa
8220gtcttgcagt tatcttagca ttgtataaac agccttaagt atagcctaag
aagagaattc 8280ctttttcttc tttagtcctt ctgccatttt ttattttcag
ttatatgtgc tgaaataatt 8340actggtaaaa tttcagggtt gtggattatc
ttccacacat gaattttctc tctcctggca 8400cgaatataaa gcacatctct
taactgcatg gtgccagtgc taatgcttca tcctgttgct 8460ggcagtggga
tgtggactta gaaaatcaag ttctagcatt ttagtaggtt aacactgaag
8520ttgtggttgt taggttcaca ccctgtttta taaacaacat caaaatggca
gaaccattgc 8580tgactttagg ttcacatgag gaatgtactt ttaacaattc
ccagtactat cagtattgtg 8640aaataattcc tctgaaagat aagaatcact
ggcttctatg cgcttctttt ctctcatcat 8700catgttcttt taccccagtt
tccttacatt tttttaaatt gtttcagagt ttgttttttt 8760tttagtttag
attgtgaggc aattattaaa tcaaaattaa ttcatccaat acccctttac
8820tagaagtttt actagaaaat gtattacatt ttattttttc ttaatccagt
tctgcaaaaa 8880tgacctataa atttattcat gtacaatttt ggttacttga
attgttaaag aaaacattgt 8940ttttgactat gggagtcaac tcaacatggc
agaaccattt ttgagatgat gatacaacag 9000gtagtgaaac agcttaagaa
ttccaaaaaa aaaaaaaaaa aaaaaaaaaa gaaaactggg 9060tttgggcttt
gctttaggta tcactggatt agaatgagtt taacattagc taaaactgct
9120ttgagttgtt tggatgatta agagattgcc atttttatct tggaagaact
agtggtaaaa 9180catccaagag cactaggatt gtgatacaga atttgtgagg
tttggtggat ccacgcccct 9240ctcccccact ttcccatgat gaaatatcac
taataaatcc tgtatattta gatattatgc 9300tagccatgta atcagattta
tttaattggg tggggcaggt gtgtatttac tttagaaaaa 9360atgaaaaaga
caagatttat gagaaatatt tgaaggcagt acactctggc caactgttac
9420cagttggtat ttctacaagt tcagaatatt ttaaacctga tttactagac
ctgggaattt 9480tcaacatggt ctaattattt actcaaagac atagatgtga
aaattttagg caaccttcta 9540aatctttttc accatggatg aaactataac
ttaaagaata atacttagaa gggttaattg 9600gaaatcagag tttgaaataa
aacttggacc actttgtata cactcttctc acttgacatt 9660ttagctatat
aatatgtact ttgagtataa catcaagctt taacaaatat ttaaagacaa
9720aaaaatcacg tcagtaaaat actaaaaggc tcatttttat atttgtttta
gatgttttaa 9780atagttgcaa tggattaaaa atgatgattt aaaatgttgc
ttgtaataca gttttgcctg 9840ctaaattctc cacattttgt aacctgtttt
atttctttgg gtgtaaagcg tttttgctta 9900gtattgtgat attgtatatg
ttttgtccca gttgtatagt aatgtttcag tccatcatcc 9960agctttggct
gctgaaatca tacagctgtg aagacttgcc tttgtttctg ttagactgct
10020tttcagttct gtattgagta tcttaagtac tgtagaaaag atgtcacttc
ttcctttaag 10080gctgttttgt aatatatata aggactggaa ttgtgttttt
aaagaaaagc attcaagtat 10140gacaatatac tatctgtgtt ttcaccattc
aaagtgctgt ttagtagttg aaacttaaac 10200tatttaatgt catttaataa
agtgaccaaa atgtgttgtg ctctttattg tattttcaca 10260gctttgaaaa
tctgtgcaca tactgtttca tagaaaatgt atagcttttg ttgtcctata
10320taatggtggt tcttttgcac atttagttat ttaatattga gaggtcacga
agtttggtta
10380ttgaatctgt tatatactaa attctgtaaa gggagatctc tcatctcaaa
aagaatttac 10440ataccaggaa gtccatgtgt gtttgtgtta gttttggatg
tctttgtgta atccagcccc 10500atttcctgtt tcccaacagc tgtaacactc
attttaagtc aagcagggct accaacccac 10560acttgataga aaagctgctt
accattcaga agcttcctta ttacctggcc tccaaatgag 10620ctgaatattt
tgtagccttc ccttagctat gttcattttc cctccattat cataaaatca
10680gatcgatatt tatgtgcccc aaacaaaact ttaagagcag ttacattctg
tcccagtagc 10740ccttgtttcc tttgagagta gcatgttgtg aggctataga
gacttattct accagtaaaa 10800caggtcaatc cttttacatg tttattatac
taaaaattat gttcagggta tttactactt 10860tatttcacca gactcagtct
caagtgactt ggctatctcc aaatcagatc tacccttaga 10920gaataaacat
ttttctaccg ttattttttt tcaagtctat aatctgagcc agtcccaaag
10980gagtgatcaa gtttcagaaa tgctttcatc ttcacaacat tttatatata
ctattatatg 11040gggtgaataa agttttaaat ccgaaatata aaaaaaaaaa
aaaaaaaa 1108892285PRTHomo sapiens 9Met Ala Ala Gln Val Ala Pro Ala
Ala Ala Ser Ser Leu Gly Asn Pro1 5 10 15Pro Pro Pro Pro Pro Ser Glu
Leu Lys Lys Ala Glu Gln Gln Gln Arg 20 25 30Glu Glu Ala Gly Gly Glu
Ala Ala Ala Ala Ala Ala Ala Glu Arg Gly 35 40 45Glu Met Lys Ala Ala
Ala Gly Gln Glu Ser Glu Gly Pro Ala Val Gly 50 55 60Pro Pro Gln Pro
Leu Gly Lys Glu Leu Gln Asp Gly Ala Glu Ser Asn65 70 75 80Gly Gly
Gly Gly Gly Gly Gly Ala Gly Ser Gly Gly Gly Pro Gly Ala 85 90 95Glu
Pro Asp Leu Lys Asn Ser Asn Gly Asn Ala Gly Pro Arg Pro Ala 100 105
110Leu Asn Asn Asn Leu Thr Glu Pro Pro Gly Gly Gly Gly Gly Gly Ser
115 120 125Ser Asp Gly Val Gly Ala Pro Pro His Ser Ala Ala Ala Ala
Leu Pro 130 135 140Pro Pro Ala Tyr Gly Phe Gly Gln Pro Tyr Gly Arg
Ser Pro Ser Ala145 150 155 160Val Ala Ala Ala Ala Ala Ala Val Phe
His Gln Gln His Gly Gly Gln 165 170 175Gln Ser Pro Gly Leu Ala Ala
Leu Gln Ser Gly Gly Gly Gly Gly Leu 180 185 190Glu Pro Tyr Ala Gly
Pro Gln Gln Asn Ser His Asp His Gly Phe Pro 195 200 205Asn His Gln
Tyr Asn Ser Tyr Tyr Pro Asn Arg Ser Ala Tyr Pro Pro 210 215 220Pro
Ala Pro Ala Tyr Ala Leu Ser Ser Pro Arg Gly Gly Thr Pro Gly225 230
235 240Ser Gly Ala Ala Ala Ala Ala Gly Ser Lys Pro Pro Pro Ser Ser
Ser 245 250 255Ala Ser Ala Ser Ser Ser Ser Ser Ser Phe Ala Gln Gln
Arg Phe Gly 260 265 270Ala Met Gly Gly Gly Gly Pro Ser Ala Ala Gly
Gly Gly Thr Pro Gln 275 280 285Pro Thr Ala Thr Pro Thr Leu Asn Gln
Leu Leu Thr Ser Pro Ser Ser 290 295 300Ala Arg Gly Tyr Gln Gly Tyr
Pro Gly Gly Asp Tyr Ser Gly Gly Pro305 310 315 320Gln Asp Gly Gly
Ala Gly Lys Gly Pro Ala Asp Met Ala Ser Gln Cys 325 330 335Trp Gly
Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ser Gly Gly 340 345
350Ala Gln Gln Arg Ser His His Ala Pro Met Ser Pro Gly Ser Ser Gly
355 360 365Gly Gly Gly Gln Pro Leu Ala Arg Thr Pro Gln Pro Ser Ser
Pro Met 370 375 380Asp Gln Met Gly Lys Met Arg Pro Gln Pro Tyr Gly
Gly Thr Asn Pro385 390 395 400Tyr Ser Gln Gln Gln Gly Pro Pro Ser
Gly Pro Gln Gln Gly His Gly 405 410 415Tyr Pro Gly Gln Pro Tyr Gly
Ser Gln Thr Pro Gln Arg Tyr Pro Met 420 425 430Thr Met Gln Gly Arg
Ala Gln Ser Ala Met Gly Gly Leu Ser Tyr Thr 435 440 445Gln Gln Ile
Pro Pro Tyr Gly Gln Gln Gly Pro Ser Gly Tyr Gly Gln 450 455 460Gln
