Therapeutic Anticancer Neoepitope Vaccine
Granum; Stine ; et al.
U.S. patent application number 16/068449 was filed with the patent office on 2019-01-24 for therapeutic anticancer neoepitope vaccine. This patent application is currently assigned to VACCIBODY AS. The applicant listed for this patent is VACCIBODY AS. Invention is credited to Agnete Brunsvik Fredriksen, Stine Granum, Elisabeth Stubsrud.
| Application Number | 20190022202 16/068449 |
| Document ID | / |
| Family ID | 57821954 |
| Filed Date | 2019-01-24 |








View All Diagrams
| United States Patent Application | 20190022202 |
| Kind Code | A1 |
| Granum; Stine ; et al. | January 24, 2019 |
THERAPEUTIC ANTICANCER NEOEPITOPE VACCINE
Abstract
The present invention relates to an anticancer vaccine comprising polynucleotides or polypeptides, methods of treatment of targeting module cancer wherein such an anticancer vaccine is used as well as methods for producing the vaccine. The vaccine comprises a polynucleotide comprising a nucleotide sequence encoding a targeting unit, a dimerization unit, a first linker and an antigenic unit, wherein said antigenic unit comprises n-1 antigenic subunits, each subunit comprising at least a part of a cancer neoepitope sequence and a second linker and said antigenic unit further comprising a final cancer neoepitope sequence, wherein n is an integer of from 3 to 50, or the vaccine comprises a polypeptide encoded by the polynucleotide or a dimeric protein consisting of two polypeptides encoded by the polynucleotide.
| Inventors: | Granum; Stine; (Oslo, NO) ; Stubsrud; Elisabeth; (Oslo, NO) ; Fredriksen; Agnete Brunsvik; (R.ae butted.lingen, NO) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | VACCIBODY AS Oslo NO |
||||||||||
| Family ID: | 57821954 | ||||||||||
| Appl. No.: | 16/068449 | ||||||||||
| Filed: | January 5, 2017 | ||||||||||
| PCT Filed: | January 5, 2017 | ||||||||||
| PCT NO: | PCT/EP2017/050206 | ||||||||||
| 371 Date: | July 6, 2018 |
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61K 2039/6031 20130101; A61K 2039/53 20130101; A61K 2039/70 20130101; A61K 39/0011 20130101; A61P 35/00 20180101; A61K 2039/64 20130101; A61K 2039/6056 20130101; A61K 2039/627 20130101 |
| International Class: | A61K 39/00 20060101 A61K039/00; A61P 35/00 20060101 A61P035/00 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Jan 8, 2016 | EP | 16150574.8 |
| Jul 7, 2016 | EP | 16178393.1 |
Claims
1. A therapeutic anticancer neoepitope vaccine comprising an immunologically effective amount of a polynucleotide comprising a nucleotide sequence encoding a targeting unit a dimerization unit a first linker an antigenic unit, wherein said antigenic unit comprises n-1 antigenic subunits, each subunit comprising at least a part of a cancer neoepitope sequence and a second linker and said antigenic unit further comprising a final cancer neoepitope sequence, wherein n is an integer of from 3 to 50. or a polypeptide encoded by the polynucleotide as defined in 1), or a dimeric protein consisting of two polypeptides encoded by the polynucleotide as defined in 1).
2. The vaccine according to claim 1, wherein the antigenic unit comprises one copy of each cancer neoepitope.
3. The vaccine according to claim 1, wherein the antigenic unit comprises at least two copies of at least one neoepitope.
4. The vaccine according to any of the preceding claims, wherein the cancer neoepitope sequence has a length of from 7 to 30 amino acids.
5. The vaccine according to any of the preceding claims, wherein each cancer neoepitope sequence has identical length.
6. The vaccine according to any of the preceding claims, wherein the cancer neoepitope is positioned essentially in the middle of the cancer neoepitope sequence.
7. The vaccine according to any of the preceding claims, wherein the cancer neoepitope sequence is a subsequence of a cancer neoantigen.
8. The vaccine according to any of the preceding claims, wherein the antigenic subunits are in the order of more antigenic to less antigenic from the first linker.
9. The vaccine according to any of the preceding claims, wherein the most hydrophobic antigenic subunit(s) is(are) substantially the middle of the antigenic unit and the most hydrophilic antigenic subunit(s) is/are at the ends of the antigenic unit.
10. The vaccine according to any of the preceding claims, wherein the second linker is a flexible linker.
11. The vaccine according to any of the preceding claims, wherein the second linker is identical in all antigenic subunits.
12. The vaccine according to any of the preceding claims, wherein the second linker is a Serine-Glycine linker.
13. The vaccine according to any of the preceding claims, wherein the length of the antigenic unit is from about 300 amino acids to about a 1000 amino acids.
14. The vaccine according to any of the preceding claims, wherein the dimerization unit comprises an amino acid sequence having at least 80% sequence identity to the amino acid sequence 94-237 of SEQ ID NO:3.
15. The vaccine according to any of the preceding claims, wherein the targeting unit has affinity for a chemokine receptor selected from CCR1, CCR3 and CCR5.
16. The vaccine according to any of the preceding claims, wherein said targeting unit, dimerization unit and antigenic unit in said peptide are in the N-terminal to C-terminal order of targeting unit, dimerization unit and antigenic unit.
17. Use of a vaccine as defined in any of the claims 1-16 for treatment of cancer.
Description
FIELD OF INVENTION
[0001] The present invention relates to an anticancer vaccine comprising polynucleotides or polypeptides, methods of treatment of cancer wherein such an anticancer vaccine is used as well as methods for producing the vaccine.
BACKGROUND OF INVENTION
[0002] Although treatment of cancer has been improved over the past few decades in particularly due to early detection and diagnosis, which has significantly increased the survival, only about 60% of patients diagnosed with cancer are alive 5 years after the diagnosis.
[0003] Most of the cancer treatments in use are surgical procedures, radiation and cytotoxic chemotherapeutics, however they all have serious side effects. Recently also treatment using antibodies directed towards known cancer associated antigens is used.
[0004] Within the last few years cancer immune therapies targeting cancer cells with the help of the patient's own immune system, i.e. cancer vaccines, have attracted interest because such therapies may reduce or even eliminate some of the side-effects seen in the traditional cancer treatment.
[0005] The foundation of immunology is based on self-nonself discrimination. Most of the pathogens inducing infectious diseases contain molecular signatures that can be recognized by the host and trigger immune responses. However tumor cells are derived from normal cells, and do not generally express any molecular signatures, making them more difficult to be distinguished from normal cells.
[0006] Nevertheless, most tumor cells express different types of tumor antigens. One class of tumor antigens are the so-called tumor associated antigens, i.e. antigens expressed at low levels in normal tissues and expressed at a much higher level in tumor tissue. Such tumorassociated antigens have been the target for cancer vaccines for the last decade. However, immunological treatment directed towards tumor associated antigens exhibit several challenges, in that the tumor cells may evade the immune system by downregulating the antigen in question, and the treatment may also lead to toxicities due to normal cell destruction.
[0007] Recently, another class of tumor antigens have been identified, the so-called tumor neoantigens or tumor specific-antigens. Tumor neoantigens arise due to one or more mutations in the tumor genome leading to a change in the amino acid sequence of the protein in question. Since these mutations are not present in normal tissue, the side-effects of the treatment directed towards the tumor associated antigens do not arise with an immunologic treatment towards tumor neoantigens.
[0008] The average number of somatic, tumor-specific non-synonymous mutations for malignant melanoma is between 100 and 120. Some of the genetic alterations can be recognized by the immune system, representing ideal antigens. Animal models have confirmed the utility of immunization with tumor neoantigens, and two clinical trials have been initiated, one with a vaccine comprising up to 10 mutated proteins and the other with an RNA vaccine (IVAC MUTANOME). The RNA vaccine comprises 2 RNA molecules each comprising five different mutation-encoding sequences.
[0009] However, by administration of either several different proteins or several RNA sequences it is difficult to control the immunological response to the various proteins administered or expressed in vivo.
[0010] Accordingly, there is a need for a more efficient vaccine ensuring expression of the mutated proteins either in vivo or in vitro and ensure delivery of the antigen as well as activation of the antigen presenting cells needed to elicit a strong T cell response.
SUMMARY OF INVENTION
[0011] The present invention relates to a therapeutic anticancer vaccine being directed to a plurality of neoepitopes from tumor neoantigens, wherein the neoepitopes are presented to the immune system as a dimeric protein called a vaccibody. WO 2004/076489 describes dimeric proteins called vaccibodies in detail.
[0012] In one embodiment the invention relates to a therapeutic anticancer neoepitope vaccine comprising an immunologically effective amount of [0013] 1) a polynucleotide comprising a nucleotide sequence encoding [0014] a targeting unit [0015] a dimerization unit [0016] a first linker [0017] an antigenic unit, wherein said antigenic unit comprises n-1 antigenic subunits, each subunit comprising at least a part of a cancer neoepitope sequence and a second linker and said antigenic unit further comprising a final cancer neoepitope sequence, wherein n is an integer of from 3 to 50. [0018] or [0019] 2) a polypeptide encoded by the polynucleotide as defined in 1), or [0020] 3) a dimeric protein consisting of two polypeptides encoded by the polynucleotide as defined in 1).
[0021] In another aspect, the invention relates to the polynucleotide as defined above. Such polynucleotide is e.g. useful in a vaccine according to the invention.
[0022] In a third aspect the invention relates to a vector comprising the polynucleotide as defined above, and in a fourth aspect the invention relates to a host cell comprising the polynucleotide or the vector as defined above.
[0023] In a fifth aspect the invention relates to a polypeptide encoded by the polynucleotide as defined above. Such polypeptide is e.g. useful in a vaccine according to the invention, and in a sixth aspect the invention relates to a dimeric protein consisting of two polypeptides as defined above.
[0024] In a seventh aspect the invention relates to the polypeptide, the dimeric protein, or the polynucleotide as defined above for use as a medicament.
[0025] As described above, in some embodiments, the vaccine comprises a polypeptide or a dimeric protein, and accordingly, in an eighth aspect the invention relates to a method for preparing a dimeric protein or an polypeptide as defined above, wherein the method comprises [0026] a) transfecting the polynucleotide as defined above into a cell population; [0027] b) culturing the cell population; [0028] c) collecting and purifying the dimeric protein, or the polypeptide expressed from the cell population.
[0029] In other embodiments, the vaccine comprises a polynucleotide, and accordingly, in a ninth aspect the invention relates to a method for preparing a vaccine, such as a DNA or RNA vaccine, comprising an immunologically effective amount of a polynucleotide, wherein said method comprises [0030] a. preparing a polynucleotide as defined above; [0031] b. mixing the polynucleotide obtained under step a) in a pharmaceutically acceptable carrier, diluent, or buffer, thereby obtaining the vaccine.
[0032] In a tenth aspect the invention relates to a method of treating cancer in a patient, the method comprising administering to the patient in need thereof, a vaccine as defined above. In an alternative tenth aspect, the invention relates to a vaccine as defined above for use in a method of treating cancer.
DESCRIPTION OF DRAWINGS
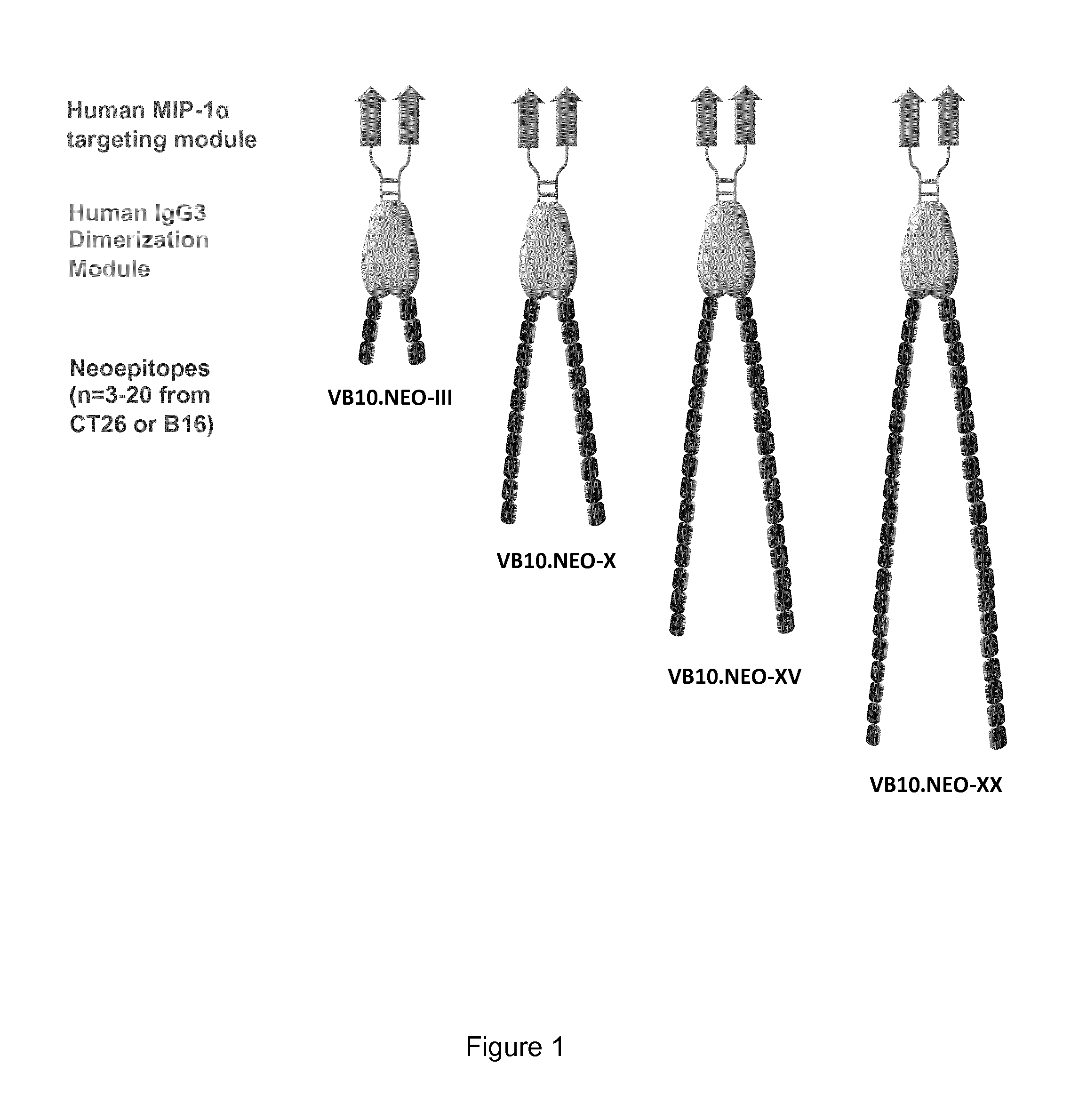
[0033] FIG. 1 shows a schematic drawing of a dimeric protein according to the invention having 3, 10 or 20 neoepitopes on each monomer, respectively.
[0034] FIG. 2 shows that neoantigen-based vaccibody proteins are produced and secreted as functional homodimers after transfection of HEK293 cells with VB10.NEO constructs. FIG. 2 upper left panels shows Western blots of VB10.NEO CT26-X (VB4001) and VB10.NEO B16-X (VB4003) comprising 10 neoepitopes and FIG. 2 lower left panels shows Western blots of VB10.NEO CT26-III (VB4002) and VB10.NEO B16-III (VB4004) comprising 3 neoepitopes. The formation of functional homodimers are shown in the left panels of the western blots for each construct (- reducing agent). The right panels illustrate the monomers (+ reducing agent). FIG. 2 right panels shows results from two ELISA experiments detecting vaccibody proteins in the supernatant from HEK293 cells transfected with the VB10.NEO constructs. Upper right panel shows the expression level of the VB10.NEO CT26 constructs, VB4001 and VB4002, and lower right panel shows the expression level of the VB10.NEO B16 constructs, VB4003 and VB4004
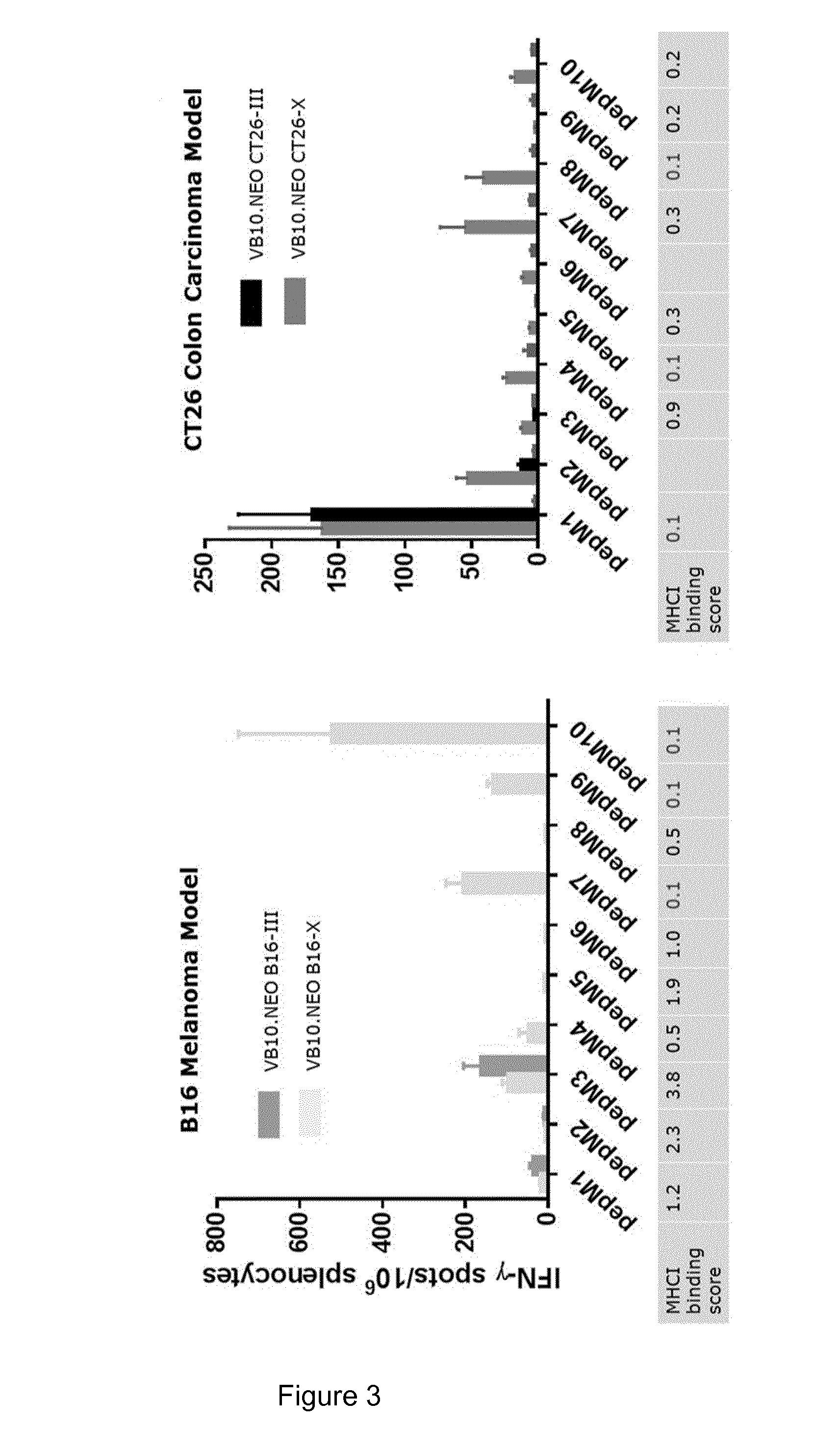
[0035] FIG. 3 illustrates that strong and broad T-cell responses are induced after a single injection with vaccibody DNA vaccines comprising 10 neoepitopes when compared to vaccibody DNA vaccines comprising 3 neoepitopes. The left panel displays IFN-.gamma. responses towards individual neoepitopes in the B16 melanoma model when injecting VB10.NEO B16-III (VB4004) or VB10.NEO B16-X (VB4003) comprising 3 and 10 neoepitopes, respectively. The right panel displays IFN-.gamma. responses towards neoepitopes in the CT26 colon carcinoma model when injecting VB10.NEO CT26-III (VB4002) or VB10.NEO CT26-X (VB4001) comprising 3 and 10 neoepitopes, respectively. The x-axis represents the 10 different neoepitopes, pepM1-M10.
[0036] VB10.NEO CT26-X=VB4001=CT26 pepM1-M10,
[0037] VB10.NEO CT26-III=VB4002=CT26 pepM1-M3,
[0038] VB10.NEO B16-X=VB4003=B16 pepM1-M10,
[0039] VB10.NEO B16-III=VB4004=B16 pepM1-M3.
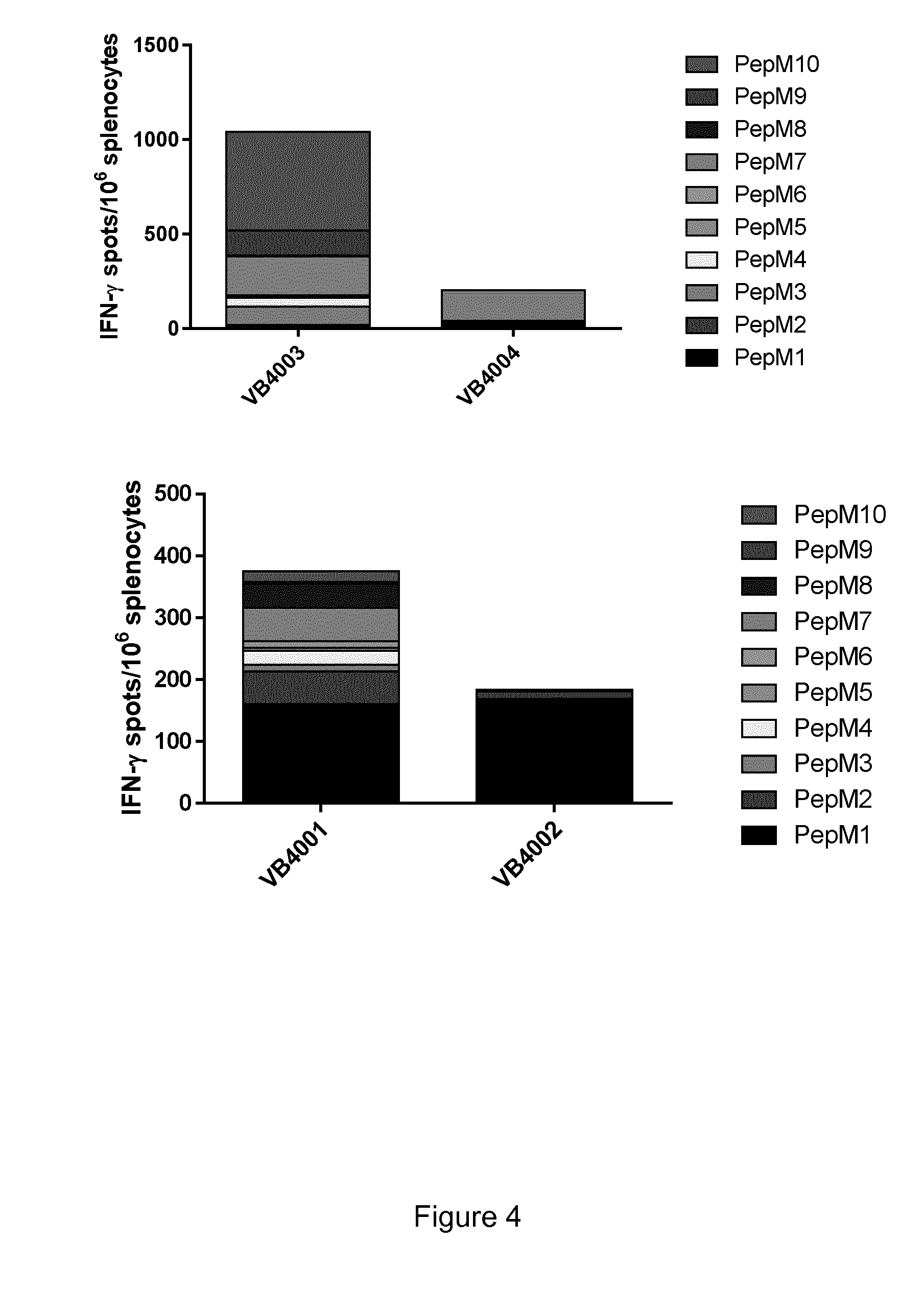
[0040] FIG. 4 illustrates that vaccibody DNA vaccines comprising 10 neoepitopes induces a stronger and broader total immune response than vaccibody DNA vaccines comprising only 3 neoepitopes. Upper panel: Comparison of the immune responses towards neoepitopes in the B16 melanoma model when injecting with VB10.NEO B16-X comprising 10 neoepitopes (VB4003) and VB10.NEO B16-III comprising 3 neoepitopes (VB4004), respectively. Lower panel: Comparison of the immune responses towards neoepitopes in the CT26 colon carcinoma model when injecting VB10.NEO CT26-X comprising 10 neoepitopes (VB4001) and VB10.NEO CT26-III comprising 3 neoepitopes (VB4002), respectively.
[0041] VB10.NEO CT26-X=VB4001=CT26 pepM1-M10,
[0042] VB10.NEO CT26-III=VB4002=CT26 pepM1-M3,
[0043] VB10.NEO B16-X=VB4003=B16 pepM1-M10,
[0044] VB10.NEO B16-III=VB4004=B16 pepM1-M3.
[0045] FIG. 5. Vaccibody DNA vaccines comprising 10 neoepitopes induce a much stronger immune response than a mix of the corresponding 10 peptides plus adjuvant. Upper panel: Comparison of the vaccibody expression level of two variants of VB10.NEO B16-X with varying order of the 10 neoepitopes (VB4003 and VB4014) in the supernatant of HEK293 cells transfected with the corresponding Vaccibody DNA constructs, detected by sandwich ELISA. In VB4003, every other neoepitope is either hydrophobic or hydrophilic, whereas in VB4014, the hydrophobic neoepitopes are placed centrally in the neoepitope antigenic module. A hydrophobic core of neoepitopes in the antigenic module may improve expression and secretion of functional vaccibody proteins in the same constructs. Lower panel: The histogram shows immune responses induced by the DNA vaccines VB10.NEO B16-X VB4003 and VB4014, and a mix of 10 peptides plus adjuvant (the same 10 neoepitopes as encoded in the VB10.NEO B16-X constructs). The order of the neoepitopes within the neoepitope antigenic module does not change the hierarchy of the immunogenicity of the individual neoepitopes.
[0046] VB10.NEO B16-X=VB4003=B16 pepM1-M10,
[0047] VB10.NEO B16-X=VB4014=B16 hydrophobic core
[0048] (pepM9+pepM5+pepM1+pepM4+pepM6+pepM8+pepM10+pepM3+pepM7+pepM2).
[0049] FIG. 6. VB10.NEO B16-X DNA vaccine where the 10 neoepitopes are spaced with 10 amino acid (aa) linkers (VB4011), induces a stronger total immune response, compared to VB10.NEO B16-X DNA vaccine where the 10 neoepitopes are spaced with 5 aa linkers (VB4003). Upper panel: Comparison of the vaccibody expression level of VB4003 and VB4011 in the supernatant of HEK293 cells transfected with the corresponding Vaccibody DNA constructs, detected by sandwich ELISA. Similar expression and secretion of functional vaccibody proteins are observed for VB4003 and VB4011. Lower panel: Histogram showing the IFN-.gamma. immune response towards neoepitopes from the B16 melanoma model in mice injected with VB4003 or VB4011. A single injection with vaccibody DNA vaccines comprising 10 neoepitopes spaced with 10 amino acid linkers resulted in the strongest total immune response. Empty vector was included as a negative control.
[0050] VB10.NEO B16-X=VB4003=B16 pepM1-M10, 5 aa linker
[0051] VB10.NEO B16-X=VB4011=B16 pepM1-M10, 10 aa linker.
