Anti-cd39 Antibodies, Compositions Comprising Anti-cd39 Antibodies And Methods Of Using Anti-cd39 Antibodies
Soros; Vanessa ; et al.
U.S. patent application number 16/049736 was filed with the patent office on 2019-02-28 for anti-cd39 antibodies, compositions comprising anti-cd39 antibodies and methods of using anti-cd39 antibodies. The applicant listed for this patent is Tizona Therapeutics. Invention is credited to Courtney Beers, John Corbin, Maria Kovalenko, Vanessa Soros, Joseph Robert Warfield, Paul Fredrick Widboom.
| Application Number | 20190062448 16/049736 |
| Document ID | / |
| Family ID | 65233024 |
| Filed Date | 2019-02-28 |





View All Diagrams
| United States Patent Application | 20190062448 |
| Kind Code | A1 |
| Soros; Vanessa ; et al. | February 28, 2019 |
ANTI-CD39 ANTIBODIES, COMPOSITIONS COMPRISING ANTI-CD39 ANTIBODIES AND METHODS OF USING ANTI-CD39 ANTIBODIES
Abstract
Provided herein are antibodies that selectively bind to CD39 and its isoforms and homologs, and compositions comprising the antibodies. Also provided are methods of using the antibodies, such as therapeutic and diagnostic methods.
| Inventors: | Soros; Vanessa; (South San Francisco, CA) ; Kovalenko; Maria; (South San Francisco, CA) ; Corbin; John; (South San Francisco, CA) ; Beers; Courtney; (South San Francisco, CA) ; Widboom; Paul Fredrick; (Lebanon, NH) ; Warfield; Joseph Robert; (Lebanon, NH) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 65233024 | ||||||||||
| Appl. No.: | 16/049736 | ||||||||||
| Filed: | July 30, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 62539527 | Jul 31, 2017 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C07K 2317/77 20130101; C07K 2317/92 20130101; C07K 2317/31 20130101; C07K 2317/76 20130101; C07K 16/2896 20130101; C07K 2317/21 20130101; A61K 2039/505 20130101; A61P 37/04 20180101; A61K 39/3955 20130101; A61K 45/06 20130101; G01N 2333/70596 20130101; G01N 2500/04 20130101; C07K 2317/24 20130101; G01N 33/566 20130101; C07K 2317/33 20130101 |
| International Class: | C07K 16/28 20060101 C07K016/28; G01N 33/566 20060101 G01N033/566; A61K 45/06 20060101 A61K045/06; A61K 39/395 20060101 A61K039/395 |
Claims
1. An antigen binding protein that binds specifically to a human CD39 (hCD39) and is capable of 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, or 18 of the following: a) inhibiting binding of CD39 to ATP; b) inhibiting conversion by CD39 of ATP to ADP and/or ADP to AMP; c) decreasing affinity of CD39 for ATP or ADP; d) inhibiting or impeding release of ADP or AMP from CD39; e) impeding or inhibiting CD39 processivity; f) inhibiting platelet aggregation; g) decreasing levels of phosphate, ADP, AMP, and/or adenosine and/or increasing levels of ATP; h) increasing T effector cell function; i) decreasing the number of regulatory T cells in tissues or in circulation; j) suppressing regulatory T cells or regulatory T cell activity; k) increasing B cell function; l) increasing antigen presenting cell function; m) inhibiting CD39 function on tumor cells; n) inhibiting processing of at least one of phospho-antigen from phosphorylated isoprenoid, phosphorylated vitamin B metabolite, and/or phosphorylated riboflavin; o) decreasing or preventing activation of phospho antigen specific T cells selected from MAIT cells and .gamma..delta. T cells; p) inhibiting angiogenesis; q) increasing proliferation of stimulated CD4.sup.+ and CD8.sup.+ T cells; r) increasing stimulated PBMC Secretion of INF-.gamma., TNF-.alpha., IL-2 and/or IL-1.beta..
2. The antigen binding protein of claim 1, wherein the antigen binding protein has 1, 2, 3, 4, 5, 6, or 7 of the following characteristics: a) is a monoclonal antibody; b) is a human antibody, a humanized antibody, or a chimeric antibody; c) is a bispecific antibody, a multi-specific antibody, a diabody, or a multivalent antibody; d) is of the IgG1, IgG2, IgG3, IgG4, or IgM type; e) is an antigen-binding antibody fragment; f) is a Fab fragment, a Fab' fragment, a F(ab')2 fragment, or an Fv fragment; and/or g) is a single chain antibody, a single domain antibody, or a nanobody.
3. A pharmaceutical composition comprising an effective amount of an antibody which binds to hCD39 and has 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, or 16 of the following of characteristics: a) blocks or decreases hydrolysis of ATP to ADP and/or ADP to AMP as determined by at least one of: (i) a decreased phosphate release (Pi), (ii) an increase in ATP levels, and (iii) a decrease of ADP, AMP, and/or adenosine levels, b) increases T effector cell activity; c) suppresses regulatory T cell or decreases a regulatory T cell activity; d) decreases number of regulatory T cells in tissues or in circulation; e) increases B cell function; f) increases antigen presenting cell function; g) inhibits CD39 function on tumor cells; h) blocks or inhibits processing of at least one of phospho-antigen from a phosphorylated isoprenoid, phosphorylated vitamin B metabolite, and phosphorylated riboflavin; i) decreases or prevents activation of phospho antigen specific T cells selected from MAIT cells and .gamma..delta. T cells; k) inhibits angiogenesis; l) decreases affinity for ATP and/or ADP; m) inhibits release of ADP or AMP from CD39; n) impedes or inhibits CD39 processivity; o) inhibits platelet aggregation; p) increases proliferation of stimulated CD4.sup.+ and CD8.sup.+ T cells; r) increases stimulated PBMC Secretion of INF-.gamma., TNF-.alpha., IL-2, and/or IL-1.beta..
4. A pharmaceutical composition comprising the antigen-binding protein of claim 1 or claim 2.
5. The pharmaceutical composition of claim 4, further comprising an effective amount of at least one of the following a) an anti-PD-1 antibody, b) an anti-PD-L1 antibody, c) an anti-CD73 antibody, d) an anti-CD38 antibody, e) an anti-A2A receptor antibody, f) an anti-A2B receptor antibody, g) an anti-A2A/A2B dual receptor antibody, or any combination thereof, or h) a small molecule inhibitor, or a combination thereof.
6. The pharmaceutical composition of claim 4 or claim 5, further comprising one or both of a) an antibody to an inhibitory receptor or ligand and/or b) an antibody to a costimulatory receptor or ligand, or a combination thereof.
7. The pharmaceutical composition of claim 6, wherein (i) the inhibitory receptor or ligand is at least one of CTLA-4, PD-L2, LAG-3, Tim3, neuritin, BTLA, CECAM-1, CECAM-5, VISTA, LAIR1, CD160, 2B4, TGF-R, HHLA2, ILT2, ILT3, ILT4, HLA-G, HLA-C, and/or a Killer-cell immunoglobulin-like receptor (KIR) and/or (ii) the costimulatory receptor or ligand is at least one of OX40, CD2, CD27, CDS, ICAM-1, LFA-1, ICOS (CD278), 4-1BB (CD137), GITR, CD28, CD30, CD40, BAFFR, HVEM, CD7, LIGHT, NKG2C, SLAMF7, NKp80, CD160, B7-H3, and/or CD83.
8. The pharmaceutical composition of claim 7, wherein the costimulatory receptor or ligand LFA-1 further comprises an LFA-1 .beta.-chain CD18 and/or an LFA-1 .alpha.-chain CD11a.
9. The antigen binding protein of claim 1, wherein the antigen binding protein has one or more of the following characteristics: a) binds to a human CD39 polypeptide or a variant thereof with a K.sub.D of less than about 20 nM; b) binds to a cyno CD39 polypeptide or a variant thereof with a K.sub.D of less than about 200 nM; c) binds to a murine CD39 polypeptide or a variant thereof with a K.sub.D of less than about 200 nM; or d) a combination of at least 2 of a), b), and c).
10. An antigen binding protein that competes or is capable of competing for binding to human CD39 with a reference antigen binding protein, wherein the reference antigen binding protein is the antigen binding protein of claim 1.
11. An antigen binding protein that binds to or is capable of competing for binding to human CD39 with a reference antigen binding protein, wherein the reference antigen binding protein binds to an epitope at positions 143-158 or 274-277 of SEQ ID NO: 249 on a human CD39 polypeptide, including, but not limited to, D150, E153, R154 or N99 alone or in combination with D150, E153, R154 or any combination thereof.
12. The antigen binding protein of claim 10 or claim 11, wherein the antigen binding protein and the reference antibody cross-compete or are capable of cross-competing for binding to a human CD39.
13. The antigen binding protein of claim 1, comprising a human heavy chain constant region or fragment or a variant thereof, wherein the constant region variant comprises up to 20 conservatively modified amino acid substitutions from any sequence set forth SEQ ID NOs: 179-218.
14. An isolated antibody molecule capable of binding to human CD39 (hCD39), comprising a heavy chain variable region (VH) and a light chain variable region (VL), VH and/or VL comprising 1, 2, 3, 4, 5, or 6 of: a) a VHCDR1 having the sequence set forth in SEQ ID NOs: 1-45, b) a VHCDR2 having the sequence set forth in SEQ ID NOs: 46-81, c) a VHCDR3 having the sequence set forth in SEQ ID NOs: 82-109, d) a VLCDR1 having the sequence set forth in SEQ ID NOs: 110-124, e) a VLCDR2 having the sequence set forth in SEQ ID NOs: 125-140, and f) a VLCDR3 having the sequence set forth in SEQ ID NOs: 141-166.
15. An isolated antibody molecule capable of binding to human CD39 (hCD39), comprising a heavy chain variable region (VH) and a light chain variable region (VL), the VH comprising, a) a VHCDR1 having a sequence set forth in SEQ ID NOs: 1-45, b) a VHCDR2 having a sequence set forth in SEQ ID NOs: 46-81, and c) a VHCDR3 having a sequence set forth in SEQ ID NOs: 82-109; and the VL comprising, a) a VLCDR1 having a sequence set forth in SEQ ID NO: 110-124, b) a VLCDR2 having a sequence set forth in SEQ ID NO: 125-140, and c) a VLCDR3 having a sequence set forth in SEQ ID NO: 141-166.
16. An isolated nucleic acid encoding an antigen binding protein according to claim 1, claim 14, or claim 15.
17. An expression vector comprising the nucleic acid according to claim 16.
18. A prokaryotic or eukaryotic host cell comprising the vector of claim 17.
19. An oncolytic virus encoding the nucleic acid of either of claim 16 or 17.
20. A method for the production of a recombinant protein comprising the steps of expressing a nucleic acid according to claim 16 in a prokaryotic or eukaryotic host cell and recovering the protein from the cell or the cell culture supernatant.
21. A method for treatment of a subject suffering from cancer, a chronic infection, or from an inflammatory disease, comprising the step of administering to the subject a pharmaceutical composition comprising an effective amount of the antigen binding protein of claim 1 or the pharmaceutical composition of claim 3.
22. The method of claim 21, wherein the cancer is a solid cancer.
23. The method of claim 21, wherein the cancer is a hematological cancer.
24. A method for modulating immune system function in a subject in need thereof, comprising the step of contacting a population of immune cells of the subject with a pharmaceutical composition comprising an effective amount of the antigen binding protein of claim 1, under conditions such that the immune system is modulated.
25. A method for inducing or enhancing an immune response in a subject in need thereof, comprising the step of administering to the subject a pharmaceutical composition comprising an antigen binding protein, wherein the immune response is generated against a tumor antigen.
26. The method of claim 24 or claim 25, wherein the subject is a human subject.
27. The method of claim 25, wherein the antigen binding protein comprises a bispecific antibody or a complexing antigen binding protein.
28. The method of claim 27, wherein the antigen binding protein, the bispecific antibody, or the complexing antigen binding protein is administered in an amount sufficient to achieve 1, 2, 3, 4, 5, 6, or 7 of the following in the subject: a) reduction of CD39 ATPase activity in a target cell population; b) reduction of regulatory T cells suppression of activity of effector T cells; c) reduction of levels of regulatory T cells; d) activation of effector T cells; e) induction or enhancement of effector T cell proliferation; f) inhibition of tumor growth; and/or g) induction of tumor regression.
29. The method of claim 28, wherein the target cell population comprises T cells, B cells, monocytes, macrophages, dendritic cells, myeloid-derived suppressor cells, and/or tumor cells.
30. The method of claim 25, wherein the method further comprises one or more of the following a) administering chemotherapy; b) administering radiation therapy; and/or c) administering one or more additional therapeutic agents.
31. The method of claim 30, wherein the one or more additional therapeutic agents comprise one or more immunostimulatory agents.
32. The method of claim 31, wherein the one or more immunostimulatory agents comprise an antagonist to an inhibitory receptor of an immune cell.
33. The method of claim 32, wherein the inhibitory receptor is at least one of CTLA-4, PD-1, PD-L1, PD-L2, LAG-3, Tim3, neuritin, BTLA, CECAM-1, CECAM-5, VISTA, LAIR1, CD160, 2B4, TGF-R, and/or a Killer-cell immunoglobulin-like receptor (KIR).
34. The method of claim 31, wherein the one or more immunostimulatory agents comprise an agonist of a co-stimulatory receptor of an immune cell.
35. The method of claim 34, wherein the co-stimulatory receptor is OX40, CD2, CD27, CDS, ICAM-1, LFA-1, ICOS (CD278), 4-1BB (CD137), GITR, CD28, CD30, CD40, BAFFR, HVEM, CD7, LIGHT, NKG2C, SLAMF7, NKp80, CD160, B7-H3, or a CD83 ligand.
36. The method of claim 35, wherein the costimulatory receptor or ligand LFA-1 further comprises an LFA-1 .beta.-chains CD18 and/or an LFA-1 .alpha.-chain CD11a.
37. The method of claim 31, wherein the one or more immunostimulatory agents comprise a cytokine.
38. The method of claim 37, wherein the cytokine is at least one of IL-2, IL-5, IL-7, IL-12, IL-15, and/or IL-21.
39. The method of claim 31, wherein the one or more immunostimulatory agents comprise an oncolytic virus.
40. The method of claim 39, wherein the oncolytic virus is a Herpes simplex virus, a Vesicular stomatitis virus, an adenovirus, a Newcastle disease virus, a vaccinia virus, or a maraba virus.
41. The method of claim 31, wherein the one or more immunostimulatory agents comprise a chimeric antigen engineered T cell.
42. The method of claim 31, wherein the one or more immunostimulatory agents comprise a bi- or multispecific T cell directed antibody.
43. The method of claim 25, wherein the one or more additional therapeutic agents comprise at least one trap for an immune suppressive agent.
44. The method of claim 43, wherein the at least one trap for an immune suppressive agent comprises at least one of an anti-TGF-beta antibody, a TGFb receptor trap, an anti-IL-10 antibody and/or anti-IL-10 receptor trap, and/or an anti-IL-35 antibody trap and/or an anti-IL-35 receptor trap.
45. The method of any one of any of claims 3-44, wherein administration of the pharmaceutical composition results in induction or enhancement of proliferation of a T-effector cell, or modulation of I-kappaB and/or NF-.kappa.B in the T cell, or modulation of CD39 activity in the T cell, or T cell receptor induced signaling in a T-effector cell, or a combination thereof.
46. A method of screening for a test compound comprising an antigen binding protein of claim 1 capable of inhibiting an activity of CD39, comprising the steps of: contacting a test sample containing CD39 with a test compound; comparing the activity of the test sample to a control sample; whereby a decrease in the activity of CD39 in the test sample compared to the control sample identifies the compound as one that inhibits the activity of CD39.
47. The method of claim 46, wherein the control sample comprises a sample not contacted with a test compound.
48. An isolated antibody molecule capable of binding to human CD39 (hCD39), comprising a heavy chain variable region (VH) and a light chain variable region (VL), VH comprising at least one of: a) a VHCDR1 having an amino acid sequence that is at least 90% identical to the sequence set forth in SEQ ID NOs: 1-45, b) a VHCDR2 having an amino acid sequence that is at least 90% identical to the sequence set forth in SEQ ID NOs: 46-81, and c) a VHCDR3 having an amino acid sequence that is at least 90% identical to the sequence set forth in SEQ ID NOs: 82-109; and VL comprising at least one of: a) a VLCDR1 having an amino acid sequence that is at least 90% identical to the sequence set forth in SEQ ID NOs: 110-124, b) a VLCDR2 having an amino acid sequence that is at least 90% identical to the sequence set forth in SEQ ID NOs: 125-140, and c) a VLCDR3 having an amino acid sequence that is at least 90% identical to the sequence set forth in SEQ ID NOs 141-166.
49. An isolated antibody molecule capable of binding to human CD39 (hCD39), comprising a heavy chain variable region (VH) and a light chain variable region (VL), VH comprising at least one of: a) a VHCDR1 having an amino acid sequence that is homologous to the sequence set forth in SEQ ID NOs: 1-45, b) a VHCDR2 having an amino acid sequence that is homologous to the sequence set forth in SEQ ID NOs: 46-81, and c) a VHCDR3 having an amino acid sequence that is homologous to the sequence set forth in SEQ ID NOs: 82-109; and VL comprising at least one of: a) a VLCDR1 having an amino acid sequence that is homologous to the sequence set forth in SEQ ID NOs: 110-124, b) a VLCDR2 having an amino acid sequence that is homologous to the sequence set forth in SEQ ID NOs: 125-140, and c) a VLCDR3 having an amino acid sequence that is homologous to the sequence set forth in SEQ ID NOs: 141-166.
50. An isolated antibody molecule capable of binding to human CD39 (hCD39), comprising a heavy chain and a light chain, the heavy chain comprising one or more molecules having a sequence consisting of one of SEQ ID NO: 255, SEQ ID NO: 257, SEQ ID NO: 259, SEQ ID NO: 261, SEQ ID NO: 263, SEQ ID NO: 265, SEQ ID NO: 267, SEQ ID NO: 269, SEQ ID NO: 271, SEQ ID NO: 273, SEQ ID NO: 275, SEQ ID NO: 277, SEQ ID NO: 279, SEQ ID NO: 281, SEQ ID NO: 283, SEQ ID NO: 285, SEQ ID NO: 287, SEQ ID NO: 289, SEQ ID NO: 291, SEQ ID NO: 293, or SEQ ID NO: 295 and the light chain comprising one or more molecules having a sequence consisting of one of SEQ ID NO: 256, SEQ ID NO: 258, SEQ ID NO: 260, SEQ ID NO: 262, SEQ ID NO: 264, SEQ ID NO: 266, SEQ ID NO: 268, SEQ ID NO: 270, SEQ ID NO: 272, SEQ ID NO: 274, SEQ ID NO: 276, SEQ ID NO: 278, SEQ ID NO: 280, SEQ ID NO: 282, SEQ ID NO: 284, SEQ ID NO: 286, SEQ ID NO: 288, SEQ ID NO: 290, SEQ ID NO: 292, SEQ ID NO: 294, or SEQ ID NO: 296.
51. An isolated antibody molecule capable of binding to human CD39 (hCD39), comprising a heavy chain and a light chain, a) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 255 and the light chain comprising one or more, each molecule having a sequence consisting of SEQ ID NO: 256; b) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 257 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 258; c) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 259 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 260; d) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 261 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 262; e) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 263 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 264; f) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 265 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 266; g) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 267 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 268; h) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 269 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 270; i) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 271 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 272; j) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 273 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 274; k) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 275 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 276; l) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 277 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 278 m) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 279 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 280; n) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 281 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 282; o) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 283 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 284; p) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 285 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 286; q) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 287 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 288; r) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 289 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 290; s) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 291 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 292; t) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 293 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 294; or u) the heavy chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 295 and the light chain comprising one or more molecules, each molecule having a sequence consisting of SEQ ID NO: 296.
52. An isolated nucleic acid encoding an antigen binding protein according to any one of claims 48-51.
53. An isolated antibody molecule capable of binding to human CD39 (hCD39), comprising a heavy chain variable region (VH) and a light chain variable region (VL), VH and/or VL comprising 1, 2, 3, 4, 5, or 6 of: a) a VHCDR1 sequence comprising: (i) a Kabat CDRH1 sequence defined by the consensus sequence S-Y-.DELTA..sub.3-M-.DELTA..sub.5 (SEQ ID NOS: 25-29 and 44-45), where .DELTA..sub.3 is E, F, Q, or Y and .DELTA..sub.5 is H or Y; (ii) a Kabat CDR-H1 sequence defined by the consensus sequence .theta..sub.1-.theta..sub.2-.theta..sub.3-I-S (SEQ ID NOS: 30-37), where .theta..sub.1 is A, H, K, L, S, or W; .theta..sub.2 is L, M, N, or T; and .theta..sub.3 is A or P; (iii) a Kabat CDR-H1 sequence defined by the consensus sequence .eta..sub.1-Y-.eta..sub.3-I-S SEQ ID NOS: 38-41), where .eta..sub.1 is S, K, N, or R and .eta..sub.3 is A or G; (iv) a Chothia CDR-H1 sequence defined by the consensus sequence G-Y-T-F-.OMEGA..sub.5-S-Y (SEQ ID NOS: 1-2 and 4-6), where .OMEGA..sub.5 is T, K, Q, F, or V; (v) a Chothia CDR-H1 sequence defined by the consensus sequence G-G-T-F-.nu..sub.5-.nu..sub.6-Y (SEQ ID NOS: 17-22 and 24), where .nu..sub.5 is S, G, or E and .nu..sub.6 is S, K, R, or S; or (vi) a Chothia CDR-H1 consensus sequence defined by the consensus sequence G-G-T-F-.kappa..sub.5-.kappa..sub.6-.kappa..sub.7 (SEQ ID NOS: 7-16), where .kappa..sub.5 is S, Q, P, or A; .kappa..sub.6 is S, K, H, L, A, or W; and .kappa..sub.7 is Y, L, T, N, or M, b) a VHCDR2 sequence comprising: (i) a Kabat CDR-H2 sequence defined by the consensus sequence .epsilon..sub.1-I-N-P-.epsilon..sub.5-.epsilon..sub.6-G-S-T-.epsilon..sub- .10-Y-A-Q-K-F-Q-G (SEQ ID NOS: 63-66 and 68), where .epsilon..sub.1 is K, S, R, or V; .epsilon..sub.5 is L, R, or S; .epsilon..sub.6 is G or V; and .epsilon..sub.10 is S or W; (ii) a Kabat CDR-H2 sequence defined by the consensus sequence G-I-.alpha..sub.3-.alpha..sub.4-.alpha..sub.5-.alpha..sub.6-G-T-A-N-Y-A-Q- -K-F-Q-G (SEQ ID NOS: 69-72), where .alpha..sub.3 is I or L or is absent; .alpha..sub.4 is P or is absent; and .alpha..sub.5 is I, G, or R; and .alpha..sub.6 is A, F, or G; (iii) a Kabat CDR-H2 sequence defined by the consensus sequence .beta..sub.1-I-I-P-.beta..sub.5-.beta..sub.6-G-.beta..sub.8-A-N-Y-A-Q-K-F- -G-Q (SEQ ID NOS: 74 and 76-79), where .beta..sub.1 is S or G; .beta..sub.5 is I, E, S, or T; .beta..sub.6 is F, I, or S; and .beta..sub.8 is I or T; (iv) a Chothia CDR-H2 sequence defined by the consensus sequence N-P-.epsilon..sub.5-.epsilon..sub.6-G-S-T (SEQ ID NOS: 46-48), where .epsilon..sub.5 is L, R, or S and .epsilon..sub.6 is G or V; (v) a Chothia CDR-H2 sequence defined by the consensus sequence .alpha..sub.3-.alpha..sub.4-.alpha..sub.5-.alpha..sub.6-G-T-A (SEQ ID NOS: 51-54), where .alpha.3 is I or L or is absent; .alpha..sub.4 is P or is absent; and .alpha..sub.5 is I, G, or R; and .alpha.6 is A, F, or G; or (vi) a Chothia CDR-H2 sequence defined by the consensus sequence I-P-.beta..sub.5-.beta..sub.6-G-.beta..sub.8-A (SEQ ID NOS: 56-60), where .beta..sub.5 is I, E, S, or T; .beta..sub.6 is F, I, or S; and .beta..sub.8 is I or T, c) a VHCDR3 sequence comprising: (i) a CDR-H3 sequence defined by the consensus sequence G-K-R-E-G-G-T-E-Y-L-R-.sub.12 (SEQ ID NOS: 82-86), where .sub.12 is H, K, S, N, or V; (ii) a CDR-H3 sequence defined by the consensus sequence E-S-G-.PHI..sub.4-Y-R-D-H-R-L-.PHI..sub.11-V (SEQ ID NOS: 94-96), where .PHI..sub.4 is G or T and .PHI..sub.11 is D or G; or (iii) a CDR-H3 sequence defined by the consensus sequence G-G-A-K-Y-A-.sub.7-.sub.8-.sub.9-G-M-D-V (SEQ ID NOS: 87-93), where .sub.7 is S, V, G, or R; .sub.8 is T, Q, K, G, or R; and .sub.9 is Y, H, L, or W, d) a VLCDR1 sequence comprising: (i) a CDR-L1 sequence defined by the consensus sequence .PHI..sub.1-A-S-.PHI..sub.4-.PHI..sub.5-V-.PHI..sub.7-.PHI..sub.8-.PHI..s- ub.9-Y-L-A (SEQ ID NOS: 1101-114), where .PHI..sub.1 is E, K, or R; .PHI..sub.4 is Q or E; .PHI..sub.5 is S or Y; .PHI..sub.7 is S or A; .PHI..sub.8 is S or Y; and .PHI..sub.9 is D or S; (ii) a CDR-L1 sequence defined by the consensus sequence .sub.1-A-S-Q-.sub.5-.sub.6-.sub.7-.sub.8-.sub.9-L-.sub.11 (SEQ ID NOS: 118 and 120-123), where .sub.1 is Q or Rl; .sub.5 is D or S; .sub.6 is I or V; .sub.7 is G or S; .sub.8 is N, R, or S; .sub.9 is N, Y, or W; and .sub.11 is A or N; or (iii) a CDR-L1 sequence defined by the consensus sequence K-S-S-.GAMMA..sub.4-S-V-L-.GAMMA..sub.8-S-.GAMMA..sub.10-N-N-K-N- -Y-L-A (SEQ ID NOS: 115-117), where .GAMMA..sub.4 is Q, R or K; .GAMMA..sub.8 is F or Y; and .GAMMA..sub.10 is S or N, e) a VLCDR2 sequence comprising: (i) a CDR-L2 sequence defined by the consensus sequence .psi..sub.1-A-S-.psi..sub.4-R-.psi..sub.6-.psi..sub.7 (SEQ ID NOS: 125-136), where .psi..sub.1 is G or Y, .psi..sub.4 is S or N; .psi..sub.6 is A or H; and .psi..sub.7 is T, Y, or N; (ii) a CDR-L2 sequence defined by the consensus sequence D-A-S-.chi..sub.4-R-A-T (SEQ ID NOS: 138 and 139), where .English Pound..sub.4 is N or K; or (iii) a CDR-L2 sequence defined by the consensus sequence W-A-S-T-R-.sigma..sub.6-S (SEQ ID NOS: 131 and 133-134), where .sigma..sub.6 is A, E, or Q, and f) a VLCDR3 sequence comprising: (i) a CDR-L3 sequence defined by the consensus sequence Q-Q-Y-.pi..sub.4-.pi..sub.5-.pi..sub.6-.pi..sub.7-T (SEQ ID NOS: 141-147), where .pi..sub.4 is G, H, or Y; .pi..sub.5 is S, N, F, G, or R; .pi..sub.6 is S, Y, A, G, or R; and .pi..sub.7 is P, I, or L; (ii) a CDR-L3 sequence defined by consensus sequence Q-Q.lamda..sub.3-.lamda..sub.4-.lamda..sub.5-.lamda..sub.6-P-T (SEQ ID NOS: 148-150), where .lamda..sub.3 is R, F, H, S, L, D, Y, or V; .lamda..sub.4 is S, V, T, G, L, Y, or N; .lamda..sub.5 is N, L, F, K, or V; and .lamda..sub.6 is W, F, Y, or L; (iii) a CDR-L3 sequence defined by the consensus sequence Q-Q-Y-.rho..sub.3-.rho..sub.4-W-P-L-T (SEQ ID NOS: 151 and 152), where .rho..sub.3 is N or L and .rho..sub.4 is N or L; or (iv) a CDR-L3 sequence defined by the consensus sequence Q-Q-.omega..sub.3-.omega..sub.4-.omega..sub.5-.omega..sub.6-P-.omega..sub- .8-T (SEQ ID NOS: 153-156), where .omega..sub.3 is Y or F; .omega..sub.4 is Y or W; .omega..sub.5 is S, L, T, or F; .omega..sub.6 is T, Y, or F; and .omega..sub.8 is L or P.
Description
RELATED APPLICATION
[0001] This application claims priority to U.S. provisional application No. 62/539,527, filed Jul. 31, 2017, which is incorporated by reference herein in its entirety.
FIELD
[0002] Provided herein are antibodies with binding specificity for CD39 and compositions comprising the antibodies, including pharmaceutical compositions, diagnostic compositions and kits. Also provided are methods of using anti-CD39 antibodies for therapeutic and diagnostic purposes.
BACKGROUND
[0003] CD39 is an integral membrane protein that phosphohydrolyzes ATP to yield ADP and AMP. Human CD39 is a 510-amino acid protein with seven potential N-linked glycosylation sites, 11 cysteine residues, and two transmembrane regions. Structurally, it is characterized by two transmembrane domains, a small cytoplasmic domain comprising the NH.sub.2- and COOH-terminal segments, and a large extracellular hydrophobic domain consisting of five highly conserved domains, known as apyrase conserved regions (ACR) 1-5, which are pivotal for the catabolic activity of the enzyme. CD39 becomes catalytically active upon its localization on the cell surface, and its glycosylation is important for protein folding, membrane targeting, and enzyme activity.
[0004] CD39 is constitutively expressed in spleen, thymus, lung, and placenta and in these tissues it is associated primarily with endothelial cells and immune cell populations, such as B cells, natural killer (NK) cells, dendritic cells, Langerhans cells, monocytes, macrophages, mesangial cells, neutrophils, and regulatory T cells (Tregs). Given that CD39, along with other enzymes, degrades ATP, ADP, and AMP to adenosine, CD39 can be viewed as an immunological switch that shifts ATP-driven pro-inflammatory immune cell activity toward an anti-inflammatory state mediated by adenosine.
[0005] Within a neoplastic milieu, cancer and immune cells can closely interact to generate an immunosuppressive environment by releasing immunomodulatory factors, which support neoplastic growth. The expression of CD39 is increased in many solid tumors (for example, colorectal cancer, head and neck cancer, pancreatic cancer (Kunzli et al., Am J Physiol, 2006, 292: 223-230), bladder cancer, brain cancer, breast cancer, gastric cancer, hepatocellular carcinoma, lung cancer, non-small cell lung cancer (Li et al., Oncoimmunology, 2017, 6: 6), chronic lymphocytic leukemia (Pulte et al., Clin Lymphoma Myeloma Leuk, 2011, 11(4): 367-372) and lymphoma, melanoma (Dzhandzhugazyan et al., FEBS Letters, 1998, 430: 227-230), ovarian cancer, and prostate cancer, among others) suggesting this enzyme is involved in the development and progression of malignancies. Modulators of CD39 may provide potential therapies for these types of cancers.
[0006] Interactions between tumor cells and their microenvironment are important for tumorigenesis. CD39 can participate in tumor immunoescape by inhibiting the activation, clonal expansion, and homing of tumor-specific T cells, impairing tumor cell killing by effector T lymphocytes. In addition to these immunoregulatory roles, CD39 can contribute directly to the modulation of cancer cell growth, differentiation, invasion, migration, metastasis, and angiogenesis. CD39 is important for both the initiation of angiogenesis and the progression of neovascularization. CD39 on vasculature mediates the angiogenic process in mouse models of melanoma, lung, and liver malignancy.
[0007] Modulators of CD39 activity may also provide potential therapeutics for the treatment of CD39 conditions including, but not limited to, autoimmune diseases and infections. In particular, modulators of CD39 activity may provide potential therapeutics for diseases such as, for example, without limitation, Celiac disease (Cook et al., American Academy of Allergy, Asthma & Immunology, 2017, Article in Press), colitis (Longhi et al., JCI Insight. 2017, 2(9)), thrombotic disease (Marcus et al., Journal of Pharmacology and Experimental Therapeutics, 2003, 305, 1: 9-16), HIV infection (zur Wiesch et al., Journal of Virology, 2011, February: 1287-1297), HBV infection, HCV infection, and inflammatory bowel disease (Friedman et al. PNAS, 2009, 106, 39: 16788-16793) and Crohn's disease (Bai et al., J Immunol, 2014, 3366-3377).
SUMMARY
[0008] Provided herein are antibodies that selectively bind CD39. In some embodiments, the antibodies bind human CD39. In some embodiments, the antibodies also bind homologs of human CD39.
[0009] In some embodiments, the antibodies comprise at least one CDR sequence defined by a consensus sequence provided in this disclosure. In some embodiments, the antibodies comprise an illustrative CDR, V.sub.H, or V.sub.L sequence provided in this disclosure, heavy chain or light chain provided in the disclosure, or a variant thereof. In some aspects, the variant is a variant with one or more conservative amino acid substitutions.
[0010] Also provided are compositions and kits comprising the antibodies. In some embodiments, the compositions are pharmaceutical compositions. Any suitable pharmaceutical composition may be used. In some embodiments, the pharmaceutical composition is a composition for parenteral administration.
[0011] This disclosure also provides methods of using the anti-CD39 antibodies provided herein. In some embodiments, the method is a method of treatment. In some embodiments, the method is a diagnostic method. In some embodiments, the method is an analytical method. In some embodiments, the method is a method of purifying and/or quantifying CD39.
[0012] In some embodiments, the antibodies are used to treat a disease or condition. In some aspects, the disease or condition is selected from a cancer, autoimmune disease, and infection.
BRIEF DESCRIPTION OF THE DRAWINGS
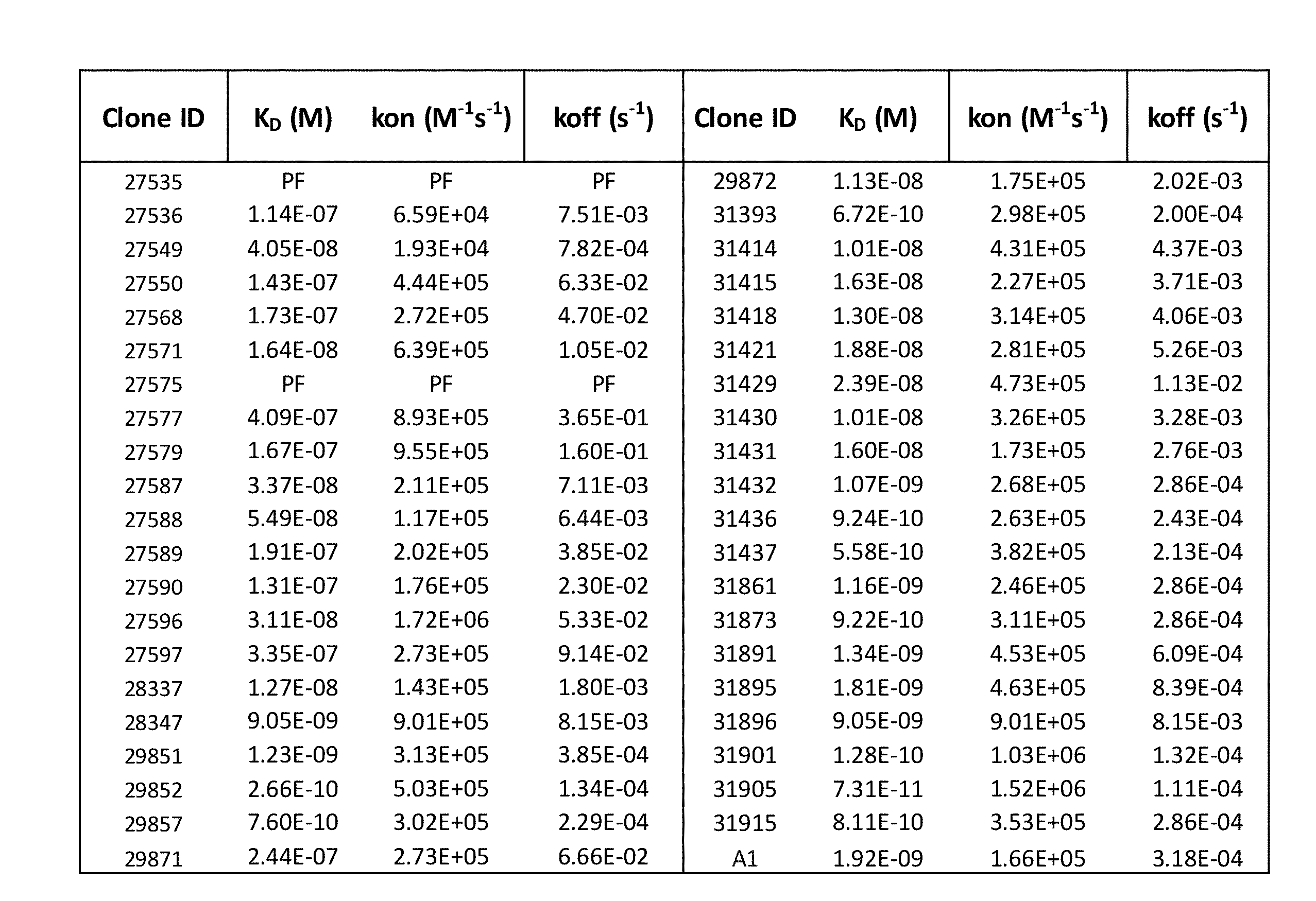
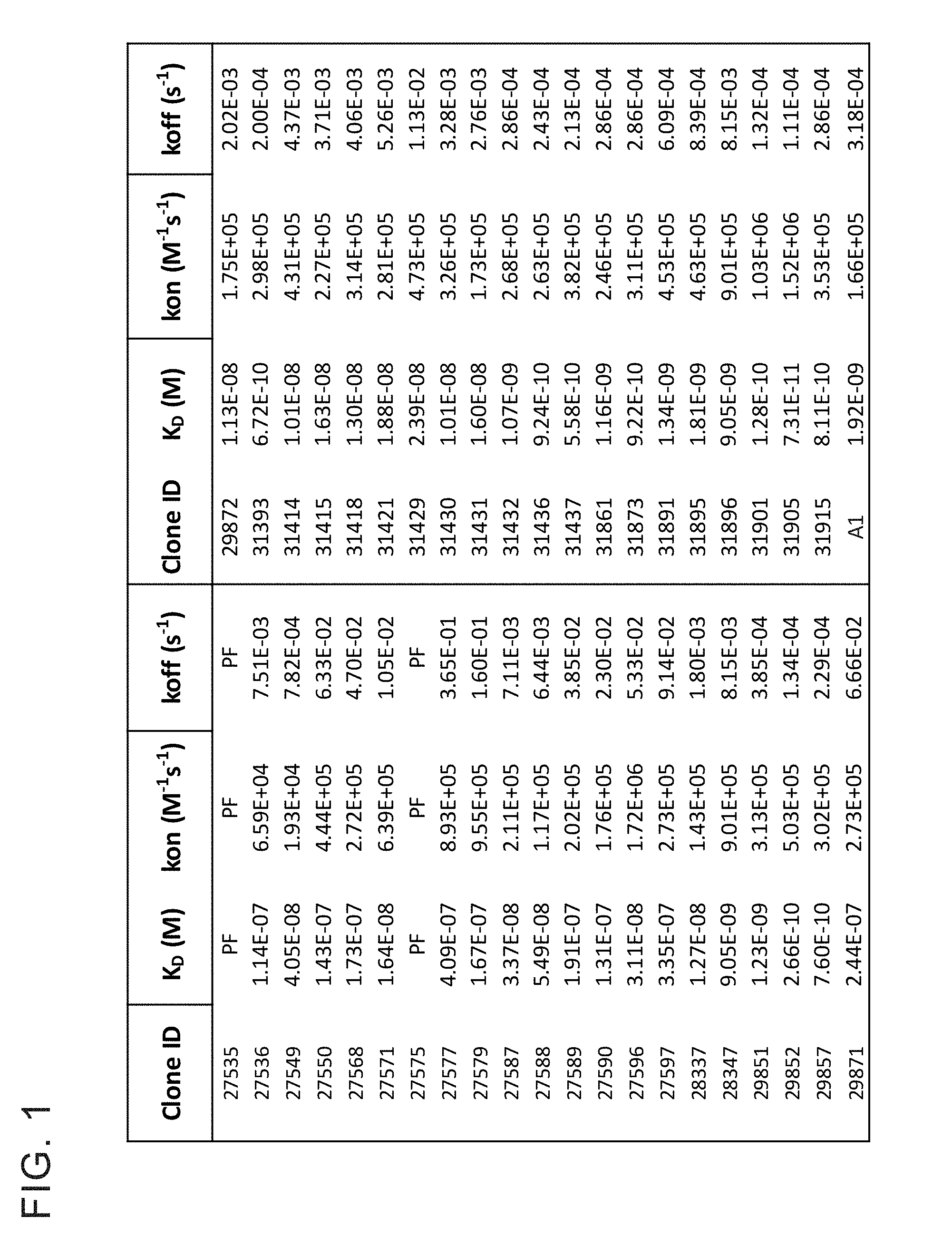
[0013] FIG. 1 provides a table showing monovalent affinity of anti-huCD39 antibodies to recombinant human CD39 extracellular domain. The table provides binding kinetics of anti-CD39 antibodies interacting with soluble recombinant human CD39 (ENTDP1) extracellular domain (ECD) by biolayer interferometry (ForteBio Octet). Anti-CD39 antibodies were captured on an anti-human Fc sensor and exposed to recombinant human CD39 ECD at concentration ranging from 10-300 nanomolar. The kinetic data was globally fit with a simple 1:1 Langmuir binding model to yield on-rate (kon) and off-rate (koff) values. The equilibrium dissociation constants (K.sub.D) were calculated from the kon and koff values.
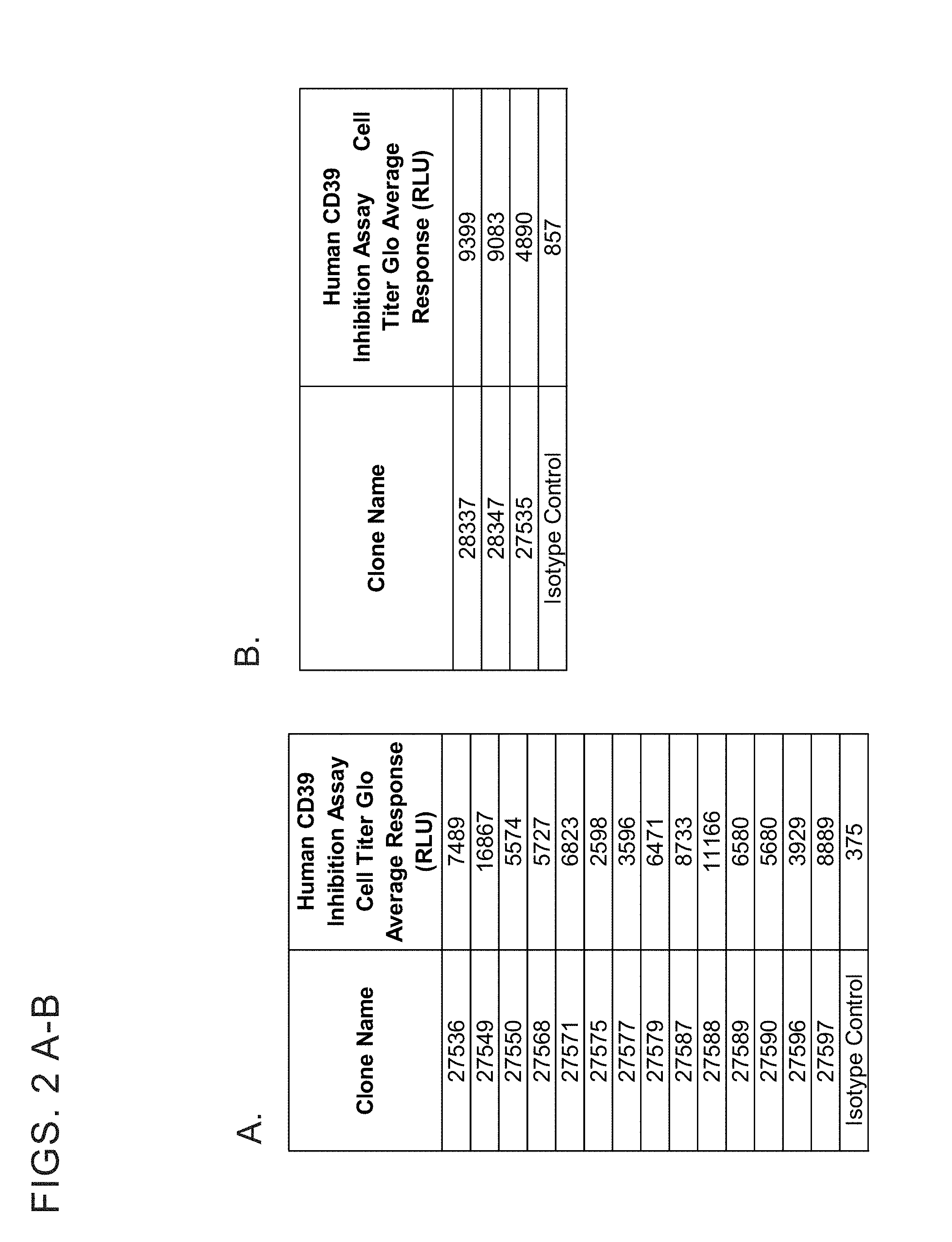
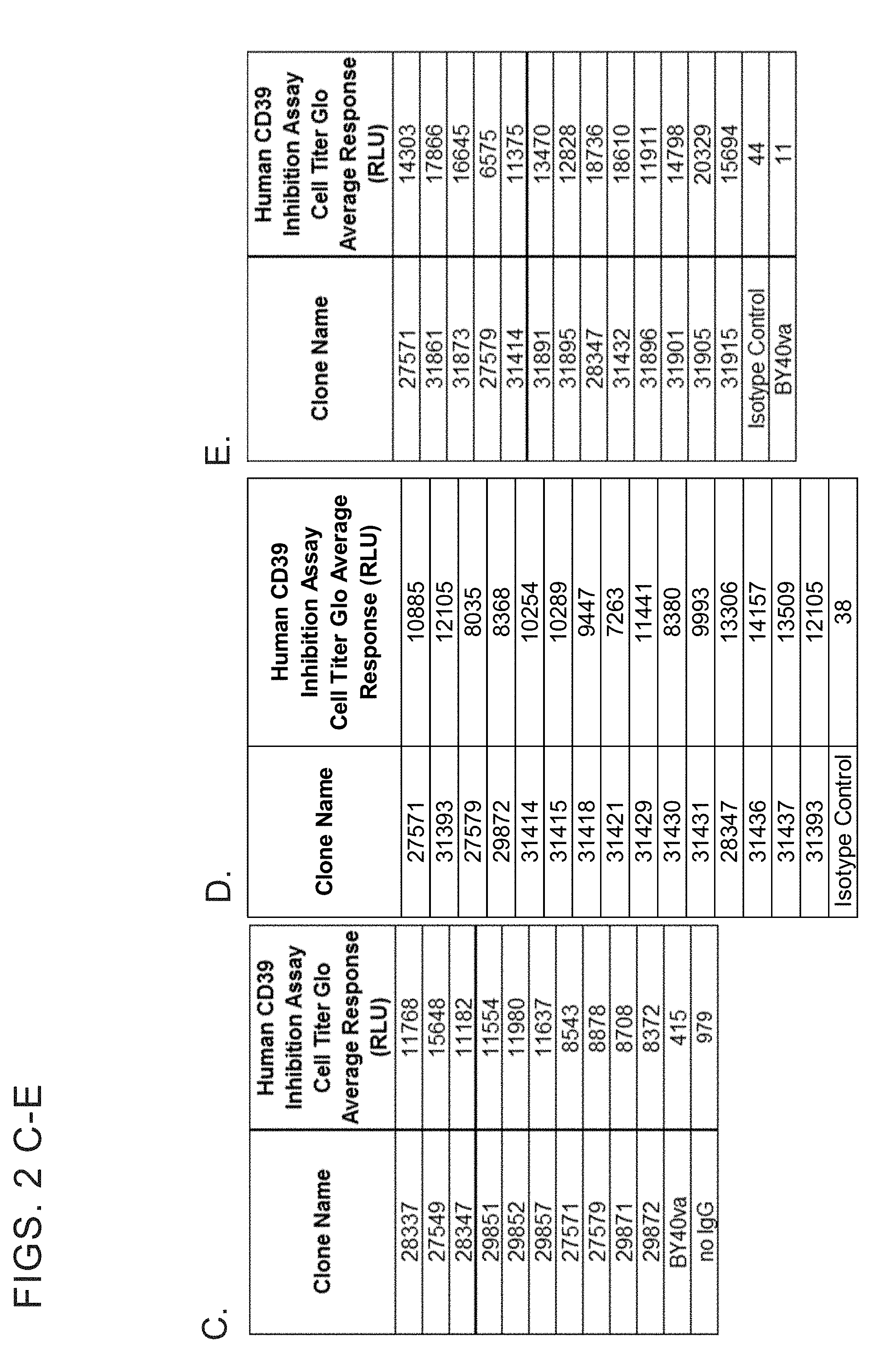
[0014] FIGS. 2 A-E show inhibition of enzymatic catabolism of ATP and ADP to Pi by human CD39 extracellular domain (ECD). Recombinant human CD39 at a final concentration of either 10 nanomolar (A-D) or 5 nanomolar (E) was incubated with anti-CD39 IgGs at a final concentration of either 1 micromolar (A-D) or 0.25 micromolar (E) in 25 mM Tris, 5 mM CaCl.sub.2), pH 7.5 at room temperature for 2 hours. ATP (500 micromolar) was added to the reaction and incubated at 37.degree. C. for 60 minutes. Residual ATP levels in the reaction were measured using the Cell Titer-Glo assay. Data values are the average of two replicates.
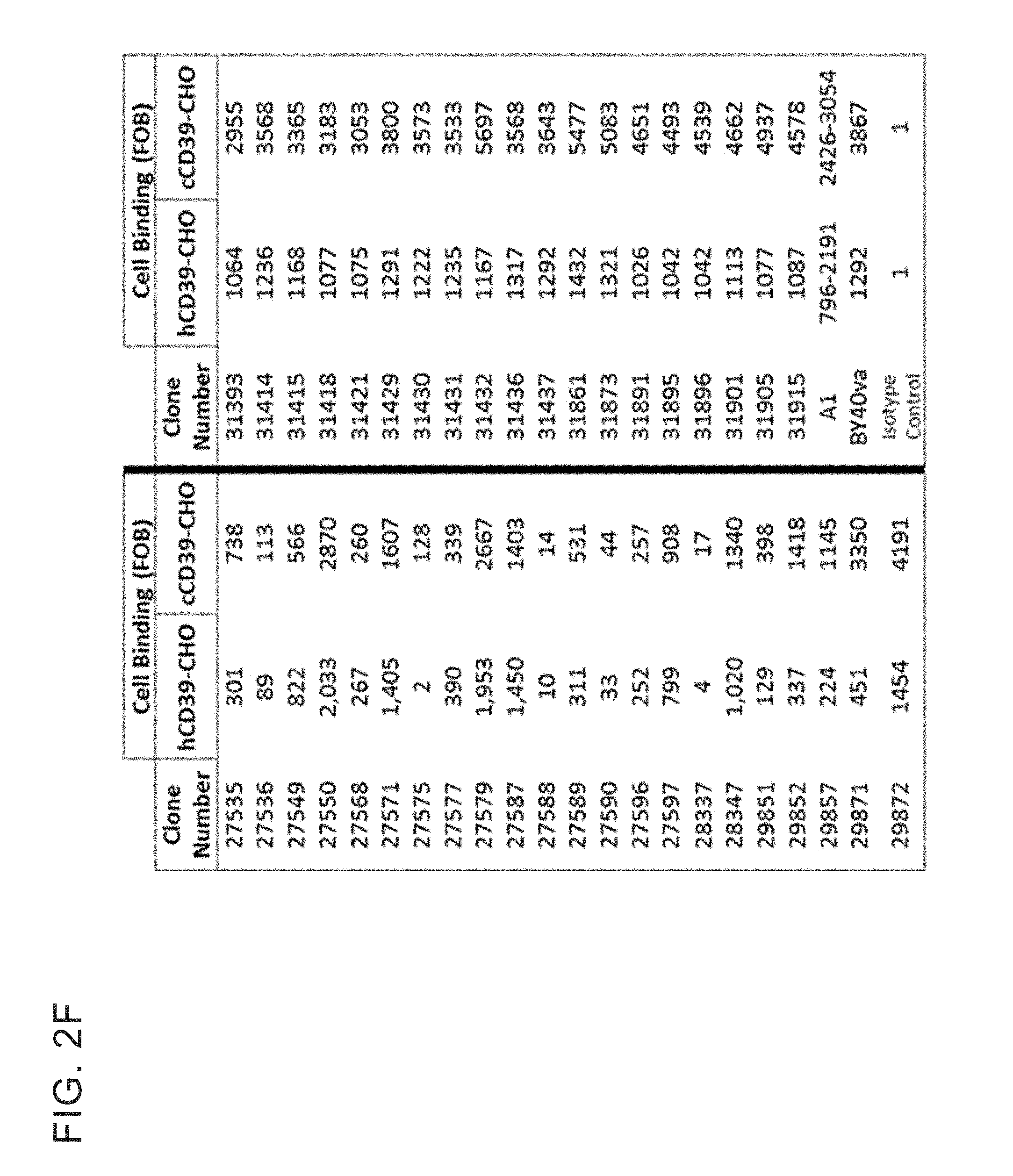
[0015] FIG. 2 F shows antibodies that bind to CHO cells expressing human or cyno CD39.
[0016] FIG. 3 shows evaluation of antibody binding to MEL-28 (A) and 721 (B) cells. Anti-CD39 antibodies (each antibody clone number indicated in the figure) were titrated from 15 to 0.001 .mu.g/ml. EC.sub.50s were calculated using GraphPad Prism Software. The figure represents three independent experiments.
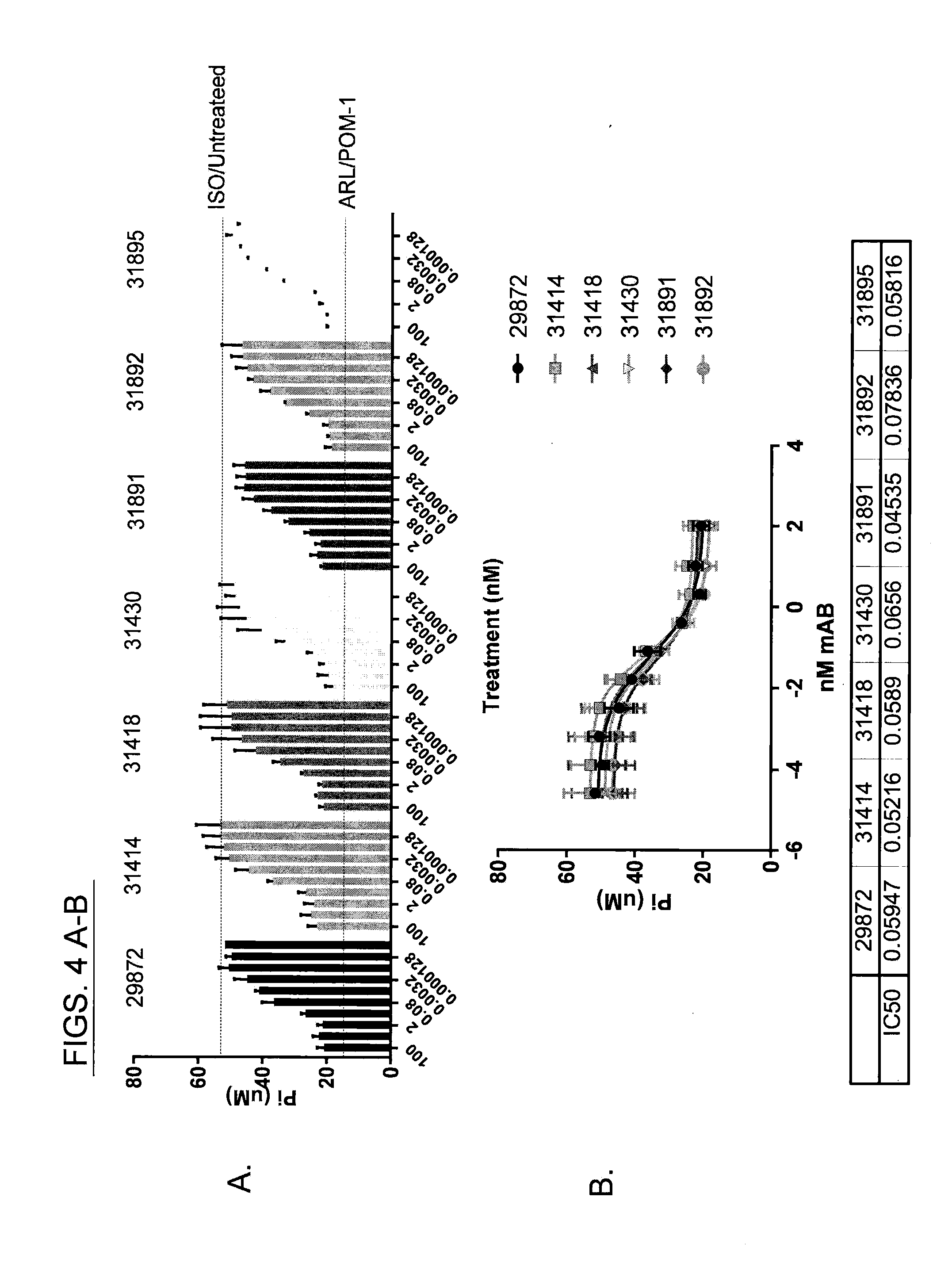
[0017] FIGS. 4 A-B provide evaluation of antibody driven inhibition of ATP hydrolysis of CD39 on MEL-28 cells in a short term ATP assay. FIG. 4A shows the results when anti-CD39 antibodies were compared to the non-specific small molecule inhibitors POM-1 and ARL. Inhibition is determined by decreased phosphate release (Pi). Data is representative of at least 10 independent experiments. Anti-CD39 antibodies (each antibody clone number is indicated in the figure) were titrated from 15 to 0.001 .mu.g/ml. IC50 values were calculated using GraphPad Prism Software, as can be seen in FIG. 4B. Three independent experiments were performed.
[0018] FIGS. 5 A-B provide an illustration of quantification when MEL-28 cells are treated with CD39 enzymatic inhibitors. FIG. 5A shows evaluation of ATP levels after incubating MEL-28 cells with a dose titration of anti-CD39 antibodies that inhibit enzymatic activity. FIG. 5B provides IC50 values calculated using GrapPad Prism. At least three independent experiments were performed.
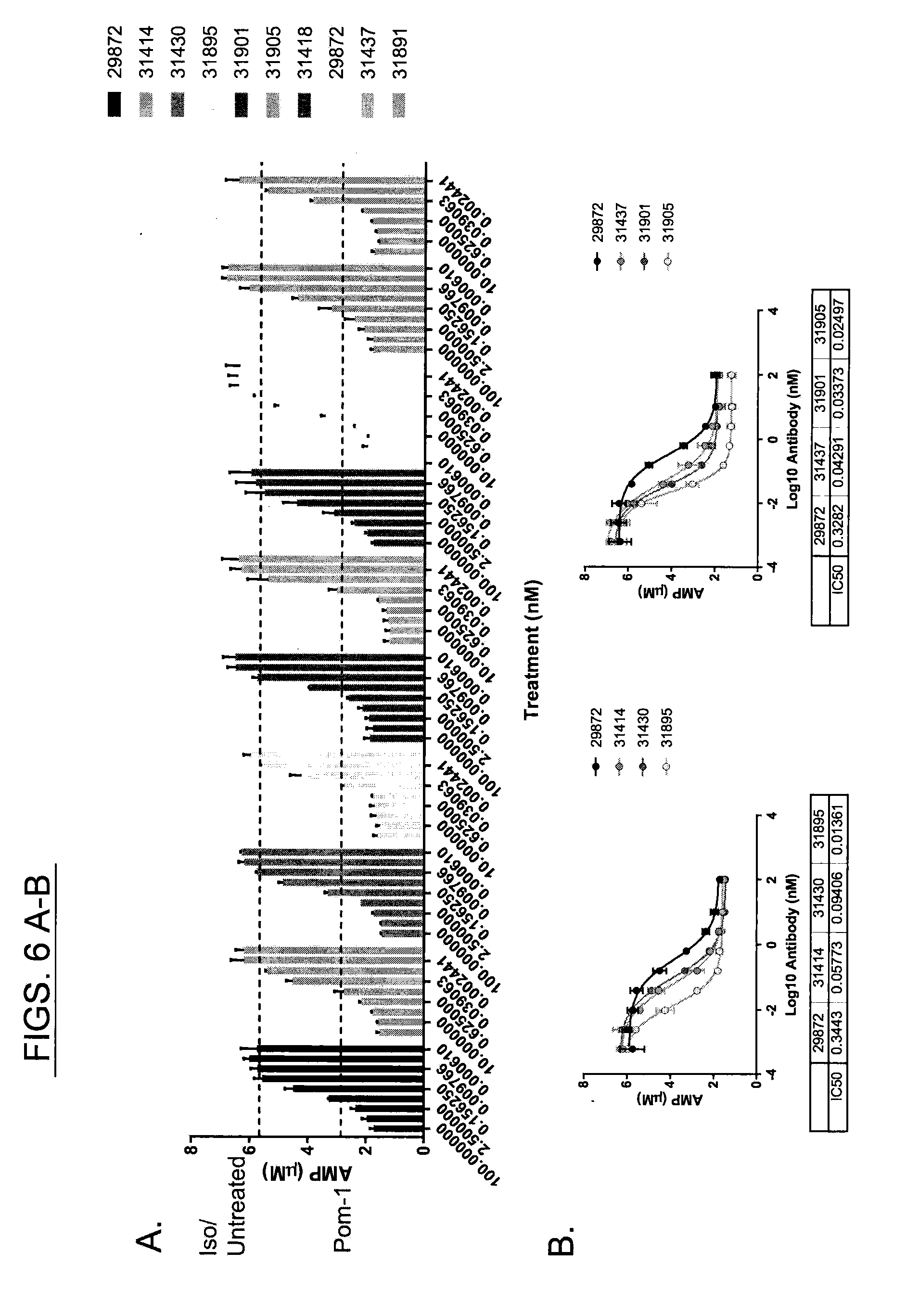
[0019] FIGS. 6 A-B show that antibodies were evaluated for inhibiting CD39 enzymatic activity overnight to MEL-28 cells. FIG. 6A shows anti-CD39 antibodies (each number represents a unique clone indicated in the figure) were titrated from 100 nM to 0.000610 nM. FIG. 6B provide IC50 values calculated using GraphPad Prism Software. Three independent experiments are represented.
[0020] FIGS. 7 A-B show results of testing of antibodies for binding to CD39 on human and cyno primary B cells. In FIG. 7A, anti-CD39 antibodies were titrated on purified B cells from healthy donor and detected with anti-human IgG-PE secondary antibody. EC50 was calculated using GrapPad Prism software. In FIG. 7B, anti-CD39 antibodies were titrated on cyno PBMCs and detected using a-human IgG PE secondary antibody. B cells were gated using FlowJo software and EC.sub.50s were calculated using GraphPad Prism.
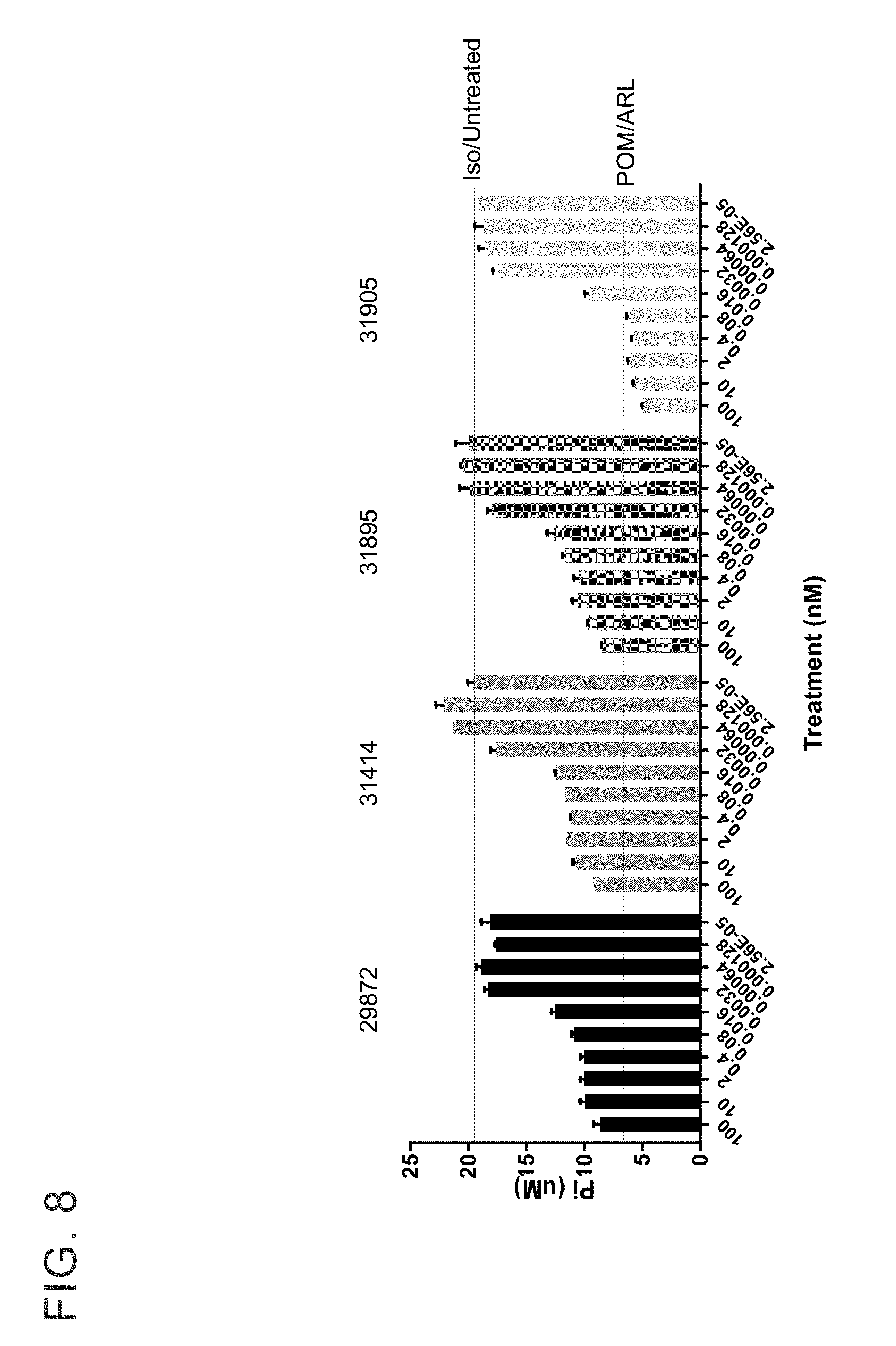
[0021] FIG. 8 sets forth antibodies that were evaluated for inhibition of CD39 activity on primary human B cells. Anti-CD39 antibodies (each unique antibody clone number indicated is in the figure) were titrated from 100 to 0.00013 nM.
[0022] FIG. 9 provides evaluation of inhibition of CD39 activity on primary human (a) and cyno (b) monocytes. Anti-CD39 antibodies (each number represents a unique antibody clone number as indicated in the figure) were titrated from 100 to 0.00013 nM and incubated with monocytes in presence of ATP. Phosphate release by CD39 processing of ATP was quantified using Malachite Green assay.
[0023] FIG. 10 shows binding of anti-CD39 antibodies on purified human CD4.sup.+CD25.sup.+CD127.sup.dim Treg cells by FACS. Anti-CD39 antibodies were titrated on purified Treg from healthy donor and detected with anti-human IgG secondary antibody. EC.sub.50s were calculated using GraphPad Prism software.
[0024] FIG. 11 shows results for antibody ability to inhibit primary Treg CD39 activity. CD24.sup.+CD25.sup.+CD127.sup.dim m T regulatory cells were incubated with serially diluted anti-CD39 antibodies and tested for ATPase activity after addition of exogenous ATP. Free phosphate (Pi) was used as a readout of CD39 activity.
[0025] FIG. 12 shows treatment with anti-CD39 antibodies increase the percent of IFN gamma producing CD8.sup.+ T cells that respond to CMV peptides in an antigen recall response assay.
[0026] FIGS. 13 A-B show evaluation of antibodies for inhibition of CD39 activity on MEL-28 (FIG. 13A) and human monocytes (FIG. 13B) as compared to the anti-CD39 antibodies generated based on Innate/Orega (BY-40v9) and Igenica (9-8B) and variants thereof. Anti-CD39 antibodies were titrated as indicated and incubated in presence of ATP. Phosphate release by CD39 processing of ATP was quantified using Malachite Green assay.
[0027] FIGS. 14 A-G provides examples of antibodies. FIG. 14A provides examples of antibodies that bind soluble recombinant CD39 ECD and cellular CD39 but do not inhibit ATPase activity and do not compete with cellular inhibitors for binding to ECD inhibitors. FIG. 14B provides examples of antibodies that have limited ability to inhibit the ATPase activity of both soluble recombinant and cellular CD39 and bin separately from other cellular CD39 inhibitors. FIG. 14C provides example of antibodies that inhibit the ATPase activity of soluble recombinant CD39 ECD but do not inhibit cellular CD39 and bin separately from other CD39 ECD inhibitors. FIG. 14D provides examples of antibodies that inhibit the ATPase activity of ECD and cellular CD39 and bin separately from other CD39 ECD and/or cellular inhibitors. FIG. 14E provides examples of inhibitory antibodies that make distinct contacts with CD39. FIG. 14E has two tables, Table 1 and Table 2. Table 1 provides examples of inhibitory antibodies that make distinct contacts with CD39. Table 2 provides examples of inhibitory antibodies that bind critical yet distinct contacts residues with CD39. FIG. 14F and FIG. 14G provide FACS plotting highlighting the importance of certain human CD39 residues.
[0028] FIG. 15 shows that anti-CD39 antibodies inhibit CD39 by 75-90%. Anti-CD39 antibodies (100 nanomolar), isotype control antibody (100 nanomolar), or ARL (200 micromolar) were incubated with MEL-28 cells endogenously expressing human CD39 for 2 hours. ATP was then added and the rate of ATP hydrolysis to Pi by CD39 was monitored using the EnzChek kinetic Pi detection assay. The initial enzyme velocity, .nu.0, was determined from the linear region of Pi vs. time curve over the first 15 minutes post-ATP addition. Each value is the mean of 3 replicates.
[0029] FIGS. 16A-B shows that the CD39 inhibitor 29872 is not a competitive inhibitor due to suppression of V.sub.max suppression.
[0030] FIG. 17 shows that anti-CD39 antibodies can induce internalization of CD39 on cyno monocytes.
[0031] FIG. 18 provides a comparison of the Kabat and Chothia numbering systems for CDR-H1. Adapted from Martin A. C. R. (2010). Protein Sequence and Structure Analysis of Antibody Variable Domains. In R. Kontermann & S. Dubel (Eds.), Antibody Engineering vol. 2 (pp. 33-51). Springer-Verlag, Berlin Heidelberg.
[0032] FIG. 19 shows an anti-CD39 antibody increases proliferation of stimulated CD4.sup.+ and CD8.sup.+ T cells.
[0033] FIG. 20 shows anti-CD39 antibody increases stimulated PBMC secretion of INF-.gamma., TNF-.alpha. and IL-2.
[0034] FIG. 21 shows anti-CD39 antibody increases stimulated PBMC secretion of INF-.gamma., TNF-.alpha., IL-2 and IL-1.beta..
[0035] FIG. 22 shows CD39 inhibition leads to accumulation of ATP and blocks generation of adenosine.
DETAILED DESCRIPTION
1. Definitions
[0036] Unless otherwise defined, all terms of art, notations and other scientific terminology used herein are intended to have the meanings commonly understood by those of skill in the art to which this invention pertains. In some cases, terms with commonly understood meanings are defined herein for clarity and/or for ready reference, and the inclusion of such definitions herein should not necessarily be construed to represent a difference over what is generally understood in the art. The techniques and procedures described or referenced herein are generally well understood and commonly employed using conventional methodologies by those skilled in the art, such as, for example, the widely utilized molecular cloning methodologies described in Sambrook et al., Molecular Cloning: A Laboratory Manual 2nd ed. (1989) Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. As appropriate, procedures involving the use of commercially available kits and reagents are generally carried out in accordance with manufacturer defined protocols and/or parameters unless otherwise noted.
[0037] As used herein, the singular forms "a," "an," and "the" include the plural referents unless the context clearly indicates otherwise.
[0038] The term "about" indicates and encompasses an indicated value and a range above and below that value. In certain embodiments, the term "about" indicates the designated value .+-.10%, .+-.5%, or .+-.1%. In certain embodiments, the term "about" indicates the designated value .+-.one standard deviation of that value.
[0039] The term "combinations thereof" includes every possible combination of elements to which the term refers.
[0040] The terms "CD39" and "CD39 antigen" and "Cluster of Differentiation 39" are used interchangeably herein. CD39 is also known as also known as ectonucleoside triphosphate diphosphohydrolase-1 (gene: ENTPD1; protein: NTPDase1, See www.ncbi.nlm.nih.gov/gene/953). CD39 has also been referred to as ATPDase and SPG64. Each of the terms set forth may be used interchangeably. Unless specified otherwise, the terms include any variants, isoforms and species homologs of human CD39 that are naturally expressed by cells, or that are expressed by cells transfected with a CD39 gene. In some embodiments, CD39 proteins include murine CD39. In some embodiments, CD39 proteins include cynomolgus CD39.
[0041] The term "immunoglobulin" refers to a class of structurally related proteins generally comprising two pairs of polypeptide chains: one pair of light (L) chains and one pair of heavy (H) chains. In an "intact immunoglobulin," all four of these chains are interconnected by disulfide bonds. The structure of immunoglobulins has been well characterized. See, e.g., Paul, Fundamental Immunology 7th ed., Ch. 5 (2013) Lippincott Williams & Wilkins, Philadelphia, Pa. Briefly, each heavy chain typically comprises a heavy chain variable region (V.sub.H) and a heavy chain constant region (C.sub.H). The heavy chain constant region typically comprises three domains, C.sub.H1, C.sub.H2, and C.sub.H3. Each light chain typically comprises a light chain variable region (V.sub.L) and a light chain constant region. The light chain constant region typically comprises one domain, abbreviated C.sub.L.
[0042] The term "antibody" describes a type of immunoglobulin molecule and is used herein in its broadest sense. An antibody specifically includes intact antibodies (e.g., intact immunoglobulins), and antibody fragments. Antibodies comprise at least one antigen-binding domain. One example of an antigen-binding domain is an antigen binding domain formed by a V.sub.H-V.sub.L dimer. A "CD39 antibody," "anti-CD39 antibody," "CD39 Ab," "CD39-specific antibody" or "anti-CD39 Ab" is an antibody, as described herein, which binds specifically to the antigen CD39. In some embodiments, the antibody binds the extracellular domain of CD39.
[0043] The V.sub.H and V.sub.L regions may be further subdivided into regions of hypervariability ("hypervariable regions (HVRs);" also called "complementarity determining regions" (CDRs)) interspersed with regions that are more conserved. The more conserved regions are called framework regions (FRs). Each V.sub.H and V.sub.L generally comprises three CDRs and four FRs, arranged in the following order (from N-terminus to C-terminus): FR1-CDR1-FR2-CDR2-FR3-CDR3-FR4. The CDRs are involved in antigen binding, and confer antigen specificity and binding affinity to the antibody. See Kabat et al., Sequences of Proteins of Immunological Interest 5th ed. (1991) Public Health Service, National Institutes of Health, Bethesda, Md., incorporated by reference in its entirety.
[0044] The light chain from any vertebrate species can be assigned to one of two types, called kappa and lambda, based on the sequence of the constant domain.
[0045] The heavy chain from any vertebrate species can be assigned to one of five different classes (or isotypes): IgA, IgD, IgE, IgG, and IgM. These classes are also designated .alpha., .delta., .epsilon., .gamma., and .mu., respectively. The IgG and IgA classes are further divided into subclasses on the basis of differences in sequence and function. Humans express the following subclasses: IgG1, IgG2, IgG3, IgG4, IgA1, and IgA2.
[0046] The amino acid sequence boundaries of a CDR can be determined by one of skill in the art using any of a number of known numbering schemes, including those described by Kabat et al., supra ("Kabat" numbering scheme); Al-Lazikani et al., 1997, J. Mol. Biol., 273:927-948 ("Chothia" numbering scheme); MacCallum et al., 1996, J. Mol. Biol. 262:732-745 ("Contact" numbering scheme); Lefranc et al., Dev. Comp. Immunol., 2003, 27:55-77 ("IMGT" numbering scheme); and Honegge and Pluckthun, J. Mol. Biol., 2001, 309:657-70 ("AHo" numbering scheme), each of which is incorporated by reference in its entirety.
[0047] Table 1 provides the positions of CDR-L1, CDR-L2, CDR-L3, CDR-H1, CDR-H2, and CDR-H3 as identified by the Kabat and Chothia schemes. For CDR-H1, residue numbering is provided using both the Kabat and Chothia numbering schemes. FIG. 1 provides a comparison of the Kabat and Chothia numbering schemes for CDR-H1. See Martin (2010), supra.
[0048] Unless otherwise specified, the numbering scheme used for identification of a particular CDR herein is the Kabat/Chothia numbering scheme. Where the residues encompassed by these two numbering schemes diverge, the numbering scheme is specified as either Kabat or Chothia.
TABLE-US-00001 TABLE 1 Residues in CDRs according to Kabat and Chothia numbering schemes. CDR Kabat Chothia L1 L24-L34 L24-L34 L2 L50-L56 L50-L56 L3 L89-L97 L89-L97 H1 (Kabat Numbering) H31-H35B H26-H32 or H34* H1 (Chothia Numbering) H31-H35 H26-H32 H2 H50-H65 H52-H56 H3 H95-H102 H95-H102 *The C-terminus of CDR-H1, when numbered using the Kabat numbering convention, varies between H32 and H34, depending on the length of the CDR, as illustrated in FIG. 1.
[0049] The "EU numbering scheme" is generally used when referring to a residue in an antibody heavy chain constant region (e.g., as reported in Kabat et al., supra). Unless stated otherwise, the EU numbering scheme is used to refer to residues in antibody heavy chain constant regions described herein.
[0050] An "antibody fragment" comprises a portion of an intact antibody, such as the antigen binding or variable region of an intact antibody. Antibody fragments include, for example, Fv fragments, Fab fragments, F(ab')2 fragments, Fab' fragments, scFv (sFv) fragments, and scFv-Fc fragments.
[0051] "Fv" fragments comprise a non-covalently-linked dimer of one heavy chain variable domain and one light chain variable domain.
[0052] "Fab" fragments comprise, in addition to the heavy and light chain variable domains, the constant domain of the light chain and the first constant domain (C.sub.H1) of the heavy chain. Fab fragments may be generated, for example, by papain digestion of a full-length antibody.
[0053] "F(ab')2" fragments contain two Fab' fragments joined, near the hinge region, by disulfide bonds. F(ab')2 fragments may be generated, for example, by pepsin digestion of an intact antibody. The F(ab') fragments can be dissociated, for example, by treatment with .beta.-mercaptoethanol.
[0054] "Single-chain Fv" or "sFv" or "scFv" antibody fragments comprise a V.sub.H domain and a V.sub.L domain in a single polypeptide chain. The V.sub.H and V.sub.L are generally linked by a peptide linker. See Pluckthun A. (1994). Antibodies from Escherichia coli. In Rosenberg M. & Moore G. P. (Eds.), The Pharmacology of Monoclonal Antibodies vol. 113 (pp. 269-315). Springer-Verlag, New York, incorporated by reference in its entirety. "scFv-Fc" fragments comprise an scFv attached to an Fc domain. For example, an Fc domain may be attached to the C-terminal of the scFv. The Fc domain may follow the V.sub.H or V.sub.L, depending on the orientation of the variable domains in the scFv (i.e., V.sub.H-V.sub.L or V.sub.L-V.sub.H). Any suitable Fc domain known in the art or described herein may be used.
[0055] The term "monoclonal antibody" refers to an antibody from a population of substantially homogeneous antibodies. A population of substantially homogeneous antibodies comprises antibodies that are substantially similar and that bind the same epitope(s), except for variants that may normally arise during production of the monoclonal antibody. Such variants are generally present in only minor amounts. A monoclonal antibody is typically obtained by a process that includes the selection of a single antibody from a plurality of antibodies. For example, the selection process can be the selection of a unique clone from a plurality of clones, such as a pool of hybridoma clones, phage clones, yeast clones, bacterial clones, or other recombinant DNA clones. The selected antibody can be further altered, for example, to improve affinity for the target ("affinity maturation"), to humanize the antibody, to improve its production in cell culture, and/or to reduce its immunogenicity in a subject.
[0056] The term "chimeric antibody" refers to an antibody in which a portion of the heavy and/or light chain is derived from a particular source or species, while the remainder of the heavy and/or light chain is derived from a different source or species.
[0057] "Humanized" forms of non-human antibodies are chimeric antibodies that contain minimal sequence derived from the non-human antibody. A humanized antibody is generally a human immunoglobulin (recipient antibody) in which residues from one or more CDRs are replaced by residues from one or more CDRs of a non-human antibody (donor antibody). The donor antibody can be any suitable non-human antibody, such as a mouse, rat, rabbit, chicken, or non-human primate antibody having a desired specificity, affinity, or biological effect. In some instances, selected framework region residues of the recipient antibody are replaced by the corresponding framework region residues from the donor antibody. Humanized antibodies may also comprise residues that are not found in either the recipient antibody or the donor antibody. Such modifications may be made to further refine antibody function. For further details, see Jones et al., Nature, 1986, 321:522-525; Riechmann et al., Nature, 1988, 332:323-329; and Presta, Curr. Op. Struct. Biol., 1992, 2:593-596, each of which is incorporated by reference in its entirety.
[0058] A "human antibody" is one which possesses an amino acid sequence corresponding to that of an antibody produced by a human or a human cell, or derived from a non-human source that utilizes a human antibody repertoire or human antibody-encoding sequences (e.g., obtained from human sources or designed de novo). Human antibodies specifically exclude humanized antibodies.
[0059] An "isolated antibody" is one that has been separated and/or recovered from a component of its natural environment. Components of the natural environment may include enzymes, hormones, and other proteinaceous or nonproteinaceous materials. In some embodiments, an isolated antibody is purified to a degree sufficient to obtain at least 15 residues of N-terminal or internal amino acid sequence, for example by use of a spinning cup sequenator. In some embodiments, an isolated antibody is purified to homogeneity by gel electrophoresis (e.g., SDS-PAGE) under reducing or nonreducing conditions, with detection by Coomassie blue or silver stain. An isolated antibody includes an antibody in situ within recombinant cells, since at least one component of the antibody's natural environment is not present. In some aspects, an isolated antibody is prepared by at least one purification step.
[0060] In some embodiments, an isolated antibody is purified to at least 80%, 85%, 90%, 95%, or 99% by weight. In some embodiments, an isolated antibody is provided as a solution comprising at least 85%, 90%, 95%, 98%, 99% to 100% by weight of an antibody, the remainder of the weight comprising the weight of other solutes dissolved in the solvent.
[0061] "Affinity" refers to the strength of the sum total of non-covalent interactions between a single binding site of a molecule (e.g., an antibody) and its binding partner (e.g., an antigen). Unless indicated otherwise, as used herein, "binding affinity" refers to intrinsic binding affinity, which reflects a 1:1 interaction between members of a binding pair (e.g., antibody and antigen). The affinity of a molecule X for its partner Y can generally be represented by the dissociation constant (K.sub.D). Affinity can be measured by common methods known in the art, including those described herein. Affinity can be determined, for example, using surface plasmon resonance (SPR) technology, such as a Biacore.RTM. instrument.
[0062] With regard to the binding of an antibody to a target molecule, the terms "specific binding," "specifically binds to," "specific for," "selectively binds," and "selective for" a particular antigen (e.g., a polypeptide target) or an epitope on a particular antigen mean binding that is measurably different from a non-specific or non-selective interaction. Specific binding can be measured, for example, by determining binding of a molecule compared to binding of a control molecule. Specific binding can also be determined by competition with a control molecule that is similar to the target, such as an excess of non-labeled target. In that case, specific binding is indicated if the binding of the labeled target to a probe is competitively inhibited by the excess non-labeled target.
[0063] The term "k.sub.d" (sec.sup.-1), as used herein, refers to the dissociation rate constant of a particular antibody-antigen interaction. This value is also referred to as the k.sub.off value.
[0064] The term "k.sub.a" (M.sup.-1.times.sec.sup.-1), as used herein, refers to the association rate constant of a particular antibody-antigen interaction. This value is also referred to as the k.sub.on value.
[0065] The term "K.sub.D" (M), as used herein, refers to the dissociation equilibrium constant of a particular antibody-antigen interaction. K.sub.D=k.sub.d/k.sub.a.
[0066] The term "K.sub.A" (M.sup.-1), as used herein, refers to the association equilibrium constant of a particular antibody-antigen interaction. K.sub.A=k.sub.a/k.sub.d.
[0067] An "affinity matured" antibody is one with one or more alterations in one or more CDRs or FRs that result in an improvement in the affinity of the antibody for its antigen, compared to a parent antibody which does not possess the alteration(s). In one embodiment, an affinity matured antibody has nanomolar or picomolar affinity for the target antigen. Affinity matured antibodies may be produced using a variety of methods known in the art. For example, Marks et al. (Bio/Technology, 1992, 10:779-783, incorporated by reference in its entirety) describes affinity maturation by V.sub.H and V.sub.L domain shuffling. Random mutagenesis of CDR and/or framework residues is described by, for example, Barbas et al. (Proc. Nat. Acad. Sci. U.S.A., 1994, 91:3809-3813); Schier et al., Gene, 1995, 169:147-155; Yelton et al., J. Immunol., 1995, 155:1994-2004; Jackson et al., J. Immunol., 1995, 154:3310-33199; and Hawkins et al, J. Mol. Biol., 1992, 226:889-896, each of which is incorporated by reference in its entirety.
[0068] When used herein in the context of two or more antibodies, the term "competes with" or "cross-competes with" indicates that the two or more antibodies compete for binding to an antigen (e.g., CD39). In one exemplary assay, CD39 is coated on a plate and allowed to bind a first antibody, after which a second, labeled antibody is added. If the presence of the first antibody reduces binding of the second antibody, then the antibodies compete. The term "competes with" also includes combinations of antibodies where one antibody reduces binding of another antibody, but where no competition is observed when the antibodies are added in the reverse order. However, in some embodiments, the first and second antibodies inhibit binding of each other, regardless of the order in which they are added. In some embodiments, one antibody reduces binding of another antibody to its antigen by at least 50%, at least 60%, at least 70%, at least 80%, or at least 90%.
[0069] The term "epitope" means a portion of an antigen capable of specific binding to an antibody. Epitopes frequently consist of surface-accessible amino acid residues and/or sugar side chains and may have specific three dimensional structural characteristics, as well as specific charge characteristics. Conformational and non-conformational epitopes are distinguished in that the binding to the former but not the latter is lost in the presence of denaturing solvents. An epitope may comprise amino acid residues that are directly involved in the binding, and other amino acid residues, which are not directly involved in the binding. The epitope to which an antibody binds can be determined using known techniques for epitope determination such as, for example, testing for antibody binding to CD39 variants with different point-mutations.
[0070] Percent "identity" between a polypeptide sequence and a reference sequence is defined as the percentage of amino acid residues in the polypeptide sequence that are identical to the amino acid residues in the reference sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN, MEGALIGN (DNASTAR), CLUSTALW, or CLUSTAL OMEGA software. Those skilled in the art can determine appropriate parameters for aligning sequences, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared.
[0071] A "conservative substitution" or a "conservative amino acid substitution," refers to the substitution of one or more amino acids with one or more chemically or functionally similar amino acids. Conservative substitution tables providing similar amino acids are well known in the art. Polypeptide sequences having such substitutions are known as "conservatively modified variants." Such conservatively modified variants are in addition to and do not exclude polymorphic variants, interspecies homologs, and alleles. By way of example, the following groups of amino acids are considered conservative substitutions for one another.
TABLE-US-00002 Acidic Residues D and E Basic Residues K, R, and H Hydrophilic Uncharged Residues S, T, N, and Q Aliphatic Uncharged Residues G, A, V, L, and I Non-polar Uncharged Residues C, M, and P Aromatic Residues F, Y, and W Alcohol Group-Containing Residues S and T Aliphatic Residues I, L, V, and M Cycloalkenyl-associated Residues F, H, W, and Y Hydrophobic Residues A, C, F, G, H, I, L, M, R, T, V, W, and Y Negatively Charged Residues D and E Polar Residues C, D, E, H, K, N, Q, R, S, and T Positively Charged Residues H, K, and R Small Residues A, C, D, G, N, P, S, T, and V Very Small Residues A, G, and S Residues Involved in Turn Formation A, C, D, E, G, H, K, N, Q, R, S, P, and T Flexible Residues Q, T, K, S, G, P, D, E, and R Group 1 A, S, and T Group 2 D and E Group 3 N and Q Group 4 R and K Group 5 I, L, and M Group 6 F, Y, and W Group A A and G Group B D and E Group C N and Q Group D R, K, and H Group E I, L, M, V Group F F, Y, and W Group G S and T Group H C and M
Additional conservative substitutions may be found, for example, in Creighton, Proteins: Structures and Molecular Properties 2nd ed. (1993) W. H. Freeman & Co., New York, N.Y. An antibody generated by making one or more conservative substitutions of amino acid residues in a parent antibody is referred to as a "conservatively modified variant."
[0072] The term "amino acid" refers to the twenty common naturally occurring amino acids. Naturally occurring amino acids include alanine (Ala; A), arginine (Arg; R), asparagine (Asn; N), aspartic acid (Asp; D), cysteine (Cys; C); glutamic acid (Glu; E), glutamine (Gln; Q), Glycine (Gly; G); histidine (His; H), isoleucine (Ile; I), leucine (Leu; L), lysine (Lys; K), methionine (Met; M), phenylalanine (Phe; F), proline (Pro; P), serine (Ser; S), threonine (Thr; T), tryptophan (Trp; W), tyrosine (Tyr; Y), and valine (Val; V).
[0073] "Treating" or "treatment" of any disease or disorder refers, in certain embodiments, to ameliorating a disease or disorder that exists in a subject. In another embodiment, "treating" or "treatment" includes ameliorating at least one physical parameter, which may be indiscernible by the subject. In yet another embodiment, "treating" or "treatment" includes modulating the disease or disorder, either physically (e.g., stabilization of a discernible symptom) or physiologically (e.g., stabilization of a physical parameter) or both. In yet another embodiment, "treating" or "treatment" includes delaying or preventing the onset of the disease or disorder.
[0074] As used herein, the term "therapeutically effective amount" or "effective amount" refers to an amount of an antibody or composition that when administered to a subject is effective to treat a disease or disorder.
[0075] As used herein, the term "subject" means a mammalian subject. Exemplary subjects include, but are not limited to humans, monkeys, dogs, cats, mice, rats, cows, horses, camels, avians, goats, and sheep. In certain embodiments, the subject is a human. In some embodiments, the subject has cancer, an autoimmune disease or condition, and/or an infection that can be treated with an antibody provided herein. In some embodiments, the subject is a human that is suspected to have cancer, an autoimmune disease or condition, and/or an infection.
2. Antibodies
[0076] Provided herein are antibodies that selectively bind human CD39, as well as the nucleic acids that encode the antibodies. In some aspects, the antibody selectively binds to the extracellular domain of human CD39.
[0077] In some embodiments, the antibody binds to homologs of human CD39. In some aspects, the antibody binds to a homolog of human CD39 from a species selected from monkeys, mice, dogs, cats, rats, cows, horses, goats, and sheep. In some aspects, the homolog is a cynomolgus monkey homolog. In some aspects, the homolog is a murine homolog.
