Isolated dsRNA Molecules And Methods Of Using Same For Silencing Target Molecules Of Interest
Avniel; Amir ; et al.
U.S. patent application number 16/024203 was filed with the patent office on 2019-01-03 for isolated dsrna molecules and methods of using same for silencing target molecules of interest. The applicant listed for this patent is A.B. Seeds Ltd.. Invention is credited to Amir Avniel, Efrat Lidor-Nili, Rudy Maor, Ofir Meir, Orly Noivirt-Brik, Osnat Yanai-Azulay.
| Application Number | 20190002878 16/024203 |
| Document ID | / |
| Family ID | 50069265 |
| Filed Date | 2019-01-03 |









View All Diagrams
| United States Patent Application | 20190002878 |
| Kind Code | A1 |
| Avniel; Amir ; et al. | January 3, 2019 |
Isolated dsRNA Molecules And Methods Of Using Same For Silencing Target Molecules Of Interest
Abstract
An isolated dsRNA molecule comprising an antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant, wherein the dsRNA sequence is flanked by two complementary sites to an smRNA or smRNAs expressed in the plant and wherein the dsRNA molecule further comprises a helicase binding site positioned so as to allow unwinding of the strands of the isolated dsRNA molecule to single stranded RNA (ssRNA) and recruitment of an RNA-dependent RNA polymerase so as to amplify the dsRNA molecule in the plant cell and generate secondary siRNA products of the dsRNA sequence.
| Inventors: | Avniel; Amir; (Tel-Aviv, IL) ; Lidor-Nili; Efrat; (Nes Ziona, IL) ; Maor; Rudy; (Rechovot, IL) ; Meir; Ofir; (Doar-Na Emek Soreq, IL) ; Noivirt-Brik; Orly; (Givataim, IL) ; Yanai-Azulay; Osnat; (Rishon-LeZion, IL) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 50069265 | ||||||||||
| Appl. No.: | 16/024203 | ||||||||||
| Filed: | June 29, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 14143748 | Dec 30, 2013 | 10041068 | ||
| 16024203 | ||||
| 61748095 | Jan 1, 2013 | |||
| 61748101 | Jan 1, 2013 | |||
| 61748094 | Jan 1, 2013 | |||
| 61748099 | Jan 1, 2013 | |||
| 61814888 | Apr 23, 2013 | |||
| 61814892 | Apr 23, 2013 | |||
| 61814899 | Apr 23, 2013 | |||
| 61814890 | Apr 23, 2013 | |||
| 61908965 | Nov 26, 2013 | |||
| 61908855 | Nov 26, 2013 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A01N 63/10 20200101; C12N 15/113 20130101; C12N 15/8206 20130101; C12N 15/8207 20130101; C12N 15/8218 20130101; C12N 15/8279 20130101; A01H 3/04 20130101 |
| International Class: | C12N 15/113 20060101 C12N015/113; C12N 15/82 20060101 C12N015/82; A01N 63/02 20060101 A01N063/02; A01H 3/04 20060101 A01H003/04 |
Claims
1. An isolated double-stranded RNA (dsRNA) molecule comprising (a) a first RNA strand having at least one antisense RNA sequence for suppressing expression of a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous smRNA-binding sequence for binding to a first small RNA (smRNA) expressed in said plant or phytopathogen, and a helicase-binding sequence comprising the helicase binding site of SEQ ID NO: 14; and (b) a second RNA strand that is a reverse complement of said first RNA strand.
2. The isolated dsRNA molecule of claim 1, wherein said first smRNA comprises a nucleic acid sequence wherein said nucleic acid sequence shares between 100% and 90% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NOs:1 to 288, and complements thereof.
3.-5. (canceled)
6. The isolated dsRNA molecule of claim 1, wherein said first RNA strand further comprises a second heterologous smRNA-binding sequence for binding a second smRNA expressed in said plant or phytopathogen, and said first heterologous smRNA-binding sequence and said second heterologous smRNA-binding sequence flank said at least one antisense RNA sequence.
7. The isolated dsRNA molecule of claim 6, wherein said first smRNA and said second smRNA comprise a nucleic acid sequence having at least 90% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NOs:1 to 288, and complements thereof.
8. The isolated dsRNA molecule of claim 6, wherein said second heterologous smRNA-binding sequence is the complement of said second smRNA.
9. The isolated dsRNA molecule of claim 6, wherein said second smRNA is identical to said first smRNA.
10. The isolated dsRNA molecule of claim 6, wherein said second smRNA is non-identical to said first smRNA.
11. The isolated dsRNA molecule of claim 6, wherein said first heterologous smRNA-binding sequence and said second heterologous smRNA-binding sequence comprise a nucleotide sequence selected from the group consisting of: a direct sequence of said first smRNA and a direct sequence of said second smRNA; a reverse complement of said first smRNA and a direct sequence of said second smRNA; a reverse complement of said first smRNA and a reverse complement of said second smRNA; a direct sequence of said first smRNA and a reverse complement of said second smRNA; a direct sequence of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a direct sequence of said second smRNA; a reverse complement of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a direct sequence of said second smRNA; a reverse complement of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a reverse complement of said second smRNA; a direct sequence of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a reverse complement of said second smRNA; a direct sequence of said first smRNA and a direct sequence of said second smRNA and further comprising a mutation rendering it resistant to cleavage; a reverse complement of said first smRNA and a direct sequence of said second smRNA and further comprising a mutation rendering it resistant to cleavage; a reverse complement of said first smRNA and a reverse complement of said second smRNA and further comprising a mutation rendering it resistant to cleavage; a direct sequence of said first smRNA and a reverse complement of said second smRNA and further comprising a mutation rendering it resistant to cleavage; a direct sequence of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a direct sequence of said second smRNA and further comprising a mutation rendering it resistant to cleavage; a reverse complement of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a direct sequence of said second smRNA and further comprising a mutation rendering it resistant to cleavage; a reverse complement of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a reverse complement of said second smRNA and further comprising a mutation rendering it resistant to cleavage; and a direct sequence of said first smRNA and further comprising a mutation rendering it resistant to cleavage and a reverse complement of said second smRNA and further comprising a mutation rendering it resistant to cleavage.
12.-14. (canceled)
15. The isolated dsRNA molecule of claim 1, wherein said first smRNA has a nucleotide sequence selected from the group consisting of an RNA sequence of a microRNA (miRNA) and an RNA sequence of an siRNA.
16. The isolated dsRNA molecule of claim 6, wherein said first smRNA has a nucleotide sequence selected from the group consisting of an RNA sequence of a miRNA and an RNA sequence of an siRNA, and said second smRNA has a nucleotide sequence selected from the group consisting of an RNA sequence of a miRNA and an RNA sequence of an siRNA.
17. (canceled)
18. The isolated dsRNA molecule of claim 1, wherein said first smRNA is a miRNA.
19. The isolated dsRNA molecule of claim 6, wherein said first smRNA is a miRNA and said second smRNA is a miRNA.
20. (canceled)
21. The isolated dsRNA molecule of claim 1, wherein said first smRNA is a miRNA selected from the group consisting of miR390, miR161.1, miR168, miR393, miR828, and miR173.
22. The isolated dsRNA molecule of claim 6, wherein said first smRNA is a miRNA selected from the group consisting of miR390, miR161.1, miR168, miR393, miR828, and miR173.
23.-48. (canceled)
49. A method of suppressing gene expression in a plant or a phytopathogen of the plant comprising: a. contacting a seed with an isolated double-stranded RNA (dsRNA) molecule of claim 1 under conditions which allow penetration of said dsRNA molecule into said seed, thereby introducing said dsRNA molecule into said seed; and optionally b. generating a plant of said seed.
50. The method of claim 49, wherein said dsRNA molecule penetrates a cell of said seed selected from the group consisting of an endosperm cell, an embryo cell, and combinations thereof.
51.-69. (canceled)
70. An isolated double-stranded RNA (dsRNA) molecule comprising: a first RNA strand having a nucleic acid sequence comprising in a sequential order from 5' to 3'; an endovirus 5' UTR sequence; an endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence; a multiple cloning site; an endovirus 3' UTR sequence; and a second RNA strand that is a reverse complement of said first RNA strand.
71. An isolated dsRNA molecule comprising: a first RNA strand having a nucleic acid sequence comprising in a sequential order from 5' to 3'; an endovirus 5' untranslated region (UTR) sequence; an endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence; an antisense nucleic acid sequence for regulating a target gene; an endovirus 3' UTR sequence; and a second RNA strand that is a reverse complement of said first RNA strand.
72. The isolated dsRNA molecule of claim 70, wherein said endovirus 5' UTR sequence, said endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence and said endovirus 3' UTR sequence are capable of autonomous replication when introduced into a plant cell.
73. The isolated dsRNA molecule of claim 71, wherein said endovirus 5' UTR sequence, said endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence and said endovirus 3' UTR sequence are capable of autonomous replication when introduced into a plant cell.
74.-113. (canceled)
Description
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of U.S. application Ser. No. 14/143,748, filed Dec. 30, 2013, which claims benefit of provisional applications 61/748,095, filed Jan. 1, 2013, 61/748,101 filed Jan. 1, 2013; 61/748,094 filed Jan. 1, 2013; 61/748,099, filed Jan. 1, 2013; 61/814,888, filed Apr. 23, 2013; 61/814,892, filed Apr. 23, 2013; 61/814,899, filed Apr. 23, 2013; 61/814,890, filed Apr. 23, 2013; 61/908,965, filed Nov. 26, 2013; and 61/908,855, filed Nov. 26, 2013, each of which is herein incorporated by reference.
SEQUENCE LISTING STATEMENT
[0002] The ASCII file, entitled P34097US03_SL.txt, created on Jun. 29, 2018, comprising 73,728 bytes, is incorporated herein by reference.
FIELD OF THE DISCLOSURE
[0003] The present disclosure provides for, and includes, methods and compositions for silencing target molecules of plants and plant pathogens. Also provided are plants, plant parts and seeds having dsRNAs and methods of introducing dsRNAs into seeds.
BACKGROUND
[0004] The present disclosure, in some embodiments thereof, relates to isolated dsRNA molecules and methods of using same for silencing target molecules of interest.
[0005] RNA interference (RNAi) has been shown effective in silencing gene expression in a broad variety of species, including plants, with wide ranging implications for cancer, inherited disease, infectious disease in plants and animals. Studies in a variety of organisms have shown that effectors of RNAi include dsRNA and related small interfering RNAs (siRNAs; also called "short interfering RNAs" and "silencing RNAs"). Studies have also shown in a variety of organisms that dsRNA or their siRNA derivatives can be used to arrest, retard or even prevent a variety of pathogens, most notably viral diseases (see, for example, PCT Patent Application Publication No. WO/2003/004649).
[0006] It has been shown in some species that RNAi mediated interference spreads from the initial site of dsRNA delivery, producing interference phenotypes throughout the injected animal. Recently the same spreading effect of dsRNA has been demonstrated in bee larva. In addition, homologs of transmembrane proteins called systemic RNA interference defective proteins (SID) have been detected in, for example, humans, mouse and C. elegans. It is thought that SID transmembrane channels are responsible for endocytic uptake and spreading effect of dsRNA (Aronstein et al., J. Apic Res and Bee World, 2006; 45:20-24; see also van Roessel P, Brand A H., "Spreading silence with Sid," Genome Biol. 5(2):208 (2004)).
[0007] Application of RNA interference technology for insects that are plant pests and other plant pests has been suggested. Moderate RNAi-type silencing of insect genes by feeding has been demonstrated (Turner et al., Insect Mol Biol 2006; 15:383; and Araujo et al., Insect Mol. Biol 2006; 36:683). Various publications have since then focused on the incorporation of dsRNA in plants as pesticides. Such incorporation methods can be divided into transgenic gene expression and coating such as a seed coating.
[0008] U.S. Pat. No. 6,326,193 refers to the use of recombinant insect viruses such as baculoviruses expressing dsRNA to silence selected insect genes for pest control. PCT Patent Application Publication No. WO 99/32619 describes the use of dsRNA for reducing crop destruction by plant pathogens or pests such as arachnids, insects, nematodes, protozoans, bacteria, or fungi. PCT Patent Application Publication No. WO 2004/005485 describes RNAi sequences and transgenic plants designed to control plant-parasitic nematodes.
[0009] U.S. Patent Application Publication No. 20030154508 describes pest control with a dsRNA against a cation-amino acid transporter/channel protein. PCT Patent Application Publication No. WO 02/14472 describes an inverted repeat and a sense or antisense nucleic acids for inhibiting target gene expression in a sucking insect. U.S. Patent Application Publication No. 20030150017 describes the use of RNA molecules homologous or complementary to a nucleotide sequence of a plant pest such as nematodes and insects.
[0010] Raemakers et al. (PCT Patent Application Publication Nos. WO 2007/080127 and WO 2007/080126) have disclosed transgenic plants expressing RNAi for controlling pest infestation by insects, nematodes, fungus and other plant pests. Among the sequences taught are sequences targeting essential genes of insects. Waterhouse et al. (U.S. Patent Application Publication No. 20060272049) and Van De Craen (U.S. Patent Application Publication No. 2010068172) also disclosed transgenic plants expressing dsRNA directed to essential genes of plant insect pests, for use as pesticides and insecticides. Boukharov et al. (U.S. Patent Application Publication No. 20070250947) disclosed dsRNA in transgenic plants for targeting plant parasitic nematodes.
[0011] U.S. Patent Application Publication No. 20080022423 describes the control of fungal and oomycete plant pathogens by inhibiting one or more biological functions. The disclosure provides methods and compositions for such control. By feeding one or more recombinant double stranded RNA molecules provided by the disclosure to the pathogen, a reduction in disease may be obtained through suppression of gene expression. The disclosure is also directed to methods for making transgenic plants that express the double stranded RNA molecules, and to particular combinations of transgenic agents for use in protecting plants from pathogen infection. Also described is a seed coating with the dsRNA anti-pathogenic compositions.
[0012] PCT Patent Application Publication No. WO 2011112570 describes a method of regulating target endogenous gene expression in growing plants/plant organs involving topically coating onto plants/organs, a composition comprising polynucleotide having sequence of specific contiguous nucleotides, and a transferring agent.
[0013] U.S. Pat. No. 8,143,480 refers to methods for knock-down of a target genes in plants, particularly efficient and specific methods for knock-down of a target gene in plants. This disclosure also relates to methods for silencing endogenous plant genes or plant pathogen genes. It further relates to nucleic acid constructs (DNA, RNA) which comprise a nucleic acid sequence that corresponds to a target gene or fragment thereof flanked by two complementary sites to an smRNA, e.g., a miRNA (one complementary site is on either side of the nucleic acid sequence), resulting in, for example the configuration: complementary site--nucleic acid sequence that corresponds to a target gene--complementary site. Axtell and Bartell describe siRNA biogenesis in Arabidopsis (Axtell and Bartel Cell. 2006 Nov. 3; 127(3):565-77.).
[0014] It has been reported that an autonomous dsRNA sequence derived from endovirus is found in every tissue of an infected plant and at every developmental stage. Thus, in 1993 Fukuhara et al. (Plant Mol. Biol. 21(6):1121-1130) identified a linear, 16 kb, dsRNA in symptomless Japonica rice that is not found in Indica rice. The dsRNA was detected in every tissue and at every developmental stage and its copy number was approximately constant (about 20 copies/cell). A sequence of about 13.2 kb of the dsRNA was determined and two open reading frames (ORFs) were found. The larger ORF (ORF B) was more than 12,351 nucleotides long and encoded a polypeptide of more than 4,117 amino acid residues having an RNA helicase-like domain followed by an RNA dependent RNA Polymerase-like domain, as characterized in subsequent works published as Fukuhara et al. 1995 J. Biol. Chem. 270(30):18147-18149; and Moriyama et al. 1995 Mol. Gen. Genet. 248(3):364-369.
[0015] While not limited by theory, during RNA silencing, RNAs of about 21 to 24 nucleotides (nt) in length are generated, which are incorporated into a protein complex where they serve as guide RNAs to direct the down-regulation of gene expression at the transcriptional or posttranscriptional level. These small interfering RNAs, small silencing RNAs, or short interfering RNAs are called "siRNAs" or "microRNAs", depending upon their biogenesis: endogenous siRNAs derive from long double-stranded RNA and miRNAs derive from local hairpin structures within longer transcripts.
[0016] RNA silencing occurs in plants, insects, nematodes and other animals. In addition, new compositions (e.g., nucleic acid constructs) and methods of achieving RNA-based silencing would be useful, and plants in which expression of one or more genes of interest is modulated, e.g., inhibited, would be of great use. New compositions and rapid cost-effective methods of achieving RNA-based silencing by directly manipulating the plant seed are highly desirable.
SUMMARY OF THE INVENTION
[0017] The present disclosure provides for, and includes, methods and compositions for the regulation of gene expression in plants.
[0018] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant and a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in a plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant) and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence.
[0019] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in the plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a helicase binding sequence and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence.
[0020] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant and a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in the plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant) and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence and the first heterologous RNA sequence.
[0021] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in a plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a helicase binding sequence and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence and the first heterologous RNA sequence.
[0022] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in said plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a helicase binding sequence and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence and the first heterologous RNA sequence.
[0023] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in said plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a helicase binding sequence and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence, the first heterologous RNA sequence, and helicase binding sequence.
[0024] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in the plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a second heterologous RNA sequence corresponding to a second smRNA expressed in the plant (e.g., a second heterologous smRNA-binding sequence for binding a second smRNA expressed in a plant), where the first heterologous smRNA and said second heterologous smRNA flank the at least one antisense RNA sequence, and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence.
[0025] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in the plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a second heterologous RNA sequence corresponding to a second smRNA expressed in the plant (e.g., a second heterologous smRNA-binding sequence for binding a second smRNA expressed in a plant), where the first heterologous smRNA and said second heterologous smRNA flank the at least one antisense RNA sequence, and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence, and first heterologous RNA sequence.
[0026] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in the plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a second heterologous RNA sequence corresponding to a second smRNA expressed in the plant (e.g., a second heterologous smRNA-binding sequence for binding a second smRNA expressed in a plant), where the first heterologous smRNA and said second heterologous smRNA flank the at least one antisense RNA sequence, and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence, first heterologous RNA sequence, and second heterologous RNA sequence.
[0027] The present disclosure provides for, and includes, isolated double-stranded RNA molecules having a first RNA strand of at least one antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of a plant, a helicase binding sequence, a first heterologous RNA sequence corresponding to a first small RNA (smRNA) expressed in the plant (e.g., a first heterologous smRNA-binding sequence for binding a first smRNA expressed in a plant), a second heterologous RNA sequence corresponding to a second smRNA expressed in the plant (e.g., a second heterologous smRNA-binding sequence for binding a second smRNA expressed in a plant), where the first heterologous smRNA and said second heterologous smRNA flank the at least one antisense RNA sequence, and a second RNA strand that is a reverse complement of the at least one antisense RNA sequence, first heterologous RNA sequence, and second heterologous RNA sequence.
[0028] According to some embodiments of the present disclosure there is provided an isolated dsRNA molecule comprising an antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant, wherein the dsRNA sequence is flanked by two complementary sites to an smRNA or smRNAs expressed in the plant and wherein the dsRNA molecule further comprises a helicase binding site positioned so as to allow unwinding of the strands of the isolated dsRNA molecule to single stranded RNA (ssRNA) and recruitment of an RNA-dependent RNA Polymerase so as to amplify the dsRNA molecule in the plant cell and generate secondary siRNA products of the dsRNA sequence.
[0029] According to some embodiments of the present disclosure there is provided an isolated dsRNA molecule comprising an antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant, wherein the dsRNA sequence is flanked by two complementary sites to an smRNA or smRNAs expressed in the plant.
[0030] According to an embodiment of some embodiments of the present disclosure there is provided an isolated dsRNA molecule comprising an antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant, wherein the dsRNA molecule further comprises a complementary site to an smRNA expressed in the plant located upstream or downstream the dsRNA.
[0031] According to some embodiments of the disclosure, the isolated dsRNA molecule further comprises a helicase binding site positioned so as to allow unwinding of the strands of the isolated dsRNA molecule to single stranded RNA (ssRNA) and recruitment of an RNA-dependent RNA Polymerase so as to amplify the dsRNA molecule in the plant cell.
[0032] According to some embodiments of the disclosure, the complementary site to the smRNA is located downstream of the dsRNA sequence.
[0033] According to some embodiments of the disclosure, the complementary site to the smRNA is located upstream of the dsRNA sequence.
[0034] According to some embodiments of the disclosure, one of the two complementary sites to the smRNA or smRNAs comprises a mutation rendering it resistant to cleavage by the complementary smRNA.
[0035] According to some embodiments of the disclosure, the helicase binding site is positioned upstream of the dsRNA sequence.
[0036] According to some embodiments of the disclosure, wherein the helicase binding site is position in the dsRNA sequence for regulating a target gene of interest in the plant or the phytopathogen of the plant.
[0037] According to some embodiments of the disclosure, the helicase binding site is positioned upstream of the dsRNA sequence and the two complementary sites to the smRNA or smRNAs flank the helicase binding site.
[0038] According to some embodiments of the disclosure, the smRNA or smRNAs is selected from the group consisting of a miRNA and a siRNA.
[0039] According to some embodiments of the disclosure, the smRNA or smRNAs is a miRNA.
[0040] According to some embodiments of the disclosure, the miRNA is smRNA390.
[0041] According to some embodiments of the disclosure, the plant comprises a TAS locus that has a second smRNA complementary site.
[0042] According to some embodiments of the disclosure, the first and second complementary sites are naturally found flanking the TAS locus in the plant.
[0043] According to some embodiments of the disclosure, the smRNA is an smRNA for which complementary sites are naturally found flanking a TAS locus in a plant.
[0044] According to some embodiments of the disclosure, the two complementary sites are complementary sites for the same smRNA.
[0045] According to some embodiments of the disclosure, the two complementary sites comprise difference sequences.
[0046] According to some embodiments of the disclosure, the two complementary sites comprise the same sequence.
[0047] According to some embodiments of the disclosure, the smRNAs are non-identical.
[0048] According to some embodiments of the disclosure, the smRNA or smRNAs is selected from the group consisting of miR390, miR161.1, miR168, miR393, miR828 and miR173. According to some embodiments of the disclosure, the plant is a crop plant.
[0049] According to an embodiment of some embodiments of the present disclosure there is provided a method of silencing expression of a target gene of interest in a plant, the method comprising introducing the isolated dsRNA molecule, and wherein the dsRNA sequence is for silencing the target gene of interest in the plant, thereby silencing expression of the target gene of interest in the plant.
[0050] According to an embodiment of some embodiments of the present disclosure there is provided a method of introducing dsRNA molecule into a seed, the method comprising contacting the seed with the isolated dsRNA molecule under conditions which allow penetration of the dsRNA molecule into the seed, thereby introducing the dsRNA molecule into the seed.
[0051] According to an embodiment of some embodiments of the present disclosure there is provided an isolated seed comprising the isolated dsRNA molecule.
[0052] According to some embodiments of the disclosure, the isolated seed is devoid of a heterologous promoter for driving expression of the dsRNA molecule in the plant.
[0053] According to an embodiment of some embodiments of the present disclosure there is provided a seed comprising the isolated dsRNA molecule and the secondary siRNA products.
[0054] According to an embodiment of some embodiments of the present disclosure there is provided a plant or plant part generated from the seed.
[0055] According to an embodiment of some embodiments of the present disclosure there is provided a seed containing device comprising a plurality of the seeds.
[0056] According to an embodiment of some embodiments of the present disclosure there is provided a sown field comprising a plurality of the seeds.
[0057] According to an embodiment of some embodiments of the present disclosure there is provided a method of producing a plant the method comprising: (a) providing the seed; and (b) germinating the seed so as to produce the plant.
[0058] According to an embodiment of some embodiments of the present disclosure there is provided a method of modulating gene expression in a plant, the method comprising: (a) contacting a seed of the plant with the dsRNA molecule, under conditions which allow penetration of the dsRNA molecule into the seed, thereby introducing the dsRNA molecule into the seed; and optionally (b) generating a plant of the seed.
[0059] According to some embodiments of the disclosure, the penetration is to an endosperm and alternatively or additionally an embryo of the seed.
[0060] According to an embodiment of some embodiments of the present disclosure there is provided a method of silencing expression of a target gene in a phytopathogenic organism, the method comprising providing to the phytopathogenic organism the plant or plant part, thereby silencing expression of a target gene in the phytopathogenic organism.
[0061] According to some embodiments of the disclosure, the phytopathogenic organism is selected from the group consisting of a fungus, a nematode, an insect, a bacteria and a virus.
[0062] According to an embodiment of some embodiments of the present disclosure there is provided a kit for introducing a dsRNA molecule to seeds comprising; (i) the dsRNA molecule; and (ii) a priming solution.
[0063] According to some embodiments of the disclosure, the dsRNA molecule and the priming solution are comprised in separate containers.
[0064] According to an embodiment of some embodiments of the present disclosure there is provided a pesticidal composition comprising the isolated dsRNA molecule.
[0065] According to some embodiments of the disclosure, the contacting is effected by inoculating the seed with the dsRNA molecule.
[0066] According to some embodiments of the disclosure, the method further comprises priming the seed prior to the contacting.
[0067] According to some embodiments of the disclosure, the priming is effected by: (i) washing the seed prior to the contacting; and (ii) drying the seed following step (i).
[0068] According to some embodiments of the disclosure, the washing is effected in the presence of double deionized water.
[0069] According to some embodiments of the disclosure, the washing is effected for 2-6 hours. According to some embodiments of the disclosure, the washing is effected at 4-28.degree. C. According to some embodiments of the disclosure, the drying is effected at 25-30.degree. C. for 10-16 hours.
[0070] According to some embodiments of the disclosure, the contacting is effected in a presence of the dsRNA molecule at a final concentration of 0.1-100 .mu.g/.mu.l.
[0071] According to some embodiments of the disclosure, the contacting is effected in a presence of the dsRNA molecule at a final concentration of 0.1-0.5 .mu.g/.mu.l.
[0072] According to some embodiments of the disclosure, the method further comprises treating the seed with an agent selected from the group consisting of a pesticide, a fungicide, an insecticide, a fertilizer, a coating agent and a coloring agent following the contacting.
[0073] According to some embodiments of the disclosure, the treating comprises coating the seed with the agent.
[0074] According to some embodiments of the disclosure, the conditions allow accumulation of the dsRNA molecule in the endosperm and alternatively or additionally embryo of the seed.
[0075] According to some embodiments of the disclosure, a concentration of the dsRNA molecule is adjusted according to a parameter selected from the group consisting of, seed size, seed weight, seed volume, seed surface area, seed density and seed permeability.
[0076] According to some embodiments of the disclosure, the contacting is effected prior to breaking of seed dormancy and embryo emergence.
[0077] According to some embodiments of the disclosure, the seed is a primed seed.
[0078] According to some embodiments of the disclosure, the seed comprises RNA dependent RNA polymerase activity for amplifying expression of the dsRNA molecule.
[0079] According to some embodiments of the disclosure, the seed is a hybrid seed.
[0080] According to an aspect of some embodiments of the present disclosure there is provided an isolated dsRNA molecule comprising a nucleic acid sequence which comprises in a sequential order from 5' to 3', an endovirus 5' UTR, an endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence, an endovirus 3' UTR and a multiple cloning site flanked by the RDRP and the 3' UTR.
[0081] According to an aspect of some embodiments of the present disclosure there is provided an isolated dsRNA molecule comprising a nucleic acid sequence which comprises in a sequential order from 5' to 3', an endovirus 5' UTR, an endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence, an endovirus 3' UTR and a nucleic acid sequence for regulating a target gene flanked by the RDRP and the 3' UTR.
[0082] According to some embodiments of the disclosure, the endovirus 5' UTR, endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence and the endovirus 3' UTR are selected capable of autonomous replication in the plant cell.
[0083] According to some embodiments of the disclosure, the 5' UTR is as set forth in SEQ ID NO: 14.
[0084] According to some embodiments of the disclosure, the 3' UTR is as set forth in SEQ ID NO: 22.
[0085] According to some embodiments of the disclosure, the endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence is as set forth in SEQ ID NO: 23.
[0086] According to some embodiments of the disclosure, the nucleic acid sequence for regulating a target gene is 17-600 bp long.
[0087] According to some embodiments of the disclosure, the nucleic acid sequence for regulating a target gene is selected from the group consisting of a miRNA and a siRNA.
[0088] According to some embodiments of the disclosure, the nucleic acid sequence for regulating a target gene is a miRNA.
[0089] Unless otherwise defined, all technical and/or scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which the disclosure pertains. Although methods and materials similar or equivalent to those described herein can be used in the practice or testing of embodiments of the disclosure, examples of methods and/or materials are described below. In case of conflict, the patent specification, including definitions, will control. In addition, the materials, methods, and examples are illustrative only and are not intended to be necessarily limiting.