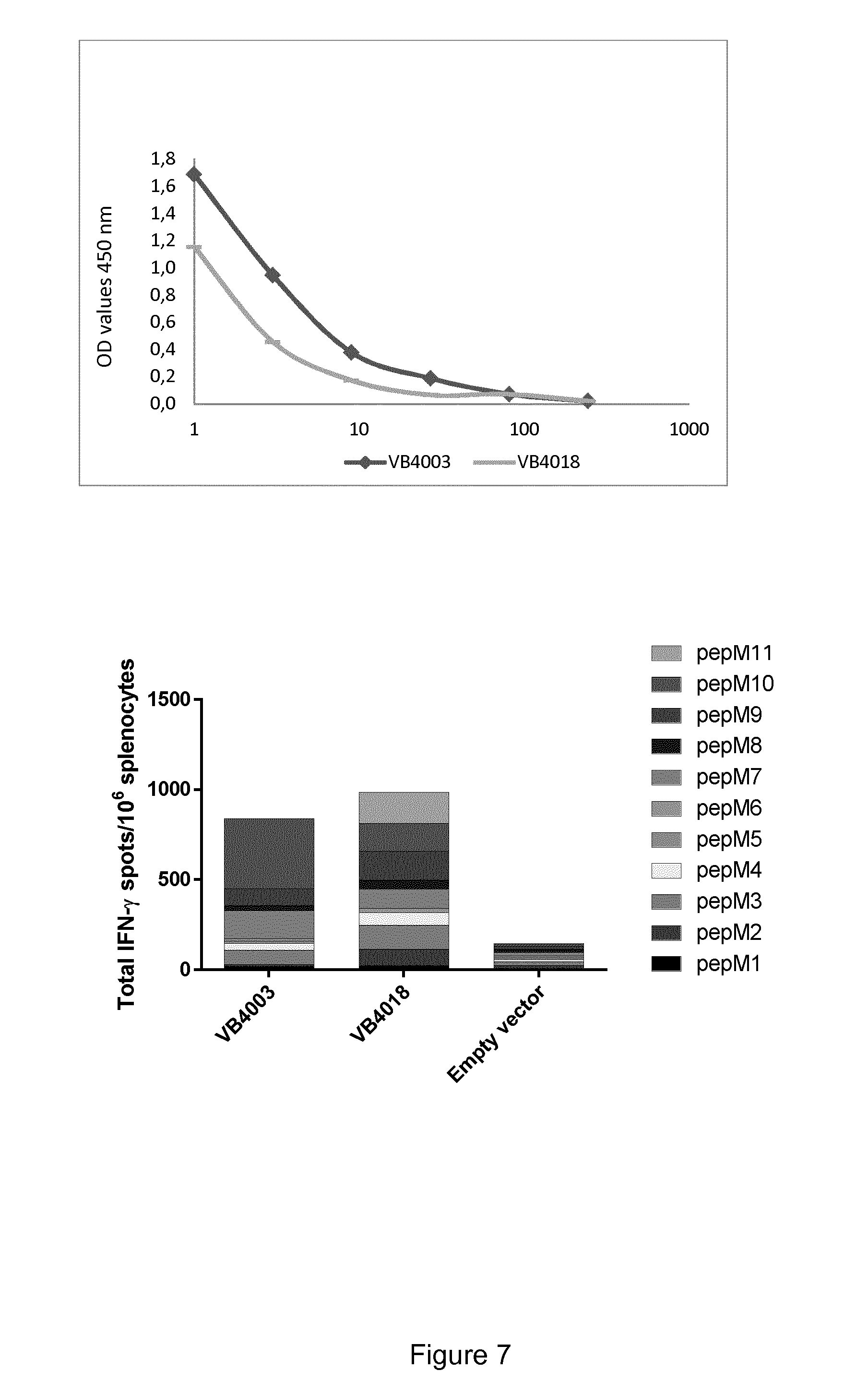
[0052] FIG. 7. Vaccibody DNA vaccine comprising 2.times.10 neoepitopes (VB4018) induces a broader immune response against individual neoepitopes compared to vaccibody DNA vaccine comprising 1.times.10 neoepitopes (VB4003). Upper panel: Comparison of vaccibody expression levels of VB10.NEO B16-X (VB4003) and VB10.NEO B16-XX (VB4018) in the supernatant of HEK293 cells transfected with the corresponding vaccibody DNA constructs, detected by sandwich ELISA. Lower panel: Histogram showing the IFN-.gamma. immune response towards neoepitopes from the B16 melanoma model in mice injected with VB4003 or VB4018. The benefit of including 2 copies of each neoepitope is limited on the total immune response, however, a broader immune response is observed towards individual neoepitopes. Empty vector is included as a negative control.
[0053] VB10.NEO B16-X=VB4003=B16 pepM1-M10, 5 aa linker
[0054] VB10.NEO B16-XX=VB4018=B16 pepM1-M4+M11+M6-M10.times.2, 5 aa linker
[0055] FIG. 8. Several copies of each neoeptiope in a vaccibody construct gives a more uniform immune response against the 5 selected best neoepitopes. Upper panel: Comparison of vaccibody expression level of VB10.NEO B16-X (VB4003 and VB4011), VB10.NEO B16-XX (VB4018), VB10.NEO B16-Vx2 (VB4019) and VB10.NEO B16-Vx4 in the supernatant of HEK293 cells transfected with the corresponding vaccibody DNA constructs, detected by sandwich ELISA. Lower panel: Histogram showing the IFN-.gamma. immune responses towards 5 neoepitopes from the B16 melanoma model (PepM3, PepM4, PepM7, PepM9 and PepM10) in mice injected with 5 different vaccibody DNA vaccines that all include these 5 neoepitopes, but in different context. Empty vector is included as a negative control. The figure illustrates that several copies of each neoepitope as observed with the vaccibody constructs VB4019 (Vx2) and VB4021 (Vx4) mediate a more evenly immune response towards the 5 shared neoepitopes compared to the decatope VB4003, where the 5 selected neoepitopes are presented once. However, the construct holding 10 different neoepitopes (i.e. just a single copy of the 5 neoepitopes tested in this assay), thus, importantly with an increased length of the linker (10 amino acids, VB4011) induced the strongest total immune response towards the 5 shared neoepitopes.
[0056] VB10.NEO B16-X=VB4003=B16 pepM1-M10, 5 aa linker
[0057] VB10.NEO B16-X=VB4011=B16 pepM1-M10, 10 aa linker
[0058] VB10.NEO B16-XX=VB4018=B16 pepM1-M4+M11+M6-M10.times.2, 5 aa linker
[0059] VB10.NEO B16-Vx2=VB4019=B16 pepM3+M4+M7+M9+M10.times.2, 5 aa linker
[0060] VB10.NEO B16-Vx4=VB4021=B16 pepM3+M4+M7+M9+M10.times.4, 5 aa linker
[0061] FIG. 9 illustrates that vaccibodiy VB4018 comprising 20 neoepitopes are expressed to the same level as vaccibody VB4017 comprising 10 neoepitopes. The vaccibody proteins are detected in the supernatant of HEK293 cells transfected with the different Vaccibody DNA constructs by sandwich ELISA.
[0062] VB10.NEO B16-X=VB4017=B16 pepM1-M4+M11+M6-M10, 5 aa linker
[0063] VB10.NEO B16-XX=VB4018=B16 pepM1-M4+M11+M6-M10.times.2, 5 aa linker
[0064] FIG. 10. Expression levels of different vaccibody constructs comprising 3-neoepitopes are compared. The vaccibody proteins are detected in the supernatant of HEK293 cells transfected with the different Vaccibody DNA constructs by sandwich ELISA Upper panel: Improved expression and secretion of functional vaccibody proteins are observed when the 3 neoepitopes are spaced with an 10 aa linker (VB4012) compared to a 5 aa linker (VB4004). Lower panel: The figure illustrates that changing the order of the neoepitopes may affect expression of the vaccibodies.
[0065] VB10.NEO B16-III=VB4004=B16 pepM1-M3, 5 aa linker
[0066] VB10.NEO B16-III=VB4012=B16 pepM1-M3, 10 aa linker
[0067] VB10.NEO B16-III=VB4015=B16 pepM1+M8+M3, 5 aa linker
[0068] VB10.NEO B16-III=VB4016=B16 pepM1+M3+M2, 5 aa linker
[0069] FIG. 11 illustrates immune responses in B16 melanoma mice that are induced after a single injection with vaccibody DNA vaccines comprising either 10 neoepitopes (VB4011), 15 neoepitopes (VB4024) or 20 neoepitopes (VB4025). Upper panel: Expression levels of the tested vaccibody constructs comprising 10-, 15- or 20 neoepitopes. The vaccibody proteins are detected in the supernatant of HEK293 cells transfected with the different Vaccibody DNA constructs by sandwich ELISA. Lower panel: Total immune response against neoepitopes in mice injected with the DNA vaccine candidates VB10.NEO B16-XV comprising 15 neoepitopes (VB4024) or VB10.NEO B16-XX comprising 20 neoepitopes (VB4025) compared to the VB10.NEO B16-X comprising 10 neoepitopes (VB4011). The figure shows the total number of IFN.gamma.-spots per 10.sup.6 splenocytes. As a negative control, mice were injected with empty vector not comprising the neoepitopes. The figure illustrates that vaccibody DNA vaccines comprising 20 neoepitopes induces a stronger and broader total immune response than vaccibody DNA vaccines comprising only 10 neoepitopes.
[0070] FIG. 12 illustrates immune responses in CT26 colon carcinoma mice that are induced after a single injection with vaccibody DNA vaccines comprising either 10 neoepitopes (VB4009), 15 neoepitopes (VB4026) or 20 neoepitopes (VB4027). Upper panel: Expression levels of the tested vaccibody construct VB10.NEO CT26-X comprising 10 neoepitopes (left panel) and vaccibody constructs VB10.NEO CT26-XV and XX comprising 15 and 20 neoepitopes, respectively (right panel). Lower panel: Total immune response towards neoepitopes in the CT26 colon carcinoma model in mice injected with the DNA vaccine candidates VB10.NEO CT26-XV comprising 15 neoepitopes (VB4026) or VB10.NEO CT26-XX comprising 20 neoepitopes (VB4027) compared to the VB10.NEO CT26-X comprising 10 neoepitopes (VB4009). The figure shows the total number of IFN.gamma.-spots per 10.sup.6 splenocytes. As a negative control, mice were injected with empty vector not comprising the neoepitopes. The figure illustrates that vaccibody DNA vaccines comprising 20 or 15 neoepitopes induces a stronger and broader total immune response than vaccibody DNA vaccines comprising only 10 neoepitopes.
[0071] NEO CT26-X=VB4009=CT26 pepM1-M10, 10 aa linker
[0072] NEO CT26-XV=VB4026=CT26 pepM1-M15, 10 aa linker
[0073] NEO CT26-XX=VB4027=CT26 pepM1-M20, 10 aa linker
[0074] FIG. 13 illustrates that mice immunized twice with VB10.NEO vaccine candidates comprising 10 neoepitopes are able to significantly delay and reduce tumour growth in the a) B16 melanoma model and b) the CT26 colon carcinoma model compared to negative control mice receiving PBS only. The figure shows the tumour volume development over time. In the CT26 colon carcinoma experiment, mice were divided into responders that were able to stabilize tumour growth and non-responders.
DEFINITIONS
[0075] Tumor is used in the present context for both a solid tumor as well as for tumor cells found in a bodily fluid, such as blood.
[0076] Tumor neoantigen is used for any tumor specific antigen comprising one or more mutations as compared to the host's exome and is used synonymously with the term cancer neoantigen.
[0077] Tumor neoepitope is used for any immunogenic mutation in a tumor antigen and is used synonymously with the term cancer neoepitope.
[0078] Tumor neoepitope sequence is used to describe the sequence comprising the neoepitope in an antigenic subunit, and is used synonymously with the term cancer neoepitope sequence.
[0079] Therapeutic anticancer vaccine is used to describe that the vaccine is used for reducing or destroying tumor cells already present in the patient.
DETAILED DESCRIPTION OF THE INVENTION
[0080] Cancers develop from the patient's normal tissue by one or a few cells starting an abnormal uncontrolled proliferation of the cells due to mutations. Although the cancer cells are mutated, most of the genome is intact and identical to the remaining cells in the patient. This is also the explanation of some of the failures in prior attempts to develop an anticancer vaccine, namely that the vaccine to some extent is also directed to the normal cells in the patient. As discussed above, the approach of attacking a tumor as defined by the present invention is to use the knowledge that any tumor, due to the mutations, expresses mutated proteins, so-called neoantigens that are not identical to any proteins in the normal cells of the patient, and therefore the neoantigens are efficient targets for a therapeutic anticancer vaccine. The mutations found in a tumor are normally highly individual, and accordingly, the vaccine according to the present invention is personalized for use only in the patient having the mutation in question.
[0081] The vaccines according to the present invention use the normal adaptive immune system to provide immunity against the tumor cells. The adaptive immune system is specific in that every foreign antigen evokes an immune response specifically towards said foreign antigen by the recognition of specific "non-self" antigens during a process called antigen presentation. The cells of the adaptive immune system are lymphocytes, in particularly B cells and T cells. B cells are involved in the humoral immune response, whereas T cells are involved in cell-mediated immune response.
[0082] In particularly, the vaccine according to the present invention is designed for evoking a cell-mediated immune response through activation of T cells against the neoantigens. T cells recognize neoepitopes when they have been processed and presented complexed to a MHC molecule as discussed below.
[0083] Major Histocompatibility Complex (MHC)
[0084] The neoepitopes according to the present invention are designed to be presented in MHC-neoepitope complexes. There are two primary classes of major histocompatibility complex (MHC) molecules, MHC I and MHC II.
[0085] MHC I is found on the cell surface of all nucleated cells in the body. One function of MHC I is to display peptides of non-self proteins from within the cell to cytotoxic T cells. The MHC I complex-peptide complex is inserted into the plasma membrane of the cell presenting the peptide to the cytotoxic T cells, whereby an activation of cytotoxic T cells against the particular MHC-peptide complex is triggered. The peptide is positioned in a groove in the MHC I molecule, allowing the peptide to be about 8-10 amino acids long.
[0086] MHC II molecules are a family of molecules normally found only on antigen-presenting cells such as dendritic cells, mononuclear phagocytes, some endothelial cells, thymic epithelial cells, and B cells.
[0087] As opposed to MHC I, the antigens presented by class II peptides are derived from extracellular proteins. Extracellular proteins are endocytosed, digested in lysosomes, and the resulting antigenic peptides are loaded onto MHC class II molecules and then presented at the cell surface. The antigen-binding groove of MHC class II molecules is open at both ends and is able to present longer peptides, generally between 15 and 24 amino acid residues long.
[0088] Class I MHC molecules are recognized by CD8 and co-receptors on the T cells, normally called CD8+ cells, whereas class II MHC molecules are recognized by CD4 and co-receptors on the T cells, normally called CD4+ cells.
[0089] Vaccines
[0090] The neoantigen vaccines of the present invention comprise a polynucleotide encoding a polypeptide comprising three units, i.e. a targeting unit, a dimerization unit and an antigenic unit. Due to the dimerization unit the polypeptide forms a dimeric protein called a vaccibody.
[0091] The genes encoding the three units are genetically engineered to be expressed as one gene. When expressed in vivo, the polypeptides/dimeric proteins target antigen presenting cells (APCs), which results in enhanced vaccine potency compared to identical non-targeted antigens.
[0092] The present invention relates to vaccines where the antigenic unit comprises antigenic subunits, wherein each subunit comprises a cancer neoepitope sequence or at least a part of a cancer neoepitope sequence. The neoepitope sequence is obtained by sequencing tumor DNA or RNA and identifying tumor specific mutations representing neoantigens. Thereby, a personalized neoantigen vaccine is obtained that specifically targets the identified tumor antigens.
[0093] One aspect of the present invention relates to a therapeutic anticancer neoepitope vaccine comprising an immunologically effective amount of [0094] a polynucleotide comprising a nucleotide sequence encoding [0095] a targeting unit [0096] a dimerization unit [0097] a first linker [0098] an antigenic unit, wherein said antigenic unit comprises n-1 antigenic subunits, each subunit comprising at least a part of a cancer neoepitope sequence and a second linker and said antigenic unit further comprising a final cancer neoepitope sequence, wherein n is an integer of from 3 to 50. [0099] or [0100] a polypeptide encoded by the polynucleotide as defined in 1), or [0101] a dimeric protein consisting of two polypeptides encoded by the polynucleotide as defined in 1).
[0102] Thus, the vaccine comprises n neoepitopes or neoepitope sequences and n-1 second linkers, wherein n is an integer from 3 to 50.
[0103] Antigenic Unit
[0104] The antigenic unit according to the invention comprises a plurality of tumor neoepitopes, wherein each neoepitope corresponds to a mutation identified in a tumor neoantigen. The mutation may be any mutation leading to a change in at least one amino acid. Accordingly, the mutation may be one of the following: [0105] a non-synonymous mutation leading to a change in the amino acid [0106] a mutation leading to a frame shift and thereby a completely different open reading frame in the direction after the mutation [0107] a read-through mutation in which a stop codon is modified or deleted leading to a longer protein with a tumor-specific neoepitope [0108] splice mutations that lead to a unique tumor-specific protein sequence [0109] chromosomal rearrangements that give rise to a chimeric protein with a tumor-specific neoepitope at the junction of the two proteins
[0110] In the antigenic unit, all but the last of the tumor neoepitopes are arranged in antigenic subunits, wherein each subunit consists of a tumor neoepitope sequence and a second linker, whereas the last subunit comprises a neoepitope only, i.e. no such second linker. Due to the separation of the tumor neoepitope sequences by said second linker, each neoepitope is presented in an optimal way to the immune system, whereby the efficiency of the vaccine is ensured as discussed below.
[0111] The cancer neoepitope sequence preferably has a length suitable for presentation by the MHC molecules discussed above. Thus, in a preferred embodiment the cancer neoepitope is from 7 to 30 amino acids long. More preferred are cancer neoepitope sequences having a length of from 7 to 10 amino acids or cancer neoepitope sequences having a length of from 13 to 30 amino acids.
[0112] In order to avoid that tumors escape the immune system by shutting down expression of a mutated gene if the vaccine is directed towards the expression product of said gene, it is preferred to include a plurality of different neoepitopes into the antigenic unit. In general the more genes the tumor has to shut down the less likely is it that the tumor is capable of shutting down all of them and still be able to proliferate or even survive. Furthermore, the tumor may be heterogeneous in that not each and every neoantigen is expressed by all the tumor cells. Accordingly, in accordance with the present invention, the approach is to include as many neoepitopes as possible into the vaccine in order to attack the tumor efficiently. Also, in order to secure that all neoepitopes are loaded efficiently to the same antigen presenting cell they are arranged as one amino acid chain instead of as discrete peptides. However, as described above, the object of the vaccine is to activate the T cells against the neoepitopes, and the T cells may be diluted in case too many neoepitopes are included into the vaccine, and therefore it is a balance to provide the vaccine with an optimal number of neoepitopes in the antigenic unit.
[0113] As discussed below in more details, the tumor exome is analysed to identify neoantigens and subsequently the most antigenic neoepitopes are selected. The present inventor has found that at least 3 neoepitopes should be selected to be incorporated into the vaccine, such as at least 5 neoepitopes, such as at least 7 neoepitopes, such as at least 10 neoepitopes, in order to efficiently be able to "hit" substantially all tumor cells.
[0114] In addition, the inventors of the present invention have found that increasing the numbers of neoepitopes in the vaccine constructs from 3 neoepitopes to 10 neoepitopes leads to a surprising increase in the immune response (see FIG. 4). In addition, it has been found that increasing the number of neoepitopes in the vaccine constructs from 10 neoepitopes to 15 or 20 neoepitopes leads to a further increase in the immune response (see FIGS. 11 and 12).
[0115] Thus, in a preferred embodiment the vaccine according to the present invention comprises at least 10 neoepitopes. In another preferred embodiment the vaccine according to the present invention comprises at least 15 neoepitopes, such as at least 20 neoepitopes.
[0116] In one embodiment from 3 to 50 neoepitopes are included in the vaccine in order to obtain the most efficient immune response without diluting the T cells, such as from 3 to 30 neoepitopes, such as from 3 to 20 neoepitopes, such as from 3 to 15 neoepitopes, such as from 3 to 10 neoepitopes, and consequently n is preferably an integer of from 3 to 50, such as from 3 to 30, such as from 5 to 25, such as from 3 to 20, such as from 3 to 15, such as from 3 to 10.
[0117] In another embodiment 5 to 50 neoepitopes may be included in the vaccine in order to obtain the most efficient immune response without diluting the T cells, such as from 5 to 30 neoepitopes, such as for example from 5 to 25 neoepitopes, such as from 5 to 20 neoepitopes, such as from 5 to 15 neoepitopes, such as from 5 to 10 neoepitopes, and consequently n is preferably an integer of from 5 to 50, such as from 5 to 30, such as from 5 to 25, such as from 5 to 20, such as from 5 to 15, such as from 5 to 10.
[0118] In a further embodiment 10 to 50 neoepitopes may be included in the vaccine in order to obtain the most efficient immune response without diluting the T cells, such as from 10 to 40 neoepitopes, such as from 10 to 30 neoepitopes, such as from 10 to 25 neoepitopes, such as from 10 to 20 neoepitopes, such as from 10 to 15 neoepitopes, and consequently n is preferably an integer of from 10 to 50, such as from 10 to 30, such as from 10 to 20, such as from 10 to 15 neoepitopes.
[0119] The inventors of the present invention have shown that vaccibody DNA vaccines comprising 10 neoepitopes induces a stronger and broader total immune response than vaccibody DNA vaccines comprising only 3 neoepitopes (see FIG. 4 and Example 2). Further, increasing the number of neoepitopes to more than 20 may result in a less efficient vaccine due to a dilution of the T cells. Further, it can be associated with technical difficulties to include more than 20 neoepitopes.
[0120] Accordingly, in a preferred embodiment of the present invention the vaccine comprises from 10 to 20 neoepitopes.
[0121] In yet another embodiment 15 to 50 neoepitopes are included in the vaccine in order to obtain the most efficient immune response without diluting the T cells, such as from 15 to 30 neoepitopes or such as from 15 to 20 neoepitopes and consequently n is preferably an integer of from 15 to 50, such as from 15 to 30 or such as from 15 to 20 neoepitopes.
[0122] In one embodiment, the antigenic unit comprises one copy of each cancer neoepitope, so that when 10 neoepitopes are included in the vaccine a cell-mediated immune response against 10 different neoepitopes can be evoked.
[0123] If however only a few relevant antigenic mutations are identified, then the antigenic unit may comprise at least two copies of at least one neoepitope in order to strengthen the immune response to these neoepitopes. Also for manufacturing and regulatory reasons it may be an advantage to keep the length of plasmid and i.e. the antigenic unit constant, and therefore it may be advantageously to include more than one copy of the same neoepitope in the antigenic unit.
[0124] As discussed above, it may be an advantage to keep the length of the antigenic unit constant, and therefore it is preferred in one embodiment that all the cancer neoepitope sequences have identical length. However, if one or more of the neoepitopes result from a mutation leading to a frame shift or stop codon mutation, the neoepitope may have a substantial length, such as consisting of at least the mutated part of the protein, the most antigenic portion of the mutated protein or maybe of the whole mutated protein, whereby the length of at least one of the neoepitopes is substantially longer than the neoepitopes arising from a non-synonymous point mutation.
[0125] The length of the antigenic unit is primarily determined by the length of the neoepitopes and the number of neoepitopes arranged in the antigenic unit and is from about 21 to 1500, preferably from about 30 amino acids to about a 1000 amino acids, more preferably from about 50 to about 500 amino acids, such as from about 100 to about 400 amino acids, from about 100 to about 300 amino acids.
[0126] In particularly when the neoepitope is short, such as a few amino acids long, the cancer neoepitope sequence comprises the neoepitope flanked at both sides by an amino acid sequence. Preferably, the neoepitope is positioned essentially in the middle of a cancer neoepitope sequence, in order to ensure that the neoepitope is presented by the antigen presenting cells after processing. The amino acid sequences flanking the neoepitope are preferably the amino acid sequences flanking the neoepitope in the neoantigen, whereby the cancer neoepitope sequence is a true subsequence of the cancer neoantigen amino acid sequence.
[0127] Although it is possible to obtain a relevant immune response towards the tumor if the neoepitopes are randomly arranged in the antigenic subunit, it is preferred to follow at least one of the following methods for ordering the neoepitopes in the antigenic unit in order to enhance the immune response.
[0128] In one embodiment, depending on the selected neoepitopes, the antigenic subunits are arranged in the order of more antigenic to less antigenic in the direction from the first linker towards the final neoepitope.
[0129] In another embodiment, in particularly if the hydrophilicity/hydrophobicity varies greatly among the neoepitopes, it is preferred that the most hydrophobic antigenic subunit(s) is/are substantially positioned in the middle of the antigenic unit and the most hydrophilic antigenic subunit(s) is/are positioned at the beginning and/or end of the antigenic unit. Alternatively, the neoepitopes may be arranged alternating between a hydrophilic and a hydrophobic neoepitope. Furthermore, GC rich neoepitopes should be spaced so that GC clusters are avoided, preferably GC rich neoepitopes are spaced by at least one subunit.
[0130] The second linker is designed to be non-immunogenic and is preferably also a flexible linker, whereby the tumor neoepitopes, in spite of the high numbers of antigenic subunits present in the antigenic unit, are presented in an optimal manner to the T cells. Preferably, the length of the second linker is from 4 to 20 amino acids to secure the flexibility. In another preferred embodiment, the length of the second linker is from 8 to 20 amino acids, such as from 8 to 15 amino acids, for example 8 to 12 amino acids or such as for example from 10 to 15 amino acids. In a particular embodiment, the length of the second linker is 10 amino acids.
[0131] In a specific embodiment, the vaccine of the present invention comprises 10 neoepitopes, wherein the second linkers have a length of from 8 to 20 amino acids, such as from 8 to 15 amino acids, for example 8 to 12 amino acids or such as for example from 10 to 15 amino acids. In a particular embodiment, the vaccine of the present invention comprises 10 neoepitopes and wherein the second linkers have a length of 10 amino acids.
[0132] The second linker is preferably identical in all antigenic subunits. If, however, one or more of the neoepitopes comprise an amino acid motif similar to the linker, it may be an advantage to substitute the neighbouring second linkers with a second linker of a different sequence. Also, if a neoepitope-second linker junction is predicted to constitute an epitope in itself, then a second linker of a different sequence might be used.
[0133] The second linker is preferably a serine-glycine linker, such as a flexible GGGGS linker, such as GGGSS, GGGSG, GGGGS or multiple variants thereof such as GGGGSGGGGS or (GGGGS).sub.m, (GGGSS).sub.m, (GGGSG).sub.m, where m is an integer from 1 to 5, from 1 to 4 or from 1 to 3. In a preferred embodiment m is 2.
[0134] In a preferred embodiment the serine-glycine linker further comprises at least one leucine (L), such as at least 2 or at least 3 leucines. The serine-glycine linker may for example comprise 1, 2, 3 or 4 leucine. Preferably, the serine-glycine linker comprises 1 leucine or 2 leucines.
[0135] In one embodiment the second linker comprises or consists of the sequence LGGGS, GLGGS, GGLGS, GGGLS or GGGGL. In another embodiment the second linker comprises or consists of the sequence LGGSG, GLGSG, GGLSG, GGGLG or GGGSL. In yet another embodiment the second linker comprises or consists of the sequence LGGSS, GLGSS, GGLSS, GGGLS or GGGSL.
[0136] In yet another embodiment the second linker comprises or consists of the sequence LGLGS, GLGLS, GLLGS, LGGLS or GLGGL. In another embodiment the second linker comprises or consists of the sequence LGLSG, GLLSG, GGLSL, GGLLG or GLGSL. In yet another embodiment the second linker comprises or consists of the sequence LGLSS, GLGLS, GGLLS, GLGSL or GLGSL.
[0137] In another embodiment of the present invention the second serine-glycine linker has a length of 10 amino acids and comprises 1 leucine or 2 leucines.
[0138] In one embodiment the second linker comprises or consists of the sequence LGGGSGGGGS, GLGGSGGGGS, GGLGSGGGGS, GGGLSGGGGS or GGGGLGGGGS. In another embodiment the second linker comprises or consists of the sequence LGGSG GGGSG, GLGSGGGGSG, GGLSGGGGSG, GGGLGGGGSG or GGGSLGGGSG. In yet another embodiment the second linker comprises or consists of the sequence LGGSSGGGSS, GLGSSGGGSS, GGLSSGGGSS, GGGLSGGGSS or GGGSLGGGSS.
[0139] In a further embodiment the second linker comprises or consists of the sequence LGGGSLGGGS, GLGGSGLGGS, GGLGSGGLGS, GGGLSGGGLS or GGGGLGGGGL. In another embodiment the second linker comprises or consists of the sequence LGGSGLGGSG, GLGSGGLGSG, GGLSGGGLSG, GGGLGGGGLG or GGGSLGGGSL. In yet another embodiment the second linker comprises or consists of the sequence LGGSSLGGSS, GLGSSGLGSS, GGLSSGGLSS, GGGLSGGGLS or GGGSLGGGSL.
[0140] In a preferred embodiment the vaccine according to the present invention comprises at least 10 neoepitopes that are separated by 10 amino acid linkers. In another preferred embodiment the vaccine according to the present invention comprises at least 15 neoepitopes that are separated by 10 amino acid linkers, such as at least 20 neoepitopes that are separated by 10 amino acid linkers.
[0141] In another preferred embodiment the vaccine comprises from 10 to 20 or from 10 to 25 neoepitopes that are separated by second linkers. Preferably, said second linkers are 10 amino acids. The second linker may also have any length as defined herein above, such as for example from 8 to 12 amino acids.
[0142] Alternative linkers may be selected from the group consisting of GSAT linkers and SEG linkers, or multiple variants thereof.
[0143] Targeting Unit
[0144] Due to the targeting unit, the polypeptide/dimeric protein of the invention leads to attraction of dendritic cells (DCs), neutrophils and other immune cells. Thus, the polypeptide/dimeric protein comprising the targeting module will not only target the antigens to specific cells, but in addition facilitate a response-amplifying effect (adjuvant effect) by recruiting specific immune cells to the administration site of the vaccine. This unique mechanism is of great importance in a clinical setting where patients can receive the vaccine without any additional adjuvants since the vaccine itself gives the adjuvant effect.