[0078] In some embodiments, the antibody has one or more CDRs having particular lengths, in terms of the number of amino acid residues. In some embodiments, the Chothia CDR-H1 of the antibody is 6, 7, 8, or 9 residues in length. In some embodiments, the Kabat CDR-H1 of the antibody is 4, 5, 6, or 7 residues in length. In some embodiments, the Chothia CDR-H2 of the antibody is 5, 6, or 7 residues in length. In some embodiments, the Kabat CDR-H2 of the antibody is 15, 16, 17, or 18 residues in length. In some embodiments, the Kabat/Chothia CDR-H3 of the antibody is 5, 6, 7, 8, 9, 10, 11, or 12 residues in length.
[0079] In some aspects, the Kabat/Chothia CDR-L1 of the antibody is 9, 10, 11, 12, 13, 14, 15, or 16 residues in length. In some aspects, the Kabat/Chothia CDR-L2 of the antibody is 6, 7, or 8 residues in length. In some aspects, the Kabat/Chothia CDR-L3 of the antibody is 8, 9, 10, 11, or 12 residues in length.
[0080] In some embodiments, the antibody comprises a light chain. In some aspects, the light chain is a kappa light chain. In some aspects, the light chain is a lambda light chain.
[0081] In some embodiments, the antibody comprises a heavy chain. In some aspects, the heavy chain is an IgA. In some aspects, the heavy chain is an IgD. In some aspects, the heavy chain is an IgE. In some aspects, the heavy chain is an IgG. In some aspects, the heavy chain is an IgM. In some aspects, the heavy chain is an IgG1. In some aspects, the heavy chain is an IgG2. In some aspects, the heavy chain is an IgG3. In some aspects, the heavy chain is an IgG4. In some aspects, the heavy chain is an IgA1. In some aspects, the heavy chain is an IgA2.
[0082] In some embodiments, the antibody is an antibody fragment. In some aspects, the antibody fragment is an Fv fragment. In some aspects, the antibody fragment is a Fab fragment. In some aspects, the antibody fragment is a F(ab').sub.2 fragment. In some aspects, the antibody fragment is a Fab' fragment. In some aspects, the antibody fragment is an scFv (sFv) fragment. In some aspects, the antibody fragment is an scFv-Fc fragment.
[0083] In some embodiments, the antibody is a monoclonal antibody. In some embodiments, the antibody is a polyclonal antibody.
[0084] In some embodiments, the antibody is a chimeric antibody. In some embodiments, the antibody is a humanized antibody. In some embodiments, the antibody is a human antibody.
[0085] In some embodiments, the antibody is an affinity matured antibody. In some aspects, the antibody is an affinity matured antibody derived from an illustrative sequence provided in this disclosure.
[0086] In some aspects, the antibody inhibits conversion by CD39 of ATP to ADP and/or ADP to AMP. In some aspects, the antibody decreases the levels of phosphate, ADP, AMP, and/or adenosine and/or increases the levels of ATP.
[0087] In some embodiments, the antibody increases proliferation of stimulated CD4.sup.+ and CD8.sup.+ T cells. In some embodiments, the antibody increases stimulated PBMC secretion of INF-.gamma., TNF-.alpha., IL-2, and/or IL-1.beta..
[0088] In some embodiments, the antibody increases a T effector cell function. In some embodiments, the antibody decreases the number of regulatory T cells in tissues or in circulation. In some embodiments, the antibody suppresses a regulatory or T cell activity. In some embodiments, the antibody increase B cell function. In some embodiments, the antibody increases antigen presenting cell function. In some embodiments, the antibody decreases or prevents activation of phospho antigen specific T cells selected from MAIT cells and gamma delta T cells.
[0089] In some aspects, the decrease is about or less than a 10% decrease, about or less than a 20% decrease, about or less than a 30% decrease, about or less than a 40% decrease, about or less than a 50% decrease, about or less than a 60% decrease, about or less than a 70% decrease, about or less than an 80% decrease, about or less than a 90% decrease, or about a complete decrease. In some aspects, the increase is about or greater than a 10% increase, about or greater than a 20% increase, about or greater than a 30% increase, about or greater than a 40% increase, about or greater than a 50% increase, about or greater than a 60% increase, about or greater than a 70% increase, about or greater than an 80% increase, about or greater than a 90% increase, or a complete increase.
[0090] Given that CD39 degrades ATP and ADP to adenosine, CD39 can be viewed as an immunological switch that shifts ATP-driven pro-inflammatory immune cell activity toward an anti-inflammatory state mediated by adenosine. CD39 has a role in regulating the function of several immune cell types, including lymphocytes, neutrophils, monocytes/macrophages, dendritic cells, and endothelial cells and shifting the switch can have a significant impact on disease. For example, the generation of adenosine via CD39 is recognized as a major mechanism of regulatory T cell (Treg) immunosuppressive function.
[0091] The antibodies provided herein may be useful for the treatment of a variety of diseases and conditions, including cancers, autoimmune diseases, and infections. In some embodiments, the antibody inhibits CD39 function on tumor cells. In some embodiments, the antibody inhibits angiogenesis.
[0092] The frequency of CD39.sup.+ Tregs and the expression on the cell surface is increased in some human cancers, and the importance of CD39.sup.+ Tregs in promoting tumor growth and metastasis has been demonstrated using several in vivo models. Immunohistochemical staining of normal and tumor tissues has revealed that CD39 expression is significantly higher in several types of human cancer than in normal tissues. In cancer specimens, CD39 is expressed by infiltrating lymphocytes, the tumor stroma, and/or tumor cells. CD39 in cancer cells displays ATPase activity and generates adenosine. CD39+ cancer cells inhibited the proliferation of CD4 and CD8 T cells and the generation of cytotoxic effector CD8 T cells (CTL) in a CD39- and adenosine-dependent manner.
2.1. CDR-H3 Sequences
[0093] In some embodiments, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 82. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 83. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 84. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 85. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 86. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 87. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 88. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 89. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 90. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 91. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 92. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 93. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 94. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 95. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 96. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 97. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 98. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 99. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 100. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 101. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 102. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 103. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 104. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 105. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 106. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 107. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 108. In some aspects, the antibody comprises a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 109.
[0094] In some aspects, the CDR-H3 sequence comprises, consists of, or consists essentially of a variant of an illustrative CDR-H3 sequence provided in this disclosure. In some aspects, the CDR-H3 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative CDR-H3 sequences provided in this disclosure. In some aspects, the CDR-H3 sequence comprises, consists of, or consists essentially of any of the illustrative CDR-H3 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.2. V.sub.H Sequences Comprising Illustrative CDRs
[0095] In some embodiments, the antibody comprises a V.sub.H sequence comprising one or more CDR-H sequences comprising, consisting of, or consisting essentially of one or more illustrative CDR-H sequences provided in this disclosure, and variants thereof.
2.2.1. V.sub.H Sequences Comprising Illustrative Kabat CDRs
[0096] In some embodiments, the antibody comprises a V.sub.H sequence comprising one or more Kabat CDR-H sequences comprising, consisting of, or consisting essentially of one or more illustrative Kabat CDR-H sequences provided in this disclosure, and variants thereof.
2.2.1.1. Kabat CDR-H3
[0097] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 83. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 84. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 85. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 86. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 88. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 89. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 90. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 91. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 92. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 93. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 94. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 95. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 96. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 97. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 98. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 99. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 100. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 101. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 102. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 103. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 104. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 105. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 106. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 107. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 108. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 109.
2.2.1.2. Kabat CDR-H2
[0098] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 63-81. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 63. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 64. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 65. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 66. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 67. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 68. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 69. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 70. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 71. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 72. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 73. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 74. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 75. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 76. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 77. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 78. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 79. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 80. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 81.
2.2.1.3. Kabat CDR-H1
[0099] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 25-45. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 25. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 26. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 27. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 28. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 29. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 30. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 31. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 32. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 33. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 34. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 35. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 36. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 37. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 38. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 39. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 40. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 41. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 42. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 43. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 44. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 45.
2.2.1.4. Kabat CDR-H3+Kabat CDR-H2
[0100] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109, and a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 63-81. In some aspects, the Kabat CDR-H3 sequence and the Kabat CDR-H2 sequence are both from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Kabat CDR-H3 and Kabat CDR-H2 are both from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
2.2.1.5. Kabat CDR-H3+Kabat CDR-H1
[0101] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109, and a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 25-45. In some aspects, the Kabat CDR-H3 sequence and the Kabat CDR-H1 sequence are both from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Kabat CDR-H3 and Kabat CDR-H1 are both from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
2.2.1.6. Kabat CDR-H1+Kabat CDR-H2
[0102] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 25-45 and a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 63-81. In some aspects, the Kabat CDR-H1 sequence and the Kabat CDR-H2 sequence are both from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Kabat CDR-H1 and Kabat CDR-H2 are both from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
2.2.1.7. Kabat CDR-H1+Kabat CDR-H2+Kabat CDR-H3
[0103] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 25-45, a Kabat CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 63-81, and a Kabat CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109. In some aspects, the Kabat CDR-H1 sequence, Kabat CDR-H2 sequence, and Kabat CDR-H3 sequence are all from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Kabat CDR-H1, Kabat CDR-H2, and Kabat CDR-H3 are all from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
[0104] In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 25, a Kabat CDR-H2 sequence comprising SEQ ID NO: 63, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 83. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 27, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 84. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 85. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 86. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 25, a Kabat CDR-H2 sequence comprising SEQ ID NO: 66, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 27, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 28, a Kabat CDR-H2 sequence comprising SEQ ID NO: 67, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 29, a Kabat CDR-H2 sequence comprising SEQ ID NO: 68, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 31, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 88. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 32, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 89. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 90. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 34, a Kabat CDR-H2 sequence comprising SEQ ID NO: 72, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 91. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 92. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 35, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 93. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 92. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 36, a Kabat CDR-H2 sequence comprising SEQ ID NO: 72, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 73, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 37, a Kabat CDR-H2 sequence comprising SEQ ID NO: 74, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 75, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 94. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 39, a Kabat CDR-H2 sequence comprising SEQ ID NO: 76, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 95. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 40, a Kabat CDR-H2 sequence comprising SEQ ID NO: 76, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 96. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 39, a Kabat CDR-H2 sequence comprising SEQ ID NO: 76, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 94. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 97. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 41, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 98. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 41, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 99. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 41, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 100. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 75, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 101. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 42, a Kabat CDR-H2 sequence comprising SEQ ID NO: 78, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 102. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 75, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 103. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 43, a Kabat CDR-H2 sequence comprising SEQ ID NO: 79, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 104. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 103. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 106. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 107. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 44, a Kabat CDR-H2 sequence comprising SEQ ID NO: 80, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 108. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 45, a Kabat CDR-H2 sequence comprising SEQ ID NO: 81, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 109.
[0105] In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 83. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 84. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 85. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 86. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 84. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 86. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 27, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 83. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 27, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 86. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 93. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 73, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 72, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 31, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 31, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 32, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 72, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 93. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 93. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 34, a Kabat CDR-H2 sequence comprising SEQ ID NO: 72, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 35, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 35, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 90. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 39, a Kabat CDR-H2 sequence comprising SEQ ID NO: 76, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 94.
2.2.1.8. Variants of V.sub.H Sequences Comprising Illustrative Kabat CDRs
[0106] In some embodiments, the V.sub.H sequences provided herein comprise a variant of an illustrative Kabat CDR-H3, CDR-H2, and/or CDR-H1 sequence provided in this disclosure.
[0107] In some aspects, the Kabat CDR-H3 sequence comprises, consists of, or consists essentially of a variant of an illustrative Kabat CDR-H3 sequence provided in this disclosure. In some aspects, the Kabat CDR-H3 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative Kabat CDR-H3 sequences provided in this disclosure. In some aspects, the Kabat CDR-H3 sequence comprises, consists of, or consists essentially of any of the illustrative Kabat CDR-H3 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0108] In some aspects, the Kabat CDR-H2 sequence comprises, consists of, or consists essentially of a variant of an illustrative Kabat CDR-H2 sequence provided in this disclosure. In some aspects, the Kabat CDR-H2 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative Kabat CDR-H2 sequences provided in this disclosure. In some aspects, the Kabat CDR-H2 sequence comprises, consists of, or consists essentially of any of the illustrative Kabat CDR-H2 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0109] In some aspects, the Kabat CDR-H1 sequence comprises, consists of, or consists essentially of a variant of an illustrative Kabat CDR-H1 sequence provided in this disclosure. In some aspects, the Kabat CDR-H1 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative Kabat CDR-H1 sequences provided in this disclosure. In some aspects, the Kabat CDR-H1 sequence comprises, consists of, or consists essentially of any of the illustrative Kabat CDR-H1 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.2.2. V.sub.H Sequences Comprising Illustrative Chothia CDRs
[0110] In some embodiments, the antibody comprises a V.sub.H sequence comprising one or more Chothia CDR-H sequences comprising, consisting of, or consisting essentially of one or more illustrative Chothia CDR-H sequences provided in this disclosure, and variants thereof.
2.2.2.1. Chothia CDR-H3
[0111] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 82. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 83. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 84. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 85. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 86. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 87. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 88. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 89. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 90. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 91. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 92. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 93. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 94. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 95. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 96. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 97. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 98. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 99. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 100. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 101. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 102. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 103. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 104. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 105. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 106. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 107. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 108. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 109.
2.2.2.2. Chothia CDR-H2
[0112] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 46-62. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 46. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 47. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 48. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 49. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 50. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 51. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 52. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 53. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 54. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 55. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 56. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 57. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 58. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 59. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 60. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 61. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 62.
2.2.2.3. Chothia CDR-H1
[0113] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 1-24. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 1. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 2. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 3. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 4. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 5. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 6. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 7. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 8. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 9. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 10. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 11. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 12. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 13. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 14. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 15. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 16. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 17. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 18. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 19. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 20. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 21. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 22. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 23. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 24.
2.2.2.4. Chothia CDR-H3+Chothia CDR-H2
[0114] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109, and a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 46-62. In some aspects, the Chothia CDR-H3 sequence and the Chothia CDR-H2 sequence are both from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Chothia CDR-H3 and Chothia CDR-H2 are both from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
2.2.2.5. Chothia CDR-H3+Chothia CDR-H1
[0115] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109, and a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 1-24. In some aspects, the Chothia CDR-H3 sequence and the Chothia CDR-H1 sequence are both from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Chothia CDR-H3 and Chothia CDR-H1 are both from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
2.2.2.6. Chothia CDR-H1+Chothia CDR-112
[0116] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 1-24 and a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 46-62. In some aspects, the Chothia CDR-H1 sequence and the Chothia CDR-H2 sequence are both from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Chothia CDR-H1 and Chothia CDR-H2 are both from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
2.2.2.7. Chothia CDR-H1+Chothia CDR-H2+Chothia CDR-H3
[0117] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 1-24, a Chothia CDR-H2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 46-62, and a Chothia CDR-H3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 82-109. In some aspects, the Chothia CDR-H1 sequence, Chothia CDR-H2 sequence, and Chothia CDR-H3 sequence are all from a single illustrative V.sub.H sequence provided in this disclosure. For example, in some aspects, the Chothia CDR-H1, Chothia CDR-H2, and Chothia CDR-H3 are all from a single illustrative V.sub.H sequence selected from SEQ ID NOs: 179-218.
[0118] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 1, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 83. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 1, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 84. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 85. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 3, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 86. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 4, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 5, a Chothia CDR-H2 sequence comprising SEQ ID NO: 49, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 6, a Chothia CDR-H2 sequence comprising SEQ ID NO: 50, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 7, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 8, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 88. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 9, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 89. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 90. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 11, a Chothia CDR-H2 sequence comprising SEQ ID NO: 54, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 91. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 12, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 92. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 13, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 93. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 92. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 14, a Chothia CDR-H2 sequence comprising SEQ ID NO: 54, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 15, a Chothia CDR-H2 sequence comprising SEQ ID NO: 55, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 16, a Chothia CDR-H2 sequence comprising SEQ ID NO: 56, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 94. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 18, a Chothia CDR-H2 sequence comprising SEQ ID NO: 57, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 95. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 19, a Chothia CDR-H2 sequence comprising SEQ ID NO: 57, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 96. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 21, a Chothia CDR-H2 sequence comprising SEQ ID NO: 57, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 94. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 97. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 22, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 98. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 22, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 99. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 22, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 100. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 101. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 59, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 102. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 103. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 23, a Chothia CDR-H2 sequence comprising SEQ ID NO: 60, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 104. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 105. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 106. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 107. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 61, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 108. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 62, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 109.
[0119] In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 83. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 84. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 85. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 86. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 84. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 86. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 1, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 83. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 1, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 86. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 93. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 55, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 54, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 8, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 8, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 9, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 54, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 93. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 93. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 11, a Chothia CDR-H2 sequence comprising SEQ ID NO: 54, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 12, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 13, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 13, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 90. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 15, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87. In some embodiments, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 18, a Chothia CDR-H2 sequence comprising SEQ ID NO: 57, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 94.
2.2.2.8. Variants of V.sub.H Sequences Comprising Illustrative Chothia CDRs
[0120] In some embodiments, the V.sub.H sequences provided herein comprise a variant of an illustrative Chothia CDR-H3, CDR-H2, and/or CDR-H1 sequence provided in this disclosure.
[0121] In some aspects, the Chothia CDR-H3 sequence comprises, consists of, or consists essentially of a variant of an illustrative Chothia CDR-H3 sequence provided in this disclosure. In some aspects, the Chothia CDR-H3 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative Chothia CDR-H3 sequences provided in this disclosure. In some aspects, the Chothia CDR-H3 sequence comprises, consists of, or consists essentially of any of the illustrative Chothia CDR-H3 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0122] In some aspects, the Chothia CDR-H2 sequence comprises, consists of, or consists essentially of a variant of an illustrative Chothia CDR-H2 sequence provided in this disclosure. In some aspects, the Chothia CDR-H2 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative Chothia CDR-H2 sequences provided in this disclosure. In some aspects, the Chothia CDR-H2 sequence comprises, consists of, or consists essentially of any of the illustrative Chothia CDR-H2 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0123] In some aspects, the Chothia CDR-H1 sequence comprises, consists of, or consists essentially of a variant of an illustrative Chothia CDR-H1 sequence provided in this disclosure. In some aspects, the Chothia CDR-H1 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative Chothia CDR-H1 sequences provided in this disclosure. In some aspects, the Chothia CDR-H1 sequence comprises, consists of, or consists essentially of any of the illustrative Chothia CDR-H1 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.3. V.sub.H Sequences
[0124] In some embodiments, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 179-218. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 179. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 180. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 181. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 182. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 183. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 184. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 185. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 186. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 187. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 188. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 189. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 190. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 191. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 192. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 193. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 194. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 195. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 196. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 197. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 198. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 199. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 200. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 201. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 202. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 203. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 204. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 205. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 206. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 207. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 208. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 209. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 210. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 211. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 212. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 213. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 214. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 215. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 216. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 217. In some aspects, the antibody comprises a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 218.
2.3.1. Variants of V.sub.H Sequences
[0125] In some embodiments, the V.sub.H sequences provided herein comprise, consist of, or consist essentially of a variant of an illustrative V.sub.H sequence provided in this disclosure.
[0126] In some aspects, the V.sub.H sequence comprises, consists of, or consists essentially of a variant of an illustrative V.sub.H sequence provided in this disclosure. In some aspects, the V.sub.H sequence comprises, consists of, or consists essentially of a sequence having at least 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.5% identity with any of the illustrative V.sub.H sequences provided in this disclosure.
[0127] In some embodiments, the V.sub.H sequence comprises, consists of, or consists essentially of any of the illustrative V.sub.H sequences provided in this disclosure, 20 or fewer, 19 or fewer, 18 or fewer, 17 or fewer, 16 or fewer, 15 or fewer, 14 or fewer, 13 or fewer, 12 or fewer, 11 or fewer, 10 or fewer, 9 or fewer, 8 or fewer, 7 or fewer, 6 or fewer, 5 or fewer, 4 or fewer, 3 or fewer, 2 or fewer, or 1 or fewer amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.4. CDR-L3 Sequences
[0128] In some embodiments, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 141-166. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 141. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 142. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 143. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 144. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 145. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 146. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 147. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 148. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 149. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 150. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 151. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 152. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 153. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 154. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 155. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 156. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 157. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 158. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 159. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 160. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 161. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 162. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 163. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 164. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 165. In some aspects, the antibody comprises a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 166.
[0129] In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of a variant of an illustrative CDR-L3 sequence provided in this disclosure. In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative CDR-L3 sequences provided in this disclosure. In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of any of the illustrative CDR-L3 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.5. V.sub.L Sequences Comprising Illustrative CDRs
[0130] In some embodiments, the antibody comprises a V.sub.L sequence comprising one or more CDR-L sequences comprising, consisting of, or consisting essentially of one or more illustrative CDR-L sequences provided in this disclosure, and variants thereof.
2.5.1. CDR-L3
[0131] In some embodiments, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 141-166. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 142. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 143. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 144. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 145. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 146. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 147. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 149. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 150. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 151. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 152. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 153. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 154. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 155. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 156. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 157. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 158. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 159. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 160. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 161. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 162. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 163. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 164. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 165. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 166.
2.5.2. CDR-L2
[0132] In some embodiments, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 125-140. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 125. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 126. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 127. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 128. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 129. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 130. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 131. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 132. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 133. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 134. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 135. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 136. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 137. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 138. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 139. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L2 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 140.
2.5.3. CDR-L1
[0133] In some embodiments, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 110-124. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 110. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 111. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 112. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 113. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 114. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 115. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 116. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 117. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 118. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 119. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 120. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 121. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 122. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 123. In some aspects, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 124.
2.5.4. CDR-L3+CDR-L2
[0134] In some embodiments, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 141-166 and a CDR-L2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 125-140. In some aspects, the CDR-L3 sequence and the CDR-L2 sequence are both from a single illustrative V.sub.L sequence provided in this disclosure. For example, in some aspects, the CDR-L3 and CDR-L2 are both from a single illustrative V.sub.L sequence selected from SEQ ID NOs: 219-248.
2.5.5. CDR-L3+CDR-L1
[0135] In some embodiments, the antibody comprises a V.sub.L sequence comprising a CDR-L3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 141-166 and a CDR-L1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 110-124. In some aspects, the CDR-L3 sequence and the CDR-L1 sequence are both from a single illustrative V.sub.L sequence provided in this disclosure. For example, in some aspects, the CDR-L3 and CDR-L1 are both from a single illustrative V.sub.L sequence selected from SEQ ID NOs: 219-248.
2.5.6. CDR-L1+CDR-L2
[0136] In some embodiments, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 110-124 and a CDR-L2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 125-140. In some aspects, the CDR-L1 sequence and the CDR-L2 sequence are both from a single illustrative V.sub.L sequence provided in this disclosure. For example, in some aspects, the CDR-L1 and CDR-L2 are both from a single illustrative V.sub.L sequence selected from SEQ ID NOs: 219-248.
2.5.7. CDR-L1+CDR-L2+CDR-L3
[0137] In some embodiments, the antibody comprises a V.sub.L sequence comprising a CDR-L1 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 110-124, a CDR-L2 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 125-140, and a CDR-L3 sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L1 sequence, CDR-L2 sequence, and CDR-L3 sequence are all from a single illustrative V.sub.L sequence provided in this disclosure. For example, in some aspects, the CDR-L1, CDR-L2, and CDR-L3 are all from a single illustrative V.sub.L sequence selected from SEQ ID NOs: 219-248.
[0138] In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 111, a CDR-L2 sequence comprising SEQ ID NO: 126, and a CDR-L3 sequence comprising SEQ ID NO: 142. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 127, and a CDR-L3 sequence comprising sequence selected from SEQ ID NO: 142. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence comprising SEQ ID NO: 143. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 112, a CDR-L2 sequence comprising SEQ ID NO: 128, and a CDR-L3 sequence comprising SEQ ID NO: 144. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 111, a CDR-L2 sequence comprising SEQ ID NO: 126, and a CDR-L3 sequence comprising SEQ ID NO: 145. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 113, a CDR-L2 sequence comprising SEQ ID NO: 129, and a CDR-L3 sequence comprising SEQ ID NO: 146. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 114, a CDR-L2 sequence comprising SEQ ID NO: 130, and a CDR-L3 sequence comprising SEQ ID NO: 147. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence comprising SEQ ID NO: 148. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence comprising SEQ ID NO: 148. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 116, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence comprising SEQ ID NO: 149. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence comprising SEQ ID NO: 148. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 132, and a CDR-L3 sequence comprising SEQ ID NO: 149. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 117, a CDR-L2 sequence comprising SEQ ID NO: 133, and a CDR-L3 sequence comprising SEQ ID NO: 150. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 134, and a CDR-L3 sequence comprising SEQ ID NO: 148. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence comprising SEQ ID NO: 151. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence comprising SEQ ID NO: 152. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 136, and a CDR-L3 sequence comprising SEQ ID NO: 152. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence comprising SEQ ID NO: 153. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 119, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence comprising SEQ ID NO: 154. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 120, a CDR-L2 sequence comprising SEQ ID NO: 137, and a CDR-L3 sequence comprising SEQ ID NO: 155. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence comprising SEQ ID NO: 156. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence comprising SEQ ID NO: 157. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 122, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence comprising SEQ ID NO: 158. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence comprising SEQ ID NO: 159. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence comprising SEQ ID NO: 160. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 139, and a CDR-L3 sequence comprising SEQ ID NO: 161. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 123, a CDR-L2 sequence comprising SEQ ID NO: 140, and a CDR-L3 sequence comprising SEQ ID NO: 162. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence comprising SEQ ID NO: 163. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 139, and a CDR-L3 sequence comprising SEQ ID NO: 164. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 120, a CDR-L2 sequence comprising SEQ ID NO: 137, and a CDR-L3 sequence comprising SEQ ID NO: 165. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 124, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence comprising SEQ ID NO: 166.
[0139] In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 128, and a CDR-L3 sequence SEQ ID NO: 144. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence comprising SEQ ID NO: 144. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 126, and a CDR-L3 sequence comprising sequence selected from SEQ ID NO: 144. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 116, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence comprising SEQ ID NO: 150. In some aspects, the antibody comprises a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 117, a CDR-L2 sequence comprising SEQ ID NO: 134, and a CDR-L3 sequence comprising SEQ ID NO: 150.
2.5.8. Variants of VL Sequences Comprising Illustrative CDR-Ls
[0140] In some embodiments, the V.sub.L sequences provided herein comprise a variant of an illustrative CDR-L3, CDR-L2, and/or CDR-L1 sequence provided in this disclosure.
[0141] In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of a variant of an illustrative CDR-L3 sequence provided in this disclosure. In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative CDR-L3 sequences provided in this disclosure. In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of any of the illustrative CDR-L3 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0142] In some aspects, the CDR-L2 sequence comprises, consists of, or consists essentially of a variant of an illustrative CDR-L2 sequence provided in this disclosure. In some aspects, the CDR-L2 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative CDR-L2 sequences provided in this disclosure. In some aspects, the CDR-L2 sequence comprises, consists of, or consists essentially of any of the illustrative CDR-L2 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0143] In some aspects, the CDR-L1 sequence comprises, consists of, or consists essentially of a variant of an illustrative CDR-L1 sequence provided in this disclosure. In some aspects, the CDR-L1 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative CDR-L1 sequences provided in this disclosure. In some aspects, the CDR-L1 sequence comprises, consists of, or consists essentially of any of the illustrative CDR-L1 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.6. V.sub.L Sequences
[0144] In some embodiments, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of a sequence selected from SEQ ID NOs: 219-248. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 219. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 220. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 221. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 222. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 223. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 224. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 225. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 226. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 227. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 228. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 229. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 230. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 231. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 232. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 233. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 234. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 235. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 236. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 237. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 238. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 239. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 240. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 241. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 242. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 243. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 244. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 245. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 246. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 247. In some aspects, the antibody comprises a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 248.
2.6.1. Variants of V.sub.L Sequences
[0145] In some embodiments, the V.sub.L sequences provided herein comprise, consist of, or consist essentially of a variant of an illustrative V.sub.L sequence provided in this disclosure.
[0146] In some aspects, the V.sub.L sequence comprises, consists of, or consists essentially of a variant of an illustrative V.sub.L sequence provided in this disclosure. In some aspects, the V.sub.L sequence comprises, consists of, or consists essentially of a sequence having at least 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.05% identity with any of the illustrative V.sub.L sequences provided in this disclosure.
[0147] In some embodiments, the V.sub.L sequence comprises, consists of, or consists essentially of any of the illustrative V.sub.L sequences provided in this disclosure, 20 or fewer, 19 or fewer, 18 or fewer, 17 or fewer, 16 or fewer, 15 or fewer, 14 or fewer, 13 or fewer, 12 or fewer, 11 or fewer, 10 or fewer, 9 or fewer, 8 or fewer, 7 or fewer, 6 or fewer, 5 or fewer, 4 or fewer, 3 or fewer, 2 or fewer, or 1 or fewer amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.7. Pairs
2.7.1. CDR-H3-CDR-L3 Pairs
[0148] In some embodiments, the antibody comprises a CDR-H3 sequence and a CDR-L3 sequence. In some aspects, the CDR-H3 sequence is part of a V.sub.H and the CDR-L3 sequence is part of a V.sub.L.
[0149] In some aspects, the CDR-H3 sequence is a CDR-H3 sequence comprising, consisting of, or consisting essentially of SEQ ID NOs: 82-109, and the CDR-L3 sequence is a CDR-L3 sequence comprising, consisting of, or consisting essentially of SEQ ID NOs: 141-166.
[0150] In some aspects, the CDR-H3 sequence is SEQ ID NO: 82 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0151] In some aspects, the CDR-H3 sequence is SEQ ID NO: 83 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0152] In some aspects, the CDR-H3 sequence is SEQ ID NO: 84 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0153] In some aspects, the CDR-H3 sequence is SEQ ID NO: 85 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0154] In some aspects, the CDR-H3 sequence is SEQ ID NO: 86 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0155] In some aspects, the CDR-H3 sequence is SEQ ID NO: 87 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0156] In some aspects, the CDR-H3 sequence is SEQ ID NO: 88 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0157] In some aspects, the CDR-H3 sequence is SEQ ID NO: 89 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0158] In some aspects, the CDR-H3 sequence is SEQ ID NO: 90 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0159] In some aspects, the CDR-H3 sequence is SEQ ID NO: 91 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0160] In some aspects, the CDR-H3 sequence is SEQ ID NO: 92 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0161] In some aspects, the CDR-H3 sequence is SEQ ID NO: 93 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0162] In some aspects, the CDR-H3 sequence is SEQ ID NO: 94 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0163] In some aspects, the CDR-H3 sequence is SEQ ID NO: 95 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0164] In some aspects, the CDR-H3 sequence is SEQ ID NO: 96 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0165] In some aspects, the CDR-H3 sequence is SEQ ID NO: 97 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0166] In some aspects, the CDR-H3 sequence is SEQ ID NO: 98 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0167] In some aspects, the CDR-H3 sequence is SEQ ID NO: 99 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0168] In some aspects, the CDR-H3 sequence is SEQ ID NO: 100 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0169] In some aspects, the CDR-H3 sequence is SEQ ID NO: 101 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0170] In some aspects, the CDR-H3 sequence is SEQ ID NO: 102 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0171] In some aspects, the CDR-H3 sequence is SEQ ID NO: 103 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0172] In some aspects, the CDR-H3 sequence is SEQ ID NO: 104 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0173] In some aspects, the CDR-H3 sequence is SEQ ID NO: 105 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0174] In some aspects, the CDR-H3 sequence is SEQ ID NO: 106 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0175] In some aspects, the CDR-H3 sequence is SEQ ID NO: 107 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0176] In some aspects, the CDR-H3 sequence is SEQ ID NO: 108 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
[0177] In some aspects, the CDR-H3 sequence is SEQ ID NO: 109 and the CDR-L3 sequence is selected from SEQ ID NOs: 141-166. In some aspects, the CDR-L3 sequence is SEQ ID NO: 141. In some aspects, the CDR-L3 sequence is SEQ ID NO: 142. In some aspects, the CDR-L3 sequence is SEQ ID NO: 143. In some aspects, the CDR-L3 sequence is SEQ ID NO: 144. In some aspects, the CDR-L3 sequence is SEQ ID NO: 145. In some aspects, the CDR-L3 sequence is SEQ ID NO: 146. In some aspects, the CDR-L3 sequence is SEQ ID NO: 147. In some aspects, the CDR-L3 sequence is SEQ ID NO: 148. In some aspects, the CDR-L3 sequence is SEQ ID NO: 149. In some aspects, the CDR-L3 sequence is SEQ ID NO: 150. In some aspects, the CDR-L3 sequence is SEQ ID NO: 151. In some aspects, the CDR-L3 sequence is SEQ ID NO: 152. In some aspects, the CDR-L3 sequence is SEQ ID NO: 153. In some aspects, the CDR-L3 sequence is SEQ ID NO: 154. In some aspects, the CDR-L3 sequence is SEQ ID NO: 155. In some aspects, the CDR-L3 sequence is SEQ ID NO: 156. In some aspects, the CDR-L3 sequence is SEQ ID NO: 157. In some aspects, the CDR-L3 sequence is SEQ ID NO: 158. In some aspects, the CDR-L3 sequence is SEQ ID NO: 159. In some aspects, the CDR-L3 sequence is SEQ ID NO: 160. In some aspects, the CDR-L3 sequence is SEQ ID NO: 161. In some aspects, the CDR-L3 sequence is SEQ ID NO: 162. In some aspects, the CDR-L3 sequence is SEQ ID NO: 163. In some aspects, the CDR-L3 sequence is SEQ ID NO: 164. In some aspects, the CDR-L3 sequence is SEQ ID NO: 165. In some aspects, the CDR-L3 sequence is SEQ ID NO: 166.
2.7.1.1. Variants of CDR-H3-CDR-L3 Pairs
[0178] In some embodiments, the CDR-H3-CDR-L3 pairs provided herein comprise a variant of an illustrative CDR-H3 and/or CDR-L1 sequence provided in this disclosure.
[0179] In some aspects, the CDR-H3 sequence comprises, consists of, or consists essentially of a variant of an illustrative CDR-H3 sequence provided in this disclosure. In some aspects, the CDR-H3 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative CDR-H3 sequences provided in this disclosure. In some aspects, the CDR-H3 sequence comprises, consists of, or consists essentially of any of the illustrative CDR-H3 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0180] In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of a variant of an illustrative CDR-L3 sequence provided in this disclosure. In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of a sequence having at least 70%, 75%, 80%, 85%, 90%, or 95% identity with any of the illustrative CDR-L3 sequences provided in this disclosure. In some aspects, the CDR-L3 sequence comprises, consists of, or consists essentially of any of the illustrative CDR-L3 sequences provided in this disclosure, with 1, 2, or 3 amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.7.2. V.sub.H-V.sub.L Pairs
[0181] In some embodiments, the antibody comprises a V.sub.H sequence and a V.sub.L sequence.
[0182] In some aspects, the V.sub.H sequence is a V.sub.H sequence comprising, consisting of, or consisting essentially of SEQ ID NOs: 179-218 and the V.sub.L sequence is a V.sub.L sequence comprising, consisting of, or consisting essentially of SEQ ID NOs: 219-248.
[0183] In some aspects, the V.sub.H sequence is SEQ ID NO: 179 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0184] In some aspects, the V.sub.H sequence is SEQ ID NO: 180 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0185] In some aspects, the V.sub.H sequence is SEQ ID NO: 181 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0186] In some aspects, the V.sub.H sequence is SEQ ID NO: 182 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0187] In some aspects, the V.sub.H sequence is SEQ ID NO: 183 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0188] In some aspects, the V.sub.H sequence is SEQ ID NO: 184 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0189] In some aspects, the V.sub.H sequence is SEQ ID NO: 185 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0190] In some aspects, the V.sub.H sequence is SEQ ID NO: 186 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0191] In some aspects, the V.sub.H sequence is SEQ ID NO: 187 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0192] In some aspects, the V.sub.H sequence is SEQ ID NO: 188 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0193] In some aspects, the V.sub.H sequence is SEQ ID NO: 189 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0194] In some aspects, the V.sub.H sequence is SEQ ID NO: 190 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0195] In some aspects, the V.sub.H sequence is SEQ ID NO: 191 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0196] In some aspects, the V.sub.H sequence is SEQ ID NO: 192 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0197] In some aspects, the V.sub.H sequence is SEQ ID NO: 193 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0198] In some aspects, the V.sub.H sequence is SEQ ID NO: 194 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0199] In some aspects, the V.sub.H sequence is SEQ ID NO: 195 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0200] In some aspects, the V.sub.H sequence is SEQ ID NO: 196 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0201] In some aspects, the V.sub.H sequence is SEQ ID NO: 197 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0202] In some aspects, the V.sub.H sequence is SEQ ID NO: 198 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0203] In some aspects, the V.sub.H sequence is SEQ ID NO: 199 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0204] In some aspects, the V.sub.H sequence is SEQ ID NO: 200 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0205] In some aspects, the V.sub.H sequence is SEQ ID NO: 201 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0206] In some aspects, the V.sub.H sequence is SEQ ID NO: 202 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0207] In some aspects, the V.sub.H sequence is SEQ ID NO: 203 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0208] In some aspects, the V.sub.H sequence is SEQ ID NO: 204 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0209] In some aspects, the V.sub.H sequence is SEQ ID NO: 205 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0210] In some aspects, the V.sub.H sequence is SEQ ID NO: 206 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0211] In some aspects, the V.sub.H sequence is SEQ ID NO: 207 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0212] In some aspects, the V.sub.H sequence is SEQ ID NO: 208 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0213] In some aspects, the V.sub.H sequence is SEQ ID NO: 209 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0214] In some aspects, the V.sub.H sequence is SEQ ID NO: 210 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0215] In some aspects, the V.sub.H sequence is SEQ ID NO: 211 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0216] In some aspects, the V.sub.H sequence is SEQ ID NO: 212 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0217] In some aspects, the V.sub.H sequence is SEQ ID NO: 213 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0218] In some aspects, the V.sub.H sequence is SEQ ID NO: 214 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0219] In some aspects, the V.sub.H sequence is SEQ ID NO: 215 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0220] In some aspects, the V.sub.H sequence is SEQ ID NO: 216 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0221] In some aspects, the V.sub.H sequence is SEQ ID NO: 217 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
