Computationally Optimized Broadly Reactive Antigens For Influenza
Ross; Ted M. ; et al.
U.S. patent application number 16/128917 was filed with the patent office on 2019-01-10 for computationally optimized broadly reactive antigens for influenza. This patent application is currently assigned to University of Pittsburgh - Of the Commonwealth System of Higher Education. The applicant listed for this patent is University of Pittsburgh -Of the Commonwealth System of Higher Education. Invention is credited to Brendan M. Giles, Ted M. Ross.
| Application Number | 20190008949 16/128917 |
| Document ID | / |
| Family ID | 45831918 |
| Filed Date | 2019-01-10 |








View All Diagrams
| United States Patent Application | 20190008949 |
| Kind Code | A1 |
| Ross; Ted M. ; et al. | January 10, 2019 |
COMPUTATIONALLY OPTIMIZED BROADLY REACTIVE ANTIGENS FOR INFLUENZA
Abstract
The development of a computationally optimized influenza HA protein that elicits broadly reactive immune response to all H5N1 influenza virus isolates is described. The optimized HA protein was developed through a series of HA protein alignments, and subsequent generation of consensus sequences, for clade 2 H5N1 influenza virus isolates. The final consensus HA amino acid sequence was reverse translated and optimized for expression in mammalian cells. Influenza virus-like particles containing the optimized HA protein are an effective vaccine against H5N1 influenza virus infection in animals.
| Inventors: | Ross; Ted M.; (Athens, GA) ; Giles; Brendan M.; (Denver, CO) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | University of Pittsburgh - Of the
Commonwealth System of Higher Education Pittsburgh PA |
||||||||||
| Family ID: | 45831918 | ||||||||||
| Appl. No.: | 16/128917 | ||||||||||
| Filed: | September 12, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15463960 | Mar 20, 2017 | 10098946 | ||
| 16128917 | ||||
| 14511930 | Oct 10, 2014 | |||
| 15463960 | ||||
| 13822844 | Mar 13, 2013 | 8883171 | ||
| PCT/US2011/051072 | Sep 9, 2011 | |||
| 14511930 | ||||
| 61403407 | Sep 14, 2010 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61K 2039/54 20130101; A61K 39/00 20130101; C12N 2760/16223 20130101; C12N 2760/16323 20130101; C12N 2760/16234 20130101; A61K 2039/575 20130101; C12Y 302/01018 20130101; A61K 39/39 20130101; A61K 2039/55505 20130101; C07K 14/11 20130101; C12N 2760/16143 20130101; C12N 2760/16171 20130101; C12N 2800/22 20130101; C12N 2760/16123 20130101; C12N 9/2402 20130101; A61K 39/12 20130101; A61K 2039/5258 20130101; C12N 2760/16134 20130101; A61K 39/145 20130101; C07K 2319/00 20130101; C12N 7/00 20130101; A61K 2039/552 20130101; C12N 2760/16122 20130101; C12N 2760/16334 20130101; C12N 15/86 20130101; A61K 2039/545 20130101; C07K 14/005 20130101 |
| International Class: | A61K 39/145 20060101 A61K039/145; C12N 9/24 20060101 C12N009/24; A61K 39/12 20060101 A61K039/12; A61K 39/39 20060101 A61K039/39; C07K 14/005 20060101 C07K014/005; C07K 14/11 20060101 C07K014/11; C12N 7/00 20060101 C12N007/00; C12N 15/86 20060101 C12N015/86 |
Claims
1. A method of eliciting a broadly reactive immune response against influenza virus in a subject, comprising generating an optimized influenza virus polypeptide sequence comprising the steps of: (i) obtaining the amino acid sequences of the polypeptide from a group of influenza virus isolates, wherein the influenza virus isolates are from the same subtype; (ii) organizing the amino acid sequences of the polypeptide from the group of influenza virus isolates by clade or sub-clade and then by geographical region within each clade or sub-clade; (iii) aligning the amino acid sequences within each geographical region to generate primary consensus sequences, wherein each geographic region is represented by a primary consensus sequence; (iv) aligning the primary consensus sequences to generate secondary consensus sequences, wherein each clade or sub-clade is represented by a secondary consensus sequence; and (v) aligning the secondary consensus sequences, thereby generating the optimized influenza virus polypeptide sequence; and administering the optimized influenza virus polypeptide to the subject.
2. The method of claim 1, wherein generating the optimized influenza virus polypeptide sequence further comprises: (vi) reverse translating the optimized influenza virus polypeptide sequence to generate a coding sequence; and (vii) optimizing the coding sequence for expression in mammalian cells.
3. The method of claim 1, comprising administering a virus-like particle (VLP) comprising the optimized influenza virus polypeptide sequence to the subject.
4. The method of claim 3, wherein the influenza virus is an H5N1 virus and the VLP elicits a broadly reactive immune response against H5N1 influenza.
5. The method of claim 4, wherein the VLP elicits a protective immune response against at least 80% of known H5N1 isolates.
6. The method of claim 1, wherein the geographical region is a continent.
7. The method of claim 1, wherein the geographical region is a country.
Description
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This is a continuation of U.S. application Ser. No. 15/463,960, filed Mar. 20, 2017, which is a divisional of U.S. application Ser. No. 14/511,930, filed Oct. 10, 2014, now abandoned, which is a divisional of U.S. application Ser. No. 13/822,844, filed Mar. 13, 2013, issued as U.S. Pat. No. 8,883,171 on Nov. 11, 2014, which is the U.S. National Stage of International Application No. PCT/US2011/051072, filed Sep. 9, 2011, published in English under PCT Article 21(2), which claims the benefit of U.S. Provisional Application No. 61/403,407, filed Sep. 14, 2010. The above-listed applications are herein incorporated by reference in their entirety.
FIELD
[0002] This disclosure concerns an optimized influenza hemagglutinin protein that elicits broadly reactive immune responses to H5N1 virus isolates and its use as a vaccine.
BACKGROUND
[0003] Influenza virus is a member of Orthomyxoviridae family. There are three subtypes of influenza viruses, designated influenza A, influenza B, and influenza C. The influenza virion contains a segmented negative-sense RNA genome, which encodes the following proteins: hemagglutinin (HA), neuraminidase (NA), matrix (M1), proton ion-channel protein (M2), nucleoprotein (NP), polymerase basic protein 1 (PB1), polymerase basic protein 2 (PB2), polymerase acidic protein (PA), and nonstructural protein 2 (NS2). The HA, NA, M1, and M2 are membrane associated, whereas NP, PB1, PB2, PA, and NS2 are nucleocapsid associated proteins. The M1 protein is the most abundant protein in influenza particles. The HA and NA proteins are envelope glycoproteins, responsible for virus attachment and penetration of the viral particles into the cell, and the sources of the major immunodominant epitopes for virus neutralization and protective immunity. Both HA and NA proteins are considered the most important components for prophylactic influenza vaccines.
[0004] Each year, seasonal influenza causes over 300,000 hospitalizations and 36,000 deaths in the U.S. alone (Simonsen et al., Lancet Infect Dis 7:658-66, 2007). The emergence of the novel H1N1 influenza virus in 2009 demonstrated how quickly a new influenza pandemic can sweep across the world. The spread of highly pathogenic H5N1 viruses in birds and coincident infections in humans have raised the concerns that H5N1 viruses may cause a new pandemic in humans. Vaccination is an effective method to prevent influenza infection. There are two influenza vaccine approaches licensed in the United States; the inactivated, split vaccine and the live-attenuated virus vaccine. Inactivated vaccines can efficiently induce humoral immune responses but generally only poor cellular immune responses. Thus, a need exists for a broadly protective influenza virus vaccine.
SUMMARY
[0005] Disclosed herein is the development of an optimized influenza HA protein that elicits broadly reactive immune response to H5N1 influenza virus isolates. The optimized HA protein was developed through a series of HA protein alignments, and subsequent generation of consensus sequences for clade 2 H5N1 influenza virus isolates (FIG. 1). The final consensus HA amino acid sequence was reverse translated and optimized for expression in mammalian cells. The optimized HA coding sequence is set forth herein as SEQ ID NO: 1, and the optimized HA protein sequence is set forth herein as SEQ ID NO: 2.
[0006] Provided herein is an isolated nucleic acid molecule comprising a nucleotide sequence encoding an optimized influenza HA polypeptide, wherein the nucleotide sequence encoding the HA polypeptide is at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 1. Optimized influenza HA polypeptides encoded by the nucleic acid molecule, vectors comprising the nucleic acid molecule, and host cells containing the disclosed vectors are also provided herein.
[0007] Further provided is an optimized influenza HA polypeptide, wherein the amino acid sequence of the polypeptide is at least 99% identical to SEQ ID NO: 2. Also provided are fusion proteins comprising the optimized HA polypeptide, virus-like particles (VLPs) containing the optimized HA polypeptides, and compositions comprising the optimized HA polypeptide.
[0008] Collections of plasmids are also provided herein. In some embodiments, the collections of plasmids include a plasmid encoding an influenza NA, a plasmid encoding an influenza MA, and a plasmid encoding the optimized HA protein disclosed herein.
[0009] Further provided is a method of eliciting an immune response to influenza virus in a subject by administering the optimized influenza HA protein, fusion proteins containing the optimized influenza HA, or VLPs containing the optimized influenza HA, as disclosed herein. Also provided is a method of immunizing a subject against influenza virus by administering to the subject VLPs containing the optimized influenza HA protein disclosed herein.
[0010] The foregoing and other objects and features of the disclosure will become more apparent from the following detailed description, which proceeds with reference to the accompanying figures.
BRIEF DESCRIPTION OF THE FIGURES
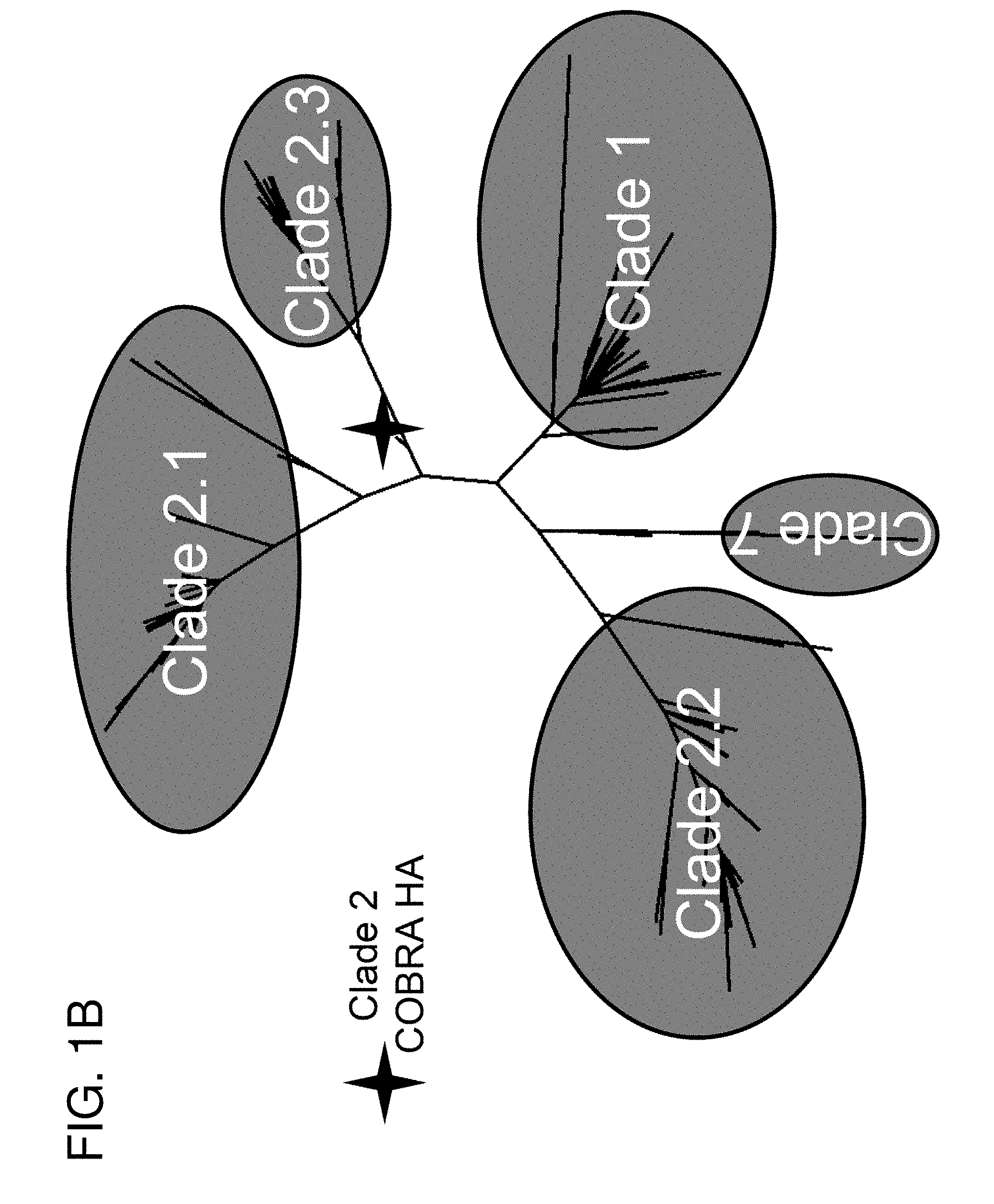
[0011] FIGS. 1A-1B: COBRA HA Design. (FIG. 1A) Schematic illustrating the design of the COBRA HA molecule. The phylogenetic tree was inferred from hemagglutinin amino acid sequences using the maximum likelihood method and clade/sub-clade groupings were identified. Primary consensus sequences were generated for each outbreak group. Secondary consensus sequences were then generated for each sub-clade using the primary sequences as input. The secondary consensus sequences were then aligned and the resulting consensus, designated COBRA, was generated. (FIG. 1B) Phylogenetic analysis of the COBRA HA. The unrooted phylogenetic tree was inferred from hemagglutinin amino acid sequences from human H5N1 infections isolated from 2004 to 2009 and the clade/sub-clade groupings are indicated. The star represents the COBRA HA sequence relative to human H5N1 infections.
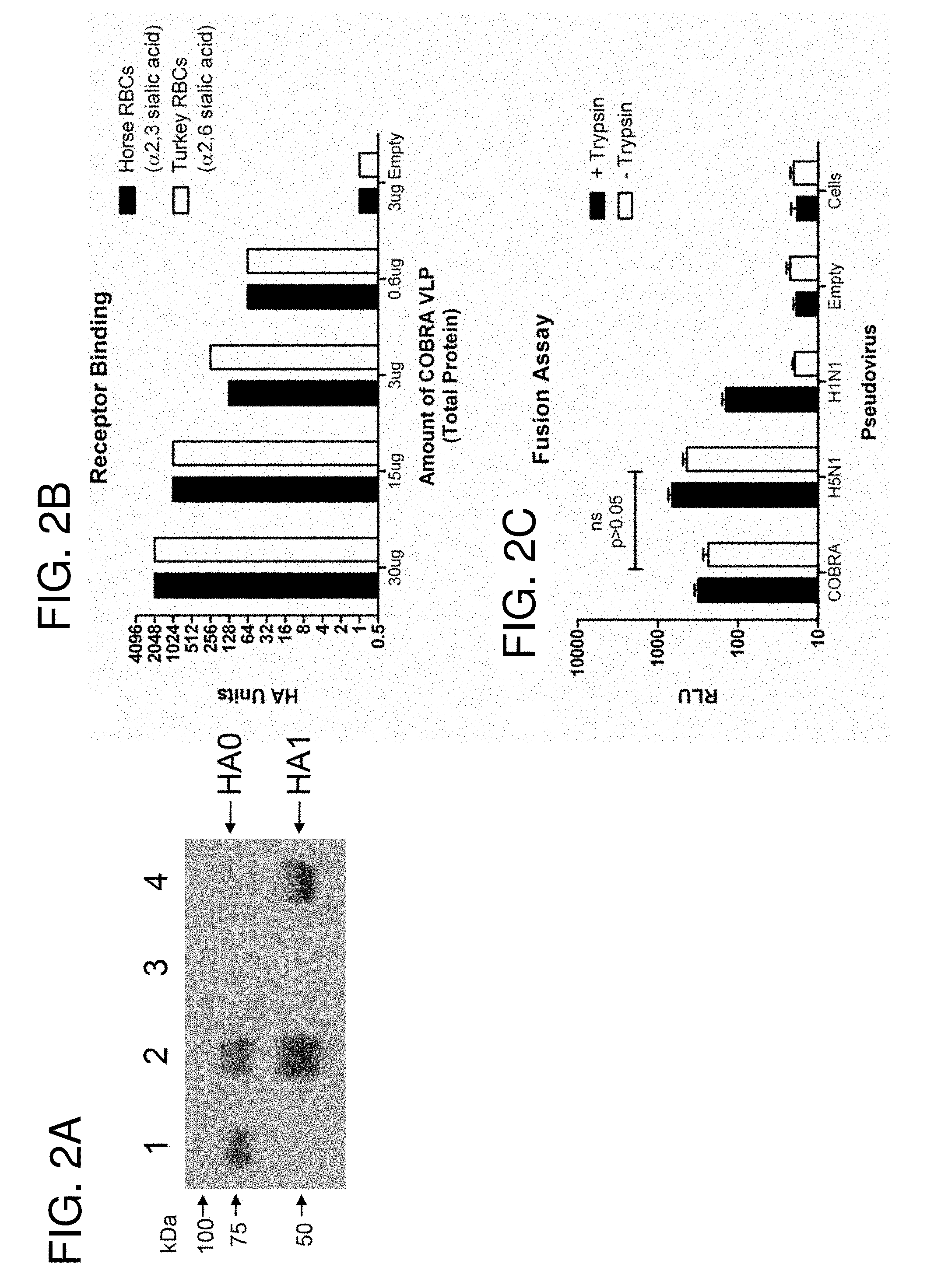
[0012] FIGS. 2A-2C: COBRA HA Functional Characterization. COBRA HA was translated in vitro and the cell culture lysates were analyzed by SDS-PAGE (FIG. 2A). Lane designations: 1) H5N1 recombinant HA; 2) COBRA HA; 3) Expression vector; 4) H5N1 reassortant virus. The COBRA HA (lane 2) migrates at its expected molecular weight confirming expression of the synthetic protein. COBRA HA VLPs were prepared in various amounts, serially diluted, and incubated with 1% erythrocytes to evaluate receptor binding (FIG. 2B). HA titer was determined as the last well in which the RBCs remained suspended in a lattice structure. COBRA HA and control lentiviral pseudoparticles packaging a CMV-Luc gene were generated in HEK 293T cells and used to infect MDCK cells with or without trypsin (FIG. 2C). Particle fusion was determined by luciferase production by infected cells.
[0013] FIGS. 3A-3F: COBRA HA Mouse Dosing Immunogenicity. BALB/c mice (n=5/group) were vaccinated at 0 and 3 weeks with blood collected at 14 to 21 days after each vaccination. Vaccines were formulated at high (1.5 .mu.g HA), and low (0.03 .mu.g HA) doses, with and without Imject.RTM. alum, and delivered intramuscularly. Total IgG at week 5 was determined via ELISA for each vaccine group (FIGS. 3A and 3B). Values represent the geometric mean titer (.+-.95% confidence interval) of log.sub.10 transformed endpoint titers. IgG isotypes were evaluated via ELISA for each vaccine group (FIGS. 3C and 3D). Values represent the mean OD.sub.450 of a 1:200 dilution of serum. Hemagglutination inhibition (HAI) serum antibody titer for each vaccine group was determined at week 5 using representative reassortant viruses (FIGS. 3E and 3F). Values represent the geometric mean titer (.+-.95% confidence interval) of log 2 transformed titers. The dotted line represents the 1:40 titer. Significant differences were determined by two-way ANOVA with Bonferroni's post-test to evaluate differences between the vaccine formulations for each test antigen. A p value of less than 0.05 was considered significant.
[0014] FIGS. 4A-4D: COBRA HA Mouse Dosing Efficacy. BALB/c mice (n=5/group) were vaccinated with COBRA HA VLPs with or without adjuvant. Mice were infected with 5.times.10.sup.3 PFU of the highly pathogenic clade 2.2 H5N1 virus A/Whooper Swan/Mongolia/244/2005. Mice were followed to monitor weight loss (FIGS. 4A and 4B) and sickness (FIGS. 4C and 4D). Sickness score was determined by evaluating activity (0=normal, 1=reduced, 2=severely reduced), hunched back (0=absent, 1=present) and ruffled fur (0=absent, 1=present). All mock vaccinated mice reached the experimental endpoint and required euthanasia by 6 days post infection.
[0015] FIGS. 5A-5B: Mouse Comparison Immunogenicity. BALB/c mice (n=20/group) were vaccinated at 0 and 3 weeks with blood collected at 14 to 21 days after each vaccination. Vaccines were formulated at a high dose (3 .mu.g HA) with Imject.RTM. alum and delivered intramuscularly. Total IgG at week 5 was determined via ELISA for each vaccine group (FIG. 5A). Values represent the geometric mean titer (.+-.95% confidence interval) of log.sub.10 transformed endpoint titers. Hemagglutination inhibition (HAI) serum antibody titer for each vaccine group was determined at week 5 using representative reassortant viruses (FIG. 5B). Values represent the geometric mean titer (.+-.95% confidence interval) of log 2 transformed titers. The dotted line represents the 1:40 titer. Significant differences were determined by two-way ANOVA with Bonferroni's post-test to evaluate differences between the vaccine formulations for each test antigen. A p value of less than 0.05 was considered significant.
[0016] FIGS. 6A-6B: Mouse Comparison Efficacy. BALB/c mice (n=20/group) were vaccinated with VLPs and adjuvant. Mice were infected with 5.times.10.sup.3 PFU of the highly pathogenic clade 2.2 H5N1 virus A/Whooper Swan/Mongolia/244/2005. Mice were followed to monitor weight loss (FIG. 6A) and sickness (FIG. 6B). Sickness score was determined by evaluating activity (0=normal, 1=reduced, 2=severely reduced), hunched back (0=absent, 1=present) and ruffled fur (0=absent, 1=present). All mock (adjuvant-only) vaccinated mice reached the experimental endpoint and required euthanasia by 6 days post infection.
[0017] FIGS. 7A-7B: Ferret Immunogenicity. Ferrets (n=9/group) were vaccinated with VLPs (15 .mu.g HA) with Imject.RTM. alum at weeks 0 and 3 and serum collected at week 5. Total IgG at week 5 was determined via ELISA for each vaccine group (FIG. 7A). Values represent the geometric mean titer (.+-.95% confidence interval) of login transformed endpoint titers. Hemagglutination inhibition (HAI) serum antibody titer for each vaccine group was determined at week 5 using representative reassortant viruses (FIG. 7B). Values represent the geometric mean titer (.+-.95% confidence interval) of log 2 transformed titers. The dotted line represents the 1:40 titer. Significant differences were determined by two-way ANOVA with Bonferroni's post-test to evaluate differences between the vaccine formulations for each test antigen. A p value of less than 0.05 was considered significant.
[0018] FIGS. 8A-8E: Ferret Efficacy. Ferrets (n=9/group) were vaccinated with VLPs formulated with adjuvant. Ferrets were challenged with 1.times.10.sup.6 PFU of the highly pathogenic clade 2.2 H5N1 virus A/Whooper Swan/Mongolia/244/2005. Animals were monitored daily for weight loss (FIG. 8A), survival (FIG. 8B), temperature (FIG. 8C) and clinical symptoms (FIG. 8D). Relative sickness scores were determined by measuring lethargy (0-3), runny nose (0-1), sneezing (0-1), loss of appetite (0-1) and diarrhea (0-1). Animals reaching experimental endpoint were euthanized according to institutional guidelines. Nasal washes were collected serially post infection and virus titers determined via plaque assay (FIG. 8E). Statistical significance was determined using a two-way ANOVA with Bonferroni's post test. A p value of less than 0.05 was considered significant.
[0019] FIG. 9: Phylogenetic diversity of H5N1 influenza. The unrooted phylogenetic tree was inferred from HA amino acid sequences derived from 8 to 10 representative isolates in all clades and sub-clades and the COBRA HA using the maximum likelihood method. Clade/sub-clade clusters were identified and are indicated in the shaded ovals. The star identifies where the COBRA antigen is located relative to the various representative isolates. Sequences were aligned with MUSCLE 3.7 software and the alignment was refined by Gblocks 0.91b software. Phylogeny was determined using the maximum likelihood method with PhyML software. Trees were rendered using TreeDyn 198.3 software (Dereeper et al., Nucleic Acids Res 36:W465-W469, 2008). The NCBI accession numbers for the HA sequences used in phylogeny inference were obtained through the Influenza Virus Resource (Bao et al., J Virol 82:596-601, 2008).
[0020] FIGS. 10A-10F: Serology. Total IgG at week 3 (FIG. 10A) and week 6 (FIG. 10B) was determined via ELISA for each vaccine group. Each collected antiserum was assayed for antibody binding to representative HA molecules from clade 2.1 (A/Indonesia/5/2005), clade 2.2 (A/Whooper Swan/Mongolia/244/2005), and clade 2.3 (A/Anhui/1/2005). Values represent the geometric mean titer (.+-.95% confidence interval) of login transformed endpoint titers. Statistical significance of the antibody titer data was determined using a two-way analysis of variance (ANOVA) followed by Bonferroni's post-test to analyze differences between each vaccine group for each of the different antigens that were tested (multiparametric). Significance was defined as p<0.05. Statistical analyses were performed with GraphPad Prism software. HAI titer for each vaccine group was determined at week 3 (FIG. 10C) and week 6 (FIG. 10D) using representative H5N1 influenza viruses: clade 2.1 (A/Indonesia/5/2005), clade 2.2 (A/Whooper swan/Mongolia/244/2005), and clade 2.3 (A/Anhui/1/2005). Values represent the geometric mean titer (.+-.95% confidence interval) of log 2 transformed titers. The dotted line represents the 1:40 titer. Significant differences were determined by two-way ANOVA with Bonferroni's post-test to evaluate differences between the vaccine formulations for each test antigen. A p value of less than 0.05 was considered significant. The number of monkeys that responded with a titer greater than 1:40 is listed above each bar. Neutralizing antibody at week 3 (FIG. 10E) and week 6 (FIG. 10F) was determined via microneutralization assay for each vaccine group. Values represent the geometric mean titer (.+-.95% confidence interval).
[0021] FIG. 11: HAI serum antibody titers from vaccinated monkeys against a panel of clade 0, 1, 2, 4, and 7 isolates. HAI titer for each vaccine group was determined at week 9 using H5N1 influenza viruses. Values represent the geometric mean titer (.+-.95% confidence interval) of log 2 transformed titers. Significant differences were determined by two-way ANOVA with Bonferroni's post-test to evaluate differences between the vaccine formulations for each test antigen. A p value of less than 0.05 was considered significant as described in FIG. 10.
[0022] FIGS. 12A-12D: Vaccine induced serum antibody responses. BALB/c mice (n=30/group) or Fitch ferrets (n=6/group) were vaccinated at 0 and 3 weeks with blood collected 14 to 21 days after each vaccination. Total IgG after the second vaccination was determined via ELISA for each vaccine group (FIGS. 12A and 12C). Receptor blocking antibody titers after the second vaccination were determined via hemagglutination inhibition (HAI) for each vaccine group (FIGS. 12B and 12D). Values represent the geometric mean of the reciprocal dilution (+/-95% confidence interval) of the last positive well. Significant differences between COBRA and polyvalent vaccines were determined by a two-tailed Student's T test and a p value of less than 0.05 was considered significant (*).
[0023] FIGS. 13A-13D: Highly pathogenic Clade 2.2 challenge. Vaccinated BALB/c mice (n=5/group) were infected with 5.times.10.sup.3 PFU of the highly pathogenic clade 2.2 H5N1 virus A/Whooper Swan/Mongolia/244/2005 (WS/05). Mice were monitored daily for weight loss (FIG. 13A) and sickness (FIG. 13B). Vaccinated Fitch ferrets (n=6/group) were infected with 1.times.10.sup.6 PFU of the highly pathogenic clade WS/05 virus. Ferrets were monitored daily for weight loss (FIG. 13C) and sickness (FIG. 13D). Values represent mean (+/-SEM) for each group.
[0024] FIGS. 14A-14B: Clade 2.2 viral loads. Vaccinated BALB/c mice (n=15/group) were infected with 5.times.10.sup.3 PFU of the highly pathogenic clade 2.2 H5N1 virus A/Whooper Swan/Mongolia/244/2005 (WS/05). Cohorts of mice (n=5/group) were sacrificed at 1, 3 and 5 days post infection, lungs harvested, and viral load determined by plaque assay (FIG. 14A). Vaccinated Fitch ferrets (n=6/group) were infected with 1.times.10.sup.6 PFU of the highly pathogenic WS/05 virus. Nasal washes were collected and viral load determined by plaque assay (FIG. 14B). Values represent mean (+/-SEM) viral titer for each group. Significant differences between COBRA and polyvalent vaccines were determined by a two-tailed Student's T test and a p value of less than 0.05 was considered significant (*).
[0025] FIGS. 15A-15B: Histopathology of infected lungs. Vaccinated BALB/c mice (n=15/group) were infected with 5.times.10.sup.3 PFU of the highly pathogenic clade 2.2 H5N1 virus A/Whooper Swan/Mongolia/244/2005 (WS/05). Cohorts of mice (n=5/group) were sacrificed at 3 days post infection and in situ hybridization (ISH) for influenza matrix protein (MP) was performed on sections from paraffin embedded lung tissue (FIG. 15A). Severity of influenza ISH foci was accessed in the bronchi (FIG. 15B). Scoring: 0=no definitive signal; 1=occasional focus; 2=focus in most fields; 3=more than one focus per field.
[0026] FIGS. 16A-16C: Clade 1 challenge. Vaccinated BALB/c mice (n=4/group) were infected with 5.times.10.sup.3 PFU of reassortant virus containing the HA and NA genes from the clade 1 H5N1 virus A/Vietnam/1203/2004 (VN/04). Mice were monitored daily for weight loss (FIG. 16A) and sickness (FIG. 16B). Values represent mean (+/-SD) for each group. An additional cohort of vaccinated mice (n=3/group) were infected and lungs were harvested 3 days post infection for analysis of viral burden (FIG. 16C). Values represent mean (+/-SEM) viral titer for each group.
[0027] FIGS. 17A-17B: Post-challenge cellular immune responses. Vaccinated BALB/c mice (n=3/group) were infected with 5.times.10.sup.3 PFU of reassortant virus containing the HA and NA genes from the clade 1 H5N1 virus A/Vietnam/1203/2004 (VN/04). Mice were sacrificed 6 days post infection, lungs were harvested and the numbers of antibody secreting cells (FIG. 17A) and IFN-.gamma. producing cells (FIG. 17B) were determined by ELISpot assay. Values represent the mean (+/-SEM) spots for each group.
[0028] FIGS. 18A-18B: Passive transfer clade 1 challenge. BALB/c mice (n=10/group) were vaccinated at 0 and 3 weeks with blood collected 14 to 21 days after each vaccination. Serum collected after the second vaccination was pooled for each vaccine group and administered to naive recipient mice (n=5/group). One day after passive transfer, recipient mice were infected with 5.times.10.sup.3 PFU of reassortant virus containing the HA and NA genes from the clade 1 H5N1 virus A/Vietnam/1203/2004 (VN/04). Mice were monitored daily for weight loss (FIG. 18A) and sickness (FIG. 18B). Values represent mean (+/-SD) for each group. Significant differences were determined by two-way ANOVA with Bonferroni's post-test to evaluate differences between vaccines at each day. A p value of less than 0.05 was considered significant (*).