[0145] The term "targeting unit" as used herein refers to a unit that delivers the polypeptide/protein with its antigen to an antigen presenting cell for MHC class II-restricted presentation to CD4+ T cells or for providing cross presentation to CD8+ T cells by MHC class I restriction.
[0146] The targeting unit is connected through the dimerization unit to the antigenic unit, wherein the latter is in either the COOH-terminal or the NH2-terminal end of the polypeptide/dimeric protein.
[0147] It is preferred that the antigenic unit is in the COOH-terminal end of the polypeptide/dimeric protein.
[0148] The targeting unit is designed to target the polypeptide/dimeric protein of the invention to surface molecules expressed on the relevant antigen presenting cells, such as molecules expressed exclusively on subsets of dendritic cells (DC).
[0149] Examples of such target surface molecules on APC are human leukocyte antigen (HLA), cluster of differentiation 14 (CD14), cluster of differentiation 40 (CD40), chemokine receptors and Toll-like receptors (TLRs). HLA is a major histocompatibility complex (MHC) in humans. The Toll-like receptors may for example include TLR-2, TLR-4 and/or TLR-5.
[0150] The polypeptide/dimeric protein of the invention can be targeted to said surface molecules by means of targeting units comprising for example antibody binding regions with specificity for CD14, CD40, or Toll-like receptor; ligands, e.g. soluble CD40 ligand; natural ligands like chemokines, e.g. RANTES or MIP-1a; or bacterial antigens like for example flagellin.
[0151] In one embodiment the targeting unit has affinity for an MHC class II protein. Thus, in one embodiment the nucleotide sequence encoding the targeting unit encodes an the antibody variable domains (VL and VH) with specificity for MHC class II proteins, selected from the group consisting of anti-HLA-DP, anti-HLA-DR and anti-HLA-II.
[0152] In another embodiment the targeting unit has affinity for a surface molecule selected from the group consisting of CD40, TLR-2, TLR-4 and TLR-5, Thus, in one embodiment the nucleotide sequence encoding the targeting unit encodes the antibody variable domains (VL and VH) with specificity for anti-CD40, anti-TLR-2, anti-TLR-4 and anti-TLR-5. In one embodiment the nucleotide sequence encoding the targeting unit encodes Flagellin. Flagellin has affinity for TLR-5.
[0153] Preferably, the targeting unit has affinity for a chemokine receptor selected from CCR1, CCR3 and CCR5. More preferably, the nucleotide sequence encoding the targeting unit encodes the chemokine hMIP-1alpha (LD78beta), which binds to its cognate receptors, CCR1, CCR3 and CCR5 expressed on the cell surface of APCs.
[0154] The binding of the polypeptide/dimeric protein of the invention to its cognate receptors leads to internalization in the APC and degradation of the proteins into small peptides that are loaded onto MHC molecules and presented to CD4+ and CD8+ T cells to induce tumor specific immune responses. Once stimulated and with help from activated CD4+ T cells, CD8+ T cells will target and kill tumor cells expressing the same neoantigens.
[0155] In one embodiment of the present invention, the targeting unit comprises an amino acid sequence having at least 80% sequence identity to the amino acid sequence 24-93 of SEQ ID NO:1. In a preferred embodiment, the targeting unit comprises an amino acid sequence having at least 85% sequence identity to the amino acid sequence 24-93 of SEQ ID NO:1, such as at least 86%, such as at least 87%, such as at least 88%, such as at least 89%, such as at least 90%, such as at least 91%, such as at least 92%, such as at least 93%, such as at least 94%, such as at least 95%, such as at least 96%, such as at least 97%, such as at least 98%, such as at least 99% sequence identity.
[0156] In a more preferred embodiment the targeting unit consists of an amino acid sequence having at least 80% sequence identity to the amino acid sequence 24-93 of SEQ ID NO:1, such as at least 85%, such as at least 86%, such as at least 87%, such as at least 88%, such as at least 89%, such as at least 90%, such as at least 91%, such as at least 92%, such as at least 93%, such as at least 94%, such as at least 95%, such as at least 96%, such as at least 97%, such as at least 98%, such as at least 99%, such as at least 100% sequence identity to the amino acid sequence 24-93 of SEQ ID NO:1.
[0157] Dimerization Unit
[0158] The term "dimerization unit" as used herein, refers to a sequence of amino acids between the antigenic unit and the targeting unit. Thus, the dimerization unit serves to connect the antigenic unit and the targeting unit, and facilitates dimerization of two monomeric polypeptides into a dimeric protein. Furthermore, the dimerization unit also provides the flexibility in the polpeptide/dimeric protein to allow optimal binding of the targeting unit to the surface molecules on the antigen presenting cells (APCs), even if they are located at variable distances. The dimerization unit may be any unit that fulfils these requirements.
[0159] Accordingly, in one embodiment the dimerization unit may comprise a hinge region and optionally another domain that facilitates dimerization, and the hinge region and the other domain may be connected through a third linker.
[0160] The term "hinge region" refers to a peptide sequence of the dimeric protein that facilitates the dimerization. The hinge region functions as a flexible spacer between the units allowing the two targeting units to bind simultaneously to two target molecules on APCs, even if they are expressed with variable distances. The hinge region may be Ig derived, such as derived from IgG3. The hinge region may contribute to the dimerization through the formation of covalent bond(s), e.g. disulfide bridge(s). Thus, in one embodiment the hinge region has the ability to form one or more covalent bonds. The covalent bond can for example be a disulfide bridge.
[0161] In one embodiment, the other domain that facilitates dimerization is an immunoglobulin domain, such as a carboxyterminal C domain, or a sequence that is substantially identical to the C domain or a variant thereof. Preferably, the other domain that facilitates dimerization is a carboxyterminal C domain derived from IgG.
[0162] The immunoglobulin domain contributes to dimerization through non-covalent interactions, e.g. hydrophobic interactions. For example, the immunoglobulin domain has the ability to form dimers via noncovalent interactions. Preferably, the noncovalent interactions are hydrophobic interactions.
[0163] It is preferred that the dimerization unit does not comprise a CH2 domain.
[0164] In a preferred embodiment, the dimerization unit consists of hinge exons h1 and h4 connected through a third linker to a CH3 domain of human IgG3.
[0165] In one embodiment of the present invention, the dimerization unit comprises an amino acid sequence having at least 80% sequence identity to the amino acid sequence 94-237 of SEQ ID NO:3. In a preferred embodiment, the dimerization unit comprises an amino acid sequence having at least 85% sequence identity to the amino acid sequence 94-237 of SEQ ID NO:3, such as at least 86%, such as at least 87%, such as at least 88%, such as at least 89%, such as at least 90%, such as at least 91%, such as at least 92%, such as at least 93%, such as at least 94%, such as at least 95%, such as at least 96%, such as at least 97%, such as at least 98%, such as at least 99% sequence identity.
[0166] In a more preferred embodiment the dimerization unit consists of an amino acid sequence having at least 80% sequence identity to the amino acid sequence 94-237 of SEQ ID NO:3, such as at least 85%, such as at least 86%, such as at least 87%, such as at least 88%, such as at least 89%, such as at least 90%, such as at least 91%, such as at least 92%, such as at least 93%, such as at least 94%, such as at least 95%, such as at least 96%, such as at least 97%, such as at least 98%, such as at least 99%, such as at least 100% sequence identity to the amino acid sequence 94-237 of SEQ ID NO:3.
[0167] In one embodiment the third linker is a G3S2G3SG linker.
[0168] It is to be understood that the dimerization unit may have any orientation with respect to antigenic unit and targeting unit. In one embodiment, the antigenic unit is in the COOH-- terminal end of the dimerization unit with the targeting unit in the N-terminal end of the dimerization unit.
[0169] In another embodiment, the antigenic unit is in the N-terminal end of the dimerization unit with the targeting unit in the COOH-terminal end of the dimerization unit. It is preferred that the antigenic unit is in the COOH end of the dimerization unit.
[0170] First Linker
[0171] The antigenic unit and the dimerization unit are preferably connected through a first linker. The first linker may comprise a restriction site in order to facilitate the construction of the polynucleotide. It is preferred that the first linker is a GLGGL linker or a GLSGL linker.
[0172] Signal Peptide
[0173] In a preferred embodiment, the polynucleotide further comprises a nucleotide sequence encoding a signal peptide. The signal peptide is constructed to allow secretion of the polypeptide encoded by the polynucleotide of the invention in the cells transfected with said polynucleotide.
[0174] Any suitable signal peptide may be used. Examples of suitable peptides are an Ig VH signal peptide, such as SEQ ID NO: 31, a human TPA signal peptide, such as SEQ ID NO: 32, and a signal peptide comprising an amino acid sequence having at least 80% sequence identity to the amino acid sequence 1-23 of SEQ ID NO:1.
[0175] In a preferred embodiment, the signal peptide comprises an amino acid sequence having at least 85%, such as at least 86%, such as at least 87%, such as at least 88%, such as at least 89%, such as at least 90%, such as at least 91%, such as at least 92%, such as at least 93%, such as at least 94%, such as at least 95%, such as at least 96%, such as at least 97%, such as at least 98%, such as at least 99%, such as 100% sequence identity to the amino acid sequence 1-23 of SEQ ID NO:1.
[0176] In a more preferred embodiment, the signal peptide consists of an amino acid sequence having at least 80%, preferably at least 85%, such as at least 86%, such as at least 87%, such as at least 88%, such as at least 89%, such as at least 90%, such as at least 91%, such as at least 92%, such as at least 93%, such as at least 94%, such as at least 95%, such as at least 96%, such as at least 97%, such as at least 98%, such as at least 99%, such as 100% sequence identity to the amino acid sequence 1-23 of SEQ ID NO:1.
[0177] Sequence Identity
[0178] Sequence identity may be determined as follows: A high level of sequence identity indicates likelihood that the first sequence is derived from the second sequence. Amino acid sequence identity requires identical amino acid sequences between two aligned sequences. Thus, a candidate sequence sharing 70% amino acid identity with a reference sequence requires that, following alignment, 70% of the amino acids in the candidate sequence are identical to the corresponding amino acids in the reference sequence. Identity may be determined by aid of computer analysis, such as, without limitations, the ClustalW computer alignment program (Higgins D., Thompson J., Gibson T., Thompson J. D., Higgins D. G., Gibson T. J., 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680), and the default parameters suggested therein. Using this program with its default settings, the mature (bioactive) part of a query and a reference polypeptide are aligned. The number of fully conserved residues is counted and divided by the length of the reference polypeptide. In doing so, any tags or fusion protein sequences, which form part of the query sequence, are disregarded in the alignment and subsequent determination of sequence identity.
[0179] The ClustalW algorithm may similarly be used to align nucleotide sequences. Sequence identities may be calculated in a similar way as indicated for amino acid sequences.
[0180] Another preferred, non-limiting example of a mathematical algorithm utilized for the comparison of sequences is the algorithm of Myers and Miller, CABIOS (1989). Such an algorithm is incorporated into the ALIGN program (version 2.0) which is part of the FASTA sequence alignment software package (Pearson W R, Methods Mol Biol, 2000, 132:185-219). Align calculates sequence identities based on a global alignment. Align0 does not penalise to gaps in the end of the sequences. When utilizing the ALIGN og Align0 program for comparing amino acid sequences, a BLOSUM50 substitution matrix with gap opening/extension penalties of -12/-2 is preferably used.
[0181] Polynucleotides
[0182] The invention also relates to a polynucleotide as described above. The polynucleotide may comprise a DNA nucleotide sequence or a RNA nucleotide sequence, such as genomic DNA, cDNA, and RNA sequences, either double stranded or single stranded.
[0183] It is preferred that the polynucleotide is optimized to the species to express the polypeptide according to the invention, i.e. it is preferred that the polynucleotide sequence is human codon optimized.
[0184] Polypeptides and Dimeric Proteins
[0185] The invention further relates to a polypeptide encoded by the polynucleotide sequence as defined above. The polypeptide may be expressed in vitro for production of the vaccine according to the invention, or the polypeptide may be expressed in vivo as a result of administration of the polynucleotide as defined above.
[0186] Due to the presence of the dimerization unit, dimeric proteins are formed when the polypeptide is expressed. The dimeric protein may be a homodimer, i.e. wherein the two polypeptide chains are identical and consequently comprise identical neoepitopes, or the dimeric protein may be a heterodimer comprising two different monomeric polypeptides encoded in the antigenic units. The latter may be relevant if the amount of neoepitopes exceeds an upper size limit for the antigenic unit. It is however preferred that the dimeric protein is a homodimeric protein.
[0187] Vector
[0188] Furthermore, the invention relates to a vector comprising a nucleotide sequence as defined above. It is preferred that the vector allows for easy exchange of the various units described above, in particularly the antigenic unit. In particularly, the expression vector may be pUMVC4a vector or NTC9385R vector backbones. The antigenic unit may be exchanged with an antigenic unit cassette restricted by the Sfil restriction enzyme cassette where the 5' site is incorporated in the GLGGL/GLSGL linker and the 3' site is included after the stop codon in the vector.
[0189] Host Cell
[0190] The invention also relates to a host cell comprising a nucleotide sequence as defined above or comprising a vector as defined above for expression of the polypeptide according to the invention.
[0191] Suitable host cells include prokaryotes, yeast, insect or higher eukaryotic cells.
[0192] Methods for Preparing the Vaccine
[0193] The vaccine according to the invention is preferably a personalized vaccine in the sense that the neoantigens are identified in the patient's tumor and accordingly, the vaccine is directed exactly against the specific mutated proteins in the patient's tumor.
[0194] Accordingly, in one aspect the invention relates to a method for preparing a vaccine comprising an immunologically effective amount of the dimeric protein, or the polypeptide as defined above by producing the polypeptides in vitro. The in vitro synthesis of the polypeptides and proteins may be carried out by any suitable method known to the person skilled in the art, such a through peptide synthesis or expression of the polypeptide in any of a variety of expressions systems followed by purification. Accordingly, in one embodiment the method comprises [0195] a) transfecting the polynucleotide as defined above into a cell population; [0196] b) culturing the cell population; [0197] c) collecting and purifying the dimeric protein, or the polypeptide expressed from the cell population, and [0198] d) mixing the dimeric protein or polypeptide obtained under step c) with a pharmaceutically acceptable carrier, thereby obtaining the vaccine.
[0199] In a preferred embodiment, the dimeric protein or polypeptide obtained under step c) is dissolved in said pharmaceutically acceptable carrier.
[0200] Furthermore, an adjuvant or buffer may be added to the vaccine.
[0201] Purification may be carried out according to any suitable method, such as chromatography, centrifugation, or differential solubility.
[0202] In another aspect the invention relates to a method for preparing a vaccine comprising an immunologically effective amount of the polynucleotide as defined above. In one embodiment the method comprises [0203] a. preparing the polynucleotide as defined above; [0204] b. mixing the polynucleotide obtained under step a) with a pharmaceutically acceptable carrier thereby obtaining the vaccine.
[0205] The polynucleotide may be prepared by any suitable method known to the skilled person. For example, the polynucleotide may be prepared by chemical synthesis using an oligonucleotide synthesizer.
[0206] In particularly, smaller nucleotide sequences, such as for example nucleotide sequences encoding the targeting unit, the dimerization unit and/or the subunits of the antigenic unit may be synthesized individually and then ligated to produce the final polynucleotide into the vector backbone.
[0207] For the design of a personalized vaccine the methods above are preceded by a method of identifying the neoepitopes to be included into the polynucleotide.
[0208] This method preferably includes the steps of [0209] sequencing the genome, or exome of a tumor [0210] identifying tumor neoantigens comprising neoepitopes from said tumor, [0211] selecting neoepitopes based on predicted antigenicity.
[0212] The tumor or tumor part may be by through any suitable method, such as by obtaining a biopsy of the tumor or by excision of the tumor, or from any suitable body fluid, such as a blood sample or a urine sample.
[0213] Sequencing of Tumor Genome or Exome
[0214] The genome or the exome, i.e. the coding part of the genome, may be sequenced using any suitable method, such as whole exome sequencing. In particularly the sequencer may be an Illumina HiSeq2500), using Paired-end 2.times.100-125 or PE100-125 (read length), multiplex.
[0215] Identifying Tumor Antigens
[0216] Once the tumor specific mutations are identified the next step is to select predicted antigenic peptides comprising the neoepitopes.
[0217] Tumor mutations are discovered by sequencing of tumor and normal tissue and make a comparison of the obtained sequences. A variety of methods are available for detecting the presence of a particular mutation or allele in an individual's DNA or RNA. For example techniques including dynamic allele-specific hybridization (DASH), microplate array diagonal gel electrophoresis (MADGE), pyrosequencing, oligonucleotide-specific ligation, the TaqMan system as well as various DNA "chip" technologies such as the Affymetrix SNP chips may be applied.
[0218] Alternatively, a method for identifying mutations by direct protein sequencing may be carried out.
[0219] Out of the maybe hundreds or thousands of mutations in the tumor exome, the neoepitopes are selected in silico on the basis of predictive HLA-binding algorithms. The intention is to identify all relevant neoepitopes and after a ranking or scoring determine the neoepitopes to be included in the vaccine for the specific patient in question.
[0220] Any suitable algorithms may be used, such as one of the following:
[0221] Available free software analysis of peptide-MHC binding (IEDB and NetMHC) may be downloaded from the following websites: [0222] http://www.iedb.org/ [0223] http://www.cbs.dtu.dk/services/NetMHC/
[0224] Commercially available advanced software to predict optimal peptides for vaccine design are found here:
[0225] http://www.oncoimmunity.com/ [0226] https://omictools.com/t-cell-epitopes-category [0227] https://github.com/griffithlab/pVAC-Seq [0228] http://crdd.osdd.net/raghava/cancertope/help.php [0229] http://www.epivax.com/tag/neoantigen/
[0230] Each mutation is scored with respect to its antigenicity, and the most antigenic neoepitopes are selected and optimally designed in the polynucleotide. As discussed above from 3 to 50 neoepitopes are preferred according to the present invention.
[0231] Vaccine
[0232] The final vaccine is then produced to comprise one of the following: [0233] the polynucleotide as defined above [0234] the polypeptide encoded by the polynucleotide as defined above [0235] the dimeric protein comprising to polypeptide chains
[0236] The vaccine may further comprise a pharmaceutically acceptable carrier, diluent, adjuvant or buffer.
[0237] Pharmaceutically acceptable carriers, diluents, and buffers include, but are not limited to, saline, buffered saline, dextrose, water, glycerol, ethanol, sterile isotonic aqueous buffer, and combinations thereof.
[0238] In particularly for vaccines comprising polypeptides/proteins, pharmaceutically acceptable adjuvants include, but are not limited to poly-ICLC, 1018 ISS, aluminum salts, Amplivax, AS 15, BCG, CP-870,893, CpG7909, CyaA, dSLIM, GM-CSF, IC30, IC31, Imiquimod, ImuFact EV1P321, IS Patch, ISS, ISCOMATRIX, JuvImmune, LipoVac, MF59, monophosphoryl lipid A, Montanide IMS 1312, Montanide ISA 206, Montanide ISA 50V, Montanide ISA-51, OK-432, OM-174, OM-197-MP-EC, ONTAK, PepTel.RTM., vector system, PLGA microparticles, resiquimod, SRL172, Virosomes and other Virus-like particles, YF-17D, VEGF trap, R848, beta-glucan, Pam3Cys, Aquila's QS21 stimulon, vadimezan, and/or AsA404 (DMXAA).
[0239] In particularly for vaccines comprising polynucleotides the carriers may include molecules that ease transfection of cells and adjuvants may include plasmids comprising nucleotide sequences encoding chemokines or cytokines in order to enhance the immune response.
[0240] The vaccine is formulated into any suitable formulation, such as a liquid formulation for intradermal or intramuscular injection.
[0241] Administration
[0242] The vaccine may be administered in any suitable way for either a polypeptide/protein vaccine or a polynucleotide vaccine, such as administered by injection intradermally, intramuscular, subcutaneously, or by mucosal or epithelial application, such as intranasally, orally, enteral or to the bladder.
[0243] In particularly the vaccine is preferably administered intramuscular or intradermally when the vaccine is a polynucleotide vaccine.
[0244] In a specific embodiment the vaccine is administered by intranodal injection. As used herein, the term "intranodal injection" means that the vaccine is injected into the lymph nodes.
[0245] Treatment
[0246] The polynucleotides, polypeptides and dimeric proteins are preferably for use in the treatment of cancer, and formulated in a vaccine as discussed above. By the methods described herein it is possible to treat a patient suffering from cancer by examining any mutations present in the tumor in the patient, producing the vaccine and then immunizing the patient with the vaccine directed exactly to neoantigens present in his or her tumor. Due to the fast and reliable methods for sequencing, epitope-determining and producing nucleotide sequences today, it has become likely that a patient may receive the vaccine within 12 weeks from having the tumor resected
[0247] The cancer may be any cancer wherein the cancer cells comprise mutations. The cancer may be a primary tumor, metastasis or both. The tumor examined for mutations may be a primary tumor or a metastasis. The cancers to be treated are in particularly the cancers known to have a high mutational load, such as melanomas, lung cancer, breast cancer, prostate cancer or colonic cancer.
[0248] In a preferred embodiment the treatment is performed with a vaccine comprising a polynucleotide as described above, for example wherein the polynucleotide is DNA or RNA.
[0249] It is preferred to inject a polynucleotide vaccine intramuscular, such as in the big muscles, for example in the shoulder, buttock or thigh. It has been found that the polypeptides are produced locally and relevant immune cells internalize the polypeptides/proteins essentially at the site of production, and substantially no polypeptides or proteins reach the blood stream.
[0250] Any suitable method for injecting the polynucleotide may be used, such as by the use of a jet injector or assisted by electroporation.
[0251] Dosage Regimen
[0252] The vaccine may be administered as a single dosage, or may be repeated. When the vaccine administration is repeated it is preferred that it is administered with at least 3 week intervals, to avoid exhaustion of the T cells.
[0253] Accordingly, in one embodiment the dosage regimen would be vaccination week 0, 3, 6 and then every 4 weeks as long as the patient has clinical benefit. The vaccine may be administered for at least a year.
[0254] The vaccine is administered in an immunologically effective amount. By "immunologically effective amount" is meant the amount of the vaccine required to establish a tumor reducing effect. Ultimately, the physician determines the dosage that typically is in the range of 0.3-6 mg for DNA vaccines, and in the range of 5 .mu.g-5 mg for polypeptide/protein vaccines.
[0255] Combination Treatments
[0256] The vaccine treatment according to the present invention may be combined with any other anticancer treatment, such as radiation therapy, chemotherapy, and surgical treatment.
[0257] The vaccine treatment according to the invention may also be combined with checkpoint-blockade inhibitor treatment.
Specific Embodiments
[0258] 1. A therapeutic anticancer neoepitope vaccine comprising an immunologically effective amount of [0259] a polynucleotide comprising a nucleotide sequence encoding [0260] a targeting unit [0261] a dimerization unit [0262] a first linker [0263] an antigenic unit, wherein said antigenic unit comprises n-1 antigenic subunits, each subunit comprising at least a part of a cancer neoepitope sequence and a second linker and said antigenic unit further comprising a final cancer neoepitope sequence, wherein n is an integer of from 3 to 50. [0264] or [0265] a polypeptide encoded by the polynucleotide as defined in 1), or [0266] a dimeric protein consisting of two polypeptides encoded by the polynucleotide as defined in 1). [0267] 2. The vaccine according to embodiment 1, wherein the antigenic unit comprises one copy of each cancer neoepitope. [0268] 3. The vaccine according to embodiment 1, wherein the antigenic unit comprises at least two copies of at least one neoepitope. [0269] 4. The vaccine according to any of the preceding embodiments, wherein the cancer neoepitope sequence has a length of from 7 to 30 amino acids. [0270] 5. The vaccine according to embodiment 4, wherein the cancer neoepitope sequence has a length of from 7 to 10 amino acids. [0271] 6. The vaccine according to embodiment 4, wherein the cancer neoepitope sequence has a length of from 13 to 30 amino acids. [0272] 7. The vaccine according to any of the preceding embodiments, wherein each cancer neoepitope sequence has identical length. [0273] 8. The vaccine according to any of the preceding embodiments, wherein the cancer neoepitope is positioned essentially in the middle of the cancer neoepitope sequence. [0274] 9. The vaccine according to any of the preceding embodiments, wherein the cancer neoepitope sequence is a subsequence of a cancer neoantigen. [0275] 10. The vaccine according to any of the preceding embodiments, wherein the antigenic subunits are in the order of more antigenic to less antigenic from the first linker. [0276] 11. The vaccine according to any of the preceding embodiments, wherein the most hydrophobic antigenic subunit(s) is(are) substantially the middle of the antigenic unit and the most hydrophilic antigenic subunit(s) is/are at the ends of the antigenic unit. [0277] 12. The vaccine according to any of the preceding embodiments, wherein the second linker is a flexible linker. [0278] 13. The vaccine according to any of the preceding embodiments, wherein the second linker is non-immunogenic. [0279] 14. The vaccine according to any of the preceding embodiments, wherein the second linker is identical in all antigenic subunits. [0280] 15. The vaccine according to any of the preceding embodiments, wherein the second linker is a Serine-Glycine linker. [0281] 16. The vaccine according to any of the preceding embodiments, wherein the length of the second linker is from 4 to 20 amino acids. [0282] 17. The vaccine according to any of the preceding embodiments, wherein the length of the second linker is 10 amino acids. [0283] 18. The vaccine according to any of the preceding embodiments, wherein the length of the antigenic unit is from about 100 amino acids to about a 1000 amino acids. [0284] 19. The vaccine according to any of the preceding embodiments, wherein n is an integer between 3 and 30. [0285] 20. The vaccine according to any of the preceding embodiments, wherein the dimerization unit comprises a hinge region and optionally another domain that facilitates dimerization, optionally connected through a third linker. [0286] 21. The vaccine according to embodiment 20, wherein the hinge region is Ig derived. [0287] 22. The vaccine according to any one of embodiments 20-21, wherein the hinge region has the ability to form one or more covalent bonds. [0288] 23. The vaccine according to embodiment 22, wherein the covalent bond is a disulfide bridge. [0289] 24. The vaccine according to any one of embodiments 20-23, wherein the another domain that facilitates dimerization is an immunoglobulin domain, preferably a carboxyterminal C domain, or a sequence that is substantially identical to said C domain or a variant thereof. [0290] 25. The vaccine according to embodiment 24, wherein the carboxyterminal C domain is derived from IgG. [0291] 26. The vaccine according to any one of embodiments 24-25, wherein the immunoglobulin domain of the dimerization unit has the ability to homodimerize. [0292] 27. The vaccine according to any one of embodiments 24-26, wherein said immunoglobulin domain has the ability to homodimerize via noncovalent interactions. [0293] 28. The vaccine according to embodiment 27, wherein said noncovalent interactions are hydrophobic interactions. [0294] 29. The vaccine according to any one of embodiments 20-28, wherein said dimerization unit does not comprise a CH2 domain. [0295] 30. The vaccine according to any one of embodiments 20-29, wherein the dimerization unit consist of hinge exons h1 and h4 connected through said third linker to a C.sub.H3 domain of human IgG3. [0296] 31. The vaccine according to any one of embodiments 20-30, wherein the dimerization unit comprises an amino acid sequence having at least 80% sequence identity to the amino acid sequence 94-237 of SEQ ID NO:3. [0297] 32. The vaccine according to any one of embodiments 30-31, wherein said third linker is a G.sub.3S.sub.2G.sub.3SG linker. [0298] 33. The vaccine according to any of the preceding embodiments, wherein said antigenic unit and the dimerization unit is connected through said first linker. [0299] 34. The vaccine according to embodiment 33, wherein the first linker comprises a restriction site. [0300] 35. The vaccine according to embodiment 33 or 34, wherein the first linker is a GLGGL linker or a GLSGL linker. [0301] 36. The vaccine according to any of the preceding embodiments, wherein the targeting unit has affinity for a chemokine receptor selected from CCR1, CCR3 and CCR5. [0302] 37. The vaccine according to any of the preceding embodiments, wherein said targeting unit comprises an amino acid sequence having at least 80% sequence identity to the amino acid sequence 24-93 of SEQ ID NO:1. [0303] 38. The vaccine according to any of the preceding embodiments, wherein said targeting unit consists of an amino acid sequence having at least 85% sequence identity to the amino acid sequence 24-93 of SEQ ID NO:1. [0304] 39. The vaccine according to any of the preceding embodiments, wherein said nucleotide sequence further encodes a signal peptide. [0305] 40. The vaccine according to embodiment 39, wherein said signal peptide comprises an amino acid sequence having at least 80% sequence identity to the amino acid sequence 1-23 of SEQ ID NO:1. [0306] 41. The vaccine according to embodiment 39 or 40, wherein said signal peptide consists of an amino acid sequence having at least 85% sequence identity to the amino acid sequence 1-23 of SEQ ID NO:1. [0307] 42. The vaccine according to any of the preceding embodiments, wherein said targeting unit, dimerization unit and antigenic unit in said peptide are in the N-terminal to C-terminal order of targeting unit, dimerization unit and antigenic unit. [0308] 43. The vaccine according to any of the preceding embodiments, wherein said polynucleotide sequence is human codon optimized. [0309] 44. The vaccine according to any of the preceding embodiments, wherein said polynucleotide sequence is a DNA nucleotide sequence or a RNA nucleotide sequence. [0310] 45. The vaccine according to any of the preceding embodiments, further comprising a pharmaceutically acceptable carrier and/or adjuvant. [0311] 46. A polynucleotide as defined in any of the embodiments 1-45. [0312] 47. A vector comprising the nucleotide sequence as defined in any of the embodiments 1-45. [0313] 48. A host cell comprising the nucleotide sequence as defined in any of the embodiments 1-45 or comprising the vector as defined in embodiment 47. [0314] 49. The polynucleotide according to embodiment 46 formulated for administration to a patient to induce production of the dimeric protein in said patient. [0315] 50. A polypeptide encoded by the nucleotide sequence as defined in any of the embodiments 1-45. [0316] 51. A dimeric protein consisting of two polypeptides as defined by embodiment 50. [0317] 52. The dimeric protein according to embodiment 51, being a homodimeric protein. [0318] 53. The polypeptide as defined in embodiment 50, the dimeric protein as defined in embodiment 51-52, or the polynucleotide as defined in embodiment 46 for use as a medicament. [0319] 54. A method for preparing a vaccine comprising an immunologically effective amount of the dimeric protein as defined in embodiment 50, or the polypeptide as defined in embodiment 50, the method comprising [0320] e) transfecting the polynucleotide as defined in embodiment 46 into a cell population; [0321] f) culturing the cell population; [0322] g) collecting and purifying the dimeric protein, or the polypeptide expressed from the cell population [0323] h) mixing the dimeric protein or polypeptide obtained under step c) with a pharmaceutically acceptable carrier thereby obtaining the vaccine. [0324] 55. A method for preparing a vaccine comprising an immunologically effective amount of the polynucleotide according to embodiment 46, said method comprising [0325] a. preparing the polynucleotide according to embodiment 46; [0326] b. mixing the polynucleotide obtained under step a) with a pharmaceutically acceptable carrier, thereby obtaining the vaccine. [0327] 56. The method according to embodiment 55, including the steps of: [0328] sequencing the exome of a tumor [0329] identifying tumor neoantigens comprising neoepitopes from said tumor, [0330] selecting neoepitopes based on antigenicity, prior to the step of preparing the polynucleotide. [0331] 57. A method of treating cancer in a patient, the method comprising administering to the patient in need thereof, the vaccine as defined in any of the embodiments 1-45. [0332] 58. The method according to embodiments 57, wherein the vaccine comprises a polynucleotide and is administered intradermally or intramuscular. [0333] 59. The method according to embodiment 58 wherein the polynucleotide is a DNA. [0334] 60. The method according to embodiment 59 wherein the polynucleotide is a RNA. [0335] 61. The method according to embodiments 57 to 60, wherein administration is carried out with a jet injector. [0336] 62. The method according to embodiments 57 to 60, wherein administration is assisted by electroporation.