[0222] In some aspects, the V.sub.H sequence is SEQ ID NO: 218 and the V.sub.L sequence is selected from SEQ ID NOs: 219-248. In some aspects, the V.sub.L sequence is SEQ ID NO: 219. In some aspects, the V.sub.L sequence is SEQ ID NO: 220. In some aspects, the V.sub.L sequence is SEQ ID NO: 221. In some aspects, the V.sub.L sequence is SEQ ID NO: 222. In some aspects, the V.sub.L sequence is SEQ ID NO: 223. In some aspects, the V.sub.L sequence is SEQ ID NO: 224. In some aspects, the V.sub.L sequence is SEQ ID NO: 225. In some aspects, the V.sub.L sequence is SEQ ID NO: 226. In some aspects, the V.sub.L sequence is SEQ ID NO: 227. In some aspects, the V.sub.L sequence is SEQ ID NO: 228. In some aspects, the V.sub.L sequence is SEQ ID NO: 229. In some aspects, the V.sub.L sequence is SEQ ID NO: 230. In some aspects, the V.sub.L sequence is SEQ ID NO: 231. In some aspects, the V.sub.L sequence is SEQ ID NO: 232. In some aspects, the V.sub.L sequence is SEQ ID NO: 233. In some aspects, the V.sub.L sequence is SEQ ID NO: 234. In some aspects, the V.sub.L sequence is SEQ ID NO: 235. In some aspects, the V.sub.L sequence is SEQ ID NO: 236. In some aspects, the V.sub.L sequence is SEQ ID NO: 237. In some aspects, the V.sub.L sequence is SEQ ID NO: 238. In some aspects, the V.sub.L sequence is SEQ ID NO: 239. In some aspects, the V.sub.L sequence is SEQ ID NO: 240. In some aspects, the V.sub.L sequence is SEQ ID NO: 241. In some aspects, the V.sub.L sequence is SEQ ID NO: 242. In some aspects, the V.sub.L sequence is SEQ ID NO: 243. In some aspects, the V.sub.L sequence is SEQ ID NO: 244. In some aspects, the V.sub.L sequence is SEQ ID NO: 245. In some aspects, the V.sub.L sequence is SEQ ID NO: 246. In some aspects, the V.sub.L sequence is SEQ ID NO: 247. In some aspects, the V.sub.L sequence is SEQ ID NO: 248.
2.7.3. CDR-H1+CDR-112+CDR-113+CDR-L1+CDR-L2+CDR-L3
[0223] In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 25, a Kabat CDR-H2 sequence comprising SEQ ID NO: 63, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 83 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 111, a CDR-L2 sequence comprising SEQ ID NO: 126, and a CDR-L3 sequence SEQ ID NO: 142. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 27, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 84 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 85 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 86 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 127, and a CDR-L3 sequence SEQ ID NO: 142. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 25, a Kabat CDR-H2 sequence comprising SEQ ID NO: 66, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 143. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 112, a CDR-L2 sequence comprising SEQ ID NO: 128, and a CDR-L3 sequence SEQ ID NO: 144. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 27, a Kabat CDR-H2 sequence comprising SEQ ID NO: 65, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 111, a CDR-L2 sequence comprising SEQ ID NO: 126, and a CDR-L3 sequence SEQ ID NO: 145. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 28, a Kabat CDR-H2 sequence comprising SEQ ID NO: 67, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 113, a CDR-L2 sequence comprising SEQ ID NO: 129, and a CDR-L3 sequence SEQ ID NO: 146. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 29, a Kabat CDR-H2 sequence comprising SEQ ID NO: 68, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 114, a CDR-L2 sequence comprising SEQ ID NO: 130, and a CDR-L3 sequence SEQ ID NO: 147. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 26, a Kabat CDR-H2 sequence comprising SEQ ID NO: 64, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 31, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 88 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 116, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 149. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 32, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 89 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 90 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 132, and a CDR-L3 sequence SEQ ID NO: 149. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 34, a Kabat CDR-H2 sequence comprising SEQ ID NO: 72, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 91 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 71, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 92 and a V.sub.L sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 35, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 93 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 117, a CDR-L2 sequence comprising SEQ ID NO: 133, and a CDR-L3 sequence SEQ ID NO: 150. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 33, a Kabat CDR-H2 sequence comprising SEQ ID NO: 70, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 92 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 134, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 36, a Kabat CDR-H2 sequence comprising SEQ ID NO: 72, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 30, a Kabat CDR-H2 sequence comprising SEQ ID NO: 73, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 37, a Kabat CDR-H2 sequence comprising SEQ ID NO: 74, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 75, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 94 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence SEQ ID NO: 151. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 39, a Kabat CDR-H2 sequence comprising SEQ ID NO: 76, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 95 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence SEQ ID NO: 152. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 40, a Kabat CDR-H2 sequence comprising SEQ ID NO: 76, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 96 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 136, and a CDR-L3 sequence SEQ ID NO: 152. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 39, a Kabat CDR-H2 sequence comprising SEQ ID NO: 76, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 94 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence SEQ ID NO: 152. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 97 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 153. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 41, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 98 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 119, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 154. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 41, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 99 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 120, a CDR-L2 sequence comprising SEQ ID NO: 137, and a CDR-L3 sequence SEQ ID NO: 155. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 41, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 100 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 156. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 75, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 101 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 157. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 42, a Kabat CDR-H2 sequence comprising SEQ ID NO: 78, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 102 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 122, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 158. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 75, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 103 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 159. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 43, a Kabat CDR-H2 sequence comprising SEQ ID NO: 79, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 104 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence SEQ ID NO: 160. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 75, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 103 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 139, and a CDR-L3 sequence SEQ ID NO: 161. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 105 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 123, a CDR-L2 sequence comprising SEQ ID NO: 140, and a CDR-L3 sequence SEQ ID NO: 162. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 106 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 163. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 38, a Kabat CDR-H2 sequence comprising SEQ ID NO: 69, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 107 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 139, and a CDR-L3 sequence SEQ ID NO: 164. In some aspects, the antibody comprises a V.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 44, a Kabat CDR-H2 sequence comprising SEQ ID NO: 80, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 108 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 120, a CDR-L2 sequence comprising SEQ ID NO: 137, and a CDR-L3 sequence SEQ ID NO: 165. In some aspects, the antibody comprises a V
.sub.H sequence comprising a Kabat CDR-H1 sequence comprising SEQ ID NO: 45, a Kabat CDR-H2 sequence comprising SEQ ID NO: 81, and a Kabat CDR-H3 sequence comprising SEQ ID NO: 109 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 124, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 166.
[0224] In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 1, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 83 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 111, a CDR-L2 sequence comprising SEQ ID NO: 126, and a CDR-L3 sequence SEQ ID NO: 142. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 1, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 84 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 85 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 3, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 86 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 127, and a CDR-L3 sequence SEQ ID NO: 142. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 4, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 143. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 112, a CDR-L2 sequence comprising SEQ ID NO: 128, and a CDR-L3 sequence SEQ ID NO: 144. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 1, a Chothia CDR-H2 sequence comprising SEQ ID NO: 46, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 111, a CDR-L2 sequence comprising SEQ ID NO: 126, and a CDR-L3 sequence SEQ ID NO: 145. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 5, a Chothia CDR-H2 sequence comprising SEQ ID NO: 49, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 113, a CDR-L2 sequence comprising SEQ ID NO: 129, and a CDR-L3 sequence SEQ ID NO: 146. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 6, a Chothia CDR-H2 sequence comprising SEQ ID NO: 50, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 114, a CDR-L2 sequence comprising SEQ ID NO: 130, and a CDR-L3 sequence SEQ ID NO: 147. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 2, a Chothia CDR-H2 sequence comprising SEQ ID NO: 47, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 82 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 110, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 141. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 7, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 8, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 88 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 116, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 149. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 9, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 89 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 90 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 132, and a CDR-L3 sequence SEQ ID NO: 149. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 11, a Chothia CDR-H2 sequence comprising SEQ ID NO: 54, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 91 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 12, a Chothia CDR-H2 sequence comprising SEQ ID NO: 53, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 92 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 13, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 93 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 117, a CDR-L2 sequence comprising SEQ ID NO: 133, and a CDR-L3 sequence SEQ ID NO: 150. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 10, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 92 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 134, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 14, a Chothia CDR-H2 sequence comprising SEQ ID NO: 54, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 15, a Chothia CDR-H2 sequence comprising SEQ ID NO: 55, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 16, a Chothia CDR-H2 sequence comprising SEQ ID NO: 56, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 87 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 148. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 94 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence SEQ ID NO: 151. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 18, a Chothia CDR-H2 sequence comprising SEQ ID NO: 57, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 95 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135 and a CDR-L3 sequence SEQ ID NO: 152. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 19, a Chothia CDR-H2 sequence comprising SEQ ID NO: 57, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 96 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 136, and a CDR-L3 sequence SEQ ID NO: 152. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 21, a Chothia CDR-H2 sequence comprising SEQ ID NO: 57, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 94 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence SEQ ID NO: 152. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 97 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 115, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 153. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 22, a Chothia CDR-H2 sequence comprising SEQ ID NO: 52, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 98 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 119, a CDR-L2 sequence comprising SEQ ID NO: 131, and a CDR-L3 sequence SEQ ID NO: 154. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 22, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 99 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 120, a CDR-L2 sequence comprising SEQ ID NO: 137, and a CDR-L3 sequence SEQ ID NO: 155. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 22, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 100 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 156. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 101 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 157. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 59, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 102 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 122, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 158. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 103 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 159. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 23, a Chothia CDR-H2 sequence comprising SEQ ID NO: 60, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 104 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 118, a CDR-L2 sequence comprising SEQ ID NO: 135, and a CDR-L3 sequence SEQ ID NO: 160. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 103 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 139, and a CDR-L3 sequence SEQ ID NO: 161. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 105 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 123, a CDR-L2 sequence comprising SEQ ID NO: 140, and a CDR-L3 sequence SEQ ID NO: 162. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 106 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 138, and a CDR-L3 sequence SEQ ID NO: 163. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 51, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 107 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 121, a CDR-L2 sequence comprising SEQ ID NO: 139, and a CDR-L3 sequence SEQ ID NO: 164. In some aspects, the antibody comprises a V
.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 61, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 108 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 120, a CDR-L2 sequence comprising SEQ ID NO: 137, and a CDR-L3 sequence SEQ ID NO: 165. In some aspects, the antibody comprises a V.sub.H sequence comprising a Chothia CDR-H1 sequence comprising SEQ ID NO: 17, a Chothia CDR-H2 sequence comprising SEQ ID NO: 62, and a Chothia CDR-H3 sequence comprising SEQ ID NO: 109 and a VL sequence comprising a CDR-L1 sequence comprising SEQ ID NO: 124, a CDR-L2 sequence comprising SEQ ID NO: 125, and a CDR-L3 sequence SEQ ID NO: 166.
2.7.3.1. Variants of V.sub.H-V.sub.L Pairs
[0225] In some embodiments, the V.sub.H-V.sub.L pairs provided herein comprise a variant of an illustrative V.sub.H and/or V.sub.L sequence provided in this disclosure.
[0226] In some aspects, the V.sub.H sequence comprises, consists of, or consists essentially of a variant of an illustrative V.sub.H sequence provided in this disclosure. In some aspects, the V.sub.H sequence comprises, consists of, or consists essentially of a sequence having at least 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.1% identity with any of the illustrative V.sub.H sequences provided in this disclosure.
[0227] In some embodiments, the V.sub.H sequence comprises, consists of, or consists essentially of any of the illustrative V.sub.H sequences provided in this disclosure, 20 or fewer, 19 or fewer, 18 or fewer, 17 or fewer, 16 or fewer, 15 or fewer, 14 or fewer, 13 or fewer, 12 or fewer, 11 or fewer, 10 or fewer, 9 or fewer, 8 or fewer, 7 or fewer, 6 or fewer, 5 or fewer, 4 or fewer, 3 or fewer, 2 or fewer, or 1 or fewer amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
[0228] In some aspects, the V.sub.L sequence comprises, consists of, or consists essentially of a variant of an illustrative V.sub.L sequence provided in this disclosure. In some aspects, the V.sub.L sequence comprises, consists of, or consists essentially of a sequence having at least 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.05% identity with any of the illustrative V.sub.L sequences provided in this disclosure.
[0229] In some embodiments, the V.sub.L sequence comprises, consists of, or consists essentially of any of the illustrative V.sub.L sequences provided in this disclosure, 20 or fewer, 19 or fewer, 18 or fewer, 17 or fewer, 16 or fewer, 15 or fewer, 14 or fewer, 13 or fewer, 12 or fewer, 11 or fewer, 10 or fewer, 9 or fewer, 8 or fewer, 7 or fewer, 6 or fewer, 5 or fewer, 4 or fewer, 3 or fewer, 2 or fewer, or 1 or fewer amino acid substitutions. In some aspects, the amino acid substitutions are conservative amino acid substitutions.
2.7.4 HC+LC
[0230] In some embodiments, the antibody comprises or consists of one or more heavy chains consisting of an HC sequence and one or more light chains consisting of an LC sequence. In some embodiments, the antibody comprises or consists of two identical heavy chains consisting of an HC sequence and two identical light chains consisting of an LC sequence.
[0231] In some embodiments, the HC sequence is an HC sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 255, SEQ ID NO: 257, SEQ ID NO: 259, SEQ ID NO: 261, SEQ ID NO: 263, SEQ ID NO: 265, SEQ ID NO: 267, SEQ ID NO: 269, SEQ ID NO: 271, SEQ ID NO: 273, SEQ ID NO: 275, SEQ ID NO: 277, SEQ ID NO: 279, SEQ ID NO: 281, SEQ ID NO: 283, SEQ ID NO: 285, SEQ ID NO: 287, SEQ ID NO: 289, SEQ ID NO: 291, SEQ ID NO: 293, SEQ ID NO: 295, or SEQ ID NO: 297 and the LC sequence is a LC sequence comprising, consisting of, or consisting essentially of SEQ ID NO: 256, SEQ ID NO: 258, SEQ ID NO: 260, SEQ ID NO: 262, SEQ ID NO: 264, SEQ ID NO: 266, SEQ ID NO: 268, SEQ ID NO: 270, SEQ ID NO: 272, SEQ ID NO: 274, SEQ ID NO: 276, SEQ ID NO: 278, SEQ ID NO: 280, SEQ ID NO: 282, SEQ ID NO: 284, SEQ ID NO: 286, SEQ ID NO: 288, SEQ ID NO: 290, SEQ ID NO: 292, SEQ ID NO: 294, SEQ ID NO: 296, or SEQ ID NO: 298. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 255, SEQ ID NO: 257, SEQ ID NO: 259, SEQ ID NO: 261, SEQ ID NO: 263, SEQ ID NO: 265, SEQ ID NO: 267, SEQ ID NO: 269, SEQ ID NO: 271, SEQ ID NO: 273, SEQ ID NO: 275, SEQ ID NO: 277, SEQ ID NO: 279, SEQ ID NO: 281, SEQ ID NO: 283, SEQ ID NO: 285, SEQ ID NO: 287, SEQ ID NO: 289, SEQ ID NO: 291, SEQ ID NO: 293, SEQ ID NO: 295, or SEQ ID NO: 297 and the LC sequence is an LC sequence consisting of SEQ ID NO: 256, SEQ ID NO: 258, SEQ ID NO: 260, SEQ ID NO: 262, SEQ ID NO: 264, SEQ ID NO: 266, SEQ ID NO: 268, SEQ ID NO: 270, SEQ ID NO: 272, SEQ ID NO: 274, SEQ ID NO: 276, SEQ ID NO: 278, SEQ ID NO: 280, SEQ ID NO: 282, SEQ ID NO: 284, SEQ ID NO: 286, SEQ ID NO: 288, SEQ ID NO: 290, SEQ ID NO: 292, SEQ ID NO: 294, SEQ ID NO: 296, or SEQ ID NO: 298.
[0232] In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 255 and the LC sequence is an LC sequence consisting of SEQ ID NO: 256. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 257 and the LC sequence is an LC sequence consisting of SEQ ID NO: 258. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 259 and the LC sequence is an LC sequence consisting of SEQ ID NO: 260. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 261 and the LC sequence is an LC sequence consisting of SEQ ID NO: 262. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 263 and the LC sequence is an LC sequence consisting of SEQ ID NO: 264. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 265 and the LC sequence is an LC sequence consisting of SEQ ID NO: 266. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 267 and the LC sequence is an LC sequence consisting of SEQ ID NO: 268. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 269 and the LC sequence is an LC sequence consisting of SEQ ID NO: 270. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 271 and the LC sequence is an LC sequence consisting of SEQ ID NO: 272. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 273 and the LC sequence is an LC sequence consisting of SEQ ID NO: 274. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 275 and the LC sequence is an LC sequence consisting of SEQ ID NO: 276. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 277 and the LC sequence is an LC sequence consisting of SEQ ID NO: 278. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 279 and the LC sequence is an LC sequence consisting of SEQ ID NO: 280. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 281 and the LC sequence is an LC sequence consisting of SEQ ID NO: 282. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 283 and the LC sequence is an LC sequence consisting of SEQ ID NO: 284. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 285 and the LC sequence is an LC sequence consisting of SEQ ID NO: 286. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 287 and the LC sequence is an LC sequence consisting of SEQ ID NO: 288. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 289 and the LC sequence is an LC sequence consisting of SEQ ID NO: 290. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 291 and the LC sequence is an LC sequence consisting of SEQ ID NO: 292. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 293 and the LC sequence is an LC sequence consisting of SEQ ID NO: 294. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 295 and the LC sequence is an LC sequence consisting of SEQ ID NO: 296. In some embodiments, the HC sequence is an HC sequence consisting of SEQ ID NO: 297 and the LC sequence is an LC sequence consisting of SEQ ID NO: 298.
2.8. Consensus Sequences
[0233] In some embodiments, provided herein are anti-CD39 antibodies comprising one or more sequences defined by consensus sequences. Each consensus sequence is based, at least in part, on one or more alignments of two or more useful anti-CD39 CDR sequences provided in this disclosure. Based on such alignments, a person of skill in the art would recognize that different amino acid residues may useful in certain positions of the CDRs. Accordingly, each consensus sequence encompasses two or more useful anti-CD39 CDR sequences.
2.8.1. CDR-113 Consensus Sequences
[0234] In some embodiments, the antibody comprises a CDR-H3 sequence defined by the consensus sequence G-K-R-E-G-G-T-E-Y-L-R-.sub.12 (SEQ ID NOS: 82-86), where .sub.12 is H, K, S, N, or V.
[0235] In some aspects, .sub.12 is H. In some aspects, .sub.12 is K. In some aspects, .sub.12 is S. In some aspects, .sub.12 is N. In some aspects, .sub.12 is V.
[0236] In some embodiments, the antibody comprises a CDR-H3 sequence defined by the consensus sequence E-S-G-.PHI..sub.4-Y-R-D-H-R-L-.PHI..sub.11-V (SEQ ID NOS: 94-96), where .PHI..sub.4 is G or T and .PHI..sub.11 is D or G.
[0237] In some aspects, .PHI..sub.4 is G when .PHI..sub.11 is D or G. In some aspects, .PHI..sub.11 is D when .PHI..sub.4 is G or T.
[0238] In some aspects, .psi..sub.4 is G and .PHI..sub.11 is D. In some aspects, .PHI..sub.4 is T and .PHI..sub.11 is D. In some aspects, .PHI..sub.4 is G and .PHI..sub.11 is G.
[0239] In some embodiments, the antibody comprises a CDR-H3 sequence defined by the consensus sequence G-G-A-K-Y-A-.sub.7-.sub.8-.sub.9-G-M-D-V (SEQ ID NOS: 87-93), where .sub.7 is S, V, G, or R; .sub.8 is T, Q, K, G, or R; and .sub.9 is Y, H, L, or W.
[0240] In some aspects, .sub.7 is S when .sub.8 is T, Q, or K and .sub.9 is Y, H, L, or W. In some aspects, .sub.8 is T when .sub.7 is S or R and .sub.9 is Y or H. In some aspects, .sub.9 is Y when .sub.7 is S, V, G, or R and .sub.8 is T, G, or R.
[0241] In some aspects, .sub.7 is S when .sub.8 is T and .sub.9 is Y. In some aspects, .sub.7 is S when .sub.8 is T and .sub.9 is H. In some aspects, .sub.7 is S when .sub.8 is Q and .sub.9 is L. In some aspects, .sub.7 is S when .sub.8 is K and .sub.9 is W. In some aspects, .sub.7 is V when .sub.8 is G and .sub.9 is Y. In some aspects, .sub.7 is G when .sub.8 is R and .sub.9 is Y. In some aspects, .sub.7 is R when .sub.8 is T and .sup.9 is Y.
2.8.2. Chothia CDR-H2 Consensus Sequences
[0242] In some embodiments, the antibody comprises a Chothia CDR-H2 sequence defined by the consensus sequence N-P-.epsilon..sub.5-.epsilon..sub.6-G-S-T (SEQ ID NOS: 46-48), where .epsilon..sub.5 is L, R, or S and .epsilon..sub.6 is G or V.
[0243] In some aspects, when .epsilon..sub.5 is S, .epsilon..sub.6 is G or V. In some aspects, when .epsilon..sub.6 is G, .epsilon..sub.5 is S, L, or R.
[0244] In some aspects, when .epsilon..sub.5 is L, .epsilon..sub.6 is G. In some aspects, when .epsilon..sub.5 is S, .epsilon..sub.6 is G. In some aspects, when .epsilon..sub.5 is S, .epsilon..sub.6 is V. In some aspects, when .epsilon..sub.5 is R, .epsilon..sub.6 is G.
[0245] In some embodiments, the antibody comprises a Chothia CDR-H2 sequence defined by the consensus sequence .alpha..sub.3-.alpha..sub.4-.alpha..sub.5-.alpha..sub.6-G-T-A (SEQ ID NOS: 51-54), where .alpha..sub.3 is I or L or is absent; .alpha..sub.4 is P or is absent; and .alpha..sub.5 is I, G, or R; and .alpha..sub.6 is A, F, or G.
[0246] In some aspects, when .alpha..sub.3, is I, .alpha..sub.4 is P; .alpha..sub.5 is I or R; and .alpha..sub.6 is F or G. In some aspects, when .alpha..sub.3, is L, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is A or G. In some aspects, when .alpha..sub.4, is P, .alpha..sub.3 is I or L; .alpha..sub.5 is I or R; and .alpha..sub.6 is A, F, or G. In some aspects, when as, is I, .alpha..sub.3 is I or L; .alpha..sub.4 is P; and .alpha..sub.6 is A, F, or G. In some aspects, when .alpha..sub.6, is F, .alpha..sub.3 is I or is absent; .alpha..sub.4 is P or is absent; and .alpha..sub.5 is I or G. In some aspects, when .alpha..sub.6, is G, .alpha..sub.3 is I or L; .alpha..sub.4 is P; and .alpha..sub.5 is I or R.
[0247] In some aspects, when .alpha..sub.3 is L, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is A. In some aspects, when .alpha..sub.3 is I, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is F. In some aspects, when .alpha..sub.3 is absent, .alpha..sub.4 is absent; .alpha..sub.5 is G; and .alpha..sub.6 is F. In some aspects, when .alpha..sub.3 is L, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is G. In some aspects, when .alpha..sub.3 is I, .alpha..sub.4 is P; .alpha..sub.5 is R; and .alpha..sub.6 is G.
[0248] In some embodiments, the antibody comprises a Chothia CDR-H2 sequence defined by the consensus sequence I-P-.beta..sub.5-.beta..sub.6-G-.beta..sub.8-A (SEQ ID NOS: 56-60), where .beta..sub.5 is I, E, S, or T; .beta..sub.6 is F, I, or S; and .beta..sub.8 is I or T.
[0249] In some aspects, when .beta..sub.5 is I, .beta..sub.6 is F or S and .beta..sub.8 is T. In some aspects, when .beta..sub.6 is F, .beta..sub.5 is E, I, or T and .beta..sub.8 is I or T. In some aspects, when .beta..sub.8 is T, .beta..sub.5 is I, S, or T and .beta..sub.6 is F, I, or S.
[0250] In some aspects, when .beta..sub.5 is I, .beta..sub.6 is F and .beta..sub.8 is T. In some aspects, when .beta..sub.5 is E, .beta..sub.6 is F and .beta..sub.8 is I. In some aspects, when .beta..sub.5 is S, .beta..sub.6 is I and .beta..sub.8 is T. In some aspects, when .beta..sub.5 is I, .beta..sub.6 is S and .beta..sub.8 is T. In some aspects, when .beta..sub.5 is T, .beta..sub.6 is S and .beta..sub.8 is T.
2.8.3. Chothia CDR-H1 Consensus Sequences
[0251] In some embodiments, the antibody comprises a Chothia CDR-H1 sequence defined by the consensus sequence G-Y-T-F-.OMEGA..sub.5-S-Y (SEQ ID NOS: 1-2 and 4-6), where .OMEGA..sub.5 is T, K, Q, F, or V.
[0252] In some aspects, .OMEGA..sub.5 is T. In some aspects, .OMEGA..sub.5 is K. In some aspects, .OMEGA..sub.5 is Q. In some aspects, .OMEGA..sub.5 is F. In some aspects, .OMEGA..sub.5 is V.
[0253] In some embodiments, the antibody comprises a Chothia CDR-H1 sequence defined by the consensus sequence G-G-T-F-.nu..sub.5-.nu..sub.6-Y (SEQ ID NOS: 17-22 and 24), where .nu..sub.5 is S, G, or E and .nu..sub.6 is S, K, R, or S.
[0254] In some aspects, .nu..sub.5 is S when .nu..sub.6 is S or K. In some aspects, .nu..sub.6 is S when .nu..sub.5 is S or E.
[0255] In some aspects, .nu..sub.5 is S when .nu..sub.6 is S. In some embodiments, .nu..sub.5 is S when .nu..sub.6 is K. In some aspects, .nu..sub.5 is G when .nu..sub.6 is R. In some aspects .nu..sub.5 is E when .nu..sub.6 is S.
[0256] In some embodiments, the antibody comprises a Chothia CDR-H1 sequence defined by the consensus sequence G-G-T-F-.kappa..sub.5-.kappa..sub.6-.kappa..sub.7 (SEQ ID NOS: 7-16), where .kappa..sub.5 is S, Q, P, or A; .kappa..sub.6 is S, K, H, L, A, or W; and .kappa..sub.7 is Y, L, T, N, or M.
[0257] In some aspects, when .kappa..sub.5 is S, .kappa.6 is S, K, H, L, A, or W and .kappa..sub.7 is Y, L, T, or M. In some aspects, when .kappa..sub.6 is S, .kappa..sub.5 is S, Q, P, or A and .kappa..sub.7 is Y, L, or N. In some aspects, when .kappa..sub.7 is L, .kappa..sub.5 is S, Q, or A and .kappa..sub.6 is S, K, L, or W.
[0258] In some aspects, when .kappa..sub.5 is S, .kappa..sub.6 is S and .kappa..sub.7 is Y. In some aspects, when .kappa..sub.5 is S, is S and .kappa..sub.7 is L. In some aspects, when .kappa..sub.5 is S, .kappa..sub.6 is K and .kappa..sub.7 is L. In some aspects, when .kappa..sub.5 is S, .kappa..sub.6 is H and .kappa..sub.7 is T. In some aspects, when .kappa..sub.5 is S, .kappa..sub.6 is L and .kappa..sub.7 is L. In some aspects, when .kappa..sub.5 is Q, .kappa..sub.6 is S and .kappa..sub.7 is L. In some aspects, when .kappa..sub.5 is P, .kappa..sub.6 is S and .kappa..sub.7 is N. In some aspects, when .kappa..sub.5 is S, .kappa..sub.6 is A and .kappa..sub.7 is M. In some aspects, when .kappa..sub.5 is A, .kappa..sub.6 is S and .kappa..sub.7 is L. In some aspects, when .kappa..sub.5 is S, .kappa..sub.6 is W and .kappa..sub.7 is L.
2.8.4. Kabat CDR-H2 Consensus Sequences
[0259] In some embodiments, the antibody comprises a Kabat CDR-H2 sequence defined by the consensus sequence .epsilon..sub.1-I-N-P-.epsilon..sub.5-.epsilon..sub.6-G-S-T-.epsilon..sub- .10-Y-A-Q-K-F-Q-G (SEQ ID NOS: 63-66 and 68), where .epsilon..sub.1 is K, S, R, or V; .epsilon..sub.5 is L, R, or S; .epsilon..sub.6 is G or V; and .epsilon..sub.10 is S or W.
[0260] In some aspects, when .epsilon..sub.1 is V; .epsilon..sub.5 is L or S; .epsilon..sub.6 is G; and .epsilon..sub.10 is S. In some aspects, when .epsilon..sub.1 is R; .epsilon..sub.5 is S; .epsilon..sub.6 is V or G; and .epsilon..sub.10 is W. In some aspects, when .epsilon..sub.5 is S; .epsilon..sub.1 is R or V; .epsilon..sub.6 is G or V; and .epsilon..sub.10 is S or W. In some aspects, when .epsilon..sub.6 is G; .epsilon..sub.1 is R or V; .epsilon..sub.5 is S; and .epsilon..sub.10 is S or W. In some aspects, when .epsilon..sub.10 is S; .epsilon..sub.1 is V or S; .epsilon..sub.5 is L, R, or S; and .epsilon..sub.6 is G. In some aspects, when .epsilon..sub.10 is W; .epsilon..sub.1 is K or R; .epsilon..sub.5 is S; and .epsilon..sub.6 is G or V.
[0261] In some aspects, when .epsilon..sub.1 is V; .epsilon..sub.5 is L; .epsilon..sub.6 is G; and .epsilon..sub.10 is S. In some aspects, when .epsilon..sub.1 is V; .epsilon..sub.5 is S; .epsilon..sub.6 is G; and .epsilon..sub.10 is S. In some aspects, when .epsilon..sub.1 is R; .epsilon..sub.5 is S; .epsilon..sub.6 is V; and .epsilon..sub.10 is W. In some aspects, when .epsilon..sub.1 is R; .epsilon..sub.5 is S; .epsilon..sub.6 is G; and .epsilon..sub.10 is W. In some aspects, when .epsilon..sub.1 is K; .epsilon..sub.5 is S; .epsilon..sub.6 is G; and .epsilon..sub.10 is W. In some aspects, when .epsilon..sub.1 is S; .epsilon..sub.5 is R; .epsilon..sub.6 is G; and .epsilon..sub.10 is S.
[0262] In some embodiments, the antibody comprises a Kabat CDR-H2 sequence defined by the consensus sequence G-I-.alpha..sub.3-.alpha..sub.4-.alpha..sub.5-.alpha..sub.6-G-T-A-N-Y-A-Q- -K-F-Q-G (SEQ ID NOS: 69-72), where .alpha..sub.3 is I or L or is absent; .alpha..sub.4 is P or is absent; and .alpha..sub.5 is I, G, or R; and .alpha..sub.6 is A, F, or G.
[0263] In some aspects, when .alpha..sub.3, is I, .alpha..sub.4 is P; as is I or R; and .alpha..sub.6 is F or G. In some aspects, when .alpha..sub.3, is L, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is A or G. In some aspects, when .alpha..sub.4, is P, .alpha..sub.3 is I or L; as is I or R; and .alpha..sub.6 is A, F, or G. In some aspects, when as, is I, .alpha..sub.3 is I or L; .alpha..sub.4 is P; and .alpha..sub.6 is A, F, or G. In some aspects, when .alpha..sub.6, is F, .alpha..sub.3 is I or is absent; .alpha..sub.4 is P or is absent; and .alpha..sub.5 is I or G. In some aspects, when .alpha..sub.6, is G, .alpha..sub.3 is I or L; .alpha..sub.4 is P; and .alpha..sub.5 is I or R.
[0264] In some aspects, when .alpha..sub.3 is L, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is A. In some aspects, when .alpha..sub.3 is I, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is F. In some aspects, when .alpha..sub.3 is absent, .alpha..sub.4 is absent; as is G; and .alpha..sub.6 is F. In some aspects, when .alpha..sub.3 is L, .alpha..sub.4 is P; .alpha..sub.5 is I; and .alpha..sub.6 is G. In some aspects, when .alpha..sub.3 is I, .alpha..sub.4 is P; as is R; and .alpha..sub.6 is G.
[0265] In some embodiments, the antibody comprises a Kabat CDR-H2 sequence defined by the consensus sequence .beta..sub.1-I-P-.beta..sub.5-.beta..sub.6-G-.beta..sub.8-A-N-Y-A-Q-K-F-G- -Q (SEQ ID NOS: 74 and 76-79) where .beta..sub.1 is S or G; .beta..sub.5 is I, E, S, or T; .beta..sub.6 is F, I, or S; and .beta..sub.8 is I or T.
[0266] In some aspects, when .beta..sub.1 is S, .beta..sub.5 is E, I, or S; .beta..sub.6 is I or F; and .beta..sub.8 is I or T. In some aspects, when .beta..sub.1 is G, .beta..sub.5 is I or T; .beta..sub.6 is F or S; and .beta..sub.8 is T. In some aspects, when .beta..sub.5 is I, .beta..sub.1 is G or S; .beta..sub.6 is F or S; and .beta..sub.8 is T. In some aspects, when .beta..sub.6 is F, .beta..sub.1 is G or S; .beta..sub.5 is E, I, or T; and .beta..sub.8 is I or T. In some aspects, when .beta..sub.8 is T, .beta..sub.1 is G or S; .beta..sub.5 is I, S, or T; and .beta..sub.6 is F, I, or S.
[0267] In some aspects, when .beta..sub.1 is S, .beta..sub.5 is I; .beta..sub.5 is F; and .beta..sub.8 is T. In some aspects, when .beta..sub.1 is S, .beta..sub.5 is E; .beta..sub.5 is F; and .beta..sub.8 is I. In some aspects, when .beta..sub.1 is S, .beta..sub.5 is S; .beta..sub.5 is I; and .beta..sub.8 is T. In some aspects, when .beta..sub.1 is G, .beta..sub.5 is I; .beta..sub.5 is F; and .beta..sub.8 is T. In some aspects, when .beta..sub.1 is G, .beta..sub.5 is I; .beta..sub.5 is S; and .beta..sub.8 is T. In some aspects, when .beta..sub.1 is G, .beta..sub.5 is T; .beta..sub.5 is F; and .beta..sub.8 is T.
2.8.5. Kabat CDR-H1 Consensus Sequences
[0268] In some embodiments, the antibody comprises a Kabat CDR-H1 sequence defined by the consensus sequence S-Y-.DELTA..sub.3-M-.DELTA..sub.5 (SEQ ID NOS: 25-29 and 44-45), where .DELTA..sub.3 is E, F, Q, or Y and .DELTA..sub.5 is H or Y.
[0269] In some aspects, when .DELTA..sub.3 is Y, .DELTA..sub.5 is H or Y. In some aspects, when .DELTA..sub.5 is H, .DELTA..sub.3 is E, F, Q, or Y.
[0270] In some aspects, .DELTA..sub.3 is Y when .DELTA..sub.5 H. In some aspects, .DELTA..sub.3 is Y when .DELTA..sub.5 is Y. In some aspects, .DELTA..sub.3 is E when .DELTA..sub.5 is H. In some aspects, .DELTA..sub.3 is Q when .DELTA..sub.5 is H. In some aspects, .DELTA..sub.3 is F when .DELTA..sub.5 is H.
[0271] In some embodiments, the antibody comprises a Kabat CDR-H1 sequence defined by the consensus sequence .theta..sub.1-.theta..sub.2-.theta..sub.3-I-S(SEQ ID NOS: 30-37), where .theta..sub.1 is A, H, K, L, S, or W; .theta..sub.2 is L, M, N, or T; and .theta..sub.3 is A or P.
[0272] In some aspects, when .theta..sub.1 is S, .theta..sub.2 is L or N and .theta..sub.3 is A or P. In some aspects, when .theta..sub.2 is L, .theta..sub.1 is K, L, S, or W and .theta..sub.3 is A or P. In some aspects, when .theta..sub.3 is A, .theta..sub.1 is A, H, K, L, S, or W and .theta..sub.2 is L, M, N, or T.
[0273] In some aspects, .theta..sub.1 is S, when .theta..sub.2 is L and .theta..sub.3 is A. In some aspects, .theta..sub.1 is K, when .theta..sub.2 is L and .theta..sub.3 is A. In some aspects, .theta..sub.1 is H when .theta..sub.2 is T and .theta..sub.3 is A. In some aspects, .theta..sub.1 is S when .theta..sub.2 is L and .theta..sub.3 is P. In some aspects, .theta..sub.1 is L when .theta..sub.2 is L and .theta..sub.3 is A. In some aspects, .theta..sub.1 is S when .theta..sub.2 is N and .theta..sub.3 is A. In some aspects, .theta..sub.1 is A when .theta..sub.2 is M and .theta..sub.3 is A. In some aspects, .theta..sub.1 is W when .theta..sub.2 is L and .theta..sub.3 is A.
[0274] In some embodiments, the antibody comprises a Kabat CDR-H1 sequence defined by the consensus sequence .eta..sub.1-Y-.eta..sub.3-I-S SEQ ID NOS: 38-41), where .eta..sub.1 is S, K, N, or R and .eta..sub.3 is A or G.
[0275] In some aspects, .eta..sub.1 is S where .eta..sub.3 is A or G. In some aspects, .eta..sub.3 is A where .eta..sub.1 is N or S. In some aspects, .eta..sub.3 is G where .eta..sub.1 is K, R, or S.
[0276] In some aspects, when .eta..sub.1 is S, .eta..sub.3 is A. In some aspects, when .eta..sub.1 is S, .eta..sub.3 is G. In some aspects, when .eta..sub.1 is K, .eta..sub.3 is G. In some aspects, when .eta..sub.1 is R, .eta..sub.3 is G. In some aspects, when .eta..sub.1 is N, .eta..sub.3 is A.
2.8.6. CDR-L3 Consensus Sequences
[0277] In some embodiments, the antibody comprises a CDR-L3 sequence defined by the consensus sequence Q-Q-Y-.pi..sub.4-.pi..sub.5-.pi..sub.6-.pi..sub.7-T (SEQ ID NOS: 141-147), where .pi..sub.4 is G, H, or Y; .pi..sub.5 is S, N, F, G, or R; .pi..sub.6 is S, Y, A, G, or R; and .pi..sub.7 is P, I, or L.
[0278] In some aspects, .pi..sub.4 is H when .pi..sub.5 is S, N, G, or R; .pi..sub.6 is Y, A, G, or R; and .pi..sub.7 is I or L. In some aspects, .pi..sub.5 is S, when .pi..sub.4 is G or H; .pi..sub.6 is S, Y, or A; and .pi..sub.7 is P, I or L. In some aspects, .pi..sub.6 is Y, when .pi..sub.4 is H or Y; .pi..sub.5 is S or F; and .pi..sub.7 is I. In some aspects, .pi..sub.6 is A when .pi..sub.4 is H; .pi..sub.5 is N or S; and .pi..sub.7 is I or L. In some aspects, .pi..sub.7 is I when .pi..sub.4 is H or Y; .pi..sub.5 is S, N, F, G, or R; and .pi..sub.6 is Y, A, G, or R.
[0279] In some aspects, .pi..sub.4 is G when .pi..sub.5 is S; .pi..sub.6 is S; and .pi..sub.7 is P. In some aspects, .pi..sub.4 is H when .pi..sub.5 is S; .pi..sub.6 is Y; and .pi..sub.7 is I. In some aspects, .pi..sub.4 is H when .pi..sub.5 is N; .pi..sub.6 is I; and .pi..sub.7 is A. In some aspects, .pi..sub.4 is Y when .pi..sub.5 is F; .pi..sub.6 is Y; and .pi..sub.7 is I. In some aspects, .pi..sub.4 is H when .pi..sub.5 is S; .pi..sub.6 is A; and .pi..sub.7 is L. In some aspects, .pi..sub.4 is H when .pi..sub.5 is G; .pi..sub.6 is G; and .pi..sub.7 is I. In some aspects, .pi..sub.4 is H when .pi..sub.5 is R; .pi..sub.6 is R; and .pi..sub.7 is I.
[0280] In some embodiments, the antibody comprises a CDR-L3 sequence defined by consensus sequence Q-Q-.lamda..sub.3-.lamda..sub.4-.lamda..sub.5-.lamda..sub.6-P-T (SEQ ID NOS: 148-150), where .lamda..sub.3 is R, F, H, S, L, D, Y, or V; .lamda..sub.4 is S, V, T, G, L, Y, or N; .lamda..sub.5 is N, L, F, K, or V; and .lamda..sub.6 is W, F, Y, or L.
[0281] In some aspects, .lamda..sub.3 is R, when .lamda..sub.4 is S or N; .lamda..sub.5 is N or F; and .lamda..sub.6 is W or Y. In some aspects, .lamda..sub.3 is H when .lamda..sub.4 is V or T; .lamda..sub.5 is N or V; and .lamda..sub.6 is F or W. In some aspects, .lamda..sub.3 is S when .lamda..sub.4 is V or Y; .lamda..sub.5 is F; and .lamda..sub.6 is W or L. In some aspects, .lamda..sub.4 is V when .lamda..sub.3 is F, H, S, or D; .lamda..sub.5 is L, N, or F; and .lamda..sub.6 is W or F. In some aspects, .lamda..sub.4 is T when .lamda..sub.3 is L or H; .lamda..sub.5 is K or V; and .lamda..sub.6 is W. In some aspects, .lamda..sub.5 is N when .lamda..sub.3 is R, H, or V; .lamda..sub.4 is S, V, or L; and .lamda..sub.6 is W, F, or Y. In some aspects, .lamda..sub.5 is L when .lamda..sub.3 is F, D, or Y; .lamda..sub.4 is V or G; and .lamda..sub.6 is W or F. In some aspects, .lamda..sub.5 is F when .lamda..sub.3 is S or R; .lamda..sub.4 is V, Y, or N; and .lamda..sub.6 is W, L, or Y. In some aspects, .lamda..sub.6 is W when .lamda..sub.3 is R, F, S, L, D, or H; .lamda..sub.4 is S, V, or T; and .lamda..sub.5 is N, L, F, K, or V. In some aspects, .lamda..sub.6 is F when .lamda..sub.3 is H or Y; .lamda..sub.4 is V or G; and .lamda..sub.5 is N or L. In some aspects, .lamda..sub.6 is Y when .lamda..sub.3 is V or R; .lamda..sub.4 is L or N; and .lamda..sub.5 is N or F.
[0282] In some aspects, .lamda..sub.3 is R when .lamda..sub.4 is S; .lamda..sub.5 is N; and .lamda..sub.6 is W. In some aspects, .lamda..sub.3 is F when .lamda..sub.4 is V; .lamda..sub.5 is L; and .lamda..sub.6 is W. In some aspects, .lamda..sub.3 is H when .lamda..sub.4 is V; .lamda..sub.5 is N; and .lamda..sub.6 is F. In some aspects, .lamda..sub.3 is S when .lamda..sub.4 is V; .lamda..sub.5 is F; and .lamda..sub.6 is W. In some aspects, .lamda..sub.3 is L when .lamda..sub.4 is T; .lamda..sub.5 is K; and .lamda..sub.6 is W. In some aspects, .lamda..sub.3 is D when .lamda..sub.4 is V; .lamda..sub.5 is L; and .lamda..sub.6 is W. In some aspects, .lamda..sub.3 is Y when .lamda..sub.4 is G; .lamda..sub.5 is L; and .lamda..sub.6 is F. In some aspects, .lamda..sub.3 is H when .lamda..sub.4 is T; .lamda..sub.5 is V; and .lamda..sub.6 is W. In some aspects, .lamda..sub.3 is V when .lamda..sub.4 is L; .lamda..sub.5 is N; and .lamda..sub.6 is Y. In some aspects, .lamda..sub.3 is S when .lamda..sub.4 is Y; .lamda..sub.5 is F; and .lamda..sub.6 is L. In some aspects, .lamda..sub.3 is R when .lamda..sub.4 is N; .lamda..sub.5 is F; and .lamda..sub.6 is Y.
[0283] In some embodiments, the antibody comprises a CDR-L3 sequence defined by the consensus sequence Q-Q-Y-.rho..sub.3-.rho..sub.4-W-P-L-T (SEQ ID NOS: 151 and 152), where .rho..sub.3 is N or L and .rho..sub.4 is N or L.
[0284] In some aspects, .rho..sub.3 is N when .rho..sub.4 is L In some aspects, .rho..sub.3 is L when .rho..sub.4 is L.
[0285] In some embodiments, the antibody comprises a CDR-L3 sequence defined by the consensus sequence Q-Q-.omega..sub.3-.omega..sub.4-.omega..sub.5-.omega..sub.6-P-.omega..sub- .8-T (SEQ ID NOS: 153-156), where .omega..sub.3 is Y or F; .omega..sub.4 is Y or W; .omega..sub.5 is S, L, T, or F; .omega..sub.6 is T, Y, or F; and .omega..sub.8 is L or P.
[0286] In some aspects, .omega..sub.3 is Y when .omega..sub.4 is Y or W; .omega..sub.5 is S, L, or T; .omega..sub.6 is T or Y; and .omega..sub.8 is L. In some aspects, .omega..sub.4 is Y when .omega..sub.3 is Y or F; .omega..sub.5 is S, L, or F; .omega..sub.6 is T or Y; and .omega..sub.8 is L or P. In some aspects, .omega..sub.6 is Y when .omega..sub.3 is Y; .omega..sub.4 is Y or W; .omega..sub.5 is L or T; and .omega..sub.6 is L. In some aspects, .omega..sub.8 is L when .omega..sub.3 is Y; .omega..sub.4 is Y or W; .omega..sub.5 is S, L, or T; and .omega..sub.6 is T or Y.
[0287] In some aspects, .omega..sub.3 is Y when .omega..sub.4 is Y; .omega..sub.5 is S; .omega..sub.6 is T; and .omega..sub.8 is L. In some aspects, .omega..sub.3 is Y when .omega..sub.4 is Y; .omega..sub.5 is L; .omega..sub.6 is Y; and .omega..sub.8 is L. In some aspects, .omega..sub.3 is Y when .omega..sub.4 is W; .omega..sub.5 is T; .omega..sub.6 is Y; and .omega..sub.8 is L. In some aspects, .omega..sub.3 is F when .omega..sub.4 is Y; .omega..sub.5 is F; .omega..sub.6 is F; and .omega..sub.8 is P.
2.8.7. CDR-L2 Consensus Sequences
[0288] In some embodiments, the antibody comprises a CDR-L2 sequence defined by the consensus sequence .psi..sub.1-A-S-.psi..sub.4-R-.psi..sub.6-.psi..sub.7 (SEQ ID NOS: 125-136), where .psi..sub.1 is G or Y, .psi..sub.4 is S or N; .psi..sub.6 is A or H; and .psi..sub.7 is T, Y, or N.
[0289] In some aspects, .psi..sub.1 is G when .psi..sub.4 is S or N; .psi..sub.6 is A or H; and .psi..sub.7 is T or N. In some aspects, .psi..sub.1 is Y when is S or N; .psi..sub.6 is A; and .psi..sub.7 is Y or T. In some aspects, .psi..sub.4 is S when .psi..sub.1 is G or Y; .psi..sub.6 is A; and .psi..sub.7 is T, Y, or N. In some aspects, .psi..sub.4 is N when r is G or Y; .psi..sub.6 is H or A; and r is T. In some aspects, .psi..sub.6 is A when .psi..sub.1 is G or Y; .psi..sub.4 is S or N; and .psi..sub.7 is T, Y, or N. In some aspects, .psi..sub.7 is T when .psi..sub.1 is G or Y; .psi..sub.4 is S or N; and .psi..sub.6 is A or H.
[0290] In some aspects, r is G when .psi..sub.4 is S; .psi..sub.6 is A; and .psi..sub.7 is T. In some aspects, .psi..sub.1 is G when .psi..sub.4 is N; .psi..sub.6 is H; and .psi..sub.7 is T. In some aspects, .psi..sub.1 is Y when .psi..sub.4 is S; .psi..sub.6 is A; and .psi..sub.7 is Y. In some aspects, .psi..sub.1 is G when .psi..sub.4 is S; .psi..sub.6 is A; and .psi..sub.7 is N. In some aspects, .psi..sub.1 is Y when .psi..sub.4 is N; .psi..sub.6 is A; and .psi..sub.7 is T.
[0291] In some embodiments, the antibody comprises a CDR-L2 sequence defined by the consensus sequence D-A-S-.chi..sub.4-R-A-T (SEQ ID NOS: 138 and 139), where .English Pound..sub.4 is N or K.
[0292] In some aspects, .chi..sub.4 is N. In some aspects, .chi..sub.4 is K.
[0293] In some embodiments, the antibody comprises a CDR-L2 sequence defined by the consensus sequence W-A-S-T-R-.sigma..sub.6-S(SEQ ID NOS: 131 and 133-134), where .sigma..sub.6 is A, E, or Q.
[0294] In some aspects, .sigma..sub.6 is A. In some aspects, .sigma..sub.6 is E. In some aspects, .sigma..sub.6 is Q.
2.8.8. CDR-L1 Consensus Sequences
[0295] In some embodiments, the antibody comprises a CDR-L1 sequence defined by the consensus sequence .PHI..sub.1-A-S-.PHI..sub.4-.PHI..sub.5-V-.PHI..sub.7-.PHI..sub.8-.PHI..s- ub.9-Y-L-A (SEQ ID NOS: 1101-114), where .PHI..sub.1 is E, K, or R; .PHI..sub.4 is Q or E; .PHI..sub.5 is S or Y; .PHI..sub.7 is S or A; .PHI..sub.8 is S or Y; and .PHI..sub.9 is D or S.
[0296] In some aspects, .PHI..sub.1 is R when .PHI..sub.4 is Q or E; .PHI..sub.5 is S or Y; .PHI..sub.7 is S or A; .PHI..sub.8 is S or Y; and .PHI..sub.9 is S or D. In some aspects, .PHI..sub.4 is E when .PHI..sub.1 is K or R; .PHI..sub.5 is S; .PHI..sub.7 is S; .PHI..sub.8 is S; and .PHI..sub.9 is S. In some aspects, .PHI..sub.4 is Q when .PHI..sub.1 is E or R; .PHI..sub.5 is S or Y; .PHI..sub.7 is S or A; .PHI..sub.8 is S or Y; and .PHI..sub.9 is S or D. In some aspects, .PHI..sub.5 is S when .PHI..sub.1 is E, K, or R; .PHI..sub.4 is E or Q; .PHI..sub.7 is S or A; .PHI..sub.8 is S or Y; and .PHI..sub.9 is S or D. In some aspects, .PHI..sub.7 is S when .PHI..sub.1 is E, K, or R; .PHI..sub.4 is E or Q; .PHI..sub.5 is S or Y; .PHI..sub.8 is S or Y; and .PHI..sub.9 is S or D. In some aspects, .PHI..sub.8 is S when .PHI..sub.1 is K or R; .PHI..sub.4 is E or Q; .PHI..sub.5 is S or 5; .PHI..sub.7 is A or S; and .PHI..sub.9 is S or D. In some aspects, .PHI..sub.8 is R when .PHI..sub.1 is E or R; .PHI..sub.4 is Q; .PHI..sub.5 is S; .PHI..sub.7 is S; and .PHI..sub.9 is S. In some aspects, .PHI..sub.9 is S when .PHI..sub.1 is E, K, or R; .PHI..sub.4 is E or Q; .PHI..sub.5 is S or Y; .PHI..sub.7 is A or S; and .PHI..sub.8 is S or Y.
[0297] In some aspects, .PHI..sub.1 is K when .PHI..sub.4 is E; .PHI..sub.5 is S; .PHI..sub.7 is S; .PHI..sub.8 is S; and .PHI..sub.9 is S. In some aspects, .PHI..sub.1 is E, when .PHI..sub.4 is Q; .PHI..sub.5 is S; .PHI..sub.7 is S; .PHI..sub.8 is Y; and .PHI..sub.9 is S. In some aspects, .PHI..sub.1 is R when .PHI..sub.4 is Q; .PHI..sub.5 is S; .PHI..sub.7 is S; .PHI..sub.8 is S; and .PHI..sub.9 is D. In some aspects, .PHI..sub.1 is R when .PHI..sub.4 is Q; .PHI..sub.5 is S; .PHI..sub.7 is S; .PHI..sub.8 is S; and .PHI..sub.9 is S. In some aspects, .PHI..sub.1 is R when .PHI..sub.4 is Q; .PHI..sub.5 is S; .PHI..sub.7 is A; .PHI..sub.8 is S; and .PHI..sub.9 is S. In some aspects, .PHI..sub.1 is R when .PHI..sub.4 is Q; .PHI..sub.5 is S; .PHI..sub.7 is S; .PHI..sub.8 is Y; and .PHI..sub.9 is S. In some aspects, .PHI..sub.1 is R when .PHI..sub.4 is E; .PHI..sub.5 is S; .PHI..sub.7 is S; .PHI..sub.8 is S; and .PHI..sub.9 is S. In some aspects, .PHI..sub.1 is R when .PHI..sub.4 is Q; .PHI..sub.5 is Y; .PHI..sub.7 is S; .PHI..sub.8 is S; and .PHI..sub.9 is S.
[0298] In some embodiments, the antibody comprises a CDR-L1 sequence defined by the consensus sequence .sub.1-A-S-Q-.sub.5-.sub.6-.sub.7-.sub.8-.sub.9-L-.sub.11 (SEQ ID NOS: 118 and 120-123), where .sub.1 is Q or Rl; .sub.5 is D or S; .sub.6 is I or V; .sub.7 is G or S; .sub.8 is N, R, or S; .sub.9 is N, Y, or W; and .sub.11 is A or N.
[0299] In some aspects, when .sub.1 is R, .sub.5 is S; .sub.6 is I or V; .sub.7 is G or S; .sub.8 is R or S; .sub.9 is N, Y, or W; and .sub.11 is A. In some aspects, when .sub.5 is S, .sub.1 is R; .sub.6 is I or V; .sub.7 is G or S; .sub.8 is R or S; .sub.9 is N, Y, or W; and .sub.11 is A. In some aspects, when .sub.6 is I, .sub.1 is Q or R; .sub.5 is D or S; .sub.7 is S; .sub.8 is N or S; .sub.9 is Y or W; and .sub.11 is A or N. In some aspects, when .sub.6 is V, .sub.1 is R; .sub.5 is S; .sub.7 is G or S; .sub.8 is R or S; .sub.9 is N or W; and .sub.11 is A. In some aspects, when .sub.7 is S, .sub.1 is Q or R; .sub.5 is D or S; .sub.6 is I or V; .sub.8 is N, S, or R; .sub.9 is Y or W; and .sub.11 is A or N. In some aspects, when .sub.8 is S, .sub.1 is R; .sub.5 is S; .sub.6 is I or V; .sub.7 is S; .sub.9 is N, Y, or W; and .sub.11 is A. In some aspects, when .sub.9 is Y, .sub.1 is Q or R; .sub.5 is D or S; .sub.6 is I or V; .sub.7 is S; .sub.8 is N, S, or R; and .sub.11 is A or N. In some aspects, when .sub.11 is A, .sub.1 is R; .sub.5 is S; .sub.6 is I or V; .sub.7 is S or G; .sub.8 is S, or R; and .sub.9 is N, W, or Y.
[0300] In some aspects, when .sub.1 is R, .sub.5 is S; .sub.6 is V; .sub.7 is S; .sub.8 is S; .sub.9 is Y; and .sub.11 is A. In some aspects, when .sub.1 is Q, .sub.5 is D; .sub.6 is I; .sub.7 is S; .sub.8 is N; .sub.9 is Y; and .sub.11 is N. In some aspects, when .sub.1 is R, .sup.5 is S; .sub.6 is V; .sub.7 is S; .sub.8 is R; .sub.9 is Y; and .sub.11 is A. In some aspects, when .sub.1 is R, .sub.5 is S; .sub.6 is V; .sub.7 is G; .sub.8 is S; .sub.9 is N; and .sub.11 is A. In some aspects, when .sub.1 is R, .sub.5 is S; .sub.6 is I; .sub.7 is S; .sub.8 is S; .sub.9 is W; and .sub.11 is A.
[0301] In some embodiments, the antibody comprises a CDR-L1 sequence defined by the consensus sequence K-S-S-.GAMMA..sub.4-S-V-L-.GAMMA..sub.8-S-.GAMMA..sub.10-N-N-K-N-Y-L-A (SEQ ID NOS: 115-117), where .GAMMA..sub.4 is Q, R or K; .GAMMA..sub.8 is F or Y; and .GAMMA..sub.10 is S or N.
[0302] In some aspects, .GAMMA..sub.4 is Q when .GAMMA..sub.8 is F or Y and .GAMMA..sub.10 is S. In some aspects, .GAMMA..sub.8 is F when .GAMMA.4 is Q or R and .GAMMA..sub.10 is S. In some aspects, .GAMMA..sub.8 is Y when .GAMMA..sub.4 is K or Q and .GAMMA..sub.10 is S or N. In some aspects, .GAMMA..sub.10 is S when .GAMMA..sub.4 is R or Q and .GAMMA..sub.8 is F or Y.
[0303] In some aspects, .GAMMA..sub.4 is Q when .GAMMA..sub.8 is Y and .GAMMA..sub.10 is S. In some aspects, .GAMMA..sub.4 is K when .GAMMA..sub.8 is Y and .GAMMA..sub.10 is N. In some aspects, .GAMMA..sub.4 is Q when .GAMMA..sub.8 is F and .GAMMA..sub.10 is S. In some aspects, .GAMMA..sub.4 is R when .GAMMA..sub.8 is F and .GAMMA..sub.10 is S.
3. Germline
[0304] In some embodiments, the antibody that specifically binds CD39 is an antibody comprising a variable region that is encoded by a particular germline gene, or a variant thereof. The illustrative antibodies provided herein comprise variable regions that are encoded by the heavy chain variable region germline genes VH1-46, VH1-69, 1-69, and VH1-46, or variants thereof; and the light chain variable region germline genes VK3-20, VK3-11, VK4-01, VK3, and VK3-15, or variants thereof. One of skill in the art would recognize that the CDR sequences provided herein may also be useful when combined with variable regions encoded by other variable region germline genes, or variants thereof. In particular, the CDR sequences provided herein may be useful when combined with variable regions encoded by variable region germline genes, or variants thereof, that are structurally similar to the variable region germline genes recited above. For example, in some embodiments, a CDR-H sequence provided herein may be combined with a variable region encoded by a variable region germline gene selected from the VH1 or VH3 family, or a variant thereof. In some embodiments, a CDR-L sequence provided herein may be combined with a variable region encoded by a variable region germline gene selected from the V.lamda.3, V.kappa.1, V.kappa.3, and V.kappa.4 families, or a variant thereof.
4. Affinity
[0305] In some embodiments, the affinity of the antibody for CD39, as indicated by K.sub.D, is less than about 10.sup.-5 M, less than about 10.sup.-6 M, less than about 10.sup.-7 M, less than about 10.sup.-8 M, less than about 10.sup.-9 M, less than about 10.sup.-10 M, less than about 10.sup.-11 M, or less than about 10.sup.-12 M. In some embodiments, the affinity of the antibody is between about 10.sup.-7 M and 10.sup.-11 M. In some embodiments, the affinity of the antibody is between about 10.sup.-7 M and 10.sup.-10 M. In some embodiments, the affinity of the antibody is between about 10.sup.-7 M and 10.sup.-9 M. In some embodiments, the affinity of the antibody is between about 10.sup.-7 M and 10.sup.-8 M. In some embodiments, the affinity of the antibody is between about 10.sup.-8 M and 10.sup.-11 M. In some embodiments, the affinity of the antibody is between about 10.sup.-8 M and 10.sup.-10 M. In some embodiments, the affinity of the antibody is between about 10.sup.-9 M and 10.sup.-11 M. In some embodiments, the affinity of the antibody is between about 10.sup.-10 M and 10.sup.-11 M.
[0306] In some embodiments, the affinity of the antibody for human CD39 is between about 4.09.times.10.sup.-7 M and 7.31.times.10.sup.-11 M. In some embodiment, the affinity of the antibody for human CD39 is about 1.14.times.10.sup.-7 M, about 1.31.times.10.sup.-7 M, about 1.67.times.10.sup.-7 M, about 1.43.times.10.sup.-7 M, about 1.30.times.10.sup.-8 M, about 1.27.times.10.sup.-8 M, about 1.13.times.10.sup.-8 M, about 1.60.times.10.sup.-8 M, about 1.34.times.10.sup.-9 M, about 1.16.times.10.sup.-9 M, about 7.31.times.10.sup.-11 M, about 7.60.times.10.sup.-10 M, about 2.66.times.10.sup.-10 M, about 9.22.times.10.sup.-10 M, about 6.72.times.10.sup.-10 M, about 9.24.times.10.sup.-10 M, about 5.58.times.10.sup.-10 M, about 5.48.times.10.sup.-8 M, about 3.37.times.10.sup.-8 M, about 3.11.times.10.sup.-8 M, about 1.88.times.10.sup.-8 M, about 1.63.times.10.sup.-8 M about 1.64.times.10.sup.-8 M, about 1.01.times.10.sup.-8 M, about 2.44.times.10.sup.-7 M, about 4.09.times.10.sup.-7 M, about 3.35.times.10.sup.-8 M, about 1.91.times.10.sup.-8 M, about 1.73.times.10.sup.-8 M, or about 2.39.times.10.sup.-8 M.
[0307] In some embodiments the antibody has a k.sub.on when associating with human CD39 of between about 1.93.times.10.sup.4 M.sup.-1.times.sec.sup.-1 and about 1.72.times.10.sup.6 M.sup.-1.times.sec.sup.-1. In some embodiments the antibody has a k.sub.a when associating with human CD39 of about 6.59.times.10.sup.4 M.sup.-1.times.sec.sup.-1, about 1.93.times.10.sup.4 M.sup.-1.times.sec.sup.-1, about 4.44.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.72.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 6.39.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 8.93.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 9.55.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.11.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 1.17.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.02.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 1.76.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 1.72.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.73.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 1.43.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 9.01.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 3.13.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 5.03.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 3.02.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.73.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 1.78.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.98.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 4.31.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.27.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 3.14.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.81.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 4.73.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 3.26.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 1.73.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.68.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.63.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 3.82.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 2.46.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 3.11.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 4.53.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 4.63.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 9.01.times.10.sup.5 M.sup.-1.times.sec.sup.-1, about 1.03.times.10.sup.6 M.sup.-1.times.sec.sup.-1, about 1.52.times.10.sup.6 M.sup.-1.times.sec.sup.-1, or about 3.53.times.10.sup.5 M.sup.-1.times.sec.sup.-1.
[0308] In some embodiments the antibody has a k.sub.off of about 7.51.times.10.sup.-3 sec.sup.-1, about 6.33.times.10.sup.-2 sec.sup.-1, about 4.70.times.10.sup.-2 sec.sup.-1, about 7.82.times.10.sup.-4 sec.sup.-1, about 4.70.times.10.sup.-2 sec.sup.-1, about 1.05.times.10.sup.-2 sec.sup.-1, about 3.65.times.10.sup.-1 sec.sup.-1, about 1.60.times.10.sup.-1 sec.sup.-1, about 7.11.times.10.sup.-3 sec.sup.-1, about 6.44.times.10.sup.-3 sec.sup.-1, about 3.85.times.10.sup.-2 sec.sup.-1, about 2.30.times.10.sup.-2 sec.sup.-1, about 5.33.times.10.sup.-2 sec.sup.-1, about 9.14.times.10.sup.-2 sec.sup.-1, about 1.80.times.10.sup.-3 sec.sup.-1, about 8.15.times.10.sup.-3 sec.sup.-1, about 3.85.times.10.sup.-4 sec.sup.-1, about 1.34.times.10.sup.-4 sec.sup.-1, about 2.29.times.10.sup.-4 sec.sup.-1, about 4.37.times.10.sup.-3 sec.sup.-1, about 3.71.times.10.sup.-3 sec.sup.-1, about 4.06.times.10.sup.-3 sec.sup.-1, about 6.66.times.10.sup.-2 sec.sup.-1, about 2.02.times.10.sup.-3 sec.sup.-1, about 2.00.times.10.sup.-4 sec.sup.-1, about 5.26.times.10.sup.-3 sec.sup.-1, about 1.13.times.10.sup.-2 sec.sup.-1, about 3.28.times.10.sup.-3 sec.sup.-1, about 2.76.times.10.sup.-3 sec.sup.-1, about 2.86.times.10.sup.-4 sec.sup.-1, about 2.43.times.10.sup.-4 sec.sup.-1, about 2.13.times.10.sup.-4 sec.sup.-1, about 6.09.times.10.sup.-4 sec.sup.-1, about 8.39.times.10.sup.-4 sec.sup.-1, about 8.15.times.10.sup.-3 sec.sup.-1, about 1.32.times.10.sup.-4 sec.sup.-1, about 1.11.times.10.sup.-4 sec.sup.-1, about 2.43.times.10.sup.-4 sec.sup.-1, about 2.13.times.10.sup.-4 sec.sup.-1, about 6.09.times.10.sup.-4 sec.sup.-1, about 8.15.times.10.sup.-3 sec.sup.-1, about 1.32.times.10.sup.-4 sec.sup.-1, or about 1.11.times.10.sup.-4 sec.sup.-1.
[0309] In some aspects, the K.sub.D, k.sub.a, and k.sub.d are determined at 25.degree. C. In some embodiments, the K.sub.D, k.sub.a, and k.sub.d are determined by surface plasmon resonance. In some embodiments, the K.sub.D, k.sub.a, and k.sub.d are determined according to the methods described in the examples.
5. Inhibition of CD39
[0310] In some aspects, the antibody decreases affinity of CD39 to its substrate. In some aspects, the antibody inhibits CD39 function on tumor cells. In some aspects, the antibody inhibits or impedes the release of ADP or AMP from CD39. In some aspects, the antibody inhibits or impedes CD39 processivity.
[0311] In some aspects, the antibody binds CD39 but does not inhibit ATPase. In some aspects, the antibody binds CD39 and inhibits extracellular CD39 activity but not cellular ATPase activity. In some aspects, the antibody binds both the extracellular domain of CD39 and cellular CD39 and can inhibit both the extracellular domain of CD39 and cellular CD39. In some aspects, the antibodies do not compete with A1 and/or others in binding to the extracellular domain.
6. CD39 Assays
[0312] In some embodiments, the antibody binds to an epitope of CD39. In some aspects, CD39 has a sequence identical to the amino acid sequence set forth in SEQ ID NO: 249. In some aspects, the epitope has an amino acid sequence that is identical to the amino acid sequence set forth in SEQ ID NO: 249. In some aspects, the epitope is in an extracellular domain of CD39. In some aspects, the extracellular domain corresponds to all or at least a portion of amino acids 38-478 of SEQ ID NO: 249. In some aspects, the epitope has an amino acid sequence that is 60%, 65%, 70%, 75%, 80%, 85%, 90%, or 95% identical to the sequence set forth in SEQ ID NO: 249 or all or a portion of the extracellular domain. In some aspects, the epitope has a sequence that is identical or corresponds to residues 143-158 and/or residues 274-277 of SEQ ID NO: 249. In some aspects, the epitope is in the region of E143 to N158 on the human CD39 polypeptide having the sequence set forth in SEQ ID NO: 249. In some aspects, the epitope has a sequence that has a 60%, 65%, 70%, 75%, 80%, 85%, 90%, or 95% identity to residues 143-158 or 274-277 of the sequence set forth in SEQ ID NO: 249. In some aspects, the epitope has 1, 2, 3, 4, 5, 6, 7, 8, or 9 substitutions from residues 143-158 of the sequence forth in SEQ ID NO: 249. In some aspects, the epitope has 1, 2 or 3 substitutions from residues 274-277 of SEQ ID NO: 249. In some aspects, the antibody makes contact with any of the residues set forth in FIG. 14E, Table 1. In some aspects, the antibody makes contact with any of the residues set forth in FIG. 14E, Table 2. In some aspects, the antibody binds to D150, E153, and/or R154 or to N99 and none, one, two, or three of D150, E153, and R154 or to any of the above alone or in combination.
