Cell
Pule; Martin ; et al.
U.S. patent application number 16/100832 was filed with the patent office on 2019-02-07 for cell. The applicant listed for this patent is UCL BUSINESS PLC. Invention is credited to Shaun Cordoba, Khai Kong, Martin Pule.
| Application Number | 20190038672 16/100832 |
| Document ID | / |
| Family ID | 52003983 |
| Filed Date | 2019-02-07 |











View All Diagrams
| United States Patent Application | 20190038672 |
| Kind Code | A1 |
| Pule; Martin ; et al. | February 7, 2019 |
Cell
Abstract
The present invention provides a cell which co-expresses a first chimeric antigen receptor (CAR) and second CAR at the cell surface, each CAR comprising: (i) an antigen-binding domain; (ii) a spacer (iii) a trans-membrane domain; and (iv) an endodomain wherein the antigen binding domains of the first and second CARs bind to different antigens, and wherein each of the first and second CARs is an activating CAR comprising an activating endodomain.
| Inventors: | Pule; Martin; (London, GB) ; Kong; Khai; (London, GB) ; Cordoba; Shaun; (London, GB) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 52003983 | ||||||||||
| Appl. No.: | 16/100832 | ||||||||||
| Filed: | August 10, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15037391 | May 18, 2016 | |||
| PCT/GB2014/053451 | Nov 21, 2014 | |||
| 16100832 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61K 2039/5158 20130101; C12N 2740/15043 20130101; C07K 2319/02 20130101; C07K 2319/01 20130101; C12N 5/0636 20130101; A61P 35/00 20180101; A61P 31/00 20180101; A61P 37/06 20180101; C12N 9/16 20130101; C07K 16/2803 20130101; C12Y 301/03048 20130101; C12N 2740/10043 20130101; C07K 2319/74 20130101; C12N 5/0637 20130101; C07K 14/70517 20130101; C07K 14/70521 20130101; A61K 35/17 20130101; A61P 37/02 20180101; A61K 38/00 20130101; A61P 31/12 20180101; C12N 5/0638 20130101; C12N 2510/00 20130101; A61P 37/08 20180101; C07K 14/70589 20130101; C07K 2317/622 20130101; A61K 39/0011 20130101; A61P 35/02 20180101; C07K 2317/52 20130101; C07K 14/7051 20130101; C12N 5/0646 20130101; C07K 2319/03 20130101; C12N 15/86 20130101; C07K 14/70503 20130101 |
| International Class: | A61K 35/17 20150101 A61K035/17; A61K 39/00 20060101 A61K039/00; C12N 5/0783 20100101 C12N005/0783; C07K 14/705 20060101 C07K014/705; C07K 14/725 20060101 C07K014/725; C07K 16/28 20060101 C07K016/28; C12N 15/86 20060101 C12N015/86; C12N 9/16 20060101 C12N009/16 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Nov 21, 2013 | GB | 1320573.7 |
| Jun 19, 2014 | GB | 1410934.2 |
Claims
1. A T cell which co-expresses a first chimeric antigen receptor (CAR) and second CAR at the cell surface, each CAR comprising: (i) an antigen-binding domain; (ii) a spacer (iii) a trans-membrane domain; and (iv) an endodomain; wherein the first CAR binds CD19 and the second CAR binds CD20.
2-4. (canceled)
5. A T cell according to claim 1 which comprises more than two CARs such that it is specifically stimulated by a cell, such as a target cell, bearing a distinct pattern of more than two antigens.
6. A nucleic acid sequence encoding first and second chimeric antigen receptors (CARs) each CAR comprising: (i) an antigen-binding domain; (ii) a spacer (iii) a trans-membrane domain; and (iv) an endodomain; wherein the first CAR binds CD19 and the second CAR binds CD20.
7. A nucleic acid sequence according to claim 6, which has the following structure: AgB1-spacer1-TM1-endo1-coexpr-AbB2-spacer2-TM2-endo2 in which AgB1 is a nucleic acid sequence encoding the antigen-binding domain of the first CAR; spacer 1 is a nucleic acid sequence encoding the spacer of the first CAR; TM1 is a a nucleic acid sequence encoding the transmembrane domain of the first CAR; endo 1 is a nucleic acid sequence encoding the endodomain of the first CAR; coexpr is a nucleic acid sequence enabling co-expression of both CARs AgB2 is a nucleic acid sequence encoding the antigen-binding domain of the second CAR; spacer 2 is a nucleic acid sequence encoding the spacer of the second CAR; TM2 is a a nucleic acid sequence encoding the transmembrane domain of the second CAR; endo 2 is a nucleic acid sequence encoding the endodomain of the second CAR; which nucleic acid sequence, when expressed in a T cell, encodes a polypeptide which is cleaved at the cleavage site such that the first and second CARs are co-expressed at the T cell surface.
8. A nucleic acid sequence according to claim 7, wherein coexpr encodes a sequence comprising a self-cleaving peptide.
9. A nucleic acid sequence according to claim 7 or 8, wherein alternative codons are used in regions of sequence encoding the same or similar amino acid sequences, in order to avoid homologous recombination.
10-12. (canceled)
13. A vector comprising a nucleic acid sequence according to claim 6.
14. A retroviral vector or a lentiviral vector or a transposon according to claim 13.
15. A method for making a T cell according to claim 1, which comprises the step of introducing into a T cell: a nucleic acid sequence according to claim 6 or a vector according to claim 13.
16. A method according to claim 15, wherein the T cell is from a sample isolated from a subject.
17. A pharmaceutical composition comprising a plurality of T cells according claim 1.
18. A method for treating and/or preventing a disease, which comprises the step of administering a pharmaceutical composition according to claim 17 to a subject.
19. A method according to claim 18, which comprises the following steps: (i) isolation of a T cell-containing sample from a subject; (ii) transduction or transfection of the T cells with: a nucleic acid sequence according to claim 6; or a vector according to claim 13; and (iii) administering the T cells from (ii) to a the subject.
20. A method according to claim 18, wherein the disease is a cancer.
21-26. (canceled)
27. A natural killer (NK) cell which co-expresses a first chimeric antigen receptor (CAR) and second CAR at the cell surface, each CAR comprising: (i) an antigen-binding domain; (ii) a spacer (iii) a trans-membrane domain; and (iv) an endodomain; wherein the first CAR binds CD19 and the second CAR binds CD20.
28. A cell composition comprising CAR expressing T cells according to claim 1 and/or CAR expressing NK cells according to claim 27 made by transducing a blood-sample ex vivo with a nucleic acid encoding the first and second CARs.
Description
FIELD OF THE INVENTION
[0001] The present invention relates to a cell which comprises more than one chimeric antigen receptor (CAR). The cell may be capable of specifically recognising a target cell, due to a differential pattern of expression (or non-expression) of two or more antigens by the target cell.
BACKGROUND TO THE INVENTION
[0002] A number of immunotherapeutic agents have been described for use in cancer treatment, including therapeutic monoclonal antibodies (mAbs), immunoconjugated mAbs, radioconjugated mAbs and bi-specific T-cell engagers.
[0003] Typically these immunotherapeutic agents target a single antigen: for instance, Rituximab targets CD20; Myelotarg targets CD33; and Alemtuzumab targets CD52.
[0004] However, it is relatively rare for the presence (or absence) of a single antigen effectively to describe a cancer.
[0005] A particular problem in the field of oncology is provided by the Goldie-Coldman hypothesis: which describes that the sole targeting of a single antigen may result in tumour escape by modulation of said antigen due to the high mutation rate inherent in most cancers. This modulation of antigen expression may reduce the efficacy of known immunotherapeutics.
[0006] Tumour heterogeneity describes the observation that different tumour cells can show distinct morphological and phenotypic profiles, including cellular morphology, gene expression, metabolism, proliferation and metastatic potential. This phenomenon occurs both between tumours and within tumours. Tumour heterogeneity has been observed in leukaemias, breast, prostate, colon, brain, head and neck, bladder and gynecological cancers, for example. Tumour heterogeneity may result in the expression of different antigens on the surface of cells within a tumour or between tumours. This heterogeneity of cancer cells introduces significant challenges in designing effective treatment strategies.
[0007] There is thus a need for immunotherapeutic agents which are capable of more targeting to reflect the complex pattern of marker expression that is associated with many cancers.
[0008] Chimeric Antigen Receptors (CARs)
[0009] Chimeric antigen receptors are proteins which graft the specificity of a monoclonal antibody (mAb) to the effector function of a T-cell. Their usual form is that of a type I transmembrane domain protein with an antigen recognizing amino terminus, a spacer, a transmembrane domain all connected to a compound endodomain which transmits T-cell survival and activation signals (see FIG. 1A).
[0010] The most common form of these molecules are fusions of single-chain variable fragments (scFv) derived from monoclonal antibodies which recognize a target antigen, fused via a spacer and a trans-membrane domain to a signaling endodomain. Such molecules result in activation of the T-cell in response to recognition by the scFv of its target. When T cells express such a CAR, they recognize and kill target cells that express the target antigen. Several CARs have been developed against tumour associated antigens, and adoptive transfer approaches using such CAR-expressing T cells are currently in clinical trial for the treatment of various cancers.
[0011] It has been observed that using a CAR approach for cancer treatment, tumour heterogeneity and immunoediting can cause escape from CAR treatment. For example, in the study described by Grupp et al (2013; New Eng. J. Med 368:1509-1518) CAR-modified T cell approach was used for the treatment of acute lymphoid leukemia. It was found that one patient of the anti-CD19 CAR trial relapsed with CD19-negative disease two months after treatment.
[0012] There is thus a need for alternative CAR treatment approaches which address the problems of cancer escape and tumour heterogeneity.
[0013] Expression of Two CAR Binding Specificities
[0014] Bispecific CARs known as tandem CARs or TanCARs have been developed in an attempt to target multiple cancer specific markers simultaneously. In a TanCAR, the extracellular domain comprises two antigen binding specificities in tandem, joined by a linker. The two binding specificities (scFvs) are thus both linked to a single transmembrane portion: one scFv being juxtaposed to the membrane and the other being in a distal position.
[0015] Grada et al (2013, Mol Ther Nucleic Acids 2:e105 describes a TanCAR which includes a CD19-specific scFv, followed by a Gly-Ser linker and then a HER2-specific scFv. The HER2-scFv was in the juxta-membrane position, and the CD19-scFv in the distal position. The Tan CAR was shown to induce distinct T cell reactivity against each of the two tumour restricted antigens. This arrangement was chosen because the respective lengths of HER2 (632 aa/125 .ANG.) and CD19 (280aa, 65 .ANG.) lends itself to that particular spatial arrangement. It was also known that the HER2 scFv bound the distal-most 4 loops of HER2.
[0016] The problem with this approach is that the juxta-membrane scFv may be inaccessible due to the presence of the distal scFv, especially which it is bound to the antigen. In view of the need to choose the relative positions of the two scFvs in view of the spatial arrangement of the antigen on the target cell, it may not be possible to use this approach for all scFv binding pairs. Moreover, it is unlikely that the TanCar approach could be used for more than two scFvs, a TanCAR with three or more scFvs would be a very large molecule and the scFvs may well fold back on each other, obscuring the antigen-binding sites. It is also doubtful that antigen-binding by the most distal scFv, which is separated from the transmembrane domain by two or more further scFvs, would be capable of triggering T cell activation.
[0017] There is thus a need for an alternative approach to express two CAR binding specificities on the surface of a cell such as a T cell.
DESCRIPTION OF THE FIGURES
[0018] FIG. 1A-1D: FIG. 1A. Generalized architecture of a CAR: A binding domain recognizes antigen; the spacer elevates the binding domain from the cell surface; the trans-membrane domain anchors the protein to the membrane and the endodomain transmits signals. FIG. 1B-1D. Different generations and permutations of CAR endodomains: FIG. 1B. initial designs transmitted ITAM signals alone through Fc.epsilon.R1-.gamma. or CD3.zeta. endodomain, while later designs transmitted additional FIG. 1C. one or FIG. 1D. two co-stimulatory signals in cis.
[0019] FIG. 2: Schematic diagram illustrating the invention
[0020] The invention relates to engineering T-cells to respond to logical rules of target cell antigen expression. This is best illustrated with an imaginary FACS scatter-plot. Target cell populations express both, either or neither of antigens "A" and "B". Different target populations (marked in red) are killed by T-cells transduced with a pair of CARs connected by different gates. With OR gated receptors, both single-positive and double-positive cells will be killed. With AND gated receptors, only double-positive target cells are killed. With AND NOT gating, double-positive targets are preserved while single-positive targets
[0021] FIG. 3: Creation of target cell populations
[0022] SupT1 cells were used as target cells. These cells were transduced to express either CD19, CD33 or both CD19 and CD33. Target cells were stained with appropriate antibodies and analysed by flow cytometry.
[0023] FIG. 4: Cassette design for an OR gate
[0024] A single open reading frame provides both CARs with an in-frame FMD-2A sequence resulting in two proteins. Signal1 is a signal peptide derived from IgG1 (but can be any effective signal peptide). scFv1 is the single-chain variable segment which recognizes CD19 (but can be a scFv or peptide loop or ligand or in fact any domain which recognizes any desired arbitrary target). STK is the CD8 stalk but may be any suitable extracellular domain. CD28tm is the CD28 trans-membrane domain but can by any stable type I protein transmembrane domain and CD3Z is the CD3 Zeta endodomain but can be any endodomain which contains ITAMs. Signal2 is a signal peptide derived from CD8 but can be any effective signal peptide which is different in DNA sequence from signal1. scFv recognizes CD33 but as for scFv1 is arbitrary. HC2CH3 is the hinge-CH2-CH3 of human IgG1 but can be any extracellular domain which does not cross-pair with the spacer used in the first CAR. CD28tm' and CD3Z' code for the same protein sequence as CD28tm and CD3Z but are codon-wobbled to prevent homologous recombination.
[0025] FIG. 5A-5B: Schematic representation of the chimeric antigen receptors (CARs) for an OR gate
[0026] Stimulatory CARs were constructed consisting of either an N-terminal FIG. 5A. anti-CD19 scFv domain followed by the extracellular hinge region of human CD8 or FIG. 5B. anti-CD33 scFv domain followed by the extracellular hinge, CH2 and CH3 (containing a pvaa mutation to reduce FcR binding) region of human IgG1. Both receptors contain a human CD28 transmembrane domain and a human CD3 Zeta (CD247) intracellular domain. "S" depicts the presence of disulphide bonds.
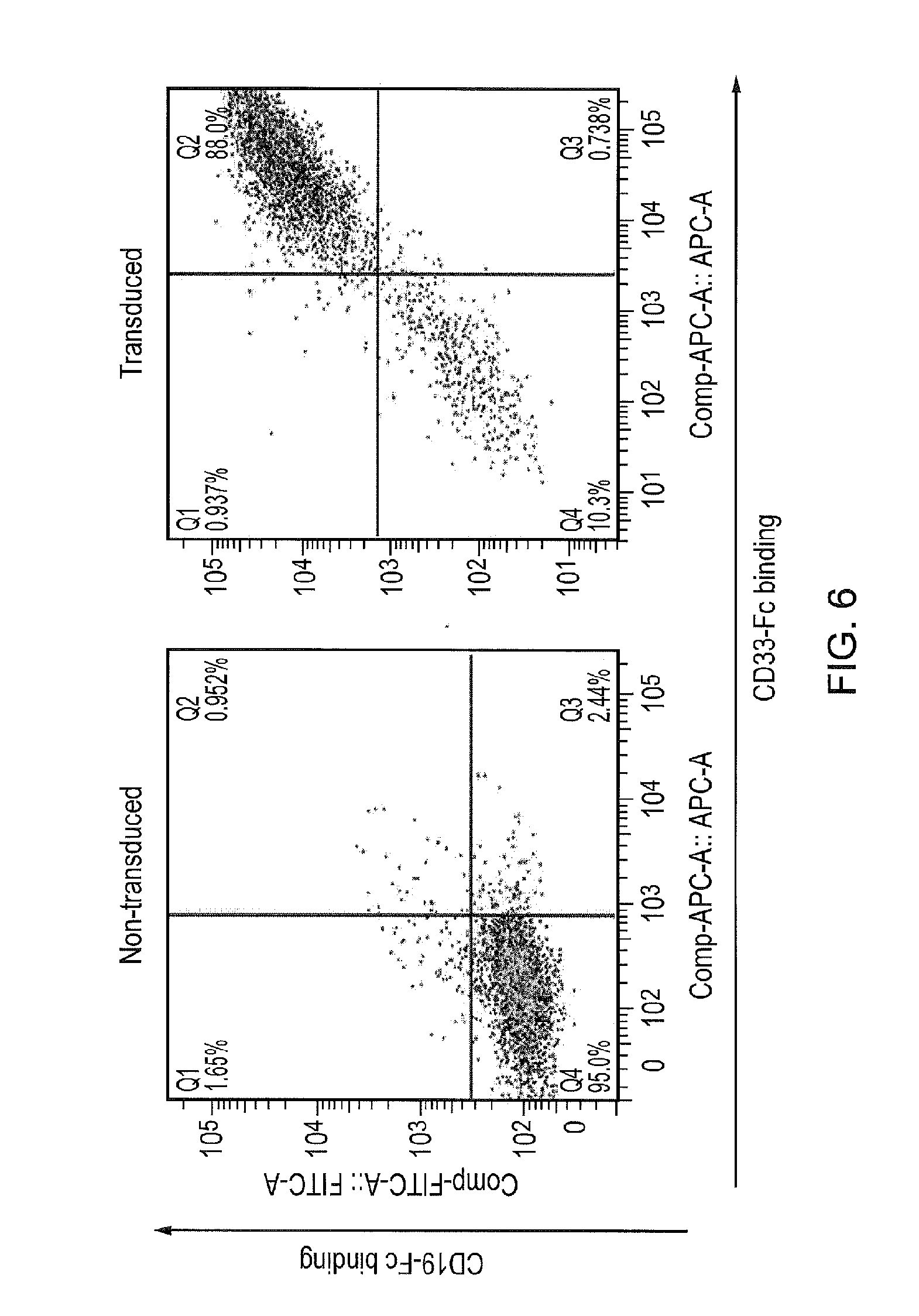
[0027] FIG. 6: Expression data showing co-expression of both CARs on the surface of one T-cell.
[0028] FIG. 7: Functional analysis of the OR gate
[0029] Effector cells (5.times.10 4 cells) expressing the OR gate construct were co-incubated with a varying number of target cells and IL-2 was analysed after 16 hours by ELISA. The graph displays the average maximum IL-2 secretion from a chemical stimulation (PMA and lonomycin) of the effector cells alone and the average background IL-2 from effector cells without any stimulus from three replicates.
[0030] FIG. 8: Cartoon showing both versions of the cassette used to express both AND gates Activating and inhibiting CARs were co-expressed once again using a FMD-2A sequence. Signal1 is a signal peptide derived from IgG1 (but can be any effective signal peptide). scFv1 is the single-chain variable segment which recognizes CD19 (but can be a scFv or peptide loop or ligand or in fact any domain which recognizes any desired arbitrary target). STK is the CD8 stalk but may be any non-bulky extracellular domain. CD28tm is the CD28 trans-membrane domain but can by any stable type I protein transmembrane domain and CD3Z is the CD3 Zeta endodomain but can be any endodomain which contains ITAMs. Signal2 is a signal peptide derived from CD8 but can be any effective signal peptide which is different in DNA sequence from signal1. scFv recognizes CD33 but as for scFv1 is arbitrary. HC2CH3 is the hinge-CH2-CH3 of human IgG1 but can be any bulky extracellular domain. CD45 and CD148 are the transmembrane and endodomains of CD45 and CD148 respectively but can be derived from any of this class of protein.
[0031] FIG. 9: Schematic representation of the protein structure of chimeric antigen receptors (CARs) for the AND gates
[0032] The stimulatory CAR consisting of an N-terminal anti-CD19 scFv domain followed by the extracellular stalk region of human CD8, human CD28 transmembrane domain and human CD3 Zeta (CD247) intracellular domain. Two inhibitory CARs were tested. These consist of an N-terminal anti-CD33 scFv domain followed by the extracellular hinge, CH2 and CH3 (containing a pvaa mutation to reduce FcR binding) region of human IgG1 followed by the transmembrane and intracellular domain of either human CD148 or CD45. "S" depicts the presence of disulphide bonds.
[0033] FIG. 10: Co-expression of activation and inhibitory CARs
[0034] BW5147 cells were used as effector cells and were transduced to express both the activation anti-CD19 CAR and one of the inhibitory anti-CD33 CARs. Effector cells were stained with CD19-mouse-Fc and CD33-rabbit-Fc and with appropriate secondary antibodies and analysed by flow cytometry.
[0035] FIG. 11A-11 B: Functional analysis of the AND gates
[0036] Effector cells (5.times.10 4 cells) expressing activation anti-CD19 CAR and the inhibitory anti-CD33 CAR with the FIG. 11A. CD148 or FIG. 11B. CXD45 intracellular domain were co-incubated with a varying number of target cells and IL-2 was analysed after 16 hours by ELISA. The graph displays the maximum IL-2 secretion from a chemical stimulation (PMA and lonomycin) of the effector cells alone and the background IL-2 from effector cells without any stimulus from three replicates.
[0037] FIG. 12: Cartoon showing three versions of the cassette used to generate the AND NOT gate
[0038] Activating and inhibiting CARs were co-expressed once again using a FMD-2A sequence. Signal1 is a signal peptide derived from IgG1 (but can be any effective signal peptide). scFv1 is the single-chain variable segment which recognizes CD19 (but can be a scFv or peptide loop or ligand or in fact any domain which recognizes any desired arbitrary target). STK is the human CD8 stalk but may be any non-bulky extracellular domain. CD28tm is the CD28 trans-membrane domain but can by any stable type I protein transmembrane domain and CD3Z is the CD3 Zeta endodomain but can be any endodomain which contains ITAMs. Signal2 is a signal peptide derived from CD8 but can be any effective signal peptide which is different in DNA sequence from signal1. scFv recognizes CD33 but as for scFv1 is arbitrary. muSTK is the mouse CD8 stalk but can be any spacer which co-localises but does not cross-pair with that of the activating CAR. dPTPN6 is the phosphatase domain of PTPN6. LAIR1 is the transmembrane and endodomain of LAIR1. 2Aw is a codon-wobbled version of the FMD-2A sequence. SH2-CD148 is the SH2 domain of PTPN6 fused with the phosphatase domain of CD148.
[0039] FIG. 13A-13D: Schematic representation of the chimeric antigen receptors (CARs) for the NOT AND gates
[0040] FIG. 13A. A stimulatory CAR consisting of an N-terminal anti-CD19 scFv domain followed by the stalk region of human CD8, human CD28 transmembrane domain and human CD247 intracellular domain. FIG. 13B. An inhibitory CAR consisting of an N-terminal anti-CD33 scFv domain followed by the stalk region of mouse CD8, transmembrane region of mouse CD8 and the phosphatase domain of PTPN6. FIG. 13C. an inhibitory CAR consisting of an N-terminal anti-CD33 scFv domain followed by the stalk region of mouse CD8 and the transmembrane and intracellular segments of LAIR1. FIG. 13D. An inhibitory CAR identical to previous CAR except it is co-expressed with a fusion protein of the PTPN6 SH2 domain and the CD148 phosphatase domain.
[0041] FIG. 14A-14B: Functional analysis of the NOT AND gate
[0042] Effector cells (5.times.10 4 cells) expressing the FIG. 14A. full length SHP-1 or FIG. 14B. truncated form of SHP-1 were co-incubated with a varying number of target cells and IL-2 was analysed after 16 hours by ELISA. The graph displays the average maximum IL-2 secretion from a chemical stimulation (PMA and lonomycin) of the effector cells alone and the average background IL-2 from effector cells without any stimulus from three replicates.
[0043] FIG. 15: Amino acid sequence of an OR gate
[0044] FIG. 16: Amino acid sequence of a CD148 and a CD45 based AND gate
[0045] FIG. 17: Amino acid sequence of three AND NOT gates
[0046] FIG. 18A-18C: Dissection of AND gate function
[0047] FIG. 18A. The prototype AND gate is illustrated on the right and its function in response to CD19, CD33 single and CD19, CD33 double positive targets is shown on the left. FIG. 18B. The scFvs are swapped so the activating endodomain is triggered by CD33 and the inhibitory endodomain is activated by CD19. This AND gate remains functional despite this scFv swap. FIG. 18C. The CD8 mouse stalk replaced Fc in the spacer of the inhibitory CAR. With this modification, the gate fails to respond to either CD19 single positive or CD19, CD33 double positive targets.
[0048] FIG. 19A-19D: Expression of target antigens on artificial target cells
[0049] FIG. 19A. Shows flow cytometry scatter plots CD19 vs CD33 of the original set of artificial target cells derived from SupT1 cells. From left to right: double negative SupT1 cells, SupT1 cells positive for CD19, positive for CD33 and positive for both CD19 and CD33. FIG. 19B. Shows flow cytometry scatter plots CD19 vs GD2 of the artificial target cells generated to test the CD19 AND GD2 gate: From left to right: negative SupT1 cells, SupT1 cells expressing CD19, SupT1 cells transduced with GD2 and GM3 synthase vectors which become GD2 positive and SupT1 cells transduced with CD19 as well as GD2 and GM3 synthase which are positive for both GD2 and CD19. FIG. 19C. Shows flow cytometry scatter plots of CD19 vs EGFRvIII of the artificial targets generated to test the CD19 AND EGFRvIII gate. From left to right: negative SupT1 cells, SupT1 cells expressing CD19, SupT1 cells transduced with EGFRvIII and SupT1 cells transduced with both CD19 and EGFRvIII. FIG. 19D. Shows flow cytometry scatter plots of CD19 vs CD5 of the artificial targets generated to test the CD19 AND CD5 gate. From left to right: negative 293T cells, 293T cells transduced with CD19, 293T cells transduced with CD5, 293T cells transduced with both CD5 and CD19 vectors.
[0050] FIG. 20A-20D: Generalizability of the AND gate
[0051] FIG. 20A. Cartoon of AND gate modified so the second CAR's specificity is changed from the original specificity of CD33, to generate 3 new CARs: CD19 AND GD2, CD19 AND EGFRvIII, CD19 AND CD5. FIG. 20B. CD19 AND GD2 AND gate: Left: expression of AND gate is shown recombinant CD19-Fc staining (x-axis) for the CD19 CAR, versus anti-human-Fc staining (Y-axis) for the GD2 CAR. Right: function in response to single positive and double positive targets. FIG. 20C. CD19 AND EGFRvIII AND gate: Left: expression of AND gate is shown recombinant CD19-Fc staining (x-axis) for the CD19 CAR, versus anti-human-Fc staining (Y-axis) for the EGFRvIII CAR. Right: function in response to single positive and double positive targets. FIG. 20D. CD19 AND CD5 AND gate: Left: expression of AND gate is shown recombinant CD19-Fc staining (x-axis) for the CD19 CAR, versus anti-human-Fc staining (Y-axis) for the CD5 CAR. Right: function in response to single positive and double positive targets.
[0052] FIG. 21A-21C: Function of the AND NOT gates
[0053] Function of the three implementations of an AND NOT gate is shown. A cartoon of the gates tested is shown to the right, and function in response to single positive and double positive targets is shown to the left. FIG. 21A. PTPN6 based AND NOT gate whereby the first CAR recognizes CD19, has a human CD8 stalk spacer and an ITAM containing activating endodomain; is co-expressed with a second CAR that recognizes CD33, has a mouse CD8 stalk spacer and has an endodomain comprising of a PTPN6 phosphatase domain. FIG. 21B. ITIM based AND NOT gate is identical to the PTPN6 gate, except the endodomain is replaced by the endodomain from LAIR1. FIG. 21C. CD148 boosted AND NOT gate is identical to the ITIM based gate except an additional fusion between the PTPN6 SH2 and the endodomain of CD148 is expressed. All three gates work as expected with activation in response to CD19 but not in response to CD19 and CD33 together.
[0054] FIG. 22A-22C: Dissection of PTPN6 based AND NOT gate function The original PTPN6 based AND NOT gate is compared with several controls to demonstrate the model. A cartoon of the gates tested is shown to the right, and function in response to single positive and double positive targets is shown to the left. FIG. 22A. Original AND NOT gate whereby the first CAR recognizes CD19, has a human CD8 stalk spacer and an ITAM containing activating endodomain; is co-expressed with a second CAR recognizes CD33, has a mouse CD8 stalk spacer and has an endodomain comprising of a PTPN6 phosphatase domain. FIG. 22B. AND NOT gate modified so the mouse CD8 stalk spacer is replaced with an Fc spacer. FIG. 22C. AND NOT gate modified so that the PTPN6 phosphatase domain is replaced with the endodomain from CD148. Original AND NOT gate (FIG. 22A.) functions as expected triggering in response to CD19, but not in response to both CD19 and CD33. The gate in FIG. 22B. triggers both in response to CD19 along or CD19 and CD33 together. The gate in FIG. 22C. does not trigger in response to one or both targets.
[0055] FIG. 23A-23B: Dissection of LAIR1 based AND NOT gate
[0056] Functional activity against CD19 positive, CD33 positive and CD19, CD33 double-positive targets is shown. FIG. 23A. Structure and activity of the original ITIM based AND NOT gate. This gate is composed of two CARs: the first recognizes CD19, has a human CD8 stalk spacer and an ITAM containing endodomain; the second CAR recognizes CD33, has a mouse CD8 stalk spacer and an ITIM containing endodomain. FIG. 23B. Structure and activity of the control ITIM based gate where the mouse CD8 stalk spacer has been replaced by an Fc domain. This gate is composed of two CARs: the first recognizes CD19, has a human CD8 stalk spacer and an ITAM containing endodomain; the second CAR recognizes CD33, has an Fc spacer and an ITIM containing endodomain. Both gates respond to CD19 single positive targets, while only the original gate is inactive in response to CD19 and CD33 double positive targets.
[0057] FIG. 24A-24D: Kinetic segregation model of CAR logic gates
[0058] Model of kinetic segregation and behaviour of AND gate, NOT AND gate and controls. CARs recognize either CD19 or CD33. The immunological synapse can be imagined between the blue line, which represents the target cell membrane and the red line, which represents the T-cell membrane. `45` is the native CD45 protein present on T-cells. `H8` is a CAR ectodomain with human CD8 stalk as the spacer. `Fc` is a CAR ectodomain with human HCH2CH3 as the spacer. `M8` is a CAR ectodomain with murine CD8 stalk as the spacer. `19` represents CD19 on the target cell surface. `33` represents CD33 on the target cell surface. The symbol `.sym.` represents an activating endodomain containing ITAMS. The symbol `.crclbar.` represents a phosphatase with slow kinetics--a `ligation on` endodomain such as one comprising of the catalytic domain of PTPN6 or an ITIM. The symbol `O` represents a phosphatase with fast kinetics--a `ligation off` endodomain such as the endodomain of CD45 or CD148. This symbol is enlarged in the figure to emphasize its potent activity.
[0059] FIG. 24A. Shows the postulated behaviour of the functional AND gate which comprises of a pair of CARs whereby the first CAR recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; and the second CAR recognizes CD33, has an Fc spacer and a CD148 endodomain;
[0060] FIG. 24B. Shows the postulated behaviour of the control AND gate. Here, the first CAR recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; and the second CAR recognizes CD33, but has a mouse CD8 stalk spacer and a CD148 endodomain;
[0061] FIG. 24C. Shows the behaviour of a functional AND NOT gate which comprises of a pair of CARs whereby the first CAR recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; and the second CAR recognizes CD33, has a mouse CD8 stalk spacer and a PTPN6 endodomain;
[0062] FIG. 24D. Shows the postulated behaviour of the control AND NOT gate which comprises of a pair of CARs whereby the first CAR recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; and the second CAR recognizes CD33, but has an Fc spacer and a PTPN6 endodomain;
[0063] In the first column, target cells are both CD19 and CD33 negative. In the second column, targets are CD19 negative and CD33 positive. In the third column, target cells are CD19 positive and CD33 negative. In the fourth column, target cells are positive for both CD19 and CD33.
[0064] FIG. 25A-25C: Design of APRIL-based CARs.
[0065] The CAR design was modified so that the scFv was replaced with a modified form of A proliferation-inducing ligand (APRIL), which interacts with interacts with BCMA, TACI and proteoglycans, to act as an antigen binding domain: APRIL was truncated so that the proteoglycan binding amino-terminus is absent. FIG. 25A. signal peptide was then attached to truncated APRIL amino-terminus to direct the protein to the cell surface. Three CARs were generated with this APRIL based binding domain: FIG. 25A. In the first CAR, the human CD8 stalk domain was used as a spacer domain. B. In the second CAR, the hinge from IgG1 was used as a spacer domain. FIG. 25C. In the third CAR, the hinge, CH2 and CH3 domains of human IgG1 modified with the pva/a mutations described by Hombach et al (2010 Gene Ther. 17:1206-1213) to reduce Fc Receptor binding was used as a spacer (henceforth referred as Fc-pvaa). In all CARs, these spacers were connected to the CD28 transmembrane domain and then to a tripartite endodomain containing a fusion of the CD28, OX40 and the CD3-Zeta endodomain (Pule et al, Molecular therapy, 2005: Volume 12; Issue 5; Pages 933-41).
[0066] FIG. 26A-26C: Annotated Amino acid sequence of the above three APRIL-CARS
[0067] FIG. 26A. Shows the annotated amino acid sequence of the CD8 stalk APRIL CAR; FIG. 26B. Shows the annotated amino acid sequence of the APRIL IgG1 hinge based CAR; FIG. 26C. Shows the annotated amino acid sequence of the APRIL Fc-pvaa based CAR.
[0068] FIG. 27A-27C: Expression and ligand binding of different APRIL based CARs
[0069] FIG. 27A. The receptors were co-expressed with a marker gene truncated CD34 in a retroviral gene vector. Expression of the marker gene on transduced cells allows confirmation of transduction. FIG. 27B. T-cells were transduced with APRIL based CARs with either the CD8 stalk spacer, IgG1 hinge or Fc spacer. To test whether these receptors could be stably expressed on the cell surface, T-cells were then stained with anti-APRIL-biotin/Streptavidin APC and anti-CD34. Flow-cytometric analysis was performed. APRIL was equally detected on the cell surface in the three CARs suggesting they are equally stably expressed. FIG. 27C. Next, the capacity of the CARs to recognize TACI and BCMA was determined. The transduced T-cells were stained with either recombinant BCMA or TACI fused to mouse IgG2a Fc fusion along with an anti-mouse secondary and anti-CD34. All three receptor formats showed binding to both BCMA and TACI. A surprising finding was that binding to BCMA seemed greater than to TACI. A further surprising finding was that although all three CARs were equally expressed, the CD8 stalk and IgG1 hinge CARs appeared better at recognizing BCMA and TACI than that with the Fc spacer.
[0070] FIG. 28A-28C: Function of the different CAR constructs.
[0071] Functional assays were performed with the three different APRIL based CARs. Normal donor peripheral blood T-cells either non-transduced (NT), or transduced to express the different CARs. Transduction was performed using equal titer supernatant. These T-cells were then CD56 depleted to remove non-specific NK activity and used as effectors. SupT1 cells either non-transduced (NT), or transduced to express BCMA or TACI were used as targets. Data shown is mean and standard deviation from 5 independent experiments. FIG. 28A. Specific killing of BCMA and TACI expressing T-cells was determined using Chromium release. FIG. 28B. Interferon-.mu. release was also determined. Targets and effectors were co-cultured at a ratio of 1:1. After 24 hours, Interferon-.mu. in the supernatant was assayed by ELISA. FIG. 28C. Proliferation/survival of CAR T-cells were also determined by counting number of CAR T-cells in the same co-culture incubated for a further 6 days. All 3 CARs direct responses against BCMA and TACI expressing targets. The responses to BCMA were greater than for TACI.
[0072] FIG. 29: AND gate functionality in primary cells.
[0073] PBMCs were isolated from blood and stimulated using PHA and IL-2. Two days later the cells were transduced on retronectin coated plates with retro virus containing the CD19:CD33 AND gate construct. On day 5 the expression level of the two CARs translated by the AND gate construct was evaluated via flow cytometry and the cells were depleted of CD56+ cells (predominantly NK cells). On day 6 the PBMCs were placed in a co-culture with target cells at a 1:2 effector to target cell ratio. On day 8 the supernatant was collected and analysed for IFN-gamma secretion via ELISA
[0074] FIG. 30A-30B: FIG. 30A. DNA alignment of the CD28 transmembrane and cytosolic domain of TCRz in the OR gate platform FIG. 30B. Protein alignment of the CD28 transmembrane and cytosolic domain of TCRz in the OR gate platform.
[0075] FIG. 31: Design rules for building logic gated CAR T-cells. OR, AND NOT and AND gated CARs are shown in cartoon format with the target cell on top, and the T-cell at the bottom with the synapse in the middle. Target cells express arbitrary target antigens A, and B. T-cells express two CARs which comprise of anti-A and anti-B recognition domains, spacers and endodomains. An OR gate requires (1) spacers simply which allow antigen recognition and CAR activation, and (2) both CARs to have activatory endodomains; An AND NOT gate requires (1) spacers which result in co-segregation of both CARs upon recognition of both antigens and (2) one CAR with an activatory endodomain, and the other whose endodomain comprises or recruits a weak phosphatase; An AND gate requires (1) spacers which result in segregation of both CARs into different parts of the immunological synapse upon recognition of both antigens and (2) one CAR with an activatory endodomain, and the other whose endodomain comprises of a potent phosphatase.
SUMMARY OF ASPECTS OF THE INVENTION
[0076] The present inventors have developed a panel of "logic-gated" chimeric antigen receptor pairs which, when expressed by a cell, such as a T cell, are capable of detecting a particular pattern of expression of at least two target antigens. If the at least two target antigens are arbitrarily denoted as antigen A and antigen B, the three possible options are as follows:
[0077] "OR GATE"--T cell triggers when either antigen A or antigen B is present on the target cell "AND GATE"--T cell triggers only when both antigens A and B are present on the target cell "AND NOT GATE"--T cell triggers if antigen A is present alone on the target cell, but not if both antigens A and B are present on the target cell
[0078] Engineered T cells expressing these CAR combinations can be tailored to be exquisitely specific for cancer cells, based on their particular expression (or lack of expression) of two or more markers.
[0079] Thus in a first aspect, the present invention provides a cell which co-expresses a first chimeric antigen receptor (CAR) and second CAR at the cell surface, each CAR comprising: [0080] (i) an antigen-binding domain; [0081] (ii) a spacer [0082] (iii) a trans-membrane domain; and [0083] (iv) an intracellular T cell signaling domain (endodomain) wherein the antigen binding domains of the first and second CARs bind to different antigens, and wherein each of the first or second CARs is an activating CAR comprising an activating intracellular T cell signaling domain.
[0084] The cell may be an immune effector cell, such as a T-cell or natural killer (NK) cell. Features mentioned herein in connection with a T cell apply equally to other immune effector cells, such as NK cells.
[0085] The spacer of the first CAR may be different to the spacer of the second CAR. such that the first and second CARs do not form heterodimers
[0086] The spacer of the first CAR may have a different length and/or size and/or configuration from the spacer of the second CAR, such that each CAR is tailored for recognition of its respective target antigen.
[0087] The spacer of the first CAR may have a different length and/or charge and/or shape and/or configuration and/or glycosylation to the spacer of the second CAR, such that when the first CAR and the second CAR bind their respective target antigens, the first CAR and second CAR become spatially separated on the T cell. Ligation of the first and second CARs to their respective antigens causes them to be compartmentalized together or separately in the immunological synapse resulting in control of activation. This may be understood when one considers the kinetic separation model of T-cell activation (see below).
[0088] The first spacer or the second spacer may comprise a CD8 stalk and the other spacer may comprise the hinge, CH2 and CH3 domain of an IgG1.
[0089] In the present invention, relating to the "OR" gate, both the first and second CAR are activating CARs. An activating CAR comprises an activating endodomain, such that it causes T cell activation upon antigen binding. Since the OR gate comprises first and second CARs which are both activating, T cell activation occurs when a target cell expresses either or both target antigens.
[0090] Either the first or the second CAR may bind CD19 and the other CAR may bind CD20. This is advantageous because some lymphomas and leukemias become CD19 negative after CD19 targeting, so it gives a "back-up" antigen, should this occur.
[0091] In a second aspect, the present invention provides a nucleic acid sequence encoding both the first and second chimeric antigen receptors (CARs) as defined in the first aspect of the invention.
[0092] The nucleic acid sequence according may have the following structure: AgB1-spacer1-TM1-endo1-coexpr-AgB2-spacer2-TM2-endo2
[0093] in which
[0094] AgB1 is a nucleic acid sequence encoding the antigen-binding domain of the first CAR;
[0095] spacer 1 is a nucleic acid sequence encoding the spacer of the first CAR;
[0096] TM1 is a a nucleic acid sequence encoding the transmembrane domain of the first CAR;
[0097] endo 1 is a nucleic acid sequence encoding the endodomain of the first CAR;
[0098] coexpr is a nucleic acid sequence allowing co-expression of two CARs (e.g. a cleavage site);
[0099] AgB2 is a nucleic acid sequence encoding the antigen-binding domain of the second CAR;
[0100] spacer 2 is a nucleic acid sequence encoding the spacer of the second CAR;
[0101] TM2 is a a nucleic acid sequence encoding the transmembrane domain of the second CAR;
[0102] endo 2 is a nucleic acid sequence encoding the endodomain of the second CAR;
[0103] which nucleic acid sequence, when expressed in a T cell, encodes a polypeptide which is cleaved at the cleavage site such that the first and second CARs are co-expressed at the T cell surface.
[0104] The nucleic acid sequence allowing co-expression of two CARs may encode a self-cleaving peptide or a sequence which allows alternative means of co-expressing two CARs such as an internal ribosome entry sequence or a 2.sup.nd promoter or other such means whereby one skilled in the art can express two proteins from the same vector.
[0105] Alternative codons may be used in regions of sequence encoding the same or similar amino acid sequences, such as the transmembrane and/or intracellular T cell signalling domain (endodomain) in order to avoid homologous recombination. An example of such "codon wobbling" is shown in FIG. 30.
[0106] In a third aspect, the present invention provides a kit which comprises [0107] (i) a first nucleic acid sequence encoding the first chimeric antigen receptor (CAR) as defined in the first aspect of the invention, which nucleic acid sequence has the following structure:
[0108] AgB1-spacer1-TM1-endo1
[0109] in which
[0110] AgB1 is a nucleic acid sequence encoding the antigen-binding domain of the first CAR;
[0111] spacer 1 is a nucleic acid sequence encoding the spacer of the first CAR;
[0112] TM1 is a a nucleic acid sequence encoding the transmembrane domain of the first CAR;
[0113] endo 1 is a nucleic acid sequence encoding the endodomain of the first CAR; and [0114] (ii) a second nucleic acid sequence encoding the second chimeric antigen receptor (CAR) as defined in the first aspect of the invention, which nucleic acid sequence has the following structure:
[0115] AgB2-spacer2-TM2-endo2
[0116] AgB2 is a nucleic acid sequence encoding the antigen-binding domain of the second CAR;
[0117] spacer 2 is a nucleic acid sequence encoding the spacer of the second CAR;
[0118] TM2 is a a nucleic acid sequence encoding the transmembrane domain of the second CAR;
[0119] endo 2 is a nucleic acid sequence encoding the endodomain of the second CAR.
[0120] In a fourth aspect, the present invention provides a kit comprising: a first vector which comprises the first nucleic acid sequence as defined above; and a second vector which comprises the first nucleic acid sequence as defined above.
[0121] The vectors may be plasmid vectors, retroviral vectors or transposon vectors. The vectors may be lentiviral vectors.
[0122] In a fifth aspect, the present invention provides a vector comprising a nucleic acid sequence according to the second aspect of the invention. The vector may be a lentiviral vector.
[0123] The vector may be a plasmid vector, a retroviral vector or a transposon vector.
[0124] In a sixth aspect, the present invention involves co-expressing more than two CARs in such a fashion that a complex pattern of more than two antigens can be recognized on the target cell.
[0125] In a seventh aspect, the present invention provides a method for making a T cell according to the first aspect of the invention, which comprises the step of introducing one or more nucleic acid sequence (s) encoding the first and second CARs; or one or more vector(s) as defined above into a T cell.
[0126] The T cell may be from a sample isolated from a patient, a related or unrelated haematopoietic transplant donor, a completely unconnected donor, from cord blood, differentiated from an embryonic cell line, differentiated from an inducible progenitor cell line, or derived from a transformed T cell line.
[0127] In an eighth aspect, the present invention provides a pharmaceutical composition comprising a plurality of T cells according to the first aspect of the invention.
[0128] In a ninth aspect, the present invention provides a method for treating and/or preventing a disease, which comprises the step of administering a pharmaceutical composition according to the eighth aspect of the invention to a subject.
[0129] The method may comprise the following steps: [0130] (i) isolation of a T cell as listed above. [0131] (ii) transduction or transfection of the T cells with one or more nucleic acid sequence(s) encoding the first and second CAR or one or more vector(s) comprising such nucleic acid sequence(s); and [0132] (iii) administering the T cells from (ii) to the subject.
[0133] The disease may be a cancer.
[0134] In a tenth aspect, the present invention provides a pharmaceutical composition according to the eighth aspect of the invention for use in treating and/or preventing a disease.
[0135] The disease may be a cancer.
[0136] In an eleventh aspect, the present invention provides use of a T cell according to the first aspect of the invention in the manufacture of a medicament for treating and/or preventing a disease.
[0137] The disease may be a cancer.
[0138] The present invention also provides a nucleic acid sequence which comprises:
[0139] a) a first nucleotide sequence encoding a first chimeric antigen receptor (CAR);
[0140] b) a second nucleotide sequence encoding a second CAR;
[0141] c) a sequence encoding a self-cleaving peptide positioned between the first and second nucleotide sequences, such that the two CARs are expressed as separate entities.
[0142] Alternative codons may be used in one or more portion(s) of the first and second nucleotide sequences in regions which encode the same or similar amino acid sequence(s).
[0143] The present invention also provides a vector and a cell comprising such a nucleic acid.
[0144] By providing two or more CARs on the surface of the T cell, it is possible to target multiple cancer markers simultaneously, providing better therapeutic efficacy for heterogeneic tumours and avoiding the problem of cancer escape.
[0145] Because the CARs are expressed on the surface of the T cell as separate molecules, this approach overcomes the spatial and accessibility issues associated with TanCARs. T-cell activation efficiency is also improved. As each CAR has its own spacer, it is possible to tailor the spacer and therefore the distance that the binding domain projects from the T cell surface and its flexibility etc to the particular target antigen. This choice is unfettered by the design considerations associated with TanCARs, i.e. that one CAR needs to be juxta-posed to the T cell membrane and one CAR needs to be distal, positioned in tandem to the first CAR.
[0146] By providing a single nucleic acid which encodes the two CARs separated by a cleavage site, it is possible to engineer T cells to co-express the two CARs using a simple single transduction procedure. A double transfection procedure could be used with CAR-encoding sequences in separate constructs, but this would be more complex and expensive and requires more integration sites for the nucleic acids. A double transfection procedure would also be associated with uncertainty as to whether both CAR-encoding nucleic acids had been transduced and expressed effectively. This is especially true for a multiple CAR approach where three or more CARs are introduced to the cell.
[0147] The two or more CARs will have portions of high homology, for example the transmembrane and/or intracellular signalling domains are likely to be highly homologous. If the same or similar linkers are use for the two CARs, then they will also be highly homologous. This would suggest that an approach where both CARs are provided on a single nucleic acid sequence would be inappropriate, because of the likelihood of homologous recombination between the sequences. However, the present inventors have found that by "codon wobbling" the portions of sequence encoding areas of high homology, it is possible to express two CARs from a single construct with high efficiency. Codon wobbling involves using alternative codons in regions of sequence encoding the same or similar amino acid sequences.
FURTHER ASPECTS OF THE INVENTION
[0148] The present invention also relates to the aspects listed in the following numbered paragraphs:
[0149] 1. A T cell which co-expresses a first chimeric antigen receptor (CAR) and second CAR at the cell surface, each CAR comprising: [0150] (i) an antigen-binding domain; [0151] (ii) a spacer [0152] (iii) a trans-membrane domain; and [0153] (iv) an endodomain
[0154] wherein the antigen binding domains of the first and second CARs bind to different antigens, wherein the spacer of the first CAR is different to the spacer of the second CAR and wherein one of the first or second CARs is an activating CAR comprising an activating endodomain and the other CAR is either an activating CAR comprising an activating endodomain or an inhibitory CAR comprising a ligation-on or ligation-off inhibitory endodomain.
[0155] 2. A T cell according to paragraph 1, wherein the spacer of the first CAR has a different length and/or charge and/or size and/or configuration and/or glycosylation of the spacer of the second CAR, such that when the first CAR and the second CAR bind their respective target antigens, the first CAR and second CAR become spatially separated on the T cell membrane.
[0156] 3. A T cell according to paragraph 2, wherein either the first spacer or the second spacer comprises a CD8 stalk and the other spacer comprises the hinge, CH2 and CH3 domain of IgG1.
[0157] 4. A T cell according to paragraph 1, wherein both the first and second CARs are activating CARs.
[0158] 5. A T cell according to paragraph 4, wherein one CAR binds CD19 and the other CAR binds CD20.
[0159] 6. A T cell according to paragraph 2 or 3, wherein one of the first or second CARs is an activating CAR comprising an activating endodomain, and the other CAR is an inhibitory CAR comprising a ligation-off inhibitory endodomain, which inhibitory CAR inhibits T-cell activation by the activating CAR in the absence of inhibitory CAR ligation, but does not significantly inhibit T-cell activation by the activating CAR when the inhibitory CAR is ligated.
[0160] 7. A T cell according to paragraph 6, wherein the inhibitory endodomain comprises all or part of the endodomain from CD148 or CD45.
[0161] 8. A T cell according to paragraph 6 or 7, wherein the antigen-binding domain of the first CAR binds CD5 and the antigen-binding domain of the second CAR binds CD19.
[0162] 9. A T cell according to paragraph 1 wherein the first and second spacers are sufficiently different so as to prevent cross-pairing of the first and second CARs but are sufficiently similar to result in co-localisation of the first and second CARs following ligation.
[0163] 10. A T cell according to paragraph 9, wherein one of the first or second CARs in an activating CAR comprising an activating endodomain, and the other CAR is an inhibitory CAR comprising a ligation-on inhibitory endodomain, which inhibitory CAR does not significantly inhibit T-cell activation by the activating CAR in the absence of inhibitory CAR ligation, but inhibits T-cell activation by the activating CAR when the inhibitory CAR is ligated.
[0164] 11. A T cell according to paragraph 10, wherein the ligation-on inhibitory endodomain comprises at least part of a phosphatase.
[0165] 12. A T cell according to paragraph 11, wherein the ligation-on inhibitory endodomain comprises all or part of PTPN6.
[0166] 13. A T cell according to paragraph 10, wherein the ligation-on inhibitory endodomain comprises at least one ITIM domain.
[0167] 14. A T cell according to paragraph 13, wherein activity of the ligation-on inhibitory endodomain is enhanced by co-expression of a PTPN6-CD45 or -CD148 fusion protein.
[0168] 15. A T cell according to any of paragraphs 10 to 14, wherein the CAR comprising the activating endodomain comprises an antigen-binding domain which binds CD33 and the CAR which comprises the ligation-on inhibitory endodomain comprises an antigen-binding domain which binds CD34.
[0169] 16. A T cell which comprises more than two CARs as defined in the preceding paragraphs such that it is specifically stimulated by a cell, such as a T cell, bearing a distinct pattern of more than two antigens.
[0170] 17. A nucleic acid sequence encoding both the first and second chimeric antigen receptors (CARs) as defined in any of paragraphs 1 to 16.
[0171] 18. A nucleic acid sequence according to paragraph 17, which has the following structure:
[0172] AgB1-spacer1-TM1-endo1-coexpr-AbB2-spacer2-TM2-endo2 in which
[0173] AgB1 is a nucleic acid sequence encoding the antigen-binding domain of the first CAR;
[0174] spacer 1 is a nucleic acid sequence encoding the spacer of the first CAR;
[0175] TM1 is a a nucleic acid sequence encoding the transmembrane domain of the first CAR;
[0176] endo 1 is a nucleic acid sequence encoding the endodomain of the first CAR;
[0177] coexpr is a nucleic acid sequence enabling co-expression of both CARs
[0178] AgB2 is a nucleic acid sequence encoding the antigen-binding domain of the second CAR;
[0179] spacer 2 is a nucleic acid sequence encoding the spacer of the second CAR;
[0180] TM2 is a a nucleic acid sequence encoding the transmembrane domain of the second CAR;
[0181] endo 2 is a nucleic acid sequence encoding the endodomain of the second CAR;
[0182] which nucleic acid sequence, when expressed in a T cell, encodes a polypeptide which is cleaved at the cleavage site such that the first and second CARs are co-expressed at the T cell surface.