EXAMPLES
Example 1: Construction and Expression of the Vaccines
[0337] Gene sequences were designed according to the following structure:
TABLE-US-00001 1: Native leader sequence for human LD78b. Signal peptide 2: Full length LD78b sequence. Targeting unit 3: Human hinge-region 1 from IgG3. Dimerization unit 4: Human hinge region 4 from IgG3. 5: Glycine-Serine linker. 6: Human CH3 domain from IgG3. 7: Glycine-Leucine linker. First linker 8: Neoepitope sequence (see below) Antigenic unit
[0338] Previously described exome sequencing and RNA sequencing of the mouse melanoma cancer cell line B16-F10 and the mouse colon cancer cell line CT26 revealed hundreds to thousands of tumor-specific non-synonymous mutations (Castle et al 2012, Castle et al 2014 and Kreiter et al 2015). In silico-based methods were used to identify potential immunogenic neo-epitopes. Mice were immunized with peptides encoding the mutated epitopes, and their immunogenicity was observed as specific T cell immune responses (ELISpot assay). Furthermore, vaccination of mice with the most immunogenic epitopes selected from the ELISpot conferred strong anti-tumor activity (Castle et al 2012 and Kreiter et al 2015).
[0339] Each of the neoepitopes are peptides of 27 amino acids separated by a flexible GGGGS linker. Short peptides (<20 amino acids) are processed and novel epitopes may be presented on MHC class I molecules and activate CD8+ T cells. However, it is preferred that the vaccine activates CD8+ and CD4+ T cells and therefore neoepitopes encoding for long peptides (>20 amino acids) are chosen. That may allow for efficient peptide processing and presentation on both MHC class I and II (Kreiter et al 2015). In the first two VB10.NEO-X constructs the selected hydrophobic and hydrophilic neoepitopes are evenly distributed. A neutral, flexible GGGGS linker between the 27mer neoepitopes is important to avoid generation of new immunogenic epitopes in the junctions of the combined neoepitopes.
[0340] The sequences of the neoepitopes found in the B16-F10 and CT26 cell lines are shown in Table 1 and 2.
TABLE-US-00002 TABLE 1 CT26 cell line Mutation number polypep- MHC I tide Sub.WT, Reactive score (Vacci- Mutated sequence used AA#, T cell (best pre- body) Gene for vaccination Mut) subtype diction) CT26- E2f8 VILPQAPSGPSYATYLQPAQA I522T CD8+ 0.1 PepM1 QMLTPP (SEQ ID NO: 14) CT26- Aldh18a1 LHSGQNHLKEMAISVLEARA P154S PepM2 CAAAGQS (SEQ ID NO: 15) CT26- Slc4a3 PLLPFYPPDEALEIGLELNSS T373I CD4+ 0.9 PepM3 ALPPTE (SEQ ID NO: 16) CT26- Nphp3 AGTQCEYWASRALDSEHSIG G234D CD4+ 0.1 PepM4 SMIQLPQ (SEQ ID NO: 17) CT26- Tdg AAYKGHHYPGPGNYFWKCL H169Y CD4+ 0.3 PepM5 FMSGLSEV (SEQ ID NO: 18) CT26- Ubqln1 DTLSAMSNPRAMQVLLQIQQ A62V PepM6 GLQTLAT (SEQ ID NO: 19) CT26- Slc20a1 DKPLRRNNSYTSYIMAICGMP T425I CD4+ 0.3 PepM7 LDSFRA (SEQ ID NO: 20) CT26- Dhx35 EVIQTSKYYMRDVIAIESAWLL T646I CD4+ 0.1 PepM8 ELAPH (SEQ ID NO: 21) CT26- Als2 GYISRVTAGKDSYIALVDKNI L675I CD8+ 0.2 PepM9 MGYIAS (SEQ ID NO: 22) CT26- Agxt2I2 EHIHRAGGLFVADAIQVGFGR E247A CD4+ 0.2 PepM10 IGKHFW (SEQ ID NO: 23) CT26- Tmem87a QAIVRGCSMPGPWRSGRLLV G63R CD8+ 0.7 PepM11 SRRWSVE (SEQ ID NO: 50) CT26- Ppp6r1 DGQLELLAQGALDNALSSMG D309N CD4+ PepM12 ALHALRP (SEQ ID NO: 51) CT26- Deptor SHDSRKSTSFMSVNPSKEIKI S253N CD4+ 0.3 PepM13 VSAVRR (SEQ ID NO: 52) CT26- Nap1I4 HTPSSYIETLPKAIKRRINALK V63I CD4+ 0.7 PepM14 QLQVR (SEQ ID NO: 53) CT26- Cxcr7 MKAFIFKYSAKTGFTKLIDASR L340F CD4+ 1.8 PepM15 VSETE (SEQ ID NO: 54) CT26- Dkk2 EGDPCLRSSDCIDEFCCARH G192E CD4+ 9.7 PepM16 FWTKICK (SEQ ID NO: 55) CT26- Trip12 WKGGPVKIDPLALMQAIERYL V1328M CD8+ PepM17 VVRGYG (SEQ ID NO: 56) CT26- Steap2 VTSIPSVSNALNWKEFSFIQS R388K CD4+ 6.8 PepM18 TLGYVA (SEQ ID NO: 57) Ct26- Gpc1 YRGANLHLEETLAGFWARLL E165G CD8+ 1.9 PepM19 ERLFKQL (SEQ ID NO: 58) CT26- Usp26 KTTLSHTQDSSQSLQSSSDS S715L n.d. 5.8 PepM20 SKSSRCS (SEQ ID NO: 59)
TABLE-US-00003 TABLE 2 B16-F10 cell line Mutation number polypep- Substi. MHC I tide WT, Reactive score (Vacci- Mutated sequence used AA#, T cell (best pre- body) Gene for vaccination Mut) subtype diction) B16-PepM1 Kif18b PSKPSFQEFVDWENVSPELNSTD K739N CD4+ 1.2 QPFL (SEQ ID NO: 4) B16-PepM2 Obsl1 REGVELCPGNKYEMRRHGTTHSL T176M CD8+ 2.3 VIHD (SEQ ID NO: 5) B16-PepM3 Def8 SHCHWNDLAVIPAGVVHNWDFEP R255G CD4+ 3.8 RKVS (SEQ ID NO: 6) B16-PepM4 Rpl13a GRGHLLGRLAAIVGKQVLLGRKVV A24G CD4+ 0.5 VVR (SEQ ID NO: 7) B16-PepM5 Tubb3 FRRKAFLHWYTGEAMDEMEFTEA G402A CD4+ 1.9 ESNM (SEQ ID NO: 8) B16-PepM6 Tnpo3 VVDRNPQFLDPVLAYLMKGLCEK G504A CD4+ 1 PLAS (SEQ ID NO: 9) B16-PepM7 Atp11a SSPDEVALVEGVQSLGFTYLRLKD R552S CD4+ 0.1 NYM (SEQ ID NO: 10) B16-PepM8 Cpsf3I EFKHIKAFDRTFANNPGPMVVFAT D314N CD4+ 0.5 PGM (SEQ ID NO: 11) B16-PepM9 Plod1 STANYNTSHLNNDVWQIFENPVD F530V CD4+ 0.1 WKEK (SEQ ID NO: 12) B16- Pbk DSGSPFPAAVILRDALHMARGLKY V145D CD8+ 0.1 PepM10 LHQ (SEQ ID NO: 13) B16- Ddx23 ANFESGKHKYRQTAMFTATMPPA V602A CD4+ 1.3 PepM11 VERL (SEQ ID NO: 36) B16- Actn4 NHSGLVTFQAFIDVMSRETTDTDT F835V CD4+ 0.2 PepM12 ADQ (SEQ ID NO: 60) B16- Tm9sf3 CGTAFFINFIAIYHHASRAIPFGTM Y382H CD4+ 0.2 PepM13 VA (SEQ ID NO: 61) B16- Eef2 FVVKAYLPVNESFAFTADLRSNTG G795A CD4+ 1.1 PepM14 GQA (SEQ ID NO: 62) B16- Gnas TPPPEEAMPFEFNGPAQGDHSQP S111G CD4+ 1.2 PepM15 PLQV (SEQ ID NO: 63) B16- Asf1b PKPDFSQLQRNILPSNPRVTRFHI A141P CD4+ 1.7 PepM16 NWD (SEQ ID NO: 64) B16- Mthfd1I IPSGTTILNCFHDVLSGKLSGGSP F294V CD4+ 1.7 PepM17 GVP (SEQ ID NO: 65) B16- Sema3b GFSQPLRRLVLHVVSAAQAERLA L663V CD4+ 2.9 PepM18 RAEE (SEQ ID NO: 66) B16- Mkm1 ECRITSNFVIPSEYWVEEKEEKQK N346Y CD4+ 1.4 PepM19 LIQ (SEQ ID NO: 67) B16- Ppp1r7 NIEGIDKLTQLKKPFLVNNKINKIEN L170P CD4+ 3.2 PepM20 I (SEQ ID NO: 68)
Example 2: Comparing Vaccibodies Comprising 3 or 10 Neoepitopes
[0341] Vaccibody vaccines containing either 3 or 10 neoepitopes were compared. In the 10 neoepitope Vaccibody DNA construct the place and order for the 3 first (N-terminal) peptides are similar as in the 3 neoepitope Vaccibody DNA construct. This is done to be able to compare the immunogenicity of these 3 neoepitopes in the context with 3 and in the context containing 7 more epitopes.
[0342] VB4001 (VB10.NEO CT26-X), VB4002 (VB10.NEO CT26-III), VB4003 (VB10.NEO B16-X) and
[0343] VB4004 (VB10.NEO B16-III) were selected as vaccine candidates. A schematic drawing of the vaccibodies are shown in FIG. 1.
[0344] The neoepitopes used for the vaccines VB4001-VB4021 are shown below. For example, VB4015 comprises three neoepitopes, B16 pepM1+pepM8+pepM3 that are separated by 5 amino acid linkers. VB4018 comprises 2 copies of the 10 neoepitopes, B16 pepM1+pepM2+pepM3+pepM4+pepM11+pepM6+pepM7+pepM8+pepM9+pepM10 that are separated by 5 amino acid linkers. The neoepitope sequences are shown in Tables 1 and 2.
[0345] VB4001=VB10.NEO CT26-X=CT26 pepM1-M10, 5 aa linker
[0346] VB4002=VB10.NEO CT26-III=CT26 pepM1-M3, 5 aa linker
[0347] VB4003=VB10.NEO B16-X=B16 pepM1-M10, 5 aa linker
[0348] VB4004=VB10.NEO B16-III=B16 pepM1-M3, 5 aa linker
[0349] VB4011=VB10.NEO B16-X=B16 pepM1-M10, 10 aa linker
[0350] VB4012=VB10.NEO B16-III=B16 pepM1-M3, 10 aa linker
[0351] VB4014=VB10.NEO B16-X=B16 hydrophobic core,
[0352] (pepM9+pepM5+pepM1+pepM4+pepM6+pepM8+pepM10+pepM3+pepM7+pepM2), 5 aa linker
[0353] VB4015=VB10.NEO B16-III=B16 pepM1+M8+M3, 5 aa linker
[0354] VB4016=VB10.NEO B16-III=B16 pepM1+M3+M2, 5 aa linker
[0355] VB4017=VB10.NEO B16-X=B16 pepM1-M4+M11+M6-M10, 5 aa linker
[0356] VB4018=VB10.NEO B16-XX=B16 pepM1-M4+M11+M6-M10.times.2, 5 aa linker
[0357] VB4019=VB10.NEO B16-Vx2=B16 pepM3+M4+M7+M9+M10.times.2, 5 aa linker
[0358] VB4021=VB10.NEO B16-Vx4=B16 pepM3+M4+M7+M9-M10.times.4, 5 aa linker
[0359] All neoepitope gene sequences were ordered from Genescript (New Jersey, US) and cloned into the expression vector pUMVC4a holding the LD78beta targeting unit and the hlgG3 dimerization unit.
[0360] All constructs were transfected into HEK293 cells and Vaccibody proteins in the supernatant were verified by Western blot and/or sandwich ELISA. Empty pUMVC4a vector was included as a negative control. FIG. 2, left panels: To illustrate the formation of intact homodimeric proteins, the proteins in the supernatant from transfected cells were detected in a Western blot by an anti-hMIP-1alpha antibody, in either the presence or absence of reducing agents. The formation of homodimers are shown in the left lane (- reducing agent) whereas the monomers are illustrated in the right lane (+ reducing agent). FIG. 2, right panel shows the expression level of the Vaccibody proteins in the supernatant of HEK293 cells transfected with the different VB10.NEO constructs detected by a sandwich ELISA using antibodies against both hMIP-1alpha and hlgG3. Right, upper panel shows the expression level of the VB10.NEO CT26-X (VB4001) and VB10.NEO CT26-III (VB4002) constructs, comprising 10 or 3 neoepitopes, respectively. Right, lower panel shows the expression level of the VB10.NEO B16-X (VB4003) and VB10.NEO B16-III (VB4004) constructs, comprising 10 or 3 neoepitopes, respectively. To compare the immunogenicity of vaccibodies comprising 3 or 10 neoepitopes, 20 .mu.g plasmid DNA of each vaccibody candidate were injected intramuscularly in the tibial anterior muscle of C57Bl/6-mice (for B16 constructs) or BALB/c-mice (for CT26 constructs), followed by electroporation using TriGrid, Ichor, (US). At day 13, the mice were euthanized and spleens were harvested.
[0361] The T cell responses were evaluated by IFN-gamma ELISpot. The results are shown in FIG. 3 where the T cell responses are indicated as the number of IFN-.gamma. spots/10.sup.6 splenocytes. We observe that vaccibodies comprising 10 neoepitopes induces significant T cell responses towards 4-6 of 10 included neoepitopes in the same mice. The peptides stimulating the strongest IFN-.gamma. response generally have the best MHC I binding score.
[0362] The total neoantigen-specific immune responses induced by vaccibody constructs comprising 3 or 10 neoepitopes are depicted in FIG. 4. Vaccibodies comprising 10 neoepitopes (VB10.NEO B16-X and VB10.NEO CT26-X) resulted in an increased total neoantigen-specific immune response when compared with vaccibodies comprising 3 neoepitopes (VB10.NEO B16-III and VB10.NEO CT26-III).
Example 3: Comparing Immunogenicity of Vaccibody DNA Vaccines and Corresponding Peptide Plus Adjuvant Vaccines
[0363] Before the VB10.NEO constructs are used in mice vaccination studies, Vaccibody protein expression and secretion in HEK293 cells are verified using a sandwich ELISA assay, as previously described in detail in the text for FIG. 2. The order of the neoepitopes could have an impact on the expression and secretion of functional Vaccibodies. In FIG. 5, upper panel we observe that the VB10.NEO B16-X construct VB4014 has a slightly improved expression and secretion of functional vaccibody proteins compared to the VB10.NEO B16-X construct VB4003. The 10 neopitopes in VB4014 is similar as for VB4003, however the order of the neoepitopes are changed and the most hydrophobic neoepitopes are located in the core in the neoepitope antigenic module. To test immunogenicity of Vaccibody DNA vaccines VB4003 and VB4014 compared with peptides comprising only neoepitopes delivered in combination with the poly (I:C) adjuvant, C57/Bl6 mice were injected with 20 .mu.g of the VB10.NEO B16-X constructs VB4003 and VB4014 (The induced immune responses were compared with immune responses of mice s.c. injected with 20 .mu.g or 200 .mu.g peptide mix+50 .mu.g poly I:C comprising the 10 neoepitopes encoded by VB4003 and VB4014. The T cell responses were evaluated by IFN-gamma ELISpot. The results, shown in FIG. 5 lower panel, illustrate that the vaccibodies clearly induces a much stronger response than peptide+adjuvant. Moreover, some of the animals immunized with the VB10.NEO B16-X VB4014 construct responded to all 10 neoepitopes included in the vaccine.
Example 4: Comparing Vaccibodies Comprising Second Linkers with a Length of 5 or 10 Amino Acids
[0364] Each of the neoepitopes is separated by a second linker. In the present example the second linker is a flexible GGGGS linker. To test if the length of the second linker has any effect on the expression level, HEK293 cells were transfected with VB10.NEO B16-X constructs comprising second linkers with a length of either 5 or 10 amino acids. FIG. 6 illustrates that changing the linker length from 5 (VB4003) to 10 (VB4011) amino acids does not affect expression of vaccibodies comprising 10 neoepitopes (FIG. 6, upper panel). To test if the length of the second linker has any effect on the immune response, C57Bl/6 mice were injected with VB10.NEO B16-X constructs comprising 10 neoepitopes with either 5 (VB4003) or 10 (VB4011) amino acid linkers. At day 13, the mice were sacrificed and splenocytes harvested, stimulated with the individual corresponding neoepitope peptides for 24 hours and T cell responses were quantified in an IFN-gamma ELISpot assay. The results are shown in FIG. 6, lower panel, and demonstrate that vaccibody constructs comprising 10 amino acid linkers (VB4011) lead to an increased total immune response when compared to vaccibodies comprising 5 amino acid linkers (VB4003). Empty vector was included as a negative control.
Example 5: Comparing Vaccibodies Comprising Different Number of Copies of Identical Neoepitopes
[0365] The following constructs were tested:
[0366] VB4003=VB10.NEO B16-X=B16 pepM1-M10, 5 aa linker
[0367] VB4018=VB10.NEO B16-XX=B16 pepM1-M4+M11+M6-M10.times.2, 5 aa linker
[0368] The expression level of VB10.NEO B16-X (VB4003) construct comprising 10 neoepitopes was compared to the expression level of VB10.NEO B16-XX (VB4018) comprising 2.times.10 neoepitopes. The results demonstrate that VB10.NEO B16-XX (VB4018) comprising 20 neoepitopes are slightly less expressed compared to VB10.NEO B16-X (VB4003) comprising 10 neoepitopes (FIG. 7, upper panel).
[0369] The immunogenicity of Vaccibodies comprising either 10 or 20 neoepitopes was tested by intramuscular injection of C57Bl/6 mice with the Vaccibody DNA vaccine VB10.NEO B16-X (VB4003) and VB10.NEO B16-XX (VB4018) At day 13, the mice were sacrificed and splenocytes harvested, stimulated with the individual corresponding neoepitope peptides for 24 hours and T cell responses were quantified in an IFN-gamma ELISpot assay. The results shown in FIG. 7, lower panel illustrate that the benefit of including 2 copies per neoepitope (2.times.10 neoepitopes) is limited on the total immune response, however, a broader immune response is observed towards individual neoepitopes.
[0370] Next, the expression levels of Vaccibody constructs comprising one or more copies of the 5 selected neoepitopes, PepM3, PepM4, PepM7, PepM9 and PepM10, were tested (FIG. 8, upper panel).
[0371] C57Bl/6 mice were injected with the following Vaccibody constructs:
[0372] VB4003=VB10.NEO B16-X=B16 pepM1-M10, 5 aa linker
[0373] VB4011=VB10.NEO B16-X=B16 pepM1-M10, 10 aa linker
[0374] VB4018=VB10.NEO B16-XX=B16 pepM1-M4+M11+M6-M10.times.2, 5 aa linker
[0375] VB4019=VB10.NEO B16-Vx2=B16 pepM3+M4+M7+M9+M10.times.2, 5 aa linker
[0376] VB4021=VB10.NEO B16-Vx4=B16 pepM3+M4+M7+M9+M10.times.4, 5 aa linker
[0377] The immune responses of the Vaccibody candidates for each of the five selected neoepitopes are shown in FIG. 8, lower panel. Multiple copies of the five neoepitopes had limited effect on the total immune response. However, several copies of each neoepitope (VB4018, VB4019 and VB4021) gives a more evenly immune response towards the 5 shared neoepitopes compared to the decatope VB4003, where the 5 neoepitopes are presented once. Interestingly, Vaccibodies comprising a 10 amino acid second linker and the neoepitopes only once (VB4011) displayed a better total immune response than Vaccibodies comprising multiple copies of the five neoepitopes.
Example 6: Comparing Vaccibodies Comprising Different Number of Neoepitopes
[0378] The immune response of vaccibody constructs comprising different numbers of neoepitopes were compared to test the immunological effect of adding further neoepitopes.
[0379] The total immune response was tested in the B16 melanoma mouse model using the following constructs:
[0380] NEO B16-X=VB4011=B16 pepM1-M10, 10 aa linker
[0381] NEO B16-XV=VB4024=B16 pepM1-M15, 10 aa linker
[0382] NEO B16-XX=VB4025=B16 pepM1-M20, 10 aa linker
[0383] The neoepitope sequences are shown in Table 2.
[0384] The expression levels of the three tested vaccibody constructs are shown in FIG. 11, upper panel.
[0385] C57Bl/6 mice were injected with the DNA vaccine candidates VB10.NEO B16-XV comprising 15 neoepitopes (VB4024) or VB10.NEO B16-XX comprising 20 neoepitopes (VB4025) compared to the VB10.NEO B16-X comprising 10 neoepitopes (VB4011). FIG. 11, lower panel, shows the total number of IFN.gamma.-spots per 10.sup.6 splenocytes. Constructs with 15 and 20 neoepitopes resulted in a broader immune response against more individual neoepitopes and a higher total T cell response when compared to constructs with only 10 neoepitopes. As a negative control, mice were injected with empty vector not comprising the neoepitopes. As seen from FIG. 11, lower panel, injections with empty vector did not lead to any significant immune response against the individual neoepitopes.
[0386] Further, the total immune response was tested in the CT26 melanoma mouse model using the following constructs
[0387] NEO CT26-X=VB4009=CT26 pepM1-M10, 10 aa linker
[0388] NEO CT26-XV=VB4026=CT26 pepM1-M15, 10 aa linker
[0389] NEO CT26-XX=VB4027=CT26 pepM1-M20, 10 aa linker
[0390] The neoepitope sequences are shown in Table 1.
[0391] BALB/c mice were injected with the DNA vaccine candidates VB10.NEO CT26-XV comprising 15 neoepitopes (VB4026) or VB10.NEO CT26-XX comprising 20 neoepitopes (VB4027) compared to the VB10.NEO CT26-X comprising 10 neoepitopes (VB4009). FIG. 12, lower panel, shows the total number of IFN.gamma.-spots per 10.sup.6 splenocytes. Constructs with 15 and 20 neoepitopes resulted in a broader immune response against more individual neoepitopes and a higher total T cell response when compared to constructs with only 10 neoepitopes. As a negative control, mice were injected with empty vector not comprising the neoepitopes. As seen from FIG. 12, lower panel, injections with empty vector did not lead to any significant immune response against the individual neoepitopes.
Example 7: Expression Levels of Different Vaccibody Constructs--are Compared
[0392] The following constructs were tested:
[0393] VB4004=VB10.NEO B16-III=B16 pepM1-M3, 5 aa linker
[0394] VB4012=VB10.NEO B16-III=B16 pepM1-M3, 10 aa linker
[0395] VB4015=VB10.NEO B16-III=B16 pepM1+M8+M3, 5 aa linker
[0396] VB4016=VB10.NEO B16-III=B16 pepM1+M3+M2, 5 aa linker
[0397] VB4017=VB10.NEO B16-X=B16 pepM1-M4+M11+M6-M10, 5 aa linker
[0398] VB4018=VB10.NEO B16-XX=B16 pepM1-M4+M11+M6-M10.times.2, 5 aa linker
[0399] Similar expression and secretion of functional vaccibody proteins are observed for VB10.NEO B16-X (VB4017) and VB10.NEO B16-XX (VB4018) (FIG. 9).
[0400] Improved expression and secretion of functional vaccibody proteins are observed when the 3 neoepitopes are spaced with a 10 aa linker as in the VB10.NEO B16-III (VB4012) construct compared to a 5 aa linker in the VB10.NEO B16-III (VB4004) construct (FIG. 10, upper panel). Moreover, by changing the order of the three neoepitopes as shown by comparing VB4004, VB4015 and VB4016 (FIG. 10, lower panel), may affect the expression levels of the vaccibodies.
Example 8: Therapeutic Effect
[0401] VB10.NEO were used as vaccine candidates for therapeutic vaccine studies.
[0402] 7.5.times.10.sup.4 B16.F10 cells or 1.times.10.sup.5 CT26 cells (ATCC) was injected in the thigh region of C57Bl/6 mice or BALB/c mice. After 1 and 8 days, the mice were vaccinated with 20 .mu.g plasmid DNA followed by electroporation, TriGrid, Ichor, US. Tumor sizes were measured two to three times a week. FIG. 13 shows that VB10.NEO DNA vaccine candidates comprising 10 neoepitopes are able to significantly delay and reduce tumour growth.
Example 9: Therapeutic DNA Vaccine
[0403] A therapeutic DNA vaccine to be used may be prepared by GMP manufacturing of the plasmid vaccine according to regulatory authorities' guidelines, and Fill & Finish of the DNA vaccine. The DNA vaccine may be formulated by dissolving in a saline solution, such as PBS at a concentration of 2-6 mg/ml. The vaccine may be administered either intradermal or intramuscular with or without following electroporation or alternatively with a jet injector.