Gly Gln Thr Pro Tyr Tyr Asn Gln Gln Ser Pro His Pro Gln Gln465 470
475 480Gln Gln Pro Pro Tyr Ser Gln Gln Pro Pro Ser Gln Thr Pro His
Ala 485 490 495Gln Pro Ser Tyr Gln Gln Gln Pro Gln Ser Gln Pro Pro
Gln Leu Gln 500 505 510Ser Ser Gln Pro Pro Tyr Ser Gln Gln Pro Ser
Gln Pro Pro His Gln 515 520 525Gln Ser Pro Ala Pro Tyr Pro Ser Gln
Gln Ser Thr Thr Gln Gln His 530 535 540Pro Gln Ser Gln Pro Pro Tyr
Ser Gln Pro Gln Ala Gln Ser Pro Tyr545 550 555 560Gln Gln Gln Gln
Pro Gln Gln Pro Ala Pro Ser Thr Leu Ser Gln Gln 565 570 575Ala Ala
Tyr Pro Gln Pro Gln Ser Gln Gln Ser Gln Gln Thr Ala Tyr 580 585
590Ser Gln Gln Arg Phe Pro Pro Pro Gln Glu Leu Ser Gln Asp Ser Phe
595 600 605Gly Ser Gln Ala Ser Ser Ala Pro Ser Met Thr Ser Ser Lys
Gly Gly 610 615 620Gln Glu Asp Met Asn Leu Ser Leu Gln Ser Arg Pro
Ser Ser Leu Pro625 630 635 640Asp Leu Ser Gly Ser Ile Asp Asp Leu
Pro Met Gly Thr Glu Gly Ala 645 650 655Leu Ser Pro Gly Val Ser Thr
Ser Gly Ile Ser Ser Ser Gln Gly Glu 660 665 670Gln Ser Asn Pro Ala
Gln Ser Pro Phe Ser Pro His Thr Ser Pro His 675 680 685Leu Pro Gly
Ile Arg Gly Pro Ser Pro Ser Pro Val Gly Ser Pro Ala 690 695 700Ser
Val Ala Gln Ser Arg Ser Gly Pro Leu Ser Pro Ala Ala Val Pro705 710
715 720Gly Asn Gln Met Pro Pro Arg Pro Pro Ser Gly Gln Ser Asp Ser
Ile 725 730 735Met His Pro Ser Met Asn Gln Ser Ser Ile Ala Gln Asp
Arg Gly Tyr 740 745 750Met Gln Arg Asn Pro Gln Met Pro Gln Tyr Ser
Ser Pro Gln Pro Gly 755 760 765Ser Ala Leu Ser Pro Arg Gln Pro Ser
Gly Gly Gln Ile His Thr Gly 770 775 780Met Gly Ser Tyr Gln Gln Asn
Ser Met Gly Ser Tyr Gly Pro Gln Gly785 790 795 800Gly Gln Tyr Gly
Pro Gln Gly Gly Tyr Pro Arg Gln Pro Asn Tyr Asn 805 810 815Ala Leu
Pro Asn Ala Asn Tyr Pro Ser Ala Gly Met Ala Gly Gly Ile 820 825
830Asn Pro Met Gly Ala Gly Gly Gln Met His Gly Gln Pro Gly Ile Pro
835 840 845Pro Tyr Gly Thr Leu Pro Pro Gly Arg Met Ser His Ala Ser
Met Gly 850 855 860Asn Arg Pro Tyr Gly Pro Asn Met Ala Asn Met Pro
Pro Gln Val Gly865 870 875 880Ser Gly Met Cys Pro Pro Pro Gly Gly
Met Asn Arg Lys Thr Gln Glu 885 890 895Thr Ala Val Ala Met His Val
Ala Ala Asn Ser Ile Gln Asn Arg Pro 900 905 910Pro Gly Tyr Pro Asn
Met Asn Gln Gly Gly Met Met Gly Thr Gly Pro 915 920 925Pro Tyr Gly
Gln Gly Ile Asn Ser Met Ala Gly Met Ile Asn Pro Gln 930 935 940Gly
Pro Pro Tyr Ser Met Gly Gly Thr Met Ala Asn Asn Ser Ala Gly945 950
955 960Met Ala Ala Ser Pro Glu Met Met Gly Leu Gly Asp Val Lys Leu
Thr 965 970 975Pro Ala Thr Lys Met Asn Asn Lys Ala Asp Gly Thr Pro
Lys Thr Glu 980 985 990Ser Lys Ser Lys Lys Ser Ser Ser Ser Thr Thr
Thr Asn Glu Lys Ile 995 1000 1005Thr Lys Leu Tyr Glu Leu Gly Gly
Glu Pro Glu Arg Lys Met Trp 1010 1015 1020Val Asp Arg Tyr Leu Ala
Phe Thr Glu Glu Lys Ala Met Gly Met 1025 1030 1035Thr Asn Leu Pro
Ala Val Gly Arg Lys Pro Leu Asp Leu Tyr Arg 1040 1045 1050Leu Tyr
Val Ser Val Lys Glu Ile Gly Gly Leu Thr Gln Val Asn 1055 1060
1065Lys Asn Lys Lys Trp Arg Glu Leu Ala Thr Asn Leu Asn Val Gly
1070 1075 1080Thr Ser Ser Ser Ala Ala Ser Ser Leu Lys Lys Gln Tyr
Ile Gln 1085 1090 1095Cys Leu Tyr Ala Phe Glu Cys Lys Ile Glu Arg
Gly Glu Asp Pro 1100 1105 1110Pro Pro Asp Ile Phe Ala Ala Ala Asp
Ser Lys Lys Ser Gln Pro 1115 1120 1125Lys Ile Gln Pro Pro Ser Pro
Ala Gly Ser Gly Ser Met Gln Gly 1130 1135 1140Pro Gln Thr Pro Gln
Ser Thr Ser Ser Ser Met Ala Glu Gly Gly 1145 1150 1155Asp Leu Lys
Pro Pro Thr Pro Ala Ser Thr Pro His Ser Gln Ile 1160 1165 1170Pro
Pro Leu Pro Gly Met Ser Arg Ser Asn Ser Val Gly Ile Gln 1175 1180
1185Asp Ala Phe Asn Asp Gly Ser Asp Ser Thr Phe Gln Lys Arg Asn
1190 1195 1200Ser Met Thr Pro Asn Pro Gly Tyr Gln Pro Ser Met Asn
Thr Ser 1205 1210 1215Asp Met Met Gly Arg Met Ser Tyr Glu Pro Asn
Lys Asp Pro Tyr 1220 1225 1230Gly Ser Met Arg Lys Ala Pro Gly Ser
Asp Pro Phe Met Ser Ser 1235 1240 1245Gly Gln Gly Pro Asn Gly Gly
Met Gly Asp Pro Tyr Ser Arg Ala 1250 1255 1260Ala Gly Pro Gly Leu
Gly Asn Val Ala Met Gly Pro Arg Gln His 1265 1270 1275Tyr Pro Tyr
Gly Gly Pro Tyr Asp Arg Val Arg Thr Glu Pro Gly 1280 1285 1290Ile
Gly Pro Glu Gly Asn Met Ser Thr Gly Ala Pro Gln Pro Asn 1295 1300
1305Leu Met Pro Ser Asn Pro Asp Ser Gly Met Tyr Ser Pro Ser Arg
1310 1315 1320Tyr Pro Pro Gln Gln Gln Gln Gln Gln Gln Gln Arg His
Asp Ser 1325 1330 1335Tyr Gly Asn Gln Phe Ser Thr Gln Gly Thr Pro
Ser Gly Ser Pro 1340 1345 1350Phe Pro Ser Gln Gln Thr Thr Met Tyr
Gln Gln Gln Gln Gln Asn 1355 1360 1365Tyr Lys Arg Pro Met Asp Gly
Thr Tyr Gly Pro Pro Ala Lys Arg 1370 1375 1380His Glu Gly Glu Met
Tyr Ser Val Pro Tyr Ser Thr Gly Gln Gly 1385 1390 1395Gln Pro Gln
Gln Gln Gln Leu Pro Pro Ala Gln Pro Gln Pro Ala 1400 1405 1410Ser
Gln Gln Gln Ala Ala Gln Pro Ser Pro Gln Gln Asp Val Tyr 1415 1420
1425Asn Gln Tyr Gly Asn Ala Tyr Pro Ala Thr Ala Thr Ala Ala Thr
1430 1435 1440Glu Arg Arg Pro Ala Gly Gly Pro Gln Asn Gln Phe Pro
Phe Gln 1445 1450 1455Phe Gly Arg Asp Arg Val Ser Ala Pro Pro Gly
Thr Asn Ala Gln 1460 1465 1470Gln Asn Met Pro Pro Gln Met Met Gly
Gly Pro Ile Gln Ala Ser 1475 1480 1485Ala Glu Val Ala Gln Gln Gly
Thr Met Trp Gln Gly Arg Asn Asp 1490 1495 1500Met Thr Tyr Asn Tyr
Ala Asn Arg Gln Ser Thr Gly Ser Ala Pro 1505 1510 1515Gln Gly Pro
Ala Tyr His Gly Val Asn Arg Thr Asp Glu Met Leu 1520 1525 1530His
Thr Asp Gln Arg Ala Asn His Glu Gly Ser Trp Pro Ser His 1535 1540
1545Gly Thr Arg Gln Pro Pro Tyr Gly Pro Ser Ala Pro Val Pro Pro
1550 1555 1560Met Thr Arg Pro Pro Pro Ser Asn Tyr Gln Pro Pro Pro
Ser Met 1565 1570 1575Gln Asn His Ile Pro Gln Val Ser Ser Pro Ala
Pro Leu Pro Arg 1580 1585 1590Pro Met Glu Asn Arg Thr Ser Pro Ser
Lys Ser Pro Phe Leu His 1595 1600 1605Ser Gly Met Lys Met Gln Lys
Ala Gly Pro Pro Val Pro Ala Ser 1610 1615 1620His Ile Ala Pro Ala