[0183] 19. A nucleic acid sequence according to paragraph 18, wherein coexpr encodes a sequence comprising a self-cleaving peptide.
[0184] 20. A nucleic acid sequence according to paragraph 18 or 19, wherein alternative codons are used in regions of sequence encoding the same or similar amino acid sequences, in order to avoid homologous recombination.
[0185] 21. A kit which comprises [0186] (i) a first nucleic acid sequence encoding the first chimeric antigen receptor (CAR) as defined in any of paragraphs 1 to 16, which nucleic acid sequence has the following structure:
[0187] AgB1-spacer1-TM1-endo1
[0188] in which
[0189] AgB1 is a nucleic acid sequence encoding the antigen-binding domain of the first CAR;
[0190] spacer 1 is a nucleic acid sequence encoding the spacer of the first CAR;
[0191] TM1 is a nucleic acid sequence encoding the transmembrane domain of the first CAR;
[0192] endo 1 is a nucleic acid sequence encoding the endodomain of the first CAR; and [0193] (ii) a second nucleic acid sequence encoding the second chimeric antigen receptor (CAR) as defined in any of paragraphs 1 to 16, which nucleic acid sequence has the following structure:
[0194] AgB2-spacer2-TM2-endo2
[0195] AgB2 is a nucleic acid sequence encoding the antigen-binding domain of the second CAR;
[0196] spacer 2 is a nucleic acid sequence encoding the spacer of the second CAR;
[0197] TM2 is a a nucleic acid sequence encoding the transmembrane domain of the second CAR;
[0198] endo 2 is a nucleic acid sequence encoding the endodomain of the second CAR.
[0199] 22. A kit comprising: a first vector which comprises the first nucleic acid sequence as defined in paragraph 21; and a second vector which comprises the first nucleic acid sequence as defined in paragraph 21.
[0200] 23. A kit according to paragraph 22, wherein the vectors are integrating viral vectors or transposons.
[0201] 24. A vector comprising a nucleic acid sequence according to any of paragraphs 17 to 20.
[0202] 25. A retroviral vector or a lentiviral vector or a transposon according to paragraph 24.
[0203] 26. A method for making a T cell according to any of paragraphs 1 to 16, which comprises the step of introducing: a nucleic acid sequence according to any of paragraphs 17 to 20; a first nucleic acid sequence and a second nucleic acid sequence as defined in paragraph 21; and/or a first vector and a second vector as defined in paragraph 22 or a vector according to paragraph 24 or 25, into a T cell.
[0204] 27. A method according to paragraph 24, wherein the T cell is from a sample isolated from a subject.
[0205] 28. A pharmaceutical composition comprising a plurality of T cells according to any of paragraphs 1 to 16.
[0206] 29. A method for treating and/or preventing a disease, which comprises the step of administering a pharmaceutical composition according to paragraph 28 to a subject.
[0207] 30. A method according to paragraph 29, which comprises the following steps: [0208] (i) isolation of a T cell-containing sample from a subject; [0209] (ii) transduction or transfection of the T cells with: a nucleic acid sequence according to any of paragraphs 17 to 20; a first nucleic acid sequence and a second nucleic acid sequence as defined in paragraph 21; a first vector and a second vector as defined in paragraph 22 or 23 or a vector according to paragraph 24 or 25; and [0210] (iii) administering the T cells from (ii) to a the subject.
[0211] 31. A method according to paragraph 29 or 30, wherein the disease is a cancer.
[0212] 32. A pharmaceutical composition according to paragraph 28 for use in treating and/or preventing a disease.
[0213] 33. The use of a T cell according to any of paragraphs 1 to 16 in the manufacture of a medicament for treating and/or preventing a disease.
DETAILED DESCRIPTION
[0214] Chimeric Antigen Receptors (Cars)
[0215] CARs, which are shown schematically in FIG. 1, are chimeric type I trans-membrane proteins which connect an extracellular antigen-recognizing domain (binder) to an intracellular signalling domain (endodomain). The binder is typically a single-chain variable fragment (scFv) derived from a monoclonal antibody (mAb), but it can be based on other formats which comprise an antibody-like antigen binding site. A spacer domain is usually necessary to isolate the binder from the membrane and to allow it a suitable orientation. A common spacer domain used is the Fc of IgG1. More compact spacers can suffice e.g. the stalk from CD8.alpha. and even just the IgG1 hinge alone, depending on the antigen. A trans-membrane domain anchors the protein in the cell membrane and connects the spacer to the endodomain.
[0216] Early CAR designs had endodomains derived from the intracellular parts of either the .gamma. chain of the Fc.epsilon.R1 or CD3.zeta.. Consequently, these first generation receptors transmitted immunological signal 1, which was sufficient to trigger T-cell killing of cognate target cells but failed to fully activate the T-cell to proliferate and survive. To overcome this limitation, compound endodomains have been constructed: fusion of the intracellular part of a T-cell co-stimulatory molecule to that of CD3.zeta. results in second generation receptors which can transmit an activating and co-stimulatory signal simultaneously after antigen recognition. The co-stimulatory domain most commonly used is that of CD28. This supplies the most potent co-stimulatory signal--namely immunological signal 2, which triggers T-cell proliferation. Some receptors have also been described which include TNF receptor family endodomains, such as the closely related OX40 and 41BB which transmit survival signals. Even more potent third generation CARs have now been described which have endodomains capable of transmitting activation, proliferation and survival signals.
[0217] CAR-encoding nucleic acids may be transferred to T cells using, for example, retroviral vectors. Lentiviral vectors may be employed. In this way, a large number of cancer-specific T cells can be generated for adoptive cell transfer. When the CAR binds the target-antigen, this results in the transmission of an activating signal to the T-cell it is expressed on. Thus the CAR directs the specificity and cytotoxicity of the T cell towards tumour cells expressing the targeted antigen.
[0218] The first aspect of the invention relates to a T-cell which co-expresses a first CAR and a second CAR such that a T-cell can recognize a desired pattern of expression on target cells in the manner of a logic gate as detailed in the truth tables: table 1, 2 and 3.
[0219] Both the first and second (and optionally subsequent) CARs comprise:
[0220] (i) an antigen-binding domain;
[0221] (ii) a spacer;
[0222] (iii) a transmembrane domain; and
[0223] (iii) an intracellular domain.
TABLE-US-00001 TABLE 1 Truth Table for CAR OR GATE Antigen A Antigen B Response Absent Absent No activation Absent Present Activation Present Absent Activation Present Present Activation
TABLE-US-00002 TABLE 2 Truth Table for CAR AND GATE Antigen A Antigen B Response Absent Absent No activation Absent Present No Activation Present Absent No Activation Present Present Activation
TABLE-US-00003 TABLE 3 Truth Table for CAR AND NOT GATE Antigen A Antigen B Response Absent Absent No activation Absent Present No Activation Present Absent Activation Present Present No Activation
[0224] The first and second CAR of the T cell of the present invention may be produced as a polypeptide comprising both CARs, together with a cleavage site.
[0225] SEQ ID No. 1 to 5 give examples of such polypeptides, which each comprise two CARs. The CAR may therefore comprise one or other part of the following amino acid sequences, which corresponds to a single CAR.
[0226] SEQ ID No 1 is a CAR OR gate which recognizes CD19 OR CD33
[0227] SEQ ID No 2 Is a CAR AND gate which recognizes CD19 AND CD33 using a CD148 phosphatase
[0228] SEQ ID No 3 Is an alternative implementation of the CAR AND GATE which recognizes CD19 AND CD33 which uses a CD45 phosphatase
[0229] SEQ ID No 4 Is a CAR AND NOT GATE which recognizes CD19 AND NOT CD33 based on PTPN6 phosphatase
[0230] SEQ ID No 5 Is an alternative implementation of the CAR AND NOT gate which recognizes CD19 AND NOT CD33 and is based on an ITIM containing endodomain from LAIR1
[0231] SEQ ID No 6. Is a further alternative implementation of the CAR AND NOT gate which recognizes CD19 AND NOT CD33 and recruits a PTPN6-CD148 fusion protein to an ITIM containing endodomain.
TABLE-US-00004 SEQ ID No. 1 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDI SKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLE QEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSGGGGSGGGGSGGGGS EVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGV IWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYY YGGSYAMDYWGQGTSVTVSSDPTTTPAPRPPTPAPTIASQPLSLRPEACR PAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAFIIFWVRRVKF SRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNP QEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPRRAEGRGSLLTCGDVEENPGPMAVPTQVLGLLLLWLTDARCDI QMTQSPSSLSASVGDRVTITCRASEDIYFNLVWYQQKPGKAPKLLIYDTN RLADGVPSRFSGSGSGTQYTLTISSLQPEDFATYYCQHYKNYPLTFGQGT KLEIKRSGGGGSGGGGSGGGGSGGGGSRSEVQLVESGGGLVQPGGSLRLS CAASGFTLSNYGMHWIRQAPGKGLEWVSSISLNGGSTYYRDSVKGRFTIS RDNAKSTLYLQMNSLRAEDTAVYYCAAQDAYTGGYFDYWGQGTLVTVSSM DPAEPKSPDKTHTCPPCPAPPVAGPSVFLFPPKPKDTLMIARTPEVTCVV VDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVD KSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGKKDPKFWVLVVVGGVLA CYSLLVTVAFIIFWVRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYD VLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRG KGHDGLYQGLSTATKDTYDALHMQALPPR SEQ ID No. 2 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDI SKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLE QEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSGGGGSGGGGSGGGGS EVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGV IWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYY YGGSYAMDYWGQGTSVTVSSDPTTTPAPRPPTPAPTIASQPLSLRPEACR PAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAFIIFWVRRVKF SRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNP QEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPRRAEGRGSLLTCGDVEENPGPMAVPTQVLGLLLLWLTDARCDI QMTQSPSSLSASVGDRVTITCRASEDIYFNLVWYQQKPGKAPKLLIYDTN RLADGVPSRFSGSGSGTQYTLTISSLQPEDFATYYCQHYKNYPLTFGQGT KLEIKRSGGGGSGGGGSGGGGSGGGGSRSEVQLVESGGGLVQPGGSLRLS CAASGFTLSNYGMHWIRQAPGKGLEWVSSISLNGGSTYYRDSVKGRFTIS RDNAKSTLYLQMNSLRAEDTAVYYCAAQDAYTGGYFDYWGQGTLVTVSSM DPAEPKSPDKTHTCPPCPAPPVAGPSVFLFPPKPKDTLMIARTPEVTCVV VDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVD KSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGKKDPKAVFGCIFGALVI VTVGGFIFWRKKRKDAKNNEVSFSQIKPKKSKLIRVENFEAYFKKQQADS NCGFAEEYEDLKLVGISQPKYAAELAENRGKNRYNNVLPYDISRVKLSVQ THSTDDYINANYMPGYHSKKDFIATQGPLPNTLKDFWRMVWEKNVYAIIM LTKCVEQGRTKCEEYWPSKQAQDYGDITVAMTSEIVLPEWTIRDFTVKNI QTSESHPLRQFHFTSWPDHGVPDTTDLLINFRYLVRDYMKQSPPESPILV HCSAGVGRTGTFIAIDRLIYQIENENTVDVYGIVYDLRMHRPLMVQTEDQ YVFLNQCVLDIVRSQKDSKVDLIYQNTTAMTIYENLAPVTTFGKTNGYIA SEQ ID No. 3 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDI SKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLE QEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSGGGGSGGGGSGGGGS EVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGV IWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYY YGGSYAMDYWGQGTSVTVSSDPTTTPAPRPPTPAPTIASQPLSLRPEACR PAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAFIIFWVRRVKF SRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNP QEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPRRAEGRGSLLTCGDVEENPGPMAVPTQVLGLLLLWLTDARCDI QMTQSPSSLSASVGDRVTITCRASEDIYFNLVWYQQKPGKAPKLLIYDTN RLADGVPSRFSGSGSGTQYTLTISSLQPEDFATYYCQHYKNYPLTFGQGT KLEIKRSGGGGSGGGGSGGGGSGGGGSRSEVQLVESGGGLVQPGGSLRLS CAASGFTLSNYGMHWIRQAPGKGLEWVSSISLNGGSTYYRDSVKGRFTIS RDNAKSTLYLQMNSLRAEDTAVYYCAAQDAYTGGYFDYWGQGTLVTVSSM DPAEPKSPDKTHTCPPCPAPPVAGPSVFLFPPKPKDTLMIARTPEVTCVV VDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVD KSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGKKDPKALIAFLAFLIIV TSIALLVVLYKIYDLHKKRSCNLDEQQELVERDDEKQLMNVEPIHADILL ETYKRKIADEGRLFLAEFQSIPRVFSKFPIKEARKPFNQNKNRYVDILPY DYNRVELSEINGDAGSNYINASYIDGFKEPRKYIAAQGPRDETVDDFWRM IWEQKATVIVMVTRCEEGNRNKCAEYWPSMEEGTRAFGDVVVKINQHKRC PDYIIQKLNIVNKKEKATGREVTHIQFTSWPDHGVPEDPHLLLKLRRRVN AFSNFFSGPIVVHCSAGVGRTGTYIGIDAMLEGLEAENKVDVYGYVVKLR RQRCLMVQVEAQYILIHQALVEYNQFGETEVNLSELHPYLHNMKKRDPPS EPSPLEAEFQRLPSYRSWRTQHIGNQEENKSKNRNSNVIPYDYNRVPLKH ELEMSKESEHDSDESSDDDSDSEEPSKYINASFIMSYWKPEVMIAAQGPL KETIGDFWQMIFQRKVKVIVMLTELKHGDQEICAQYWGEGKQTYGDIEVD LKDTDKSSTYTLRVFELRHSKRKDSRTVYQYQYTNWSVEQLPAEPKELIS MIQVVKQKLPQKNSSEGNKHHKSTPLLIHCRDGSQQTGIFCALLNLLESA ETEEVVDIFQVVKALRKARPGMVSTFEQYQFLYDVIASTYPAQNGQVKKN NHQEDKIEFDNEVDKVKQDANCVNPLGAPEKLPEAKEQAEGSEPTSGTEG PEHSVNGPASPALNQGS SEQ ID No. 4 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDI SKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLE QEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSGGGGSGGGGSGGGGS EVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGV IWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYY YGGSYAMDYWGQGTSVTVSSDPTTTPAPRPPTPAPTIASQPLSLRPEACR PAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAFIIFWVRRVKF SRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNP QEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPRRAEGRGSLLTCGDVEENPGPMAVPTQVLGLLLLWLTDARCDI QMTQSPSSLSASVGDRVTITCRASEDIYFNLVWYQQKPGKAPKLLIYDTN RLADGVPSRFSGSGSGTQYTLTISSLQPEDFATYYCQHYKNYPLTFGQGT KLEIKRSGGGGSGGGGSGGGGSGGGGSRSEVQLVESGGGLVQPGGSLRLS CAASGFTLSNYGMHWIRQAPGKGLEWVSSISLNGGSTYYRDSVKGRFTIS RDNAKSTLYLQMNSLRAEDTAVYYCAAQDAYTGGYFDYWGQGTLVTVSSM DPATTTKPVLRTPSPVHPTGTSQPQRPEDCRPRGSVKGTGLDFACDIYWA PLAGICVALLLSLIITLICYHRSRKRVCKSGGGSFWEEFESLQKQEVKNL HQRLEGQRPENKGKNRYKNILPFDHSRVILQGRDSNIPGSDYINANYIKN QLLGPDENAKTYIASQGCLEATVNDFWQMAWQENSRVIVMTTREVEKGRN KCVPYWPEVGMQRAYGPYSVTNCGEHDTTEYKLRTLQVSPLDNGDLIREI WHYQYLSWPDHGVPSEPGGVLSFLDQINQRQESLPHAGPIIVHCSAGIGR TGTIIVIDMLMENISTKGLDCDIDIQKTIQMVRAQRSGMVQTEAQYKFIY VAIAQFIETTKKKL SEQ ID No. 5 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDI SKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLE QEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSGGGGSGGGGSGGGGS EVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGV IWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYY YGGSYAMDYWGQGTSVTVSSDPTTTPAPRPPTPAPTIASQPLSLRPEACR PAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAFIIFWVRRVKF SRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNP QEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPRRAEGRGSLLTCGDVEENPGPMAVPTQVLGLLLLWLTDARCDI QMTQSPSSLSASVGDRVTITCRASEDIYFNLVWYQQKPGKAPKLLIYDTN RLADGVPSRFSGSGSGTQYTLTISSLQPEDFATYYCQHYKNYPLTFGQGT KLEIKRSGGGGSGGGGSGGGGSGGGGSRSEVQLVESGGGLVQPGGSLRLS CAASGFTLSNYGMHWIRQAPGKGLEWVSSISLNGGSTYYRDSVKGRFTIS RDNAKSTLYLQMNSLRAEDTAVYYCAAQDAYTGGYFDYWGQGTLVTVSSM
DPATTTKPVLRTPSPVHPTGTSQPQRPEDCRPRGSVKGTGLDFACDILIG VSVVFLFCLLLLVLFCLHRQNQIKQGPPRSKDEEQKPQQRPDLAVDVLER TADKATVNGLPEKDRETDTSALAAGSSQEVTYAQLDHWALTQRTARAVSP QSTKPMAESITYAAVARH SEQ ID No. 6 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDI SKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLE QEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSGGGGSGGGGSGGGGS EVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGV IWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYY YGGSYAMDYWGQGTSVTVSSDPTTTPAPRPPTPAPTIASQPLSLRPEACR PAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAFIIFWVRRVKF SRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNP QEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDAL HMQALPPRRAEGRGSLLTCGDVEENPGPMAVPTQVLGLLLLWLTDARCDI QMTQSPSSLSASVGDRVTITCRASEDIYFNLVWYQQKPGKAPKLLIYDTN RLADGVPSRFSGSGSGTQYTLTISSLQPEDFATYYCQHYKNYPLTFGQGT KLEIKRSGGGGSGGGGSGGGGSGGGGSRSEVQLVESGGGLVQPGGSLRLS CAASGFTLSNYGMHWIRQAPGKGLEWVSSISLNGGSTYYRDSVKGRFTIS RDNAKSTLYLQMNSLRAEDTAVYYCAAQDAYTGGYFDYWGQGTLVTVSSM DPATTTKPVLRTPSPVHPTGTSQPQRPEDCRPRGSVKGTGLDFACDILIG VSVVFLFCLLLLVLFCLHRQNQIKQGPPRSKDEEQKPQQRPDLAVDVLER TADKATVNGLPEKDRETDTSALAAGSSQEVTYAQLDHWALTQRTARAVSP QSTKPMAESITYAAVARHRAEGRGSLLTCGDVEENPGPWYHGHMSGGQAE TLLQAKGEPWTFLVRESLSQPGDFVLSVLSDQPKAGPGSPLRVTHIKVMC EGGRYTVGGLETFDSLTDLVEHFKKTGIEEASGAFVYLRQPYSGGGGSFE AYFKKQQADSNCGFAEEYEDLKLVGISQPKYAAELAENRGKNRYNNVLPY DISRVKLSVQTHSTDDYINANYMPGYHSKKDFIATQGPLPNTLKDFWRMV WEKNVYAIIMLTKCVEQGRTKCEEYWPSKQAQDYGDITVAMTSEIVLPEW TIRDFTVKNIQTSESHPLRQFHFTSWPDHGVPDTTDLLINFRYLVRDYMK QSPPESPILVHCSAGVGRTGTFIAIDRLIYQIENENTVDVYGIVYDLRMH RPLMVQTEDQYVFLNQCVLDIVRSQKDSKVDLIYQNTTAMTIYENLAPVT TFGKTNGYIASGS
[0232] The CAR may comprise a variant of the CAR-encoding part of the sequence shown as SEQ ID No. 1, 2, 3, 4, 5 or 6 having at least 80, 85, 90, 95, 98 or 99% sequence identity, provided that the variant sequence is a CAR having the required properties.
[0233] Methods of sequence alignment are well known in the art and are accomplished using suitable alignment programs. The % sequence identity refers to the percentage of amino acid or nucleotide residues that are identical in the two sequences when they are optimally aligned. Nucleotide and protein sequence homology or identity may be determined using standard algorithms such as a BLAST program (Basic Local Alignment Search Tool at the National Center for Biotechnology Information) using default parameters, which is publicly available at http://blast.ncbi.nlm.nih.gov. Other algorithms for determining sequence identity or homology include: LALIGN (http://www.ebi.ac.uk/Tools/psa/lalign/ and http://www.ebi.ac.uk/Tools/psa/lalign/nucleotide.html), AMAS (Analysis of Multiply Aligned Sequences, at http://www.compbio.dundee.ac.uk/Software/Amas/amas.html), FASTA (http://www.ebi.ac.uk/Tools/sss/fasta/), Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo/), SIM (http://web.expasy.org/sim/), and EMBOSS Needle (http://www.ebi.ac.uk/Tools/psa/emboss_needle/nucleotide.html).
[0234] Car Logical or Gate
[0235] In this embodiment, the antigen binding domains of the first and second CARs of the present invention bind to different antigens and both CARs comprise an activating endodomain. The spacer domains may be the same, or sufficiently different to prevent cross-pairing of the two different receptors. A T cell can hence be engineered to activate upon recognition of either or both antigens. This is useful in the field of oncology as indicated by the Goldie-Coldman hypothesis: sole targeting of a single antigen may result in tumour escape by modulation of said antigen due to the high mutation rate inherent in most cancers. By simultaneously targeting two antigens, the probably of such escape is exponentially reduced.
[0236] Various tumour associated antigens are known as shown in the following Table 4. For a given disease, the first CAR and second CAR may bind to two different TAAs associated with that disease. In this way, tumour escape by modulating a single antigen is prevented, since a second antigen is also targeted. For example, when targeting a B-cell malignancy, both CD19 and CD20 can be simultaneously targeted. In this embodiment, it is important that the two CARs do not heterodimerize.
TABLE-US-00005 TABLE 4 Cancer type TAA Diffuse Large B-cell Lymphoma CD19, CD20 Breast cancer ErbB2, MUC1 AML CD13, CD33 Neuroblastoma GD2, NCAM B-CLL CD19, CD52 Colorectal cancer Folate binding protein, CA-125
[0237] Kinetic Segregation Model
[0238] Subsequent pairing of CARs to generate the AND gate and the AND NOT gate are based on the kinetic segregation model (KS) of T-cell activation. This is a functional model, backed by experimental data, which explains how antigen recognition by a T-cell receptor is converted into down-stream activation signals. Briefly: at the ground state, the signalling components on the T-cell membrane are in dynamic homeostasis whereby dephosphorylated ITAMs are favoured over phosphorylated ITAMs. This is due to greater activity of the transmembrane CD45/CD148 phosphatases over membrane-tethered kinases such as Ick. When a T-cell engages a target cell through a T-cell receptor (or CAR) recognition of cognate antigen, tight immunological synapses form. This close juxtapositioning of the T-cell and target membranes excludes CD45/CD148 due to their large ectodomains which cannot fit into the synapse. Segregation of a high concentration of T-cell receptor associated ITAMs and kinases in the synapse, in the absence of phosphatases, leads to a state whereby phosphorylated ITAMs are favoured. ZAP70 recognizes a threshold of phosphorylated ITAMs and propagates a T-cell activation signal. This advanced understanding of T-cell activation is exploited by the present invention. In particular, the invention is based on this understanding of how ectodomains of different length and/or bulk and/or charge and/or configuration and/or glycosylation result in differential segregation upon synapse formation.
[0239] The Car Logical and Gate
[0240] In this embodiment, one CAR comprises an activating endodomain and one CAR comprises an inhibitory endodomain whereby the inhibitory CAR constitutively inhibits the first activating CAR, but upon recognition of its cognate antigen releases its inhibition of the activating CAR. In this manner, a T-cell can be engineered to trigger only if a target cell expresses both cognate antigens. This behaviour is achieved by the activating CAR comprising an activating endodomain containing ITAM domains for example the endodomain of CD3 Zeta, and the inhibitory CAR comprising the endodomain from a phosphatase able to dephosphorylate an ITAM (e.g. CD45 or CD148). Crucially, the spacer domains of both CARs are significantly different in size and/or shape and/or charge etc. When only the activating CAR is ligated, the inhibitory CAR is in solution on the T-cell surface and can diffuse in and out of the synapse inhibiting the activating CAR. When both CARs are ligated, due to differences in spacer properties, the activating and inhibiting CAR are differentially segregated allowing the activating CAR to trigger T-cell activation unhindered by the inhibiting CAR.
[0241] This is of considerable utility in the field of cancer therapy. Currently, immunotherapies typically target a single antigen. Most cancers cannot be differentiated from normal tissues on the basis of a single antigen. Hence, considerable "on-target off-tumour" toxicity occurs whereby normal tissues are damaged by the therapy. For instance, whilst targeting CD20 to treat B-cell lymphomas with Rituximab, the entire normal B-cell compartment is depleted. For instance, whilst targeting CD52 to treat chronic lymphocytic leukaemia, the entire lymphoid compartment is depleted. For instance, whilst targeting CD33 to treat acute myeloid leukaemia, the entire myeloid compartment is damaged etc. By restricting activity to a pair of antigens, much more refined targeting, and hence less toxic therapy can be developed. A practical example is targeting of CLL which expresses both CD5 and CD19. Only a small proportion of normal B-cells express both antigens, so the off-target toxicity of targeting both antigens with a logical AND gate is substantially less than targeting each antigen individually.
[0242] The design of the present invention is a considerable improvement on previous implementation as described by Wilkie et al. ((2012). J. Clin. Immunol. 32, 1059-1070) and then tested in vivo (Kloss et al (2013) Nat. Biotechnol. 31, 71-75). In this implementation, the first CAR comprises of an activating endodomain, and the second a co-stimulatory domain. This way, a T-cell only receives an activating and co-stimulatory signal when both antigens are present. However, the T-cell still will activate in the sole presence of the first antigen resulting in the potential for off-target toxicity. Further, the implementation of the present invention allows for multiple compound linked gates whereby a cell can interpret a complex pattern of antigens.
TABLE-US-00006 TABLE 5 Cancer Type Antigens Chronic Lymphocytic Leukaemia CD5, CD19 Neuroblastoma ALK, GD2 Glioma EGFR, Vimentin Multiple myeloma BCMA, CD138 Renal Cell Carcinoma Carbonic anhydrase IX, G250 T-ALL CD2, N-Cadherin Prostate Cancer PSMA, hepsin (or others)
[0243] The Car Logical and not Gate
[0244] In this embodiment, one CAR comprises an activating endodomain and one CAR comprises an inhibitory endodomain such that this inhibitory CAR is only active when it recognizes its cognate antigen. Hence a T-cell engineered in this manner is activated in response to the sole presence of the first antigen but is not activated when both antigens are present. This invention is implemented by inhibitory CARs with a spacer that co-localise with the first CAR but either the phosphatase activity of the inhibitory CAR should not be so potent that it inhibits in solution, or the inhibitory endodomain in fact recruits a phosphatase solely when the inhibitory CAR recognizes its cognate target. Such endodomains are termed "ligation-on" or semi-inhibitory herein.
[0245] This invention is of use in refining targeting when a tumour can be distinguished from normal tissue by the presence of tumour associated antigen and the loss of an antigen expressed on normal tissue. The AND NOT gate is of considerable utility in the field of oncology as it allows targeting of an antigen which is expressed by a normal cell, which normal cell also expresses the antigen recognised by the CAR comprising the activating endodomain. An example of such an antigen is CD33 which is expressed by normal stem cells and acute myeloid leukaemia (AML) cells. CD34 is expressed on stem cells but not typically expressed on AML cells. A T-cell recognizing CD33 AND NOT CD34 would result in destruction of leukaemia cells but sparing of normal stem cells.
[0246] Potential antigen pairs for use with AND NOT gates are shown in Table 6.
TABLE-US-00007 TABLE 6 Antigen expressed Normal cell which by normal cell Disease TAA expresses TAA but not cancer cell AML CD33 stem cells CD34 Myeloma BCMA Dendritic cells CD1c B-CLL CD160 Natural Killer cells CD56 Prostate PSMA Neural Tissue NCAM cancer Bowel cancer A33 Normal bowel HLA class I epithelium
[0247] Compound Gates
[0248] The kinetic segregation model with the above components allows compound gates to be made e.g. a T-cell which triggers in response to patterns of more than two target antigens. For example, it is possible to make a T cell which only triggers when three antigens are present (A AND B AND C). Here, a cell expresses three CARs, each recognizing antigens A, B and C. One CAR is excitatory and two are inhibitory, which each CAR having spacer domains which result in differential segregation. Only when all three are ligated, will the T-cell activate. A further example: (A OR B) AND C: here, CARs recognizing antigens A and B are activating and have spacers which co-localise, while CAR recognizing antigen C is inhibitory and has a spacer which results in different co-segregation. A further example (A AND NOT B) AND C: Here CAR against antigen A has an activating endodomain and co-localises with CAR against antigen B which has a conditionally inhibiting endodomain. CAR against antigen C has a spacer who segregates differently from A or B and is inhibitory. In fact, ever more complex boolean logic can be programmed with these simple components of the invention with any number of CARs and spacers.
[0249] Signal Peptide
[0250] The CARs of the T cell of the present invention may comprise a signal peptide so that when the CAR is expressed inside a cell, such as a T-cell, the nascent protein is directed to the endoplasmic reticulum and subsequently to the cell surface, where it is expressed.
[0251] The core of the signal peptide may contain a long stretch of hydrophobic amino acids that has a tendency to form a single alpha-helix. The signal peptide may begin with a short positively charged stretch of amino acids, which helps to enforce proper topology of the polypeptide during translocation. At the end of the signal peptide there is typically a stretch of amino acids that is recognized and cleaved by signal peptidase. Signal peptidase may cleave either during or after completion of translocation to generate a free signal peptide and a mature protein. The free signal peptides are then digested by specific proteases.
[0252] The signal peptide may be at the amino terminus of the molecule.
[0253] The signal peptide may comprise the SEQ ID No. 7, 8 or 9 or a variant thereof having 5, 4, 3, 2 or 1 amino acid mutations (insertions, substitutions or additions) provided that the signal peptide still functions to cause cell surface expression of the CAR.
TABLE-US-00008 SEQ ID No. 7: MGTSLLCWMALCLLGADHADG
[0254] The signal peptide of SEQ ID No. 7 is compact and highly efficient. It is predicted to give about 95% cleavage after the terminal glycine, giving efficient removal by signal peptidase.
TABLE-US-00009 SEQ ID No. 8: MSLPVTALLLPLALLLHAARP
[0255] The signal peptide of SEQ ID No. 8 is derived from IgG1.
TABLE-US-00010 SEQ ID No. 9: MAVPTQVLGLLLLWLTDARC
[0256] The signal peptide of SEQ ID No. 9 is derived from CD8.
[0257] The signal peptide for the first CAR may have a different sequence from the signal peptide of the second CAR (and from the 3.sup.rd CAR and 4.sup.th CAR etc).
[0258] Antigen Binding Domain
[0259] The antigen binding domain is the portion of the CAR which recognizes antigen. Numerous antigen-binding domains are known in the art, including those based on the antigen binding site of an antibody, antibody mimetics, and T-cell receptors. For example, the antigen-binding domain may comprise: a single-chain variable fragment (scFv) derived from a monoclonal antibody; a natural ligand of the target antigen; a peptide with sufficient affinity for the target; a single domain antibody; an artificial single binder such as a Darpin (designed ankyrin repeat protein); or a single-chain derived from a T-cell receptor.
[0260] The antigen binding domain may comprise a domain which is not based on the antigen binding site of an antibody. For example the antigen binding domain may comprise a domain based on a protein/peptide which is a soluble ligand for a tumour cell surface receptor (e.g. a soluble peptide such as a cytokine or a chemokine); or an extracellular domain of a membrane anchored ligand or a receptor for which the binding pair counterpart is expressed on the tumour cell.
[0261] Examples 11 to 13 relate to a CAR which binds BCMA, in which the antigen binding doaimn comprises APRIL, a ligand for BCMA.
[0262] The antigen binding domain may be based on a natural ligand of the antigen.
[0263] The antigen binding domain may comprise an affinity peptide from a combinatorial library or a de novo designed affinity protein/peptide.
[0264] Spacer Domain CARs comprise a spacer sequence to connect the antigen-binding domain with the transmembrane domain and spatially separate the antigen-binding domain from the endodomain. A flexible spacer allows the antigen-binding domain to orient in different directions to facilitate binding.
[0265] In the T cell of the present invention, the first and second CARs comprise different spacer molecules. For example, the spacer sequence may, for example, comprise an IgG1 Fc region, an IgG1 hinge or a human CD8 stalk or the mouse CD8 stalk. The spacer may alternatively comprise an alternative linker sequence which has similar length and/or domain spacing properties as an IgG1 Fc region, an IgG1 hinge or a CD8 stalk. A human IgG1 spacer may be altered to remove Fc binding motifs.
[0266] Examples of amino acid sequences for these spacers are given below:
TABLE-US-00011 (hinge-CH2CH3 of human IgG1) SEQ ID No. 10 AEPKSPDKTHTCPPCPAPPVAGPSVFLFPPKPKDTLMIARTPEVTCVVVD VSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLN GKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSL TCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKS RWQQGNVFSCSVMHEALHNHYTQKSLSLSPGKKD (human CD8 stalk): SEQ ID No. 11 TTTPAPRPPTPAPTIASQPLSLRPEACRPAAGGAVHTRGLDFACDI (human IgG1 hinge): SEQ ID No. 12 AEPKSPDKTHTCPPCPKDPK (CD2 ectodomain) SEQ ID No. 13 KEITNALETWGALGQDINLDIPSFQMSDDIDDIKWEKTSDKKKIAQFRKE KETFKEKDTYKLFKNGTLKIKHLKTDDQDIYKVSIYDTKGKNVLEKIFDL KIQERVSKPKISWTCINTTLTCEVMNGTDPELNLYQDGKHLKLSQRVITH KWTTSLSAKFKCTAGNKVSKESSVEPVSCPEKGLD (CD34 ectodomain) SEQ ID no. 14 SLDNNGTATPELPTQGTFSNVSTNVSYQETTTPSTLGSTSLHPVSQHGNE ATTNITETTVKFTSTSVITSVYGNTNSSVQSQTSVISTVFTTPANVSTPE TTLKPSLSPGNVSDLSTTSTSLATSPTKPYTSSSPILSDIKAEIKCSGIR EVKLTQGICLEQNKTSSCAEFKKDRGEGLARVLCGEEQADADAGAQVCSL LLAQSEVRPQCLLLVLANRTEISSKLQLMKKHQSDLKKLGILDFTEQDVA SHQSYSQKT
[0267] Since CARs are typically homodimers (see FIG. 1a), cross-pairing may result in a heterodimeric chimeric antigen receptor. This is undesirable for various reasons, for example: (1) the epitope may not be at the same "level" on the target cell so that a cross-paired CAR may only be able to bind to one antigen; (2) the VH and VL from the two different scFv could swap over and either fail to recognize target or worse recognize an unexpected and unpredicted antigen. For the "OR" gate and the "AND NOT" gate, the spacer of the first CAR may be sufficiently different from the spacer of the second CAR in order to avoid cross-pairing. The amino acid sequence of the first spacer may share less that 50%, 40%, 30% or 20% identity at the amino acid level with the second spacer.
[0268] In other aspects of the invention (for example the AND gate) it is important that the spacer of the first CAR has a different length, and/or charge and/or shape and/or configuration and/or glycosylation, such that when both first and second CARs bind their target antigen, the difference in spacer charge or dimensions results in spatial separation of the two types of CAR to different parts of the membrane to result in activation as predicted by the kinetic separation model. In these aspects, the different length, shape and/or configuration of the spacers is carefully chosen bearing in mind the size and binding epitope on the target antigen to allow differential segregation upon cognate target recognition. For example the IgG1 Hinge, CD8 stalk, IgG1 Fc, ectodomain of CD34, ectodomain of CD45 are expected to differentially segregate.
[0269] Examples of spacer pairs which differentially segregate and are therefore suitable for use with the AND gate are shown in the following Table:
TABLE-US-00012 Stimulatory CAR spacer Inhibitory CAR spacer Human-CD8STK Human-IgG-Hinge-CH2CH3 Human-CD3z ectodomain Human-IgG-Hinge-CH2CH3 Human-IgG-Hinge Human-IgG-Hinge-CH2CH3 Human-CD28STK Human-IgG-Hinge-CH2CH3 Human-CD8STK Human-IgM-Hinge-CH2CH3CD4 Human-CD3z ectodomain Human-IgM-Hinge-CH2CH3CD4 Human-IgG-Hinge Human-IgM-Hinge-CH2CH3CD4 Human-CD28STK Human-IgM-Hinge-CH2CH3CD4
[0270] In other aspects of the invention (for example the AND NOT gate), it is important that the spacer be sufficiently different as to prevent cross-pairing, but to be sufficiently similar to co-localise. Pairs of orthologous spacer sequences may be employed. Examples are murine and human CD8 stalks, or alternatively spacer domains which are monomeric--for instance the ectodomain of CD2.
[0271] Examples of spacer pairs which co-localise and are therefore suitable for use with the AND NOT gate are shown in the following Table:
TABLE-US-00013 Stimulatory CAR spacer Inhibitory CAR spacer Human-CD8aSTK Mouse CD8aSTK Human-CD28STK Mouse CD8aSTK Human-IgG-Hinge Human-CD3z ectodomain Human-CD8aSTK Mouse CD28STK Human-CD28STK Mouse CD28STK Human-IgG-Hinge-CH2CH3 Human-IgM-Hinge-CH2CH3CD4
[0272] All the spacer domains mentioned above form homodimers. However the mechanism is not limited to using homodimeric receptors and should work with monomeric receptors as long as the spacer is sufficiently rigid. An example of such a spacer is CD2 or truncated CD22.
[0273] Transmembrane Domain
[0274] The transmembrane domain is the sequence of the CAR that spans the membrane.
[0275] A transmembrane domain may be any protein structure which is thermodynamically stable in a membrane. This is typically an alpha helix comprising of several hydrophobic residues. The transmembrane domain of any transmembrane protein can be used to supply the transmembrane portion of the invention. The presence and span of a transmembrane domain of a protein can be determined by those skilled in the art using the TMHMM algorithm (http://www.cbs.dtu.dk/services/TMHMM-2.0/). Further, given that the transmembrane domain of a protein is a relatively simple structure, i.e a polypeptide sequence predicted to form a hydrophobic alpha helix of sufficient length to span the membrane, an artificially designed TM domain may also be used (U.S. Pat. No. 7,052,906 B1 describes synthetic transmembrane components).
[0276] The transmembrane domain may be derived from CD28, which gives good receptor stability.
[0277] Activating Endodomain
[0278] The endodomain is the signal-transmission portion of the CAR. After antigen recognition, receptors cluster, native CD45 and CD148 are excluded from the synapse and a signal is transmitted to the cell. The most commonly used endodomain component is that of CD3-zeta which contains 3 ITAMs. This transmits an activation signal to the T cell after antigen is bound. CD3-zeta may not provide a fully competent activation signal and additional co-stimulatory signaling may be needed. For example, chimeric CD28 and OX40 can be used with CD3-Zeta to transmit a proliferative/survival signal, or all three can be used together.
[0279] In the "OR gate", the cell of the present invention comprises two CARs, each with an activating endodomain. The activating endodomain is capable of transmitting both immunological signal 1 and immunological signal 2. An endodomain is not considered "activating" if it just comprises CD3.zeta., which is sufficient to trigger T-cell killing of cognate target cells does not fully activate the T-cell to proliferate and survive. An "activating endodomain" is a compound endodomains: in which the intracellular part of a T-cell co-stimulatory molecule is fused to that of CD3 .zeta. resulting in a "second generation" receptor which can transmit an activating and co-stimulatory signal simultaneously after antigen recognition. The co-stimulatory domain most commonly used is that of CD28. This supplies the most potent co-stimulatory signal--namely immunological signal 2, which triggers T-cell proliferation. In addition to a co-stimulatory domain, the activating endodomain may also include TNF receptor family endodomains, such as the closely related OX40 and 41BB which transmit survival signals.
[0280] An "activating" endodomain .may therefore not comprise the CD3-Zeta endodomain or the CD28 endodomain alone. An activating endodomain may, for example, comprise the CD3-Zeta endodomain with that of either CD28 or OX40 or the CD28 endodomain and OX40 and CD3-Zeta endodomain.
[0281] Each of the CARs in the OR gate is independently capable of activating the T cell. The T cell is thus activated by the presence of either antigen alone. The two CARs are not "complementary" in the sense that activation of both CARs is necessary to provide activation and co-stimulatory signals.
[0282] A endodomain which contains an ITAM motif can act as an activation endodomain in this invention. Several proteins are known to contain endodomains with one or more ITAM motifs. Examples of such proteins include the CD3 epsilon chain, the CD3 gamma chain and the CD3 delta chain to name a few. The ITAM motif can be easily recognized as a tyrosine separated from a leucine or isoleucine by any two other amino acids, giving the signature YxxL/I. Typically, but not always, two of these motifs are separated by between 6 and 8 amino acids in the tail of the molecule (YxxL/Ix(6-8)YxxL/I). Hence, one skilled in the art can readily find existing proteins which contain one or more ITAM to transmit an activation signal. Further, given the motif is simple and a complex secondary structure is not required, one skilled in the art can design polypeptides containing artificial ITAMs to transmit an activation signal (see WO 2000063372, which relates to synthetic signalling molecules).
[0283] The transmembrane and intracellular T-cell signalling domain (endodomain) of a CAR with an activating endodomain may comprise the sequence shown as SEQ ID No. 16 or 17 or a variant thereof having at least 80% sequence identity.
TABLE-US-00014 comprising CD28 transmembrane domain and CD3 Z endodomain SEQ ID No. 15 FWVLVVVGGVLACYSLLVTVAFIIFWVRRVKFSRSADAPAYQQGQNQLYN ELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYS EIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR comprising CD28 transmembrane domain and CD28 and CD3 Zeta endodomains SEQ ID No. 16 FWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPT RKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEY DVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRR GKGHDGLYQGLSTATKDTYDALHMQALPPR comprising CD28 transmembrane domain and CD28, OX40 and CD3 Zeta endodomains. SEQ ID No. 17 FWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPT RKHYQPYAPPRDFAAYRSRDQRLPPDAHKPPGGGSFRTPIQEEQADAHST LAKIRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMG GKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTA TKDTYDALHMQALPPR
[0284] A variant sequence may have at least 80%, 85%, 90%, 95%, 98% or 99% sequence identity to SEQ ID No. 16 or 17, provided that the sequence provides an effective trans-membrane domain and an effective intracellular T cell signaling domain.
[0285] "Ligation-Off" Inhibitory Endodomain
[0286] In the embodiment referred above as the AND gate, one of the CARs comprises an inhibitory endodomain such that the inhibitory CAR inhibits T-cell activation by the activating CAR in the absence of inhibitory CAR ligation, but does not significantly inhibit T-cell activation by the activating CAR when the inhibitory CAR is ligated. This is termed a "ligation-off" inhibitory endodomain.
[0287] In this case, the spacer of the inhibitory CAR is of a different length, charge, shape and/or configuration and/or glycosylation from the spacer of the activating CAR, such that when both receptors are ligated, the difference in spacer dimensions results in isolation of the activating CARs and the inhibitory CARs in different membrane compartments of the immunological synapse, so that the activating endodomain is released from inhibition by the inhibitory endodomain.
[0288] The inhibitory endodomains for use in a ligation-off inhibitory CAR may therefore comprise any sequence which inhibits T-cell signaling by the activating CAR when it is in the same membrane compartment (i.e. in the absence of the antigen for the inhibitory CAR) but which does not significantly inhibit T cell signaling when it is isolated in a separate part of the membrane from the inhibitory CAR.
[0289] The ligation-off inhibitory endodomain may be or comprise a tyrosine phosphatase, such as a receptor-like tyrosine phosphatase. An inhibitory endodomain may be or comprise any tyrosine phosphatase that is capable of inhibiting the TCR signalling when only the stimulatory receptor is ligated. An inhibitory endodomain may be or comprise any tyrosine phosphatase with a sufficiently fast catalytic rate for phosphorylated ITAMs that is capable of inhibiting the TCR signalling when only the stimulatory receptor is ligated.
[0290] For example, the inhibitory endodomain of an AND gate may comprise the endodomain of CD148 or CD45. CD148 and CD45 have been shown to act naturally on the phosphorylated tyrosines up-stream of TCR signalling.
[0291] CD148 is a receptor-like protein tyrosine phosphatase which negatively regulates TCR signaling by interfering with the phosphorylation and function of PLC.gamma.1 and LAT.
[0292] CD45 present on all hematopoetic cells, is a protein tyrosine phosphatase which is capable of regulating signal transduction and functional responses, again by phosphorylating PLC.gamma.1.
[0293] An inhibitory endodomain may comprise all of part of a receptor-like tyrosine phosphatase. The phospatase may interfere with the phosphorylation and/or function of elements involved in T-cell signalling, such as PLC.gamma.1 and/or LAT.
[0294] The transmembrane and endodomain of CD45 and CD148 is shown as SEQ ID No. 18 and No. 19 respectively.
TABLE-US-00015 CD45 trans-membrane and endodomain sequence SEQ ID 18 ALIAFLAFLIIVTSIALLVVLYKIYDLHKKRSCNLDEQQELVERDDEKQL MNVEPIHADILLETYKRKIADEGRLFLAEFQSIPRVFSKFPIKEARKPFN QNKNRYVDILPYDYNRVELSEINGDAGSNYINASYIDGFKEPRKYIAAQG PRDETVDDFWRMIWEQKATVIVMVTRCEEGNRNKCAEYWPSMEEGTRAFG DVVVKINQHKRCPDYIIQKLNIVNKKEKATGREVTHIQFTSWPDHGVPED PHLLLKLRRRVNAFSNFFSGPIVVHCSAGVGRTGTYIGIDAMLEGLEAEN KVDVYGYVVKLRRQRCLMVQVEAQYILIHQALVEYNQFGETEVNLSELHP YLHNMKKRDPPSEPSPLEAEFQRLPSYRSWRTQHIGNQEENKSKNRNSNV IPYDYNRVPLKHELEMSKESEHDSDESSDDDSDSEEPSKYINASFIMSYW KPEVMIAAQGPLKETIGDFWQMIFQRKVKVIVMLTELKHGDQEICAQYWG EGKQTYGDIEVDLKDTDKSSTYTLRVFELRHSKRKDSRTVYQYQYTNWSV EQLPAEPKELISMIQVVKQKLPQKNSSEGNKHHKSTPLLIHCRDGSQQTG IFCALLNLLESAETEEVVDIFQVVKALRKARPGMVSTFEQYQFLYDVIAS TYPAQNGQVKKNNHQEDKIEFDNEVDKVKQDANCVNPLGAPEKLPEAKEQ AEGSEPTSGTEGPEHSVNGPASPALNQGS CD148 trans-membrane and endodomain sequence SEQ ID 19 AVFGCIFGALVIVTVGGFIFWRKKRKDAKNNEVSFSQIKPKKSKLIRVEN FEAYFKKQQADSNCGFAEEYEDLKLVGISQPKYAAELAENRGKNRYNNVL PYDISRVKLSVQTHSTDDYINANYMPGYHSKKDFIATQGPLPNTLKDFWR MVWEKNVYAIIMLTKCVEQGRTKCEEYWPSKQAQDYGDITVAMTSEIVLP EWTIRDFTVKNIQTSESHPLRQFHFTSWPDHGVPDTTDLLINFRYLVRDY MKQSPPESPILVHCSAGVGRTGTFIAIDRLIYQIENENTVDVYGIVYDLR MHRPLMVQTEDQYVFLNQCVLDIVRSQKDSKVDLIYQNTTAMTIYENLAP VTTFGKTNGYIA
[0295] An inhibitory CAR may comprise all or part of SEQ ID No 18 or 19 (for example, it may comprise the phosphatase function of the endodomain). It may comprise a variant of the sequence or part thereof having at least 80% sequence identity, as long as the variant retains the capacity to basally inhibit T cell signalling by the activating CAR.
[0296] Other spacers and endodomains may be tested for example using the model system exemplified herein. Target cell populations can be created by transducing a suitable cell line such as a SupT1 cell line either singly or doubly to establish cells negative for both antigens (the wild-type), positive for either and positive for both (e.g. CD19-CD33-, CD19+CD33-, CD19-CD33+ and CD19+CD33+). T cells such as the mouse T cell line BW5147 which releases IL-2 upon activation may be transduced with pairs of CARs and their ability to function in a logic gate measured through measurement of IL-2 release (for example by ELISA). For example, it is shown in Example 4 that both CD148 and CD45 endodomains can function as inhibitory CARs in combination with an activating CAR containing a CD3 Zeta endodomain. These CARs rely upon a short/non-bulky CD8 stalk spacer on one CAR and a bulky Fc spacer on the other CAR to achieve AND gating. When both receptors are ligated, the difference in spacer dimensions results in isolation of the different receptors in different membrane compartments, releasing the CD3 Zeta receptor from inhibition by the CD148 or CD45 endodomains. In this way, activation only occurs once both receptors are activated. It can be readily seen that this modular system can be used to test alternative spacer pairs and inhibitory endodomains. If the spacers do not achieve isolation following ligation of both receptors, the inhibition would not be released and so no activation would occur. If the inhibitory endodomain under test is ineffective, activation would be expected in the presence of ligation of the activating CAR irrespective of the ligation status of the inhibitory CAR.
[0297] "Ligation-On" Endodomain
[0298] In the embodiment referred above as the AND NOT gate, one of the CARs comprises a "ligation-on" inhibitory endodomain such that the inhibitory CAR does not significantly inhibit T-cell activation by the activating CAR in the absence of inhibitory CAR ligation, but inhibits T-cell activation by the activating CAR when the inhibitory CAR is ligated.
[0299] The "ligation-on" inhibitory endodomain may be or comprise a tyrosine phosphatase that is incapable of inhibiting the TCR signalling when only the stimulatory receptor is ligated.
[0300] The "ligation-on" inhibitory endodomain may be or comprise a tyrosine phosphatase with a sufficiently slow catalytic rate for phosphorylated ITAMs that is incapable of inhibiting the TCR signalling when only the stimulatory receptor is ligated but it is capable of inhibiting the TCR signalling response when concentrated at the synapse. Concentration at the synapse is achieved through inhibitory receptor ligation.
[0301] If a tyrosine phosphatase has a catalytic rate which is too fast for a "ligation-on" inhibitory endodomain, then it is possible to tune-down the catalytic rates of phosphatase through modification such as point mutations and short linkers (which cause steric hindrance) to make it suitable for a "ligation-on" inhibitory endodomain.
[0302] In this first embodiment the endodomain may be or comprise a phosphatase which is considerably less active than CD45 or CD148, such that significant dephosphorylation of ITAMS only occurs when activating and inhibitory endodomains are co-localised. Many suitable sequences are known in the art. For example, the inhibitory endodomain of a NOT AND gate may comprise all or part of a protein-tyrosine phosphatase such as PTPN6.
[0303] Protein tyrosine phosphatases (PTPs) are signaling molecules that regulate a variety of cellular processes including cell growth, differentiation, mitotic cycle, and oncogenic transformation. The N-terminal part of this PTP contains two tandem Src homolog (SH2) domains, which act as protein phospho-tyrosine binding domains, and mediate the interaction of this PTP with its substrates. This PTP is expressed primarily in hematopoietic cells, and functions as an important regulator of multiple signaling pathways in hematopoietic cells.
[0304] The inhibitor domain may comprise all of PTPN6 (SEQ ID No. 20) or just the phosphatase domain (SEQ ID No. 21).