[0313] In some aspects, the antibody competes with 1, 2, 3, 4, or 5 of antibodies 27536, 27571, 28347, 27579, or 27597 as set forth in FIG. 14A. In some aspects, the antibody competes with 1, 2, 3, 4, or 5 antibodies 27536, 27571, 28347, 27579, or 27597 as set forth in FIG. 14A. In some aspects, the antibody competes with 1, 2, or 3 of antibodies 25571, 27536, or 27549 as set forth in FIG. 14C. In some aspects, the antibody competes with 1, 2, or 3 of antibodies 25571, 27536, or 27549 set forth in FIG. 14C.
[0314] In some aspects, the antibody inhibits conversion by CD39 of ATP to ADP and/or ADP to AMP. In some aspects, the antibody inhibits platelet aggregation. In some aspects, the antibody decreases or prevents activation of phospho antigen specific T cells selected from MAIT cells and .gamma..delta. T cells. In some aspects, the antibody inhibits angiogenesis. In some aspects, the antibody decreases levels of phosphate, ADP, AMP, and/or adenosine and/or increasing levels of ATP. In some aspects, the antibody increases T effector cell function. In some aspects, the antibody decreases the number of regulatory T cells in tissues or in circulation. In some aspects, the antibody decreases the regulatory T cells or regulatory T cell activity. In some aspects, the antibody increases B cell function. In some aspects, the antibody increases antigen presenting cell function. In some aspects, the antibody inhibits processing of at least one of phospho-antigen from phosphorylated isoprenoid, phosphorylated vitamin B metabolite, and/or phosphorylated riboflavin.
[0315] In some aspects, the antibody has limited ability to limit ATPase of the soluble or extracellular domain. In some aspects, the antibody has limited ability to inhibit ATPase of the cellular and/or extracellular domain of CD39.
7. Glycosylation Variants
[0316] In certain embodiments, an antibody may be altered to increase, decrease or eliminate the extent to which it is glycosylated. Glycosylation of polypeptides is typically either "N-linked" or "O-linked."
[0317] "N-linked" glycosylation refers to the attachment of a carbohydrate moiety to the side chain of an asparagine residue. The tripeptide sequences asparagine-X-serine and asparagine-X-threonine, where X is any amino acid except proline, are the recognition sequences for enzymatic attachment of the carbohydrate moiety to the asparagine side chain. Thus, the presence of either of these tripeptide sequences in a polypeptide creates a potential glycosylation site.
[0318] "O-linked" glycosylation refers to the attachment of one of the sugars N-acetylgalactosamine, galactose, or xylose to a hydroxyamino acid, most commonly serine or threonine, although 5-hydroxyproline or 5-hydroxylysine may also be used.
[0319] Addition or deletion of N-linked glycosylation sites to the antibody may be accomplished by altering the amino acid sequence such that one or more of the above-described tripeptide sequences is created or removed. Addition or deletion of O-linked glycosylation sites may be accomplished by addition, deletion, or substitution of one or more serine or threonine residues in or to (as the case may be) the sequence of an antibody.
[0320] In certain embodiments, the antibody is glycosylated. In certain embodiments, the antibody is deglycosylated. Carbohydrates may be removed by standard techniques. In certain embodiments, the antibody is aglycosylated, for instance by expression in a system that does not glycosylate.
8. Fc Variants
[0321] In certain embodiments, amino acid modifications may be introduced into the Fc region of an antibody provided herein to generate an Fc region variant. In certain embodiments, the Fc region variant possesses some, but not all, effector functions. Such antibodies may be useful, for example, in applications in which the half-life of the antibody in vivo is important, yet certain effector functions are unnecessary or deleterious. Examples of effector functions include complement-dependent cytotoxicity (CDC) and antibody-directed complement-mediated cytotoxicity (ADCC). Numerous substitutions or substitutions or deletions with altered effector function are known in the art.
[0322] An alteration in in CDC and/or ADCC activity can be confirmed using in vitro and/or in vivo assays. For example, Fc receptor (FcR) binding assays can be conducted to measure Fc.gamma.R binding. The primary cells for mediating ADCC, NK cells, express Fc.gamma.RIII only, whereas monocytes express Fc.gamma.RI, Fc.gamma.RII and Fc.gamma.RIII FcR expression on hematopoietic cells is summarized in Ravetch and Kinet, Ann. Rev. Immunol., 1991, 9:457-492.
[0323] Non-limiting examples of in vitro assays to assess ADCC activity of a molecule of interest are provided in U.S. Pat. Nos. 5,500,362 and 5,821,337; Hellstrom et al., Proc. Natl. Acad. Sci. U.S.A., 1986, 83:7059-7063; Hellstrom et al., Proc. Natl. Acad. Sci. U.S.A., 1985, 82:1499-1502; and Bruggemann et al., J. Exp. Med., 1987, 166:1351-1361. Useful effector cells for such assays include peripheral blood mononuclear cells (PBMC) and Natural Killer (NK) cells. Alternatively, or additionally, ADCC activity of the molecule of interest may be assessed in vivo, using an animal model such as that disclosed in Clynes et al. Proc. Natl. Acad. Sci. U.S.A., 1998, 95:652-656.
[0324] C1q binding assays may also be carried out to confirm that the antibody is unable to bind C1q and hence lacks CDC activity. Examples of C1q binding assays include those described in WO 2006/029879 and WO 2005/100402.
[0325] Complement activation assays include those described, for example, in Gazzano-Santoro et al., J. Immunol. Methods, 1996, 202:163-171; Cragg et al., Blood, 2003, 101:1045-1052; and Cragg and Glennie, Blood, 2004, 103:2738-2743.
[0326] FcRn binding and in vivo clearance (half-life determination) can also be measured, for example, using the methods described in Petkova et al., Intl. Immunol., 2006, 18:1759-1769.
9. Preparation of Antibodies
9.1. Antigen Preparation
[0327] The CD39 antigen to be used for production of antibodies may be intact CD39 or a fragment of CD39. The intact CD39, or fragment of CD39, may be in the form of an isolated protein or expressed by a cell. Other forms of CD39 useful for generating antibodies will be apparent to those skilled in the art.
9.2. Monoclonal Antibodies
[0328] Monoclonal antibodies may be obtained, for example, using the hybridoma method first described by Kohler et al., Nature, 1975, 256:495-497, and/or by recombinant DNA methods (see e.g., U.S. Pat. No. 4,816,567). Monoclonal antibodies may also be obtained, for example, using phage or yeast-based libraries. See e.g., U.S. Pat. Nos. 8,258,082 and 8,691,730.
[0329] In the hybridoma method, a mouse or other appropriate host animal is immunized to elicit lymphocytes that produce or are capable of producing antibodies that will specifically bind to the protein used for immunization. Alternatively, lymphocytes may be immunized in vitro. Lymphocytes are then fused with myeloma cells using a suitable fusing agent, such as polyethylene glycol, to form a hybridoma cell. See Goding J. W., Monoclonal Antibodies: Principles and Practice 3.sup.rd ed. (1986) Academic Press, San Diego, Calif.
[0330] The hybridoma cells are seeded and grown in a suitable culture medium that contains one or more substances that inhibit the growth or survival of the unfused, parental myeloma cells. For example, if the parental myeloma cells lack the enzyme hypoxanthine guanine phosphoribosyl transferase (HGPRT or HPRT), the culture medium for the hybridomas typically will include hypoxanthine, aminopterin, and thymidine (HAT medium), which substances prevent the growth of HGPRT-deficient cells.
[0331] Useful myeloma cells are those that fuse efficiently, support stable high-level production of antibody by the selected antibody-producing cells, and are sensitive media conditions, such as the presence or absence of HAT medium. Among these, preferred myeloma cell lines are murine myeloma lines, such as those derived from MOP-21 and MC-11 mouse tumors (available from the Salk Institute Cell Distribution Center, San Diego, Calif.), and SP-2 or X63-Ag8-653 cells (available from the American Type Culture Collection, Rockville, Md.). Human myeloma and mouse-human heteromyeloma cell lines also have been described for the production of human monoclonal antibodies. See e.g., Kozbor, J. Immunol., 1984, 133:3001.
[0332] After the identification of hybridoma cells that produce antibodies of the desired specificity, affinity, and/or biological activity, selected clones may be subcloned by limiting dilution procedures and grown by standard methods. See Goding, supra. Suitable culture media for this purpose include, for example, D-MEM or RPMI-1640 medium. In addition, the hybridoma cells may be grown in vivo as ascites tumors in an animal.
[0333] DNA encoding the monoclonal antibodies may be readily isolated and sequenced using conventional procedures (e.g., by using oligonucleotide probes that are capable of binding specifically to genes encoding the heavy and light chains of the monoclonal antibodies). Thus, the hybridoma cells can serve as a useful source of DNA encoding antibodies with the desired properties. Once isolated, the DNA may be placed into expression vectors, which are then transfected into host cells such as bacteria (e.g., E. coli), yeast (e.g., Saccharomyces or Pichia sp.), COS cells, Chinese hamster ovary (CHO) cells, or myeloma cells that do not otherwise produce antibody, to produce the monoclonal antibodies.
9.3. Humanized Antibodies
[0334] Humanized antibodies may be generated by replacing most, or all, of the structural portions of a monoclonal antibody with corresponding human antibody sequences. Consequently, a hybrid molecule is generated in which only the antigen-specific variable, or CDR, is composed of non-human sequence. Methods to obtain humanized antibodies include those described in, for example, Winter and Milstein, Nature, 1991, 349:293-299; Rader et al., Proc. Nat. Acad. Sci. U.S.A., 1998, 95:8910-8915; Steinberger et al., J. Biol. Chem., 2000, 275:36073-36078; Queen et al., Proc. Natl. Acad. Sci. U.S.A., 1989, 86:10029-10033; and U.S. Pat. Nos. 5,585,089, 5,693,761, 5,693,762, and 6,180,370.
9.4. Human Antibodies
[0335] Human antibodies can be generated by a variety of techniques known in the art, for example by using transgenic animals (e.g., humanized mice). See, e.g., Jakobovits et al., Proc. Natl. Acad. Sci. U.S.A., 1993, 90:2551; Jakobovits et al., Nature, 1993, 362:255-258; Bruggermann et al., Year in Immuno., 1993, 7:33; and U.S. Pat. Nos. 5,591,669, 5,589,369 and 5,545,807. Human antibodies can also be derived from phage-display libraries (see e.g., Hoogenboom et al., J. Mol. Biol., 1991, 227:381-388; Marks et al., J. Mol. Biol., 1991, 222:581-597; and U.S. Pat. Nos. 5,565,332 and 5,573,905). Human antibodies may also be generated by in vitro activated B cells (see e.g., U.S. Pat. Nos. 5,567,610 and 5,229,275). Human antibodies may also be derived from yeast-based libraries (see e.g., U.S. Pat. No. 8,691,730).
10. Vectors, Host Cells, and Recombinant Methods
[0336] The invention also provides isolated nucleic acids encoding anti-CD39 antibodies, vectors and host cells comprising the nucleic acids, and recombinant techniques for the production of the antibodies.
[0337] For recombinant production of the antibody, the nucleic acid encoding it may be isolated and inserted into a replicable vector for further cloning (i.e., amplification of the DNA) or expression. In some aspects, the nucleic acid may be produced by homologous recombination, for example as described in U.S. Pat. No. 5,204,244.
[0338] Many different vectors are known in the art. The vector components generally include, but are not limited to, one or more of the following: a signal sequence, an origin of replication, one or more marker genes, an enhancer element, a promoter, and a transcription termination sequence, for example as described in U.S. Pat. No. 5,534,615.
[0339] Illustrative examples of suitable host cells are provided below. these host cells are not meant to be limiting.
[0340] Suitable host cells include any prokaryotic (e.g., bacterial), lower eukaryotic (e.g., yeast), or higher eukaryotic (e.g., mammalian) cells. Suitable prokaryotes include eubacteria, such as Gram-negative or Gram-positive organisms, for example, Enterobacteriaceae such as Escherichia (E. coli), Enterobacter, Envinia, Klebsiella, Proteus, Salmonella (S. typhimurium), Serratia (S. marcescans), Shigella, Bacilli (B. subtilis and B. licheniformis), Pseudomonas (P. aeruginosa), and Streptomyces. One useful E. coli cloning host is E. coli 294, although other strains such as E. coli B, E. coli X1776, and E. coli W3110 are suitable.
[0341] In addition to prokaryotes, eukaryotic microbes such as filamentous fungi or yeast are also suitable cloning or expression hosts for anti-CD39 antibody-encoding vectors. Saccharomyces cerevisiae, or common baker's yeast, is a commonly used lower eukaryotic host microorganism. However, a number of other genera, species, and strains are available and useful, such as Schizosaccharomyces pombe, Kluyveromyces (K. lactis, K. fragilis, K. bulgaricus K. wickeramii, K. waltii, K. drosophilarum, K. thermotolerans, and K. marxianus), Yarrowia, Pichia pastoris, Candida (C. albicans), Trichoderma reesia, Neurospora crassa, Schwanniomyces (S. occidentalis), and filamentous fungi such as, for example Penicillium, Tolypocladium, and Aspergillus (A. nidulans and A. niger).
[0342] Useful mammalian host cells include COS-7 cells, HEK293 cells; baby hamster kidney (BHK) cells; Chinese hamster ovary (CHO); mouse sertoli cells; African green monkey kidney cells (VERO-76), and the like.
[0343] The host cells used to produce the anti-CD39 antibody of this invention may be cultured in a variety of media. Commercially available media such as, for example, Ham's F10, Minimal Essential Medium (MEM), RPMI-1640, and Dulbecco's Modified Eagle's Medium (DMEM) are suitable for culturing the host cells. In addition, any of the media described in Ham et al., Meth. Enz., 1979, 58:44; Barnes et al., Anal. Biochem., 1980, 102:255; and U.S. Pat. Nos. 4,767,704, 4,657,866, 4,927,762, 4,560,655, and 5,122,469, or WO 90/03430 and WO 87/00195 may be used.
[0344] Any of these media may be supplemented as necessary with hormones and/or other growth factors (such as insulin, transferrin, or epidermal growth factor), salts (such as sodium chloride, calcium, magnesium, and phosphate), buffers (such as HEPES), nucleotides (such as adenosine and thymidine), antibiotics, trace elements (defined as inorganic compounds usually present at final concentrations in the micromolar range), and glucose or an equivalent energy source. Any other necessary supplements may also be included at appropriate concentrations that would be known to those skilled in the art.
[0345] The culture conditions, such as temperature, pH, and the like, are those previously used with the host cell selected for expression, and will be apparent to the ordinarily skilled artisan.
[0346] When using recombinant techniques, the antibody can be produced intracellularly, in the periplasmic space, or directly secreted into the medium. If the antibody is produced intracellularly, as a first step, the particulate debris, either host cells or lysed fragments, is removed, for example, by centrifugation or ultrafiltration. For example, Carter et al. (Bio/Technology, 1992, 10:163-167) describes a procedure for isolating antibodies which are secreted to the periplasmic space of E. coli. Briefly, cell paste is thawed in the presence of sodium acetate (pH 3.5), EDTA, and phenylmethylsulfonylfluoride (PMSF) over about 30 minutes. Cell debris can be removed by centrifugation.
[0347] In some embodiments, the antibody is produced in a cell-free system. In some aspects, the cell-free system is an in vitro transcription and translation system as described in Yin et al., mAbs, 2012, 4:217-225, incorporated by reference in its entirety. In some aspects, the cell-free system utilizes a cell-free extract from a eukaryotic cell or from a prokaryotic cell. In some aspects, the prokaryotic cell is E. coli. Cell-free expression of the antibody may be useful, for example, where the antibody accumulates in a cell as an insoluble aggregate, or where yields from periplasmic expression are low.
[0348] Where the antibody is secreted into the medium, supernatants from such expression systems are generally first concentrated using a commercially available protein concentration filter, for example, an Amicon.RTM. or Millipore.RTM. Pellcon.RTM. ultrafiltration unit. A protease inhibitor such as PMSF may be included in any of the foregoing steps to inhibit proteolysis and antibiotics may be included to prevent the growth of adventitious contaminants.
[0349] The antibody composition prepared from the cells can be purified using, for example, hydroxylapatite chromatography, gel electrophoresis, dialysis, and affinity chromatography, with affinity chromatography being a particularly useful purification technique. The suitability of protein A as an affinity ligand depends on the species and isotype of any immunoglobulin Fc domain that is present in the antibody. Protein A can be used to purify antibodies that are based on human .gamma.1, .gamma.2, or .gamma.4 heavy chains (Lindmark et al., J. Immunol. Meth., 1983, 62:1-13). Protein G is useful for all mouse isotypes and for human .gamma.3 (Guss et al., EMBO J., 1986, 5:1567-1575).
[0350] The matrix to which the affinity ligand is attached is most often agarose, but other matrices are available. Mechanically stable matrices such as controlled pore glass or poly(styrenedivinyl)benzene allow for faster flow rates and shorter processing times than can be achieved with agarose. Where the antibody comprises a C.sub.H3 domain, the BakerBond ABX.RTM. resin is useful for purification.
[0351] Other techniques for protein purification, such as fractionation on an ion-exchange column, ethanol precipitation, Reverse Phase HPLC, chromatography on silica, chromatography on heparin Sepharos.RTM., chromatofocusing, SDS-PAGE, and ammonium sulfate precipitation are also available, and can be applied by one of skill in the art.
[0352] Following any preliminary purification step(s), the mixture comprising the antibody of interest and contaminants may be subjected to low pH hydrophobic interaction chromatography using an elution buffer at a pH between about 2.5 to about 4.5, generally performed at low salt concentrations (e.g., from about 0 to about 0.25 M salt).
11. Pharmaceutical Compositions and Methods of Administration
[0353] Any of the antibodies provided herein can be provided in any appropriate pharmaceutical composition and be administered by any suitable route of administration. Suitable routes of administration include, but are not limited to, the inhalation, intraarterial, intradermal, intramuscular, intraperitoneal, intravenous, nasal, parenteral, pulmonary, and subcutaneous routes.
[0354] The pharmaceutical composition may comprise one or more pharmaceutical excipients. Any suitable pharmaceutical excipient may be used, and one of ordinary skill in the art is capable of selecting suitable pharmaceutical excipients. Accordingly, the pharmaceutical excipients provided below are intended to be illustrative, and not limiting. Additional pharmaceutical excipients include, for example, those described in the Handbook of Pharmaceutical Excipients, Rowe et al. (Eds.) 6th Ed. (2009), incorporated by reference in its entirety.
[0355] In some embodiments, the pharmaceutical composition comprises an anti-foaming agent. Any suitable anti-foaming agent may be used. In some aspects, the anti-foaming agent is selected from an alcohol, an ether, an oil, a wax, a silicone, a surfactant, and combinations thereof. In some aspects, the anti-foaming agent is selected from a mineral oil, a vegetable oil, ethylene bis stearamide, a paraffin wax, an ester wax, a fatty alcohol wax, a long chain fatty alcohol, a fatty acid soap, a fatty acid ester, a silicon glycol, a fluorosilicone, a polyethylene glycol-polypropylene glycol copolymer, polydimethylsiloxane-silicon dioxide, ether, octyl alcohol, capryl alcohol, sorbitan trioleate, ethyl alcohol, 2-ethyl-hexanol, dimethicone, oleyl alcohol, simethicone, and combinations thereof.
[0356] In some embodiments, the pharmaceutical composition comprises a cosolvent. Illustrative examples of cosolvents include ethanol, poly(ethylene) glycol, butylene glycol, dimethylacetamide, glycerin, and propylene glycol.
[0357] In some embodiments, the pharmaceutical composition comprises a buffer. Illustrative examples of buffers include acetate, borate, carbonate, lactate, malate, phosphate, citrate, hydroxide, diethanolamine, monoethanolamine, glycine, methionine, guar gum, and monosodium glutamate.
[0358] In some embodiments, the pharmaceutical composition comprises a carrier or filler. Illustrative examples of carriers or fillers include lactose, maltodextrin, mannitol, sorbitol, chitosan, stearic acid, xanthan gum, and guar gum.
[0359] In some embodiments, the pharmaceutical composition comprises a surfactant. Illustrative examples of surfactants include d-alpha tocopherol, benzalkonium chloride, benzethonium chloride, cetrimide, cetylpyridinium chloride, docusate sodium, glyceryl behenate, glyceryl monooleate, lauric acid, macrogol 15 hydroxystearate, myristyl alcohol, phospholipids, polyoxyethylene alkyl ethers, polyoxyethylene sorbitan fatty acid esters, polyoxyethylene stearates, polyoxylglycerides, sodium lauryl sulfate, sorbitan esters, and vitamin E polyethylene(glycol) succinate.
[0360] In some embodiments, the pharmaceutical composition comprises an anti-caking agent. Illustrative examples of anti-caking agents include calcium phosphate (tribasic), hydroxymethyl cellulose, hydroxypropyl cellulose, and magnesium oxide.
[0361] Other excipients that may be used with the pharmaceutical compositions include, for example, albumin, antioxidants, antibacterial agents, antifungal agents, bioabsorbable polymers, chelating agents, controlled release agents, diluents, dispersing agents, dissolution enhancers, emulsifying agents, gelling agents, ointment bases, penetration enhancers, preservatives, solubilizing agents, solvents, stabilizing agents, and sugars. Specific examples of each of these agents are described, for example, in the Handbook of Pharmaceutical Excipients, Rowe et al. (Eds.) 6th Ed. (2009), The Pharmaceutical Press, incorporated by reference in its entirety.
[0362] In some embodiments, the pharmaceutical composition comprises a solvent. In some aspects, the solvent is saline solution, such as a sterile isotonic saline solution or dextrose solution. In some aspects, the solvent is water for injection.
[0363] In some embodiments, the pharmaceutical compositions are in a particulate form, such as a microparticle or a nanoparticle. Microparticles and nanoparticles may be formed from any suitable material, such as a polymer or a lipid. In some aspects, the microparticles or nanoparticles are micelles, liposomes, or polymersomes. In certain embodiments, a composition provided herein is a pharmaceutical composition or a single unit dosage form. Pharmaceutical compositions and single unit dosage forms provided herein comprise a prophylactically or therapeutically effective amount of one or more prophylactic or therapeutic antibodies.
[0364] Further encompassed herein are anhydrous pharmaceutical compositions and dosage forms comprising an antibody, since water can facilitate the degradation of some antibodies.
[0365] Anhydrous pharmaceutical compositions and dosage forms provided herein can be prepared using anhydrous or low moisture containing ingredients and low moisture or low humidity conditions. Pharmaceutical compositions and dosage forms that comprise lactose and at least one active ingredient that comprises a primary or secondary amine can be anhydrous if substantial contact with moisture and/or humidity during manufacturing, packaging, and/or storage is expected.
[0366] An anhydrous pharmaceutical composition should be prepared and stored such that its anhydrous nature is maintained. Accordingly, anhydrous compositions can be packaged using materials known to prevent exposure to water such that they can be included in suitable formulary kits. Examples of suitable packaging include, but are not limited to, hermetically sealed foils, plastics, unit dose containers (e.g., vials), blister packs, and strip packs.
11.1. Parenteral Dosage Forms
[0367] In certain embodiments, provided are parenteral dosage forms. Parenteral dosage forms can be administered to subjects by various routes including, but not limited to, subcutaneous, intravenous (including bolus injection), intramuscular, and intraarterial. Because their administration typically bypasses subjects' natural defenses against contaminants, parenteral dosage forms are typically, sterile or capable of being sterilized prior to administration to a subject. Examples of parenteral dosage forms include, but are not limited to, solutions ready for injection, dry products ready to be dissolved or suspended in a pharmaceutically acceptable vehicle for injection, suspensions ready for injection, and emulsions.
[0368] Suitable vehicles that can be used to provide parenteral dosage forms are well known to those skilled in the art. Examples include, but are not limited to: Water for Injection USP; aqueous vehicles such as, but not limited to, Sodium Chloride Injection, Ringer's Injection, Dextrose Injection, Dextrose and Sodium Chloride Injection, and Lactated Ringer's Injection; water miscible vehicles such as, but not limited to, ethyl alcohol, polyethylene glycol, and polypropylene glycol; and non-aqueous vehicles such as, but not limited to, corn oil, cottonseed oil, peanut oil, sesame oil, ethyl oleate, isopropyl myristate, and benzyl benzoate.
[0369] Excipients that increase the solubility of one or more of the antibodies disclosed herein can also be incorporated into the parenteral dosage forms.
11.2. Dosage and Unit Dosage Forms
[0370] In human therapeutics, the doctor will determine the posology which he considers most appropriate according to a preventive or curative treatment and according to the age, weight, condition and other factors specific to the subject to be treated.
[0371] The amount of the antibody or composition which will be effective in the prevention or treatment of a disorder or one or more symptoms thereof will vary with the nature and severity of the disease or condition, and the route by which the antibody is administered. The frequency and dosage will also vary according to factors specific for each subject depending on the specific therapy (e.g., therapeutic or prophylactic agents) administered, the severity of the disorder, disease, or condition, the route of administration, as well as age, body, weight, response, and the past medical history of the subject. Effective doses may be extrapolated from dose-response curves derived from in vitro or animal model test systems.
[0372] In certain embodiments, exemplary doses of a composition include milligram or microgram amounts of the antibody per kilogram of subject or sample weight (e.g., about 10 micrograms per kilogram to about 50 milligrams per kilogram, about 100 micrograms per kilogram to about 25 milligrams per kilogram, or about 100 microgram per kilogram to about 10 milligrams per kilogram). In certain embodiment, the dosage of the antibody provided herein, based on weight of the antibody, administered to prevent, treat, manage, or ameliorate a disorder, or one or more symptoms thereof in a subject is 0.1 mg/kg, 1 mg/kg, 2 mg/kg, 3 mg/kg, 4 mg/kg, 5 mg/kg, 6 mg/kg, 10 mg/kg, or 15 mg/kg or more of a subject's body weight. In another embodiment, the dosage of the composition or a composition provided herein administered to prevent, treat, manage, or ameliorate a disorder, or one or more symptoms thereof in a subject is 0.1 mg to 200 mg, 0.1 mg to 100 mg, 0.1 mg to 50 mg, 0.1 mg to 25 mg, 0.1 mg to 20 mg, 0.1 mg to 15 mg, 0.1 mg to 10 mg, 0.1 mg to 7.5 mg, 0.1 mg to 5 mg, 0.1 to 2.5 mg, 0.25 mg to 20 mg, 0.25 to 15 mg, 0.25 to 12 mg, 0.25 to 10 mg, 0.25 mg to 7.5 mg, 0.25 mg to 5 mg, 0.25 mg to 2.5 mg, 0.5 mg to 20 mg, 0.5 to 15 mg, 0.5 to 12 mg, 0.5 to 10 mg, 0.5 mg to 7.5 mg, 0.5 mg to 5 mg, 0.5 mg to 2.5 mg, 1 mg to 20 mg, 1 mg to 15 mg, 1 mg to 12 mg, 1 mg to 10 mg, 1 mg to 7.5 mg, 1 mg to 5 mg, or 1 mg to 2.5 mg.
[0373] The dose can be administered according to a suitable schedule, for example, once, two times, three times, or for times weekly. It may be necessary to use dosages of the antibody outside the ranges disclosed herein in some cases, as will be apparent to those of ordinary skill in the art. Furthermore, it is noted that the clinician or treating physician will know how and when to interrupt, adjust, or terminate therapy in conjunction with subject response.
[0374] Different therapeutically effective amounts may be applicable for different diseases and conditions, as will be readily known by those of ordinary skill in the art. Similarly, amounts sufficient to prevent, manage, treat or ameliorate such disorders, but insufficient to cause, or sufficient to reduce, adverse effects associated with the antibodies provided herein are also encompassed by the herein described dosage amounts and dose frequency schedules. Further, when a subject is administered multiple dosages of a composition provided herein, not all of the dosages need be the same. For example, the dosage administered to the subject may be increased to improve the prophylactic or therapeutic effect of the composition or it may be decreased to reduce one or more side effects that a particular subject is experiencing.
[0375] In certain embodiments, treatment or prevention can be initiated with one or more loading doses of an antibody or composition provided herein followed by one or more maintenance doses.
[0376] In certain embodiments, a dose of an antibody or composition provided herein can be administered to achieve a steady-state concentration of the antibody in blood or serum of the subject. The steady-state concentration can be determined by measurement according to techniques available to those of skill or can be based on the physical characteristics of the subject such as height, weight and age.
[0377] In certain embodiments, administration of the same composition may be repeated and the administrations may be separated by at least 1 day, 2 days, 3 days, 5 days, 10 days, 15 days, 30 days, 45 days, 2 months, 75 days, 3 months, or 6 months. In other embodiments, administration of the same prophylactic or therapeutic agent may be repeated and the administration may be separated by at least 1 day, 2 days, 3 days, 5 days, 10 days, 15 days, 30 days, 45 days, 2 months, 75 days, 3 months, or 6 months.
12. Therapeutic Applications
[0378] For therapeutic applications, the antibodies of the invention are administered to a mammal, generally a human, in a pharmaceutically acceptable dosage form such as those known in the art and those discussed above. For example, the antibodies of the invention may be administered to a human intravenously as a bolus or by continuous infusion over a period of time, by intramuscular, intraperitoneal, intra-cerebrospinal, subcutaneous, intra-articular, intrasynovial, intrathecal, or intratumoral routes. The antibodies also are suitably administered by peritumoral, intralesional, or perilesional routes, to exert local as well as systemic therapeutic effects. The intraperitoneal route may be particularly useful, for example, in the treatment of ovarian tumors.
[0379] The antibodies provided herein may be useful for the treatment of any disease or condition involving CD39, such as cancer, autoimmune disease, and infection.
[0380] Any suitable cancer may be treated with the antibodies provided herein. Illustrative suitable cancers include, for example, acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), adrenocortical carcinoma, anal cancer, appendix cancer, astrocytoma, basal cell carcinoma, brain tumor, bile duct cancer, bladder cancer, bone cancer, breast cancer, bronchial tumor, Burkitt Lymphoma, carcinoma of unknown primary origin, cardiac tumor, cervical cancer, chordoma, chronic lymphocytic leukemia (CLL), chronic myelogenous leukemia (CML), chronic myeloproliferative neoplasm, colon cancer, colorectal cancer, craniopharyngioma, cutaneous T-cell lymphoma, ductal carcinoma, embryonal tumor, endometrial cancer, ependymoma, esophageal cancer, esthesioneuroblastoma, fibrous histiocytoma, Ewing sarcoma, eye cancer, germ cell tumor, gallbladder cancer, gastric cancer, gastrointestinal carcinoid tumor, gastrointestinal stromal tumor, gestational trophoblastic disease, glioma, head and neck cancer, hairy cell leukemia, hepatocellular cancer, histiocytosis, Hodgkin lymphoma, hypopharyngeal cancer, intraocular melanoma, islet cell tumor, Kaposi sarcoma, kidney cancer, Langerhans cell histiocytosis, laryngeal cancer, leukemia, lip and oral cavity cancer, liver cancer, lobular carcinoma in situ, lung cancer, lymphoma, macroglobulinemia, malignant fibrous histiocytoma, melanoma, Merkel cell carcinoma, mesothelioma, metastatic squamous neck cancer with occult primary, midline tract carcinoma involving NUT gene, mouth cancer, multiple endocrine neoplasia syndrome, multiple myeloma, mycosis fungoides, myelodysplastic syndrome, myelodysplastic/myeloproliferative neoplasm, nasal cavity and par nasal sinus cancer, nasopharyngeal cancer, neuroblastoma, non-Hodgkin lymphoma, non-small cell lung cancer, oropharyngeal cancer, osteosarcoma, ovarian cancer, pancreatic cancer, papillomatosis, paraganglioma, parathyroid cancer, penile cancer, pharyngeal cancer, pheochromocytomas, pituitary tumor, pleuropulmonary blastoma, primary central nervous system lymphoma, prostate cancer, rectal cancer, renal cell cancer, renal pelvis and ureter cancer, retinoblastoma, rhabdoid tumor, salivary gland cancer, Sezary syndrome, skin cancer, small cell lung cancer, small intestine cancer, soft tissue sarcoma, spinal cord tumor, stomach cancer, T-cell lymphoma, teratoid tumor, testicular cancer, throat cancer, thymoma and thymic carcinoma, thyroid cancer, urethral cancer, uterine cancer, vaginal cancer, vulvar cancer, and Wilms tumor.
[0381] Any suitable autoimmune disease may be treated with the antibodies provided herein. Illustrative suitable autoimmune diseases, or diseases with an autoimmune component, include, for example, acute disseminated encephalomyelitis (ADEM), acute necrotizing hemorrhagic leukoencephalitis, Addison's disease, agammaglobulinemia, alopecia areata, amyloidosis, ankylosing spondylitis, anti-GBM/anti-TBM nephritis, antiphospholipid syndrome (APS), autoimmune angioedema, autoimmune aplastic anemia, autoimmune dysautonomia, autoimmune hepatitis, autoimmune hyperlipidemia, autoimmune immunodeficiency, autoimmune inner ear disease (AIED), autoimmune myocarditis, autoimmune oophoritis, autoimmune pancreatitis, autoimmune retinopathy, autoimmune thrombocytopenic purpura (ATP), autoimmune thyroid disease, autoimmune urticarial, axonal & neuronal neuropathies, Balo disease, Behcet's disease, bullous pemphigoid, cardiomyopathy, Castleman disease, Celiac disease, Chagas disease, chronic fatigue syndrome, chronic inflammatory demyelinating polyneuropathy (CIDP), chronic recurrent multifocal ostomyelitis (CRMO), Churg-Strauss syndrome, cicatricial pemphigoid/benign mucosal pemphigoid, Crohn's disease, Cogans syndrome, cold agglutinin disease, colitis, congenital heart block, coxsackie myocarditis, CREST disease, essential mixed cryoglobulinemia, demyelinating neuropathies, dermatitis herpetiformis, dermatomyositis, Devic's disease (neuromyelitis optica), discoid lupus, Dressler's syndrome, endometriosis, eosinophilic esophagitis, eosinophilic fasciitis, erythema nodosum, experimental allergic encephalomyelitis, Evans syndrome, fibromyalgia, fibrosing alveolitis, giant cell arteritis (temporal arteritis), giant cell myocarditis, glomerulonephritis, Goodpasture's syndrome, granulomatosis with polyangiitis (GPA) (formerly called Wegener's Granulomatosis), Graves' disease, Guillain-Barre syndrome, Hashimoto's encephalitis, Hashimoto's thyroiditis, hemolytic anemia, Henoch-Schonlein purpura, herpes gestationis, hypogammaglobulinemia, idiopathic thrombocytopenic purpura (ITP), IgA nephropathy, IgG4-related sclerosing disease, immunoregulatory lipoproteins, inclusion body myositis, inflammatory bowel disease. interstitial cystitis, juvenile arthritis, juvenile diabetes (Type 1 diabetes), juvenile myositis, Kawasaki syndrome, Lambert-Eaton syndrome, leukocytoclastic vasculitis, lichen planus, lichen sclerosus, ligneous conjunctivitis, linear IgA disease (LAD), lupus (SLE), Lyme disease (chronic), Meniere's disease, microscopic polyangiitis, mixed connective tissue disease (MCTD), Mooren's ulcer, Mucha-Habermann disease, multiple sclerosis, myasthenia gravis, myositis, narcolepsy, neuromyelitis optica (Devic's), neutropenia, ocular cicatricial pemphigoid, optic neuritis, palindromic rheumatism, PANDAS (Pediatric Autoimmune Neuropsychiatric Disorders Associated with Streptococcus), paraneoplastic cerebellar degeneration, paroxysmal nocturnal hemoglobinuria (PNH), Parry Romberg syndrome, Parsonnage-Turner syndrome, pars planitis (peripheral uveitis), pemphigus, peripheral neuropathy, perivenous encephalomyelitis, pernicious anemia, POEMS syndrome, polyarteritis nodosa, type I, II, & III autoimmune polyglandular syndromes, polymyalgia rheumatic, polymyositis, postmyocardial infarction syndrome, postpericardiotomy syndrome, progesterone dermatitis, primary biliary cirrhosis, rimary sclerosing cholangitis, psoriasis, psoriatic arthritis, idiopathic pulmonary fibrosis, pyoderma gangrenosum, pure red cell aplasia, Raynauds phenomenon, reactive arthritis, reflex sympathetic dystrophy, Reiter's syndrome, relapsing polychondritis, restless legs syndrome, retroperitoneal fibrosis, rheumatic fever, rheumatoid arthritis, sarcoidosis, Schmidt syndrome, scleritis, scleroderma, Sjogren's syndrome, sperm & testicular autoimmunity, stiff person syndrome, subacute bacterial endocarditis (SBE), Susac's syndrome, sympathetic ophthalmia, Takayasu's arteritis, temporal arteritis/giant cell arteritis, thrombotic disease, thrombocytopenic purpura (TTP), Tolosa-Hunt syndrome, transverse myelitis, type 1 diabetes, ulcerative colitis, undifferentiated connective tissue disease (UCTD), uveitis, vasculitis, vesiculobullous dermatosis, vitiligo, and Wegener's granulomatosis (now termed Granulomatosis with Polyangiitis (GPA).
[0382] Any suitable infection may be treated with the antibodies provided herein. Illustrative suitable infections include, for example, hepatitis A virus, hepatitis B virus, hepatitis C virus (HCV), human immunodeficiency virus (HIV), and other viral infections.
13. Diagnostic Applications
[0383] In some embodiments, the antibodies provided herein are used in diagnostic applications. For example, an ant-CD39 antibody may be useful in assays for CD39 protein. In some aspects, the antibody can be used to detect the expression of CD39 in various cells and tissues. These assays may be useful, for example, evaluating cancer and autoimmune disease.
[0384] In some diagnostic applications, the antibody may be labeled with a detectable moiety. Suitable detectable moieties include, but are not limited to radioisotopes, fluorescent labels, and enzyme-substrate labels. In another embodiment of the invention, the anti-CD39 antibody need not be labeled, and the presence thereof can be detected using a labeled antibody which specifically binds to the anti-CD39 antibody.
14. Affinity Purification Reagents
[0385] The antibodies of the invention may be used as affinity purification agents. In this process, the antibodies may be immobilized on a solid phase such a resin or filter paper, using methods well known in the art. The immobilized antibody is contacted with a sample containing the CD39 protein (or fragment thereof) to be purified, and thereafter the support is washed with a suitable solvent that will remove substantially all the material in the sample except the CD39 protein, which is bound to the immobilized antibody. Finally, the support is washed with another suitable solvent, such as glycine buffer, pH 5.0, that will release the CD39 protein from the antibody.
15. Kits
[0386] In some embodiments, an anti-CD39 antibody provided herein is provided in the form of a kit, i.e., a packaged combination of reagents in predetermined amounts with instructions for performing a procedure. In some embodiments, the procedure is a diagnostic assay. In other embodiments, the procedure is a therapeutic procedure.
[0387] In some embodiments, the kit further comprises a solvent for the reconstitution of the anti-CD39 antibody. In some embodiments, the anti-CD39 antibody is provided in the form of a pharmaceutical composition.
EXAMPLES
Example 1: Selection of CD39 Antigen-Binding Proteins
[0388] CD39 ABPs were selected from a synthetic library of human antibodies presented on the surface of yeast cells in IgG format, as generally described, e.g., in WO2009036379; WO2010105256; WO2012009568; and Xu et al., Protein Eng. Des. Sel., 2013, 26:663-670 (each incorporated by reference in its entirety), and more specifically as provided below. The sequences and characteristics of the ABPs isolated from the recombinant library are provided in Table S.
[0389] Eight naive human synthetic yeast libraries each of .sup..about.10E+09 diversity were propagated as described in WO2009036379; WO2010105256; WO2012009568; and Xu et al., Protein Eng. Des. Sel., 2013, 26:663-670; each incorporated by reference in its entirety. For the first two rounds of selection, a magnetic bead sorting technique utilizing the Miltenyi MACS.RTM. system was performed, as described in Siegel et al., J. Immunol. Meth., 2004, 286:141-153. The following rounds of selection were performed using flow cytometry based sorting. For all round of selection, the antigen was biotinylated human CD39 extracellular domain (heretofore "ECD"), and decreasing concentrations of antigen were used in each subsequent round of selection. In addition to selection on antigen, some rounds of selection were employed in order to reduce the number of non-specific binders utilizing soluble membrane proteins from CHO cells (see WO2014179363 and Xu et al., Protein Eng. Des. Sel., 2013, 26:663-670, each incorporated by reference in its entirety). After the final round of sorting, yeast were plated and individual colonies were picked for characterization and for nomination of clones for affinity maturation.
[0390] Antibody variable domains of interest were synthesized, with codon optimization to maximize transient expression in host cells. The variable regions were cloned in to expression vectors containing human immunoglobulin constant domains and their sequence confirmed. Antibody heavy and light chain vector pairings were transfected into Expi293 cells using the Expifectamine system (Invitrogen). Transient cultures were harvested on day 4 and clarified cell culture supernatant IgG titer was estimated using Bio-Layer Interferometry (BLI) using Octet (ForteBio) alongside standards. Antibodies were subsequently purified on a Protein A column and eluted using low pH glycine. Purified antibody samples were then buffer-exchanged or dialyzed into downstream assay-compatible buffers.
[0391] Antibody purity was assessed by running samples on SDS-PAGE and on an analytical size exclusion chromatography column.
[0392] Light Chain Shuffling: Heavy chain plasmids were extracted from naive outputs (described herein) and transformed into a pre-made naive light chain library with a diversity of 10E+06. Selections were performed as described above with one round of MACS sorting and three rounds of FACS sorting using decreasing amounts of biotinylated ECD antigen for respective rounds. Selected individual heavy chains from the primary discovery process were also independently transformed into separate pre-made light chain libraries with a diversity of 10E+06 and selections performed as described above with one round of MACS sorting and three rounds of FACS sorting using decreasing amount of biotinylated ECD antigen for respective rounds.
Example 2: Affinity Maturation
[0393] Optimization of naive clones was carried out utilizing three maturation strategies; diversification of CDR-H1 and CDR-H2; diversification of CDR-H3; diversification of CDR-L1, L2 and L3; shuffling of diversified heavy and light chains.
[0394] CDR-H1 and CDR-H2 Selection: The CDR-H3s from clones selected from each of the light chain batch diversification, light chain diversification, and naive discovery efforts were independently recombined into premade libraries with CDR-H1 and CDR-H2 variants of a diversity of >10E+8 and selections were performed using ECD antigen. Affinity pressures were applied by using decreasing concentrations of antigen.
[0395] CDR-H3 Selection: Clones obtained from the CDR-H1 and CDR-H2 selection procedure were subject to additional rounds of affinity maturation via walking dimer mutagenesis of the heavy chain. Selections were performed using ECD as antigen generally as described above but with the addition of employing FACS sorting for all selection rounds.
[0396] CDR-L1, L2, L3 Selection: Clones obtained from the CDR-H1 and CDR-H2 selection procedure were subject to additional rounds of affinity maturation via mutagenesis of the light chain. The CDR-L1 and CDR-L2 diversity derived from a pre-made library while CDR-L3 diversity derived from walking dimer mutagenesis. Selections were performed using ECD as antigen generally as described above but with the addition of employing FACS sorting for all selection rounds, with one round of MACS followed by three rounds of FACS in the CDR-L1, L2, L3 process described here.
[0397] Diversified Heavy Chain and Light Chain Shuffling: Outputs from heavy chain diversification and light diversification described above were recombined and selections were performed using ECD as antigen generally as described above but with the addition of employing FACS sorting for all selection rounds.
Example 3: Monovalent Affinity of Anti-hCD39 Antibodies to Recombinant CD39 Extracellular Domain
[0398] Binding kinetics were measured using the Octet Red96 system (ForteBio) at 25.degree. C. in running buffer (1.times. Pall ForteBio Kinetics Buffer diluted into PBS or Tris pH 7.4). In brief, 1.25 mg/ml of unlabeled anti-hCD39 antibodies were immobilized onto anti-human Fc sensors. After a short baseline step in running buffer, the sensors were exposed to varying concentrations (10-300 nM) of rhCD39-ECD-His (R&D Systems) for the association step. Dissociation of the complex was monitored upon exposure of the sensors to running buffer once again. Data was processed using ForteBio Octet software with baseline subtraction, global fit and 1:1 binding model to obtain association and dissociation rates. K.sub.D was calculated from the ratio of k.sub.d to k.sub.a.
[0399] Data shown in FIG. 1 had R.sup.2>0.980. PF=poor fit. The association and dissociation time course data was globally fit with a simple 1:1 Langmuir binding model to yield on-rate (kon) and off-rate (koff) values. The equilibrium dissociation constants (K.sub.D) were calculated from the kon and koff values. The kon values ranged from 1.93E+04 to 1.72E+06 M.sup.-1s.sup.-1 and the off rate values ranges from 3.65E-01 to 1.11E-04 s.sup.-1. The K.sub.D values ranged from 4.09E-07 to 7.31E-011 molar indicating that all of the antibodies bound with moderate or high affinity to human CD39 ECD.
[0400] The paralog specificity of the anti-CD39 antibodies was assessed by biolayer interferometry using soluble recombinant human ENTDP2 and soluble recombinant human ENTDP3 (both from R&D Systems). ENTDP2 and ENTDP3 are enzymes with functions similar to CD39. None of the antibodies exhibited detectable binding to ENTDP2 or ENTDP3 (data not shown). Thus all of the antibodies exhibit specific binding to human CD39.
Example 4: Inhibition of Recombinant Human CD39 Extracellular Domain
[0401] The inhibition of recombinant human CD39 ECD by anti-CD39 antibodies was measured as follows. Recombinant human CD39/ENTPD1 (4397-EN from R&D systems), (either 5 or 10 nM final concentration) was combined with anti-CD39 IgGs (0.25 or 1 micromolar final concentration) in 25 mM Tris, 5 mM CaCl.sub.2), pH 7.5 in a 96-well plate and incubated at room temperature for 2 hrs. ATP (Sigma A1852-1VL) was then added to a final concentration of 500 micromolar and incubated at 37.degree. C. for 60 minutes. The plate was then placed at room temperature and CellTiter-Glo Luminescent Cell Viability Assay solution was added to each well of the assay plate, mixed and read on a microplate reader using "CellTiter-Glo luminescent" preset. Control reactions consisting of negative control IgG, IgG only (no ATP), ATP only (no CD39) were run using the same method. Data values are the average of 2 replicates.
[0402] Inhibition of human CD39 ECD enzymatic activity by anti-CD39 antibodies was determined by measuring ATP levels using the CellTiter-Glo assay (FIGS. 2 A-E). The enzymatic catabolism of ATP by CD39 ECD was observed in the presence of an isotype control antibody or no IgG with average RLU values ranging from 38 to 857. All of the anti-CD39 antibodies showed marked inhibition ATP catabolism by CD39, having much higher average RLU values than the isotype control antibody (average RLU values range from 4890 to 20329). In contrast, Benchmark antibody BY40va did not show significant inhibition of CC39 ECD in this assay having RLU values similar to the isotype control antibody (average RLU values 11 and 415).
Example 5: Antibodies Bind to CHO Cells Expressing Human and Cyno CD39
[0403] Binding of anti-CD39 IgGs to Chinese Hamster Ovary K1 (CHO) CD39 cells. 100 nanomolar IgGs (each antibody is indicated as a unique clone number in the FIG.) were incubated at 25.degree. C. for 30 minutes on ice in phosphate buffered saline (PBS) with parental CHO cells or CHO cells engineered to express either human or cynomolgus macaque (Macaca fascicularis) CD39 (CHO CD39 cells). Cells were then washed with ice cold PBS and incubated with a fluorescently labeled goat-anti-human IgG for 20 minutes on ice. Cells were washed and resuspended in ice cold PBS prior to analysis by flow cytrometry. Fold over background binding levels represent the ratio of median fluorescence intensity (MFI) values for anti-CD39 antibodies binding to CHO CD39 to MFI values for anti-CD39 antibodies binding to the parental CHO cells.
[0404] The anti-CD39 antibodies bound to CHO cells expressing cellular human CD39 (CHO CD39 cells) and did not exhibit significant binding to parental CHO cells (FIGS. 2F-J). The binding of these antibodies to CHO CD39 cells ranged from 10 to 2033-fold over background. Antibodies 28337 and 27575 did not show significant binding to CHO CD39 cells (only 2 to 4-fold over background) (FIGS. 2F-J) indicating that these antibodies do not have low affinity for the cellular form of human CD39.
[0405] The ortholog specificity of the anti-CD39 antibodies was assessed with flow cytrometry using CHO cells engineered to express either cynomolgus macaque or mouse CD39. The anti-CD39 antibodies bound to cynomolgus macaque CD39 to a similar extent as human CD39 (FIGS. 2 F-J). None of the anti-CD39 antibodies showed detectable binding to mouse CD39. Thus, the anti-CD39 antibodies are cross reactive to cynomolgus macaque CD39 but not to mouse CD39.
Example 6: Binding of Antibodies to Cell Surface CD39 in MEL-28 or 721 Cells and Antibodies that Inhibit CD39 on MEL-28 Cells a Short Term ATPAse Assay
[0406] Cells were incubated with serially diluted anti-CD39 antibodies for 30 minutes at 4 degrees C. Cells were washed 3 times in FACS buffer (PS, 2% FBS, and 2 mM EDTA) and next incubated with secondary antibody (goat anti-human IgG Southern Biotech) at 1:100 for 30 minutes at 4 degrees C. Cells were washed, resuspended in FACS buffer, and analyzed for binding by flow cytometry analysis on BD Celesta.
[0407] 3.5.times.10.sup.4 MEL-28 cells/well were washed with Tris buffer and incubated with serially diluted (100-0.00013 nM) antibody for 30 minutes at 37 degrees C. 50 .mu.M ATP was added to each well and incubated with cells for 15 minutes. The supernatants were collected and analyzed in Malachite Green Assay (R&D) according to manufacturer's protocol. Phosphate released from CD39 processing of ATP was used as a readout of enzyme activity. Palivizumab was used as an isotype control and ARL (Tocris) and POM-1 (Alpha Aesar), non-specific small molecule inhibitors of CD39, were used as positive controls at 100 .mu.m.
[0408] All antibodies bound to endogenously expressed CD39 on both cell lines with similar affinity with EC50 ranges from 0.05-0.28 .mu.g/ml on MEL-28 (see FIG. 3 A) and with EC50 ranges of 0.2-7.5 .mu.g/ml on 721.22 cell line (see FIG. 3B). Maximum signal (MFI) differed between antibodies tested even when EC50 values were similar (FIGS. 3A-B).
[0409] After confirmation of cellular binding, anti-CD39 antibodies were evaluated for inhibition of ATPase activity on MEL-28 cell in short term 30 minute Malachite Green phosphate readout assay. Isotype control was used to establish maximum possible signal from ATP processing in MEL-28 cells-50 .mu.m ATP addition to cells typically resulted in 55-60 .mu.M phosphate signal in this assay (see FIGS. 4 A-B). Anti-CD39 antibodies inhibited ATPase activity in MEL-28 cells by 60-80% at the highest concentration of antibodies tested (100 nM)--this level of inhibition was similar to non-specific ATPase inhibitors ARL and POM1 (see FIG. 4 A). IC50 values for anti-CD39 antibodies in MEL-28 malachite green assay were all in sub-nanomolar range (see FIG. 4 B).
Example 7: ATP Preservation Quantified when MEL-28 Cells are Treated with CD39 Enzymatic Inhibitors
[0410] 3.5.times.10.sup.4 MEL-28 cells were plated overnight at 37 degrees C. Cells were washed with Tris assay buffer to remove phosphate. 100 nM titrated down to 0.005 pM of monoclonal antibodies were incubated with cells for 30 minutes at 37 degrees C. 50 .mu.M ATP was added and incubated for 15 minutes. Supernates were harvested and frozen. Supernates were thawed and evaluated for ATP using the EnzyLight (EnzyLight ATP Assay Kit, BioAssay Systems). Palivizumab was used as an isotype control and ARL (Tocris) and POM-1 (Alpha Aesar) used at a concentration of 100 .mu.M are non-specific small molecule inhibitors of CD39 as positive controls.
[0411] ATP was almost undetectable after 30 minutes post ATP addition to the cells in untreated and/or isotype treated samples (see FIG. 5 A) while all of the anti-CD39 antibodies tested prevented processing of ATP in dose dependent manner (see FIGS. 5 A-B). Most of the anti-CD39 antibodies tested in this assay prevented ATP processing by CD39 to a similar extent as ARL (see FIG. 5 A). IC50s of anti-CD39 antibodies in ATP preservation assay ranged from 0.02-0.1 nM. Overall potency of antibodies in this assay was consistent with what was observed in Malachite Green phosphate readout assay.
Example 8: Antibodies Inhibit CD39 Activity on MEL-28 in an Overnight Assay
[0412] 3.5.times.10.sup.4 MEL-28 cells/well were plated and incubated with antibodies overnight at 37 degrees C. Cells were washed to remove FBS. Cells next were pre-treated with antibodies in X-VIVO 15 FBS free media overnight at 37 degrees C. ATP was then spiked in at 50 .mu.M for 15 minutes. Supernatants were collected and analyzed using AMP-Glo kit according to manufacturer's instructions (Promega). Palivizumab was used as an isotype control and POM-1 (Alpha Aesar), a non-specific small molecule inhibitor of CD39, was used as positive control at 100 .mu.M.
[0413] Anti-CD39 antibodies tested in overnight AMPGlo assay in MEL-28 cells demonstrated sustained inhibition of ATPase activity as indicated by decreased AMP levels present in the supernatants (see FIG. 6 A). Inhibition of CD39 activity by antibodies was equivalent to or more potent compared to POM-1 treatment. The data is consistent with results obtained in CD39 short-term Malachite Green assay in MEL-28 (see FIG. 4). Antibodies tested in an overnight assay had IC50 values in an AMPGlo CD39 inhibition assay ranging from 0.01 to 0.3 nM. (see FIG. 6 B)
Example 9: Anti-CD 39 Antibodies Bind to Primary Human and Cyno B Cells
[0414] B cells were isolated from human donor leukopak using EasySep B cell isolation kit (STEMCELL Technologies). Cyno monocytes were purified from fresh cyno blood using NHP CD14 positive selection kit (Miltenyi) and flow through was collected and stained with CD4, CD8, CD20, CD16, and CD3 antibodies (BD). Human B cells or cyno cells were incubated with serially diluted anti-CD39 antibodies (15 .mu.g/m17.5 fold serial dilution, 8-point) for 30 minutes at 4 degrees C. Cell were washed 3 times in FACS buffer (PS, 2% FBS, and 2 mM EDTA) and incubated with secondary antibody (mouse anti-human IgG southern biotech) at 1:100 for 30 minutes at 4 degrees C. Cells were washed 2 times in FACS buffer and resuspended in FACS buffer and analyzed on BD Celesta.
[0415] Detection of antibody binding is as described in FIGS. 7 A-B where B cells were incubated with serially diluted antibodies and detected using a fluorescently tagged antibody and analyzed by flow cytometry. The results are shown in FIGS. 7 A-B and appear to indicate that the antibodies bind specifically to both human and cyno B cells with EC50s that range from 0.02 .mu.g/ml to 3.18 .mu.g/ml (human) and 0.03 .mu.g/ml to 0.17 .mu.g/ml (cyno). Similar binding was observed on human tumor cells lines (FIG. 3) where subset of the antibodies had a low maximal MFI and a subset had a high MFI to both the human and cyno B cells.
Example 10: Antibodies Inhibit CD39 Activity on Human B Cells
[0416] B cells were isolated from human leukopak using EasySep B cell isolation kit (STEMCELL Technologies). 5.times.10.sup.4B cells/well were washed with Tris buffer and incubated with serially diluted (100-0.00013 nM) antibodies for 30 minutes at 37 degrees C. 50 .mu.M ATP was added to each well and incubated with cells for 2 hrs. The supernatants were collected and analyzed in Malachite Green Assay (R&D) according to manufacturer's protocol. Phosphate released from CD39 processing of ATP was used as a readout of enzyme activity. Palivizumab was used as an isotype control and ARL (Tocris) and POM-1 (Alpha Aesar), non-specific small molecule inhibitors of CD39, were used as positive controls at 100 .mu.M.
[0417] The antibodies were demonstrated to bind to primary human and cyno B cells (see FIGS. 7 A-B) and the next step was to evaluate the inhibition of ATP hydrolysis by detection of free phosphate (Pi) using a malachite green assay. The results are shown in FIG. 8 and indicate the anti-CD39 antibodies inhibit the enzymatic inhibition/dephosphorylation of ATP by primary human B cells. The ability of the antibodies to inhibit enzymatic activity was comparable regardless of high vs. low max MFI detected in the binding to human B cells (FIGS. 7 A-B).
Example 11: Anti-CD39 Antibodies Inhibit ATPAse Activity on Human and Cyano Monocytes
[0418] Human monocytes were purified from leukopak using EasySep Human monocytes isolation kit (STEMCELL). Cyno monocytes were isolated from whole cyno blood using NHP CD14 positive selection kit (Miltenyi). Monocytes at 5.times.10.sup.4 cells/well were washed with Tris buffer and incubated with serially diluted (100-0.00013 nM) anti-CD39 antibodies for 30 minutes at 37 C. 50 .mu.M ATP was added to the cells for 15 minutes at 37 C and supernatants were harvested and analyzed in Malachite Green Assay (R&D) for phosphate levels. Paliviz.mu.tmab was used as an isotype control and ARL (Tocris) and POM-1 (Alpha Aesar), non-specific small molecule inhibitors of CD39, were used as positive controls at 100 .mu.M.
[0419] CD39 expression has been detected on human leukocytes with the highest expression detected on monocytes (Thromb Res. 2007; 121(3):309-17). Because of this information, it was important to evaluate the ability of the anti-CD39 antibodies to inhibit ATPase activity on the cell surface. As demonstrated in FIGS. 7 A-B, anti-CD39 antibodies bind to both human and cyno B cells. It is appropriate to evaluate the inhibition of enzymatic activity on human and cyno monocytes. The results indicate that all the antibodies are able to inhibit ATPase activity of CD39 on human and cyno monocytes with similar potencies.
Example 12: Anti-CD39 Antibodies Bind to Primary Human TRegs and Inhibit CD39 Enzymatic Activity
[0420] Treg cells were isolated from human donor leukopak using CD4.sup.+CD25.sup.+CD127.sup.dim regulatory T cell isolation kit II (Miltenyi). Human Treg cells were incubated with serially diluted anti-CD39 antibodies (100 nM-0.00064 nM) for 30 minutes at 4 degrees C. Cell were washed 3 times in FACS buffer (PS, 2% FBS, and 2 mM EDTA) and incubated with secondary antibody (mouse anti-human IgG southern biotech) at 1:100 for 30 minutes at 4 degrees C. Cells were washed 2 times in FACS buffer, resuspended in FACS buffer and analyzed on BD Fortessa.
[0421] CD4.sup.+CD25.sup.+CD127.sup.dim human Treg cells were washed 3.times. with Tris buffer. Cells were incubated with anti-CD39 antibodies (100 nM-0.00064 nM) for 30 minutes at 37 C. The cells were spiked with 50 .mu.M ATP and supernatants were collected after 15 minutes incubation at 37 C. Supernatants were analyzed for phosphate levels in Malachite Green Assay kit (R&D). Palivizumab was used as an isotype control and ARL (Tocris) and POM-1 (Alpha Aesar), non-specific small molecule inhibitors of CD39, were used as positive controls at 100 .mu.M.
[0422] CD39 had been shown to be expressed on human regulatory T cells (Treg) and important for their suppressive function by hydrolysis of ATP to immune suppressive adenosine. (Blood. 2007 Aug. 15; 110(4):1225-32, Cellular & Molecular Immunology (2017) 14, 521-528; doi:10.1038/cmi.2016.30). In order to determine whether the anti-CD39 antibodies were capable of inhibiting CD39 enzymatic activity, it was important to evaluate the binding to human Tregs. Tregs were isolated from human PBMCs and the Tregs were purified and the anti-bodies were evaluated for binding by flow cytometry. The result indicate that the anti-CD39 antibodies bind to human Tregs (FIG. 10). Of note, both the high maximum MFI and low MFI profiles were observed similar to the human B cell staining (see FIG. 8). The ability of the antibodies to inhibit the ATPase activity on human Tregs was also evaluated. All the antibodies inhibit Treg CD39 enzymatic activity (see FIG. 11) equally well regardless of the maximal MFI staining observed.
Example 13: Anti-CD39 Antibodies Increase CD8.sup.+ T Cell Response in a CMV Recall Response Assay
[0423] Frozen PBMCs are thawed and resuspended at 3.times.10.sup.6/ml and cultured in presence of CMV peptides (Miltenyi, PeptTivator CMV pp65) for 3 days in complete (10% FBS) media at 37 degrees C. T cells were then purified (STEMCELL, EasyStep) and rested for 24 hours at 37 degrees C. APCs were generated by depleting CD2 positive cells (STEMCELL, Easystep) from PBMCs from the same donor and plated at 5.times.10.sup.4/well overnight at 37 degrees C. The following day 5.times.10.sup.4 rested T cells were added to the APCs. Antibodies were added at 25 .mu.g/ml plus 100.mu. M ATP+1 .mu.M EHNA and Golgi plug/stop with CMV peptides and incubated at 37 degrees C. for 5 hours. T cells were stained and analyzed for intracellular IFN gamma on a Fortessa (Becton Dickinson) flow cytometer.
[0424] Adenosine has been shown to inhibit T cell activation. (Int J Oncol. 2008 March; 32(3):527-35). As the rate-limiting enzyme in ATP/ADP-AMP-adenosine pathway, inhibiting CD39 would diminish the levels of immune suppressive adenosine and increase immune activating ATP resulting in enhanced T cell activity. In order to evaluate the role of anti-CD39 antibodies inhibiting ATP/ADP-AMP hydrolysis, preventing the generation of adenosine which could result in an increased T cell response, a CMV recall assay was used. PBMCs were cultured in the presence of a pool of CMV peptides for 3 days and then the T cells were purified cultured with autologous PBMC plus CMV peptides and ATP+EHNA. After 5 hours the T cells were evaluated for the production of IFN gamma. The results indicate that some of the anti-CD39 antibodies were able to increase CD8.sup.+ T cells activity in a CMV recall responses assay. This demonstrates that inhibiting the enzymatic activity of CD39 prevents the generation of adenosine and preserves ATP levels and allowing for a robust T cell response to a peptide:MHC complex. In addition, not all antibodies that bind to CD39 and inhibit cell surface enzymatic activity are capable of increasing T cell response and the specific interaction of the antibody to CD39 is important.