SEQUENCE LISTING
[0029] The nucleic and amino acid sequences listed in the accompanying sequence listing are shown using standard letter abbreviations for nucleotide bases, and three letter code for amino acids, as defined in 37 C.F.R. 1.822. Only one strand of each nucleic acid sequence is shown, but the complementary strand is understood as included by any reference to the displayed strand. The Sequence Listing is submitted as an ASCII text file, created on Sep. 7, 2018, 51.0 KB, which is incorporated by reference herein. In the accompanying sequence listing:
[0030] SEQ ID NOs: 1 and 2 are the nucleotide and amino acid sequences, respectively, of a codon-optimized influenza HA (designated COBRA).
[0031] SEQ ID NOs: 3 and 4 are the nucleotide and amino acid sequences, respectively, of a codon-optimized influenza NA.
[0032] SEQ ID NOs: 5 and 6 are the nucleotide and amino acid sequences, respectively, of a codon-optimized influenza M1.
[0033] SEQ ID NO: 7 is the nucleotide sequence of a plasmid encoding a codon-optimized influenza HA.
[0034] SEQ ID NO: 8 is the nucleotide sequence of a plasmid encoding a codon-optimized influenza NA.
[0035] SEQ ID NO: 9 is the nucleotide sequence of a plasmid encoding a codon-optimized influenza M1.
[0036] SEQ ID NO: 10 is the amino acid sequence of a T cell epitope in H5 HA (HA.sub.533).
[0037] SEQ ID NO: 11 is the amino acid sequence of an ovalbumin T cell epitope (Ova.sub.257).
DETAILED DESCRIPTION
I. Abbreviations
[0038] ASC: antibody secreting cell
[0039] DPI: days post infection
[0040] HA: hemagglutinin or hemagglutination assay
[0041] HAI: hemagglutination inhibition
[0042] hRBC: horse red blood cell
[0043] IFU: infectious unit
[0044] LD.sub.50: lethal dose 50
[0045] M1: matrix protein 1
[0046] MN: microneutralization
[0047] MOI: multiplicity of infection
[0048] NA: neuraminidase
[0049] PFU: plaque form unit
[0050] RDE: receptor destroying enzyme
[0051] TCID: tissue culture infectious dose
[0052] tRBC: turkey red blood cell
[0053] VLP: virus-like particle
II. Terms and Methods
[0054] Unless otherwise noted, technical terms are used according to conventional usage. Definitions of common terms in molecular biology may be found in Benjamin Lewin, Genes V, published by Oxford University Press, 1994 (ISBN 0-19-854287-9); Kendrew et al. (eds.), The Encyclopedia of Molecular Biology, published by Blackwell Science Ltd., 1994 (ISBN 0-632-02182-9); and Robert A. Meyers (ed.), Molecular Biology and Biotechnology: a Comprehensive Desk Reference, published by VCH Publishers, Inc., 1995 (ISBN 1-56081-569-8).
[0055] In order to facilitate review of the various embodiments of the disclosure, the following explanations of specific terms are provided:
[0056] Adjuvant: A substance or vehicle that non-specifically enhances the immune response to an antigen. Adjuvants can include a suspension of minerals (alum, aluminum hydroxide, or phosphate) on which antigen is adsorbed; or water-in-oil emulsion in which antigen solution is emulsified in mineral oil (for example, Freund's incomplete adjuvant), sometimes with the inclusion of killed mycobacteria (Freund's complete adjuvant) to further enhance antigenicity. Immunostimulatory oligonucleotides (such as those including a CpG motif) can also be used as adjuvants (for example, see U.S. Pat. Nos. 6,194,388; 6,207,646; 6,214,806; 6,218,371; 6,239,116; 6,339,068; 6,406,705; and 6,429,199). Adjuvants also include biological molecules, such as costimulatory molecules. Exemplary biological adjuvants include IL-2, RANTES, GM-CSF, TNF-.alpha., IFN-.gamma., G-CSF, LFA-3, CD72, B7-1, B7-2, OX-40L and 41 BBL.
[0057] Administer: As used herein, administering a composition to a subject means to give, apply or bring the composition into contact with the subject. Administration can be accomplished by any of a number of routes, such as, for example, topical, oral, subcutaneous, intramuscular, intraperitoneal, intravenous, intrathecal and intramuscular.
[0058] Antibody: An immunoglobulin molecule produced by B lymphoid cells with a specific amino acid sequence. Antibodies are evoked in humans or other animals by a specific antigen (immunogen). Antibodies are characterized by reacting specifically with the antigen in some demonstrable way, antibody and antigen each being defined in terms of the other. "Eliciting an antibody response" refers to the ability of an antigen or other molecule to induce the production of antibodies.
[0059] Antigen: A compound, composition, or substance that can stimulate the production of antibodies or a T-cell response in an animal, including compositions that are injected or absorbed into an animal. An antigen reacts with the products of specific humoral or cellular immunity, including those induced by heterologous immunogens. In some embodiments of the disclosed compositions and methods, the antigen is an influenza HA protein.
[0060] Attenuated: In the context of a live virus, the virus is attenuated if its ability to infect a cell or subject and/or its ability to produce disease is reduced (for example, eliminated) compared to a wild-type virus. Typically, an attenuated virus retains at least some capacity to elicit an immune response following administration to an immunocompetent subject. In some cases, an attenuated virus is capable of eliciting a protective immune response without causing any signs or symptoms of infection. In some embodiments, the ability of an attenuated virus to cause disease in a subject is reduced at least about 10%, at least about 25%, at least about 50%, at least about 75% or at least about 90% relative to wild-type virus.
[0061] Clade: Refers to the different categorizations of the known influenza viruses, such as influenza A H5N1 viruses. Viruses in an H5N1 clade are genetically related, but do not share the exact viral genome. There are at least ten different clades of H5N1 subtypes designated in the art: clade 0 clade 1, clade 2, clade 3, clade 4, clade 5, clade 6, clade 7, clade 8 and clade 9 (Abdel-Ghafar et al., N Engl J Med 358:261-273, 2008). Clade 2 is further divided into sub-clades (including clade 2.1, clade 2.2, clade 2.3, clade 2.4 and clade 2.5).
[0062] Codon-optimized: A "codon-optimized" nucleic acid refers to a nucleic acid sequence that has been altered such that the codons are optimal for expression in a particular system (such as a particular species of group of species). For example, a nucleic acid sequence can be optimized for expression in mammalian cells. Codon optimization does not alter the amino acid sequence of the encoded protein.
[0063] Fusion protein: A protein generated by expression of a nucleic acid sequence engineered from nucleic acid sequences encoding at least a portion of two different (heterologous) proteins. To create a fusion protein, the nucleic acid sequences must be in the same reading frame and contain to internal stop codons. For example, a fusion protein includes an influenza HA fused to a heterologous protein.
[0064] Geographical location or geographical region: Refers to preselected divisions of geographical areas of the earth, for example, by continent or other preselected territory or subdivision (e.g., the Middle East, which spans more than one continent). Examples of different geographical regions include countries (e.g., Turkey, Egypt, Iraq, Azerbaijan, China, United States), continents (e.g., Asia, Europe, North America, South America, Oceania, Africa), and recognized geopolitical subdivisions (such as the Middle East).
[0065] Hemagglutinin (HA): An influenza virus surface glycoprotein. HA mediates binding of the virus particle to a host cells and subsequent entry of the virus into the host cell. The nucleotide and amino acid sequences of numerous influenza HA proteins are known in the art and are publically available, such as those deposited with GenBank (see Table 1 for a list of GenBank Accession Nos. of H5N1 HA sequences). HA (along with NA) is one of the two major influenza virus antigenic determinants.
[0066] Immune response: A response of a cell of the immune system, such as a B-cell, T-cell, macrophage or polymorphonucleocyte, to a stimulus such as an antigen or vaccine. An immune response can include any cell of the body involved in a host defense response, including for example, an epithelial cell that secretes an interferon or a cytokine. An immune response includes, but is not limited to, an innate immune response or inflammation. As used herein, a protective immune response refers to an immune response that protects a subject from infection (prevents infection or prevents the development of disease associated with infection). Methods of measuring immune responses are well known in the art and include, for example, measuring proliferation and/or activity of lymphocytes (such as B or T cells), secretion of cytokines or chemokines, inflammation, antibody production and the like.
[0067] Immunogen: A compound, composition, or substance which is capable, under appropriate conditions, of stimulating an immune response, such as the production of antibodies or a T-cell response in an animal, including compositions that are injected or absorbed into an animal. As used herein, as "immunogenic composition" is a composition comprising an immunogen (such as an HA polypeptide).
[0068] Immunize: To render a subject protected from an infectious disease, such as by vaccination.
[0069] Influenza virus: A segmented negative-strand RNA virus that belongs to the Orthomyxoviridae family. There are three types of Influenza viruses, A, B and C. Influenza A viruses infect a wide variety of birds and mammals, including humans, horses, marine mammals, pigs, ferrets, and chickens. In animals, most influenza A viruses cause mild localized infections of the respiratory and intestinal tract. However, highly pathogenic influenza A strains, such as H5N1, cause systemic infections in poultry in which mortality may reach 100%. H5N1 is also referred to as "avian influenza."
[0070] Isolated: An "isolated" biological component (such as a nucleic acid, protein or virus) has been substantially separated or purified away from other biological components (such as cell debris, or other proteins or nucleic acids). Biological components that have been "isolated" include those components purified by standard purification methods. The term also embraces recombinant nucleic acids, proteins or viruses, as well as chemically synthesized nucleic acids or peptides.
[0071] Linker: One or more amino acids that serve as a spacer between two polypeptides of a fusion protein.
[0072] Matrix (M1) protein: An influenza virus structural protein found within the viral envelope. M1 is thought to function in assembly and budding.
[0073] Neuraminidase (NA): An influenza virus membrane glycoprotein. NA is involved in the destruction of the cellular receptor for the viral HA by cleaving terminal sialic acid residues from carbohydrate moieties on the surfaces of infected cells. NA also cleaves sialic acid residues from viral proteins, preventing aggregation of viruses. NA (along with HA) is one of the two major influenza virus antigenic determinants.
[0074] Operably linked: A first nucleic acid sequence is operably linked with a second nucleic acid sequence when the first nucleic acid sequence is placed in a functional relationship with the second nucleic acid sequence. For instance, a promoter is operably linked to a coding sequence if the promoter affects the transcription or expression of the coding sequence. Generally, operably linked DNA sequences are contiguous and, where necessary to join two protein-coding regions, in the same reading frame.
[0075] Optimized influenza HA protein: As used herein, "optimized influenza HA protein" refers to the HA protein consensus sequence generated by sequence alignments of clade 2 H5N1 influenza viruses (as described in Example 1 below). The nucleotide sequence encoding the optimized HA protein was further optimized for expression in mammalian cells via codon-optimization and RNA optimization (such as to increase RNA stability). The optimized influenza HA protein disclosed herein (and set forth herein as SEQ ID NO: 2) is also referred to as "COBRA."
[0076] ORF (open reading frame): A series of nucleotide triplets (codons) coding for amino acids without any termination codons. These sequences are usually translatable into a peptide.
[0077] Outbreak: As used herein, an influenza virus "outbreak" refers to a collection of virus isolates from within a single country in a given year.
[0078] Pharmaceutically acceptable vehicles: The pharmaceutically acceptable carriers (vehicles) useful in this disclosure are conventional. Remington's Pharmaceutical Sciences, by E. W. Martin, Mack Publishing Co., Easton, Pa., 15th Edition (1975), describes compositions and formulations suitable for pharmaceutical delivery of one or more therapeutic compositions, such as one or more influenza vaccines, and additional pharmaceutical agents.
[0079] In general, the nature of the carrier will depend on the particular mode of administration being employed. For instance, parenteral formulations usually comprise injectable fluids that include pharmaceutically and physiologically acceptable fluids such as water, physiological saline, balanced salt solutions, aqueous dextrose, glycerol or the like as a vehicle. For solid compositions (for example, powder, pill, tablet, or capsule forms), conventional non-toxic solid carriers can include, for example, pharmaceutical grades of mannitol, lactose, starch, or magnesium stearate. In addition to biologically-neutral carriers, pharmaceutical compositions to be administered can contain minor amounts of non-toxic auxiliary substances, such as wetting or emulsifying agents, preservatives, and pH buffering agents and the like, for example sodium acetate or sorbitan monolaurate.
[0080] Plasmid: A circular nucleic acid molecule capable of autonomous replication in a host cell.
[0081] Polypeptide: A polymer in which the monomers are amino acid residues which are joined together through amide bonds. When the amino acids are alpha-amino acids, either the L-optical isomer or the D-optical isomer can be used. The terms "polypeptide" or "protein" as used herein are intended to encompass any amino acid sequence and include modified sequences such as glycoproteins. The term "polypeptide" is specifically intended to cover naturally occurring proteins, as well as those which are recombinantly or synthetically produced. The term "residue" or "amino acid residue" includes reference to an amino acid that is incorporated into a protein, polypeptide, or peptide.
[0082] Conservative amino acid substitutions are those substitutions that, when made, least interfere with the properties of the original protein, that is, the structure and especially the function of the protein is conserved and not significantly changed by such substitutions. Examples of conservative substitutions are shown below.
TABLE-US-00001 Original Residue Conservative Substitutions Ala Ser Arg Lys Asn Gln, His Asp Glu Cys Ser Gln Asn Glu Asp His Asn; Gln Ile Leu, Val Leu Ile; Val Lys Arg; Gln; Glu Met Leu; Ile Phe Met; Leu; Tyr Ser Thr Thr Ser Trp Tyr Tyr Trp; Phe Val Ile; Leu
[0083] Conservative substitutions generally maintain (a) the structure of the polypeptide backbone in the area of the substitution, for example, as a sheet or helical conformation, (b) the charge or hydrophobicity of the molecule at the target site, or (c) the bulk of the side chain.
[0084] The substitutions which in general are expected to produce the greatest changes in protein properties will be non-conservative, for instance changes in which (a) a hydrophilic residue, for example, seryl or threonyl, is substituted for (or by) a hydrophobic residue, for example, leucyl, isoleucyl, phenylalanyl, valyl or alanyl; (b) a cysteine or proline is substituted for (or by) any other residue; (c) a residue having an electropositive side chain, for example, lysyl, arginyl, or histadyl, is substituted for (or by) an electronegative residue, for example, glutamyl or aspartyl; or (d) a residue having a bulky side chain, for example, phenylalanine, is substituted for (or by) one not having a side chain, for example, glycine.
[0085] Preventing, treating or ameliorating a disease: "Preventing" a disease refers to inhibiting the full development of a disease. "Treating" refers to a therapeutic intervention that ameliorates a sign or symptom of a disease or pathological condition after it has begun to develop. "Ameliorating" refers to the reduction in the number or severity of signs or symptoms of a disease.
[0086] Promoter: An array of nucleic acid control sequences which direct transcription of a nucleic acid. A promoter includes necessary nucleic acid sequences near the start site of transcription. A promoter also optionally includes distal enhancer or repressor elements. A "constitutive promoter" is a promoter that is continuously active and is not subject to regulation by external signals or molecules. In contrast, the activity of an "inducible promoter" is regulated by an external signal or molecule (for example, a transcription factor). In some embodiments herein, the promoter is a CMV promoter.
[0087] Purified: The term "purified" does not require absolute purity; rather, it is intended as a relative term. Thus, for example, a purified peptide, protein, virus, or other active compound is one that is isolated in whole or in part from naturally associated proteins and other contaminants. In certain embodiments, the term "substantially purified" refers to a peptide, protein, virus or other active compound that has been isolated from a cell, cell culture medium, or other crude preparation and subjected to fractionation to remove various components of the initial preparation, such as proteins, cellular debris, and other components.
[0088] Recombinant: A recombinant nucleic acid, protein or virus is one that has a sequence that is not naturally occurring or has a sequence that is made by an artificial combination of two otherwise separated segments of sequence. This artificial combination is often accomplished by chemical synthesis or by the artificial manipulation of isolated segments of nucleic acids, for example, by genetic engineering techniques.
[0089] Sequence identity: The similarity between amino acid or nucleic acid sequences is expressed in terms of the similarity between the sequences, otherwise referred to as sequence identity. Sequence identity is frequently measured in terms of percentage identity (or similarity or homology); the higher the percentage, the more similar the two sequences are. Homologs or variants of a given gene or protein will possess a relatively high degree of sequence identity when aligned using standard methods.
[0090] Methods of alignment of sequences for comparison are well known in the art. Various programs and alignment algorithms are described in: Smith and Waterman, Adv. Appl. Math. 2:482, 1981; Needleman and Wunsch, J. Mol. Biol. 48:443, 1970; Pearson and Lipman, Proc. Natl. Acad. Sci. U.S.A. 85:2444, 1988; Higgins and Sharp, Gene 73:237-244, 1988; Higgins and Sharp, CABIOS 5:151-153, 1989; Corpet et al., Nucleic Acids Research 16:10881-10890, 1988; and Pearson and Lipman, Proc. Natl. Acad. Sci. U.S.A. 85:2444, 1988. Altschul et al., Nature Genet. 6:119-129, 1994.
[0091] The NCBI Basic Local Alignment Search Tool (BLAST.TM.) (Altschul et al., J. Mol. Biol. 215:403-410, 1990) is available from several sources, including the National Center for Biotechnology Information (NCBI, Bethesda, Md.) and on the Internet, for use in connection with the sequence analysis programs blastp, blastn, blastx, tblastn and tblastx.
[0092] Subject: Living multi-cellular vertebrate organisms, a category that includes both human and non-human mammals, such as non-human primates. In one example, a subject is one who is infected with H5N1 or is susceptible to such infection.
[0093] Therapeutically effective amount: A quantity of a specified agent sufficient to achieve a desired effect in a subject being treated with that agent. For example, this may be the amount of an influenza virus vaccine useful for eliciting an immune response in a subject and/or for preventing infection by influenza virus. Ideally, in the context of the present disclosure, a therapeutically effective amount of an influenza vaccine is an amount sufficient to increase resistance to, prevent, ameliorate, and/or treat infection caused by influenza virus in a subject without causing a substantial cytotoxic effect in the subject. The effective amount of an influenza vaccine useful for increasing resistance to, preventing, ameliorating, and/or treating infection in a subject will be dependent on, for example, the subject being treated, the manner of administration of the therapeutic composition and other factors.
[0094] Transformed: A transformed cell is a cell into which has been introduced a nucleic acid molecule by molecular biology techniques. As used herein, the term transformation encompasses all techniques by which a nucleic acid molecule might be introduced into such a cell, including transfection with viral vectors, transformation with plasmid vectors, and introduction of naked DNA by electroporation, lipofection, and particle gun acceleration.
[0095] Vaccine: A preparation of immunogenic material capable of stimulating an immune response, administered for the prevention, amelioration, or treatment of disease, such as an infectious disease. The immunogenic material may include, for example, attenuated or killed microorganisms (such as attenuated viruses), or antigenic proteins, peptides or DNA derived from them. Vaccines may elicit both prophylactic (preventative) and therapeutic responses. Methods of administration vary according to the vaccine, but may include inoculation, ingestion, inhalation or other forms of administration. Inoculations can be delivered by any of a number of routes, including parenteral, such as intravenous, subcutaneous or intramuscular. Vaccines may be administered with an adjuvant to boost the immune response.
[0096] Vector: A vector is a nucleic acid molecule allowing insertion of foreign nucleic acid without disrupting the ability of the vector to replicate and/or integrate in a host cell. A vector can include nucleic acid sequences that permit it to replicate in a host cell, such as an origin of replication. An insertional vector is capable of inserting itself into a host nucleic acid. A vector can also include one or more selectable marker genes and other genetic elements. An expression vector is a vector that contains the necessary regulatory sequences to allow transcription and translation of inserted gene or genes. In some embodiments of the present disclosure, the vector encodes an influenza HA, NA or M1 protein. In some embodiments, the vector is the pTR600 expression vector (U.S. Patent Application Publication No. 2002/0106798; Ross et al., Nat Immunol. 1(2):102-103, 2000; Green et al., Vaccine 20:242-248, 2001).
[0097] Virus-like particle (VLP): Virus particles made up of one of more viral structural proteins, but lacking the viral genome. Because VLPs lack a viral genome, they are non-infectious. In addition, VLPs can often be produced by heterologous expression and can be easily purified. Most VLPs comprise at least a viral core protein that drives budding and release of particles from a host cell. One example of such a core protein is influenza M1. In some embodiments herein, an influenza VLP comprises the HA, NA and M1 proteins. As described herein, influenza VLPs can be produced by transfection of host cells with plasmids encoding the HA, NA and M1 proteins. After incubation of the transfected cells for an appropriate time to allow for protein expression (such as for approximately 72 hours), VLPs can be isolated from cell culture supernatants. Example 1 provides an exemplary protocol for purifying influenza VLPs from cell supernatants. In this example, VLPs are isolated by low speed centrifugation (to remove cell debris), vacuum filtration and ultracentrifugation through 20% glycerol.
[0098] Unless otherwise explained, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this disclosure belongs. The singular terms "a," "an," and "the" include plural referents unless context clearly indicates otherwise. Similarly, the word "or" is intended to include "and" unless the context clearly indicates otherwise. Hence "comprising A or B" means including A, or B, or A and B. It is further to be understood that all base sizes or amino acid sizes, and all molecular weight or molecular mass values, given for nucleic acids or polypeptides are approximate, and are provided for description. Although methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present disclosure, suitable methods and materials are described below. All publications, patent applications, patents, and other references mentioned herein are incorporated by reference in their entirety. All GenBank Accession numbers are incorporated by reference herein as they appeared in the database on Sep. 9, 2010. In case of conflict, the present specification, including explanations of terms, will control. In addition, the materials, methods, and examples are illustrative only and not intended to be limiting.
III. Overview of Several Embodiments
[0099] Disclosed herein is the development of a computationally optimized influenza HA protein that elicits broadly reactive immune responses to H5N1 influenza virus isolates, such as the isolates listed in Table 1. The optimized HA protein was developed through a series of HA protein alignments, and subsequent generation of consensus sequences, for clade 2 H5N1 influenza virus isolates (described in detail in Example 1 below; see also FIG. 1). The final consensus HA amino acid sequence was reverse translated and optimized for expression in mammalian cells. Optimization of the nucleic acid sequence included optimization of the codons for expression in mammalian cells and RNA optimization (such as RNA stability). The optimized HA coding sequence is set forth herein as SEQ ID NO: 1, and the optimized HA protein sequence is set forth herein as SEQ ID NO: 2.
[0100] Thus, provided herein is an isolated nucleic acid molecule comprising a nucleotide sequence encoding an influenza HA polypeptide. In some embodiments, the nucleotide sequence encoding the HA polypeptide is at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 1.
[0101] In some examples, the nucleotide sequence encoding the influenza HA polypeptide that is at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% identical to SEQ ID NO: 1 lacks the start codon (nucleotides 1-3 of SEQ ID NO: 1), encoding a N-terminal methionine. In particular examples, the nucleotide sequence encoding the influenza HA polypeptide is at least 94% identical to nucleotides 4-1707 of SEQ ID NO: 1. In other examples, the nucleotide sequence encoding the HA polypeptide comprises or consists of nucleotides 4-1707 of SEQ ID NO: 1.
[0102] In some examples, the nucleotide sequence encoding the HA polypeptide comprises SEQ ID NO: 1. In particular examples, the nucleotide sequence encoding the HA polypeptide consists of SEQ ID NO: 1. Also provided herein are influenza HA polypeptides encoded by the disclosed nucleic acid molecules.
[0103] Further provided are vectors containing a nucleotide sequence encoding an optimized HA polypeptide. In some embodiments of the vectors provided herein, the nucleotide sequence encoding the HA polypeptide is at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 1. In some examples, the vector further includes a promoter operably linked to the nucleotide sequence encoding the HA polypeptide. In particular examples, the promoter is a cytomegalovirus (CMV) promoter. In some embodiments, the nucleotide sequence of the vector is at least 85%, at least 90%, at least 95%, at least 98% or at least 99% identical to the nucleotide sequence of SEQ ID NO: 7. In some examples, the nucleotide sequence of the vector comprises the nucleotide sequence of SEQ ID NO: 7. In particular examples, the nucleotide sequence of the vector consists of the nucleotide sequence of SEQ ID NO: 7.
[0104] Also provided herein are influenza HA polypeptides produced by transfecting a host cell with a vector provided herein under conditions sufficient to allow for expression of the HA polypeptide. Isolated cells containing the disclosed vectors are also provided.
[0105] Also provided herein are optimized influenza HA polypeptides. In some embodiments, the amino acid sequence of the polypeptide is at least 99% identical to SEQ ID NO: 2. In some examples, the amino acid sequence of the influenza HA polypeptide that is at least 99% identical to SEQ ID NO: 2 lacks the N-terminal methionine residue. In particular examples, the amino acid sequence of the influenza HA polypeptide is at least 99% identical to amino acids 2-568 of SEQ ID NO: 2. In other examples, the amino acid sequence of the HA polypeptides comprises or consists of amino acids 2-568 of SEQ ID NO: 2.
[0106] In some examples, the amino acid sequence of the polypeptide comprises the amino acid sequence of SEQ ID NO: 2. In particular examples, the amino acid sequence of the polypeptide consists of the amino acid sequence of SEQ ID NO: 2. Fusion proteins comprising the influenza HA polypeptides disclosed herein are also provided. The influenza HA polypeptide can be fused to any heterologous amino acid sequence to form the fusion protein.
[0107] Further provided herein are influenza virus-like particles (VLPs) containing an optimized influenza HA protein disclosed herein. In some embodiments, the HA protein of the VLP is at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99% or 100% identical to SEQ ID NO: 2. The influenza VLPs can further include any additional influenza proteins necessary to form the virus particle. In some embodiments, the influenza VLPs further include influenza neuraminidase (NA) protein, influenza matrix (M1) protein, or both.
[0108] In some embodiments of the influenza VLPs, the amino acid sequence of the influenza NA protein is at least 85%, at least 90%, at least 95%, at least 98% or at least 99% identical to SEQ ID NO: 4. In some examples, the amino acid sequence of the influenza NA protein comprises SEQ ID NO: 4. In particular examples, the amino acid sequence of the influenza NA protein consists of SEQ ID NO: 4. In some embodiments, the amino acid sequence of the influenza M1 protein is at least 85%, at least 90%, at least 95%, at least 98% or at least 99% identical to SEQ ID NO: 6. In some examples, the amino acid sequence of the influenza M1 protein comprises SEQ ID NO: 6. In particular examples, the amino acid sequence of the influenza M1 protein consists of SEQ ID NO: 6.
[0109] Also provided is an influenza VLP containing an influenza HA polypeptide as described herein, produced by transfecting a host cell with a vector encoding the HA polypeptide, a vector encoding an influenza NA protein and a vector encoding an influenza M1 protein, under conditions sufficient to allow for expression of the HA, M1 and NA proteins.
[0110] The vectors used to express the HA, NA and M1 proteins can be any suitable expression vectors known in the art. The vectors can be, for example, mammalian expression vectors, or viral vectors. In some embodiments, the vector is the pTR600 expression vector (U.S. Patent Application Publication No. 2002/0106798, herein incorporated by reference; Ross et al., Nat Immunol. 1(2):102-103, 2000; Green et al., Vaccine 20:242-248, 2001).
[0111] In some embodiments, the nucleotide sequence of the vector encoding the HA polypeptide is at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 7. In some examples, the nucleotide sequence of the vector encoding the HA polypeptide comprises SEQ ID NO: 7. In particular examples, the nucleotide sequence of the vector encoding the HA polypeptide consists of SEQ ID NO: 7.
[0112] In some embodiments, the nucleotide sequence of the vector encoding the NA protein is at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 8. In some examples, the nucleotide sequence of the vector encoding the NA protein comprises SEQ ID NO: 8. In particular examples, the nucleotide sequence of the vector encoding the NA protein consists of SEQ ID NO: 8.
[0113] In some embodiments, the nucleotide sequence of the vector encoding the M1 protein is at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 9. In some examples, the nucleotide sequence of the vector encoding the M1 protein comprises SEQ ID NO: 9. In particular examples, the nucleotide sequence of the vector encoding the M1 protein consists of SEQ ID NO: 9.
[0114] Collections of plasmids are also provided herein. In some embodiments, the collection of plasmids includes a plasmid encoding an influenza NA, a plasmid encoding an influenza MA, and a plasmid encoding the optimized HA protein disclosed herein. In some embodiments, the nucleotide sequence encoding the codon-optimized influenza HA of the HA-encoding plasmid is at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 1. Also provided are kits comprising the collection of plasmids.
[0115] In some embodiments of the collections of plasmids, the influenza NA is codon-optimized and/or the influenza M1 is codon-optimized. In some examples, the nucleotide sequence encoding the codon-optimized influenza NA is at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 3. In particular examples, the nucleotide sequence encoding the codon-optimized influenza NA comprises, or consists of, SEQ ID NO: 3. In some examples, the nucleotide sequence encoding the codon-optimized influenza M1 is at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical to SEQ ID NO: 5. In particular examples, the nucleotide sequence encoding the codon-optimized influenza M1 comprises, or consists of, SEQ ID NO: 5.
[0116] In one non-limiting example, the plasmid encoding influenza NA comprises SEQ ID NO: 8; the plasmid encoding influenza M1 comprises SEQ ID NO: 9; and the plasmid encoding influenza HA comprises SEQ ID NO: 10.
[0117] In some embodiments, transfection of the collection of plasmids into host cells under conditions sufficient to allow for expression of the HA, NA and M1 proteins produces influenza VLPs comprising the HA, NA and M1 proteins.
[0118] Also provided herein are compositions comprising an optimized influenza HA protein as disclosed herein, or a fusion protein or VLP comprising the optimized influenza HA protein. In some embodiments, the compositions further comprise a pharmaceutically acceptable carrier and/or an adjuvant. For example, the adjuvant can be alum, Freund's complete adjuvant, a biological adjuvant or immunostimulatory oligonucleotides (such as CpG oligonucleotides).
[0119] Further provided is a method of eliciting an immune response to influenza virus in a subject by administering an influenza HA protein disclosed herein, fusion proteins containing the influenza HA, or VLPs containing the influenza HA, as disclosed herein. In some embodiments, the influenza virus is an H5N1 influenza virus. In some embodiments, the HA protein, HA fusion protein or VLP can be administered using any suitable route of administration, such as, for example, intramuscular. In some embodiments, the HA protein, fusion protein or VLP is administered as a composition further comprising a pharmaceutically acceptable carrier and/or an adjuvant. For example, the adjuvant can be alum, Freund's complete adjuvant, a biological adjuvant or immunostimulatory oligonucleotides (such as CpG oligonucleotides).
[0120] Also provided is a method of immunizing a subject against influenza virus by administering to the subject VLPs containing the optimized influenza HA protein disclosed herein, or administering a composition thereof. In some embodiments of the method, the composition further comprises a pharmaceutically acceptable carrier and/or an adjuvant. For example, the adjuvant can be alum, Freund's complete adjuvant, a biological adjuvant or immunostimulatory oligonucleotides (such as CpG oligonucleotides). In some embodiments, the VLPs (or compositions thereof) are administered intramuscularly.