TABLE-US-00004 SEQUENCES SEQ ID NO: 1 C-C motif chemokine 3-like 1 precursor including signal peptide and mature peptide (LD78-beta), aa 24-93: MQVSTAALAVLLCTMALCNQVLSAPLAADTPTACCFSYTSRQIPQNFIADYFETSSQCSKPSVIF LTKRGRQVCADPSEEWVQKYVSDLELSA SEQ ID NO: 2 DNA sequence of constant coding part of all VB10.NEO constructs For the purpose of illustration only, the different domains of the constructs are separated by an ''|'' with the domains in the following order: Signal peptide|human MIP-I.alpha.|Hinge hi|Hinge h4| Gly-Ser Linker or Gly-Leu linker|hCH3 IgG3|Gly-Ser Linker or Gly-Leu linker| The construct is a standard construct that can be used to insert neoepitopes. Neoepitope sequences can be added after the linker GGCCTCGGTGGCCTG. ATGCAGGTCTCCACTGCTGCCCTTGCCGTCCTCCTCTGCACCATGGCTCTCTGCAACCAG GTCCTCTCT|GCACCACTT GCTGCTGACACGCCGACCGCCTGCTGCTTCAGCTACACCTCCCGACAGATTCCACAGAAT TTCATAGCTGACTACTTTG AGACGAGCAGCCAGTGCTCCAAGCCCAGTGTCATCTTCCTAACCAAGAGAGGCCGGCAGG TCTGTGCTGACCCCAGTGA GGAGTGGGTCCAGAAATACGTCAGTGACCTGGAGCTGAGTGCC| GAGCTCAAAACCCCACTTGGTGACACAACTCACACA| GAGCCCAAATCTTGTGACACACCTCCCCCGTGCCCAAGGTGCCC A| GGCGGTGGAAGCAGCGGAGGTGGAAGTGGA| GGACAGCCCCGAGAACCACAGGTGTACACCCTGCCCCCATCCCGGGAGGAGATGACCAA GAACCAGGTCAGCCTGACCT GCCTGGTCAAAGGCTTCTACCCCAGCGACATCGCCGTGGAGTGGGAGAGCAGCGGGCAG CCGGAGAACAACTACAACAC CACGCCTCCCATGCTGGACTCCGACGGCTCCTTCTTCCTCTACAGCAAGCTCACCGTGGA CAAGAGCAGGTGGCAGCAG GGGAACATCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCGCTTCACGCAGAAG AGCCTCTCCCTGTCTCCGG GTAAA|GGCCTCGGTGGCCTG| SEQ ID NO: 3 Amino acid sequence of constant coding part of all VB10.NEO proteins: B4001 MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL| SEQ ID NO: 4 B16-F10 mutated epitope, B16-PepM1, amino acid sequence PSKPSFQEFVDWENVSPELNSTDQPFL SEQ ID NO: 5 B16-F10 mutated epitope, B16-PepM2, amino acid sequence REGVELCPGNKYEMRRHGTTHSLVIHD SEQ ID NO: 6 B16-F10 mutated epitope, B16-PepM3, amino acid sequence SHCHWNDLAVIPAGVVHNWDFEPRKVS SEQ ID NO: 7 B16-F10 mutated epitope, B16-PepM4, amino acid sequence GRGHLLGRLAAIVGKQVLLGRKVVVVR SEQ ID NO: 8 B16-F10 mutated epitope, B16-PepM5, amino acid sequence FRRKAFLHWYTGEAMDEMEFTEAESNM SEQ ID NO: 9 B16-F10 mutated epitope, B16-PepM6, amino acid sequence VVDRNPQFLDPVLAYLMKGLCEKPLAS SEQ ID NO: 10 B16-F10 mutated epitope, B16-PepM7, amino acid sequence SSPDEVALVEGVQSLGFTYLRLKDNYM SEQ ID NO: 11 B16-F10 mutated epitope, B16-PepM8, amino acid sequence EFKHIKAFDRTFANNPGPMVVFATPGM SEQ ID NO: 12 B16-F10 mutated epitope, B16-PepM9, amino acid sequence STANYNTSHLNNDVWQIFENPVDWKEK SEQ ID NO: 13 B16-F10 mutated epitope, B16-PepM10, amino acid sequence DSGSPFPAAVILRDALHMARGLKYLHQ SEQ ID NO: 14 CT26 mutated epitope, CT26-PepM1, amino acid sequence VILPQAPSGPSYATYLQPAQAQMLTPP SEQ ID NO: 15 CT26 mutated epitope, CT26-PepM2, amino acid sequence LHSGQNHLKEMAISVLEARACAAAGQS SEQ ID NO: 16 CT26 mutated epitope, CT26-PepM3, amino acid sequence PLLPFYPPDEALEIGLELNSSALPPTE SEQ ID NO: 17 CT26 mutated epitope, CT26-PepM4, amino acid sequence AGTQCEYWASRALDSEHSIGSMIQLPQ SEQ ID NO: 18 CT26 mutated epitope, CT26-PepM5, amino acid sequence AAYKGHHYPGPGNYFWKCLFMSGLSEV SEQ ID NO: 19 CT26 mutated epitope, CT26-PepM6, amino acid sequence DTLSAMSNPRAMQVLLQIQQGLQTLAT SEQ ID NO: 20 CT26 mutated epitope, CT26-PepM7, amino acid sequence DKPLRRNNSYTSYIMAICGMPLDSFRA SEQ ID NO: 21 CT26 mutated epitope, CT26-PepM8, amino acid sequence EVIQTSKYYMRDVIAIESAWLLELAPH SEQ ID NO: 22 CT26 mutated epitope, CT26-PepM9, amino acid sequence GYISRVTAGKDSYIALVDKNIMGYIAS SEQ ID NO: 23 CT26 mutated epitope, CT26-PepM10, amino acid sequence EHIHRAGGLFVADAIQVGFGRIGKHFW SEQ ID NO: 24 First linker, amino acid sequence: GLSGL SEQ ID NO: 25 First linker, amino acid sequence: GLGGL SEQ ID NO: 26 Hinge regions (IgG3 UH hinge), 12 amino acids: ELKTPLGDTTHT SEQ ID NO: 27 Hinge region (IgG3, MH hinge, 15 amino acids): EPKSCDTPPPCPRCP SEQ ID NO: 28 Gly-Ser Linker: GGGSSGGGSG SEQ ID NO: 29 hCH3 IgG3, amino acid sequence: GQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDG SFFLYSKLTVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK SEQ ID NO: 30 Amino acid sequence of VB4001 = VB10.NEO CT26-X = CT26 pepM1-M10, 5 aa linker The neoepitope sequences are inserted after GGGSSGGGSG. MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL| MHGDTPTLHEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| VILPQAPSGPSYATYLQPAQAQMLTPPGGGGSLHSGQNHLKEMAISVLEARACAAAGQSGGG GSPLLPFYPPDEALEIGLELNSSALPPTEGGGGSAGTQCEYWASRALDSEHSIGSMIQLPQGG GGSAAYKGHHYPGPGNYFWKCLFMSGLSEVGGGGSDTLSAMSNPRAMQVLLQIQQGLQTLA TGGGGSDKPLRRNNSYTSYIMAICGMPLDSFRAGGGGSEVIQTSKYYMRDVIAIESAWLLELAP HGGGGSGYISRVTAGKDSYIALVDKNIMGYIASGGGGSEHIHRAGGLFVADAIQVGFGRIGKHF W SEQ ID NO: 31 Amino acid sequence of VB4002 VB10.NEO CT26-III = CT26 pepM1-M3, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| VILPQAPSGPSYATYLQPAQAQMLTPPGGGGSLHSGQNHLKEMAISVLEARACAAAGQSGGG GSPLLPFYPPDEALEIGLELNSSALPPTE SEQ ID NO: 32 Amino acid sequence of VB4003 = VB10.NEO B16-X = B16 pepM1-M10, 5 aa linker (VB10.Neo-10B) MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSREGVELCPGNKYEMRRHGTTHSLVIHDGG GGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVV RGGGGSFRRKAFLHWYTGEAMDEMEFTEAESNMGGGGSVVDRNPQFLDPVLAYLMKGLCE KPLASGGGGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSEFKHIKAFDRTFANNPGPMV VFATPGMGGGGSSTANYNTSHLNNDVWQIFENPVDWKEKGGGGSDSGSPFPAAVILRDALH MARGLKYLHQ SEQ ID NO: 33 Amino acid sequence of VB4004 = VB10.NEO B16-III = B16 pepM1-M3, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSREGVELCPGNKYEMRRHGTTHSLVIHDGG GGSSHCHWNDLAVIPAGVVHNWDFEPRKVS SEQ ID NO: 34 Signal peptide MNFGLRLIFLVLTLKGVQC SEQ ID NO: 35 Signal peptide MDAMKRGLCCVLLLCGAVFVSP SEQ ID NO: 36 B16-F10 mutated epitope, B16-pepM11, amino acid sequence ANFESGKHKYRQTAMFTATMPPAVERL SEQ ID NO: 37 Amino acid sequence of VB4011 = VB10.NE0 B16-X = B16 pepM1-M10, 10 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSGGGGSREGVELCPGNKYEMRRHGTTHSLV IHDGGGGSGGGGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGGGGSGRGHLLGRLA AIVGKQVLLGRKVVVVRGGGGSGGGGSFRRKAFLHWYTGEAMDEMEFTEAESNMGGGGSG GGGSVVDRNPQFLDPVLAYLMKGLCEKPLASGGGGSGGGGSSSPDEVALVEGVQSLGFTYL RLKDNYMGGGGSGGGGSEFKHIKAFDRTFANNPGPMVVFATPGMGGGGSGGGGSSTANYN TSHLNNDVWQIFENPVDWKEKGGGGSGGGGSDSGSPFPAAVILRDALHMARGLKYLHQ SEQ ID NO: 38 Amino acid sequence of VB4012 = VB10.NE0 B16-III = B16 pepM1-M3, 10 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG|
PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSGGGGSREGVELCPGNKYEMRRHGTTHSLV IHDGGGGSGGGGSSHCHWNDLAVIPAGVVHNWDFEPRKVS SEQ ID NO: 39 Amino acid sequence of VB4014 = VB10.NEO B16-X = B16 hydrophobic core, (pepM9 + M5 + M1 + M4 + M6 + M8 + M10 + M3 + M7 + M2), 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEVVVQKYVSDLELSA|ELKTPLG DTTHT|EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| STANYNTSHLNNDVWQIFENPVDWKEKGGGGSFRRKAFLHWYTGEAMDEMEFTEAESNMG GGGSPSKPSFQEFVDWENVSPELNSTDQPFLGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVV RGGGGSVVDRNPQFLDPVLAYLMKGLCEKPLASGGGGSEFKHIKAFDRTFANNPGPMVVFAT PGMGGGGSDSGSPFPAAVILRDALHMARGLKYLHQGGGGSSHCHWNDLAVIPAGVVHNWDF EPRKVSGGGGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSREGVELCPGNKYEMRRHG TTHSLVIHD SEQ ID NO: 40 Amino acid sequence of VB4015 = VB10.NEO B16-III = B16 pepM1-M8-M3, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSEFKHIKAFDRTFANNPGPMVVFATPGMGGG GSSHCHWNDLAVIPAGVVHNWDFEPRKVS SEQ ID NO: 41 Amino acid sequence of VB4016 = VB10.NEO B16-III = B16 pepM1-M3-M2, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGG GGSREGVELCPGNKYEMRRHGTTHSLVIHD SEQ ID NO: 42 Amino acid sequence of VB4017 = VB10.NEO B16-X = B16 pepM1-M4 + M11 + M6-M10, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSREGVELCPGNKYEMRRHGTTHSLVIHDGG GGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVV RGGGGSANFESGKHKYRQTAMFTATMPPAVERLGGGGSVVDRNPQFLDPVLAYLMKGLCEK PLASGGGGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSEFKHIKAFDRTFANNPGPMVV FATPGMGGGGSSTANYNTSHLNNDVWQIFENPVDWKEKGGGGSDSGSPFPAAVILRDALHM ARGLKYLHQ SEQ ID NO: 43 Amino acid sequence of VB4018 = VB10.NEO B16-XX = B16 pepM1-M4 + M11 + M6-M10 x 2, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSREGVELCPGNKYEMRRHGTTHSLVIHDGG GGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVV RGGGGSANFESGKHKYRQTAMFTATMPPAVERLGGGGSVVDRNPQFLDPVLAYLMKGLCEK PLASGGGGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSEFKHIKAFDRTFANNPGPMVV FATPGMGGGGSSTANYNTSHLNNDVWQIFENPVDWKEKGGGGSDSGSPFPAAVILRDALHM ARGLKYLHQGGGGSPSKPSFQEFVDWENVSPELNSTDQPFLGGGGSREGVELCPGNKYEMR RHGTTHSLVIHDGGGGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGRGHLLGRLAAIV GKQVLLGRKVVVVRGGGGSANFESGKHKYRQTAMFTATMPPAVERLGGGGSVVDRNPQFLD PVLAYLMKGLCEKPLASGGGGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSEFKHIKAF DRTFANNPGPMVVFATPGMGGGGSSTANYNTSHLNNDVWQIFENPVDWKEKGGGGSDSGS PFPAAVILRDALHMARGLKYLHQ SEQ ID NO: 44 Amino acid sequence of VB4019 = VB10.NEO B16-Vx2 = B16 pepM3-M4-M7-M9-M10 x 2, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| SHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVVRGG GGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSSTANYNTSHLNNDVWQIFENPVDWKE KGGGGSDSGSPFPAAVILRDALHMARGLKYLHQGGGGSSHCHWNDLAVIPAGVVHNWDFEP RKVSGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVVRGGGGSSSPDEVALVEGVQSLGFTYLR LKDNYMGGGGSSTANYNTSHLNNDVWQIFENPVDWKEKGGGGSDSGSPFPAAVILRDALHM ARGLKYLHQ SEQ ID NO: 45 Amino acid sequence of VB4021 = VB10.NEO B16-Vx4 = B16 pepM3-M4-M7-M9-M10 x 4, 5 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| SHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVVRGG GGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSSTANYNTSHLNNDVWQIFENPVDWKE KGGGGSDSGSPFPAAVILRDALHMARGLKYLHQGGGGSSHCHWNDLAVIPAGVVHNWDFEP RKVSGGGGSGRGHLLGRLAAIVGKQVLLGRKVVVVRGGGGSSSPDEVALVEGVQSLGFTYLR LKDNYMGGGGSSTANYNTSHLNNDVWQIFENPVDWKEKGGGGSDSGSPFPAAVILRDALHM ARGLKYLHQGGGGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGRGHLLGRLAAIVGK QVLLGRKVVVVRGGGGSSSPDEVALVEGVQSLGFTYLRLKDNYMGGGGSSTANYNTSHLNN DVWQIFENPVDWKEKGGGGSDSGSPFPAAVILRDALHMARGLKYLHQ SEQ ID NO: 46 Amino acid sequence of VB4024 = VB10.NEO B16-XV = B16 pepM1-M15, 10 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSGGGGSREGVELCPGNKYEMRRHGTTHSLV IHDGGGGSGGGGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGGGGSGRGHLLGRLA AIVGKQVLLGRKVVVVRGGGGSGGGGSFRRKAFLHWYTGEAMDEMEFTEAESNMGGGGSG GGGSVVDRNPQFLDPVLAYLMKGLCEKPLASGGGGSGGGGSSSPDEVALVEGVQSLGFTYL RLKDNYMGGGGSGGGGSEFKHIKAFDRTFANNPGPMVVFATPGMGGGGSGGGGSSTANYN TSHLNNDVWQIFENPVDWKEKGGGGSGGGGSDSGSPFPAAVILRDALHMARGLKYLHQGGG GSGGGGSANFESGKHKYRQTAMFTATMPPAVERLGGGGSGGGGSNHSGLVTFQAFIDVMSR ETTDTDTADQGGGGSGGGGSCGTAFFINFIAIYHHASRAIPFGTMVAGGGGSGGGGSFVVKA YLPVNESFAFTADLRSNTGGQAGGGGSGGGGSTPPPEEAMPFEFNGPAQGDHSQPPLQV SEQ ID NO: 47 Amino acid sequence of VB4025 = VB10.NEO B16-XX = B16 pepM1-M20, 10 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| PSKPSFQEFVDWENVSPELNSTDQPFLGGGGSGGGGSREGVELCPGNKYEMRRHGTTHSLV IHDGGGGSGGGGSSHCHWNDLAVIPAGVVHNWDFEPRKVSGGGGSGGGGSGRGHLLGRLA AIVGKQVLLGRKVVVVRGGGGSGGGGSFRRKAFLHWYTGEAMDEMEFTEAESNMGGGGSG GGGSVVDRNPQFLDPVLAYLMKGLCEKPLASGGGGSGGGGSSSPDEVALVEGVQSLGFTYL RLKDNYMGGGGSGGGGSEFKHIKAFDRTFANNPGPMVVFATPGMGGGGSGGGGSSTANYN TSHLNNDVWQIFENPVDWKEKGGGGSGGGGSDSGSPFPAAVILRDALHMARGLKYLHQGGG GSGGGGSANFESGKHKYRQTAMFTATMPPAVERLGGGGSGGGGSNHSGLVTFQAFIDVMSR ETTDTDTADQGGGGSGGGGSCGTAFFINFIAIYHHASRAIPFGTMVAGGGGSGGGGSFVVKA YLPVNESFAFTADLRSNTGGQAGGGGSGGGGSTPPPEEAMPFEFNGPAQGDHSQPPLQVGG GGSGGGGSPKPDFSQLQRNILPSNPRVTRFHINWDGGGGSGGGGSIPSGTTILNCFHDVLSG KLSGGSPGVPGGGGSGGGGSGFSQPLRRLVLHVVSAAQAERLARAEEGGGGSGGGGSECRI TSNFVIPSEYWVEEKEEKQKLIQGGGGSGGGGSNIEGIDKLTQLKKPFLVNNKINKIENI SEQ ID NO: 48 Amino acid sequence of VB4026 = VB10.NEO CT26-XV = CT26 pepM1-M15, 10 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| VILPQAPSGPSYATYLQPAQAQMLTPPGGGGSGGGGSLHSGQNHLKEMAISVLEARACAAAG QSGGGGSGGGGSPLLPFYPPDEALEIGLELNSSALPPTEGGGGSGGGGSAGTQCEYWASRA LDSEHSIGSMIQLPQGGGGSGGGGSAAYKGHHYPGPGNYFWKCLFMSGLSEVGGGGSGGG GSDTLSAMSNPRAMQVLLQIQQGLQTLATGGGGSGGGGSDKPLRRNNSYTSYIMAICGMPLD SFRAGGGGSGGGGSEVIQTSKYYMRDVIAIESAWLLELAPHGGGGSGGGGSGYISRVTAGKD SYIALVDKNIMGYIASGGGGSGGGGSEHIHRAGGLFVADAIQVGFGRIGKHFWGGGGSGGGG SQAIVRGCSMPGPWRSGRLLVSRRWSVEGGGGSGGGGSDGQLELLAQGALDNALSSMGAL HALRPGGGGSGGGGSSHDSRKSTSFMSVNPSKEIKIVSAVRRGGGGSGGGGSHTPSSYIETL PKAIKRRINALKQLQVRGGGGSGGGGSMKAFIFKYSAKTGFTKLIDASRVSETE SEQ ID NO: 49 Amino acid sequence of VB4027 = VB10.NEO CT26-XX = CT26 pepM1-M20, 10 aa linker MQVSTAALAVLLCTMALCNQVLS|APLAADTPTACCFSYTSRQIPQNFIAD YFETSSQCSKPSVIFLTKRGRQVCADPSEEWVQKYVSDLELSA|ELKTPLG DTTHT I EPKSCDTPPPCPRCP|GGGSSGGGSG|GQPREPQVYTLPPSREEMTK NQVSLTCLVKGFYPSDIAVEWESSGQPENNYNTTPPMLDSDGSFFLYSKL TVDKSRWQQGNIFSCSVMHEALHNRFTQKSLSLSPGK|GLGGL|MHGDTPTL HEYMLDLQPETTDLYGYGQLNDSSEEEDEIDGPAGQAEPDRAHYNIVTFC CKCDSTLRLCVQSTHVDIRTLEDLLMGTLGIVCPICSQKP|GGGSSGGGSG| VILPQAPSGPSYATYLQPAQAQMLTPPGGGGSGGGGSLHSGQNHLKEMAISVLEARACAAAG QSGGGGSGGGGSPLLPFYPPDEALEIGLELNSSALPPTEGGGGSGGGGSAGTQCEYWASRA LDSEHSIGSMIQLPQGGGGSGGGGSAAYKGHHYPGPGNYFWKCLFMSGLSEVGGGGSGGG GSDTLSAMSNPRAMQVLLQIQQGLQTLATGGGGSGGGGSDKPLRRNNSYTSYIMAICGMPLD SFRAGGGGSGGGGSEVIQTSKYYMRDVIAIESAWLLELAPHGGGGSGGGGSGYISRVTAGKD SYIALVDKNIMGYIASGGGGSGGGGSEHIHRAGGLFVADAIQVGFGRIGKHFWGGGGSGGGG SQAIVRGCSMPGPWRSGRLLVSRRWSVEGGGGSGGGGSDGQLELLAQGALDNALSSMGAL HALRPGGGGSGGGGSSHDSRKSTSFMSVNPSKEIKIVSAVRRGGGGSGGGGSHTPSSYIETL PKAIKRRINALKQLQVRGGGGSGGGGSMKAFIFKYSAKTGFTKLIDASRVSETEGGGGSGGGG SEGDPCLRSSDCIDEFCCARHFWTKICKGGGGSGGGGSWKGGPVKIDPLALMQAIERYLVVR GYGGGGGSGGGGSVTSIPSVSNALNWKEFSFIQSTLGYVAGGGGSGGGGSYRGANLHLEET LAGFWARLLERLFKQLGGGGSGGGGSKTTLSHTQDSSQSLQSSSDSSKSSRCS SEQ ID NO: 50 CT26 mutated epitope, CT26-PepM11, amino acid sequence QAIVRGCSMPGPWRSGRLLVSRRWSVE SEQ ID NO: 51 CT26 mutated epitope, CT26-PepM12, amino acid sequence DGQLELLAQGALDNALSSMGALHALRP SEQ ID NO: 52 CT26 mutated epitope, CT26-PepM13, amino acid sequence SHDSRKSTSFMSVNPSKEIKIVSAVRR SEQ ID NO: 53 CT26 mutated epitope, CT26-PepM14, amino acid sequence HTPSSYIETLPKAIKRRINALKQLQVR SEQ ID NO: 54 CT26 mutated epitope, CT26-PepM15, amino acid sequence MKAFIFKYSAKTGFTKLIDASRVSETE SEQ ID NO: 55 CT26 mutated epitope, CT26-PepM16, amino acid sequence EGDPCLRSSDCIDEFCCARHFWTKICK SEQ ID NO: 56 CT26 mutated epitope, CT26-PepM17, amino acid sequence WKGGPVKIDPLALMQAIERYLVVRGYG SEQ ID NO: 57 CT26 mutated epitope, CT26-PepM18, amino acid sequence VTSIPSVSNALNWKEFSFIQSTLGYVA SEQ ID NO: 58 CT26 mutated epitope, CT26-PepM19, amino acid sequence YRGANLHLEETLAGFWARLLERLFKQL SEQ ID NO: 59 CT26 mutated epitope, CT26-PepM20, amino acid sequence KTTLSHTQDSSQSLQSSSDSSKSSRCS
SEQ ID NO: 60 B16-F10 mutated epitope, B16-PepM12, amino acid sequence NHSGLVTFQAFIDVMSRETTDTDTADQ SEQ ID NO: 61 B16-F10 mutated epitope, B16-PepM13, amino acid sequence CGTAFFINFIAIYHHASRAIPFGTMVA SEQ ID NO: 62 B16-F10 mutated epitope, B16-PepM14, amino acid sequence FVVKAYLPVNESFAFTADLRSNTGGQA SEQ ID NO: 63 B16-F10 mutated epitope, B16-PepM15, amino acid sequence TPPPEEAMPFEFNGPAQGDHSQPPLQV SEQ ID NO: 64 B16-F10 mutated epitope, B16-PepM16, amino acid sequence PKPDFSQLQRNILPSNPRVTRFHINWD SEQ ID NO: 65 B16-F10 mutated epitope, B16-PepM17, amino acid sequence IPSGTTILNCFHDVLSGKLSGGSPGVP SEQ ID NO: 66 B16-F10 mutated epitope, B16-PepM18, amino acid sequence GFSQPLRRLVLHVVSAAQAERLARAEE SEQ ID NO: 67 B16-F10 mutated epitope, B16-PepM19, amino acid sequence ECRITSNFVIPSEYVVVEEKEEKQKLIQ SEQ ID NO: 68 B16-F10 mutated epitope, B16-PepM20, amino acid sequence NIEGIDKLTQLKKPFLVNNKINKIENI SEQ ID NO: 69. Linker: GGGSS SEQ ID NO: 70. Linker: GGGSG SEQ ID NO: 71. Linker: GGGGS SEQ ID NO: 72. Linker: LGGGS SEQ ID NO: 73. Linker: GLGGS SEQ ID NO: 74. Linker: GGLGS SEQ ID NO: 75. Linker: GGGLS SEQ ID NO: 76. Linker: GGGGL SEQ ID NO: 77. Linker: LGGSG SEQ ID NO: 78. Linker: GLGSG SEQ ID NO: 79. Linker: GGLSG SEQ ID NO: 80. Linker: GGGLG SEQ ID NO: 81. Linker: GGGSL SEQ ID NO: 82. Linker: LGGSS SEQ ID NO: 83. Linker: GLGSS SEQ ID NO: 84. Linker: GGLSS SEQ ID NO: 85. Linker: GGGLS SEQ ID NO: 86. Linker: GGGSL SEQ ID NO: 87. Linker: LGLGS SEQ ID NO: 88. Linker: GLGLS SEQ ID NO: 89. Linker: GLLGS SEQ ID NO: 90. Linker: LGGLS SEQ ID NO: 91. Linker: GLGGL SEQ ID NO: 92. Linker: LGLSG SEQ ID NO: 93. Linker: GLLSG SEQ ID NO: 94. Linker: GGLSL SEQ ID NO: 95. Linker: GGLLG SEQ ID NO: 96. Linker: GLGSL SEQ ID NO: 97. Linker: LGLSS SEQ ID NO: 98. Linker: GLGLS SEQ ID NO: 99. Linker: GGLLS SEQ ID NO: 100. Linker: GLGSL SEQ ID NO: 101. Linker: GLGSL SEQ ID NO: 102. Linker: LGGGSGGGGS SEQ ID NO: 103. Linker: GLGGSGGGGS SEQ ID NO: 104. Linker: GGLGSGGGGS SEQ ID NO: 105. Linker: GGGLSGGGGS SEQ ID NO: 106. Linker: GGGGLGGGGS SEQ ID NO: 107. Linker: LGGSGGGGSG SEQ ID NO: 108. Linker: GLGSGGGGSG SEQ ID NO: 109. Linker: GGLSGGGGSG SEQ ID NO: 110. Linker: GGGLGGGGSG SEQ ID NO: 111. Linker: GGGSLGGGSG SEQ ID NO: 112. Linker: GGGSLGGGSG SEQ ID NO: 113. Linker: GLGSSGGGSS SEQ ID NO: 114. Linker: GGLSSGGGSS SEQ ID NO: 115. Linker: GGGLSGGGSS SEQ ID NO: 116. Linker: GGGSLGGGSS SEQ ID NO: 117. Linker: LGGGSLGGGS SEQ ID NO: 118. Linker: GLGGSGLGGS SEQ ID NO: 119. Linker: GGLGSGGLGS SEQ ID NO: 120. Linker: GGGLSGGGLS SEQ ID NO: 121. Linker: GGGGLGGGGL SEQ ID NO: 122. Linker: LGGSGLGGSG SEQ ID NO: 123. Linker: GLGSGGLGSG SEQ ID NO: 124. Linker: GGLSGGGLSG SEQ ID NO: 125. Linker: GGGLGGGGLG SEQ ID NO: 126. Linker: GGGSLGGGSL SEQ ID NO: 127. Linker: LGGSSLGGSS SEQ ID NO: 128. Linker: GLGSSGLGSS SEQ ID NO: 129. Linker: GGLSSGGLSS SEQ ID NO: 130. Linker: GGGLSGGGLS SEQ ID NO: 131. Linker: GGGSLGGGSL
Sequence CWU 1
1
131193PRTHomo sapiens 1Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu
Leu Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro
Leu Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr
Ser Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu
Thr Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr
Lys Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp
Val Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala 85 90
2726DNAArtificial SequenceVector 2atgcaggtct ccactgctgc ccttgccgtc
ctcctctgca ccatggctct ctgcaaccag 60gtcctctctg caccacttgc tgctgacacg
ccgaccgcct gctgcttcag ctacacctcc 120cgacagattc cacagaattt
catagctgac tactttgaga cgagcagcca gtgctccaag 180cccagtgtca
tcttcctaac caagagaggc cggcaggtct gtgctgaccc cagtgaggag
240tgggtccaga aatacgtcag tgacctggag ctgagtgccg agctcaaaac
cccacttggt 300gacacaactc acacagagcc caaatcttgt gacacacctc
ccccgtgccc aaggtgccca 360ggcggtggaa gcagcggagg tggaagtgga
ggacagcccc gagaaccaca ggtgtacacc 420ctgcccccat cccgggagga
gatgaccaag aaccaggtca gcctgacctg cctggtcaaa 480ggcttctacc
ccagcgacat cgccgtggag tgggagagca gcgggcagcc ggagaacaac
540tacaacacca cgcctcccat gctggactcc gacggctcct tcttcctcta
cagcaagctc 600accgtggaca agagcaggtg gcagcagggg aacatcttct
catgctccgt gatgcatgag 660gctctgcaca accgcttcac gcagaagagc
ctctccctgt ctccgggtaa aggcctcggt 720ggcctg 7263243PRTArtificial
SequenceHomodimeric construct 3Met Gln Val Ser Thr Ala Ala Leu Ala
Val Leu Leu Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser
Ala Pro Leu Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser
Tyr Thr Ser Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr
Phe Glu Thr Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe
Leu Thr Lys Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70
75 80 Trp Val Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu
Lys 85 90 95 Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys
Ser Cys Asp 100 105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly
Gly Ser Ser Gly Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro
Gln Val Tyr Thr Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro
Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185 190
Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195
200 205 Gln Gln Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu
His 210 215 220 Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly
Lys Gly Leu 225 230 235 240 Gly Gly Leu 427PRTMus musculus 4Pro Ser
Lys Pro Ser Phe Gln Glu Phe Val Asp Trp Glu Asn Val Ser 1 5 10 15
Pro Glu Leu Asn Ser Thr Asp Gln Pro Phe Leu 20 25 527PRTMus
musculus 5Arg Glu Gly Val Glu Leu Cys Pro Gly Asn Lys Tyr Glu Met
Arg Arg 1 5 10 15 His Gly Thr Thr His Ser Leu Val Ile His Asp 20 25
627PRTMus musculus 6Ser His Cys His Trp Asn Asp Leu Ala Val Ile Pro
Ala Gly Val Val 1 5 10 15 His Asn Trp Asp Phe Glu Pro Arg Lys Val
Ser 20 25 727PRTMus musculus 7Gly Arg Gly His Leu Leu Gly Arg Leu
Ala Ala Ile Val Gly Lys Gln 1 5 10 15 Val Leu Leu Gly Arg Lys Val
Val Val Val Arg 20 25 827PRTMus musculus 8Phe Arg Arg Lys Ala Phe
Leu His Trp Tyr Thr Gly Glu Ala Met Asp 1 5 10 15 Glu Met Glu Phe
Thr Glu Ala Glu Ser Asn Met 20 25 927PRTMus musculus 9Val Val Asp
Arg Asn Pro Gln Phe Leu Asp Pro Val Leu Ala Tyr Leu 1 5 10 15 Met
Lys Gly Leu Cys Glu Lys Pro Leu Ala Ser 20 25 1027PRTMus musculus
10Ser Ser Pro Asp Glu Val Ala Leu Val Glu Gly Val Gln Ser Leu Gly 1
5 10 15 Phe Thr Tyr Leu Arg Leu Lys Asp Asn Tyr Met 20 25
1127PRTMus musculus 11Glu Phe Lys His Ile Lys Ala Phe Asp Arg Thr
Phe Ala Asn Asn Pro 1 5 10 15 Gly Pro Met Val Val Phe Ala Thr Pro
Gly Met 20 25 1227PRTMus musculus 12Ser Thr Ala Asn Tyr Asn Thr Ser
His Leu Asn Asn Asp Val Trp Gln 1 5 10 15 Ile Phe Glu Asn Pro Val
Asp Trp Lys Glu Lys 20 25 1327PRTMus musculus 13Asp Ser Gly Ser Pro
Phe Pro Ala Ala Val Ile Leu Arg Asp Ala Leu 1 5 10 15 His Met Ala
Arg Gly Leu Lys Tyr Leu His Gln 20 25 1427PRTMus musculus 14Val Ile
Leu Pro Gln Ala Pro Ser Gly Pro Ser Tyr Ala Thr Tyr Leu 1 5 10 15
Gln Pro Ala Gln Ala Gln Met Leu Thr Pro Pro 20 25 1527PRTMus
musculus 15Leu His Ser Gly Gln Asn His Leu Lys Glu Met Ala Ile Ser
Val Leu 1 5 10 15 Glu Ala Arg Ala Cys Ala Ala Ala Gly Gln Ser 20 25
1627PRTMus musculus 16Pro Leu Leu Pro Phe Tyr Pro Pro Asp Glu Ala
Leu Glu Ile Gly Leu 1 5 10 15 Glu Leu Asn Ser Ser Ala Leu Pro Pro
Thr Glu 20 25 1727PRTMus musculus 17Ala Gly Thr Gln Cys Glu Tyr Trp
Ala Ser Arg Ala Leu Asp Ser Glu 1 5 10 15 His Ser Ile Gly Ser Met
Ile Gln Leu Pro Gln 20 25 1827PRTMus musculus 18Ala Ala Tyr Lys Gly
His His Tyr Pro Gly Pro Gly Asn Tyr Phe Trp 1 5 10 15 Lys Cys Leu
Phe Met Ser Gly Leu Ser Glu Val 20 25 1927PRTMus musculus 19Asp Thr
Leu Ser Ala Met Ser Asn Pro Arg Ala Met Gln Val Leu Leu 1 5 10 15