BRIEF DESCRIPTION OF THE DRAWINGS
[0090] Some embodiments of the disclosure are herein described, by way of example only, with reference to the accompanying drawings. With specific reference now to the drawings in detail, it is stressed that the particulars shown are by way of example and for purposes of illustrative discussion of embodiments of the disclosure. In this regard, the description taken with the drawings makes apparent to those skilled in the art how embodiments of the disclosure may be practiced.
[0091] FIG. 1 presents fluorescent images of siGLO-treatment rice seeds over a 24 hour period according to embodiment of the present disclosure.
[0092] FIG. 2A presents a schematic representation of the Solanum Lycopersicum (Lycopersicon esculentum) TAS3 gene according to an embodiment of the present disclosure. Mir390BS is displayed in a darker gray box (SEQ ID NO: 319). The light gray box represents the 5' Mut Mir390BS (SEQ ID NO: 319).
[0093] FIG. 2B presents a schematic representation of dsRNA construct #1 having an exogenous trigger control according to an embodiment of the present disclosure. The construct includes a 234 bp exogenous sequence provided in Table 5 (e.g., Trigger #1).
[0094] FIG. 2C presents a schematic representation of dsRNA construct #2 having a dual Mir390BS sequence on the sense strand and an exogenous sequence according to an embodiment of the present disclosure. The construct is comprises 3 parts from 5' to 3': a 5' Mut Mir390BS sequence, a 234 bp exogenous sequence in reverse complement orientation, and a 3' Mir390BS. The sequences are presented in Table 5 (e.g., Trigger #2).
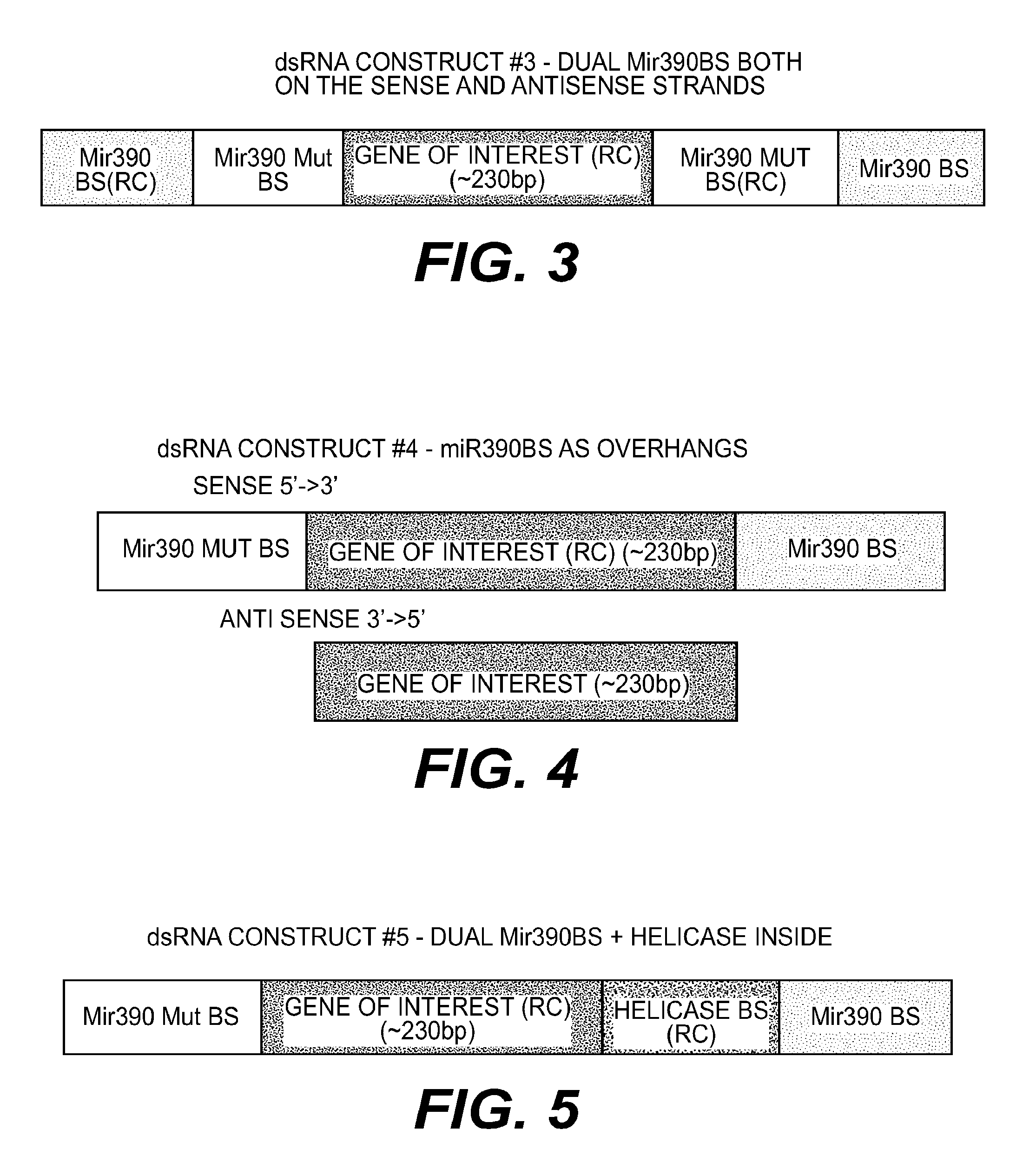
[0095] FIG. 3 presents a schematic representation of dsRNA construct#3 having a dual Mir390BS on both on the sense and antisense strands. The construct is composed of 5 parts from 5' to 3': 3' Mir390BS in the reverse complement orientation, 5' Mut Mir390BS, a 234 bp exogenous sequence in the reverse complement orientation, 5' Mut Mir390BS in the reverse complement orientation and 3' Mir390BS. For sequence, see Table 5 (Trigger #3).
[0096] FIG. 4 presents a schematic representation of dsRNA construct #4 having miR390S as overhangs. This construct is composed of two different strands. The sense strand is composed of 3 parts from 5' to 3': 5' Mut Mir390BS, a 234 bp exogenous sequence in the reverse complement orientation, 3' Mir390BS. The antisense is composed of only one part: a 234 bp exogenous sequence in the sense orientation. For sequences, see Table 5 (Sense-Trigger#4, Antisense-Trigger #5).
[0097] FIG. 5 presents a schematic representation of dsRNA construct #5 having miR390BS and a helicase binding sequence (Helicase BS). This construct is composed of 4 parts from 5' to 3': 5' Mut Mir390BS, a 234 bp exogenous sequence in the reverse complement orientation, Helicase BS in the reverse complement orientation, 3' Mir390BS. For sequence see Table 5 (Trigger #6).
[0098] FIG. 6 presents a schematic representation of dsRNA construct #6 having Mir390BS on both strands and Helicase BS as an overhang. This construct is composed of two different strands. The sense strand is composed of 5 parts from 5' to 3: `3` Mir390BS in the reverse complement orientation, 5' Mut Mir390BS, a 234 bp exogenous sequence in the reverse complement orientation, 5' Mut Mir390BS in the reverse complement orientation, and 3'Mir390BS. The antisense is composed of 6 parts from 5' to 3': 3'Mir390BS in the reverse complement orientation, 5' Mut Mir390BS, a 234 bp exogenous sequence in the sense orientation, 5' Mut Mir390BS in the reverse complement orientation, 3' Mir390BS and an Helicase BS as an overhang. For sequences, see Table 5 (Sense-Trigger #7, Antisense-Trigger #8).
[0099] FIG. 7 presents a schematic representation of dsRNA construct #7 having Sense dual Mir390BS coupled with Antisense Mir4376BS. This construct is composed of 5 parts from 5' to 3': 5' Mut Mir390BS, a 234 bp exogenous sequence in the reverse complement orientation, Mir4376BS in the reverse complement orientation and 3' Mir390BS. For sequence, see Table 5 (Trigger #9).
[0100] FIG. 8 presents a schematic representation of dsRNA construct #8 an Endogenous Trigger Control. This construct is composed of one part: a 234 bp of the endogenous TAS3 sequence. For sequence, see Table 5 (Trigger #10).
[0101] FIG. 9 presents a schematic representation of dsRNA construct #9--Mir390BS+Endogenous insert. This construct is composed of 3 parts from 5' to 3': 5' Mut Mir390BS, a 234 bp of the endogenous TAS3 sequence and 3' Mir390BS. For sequence, see Table 5 (Trigger #11).
[0102] FIGS. 10A-E are schematic representations of dsRNA constructs of the present disclosure.
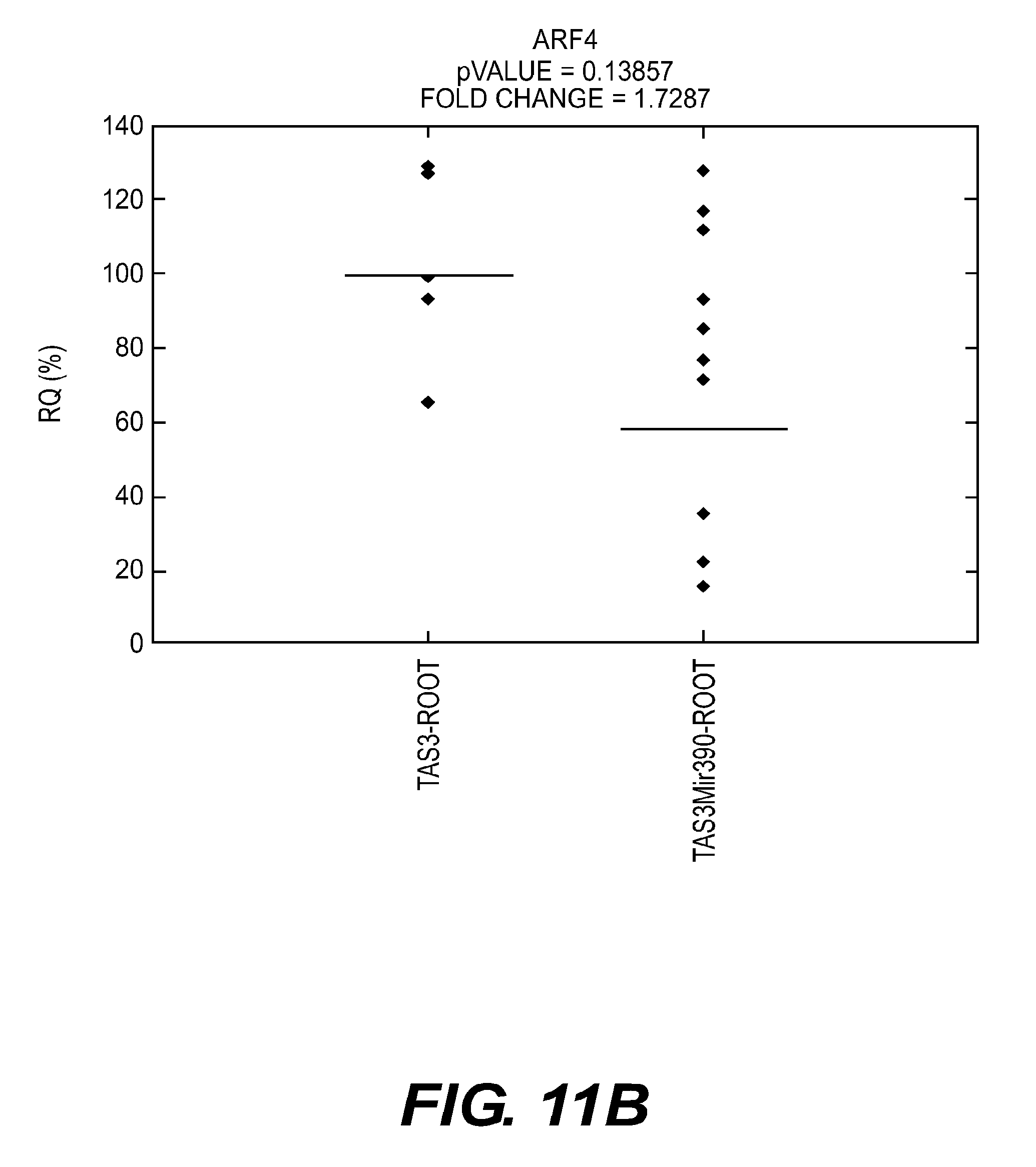
[0103] FIGS. 11A-B present graphs showing real-time PCR analyses of ARF3 and ARF4 mRNA expression in roots 14 days after seed treatment according to an embodiment of the present disclosure.
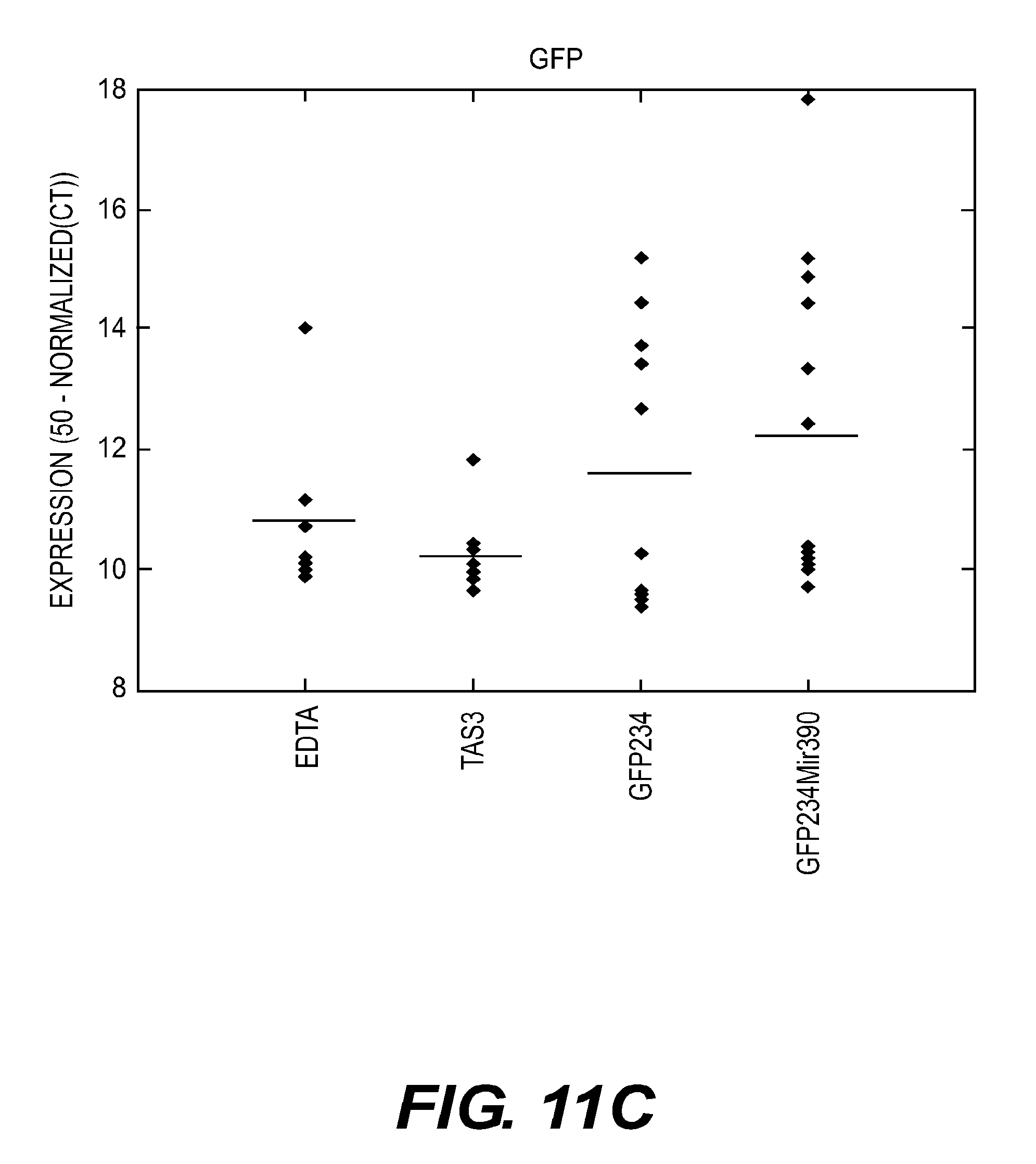
[0104] FIGS. 11C-E present graphs showing the results of real-time PCR analyses of GFP in seedlings seven days after seed treatment according to an embodiment of the present disclosure.
[0105] FIG. 11F presents a graph showing the results of real-time PCR analyses of GFP in leaves 30 days after seed treatment according to an embodiment of the present disclosure.
[0106] FIGS. 12A-E present graphs showing the results of real-time PCR analyses of GFP in shoots seven days after seed treatment according to an embodiment of the present disclosure.
[0107] FIGS. 13A-B presents graphs showing the results of real-time PCR analyses of GFP in shoots 14 days after seed treatment according to an embodiment of the present disclosure.
[0108] FIGS. 14A-B presents graphs showing the results of real-time PCR analyses of GFP in shoots seven days (A) and 14 days (B), according to an embodiment of the present disclosure.
[0109] FIGS. 15A-B presents graphs showing the results of real-time PCR analyses of GUS in shoots seven days (A) and 14 days (B) after seed treatment according to an embodiment of the present disclosure.
DETAILED DESCRIPTION
[0110] The present disclosure, in some embodiments thereof, relates to and provides for isolated dsRNA molecules and methods of using same for silencing target molecules of interest.
[0111] The present disclosure further includes and provides for compositions and methods for silencing gene expression.
[0112] The present disclosure provides for, and includes tools for overcoming the delivery obstacle and amplifying the small interfering RNA (siRNA) levels within the plant cell to thereby efficiently down-regulate target genes of interest.
[0113] Before explaining at least one embodiment of the disclosure in detail, it is to be understood that the disclosure is not necessarily limited in its application to the details set forth in the following description or exemplified by the Examples. The disclosure is capable of other embodiments or of being practiced or carried out in various ways.
[0114] It is understood that any Sequence Identification Number (SEQ ID NO) disclosed in the instant application can refer to either a DNA sequence or a RNA sequence, depending on the context where that SEQ ID NO is mentioned, even if that SEQ ID NO is expressed only in a DNA sequence format or a RNA sequence format. For example, SEQ ID NO: 1 is expressed in a DNA sequence format (e.g., reciting T for thymine), but it can refer to either a DNA sequence that corresponds to a T7 DNA Dependent RNA Polymerase primer nucleic acid sequence, or the RNA sequence of an RNA molecule nucleic acid sequence. Similarly, though SEQ ID NO: 25 is expressed in a RNA sequence format (e.g., reciting U for uracil), depending on the actual type of molecule being described, SEQ ID NO: 25 can refer to either the sequence of a RNA molecule comprising a dsRNA, or the sequence of a DNA molecule that corresponds to the RNA sequence shown. In any event, both DNA and RNA molecules having the sequences disclosed with any substitutes are envisioned.
[0115] As used herein, the terms "homology" and "identity" when used in relation to nucleic acids, describe the degree of similarity between two or more nucleotide sequences. The percentage of "sequence identity" between two sequences is determined by comparing two optimally aligned sequences over a comparison window, such that the portion of the sequence in the comparison window may comprise additions or deletions (gaps) as compared to the reference sequence (which does not comprise additions or deletions) for optimal alignment of the two sequences. The percentage is calculated by determining the number of positions at which the identical nucleic acid base or amino acid residue occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison, and multiplying the result by 100 to yield the percentage of sequence identity. A sequence that is identical at every position in comparison to a reference sequence is said to be identical to the reference sequence and vice-versa. An alignment of two or more sequences may be performed using any suitable computer program. For example, a widely used and accepted computer program for performing sequence alignments is CLUSTALW v1.6 (Thompson, et al. Nucl. Acids Res., 22: 4673-4680, 1994).
[0116] Homologous sequences include both orthologous and paralogous sequences. The term "paralogous" paralogous" relates to gene-duplications within the genome of a species leading to paralogous genes. The term "orthologous" relates to homologous genes in different organisms due to ancestral relationship. For instance in this case, other plant RNA viruses.
[0117] Identity (e.g., percent homology) can be determined using any homology comparison software, including for example, the BlastN software of the National Center of Biotechnology Information (NCBI) such as by using default parameters.
[0118] According to some embodiments of the disclosure, the identity is a global identity, i.e., an identity over the entire nucleic acid sequences of the disclosure and not over portions thereof.
[0119] The degree of homology or identity between two or more sequences can be determined using various known sequence comparison tools. Following is a non-limiting description of such tools which can be used along with some embodiments of the disclosure.
[0120] As used herein, the terms "exogenous polynucleotide" and "exogenous nucleic acid molecule" relative to an organism refer to a heterologous nucleic acid sequence which is not naturally expressed within that organism, for example a plant. An exogenous nucleic acid molecule may be introduced into an organism in a stable or transient manner. An exogenous nucleic acid molecule may comprise a nucleic acid sequence which is identical or partially homologous to an endogenous nucleic acid sequence of the organism. In certain embodiments, an "exogenous polynucleotide" and "exogenous nucleic acid molecule" may refer to a nucleic acid sequence expressed or present in a plant, either transiently or stably. As used herein, the terms "endogenous polynucleotide" and "endogenous nucleic acid" refers to nucleic acid sequences that are found in an organism's cell. In certain aspects, an endogenous nucleic acid may be part of the nuclear genome or the plastid genome. In other aspects, an endogenous nucleic acid may be found outside the nuclear or plastid genomes. As used herein, endogenous nucleic acids do not include viral, parasite or pathogen nucleic acids, for example an endovirus sequence. The present disclosure provides for, and includes, compositions comprising exogenous polynucleotides and exogenous nucleic acid molecules and methods for introducing them into a target organism. The present disclosure provides for, and includes, compositions comprising exogenous polynucleotides and exogenous nucleic acid molecules in combination with endogenous nucleic acids and polynucleotides and methods for introducing them into a target organism. The present disclosure provides for, and includes, compositions comprising recombinant endogenous nucleic acids and polynucleotides and methods for introducing them into a target organism.
[0121] The present disclosure provides for, and includes dsRNA molecules which are processed through the trans-acting siRNA (ta-siRNA) pathway. Transacting siRNAs are a subclass of siRNAs that function like miRNAs to repress expression of target genes. While not limited to any particular theory, trans-acting siRNAs form by transcription of ta-siRNA-generating genes found at trans-acting (TAS) loci. A ta-siRNA precursor is any nucleic acid molecule, including single-stranded or double-stranded DNA or RNA, that can be transcribed and/or processed to release a ta-siRNA. Cleavage of the primary transcript occurs through a guided RISC mechanism, conversion of one of the cleavage products to dsRNA, and processing of the dsRNA by dicer or dicer-like (DCL) enzymes. While not limited by any particular theory, it is thought that RNA-dependent RNA polymerase 6 (RDR6) (or related enzymes) function in posttranscriptional RNAi of sense transgenes, some viruses, and specific endogenous mRNAs that are targeted by trans-acting siRNAs (ta-siRNAs) (see Dalmay et al., Cell 101:543-553, 2000; Mourrain et al., Cell 101:533-542, 2000; Peragine et al., Genes & Dev 18:2369-2379, 2004; Vazquez et al., Mol Cell 16:69-79, 2004b; Yu et al., Mol Plant Microbe Interact 16:206-216, 2003). Again, while not being limited to any particular theory, it is thought that ta-siRNAs arise from transcripts that are recognized by RDR6, in cooperation with SGS3, as a substrate to form dsRNA. The dsRNA is processed accurately in 21-nucleotide steps by DCL1 to yield a set of "phased" ta-siRNAs. These ta-siRNAs interact with target mRNAs to guide cleavage by the same mechanism as do plant miRNAs (Peragine et al., Genes & Dev 18:2369-2379, 2004; Vazquez et al., Mol Cell 16:69-79, 2004; Allen et al., Cell 121:207-221, 2005). Trans-acting siRNAs are conserved among distantly related plant species and have been maintained over a long evolutionary period. The design and construction of ta-siRNA constructs and their use in the modulation of protein in transgenic plant cells is disclosed by Allen and Carrington in US Patent Application Publication US 2006/0174380 A1 (now U.S. Pat. No. 8,030,473) which is incorporated herein by reference.
[0122] As used herein, the term "dsRNA sequence" refers to, and includes, a double-stranded sequences having a first strand and a second strand that is a reverse complement of the first strand. It will be understood that reference to an antisense RNA sequence for regulating a target gene of interest and a sense RNA sequence for regulating a target gene of interest, would necessarily include a dsRNA sequences when included in a dsRNA molecule. For clarity, the sequences for targeting a gene of interest for regulation will be generally referenced as the antisense RNA sequence and provides for a standard reference point for the 5' and 3' ends. As used herein, the `antisense strand` refers to the strand having the antisense RNA sequence for regulating (e.g., suppressing or silencing) a target gene of interest. One of ordinary skill in the art would further understand that reference to a single strand, whether the sense or antisense strand, provides a definition and sequence for the reverse complement strand. Further, it is well understood that a single nucleic acid strand and its reverse complement provide for a double-stranded nucleic acid. One of ordinary skill in the art would understand that an RNA and DNA sequence may be readily substituted using the well-known base pairing rules and as provided above. One of ordinary skill in the art would further understand that binding can occur between two polynucleotide sequences that are characterized by having sufficient sequence complementarity (which need not be 100% complementarity) to allow hybridization between the two polynucleotides (e.g., binding or hybridization under common physiological conditions). Thus, a "heterologous smRNA-binding sequence for binding a first small RNA" need not be 100% complementary to the sequence of the first small RNA (for example, where the heterologous smRNA-binding sequence is complementary to the sequence of the first small RNA except for one or more mutations or mismatches at the site where cleavage mediated by the small RNA would normally occur), although in some embodiments the complementarity is 100%. The present disclosure provides for, and includes, an isolated dsRNA molecule comprising an antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant, wherein the dsRNA sequence is flanked by two complementary sites to an smRNA expressed in the plant. In some embodiments, the dsRNA sequence may be flanked by two complementary sites from the same smRNA expressed in the plant. In other embodiments, the dsRNA sequence may be flanked by complementary sites from two different smRNAs. In yet other embodiments, the dsRNA sequence may be flanked by four complementary sites corresponding to one or more smRNAs expressed in a plant (e.g., two heterologous sequences on one side and two heterologous sequences on the other side of the dsRNA sequence). In certain embodiments, the dsRNA molecule further comprises a helicase binding site positioned so as to allow unwinding of the strands of the isolated dsRNA molecule to single stranded RNA (ssRNA) and amplification by recruitment of an RNA-dependent RNA Polymerase (RDRP) when introduced into a host cell. In other embodiments, the helicase and other proteins may be provided in vitro, for example as part of a cell extract. Methods of in vitro analysis are known in the art. In certain embodiments, the host cell is a plant cell. In some embodiments, introduction of the dsRNA molecule into a plant cell results in the recruitment of a helicase and RDRP and the generation of secondary siRNA products corresponding to the dsRNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant. In certain embodiments, the target gene is silenced. In other embodiments, expression of the target gene is enhanced.
[0123] According to another embodiment of the disclosure there is provided, and included, an isolated dsRNA molecule comprising an antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant, wherein the dsRNA sequence is flanked by two complementary sites to an smRNA or smRNAs expressed in the plant. In certain embodiments, the target gene is silenced. In other embodiments, expression of the target gene is enhanced.
[0124] According to a further embodiment of the disclosure there is provided, and included, an isolated dsRNA molecule comprising an antisense RNA sequence for regulating a target gene of interest in a plant or a phytopathogen of the plant, wherein the dsRNA molecule further comprises a complementary site to an smRNA expressed in the plant located upstream or downstream the dsRNA. In certain embodiments, the target gene is silenced. In other embodiments, expression of the target gene is enhanced.
[0125] Not to be limited by theory, a possible downstream mechanism for a dsRNA construct of the present disclosure having two flanking heterologous RNA sequences corresponding to an smRNA, with one sequence being a non-cleavable mutant (for example, mir390 BS and Mir390 Mut BS of FIG. 2C) includes unwinding of the dsRNA in the cell. Following the opening of the double stranded RNA into two single strands the sense strand may recruit an AGO7, or AGO7-like, protein to the flanking heterologous RNA sequences. This binding, in turn, may lead to cleavage at a the non-mutated heterologous RNA sequence (for example Mir390 BS of FIG. 2C) and localization of this single stranded RNA inside a cytoplasmic processing center (Evidence for such a processing center was reported in Kumakura et al. (2009). SGS3 and RDR6 interact and colocalize in cytoplasmic SGS3/RDR6-bodies. (2009). FEBS Letters, 583, 1261-1266 and Jouannet et al. (2012). Cytoplasmic Arabidopsis AGO7 accumulates in membrane-associated siRNA bodies and is required for to-siRNA biogenesis. EMBO Journal, 31, 17041713.). The antisense strand may then either be diced or cleaved but is not expected to take part in additional amplification of the exogenous sequence (e.g., the gene of interest of FIG. 2C).