Example 14: Evaluation of Antibodies for Inhibition of CD39 Activity on Mel-28 and Human Monocytes
[0425] 3.5.times.10.sup.4 MEL-28 cells/well were incubated with 100 nM down to 0.32 pM of monoclonal antibodies for 30 minutes. 50 .mu.M ATP was added and incubated for 15 minutes. Supernate was evaluated for free phosphate (Pi) using the Malachite Green Phosphate Detection Kit (R&D Systems cat#DY996). Palivizumab was used as an isotype control and ARL (Tocris) and POM-1 (Alpha Aesar) used at a concentration of 100 .mu.M as non-specific small molecule inhibitors of CD39 as positive controls. Cyno monocytes were isolated from whole cyno blood using NHP CD14 positive selection kit (Miltenyi). Monocytes at 5.times.10.sup.4 cells/well were washed with Tris buffer and incubated as described above.
[0426] When comparing anti-CD39 antibodies for enzymatic inhibition in short term assays, differences were observed. Some were able to block the release of free phosphate (Pi) measured in a malachite green assay and others demonstrated very little activity. For example, BY40v9 and 9-8B have little to no enzymatic inhibition at concentrations of 100 nM using both the MEL-28 cell or primary human monocyte compared to other anti-CD39 antibodies (e.g., 29579 and 28347) that are able to inhibit ATPase activity at low nM concentrations.
[0427] 9-8B was produced as described in US 2017/0335007 A1, using SEQ ID Nos. 22 and 23, as hIgG4. BY40v9 is an engineered variant of the antibody BY40 described in WO 2009/095478 A1 SEQ ID Nos. 1 and 5. We tried to express BY40 as described as a human IgG4 but repeated attempts failed. The VH described in SEQ No. 1 appears to be missing several N-terminal residues compared to germline VH's, so we engineered in the missing residues with closest germline sequence from IMGT (http://imgt.org/), resulting in BY40-v9, which expressed and was determined to specifically bind rhCD39-ECD and cellular ECD.
Example 15: Design of Chimera and Region of Distinct Binding
[0428] (a) Examples of Antibodies that Bind Soluble Recombinant CD39 ECD and Cellular CD39 but do not Inhibit ATPase Activity and do not Compete with Cellular Inhibitors for Binding to ECD Inhibitors
[0429] 0.04-3.3 nM rhCD39-ECD (R&D Systems) was incubated with buffer, 50 .mu.g/ml antibody or 100 .mu.M POM-1 (Alpha Aesar) for 1 hr at 37.degree. C. in assay buffer (25 mM Tris pH 7.5, 5 mM CaCl.sub.2) at which point ATP (Sigma) was spiked in to a final concentration of 10 .mu.M, and the reaction further incubated for 30 min 37.degree. C. Production of free phosphate (Pi) was subsequently measured using Malachite Green Phosphate Detection kit (R&D Systems).
[0430] Indicated cells were incubated in assay buffer with 10-25 .mu.g/ml antibody, 100 .mu.M POM-1 or 100 .mu.M ARL for 30 min at 37.degree. C. 5% CO.sub.2. ATP was added to a final concentration of 50 .mu.M and the incubation continued for 15 min. Supernatant was evaluated for free phosphate (Pi) using the Malachite Green Phosphate Detection kit. Palivizumab is used as an isotype control and POM-1 and ARL are non-specific small molecule inhibitors of CD39.
[0431] A1 antibody was immobilized onto an Anti-Mouse IgG Fc Capture (AMC) biosensor (ForteBio). Association of hCD39-ECD was then monitored for 180 seconds via Bio-Layer Interferometry (BLI) using the Octet system (ForteBio), at which point the biosensor was dipped into competitor antibody and monitored for another 180 seconds. Association of the second antibody was recorded as an upward shift in the interference pattern and indicates that the antibodies bind to different epitopes on CD39. No change in the interference pattern indicated that A1 blocks the second antibody from binding to CD39.
[0432] Although A1 does bind hCD39 ECD (FIG. 1), it does not inhibit its ATPase activity. Although A1 does bind cellular CD39 (FIG. 2), it does not appreciably directly inhibit the ATPase activity of CD39 expressed on OAW42. Capture of hCD39 ECD by A1 blocks subsequent binding by A1 (FIG. 14A, bottom right hand sensorgram), but does not block binding of inhibitory antibodies such as 27536, 27571, 27579, 27597, or 38347.
[0433] The commercially available A1 antibody represents a group of anti-human CD39 monoclonal antibodies that do not directly inhibit the ATPase activity of CD39 and do not bin with any of the other anti-CD39 antibodies described here. More than 30 antibodies were discovered that do not inhibit ECD ATPase activity, do not inhibit cellular CD39 ATPase activity, but do compete with A1 for binding to ECD. These antibodies may be considered to bin with anti-hCD39 antibody A1.
(b) Example of Antibodies that have Limited Ability to Inhibit the ATPase Activity of Both Soluble Recombinant and Cellular CD39 and Bin Separately from Other Cellular CD39 Inhibitors
[0434] The methods are the same as set forth above except that instead of A1, the representative antibody was immobilized using anti-human IgG Fc Capture (AHC) biosensor.
[0435] Anti-hCD39 antibodies exemplified by 27536 and 28337 represent a group of antibodies that bind both soluble ECD and cellular CD39 and have the ability to inhibit their ATPase activity yet do not compete with other inhibitors for CD39 binding. Both 27536 and 28337 are able to inhibit the hydrolysis of ATP by hCD39 ECD compared to isotype or buffer controls as shown by .mu.M Pi, as described herein (top left panel). 27536 and 28337 inhibit the hydrolysis of CD39 expressed on MEL-28 and OAW42 cells, as shown, compared to isotype control. Antibodies 27536 and 28337 represent a distinct bin group of anti-hCD39 inhibitory antibodies since they are blocked from binding CD39 ECD by another bin member that does not block other inhibitory antibodies such as 27571, 27579, 27597, or 38347 (see insert (c) at the bottom of FIG. 14B).
(c) Example of Antibodies that Inhibit the ATPase Activity of Soluble Recombinant CD39ECD but do not Inhibit Cellular CD39 and Bin Separately from Other CD39 ECD Inibitors
[0436] The methods are as provided above, with the addition of experimental design where inhibition of 0.37 nM CD39 by 50 .mu.g/ml antibody was challenged by ATP concentrations of 3-100 .mu.M. Also, an 18 hour incubation was used with the antibody or isotype control. Finally, an additional 30 second baseline dip in buffer between antigen capture and dipping into a second competitor antibody for 300 seconds was used.
[0437] Anti-hCD39 antibodies exemplified by 27549 represent a group of antibodies that can bind ECD and cellular CD39 but can only inhibit ECD ATPase activity and not cellular CD39 ATPase activity. As can be seen in FIG. 14C (top left), 27549 is able to inhibit the hydrolysis of ATP by hCD39 ECD compared to isotype or buffer controls. Also from FIG. 14C (top right), 27549 is unable to inhibit the hydrolysis of CD39 expressed on MEL-28 cells compared to isotype control. Antibodies 27536 and 28337 represent a distinct bin group of anti-hCD39 inhibitory antibodies since they are blocked from binding CD39 ECD by another bin member that does not block other inhibitory antibodies such as 27571, 27579, 27597, or 38347 (see FIG. 14C, bottom).
[0438] Antibody 27549 represents a group of anti-human CD39 monoclonal antibodies that directly inhibit the ATPase activity of sol CD39 ECD yet does inhibit cellular CD39 despite being able to bind to CD39 expressed on cells (FIG. 2). It does not compete with A1 or with any of the other inhibitory anti-CD39 antibodies described here for binding to ECD, so 27549 represents another bin of anti-hCD39 antibodies (see FIG. 14C).
(d) Example of Antibodies that Inhibit the ATPase Activity of ECD and Cellular CD39 and Bin Separately from Other CD39 ECD and/or Cellular Inhibitors
[0439] The methods are as provided above.
[0440] Antibodies 27571, 27579 and 28347 represent a group of anti-human CD39 antibodies that directly inhibit the ATPase activity of soluble CD39 ECD and inhibit cellular CD39 and yet do not compete with A1, cellular inhibitors 27536 or 28337, or ECD inhibitor 27549 (FIGS. 14A-C) for binding to ECD. 27571, 27579, and 28347 are able to inhibit the hydrolysis of ATP by hCD39 ECD compared to isotype or buffer controls (see FIG. 14D, left). 27571, 27579, and 28347 also inhibit the hydrolysis of CD39 expressed on MEL-28 cells compared to isotype control (see FIG. 14D, right). These cellular inhibitors represent another bin of anti-hCD39 antibodies (see FIG. 14D).
(e) Examples of Inhibitory Antibodies that Make Distinct Contacts with CD39
[0441] Chimeras were generated by replacing human CD39 with mouse CD39 sequence in mammalian expression vectors. The chimeras were expressed in CHO cells and the ability of anti-human CD39 antibodies to bind the chimeras tested via FACS. In brief, cells were washed and blocked in FACS buffer (PBS/2% FBS) for 30 min on ice. They were then incubated with 15 .mu.g/ml anti-human CD39 antibodies diluted in FACS buffer.times.1 h on ice. After 2 washes, cells were incubated with fluorescently labeled anti-human Fc antibodies, anti-mouse Fc antibodies, or anti-mCD39 antibody (R&D Systems 495826) diluted in FACS buffer per manufacturers' instructions.times.30 min on ice. After 2 washes, cells were resuspended in FACS buffer and analyzed on BD Fortessa. Data was processed with FlowJo. Positive binding was scored as "yes" and lack of binding was scored as "NO" and examples of FACS plots used to generate this table are in FIGS. 14.F and 14.G.
[0442] Chimeras formed between human and mouse CD39 distinguish regions critical for antibody contact that are unique to inhibitory antibodies such as 31414, 31895, 31873, 31901, 31905, reference antibodies such as BY40v9 and 9-8B, and commercial antibodies such as A1 and 498403. Columns 1 and 3 describe the human CD39 sequences flanking the mouse CD39 sequence described in column 2. Column 4 lists the exact mouse amino sequence in the chimera.
[0443] The antibodies described thus far have no/minimal cross-reactivity to mouse CD39, which shares 78% percent identity with human CD39 in the extracellular domain. Chimeras were therefore made between human and mouse CD39 in order to identify those region(s) critical to antibody recognition and binding. To that end, 8 chimeras were generated with the sequence swaps chosen based on sequence diversity between human and mouse CD39 and potential for surface exposure based on rat ENTPD1 and rat ENTPD2 crystal structures. Thus, these 8 chimeras do not comprehensively interrogate all potential contact residues. All tested antibodies were able to bind Chimeras #1,3,5,6,7, and 8, suggesting that these chimeras maintained overall global structural integrity. A1 and 498403 lost the ability to bind Chimera #4, indicating that residues critical to their contact with human CD39 had been lost. A1, 498403, BY40v9 and 9-8B were able to bind Chimera #2, suggesting that this chimera maintained overall global structural integrity. In contrast, inhibitory antibodies 31414, 31895, 31901, 31905, and 31873 were able to bind all chimeras but Chimera #2. Thus, Chimera #2 has lost residues critical to making contacts with these antibodies and therefore E143-N158 constitute part or all of the human CD39 epitope for these antibodies. Notably, these antibodies belong to the bin group exemplified by 27571, 27579 and 28347 in FIG. 14D, i.e. they do not bin with anti-CD39 antibody bin groups represented by antibodies A1, 27536, or 27549.
[0444] Chimeras were generated by mutagenesis of Chimera #2 or WT hCD39 expression vectors. Residues for mutagenesis were chosen based on divergence between human and mouse CD39. The chimeras were expressed in CHO cells and the ability of anti-human CD39 antibodies to bind the chimeras tested via FACS, as described in FIG. 14E. Table 2. Positive binding was scored as "Yes" and lack of binding was scored as "No" and examples of FACS plots used to generate this table are in FIGS. 14.F and 14.G.
[0445] Specific residues critical for antibody contact can distinguish inhibitory antibodies 31414, 31418, and 31895 from 31901, 31905, and 31873. Individual amino acids in the mouse sequence of human-mouse Chimera #2, when reverted back to the respective human residue (resulting in Chimera #9,10), restore the ability of antibodies 31895, 31414, 31418 to bind to CD39 but not the ability of 31901, 31905, or 31873 to bind CD39. Mutation of individual residues in the context of otherwise fully human CD39 (Chimeras #11,12) did not impair the binding ability of any of the tested antibodies. Co-mutation of two human residues to the cognate mouse residues (Chimeras #13,14) did not impair binding by antibodies 31414, 31418 or 31895. In stark contrast, antibodies 31901, 31905, and 31873 completely lost their ability to bind Chimeras #13 and 14. The ability of indicated antibodies to bind a given chimera was determined by FACS. In particular, FIG. 14F and FIG. 14G show representative FACS plots used to populate this table and FIG. 14G shows the importance of human CD39 residues N99, R154, and/or S153 in recognition by antibodies represented by 31901, 31905, and 31873.
[0446] As demonstrated in FIG. 14E, Table 1, inhibitory antibodies 31414, 31418, 31895, 31901, 31905 and 31873 can be distinguished from other monoclonal anti-CD39 antibodies based on their inability to recognize Chimera #2. In Table 2, these antibodies are further distinguished by the gain or loss of the ability to bind specific point mutants of hCD39 or Chimera #2. Thus, these antibodies, while binding the same general region of human CD39, form distinct contacts with different critical residues in hCD39. Since these antibodies share the requirement for residues E143-N158, they may share some common contact residues yet can be differentiated based on the demonstrated unique points of contact.
(f) FACS Plots Highlighting the Importance of Human CD39 Residue D150 and/or E153 in Recognition of Antibodies Represented by 31414, 31418, and 31895 and FACS Plots Highlighting the Importance of Human CD39 Residues N99, R154, and/or E153 in Recognition by Antibodies Represented by 31901, 31905, and 31873
[0447] The methods are as provided above for the examples of inhibitory antibodies that make distinct contacts with CD39.
[0448] The ability of indicated antibodies to bind to a given chimera was determined by FACS.
[0449] While antibodies 31414, 31418, 31895, 31873, 31901, and 31905 all share a common requirement for human CD39 residues E143-N158, these antibodies make different critical contacts with residues within and outside of this region. Thus, antibodies, which belong to the bin group represented, by antibodies 27571, 27579, and 28347 (FIG. 14.D) can be further distinguished into at least two more groups based on sensitivity to distinct residues. There are at least 5 bin groups: (1) A1-like antibodies that compete each other for ECD binding yet do not inhibit ECD activity, but do not compete with antibodies in bin groups 2, 3, 4, or 5 for ECD binding and make critical contacts with N275-1277 as evidenced by loss of binding to Chimera #4; (2) 27536/28337-like antibodies that compete each other for ECD binding and can inhibit both ECD and cellular CD39 ATPase activity, but do not compete with antibodies in bin groups 1, 3, 4 or 5 for ECD binding; (3) 27549-like antibodies that compete each other for ECD binding and can inhibit CD39 ECD but not cellular CD39, and do not compete with antibodies in bin groups 1, 2, 4 or 5 for ECD binding; (4) 31414/31418/31895-like antibodies that make critical contacts with E143-N158 in cellular CD39 including (but not limited to) D150 and E153, inhibit the ATPase activity of both ECD and cellular CD39, compete with each other for ECD binding, but do not compete with antibodies in bins 1, 2 or 3 for ECD binding; and (5) 31873/31901/31905-like antibodies that make critical contacts with E143-N158 in cellular CD39 including (but not limited to) E153 and R154, as well as a sensitivity to residue N99, inhibit the ATPase activity of both ECD and cellular CD39, compete with each other for ECD binding, but do not compete with antibodies in bins 1, 2, or 3 for ECD binding. Thus, there are anti-human CD39 antibodies that can be distinguished based on their affinity for ECD, affinity for cells, ability to inhibit ECD ATPase activity, ability to inhibit cellular ATPase activity, and points of contact with CD39. For example, 27549 binds both ECD (FIG. 1) and cells well (FIG. 2) yet can only inhibit ECD ATPase activity. 27579 binds ECD weakly but is a potent cellular CD39 inhibitor.
Example 16: Anti-CD39 Antibodies are Reversible Allosteric, not Competitive, Inhibitors Due to V.sub.max Suppression of V.sub.max
[0450] Anti-CD39 antibodies or isotype control antibody (100 nanomolar final concentration) were incubated with MEL-28 cells (35,000 MEL-28 cells/well) endogenously expressing human CD39 at 37.degree. C. for 20 minutes in the presence of EnzChek reagents (PNP & MESG). Immediately following the addition of ATP (final concentrations ranging from 0-450 micromolar), the rate of ATP hydrolysis to free phosphate (Pi) by CD39 was monitored over time at Abs360 nm using SpectraMax i3x plate reader. The initial enzyme velocity, .nu.0, was determined from the linear region of Pi vs. time curve for each ATP concentration. The plot of .nu.0 vs. [ATP] was curve fit using non-linear regression of the Michaelis-Menten kinetic model.
[0451] The ATP hydrolysis rate by CD39 expressed on MEL-28 cells can be markedly reduced by the anti-CD39 antibodies (see FIG. 15). The ATP hydrolysis rate in the presence of the anti-CD39 antibodies ranges from 1.2 to 2.7 micromolar Pi per minute, which is much less than the ATP hydrolysis rate observed in the presence of an isotype control antibody (7.7 micromolar Pi per minute) and similar in magnitude to the pan ATPase inhibitor ARL (1.2 micromolar Pi per minute).
[0452] The plot of initial velocity versus ATP concentration for multiple concentrations of the anti-CD39 antibody 29872 indicates that the mechanism of inhibition of this IgG antibody is not competitive (see FIG. 16 A.). The reduction in initial velocity is constant for ATP concentrations above 100 micromolar (100, 200, 300 and 500 micromolar) in the presence of 5 nanomolar 29872. 29872 could be a non-competitive inhibitor, an un-competitive inhibitor, or a mixed inhibitor of CD39 (having properties of both non-competitive and un-competitive inhibition). The monovalent Fab form of anti-CD39 antibody 29872 showed a very similar inhibition profile (see FIG. 16 B), indicating that the CD39 inhibition activity of 29872 is not dependent on the bivalent structure of the IgG and the potential properties that could result from IgG bivalency such as CD39 crosslinking.
[0453] Many of the other anti-CD39 antibodies tested in this assay showed a similar profile for CD39 enzymatic inhibition (data not shown), indicating that they also may be non-competitive inhibitors, un-competitive inhibitors, or mixed inhibitors of CD39. The mechanism of enzymatic inhibition of these antibodies suggests that they retain their full inhibition of CD39 enzymatic inhibition at high ATP concentrations. The anti-CD39 antibodies appear to be reversible inhibitors.
Example 17: Anti-CD39 can Induce Internalization of CD39 on Cyno Monocytes
[0454] Anti-CD39 antibodies were injected into cynomolgus monkeys to test for ability to downregulate CD39 on cell surface of cyno monocytes. CD39 internalization was assessed by FACS of whole blood samples collected pre-dose, Day 1, Day 7, and Day 14 after antibody treatment. Non-competing anti-CD39-PE antibody (clone A1) was used to measure CD39 levels on CD14+ gated monocytes. Data is shown as MFI of A1-PE on gated cyno monocytes in FIG. 17.
[0455] Cynomolgus monkeys were injected with anti-CD39 antibodies at 10 mg/kg. Blood samples were collected prior to injection and on Day 1, Day 7, and Day 14 after treatment. For each time-point, 50 uL of cynomolgus whole blood was incubated with 30 uL of staining buffer (PBS, 2% FBS, 2 mM EDTA, 3% mouse serum, 5% goat serum) containing CD14 and CD20 antibodies (eBioscience) and an Fc blocking reagent (BD) for 30 minutes at 4 C. CD39 antibody (clone eBioA1, eBioscience) was added to the sample and incubated for an additional 40 minutes at 4 degrees C. Following incubation, the samples were treated with ACK lysis buffer (ThermoFisher) for 10 minutes at room temperature to lyse red blood cells. Samples were washed several times with staining buffer and fixed with 1% PFA (Sigma) before acquisition on a BD Fortessa X-20 flow cytometer. Total CD39 receptor expression on CD14 positive monocytes was determined by mean fluorescence intensity.
[0456] CD39 was highly expressed on cyno monocytes prior to antibody treatment in all blood samples tested (pre-dose). Anti-CD39 antibodies tested in vivo had distinct internalization profiles where antibody 1 treatment led to downregulation of CD39 on cell surface of monocytes and antibody 2 had no effect on overall CD39 levels. Apparent decrease of CD39 levels after antibody 1 treatment was not due to A1 antibody competing with antibody 1 for binding to CD39 as the epitopes for A1 and antibody 1 are distinct from each other.
Example 18: Anti-CD39 Antibody Increases Stimulated Human CD4.sup.+ and CD8.sup.+ T Cell in the Presence of Exogenous ATP
[0457] Anti-CD3+anti-CD28 stimulated PBMCs were treated with anti-CD39 antibody 31895 or isotype control in the presence of 50 .mu.M ATP for 96 hours. Proliferation of CD4.sup.+ and CD8.sup.+ T cells was measured by Cell Trace Violet by flow cytometry.
[0458] FIG. 19 shows that anti-CD39 antibody increases proliferation of stimulated human CD4.sup.+ and CD8.sup.+ T cell in the presence of exogenous ATP. The left side shows CD8.sup.+ T cells, with the x-axis showing antibody (nM) and the y-axis showing % CD8.sup.+ T cell proliferation. The right side shows CD4.sup.+ T cells, with the x-axis showing antibody (nM) and the y-axis showing % CD4.sup.+ T cell proliferation. The inset to the right shows symbols for the respective antibodies and controls.
Example 19: Anti-CD39 Antibody Increase Stimulated PBMC Secretion of INF-.gamma., TNF-.alpha. and IL-2
[0459] Human PBMCs were treated with anti-CD3+anti-CD28 and incubated with anti-CD39 antibody 31895 or isotype control in presence (B) or absence (A) of exogenous ATP (50 .mu.M). Supernatants were harvested after 96 hours and cytokines were measured using a Meso Scale Discovery human cytokine kit.
[0460] FIG. 20 shows anti-CD39 antibody 31895 increases cytokine secretion by anti-CD3+anti-CD28 activated PBMC in absence (A) or presence (B) of exogenous ATP in a dose dependent manner. The top row shows results with exogenous ATP added and the bottom row shows results with no exogeonous ATP added. The x-axis shows antibody (nM) and the y-axis shows secretion of INF-.gamma., TNF-.alpha. and IL-2, respectively.
Example 20: Anti-CD39 Antibody Increases Stimulated PBMC Secretion of INF-.gamma., TNF-.alpha., IL-2 and IL-1.beta.
[0461] Human PBMCs were stimulated with anti-CD3+anti-CD28 and incubated with anti-CD39 antibodies 31895, HAO-391 (See, SEQ ID 10/SEQ ID 11 from WO 2017/089334), HAO mAb4 (See SEQ ID 12/SEQ ID 13 from WO2017157948) or isotype control at a fixed concentration of 50 .mu.g/ml in presence of ATP. Supernatants were harvested after 96 hours and analyzed by Meso Scale Discovery human cytokine kit.
[0462] FIG. 21 shows anti-CD39 antibody 31895 increased cytokine release by activated PBMCs to a higher degree compared to anti-CD39 antibodies HAO-391 (VL SEQ ID No. 10; VH SEQ ID No. 11 from WO 2017/089334) and HAO mAb4 (VL SEQ ID No. 12; VH SEQ ID No. 13 from WO2017157948). The x-axis indicates the andtibody and/or conditions and the y-axis shows INF-.gamma., TNF-.alpha., IL-2 and IL-1.beta., respectively.
Example 21: Anti-39 Antibody 31895 Increase Extracellular ATP Accumulation and Reduces Adenosine Generation by CD39.sup.+CD73.sup.+SK-MEL-28 Cells
[0463] SK-MEL-28 cells were treated with 31895, isotype control, or small molecule inhibitors (EHNA, ARL, or POM-1) for 1 hour and 50 .mu.M ATP was added for 15 min prior to harvesting of the supernatants. The supernatants were analyzed for ATP levels using AmpGlo Kit (A) and for adenosine levels using LC/MS analysis (B). % Adenosine levels were normalized to isotype control (100%) and SK-MEL-28 CD39 KO cells (0%).
[0464] The results are shown in FIG. 22. The left graph shows ATP accumulation and the right graph shows adenosine generation (LC/MC). The x-axis for each graph shows conditions. The y-axis for the left graph shows ATP (.mu.M) and the y-axis for the right graph shows % of adenosine levels.
Example S: Sequences
[0465] Table S provides sequences referred to herein.
TABLE-US-00003 TABLE S Sequences. SEQ Scheme/ ID NO: Region Clone Sequence 1 CDR-H1 Chothia GYTFTSY 2 CDR-H1 Chothia GYTFKSY 3 CDR-H1 Chothia GYIFKSY 4 CDR-H1 Chothia GYTFQSY 5 CDR-H1 Chothia GYTFFSY 6 CDR-H1 Chothia GYTFVSY 7 CDR-H1 Chothia GGTFSSLAIS 8 CDR-H1 Chothia GGTFSKLAIS 9 CDR-H1 Chothia GGTFSHT 10 CDR-H1 Chothia GGTFSSL 11 CDR-H1 Chothia GGTFSLL 12 CDR-H1 Chothia GGTFQSL 13 CDR-H1 Chothia GGTFPSN 14 CDR-H1 Chothia GGTFSAM 15 CDR-H1 Chothia GGTFASL 16 CDR-H1 Chothia GGTFSWL 17 CDR-H1 Chothia GGTFSSY 18 CDR-H1 Chothia GGTFGSY 19 CDR-H1 Chothia GGTFSKY 20 CDR-H1 Chothia GGTFGRY 21 CDR-H1 Chothia GGTFESY 22 CDR-H1 Chothia GGTFSNY 23 CDR-H1 Chothia GGAFSSY 24 CDR-H1 Chothia GFTFSSY 25 CDR-H1 Kabat SYYMH 26 CDR-H1 Kabat SYEMH 27 CDR-H1 Kabat SYQMH 28 CDR-H1 Kabat SYYMY 29 CDR-H1 Kabat SYFMH 30 CDR-H1 Kabat SLAIS 31 CDR-H1 Kabat KLAIS 32 CDR-H1 Kabat HTAIS 33 CDR-H1 Kabat SLPIS 34 CDR-H1 Kabat LLAIS 35 CDR-H1 Kabat SNAIS 36 CDR-H1 Kabat AMAIS 37 CDR-H1 Kabat WLAIS 38 CDR-H1 Kabat SYAIS 39 CDR-H1 Kabat SYGIS 40 CDR-H1 Kabat KYGIS 41 CDR-H1 Kabat NYAIS 42 CDR-H1 Kabat SYATS 43 CDR-H1 Kabat SYAIG 44 CDR-H1 Kabat SYSMN 45 CDR-H1 Kabat SYGMN 46 CDR-H2 Chothia NPSGGST 47 CDR-H2 Chothia NPSVGS 48 CDR-H2 Chothia NPSGGS 49 CDR-H2 Chothia NPLGGG 50 CDR-H2 Chothia NPRGGS 51 CDR-H2 Chothia IPIFGT 52 CDR-H2 Chothia GFGT 53 CDR-H2 Chothia LPIGGT 54 CDR-H2 Chothia LPIAGT 55 CDR-H2 Chothia LPIFGE 56 CDR-H2 Chothia IPRGGT 57 CDR-H2 Chothia IPEFGI 58 CDR-H2 Chothia IPSIGT 59 CDR-H2 Chothia IPISGT 60 CDR-H2 Chothia IPTFGT 61 CDR-H2 Chothia SSSSSY 62 CDR-H2 Chothia WYDGSN 63 CDR-H2 Kabat VINPSGGSTSYAQKFQG 64 CDR-H2 Kabat RINPSVGSTWYAQKFQG 65 CDR-H2 Kabat RINPSGGSTWYAQKFQG 66 CDR-H2 Kabat KINPSGGSTWYAQKFQG 67 CDR-H2 Kabat VINPLGGGTSYAQKFQG 68 CDR-H2 Kabat SINPRGGSTSYAQKFQG 69 CDR-H2 Kabat GIIPIFGTANYAQKFQG 70 CDR-H2 Kabat GI--GFGTANYAQKFQG 71 CDR-H2 Kabat GILPIGGTANYAQKFQG 72 CDR-H2 Kabat GILPIAGTANYAQKFQG 73 CDR-H2 Kabat GILPIFGEANYAQKFQG 74 CDR-H2 Kabat GIIPRGGTANYAQKFQG 75 CDR-H2 Kabat SIIPIFGTANYAQKFRG 76 CDR-H2 Kabat SIIPEFGIANYAQKFQG 77 CDR-H2 Kabat SIIPIFGTANYAQKFQG 78 CDR-H2 Kabat GIIPISGTANYAQEFQG 79 CDR-H2 Kabat GIIPTFGTANYAQKFQG 80 CDR-H2 Kabat SISSSSSYIYYADSVKG 81 CDR-H2 Kabat VIWYDGSNKYYADSVKG 82 CDR-H3 GKREGGTEYLRH 83 CDR-H3 GKREGGTEYLRK 84 CDR-H3 GKREGGTEYLRS 85 CDR-H3 GKREGGTEYLRN 86 CDR-H3 GKREGGTEYLRV 87 CDR-H3 GGAKYASTYGMDV 88 CDR-H3 GGAKYASTHGMDV 89 CDR-H3 GGAKYASQLGMDV 90 CDR-H3 GGAKYASKWGMDV 91 CDR-H3 GGAKYAVGYGMDV 92 CDR-H3 GGAKYAGRYGMDV 93 CDR-H3 GGAKYARTYGMDV 94 CDR-H3 ESGGYRDHRLDV 95 CDR-H3 ESGTYRDHRLDV 96 CDR-H3 ESGGYRDHRLGV 97 CDR-H3 DFTDYSSGYSSGWTY 98 CDR-H3 DTLYSSGAYYGYNV 99 CDR-H3 AKRGYDSYGGVYFDY 100 CDR-H3 GPTVTATTSIGTHNWFDP 101 CDR-H3 EGRGYDSSRYYKFWFDPWGQGTLVTVSS 102 CDR-H3 DGGGYRHHYFDL 103 CDR-H3 ESGGYRDHKLDV 104 CDR-H3 DGGGYQHHYFDL 105 CDR-H3 DSGYHRHYSDY 106 CDR-H3 DPLGIRKHWFDP 107 CDR-H3 DTPRWRYHYFDY 108 CDR-H3 ERRGSLALGMDV 109 CDR-H3 DLGGYSYGEPYYYYYGMDV 110 CDR-L1 RASQSVSSSYLA 111 CDR-L1 RASQSVASSYLA 112 CDR-L1 EASQSVSYSYLA 113 CDR-L1 KASESVSSSYLA 114 CDR-L1 RASQYVSSSYLA 115 CDR-L1 KSSQSVLFSSNNKNYLA 116 CDR-L1 KSSRSVLFSSNNKNYLA 117 CDR-L1 KSSKSVLYSNNNKNYLA 118 CDR-L1 RASQSVGSNLA 119 CDR-L1 KSSQSVLYSSNNKNYLA 120 CDR-L1 QASQDISNYLN 121 CDR-L1 RASQSVSSYLA 122 CDR-L1 RASQSVSRYLA
123 CDR-L1 RASQSISSWLA 124 CDR-L1 RASQSVSSDYLA 125 CDR-L2 GASSRAT 126 CDR-L2 GASNRHT 127 CDR-L2 YASSRAY 128 CDR-L2 GASSRAN 129 CDR-L2 YASSRAT 130 CDR-L2 YASNRAT 131 CDR-L2 WASTRES 132 CDR-L2 WASSRES 133 CDR-L2 WASTRQS 134 CDR-L2 WASTRAS 135 CDR-L2 GASTRAT 136 CDR-L2 GASTRAS 137 CDR-L2 DASNLET 138 CDR-L2 DASNRAT 139 CDR-L2 DASKRAT 140 CDR-L2 KASSLES 141 CDR-L3 QQYHSYIT 142 CDR-L3 QQYHNAIT 143 CDR-L3 QQYYFYIT 144 CDR-L3 QQYHSALT 145 CDR-L3 QQYHGGIT 146 CDR-L3 QQYHRRIT 147 CDR-L3 QQYHSGIT 148 CDR-L3 QQYYLYPLT 149 CDR-L3 QQYWTYPLT 150 CDR-L3 QQYLLYPLT 151 CDR-L3 QQYLIWPLT 152 CDR-L3 QQYLLWPLT 153 CDR-L3 QQFYFFPPT 154 CDR-L3 QQAYTFPPT 155 CDR-L3 QQYYIFPPT 156 CDR-L3 QQRNFYPPT 157 CDR-L3 QQFVLWPRT 158 CDR-L3 QQHVNFPLT 159 CDR-L3 QQSVFWPIT 160 CDR-L3 QQLTKWPLT 161 CDR-L3 QQDVLWPLT 162 CDR-L3 QQYGLFPIT 163 CDR-L3 QQHTVWPIT 164 CDR-L3 QQVLNYPLT 165 CDR-L3 QQSYFLPPT 166 CDR-L3 QQAHSSPYT 167 Leader for Leader MKYLLPTAAAGLLLLAAQPAMA scFV, scFv-Fc 168 Linker for Linker GGGGSGGGGSGGGGS scFV, scFV-FC 169 C-Term Tag for C-Term Tag GPGGQHHHHHH scFV, scFV-FC 170 C-Term Tag for C-Term Tag PKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKD scFv, scFV-FC TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGV EVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLN GKEYKCKVSNKALPAPIEKTISKAKGQPREPQV YTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEW ESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDK SRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK 171 scFv 29872 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KEPGASVKVSCKAPGYTFTSYYMHWVRQAPGQG LEWMGVINPSGGSTSYAQKFQGRVIMIRDTSTS TVYMELSSLRSEDTAVYYCARGKREGGTEYLRH WGQGTLVTVSSGGGGSGGGGSGGGGSEIVLTQS PGTLSLSPGERATLSCRASQSVSSSYLAWYQQK PGQAPRLLIYGASSRATGIPDRFSGSGSGTDFT LTISRLEPEDFAVYYCQQYHSYITFGGGTKVEI KGPGGQHHHHHH 172 scFv 31895 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KKPGASVKVSCKASGYTFKSYEMHWVRQAPGQG LEWMGRINPSVGSTWYAQKFQGRVTMTRDTSTS TVYMELSSLRSEDTAVYYCARGKREGGTEYLRK WGQGTLVTVSSGGGGSGGGGSGGGGSEIVLTQS PGTLSLSPGERATLSCRASQSVASSYLAWYQQK PGQAPRLLIYGASNRHTGIPDRFSGSGSGTDFT LIISRLEPEDFAVYYCQQYHNAITFGGGTKVEI K GPGGQHHHHHH 173 scFv 31414 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KKPGASVKVSCKASGYTFKSYEMHWVRQAPGQG LEWMGRINPSVGSTWYAQKFQGRVTMTRDTSTS TVYMELSSLRSEDTAVYYCARGKREGGTEYLRN WGQGTLVTVSSGGGGSGGGGSGGGGSEIVLTQS PGTLSLSPGERATLSCRASQSVSSSYLAWYQQK PGQAPRLLIYGASSRATGIPDRFSGSGSGTDFT LTISRLEPEDFAVYYCQQYHSYITFGGGTKVEI KGPGGQHHHHHH 174 scFv 31905 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KKPGSSVKVSCKASGGTFPSNAISWVRQAPGQG LEWMGGIGFGTANYAQKFQGRVTITADESTSTA YMELSSLRSEDTAVYYCARGGAKYARTYGMDVW GQGTTVTVSSGGGGSGGGGSGGGGSDIVMTQSP DSLAVSLGERATINCKSSKSVLYSNNNKNYLAW YQQKPGQPPKLLIYWASTRQSGVPDRFSGSGSG TDFTLTISSLQAEDVAVYYCQQYLLYPLTFGGG TKVEIKGPGGQHHHHHH 175 scFv-Fc 29872 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KEPGASVKVSCKAPGYTFTSYYMHWVRQAPGQG LEWMGVINPSGGSTSYAQKFQGRVIMIRDTSTS TVYMELSSLRSEDTAVYYCARGKREGGTEYLRH WGQGTLVTVSSGGGGSGGGGSGGGGSEIVLTQS PGTLSLSPGERATLSCRASQSVSSSYLAWYQQK PGQAPRLLIYGASSRATGIPDRFSGSGSGTDFT LTISRLEPEDFAVYYCQQYHSYITFGGGTKVEI KPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPK DTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQ VYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVD KSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPG K 176 scFv-Fc 31895 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KKPGASVKVSCKASGYTFKSYEMHWVRQAPGQG LEWMGRINPSVGSTWYAQKFQGRVTMTRDTSTS TVYMELSSLRSEDTAVYYCARGKREGGTEYLRK WGQGTLVTVSSGGGGSGGGGSGGGGSEIVLTQS PGTLSLSPGERATLSCRASQSVASSYLAWYQQK PGQAPRLLIYGASNRHTGIPDRFSGSGSGTDFT LIISRLEPEDFAVYYCQQYHNAITFGGGTKVEI KPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPK DTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQ VYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVD KSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPG K 177 scFv-Fc 31414 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KKPGASVKVSCKASGYTFKSYEMHWVRQAPGQG LEWMGRINPSVGSTWYAQKFQGRVTMTRDTSTS TVYMELSSLRSEDTAVYYCARGKREGGTEYLRN WGQGTLVTVSSGGGGSGGGGSGGGGSEIVLTQS PGTLSLSPGERATLSCRASQSVSSSYLAWYQQK PGQAPRLLIYGASSRATGIPDRFSGSGSGTDFT LTISRLEPEDFAVYYCQQYHSYITFGGGTKVEI KPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPK DTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDG VEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQ VYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVE WESNGQPENNYKTTPPVLDSDGSFFLYSKLTVD KSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPG K 178 scFv-Fc 31905 MKYLLPTAAAGLLLLAAQPAMAQVQLVQSGAEV KKPGSSVKVSCKASGGTFPSNAISWVRQAPGQG LEWMGGIGFGTANYAQKFQGRVTITADESTSTA YMELSSLRSEDTAVYYCARGGAKYARTYGMDVW GQGTTVTVSSGGGGSGGGGSGGGGSDIVMTQSP DSLAVSLGERATINCKSSKSVLYSNNNKNYLAW YQQKPGQPPKLLIYWASTRQSGVPDRFSGSGSG TDFTLTISSLQAEDVAVYYCQQYLLYPLTFGGG TKVEIKPKSCDKTHTCPPCPAPELLGGPSVFLF PPKPKDTLMISRIPEVTCVVVDVSHEDPEVKFN WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVL HQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQ PREPQVYTLPPSREEMTKNQVSLTCLVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLYS KLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSL SLSPGK 179 VH 27579 QVQLVQSGAEVKEPGASVKVSCKAPGYTFTSYY MHWVRQAPGQGLEWMGVINPSGGSTSYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSS 180 VH 31895 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSVGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRKWGQGTLVTVSS 181 VH 31415 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYQ MHWVRQAPGQGLEWMGRINPSGGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRSWGQGTLVTVSS 182 VH 31414 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSVGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRNWGQGTLVTVSS 183 VH 31891 QVQLVQSGAEVKKPGASVKVSCKASGYIFKSYE MHWVRQAPGQGLEWMGRINPSVGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRVWGQGTLVTVSS 184 VH 29871 QVQLVQSGAEVKKPGASVKVSCKASGYTFQSYY MHWVRQAPGQGLEWMGKINPSGGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSS 185 VH 31418 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSGGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSS 186 VH 31431 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYQ MHWVRQAPGQGLEWMGRINPSGGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG
KREGGTEYLRHWGQGTLVTVSS 187 VH 31421 QVQLVQSGAEVKKPGASVKVSCKASGYTFFSYY MYWVRQAPGQGLEWMGVINPLGGGTSYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSS 188 VH 31429 QVQLVQSGAEVKKPGASVKVSCKASGYTFVSYF MHWVRQAPGQGLEWMGSINPRGGSTSYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSS 189 VH 29872 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSVGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSS 190 VH 28347 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSLA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTNTAYMELSSLRSEDTAVYYCARG GAKYASTYGMDVWGQGTTVTVSS 191 VH 31896 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSKLA ISWVRQAPGQGLEWMGGIGFGTANYAQKFQGRV TITADESASTAYMELSSLRSEDTAVYYCARGGA KYASTHGMDVWGQGTTVTVSS 192 VH 31432 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSHTA ISWVRQAPGQGLEWMGGILPIGGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARG GAKYASQLGMDVWGQGTTVTVSS 193 VH 31915 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSLP ISWVRQAPGQGLEWMGGIGFGTANYAQKFQGRV TITADESTSTAYMELSSLRSEDTAVYYCARGGA KYASKWGMDVWGQGTTVTVSS 194 VH 31436 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSLLA ISWVRQAPGQGLEWMGGILPIAGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARG GAKYAVGYGMDVWGQGTTVTVSS 195 VH 31437 QVQLVQSGAEVKKPGASVKVSCKASGGTFQSLA ISWVRQAPGQGLEWMGGILPIGGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARG GAKYAGRYGMDVWGQGTTVTVSS 196 VH 31905 QVQLVQSGAEVKKPGSSVKVSCKASGGTFPSNA ISWVRQAPGQGLEWMGGIGFGTANYAQKFQGRV TITADESTSTAYMELSSLRSEDTAVYYCARGGA KYARTYGMDVWGQGTTVTVSS 197 VH 31901 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSLP ISWVRQAPGQGLEWMGGIGFGTANYAQKFQGRV TITADESTSTAYMELSSLRSEDTAVYYCARGGA KYAGRYGMDVWGQGTTVTVSS 198 VH 29852 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSAMA ISWVRQAPGQGLEWMGGILPIAGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARG GAKYASTYGMDVWGQGTTVTVSS 199 VH 29851 QVQLVQSGAEVKKPGSSVKVSCKASGGTFASLA ISWVRQAPGQGLEWMGGILPIFGEANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARG GAKYASTYGMDVWGQGTTVTVSS 200 VH 29857 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSWLA ISWVRQAPGQGLEWMGGIIPRGGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARG GAKYASTYGMDVWGQGTTVTVSS 201 VH 27571 QVQLVQSGAEVKKPGSSVKASCKASGGTFSSYA ISWVRQAPGQGLEWMGSIIPIFGTANYAQKFRG RVTITADESTSTTYMELSSLRSEDTAVYYCARE SGGYRDHRLDVWGQGTMVTVSS 202 VH 31861 QVQLVQSGAEVKKPGSSVKVSCKASGGTFGSYG ISWVRQAPGQGLEWMGSIIPEFGIANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE SGTYRDHRLDVWGQGTMVTVSS 203 VH 31873 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSKYG ISWVRQAPGQGLEWMGSIIPEFGIANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE SGGYRDHRLGVWGQGTMVTVSS 204 VH 31393 QVQLVQSGAEVKKPGSSVKVSCKASGGTFESYG ISWVRQAPGQGLEWMGSIIPEFGIANYAQKFQG RVTITADESTSTTYMELSSLRSEDTAVYYCARE SGGYRDHRLDVWGQGTMVTVSS 205 VH 27534 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD FTDYSSGYSSGWTYWGQGTLVTVSS 206 VH 27536 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSNYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD TLYSSGAYYGYNVWGQGTMVTVSS 207 VH 27588 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSNYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARA KRGYDSYGGVYFDYWGQGTLVTVSS 208 VH 27590 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSNYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARG PTVTATTSIGTHNWFDPWGQGTLVTVSS 209 VH 27597 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGSIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE GRGYDSSRYYKFWFDPWGQGTLVTVSS 210 VH 27575 QVQLVQSGAEVKEPGSSVKVSCKASGGTFSSYA TSWVRQAPGQGLEWMGGIIPISGTANYAQEFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD GGGYRHHYFDLWGRGTLVTVSS 211 VH 27568 QVQLVQSGAEVKKPGSSVKVPCKASGGTFSSYA ISWVRQAPEQGLEWMGSIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCAGE SGGYRDHKLDVWGQGTVVTVSS 212 VH 27577 QVQLVQSGAEVKKPGSSVKVSCKASGGAFSSYA IGWVRQAPGQGLEWMGGIIPTFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD GGGYQHHYFDLWGRGTLVTVSS 213 VH 27587 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGSIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE SGGYRDHKLDVWGQGTMVTVSS 214 VH 27589 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD SGYHRHYSDYWGQGTLVTVSS 215 VH 27596 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD PLGIRKHWFDPWGQGTLVTVSS 216 VH 27535 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD TPRWRYHYFDYWGQGTLVTVSS 217 VH 27550 EVQLVESGGGLVKPGGSLRLSCAASGFTFSSYS MNWVRQAPGKGLEWVSSISSSSSYIYYADSVKG RFTISRDNAKNSLYLQMNSLRAEDTAVYYCARE RRGSLALGMDVWGQGTLVTVSS 218 VH 27549 QVQLVESGGGVVQPGRSLRLSCAASGFTFSSYG MNWVRQAPGKGLEWVAVIWYDGSNKYYADSVKG RFTISRDNSKNTLYLQMNSLRAEDTAVYYCARD LGGYSYGEPYYYYYGMDVWGQGTTVTVSS 219 VL 27579 EIVLIQSPGILSLSPGERATLSCRASQSVSSSY LAWYQQKPGQAPRLLIYGASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHSYITFG GGTKVEIK 220 VL 31895 EIVLIQSPGILSLSPGERATLSCRASQSVASSY LAWYQQKPGQAPRLLIYGASNRHTGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHNAITFG GGTKVEIK 221 VL 31891 EIVLIQSPGILSLSPGERATLSCRASQSVSSSY LAWYQQKPGQAPRLLIYYASSRAYGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHNAITFG GGTKVEIK 222 VL 31418 EIVLIQSPGILSLSPGERATLSCRASQSVSSSY LAWYQQKPGQAPRLLIYGASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYYFYITFG GGTKVEIK 223 VL 31430 EIVLIQSPGILSLSPGERATLSCEASQSVSYSY LAWYQQKPGQAPRLLIYGASSRANGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHSALTFG GGTKVEIK 224 VL 31431 EIVLIQSPGILSLSPGERATLSCRASQSVASSY LAWYQQKPGQAPRLLIYGASNRHTGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHGGITFG GGTKVEIK 225 VL 31421 EIVLIQSPGILSLSPGERATLSCKASESVSSSY LAWYQQKPGQAPRLLIYYASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHRRITFG GGTKVEIK 226 VL 31429 EIVLIQSPGILSLSPGERATLSCRASQYVSSSY LAWYQQKPGQAPRLLIYYASNRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHSGITFG GGTKVEIK 227 VL 28347 DIVMTQSPDSLAVSLGERATINCKSSQSVLFSS NNKNYLAWYQQKPGQPPKLLIYWASTRESGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYL YPLTFGGGTKVEIK 228 VL 31896 DIVMTQSPDSLAVSLGERATINCKSSRSVLFSS NNKNYLAWYQQKPGQPPKLLIYWASTRESGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYWT YPLTFGGGTKVEIK 229 VL 31915 DIVMTQSPDSLAVSLGERATINCKSSQSVLFSS NNKNYLAWYQQKPGQPPKLLIYWASSRESGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYWT YPLTFGGGTKVEIK 230 VL 31905 DIVMTQSPDSLAVSLGERATINCKSSKSVLYSN NNKNYLAWYQQKPGQPPKLLIYWASTRQSGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYLL YPLTFGGGTKVEIK 231 VL 31901 GIVMTQSPDSLAVSLGERATINCKSSQSVLFSS NNKNYLAWYQQKPGQPPKLLIYWASTRASGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYL YPLTFGGGTKVEIK 232 VL 27571 EIVMTQSPATLSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRATGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQYLIWPLTFG GGTKVEIK 233 VL 31861 EIVMTQSPATLSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRATGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQYLLWPLTFG GGTKVEIK 234 VL 31873 EIVMTQSPATLSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRASGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQYLLWPLTFG GGTKVEIK 235 VL 28337 DIVMTQSPDSLAVSLGERATINCKSSQSVLFSS NNKNYLAWYQQKPGQPPKLLIYWASTRESGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQFYF YPPTFGGGTKVEIK 236 VL 27536 DIVMTQSPDSLAVSLGERATINCKSSQSVLYSS NNKNYLAWYQQKPGQPPKLLIYWASTRESGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQAYT FPPTFGGGTKVEIK
237 VL 27588 DIQMTQSPSSLSASVGDRVTITCQASQDISNYL NWYQQKPGKAPKLLIYDASNLETGVPSRFSGSG SGTDFTFTISSLQPEDIATYYCQQYYIFPPTFG GGTKVEIK 238 VL 27590 EIVLTQSPATLSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQRNFYPPTFG GGTKVEIK 239 VL 27597 EIVLTQSPATLSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQFVLWPRTFG GGTKVEIK 240 VL 27575 EIVLTQSPATLSLSPGERATLSCRASQSVSRYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQHVNFPLTFG GGTKVEIK 241 VL 27568 EIVLTQSPATLSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQSVFWPITFG GGTKVEIK 242 VL 27577 EIVMTQSPATLSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRATGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQLTKWPLTFG GGTKVEIK 243 VL 27587 EIVLTQSPATLSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASKRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQDVLWPLTFG GGTKVEIK 244 VL 27589 DIQMTQSPSTLSASVGDRVTITCRASQSISSWL AWYQQKPGKAPKLLIYKASSLESGVPSRFSGSG SGTEFTLTISSLQPDDFATYYCQQYGLFPITFG GGTKVEIK 245 VL 27596 EIVMTQSPATLSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQHTVWPITFG GGTKVEIK 246 VL 27535 EIVLTQSPATLSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASKRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQVLNYPLTFG GGTKVEIK 247 VL 27550 DIQMTQSPSSLSASVGDRVTITCQASQDISNYL NWYQQKPGKAPKLLIYDASNLETGVPSRFSGSG SGTDFTFTISSLQPEDIATYYCQQSYFLPPTFG GGTKVEIK 248 VL 27549 EIVLIQSPGILSLSPGERATLSCRASQSVSSDY LAWYQQKPGQAPRLLIYGASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQAHSSPYTF GGGTKVEIK 249 hCD39 MEDTKESNVKTFCSKNILAILGFSSIIAVIALL AVGLTQNKALPENVKYGIVLDAGSSHTSLYIYK WPAEKENDTGVVHQVEECRVKGPGISKFVQKVN EIGIYLTDCMERAREVIPRSQHQETPVYLGATA GMRLLRMESEELADRVLDVVERSLSNYPFDFQG ARIITGQEEGAYGWITINYLLGKFSQKTRWFSI VPYETNNQETFGALDLGGASTQVTFVPQNQTIE SPDNALQFRLYGKDYNVYTHSFLCYGKDQALWQ KLAKDIQVASNEILRDPCFHPGYKKVVNVSDLY KTPCTKRFEMTLPFQQFEIQGIGNYQQCHQSIL ELFNTSYCPYSQCAFNGIFLPPLQGDFGAFSAF YFVMKFLNLTSEKVSQEKVTEMMKKFCAQPWEE IKTSYAGVKEKYLSEYCFSGTYILSLLLQGYHF TADSWEHIHFIGKIQGSDAGWTLGYMLNLTNMI PAEQPLSTPLSHSTYVFLMVLFSLVLFTVAIIG LLIFHKPSYFWKDMV 250 mCD39 MEDIKDSKVKRFCSKNILIILGFTSILAVIALI AVGLTQNKPLPENVKYGIVLDAGSSHTNLYIYK WPAEKENDTGVVQQLEECQVKGPGISKYAQKTD EIGAYLAECMELSTELIPTSKHHQTPVYLGATA GMRLLRMESEQSADEVLAAVSTSLKSYPFDFQG AKIITGQEEGAYGWITINYLLGRFTQEQSWLSL ISDSQKQETFGALDLGGASTQITFVPQNSTIES PENSLQFRLYGEDYTVYTHSFLCYGKDQALWQK LAKDIQVSSGGVLKDPCFNPGYEKVVNVSELYG TPCTKRFEKKLPFDQFRIQGTGDYEQCHQSILE LFNNSHCPYSQCAFNGVFLPPLHGSFGAFSAFY FVMDFFKKVAKNSVISQEKMTEITKNFCSKSWE ETKTSYPSVKEKYLSEYCFSGAYILSLLQGYNF TDSSWEQIHFMGKIKDSNAGWTLGYMLNLTNMI PAEQPLSPPLPHSTYIGLMVLFSLLLVAVAITG LFIYSKPSYFWKEAV 251 Macaca MLFDSILSTVGLSKLVSVVSSPAAALSKSNVKT fascicularis FCSKNILAILGFSSIIAVIALLAVGLTQNKALP cCD39 ENIKYGIVLDAGSSHTSLYIYKWPAEKENDTGV VHQVEECRVKGPGISKYVQKVNEIGIYLTDCME RAREVIPRSQHQETPVYLGATAGMRLLRMESEE LADRVLDVVERSLSNYPFDFQGARIITGQEEGA YGWITINYLLGKFSQKTRWFSIVPYETNNQETF GALDLGGASTQITFVPQNQTTESPDNALQFRLY GKDYNVYTHSFLCYGKDQALWQKLAKDIQVASN EILRDPCFHPGYKKVVNVSDLYKTPCTKRFEMT LPFQQFEIQGIGNYQQCHQSVLELFNTSYCPYS QCAFNGIFLPPLQGDFGAFSAFYFVMNFLNLTS EKVSQEKVTEMMKKFCSQPWEEIKTSYAGVKEK YLSEYCFSGTYILSLLLQGYHFTADSWEHIHFI GKIQGSDAGWTLGYMLNLTNMIPAEQPLSTPLS HSTYVFLMVLFSLVLVIVAIIGLLIFHKPSYFW KDMV 252 hCD39 ECD TQNKALPENVKYGIVLDAGSSHTSLYIYKWPAE KENDTGVVHQVEECRVKGPGISKFVQKVNEIGI YLTDCMERAREVIPRSQHQETPVYLGATAGMRL LRMESEELADRVLDVVERSLSNYPFDFQGARII TGQEEGAYGWITINYLLGKFSQKTRWFSIVPYE TNNQETFGALDLGGASTQVTFVPQNQTIESPDN ALQFRLYGKDYNVYTHSFLCYGKDQALWQKLAK DIQVASNEILRDPCFHPGYKKVVNVSDLYKTPC TKRFEMTLPFQQFEIQGIGNYQQCHQSILELFN TSYCPYSQCAFNGIFLPPLQGDFGAFSAFYFVM KFLNLTSEKVSQEKVTEMMKKFCAQPWEEIKTS YAGVKEKYLSEYCFSGTYILSLLLQGYHFTADS WEHIHFIGKIQGSDAGWTLGYMLNLTNMIPAEQ PLSTPLSHSTYVFLMVLFSLVLFTVAIIGLLIF HKPSYFWKDMV 253 mCD39 ECD TQNKPLPENVKYGIVLDAGSSHTNLYIYKWPAE KENDTGVVQQLEECQVKGPGISKYAQKTDEIGA YLAECMELSTELIPTSKHHQTPVYLGATAGMRL LRMESEQSADEVLAAVSTSLKSYPFDFQGAKII TGQEEGAYGWITINYLLGRFTQEQSWLSLISDS QKQETFGALDLGGASTQITFVPQNSTIESPENS LQFRLYGEDYTVYTHSFLCYGKDQALWQKLAKD IQVSSGGVLKDPCFNPGYEKVVNVSELYGTPCT KRFEKKLPFDQFRIQGTGDYEQCHQSILELFNN SHCPYSQCAFNGVFLPPLHGSFGAFSAFYFVMD FFKKVAKNSVISQEKMTEITKNFCSKSWEETKT SYPSVKEKYLSEYCFSGAYILSLLQGYNFTDSS WEQIHFMGKIKDSNAGWTLGYMLNLTNMIPAEQ PLSPPLPHSTYIGLMVLFSLLLVAVAITGLFIY SKPSYFWKEAV 254 cCD39 ECD TQNKALPENIKYGIVLDAGSSHTSLYIYKWPAE KENDTGVVHQVEECRVKGPGISKYVQKVNEIGI YLTDCMERAREVIPRSQHQETPVYLGATAGMRL LRMESEELADRVLDVVERSLSNYPFDFQGARII TGQEEGAYGWITINYLLGKFSQKTRWFSIVPYE TNNQETFGALDLGGASTQITFVPQNQTTESPDN ALQFRLYGKDYNVYTHSFLCYGKDQALWQKLAK DIQVASNEILRDPCFHPGYKKVVNVSDLYKTPC TKRFEMTLPFQQFEIQGIGNYQQCHQSVLELFN TSYCPYSQCAFNGIFLPPLQGDFGAFSAFYFVM NFLNLTSEKVSQEKVTEMMKKFCSQPWEEIKTS YAGVKEKYLSEYCFSGTYILSLLLQGYHFTADS WEHIHFIGKIQGSDAGWTLGYMLNLTNMIPAEQ PLSTPLSHSTYVFLMVLFSLVLVIVAIIGLLIF HKPSYFWKDMV 255 HC 31895 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSVGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRKWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDILMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 256 LC 31895 EIVLIQSPGILSLSPGERATLSCRASQSVASSY LAWYQQKPGQAPRLLIYGASNRHTGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHNAITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 257 HC 31415 QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYQ MHWVRQAPGQGLEWMGRINPSGGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRSWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDILMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 258 LC 31415 EIVLIQSPGILSLSPGERATLSCRASQSVSSSY LAWYQQKPGQAPRLLIYGASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHSYITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 259 HC 31891 QVQLVQSGAEVKKPGASVKVSCKASGYIFKSYE MHWVRQAPGQGLEWMGRINPSVGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRVWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDILMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 260 LC 31891 EIVLIQSPGILSLSPGERATLSCRASQSVSSSY LAWYQQKPGQAPRLLIYYASSRAYGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHNAITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 261 HC 31418 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSGGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDILMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 262 LC 31418 EIVLIQSPGILSLSPGERATLSCRASQSVSSSY LAWYQQKPGQAPRLLIYGASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYYFYITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH
QGLSSPVTKSFNRGEC 263 HC 31430 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSGGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRHWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDILMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 264 LC 31430 EIVLIQSPGILSLSPGERATLSCEASQSVSYSY LAWYQQKPGQAPRLLIYGASSRANGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHSALTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 265 HC 31915 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSLP ISWVRQAPGQGLEWMGGIGFGTANYAQKFQGRV TITADESTSTAYMELSSLRSEDTAVYYCARGGA KYASKWGMDVWGQGTTVTVSSASTKGPSVFPLA PCSRSTSESTAALGCLVKDYFPEPVTVSWNSGA LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGT KTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPA PEFLGGPSVFLFPPKPKDILMISRIPEVICVVV DVSQEDPEVQFNWYVDGVEVHNAKTKPREEQFN STYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPS SIEKTISKAKGQPREPQVYTLPPSQEEMTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPP VLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMH EALHNHYTQKSLSLSLGK 266 LC 31915 DIVMTQSPDSLAVSLGERATINCKSSQSVLFSS NNKNYLAWYQQKPGQPPKLLIYWASSRESGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYWT YPLTFGGGTKVEIKRTVAAPSVFIFPPSDEQLK SGTASVVCLLNNFYPREAKVQWKVDNALQSGNS QESVTEQDSKDSTYSLSSTLTLSKADYEKHKVY ACEVTHQGLSSPVTKSFNRGEC 267 HC 31905 QVQLVQSGAEVKKPGSSVKVSCKASGGTFPSNA ISWVRQAPGQGLEWMGGIGFGTANYAQKFQGRV TITADESTSTAYMELSSLRSEDTAVYYCARGGA KYARTYGMDVWGQGTIVIVSSASTKGPSVFPLA PCSRSTSESTAALGCLVKDYFPEPVTVSWNSGA LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGT KTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPA PEFLGGPSVFLFPPKPKDTLMISRIPEVTCVVV DVSQEDPEVQFNWYVDGVEVHNAKTKPREEQFN STYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPS SIEKTISKAKGQPREPQVYTLPPSQEEMTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPP VLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMH EALHNHYTQKSLSLSLGK 268 LC 31905 DIVMTQSPDSLAVSLGERATINCKSSKSVLYSN NNKNYLAWYQQKPGQPPKLLIYWASTRQSGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYLL YPLTFGGGTKVEIKRTVAAPSVFIFPPSDEQLK SGTASVVCLLNNFYPREAKVQWKVDNALQSGNS QESVTEQDSKDSTYSLSSTLTLSKADYEKHKVY ACEVTHQGLSSPVTKSFNRGEC 269 HC 31901 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSLP ISWVRQAPGQGLEWMGGIGFGTANYAQKFQGRV TITADESTSTAYMELSSLRSEDTAVYYCARGGA KYAGRYGMDVWGQGTIVIVSSASTKGPSVFPLA PCSRSTSESTAALGCLVKDYFPEPVTVSWNSGA LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGT KTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPA PEFLGGPSVFLFPPKPKDTLMISRIPEVTCVVV DVSQEDPEVQFNWYVDGVEVHNAKTKPREEQFN STYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPS SIEKTISKAKGQPREPQVYTLPPSQEEMTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPP VLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMH EALHNHYTQKSLSLSLGK 270 LC 31901 GIVMTQSPDSLAVSLGERATINCKSSQSVLFSS NNKNYLAWYQQKPGQPPKLLIYWASTRASGVPD RFSGSGSGTDFTLTISSLQAEDVAVYYCQQYYL YPLTFGGGTKVEIKRTVAAPSVFIFPPSDEQLK SGTASVVCLLNNFYPREAKVQWKVDNALQSGNS QESVTEQDSKDSTYSLSSTLTLSKADYEKHKVY ACEVTHQGLSSPVTKSFNRGEC 271 HC 31861 QVQLVQSGAEVKKPGSSVKVSCKASGGTFGSYG ISWVRQAPGQGLEWMGSIIPEFGIANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE SGTYRDHRLDVWGQGTMVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 272 LC 31861 EIVMTQSPAILSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRATGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQYLLWPLTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 273 HC 31873 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSKYG ISWVRQAPGQGLEWMGSIIPEFGIANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE SGGYRDHRLGVWGQGTMVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 274 LC 31873 EIVMTQSPAILSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRASGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQYLLWPLTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 275 HC 31393 QVQLVQSGAEVKKPGSSVKVSCKASGGTFESYG ISWVRQAPGQGLEWMGSIIPEFGIANYAQKFQG RVTITADESTSTTYMELSSLRSEDTAVYYCARE SGGYRDHRLDVWGQGTMVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 276 LC 31393 EIVMTQSPAILSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRATGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQYLLWPLTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 277 HC 27597 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGSIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE GRGYDSSRYYKFWFDPWGQGTLVTVSSASTKGP SVFPLAPCSRSTSESTAALGCLVKDYFPEPVTV SWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVP SSSLGTKTYTCNVDHKPSNTKVDKRVESKYGPP CPPCPAPEFLGGPSVFLFPPKPKDTLMISRTPE VTCVVVDVSQEDPEVQFNWYVDGVEVHNAKTKP REEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVS NKGLPSSIEKTISKAKGQPREPQVYTLPPSQEE MTKNQVSLTCLVKGFYPSDIAVEWESNGQPENN YKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVF SCSVMHEALHNHYTQKSLSLSLGK 278 LC 27597 EIVLTQSPAILSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQFVLWPRTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 279 HC 27575 QVQLVQSGAEVKEPGSSVKVSCKASGGTFSSYA TSWVRQAPGQGLEWMGGIIPISGTANYAQEFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD GGGYRHHYFDLWGRGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 280 LC 27575 EIVLTQSPAILSLSPGERATLSCRASQSVSRYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQHVNFPLTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 281 HC 27568 QVQLVQSGAEVKKPGSSVKVPCKASGGTFSSYA ISWVRQAPEQGLEWMGSIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCAGE SGGYRDHKLDVWGQGTVVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 282 LC 27568 EIVLTQSPAILSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQSVFWPITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 283 HC 27577 QVQLVQSGAEVKKPGSSVKVSCKASGGAFSSYA IGWVRQAPGQGLEWMGGIIPTFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD GGGYQHHYFDLWGRGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 284 LC 27577 EIVMTQSPAILSVSPGERATLSCRASQSVGSNL AWYQQKPGQAPRLLIYGASTRATGIPARFSGSG SGTEFTLTISSLQSEDFAVYYCQQLTKWPLTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV
VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 285 HC 27587 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGSIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARE SGGYRDHKLDVWGQGTMVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 286 LC 27587 EIVLTQSPAILSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASKRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQDVLWPLTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 287 HC 27589 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD SGYHRHYSDYWGQGTLVTVSSASTKGPSVFPLA PCSRSTSESTAALGCLVKDYFPEPVTVSWNSGA LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGT KTYTCNVDHKPSNTKVDKRVESKYGPPCPPCPA PEFLGGPSVFLFPPKPKDTLMISRIPEVTCVVV DVSQEDPEVQFNWYVDGVEVHNAKTKPREEQFN STYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPS SIEKTISKAKGQPREPQVYTLPPSQEEMTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPP VLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMH EALHNHYTQKSLSLSLGK 288 LC 27589 DIQMTQSPSTLSASVGDRVTITCRASQSISSWL AWYQQKPGKAPKLLIYKASSLESGVPSRFSGSG SGTEFTLTISSLQPDDFATYYCQQYGLFPITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 289 HC 27596 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD PLGIRKHWFDPWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 290 LC 27596 EIVMTQSPAILSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASNRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQHTVWPITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 291 HC 27535 QVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYA ISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQG RVTITADESTSTAYMELSSLRSEDTAVYYCARD TPRWRYHYFDYWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 292 LC 27535 EIVLTQSPAILSLSPGERATLSCRASQSVSSYL AWYQQKPGQAPRLLIYDASKRATGIPARFSGSG SGTDFTLTISSLEPEDFAVYYCQQVLNYPLTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 293 HC 27550 EVQLVESGGGLVKPGGSLRLSCAASGFTFSSYS MNWVRQAPGKGLEWVSSISSSSSYIYYADSVKG RFTISRDNAKNSLYLQMNSLRAEDTAVYYCARE RRGSLALGMDVWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDTLMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 294 LC 27550 DIQMTQSPSSLSASVGDRVTITCQASQDISNYL NWYQQKPGKAPKLLIYDASNLETGVPSRFSGSG SGTDFTFTISSLQPEDIATYYCQQSYFLPPTFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC 295 HC 27549 QVQLVESGGGVVQPGRSLRLSCAASGFTFSSYG MNWVRQAPGKGLEWVAVIWYDGSNKYYADSVKG RFTISRDNSKNTLYLQMNSLRAEDTAVYYCARD LGGYSYGEPYYYYYGMDVWGQGTTVTVSSASTK GPSVFPLAPCSRSTSESTAALGCLVKDYFPEPV TVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVT VPSSSLGTKTYTCNVDHKPSNTKVDKRVESKYG PPCPPCPAPEFLGGPSVFLFPPKPKDTLMISRT PEVTCVVVDVSQEDPEVQFNWYVDGVEVHNAKT KPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCK VSNKGLPSSIEKTISKAKGQPREPQVYTLPPSQ EEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGN VFSCSVMHEALHNHYTQKSLSLSLGK 296 LC 27549 EIVLIQSPGILSLSPGERATLSCRASQSVSSDY LAWYQQKPGQAPRLLIYGASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQAHSSPYTF GGGTKVEIKRTVAAPSVFIFPPSDEQLKSGTAS VVCLLNNFYPREAKVQWKVDNALQSGNSQESVT EQDSKDSTYSLSSTLTLSKADYEKHKVYACEVT HQGLSSPVTKSFNRGEC 297 HC 31414 QVQLVQSGAEVKKPGASVKVSCKASGYTFKSYE MHWVRQAPGQGLEWMGRINPSVGSTWYAQKFQG RVIMIRDTSTSTVYMELSSLRSEDTAVYYCARG KREGGTEYLRNWGQGTLVTVSSASTKGPSVFPL APCSRSTSESTAALGCLVKDYFPEPVTVSWNSG ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLG TKTYTCNVDHKPSNTKVDKRVESKYGPPCPPCP APEFLGGPSVFLFPPKPKDILMISRTPEVICVV VDVSQEDPEVQFNWYVDGVEVHNAKTKPREEQF NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLP SSIEKTISKAKGQPREPQVYTLPPSQEEMTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVM HEALHNHYTQKSLSLSLGK 298 LC 31414 EIVLIQSPGILSLSPGERATLSCRASQSVSSSY LAWYQQKPGQAPRLLIYGASSRATGIPDRFSGS GSGTDFTLTISRLEPEDFAVYYCQQYHSYITFG GGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASV VCLLNNFYPREAKVQWKVDNALQSGNSQESVTE QDSKDSTYSLSSTLTLSKADYEKHKVYACEVTH QGLSSPVTKSFNRGEC
EQUIVALENTS
[0466] The disclosure set forth above may encompass multiple distinct inventions with independent utility. Although each of these inventions has been disclosed in its preferred form(s), the specific embodiments thereof as disclosed and illustrated herein are not to be considered in a limiting sense, because numerous variations are possible. The subject matter of the inventions includes all novel and nonobvious combinations and subcombinations of the various elements, features, functions, and/or properties disclosed herein. The following claims particularly point out certain combinations and subcombinations regarded as novel and nonobvious. Inventions embodied in other combinations and subcombinations of features, functions, elements, and/or properties may be claimed in this application, in applications claiming priority from this application, or in related applications. Such claims, whether directed to a different invention or to the same invention, and whether broader, narrower, equal, or different in scope in comparison to the original claims, also are regarded as included within the subject matter of the inventions of the present disclosure.
Sequence CWU 1
1
29817PRTArtificial SequenceSynthetic CDR-H1, Chothia 1Gly Tyr Thr
Phe Thr Ser Tyr 1 5 27PRTArtificial SequenceSynthetic CDR-H1,
Chothia 2Gly Tyr Thr Phe Lys Ser Tyr 1 5 37PRTArtificial
SequenceSynthetic CDR-H1, Chothia 3Gly Tyr Ile Phe Lys Ser Tyr 1 5
47PRTArtificial SequenceSynthetic CDR-H1, Chothia 4Gly Tyr Thr Phe
Gln Ser Tyr 1 5 57PRTArtificial SequenceSynthetic CDR-H1, Chothia
5Gly Tyr Thr Phe Phe Ser Tyr 1 5 67PRTArtificial SequenceSynthetic
CDR-H1, Chothia 6Gly Tyr Thr Phe Val Ser Tyr 1 5 710PRTArtificial
SequenceSynthetic CDR-H1, Chothia 7Gly Gly Thr Phe Ser Ser Leu Ala
Ile Ser 1 5 10 810PRTArtificial SequenceSynthetic CDR-H1, Chothia
8Gly Gly Thr Phe Ser Lys Leu Ala Ile Ser 1 5 10 97PRTArtificial
SequenceSynthetic CDR-H1, Chothia 9Gly Gly Thr Phe Ser His Thr 1 5
107PRTArtificial SequenceSynthetic CDR-H1, Chothia 10Gly Gly Thr
Phe Ser Ser Leu 1 5 117PRTArtificial SequenceSynthetic CDR-H1,
Chothia 11Gly Gly Thr Phe Ser Leu Leu 1 5 127PRTArtificial
SequenceSynthetic CDR-H1, Chothia 12Gly Gly Thr Phe Gln Ser Leu 1 5
137PRTArtificial SequenceSynthetic CDR-H1, Chothia 13Gly Gly Thr
Phe Pro Ser Asn 1 5 147PRTArtificial SequenceSynthetic CDR-H1,
Chothia 14Gly Gly Thr Phe Ser Ala Met 1 5 157PRTArtificial
SequenceSynthetic CDR-H1, Chothia 15Gly Gly Thr Phe Ala Ser Leu 1 5
167PRTArtificial SequenceSynthetic CDR-H1, Chothia 16Gly Gly Thr
Phe Ser Trp Leu 1 5 177PRTArtificial SequenceSynthetic CDR-H1,
Chothia 17Gly Gly Thr Phe Ser Ser Tyr 1 5 187PRTArtificial
SequenceSynthetic CDR-H1, Chothia 18Gly Gly Thr Phe Gly Ser Tyr 1 5
197PRTArtificial SequenceSynthetic CDR-H1, Chothia 19Gly Gly Thr
Phe Ser Lys Tyr 1 5 207PRTArtificial SequenceSynthetic CDR-H1,
Chothia 20Gly Gly Thr Phe Gly Arg Tyr 1 5 217PRTArtificial
SequenceSynthetic CDR-H1, Chothia 21Gly Gly Thr Phe Glu Ser Tyr 1 5
227PRTArtificial SequenceSynthetic CDR-H1, Chothia 22Gly Gly Thr
Phe Ser Asn Tyr 1 5 237PRTArtificial SequenceSynthetic CDR-H1,
Chothia 23Gly Gly Ala Phe Ser Ser Tyr 1 5 247PRTArtificial
SequenceSynthetic CDR-H1, Chothia 24Gly Phe Thr Phe Ser Ser Tyr 1 5
255PRTArtificial SequenceSynthetic CDR-H1, Kabat 25Ser Tyr Tyr Met
His 1 5 265PRTArtificial SequenceSynthetic CDR-H1, Kabat 26Ser Tyr
Glu Met His 1 5 275PRTArtificial SequenceSynthetic CDR-H1, Kabat
27Ser Tyr Gln Met His 1 5 285PRTArtificial SequenceSynthetic
CDR-H1, Kabat 28Ser Tyr Tyr Met Tyr 1 5 295PRTArtificial
SequenceSynthetic CDR-H1, Kabat 29Ser Tyr Phe Met His 1 5
305PRTArtificial SequenceSynthetic CDR-H1, Kabat 30Ser Leu Ala Ile
Ser 1 5 315PRTArtificial SequenceSynthetic CDR-H1, Kabat 31Lys Leu
Ala Ile Ser 1 5 325PRTArtificial SequenceSynthetic CDR-H1, Kabat
32His Thr Ala Ile Ser 1 5 335PRTArtificial SequenceSynthetic
CDR-H1, Kabat 33Ser Leu Pro Ile Ser 1 5 345PRTArtificial
SequenceSynthetic CDR-H1, Kabat 34Leu Leu Ala Ile Ser 1 5
355PRTArtificial SequenceSynthetic CDR-H1, Kabat 35Ser Asn Ala Ile
Ser 1 5 365PRTArtificial SequenceSynthetic CDR-H1, Kabat 36Ala Met
Ala Ile Ser 1 5 375PRTArtificial SequenceSynthetic CDR-H1, Kabat
37Trp Leu Ala Ile Ser 1 5 385PRTArtificial SequenceSynthetic
CDR-H1, Kabat 38Ser Tyr Ala Ile Ser 1 5 395PRTArtificial
SequenceSynthetic CDR-H1, Kabat 39Ser Tyr Gly Ile Ser 1 5
405PRTArtificial SequenceSynthetic CDR-H1, Kabat 40Lys Tyr Gly Ile
Ser 1 5 415PRTArtificial SequenceSynthetic CDR-H1, Kabat 41Asn Tyr
Ala Ile Ser 1 5 425PRTArtificial SequenceSynthetic CDR-H1, Kabat
42Ser Tyr Ala Thr Ser 1 5 435PRTArtificial SequenceSynthetic
CDR-H1, Kabat 43Ser Tyr Ala Ile Gly 1 5 445PRTArtificial
SequenceSynthetic CDR-H1, Kabat 44Ser Tyr Ser Met Asn 1 5
455PRTArtificial SequenceSynthetic CDR-H1, Kabat 45Ser Tyr Gly Met
Asn 1 5 467PRTArtificial SequenceSynthetic CDR-H2, Chothia 46Asn
Pro Ser Gly Gly Ser Thr 1 5 476PRTArtificial SequenceSynthetic
CDR-H2, Chothia 47Asn Pro Ser Val Gly Ser 1 5 486PRTArtificial
SequenceSynthetic CDR-H2, Chothia 48Asn Pro Ser Gly Gly Ser 1 5
496PRTArtificial SequenceSynthetic CDR-H2, Chothia 49Asn Pro Leu
Gly Gly Gly 1 5 506PRTArtificial SequenceSynthetic CDR-H2, Chothia
50Asn Pro Arg Gly Gly Ser 1 5 516PRTArtificial SequenceSynthetic
CDR-H2, Chothia 51Ile Pro Ile Phe Gly Thr 1 5 524PRTArtificial
SequenceSynthetic CDR-H2, Chothia 52Gly Phe Gly Thr 1
536PRTArtificial SequenceSynthetic CDR-H2, Chothia 53Leu Pro Ile
Gly Gly Thr 1 5 546PRTArtificial SequenceSynthetic CDR-H2, Chothia
54Leu Pro Ile Ala Gly Thr 1 5 556PRTArtificial SequenceSynthetic
CDR-H2, Chothia 55Leu Pro Ile Phe Gly Glu 1 5 566PRTArtificial
SequenceSynthetic CDR-H2, Chothia 56Ile Pro Arg Gly Gly Thr 1 5
576PRTArtificial SequenceSynthetic CDR-H2, Chothia 57Ile Pro Glu
Phe Gly Ile 1 5 586PRTArtificial SequenceSynthetic CDR-H2, Chothia
58Ile Pro Ser Ile Gly Thr 1 5 596PRTArtificial SequenceSynthetic
CDR-H2, Chothia 59Ile Pro Ile Ser Gly Thr 1 5 606PRTArtificial
SequenceSynthetic CDR-H2, Chothia 60Ile Pro Thr Phe Gly Thr 1 5
616PRTArtificial SequenceSynthetic CDR-H2, Chothia 61Ser Ser Ser
Ser Ser Tyr 1 5 626PRTArtificial SequenceSynthetic CDR-H2, Chothia
62Trp Tyr Asp Gly Ser Asn 1 5 6317PRTArtificial SequenceSynthetic
CDR-H2, Kabat 63Val Ile Asn Pro Ser Gly Gly Ser Thr Ser Tyr Ala Gln
Lys Phe Gln 1 5 10 15 Gly 6417PRTArtificial SequenceSynthetic
CDR-H2, Kabat 64Arg Ile Asn Pro Ser Val Gly Ser Thr Trp Tyr Ala Gln
Lys Phe Gln 1 5 10 15 Gly 6517PRTArtificial SequenceSynthetic
CDR-H2, Kabat 65Arg Ile Asn Pro Ser Gly Gly Ser Thr Trp Tyr Ala Gln
Lys Phe Gln 1 5 10 15 Gly 6617PRTArtificial SequenceSynthetic
CDR-H2, Kabat 66Lys Ile Asn Pro Ser Gly Gly Ser Thr Trp Tyr Ala Gln
Lys Phe Gln 1 5 10 15 Gly 6717PRTArtificial SequenceSynthetic
CDR-H2, Kabat 67Val Ile Asn Pro Leu Gly Gly Gly Thr Ser Tyr Ala Gln
Lys Phe Gln 1 5 10 15 Gly 6817PRTArtificial SequenceSynthetic
CDR-H2, Kabat 68Ser Ile Asn Pro Arg Gly Gly Ser Thr Ser Tyr Ala Gln
Lys Phe Gln 1 5 10 15 Gly 6917PRTArtificial SequenceSynthetic
CDR-H2, Kabat 69Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln
Lys Phe Gln 1 5 10 15 Gly 7015PRTArtificial SequenceSynthetic
CDR-H2, Kabat 70Gly Ile Gly Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe
Gln Gly 1 5 10 15 7117PRTArtificial SequenceSynthetic CDR-H2, Kabat
71Gly Ile Leu Pro Ile Gly Gly Thr Ala Asn Tyr Ala Gln Lys Phe Gln 1