[0121] In some embodiments of the methods of eliciting an immune response or immunizing a subject, the subject is administered at least 1 .mu.g of the VLPs containing the optimized HA protein, such as at least 5 .mu.g, at least 10 .mu.g, at least 15 .mu.g, at least 20 .mu.g, at least 25 .mu.g, at least 30 .mu.g, at least 40 .mu.g or at least 50 .mu.g of the VLPs containing the optimized HA protein, for example about 1 to about 50 .mu.g or about 1 to about 25 .mu.g of the VLPs containing the optimized HA protein. In particular examples, the subject is administered about 5 to about 20 .mu.g of the VLPs, or about 10 to about 15 .mu.g of the VLPs. In one specific non-limiting example, the subject is administered about 15 .mu.g of the VLPs. However, one of skill in the art is capable of determining a therapeutically effective amount (for example an amount that provides protection against H5N1 influenza virus infection) of VLPs to administer to a subject.
[0122] It is disclosed herein that administration of VLPs comprising the COBRA HA disclosed herein (SEQ ID NO: 2) elicits protective levels of HAI antibodies to a number of representative clade 2 isolates and provides complete protection against lethal challenge with a clade 2.2 H5N1 virus. In some embodiments, administration of VLPs containing an optimized influenza HA results in production of high HAI titers (.gtoreq.1:40) to H5N1 clade 1, clade 2.1, clade 2.2 and clade 2.3 isolates. In some examples, the VLPs containing an optimized influenza HA elicit high HAI titers against clade 1 and/or clade 7 viruses. The VLPs containing an optimized influenza HA disclosed herein elicit a broader immune response (e.g., elicit neutralizing antibodies to a broader range is H5N1 virus isolates, such as those from clade 1, sub-clades of clade 2, and clade 7) than a polyvalent influenza virus vaccine.
[0123] Also provided herein is a method of optimizing an influenza protein sequence to elicit broadly reactive immune responses in a subject. In the context of the present disclosure, "broadly reactive" means the protein sequence elicits an immune response in a subject that is sufficient to inhibit, neutralize or prevent infection of a broad range of influenza viruses (such as most or all influenza viruses within a specific subtype). In some embodiments, the influenza protein is influenza HA or influenza NA. In one example, the optimized influenza protein is capable of eliciting a protective immune response against most or all known H5N1 influenza virus isolates (such as those listed in Table 1), such as about 80%, about 85%, about 90% or about 95% of known H5N1 influenza virus isolates.
[0124] In some embodiments, the method of optimizing an influenza protein sequence includes obtaining the amino acid sequences of a group of influenza virus isolates. For example, the group can consist of influenza virus isolates from a specific subtype (such as, for example, H5N1 or H1N1), and/or from one or more clades/sub-clades of a specific influenza subtype (for example, one or more of clades/sub-clades 0, 1, 2.1, 2.2, 2.3, 2.4, 3, 4, 5, 6, 7, 8 and 9 of H5N1). Amino acid sequences of the group of influenza viruses are first organized by clade or sub-clade and then by geographic location within each clade or sub-clade. The amino acid sequences for each geographic location are aligned to generate a primary consensus sequence for each geographical region. Grouping virus isolates by geographical region controls for single outbreak dominance and incomplete reporting and sequencing. The primary consensus sequence can be generated, for example, by multiple alignment analysis using AlignX (Vector NTI), or by any other method known in the art. The primary geographically-based consensus sequences for each clade or sub-clade are then aligned, and a secondary consensus sequence is generated for each clade or sub-clade. The secondary consensus sequences for each clade or sub-clade are then aligned to generate the optimized, broadly reactive, consensus sequence (see FIG. 1). In some embodiments, the optimized influenza virus polypeptide sequence is further optimized for expression in mammalian cells. In some examples, optimization includes reverse translation of the optimized influenza virus polypeptide sequence to generate a coding sequence, followed by codon-optimization and/or optimization of the RNA (such as for stability).
[0125] In one non-limiting example, the method of optimization includes: (i) obtaining the amino acid sequences of the polypeptide from a group of influenza virus isolates, wherein the influenza virus isolates are from the same subtype; (ii) organizing the amino acid sequences of the polypeptide from the group of influenza virus isolates by clade or sub-clade and then by geographical region within each clade or sub-clade; (iii) aligning the amino acid sequences within each geographical region to generate primary consensus sequences, wherein each geographic region is represented by a primary consensus sequence; (iv) aligning the primary consensus sequences to generate secondary consensus sequences, wherein each clade or sub-clade is represented by a secondary consensus sequence; and (v) aligning the secondary consensus sequences to generate the optimized influenza virus polypeptide sequence. In some cases, the method further includes (i) reverse translating the optimized influenza virus polypeptide sequence to generate a coding sequence; and (ii) optimizing the coding sequence for expression in mammalian cells.
[0126] In an alternative embodiment, the primary consensus sequence is obtained by aligning influenza protein sequences (such as HA or NA sequences) from viral isolates from a single outbreak (a collection of influenza virus isolates within a single country within a given year). Thus, in one non-limiting example, the method of optimization includes: (i) obtaining the amino acid sequences of the polypeptide from a group of influenza virus isolates, wherein the influenza virus isolates are from the same subtype; (ii) organizing the amino acid sequences of the polypeptide from the group of influenza virus isolates by clade or sub-clade and then by outbreak; (iii) aligning the amino acid sequences within each outbreak to generate primary consensus sequences, wherein each outbreak is represented by a primary consensus sequence; (iv) aligning the primary consensus sequences to generate secondary consensus sequences, wherein each clade or sub-clade is represented by a secondary consensus sequence; and (v) aligning the secondary consensus sequences to generate the optimized influenza virus polypeptide sequence. In some cases, the method further includes (i) reverse translating the optimized influenza virus polypeptide sequence to generate a coding sequence; and (ii) optimizing the coding sequence for expression in mammalian cells.
VI. Influenza
[0127] Influenza viruses are segmented negative-strand RNA viruses that belong to the Orthomyxoviridae family. There are three types of Influenza viruses, A, B and C. Influenza A viruses infect a wide variety of birds and mammals, including humans, horses, marine mammals, pigs, ferrets, and chickens. In animals, most influenza A viruses cause mild localized infections of the respiratory and intestinal tract. However, highly pathogenic influenza A strains, such as H5N1, cause systemic infections in poultry in which mortality may reach 100%. Animals infected with influenza A often act as a reservoir for the influenza viruses and certain subtypes have been shown to cross the species barrier to humans.
[0128] Influenza A viruses can be classified into subtypes based on allelic variations in antigenic regions of two genes that encode surface glycoproteins, namely, hemagglutinin (HA) and neuraminidase (NA) which are required for viral attachment and cellular release. Currently, sixteen subtypes of HA (H1-H16) and nine NA (N1-N9) antigenic variants are known for influenza A virus. Previously, only three subtypes were known to circulate in humans (H1N1, H1N2, and H3N2). However, in recent years, the pathogenic H5N1 subtype of avian influenza A has been reported to cross the species barrier and infect humans as documented in Hong Kong in 1997 and 2003, leading to the death of several patients.
[0129] In humans, the avian influenza virus infects cells of the respiratory tract as well as the intestinal tract, liver, spleen, kidneys and other organs. Symptoms of avian influenza infection include fever, respiratory difficulties including shortness of breath and cough, lymphopenia, diarrhea and difficulties regulating blood sugar levels. In contrast to seasonal influenza, the group most at risk is healthy adults which make up the bulk of the population. Due to the high pathogenicity of certain avian influenza A subtypes, particularly H5N1, and their demonstrated ability to cross over to infect humans, there is a significant economic and public health risk associated with these viral strains, including a real epidemic and pandemic threat. Currently, no effective vaccines for H5N1 infection are available.
[0130] The influenza A virus genome encodes nine structural proteins and one nonstructural (NS1) protein with regulatory functions. The influenza virus segmented genome contains eight negative-sense RNA (nsRNA) gene segments (PB2, PB1, PA, NP, M, NS, HA and NA) that encode at least ten polypeptides, including RNA-directed RNA polymerase proteins (PB2, PB1 and PA), nucleoprotein (NP), neuraminidase (NA), hemagglutinin (subunits HA1 and HA2), the matrix proteins (M1 and M2) and the non-structural proteins (NS1 and NS2) (Krug et al., In "The Influenza Viruses," R. M. Krug, ed., Plenum Press, N.Y., 1989, pp. 89 152).
[0131] Influenza virus' ability to cause widespread disease is due to its ability to evade the immune system by undergoing antigenic change, which is believed to occur when a host is infected simultaneously with both an animal influenza virus and a human influenza virus. During mutation and reassortment in the host, the virus may incorporate an HA and/or NA surface protein gene from another virus into its genome, thereby producing a new influenza subtype and evading the immune system.
[0132] HA is a viral surface glycoprotein generally comprising approximately 560 amino acids and representing 25% of the total virus protein. It is responsible for adhesion of the viral particle to, and its penetration into, a host cell in the early stages of infection. Cleavage of the virus HA0 precursor into the HA1 and HA2 sub-fragments is a necessary step in order for the virus to infect a cell. Thus, cleavage is required in order to convert new virus particles in a host cell into virions capable of infecting new cells. Cleavage is known to occur during transport of the integral HA0 membrane protein from the endoplasmic reticulum of the infected cell to the plasma membrane. In the course of transport, hemagglutinin undergoes a series of co- and post-translational modifications including proteolytic cleavage of the precursor HA into the amino-terminal fragment HA1 and the carboxy terminal HA2. One of the primary difficulties in growing influenza strains in primary tissue culture or established cell lines arises from the requirement for proteolytic cleavage activation of the influenza hemagglutinin in the host cell.
[0133] Although it is known that an uncleaved HA can mediate attachment of the virus to its neuraminic acid-containing receptors on a cell surface, it is not capable of the next step in the infectious cycle, which is fusion. It has been reported that exposure of the hydrophobic amino terminus of HA2 by cleavage is required so that it can be inserted into the target cell, thereby forming a bridge between virus and target cell membrane. This process is followed by fusion of the two membranes and entry of the virus into the target cell.
[0134] Proteolytic activation of HA involves cleavage at an arginine residue by a trypsin-like endoprotease, which is often an intracellular enzyme that is calcium dependent and has a neutral pH optimum. Since the activating proteases are cellular enzymes, the infected cell type determines whether the HA is cleaved. The HA of the mammalian influenza viruses and the nonpathogenic avian influenza viruses are susceptible to proteolytic cleavage only in a restricted number of cell types. On the other hand, HA of pathogenic avian viruses among the H5 and H7 subtypes are cleaved by proteases present in a broad range of different host cells. Thus, there are differences in host range resulting from differences in hemagglutinin cleavability which are correlated with the pathogenic properties of the virus.
[0135] Neuraminidase (NA) is a second membrane glycoprotein of the influenza viruses. The presence of viral NA has been shown to be important for generating a multi-faceted protective immune response against an infecting virus. For most influenza A viruses, NA is 413 amino acid in length, and is encoded by a gene of 1413 nucleotides. Nine different NA subtypes have been identified in influenza viruses (N1, N2, N3, N4, N5, N6, N7, N8 and N9), all of which have been found among wild birds. NA is involved in the destruction of the cellular receptor for the viral HA by cleaving terminal neuraminic acid (also called sialic acid) residues from carbohydrate moieties on the surfaces of infected cells. NA also cleaves sialic acid residues from viral proteins, preventing aggregation of viruses. Using this mechanism, it is hypothesized that NA facilitates release of viral progeny by preventing newly formed viral particles from accumulating along the cell membrane, as well as by promoting transportation of the virus through the mucus present on the mucosal surface. NA is an important antigenic determinant that is subject to antigenic variation.
[0136] In addition to the surface proteins HA and NA, influenza virus comprises six additional internal genes, which give rise to eight different proteins, including polymerase genes PB1, PB2 and PA, matrix proteins M1 and M2, nucleoprotein (NP), and non-structural proteins NS1 and NS2 (Horimoto et al., Clin Microbiol Rev. 14(1):129-149, 2001).
[0137] In order to be packaged into progeny virions, viral RNA is transported from the nucleus as a ribonucleoprotein (RNP) complex composed of the three influenza virus polymerase proteins, the nucleoprotein (NP), and the viral RNA, in association with the influenza virus matrix 1 (M1) protein and nuclear export protein (Marsh et al., J Virol, 82:2295-2304, 2008). The M1 protein that lies within the envelope is thought to function in assembly and budding. A limited number of M2 proteins are integrated into the virions (Zebedee, J. Virol. 62:2762-2772, 1988). They form tetramers having H+ ion channel activity, and when activated by the low pH in endosomes, acidify the inside of the virion, facilitating its uncoating (Pinto et al., Cell 69:517-528, 1992). Amantadine is an anti-influenza drug that prevents viral infection by interfering with M2 ion channel activity, thus inhibiting virus uncoating.
[0138] NS1, a nonstructural protein, has multiple functions, including regulation of splicing and nuclear export of cellular mRNAs as well as stimulation of translation. The major function of NS1 seems to be to counteract the interferon activity of the host, since an NS1 knockout virus was viable although it grew less efficiently than the parent virus in interferon-nondefective cells (Garcia-Sastre, Virology 252:324-330, 1998).
[0139] NS2 has been detected in virus particles (Richardson et al., Arch. Virol. 116:69-80, 1991; Yasuda et al., Virology 196:249-255, 1993). The average number of NS2 proteins in a virus particle was estimated to be 130-200 molecules. An in vitro binding assay shows direct protein-protein contact between M1 and NS2. NS2-M1 complexes have also been detected by immunoprecipitation in virus-infected cell lysates. The NS2 protein is thought to play a role in the export of RNP from the nucleus through interaction with M1 protein (Ward et al., Arch. Virol. 140:2067-2073, 1995).
V. Influenza Proteins, VLPs and Administration Thereof
[0140] Optimized influenza HA polypeptides and influenza VLPs comprising an optimized HA (such as the HA having the sequence set forth as SEQ ID NO: 2) are provided herein. The optimized HA polypeptides can be administered to elicit an immune response against influenza. In particular examples, the optimized HA polypeptides are administered as part of a VLP.
[0141] The influenza VLPs are made up of the HA, NA and M1 proteins. The production of influenza VLPs has been described in the art and is within the abilities of one of ordinary skill in the art. As described herein, influenza VLPs can be produced by transfection of host cells with plasmids encoding the HA, NA and M1 proteins. After incubation of the transfected cells for an appropriate time to allow for protein expression (such as for approximately 72 hours), VLPs can be isolated from cell culture supernatants. Example 1 below provides an exemplary protocol for purifying influenza VLPs from cell supernatants. In this example, VLPs are isolated by low speed centrifugation (to remove cell debris), vacuum filtration and ultracentrifugation through 20% glycerol.
[0142] The influenza VLPs disclosed herein can be used as influenza vaccines to elicit a protective immune response against H5N1 influenza viruses.
[0143] Influenza HA polypeptides and VLPs comprising HA polypeptides, or compositions thereof, can be administered to a subject by any of the routes normally used for introducing recombinant virus into a subject. Methods of administration include, but are not limited to, intradermal, intramuscular, intraperitoneal, parenteral, intravenous, subcutaneous, vaginal, rectal, intranasal, inhalation or oral. Parenteral administration, such as subcutaneous, intravenous or intramuscular administration, is generally achieved by injection. Injectables can be prepared in conventional forms, either as liquid solutions or suspensions, solid forms suitable for solution or suspension in liquid prior to injection, or as emulsions. Injection solutions and suspensions can be prepared from sterile powders, granules, and tablets of the kind previously described. Administration can be systemic or local.
[0144] Influenza VLPs, or compositions thereof, are administered in any suitable manner, such as with pharmaceutically acceptable carriers. Pharmaceutically acceptable carriers are determined in part by the particular composition being administered, as well as by the particular method used to administer the composition. Accordingly, there is a wide variety of suitable formulations of pharmaceutical compositions of the present disclosure.
[0145] Preparations for parenteral administration include sterile aqueous or non-aqueous solutions, suspensions, and emulsions. Examples of non-aqueous solvents are propylene glycol, polyethylene glycol, vegetable oils such as olive oil, and injectable organic esters such as ethyl oleate. Aqueous carriers include water, alcoholic/aqueous solutions, emulsions or suspensions, including saline and buffered media. Parenteral vehicles include sodium chloride solution, Ringer's dextrose, dextrose and sodium chloride, lactated Ringer's, or fixed oils. Intravenous vehicles include fluid and nutrient replenishers, electrolyte replenishers (such as those based on Ringer's dextrose), and the like. Preservatives and other additives may also be present such as, for example, antimicrobials, anti-oxidants, chelating agents, and inert gases and the like.
[0146] Some of the compositions may potentially be administered as a pharmaceutically acceptable acid- or base-addition salt, formed by reaction with inorganic acids such as hydrochloric acid, hydrobromic acid, perchloric acid, nitric acid, thiocyanic acid, sulfuric acid, and phosphoric acid, and organic acids such as formic acid, acetic acid, propionic acid, glycolic acid, lactic acid, pyruvic acid, oxalic acid, malonic acid, succinic acid, maleic acid, and fumaric acid, or by reaction with an inorganic base such as sodium hydroxide, ammonium hydroxide, potassium hydroxide, and organic bases such as mono-, di-, trialkyl and aryl amines and substituted ethanolamines.
[0147] Administration can be accomplished by single or multiple doses. The dose administered to a subject in the context of the present disclosure should be sufficient to induce a beneficial therapeutic response in a subject over time, or to inhibit or prevent H5N1 influenza virus infection. The dose required will vary from subject to subject depending on the species, age, weight and general condition of the subject, the severity of the infection being treated, the particular composition being used and its mode of administration. An appropriate dose can be determined by one of ordinary skill in the art using only routine experimentation.
[0148] Provided herein are pharmaceutical compositions which include a therapeutically effective amount of the influenza VLPs alone or in combination with a pharmaceutically acceptable carrier. Pharmaceutically acceptable carriers include, but are not limited to, saline, buffered saline, dextrose, water, glycerol, ethanol, and combinations thereof. The carrier and composition can be sterile, and the formulation suits the mode of administration. The composition can also contain minor amounts of wetting or emulsifying agents, or pH buffering agents. The composition can be a liquid solution, suspension, emulsion, tablet, pill, capsule, sustained release formulation, or powder. The composition can be formulated as a suppository, with traditional binders and carriers such as triglycerides. Oral formulations can include standard carriers such as pharmaceutical grades of mannitol, lactose, starch, magnesium stearate, sodium saccharine, cellulose, and magnesium carbonate. Any of the common pharmaceutical carriers, such as sterile saline solution or sesame oil, can be used. The medium can also contain conventional pharmaceutical adjunct materials such as, for example, pharmaceutically acceptable salts to adjust the osmotic pressure, buffers, preservatives and the like. Other media that can be used with the compositions and methods provided herein are normal saline and sesame oil.
[0149] The influenza VLPs described herein can be administered alone or in combination with other therapeutic agents to enhance antigenicity. For example, the influenza VLPs can be administered with an adjuvant, such as Freund incomplete adjuvant or Freund's complete adjuvant.
[0150] Optionally, one or more cytokines, such as IL-2, IL-6, IL-12, RANTES, GM-CSF, TNF-.alpha., or IFN-.gamma., one or more growth factors, such as GM-CSF or G-CSF; one or more molecules such as OX-40L or 41 BBL, or combinations of these molecules, can be used as biological adjuvants (see, for example, Salgaller et al., 1998, J. Surg. Oncol. 68(2):122-38; Lotze et al., 2000, Cancer J. Sci. Am. 6(Suppl 1):S61-6; Cao et al., 1998, Stem Cells 16(Suppl 1):251-60; Kuiper et al., 2000, Adv. Exp. Med. Biol. 465:381-90). These molecules can be administered systemically (or locally) to the host.
[0151] A number of means for inducing cellular responses, both in vitro and in vivo, are known. Lipids have been identified as agents capable of assisting in priming CTL in vivo against various antigens. For example, as described in U.S. Pat. No. 5,662,907, palmitic acid residues can be attached to the alpha and epsilon amino groups of a lysine residue and then linked (for example, via one or more linking residues, such as glycine, glycine-glycine, serine, serine-serine, or the like) to an immunogenic peptide. The lipidated peptide can then be injected directly in a micellar form, incorporated in a liposome, or emulsified in an adjuvant. As another example, E. coli lipoproteins, such as tripalmitoyl-S-glycerylcysteinlyseryl-serine can be used to prime tumor specific CTL when covalently attached to an appropriate peptide (see, Deres et al., Nature 342:561, 1989). Further, as the induction of neutralizing antibodies can also be primed with the same molecule conjugated to a peptide which displays an appropriate epitope, two compositions can be combined to elicit both humoral and cell-mediated responses where that is deemed desirable.
[0152] Although administration of VLPs containing the optimized HA protein, one of skill in the art would understand that it is also possible to administer the optimized influenza HA protein itself (in the absence of a viral particle) or as a fusion protein to elicit an immune response in a subject.
[0153] The following examples are provided to illustrate certain particular features and/or embodiments. These examples should not be construed to limit the disclosure to the particular features or embodiments described.
EXAMPLES
Example 1: A Computationally Optimized Broadly Reactive Antigen (COBRA) Based H5N1 VLP Vaccine Elicits Broadly Reactive Antibodies in Mice and Ferrets
[0154] This example describes the finding that mice and ferrets vaccinated with COBRA hemagglutinin (HA) H5N1 VLPs exhibited protective levels of HAI antibodies to representative isolates from each sub-clade of clade 2 and were completely protected from lethal challenge with a clade 2.2 H5N1 virus.
Materials and Methods
COBRA Hemagglutinin (HA) Construction and Synthesis
[0155] Influenza A HA amino acid sequences isolated from human H5N1 infections were downloaded from the NCBI Influenza Virus Resource database (Bao et al., J Virol 82:596-601, 2008; see Table 1 for a complete list of accession numbers and isolate descriptions). Nucleotide sequences were translated into protein sequences using the standard genetic code. All available full length sequences from H5N1 clade 2 human infections from 2004 to 2006 were acquired and used for subsequent consensus generations. For each round of consensus generation, multiple alignment analysis was applied and the consensus sequence was generated using AlignX (Vector NTI). The final amino acid sequence, termed computationally optimized broadly reactive antigen (COBRA), was reverse translated and optimized for expression in mammalian cells, including codon usage and RNA optimization (GeneArt; Regensburg, Germany). This construct was then synthesized and inserted into the pTR600 expression vector (U.S. Patent Application Publication No. 2002/0106798; Ross et al., Nat Immunol. 1(2):102-103, 2000; Green et al., Vaccine 20:242-248, 2001).
TABLE-US-00002 TABLE 1 COBRA input sequences Strain Clade Accession Host Country Year A/Indonesia/534H/2006 2.1.2 EU146737 Human Indonesia 2006 A/Indonesia/535H/2006 2.1.2 EU146753 Human Indonesia 2006 A/Indonesia/536H/2006 2.1.2 EU146754 Human Indonesia 2006 A/Indonesia/538H/2006 2.1.2 EU146745 Human Indonesia 2006 A/Indonesia/546bH/2006 2.1.2 EU146793 Human Indonesia 2006 A/Indonesia/546H/2006 2.1.2 EU146755 Human Indonesia 2006 A/Indonesia/560H/2006 2.1.2 EU146785 Human Indonesia 2006 A/Indonesia/CDC594/2006 2.1.2 CY014272 Human Indonesia 2006 A/Indonesia/CDC595/2006 2.1.2 CY014280 Human Indonesia 2006 A/Indonesia/CDC596/2006 2.1.2 CY014288 Human Indonesia 2006 A/Indonesia/CDC597/2006 2.1.2 CY014296 Human Indonesia 2006 A/Indonesia/CDC599/2006 2.1.2 CY014303 Human Indonesia 2006 A/Indonesia/CDC599N/2006 2.1.2 CY014477 Human Indonesia 2006 A/Indonesia/CDC625/2006 2.1.2 CY014433 Human Indonesia 2006 A/Indonesia/CDC625L/2006 2.1.2 CY014465 Human Indonesia 2006 A/Indonesia/160H/2005 2.1.3 EU146648 Human Indonesia 2005 A/Indonesia/175H/2005 2.1.3 EU146640 Human Indonesia 2005 A/Indonesia/177H/2005 2.1.3 EU146680 Human Indonesia 2005 A/Indonesia/195H/2005 2.1.3 EU146656 Human Indonesia 2005 A/Indonesia/239H/2005 2.1.3 EU146664 Human Indonesia 2005 A/Indonesia/245H/2005 2.1.3 EU146672 Human Indonesia 2005 A/Indonesia/283H/2006 2.1.3 EU146681 Human Indonesia 2006 A/Indonesia/286H/2006 2.1.3 EU146688 Human Indonesia 2006 A/Indonesia/292H/2006 2.1.3 EU146713 Human Indonesia 2006 A/Indonesia/298H/2006 2.1.3 EU146697 Human Indonesia 2006 A/Indonesia/304H/2006 2.1.3 EU146705 Human Indonesia 2006 A/Indonesia/321H/2006 2.1.3 EU146721 Human Indonesia 2006 A/Indonesia/341H/2006 2.1.3 EU146729 Human Indonesia 2006 A/Indonesia/5/2005 2.1.3 EF541394 Human Indonesia 2005 A/Indonesia/542H/2006 2.1.3 EU146777 Human Indonesia 2006 A/Indonesia/567H/2006 2.1.3 EU146801 Human Indonesia 2006 A/Indonesia/569H/2006 2.1.3 EU146809 Human Indonesia 2006 A/Indonesia/583H/2006 2.1.3 EU146817 Human Indonesia 2006 A/Indonesia/604H/2006 2.1.3 EU146825 Human Indonesia 2006 A/Indonesia/7/2005 2.1.3 EU146632 Human Indonesia 2005 A/Indonesia/CDC184/2005 2.1.3 CY014197 Human Indonesia 2005 A/Indonesia/CDC194P/2005 2.1.3 CY014168 Human Indonesia 2005 A/Indonesia/CDC287E/2005 2.1.3 CY014198 Human Indonesia 2005 A/Indonesia/CDC287T/2005 2.1.3 CY014199 Human Indonesia 2005 A/Indonesia/CDC292N/2005 2.1.3 CY014200 Human Indonesia 2005 A/Indonesia/CDC292T/2005 2.1.3 CY014201 Human Indonesia 2005 A/Indonesia/CDC326/2006 2.1.3 CY014204 Human Indonesia 2006 A/Indonesia/CDC326N/2006 2.1.3 CY014202 Human Indonesia 2006 A/Indonesia/CDC326N2/2006 2.1.3 CY014203 Human Indonesia 2006 A/Indonesia/CDC326T/2006 2.1.3 CY014205 Human Indonesia 2006 A/Indonesia/CDC329/2006 2.1.3 CY014206 Human Indonesia 2006 A/Indonesia/CDC357/2006 2.1.3 CY014207 Human Indonesia 2006 A/Indonesia/CDC370/2006 2.1.3 CY014209 Human Indonesia 2006 A/Indonesia/CDC370E/2006 2.1.3 CY014210 Human Indonesia 2006 A/Indonesia/CDC370P/2006 2.1.3 CY014211 Human Indonesia 2006 A/Indonesia/CDC370T/2006 2.1.3 CY014212 Human Indonesia 2006 A/Indonesia/CDC390/2006 2.1.3 CY014213 Human Indonesia 2006 A/Indonesia/CDC523/2006 2.1.3 CY014311 Human Indonesia 2006 A/Indonesia/CDC523E/2006 2.1.3 CY014368 Human Indonesia 2006 A/Indonesia/CDC523T/2006 2.1.3 CY014376 Human Indonesia 2006 A/Indonesia/CDC582/2006 2.1.3 CY014384 Human Indonesia 2006 A/Indonesia/CDC610/2006 2.1.3 CY014393 Human Indonesia 2006 A/Indonesia/CDC623/2006 2.1.3 CY014401 Human Indonesia 2006 A/Indonesia/CDC623E/2006 2.1.3 CY014409 Human Indonesia 2006 A/Indonesia/CDC624/2006 2.1.3 CY014417 Human Indonesia 2006 A/Indonesia/CDC624E/2006 2.1.3 CY014425 Human Indonesia 2006 A/Indonesia/CDC634/2006 2.1.3 CY014441 Human Indonesia 2006 A/Indonesia/CDC634P/2006 2.1.3 CY014449 Human Indonesia 2006 A/Indonesia/CDC634T/2006 2.1.3 CY014457 Human Indonesia 2006 A/Indonesia/CDC644/2006 2.1.3 CY014518 Human Indonesia 2006 A/Indonesia/CDC644T/2006 2.1.3 CY014510 Human Indonesia 2006 A/Indonesia/CDC669/2006 2.1.3 CY014481 Human Indonesia 2006 A/Indonesia/CDC669P/2006 2.1.3 CY014489 Human Indonesia 2006 A/Indonesia/CDC699/2006 2.1.3 CY014497 Human Indonesia 2006 A/Indonesia/CDC7/2005 2.1.3 CY014177 Human Indonesia 2005 A/Indonesia/CDC739/2006 2.1.3 CY014529 Human Indonesia 2006 A/Indonesia/CDC759/2006 2.1.3 CY014543 Human Indonesia 2006 A/Indonesia/CDC835/2006 2.1.3 CY017662 Human Indonesia 2006 A/Indonesia/CDC836/2006 2.1.3 CY017670 Human Indonesia 2006 A/Indonesia/CDC836T/2006 2.1.3 CY017678 Human Indonesia 2006 A/Indonesia/CDC887/2006 2.1.3 CY017688 Human Indonesia 2006 A/Indonesia/CDC938/2006 2.1.3 CY017638 Human Indonesia 2006 A/Indonesia/CDC938E/2006 2.1.3 CY017646 Human Indonesia 2006 A/Indonesia/CDC940/2006 2.1.3 CY017654 Human Indonesia 2006 A/Indonesia/TLL001/2006 2.1.3 EU015403 Human Indonesia 2006 A/Indonesia/TLL002/2006 2.1.3 EU015404 Human Indonesia 2006 A/Indonesia/TLL003/2006 2.1.3 EU015405 Human Indonesia 2006 A/Indonesia/TLL004/2006 2.1.3 EU015406 Human Indonesia 2006 A/Indonesia/TLL005/2006 2.1.3 EU015407 Human Indonesia 2006 A/Indonesia/TLL006/2006 2.1.3 EU015408 Human Indonesia 2006 A/Indonesia/TLL007/2006 2.1.3 EU015409 Human Indonesia 2006 A/Indonesia/TLL008/2006 2.1.3 EU015410 Human Indonesia 2006 A/Indonesia/TLL009/2006 2.1.3 EU015411 Human Indonesia 2006 A/Indonesia/TLL010/2006 2.1.3 EU015412 Human Indonesia 2006 A/Indonesia/TLL011/2006 2.1.3 EU015413 Human Indonesia 2006 A/Indonesia/TLL012/2006 2.1.3 EU015414 Human Indonesia 2006 A/Indonesia/TLL013/2006 2.1.3 EU015415 Human Indonesia 2006 A/Indonesia/TLL014/2006 2.1.3 EU015416 Human Indonesia 2006 A/Djibouti/5691NAMRU3/2006 2.2 DQ666146 Human Djibouti 2006 A/Egypt/7021-NAMRU3/2006 2.2 CY062439 Human Egypt 2006 A/human/Iraq/207-NAMRU3/2006 2.2 DQ435202 Human Iraq 2006 A/Iraq/1/2006 2.2 EU146870 Human Iraq 2006 A/Iraq/659/2006 2.2 EU146876 Human Iraq 2006 A/Iraq/754/2006 2.2 EU146877 Human Iraq 2006 A/Iraq/755/2006 2.2 EU146869 Human Iraq 2006 A/Iraq/756/2006 2.2 EU146878 Human Iraq 2006 A/Turkey/12/2006 2.2 EF619982 Human Turkey 2006 A/Turkey/15/2006 2.2 EF619989 Human Turkey 2006 A/Turkey/651242/2006 2.2 EF619990 Human Turkey 2006 A/Turkey/65596/2006 2.2 EF619998 Human Turkey 2006 A/Xinjiang/1/2006 2.2 FJ492886 Human China 2006 A/Egypt/14724-NAMRU3/2006 2.2.1 EF200512 Human Egypt 2006 A/Egypt/14725-NAMRU3/2006 2.2.1 EF200513 Human Egypt 2006 A/Egypt/2782-NAMRU3/2006 2.2.1 DQ464377 Human Egypt 2006 A/Egypt/2991-NAMRU3/2006 2.2.1 EU095023 Human Egypt 2006 A/Egypt/2992-NAMRU3/2006 2.2.1 EU095024 Human Egypt 2006 A/Egypt/902782/2006 2.2.1 EU146867 Human Egypt 2006 A/Egypt/902786/2006 2.2.1 EU146868 Human Egypt 2006 A/Anhui/1/2005 2.3.4 DQ371928 Human China 2005 A/Anhui/2/2005 2.3.4 DQ371929 Human China 2005 A/China/2006 2.3.4 EF624256 Human China 2006 A/China/GD01/2006 2.3.4 DQ835313 Human China 2006 A/Fujian/1/2005 2.3.4 FJ492882 Human China 2005 A/Guangdong/1/2006 2.3.4 FJ492884 Human China 2006 A/Guangxi/1/2005 2.3.4 DQ371930 Human China 2005 A/human/China/GD02/2006 2.3.4 EU263981 Human China 2006 A/Hunan/1/2006 2.3.4 FJ492879 Human China 2006 A/Jiangxi/1/2005 2.3.4 FJ492885 Human China 2005 A/Shanghai/1/2006 2.3.4 AB462295 Human China 2006 A/Shenzhen/406H/2006 2.3.4 EF137706 Human China 2006 A/Sichuan/1/2006 2.3.4 FJ492881 Human China 2006 A/Vietnam/UT30850/2005 2.3.4 HM114537 Human Viet Nam 2005 A/Zhejiang/1/2006 2.3.4 FJ492880 Human China 2006 A/Zhejiang/16/2006 2.3.4 DQ643809 Human China 2006
COBRA HA Antigenic Modeling
[0156] Influenza hemagglutinin (HA) protein sequences representing COBRA, A/Indonesia/5/2005 (Clade 2.1), A/Whooper Swan/Mongolia/244/2005 (Clade 2.2) and A/Anhui/1/2005 (Clade 2.3) were submitted to the 3D-JIGSAW Protein Comparative Modeling website for rendering (Bates et al., Proteins 45(55):39-46, 2001; Bates and Sternberg, Proteins 37(53):47-54, 1999; Contreras-Moreira and Bates, Bioinformatics 18(8):1141-1142, 2002). Structures were overlaid and analyzed using Swiss-Pdb viewer software (Guex and Peitsch, Electrophoresis 18(15):2714-23, 1998). Antigenic regions and designations are based on the original description of the antigenic structure of the H3N2 virus A/Hong Kong/1/1968 (Wiley et al., Nature 289(5796):373-378, 1981). No significant alterations were observed in region B of the COBRA HA relative to the primary influenza isolates; however, some divergent structures in HA regions A and C were identified in primary isolates.