[0126] Not to be limited by theory, an alternative downstream mechanism for dsRNA construct of the present disclosure having two flanking heterologous RNA sequences corresponding to an smRNA (for example, as provided in FIG. 2C) similarly starts with the unwinding of the dsRNA in a cell. In this non-limiting theoretical mechanism, the sense strand is translocated to a processing center that may have an accumulation of a RNA Dependent RNA Polymerase (RDRP) that is predicted to lead to the formation of antisense transcripts. Preferably, each template of sense RNA will serve for multiple rounds of antisense RNA production. Following antisense RNA accumulation, it may be that the mere localization of this transcript inside the processing center enables RDRP recruitment and creation of double-stranded RNAs (even though this strand may lack a recognizable element of the TAS system). Some of these double stranded RNAs may be translocated to the nucleus where to be diced into ta-siRNAs against an exogenous sequence and some of the double stranded RNA may remain in the processing center where it will unwind again and lead to further cycles of amplification. One possible mediator of the unwinding process inside the processing center is the SDE3 RNA helicase (see Garcia et al. (2012). Ago Hook and RNA Helicase Motifs Underpin Dual Roles for SDE3 in Antiviral Defense and Silencing of Nonconserved Intergenic Regions. Mol Cell, 48, 109-120.).
[0127] Not to be limited by theory, a possible downstream mechanism for dsRNA construct #3 (FIG. 3) also begins with unwinding of the dsRNA into two single strands.
[0128] Focusing on the outcome of the sense strand, it may be recognized by Mir390-Ago7 at both Mir390 Binding sites. The binding of this complex may lead to cleavage at the 3' Mir390BS and to the translocation of this truncated transcript into a processing center. Inside the processing center it may serve as a template for the creation of multiple transcripts of antisense strands. The newly created antisense strands may contain recognizable Mir390 binding sites and therefore may be able to recruit Ago7 and Mir390 to the 5' Mut Mir390BS. This binding is may enable efficient recruitment of RDRP and creation of double stranded RNAs. Some of this double stranded RNA may be translocated to the nucleus and diced into ta-siRNAs whereas other dsRNA may be expected to continue to additional rounds of unwinding and amplification.
[0129] Not to be limited by theory, in the alternative, dsRNA construct #3 may be unwound in the cell to a ssRNA. Focusing on the outcome of the antisense strand, it may be recognized by Mir390-Ago7 at both Mir390 Binding sites. The binding of this complex may lead to cleavage at the 3' Mir390BS and to the translocation of this truncated transcript into the processing center. Inside the processing center it may serve as a template for the creation of multiple transcripts of sense strands. The newly created sense strands will contain recognizable Mir390 binding sites and therefore may be able to recruit Ago7 and Mir390 to the 5' Mut Mir390BS. This binding may enable efficient recruitment of RDRP and creation of double stranded RNAs. Some of this double stranded RNA may be translocated to the nucleus and diced into ta-siRNAs whereas other dsRNA may continue to additional rounds of unwinding and amplification.
[0130] Not to be limited by theory, in another alternative mechanism, dsRNA construct #3 is undergoes strand unwinding in a cell. Newly synthesized sense and antisense strands (possibly resulting from the mechanisms described above may serve as templates for multiple rounds of RDRP recruitment and dsRNA amplification. This construct may lead to an optimal amplification due to the presence of the 5' Mut Mir390BS on both strands enabling ongoing recruitment of Mir390-Ago7 complex.
[0131] As used herein, the term "upstream" refers to positions that are 5' end of the polynucleotide. In certain aspects, upstream refers to the 5' location of sequences relative to an antisense sequence for regulating a target gene.
[0132] As used herein the term "isolated" refers to separated from its natural environment. In the case of a dsRNA molecule, separated from the cytoplasm or the nucleus, conversely, in the case of a plant part such as a seed, separated from the rest of the plant.
[0133] As used herein the term "isolated dsRNA molecule" refers to an isolated RNA molecule which is substantially in a double stranded form. As used herein, an isolated dsRNA molecule may be in solution and may include buffers. An isolated dsRNA molecule is substantially separated from other nucleic acid molecules including DNA.
[0134] As used herein the term "dsRNA" refers to two strands of anti-parallel polyribonucleic acids held together by base pairing (e.g., two sequences that are the reverse complement of each other in the region of base pairing). The two strands can be of identical length or of different lengths provided there is enough sequence homology between the two strands that a double stranded structure is formed with at least 80%, 90%, 95% or 100% complementarity over the entire length. As used herein, the term "overhang" refers to non-double stranded regions of a dsRNA molecule (i.e., single stranded RNA). According to an embodiment of the disclosure, there are no overhangs for the dsRNA molecule. According to another embodiment of the disclosure, the dsRNA molecule comprises one overhang. According to other embodiments, a dsRNA molecule may comprise two overhangs.
[0135] In embodiments according to the present disclosure, an isolated dsRNA molecule comprises a second strand having an RNA sequence that is at least 80%, 90%, 95% or 100% complementary over its entire length to an antisense RNA sequence. In some embodiments, an isolated dsRNA molecule comprises a second strand that is 99% complementary over its entire length to an antisense RNA sequence. In other embodiments, the double stranded region is 98% complementary over the entire length of an antisense RNA sequence. In yet other embodiments, the double stranded region is 97% complementary over the entire length of an antisense RNA sequence. In further embodiments, the double stranded region may comprise 96% of the entire length of an antisense RNA sequences. In certain embodiments the double stranded region is between 90 and 100% complementary over the entire length of antisense RNA sequence. In certain embodiments the double stranded region is between 95 and 100% complementary over the entire length of antisense RNA sequence.
[0136] The present disclosure provides for, and includes, embodiments of an isolated dsRNA molecule comprising a second strand having an RNA sequence that is nearly 100% complementary over its entire length to an antisense RNA sequence but having 1 mismatch. In some embodiments, the nearly 100% complementary dsRNA region may have 2 mismatches. In some embodiments, the nearly 100% complementary dsRNA region may have 3 mismatches. Some embodiments according to the present disclosure provide for 4, 5 or 6 mismatches in a dsRNA region. In some embodiments, the nearly 100% complementary dsRNA region may have 1 or more, 2 or more, or 3 or more mismatches.
[0137] According to an embodiment, an overhang may be 5' to a double stranded region comprising at least one antisense RNA sequence and its reverse complement (e.g., 5' to said antisense RNA sequence). According to an embodiment, an overhang may be 3' to a double stranded region comprising at least one antisense RNA sequence and its reverse complement (e.g., 3' to said antisense RNA sequence). In other embodiments according to the present disclosure, a dsRNA molecule may comprise two overhang regions flanking a double stranded region.
[0138] According to other embodiments, an overhang region comprises less than 10 bases. In certain embodiments, the strands are aligned such that there are at least 1, 2, or 3 bases at the end of the strands which do not align (i.e., for which no complementary bases occur in the opposing strand) such that an overhang of 1, 2 or 3 residues occurs at one or both ends of the duplex when strands are annealed. In an embodiment, a less than 10 base overhang may be a 5' overhang (relative to the 5' and 3' positions on the end of a double stranded RNA region). In another embodiment, a less than 10 base overhang may be a 3' overhang. Relative to a dsRNA molecule having at least one antisense RNA sequence, the 5' overhang may be located at 5' of said antisense RNA sequences. In other embodiments, the 5' overhang may be located 3' of said antisense RNA sequence (e.g., the 5' overhang is on the complementary strand). Also provided by the present disclosure are embodiments wherein the 3' overhang is located 3' of said antisense RNA sequence or wherein the 3' overhang is located 5' of said antisense RNA sequence. According to embodiments of the present disclosure, a 5' overhanging sequence may be 9 bases. In an embodiment, a 3' overhanging sequence may be 9 bases. According to embodiments of the present disclosure, a 5' overhanging sequence may be 8 bases. In an embodiment, a 3' overhanging sequence may be 8 bases. According to embodiments of the present disclosure, a 5' overhanging sequence may be 7 bases. In an embodiment, a 3' overhanging sequence may be 7 bases. According to embodiments of the present disclosure, a 5' overhanging sequence may be 6 bases. In an embodiment, a 3' overhanging sequence may be 6 bases. In some embodiments, a single stranded overhanging sequence may be less than 5 bases. According to embodiments of the present disclosure, a 5' overhanging sequence may be 5 bases. In an embodiment, a 3' overhanging sequence may be 5 bases. According to embodiments of the present disclosure, a 5' overhanging sequence may be 4 bases. In an embodiment, a 3' overhanging sequence may be 4 bases. According to embodiments of the present disclosure, a 5' overhanging sequence may be 3 bases. In an embodiment, a 3' overhanging sequence may be 3 bases. According to embodiments of the present disclosure, a 5' overhanging sequence may be 2 bases. In an embodiment, a 3' overhanging sequence may be 2 bases.
[0139] As will be appreciated by one of ordinary skill in the art, a dsRNA molecule of the present disclosure may refer to either strand of the anti-parallel nucleic acids. As will also be appreciated by one of ordinary skill in the art, a dsRNA molecule of the present disclosure includes both a `sense` and `antisense` strand and that the sense and antisense strands are reverse complements of each other in a region of base pairing. As used herein the sequence of a dsRNA molecule for regulating a target gene of interest is provided as the `antisense` orientation with respect to the target gene of interest. Thus, one of ordinary skill in the art would appreciate that the 5' end of a dsRNA molecule for regulating a target gene of interest corresponds to sequences towards the 3' end of the target gene of interest. Similarly, the 3' end of a dsRNA molecule for regulating a target gene of interest corresponds to sequences towards the 5' end of a target gene of interest. As used herein, "the reverse complement of a dsRNA molecule for regulating a target gene of interest" refers to a nucleic acid sequence in the `sense` orientation.
[0140] The term "corresponding to the target gene of interest" or "dsRNA for regulating a target gene of interest" means that the dsRNA sequence contains an RNA silencing agent to the target gene.
[0141] As used herein, the term "RNA silencing agent" refers to a nucleic acid which is capable of inhibiting or "silencing" the expression of a target gene. In certain aspects, the RNA silencing agent is capable of preventing complete processing (e.g., the full translation and/or expression) of an mRNA molecule through a post-transcriptional silencing mechanism. RNA silencing agents can be single- or double-stranded RNA or single- or double-stranded DNA or double-stranded DNA/RNA hybrids or modified analogues thereof. In some aspects, the RNA silencing agents are selected from the group consisting of (a) a single-stranded RNA molecule (ssRNA), (b) a ssRNA molecule that self-hybridizes to form a double-stranded RNA molecule, (c) a double-stranded RNA molecule (dsRNA), (d) a single-stranded DNA molecule (ssDNA), (e) a ssDNA molecule that self-hybridizes to form a double-stranded DNA molecule, and (f) a single-stranded DNA molecule including a modified Pol III gene that is transcribed to an RNA molecule, (g) a double-stranded DNA molecule (dsDNA), (h) a double-stranded DNA molecule including a modified Pol III promoter that is transcribed to an RNA molecule, (i) a double-stranded, hybridized RNA/DNA molecule, or combinations thereof. In some aspects these polynucleotides include chemically modified nucleotides or non-canonical nucleotides. In some aspects, the RNA silencing agents are noncoding RNA molecules, for example RNA duplexes comprising paired strands, as well as precursor RNAs from which such small non-coding RNAs can be generated. In some aspects, the RNA silencing agents are dsRNAs such as siRNAs, miRNAs and shRNAs. In one aspect, the RNA silencing agent is capable of inducing RNA interference. In another aspect, the RNA silencing agent is capable of mediating translational repression. As used herein, an RNA silencing agent is a type of agent for regulating a target gene.
[0142] In some embodiments, the dsRNA molecule is subject to amplification by RNA-Dependent RNA Polymerase (RDRP). According to some embodiments, a dsRNA molecule comprises a first strand having at least one antisense RNA sequence for regulating a target gene, one or two heterologous RNA sequences corresponding to a smRNA, a helicase binding site and a sequence encoding an RDRP, and a second complementary strand. According to some embodiments, a dsRNA molecule comprises a first strand having at least one antisense RNA sequence for regulating a target gene, one or two heterologous RNA sequences corresponding to a smRNA, a helicase binding site and a sequence encoding an RDRP and further including flanking 3' UTR and 5' UTR sequences from an endovirus and a second RNA strand that is the reverse complement.
[0143] As used herein, "small RNA" or "smRNA" refers to RNA molecules that function to modulate (e.g., inhibit), gene expression, and are present in diverse eukaryotic organisms, including plants. As known to those of skill in the art, smRNAs may be defined as low-molecular weight RNAs associated with gene silencing and in some embodiments may be further described as short (generally 21 to 26 nucleotides). Small RNAs include siRNAs and miRNAs, which function in RNA silencing, also sometimes referred to as RNA interference (RNAi). RNA silencing encompasses a broad range of phenomena in which large, double-stranded RNA, fold-back structures, or stem-loop precursors are processed to about 21-26 nucleotide (nt) small RNAs (e.g., siRNAs or miRNAs, which are described further below) that then guide the cleavage of cognate RNAs, block productive translation thereof, or induce methylation of specific target DNAs (Meins, F., et al., Annu Rev. Cell Dev. Biol., 21:297-318, 2005).
[0144] As used herein, a small RNA is an RNA molecule that is at least 15 base pairs in length, generally 15-30 nucleotides long, preferably 20-24 nucleotides long. In some aspects, In aspects according to the present disclosure, a "small RNA" is greater than 30 base pairs in length. In an aspect, the small RNA is greater than 30 base pairs in length but less than about 600 base pairs. In an aspect, the small RNA is greater than 100 base pairs in length but less than about 600 base pairs. In an aspect, the small RNA is greater than 200 base pairs in length but less than about 600 base pairs. A small RNA can be either double-stranded or single-stranded. Small RNA includes, without limitation, miRNA (microRNA), ta-siRNA (trans activating siRNA), siRNA, activating RNA (RNAa), nat-siRNA (natural anti-sense siRNA), he-siRNA (heterochromatic siRNA), cis-acting siRNA, lmiRNA (long miRNA), lsiRNA (long siRNA) and easiRNA (epigenetically activated siRNA) and their respective precursors. Preferred siRNA molecules of the disclosure are miRNA molecules, to-siRNA molecules and RNAa molecules and their respective precursors. A small RNA may be processed in vivo by an organism to an active form. According to aspects of the present disclosure, a selective insecticide may be a small RNA. In embodiments according to the present disclosure a small RNA is a dsRNA.
[0145] As provided for and included in the present disclosure, a dsRNA molecule may comprise an antisense RNA sequence for regulating a target gene of interest. In some embodiments, a dsRNA molecule for regulating a target gene of interest may comprise an antisense RNA sequence that is greater than 30 base pairs in length to allow processing of the dsRNA in a plant cell and generation of secondary siRNA molecules. In other embodiments, a dsRNA molecule for regulating a target gene of interest may comprise an antisense RNA sequence that is from 30 to 600 bp in length to allow processing of the dsRNA in a plant cell and generation of secondary siRNA molecules. As used herein, "secondary siRNA", "phase RNA" and "ta-siRNA" or refer to dsRNA molecules generated after processing a dsRNA molecule. In certain embodiments, the target gene regulation is silencing. In other embodiments, expression of the target gene is enhanced.
[0146] The present disclosure also includes and provides for embodiments having dsRNA molecules having various lengths of dsRNA sequences, whereby the shorter version i.e., x is shorter or equals 50 bp (e.g., 17-50), is referred to as siRNA or miRNA sequences. Longer dsRNA sequences of 51-600 nucleotides are referred to herein as dsRNA, which can be further processed for siRNA molecules.
[0147] The term "siRNA" generally refers to small inhibitory RNA duplexes (generally between 17-30 base pairs, but also longer e.g., 31-50 bp) that induce the RNA interference (RNAi) pathway. In certain embodiments, siRNAs are chemically synthesized as 2 lmers with a central 19 bp duplex region and symmetric 2-base 3'-overhangs on the termini, although it has been recently described that chemically synthesized RNA duplexes of 25-30 base length can have as much as a 100-fold increase in potency compared with 21mers at the same location. Without being limited by any theory, a role of siRNA is its involvement in the RNA interference (RNAi) pathway, where it interferes with the expression of a specific gene. Though not to be limiting, the observed increased potency obtained using longer RNAs in triggering RNAi is theorized to result from providing Dicer with a substrate (27mer) instead of a product (21mer) and that this improves the rate or efficiency of entry of the siRNA duplex into the RNA-induced silencing complex (RISC).
[0148] It has been found that position of the 3'-overhang influences potency of an siRNA and asymmetric duplexes having a 3'-overhang on the antisense strand are generally more potent than those with the 3'-overhang on the sense strand (Rose et al., 2005). This can be attributed to asymmetrical strand loading into RISC, as the opposite efficacy patterns are observed when targeting the antisense transcript.
[0149] In some embodiments, the strands of a double-stranded interfering RNA (e.g., an siRNA) may be connected to form a hairpin or stem-loop structure (e.g., an shRNA). Thus, as mentioned the RNA silencing agent of some embodiments of the disclosure may also be a short hairpin RNA (shRNA).
[0150] The term "shRNA", as used herein, refers to an RNA agent having a stem-loop structure, comprising a first and second region of complementary sequence, the degree of complementarity and orientation of the regions being sufficient such that base pairing occurs between the regions, the first and second regions being joined by a loop region, the loop resulting from a lack of base pairing between nucleotides (or nucleotide analogs) within the loop region. The number of nucleotides in the loop is a number between and including 3 to 23, 5 to 15, 7 to 13, 4 to 9, or 9 to 11. Some of the nucleotides in the loop can be involved in base-pair interactions with other nucleotides in the loop. Examples of oligonucleotide sequences that can be used to form the loop include 5'-UUCAAGAGA-3' (Brummelkamp, T. R. et al. (2002) Science 296: 550) and 5'-UUUGUGUAG-3' (Castanotto, D. et al. (2002) RNA 8:1454). It will be recognized by one of skill in the art that the resulting single chain oligonucleotide forms a stem-loop or hairpin structure comprising a double-stranded region capable of interacting with the RNAi machinery.
[0151] As used herein, the phrase "microRNA (also referred to herein interchangeably as "miRNA" or "miR") or a precursor thereof" refers to a microRNA (miRNA) molecule acting as a post-transcriptional regulator. Typically, the miRNA molecules are RNA molecules of about 20 to 22 nucleotides in length which can be loaded into a RISC complex and which direct the cleavage of another RNA molecule, wherein the other RNA molecule comprises a nucleotide sequence essentially complementary to the nucleotide sequence of the miRNA molecule.
[0152] While not limited by a particular theory, a miRNA molecule is often processed from a "pre-miRNA" or as used herein a precursor of a miRNA molecule by proteins, such as DCL proteins. Pre-microRNA molecules are typically processed from pri-microRNA molecules (primary transcripts). The single-stranded RNA segments flanking the pre-microRNA are important for processing of the pri-miRNA into the pre-miRNA. The cleavage site appears to be determined by the distance from the stem-ssRNA junction (Han et al., 2006, Cell, 125:887-901). In some embodiments, a miRNA molecule is loaded onto a RISC complex where it can guide the cleavage of the target gene of interest.
[0153] Pre-microRNA molecules are typically processed from pri-microRNA molecules (primary transcripts). The single stranded RNA segments flanking the pre-microRNA are important for processing of the pri-miRNA into the pre-miRNA. The cleavage site appears to be determined by the distance from the stem-ssRNA junction (Han et al. 2006, Cell 125, 887-901, 887-901).
[0154] As used herein, a "pre-miRNA" molecule is an RNA molecule of about 100 to about 200 nucleotides, preferably about 100 to about 130 nucleotides which can adopt a secondary structure comprising an imperfect double stranded RNA stem and a single stranded RNA loop (also referred to as "hairpin") and further comprising the nucleotide sequence of the miRNA (and its complement sequence) in the double stranded RNA stem. According to a specific embodiment, the miRNA and its complement are located about 10 to about 20 nucleotides from the free ends of the miRNA double stranded RNA stem. The length and sequence of the single stranded loop region are not critical and may vary considerably, e.g. between 30 and 50 nucleotides in length. The complementarity between the miRNA and its complement need not be perfect and about 1 to 3 bulges of unpaired nucleotides can be tolerated. The secondary structure adopted by an RNA molecule can be predicted by computer algorithms conventional in the art such as mFOLD. The particular strand of the double stranded RNA stem from the pre-miRNA which is released by DCL activity and loaded onto the RISC complex is determined by the degree of complementarity at the 5' end, whereby the strand which at its 5' end is the least involved in hydrogen bounding between the nucleotides of the different strands of the cleaved dsRNA stem is loaded onto the RISC complex and will determine the sequence specificity of the target RNA molecule degradation. However, if empirically the miRNA molecule from a particular synthetic pre-miRNA molecule is not functional (because the "wrong" strand is loaded on the RISC complex), it will be immediately evident that this problem can be solved by exchanging the position of the miRNA molecule and its complement on the respective strands of the dsRNA stem of the pre-miRNA molecule. As is known in the art, binding between A and U involving two hydrogen bounds, or G and U involving two hydrogen bounds is less strong that between G and C involving three hydrogen bounds.
[0155] In some embodiments according to the present disclosure, naturally occurring miRNA molecules may be comprised within their naturally occurring pre-miRNA molecules. In other embodiments, a miRNA can be introduced into a non-natural heterologous pre-miRNA molecule scaffold by exchanging the nucleotide sequence of the miRNA molecule. Thus, when processed the recombinant pre-miRNA produces an miRNA having a replaced sequence. In some embodiments, the scaffold of the pre-miRNA can also be completely synthetic. Likewise, synthetic miRNA molecules may be comprised within, and processed from, existing pre-miRNA molecule scaffolds or synthetic pre-miRNA scaffolds. Some pre-miRNA scaffolds may be preferred over others for their efficiency to be correctly processed into the designed microRNAs, particularly when expressed as a chimeric gene. In some aspects a chimeric pre-miRNA gene may include other DNA regions, such as untranslated leader sequences, transcription termination and polyadenylation regions that are incorporated in the primary transcript in addition to the pre-microRNA.
[0156] According to the present teachings, the dsRNA sequences may be naturally occurring or synthetic.
[0157] The dsRNA sequence for regulating a target gene of interest may contain multiple discrete portions or regions that correspond to the target gene, separated by portions that do not correspond to the target gene. In some embodiments, portions that do not correspond to the target gene may optionally correspond to a second, third, or fourth target gene. It will be appreciated that the portions that correspond to the target gene may have different lengths, different degrees of sequence identity to the target gene, and may correspond to regions located anywhere within the target gene.
[0158] In embodiments according to the present disclosure, an antisense RNA sequence may be flanked by two nucleic acid sequences that are complementary to an smRNA expressed in the plant. In some embodiments, the two flanking nucleic acid sequences may be complementary to two different smRNAs. In yet other embodiments, the two flanking nucleic acid sequences may comprise sequences that are complementary to more than two smRNAs. In further embodiments, the smRNAs may comprise two copies of one smRNA and a nucleic acid sequence complementary to a different smRNA.
[0159] The present disclosure provides for an includes dsRNA molecules comprising an antisense RNA sequence and one or two nucleic acid sequences that are complementary to an smRNA expressed in the plant. The present disclosure provides for embodiments having the composition and orientation of one or two nucleic acid sequences that are complementary to an smRNA expressed in the plant of dsRNA molecules as shown in Table 1 below. In certain embodiments, the dsRNA molecules of Table 1 further comprise a helicase binding sequence. In some embodiments, the dsRNA molecules of Table 1, further comprise a helicase binding sequence and a RDRP polypeptide encoding sequence.