TABLE-US-00016 sequence of PTPN6 SEQ ID 20 MVRWFHRDLSGLDAETLLKGRGVHGSFLARPSRKNQGDFSLSVRVGDQVT HIRIQNSGDFYDLYGGEKFATLTELVEYYTQQQGVLQDRDGTIIHLKYPL NCSDPTSERWYHGHMSGGQAETLLQAKGEPWTFLVRESLSQPGDFVLSVL SDQPKAGPGSPLRVTHIKVMCEGGRYTVGGLETFDSLTDLVEHFKKTGIE EASGAFVYLRQPYYATRVNAADIENRVLELNKKQESEDTAKAGFWEEFES LQKQEVKNLHQRLEGQRPENKGKNRYKNILPFDHSRVILQGRDSNIPGSD YINANYIKNQLLGPDENAKTYIASQGCLEATVNDFWQMAWQENSRVIVMT TREVEKGRNKCVPYWPEVGMQRAYGPYSVTNCGEHDTTEYKLRTLQVSPL DNGDLIREIWHYQYLSWPDHGVPSEPGGVLSFLDQINQRQESLPHAGPII VHCSAGIGRTGTIIVIDMLMENISTKGLDCDIDIQKTIQMVRAQRSGMVQ TEAQYKFIYVAIAQFIETTKKKLEVLQSQKGQESEYGNITYPPAMKNAHA KASRTSSKHKEDVYENLHTKNKREEKVKKQRSADKEKSKGSLKRK sequence of phosphatase domain of PTPN6 SEQ ID 21 FWEEFESLQKQEVKNLHQRLEGQRPENKGKNRYKNILPFDHSRVILQGRD SNIPGSDYINANYIKNQLLGPDENAKTYIASQGCLEATVNDFWQMAWQEN SRVIVMTTREVEKGRNKCVPYWPEVGMQRAYGPYSVTNCGEHDTTEYKLR TLQVSPLDNGDLIREIWHYQYLSWPDHGVPSEPGGVLSFLDQINQRQESL PHAGPIIVHCSAGIGRTGTIIVIDMLMENISTKGLDCDIDIQKTIQMVRA QRSGMVQTEAQYKFIYVAIAQF
[0305] A second embodiment of a ligation-on inhibitory endodomain is an ITIM (Immunoreceptor Tyrosine-based Inhibition motif) containing endodomain such as that from CD22, LAIR-1, the Killer inhibitory receptor family (KIR), LILRB1, CTLA4, PD-1, BTLA etc. When phosphorylated, ITIMs recruits endogenous PTPN6 through its SH2 domain. If co-localised with an ITAM containing endodomain, dephosphorylation occurs and the activating CAR is inhibited.
[0306] An ITIM is a conserved sequence of amino acids (S/I/V/LxYxxI/V/L) that is found in the cytoplasmic tails of many inhibitory receptors of the immune system. One skilled in the art can easily find protein domains containing an ITIM. A list of human candidate ITIM-containing proteins has been generated by proteome-wide scans (Staub, et al (2004) Cell. Signal. 16, 435-456). Further, since the consensus sequence is well known and little secondary structure appears to be required, one skilled in the art could generate an artificial ITIM.
[0307] ITIM endodomains from PDCD1, BTLA4, LILRB1, LAIR1, CTLA4, KIR2DL1, KIR2DL4, KIR2DL5, KIR3DL1 and KIR3DL3 are shown in SEQ ID 22 to 31 respectively
TABLE-US-00017 PDCD1 endodomain SEQ ID 22 CSRAARGTIGARRTGQPLKEDPSAVPVFSVDYGELDFQWREKTPEPPVPC VPEQTEYATIVFPSGMGTSSPARRGSADGPRSAQPLRPEDGHCSWPL BTLA4 SEQ ID 23 KLQRRWKRTQSQQGLQENSSGQSFFVRNKKVRRAPLSEGPHSLGCYNPMM EDGISYTTLRFPEMNIPRTGDAESSEMQRPPPDCDDTVTYSALHKRQVGD YENVIPDFPEDEGIHYSELIQFGVGERPQAQENVDYVILKH LILRB1 SEQ ID 24 LRHRRQGKHWTSTQRKADFQHPAGAVGPEPTDRGLQWRSSPAADAQEENL YAAVKHTQPEDGVEMDTRSPHDEDPQAVTYAEVKHSRPRREMASPPSPLS GEFLDTKDRQAEEDRQMDTEAAASEAPQDVTYAQLHSLTLRREATEPPPS QEGPSPAVPSIYATLAIH LAIR1 SEQ ID 25 HRQNQIKQGPPRSKDEEQKPQQRPDLAVDVLERTADKATVNGLPEKDRET DTSALAAGSSQEVTYAQLDHWALTQRTARAVSPQSTKPMAESITYAAVAR H CTLA4 SEQ ID 26 FLLWILAAVSSGLFFYSFLLTAVSLSKMLKKRSPLTTGVYVKMPPTEPEC EKQFQPYFIPIN KIR2DL1 SEQ ID 27 GNSRHLHVLIGTSVVIIPFAILLFFLLHRWCANKKNAVVMDQEPAGNRTV NREDSDEQDPQEVTYTQLNHCVFTQRKITRPSQRPKTPPTDIIVYTELPN AESRSKVVSCP KIR2DL4 SEQ ID 28 GIARHLHAVIRYSVAIILFTILPFFLLHRWCSKKKENAAVMNQEPAGHRT VNREDSDEQDPQEVTYAQLDHCIFTQRKITGPSQRSKRPSTDTSVCIELP NAEPRALSPAHEHHSQALMGSSRETTALSQTQLASSNVPAAGI KIR2DL5 SEQ ID 29 TGIRRHLHILIGTSVAIILFIILFFFLLHCCCSNKKNAAVMDQEPAGDRT VNREDSDDQDPQEVTYAQLDHCVFTQTKITSPSQRPKTPPTDTTMYMELP NAKPRSLSPAHKHHSQALRGSSRETTALSQNRVASSHVPAAGI KIR3DL1 SEQ ID 30 KDPRHLHILIGTSVVIILFILLLFFLLHLWCSNKKNAAVMDQEPAGNRTA NSEDSDEQDPEEVTYAQLDHCVFTQRKITRPSQRPKTPPTDTILYTELPN AKPRSKVVSCP KIR3DL3 SEQ ID 31 KDPGNSRHLHVLIGTSVVIIPFAILLFFLLHRWCANKKNAVVMDQEPAGN RTVNREDSDEQDPQEVTYAQLNHCVFTQRKITRPSQRPKTPPTDTSV
[0308] A third embodiment of a ligation-on inhibitory endodomain is an ITIM containing endodomain co-expressed with a fusion protein. The fusion protein may comprise at least part of a protein-tyrosine phosphatase and at least part of a receptor-like tyrosine phosphatase. The fusion may comprise one or more SH2 domains from the protein-tyrosine phosphatase. For example, the fusion may be between a PTPN6 SH2 domain and CD45 endodomain or between a PTPN6 SH2 domain and CD148 endodomain. When phosphorylated, the ITIM domains recruit the fusion protein bring the highly potent CD45 or CD148 phosphatase to proximity to the activating endodomain blocking activation.
[0309] SEQUENCES of fusion proteins are listed 32 and 33
TABLE-US-00018 PTPN6-CD45 fusion protein SEQ ID 32 WYHGHMSGGQAETLLQAKGEPWTFLVRESLSQPGDFVLSVLSDQPKAGPG SPLRVTHIKVMCEGGRYTVGGLETFDSLTDLVEHFKKTGIEEASGAFVYL RQPYKIYDLHKKRSCNLDEQQELVERDDEKQLMNVEPIHADILLETYKRK IADEGRLFLAEFQSIPRVFSKFPIKEARKPFNQNKNRYVDILPYDYNRVE LSEINGDAGSNYINASYIDGFKEPRKYIAAQGPRDETVDDFWRMIWEQKA TVIVMVTRCEEGNRNKCAEYWPSMEEGTRAFGDVVVKINQHKRCPDYIIQ KLNIVNKKEKATGREVTHIQFTSWPDHGVPEDPHLLLKLRRRVNAFSNFF SGPIVVHCSAGVGRTGTYIGIDAMLEGLEAENKVDVYGYVVKLRRQRCLM VQVEAQYILIHQALVEYNQFGETEVNLSELHPYLHNMKKRDPPSEPSPLE AEFQRLPSYRSWRTQHIGNQEENKSKNRNSNVIPYDYNRVLKHELEMSKE SEHDSDESSDDDSDSEEPSKYINASFIMSYWKPEVMIAAQGPLKETIGDF MIQRKVKVIVMLTELKHGDQEICAQYWGEGKQTYGDIEVDLKDTDKSSTY TLRVFELRHSKRKDSRTVYQYQYTNWSVEQLPAEPKELISMIQVVKQKLP QKNSSEGNKHHKSTPLLIHCRDGSQQTGIFCALLNLLESAETEEVVDIFQ VVKALRKARPGMVSTFEQYQFLYDVIASTYPAQNGQVKKNNHQEDKIEFD NEVDKVKQDANCVNPLGAPEKLPEAKEQAEGSEPTSGTEGPEHSVNGPAS PALNQGS PTPN6-CD148 fusion SEQ ID 33 ETLLQAKGEPWTFLVRESLSQPGDFVLSVLSDQPKAGPGSPLRVTHIKVM CEGGRYTVGGLETFDSLTDLVEHFKKTGIEEASGAFVYLRQPYRKKRKDA KNNEVSFSQIKPKKSKLIRVENFEAYFKKQQADSNCGFAEEYEDLKLVGI SQPKYAAELAENRGKNRYNNVLPYDISRVKLSVQTHSTDDYINANYMPGY HSKKDFIATQGPLPNTLKDFWRMVWEKNVYAIIMLTKCVEQGRTKCEEYW PSKQAQDYGDITVAMTSEIVLPEWTIRDFTVKNIQTSESHPLRQFHFTSW PDHGVPDTTDLLINFRYLVRDYMKQSPPESPILVHCSAGVGRTGTFIAID RLIYQIENENTVDVYGIVYDLRMHRPLMVQTEDQYVFLNQCVLDIVRSQK DSKVDLIYQNTTAMTIYENLAPVTTFGKTNGYIA
[0310] A ligation-on inhibitory CAR may comprise all or part of SEQ ID No 20 or 21. It may comprise all or part of SEQ ID 22 to 31. It may comprise all or part of SEQ ID 22 to 31 co-expressed with either SEQ ID 32 or 33. It may comprise a variant of the sequence or part thereof having at least 80% sequence identity, as long as the variant retains the capacity to inhibit T cell signaling by the activating CAR upon ligation of the inhibitory CAR.
[0311] As above, alternative spacers and endodomains may be tested for example using the model system exemplified herein. It is shown in Example 5 that the PTPN6 endodomain can function as a semi-inhibitory CAR in combination with an activating CAR containing a CD3 Zeta endodomain. These CARs rely upon a human CD8 stalk spacer on one CAR and a mouse CD8 stalk spacer on the other CAR. The orthologous sequences prevent cross pairing. However, when both receptors are ligated, the similarity between the spacers results in co-segregation of the different receptors in the same membrane compartments. This results in inhibition of the CD3 Zeta receptor by the PTPN6 endodomain. If only the activating CAR is ligated the PTPN6 endodomain is not sufficiently active to prevent T cell activation. In this way, activation only occurs if the activating CAR is ligated and the inhibitory CAR is not ligated (AND NOT gating). It can be readily seen that this modular system can be used to test alternative spacer pairs and inhibitory domains. If the spacers do not achieve co-segregation following ligation of both receptors, the inhibition would not be effective and so activation would occur. If the semi-inhibitory endodomain under test is ineffective, activation would be expected in the presence of ligation of the activating CAR irrespective of the ligation status of the semi-inhibitory CAR.
[0312] Co-Expression Site
[0313] The second aspect of the invention relates to a nucleic acid which encodes the first and second CARs.
[0314] The nucleic acid may produce a polypeptide which comprises the two CAR molecules joined by a cleavage site. The cleavage site may be self-cleaving, such that when the polypeptide is produced, it is immediately cleaved into the first and second CARs without the need for any external cleavage activity.
[0315] Various self-cleaving sites are known, including the Foot-and-Mouth disease virus (FMDV) 2a self-cleaving peptide, which has the sequence shown as SEQ ID No. 34:
TABLE-US-00019 SEQ ID No. 34 RAEGRGSLLTCGDVEENPGP.
[0316] The co-expressing sequence may be an internal ribosome entry sequence (IRES). The co-expressing sequence may be an internal promoter.
[0317] Cell
[0318] The first aspect of the invention relates to a cell which co-expresses a first CAR and a second CAR at the cell surface.
[0319] The cell may be any eukaryotic cell capable of expressing a CAR at the cell surface, such as an immunological cell.
[0320] In particular the cell may be an immune effector cell such as a T cell or a natural killer (NK) cell.
[0321] T cells or T lymphocytes are a type of lymphocyte that play a central role in cell-mediated immunity. They can be distinguished from other lymphocytes, such as B cells and natural killer cells (NK cells), by the presence of a T-cell receptor (TCR) on the cell surface. There are various types of T cell, as summarised below.
[0322] Helper T helper cells (TH cells) assist other white blood cells in immunologic processes, including maturation of B cells into plasma cells and memory B cells, and activation of cytotoxic T cells and macrophages. TH cells express CD4 on their surface. TH cells become activated when they are presented with peptide antigens by MHC class II molecules on the surface of antigen presenting cells (APCs). These cells can differentiate into one of several subtypes, including TH1, TH2, TH3, TH17, Th9, or TFH, which secrete different cytokines to facilitate different types of immune responses.
[0323] Cytotoxic T cells (TC cells, or CTLs) destroy virally infected cells and tumor cells, and are also implicated in transplant rejection. CTLs express the CD8 at their surface. These cells recognize their targets by binding to antigen associated with MHC class I, which is present on the surface of all nucleated cells. Through IL-10, adenosine and other molecules secreted by regulatory T cells, the CD8+ cells can be inactivated to an anergic state, which prevent autoimmune diseases such as experimental autoimmune encephalomyelitis.
[0324] Memory T cells are a subset of antigen-specific T cells that persist long-term after an infection has resolved. They quickly expand to large numbers of effector T cells upon re-exposure to their cognate antigen, thus providing the immune system with "memory" against past infections. Memory T cells comprise three subtypes: central memory T cells (TCM cells) and two types of effector memory T cells (TEM cells and TEMRA cells). Memory cells may be either CD4+ or CD8+. Memory T cells typically express the cell surface protein CD45RO.
[0325] Regulatory T cells (Treg cells), formerly known as suppressor T cells, are crucial for the maintenance of immunological tolerance. Their major role is to shut down T cell-mediated immunity toward the end of an immune reaction and to suppress auto-reactive T cells that escaped the process of negative selection in the thymus.
[0326] Two major classes of CD4+ Treg cells have been described--naturally occurring Treg cells and adaptive Treg cells.
[0327] Naturally occurring Treg cells (also known as CD4+CD25+FoxP3+ Treg cells) arise in the thymus and have been linked to interactions between developing T cells with both myeloid (CD11c+) and plasmacytoid (CD123+) dendritic cells that have been activated with TSLP. Naturally occurring Treg cells can be distinguished from other T cells by the presence of an intracellular molecule called FoxP3. Mutations of the FOXP3 gene can prevent regulatory T cell development, causing the fatal autoimmune disease IPEX.
[0328] Adaptive Treg cells (also known as Tr1 cells or Th3 cells) may originate during a normal immune response.
[0329] The T cell of the invention may be any of the T cell types mentioned above, in particular a CTL.
[0330] Natural killer (NK) cells are a type of cytolytic cell which forms part of the innate immune system. NK cells provide rapid responses to innate signals from virally infected cells in an MHC independent manner NK cells (belonging to the group of innate lymphoid cells) are defined as large granular lymphocytes (LGL) and constitute the third kind of cells differentiated from the common lymphoid progenitor generating B and T lymphocytes. NK cells are known to differentiate and mature in the bone marrow, lymph node, spleen, tonsils and thymus where they then enter into the circulation.
[0331] The CAR cells of the invention may be any of the cell types mentioned above.
[0332] CAR-expressing cells, such as CAR-expressing T or NK cells may either be created ex vivo either from a patient's own peripheral blood (1.sup.st party), or in the setting of a haematopoietic stem cell transplant from donor peripheral blood (2.sup.nd party), or peripheral blood from an unconnected donor (3.sup.rd party).
[0333] The present invention also provide a cell composition comprising CAR expressing T cells and/or CAR expressing NK cells according to the present invention. The cell composition may be made by tranducing a blood-sample ex vivo with a nucleic acid according to the present invention.
[0334] Alternatively, CAR-expressing cells may be derived from ex vivo differentiation of inducible progenitor cells or embryonic progenitor cells to the relevant cell type, such as T cells. Alternatively, an immortalized cell line such as a T-cell line which retains its lytic function and could act as a therapeutic may be used.
[0335] In all these embodiments, CAR cells are generated by introducing DNA or RNA coding for the CARs by one of many means including transduction with a viral vector, transfection with DNA or RNA.
[0336] A CAR T cell of the invention may be an ex vivo T cell from a subject. The T cell may be from a peripheral blood mononuclear cell (PBMC) sample. T cells may be activated and/or expanded prior to being transduced with CAR-encoding nucleic acid, for example by treatment with an anti-CD3 monoclonal antibody.
[0337] A CAR T cell of the invention may be made by: [0338] (i) isolation of a T cell-containing sample from a subject or other sources listed above; and [0339] (ii) transduction or transfection of the T cells with one or more nucleic acid sequence(s) encoding the first and second CAR.
[0340] The T cells may then by purified, for example, selected on the basis of co-expression of the first and second CAR.
[0341] Nucleic Acid Sequences
[0342] The second aspect of the invention relates to one or more nucleic acid sequence(s) which codes for a first CAR and a second CAR as defined in the first aspect of the invention.
[0343] The nucleic acid sequence may comprise one of the following sequences, or a variant thereof
[0344] SEQ ID 35 OR gate
[0345] SEQ ID 36 AND gate using CD45
[0346] SEQ ID 37 AND gate using CD148
[0347] SEQ ID 38 AND NOT gate using PTPN6 as endodomain
[0348] SEQ ID 39 AND NOT gate using LAIR1 endodomain
[0349] SEQ ID 40 AND NOT gate using LAIR1 and PTPN6 SH2 fusion with CD148 phosphatase
TABLE-US-00020 SEQ ID No. 35: >MP13974.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-HCH2CH3pvaa- CD28tmZw ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCC AGACCAGACATCCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGAC CGGGTGACCATCAGCTGCAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTACC AGCAGAAGCCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCCGGCTGCACA GCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGACTACAGCCTGACC ATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACACCC TGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAG GCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGT GAAGCTGCAGGAGTCTGGCCCAGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGAC CTGCACCGTGAGCGGCGTGAGCCTGCCCGACTACGGCGTGAGCTGGATCAGGCAGCC CCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGCGAGACCACCTACTA CAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCAGGT GTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAG CACTACTACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTG ACCGTGAGCTCAGATCCCACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCC ACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGG GGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTTTTGGGTGCTGGT GGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTTATTATTT TCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGG GCCAGAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTT GGACAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACC CTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGGAGGCCTACAGTGA GATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGCACGATGGCCTTTACCAGG GTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGGCCCTGCCTCC TCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCC CGGGCCCATGGCCGTGCCCACTCAGGTCCTGGGGTTGTTGCTACTGTGGCTTACAGAT GCCAGATGTGACATCCAGATGACACAGTCTCCATCTTCCCTGTCTGCATCTGTCGGAGA TCGCGTCACCATCACCTGTCGAGCAAGTGAGGACATTTATTTTAATTTAGTGTGGTATCA GCAGAAACCAGGAAAGGCCCCTAAGCTCCTGATCTATGATACAAATCGCTTGGCAGAT GGGGTCCCATCACGGTTCAGTGGCTCTGGATCTGGCACACAGTATACTCTAACCATAA GTAGCCTGCAACCCGAAGATTTCGCAACCTATTATTGTCAACACTATAAGAATTATCCGC TCACGTTCGGTCAGGGGACCAAGCTGGAAATCAAAAGATCTGGTGGCGGAGGGTCAG GAGGCGGAGGCAGCGGAGGCGGTGGCTCGGGAGGCGGAGGCTCGAGATCTGAGGTG CAGTTGGTGGAGTCTGGGGGCGGCTTGGTGCAGCCTGGAGGGTCCCTGAGGCTCTCC TGTGCAGCCTCAGGATTCACTCTCAGTAATTATGGCATGCACTGGATCAGGCAGGCTC CAGGGAAGGGTCTGGAGTGGGTCTCGTCTATTAGTCTTAATGGTGGTAGCACTTACTAT CGAGACTCCGTGAAGGGCCGATTCACTATCTCCAGGGACAATGCAAAAAGCACCCTCT ACCTTCAAATGAATAGTCTGAGGGCCGAGGACACGGCCGTCTATTACTGTGCAGCACA GGACGCTTATACGGGAGGTTACTTTGATTACTGGGGCCAAGGAACGCTGGTCACAGTC TCGTCTATGGATCCCGCCGAGCCCAAATCTCCTGACAAAACTCACACATGCCCACCGT GCCCAGCACCTCCCGTGGCCGGCCCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGGA CACCCTCATGATCGCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCA CGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCC AAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGTGTGGTCAGCGTCCTC ACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACA AAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGATGAGCTGACCAAGAACCAGGTCAG CCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAG CAATGGGCAACCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGG CTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAAC GTCTTCTCATGCTCCGTGATGCATGAGGCCCTGCACAATCACTATACCCAGAAATCTCT GAGTCTGAGCCCAGGCAAGAAGGACCCCAAGTTCTGGGTCCTGGTGGTGGTGGGAGG CGTGCTGGCCTGTTACTCTCTCCTGGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCT CCCGGGTGAAGTTTTCTCGCTCTGCCGATGCCCCAGCCTATCAGCAGGGCCAGAATCA GCTGTACAATGAACTGAACCTGGGCAGGCGGGAGGAGTACGACGTGCTGGATAAGCG GAGAGGCAGAGACCCCGAGATGGGCGGCAAACCACGGCGCAAAAATCCCCAGGAGG GACTCTATAACGAGCTGCAGAAGGACAAAATGGCCGAGGCCTATTCCGAGATCGGCAT GAAGGGAGAGAGAAGACGCGGAAAGGGCCACGACGGCCTGTATCAGGGATTGTCCAC CGCTACAAAAGATACATATGATGCCCTGCACATGCAGGCCCTGCCACCCAGATGA SEQ ID No. 36 >MP14802.SFG.aCD19fmc63_clean-CD8STK-CD28tmZ-2A-aCD33glx-HCH2CH3pvaa- dCD45 ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCC AGACCAGACATCCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGAC CGGGTGACCATCAGCTGCAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTACC AGCAGAAGCCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCCGGCTGCACA GCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGACTACAGCCTGACC ATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACACCC TGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAG GCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGT GAAGCTGCAGGAGTCTGGCCCAGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGAC CTGCACCGTGAGCGGCGTGAGCCTGCCCGACTACGGCGTGAGCTGGATCAGGCAGCC CCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGCGAGACCACCTACTA CAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCAGGT GTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAG CACTACTACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTG ACCGTGAGCTCAGATCCCACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCC ACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGG GGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTTTTGGGTGCTGGT GGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTTATTATTT TCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGG GCCAGAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTT GGACAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACC CTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGGAGGCCTACAGTGA GATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGCACGATGGCCTTTACCAGG GTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGGCCCTGCCTCC TCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCC CGGGCCCATGGCCGTGCCCACTCAGGTCCTGGGGTTGTTGCTACTGTGGCTTACAGAT GCCAGATGTGACATCCAGATGACACAGTCTCCATCTTCCCTGTCTGCATCTGTCGGAGA TCGCGTCACCATCACCTGTCGAGCAAGTGAGGACATTTATTTTAATTTAGTGTGGTATCA GCAGAAACCAGGAAAGGCCCCTAAGCTCCTGATCTATGATACAAATCGCTTGGCAGAT GGGGTCCCATCACGGTTCAGTGGCTCTGGATCTGGCACACAGTATACTCTAACCATAA GTAGCCTGCAACCCGAAGATTTCGCAACCTATTATTGTCAACACTATAAGAATTATCCGC TCACGTTCGGTCAGGGGACCAAGCTGGAAATCAAAAGATCTGGTGGCGGAGGGTCAG GAGGCGGAGGCAGCGGAGGCGGTGGCTCGGGAGGCGGAGGCTCGAGATCTGAGGTG CAGTTGGTGGAGTCTGGGGGCGGCTTGGTGCAGCCTGGAGGGTCCCTGAGGCTCTCC TGTGCAGCCTCAGGATTCACTCTCAGTAATTATGGCATGCACTGGATCAGGCAGGCTC CAGGGAAGGGTCTGGAGTGGGTCTCGTCTATTAGTCTTAATGGTGGTAGCACTTACTAT CGAGACTCCGTGAAGGGCCGATTCACTATCTCCAGGGACAATGCAAAAAGCACCCTCT ACCTTCAAATGAATAGTCTGAGGGCCGAGGACACGGCCGTCTATTACTGTGCAGCACA GGACGCTTATACGGGAGGTTACTTTGATTACTGGGGCCAAGGAACGCTGGTCACAGTC TCGTCTATGGATCCCGCCGAGCCCAAATCTCCTGACAAAACTCACACATGCCCACCGT GCCCAGCACCTCCCGTGGCCGGCCCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGGA CACCCTCATGATCGCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCA CGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCC AAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGTGTGGTCAGCGTCCTC ACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACA AAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGATGAGCTGACCAAGAACCAGGTCAG CCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAG CAATGGGCAACCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGG CTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAAC GTCTTCTCATGCTCCGTGATGCATGAGGCCCTGCACAATCACTATACCCAGAAATCTCT GAGTCTGAGCCCAGGCAAGAAGGACCCCAAGGCACTGATAGCATTTCTGGCATTTCTG ATTATTGTGACATCAATAGCCCTGCTTGTTGTTCTCTACAAAATCTATGATCTACATAAGA AAAGATCCTGCAATTTAGATGAACAGCAGGAGCTTGTTGAAAGGGATGATGAAAAACAA CTGATGAATGTGGAGCCAATCCATGCAGATATTTTGTTGGAAACTTATAAGAGGAAGAT TGCTGATGAAGGAAGACTTTTTCTGGCTGAATTTCAGAGCATCCCGCGGGTGTTCAGCA AGTTTCCTATAAAGGAAGCTCGAAAGCCCTTTAACCAGAATAAAAACCGTTATGTTGACA TTCTTCCTTATGATTATAACCGTGTTGAACTCTCTGAGATAAACGGAGATGCAGGGTCAA ACTACATAAATGCCAGCTATATTGATGGTTTCAAAGAACCCAGGAAATACATTGCTGCAC AAGGTCCCAGGGATGAAACTGTTGATGATTTCTGGAGGATGATTTGGGAACAGAAAGC CACAGTTATTGTCATGGTCACTCGATGTGAAGAAGGAAACAGGAACAAGTGTGCAGAAT ACTGGCCGTCAATGGAAGAGGGCACTCGGGCTTTTGGAGATGTTGTTGTAAAGATCAA CCAGCACAAAAGATGTCCAGATTACATCATTCAGAAATTGAACATTGTAAATAAAAAAGA
AAAAGCAACTGGAAGAGAGGTGACTCACATTCAGTTCACCAGCTGGCCAGACCACGGG GTGCCTGAGGATCCTCACTTGCTCCTCAAACTGAGAAGGAGAGTGAATGCCTTCAGCA ATTTCTTCAGTGGTCCCATTGTGGTGCACTGCAGTGCTGGTGTTGGGCGCACAGGAAC CTATATCGGAATTGATGCCATGCTAGAAGGCCTGGAAGCCGAGAACAAAGTGGATGTTT ATGGTTATGTTGTCAAGCTAAGGCGACAGAGATGCCTGATGGTTCAAGTAGAGGCCCA GTACATCTTGATCCATCAGGCTTTGGTGGAATACAATCAGTTTGGAGAAACAGAAGTGA ATTTGTCTGAATTACATCCATATCTACATAACATGAAGAAAAGGGATCCACCCAGTGAGC CGTCTCCACTAGAGGCTGAATTCCAGAGACTTCCTTCATATAGGAGCTGGAGGACACA GCACATTGGAAATCAAGAAGAAAATAAAAGTAAAAACAGGAATTCTAATGTCATCCCATA TGACTATAACAGAGTGCCACTTAAACATGAGCTGGAAATGAGTAAAGAGAGTGAGCATG ATTCAGATGAATCCTCTGATGATGACAGTGATTCAGAGGAACCAAGCAAATACATCAAT GCATCTTTTATAATGAGCTACTGGAAACCTGAAGTGATGATTGCTGCTCAGGGACCACT GAAGGAGACCATTGGTGACTTTTGGCAGATGATCTTCCAAAGAAAAGTCAAAGTTATTG TTATGCTGACAGAACTGAAACATGGAGACCAGGAAATCTGTGCTCAGTACTGGGGAGA AGGAAAGCAAACATATGGAGATATTGAAGTTGACCTGAAAGACACAGACAAATCTTCAA CTTATACCCTTCGTGTCTTTGAACTGAGACATTCCAAGAGGAAAGACTCTCGAACTGTG TACCAGTACCAATATACAAACTGGAGTGTGGAGCAGCTTCCTGCAGAACCCAAGGAATT AATCTCTATGATTCAGGTCGTCAAACAAAAACTTCCCCAGAAGAATTCCTCTGAAGGGA ACAAGCATCACAAGAGTACACCTCTACTCATTCACTGCAGGGATGGATCTCAGCAAACG GGAATATTTTGTGCTTTGTTAAATCTCTTAGAAAGTGCGGAAACAGAAGAGGTAGTGGA TATTTTTCAAGTGGTAAAAGCTCTACGCAAAGCTAGGCCAGGCATGGTTTCCACATTCG AGCAATATCAATTCCTATATGACGTCATTGCCAGCACCTACCCTGCTCAGAATGGACAA GTAAAGAAAAACAACCATCAAGAAGATAAAATTGAATTTGATAATGAAGTGGACAAAGTA AAGCAGGATGCTAATTGTGTTAATCCACTTGGTGCCCCAGAAAAGCTCCCTGAAGCAAA GGAACAGGCTGAAGGTTCTGAACCCACGAGTGGCACTGAGGGGCCAGAACATTCTGTC AATGGTCCTGCAAGTCCAGCTTTAAATCAAGGTTCATAG SEQ ID No. 37: >MP14801.SFG.aCD19fmc63_clean-CD8STK-CD28tmZ-2A-aCD33glx-HCH2CH3pvaa- dCD148 ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCC AGACCAGACATCCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGAC CGGGTGACCATCAGCTGCAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTACC AGCAGAAGCCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCCGGCTGCACA GCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGACTACAGCCTGACC ATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACACCC TGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAG GCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGT GAAGCTGCAGGAGTCTGGCCCAGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGAC CTGCACCGTGAGCGGCGTGAGCCTGCCCGACTACGGCGTGAGCTGGATCAGGCAGCC CCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGCGAGACCACCTACTA CAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCAGGT GTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAG CACTACTACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTG ACCGTGAGCTCAGATCCCACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCC ACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGG GGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTTTTGGGTGCTGGT GGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTTATTATTT TCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGG GCCAGAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTT GGACAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACC CTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGGAGGCCTACAGTGA GATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGCACGATGGCCTTTACCAGG GTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGGCCCTGCCTCC TCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCC CGGGCCCATGGCCGTGCCCACTCAGGTCCTGGGGTTGTTGCTACTGTGGCTTACAGAT GCCAGATGTGACATCCAGATGACACAGTCTCCATCTTCCCTGTCTGCATCTGTCGGAGA TCGCGTCACCATCACCTGTCGAGCAAGTGAGGACATTTATTTTAATTTAGTGTGGTATCA GCAGAAACCAGGAAAGGCCCCTAAGCTCCTGATCTATGATACAAATCGCTTGGCAGAT GGGGTCCCATCACGGTTCAGTGGCTCTGGATCTGGCACACAGTATACTCTAACCATAA GTAGCCTGCAACCCGAAGATTTCGCAACCTATTATTGTCAACACTATAAGAATTATCCGC TCACGTTCGGTCAGGGGACCAAGCTGGAAATCAAAAGATCTGGTGGCGGAGGGTCAG GAGGCGGAGGCAGCGGAGGCGGTGGCTCGGGAGGCGGAGGCTCGAGATCTGAGGTG CAGTTGGTGGAGTCTGGGGGCGGCTTGGTGCAGCCTGGAGGGTCCCTGAGGCTCTCC TGTGCAGCCTCAGGATTCACTCTCAGTAATTATGGCATGCACTGGATCAGGCAGGCTC CAGGGAAGGGTCTGGAGTGGGTCTCGTCTATTAGTCTTAATGGTGGTAGCACTTACTAT CGAGACTCCGTGAAGGGCCGATTCACTATCTCCAGGGACAATGCAAAAAGCACCCTCT ACCTTCAAATGAATAGTCTGAGGGCCGAGGACACGGCCGTCTATTACTGTGCAGCACA GGACGCTTATACGGGAGGTTACTTTGATTACTGGGGCCAAGGAACGCTGGTCACAGTC TCGTCTATGGATCCCGCCGAGCCCAAATCTCCTGACAAAACTCACACATGCCCACCGT GCCCAGCACCTCCCGTGGCCGGCCCGTCAGTCTTCCTCTTCCCCCCAAAACCCAAGGA CACCCTCATGATCGCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACGTGAGCCA CGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCC AAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGTGTGGTCAGCGTCCTC ACCGTCCTGCACCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACA AAGCCCTCCCAGCCCCCATCGAGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGA ACCACAGGTGTACACCCTGCCCCCATCCCGGGATGAGCTGACCAAGAACCAGGTCAG CCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGTGGAGTGGGAGAG CAATGGGCAACCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACGG CTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAAC GTCTTCTCATGCTCCGTGATGCATGAGGCCCTGCACAATCACTATACCCAGAAATCTCT GAGTCTGAGCCCAGGCAAGAAGGACCCCAAGGCGGTTTTTGGCTGTATCTTTGGTGCC CTGGTTATTGTGACTGTGGGAGGCTTCATCTTCTGGAGAAAGAAGAGGAAAGATGCAAA GAATAATGAAGTGTCCTTTTCTCAAATTAAACCTAAAAAATCTAAGTTAATCAGAGTGGA GAATTTTGAGGCCTACTTCAAGAAGCAGCAAGCTGACTCCAACTGTGGGTTCGCAGAG GAATACGAAGATCTGAAGCTTGTTGGAATTAGTCAACCTAAATATGCAGCAGAACTGGC TGAGAATAGAGGAAAGAATCGCTATAATAATGTTCTGCCCTATGATATTTCCCGTGTCAA ACTTTCGGTCCAGACCCATTCAACGGATGACTACATCAATGCCAACTACATGCCTGGCT ACCACTCCAAGAAAGATTTTATTGCCACACAAGGACCTTTACCGAACACTTTGAAAGATT TTTGGCGTATGGTTTGGGAGAAAAATGTATATGCCATCATTATGTTGACTAAATGTGTTG AACAGGGAAGAACCAAATGTGAGGAGTATTGGCCCTCCAAGCAGGCTCAGGACTATGG AGACATAACTGTGGCAATGACATCAGAAATTGTTCTTCCGGAATGGACCATCAGAGATT TCACAGTGAAAAATATCCAGACAAGTGAGAGTCACCCTCTGAGACAGTTCCATTTCACC TCCTGGCCAGACCACGGTGTTCCCGACACCACTGACCTGCTCATCAACTTCCGGTACC TCGTTCGTGACTACATGAAGCAGAGTCCTCCCGAATCGCCGATTCTGGTGCATTGCAGT GCTGGGGTCGGAAGGACGGGCACTTTCATTGCCATTGATCGTCTCATCTACCAGATAG AGAATGAGAACACCGTGGATGTGTATGGGATTGTGTATGACCTTCGAATGCATAGGCCT TTAATGGTGCAGACAGAGGACCAGTATGTTTTCCTCAATCAGTGTGTTTTGGATATTGTC AGATCCCAGAAAGACTCAAAAGTAGATCTTATCTACCAGAACACAACTGCAATGACAAT CTATGAAAACCTTGCGCCCGTGACCACATTTGGAAAGACCAATGGTTACATCGCCTAA SEQ ID No. 38 >16076.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-tm-dPTPN6 ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCC AGACCAGACATCCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGAC CGGGTGACCATCAGCTGCAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTACC AGCAGAAGCCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCCGGCTGCACA GCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGACTACAGCCTGACC ATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACACCC TGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAG GCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGT GAAGCTGCAGGAGTCTGGCCCAGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGAC CTGCACCGTGAGCGGCGTGAGCCTGCCCGACTACGGCGTGAGCTGGATCAGGCAGCC CCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGCGAGACCACCTACTA CAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCAGGT GTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAG CACTACTACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTG ACCGTGAGCTCAGATCCCACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCC ACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGG GGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTTTTGGGTGCTGGT GGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTTATTATTT TCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGG GCCAGAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTT GGACAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACC CTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGGAGGCCTACAGTGA GATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGCACGATGGCCTTTACCAGG GTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGGCCCTGCCTCC TCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCC CGGGCCCATGGCCGTGCCCACTCAGGTCCTGGGGTTGTTGCTACTGTGGCTTACAGAT GCCAGATGTGACATCCAGATGACACAGTCTCCATCTTCCCTGTCTGCATCTGTCGGAGA
TCGCGTCACCATCACCTGTCGAGCAAGTGAGGACATTTATTTTAATTTAGTGTGGTATCA GCAGAAACCAGGAAAGGCCCCTAAGCTCCTGATCTATGATACAAATCGCTTGGCAGAT GGGGTCCCATCACGGTTCAGTGGCTCTGGATCTGGCACACAGTATACTCTAACCATAA GTAGCCTGCAACCCGAAGATTTCGCAACCTATTATTGTCAACACTATAAGAATTATCCGC TCACGTTCGGTCAGGGGACCAAGCTGGAAATCAAAAGATCTGGTGGCGGAGGGTCAG GAGGCGGAGGCAGCGGAGGCGGTGGCTCGGGAGGCGGAGGCTCGAGATCTGAGGTG CAGTTGGTGGAGTCTGGGGGCGGCTTGGTGCAGCCTGGAGGGTCCCTGAGGCTCTCC TGTGCAGCCTCAGGATTCACTCTCAGTAATTATGGCATGCACTGGATCAGGCAGGCTC CAGGGAAGGGTCTGGAGTGGGTCTCGTCTATTAGTCTTAATGGTGGTAGCACTTACTAT CGAGACTCCGTGAAGGGCCGATTCACTATCTCCAGGGACAATGCAAAAAGCACCCTCT ACCTTCAAATGAATAGTCTGAGGGCCGAGGACACGGCCGTCTATTACTGTGCAGCACA GGACGCTTATACGGGAGGTTACTTTGATTACTGGGGCCAAGGAACGCTGGTCACAGTC TCGTCTATGGATCCCGCCACCACAACCAAGCCCGTGCTGCGGACCCCAAGCCCTGTGC ACCCTACCGGCACCAGCCAGCCTCAGAGACCCGAGGACTGCCGGCCTCGGGGCAGC GTGAAGGGCACCGGCCTGGACTTCGCCTGCGACATCTACTGGGCACCTCTGGCCGGA ATATGCGTGGCACTGCTGCTGAGCCTCATCATCACCCTGATCTGTTATCACCGAAGCCG CAAGCGGGTGTGTAAAAGTGGAGGCGGAAGCTTCTGGGAGGAGTTTGAGAGTTTGCA GAAGCAGGAGGTGAAGAACTTGCACCAGCGTCTGGAAGGGCAGCGGCCAGAGAACAA GGGCAAGAACCGCTACAAGAACATTCTCCCCTTTGACCACAGCCGAGTGATCCTGCAG GGACGGGACAGTAACATCCCCGGGTCCGACTACATCAATGCCAACTACATCAAGAACC AGCTGCTAGGCCCTGATGAGAACGCTAAGACCTACATCGCCAGCCAGGGCTGTCTGGA GGCCACGGTCAATGACTTCTGGCAGATGGCGTGGCAGGAGAACAGCCGTGTCATCGT CATGACCACCCGAGAGGTGGAGAAAGGCCGGAACAAATGCGTCCCATACTGGCCCGA GGTGGGCATGCAGCGTGCTTATGGGCCCTACTCTGTGACCAACTGCGGGGAGCATGA CACAACCGAATACAAACTCCGTACCTTACAGGTCTCCCCGCTGGACAATGGAGACCTG ATTCGGGAGATCTGGCATTACCAGTACCTGAGCTGGCCCGACCACGGGGTCCCCAGT GAGCCTGGGGGTGTCCTCAGCTTCCTGGACCAGATCAACCAGCGGCAGGAAAGTCTG CCTCACGCAGGGCCCATCATCGTGCACTGCAGCGCCGGCATCGGCCGCACAGGCACC ATCATTGTCATCGACATGCTCATGGAGAACATCTCCACCAAGGGCCTGGACTGTGACAT TGACATCCAGAAGACCATCCAGATGGTGCGGGCGCAGCGCTCGGGCATGGTGCAGAC GGAGGCGCAGTACAAGTTCATCTACGTGGCCATCGCCCAGTTCATTGAAACCACTAAG AAGAAGCTGTGA SEQ ID No. 39 >MP16091.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-LAIR1tm-end- o ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCC AGACCAGACATCCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGAC CGGGTGACCATCAGCTGCAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTACC AGCAGAAGCCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCCGGCTGCACA GCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGACTACAGCCTGACC ATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACACCC TGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAG GCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGT GAAGCTGCAGGAGTCTGGCCCAGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGAC CTGCACCGTGAGCGGCGTGAGCCTGCCCGACTACGGCGTGAGCTGGATCAGGCAGCC CCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGCGAGACCACCTACTA CAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCAGGT GTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAG CACTACTACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTG ACCGTGAGCTCAGATCCCACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCC ACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGG GGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTTTTGGGTGCTGGT GGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTTATTATTT TCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGG GCCAGAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTT GGACAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACC CTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGGAGGCCTACAGTGA GATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGCACGATGGCCTTTACCAGG GTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGGCCCTGCCTCC TCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCC CGGGCCCATGGCCGTGCCCACTCAGGTCCTGGGGTTGTTGCTACTGTGGCTTACAGAT GCCAGATGTGACATCCAGATGACACAGTCTCCATCTTCCCTGTCTGCATCTGTCGGAGA TCGCGTCACCATCACCTGTCGAGCAAGTGAGGACATTTATTTTAATTTAGTGTGGTATCA GCAGAAACCAGGAAAGGCCCCTAAGCTCCTGATCTATGATACAAATCGCTTGGCAGAT GGGGTCCCATCACGGTTCAGTGGCTCTGGATCTGGCACACAGTATACTCTAACCATAA GTAGCCTGCAACCCGAAGATTTCGCAACCTATTATTGTCAACACTATAAGAATTATCCGC TCACGTTCGGTCAGGGGACCAAGCTGGAAATCAAAAGATCTGGTGGCGGAGGGTCAG GAGGCGGAGGCAGCGGAGGCGGTGGCTCGGGAGGCGGAGGCTCGAGATCTGAGGTG CAGTTGGTGGAGTCTGGGGGCGGCTTGGTGCAGCCTGGAGGGTCCCTGAGGCTCTCC TGTGCAGCCTCAGGATTCACTCTCAGTAATTATGGCATGCACTGGATCAGGCAGGCTC CAGGGAAGGGTCTGGAGTGGGTCTCGTCTATTAGTCTTAATGGTGGTAGCACTTACTAT CGAGACTCCGTGAAGGGCCGATTCACTATCTCCAGGGACAATGCAAAAAGCACCCTCT ACCTTCAAATGAATAGTCTGAGGGCCGAGGACACGGCCGTCTATTACTGTGCAGCACA GGACGCTTATACGGGAGGTTACTTTGATTACTGGGGCCAAGGAACGCTGGTCACAGTC TCGTCTATGGATCCCGCCACCACAACCAAGCCCGTGCTGCGGACCCCAAGCCCTGTGC ACCCTACCGGCACCAGCCAGCCTCAGAGACCCGAGGACTGCCGGCCTCGGGGCAGC GTGAAGGGCACCGGCCTGGACTTCGCCTGCGACATTCTCATCGGGGTCTCAGTGGTCT TCCTCTTCTGTCTCCTCCTCCTGGTCCTCTTCTGCCTCCATCGCCAGAATCAGATAAAG CAGGGGCCCCCCAGAAGCAAGGACGAGGAGCAGAAGCCACAGCAGAGGCCTGACCT GGCTGTTGATGTTCTAGAGAGGACAGCAGACAAGGCCACAGTCAATGGACTTCCTGAG AAGGACCGGGAGACCGACACCAGCGCCCTGGCTGCAGGGAGTTCCCAGGAGGTGAC GTATGCTCAGCTGGACCACTGGGCCCTCACACAGAGGACAGCCCGGGCTGTGTCCCC ACAGTCCACAAAGCCCATGGCCGAGTCCATCACGTATGCAGCCGTTGCCAGACACTGA SEQ ID no. 40 >MP16092.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-LAIR1tm- endo-2A-PTPN6_SH2-dCD148 ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCC AGACCAGACATCCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGAC CGGGTGACCATCAGCTGCAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTACC AGCAGAAGCCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCCGGCTGCACA GCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGACTACAGCCTGACC ATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACACCC TGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAG GCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGT GAAGCTGCAGGAGTCTGGCCCAGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGAC CTGCACCGTGAGCGGCGTGAGCCTGCCCGACTACGGCGTGAGCTGGATCAGGCAGCC CCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGCGAGACCACCTACTA CAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCAGGT GTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAG CACTACTACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTG ACCGTGAGCTCAGATCCCACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCC ACCATCGCGTCGCAGCCCCTGTCCCTGCGCCCAGAGGCGTGCCGGCCAGCGGCGGG GGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATATCTTTTGGGTGCTGGT GGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTTATTATTT TCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGG GCCAGAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTT GGACAAGAGACGTGGCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACC CTCAGGAAGGCCTGTACAATGAACTGCAGAAAGATAAGATGGCGGAGGCCTACAGTGA GATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGGGCACGATGGCCTTTACCAGG GTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGGCCCTGCCTCC TCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCC CGGGCCCATGGCCGTGCCCACTCAGGTCCTGGGGTTGTTGCTACTGTGGCTTACAGAT GCCAGATGTGACATCCAGATGACACAGTCTCCATCTTCCCTGTCTGCATCTGTCGGAGA TCGCGTCACCATCACCTGTCGAGCAAGTGAGGACATTTATTTTAATTTAGTGTGGTATCA GCAGAAACCAGGAAAGGCCCCTAAGCTCCTGATCTATGATACAAATCGCTTGGCAGAT GGGGTCCCATCACGGTTCAGTGGCTCTGGATCTGGCACACAGTATACTCTAACCATAA GTAGCCTGCAACCCGAAGATTTCGCAACCTATTATTGTCAACACTATAAGAATTATCCGC TCACGTTCGGTCAGGGGACCAAGCTGGAAATCAAAAGATCTGGTGGCGGAGGGTCAG GAGGCGGAGGCAGCGGAGGCGGTGGCTCGGGAGGCGGAGGCTCGAGATCTGAGGTG CAGTTGGTGGAGTCTGGGGGCGGCTTGGTGCAGCCTGGAGGGTCCCTGAGGCTCTCC TGTGCAGCCTCAGGATTCACTCTCAGTAATTATGGCATGCACTGGATCAGGCAGGCTC CAGGGAAGGGTCTGGAGTGGGTCTCGTCTATTAGTCTTAATGGTGGTAGCACTTACTAT CGAGACTCCGTGAAGGGCCGATTCACTATCTCCAGGGACAATGCAAAAAGCACCCTCT ACCTTCAAATGAATAGTCTGAGGGCCGAGGACACGGCCGTCTATTACTGTGCAGCACA GGACGCTTATACGGGAGGTTACTTTGATTACTGGGGCCAAGGAACGCTGGTCACAGTC TCGTCTATGGATCCCGCCACCACAACCAAGCCCGTGCTGCGGACCCCAAGCCCTGTGC ACCCTACCGGCACCAGCCAGCCTCAGAGACCCGAGGACTGCCGGCCTCGGGGCAGC GTGAAGGGCACCGGCCTGGACTTCGCCTGCGACATTCTCATCGGGGTCTCAGTGGTCT TCCTCTTCTGTCTCCTCCTCCTGGTCCTCTTCTGCCTCCATCGCCAGAATCAGATAAAG
CAGGGGCCCCCCAGAAGCAAGGACGAGGAGCAGAAGCCACAGCAGAGGCCTGACCT GGCTGTTGATGTTCTAGAGAGGACAGCAGACAAGGCCACAGTCAATGGACTTCCTGAG AAGGACCGGGAGACCGACACCAGCGCCCTGGCTGCAGGGAGTTCCCAGGAGGTGAC GTATGCTCAGCTGGACCACTGGGCCCTCACACAGAGGACAGCCCGGGCTGTGTCCCC ACAGTCCACAAAGCCCATGGCCGAGTCCATCACGTATGCAGCCGTTGCCAGACACAGG GCAGAAGGAAGAGGTAGCCTGCTGACTTGCGGGGACGTGGAAGAGAACCCAGGGCCA TGGTATCATGGCCACATGTCTGGCGGGCAGGCAGAGACGCTGCTGCAGGCCAAGGGC GAGCCCTGGACGTTTCTTGTGCGTGAGAGCCTCAGCCAGCCTGGAGACTTCGTGCTTT CTGTGCTCAGTGACCAGCCCAAGGCTGGCCCAGGCTCCCCGCTCAGGGTCACCCACA TCAAGGTCATGTGCGAGGGTGGACGCTACACAGTGGGTGGTTTGGAGACCTTCGACAG CCTCACGGACCTGGTGGAGCATTTCAAGAAGACGGGGATTGAGGAGGCCTCAGGCGC CTTTGTCTACCTGCGGCAGCCGTACAGCGGTGGCGGTGGCAGCTTTGAGGCCTACTTC AAGAAGCAGCAAGCTGACTCCAACTGTGGGTTCGCAGAGGAATACGAAGATCTGAAGC TTGTTGGAATTAGTCAACCTAAATATGCAGCAGAACTGGCTGAGAATAGAGGAAAGAAT CGCTATAATAATGTTCTGCCCTATGATATTTCCCGTGTCAAACTTTCGGTCCAGACCCAT TCAACGGATGACTACATCAATGCCAACTACATGCCTGGCTACCACTCCAAGAAAGATTT TATTGCCACACAAGGACCTTTACCGAACACTTTGAAAGATTTTTGGCGTATGGTTTGGG AGAAAAATGTATATGCCATCATTATGTTGACTAAATGTGTTGAACAGGGAAGAACCAAAT GTGAGGAGTATTGGCCCTCCAAGCAGGCTCAGGACTATGGAGACATAACTGTGGCAAT GACATCAGAAATTGTTCTTCCGGAATGGACCATCAGAGATTTCACAGTGAAAAATATCC AGACAAGTGAGAGTCACCCTCTGAGACAGTTCCATTTCACCTCCTGGCCAGACCACGG TGTTCCCGACACCACTGACCTGCTCATCAACTTCCGGTACCTCGTTCGTGACTACATGA AGCAGAGTCCTCCCGAATCGCCGATTCTGGTGCATTGCAGTGCTGGGGTCGGAAGGA CGGGCACTTTCATTGCCATTGATCGTCTCATCTACCAGATAGAGAATGAGAACACCGTG GATGTGTATGGGATTGTGTATGACCTTCGAATGCATAGGCCTTTAATGGTGCAGACAGA GGACCAGTATGTTTTCCTCAATCAGTGTGTTTTGGATATTGTCAGATCCCAGAAAGACTC AAAAGTAGATCTTATCTACCAGAACACAACTGCAATGACAATCTATGAAAACCTTGCGCC CGTGACCACATTTGGAAAGACCAATGGTTACATCGCCAGCGGTAGCTAA
[0350] The nucleic acid sequence may encode the same amino acid sequence as that encoded by SEQ ID No. 35, 36, 37, 38, 39 or 40, but may have a different nucleic acid sequence, due to the degeneracy of the genetic code. The nucleic acid sequence may have at least 80, 85, 90, 95, 98 or 99% identity to the sequence shown as SEQ ID No. 35, 36, 37, 38, 39 or 40, provided that it encodes a first CAR and a second CAR as defined in the first aspect of the invention.