5 10 15 Gly 7217PRTArtificial SequenceSynthetic CDR-H2, Kabat 72Gly
Ile Leu Pro Ile Ala Gly Thr Ala Asn Tyr Ala Gln Lys Phe Gln 1 5 10
15 Gly 7317PRTArtificial SequenceSynthetic CDR-H2, Kabat 73Gly Ile
Leu Pro Ile Phe Gly Glu Ala Asn Tyr Ala Gln Lys Phe Gln 1 5 10 15
Gly 7417PRTArtificial SequenceSynthetic CDR-H2, Kabat 74Gly Ile Ile
Pro Arg Gly Gly Thr Ala Asn Tyr Ala Gln Lys Phe Gln 1 5 10 15 Gly
7517PRTArtificial SequenceSynthetic CDR-H2, Kabat 75Ser Ile Ile Pro
Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe Arg 1 5 10 15 Gly
7617PRTArtificial SequenceSynthetic CDR-H2, Kabat 76Ser Ile Ile Pro
Glu Phe Gly Ile Ala Asn Tyr Ala Gln Lys Phe Gln 1 5 10 15 Gly
7717PRTArtificial SequenceSynthetic CDR-H2, Kabat 77Ser Ile Ile Pro
Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe Gln 1 5 10 15 Gly
7817PRTArtificial SequenceSynthetic CDR-H2, Kabat 78Gly Ile Ile Pro
Ile Ser Gly Thr Ala Asn Tyr Ala Gln Glu Phe Gln 1 5 10 15 Gly
7917PRTArtificial SequenceSynthetic CDR-H2, Kabat 79Gly Ile Ile Pro
Thr Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe Gln 1 5 10 15 Gly
8017PRTArtificial SequenceSynthetic CDR-H2, Kabat 80Ser Ile Ser Ser
Ser Ser Ser Tyr Ile Tyr Tyr Ala Asp Ser Val Lys 1 5 10 15 Gly
8117PRTArtificial SequenceSynthetic CDR-H2, Kabat 81Val Ile Trp Tyr
Asp Gly Ser Asn Lys Tyr Tyr Ala Asp Ser Val Lys 1 5 10 15 Gly
8212PRTArtificial SequenceSynthetic CDR-H3 82Gly Lys Arg Glu Gly
Gly Thr Glu Tyr Leu Arg His 1 5 10 8312PRTArtificial
SequenceSynthetic CDR-H3 83Gly Lys Arg Glu Gly Gly Thr Glu Tyr Leu
Arg Lys 1 5 10 8412PRTArtificial SequenceSynthetic CDR-H3 84Gly Lys
Arg Glu Gly Gly Thr Glu Tyr Leu Arg Ser 1 5 10 8512PRTArtificial
SequenceSynthetic CDR-H3 85Gly Lys Arg Glu Gly Gly Thr Glu Tyr Leu
Arg Asn 1 5 10 8612PRTArtificial SequenceSynthetic CDR-H3 86Gly Lys
Arg Glu Gly Gly Thr Glu Tyr Leu Arg Val 1 5 10 8713PRTArtificial
SequenceSynthetic CDR-H3 87Gly Gly Ala Lys Tyr Ala Ser Thr Tyr Gly
Met Asp Val 1 5 10 8813PRTArtificial SequenceSynthetic CDR-H3 88Gly
Gly Ala Lys Tyr Ala Ser Thr His Gly Met Asp Val 1 5 10
8913PRTArtificial SequenceSynthetic CDR-H3 89Gly Gly Ala Lys Tyr
Ala Ser Gln Leu Gly Met Asp Val 1 5 10 9013PRTArtificial
SequenceSynthetic CDR-H3 90Gly Gly Ala Lys Tyr Ala Ser Lys Trp Gly
Met Asp Val 1 5 10 9113PRTArtificial SequenceSynthetic CDR-H3 91Gly
Gly Ala Lys Tyr Ala Val Gly Tyr Gly Met Asp Val 1 5 10
9213PRTArtificial SequenceSynthetic CDR-H3 92Gly Gly Ala Lys Tyr
Ala Gly Arg Tyr Gly Met Asp Val 1 5 10 9313PRTArtificial
SequenceSynthetic CDR-H3 93Gly Gly Ala Lys Tyr Ala Arg Thr Tyr Gly
Met Asp Val 1 5 10 9412PRTArtificial SequenceSynthetic CDR-H3 94Glu
Ser Gly Gly Tyr Arg Asp His Arg Leu Asp Val 1 5 10
9512PRTArtificial SequenceSynthetic CDR-H3 95Glu Ser Gly Thr Tyr
Arg Asp His Arg Leu Asp Val 1 5 10 9612PRTArtificial
SequenceSynthetic CDR-H3 96Glu Ser Gly Gly Tyr Arg Asp His Arg Leu
Gly Val 1 5 10 9715PRTArtificial SequenceSynthetic CDR-H3 97Asp Phe
Thr Asp Tyr Ser Ser Gly Tyr Ser Ser Gly Trp Thr Tyr 1 5 10 15
9814PRTArtificial SequenceSynthetic CDR-H3 98Asp Thr Leu Tyr Ser
Ser Gly Ala Tyr Tyr Gly Tyr Asn Val 1 5 10 9915PRTArtificial
SequenceSynthetic CDR-H3 99Ala Lys Arg Gly Tyr Asp Ser Tyr Gly Gly
Val Tyr Phe Asp Tyr 1 5 10 15 10018PRTArtificial SequenceSynthetic
CDR-H3 100Gly Pro Thr Val Thr Ala Thr Thr Ser Ile Gly Thr His Asn
Trp Phe 1 5 10 15 Asp Pro 10128PRTArtificial SequenceSynthetic
CDR-H3 101Glu Gly Arg Gly Tyr Asp Ser Ser Arg Tyr Tyr Lys Phe Trp
Phe Asp 1 5 10 15 Pro Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser
20 25 10212PRTArtificial SequenceSynthetic CDR-H3 102Asp Gly Gly
Gly Tyr Arg His His Tyr Phe Asp Leu 1 5 10 10312PRTArtificial
SequenceSynthetic CDR-H3 103Glu Ser Gly Gly Tyr Arg Asp His Lys Leu
Asp Val 1 5 10 10412PRTArtificial SequenceSynthetic CDR-H3 104Asp
Gly Gly Gly Tyr Gln His His Tyr Phe Asp Leu 1 5 10
10511PRTArtificial SequenceSynthetic CDR-H3 105Asp Ser Gly Tyr His
Arg His Tyr Ser Asp Tyr 1 5 10 10612PRTArtificial SequenceSynthetic
CDR-H3 106Asp Pro Leu Gly Ile Arg Lys His Trp Phe Asp Pro 1 5 10
10712PRTArtificial SequenceSynthetic CDR-H3 107Asp Thr Pro Arg Trp
Arg Tyr His Tyr Phe Asp Tyr 1 5 10 10812PRTArtificial
SequenceSynthetic CDR-H3 108Glu Arg Arg Gly Ser Leu Ala Leu Gly Met
Asp Val 1 5 10 10919PRTArtificial SequenceSynthetic CDR-H3 109Asp
Leu Gly Gly Tyr Ser Tyr Gly Glu Pro Tyr Tyr Tyr Tyr Tyr Gly 1 5 10
15 Met Asp Val 11012PRTArtificial SequenceSynthetic CDR-L1 110Arg
Ala Ser Gln Ser Val Ser Ser Ser Tyr Leu Ala 1 5 10
11112PRTArtificial SequenceSynthetic CDR-L1 111Arg Ala Ser Gln Ser
Val Ala Ser Ser Tyr Leu Ala 1 5 10 11212PRTArtificial
SequenceSynthetic CDR-L1 112Glu Ala Ser Gln Ser Val Ser Tyr Ser Tyr
Leu Ala 1 5 10 11312PRTArtificial SequenceSynthetic CDR-L1 113Lys
Ala Ser Glu Ser Val Ser Ser Ser Tyr Leu Ala 1 5 10
11412PRTArtificial SequenceSynthetic CDR-L1 114Arg Ala Ser Gln Tyr
Val Ser Ser Ser Tyr Leu Ala 1 5 10 11517PRTArtificial
SequenceSynthetic CDR-L1 115Lys Ser Ser Gln Ser Val Leu Phe Ser Ser
Asn Asn Lys Asn Tyr Leu 1 5 10 15 Ala 11617PRTArtificial
SequenceSynthetic CDR-L1 116Lys Ser Ser Arg Ser Val Leu Phe Ser Ser
Asn Asn Lys Asn Tyr Leu 1 5 10 15 Ala 11717PRTArtificial
SequenceSynthetic CDR-L1 117Lys Ser Ser Lys Ser Val Leu Tyr Ser Asn
Asn Asn Lys Asn Tyr Leu 1 5 10 15 Ala 11811PRTArtificial
SequenceSynthetic CDR-L1 118Arg Ala Ser Gln Ser Val Gly Ser Asn Leu
Ala 1 5 10 11917PRTArtificial SequenceSynthetic CDR-L1 119Lys Ser
Ser Gln Ser Val Leu Tyr Ser Ser Asn Asn Lys Asn Tyr Leu 1 5 10 15
Ala 12011PRTArtificial SequenceSynthetic CDR-L1 120Gln Ala Ser Gln
Asp Ile Ser Asn Tyr Leu Asn 1 5 10 12111PRTArtificial
SequenceSynthetic CDR-L1 121Arg Ala Ser Gln Ser Val Ser Ser Tyr Leu
Ala 1 5 10 12211PRTArtificial SequenceSynthetic CDR-L1 122Arg Ala
Ser Gln Ser Val Ser Arg Tyr Leu Ala 1 5 10 12311PRTArtificial
SequenceSynthetic CDR-L1 123Arg Ala Ser Gln Ser Ile Ser Ser Trp Leu
Ala 1 5 10 12412PRTArtificial SequenceSynthetic CDR-L1 124Arg Ala
Ser Gln Ser Val Ser Ser Asp Tyr Leu Ala 1 5 10 1257PRTArtificial
SequenceSynthetic CDR-L2 125Gly Ala Ser Ser Arg Ala Thr 1 5
1267PRTArtificial SequenceSynthetic CDR-L2 126Gly Ala Ser Asn Arg
His Thr 1 5 1277PRTArtificial SequenceSynthetic CDR-L2 127Tyr Ala
Ser Ser Arg Ala Tyr 1 5 1287PRTArtificial SequenceSynthetic CDR-L2
128Gly Ala Ser Ser Arg Ala Asn 1 5 1297PRTArtificial
SequenceSynthetic CDR-L2 129Tyr Ala Ser Ser Arg Ala Thr 1 5
1307PRTArtificial SequenceSynthetic CDR-L2 130Tyr Ala Ser Asn Arg
Ala Thr 1 5 1317PRTArtificial SequenceSynthetic CDR-L2 131Trp Ala
Ser Thr Arg Glu Ser 1 5 1327PRTArtificial SequenceSynthetic CDR-L2
132Trp Ala Ser Ser Arg Glu Ser 1 5 1337PRTArtificial
SequenceSynthetic CDR-L2 133Trp Ala Ser Thr Arg Gln Ser 1 5
1347PRTArtificial SequenceSynthetic CDR-L2 134Trp Ala Ser Thr Arg
Ala Ser 1 5 1357PRTArtificial SequenceSynthetic CDR-L2 135Gly Ala
Ser Thr Arg Ala Thr 1 5 1367PRTArtificial SequenceSynthetic CDR-L2
136Gly Ala Ser Thr Arg Ala Ser 1 5 1377PRTArtificial
SequenceSynthetic CDR-L2 137Asp Ala Ser Asn Leu Glu Thr 1 5
1387PRTArtificial SequenceSynthetic CDR-L2 138Asp Ala Ser Asn Arg
Ala Thr 1 5 1397PRTArtificial SequenceSynthetic CDR-L2 139Asp Ala
Ser Lys Arg Ala Thr 1 5 1407PRTArtificial SequenceSynthetic
CDR-L2
140Lys Ala Ser Ser Leu Glu Ser 1 5 1418PRTArtificial
SequenceSynthetic CDR-L3 141Gln Gln Tyr His Ser Tyr Ile Thr 1 5
1428PRTArtificial SequenceSynthetic CDR-L3 142Gln Gln Tyr His Asn
Ala Ile Thr 1 5 1438PRTArtificial SequenceSynthetic CDR-L3 143Gln
Gln Tyr Tyr Phe Tyr Ile Thr 1 5 1448PRTArtificial SequenceSynthetic
CDR-L3 144Gln Gln Tyr His Ser Ala Leu Thr 1 5 1458PRTArtificial
SequenceSynthetic CDR-L3 145Gln Gln Tyr His Gly Gly Ile Thr 1 5
1468PRTArtificial SequenceSynthetic CDR-L3 146Gln Gln Tyr His Arg
Arg Ile Thr 1 5 1478PRTArtificial SequenceSynthetic CDR-L3 147Gln
Gln Tyr His Ser Gly Ile Thr 1 5 1489PRTArtificial SequenceSynthetic
CDR-L3 148Gln Gln Tyr Tyr Leu Tyr Pro Leu Thr 1 5 1499PRTArtificial
SequenceSynthetic CDR-L3 149Gln Gln Tyr Trp Thr Tyr Pro Leu Thr 1 5
1509PRTArtificial SequenceSynthetic CDR-L3 150Gln Gln Tyr Leu Leu
Tyr Pro Leu Thr 1 5 1519PRTArtificial SequenceSynthetic CDR-L3
151Gln Gln Tyr Leu Ile Trp Pro Leu Thr 1 5 1529PRTArtificial
SequenceSynthetic CDR-L3 152Gln Gln Tyr Leu Leu Trp Pro Leu Thr 1 5
1539PRTArtificial SequenceSynthetic CDR-L3 153Gln Gln Phe Tyr Phe
Phe Pro Pro Thr 1 5 1549PRTArtificial SequenceSynthetic CDR-L3
154Gln Gln Ala Tyr Thr Phe Pro Pro Thr 1 5 1559PRTArtificial
SequenceSynthetic CDR-L3 155Gln Gln Tyr Tyr Ile Phe Pro Pro Thr 1 5
1569PRTArtificial SequenceSynthetic CDR-L3 156Gln Gln Arg Asn Phe
Tyr Pro Pro Thr 1 5 1579PRTArtificial SequenceSynthetic CDR-L3
157Gln Gln Phe Val Leu Trp Pro Arg Thr 1 5 1589PRTArtificial
SequenceSynthetic CDR-L3 158Gln Gln His Val Asn Phe Pro Leu Thr 1 5
1599PRTArtificial SequenceSynthetic CDR-L3 159Gln Gln Ser Val Phe
Trp Pro Ile Thr 1 5 1609PRTArtificial SequenceSynthetic CDR-L3
160Gln Gln Leu Thr Lys Trp Pro Leu Thr 1 5 1619PRTArtificial
SequenceSynthetic CDR-L3 161Gln Gln Asp Val Leu Trp Pro Leu Thr 1 5
1629PRTArtificial SequenceSynthetic CDR-L3 162Gln Gln Tyr Gly Leu
Phe Pro Ile Thr 1 5 1639PRTArtificial SequenceSynthetic CDR-L3
163Gln Gln His Thr Val Trp Pro Ile Thr 1 5 1649PRTArtificial
SequenceSynthetic CDR-L3 164Gln Gln Val Leu Asn Tyr Pro Leu Thr 1 5
1659PRTArtificial SequenceSynthetic CDR-L3 165Gln Gln Ser Tyr Phe
Leu Pro Pro Thr 1 5 1669PRTArtificial SequenceSynthetic CDR-L3
166Gln Gln Ala His Ser Ser Pro Tyr Thr 1 5 16722PRTArtificial
SequenceSynthetic Leader for scFV, scFv-Fc, Leader 167Met Lys Tyr
Leu Leu Pro Thr Ala Ala Ala Gly Leu Leu Leu Leu Ala 1 5 10 15 Ala
Gln Pro Ala Met Ala 20 16815PRTArtificial SequenceSynthetic Linker
for scFV, scFV-FC, Linker 168Gly Gly Gly Gly Ser Gly Gly Gly Gly
Ser Gly Gly Gly Gly Ser 1 5 10 15 16911PRTArtificial
SequenceSynthetic C-Term Tag for scFV, scFV-FC, C-Term Tag 169Gly
Pro Gly Gly Gln His His His His His His 1 5 10 170231PRTArtificial
SequenceSynthetic C-Term Tag for scFv, scFV-FC, C-Term Tag 170Pro
Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro 1 5 10
15 Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys
20 25 30 Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val
Val Val 35 40 45 Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn
Trp Tyr Val Asp 50 55 60 Gly Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Tyr 65 70 75 80 Asn Ser Thr Tyr Arg Val Val Ser
Val Leu Thr Val Leu His Gln Asp 85 90 95 Trp Leu Asn Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu 100 105 110 Pro Ala Pro Ile
Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg 115 120 125 Glu Pro
Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr Lys 130 135 140
Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp 145
150 155 160 Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys 165 170 175 Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser 180 185 190 Lys Leu Thr Val Asp Lys Ser Arg Trp Gln
Gln Gly Asn Val Phe Ser 195 200 205 Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser 210 215 220 Leu Ser Leu Ser Pro Gly
Lys 225 230 171276PRTArtificial SequenceSynthetic scFv, 29872
171Met Lys Tyr Leu Leu Pro Thr Ala Ala Ala Gly Leu Leu Leu Leu Ala
1 5 10 15 Ala Gln Pro Ala Met Ala Gln Val Gln Leu Val Gln Ser Gly
Ala Glu 20 25 30 Val Lys Glu Pro Gly Ala Ser Val Lys Val Ser Cys
Lys Ala Pro Gly 35 40 45 Tyr Thr Phe Thr Ser Tyr Tyr Met His Trp
Val Arg Gln Ala Pro Gly 50 55 60 Gln Gly Leu Glu Trp Met Gly Val
Ile Asn Pro Ser Gly Gly Ser Thr 65 70 75 80 Ser Tyr Ala Gln Lys Phe
Gln Gly Arg Val Thr Met Thr Arg Asp Thr 85 90 95 Ser Thr Ser Thr
Val Tyr Met Glu Leu Ser Ser Leu Arg Ser Glu Asp 100 105 110 Thr Ala
Val Tyr Tyr Cys Ala Arg Gly Lys Arg Glu Gly Gly Thr Glu 115 120 125
Tyr Leu Arg His Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly 130
135 140 Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Glu
Ile 145 150 155 160 Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser
Pro Gly Glu Arg 165 170 175 Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser
Val Ser Ser Ser Tyr Leu 180 185 190 Ala Trp Tyr Gln Gln Lys Pro Gly
Gln Ala Pro Arg Leu Leu Ile Tyr 195 200 205 Gly Ala Ser Ser Arg Ala
Thr Gly Ile Pro Asp Arg Phe Ser Gly Ser 210 215 220 Gly Ser Gly Thr
Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu Pro Glu 225 230 235 240 Asp
Phe Ala Val Tyr Tyr Cys Gln Gln Tyr His Ser Tyr Ile Thr Phe 245 250
255 Gly Gly Gly Thr Lys Val Glu Ile Lys Gly Pro Gly Gly Gln His His
260 265 270 His His His His 275 172276PRTArtificial
SequenceSynthetic scFv, 31895 172Met Lys Tyr Leu Leu Pro Thr Ala
Ala Ala Gly Leu Leu Leu Leu Ala 1 5 10 15 Ala Gln Pro Ala Met Ala
Gln Val Gln Leu Val Gln Ser Gly Ala Glu 20 25 30 Val Lys Lys Pro
Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly 35 40 45 Tyr Thr
Phe Lys Ser Tyr Glu Met His Trp Val Arg Gln Ala Pro Gly 50 55 60
Gln Gly Leu Glu Trp Met Gly Arg Ile Asn Pro Ser Val Gly Ser Thr 65
70 75 80 Trp Tyr Ala Gln Lys Phe Gln Gly Arg Val Thr Met Thr Arg
Asp Thr 85 90 95 Ser Thr Ser Thr Val Tyr Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp 100 105 110 Thr Ala Val Tyr Tyr Cys Ala Arg Gly Lys
Arg Glu Gly Gly Thr Glu 115 120 125 Tyr Leu Arg Lys Trp Gly Gln Gly
Thr Leu Val Thr Val Ser Ser Gly 130 135 140 Gly Gly Gly Ser Gly Gly
Gly Gly Ser Gly Gly Gly Gly Ser Glu Ile 145 150 155 160 Val Leu Thr
Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly Glu Arg 165 170 175 Ala
Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ala Ser Ser Tyr Leu 180 185
190 Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
195 200 205 Gly Ala Ser Asn Arg His Thr Gly Ile Pro Asp Arg Phe Ser
Gly Ser 210 215 220 Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg
Leu Glu Pro Glu 225 230 235 240 Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Tyr His Asn Ala Ile Thr Phe 245 250 255 Gly Gly Gly Thr Lys Val Glu
Ile Lys Gly Pro Gly Gly Gln His His 260 265 270 His His His His 275
173276PRTArtificial SequenceSynthetic scFv, 31414 173Met Lys Tyr
Leu Leu Pro Thr Ala Ala Ala Gly Leu Leu Leu Leu Ala 1 5 10 15 Ala
Gln Pro Ala Met Ala Gln Val Gln Leu Val Gln Ser Gly Ala Glu 20 25
30 Val Lys Lys Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly
35 40 45 Tyr Thr Phe Lys Ser Tyr Glu Met His Trp Val Arg Gln Ala
Pro Gly 50 55 60 Gln Gly Leu Glu Trp Met Gly Arg Ile Asn Pro Ser
Val Gly Ser Thr 65 70 75 80 Trp Tyr Ala Gln Lys Phe Gln Gly Arg Val
Thr Met Thr Arg Asp Thr 85 90 95 Ser Thr Ser Thr Val Tyr Met Glu
Leu Ser Ser Leu Arg Ser Glu Asp 100 105 110 Thr Ala Val Tyr Tyr Cys
Ala Arg Gly Lys Arg Glu Gly Gly Thr Glu 115 120 125 Tyr Leu Arg Asn
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly 130 135 140 Gly Gly
Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Glu Ile 145 150 155
160 Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly Glu Arg
165 170 175 Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Ser
Tyr Leu 180 185 190 Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg
Leu Leu Ile Tyr 195 200 205 Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro
Asp Arg Phe Ser Gly Ser 210 215 220 Gly Ser Gly Thr Asp Phe Thr Leu
Thr Ile Ser Arg Leu Glu Pro Glu 225 230 235 240 Asp Phe Ala Val Tyr
Tyr Cys Gln Gln Tyr His Ser Tyr Ile Thr Phe 245 250 255 Gly Gly Gly
Thr Lys Val Glu Ile Lys Gly Pro Gly Gly Gln His His 260 265 270 His
His His His 275 174281PRTArtificial SequenceSynthetic scFv, 31905
174Met Lys Tyr Leu Leu Pro Thr Ala Ala Ala Gly Leu Leu Leu Leu Ala
1 5 10 15 Ala Gln Pro Ala Met Ala Gln Val Gln Leu Val Gln Ser Gly
Ala Glu 20 25 30 Val Lys Lys Pro Gly Ser Ser Val Lys Val Ser Cys
Lys Ala Ser Gly 35 40 45 Gly Thr Phe Pro Ser Asn Ala Ile Ser Trp
Val Arg Gln Ala Pro Gly 50 55 60 Gln Gly Leu Glu Trp Met Gly Gly
Ile Gly Phe Gly Thr Ala Asn Tyr 65 70 75 80 Ala Gln Lys Phe Gln Gly
Arg Val Thr Ile Thr Ala Asp Glu Ser Thr 85 90 95 Ser Thr Ala Tyr
Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala 100 105 110 Val Tyr
Tyr Cys Ala Arg Gly Gly Ala Lys Tyr Ala Arg Thr Tyr Gly 115 120 125
Met Asp Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser Gly Gly 130
135 140 Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Asp Ile
Val 145 150 155 160 Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu
Gly Glu Arg Ala 165 170 175 Thr Ile Asn Cys Lys Ser Ser Lys Ser Val
Leu Tyr Ser Asn Asn Asn 180 185 190 Lys Asn Tyr Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Pro Pro Lys 195 200 205 Leu Leu Ile Tyr Trp Ala
Ser Thr Arg Gln Ser Gly Val Pro Asp Arg 210 215 220 Phe Ser Gly Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser 225 230 235 240 Leu
Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln Tyr Leu Leu 245 250
255 Tyr Pro Leu Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Gly Pro
260 265 270 Gly Gly Gln His His His His His His 275 280
175496PRTArtificial SequenceSynthetic scFv-Fc, 29872 175Met Lys Tyr
Leu Leu Pro Thr Ala Ala Ala Gly Leu Leu Leu Leu Ala 1 5 10 15 Ala
Gln Pro Ala Met Ala Gln Val Gln Leu Val Gln Ser Gly Ala Glu 20 25
30 Val Lys Glu Pro Gly Ala Ser Val Lys Val Ser Cys Lys Ala Pro Gly
35 40 45 Tyr Thr Phe Thr Ser Tyr Tyr Met His Trp Val Arg Gln Ala
Pro Gly 50 55 60 Gln Gly Leu Glu Trp Met Gly Val Ile Asn Pro Ser
Gly Gly Ser Thr 65 70 75 80 Ser Tyr Ala Gln Lys Phe Gln Gly Arg Val
Thr Met Thr Arg Asp Thr 85 90 95 Ser Thr Ser Thr Val Tyr Met Glu
Leu Ser Ser Leu Arg Ser Glu Asp 100 105 110 Thr Ala Val Tyr Tyr Cys
Ala Arg Gly Lys Arg Glu Gly Gly Thr Glu 115 120 125 Tyr Leu Arg His
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Gly 130 135 140 Gly Gly
Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Glu Ile 145 150 155
160 Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly Glu Arg
165 170 175 Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Ser
Tyr Leu 180 185 190 Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg
Leu Leu Ile Tyr 195 200 205 Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro
Asp Arg Phe Ser Gly Ser 210 215 220 Gly Ser Gly Thr Asp Phe Thr Leu
Thr Ile Ser Arg Leu Glu Pro Glu 225 230 235 240 Asp Phe Ala Val Tyr
Tyr Cys Gln Gln Tyr His Ser Tyr Ile Thr Phe 245 250 255 Gly Gly Gly
Thr Lys Val Glu Ile Lys Pro Lys Ser Cys Asp Lys Thr 260 265 270 His
Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser 275 280
285 Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
290 295 300 Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu
Asp Pro 305 310 315 320 Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val
Glu Val His Asn Ala 325 330 335 Lys Thr Lys Pro Arg Glu Glu Gln Tyr
Asn Ser Thr Tyr Arg Val Val 340 345 350 Ser Val Leu Thr Val Leu His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr 355 360 365 Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr 370 375 380 Ile Ser Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu 385 390 395 400
Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 405
410 415 Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu
Ser 420 425 430 Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro
Val Leu Asp 435
440 445 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser 450 455 460 Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala 465 470 475 480 Leu His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Pro Gly Lys 485 490 495 176496PRTArtificial
SequenceSynthetic scFv-Fc, 31895 176Met Lys Tyr Leu Leu Pro Thr Ala
Ala Ala Gly Leu Leu Leu Leu Ala 1 5 10 15 Ala Gln Pro Ala Met Ala
Gln Val Gln Leu Val Gln Ser Gly Ala Glu 20 25 30 Val Lys Lys Pro
Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly 35 40 45 Tyr Thr
Phe Lys Ser Tyr Glu Met His Trp Val Arg Gln Ala Pro Gly 50 55 60
Gln Gly Leu Glu Trp Met Gly Arg Ile Asn Pro Ser Val Gly Ser Thr 65
70 75 80 Trp Tyr Ala Gln Lys Phe Gln Gly Arg Val Thr Met Thr Arg
Asp Thr 85 90 95 Ser Thr Ser Thr Val Tyr Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp 100 105 110 Thr Ala Val Tyr Tyr Cys Ala Arg Gly Lys
Arg Glu Gly Gly Thr Glu 115 120 125 Tyr Leu Arg Lys Trp Gly Gln Gly
Thr Leu Val Thr Val Ser Ser Gly 130 135 140 Gly Gly Gly Ser Gly Gly
Gly Gly Ser Gly Gly Gly Gly Ser Glu Ile 145 150 155 160 Val Leu Thr
Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly Glu Arg 165 170 175 Ala
Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ala Ser Ser Tyr Leu 180 185
190 Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
195 200 205 Gly Ala Ser Asn Arg His Thr Gly Ile Pro Asp Arg Phe Ser
Gly Ser 210 215 220 Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg
Leu Glu Pro Glu 225 230 235 240 Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Tyr His Asn Ala Ile Thr Phe 245 250 255 Gly Gly Gly Thr Lys Val Glu
Ile Lys Pro Lys Ser Cys Asp Lys Thr 260 265 270 His Thr Cys Pro Pro
Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser 275 280 285 Val Phe Leu
Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 290 295 300 Thr
Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro 305 310
315 320 Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn
Ala 325 330 335 Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val 340 345 350 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr 355 360 365 Lys Cys Lys Val Ser Asn Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr 370 375 380 Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu 385 390 395 400 Pro Pro Ser Arg
Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 405 410 415 Leu Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 420 425 430
Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 435
440 445 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser 450 455 460 Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala 465 470 475 480 Leu His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Pro Gly Lys 485 490 495 177496PRTArtificial
SequenceSynthetic scFv-Fc, 31414 177Met Lys Tyr Leu Leu Pro Thr Ala
Ala Ala Gly Leu Leu Leu Leu Ala 1 5 10 15 Ala Gln Pro Ala Met Ala
Gln Val Gln Leu Val Gln Ser Gly Ala Glu 20 25 30 Val Lys Lys Pro
Gly Ala Ser Val Lys Val Ser Cys Lys Ala Ser Gly 35 40 45 Tyr Thr
Phe Lys Ser Tyr Glu Met His Trp Val Arg Gln Ala Pro Gly 50 55 60
Gln Gly Leu Glu Trp Met Gly Arg Ile Asn Pro Ser Val Gly Ser Thr 65
70 75 80 Trp Tyr Ala Gln Lys Phe Gln Gly Arg Val Thr Met Thr Arg
Asp Thr 85 90 95 Ser Thr Ser Thr Val Tyr Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp 100 105 110 Thr Ala Val Tyr Tyr Cys Ala Arg Gly Lys
Arg Glu Gly Gly Thr Glu 115 120 125 Tyr Leu Arg Asn Trp Gly Gln Gly
Thr Leu Val Thr Val Ser Ser Gly 130 135 140 Gly Gly Gly Ser Gly Gly
Gly Gly Ser Gly Gly Gly Gly Ser Glu Ile 145 150 155 160 Val Leu Thr
Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly Glu Arg 165 170 175 Ala
Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Ser Tyr Leu 180 185
190 Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
195 200 205 Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe Ser
Gly Ser 210 215 220 Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg
Leu Glu Pro Glu 225 230 235 240 Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Tyr His Ser Tyr Ile Thr Phe 245 250 255 Gly Gly Gly Thr Lys Val Glu
Ile Lys Pro Lys Ser Cys Asp Lys Thr 260 265 270 His Thr Cys Pro Pro
Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser 275 280 285 Val Phe Leu
Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 290 295 300 Thr
Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro 305 310
315 320 Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn
Ala 325 330 335 Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val 340 345 350 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr 355 360 365 Lys Cys Lys Val Ser Asn Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr 370 375 380 Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu 385 390 395 400 Pro Pro Ser Arg
Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 405 410 415 Leu Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 420 425 430
Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 435
440 445 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser 450 455 460 Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala 465 470 475 480 Leu His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Pro Gly Lys 485 490 495 178501PRTArtificial
SequenceSynthetic scFv-Fc, 31905 178Met Lys Tyr Leu Leu Pro Thr Ala
Ala Ala Gly Leu Leu Leu Leu Ala 1 5 10 15 Ala Gln Pro Ala Met Ala
Gln Val Gln Leu Val Gln Ser Gly Ala Glu 20 25 30 Val Lys Lys Pro
Gly Ser Ser Val Lys Val Ser Cys Lys Ala Ser Gly 35 40 45 Gly Thr
Phe Pro Ser Asn Ala Ile Ser Trp Val Arg Gln Ala Pro Gly 50 55 60
Gln Gly Leu Glu Trp Met Gly Gly Ile Gly Phe Gly Thr Ala Asn Tyr 65
70 75 80 Ala Gln Lys Phe Gln Gly Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr 85 90 95 Ser Thr Ala Tyr Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala 100 105 110 Val Tyr Tyr Cys Ala Arg Gly Gly Ala Lys
Tyr Ala Arg Thr Tyr Gly 115 120 125 Met Asp Val Trp Gly Gln Gly Thr
Thr Val Thr Val Ser Ser Gly Gly 130 135 140 Gly Gly Ser Gly Gly Gly
Gly Ser Gly Gly Gly Gly Ser Asp Ile Val 145 150 155 160 Met Thr Gln
Ser Pro Asp Ser Leu Ala Val Ser Leu Gly Glu Arg Ala 165 170 175 Thr
Ile Asn Cys Lys Ser Ser Lys Ser Val Leu Tyr Ser Asn Asn Asn 180 185
190 Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Pro Pro Lys
195 200 205 Leu Leu Ile Tyr Trp Ala Ser Thr Arg Gln Ser Gly Val Pro
Asp Arg 210 215 220 Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu
Thr Ile Ser Ser 225 230 235 240 Leu Gln Ala Glu Asp Val Ala Val Tyr
Tyr Cys Gln Gln Tyr Leu Leu 245 250 255 Tyr Pro Leu Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys Pro Lys 260 265 270 Ser Cys Asp Lys Thr
His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu 275 280 285 Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr 290 295 300 Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val 305 310
315 320 Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
Val 325 330 335 Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn Ser 340 345 350 Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu 355 360 365 Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser Asn Lys Ala Leu Pro Ala 370 375 380 Pro Ile Glu Lys Thr Ile Ser
Lys Ala Lys Gly Gln Pro Arg Glu Pro 385 390 395 400 Gln Val Tyr Thr
Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln 405 410 415 Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala 420 425 430
Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr 435
440 445 Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
Leu 450 455 460 Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe
Ser Cys Ser 465 470 475 480 Val Met His Glu Ala Leu His Asn His Tyr
Thr Gln Lys Ser Leu Ser 485 490 495 Leu Ser Pro Gly Lys 500
179121PRTArtificial SequenceSynthetic VH, 27579 179Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Glu Pro Gly Ala 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Pro Gly Tyr Thr Phe Thr Ser Tyr 20 25 30
Tyr Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Val Ile Asn Pro Ser Gly Gly Ser Thr Ser Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser
Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr
Glu Tyr Leu Arg His Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser 115 120 180121PRTArtificial SequenceSynthetic VH, 31895
180Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Lys
Ser Tyr 20 25 30 Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Arg Ile Asn Pro Ser Val Gly Ser Thr
Trp Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg
Asp Thr Ser Thr Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys
Arg Glu Gly Gly Thr Glu Tyr Leu Arg Lys Trp Gly 100 105 110 Gln Gly
Thr Leu Val Thr Val Ser Ser 115 120 181121PRTArtificial
SequenceSynthetic VH, 31415 181Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30 Gln Met His Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Arg Ile
Asn Pro Ser Gly Gly Ser Thr Trp Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr Glu Tyr Leu Arg
Ser Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val Ser Ser 115 120
182121PRTArtificial SequenceSynthetic VH, 31414 182Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Lys Ser Tyr 20 25 30
Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Arg Ile Asn Pro Ser Val Gly Ser Thr Trp Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser
Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr
Glu Tyr Leu Arg Asn Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser 115 120 183121PRTArtificial SequenceSynthetic VH, 31891
183Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Ile Phe Lys
Ser Tyr 20 25 30 Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Arg Ile Asn Pro Ser Val Gly Ser Thr
Trp Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg
Asp Thr Ser Thr Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys
Arg Glu Gly Gly Thr Glu Tyr Leu Arg Val Trp Gly 100 105 110 Gln Gly
Thr Leu Val Thr Val Ser Ser 115 120 184121PRTArtificial
SequenceSynthetic VH, 29871 184Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Gln Ser Tyr 20 25 30 Tyr Met His Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Lys Ile
Asn Pro Ser Gly Gly Ser Thr Trp Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr
Glu Tyr Leu Arg His Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser 115 120 185121PRTArtificial SequenceSynthetic VH, 31418
185Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Lys
Ser Tyr 20 25 30 Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Arg Ile Asn Pro Ser Gly Gly Ser Thr
Trp Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg
Asp Thr Ser Thr Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys
Arg Glu Gly Gly Thr Glu Tyr Leu Arg His Trp Gly 100 105 110 Gln Gly
Thr Leu Val Thr Val Ser Ser 115 120 186121PRTArtificial
SequenceSynthetic VH, 31431 186Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30 Gln Met His Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Arg Ile
Asn Pro Ser Gly Gly Ser Thr Trp Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr Glu Tyr Leu Arg
His Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val Ser Ser 115 120
187121PRTArtificial SequenceSynthetic VH, 31421 187Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Phe Ser Tyr 20 25 30
Tyr Met Tyr Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Val Ile Asn Pro Leu Gly Gly Gly Thr Ser Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser
Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr
Glu Tyr Leu Arg His Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser 115 120 188121PRTArtificial SequenceSynthetic VH, 31429
188Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Val
Ser Tyr 20 25 30 Phe Met His Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Ser Ile Asn Pro Arg Gly Gly Ser Thr
Ser Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg
Asp Thr Ser Thr Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys
Arg Glu Gly Gly Thr Glu Tyr Leu Arg His Trp Gly 100 105 110 Gln Gly
Thr Leu Val Thr Val Ser Ser 115 120 189121PRTArtificial
SequenceSynthetic VH, 29872 189Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Lys Ser Tyr 20 25 30 Glu Met His Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Arg Ile
Asn Pro Ser Val Gly Ser Thr Trp Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr Glu Tyr Leu Arg
His Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val Ser Ser 115 120
190122PRTArtificial SequenceSynthetic VH, 28347 190Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Leu 20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Asn
Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Gly Ala Lys Tyr Ala Ser
Thr Tyr Gly Met Asp Val Trp 100 105 110 Gly Gln Gly Thr Thr Val Thr
Val Ser Ser 115 120 191120PRTArtificial SequenceSynthetic VH, 31896
191Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser
Lys Leu 20 25 30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Gly Ile Gly Phe Gly Thr Ala Asn Tyr
Ala Gln Lys Phe Gln Gly 50 55 60 Arg Val Thr Ile Thr Ala Asp Glu
Ser Ala Ser Thr Ala Tyr Met Glu 65 70 75 80 Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg 85 90 95 Gly Gly Ala Lys
Tyr Ala Ser Thr His Gly Met Asp Val Trp Gly Gln 100 105 110 Gly Thr
Thr Val Thr Val Ser Ser 115 120 192122PRTArtificial
SequenceSynthetic VH, 31432 192Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser His Thr 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Leu Pro Ile Gly Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Gly Gly Ala Lys Tyr Ala Ser Gln Leu Gly Met
Asp Val Trp 100 105 110 Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115
120 193120PRTArtificial SequenceSynthetic VH, 31915 193Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Leu 20 25
30 Pro Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Gly Ile Gly Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe
Gln Gly 50 55 60 Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr
Ala Tyr Met Glu 65 70 75 80 Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala
Val Tyr Tyr Cys Ala Arg 85 90 95 Gly Gly Ala Lys Tyr Ala Ser Lys
Trp Gly Met Asp Val Trp Gly Gln 100 105 110 Gly Thr Thr Val Thr Val
Ser Ser 115 120 194122PRTArtificial SequenceSynthetic VH, 31436
194Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser
Leu Leu 20 25 30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Gly Ile Leu Pro Ile Ala Gly Thr Ala
Asn Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala
Asp Glu Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Gly
Ala Lys Tyr Ala Val Gly Tyr Gly Met Asp Val Trp 100 105 110 Gly Gln
Gly Thr Thr Val Thr Val Ser Ser 115 120 195122PRTArtificial
SequenceSynthetic VH, 31437 195Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Gln Ser Leu 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Leu Pro Ile Gly Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Gly Gly Ala Lys Tyr Ala Gly Arg Tyr Gly Met
Asp Val Trp 100 105 110 Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115
120 196120PRTArtificial SequenceSynthetic VH, 31905 196Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Pro Ser Asn 20 25
30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Gly Ile Gly Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe
Gln Gly 50 55 60 Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr
Ala Tyr Met Glu 65 70 75 80 Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala
Val Tyr Tyr Cys Ala Arg 85 90 95 Gly Gly Ala Lys Tyr Ala Arg Thr
Tyr Gly Met Asp Val Trp Gly Gln 100 105 110 Gly Thr Thr Val Thr Val
Ser Ser 115 120 197120PRTArtificial SequenceSynthetic VH, 31901
197Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser
Ser Leu 20 25 30 Pro Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Gly Ile Gly Phe Gly Thr Ala Asn Tyr
Ala Gln Lys Phe Gln Gly 50 55 60 Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr Ser Thr Ala Tyr Met Glu 65 70 75 80 Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg 85 90 95 Gly Gly Ala Lys
Tyr Ala Gly Arg Tyr Gly Met Asp Val Trp Gly Gln 100 105 110 Gly Thr
Thr Val Thr Val Ser Ser 115 120 198122PRTArtificial
SequenceSynthetic VH, 29852 198Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ala Met 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Leu Pro Ile Ala Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Gly Gly Ala Lys Tyr Ala Ser Thr Tyr Gly Met
Asp Val Trp 100 105 110 Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115
120 199122PRTArtificial SequenceSynthetic VH, 29851 199Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ala Ser Leu 20 25
30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Gly Ile Leu Pro Ile Phe Gly Glu Ala Asn Tyr Ala Gln
Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr
Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Gly Ala Lys Tyr Ala
Ser Thr Tyr Gly Met Asp Val Trp 100 105 110 Gly Gln Gly Thr Thr Val
Thr Val Ser Ser 115 120 200122PRTArtificial SequenceSynthetic VH,
29857 200Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro
Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr
Phe Ser Trp Leu 20 25 30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly
Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile Ile Pro Arg Gly Gly
Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile
Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser
Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg
Gly Gly Ala Lys Tyr Ala Ser Thr Tyr Gly Met Asp Val Trp 100 105 110
Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120 201121PRTArtificial
SequenceSynthetic VH, 27571 201Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Ala Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Ser Ile
Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Arg
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Thr Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Glu Ser Gly Gly Tyr Arg Asp His Arg Leu Asp
Val Trp Gly 100 105 110 Gln Gly Thr Met Val Thr Val Ser Ser 115 120
202121PRTArtificial SequenceSynthetic VH, 31861 202Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Gly Ser Tyr 20 25 30
Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Ser Ile Ile Pro Glu Phe Gly Ile Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Glu Ser Gly Thr Tyr Arg Asp His Arg Leu Asp
Val Trp Gly 100 105 110 Gln Gly Thr Met Val Thr Val Ser Ser 115 120
203121PRTArtificial SequenceSynthetic VH, 31873 203Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Lys Tyr 20 25 30
Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Ser Ile Ile Pro Glu Phe Gly Ile Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Glu Ser Gly Gly Tyr Arg Asp
His Arg Leu Gly Val Trp Gly 100 105 110 Gln Gly Thr Met Val Thr Val
Ser Ser 115 120 204121PRTArtificial SequenceSynthetic VH, 31393
204Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Glu
Ser Tyr 20 25 30 Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Ser Ile Ile Pro Glu Phe Gly Ile Ala
Asn Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala
Asp Glu Ser Thr Ser Thr Thr Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Glu Ser
Gly Gly Tyr Arg Asp His Arg Leu Asp Val Trp Gly 100 105 110 Gln Gly
Thr Met Val Thr Val Ser Ser 115 120 205124PRTArtificial
SequenceSynthetic VH, 27534 205Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Asp Phe Thr Asp Tyr Ser Ser Gly Tyr Ser Ser
Gly Trp Thr 100 105 110 Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser
Ser 115 120 206123PRTArtificial SequenceSynthetic VH, 27536 206Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10
15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Asn Tyr
20 25 30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45 Gly Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr
Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Thr Leu Tyr
Ser Ser Gly Ala Tyr Tyr Gly Tyr Asn Val 100 105 110 Trp Gly Gln Gly
Thr Met Val Thr Val Ser Ser 115 120 207124PRTArtificial
SequenceSynthetic VH, 27588 207Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Asn Tyr 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Ala Lys Arg Gly Tyr Asp Ser Tyr Gly Gly Val
Tyr Phe Asp 100 105 110 Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser
Ser 115 120 208127PRTArtificial SequenceSynthetic VH, 27590 208Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10
15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Asn Tyr
20 25 30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45 Gly Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr
Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Pro Thr Val
Thr Ala Thr Thr Ser Ile Gly Thr His Asn 100 105 110 Trp Phe Asp Pro
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser 115 120 125
209126PRTArtificial SequenceSynthetic VH, 27597 209Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Ser Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Glu Gly Arg Gly Tyr Asp Ser
Ser Arg Tyr Tyr Lys Phe Trp 100 105 110 Phe Asp Pro Trp Gly Gln Gly
Thr Leu Val Thr Val Ser Ser 115 120 125 210121PRTArtificial
SequenceSynthetic VH, 27575 210Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Glu Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30 Ala Thr Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Ile Pro Ile Ser Gly Thr Ala Asn Tyr Ala Gln Glu Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Asp Gly Gly Gly Tyr Arg His His Tyr Phe Asp
Leu Trp Gly 100 105 110 Arg Gly Thr Leu Val Thr Val Ser Ser 115 120
211121PRTArtificial SequenceSynthetic VH, 27568 211Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Pro Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Glu Gln Gly Leu Glu Trp Met 35
40 45 Gly Ser Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Gly Glu Ser Gly Gly Tyr Arg Asp
His Lys Leu Asp Val Trp Gly 100 105 110 Gln Gly Thr Val Val Thr Val
Ser Ser 115 120 212121PRTArtificial SequenceSynthetic VH, 27577
212Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Ala Phe Ser
Ser Tyr 20 25 30 Ala Ile Gly Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Gly Ile Ile Pro Thr Phe Gly Thr Ala
Asn Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala
Asp Glu Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu
Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Gly
Gly Gly Tyr Gln His His Tyr Phe Asp Leu Trp Gly 100 105 110 Arg Gly
Thr Leu Val Thr Val Ser Ser 115 120 213121PRTArtificial
SequenceSynthetic VH, 27587 213Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Ser Ile
Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Glu Ser Gly Gly Tyr Arg Asp His Lys Leu Asp
Val Trp Gly 100 105 110 Gln Gly Thr Met Val Thr Val Ser Ser 115 120
214120PRTArtificial SequenceSynthetic VH, 27589 214Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Ser Gly Tyr His Arg His
Tyr Ser Asp Tyr Trp Gly Gln 100 105 110 Gly Thr Leu Val Thr Val Ser
Ser 115 120 215121PRTArtificial SequenceSynthetic VH, 27596 215Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10
15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr
20 25 30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45 Gly Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr
Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Pro Leu Gly
Ile Arg Lys His Trp Phe Asp Pro Trp Gly 100 105 110 Gln Gly Thr Leu
Val Thr Val Ser Ser 115 120 216121PRTArtificial SequenceSynthetic
VH, 27535 216Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys
Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly
Thr Phe Ser Ser Tyr 20 25 30 Ala Ile Ser Trp Val Arg Gln Ala Pro
Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile Ile Pro Ile Phe
Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr
Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu
Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala
Arg Asp Thr Pro Arg Trp Arg Tyr His Tyr Phe Asp Tyr Trp Gly 100 105
110 Gln Gly Thr Leu Val Thr Val Ser Ser 115 120 217121PRTArtificial
SequenceSynthetic VH, 27550 217Glu Val Gln Leu Val Glu Ser Gly Gly
Gly Leu Val Lys Pro Gly Gly 1 5 10 15 Ser Leu Arg Leu Ser Cys Ala
Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30 Ser Met Asn Trp Val
Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40 45 Ser Ser Ile
Ser Ser Ser Ser Ser Tyr Ile Tyr Tyr Ala Asp Ser Val 50 55 60 Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr 65 70
75 80 Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Glu Arg Arg Gly Ser Leu Ala Leu Gly Met Asp
Val Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val Ser Ser 115 120
218128PRTArtificial SequenceSynthetic VH, 27549 218Gln Val Gln Leu
Val Glu Ser Gly Gly Gly Val Val Gln Pro Gly Arg 1 5 10 15 Ser Leu
Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30
Gly Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35
40 45 Ala Val Ile Trp Tyr Asp Gly Ser Asn Lys Tyr Tyr Ala Asp Ser
Val 50 55 60 Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn
Thr Leu Tyr 65 70 75 80 Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Leu Gly Gly Tyr Ser Tyr
Gly Glu Pro Tyr Tyr Tyr Tyr 100 105 110 Tyr Gly Met Asp Val Trp Gly
Gln Gly Thr Thr Val Thr Val Ser Ser 115 120 125 219107PRTArtificial
SequenceSynthetic VL, 27579 219Glu Ile Val Leu Thr Gln Ser Pro Gly
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Ser Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Gly
Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70
75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr His Ser Tyr
Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105
220107PRTArtificial SequenceSynthetic VL, 31895 220Glu Ile Val Leu
Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg
Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ala Ser Ser 20 25 30
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35
40 45 Ile Tyr Gly Ala Ser Asn Arg His Thr Gly Ile Pro Asp Arg Phe
Ser 50 55
60 Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu
65 70 75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr His Asn
Ala Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100
105 221107PRTArtificial SequenceSynthetic VL, 31891 221Glu Ile Val
Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Ser 20 25
30 Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu
35 40 45 Ile Tyr Tyr Ala Ser Ser Arg Ala Tyr Gly Ile Pro Asp Arg
Phe Ser 50 55 60 Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile
Ser Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln
Gln Tyr His Asn Ala Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val
Glu Ile Lys 100 105 222107PRTArtificial SequenceSynthetic VL, 31418
222Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly
1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser
Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala
Pro Arg Leu Leu 35 40 45 Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly
Ile Pro Asp Arg Phe Ser 50 55 60 Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe Ala Val
Tyr Tyr Cys Gln Gln Tyr Tyr Phe Tyr Ile 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys 100 105 223107PRTArtificial
SequenceSynthetic VL, 31430 223Glu Ile Val Leu Thr Gln Ser Pro Gly
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Glu Ala Ser Gln Ser Val Ser Tyr Ser 20 25 30 Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Gly
Ala Ser Ser Arg Ala Asn Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70
75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr His Ser Ala
Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105
224107PRTArtificial SequenceSynthetic VL, 31431 224Glu Ile Val Leu
Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg
Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ala Ser Ser 20 25 30
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35
40 45 Ile Tyr Gly Ala Ser Asn Arg His Thr Gly Ile Pro Asp Arg Phe
Ser 50 55 60 Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Tyr His Gly Gly Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys 100 105 225107PRTArtificial SequenceSynthetic VL, 31421
225Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly
1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Lys Ala Ser Glu Ser Val Ser
Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala
Pro Arg Leu Leu 35 40 45 Ile Tyr Tyr Ala Ser Ser Arg Ala Thr Gly
Ile Pro Asp Arg Phe Ser 50 55 60 Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe Ala Val
Tyr Tyr Cys Gln Gln Tyr His Arg Arg Ile 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys 100 105 226107PRTArtificial
SequenceSynthetic VL, 31429 226Glu Ile Val Leu Thr Gln Ser Pro Gly
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Tyr Val Ser Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Tyr
Ala Ser Asn Arg Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70
75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr His Ser Gly
Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105
227113PRTArtificial SequenceSynthetic VL, 28347 227Asp Ile Val Met
Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly 1 5 10 15 Glu Arg
Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val Leu Phe Ser 20 25 30
Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35
40 45 Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly
Val 50 55 60 Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr 65 70 75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val
Tyr Tyr Cys Gln Gln 85 90 95 Tyr Tyr Leu Tyr Pro Leu Thr Phe Gly
Gly Gly Thr Lys Val Glu Ile 100 105 110 Lys 228113PRTArtificial
SequenceSynthetic VL, 31896 228Asp Ile Val Met Thr Gln Ser Pro Asp
Ser Leu Ala Val Ser Leu Gly 1 5 10 15 Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Arg Ser Val Leu Phe Ser 20 25 30 Ser Asn Asn Lys Asn
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45 Pro Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60 Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr 65 70
75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln
Gln 85 90 95 Tyr Trp Thr Tyr Pro Leu Thr Phe Gly Gly Gly Thr Lys
Val Glu Ile 100 105 110 Lys 229113PRTArtificial SequenceSynthetic
VL, 31915 229Asp Ile Val Met Thr Gln Ser Pro Asp Ser Leu Ala Val
Ser Leu Gly 1 5 10 15 Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln
Ser Val Leu Phe Ser 20 25 30 Ser Asn Asn Lys Asn Tyr Leu Ala Trp
Tyr Gln Gln Lys Pro Gly Gln 35 40 45 Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Ser Arg Glu Ser Gly Val 50 55 60 Pro Asp Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr 65 70 75 80 Ile Ser Ser
Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95 Tyr
Trp Thr Tyr Pro Leu Thr Phe Gly Gly Gly Thr Lys Val Glu Ile 100 105
110 Lys 230113PRTArtificial SequenceSynthetic VL, 31905 230Asp Ile
Val Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly 1 5 10 15
Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Lys Ser Val Leu Tyr Ser 20
25 30 Asn Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly
Gln 35 40 45 Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Gln
Ser Gly Val 50 55 60 Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr
Asp Phe Thr Leu Thr 65 70 75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val
Ala Val Tyr Tyr Cys Gln Gln 85 90 95 Tyr Leu Leu Tyr Pro Leu Thr
Phe Gly Gly Gly Thr Lys Val Glu Ile 100 105 110 Lys
231113PRTArtificial SequenceSynthetic VL, 31901 231Gly Ile Val Met
Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly 1 5 10 15 Glu Arg
Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val Leu Phe Ser 20 25 30
Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35
40 45 Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Ala Ser Gly
Val 50 55 60 Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr 65 70 75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val
Tyr Tyr Cys Gln Gln 85 90 95 Tyr Tyr Leu Tyr Pro Leu Thr Phe Gly
Gly Gly Thr Lys Val Glu Ile 100 105 110 Lys 232107PRTArtificial
SequenceSynthetic VL, 27571 232Glu Ile Val Met Thr Gln Ser Pro Ala
Thr Leu Ser Val Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Gly Ser Asn 20 25 30 Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Gly Ala
Ser Thr Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser
Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Ser 65 70
75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr Leu Ile Trp Pro
Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105
233107PRTArtificial SequenceSynthetic VL, 31861 233Glu Ile Val Met
Thr Gln Ser Pro Ala Thr Leu Ser Val Ser Pro Gly 1 5 10 15 Glu Arg
Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Gly Ser Asn 20 25 30
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35
40 45 Tyr Gly Ala Ser Thr Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser
Gly 50 55 60 Ser Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser
Leu Gln Ser 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr
Leu Leu Trp Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys 100 105 234107PRTArtificial SequenceSynthetic VL, 31873
234Glu Ile Val Met Thr Gln Ser Pro Ala Thr Leu Ser Val Ser Pro Gly
1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Gly
Ser Asn 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro
Arg Leu Leu Ile 35 40 45 Tyr Gly Ala Ser Thr Arg Ala Ser Gly Ile
Pro Ala Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Glu Phe Thr
Leu Thr Ile Ser Ser Leu Gln Ser 65 70 75 80 Glu Asp Phe Ala Val Tyr
Tyr Cys Gln Gln Tyr Leu Leu Trp Pro Leu 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys 100 105 235113PRTArtificial
SequenceSynthetic VL, 28337 235Asp Ile Val Met Thr Gln Ser Pro Asp
Ser Leu Ala Val Ser Leu Gly 1 5 10 15 Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Val Leu Phe Ser 20 25 30 Ser Asn Asn Lys Asn
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40 45 Pro Pro Lys
Leu Leu Ile Tyr Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60 Pro
Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr 65 70
75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln
Gln 85 90 95 Phe Tyr Phe Tyr Pro Pro Thr Phe Gly Gly Gly Thr Lys
Val Glu Ile 100 105 110 Lys 236113PRTArtificial SequenceSynthetic
VL, 27536 236Asp Ile Val Met Thr Gln Ser Pro Asp Ser Leu Ala Val
Ser Leu Gly 1 5 10 15 Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln
Ser Val Leu Tyr Ser 20 25 30 Ser Asn Asn Lys Asn Tyr Leu Ala Trp
Tyr Gln Gln Lys Pro Gly Gln 35 40 45 Pro Pro Lys Leu Leu Ile Tyr
Trp Ala Ser Thr Arg Glu Ser Gly Val 50 55 60 Pro Asp Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr 65 70 75 80 Ile Ser Ser
Leu Gln Ala Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95 Ala
Tyr Thr Phe Pro Pro Thr Phe Gly Gly Gly Thr Lys Val Glu Ile 100 105
110 Lys 237107PRTArtificial SequenceSynthetic VL, 27588 237Asp Ile
Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly 1 5 10 15
Asp Arg Val Thr Ile Thr Cys Gln Ala Ser Gln Asp Ile Ser Asn Tyr 20
25 30 Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu
Ile 35 40 45 Tyr Asp Ala Ser Asn Leu Glu Thr Gly Val Pro Ser Arg
Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr Phe Thr Ile
Ser Ser Leu Gln Pro 65 70 75 80 Glu Asp Ile Ala Thr Tyr Tyr Cys Gln
Gln Tyr Tyr Ile Phe Pro Pro 85 90 95 Thr Phe Gly Gly Gly Thr Lys
Val Glu Ile Lys 100 105 238107PRTArtificial SequenceSynthetic VL,
27590 238Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser
Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser
Val Ser Ser Tyr 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln
Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Asp Ala Ser Asn Arg Ala Thr
Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala
Val Tyr Tyr Cys Gln Gln Arg Asn Phe Tyr Pro Pro 85 90 95 Thr Phe
Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105 239107PRTArtificial
SequenceSynthetic VL, 27597 239Glu Ile Val Leu Thr Gln Ser Pro Ala
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Ser Ser Tyr 20 25 30 Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Asp Ala
Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro 65 70
75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Val Leu Trp Pro
Arg 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105
240107PRTArtificial SequenceSynthetic VL, 27575 240Glu Ile Val Leu
Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg
Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Arg Tyr 20 25 30
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35
40 45 Tyr Asp Ala Ser Asn Arg Ala Thr Gly Ile Pro Ala
Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr
Ile Ser Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys
Gln Gln His Val Asn Phe Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr
Lys Val Glu Ile Lys 100 105 241107PRTArtificial SequenceSynthetic
VL, 27568 241Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu
Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln
Ser Val Ser Ser Tyr 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly
Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Asp Ala Ser Asn Arg Ala
Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr
Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe
Ala Val Tyr Tyr Cys Gln Gln Ser Val Phe Trp Pro Ile 85 90 95 Thr
Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105 242107PRTArtificial
SequenceSynthetic VL, 27577 242Glu Ile Val Met Thr Gln Ser Pro Ala
Thr Leu Ser Val Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Gly Ser Asn 20 25 30 Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Gly Ala
Ser Thr Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser
Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Ser 65 70
75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Leu Thr Lys Trp Pro
Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105
243107PRTArtificial SequenceSynthetic VL, 27587 243Glu Ile Val Leu
Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg
Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Tyr 20 25 30
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35
40 45 Tyr Asp Ala Ser Lys Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser
Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser
Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Asp
Val Leu Trp Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys 100 105 244107PRTArtificial SequenceSynthetic VL, 27589
244Asp Ile Gln Met Thr Gln Ser Pro Ser Thr Leu Ser Ala Ser Val Gly
1 5 10 15 Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser
Ser Trp 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro
Lys Leu Leu Ile 35 40 45 Tyr Lys Ala Ser Ser Leu Glu Ser Gly Val
Pro Ser Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Glu Phe Thr
Leu Thr Ile Ser Ser Leu Gln Pro 65 70 75 80 Asp Asp Phe Ala Thr Tyr
Tyr Cys Gln Gln Tyr Gly Leu Phe Pro Ile 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys 100 105 245107PRTArtificial
SequenceSynthetic VL, 27596 245Glu Ile Val Met Thr Gln Ser Pro Ala
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Ser Ser Tyr 20 25 30 Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Asp Ala
Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro 65 70
75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln His Thr Val Trp Pro
Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100 105
246107PRTArtificial SequenceSynthetic VL, 27535 246Glu Ile Val Leu
Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg
Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Tyr 20 25 30
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35
40 45 Tyr Asp Ala Ser Lys Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser
Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser
Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Val
Leu Asn Tyr Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu
Ile Lys 100 105 247107PRTArtificial SequenceSynthetic VL, 27550
247Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly
1 5 10 15 Asp Arg Val Thr Ile Thr Cys Gln Ala Ser Gln Asp Ile Ser
Asn Tyr 20 25 30 Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro
Lys Leu Leu Ile 35 40 45 Tyr Asp Ala Ser Asn Leu Glu Thr Gly Val
Pro Ser Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr
Phe Thr Ile Ser Ser Leu Gln Pro 65 70 75 80 Glu Asp Ile Ala Thr Tyr
Tyr Cys Gln Gln Ser Tyr Phe Leu Pro Pro 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys 100 105 248108PRTArtificial
SequenceSynthetic VL, 27549 248Glu Ile Val Leu Thr Gln Ser Pro Gly
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Ser Ser Asp 20 25 30 Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Gly
Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70
75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Ala His Ser Ser
Pro 85 90 95 Tyr Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100
105 249510PRTArtificial SequenceSynthetic hCD39 249Met Glu Asp Thr
Lys Glu Ser Asn Val Lys Thr Phe Cys Ser Lys Asn 1 5 10 15 Ile Leu
Ala Ile Leu Gly Phe Ser Ser Ile Ile Ala Val Ile Ala Leu 20 25 30
Leu Ala Val Gly Leu Thr Gln Asn Lys Ala Leu Pro Glu Asn Val Lys 35
40 45 Tyr Gly Ile Val Leu Asp Ala Gly Ser Ser His Thr Ser Leu Tyr
Ile 50 55 60 Tyr Lys Trp Pro Ala Glu Lys Glu Asn Asp Thr Gly Val
Val His Gln 65 70 75 80 Val Glu Glu Cys Arg Val Lys Gly Pro Gly Ile
Ser Lys Phe Val Gln 85 90 95 Lys Val Asn Glu Ile Gly Ile Tyr Leu
Thr Asp Cys Met Glu Arg Ala 100 105 110 Arg Glu Val Ile Pro Arg Ser
Gln His Gln Glu Thr Pro Val Tyr Leu 115 120 125 Gly Ala Thr Ala Gly
Met Arg Leu Leu Arg Met Glu Ser Glu Glu Leu 130 135 140 Ala Asp Arg
Val Leu Asp Val Val Glu Arg Ser Leu Ser Asn Tyr Pro 145 150 155 160
Phe Asp Phe Gln Gly Ala Arg Ile Ile Thr Gly Gln Glu Glu Gly Ala 165
170 175 Tyr Gly Trp Ile Thr Ile Asn Tyr Leu Leu Gly Lys Phe Ser Gln
Lys 180 185 190 Thr Arg Trp Phe Ser Ile Val Pro Tyr Glu Thr Asn Asn
Gln Glu Thr 195 200 205 Phe Gly Ala Leu Asp Leu Gly Gly Ala Ser Thr
Gln Val Thr Phe Val 210 215 220 Pro Gln Asn Gln Thr Ile Glu Ser Pro
Asp Asn Ala Leu Gln Phe Arg 225 230 235 240 Leu Tyr Gly Lys Asp Tyr
Asn Val Tyr Thr His Ser Phe Leu Cys Tyr 245 250 255 Gly Lys Asp Gln
Ala Leu Trp Gln Lys Leu Ala Lys Asp Ile Gln Val 260 265 270 Ala Ser
Asn Glu Ile Leu Arg Asp Pro Cys Phe His Pro Gly Tyr Lys 275 280 285
Lys Val Val Asn Val Ser Asp Leu Tyr Lys Thr Pro Cys Thr Lys Arg 290
295 300 Phe Glu Met Thr Leu Pro Phe Gln Gln Phe Glu Ile Gln Gly Ile
Gly 305 310 315 320 Asn Tyr Gln Gln Cys His Gln Ser Ile Leu Glu Leu
Phe Asn Thr Ser 325 330 335 Tyr Cys Pro Tyr Ser Gln Cys Ala Phe Asn
Gly Ile Phe Leu Pro Pro 340 345 350 Leu Gln Gly Asp Phe Gly Ala Phe
Ser Ala Phe Tyr Phe Val Met Lys 355 360 365 Phe Leu Asn Leu Thr Ser
Glu Lys Val Ser Gln Glu Lys Val Thr Glu 370 375 380 Met Met Lys Lys
Phe Cys Ala Gln Pro Trp Glu Glu Ile Lys Thr Ser 385 390 395 400 Tyr
Ala Gly Val Lys Glu Lys Tyr Leu Ser Glu Tyr Cys Phe Ser Gly 405 410
415 Thr Tyr Ile Leu Ser Leu Leu Leu Gln Gly Tyr His Phe Thr Ala Asp
420 425 430 Ser Trp Glu His Ile His Phe Ile Gly Lys Ile Gln Gly Ser
Asp Ala 435 440 445 Gly Trp Thr Leu Gly Tyr Met Leu Asn Leu Thr Asn
Met Ile Pro Ala 450 455 460 Glu Gln Pro Leu Ser Thr Pro Leu Ser His
Ser Thr Tyr Val Phe Leu 465 470 475 480 Met Val Leu Phe Ser Leu Val
Leu Phe Thr Val Ala Ile Ile Gly Leu 485 490 495 Leu Ile Phe His Lys
Pro Ser Tyr Phe Trp Lys Asp Met Val 500 505 510 250510PRTArtificial
SequenceSynthetic mCD39 250Met Glu Asp Ile Lys Asp Ser Lys Val Lys
Arg Phe Cys Ser Lys Asn 1 5 10 15 Ile Leu Ile Ile Leu Gly Phe Thr
Ser Ile Leu Ala Val Ile Ala Leu 20 25 30 Ile Ala Val Gly Leu Thr
Gln Asn Lys Pro Leu Pro Glu Asn Val Lys 35 40 45 Tyr Gly Ile Val
Leu Asp Ala Gly Ser Ser His Thr Asn Leu Tyr Ile 50 55 60 Tyr Lys
Trp Pro Ala Glu Lys Glu Asn Asp Thr Gly Val Val Gln Gln 65 70 75 80
Leu Glu Glu Cys Gln Val Lys Gly Pro Gly Ile Ser Lys Tyr Ala Gln 85
90 95 Lys Thr Asp Glu Ile Gly Ala Tyr Leu Ala Glu Cys Met Glu Leu
Ser 100 105 110 Thr Glu Leu Ile Pro Thr Ser Lys His His Gln Thr Pro
Val Tyr Leu 115 120 125 Gly Ala Thr Ala Gly Met Arg Leu Leu Arg Met
Glu Ser Glu Gln Ser 130 135 140 Ala Asp Glu Val Leu Ala Ala Val Ser
Thr Ser Leu Lys Ser Tyr Pro 145 150 155 160 Phe Asp Phe Gln Gly Ala
Lys Ile Ile Thr Gly Gln Glu Glu Gly Ala 165 170 175 Tyr Gly Trp Ile
Thr Ile Asn Tyr Leu Leu Gly Arg Phe Thr Gln Glu 180 185 190 Gln Ser
Trp Leu Ser Leu Ile Ser Asp Ser Gln Lys Gln Glu Thr Phe 195 200 205
Gly Ala Leu Asp Leu Gly Gly Ala Ser Thr Gln Ile Thr Phe Val Pro 210
215 220 Gln Asn Ser Thr Ile Glu Ser Pro Glu Asn Ser Leu Gln Phe Arg
Leu 225 230 235 240 Tyr Gly Glu Asp Tyr Thr Val Tyr Thr His Ser Phe
Leu Cys Tyr Gly 245 250 255 Lys Asp Gln Ala Leu Trp Gln Lys Leu Ala
Lys Asp Ile Gln Val Ser 260 265 270 Ser Gly Gly Val Leu Lys Asp Pro
Cys Phe Asn Pro Gly Tyr Glu Lys 275 280 285 Val Val Asn Val Ser Glu
Leu Tyr Gly Thr Pro Cys Thr Lys Arg Phe 290 295 300 Glu Lys Lys Leu
Pro Phe Asp Gln Phe Arg Ile Gln Gly Thr Gly Asp 305 310 315 320 Tyr
Glu Gln Cys His Gln Ser Ile Leu Glu Leu Phe Asn Asn Ser His 325 330
335 Cys Pro Tyr Ser Gln Cys Ala Phe Asn Gly Val Phe Leu Pro Pro Leu
340 345 350 His Gly Ser Phe Gly Ala Phe Ser Ala Phe Tyr Phe Val Met
Asp Phe 355 360 365 Phe Lys Lys Val Ala Lys Asn Ser Val Ile Ser Gln
Glu Lys Met Thr 370 375 380 Glu Ile Thr Lys Asn Phe Cys Ser Lys Ser
Trp Glu Glu Thr Lys Thr 385 390 395 400 Ser Tyr Pro Ser Val Lys Glu
Lys Tyr Leu Ser Glu Tyr Cys Phe Ser 405 410 415 Gly Ala Tyr Ile Leu
Ser Leu Leu Gln Gly Tyr Asn Phe Thr Asp Ser 420 425 430 Ser Trp Glu
Gln Ile His Phe Met Gly Lys Ile Lys Asp Ser Asn Ala 435 440 445 Gly
Trp Thr Leu Gly Tyr Met Leu Asn Leu Thr Asn Met Ile Pro Ala 450 455
460 Glu Gln Pro Leu Ser Pro Pro Leu Pro His Ser Thr Tyr Ile Gly Leu
465 470 475 480 Met Val Leu Phe Ser Leu Leu Leu Val Ala Val Ala Ile
Thr Gly Leu 485 490 495 Phe Ile Tyr Ser Lys Pro Ser Tyr Phe Trp Lys
Glu Ala Val 500 505 510 251532PRTMacaca
fascicularismisc_feature(1)..(532)Macaca fascicularis cCD39 251Met
Leu Phe Asp Ser Ile Leu Ser Thr Val Gly Leu Ser Lys Leu Val 1 5 10
15 Ser Val Val Ser Ser Pro Ala Ala Ala Leu Ser Lys Ser Asn Val Lys
20 25 30 Thr Phe Cys Ser Lys Asn Ile Leu Ala Ile Leu Gly Phe Ser
Ser Ile 35 40 45 Ile Ala Val Ile Ala Leu Leu Ala Val Gly Leu Thr
Gln Asn Lys Ala 50 55 60 Leu Pro Glu Asn Ile Lys Tyr Gly Ile Val
Leu Asp Ala Gly Ser Ser 65 70 75 80 His Thr Ser Leu Tyr Ile Tyr Lys
Trp Pro Ala Glu Lys Glu Asn Asp 85 90 95 Thr Gly Val Val His Gln
Val Glu Glu Cys Arg Val Lys Gly Pro Gly 100 105 110 Ile Ser Lys Tyr
Val Gln Lys Val Asn Glu Ile Gly Ile Tyr Leu Thr 115 120 125 Asp Cys
Met Glu Arg Ala Arg Glu Val Ile Pro Arg Ser Gln His Gln 130 135 140
Glu Thr Pro Val Tyr Leu Gly Ala Thr Ala Gly Met Arg Leu Leu Arg 145
150 155 160 Met Glu Ser Glu Glu Leu Ala Asp Arg Val Leu Asp Val Val
Glu Arg 165 170 175 Ser Leu Ser Asn Tyr Pro Phe Asp Phe Gln Gly Ala
Arg Ile Ile Thr 180 185 190 Gly Gln Glu Glu Gly Ala Tyr Gly Trp Ile
Thr Ile Asn Tyr Leu Leu 195 200 205 Gly Lys Phe Ser Gln Lys Thr Arg
Trp Phe Ser Ile Val Pro Tyr Glu 210 215 220 Thr Asn Asn Gln Glu Thr
Phe Gly Ala Leu Asp Leu Gly Gly Ala Ser 225 230 235 240 Thr Gln Ile
Thr Phe Val Pro Gln Asn Gln Thr Thr Glu Ser Pro Asp 245 250 255 Asn
Ala Leu Gln Phe Arg Leu Tyr Gly Lys Asp Tyr Asn Val Tyr Thr 260 265
270 His Ser Phe Leu Cys Tyr Gly Lys Asp Gln Ala Leu Trp Gln Lys Leu