TABLE-US-00001 TABLE 1 Embodiments of first strands of dsRNA molecules having a first and second nucleic acid sequence complementary to an smRNA expressed in a plant 5' 5' Antisense RNA 3' 3' Construct smRNA.sub.2 smRNA.sub.1 sequence smRNA.sub.1 smRNA.sub.2 A01.sup.1 None None Present None None A02 None Direct Present Direct None A03 None Direct Present Direct Mut None A04 None Direct Present R/C None A05 None Direct Present R/C Mut None A06 None Direct Mut Present Direct None A07 None Direct Mut Present Direct Mut None A08 None Direct Mut Present R/C None A09 None Direct Mut Present R/C Mut None A10 None R/C Present Direct None A11 None R/C Present Direct Mut None A12 None R/C Present R/C None A13 None R/C Present R/C Mut None A14 None R/C Mut Present Direct None A15 None R/C Mut Present Direct Mut None A16 None R/C Mut Present R/C None A17 None R/C Mut Present R/C Mut None B01 Direct Direct Present Direct Direct B02 Direct Direct Present Direct Direct Mut B03 Direct Direct Present Direct R/C B04 Direct Direct Present Direct R/C Mut B05 Direct Direct Present Direct Mut Direct B06 Direct Direct Present Direct Mut Direct Mut B07 Direct Direct Present Direct Mut R/C B08 Direct Direct Present Direct Mut R/C Mut B09 Direct Direct Present R/C Direct B10 Direct Direct Present R/C Direct Mut B11 Direct Direct Present R/C R/C B12 Direct Direct Present R/C R/C Mut B13 Direct Direct Present R/C Mut Direct B14 Direct Direct Present R/C Mut Direct Mut B15 Direct Direct Present R/C Mut R/C B16 Direct Direct Present R/C Mut R/C Mut B17 Direct Direct Mut Present Direct Direct B18 Direct Direct Mut Present Direct Direct Mut B19 Direct Direct Mut Present Direct R/C B20 Direct Direct Mut Present Direct R/C Mut B21 Direct Direct Mut Present Direct Mut Direct B22 Direct Direct Mut Present Direct Mut Direct Mut B23 Direct Direct Mut Present Direct Mut R/C B24 Direct Direct Mut Present Direct Mut R/C Mut B25 Direct Direct Mut Present R/C Direct B26 Direct Direct Mut Present R/C Direct Mut B27 Direct Direct Mut Present R/C R/C B28 Direct Direct Mut Present R/C R/C Mut B29 Direct Direct Mut Present R/C Mut Direct B30 Direct Direct Mut Present R/C Mut Direct Mut B31 Direct Direct Mut Present R/C Mut R/C B32 Direct Direct Mut Present R/C Mut R/C Mut B33 Direct R/C Present Direct Direct B34 Direct R/C Present Direct Direct Mut B35 Direct R/C Present Direct R/C B36 Direct R/C Present Direct R/C Mut B37 Direct R/C Present R/C Direct B38 Direct R/C Present R/C Direct Mut B39 Direct R/C Present R/C R/C B40 Direct R/C Present R/C R/C Mut B41 Direct R/C Present R/C Mut Direct B42 Direct R/C Present R/C Mut Direct Mut B43 Direct R/C Present R/C Mut R/C B44 Direct R/C Present R/C Mut R/C Mut B45 Direct R/C Mut Present Direct Direct B46 Direct R/C Mut Present Direct Direct Mut B47 Direct R/C Mut Present Direct R/C B48 Direct R/C Mut Present Direct R/C Mut B49 Direct R/C Mut Present Direct Mut Direct B50 Direct R/C Mut Present Direct Mut Direct Mut B51 Direct R/C Mut Present Direct Mut R/C B52 Direct R/C Mut Present Direct Mut R/C Mut B53 Direct R/C Mut Present R/C Direct B54 Direct R/C Mut Present R/C Direct Mut B55 Direct R/C Mut Present R/C R/C B56 Direct R/C Mut Present R/C R/C Mut B57 Direct R/C Mut Present R/C Mut Direct B58 Direct R/C Mut Present R/C Mut Direct Mut B59 Direct R/C Mut Present R/C Mut R/C B60 Direct R/C Mut Present R/C Mut R/C Mut C01 Direct Mut Direct Present Direct Direct C02 Direct Mut Direct Present Direct Direct Mut C03 Direct Mut Direct Present Direct R/C C04 Direct Mut Direct Present Direct R/C Mut C05 Direct Mut Direct Present Direct Mut Direct C06 Direct Mut Direct Present Direct Mut Direct Mut C07 Direct Mut Direct Present Direct Mut R/C C08 Direct Mut Direct Present Direct Mut R/C Mut C09 Direct Mut Direct Present R/C Direct C10 Direct Mut Direct Present R/C Direct Mut C11 Direct Mut Direct Present R/C R/C C12 Direct Mut Direct Present R/C R/C Mut C13 Direct Mut Direct Present R/C Mut Direct C14 Direct Mut Direct Present R/C Mut Direct Mut C15 Direct Mut Direct Present R/C Mut R/C C16 Direct Mut Direct Present R/C Mut R/C Mut C17 Direct Mut Direct Mut Present Direct Direct C18 Direct Mut Direct Mut Present Direct Direct Mut C19 Direct Mut Direct Mut Present Direct R/C C20 Direct Mut Direct Mut Present Direct R/C Mut C21 Direct Mut Direct Mut Present Direct Mut Direct C22 Direct Mut Direct Mut Present Direct Mut Direct Mut C23 Direct Mut Direct Mut Present Direct Mut R/C C24 Direct Mut Direct Mut Present Direct Mut R/C Mut C25 Direct Mut Direct Mut Present R/C Direct C26 Direct Mut Direct Mut Present R/C Direct Mut C27 Direct Mut Direct Mut Present R/C R/C C28 Direct Mut Direct Mut Present R/C R/C Mut C29 Direct Mut Direct Mut Present R/C Mut Direct C30 Direct Mut Direct Mut Present R/C Mut Direct Mut C31 Direct Mut Direct Mut Present R/C Mut R/C C32 Direct Mut Direct Mut Present R/C Mut R/C Mut C33 Direct Mut R/C Present Direct Direct C34 Direct Mut R/C Present Direct Direct Mut C35 Direct Mut R/C Present Direct R/C C36 Direct Mut R/C Present Direct R/C Mut C37 Direct Mut R/C Present R/C Direct C38 Direct Mut R/C Present R/C Direct Mut C39 Direct Mut R/C Present R/C R/C C40 Direct Mut R/C Present R/C R/C Mut C41 Direct Mut R/C Present R/C Mut Direct C42 Direct Mut R/C Present R/C Mut Direct Mut C43 Direct Mut R/C Present R/C Mut R/C C44 Direct Mut R/C Present R/C Mut R/C Mut C45 Direct Mut R/C Mut Present Direct Direct C46 Direct Mut R/C Mut Present Direct Direct Mut C47 Direct Mut R/C Mut Present Direct R/C C48 Direct Mut R/C Mut Present Direct R/C Mut C49 Direct Mut R/C Mut Present Direct Mut Direct C50 Direct Mut R/C Mut Present Direct Mut Direct Mut C51 Direct Mut R/C Mut Present Direct Mut R/C C52 Direct Mut R/C Mut Present Direct Mut R/C Mut C53 Direct Mut R/C Mut Present R/C Direct C54 Direct Mut R/C Mut Present R/C Direct Mut C55 Direct Mut R/C Mut Present R/C R/C C56 Direct Mut R/C Mut Present R/C R/C Mut C57 Direct Mut R/C Mut Present R/C Mut Direct C58 Direct Mut R/C Mut Present R/C Mut Direct Mut C59 Direct Mut R/C Mut Present R/C Mut R/C C60 Direct Mut R/C Mut Present R/C Mut R/C Mut D01 R/C Direct Present Direct Direct D02 R/C Direct Present Direct Direct Mut D03 R/C Direct Present Direct R/C D04 R/C Direct Present Direct R/C Mut D05 R/C Direct Present Direct Mut Direct D07 R/C Direct Present Direct Mut R/C D08 R/C Direct Present Direct Mut R/C Mut D10 R/C Direct Present R/C Direct Mut D11 R/C Direct Present R/C R/C D12 R/C Direct Present R/C R/C Mut D13 R/C Direct Present R/C Mut Direct D14 R/C Direct Present R/C Mut Direct Mut D15 R/C Direct Present R/C Mut R/C D16 R/C Direct Present R/C Mut R/C Mut D17 R/C Direct Mut Present Direct Direct D18 R/C Direct Mut Present Direct Direct Mut D19 R/C Direct Mut Present Direct R/C D20 R/C Direct Mut Present Direct R/C Mut D21 R/C Direct Mut Present Direct Mut Direct D22 R/C Direct Mut Present Direct Mut Direct Mut D23 R/C Direct Mut Present Direct Mut R/C D24 R/C Direct Mut Present Direct Mut R/C Mut D25 R/C Direct Mut Present R/C Direct D26 R/C Direct Mut Present R/C Direct Mut D27 R/C Direct Mut Present R/C R/C D28 R/C Direct Mut Present R/C R/C Mut D29.sup.3 R/C Direct Mut Present R/C Mut Direct D30 R/C Direct Mut Present R/C Mut Direct Mut D31 R/C Direct Mut Present R/C Mut R/C D32 R/C Direct Mut Present R/C Mut R/C Mut D33 R/C R/C Present Direct Direct D34 R/C R/C Present Direct Direct Mut D35 R/C R/C Present Direct R/C D36 R/C R/C Present Direct R/C Mut D37 R/C R/C Present R/C Direct D38 R/C R/C Present R/C Direct Mut D39 R/C R/C Present R/C R/C D40 R/C R/C Present R/C R/C Mut D41 R/C R/C Present R/C Mut Direct D42 R/C R/C Present R/C Mut Direct Mut D43 R/C R/C Present R/C Mut R/C D44 R/C R/C Present R/C Mut R/C Mut D45 R/C R/C Mut Present Direct Direct D46 R/C R/C Mut Present Direct Direct Mut D47 R/C R/C Mut Present Direct R/C D48 R/C R/C Mut Present Direct R/C Mut D49 R/C R/C Mut Present Direct Mut Direct D50 R/C R/C Mut Present Direct Mut Direct Mut D51 R/C R/C Mut Present Direct Mut R/C D52 R/C R/C Mut Present Direct Mut R/C Mut D53 R/C R/C Mut Present R/C Direct D54 R/C R/C Mut Present R/C Direct Mut D55 R/C R/C Mut Present R/C R/C D56 R/C R/C Mut Present R/C R/C Mut D57 R/C R/C Mut Present R/C Mut Direct D58 R/C R/C Mut Present R/C Mut Direct Mut D59 R/C R/C Mut Present R/C Mut R/C D6 R/C Direct Present Direct Mut Direct Mut D60 R/C R/C Mut Present R/C Mut R/C Mut D9 R/C Direct Present R/C Direct E01 R/C Mut Direct Present Direct Direct E02 R/C Mut Direct Present Direct Direct Mut E03 R/C Mut Direct Present Direct R/C E04 R/C Mut Direct Present Direct R/C Mut E05 R/C Mut Direct Present Direct Mut Direct E06 R/C Mut Direct Present Direct Mut Direct Mut E07 R/C Mut Direct Present Direct Mut R/C E08 R/C Mut Direct Present Direct Mut R/C Mut E09 R/C Mut Direct Present R/C Direct E10 R/C Mut Direct Present R/C Direct Mut E11 R/C Mut Direct Present R/C R/C E12 R/C Mut Direct Present R/C R/C Mut E13 R/C Mut Direct Present R/C Mut Direct E14 R/C Mut Direct Present R/C Mut Direct Mut E15 R/C Mut Direct Present R/C Mut R/C E16 R/C Mut Direct Present R/C Mut R/C Mut E17 R/C Mut Direct Mut Present Direct Direct E18 R/C Mut Direct Mut Present Direct Direct Mut E19 R/C Mut Direct Mut Present Direct R/C E20 R/C Mut Direct Mut Present Direct R/C Mut E21 R/C Mut Direct Mut Present Direct Mut Direct E22 R/C Mut Direct Mut Present Direct Mut Direct Mut E23 R/C Mut Direct Mut Present Direct Mut R/C E24 R/C Mut Direct Mut Present Direct Mut R/C Mut E25 R/C Mut Direct Mut Present R/C Direct E26 R/C Mut Direct Mut Present R/C Direct Mut E27 R/C Mut Direct Mut Present R/C R/C E28 R/C Mut Direct Mut Present R/C R/C Mut E29 R/C Mut Direct Mut Present R/C Mut Direct E30 R/C Mut Direct Mut Present R/C Mut Direct Mut E31 R/C Mut Direct Mut Present R/C Mut R/C E32 R/C Mut Direct Mut Present R/C Mut R/C Mut E33 R/C Mut R/C Present Direct Direct E34 R/C Mut R/C Present Direct Direct Mut E35 R/C Mut R/C Present Direct R/C E36 R/C Mut R/C Present Direct R/C Mut E37 R/C Mut R/C Present R/C Direct E38 R/C Mut R/C Present R/C Direct Mut E39 R/C Mut R/C Present R/C R/C E40 R/C Mut R/C Present R/C R/C Mut E41 R/C Mut R/C Present R/C Mut Direct E42 R/C Mut R/C Present R/C Mut Direct Mut E43 R/C Mut R/C Present R/C Mut R/C E44 R/C Mut R/C Present R/C Mut R/C Mut E45 R/C Mut R/C Mut Present Direct Direct E46 R/C Mut R/C Mut Present Direct Direct Mut
E47 R/C Mut R/C Mut Present Direct R/C E48 R/C Mut R/C Mut Present Direct R/C Mut E49 R/C Mut R/C Mut Present Direct Mut Direct E50 R/C Mut R/C Mut Present Direct Mut Direct Mut E51 R/C Mut R/C Mut Present Direct Mut R/C E52 R/C Mut R/C Mut Present Direct Mut R/C Mut E53 R/C Mut R/C Mut Present R/C Direct E54 R/C Mut R/C Mut Present R/C Direct Mut E55 R/C Mut R/C Mut Present R/C R/C E56 R/C Mut R/C Mut Present R/C R/C Mut E57 R/C Mut R/C Mut Present R/C Mut Direct E58 R/C Mut R/C Mut Present R/C Mut Direct Mut E59 R/C Mut R/C Mut Present R/C Mut R/C E60 R/C Mut R/C Mut Present R/C Mut R/C Mut .sup.1see construct #1, FIG. 2B and construct #8, FIG. 8; 2: See construct #2, FIG. 2A and 2C; construct #4 FIG. 4; construct #9 FIG. 9 .sup.3see construct, #3 and construct #6 As used herein, "Direct" means the direct sequence (i.e., a sequence having the same order of nucleotides and in the same orientation) of a smRNA; "Direct Mut" means the direct sequence of an smRNA having a mutation that renders it resistant to cleavage; "R/C" means the reverse complement of an smRNA; and "R/C Mut" means a reverse complement of an smRNA having a mutation that renders it resistant to cleavage.
[0160] In embodiments according to the present disclosure, the sequences of the embodiments of Table 1 include, but are not limited to combinations of SEQ ID NOs: 26 to 35, and 41 to 288, their complements, and non-cleavable mutants thereof. In some embodiments, RNA sequence for regulating a target gene of interest comprises a nucleic acid having 90 to 100% homology to a sequence selected from the group consisting of SEQ ID NOs:8, 11, 12, 36 to 38, and their complements thereof. It is understood that the present disclosure provides for, and includes, dsRNA constructs of Table 1 having a second reverse complementary strand at least to the antisense RNA sequence. The present disclosure further provides dsRNA constructs having a second reverse complimentary strand comprising an antisense RNA sequence and a 5' smRNA.sub.1 sequence. The present disclosure further provides dsRNA constructs having a second reverse complimentary strand comprising an antisense RNA sequence, a 5' smRNA.sub.1 sequence and a 3' smRNA.sub.1 sequence. The present disclosure further provides dsRNA constructs having a second reverse complimentary strand comprising an antisense RNA sequence, a 5' smRNA.sub.1 sequence, a 5' smRNA.sub.2 sequence and a 3' smRNA.sub.1 sequence. The present disclosure further provides dsRNA constructs having a second reverse complimentary strand comprising an antisense RNA sequence, a 5' smRNA.sub.1 sequence, a 5' smRNA.sub.2 sequence, a 3' smRNA.sub.1 sequence and a 4' smRNA.sub.2 sequence.
[0161] The present disclosure provides for and includes second reverse complementary strand of the constructs of Table 1 having mismatches. In some embodiments, the second reverse complementary strand provides for a double stranded region comprising a smRNA and its non-cleavable mutant. Accordingly it is understood, the dsRNA comprises one or more mismatches corresponding to the mismatch between the smRNA and its non-cleavable mutant. The present disclosure provides for combinations of the first strands of Table 1 to produce dsRNA molecules.
[0162] Accordingly, it is understood that construct A02 may be combined with, for example, the reverse complement of A02 to prepare a dsRNA of the present disclosure. In other embodiments, for example, construct A02 may be combined with the reverse complement of construct A06 to prepare a dsRNA of the present disclosure having a mismatch sequence at the non-cleavable site. One of ordinary skill in the art would recognize that additional combinations of the constructs of Table 1 may be prepared in accordance with the present disclosure.
[0163] In embodiments according to the present disclosure, the sequence complementarity may be, but are not required to be, 100%. In certain embodiments of the disclosure the degree of complementarity, e.g., percent complementarity, need only be sufficient to provide for stable binding of a smRNA to the complementary site. In certain embodiments of the disclosure the degree of complementarity need only be sufficient such that the smRNA pairs to the complementary site and mediates cleavage of the target mRNA. For example, in certain embodiments of the disclosure the degree of complementarity is at least 70%, at least 80%, or at least 90%. In certain embodiments of the disclosure the number of mismatched or unpaired nucleotides in the siRNA strand or miRNA, following binding to the complementary site, is between 0 and 5, e.g., 1, 2, 3, 4, or 5.
[0164] In other embodiments according to the present disclosure, the sequence complementarity of an smRNA may be greater than 90%. In some embodiments, the sequence complementarity of an smRNA may be greater than 91%. In some embodiments, the sequence complementarity of an smRNA may be greater than 92%. In some embodiments, the sequence complementarity of an smRNA may be greater than 93%. In some embodiments, the sequence complementarity of an smRNA may be greater than 94%. In some embodiments, the sequence complementarity of an smRNA may be greater than 95%. In some embodiments, the sequence complementarity of an smRNA may be greater than 96%. In some embodiments, the sequence complementarity of an smRNA may be greater than 97%. In some embodiments, the sequence complementarity of an smRNA may be greater than 98%. In some embodiments, the sequence complementarity of an smRNA may be greater than 99%. In some embodiments, the sequence complementarity of an smRNA may be 100%. In embodiments according to the present disclosure, sequence complementarity may be between 90 and 100% or 95 and 100%. According to embodiments of the present disclosure the smRNAs may be selected from the group consisting of SEQ ID NOs 26 to 35, 41 to 288, and non-cleavable mutants thereof.
[0165] As used herein, the terms "complementarity" and "complementary" refer to a nucleic acid that can form one or more hydrogen bonds with another nucleic acid sequence by either traditional Watson-Crick or other non-traditional types of interactions. It will be recognized that complementarity and homology or identity are related terms. That is, a homology describes the degree of similarity between two or more nucleotide sequences when examined in the same 5' to 3' orientation. In contrast, complementarity describes the degree of similarity between two or more nucleotide sequences when comparing a sequence having a 5' to 3' orientation to a sequence having a 3' to 5' orientation. Thus, a first and second sequence having 90% homology will also have 90% complementarity when the first sequence is compared to the reverse complement of the second sequences. In reference to the nucleic molecules of the presently disclosed subject matter, the binding free energy of a nucleic acid molecule with its complementary sequence is sufficient to allow the relevant function of the nucleic acid to proceed, in some embodiments, to form a duplex structure under physiological conditions in a plant cell, to mediate ribonuclease activity, etc. For example, the degree of complementarity between the sense and antisense strands of an miRNA precursor can be the same or different from the degree of complementarity between the miRNA-containing strand of an miRNA precursor and the target nucleic acid sequence. Determination of binding free energies for nucleic acid molecules is well known in the art. See e.g., Freier et al., 1986; Turner et al., 1987. One of ordinary skill in the art would be able to test for sufficiency of complementarity by random or site directed mutagenesis and screening of silencing activity and dsRNA molecule stability in vivo.
[0166] In certain embodiments, the phrase "percent complementarity" refers to the percentage of residues in a nucleic acid molecule that can form hydrogen bonds (e.g., Watson-Crick base pairing) with a second nucleic acid sequence (e.g., 5, 6, 7, 8, 9, 10 out of 10 being 50%, 60%, 70%, 80%, 90%, and 100% complementary). The terms "100% complementary", "fully complementary", and "perfectly complementary" indicate that all of the contiguous residues of a nucleic acid sequence can hydrogen bond with the same number of contiguous residues in a second nucleic acid sequence. It will be appreciated that the nucleic acids may have different lengths and/or that there may be bulges when the two nucleic acids are optimally aligned for maximum complementarity over a given portion of either sequence. Percent complementarity can, in various embodiments of the disclosure, disregard such bulges in the computation or consider the percentage complementarity to be the number of paired (hydrogen bonded) residues divided by the total number of residues over a given length, which may be the length of the shorter or the longer nucleic acid in different embodiments.
[0167] Any complementary sequence for a smRNA may be used in various embodiments of the present disclosure. The sequence may be complementary to an miRNA or siRNA. The sequence may be perfectly (100%) complementary or may have imperfect complementarity as described herein and known in the art. The complementary sequence may be one that is naturally found in a trans-acting-siRNA-producing (TAS) locus or to-siRNA precursor RNA, flanking the portion of the RNA that is cleaved to produce ta-siRNAs. The complementary sequence may be any smRNA complementary sequence that is found on one side of a nucleic acid sequence that is cleaved to produce siRNA, wherein a second smRNA complementary sequence is found on the other side of the nucleic acid sequence. In various embodiments of the disclosure the complementary sequence is recognized by a smRNA selected from the group consisting of:
TABLE-US-00002 miR390: (SEQ ID NO: 25) AAGCUCAGGAGGGAUAGCGCC; miR161.1: (SEQ ID NO: 26) UUGAAAGUGACUACAUCGGGG; miR400: (SEQ ID NO: 27) UAUGAGAGUAUUAUAAGUCAC; TAS2 3'D6(-): (SEQ ID NO: 28) AUAUCCCAUUUCUACCAUCUG; TAS 1b 3'D4(-): (SEQ ID NO: 29) UUCUUCUACCAUCCUAUCAAU; TAS3 5'D7(+): (SEQ ID NO: 30) UUCUUGACCUUGUAAGACCCC; TAS3 5'D8(+): (SEQ ID NO: 31) UUCUUGACCUUGUAAGGCCUU; miR168: (SEQ ID NO: 32) UCGCUUGGUGCAGGUCGGGAA; miR828 (SEQ ID NO: 33) UCUUGCUUAAAUGAGUAUUCCA; and miR393: (SEQ ID NO: 34) UCCAAAGGGAUCGCAUUGAUC.
[0168] In one embodiment, the miRNA is UUCGCUUGCAGAGAGAAAUCAC (SEQ ID NO: 35). Note that these sequences may have been identified in one or more plants, e.g., Arabidopsis, most land plants, moss, etc. It will be appreciated that in some cases the sequences are conserved across multiple species while in other cases there could be minor variations. Such variations are encompassed within the present disclosure. It will be appreciated that homologous siRNAs or miRNAs from other plant species than those listed could be used. Optionally, recognition of the complementary sequence by the cognate miRNA or siRNA leads to cleavage. One of skill in the art could determine whether binding and/or cleavage of a smRNA to a candidate complementary sequence occurs in vivo (in living cells or organisms) or in vitro, e.g., under conditions approximating physiological intercellular conditions. In other embodiments, an smRNA may be selected from the group consisting of SEQ ID NOs:41 to 288 (see, Table 1 of U.S. Pat. No. 8,143,480).
[0169] The length of the complementary sequence could vary. The length of a complementary sequence may be defined as equal to the length of the smRNA that binds to it, but it will be appreciated that a complementary sequence could differ in length from that of the smRNA, e.g., it may be shorter than the length of the smRNA. Typically the complementary sequence is sufficiently long such that the smRNA can bind (e.g., hybridize) to the sequence with reasonable specificity and, optionally, direct cleavage within a duplex structure formed upon binding. Such cleavage may occur at a position within the duplex typical of cleavage directed by smRNAs, (e.g., in certain embodiments at position 10 or 11 of the smRNA). For example, a complementary sequence could be between 15 and 24 nucleotides in length, or any intervening number, wherein there are 1, 2, 3, 4, or 5 mismatches when the smRNA is paired with the complementary sequence in the case of a 15 nucleotide sequence and up to 6, 7, or 8 mismatches in the case of a 24 nucleotide complementary sequence. Similar considerations would apply for other smRNA complementary sequence. It will be appreciated that there may be "bulges" in the duplex formed when an smRNA pairs with its complementary sequence. In such instances a bulge could be considered equivalent to a single mismatch or, in various embodiments of the disclosure a bulge of X nucleotides could be considered equivalent to X mismatches. It will also be appreciated that the specificity of binding of the smRNA to the complementary sequence need not be completely specific, e.g., the smRNA may bind to different sequence having either a lesser or greater degree of complementarity.
[0170] In embodiments according to the present disclosure, a first or second complementary sequence may be between 15 and 30 nt, between 18 and 24 nt, between 20 and 22, or exactly 21 nt in length. In some embodiments a first or second complementary sequence may comprise any intervening range or specific value within the foregoing ranges in certain embodiments of the disclosure. In certain non-limiting embodiments of the disclosure the number of mismatched or unpaired nucleotides in the siRNA strand or miRNA, following binding to the complementary sequence, is between 0 and 5, e.g., 1, 2, 3, 4, or 5. In certain non-limiting embodiments of the disclosure the number of mismatched or unpaired nucleotides (including those in both strands) in a duplex structure formed between the smRNA and its complementary sequence, is between 0 and 10, e.g., 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. The mismatches or bulges may occur at any position within the duplex structure, in various embodiments of the disclosure. In certain embodiments the mismatches or bulges are located at positions known in the art not to typically inhibit or prevent smRNA-directed cleavage. In other embodiments the mismatches or bulges are located at such positions. One or more mismatches or bulges may occur, for example, at any position with respect to the 5' end of a smRNA depicted in the figures herein or contemplated when an smRNA described herein pairs with a sequence complementary to it. The mismatch may be any mismatch known in the art. In certain embodiments a mismatch is said to occur when a nucleotide within an at least partly double-stranded structure is not paired in a conventional G-C, A-T, or A-U base pair. In certain embodiments a mismatch is said to occur when a nucleotide in an at least partly double-stranded structure is not paired in a Watson-Crick base pair. It will be appreciated that the aforementioned mismatches may exclude "bulges", wherein a nucleotide bulges outward from an otherwise duplex region by being located between two nucleotides that are base paired with adjacent nucleotides on the opposite strand of the duplex.
[0171] The present disclosure further includes and provides for embodiments wherein a first and a second complementary sequence are the same length. In other embodiments, a first complementary sequence may be a different length than a second complementary sequence.
[0172] The portion of the smRNA that is complementary to the complementary sequence could vary. For example, in certain embodiments the complementary site is at least 70%, at least 80%, or at least 90% complementary to the first 16 nucleotides of the smRNA. In certain embodiments the complementary sequence is at least 70%, at least 80%, or at least 90% complementary to the first 17, 18, or 19 nucleotides of the smRNA. In certain embodiments the complementary sequence is a subsequence of a complementary sequence, wherein said subsequence is at least 16, 17, 18, 19, 20, or 21 nucleotides in length. In some embodiments the subsequence is the last 16, 17, 18, 19, 20, or 21 nucleotides of the listed sequence. In some embodiments the subsequence is at least 70%, at least 80%, at least 90%, or 100% complementary to the first 16, 17, 18, 19, 20, or 21 nucleotides of an smRNA.
[0173] The present disclosure also includes, and provides for, dsRNA molecules having a single smRNA complementary sequence and an antisense RNA sequence for regulating a target gene of interest. In embodiments according to the present disclosure, where the dsRNA molecule is presented as an antisense sequence, an smRNA sequence on the 5' end is an upstream sequences. In other embodiments, an smRNA sequence on the 3' end is a downstream sequence. Not to be limited by theory, a smRNA complementary sequence binds (hybridizes) to a complementary smRNA and the duplex recruits an RDRP, such as RNA-dependent RNA polymerase 6 (RDR6). The dsRNA molecule can be further processed by Dicer-like enzymes such as dicer-like protein 4 (DCL4) to produce the siRNAs. It will be appreciated that the presence of a second complementary sequence (e.g., flanking sequences) depends on the sequence of smRNA that the complementary sequence recognizes. In some instances (i.e., miR390) it is required whereas in others it is not required (i.e., miR173). In some aspects, for smRNAs that do not require a second complementary sequence (flanking sequence), the inclusion of a second complementary sequence may increase the propensity, efficiency or rate of generating siRNAs.
[0174] When only one smRNA complementary sequence is present in a dsRNA molecule, then it can be upstream (5') or downstream (3') to the dsRNA sequence for silencing the target gene (where the dsRNA sequence is referenced in the antisense orientation). According to a specific embodiment, this smRNA complementary sequence is functional and is located 3' to the dsRNA sequence for silencing the target gene. According to another specific embodiment, this smRNA complementary sequence is functional (e.g., miR173) and is located 5' to the dsRNA sequence for silencing the target gene.
[0175] When two complementary sequences to one or more an smRNAs are included, one sequence (e.g., the first site) is located 5' to the dsRNA sequence for silencing the target gene (where the dsRNA sequence is referenced in the antisense orientation), and the second site is located 3' to the dsRNA sequence for silencing the target gene (e.g., flanking smRNA sequences). Alternatively, the present disclosure also provides for dsRNA molecules having the second site located 5' to the dsRNA sequence for silencing the target gene, and the first site located 3' to dsRNA sequence for silencing the target gene.
[0176] It will be appreciated that the complementary sequences can be positioned on one strand (sense) and the other on the other strand (antisense). It will be further appreciated that in the presence of two complementary sequences to an smRNA or smRNAs, one of said sequences can mediate binding of the smRNA but not cleavage of the dsRNA sequence for silencing the target gene. Thus, one of the complementary binding sequences is essentially an smRNA mimic sequence (e.g. sufficient for binding but not cleavage).
[0177] The smRNA mimic sequence is essentially complementary to the microRNA or siRNA provided that one or more mismatches are allowed: thus, a mismatch between the complementary nucleotides at position 10 or position 11 of the microRNA and the corresponding nucleotide sequence in the micro-RNA resistant site. As used herein, the term "smRNA mimic," "smRNA mutant," and "miRNA Mut" are used interchangeably and refer to smRNAs that are not cleaved in a cell. Not to be limited by any particular theory, mimic or mutant are thought mediate binding of the machinery, such as the Argonaute protein family, but are not processed, for example by a dicer-like protein. Accordingly, smRNA mimics or mutants interact with the RISC complex but can not be cleaved. Thus a non-cleavable target mimic of a smRNA acts to sequester the corresponding target miRNA and arrest its activity. By incorporating a miRNA target mimic having a non-cleavable target site the accumulation of all MIR gene family members may be reduced. Methods for preparing smRNA mutants or mimics are known. See Todesco et al., "A Collection of Target Mimics for Comprehensive Analysis of MicroRNA Function in Arabidopsis thaliana," PLOS Genetics 6(7):e1001031 (2010); Wang Z., "The guideline of the design and validation of MiRNA mimics," Methods Mol Biol. 676:211-23 (2011);
[0178] The complementary sequences may be identical or different and may be recognized by the same or different smRNAs, which may be miRNA or siRNA, or both, in any combination.
[0179] The complementary sequences(s) can be immediately adjacent to (i.e., contiguous with) the dsRNA sequence for silencing the target gene. Alternatively, the complementary sequence (or at least one of same) may be separated from the dsRNA sequence for silencing the target gene by an intervening spacer or functional sequence (e.g., Helicase binding site).
[0180] As used herein a "helicase binding site" refers to a binding site of an RNA helicase. RNA helicases are essential for most processes of RNA metabolism such as ribosome biogenesis, pri-mRNA splicing and translation initiation. Sequence information is available from the RNA Helicase Database, available on the internet at wwwdotrnahelicasedotorg/. According to a specific embodiment, the helicase may be a DEAD RNA helicase (DEAD RH), such as described in Chi et al. (2012). "The Function of RH22, a DEAD RNA Helicase, in the Biogenesis of the 50S Ribosomal Subunits of Arabidopsis Chloroplasts," Plant Physiology, 158, 693-707. In certain embodiments, a helicase binding site may be positioned so as to allow unwinding of the strands of a dsRNA molecule to single stranded RNA (ssRNA) and allow recruitment of an RNA-dependent RNA Polymerase such as RDR6. Unwinding and recruitment of an RDRP provides for amplification of a dsRNA molecule in the plant cell. Other proteins that are known to be cytosolic proteins and have helicase or helicase-like activity include the Argonaute protein family, which are a key components of the RISC complex (RNA Induced Silencing Complex). Alternatively a helicase binding site may includes sequences recognized by the plant homolog of RNA helicase RIG-I (Yoneyama et al. 2004 Nat. Immun. 5:730-737). Alternatively or additionally, the present disclosure provides for helicase binding site sequences as described in Garcia et al. 2012 Mol. Cell 48(1):109-20, which is hereby incorporated by reference in its entirety.
[0181] According to a specific embodiment, the helicase binding site is positioned upstream or downstream of the dsRNA sequence for silencing expression of the target gene (where the dsRNA sequence is referenced in the antisense orientation).
[0182] According to a specific embodiment, the helicase binding site is positioned upstream or downstream of the dsRNA sequence (where the dsRNA sequence is referenced in the antisense orientation) and the two complementary sites to the smRNA or smRNAs flank the helicase binding site.
[0183] According to a further specific embodiment, the helicase binding site is located within the dsRNA sequence corresponding to the target site.