In Vitro Expression
[0157] COBRA HA protein expression was confirmed by transfecting mammalian cells. Human embryonic kidney (HEK) 293T cells (1.times.10.sup.6) were transiently transfected with 3 .mu.g of DNA expressing COBRA. Cells were incubated for 72 hours at 37.degree. C., supernatants were removed, the cells were lysed with 1% Triton-X 100 and cell lysates were collected. Cell lysates were electrophoresed on a 10% SDS-PAGE gel and transferred to a PVDF membrane. The blot was probed with mouse polyclonal antisera pooled from mice infected with 6:2 reassortant H5N1 viruses with the surface glycoproteins derived from either A/Vietnam/1203/2004 or A/Whooper Swan/244/2005, and the HA-antibody complexes were detected using a goat anti-mouse IgG conjugated to horse radish peroxidase (HRP) (Southern Biotech; Birmingham, Ala., USA). HRP was detected by chemiluminescent substrate (Pierce Biotechnology; Rockford Ill., USA) and exposed to X-ray film (ThermoFisher; Pittsburgh, Pa., USA).
COBRA HA Functional Characterization
[0158] To determine receptor binding characteristics, virus-like particles (VLPs) containing COBRA HA proteins were purified from the supernatants of mammalian cell lines. HEK 293T cells were transiently transfected with plasmids expressing HIV Gag, COBRA HA and neuraminidase (NA, A/Thailand/1(KAN-1)/2004) and incubated for 72 hours at 37.degree. C. Supernatants were collected and VLPs were purified via ultracentrifugation (100,000.times.g through 20% glycerol, weight per volume) for 4 hours at 4.degree. C. The pellets were subsequently resuspended in phosphate buffered saline PBS, pH 7.2 and stored at -80.degree. C. until use. Protein concentration was determined by Micro BCA.TM. Protein Assay Reagent Kit (Pierce Biotechnology, Rockford, Ill., USA). COBRA HA VLPs were prepared in various amounts as measured by total protein and each individual preparation was two-fold serially diluted in v-bottom microtiter plates. An equal volume of either 1% turkey or 1% horse erythrocytes (RBC) (Lampire; Pipersville, Pa., USA) in PBS was added to the diluted VLPs and incubated for 30-60 minutes at room temperature. The HA titer was determined by the reciprocal dilution of the last well which contained agglutinated RBC.
[0159] To determine endosomal fusion characteristics, COBRA-pseudotyped lentiviral vectors expressing a luciferase reporter gene were produced as described (Yang et al., J Virol 78(8):4029-4036). Briefly, 293T cells were co-transfected by using the following plasmids: 7 .mu.g of pCMVdeltaR8.2, 7 .mu.g of pHRCMV-Luc, 3 .mu.g pCMV/R N1(Kan-1) and 3 .mu.g pTR600 COBRA. Cells were transiently transfected and incubated for 72 hours at 37.degree. C. Supernatants were harvested, centrifuged to clear cell debris, filtered through a 0.22 .mu.m syringe filter, aliquotted, and used immediately or frozen at -80.degree. C. For fusion assays, 100 .mu.l of pseudoviruses were added to confluent MDCK cells in 48-well plates (30,000 cells per well). Plates were incubated at room temperature for 30 minutes, washed, and fresh medium added. Forty-eight hours after infection, cells were lysed in mammalian cell lysis buffer (Promega; Madison, Wis., USA). A standard quantity of cell lysate was used in a luciferase assay with luciferase assay reagent (Promega; Madison, Wis., USA) according to the manufacturer's protocol.
Vaccine Preparation and Dose Determination
[0160] HEK 293T cells were transiently transfected with plasmids expressing M1 (A/Puerto Rico/8/1934, optimized for expression in mammalian cells; SEQ ID NO: 9), NA (A/Thailand/1(KAN-1)/2004, optimized for expression in mammalian cells; SEQ ID NO: 8) and COBRA HA (SEQ ID NO: 7) and incubated for 72 hours at 37.degree. C. Supernatants were collected and cell debris removed by low speed centrifugation followed by vacuum filtration through a 0.22 .mu.m sterile filter. VLPs were purified via ultracentrifugation (100,000.times.g through 20% glycerol, weight per volume) for 4 hours at 4.degree. C. The pellets were subsequently resuspended in PBS pH 7.2 and stored in single use aliquots at -80.degree. C. until use. Total protein concentration was determined by Micro BCA.TM. Protein Assay Reagent Kit (Pierce Biotechnology, Rockford, Ill., USA).
[0161] HA specific content was determined by western blot and densitometry. Purified recombinant COBRA HA and purified VLPs were prepared in standard total protein amounts and were electrophoresed on a 10% SDS-PAGE gel and transferred to a PVDF membrane. The blot was probed with mouse polyclonal antisera pooled from mice infected with 6:2 reassortant H5N1 viruses with the surface glycoproteins derived from either A/Vietnam/1203/2004 or A/Whooper Swan/244/2005 and the HA-antibody complexes were detected using a goat anti-mouse IgG conjugated to horse radish peroxidase (HRP) (Southern Biotech; Birmingham, Ala., USA). HRP was detected by chemiluminescent substrate (Pierce Biotechnology; Rockford Ill., USA) and exposed to X-ray film (ThermoFisher; Pittsburgh, Pa., USA). Density of bands was determined using ImageJ software (NIH) (Abramoff et al., Biophotonics International 11(7):36-42, 2004). Density of recombinant HA bands were used to calculate a standard curve and the density of the purified VLPs was interpolated using the results from the recombinant HA. Experiments were performed in triplicate and multiple exposure times were analyzed for all iterations.
Codon-Optimized Influenza HA, NA and M1 Genes
[0162] The nucleotide sequences of the codon-optimized HA (SEQ ID NO: 1), codon-optimized NA (SEQ ID NO: 3) and codon-optimized M1 (SEQ ID NO: 5) genes are set forth in the Sequence Listing. The corresponding amino acid sequences of the encoded HA, NA and M1 proteins are set forth in the Sequence Listing as SEQ ID NO: 2, SEQ ID NO: 4 and SEQ ID NO: 6, respectively.
Mouse Studies
[0163] BALB/c mice (Mus musculis, females, 6-8 weeks old) were purchased from Harlan Sprague Dawley (Indianapolis, Ind., USA) and housed in microisolator units and allowed free access to food and water and were cared for under USDA guidelines for laboratory animals. Mice (5 mice per group) were vaccinated with one of three doses of purified COBRA HA VLPs (1.5 .mu.g, 0.3 .mu.g or 0.06 .mu.g), based upon HA content from a densitometry assay, via intramuscular injection at week 0 and then boosted with the same dose at week 3. For comparison studies, mice (20 mice per group) were vaccinated with purified VLPs (3 .mu.g) via intramuscular injection at week 0 and then boosted with the same dose at week 3. Vaccines at each dose were formulated with Imject.RTM. alum adjuvant (Imject.RTM. Alum, Pierce Biotechnology; Rockford, Ill., USA) according to the manufacturer's protocol or vehicle alone. Fourteen to twenty-one days after each vaccination, blood was collected from anesthetized mice via the retro-orbital plexus and transferred to a microfuge tube. Tubes were centrifuged and sera was removed and frozen at -20.+-.5.degree. C.
[0164] Three weeks after final vaccination, mice were challenged intranasally with 5.times.10.sup.3 plaque forming units (PFU) of the highly pathogenic H5N1 virus A/Whooper Swan/Mongolia/244/2005 (clade 2.2) in a volume of 50 .mu.l. The challenge dose represents approximately 50LD.sub.50 in mice. After infection, mice were monitored daily for weight loss, disease signs and death for 14 days after infection. Individual body weights, sickness scores and death were recorded for each group on each day after inoculation. Sickness score was determined by evaluating activity (0=normal, 1=reduced, 2=severely reduced), hunched back (0=absent, 1=present) and ruffled fur (0=absent, 1=present) (Toapanta and Ross, Respiratory Res 10(1):112, 2009). Experimental endpoint was defined as >20% weight loss or display of neurological disease such as hind limb paralysis. All H5N1 influenza virus studies were performed under high-containment biosafety level 3 enhanced conditions (BSL3+).
Ferret Studies
[0165] Fitch ferrets (Mustela putorius faro, female, 6-12-months of age), influenza naive and descented, were purchased from Marshall Farms (Sayre, Pa., USA). Ferrets were pair housed in stainless steel cages (Shor-line, Kansas City, Kans., USA) containing Sani-chips Laboratory Animal Bedding (P.J. Murphy Forest Products, Montville, N.J., USA). Ferrets were provided with Teklad Global Ferret Diet (Harlan Teklad, Madison, Wis., USA) and fresh water ad libitum. The COBRA HA VLPs were diluted in PBS, pH 7.2 to achieve final concentration. Ferrets (n=3) were vaccinated with 15 .mu.g of purified COBRA VLPs, based upon HA content as determined by densitometry assay, via intramuscular injection in the quadriceps muscle in a volume of 0.25 ml at week 0 and then boosted with the same dose at week 3. Vaccines were stored at -80.degree. C. prior to use and formulated with Imject.RTM. alum adjuvant (Imject.RTM. Alum; Pierce Biotechnology, Rockford, Ill., USA) immediately prior to use. Animals were monitored for adverse events including weight loss, temperature, loss of activity, nasal discharge, sneezing and diarrhea weekly during the vaccination regimen. Prior to vaccination, animals were confirmed by HAI assay to be seronegative for circulating influenza A (H1N1 and H3N2) and influenza B viruses. Fourteen to twenty-one days after each vaccination, blood was collected from anesthetized ferrets via the anterior vena cava and transferred to a microfuge tube. Tubes were centrifuged and sera was removed and frozen at -20.+-.5.degree. C.
[0166] Three weeks after final vaccination, ferrets were challenged intranasally with 1.times.10.sup.6 plaque forming units (PFU) of the highly pathogenic H5N1 virus A/Whooper Swan/Mongolia/244/2005 (clade 2.2) in a volume of 0.5 ml in each nostril for a total infection volume of 1 ml. After infection, ferrets were monitored daily for weight loss, disease signs and death for 14 days after infection. Individual body weights, sickness scores, and death were recorded for each group on each day after inoculation. Sickness score was determined by evaluating activity (0=normal, 1=alert and active with stimulation, 2=alert but not active after stimulation, 3=not alert or active after stimulation), nasal discharge (0=absent, 1=present), sneezing (0=absent, 1=present), decreased food intake (0=absent, 1=present), diarrhea (0=absent, 1=present), dyspnea (0=absent, 1=present) and neurological symptoms (0=absent, 1=present). Nasal washes were performed by instilling 3 ml of PBS into the nares of anesthetized ferrets each day for 14 days after inoculation. Washes were collected and stored at -80.degree. C. until use. Experimental endpoint was defined as >20% weight loss, development of neurological symptoms, or an activity score of 3 (not active or alert after stimulation). All H5N1 influenza virus studies were performed under high-containment biosafety level 3 enhanced conditions (BSL3+).
ELISA
[0167] The ELISA assay was used to assess total antibody titer and IgG isotype titer to the HA. High binding, 96-well polystyrene plates (Costar; Lowell, Mass., USA) were coated overnight with 50 ng/well of recombinant HA. Coating antigens were derived from the following representative viral isolates: A/Vietnam/1203/2004 (clade 1), A/Indonesia/5/2005 (clade 2.1), A/Whooper Swan/244/2005 (clade 2.2) and A/Anhui/1/2005 (clade 2.3). Plates were blocked with 5% milk diluted in PBS with 0.05% Tween 20. Serum samples were diluted in blocking buffer and added to plates. Serum was two-fold serially diluted and allowed to incubate for 1 hour at room temperature. Plates were washed and species specific antibody against IgG, IgG1, IgG2a, IgG2b or IgG3 and linked to horseradish peroxidase (HRP) (Southern Biotech; Birmingham, Ala., USA) were diluted in blocking buffer and added to plates. Plates were incubated for 1 hour at room temperature. Plates were washed and HRP was developed with TMB substrate (Sigma-Aldrich; St. Louis, Mo., USA). Plates were incubated in the dark for 15 minutes and then the reaction was stopped with 2N H.sub.2SO.sub.4. Optical densities at a wavelength of 450 nm (OD.sub.450) were read by a spectrophotometer (BioTek; Winooski, Vt., USA) and end point dilution titers were determined. End point titers were determined as the reciprocal dilution of the last well which had an OD.sub.450 above the mean OD.sub.450 plus two standard deviations of naive animal sera.
Hemagglutination Inhibition (HAI)
[0168] The HAI assay was used to assess functional antibodies to HA able to inhibit agglutination of horse erythrocytes. The protocol was adapted from the CDC laboratory-based influenza surveillance manual (Gillim-Ross and Subbarao, Clin Microbiol Rev 19(4):614-636, 2006). To inactivate non-specific inhibitors, sera were treated with receptor destroying enzyme (RDE; Denka Seiken, Co., Japan) prior to being tested (Bright et al., Lancet 366(9492):1175-1181, 2005; Bright et al., Virology 308(2):270-278, 2003; Bright et al., JAMA 295(8):891-894, 2006; Mitchell et al., Vaccine 21(9-10):902-914, 2004; Ross et al., Nat Immunol 1(2):127-131, 2000). Briefly, three parts RDE was added to one part sera and incubated overnight at 37.degree. C. RDE was inactivated by incubation at 56.degree. C. for .about.30 minutes. RDE-treated sera was two-fold serially diluted in v-bottom microtiter plates. An equal volume of reassortant virus, adjusted to approximately 8 HAU/50 .mu.l, was added to each well. The reassortant viruses contained the internal genes from the mouse adapted strain A/Puerto Rico/8/1934 and the surface proteins HA and NA from the following representative viral isolates: A/Vietnam/1203/2004 (clade 1), A/Indonesia/5/2005 (clade 2.1), A/Whooper Swan/244/2005 (clade 2.2) and A/Anhui/1/2005 (clade 2.3). The plates were covered and incubated at room temperature for 20 minutes followed by the addition of 1% horse erythrocytes (hRBC) (Lampire Biologicals, Pipersville, Pa., USA) in PBS. Red blood cells were stored at 4.degree. C. and used within 72 hours of preparation. The plates were mixed by agitation, covered, and the RBCs were allowed to settle for 1 hour at room temperature (Askonas B, McMichael A, Webster R. The immune response to influenza viruses and the problem of protection against infection. In: Beare AS, editor. Basic and applied influenza research: CRC Press 1982: 159-188). The HAI titer was determined by the reciprocal dilution of the last row which contained non-agglutinated RBCs. Positive and negative serum controls were included for each plate. All mice were negative (HAI.ltoreq.1:10) for pre-existing antibodies to currently circulating human influenza viruses prior to vaccination.
Plaque Assay
[0169] Madin-Darby Canine Kidney (MDCK) cells were plated (5.times.10.sup.5) in each well of a 6-well plate. Samples were diluted (final dilution factors of 10.degree. to 10.sup.-6) and overlayed onto the cells in 100 .mu.l of DMEM supplemented with penicillin-streptomycin and incubated for 1 hour. Samples were removed, cells were washed twice and media was replaced with 2 ml of L15 medium plus 0.8% agarose (Cambrex; East Rutherford, N.J., USA) and incubated for 72 hours at 37.degree. C. with 5% CO.sub.2. Agarose was removed and discarded. Cells were fixed with 10% buffered formalin, and then stained with 1% crystal violet for 15 minutes. Following thorough washing in dH.sub.2O to remove excess crystal violet, plates were allowed to dry, plaques counted, and the plaque forming units (PFU)/ml were calculated.
Statistical Analysis
[0170] Statistical significance of the antibody data was determined using a two-way analysis of variance (ANOVA) with Bonferroni's post-test to analyze differences between each vaccine group for the different test antigens (multiparametric). Differences in weight loss, sickness score, and viral titers were analyzed by two-way ANOVA, followed by Bonferroni's post test for each vaccine group at multiple time points. Significance was defined asp<0.05. Statistical analyses were done using GraphPad Prism software.
Results
Computationally Optimized Broadly Reactive Antigen Design
[0171] To address the challenge of antigenic diversity present in H5N1 influenza, a computationally optimized broadly reactive antigen (COBRA) was designed. For the first step of antigen generation, 129 unique hemagglutinin (HA) sequences were downloaded from the NCBI Influenza Virus Resource (IVR) sequence database (Bao et al., J Virol 82:596-601, 2008) representing clade 2 H5N1 viruses isolated from human infections between 2004 and 2006. The sequences were first grouped into phylogenetic sub-clades and then further divided into individual outbreak groups within each sub-clade based upon geographic location and time of isolation. HA amino acid sequences for each individual outbreak group were aligned and the most common amino acid at each position was determined resulting in primary consensus sequences representing each outbreak group within each sub-clade (FIG. 1A). Primary consensus sequences within each sub-clade were then aligned and the most common amino acid was chosen resulting in secondary consensus sequences representing each sub-clade (FIG. 1A). The secondary consensus sequences were aligned and the most common amino acid at each position was selected resulting in the final consensus sequence referred to as clade 2 COBRA HA (FIG. 1A). Phylogenetic analysis of the COBRA HA with all human isolates of H5N1 HA proteins indicated that COBRA retained a clade 2-like sequence without being grouped specifically within any clade 2 sub-clade cluster (FIG. 1B). Furthermore, a BLAST search using the COBRA HA sequence revealed that it is a unique sequence that has not been isolated from the environment.
Characterization of COBRA
[0172] Since COBRA is a fully synthetic protein, the retention of natural hemagglutinin function was confirmed. Initially, COBRA expression was verified by transient transfection of mammalian cells. Analysis of the total cell lysate demonstrated that the COBRA HA migrates at its predicted molecular weight of approximately 73 kDa (FIG. 2A). Because the poly-basic cleavage site was retained in the COBRA HA sequence, both HA0 and the HA1 subunits were detected by immunoblot at similar molecular weights as recombinant HA and the HA on the H5N1 virion (FIG. 2A). Virus-like particles (VLPs) with COBRA HA on the surface bound sialic acid in a dose-dependent manner and this binding was specific to COBRA, since empty lentiviral core alone did not bind to the receptor (FIG. 2B).
[0173] To determine if the COBRA HA was functional, the protein was pseudotyped onto lentiviral Gag.sub.p24 to generate pseudoparticles (Nefkens et al., J Clin Virol 39(1):27-33, 2007; Haynes et al., Vaccine 27(4):530-541, 2009). COBRA HA containing pseudoparticles mediated cell fusion as efficiently as H5N1 control pseudoparticles without the requirement for trypsin. In contrast, H1N1 pseudoparticles did require trypsin and pseudoparticles without surface HA produced luciferase at similar levels as the cell only controls (FIG. 2C). Taken together, these results demonstrate that although the COBRA HA is a synthetic protein not found in nature, it retains all of the functions of a natural hemagglutinin protein.
Mouse Dosing Immunizations
[0174] Mice (BALB/c; n=5) were vaccinated (week 0 and 3) via intramuscular injection with purified COBRA VLPs at either a high dose (1.5 .mu.g HA) or low dose (0.3 .mu.g HA) with and without Imject.RTM. alum adjuvant. At week 5, all COBRA VLP-vaccinated animals had anti-HA antibodies that recognized heterologous recombinant HA derived from both clade 1 and also sub-clades of clade 2 (FIGS. 3A and 3B). Imject.RTM. alum significantly increased anti-HA antibody titers in both low and high dose groups as compared to the non-adjuvanted groups (p<0.01). The IgG isotype subclasses elicited by the VLP vaccines against a clade 2.1 coating antigen consisted mainly of IgG1 and IgG2a, indicating a mixed T helper response (FIGS. 3C and 3D). Similar results were found for additional coating antigens representing clade 1, clade 2.2 and clade 2.3. In addition to recognizing HA, antibodies were also evaluated for the ability to block virus from binding its receptor via inhibition of viral-induced agglutination of horse erythrocytes (HAI). All mice administered Imject.RTM. alum adjuvanted vaccines, regardless of dose, had HAI titers.gtoreq.1:40 to viruses expressing HA from clades 2.1 and 2.2 and 90% of the mice had titers.gtoreq.1:40 to a clade 2.3 representative virus (FIGS. 3E and 3F). Non-adjuvanted vaccines elicited generally lower HAI antibody titers with 100% of high dose animals achieving titers.gtoreq.1:40 only against clade 2.1 viruses. Imject.RTM. alum adjuvanted vaccines elicited significantly higher HAI antibody titers to clade 2.2 and clade 2.3 viruses regardless of dose as compared to non-adjuvanted vaccines (p<0.05 for high dose and p<0.001 for low dose, respectively). None of the vaccines elicited high HAI titer antibodies to a clade 1 virus.
Mouse Dosing Challenge
[0175] Mice that received the COBRA VLP vaccines or mock vaccinated control mice were challenged intranasally with a lethal dose of clade 2.2 H5N1 highly pathogenic avian influenza (A/Mongolia/whooper swan/244/2005) to evaluate the protective efficacy of the different COBRA vaccine formulations. All COBRA vaccinated mice, regardless of dose or the presence of adjuvant, were protected from weight loss and death following lethal challenge, while all mock vaccinated animals rapidly lost weight and required euthanasia by day 6 post infection (FIGS. 4A and 4B). Additionally, COBRA VLP vaccinated mice had no signs of disease, while mock vaccinated animals developed such symptoms as ruffled fur, hunched back, and lethargy (FIGS. 4C and 4D).
Mouse Comparison Immunizations
[0176] To determine if the COBRA HA vaccine elicits a broader antibody response compared to a vaccine derived from a primary isolate, an additional set of mice were vaccinated with either COBRA VLPs or clade 2.2 (A/Mongolia/whooper swan/244/2005) VLPs. Mice (BALB/c; n=20) were vaccinated (week 0 and 3) via intramuscular injection with either COBRA VLPs or clade 2.2 VLPs at a high dose (3 .mu.g HA) with Imject.RTM. alum adjuvant. At week 5, all COBRA VLP-vaccinated mice and all clade 2.2 VLP-vaccinated mice had anti-HA antibodies that recognized heterologous recombinant HA derived from both clade 1 and various sub-clades of clade 2 (FIG. 5A). Although no significant differences were found in total IgG titers between vaccine groups, COBRA VLP-vaccinated animals had higher HAI antibody titers against all viruses tested as compared to clade 2.2 VLP-vaccinated animals (p<0.01; FIG. 5B). Furthermore, COBRA VLP-vaccinated animals had an increased frequency of HAI titers of .gtoreq.1:40 compared to clade 2.2 VLP-vaccinated animals (Table 2).
TABLE-US-00003 TABLE 2 Mouse seroconversion frequency Vaccine Antigen Clade 1.sup.a Clade 2.1.sup.b Clade 2.2.sup.c Clade 2.3.sup.d COBRA 45% (9/20) 100% (20/20) 100% (20/20) 100% (20/20) Clade 2.2.sup.c 0% (0/20) 0% (0/20) 0% (0/20) 0% (0/20) Percentage of VLP-vaccinated animals achieving an HAI titer of .gtoreq.1:40 to each test antigen. .sup.aA/Vietnam/1203/2004 .sup.bA/Indonesia/5/2005 .sup.cA/Whooper Swan/Mongolia/244/2005 .sup.dA/Anhui/1/2005
Mouse Comparison Challenge
[0177] Mice that received the COBRA VLP vaccine, clade 2.2 VLP vaccine or mock vaccinated control mice were challenged intranasally with a lethal dose of clade 2.2 H5N1 highly pathogenic avian influenza (A/Mongolia/whooper swan/244/2005) to evaluate the protective efficacy of the VLP vaccines. All VLP-vaccinated mice were protected from weight loss and death following lethal challenge while all mock vaccinated animals rapidly lost weight and required euthanasia by day 6 post infection (FIG. 6A). Additionally, VLP vaccinated mice did not show signs of disease, while mock vaccinated animals developed ruffled fur, hunched back, and lethargy (FIG. 6B). Even though the clade 2.2 VLP was matched to the challenge virus, no significant differences were found between COBRA VLP and clade 2.2 VLP vaccinated mice in any of the parameters analyzed indicating that the COBRA VLP vaccine protected animals as efficiently as the homologous vaccine.
Ferret Comparison Immunizations
[0178] Ferrets are the most relevant model for influenza disease and as such the COBRA vaccine was tested in this more rigorous animal model. Ferrets (Fitch; n=9) were vaccinated (week 0 and 3) via intramuscular injection with COBRA VLPs or clade 2.2 VLPs at a high dose (15 .mu.g HA) with Imject.RTM. alum adjuvant. Serum was collected from ferrets at week 5 and antibody responses to the COBRA vaccines were evaluated. All vaccinated ferrets had anti-HA antibodies that recognized heterologous recombinant HA derived from both clade 1 and also sub-clades of clade 2 (FIG. 7A). No significant difference in anti-HA antibody was found between the COBRA VLP vaccine and the clade 2.2 VLP vaccine for any of the antigens tested (p>0.05). In addition to recognizing HA, antibodies were also evaluated for HAI activity. COBRA VLP-vaccinated animals had higher HAI antibody titers against clade 2.1 and clade 2.3 viruses as compared to clade 2.2 VLP-vaccinated animals (p<0.01 FIG. 7B). Similar to the mice, COBRA VLP-vaccinated ferrets displayed an increased rate of achieving HAI titers.gtoreq.1:40 when compared to clade 2.2 VLP-vaccinated ferrets (Table 3).
TABLE-US-00004 TABLE 3 Ferret seroconversion frequency Vaccine Antigen Clade 1.sup.a Clade 2.1.sup.b Clade 2.2.sup.c Clade 2.3.sup.d COBRA 0% (0/9) 78% (7/9) 56% (5/9) 56% (5/9) Clade 2.2.sup.c 0% (0/9) 0% (0/9) 22% (2/9) 0% (0/9) Percentage of VLP-vaccinated animals achieving an HAI titer of .gtoreq.1:40 to each test antigen. .sup.aA/Vietnam/1203/2004 .sup.bA/Indonesia/5/2005 .sup.cA/Whooper Swan/Mongolia/244/2005 .sup.dA/Anhui/1/2005
Ferret Comparison Challenge
[0179] Ferrets that received the COBRA VLP vaccines, clade 2.2 VLP vaccines or mock vaccinated control animals were challenged intranasally with clade 2.2 H5N1 highly pathogenic avian influenza (A/Mongolia/whooper swan/244/2005) to evaluate the protective efficacy of the COBRA vaccine in the ferret model of influenza infection. All VLP vaccinated ferrets were protected from weight loss and death following viral challenge, while all mock vaccinated animals rapidly lost weight and 78% (7/9) of mock vaccinated animals required euthanasia by day 7 post-infection (FIGS. 8A and 8B). Additionally, both COBRA VLP-vaccinated and clade 2.2-vaccinated ferrets were protected from acute fever and failed to develop significant signs of disease while mock vaccinated animals had an elevated body temperature and developed such symptoms as lethargy, diarrhea and decreased food and water intake (FIGS. 8C and 8D). In addition to monitoring outward signs of disease progression, nasal washes were collected for determination of viral replication in the upper respiratory tract. Ferrets vaccinated with either COBRA VLPs or clade 2.2 VLPs did not have detectable virus at any point after infection, while mock vaccinated animals had high levels of viral replication for the first five days of the infection (FIG. 8E). No significant differences were found between COBRA VLP and clade 2.2 VLP vaccinated ferrets in any of the challenge parameters analyzed confirming the findings in mice that the COBRA VLP vaccine protected animals as efficiently as the homologous vaccine.
[0180] The percent identity of COBRA HA and the test antigens used in the mouse and ferret studies described above are shown in Table 4.