Pro Val Gln Pro Pro Met Ile Arg Arg Asp 1625 1630 1635Ile Thr Phe
Pro Pro Gly Ser Val Glu Ala Thr Gln Pro Val Leu 1640 1645 1650Lys
Gln Arg Arg Arg Leu Thr Met Lys Asp Ile Gly Thr Pro Glu 1655 1660
1665Ala Trp Arg Val Met Met Ser Leu Lys Ser Gly Leu Leu Ala Glu
1670 1675 1680Ser Thr Trp Ala Leu Asp Thr Ile Asn Ile Leu Leu Tyr
Asp Asp 1685 1690 1695Asn Ser Ile Met Thr Phe Asn Leu Ser Gln Leu
Pro Gly Leu Leu 1700 1705 1710Glu Leu Leu Val Glu Tyr Phe Arg Arg
Cys Leu Ile Glu Ile Phe 1715 1720 1725Gly Ile Leu Lys Glu Tyr Glu
Val Gly Asp Pro Gly Gln Arg Thr 1730 1735 1740Leu Leu Asp Pro Gly
Arg Phe Ser Lys Val Ser Ser Pro Ala Pro 1745 1750 1755Met Glu Gly
Gly Glu Glu Glu Glu Glu Leu Leu Gly Pro Lys Leu 1760 1765 1770Glu
Glu Glu Glu Glu Glu Glu Val Val Glu Asn Asp Glu Glu Ile 1775 1780
1785Ala Phe Ser Gly Lys Asp Lys Pro Ala Ser Glu Asn Ser Glu Glu
1790 1795 1800Lys Leu Ile Ser Lys Phe Asp Lys Leu Pro Val Lys Ile
Val Gln 1805 1810 1815Lys Asn Asp Pro Phe Val Val Asp Cys Ser Asp
Lys Leu Gly Arg 1820 1825 1830Val Gln Glu Phe Asp Ser Gly Leu Leu
His Trp Arg Ile Gly Gly 1835 1840 1845Gly Asp Thr Thr Glu His Ile
Gln Thr His Phe Glu Ser Lys Thr 1850 1855 1860Glu Leu Leu Pro Ser
Arg Pro His Ala Pro Cys Pro Pro Ala Pro 1865 1870 1875Arg Lys His
Val Thr Thr Ala Glu Gly Thr Pro Gly Thr Thr Asp 1880 1885 1890Gln
Glu Gly Pro Pro Pro Asp Gly Pro Pro Glu Lys Arg Ile Thr 1895 1900
1905Ala Thr Met Asp Asp Met Leu Ser Thr Arg Ser Ser Thr Leu Thr
1910 1915 1920Glu Asp Gly Ala Lys Ser Ser Glu Ala Ile Lys Glu Ser
Ser Lys 1925 1930 1935Phe Pro Phe Gly Ile Ser Pro Ala Gln Ser His
Arg Asn Ile Lys 1940 1945 1950Ile Leu Glu Asp Glu Pro His Ser Lys
Asp Glu Thr Pro Leu Cys 1955 1960 1965Thr Leu Leu Asp Trp Gln Asp
Ser Leu Ala Lys Arg Cys Val Cys 1970 1975 1980Val Ser Asn Thr Ile
Arg Ser Leu Ser Phe Val Pro Gly Asn Asp 1985 1990 1995Phe Glu Met
Ser Lys His Pro Gly Leu Leu Leu Ile Leu Gly Lys 2000 2005 2010Leu
Ile Leu Leu His His Lys His Pro Glu Arg Lys Gln Ala Pro 2015 2020
2025Leu Thr Tyr Glu Lys Glu Glu Glu Gln Asp Gln Gly Val Ser Cys
2030 2035 2040Asn Lys Val Glu Trp Trp Trp Asp Cys Leu Glu Met Leu
Arg Glu 2045 2050 2055Asn Thr Leu Val Thr Leu Ala Asn Ile Ser Gly
Gln Leu Asp Leu 2060 2065 2070Ser Pro Tyr Pro Glu Ser Ile Cys Leu
Pro Val Leu Asp Gly Leu 2075 2080 2085Leu His Trp Ala Val Cys Pro
Ser Ala Glu Ala Gln Asp Pro Phe 2090 2095 2100Ser Thr Leu Gly Pro
Asn Ala Val Leu Ser Pro Gln Arg Leu Val 2105 2110 2115Leu Glu Thr
Leu Ser Lys Leu Ser Ile Gln Asp Asn Asn Val Asp 2120 2125 2130Leu
Ile Leu Ala Thr Pro Pro Phe Ser Arg Leu Glu Lys Leu Tyr 2135 2140
2145Ser Thr Met Val Arg Phe Leu Ser Asp Arg Lys Asn Pro Val Cys
2150 2155 2160Arg Glu Met Ala Val Val Leu Leu Ala Asn Leu Ala Gln
Gly Asp 2165 2170 2175Ser Leu Ala Ala Arg Ala Ile Ala Val Gln Lys
Gly Ser Ile Gly 2180 2185 2190Asn Leu Leu Gly Phe Leu Glu Asp Ser
Leu Ala Ala Thr Gln Phe 2195 2200 2205Gln Gln Ser Gln Ala Ser Leu
Leu His Met Gln Asn Pro Pro Phe 2210 2215 2220Glu Pro Thr Ser Val
Asp Met Met Arg Arg Ala Ala Arg Ala Leu 2225 2230 2235Leu Ala Leu
Ala Lys Val Asp Glu Asn His Ser Glu Phe Thr Leu 2240 2245 2250Tyr
Glu Ser Arg Leu Leu Asp Ile Ser Val Ser Pro Leu Met Asn 2255 2260
2265Ser Leu Val Ser Gln Val Ile Cys Asp Val Leu Phe Leu Ile Gly
2270 2275 2280Gln Ser 2285108585DNAHomo sapiens 10cagaaagcgg
agagtcacag cggggccagg ccctggggag cggagcctcc accgcccccc 60tcattcccag
gcaagggctt ggggggaatg agccgggaga gccgggtccc gagcctacag
120agccgggagc agctgagccg ccggcgcctc ggccgccgcc gccgcctcct
cctcctccgc 180cgccgccagc ccggagcctg agccggcggg gcggggggga
gaggagcgag cgcagcgcag 240cagcggagcc ccgcgaggcc cgcccgggcg
ggtggggagg gcagcccggg ggactgggcc 300ccggggcggg gtgggagggg
gggagaagac gaagacaggg ccgggtctct ccgcggacga 360gacagcgggg
atcatggccg cgcaggtcgc ccccgccgcc gccagcagcc tgggcaaccc
420gccgccgccg ccgccctcgg agctgaagaa agccgagcag cagcagcggg
aggaggcggg 480gggcgaggcg gcggcggcgg cagcggccga gcgcggggaa
atgaaggcag ccgccgggca 540ggaaagcgag ggccccgccg tggggccgcc
gcagccgctg ggaaaggagc tgcaggacgg 600ggccgagagc aatgggggtg
gcggcggcgg cggagccggc agcggcggcg ggcccggcgc 660ggagccggac
ctgaagaact cgaacgggaa cgcgggccct aggcccgccc tgaacaataa
720cctcacggag ccgcccggcg gcggcggtgg cggcagcagc gatggggtgg
gggcgcctcc 780tcactcagcc gcggccgcct tgccgccccc agcctacggc
ttcgggcaac cctacggccg 840gagcccgtct gccgtcgccg ccgccgcggc
cgccgtcttc caccaacaac atggcggaca 900acaaagccct ggcctggcag
cgctgcagag cggcggcggc gggggcctgg agccctacgc 960ggggccccag
cagaactctc acgaccacgg cttccccaac caccagtaca actcctacta
1020ccccaaccgc agcgcctacc ccccgcccgc cccggcctac gcgctgagct
ccccgagagg 1080tggcactccg ggctccggcg cggcggcggc tgccggctcc
aagccgcctc cctcctccag 1140cgcctccgcc tcctcgtcgt cttcgtcctt
cgctcagcag cgcttcgggg ccatgggggg 1200aggcggcccc tccgcggccg
gcgggggaac tccccagccc accgccaccc ccaccctcaa 1260ccaactgctc
acgtcgccca gctcggcccg gggctaccag ggctaccccg ggggcgacta
1320cagtggcggg ccccaggacg ggggcgccgg caagggcccg gcggacatgg
cctcgcagtg 1380ttggggggct gcggcggcgg cagctgcggc ggcggccgcc
tcgggagggg cccaacaaag 1440gagccaccac gcgcccatga gccccgggag
cagcggcggc ggggggcagc cgctcgcccg 1500gacccctcag ccatccagtc
caatggatca gatgggcaag atgagacctc agccatatgg 1560cgggactaac
ccatactcgc agcaacaggg acctccgtca ggaccgcagc aaggacatgg
1620gtacccaggg cagccatacg ggtcccagac cccgcagcgg tacccgatga
ccatgcaggg 1680ccgggcgcag agtgccatgg gcggcctctc ttatacacag
cagattcctc cttatggaca 1740acaaggcccc agcgggtatg gtcaacaggg
ccagactcca tattacaacc agcaaagtcc 1800tcaccctcag cagcagcagc
caccctactc ccagcaacca ccgtcccaga cccctcatgc 1860ccaaccttcg
tatcagcagc agccacagtc tcaaccacca cagctccagt cctctcagcc
1920tccatactcc cagcagccat cccagcctcc acatcagcag tccccggctc