[0184] "Flanked by" as used herein, does not require that the smRNA complementary sequences are contiguous with the dsRNA sequence for silencing the target gene. All that is necessary is that there is an smRNA complementary sequences on at least one side or in case of two complementary sequences on each side of the dsRNA sequence for silencing the target gene. Either or both smRNA complementary sequences may, in various embodiments of the disclosure, be located contiguously with the dsRNA sequence for silencing the target gene. In certain embodiments either or both smRNA complementary sequences may, in various embodiments of the disclosure, be separated from the dsRNA sequence for silencing the target gene by between 1 nt and 2 kB, e.g., between 1 nt and 1 kB, between 1 nt and 500 nt, between 1 nt and 250 nt, between 1 nt and 100 nt, etc. In certain embodiments either or both smRNA complementary sequences are separated from a portion of the nucleic acid sequence that corresponds to the target gene by between 10 and 20 nt, between 10 and 50 nt, or between 10 and 100 nt. Thus the spacer between either smRNA complementary sequences and the closest nucleotide that corresponds to a portion of a target gene may, in various embodiments of the disclosure, be between 1 nt and 2 kB, e.g., between 1 nt and 1 kB, between 1 nt and 500 nt, between 1 nt and 250 nt, between 1 nt and 100 nt, between 10 and 20 nt, between 10 and 50 nt, or between 10 and 100 nt in length.
[0185] Without being bound to theory, it is suggested that following introduction into the plant, the dsRNA molecule is unwound either by the binding of a helicase to a "helicase binding site" when present or by endogenous RNA helicases that recognize and unwind the dsRNA molecule in a manner similar to antiviral response in a case where it is absent.
[0186] Without being bound to a particular theory, once a single stranded molecule is formed is processed by miRNA-guided-cleavage. One product of the cleaved transcript may be stabilized possibly by Suppressor of Gene Silencing 3 (SGS3) and converted to dsRNA by RNA-Dependent RNA Polymerase 6 (RDR6). The resulting dsRNA may be processed through Dicer-Like 4 (DCL4) into 21-nt siRNA duplexes in register with the miRNA-cleavage site. One strand of each smRNA duplex may be selectively sorted to one or more Argonaute (AGO) proteins according to the 5' nucleotide sequence while the other is used as a template for RNA dependent RNA polymerase, thereby constantly generating more phase siRNA molecule.
[0187] Thus, the dsRNA molecule is designed for specifically targeting a target gene of interest. It will be appreciated that the dsRNA can be used to down-regulate one or more target genes. In some embodiments, a single isolated dsRNA molecule can target a number of different genes.
[0188] The present disclosure provides for and includes heterogenic compositions of dsRNA molecules. In certain embodiments wherein a dsRNA molecule targets a single target gene of interest, heterogenic compositions comprising two or more dsRNA molecules that target two or more target genes of interest may be prepared. A heterogenic composition comprises a plurality of dsRNA molecules for targeting a number of target genes may be prepared. In some embodiments, a plurality of dsRNA molecules may be separately applied to the seeds (but not as a single composition).
[0189] The present disclosure provides for an includes dsRNA molecules comprising a sequence, wherein said nucleic acid sequence shares between 100% and 90% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 90% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 91% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 92% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 93% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 94% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 95% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 96% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 97% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 98% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 99% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299. In other embodiments according to the present disclosure, a dsRNA molecule comprises a sequence that shares more than 100% sequence identity to a nucleic acid sequence selected from the group consisting of SEQ ID NO: 289-299.
[0190] While conceiving the present disclosure, the present inventors realized that the long dsRNA identified by Fukuhara et al. is able to survive autonomously in rice cells, and as such can be used as a cassette (a building block) for introducing dsRNA sequenced for RNA silencing of a target gene of interest (endogenous to the plant or exogenous thereto). Such a dsRNA molecule is expressed throughout the plant's life cycle, does not become integrated into the plant genome (plastid or nuclear), and does not get reverse-transcribed into DNA.
[0191] Thus, in some embodiments of the present disclosure provide for endovirus-derived sequences which have evolved to co-exist in plant cells in a dsRNA form, maintaining a near exact copy number in all cells.
[0192] As used herein the term "endovirus" refers to a dsRNA symbiotic virus which propagates in the plant cell and maintains a relatively stable copy number throughout the life cycle of the plant.
[0193] As used herein the term "5' UTR" refers to an untranslated region derived from the endovirus sequence (13,716 nucleotides, available from GSDB, DDBJ, EMBL, and NCBI nucleotide sequence data bases with accession number D32136, according to Fukuhara 1995 supra), adjacent (in a 5' orientation) to its RDRP sequence.
[0194] As used herein the term "3' UTR" refers to an untranslated region derived from the endovirus sequence (13,716 nucleotides, available from GSDB, DDBJ, EMBL, and NCBI nucleotide sequence data bases with accession number D32136, according to Fukuhara 1995 supra), adjacent (in a 3' orientation) to its RDRP sequence.
[0195] As used herein the term "RNA dependent RNA Polymerase" refers to the RDRP-like sequence derived from the 13,716 nucleotides described in Fukuhara et al. 1995 supra.
[0196] According to a specific embodiment said 5' UTR is as set forth in SEQ ID NO: 14. It will be appreciated that the sequences are provided in the form of DNA but will be made RNA upon subjecting to T7 activity. In some embodiments, the 5' UTR shares between 90% and 100% sequences identity to a nucleic acid sequence of SEQ ID NO:14.
[0197] According to a specific embodiment said 3' UTR is as set forth in SEQ ID NO: 22. According to a specific embodiment said endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence is as set forth in SEQ ID NO: 23.
[0198] According to some embodiments of the disclosure, the nucleic acid sequence is at least about 80%, at least about 81%, at least about 82%, at least about 83%, at least about 84%, at least about 85%, at least about 86%, at least about 87%, at least about 88%, at least about 89%, at least about 90%, at least about 91%, at least about 92%, at least about 93%, at least about 94%, at least about 95%, at least about 96%, at least about 97%, at least about 98%, at least about 99%, or more say 100% identical to the nucleic acid sequence of SEQ ID NO: 14, 22 or 23 (or a combination of same). It is understood that combinations of SEQ ID NOs: 14, 22, or 23 having differing percent homology are envisioned. As a non-limiting example, a combination dsRNA molecule may comprise a sequence having 85% homology to SEQ ID NO:14, 90% homology to SEQ ID NO:22 and 99% homology to SEQ ID NO:23. Any number of such combinations are contemplated and provided for in the present disclosure.
[0199] One of ordinary skill in the art would understand that certain nucleotide positions of a polynucleotide sequence play critical roles while other nucleotide positions may vary without significant effect. This understanding is well illustrated for polypeptide encoding sequences. For example, the first and second nucleotide of an amino acid encoding codon largely determine the identity of an amino acid in a polypeptide chain. In contrast, the third "wobble" position can vary, sometimes without limitation. Accordingly, one of ordinary skill in the art can substitute approximately 30% of the nucleic acid sequence without affecting the amino acid sequence. A similar dependence for non-coding sequences, including 3' and 5' UTRs exists, though the positions of importance may not be predictable. However, one of ordinary skill in the art may mutagenize (randomly or through site directed) positions in a nucleic acid sequence and confirm their activity using the functional assays disclosed in the present application (See, Examples below).
[0200] According to some embodiments, a nucleic acid sequence may encode a protein that shares between 70% and 100% homology with an RNA Dependent RNA Polymerase (RDRP) polypeptide sequence according to SEQ ID NO: 300. A nucleic acid sequence according to the present disclosure may encode a protein having 70 to 75% homology to a polypeptide sequence according to SEQ ID NO: 300. In another embodiment, a nucleic acid sequence may encode a protein having 75 to 80% homology to a polypeptide sequence according to SEQ ID NO: 300. In another embodiment, a nucleic acid sequence may encode a protein having 80 to 85% homology to a polypeptide sequence according to SEQ ID NO: 300. In an embodiment, a nucleic acid sequence may encode a protein having 85 to 90% homology to a polypeptide sequence according to SEQ ID NO: 300. In another embodiment, a nucleic acid sequence may encode a protein having 90 to 95% homology to a polypeptide sequence according to SEQ ID NO: 300. In an embodiment, a nucleic acid sequence may encode a protein having 95 to 100% homology to a polypeptide sequence according to SEQ ID NO: 300. In another embodiment, a nucleic acid sequence may encode a protein having 97 to 100% homology to a polypeptide sequence according to SEQ ID NO: 300. In an embodiment, a nucleic acid sequence may encode a protein having 98 to 100% homology to a polypeptide sequence according to SEQ ID NO: 300. In another embodiment, a nucleic acid sequence may encode a protein having 99 to 100% homology to a polypeptide sequence according to SEQ ID NO: 300.
[0201] Homologous sequences to the above can also be used according to the present teachings, as long as their main characteristics are maintained, i.e., amplification by RDRP and maintenance of stable copy number in the cell throughout the plant life cycle.
[0202] The present disclosure further provides for, and includes, an isolated dsRNA molecule comprising a nucleic acid sequence in a sequential order from 5' to 3', an endovirus 5' UTR, an endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence, an endovirus 3' UTR and a cloning site flanked by said RDRP and said 3' UTR.
[0203] The present disclosure further provides for, and includes, an isolated dsRNA molecule comprising a nucleic acid sequence in a sequential order from 5' to 3', an endovirus 5' UTR, an endovirus RNA Dependent RNA Polymerase (RDRP) coding sequence, an endovirus 3' UTR and a nucleic acid sequence for regulating a target gene flanked by said RDRP and said 3' UTR.
[0204] In some embodiments, a heterologous dsRNA sequence corresponding to a target gene is constructed such that it is flanked by the RDRP sequence and the 3' UTR. Alternatively, in other embodiments, a heterologous dsRNA sequence corresponding to the target gene is constructed such that it is flanked by the RDRP sequence and the 5' UTR. When introduced into the plant (e.g., directly to the seed), and once germination has initiated, gene expression occurs including expression of endogenous plant helicases, RDRPs and other components of the silencing machinery. Not to be limited by theory, at any given time, a portion of the dsRNA molecules is recognized and processed by the plant's dicer like (DCL) proteins into siRNA of different lengths. In certain embodiments, this recognition and processing includes processing of the gene targeted for silencing, which is flanked between the 5' and the 3' UTR. In certain embodiments, the gene targeted for silencing and the RDRP sequence is flanked between the 5' and the 3' UTR. Not all of the heterologous dsRNA sequence is processed and a portion remains in double-stranded, full length form. Not to be limited by theory, it is thought that plant helicases recognize unique features in the 5' UTR and the 3' UTR of the dsRNA and unwind the dsRNA into two ssRNA molecules. Again, not limited by theory, the same or other feature in the 5' and 3' UTR are thought to also recruit and activate an RDRP. In some embodiments, the RDRP may be an RDRP that is encoded by the dsRNA. In other embodiments, the RDRP may be an endogenous RDRP, naturally occurring in the plant or introduced as a transgene.
[0205] Though not limited by theory, it is thought that the RDRP uses each of the ssRNA molecules thought to be produced by the activity of a helicase as templates to produce a dsRNA molecule identical to the original dsRNA molecule.
[0206] Accordingly, and not to be limited by theory, as long as a ratio between processed and un-processed dsRNA molecules is maintained, the cycle can go on and repeat throughout the plant's life cycle. Also not to be limited by theory, it is thought that some features in the 5' and 3' UTR can assist to maintain a stable copy number of the dsRNA in cells, in a similar manner to the endovirus which is maintained at a stable copy number.
[0207] As used herein the term "sequential" refers to multiple nucleic acid segments (e.g., 5' UTR, RDRP and 3' UTR) arranged in sequence. In this case, from 5' to 3. Each of the specified nucleic acid segments can be directly attached to each other and contiguous, however intervening nucleic acids can be implanted there between such that the segments are not directly attached to each other and are discontinuous.
[0208] According to an embodiment of the disclosure the target gene is endogenous to the plant. Downregulating such a gene is typically important for conferring the plant with an improved, agricultural, horticultural, nutritional trait ("improvement" or an "increase" is further defined hereinbelow).
[0209] As used herein, the terms "suppress," "repress," and "downregulate" when referring to the expression or activity of a nucleic acid molecule in a plant cell are used equivalently herein and mean that the level of expression or activity of the nucleic acid molecule in a plant, a plant part, or plant cell after applying a method of the present disclosure is lower than its expression or activity in the plant, part of the plant, or plant cell before applying the method, or compared to a control plant lacking a dsRNA molecule of the disclosure.
[0210] The terms "suppressed," "repressed" and "downregulated" as used herein are synonymous and mean herein lower, preferably significantly lower, expression or activity of the nucleic acid molecule to be expressed.
[0211] As used herein, a "suppression," "repression," or "downregulation" of the level or activity of an agent such as a protein, mRNA, or RNA means that the level or activity is reduced relative to a substantially identical plant, part of a plant, or plant cell grown under substantially identical conditions, lacking a dsRNA molecule of the disclosure, for example, lacking an RNA sequence for regulating a target gene of interest. As used herein, "suppression," "repression," or "downregulation" of the level or activity of an agent, such as, for example, a preRNA, mRNA, rRNA, tRNA, snoRNA, snRNA expressed by the target gene, and/or of the protein product encoded by it, means that the amount is reduced by 10% or more, for example, 20% or more, preferably 30% or more, more preferably 50% or more, even more preferably 70% or more, most preferably 80% or more, for example, 90%, relative to a cell or organism lacking a dsRNA molecule of the disclosure.
[0212] As used herein "a suppressive amount" refers to an amount of dsRNA molecule which is sufficient to down regulate the target gene by at least 20%. In an aspect, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by 30% or more. In an aspect, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by 40% or more. In an aspect, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by at least 50%. In other aspects, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by 60% or more. In an aspect, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by at least 70%. In an aspect, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by 80% or more. In an aspect, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by greater than 90%. In certain aspects, a suppressive amount according to the present disclosure is an amount sufficient to downregulate a target gene by 100% (e.g., wherein the remaining amount of the target gene is not detectable). The suppressive amount can be a result of the formation of amplification in the plant or the phytopathogen.
[0213] As used herein "endogenous" refers to a gene which expression (mRNA or protein) takes place in the plant. Typically, the endogenous gene is naturally expressed in the plant or originates from the plant. Thus, the plant may be a wild-type plant. However, the plant may also be a genetically modified plant (transgenic).
[0214] Downregulation of the target gene may be important for conferring improved one of-, or at least one of (e.g., two of- or more), biomass, vigor, yield, abiotic stress tolerance, biotic stress tolerance or improved nitrogen use efficiency.
[0215] Examples of target genes include, but are not limited to, an enzyme, a structural protein, a plant regulatory protein, an miRNA target gene, or a non-coding RNA such as a miRNA of the plant. WO2011067745, WO 2009125401 and WO 2012056401 provide examples of miRNA sequences or targets of miRNAs (e.g., miRNA167, miRNA169, miRNA 156, miR164 and targets thereof ARF, NFY, SPL17 and NAC, respectively) which expression can be silenced to improve a plant trait.
[0216] The target gene may comprise a nucleic acid sequence which is transcribed to an mRNA which codes for a polypeptide.
[0217] Alternatively, the target gene can be a non-coding gene such as an miRNA or a siRNA.
[0218] For example, in order to silence the expression of an mRNA of interest, synthesis of the dsRNA molecule suitable for use with some embodiments of the disclosure can be selected as follows. First, the mRNA sequence may be scanned including the 3' UTR and the 5' UTR.
[0219] Second, the mRNA sequence may be compared to an appropriate genomic database using any sequence alignment software, such as the BLAST software available from the NCBI server (available on the internet at wwwdotncbidotnlmdotnihdotgov/BLAST/). Putative regions in the mRNA sequence which exhibit significant homology to other coding sequences may be filtered out.
[0220] Qualifying target sequences may be selected as for the preparation of dsRNA templates for dsRNA molecule synthesis. Preferred sequences are those that have as little homology to other genes in the genome to reduce an "off-target" effect.
[0221] It will be appreciated that the RNA regulating or silencing agent of some embodiments of the disclosure need not be limited to those molecules containing only RNA, but further encompasses chemically-modified nucleotides and non-nucleotides.
[0222] The dsRNA molecules of the present disclosure may be synthesized using any method known in the art, including either enzymatic syntheses or solid-phase syntheses. These are especially useful in the case of short polynucleotide sequences with or without modifications as explained above. Equipment and reagents for executing solid-phase synthesis are commercially available from, for example, Applied Biosystems. Any other means for such synthesis may also be employed; the actual synthesis of the oligonucleotides is well within the capabilities of one skilled in the art and can be accomplished via established methodologies as detailed in, for example: Sambrook, J. and Russell, D. W. (2001), "Molecular Cloning: A Laboratory Manual"; Ausubel, R. M. et al., eds. (1994, 1989), "Current Protocols in Molecular Biology," Volumes I-III, John Wiley & Sons, Baltimore, Md.; Perbal, B. (1988), "A Practical Guide to Molecular Cloning", John Wiley & Sons, New York; and Gait, M. J., ed. (1984), "Oligonucleotide Synthesis"; utilizing solid-phase chemistry, e.g., cyanoethyl phosphoramidite followed by deprotection, desalting, and purification by, for example, an automated trityl-on method or HPLC.
[0223] The nucleic acids of the present disclosure may comprise heterocylic nucleosides consisting of purines and the pyrimidines bases, bonded in a 3' to 5' 5phosphodiester linkage. Preferably used nucleic acids are those modified in either backbone, internucleoside linkages or bases, as is broadly described hereinunder.
[0224] Specific examples of preferred nucleic acids useful according to this aspect of the present disclosure include nucleic acids containing modified backbones or non-natural internucleoside linkages. Nucleic acids having modified backbones include those that retain a phosphorus atom in the backbone, as disclosed in U.S. Pat. Nos. 4,469,863; 4,476,301; 5,023,243; 5,177,196; 5,188,897; 5,264,423; 5,276,019; 5,278,302; 5,286,717; 5,321,131; 5,399,676; 5,405,939; 5,453,496; 5,455,233; 5,466,677; 5,476,925; 5,519,126; 5,536,821; 5,541,306; 5,550,111; 5,563,253; 5,571,799; 5,587,361; and 5,625,050.
[0225] Preferred modified polynucleotide backbones include, for example, phosphorothioates, chiral phosphorothioates, phosphorodithioates, phosphotriesters, aminoalkyl phosphotriesters, methyl and other alkyl phosphonates including 3'-alkylene phosphonates and chiral phosphonates, phosphinates, phosphoramidates including 3'-amino phosphoramidate and aminoalkylphosphoramidates, thionophosphoramidates, thionoalkylphosphonates, thionoalkylphosphotriesters, and boranophosphates having normal 3'-5' linkages, 2'-5' linked analogs of these, and those having inverted polarity wherein the adjacent pairs of nucleoside units are linked 3'-5' to 5'-3' or 2'-5' to 5'-2'. Various salts, mixed salts and free acid forms can also be used.
[0226] Alternatively, modified polynucleotide backbones that do not include a phosphorus atom therein have backbones that are formed by short chain alkyl or cycloalkyl internucleoside linkages, mixed heteroatom and alkyl or cycloalkyl internucleoside linkages, or one or more short chain heteroatomic or heterocyclic internucleoside linkages. These include those having morpholino linkages (formed in part from the sugar portion of a nucleoside); siloxane backbones; sulfide, sulfoxide and sulfone backbones; formacetyl and thioformacetyl backbones; methylene formacetyl and thioformacetyl backbones; alkene containing backbones; sulfamate backbones; methyleneimino and methylenehydrazino backbones; sulfonate and sulfonamide backbones; amide backbones; and others having mixed N, 0, S and CH2 component parts, as disclosed in U.S. Pat. Nos. 5,034,506; 5,166,315; 5,185,444; 5,216,141; 5,235,033; 5,264,562; 5,264,564; 5,405,938; 5,470,967; 5,489,677; 5,541,307; 5,561,225; 5,596,086; 5,602,240; 5,602,240; 5,608,046; 5,610,289; 5,618,704; 5,623,070; 5,663,312; 5,214,134; 5,466,677; 5,610,289; 5,633,360; 5,677,437; and 5,677,439.
[0227] Other nucleic acids which can be used according to the present disclosure, are those modified in both sugar and the internucleoside linkage, i.e., the backbone, of the nucleotide units are replaced with novel groups. The base units are maintained for complementation with the appropriate polynucleotide target. An example for such an polynucleotide mimetic, includes peptide nucleic acid (PNA). A PNA polynucleotide refers to a polynucleotide where the sugar-backbone is replaced with an amide containing backbone, in particular an aminoethylglycine backbone. The bases are retained and are bound directly or indirectly to aza nitrogen atoms of the amide portion of the backbone. United States patents that teach the preparation of PNA compounds include, but are not limited to, U.S. Pat. Nos. 5,539,082; 5,714,331; and 5,719,262, each of which is herein incorporated by reference. Other backbone modifications, which can be used in the present disclosure are disclosed in U.S. Pat. No. 6,303,374.
[0228] Polynucleotide agents of the present disclosure may also include base modifications or substitutions. As used herein, "unmodified" or "natural" bases include the purine bases adenine (A) and guanine (G), and the pyrimidine bases thymine (T), cytosine (C) and uracil (U). Modified bases include but are not limited to other synthetic and natural bases such as 5-methyl cytosine (5-me-C), 5-hydroxymethyl cytosine, xanthine, hypoxanthine, 2-aminoadenine, 6-methyl and other alkyl derivatives of adenine and guanine, 2-propyl and other alkyl derivatives of adenine and guanine, 2-thiouracil, 2-thiothymine and 2-thiocytosine, 5-halouracil and cytosine, 5-propynyl uracil and cytosine, 6-azo uracil, cytosine and thymine, 5-uracil (pseudouracil), 4-thiouracil, 8-halo, 8-amino, 8-thiol, 8-thioalkyl, 8-hydroxyl and other 8-substituted adenines and guanines, 5-halo particularly 5-bromo, 5-trifluoromethyl and other 5-substituted uracils and cytosines, 7-methylguanine and 7-methyladenine, 8-azaguanine and 8-azaadenine, 7-deazaguanine and 7-deazaadenine and 3-deazaguanine and 3-deazaadenine. Further bases include those disclosed in U.S. Pat. No. 3,687,808, those disclosed in The Concise Encyclopedia Of Polymer Science And Engineering, pages 858-859, Kroschwitz, J. I., ed. John Wiley & Sons, 1990, those disclosed by Englisch et al., Angewandte Chemie, International Edition, 1991, 613, and those disclosed by Sanghvi, Y. S., Chapter 15, Antisense Research and Applications, pages 289-2, Crooke, S. T. and Lebleu, B., ed., CRC Press, 1993. Such bases are particularly useful for increasing the binding affinity of the oligomeric compounds of the disclosure. These include 5-substituted pyrimidines, 6-azapyrimidines and N-2, N-6 and 0-6 substituted purines, including 2-aminopropyladenine, 5-propynyluracil and 5-propynylcytosine. 5-methylcytosine substitutions have been shown to increase nucleic acid duplex stability by 0.6-1.2.degree. C. (Sanghvi Y S et al. (1993) Antisense Research and Applications, CRC Press, Boca Raton 276-278) and are presently preferred base substitutions, even more particularly when combined with 2'-O-methoxyethyl sugar modifications.
[0229] Following synthesis, the polynucleotide agents of the present disclosure may optionally be purified. For example, polynucleotides can be purified from a mixture by extraction with a solvent or resin, precipitation, electrophoresis, chromatography, or a combination thereof. Alternatively, polynucleotides may be used with no, or a minimum of, purification to avoid losses due to sample processing. The polynucleotides may be dried for storage or dissolved in an aqueous solution. The solution may contain buffers or salts to promote annealing, and/or stabilization of the duplex strands.
[0230] It will be appreciated that a polynucleotide agent of the present disclosure may be provided per se, or as a nucleic acid construct comprising a nucleic acid sequence encoding the polynucleotide agent. Typically, the nucleic acid construct comprises a promoter sequence which is functional in the host cell, as detailed herein below.
[0231] The polynucleotide sequences of the present disclosure, under the control of an operably linked promoter sequence, may further be flanked by additional sequences that advantageously affect its transcription and/or the stability of a resulting transcript. Such sequences are generally located upstream of the promoter and/or downstream of the 3' end of the expression construct.
[0232] The term "operably linked", as used in reference to a regulatory sequence and a structural nucleotide sequence, means that the regulatory sequence causes regulated expression of the linked structural nucleotide sequence. "Regulatory sequences" or "control elements" refer to nucleotide sequences located upstream, within, or downstream of a structural nucleotide sequence, and which influence the timing and level or amount of transcription, RNA processing or stability, or translation of the associated structural nucleotide sequence. Regulatory sequences may include promoters, translation leader sequences, introns, enhancers, stem-loop structures, repressor binding sequences, termination sequences, pausing sequences, polyadenylation recognition sequences, and the like.
[0233] The present disclosure provides for and includes DNA templates for the preparation of dsRNA molecules. As used herein, "dsRNA template" refers to a DNA sequence having the same sequence as the corresponding dsRNA molecule. In certain embodiments, a dsRNA template may further include additional sequences, such as promoters, sufficient for the expression of one or more RNA molecules via transcription. In some embodiments the promoters are bacteriophage promoters, for example but not limited to, SP6, T3 and T7. In some embodiments a promoter may be a bacterial or a eukaryotic promoters.
[0234] In certain embodiments, a dsRNA template according to the present teachings may be used as a cassette for the cloning of a nucleic acid sequence corresponding to a target gene of interest (exogenous to the plant or endogenous thereto) for silencing expression of same when expressed as a dsRNA molecule.
[0235] Thus according to an embodiment of the disclosure, a dsRNA template may comprise a cloning site (multiple cloning site for instance) to which a nucleic sequence for silencing a target gene of interest is ligated while being flanked by the RDRP encoding and the 3'UTR encoding nucleic acid sequences. In other aspects according to present disclosure, a dsRNA molecule may be prepared by chemical synthesis using methods known in the art. In other embodiments, a dsRNA template may comprise a plasmid vector having the dsRNA molecule coding sequences. In certain embodiments, the dsRNA template may be a linear polynucleotide having a RNA polymerase promoter at one end. It will be appreciated that such a template produces a single strand RNA.
[0236] In certain embodiments, a dsRNA template may comprise a mixture of two linear polynucleotides having the same dsRNA coding sequence but promoters at opposite ends. It will be appreciated that transcription results in the production of two complementary RNA strands from the separate template that may be annealed and recovered, or recovered and annealed. By providing separate transcription templates, and recovering annealed double stranded dsRNA molecules according to the present disclosure, asymmetric dsRNA molecules may be produced. As provided below, dsRNA construct #4 provides for a dsRNA region corresponding to a region of a target gene of interest and having non-double stranded Mir390 Mut BS and Mir390BS sequences. As used herein, the term "overhang" refers to non-double stranded regions of a dsRNA molecule (i.e., single stranded RNA). Accordingly dsRNA construct #4 has two overhang regions comprising Mir390 Mut BS and Mir390 BS sequences respectively. Similarly, dsRNA construct #6 provides for an asymmetric dsRNA having a non-double stranded helicase binding sequence or overhang.
[0237] In other aspects, a dsRNA template may have two promoters flanking the dsRNA coding sequences. It will be appreciated that, like the separate templates, two complementary strands are produced that may be annealed and recovered, or recovered and annealed. The promoters of the dsRNA templates of some embodiments may be the same or different.
[0238] As mentioned, in certain embodiments, the dsRNA molecule may be directly contacted with the seed.
[0239] The seed may be of any plant, such as of the Viridiplantae super family including monocotyledon and dicotyledon plants. Other plants are listed below. According to an embodiment of the disclosure, the cells of the plant comprise RNA dependent RNA polymerase activity and the target RNA molecule of the dsRNA to ensure amplification of the dsRNA.
[0240] The term "plant" as used herein encompasses whole plants, ancestors and progeny of the plants and plant parts, including seeds, shoots, stems, roots (including tubers), and isolated plant cells, tissues and organs. The plant may be in any form including suspension cultures, embryos, meristematic regions, callus tissue, leaves, gametophytes, sporophytes, pollen, and microspores. It will be appreciated, that the plant or seed thereof may be transgenic plants.
[0241] As used herein the phrase "plant cell" refers to plant cells which are derived and isolated from disintegrated plant cell tissue or plant cell cultures.
[0242] As used herein the phrase "plant cell culture" refers to any type of native (naturally occurring) plant cells, plant cell lines and genetically modified plant cells, which are not assembled to form a complete plant, such that at least one biological structure of a plant is not present. Optionally, the plant cell culture of this embodiment of the present disclosure may comprise a particular type of a plant cell or a plurality of different types of plant cells. It should be noted that optionally plant cultures featuring a particular type of plant cell may be originally derived from a plurality of different types of such plant cells. In certain embodiments according to the present disclosure, the plant cell is a non-sexually producing plant cell. In other aspects, a plant cell of the present disclosure is a non-photosynthetic plant cell.