TABLE-US-00005 TABLE 4 Percent Identity of Test Antigens Vaccine Antigen Clade 1.sup.a Clade 2.1.sup.b Clade 2.2.sup.c Clade 2.3.sup.d COBRA 97% 97% 95% 97% Clade 2.2.sup.c 94% 97% 100% 94% HA amino acid sequences were aligned and percent identity across the entire protein was determined for the vaccine immunogens compared to the representative test antigens. .sup.aA/Vietnam/1203/2004 .sup.bA/Indonesia/5/2005 .sup.cA/Whooper Swan/Mongolia/244/2005 .sup.dA/Anhui/1/2005
Example 2: A Computationally-Optimized HA VLP Vaccines Elicits Broadly-Reactive Antibodies that Protect Monkeys from H5N1 Infection
[0181] This example describes the finding that a COBRA clade 2 HA H5N1 VLP elicits broad humoral immunity against multiple H5N1 isolates from different clades.
Materials and Methods
Expression and Purification of Virus-Like Particles
[0182] The COBRA HA sequence is described above in Example 1. 293T cells were transiently transfected with plasmids expressing HA, M1, and NA in low serum media, incubated for 72 h at 37.degree. C., and purified by ultracentrifugation through a 20% glycerol cushion as previously described (Giles and Ross, Vaccine 29:3043-3054, 2011). All VLP vaccines were engineered using the same NA from A/Thailand/1(KAN-1)/2004. HA content was quantified as previously described (Giles and Ross, Vaccine 29:3043-3054, 2011). Two different VLP preparations were purified, each containing one of the HA influenza gene products: WS/05 or the COBRA HA.
Primate Immunizations and H5N1 Challenges
[0183] Cynomolgus macaques (Macaca fascicularis, male, 3-5 years old) were vaccinated with 15 .mu.g (based upon HA content) of purified COBRA HA VLPs (n=7) or WS/05 VLPs (n=7), via intramuscular injection at weeks 0, 3 and 6. Vaccines at each dose were formulated with alum adjuvant (Imject.RTM. Alum, Pierce Biotechnology; Rockford, Ill., USA) immediately prior to use. Twenty-one days after each vaccination, blood was collected from anesthetized macaques. All procedures were in accordance with the NRC Guide for the Care and Use of Laboratory Animals.
[0184] Three weeks after final vaccination, macaques were placed into BSL3+ isolator units (Bioqual, Inc., Rockville, Md.) and then challenged by a multi-route of infection (ocular, nasal, tracheal) as previously described (Kobasa et al., Nature 445:319-323, 2007; Kuiken et al., Vet Pathol 40:304-310, 2003; Rimmelzwaan et al., Avian Dis 47:931-933, 2003) using 1.times.10.sup.6 plaque forming units (PFU) of the highly pathogenic H5N1 virus, A/Whooper Swan/Mongolia/244/2005 (clade 2.2), at each location. Monkeys were monitored daily for weight loss, signs of disease, and mortality until 7 days after infection. Individual body weights, sickness scores (based upon lethargy, temperature change, nasal discharge, lack of appetite, dehydration, lack of responsiveness), and death were recorded for each group.
[0185] Nasal and tracheal washes were performed at day 0, 1, 3, 5, and 7 post-infection. In addition, subsets of monkeys were sacrificed following administration of anesthesia and necropsies were performed according to standard procedures for assessment of gross pathologic and histopathologic changes, as well as the extent of virus replication.
Serological Assays
[0186] A quantitative ELISA was performed to assess anti-HA specific IgG in immune serum as previously described (Bright et al., PLoS One 3:e1501, 2008; Giles and Ross, Vaccine 29:3043-3054, 2011). The hemagglutination inhibition (HAI) assay was used on sera treated with receptor destroying enzyme (RDE; Denka Seiken, Co., Japan) prior to being tested (Bright et al., Vaccine 25:3871-3878, 2007; Mitchell et al., Vaccine 21:902-914, 2003; Bright et al., PLoS One 3:e1501, 2008) to assess functional antibodies to the HA able to inhibit agglutination of horse red blood (Askonas B, McMichael A, Webster R. The immune response to influenza viruses and the problem of protection against infection. In: Beare AS, editor. Basic and applied influenza research: CRC Press 1982: 159-188). The protocol was adapted from the CDC laboratory-based influenza surveillance manual and performed as previously described (Gillim-Ross and Subbarao, Clin Microbiol Rev 19:614-636, 2006; Bright et al., PLoS One 3:e1501, 2008). The HAI titer was determined by the reciprocal dilution of the last row which contained non-agglutinated RBC. Positive and negative serum controls were included for each plate. All monkeys were negative (HAI.ltoreq.1:10) for pre-existing antibodies to currently circulating human influenza viruses prior to vaccination. Serum neutralizing antibody titers were determined by microneutralization (MN) assays performed on Madin Darby Canine Kidney (MDCK) cells following the procedure until CPE was observed (Rowe et al., J Clin Microbiol 37:937-943, 1999). Cells were then fixed in 10% formalin and stained with 1% crystal violet to quantify CPE. The neutralizing antibody titers are expressed as the reciprocal of the highest dilution of serum that gave 50% neutralization of 100 TCID.sub.50 of virus in MDCK cells. Geometric mean neutralizing antibody titers were calculated for each group.
Histopathologic Evaluation and Immunohistochemical Analysis
[0187] Formalin-inflated lungs and trachea were fixed in 10% neutral buffered formalin. Cross-sections of upper and lower left and right lung lobes and trachea were made, concentrating on gross-lesions. Tissue was paraffin-embedded and 6 .mu.m sections were stained with hematoxylin and eosin for histologic evaluation. Sequential sections were processed for immunohistochemistry or in situ hybridization (ISH). Immunohistochemistry was performed as described previously (Bissel et al., Am J Pathol 160:927-941, 2002) using an immunoperoxidase method with a polyclonal antibody (Maine Biotechnology Services, Portland, Me.) directed against influenza A virus. A biotinylated donkey anti-goat IgG (Rockland Immunochemicals, Gilbertsville, Pa.) was used as the secondary antibody. ISH was performed as described previously (Fallert et al., J Virol Methods 99:23-32, 2002) using a 35S-labeled riboprobe synthesized using templates derived from 760 bp of influenza A/California/04/2009 matrix protein.
Results
Vaccine Induced Antibody Responses
[0188] Cynomolgus macaques were vaccinated with COBRA VLPs or WS/05 VLPs formulated with Imject.RTM. alum at 0, 3 and 6 weeks. At week 3 post-vaccination, all COBRA VLP-vaccinated animals had anti-HA antibodies that recognized recombinant HA derived from three sub-clades of clade 2, which were boosted at week 6 (FIGS. 10A and 10B). There was no statistical difference (p>0.05) in the anti-HA titers elicited against any of the HA proteins, except monkeys vaccinated with COBRA VLPs had a statistically higher titer against the Indo/05 HA (clade 2.1) compared with monkeys vaccinated with the WS/05 VLP (derived from clade 2.2) on week 6.
A Single COBRA VLP Vaccination Induced High Titer HAI and MN Antibodies to Clade 2 H5N1 Viruses
[0189] Monkeys vaccinated with COBRA VLPs (but not with WS/05 VLPs) had HAI activity against viruses representing all three clade 2 sub-clades after a single vaccination (FIG. 10C). Four to six monkeys responded to the COBRA VLP vaccine with an HAI titer.gtoreq.1:40 for the all of the various test antigens. In contrast, 4 of 7 monkeys vaccinated with the WS/05 VLP responded to the homologous clade 2.2 virus, but none of these vaccinated monkeys responded to the clade 2.1 or 2.3 virus. Following a second vaccination, almost all the monkeys vaccinated with either vaccine responded to all three viruses (FIG. 11D). These results were confirmed by microneutralization assay (FIGS. 11E and 11F). However, monkeys vaccinated with COBRA VLPs showed boosted HAI titers to all three clade 2 viruses (FIG. 11).
COBRA VLPs Induced HAI Antibodies that Recognize Broader Numbers of H5N1 Viruses
[0190] In order to determine if the COBRA HA elicited antibodies that recognized a broader number of H5N1 isolates, serum was collected and tested for the ability to inhibit influenza virus induced hemagglutination of red blood cells in vitro. Antisera collected from both vaccinated and unvaccinated monkeys were then tested against a broad panel of H5N1 viruses representing not only sub-clades of clade 2, but also non-clade 2 H5N1 virus strains (0, 1, 4, and 7) by HAI. Monkeys vaccinated with the COBRA VLP had high average HAI titers against all clade 2 isolates, regardless of sub-clade (FIG. 11). In general, all 7 monkeys responded to the COBRA VLP vaccine and seroconverted with an HAI titer.gtoreq.1:40 against all the clade 2 viruses. In contrast, monkeys vaccinated with the WS/05 VLP vaccine had lower HAI titers against clade 2 viruses (FIG. 10) and fewer number of monkeys responded to the vaccine. Of the 10 clade 2 viruses tested in the HAI assay, WS/05 VLP vaccinated monkeys responded more poorly (fewer than 4 of 7 monkeys) to 4 of the isolates and none of these monkeys had antibodies that responded to the Dk/HU/02 (clade 2.1.1) or Eg/3300/08 (clade 2.2.1) isolates. The COBRA VLPs elicited significantly higher HAI titers against almost all of the clade 2 viruses than the WS/05 VLPs (FIG. 11).
[0191] In addition to clade 2 isolates, a minimum of five COBRA VLP vaccinated monkeys had HAI antibodies against both clade 1 and 7 virus isolates (FIG. 11). In comparison, almost none of the WS/05 VLP vaccinated monkeys had HAI antibodies against clade 1 and clade 7 viruses. None of the monkeys, regardless of the vaccine, had antibodies that responded to the clade 0 or 4 isolates. All mock vaccinated monkeys did not recognize any of the H5N1 isolates.
Challenge of Vaccinated and Unvaccinated Primates with H5N1 Clade 2.2 Virus
[0192] Three weeks after final vaccination, both VLP vaccinated and mock-vaccinated monkeys were transferred to ABSL3+ isolator units and then challenged with highly pathogenic H5N1 virus, A/Whooper Swan/Mongolia/244/2005 (clade 2.2) (1.times.10.sup.6 pfu), by a multi-route (ocular, nasal, tracheal, oral) of infection (Kobasa et al., Nature 445:319-323, 2007; Kuiken et al., Vet Pathol 40:304-310, 2003; Rimmelzwaan et al., Avian Dis 47:931-933, 2003). There was no significant weight loss or mortality in any of the monkeys over the 7 day period of observation. Unvaccinated monkeys had an elevated temperature of .about.2.degree. C. that was sustained for 5 days post-infection and higher gross pathology scores by day 3 post-infection (Table 5).
TABLE-US-00006 TABLE 5 Lung pathology, temperature and viral titer of vaccinated macaques Lung Pathology Elevated Peak Viral Titer (pfu/ml) Vaccine Score (day 3) temperature (days) (day) Mock 5.3 1.9.degree. C. (1-5 DPI) Nasal wash: 2.2-2.5 (5 DPI) Trachea wash: 2.0-4.4 (3 DPI) WS/05 VLP 3.3 1.1.degree. C.-1.3.degree. C. (1-5 DPI) Nasal wash: <2 Trachea wash: <2 COBRA 2.1 1.3.degree. C. (2 DPI) Nasal wash: <2 VLP Trachea wash: <2
[0193] The lungs of unvaccinated monkeys had mild to moderate acute pneumonia with alveolar pulmonary exudate by day 3 post-infection by H&E staining. ISH showed focal collections of H5N1 infected cells present at day 3 post-infection in alveolar spaces and to a lesser extent in bronchial epithelium. These results were similar to unvaccinated monkeys infected with the clade 1 H5N1 virus, A/Vietnam/1203/2004. In contrast, monkeys vaccinated with either the COBRA VLP or the WS/05 VLP vaccine had a reduced gross pathology scores of 2.1-3.3 at day 3 post-infection with a milder increase in body temperature (1.1-1.3.degree. C.) that spiked between days 2-3 post-infection and then returned to pre-infection temperatures. Vaccinated animals had fewer H5N1 infected cells that were detected primarily on day 1 post-infection (Table 6).
TABLE-US-00007 TABLE 6 H5N1 lung infection scores Alveolar infection Submucosal Vaccine score infection score 1 3 5 1 3 5 1 3 5 Mock 1.00 0.05 0 1.10 0.48 0.25 0 0 0 WS/05 VLP 0.05 0 0 0.55 0.10 0 0 0 0 COBRA 0 0 0 0.60 0.03 0.05 0 0 0 VLP
ISH for influenza was performed on tissue sections of from upper and lower left and right lung. A semi-quantitative scoring system was developed to evaluate the presence of influenza infected cells. Scores were then averaged: 0.2=rare or occasional cells but<5% of fields; 1=>1/2 to 1/4 low power fields; 2=>1/4 low power fields; 3=essentially all low power fields.
[0194] However, monkeys vaccinated with the COBRA VLP had little to no signs of lung inflammation by H&E staining, while animals vaccinated with the WS/05 VLP vaccine had similar signs of inflammation as non-vaccinated monkeys (Table 7). In addition, unvaccinated monkeys had high titers of virus in both the nasal and tracheal washes between days 3 and 5 post-infection. In contrast, no virus was detected in either vaccinated groups.
TABLE-US-00008 TABLE 7 Lung involvement and inflammation scores % lung Bronchial Alveolar involvement.sup.a inflammation.sup.b inflammation.sup.b Vaccine 1 3 5 1 3 5 1 3 5 Mock 0.38 1.13 1.25 0.63 0.75 1.25 0.63 1.00 1.25 (0-1) (0-2) (0-2) (0-1) (0-2) (0-2) (0-1) (0-2) (0-2) WS/05 VLP 0.75 1.50 0.88 1.00 1.42 0.63 1.00 1.25 1.00 (0-2) (0-3) (0-3) (1) (1-2) (0-2) (0-2) (0-2) (0-2) COBRA VLP 0.88 0.50 0.38 1.13 0.75 0.88 1.13 0.67 0.25 (0-2) (0-2) (0-2) (1-2) (0-2) (0-2) (0-2) (0-2) (0-1) .sup.a% Lung involvement. Tissue sections from upper and lower left and right lung were evaluated for percent area demonstrating pneumonia. Scores were then averaged. Range in parentheses. 0 = <10%, 1 = 10-24%, 2 = 25-50%, 3 = >50%. .sup.bBronchial and alveolar inflammation scores. Tissue sections from upper and lower left and right lung were evaluated for presence of bronchial inflammation and denudation and alveolar immune cell infiltration. Scores were then averages: 0 = absent, 1 = present, 2 = abundant.
Example 3: Comparison of Protective Efficacy by Vaccination with Computationally Optimized HA and Polyvalent HA Based H5N1 VLP Vaccines
[0195] This example describes a comparison of the COBRA HA vaccine to a polyvalent H5N1 vaccine. The results demonstrate that a single COBRA antigen elicits broader antibodies and is more effective than a polyvalent mixture of primary antigens.
Materials and Methods
Vaccine Antigens and Preparation
[0196] The design and characterization of the computationally optimized broadly reactive antigen (COBRA) is described in Example 1. Polyvalent vaccine HA antigens were derived via reverse transcription from the following 6:2 reassortant H5N1 viruses: A/Indonesia/5/2005 (clade 2.1; IN/05), A/Whooper Swan/Mongolia/244/2005 (clade 2.2; WS/05) and A/Anhui/1/2005 (clade 2.3; AN/05). All HA antigens were cloned into the pTR600 expression vector.
[0197] Virus-like particles (VLPs) were generated by transiently transfecting HEK 293T cells with plasmids expressing M1 (A/Puerto Rico/8/1934), NA (A/Thailand/1(KAN-1)/2004), and a single HA for each preparation. Cells were incubated for 72 h at 37.degree. C. after which supernatants were harvested. Cell debris was cleared by low speed centrifugation followed by vacuum filtration through a 0.22 .mu.m sterile filter. VLPs were purified by ultracentrifugation (100,000.times.g through 20% glycerol, weight to volume) for 4 hours at 4.degree. C. Pellets were then resuspended in PBS pH 7.2 and stored in single use aliquots at -80.degree. C. until use. Total protein concentration was determined by MicroBCA.TM. Protein Assay Reagent Kit (Pierce Biotechnology, Rockford, Ill., USA). HA specific content of each VLP was determined by scanning densitometry as described previously (Giles and Ross, Vaccine 29:3043-3054, 2011). Briefly, purified HA matched to each VLP was electrophoresed with purified VLPs, transferred to a PVDF membrane and probed by western blot with H5-specific antisera. The relative density of the HA band in the purified protein lanes was used to calculate a standard curve and the density of the HA in the VLP lanes was interpolated. In total, four different VLP preparations were purified and HA content quantified independently, each containing one of the three wild-type influenza gene products (IN/05, WS/05, AN/05) or the COBRA HA.
Mouse Studies
[0198] BALB/c mice (Mus musculis, females, 6-8 weeks) were purchased from Harlan Sprague Dawley, (Indianapolis, Ind., USA) and housed in microisolator units and allowed free access to food and water and were cared for under USDA guidelines for laboratory animals. Mice were vaccinated with purified COBRA VLPs (3 .mu.g HA) or a polyvalent formulation of VLPs consisting of 1 .mu.g HA each IN/05, WS/05 and AN/05 (3 .mu.g HA total) via intramuscular injection at week 0 and then boosted at week 3. Vaccines were formulated with Imject.RTM. alum adjuvant (Imject.RTM. Alum, Pierce Biotechnology; Rockford, Ill., USA) according to the manufacturer's protocol. Fourteen to twenty-one days after each vaccination, blood was collected from anesthetized mice via the retro-orbital plexus and transferred to a microfuge tube. Tubes were centrifuged and sera was removed and frozen at -20.+-.5.degree. C.
[0199] Three weeks after final vaccination, mice were challenged intranasally with 5.times.10.sup.3 plaque forming units (PFU) of either highly pathogenic wild type H5N1 virus A/Whooper Swan/Mongolia/244/2005 (n=20/group) or 6:2 reassortant virus with internal genes from the mouse adapted virus A/Puerto Rico/8/1934 and the surface proteins HA and NA from A/Vietnam/1203/2004 (n=10/group) in a total volume of 50 .mu.l. Challenge doses for both viruses were established independently and represent approximately 50LD.sub.50. After infection, mice were monitored daily for weight loss, disease signs and death for 14 days after infection. Individual body weights, sickness scores and death were recorded for each group on each day after inoculation. Sickness score was determined by evaluating activity (0=normal, 1=reduced, 2=severely reduced), hunched back (0=absent, 1=present) and ruffled fur (0=absent, 1=present) (Toapanta and Ross, Respiratory Res 10(1):112, 2009). Experimental endpoint was determined by >20% weight loss or display of neurological disease such as hind limb paralysis. All highly pathogenic wild type H5N1 influenza virus studies were performed under high-containment biosafety level 3 enhanced conditions (BSL3+).
Ferret Studies
[0200] Fitch ferrets (Mustela putorius faro, female, 6-12-months of age), influenza naive and descented, were purchased from Marshall Farms (Sayre, Pa., USA). Ferrets were pair housed in stainless steel cages (Shor-line, Kansas City, Kans., USA) containing Sani-chips Laboratory Animal Bedding (P.J. Murphy Forest Products, Montville, N.J., USA). Ferrets were provided with Teklad Global Ferret Diet (Harlan Teklad, Madison, Wis., USA) and fresh water ad libitum. The VLPs were diluted in PBS, pH 7.2 to achieve final concentration. Ferrets (n=6) were vaccinated with purified COBRA VLPs (15 .mu.g HA) or a polyvalent formulation of VLPs consisting of 5 .mu.g HA each IN/05, WS/05 and AN/05 (15 .mu.g HA total) via intramuscular injection at week 0 and then boosted at week 3. Vaccines were formulated with Imject.RTM. alum adjuvant (Imject.RTM. Alum, Pierce Biotechnology; Rockford, Ill., USA) immediately prior to use according to the manufacturer's protocol. Animals were monitored for adverse events including weight loss, temperature, loss of activity, nasal discharge, sneezing and diarrhea weekly during the vaccination regimen. Prior to vaccination, animals were confirmed by HAI assay to be seronegative for circulating influenza A (H1N1 and H3N2) and influenza B viruses. Fourteen to twenty-one days after each vaccination, blood was collected from anesthetized ferrets via the anterior vena cava and transferred to a microfuge tube. Tubes were centrifuged and sera was removed and frozen at -20.+-.5.degree. C.
[0201] Three weeks after final vaccination, ferrets were challenged intranasally with 1.times.10.sup.6 plaque forming units (PFU) of the highly pathogenic H5N1 virus A/Whooper Swan/Mongolia/244/2005 (clade 2.2) in a volume of 0.5 ml in each nostril for a total infection volume of 1 ml. After infection, ferrets were monitored daily for weight loss, disease signs and death for 14 days after infection. Individual body weights, sickness scores, and death were recorded for each group on each day after inoculation. Sickness score was determined by evaluating activity (0=normal, 1=alert and active after stimulation, 2=alert but not active after stimulation, 3=neither active nor alert after stimulation), nasal discharge (0=absent, 1=present), sneezing (0=absent, 1=present), decreased food intake (0=absent, 1=present), diarrhea (0=absent, 1=present), dyspnea (0=absent, 1=present) and neurological symptoms (0=absent, 1=present) as previously described (Giles and Ross, Vaccine 29:3043-3054, 2011). Experimental endpoint was defined as >20% weight loss, development of neurological disease or an activity score of 3 (neither active nor alert after stimulation). Nasal washes were performed by instilling 3 ml of PBS into the nares of anesthetized ferrets each day for 14 days after inoculation. Washes were collected and stored at -80.degree. C. until use. All highly pathogenic wild type H5N1 influenza virus studies were performed under high-containment biosafety level 3 enhanced conditions (BSL3+).
ELISA Assay
[0202] The ELISA assay was used to assess total antibody titer to the HA. High binding, 96-well polystyrene plates (Costar; Lowell, Mass., USA) were coated overnight with 50 ng/well of recombinant HA. Coating antigens were derived from the following representative viral isolates: A/Vietnam/1203/2004 (clade 1), A/Indonesia/5/2005 (clade 2.1), A/Whooper Swan/Mongolia/244/2005 (clade 2.2) and A/Anhui/1/2005 (clade 2.3). Plates were blocked with 5% milk diluted in PBS with 0.05% Tween 20. Serum samples were diluted in blocking buffer and added to plates. Serum was two-fold serially diluted and allowed to incubate for 1 hour at room temperature. Plates were washed and species specific antibody against IgG linked to horseradish peroxidase (HRP) was diluted in blocking buffer and added to plates. Plates were incubated for 1 hour at room temperature. Plates were washed and HRP was developed with TMB substrate (Sigma-Aldrich; St. Louis, Mo., USA). Plates were incubated in the dark for 15 minutes and then the reaction was stopped with 2N H.sub.2SO.sub.4. Optical densities at a wavelength of 450 nm (OD.sub.450) were read by a spectrophotometer (BioTek; Winooski, Vt., USA) and end point dilution titers were determined as the reciprocal dilution of the last well which had an OD.sub.450 above the mean OD.sub.450 plus two standard deviations of naive animal sera.
Hemagglutination Inhibition (HAI) Assay
[0203] The HAI assay was used to assess functional antibodies to the HA able to inhibit agglutination of horse erythrocytes. The protocol was adapted from the CDC laboratory-based influenza surveillance manual (Gillim-Ross and Subbarao, Clin Microbiol Rev 19(4):614-636, 2006). To inactivate non-specific inhibitors, sera were treated with receptor destroying enzyme (RDE; Denka Seiken, Co., Japan) prior to being tested. Briefly, three parts RDE was added to one part sera and incubated overnight at 37.degree. C. RDE was inactivated by incubation at 56.degree. C. for .about.30 min. RDE treated sera was two-fold serially diluted in v-bottom microtiter plates. An equal volume of reassortant virus, adjusted to approximately 8 HAU/50 .mu.l, was added to each well. The reassortant viruses contained the internal genes from the mouse adapted strain A/Puerto Rico/8/1934 and the surface proteins HA and NA from the following representative viral isolates: A/Vietnam/1203/2004 (clade 1), A/Indonesia/5/2005 (clade 2.1), A/Whooper Swan/Mongolia/244/2005 (clade 2.2) and A/Anhui/1/2005 (clade 2.3). The plates were covered and incubated at room temperature for 20 minutes followed by the addition of 1% horse erythrocytes (HRBC) (Lampire Biologicals, Pipersville, Pa., USA) in PBS. Red blood cells were stored at 4.degree. C. and used within 72 h of preparation. The plates were mixed by agitation, covered, and the RBCs were allowed to settle for 1 h at room temperature (Askonas B, McMichael A, Webster R. The immune response to influenza viruses and the problem of protection against infection. In: Beare AS, editor. Basic and applied influenza research: CRC Press 1982: 159-188). The HAI titer was determined by the reciprocal dilution of the last well which contained non-agglutinated RBC. Positive and negative serum controls were included for each plate. All mice and ferrets were negative (HAI.ltoreq.1:10) for pre-existing antibodies to currently circulating human influenza viruses prior to vaccination.
Plaque Assay
[0204] For mouse infections, lung virus titers were evaluated. For ferret infections, nasal wash virus titers were used to assess viral burden. Both lungs and nasal wash virus titers were determined using a plaque assay (Tobita et al., Med Microbiol Immunol 162:23-27, 1975; Tobita et al., Med Microbiol Immunol 162:9-14, 1975). Briefly, lungs from mice infected with virus were harvested post infection, snap-frozen and stored at -80.degree. C. until use. Samples were thawed, weighed and single cell suspensions were prepared via passage through a 70 .mu.m mesh (BD Falcon, Bedford, Mass., USA) in an appropriate volume of DMEM supplemented with penicillin-streptomycin (iDEME) as to achieve 100 mg/ml final concentration. Cell suspensions were centrifuged at 2000 rpm for 5 minutes and the supernatants were collected.
[0205] Madin-Darby Canine Kidney (MDCK) cells were plated (5.times.10.sup.5) in each well of a 6 well plate. Samples (lung supernatants for mice and nasal washes for ferrets) were diluted (dilution factors of 1.times.10.sup.1 to 10.sup.6) and overlayed onto the cells in 100 .mu.l of iDMEM and incubated for 1 hour. Virus-containing medium was removed and replaced with 2 ml of L15 medium plus 0.8% agarose (Cambrex, East Rutherford, N.J., USA) and incubated for 96 hours at 37.degree. C. with 5% CO.sub.2. Agarose was removed and discarded. Cells were fixed with 10% buffered formalin, and then stained with 1% crystal violet for 15 minutes. Following thorough washing in dH.sub.2O to remove excess crystal violet, plates were allowed to dry, plaques counted, and the plaque forming units (PFU)/g for or PFU/ml for nasal washes were calculated.
Histopathological Analysis
[0206] Left lobes of lungs from infected mice were collected 4 days post-infection and placed into 10% buffered formalin. After fixation, lungs were paraffin embedded and 6 .mu.m sections were prepared for histopathological analysis. For in situ hybridization (ISH), vectors containing 760 bp of Influenza/California/04/2009 matrix protein were linearized to create antisense and sense templates. .sup.35S-labeled riboprobes were generated using MAXIscript in vitro transcription kit (Ambion, Austin, Tex.). ISH was performed as described before (Bissel et al., Brain Pathol, Accepted Article doi: 10.1111/j.1750-3639.2010.00514.x). Control riboprobes did not hybridize to lung tissue at any time point post-infection and non-infected tissue did not show hybridization with viral probes. Hybridized slides were assessed and scored for abundance of foci.
Cellular Assays
[0207] The number of anti-influenza specific cells secreting interferon gamma (IFN-.gamma.) was determined by enzyme-linked immunospot (ELISpot) assay (R&D systems, Minneapolis, Minn., USA) following the manufacturer's protocol. Mice were sacrificed at 6 days post infection (DPI) and spleens and lungs were harvested and prepared in single cell suspensions. Briefly, pre-coated anti-IFN.gamma. plates were blocked with RPMI plus 10% FCS and antibiotics (cRPMI) for 30 minutes at room temperature. Media was removed from wells and 10.sup.5 cells were added to each well. Cells were stimulated with purified recombinant HA from A/Vietnam/1203/2004 (truncated at residue 530; 1 .mu.g/well), inactivated 6:2 reassortant virus A/Vietnam/1203/2004 (1:100 dilution of inactivated stock; 100 .mu.l/well) or the immunodominant H2-K.sup.d CD8.sup.+ T cell epitope in H5 HA: HA533 (IYSTVASSL; SEQ ID NO: 10; 1 .mu.g/well) (Pepscan Presto, Leystad, Netherlands). Additional wells were stimulated with PMA (50 ng/well) and ionomycin (500 ng/well) as positive controls or Ova.sub.257 (SIINFEKL; SEQ ID NO: 11; 1 .mu.g/well) (Pepscan Presto, Leystad, Netherlands) as negative controls. Additionally, IL-2 (10 U/ml) was added to each well. Plates were incubated at 37.degree. C. for 48 hours. After incubation, plates were washed four times with R&D wash buffer and were incubated at 4.degree. C. overnight with biotinylated anti-mouse IFN.gamma.. Plates were washed as before and incubated at room temperature for 2 hours with streptavidin conjugated to alkaline phosphatase. Plates were washed as before and spots were developed by incubating at room temperature for 1 hour in the dark with BCIP/NBT chromogen substrate. The plates were washed extensively with DI H.sub.2O and allowed to dry overnight prior to spots being counted using an ImmunoSpot ELISpot reader (Cellular Technology Ltd., Cleveland, Ohio, USA).
[0208] The number of anti-HA and anti-NA specific antibody secreting cells was determined by B cell ELISpot assay as previously described (Joo et al., Vaccine 28:2186-2194, 2009; Sasaki et al., PLoS ONE 3:e2975, 2008; Sasaki et al., J Virol 81:215-228, 2007). Mice were sacrificed at 6 DPI and spleens and lungs were harvested and prepared in single cell suspensions. Briefly, 0.45 .mu.m PVDF membrane plates (Millipore, Billerica, Mass., USA) were coated with either purified recombinant HA from A/Vietnam/1203/2004 or purified recombinant NA from A/Thailand/1(KAN-1)/2004 (250 ng/well) and incubated at 4.degree. C. overnight. Plates were washed three times with PBS and blocked with cRPMI for at 37.degree. C. for 3-4 hours. Media was removed from wells and 10.sup.5 cells were added to each well. Plates were incubated at 37.degree. C. for 48 hours. After incubation, plates were washed as before and incubated at room temperature for 2 hours with horse radish peroxidase conjugated anti-mouse IgG or IgA (Southern Biotech, Birmingham, Ala., USA). Plates were washed as before and spots were developed at room temperature for 1 hour in the dark with detection substrate (NovaRED.TM.; Vector Labs, Burlingame, Calif., USA). The plates were washed extensively with DI H.sub.2O and allowed to dry overnight prior to spots being counted using an ImmunoSpot ELISpot reader (Cellular Technology Ltd., Cleveland, Ohio, USA).
Passive Transfer of Sera
[0209] Serum from vaccinated mice was pooled and passively transferred into 9 week old recipient BALB/c mice (n=5/group). Equal amounts of serum from each mouse in a particular vaccine group were pooled and heat inactivated for 30 minutes at 56.degree. C. 200 .mu.l of pooled and inactivated serum was transferred to recipient mice via IP injection. 24 hours post transfer, mice were infected with 6:2 reassortant virus with internal genes from the mouse adapted virus A/Puerto Rico/8/1934 and surface antigens from A/Vietnam/1203/2004.