catacccctc 1980ccagcagtcg acgacacagc agcaccccca gagccagccc
ccctactcac agccacaggc 2040tcagtctcct taccagcagc agcaacctca
gcagccagca ccctcgacgc tctcccagca 2100ggctgcgtat cctcagcccc
agtctcagca gtcccagcaa actgcctatt cccagcagcg 2160cttccctcca
ccgcaggagc tatctcaaga ttcatttggg tctcaggcat cctcagcccc
2220ctcaatgacc tccagtaagg gagggcaaga agatatgaac ctgagccttc
agtcaagacc 2280ctccagcttg cctgatctat ctggttcaat agatgacctc
cccatgggga cagaaggagc 2340tctgagtcct ggagtgagca catcagggat
ttccagcagc caaggagagc agagtaatcc 2400agctcagtct cctttctctc
ctcatacctc ccctcacctg cctggcatcc gaggcccttc 2460cccgtcccct
gttggctctc ccgccagtgt tgctcagtct cgctcaggac cactctcgcc
2520tgctgcagtg ccaggcaacc agatgccacc tcggccaccc agtggccagt
cggacagcat 2580catgcatcct tccatgaacc aatcaagcat tgcccaagat
cgaggttata tgcagaggaa 2640cccccagatg ccccagtaca gttcccccca
gcccggctca gccttatctc cgcgtcagcc 2700ttccggagga cagatacaca
caggcatggg ctcctaccag cagaactcca tggggagcta 2760tggtccccag
gggggtcagt atggcccaca aggtggctac cccaggcagc caaactataa
2820tgccttgccc aatgccaact accccagtgc aggcatggct ggaggcataa
accccatggg 2880tgccggaggt caaatgcatg gacagcctgg catcccacct
tatggcacac tccctccagg 2940gaggatgagt cacgcctcca tgggcaaccg
gccttatggc cctaacatgg ccaatatgcc 3000acctcaggtt gggtcaggga
tgtgtccccc accagggggc atgaaccgga aaacccaaga 3060aactgctgtc
gccatgcatg ttgctgccaa ctctatccaa aacaggccgc caggctaccc
3120caatatgaat caagggggca tgatgggaac tggacctcct tatggacaag
ggattaatag 3180tatggctggc atgatcaacc ctcagggacc cccatattcc
atgggtggaa ccatggccaa 3240caattctgca gggatggcag ccagcccaga
gatgatgggc cttggggatg taaagttaac 3300tccagccacc aaaatgaaca
acaaggcaga tgggacaccc aagacagaat ccaaatccaa 3360gaaatccagt
tcttctacta caaccaatga gaagatcacc aagttgtatg agctgggtgg
3420tgagcctgag aggaagatgt gggtggaccg ttatctggcc ttcactgagg
agaaggccat 3480gggcatgaca aatctgcctg ctgtgggtag gaaacctctg
gacctctatc gcctctatgt 3540gtctgtgaag gagattggtg gattgactca
ggtcaacaag aacaaaaaat ggcgggaact 3600tgcaaccaac ctcaatgtgg
gcacatcaag cagtgctgcc agctccttga aaaagcagta 3660tatccagtgt
ctctatgcct ttgaatgcaa gattgaacgg ggagaagacc ctcccccaga
3720catctttgca gctgctgatt ccaagaagtc ccagcccaag atccagcctc
cctctcctgc 3780gggatcagga tctatgcagg ggccccagac tccccagtca
accagcagtt ccatggcaga 3840aggaggagac ttaaagccac caactccagc
atccacacca cacagtcaga tccccccatt 3900gccaggcatg agcaggagca
attcagttgg gatccaggat gcctttaatg atggaagtga 3960ctccacattc
cagaagcgga attccatgac tccaaaccct gggtatcagc ccagtatgaa
4020tacctctgac atgatggggc gcatgtccta tgagccaaat aaggatcctt
atggcagcat 4080gaggaaagct ccagggagtg atcccttcat gtcctcaggg
cagggcccca acggcgggat 4140gggtgacccc tacagtcgtg ctgccggccc
tgggctagga aatgtggcga tgggaccacg 4200acagcactat ccctatggag
gtccttatga cagagtgagg acggagcctg gaatagggcc 4260tgagggaaac
atgagcactg gggccccaca gccgaatctc atgccttcca acccagactc
4320ggggatgtat tctcctagcc gctacccccc gcagcagcag cagcagcagc
agcaacgaca 4380tgattcctat ggcaatcagt tctccaccca aggcacccct
tctggcagcc ccttccccag 4440ccagcagact acaatgtatc aacagcaaca
gcagaattac aagcggccaa tggatggcac 4500atatggccct cctgccaagc
ggcacgaagg ggagatgtac agcgtgccat acagcactgg 4560gcaggggcag
cctcagcagc agcagttgcc cccagcccag ccccagcctg ccagccagca
4620acaagctgcc cagccttccc ctcagcaaga tgtatacaac cagtatggca
atgcctatcc 4680tgccactgcc acagctgcta ctgagcgccg accagcaggc
ggcccccaga accaatttcc 4740attccagttt ggccgagacc gtgtctctgc
accccctggc accaatgccc agcaaaacat 4800gccaccacaa atgatgggcg
gccccataca ggcatcagct gaggttgctc agcaaggcac 4860catgtggcag
gggcgtaatg acatgaccta taattatgcc aacaggcaga gcacgggctc
4920tgccccccag ggccccgcct atcatggcgt gaaccgaaca gatgaaatgc
tgcacacaga 4980tcagagggcc aaccacgaag gctcgtggcc ttcccatggc
acacgccagc ccccatatgg 5040tccctctgcc cctgtgcccc ccatgacaag
gccccctcca tctaactacc agcccccacc 5100aagcatgcag aatcacattc
ctcaggtatc cagccctgct cccctgcccc ggccaatgga 5160gaaccgcacc
tctcctagca agtctccatt cctgcactct gggatgaaaa tgcagaaggc
5220aggtccccca gtacctgcct cgcacatagc acctgcccct gtgcagcccc
ccatgattcg 5280gcgggatatc accttcccac ctggctctgt tgaagccaca
cagcctgtgt tgaagcagag 5340gaggcggctc acaatgaaag acattggaac
cccggaggca tggcgggtaa tgatgtccct 5400caagtctggt ctcctggcag
agagcacatg ggcattagat accatcaaca tcctgctgta 5460tgatgacaac
agcatcatga ccttcaacct cagtcagctc ccagggttgc tagagctcct
5520tgtagaatat ttccgacgat gcctgattga gatctttggc attttaaagg
agtatgaggt 5580gggtgaccca ggacagagaa cgctactgga tcctgggagg
ttcagcaagg tgtctagtcc 5640agctcccatg gagggtgggg aagaagaaga
agaacttcta ggtcctaaac tagaagagga 5700agaagaagag gaagtagttg
aaaatgatga ggagatagcc ttttcaggca aggacaagcc 5760agcttcagag
aatagtgagg agaagctgat cagtaagttt gacaagcttc cagtaaagat
5820cgtacagaag aatgatccat ttgtggtgga ctgctcagat aagcttgggc
gtgtgcagga 5880gtttgacagt ggcctgctgc actggcggat tggtgggggg
gacaccactg agcatatcca 5940gacccacttc gagagcaaga cagagctgct
gccttcccgg cctcacgcac cctgcccacc 6000agcccctcgg aagcatgtga
caacagcaga gggtacacca gggacaacag accaggaggg 6060gcccccacct
gatggacctc cagaaaaacg gatcacagcc actatggatg acatgttgtc
6120tactcggtct agcaccttga ccgaggatgg agctaagagt tcagaggcca
tcaaggagag 6180cagcaagttt ccatttggca ttagcccagc acagagccac
cggaacatca agatcctaga 6240ggacgaaccc cacagtaagg atgagacccc
actgtgtacc cttctggact ggcaggattc 6300tcttgccaag cgctgcgtct
gtgtgtccaa taccattcga agcctgtcat ttgtgccagg 6360caatgacttt
gagatgtcca aacacccagg gctgctgctc atcctgggca agctgatcct
6420gctgcaccac aagcacccag aacggaagca ggcaccacta acttatgaaa
aggaggagga 6480acaggaccaa ggggtgagct gcaacaaagt ggagtggtgg
tgggactgct tggagatgct 6540ccgggaaaac accttggtta cactcgccaa
catctcgggg cagttggacc tatctccata 6600ccccgagagc atttgcctgc
ctgtcctgga cggactccta cactgggcag tttgcccttc 6660agctgaagcc
caggacccct tttccaccct gggccccaat gccgtccttt ccccgcagag
6720actggtcttg gaaaccctca gcaaactcag catccaggac aacaatgtgg