[0243] Any commercially or scientifically valuable plant is envisaged in accordance with some embodiments of the disclosure. Plants that are particularly useful in the methods of the disclosure include all plants which belong to the super family Viridiplantae, in particular monocotyledonous and dicotyledonous plants including a fodder or forage legume, ornamental plant, food crop, tree, or shrub selected from the list comprising Acacia spp., Acer spp., Actinidia spp., Aesculus spp., Agathis australis, Albizia amara, Alsophila tricolor, Andropogon spp., Arachis spp, Areca catechu, Astelia fragrans, Astragalus cicer, Baikiaea plurijuga, Betula spp., Brassica spp., Bruguiera gymnorrhiza, Burkea africana, Butea frondosa, Cadaba farinosa, Calliandra spp, Camellia sinensis, Canna indica, Capsicum spp., Cassia spp., Centroema pubescens, Chacoomeles spp., Cinnamomum cassia, Coffea arabica, Colophospermum mopane, Coronillia varia, Cotoneaster serotina, Crataegus spp., Cucumis spp., Cupressus spp., Cyathea dealbata, Cydonia oblonga, Cryptomeria japonica, Cymbopogon spp., Cynthea dealbata, Cydonia oblonga, Dalbergia monetaria, Davallia divaricata, Desmodium spp., Dicksonia squarosa, Dibeteropogon amplectens, Dioclea spp, Dolichos spp., Dorycnium rectum, Echinochloa pyramidalis, Ehraffia spp., Eleusine coracana, Eragrestis spp., Erythrina spp., Eucalyptus spp., Euclea schimperi, Eulalia vi/losa, Pagopyrum spp., Feijoa sellowlana, Fragaria spp., Flemingia spp, Freycinetia banksli, Geranium thunbergii, GinAgo biloba, Glycine javanica, Gliricidia spp, Gossypium hirsutum, Grevillea spp., Guibourtia coleosperma, Hedysarum spp., Hemaffhia alassima, Heteropogon contoffus, Hordeum vulgare, Hyparrhenia rufa, Hypericum erectum, Hypeffhelia dissolute, Indigo incamata, Iris spp., Leptarrhena pyrolifolia, Lespediza spp., Lettuca spp., Leucaena leucocephala, Loudetia simplex, Lotonus bainesli, Lotus spp., Macrotyloma axillare, Malus spp., Manihot esculenta, Medicago saliva, Metasequoia glyptostroboides, Musa sapientum, Nicotianum spp., Onobrychis spp., Ornithopus spp., Oryza spp., Peltophorum africanum, Pennisetum spp., Persea gratissima, Petunia spp., Phaseolus spp., Phoenix canariensis, Phormium cookianum, Photinia spp., Picea glauca, Pinus spp., Pisum sativam, Podocarpus totara, Pogonarthria fleckii, Pogonaffhria squarrosa, Populus spp., Prosopis cineraria, Pseudotsuga menziesii, Pterolobium stellatum, Pyrus communis, Quercus spp., Rhaphiolepsis umbellata, Rhopalostylis sapida, Rhus natalensis, Ribes grossularia, Ribes spp., Robinia pseudoacacia, Rosa spp., Rubus spp., Salix spp., Schyzachyrium sanguineum, Sciadopitys vefficillata, Sequoia sempervirens, Sequoiadendron giganteum, Sorghum bicolor, Spinacia spp., Sporobolus fimbriatus, Stiburus alopecuroides, Stylosanthos humilis, Tadehagi spp, Taxodium distichum, Themeda triandra, Trifolium spp., Triticum spp., Tsuga heterophylla, Vaccinium spp., Vicia spp., Vitis vinifera, Watsonia pyramidata, Zantedeschia aethiopica, Zea mays, amaranth, artichoke, asparagus, broccoli, Brussels sprouts, cabbage, canola, carrot, cauliflower, celery, collard greens, flax, kale, lentil, oilseed rape, okra, onion, potato, rice, soybean, straw, sugar beet, sugar cane, sunflower, tomato, squash tea, maize, wheat, barley, rye, oat, peanut, pea, lentil and alfalfa, cotton, rapeseed, canola, pepper, sunflower, tobacco, eggplant, eucalyptus, a tree, an ornamental plant, a perennial grass and a forage crop. Alternatively algae and other non-Viridiplantae can be used for the methods of the present disclosure.
[0244] According to some embodiments of the disclosure, the plant used by the method of the disclosure is a crop plant including, but not limited to, cotton, Brassica vegetables, oilseed rape, sesame, olive tree, palm oil, banana, wheat, corn or maize, barley, alfalfa, peanuts, sunflowers, rice, oats, sugarcane, soybean, turf grasses, barley, rye, sorghum, sugar cane, chicory, lettuce, tomato, zucchini, bell pepper, eggplant, cucumber, melon, watermelon, beans, hibiscus, okra, apple, rose, strawberry, chili, garlic, pea, lentil, canola, mums, Arabidopsis, broccoli, cabbage, beet, quinoa, spinach, squash, onion, leek, tobacco, potato, sugar beet, papaya, pineapple, mango, Arabidopsis thaliana, and also plants used in horticulture, floriculture or forestry, such as, but not limited to, poplar, fir, eucalyptus, pine, an ornamental plant, a perennial grass and a forage crop, coniferous plants, moss, algae, as well as other plants available on the internet at, for example, wwwdotnationmasterdotcom/encyclopedia/Plantae.
[0245] According to a specific embodiment, the plant is selected from the group consisting of corn, rice, wheat, tomato, cotton and sorghum. In certain embodiments, the plant is a corn plant. In certain embodiments, the plant is a rice plant. In certain embodiments, the plant is a wheat plant. In certain embodiments, the plant is a cotton plant. In certain embodiments, the plant is a sorghum plant.
[0246] Introduction of the compositions of the present disclosure can be performed to any organs/cells of the plant (as opposed to seeds) using conventional delivery methods such as particle bombardment, grafting, soaking and the like.
[0247] According to a specific embodiment, the composition is introduced directly to the seed. According to a specific embodiment, the seed is an uncoated or fresh seed that hasn't been subjected to chemical/physical treatments. In certain embodiments, the seed is a corn seed. In certain embodiments, the seed is a rice seed. In certain embodiments, the seed is a wheat seed. In certain embodiments, the seed is a cotton seed. In certain embodiments, the seed is a sorghum seed.
[0248] The seed may be subjected to priming or washing prior to contacting with the dsRNA.
[0249] In embodiments according to the present disclosure, washing of the seeds is effected for 30 min to 4 hours. In other embodiments, the seeds may be washed up to 5 hours. In an embodiment a seed may be washed up 6, 7 or 8 hours. In another embodiment, seeds are washed for less than 4 or less than 3 hours. In some embodiments, a seed may be washed less than two hours or less than one hour.
[0250] The present disclosure provides for and includes washing of a seed between 1 minute and 1 hours. Also included are brief washes comprising less than 1 minute wherein the seed is completely wet and then removed from the wash. In some embodiments, a seed may be washed from 1 minute to 10 minutes. In another embodiment, a seed may be washed from 1 minute to 10 minutes. In an embodiment, a seed may be washed from 10 to 30 minutes. In yet another embodiment, a seed may be washed from 1 to 30 minutes. In certain embodiments, a seed may be washed from 5 to 10 minutes or 5 to 15 minutes. In some embodiments, a seed may be washed from 15 to 30 minutes or 10 to 25 minutes.
[0251] In some embodiments according to the present disclosure, the wash solution may include a weak detergent such as Tween-20, or its equivalents. In some embodiments, the detergent may be less than 1% by volume. In other embodiments, the detergent may be less than 0.5% by volume. In some embodiments, the detergent may be less than 0.25% or 0.2% by volume. In other embodiments, the detergent may be less than 0.1% or 0.05% by volume. In embodiments according to the present disclosure, the wash solution may contain a detergent at between 0.01 to 0.2% or 0.2 to 1%. In other embodiments, the wash solution may contain a detergent at between 0.05 to 0.5% or 0.5 to 1.5%.
[0252] As used herein the term "priming" refers to controlling the hydration level within seeds so that the metabolic activity necessary for germination can occur but radicle emergence is prevented. Different physiological activities within the seed occur at different moisture levels (Leopold and Vertucci, 1989; Taylor, 1997). The last physiological activity in the germination process is radicle emergence. The initiation of radicle emergence requires a high seed water content. By limiting seed water content, all the metabolic steps necessary for germination can occur without the irreversible act of radicle emergence. Prior to radicle emergence, the seed is considered desiccation tolerant, thus the primed seed moisture content can be decreased by drying. After drying, primed seeds can be stored until time of sowing.
[0253] Several different priming methods are used commercially. Among them, liquid or osmotic priming and solid matrix priming appear to have the greatest following (Khan et al., 1991).
[0254] According to an embodiment of the disclosure, priming is effected in the presence of salt, a chelating agent, polyethylene glycol or a combination of same (e.g., chelating agent and salt).
[0255] Alternatively priming is effected in the presence of water such as deionized water or double deionized water (ddW). According to a specific embodiment, the priming is effected in the presence of 100% ddW.
[0256] Several types of seed priming are commonly used:
[0257] Osmopriming (osmoconditioning) is a standard priming technique. Seeds are incubated in well aerated solutions with a low water potential, and afterwards washes and dried. The low water potential of the solutions can be achieved by adding osmotica like mannitol, polyethyleneglycol (PEG) or salts like KCl. In embodiments according to the present disclosure, the seeds are osmoprimed. In certain embodiments, the osmoprimed seed is a corn seed. In certain embodiments, the osmoprimed seed is a rice seed. In certain embodiments, the osmoprimed seed is a wheat seed. In certain embodiments, the osmoprimed seed is a cotton seed. In certain embodiments, the osmoprimed seed is a sorghum seed.
[0258] Hydropriming (drum priming) is achieved by continuous or successive addition of a limited amount of water to the seeds. A drum is used for this purpose and the water can also be applied by humid air. `On-farm steeping` is a cheap and useful technique that is practiced by incubating seeds (cereals, legumes) for a limited time in warm water. In embodiments according to the present disclosure, the seeds are hydroprimed. In certain embodiments, the hydroprimed seed is a corn seed. In certain embodiments, the hydroprimed seed is a rice seed. In certain embodiments, the hydroprimed seed is a wheat seed. In certain embodiments, the hydroprimed seed is a cotton seed. In certain embodiments, the hydroprimed seed is a sorghum seed.
[0259] Matrixpriming (matriconditioning) is the incubation of seeds in a solid, insoluble matrix (vermiculite, diatomaceous earth, cross-linked highly water-absorbent polymers) with a limited amount of water. This method confers a slow imbibition. In embodiments according to the present disclosure, the seeds are matriconditioned. In certain embodiments, the matriconditioned seed is a corn seed. In certain embodiments, the matriconditioned seed is a rice seed. In certain embodiments, the matriconditioned seed is a wheat seed. In certain embodiments, the matriconditioned seed is a cotton seed. In certain embodiments, the matriconditioned seed is a sorghum seed.
[0260] Pregerminated seeds may be used in certain embodiments however not all species can be primed using this method. In contrast to normal priming, seeds are allowed to perform radicle protrusion. This is followed by sorting for specific stages, a treatment that reinduces desiccation tolerance, and drying. The use of pregerminated seeds causes rapid and uniform seedling development.
[0261] Thus, according to one embodiment, the seeds are primed seeds.
[0262] Of note, it may be possible that the seeds are treated with water (double-distilled water, ddW), prior to contacting with the dsRNA without effecting any priming on the seeds. For instance, treatment for a short while with water (e.g., 30 seconds to 1 hours, 30 seconds to 0.5 hour, 30 seconds to 10 min, 30 seconds to 5 min or 45 seconds to 5 min).
[0263] Thus, according to one embodiment, the seeds are non-primed seeds.
[0264] A non-limiting example of a method of introducing the dsRNA into the seed is provided in Example 1, which is considered as an integral part of the specification.
[0265] The temperature at the washing/priming and drying steps may be the same or differ.
[0266] According to one embodiment, the temperature for washing/priming is between 4 and 28.degree. C. In some embodiments, the washing/priming temperature is less than 28.degree. C. In some embodiments, the washing/priming temperature is less than 25.degree. C. In some embodiments, the washing/priming temperature is less than 20.degree. C. In some embodiments, the washing/priming temperature is less than 15.degree. C. In some embodiments, the washing/priming temperature is less than 10.degree. C. In some embodiments, the washing/priming temperature is between 4 and 10.degree. C. In an embodiment the washing/priming temperature is between 10 and 15.degree. C. In an another embodiment the washing/priming temperature is between 15 and 20.degree. C. or 15 and 25.degree. C. In an another embodiment the washing/priming temperature is between 10 and 20.degree. C. or 10 and 25.degree. C.
[0267] According to one embodiment, the priming/washing solution or the dsRNA containing solution is devoid of a solid carrier.
[0268] According to one embodiment, the priming/washing solution or the dsRNA containing solution is devoid of a transferring agent such as a surfactant or a salt.
[0269] According to a further embodiment of the disclosure, the seeds subject to contacting with the dsRNA molecule are washed in order to remove agents, to which the seeds have been subjected, such as a pesticide, a fungicide, an insecticide, a fertilizer, a coating agent and a coloring agent.
[0270] Thus, according to one embodiment, the seeds (prior to treatment with dsRNA) are substantially free (i.e., do not comprise effective amounts) of pesticide, a fungicide, an insecticide, a fertilizer, a coating agent and a coloring agent.
[0271] The seeds are then subjected to drying.
[0272] According to one embodiment, the drying temperature is between 20 to 37.degree. C. In another embodiment, the drying temperature is between 20 to 30.degree. C. In another embodiment, the drying temperature is between 22 and 37.degree. C. In another embodiment, the drying temperature is between 15 to 22.degree. C. or 20 to 25.degree. C. In embodiments according to the present disclosure, the drying time at a temperature of the present disclosure may be for 10 to 20 hours. In other embodiments the drying time may be from 10 to 16 hours. In other embodiments, the drying time may be 2 to 5 hours.
[0273] Various considerations are to be taken when calculating the concentration of the dsRNA in the contacting solution. Considerations include, but are not limited to, at least one of the group consisting of seed size, seed weight, seed volume, seed surface area, seed density and seed permeability.
[0274] For example, related to seed size, weight, volume and surface area, it is estimated that corn seeds will require longer treatment than Arabidopsis and tomato seeds. Regarding permeability and density, it is estimated that wheat seeds will require longer treatments at higher concentrations than tomato seeds.
[0275] Examples of concentrations of dsRNA in the treating solution include, but are not limited to 0.1 to 100 micrograms (1.times.10.sup.-6 grams) per microliter (1.times.10.sup.-6 liter) (.mu.g/.mu.l). In an embodiment the dsRNA concentration in the treating solution may be 0.04 to 0.15 .mu.g/.mu.l. In an another embodiment the dsRNA concentration in the treating solution may be 0.1 to 50 .mu.g/.mu.l. In certain embodiments, the dsRNA concentration in the treating solution may be 0.1 to 10 .mu.g/.mu.l. In yet other embodiments the dsRNA concentration in the treating solution may be 0.1 to 5 .mu.g/.mu.l. In some embodiments the dsRNA concentration in the treating solution may be 0.1 to 1 .mu.g/.mu.l. In an embodiment the dsRNA concentration in the treating solution may be 0.1 to 0.5 .mu.g/.mu.l. Also included and provided for in the present disclosure are embodiments having a dsRNA concentration in the treating solution of between 0.15 and 0.5 .mu.g/.mu.l. In an embodiment the dsRNA concentration in the treating solution may be 0.1 to 0.3 .mu.g/.mu.l. In an embodiment the dsRNA concentration in the treating solution may be 0.01 to 0.1 .mu.g/.mu.l. In an embodiment the dsRNA concentration in the treating solution may be 0.01 to 0.05 .mu.g/.mu.l. In an embodiment the dsRNA concentration in the treating solution may be 0.02 to 0.04 .mu.g/.mu.l. In an embodiment the dsRNA concentration in the treating solution may be 0.001 to 0.02 1.1 g/.mu.l. According to a specific embodiment, the concentration of the dsRNA in the treating solution is 0.04 to 0.15 .mu.g/.mu.l.
[0276] According to a specific embodiment, the contacting with the dsRNA is effected in the presence of a chelating agent such as EDTA or another chelating agent such as DTPA (0.01 to 0.1 mM).
[0277] The contacting solution may comprise a transferring agent such as a surfactant or a salt.
[0278] Examples of such transferring agents include but are not limited salts such as sodium or lithium salts of fatty acids (such as tallow or tallowamines or phospholipids lipofectamine or lipofectin (1 to 20 nM, or 0.1 to 1 nM)) and organosilicone surfactants. Other useful surfactants include organosilicone surfactants including nonionic organosilicone surfactants, e.g., trisiloxane ethoxylate surfactants or a silicone polyether copolymer such as a copolymer of polyalkylene oxide modified heptamethyl trisiloxane and allyloxypolypropylene glycol methylether (commercially available as Silwet.TM. L-77 surfactant having CAS Number 27306-78-1 and EPA Number: CAL.REG.NO. 5905-50073-AA, currently available from Momentive Performance Materials, Albany, N.Y.).
[0279] Useful physical agents can include (a) abrasives such as carborundum, corundum, sand, calcite, pumice, garnet, and the like, (b) nanoparticles such as carbon nanotubes or (c) a physical force. Carbon nanotubes are disclosed by Kam et al. (2004) J. Am. Chem. Soc., 126 (22):68506851, Liu et al. (2009) Nano Lett., 9(3):1007-1010, and Khodakovskaya et al. (2009) ACS Nano, 3(10):3221-3227. Physical force agents can include heating, chilling, the application of positive pressure, or ultrasound treatment. Agents for laboratory conditioning of a plant to permeation by polynucleotides include, e.g., application of a chemical agent, enzymatic treatment, heating or chilling, treatment with positive or negative pressure, or ultrasound treatment. Agents for conditioning plants in a field include chemical agents such as surfactants and salts.
[0280] Contacting of the seeds with the dsRNA molecule can be effected using any method known in the art as long as a suppressive amount of the dsRNA molecule enters the seeds. These examples include, but are not limited to, soaking, spraying or coating with powder, emulsion, suspension, or solution; similarly, the polynucleotide molecules are applied to the plant by any convenient method, e.g., spraying or wiping a solution, emulsion, or suspension.
[0281] According to a specific embodiment contacting may be effected by soaking (i.e., inoculation) so that shaking the seeds with the treating solution may improve penetration and soaking and therefore reduce treatment time. Shaking is typically performed at 50 to 150 RPM and depends on the volume of the treating solution. Shaking may be effected for 4 to 24 hours (1 to 4 hours, 10 minutes to 1 hour or 30 seconds to 10 minutes). The incubation takes place in the dark at 4 to 28.degree. C. or 15 to 22.degree. C. (e.g., 8 to 15.degree. C., 4 to 8.degree. C., 22 to 28.degree. C.).
[0282] According to a specific embodiment, contacting occurs prior to breaking of seed dormancy and embryo emergence.
[0283] Following contacting, preferably prior to breaking of seed dormancy and embryo emergence, the seeds may be subjected to treatments (e.g., coating) with the above agents (e.g., pesticide, fungicide etc.).
[0284] Contacting is effected such that the dsRNA molecule enters the embryo, endosperm, the coat, or a combination of the three.
[0285] After contacting with the treatment solution, the seeds may be subjected to drying for up to 30 hours at 25 to 37.degree. C.
[0286] According to a specific embodiment, the seed (e.g., isolated seed) comprises the exogenous dsRNA and wherein at least 10 or 20 molecules of the dsRNA are in the endosperm of the isolated seed.
[0287] As used herein the term "isolated" refers to separation from the natural physiological environment. In the case of seed, the isolated seed is separated from other parts of the plant. In the case of a nucleic acid molecule (e.g., dsRNA) separated from the cytoplasm.
[0288] According to a specific embodiment, the dsRNA molecule is not expressed from the plant genome, thereby not being an integral part of the genome.
[0289] Methods of qualifying successful introduction of the dsRNA molecule include but are not limited to, RT-PCR (e.g., quantifying the level of the target gene or the dsRNA), phenotypic analysis such as biomass, vigor, yield and stress tolerance, root architecture, leaf dimensions, grain size and weight, oil content, cellulose, as well as cell biology techniques.
[0290] Seeds may be stored for 1 day to several months prior to planting (e.g., at 4 to 10.degree. C.).
[0291] The resultant seed can be germinated in the dark so as to produce a plant.
[0292] Thus there is provided a plant or plant part comprising an exogenous dsRNA molecule and devoid of a heterologous promoter for driving expression of the dsRNA molecule in the plant.
[0293] As used herein "devoid of a heterologous promoter for driving expression of the dsRNA" means that the plant or plant cell doesn't include a cis-acting regulatory sequence (e.g., heterologous) transcribing the dsRNA in the plant.
[0294] As used herein, the term "heterologous" means not naturally occurring together. In some embodiments, the term "heterologous" refers to exogenous, not-naturally occurring within the native plant cell (such as by position of integration, or being non-naturally found within the plant cell). Thus the isolated seed in the absence of a heterologous promoter sequence for driving expression of the dsRNA in the plant, comprises a homogenic (prior to amplification) or heterogenic (secondary siRNAs, following amplification) population of plant non-transcribable dsRNA. In embodiments according to the present disclosure, an antisense RNA sequence may be a heterologous sequence.
[0295] The present methodology can be used for modulating gene expression such as in a plant, the method comprising: (a) contacting a seed of the plant with a dsRNA, under conditions which allow penetration of the dsRNA into the seed, thereby introducing the dsRNA into the seed; and optionally (b) generating a plant of the seed.
[0296] When used for down-regulating a plant gene, the dsRNA is designed of the desired specificity using bioinformatic tools which are well known in the art (e.g., BLAST).
[0297] This methodology can be used in various applications starting from basic research such as in order to asses gene function and lasting in generating plants with altered traits which have valuable commercial use.
[0298] Such plants can exhibit agricultural beneficial traits including altered morphology, altered flowering, altered tolerance to stress (i.e., biotic and/or abiotic), altered biomass vigor and/or yield and the like.
[0299] The phrase "abiotic stress" as used herein refers to any adverse effect on metabolism, growth, viability and/or reproduction of a plant. Abiotic stress can be induced by any of suboptimal environmental growth conditions such as, for example, water deficit or drought, flooding, freezing, low or high temperature, strong winds, heavy metal toxicity, anaerobiosis, high or low nutrient levels (e.g. nutrient deficiency), high or low salt levels (e.g. salinity), atmospheric pollution, high or low light intensities (e.g. insufficient light) or UV irradiation. Abiotic stress may be a short term effect (e.g. acute effect, e.g. lasting for about a week) or alternatively may be persistent (e.g. chronic effect, e.g. lasting for example 10 days or more). The present disclosure contemplates situations in which there is a single abiotic stress condition or alternatively situations in which two or more abiotic stresses occur.
[0300] According to one embodiment the abiotic stress refers to salinity.
[0301] According to another embodiment the abiotic stress refers to drought.
[0302] According to another embodiment the abiotic stress refers to a temperature stress.
[0303] As used herein the phrase "abiotic stress tolerance" refers to the ability of a plant toendure an abiotic stress without exhibiting substantial physiological or physical damage (e.g. alteration in metabolism, growth, viability and/or reproducibility of the plant).
[0304] As used herein the phrase "nitrogen use efficiency (NUE)" refers to a measure of crop production per unit of nitrogen fertilizer input. Fertilizer use efficiency (FUE) is a measure of NUE. Crop production can be measured by biomass, vigor or yield. The plant's nitrogen use efficiency is typically a result of an alteration in at least one of the uptake, spread, absorbance, accumulation, relocation (within the plant) and use of nitrogen absorbed by the plant. Improved NUE is with respect to that of a non-transgenic plant (i.e., lacking the transgene of the transgenic plant) of the same species and of the same developmental stage and grown under the same conditions.
[0305] As used herein the phrase "nitrogen-limiting conditions" refers to growth conditions which include a level (e.g., concentration) of nitrogen (e.g., ammonium or nitrate) applied which is below the level needed for optimal plant metabolism, growth, reproduction and/or viability.
[0306] As used herein the term/phrase "biomass", "biomass of a plant" or "plant biomass" refers to the amount (e.g., measured in grams of air-dry tissue) of a tissue produced from the plant in a growing season. An increase in plant biomass can be in the whole plant or in parts thereof such as aboveground (e.g. harvestable) parts, vegetative biomass, roots and/or seeds or contents thereof (e.g., oil, starch etc.).
[0307] As used herein the term/phrase "vigor", "vigor of a plant" or "plant vigor" refers to the amount (e.g., measured by weight) of tissue produced by the plant in a given time. Increased vigor could determine or affect the plant yield or the yield per growing time or growing area. In addition, early vigor (e.g. seed and/or seedling) results in improved field stand.
[0308] As used herein the term/phrase "yield", "yield of a plant" or "plant yield" refers to the amount (e.g., as determined by weight or size) or quantity (e.g., numbers) of tissues or organs produced per plant or per growing season. Increased yield of a plant can affect the economic benefit one can obtain from the plant in a certain growing area and/or growing time.
[0309] According to one embodiment the yield is measured by cellulose content, oil content, starch content and the like.
[0310] According to another embodiment the yield is measured by oil content.
[0311] According to another embodiment the yield is measured by protein content.
[0312] According to another embodiment, the yield is measured by seed number, seed weight, fruit number or fruit weight per plant or part thereof (e.g., kernel, bean).
[0313] A plant yield can be affected by various parameters including, but not limited to, plant biomass; plant vigor; plant growth rate; seed yield; seed or grain quantity; seed or grain quality; oil yield; content of oil, starch and/or protein in harvested organs (e.g., seeds or vegetative parts of the plant); number of flowers (e.g. florets) per panicle (e.g. expressed as a ratio of number of filled seeds over number of primary panicles); harvest index; number of plants grown per area; number and size of harvested organs per plant and per area; number of plants per growing area (e.g. density); number of harvested organs in field; total leaf area; carbon assimilation and carbon partitioning (e.g. the distribution/allocation of carbon within the plant); resistance to shade; number of harvestable organs (e.g. seeds), seeds per pod, weight per seed; and modified architecture [such as increase stalk diameter, thickness or improvement of physical properties (e.g. elasticity)].
[0314] Improved plant NUE is translated in the field into either harvesting similar quantities of yield, while implementing less fertilizers, or increased yields gained by implementing the same levels of fertilizers. Thus, improved NUE or FUE has a direct effect on plant yield in the field.
[0315] As used herein "biotic stress" refers stress that occurs as a result of damage done to plants by other living organisms, such as bacteria, viruses, fungi, parasites, beneficial and harmful insects, weeds, and cultivated or native plants. Example 7 of the Examples section which follows, implements the present teachings towards conferring resistance to Spodoptera littoralis.
[0316] As used herein the term "improving" or "increasing" refers to at least about 2%, at least about 3%, at least about 4%, at least about 5%, at least about 10%, at least about 15%, at least about 20%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 90% or greater increase in NUE, in tolerance to stress, in yield, in biomass or in vigor of a plant, as compared to a native or wild-type plants (i.e., isogenic plants (not modified to comprise the dsRNA)) of the disclosure.
[0317] As mentioned the target gene of the dsRNA may not be an endogenous plant gene but rather a gene exogenous to the plant such as of a phytopathogenic organism which feeds on the plant or depends thereon for growth/replication (e.g., bacteria or viruses) and survival.
[0318] As used herein. The term "phytopathogen" refers to an organism that benefits from an interaction with a plant, and has a negative effect on that plant.
[0319] Thus, according to an embodiment of the disclosure there is provided a method of inhibiting expression of a target gene in a phytopathogenic organism, the method comprising providing (e.g., feeding or contacting under infecting conditions) to the phytopathogenic organism the plant as described herein (at least part thereof includes the dsRNA), thereby inhibiting expression of a target gene in the phytopathogenic organism.
[0320] The phytopathogenic organism refers to a multicellular organism e.g., insects, fungi, animals or a microorganism that can cause plant disease, including viruses, bacteria, fungi as well as oomycetes, chytrids, algae, and nematodes.
[0321] Reference herein to a "nematode" refers to a member of the phylum Nematoda. Members of the family Heteroderidae are sedentary parasites that form elaborate permanent associations with the target host organism. They deprive nutrients from cells of an infected organism through a specialized stylet. The cyst nematodes (genera Heterodera and Globodera) and root-knot nematodes (genus Meliodogyne), in particular, cause significant economic loss in plants, especially crop plants. Examples of cyst nematodes include, inter alia, H. avenae (cereal cyst nematodes), H. glycines (beet cyst nematode) and G. pallida (potato cyst nematode). Root-knot nematodes include, for example, M. javanica, M. incognita and M. arenaria. These pathogens establish "feeding sites" in the plant, by causing the morphological transformation of root cells into giant cells. Hence, nematode "infestation" or "infection" refers to invasion of and feeding upon the tissues of the host plant. Other nematodes that cause significant damage include the lesion nematodes such as Pratylenchus, particularly P. penetrans, which infects maize, rice and vegetables, P. brachyurus which infects pineapple and P. thornei which infects inter alia, wheat.