[0351] Vector
[0352] The present invention also provides a vector, or kit of vectors which comprises one or more CAR-encoding nucleic acid sequence(s). Such a vector may be used to introduce the nucleic acid sequence(s) into a host cell so that it expresses the first and second CARs.
[0353] The vector may, for example, be a plasmid or a viral vector, such as a retroviral vector or a lentiviral vector, or a transposon based vector or synthetic mRNA.
[0354] The vector may be capable of transfecting or transducing a T cell.
[0355] Pharmaceutical Composition
[0356] The present invention also relates to a pharmaceutical composition containing a plurality of CAR-expressing T cells or NK cells according to the first aspect of the invention. The pharmaceutical composition may additionally comprise a pharmaceutically acceptable carrier, diluent or excipient. The pharmaceutical composition may optionally comprise one or more further pharmaceutically active polypeptides and/or compounds. Such a formulation may, for example, be in a form suitable for intravenous infusion.
[0357] Method of Treatment
[0358] The T cells of the present invention may be capable of killing target cells, such as cancer cells. The target cell may be recognisable by a defined pattern of antigen expression, for example the expression of antigen A AND antigen B; the expression of antigen A OR antigen B; or the expression of antigen A AND NOT antigen B or complex iterations of these gates.
[0359] T cells of the present invention may be used for the treatment of an infection, such as a viral infection.
[0360] T cells of the invention may also be used for the control of pathogenic immune responses, for example in autoimmune diseases, allergies and graft-vs-host rejection.
[0361] T cells of the invention may be used for the treatment of a cancerous disease, such as bladder cancer, breast cancer, colon cancer, endometrial cancer, kidney cancer (renal cell), leukemia, lung cancer, melanoma, non-Hodgkin lymphoma, pancreatic cancer, prostate cancer and thyroid cancer.
[0362] It is particularly suited for treatment of solid tumours where the availability of good selective single targets is limited.
[0363] T cells of the invention may be used to treat: cancers of the oral cavity and pharynx which includes cancer of the tongue, mouth and pharynx; cancers of the digestive system which includes oesophageal, gastric and colorectal cancers; cancers of the liver and biliary tree which includes hepatocellular carcinomas and cholangiocarcinomas; cancers of the respiratory system which includes bronchogenic cancers and cancers of the larynx; cancers of bone and joints which includes osteosarcoma; cancers of the skin which includes melanoma; breast cancer; cancers of the genital tract which include uterine, ovarian and cervical cancer in women, prostate and testicular cancer in men; cancers of the renal tract which include renal cell carcinoma and transitional cell carcinomas of the utterers or bladder; brain cancers including gliomas, glioblastoma multiforme and medullobastomas; cancers of the endocrine system including thyroid cancer, adrenal carcinoma and cancers associated with multiple endocrine neoplasm syndromes; lymphomas including Hodgkin's lymphoma and non-Hodgkin lymphoma; Multiple Myeloma and plasmacytomas; leukaemias both acute and chronic, myeloid or lymphoid; and cancers of other and unspecified sites including neuroblastoma.
[0364] Treatment with the T cells of the invention may help prevent the escape or release of tumour cells which often occurs with standard approaches.
[0365] The invention will now be further described by way of Examples, which are meant to serve to assist one of ordinary skill in the art in carrying out the invention and are not intended in any way to limit the scope of the invention.
EXAMPLES
Example 1--Creation of Target Cell Populations
[0366] For the purposes of proving the principle of the invention, receptors based on anti-CD19 and anti-CD33 were arbitrarily chosen. Using retroviral vectors, CD19 and CD33 were cloned. These proteins were truncated so that they do not signal and could be stably expressed for prolonged periods. Next, these vectors were used to transduce the SupT1 cell line either singly or doubly to establish cells negative for both antigen (the wild-type), positive for either and positive for both. The expression data are shown in FIG. 3.
Example 2--Design and Function of the OR Gate
[0367] To construct the OR gate, a pair of receptors recognizing CD19 and CD33 were co-expressed. Different spacers were used to prevent cross-pairing. Both receptors had a trans-membrane domain derived from CD28 to improve surface stability and an endodomain derived from that of CD3 Zeta to provide a simple activating signal. In this way, a pair of independent 1.sup.st generation CARs were co-expressed. The retroviral vector cassette used to co-express the sequences utilizes a foot-and-mouth 2A self-cleaving peptide to allow co-expression 1:1 of both receptors. The cassette design is shown in FIG. 4, and the protein structures in FIG. 5. The nucleotide sequence of homologous regions was codon-wobbled to prevent recombination during retroviral vector reverse transcription.
Example 3--Testing the OR Gate
[0368] Expression of both CARs was tested on the T-cell surface by staining with cognate antigen fused to Fc. By using different species of Fc domains (mouse for CD19 and rabbit for CD33), co-expression of both CARs was determined on the cell surface by staining with different secondary antibodies conjugated with different fluorophores. This is shown in FIG. 6.
[0369] Functional testing was then carried out using the mouse T-cell line BW5147. This cell line releases IL2 upon activation allowing a simple quantitative readout. These T-cells were co-cultured with increasing amounts of the artificial target cells described above. T-cells responded to target cells expressing either antigen, as shown by IL2 release measured by ELISA. Both CARs were shown to be expressed on the cell surfaces and the T-cells were shown to respond to either or both antigens. These data are show in FIG. 7.
Example 4--Design and Function of the AND Gate
[0370] The AND gate combines a simple activating receptor with a receptor which basally inhibits activity, but whose inhibition is turned off once the receptor is ligated. This was achieved by combining a standard 1.sup.st generation CAR with a short/non-bulky CD8 stalk spacer and a CD3 Zeta endodomain with a second receptor with a bulky Fc spacer whose endodomain contained either CD148 or CD45 endodomains. When both receptors are ligated, the difference in spacer dimensions results in isolation of the different receptors in different membrane compartments, releasing the CD3 Zeta receptor from inhibition by the CD148 or CD45 endodomains. In this way, activation only occurs once both receptors are activated. CD148 and CD45 were chosen for this as they function in this manner natively: for instance, the very bulky CD45 ectodomain excludes the entire receptor from the immunological synapse. The expression cassette is depicted in FIG. 8 and the subsequent proteins in FIG. 9.
[0371] Surface staining for the different specificity showed that both receptor pairs could be effectively expressed on the cell surface shown in FIG. 10. Function in BW5147 shows that the T-cell is only activated in the presence of both antigens (FIG. 11).
Example 5: Demonstration of Generalizability of the AND Gate
[0372] To ensure that the observations were not a manifestation of some specific characteristic of CD19/CD33 and their binders which had been used, the two targeting scFvs were swapped such that now, the activation (ITAM) signal was transmitted upon recognition of CD33, rather than CD19; and the inhibitory (CD148) signal was transmitted upon recognition of CD19, rather than of CD33. Since CD45 and CD148 endodomains are considered to be functionally similar, experimentation was restricted to AND gates with CD148 endodomain. This should still result in a functional AND gate. T-cells expressing the new logic gate where challenged with targets bearing either CD19 or CD33 alone, or both. The T-cells responded to targets expressing both CD19 and CD33, but not to targets expressing only one or none of these antigens. This shows that the AND gate is still functional in this format (FIG. 18B).
[0373] On the same lines, it was sought to establish how generalizable our AND gate is: the AND gate should be generalizable across different targets. While there may be lesser or greater fidelity of the gate given relative antigen density, cognate scFv binding kinetics and precise distance of the scFv binding epitope, one would expect to see some AND gate manifestations with a wide set of targets and binders. To test this, three additional AND gates were generated. Once again, experimentation was restricted to the CD148 version of the AND gate. The second scFv from the original CD148 AND gate was replaced with the anti-GD2 scFv huK666 (SEQ ID 41 and SEQ ID 42), or with the anti-CD5 scFv (SEQ ID 43 and SEQ ID 44), or the anti-EGFRvIII scFv MR1.1 (SEQ ID 45 AND SEQ ID 46) to generate the following CAR AND gates: CD19 AND GD2; CD19 AND CD5; CD19 AND EGFRvIII. The following artificial antigen expressing cell lines were also generated: by transducing SupT1, and our SupT1.CD19 with GM3 and GD2 synthases SupT1.GD2 and SupT1.CD19.GD2 were generated. By transducing SupT1 and SupT1.CD19 with a retroviral vector coding for EGFRvIII SupT1.EGFRvII and SupT1.CD19.EGFRvII were generated. Since CD5 is expressed on SupT1 cells, a different cell line was used to generate the target cells: 293T cells were generated which express CD19 alone, CD5 alone and both CD5 and CD19 together. Expression was confirmed by flow-cytometry (FIG. 19). T-cells expressing the three new CAR AND gates were challenged with SupT1.CD19 and respective cognate double positive and single positive target cells. All three AND gates demonstrated reduced activation by the double positive cell lines in comparison with the single positive targets (FIG. 20). This demonstrates generalizability of the AND gate design to arbitrary targets and cognate binders.
Example 6: Experimental Proof of Kinetic Segregation Model of CAR AND Gate
[0374] The aim was to prove the model that differential segregation caused by different spacers is the central mechanism behind the ability to generate these logic CAR gates. The model is that if only the activating CAR is ligated, the potent inhibiting `ligation off` type CAR is in solution in the membrane and can inhibit the activating CAR. Once both CARs are ligated, if both CAR spacers are sufficiently different, they will segregate within the synapse and not co-localize. Hence, a key requirement is that the spacers are sufficiently different. If the model is correct, if both spacers are sufficiently similar so they co-localize when both receptors are ligated, the gate will fail to function. To test this, the "bulky" Fc spacer in the original CAR we replaced with a murine CD8 spacer. It was predicted that this has the similar length, bulk and charge as human CD8 but so should not cross-pair with it. Hence, the new gate had a first CAR which recognizes CD19, a human CD8 stalk spacer and an activatory endodomain; while the second CAR recognizes CD33, has a mouse CD8 stalk spacer and a CD148 endodomain (FIG. 18C). T-cells were transduced to express this new CAR gate. These T-cells were then challenged with SupT1 cells expressing CD19 alone, CD33 alone or CD19 and CD33 together. T-cells did not respond to SupT1 cells expressing either antigen alone as per the original AND gate. However, CAR T-cells failed to respond to SupT1 cells expressing both antigens, thereby confirming the model (FIG. 18C). A functional AND gate requires both CARs to have spacers sufficiently different so that they do not co-localize within an immunological synapse (FIGS. 23A and B).
Example 7--Design and Function of an AND NOT Gate
[0375] Phosphatases such as CD45 and CD148 are so potent that even a small amount entering an immunological synapse can inhibit ITAM activation. This is the basis of inhibition of the logical AND gate. Other classes of phosphatases are not as potent e.g. PTPN6 and related phosphatases. It was predicted that a small amount of PTPN6 entering a synapse by diffusion would not inhibit activation. In addition, it was predicted that if an inhibitory CAR had a sufficiently similar spacer to an activating CAR, it could co-localize within a synapse if both CARs were ligated. In this case, large amounts of the inhibitory endodomain would be sufficient to stop the ITAMS from activating when both antigens were present. In this way, an AND NOT gate could be created.
[0376] For the NOT AND gate, the second signal needs to "veto" activation. This is done by bringing an inhibitory signal into the immunological synapse, for example by bringing in the phosphatase of an enzyme such as PTPN6. We hence generated an initial AND NOT gate as follows: two CARs co-expressed whereby the first recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; co-expressed with an anti-CD33 CAR with a mouse CD8 stalk spacer and an endodomain comprising of the catalytic domain of PTPN6 (SEQ ID 38, FIG. 13 A with B). A suitable cassette is shown in FIG. 12 and preliminary functional data are shown in FIG. 14.
[0377] In addition, an alternative strategy was developed for generating an AND NOT gate. Immune Tyrosinase Inhibitory Motifs (ITIMs) are activated in a similar manner to ITAMS, in that they become phosphorylated by Ick upon clustering and exclusion of phosphatases. Instead of triggering activation by binding ZAP70, phosphorylated ITIMs recruit phosphatases like PTPN6 through their cognate SH2 domains. An ITIM can function as an inhibitory endodomain, as long as the spacers on the activating and inhibiting CARs can co-localize. To generate this construct, an AND NOT gate was generated as follows: two CARs co-expressed--the first recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; co-expressed with an anti-CD33 CAR with a mouse CD8 stalk spacer and an ITIM containing endodomain derived from that of LAIR1 (SEQ ID 39, FIG. 13 A with C).
[0378] A further, more complex AND NOT gate was also developed, whereby an ITIM is enhanced by the presence of an additional chimeric protein: an intracellular fusion of the SH2 domain of PTPN6 and the endodomain of CD148. In this design three proteins are expressed--the first recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; co-expressed with an anti-CD33 CAR with a mouse CD8 stalk spacer and an ITIM containing endodomain derived from that of LAIR1. A further 2A peptide, allows co-expression of the PTPN6-CD148 fusion (SEQ ID 40, FIGS. 13 A and D). It was predicted that these AND NOT gates would have a different range of inhibition: PTPN6-CD148>PTPN6>>ITIM.
[0379] T-cells were transduced with these gates and challenged with targets expressing either CD19 or CD33 alone, or both CD19 and CD33 together. All three gates responded to targets expressing only CD19, but not targets expressing both CD19 and CD33 together (FIG. 21), confirming that all three of the AND NOT gates were functional.
Example 8: Experimental Proof of Kinetic Segregation Model of PTPN6 Based AND NOT Gate
[0380] The model of the AND NOT gate centres around the fact that the nature of the spacers used in both CARs is pivotal for the correct function of the gate. In the functional AND NOT gate with PTPN6, both CAR spacers are sufficiently similar that when both CARs are ligated, both co-localize within the synapse so the high concentration even the weak PTPN6 is sufficient to inhibit activation. If the spacers were different, segregation in the synapse will isolate the PTPN6 from the ITAM allowing activation disrupting the AND NOT gate. To test this, a control was generated replacing the murine CD8 stalk spacer with that of Fc. In this case, the test gate consisted of two CARs, the first recognizes CD19, has a human CD8 stalk spacer and an ITAM endodomain; while the second CAR recognizes CD33, has an Fc spacer and an endodomain comprising of the phosphatase from PTPN6. This gate activates in response to CD19, but also activates in response to CD19 and CD33 together (FIG. 22B, where function of this gate is compared with that of the original AND NOT, and the control AND gate variant described in Example 6). This experimental data proves the model that for a functional AND NOT gate with PTPN6, co-localizing spacers are needed.
Example 9: Experimental Proof of Kinetic Segregation Model of ITIM Based AND NOT Gate
[0381] Similar to the PTPN6 based AND NOT gate, the ITIM based gate also requires co-localization in an immunological synapse to function as an AND NOT gate. To prove this hypothesis, a control ITIM based gate was generated as follows: two CARs co-expressed--the first recognizes CD19, has a human CD8 stalk spacer and an activating endodomain; co-expressed with an anti-CD33 CAR with an Fc spacer and an ITIM containing endodomain derived from that of LAIR1. The activity of this gate was compared with that of the original ITIM based AND NOT gate. In this case, the modified gate activated in response to targets expressing CD19, but also activated in response to cells expressing both CD19 and CD33. These data indicate that ITIM based AND NOT gates follow the kinetic segregation based model and a correct spacer must be selected to create a functional gate (FIG. 23B).
Example 10: Summary of Model of CAR Logic Gates Generated by Kinetic Segregation
[0382] Based on current understanding of the kinetic-segregation model and the experimental data described herein, a summary of the model for a two-CAR gate is presented in FIG. 24. The Figure shows a cell expressing two CARs, each recognizing a different antigen. When either or both CARs recognize a target antigen on a cell, a synapse forms and native CD45 and CD148 are excluded from the synapse due to the bulk of their ectodomain. This sets the stage for T-cell activation. In the case that the target cell bears only one cognate antigen, the cognate CAR is ligated and the cognate CAR segregates into the synapse. The unligated CAR remains in solution on the T-cell membrane and can diffuse in and out of the synapse so that an area of high local concentration of ligated CAR with low concentration of unligated CAR forms. In this case, if the ligated CAR has an ITAM and the non-ligated CAR has `ligation off" type inhibitory endodomain such as that of CD148, the amount of non-ligated CAR is sufficient to inhibit activation and the gate is off. In contrast, in this case, if the ligated CAR has an ITAM and the non-ligated CAR has a `ligation on` type inhibitory endodomain such as PTPN6, the amount of non-ligated CAR is insufficient to inhibit and the gate is on. When challenged by a target cell bearing both cognate antigens, both cognate CARs are ligated and form part of an immunological synapse. Importantly, if the CAR spacers are sufficiently similar, the CARs co-localize in the synapse but if the CAR spacers are sufficiently different the CARs segregate within the synapse. In this latter case, areas of membrane form whereby high concentrations of one CAR are present but the other CAR is absent. In this case since segregation is complete, even if the inhibitory endodomain is a `ligation off` type, the gate is on. In the former case, areas of membrane form with high concentrations of both CARs mixed together. In this case, since both endodomains are concentrated, even if the inhibitory endodomain is `ligation on` type, the gate is off. By selecting the correct combination of spacer and endodomain logic can be programmed into a CAR T-cell.
[0383] Based on our work above, we have established a series of design rules to allow generation of logic-gated CARs (illustrated in FIG. 31). To generate an "antigen A OR antigen B" gated CAR T-cell, anti-A and anti-B CARs must be generated such that (1) each CAR has a spacer which simply allows antigen access and synapse formation such that the CAR functions, and (2) Each CAR has an activating endodomain; To generate an "antigen A AND NOT B" gated CAR T-cell, anti-A and anti-B CARs must be generated such that (1) both CARs have spacers which do not cross-pair, but which will allow the CARs to co-segregate upon recognition of both cognate antigens on the target cell, (2) and one CAR has an activating endodomain, while the other CAR has an endodomain which comprises or recruits a weak phosphatase (e.g. PTPN6); (3) To generate an "antigen A AND antigen B" gated CAR T-cell, anti-A and anti-B CARs must be generated such that (1) one CAR has a spacer sufficiently different from the other CAR such that both CARs will not co-segregate upon recognition of both cognate antigens on the target cell, (2) one CAR has an activating endodomain, while the other car has an endodomain which comprises of a potent phosphatase (e.g. that of CD45 or CD148). The correct spacers to achieve the desired effect can be selected from a set of spacers with known size/shape etc as well as taking into consideration size/shape etc of the target antigen and the location of the cognate epitope on the target antigen.
TABLE-US-00021 SEQ ID No 41: SFG.aCD19-CD8STK-CD28tmZ-2A-aGD2-HCH2CH3pvaa-dCD148 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDISKYLNWYQQKPDGTVKLL IYHTSRLHSGVPSRFSGSGSGTDYSLTISNLEQEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSG GGGSGGGGSGGGGSEVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGVIWGS ETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYYYGGSYAMDYWGQGTSVTVSSDP TTTPAPRPPTPAPTIASQPLSLRPEACRPAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAF IIFWVRRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNEL QKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPRRAEGRGSLLTCGDVEENP GPMETDTLLLWVLLLWVPGSTGQVQLQESGPGLVKPSQTLSITCTVSGFSLASYNIHWVRQPPGKGLE WLGVIWAGGSTNYNSALMSRLTISKDNSKNQVFLKMSSLTAADTAVYYCAKRSDDYSWFAYWGQGTLV TVSSGGGGSGGGGSGGGGSENQMTQSPSSLSASVGDRVTMTCRASSSVSSSYLHWYQQKSGKAPKVWI YSTSNLASGVPSRFSGSGSGTDYTLTISSLQPEDFATYYCQQYSGYPITFGQGTKVEIKRSDPAEPKS PDKTHTCPPCPAPPVAGPSVFLFPPKPKDTLMIARTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRD ELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFS CSVMHEALHNHYTQKSLSLSPGKKDPKAVFGCIFGALVIVTVGGFIFWRKKRKDAKNNEVSFSQIKPK KSKLIRVENFEAYFKKQQADSNCGFAEEYEDLKLVGISQPKYAAELAENRGKNRYNNVLPYDISRVKL SVQTHSTDDYINANYMPGYHSKKDFIATQGPLPNTLKDFWRMVWEKNVYAIIMLTKCVEQGRTKCEEY WPSKQAQDYGDITVAMTSEIVLPEWTIRDFTVKNIQTSESHPLRQFHFTSWPDHGVPDTTDLLINFRY LVRDYMKQSPPESPILVHCSAGVGRTGTFIAIDRLIYQIENENTVDVYGIVYDLRMHRPLMVQTEDQY VFLNQCVLDIVRSQKDSKVDLIYQNTTAMTIYENLAPVTTFGKTNGYIA SEQ ID No. 42: SFG.aCD19-CD8STK-CD28tmZ-2A-aGD2-HCH2CH3pvaa-dCD148 ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCCAGACCAGACAT CCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGACCGGGTGACCATCAGCTGCAGAG CCAGCCAGGACATCAGCAAGTACCTGAACTGGTACCAGCAGAAGCCCGACGGCACCGTGAAGCTGCTG ATCTACCACACCAGCCGGCTGCACAGCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGA CTACAGCCTGACCATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACA CCCTGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAGGCTCTGGC GGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGTGAAGCTGCAGGAGTCTGGCCC AGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGACCTGCACCGTGAGCGGCGTGAGCCTGCCCGACT ACGGCGTGAGCTGGATCAGGCAGCCCCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGC GAGACCACCTACTACAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCA GGTGTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAGCACTACT ACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTGACCGTGAGCTCAGATCCC ACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCCACCATCGCGTCGCAGCCCCTGTCCCTGCG CCCAGAGGCGTGCCGGCCAGCGGCGGGGGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATA TCTTTTGGGTGCTGGTGGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTT ATTATTTTCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGGGCCA GAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTTGGACAAGAGACGTG GCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACCCTCAGGAAGGCCTGTACAATGAACTG CAGAAAGATAAGATGGCGGAGGCCTACAGTGAGATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGG GCACGATGGCCTTTACCAGGGTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGG CCCTGCCTCCTCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCC GGGCCCATGGAGACCGACACCCTGCTGCTGTGGGTGCTGCTGCTGTGGGTGCCAGGCAGCACCGGCCA GGTGCAGCTGCAGGAGTCTGGCCCAGGCCTGGTGAAGCCCAGCCAGACCCTGAGCATCACCTGCACCG TGAGCGGCTTCAGCCTGGCCAGCTACAACATCCACTGGGTGCGGCAGCCCCCAGGCAAGGGCCTGGAG TGGCTGGGCGTGATCTGGGCTGGCGGCAGCACCAACTACAACAGCGCCCTGATGAGCCGGCTGACCAT CAGCAAGGACAACAGCAAGAACCAGGTGTTCCTGAAGATGAGCAGCCTGACAGCCGCCGACACCGCCG TGTACTACTGCGCCAAGCGGAGCGACGACTACAGCTGGTTCGCCTACTGGGGCCAGGGCACCCTGGTG ACCGTGAGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGAACCAGAT GACCCAGAGCCCCAGCAGCTTGAGCGCCAGCGTGGGCGACCGGGTGACCATGACCTGCAGAGCCAGCA GCAGCGTGAGCAGCAGCTACCTGCACTGGTACCAGCAGAAGAGCGGCAAGGCCCCAAAGGTGTGGATC TACAGCACCAGCAACCTGGCCAGCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGACTA CACCCTGACCATCAGCAGCCTGCAGCCCGAGGACTTCGCCACCTACTACTGCCAGCAGTACAGCGGCT ACCCCATCACCTTCGGCCAGGGCACCAAGGTGGAGATCAAGCGGTCGGATCCCGCCGAGCCCAAATCT CCTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTCCCGTGGCCGGCCCGTCAGTCTTCCTCTT CCCCCCAAAACCCAAGGACACCCTCATGATCGCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACG TGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAG ACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGTGTGGTCAGCGTCCTCACCGTCCTGCACCA GGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCATCGAGA AAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGGGAT GAGCTGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGT GGAGTGGGAGAGCAATGGGCAACCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACG GCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCA TGCTCCGTGATGCATGAGGCCCTGCACAATCACTATACCCAGAAATCTCTGAGTCTGAGCCCAGGCAA GAAGGACCCCAAGGCGGTTTTTGGCTGTATCTTTGGTGCCCTGGTTATTGTGACTGTGGGAGGCTTCA TCTTCTGGAGAAAGAAGAGGAAAGATGCAAAGAATAATGAAGTGTCCTTTTCTCAAATTAAACCTAAA AAATCTAAGTTAATCAGAGTGGAGAATTTTGAGGCCTACTTCAAGAAGCAGCAAGCTGACTCCAACTG TGGGTTCGCAGAGGAATACGAAGATCTGAAGCTTGTTGGAATTAGTCAACCTAAATATGCAGCAGAAC TGGCTGAGAATAGAGGAAAGAATCGCTATAATAATGTTCTGCCCTATGATATTTCCCGTGTCAAACTT TCGGTCCAGACCCATTCAACGGATGACTACATCAATGCCAACTACATGCCTGGCTACCACTCCAAGAA AGATTTTATTGCCACACAAGGACCTTTACCGAACACTTTGAAAGATTTTTGGCGTATGGTTTGGGAGA AAAATGTATATGCCATCATTATGTTGACTAAATGTGTTGAACAGGGAAGAACCAAATGTGAGGAGTAT TGGCCCTCCAAGCAGGCTCAGGACTATGGAGACATAACTGTGGCAATGACATCAGAAATTGTTCTTCC GGAATGGACCATCAGAGATTTCACAGTGAAAAATATCCAGACAAGTGAGAGTCACCCTCTGAGACAGT TCCATTTCACCTCCTGGCCAGACCACGGTGTTCCCGACACCACTGACCTGCTCATCAACTTCCGGTAC CTCGTTCGTGACTACATGAAGCAGAGTCCTCCCGAATCGCCGATTCTGGTGCATTGCAGTGCTGGGGT CGGAAGGACGGGCACTTTCATTGCCATTGATCGTCTCATCTACCAGATAGAGAATGAGAACACCGTGG ATGTGTATGGGATTGTGTATGACCTTCGAATGCATAGGCCTTTAATGGTGCAGACAGAGGACCAGTAT GTTTTCCTCAATCAGTGTGTTTTGGATATTGTCAGATCCCAGAAAGACTCAAAAGTAGATCTTATCTA CCAGAACACAACTGCAATGACAATCTATGAAAACCTTGCGCCCGTGACCACATTTGGAAAGACCAATG GTTACATCGCCTAA SEQ ID No. 43: SFG.aCD19-CD8STK-CD28tmZ-2A-aCD5-HCH2CH3pvaa-dCD148 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDISKYLNWYQQKPDGTVKLL IYHTSRLHSGVPSRFSGSGSGTDYSLTISNLEQEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSG GGGSGGGGSGGGGSEVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGVIWGS ETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYYYGGSYAMDYWGQGTSVTVSSDP TTTPAPRPPTPAPTIASQPLSLRPEACRPAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAF IIFWVRRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNEL QKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPRRAEGRGSLLTCGDVEENP GPMETDTLLLWVLLLWVPGSTGQVTLKESGPGILKPSQTLSLTCSFSGFSLSTSGMGVGWIRQPSGKG LEWLAHIWWDDDVYYNPSLKNQLTISKDASRDQVFLKITNLDTADTATYYCVRRRATGTGFDYWGQGT TLTVSSGGGGSGGGGSGGGGSNIVMTQSHKFMSTSVGDRVSIACKASQDVGTAVAWYQQKPGQSPKLL IYWTSTRHTGVPDRFTGSGSGTDFTLTITNVQSEDLADYFCHQYNSYNTFGSGTRLELKRSDPAEPKS PDKTHTCPPCPAPPVAGPSVFLFPPKPKDTLMIARTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRD ELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFS CSVMHEALHNHYTQKSLSLSPGKKDPKAVFGCIFGALVIVTVGGFIFWRKKRKDAKNNEVSFSQIKPK KSKLIRVENFEAYFKKQQADSNCGFAEEYEDLKLVGISQPKYAAELAENRGKNRYNNVLPYDISRVKL SVQTHSTDDYINANYMPGYHSKKDFIATQGPLPNTLKDFWRMVWEKNVYAIIMLTKCVEQGRTKCEEY WPSKQAQDYGDITVAMTSEIVLPEWTIRDFTVKNIQTSESHPLRQFHFTSWPDHGVPDTTDLLINFRY LVRDYMKQSPPESPILVHCSAGVGRTGTFIAIDRLIYQIENENTVDVYGIVYDLRMHRPLMVQTEDQY VFLNQCVLDIVRSQKDSKVDLIYQNTTAMTIYENLAPVTTFGKTNGYIA SEQ ID No. 44: SFG.aCD19-CD8STK-CD28tmZ-2A-aCD5-HCH2CH3pvaa-dCD148 ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCCAGACCAGACAT CCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGACCGGGTGACCATCAGCTGCAGAG CCAGCCAGGACATCAGCAAGTACCTGAACTGGTACCAGCAGAAGCCCGACGGCACCGTGAAGCTGCTG ATCTACCACACCAGCCGGCTGCACAGCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGA CTACAGCCTGACCATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACA CCCTGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAGGCTCTGGC GGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGTGAAGCTGCAGGAGTCTGGCCC AGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGACCTGCACCGTGAGCGGCGTGAGCCTGCCCGACT ACGGCGTGAGCTGGATCAGGCAGCCCCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGC GAGACCACCTACTACAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCA GGTGTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAGCACTACT ACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTGACCGTGAGCTCAGATCCC ACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCCACCATCGCGTCGCAGCCCCTGTCCCTGCG CCCAGAGGCGTGCCGGCCAGCGGCGGGGGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATA TCTTTTGGGTGCTGGTGGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTT ATTATTTTCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGGGCCA GAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTTGGACAAGAGACGTG GCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACCCTCAGGAAGGCCTGTACAATGAACTG CAGAAAGATAAGATGGCGGAGGCCTACAGTGAGATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGG GCACGATGGCCTTTACCAGGGTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGG CCCTGCCTCCTCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCC GGGCCCATGGAGACCGACACCCTGCTGCTGTGGGTGCTGCTGCTGTGGGTGCCCGGCAGCACCGGCCA GGTGACCCTGAAGGAGAGCGGTCCCGGCATCCTGAAGCCCAGCCAGACCCTGAGCCTGACCTGCAGCT
TCAGCGGCTTCAGCCTGAGCACCAGCGGCATGGGCGTGGGCTGGATTCGGCAGCCCAGCGGCAAGGGC CTGGAGTGGCTGGCCCACATCTGGTGGGACGACGACGTGTACTACAACCCCAGCCTGAAGAACCAGCT GACCATCAGCAAGGACGCCAGCCGGGACCAGGTGTTCCTGAAGATCACCAACCTGGACACCGCCGACA CCGCCACCTACTACTGCGTGCGGCGCCGGGCCACCGGCACCGGCTTCGACTACTGGGGCCAGGGCACC ACCCTGACCGTGAGCAGCGGTGGCGGTGGCAGCGGCGGCGGCGGAAGCGGAGGTGGTGGCAGCAACAT CGTGATGACCCAGAGCCACAAGTTCATGAGCACCAGCGTGGGCGACCGGGTGAGCATCGCCTGCAAGG CCAGCCAGGACGTGGGCACCGCCGTGGCCTGGTACCAGCAGAAGCCTGGCCAGAGCCCCAAGCTGCTG ATCTACTGGACCAGCACCCGGCACACCGGCGTGCCCGACCGGTTCACCGGCAGCGGCAGCGGCACCGA CTTCACCCTGACCATCACCAACGTGCAGAGCGAGGACCTGGCCGACTACTTCTGCCACCAGTACAACA GCTACAACACCTTCGGCAGCGGCACCCGGCTGGAGCTGAAGCGGTCGGATCCCGCCGAGCCCAAATCT CCTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTCCCGTGGCCGGCCCGTCAGTCTTCCTCTT CCCCCCAAAACCCAAGGACACCCTCATGATCGCCCGGACCCCTGAGGTCACATGCGTGGTGGTGGACG TGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCCAAG ACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGTGTGGTCAGCGTCCTCACCGTCCTGCACCA GGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCATCGAGA AAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGGGAT GAGCTGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGCCGT GGAGTGGGAGAGCAATGGGCAACCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCGACG GCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTCTCA TGCTCCGTGATGCATGAGGCCCTGCACAATCACTATACCCAGAAATCTCTGAGTCTGAGCCCAGGCAA GAAGGACCCCAAGGCGGTTTTTGGCTGTATCTTTGGTGCCCTGGTTATTGTGACTGTGGGAGGCTTCA TCTTCTGGAGAAAGAAGAGGAAAGATGCAAAGAATAATGAAGTGTCCTTTTCTCAAATTAAACCTAAA AAATCTAAGTTAATCAGAGTGGAGAATTTTGAGGCCTACTTCAAGAAGCAGCAAGCTGACTCCAACTG TGGGTTCGCAGAGGAATACGAAGATCTGAAGCTTGTTGGAATTAGTCAACCTAAATATGCAGCAGAAC TGGCTGAGAATAGAGGAAAGAATCGCTATAATAATGTTCTGCCCTATGATATTTCCCGTGTCAAACTT TCGGTCCAGACCCATTCAACGGATGACTACATCAATGCCAACTACATGCCTGGCTACCACTCCAAGAA AGATTTTATTGCCACACAAGGACCTTTACCGAACACTTTGAAAGATTTTTGGCGTATGGTTTGGGAGA AAAATGTATATGCCATCATTATGTTGACTAAATGTGTTGAACAGGGAAGAACCAAATGTGAGGAGTAT TGGCCCTCCAAGCAGGCTCAGGACTATGGAGACATAACTGTGGCAATGACATCAGAAATTGTTCTTCC GGAATGGACCATCAGAGATTTCACAGTGAAAAATATCCAGACAAGTGAGAGTCACCCTCTGAGACAGT TCCATTTCACCTCCTGGCCAGACCACGGTGTTCCCGACACCACTGACCTGCTCATCAACTTCCGGTAC CTCGTTCGTGACTACATGAAGCAGAGTCCTCCCGAATCGCCGATTCTGGTGCATTGCAGTGCTGGGGT CGGAAGGACGGGCACTTTCATTGCCATTGATCGTCTCATCTACCAGATAGAGAATGAGAACACCGTGG ATGTGTATGGGATTGTGTATGACCTTCGAATGCATAGGCCTTTAATGGTGCAGACAGAGGACCAGTAT GTTTTCCTCAATCAGTGTGTTTTGGATATTGTCAGATCCCAGAAAGACTCAAAAGTAGATCTTATCTA CCAGAACACAACTGCAATGACAATCTATGAAAACCTTGCGCCCGTGACCACATTTGGAAAGACCAATG GTTACATCGCCTAA SEQ ID No. 45: SFG.aCD19-CD8STK-CD28tmZ-2A-aEGFRvIII-HCH2CH3pvaa- dCD148 MSLPVTALLLPLALLLHAARPDIQMTQTTSSLSASLGDRVTISCRASQDISKYLNWYQQKPDGTVKLL IYHTSRLHSGVPSRFSGSGSGTDYSLTISNLEQEDIATYFCQQGNTLPYTFGGGTKLEITKAGGGGSG GGGSGGGGSGGGGSEVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGVIWGS ETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYYYGGSYAMDYWGQGTSVTVSSDP TTTPAPRPPTPAPTIASQPLSLRPEACRPAAGGAVHTRGLDFACDIFWVLVVVGGVLACYSLLVTVAF IIFWVRRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNEL QKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPRRAEGRGSLLTCGDVEENP GPMETDTLLLWVLLLWVPGSTGQVKLQQSGGGLVKPGASLKLSCVTSGFTFRKFGMSWVRQTSDKRLE WVASISTGGYNTYYSDNVKGRFTISRENAKNTLYLQMSSLKSEDTALYYCTRGYSSTSYAMDYWGQGT TVTVSSGGGGSGGGGSGGGGSDIELTQSPASLSVATGEKVTIRCMTSTDIDDDMNWYQQKPGEPPKFL ISEGNTLRPGVPSRFSSSGTGTDFVFTIENTLSEDVGDYYCLQSFNVPLTFGDGTKLEIKRSDPAEPK SPDKTHTCPPCPAPPVAGPSVFLFPPKPKDTLMIARTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNA KTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSR DELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVF SCSVMHEALHNHYTQKSLSLSPGKKDPKAVFGCIFGALVIVTVGGFIFWRKKRKDAKNNEVSFSQIKP KKSKLIRVENFEAYFKKQQADSNCGFAEEYEDLKLVGISQPKYAAELAENRGKNRYNNVLPYDISRVK LSVQTHSTDDYINANYMPGYHSKKDFIATQGPLPNTLKDFWRMVWEKNVYAIIMLTKCVEQGRTKCEE YWPSKQAQDYGDITVAMTSEIVLPEWTIRDFTVKNIQTSESHPLRQFHFTSWPDHGVPDTTDLLINFR YLVRDYMKQSPPESPILVHCSAGVGRTGTFIAIDRLIYQIENENTVDVYGIVYDLRMHRPLMVQTEDQ YVFLNQCVLDIVRSQKDSKVDLIYQNTTAMTIYENLAPVTTFGKTNGYIA SEQ ID No. 46: SFG.aCD19-CD8STK-CD28tmZ-2A-aEGFRvIII-HCH2CH3pvaa- dCD148 ATGAGCCTGCCCGTGACCGCCCTGCTGCTGCCCCTGGCCCTGCTGCTGCACGCCGCCAGACCAGACAT CCAGATGACCCAGACCACCAGCAGCCTGAGCGCCAGCCTGGGCGACCGGGTGACCATCAGCTGCAGAG CCAGCCAGGACATCAGCAAGTACCTGAACTGGTACCAGCAGAAGCCCGACGGCACCGTGAAGCTGCTG ATCTACCACACCAGCCGGCTGCACAGCGGCGTGCCCAGCCGGTTCAGCGGCAGCGGCAGCGGCACCGA CTACAGCCTGACCATCAGCAACCTGGAGCAGGAGGACATCGCCACCTACTTCTGCCAGCAGGGCAACA CCCTGCCCTACACCTTCGGAGGCGGCACCAAGCTGGAGATCACCAAGGCCGGAGGCGGAGGCTCTGGC GGAGGCGGCTCTGGCGGAGGCGGCTCTGGCGGAGGCGGCAGCGAGGTGAAGCTGCAGGAGTCTGGCCC AGGCCTGGTGGCCCCAAGCCAGAGCCTGAGCGTGACCTGCACCGTGAGCGGCGTGAGCCTGCCCGACT ACGGCGTGAGCTGGATCAGGCAGCCCCCACGGAAGGGCCTGGAGTGGCTGGGCGTGATCTGGGGCAGC GAGACCACCTACTACAACAGCGCCCTGAAGAGCCGGCTGACCATCATCAAGGACAACAGCAAGAGCCA GGTGTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTACTGCGCCAAGCACTACT ACTATGGCGGCAGCTACGCTATGGACTACTGGGGCCAGGGCACCAGCGTGACCGTGAGCTCAGATCCC ACCACGACGCCAGCGCCGCGACCACCAACACCGGCGCCCACCATCGCGTCGCAGCCCCTGTCCCTGCG CCCAGAGGCGTGCCGGCCAGCGGCGGGGGGCGCAGTGCACACGAGGGGGCTGGACTTCGCCTGTGATA TCTTTTGGGTGCTGGTGGTGGTTGGTGGAGTCCTGGCTTGCTATAGCTTGCTAGTAACAGTGGCCTTT ATTATTTTCTGGGTGAGGAGAGTGAAGTTCAGCAGGAGCGCAGACGCCCCCGCGTACCAGCAGGGCCA GAACCAGCTCTATAACGAGCTCAATCTAGGACGAAGAGAGGAGTACGATGTTTTGGACAAGAGACGTG GCCGGGACCCTGAGATGGGGGGAAAGCCGAGAAGGAAGAACCCTCAGGAAGGCCTGTACAATGAACTG CAGAAAGATAAGATGGCGGAGGCCTACAGTGAGATTGGGATGAAAGGCGAGCGCCGGAGGGGCAAGGG GCACGATGGCCTTTACCAGGGTCTCAGTACAGCCACCAAGGACACCTACGACGCCCTTCACATGCAGG CCCTGCCTCCTCGCAGAGCCGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCC GGGCCCATGGAGACCGACACCCTGCTGCTGTGGGTGCTGCTGCTGTGGGTGCCCGGCAGCACCGGCCA GGTGAAGCTGCAGCAGAGCGGCGGAGGCCTGGTGAAGCCCGGCGCCAGCCTGAAGCTGAGCTGCGTGA CCAGCGGCTTCACCTTCCGGAAGTTCGGCATGAGCTGGGTGCGGCAGACCAGCGACAAGCGGCTGGAG TGGGTGGCCAGCATCAGCACCGGCGGCTACAACACCTACTACAGCGACAACGTGAAGGGCCGGTTCAC CATCAGCCGGGAGAACGCCAAGAACACCCTGTACCTGCAGATGAGCAGCCTGAAGAGCGAGGACACCG CCCTGTACTACTGCACCCGGGGCTACAGCAGCACCAGCTACGCTATGGACTACTGGGGCCAGGGCACC ACCGTGACAGTGAGCAGCGGCGGAGGAGGCAGTGGTGGGGGTGGATCTGGCGGAGGTGGCAGCGACAT CGAGCTGACCCAGAGCCCCGCCAGCCTGAGCGTGGCCACCGGCGAGAAGGTGACCATCCGGTGCATGA CCAGCACCGACATCGACGACGACATGAACTGGTACCAGCAGAAGCCCGGCGAGCCCCCAAAGTTCCTG ATCAGCGAGGGCAACACCCTGCGGCCCGGCGTGCCCAGCCGGTTCAGCAGCAGCGGCACCGGCACCGA CTTCGTGTTCACCATCGAGAACACCCTGAGCGAGGACGTGGGCGACTACTACTGCCTGCAGAGCTTCA ACGTGCCCCTGACCTTCGGCGACGGCACCAAGCTGGAGATCAAGCGGTCGGATCCCGCCGAGCCCAAA TCTCCTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTCCCGTGGCCGGCCCGTCAGTCTTCCT CTTCCCCCCAAAACCCAAGGACACCCTCATGATCGCCCGGACCCCTGAGGTCACATGCGTGGTGGTGG ACGTGAGCCACGAAGACCCTGAGGTCAAGTTCAACTGGTACGTGGACGGCGTGGAGGTGCATAATGCC AAGACAAAGCCGCGGGAGGAGCAGTACAACAGCACGTACCGTGTGGTCAGCGTCCTCACCGTCCTGCA CCAGGACTGGCTGAATGGCAAGGAGTACAAGTGCAAGGTCTCCAACAAAGCCCTCCCAGCCCCCATCG AGAAAACCATCTCCAAAGCCAAAGGGCAGCCCCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGG GATGAGCTGACCAAGAACCAGGTCAGCCTGACCTGCCTGGTCAAAGGCTTCTATCCCAGCGACATCGC CGTGGAGTGGGAGAGCAATGGGCAACCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGACTCCG ACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAGGGGAACGTCTTC TCATGCTCCGTGATGCATGAGGCCCTGCACAATCACTATACCCAGAAATCTCTGAGTCTGAGCCCAGG CAAGAAGGACCCCAAGGCGGTTTTTGGCTGTATCTTTGGTGCCCTGGTTATTGTGACTGTGGGAGGCT TCATCTTCTGGAGAAAGAAGAGGAAAGATGCAAAGAATAATGAAGTGTCCTTTTCTCAAATTAAACCT AAAAAATCTAAGTTAATCAGAGTGGAGAATTTTGAGGCCTACTTCAAGAAGCAGCAAGCTGACTCCAA CTGTGGGTTCGCAGAGGAATACGAAGATCTGAAGCTTGTTGGAATTAGTCAACCTAAATATGCAGCAG AACTGGCTGAGAATAGAGGAAAGAATCGCTATAATAATGTTCTGCCCTATGATATTTCCCGTGTCAAA CTTTCGGTCCAGACCCATTCAACGGATGACTACATCAATGCCAACTACATGCCTGGCTACCACTCCAA GAAAGATTTTATTGCCACACAAGGACCTTTACCGAACACTTTGAAAGATTTTTGGCGTATGGTTTGGG AGAAAAATGTATATGCCATCATTATGTTGACTAAATGTGTTGAACAGGGAAGAACCAAATGTGAGGAG TATTGGCCCTCCAAGCAGGCTCAGGACTATGGAGACATAACTGTGGCAATGACATCAGAAATTGTTCT TCCGGAATGGACCATCAGAGATTTCACAGTGAAAAATATCCAGACAAGTGAGAGTCACCCTCTGAGAC AGTTCCATTTCACCTCCTGGCCAGACCACGGTGTTCCCGACACCACTGACCTGCTCATCAACTTCCGG TACCTCGTTCGTGACTACATGAAGCAGAGTCCTCCCGAATCGCCGATTCTGGTGCATTGCAGTGCTGG GGTCGGAAGGACGGGCACTTTCATTGCCATTGATCGTCTCATCTACCAGATAGAGAATGAGAACACCG TGGATGTGTATGGGATTGTGTATGACCTTCGAATGCATAGGCCTTTAATGGTGCAGACAGAGGACCAG TATGTTTTCCTCAATCAGTGTGTTTTGGATATTGTCAGATCCCAGAAAGACTCAAAAGTAGATCTTAT CTACCAGAACACAACTGCAATGACAATCTATGAAAACCTTGCGCCCGTGACCACATTTGGAAAGACCA ATGGTTACATCGCCTAA
Example 11: Design and Construction of APRIL Based CARs
[0384] APRIL in its natural form is a secreted type II protein. The use of APRIL as a BCMA binding domain for a CAR requires conversion of this type II secreted protein to a type I membrane bound protein and for this protein to be stable and to retain binding to BCMA in this form. To generate candidate molecules, the extreme amino-terminus of APRIL was deleted to remove binding to proteoglycans. Next, a signal peptide was added to direct the nascent protein to the endoplasmic reticulum and hence the cell surface. Also, because the nature of spacer used can alter the function of a CAR, three different spacer domains were tested: an APRIL based CAR was generated comprising (i) a human IgG1 spacer altered to remove Fc binding motifs; (ii) a CD8 stalk; and (iii) the IgG1 hinge alone (cartoon in FIG. 25 and amino acid sequences in FIG. 26). These CARs were expressed in a bicistronic retroviral vector (FIG. 27A) so that a marker protein--truncated CD34 could be co-expressed as a convenient marker gene.
Example 12: Expression and Function of APRIL Based CARs
[0385] The aim of this study was to test whether the APRIL based CARs which had been constructed were expressed on the cell surface and whether APRIL had folded to form the native protein. T-cells were transduced with these different CAR constructs and stained using a commercially available anti-APRIL mAb, along with staining for the marker gene and analysed by flow-cytometry. The results of this experiment are shown in FIG. 27B where APRIL binding is plotting against marker gene fluorescence. These data show that in this format, the APRIL based CARs are expressed on the cell surface and APRIL folds sufficiently to be recognized by an anti-APRIL mAb.
[0386] Next, it was determined whether APRIL in this format could recognize BCMA and TACI. Recombinant BCMA and TACI were generated as fusions with mouse IgG2a-Fc. These recombinant proteins were incubated with the transduced T-cells. After this, the cells were washed and stained with an anti-mouse fluorophore conjugated antibody and an antibody to detect the marker gene conjugated to a different fluorophore. The cells were analysed by flow cytometry and the results are presented in FIG. 27C. The different CARs were able to bind both BCMA and TACI. Surprisingly, the CARs were better able to bind BCMA than TACI. Also, surprisingly CARs with a CD8 stalk or IgG1 hinge spacer were better able to bind BCMA and TACI than CAR with an Fc spacer.
Example 13: APRIL Based Chimeric Antigen Receptors are Active Against BCMA Expressing Cells
[0387] T-cells from normal donors were transduced with the different APRIL CARs and tested against SupT1 cells either wild-type, or engineered to express BCMA and TACI. Several different assays were used to determine function. A classical chromium release assay was performed. Here, the target cells (the SupT1 cells) were labelled with 51Cr and mixed with effectors (the transduced T-cells) at different ratio. Lysis of target cells was determined by counting 51Cr in the co-culture supernatant (FIG. 28A shows the cumulative data).
[0388] In addition, supernatant from T-cells cultured 1:1 with SupT1 cells was assayed by ELISA for Interferon-gamma (FIG. 28B shows cumulative data). Measurement of T-cell expansion after one week of co-culture with SupT1 cells was also performed (FIG. 28C). T-cells were counted by flow-cytometry calibrated with counting beads. These experimental data show that APRIL based CARs can kill BCMA expressing targets. Further, these data show that CARs based on the CD8 stalk or IgG1 hinge performed better than the Fc-pvaa based CAR.
Example 14: Functional Analysis of the AND Gate in Primary Cells
[0389] PBMCs were isolated from blood and stimulated using PHA and IL-2. Two days later the cells were transduced on retronectin coated plates with retro virus containing the CD19:CD33 AND gate construct. On day 5 the expression level of the two CARs translated by the AND gate construct was evaluated via flow cytometry and the cells were depleted of CD56+ cells (predominantly NK cells). On day 6 the PBMCs were placed in a co-culture with target cells at a 1:2 effector to target cell ratio. On day 8 the supernatant was collected and analysed for IFN-gamma secretion via ELISA (FIG. 29).
[0390] These data demonstrate that the AND gate functions in primary cells.
[0391] All publications mentioned in the above specification are herein incorporated by reference. Various modifications and variations of the described methods and system of the invention will be apparent to those skilled in the art without departing from the scope and spirit of the invention. Although the invention has been described in connection with specific preferred embodiments, it should be understood that the invention as claimed should not be unduly limited to such specific embodiments. Indeed, various modifications of the described modes for carrying out the invention which are obvious to those skilled in molecular biology, cell biology or related fields are intended to be within the scope of the following claims.