acctgattct 6780ggccacaccc cccttcagcc gcctggagaa gttgtatagc
actatggtgc gcttcctcag 6840tgaccgaaag aacccggtgt gccgggagat
ggctgtggta ctgctggcca acctggctca 6900gggggacagc ctggcagctc
gtgccattgc agtgcagaag ggcagtatcg gcaacctcct 6960gggcttccta
gaggacagcc ttgccgccac acagttccag cagagccagg ccagcctcct
7020ccacatgcag aacccaccct ttgagccaac tagtgtggac atgatgcggc
gggctgcccg 7080cgcgctgctt gccttggcca aggtggacga gaaccactca
gagtttactc tgtacgaatc 7140acggctgttg gacatctcgg tatcaccgtt
gatgaactca ttggtttcac aagtcatttg 7200tgatgtactg tttttgattg
gccagtcatg acagccgtgg gacacctccc ccccccgtgt 7260gtgtgtgcgt
gtgtggagaa cttagaaact gactgttgcc ctttatttat gcaaaaccac
7320ctcagaatcc agtttaccct gtgctgtcca gcttctccct tgggaaaaag
tctctcctgt 7380ttctctctcc tccttccacc tcccctccct ccatcacctc
acgcctttct gttccttgtc 7440ctcaccttac tcccctcagg accctacccc
accctctttg aaaagacaaa gctctgccta 7500catagaagac tttttttatt
ttaaccaaag ttactgttgt ttacagtgag tttggggaaa 7560aaaaataaaa
taaaaatggc tttcccagtc cttgcatcaa cgggatgcca catttcataa
7620ctgtttttaa tggtaaaaaa aaaaaaaaaa aatacaaaaa aaaattctga
aggacaaaaa 7680aggtgactgc tgaactgtgt gtggtttatt gttgtacatt
cacaatcttg caggagccaa 7740gaagttcgca gttgtgaaca gaccctgttc
actggagagg cctgtgcagt agagtgtaga 7800ccctttcatg tactgtactg
tacacctgat actgtaaaca tactgtaata ataatgtctc 7860acatggaaac
agaaaacgct gggtcagcag caagctgtag tttttaaaaa tgtttttagt
7920taaacgttga ggagaaaaaa aaaaaaggct tttcccccaa agtatcatgt
gtgaacctac 7980aacaccctga cctctttctc tcctccttga ttgtatgaat
aaccctgaga tcacctctta 8040gaactggttt taacctttag ctgcagcggc
tacgctgcca cgtgtgtata tatatgacgt 8100tgtacattgc acataccctt
ggatccccac agtttggtcc tcctcccagc taccccttta 8160tagtatgacg
agttaacaag ttggtgacct gcacaaagcg agacacagct atttaatctc
8220ttgccagata tcgcccctct tggtgcgatg ctgtacaggt ctctgtaaaa
agtccttgct 8280gtctcagcag ccaatcaact tatagtttat ttttttctgg
gtttttgttt tgttttgttt 8340tctttctaat cgaggtgtga aaaagttcta
ggttcagttg aagttctgat gaagaaacac 8400aattgagatt ttttcagtga
taaaatctgc atatttgtat ttcaacaatg tagctaaaac 8460ttgatgtaaa
ttcctccttt ttttcctttt ttggcttaat gaatatcatt tattcagtat
8520gaaatcttta tactatatgt tccacgtgtt aagaataaat gtacattaaa
tcttggtaag 8580acttt 8585112068PRTHomo sapiens 11Met Ala Ala Gln
Val Ala Pro Ala Ala Ala Ser Ser Leu Gly Asn Pro1 5 10 15Pro Pro Pro
Pro Pro Ser Glu Leu Lys Lys Ala Glu Gln Gln Gln Arg 20 25 30Glu Glu
Ala Gly Gly Glu Ala Ala Ala Ala Ala Ala Ala Glu Arg Gly 35 40 45Glu
Met Lys Ala Ala Ala Gly Gln Glu Ser Glu Gly Pro Ala Val Gly 50 55
60Pro Pro Gln Pro Leu Gly Lys Glu Leu Gln Asp Gly Ala Glu Ser Asn65
70 75 80Gly Gly Gly Gly Gly Gly Gly Ala Gly Ser Gly Gly Gly Pro Gly
Ala 85 90 95Glu Pro Asp Leu Lys Asn Ser Asn Gly Asn Ala Gly Pro Arg
Pro Ala 100 105 110Leu Asn Asn Asn Leu Thr Glu Pro Pro Gly Gly Gly
Gly Gly Gly Ser 115 120 125Ser Asp Gly Val Gly Ala Pro Pro His Ser
Ala Ala Ala Ala Leu Pro 130 135 140Pro Pro Ala Tyr Gly Phe Gly Gln
Pro Tyr Gly Arg Ser Pro Ser Ala145 150 155 160Val Ala Ala Ala Ala
Ala Ala Val Phe His Gln Gln His Gly Gly Gln 165 170 175Gln Ser Pro
Gly Leu Ala Ala Leu Gln Ser Gly Gly Gly Gly Gly Leu 180 185 190Glu
Pro Tyr Ala Gly Pro Gln Gln Asn Ser His Asp His Gly Phe Pro 195 200
205Asn His Gln Tyr Asn Ser Tyr Tyr Pro Asn Arg Ser Ala Tyr Pro Pro
210 215 220Pro Ala Pro Ala Tyr Ala Leu Ser Ser Pro Arg Gly Gly Thr
Pro Gly225 230 235 240Ser Gly Ala Ala Ala Ala Ala Gly Ser Lys Pro
Pro Pro Ser Ser Ser 245 250 255Ala Ser Ala Ser Ser Ser Ser Ser Ser
Phe Ala Gln Gln Arg Phe Gly 260 265 270Ala Met Gly Gly Gly Gly Pro
Ser Ala Ala Gly Gly Gly Thr Pro Gln 275 280 285Pro Thr Ala Thr Pro
Thr Leu Asn Gln Leu Leu Thr Ser Pro Ser Ser 290 295 300Ala Arg Gly
Tyr Gln Gly Tyr Pro Gly Gly Asp Tyr Ser Gly Gly Pro305 310 315
320Gln Asp Gly Gly Ala Gly Lys Gly Pro Ala Asp Met Ala Ser Gln Cys
325 330 335Trp Gly Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ser
Gly Gly 340 345 350Ala Gln Gln Arg Ser His His Ala Pro Met Ser Pro
Gly Ser Ser Gly 355 360 365Gly Gly Gly Gln Pro Leu Ala Arg Thr Pro
Gln Pro Ser Ser Pro Met 370 375 380Asp Gln Met Gly Lys Met Arg Pro
Gln Pro Tyr Gly Gly Thr Asn Pro385 390 395 400Tyr Ser Gln Gln Gln
Gly Pro Pro Ser Gly Pro Gln Gln Gly His Gly 405 410 415Tyr Pro Gly
Gln Pro Tyr Gly Ser Gln Thr Pro Gln Arg Tyr Pro Met 420 425 430Thr
Met Gln Gly Arg Ala Gln Ser Ala Met Gly Gly Leu Ser Tyr Thr 435 440
445Gln Gln Ile Pro Pro Tyr Gly Gln Gln Gly Pro Ser Gly Tyr Gly Gln
450 455 460Gln Gly Gln Thr Pro Tyr Tyr Asn Gln Gln Ser Pro His Pro
Gln Gln465 470 475 480Gln Gln Pro Pro Tyr Ser Gln Gln Pro Pro Ser
Gln Thr Pro His Ala 485 490 495Gln Pro Ser Tyr Gln Gln Gln Pro Gln
Ser Gln Pro Pro Gln Leu Gln 500 505 510Ser Ser Gln Pro Pro Tyr Ser
Gln Gln Pro Ser Gln Pro Pro His Gln 515 520 525Gln Ser Pro Ala Pro
Tyr Pro Ser Gln Gln Ser Thr Thr Gln Gln His 530 535 540Pro Gln Ser
Gln Pro Pro Tyr Ser Gln Pro Gln Ala Gln Ser Pro Tyr545 550 555
560Gln Gln Gln Gln Pro Gln Gln Pro Ala Pro Ser Thr Leu Ser Gln Gln
565 570 575Ala Ala Tyr Pro Gln Pro Gln Ser Gln Gln Ser Gln Gln Thr
Ala Tyr 580 585 590Ser Gln Gln Arg Phe Pro Pro Pro Gln Glu Leu Ser
Gln Asp Ser Phe 595 600 605Gly Ser Gln Ala Ser Ser Ala Pro Ser Met
Thr Ser Ser Lys Gly Gly 610 615 620Gln Glu Asp Met Asn Leu Ser Leu
Gln Ser Arg Pro Ser Ser Leu Pro625 630 635 640Asp Leu Ser Gly Ser
Ile Asp Asp Leu Pro Met Gly Thr Glu Gly Ala 645 650 655Leu Ser Pro
Gly Val Ser Thr Ser Gly Ile Ser Ser Ser Gln Gly Glu 660 665 670Gln
Ser Asn Pro Ala Gln Ser Pro Phe Ser Pro His Thr Ser Pro His 675 680
685Leu Pro Gly Ile Arg Gly Pro Ser Pro Ser Pro Val Gly Ser Pro Ala
690 695 700Ser Val Ala Gln Ser Arg Ser Gly Pro Leu Ser Pro Ala Ala
Val Pro705 710 715 720Gly Asn Gln Met Pro Pro Arg Pro Pro Ser Gly