275 280 285 Ala Lys Asp Ile Gln Val Ala Ser Asn Glu Ile Leu Arg
Asp Pro Cys 290 295 300 Phe His Pro Gly Tyr Lys Lys Val Val Asn Val
Ser Asp Leu Tyr Lys 305 310 315 320 Thr Pro Cys Thr Lys Arg Phe Glu
Met Thr Leu Pro Phe Gln Gln Phe 325 330 335 Glu Ile Gln Gly Ile Gly
Asn Tyr Gln Gln Cys His Gln Ser Val Leu 340 345 350 Glu Leu Phe Asn
Thr Ser Tyr Cys Pro Tyr Ser Gln Cys Ala Phe Asn 355 360 365 Gly Ile
Phe Leu Pro Pro Leu Gln Gly Asp Phe Gly Ala Phe Ser Ala 370 375 380
Phe Tyr Phe Val Met Asn Phe Leu Asn Leu Thr Ser Glu Lys Val Ser 385
390 395 400 Gln Glu Lys Val Thr Glu Met Met Lys Lys Phe Cys Ser Gln
Pro Trp 405 410 415 Glu Glu Ile Lys Thr Ser Tyr Ala Gly Val Lys Glu
Lys Tyr Leu Ser 420 425 430 Glu Tyr Cys Phe Ser Gly Thr Tyr Ile Leu
Ser Leu Leu Leu Gln Gly 435 440 445 Tyr His Phe Thr Ala Asp Ser Trp
Glu His Ile His Phe Ile Gly Lys 450 455 460 Ile Gln Gly Ser Asp Ala
Gly Trp Thr Leu Gly Tyr Met Leu Asn Leu 465 470 475 480 Thr Asn Met
Ile Pro Ala Glu Gln Pro Leu Ser Thr Pro Leu Ser His 485 490 495 Ser
Thr Tyr Val Phe Leu Met Val Leu Phe Ser Leu Val Leu Val Ile 500 505
510 Val Ala Ile Ile Gly Leu Leu Ile Phe His Lys Pro Ser Tyr Phe Trp
515 520 525 Lys Asp Met Val 530 252473PRTArtificial
SequenceSynthetic hCD39 ECD 252Thr Gln Asn Lys Ala Leu Pro Glu Asn
Val Lys Tyr Gly Ile Val Leu 1 5 10 15 Asp Ala Gly Ser Ser His Thr
Ser Leu Tyr Ile Tyr Lys Trp Pro Ala 20 25 30 Glu Lys Glu Asn Asp
Thr Gly Val Val His Gln Val Glu Glu Cys Arg 35 40 45 Val Lys Gly
Pro Gly Ile Ser Lys Phe Val Gln Lys Val Asn Glu Ile 50 55 60 Gly
Ile Tyr Leu Thr Asp Cys Met Glu Arg Ala Arg Glu Val Ile Pro 65 70
75 80 Arg Ser Gln His Gln Glu Thr Pro Val Tyr Leu Gly Ala Thr Ala
Gly 85 90 95 Met Arg Leu Leu Arg Met Glu Ser Glu Glu Leu Ala Asp
Arg Val Leu 100 105 110 Asp Val Val Glu Arg Ser Leu Ser Asn Tyr Pro
Phe Asp Phe Gln Gly 115 120 125 Ala Arg Ile Ile Thr Gly Gln Glu Glu
Gly Ala Tyr Gly Trp Ile Thr 130 135 140 Ile Asn Tyr Leu Leu Gly Lys
Phe Ser Gln Lys Thr Arg Trp Phe Ser 145 150 155 160 Ile Val Pro Tyr
Glu Thr Asn Asn Gln Glu Thr Phe Gly Ala Leu Asp 165 170 175 Leu Gly
Gly Ala Ser Thr Gln Val Thr Phe Val Pro Gln Asn Gln Thr 180 185 190
Ile Glu Ser Pro Asp Asn Ala Leu Gln Phe Arg Leu Tyr Gly Lys Asp 195
200 205 Tyr Asn Val Tyr Thr His Ser Phe Leu Cys Tyr Gly Lys Asp Gln
Ala 210 215 220 Leu Trp Gln Lys Leu Ala Lys Asp Ile Gln Val Ala Ser
Asn Glu Ile 225 230 235 240 Leu Arg Asp Pro Cys Phe His Pro Gly Tyr
Lys Lys Val Val Asn Val 245 250 255 Ser Asp Leu Tyr Lys Thr Pro Cys
Thr Lys Arg Phe Glu Met Thr Leu 260 265 270 Pro Phe Gln Gln Phe Glu
Ile Gln Gly Ile Gly Asn Tyr Gln Gln Cys 275 280 285 His Gln Ser Ile
Leu Glu Leu Phe Asn Thr Ser Tyr Cys Pro Tyr Ser 290 295 300 Gln Cys
Ala Phe Asn Gly Ile Phe Leu Pro Pro Leu Gln Gly Asp Phe 305 310 315
320 Gly Ala Phe Ser Ala Phe Tyr Phe Val Met Lys Phe Leu Asn Leu Thr
325 330 335 Ser Glu Lys Val Ser Gln Glu Lys Val Thr Glu Met Met Lys
Lys Phe 340 345 350 Cys Ala Gln Pro Trp Glu Glu Ile Lys Thr Ser Tyr
Ala Gly Val Lys 355 360 365 Glu Lys Tyr Leu Ser Glu Tyr Cys Phe Ser
Gly Thr Tyr Ile Leu Ser 370 375 380 Leu Leu Leu Gln Gly Tyr His Phe
Thr Ala Asp Ser Trp Glu His Ile 385 390 395 400 His Phe Ile Gly Lys
Ile Gln Gly Ser Asp Ala Gly Trp Thr Leu Gly 405 410 415 Tyr Met Leu
Asn Leu Thr Asn Met Ile Pro Ala Glu Gln Pro Leu Ser 420 425 430 Thr
Pro Leu Ser His Ser Thr Tyr Val Phe Leu Met Val Leu Phe Ser 435 440
445 Leu Val Leu Phe Thr Val Ala Ile Ile Gly Leu Leu Ile Phe His Lys
450 455 460 Pro Ser Tyr Phe Trp Lys Asp Met Val 465 470
253473PRTArtificial SequenceSynthetic mCD39 ECD 253Thr Gln Asn Lys
Pro Leu Pro Glu Asn Val Lys Tyr Gly Ile Val Leu 1 5 10 15 Asp Ala
Gly Ser Ser His Thr Asn Leu Tyr Ile Tyr Lys Trp Pro Ala 20 25 30
Glu Lys Glu Asn Asp Thr Gly Val Val Gln Gln Leu Glu Glu Cys Gln 35
40 45 Val Lys Gly Pro Gly Ile Ser Lys Tyr Ala Gln Lys Thr Asp Glu
Ile 50 55 60 Gly Ala Tyr Leu Ala Glu Cys Met Glu Leu Ser Thr Glu
Leu Ile Pro 65 70 75 80 Thr Ser Lys His His Gln Thr Pro Val Tyr Leu
Gly Ala Thr Ala Gly 85 90 95 Met Arg Leu Leu Arg Met Glu Ser Glu
Gln Ser Ala Asp Glu Val Leu 100 105 110 Ala Ala Val Ser Thr Ser Leu
Lys Ser Tyr Pro Phe Asp Phe Gln Gly 115 120 125 Ala Lys Ile Ile Thr
Gly Gln Glu Glu Gly Ala Tyr Gly Trp Ile Thr 130 135 140 Ile Asn Tyr
Leu Leu Gly Arg Phe Thr Gln Glu Gln Ser Trp Leu Ser 145 150 155 160
Leu Ile Ser Asp Ser Gln Lys Gln Glu Thr Phe Gly Ala Leu Asp Leu 165
170 175 Gly Gly Ala Ser Thr Gln Ile Thr Phe Val Pro Gln Asn Ser Thr
Ile 180 185 190 Glu Ser Pro Glu Asn Ser Leu Gln Phe Arg Leu Tyr Gly
Glu Asp Tyr 195 200 205 Thr Val Tyr Thr His Ser Phe Leu Cys Tyr Gly
Lys Asp Gln Ala Leu 210 215 220 Trp Gln Lys Leu Ala Lys Asp Ile Gln
Val Ser Ser Gly Gly Val Leu 225 230 235 240 Lys Asp Pro Cys Phe Asn
Pro Gly Tyr Glu Lys Val Val Asn Val Ser 245 250 255 Glu Leu Tyr Gly
Thr Pro Cys Thr Lys Arg Phe Glu Lys Lys Leu Pro 260 265 270 Phe Asp
Gln Phe Arg Ile Gln Gly Thr Gly Asp Tyr Glu Gln Cys His 275 280 285
Gln Ser Ile Leu Glu Leu Phe Asn Asn Ser His Cys Pro Tyr Ser Gln 290
295 300 Cys Ala Phe Asn Gly Val Phe Leu Pro Pro Leu His Gly Ser Phe
Gly 305 310 315 320 Ala Phe Ser Ala Phe Tyr Phe Val Met Asp Phe Phe
Lys Lys Val Ala 325 330 335 Lys Asn Ser Val Ile Ser Gln Glu Lys Met
Thr Glu Ile Thr Lys Asn 340 345 350 Phe Cys Ser Lys Ser Trp Glu Glu
Thr Lys Thr Ser Tyr Pro Ser Val 355 360 365 Lys Glu Lys Tyr Leu Ser
Glu Tyr Cys Phe Ser Gly Ala Tyr Ile Leu 370 375 380 Ser Leu Leu Gln
Gly Tyr Asn Phe Thr Asp Ser Ser Trp Glu Gln Ile 385 390 395 400 His
Phe Met Gly Lys Ile Lys Asp Ser Asn Ala Gly Trp Thr Leu Gly 405 410
415 Tyr Met Leu Asn Leu Thr Asn Met Ile Pro Ala Glu Gln Pro Leu Ser
420 425 430 Pro Pro Leu Pro His Ser Thr Tyr Ile Gly Leu Met Val Leu
Phe Ser 435 440 445 Leu Leu Leu Val Ala Val Ala Ile Thr Gly Leu Phe
Ile Tyr Ser Lys 450 455 460 Pro Ser Tyr Phe Trp Lys Glu Ala Val 465
470 254473PRTArtificial SequenceSynthetic cCD39 ECD 254Thr Gln Asn
Lys Ala Leu Pro Glu Asn Ile Lys Tyr Gly Ile Val Leu 1 5 10 15 Asp
Ala Gly Ser Ser His Thr Ser Leu Tyr Ile Tyr Lys Trp Pro Ala 20 25
30 Glu Lys Glu Asn Asp Thr Gly Val Val His Gln Val Glu Glu Cys Arg
35 40 45 Val Lys Gly Pro Gly Ile Ser Lys Tyr Val Gln Lys Val Asn
Glu Ile 50 55 60 Gly Ile Tyr Leu Thr Asp Cys Met Glu Arg Ala Arg
Glu Val Ile Pro 65 70 75 80 Arg Ser Gln His Gln Glu Thr Pro Val Tyr
Leu Gly Ala Thr Ala Gly 85 90 95 Met Arg Leu Leu Arg Met Glu Ser
Glu Glu Leu Ala Asp Arg Val Leu 100 105 110 Asp Val Val Glu Arg Ser
Leu Ser Asn Tyr Pro Phe Asp Phe Gln Gly 115 120 125 Ala Arg Ile Ile
Thr Gly Gln Glu Glu Gly Ala Tyr Gly Trp Ile Thr 130 135 140 Ile Asn
Tyr Leu Leu Gly Lys Phe Ser Gln Lys Thr Arg Trp Phe Ser 145 150 155
160 Ile Val Pro Tyr Glu Thr Asn Asn Gln Glu Thr Phe Gly Ala Leu Asp
165 170 175 Leu Gly Gly Ala Ser Thr Gln Ile Thr Phe Val Pro Gln Asn
Gln Thr 180 185 190 Thr Glu Ser Pro Asp Asn Ala Leu Gln Phe Arg Leu
Tyr Gly Lys Asp 195 200 205 Tyr Asn Val Tyr Thr His Ser Phe Leu Cys
Tyr Gly Lys Asp Gln Ala 210 215 220 Leu Trp Gln Lys Leu Ala Lys Asp
Ile Gln Val Ala Ser Asn Glu Ile 225 230 235 240 Leu Arg Asp Pro Cys
Phe His Pro Gly Tyr Lys Lys Val Val Asn Val 245 250 255 Ser Asp Leu
Tyr Lys Thr Pro Cys Thr Lys Arg Phe Glu Met Thr Leu 260 265 270 Pro
Phe Gln Gln Phe Glu Ile Gln Gly Ile Gly Asn Tyr Gln Gln Cys 275 280
285 His Gln Ser Val Leu Glu Leu Phe Asn Thr Ser Tyr Cys Pro Tyr Ser
290 295 300 Gln Cys Ala Phe Asn Gly Ile Phe Leu Pro Pro Leu Gln Gly
Asp Phe 305 310 315 320 Gly Ala Phe Ser Ala Phe Tyr Phe Val Met Asn
Phe Leu Asn Leu Thr 325 330 335 Ser Glu Lys Val Ser Gln Glu Lys Val
Thr Glu Met Met Lys Lys Phe 340 345 350 Cys Ser Gln Pro Trp Glu Glu
Ile Lys Thr Ser Tyr Ala Gly Val Lys 355 360 365 Glu Lys Tyr Leu Ser
Glu Tyr Cys Phe Ser Gly Thr Tyr Ile Leu Ser 370 375 380 Leu Leu Leu
Gln Gly Tyr His Phe Thr Ala Asp Ser Trp Glu His Ile 385 390 395 400
His Phe Ile Gly Lys Ile Gln Gly Ser Asp Ala Gly Trp Thr Leu Gly 405
410 415 Tyr Met Leu Asn Leu Thr Asn Met Ile Pro Ala Glu Gln Pro Leu
Ser 420 425 430 Thr Pro Leu Ser His Ser Thr Tyr Val Phe Leu Met Val
Leu Phe Ser 435 440 445 Leu Val Leu Val Ile Val Ala Ile Ile Gly Leu
Leu Ile Phe His Lys 450 455 460 Pro Ser Tyr Phe Trp Lys Asp Met Val
465 470 255448PRTArtificial SequenceSynthetic HC, 31895 255Gln Val
Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15
Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Lys Ser Tyr 20
25 30 Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp
Met 35 40 45 Gly Arg Ile Asn Pro Ser Val Gly Ser Thr Trp Tyr Ala
Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser
Thr Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu
Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly
Gly Thr Glu Tyr Leu Arg Lys Trp Gly 100 105 110 Gln Gly Thr Leu Val
Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro
Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala
Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150
155 160 Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro
Ala 165 170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val
Val Thr Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr
Cys Asn Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys
Arg Val Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro
Ala Pro Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270
Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275
280 285 Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val
Val 290 295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly
Lys Glu Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro
Ser Ser Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu
Met Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly
Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395
400 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser
405 410 415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His
Glu Ala 420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu
Ser Leu Gly Lys 435 440 445 256214PRTArtificial SequenceSynthetic
LC, 31895 256Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu
Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln
Ser Val Ala Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Gly Ala Ser Asn Arg
His Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly Ser Gly Ser Gly
Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70 75 80 Pro Glu Asp
Phe Ala Val Tyr Tyr Cys Gln Gln Tyr His Asn Ala Ile 85 90 95 Thr
Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105
110 Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly
115 120 125 Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg
Glu Ala 130 135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser
Gly Asn Ser Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys
Asp Ser Thr Tyr Ser Leu Ser
165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys 210
257448PRTArtificial SequenceSynthetic HC, 31415 257Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25 30
Gln Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Arg Ile Asn Pro Ser Gly Gly Ser Thr Trp Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser
Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr
Glu Tyr Leu Arg Ser Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160
Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165
170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn
Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val
Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro
Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val
Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285
Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290
295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser
Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410
415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala
420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu
Gly Lys 435 440 445 258214PRTArtificial SequenceSynthetic LC, 31415
258Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly
1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser
Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala
Pro Arg Leu Leu 35 40 45 Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly
Ile Pro Asp Arg Phe Ser 50 55 60 Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Ser Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe Ala Val
Tyr Tyr Cys Gln Gln Tyr His Ser Tyr Ile 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130
135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser
Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser Pro Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys
210 259448PRTArtificial SequenceSynthetic HC, 31891 259Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Ile Phe Lys Ser Tyr 20 25
30 Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Arg Ile Asn Pro Ser Val Gly Ser Thr Trp Tyr Ala Gln
Lys Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr
Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly
Thr Glu Tyr Leu Arg Val Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr
Val Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu
Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu
Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155
160 Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala
165 170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val
Thr Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys
Asn Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg
Val Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala
Pro Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu
Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu
Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280
285 Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val
290 295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys
Glu Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met
Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400
Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405
410 415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu
Ala 420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
Leu Gly Lys 435 440 445 260214PRTArtificial SequenceSynthetic LC,
31891 260Glu Ile Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser
Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser
Val Ser Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly
Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Tyr Ala Ser Ser Arg Ala
Tyr Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly Ser Gly Ser Gly Thr
Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe
Ala Val Tyr Tyr Cys Gln Gln Tyr His Asn Ala Ile 85 90 95 Thr Phe
Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110
Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115
120 125 Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu
Ala 130 135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly
Asn Ser Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp
Ser Thr Tyr Ser Leu Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala
Asp Tyr Glu Lys His Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His
Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly
Glu Cys 210 261448PRTArtificial SequenceSynthetic HC, 31418 261Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10
15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Lys Ser Tyr
20 25 30 Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45 Gly Arg Ile Asn Pro Ser Gly Gly Ser Thr Trp Tyr
Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr
Ser Thr Ser Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu
Gly Gly Thr Glu Tyr Leu Arg His Trp Gly 100 105 110 Gln Gly Thr Leu
Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe
Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140
Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145
150 155 160 Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe
Pro Ala 165 170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser
Val Val Thr Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr
Thr Cys Asn Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp
Lys Arg Val Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys
Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu
Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr
Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265
270 Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
275 280 285 Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg
Val Val 290 295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn
Gly Lys Glu Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu
Pro Ser Ser Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu
Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn
Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390
395 400 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys
Ser 405 410 415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala 420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Leu Gly Lys 435 440 445 262214PRTArtificial
SequenceSynthetic LC, 31418 262Glu Ile Val Leu Thr Gln Ser Pro Gly
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Ser Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Gly
Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70
75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr Tyr Phe Tyr
Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr
Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu
Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val Val Cys Leu Leu Asn
Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155 160 Glu Ser Val Thr
Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175 Ser Thr
Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190
Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195
200 205 Phe Asn Arg Gly Glu Cys 210 263448PRTArtificial
SequenceSynthetic HC, 31430 263Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Lys Ser Tyr 20 25 30 Glu Met His Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Arg Ile
Asn Pro Ser Gly Gly Ser Thr Trp Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr Glu Tyr Leu Arg
His Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser
Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala Pro Cys Ser Arg
Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160 Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170 175 Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val 180 185 190
Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val Asp His 195
200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys Tyr
Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Leu
Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro Lys Pro Lys
Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val Thr Cys Val
Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val Gln Phe Asn
Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285 Lys Thr Lys
Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290 295 300 Ser
Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 305 310
315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys
Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val
Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln
Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser Asp Gly Ser
Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410 415 Arg Trp
Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala 420 425 430
Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys 435
440 445 264214PRTArtificial SequenceSynthetic LC, 31430 264Glu Ile
Val Leu Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly 1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Glu Ala Ser Gln Ser Val Ser Tyr Ser 20
25 30 Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu
Leu 35 40 45 Ile Tyr Gly Ala Ser Ser Arg Ala Asn Gly Ile Pro Asp
Arg Phe Ser 50 55 60 Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr
Ile Ser Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys
Gln Gln Tyr His Ser Ala Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys
Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser Val Phe Ile
Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser
Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys
Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150
155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu
Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His
Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser
Pro Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys 210
265447PRTArtificial SequenceSynthetic HC, 31915 265Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Leu 20 25 30
Pro Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Gly Ile Gly Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe Gln
Gly 50 55 60 Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala
Tyr Met Glu 65 70 75 80 Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val
Tyr Tyr Cys Ala Arg 85 90 95 Gly Gly Ala Lys Tyr Ala Ser Lys Trp
Gly Met Asp Val Trp Gly Gln 100 105 110 Gly Thr Thr Val Thr Val Ser
Ser Ala Ser Thr Lys Gly Pro Ser Val 115 120 125 Phe Pro Leu Ala Pro
Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala 130 135 140 Leu Gly Cys
Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser 145 150 155 160
Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val 165
170 175 Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro 180 185 190 Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val
Asp His Lys 195 200 205 Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu
Ser Lys Tyr Gly Pro 210 215 220 Pro Cys Pro Pro Cys Pro Ala Pro Glu
Phe Leu Gly Gly Pro Ser Val 225 230 235 240 Phe Leu Phe Pro Pro Lys
Pro Lys Asp Thr Leu Met Ile Ser Arg Thr 245 250 255 Pro Glu Val Thr
Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu 260 265 270 Val Gln
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys 275 280 285
Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser 290
295 300 Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr
Lys 305 310 315 320 Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile
Glu Lys Thr Ile 325 330 335 Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro
Gln Val Tyr Thr Leu Pro 340 345 350 Pro Ser Gln Glu Glu Met Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu 355 360 365 Val Lys Gly Phe Tyr Pro
Ser Asp Ile Ala Val Glu Trp Glu Ser Asn 370 375 380 Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser 385 390 395 400 Asp
Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg 405 410
415 Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu
420 425 430 His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly
Lys 435 440 445 266220PRTArtificial SequenceSynthetic LC, 31915
266Asp Ile Val Met Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly
1 5 10 15 Glu Arg Ala Thr Ile Asn Cys Lys Ser Ser Gln Ser Val Leu
Phe Ser 20 25 30 Ser Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln
Lys Pro Gly Gln 35 40 45 Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser
Ser Arg Glu Ser Gly Val 50 55 60 Pro Asp Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr 65 70 75 80 Ile Ser Ser Leu Gln Ala
Glu Asp Val Ala Val Tyr Tyr Cys Gln Gln 85 90 95 Tyr Trp Thr Tyr
Pro Leu Thr Phe Gly Gly Gly Thr Lys Val Glu Ile 100 105 110 Lys Arg
Thr Val Ala Ala Pro Ser Val Phe Ile Phe Pro Pro Ser Asp 115 120 125
Glu Gln Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn 130
135 140 Phe Tyr Pro Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala
Leu 145 150 155 160 Gln Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln
Asp Ser Lys Asp 165 170 175 Ser Thr Tyr Ser Leu Ser Ser Thr Leu Thr
Leu Ser Lys Ala Asp Tyr 180 185 190 Glu Lys His Lys Val Tyr Ala Cys
Glu Val Thr His Gln Gly Leu Ser 195 200 205 Ser Pro Val Thr Lys Ser
Phe Asn Arg Gly Glu Cys 210 215 220 267447PRTArtificial
SequenceSynthetic HC, 31905 267Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Pro Ser Asn 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Gly Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe Gln Gly 50 55 60 Arg
Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr Met Glu 65 70
75 80 Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys Ala
Arg 85 90 95 Gly Gly Ala Lys Tyr Ala Arg Thr Tyr Gly Met Asp Val
Trp Gly Gln 100 105 110 Gly Thr Thr Val Thr Val Ser Ser Ala Ser Thr
Lys Gly Pro Ser Val 115 120 125 Phe Pro Leu Ala Pro Cys Ser Arg Ser
Thr Ser Glu Ser Thr Ala Ala 130 135 140 Leu Gly Cys Leu Val Lys Asp
Tyr Phe Pro Glu Pro Val Thr Val Ser 145 150 155 160 Trp Asn Ser Gly
Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val 165 170 175 Leu Gln
Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro 180 185 190
Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val Asp His Lys 195
200 205 Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys Tyr Gly
Pro 210 215 220 Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Leu Gly Gly
Pro Ser Val 225 230 235 240 Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr
Leu Met Ile Ser Arg Thr 245 250 255 Pro Glu Val Thr Cys Val Val Val
Asp Val Ser Gln Glu Asp Pro Glu 260 265 270 Val Gln Phe Asn Trp Tyr
Val Asp Gly Val Glu Val His Asn Ala Lys 275 280 285 Thr Lys Pro Arg
Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser 290 295 300 Val Leu
Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys 305 310 315
320 Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile
325 330 335 Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu Pro 340 345 350 Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu 355 360 365 Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala
Val Glu Trp Glu Ser Asn 370 375 380 Gly Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser 385 390 395 400 Asp Gly Ser Phe Phe
Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg 405 410 415 Trp Gln Glu
Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu 420 425 430 His
Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys 435 440 445
268220PRTArtificial SequenceSynthetic LC, 31905 268Asp Ile Val Met
Thr Gln Ser Pro Asp Ser Leu Ala Val Ser Leu Gly 1 5 10 15 Glu Arg
Ala Thr Ile Asn Cys Lys Ser Ser Lys Ser Val Leu Tyr Ser 20 25 30
Asn Asn Asn Lys Asn Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35
40 45 Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Gln Ser Gly
Val 50 55 60 Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr 65 70 75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val
Tyr Tyr Cys Gln Gln 85 90 95 Tyr Leu Leu Tyr Pro Leu Thr Phe Gly
Gly Gly Thr Lys Val Glu Ile 100 105 110 Lys Arg Thr Val Ala Ala Pro
Ser Val Phe Ile Phe Pro Pro Ser Asp 115 120 125 Glu Gln Leu Lys Ser
Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn 130 135 140 Phe Tyr Pro
Arg Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu 145 150 155 160
Gln Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp 165
170 175 Ser Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp
Tyr 180 185 190 Glu Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln
Gly Leu Ser 195 200 205 Ser Pro Val Thr Lys Ser Phe Asn Arg Gly Glu
Cys 210 215 220 269447PRTArtificial SequenceSynthetic HC, 31901
269Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser
1 5 10 15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser
Ser Leu 20 25 30 Pro Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly
Leu Glu Trp Met 35 40 45 Gly Gly Ile Gly Phe Gly Thr Ala Asn Tyr
Ala Gln Lys Phe Gln Gly 50 55 60 Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr Ser Thr Ala Tyr Met Glu 65 70 75 80 Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys Ala Arg 85 90 95 Gly Gly Ala Lys
Tyr Ala Gly Arg Tyr Gly Met Asp Val Trp Gly Gln 100 105 110 Gly Thr
Thr Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val 115 120 125
Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala 130
135 140 Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val
Ser 145 150 155 160 Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr
Phe Pro Ala Val 165 170 175 Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser
Ser Val Val Thr Val Pro 180 185 190 Ser Ser Ser Leu Gly Thr Lys Thr
Tyr Thr Cys Asn Val Asp His Lys 195 200 205 Pro Ser Asn Thr Lys Val
Asp Lys Arg Val Glu Ser Lys Tyr Gly Pro 210 215 220 Pro Cys Pro Pro
Cys Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser Val 225 230 235 240 Phe
Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr 245 250
255 Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu
260 265 270 Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn
Ala Lys 275 280 285 Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr
Arg Val Val Ser 290 295 300 Val Leu Thr Val Leu His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr Lys 305 310 315 320 Cys Lys Val Ser Asn Lys Gly
Leu Pro Ser Ser Ile Glu Lys Thr Ile 325 330 335 Ser Lys Ala Lys Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro 340 345 350 Pro Ser Gln
Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu 355 360 365 Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn 370 375
380 Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser
385 390 395 400 Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp
Lys Ser Arg 405 410 415 Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu 420 425 430 His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Leu Gly Lys 435 440 445 270220PRTArtificial
SequenceSynthetic LC, 31901 270Gly Ile Val Met Thr Gln Ser Pro Asp
Ser Leu Ala Val Ser Leu Gly 1 5 10 15 Glu Arg Ala Thr Ile Asn Cys
Lys Ser Ser Gln Ser Val Leu Phe Ser 20 25 30 Ser Asn Asn Lys Asn
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln 35 40
45 Pro Pro Lys Leu Leu Ile Tyr Trp Ala Ser Thr Arg Ala Ser Gly Val
50 55 60 Pro Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr
Leu Thr 65 70 75 80 Ile Ser Ser Leu Gln Ala Glu Asp Val Ala Val Tyr
Tyr Cys Gln Gln 85 90 95 Tyr Tyr Leu Tyr Pro Leu Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile 100 105 110 Lys Arg Thr Val Ala Ala Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp 115 120 125 Glu Gln Leu Lys Ser Gly
Thr Ala Ser Val Val Cys Leu Leu Asn Asn 130 135 140 Phe Tyr Pro Arg
Glu Ala Lys Val Gln Trp Lys Val Asp Asn Ala Leu 145 150 155 160 Gln
Ser Gly Asn Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp 165 170
175 Ser Thr Tyr Ser Leu Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
180 185 190 Glu Lys His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly
Leu Ser 195 200 205 Ser Pro Val Thr Lys Ser Phe Asn Arg Gly Glu Cys
210 215 220 271448PRTArtificial SequenceSynthetic HC, 31861 271Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10
15 Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Gly Ser Tyr
20 25 30 Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45 Gly Ser Ile Ile Pro Glu Phe Gly Ile Ala Asn Tyr
Ala Gln Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu
Ser Thr Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser
Glu Asp Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Glu Ser Gly Thr
Tyr Arg Asp His Arg Leu Asp Val Trp Gly 100 105 110 Gln Gly Thr Met
Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe
Pro Leu Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140
Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145
150 155 160 Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe
Pro Ala 165 170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser
Val Val Thr Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr
Thr Cys Asn Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp
Lys Arg Val Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys
Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu
Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr
Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265
270 Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
275 280 285 Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg
Val Val 290 295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn
Gly Lys Glu Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu
Pro Ser Ser Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu
Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn
Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390
395 400 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys
Ser 405 410 415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala 420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Leu Gly Lys 435 440 445 272214PRTArtificial
SequenceSynthetic LC, 31861 272Glu Ile Val Met Thr Gln Ser Pro Ala
Thr Leu Ser Val Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Gly Ser Asn 20 25 30 Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Gly Ala
Ser Thr Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser
Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Ser 65 70
75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr Leu Leu Trp Pro
Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr
Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu
Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val Val Cys Leu Leu Asn
Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155 160 Glu Ser Val Thr
Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175 Ser Thr
Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190
Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195
200 205 Phe Asn Arg Gly Glu Cys 210 273448PRTArtificial
SequenceSynthetic HC, 31873 273Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Lys Tyr 20 25 30 Gly Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Ser Ile
Ile Pro Glu Phe Gly Ile Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Glu Ser Gly Gly Tyr Arg Asp His Arg Leu Gly
Val Trp Gly 100 105 110 Gln Gly Thr Met Val Thr Val Ser Ser Ala Ser
Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala Pro Cys Ser Arg
Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160 Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170 175 Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val 180 185 190
Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val Asp His 195
200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys Tyr
Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Leu Gly
Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val Gln Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285 Lys Thr Lys Pro
Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290 295 300 Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 305 310 315
320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr
325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410 415 Arg Trp Gln
Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala 420 425 430 Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys 435 440
445 274214PRTArtificial SequenceSynthetic LC, 31873 274Glu Ile Val
Met Thr Gln Ser Pro Ala Thr Leu Ser Val Ser Pro Gly 1 5 10 15 Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Gly Ser Asn 20 25
30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45 Tyr Gly Ala Ser Thr Arg Ala Ser Gly Ile Pro Ala Arg Phe
Ser Gly 50 55 60 Ser Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser
Ser Leu Gln Ser 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Tyr Leu Leu Trp Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val
Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe
Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val
Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val
Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155
160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys 210
275448PRTArtificial SequenceSynthetic HC, 31393 275Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Glu Ser Tyr 20 25 30
Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Ser Ile Ile Pro Glu Phe Gly Ile Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Thr Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Glu Ser Gly Gly Tyr Arg Asp
His Arg Leu Asp Val Trp Gly 100 105 110 Gln Gly Thr Met Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160
Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165
170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn
Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val
Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro
Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val
Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285
Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290
295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser
Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410
415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala
420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu
Gly Lys 435 440 445 276214PRTArtificial SequenceSynthetic LC, 31393
276Glu Ile Val Met Thr Gln Ser Pro Ala Thr Leu Ser Val Ser Pro Gly
1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Gly
Ser Asn 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro
Arg Leu Leu Ile 35 40 45 Tyr Gly Ala Ser Thr Arg Ala Thr Gly Ile
Pro Ala Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Glu Phe Thr
Leu Thr Ile Ser Ser Leu Gln Ser 65 70 75 80 Glu Asp Phe Ala Val Tyr
Tyr Cys Gln Gln Tyr Leu Leu Trp Pro Leu 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130
135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser
Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser Pro Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys
210 277453PRTArtificial SequenceSynthetic HC, 27597 277Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25
30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Ser Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln
Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr
Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Glu Gly Arg Gly Tyr Asp
Ser Ser
Arg Tyr Tyr Lys Phe Trp 100 105 110 Phe Asp Pro Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Ser Ala Ser 115 120 125 Thr Lys Gly Pro Ser Val
Phe Pro Leu Ala Pro Cys Ser Arg Ser Thr 130 135 140 Ser Glu Ser Thr
Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr Phe Pro 145 150 155 160 Glu
Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val 165 170
175 His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser
180 185 190 Ser Val Val Thr Val Pro Ser Ser Ser Leu Gly Thr Lys Thr
Tyr Thr 195 200 205 Cys Asn Val Asp His Lys Pro Ser Asn Thr Lys Val
Asp Lys Arg Val 210 215 220 Glu Ser Lys Tyr Gly Pro Pro Cys Pro Pro
Cys Pro Ala Pro Glu Phe 225 230 235 240 Leu Gly Gly Pro Ser Val Phe
Leu Phe Pro Pro Lys Pro Lys Asp Thr 245 250 255 Leu Met Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val Asp Val 260 265 270 Ser Gln Glu
Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val 275 280 285 Glu
Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser 290 295
300 Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu
305 310 315 320 Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly
Leu Pro Ser 325 330 335 Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly
Gln Pro Arg Glu Pro 340 345 350 Gln Val Tyr Thr Leu Pro Pro Ser Gln
Glu Glu Met Thr Lys Asn Gln 355 360 365 Val Ser Leu Thr Cys Leu Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala 370 375 380 Val Glu Trp Glu Ser
Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr 385 390 395 400 Pro Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu 405 410 415
Thr Val Asp Lys Ser Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser 420
425 430 Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu
Ser 435 440 445 Leu Ser Leu Gly Lys 450 278214PRTArtificial
SequenceSynthetic LC, 27597 278Glu Ile Val Leu Thr Gln Ser Pro Ala
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Ser Ser Tyr 20 25 30 Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Asp Ala
Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro 65 70
75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Val Leu Trp Pro
Arg 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr
Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu
Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val Val Cys Leu Leu Asn
Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155 160 Glu Ser Val Thr
Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175 Ser Thr
Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190
Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195
200 205 Phe Asn Arg Gly Glu Cys 210 279448PRTArtificial
SequenceSynthetic HC, 27575 279Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Glu Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30 Ala Thr Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Ile Pro Ile Ser Gly Thr Ala Asn Tyr Ala Gln Glu Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Asp Gly Gly Gly Tyr Arg His His Tyr Phe Asp
Leu Trp Gly 100 105 110 Arg Gly Thr Leu Val Thr Val Ser Ser Ala Ser
Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala Pro Cys Ser Arg
Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160 Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170 175 Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val 180 185 190
Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val Asp His 195
200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys Tyr
Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Leu Gly
Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val Gln Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285 Lys Thr Lys Pro
Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290 295 300 Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 305 310 315
320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr
325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410 415 Arg Trp Gln
Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala 420 425 430 Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys 435 440
445 280214PRTArtificial SequenceSynthetic LC, 27575 280Glu Ile Val
Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Arg Tyr 20 25
30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45 Tyr Asp Ala Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe
Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln
His Val Asn Phe Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val
Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe
Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val
Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val
Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155
160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys 210
281448PRTArtificial SequenceSynthetic HC, 27568 281Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Pro Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Glu Gln Gly Leu Glu Trp Met 35
40 45 Gly Ser Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Gly Glu Ser Gly Gly Tyr Arg Asp
His Lys Leu Asp Val Trp Gly 100 105 110 Gln Gly Thr Val Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160
Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165
170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn
Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val
Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro
Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val
Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285
Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290
295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser
Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410
415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala
420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu
Gly Lys 435 440 445 282214PRTArtificial SequenceSynthetic LC, 27568
282Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser
Ser Tyr 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro
Arg Leu Leu Ile 35 40 45 Tyr Asp Ala Ser Asn Arg Ala Thr Gly Ile
Pro Ala Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr
Leu Thr Ile Ser Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala Val Tyr
Tyr Cys Gln Gln Ser Val Phe Trp Pro Ile 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130
135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser
Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser Pro Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys
210 283448PRTArtificial SequenceSynthetic HC, 27577 283Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Ala Phe Ser Ser Tyr 20 25
30 Ala Ile Gly Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Gly Ile Ile Pro Thr Phe Gly Thr Ala Asn Tyr Ala Gln
Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr
Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Gly Gly Gly Tyr Gln
His His Tyr Phe Asp Leu Trp Gly 100 105 110 Arg Gly Thr Leu Val Thr
Val Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu
Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu
Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155
160 Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala
165 170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val
Thr Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys
Asn Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg
Val Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala
Pro Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu
Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu
Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280
285 Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val
290 295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys
Glu Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met
Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370
375 380 Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu
Asp 385 390 395 400 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr
Val Asp Lys Ser 405 410 415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys
Ser Val Met His Glu Ala 420 425 430 Leu His Asn His Tyr Thr Gln Lys
Ser Leu Ser Leu Ser Leu Gly Lys 435 440 445 284214PRTArtificial
SequenceSynthetic LC, 27577 284Glu Ile Val Met Thr Gln Ser Pro Ala
Thr Leu Ser Val Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Gly Ser Asn 20 25 30 Leu Ala Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Gly Ala
Ser Thr Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser
Gly Ser Gly Thr Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Ser 65 70
75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Leu Thr Lys Trp Pro
Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr
Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu
Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val Val Cys Leu Leu Asn
Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155 160 Glu Ser Val Thr
Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175 Ser Thr
Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190
Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195
200 205 Phe Asn Arg Gly Glu Cys 210 285448PRTArtificial
SequenceSynthetic HC, 27587 285Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Ser Ile
Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Glu Ser Gly Gly Tyr Arg Asp His Lys Leu Asp
Val Trp Gly 100 105 110 Gln Gly Thr Met Val Thr Val Ser Ser Ala Ser
Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala Pro Cys Ser Arg
Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160 Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170 175 Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val 180 185 190
Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val Asp His 195
200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys Tyr
Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Leu Gly
Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val Gln Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285 Lys Thr Lys Pro
Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290 295 300 Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 305 310 315
320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr
325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410 415 Arg Trp Gln
Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala 420 425 430 Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys 435 440
445 286214PRTArtificial SequenceSynthetic LC, 27587 286Glu Ile Val
Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Tyr 20 25
30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45 Tyr Asp Ala Ser Lys Arg Ala Thr Gly Ile Pro Ala Arg Phe
Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Asp Val Leu Trp Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val
Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe
Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val
Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val
Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155
160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys 210
287447PRTArtificial SequenceSynthetic HC, 27589 287Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser
Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Ser Gly Tyr His Arg His
Tyr Ser Asp Tyr Trp Gly Gln 100 105 110 Gly Thr Leu Val Thr Val Ser
Ser Ala Ser Thr Lys Gly Pro Ser Val 115 120 125 Phe Pro Leu Ala Pro
Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala Ala 130 135 140 Leu Gly Cys
Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser 145 150 155 160
Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val 165
170 175 Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val
Pro 180 185 190 Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val
Asp His Lys 195 200 205 Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu
Ser Lys Tyr Gly Pro 210 215 220 Pro Cys Pro Pro Cys Pro Ala Pro Glu
Phe Leu Gly Gly Pro Ser Val 225 230 235 240 Phe Leu Phe Pro Pro Lys
Pro Lys Asp Thr Leu Met Ile Ser Arg Thr 245 250 255 Pro Glu Val Thr
Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu 260 265 270 Val Gln
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys 275 280 285
Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser 290
295 300 Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr
Lys 305 310 315 320 Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile
Glu Lys Thr Ile 325 330 335 Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro
Gln Val Tyr Thr Leu Pro 340 345 350 Pro Ser Gln Glu Glu Met Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu 355 360 365 Val Lys Gly Phe Tyr Pro
Ser Asp Ile Ala Val Glu Trp Glu Ser Asn 370 375 380 Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser 385 390 395 400 Asp
Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg 405 410
415 Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu
420 425 430 His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly
Lys 435 440 445 288214PRTArtificial SequenceSynthetic LC, 27589
288Asp Ile Gln Met Thr Gln Ser Pro Ser Thr Leu Ser Ala Ser Val Gly
1 5 10 15 Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser
Ser Trp 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro
Lys Leu Leu Ile 35 40 45 Tyr Lys Ala Ser Ser Leu Glu Ser Gly Val
Pro Ser Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Glu Phe Thr
Leu Thr Ile Ser Ser Leu Gln Pro 65 70 75 80 Asp Asp Phe Ala Thr Tyr
Tyr Cys Gln Gln Tyr Gly Leu Phe Pro Ile 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130
135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser
Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser Pro Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys
210 289448PRTArtificial SequenceSynthetic HC, 27596 289Gln Val Gln
Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser
Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25
30 Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45 Gly Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln
Lys Phe 50 55 60 Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr
Ser Thr Ala Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Pro Leu Gly Ile Arg
Lys His Trp Phe Asp Pro Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr
Val Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu
Ala Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu
Gly Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155
160 Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala
165 170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val
Thr Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys
Asn Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg
Val Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala
Pro Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu
Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu
Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280
285 Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val
290 295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys
Glu Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met
Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400
Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405
410 415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu
Ala 420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
Leu Gly Lys 435 440 445 290214PRTArtificial SequenceSynthetic LC,
27596 290Glu Ile Val Met Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser
Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser
Val Ser Ser Tyr 20 25 30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln
Ala Pro Arg Leu Leu Ile 35 40 45 Tyr Asp Ala Ser Asn Arg Ala Thr
Gly Ile Pro Ala Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala
Val Tyr Tyr Cys Gln Gln His Thr Val Trp Pro Ile 85 90 95 Thr Phe
Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110
Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115
120 125 Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu
Ala 130 135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly
Asn Ser Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp
Ser Thr Tyr Ser Leu Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala
Asp Tyr Glu Lys His Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His
Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195
200 205 Phe Asn Arg Gly Glu Cys 210 291448PRTArtificial
SequenceSynthetic HC, 27535 291Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ser 1 5 10 15 Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Gly Thr Phe Ser Ser Tyr 20 25 30 Ala Ile Ser Trp Val
Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40 45 Gly Gly Ile
Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe 50 55 60 Gln
Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr 65 70
75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr
Cys 85 90 95 Ala Arg Asp Thr Pro Arg Trp Arg Tyr His Tyr Phe Asp
Tyr Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val Ser Ser Ala Ser
Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala Pro Cys Ser Arg
Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly Cys Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160 Ser Trp Asn Ser
Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165 170 175 Val Leu
Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val 180 185 190
Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn Val Asp His 195
200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val Glu Ser Lys Tyr
Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro Glu Phe Leu Gly
Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val Gln Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285 Lys Thr Lys Pro
Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290 295 300 Ser Val
Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 305 310 315
320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr
325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410 415 Arg Trp Gln
Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala 420 425 430 Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys 435 440
445 292214PRTArtificial SequenceSynthetic LC, 27535 292Glu Ile Val
Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu
Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Tyr 20 25
30 Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45 Tyr Asp Ala Ser Lys Arg Ala Thr Gly Ile Pro Ala Arg Phe
Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Glu Pro 65 70 75 80 Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Val Leu Asn Tyr Pro Leu 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val
Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe
Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val
Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val
Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155
160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser
165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys 210
293448PRTArtificial SequenceSynthetic HC, 27550 293Glu Val Gln Leu
Val Glu Ser Gly Gly Gly Leu Val Lys Pro Gly Gly 1 5 10 15 Ser Leu
Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25 30
Ser Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35
40 45 Ser Ser Ile Ser Ser Ser Ser Ser Tyr Ile Tyr Tyr Ala Asp Ser
Val 50 55 60 Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn
Ser Leu Tyr 65 70 75 80 Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Glu Arg Arg Gly Ser Leu Ala
Leu Gly Met Asp Val Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160
Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165
170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn
Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val
Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro
Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250 255 Thr Pro Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro 260 265 270 Glu Val
Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala 275 280 285
Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val 290
295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser Ser
Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser Gln Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu Val Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375 380 Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp 385 390 395 400 Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser 405 410
415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala
420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu
Gly Lys 435 440 445 294214PRTArtificial SequenceSynthetic LC, 27550
294Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly
1 5 10 15 Asp Arg Val Thr Ile Thr Cys Gln Ala Ser Gln Asp Ile Ser
Asn Tyr 20 25 30 Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro
Lys Leu Leu Ile 35 40 45 Tyr Asp Ala Ser Asn Leu Glu Thr Gly Val
Pro Ser Arg Phe Ser Gly 50 55 60 Ser Gly Ser Gly Thr Asp Phe Thr
Phe Thr Ile Ser Ser Leu Gln Pro 65 70 75 80 Glu Asp Ile Ala Thr Tyr
Tyr Cys Gln Gln Ser Tyr Phe Leu Pro Pro 85 90 95 Thr Phe Gly Gly
Gly Thr Lys Val Glu Ile Lys Arg Thr Val Ala Ala 100 105 110 Pro Ser
Val Phe Ile Phe Pro Pro Ser Asp Glu Gln Leu Lys Ser Gly 115 120 125
Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala 130
135 140 Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser
Gln 145 150 155 160 Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr
Tyr Ser Leu Ser 165 170 175 Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr
Glu Lys His Lys Val Tyr 180 185 190 Ala Cys Glu Val Thr His Gln Gly
Leu Ser Ser Pro Val Thr Lys Ser 195 200 205 Phe Asn Arg Gly Glu Cys
210 295455PRTArtificial SequenceSynthetic HC, 27549 295Gln Val Gln
Leu Val Glu Ser Gly Gly Gly Val Val Gln Pro Gly Arg 1 5 10 15 Ser
Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25
30 Gly Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45 Ala Val Ile Trp Tyr Asp Gly Ser Asn Lys Tyr Tyr Ala Asp
Ser Val 50 55 60 Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys
Asn Thr Leu Tyr 65 70 75 80 Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Asp Leu Gly Gly Tyr Ser
Tyr Gly Glu Pro Tyr Tyr Tyr Tyr 100 105 110 Tyr Gly Met Asp Val Trp
Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115 120 125 Ala Ser Thr Lys
Gly Pro Ser Val Phe Pro Leu Ala Pro Cys Ser Arg 130 135 140 Ser Thr
Ser Glu Ser Thr Ala Ala Leu Gly Cys Leu Val Lys Asp Tyr 145 150 155
160 Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser Gly Ala Leu Thr Ser
165 170 175 Gly Val His Thr Phe Pro Ala Val Leu Gln Ser Ser Gly Leu
Tyr Ser 180 185 190 Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser Leu
Gly Thr Lys Thr 195 200 205 Tyr Thr Cys Asn Val Asp His Lys Pro Ser
Asn Thr Lys Val Asp Lys 210 215 220 Arg Val Glu Ser Lys Tyr Gly Pro
Pro Cys Pro Pro Cys Pro Ala Pro 225 230 235 240 Glu Phe Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys 245 250 255 Asp Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val 260 265 270 Asp
Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp 275 280
285 Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe
290 295 300 Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His
Gln Asp 305 310 315 320 Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser Asn Lys Gly Leu 325 330 335 Pro Ser Ser Ile Glu Lys Thr Ile Ser
Lys Ala Lys Gly Gln Pro Arg 340 345 350 Glu Pro Gln Val Tyr Thr Leu
Pro Pro Ser Gln Glu Glu Met Thr Lys 355 360 365 Asn Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp 370 375 380 Ile Ala Val
Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys 385 390 395 400
Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser 405
410 415 Arg Leu Thr Val Asp Lys Ser Arg Trp Gln Glu Gly Asn Val Phe
Ser 420 425 430 Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr
Gln Lys Ser 435 440 445 Leu Ser Leu Ser Leu Gly Lys 450 455
296215PRTArtificial SequenceSynthetic LC, 27549 296Glu Ile Val Leu
Thr Gln Ser Pro Gly Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg
Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Asp 20 25 30
Tyr Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35
40 45 Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe
Ser 50 55 60 Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Arg Leu Glu 65 70 75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln
Ala His Ser Ser Pro 85 90 95 Tyr Thr Phe Gly Gly Gly Thr Lys Val
Glu Ile Lys Arg Thr Val Ala 100 105 110 Ala Pro Ser Val Phe Ile Phe
Pro Pro Ser Asp Glu Gln Leu Lys Ser 115 120 125 Gly Thr Ala Ser Val
Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu 130 135 140 Ala Lys Val
Gln Trp Lys Val Asp Asn Ala Leu Gln Ser Gly Asn Ser 145 150 155 160
Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu 165
170 175 Ser Ser Thr Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys
Val 180 185 190 Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro
Val Thr Lys 195 200 205 Ser Phe Asn Arg Gly Glu Cys 210 215
297448PRTArtificial SequenceSynthetic HC, 31414 297Gln Val Gln Leu
Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala 1 5 10 15 Ser Val
Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Lys Ser Tyr 20 25 30
Glu Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45 Gly Arg Ile Asn Pro Ser Val Gly Ser Thr Trp Tyr Ala Gln Lys
Phe 50 55 60 Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser
Thr Val Tyr 65 70 75 80 Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90 95 Ala Arg Gly Lys Arg Glu Gly Gly Thr
Glu Tyr Leu Arg Asn Trp Gly 100 105 110 Gln Gly Thr Leu Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser 115 120 125 Val Phe Pro Leu Ala
Pro Cys Ser Arg Ser Thr Ser Glu Ser Thr Ala 130 135 140 Ala Leu Gly
Cys Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val 145 150 155 160
Ser Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala 165
170 175 Val Leu Gln Ser Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val 180 185 190 Pro Ser Ser Ser Leu Gly Thr Lys Thr Tyr Thr Cys Asn
Val Asp His 195 200 205 Lys Pro Ser Asn Thr Lys Val Asp Lys Arg Val
Glu Ser Lys Tyr Gly 210 215 220 Pro Pro Cys Pro Pro Cys Pro Ala Pro
Glu Phe Leu Gly Gly Pro Ser 225 230 235 240 Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg 245 250
255 Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro
260 265 270 Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His
Asn Ala 275 280 285 Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr
Tyr Arg Val Val 290 295 300 Ser Val Leu Thr Val Leu His Gln Asp Trp
Leu Asn Gly Lys Glu Tyr 305 310 315 320 Lys Cys Lys Val Ser Asn Lys
Gly Leu Pro Ser Ser Ile Glu Lys Thr 325 330 335 Ile Ser Lys Ala Lys
Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu 340 345 350 Pro Pro Ser
Gln Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys 355 360 365 Leu
Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser 370 375
380 Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp
385 390 395 400 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val
Asp Lys Ser 405 410 415 Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala 420 425 430 Leu His Asn His Tyr Thr Gln Lys Ser
Leu Ser Leu Ser Leu Gly Lys 435 440 445 298214PRTArtificial
SequenceSynthetic LC, 31414 298Glu Ile Val Leu Thr Gln Ser Pro Gly
Thr Leu Ser Leu Ser Pro Gly 1 5 10 15 Glu Arg Ala Thr Leu Ser Cys
Arg Ala Ser Gln Ser Val Ser Ser Ser 20 25 30 Tyr Leu Ala Trp Tyr
Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35 40 45 Ile Tyr Gly
Ala Ser Ser Arg Ala Thr Gly Ile Pro Asp Arg Phe Ser 50 55 60 Gly
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu 65 70
75 80 Pro Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr His Ser Tyr
Ile 85 90 95 Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys Arg Thr
Val Ala Ala 100 105 110 Pro Ser Val Phe Ile Phe Pro Pro Ser Asp Glu
Gln Leu Lys Ser Gly 115 120 125 Thr Ala Ser Val Val Cys Leu Leu Asn
Asn Phe Tyr Pro Arg Glu Ala 130 135 140 Lys Val Gln Trp Lys Val Asp
Asn Ala Leu Gln Ser Gly Asn Ser Gln 145 150 155 160 Glu Ser Val Thr
Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser 165 170 175 Ser Thr
Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys His Lys Val Tyr 180 185 190
Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser 195
200 205 Phe Asn Arg Gly Glu Cys 210
References
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012
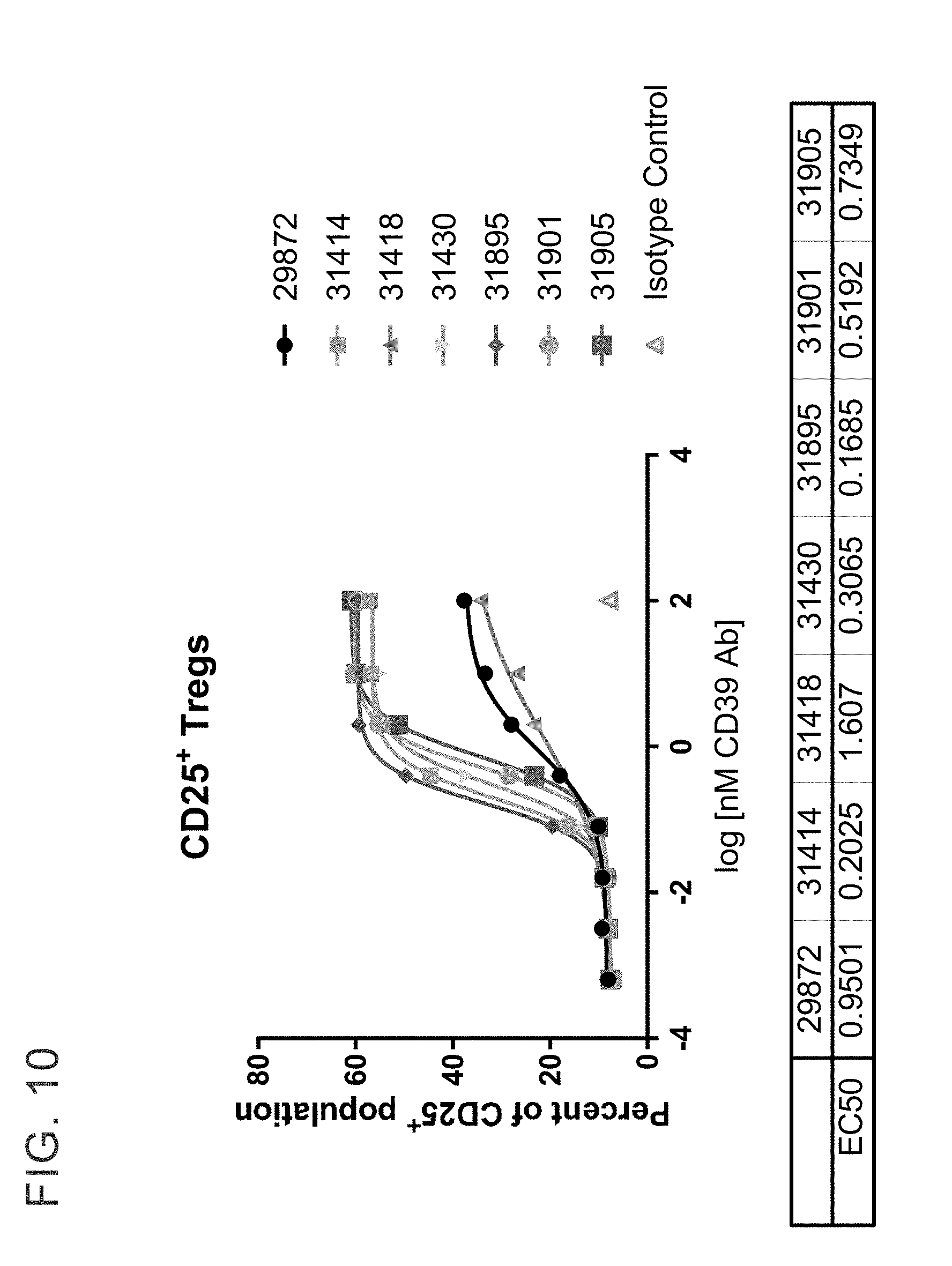
D00013