Sequence CWU 1
1
5611129PRTArtificial SequenceChimeric antigen receptor (CAR) 1Met
Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1 5 10
15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser Leu
20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg Ala
Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln Lys
Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His Thr Ser Arg
Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly Ser
Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln Glu
Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu Pro
Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys Ala
Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135 140
Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly 145
150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr Val
Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile Arg
Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile Trp
Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser Arg
Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe Leu
Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile Tyr
Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255 Met
Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260 265
270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala
275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala
Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala Cys
Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly Gly Val Leu
Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe Ile Ile Phe
Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala Asp Ala Pro
Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365 Glu Leu Asn
Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370 375 380 Arg
Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro 385 390
395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu
Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly
Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr
Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala Leu Pro Pro
Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr Cys Gly Asp
Val Glu Glu Asn Pro Gly Pro Met Ala 465 470 475 480 Val Pro Thr Gln
Val Leu Gly Leu Leu Leu Leu Trp Leu Thr Asp Ala 485 490 495 Arg Cys
Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser 500 505 510
Val Gly Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu Asp Ile Tyr 515
520 525 Phe Asn Leu Val Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys
Leu 530 535 540 Leu Ile Tyr Asp Thr Asn Arg Leu Ala Asp Gly Val Pro
Ser Arg Phe 545 550 555 560 Ser Gly Ser Gly Ser Gly Thr Gln Tyr Thr
Leu Thr Ile Ser Ser Leu 565 570 575 Gln Pro Glu Asp Phe Ala Thr Tyr
Tyr Cys Gln His Tyr Lys Asn Tyr 580 585 590 Pro Leu Thr Phe Gly Gln
Gly Thr Lys Leu Glu Ile Lys Arg Ser Gly 595 600 605 Gly Gly Gly Ser
Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly 610 615 620 Gly Gly
Ser Arg Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu 625 630 635
640 Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe
645 650 655 Thr Leu Ser Asn Tyr Gly Met His Trp Ile Arg Gln Ala Pro
Gly Lys 660 665 670 Gly Leu Glu Trp Val Ser Ser Ile Ser Leu Asn Gly
Gly Ser Thr Tyr 675 680 685 Tyr Arg Asp Ser Val Lys Gly Arg Phe Thr
Ile Ser Arg Asp Asn Ala 690 695 700 Lys Ser Thr Leu Tyr Leu Gln Met
Asn Ser Leu Arg Ala Glu Asp Thr 705 710 715 720 Ala Val Tyr Tyr Cys
Ala Ala Gln Asp Ala Tyr Thr Gly Gly Tyr Phe 725 730 735 Asp Tyr Trp
Gly Gln Gly Thr Leu Val Thr Val Ser Ser Met Asp Pro 740 745 750 Ala
Glu Pro Lys Ser Pro Asp Lys Thr His Thr Cys Pro Pro Cys Pro 755 760
765 Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro
770 775 780 Lys Asp Thr Leu Met Ile Ala Arg Thr Pro Glu Val Thr Cys
Val Val 785 790 795 800 Val Asp Val Ser His Glu Asp Pro Glu Val Lys
Phe Asn Trp Tyr Val 805 810 815 Asp Gly Val Glu Val His Asn Ala Lys
Thr Lys Pro Arg Glu Glu Gln 820 825 830 Tyr Asn Ser Thr Tyr Arg Val
Val Ser Val Leu Thr Val Leu His Gln 835 840 845 Asp Trp Leu Asn Gly
Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala 850 855 860 Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro 865 870 875 880
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr 885
890 895 Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro
Ser 900 905 910 Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu
Asn Asn Tyr 915 920 925 Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly
Ser Phe Phe Leu Tyr 930 935 940 Ser Lys Leu Thr Val Asp Lys Ser Arg
Trp Gln Gln Gly Asn Val Phe 945 950 955 960 Ser Cys Ser Val Met His
Glu Ala Leu His Asn His Tyr Thr Gln Lys 965 970 975 Ser Leu Ser Leu
Ser Pro Gly Lys Lys Asp Pro Lys Phe Trp Val Leu 980 985 990 Val Val
Val Gly Gly Val Leu Ala Cys Tyr Ser Leu Leu Val Thr Val 995 1000
1005 Ala Phe Ile Ile Phe Trp Val Arg Ser Arg Val Lys Phe Ser Arg
1010 1015 1020 Ser Ala Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln
Leu Tyr 1025 1030 1035 Asn Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr
Asp Val Leu Asp 1040 1045 1050 Lys Arg Arg Gly Arg Asp Pro Glu Met
Gly Gly Lys Pro Arg Arg 1055 1060 1065 Lys Asn Pro Gln Glu Gly Leu
Tyr Asn Glu Leu Gln Lys Asp Lys 1070 1075 1080 Met Ala Glu Ala Tyr
Ser Glu Ile Gly Met Lys Gly Glu Arg Arg 1085 1090 1095 Arg Gly Lys
Gly His Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala 1100 1105 1110 Thr
Lys Asp Thr Tyr Asp Ala Leu His Met Gln Ala Leu Pro Pro 1115 1120
1125 Arg 21350PRTArtificial SequenceChimeric antigen receptor (CAR)
2Met Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1
5 10 15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser
Leu 20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg
Ala Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln
Lys Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His Thr Ser
Arg Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln
Glu Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu
Pro Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys
Ala Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135
140 Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly
145 150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr
Val Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile
Arg Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile
Trp Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser
Arg Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe
Leu Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile
Tyr Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255
Met Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260
265 270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile
Ala 275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro
Ala Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala
Cys Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly Gly Val
Leu Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe Ile Ile
Phe Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala Asp Ala
Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365 Glu Leu
Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370 375 380
Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro 385
390 395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala
Glu Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg
Gly Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala
Thr Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala Leu Pro
Pro Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr Cys Gly
Asp Val Glu Glu Asn Pro Gly Pro Met Ala 465 470 475 480 Val Pro Thr
Gln Val Leu Gly Leu Leu Leu Leu Trp Leu Thr Asp Ala 485 490 495 Arg
Cys Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser 500 505
510 Val Gly Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu Asp Ile Tyr
515 520 525 Phe Asn Leu Val Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro
Lys Leu 530 535 540 Leu Ile Tyr Asp Thr Asn Arg Leu Ala Asp Gly Val
Pro Ser Arg Phe 545 550 555 560 Ser Gly Ser Gly Ser Gly Thr Gln Tyr
Thr Leu Thr Ile Ser Ser Leu 565 570 575 Gln Pro Glu Asp Phe Ala Thr
Tyr Tyr Cys Gln His Tyr Lys Asn Tyr 580 585 590 Pro Leu Thr Phe Gly
Gln Gly Thr Lys Leu Glu Ile Lys Arg Ser Gly 595 600 605 Gly Gly Gly
Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly 610 615 620 Gly
Gly Ser Arg Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu 625 630
635 640 Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly
Phe 645 650 655 Thr Leu Ser Asn Tyr Gly Met His Trp Ile Arg Gln Ala
Pro Gly Lys 660 665 670 Gly Leu Glu Trp Val Ser Ser Ile Ser Leu Asn
Gly Gly Ser Thr Tyr 675 680 685 Tyr Arg Asp Ser Val Lys Gly Arg Phe
Thr Ile Ser Arg Asp Asn Ala 690 695 700 Lys Ser Thr Leu Tyr Leu Gln
Met Asn Ser Leu Arg Ala Glu Asp Thr 705 710 715 720 Ala Val Tyr Tyr
Cys Ala Ala Gln Asp Ala Tyr Thr Gly Gly Tyr Phe 725 730 735 Asp Tyr
Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Met Asp Pro 740 745 750
Ala Glu Pro Lys Ser Pro Asp Lys Thr His Thr Cys Pro Pro Cys Pro 755
760 765 Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro 770 775 780 Lys Asp Thr Leu Met Ile Ala Arg Thr Pro Glu Val Thr
Cys Val Val 785 790 795 800 Val Asp Val Ser His Glu Asp Pro Glu Val
Lys Phe Asn Trp Tyr Val 805 810 815 Asp Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln 820 825 830 Tyr Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln 835 840 845 Asp Trp Leu Asn
Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala 850 855 860 Leu Pro
Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro 865 870 875
880 Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr
885 890 895 Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr
Pro Ser 900 905 910 Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn Asn Tyr 915 920 925 Lys Thr Thr Pro Pro Val Leu Asp Ser Asp
Gly Ser Phe Phe Leu Tyr 930 935 940 Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp Gln Gln Gly Asn Val Phe 945 950 955 960 Ser Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys 965 970 975 Ser Leu Ser
Leu Ser Pro Gly Lys Lys Asp Pro Lys Ala Val Phe Gly 980 985 990 Cys
Ile Phe Gly Ala Leu Val Ile Val Thr Val Gly Gly Phe Ile Phe 995
1000 1005 Trp Arg Lys Lys Arg Lys Asp Ala Lys Asn Asn Glu Val Ser
Phe 1010 1015 1020 Ser Gln Ile Lys Pro Lys Lys Ser Lys Leu Ile Arg
Val Glu Asn 1025 1030 1035 Phe Glu Ala Tyr Phe Lys Lys Gln Gln Ala
Asp Ser Asn Cys Gly 1040 1045 1050 Phe Ala Glu Glu Tyr Glu Asp Leu
Lys Leu Val Gly Ile Ser Gln 1055 1060 1065 Pro Lys Tyr Ala Ala Glu
Leu Ala Glu Asn Arg Gly Lys Asn Arg 1070 1075 1080 Tyr Asn Asn Val
Leu Pro Tyr Asp Ile Ser Arg Val Lys Leu Ser 1085 1090 1095 Val Gln
Thr His Ser Thr Asp Asp Tyr Ile Asn Ala Asn Tyr Met 1100 1105 1110
Pro Gly Tyr His Ser Lys Lys Asp Phe Ile Ala Thr Gln Gly Pro 1115
1120 1125 Leu Pro Asn Thr Leu Lys Asp Phe Trp Arg Met Val Trp Glu
Lys 1130 1135 1140 Asn Val Tyr Ala Ile Ile Met Leu Thr Lys Cys Val
Glu Gln Gly
1145 1150 1155 Arg Thr Lys Cys Glu Glu Tyr Trp Pro Ser Lys Gln Ala
Gln Asp 1160 1165 1170 Tyr Gly Asp Ile Thr Val Ala Met Thr Ser Glu
Ile Val Leu Pro 1175 1180 1185 Glu Trp Thr Ile Arg Asp Phe Thr Val
Lys Asn Ile Gln Thr Ser 1190 1195 1200 Glu Ser His Pro Leu Arg Gln
Phe His Phe Thr Ser Trp Pro Asp 1205 1210 1215 His Gly Val Pro Asp
Thr Thr Asp Leu Leu Ile Asn Phe Arg Tyr 1220 1225 1230 Leu Val Arg
Asp Tyr Met Lys Gln Ser Pro Pro Glu Ser Pro Ile 1235 1240 1245 Leu
Val His Cys Ser Ala Gly Val Gly Arg Thr Gly Thr Phe Ile 1250 1255
1260 Ala Ile Asp Arg Leu Ile Tyr Gln Ile Glu Asn Glu Asn Thr Val
1265 1270 1275 Asp Val Tyr Gly Ile Val Tyr Asp Leu Arg Met His Arg
Pro Leu 1280 1285 1290 Met Val Gln Thr Glu Asp Gln Tyr Val Phe Leu
Asn Gln Cys Val 1295 1300 1305 Leu Asp Ile Val Arg Ser Gln Lys Asp
Ser Lys Val Asp Leu Ile 1310 1315 1320 Tyr Gln Asn Thr Thr Ala Met
Thr Ile Tyr Glu Asn Leu Ala Pro 1325 1330 1335 Val Thr Thr Phe Gly
Lys Thr Asn Gly Tyr Ile Ala 1340 1345 1350 31717PRTArtificial
SequenceChimeric antigen receptor (CAR) 3Met Ser Leu Pro Val Thr
Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1 5 10 15 His Ala Ala Arg
Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser Leu 20 25 30 Ser Ala
Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg Ala Ser Gln 35 40 45
Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln Lys Pro Asp Gly Thr 50
55 60 Val Lys Leu Leu Ile Tyr His Thr Ser Arg Leu His Ser Gly Val
Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Tyr Ser
Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln Glu Asp Ile Ala Thr Tyr
Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu Pro Tyr Thr Phe Gly Gly
Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys Ala Gly Gly Gly Gly Ser
Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135 140 Ser Gly Gly Gly Gly
Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly 145 150 155 160 Leu Val
Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr Val Ser Gly 165 170 175
Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile Arg Gln Pro Pro Arg 180
185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile Trp Gly Ser Glu Thr Thr
Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser Arg Leu Thr Ile Ile Lys
Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe Leu Lys Met Asn Ser Leu
Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile Tyr Tyr Cys Ala Lys His
Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255 Met Asp Tyr Trp Gly Gln
Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260 265 270 Thr Thr Thr Pro
Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala 275 280 285 Ser Gln
Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala Ala Gly 290 295 300
Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala Cys Asp Ile Phe Trp 305
310 315 320 Val Leu Val Val Val Gly Gly Val Leu Ala Cys Tyr Ser Leu
Leu Val 325 330 335 Thr Val Ala Phe Ile Ile Phe Trp Val Arg Arg Val
Lys Phe Ser Arg 340 345 350 Ser Ala Asp Ala Pro Ala Tyr Gln Gln Gly
Gln Asn Gln Leu Tyr Asn 355 360 365 Glu Leu Asn Leu Gly Arg Arg Glu
Glu Tyr Asp Val Leu Asp Lys Arg 370 375 380 Arg Gly Arg Asp Pro Glu
Met Gly Gly Lys Pro Arg Arg Lys Asn Pro 385 390 395 400 Gln Glu Gly
Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu Ala 405 410 415 Tyr
Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly Lys Gly His 420 425
430 Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp Thr Tyr Asp
435 440 445 Ala Leu His Met Gln Ala Leu Pro Pro Arg Arg Ala Glu Gly
Arg Gly 450 455 460 Ser Leu Leu Thr Cys Gly Asp Val Glu Glu Asn Pro
Gly Pro Met Ala 465 470 475 480 Val Pro Thr Gln Val Leu Gly Leu Leu
Leu Leu Trp Leu Thr Asp Ala 485 490 495 Arg Cys Asp Ile Gln Met Thr
Gln Ser Pro Ser Ser Leu Ser Ala Ser 500 505 510 Val Gly Asp Arg Val
Thr Ile Thr Cys Arg Ala Ser Glu Asp Ile Tyr 515 520 525 Phe Asn Leu
Val Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu 530 535 540 Leu
Ile Tyr Asp Thr Asn Arg Leu Ala Asp Gly Val Pro Ser Arg Phe 545 550
555 560 Ser Gly Ser Gly Ser Gly Thr Gln Tyr Thr Leu Thr Ile Ser Ser
Leu 565 570 575 Gln Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Gln His Tyr
Lys Asn Tyr 580 585 590 Pro Leu Thr Phe Gly Gln Gly Thr Lys Leu Glu
Ile Lys Arg Ser Gly 595 600 605 Gly Gly Gly Ser Gly Gly Gly Gly Ser
Gly Gly Gly Gly Ser Gly Gly 610 615 620 Gly Gly Ser Arg Ser Glu Val
Gln Leu Val Glu Ser Gly Gly Gly Leu 625 630 635 640 Val Gln Pro Gly
Gly Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe 645 650 655 Thr Leu
Ser Asn Tyr Gly Met His Trp Ile Arg Gln Ala Pro Gly Lys 660 665 670
Gly Leu Glu Trp Val Ser Ser Ile Ser Leu Asn Gly Gly Ser Thr Tyr 675
680 685 Tyr Arg Asp Ser Val Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn
Ala 690 695 700 Lys Ser Thr Leu Tyr Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr 705 710 715 720 Ala Val Tyr Tyr Cys Ala Ala Gln Asp Ala
Tyr Thr Gly Gly Tyr Phe 725 730 735 Asp Tyr Trp Gly Gln Gly Thr Leu
Val Thr Val Ser Ser Met Asp Pro 740 745 750 Ala Glu Pro Lys Ser Pro
Asp Lys Thr His Thr Cys Pro Pro Cys Pro 755 760 765 Ala Pro Pro Val
Ala Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro 770 775 780 Lys Asp
Thr Leu Met Ile Ala Arg Thr Pro Glu Val Thr Cys Val Val 785 790 795
800 Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val
805 810 815 Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu
Glu Gln 820 825 830 Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr
Val Leu His Gln 835 840 845 Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn Lys Ala 850 855 860 Leu Pro Ala Pro Ile Glu Lys Thr
Ile Ser Lys Ala Lys Gly Gln Pro 865 870 875 880 Arg Glu Pro Gln Val
Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr 885 890 895 Lys Asn Gln
Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser 900 905 910 Asp
Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr 915 920
925 Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr
930 935 940 Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn
Val Phe 945 950 955 960 Ser Cys Ser Val Met His Glu Ala Leu His Asn
His Tyr Thr Gln Lys 965 970 975 Ser Leu Ser Leu Ser Pro Gly Lys Lys
Asp Pro Lys Ala Leu Ile Ala 980 985 990 Phe Leu Ala Phe Leu Ile Ile
Val Thr Ser Ile Ala Leu Leu Val Val 995 1000 1005 Leu Tyr Lys Ile
Tyr Asp Leu His Lys Lys Arg Ser Cys Asn Leu 1010 1015 1020 Asp Glu
Gln Gln Glu Leu Val Glu Arg Asp Asp Glu Lys Gln Leu 1025 1030 1035
Met Asn Val Glu Pro Ile His Ala Asp Ile Leu Leu Glu Thr Tyr 1040
1045 1050 Lys Arg Lys Ile Ala Asp Glu Gly Arg Leu Phe Leu Ala Glu
Phe 1055 1060 1065 Gln Ser Ile Pro Arg Val Phe Ser Lys Phe Pro Ile
Lys Glu Ala 1070 1075 1080 Arg Lys Pro Phe Asn Gln Asn Lys Asn Arg
Tyr Val Asp Ile Leu 1085 1090 1095 Pro Tyr Asp Tyr Asn Arg Val Glu
Leu Ser Glu Ile Asn Gly Asp 1100 1105 1110 Ala Gly Ser Asn Tyr Ile
Asn Ala Ser Tyr Ile Asp Gly Phe Lys 1115 1120 1125 Glu Pro Arg Lys
Tyr Ile Ala Ala Gln Gly Pro Arg Asp Glu Thr 1130 1135 1140 Val Asp
Asp Phe Trp Arg Met Ile Trp Glu Gln Lys Ala Thr Val 1145 1150 1155
Ile Val Met Val Thr Arg Cys Glu Glu Gly Asn Arg Asn Lys Cys 1160
1165 1170 Ala Glu Tyr Trp Pro Ser Met Glu Glu Gly Thr Arg Ala Phe
Gly 1175 1180 1185 Asp Val Val Val Lys Ile Asn Gln His Lys Arg Cys
Pro Asp Tyr 1190 1195 1200 Ile Ile Gln Lys Leu Asn Ile Val Asn Lys
Lys Glu Lys Ala Thr 1205 1210 1215 Gly Arg Glu Val Thr His Ile Gln
Phe Thr Ser Trp Pro Asp His 1220 1225 1230 Gly Val Pro Glu Asp Pro
His Leu Leu Leu Lys Leu Arg Arg Arg 1235 1240 1245 Val Asn Ala Phe
Ser Asn Phe Phe Ser Gly Pro Ile Val Val His 1250 1255 1260 Cys Ser
Ala Gly Val Gly Arg Thr Gly Thr Tyr Ile Gly Ile Asp 1265 1270 1275
Ala Met Leu Glu Gly Leu Glu Ala Glu Asn Lys Val Asp Val Tyr 1280
1285 1290 Gly Tyr Val Val Lys Leu Arg Arg Gln Arg Cys Leu Met Val
Gln 1295 1300 1305 Val Glu Ala Gln Tyr Ile Leu Ile His Gln Ala Leu
Val Glu Tyr 1310 1315 1320 Asn Gln Phe Gly Glu Thr Glu Val Asn Leu
Ser Glu Leu His Pro 1325 1330 1335 Tyr Leu His Asn Met Lys Lys Arg
Asp Pro Pro Ser Glu Pro Ser 1340 1345 1350 Pro Leu Glu Ala Glu Phe
Gln Arg Leu Pro Ser Tyr Arg Ser Trp 1355 1360 1365 Arg Thr Gln His
Ile Gly Asn Gln Glu Glu Asn Lys Ser Lys Asn 1370 1375 1380 Arg Asn
Ser Asn Val Ile Pro Tyr Asp Tyr Asn Arg Val Pro Leu 1385 1390 1395
Lys His Glu Leu Glu Met Ser Lys Glu Ser Glu His Asp Ser Asp 1400
1405 1410 Glu Ser Ser Asp Asp Asp Ser Asp Ser Glu Glu Pro Ser Lys
Tyr 1415 1420 1425 Ile Asn Ala Ser Phe Ile Met Ser Tyr Trp Lys Pro
Glu Val Met 1430 1435 1440 Ile Ala Ala Gln Gly Pro Leu Lys Glu Thr
Ile Gly Asp Phe Trp 1445 1450 1455 Gln Met Ile Phe Gln Arg Lys Val
Lys Val Ile Val Met Leu Thr 1460 1465 1470 Glu Leu Lys His Gly Asp
Gln Glu Ile Cys Ala Gln Tyr Trp Gly 1475 1480 1485 Glu Gly Lys Gln
Thr Tyr Gly Asp Ile Glu Val Asp Leu Lys Asp 1490 1495 1500 Thr Asp
Lys Ser Ser Thr Tyr Thr Leu Arg Val Phe Glu Leu Arg 1505 1510 1515
His Ser Lys Arg Lys Asp Ser Arg Thr Val Tyr Gln Tyr Gln Tyr 1520
1525 1530 Thr Asn Trp Ser Val Glu Gln Leu Pro Ala Glu Pro Lys Glu
Leu 1535 1540 1545 Ile Ser Met Ile Gln Val Val Lys Gln Lys Leu Pro
Gln Lys Asn 1550 1555 1560 Ser Ser Glu Gly Asn Lys His His Lys Ser
Thr Pro Leu Leu Ile 1565 1570 1575 His Cys Arg Asp Gly Ser Gln Gln
Thr Gly Ile Phe Cys Ala Leu 1580 1585 1590 Leu Asn Leu Leu Glu Ser
Ala Glu Thr Glu Glu Val Val Asp Ile 1595 1600 1605 Phe Gln Val Val
Lys Ala Leu Arg Lys Ala Arg Pro Gly Met Val 1610 1615 1620 Ser Thr
Phe Glu Gln Tyr Gln Phe Leu Tyr Asp Val Ile Ala Ser 1625 1630 1635
Thr Tyr Pro Ala Gln Asn Gly Gln Val Lys Lys Asn Asn His Gln 1640
1645 1650 Glu Asp Lys Ile Glu Phe Asp Asn Glu Val Asp Lys Val Lys
Gln 1655 1660 1665 Asp Ala Asn Cys Val Asn Pro Leu Gly Ala Pro Glu
Lys Leu Pro 1670 1675 1680 Glu Ala Lys Glu Gln Ala Glu Gly Ser Glu
Pro Thr Ser Gly Thr 1685 1690 1695 Glu Gly Pro Glu His Ser Val Asn
Gly Pro Ala Ser Pro Ala Leu 1700 1705 1710 Asn Gln Gly Ser 1715
41114PRTArtificial SequenceChimeric antigen receptor (CAR) 4Met Ser
Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1 5 10 15
His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser Leu 20
25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg Ala Ser
Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln Lys Pro
Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His Thr Ser Arg Leu
His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln Glu Asp
Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu Pro Tyr
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys Ala Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135 140 Ser
Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly 145 150
155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr Val Ser
Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile Arg Gln
Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile Trp Gly
Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser Arg Leu
Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe Leu Lys
Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile Tyr Tyr
Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255 Met Asp
Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260 265 270
Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala 275
280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala Ala
Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala Cys Asp
Ile Phe Trp 305 310 315 320 Val Leu Val Val Val
Gly Gly Val Leu Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala
Phe Ile Ile Phe Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser
Ala Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360
365 Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg
370 375 380 Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys
Asn Pro 385 390 395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp
Lys Met Ala Glu Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu
Arg Arg Arg Gly Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu
Ser Thr Ala Thr Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln
Ala Leu Pro Pro Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu
Thr Cys Gly Asp Val Glu Glu Asn Pro Gly Pro Met Ala 465 470 475 480
Val Pro Thr Gln Val Leu Gly Leu Leu Leu Leu Trp Leu Thr Asp Ala 485
490 495 Arg Cys Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala
Ser 500 505 510 Val Gly Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu
Asp Ile Tyr 515 520 525 Phe Asn Leu Val Trp Tyr Gln Gln Lys Pro Gly
Lys Ala Pro Lys Leu 530 535 540 Leu Ile Tyr Asp Thr Asn Arg Leu Ala
Asp Gly Val Pro Ser Arg Phe 545 550 555 560 Ser Gly Ser Gly Ser Gly
Thr Gln Tyr Thr Leu Thr Ile Ser Ser Leu 565 570 575 Gln Pro Glu Asp
Phe Ala Thr Tyr Tyr Cys Gln His Tyr Lys Asn Tyr 580 585 590 Pro Leu
Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys Arg Ser Gly 595 600 605
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly 610
615 620 Gly Gly Ser Arg Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu 625 630 635 640 Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala
Ala Ser Gly Phe 645 650 655 Thr Leu Ser Asn Tyr Gly Met His Trp Ile
Arg Gln Ala Pro Gly Lys 660 665 670 Gly Leu Glu Trp Val Ser Ser Ile
Ser Leu Asn Gly Gly Ser Thr Tyr 675 680 685 Tyr Arg Asp Ser Val Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala 690 695 700 Lys Ser Thr Leu
Tyr Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr 705 710 715 720 Ala
Val Tyr Tyr Cys Ala Ala Gln Asp Ala Tyr Thr Gly Gly Tyr Phe 725 730
735 Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Met Asp Pro
740 745 750 Ala Thr Thr Thr Lys Pro Val Leu Arg Thr Pro Ser Pro Val
His Pro 755 760 765 Thr Gly Thr Ser Gln Pro Gln Arg Pro Glu Asp Cys
Arg Pro Arg Gly 770 775 780 Ser Val Lys Gly Thr Gly Leu Asp Phe Ala
Cys Asp Ile Tyr Trp Ala 785 790 795 800 Pro Leu Ala Gly Ile Cys Val
Ala Leu Leu Leu Ser Leu Ile Ile Thr 805 810 815 Leu Ile Cys Tyr His
Arg Ser Arg Lys Arg Val Cys Lys Ser Gly Gly 820 825 830 Gly Ser Phe
Trp Glu Glu Phe Glu Ser Leu Gln Lys Gln Glu Val Lys 835 840 845 Asn
Leu His Gln Arg Leu Glu Gly Gln Arg Pro Glu Asn Lys Gly Lys 850 855
860 Asn Arg Tyr Lys Asn Ile Leu Pro Phe Asp His Ser Arg Val Ile Leu
865 870 875 880 Gln Gly Arg Asp Ser Asn Ile Pro Gly Ser Asp Tyr Ile
Asn Ala Asn 885 890 895 Tyr Ile Lys Asn Gln Leu Leu Gly Pro Asp Glu
Asn Ala Lys Thr Tyr 900 905 910 Ile Ala Ser Gln Gly Cys Leu Glu Ala
Thr Val Asn Asp Phe Trp Gln 915 920 925 Met Ala Trp Gln Glu Asn Ser
Arg Val Ile Val Met Thr Thr Arg Glu 930 935 940 Val Glu Lys Gly Arg
Asn Lys Cys Val Pro Tyr Trp Pro Glu Val Gly 945 950 955 960 Met Gln
Arg Ala Tyr Gly Pro Tyr Ser Val Thr Asn Cys Gly Glu His 965 970 975
Asp Thr Thr Glu Tyr Lys Leu Arg Thr Leu Gln Val Ser Pro Leu Asp 980
985 990 Asn Gly Asp Leu Ile Arg Glu Ile Trp His Tyr Gln Tyr Leu Ser
Trp 995 1000 1005 Pro Asp His Gly Val Pro Ser Glu Pro Gly Gly Val
Leu Ser Phe 1010 1015 1020 Leu Asp Gln Ile Asn Gln Arg Gln Glu Ser
Leu Pro His Ala Gly 1025 1030 1035 Pro Ile Ile Val His Cys Ser Ala
Gly Ile Gly Arg Thr Gly Thr 1040 1045 1050 Ile Ile Val Ile Asp Met
Leu Met Glu Asn Ile Ser Thr Lys Gly 1055 1060 1065 Leu Asp Cys Asp
Ile Asp Ile Gln Lys Thr Ile Gln Met Val Arg 1070 1075 1080 Ala Gln
Arg Ser Gly Met Val Gln Thr Glu Ala Gln Tyr Lys Phe 1085 1090 1095
Ile Tyr Val Ala Ile Ala Gln Phe Ile Glu Thr Thr Lys Lys Lys 1100
1105 1110 Leu 5918PRTArtificial SequenceChimeric antigen receptor
(CAR) 5Met Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu
Leu 1 5 10 15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr
Ser Ser Leu 20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser
Cys Arg Ala Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr
Gln Gln Lys Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His
Thr Ser Arg Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu
Glu Gln Glu Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn
Thr Leu Pro Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120
125 Lys Ala Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly
130 135 140 Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly
Pro Gly 145 150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr
Cys Thr Val Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser
Trp Ile Arg Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly
Val Ile Trp Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu
Lys Ser Arg Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln
Val Phe Leu Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240
Ala Ile Tyr Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245
250 255 Met Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp
Pro 260 265 270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro
Thr Ile Ala 275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys
Arg Pro Ala Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp
Phe Ala Cys Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly
Gly Val Leu Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe
Ile Ile Phe Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala
Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365
Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370
375 380 Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn
Pro 385 390 395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys
Met Ala Glu Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg
Arg Arg Gly Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser
Thr Ala Thr Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala
Leu Pro Pro Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr
Cys Gly Asp Val Glu Glu Asn Pro Gly Pro Met Ala 465 470 475 480 Val
Pro Thr Gln Val Leu Gly Leu Leu Leu Leu Trp Leu Thr Asp Ala 485 490
495 Arg Cys Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser
500 505 510 Val Gly Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu Asp
Ile Tyr 515 520 525 Phe Asn Leu Val Trp Tyr Gln Gln Lys Pro Gly Lys
Ala Pro Lys Leu 530 535 540 Leu Ile Tyr Asp Thr Asn Arg Leu Ala Asp
Gly Val Pro Ser Arg Phe 545 550 555 560 Ser Gly Ser Gly Ser Gly Thr
Gln Tyr Thr Leu Thr Ile Ser Ser Leu 565 570 575 Gln Pro Glu Asp Phe
Ala Thr Tyr Tyr Cys Gln His Tyr Lys Asn Tyr 580 585 590 Pro Leu Thr
Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys Arg Ser Gly 595 600 605 Gly
Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly 610 615
620 Gly Gly Ser Arg Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
625 630 635 640 Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly Phe 645 650 655 Thr Leu Ser Asn Tyr Gly Met His Trp Ile Arg
Gln Ala Pro Gly Lys 660 665 670 Gly Leu Glu Trp Val Ser Ser Ile Ser
Leu Asn Gly Gly Ser Thr Tyr 675 680 685 Tyr Arg Asp Ser Val Lys Gly
Arg Phe Thr Ile Ser Arg Asp Asn Ala 690 695 700 Lys Ser Thr Leu Tyr
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr 705 710 715 720 Ala Val
Tyr Tyr Cys Ala Ala Gln Asp Ala Tyr Thr Gly Gly Tyr Phe 725 730 735
Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Met Asp Pro 740
745 750 Ala Thr Thr Thr Lys Pro Val Leu Arg Thr Pro Ser Pro Val His
Pro 755 760 765 Thr Gly Thr Ser Gln Pro Gln Arg Pro Glu Asp Cys Arg
Pro Arg Gly 770 775 780 Ser Val Lys Gly Thr Gly Leu Asp Phe Ala Cys
Asp Ile Leu Ile Gly 785 790 795 800 Val Ser Val Val Phe Leu Phe Cys
Leu Leu Leu Leu Val Leu Phe Cys 805 810 815 Leu His Arg Gln Asn Gln
Ile Lys Gln Gly Pro Pro Arg Ser Lys Asp 820 825 830 Glu Glu Gln Lys
Pro Gln Gln Arg Pro Asp Leu Ala Val Asp Val Leu 835 840 845 Glu Arg
Thr Ala Asp Lys Ala Thr Val Asn Gly Leu Pro Glu Lys Asp 850 855 860
Arg Glu Thr Asp Thr Ser Ala Leu Ala Ala Gly Ser Ser Gln Glu Val 865
870 875 880 Thr Tyr Ala Gln Leu Asp His Trp Ala Leu Thr Gln Arg Thr
Ala Arg 885 890 895 Ala Val Ser Pro Gln Ser Thr Lys Pro Met Ala Glu
Ser Ile Thr Tyr 900 905 910 Ala Ala Val Ala Arg His 915
61363PRTArtificial SequenceChimeric antigen receptor (CAR) 6Met Ser
Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1 5 10 15
His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser Leu 20
25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg Ala Ser
Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln Lys Pro
Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His Thr Ser Arg Leu
His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly Ser Gly
Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln Glu Asp
Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu Pro Tyr
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys Ala Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135 140 Ser
Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly 145 150
155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr Val Ser
Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile Arg Gln
Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile Trp Gly
Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser Arg Leu
Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe Leu Lys
Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile Tyr Tyr
Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255 Met Asp
Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260 265 270
Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala 275
280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala Ala
Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala Cys Asp
Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly Gly Val Leu Ala
Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe Ile Ile Phe Trp
Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala Asp Ala Pro Ala
Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365 Glu Leu Asn Leu
Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370 375 380 Arg Gly
Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro 385 390 395
400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu Ala
405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly Lys
Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys
Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala Leu Pro Pro Arg
Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr Cys Gly Asp Val
Glu Glu Asn Pro Gly Pro Met Ala 465 470 475 480 Val Pro Thr Gln Val
Leu Gly Leu Leu Leu Leu Trp Leu Thr Asp Ala 485 490 495 Arg Cys Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser 500 505 510 Val
Gly Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu Asp Ile Tyr 515 520
525 Phe Asn Leu Val Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
530 535 540 Leu Ile Tyr Asp Thr Asn Arg Leu Ala Asp Gly Val Pro Ser
Arg Phe 545 550 555 560 Ser Gly Ser Gly Ser Gly Thr Gln Tyr Thr Leu
Thr Ile Ser Ser Leu 565 570 575 Gln Pro
Glu Asp Phe Ala Thr Tyr Tyr Cys Gln His Tyr Lys Asn Tyr 580 585 590
Pro Leu Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys Arg Ser Gly 595
600 605 Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly
Gly 610 615 620 Gly Gly Ser Arg Ser Glu Val Gln Leu Val Glu Ser Gly
Gly Gly Leu 625 630 635 640 Val Gln Pro Gly Gly Ser Leu Arg Leu Ser
Cys Ala Ala Ser Gly Phe 645 650 655 Thr Leu Ser Asn Tyr Gly Met His
Trp Ile Arg Gln Ala Pro Gly Lys 660 665 670 Gly Leu Glu Trp Val Ser
Ser Ile Ser Leu Asn Gly Gly Ser Thr Tyr 675 680 685 Tyr Arg Asp Ser
Val Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala 690 695 700 Lys Ser
Thr Leu Tyr Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr 705 710 715
720 Ala Val Tyr Tyr Cys Ala Ala Gln Asp Ala Tyr Thr Gly Gly Tyr Phe
725 730 735 Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Met
Asp Pro 740 745 750 Ala Thr Thr Thr Lys Pro Val Leu Arg Thr Pro Ser
Pro Val His Pro 755 760 765 Thr Gly Thr Ser Gln Pro Gln Arg Pro Glu
Asp Cys Arg Pro Arg Gly 770 775 780 Ser Val Lys Gly Thr Gly Leu Asp
Phe Ala Cys Asp Ile Leu Ile Gly 785 790 795 800 Val Ser Val Val Phe
Leu Phe Cys Leu Leu Leu Leu Val Leu Phe Cys 805 810 815 Leu His Arg
Gln Asn Gln Ile Lys Gln Gly Pro Pro Arg Ser Lys Asp 820 825 830 Glu
Glu Gln Lys Pro Gln Gln Arg Pro Asp Leu Ala Val Asp Val Leu 835 840
845 Glu Arg Thr Ala Asp Lys Ala Thr Val Asn Gly Leu Pro Glu Lys Asp
850 855 860 Arg Glu Thr Asp Thr Ser Ala Leu Ala Ala Gly Ser Ser Gln
Glu Val 865 870 875 880 Thr Tyr Ala Gln Leu Asp His Trp Ala Leu Thr
Gln Arg Thr Ala Arg 885 890 895 Ala Val Ser Pro Gln Ser Thr Lys Pro
Met Ala Glu Ser Ile Thr Tyr 900 905 910 Ala Ala Val Ala Arg His Arg
Ala Glu Gly Arg Gly Ser Leu Leu Thr 915 920 925 Cys Gly Asp Val Glu
Glu Asn Pro Gly Pro Trp Tyr His Gly His Met 930 935 940 Ser Gly Gly
Gln Ala Glu Thr Leu Leu Gln Ala Lys Gly Glu Pro Trp 945 950 955 960
Thr Phe Leu Val Arg Glu Ser Leu Ser Gln Pro Gly Asp Phe Val Leu 965
970 975 Ser Val Leu Ser Asp Gln Pro Lys Ala Gly Pro Gly Ser Pro Leu
Arg 980 985 990 Val Thr His Ile Lys Val Met Cys Glu Gly Gly Arg Tyr
Thr Val Gly 995 1000 1005 Gly Leu Glu Thr Phe Asp Ser Leu Thr Asp
Leu Val Glu His Phe 1010 1015 1020 Lys Lys Thr Gly Ile Glu Glu Ala
Ser Gly Ala Phe Val Tyr Leu 1025 1030 1035 Arg Gln Pro Tyr Ser Gly
Gly Gly Gly Ser Phe Glu Ala Tyr Phe 1040 1045 1050 Lys Lys Gln Gln
Ala Asp Ser Asn Cys Gly Phe Ala Glu Glu Tyr 1055 1060 1065 Glu Asp
Leu Lys Leu Val Gly Ile Ser Gln Pro Lys Tyr Ala Ala 1070 1075 1080
Glu Leu Ala Glu Asn Arg Gly Lys Asn Arg Tyr Asn Asn Val Leu 1085
1090 1095 Pro Tyr Asp Ile Ser Arg Val Lys Leu Ser Val Gln Thr His
Ser 1100 1105 1110 Thr Asp Asp Tyr Ile Asn Ala Asn Tyr Met Pro Gly
Tyr His Ser 1115 1120 1125 Lys Lys Asp Phe Ile Ala Thr Gln Gly Pro
Leu Pro Asn Thr Leu 1130 1135 1140 Lys Asp Phe Trp Arg Met Val Trp
Glu Lys Asn Val Tyr Ala Ile 1145 1150 1155 Ile Met Leu Thr Lys Cys
Val Glu Gln Gly Arg Thr Lys Cys Glu 1160 1165 1170 Glu Tyr Trp Pro
Ser Lys Gln Ala Gln Asp Tyr Gly Asp Ile Thr 1175 1180 1185 Val Ala
Met Thr Ser Glu Ile Val Leu Pro Glu Trp Thr Ile Arg 1190 1195 1200
Asp Phe Thr Val Lys Asn Ile Gln Thr Ser Glu Ser His Pro Leu 1205
1210 1215 Arg Gln Phe His Phe Thr Ser Trp Pro Asp His Gly Val Pro
Asp 1220 1225 1230 Thr Thr Asp Leu Leu Ile Asn Phe Arg Tyr Leu Val
Arg Asp Tyr 1235 1240 1245 Met Lys Gln Ser Pro Pro Glu Ser Pro Ile
Leu Val His Cys Ser 1250 1255 1260 Ala Gly Val Gly Arg Thr Gly Thr
Phe Ile Ala Ile Asp Arg Leu 1265 1270 1275 Ile Tyr Gln Ile Glu Asn
Glu Asn Thr Val Asp Val Tyr Gly Ile 1280 1285 1290 Val Tyr Asp Leu
Arg Met His Arg Pro Leu Met Val Gln Thr Glu 1295 1300 1305 Asp Gln
Tyr Val Phe Leu Asn Gln Cys Val Leu Asp Ile Val Arg 1310 1315 1320
Ser Gln Lys Asp Ser Lys Val Asp Leu Ile Tyr Gln Asn Thr Thr 1325
1330 1335 Ala Met Thr Ile Tyr Glu Asn Leu Ala Pro Val Thr Thr Phe
Gly 1340 1345 1350 Lys Thr Asn Gly Tyr Ile Ala Ser Gly Ser 1355
1360 721PRTArtificial SequenceSignal peptide 7Met Gly Thr Ser Leu
Leu Cys Trp Met Ala Leu Cys Leu Leu Gly Ala 1 5 10 15 Asp His Ala
Asp Gly 20 821PRTArtificial SequenceSignal peptide 8Met Ser Leu Pro
Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1 5 10 15 His Ala
Ala Arg Pro 20 920PRTArtificial SequenceSignal peptide 9Met Ala Val
Pro Thr Gln Val Leu Gly Leu Leu Leu Leu Trp Leu Thr 1 5 10 15 Asp
Ala Arg Cys 20 10234PRTArtificial SequenceSpacer (hinge-CH2CH3 of
human IgG1) 10Ala Glu Pro Lys Ser Pro Asp Lys Thr His Thr Cys Pro
Pro Cys Pro 1 5 10 15 Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu
Phe Pro Pro Lys Pro 20 25 30 Lys Asp Thr Leu Met Ile Ala Arg Thr
Pro Glu Val Thr Cys Val Val 35 40 45 Val Asp Val Ser His Glu Asp
Pro Glu Val Lys Phe Asn Trp Tyr Val 50 55 60 Asp Gly Val Glu Val
His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln 65 70 75 80 Tyr Asn Ser
Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln 85 90 95 Asp
Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala 100 105
110 Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro
115 120 125 Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu
Leu Thr 130 135 140 Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly
Phe Tyr Pro Ser 145 150 155 160 Asp Ile Ala Val Glu Trp Glu Ser Asn
Gly Gln Pro Glu Asn Asn Tyr 165 170 175 Lys Thr Thr Pro Pro Val Leu
Asp Ser Asp Gly Ser Phe Phe Leu Tyr 180 185 190 Ser Lys Leu Thr Val
Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe 195 200 205 Ser Cys Ser
Val Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys 210 215 220 Ser
Leu Ser Leu Ser Pro Gly Lys Lys Asp 225 230 1146PRTArtificial
SequenceSpacer (human CD8 stalk) 11Thr Thr Thr Pro Ala Pro Arg Pro
Pro Thr Pro Ala Pro Thr Ile Ala 1 5 10 15 Ser Gln Pro Leu Ser Leu
Arg Pro Glu Ala Cys Arg Pro Ala Ala Gly 20 25 30 Gly Ala Val His
Thr Arg Gly Leu Asp Phe Ala Cys Asp Ile 35 40 45 1220PRTArtificial
SequenceSpacer (human IgG1 hinge) 12Ala Glu Pro Lys Ser Pro Asp Lys
Thr His Thr Cys Pro Pro Cys Pro 1 5 10 15 Lys Asp Pro Lys 20
13185PRTArtificial SequenceSpacer (CD2 ectodomain) 13Lys Glu Ile
Thr Asn Ala Leu Glu Thr Trp Gly Ala Leu Gly Gln Asp 1 5 10 15 Ile
Asn Leu Asp Ile Pro Ser Phe Gln Met Ser Asp Asp Ile Asp Asp 20 25
30 Ile Lys Trp Glu Lys Thr Ser Asp Lys Lys Lys Ile Ala Gln Phe Arg
35 40 45 Lys Glu Lys Glu Thr Phe Lys Glu Lys Asp Thr Tyr Lys Leu
Phe Lys 50 55 60 Asn Gly Thr Leu Lys Ile Lys His Leu Lys Thr Asp
Asp Gln Asp Ile 65 70 75 80 Tyr Lys Val Ser Ile Tyr Asp Thr Lys Gly
Lys Asn Val Leu Glu Lys 85 90 95 Ile Phe Asp Leu Lys Ile Gln Glu
Arg Val Ser Lys Pro Lys Ile Ser 100 105 110 Trp Thr Cys Ile Asn Thr
Thr Leu Thr Cys Glu Val Met Asn Gly Thr 115 120 125 Asp Pro Glu Leu
Asn Leu Tyr Gln Asp Gly Lys His Leu Lys Leu Ser 130 135 140 Gln Arg
Val Ile Thr His Lys Trp Thr Thr Ser Leu Ser Ala Lys Phe 145 150 155
160 Lys Cys Thr Ala Gly Asn Lys Val Ser Lys Glu Ser Ser Val Glu Pro
165 170 175 Val Ser Cys Pro Glu Lys Gly Leu Asp 180 185
14259PRTArtificial SequenceSpacer (CD34 ectodomain) 14Ser Leu Asp
Asn Asn Gly Thr Ala Thr Pro Glu Leu Pro Thr Gln Gly 1 5 10 15 Thr
Phe Ser Asn Val Ser Thr Asn Val Ser Tyr Gln Glu Thr Thr Thr 20 25
30 Pro Ser Thr Leu Gly Ser Thr Ser Leu His Pro Val Ser Gln His Gly
35 40 45 Asn Glu Ala Thr Thr Asn Ile Thr Glu Thr Thr Val Lys Phe
Thr Ser 50 55 60 Thr Ser Val Ile Thr Ser Val Tyr Gly Asn Thr Asn
Ser Ser Val Gln 65 70 75 80 Ser Gln Thr Ser Val Ile Ser Thr Val Phe
Thr Thr Pro Ala Asn Val 85 90 95 Ser Thr Pro Glu Thr Thr Leu Lys
Pro Ser Leu Ser Pro Gly Asn Val 100 105 110 Ser Asp Leu Ser Thr Thr
Ser Thr Ser Leu Ala Thr Ser Pro Thr Lys 115 120 125 Pro Tyr Thr Ser
Ser Ser Pro Ile Leu Ser Asp Ile Lys Ala Glu Ile 130 135 140 Lys Cys
Ser Gly Ile Arg Glu Val Lys Leu Thr Gln Gly Ile Cys Leu 145 150 155
160 Glu Gln Asn Lys Thr Ser Ser Cys Ala Glu Phe Lys Lys Asp Arg Gly
165 170 175 Glu Gly Leu Ala Arg Val Leu Cys Gly Glu Glu Gln Ala Asp
Ala Asp 180 185 190 Ala Gly Ala Gln Val Cys Ser Leu Leu Leu Ala Gln
Ser Glu Val Arg 195 200 205 Pro Gln Cys Leu Leu Leu Val Leu Ala Asn
Arg Thr Glu Ile Ser Ser 210 215 220 Lys Leu Gln Leu Met Lys Lys His
Gln Ser Asp Leu Lys Lys Leu Gly 225 230 235 240 Ile Leu Asp Phe Thr
Glu Gln Asp Val Ala Ser His Gln Ser Tyr Ser 245 250 255 Gln Lys Thr
15140PRTArtificial SequenceCD28 transmembrane domain and CD3 Z
endodomains 15Phe Trp Val Leu Val Val Val Gly Gly Val Leu Ala Cys
Tyr Ser Leu 1 5 10 15 Leu Val Thr Val Ala Phe Ile Ile Phe Trp Val
Arg Arg Val Lys Phe 20 25 30 Ser Arg Ser Ala Asp Ala Pro Ala Tyr
Gln Gln Gly Gln Asn Gln Leu 35 40 45 Tyr Asn Glu Leu Asn Leu Gly
Arg Arg Glu Glu Tyr Asp Val Leu Asp 50 55 60 Lys Arg Arg Gly Arg
Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys 65 70 75 80 Asn Pro Gln
Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala 85 90 95 Glu
Ala Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly Lys 100 105
110 Gly His Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp Thr
115 120 125 Tyr Asp Ala Leu His Met Gln Ala Leu Pro Pro Arg 130 135
140 16180PRTArtificial SequenceCD28 transmembrane domain and CD28
and CD3 Zeta endodomains 16Phe Trp Val Leu Val Val Val Gly Gly Val
Leu Ala Cys Tyr Ser Leu 1 5 10 15 Leu Val Thr Val Ala Phe Ile Ile
Phe Trp Val Arg Ser Lys Arg Ser 20 25 30 Arg Leu Leu His Ser Asp
Tyr Met Asn Met Thr Pro Arg Arg Pro Gly 35 40 45 Pro Thr Arg Lys
His Tyr Gln Pro Tyr Ala Pro Pro Arg Asp Phe Ala 50 55 60 Ala Tyr
Arg Ser Arg Val Lys Phe Ser Arg Ser Ala Asp Ala Pro Ala 65 70 75 80
Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn Glu Leu Asn Leu Gly Arg 85
90 95 Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg Arg Gly Arg Asp Pro
Glu 100 105 110 Met Gly Gly Lys Pro Arg Arg Lys Asn Pro Gln Glu Gly
Leu Tyr Asn 115 120 125 Glu Leu Gln Lys Asp Lys Met Ala Glu Ala Tyr
Ser Glu Ile Gly Met 130 135 140 Lys Gly Glu Arg Arg Arg Gly Lys Gly
His Asp Gly Leu Tyr Gln Gly 145 150 155 160 Leu Ser Thr Ala Thr Lys
Asp Thr Tyr Asp Ala Leu His Met Gln Ala 165 170 175 Leu Pro Pro Arg
180 17216PRTArtificial SequenceCD28 transmembrane domain and CD28,
OX40 and CD3 Zeta endodomains 17Phe Trp Val Leu Val Val Val Gly Gly
Val Leu Ala Cys Tyr Ser Leu 1 5 10 15 Leu Val Thr Val Ala Phe Ile
Ile Phe Trp Val Arg Ser Lys Arg Ser 20 25 30 Arg Leu Leu His Ser
Asp Tyr Met Asn Met Thr Pro Arg Arg Pro Gly 35 40 45 Pro Thr Arg
Lys His Tyr Gln Pro Tyr Ala Pro Pro Arg Asp Phe Ala 50 55 60 Ala
Tyr Arg Ser Arg Asp Gln Arg Leu Pro Pro Asp Ala His Lys Pro 65 70
75 80 Pro Gly Gly Gly Ser Phe Arg Thr Pro Ile Gln Glu Glu Gln Ala
Asp 85 90 95 Ala His Ser Thr Leu Ala Lys Ile Arg Val Lys Phe Ser
Arg Ser Ala 100 105 110 Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln
Leu Tyr Asn Glu Leu 115 120 125 Asn Leu Gly Arg Arg Glu Glu Tyr Asp
Val Leu Asp Lys Arg Arg Gly 130 135 140 Arg Asp Pro Glu Met Gly Gly
Lys Pro Arg Arg Lys Asn Pro Gln Glu 145 150 155 160 Gly Leu Tyr Asn
Glu Leu Gln Lys Asp Lys Met Ala Glu Ala Tyr Ser 165 170 175 Glu Ile
Gly Met Lys Gly Glu Arg Arg Arg Gly Lys Gly His Asp Gly 180 185 190
Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp Thr Tyr Asp Ala Leu 195
200 205 His Met Gln Ala Leu Pro Pro Arg 210 215 18729PRTArtificial
SequenceCD45 transmembrane and endodomain 18Ala Leu Ile Ala Phe Leu
Ala Phe Leu Ile Ile Val Thr Ser Ile Ala 1 5 10 15 Leu Leu Val Val
Leu Tyr Lys Ile Tyr Asp Leu His Lys Lys Arg Ser 20 25 30 Cys Asn
Leu Asp Glu Gln Gln Glu Leu Val Glu Arg Asp Asp Glu Lys 35 40 45
Gln Leu Met Asn Val Glu Pro Ile His Ala Asp Ile Leu Leu Glu Thr 50
55
60 Tyr Lys Arg Lys Ile Ala Asp Glu Gly Arg Leu Phe Leu Ala Glu Phe
65 70 75 80 Gln Ser Ile Pro Arg Val Phe Ser Lys Phe Pro Ile Lys Glu
Ala Arg 85 90 95 Lys Pro Phe Asn Gln Asn Lys Asn Arg Tyr Val Asp
Ile Leu Pro Tyr 100 105 110 Asp Tyr Asn Arg Val Glu Leu Ser Glu Ile
Asn Gly Asp Ala Gly Ser 115 120 125 Asn Tyr Ile Asn Ala Ser Tyr Ile
Asp Gly Phe Lys Glu Pro Arg Lys 130 135 140 Tyr Ile Ala Ala Gln Gly
Pro Arg Asp Glu Thr Val Asp Asp Phe Trp 145 150 155 160 Arg Met Ile
Trp Glu Gln Lys Ala Thr Val Ile Val Met Val Thr Arg 165 170 175 Cys
Glu Glu Gly Asn Arg Asn Lys Cys Ala Glu Tyr Trp Pro Ser Met 180 185
190 Glu Glu Gly Thr Arg Ala Phe Gly Asp Val Val Val Lys Ile Asn Gln
195 200 205 His Lys Arg Cys Pro Asp Tyr Ile Ile Gln Lys Leu Asn Ile
Val Asn 210 215 220 Lys Lys Glu Lys Ala Thr Gly Arg Glu Val Thr His
Ile Gln Phe Thr 225 230 235 240 Ser Trp Pro Asp His Gly Val Pro Glu
Asp Pro His Leu Leu Leu Lys 245 250 255 Leu Arg Arg Arg Val Asn Ala
Phe Ser Asn Phe Phe Ser Gly Pro Ile 260 265 270 Val Val His Cys Ser
Ala Gly Val Gly Arg Thr Gly Thr Tyr Ile Gly 275 280 285 Ile Asp Ala
Met Leu Glu Gly Leu Glu Ala Glu Asn Lys Val Asp Val 290 295 300 Tyr
Gly Tyr Val Val Lys Leu Arg Arg Gln Arg Cys Leu Met Val Gln 305 310
315 320 Val Glu Ala Gln Tyr Ile Leu Ile His Gln Ala Leu Val Glu Tyr
Asn 325 330 335 Gln Phe Gly Glu Thr Glu Val Asn Leu Ser Glu Leu His
Pro Tyr Leu 340 345 350 His Asn Met Lys Lys Arg Asp Pro Pro Ser Glu
Pro Ser Pro Leu Glu 355 360 365 Ala Glu Phe Gln Arg Leu Pro Ser Tyr
Arg Ser Trp Arg Thr Gln His 370 375 380 Ile Gly Asn Gln Glu Glu Asn
Lys Ser Lys Asn Arg Asn Ser Asn Val 385 390 395 400 Ile Pro Tyr Asp
Tyr Asn Arg Val Pro Leu Lys His Glu Leu Glu Met 405 410 415 Ser Lys
Glu Ser Glu His Asp Ser Asp Glu Ser Ser Asp Asp Asp Ser 420 425 430
Asp Ser Glu Glu Pro Ser Lys Tyr Ile Asn Ala Ser Phe Ile Met Ser 435
440 445 Tyr Trp Lys Pro Glu Val Met Ile Ala Ala Gln Gly Pro Leu Lys
Glu 450 455 460 Thr Ile Gly Asp Phe Trp Gln Met Ile Phe Gln Arg Lys
Val Lys Val 465 470 475 480 Ile Val Met Leu Thr Glu Leu Lys His Gly
Asp Gln Glu Ile Cys Ala 485 490 495 Gln Tyr Trp Gly Glu Gly Lys Gln
Thr Tyr Gly Asp Ile Glu Val Asp 500 505 510 Leu Lys Asp Thr Asp Lys
Ser Ser Thr Tyr Thr Leu Arg Val Phe Glu 515 520 525 Leu Arg His Ser
Lys Arg Lys Asp Ser Arg Thr Val Tyr Gln Tyr Gln 530 535 540 Tyr Thr
Asn Trp Ser Val Glu Gln Leu Pro Ala Glu Pro Lys Glu Leu 545 550 555
560 Ile Ser Met Ile Gln Val Val Lys Gln Lys Leu Pro Gln Lys Asn Ser
565 570 575 Ser Glu Gly Asn Lys His His Lys Ser Thr Pro Leu Leu Ile
His Cys 580 585 590 Arg Asp Gly Ser Gln Gln Thr Gly Ile Phe Cys Ala
Leu Leu Asn Leu 595 600 605 Leu Glu Ser Ala Glu Thr Glu Glu Val Val
Asp Ile Phe Gln Val Val 610 615 620 Lys Ala Leu Arg Lys Ala Arg Pro
Gly Met Val Ser Thr Phe Glu Gln 625 630 635 640 Tyr Gln Phe Leu Tyr
Asp Val Ile Ala Ser Thr Tyr Pro Ala Gln Asn 645 650 655 Gly Gln Val
Lys Lys Asn Asn His Gln Glu Asp Lys Ile Glu Phe Asp 660 665 670 Asn
Glu Val Asp Lys Val Lys Gln Asp Ala Asn Cys Val Asn Pro Leu 675 680
685 Gly Ala Pro Glu Lys Leu Pro Glu Ala Lys Glu Gln Ala Glu Gly Ser
690 695 700 Glu Pro Thr Ser Gly Thr Glu Gly Pro Glu His Ser Val Asn
Gly Pro 705 710 715 720 Ala Ser Pro Ala Leu Asn Gln Gly Ser 725
19362PRTArtificial SequenceCD148 transmembrane and endodomain 19Ala
Val Phe Gly Cys Ile Phe Gly Ala Leu Val Ile Val Thr Val Gly 1 5 10
15 Gly Phe Ile Phe Trp Arg Lys Lys Arg Lys Asp Ala Lys Asn Asn Glu
20 25 30 Val Ser Phe Ser Gln Ile Lys Pro Lys Lys Ser Lys Leu Ile
Arg Val 35 40 45 Glu Asn Phe Glu Ala Tyr Phe Lys Lys Gln Gln Ala
Asp Ser Asn Cys 50 55 60 Gly Phe Ala Glu Glu Tyr Glu Asp Leu Lys
Leu Val Gly Ile Ser Gln 65 70 75 80 Pro Lys Tyr Ala Ala Glu Leu Ala
Glu Asn Arg Gly Lys Asn Arg Tyr 85 90 95 Asn Asn Val Leu Pro Tyr
Asp Ile Ser Arg Val Lys Leu Ser Val Gln 100 105 110 Thr His Ser Thr
Asp Asp Tyr Ile Asn Ala Asn Tyr Met Pro Gly Tyr 115 120 125 His Ser
Lys Lys Asp Phe Ile Ala Thr Gln Gly Pro Leu Pro Asn Thr 130 135 140
Leu Lys Asp Phe Trp Arg Met Val Trp Glu Lys Asn Val Tyr Ala Ile 145
150 155 160 Ile Met Leu Thr Lys Cys Val Glu Gln Gly Arg Thr Lys Cys
Glu Glu 165 170 175 Tyr Trp Pro Ser Lys Gln Ala Gln Asp Tyr Gly Asp
Ile Thr Val Ala 180 185 190 Met Thr Ser Glu Ile Val Leu Pro Glu Trp
Thr Ile Arg Asp Phe Thr 195 200 205 Val Lys Asn Ile Gln Thr Ser Glu
Ser His Pro Leu Arg Gln Phe His 210 215 220 Phe Thr Ser Trp Pro Asp
His Gly Val Pro Asp Thr Thr Asp Leu Leu 225 230 235 240 Ile Asn Phe
Arg Tyr Leu Val Arg Asp Tyr Met Lys Gln Ser Pro Pro 245 250 255 Glu
Ser Pro Ile Leu Val His Cys Ser Ala Gly Val Gly Arg Thr Gly 260 265
270 Thr Phe Ile Ala Ile Asp Arg Leu Ile Tyr Gln Ile Glu Asn Glu Asn
275 280 285 Thr Val Asp Val Tyr Gly Ile Val Tyr Asp Leu Arg Met His
Arg Pro 290 295 300 Leu Met Val Gln Thr Glu Asp Gln Tyr Val Phe Leu
Asn Gln Cys Val 305 310 315 320 Leu Asp Ile Val Arg Ser Gln Lys Asp
Ser Lys Val Asp Leu Ile Tyr 325 330 335 Gln Asn Thr Thr Ala Met Thr
Ile Tyr Glu Asn Leu Ala Pro Val Thr 340 345 350 Thr Phe Gly Lys Thr
Asn Gly Tyr Ile Ala 355 360 20595PRTArtificial Sequencesequence of
PTPN6 20Met Val Arg Trp Phe His Arg Asp Leu Ser Gly Leu Asp Ala Glu
Thr 1 5 10 15 Leu Leu Lys Gly Arg Gly Val His Gly Ser Phe Leu Ala
Arg Pro Ser 20 25 30 Arg Lys Asn Gln Gly Asp Phe Ser Leu Ser Val
Arg Val Gly Asp Gln 35 40 45 Val Thr His Ile Arg Ile Gln Asn Ser
Gly Asp Phe Tyr Asp Leu Tyr 50 55 60 Gly Gly Glu Lys Phe Ala Thr
Leu Thr Glu Leu Val Glu Tyr Tyr Thr 65 70 75 80 Gln Gln Gln Gly Val
Leu Gln Asp Arg Asp Gly Thr Ile Ile His Leu 85 90 95 Lys Tyr Pro
Leu Asn Cys Ser Asp Pro Thr Ser Glu Arg Trp Tyr His 100 105 110 Gly
His Met Ser Gly Gly Gln Ala Glu Thr Leu Leu Gln Ala Lys Gly 115 120
125 Glu Pro Trp Thr Phe Leu Val Arg Glu Ser Leu Ser Gln Pro Gly Asp
130 135 140 Phe Val Leu Ser Val Leu Ser Asp Gln Pro Lys Ala Gly Pro
Gly Ser 145 150 155 160 Pro Leu Arg Val Thr His Ile Lys Val Met Cys
Glu Gly Gly Arg Tyr 165 170 175 Thr Val Gly Gly Leu Glu Thr Phe Asp
Ser Leu Thr Asp Leu Val Glu 180 185 190 His Phe Lys Lys Thr Gly Ile
Glu Glu Ala Ser Gly Ala Phe Val Tyr 195 200 205 Leu Arg Gln Pro Tyr
Tyr Ala Thr Arg Val Asn Ala Ala Asp Ile Glu 210 215 220 Asn Arg Val
Leu Glu Leu Asn Lys Lys Gln Glu Ser Glu Asp Thr Ala 225 230 235 240
Lys Ala Gly Phe Trp Glu Glu Phe Glu Ser Leu Gln Lys Gln Glu Val 245
250 255 Lys Asn Leu His Gln Arg Leu Glu Gly Gln Arg Pro Glu Asn Lys
Gly 260 265 270 Lys Asn Arg Tyr Lys Asn Ile Leu Pro Phe Asp His Ser
Arg Val Ile 275 280 285 Leu Gln Gly Arg Asp Ser Asn Ile Pro Gly Ser
Asp Tyr Ile Asn Ala 290 295 300 Asn Tyr Ile Lys Asn Gln Leu Leu Gly
Pro Asp Glu Asn Ala Lys Thr 305 310 315 320 Tyr Ile Ala Ser Gln Gly
Cys Leu Glu Ala Thr Val Asn Asp Phe Trp 325 330 335 Gln Met Ala Trp
Gln Glu Asn Ser Arg Val Ile Val Met Thr Thr Arg 340 345 350 Glu Val
Glu Lys Gly Arg Asn Lys Cys Val Pro Tyr Trp Pro Glu Val 355 360 365
Gly Met Gln Arg Ala Tyr Gly Pro Tyr Ser Val Thr Asn Cys Gly Glu 370
375 380 His Asp Thr Thr Glu Tyr Lys Leu Arg Thr Leu Gln Val Ser Pro
Leu 385 390 395 400 Asp Asn Gly Asp Leu Ile Arg Glu Ile Trp His Tyr
Gln Tyr Leu Ser 405 410 415 Trp Pro Asp His Gly Val Pro Ser Glu Pro
Gly Gly Val Leu Ser Phe 420 425 430 Leu Asp Gln Ile Asn Gln Arg Gln
Glu Ser Leu Pro His Ala Gly Pro 435 440 445 Ile Ile Val His Cys Ser
Ala Gly Ile Gly Arg Thr Gly Thr Ile Ile 450 455 460 Val Ile Asp Met
Leu Met Glu Asn Ile Ser Thr Lys Gly Leu Asp Cys 465 470 475 480 Asp
Ile Asp Ile Gln Lys Thr Ile Gln Met Val Arg Ala Gln Arg Ser 485 490
495 Gly Met Val Gln Thr Glu Ala Gln Tyr Lys Phe Ile Tyr Val Ala Ile
500 505 510 Ala Gln Phe Ile Glu Thr Thr Lys Lys Lys Leu Glu Val Leu
Gln Ser 515 520 525 Gln Lys Gly Gln Glu Ser Glu Tyr Gly Asn Ile Thr
Tyr Pro Pro Ala 530 535 540 Met Lys Asn Ala His Ala Lys Ala Ser Arg
Thr Ser Ser Lys His Lys 545 550 555 560 Glu Asp Val Tyr Glu Asn Leu
His Thr Lys Asn Lys Arg Glu Glu Lys 565 570 575 Val Lys Lys Gln Arg
Ser Ala Asp Lys Glu Lys Ser Lys Gly Ser Leu 580 585 590 Lys Arg Lys
595 21272PRTArtificial Sequencesequence of phosphatase domain of
PTPN6 21Phe Trp Glu Glu Phe Glu Ser Leu Gln Lys Gln Glu Val Lys Asn
Leu 1 5 10 15 His Gln Arg Leu Glu Gly Gln Arg Pro Glu Asn Lys Gly
Lys Asn Arg 20 25 30 Tyr Lys Asn Ile Leu Pro Phe Asp His Ser Arg
Val Ile Leu Gln Gly 35 40 45 Arg Asp Ser Asn Ile Pro Gly Ser Asp
Tyr Ile Asn Ala Asn Tyr Ile 50 55 60 Lys Asn Gln Leu Leu Gly Pro
Asp Glu Asn Ala Lys Thr Tyr Ile Ala 65 70 75 80 Ser Gln Gly Cys Leu
Glu Ala Thr Val Asn Asp Phe Trp Gln Met Ala 85 90 95 Trp Gln Glu
Asn Ser Arg Val Ile Val Met Thr Thr Arg Glu Val Glu 100 105 110 Lys
Gly Arg Asn Lys Cys Val Pro Tyr Trp Pro Glu Val Gly Met Gln 115 120
125 Arg Ala Tyr Gly Pro Tyr Ser Val Thr Asn Cys Gly Glu His Asp Thr
130 135 140 Thr Glu Tyr Lys Leu Arg Thr Leu Gln Val Ser Pro Leu Asp
Asn Gly 145 150 155 160 Asp Leu Ile Arg Glu Ile Trp His Tyr Gln Tyr
Leu Ser Trp Pro Asp 165 170 175 His Gly Val Pro Ser Glu Pro Gly Gly
Val Leu Ser Phe Leu Asp Gln 180 185 190 Ile Asn Gln Arg Gln Glu Ser
Leu Pro His Ala Gly Pro Ile Ile Val 195 200 205 His Cys Ser Ala Gly
Ile Gly Arg Thr Gly Thr Ile Ile Val Ile Asp 210 215 220 Met Leu Met
Glu Asn Ile Ser Thr Lys Gly Leu Asp Cys Asp Ile Asp 225 230 235 240
Ile Gln Lys Thr Ile Gln Met Val Arg Ala Gln Arg Ser Gly Met Val 245
250 255 Gln Thr Glu Ala Gln Tyr Lys Phe Ile Tyr Val Ala Ile Ala Gln
Phe 260 265 270 2297PRTArtificial SequencePDCD1 endodomain 22Cys
Ser Arg Ala Ala Arg Gly Thr Ile Gly Ala Arg Arg Thr Gly Gln 1 5 10
15 Pro Leu Lys Glu Asp Pro Ser Ala Val Pro Val Phe Ser Val Asp Tyr
20 25 30 Gly Glu Leu Asp Phe Gln Trp Arg Glu Lys Thr Pro Glu Pro
Pro Val 35 40 45 Pro Cys Val Pro Glu Gln Thr Glu Tyr Ala Thr Ile
Val Phe Pro Ser 50 55 60 Gly Met Gly Thr Ser Ser Pro Ala Arg Arg
Gly Ser Ala Asp Gly Pro 65 70 75 80 Arg Ser Ala Gln Pro Leu Arg Pro
Glu Asp Gly His Cys Ser Trp Pro 85 90 95 Leu 23141PRTArtificial
SequenceBTLA4 endodomain 23Lys Leu Gln Arg Arg Trp Lys Arg Thr Gln
Ser Gln Gln Gly Leu Gln 1 5 10 15 Glu Asn Ser Ser Gly Gln Ser Phe
Phe Val Arg Asn Lys Lys Val Arg 20 25 30 Arg Ala Pro Leu Ser Glu
Gly Pro His Ser Leu Gly Cys Tyr Asn Pro 35 40 45 Met Met Glu Asp
Gly Ile Ser Tyr Thr Thr Leu Arg Phe Pro Glu Met 50 55 60 Asn Ile
Pro Arg Thr Gly Asp Ala Glu Ser Ser Glu Met Gln Arg Pro 65 70 75 80
Pro Pro Asp Cys Asp Asp Thr Val Thr Tyr Ser Ala Leu His Lys Arg 85
90 95 Gln Val Gly Asp Tyr Glu Asn Val Ile Pro Asp Phe Pro Glu Asp
Glu 100 105 110 Gly Ile His Tyr Ser Glu Leu Ile Gln Phe Gly Val Gly
Glu Arg Pro 115 120 125 Gln Ala Gln Glu Asn Val Asp Tyr Val Ile Leu
Lys His 130 135 140 24168PRTArtificial SequenceLILRB1 endodomain
24Leu Arg His Arg Arg Gln Gly Lys His Trp Thr Ser Thr Gln Arg Lys 1
5 10 15 Ala Asp Phe Gln His Pro Ala Gly Ala Val Gly Pro Glu Pro Thr
Asp 20 25 30 Arg Gly Leu Gln Trp Arg Ser Ser Pro Ala Ala Asp Ala
Gln Glu Glu 35 40 45 Asn Leu Tyr Ala Ala Val Lys His Thr Gln Pro
Glu Asp Gly Val Glu 50 55 60 Met Asp Thr Arg Ser Pro His Asp Glu
Asp Pro Gln Ala Val Thr Tyr 65 70 75 80 Ala Glu Val Lys His Ser Arg
Pro Arg Arg Glu Met Ala Ser Pro Pro 85 90 95 Ser Pro Leu Ser Gly
Glu Phe Leu Asp Thr Lys Asp Arg Gln Ala Glu 100 105 110 Glu Asp Arg
Gln Met Asp Thr Glu Ala Ala Ala Ser Glu Ala Pro Gln 115 120
125 Asp Val Thr Tyr Ala Gln Leu His Ser Leu Thr Leu Arg Arg Glu Ala
130 135 140 Thr Glu Pro Pro Pro Ser Gln Glu Gly Pro Ser Pro Ala Val
Pro Ser 145 150 155 160 Ile Tyr Ala Thr Leu Ala Ile His 165
25101PRTArtificial SequenceLAIR1 endodomain 25His Arg Gln Asn Gln
Ile Lys Gln Gly Pro Pro Arg Ser Lys Asp Glu 1 5 10 15 Glu Gln Lys
Pro Gln Gln Arg Pro Asp Leu Ala Val Asp Val Leu Glu 20 25 30 Arg
Thr Ala Asp Lys Ala Thr Val Asn Gly Leu Pro Glu Lys Asp Arg 35 40
45 Glu Thr Asp Thr Ser Ala Leu Ala Ala Gly Ser Ser Gln Glu Val Thr
50 55 60 Tyr Ala Gln Leu Asp His Trp Ala Leu Thr Gln Arg Thr Ala
Arg Ala 65 70 75 80 Val Ser Pro Gln Ser Thr Lys Pro Met Ala Glu Ser
Ile Thr Tyr Ala 85 90 95 Ala Val Ala Arg His 100 2662PRTArtificial
SequenceCTLA4 endodomain 26Phe Leu Leu Trp Ile Leu Ala Ala Val Ser
Ser Gly Leu Phe Phe Tyr 1 5 10 15 Ser Phe Leu Leu Thr Ala Val Ser
Leu Ser Lys Met Leu Lys Lys Arg 20 25 30 Ser Pro Leu Thr Thr Gly
Val Tyr Val Lys Met Pro Pro Thr Glu Pro 35 40 45 Glu Cys Glu Lys
Gln Phe Gln Pro Tyr Phe Ile Pro Ile Asn 50 55 60 27111PRTArtificial
SequenceKIR2DL1 endodomain 27Gly Asn Ser Arg His Leu His Val Leu
Ile Gly Thr Ser Val Val Ile 1 5 10 15 Ile Pro Phe Ala Ile Leu Leu
Phe Phe Leu Leu His Arg Trp Cys Ala 20 25 30 Asn Lys Lys Asn Ala
Val Val Met Asp Gln Glu Pro Ala Gly Asn Arg 35 40 45 Thr Val Asn
Arg Glu Asp Ser Asp Glu Gln Asp Pro Gln Glu Val Thr 50 55 60 Tyr
Thr Gln Leu Asn His Cys Val Phe Thr Gln Arg Lys Ile Thr Arg 65 70
75 80 Pro Ser Gln Arg Pro Lys Thr Pro Pro Thr Asp Ile Ile Val Tyr
Thr 85 90 95 Glu Leu Pro Asn Ala Glu Ser Arg Ser Lys Val Val Ser
Cys Pro 100 105 110 28143PRTArtificial SequenceKIR2DL4 endodomain
28Gly Ile Ala Arg His Leu His Ala Val Ile Arg Tyr Ser Val Ala Ile 1
5 10 15 Ile Leu Phe Thr Ile Leu Pro Phe Phe Leu Leu His Arg Trp Cys
Ser 20 25 30 Lys Lys Lys Glu Asn Ala Ala Val Met Asn Gln Glu Pro
Ala Gly His 35 40 45 Arg Thr Val Asn Arg Glu Asp Ser Asp Glu Gln
Asp Pro Gln Glu Val 50 55 60 Thr Tyr Ala Gln Leu Asp His Cys Ile
Phe Thr Gln Arg Lys Ile Thr 65 70 75 80 Gly Pro Ser Gln Arg Ser Lys
Arg Pro Ser Thr Asp Thr Ser Val Cys 85 90 95 Ile Glu Leu Pro Asn
Ala Glu Pro Arg Ala Leu Ser Pro Ala His Glu 100 105 110 His His Ser
Gln Ala Leu Met Gly Ser Ser Arg Glu Thr Thr Ala Leu 115 120 125 Ser
Gln Thr Gln Leu Ala Ser Ser Asn Val Pro Ala Ala Gly Ile 130 135 140
29143PRTArtificial SequenceKIR2DL5 endodomain 29Thr Gly Ile Arg Arg
His Leu His Ile Leu Ile Gly Thr Ser Val Ala 1 5 10 15 Ile Ile Leu
Phe Ile Ile Leu Phe Phe Phe Leu Leu His Cys Cys Cys 20 25 30 Ser
Asn Lys Lys Asn Ala Ala Val Met Asp Gln Glu Pro Ala Gly Asp 35 40
45 Arg Thr Val Asn Arg Glu Asp Ser Asp Asp Gln Asp Pro Gln Glu Val
50 55 60 Thr Tyr Ala Gln Leu Asp His Cys Val Phe Thr Gln Thr Lys
Ile Thr 65 70 75 80 Ser Pro Ser Gln Arg Pro Lys Thr Pro Pro Thr Asp
Thr Thr Met Tyr 85 90 95 Met Glu Leu Pro Asn Ala Lys Pro Arg Ser
Leu Ser Pro Ala His Lys 100 105 110 His His Ser Gln Ala Leu Arg Gly
Ser Ser Arg Glu Thr Thr Ala Leu 115 120 125 Ser Gln Asn Arg Val Ala
Ser Ser His Val Pro Ala Ala Gly Ile 130 135 140 30111PRTArtificial
SequenceKIR3DL1 endodomain 30Lys Asp Pro Arg His Leu His Ile Leu
Ile Gly Thr Ser Val Val Ile 1 5 10 15 Ile Leu Phe Ile Leu Leu Leu
Phe Phe Leu Leu His Leu Trp Cys Ser 20 25 30 Asn Lys Lys Asn Ala
Ala Val Met Asp Gln Glu Pro Ala Gly Asn Arg 35 40 45 Thr Ala Asn
Ser Glu Asp Ser Asp Glu Gln Asp Pro Glu Glu Val Thr 50 55 60 Tyr
Ala Gln Leu Asp His Cys Val Phe Thr Gln Arg Lys Ile Thr Arg 65 70
75 80 Pro Ser Gln Arg Pro Lys Thr Pro Pro Thr Asp Thr Ile Leu Tyr
Thr 85 90 95 Glu Leu Pro Asn Ala Lys Pro Arg Ser Lys Val Val Ser
Cys Pro 100 105 110 3197PRTArtificial SequenceKIR3DL3 endodomain
31Lys Asp Pro Gly Asn Ser Arg His Leu His Val Leu Ile Gly Thr Ser 1
5 10 15 Val Val Ile Ile Pro Phe Ala Ile Leu Leu Phe Phe Leu Leu His
Arg 20 25 30 Trp Cys Ala Asn Lys Lys Asn Ala Val Val Met Asp Gln
Glu Pro Ala 35 40 45 Gly Asn Arg Thr Val Asn Arg Glu Asp Ser Asp
Glu Gln Asp Pro Gln 50 55 60 Glu Val Thr Tyr Ala Gln Leu Asn His
Cys Val Phe Thr Gln Arg Lys 65 70 75 80 Ile Thr Arg Pro Ser Gln Arg
Pro Lys Thr Pro Pro Thr Asp Thr Ser 85 90 95 Val 32807PRTArtificial
SequencePTPN6-CD45 fusion protein 32Trp Tyr His Gly His Met Ser Gly
Gly Gln Ala Glu Thr Leu Leu Gln 1 5 10 15 Ala Lys Gly Glu Pro Trp
Thr Phe Leu Val Arg Glu Ser Leu Ser Gln 20 25 30 Pro Gly Asp Phe
Val Leu Ser Val Leu Ser Asp Gln Pro Lys Ala Gly 35 40 45 Pro Gly
Ser Pro Leu Arg Val Thr His Ile Lys Val Met Cys Glu Gly 50 55 60
Gly Arg Tyr Thr Val Gly Gly Leu Glu Thr Phe Asp Ser Leu Thr Asp 65
70 75 80 Leu Val Glu His Phe Lys Lys Thr Gly Ile Glu Glu Ala Ser
Gly Ala 85 90 95 Phe Val Tyr Leu Arg Gln Pro Tyr Lys Ile Tyr Asp
Leu His Lys Lys 100 105 110 Arg Ser Cys Asn Leu Asp Glu Gln Gln Glu
Leu Val Glu Arg Asp Asp 115 120 125 Glu Lys Gln Leu Met Asn Val Glu
Pro Ile His Ala Asp Ile Leu Leu 130 135 140 Glu Thr Tyr Lys Arg Lys
Ile Ala Asp Glu Gly Arg Leu Phe Leu Ala 145 150 155 160 Glu Phe Gln
Ser Ile Pro Arg Val Phe Ser Lys Phe Pro Ile Lys Glu 165 170 175 Ala
Arg Lys Pro Phe Asn Gln Asn Lys Asn Arg Tyr Val Asp Ile Leu 180 185
190 Pro Tyr Asp Tyr Asn Arg Val Glu Leu Ser Glu Ile Asn Gly Asp Ala
195 200 205 Gly Ser Asn Tyr Ile Asn Ala Ser Tyr Ile Asp Gly Phe Lys
Glu Pro 210 215 220 Arg Lys Tyr Ile Ala Ala Gln Gly Pro Arg Asp Glu
Thr Val Asp Asp 225 230 235 240 Phe Trp Arg Met Ile Trp Glu Gln Lys
Ala Thr Val Ile Val Met Val 245 250 255 Thr Arg Cys Glu Glu Gly Asn
Arg Asn Lys Cys Ala Glu Tyr Trp Pro 260 265 270 Ser Met Glu Glu Gly
Thr Arg Ala Phe Gly Asp Val Val Val Lys Ile 275 280 285 Asn Gln His
Lys Arg Cys Pro Asp Tyr Ile Ile Gln Lys Leu Asn Ile 290 295 300 Val
Asn Lys Lys Glu Lys Ala Thr Gly Arg Glu Val Thr His Ile Gln 305 310
315 320 Phe Thr Ser Trp Pro Asp His Gly Val Pro Glu Asp Pro His Leu
Leu 325 330 335 Leu Lys Leu Arg Arg Arg Val Asn Ala Phe Ser Asn Phe
Phe Ser Gly 340 345 350 Pro Ile Val Val His Cys Ser Ala Gly Val Gly
Arg Thr Gly Thr Tyr 355 360 365 Ile Gly Ile Asp Ala Met Leu Glu Gly
Leu Glu Ala Glu Asn Lys Val 370 375 380 Asp Val Tyr Gly Tyr Val Val
Lys Leu Arg Arg Gln Arg Cys Leu Met 385 390 395 400 Val Gln Val Glu
Ala Gln Tyr Ile Leu Ile His Gln Ala Leu Val Glu 405 410 415 Tyr Asn
Gln Phe Gly Glu Thr Glu Val Asn Leu Ser Glu Leu His Pro 420 425 430
Tyr Leu His Asn Met Lys Lys Arg Asp Pro Pro Ser Glu Pro Ser Pro 435
440 445 Leu Glu Ala Glu Phe Gln Arg Leu Pro Ser Tyr Arg Ser Trp Arg
Thr 450 455 460 Gln His Ile Gly Asn Gln Glu Glu Asn Lys Ser Lys Asn
Arg Asn Ser 465 470 475 480 Asn Val Ile Pro Tyr Asp Tyr Asn Arg Val
Leu Lys His Glu Leu Glu 485 490 495 Met Ser Lys Glu Ser Glu His Asp
Ser Asp Glu Ser Ser Asp Asp Asp 500 505 510 Ser Asp Ser Glu Glu Pro
Ser Lys Tyr Ile Asn Ala Ser Phe Ile Met 515 520 525 Ser Tyr Trp Lys
Pro Glu Val Met Ile Ala Ala Gln Gly Pro Leu Lys 530 535 540 Glu Thr
Ile Gly Asp Phe Met Ile Gln Arg Lys Val Lys Val Ile Val 545 550 555
560 Met Leu Thr Glu Leu Lys His Gly Asp Gln Glu Ile Cys Ala Gln Tyr
565 570 575 Trp Gly Glu Gly Lys Gln Thr Tyr Gly Asp Ile Glu Val Asp
Leu Lys 580 585 590 Asp Thr Asp Lys Ser Ser Thr Tyr Thr Leu Arg Val
Phe Glu Leu Arg 595 600 605 His Ser Lys Arg Lys Asp Ser Arg Thr Val
Tyr Gln Tyr Gln Tyr Thr 610 615 620 Asn Trp Ser Val Glu Gln Leu Pro
Ala Glu Pro Lys Glu Leu Ile Ser 625 630 635 640 Met Ile Gln Val Val
Lys Gln Lys Leu Pro Gln Lys Asn Ser Ser Glu 645 650 655 Gly Asn Lys
His His Lys Ser Thr Pro Leu Leu Ile His Cys Arg Asp 660 665 670 Gly
Ser Gln Gln Thr Gly Ile Phe Cys Ala Leu Leu Asn Leu Leu Glu 675 680
685 Ser Ala Glu Thr Glu Glu Val Val Asp Ile Phe Gln Val Val Lys Ala
690 695 700 Leu Arg Lys Ala Arg Pro Gly Met Val Ser Thr Phe Glu Gln
Tyr Gln 705 710 715 720 Phe Leu Tyr Asp Val Ile Ala Ser Thr Tyr Pro
Ala Gln Asn Gly Gln 725 730 735 Val Lys Lys Asn Asn His Gln Glu Asp
Lys Ile Glu Phe Asp Asn Glu 740 745 750 Val Asp Lys Val Lys Gln Asp
Ala Asn Cys Val Asn Pro Leu Gly Ala 755 760 765 Pro Glu Lys Leu Pro
Glu Ala Lys Glu Gln Ala Glu Gly Ser Glu Pro 770 775 780 Thr Ser Gly
Thr Glu Gly Pro Glu His Ser Val Asn Gly Pro Ala Ser 785 790 795 800
Pro Ala Leu Asn Gln Gly Ser 805 33434PRTArtificial
SequencePTPN6-CD148 fusion protein 33Glu Thr Leu Leu Gln Ala Lys
Gly Glu Pro Trp Thr Phe Leu Val Arg 1 5 10 15 Glu Ser Leu Ser Gln
Pro Gly Asp Phe Val Leu Ser Val Leu Ser Asp 20 25 30 Gln Pro Lys
Ala Gly Pro Gly Ser Pro Leu Arg Val Thr His Ile Lys 35 40 45 Val
Met Cys Glu Gly Gly Arg Tyr Thr Val Gly Gly Leu Glu Thr Phe 50 55
60 Asp Ser Leu Thr Asp Leu Val Glu His Phe Lys Lys Thr Gly Ile Glu
65 70 75 80 Glu Ala Ser Gly Ala Phe Val Tyr Leu Arg Gln Pro Tyr Arg
Lys Lys 85 90 95 Arg Lys Asp Ala Lys Asn Asn Glu Val Ser Phe Ser
Gln Ile Lys Pro 100 105 110 Lys Lys Ser Lys Leu Ile Arg Val Glu Asn
Phe Glu Ala Tyr Phe Lys 115 120 125 Lys Gln Gln Ala Asp Ser Asn Cys
Gly Phe Ala Glu Glu Tyr Glu Asp 130 135 140 Leu Lys Leu Val Gly Ile
Ser Gln Pro Lys Tyr Ala Ala Glu Leu Ala 145 150 155 160 Glu Asn Arg
Gly Lys Asn Arg Tyr Asn Asn Val Leu Pro Tyr Asp Ile 165 170 175 Ser
Arg Val Lys Leu Ser Val Gln Thr His Ser Thr Asp Asp Tyr Ile 180 185
190 Asn Ala Asn Tyr Met Pro Gly Tyr His Ser Lys Lys Asp Phe Ile Ala
195 200 205 Thr Gln Gly Pro Leu Pro Asn Thr Leu Lys Asp Phe Trp Arg
Met Val 210 215 220 Trp Glu Lys Asn Val Tyr Ala Ile Ile Met Leu Thr
Lys Cys Val Glu 225 230 235 240 Gln Gly Arg Thr Lys Cys Glu Glu Tyr
Trp Pro Ser Lys Gln Ala Gln 245 250 255 Asp Tyr Gly Asp Ile Thr Val
Ala Met Thr Ser Glu Ile Val Leu Pro 260 265 270 Glu Trp Thr Ile Arg
Asp Phe Thr Val Lys Asn Ile Gln Thr Ser Glu 275 280 285 Ser His Pro
Leu Arg Gln Phe His Phe Thr Ser Trp Pro Asp His Gly 290 295 300 Val
Pro Asp Thr Thr Asp Leu Leu Ile Asn Phe Arg Tyr Leu Val Arg 305 310
315 320 Asp Tyr Met Lys Gln Ser Pro Pro Glu Ser Pro Ile Leu Val His
Cys 325 330 335 Ser Ala Gly Val Gly Arg Thr Gly Thr Phe Ile Ala Ile
Asp Arg Leu 340 345 350 Ile Tyr Gln Ile Glu Asn Glu Asn Thr Val Asp
Val Tyr Gly Ile Val 355 360 365 Tyr Asp Leu Arg Met His Arg Pro Leu
Met Val Gln Thr Glu Asp Gln 370 375 380 Tyr Val Phe Leu Asn Gln Cys
Val Leu Asp Ile Val Arg Ser Gln Lys 385 390 395 400 Asp Ser Lys Val
Asp Leu Ile Tyr Gln Asn Thr Thr Ala Met Thr Ile 405 410 415 Tyr Glu
Asn Leu Ala Pro Val Thr Thr Phe Gly Lys Thr Asn Gly Tyr 420 425 430
Ile Ala 3420PRTFoot-and-mouth disease virus 34Arg Ala Glu Gly Arg
Gly Ser Leu Leu Thr Cys Gly Asp Val Glu Glu 1 5 10 15 Asn Pro Gly
Pro 20 353390DNAArtificial SequenceNucleic acid sequences coding
for CARs
(MP13974.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-HCH2CH3pvaa-CD
28tmZw) 35atgagcctgc ccgtgaccgc cctgctgctg cccctggccc tgctgctgca
cgccgccaga 60ccagacatcc agatgaccca gaccaccagc agcctgagcg ccagcctggg
cgaccgggtg 120accatcagct gcagagccag ccaggacatc agcaagtacc
tgaactggta ccagcagaag 180cccgacggca ccgtgaagct gctgatctac
cacaccagcc ggctgcacag cggcgtgccc 240agccggttca gcggcagcgg
cagcggcacc gactacagcc tgaccatcag caacctggag 300caggaggaca
tcgccaccta cttctgccag cagggcaaca ccctgcccta caccttcgga
360ggcggcacca agctggagat caccaaggcc ggaggcggag gctctggcgg
aggcggctct 420ggcggaggcg gctctggcgg aggcggcagc gaggtgaagc
tgcaggagtc tggcccaggc 480ctggtggccc caagccagag cctgagcgtg
acctgcaccg tgagcggcgt gagcctgccc 540gactacggcg tgagctggat
caggcagccc ccacggaagg gcctggagtg gctgggcgtg 600atctggggca
gcgagaccac ctactacaac agcgccctga agagccggct gaccatcatc
660aaggacaaca gcaagagcca ggtgttcctg aagatgaaca gcctgcagac
cgacgacacc 720gccatctact actgcgccaa gcactactac tatggcggca
gctacgctat ggactactgg 780ggccagggca ccagcgtgac cgtgagctca
gatcccacca cgacgccagc gccgcgacca 840ccaacaccgg cgcccaccat
cgcgtcgcag cccctgtccc tgcgcccaga
ggcgtgccgg 900ccagcggcgg ggggcgcagt gcacacgagg gggctggact
tcgcctgtga tatcttttgg 960gtgctggtgg tggttggtgg agtcctggct
tgctatagct tgctagtaac agtggccttt 1020attattttct gggtgaggag
agtgaagttc agcaggagcg cagacgcccc cgcgtaccag 1080cagggccaga
accagctcta taacgagctc aatctaggac gaagagagga gtacgatgtt
1140ttggacaaga gacgtggccg ggaccctgag atggggggaa agccgagaag
gaagaaccct 1200caggaaggcc tgtacaatga actgcagaaa gataagatgg
cggaggccta cagtgagatt 1260gggatgaaag gcgagcgccg gaggggcaag
gggcacgatg gcctttacca gggtctcagt 1320acagccacca aggacaccta
cgacgccctt cacatgcagg ccctgcctcc tcgcagagcc 1380gagggcaggg
gaagtcttct aacatgcggg gacgtggagg aaaatcccgg gcccatggcc
1440gtgcccactc aggtcctggg gttgttgcta ctgtggctta cagatgccag
atgtgacatc 1500cagatgacac agtctccatc ttccctgtct gcatctgtcg
gagatcgcgt caccatcacc 1560tgtcgagcaa gtgaggacat ttattttaat
ttagtgtggt atcagcagaa accaggaaag 1620gcccctaagc tcctgatcta
tgatacaaat cgcttggcag atggggtccc atcacggttc 1680agtggctctg
gatctggcac acagtatact ctaaccataa gtagcctgca acccgaagat
1740ttcgcaacct attattgtca acactataag aattatccgc tcacgttcgg
tcaggggacc 1800aagctggaaa tcaaaagatc tggtggcgga gggtcaggag
gcggaggcag cggaggcggt 1860ggctcgggag gcggaggctc gagatctgag
gtgcagttgg tggagtctgg gggcggcttg 1920gtgcagcctg gagggtccct
gaggctctcc tgtgcagcct caggattcac tctcagtaat 1980tatggcatgc
actggatcag gcaggctcca gggaagggtc tggagtgggt ctcgtctatt
2040agtcttaatg gtggtagcac ttactatcga gactccgtga agggccgatt
cactatctcc 2100agggacaatg caaaaagcac cctctacctt caaatgaata
gtctgagggc cgaggacacg 2160gccgtctatt actgtgcagc acaggacgct
tatacgggag gttactttga ttactggggc 2220caaggaacgc tggtcacagt
ctcgtctatg gatcccgccg agcccaaatc tcctgacaaa 2280actcacacat
gcccaccgtg cccagcacct cccgtggccg gcccgtcagt cttcctcttc
2340cccccaaaac ccaaggacac cctcatgatc gcccggaccc ctgaggtcac
atgcgtggtg 2400gtggacgtga gccacgaaga ccctgaggtc aagttcaact
ggtacgtgga cggcgtggag 2460gtgcataatg ccaagacaaa gccgcgggag
gagcagtaca acagcacgta ccgtgtggtc 2520agcgtcctca ccgtcctgca
ccaggactgg ctgaatggca aggagtacaa gtgcaaggtc 2580tccaacaaag
ccctcccagc ccccatcgag aaaaccatct ccaaagccaa agggcagccc
2640cgagaaccac aggtgtacac cctgccccca tcccgggatg agctgaccaa
gaaccaggtc 2700agcctgacct gcctggtcaa aggcttctat cccagcgaca
tcgccgtgga gtgggagagc 2760aatgggcaac cggagaacaa ctacaagacc
acgcctcccg tgctggactc cgacggctcc 2820ttcttcctct acagcaagct
caccgtggac aagagcaggt ggcagcaggg gaacgtcttc 2880tcatgctccg
tgatgcatga ggccctgcac aatcactata cccagaaatc tctgagtctg
2940agcccaggca agaaggaccc caagttctgg gtcctggtgg tggtgggagg
cgtgctggcc 3000tgttactctc tcctggtgac cgtggccttc atcatctttt
gggtgcgctc ccgggtgaag 3060ttttctcgct ctgccgatgc cccagcctat
cagcagggcc agaatcagct gtacaatgaa 3120ctgaacctgg gcaggcggga
ggagtacgac gtgctggata agcggagagg cagagacccc 3180gagatgggcg
gcaaaccacg gcgcaaaaat ccccaggagg gactctataa cgagctgcag
3240aaggacaaaa tggccgaggc ctattccgag atcggcatga agggagagag
aagacgcgga 3300aagggccacg acggcctgta tcagggattg tccaccgcta
caaaagatac atatgatgcc 3360ctgcacatgc aggccctgcc acccagatga
3390365154DNAArtificial SequenceNucleic acid sequences coding for
CARs
(MP14802.SFG.aCD19fmc63_clean-CD8STK-CD28tmZ-2A-aCD33glx-HCH2CH3p
vaa-dCD45) 36atgagcctgc ccgtgaccgc cctgctgctg cccctggccc tgctgctgca
cgccgccaga 60ccagacatcc agatgaccca gaccaccagc agcctgagcg ccagcctggg
cgaccgggtg 120accatcagct gcagagccag ccaggacatc agcaagtacc
tgaactggta ccagcagaag 180cccgacggca ccgtgaagct gctgatctac
cacaccagcc ggctgcacag cggcgtgccc 240agccggttca gcggcagcgg
cagcggcacc gactacagcc tgaccatcag caacctggag 300caggaggaca
tcgccaccta cttctgccag cagggcaaca ccctgcccta caccttcgga
360ggcggcacca agctggagat caccaaggcc ggaggcggag gctctggcgg
aggcggctct 420ggcggaggcg gctctggcgg aggcggcagc gaggtgaagc
tgcaggagtc tggcccaggc 480ctggtggccc caagccagag cctgagcgtg
acctgcaccg tgagcggcgt gagcctgccc 540gactacggcg tgagctggat
caggcagccc ccacggaagg gcctggagtg gctgggcgtg 600atctggggca
gcgagaccac ctactacaac agcgccctga agagccggct gaccatcatc
660aaggacaaca gcaagagcca ggtgttcctg aagatgaaca gcctgcagac
cgacgacacc 720gccatctact actgcgccaa gcactactac tatggcggca
gctacgctat ggactactgg 780ggccagggca ccagcgtgac cgtgagctca
gatcccacca cgacgccagc gccgcgacca 840ccaacaccgg cgcccaccat
cgcgtcgcag cccctgtccc tgcgcccaga ggcgtgccgg 900ccagcggcgg
ggggcgcagt gcacacgagg gggctggact tcgcctgtga tatcttttgg
960gtgctggtgg tggttggtgg agtcctggct tgctatagct tgctagtaac
agtggccttt 1020attattttct gggtgaggag agtgaagttc agcaggagcg
cagacgcccc cgcgtaccag 1080cagggccaga accagctcta taacgagctc
aatctaggac gaagagagga gtacgatgtt 1140ttggacaaga gacgtggccg
ggaccctgag atggggggaa agccgagaag gaagaaccct 1200caggaaggcc
tgtacaatga actgcagaaa gataagatgg cggaggccta cagtgagatt
1260gggatgaaag gcgagcgccg gaggggcaag gggcacgatg gcctttacca
gggtctcagt 1320acagccacca aggacaccta cgacgccctt cacatgcagg
ccctgcctcc tcgcagagcc 1380gagggcaggg gaagtcttct aacatgcggg
gacgtggagg aaaatcccgg gcccatggcc 1440gtgcccactc aggtcctggg
gttgttgcta ctgtggctta cagatgccag atgtgacatc 1500cagatgacac
agtctccatc ttccctgtct gcatctgtcg gagatcgcgt caccatcacc
1560tgtcgagcaa gtgaggacat ttattttaat ttagtgtggt atcagcagaa
accaggaaag 1620gcccctaagc tcctgatcta tgatacaaat cgcttggcag
atggggtccc atcacggttc 1680agtggctctg gatctggcac acagtatact
ctaaccataa gtagcctgca acccgaagat 1740ttcgcaacct attattgtca
acactataag aattatccgc tcacgttcgg tcaggggacc 1800aagctggaaa
tcaaaagatc tggtggcgga gggtcaggag gcggaggcag cggaggcggt
1860ggctcgggag gcggaggctc gagatctgag gtgcagttgg tggagtctgg
gggcggcttg 1920gtgcagcctg gagggtccct gaggctctcc tgtgcagcct
caggattcac tctcagtaat 1980tatggcatgc actggatcag gcaggctcca
gggaagggtc tggagtgggt ctcgtctatt 2040agtcttaatg gtggtagcac
ttactatcga gactccgtga agggccgatt cactatctcc 2100agggacaatg
caaaaagcac cctctacctt caaatgaata gtctgagggc cgaggacacg
2160gccgtctatt actgtgcagc acaggacgct tatacgggag gttactttga
ttactggggc 2220caaggaacgc tggtcacagt ctcgtctatg gatcccgccg
agcccaaatc tcctgacaaa 2280actcacacat gcccaccgtg cccagcacct
cccgtggccg gcccgtcagt cttcctcttc 2340cccccaaaac ccaaggacac
cctcatgatc gcccggaccc ctgaggtcac atgcgtggtg 2400gtggacgtga
gccacgaaga ccctgaggtc aagttcaact ggtacgtgga cggcgtggag
2460gtgcataatg ccaagacaaa gccgcgggag gagcagtaca acagcacgta
ccgtgtggtc 2520agcgtcctca ccgtcctgca ccaggactgg ctgaatggca
aggagtacaa gtgcaaggtc 2580tccaacaaag ccctcccagc ccccatcgag
aaaaccatct ccaaagccaa agggcagccc 2640cgagaaccac aggtgtacac
cctgccccca tcccgggatg agctgaccaa gaaccaggtc 2700agcctgacct
gcctggtcaa aggcttctat cccagcgaca tcgccgtgga gtgggagagc
2760aatgggcaac cggagaacaa ctacaagacc acgcctcccg tgctggactc
cgacggctcc 2820ttcttcctct acagcaagct caccgtggac aagagcaggt
ggcagcaggg gaacgtcttc 2880tcatgctccg tgatgcatga ggccctgcac
aatcactata cccagaaatc tctgagtctg 2940agcccaggca agaaggaccc
caaggcactg atagcatttc tggcatttct gattattgtg 3000acatcaatag
ccctgcttgt tgttctctac aaaatctatg atctacataa gaaaagatcc
3060tgcaatttag atgaacagca ggagcttgtt gaaagggatg atgaaaaaca
actgatgaat 3120gtggagccaa tccatgcaga tattttgttg gaaacttata
agaggaagat tgctgatgaa 3180ggaagacttt ttctggctga atttcagagc
atcccgcggg tgttcagcaa gtttcctata 3240aaggaagctc gaaagccctt
taaccagaat aaaaaccgtt atgttgacat tcttccttat 3300gattataacc
gtgttgaact ctctgagata aacggagatg cagggtcaaa ctacataaat
3360gccagctata ttgatggttt caaagaaccc aggaaataca ttgctgcaca
aggtcccagg 3420gatgaaactg ttgatgattt ctggaggatg atttgggaac
agaaagccac agttattgtc 3480atggtcactc gatgtgaaga aggaaacagg
aacaagtgtg cagaatactg gccgtcaatg 3540gaagagggca ctcgggcttt
tggagatgtt gttgtaaaga tcaaccagca caaaagatgt 3600ccagattaca
tcattcagaa attgaacatt gtaaataaaa aagaaaaagc aactggaaga
3660gaggtgactc acattcagtt caccagctgg ccagaccacg gggtgcctga
ggatcctcac 3720ttgctcctca aactgagaag gagagtgaat gccttcagca
atttcttcag tggtcccatt 3780gtggtgcact gcagtgctgg tgttgggcgc
acaggaacct atatcggaat tgatgccatg 3840ctagaaggcc tggaagccga
gaacaaagtg gatgtttatg gttatgttgt caagctaagg 3900cgacagagat
gcctgatggt tcaagtagag gcccagtaca tcttgatcca tcaggctttg
3960gtggaataca atcagtttgg agaaacagaa gtgaatttgt ctgaattaca
tccatatcta 4020cataacatga agaaaaggga tccacccagt gagccgtctc
cactagaggc tgaattccag 4080agacttcctt catataggag ctggaggaca
cagcacattg gaaatcaaga agaaaataaa 4140agtaaaaaca ggaattctaa
tgtcatccca tatgactata acagagtgcc acttaaacat 4200gagctggaaa
tgagtaaaga gagtgagcat gattcagatg aatcctctga tgatgacagt
4260gattcagagg aaccaagcaa atacatcaat gcatctttta taatgagcta
ctggaaacct 4320gaagtgatga ttgctgctca gggaccactg aaggagacca
ttggtgactt ttggcagatg 4380atcttccaaa gaaaagtcaa agttattgtt
atgctgacag aactgaaaca tggagaccag 4440gaaatctgtg ctcagtactg
gggagaagga aagcaaacat atggagatat tgaagttgac 4500ctgaaagaca
cagacaaatc ttcaacttat acccttcgtg tctttgaact gagacattcc
4560aagaggaaag actctcgaac tgtgtaccag taccaatata caaactggag
tgtggagcag 4620cttcctgcag aacccaagga attaatctct atgattcagg
tcgtcaaaca aaaacttccc 4680cagaagaatt cctctgaagg gaacaagcat
cacaagagta cacctctact cattcactgc 4740agggatggat ctcagcaaac
gggaatattt tgtgctttgt taaatctctt agaaagtgcg 4800gaaacagaag
aggtagtgga tatttttcaa gtggtaaaag ctctacgcaa agctaggcca
4860ggcatggttt ccacattcga gcaatatcaa ttcctatatg acgtcattgc
cagcacctac 4920cctgctcaga atggacaagt aaagaaaaac aaccatcaag
aagataaaat tgaatttgat 4980aatgaagtgg acaaagtaaa gcaggatgct
aattgtgtta atccacttgg tgccccagaa 5040aagctccctg aagcaaagga
acaggctgaa ggttctgaac ccacgagtgg cactgagggg 5100ccagaacatt
ctgtcaatgg tcctgcaagt ccagctttaa atcaaggttc atag
5154374053DNAArtificial SequenceNucleic acid sequences coding for
CARs
(MP14801.SFG.aCD19fmc63_clean-CD8STK-CD28tmZ-2A-aCD33glx-HCH2CH3p
vaa-dCD148) 37atgagcctgc ccgtgaccgc cctgctgctg cccctggccc
tgctgctgca cgccgccaga 60ccagacatcc agatgaccca gaccaccagc agcctgagcg
ccagcctggg cgaccgggtg 120accatcagct gcagagccag ccaggacatc
agcaagtacc tgaactggta ccagcagaag 180cccgacggca ccgtgaagct
gctgatctac cacaccagcc ggctgcacag cggcgtgccc 240agccggttca
gcggcagcgg cagcggcacc gactacagcc tgaccatcag caacctggag
300caggaggaca tcgccaccta cttctgccag cagggcaaca ccctgcccta
caccttcgga 360ggcggcacca agctggagat caccaaggcc ggaggcggag
gctctggcgg aggcggctct 420ggcggaggcg gctctggcgg aggcggcagc
gaggtgaagc tgcaggagtc tggcccaggc 480ctggtggccc caagccagag
cctgagcgtg acctgcaccg tgagcggcgt gagcctgccc 540gactacggcg
tgagctggat caggcagccc ccacggaagg gcctggagtg gctgggcgtg