[0322] Insects that may cause damage and disease in plants belong to three categories, according to their method of feeding: chewing, sucking and boring. Major damage is caused by chewing insects that eat plant tissue, such as leaves, flowers, buds and twigs. Examples from this large insect category include beetles and their larvae (grubs), web-worms, bagworms and larvae of moths and sawflies (caterpillars). By comparison, sucking insects insert their mouth parts into the tissues of leaves, twigs, branches, flowers or fruit and suck out the plant's juices. Typical examples of sucking insects include but are not limited to aphids, mealy bugs, thrips and leafhoppers. Damage caused by these pests is often indicated by discoloration, drooping, wilting and general lack of vigor in the affected plant.
[0323] According to a specific embodiment, the phytopathogen is prodentia of the family Noctuidae e.g., Spodoptera littoralis.
[0324] Examples of significant bacterial plant pathogens include, but are not limited to, Burkholderia, Proteobacteria (Xanthomonas spp. and Pseudomonas spp, Pseudomonas syringae pv. tomato).
[0325] A number of virus genera are transmitted, both persistently and non-persistently, by soil borne zoosporic protozoa. These protozoa are not phytopathogenic themselves, but parasitic. Transmission of the virus takes place when they become associated with the plant roots. Examples include Polymyxa graminis, which has been shown to transmit plant viral diseases in cereal crops and Polymyxa betae which transmits Beet necrotic yellow vein virus. Plasmodiophorids also create wounds in the plant's root through which other viruses can enter.
[0326] Specific examples of viruses which can be targeted according to the present teachings includes, but are not limited to:
[0327] (1) Tobacco mosaic virus (TMV, RNA virus) which infects plants, especially tobacco and other members of the family Solanaceae.
[0328] (2) Tomato spotted wilt virus (TSWV, RNA virus) which causes serious diseases of many economically important plants representing 35 plant families, including dicots and monocots. This wide host range of ornamentals, vegetables, and field crops is unique among plant-infecting viruses. Belongs to tospoviruses in the Mediterranean area, affect vegetable crops, especially tomato, pepper and lettuce (Turina et al., 2012, Adv Virus Res 84; 403-437).
[0329] (3) Tomato yellow leaf curl virus (TYLCV) which is transmitted by whitefly, mostly affects tomato plants. Geminiviruses (DNA viruses) in the genus Begomovirus (including sweepoviruses and legumoviruses)--most devastating pathogens affecting a variety of cultivated crops, including cassava, sweet potato, beans, tomato, cotton and grain legumes (Rey et al. 2012, Viruses 4; 1753-1791). Members include TYLCV above and tomato leaf curl virus (ToLCV).
[0330] (4) Cucumber mosaic virus (CMV)--CMV has a wide range of hosts and attacks a great variety of vegetables, ornamentals, and other plants (as many as 191 host species in 40 families). Among the most important vegetables affected by cucumber mosaic are peppers (Capsicum annuum L.), cucurbits, tomatoes (Lycopersicon esculentum Mill.), and bananas (Musa L. spp.).
[0331] Other vegetable hosts include: cucumber, muskmelon, squash, tomato, spinach, celery, peppers, water cress, beet, sweet potato, turnip, chayote, gherkin, watermelon, pumpkin, citron, gourd, lima bean, broad bean, onion, ground-cherry, eggplant, potato, rhubarb, carrot, dill, fennel, parsnip, parsley, luffa, and artichoke (Chabbouh and Cherif, 1990, FAO Plant Prot. Bull. 38:52-53.).
[0332] Ornamental hosts include: China aster, chrysanthemum, delphinium, salvia, geranium, gilia, gladiolus, heliotrope, hyacinth, larkspur, lily, marigold, morning glory, nasturtium, periwinkle, petunia, phlox, snapdragon, tulip, and zinnia (Chupp and Sherf, 1960; Agrios, 1978).
[0333] (5) Potato virus Y (PVY)--one of the most important plant viruses affecting potato production.
[0334] (6) Cauliflower mosaic virus (CaMV, DNA virus (Rothnie et al., 1994)).
[0335] (7) African cassava mosaic virus (ACMV).
[0336] (8) Plum pox virus (PPV) is the most devastating viral disease of stone fruit from the genus Prunus.
[0337] (9) Brome mosaic virus (BMV)--commonly infects Bromus inermis and other grasses, can be found almost anywhere wheat is grown.
[0338] (10) Potato virus X (PVX) There are no insect or fungal vectors for this virus. This virus causes mild or no symptoms in most potato varieties, but when Potato virus Y is present, synergy between these two viruses causes severe symptoms in potato.
[0339] Additional Viruses:
[0340] Citrus tristeza virus (CTV)--causes the most economically damaging disease to Citrus, including sour orange (Citrus aurantium), and any Citrus species grafted onto sour orange root stock, sweet orange (C. sinensis), grapefruit (C. paradisi), lime and Seville orange (C. aurantifolia), and mandarin (C. reticulata). CTV is also known to infect Aeglopsis chevalieri, Afraegle paniculata, Pamburus missionis, and Passiflora gracilis. CTV is distributed worldwide and can be found wherever citrus trees grow.
[0341] Barley yellow dwarf virus (BYDV)--most widely distributed viral disease of cereals. It affects the economically important crop species barley, oats, wheat, maize, triticale and rice.
[0342] Potato leafroll virus (PLRV) infects potatoes and other members of the family Solanaceae.
[0343] Tomato bushy stunt virus (TBSV), RNA virus, a member of the genus Tombusvirus and mostly affects tomatoes and eggplant.
[0344] Additional Reviews:
[0345] Hamilton et al., 1981, J Gen Virol 54; 223-241--mentions TMV, PVX, PVY, CMV, CaMV
[0346] Additional Scientific Papers:
[0347] Makkouk et al., 2012, Adv Virus Res 84; 367-402--Viruses affecting peas and beans with narrow (Faba bean necrotic yellow virus (FBNYN)) and wide (alfalfa mosaic virus (AMV) and CMV) host range.
[0348] Insect pests causing plant disease include those from the families of, for example, Apidae, Curculionidae, Scarabaeidae, Tephritidae, Tortricidae, amongst others.
[0349] The target gene of the phytopathogenic organism encodes a product essential to the viability and/or infectivity of the pathogen, therefore its down-regulation (by the dsRNA) results in a reduced capability of the pathogen to survive and infect host cells. Hence, such down-regulation results in a "deleterious effect" on the maintenance viability and/or infectivity of the phytopathogen, in that it prevents or reduces the pathogen's ability to feed off and survive on nutrients derived from host cells. By virtue of this reduction in the phytopathogen's viability and/or infectivity, resistance and/or enhanced tolerance to infection by a pathogen is facilitated in the cells of the plant. Genes in the pathogen may be targeted at the mature (adult), immature (juvenile) or embryo stages.
[0350] Examples of genes essential to the viability and/or infectivity of the pathogen are provided herein. Such genes may include genes involved in development and reproduction, e.g. transcription factors (see, e.g. Xue et al., 1993; Finney et al., 1988), cell cycle regulators such as wee-1 and ncc-1 proteins (see, e.g. Wilson et al., 1999; Boxem et al., 1999) and embryo-lethal mutants (see, e.g. Schnabel et al., 1991); proteins required for modeling such as collagen, ChR3 and LRP-1 (see, e.g. Yochem et al., 1999; Kostrouchova et al., 1998; Ray et al., 1989); genes encoding proteins involved in the motility/nervous system, e.g. acetycholinesterase (see, e.g. Piotee et al., 1999; Talesa et al., 1995; Arpagaus et al., 1998), ryanodine receptor such as unc-68 (see, e.g. Maryon et al., 1998; Maryon et al., 1996) and glutamate-gated chloride channels or the avermeetin receptor (see, e.g., Cully et al., 1994; Vassilatis et al., 1997; Dent et al., 1997); hydrolytic enzymes required for deriving nutrition from the host, e.g. serine proteinases such as HGSP-1 and HGSP-III (see, e.g. Lilley et al., 1997); parasitic genes encoding proteins required for invasion and establishment of the feeding site, e.g. cellulases (see, e.g. de Boer et al., 1999; Rosso et al., 1999) and genes encoding proteins that direct production of stylar or amphidial secretions such as sec-1 protein (see, e.g. Ray et al., 1994; Ding et al., 1998); genes encoding proteins required for sex or female determination, e.g. tra-1, tra-2 and egl-1, a suppressor of ced9 (see, e.g. Hodgkin, 1980; Hodgkin, 1977; Hodgkin, 1999; Gumienny et al., 1999; Zarkower et al., 1992); and genes encoding proteins required for maintenance of normal metabolic function and homeostasis, e.g. sterol metabolism, embryo lethal mutants (see, e.g. Schnabel et al., 1991) and trans-spliced leader sequences (see, e.g. Ferguson et al, 1996), pos-1, cytoplasmic Zn finger protein; pie-1, cytoplasmic Zn finger protein; mei-1, ATPase; dif-1, mitochondrial energy transfer protein; rba-2, chromatin assembly factor; skn-1, transcription factor; plk-1, kinase; gpb-1, G-protein B subunit; par-1, kinase; bir-1, inhibitor of apoptosis; mex-3, RNA-binding protein, unc-37, G-protein B subunit; hlh-2, transcription factor; par-2, dnc-1, dynactin; par-6, dhc-1, dynein heavy chain; and pal-1, homeobox. Such genes have been cloned from parasitic nematodes such as Meliodogyne and Heterodera species or can be identified by one of skill in the art using sequence information from cloned C. elegans orthologs (the genome of C. elegans has been sequenced and is available, see The C. elegans Sequencing Consortium (1998)).
[0351] As used herein, a "pathogen resistance" trait is a characteristic of a plant that causes the plant host to be resistant to attack from a pathogen that typically is capable of inflicting damage or loss to the plant. Once the phytopathogen is provided with the plant material comprising the dsRNA, expression of the gene within the target pathogen is suppressed by the dsRNA, and the suppression of expression of the gene in the target pathogen results in the plant being resistant to the pathogen.
[0352] In this case, the target gene can encode an essential protein or transcribe an non-coding RNA which, the predicted function is for example selected from the group consisting of ion regulation and transport, enzyme synthesis, maintenance of cell membrane potential, amino acid biosynthesis, amino acid degradation, development and differentiation, infection, penetration, development of appressoria or haustoria, mycelial growth, melanin synthesis, toxin synthesis, siderophore synthesis, sporulation, fruiting body synthesis, cell division, energy metabolism, respiration, and apoptosis, among others.
[0353] According to a specific embodiment, the phytopathogenic organism is selected from the group consisting of a fungus, a nematode, a virus, a bacteria and an insect.
[0354] To substantiate the anti-pest activity, the present teachings also contemplate observing death or growth inhibition and the degree of host symptomatology following said providing.
[0355] To improve the anti-phytopathogen activity, embodiments of the present disclosure further provide a composition that contains two or more different agents each toxic to the same plant pathogenic microorganism, at least one of which comprises a dsRNA described herein. In certain embodiments, the second agent can be an agent selected from the group consisting of inhibitors of metabolic enzymes involved in amino acid or carbohydrate synthesis; inhibitors of cell division; cell wall synthesis inhibitors; inhibitors of DNA or RNA synthesis, gyrase inhibitors, tubulin assembly inhibitors, inhibitors of ATP synthesis; oxidative phosphorylation uncouplers; inhibitors of protein synthesis; MAP kinase inhibitors; lipid synthesis or oxidation inhibitors; sterol synthesis inhibitors; and melanin synthesis inhibitors.
[0356] In addition, plants generated according to the teachings of the present disclosure or parts thereof can exhibit altered nutritional or therapeutic efficacy and as such can be employed in the food or feed and drug industries. Likewise, the plants generated according to the teachings of the present disclosure or parts thereof can exhibit altered oil or cellulose content and as such can be implemented in the construction or oil industry.
[0357] The seeds of the present disclosure can be packed in a seed containing device which comprises a plurality of seeds at least some of which (e.g., 5%, 10% or more) containing an exogenous dsRNA, wherein the seed is devoid of a heterologous promoter for driving expression of the dsRNA.
[0358] The seed containing device can be a bag, a plastic bag, a paper bag, a soft shell container or a hard shell container.
[0359] Reagents of the present disclosure can be packed in a kit including the dsRNA, instructions for introducing the dsRNA into the seeds and optionally a priming solution. According to one embodiment, the dsRNA and priming solution are comprised in separate containers.
[0360] Compositions of some embodiments of the disclosure may, if desired, be presented in a pack or dispenser device which may contain one or more unit dosage forms containing the active ingredient. The pack may, for example, comprise metal or plastic foil, such as a blister pack. The pack or dispenser device may be accompanied by instructions for introduction to the seed.
[0361] As used herein the term "about" refers to .+-.10%.
[0362] The terms "comprises", "comprising", "includes", "including", "having" and their conjugates mean "including but not limited to".
[0363] The term "consisting of" means "including and limited to".
[0364] The term "consisting essentially of" means that the composition, method or structure may include additional ingredients, steps and/or parts, but only if the additional ingredients, steps and/or parts do not materially alter the basic and novel characteristics of the claimed composition, method or structure.
[0365] As used herein, the singular form "a", "an" and "the" include plural references unless the context clearly dictates otherwise. For example, the term "a compound" or "at least one compound" may include a plurality of compounds, including mixtures thereof.
[0366] Throughout this application, various embodiments of this disclosure may be presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the disclosure. Accordingly, the description of a range should be considered to have specifically disclosed all the possible subranges as well as individual numerical values within that range. For example, description of a range such as from 1 to 6 should be considered to have specifically disclosed subranges such as from 1 to 3, from 1 to 4, from 1 to 5, from 2 to 4, from 2 to 6, from 3 to 6 etc., as well as individual numbers within that range, for example, 1, 2, 3, 4, 5, and 6. This applies regardless of the breadth of the range.
[0367] Whenever a numerical range is indicated herein, it is meant to include any cited numeral (fractional or integral) within the indicated range. The phrases "ranging/ranges between" a first indicate number and a second indicate number and "ranging/ranges from" a first indicate number "to" a second indicate number are used herein interchangeably and are meant to include the first and second indicated numbers and all the fractional and integral numerals there between.
[0368] As used herein the term "method" refers to manners, means, techniques and procedures for accomplishing a given task including, but not limited to, those manners, means, techniques and procedures either known to, or readily developed from known manners, means, techniques and procedures by practitioners of the chemical, pharmacological, biological, biochemical and medical arts.
[0369] It is appreciated that certain features of the disclosure, which are, for clarity, described in the context of separate embodiments, may also be provided in combination in a single embodiment. Conversely, various features of the disclosure, which are, for brevity, described in the context of a single embodiment, may also be provided separately or in any suitable subcombination or as suitable in any other described embodiment of the disclosure. Certain features described in the context of various embodiments are not to be considered essential features of those embodiments, unless the embodiment is inoperative without those elements.
[0370] Various embodiments and embodiments of the present disclosure as delineated hereinabove and as claimed in the claims section below find experimental support in the following examples.
EXAMPLES
[0371] Reference is now made to the following examples, which together with the above descriptions, illustrate some embodiments of the disclosure in a non limiting fashion.
[0372] Generally, the nomenclature used herein and the laboratory procedures utilized in the present disclosure include molecular, biochemical, microbiological and recombinant DNA techniques. Such techniques are thoroughly explained in the literature. See, for example, "Molecular Cloning: A laboratory Manual" Sambrook et al., (1989); "Current Protocols in Molecular Biology" Volumes I-III Ausubel, R. M., ed. (1994); Ausubel et al., "Current Protocols in Molecular Biology", John Wiley and Sons, Baltimore, Md. (1989); Perbal, "A Practical Guide to Molecular Cloning", John Wiley & Sons, New York (1988); Watson et al., "Recombinant DNA", Scientific American Books, New York; Birren et al. (eds) "Genome Analysis: A Laboratory Manual Series", Vols. 1-4, Cold Spring Harbor Laboratory Press, New York (1998); methodologies as set forth in U.S. Pat. Nos. 4,666,828; 4,683,202; 4,801,531; 5,192,659 and 5,272,057; "Cell Biology: A Laboratory Handbook", Volumes I-III Cellis, J. E., ed. (1994); "Culture of Animal Cells--A Manual of Basic Technique" by Freshney, Wiley-Liss, N.Y. (1994), Third Edition; "Current Protocols in Immunology" Volumes I-III Coligan J. E., ed. (1994); Stites et al. (eds), "Basic and Clinical Immunology" (8th Edition), Appleton & Lange, Norwalk, Conn. (1994); Mishell and Shiigi (eds), "Selected Methods in Cellular Immunology", W. H. Freeman and Co., New York (1980); available immunoassays are extensively described in the patent and scientific literature, see, for example, U.S. Pat. Nos. 3,791,932; 3,839,153; 3,850,752; 3,850,578; 3,853,987; 3,867,517; 3,879,262; 3,901,654; 3,935,074; 3,984,533; 3,996,345; 4,034,074; 4,098,876; 4,879,219; 5,011,771 and 5,281,521; "Oligonucleotide Synthesis" Gait, M. J., ed. (1984); "Nucleic Acid Hybridization" Hames, B. D., and Higgins S. J., eds. (1985); "Transcription and Translation" Hames, B. D., and Higgins S. J., eds. (1984); "Animal Cell Culture" Freshney, R. I., ed. (1986); "Immobilized Cells and Enzymes" IRL Press, (1986); "A Practical Guide to Molecular Cloning" Perbal, B., (1984) and "Methods in Enzymology" Vol. 1317, Academic Press; "PCR Protocols: A Guide To Methods And Applications", Academic Press, San Diego, Calif. (1990); Marshak et al., "Strategies for Protein Purification and Characterization--A Laboratory Course Manual" CSHL Press (1996); all of which are incorporated by reference as if fully set forth herein. Other general references are provided throughout this document. The procedures therein are believed to be well known in the art and are provided for the convenience of the reader. All the information contained therein is incorporated herein by reference.
Example 1: Protocols for dsRNA Production and Seed Treatment
[0373] Generating dsRNA/siRNA Sequences
[0374] The dsRNA sequences were custom-created for each gene using in vitro transcription of PCR products. Part of the mRNA, including either the ORF, 3' UTR or 5' UTR for which dsRNA to be produced was PCR-amplified using gene-specific primers, which contain the sequence of the T7 promoter on either side. This product was used as a template for dsRNA production using commercial kits such as the MaxiScript dsRNA kit (Life Technologies) or T7 High Yield RNA Synthesis kit (NEB). Next, the sample is treated with DNase Turbo at 37.degree. C. for 15-30 min followed by phenol treatment and nucleic acid precipitation. Next, one of two different reactions is carried out: (1) dsRNA is ready to use, (2) processing of the dsRNA with Dicer (Shortcut RNase III (NEB)) to create small interfering RNAs (siRNA).
[0375] Either dsRNA or a combination of dsRNA and siRNA were used for seed treatments as described below.
[0376] General Seed Treatment Protocol for Gene Silencing Using a dsRNA/siRNA Mixture
[0377] Uncoated organic corn seeds were from variety "popcorn", uncoated organic whole grain rice seeds, organic soybean and wheat seeds were purchased from Nitsat Haduvdevan (Israel), Fresh tomato seeds were retrieved from M82 tomato fruits, which are propagated in-house. Uncoated or fresh plant seeds were washed with double distilled water (DDW) prior to treatment for four hours. Next, seeds were dried at 30.degree. C. for 10-16 hours. Following the drying step, seeds were treated with a solution containing the dsRNA formulation, which is made of dsRNA at a final concentration of 40-150 .mu.g/ml in 0.1 mM EDTA. Treatment was performed by gently shaking the seeds in the solution for 24 hours in a dark growth chamber at 15.degree. C. Finally, seeds were washed twice briefly and planted on soil or dried for 0-30 hours and germinated at 25.degree. C. in a dark growth chamber and planted in soil or planted directly in soil. Control seeds were treated in a similar way, with a formulation that lacked the dsRNA or with non-specific dsRNA.
Example 2: Stability of the dsRNA in Seedlings of Rice, Tomato and Sorghum
[0378] As an example for an exogenous gene that is not present/expressed in plants, the ORFs encoding the replicase and coat protein of CGMMV (accession number AF417242) were used to as targets for dsRNA treatment of plant seeds using the protocol described in Example 1. Rice, tomato and sorghum seeds were washed for 4 hours at 20.degree. C., tomato and sorghum were dried at 30.degree. C. and rice at 20.degree. C. for overnight. Seeds were immediately treated at 15.degree. C. with 132.7 .mu.g/ml dsRNA (final concentration) for 39 hours for rice, 93.8 .mu.g/ml dsRNA (final concentration) for 48 hours for tomato, and 75 .mu.g/ml dsRNA (final concentration) for 40 hours for sorghum.
[0379] Briefly, the virus-derived ORFs were amplified by PCR with specifically designed forward and reverse primers that contain the T7 sequence (5'-TAATACGACTCACTATAGGG-3', SEQ ID NO: 1) at their 5' (see Table 2, below). PCR products were purified from agarose gel and since they carry T7 promoters at both ends they were used as templates for T7-dependent in-vitro transcription, resulting in dsRNA product of the CGMMV genes. PCR on a housekeeping gene, tubulin, was used as a positive control (forward primer 5'-GGTGCTCTGAACGTGGATG-3' (SEQ ID NO: 2), and reverse primer 5'-CATCATCGCCATCCTCATTCTC-3'(SEQ ID NO: 3)).
TABLE-US-00003 TABLE 2 PCR primers served as Templates for in vitro Transcription and detection of CGMMV, and CGMMV dsRNA products. Forward Reverse Virus Product primer/SEQ ID primer/SEQ Name Name Product Sequence/SEQ ID NO: NO: ID NO: 1) CGMMV CGMVV TAATACGACTCACTATAGGGGGTAAGCG TAATACGACT Set 1: (NCBI dsRNA GCATTCTAAACCTCCAAATCGGAGGTTGG CACTATAGGG TAATACGA Accession product 1 ACTCTGCTTCTGAAGAGTCCAGTTCTGTT GGTAAGCGGC CTCACTATA number TCTTTTGAAGATGGCTTACAATCCGATCA ATTCTAAACC/ GGGGAAGA AF417242) CACCTAGCAAACTTATTGCGTTTAGTGCT (SEQ ID NO: 5) CCCTCGAA TCTTATGTTCCCGTCAGGACTTTACTTAAT CTTCTTATGT ACTAAGC/ TTTCTAGTTGCTTCACAAGGTACCGCTTTC TCCCGTCAGG/ (SEQ ID NO: 4) CAGACTCAAGCGGGAAGAGATTCTTTCCG (SEQ ID NO: 7) Set 2: CGAGTCCCTGTCTGCGTTACCCTCGTCTG ACTCAGCA TCGTAGATATTAATTCTAGATTCCCAGAT GTCGTAGG GCGGGTTTTTACGCTTTCCTCAACGGTCC ATTG/(SEQ TGTGTTGAGGCCTATCTTCGTTTCGCTTCT ID NO: 6) CAGCTCCACGGATACGCGTAATAGGGTC ATTGAGGTTGTAGATCCTAGCAATCCTAC GACTGCTGAGTCGCTTAACGCCGTAAAGC GTACTGATGACGCGTCTACGGCCGCTAGG GCTGAGATAGATAATTTAATAGAGTCTAT TTCTAAGGGTTTTGATGTTTACGATAGGG CTTCATTTGAAGCCGCGTTTTCGGTAGTC TGGTCAGAGGCTACCACCTCGAAAGCTTA GTTTCGAGGGTCTTCCCCTATAGTGAGTC GTATTA/(SEQ ID NO: 8) CGMVV TAATACGACTCACTATAGGGGCTTTACCG TAATACGACT Set 3: dsRNA CCACTAAGAACTCTGTACACTCCCTTGCG CACTATAGGG TAATACGA product 2 GGTGGTCTGAGGCTTCTTGAATTGGAATA GCTTTACCGC CTCACTATA TATGATGATGCAAGTGCCCTACGGCTCAC CACTAAGAAC/ GGGCATCA CTTGTTATGACATCGGCGGTAACTATACG (SEQ ID CCATCGAC CAGCACTTGTTCAAAGGTAGATCATATGT NO: 10) CCTAAAC/ GCATTGCTGCAATCCGTGCCTAGATCTTA (SEQ ID AAGATGTTGCGAGGAATGTGATGTACAA NO: 9) CGATATGATCACGCAACATGTACAGAGG CACAAGGGATCTGGCGGGTGCAGACCTC TTCCAACTTTCCAGATAGATGCATTCAGG AGGTACGATAGTTCTCCCTGTGCGGTCAC CTGTTCAGACGTTTTCCAAGAGTGTTCCT ATGATTTTGGGAGTGGTAGGGATAATCAT GCAGTCTCGTTGCATTCAATCTACGATAT CCCTTATTCTTCGATCGGACCTGCTCTTCA TAGGAAAAATGTGCGAGTTTGTTATGCAG CCTTTCATTTCTCGGAGGCATTGCTTTTAG GTTCGCCTGTAGGTAATTTAAATAGTATT GGGGCTCAGTTTAGGGTCGATGGTGATGC CCTATAGTGAGTCGTATTA/(SEQ ID NO: 11)
[0380] dsRNA homologous to green mottle mosaic virus was observed to be stable in rice seedlings. Rice seeds were treated at 15.degree. C. with 132.7 .mu.g/ml dsRNA (final concentration) for 39 hours and dsRNA was detected_by real time polymerase chain reaction (RT-PCR) 1 week post germination. Detection of tubulin cDNA serves as a positive control for the cDNA quality. At two weeks post germination, dsRNA was detectable in 10 out of 10 seedlings. At 3 weeks post germination, dsRNA homologous to green mottle mosaic virus was detected in 5 out of 5 samples of rice seedlings.
[0381] Tomato seeds were treated at 15.degree. C. with 93.8 .mu.g/ml dsRNA (final concentration) for 48 hours and sorghum seeds treated at 5 .mu.g/ml dsRNA (final concentration) for 40 hours. CGMMV dsRNA was detected by RT-PCR in 5 out of 13 tomato seedlings tested at 10 day post-germination and 3 out of four sorghum seedlings 4 weeks after germination.
[0382] The exogenous dsRNA was found to be stable for at least three weeks in rice seedlings and at least 10 days in tomato seedlings and four weeks in Sorghum plants.
Example 3: The dsRNA is not Integrated into the Genome of Rice
[0383] Rice seeds were treated with an exogenous dsRNA as in Example 2. Plants were germinated and grown for five weeks, DNA was extracted and PCR reactions were performed to demonstrate that the dsRNA did not integrate into the Rice's genome. Two sets of primers that gave a positive reaction when checked on the RNA level were used, set 1 (see Table 2) of primers were the set of primers used to amplify the template (all the dsRNA sequence). Set 2 (see Table 3) are the primers that were used in the PCR above. A Rice endogenous housekeeping gene (tubulin) was used as a positive control for the PCR reaction (see Table 3).
[0384] Three different DNA PCR reactions were carried out on dsRNA treated and untreated plants. No amplified DNA corresponding to CGMMV was detected in any treated or untreated plant.
TABLE-US-00004 TABLE 3 Tubulin Primers Used for PCR Amplification. Primer Name and Primer Sequence/ Primer Direction (SEQ ID NO:) Length osa_TubA1_736F GGTGCTCTGAACGTGGATG (SEQ 19 ID NO: 12) osa_TubA1_1342R CATCATCGCCATCCTCATTCTC 22 (SEQ ID NO: 13)
Example 4: Exogenous dsRNA Molecules are Highly Stable in Solution and do not Get Incorporated into the Genome of Treated Plants
[0385] Corn seeds were treated using the protocol described in Example 1, seeds were washed for 4 h at 20.degree. C., dried at 30.degree. C. overnight and immediately treated with 40 .mu.g/ml dsRNA (final concentration) directed against the 13-glucuronidase (GUS) reporter gene for 60 hours at 15.degree. C., dried and were germinated. Leaves and roots were harvested from control and dsGUS-treated plants 7 and 15 days following germination. RNA was extracted from the harvested tissues and RT-PCR with specific GUS primers was run (Table 4). In addition, a corn endogenous housekeeping gene (ubiquitin) was used as a positive control for the PCR reaction. The GUS dsRNA molecules were found to be extremely stable in the treated seeds, and can be detected in corn plants 7 and 15 days post germination of the seeds.