Gln Ile Gln Gln Gly Leu Gln Thr Leu Ala Thr 20 25 2027PRTMus
musculus 20Asp Lys Pro Leu Arg Arg Asn Asn Ser Tyr Thr Ser Tyr Ile
Met Ala 1 5 10 15 Ile Cys Gly Met Pro Leu Asp Ser Phe Arg Ala 20 25
2127PRTMus musculus 21Glu Val Ile Gln Thr Ser Lys Tyr Tyr Met Arg
Asp Val Ile Ala Ile 1 5 10 15 Glu Ser Ala Trp Leu Leu Glu Leu Ala
Pro His 20 25 2227PRTMus musculus 22Gly Tyr Ile Ser Arg Val Thr Ala
Gly Lys Asp Ser Tyr Ile Ala Leu 1 5 10 15 Val Asp Lys Asn Ile Met
Gly Tyr Ile Ala Ser 20 25 2327PRTMus musculus 23Glu His Ile His Arg
Ala Gly Gly Leu Phe Val Ala Asp Ala Ile Gln 1 5 10 15 Val Gly Phe
Gly Arg Ile Gly Lys His Phe Trp 20 25 245PRTArtificial
SequenceLinker 24Gly Leu Ser Gly Leu 1 5 255PRTArtificial
SequenceLinker 25Gly Leu Gly Gly Leu 1 5 2612PRTArtificial
SequenceHinge region 26Glu Leu Lys Thr Pro Leu Gly Asp Thr Thr His
Thr 1 5 10 2715PRTArtificial SequenceHinge region 27Glu Pro Lys Ser
Cys Asp Thr Pro Pro Pro Cys Pro Arg Cys Pro 1 5 10 15
2810PRTArtificial SequenceLinker 28Gly Gly Gly Ser Ser Gly Gly Gly
Ser Gly 1 5 10 29107PRTHomo sapiens 29Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser Arg Glu 1 5 10 15 Glu Met Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe 20 25 30 Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Ser Gly Gln Pro Glu 35 40 45 Asn
Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe 50 55
60 Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly
65 70 75 80 Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn
Arg Phe 85 90 95 Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 100
105 30666PRTArtificial SequenceConstruct 30Met Gln Val Ser Thr Ala
Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln
Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys
Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45
Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50
55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu
Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala
Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu
Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro
Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met
Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val 145 150 155 160 Lys Gly
Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly 165 170 175
Gln Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180
185 190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg
Trp 195 200 205 Gln Gln Gly Asn Ile Phe Ser Cys Ser Val Met His Glu
Ala Leu His 210 215 220 Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser
Pro Gly Lys Gly Leu 225 230 235 240 Gly Gly Leu Met His Gly Asp Thr
Pro Thr Leu His Glu Tyr Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr
Thr Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu
Glu Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro
Asp Arg Ala His Tyr Asn Ile Val Thr Phe Cys Cys Lys Cys 290 295 300
Asp Ser Thr Leu Arg Leu Cys Val Gln Ser Thr His Val Asp Ile Arg 305
310 315 320 Thr Leu Glu Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys
Pro Ile 325 330 335 Cys Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly
Gly Ser Gly Val 340 345 350 Ile Leu Pro Gln Ala Pro Ser Gly Pro Ser
Tyr Ala Thr Tyr Leu Gln 355 360 365 Pro Ala Gln Ala Gln Met Leu Thr
Pro Pro Gly Gly Gly Gly Ser Leu 370 375 380 His Ser Gly Gln Asn His
Leu Lys Glu Met Ala Ile Ser Val Leu Glu 385 390 395 400 Ala Arg Ala
Cys Ala Ala Ala Gly Gln Ser Gly Gly Gly Gly Ser Pro 405 410 415 Leu
Leu Pro Phe Tyr Pro Pro Asp Glu Ala Leu Glu Ile Gly Leu Glu 420 425
430 Leu Asn Ser Ser Ala Leu Pro Pro Thr Glu Gly Gly Gly Gly Ser Ala
435 440 445 Gly Thr Gln Cys Glu Tyr Trp Ala Ser Arg Ala Leu Asp Ser
Glu His 450 455 460 Ser Ile Gly Ser Met Ile Gln Leu Pro Gln Gly Gly
Gly Gly Ser Ala 465 470 475 480 Ala Tyr Lys Gly His His Tyr Pro Gly
Pro Gly Asn Tyr Phe Trp Lys 485 490 495 Cys Leu Phe Met Ser Gly Leu
Ser Glu Val Gly Gly Gly Gly Ser Asp 500 505 510 Thr Leu Ser Ala Met
Ser Asn Pro Arg Ala Met Gln Val Leu Leu Gln 515 520 525 Ile Gln Gln
Gly Leu Gln Thr Leu Ala Thr Gly Gly Gly Gly Ser Asp 530 535 540 Lys
Pro Leu Arg Arg Asn Asn Ser Tyr Thr Ser Tyr Ile Met Ala Ile 545 550
555 560 Cys Gly Met Pro Leu Asp Ser Phe Arg Ala Gly Gly Gly Gly Ser
Glu 565 570 575 Val Ile Gln Thr Ser Lys Tyr Tyr Met Arg Asp Val Ile
Ala Ile Glu 580 585 590 Ser Ala Trp Leu Leu Glu Leu Ala Pro His Gly
Gly Gly Gly Ser Gly 595 600 605 Tyr Ile Ser Arg Val Thr Ala Gly Lys
Asp Ser Tyr Ile Ala Leu Val 610 615 620 Asp Lys Asn Ile Met Gly Tyr
Ile Ala Ser Gly Gly Gly Gly Ser Glu 625 630 635 640 His Ile His Arg
Ala Gly Gly Leu Phe Val Ala Asp Ala Ile Gln Val 645 650 655 Gly Phe
Gly Arg Ile Gly Lys His Phe Trp 660 665 31442PRTArtificial
SequenceConstruct 31Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu
Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu
Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser
Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr
Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys
Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val
Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95
Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100
105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly
Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr
Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe
Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln
Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215 220
Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225
230 235 240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu Tyr
Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr
Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp
Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr Asn
Ile Val Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu
Cys Val Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu Glu
Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys
Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Val 340 345
350 Ile Leu Pro Gln Ala Pro Ser Gly Pro Ser Tyr Ala Thr Tyr Leu Gln
355 360 365 Pro Ala Gln Ala Gln Met Leu Thr Pro Pro Gly Gly Gly Gly
Ser Leu 370 375
380 His Ser Gly Gln Asn His Leu Lys Glu Met Ala Ile Ser Val Leu Glu
385 390 395 400 Ala Arg Ala Cys Ala Ala Ala Gly Gln Ser Gly Gly Gly
Gly Ser Pro 405 410 415 Leu Leu Pro Phe Tyr Pro Pro Asp Glu Ala Leu
Glu Ile Gly Leu Glu 420 425 430 Leu Asn Ser Ser Ala Leu Pro Pro Thr
Glu 435 440 32817PRTArtificial SequenceConstruct 32Met Gln Val Ser
Thr Ala Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10 15 Leu Cys
Asn Gln Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr 20 25 30
Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn Phe Ile 35
40 45 Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys Pro Ser Val
Ile 50 55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys Ala Asp Pro
Ser Glu Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp Leu Glu Leu
Ser Ala Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr Thr His Thr
Ile Glu Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro Cys Pro Arg
Cys Pro Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser Gly Gly Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140 Ser Arg Glu
Glu Met Thr Lys Met Gln Val Ser Thr Ala Ala Leu Ala 145 150 155 160
Val Leu Leu Cys Thr Met Ala Leu Cys Asn Gln Val Leu Ser Ala Pro 165
170 175 Leu Ala Ala Asp Thr Pro Thr Ala Cys Cys Phe Ser Tyr Thr Ser
Arg 180 185 190 Gln Ile Pro Gln Asn Phe Ile Ala Asp Tyr Phe Glu Thr
Ser Ser Gln 195 200 205 Cys Ser Lys Pro Ser Val Ile Phe Leu Thr Lys
Arg Gly Arg Gln Val 210 215 220 Cys Ala Asp Pro Ser Glu Glu Trp Val
Gln Lys Tyr Val Ser Asp Leu 225 230 235 240 Glu Leu Ser Ala Glu Leu
Lys Thr Pro Leu Gly Asp Thr Thr His Thr 245 250 255 Ile Glu Pro Lys
Ser Cys Asp Thr Pro Pro Pro Cys Pro Arg Cys Pro 260 265 270 Gly Gly
Gly Ser Ser Gly Gly Gly Ser Gly Gly Gln Pro Arg Glu Pro 275 280 285
Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln 290
295 300 Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala 305 310 315 320 Val Glu Trp Glu Ser Ser Gly Gln Pro Glu Asn Asn
Tyr Asn Thr Thr 325 330 335 Pro Pro Met Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys Leu 340 345 350 Thr Val Asp Lys Ser Arg Trp Gln
Gln Gly Asn Ile Phe Ser Cys Ser 355 360 365 Val Met His Glu Ala Leu
His Asn Arg Phe Thr Gln Lys Ser Leu Ser 370 375 380 Leu Ser Pro Gly
Lys Gly Leu Gly Gly Leu Met His Gly Asp Thr Pro 385 390 395 400 Thr
Leu His Glu Tyr Met Leu Asp Leu Gln Pro Glu Thr Thr Asp Leu 405 410
415 Tyr Gly Tyr Gly Gln Leu Asn Asp Ser Ser Glu Glu Glu Asp Glu Ile
420 425 430 Asp Gly Pro Ala Gly Gln Ala Glu Pro Asp Arg Ala His Tyr
Asn Ile 435 440 445 Val Thr Phe Cys Cys Lys Cys Asp Ser Thr Leu Arg
Leu Cys Val Gln 450 455 460 Ser Thr His Val Asp Ile Arg Thr Leu Glu
Asp Leu Leu Met Gly Thr 465 470 475 480 Leu Gly Ile Val Cys Pro Ile
Cys Ser Gln Lys Pro Gly Gly Gly Ser 485 490 495 Ser Gly Gly Gly Ser
Gly Pro Ser Lys Pro Ser Phe Gln Glu Phe Val 500 505 510 Asp Trp Glu
Asn Val Ser Pro Glu Leu Asn Ser Thr Asp Gln Pro Phe 515 520 525 Leu
Gly Gly Gly Gly Ser Arg Glu Gly Val Glu Leu Cys Pro Gly Asn 530 535
540 Lys Tyr Glu Met Arg Arg His Gly Thr Thr His Ser Leu Val Ile His
545 550 555 560 Asp Gly Gly Gly Gly Ser Ser His Cys His Trp Asn Asp
Leu Ala Val 565 570 575 Ile Pro Ala Gly Val Val His Asn Trp Asp Phe
Glu Pro Arg Lys Val 580 585 590 Ser Gly Gly Gly Gly Ser Gly Arg Gly
His Leu Leu Gly Arg Leu Ala 595 600 605 Ala Ile Val Gly Lys Gln Val
Leu Leu Gly Arg Lys Val Val Val Val 610 615 620 Arg Gly Gly Gly Gly
Ser Phe Arg Arg Lys Ala Phe Leu His Trp Tyr 625 630 635 640 Thr Gly
Glu Ala Met Asp Glu Met Glu Phe Thr Glu Ala Glu Ser Asn 645 650 655
Met Gly Gly Gly Gly Ser Val Val Asp Arg Asn Pro Gln Phe Leu Asp 660
665 670 Pro Val Leu Ala Tyr Leu Met Lys Gly Leu Cys Glu Lys Pro Leu
Ala 675 680 685 Ser Gly Gly Gly Gly Ser Ser Ser Pro Asp Glu Val Ala
Leu Val Glu 690 695 700 Gly Val Gln Ser Leu Gly Phe Thr Tyr Leu Arg
Leu Lys Asp Asn Tyr 705 710 715 720 Met Gly Gly Gly Gly Ser Glu Phe
Lys His Ile Lys Ala Phe Asp Arg 725 730 735 Thr Phe Ala Asn Asn Pro
Gly Pro Met Val Val Phe Ala Thr Pro Gly 740 745 750 Met Gly Gly Gly
Gly Ser Ser Thr Ala Asn Tyr Asn Thr Ser His Leu 755 760 765 Asn Asn
Asp Val Trp Gln Ile Phe Glu Asn Pro Val Asp Trp Lys Glu 770 775 780
Lys Gly Gly Gly Gly Ser Asp Ser Gly Ser Pro Phe Pro Ala Ala Val 785
790 795 800 Ile Leu Arg Asp Ala Leu His Met Ala Arg Gly Leu Lys Tyr
Leu His 805 810 815 Gln 33442PRTArtificial SequenceConstruct 33Met
Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10
15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr
20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn
Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys
Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys
Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp
Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr
Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro
Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser
Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140
Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val 145
150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met
Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr
Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln Gly Asn Ile Phe Ser Cys
Ser Val Met His Glu Ala Leu His 210 215 220 Asn Arg Phe Thr Gln Lys
Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225 230 235 240 Gly Gly Leu
Met His Gly Asp Thr Pro Thr Leu His Glu Tyr Met Leu 245 250 255 Asp
Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260 265
270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln Ala
275 280 285 Glu Pro Asp Arg Ala His Tyr Asn Ile Val Thr Phe Cys Cys
Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu Cys Val Gln Ser Thr His
Val Asp Ile Arg 305 310 315 320 Thr Leu Glu Asp Leu Leu Met Gly Thr
Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys Ser Gln Lys Pro Gly Gly
Gly Ser Ser Gly Gly Gly Ser Gly Pro 340 345 350 Ser Lys Pro Ser Phe
Gln Glu Phe Val Asp Trp Glu Asn Val Ser Pro 355 360 365 Glu Leu Asn
Ser Thr Asp Gln Pro Phe Leu Gly Gly Gly Gly Ser Arg 370 375 380 Glu
Gly Val Glu Leu Cys Pro Gly Asn Lys Tyr Glu Met Arg Arg His 385 390
395 400 Gly Thr Thr His Ser Leu Val Ile His Asp Gly Gly Gly Gly Ser
Ser 405 410 415 His Cys His Trp Asn Asp Leu Ala Val Ile Pro Ala Gly
Val Val His 420 425 430 Asn Trp Asp Phe Glu Pro Arg Lys Val Ser 435
440 3419PRTArtificial SequenceSignal peptide 34Met Asn Phe Gly Leu
Arg Leu Ile Phe Leu Val Leu Thr Leu Lys Gly 1 5 10 15 Val Gln Cys
3522PRTArtificial SequenceSignal peptide 35Met Asp Ala Met Lys Arg
Gly Leu Cys Cys Val Leu Leu Leu Cys Gly 1 5 10 15 Ala Val Phe Val
Ser Pro 20 3627PRTMus musculus 36Ala Asn Phe Glu Ser Gly Lys His
Lys Tyr Arg Gln Thr Ala Met Phe 1 5 10 15 Thr Ala Thr Met Pro Pro
Ala Val Glu Arg Leu 20 25 37711PRTArtificial SequenceConstruct
37Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1
5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro
Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln
Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser
Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val
Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser
Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp
Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro
Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly
Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135
140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu
Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro
Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu
Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln Gly Asn Ile Phe Ser
Cys Ser Val Met His Glu Ala Leu His 210 215 220 Asn Arg Phe Thr Gln
Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225 230 235 240 Gly Gly
Leu Met His Gly Asp Thr Pro Thr Leu His Glu Tyr Met Leu 245 250 255
Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260
265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln
Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr Asn Ile Val Thr Phe Cys
Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu Cys Val Gln Ser Thr
His Val Asp Ile Arg 305 310 315 320 Thr Leu Glu Asp Leu Leu Met Gly
Thr Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys Ser Gln Lys Pro Gly
Gly Gly Ser Ser Gly Gly Gly Ser Gly Pro 340 345 350 Ser Lys Pro Ser
Phe Gln Glu Phe Val Asp Trp Glu Asn Val Ser Pro 355 360 365 Glu Leu
Asn Ser Thr Asp Gln Pro Phe Leu Gly Gly Gly Gly Ser Gly 370 375 380
Gly Gly Gly Ser Arg Glu Gly Val Glu Leu Cys Pro Gly Asn Lys Tyr 385
390 395 400 Glu Met Arg Arg His Gly Thr Thr His Ser Leu Val Ile His
Asp Gly 405 410 415 Gly Gly Gly Ser Gly Gly Gly Gly Ser Ser His Cys
His Trp Asn Asp 420 425 430 Leu Ala Val Ile Pro Ala Gly Val Val His
Asn Trp Asp Phe Glu Pro 435 440 445 Arg Lys Val Ser Gly Gly Gly Gly
Ser Gly Gly Gly Gly Ser Gly Arg 450 455 460 Gly His Leu Leu Gly Arg
Leu Ala Ala Ile Val Gly Lys Gln Val Leu 465 470 475 480 Leu Gly Arg
Lys Val Val Val Val Arg Gly Gly Gly Gly Ser Gly Gly 485 490 495 Gly
Gly Ser Phe Arg Arg Lys Ala Phe Leu His Trp Tyr Thr Gly Glu 500 505
510 Ala Met Asp Glu Met Glu Phe Thr Glu Ala Glu Ser Asn Met Gly Gly
515 520 525 Gly Gly Ser Gly Gly Gly Gly Ser Val Val Asp Arg Asn Pro
Gln Phe 530 535 540 Leu Asp Pro Val Leu Ala Tyr Leu Met Lys Gly Leu
Cys Glu Lys Pro 545 550 555 560 Leu Ala Ser Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser Ser Ser Pro 565 570 575 Asp Glu Val Ala Leu Val Glu
Gly Val Gln Ser Leu Gly Phe Thr Tyr 580 585 590 Leu Arg Leu Lys Asp
Asn Tyr Met Gly Gly Gly Gly Ser Gly Gly Gly 595 600 605 Gly Ser Glu
Phe Lys His Ile Lys Ala Phe Asp Arg Thr Phe Ala Asn 610 615 620 Asn
Pro Gly Pro Met Val Val Phe Ala Thr Pro Gly Met Gly Gly Gly 625 630
635 640 Gly Ser Gly Gly Gly Gly Ser Ser Thr Ala Asn Tyr Asn Thr Ser
His 645 650 655 Leu Asn Asn Asp Val Trp Gln Ile Phe Glu Asn Pro Val
Asp Trp Lys 660 665 670 Glu Lys Gly Gly Gly Gly Ser Gly Gly Gly Gly
Ser Asp Ser Gly Ser 675 680 685 Pro Phe Pro Ala Ala Val Ile Leu Arg
Asp Ala Leu His Met Ala Arg 690 695 700 Gly Leu Lys Tyr Leu His Gln
705 710 38452PRTArtificial SequenceConstruct 38Met Gln Val Ser Thr
Ala Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10 15 Leu Cys Asn
Gln Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr 20 25 30 Ala
Cys Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn Phe Ile 35 40
45 Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys Pro Ser Val Ile
50 55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys Ala Asp Pro Ser
Glu Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser
Ala Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr Thr His Thr Ile
Glu Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro Cys Pro Arg Cys
Pro Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser Gly Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140 Ser Arg Glu Glu
Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val 145 150 155 160 Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly 165
170
175 Gln Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp
180 185 190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp 195 200 205 Gln Gln Gly Asn Ile Phe Ser Cys Ser Val Met His
Glu Ala Leu His 210 215 220 Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu
Ser Pro Gly Lys Gly Leu 225 230 235 240 Gly Gly Leu Met His Gly Asp
Thr Pro Thr Leu His Glu Tyr Met Leu 245 250 255 Asp Leu Gln Pro Glu
Thr Thr Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260 265 270 Asp Ser Ser
Glu Glu Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln Ala 275 280 285 Glu
Pro Asp Arg Ala His Tyr Asn Ile Val Thr Phe Cys Cys Lys Cys 290 295
300 Asp Ser Thr Leu Arg Leu Cys Val Gln Ser Thr His Val Asp Ile Arg
305 310 315 320 Thr Leu Glu Asp Leu Leu Met Gly Thr Leu Gly Ile Val
Cys Pro Ile 325 330 335 Cys Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly
Gly Gly Ser Gly Pro 340 345 350 Ser Lys Pro Ser Phe Gln Glu Phe Val
Asp Trp Glu Asn Val Ser Pro 355 360 365 Glu Leu Asn Ser Thr Asp Gln
Pro Phe Leu Gly Gly Gly Gly Ser Gly 370 375 380 Gly Gly Gly Ser Arg
Glu Gly Val Glu Leu Cys Pro Gly Asn Lys Tyr 385 390 395 400 Glu Met
Arg Arg His Gly Thr Thr His Ser Leu Val Ile His Asp Gly 405 410 415
Gly Gly Gly Ser Gly Gly Gly Gly Ser Ser His Cys His Trp Asn Asp 420
425 430 Leu Ala Val Ile Pro Ala Gly Val Val His Asn Trp Asp Phe Glu
Pro 435 440 445 Arg Lys Val Ser 450 39666PRTArtificial
SequenceConstruct 39Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu
Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu
Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser
Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr
Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys
Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val
Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95
Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100
105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly
Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr
Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe
Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln
Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215 220
Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225
230 235 240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu Tyr
Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr
Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp
Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr Asn
Ile Val Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu
Cys Val Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu Glu
Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys
Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Ser 340 345
350 Thr Ala Asn Tyr Asn Thr Ser His Leu Asn Asn Asp Val Trp Gln Ile
355 360 365 Phe Glu Asn Pro Val Asp Trp Lys Glu Lys Gly Gly Gly Gly
Ser Phe 370 375 380 Arg Arg Lys Ala Phe Leu His Trp Tyr Thr Gly Glu
Ala Met Asp Glu 385 390 395 400 Met Glu Phe Thr Glu Ala Glu Ser Asn
Met Gly Gly Gly Gly Ser Pro 405 410 415 Ser Lys Pro Ser Phe Gln Glu
Phe Val Asp Trp Glu Asn Val Ser Pro 420 425 430 Glu Leu Asn Ser Thr
Asp Gln Pro Phe Leu Gly Gly Gly Gly Ser Gly 435 440 445 Arg Gly His
Leu Leu Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val 450 455 460 Leu
Leu Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly Ser Val 465 470
475 480 Val Asp Arg Asn Pro Gln Phe Leu Asp Pro Val Leu Ala Tyr Leu
Met 485 490 495 Lys Gly Leu Cys Glu Lys Pro Leu Ala Ser Gly Gly Gly
Gly Ser Glu 500 505 510 Phe Lys His Ile Lys Ala Phe Asp Arg Thr Phe
Ala Asn Asn Pro Gly 515 520 525 Pro Met Val Val Phe Ala Thr Pro Gly
Met Gly Gly Gly Gly Ser Asp 530 535 540 Ser Gly Ser Pro Phe Pro Ala
Ala Val Ile Leu Arg Asp Ala Leu His 545 550 555 560 Met Ala Arg Gly
Leu Lys Tyr Leu His Gln Gly Gly Gly Gly Ser Ser 565 570 575 His Cys
His Trp Asn Asp Leu Ala Val Ile Pro Ala Gly Val Val His 580 585 590
Asn Trp Asp Phe Glu Pro Arg Lys Val Ser Gly Gly Gly Gly Ser Ser 595
600 605 Ser Pro Asp Glu Val Ala Leu Val Glu Gly Val Gln Ser Leu Gly
Phe 610 615 620 Thr Tyr Leu Arg Leu Lys Asp Asn Tyr Met Gly Gly Gly
Gly Ser Arg 625 630 635 640 Glu Gly Val Glu Leu Cys Pro Gly Asn Lys
Tyr Glu Met Arg Arg His 645 650 655 Gly Thr Thr His Ser Leu Val Ile
His Asp 660 665 40442PRTArtificial SequenceConstruct 40Met Gln Val
Ser Thr Ala Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10 15 Leu
Cys Asn Gln Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr 20 25
30 Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn Phe Ile
35 40 45 Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys Pro Ser
Val Ile 50 55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys Ala Asp
Pro Ser Glu Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp Leu Glu
Leu Ser Ala Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr Thr His
Thr Ile Glu Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro Cys Pro
Arg Cys Pro Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser Gly Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140 Ser Arg
Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val 145 150 155
160 Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly
165 170 175 Gln Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp
Ser Asp 180 185 190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp
Lys Ser Arg Trp 195 200 205 Gln Gln Gly Asn Ile Phe Ser Cys Ser Val
Met His Glu Ala Leu His 210 215 220 Asn Arg Phe Thr Gln Lys Ser Leu
Ser Leu Ser Pro Gly Lys Gly Leu 225 230 235 240 Gly Gly Leu Met His
Gly Asp Thr Pro Thr Leu His Glu Tyr Met Leu 245 250 255 Asp Leu Gln
Pro Glu Thr Thr Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260 265 270 Asp
Ser Ser Glu Glu Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln Ala 275 280
285 Glu Pro Asp Arg Ala His Tyr Asn Ile Val Thr Phe Cys Cys Lys Cys
290 295 300 Asp Ser Thr Leu Arg Leu Cys Val Gln Ser Thr His Val Asp
Ile Arg 305 310 315 320 Thr Leu Glu Asp Leu Leu Met Gly Thr Leu Gly
Ile Val Cys Pro Ile 325 330 335 Cys Ser Gln Lys Pro Gly Gly Gly Ser
Ser Gly Gly Gly Ser Gly Pro 340 345 350 Ser Lys Pro Ser Phe Gln Glu
Phe Val Asp Trp Glu Asn Val Ser Pro 355 360 365 Glu Leu Asn Ser Thr
Asp Gln Pro Phe Leu Gly Gly Gly Gly Ser Glu 370 375 380 Phe Lys His
Ile Lys Ala Phe Asp Arg Thr Phe Ala Asn Asn Pro Gly 385 390 395 400
Pro Met Val Val Phe Ala Thr Pro Gly Met Gly Gly Gly Gly Ser Ser 405
410 415 His Cys His Trp Asn Asp Leu Ala Val Ile Pro Ala Gly Val Val
His 420 425 430 Asn Trp Asp Phe Glu Pro Arg Lys Val Ser 435 440
41442PRTArtificial SequenceConstruct 41Met Gln Val Ser Thr Ala Ala
Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val
Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys
Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala
Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55
60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu
65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu
Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro
Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly
Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln
Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185
190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp
195 200 205 Gln Gln Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala
Leu His 210 215 220 Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro
Gly Lys Gly Leu 225 230 235 240 Gly Gly Leu Met His Gly Asp Thr Pro
Thr Leu His Glu Tyr Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr Thr
Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu Glu
Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro Asp
Arg Ala His Tyr Asn Ile Val Thr Phe Cys Cys Lys Cys 290 295 300 Asp
Ser Thr Leu Arg Leu Cys Val Gln Ser Thr His Val Asp Ile Arg 305 310
315 320 Thr Leu Glu Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys Pro
Ile 325 330 335 Cys Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly
Ser Gly Pro 340 345 350 Ser Lys Pro Ser Phe Gln Glu Phe Val Asp Trp
Glu Asn Val Ser Pro 355 360 365 Glu Leu Asn Ser Thr Asp Gln Pro Phe
Leu Gly Gly Gly Gly Ser Ser 370 375 380 His Cys His Trp Asn Asp Leu
Ala Val Ile Pro Ala Gly Val Val His 385 390 395 400 Asn Trp Asp Phe
Glu Pro Arg Lys Val Ser Gly Gly Gly Gly Ser Arg 405 410 415 Glu Gly
Val Glu Leu Cys Pro Gly Asn Lys Tyr Glu Met Arg Arg His 420 425 430
Gly Thr Thr His Ser Leu Val Ile His Asp 435 440 42666PRTArtificial
SequenceConstruct 42Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu
Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu
Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser
Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr
Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys
Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val
Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95
Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100
105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly
Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr
Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe
Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln
Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215 220
Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225
230 235 240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu Tyr
Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr
Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp
Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr Asn
Ile Val Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu
Cys Val Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu Glu
Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys
Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Pro 340 345
350 Ser Lys Pro Ser Phe Gln Glu Phe Val Asp Trp Glu Asn Val Ser Pro
355 360 365 Glu Leu Asn Ser Thr Asp Gln Pro Phe Leu Gly Gly Gly Gly
Ser Arg 370 375 380 Glu Gly Val Glu Leu Cys Pro Gly Asn Lys Tyr Glu
Met Arg Arg His 385 390 395 400 Gly Thr Thr His Ser Leu Val Ile His
Asp Gly Gly Gly Gly Ser Ser 405 410 415 His Cys His Trp Asn Asp Leu
Ala Val Ile Pro Ala Gly Val Val His 420 425 430 Asn Trp Asp Phe Glu
Pro Arg Lys Val Ser Gly Gly Gly Gly Ser Gly 435 440 445 Arg Gly His
Leu Leu Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val 450 455
460 Leu Leu Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly Ser Ala
465 470 475 480 Asn Phe Glu Ser Gly Lys His Lys Tyr Arg Gln Thr Ala
Met Phe Thr 485 490 495 Ala Thr Met Pro Pro Ala Val Glu Arg Leu Gly
Gly Gly Gly Ser Val 500 505 510 Val Asp Arg Asn Pro Gln Phe Leu Asp
Pro Val Leu Ala Tyr Leu Met 515 520 525 Lys Gly Leu Cys Glu Lys Pro
Leu Ala Ser Gly Gly Gly Gly Ser Ser 530 535 540 Ser Pro Asp Glu Val
Ala Leu Val Glu Gly Val Gln Ser Leu Gly Phe 545 550 555 560 Thr Tyr
Leu Arg Leu Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser Glu 565 570 575
Phe Lys His Ile Lys Ala Phe Asp Arg Thr Phe Ala Asn Asn Pro Gly 580
585 590 Pro Met Val Val Phe Ala Thr Pro Gly Met Gly Gly Gly Gly Ser
Ser 595 600 605 Thr Ala Asn Tyr Asn Thr Ser His Leu Asn Asn Asp Val
Trp Gln Ile 610 615 620 Phe Glu Asn Pro Val Asp Trp Lys Glu Lys Gly
Gly Gly Gly Ser Asp 625 630 635 640 Ser Gly Ser Pro Phe Pro Ala Ala
Val Ile Leu Arg Asp Ala Leu His 645 650 655 Met Ala Arg Gly Leu Lys
Tyr Leu His Gln 660 665 43986PRTArtificial SequenceConstruct 43Met
Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10
15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr
20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn
Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys
Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys
Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp
Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr
Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro
Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser
Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140
Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val 145
150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met
Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr
Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln Gly Asn Ile Phe Ser Cys
Ser Val Met His Glu Ala Leu His 210 215 220 Asn Arg Phe Thr Gln Lys
Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225 230 235 240 Gly Gly Leu
Met His Gly Asp Thr Pro Thr Leu His Glu Tyr Met Leu 245 250 255 Asp
Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260 265
270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln Ala
275 280 285 Glu Pro Asp Arg Ala His Tyr Asn Ile Val Thr Phe Cys Cys
Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu Cys Val Gln Ser Thr His
Val Asp Ile Arg 305 310 315 320 Thr Leu Glu Asp Leu Leu Met Gly Thr
Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys Ser Gln Lys Pro Gly Gly
Gly Ser Ser Gly Gly Gly Ser Gly Pro 340 345 350 Ser Lys Pro Ser Phe
Gln Glu Phe Val Asp Trp Glu Asn Val Ser Pro 355 360 365 Glu Leu Asn
Ser Thr Asp Gln Pro Phe Leu Gly Gly Gly Gly Ser Arg 370 375 380 Glu
Gly Val Glu Leu Cys Pro Gly Asn Lys Tyr Glu Met Arg Arg His 385 390
395 400 Gly Thr Thr His Ser Leu Val Ile His Asp Gly Gly Gly Gly Ser
Ser 405 410 415 His Cys His Trp Asn Asp Leu Ala Val Ile Pro Ala Gly
Val Val His 420 425 430 Asn Trp Asp Phe Glu Pro Arg Lys Val Ser Gly
Gly Gly Gly Ser Gly 435 440 445 Arg Gly His Leu Leu Gly Arg Leu Ala
Ala Ile Val Gly Lys Gln Val 450 455 460 Leu Leu Gly Arg Lys Val Val
Val Val Arg Gly Gly Gly Gly Ser Ala 465 470 475 480 Asn Phe Glu Ser
Gly Lys His Lys Tyr Arg Gln Thr Ala Met Phe Thr 485 490 495 Ala Thr
Met Pro Pro Ala Val Glu Arg Leu Gly Gly Gly Gly Ser Val 500 505 510
Val Asp Arg Asn Pro Gln Phe Leu Asp Pro Val Leu Ala Tyr Leu Met 515
520 525 Lys Gly Leu Cys Glu Lys Pro Leu Ala Ser Gly Gly Gly Gly Ser
Ser 530 535 540 Ser Pro Asp Glu Val Ala Leu Val Glu Gly Val Gln Ser
Leu Gly Phe 545 550 555 560 Thr Tyr Leu Arg Leu Lys Asp Asn Tyr Met
Gly Gly Gly Gly Ser Glu 565 570 575 Phe Lys His Ile Lys Ala Phe Asp
Arg Thr Phe Ala Asn Asn Pro Gly 580 585 590 Pro Met Val Val Phe Ala
Thr Pro Gly Met Gly Gly Gly Gly Ser Ser 595 600 605 Thr Ala Asn Tyr
Asn Thr Ser His Leu Asn Asn Asp Val Trp Gln Ile 610 615 620 Phe Glu
Asn Pro Val Asp Trp Lys Glu Lys Gly Gly Gly Gly Ser Asp 625 630 635
640 Ser Gly Ser Pro Phe Pro Ala Ala Val Ile Leu Arg Asp Ala Leu His
645 650 655 Met Ala Arg Gly Leu Lys Tyr Leu His Gln Gly Gly Gly Gly
Ser Pro 660 665 670 Ser Lys Pro Ser Phe Gln Glu Phe Val Asp Trp Glu
Asn Val Ser Pro 675 680 685 Glu Leu Asn Ser Thr Asp Gln Pro Phe Leu
Gly Gly Gly Gly Ser Arg 690 695 700 Glu Gly Val Glu Leu Cys Pro Gly
Asn Lys Tyr Glu Met Arg Arg His 705 710 715 720 Gly Thr Thr His Ser
Leu Val Ile His Asp Gly Gly Gly Gly Ser Ser 725 730 735 His Cys His
Trp Asn Asp Leu Ala Val Ile Pro Ala Gly Val Val His 740 745 750 Asn
Trp Asp Phe Glu Pro Arg Lys Val Ser Gly Gly Gly Gly Ser Gly 755 760
765 Arg Gly His Leu Leu Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val
770 775 780 Leu Leu Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly
Ser Ala 785 790 795 800 Asn Phe Glu Ser Gly Lys His Lys Tyr Arg Gln
Thr Ala Met Phe Thr 805 810 815 Ala Thr Met Pro Pro Ala Val Glu Arg
Leu Gly Gly Gly Gly Ser Val 820 825 830 Val Asp Arg Asn Pro Gln Phe
Leu Asp Pro Val Leu Ala Tyr Leu Met 835 840 845 Lys Gly Leu Cys Glu
Lys Pro Leu Ala Ser Gly Gly Gly Gly Ser Ser 850 855 860 Ser Pro Asp
Glu Val Ala Leu Val Glu Gly Val Gln Ser Leu Gly Phe 865 870 875 880
Thr Tyr Leu Arg Leu Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser Glu 885
890 895 Phe Lys His Ile Lys Ala Phe Asp Arg Thr Phe Ala Asn Asn Pro
Gly 900 905 910 Pro Met Val Val Phe Ala Thr Pro Gly Met Gly Gly Gly
Gly Ser Ser 915 920 925 Thr Ala Asn Tyr Asn Thr Ser His Leu Asn Asn
Asp Val Trp Gln Ile 930 935 940 Phe Glu Asn Pro Val Asp Trp Lys Glu
Lys Gly Gly Gly Gly Ser Asp 945 950 955 960 Ser Gly Ser Pro Phe Pro
Ala Ala Val Ile Leu Arg Asp Ala Leu His 965 970 975 Met Ala Arg Gly
Leu Lys Tyr Leu His Gln 980 985 44666PRTArtificial
SequenceConstruct 44Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu
Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu
Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser
Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr
Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys
Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val
Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95
Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100
105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly
Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr
Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe
Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln
Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215 220
Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225
230 235 240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu Tyr
Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr
Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp
Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr Asn
Ile Val Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu
Cys Val Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu Glu
Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys
Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Ser 340 345
350 His Cys His Trp Asn Asp Leu Ala Val Ile Pro Ala Gly Val Val His
355 360 365 Asn Trp Asp Phe Glu Pro Arg Lys Val Ser Gly Gly Gly Gly
Ser Gly 370 375 380 Arg Gly His Leu Leu Gly Arg Leu Ala Ala Ile Val
Gly Lys Gln Val 385 390 395 400 Leu Leu Gly Arg Lys Val Val Val Val
Arg Gly Gly Gly Gly Ser Ser 405 410 415 Ser Pro Asp Glu Val Ala Leu
Val Glu Gly Val Gln Ser Leu Gly Phe 420 425 430 Thr Tyr Leu Arg Leu
Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser Ser 435 440 445 Thr Ala Asn
Tyr Asn Thr Ser His Leu Asn Asn Asp Val Trp Gln Ile 450 455 460 Phe
Glu Asn Pro Val Asp Trp Lys Glu Lys Gly Gly Gly Gly Ser Asp 465 470
475 480 Ser Gly Ser Pro Phe Pro Ala Ala Val Ile Leu Arg Asp Ala Leu
His 485 490 495 Met Ala Arg Gly Leu Lys Tyr Leu His Gln Gly Gly Gly
Gly Ser Ser 500 505 510 His Cys His Trp Asn Asp Leu Ala Val Ile Pro
Ala Gly Val Val His 515 520 525 Asn Trp Asp Phe Glu Pro Arg Lys Val
Ser Gly Gly Gly Gly Ser Gly 530 535 540 Arg Gly His Leu Leu Gly Arg
Leu Ala Ala Ile Val Gly Lys Gln Val 545 550 555 560 Leu Leu Gly Arg
Lys Val Val Val Val Arg Gly Gly Gly Gly Ser Ser 565 570 575 Ser Pro
Asp Glu Val Ala Leu Val Glu Gly Val Gln Ser Leu Gly Phe 580 585 590
Thr Tyr Leu Arg Leu Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser Ser 595
600 605 Thr Ala Asn Tyr Asn Thr Ser His Leu Asn Asn Asp Val Trp Gln
Ile 610 615 620 Phe Glu Asn Pro Val Asp Trp Lys Glu Lys Gly Gly Gly
Gly Ser Asp 625 630 635 640 Ser Gly Ser Pro Phe Pro Ala Ala Val Ile
Leu Arg Asp Ala Leu His 645 650 655 Met Ala Arg Gly Leu Lys Tyr Leu
His Gln 660 665 45826PRTArtificial SequenceConstruct 45Met Gln Val
Ser Thr Ala Ala Leu Ala Val Leu Leu Cys Thr Met Ala 1 5 10 15 Leu
Cys Asn Gln Val Leu Ser Ala Pro Leu Ala Ala Asp Thr Pro Thr 20 25
30 Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln Ile Pro Gln Asn Phe Ile
35 40 45 Ala Asp Tyr Phe Glu Thr Ser Ser Gln Cys Ser Lys Pro Ser
Val Ile 50 55 60 Phe Leu Thr Lys Arg Gly Arg Gln Val Cys Ala Asp
Pro Ser Glu Glu 65 70 75 80 Trp Val Gln Lys Tyr Val Ser Asp Leu Glu
Leu Ser Ala Glu Leu Lys 85 90 95 Thr Pro Leu Gly Asp Thr Thr His
Thr Ile Glu Pro Lys Ser Cys Asp 100 105 110 Thr Pro Pro Pro Cys Pro
Arg Cys Pro Gly Gly Gly Ser Ser Gly Gly 115 120 125 Gly Ser Gly Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro 130 135 140 Ser Arg
Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val 145 150 155
160 Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly
165 170 175 Gln Pro Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp
Ser Asp 180 185 190 Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp
Lys Ser Arg Trp 195 200 205 Gln Gln Gly Asn Ile Phe Ser Cys Ser Val
Met His Glu Ala Leu His 210 215 220 Asn Arg Phe Thr Gln Lys Ser Leu
Ser Leu Ser Pro Gly Lys Gly Leu 225 230 235 240 Gly Gly Leu Met His
Gly Asp Thr Pro Thr Leu His Glu Tyr Met Leu 245 250 255 Asp Leu Gln
Pro Glu Thr Thr Asp Leu Tyr Gly Tyr Gly Gln Leu Asn 260 265 270 Asp
Ser Ser Glu Glu Glu Asp Glu Ile Asp Gly Pro Ala Gly Gln Ala 275 280
285 Glu Pro Asp Arg Ala His Tyr Asn Ile Val Thr Phe Cys Cys Lys Cys
290 295 300 Asp Ser Thr Leu Arg Leu Cys Val Gln Ser Thr His Val Asp
Ile Arg 305 310 315 320 Thr Leu Glu Asp Leu Leu Met Gly Thr Leu Gly
Ile Val Cys Pro Ile 325 330 335 Cys Ser Gln Lys Pro Gly Gly Gly Ser
Ser Gly Gly Gly Ser Gly Ser 340 345 350 His Cys His Trp Asn Asp Leu
Ala Val Ile Pro Ala Gly Val Val His 355 360 365 Asn Trp Asp Phe Glu
Pro Arg Lys Val Ser Gly Gly Gly Gly Ser Gly 370 375 380 Arg Gly His
Leu Leu Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val 385 390 395 400
Leu Leu Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly Ser Ser 405
410 415 Ser Pro Asp Glu Val Ala Leu Val Glu Gly Val Gln Ser Leu Gly
Phe 420 425 430 Thr Tyr Leu
Arg Leu Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser Ser 435 440 445 Thr
Ala Asn Tyr Asn Thr Ser His Leu Asn Asn Asp Val Trp Gln Ile 450 455
460 Phe Glu Asn Pro Val Asp Trp Lys Glu Lys Gly Gly Gly Gly Ser Asp
465 470 475 480 Ser Gly Ser Pro Phe Pro Ala Ala Val Ile Leu Arg Asp
Ala Leu His 485 490 495 Met Ala Arg Gly Leu Lys Tyr Leu His Gln Gly
Gly Gly Gly Ser Ser 500 505 510 His Cys His Trp Asn Asp Leu Ala Val
Ile Pro Ala Gly Val Val His 515 520 525 Asn Trp Asp Phe Glu Pro Arg
Lys Val Ser Gly Gly Gly Gly Ser Gly 530 535 540 Arg Gly His Leu Leu
Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val 545 550 555 560 Leu Leu
Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly Ser Ser 565 570 575
Ser Pro Asp Glu Val Ala Leu Val Glu Gly Val Gln Ser Leu Gly Phe 580
585 590 Thr Tyr Leu Arg Leu Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser
Ser 595 600 605 Thr Ala Asn Tyr Asn Thr Ser His Leu Asn Asn Asp Val
Trp Gln Ile 610 615 620 Phe Glu Asn Pro Val Asp Trp Lys Glu Lys Gly
Gly Gly Gly Ser Asp 625 630 635 640 Ser Gly Ser Pro Phe Pro Ala Ala
Val Ile Leu Arg Asp Ala Leu His 645 650 655 Met Ala Arg Gly Leu Lys
Tyr Leu His Gln Gly Gly Gly Gly Ser Ser 660 665 670 His Cys His Trp
Asn Asp Leu Ala Val Ile Pro Ala Gly Val Val His 675 680 685 Asn Trp
Asp Phe Glu Pro Arg Lys Val Ser Gly Gly Gly Gly Ser Gly 690 695 700
Arg Gly His Leu Leu Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val 705
710 715 720 Leu Leu Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly
Ser Ser 725 730 735 Ser Pro Asp Glu Val Ala Leu Val Glu Gly Val Gln
Ser Leu Gly Phe 740 745 750 Thr Tyr Leu Arg Leu Lys Asp Asn Tyr Met
Gly Gly Gly Gly Ser Ser 755 760 765 Thr Ala Asn Tyr Asn Thr Ser His
Leu Asn Asn Asp Val Trp Gln Ile 770 775 780 Phe Glu Asn Pro Val Asp
Trp Lys Glu Lys Gly Gly Gly Gly Ser Asp 785 790 795 800 Ser Gly Ser
Pro Phe Pro Ala Ala Val Ile Leu Arg Asp Ala Leu His 805 810 815 Met
Ala Arg Gly Leu Lys Tyr Leu His Gln 820 825 46896PRTArtificial
SequenceVB construct 46Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu
Leu Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro
Leu Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr
Ser Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu
Thr Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr
Lys Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp
Val Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90
95 Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp
100 105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser
Gly Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn
Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe
Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln
Gln Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215
220 Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu
225 230 235 240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu
Tyr Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly
Tyr Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile
Asp Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr
Asn Ile Val Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg
Leu Cys Val Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu
Glu Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335
Cys Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Pro 340
345 350 Ser Lys Pro Ser Phe Gln Glu Phe Val Asp Trp Glu Asn Val Ser
Pro 355 360 365 Glu Leu Asn Ser Thr Asp Gln Pro Phe Leu Gly Gly Gly
Gly Ser Gly 370 375 380 Gly Gly Gly Ser Arg Glu Gly Val Glu Leu Cys
Pro Gly Asn Lys Tyr 385 390 395 400 Glu Met Arg Arg His Gly Thr Thr
His Ser Leu Val Ile His Asp Gly 405 410 415 Gly Gly Gly Ser Gly Gly
Gly Gly Ser Ser His Cys His Trp Asn Asp 420 425 430 Leu Ala Val Ile
Pro Ala Gly Val Val His Asn Trp Asp Phe Glu Pro 435 440 445 Arg Lys
Val Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Arg 450 455 460
Gly His Leu Leu Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val Leu 465
470 475 480 Leu Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly Ser
Gly Gly 485 490 495 Gly Gly Ser Phe Arg Arg Lys Ala Phe Leu His Trp
Tyr Thr Gly Glu 500 505 510 Ala Met Asp Glu Met Glu Phe Thr Glu Ala
Glu Ser Asn Met Gly Gly 515 520 525 Gly Gly Ser Gly Gly Gly Gly Ser
Val Val Asp Arg Asn Pro Gln Phe 530 535 540 Leu Asp Pro Val Leu Ala
Tyr Leu Met Lys Gly Leu Cys Glu Lys Pro 545 550 555 560 Leu Ala Ser
Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Ser Ser Pro 565 570 575 Asp
Glu Val Ala Leu Val Glu Gly Val Gln Ser Leu Gly Phe Thr Tyr 580 585
590 Leu Arg Leu Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser Gly Gly Gly
595 600 605 Gly Ser Glu Phe Lys His Ile Lys Ala Phe Asp Arg Thr Phe
Ala Asn 610 615 620 Asn Pro Gly Pro Met Val Val Phe Ala Thr Pro Gly
Met Gly Gly Gly 625 630 635 640 Gly Ser Gly Gly Gly Gly Ser Ser Thr
Ala Asn Tyr Asn Thr Ser His 645 650 655 Leu Asn Asn Asp Val Trp Gln
Ile Phe Glu Asn Pro Val Asp Trp Lys 660 665 670 Glu Lys Gly Gly Gly
Gly Ser Gly Gly Gly Gly Ser Asp Ser Gly Ser 675 680 685 Pro Phe Pro
Ala Ala Val Ile Leu Arg Asp Ala Leu His Met Ala Arg 690 695 700 Gly
Leu Lys Tyr Leu His Gln Gly Gly Gly Gly Ser Gly Gly Gly Gly 705 710
715 720 Ser Ala Asn Phe Glu Ser Gly Lys His Lys Tyr Arg Gln Thr Ala
Met 725 730 735 Phe Thr Ala Thr Met Pro Pro Ala Val Glu Arg Leu Gly
Gly Gly Gly 740 745 750 Ser Gly Gly Gly Gly Ser Asn His Ser Gly Leu
Val Thr Phe Gln Ala 755 760 765 Phe Ile Asp Val Met Ser Arg Glu Thr
Thr Asp Thr Asp Thr Ala Asp 770 775 780 Gln Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser Cys Gly Thr Ala Phe 785 790 795 800 Phe Ile Asn Phe
Ile Ala Ile Tyr His His Ala Ser Arg Ala Ile Pro 805 810 815 Phe Gly
Thr Met Val Ala Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser 820 825 830
Phe Val Val Lys Ala Tyr Leu Pro Val Asn Glu Ser Phe Ala Phe Thr 835
840 845 Ala Asp Leu Arg Ser Asn Thr Gly Gly Gln Ala Gly Gly Gly Gly
Ser 850 855 860 Gly Gly Gly Gly Ser Thr Pro Pro Pro Glu Glu Ala Met
Pro Phe Glu 865 870 875 880 Phe Asn Gly Pro Ala Gln Gly Asp His Ser
Gln Pro Pro Leu Gln Val 885 890 895 471081PRTArtificial SequenceVB
vector 47Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu Cys Thr
Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu Ala Ala
Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln
Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr Ser Ser
Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys Arg Gly
Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val Gln Lys
Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95 Thr Pro
Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100 105 110
Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly Gly 115
120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro
Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr
Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val
Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr Asn Thr
Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe Leu Tyr
Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln Gly Asn
Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215 220 Asn Arg
Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225 230 235
240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu Tyr Met Leu
245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr Gly Gln
Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp Gly Pro
Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr Asn Ile Val
Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu Cys Val
Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu Glu Asp Leu
Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys Ser Gln
Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Pro 340 345 350 Ser
Lys Pro Ser Phe Gln Glu Phe Val Asp Trp Glu Asn Val Ser Pro 355 360
365 Glu Leu Asn Ser Thr Asp Gln Pro Phe Leu Gly Gly Gly Gly Ser Gly
370 375 380 Gly Gly Gly Ser Arg Glu Gly Val Glu Leu Cys Pro Gly Asn
Lys Tyr 385 390 395 400 Glu Met Arg Arg His Gly Thr Thr His Ser Leu
Val Ile His Asp Gly 405 410 415 Gly Gly Gly Ser Gly Gly Gly Gly Ser
Ser His Cys His Trp Asn Asp 420 425 430 Leu Ala Val Ile Pro Ala Gly
Val Val His Asn Trp Asp Phe Glu Pro 435 440 445 Arg Lys Val Ser Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly Arg 450 455 460 Gly His Leu
Leu Gly Arg Leu Ala Ala Ile Val Gly Lys Gln Val Leu 465 470 475 480
Leu Gly Arg Lys Val Val Val Val Arg Gly Gly Gly Gly Ser Gly Gly 485
490 495 Gly Gly Ser Phe Arg Arg Lys Ala Phe Leu His Trp Tyr Thr Gly
Glu 500 505 510 Ala Met Asp Glu Met Glu Phe Thr Glu Ala Glu Ser Asn
Met Gly Gly 515 520 525 Gly Gly Ser Gly Gly Gly Gly Ser Val Val Asp
Arg Asn Pro Gln Phe 530 535 540 Leu Asp Pro Val Leu Ala Tyr Leu Met
Lys Gly Leu Cys Glu Lys Pro 545 550 555 560 Leu Ala Ser Gly Gly Gly
Gly Ser Gly Gly Gly Gly Ser Ser Ser Pro 565 570 575 Asp Glu Val Ala
Leu Val Glu Gly Val Gln Ser Leu Gly Phe Thr Tyr 580 585 590 Leu Arg
Leu Lys Asp Asn Tyr Met Gly Gly Gly Gly Ser Gly Gly Gly 595 600 605
Gly Ser Glu Phe Lys His Ile Lys Ala Phe Asp Arg Thr Phe Ala Asn 610
615 620 Asn Pro Gly Pro Met Val Val Phe Ala Thr Pro Gly Met Gly Gly
Gly 625 630 635 640 Gly Ser Gly Gly Gly Gly Ser Ser Thr Ala Asn Tyr
Asn Thr Ser His 645 650 655 Leu Asn Asn Asp Val Trp Gln Ile Phe Glu
Asn Pro Val Asp Trp Lys 660 665 670 Glu Lys Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser Asp Ser Gly Ser 675 680 685 Pro Phe Pro Ala Ala Val
Ile Leu Arg Asp Ala Leu His Met Ala Arg 690 695 700 Gly Leu Lys Tyr
Leu His Gln Gly Gly Gly Gly Ser Gly Gly Gly Gly 705 710 715 720 Ser
Ala Asn Phe Glu Ser Gly Lys His Lys Tyr Arg Gln Thr Ala Met 725 730
735 Phe Thr Ala Thr Met Pro Pro Ala Val Glu Arg Leu Gly Gly Gly Gly
740 745 750 Ser Gly Gly Gly Gly Ser Asn His Ser Gly Leu Val Thr Phe
Gln Ala 755 760 765 Phe Ile Asp Val Met Ser Arg Glu Thr Thr Asp Thr
Asp Thr Ala Asp 770 775 780 Gln Gly Gly Gly Gly Ser Gly Gly Gly Gly
Ser Cys Gly Thr Ala Phe 785 790 795 800 Phe Ile Asn Phe Ile Ala Ile
Tyr His His Ala Ser Arg Ala Ile Pro 805 810 815 Phe Gly Thr Met Val
Ala Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser 820 825 830 Phe Val Val
Lys Ala Tyr Leu Pro Val Asn Glu Ser Phe Ala Phe Thr 835 840 845 Ala
Asp Leu Arg Ser Asn Thr Gly Gly Gln Ala Gly Gly Gly Gly Ser 850 855
860 Gly Gly Gly Gly Ser Thr Pro Pro Pro Glu Glu Ala Met Pro Phe Glu
865 870 875 880 Phe Asn Gly Pro Ala Gln Gly Asp His Ser Gln Pro Pro
Leu Gln Val 885 890 895 Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Pro
Lys Pro Asp Phe Ser 900 905 910 Gln Leu Gln Arg Asn Ile Leu Pro Ser
Asn Pro Arg Val Thr Arg Phe 915 920 925 His Ile Asn Trp Asp Gly Gly
Gly Gly Ser Gly Gly Gly Gly Ser Ile 930 935 940 Pro Ser Gly Thr Thr
Ile Leu Asn Cys Phe His Asp Val Leu Ser Gly 945 950 955 960 Lys Leu
Ser Gly Gly Ser Pro Gly Val Pro Gly Gly Gly Gly Ser Gly 965 970 975
Gly Gly Gly Ser Gly Phe Ser Gln Pro Leu Arg Arg Leu Val Leu His 980
985 990 Val Val Ser Ala Ala Gln Ala Glu Arg Leu Ala Arg Ala Glu Glu
Gly 995 1000
1005 Gly Gly Gly Ser Gly Gly Gly Gly Ser Glu Cys Arg Ile Thr Ser
1010 1015 1020 Asn Phe Val Ile Pro Ser Glu Tyr Trp Val Glu Glu Lys
Glu Glu 1025 1030 1035 Lys Gln Lys Leu Ile Gln Gly Gly Gly Gly Ser
Gly Gly Gly Gly 1040 1045 1050 Ser Asn Ile Glu Gly Ile Asp Lys Leu
Thr Gln Leu Lys Lys Pro 1055 1060 1065 Phe Leu Val Asn Asn Lys Ile
Asn Lys Ile Glu Asn Ile 1070 1075 1080 48896PRTArtificial
SequenceVB construct 48Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu
Leu Cys Thr Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro
Leu Ala Ala Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr
Ser Arg Gln Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu
Thr Ser Ser Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr
Lys Arg Gly Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp
Val Gln Lys Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90
95 Thr Pro Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp
100 105 110 Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser
Gly Gly 115 120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu Pro Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn
Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe
Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln
Gln Gly Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215
220 Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu
225 230 235 240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu
Tyr Met Leu 245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly
Tyr Gly Gln Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile
Asp Gly Pro Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr
Asn Ile Val Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg
Leu Cys Val Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu
Glu Asp Leu Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335
Cys Ser Gln Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Val 340
345 350 Ile Leu Pro Gln Ala Pro Ser Gly Pro Ser Tyr Ala Thr Tyr Leu
Gln 355 360 365 Pro Ala Gln Ala Gln Met Leu Thr Pro Pro Gly Gly Gly
Gly Ser Gly 370 375 380 Gly Gly Gly Ser Leu His Ser Gly Gln Asn His
Leu Lys Glu Met Ala 385 390 395 400 Ile Ser Val Leu Glu Ala Arg Ala
Cys Ala Ala Ala Gly Gln Ser Gly 405 410 415 Gly Gly Gly Ser Gly Gly
Gly Gly Ser Pro Leu Leu Pro Phe Tyr Pro 420 425 430 Pro Asp Glu Ala
Leu Glu Ile Gly Leu Glu Leu Asn Ser Ser Ala Leu 435 440 445 Pro Pro
Thr Glu Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Ala Gly 450 455 460
Thr Gln Cys Glu Tyr Trp Ala Ser Arg Ala Leu Asp Ser Glu His Ser 465
470 475 480 Ile Gly Ser Met Ile Gln Leu Pro Gln Gly Gly Gly Gly Ser
Gly Gly 485 490 495 Gly Gly Ser Ala Ala Tyr Lys Gly His His Tyr Pro
Gly Pro Gly Asn 500 505 510 Tyr Phe Trp Lys Cys Leu Phe Met Ser Gly
Leu Ser Glu Val Gly Gly 515 520 525 Gly Gly Ser Gly Gly Gly Gly Ser
Asp Thr Leu Ser Ala Met Ser Asn 530 535 540 Pro Arg Ala Met Gln Val
Leu Leu Gln Ile Gln Gln Gly Leu Gln Thr 545 550 555 560 Leu Ala Thr
Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Asp Lys Pro 565 570 575 Leu
Arg Arg Asn Asn Ser Tyr Thr Ser Tyr Ile Met Ala Ile Cys Gly 580 585
590 Met Pro Leu Asp Ser Phe Arg Ala Gly Gly Gly Gly Ser Gly Gly Gly
595 600 605 Gly Ser Glu Val Ile Gln Thr Ser Lys Tyr Tyr Met Arg Asp
Val Ile 610 615 620 Ala Ile Glu Ser Ala Trp Leu Leu Glu Leu Ala Pro
His Gly Gly Gly 625 630 635 640 Gly Ser Gly Gly Gly Gly Ser Gly Tyr
Ile Ser Arg Val Thr Ala Gly 645 650 655 Lys Asp Ser Tyr Ile Ala Leu
Val Asp Lys Asn Ile Met Gly Tyr Ile 660 665 670 Ala Ser Gly Gly Gly
Gly Ser Gly Gly Gly Gly Ser Glu His Ile His 675 680 685 Arg Ala Gly
Gly Leu Phe Val Ala Asp Ala Ile Gln Val Gly Phe Gly 690 695 700 Arg
Ile Gly Lys His Phe Trp Gly Gly Gly Gly Ser Gly Gly Gly Gly 705 710
715 720 Ser Gln Ala Ile Val Arg Gly Cys Ser Met Pro Gly Pro Trp Arg
Ser 725 730 735 Gly Arg Leu Leu Val Ser Arg Arg Trp Ser Val Glu Gly
Gly Gly Gly 740 745 750 Ser Gly Gly Gly Gly Ser Asp Gly Gln Leu Glu
Leu Leu Ala Gln Gly 755 760 765 Ala Leu Asp Asn Ala Leu Ser Ser Met
Gly Ala Leu His Ala Leu Arg 770 775 780 Pro Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser Ser His Asp Ser Arg 785 790 795 800 Lys Ser Thr Ser
Phe Met Ser Val Asn Pro Ser Lys Glu Ile Lys Ile 805 810 815 Val Ser
Ala Val Arg Arg Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser 820 825 830
His Thr Pro Ser Ser Tyr Ile Glu Thr Leu Pro Lys Ala Ile Lys Arg 835
840 845 Arg Ile Asn Ala Leu Lys Gln Leu Gln Val Arg Gly Gly Gly Gly
Ser 850 855 860 Gly Gly Gly Gly Ser Met Lys Ala Phe Ile Phe Lys Tyr
Ser Ala Lys 865 870 875 880 Thr Gly Phe Thr Lys Leu Ile Asp Ala Ser
Arg Val Ser Glu Thr Glu 885 890 895 491081PRTArtificial SequenceVB
construct 49Met Gln Val Ser Thr Ala Ala Leu Ala Val Leu Leu Cys Thr
Met Ala 1 5 10 15 Leu Cys Asn Gln Val Leu Ser Ala Pro Leu Ala Ala
Asp Thr Pro Thr 20 25 30 Ala Cys Cys Phe Ser Tyr Thr Ser Arg Gln
Ile Pro Gln Asn Phe Ile 35 40 45 Ala Asp Tyr Phe Glu Thr Ser Ser
Gln Cys Ser Lys Pro Ser Val Ile 50 55 60 Phe Leu Thr Lys Arg Gly
Arg Gln Val Cys Ala Asp Pro Ser Glu Glu 65 70 75 80 Trp Val Gln Lys
Tyr Val Ser Asp Leu Glu Leu Ser Ala Glu Leu Lys 85 90 95 Thr Pro
Leu Gly Asp Thr Thr His Thr Ile Glu Pro Lys Ser Cys Asp 100 105 110
Thr Pro Pro Pro Cys Pro Arg Cys Pro Gly Gly Gly Ser Ser Gly Gly 115
120 125 Gly Ser Gly Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro
Pro 130 135 140 Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr
Cys Leu Val 145 150 155 160 Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val
Glu Trp Glu Ser Ser Gly 165 170 175 Gln Pro Glu Asn Asn Tyr Asn Thr
Thr Pro Pro Met Leu Asp Ser Asp 180 185 190 Gly Ser Phe Phe Leu Tyr
Ser Lys Leu Thr Val Asp Lys Ser Arg Trp 195 200 205 Gln Gln Gly Asn
Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His 210 215 220 Asn Arg
Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys Gly Leu 225 230 235
240 Gly Gly Leu Met His Gly Asp Thr Pro Thr Leu His Glu Tyr Met Leu
245 250 255 Asp Leu Gln Pro Glu Thr Thr Asp Leu Tyr Gly Tyr Gly Gln
Leu Asn 260 265 270 Asp Ser Ser Glu Glu Glu Asp Glu Ile Asp Gly Pro
Ala Gly Gln Ala 275 280 285 Glu Pro Asp Arg Ala His Tyr Asn Ile Val
Thr Phe Cys Cys Lys Cys 290 295 300 Asp Ser Thr Leu Arg Leu Cys Val
Gln Ser Thr His Val Asp Ile Arg 305 310 315 320 Thr Leu Glu Asp Leu
Leu Met Gly Thr Leu Gly Ile Val Cys Pro Ile 325 330 335 Cys Ser Gln
Lys Pro Gly Gly Gly Ser Ser Gly Gly Gly Ser Gly Val 340 345 350 Ile
Leu Pro Gln Ala Pro Ser Gly Pro Ser Tyr Ala Thr Tyr Leu Gln 355 360
365 Pro Ala Gln Ala Gln Met Leu Thr Pro Pro Gly Gly Gly Gly Ser Gly
370 375 380 Gly Gly Gly Ser Leu His Ser Gly Gln Asn His Leu Lys Glu
Met Ala 385 390 395 400 Ile Ser Val Leu Glu Ala Arg Ala Cys Ala Ala
Ala Gly Gln Ser Gly 405 410 415 Gly Gly Gly Ser Gly Gly Gly Gly Ser
Pro Leu Leu Pro Phe Tyr Pro 420 425 430 Pro Asp Glu Ala Leu Glu Ile
Gly Leu Glu Leu Asn Ser Ser Ala Leu 435 440 445 Pro Pro Thr Glu Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser Ala Gly 450 455 460 Thr Gln Cys
Glu Tyr Trp Ala Ser Arg Ala Leu Asp Ser Glu His Ser 465 470 475 480
Ile Gly Ser Met Ile Gln Leu Pro Gln Gly Gly Gly Gly Ser Gly Gly 485
490 495 Gly Gly Ser Ala Ala Tyr Lys Gly His His Tyr Pro Gly Pro Gly
Asn 500 505 510 Tyr Phe Trp Lys Cys Leu Phe Met Ser Gly Leu Ser Glu
Val Gly Gly 515 520 525 Gly Gly Ser Gly Gly Gly Gly Ser Asp Thr Leu
Ser Ala Met Ser Asn 530 535 540 Pro Arg Ala Met Gln Val Leu Leu Gln
Ile Gln Gln Gly Leu Gln Thr 545 550 555 560 Leu Ala Thr Gly Gly Gly
Gly Ser Gly Gly Gly Gly Ser Asp Lys Pro 565 570 575 Leu Arg Arg Asn
Asn Ser Tyr Thr Ser Tyr Ile Met Ala Ile Cys Gly 580 585 590 Met Pro
Leu Asp Ser Phe Arg Ala Gly Gly Gly Gly Ser Gly Gly Gly 595 600 605
Gly Ser Glu Val Ile Gln Thr Ser Lys Tyr Tyr Met Arg Asp Val Ile 610
615 620 Ala Ile Glu Ser Ala Trp Leu Leu Glu Leu Ala Pro His Gly Gly
Gly 625 630 635 640 Gly Ser Gly Gly Gly Gly Ser Gly Tyr Ile Ser Arg
Val Thr Ala Gly 645 650 655 Lys Asp Ser Tyr Ile Ala Leu Val Asp Lys
Asn Ile Met Gly Tyr Ile 660 665 670 Ala Ser Gly Gly Gly Gly Ser Gly
Gly Gly Gly Ser Glu His Ile His 675 680 685 Arg Ala Gly Gly Leu Phe
Val Ala Asp Ala Ile Gln Val Gly Phe Gly 690 695 700 Arg Ile Gly Lys
His Phe Trp Gly Gly Gly Gly Ser Gly Gly Gly Gly 705 710 715 720 Ser
Gln Ala Ile Val Arg Gly Cys Ser Met Pro Gly Pro Trp Arg Ser 725 730
735 Gly Arg Leu Leu Val Ser Arg Arg Trp Ser Val Glu Gly Gly Gly Gly
740 745 750 Ser Gly Gly Gly Gly Ser Asp Gly Gln Leu Glu Leu Leu Ala
Gln Gly 755 760 765 Ala Leu Asp Asn Ala Leu Ser Ser Met Gly Ala Leu
His Ala Leu Arg 770 775 780 Pro Gly Gly Gly Gly Ser Gly Gly Gly Gly
Ser Ser His Asp Ser Arg 785 790 795 800 Lys Ser Thr Ser Phe Met Ser
Val Asn Pro Ser Lys Glu Ile Lys Ile 805 810 815 Val Ser Ala Val Arg
Arg Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser 820 825 830 His Thr Pro
Ser Ser Tyr Ile Glu Thr Leu Pro Lys Ala Ile Lys Arg 835 840 845 Arg
Ile Asn Ala Leu Lys Gln Leu Gln Val Arg Gly Gly Gly Gly Ser 850 855
860 Gly Gly Gly Gly Ser Met Lys Ala Phe Ile Phe Lys Tyr Ser Ala Lys
865 870 875 880 Thr Gly Phe Thr Lys Leu Ile Asp Ala Ser Arg Val Ser
Glu Thr Glu 885 890 895 Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Glu
Gly Asp Pro Cys Leu 900 905 910 Arg Ser Ser Asp Cys Ile Asp Glu Phe
Cys Cys Ala Arg His Phe Trp 915 920 925 Thr Lys Ile Cys Lys Gly Gly
Gly Gly Ser Gly Gly Gly Gly Ser Trp 930 935 940 Lys Gly Gly Pro Val
Lys Ile Asp Pro Leu Ala Leu Met Gln Ala Ile 945 950 955 960 Glu Arg
Tyr Leu Val Val Arg Gly Tyr Gly Gly Gly Gly Gly Ser Gly 965 970 975
Gly Gly Gly Ser Val Thr Ser Ile Pro Ser Val Ser Asn Ala Leu Asn 980
985 990 Trp Lys Glu Phe Ser Phe Ile Gln Ser Thr Leu Gly Tyr Val Ala
Gly 995 1000 1005 Gly Gly Gly Ser Gly Gly Gly Gly Ser Tyr Arg Gly
Ala Asn Leu 1010 1015 1020 His Leu Glu Glu Thr Leu Ala Gly Phe Trp
Ala Arg Leu Leu Glu 1025 1030 1035 Arg Leu Phe Lys Gln Leu Gly Gly
Gly Gly Ser Gly Gly Gly Gly 1040 1045 1050 Ser Lys Thr Thr Leu Ser
His Thr Gln Asp Ser Ser Gln Ser Leu 1055 1060 1065 Gln Ser Ser Ser
Asp Ser Ser Lys Ser Ser Arg Cys Ser 1070 1075 1080 5027PRTMus
musculus 50Gln Ala Ile Val Arg Gly Cys Ser Met Pro Gly Pro Trp Arg
Ser Gly 1 5 10 15 Arg Leu Leu Val Ser Arg Arg Trp Ser Val Glu 20 25
5127PRTMus musculus 51Asp Gly Gln Leu Glu Leu Leu Ala Gln Gly Ala
Leu Asp Asn Ala Leu 1 5 10 15 Ser Ser Met Gly Ala Leu His Ala Leu
Arg Pro 20 25 5227PRTMus musculus 52Ser His Asp Ser Arg Lys Ser Thr
Ser Phe Met Ser Val Asn Pro Ser 1 5 10 15 Lys Glu Ile Lys Ile Val
Ser Ala Val Arg Arg 20 25 5327PRTMus musculus 53His Thr Pro Ser Ser
Tyr Ile Glu Thr Leu Pro Lys Ala Ile Lys Arg 1 5 10 15 Arg Ile Asn
Ala Leu Lys Gln Leu Gln Val Arg 20 25 5427PRTMus musculus 54Met Lys
Ala Phe Ile Phe Lys Tyr Ser Ala Lys Thr Gly Phe Thr Lys 1 5 10 15
Leu Ile Asp Ala Ser Arg Val Ser Glu Thr Glu 20 25 5527PRTMus
musculus 55Glu Gly Asp Pro Cys Leu Arg Ser Ser Asp Cys Ile Asp Glu
Phe Cys 1 5 10 15 Cys Ala Arg His Phe Trp Thr Lys Ile Cys Lys 20 25
5627PRTMus musculus 56Trp Lys Gly Gly Pro Val Lys Ile Asp Pro Leu
Ala Leu Met Gln Ala 1 5 10 15 Ile Glu Arg Tyr Leu Val Val Arg Gly
Tyr Gly 20 25 5727PRTMus musculus 57Val Thr Ser Ile Pro Ser Val Ser
Asn Ala Leu Asn Trp Lys Glu Phe 1 5 10 15 Ser Phe Ile Gln Ser Thr
Leu Gly Tyr Val
Ala 20 25 5827PRTMus musculus 58Tyr Arg Gly Ala Asn Leu His Leu Glu
Glu Thr Leu Ala Gly Phe Trp 1 5 10 15 Ala Arg Leu Leu Glu Arg Leu
Phe Lys Gln Leu 20 25 5927PRTMus musculus 59Lys Thr Thr Leu Ser His
Thr Gln Asp Ser Ser Gln Ser Leu Gln Ser 1 5 10 15 Ser Ser Asp Ser
Ser Lys Ser Ser Arg Cys Ser 20 25 6027PRTMus musculus 60Asn His Ser
Gly Leu Val Thr Phe Gln Ala Phe Ile Asp Val Met Ser 1 5 10 15 Arg
Glu Thr Thr Asp Thr Asp Thr Ala Asp Gln 20 25 6127PRTMus musculus
61Cys Gly Thr Ala Phe Phe Ile Asn Phe Ile Ala Ile Tyr His His Ala 1
5 10 15 Ser Arg Ala Ile Pro Phe Gly Thr Met Val Ala 20 25
6227PRTMus musculus 62Phe Val Val Lys Ala Tyr Leu Pro Val Asn Glu
Ser Phe Ala Phe Thr 1 5 10 15 Ala Asp Leu Arg Ser Asn Thr Gly Gly
Gln Ala 20 25 6327PRTMus musculus 63Thr Pro Pro Pro Glu Glu Ala Met
Pro Phe Glu Phe Asn Gly Pro Ala 1 5 10 15 Gln Gly Asp His Ser Gln
Pro Pro Leu Gln Val 20 25 6427PRTMus musculus 64Pro Lys Pro Asp Phe
Ser Gln Leu Gln Arg Asn Ile Leu Pro Ser Asn 1 5 10 15 Pro Arg Val
Thr Arg Phe His Ile Asn Trp Asp 20 25 6527PRTMus musculus 65Ile Pro
Ser Gly Thr Thr Ile Leu Asn Cys Phe His Asp Val Leu Ser 1 5 10 15
Gly Lys Leu Ser Gly Gly Ser Pro Gly Val Pro 20 25 6627PRTMus
musculus 66Gly Phe Ser Gln Pro Leu Arg Arg Leu Val Leu His Val Val
Ser Ala 1 5 10 15 Ala Gln Ala Glu Arg Leu Ala Arg Ala Glu Glu 20 25
6727PRTMus musculus 67Glu Cys Arg Ile Thr Ser Asn Phe Val Ile Pro
Ser Glu Tyr Trp Val 1 5 10 15 Glu Glu Lys Glu Glu Lys Gln Lys Leu
Ile Gln 20 25 6827PRTMus musculus 68Asn Ile Glu Gly Ile Asp Lys Leu
Thr Gln Leu Lys Lys Pro Phe Leu 1 5 10 15 Val Asn Asn Lys Ile Asn
Lys Ile Glu Asn Ile 20 25 695PRTArtificial SequenceLinker 69Gly Gly
Gly Ser Ser 1 5 705PRTArtificial SequenceLinker 70Gly Gly Gly Ser
Gly 1 5 715PRTArtificial SequenceLinker 71Gly Gly Gly Gly Ser 1 5
725PRTArtificial SequenceLinker 72Leu Gly Gly Gly Ser 1 5
735PRTArtificial SequenceLinker 73Gly Leu Gly Gly Ser 1 5
745PRTArtificial SequenceLinker 74Gly Gly Leu Gly Ser 1 5
755PRTArtificial SequenceLinker 75Gly Gly Gly Leu Ser 1 5
765PRTArtificial SequenceLinker 76Gly Gly Gly Gly Leu 1 5
775PRTArtificial SequenceLinker 77Leu Gly Gly Ser Gly 1 5
785PRTArtificial SequenceLinker 78Gly Leu Gly Ser Gly 1 5
795PRTArtificial SequenceLinker 79Gly Gly Leu Ser Gly 1 5
805PRTArtificial SequenceLinker 80Gly Gly Gly Leu Gly 1 5
815PRTArtificial SequenceLinker 81Gly Gly Gly Ser Leu 1 5
825PRTArtificial SequenceLinker 82Leu Gly Gly Ser Ser 1 5
835PRTArtificial SequenceLinker 83Gly Leu Gly Ser Ser 1 5
845PRTArtificial SequenceLinker 84Gly Gly Leu Ser Ser 1 5
855PRTArtificial SequenceLinker 85Gly Gly Gly Leu Ser 1 5
865PRTArtificial SequenceLinker 86Gly Gly Gly Ser Leu 1 5
875PRTArtificial SequenceLinker 87Leu Gly Leu Gly Ser 1 5
885PRTArtificial SequenceLinker 88Gly Leu Gly Leu Ser 1 5
895PRTArtificial SequenceLinker 89Gly Leu Leu Gly Ser 1 5
905PRTArtificial SequenceLinker 90Leu Gly Gly Leu Ser 1 5
915PRTArtificial SequenceLinker 91Gly Leu Gly Gly Leu 1 5
925PRTArtificial SequenceLinker 92Leu Gly Leu Ser Gly 1 5
935PRTArtificial SequenceLinker 93Gly Leu Leu Ser Gly 1 5
945PRTArtificial SequenceLinker 94Gly Gly Leu Ser Leu 1 5
955PRTArtificial SequenceLinker 95Gly Gly Leu Leu Gly 1 5
965PRTArtificial SequenceLinker 96Gly Leu Gly Ser Leu 1 5
975PRTArtificial SequenceLinker 97Leu Gly Leu Ser Ser 1 5
985PRTArtificial SequenceLinker 98Gly Leu Gly Leu Ser 1 5
995PRTArtificial SequenceLinker 99Gly Gly Leu Leu Ser 1 5
1005PRTArtificial SequenceLinker 100Gly Leu Gly Ser Leu 1 5
1015PRTArtificial SequenceLinker 101Gly Leu Gly Ser Leu 1 5
10210PRTArtificial SequenceLinker 102Leu Gly Gly Gly Ser Gly Gly
Gly Gly Ser 1 5 10 10310PRTArtificial SequenceLinker 103Gly Leu Gly
Gly Ser Gly Gly Gly Gly Ser 1 5 10 10410PRTArtificial
SequenceLinker 104Gly Gly Leu Gly Ser Gly Gly Gly Gly Ser 1 5 10
10510PRTArtificial SequenceLinker 105Gly Gly Gly Leu Ser Gly Gly
Gly Gly Ser 1 5 10 10610PRTArtificial SequenceLinker 106Gly Gly Gly
Gly Leu Gly Gly Gly Gly Ser 1 5 10 10710PRTArtificial
SequenceLinker 107Leu Gly Gly Ser Gly Gly Gly Gly Ser Gly 1 5 10
10810PRTArtificial SequenceLinker 108Gly Leu Gly Ser Gly Gly Gly
Gly Ser Gly 1 5 10 10910PRTArtificial SequenceLinker 109Gly Gly Leu
Ser Gly Gly Gly Gly Ser Gly 1 5 10 11010PRTArtificial
SequenceLinker 110Gly Gly Gly Leu Gly Gly Gly Gly Ser Gly 1 5 10
11110PRTArtificial SequenceLinker 111Gly Gly Gly Ser Leu Gly Gly
Gly Ser Gly 1 5 10 11210PRTArtificial SequenceLinker 112Gly Gly Gly
Ser Leu Gly Gly Gly Ser Gly 1 5 10 11310PRTArtificial
SequenceLinker 113Gly Leu Gly Ser Ser Gly Gly Gly Ser Ser 1 5 10
11410PRTArtificial SequenceLinker 114Gly Gly Leu Ser Ser Gly Gly
Gly Ser Ser 1 5 10 11510PRTArtificial SequenceLinker 115Gly Gly Gly
Leu Ser Gly Gly Gly Ser Ser 1 5 10 11610PRTArtificial
SequenceLinker 116Gly Gly Gly Ser Leu Gly Gly Gly Ser Ser 1 5 10
11710PRTArtificial SequenceLinker 117Leu Gly Gly Gly Ser Leu Gly
Gly Gly Ser 1 5 10 11810PRTArtificial SequenceLinker 118Gly Leu Gly
Gly Ser Gly Leu Gly Gly Ser 1 5 10 11910PRTArtificial
SequenceLinker 119Gly Gly Leu Gly Ser Gly Gly Leu Gly Ser 1 5 10
12010PRTArtificial SequenceLinker 120Gly Gly Gly Leu Ser Gly Gly
Gly Leu Ser 1 5 10 12110PRTArtificial SequenceLinker 121Gly Gly Gly
Gly Leu Gly Gly Gly Gly Leu 1 5 10 12210PRTArtificial
SequenceLinker 122Leu Gly Gly Ser Gly Leu Gly Gly Ser Gly 1 5 10
12310PRTArtificial SequenceLinker 123Gly Leu Gly Ser Gly Gly Leu
Gly Ser Gly 1 5 10 12410PRTArtificial SequenceLinker 124Gly Gly Leu
Ser Gly Gly Gly Leu Ser Gly 1 5 10 12510PRTArtificial
SequenceLinker 125Gly Gly Gly Leu Gly Gly Gly Gly Leu Gly 1 5 10
12610PRTArtificial SequenceLinker 126Gly Gly Gly Ser Leu Gly Gly
Gly Ser Leu 1 5 10 12710PRTArtificial SequenceLinker 127Leu Gly Gly
Ser Ser Leu Gly Gly Ser Ser 1 5 10 12810PRTArtificial
SequenceLinker 128Gly Leu Gly Ser Ser Gly Leu Gly Ser Ser 1 5 10
12910PRTArtificial SequenceLinker 129Gly Gly Leu Ser Ser Gly Gly
Leu Ser Ser 1 5 10 13010PRTArtificial SequenceLinker 130Gly Gly Gly
Leu Ser Gly Gly Gly Leu Ser 1 5 10 13110PRTArtificial
SequenceLinker 131Gly Gly Gly Ser Leu Gly Gly Gly Ser Leu 1 5
10
References
D00000

D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012
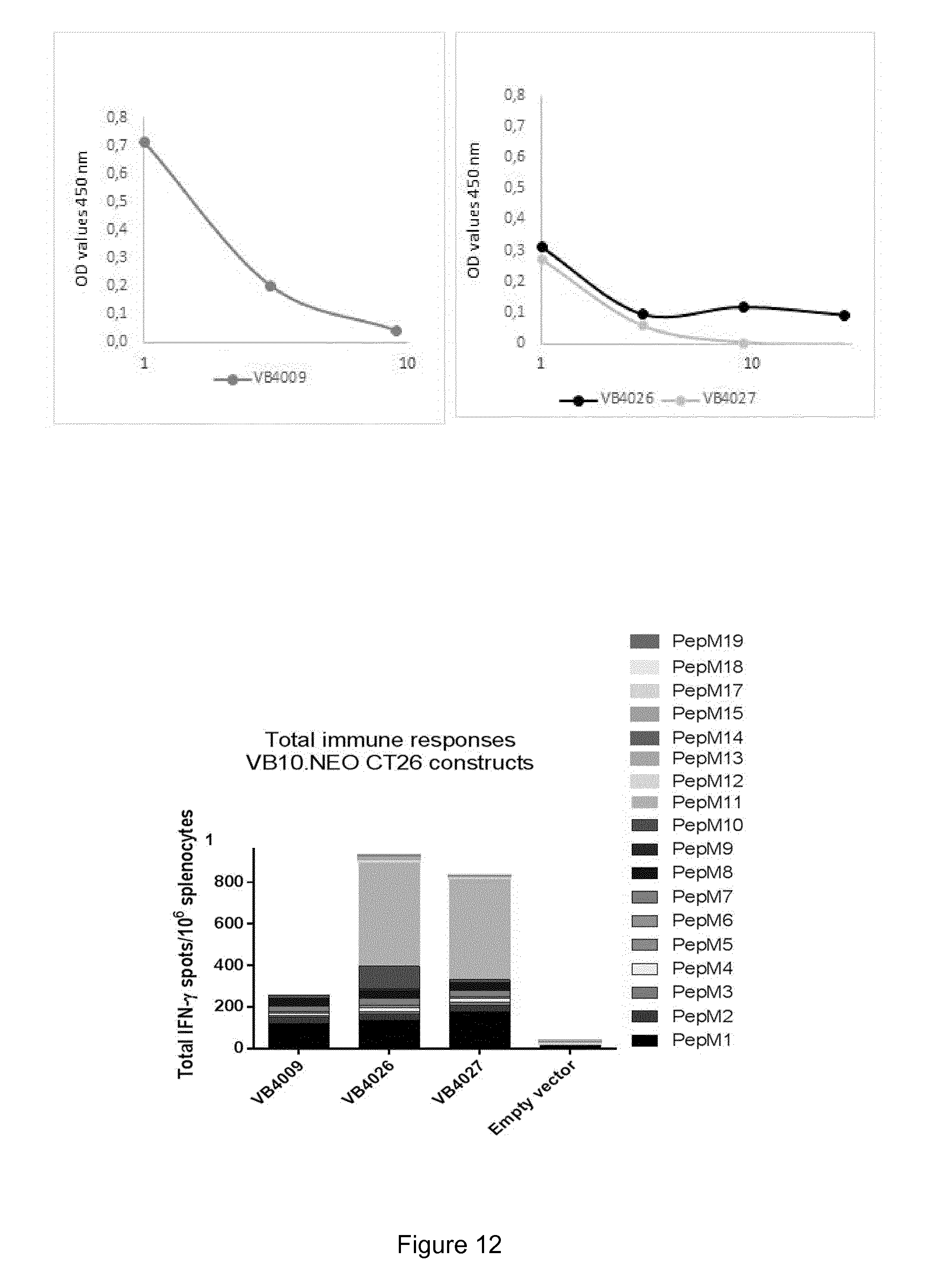
D00013

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.