600atctggggca gcgagaccac ctactacaac agcgccctga agagccggct
gaccatcatc 660aaggacaaca gcaagagcca ggtgttcctg aagatgaaca
gcctgcagac cgacgacacc 720gccatctact actgcgccaa gcactactac
tatggcggca gctacgctat ggactactgg 780ggccagggca ccagcgtgac
cgtgagctca gatcccacca cgacgccagc gccgcgacca 840ccaacaccgg
cgcccaccat cgcgtcgcag cccctgtccc tgcgcccaga ggcgtgccgg
900ccagcggcgg ggggcgcagt gcacacgagg gggctggact tcgcctgtga
tatcttttgg 960gtgctggtgg tggttggtgg agtcctggct tgctatagct
tgctagtaac agtggccttt 1020attattttct gggtgaggag agtgaagttc
agcaggagcg cagacgcccc cgcgtaccag 1080cagggccaga accagctcta
taacgagctc aatctaggac gaagagagga gtacgatgtt 1140ttggacaaga
gacgtggccg ggaccctgag atggggggaa agccgagaag gaagaaccct
1200caggaaggcc tgtacaatga actgcagaaa gataagatgg cggaggccta
cagtgagatt 1260gggatgaaag gcgagcgccg gaggggcaag gggcacgatg
gcctttacca gggtctcagt 1320acagccacca aggacaccta cgacgccctt
cacatgcagg ccctgcctcc tcgcagagcc 1380gagggcaggg gaagtcttct
aacatgcggg gacgtggagg aaaatcccgg gcccatggcc 1440gtgcccactc
aggtcctggg gttgttgcta ctgtggctta cagatgccag atgtgacatc
1500cagatgacac agtctccatc ttccctgtct gcatctgtcg gagatcgcgt
caccatcacc 1560tgtcgagcaa gtgaggacat ttattttaat ttagtgtggt
atcagcagaa accaggaaag 1620gcccctaagc tcctgatcta tgatacaaat
cgcttggcag atggggtccc atcacggttc 1680agtggctctg gatctggcac
acagtatact ctaaccataa gtagcctgca acccgaagat 1740ttcgcaacct
attattgtca acactataag aattatccgc tcacgttcgg tcaggggacc
1800aagctggaaa tcaaaagatc tggtggcgga gggtcaggag gcggaggcag
cggaggcggt 1860ggctcgggag gcggaggctc gagatctgag gtgcagttgg
tggagtctgg gggcggcttg 1920gtgcagcctg gagggtccct gaggctctcc
tgtgcagcct caggattcac tctcagtaat 1980tatggcatgc actggatcag
gcaggctcca gggaagggtc tggagtgggt ctcgtctatt 2040agtcttaatg
gtggtagcac ttactatcga gactccgtga agggccgatt cactatctcc
2100agggacaatg caaaaagcac cctctacctt caaatgaata gtctgagggc
cgaggacacg 2160gccgtctatt actgtgcagc acaggacgct tatacgggag
gttactttga ttactggggc 2220caaggaacgc tggtcacagt ctcgtctatg
gatcccgccg agcccaaatc tcctgacaaa 2280actcacacat gcccaccgtg
cccagcacct cccgtggccg gcccgtcagt cttcctcttc 2340cccccaaaac
ccaaggacac cctcatgatc gcccggaccc ctgaggtcac atgcgtggtg
2400gtggacgtga gccacgaaga ccctgaggtc aagttcaact ggtacgtgga
cggcgtggag 2460gtgcataatg ccaagacaaa gccgcgggag gagcagtaca
acagcacgta ccgtgtggtc 2520agcgtcctca ccgtcctgca ccaggactgg
ctgaatggca aggagtacaa gtgcaaggtc 2580tccaacaaag ccctcccagc
ccccatcgag aaaaccatct ccaaagccaa agggcagccc 2640cgagaaccac
aggtgtacac cctgccccca tcccgggatg agctgaccaa gaaccaggtc
2700agcctgacct gcctggtcaa aggcttctat cccagcgaca tcgccgtgga
gtgggagagc 2760aatgggcaac cggagaacaa ctacaagacc acgcctcccg
tgctggactc cgacggctcc 2820ttcttcctct acagcaagct caccgtggac
aagagcaggt ggcagcaggg gaacgtcttc 2880tcatgctccg tgatgcatga
ggccctgcac aatcactata cccagaaatc tctgagtctg 2940agcccaggca
agaaggaccc caaggcggtt tttggctgta tctttggtgc cctggttatt
3000gtgactgtgg gaggcttcat cttctggaga aagaagagga aagatgcaaa
gaataatgaa 3060gtgtcctttt ctcaaattaa acctaaaaaa tctaagttaa
tcagagtgga gaattttgag 3120gcctacttca agaagcagca agctgactcc
aactgtgggt tcgcagagga atacgaagat 3180ctgaagcttg ttggaattag
tcaacctaaa tatgcagcag aactggctga gaatagagga 3240aagaatcgct
ataataatgt tctgccctat gatatttccc gtgtcaaact ttcggtccag
3300acccattcaa cggatgacta catcaatgcc aactacatgc ctggctacca
ctccaagaaa 3360gattttattg ccacacaagg acctttaccg aacactttga
aagatttttg gcgtatggtt 3420tgggagaaaa atgtatatgc catcattatg
ttgactaaat gtgttgaaca gggaagaacc 3480aaatgtgagg agtattggcc
ctccaagcag gctcaggact atggagacat aactgtggca 3540atgacatcag
aaattgttct tccggaatgg accatcagag atttcacagt gaaaaatatc
3600cagacaagtg agagtcaccc tctgagacag ttccatttca cctcctggcc
agaccacggt 3660gttcccgaca ccactgacct gctcatcaac ttccggtacc
tcgttcgtga ctacatgaag 3720cagagtcctc ccgaatcgcc gattctggtg
cattgcagtg ctggggtcgg aaggacgggc 3780actttcattg ccattgatcg
tctcatctac cagatagaga atgagaacac cgtggatgtg 3840tatgggattg
tgtatgacct tcgaatgcat aggcctttaa tggtgcagac agaggaccag
3900tatgttttcc tcaatcagtg tgttttggat attgtcagat cccagaaaga
ctcaaaagta 3960gatcttatct accagaacac aactgcaatg acaatctatg
aaaaccttgc gcccgtgacc 4020acatttggaa agaccaatgg ttacatcgcc taa
4053383345DNAArtificial SequenceNucleic acid sequences coding for
CARs
(16076.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-tm-dPTP
N6) 38atgagcctgc ccgtgaccgc cctgctgctg cccctggccc tgctgctgca
cgccgccaga 60ccagacatcc agatgaccca gaccaccagc agcctgagcg ccagcctggg
cgaccgggtg 120accatcagct gcagagccag ccaggacatc agcaagtacc
tgaactggta ccagcagaag 180cccgacggca ccgtgaagct gctgatctac
cacaccagcc ggctgcacag cggcgtgccc 240agccggttca gcggcagcgg
cagcggcacc gactacagcc tgaccatcag caacctggag 300caggaggaca
tcgccaccta cttctgccag cagggcaaca ccctgcccta caccttcgga
360ggcggcacca agctggagat caccaaggcc ggaggcggag gctctggcgg
aggcggctct 420ggcggaggcg gctctggcgg aggcggcagc gaggtgaagc
tgcaggagtc tggcccaggc 480ctggtggccc caagccagag cctgagcgtg
acctgcaccg tgagcggcgt gagcctgccc 540gactacggcg tgagctggat
caggcagccc ccacggaagg gcctggagtg gctgggcgtg 600atctggggca
gcgagaccac ctactacaac agcgccctga agagccggct gaccatcatc
660aaggacaaca gcaagagcca ggtgttcctg aagatgaaca gcctgcagac
cgacgacacc 720gccatctact actgcgccaa gcactactac tatggcggca
gctacgctat ggactactgg 780ggccagggca ccagcgtgac cgtgagctca
gatcccacca cgacgccagc gccgcgacca 840ccaacaccgg cgcccaccat
cgcgtcgcag cccctgtccc tgcgcccaga ggcgtgccgg 900ccagcggcgg
ggggcgcagt gcacacgagg gggctggact tcgcctgtga tatcttttgg
960gtgctggtgg tggttggtgg agtcctggct tgctatagct tgctagtaac
agtggccttt 1020attattttct gggtgaggag agtgaagttc agcaggagcg
cagacgcccc cgcgtaccag 1080cagggccaga accagctcta taacgagctc
aatctaggac gaagagagga gtacgatgtt 1140ttggacaaga gacgtggccg
ggaccctgag atggggggaa agccgagaag gaagaaccct 1200caggaaggcc
tgtacaatga actgcagaaa gataagatgg cggaggccta cagtgagatt
1260gggatgaaag gcgagcgccg gaggggcaag gggcacgatg gcctttacca
gggtctcagt 1320acagccacca aggacaccta cgacgccctt cacatgcagg
ccctgcctcc tcgcagagcc 1380gagggcaggg gaagtcttct aacatgcggg
gacgtggagg aaaatcccgg gcccatggcc 1440gtgcccactc aggtcctggg
gttgttgcta ctgtggctta cagatgccag atgtgacatc 1500cagatgacac
agtctccatc ttccctgtct gcatctgtcg gagatcgcgt caccatcacc
1560tgtcgagcaa gtgaggacat ttattttaat ttagtgtggt atcagcagaa
accaggaaag 1620gcccctaagc tcctgatcta tgatacaaat cgcttggcag
atggggtccc atcacggttc 1680agtggctctg gatctggcac acagtatact
ctaaccataa gtagcctgca acccgaagat 1740ttcgcaacct attattgtca
acactataag aattatccgc tcacgttcgg tcaggggacc 1800aagctggaaa
tcaaaagatc tggtggcgga gggtcaggag gcggaggcag cggaggcggt
1860ggctcgggag gcggaggctc gagatctgag gtgcagttgg tggagtctgg
gggcggcttg 1920gtgcagcctg gagggtccct gaggctctcc tgtgcagcct
caggattcac tctcagtaat 1980tatggcatgc actggatcag gcaggctcca
gggaagggtc tggagtgggt ctcgtctatt 2040agtcttaatg gtggtagcac
ttactatcga gactccgtga agggccgatt cactatctcc 2100agggacaatg
caaaaagcac cctctacctt caaatgaata gtctgagggc cgaggacacg
2160gccgtctatt actgtgcagc acaggacgct tatacgggag gttactttga
ttactggggc 2220caaggaacgc tggtcacagt ctcgtctatg gatcccgcca
ccacaaccaa gcccgtgctg 2280cggaccccaa gccctgtgca ccctaccggc
accagccagc ctcagagacc cgaggactgc 2340cggcctcggg gcagcgtgaa
gggcaccggc ctggacttcg cctgcgacat ctactgggca 2400cctctggccg
gaatatgcgt ggcactgctg ctgagcctca tcatcaccct gatctgttat
2460caccgaagcc gcaagcgggt gtgtaaaagt ggaggcggaa gcttctggga
ggagtttgag 2520agtttgcaga agcaggaggt gaagaacttg caccagcgtc
tggaagggca gcggccagag 2580aacaagggca agaaccgcta caagaacatt
ctcccctttg accacagccg agtgatcctg 2640cagggacggg acagtaacat
ccccgggtcc gactacatca atgccaacta catcaagaac 2700cagctgctag
gccctgatga gaacgctaag acctacatcg ccagccaggg ctgtctggag
2760gccacggtca atgacttctg gcagatggcg tggcaggaga acagccgtgt
catcgtcatg 2820accacccgag aggtggagaa aggccggaac aaatgcgtcc
catactggcc cgaggtgggc 2880atgcagcgtg cttatgggcc ctactctgtg
accaactgcg gggagcatga cacaaccgaa 2940tacaaactcc gtaccttaca
ggtctccccg ctggacaatg gagacctgat tcgggagatc 3000tggcattacc
agtacctgag ctggcccgac cacggggtcc ccagtgagcc tgggggtgtc
3060ctcagcttcc tggaccagat caaccagcgg caggaaagtc tgcctcacgc
agggcccatc 3120atcgtgcact gcagcgccgg catcggccgc acaggcacca
tcattgtcat cgacatgctc 3180atggagaaca tctccaccaa gggcctggac
tgtgacattg acatccagaa gaccatccag 3240atggtgcggg cgcagcgctc
gggcatggtg cagacggagg cgcagtacaa gttcatctac 3300gtggccatcg
cccagttcat tgaaaccact aagaagaagc tgtga 3345392757DNAArtificial
SequenceNucleic acid sequences coding for CARs
(MP16091.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-LAIR1
tm-endo) 39atgagcctgc ccgtgaccgc cctgctgctg cccctggccc tgctgctgca
cgccgccaga 60ccagacatcc agatgaccca gaccaccagc agcctgagcg ccagcctggg
cgaccgggtg 120accatcagct gcagagccag ccaggacatc agcaagtacc
tgaactggta ccagcagaag 180cccgacggca ccgtgaagct gctgatctac
cacaccagcc ggctgcacag cggcgtgccc 240agccggttca gcggcagcgg
cagcggcacc gactacagcc tgaccatcag caacctggag 300caggaggaca
tcgccaccta cttctgccag cagggcaaca ccctgcccta caccttcgga
360ggcggcacca agctggagat caccaaggcc ggaggcggag gctctggcgg
aggcggctct 420ggcggaggcg gctctggcgg aggcggcagc gaggtgaagc
tgcaggagtc tggcccaggc 480ctggtggccc caagccagag cctgagcgtg
acctgcaccg tgagcggcgt gagcctgccc 540gactacggcg tgagctggat
caggcagccc ccacggaagg gcctggagtg gctgggcgtg 600atctggggca
gcgagaccac ctactacaac agcgccctga agagccggct gaccatcatc
660aaggacaaca gcaagagcca ggtgttcctg aagatgaaca gcctgcagac
cgacgacacc 720gccatctact actgcgccaa gcactactac tatggcggca
gctacgctat ggactactgg 780ggccagggca ccagcgtgac cgtgagctca
gatcccacca cgacgccagc gccgcgacca 840ccaacaccgg cgcccaccat
cgcgtcgcag cccctgtccc tgcgcccaga ggcgtgccgg 900ccagcggcgg
ggggcgcagt gcacacgagg gggctggact tcgcctgtga tatcttttgg
960gtgctggtgg tggttggtgg agtcctggct tgctatagct tgctagtaac
agtggccttt 1020attattttct gggtgaggag agtgaagttc agcaggagcg
cagacgcccc cgcgtaccag 1080cagggccaga accagctcta taacgagctc
aatctaggac gaagagagga gtacgatgtt 1140ttggacaaga gacgtggccg
ggaccctgag atggggggaa agccgagaag gaagaaccct 1200caggaaggcc
tgtacaatga actgcagaaa gataagatgg cggaggccta cagtgagatt
1260gggatgaaag gcgagcgccg gaggggcaag gggcacgatg gcctttacca
gggtctcagt 1320acagccacca aggacaccta cgacgccctt cacatgcagg
ccctgcctcc tcgcagagcc 1380gagggcaggg gaagtcttct aacatgcggg
gacgtggagg aaaatcccgg gcccatggcc 1440gtgcccactc aggtcctggg
gttgttgcta ctgtggctta cagatgccag atgtgacatc 1500cagatgacac
agtctccatc ttccctgtct gcatctgtcg gagatcgcgt caccatcacc
1560tgtcgagcaa gtgaggacat ttattttaat ttagtgtggt atcagcagaa
accaggaaag 1620gcccctaagc tcctgatcta tgatacaaat cgcttggcag
atggggtccc atcacggttc 1680agtggctctg gatctggcac acagtatact
ctaaccataa gtagcctgca acccgaagat 1740ttcgcaacct attattgtca
acactataag aattatccgc tcacgttcgg tcaggggacc 1800aagctggaaa
tcaaaagatc tggtggcgga gggtcaggag gcggaggcag cggaggcggt
1860ggctcgggag gcggaggctc gagatctgag gtgcagttgg tggagtctgg
gggcggcttg 1920gtgcagcctg gagggtccct gaggctctcc tgtgcagcct
caggattcac tctcagtaat 1980tatggcatgc actggatcag gcaggctcca
gggaagggtc tggagtgggt ctcgtctatt 2040agtcttaatg gtggtagcac
ttactatcga gactccgtga agggccgatt cactatctcc 2100agggacaatg
caaaaagcac cctctacctt caaatgaata gtctgagggc cgaggacacg
2160gccgtctatt actgtgcagc acaggacgct tatacgggag gttactttga
ttactggggc 2220caaggaacgc tggtcacagt ctcgtctatg gatcccgcca
ccacaaccaa gcccgtgctg 2280cggaccccaa gccctgtgca ccctaccggc
accagccagc ctcagagacc cgaggactgc 2340cggcctcggg gcagcgtgaa
gggcaccggc ctggacttcg cctgcgacat tctcatcggg 2400gtctcagtgg
tcttcctctt ctgtctcctc ctcctggtcc tcttctgcct ccatcgccag
2460aatcagataa agcaggggcc ccccagaagc aaggacgagg agcagaagcc
acagcagagg 2520cctgacctgg ctgttgatgt tctagagagg acagcagaca
aggccacagt caatggactt 2580cctgagaagg accgggagac cgacaccagc
gccctggctg cagggagttc ccaggaggtg 2640acgtatgctc agctggacca
ctgggccctc acacagagga cagcccgggc tgtgtcccca 2700cagtccacaa
agcccatggc cgagtccatc acgtatgcag ccgttgccag acactga
2757404092DNAArtificial SequenceNucleic acid sequences coding for
CARs
(MP16092.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-LAIR1
tm-endo-2A-PTPN6_SH2-dCD148) 40atgagcctgc ccgtgaccgc cctgctgctg
cccctggccc tgctgctgca cgccgccaga 60ccagacatcc agatgaccca gaccaccagc
agcctgagcg ccagcctggg cgaccgggtg 120accatcagct gcagagccag
ccaggacatc agcaagtacc tgaactggta ccagcagaag 180cccgacggca
ccgtgaagct gctgatctac cacaccagcc ggctgcacag cggcgtgccc
240agccggttca gcggcagcgg cagcggcacc gactacagcc tgaccatcag
caacctggag 300caggaggaca tcgccaccta cttctgccag cagggcaaca
ccctgcccta caccttcgga 360ggcggcacca agctggagat caccaaggcc
ggaggcggag gctctggcgg aggcggctct 420ggcggaggcg gctctggcgg
aggcggcagc gaggtgaagc tgcaggagtc tggcccaggc 480ctggtggccc
caagccagag cctgagcgtg acctgcaccg tgagcggcgt gagcctgccc
540gactacggcg tgagctggat caggcagccc ccacggaagg gcctggagtg
gctgggcgtg 600atctggggca gcgagaccac ctactacaac agcgccctga
agagccggct gaccatcatc 660aaggacaaca gcaagagcca ggtgttcctg
aagatgaaca gcctgcagac cgacgacacc 720gccatctact actgcgccaa
gcactactac tatggcggca gctacgctat ggactactgg 780ggccagggca
ccagcgtgac cgtgagctca gatcccacca cgacgccagc gccgcgacca
840ccaacaccgg cgcccaccat cgcgtcgcag cccctgtccc tgcgcccaga
ggcgtgccgg 900ccagcggcgg ggggcgcagt gcacacgagg gggctggact
tcgcctgtga tatcttttgg 960gtgctggtgg tggttggtgg agtcctggct
tgctatagct tgctagtaac agtggccttt 1020attattttct gggtgaggag
agtgaagttc agcaggagcg cagacgcccc cgcgtaccag 1080cagggccaga
accagctcta taacgagctc aatctaggac gaagagagga gtacgatgtt
1140ttggacaaga gacgtggccg ggaccctgag atggggggaa agccgagaag
gaagaaccct 1200caggaaggcc tgtacaatga actgcagaaa gataagatgg
cggaggccta cagtgagatt 1260gggatgaaag gcgagcgccg gaggggcaag
gggcacgatg gcctttacca gggtctcagt 1320acagccacca aggacaccta
cgacgccctt cacatgcagg ccctgcctcc tcgcagagcc 1380gagggcaggg
gaagtcttct aacatgcggg gacgtggagg aaaatcccgg gcccatggcc
1440gtgcccactc aggtcctggg gttgttgcta ctgtggctta cagatgccag
atgtgacatc 1500cagatgacac agtctccatc ttccctgtct gcatctgtcg
gagatcgcgt caccatcacc 1560tgtcgagcaa gtgaggacat ttattttaat
ttagtgtggt atcagcagaa accaggaaag 1620gcccctaagc tcctgatcta
tgatacaaat cgcttggcag atggggtccc atcacggttc 1680agtggctctg
gatctggcac acagtatact ctaaccataa gtagcctgca acccgaagat
1740ttcgcaacct attattgtca acactataag aattatccgc tcacgttcgg
tcaggggacc 1800aagctggaaa tcaaaagatc tggtggcgga gggtcaggag
gcggaggcag cggaggcggt 1860ggctcgggag gcggaggctc gagatctgag
gtgcagttgg tggagtctgg gggcggcttg 1920gtgcagcctg gagggtccct
gaggctctcc tgtgcagcct caggattcac tctcagtaat 1980tatggcatgc
actggatcag gcaggctcca gggaagggtc tggagtgggt ctcgtctatt
2040agtcttaatg gtggtagcac ttactatcga gactccgtga agggccgatt
cactatctcc 2100agggacaatg caaaaagcac cctctacctt caaatgaata
gtctgagggc cgaggacacg 2160gccgtctatt actgtgcagc acaggacgct
tatacgggag gttactttga ttactggggc 2220caaggaacgc tggtcacagt
ctcgtctatg gatcccgcca ccacaaccaa gcccgtgctg 2280cggaccccaa
gccctgtgca ccctaccggc accagccagc ctcagagacc cgaggactgc
2340cggcctcggg gcagcgtgaa gggcaccggc ctggacttcg cctgcgacat
tctcatcggg 2400gtctcagtgg tcttcctctt ctgtctcctc ctcctggtcc
tcttctgcct ccatcgccag 2460aatcagataa agcaggggcc ccccagaagc
aaggacgagg agcagaagcc acagcagagg 2520cctgacctgg ctgttgatgt
tctagagagg acagcagaca aggccacagt caatggactt 2580cctgagaagg
accgggagac cgacaccagc gccctggctg cagggagttc ccaggaggtg
2640acgtatgctc agctggacca ctgggccctc acacagagga cagcccgggc
tgtgtcccca 2700cagtccacaa agcccatggc cgagtccatc acgtatgcag
ccgttgccag acacagggca 2760gaaggaagag gtagcctgct gacttgcggg
gacgtggaag agaacccagg gccatggtat 2820catggccaca tgtctggcgg
gcaggcagag acgctgctgc aggccaaggg cgagccctgg 2880acgtttcttg
tgcgtgagag cctcagccag cctggagact tcgtgctttc tgtgctcagt
2940gaccagccca aggctggccc aggctccccg ctcagggtca cccacatcaa
ggtcatgtgc 3000gagggtggac gctacacagt gggtggtttg gagaccttcg
acagcctcac ggacctggtg 3060gagcatttca agaagacggg gattgaggag
gcctcaggcg cctttgtcta cctgcggcag 3120ccgtacagcg gtggcggtgg
cagctttgag gcctacttca agaagcagca agctgactcc 3180aactgtgggt
tcgcagagga atacgaagat ctgaagcttg ttggaattag tcaacctaaa
3240tatgcagcag aactggctga gaatagagga aagaatcgct ataataatgt
tctgccctat 3300gatatttccc gtgtcaaact ttcggtccag acccattcaa
cggatgacta catcaatgcc 3360aactacatgc ctggctacca ctccaagaaa
gattttattg ccacacaagg acctttaccg 3420aacactttga aagatttttg
gcgtatggtt tgggagaaaa atgtatatgc catcattatg 3480ttgactaaat
gtgttgaaca gggaagaacc aaatgtgagg agtattggcc ctccaagcag
3540gctcaggact atggagacat aactgtggca atgacatcag aaattgttct
tccggaatgg 3600accatcagag atttcacagt gaaaaatatc cagacaagtg
agagtcaccc tctgagacag 3660ttccatttca cctcctggcc agaccacggt
gttcccgaca ccactgacct gctcatcaac 3720ttccggtacc tcgttcgtga
ctacatgaag cagagtcctc ccgaatcgcc gattctggtg 3780cattgcagtg
ctggggtcgg aaggacgggc actttcattg ccattgatcg tctcatctac
3840cagatagaga atgagaacac cgtggatgtg tatgggattg tgtatgacct
tcgaatgcat 3900aggcctttaa tggtgcagac agaggaccag tatgttttcc
tcaatcagtg tgttttggat 3960attgtcagat cccagaaaga ctcaaaagta
gatcttatct accagaacac aactgcaatg 4020acaatctatg aaaaccttgc
gcccgtgacc acatttggaa agaccaatgg ttacatcgcc 4080agcggtagct aa
4092411341PRTArtificial SequenceSingle-chain variable fragment
(scFv) SFG.aCD19-CD8STK-CD28tmZ-2A-aGD2-HCH2CH3pvaa-dCD148 41Met
Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1 5 10
15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser Leu
20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg Ala
Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln Lys
Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His Thr Ser Arg
Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly Ser
Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln Glu
Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu Pro
Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys Ala
Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135 140
Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly 145
150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr Val
Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile Arg
Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile Trp
Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser Arg
Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe Leu
Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile Tyr
Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255 Met
Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260 265
270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala
275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala
Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala Cys
Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly Gly Val Leu
Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe Ile Ile Phe
Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala Asp Ala Pro
Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365 Glu Leu Asn
Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370 375 380 Arg
Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro 385 390
395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu
Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly
Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr
Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala Leu Pro Pro
Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr Cys Gly Asp
Val Glu Glu Asn Pro Gly Pro Met Glu 465 470 475 480 Thr Asp Thr Leu
Leu Leu Trp Val Leu Leu Leu Trp Val Pro Gly Ser 485 490 495 Thr Gly
Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro 500 505 510
Ser Gln Thr Leu Ser Ile Thr Cys Thr Val Ser Gly Phe Ser Leu Ala 515
520 525 Ser Tyr Asn Ile His Trp Val Arg Gln Pro Pro Gly Lys Gly Leu
Glu 530 535 540 Trp Leu Gly Val Ile Trp Ala Gly Gly Ser Thr Asn Tyr
Asn Ser Ala 545 550 555 560 Leu Met Ser Arg Leu Thr Ile Ser Lys Asp
Asn Ser Lys Asn Gln Val 565 570 575 Phe Leu Lys Met Ser Ser Leu Thr
Ala Ala Asp Thr Ala Val Tyr Tyr 580 585 590 Cys Ala Lys Arg Ser Asp
Asp Tyr Ser Trp Phe Ala Tyr Trp Gly Gln 595 600 605 Gly Thr Leu Val
Thr Val Ser Ser Gly Gly Gly Gly Ser Gly Gly Gly 610 615 620 Gly Ser
Gly Gly Gly Gly Ser Glu Asn Gln Met Thr Gln Ser Pro Ser 625 630 635
640 Ser Leu Ser Ala Ser Val Gly Asp Arg Val Thr Met Thr Cys Arg Ala
645 650 655 Ser Ser Ser Val Ser Ser Ser Tyr Leu His Trp Tyr Gln Gln
Lys Ser 660 665 670 Gly Lys Ala Pro Lys Val Trp Ile Tyr Ser Thr Ser
Asn Leu Ala Ser 675 680 685 Gly Val Pro Ser Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Tyr Thr 690 695 700 Leu Thr Ile Ser Ser Leu Gln Pro
Glu Asp Phe Ala Thr Tyr Tyr Cys 705 710 715 720 Gln Gln Tyr Ser Gly
Tyr Pro Ile Thr Phe Gly Gln Gly Thr Lys Val 725 730 735 Glu Ile Lys
Arg Ser Asp Pro Ala Glu Pro Lys Ser Pro Asp Lys Thr 740 745 750 His
Thr Cys Pro Pro Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val 755 760
765 Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ala Arg Thr
770 775 780 Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp
Pro Glu 785 790 795 800 Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His Asn Ala Lys 805 810 815 Thr Lys Pro Arg Glu Glu Gln Tyr Asn
Ser Thr Tyr Arg Val Val Ser 820 825 830 Val Leu Thr Val Leu His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys 835 840 845 Cys Lys Val Ser Asn
Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile 850 855 860 Ser Lys Ala
Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro 865 870 875 880
Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu 885
890 895 Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Asn 900 905 910 Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp Ser 915 920 925 Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr
Val Asp Lys Ser Arg 930 935 940 Trp Gln Gln Gly Asn Val Phe Ser Cys
Ser Val Met His Glu Ala Leu 945 950 955 960 His Asn His Tyr Thr Gln
Lys Ser Leu Ser Leu Ser Pro Gly Lys Lys 965 970 975 Asp Pro Lys Ala
Val Phe Gly Cys Ile Phe Gly Ala Leu Val Ile Val 980 985 990 Thr Val
Gly Gly Phe Ile Phe Trp Arg Lys Lys Arg Lys Asp Ala Lys 995 1000
1005 Asn Asn Glu Val Ser Phe Ser Gln Ile Lys Pro Lys Lys Ser Lys
1010 1015 1020 Leu Ile Arg Val Glu Asn Phe Glu Ala Tyr Phe Lys Lys
Gln Gln 1025 1030 1035 Ala Asp Ser Asn Cys Gly Phe Ala Glu Glu Tyr
Glu Asp Leu Lys 1040 1045 1050 Leu Val Gly Ile Ser Gln Pro Lys Tyr
Ala Ala Glu Leu Ala Glu 1055 1060 1065 Asn Arg Gly Lys Asn Arg Tyr
Asn Asn Val Leu Pro Tyr Asp Ile 1070 1075 1080
Ser Arg Val Lys Leu Ser Val Gln Thr His Ser Thr Asp Asp Tyr 1085
1090 1095 Ile Asn Ala Asn Tyr Met Pro Gly Tyr His Ser Lys Lys Asp
Phe 1100 1105 1110 Ile Ala Thr Gln Gly Pro Leu Pro Asn Thr Leu Lys
Asp Phe Trp 1115 1120 1125 Arg Met Val Trp Glu Lys Asn Val Tyr Ala
Ile Ile Met Leu Thr 1130 1135 1140 Lys Cys Val Glu Gln Gly Arg Thr
Lys Cys Glu Glu Tyr Trp Pro 1145 1150 1155 Ser Lys Gln Ala Gln Asp
Tyr Gly Asp Ile Thr Val Ala Met Thr 1160 1165 1170 Ser Glu Ile Val
Leu Pro Glu Trp Thr Ile Arg Asp Phe Thr Val 1175 1180 1185 Lys Asn
Ile Gln Thr Ser Glu Ser His Pro Leu Arg Gln Phe His 1190 1195 1200
Phe Thr Ser Trp Pro Asp His Gly Val Pro Asp Thr Thr Asp Leu 1205
1210 1215 Leu Ile Asn Phe Arg Tyr Leu Val Arg Asp Tyr Met Lys Gln
Ser 1220 1225 1230 Pro Pro Glu Ser Pro Ile Leu Val His Cys Ser Ala
Gly Val Gly 1235 1240 1245 Arg Thr Gly Thr Phe Ile Ala Ile Asp Arg
Leu Ile Tyr Gln Ile 1250 1255 1260 Glu Asn Glu Asn Thr Val Asp Val
Tyr Gly Ile Val Tyr Asp Leu 1265 1270 1275 Arg Met His Arg Pro Leu
Met Val Gln Thr Glu Asp Gln Tyr Val 1280 1285 1290 Phe Leu Asn Gln
Cys Val Leu Asp Ile Val Arg Ser Gln Lys Asp 1295 1300 1305 Ser Lys
Val Asp Leu Ile Tyr Gln Asn Thr Thr Ala Met Thr Ile 1310 1315 1320
Tyr Glu Asn Leu Ala Pro Val Thr Thr Phe Gly Lys Thr Asn Gly 1325
1330 1335 Tyr Ile Ala 1340 424026DNAArtificial SequenceSingle-chain
variable fragment (scFv)
SFG.aCD19-CD8STK-CD28tmZ-2A-aGD2-HCH2CH3pvaa-dCD148 42atgagcctgc
ccgtgaccgc cctgctgctg cccctggccc tgctgctgca cgccgccaga 60ccagacatcc
agatgaccca gaccaccagc agcctgagcg ccagcctggg cgaccgggtg
120accatcagct gcagagccag ccaggacatc agcaagtacc tgaactggta
ccagcagaag 180cccgacggca ccgtgaagct gctgatctac cacaccagcc
ggctgcacag cggcgtgccc 240agccggttca gcggcagcgg cagcggcacc
gactacagcc tgaccatcag caacctggag 300caggaggaca tcgccaccta
cttctgccag cagggcaaca ccctgcccta caccttcgga 360ggcggcacca
agctggagat caccaaggcc ggaggcggag gctctggcgg aggcggctct
420ggcggaggcg gctctggcgg aggcggcagc gaggtgaagc tgcaggagtc
tggcccaggc 480ctggtggccc caagccagag cctgagcgtg acctgcaccg
tgagcggcgt gagcctgccc 540gactacggcg tgagctggat caggcagccc
ccacggaagg gcctggagtg gctgggcgtg 600atctggggca gcgagaccac
ctactacaac agcgccctga agagccggct gaccatcatc 660aaggacaaca
gcaagagcca ggtgttcctg aagatgaaca gcctgcagac cgacgacacc
720gccatctact actgcgccaa gcactactac tatggcggca gctacgctat
ggactactgg 780ggccagggca ccagcgtgac cgtgagctca gatcccacca
cgacgccagc gccgcgacca 840ccaacaccgg cgcccaccat cgcgtcgcag
cccctgtccc tgcgcccaga ggcgtgccgg 900ccagcggcgg ggggcgcagt
gcacacgagg gggctggact tcgcctgtga tatcttttgg 960gtgctggtgg
tggttggtgg agtcctggct tgctatagct tgctagtaac agtggccttt
1020attattttct gggtgaggag agtgaagttc agcaggagcg cagacgcccc
cgcgtaccag 1080cagggccaga accagctcta taacgagctc aatctaggac
gaagagagga gtacgatgtt 1140ttggacaaga gacgtggccg ggaccctgag
atggggggaa agccgagaag gaagaaccct 1200caggaaggcc tgtacaatga
actgcagaaa gataagatgg cggaggccta cagtgagatt 1260gggatgaaag
gcgagcgccg gaggggcaag gggcacgatg gcctttacca gggtctcagt
1320acagccacca aggacaccta cgacgccctt cacatgcagg ccctgcctcc
tcgcagagcc 1380gagggcaggg gaagtcttct aacatgcggg gacgtggagg
aaaatcccgg gcccatggag 1440accgacaccc tgctgctgtg ggtgctgctg
ctgtgggtgc caggcagcac cggccaggtg 1500cagctgcagg agtctggccc
aggcctggtg aagcccagcc agaccctgag catcacctgc 1560accgtgagcg
gcttcagcct ggccagctac aacatccact gggtgcggca gcccccaggc
1620aagggcctgg agtggctggg cgtgatctgg gctggcggca gcaccaacta
caacagcgcc 1680ctgatgagcc ggctgaccat cagcaaggac aacagcaaga
accaggtgtt cctgaagatg 1740agcagcctga cagccgccga caccgccgtg
tactactgcg ccaagcggag cgacgactac 1800agctggttcg cctactgggg
ccagggcacc ctggtgaccg tgagctctgg cggaggcggc 1860tctggcggag
gcggctctgg cggaggcggc agcgagaacc agatgaccca gagccccagc
1920agcttgagcg ccagcgtggg cgaccgggtg accatgacct gcagagccag
cagcagcgtg 1980agcagcagct acctgcactg gtaccagcag aagagcggca
aggccccaaa ggtgtggatc 2040tacagcacca gcaacctggc cagcggcgtg
cccagccggt tcagcggcag cggcagcggc 2100accgactaca ccctgaccat
cagcagcctg cagcccgagg acttcgccac ctactactgc 2160cagcagtaca
gcggctaccc catcaccttc ggccagggca ccaaggtgga gatcaagcgg
2220tcggatcccg ccgagcccaa atctcctgac aaaactcaca catgcccacc
gtgcccagca 2280cctcccgtgg ccggcccgtc agtcttcctc ttccccccaa
aacccaagga caccctcatg 2340atcgcccgga cccctgaggt cacatgcgtg
gtggtggacg tgagccacga agaccctgag 2400gtcaagttca actggtacgt
ggacggcgtg gaggtgcata atgccaagac aaagccgcgg 2460gaggagcagt
acaacagcac gtaccgtgtg gtcagcgtcc tcaccgtcct gcaccaggac
2520tggctgaatg gcaaggagta caagtgcaag gtctccaaca aagccctccc
agcccccatc 2580gagaaaacca tctccaaagc caaagggcag ccccgagaac
cacaggtgta caccctgccc 2640ccatcccggg atgagctgac caagaaccag
gtcagcctga cctgcctggt caaaggcttc 2700tatcccagcg acatcgccgt
ggagtgggag agcaatgggc aaccggagaa caactacaag 2760accacgcctc
ccgtgctgga ctccgacggc tccttcttcc tctacagcaa gctcaccgtg
2820gacaagagca ggtggcagca ggggaacgtc ttctcatgct ccgtgatgca
tgaggccctg 2880cacaatcact atacccagaa atctctgagt ctgagcccag
gcaagaagga ccccaaggcg 2940gtttttggct gtatctttgg tgccctggtt
attgtgactg tgggaggctt catcttctgg 3000agaaagaaga ggaaagatgc
aaagaataat gaagtgtcct tttctcaaat taaacctaaa 3060aaatctaagt
taatcagagt ggagaatttt gaggcctact tcaagaagca gcaagctgac
3120tccaactgtg ggttcgcaga ggaatacgaa gatctgaagc ttgttggaat
tagtcaacct 3180aaatatgcag cagaactggc tgagaataga ggaaagaatc
gctataataa tgttctgccc 3240tatgatattt cccgtgtcaa actttcggtc
cagacccatt caacggatga ctacatcaat 3300gccaactaca tgcctggcta
ccactccaag aaagatttta ttgccacaca aggaccttta 3360ccgaacactt
tgaaagattt ttggcgtatg gtttgggaga aaaatgtata tgccatcatt
3420atgttgacta aatgtgttga acagggaaga accaaatgtg aggagtattg
gccctccaag 3480caggctcagg actatggaga cataactgtg gcaatgacat
cagaaattgt tcttccggaa 3540tggaccatca gagatttcac agtgaaaaat
atccagacaa gtgagagtca ccctctgaga 3600cagttccatt tcacctcctg
gccagaccac ggtgttcccg acaccactga cctgctcatc 3660aacttccggt
acctcgttcg tgactacatg aagcagagtc ctcccgaatc gccgattctg
3720gtgcattgca gtgctggggt cggaaggacg ggcactttca ttgccattga
tcgtctcatc 3780taccagatag agaatgagaa caccgtggat gtgtatggga
ttgtgtatga ccttcgaatg 3840cataggcctt taatggtgca gacagaggac
cagtatgttt tcctcaatca gtgtgttttg 3900gatattgtca gatcccagaa
agactcaaaa gtagatctta tctaccagaa cacaactgca 3960atgacaatct
atgaaaacct tgcgcccgtg accacatttg gaaagaccaa tggttacatc 4020gcctaa
4026431341PRTArtificial SequenceSingle-chain variable fragment
(scFv) SFG.aCD19-CD8STK-CD28tmZ-2A-aCD5-HCH2CH3pvaa-dCD148 43Met
Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1 5 10
15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser Leu
20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg Ala
Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln Lys
Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His Thr Ser Arg
Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly Ser
Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln Glu
Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu Pro
Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys Ala
Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135 140
Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly 145
150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr Val
Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile Arg
Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile Trp
Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser Arg
Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe Leu
Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile Tyr
Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255 Met
Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260 265
270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala
275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala
Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala Cys
Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly Gly Val Leu
Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe Ile Ile Phe
Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala Asp Ala Pro
Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365 Glu Leu Asn
Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370 375 380 Arg
Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro 385 390
395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu
Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly
Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr
Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala Leu Pro Pro
Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr Cys Gly Asp
Val Glu Glu Asn Pro Gly Pro Met Glu 465 470 475 480 Thr Asp Thr Leu
Leu Leu Trp Val Leu Leu Leu Trp Val Pro Gly Ser 485 490 495 Thr Gly
Gln Val Thr Leu Lys Glu Ser Gly Pro Gly Ile Leu Lys Pro 500 505 510
Ser Gln Thr Leu Ser Leu Thr Cys Ser Phe Ser Gly Phe Ser Leu Ser 515
520 525 Thr Ser Gly Met Gly Val Gly Trp Ile Arg Gln Pro Ser Gly Lys
Gly 530 535 540 Leu Glu Trp Leu Ala His Ile Trp Trp Asp Asp Asp Val
Tyr Tyr Asn 545 550 555 560 Pro Ser Leu Lys Asn Gln Leu Thr Ile Ser
Lys Asp Ala Ser Arg Asp 565 570 575 Gln Val Phe Leu Lys Ile Thr Asn
Leu Asp Thr Ala Asp Thr Ala Thr 580 585 590 Tyr Tyr Cys Val Arg Arg
Arg Ala Thr Gly Thr Gly Phe Asp Tyr Trp 595 600 605 Gly Gln Gly Thr
Thr Leu Thr Val Ser Ser Gly Gly Gly Gly Ser Gly 610 615 620 Gly Gly
Gly Ser Gly Gly Gly Gly Ser Asn Ile Val Met Thr Gln Ser 625 630 635
640 His Lys Phe Met Ser Thr Ser Val Gly Asp Arg Val Ser Ile Ala Cys
645 650 655 Lys Ala Ser Gln Asp Val Gly Thr Ala Val Ala Trp Tyr Gln
Gln Lys 660 665 670 Pro Gly Gln Ser Pro Lys Leu Leu Ile Tyr Trp Thr
Ser Thr Arg His 675 680 685 Thr Gly Val Pro Asp Arg Phe Thr Gly Ser
Gly Ser Gly Thr Asp Phe 690 695 700 Thr Leu Thr Ile Thr Asn Val Gln
Ser Glu Asp Leu Ala Asp Tyr Phe 705 710 715 720 Cys His Gln Tyr Asn
Ser Tyr Asn Thr Phe Gly Ser Gly Thr Arg Leu 725 730 735 Glu Leu Lys
Arg Ser Asp Pro Ala Glu Pro Lys Ser Pro Asp Lys Thr 740 745 750 His
Thr Cys Pro Pro Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val 755 760
765 Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ala Arg Thr
770 775 780 Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp
Pro Glu 785 790 795 800 Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His Asn Ala Lys 805 810 815 Thr Lys Pro Arg Glu Glu Gln Tyr Asn
Ser Thr Tyr Arg Val Val Ser 820 825 830 Val Leu Thr Val Leu His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys 835 840 845 Cys Lys Val Ser Asn
Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile 850 855 860 Ser Lys Ala
Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro 865 870 875 880
Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu 885
890 895 Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Asn 900 905 910 Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp Ser 915 920 925 Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr
Val Asp Lys Ser Arg 930 935 940 Trp Gln Gln Gly Asn Val Phe Ser Cys
Ser Val Met His Glu Ala Leu 945 950 955 960 His Asn His Tyr Thr Gln
Lys Ser Leu Ser Leu Ser Pro Gly Lys Lys 965 970 975 Asp Pro Lys Ala
Val Phe Gly Cys Ile Phe Gly Ala Leu Val Ile Val 980 985 990 Thr Val
Gly Gly Phe Ile Phe Trp Arg Lys Lys Arg Lys Asp Ala Lys 995 1000
1005 Asn Asn Glu Val Ser Phe Ser Gln Ile Lys Pro Lys Lys Ser Lys
1010 1015 1020 Leu Ile Arg Val Glu Asn Phe Glu Ala Tyr Phe Lys Lys
Gln Gln 1025 1030 1035 Ala Asp Ser Asn Cys Gly Phe Ala Glu Glu Tyr
Glu Asp Leu Lys 1040 1045 1050 Leu Val Gly Ile Ser Gln Pro Lys Tyr
Ala Ala Glu Leu Ala Glu 1055 1060 1065 Asn Arg Gly Lys Asn Arg Tyr
Asn Asn Val Leu Pro Tyr Asp Ile 1070 1075 1080 Ser Arg Val Lys Leu
Ser Val Gln Thr His Ser Thr Asp Asp Tyr 1085 1090 1095 Ile Asn Ala
Asn Tyr Met Pro Gly Tyr His Ser Lys Lys Asp Phe 1100 1105 1110 Ile
Ala Thr Gln Gly Pro Leu Pro Asn Thr Leu Lys Asp Phe Trp 1115 1120
1125 Arg Met Val Trp Glu Lys Asn Val Tyr Ala Ile Ile Met Leu Thr
1130 1135 1140 Lys Cys Val Glu Gln Gly Arg Thr Lys Cys Glu Glu Tyr
Trp Pro 1145 1150 1155 Ser Lys Gln Ala Gln Asp Tyr Gly Asp Ile Thr
Val Ala Met Thr 1160 1165 1170 Ser Glu Ile Val Leu Pro Glu Trp Thr
Ile Arg Asp Phe Thr Val 1175 1180 1185 Lys Asn Ile Gln Thr Ser Glu
Ser His Pro Leu Arg Gln Phe His 1190 1195 1200 Phe Thr Ser Trp Pro
Asp His Gly Val Pro Asp Thr Thr Asp Leu 1205 1210 1215 Leu Ile Asn
Phe Arg Tyr Leu Val Arg Asp Tyr Met Lys Gln Ser 1220 1225 1230 Pro
Pro Glu Ser Pro Ile Leu Val His Cys Ser Ala Gly Val Gly 1235 1240
1245 Arg Thr Gly Thr Phe Ile Ala Ile Asp Arg Leu Ile Tyr Gln Ile
1250 1255 1260 Glu Asn Glu Asn Thr Val Asp Val Tyr Gly Ile Val Tyr
Asp Leu 1265 1270 1275 Arg Met His Arg Pro Leu Met Val Gln Thr Glu
Asp Gln Tyr Val 1280 1285 1290 Phe Leu Asn Gln Cys Val Leu Asp Ile
Val Arg Ser Gln Lys Asp 1295 1300 1305 Ser Lys Val Asp Leu Ile Tyr
Gln Asn Thr Thr Ala Met Thr Ile 1310 1315 1320 Tyr Glu Asn Leu Ala
Pro Val Thr Thr Phe Gly Lys Thr Asn Gly 1325 1330 1335 Tyr Ile Ala
1340 444026DNAArtificial SequenceSingle-chain variable fragment
(scFv)
SFG.aCD19-CD8STK-CD28tmZ-2A-aCD5-HCH2CH3pvaa-dCD148 44atgagcctgc
ccgtgaccgc cctgctgctg cccctggccc tgctgctgca cgccgccaga 60ccagacatcc
agatgaccca gaccaccagc agcctgagcg ccagcctggg cgaccgggtg
120accatcagct gcagagccag ccaggacatc agcaagtacc tgaactggta
ccagcagaag 180cccgacggca ccgtgaagct gctgatctac cacaccagcc
ggctgcacag cggcgtgccc 240agccggttca gcggcagcgg cagcggcacc
gactacagcc tgaccatcag caacctggag 300caggaggaca tcgccaccta
cttctgccag cagggcaaca ccctgcccta caccttcgga 360ggcggcacca
agctggagat caccaaggcc ggaggcggag gctctggcgg aggcggctct
420ggcggaggcg gctctggcgg aggcggcagc gaggtgaagc tgcaggagtc
tggcccaggc 480ctggtggccc caagccagag cctgagcgtg acctgcaccg
tgagcggcgt gagcctgccc 540gactacggcg tgagctggat caggcagccc
ccacggaagg gcctggagtg gctgggcgtg 600atctggggca gcgagaccac
ctactacaac agcgccctga agagccggct gaccatcatc 660aaggacaaca
gcaagagcca ggtgttcctg aagatgaaca gcctgcagac cgacgacacc
720gccatctact actgcgccaa gcactactac tatggcggca gctacgctat
ggactactgg 780ggccagggca ccagcgtgac cgtgagctca gatcccacca
cgacgccagc gccgcgacca 840ccaacaccgg cgcccaccat cgcgtcgcag
cccctgtccc tgcgcccaga ggcgtgccgg 900ccagcggcgg ggggcgcagt
gcacacgagg gggctggact tcgcctgtga tatcttttgg 960gtgctggtgg
tggttggtgg agtcctggct tgctatagct tgctagtaac agtggccttt
1020attattttct gggtgaggag agtgaagttc agcaggagcg cagacgcccc
cgcgtaccag 1080cagggccaga accagctcta taacgagctc aatctaggac
gaagagagga gtacgatgtt 1140ttggacaaga gacgtggccg ggaccctgag
atggggggaa agccgagaag gaagaaccct 1200caggaaggcc tgtacaatga
actgcagaaa gataagatgg cggaggccta cagtgagatt 1260gggatgaaag
gcgagcgccg gaggggcaag gggcacgatg gcctttacca gggtctcagt
1320acagccacca aggacaccta cgacgccctt cacatgcagg ccctgcctcc
tcgcagagcc 1380gagggcaggg gaagtcttct aacatgcggg gacgtggagg
aaaatcccgg gcccatggag 1440accgacaccc tgctgctgtg ggtgctgctg
ctgtgggtgc ccggcagcac cggccaggtg 1500accctgaagg agagcggtcc
cggcatcctg aagcccagcc agaccctgag cctgacctgc 1560agcttcagcg
gcttcagcct gagcaccagc ggcatgggcg tgggctggat tcggcagccc
1620agcggcaagg gcctggagtg gctggcccac atctggtggg acgacgacgt
gtactacaac 1680cccagcctga agaaccagct gaccatcagc aaggacgcca
gccgggacca ggtgttcctg 1740aagatcacca acctggacac cgccgacacc
gccacctact actgcgtgcg gcgccgggcc 1800accggcaccg gcttcgacta
ctggggccag ggcaccaccc tgaccgtgag cagcggtggc 1860ggtggcagcg
gcggcggcgg aagcggaggt ggtggcagca acatcgtgat gacccagagc
1920cacaagttca tgagcaccag cgtgggcgac cgggtgagca tcgcctgcaa
ggccagccag 1980gacgtgggca ccgccgtggc ctggtaccag cagaagcctg
gccagagccc caagctgctg 2040atctactgga ccagcacccg gcacaccggc
gtgcccgacc ggttcaccgg cagcggcagc 2100ggcaccgact tcaccctgac
catcaccaac gtgcagagcg aggacctggc cgactacttc 2160tgccaccagt
acaacagcta caacaccttc ggcagcggca cccggctgga gctgaagcgg
2220tcggatcccg ccgagcccaa atctcctgac aaaactcaca catgcccacc
gtgcccagca 2280cctcccgtgg ccggcccgtc agtcttcctc ttccccccaa
aacccaagga caccctcatg 2340atcgcccgga cccctgaggt cacatgcgtg
gtggtggacg tgagccacga agaccctgag 2400gtcaagttca actggtacgt
ggacggcgtg gaggtgcata atgccaagac aaagccgcgg 2460gaggagcagt
acaacagcac gtaccgtgtg gtcagcgtcc tcaccgtcct gcaccaggac
2520tggctgaatg gcaaggagta caagtgcaag gtctccaaca aagccctccc
agcccccatc 2580gagaaaacca tctccaaagc caaagggcag ccccgagaac
cacaggtgta caccctgccc 2640ccatcccggg atgagctgac caagaaccag
gtcagcctga cctgcctggt caaaggcttc 2700tatcccagcg acatcgccgt
ggagtgggag agcaatgggc aaccggagaa caactacaag 2760accacgcctc
ccgtgctgga ctccgacggc tccttcttcc tctacagcaa gctcaccgtg
2820gacaagagca ggtggcagca ggggaacgtc ttctcatgct ccgtgatgca
tgaggccctg 2880cacaatcact atacccagaa atctctgagt ctgagcccag
gcaagaagga ccccaaggcg 2940gtttttggct gtatctttgg tgccctggtt
attgtgactg tgggaggctt catcttctgg 3000agaaagaaga ggaaagatgc
aaagaataat gaagtgtcct tttctcaaat taaacctaaa 3060aaatctaagt
taatcagagt ggagaatttt gaggcctact tcaagaagca gcaagctgac
3120tccaactgtg ggttcgcaga ggaatacgaa gatctgaagc ttgttggaat
tagtcaacct 3180aaatatgcag cagaactggc tgagaataga ggaaagaatc
gctataataa tgttctgccc 3240tatgatattt cccgtgtcaa actttcggtc
cagacccatt caacggatga ctacatcaat 3300gccaactaca tgcctggcta
ccactccaag aaagatttta ttgccacaca aggaccttta 3360ccgaacactt
tgaaagattt ttggcgtatg gtttgggaga aaaatgtata tgccatcatt
3420atgttgacta aatgtgttga acagggaaga accaaatgtg aggagtattg
gccctccaag 3480caggctcagg actatggaga cataactgtg gcaatgacat
cagaaattgt tcttccggaa 3540tggaccatca gagatttcac agtgaaaaat