Statistical Analysis
[0210] Statistical significance of the antibody and cellular immunology data was determined using a two-tailed Student's T test to analyze differences between COBRA and polyvalent vaccine groups for each of the different test antigens. Differences in weight loss and sickness score were analyzed by two-way ANOVA, followed by Bonferroni's post test for each vaccine group at multiple time points (multiparametric). Statistical significance of viral titer data was evaluated using a two-tailed Student's T test on Log.sub.10 transformed values. Significance was defined asp<0.05. Statistical analyses were done using GraphPad Prism software.
Results
Immunogenicity in Mice and Ferrets
[0211] BALB/c mice were vaccinated twice via intramuscular injection with either purified COBRA or polyvalent VLPs and two weeks after the second vaccination serum was analyzed for antibody responses. All vaccinated mice had high titer anti-HA antibodies that bound to recombinant HA derived from both clade 1 and various sub-clades of clade 2 (FIG. 12A). Although both COBRA and polyvalent vaccines elicited similar total IgG, COBRA vaccinated animals had higher HAI antibody titers for all viruses tested (p<0.001; FIG. 12B). In addition to higher HAI titer, COBRA vaccinated mice had an increased frequency of HAI titers.gtoreq.1:40 for all viruses tested, including those which were components of the polyvalent formulation (Table 8).
[0212] To confirm the results from mice in a more rigorous animal model, ferrets were vaccinated twice via intramuscular injection with either COBRA or polyvalent vaccines. Serum was collected two weeks after the second vaccination and antibody responses were evaluated. Similar to the mice, all vaccinated ferrets had anti-HA antibodies that bound to diverse recombinant HA and the relative total IgG titers were equivalent for both COBRA and polyvalent vaccines (FIG. 12C). COBRA vaccinated ferrets demonstrated increased HAI antibody titers compared to polyvalent vaccinated animals against all viruses tested, however only the antibodies to the clade 2.1 virus were significantly different (p<0.05; FIG. 12D). Furthermore, COBRA vaccinated animals displayed an increased rate of achieving an HAI titer of .gtoreq.1:40 in comparison to the polyvalent vaccinated ferrets for all test antigens (Table 8).
TABLE-US-00009 TABLE 8 Seroconversion frequency Species Vaccine antigen Clade 1 Clade 2.1 Clade 2.2 Clade 2.3 Mouse COBRA 60% (18/30) 100% (30/30) 100% (30/30) 100% (30/30) Polyvalent 3.3% (1/30) 70% (21/30) 50% (15/30) 53% (16/30) Ferret COBRA 33% (2/6) 67% (4/6) 50% (3/6) 50% (3/6) Polyvalent 0% (0/6) 33% (2/6) 0% (0/6) 0% (0/6)
Wild Type Clade 2.2 Challenge
[0213] To confirm protective efficacy against highly pathogenic H5N1 infection, vaccinated animals were challenged with a lethal dose of the wild-type clade 2.2 isolate A/Whooper Swan/Mongolia/244/2005. All VLP vaccinated mice were protected from weight loss and death while mock vaccinated animals rapidly lost weight and reached experimental end-point by 6 days post infection (DPI; FIG. 13A). COBRA and polyvalent vaccinated mice both had a mean maximum weight loss of 4% at 12 and 13 DPI, respectively. Additionally, all VLP vaccinated mice failed to develop any overt signs of disease while mock vaccinated mice developed visible illness (FIG. 13B).
[0214] Similar to the mice, all VLP vaccinated ferrets were protected from death following a lethal challenge. Vaccinated ferrets demonstrated mild weight loss in response to the infection with COBRA vaccinated animals having mean maximum weight loss of 5.5% at 2 DPI and polyvalent vaccinated animals losing 6.8% at 3 DPI (FIG. 13C). Both groups rapidly recovered weight and failed to develop any significant signs of disease (FIG. 13D). Furthermore, VLP vaccinated animals did not demonstrate any temperature spikes while mock vaccinated animals had an elevated temperature of -3.degree. C. for 1-3 DPI.
[0215] To evaluate vaccine efficacy with a more sensitive output than morbidity and mortality, the viral burden of infected animals was also determined. Both COBRA and polyvalent vaccinated mice had reduced lung viral titers as quickly as 1 DPI when compared to mock vaccinated animals. Furthermore, COBRA vaccinated mice did not have detectable virus by 3 DPI while polyvalent vaccinated mice demonstrated prolonged viral replication with 1.8.times.10.sup.3 PFU/g at 3 DPI (p<0.05; FIG. 14A). Additionally, both VLP vaccines prevented extra-pulmonary spread of the virus while mock vaccinated animals had detectable virus in both kidney and liver by 3 DPI. Control of virus replication in ferrets was similar to that observed in mice, although complete clearance of the virus was delayed (FIG. 14B). All VLP vaccinated animals had decreased recovery of virus in nasal washes compared to mock vaccinated ferrets at all timepoints tested (p<0.05). COBRA vaccinated animals did not have detectable virus by 5 DPI. In contrast, virus replication did not reach undetectable levels until 9 DPI in polyvalent vaccinated ferrets.
Histopathology of Infected Lungs
[0216] To evaluate the location and severity of influenza viral antigen and viral replication, ISH for influenza A MP was scored on 3 DPI lung sections. COBRA vaccinated animals had rare bronchial epithelium infection (FIGS. 15A and 15B). Animals receiving polyvalent vaccines had occasional bronchial epithelium infection that was comparable to the COBRA vaccinated animals (FIGS. 15A and 15B). This was in contrast to significant bronchial epithelium infection and replication observed in mock animals (FIGS. 15A and 15B).
Reassortant Clade 1 Challenge
[0217] Having established the clade 2.2 protective profile of both the COBRA and polyvalent vaccines, the efficacy of these vaccines against a more divergent clade 1 challenge in mice was evaluated. COBRA and polyvalent vaccinated mice were challenged with 6:2 reassortant virus containing the HA and NA proteins from the clade 1 virus A/Vietnam/1203/2004. All VLP vaccinated animals were protected from weight loss and death while mock vaccinated animals rapidly lost weight and reached experimental endpoint by 7 DPI (FIG. 16A). Furthermore, vaccinated mice also did not develop any signs of disease throughout the course of the study (FIG. 16B). Lungs were harvested at 3 DPI for determination of viral burden (FIG. 16C). COBRA vaccinated animals did not have detectable virus while polyvalent animals had 1.1.times.10.sup.3 PFU/g virus (p=0.12). Importantly, both vaccines had significantly less recoverable virus than mock vaccinated animals at 3 DPI (p<0.01).
Post-Challenge Cellular Immune Responses
[0218] The magnitude of influenza specific cellular immune responses in the lungs post-infection was evaluated via ELISpot assay for both antibody secreting cells (ASC) and IFN-.gamma. producing cells. Vaccinated mice were infected with reassortant A/Vietnam/1203/2994 virus as before and lungs were harvested at 6 DPI. COBRA and polyvalent vaccinated animals had statistically equivalent numbers of both IgG and IgA ASC specific for HA from the challenge virus (p>0.05; FIG. 17A). No ASC were detected in mock vaccinated animals indicating that the 6 DPI time point is likely representative of a recall response. Additionally, the majority of the ASC response to infection was specific for HA as lower numbers of cells were detected for the NA component of the vaccines.
[0219] VLP vaccine primed IFN-.gamma. secreting cells were also evaluated after infection. IFN-.gamma. responses were equivalent between VLP vaccine groups regardless of stimulating antigen (p>0.05; FIG. 17B). Recombinant HA and inactivated virus were inefficient stimulators of IFN-.gamma. production compared to the HA533 peptide. HA533 is the immunodominant CD8.sup.+ T cell epitope in BALB/c mice and is conserved in all HA vaccine antigens used in this study. Overlapping peptide pools spanning the entire HA molecule were also used to stimulate cells and no differences were observed between COBRA and polyvalent vaccines for any of the pools. Similar to the ASC data, no IFN-.gamma. responses were detectable above background in mock vaccinated animals at 6 DPI.
Passive Transfer of Immune Sera
[0220] The contribution of serum factors to protection from clade 1 challenge was evaluated using a passive transfer model. Nine-week old recipient mice were administered pooled sera via IP injection from COBRA, polyvalent and mock vaccinated mice. The next day, recipient mice were challenged with the clade 1 reassortant A/Vietnam/1203/2004 virus as before. Regardless of transferred serum, all recipient mice lost weight and became visibly ill (FIGS. 18A and 18B). COBRA serum recipient mice lost less weight than polyvalent recipient mice with maximum losses of 5.2% (6 DPI) and 11.8% (7 DPI), respectively (p<0.05 at 7 DPI). COBRA serum recipient mice also began to resolve the clinical symptoms more rapidly than polyvalent recipient mice (p<0.05 at 7 DPI). Although COBRA serum prevented recipient mice from developing illness more efficiently than polyvalent serum, both COBRA and polyvalent serum protected all recipient mice from death. Conversely, all mice receiving serum from mock vaccinated mice rapidly lost weight, became visibly ill and reached experimental endpoint by 7 DPI.
[0221] In view of the many possible embodiments to which the principles of the disclosed invention may be applied, it should be recognized that the illustrated embodiments are only examples of the disclosure and should not be taken as limiting the scope of the invention. Rather, the scope of the invention is defined by the following claims. We therefore claim as our invention all that comes within the scope and spirit of these claims.
Sequence CWU 1
1
1111707DNAArtificial SequenceSynthetic polynucleotideCDS(1)..(1707)
1atg gaa aag atc gtg ctg ctg ctg gct atc gtg agc ctg gtg aag agc
48Met Glu Lys Ile Val Leu Leu Leu Ala Ile Val Ser Leu Val Lys Ser 1
5 10 15 gac cag att tgc atc ggc tac cac gcc aac aac agc acc gag cag
gtg 96Asp Gln Ile Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Glu Gln
Val 20 25 30 gac acc atc atg gaa aag aac gtc acc gtg acc cac gcc
cag gac atc 144Asp Thr Ile Met Glu Lys Asn Val Thr Val Thr His Ala
Gln Asp Ile 35 40 45 ctg gaa aag acc cac aac ggc aag ctg tgc gac
ctg gac ggc gtg aag 192Leu Glu Lys Thr His Asn Gly Lys Leu Cys Asp
Leu Asp Gly Val Lys 50 55 60 ccc ctg atc ctg agg gac tgc agc gtg
gcc ggc tgg ctg ctg ggc aac 240Pro Leu Ile Leu Arg Asp Cys Ser Val
Ala Gly Trp Leu Leu Gly Asn 65 70 75 80 ccc atg tgc gac gag ttc atc
aac gtg ccc gag tgg agc tac atc gtg 288Pro Met Cys Asp Glu Phe Ile
Asn Val Pro Glu Trp Ser Tyr Ile Val 85 90 95 gag aag gcc aac ccc
gcc aac gac ctg tgc tac ccc ggc aac ttc aac 336Glu Lys Ala Asn Pro
Ala Asn Asp Leu Cys Tyr Pro Gly Asn Phe Asn 100 105 110 gac tac gag
gaa ctg aag cac ctg ctg tcc agg atc aac cac ttc gag 384Asp Tyr Glu
Glu Leu Lys His Leu Leu Ser Arg Ile Asn His Phe Glu 115 120 125 aag
atc cag atc atc ccc aag agc agc tgg tcc gac cac gag gcc agc 432Lys
Ile Gln Ile Ile Pro Lys Ser Ser Trp Ser Asp His Glu Ala Ser 130 135
140 agc ggc gtg agc agc gcc tgc cca tac cag ggc agc ccc agc ttc ttc
480Ser Gly Val Ser Ser Ala Cys Pro Tyr Gln Gly Ser Pro Ser Phe Phe
145 150 155 160 aga aac gtg gtg tgg ctg atc aag aag aac aac acc tac
ccc acc atc 528Arg Asn Val Val Trp Leu Ile Lys Lys Asn Asn Thr Tyr
Pro Thr Ile 165 170 175 aag agg tcc tac aac aac acc aac cag gaa gat
ctg ctg gtc ctg tgg 576Lys Arg Ser Tyr Asn Asn Thr Asn Gln Glu Asp
Leu Leu Val Leu Trp 180 185 190 ggc atc cac cac cct aat gac gcc gcc
gaa cag acc agg ctg tac cag 624Gly Ile His His Pro Asn Asp Ala Ala
Glu Gln Thr Arg Leu Tyr Gln 195 200 205 aac ccc acc acc tac atc agc
gtg ggc aca agc acc ctg aac cag agg 672Asn Pro Thr Thr Tyr Ile Ser
Val Gly Thr Ser Thr Leu Asn Gln Arg 210 215 220 ctg gtg ccc aag atc
gcc acc agg tcc aag gtg aac gga cag tcc ggc 720Leu Val Pro Lys Ile
Ala Thr Arg Ser Lys Val Asn Gly Gln Ser Gly 225 230 235 240 agg atg
gaa ttc ttc tgg acc atc ctg aag cct aac gac gcc atc aac 768Arg Met
Glu Phe Phe Trp Thr Ile Leu Lys Pro Asn Asp Ala Ile Asn 245 250 255
ttc gag agc aac ggc aac ttt atc gcc ccc gag tac gcc tac aag atc
816Phe Glu Ser Asn Gly Asn Phe Ile Ala Pro Glu Tyr Ala Tyr Lys Ile
260 265 270 gtg aag aag ggc gac agc gcc atc atg aag agc gag ctg gaa
tac ggc 864Val Lys Lys Gly Asp Ser Ala Ile Met Lys Ser Glu Leu Glu
Tyr Gly 275 280 285 aac tgc aac acc aag tgc cag acc ccc atc ggc gcc
atc aac agc agc 912Asn Cys Asn Thr Lys Cys Gln Thr Pro Ile Gly Ala
Ile Asn Ser Ser 290 295 300 atg ccc ttc cac aac atc cac ccc ctg acc
atc ggc gag tgc ccc aag 960Met Pro Phe His Asn Ile His Pro Leu Thr
Ile Gly Glu Cys Pro Lys 305 310 315 320 tac gtg aag agc aac agg ctg
gtg ctg gcc acc ggc ctg agg aac agc 1008Tyr Val Lys Ser Asn Arg Leu
Val Leu Ala Thr Gly Leu Arg Asn Ser 325 330 335 ccc cag aga gag agc
aga aga aag aag agg ggc ctg ttc ggc gct atc 1056Pro Gln Arg Glu Ser
Arg Arg Lys Lys Arg Gly Leu Phe Gly Ala Ile 340 345 350 gcc ggc ttc
atc gag ggc ggc tgg cag ggc atg gtg gac ggg tgg tac 1104Ala Gly Phe
Ile Glu Gly Gly Trp Gln Gly Met Val Asp Gly Trp Tyr 355 360 365 ggc
tac cac cac tct aac gag cag ggc agc ggc tac gcc gcc gac aaa 1152Gly
Tyr His His Ser Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Lys 370 375
380 gag agc acc cag aag gcc atc gac ggc gtc acc aac aag gtg aac agc
1200Glu Ser Thr Gln Lys Ala Ile Asp Gly Val Thr Asn Lys Val Asn Ser
385 390 395 400 atc atc gac aag atg aac acc cag ttc gag gcc gtg ggc
aga gag ttc 1248Ile Ile Asp Lys Met Asn Thr Gln Phe Glu Ala Val Gly
Arg Glu Phe 405 410 415 aac aac ctg gaa agg cgg atc gag aac ctg aac
aag aaa atg gaa gat 1296Asn Asn Leu Glu Arg Arg Ile Glu Asn Leu Asn
Lys Lys Met Glu Asp 420 425 430 ggc ttc ctg gac gtg tgg acc tac aac
gcc gag ctg ctg gtg ctg atg 1344Gly Phe Leu Asp Val Trp Thr Tyr Asn
Ala Glu Leu Leu Val Leu Met 435 440 445 gaa aac gag agg acc ctg gac
ttc cac gac agc aac gtg aag aac ctg 1392Glu Asn Glu Arg Thr Leu Asp
Phe His Asp Ser Asn Val Lys Asn Leu 450 455 460 tac gac aaa gtg cgg
ctg cag ctg agg gac aac gcc aaa gag ctg ggc 1440Tyr Asp Lys Val Arg
Leu Gln Leu Arg Asp Asn Ala Lys Glu Leu Gly 465 470 475 480 aac ggc
tgc ttc gag ttc tac cac aag tgc gac aac gag tgc atg gaa 1488Asn Gly
Cys Phe Glu Phe Tyr His Lys Cys Asp Asn Glu Cys Met Glu 485 490 495
agc gtg agg aac ggc acc tac gac tac ccc cag tac agc gag gaa gcc
1536Ser Val Arg Asn Gly Thr Tyr Asp Tyr Pro Gln Tyr Ser Glu Glu Ala
500 505 510 agg ctg aag agg gaa gag atc agc gga gtg aag ctg gaa agc
atc ggc 1584Arg Leu Lys Arg Glu Glu Ile Ser Gly Val Lys Leu Glu Ser
Ile Gly 515 520 525 acc tac cag atc ctg agc atc tac agc acc gtc gcc
agc agc ctg gcc 1632Thr Tyr Gln Ile Leu Ser Ile Tyr Ser Thr Val Ala
Ser Ser Leu Ala 530 535 540 ctg gct atc atg gtg gcc gga ctg agc ctg
tgg atg tgc agc aac ggc 1680Leu Ala Ile Met Val Ala Gly Leu Ser Leu
Trp Met Cys Ser Asn Gly 545 550 555 560 agc ctg cag tgc agg atc tgc
atc tga 1707Ser Leu Gln Cys Arg Ile Cys Ile 565 2568PRTArtificial
SequenceSynthetic Construct 2Met Glu Lys Ile Val Leu Leu Leu Ala
Ile Val Ser Leu Val Lys Ser 1 5 10 15 Asp Gln Ile Cys Ile Gly Tyr
His Ala Asn Asn Ser Thr Glu Gln Val 20 25 30 Asp Thr Ile Met Glu
Lys Asn Val Thr Val Thr His Ala Gln Asp Ile 35 40 45 Leu Glu Lys
Thr His Asn Gly Lys Leu Cys Asp Leu Asp Gly Val Lys 50 55 60 Pro
Leu Ile Leu Arg Asp Cys Ser Val Ala Gly Trp Leu Leu Gly Asn 65 70
75 80 Pro Met Cys Asp Glu Phe Ile Asn Val Pro Glu Trp Ser Tyr Ile
Val 85 90 95 Glu Lys Ala Asn Pro Ala Asn Asp Leu Cys Tyr Pro Gly
Asn Phe Asn 100 105 110 Asp Tyr Glu Glu Leu Lys His Leu Leu Ser Arg