D00014
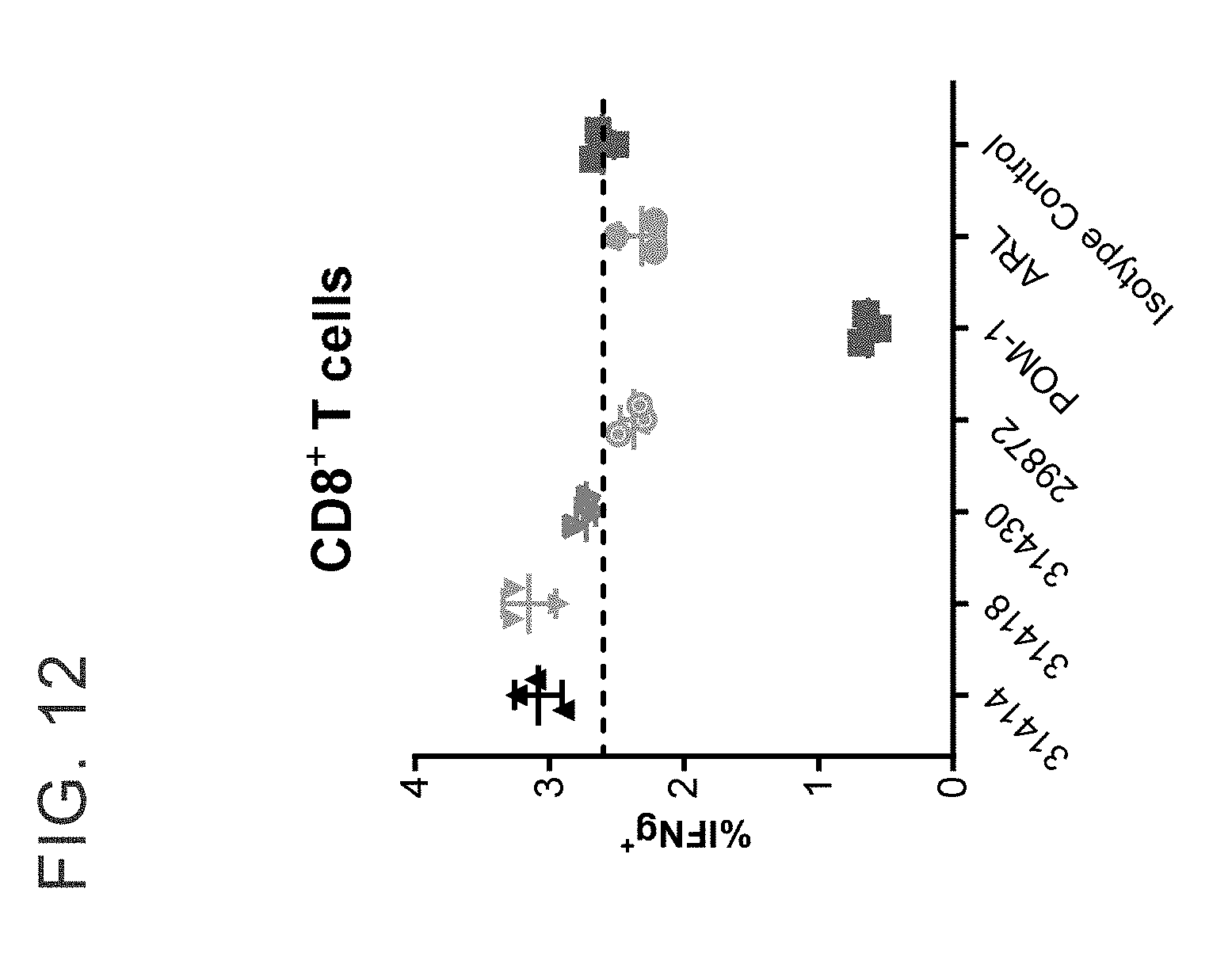
D00015

D00016
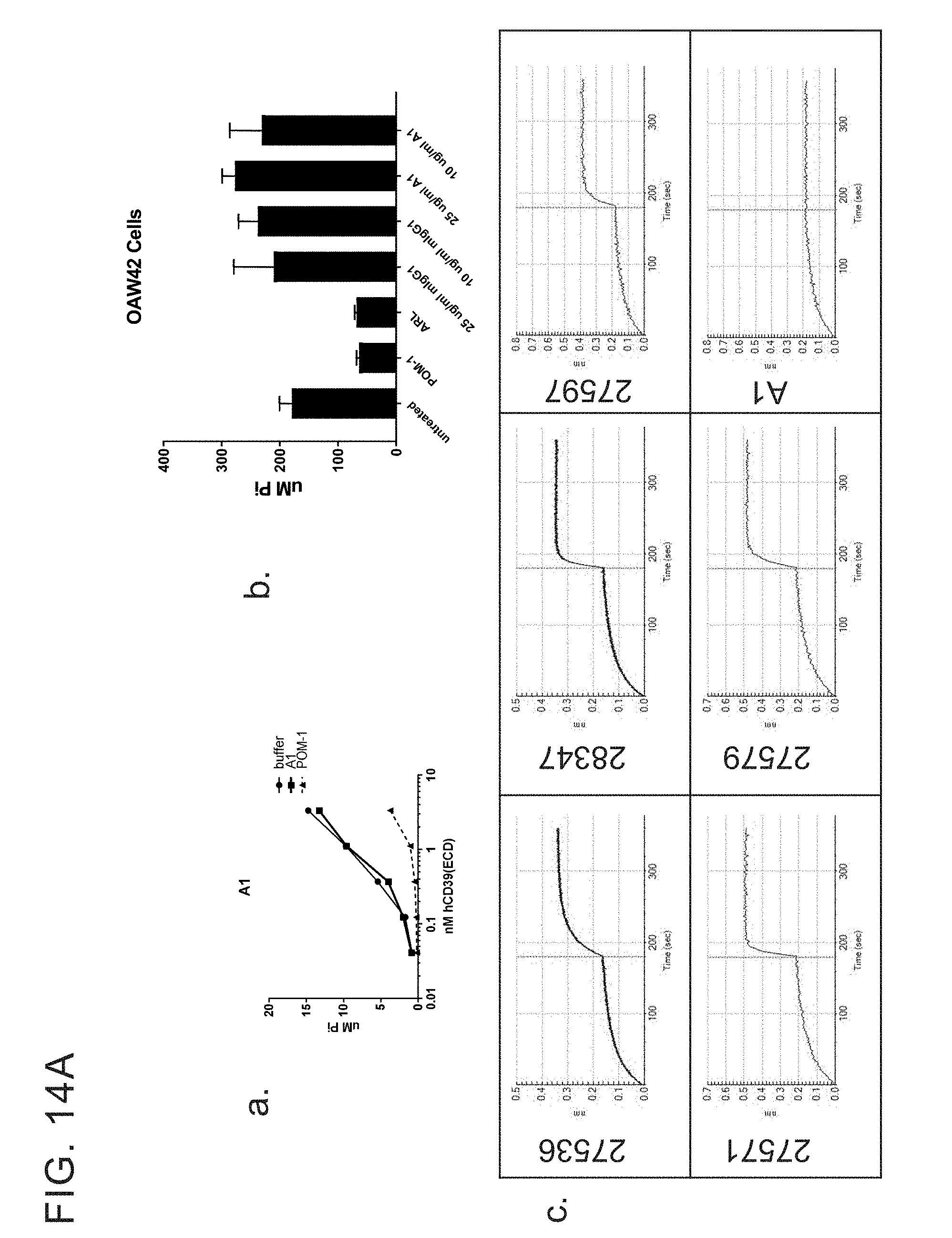
D00017
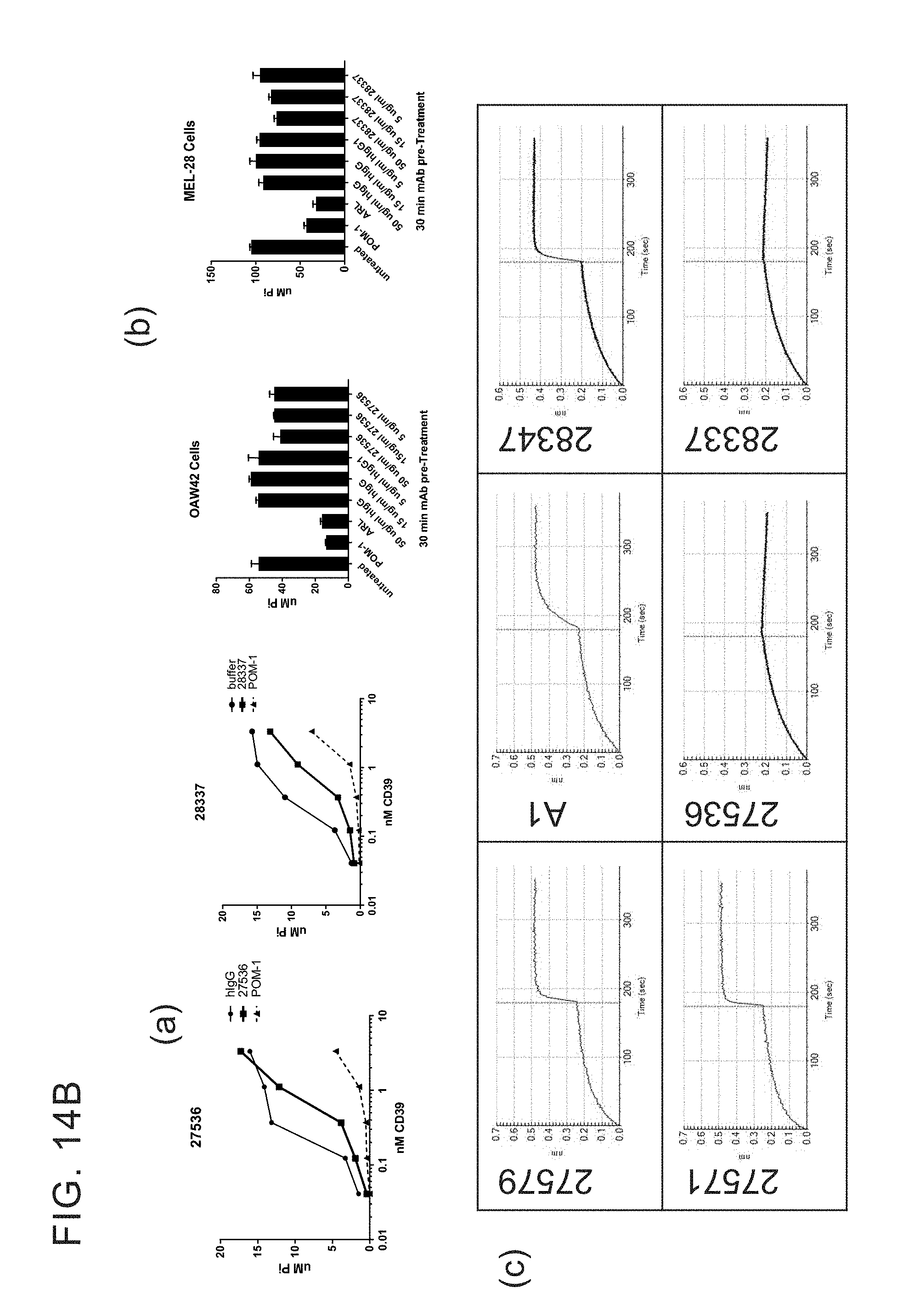
D00018

D00019
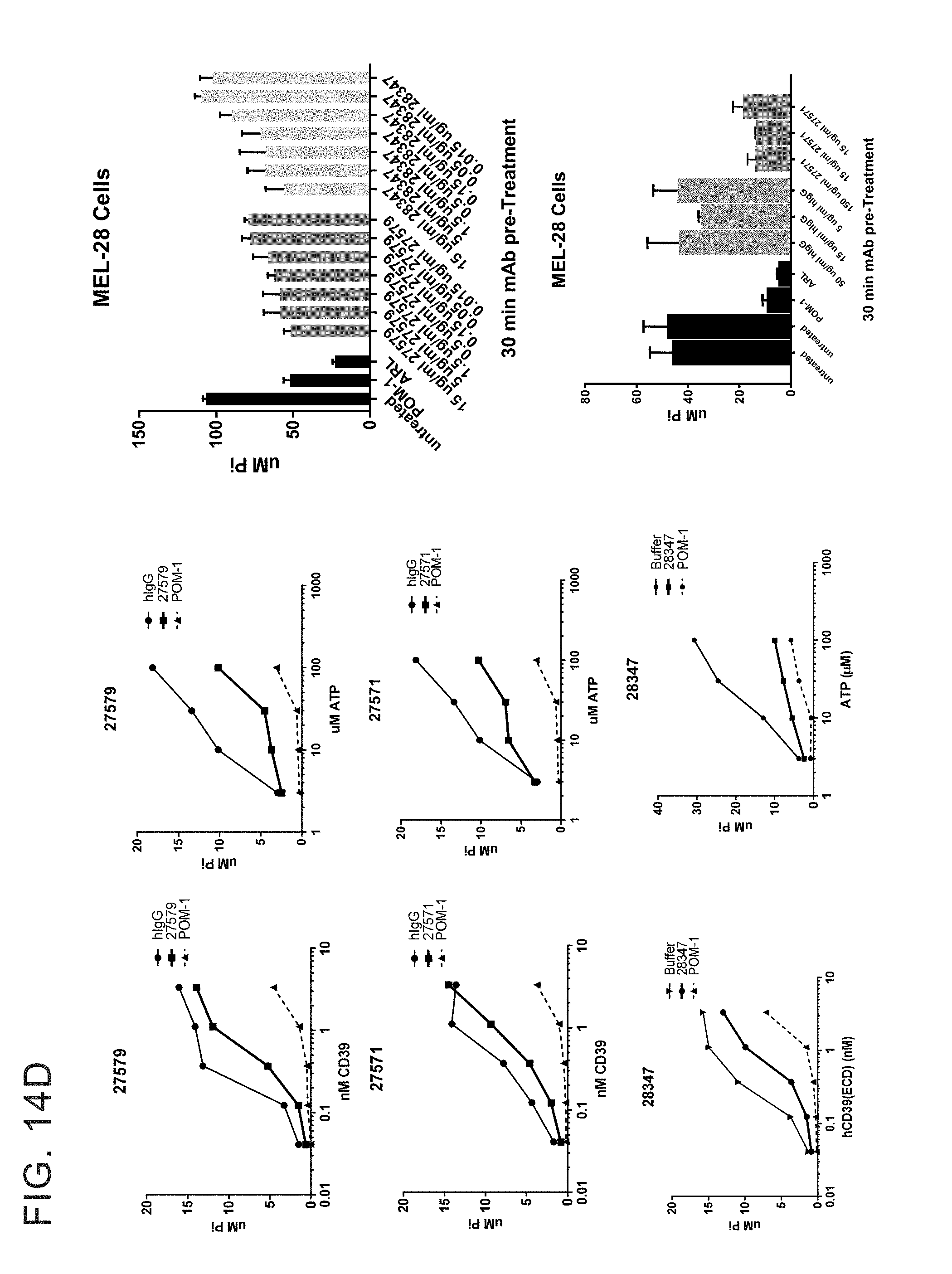
D00020

D00021

D00022
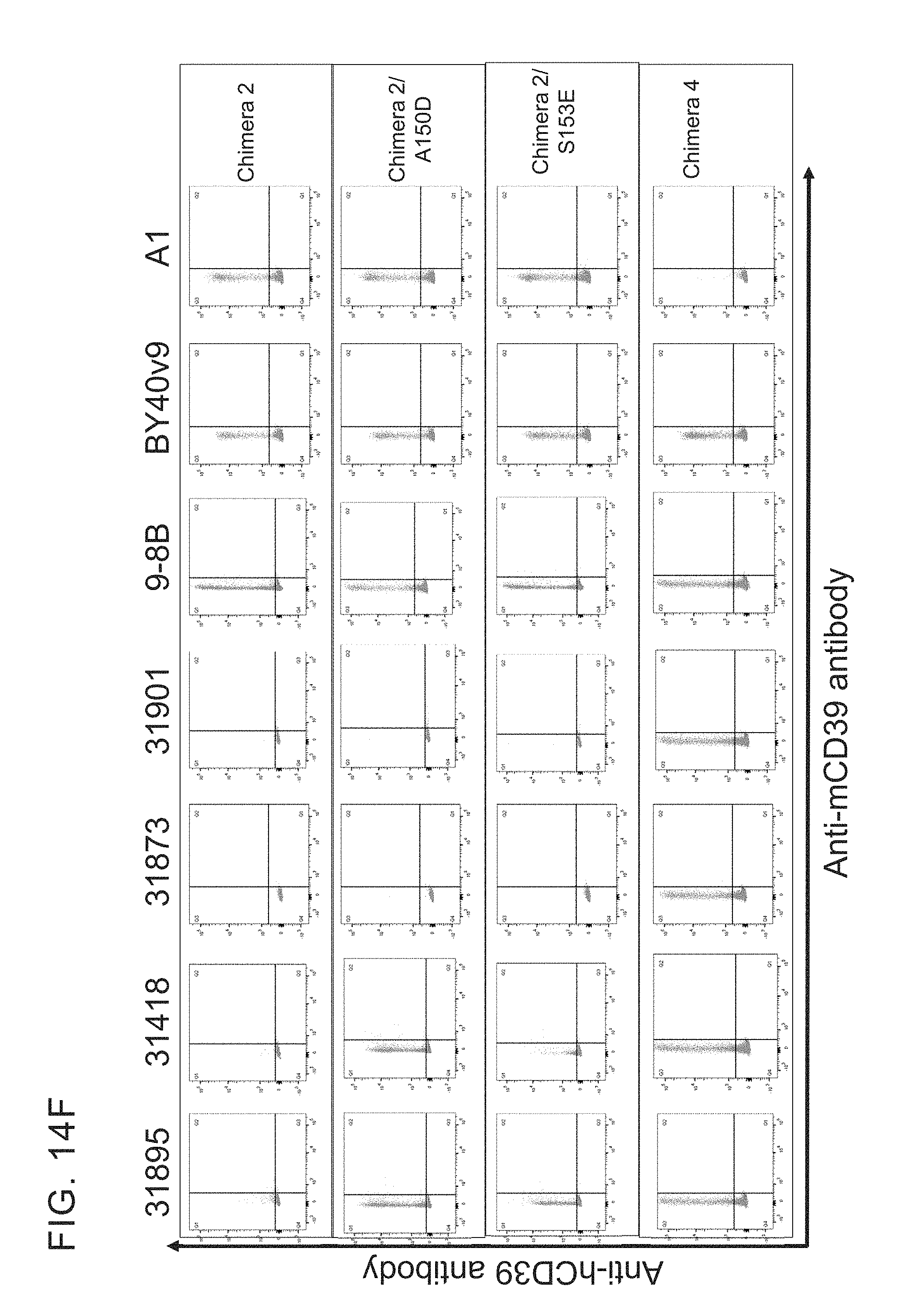
D00023

D00024

D00025

D00026
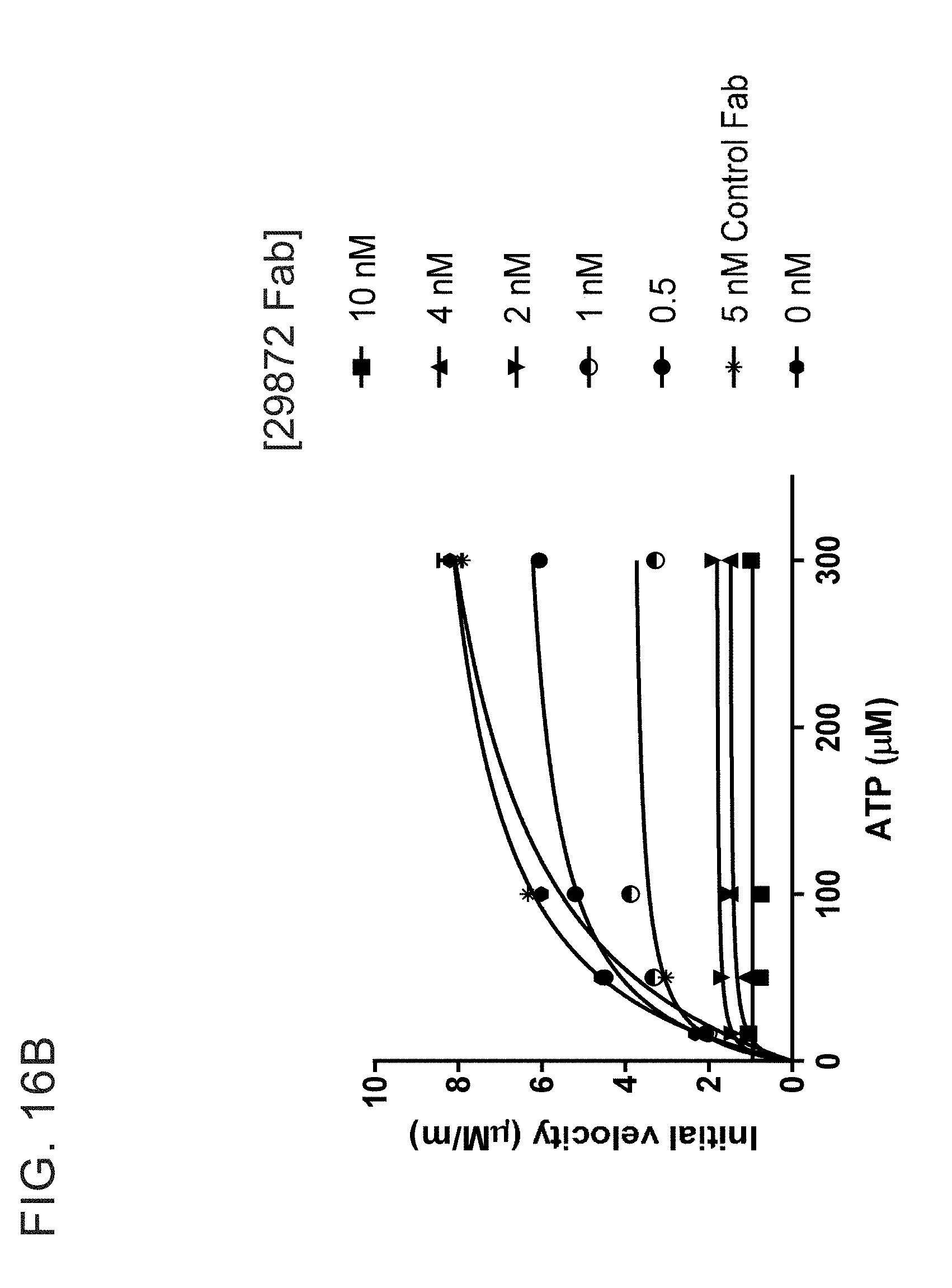
D00027

D00028

D00029

D00030

D00031

D00032

P00001

P00002

P00003
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.