Gln Ser Asp Ser Ile 725 730 735Met His Pro Ser Met Asn Gln Ser Ser
Ile Ala Gln Asp Arg Gly Tyr 740 745 750Met Gln Arg Asn Pro Gln Met
Pro Gln Tyr Ser Ser Pro Gln Pro Gly 755 760 765Ser Ala Leu Ser Pro
Arg Gln Pro Ser Gly Gly Gln Ile His Thr Gly 770 775 780Met Gly Ser
Tyr Gln Gln Asn Ser Met Gly Ser Tyr Gly Pro Gln Gly785 790 795
800Gly Gln Tyr Gly Pro Gln Gly Gly Tyr Pro Arg Gln Pro Asn Tyr Asn
805 810 815Ala Leu Pro Asn Ala Asn Tyr Pro Ser Ala Gly Met Ala Gly
Gly Ile 820 825 830Asn Pro Met Gly Ala Gly Gly Gln Met His Gly Gln
Pro Gly Ile Pro 835 840 845Pro Tyr Gly Thr Leu Pro Pro Gly Arg Met
Ser His Ala Ser Met Gly 850 855 860Asn Arg Pro Tyr Gly Pro Asn Met
Ala Asn Met Pro Pro Gln Val Gly865 870 875 880Ser Gly Met Cys Pro
Pro Pro Gly Gly Met Asn Arg Lys Thr Gln Glu 885 890 895Thr Ala Val
Ala Met His Val Ala Ala Asn Ser Ile Gln Asn Arg Pro 900 905 910Pro
Gly Tyr Pro Asn Met Asn Gln Gly Gly Met Met Gly Thr Gly Pro 915 920
925Pro Tyr Gly Gln Gly Ile Asn Ser Met Ala Gly Met Ile Asn Pro Gln
930 935 940Gly Pro Pro Tyr Ser Met Gly Gly Thr Met Ala Asn Asn Ser
Ala Gly945 950 955 960Met Ala Ala Ser Pro Glu Met Met Gly Leu Gly
Asp Val Lys Leu Thr 965 970 975Pro Ala Thr Lys Met Asn Asn Lys Ala
Asp Gly Thr Pro Lys Thr Glu 980 985 990Ser Lys Ser Lys Lys Ser Ser
Ser Ser Thr Thr Thr Asn Glu Lys Ile 995 1000 1005Thr Lys Leu Tyr
Glu Leu Gly Gly Glu Pro Glu Arg Lys Met Trp 1010 1015 1020Val Asp
Arg Tyr Leu Ala Phe Thr Glu Glu Lys Ala Met Gly Met 1025 1030
1035Thr Asn Leu Pro Ala Val Gly Arg Lys Pro Leu Asp Leu Tyr Arg
1040 1045 1050Leu Tyr Val Ser Val Lys Glu Ile Gly Gly Leu Thr Gln
Val Asn 1055 1060 1065Lys Asn Lys Lys Trp Arg Glu Leu Ala Thr Asn
Leu Asn Val Gly 1070 1075 1080Thr Ser Ser Ser Ala Ala Ser Ser Leu
Lys Lys Gln Tyr Ile Gln 1085
1090 1095Cys Leu Tyr Ala Phe Glu Cys Lys Ile Glu Arg Gly Glu Asp
Pro 1100 1105 1110Pro Pro Asp Ile Phe Ala Ala Ala Asp Ser Lys Lys
Ser Gln Pro 1115 1120 1125Lys Ile Gln Pro Pro Ser Pro Ala Gly Ser
Gly Ser Met Gln Gly 1130 1135 1140Pro Gln Thr Pro Gln Ser Thr Ser
Ser Ser Met Ala Glu Gly Gly 1145 1150 1155Asp Leu Lys Pro Pro Thr
Pro Ala Ser Thr Pro His Ser Gln Ile 1160 1165 1170Pro Pro Leu Pro
Gly Met Ser Arg Ser Asn Ser Val Gly Ile Gln 1175 1180 1185Asp Ala
Phe Asn Asp Gly Ser Asp Ser Thr Phe Gln Lys Arg Asn 1190 1195
1200Ser Met Thr Pro Asn Pro Gly Tyr Gln Pro Ser Met Asn Thr Ser
1205 1210 1215Asp Met Met Gly Arg Met Ser Tyr Glu Pro Asn Lys Asp
Pro Tyr 1220 1225 1230Gly Ser Met Arg Lys Ala Pro Gly Ser Asp Pro
Phe Met Ser Ser 1235 1240 1245Gly Gln Gly Pro Asn Gly Gly Met Gly
Asp Pro Tyr Ser Arg Ala 1250 1255 1260Ala Gly Pro Gly Leu Gly Asn
Val Ala Met Gly Pro Arg Gln His 1265 1270 1275Tyr Pro Tyr Gly Gly
Pro Tyr Asp Arg Val Arg Thr Glu Pro Gly 1280 1285 1290Ile Gly Pro
Glu Gly Asn Met Ser Thr Gly Ala Pro Gln Pro Asn 1295 1300 1305Leu
Met Pro Ser Asn Pro Asp Ser Gly Met Tyr Ser Pro Ser Arg 1310 1315
1320Tyr Pro Pro Gln Gln Gln Gln Gln Gln Gln Gln Arg His Asp Ser
1325 1330 1335Tyr Gly Asn Gln Phe Ser Thr Gln Gly Thr Pro Ser Gly
Ser Pro 1340 1345 1350Phe Pro Ser Gln Gln Thr Thr Met Tyr Gln Gln
Gln Gln Gln Val 1355 1360 1365Ser Ser Pro Ala Pro Leu Pro Arg Pro
Met Glu Asn Arg Thr Ser 1370 1375 1380Pro Ser Lys Ser Pro Phe Leu
His Ser Gly Met Lys Met Gln Lys 1385 1390 1395Ala Gly Pro Pro Val
Pro Ala Ser His Ile Ala Pro Ala Pro Val 1400 1405 1410Gln Pro Pro
Met Ile Arg Arg Asp Ile Thr Phe Pro Pro Gly Ser 1415 1420 1425Val
Glu Ala Thr Gln Pro Val Leu Lys Gln Arg Arg Arg Leu Thr 1430 1435
1440Met Lys Asp Ile Gly Thr Pro Glu Ala Trp Arg Val Met Met Ser
1445 1450 1455Leu Lys Ser Gly Leu Leu Ala Glu Ser Thr Trp Ala Leu
Asp Thr 1460 1465 1470Ile Asn Ile Leu Leu Tyr Asp Asp Asn Ser Ile
Met Thr Phe Asn 1475 1480 1485Leu Ser Gln Leu Pro Gly Leu Leu Glu
Leu Leu Val Glu Tyr Phe 1490 1495 1500Arg Arg Cys Leu Ile Glu Ile
Phe Gly Ile Leu Lys Glu Tyr Glu 1505 1510 1515Val Gly Asp Pro Gly
Gln Arg Thr Leu Leu Asp Pro Gly Arg Phe 1520 1525 1530Ser Lys Val
Ser Ser Pro Ala Pro Met Glu Gly Gly Glu Glu Glu 1535 1540 1545Glu
Glu Leu Leu Gly Pro Lys Leu Glu Glu Glu Glu Glu Glu Glu 1550 1555
1560Val Val Glu Asn Asp Glu Glu Ile Ala Phe Ser Gly Lys Asp Lys
1565 1570 1575Pro Ala Ser Glu Asn Ser Glu Glu Lys Leu Ile Ser Lys
Phe Asp 1580 1585 1590Lys Leu Pro Val Lys Ile Val Gln Lys Asn Asp
Pro Phe Val Val 1595 1600 1605Asp Cys Ser Asp Lys Leu Gly Arg Val
Gln Glu Phe Asp Ser Gly 1610 1615 1620Leu Leu His Trp Arg Ile Gly
Gly Gly Asp Thr Thr Glu His Ile 1625 1630 1635Gln Thr His Phe Glu
Ser Lys Thr Glu Leu Leu Pro Ser Arg Pro 1640 1645 1650His Ala Pro
Cys Pro Pro Ala Pro Arg Lys His Val Thr Thr Ala 1655 1660 1665Glu
Gly Thr Pro Gly Thr Thr Asp Gln Glu Gly Pro Pro Pro Asp 1670 1675
1680Gly Pro Pro Glu Lys Arg Ile Thr Ala Thr Met Asp Asp Met Leu
1685 1690 1695Ser Thr Arg Ser Ser Thr Leu Thr Glu Asp Gly Ala Lys
Ser Ser 1700 1705 1710Glu Ala Ile Lys Glu Ser Ser Lys Phe Pro Phe
Gly Ile Ser Pro 1715 1720 1725Ala Gln Ser His Arg Asn Ile Lys Ile
Leu Glu Asp Glu Pro His 1730 1735 1740Ser Lys Asp Glu Thr Pro Leu
Cys Thr Leu Leu Asp Trp Gln Asp 1745 1750 1755Ser Leu Ala Lys Arg
Cys Val Cys Val Ser Asn Thr Ile Arg Ser 1760 1765 1770Leu Ser Phe
Val Pro Gly Asn Asp Phe Glu Met Ser Lys His Pro 1775 1780 1785Gly
Leu Leu Leu Ile Leu Gly Lys Leu Ile Leu Leu His His Lys 1790 1795
1800His Pro Glu Arg Lys Gln Ala Pro Leu Thr Tyr Glu Lys Glu Glu
1805 1810 1815Glu Gln Asp Gln Gly Val Ser Cys Asn Lys Val Glu Trp
Trp Trp 1820 1825 1830Asp Cys Leu Glu Met Leu Arg Glu Asn Thr Leu
Val Thr Leu Ala 1835 1840 1845Asn Ile Ser Gly Gln Leu Asp Leu Ser
Pro Tyr Pro Glu Ser Ile 1850 1855 1860Cys Leu Pro Val Leu Asp Gly
Leu Leu His Trp Ala Val Cys Pro 1865 1870 1875Ser Ala Glu Ala Gln