Ile Asn His Phe Glu 115 120 125 Lys Ile Gln Ile Ile Pro Lys Ser Ser
Trp Ser Asp His Glu Ala Ser 130 135 140 Ser Gly Val Ser Ser Ala Cys
Pro Tyr Gln Gly Ser Pro Ser Phe Phe 145 150 155 160 Arg Asn Val Val
Trp Leu Ile Lys Lys Asn Asn Thr Tyr Pro Thr Ile 165 170 175 Lys Arg
Ser Tyr Asn Asn Thr Asn Gln Glu Asp Leu Leu Val Leu Trp 180 185 190
Gly Ile His His Pro Asn Asp Ala Ala Glu Gln Thr Arg Leu Tyr Gln 195
200 205 Asn Pro Thr Thr Tyr Ile Ser Val Gly Thr Ser Thr Leu Asn Gln
Arg 210 215 220 Leu Val Pro Lys Ile Ala Thr Arg Ser Lys Val Asn Gly
Gln Ser Gly 225 230 235 240 Arg Met Glu Phe Phe Trp Thr Ile Leu Lys
Pro Asn Asp Ala Ile Asn 245 250 255 Phe Glu Ser Asn Gly Asn Phe Ile
Ala Pro Glu Tyr Ala Tyr Lys Ile 260 265 270 Val Lys Lys Gly Asp Ser
Ala Ile Met Lys Ser Glu Leu Glu Tyr Gly 275 280 285 Asn Cys Asn Thr
Lys Cys Gln Thr Pro Ile Gly Ala Ile Asn Ser Ser 290 295 300 Met Pro
Phe His Asn Ile His Pro Leu Thr Ile Gly Glu Cys Pro Lys 305 310 315
320 Tyr Val Lys Ser Asn Arg Leu Val Leu Ala Thr Gly Leu Arg Asn Ser
325 330 335 Pro Gln Arg Glu Ser Arg Arg Lys Lys Arg Gly Leu Phe Gly
Ala Ile 340 345 350 Ala Gly Phe Ile Glu Gly Gly Trp Gln Gly Met Val
Asp Gly Trp Tyr 355 360 365 Gly Tyr His His Ser Asn Glu Gln Gly Ser
Gly Tyr Ala Ala Asp Lys 370 375 380 Glu Ser Thr Gln Lys Ala Ile Asp
Gly Val Thr Asn Lys Val Asn Ser 385 390 395 400 Ile Ile Asp Lys Met
Asn Thr Gln Phe Glu Ala Val Gly Arg Glu Phe 405 410 415 Asn Asn Leu
Glu Arg Arg Ile Glu Asn Leu Asn Lys Lys Met Glu Asp 420 425 430 Gly
Phe Leu Asp Val Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Met 435 440
445 Glu Asn Glu Arg Thr Leu Asp Phe His Asp Ser Asn Val Lys Asn Leu
450 455 460 Tyr Asp Lys Val Arg Leu Gln Leu Arg Asp Asn Ala Lys Glu
Leu Gly 465 470 475 480 Asn Gly Cys Phe Glu Phe Tyr His Lys Cys Asp
Asn Glu Cys Met Glu 485 490 495 Ser Val Arg Asn Gly Thr Tyr Asp Tyr
Pro Gln Tyr Ser Glu Glu Ala 500 505 510 Arg Leu Lys Arg Glu Glu Ile
Ser Gly Val Lys Leu Glu Ser Ile Gly 515 520 525 Thr Tyr Gln Ile Leu
Ser Ile Tyr Ser Thr Val Ala Ser Ser Leu Ala 530 535 540 Leu Ala Ile
Met Val Ala Gly Leu Ser Leu Trp Met Cys Ser Asn Gly 545 550 555 560
Ser Leu Gln Cys Arg Ile Cys Ile 565 31350DNAArtificial
SequenceSynthetic polynucleotideCDS(1)..(1350) 3atg aat cct aat aag
aag atc atc aca atc gga agc atc tgc atg gtg 48Met Asn Pro Asn Lys
Lys Ile Ile Thr Ile Gly Ser Ile Cys Met Val 1 5 10 15 aca gga atg
gtg agc ctg atg ctg cag atc gga aat ctg atc agc atc 96Thr Gly Met
Val Ser Leu Met Leu Gln Ile Gly Asn Leu Ile Ser Ile 20 25 30 tgg
gtg agc cac agc atc cac aca gga aat cag cac aag gcc gag cct 144Trp
Val Ser His Ser Ile His Thr Gly Asn Gln His Lys Ala Glu Pro 35 40
45 atc agc aat aca aat ttt ctg aca gag aag gcc gtg gcc agc gtg aag
192Ile Ser Asn Thr Asn Phe Leu Thr Glu Lys Ala Val Ala Ser Val Lys
50 55 60 ctg gcc gga aat agc agc ctg tgc cct atc aat gga tgg gcc
gtg tac 240Leu Ala Gly Asn Ser Ser Leu Cys Pro Ile Asn Gly Trp Ala
Val Tyr 65 70 75 80 agc aag gat aat agc atc aga atc gga agc aag gga
gat gtg ttt gtg 288Ser Lys Asp Asn Ser Ile Arg Ile Gly Ser Lys Gly
Asp Val Phe Val 85 90 95 atc aga gag cct ttt atc agc tgc agc cac
ctg gag tgc aga aca ttt 336Ile Arg Glu Pro Phe Ile Ser Cys Ser His
Leu Glu Cys Arg Thr Phe 100 105 110 ttt ctg aca cag gga gcc ctg ctg
aat gat aag cac agc aat gga aca 384Phe Leu Thr Gln Gly Ala Leu Leu
Asn Asp Lys His Ser Asn Gly Thr 115 120 125 gtg aag gat aga agc cct
cac aga aca ctg atg agc tgc cct gtg gga 432Val Lys Asp Arg Ser Pro
His Arg Thr Leu Met Ser Cys Pro Val Gly 130 135 140 gag gcc cct agc
cct tac aat agc aga ttt gag agc gtg gcc tgg agc 480Glu Ala Pro Ser
Pro Tyr Asn Ser Arg Phe Glu Ser Val Ala Trp Ser 145 150 155 160 gcc
agc gcc tgc cac gat gga aca agc tgg ctg aca atc gga atc agc 528Ala
Ser Ala Cys His Asp Gly Thr Ser Trp Leu Thr Ile Gly Ile Ser 165 170
175 gga cct gat aat gga gcc gtg gcc gtg ctg aag tac aat gga atc atc
576Gly Pro Asp Asn Gly Ala Val Ala Val Leu Lys Tyr Asn Gly Ile Ile
180 185 190 aca gat aca atc aag agc tgg aga aat aat atc ctg aga aca
cag gag 624Thr Asp Thr Ile Lys Ser Trp Arg Asn Asn Ile Leu Arg Thr
Gln Glu 195 200 205 agc gag tgc gcc tgc gtg aat gga agc tgc ttt aca
gtg atg aca gat 672Ser Glu Cys Ala Cys Val Asn Gly Ser Cys Phe Thr
Val Met Thr Asp 210 215 220 gga cct agc aat gga cag gcc agc cac aag
atc ttt aag atg gag aag 720Gly Pro Ser Asn Gly Gln Ala Ser His Lys
Ile Phe Lys Met Glu Lys 225 230 235 240 gga aag gtg gtg aag agc gtg
gag ctg gat gcc cct aat tac cac tac 768Gly Lys Val Val Lys Ser Val
Glu Leu Asp Ala Pro Asn Tyr His Tyr 245 250 255 gag gag tgc agc tgc
tac cct gat gcc gga gag atc aca tgc gtg tgc 816Glu Glu Cys Ser Cys
Tyr Pro Asp Ala Gly Glu Ile Thr Cys Val Cys 260 265 270 aga gat aat
tgg cac gga agc aat aga cct tgg gtg agc ttt aat cag 864Arg Asp Asn
Trp His Gly Ser Asn Arg Pro Trp Val Ser Phe Asn Gln 275 280 285 aat
ctg gag tac cag atc gga tac atc tgc agc gga gtg ttt gga gat 912Asn
Leu Glu Tyr Gln Ile Gly Tyr Ile Cys Ser Gly Val Phe Gly Asp 290 295
300 aat cct aga cct aat gat gga aca gga agc tgc gga cct gtg agc agc
960Asn Pro Arg Pro Asn Asp Gly Thr Gly Ser Cys Gly Pro Val Ser Ser
305 310 315 320 aat gga gcc tac gga gtg aag gga ttt agc ttt aag tac
gga aat gga 1008Asn Gly Ala Tyr Gly Val Lys Gly Phe Ser Phe Lys Tyr
Gly Asn Gly 325 330 335 gtg tgg atc gga aga aca aag agc aca aat agc
aga agc gga ttt gag 1056Val Trp Ile Gly Arg Thr Lys Ser Thr Asn Ser
Arg Ser Gly Phe Glu 340 345 350 atg atc tgg gac cct aat gga tgg aca
gag aca gat agc agc ttt agc 1104Met Ile Trp Asp Pro Asn Gly Trp Thr
Glu Thr Asp Ser Ser Phe Ser 355 360 365 gtg aag cag gat atc gtg gcc
atc aca gat tgg agc gga tac agc gga 1152Val Lys Gln Asp Ile Val Ala
Ile Thr Asp Trp Ser Gly Tyr Ser Gly 370 375 380 agc ttt gtg cag cac
cct gag ctg aca gga ctg gat tgc atc aga cct 1200Ser Phe Val Gln His
Pro Glu Leu Thr Gly Leu Asp Cys Ile Arg Pro 385 390 395 400 tgc ttt
tgg gtg gag ctg atc aga gga aga cct aag gag agc aca atc 1248Cys Phe
Trp Val Glu Leu Ile Arg Gly Arg Pro Lys Glu Ser Thr Ile 405 410 415
tgg aca agc gga agc agc atc agc ttt tgc gga
gtg aat agc gat aca 1296Trp Thr Ser Gly Ser Ser Ile Ser Phe Cys Gly
Val Asn Ser Asp Thr 420 425 430 gtg gga tgg agc tgg cct gat gga gcc
gag ctg cct ttt aca atc gat 1344Val Gly Trp Ser Trp Pro Asp Gly Ala
Glu Leu Pro Phe Thr Ile Asp 435 440 445 aag tga 1350Lys
4449PRTArtificial SequenceSynthetic Construct 4Met Asn Pro Asn Lys
Lys Ile Ile Thr Ile Gly Ser Ile Cys Met Val 1 5 10 15 Thr Gly Met
Val Ser Leu Met Leu Gln Ile Gly Asn Leu Ile Ser Ile 20 25 30 Trp
Val Ser His Ser Ile His Thr Gly Asn Gln His Lys Ala Glu Pro 35 40
45 Ile Ser Asn Thr Asn Phe Leu Thr Glu Lys Ala Val Ala Ser Val Lys
50 55 60 Leu Ala Gly Asn Ser Ser Leu Cys Pro Ile Asn Gly Trp Ala
Val Tyr 65 70 75 80 Ser Lys Asp Asn Ser Ile Arg Ile Gly Ser Lys Gly
Asp Val Phe Val 85 90 95 Ile Arg Glu Pro Phe Ile Ser Cys Ser His
Leu Glu Cys Arg Thr Phe 100 105 110 Phe Leu Thr Gln Gly Ala Leu Leu
Asn Asp Lys His Ser Asn Gly Thr 115 120 125 Val Lys Asp Arg Ser Pro
His Arg Thr Leu Met Ser Cys Pro Val Gly 130 135 140 Glu Ala Pro Ser
Pro Tyr Asn Ser Arg Phe Glu Ser Val Ala Trp Ser 145 150 155 160 Ala
Ser Ala Cys His Asp Gly Thr Ser Trp Leu Thr Ile Gly Ile Ser 165 170
175 Gly Pro Asp Asn Gly Ala Val Ala Val Leu Lys Tyr Asn Gly Ile Ile
180 185 190 Thr Asp Thr Ile Lys Ser Trp Arg Asn Asn Ile Leu Arg Thr
Gln Glu 195 200 205 Ser Glu Cys Ala Cys Val Asn Gly Ser Cys Phe Thr
Val Met Thr Asp 210 215 220 Gly Pro Ser Asn Gly Gln Ala Ser His Lys
Ile Phe Lys Met Glu Lys 225 230 235 240 Gly Lys Val Val Lys Ser Val
Glu Leu Asp Ala Pro Asn Tyr His Tyr 245 250 255 Glu Glu Cys Ser Cys
Tyr Pro Asp Ala Gly Glu Ile Thr Cys Val Cys 260 265 270 Arg Asp Asn
Trp His Gly Ser Asn Arg Pro Trp Val Ser Phe Asn Gln 275 280 285 Asn
Leu Glu Tyr Gln Ile Gly Tyr Ile Cys Ser Gly Val Phe Gly Asp 290 295
300 Asn Pro Arg Pro Asn Asp Gly Thr Gly Ser Cys Gly Pro Val Ser Ser
305 310 315 320 Asn Gly Ala Tyr Gly Val Lys Gly Phe Ser Phe Lys Tyr
Gly Asn Gly 325 330 335 Val Trp Ile Gly Arg Thr Lys Ser Thr Asn Ser
Arg Ser Gly Phe Glu 340 345 350 Met Ile Trp Asp Pro Asn Gly Trp Thr
Glu Thr Asp Ser Ser Phe Ser 355 360 365 Val Lys Gln Asp Ile Val Ala
Ile Thr Asp Trp Ser Gly Tyr Ser Gly 370 375 380 Ser Phe Val Gln His
Pro Glu Leu Thr Gly Leu Asp Cys Ile Arg Pro 385 390 395 400 Cys Phe
Trp Val Glu Leu Ile Arg Gly Arg Pro Lys Glu Ser Thr Ile 405 410 415
Trp Thr Ser Gly Ser Ser Ile Ser Phe Cys Gly Val Asn Ser Asp Thr 420
425 430 Val Gly Trp Ser Trp Pro Asp Gly Ala Glu Leu Pro Phe Thr Ile
Asp 435 440 445 Lys 5759DNAArtificial SequenceSynthetic
polynucleotideCDS(1)..(759) 5atg agc ctg ctg acc gag gtg gag aca
tac gtg ctg tcc atc atc ccc 48Met Ser Leu Leu Thr Glu Val Glu Thr
Tyr Val Leu Ser Ile Ile Pro 1 5 10 15 agc ggc cct ctg aag gcc gag
atc gcc cag aga ctg gaa gat gtg ttc 96Ser Gly Pro Leu Lys Ala Glu
Ile Ala Gln Arg Leu Glu Asp Val Phe 20 25 30 gcc ggc aag aac acc
gac ctg gaa gtg ctg atg gaa tgg ctg aaa acc 144Ala Gly Lys Asn Thr
Asp Leu Glu Val Leu Met Glu Trp Leu Lys Thr 35 40 45 aga ccc atc
ctg agc cct ctg acc aag ggc atc ctg ggc ttc gtg ttc 192Arg Pro Ile
Leu Ser Pro Leu Thr Lys Gly Ile Leu Gly Phe Val Phe 50 55 60 acc
ctg acc gtg ccc agc gag aga ggc ctg cag agg cgg aga ttc gtg 240Thr
Leu Thr Val Pro Ser Glu Arg Gly Leu Gln Arg Arg Arg Phe Val 65 70
75 80 cag aac gcc ctg aac ggc aac ggc gac ccc aac aac atg gac aag
gcc 288Gln Asn Ala Leu Asn Gly Asn Gly Asp Pro Asn Asn Met Asp Lys
Ala 85 90 95 gtg aag ctg tac aga aag ctg aag cgg gag atc acc ttc
cac ggc gcc 336Val Lys Leu Tyr Arg Lys Leu Lys Arg Glu Ile Thr Phe
His Gly Ala 100 105 110 aaa gag atc agc ctg agc tac agc gct ggc gcc
ctg gcc agc tgc atg 384Lys Glu Ile Ser Leu Ser Tyr Ser Ala Gly Ala
Leu Ala Ser Cys Met 115 120 125 ggc ctg atc tac aac aga atg ggc gcc
gtg acc acc gag gtg gcc ttc 432Gly Leu Ile Tyr Asn Arg Met Gly Ala
Val Thr Thr Glu Val Ala Phe 130 135 140 ggc ctg gtc tgc gcc acc tgc
gag cag atc gcc gac agc cag cac aga 480Gly Leu Val Cys Ala Thr Cys
Glu Gln Ile Ala Asp Ser Gln His Arg 145 150 155 160 tcc cac aga cag
atg gtc acc acc acc aac ccc ctg atc aga cac gag 528Ser His Arg Gln
Met Val Thr Thr Thr Asn Pro Leu Ile Arg His Glu 165 170 175 aac aga
atg gtg ctg gcc tct acc acc gcc aag gcc atg gaa cag atg 576Asn Arg
Met Val Leu Ala Ser Thr Thr Ala Lys Ala Met Glu Gln Met 180 185 190
gcc ggc agc agc gag cag gcc gcc gag gct atg gaa gtc gcc tct cag
624Ala Gly Ser Ser Glu Gln Ala Ala Glu Ala Met Glu Val Ala Ser Gln
195 200 205 gct agg cag atg gtc cag gcc atg aga acc atc ggc acc cac
ccc agc 672Ala Arg Gln Met Val Gln Ala Met Arg Thr Ile Gly Thr His
Pro Ser 210 215 220 agc tct gct ggc ctg aag aac gac ctg ctg gaa aac
ctg cag gcc tac 720Ser Ser Ala Gly Leu Lys Asn Asp Leu Leu Glu Asn
Leu Gln Ala Tyr 225 230 235 240 cag aaa aga atg ggc gtc cag atg cag
aga ttc aag tga 759Gln Lys Arg Met Gly Val Gln Met Gln Arg Phe Lys
245 250 6252PRTArtificial SequenceSynthetic Construct 6Met Ser Leu
Leu Thr Glu Val Glu Thr Tyr Val Leu Ser Ile Ile Pro 1 5 10 15 Ser
Gly Pro Leu Lys Ala Glu Ile Ala Gln Arg Leu Glu Asp Val Phe 20 25
30 Ala Gly Lys Asn Thr Asp Leu Glu Val Leu Met Glu Trp Leu Lys Thr
35 40 45 Arg Pro Ile Leu Ser Pro Leu Thr Lys Gly Ile Leu Gly Phe
Val Phe 50 55 60 Thr Leu Thr Val Pro Ser Glu Arg Gly Leu Gln Arg
Arg Arg Phe Val 65 70 75 80 Gln Asn Ala Leu Asn Gly Asn Gly Asp Pro
Asn Asn Met Asp Lys Ala 85 90 95 Val Lys Leu Tyr Arg Lys Leu Lys
Arg Glu Ile Thr Phe His Gly Ala 100 105 110 Lys Glu Ile Ser Leu Ser
Tyr Ser Ala Gly Ala Leu Ala Ser Cys Met 115 120 125 Gly Leu Ile Tyr
Asn Arg Met Gly Ala Val Thr Thr Glu Val Ala Phe 130 135 140 Gly Leu
Val Cys Ala Thr Cys Glu Gln Ile Ala Asp Ser Gln His Arg 145 150 155
160 Ser His Arg Gln Met Val Thr Thr Thr Asn Pro Leu Ile Arg His Glu
165 170 175 Asn Arg Met Val Leu Ala Ser Thr Thr Ala Lys Ala Met Glu
Gln Met 180 185 190 Ala Gly Ser Ser Glu Gln Ala Ala Glu Ala Met Glu
Val Ala Ser Gln 195 200 205 Ala Arg Gln Met Val Gln Ala Met Arg Thr
Ile Gly Thr His Pro Ser 210 215 220 Ser Ser Ala Gly Leu Lys Asn Asp
Leu Leu Glu Asn Leu Gln Ala Tyr 225 230 235 240 Gln Lys Arg Met Gly
Val Gln Met Gln Arg Phe Lys 245 250 75511DNAArtificial
SequenceSynthetic construct 7cgacaatatt ggctattggc cattgcatac
gttgtatcta tatcataata tgtacattta 60tattggctca tgtccaatat gaccgccatg
ttgacattga ttattgacta gttattaata 120gtaatcaatt acggggtcat
tagttcatag cccatatatg gagttccgcg ttacataact 180tacggtaaat
ggcccgcctc gtgaccgccc aacgaccccc gcccattgac gtcaataatg
240acgtatgttc ccatagtaac gccaataggg actttccatt gacgtcaatg
ggtggagtat 300ttacggtaaa ctgcccactt ggcagtacat caagtgtatc
atatgccaag tccgccccta 360ttgacgtcaa tgacggtaaa tggcccgcct
ggcattatgc ccagtacatg accttacggg 420actttcctac ttggcagtac
atctacgtat tagtcatcgc tattaccatg gtgatgcggt 480tttggcagta
caccaatggg cgtggatagc ggtttgactc acggggattt ccaagtctcc
540accccattga cgtcaatggg agtttgtttt ggcaccaaaa tcaacgggac
tttccaaaat 600gtcgtaataa ccccgccccg ttgacgcaaa tgggcggtag
gcgtgtacgg tgggaggtct 660atataagcag agctcgttta gtgaaccgtc
agatcgcctg gagacgccat ccacgctgtt 720ttgacctcca tagaagacac
cgggaccgat ccagcctccg cggccgggaa cggtgcattg 780gaacgcggat
tccccgtgcc aagagtgacg taagtaccgc ctatagactc tataggcaca
840cccctttggc tcttatgcat gctatactgt ttttggcttg gggcctatac
acccccgctc 900cttatgctat aggtgatggt atagcttagc ctataggtgt
gggttattga ccattattga 960ccactcccct attggtgacg atactttcca
ttactaatcc ataacatggc tctttgccac 1020aactatctct attggctata
tgccaatact ctgtccttca gagactgaca cggactctgt 1080atttttacag
gatggggtcc catttattat ttacaaattc acatatacaa caacgccgtc
1140ccccgtgccc gcagttttta ttaaacatag cgtgggatct ccacgcgaat
ctcgggtacg 1200tgttccggac atgggctctt ctccggtagc ggcggagctt
ccacatccga gccctggtcc 1260catgcctcca gcggctcatg gtcgctcggc
agctccttgc tcctaacagt ggaggccaga 1320cttaggcaca gcacaatgcc
caccaccacc agtgtgccgc acaaggccgt ggcggtaggg 1380tatgtgtctg
aaaatgagct cggagattgg gctcgcaccg tgacgcagat ggaagactta
1440aggcagcggc agaagaagat gcaggcagct gagttgttgt attctgataa
gagtcagagg 1500taactcccgt tgcggtgctg ttaacggtgg agggcagtgt
agtctgagca gtactcgttg 1560ctgccgcgcg cgccaccaga cataatagct
gacagactaa cagactgttc ctttccatgg 1620gtcttttctg cagtcaccgt
ccaagcttat ggaaaagatc gtgctgctgc tggctatcgt 1680gagcctggtg
aagagcgacc agatttgcat cggctaccac gccaacaaca gcaccgagca
1740ggtggacacc atcatggaaa agaacgtcac cgtgacccac gcccaggaca
tcctggaaaa 1800gacccacaac ggcaagctgt gcgacctgga cggcgtgaag
cccctgatcc tgagggactg 1860cagcgtggcc ggctggctgc tgggcaaccc
catgtgcgac gagttcatca acgtgcccga 1920gtggagctac atcgtggaga
aggccaaccc cgccaacgac ctgtgctacc ccggcaactt 1980caacgactac
gaggaactga agcacctgct gtccaggatc aaccacttcg agaagatcca
2040gatcatcccc aagagcagct ggtccgacca cgaggccagc agcggcgtga
gcagcgcctg 2100cccataccag ggcagcccca gcttcttcag aaacgtggtg
tggctgatca agaagaacaa 2160cacctacccc accatcaaga ggtcctacaa
caacaccaac caggaagatc tgctggtcct 2220gtggggcatc caccacccta
atgacgccgc cgaacagacc aggctgtacc agaaccccac 2280cacctacatc
agcgtgggca caagcaccct gaaccagagg ctggtgccca agatcgccac
2340caggtccaag gtgaacggac agtccggcag gatggaattc ttctggacca
tcctgaagcc 2400taacgacgcc atcaacttcg agagcaacgg caactttatc
gcccccgagt acgcctacaa 2460gatcgtgaag aagggcgaca gcgccatcat
gaagagcgag ctggaatacg gcaactgcaa 2520caccaagtgc cagaccccca
tcggcgccat caacagcagc atgcccttcc acaacatcca 2580ccccctgacc
atcggcgagt gccccaagta cgtgaagagc aacaggctgg tgctggccac
2640cggcctgagg aacagccccc agagagagag cagaagaaag aagaggggcc
tgttcggcgc 2700tatcgccggc ttcatcgagg gcggctggca gggcatggtg
gacgggtggt acggctacca 2760ccactctaac gagcagggca gcggctacgc
cgccgacaaa gagagcaccc agaaggccat 2820cgacggcgtc accaacaagg
tgaacagcat catcgacaag atgaacaccc agttcgaggc 2880cgtgggcaga
gagttcaaca acctggaaag gcggatcgag aacctgaaca agaaaatgga
2940agatggcttc ctggacgtgt ggacctacaa cgccgagctg ctggtgctga
tggaaaacga 3000gaggaccctg gacttccacg acagcaacgt gaagaacctg
tacgacaaag tgcggctgca 3060gctgagggac aacgccaaag agctgggcaa