atccagacaa gtgagagtca ccctctgaga 3600cagttccatt tcacctcctg
gccagaccac ggtgttcccg acaccactga cctgctcatc 3660aacttccggt
acctcgttcg tgactacatg aagcagagtc ctcccgaatc gccgattctg
3720gtgcattgca gtgctggggt cggaaggacg ggcactttca ttgccattga
tcgtctcatc 3780taccagatag agaatgagaa caccgtggat gtgtatggga
ttgtgtatga ccttcgaatg 3840cataggcctt taatggtgca gacagaggac
cagtatgttt tcctcaatca gtgtgttttg 3900gatattgtca gatcccagaa
agactcaaaa gtagatctta tctaccagaa cacaactgca 3960atgacaatct
atgaaaacct tgcgcccgtg accacatttg gaaagaccaa tggttacatc 4020gcctaa
4026451342PRTArtificial SequenceSingle-chain variable fragment
(scFv) SFG.aCD19-CD8STK-CD28tmZ-2A-aEGFRvIII-HCH2CH3pvaa-dCD148
45Met Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu Leu 1
5 10 15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr Ser Ser
Leu 20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser Cys Arg
Ala Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr Gln Gln
Lys Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His Thr Ser
Arg Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly Ser Gly
Ser Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu Glu Gln
Glu Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn Thr Leu
Pro Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120 125 Lys
Ala Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly 130 135
140 Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly Pro Gly
145 150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr Cys Thr
Val Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser Trp Ile
Arg Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly Val Ile
Trp Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu Lys Ser
Arg Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln Val Phe
Leu Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240 Ala Ile
Tyr Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245 250 255
Met Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp Pro 260
265 270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro Thr Ile
Ala 275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys Arg Pro
Ala Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp Phe Ala
Cys Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly Gly Val
Leu Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe Ile Ile
Phe Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala Asp Ala
Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365 Glu Leu
Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370 375 380
Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro 385
390 395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala
Glu Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg Arg Arg
Gly Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser Thr Ala
Thr Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala Leu Pro
Pro Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr Cys Gly
Asp Val Glu Glu Asn Pro Gly Pro Met Glu 465 470 475 480 Thr Asp Thr
Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro Gly Ser 485 490 495 Thr
Gly Gln Val Lys Leu Gln Gln Ser Gly Gly Gly Leu Val Lys Pro 500 505
510 Gly Ala Ser Leu Lys Leu Ser Cys Val Thr Ser Gly Phe Thr Phe Arg
515 520 525 Lys Phe Gly Met Ser Trp Val Arg Gln Thr Ser Asp Lys Arg
Leu Glu 530 535 540 Trp Val Ala Ser Ile Ser Thr Gly Gly Tyr Asn Thr
Tyr Tyr Ser Asp 545 550 555 560 Asn Val Lys Gly Arg Phe Thr Ile Ser
Arg Glu Asn Ala Lys Asn Thr 565 570 575 Leu Tyr Leu Gln Met Ser Ser
Leu Lys Ser Glu Asp Thr Ala Leu Tyr 580 585 590 Tyr Cys Thr Arg Gly
Tyr Ser Ser Thr Ser Tyr Ala Met Asp Tyr Trp 595 600 605 Gly Gln Gly
Thr Thr Val Thr Val Ser Ser Gly Gly Gly Gly Ser Gly 610 615 620 Gly
Gly Gly Ser Gly Gly Gly Gly Ser Asp Ile Glu Leu Thr Gln Ser 625 630
635 640 Pro Ala Ser Leu Ser Val Ala Thr Gly Glu Lys Val Thr Ile Arg
Cys 645 650 655 Met Thr Ser Thr Asp Ile Asp Asp Asp Met Asn Trp Tyr
Gln Gln Lys 660 665 670 Pro Gly Glu Pro Pro Lys Phe Leu Ile Ser Glu
Gly Asn Thr Leu Arg 675 680 685 Pro Gly Val Pro Ser Arg Phe Ser Ser
Ser Gly Thr Gly Thr Asp Phe 690 695 700 Val Phe Thr Ile Glu Asn Thr
Leu Ser Glu Asp Val Gly Asp Tyr Tyr 705 710 715 720 Cys Leu Gln Ser
Phe Asn Val Pro Leu Thr Phe Gly Asp Gly Thr Lys 725 730 735 Leu Glu
Ile Lys Arg Ser Asp Pro Ala Glu Pro Lys Ser Pro Asp Lys 740 745 750
Thr His Thr Cys Pro Pro Cys Pro Ala Pro Pro Val Ala Gly Pro Ser 755
760 765 Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ala
Arg 770 775 780 Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser His
Glu Asp Pro 785 790 795 800 Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
Val Glu Val His Asn Ala 805 810 815 Lys Thr Lys Pro Arg Glu Glu Gln
Tyr Asn Ser Thr Tyr Arg Val Val 820 825 830 Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu Asn Gly Lys Glu Tyr 835 840 845 Lys Cys Lys Val
Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr 850 855 860 Ile Ser
Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu 865 870 875
880 Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys
885 890 895 Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp
Glu Ser 900 905 910 Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro
Pro Val Leu Asp 915 920 925 Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
Leu Thr Val Asp Lys Ser 930 935 940 Arg Trp Gln Gln Gly Asn Val Phe
Ser Cys Ser Val Met His Glu Ala 945 950 955 960 Leu His Asn His Tyr
Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 965 970 975 Lys Asp Pro
Lys Ala Val Phe Gly Cys Ile Phe Gly Ala Leu Val Ile 980 985 990 Val
Thr Val Gly Gly Phe Ile Phe Trp Arg Lys Lys Arg Lys Asp Ala 995
1000 1005 Lys Asn Asn Glu Val Ser Phe Ser Gln Ile Lys Pro Lys Lys
Ser 1010 1015 1020 Lys Leu Ile Arg Val Glu Asn Phe Glu Ala Tyr Phe
Lys Lys Gln 1025 1030 1035 Gln Ala Asp Ser Asn Cys Gly Phe Ala Glu
Glu Tyr Glu Asp Leu 1040 1045 1050 Lys Leu Val Gly Ile Ser Gln Pro
Lys Tyr Ala Ala Glu Leu Ala 1055 1060 1065 Glu Asn Arg Gly Lys Asn
Arg Tyr Asn Asn Val Leu Pro Tyr Asp 1070 1075 1080 Ile Ser Arg Val
Lys Leu Ser Val Gln Thr His Ser Thr Asp Asp 1085 1090 1095 Tyr Ile
Asn Ala Asn Tyr Met Pro Gly Tyr His Ser Lys Lys Asp 1100 1105 1110
Phe Ile Ala Thr Gln Gly Pro Leu Pro Asn Thr Leu Lys Asp Phe 1115
1120 1125 Trp Arg Met Val Trp Glu Lys Asn Val Tyr Ala Ile Ile Met
Leu 1130 1135 1140 Thr Lys Cys Val Glu Gln Gly Arg Thr Lys Cys Glu
Glu Tyr Trp 1145 1150 1155 Pro Ser Lys Gln Ala Gln Asp Tyr Gly Asp
Ile Thr Val Ala Met 1160 1165 1170 Thr Ser Glu Ile Val Leu Pro Glu
Trp Thr Ile Arg Asp Phe Thr 1175 1180 1185 Val Lys Asn Ile Gln Thr
Ser Glu Ser His Pro Leu Arg Gln Phe 1190 1195 1200 His Phe Thr Ser
Trp Pro Asp His Gly Val Pro Asp Thr Thr Asp 1205 1210 1215 Leu Leu
Ile Asn Phe Arg Tyr Leu Val Arg Asp Tyr Met Lys Gln 1220 1225 1230
Ser Pro Pro Glu Ser Pro Ile Leu Val His Cys Ser Ala Gly Val 1235
1240 1245 Gly Arg Thr Gly Thr Phe Ile Ala Ile Asp Arg Leu Ile Tyr
Gln 1250 1255 1260 Ile Glu Asn Glu Asn Thr Val Asp Val Tyr Gly Ile
Val Tyr Asp 1265 1270 1275 Leu Arg Met His Arg Pro Leu Met Val Gln
Thr Glu Asp Gln Tyr 1280 1285 1290 Val Phe Leu Asn Gln Cys Val Leu
Asp Ile Val Arg Ser Gln Lys 1295 1300 1305 Asp Ser Lys Val Asp Leu
Ile Tyr Gln Asn Thr Thr Ala Met Thr 1310 1315 1320 Ile Tyr Glu Asn
Leu Ala Pro Val Thr Thr Phe Gly Lys Thr Asn 1325 1330 1335 Gly Tyr
Ile Ala 1340 464029DNAArtificial SequenceSingle-chain variable
fragment (scFv)
SFG.aCD19-CD8STK-CD28tmZ-2A-aEGFRvIII-HCH2CH3pvaa-dCD148
46atgagcctgc ccgtgaccgc cctgctgctg cccctggccc tgctgctgca cgccgccaga
60ccagacatcc agatgaccca gaccaccagc agcctgagcg ccagcctggg cgaccgggtg
120accatcagct gcagagccag ccaggacatc agcaagtacc tgaactggta
ccagcagaag 180cccgacggca ccgtgaagct gctgatctac cacaccagcc
ggctgcacag cggcgtgccc 240agccggttca gcggcagcgg cagcggcacc
gactacagcc tgaccatcag caacctggag 300caggaggaca tcgccaccta
cttctgccag cagggcaaca ccctgcccta caccttcgga 360ggcggcacca
agctggagat caccaaggcc ggaggcggag gctctggcgg aggcggctct
420ggcggaggcg gctctggcgg aggcggcagc gaggtgaagc tgcaggagtc
tggcccaggc 480ctggtggccc caagccagag cctgagcgtg acctgcaccg
tgagcggcgt gagcctgccc 540gactacggcg tgagctggat caggcagccc
ccacggaagg gcctggagtg gctgggcgtg 600atctggggca gcgagaccac
ctactacaac agcgccctga agagccggct gaccatcatc 660aaggacaaca
gcaagagcca ggtgttcctg aagatgaaca gcctgcagac cgacgacacc
720gccatctact actgcgccaa gcactactac tatggcggca gctacgctat
ggactactgg 780ggccagggca ccagcgtgac cgtgagctca gatcccacca
cgacgccagc gccgcgacca 840ccaacaccgg cgcccaccat cgcgtcgcag
cccctgtccc tgcgcccaga ggcgtgccgg 900ccagcggcgg ggggcgcagt
gcacacgagg gggctggact tcgcctgtga tatcttttgg 960gtgctggtgg
tggttggtgg agtcctggct tgctatagct tgctagtaac agtggccttt
1020attattttct gggtgaggag agtgaagttc agcaggagcg cagacgcccc
cgcgtaccag 1080cagggccaga accagctcta taacgagctc aatctaggac
gaagagagga gtacgatgtt 1140ttggacaaga gacgtggccg ggaccctgag
atggggggaa agccgagaag gaagaaccct 1200caggaaggcc tgtacaatga
actgcagaaa gataagatgg cggaggccta cagtgagatt 1260gggatgaaag
gcgagcgccg gaggggcaag gggcacgatg gcctttacca gggtctcagt
1320acagccacca aggacaccta cgacgccctt cacatgcagg ccctgcctcc
tcgcagagcc 1380gagggcaggg gaagtcttct aacatgcggg gacgtggagg
aaaatcccgg gcccatggag 1440accgacaccc tgctgctgtg ggtgctgctg
ctgtgggtgc ccggcagcac cggccaggtg 1500aagctgcagc agagcggcgg
aggcctggtg aagcccggcg ccagcctgaa gctgagctgc 1560gtgaccagcg
gcttcacctt ccggaagttc ggcatgagct gggtgcggca gaccagcgac
1620aagcggctgg agtgggtggc cagcatcagc accggcggct acaacaccta
ctacagcgac 1680aacgtgaagg gccggttcac catcagccgg gagaacgcca
agaacaccct gtacctgcag 1740atgagcagcc tgaagagcga ggacaccgcc
ctgtactact gcacccgggg ctacagcagc 1800accagctacg
ctatggacta ctggggccag ggcaccaccg tgacagtgag cagcggcgga
1860ggaggcagtg gtgggggtgg atctggcgga ggtggcagcg acatcgagct
gacccagagc 1920cccgccagcc tgagcgtggc caccggcgag aaggtgacca
tccggtgcat gaccagcacc 1980gacatcgacg acgacatgaa ctggtaccag
cagaagcccg gcgagccccc aaagttcctg 2040atcagcgagg gcaacaccct
gcggcccggc gtgcccagcc ggttcagcag cagcggcacc 2100ggcaccgact
tcgtgttcac catcgagaac accctgagcg aggacgtggg cgactactac
2160tgcctgcaga gcttcaacgt gcccctgacc ttcggcgacg gcaccaagct
ggagatcaag 2220cggtcggatc ccgccgagcc caaatctcct gacaaaactc
acacatgccc accgtgccca 2280gcacctcccg tggccggccc gtcagtcttc
ctcttccccc caaaacccaa ggacaccctc 2340atgatcgccc ggacccctga
ggtcacatgc gtggtggtgg acgtgagcca cgaagaccct 2400gaggtcaagt
tcaactggta cgtggacggc gtggaggtgc ataatgccaa gacaaagccg
2460cgggaggagc agtacaacag cacgtaccgt gtggtcagcg tcctcaccgt
cctgcaccag 2520gactggctga atggcaagga gtacaagtgc aaggtctcca
acaaagccct cccagccccc 2580atcgagaaaa ccatctccaa agccaaaggg
cagccccgag aaccacaggt gtacaccctg 2640cccccatccc gggatgagct
gaccaagaac caggtcagcc tgacctgcct ggtcaaaggc 2700ttctatccca
gcgacatcgc cgtggagtgg gagagcaatg ggcaaccgga gaacaactac
2760aagaccacgc ctcccgtgct ggactccgac ggctccttct tcctctacag
caagctcacc 2820gtggacaaga gcaggtggca gcaggggaac gtcttctcat
gctccgtgat gcatgaggcc 2880ctgcacaatc actataccca gaaatctctg
agtctgagcc caggcaagaa ggaccccaag 2940gcggtttttg gctgtatctt
tggtgccctg gttattgtga ctgtgggagg cttcatcttc 3000tggagaaaga
agaggaaaga tgcaaagaat aatgaagtgt ccttttctca aattaaacct
3060aaaaaatcta agttaatcag agtggagaat tttgaggcct acttcaagaa
gcagcaagct 3120gactccaact gtgggttcgc agaggaatac gaagatctga
agcttgttgg aattagtcaa 3180cctaaatatg cagcagaact ggctgagaat
agaggaaaga atcgctataa taatgttctg 3240ccctatgata tttcccgtgt
caaactttcg gtccagaccc attcaacgga tgactacatc 3300aatgccaact
acatgcctgg ctaccactcc aagaaagatt ttattgccac acaaggacct
3360ttaccgaaca ctttgaaaga tttttggcgt atggtttggg agaaaaatgt
atatgccatc 3420attatgttga ctaaatgtgt tgaacaggga agaaccaaat
gtgaggagta ttggccctcc 3480aagcaggctc aggactatgg agacataact
gtggcaatga catcagaaat tgttcttccg 3540gaatggacca tcagagattt
cacagtgaaa aatatccaga caagtgagag tcaccctctg 3600agacagttcc
atttcacctc ctggccagac cacggtgttc ccgacaccac tgacctgctc
3660atcaacttcc ggtacctcgt tcgtgactac atgaagcaga gtcctcccga
atcgccgatt 3720ctggtgcatt gcagtgctgg ggtcggaagg acgggcactt
tcattgccat tgatcgtctc 3780atctaccaga tagagaatga gaacaccgtg
gatgtgtatg ggattgtgta tgaccttcga 3840atgcataggc ctttaatggt
gcagacagag gaccagtatg ttttcctcaa tcagtgtgtt 3900ttggatattg
tcagatccca gaaagactca aaagtagatc ttatctacca gaacacaact
3960gcaatgacaa tctatgaaaa ccttgcgccc gtgaccacat ttggaaagac
caatggttac 4020atcgcctaa 4029476PRTArtificial
SequenceImmunoreceptor tyrosine-based inhibition motif
(ITIM)MISC_FEATURE(1)..(1)Xaa may be Ser, Ile, Val or
Leumisc_feature(2)..(2)Xaa can be any naturally occurring amino
acidmisc_feature(4)..(5)Xaa can be any naturally occurring amino
acidMISC_FEATURE(6)..(6)Xaa may be Ile, Val or Leu 47Xaa Xaa Tyr
Xaa Xaa Xaa 1 5 481114PRTArtificial SequenceAmino acid sequence of
a AND NOT gate
(16076.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-tm-dPTP
N6) 48Met Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu Leu
Leu 1 5 10 15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr Thr
Ser Ser Leu 20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile Ser
Cys Arg Ala Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp Tyr
Gln Gln Lys Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr His
Thr Ser Arg Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser Gly
Ser Gly Ser Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn Leu
Glu Gln Glu Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110 Asn
Thr Leu Pro Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115 120
125 Lys Ala Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly
130 135 140 Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser Gly
Pro Gly 145 150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val Thr
Cys Thr Val Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val Ser
Trp Ile Arg Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu Gly
Val Ile Trp Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala Leu
Lys Ser Arg Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser Gln
Val Phe Leu Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235 240
Ala Ile Tyr Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala 245
250 255 Met Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser Asp
Pro 260 265 270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala Pro
Thr Ile Ala 275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala Cys
Arg Pro Ala Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu Asp
Phe Ala Cys Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val Gly
Gly Val Leu Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala Phe
Ile Ile Phe Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser Ala
Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360 365
Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg 370
375 380 Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn
Pro 385 390 395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp Lys
Met Ala Glu Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu Arg
Arg Arg Gly Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu Ser
Thr Ala Thr Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln Ala
Leu Pro Pro Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu Thr
Cys Gly Asp Val Glu Glu Asn Pro Gly Pro Met Ala 465 470 475 480 Val
Pro Thr Gln Val Leu Gly Leu Leu Leu Leu Trp Leu Thr Asp Ala 485 490
495 Arg Cys Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser
500 505 510 Val Gly Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu Asp
Ile Tyr 515 520 525 Phe Asn Leu Val Trp Tyr Gln Gln Lys Pro Gly Lys
Ala Pro Lys Leu 530 535 540 Leu Ile Tyr Asp Thr Asn Arg Leu Ala Asp
Gly Val Pro Ser Arg Phe 545 550 555 560 Ser Gly Ser Gly Ser Gly Thr
Gln Tyr Thr Leu Thr Ile Ser Ser Leu 565 570 575 Gln Pro Glu Asp Phe
Ala Thr Tyr Tyr Cys Gln His Tyr Lys Asn Tyr 580 585 590 Pro Leu Thr
Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys Arg Ser Gly 595 600 605 Gly
Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly 610 615
620 Gly Gly Ser Arg Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
625 630 635 640 Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly Phe 645 650 655 Thr Leu Ser Asn Tyr Gly Met His Trp Ile Arg
Gln Ala Pro Gly Lys 660 665 670 Gly Leu Glu Trp Val Ser Ser Ile Ser
Leu Asn Gly Gly Ser Thr Tyr 675 680 685 Tyr Arg Asp Ser Val Lys Gly
Arg Phe Thr Ile Ser Arg Asp Asn Ala 690 695 700 Lys Ser Thr Leu Tyr
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr 705 710 715 720 Ala Val
Tyr Tyr Cys Ala Ala Gln Asp Ala Tyr Thr Gly Gly Tyr Phe 725 730 735
Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Met Asp Pro 740
745 750 Ala Thr Thr Thr Lys Pro Val Leu Arg Thr Pro Ser Pro Val His
Pro 755 760 765 Thr Gly Thr Ser Gln Pro Gln Arg Pro Glu Asp Cys Arg
Pro Arg Gly 770 775 780 Ser Val Lys Gly Thr Gly Leu Asp Phe Ala Cys
Asp Ile Tyr Trp Ala 785 790 795 800 Pro Leu Ala Gly Ile Cys Val Ala
Leu Leu Leu Ser Leu Ile Ile Thr 805 810 815 Leu Ile Cys Tyr His Arg
Ser Arg Lys Arg Val Cys Lys Ser Gly Gly 820 825 830 Gly Ser Phe Trp
Glu Glu Phe Glu Ser Leu Gln Lys Gln Glu Val Lys 835 840 845 Asn Leu
His Gln Arg Leu Glu Gly Gln Arg Pro Glu Asn Lys Gly Lys 850 855 860
Asn Arg Tyr Lys Asn Ile Leu Pro Phe Asp His Ser Arg Val Ile Leu 865
870 875 880 Gln Gly Arg Asp Ser Asn Ile Pro Gly Ser Asp Tyr Ile Asn
Ala Asn 885 890 895 Tyr Ile Lys Asn Gln Leu Leu Gly Pro Asp Glu Asn
Ala Lys Thr Tyr 900 905 910 Ile Ala Ser Gln Gly Cys Leu Glu Ala Thr
Val Asn Asp Phe Trp Gln 915 920 925 Met Ala Trp Gln Glu Asn Ser Arg
Val Ile Val Met Thr Thr Arg Glu 930 935 940 Val Glu Lys Gly Arg Asn
Lys Cys Val Pro Tyr Trp Pro Glu Val Gly 945 950 955 960 Met Gln Arg
Ala Tyr Gly Pro Tyr Ser Val Thr Asn Cys Gly Glu His 965 970 975 Asp
Thr Thr Glu Tyr Lys Leu Arg Thr Leu Gln Val Ser Pro Leu Asp 980 985
990 Asn Gly Asp Leu Ile Arg Glu Ile Trp His Tyr Gln Tyr Leu Ser Trp
995 1000 1005 Pro Asp His Gly Val Pro Ser Glu Pro Gly Gly Val Leu
Ser Phe 1010 1015 1020 Leu Asp Gln Ile Asn Gln Arg Gln Glu Ser Leu
Pro His Ala Gly 1025 1030 1035 Pro Ile Ile Val His Cys Ser Ala Gly
Ile Gly Arg Thr Gly Thr 1040 1045 1050 Ile Ile Val Ile Asp Met Leu
Met Glu Asn Ile Ser Thr Lys Gly 1055 1060 1065 Leu Asp Cys Asp Ile
Asp Ile Gln Lys Thr Ile Gln Met Val Arg 1070 1075 1080 Ala Gln Arg
Ser Gly Met Val Gln Thr Glu Ala Gln Tyr Lys Phe 1085 1090 1095 Ile
Tyr Val Ala Ile Ala Gln Phe Ile Glu Thr Thr Lys Lys Lys 1100 1105
1110 Leu 49918PRTArtificial SequenceAmino acid sequence of a AND
NOT gate
(MP16091.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-LAIR1
tm-endo) 49Met Ser Leu Pro Val Thr Ala Leu Leu Leu Pro Leu Ala Leu
Leu Leu 1 5 10 15 His Ala Ala Arg Pro Asp Ile Gln Met Thr Gln Thr
Thr Ser Ser Leu 20 25 30 Ser Ala Ser Leu Gly Asp Arg Val Thr Ile
Ser Cys Arg Ala Ser Gln 35 40 45 Asp Ile Ser Lys Tyr Leu Asn Trp
Tyr Gln Gln Lys Pro Asp Gly Thr 50 55 60 Val Lys Leu Leu Ile Tyr
His Thr Ser Arg Leu His Ser Gly Val Pro 65 70 75 80 Ser Arg Phe Ser
Gly Ser Gly Ser Gly Thr Asp Tyr Ser Leu Thr Ile 85 90 95 Ser Asn
Leu Glu Gln Glu Asp Ile Ala Thr Tyr Phe Cys Gln Gln Gly 100 105 110
Asn Thr Leu Pro Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Thr 115
120 125 Lys Ala Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly
Gly 130 135 140 Ser Gly Gly Gly Gly Ser Glu Val Lys Leu Gln Glu Ser
Gly Pro Gly 145 150 155 160 Leu Val Ala Pro Ser Gln Ser Leu Ser Val
Thr Cys Thr Val Ser Gly 165 170 175 Val Ser Leu Pro Asp Tyr Gly Val
Ser Trp Ile Arg Gln Pro Pro Arg 180 185 190 Lys Gly Leu Glu Trp Leu
Gly Val Ile Trp Gly Ser Glu Thr Thr Tyr 195 200 205 Tyr Asn Ser Ala
Leu Lys Ser Arg Leu Thr Ile Ile Lys Asp Asn Ser 210 215 220 Lys Ser
Gln Val Phe Leu Lys Met Asn Ser Leu Gln Thr Asp Asp Thr 225 230 235
240 Ala Ile Tyr Tyr Cys Ala Lys His Tyr Tyr Tyr Gly Gly Ser Tyr Ala
245 250 255 Met Asp Tyr Trp Gly Gln Gly Thr Ser Val Thr Val Ser Ser
Asp Pro 260 265 270 Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro Ala
Pro Thr Ile Ala 275 280 285 Ser Gln Pro Leu Ser Leu Arg Pro Glu Ala
Cys Arg Pro Ala Ala Gly 290 295 300 Gly Ala Val His Thr Arg Gly Leu
Asp Phe Ala Cys Asp Ile Phe Trp 305 310 315 320 Val Leu Val Val Val
Gly Gly Val Leu Ala Cys Tyr Ser Leu Leu Val 325 330 335 Thr Val Ala
Phe Ile Ile Phe Trp Val Arg Arg Val Lys Phe Ser Arg 340 345 350 Ser
Ala Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn Gln Leu Tyr Asn 355 360
365 Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg
370 375 380 Arg Gly Arg Asp Pro Glu Met Gly Gly Lys Pro Arg Arg Lys
Asn Pro 385 390 395 400 Gln Glu Gly Leu Tyr Asn Glu Leu Gln Lys Asp
Lys Met Ala Glu Ala 405 410 415 Tyr Ser Glu Ile Gly Met Lys Gly Glu
Arg Arg Arg Gly Lys Gly His 420 425 430 Asp Gly Leu Tyr Gln Gly Leu
Ser Thr Ala Thr Lys Asp Thr Tyr Asp 435 440 445 Ala Leu His Met Gln
Ala Leu Pro Pro Arg Arg Ala Glu Gly Arg Gly 450 455 460 Ser Leu Leu
Thr Cys Gly Asp Val Glu Glu Asn Pro Gly Pro Met Ala 465 470 475 480
Val Pro Thr Gln Val Leu Gly Leu Leu Leu Leu Trp Leu Thr Asp Ala 485
490 495 Arg Cys Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala
Ser 500 505 510 Val Gly Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Glu
Asp Ile Tyr 515 520 525 Phe Asn Leu Val Trp Tyr Gln Gln Lys Pro Gly
Lys Ala Pro Lys Leu 530 535 540 Leu Ile Tyr Asp Thr Asn Arg Leu Ala
Asp Gly Val Pro Ser Arg Phe 545 550 555 560 Ser Gly Ser Gly Ser Gly
Thr Gln Tyr Thr Leu Thr Ile Ser Ser Leu 565 570 575 Gln Pro Glu Asp
Phe Ala Thr Tyr Tyr Cys Gln His Tyr Lys Asn Tyr 580 585 590 Pro Leu
Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys Arg Ser Gly 595 600 605
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly 610
615 620 Gly Gly Ser Arg Ser Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu 625 630 635 640 Val Gln Pro Gly Gly Ser Leu Arg Leu Ser Cys Ala
Ala Ser Gly Phe 645 650 655 Thr Leu Ser Asn Tyr Gly Met His Trp Ile
Arg Gln Ala Pro Gly Lys 660 665 670 Gly Leu Glu Trp Val Ser Ser Ile
Ser Leu Asn Gly Gly Ser Thr Tyr 675 680 685 Tyr Arg Asp Ser Val Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala 690 695 700 Lys Ser Thr Leu
Tyr Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr 705 710 715 720 Ala
Val Tyr Tyr Cys Ala Ala Gln Asp Ala Tyr Thr Gly Gly Tyr Phe 725 730
735 Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Ser Met
Asp Pro 740 745 750 Ala Thr Thr Thr Lys Pro Val Leu Arg Thr Pro Ser
Pro Val His Pro 755 760 765 Thr Gly Thr Ser Gln Pro Gln Arg Pro Glu
Asp Cys Arg Pro Arg Gly 770 775 780 Ser Val Lys Gly Thr Gly Leu Asp
Phe Ala Cys Asp Ile Leu Ile Gly 785 790 795 800 Val Ser Val Val Phe
Leu Phe Cys Leu Leu Leu Leu Val Leu Phe Cys 805 810 815 Leu His Arg
Gln Asn Gln Ile Lys Gln Gly Pro Pro Arg Ser Lys Asp 820 825 830 Glu
Glu Gln Lys Pro Gln Gln Arg Pro Asp Leu Ala Val Asp Val Leu 835 840
845 Glu Arg Thr Ala Asp Lys Ala Thr Val Asn Gly Leu Pro Glu Lys Asp
850 855 860 Arg Glu Thr Asp Thr Ser Ala Leu Ala Ala Gly Ser Ser Gln
Glu Val 865 870 875 880 Thr Tyr Ala Gln Leu Asp His Trp Ala Leu Thr
Gln Arg Thr Ala Arg 885 890 895 Ala Val Ser Pro Gln Ser Thr Lys Pro
Met Ala Glu Ser Ile Thr Tyr 900 905 910 Ala Ala Val Ala Arg His 915
501362PRTArtificial SequenceAmino acid sequence of a AND NOT gate
(MP16092.SFG.aCD19fmc63-CD8STK-CD28tmZ-2A-aCD33glx-muCD8STK-LAI- R1
tm-endo-2A-PTPN6_SH2-dCD148) 50Met Ser Leu Pro Val Thr Ala Leu Leu
Leu Pro Leu Ala Leu Leu Leu 1 5 10 15 His Ala Ala Arg Pro Asp Ile
Gln Met Thr Gln Thr Thr Ser Ser Leu 20 25 30 Ser Ala Ser Leu Gly
Asp Arg Val Thr Ile Ser Cys Arg Ala Ser Gln 35 40 45 Asp Ile Ser
Lys Tyr Leu Asn Trp Tyr Gln Gln Lys Pro Asp Gly Thr 50 55 60 Val
Lys Leu Leu Ile Tyr His Thr Ser Arg Leu His Ser Gly Val Pro 65 70
75 80 Ser Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Tyr Ser Leu Thr
Ile 85 90 95 Ser Asn Leu Glu Gln Glu Asp Ile Ala Thr Tyr Phe Cys
Gln Gln Gly 100 105 110 Asn Thr Leu Pro Tyr Thr Phe Gly Gly Gly Thr
Lys Leu Glu Ile Thr 115 120 125 Lys Ala Gly Gly Gly Gly Ser Gly Gly
Gly Gly Ser Gly Gly Gly Gly 130 135 140 Ser Gly Gly Gly Gly Ser Glu
Val Lys Leu Gln Glu Ser Gly Pro Gly 145 150 155 160 Leu Val Ala Pro
Ser Gln Ser Leu Ser Val Thr Cys Thr Val Ser Gly 165 170 175 Val Ser
Leu Pro Asp Tyr Gly Val Ser Trp Ile Arg Gln Pro Pro Arg 180 185 190
Lys Gly Leu Glu Trp Leu Gly Val Ile Trp Gly Ser Glu Thr Thr Tyr 195
200 205 Tyr Asn Ser Ala Leu Lys Ser Arg Leu Thr Ile Ile Lys Asp Asn
Ser 210 215 220 Lys Ser Gln Val Phe Leu Lys Met Asn Ser Leu Gln Thr
Asp Asp Thr 225 230 235 240 Ala Ile Tyr Tyr Cys Ala Lys His Tyr Tyr
Tyr Gly Gly Ser Tyr Ala 245 250 255 Met Asp Tyr Trp Gly Gln Gly Thr
Ser Val Thr Val Ser Ser Asp Pro 260 265 270 Thr Thr Thr Pro Ala Pro
Arg Pro Pro Thr Pro Ala Pro Thr Ile Ala 275 280 285 Ser Gln Pro Leu
Ser Leu Arg Pro Glu Ala Cys Arg Pro Ala Ala Gly 290 295 300 Gly Ala
Val His Thr Arg Gly Leu Asp Phe Ala Cys Asp Ile Phe Trp 305 310 315
320 Val Leu Val Val Val Gly Gly Val Leu Ala Cys Tyr Ser Leu Leu Val
325 330 335 Thr Val Ala Phe Ile Ile Phe Trp Val Arg Arg Val Lys Phe
Ser Arg 340 345 350 Ser Ala Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn
Gln Leu Tyr Asn 355 360 365 Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr
Asp Val Leu Asp Lys Arg 370 375 380 Arg Gly Arg Asp Pro Glu Met Gly
Gly Lys Pro Arg Arg Lys Asn Pro 385 390 395 400 Gln Glu Gly Leu Tyr
Asn Glu Leu Gln Lys Asp Lys Met Ala Glu Ala 405 410 415 Tyr Ser Glu
Ile Gly Met Lys Gly Glu Arg Arg Arg Gly Lys Gly His 420 425 430 Asp
Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp Thr Tyr Asp 435 440
445 Ala Leu His Met Gln Ala Leu Pro Pro Arg Arg Ala Glu Gly Arg Gly
450 455 460 Ser Leu Leu Thr Cys Gly Asp Val Glu Glu Asn Pro Gly Pro
Met Ala 465 470 475 480 Val Pro Thr Gln Val Leu Gly Leu Leu Leu Leu
Trp Leu Thr Asp Ala 485 490 495 Arg Cys Asp Ile Gln Met Thr Gln Ser
Pro Ser Ser Leu Ser Ala Ser 500 505 510 Val Gly Asp Arg Val Thr Ile
Thr Cys Arg Ala Ser Glu Asp Ile Tyr 515 520 525 Phe Asn Leu Val Trp
Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu 530 535 540 Leu Ile Tyr
Asp Thr Asn Arg Leu Ala Asp Gly Val Pro Ser Arg Phe 545 550 555 560
Ser Gly Ser Gly Ser Gly Thr Gln Tyr Thr Leu Thr Ile Ser Ser Leu 565
570 575 Gln Pro Glu Asp Phe Ala Thr Tyr Tyr Cys Gln His Tyr Lys Asn
Tyr 580 585 590 Pro Leu Thr Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys
Arg Ser Gly 595 600 605 Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Gly
Gly Gly Ser Gly Gly 610 615 620 Gly Gly Ser Arg Ser Glu Val Gln Leu
Val Glu Ser Gly Gly Gly Leu 625 630 635 640 Val Gln Pro Gly Gly Ser
Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe 645 650 655 Thr Leu Ser Asn
Tyr Gly Met His Trp Ile Arg Gln Ala Pro Gly Lys 660 665 670 Gly Leu
Glu Trp Val Ser Ser Ile Ser Leu Asn Gly Gly Ser Thr Tyr 675 680 685
Tyr Arg Asp Ser Val Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala 690
695 700 Lys Ser Thr Leu Tyr Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr 705 710 715 720 Ala Val Tyr Tyr Cys Ala Ala Gln Asp Ala Tyr Thr
Gly Gly Tyr Phe 725 730 735 Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr
Val Ser Ser Met Asp Pro 740 745 750 Thr Thr Thr Lys Pro Val Leu Arg
Thr Pro Ser Pro Val His Pro Thr 755 760 765 Gly Thr Ser Gln Pro Gln
Arg Pro Glu Asp Cys Arg Pro Arg Gly Ser 770 775 780 Val Lys Gly Thr
Gly Leu Asp Phe Ala Cys Asp Ile Leu Ile Gly Val 785 790 795 800 Ser
Val Val Phe Leu Phe Cys Leu Leu Leu Leu Val Leu Phe Cys Leu 805 810
815 His Arg Gln Asn Gln Ile Lys Gln Gly Pro Pro Arg Ser Lys Asp Glu
820 825 830 Glu Gln Lys Pro Gln Gln Arg Pro Asp Leu Ala Val Asp Val
Leu Glu 835 840 845 Arg Thr Ala Asp Lys Ala Thr Val Asn Gly Leu Pro
Glu Lys Asp Arg 850 855 860 Glu Thr Asp Thr Ser Ala Leu Ala Ala Gly
Ser Ser Gln Glu Val Thr 865 870 875 880 Tyr Ala Gln Leu Asp His Trp
Ala Leu Thr Gln Arg Thr Ala Arg Ala 885 890 895 Val Ser Pro Gln Ser
Thr Lys Pro Met Ala Glu Ser Ile Thr Tyr Ala 900 905 910 Ala Val Ala
Arg His Arg Ala Glu Gly Arg Gly Ser Leu Leu Thr Cys 915 920 925 Gly
Asp Val Glu Glu Asn Pro Gly Pro Trp Tyr His Gly His Met Ser 930 935
940 Gly Gly Gln Ala Glu Thr Leu Leu Gln Ala Lys Gly Glu Pro Trp Thr
945 950 955 960 Phe Leu Val Arg Glu Ser Leu Ser Gln Pro Gly Asp Phe
Val Leu Ser 965 970 975 Val Leu Ser Asp Gln Pro Lys Ala Gly Pro Gly
Ser Pro Leu Arg Val 980 985 990 Thr His Ile Lys Val Met Cys Glu Gly
Gly Arg Tyr Thr Val Gly Gly 995 1000 1005 Leu Glu Thr Phe Asp Ser
Leu Thr Asp Leu Val Glu His Phe Lys 1010 1015 1020 Lys Thr Gly Ile
Glu Glu Ala Ser Gly Ala Phe Val Tyr Leu Arg 1025 1030 1035 Gln Pro
Tyr Ser Gly Gly Gly Gly Ser Phe Glu Ala Tyr Phe Lys 1040 1045 1050
Lys Gln Gln Ala Asp Ser Asn Cys Gly Phe Ala Glu Glu Tyr Glu 1055
1060 1065 Asp Leu Lys Leu Val Gly Ile Ser Gln Pro Lys Tyr Ala Ala
Glu 1070 1075 1080 Leu Ala Glu Asn Arg Gly Lys Asn Arg Tyr Asn Asn
Val Leu Pro 1085 1090 1095 Tyr Asp Ile Ser Arg Val Lys Leu Ser Val
Gln Thr His Ser Thr 1100 1105 1110 Asp Asp Tyr Ile Asn Ala Asn Tyr
Met Pro Gly Tyr His Ser Lys 1115 1120 1125 Lys Asp Phe Ile Ala Thr
Gln Gly Pro Leu Pro Asn Thr Leu Lys 1130 1135 1140 Asp Phe Trp Arg
Met Val Trp Glu Lys Asn Val Tyr Ala Ile Ile 1145 1150 1155 Met Leu
Thr Lys Cys Val Glu Gln Gly Arg Thr Lys Cys Glu Glu 1160 1165 1170
Tyr Trp Pro Ser Lys Gln Ala Gln Asp Tyr Gly Asp Ile Thr Val 1175
1180 1185 Ala Met Thr Ser Glu Ile Val Leu Pro Glu Trp Thr Ile Arg
Asp 1190 1195 1200 Phe Thr Val Lys Asn Ile Gln Thr Ser Glu Ser His
Pro Leu Arg 1205 1210 1215 Gln Phe His Phe Thr Ser Trp Pro Asp His
Gly Val Pro Asp Thr 1220 1225 1230 Thr Asp Leu Leu Ile Asn Phe Arg
Tyr Leu Val Arg Asp Tyr Met 1235 1240 1245 Lys Gln Ser Pro Pro Glu
Ser Pro Ile Leu Val His Cys Ser Ala 1250 1255 1260 Gly Val Gly Arg
Thr Gly Thr Phe Ile Ala Ile Asp Arg Leu Ile 1265 1270 1275 Tyr Gln
Ile Glu Asn Glu Asn Thr Val Asp Val Tyr Gly Ile Val 1280 1285 1290
Tyr Asp Leu Arg Met His Arg Pro Leu Met Val Gln Thr Glu Asp 1295
1300 1305 Gln Tyr Val Phe Leu Asn Gln Cys Val Leu Asp Ile Val Arg
Ser 1310 1315 1320 Gln Lys Asp Ser Lys Val Asp Leu Ile Tyr Gln Asn
Thr Thr Ala 1325 1330 1335 Met Thr Ile Tyr Glu Asn Leu Ala Pro Val
Thr Thr Phe Gly Lys 1340 1345 1350 Thr Asn Gly Tyr Ile Ala Ser Gly
Ser 1355 1360 51424PRTArtificial SequenceAPRIL-based (A
proliferation-inducing ligand- based) CAR, CD8 stalk APRIL CAR
51Met Glu Thr Asp Thr Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro 1
5 10 15 Gly Ser Thr Gly Ser Val Leu His Leu Val Pro Ile Asn Ala Thr
Ser 20 25 30 Lys Asp Asp Ser Asp Val Thr Glu Val Met Trp Gln Pro
Ala Leu Arg 35 40 45 Arg Gly Arg Gly Leu Gln Ala Gln Gly Tyr Gly
Val Arg Ile Gln Asp 50 55 60 Ala Gly Val Tyr Leu Leu Tyr Ser Gln
Val Leu Phe Gln Asp Val Thr 65 70 75 80 Phe Thr Met Gly Gln Val Val
Ser Arg Glu Gly Gln Gly Arg Gln Glu 85 90 95 Thr Leu Phe Arg Cys
Ile Arg Ser Met Pro Ser His Pro Asp Arg Ala 100 105 110 Tyr Asn Ser
Cys Tyr Ser Ala Gly Val Phe His Leu His Gln Gly Asp 115 120 125 Ile
Leu Ser Val Ile Ile Pro Arg Ala Arg Ala Lys Leu Asn Leu Ser 130 135
140 Pro His Gly Thr Phe Leu Gly Phe Val Lys Leu Ser Gly Gly Gly Ser
145 150 155 160 Asp Pro Thr Thr Thr Pro Ala Pro Arg Pro Pro Thr Pro
Ala Pro Thr 165 170 175 Ile Ala Ser Gln Pro Leu Ser Leu Arg Pro Glu
Ala Cys Arg Pro Ala 180 185 190 Ala Gly Gly Ala Val His Thr Arg Gly
Leu Asp Phe Ala Cys Asp Ile 195 200 205 Phe Trp Val Leu Val Val Val
Gly Gly Val Leu Ala Cys Tyr Ser Leu 210 215 220 Leu Val Thr Val Ala
Phe Ile Ile Phe Trp Val Arg Ser Lys Arg Ser 225 230 235 240 Arg Leu
Leu His Ser Asp Tyr Met Asn Met Thr Pro Arg Arg Pro Gly 245 250 255
Pro Thr Arg Lys His Tyr Gln Pro Tyr Ala Pro Pro Arg Asp Phe Ala 260
265 270 Ala Tyr Arg Ser Arg Asp Gln Arg Leu Pro Pro Asp Ala His Lys
Pro 275 280 285 Pro Gly Gly Gly Ser Phe Arg Thr Pro Ile Gln Glu Glu
Gln Ala Asp 290 295 300 Ala His Ser Thr Leu Ala Lys Ile Arg Val Lys
Phe Ser Arg Ser Ala 305 310 315 320 Asp Ala Pro Ala Tyr Gln Gln Gly
Gln Asn Gln Leu Tyr Asn Glu Leu 325 330 335 Asn Leu Gly Arg Arg Glu
Glu Tyr Asp Val Leu Asp Lys Arg Arg Gly 340 345 350 Arg Asp Pro Glu
Met Gly Gly Lys Pro Arg Arg Lys Asn Pro Gln Glu 355 360 365 Gly Leu
Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu Ala Tyr Ser 370 375 380
Glu Ile Gly Met Lys Gly Glu Arg Arg Arg Gly Lys Gly His Asp Gly 385
390 395 400 Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys Asp Thr Tyr Asp
Ala Leu 405 410 415 His Met Gln Ala Leu Pro Pro Arg 420
52398PRTArtificial SequenceAPRIL-based (A proliferation-inducing
ligand- based) CAR, APRIL IgG1 hinge based CAR 52Met Glu Thr Asp
Thr Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro 1 5 10 15 Gly Ser
Thr Gly Ser Val Leu His Leu Val Pro Ile Asn Ala Thr Ser 20 25 30
Lys Asp Asp Ser Asp Val Thr Glu Val Met Trp Gln Pro Ala Leu Arg 35
40 45 Arg Gly Arg Gly Leu Gln Ala Gln Gly Tyr Gly Val Arg Ile Gln
Asp 50 55 60 Ala Gly Val Tyr Leu Leu Tyr Ser Gln Val Leu Phe Gln
Asp Val Thr 65 70 75 80 Phe Thr Met Gly Gln Val Val Ser Arg Glu Gly
Gln Gly Arg Gln Glu 85 90 95 Thr Leu Phe Arg Cys Ile Arg Ser Met
Pro Ser His Pro Asp Arg Ala 100 105 110 Tyr Asn Ser Cys Tyr Ser Ala
Gly Val Phe His Leu His Gln Gly Asp 115 120 125 Ile Leu Ser Val Ile
Ile Pro Arg Ala Arg Ala Lys Leu Asn Leu Ser 130 135 140 Pro His Gly
Thr Phe Leu Gly Phe Val Lys Leu Ser Gly Gly Gly Ser 145 150 155 160
Asp Pro Ala Glu Pro Lys Ser Pro Asp Lys Thr His Thr Cys Pro Pro 165
170 175 Cys Pro Lys Asp Pro Lys Phe Trp Val Leu Val Val Val Gly Gly
Val 180 185 190 Leu Ala Cys Tyr Ser Leu Leu Val Thr Val Ala Phe Ile
Ile Phe Trp 195 200 205 Val Arg Ser Lys Arg Ser Arg Leu Leu His Ser
Asp Tyr Met Asn Met 210 215 220 Thr Pro Arg Arg Pro Gly Pro Thr Arg
Lys His Tyr Gln Pro Tyr Ala 225 230 235 240 Pro Pro Arg Asp Phe Ala
Ala Tyr Arg Ser Arg Asp Gln Arg Leu Pro 245 250 255 Pro Asp Ala His
Lys Pro Pro Gly Gly Gly Ser Phe Arg Thr Pro Ile 260 265 270 Gln
Glu Glu Gln Ala Asp Ala His Ser Thr Leu Ala Lys Ile Arg Val 275 280
285 Lys Phe Ser Arg Ser Ala Asp Ala Pro Ala Tyr Gln Gln Gly Gln Asn
290 295 300 Gln Leu Tyr Asn Glu Leu Asn Leu Gly Arg Arg Glu Glu Tyr
Asp Val 305 310 315 320 Leu Asp Lys Arg Arg Gly Arg Asp Pro Glu Met
Gly Gly Lys Pro Arg 325 330 335 Arg Lys Asn Pro Gln Glu Gly Leu Tyr
Asn Glu Leu Gln Lys Asp Lys 340 345 350 Met Ala Glu Ala Tyr Ser Glu
Ile Gly Met Lys Gly Glu Arg Arg Arg 355 360 365 Gly Lys Gly His Asp
Gly Leu Tyr Gln Gly Leu Ser Thr Ala Thr Lys 370 375 380 Asp Thr Tyr
Asp Ala Leu His Met Gln Ala Leu Pro Pro Arg 385 390 395
53614PRTArtificial SequenceAPRIL-based (A proliferation-inducing
ligand- based) CAR, APRIL Fc-pvaa based CAR 53Met Glu Thr Asp Thr
Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro 1 5 10 15 Gly Ser Thr
Gly Ser Val Leu His Leu Val Pro Ile Asn Ala Thr Ser 20 25 30 Lys
Asp Asp Ser Asp Val Thr Glu Val Met Trp Gln Pro Ala Leu Arg 35 40
45 Arg Gly Arg Gly Leu Gln Ala Gln Gly Tyr Gly Val Arg Ile Gln Asp
50 55 60 Ala Gly Val Tyr Leu Leu Tyr Ser Gln Val Leu Phe Gln Asp
Val Thr 65 70 75 80 Phe Thr Met Gly Gln Val Val Ser Arg Glu Gly Gln
Gly Arg Gln Glu 85 90 95 Thr Leu Phe Arg Cys Ile Arg Ser Met Pro
Ser His Pro Asp Arg Ala 100 105 110 Tyr Asn Ser Cys Tyr Ser Ala Gly
Val Phe His Leu His Gln Gly Asp 115 120 125 Ile Leu Ser Val Ile Ile
Pro Arg Ala Arg Ala Lys Leu Asn Leu Ser 130 135 140 Pro His Gly Thr
Phe Leu Gly Phe Val Lys Leu Ser Gly Gly Gly Ser 145 150 155 160 Asp
Pro Ala Glu Pro Lys Ser Pro Asp Lys Thr His Thr Cys Pro Pro 165 170
175 Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu Phe Pro Pro
180 185 190 Lys Pro Lys Asp Thr Leu Met Ile Ala Arg Thr Pro Glu Val
Thr Cys 195 200 205 Val Val Val Asp Val Ser His Glu Asp Pro Glu Val
Lys Phe Asn Trp 210 215 220 Tyr Val Asp Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg Glu 225 230 235 240 Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val Ser Val Leu Thr Val Leu 245 250 255 His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn 260 265 270 Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly 275 280 285 Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu 290 295
300 Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr
305 310 315 320 Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn 325 330 335 Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe 340 345 350 Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn 355 360 365 Val Phe Ser Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr 370 375 380 Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys Lys Asp Pro Lys Phe Trp 385 390 395 400 Val Leu
Val Val Val Gly Gly Val Leu Ala Cys Tyr Ser Leu Leu Val 405 410 415
Thr Val Ala Phe Ile Ile Phe Trp Val Arg Ser Lys Arg Ser Arg Leu 420
425 430 Leu His Ser Asp Tyr Met Asn Met Thr Pro Arg Arg Pro Gly Pro
Thr 435 440 445 Arg Lys His Tyr Gln Pro Tyr Ala Pro Pro Arg Asp Phe
Ala Ala Tyr 450 455 460 Arg Ser Arg Asp Gln Arg Leu Pro Pro Asp Ala
His Lys Pro Pro Gly 465 470 475 480 Gly Gly Ser Phe Arg Thr Pro Ile
Gln Glu Glu Gln Ala Asp Ala His 485 490 495 Ser Thr Leu Ala Lys Ile
Arg Val Lys Phe Ser Arg Ser Ala Asp Ala 500 505 510 Pro Ala Tyr Gln
Gln Gly Gln Asn Gln Leu Tyr Asn Glu Leu Asn Leu 515 520 525 Gly Arg
Arg Glu Glu Tyr Asp Val Leu Asp Lys Arg Arg Gly Arg Asp 530 535 540
Pro Glu Met Gly Gly Lys Pro Arg Arg Lys Asn Pro Gln Glu Gly Leu 545
550 555 560 Tyr Asn Glu Leu Gln Lys Asp Lys Met Ala Glu Ala Tyr Ser
Glu Ile 565 570 575 Gly Met Lys Gly Glu Arg Arg Arg Gly Lys Gly His
Asp Gly Leu Tyr 580 585 590 Gln Gly Leu Ser Thr Ala Thr Lys Asp Thr
Tyr Asp Ala Leu His Met 595 600 605 Gln Ala Leu Pro Pro Arg 610
54420DNAArtificial SequenceAntiCD19 CD28TM-TCRz 54ttttgggtgc
tggtggtggt tggtggagtc ctggcttgct atagcttgct agtaacagtg 60gcctttatta
ttttctgggt gaggagagtg aagttcagca ggagcgcaga cgcccccgcg
120taccagcagg gccagaacca gctctataac gagctcaatc taggacgaag
agaggagtac 180gatgttttgg acaagagacg tggccgggac cctgagatgg
ggggaaagcc gagaaggaag 240aaccctcagg aaggcctgta caatgaactg
cagaaagata agatggcgga ggcctacagt 300gagattggga tgaaaggcga
gcgccggagg ggcaaggggc acgatggcct ttaccagggt 360ctcagtacag
ccaccaagga cacctacgac gcccttcaca tgcaggccct gcctcctcgc
42055420DNAArtificial SequenceAntiCD33 CD28TM-TCRz 55ttctgggtcc
tggtggtggt gggaggcgtg ctggcctgtt actctctcct ggtgaccgtg 60gccttcatca
tcttttgggt gcgccgggtg aagttttctc gctctgccga tgccccagcc
120tatcagcagg gccagaatca gctgtacaat gaactgaacc tgggcaggcg
ggaggagtac 180gacgtgctgg ataagcggag aggcagagac cccgagatgg
gcggcaaacc acggcgcaaa 240aatccccagg agggactcta taacgagctg
cagaaggaca aaatggccga ggcctattcc 300gagatcggca tgaagggaga
gagaagacgc ggaaagggcc acgacggcct gtatcaggga 360ttgtccaccg
ctacaaaaga tacatatgat gccctgcaca tgcaggccct gccacccaga
42056298DNAArtificial SequenceAntiCD19 CD28TM-TCRz vs. AntiCD33
CD28TM-TCRz Consensus sequence 56tttgggtctg gtggtggtgg gggtctggct
gtatctgtac gtggccttat attttgggtg 60gggtgaagtt ggcgagcccc gctacagcag
ggccagaaca gcttaaagac taactggggg 120aggagtacga gttggaaagg
gggcggaccc gagatggggg aaccggaaaa cccaggaggc 180ttaaagactg
cagaagaaaa tggcgaggcc tagagatgga tgaagggagg ggggaagggc
240acgaggcctt acagggtacg cacaagaact agagccctca catgcaggcc ctgccccg
298
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025

D00026

D00027

D00028

D00029

D00030

D00031

D00032

D00033

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.