Asp Pro Phe Ser Thr Leu Gly Pro Asn Ala 1880 1885 1890Val Leu Ser
Pro Gln Arg Leu Val Leu Glu Thr Leu Ser Lys Leu 1895 1900 1905Ser
Ile Gln Asp Asn Asn Val Asp Leu Ile Leu Ala Thr Pro Pro 1910 1915
1920Phe Ser Arg Leu Glu Lys Leu Tyr Ser Thr Met Val Arg Phe Leu
1925 1930 1935Ser Asp Arg Lys Asn Pro Val Cys Arg Glu Met Ala Val
Val Leu 1940 1945 1950Leu Ala Asn Leu Ala Gln Gly Asp Ser Leu Ala
Ala Arg Ala Ile 1955 1960 1965Ala Val Gln Lys Gly Ser Ile Gly Asn
Leu Leu Gly Phe Leu Glu 1970 1975 1980Asp Ser Leu Ala Ala Thr Gln
Phe Gln Gln Ser Gln Ala Ser Leu 1985 1990 1995Leu His Met Gln Asn
Pro Pro Phe Glu Pro Thr Ser Val Asp Met 2000 2005 2010Met Arg Arg
Ala Ala Arg Ala Leu Leu Ala Leu Ala Lys Val Asp 2015 2020 2025Glu
Asn His Ser Glu Phe Thr Leu Tyr Glu Ser Arg Leu Leu Asp 2030 2035
2040Ile Ser Val Ser Pro Leu Met Asn Ser Leu Val Ser Gln Val Ile
2045 2050 2055Cys Asp Val Leu Phe Leu Ile Gly Gln Ser 2060
2065127934DNAHomo sapiens 12cagaaagcgg agagtcacag cggggccagg
ccctggggag cggagcctcc accgcccccc 60tcattcccag gcaagggctt ggggggaatg
agccgggaga gccgggtccc gagcctacag 120agccgggagc agctgagccg
ccggcgcctc ggccgccgcc gccgcctcct cctcctccgc 180cgccgccagc
ccggagcctg agccggcggg gcggggggga gaggagcgag cgcagcgcag
240cagcggagcc ccgcgaggcc cgcccgggcg ggtggggagg gcagcccggg
ggactgggcc 300ccggggcggg gtgggagggg gggagaagac gaagacaggg
ccgggtctct ccgcggacga 360gacagcgggg atcatggccg cgcaggtcgc
ccccgccgcc gccagcagcc tgggcaaccc 420gccgccgccg ccgccctcgg
agctgaagaa agccgagcag cagcagcggg aggaggcggg 480gggcgaggcg
gcggcggcgg cagcggccga gcgcggggaa atgaaggcag ccgccgggca
540ggaaagcgag ggccccgccg tggggccgcc gcagccgctg ggaaaggagc
tgcaggacgg 600ggccgagagc aatgggggtg gcggcggcgg cggagccggc
agcggcggcg ggcccggcgc 660ggagccggac ctgaagaact cgaacgggaa
cgcgggccct aggcccgccc tgaacaataa 720cctcacggag ccgcccggcg
gcggcggtgg cggcagcagc gatggggtgg gggcgcctcc 780tcactcagcc
gcggccgcct tgccgccccc agcctacggc ttcgggcaac cctacggccg
840gagcccgtct gccgtcgccg ccgccgcggc cgccgtcttc caccaacaac
atggcggaca 900acaaagccct ggcctggcag cgctgcagag cggcggcggc
gggggcctgg agccctacgc 960ggggccccag cagaactctc acgaccacgg
cttccccaac caccagtaca actcctacta 1020ccccaaccgc agcgcctacc
ccccgcccgc cccggcctac gcgctgagct ccccgagagg 1080tggcactccg
ggctccggcg cggcggcggc tgccggctcc aagccgcctc cctcctccag
1140cgcctccgcc tcctcgtcgt cttcgtcctt cgctcagcag cgcttcgggg
ccatgggggg 1200aggcggcccc tccgcggccg gcgggggaac tccccagccc
accgccaccc ccaccctcaa 1260ccaactgctc acgtcgccca gctcggcccg
gggctaccag ggctaccccg ggggcgacta 1320cagtggcggg ccccaggacg
ggggcgccgg caagggcccg gcggacatgg cctcgcagtg 1380ttggggggct
gcggcggcgg cagctgcggc ggcggccgcc tcgggagggg cccaacaaag
1440gagccaccac gcgcccatga gccccgggag cagcggcggc ggggggcagc
cgctcgcccg 1500gacccctcag ccatccagtc caatggatca gatgggcaag
atgagacctc agccatatgg 1560cgggactaac ccatactcgc agcaacaggg
acctccgtca ggaccgcagc aaggacatgg 1620gtacccaggg cagccatacg
ggtcccagac cccgcagcgg tacccgatga ccatgcaggg 1680ccgggcgcag
agtgccatgg gcggcctctc ttatacacag cagattcctc cttatggaca
1740acaaggcccc agcgggtatg gtcaacaggg ccagactcca tattacaacc
agcaaagtcc 1800tcaccctcag cagcagcagc caccctactc ccagcaacca
ccgtcccaga cccctcatgc 1860ccaaccttcg tatcagcagc agccacagtc
tcaaccacca cagctccagt cctctcagcc 1920tccatactcc cagcagccat
cccagcctcc acatcagcag tccccggctc catacccctc 1980ccagcagtcg
acgacacagc agcaccccca gagccagccc ccctactcac agccacaggc
2040tcagtctcct taccagcagc agcaacctca gcagccagca ccctcgacgc
tctcccagca 2100ggctgcgtat cctcagcccc agtctcagca gtcccagcaa
actgcctatt cccagcagcg 2160cttccctcca ccgcaggagc tatctcaaga
ttcatttggg tctcaggcat cctcagcccc 2220ctcaatgacc tccagtaagg
gagggcaaga agatatgaac ctgagccttc agtcaagacc 2280ctccagcttg
cctgatctat ctggttcaat agatgacctc cccatgggga cagaaggagc
2340tctgagtcct ggagtgagca catcagggat ttccagcagc caaggagagc
agagtaatcc 2400agctcagtct cctttctctc ctcatacctc ccctcacctg
cctggcatcc gaggcccttc 2460cccgtcccct gttggctctc ccgccagtgt
tgctcagtct cgctcaggac cactctcgcc 2520tgctgcagtg ccaggcaacc
agatgccacc tcggccaccc agtggccagt cggacagcat 2580catgcatcct
tccatgaacc aatcaagcat tgcccaagat cgaggttata tgcagaggaa
2640cccccagatg ccccagtaca gttcccccca gcccggctca gccttatctc
cgcgtcagcc 2700ttccggagga cagatacaca caggcatggg ctcctaccag
cagaactcca tggggagcta 2760tggtccccag gggggtcagt atggcccaca
aggtggctac cccaggcagc caaactataa 2820tgccttgccc aatgccaact
accccagtgc aggcatggct ggaggcataa accccatggg 2880tgccggaggt
caaatgcatg gacagcctgg catcccacct tatggcacac tccctccagg
2940gaggatgagt cacgcctcca tgggcaaccg gccttatggc cctaacatgg
ccaatatgcc 3000acctcaggtt gggtcaggga tgtgtccccc accagggggc
atgaaccgga aaacccaaga 3060aactgctgtc gccatgcatg ttgctgccaa
ctctatccaa aacaggccgc caggctaccc 3120caatatgaat caagggggca
tgatgggaac tggacctcct tatggacaag ggattaatag 3180tatggctggc
atgatcaacc ctcagggacc cccatattcc atgggtggaa ccatggccaa
3240caattctgca gggatggcag ccagcccaga gatgatgggc cttggggatg
taaagttaac 3300tccagccacc aaaatgaaca acaaggcaga tgggacaccc
aagacagaat ccaaatccaa 3360gaaatccagt tcttctacta caaccaatga
gaagatcacc aagttgtatg agctgggtgg 3420tgagcctgag aggaagatgt
gggtggaccg ttatctggcc ttcactgagg agaaggccat 3480gggcatgaca
aatctgcctg ctgtgggtag gaaacctctg gacctctatc gcctctatgt
3540gtctgtgaag gagattggtg gattgactca ggtcaacaag aacaaaaaat
ggcgggaact 3600tgcaaccaac ctcaatgtgg gcacatcaag cagtgctgcc
agctccttga aaaagcagta 3660tatccagtgt ctctatgcct ttgaatgcaa
gattgaacgg ggagaagacc ctcccccaga 3720catctttgca gctgctgatt
ccaagaagtc ccagcccaag atccagcctc cctctcctgc 3780gggatcagga
tctatgcagg ggccccagac tccccagtca accagcagtt ccatggcaga
3840aggaggagac ttaaagccac caactccagc atccacacca cacagtcaga
tccccccatt 3900gccaggcatg agcaggagca attcagttgg gatccaggat
gcctttaatg atggaagtga 3960ctccacattc cagaagcgga attccatgac