cggctgcttc gagttctacc acaagtgcga 3120caacgagtgc atggaaagcg
tgaggaacgg cacctacgac tacccccagt acagcgagga 3180agccaggctg
aagagggaag agatcagcgg agtgaagctg gaaagcatcg gcacctacca
3240gatcctgagc atctacagca ccgtcgccag cagcctggcc ctggctatca
tggtggccgg 3300actgagcctg tggatgtgca gcaacggcag cctgcagtgc
aggatctgca tcggatcctc 3360gcaatcccta gggctgtgcc ttctagttgc
cagccaaact gttgtttgcc cctcccccgt 3420gccttccttg accctggaag
gtgccactcc cactgtcctt tcctaataaa atgaggaaat 3480tgcatcgcat
tgtctgagta ggtgtcattc tattctgggg ggtggggtgg ggcaggacag
3540caagggggag gattgggaag acaatagcag gcatgctggg gatgcggtgg
gctctatgtt 3600cagaacgctc ggttgccgcc gggcgttttt tatctagagt
cgacaaattc agaagaactc 3660gtcaagaagg cgatagaagg cgatgcgctg
cgaatcggga gcggcgatac cgtaaagcac 3720gaggaagcgg tcagcccatt
cgccgccaag ctcttcagca atatcacggg tagccaacgc 3780tatgtcctga
tagcggtccg ccacacccag ccggccacag tcgatgaatc cagaaaagcg
3840gccattttcc accatgatat tcggcaagca ggcatcgcca tgggtcacga
cgagatcctc 3900gccgtcgggc atgctcgcct tgagcctggc gaacagttcg
gctggcgcga gcccctgatg 3960ctcttcgtcc agatcatcct gatcgacaag
accggcttcc atccgagtac gtgctcgctc 4020gatgcgatgt ttcgcttggt
ggtcgaatgg gcaggtagcc ggatcaagcg tatgcagccg 4080ccgcattgca
tcagccatga tggatacttt ctcggcagga gcaaggtgag atgacaggag
4140atcctgcccc ggcacttcgc ccaatagcag ccagtccctt cccgcttcag
tgacaacgtc 4200gagcacagct gcgcaaggaa cgcccgtcgt ggccagccac
gatagccgcg ctgcctcgtc 4260ttgcagttca ttcagggcac cggacaggtc
ggtcttgaca aaaagaaccg ggcgcccctg 4320cgctgacagc cggaacacgg
cggcatcaga gcagccgatt gtctgttgtg cccagtcata 4380gccgaatagc
ctctccaccc aagcggccgg agaacctgcg tgcaatccat cttgttcaat
4440catgcgaaac gatcctcatc ctgtctcttg atcagatctt gatcccctgc
gccatcagat 4500ccttggcggc gagaaagcca tccagtttac tttgcagggc
ttcccaacct taccagaggg 4560cgccccagct ggcaattccg gttcgcttgc
tgtccataaa accgcccagt ctagctatcg 4620ccatgtaagc ccactgcaag
ctacctgctt tctctttgcg cttgcgtttt cccttgtcca 4680gatagcccag
tagctgacat tcatccgggg tcagcaccgt ttctgcggac tggctttcta
4740cgtgaaaagg atctaggtga agatcctttt tgataatctc atgaccaaaa
tcccttaacg 4800tgagttttcg ttccactgag cgtcagaccc cgtagaaaag
atcaaaggat cttcttgaga 4860tccttttttt ctgcgcgtaa tctgctgctt
gcaaacaaaa aaaccaccgc taccagcggt 4920ggtttgtttg ccggatcaag
agctaccaac tctttttccg aaggtaactg gcttcagcag 4980agcgcagata
ccaaatactg tccttctagt gtagccgtag ttaggccacc acttcaagaa
5040ctctgtagca ccgcctacat acctcgctct gctaatcctg ttaccagtgg
ctgctgccag 5100tggcgataag tcgtgtctta ccgggttgga ctcaagacga
tagttaccgg ataaggcgca 5160gcggtcgggc tgaacggggg gttcgtgcac
acagcccagc ttggagcgaa cgacctacac 5220cgaactgaga tacctacagc
gtgagctatg agaaagcgcc acgcttcccg aagggagaaa 5280ggcggacagg
tatccggtaa gcggcagggt cggaacagga gagcgcacga gggagcttcc
5340agggggaaac gcctggtatc tttatagtcc tgtcgggttt cgccacctct
gacttgagcg 5400tcgatttttg tgatgctcgt caggggggcg gagcctatgg
aaaaacgcca gcaacgcggc 5460ctttttacgg ttcctgggct tttgctggcc
ttttgctcac atgttgtcga c 551185769DNAArtificial SequenceSynthetic
construct 8tcgcgcgttt cggtgatgac ggtgaaaacc tctgacacat gcagctcccg
gagacggtca 60cagcttgtct gtaagcggat gccgggagca gacaagcccg tcagggcgcg
tcagcgggtg 120ttggcgggtg tcggggctgg cttaactatg cggcatcaga
gcagattgta ctgagagtgc 180accatatgcg gtgtgaaata ccgcacagat
gcgtaaggag aaaataccgc atcagattgg 240ctattggcca ttgcatacgt
tgtatccata tcataatatg tacatttata ttggctcatg 300tccaacatta
ccgccatgtt gacattgatt attgactagt tattaatagt aatcaattac
360ggggtcatta gttcatagcc catatatgga gttccgcgtt acataactta
cggtaaatgg 420cccgcctggc tgaccgccca acgacccccg cccattgacg
tcaataatga cgtatgttcc 480catagtaacg ccaataggga ctttccattg
acgtcaatgg gtggagtatt tacggtaaac 540tgcccacttg gcagtacatc
aagtgtatca tatgccaagt acgcccccta ttgacgtcaa 600tgacggtaaa
tggcccgcct ggcattatgc ccagtacatg accttatggg actttcctac
660ttggcagtac atctacgtat tagtcatcgc tattaccatg gtgatgcggt
tttggcagta 720catcaatggg cgtggatagc ggtttgactc acggggattt
ccaagtctcc accccattga 780cgtcaatggg agtttgtttt ggcaccaaaa
tcaacgggac tttccaaaat gtcgtaacaa 840ctccgcccca ttgacgcaaa
tgggcggtag gcgtgtacgg tgggaggtct atataagcag 900agctcgttta
gtgaaccgtc agatcgcctg gagacgccat ccacgctgtt ttgacctcca
960tagaagacac cgggaccgat ccagcctcca tcggctcgca tctctccttc
acgcgcccgc 1020cgccctacct gaggccgcca tccacgccgg ttgagtcgcg
ttctgccgcc tcccgcctgt 1080ggtgcctcct gaactgcgtc cgccgtctag
gtaagtttaa agctcaggtc gagaccgggc 1140ctttgtccgg cgctcccttg
gagcctacct agactcagcc ggctctccac gctttgcctg 1200accctgcttg
ctcaactcta gttaacggtg gagggcagtg tagtctgagc agtactcgtt
1260gctgccgcgc gcgccaccag acataatagc tgacagacta acagactgtt
cctttccatg 1320ggtcttttct gcagtcaccg tcgtcgacac gatccgatat
cgccgccacc atgaatccta 1380ataagaagat catcacaatc
ggaagcatct gcatggtgac aggaatggtg agcctgatgc 1440tgcagatcgg
aaatctgatc agcatctggg tgagccacag catccacaca ggaaatcagc
1500acaaggccga gcctatcagc aatacaaatt ttctgacaga gaaggccgtg
gccagcgtga 1560agctggccgg aaatagcagc ctgtgcccta tcaatggatg
ggccgtgtac agcaaggata 1620atagcatcag aatcggaagc aagggagatg
tgtttgtgat cagagagcct tttatcagct 1680gcagccacct ggagtgcaga
acattttttc tgacacaggg agccctgctg aatgataagc 1740acagcaatgg
aacagtgaag gatagaagcc ctcacagaac actgatgagc tgccctgtgg
1800gagaggcccc tagcccttac aatagcagat ttgagagcgt ggcctggagc
gccagcgcct 1860gccacgatgg aacaagctgg ctgacaatcg gaatcagcgg
acctgataat ggagccgtgg 1920ccgtgctgaa gtacaatgga atcatcacag
atacaatcaa gagctggaga aataatatcc 1980tgagaacaca ggagagcgag
tgcgcctgcg tgaatggaag ctgctttaca gtgatgacag 2040atggacctag
caatggacag gccagccaca agatctttaa gatggagaag ggaaaggtgg
2100tgaagagcgt ggagctggat gcccctaatt accactacga ggagtgcagc
tgctaccctg 2160atgccggaga gatcacatgc gtgtgcagag ataattggca
cggaagcaat agaccttggg 2220tgagctttaa tcagaatctg gagtaccaga
tcggatacat ctgcagcgga gtgtttggag 2280ataatcctag acctaatgat
ggaacaggaa gctgcggacc tgtgagcagc aatggagcct 2340acggagtgaa
gggatttagc tttaagtacg gaaatggagt gtggatcgga agaacaaaga
2400gcacaaatag cagaagcgga tttgagatga tctgggaccc taatggatgg
acagagacag 2460atagcagctt tagcgtgaag caggatatcg tggccatcac
agattggagc ggatacagcg 2520gaagctttgt gcagcaccct gagctgacag
gactggattg catcagacct tgcttttggg 2580tggagctgat cagaggaaga
cctaaggaga gcacaatctg gacaagcgga agcagcatca 2640gcttttgcgg
agtgaatagc gatacagtgg gatggagctg gcctgatgga gccgagctgc
2700cttttacaat cgataagtga gcggccgctc tagaccaggc cctggatcca
gatctgctgt 2760gccttctagt tgccagccat ctgttgtttg cccctccccc
gtgccttcct tgaccctgga 2820aggtgccact cccactgtcc tttcctaata
aaatgaggaa attgcatcgc attgtctgag 2880taggtgtcat tctattctgg
ggggtggggt ggggcaggac agcaaggggg aggattggga 2940agacaatagc
aggcatgctg gggatgcggt gggctctatg ggtacccagg tgctgaagaa
3000ttgacccggt tcctcctggg ccagaaagaa gcaggcacat ccccttctct
gtgacacacc 3060ctgtccacgc ccctggttct tagttccagc cccactcata
ggacactcat agctcaggag 3120ggctccgcct tcaatcccac ccgctaaagt
acttggagcg gtctctccct ccctcatcag 3180cccaccaaac caaacctagc
ctccaagagt gggaagaaat taaagcaaga taggctatta 3240agtgcagagg
gagagaaaat gcctccaaca tgtgaggaag taatgagaga aatcatagaa
3300ttttaaggcc atgatttaag gccatcatgg ccttaatctt ccgcttcctc
gctcactgac 3360tcgctgcgct cggtcgttcg gctgcggcga gcggtatcag
ctcactcaaa ggcggtaata 3420cggttatcca cagaatcagg ggataacgca
ggaaagaaca tgtgagcaaa aggccagcaa 3480aaggccagga accgtaaaaa
ggccgcgttg ctggcgtttt tccataggct ccgcccccct 3540gacgagcatc
acaaaaatcg acgctcaagt cagaggtggc gaaacccgac aggactataa
3600agataccagg cgtttccccc tggaagctcc ctcgtgcgct ctcctgttcc
gaccctgccg 3660cttaccggat acctgtccgc ctttctccct tcgggaagcg
tggcgctttc tcatagctca 3720cgctgtaggt atctcagttc ggtgtaggtc
gttcgctcca agctgggctg tgtgcacgaa 3780ccccccgttc agcccgaccg
ctgcgcctta tccggtaact atcgtcttga gtccaacccg 3840gtaagacacg
acttatcgcc actggcagca gccactggta acaggattag cagagcgagg
3900tatgtaggcg gtgctacaga gttcttgaag tggtggccta actacggcta
cactagaaga 3960acagtatttg gtatctgcgc tctgctgaag ccagttacct
tcggaaaaag agttggtagc 4020tcttgatccg gcaaacaaac caccgctggt
agcggtggtt tttttgtttg caagcagcag 4080attacgcgca gaaaaaaagg
atctcaagaa gatcctttga tcttttctac ggggtctgac 4140gctcagtgga
acgaaaactc acgttaaggg attttggtca tgagattatc aaaaaggatc
4200ttcacctaga tccttttaaa ttaaaaatga agttttaaat caatctaaag
tatatatgag 4260taaacttggt ctgacagtta ccaatgctta atcagtgagg
cacctatctc agcgatctgt 4320ctatttcgtt catccatagt tgcctgactc
gggggggggg ggcgctgagg tctgcctcgt 4380gaagaaggtg ttgctgactc
ataccaggcc tgaatcgccc catcatccag ccagaaagtg 4440agggagccac
ggttgatgag agctttgttg taggtggacc agttggtgat tttgaacttt
4500tgctttgcca cggaacggtc tgcgttgtcg ggaagatgcg tgatctgatc
cttcaactca 4560gcaaaagttc gatttattca acaaagccgc cgtcccgtca
agtcagcgta atgctctgcc 4620agtgttacaa ccaattaacc aattctgatt
agaaaaactc atcgagcatc aaatgaaact 4680gcaatttatt catatcagga
ttatcaatac catatttttg aaaaagccgt ttctgtaatg 4740aaggagaaaa
ctcaccgagg cagttccata ggatggcaag atcctggtat cggtctgcga
4800ttccgactcg tccaacatca atacaaccta ttaatttccc ctcgtcaaaa
ataaggttat 4860caagtgagaa atcaccatga gtgacgactg aatccggtga
gaatggcaaa agcttatgca 4920tttctttcca gacttgttca acaggccagc
cattacgctc gtcatcaaaa tcactcgcat 4980caaccaaacc gttattcatt
cgtgattgcg cctgagcgag acgaaatacg cgatcgctgt 5040taaaaggaca
attacaaaca ggaatcgaat gcaaccggcg caggaacact gccagcgcat
5100caacaatatt ttcacctgaa tcaggatatt cttctaatac ctggaatgct
gttttcccgg 5160ggatcgcagt ggtgagtaac catgcatcat caggagtacg
gataaaatgc ttgatggtcg 5220gaagaggcat aaattccgtc agccagttta
gtctgaccat ctcatctgta acatcattgg 5280caacgctacc tttgccatgt
ttcagaaaca actctggcgc atcgggcttc ccatacaatc 5340gatagattgt
cgcacctgat tgcccgacat tatcgcgagc ccatttatac ccatataaat
5400cagcatccat gttggaattt aatcgcggcc tcgagcaaga cgtttcccgt
tgaatatggc 5460tcataacacc ccttgtatta ctgtttatgt aagcagacag
ttttattgtt catgatgata 5520tatttttatc ttgtgcaatg taacatcaga
gattttgaga cacaacgtgg ctttcccccc 5580ccccccatta ttgaagcatt
tatcagggtt attgtctcat gagcggatac atatttgaat 5640gtatttagaa
aaataaacaa ataggggttc cgcgcacatt tccccgaaaa gtgccacctg
5700acgtctaaga aaccattatt atcatgacat taacctataa aaataggcgt
atcacgaggc 5760cctttcgtc 576994598DNAArtificial SequenceArtificial
construct 9cgacaatatt ggctattggc cattgcatac gttgtatcta tatcataata
tgtacattta 60tattggctca tgtccaatat gaccgccatg ttgacattga ttattgacta
gttattaata 120gtaatcaatt acggggtcat tagttcatag cccatatatg
gagttccgcg ttacataact 180tacggtaaat ggcccgcctc gtgaccgccc
aacgaccccc gcccattgac gtcaataatg 240acgtatgttc ccatagtaac
gccaataggg actttccatt gacgtcaatg ggtggagtat 300ttacggtaaa
ctgcccactt ggcagtacat caagtgtatc atatgccaag tccgccccta
360ttgacgtcaa tgacggtaaa tggcccgcct ggcattatgc ccagtacatg
accttacggg 420actttcctac ttggcagtac atctacgtat tagtcatcgc
tattaccatg gtgatgcggt 480tttggcagta caccaatggg cgtggatagc
ggtttgactc acggggattt ccaagtctcc 540accccattga cgtcaatggg
agtttgtttt ggcaccaaaa tcaacgggac tttccaaaat 600gtcgtaataa
ccccgccccg ttgacgcaaa tgggcggtag gcgtgtacgg tgggaggtct
660atataagcag agctcgttta gtgaaccgtc agatcgcctg gagacgccat
ccacgctgtt 720ttgacctcca tagaagacac cgggaccgat ccagcctccg
cggccgggaa cggtgcattg 780gaacgcggat tccccgtgcc aagagtgacg
taagtaccgc ctatagactc tataggcaca 840cccctttggc tcttatgcat
gctatactgt ttttggcttg gggcctatac acccccgctc 900cttatgctat
aggtgatggt atagcttagc ctataggtgt gggttattga ccattattga
960ccactcccct attggtgacg atactttcca ttactaatcc ataacatggc
tctttgccac 1020aactatctct attggctata tgccaatact ctgtccttca
gagactgaca cggactctgt 1080atttttacag gatggggtcc catttattat
ttacaaattc acatatacaa caacgccgtc 1140ccccgtgccc gcagttttta
ttaaacatag cgtgggatct ccacgcgaat ctcgggtacg 1200tgttccggac
atgggctctt ctccggtagc ggcggagctt ccacatccga gccctggtcc
1260catgcctcca gcggctcatg gtcgctcggc agctccttgc tcctaacagt
ggaggccaga 1320cttaggcaca gcacaatgcc caccaccacc agtgtgccgc
acaaggccgt ggcggtaggg 1380tatgtgtctg aaaatgagct cggagattgg
gctcgcaccg tgacgcagat ggaagactta 1440aggcagcggc agaagaagat
gcaggcagct gagttgttgt attctgataa gagtcagagg 1500taactcccgt
tgcggtgctg ttaacggtgg agggcagtgt agtctgagca gtactcgttg
1560ctgccgcgcg cgccaccaga cataatagct gacagactaa cagactgttc
ctttccatgg 1620gtcttttctg cagtcaccgt ccaagcttag atctgccacc
atgagcctgc tgaccgaggt 1680ggagacatac gtgctgtcca tcatccccag
cggccctctg aaggccgaga tcgcccagag 1740actggaagat gtgttcgccg
gcaagaacac cgacctggaa gtgctgatgg aatggctgaa 1800aaccagaccc
atcctgagcc ctctgaccaa gggcatcctg ggcttcgtgt tcaccctgac
1860cgtgcccagc gagagaggcc tgcagaggcg gagattcgtg cagaacgccc
tgaacggcaa 1920cggcgacccc aacaacatgg acaaggccgt gaagctgtac
agaaagctga agcgggagat 1980caccttccac ggcgccaaag agatcagcct
gagctacagc gctggcgccc tggccagctg 2040catgggcctg atctacaaca
gaatgggcgc cgtgaccacc gaggtggcct tcggcctggt 2100ctgcgccacc
tgcgagcaga tcgccgacag ccagcacaga tcccacagac agatggtcac
2160caccaccaac cccctgatca gacacgagaa cagaatggtg ctggcctcta
ccaccgccaa 2220ggccatggaa cagatggccg gcagcagcga gcaggccgcc
gaggctatgg aagtcgcctc 2280tcaggctagg cagatggtcc aggccatgag
aaccatcggc acccacccca gcagctctgc 2340tggcctgaag aacgacctgc
tggaaaacct gcaggcctac cagaaaagaa tgggcgtcca 2400gatgcagaga
ttcaagtgat gagatatccc tcagagaggg gatcctcgca atccctaggg
2460ctgtgccttc tagttgccag ccaaactgtt gtttgcccct cccccgtgcc
ttccttgacc 2520ctggaaggtg ccactcccac tgtcctttcc taataaaatg
aggaaattgc atcgcattgt 2580ctgagtaggt gtcattctat tctggggggt
ggggtggggc aggacagcaa gggggaggat 2640tgggaagaca atagcaggca
tgctggggat gcggtgggct ctatgttcag aacgctcggt 2700tgccgccggg
cgttttttat ctagagtcga caaattcaga agaactcgtc aagaaggcga
2760tagaaggcga tgcgctgcga atcgggagcg gcgataccgt aaagcacgag
gaagcggtca 2820gcccattcgc cgccaagctc ttcagcaata tcacgggtag
ccaacgctat gtcctgatag 2880cggtccgcca cacccagccg gccacagtcg
atgaatccag aaaagcggcc attttccacc 2940atgatattcg gcaagcaggc
atcgccatgg gtcacgacga gatcctcgcc gtcgggcatg 3000ctcgccttga
gcctggcgaa cagttcggct ggcgcgagcc cctgatgctc ttcgtccaga
3060tcatcctgat cgacaagacc ggcttccatc cgagtacgtg ctcgctcgat
gcgatgtttc 3120gcttggtggt cgaatgggca ggtagccgga tcaagcgtat
gcagccgccg cattgcatca 3180gccatgatgg atactttctc ggcaggagca
aggtgagatg acaggagatc ctgccccggc 3240acttcgccca atagcagcca
gtcccttccc gcttcagtga caacgtcgag cacagctgcg 3300caaggaacgc
ccgtcgtggc cagccacgat agccgcgctg cctcgtcttg cagttcattc
3360agggcaccgg acaggtcggt cttgacaaaa agaaccgggc gcccctgcgc
tgacagccgg 3420aacacggcgg catcagagca gccgattgtc tgttgtgccc
agtcatagcc gaatagcctc 3480tccacccaag cggccggaga acctgcgtgc
aatccatctt gttcaatcat gcgaaacgat 3540cctcatcctg tctcttgatc
agatcttgat cccctgcgcc atcagatcct tggcggcgag 3600aaagccatcc
agtttacttt gcagggcttc ccaaccttac cagagggcgc cccagctggc
3660aattccggtt cgcttgctgt ccataaaacc gcccagtcta gctatcgcca
tgtaagccca 3720ctgcaagcta cctgctttct ctttgcgctt gcgttttccc
ttgtccagat agcccagtag 3780ctgacattca tccggggtca gcaccgtttc
tgcggactgg ctttctacgt gaaaaggatc 3840taggtgaaga tcctttttga
taatctcatg accaaaatcc cttaacgtga gttttcgttc 3900cactgagcgt
cagaccccgt agaaaagatc aaaggatctt cttgagatcc tttttttctg
3960cgcgtaatct gctgcttgca aacaaaaaaa ccaccgctac cagcggtggt
ttgtttgccg 4020gatcaagagc taccaactct ttttccgaag gtaactggct
tcagcagagc gcagatacca 4080aatactgtcc ttctagtgta gccgtagtta
ggccaccact tcaagaactc tgtagcaccg 4140cctacatacc tcgctctgct
aatcctgtta ccagtggctg ctgccagtgg cgataagtcg 4200tgtcttaccg
ggttggactc aagacgatag ttaccggata aggcgcagcg gtcgggctga
4260acggggggtt cgtgcacaca gcccagcttg gagcgaacga cctacaccga
actgagatac 4320ctacagcgtg agctatgaga aagcgccacg cttcccgaag
ggagaaaggc ggacaggtat 4380ccggtaagcg gcagggtcgg aacaggagag
cgcacgaggg agcttccagg gggaaacgcc 4440tggtatcttt atagtcctgt
cgggtttcgc cacctctgac ttgagcgtcg atttttgtga 4500tgctcgtcag
gggggcggag cctatggaaa aacgccagca acgcggcctt tttacggttc
4560ctgggctttt gctggccttt tgctcacatg ttgtcgac 4598109PRTArtificial
SequenceSynthetic peptide 10Ile Tyr Ser Thr Val Ala Ser Ser Leu 1 5
118PRTArtificial SequenceSynthetic peptide 11Ser Ile Ile Asn Phe
Glu Lys Leu 1 5
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013
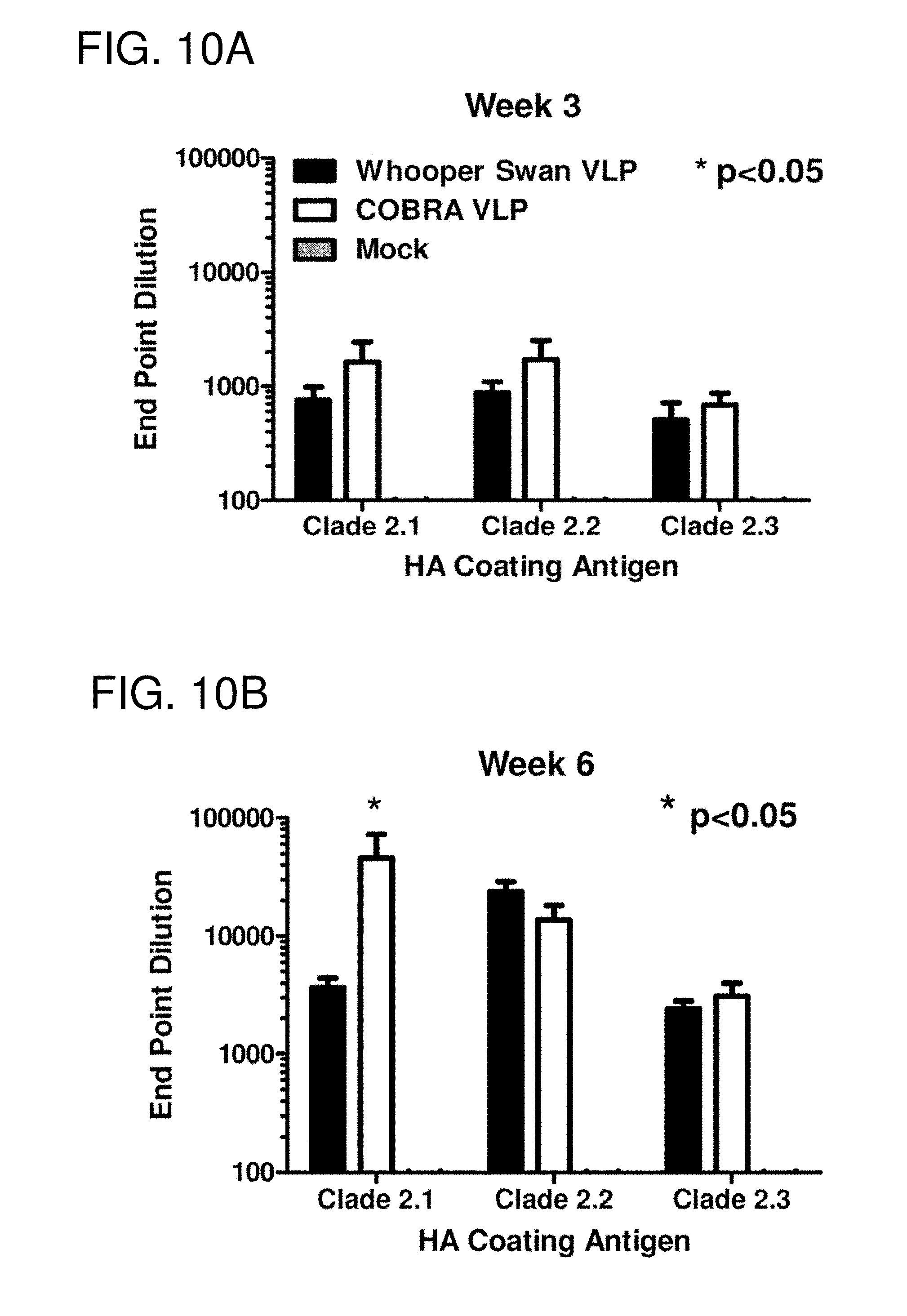
D00014

D00015
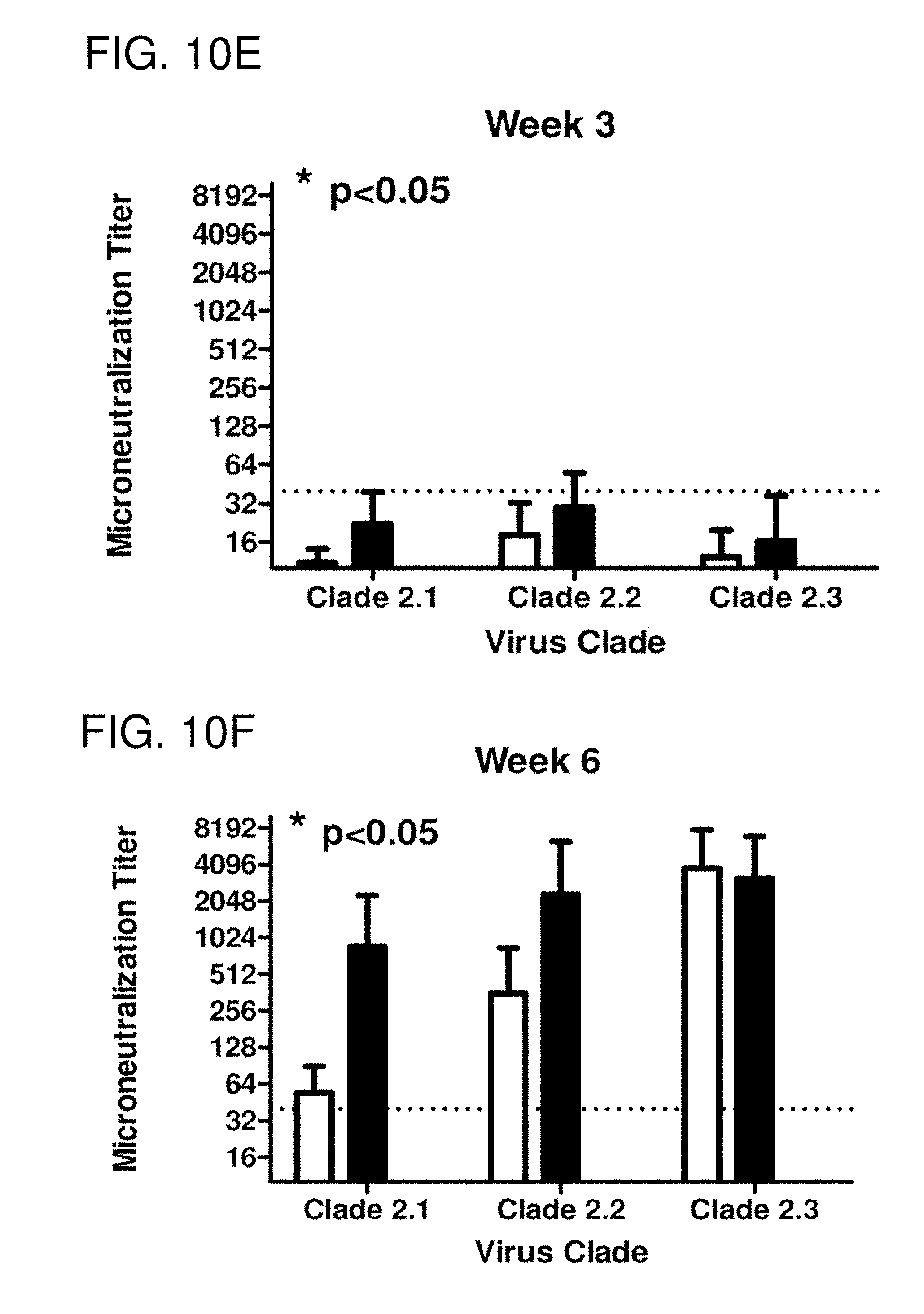
D00016
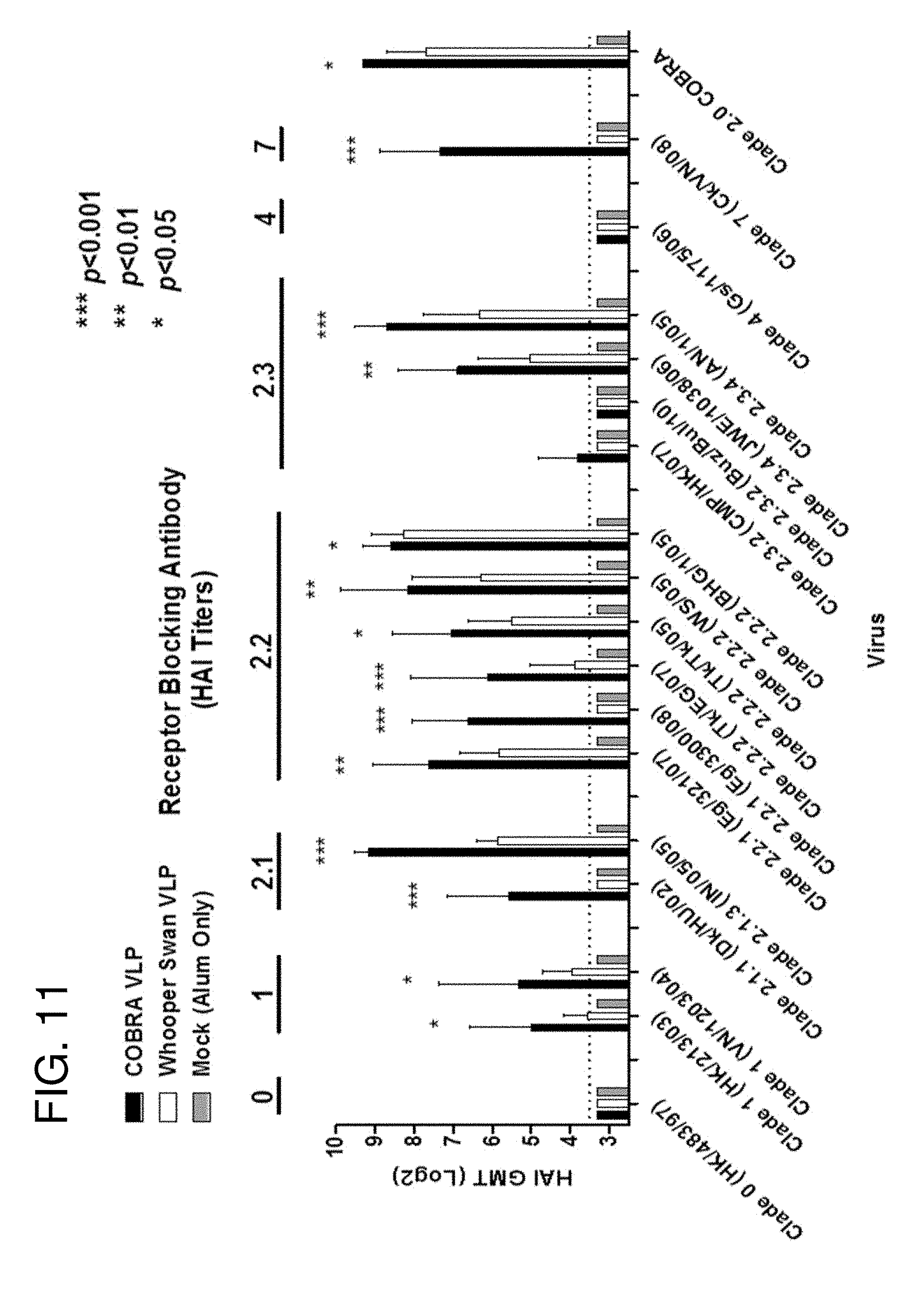
D00017

D00018

D00019
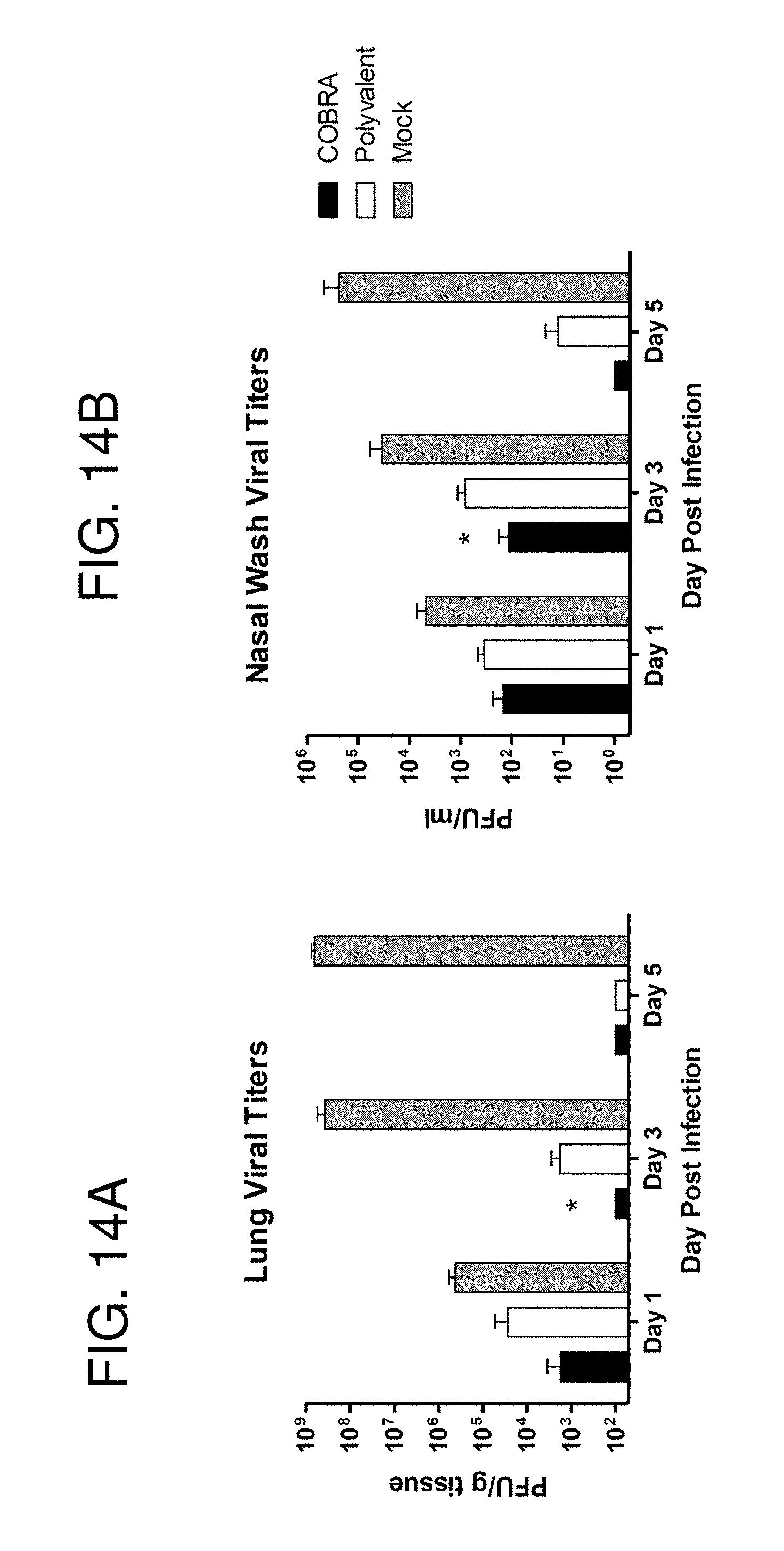
D00020

D00021
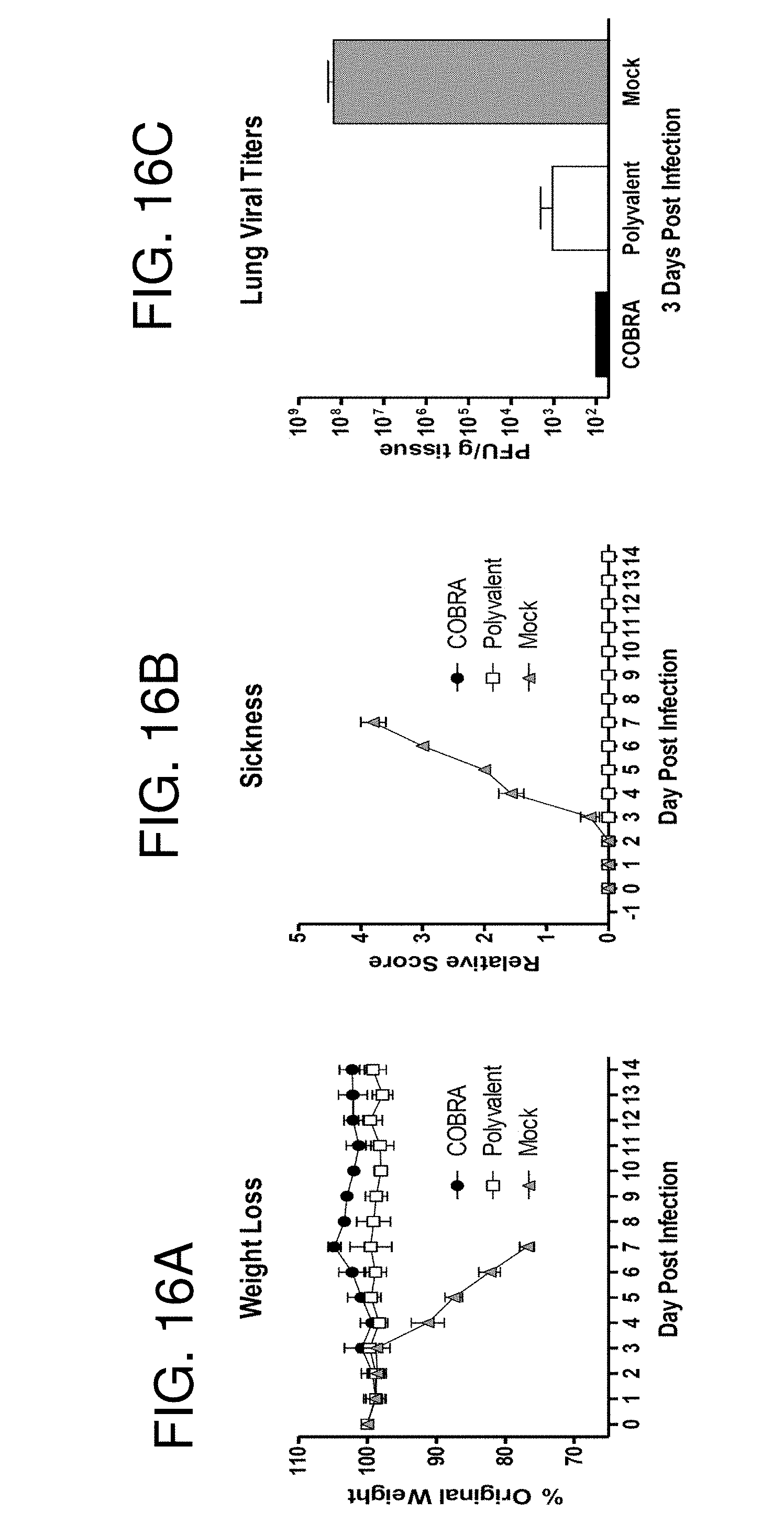
D00022
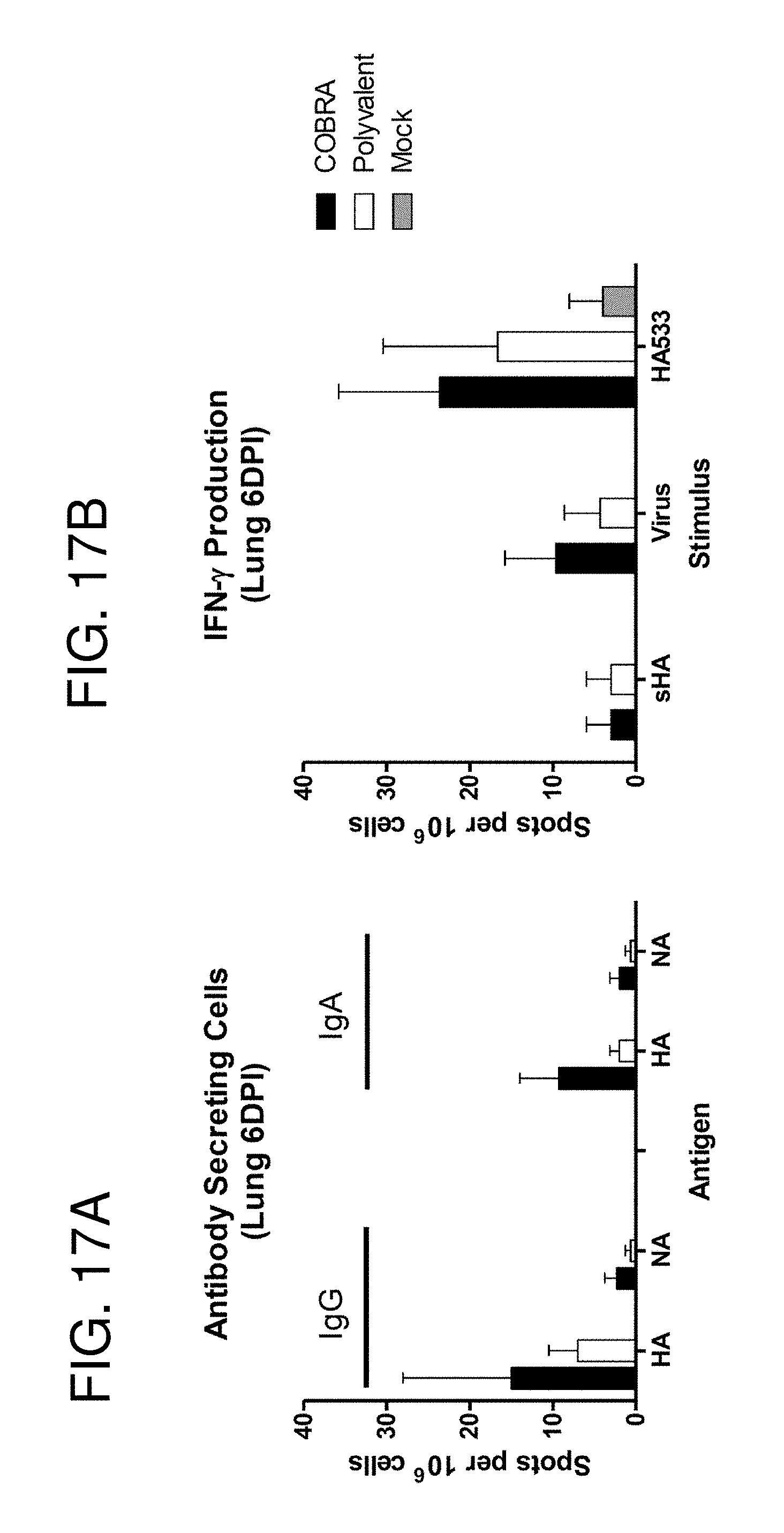
D00023
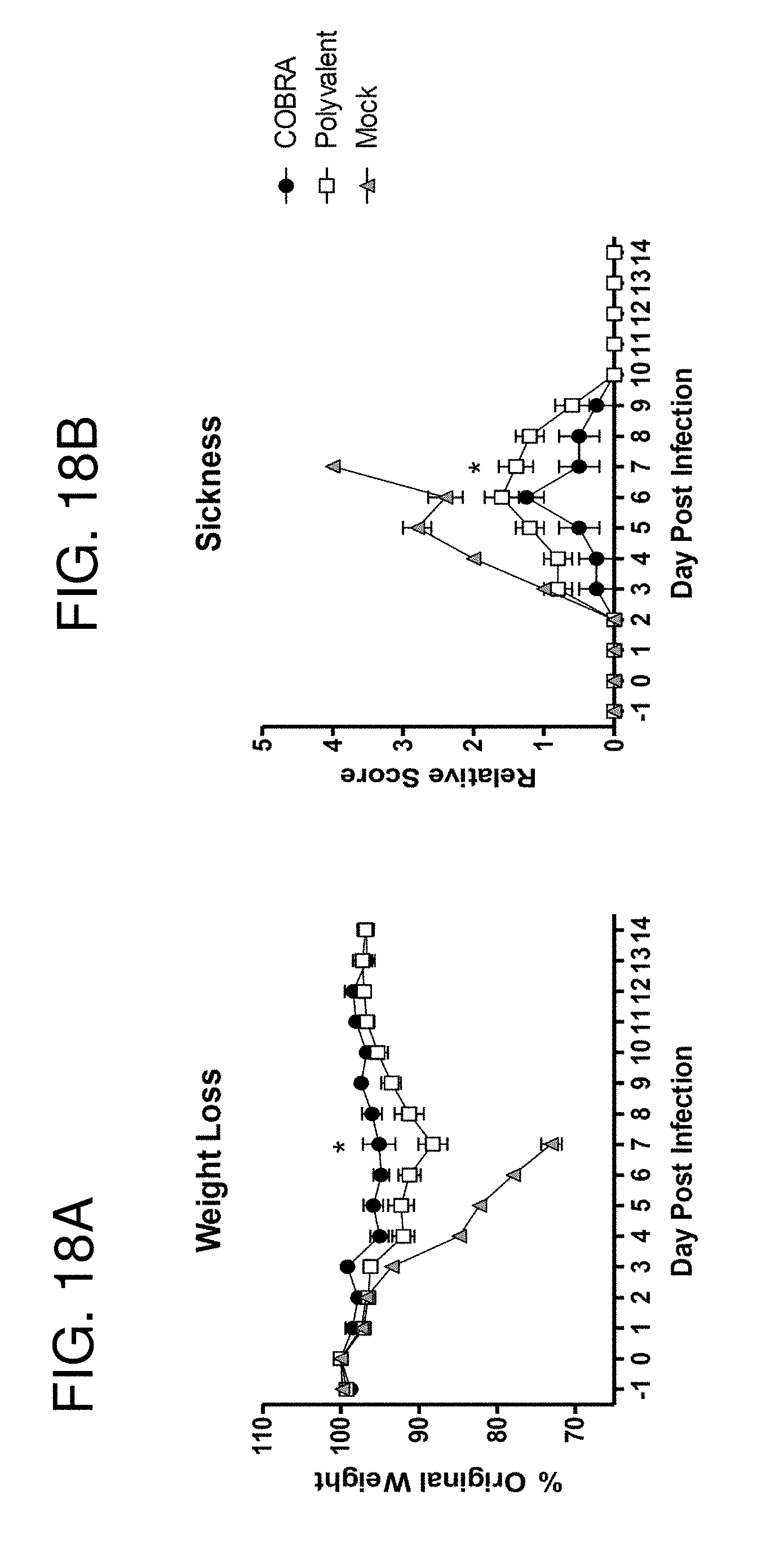
S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.