tccaaaccct gggtatcagc ccagtatgaa 4020tacctctgac atgatggggc
gcatgtccta tgagccaaat aaggatcctt atggcagcat 4080gaggaaagct
ccagggagtg atcccttcat gtcctcaggg cagggcccca acggcgggat
4140gggtgacccc tacagtcgtg ctgccggccc tgggctagga aatgtggcga
tgggaccacg 4200acagcactat ccctatggag gtccttatga cagagtgagg
acggagcctg gaatagggcc 4260tgagggaaac atgagcactg gggccccaca
gccgaatctc atgccttcca acccagactc 4320ggggatgtat tctcctagcc
gctacccccc gcagcagcag cagcagcagc agcaacgaca 4380tgattcctat
ggcaatcagt tctccaccca aggcacccct tctggcagcc ccttccccag
4440ccagcagact acaatgtatc aacagcaaca gcaggtatcc agccctgctc
ccctgccccg 4500gccaatggag aaccgcacct ctcctagcaa gtctccattc
ctgcactctg ggatgaaaat 4560gcagaaggca ggtcccccag tacctgcctc
gcacatagca cctgcccctg tgcagccccc 4620catgattcgg cgggatatca
ccttcccacc tggctctgtt gaagccacac agcctgtgtt 4680gaagcagagg
aggcggctca caatgaaaga cattggaacc ccggaggcat ggcgggtaat
4740gatgtccctc aagtctggtc tcctggcaga gagcacatgg gcattagata
ccatcaacat 4800cctgctgtat gatgacaaca gcatcatgac cttcaacctc
agtcagctcc cagggttgct 4860agagctcctt gtagaatatt tccgacgatg
cctgattgag atctttggca ttttaaagga 4920gtatgaggtg ggtgacccag
gacagagaac gctactggat cctgggaggt tcagcaaggt 4980gtctagtcca
gctcccatgg agggtgggga agaagaagaa gaacttctag gtcctaaact
5040agaagaggaa gaagaagagg aagtagttga aaatgatgag gagatagcct
tttcaggcaa 5100ggacaagcca gcttcagaga atagtgagga gaagctgatc
agtaagtttg acaagcttcc 5160agtaaagatc gtacagaaga atgatccatt
tgtggtggac tgctcagata agcttgggcg 5220tgtgcaggag tttgacagtg
gcctgctgca ctggcggatt ggtggggggg acaccactga 5280gcatatccag
acccacttcg agagcaagac agagctgctg ccttcccggc ctcacgcacc
5340ctgcccacca gcccctcgga agcatgtgac aacagcagag ggtacaccag
ggacaacaga 5400ccaggagggg cccccacctg atggacctcc agaaaaacgg
atcacagcca ctatggatga 5460catgttgtct actcggtcta gcaccttgac
cgaggatgga gctaagagtt cagaggccat 5520caaggagagc agcaagtttc
catttggcat tagcccagca cagagccacc ggaacatcaa 5580gatcctagag
gacgaacccc acagtaagga tgagacccca ctgtgtaccc ttctggactg
5640gcaggattct cttgccaagc gctgcgtctg tgtgtccaat accattcgaa
gcctgtcatt 5700tgtgccaggc aatgactttg agatgtccaa acacccaggg
ctgctgctca tcctgggcaa 5760gctgatcctg ctgcaccaca agcacccaga
acggaagcag gcaccactaa cttatgaaaa 5820ggaggaggaa caggaccaag
gggtgagctg caacaaagtg gagtggtggt gggactgctt 5880ggagatgctc
cgggaaaaca ccttggttac actcgccaac atctcggggc agttggacct
5940atctccatac cccgagagca tttgcctgcc tgtcctggac ggactcctac
actgggcagt 6000ttgcccttca gctgaagccc aggacccctt ttccaccctg
ggccccaatg ccgtcctttc 6060cccgcagaga ctggtcttgg aaaccctcag
caaactcagc atccaggaca acaatgtgga 6120cctgattctg gccacacccc
ccttcagccg cctggagaag ttgtatagca ctatggtgcg 6180cttcctcagt
gaccgaaaga acccggtgtg ccgggagatg gctgtggtac tgctggccaa
6240cctggctcag ggggacagcc tggcagctcg tgccattgca gtgcagaagg
gcagtatcgg 6300caacctcctg ggcttcctag aggacagcct tgccgccaca
cagttccagc agagccaggc 6360cagcctcctc cacatgcaga acccaccctt
tgagccaact agtgtggaca tgatgcggcg 6420ggctgcccgc gcgctgcttg
ccttggccaa ggtggacgag aaccactcag agtttactct 6480gtacgaatca
cggctgttgg acatctcggt atcaccgttg atgaactcat tggtttcaca
6540agtcatttgt gatgtactgt ttttgattgg ccagtcatga cagccgtggg
acacctcccc 6600cccccgtgtg tgtgtgcgtg tgtggagaac ttagaaactg
actgttgccc tttatttatg 6660caaaaccacc tcagaatcca gtttaccctg
tgctgtccag cttctccctt gggaaaaagt 6720ctctcctgtt tctctctcct
ccttccacct cccctccctc catcacctca cgcctttctg 6780ttccttgtcc
tcaccttact cccctcagga ccctacccca ccctctttga aaagacaaag
6840ctctgcctac atagaagact ttttttattt taaccaaagt tactgttgtt
tacagtgagt 6900ttggggaaaa aaaataaaat aaaaatggct ttcccagtcc
ttgcatcaac gggatgccac 6960atttcataac tgtttttaat ggtaaaaaaa
aaaaaaaaaa atacaaaaaa aaattctgaa 7020ggacaaaaaa ggtgactgct
gaactgtgtg tggtttattg ttgtacattc acaatcttgc 7080aggagccaag
aagttcgcag ttgtgaacag accctgttca ctggagaggc ctgtgcagta
7140gagtgtagac cctttcatgt actgtactgt acacctgata ctgtaaacat
actgtaataa 7200taatgtctca catggaaaca gaaaacgctg ggtcagcagc
aagctgtagt ttttaaaaat 7260gtttttagtt aaacgttgag gagaaaaaaa
aaaaaggctt ttcccccaaa gtatcatgtg 7320tgaacctaca acaccctgac
ctctttctct cctccttgat tgtatgaata accctgagat 7380cacctcttag
aactggtttt aacctttagc tgcagcggct acgctgccac gtgtgtatat
7440atatgacgtt gtacattgca catacccttg gatccccaca gtttggtcct
cctcccagct 7500acccctttat agtatgacga gttaacaagt tggtgacctg
cacaaagcga gacacagcta 7560tttaatctct tgccagatat cgcccctctt
ggtgcgatgc tgtacaggtc tctgtaaaaa 7620gtccttgctg tctcagcagc
caatcaactt atagtttatt tttttctggg tttttgtttt 7680gttttgtttt
ctttctaatc gaggtgtgaa aaagttctag gttcagttga agttctgatg
7740aagaaacaca attgagattt tttcagtgat aaaatctgca tatttgtatt
tcaacaatgt 7800agctaaaact tgatgtaaat tcctcctttt tttccttttt
tggcttaatg aatatcattt 7860attcagtatg aaatctttat actatatgtt
ccacgtgtta agaataaatg tacattaaat 7920cttggtaaga cttt
79341326PRTArtificial SequenceSynthesized
peptideMISC_FEATURE(26)..(26)wherein the Lys is attached to a
biotin and an amide 13Ala Thr Lys Ala Ala Arg Lys Ser Ala Pro Ala
Thr Gly Gly Val Lys1 5 10 15Lys Pro His Arg Tyr Arg Pro Gly Gly Lys
20 251426PRTArtificial SequenceSynthesized
peptideMISC_FEATURE(7)..(7)wherein the Lys is
trimethylatedMISC_FEATURE(26)..(26)wherein the Lys is attached to a
biotin and an amide 14Ala Thr Lys Ala Ala Arg Lys Ser Ala Pro Ala
Thr Gly Gly Val Lys1 5 10 15Lys Pro His Arg Tyr Arg Pro Gly Gly Lys
20 25
References









































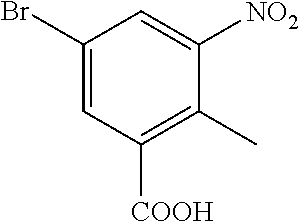



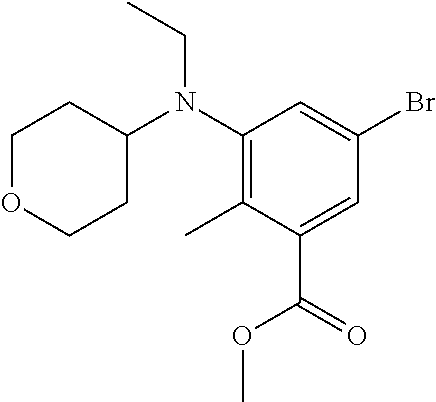






D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019
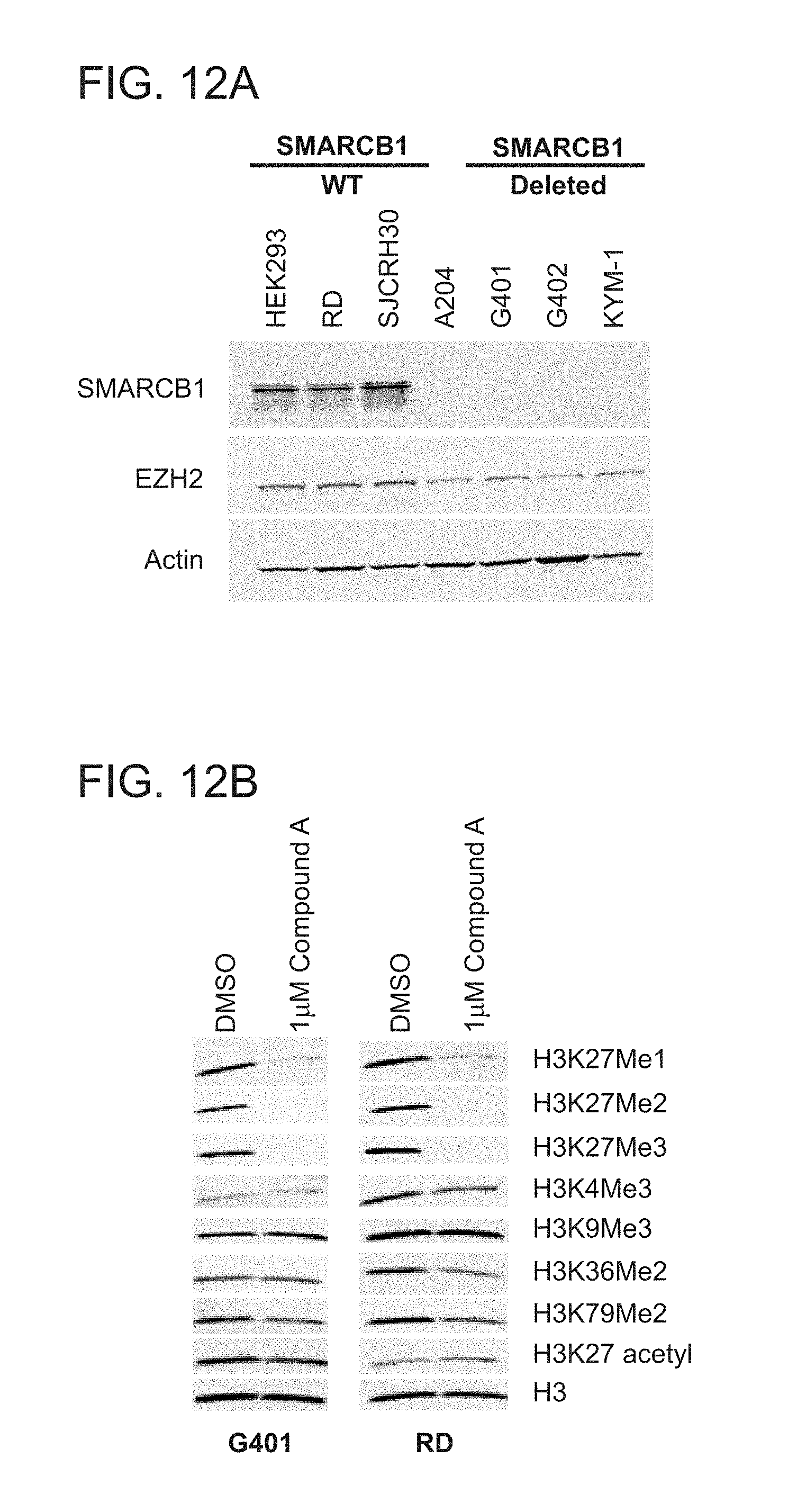
D00020

D00021

D00022

D00023

D00024
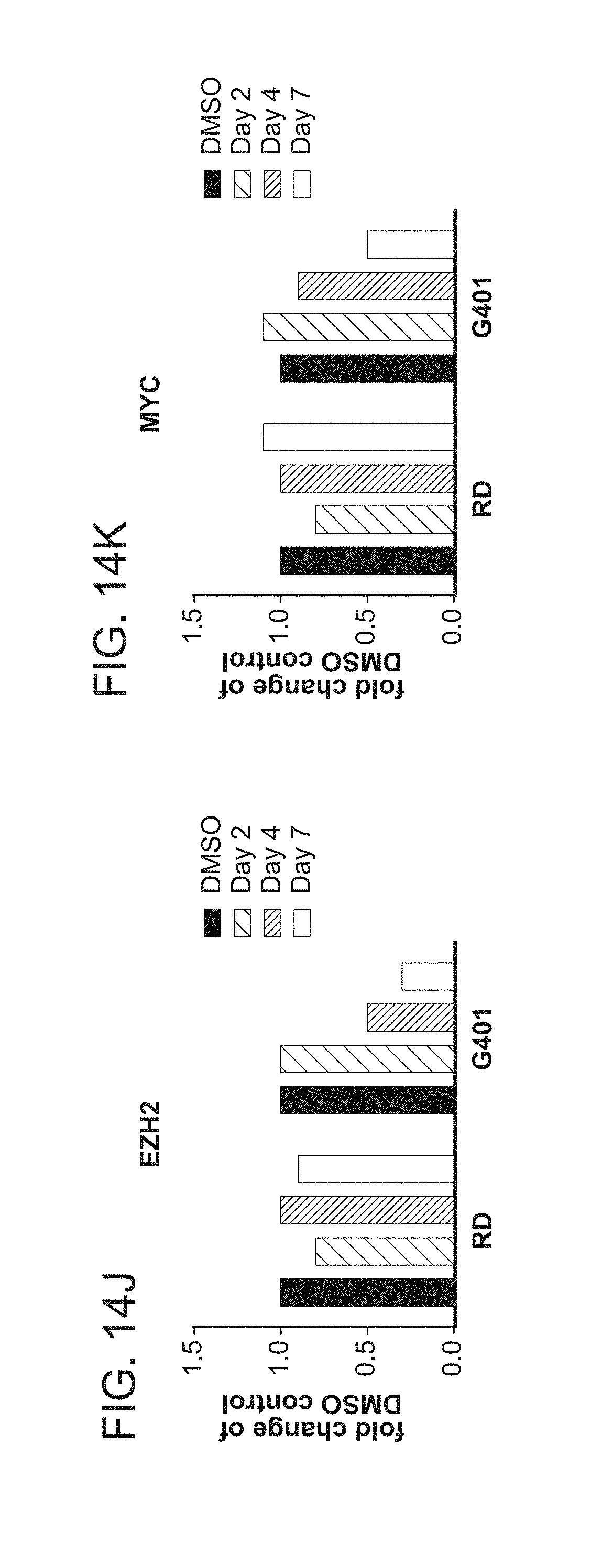
D00025

D00026

D00027

D00028

D00029
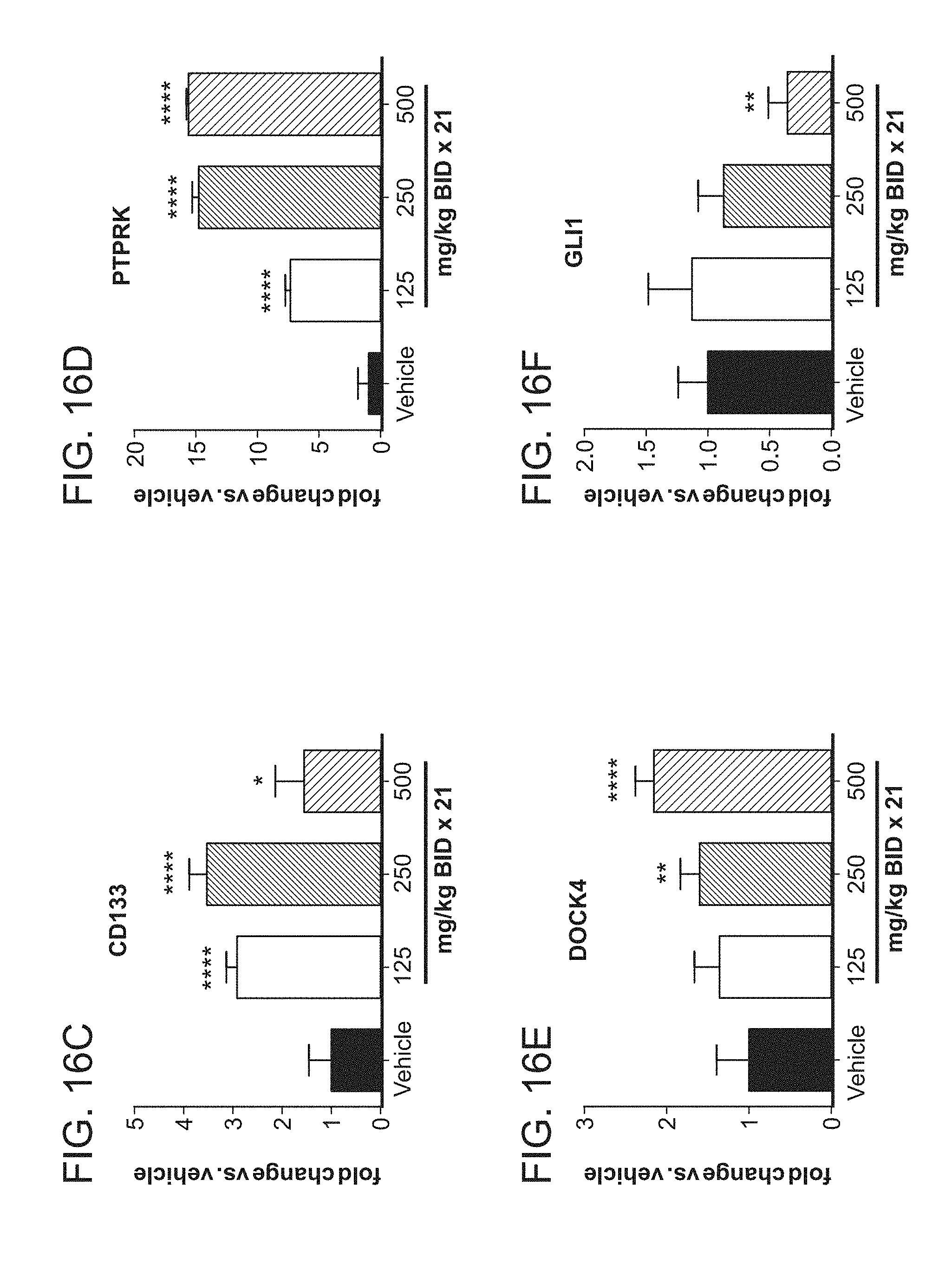




S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.