[0386] GUS dsRNA was detected in corn seedlings by RT-PCR at 7 and 15 days after germination according to an aspect of the present disclosure. At one week, GUS dsRNA was detected in shoots of nine of eleven corn seedlings tested. GUS dsRNA was not detected in untreated plants. At 1 week post-germination, GUS dsRNA was detected in five of five corn seedlings' roots. At 15 days post germination, GUS dsRNA was detected in corn seedlings' roots. GUS dsRNA molecules do not get incorporated in the genome of treated corn plants one week after germination as determined by agarose gel electrophoresis of DNA PCR reactions using GUS primers on DNA isolated from treated corn plants.
TABLE-US-00005 TABLE 4 Primers for PCR Amplification of GUS and Ubiquitin Genes and GUS dsRNA product. Primer Primer Length Primer Sequence/SEQ ID NO: Name GUS T7_For TAATACGACTCACTATAGGGAGATCGACGGCCTGTGGGCATT C /(SEQ ID NO: 15) GUS T7_Rev TAATACGACTCACTATAGGGAGCATTCCCGGCGGGATAGTCT 43 G /(SEQ ID NO: 16) GUS208For CAGCGCGAAGTCTTTATACC/(SEQ ID NO: 17) 43 GUS289Rev CTTTGCCGTAATGAGTGACC/(SEQ ID NO: 18) 20 zmaUBQ-947F CCATAACCCTGGAGGTTGAG/(SEQ ID NO: 19) 20 zmaUBQ1043R ATCAGACGCTGCTGGTCTGG/(SEQ ID NO: 20) 20 GUS dsRNA TAATACGACTCACTATAGGGAGATCGACGGCCTGTGGGCATTC product AGTCTGGATCGCGAAAACTGTGGAATTGATCAGCGTTGGTGG GAAAGCGCGTTACAAGAAAGCCGGGCTATTGCTGTGCCAGGC AGTTTTAACGATCAGTTCGCCGATGCAGATATTCGTAATTATG CGGGCAACGTCTGGTATCAGCGCGAAGTCTTTATACCGAAAG GTTGGGCAGGCCAGCGTATCGTGCTGCGTTTCGATGCGGTCAC TCATTACGGCAAAGTGTGGGTCAATAATCAGGAAGTGATGGA GCATCAGGGCGGCTATACGCCATTTGAAGCCGATGTCACGCC GTATGTTATTGCCGGGAAAAGTGTACGTATCACCGTTTGTGTG AACAACGAACTGAACTGGCAGACTATCCCGCCGGGAATGCTC CCTATAGTGAGTCGTATTA/(SEQ ID NO: 21)
Example 5: Fluorescence Microscopy of siRNA Sequences in Various Plant Seeds
[0387] Plant seeds as per the protocol described in example 1. Seeds were washed for 4 h at 20.degree. C., dried at 25.degree. C. and were immediately treated with a fluorescent siRNA (siGLO, 2 .mu.M final concentration, Thermo Scientific) at 15.degree. C. for 24 h. The quality of the siGLO before application to a plant seed was verified by gel electrophoresis analysis. Bands corresponding to the expected size of 20-24 bp of the fluorescent siRNA molecules was detected.
[0388] Fluorescent pictures of the seeds were taken 24-48 hours post treatment using an Olympus microscope at the lowest objective magnification (5.times. for bigger seeds such as rice and tomato seeds, and 10.times. for smaller seeds such as Arabidopsis seeds). To eliminate the possibility of non-specific auto-fluorescence, dsRNA-treated seeds were compared to control untreated seeds. Penetration of fluorescent siRNA molecules into plant seeds was observed at 24 hours after seed treatment with siRNA at 2 .mu.M final concentration in rice seeds and tomato seeds.
[0389] Penetration of fluorescent siRNA molecules into rice seeds was observed at 24 hours following treatment with siGLO dsRNA.
[0390] In order to evaluate the distribution efficiency of the fluorescent siRNA inside the seeds, different plant seeds were cut into slices and imaged with a fluorescent microscope 48 hours after treatment. Each treated seed was imaged alongside a control untreated seed. Light and fluorescent images were taken where applicable for rice, tomato, cucumber, bean, sorghum and wheat seed samples.
[0391] Penetration of fluorescent siRNA molecules into rice seeds was observed at 48 hours following treatment with siGLO dsRNA. siGLO-treated and control rice seeds were sliced to view the interior distribution of the fluorescent dsRNA using a fluorescent microscope and fluorescent siRNA molecules detected in the treated seed. Fluorescent siGLO RNA is detected throughout the seed.
[0392] Penetration of fluorescent siRNA molecules into tomato seeds was observed at 48 hours following treatment with siGLO dsRNA. siGLO-treated and control tomato seeds were sliced to view the interior distribution of the fluorescent dsRNA using a fluorescent microscope. Fluorescent siGLO RNA is detected in the endosperm and the embryo.
[0393] Penetration of fluorescent siRNA molecules into cucumber seeds was observed at 48 hours following treatment with siGLO dsRNA. siGLO-treated and control cucumber seeds were sliced to view the interior distribution of the florescent dsRNA using a fluorescent microscope.
[0394] Penetration of fluorescent siRNA molecules into cucumber seeds was observed at Fluorescent siGLO RNA is detected in the endosperm and the embryo.
[0395] Penetration of fluorescent siRNA molecules is detected in sliced seeds of various plant species, including bean, tomato, sorghum and wheat, 48 hours following treatment with siGLO dsRNA. siGLO-treated and control seeds were sliced to view the interior distribution of the fluorescent dsRNA using a fluorescent microscope. Light images were also taken for each seed and are shown alongside the fluorescent image of the seed for reference.
[0396] FIG. 1 presents fluorescent images of siGLO-treatment rice seeds over a 24 hour period. The effect of incubation time with siGLO dsRNA on fluorescence intensity, indicating quantity and quality of dsRNA penetration, was tested. Control seeds that were left untreated (1), were imaged along with seeds treated with siGLO dsRNA for four different incubation times; 10 min (2), 3.5 hours (3), 5.5 hours (4), and 24 hours (5).
[0397] It is clear that the siRNA is distributed at various levels between the embryo and the endosperm. Accordingly, dsRNA molecules enter the embryo directly. Though not to be limited by any particular theory, the dsRNA molecules are carried by the water-based solution used for the seed treatment. The dsRNA molecules enter the endosperm as part of the endosperm's water-absorption process. These molecules then are transferred to the embryo as it develops as part of the endosperm to embryo nutrient flow during germination and seed development.
[0398] These present findings suggest the RNA molecules used to treat the seeds both penetrate the embryo and function in the embryo as it develops and also penetrate the endosperm and feed the embryo following germination.
Example 6: Time Course Experiment with siGLO Treatment
[0399] A time course experiment was performed on rice seeds to monitor the kinetics of siGLO penetration into the seeds following the seed treatment (FIG. 1). The results indicate that the siRNA efficiently penetrates the plant seeds using the protocol described in Example 1.
Example 7: Example Embodiments of dsRNA Molecules
[0400] Example 7A provides A backbone sequence with two smRNA complementary sites and a helicase binding site:
TABLE-US-00006 (SEQ ID NO: 14) 5' GCATCCTCATCTTAATCTCGGTGCTATCCTACCTGAGCTTGATATCT AGGCGAAGCAGCCCGAATGCTGCACCCTAGATGGCGAAAGTCCAGTAGC GATATCGAATTCCTCGAGGGATCCAAGCTTCCTTGTCTATCCCTCCTGAG CTGTTGATTTTATTCCATGT 3'.
This example contains a sequence for mutated microRNA 390 binding (bold), followed by a helicase binding site (bold and underlined) and a microRNA 390 binding sequence (underlined). DNA sequences for restriction enzyme recognition are added for cloning of the sequence to be silenced.
[0401] Example 7B is the same as example 7A, without the helicase binding site:
TABLE-US-00007 (SEQ ID NO: 22) GCATCCTCATCTTAATCTCGGTGCTATCCTACCTGAGCTTGATATCGATA TCGAATTCCTCGAGGGATCCAAGCTTCCTTGTCTATCCCTCCTGAGCTGT TGATTTTATTCCATGT.
[0402] Example 7C provides a backbone sequence with two smRNA complementary sites and an helicase binding site:
TABLE-US-00008 (SEQ ID NO: 23) GCATCCTCATCTTAATCTCGTGATTTTTCTCTACAAGCGAAGATATCTAG GCGAAGCAGCCCGAATGCTGCACCCTAGATGGCGAAAGTCCAGTAGCGAT ATCGAATTCCTCGAGGGATCCAAGCTTTCTTGCTCAAATGAGTATTCCAG TTGATTTTATTCCATGT.
Example 7C contains a sequence for microRNA 173 binding (bold), followed by a helicase binding site (bold and underlined) and the reverse-complement sequence of microRNA 828 binding sequence (underlined). DNA sequences for restriction enzyme recognition are added for cloning of the sequence to be silenced. In this case a single complementary site is sufficient (i.e., miR173BS) yet a second complementary site is placed on the complementary strand (i.e., miR828BS) so as to enhance amplification from both strands.
[0403] Example 7D is the same as Example 7C, without the helicase binding site:
TABLE-US-00009 (SEQ ID NO: 24) GCATCCTCATCTTAATCTCGTGATTTTTCTCTACAAGCGAAGATATCGAT ATCGAATTCCTCGAGGGATCCAAGCTTTCTTGCTCAAATGAGTATTCCAG TTGATTTTATTCCATGT.
Example 8: Schematic Representation of the Solanum Lycopersicum TAS3 Gene
[0404] FIG. 2A presents a schematic representation of the Solanum Lycopersicum (Lycopersicon esculentum) TAS3 gene. This gene contains two Mir390 binding sites (BS). The 5' Mir390BS has mutations in critical positions for Mir390 dependent cleavage and therefore it is bound by Mir390 but not cleaved (will be referred hereafter as 5' Mut Mir390BS). The 3' Mir390 does lead to Mir390 binding and cleavage. In between these two sequences there is a 234 bp sequence that contains all the different ta-siRNAs that will be created following RDRP recruitment, RNA dependent RNA polymerization and dicing (Allen et al. (2005). MicroRNA-Directed Phasing during Trans-Acting siRNA Biogenesis in Plants. Cell, 121, 207-221., Axtell et al. (2006). A Two-Hit Trigger for siRNA Biogenesis in Plants. Cell, 127, 565-577., Montgomery et al. (2008). Specificity of ARGONAUTE7-miR390 Interaction and Dual Functionality in TAS3 Trans-Acting siRNA Formation. Cell, 133, 128-141).
Example 9: Additional dsRNA Constructs According to the Present Disclosure
[0405] Example 9A provides dsRNA Construct #1 that is an exogenous trigger control. FIG. 2B presents a schematic representation of dsRNA Construct #1 that will serve as a control for the other experiments since it contains only the exogenous sequence with no additional features that should lead to its amplification. The length of the exogenous sequence is 234 bp, the same size of the original insert between the two Mir390BS in TAS3.
[0406] Example 9B provides dsRNA Construct #2 having a dual Mir390BS on sense strand and an exogenous sequence. FIG. 2C presents schematic representation of dsRNA Construct #2 having a dual Mir390BS on sense strand and an exogenous sequence. Double-stranded RNA Construct #2 is based on the dual Mir390BS from the TAS3 gene with an exogenous sequence replacing the original insert between the two Mir390BS.
[0407] Example 9C provides dsRNA construct #3 having a Dual Mir390BS both on the sense and antisense strands. FIG. 3 presents a schematic of dsRNA construct #3. This construct contains dual Mir390BS on both the sense and antisense strands and therefore we hypothesize that it will continuously recruit Mir390-Ago7 and RDRP to both strands. As a result, it is predicted to lead to long lasting amplification of the exogenous sequence and to ongoing production of its ta-siRNAs.
[0408] Example 9D provides dsRNA construct #4 having miR390BS as overhangs. FIG. 4 presents dsRNA construct #4 composed of two different strands. Having the Mir390BS present as overhangs will ease the initial requirement for the unwinding of the dsRNA since the Mir390BS will already be accessible for mir390 and Ago7 binding. The Mir390BS sequences will facilitate the unwinding and as a result the translocation into the processing center and the initiation of the entire process as explained in example #3 above.
[0409] Example 9E provides dsRNA construct #5 having Dual miR390BS sequences and helicase binding sequences. FIG. 5 presents dsRNA construct #5. The presence of the helicase binding sequences will enable more efficient unwinding of the dsRNA through active recruitment of a helicase and therefore leading to a strong and efficient amplification.
[0410] Example 9F provides dsRNA construct #6 having Mir390BS on both strands and a helicase binding sequence (helicase BS) overhang. FIG. 6 presents dsRNA construct #6 having Mir390BS on both strands and a helicase overhang. This dsRNA construct #6 is composed of two different strands. The sense strand is the same as in construct #3 and the antisense strand is the same as construct #3 with an addition of an overhang of a helicase BS at the 3' end. The helicase BS leads to recruitment of a helicase that will unwind the dsRNA and enable efficient initiation of the entire process. Each of the strands will contain Mir390-Ago7 sequences for binding and localization into the processing center enabling long lasting amplification.
[0411] Example 9G provides dsRNA construct #7 having a sense dual Mir390BS sequence coupled with an antisense Mir4376BS. FIG. 7 presents dsRNA construct #7 containing a dualMir390BS on its sense strand and a single Mir4376BS on its antisense strand. The presence of ta-siRNA inducing miRNAs on both strands will lead to ongoing amplification.
[0412] Example 9H provides dsRNA construct #8 having an Endogenous Trigger Control. FIG. 8 presents dsRNA construct #8 that is based on an exact endogenous insert sequence (the insert is the region between the two Mir390BS) in order to serve as an endogenous trigger control for the dsRNA construct #9 of Example 91.
[0413] Example 9I presents dsRNA construct #9 having a Mir390BS sequence and the Endogenous insert of Example 9H. FIG. 9 presents a schematic of dsRNA construct #9. Construct #9 maintains the endogenous sequence of the Mir390BS including the original insert region. This construct result in production of ta-siRNAs targeting ARF3 and ARF4.
Example 10: Treatment of Seeds with ta-siRNA Constructs and Analysis of RNA Levels
[0414] Tomato seeds were treated with dsRNA molecules corresponding to the constructs and sequences of Table 5.
TABLE-US-00010 TABLE 5 dsRNA constructs SEQ Trigger Trigger ID # alias Sequence (5'-3') Length DS/SS S/AS NO: 1 GFP234 CTAATACGACTCACTATAGGGAGATTTCCG 282 DS Sense 320 TCCTCCTTGAAATCAATTCCCTTAAGCTCG ATCCTGTTGACGAGGGTGTCTCCCTCAAAC TTGACTTCAGCACGTGTCTTGTAGTTCCCG TCGTCCTTGAAAGAGATGGTCCTCTCCTGC ACGTATCCCTCAGGCATGGCGCTCTTGAAG AAGTCGTGCCGCTTCATATGATCTGGGTAT CTTGAAAAGCATTGAACACCATAAGAGAA AGTAGTGACAAGTGTTGGCTCTCCCTATAG TGAGTCGTATTAG 2 GFP234Mir390 CTAATACGACTCACTATAGGGAGAGGTGC 325 DS Sense 321 TATCCTACCTGAGCTTTTTCCGTCCTCCTTG AAATCAATTCCCTTAAGCTCGATCCTGTTG ACGAGGGTGTCTCCCTCAAACTTGACTTCA GCACGTGTCTTGTAGTTCCCGTCGTCCTTG AAAGAGATGGTCCTCTCCTGCACGTATCCC TCAGGCATGGCGCTCTTGAAGAAGTCGTGC CGCTTCATATGATCTGGGTATCTTGAAAAG CATTGAACACCATAAGAGAAAGTAGTGAC AAGTGTTGGCCCTTGTCTATCCCTCCTGAG CTTCTCCCTATAGTGAGTCGTATTAG 3 GFP234Mir390X2 CTAATACGACTCACTATAGGGAGAAGCTC 368 DS Sense 322 AGGAGGGATAGACAAGGGGTGCTATCCTA CCTGAGCTTTTTCCGTCCTCCTTGAAATCA ATTCCCTTAAGCTCGATCCTGTTGACGAGG GTGTCTCCCTCAAACTTGACTTCAGCACGT GTCTTGTAGTTCCCGTCGTCCTTGAAAGAG ATGGTCCTCTCCTGCACGTATCCCTCAGGC ATGGCGCTCTTGAAGAAGTCGTGCCGCTTC ATATGATCTGGGTATCTTGAAAAGCATTGA ACACCATAAGAGAAAGTAGTGACAAGTGT TGGCAAGCTCAGGTAGGATAGCACCCCTT GTCTATCCCTCCTGAGCTTCTCCCTATAGT GAGTCGTATTAG 4 GFP234Mir390_ CTAATACGACTCACTATAGGGAGAGGTGC 301 SS Sense 323 Sense TATCCTACCTGAGCTTTTTCCGTCCTCCTTG AAATCAATTCCCTTAAGCTCGATCCTGTTG ACGAGGGTGTCTCCCTCAAACTTGACTTCA GCACGTGTCTTGTAGTTCCCGTCGTCCTTG AAAGAGATGGTCCTCTCCTGCACGTATCCC TCAGGCATGGCGCTCTTGAAGAAGTCGTGC CGCTTCATATGATCTGGGTATCTTGAAAAG CATTGAACACCATAAGAGAAAGTAGTGAC AAGTGTTGGCCCTTGTCTATCCCTCCTGAG CT 5 GFP234Mir390_ CTAATACGACTCACTATAGGGAGAGCCAA 258 SS Antisense 324 Antisense CACTTGTCACTACTTTCTCTTATGGTGTTCA ATGCTTTTCAAGATACCCAGATCATATGAA GCGGCACGACTTCTTCAAGAGCGCCATGCC TGAGGGATACGTGCAGGAGAGGACCATCT CTTTCAAGGACGACGGGAACTACAAGACA CGTGCTGAAGTCAAGTTTGAGGGAGACAC CCTCGTCAACAGGATCGAGCTTAAGGGAA 6 GFP234Mir390_ CTAATACGACTCACTATAGGGAGAGGTGC 375 DS Sense 325 Helicase TATCCTACCTGAGCTTTTTCCGTCCTCCTTG AAATCAATTCCCTTAAGCTCGATCCTGTTG ACGAGGGTGTCTCCCTCAAACTTGACTTCA GCACGTGTCTTGTAGTTCCCGTCGTCCTTG AAAGAGATGGTCCTCTCCTGCACGTATCCC TCAGGCATGGCGCTCTTGAAGAAGTCGTGC CGCTTCATATGATCTGGGTATCTTGAAAAG CATTGAACACCATAAGAGAAAGTAGTGAC AAGTGTTGGCGCTACTGGACTTTCGCCATC TAGGGTGCAGCATTCGGGCTGCTTCGCCTA CCTTGTCTATCCCTCCTGAGCTTCTCCCTAT AGTGAGTCGTATTAG 7 GFP234Mir390_ CTAATACGACTCACTATAGGGAGAAGCTC 344 SS Sense 326 Helicase_ AGGAGGGATAGACAAGGGGTGCTATCCTA Sense CCTGAGCTTTTTCCGTCCTCCTTGAAATCA ATTCCCTTAAGCTCGATCCTGTTGACGAGG GTGTCTCCCTCAAACTTGACTTCAGCACGT GTCTTGTAGTTCCCGTCGTCCTTGAAAGAG ATGGTCCTCTCCTGCACGTATCCCTCAGGC ATGGCGCTCTTGAAGAAGTCGTGCCGCTTC ATATGATCTGGGTATCTTGAAAAGCATTGA ACACCATAAGAGAAAGTAGTGACAAGTGT TGGCAAGCTCAGGTAGGATAGCACCCCTT GTCTATCCCTCCTGAGCT 8 GFP234Mir390_ CTAATACGACTCACTATAGGGAGAAGCTC 394 SS Antisense 327 Helicase_ AGGAGGGATAGACAAGGGGTGCTATCCTA AntiSense CCTGAGCTTGCCAACACTTGTCACTACTTT CTCTTATGGTGTTCAATGCTTTTCAAGATA CCCAGATCATATGAAGCGGCACGACTTCTT CAAGAGCGCCATGCCTGAGGGATACGTGC AGGAGAGGACCATCTCTTTCAAGGACGAC GGGAACTACAAGACACGTGCTGAAGTCAA GTTTGAGGGAGACACCCTCGTCAACAGGA TCGAGCTTAAGGGAATTGATTTCAAGGAG GACGGAAAAAGCTCAGGTAGGATAGCACC CCTTGTCTATCCCTCCTGAGCTTAGGCGAA GCAGCCCGAATGCTGCACCCTAGATGGCG AAAGTCCAGTAGC 9 GFP234Mir390_ CTAATACGACTCACTATAGGGAGAGGTGC 347 DS Sense 328 Mir4376 TATCCTACCTGAGCTTTTTCCGTCCTCCTTG AAATCAATTCCCTTAAGCTCGATCCTGTTG ACGAGGGTGTCTCCCTCAAACTTGACTTCA GCACGTGTCTTGTAGTTCCCGTCGTCCTTG AAAGAGATGGTCCTCTCCTGCACGTATCCC TCAGGCATGGCGCTCTTGAAGAAGTCGTGC CGCTTCATATGATCTGGGTATCTTGAAAAG CATTGAACACCATAAGAGAAAGTAGTGAC AAGTGTTGGCTCGCAGGAGAGATGACACC AGACCTTGTCTATCCCTCCTGAGCTTCTCC CTATAGTGAGTCGTATTAG 10 TAS3 CTAATACGACTCACTATAGGGAGATTTCTC 282 DS Sense 329 ACCGCTTTTTTTTTTCTGTTGTGTATTCTCT TTTTTGACTTGTTGCCTTTCGTTCCTCTACC TACCCCATTCTTCTTGACCTTGTAAGACCTT TTCTTGACCTTGTAAGACCCCGTGTTATCT CTTACGTCTTTATGTTTTGTTTTTTTGCAAA TCTTACGTCATGACTTCTTCATGTAAGCTTT GTTTGGTCTCCTTCTTCTTTCCTACTCAACT CTCGTTCTCCTTTCTCCCTATAGTGAGTCGT ATTAG 11 TAS3Mir390 CTAATACGACTCACTATAGGGAGAGGTGC 325 DS Sense 330 TATCCTACCTGAGCTTTTTCTCACCGCTTTT TTTTTTCTGTTGTGTATTCTCTTTTTTGACTT GTTGCCTTTCGTTCCTCTACCTACCCCATTC TTCTTGACCTTGTAAGACCTTTTCTTGACCT TGTAAGACCCCGTGTTATCTCTTACGTCTTT ATGTTTTGTTTTTTTGCAAATCTTACGTCAT GACTTCTTCATGTAAGCTTTGTTTGGTCTCC TTCTTCTTTCCTACTCAACTCTCGTTCTCCT TCCTTGTCTATCCCTCCTGAGCTTCTCCCTA TAGTGAGTCGTATTAG 12 GUS234 GCCACTTGCAAAGTCCCGCTAGTGCCT 236 DS Sense 331 TGTCCAGTTGCAACCACCTGTTGATCC GCATCACGCAGTTCAACGCTGACATCA CCATTGGCCACCACCTGCCAGTCAACA GACGCGTGGTTACAGTCTTGCGCGACA TGCGTCACCACGGTGATATCGTCCACC CAGGTGTTCGGCGTGGTGTAGAGCATT ACGCTGCGATGGATTCCGGCATAGTTA AAGAAATCATGGAAGTAAGC 13 GUS234Mir390 GGTGCTATCCTACCTGAGCTTCCACTT 278 DS Sense 332 GCAAAGTCCCGCTAGTGCCTTGTCCAG TTGCAACCACCTGTTGATCCGCATCAC GCAGTTCAACGCTGACATCACCATTGG CCACCACCTGCCAGTCAACAGACGCGT GGTTACAGTCTTGCGCGACATGCGTCA CCACGGTGATATCGTCCACCCAGGTGT TCGGCGTGGTGTAGAGCATTACGCTGC GATGGATTCCGGCATAGTTAAAGAAAT CATGGAAGTAAGCCTTGTCTATCCCTC CTGAGCTC
[0415] GFP234 (FIG. 10A; Trigger #1), GFP234Mir390 (Trigger #2), TAS3 (Trigger #10) and TAS3Mir390 (FIG. 10B, Trigger #11) were prepared as provided in Example 1. A final concentration of 50 .mu.g/ml dsRNA diluted with 0.1 mM EDTA was used. Treatment was performed by gently shaking the seeds in the solution for 24 hours in a dark growth chamber at 15.degree. C. followed by washing with water three times for one minute. After treatment, seeds were germinated either on wet paper or in soil and grown at about 25.degree. C. with a 16 hour photoperiod. The plants germinated in soil were watered with tap water as necessary. Seeds that were treated with a similar solution not containing dsRNA (e.g., 0.1 mM EDTA "EDTA") were germinated and grown alongside the treated plants as a control.
[0416] Total RNA was extracted from whole seedlings, leaves or roots of germinated seeds seven, 14 and 30 days post treatment. For seeds that were germinated on paper, the entire seedling was harvested after seven days. For seeds that were germinated in soil, the leaves and roots were harvested and analyzed separately 14 and 30 days after treatment.
[0417] cDNA was prepared using oligo-dT primers and the expression levels of TAS3, ARF3 and ARF4 was determined by real-time PCR with SYBR Green (Quanta BioSciences). The results are presented in FIG. 11A-E. The house-keeping genes Expressed and Tip41 were used as endogenous control genes to normalize for input RNA amounts. ARF3 and ARF4 genes are regulated by the TAS3 system and their expression is predicted to decrease following TAS3Mir390 treatment. No significant difference in TAS3, ARF3 or ARF4 expression was detected in seedlings when comparing the TAS3 construct with the TAS3Mir390 construct seven days after treatment (t-test, p-value>0.05). Similarly, no significant difference in TAS3, ARF3 or ARF4 expression was detected in leaves when comparing the two constructs 14 days after treatment. When comparing expression levels in roots 14 days after treatment, a down-regulation trend was observed for ARF3 and ARF4 genes following treatment with the TAS3Mir390 construct (FIGS. 11A-B). FIG. 11A shows relative quantification of ARF3 mRNA following treatment with either TAS3 Insert or Mir390BS TAS3 dsRNA constructs. Each point represents the expression value per individual plant. Expression values were normalized to the average expression values of all plants treated with TAS3_Insert, which was set to 100%. The red line represents the normalized average expression values for each treatment. FIG. 11B shows the same analysis for ARF4 mRNA levels.
[0418] The RNA extracted from seven-day old seedlings, as well as RNA extracted from 30-day old roots and 30-day old leaves was used in a second cDNA reaction with random primers and then subjected to real-time PCR with SYBR Green (Quanta BioSciences). The primers used for real-time were derived from the GFP sequence that appears in the dsRNA constructs. Therefore, this analysis provides an indication for the presence of the dsRNA that was used for seed treatment and/or for RNA that was synthesized from this dsRNA in the plant tissue. Expressed and Tip41 were used as endogenous control genes. For seven day old seedlings, a 2.6 fold difference in GFP level was observed in plants treated with GFP234 compared to plants treated with TAS3 (t-test, p-value=0.08). However, no significant difference in GFP level was observed when comparing between GFP234 and GFP234Mir390 treatments (FIGS. 11C-E). FIG. 11C shows normalized Ct values for all treatments analyzed; for each RNA sample tested, the Ct value obtained from the real-time amplification plot was normalized to the average Ct value of the two endogenous control genes. This value was then subtracted from the number 50 to assign larger values for higher expression levels. Each dot represents one plant and the red line represents the average value per treatment. RNA samples that gave no Ct values were assigned a value of 40. FIG. 11D shows a comparison between GFP234 and TAS3 treatments. FIG. 11E shows a comparison between GFP234 and GFP234Mir390 treatments. For 30 day old roots, no significant difference in GFP level was observed when comparing between GFP234 and GFP234Mir390 treatments. For 30 day old leaves, a significant (t-test, p-value=0.005), 7.1-fold difference in GFP level was observed in plants treated with GFP234Mir390 compared to plants treated with GFP234 (FIG. 11F).
Example 11: Detection of GFP Sequence in Plants Following Seed Treatment with ta-si dsRNA Constructs
[0419] In a second experiment, tomato seeds were treated with dsRNA molecules corresponding to GFP234 (Trigger#1), GFP234Mir390 (Trigger#2), GFP234Mir390X2 (Trigger#3), GFP234Mir390_Helicase (Trigger#6), GFP234Mir390_Mir4376 (Trigger#9), TAS3 (Trigger#10) and TAS3Mir390 (Trigger#11) as provided above in Example 10 and Table 5, according to the protocol described in Example 10. A final concentration of 50 .mu.g/ml dsRNA diluted with 0.1 mM EDTA was used. Treatment was performed by gently shaking the seeds in the solution for 24 hours in a dark growth chamber at 15.degree. C. followed by washing with water three times for one minute. After treatment, seeds were germinated on wet paper and grown at about 25.degree. C. with 16 hours photoperiod. Seeds that were treated with EDTA solution alone were germinated and grown alongside the treated plants as a control.
[0420] Total RNA was extracted from shoots (including hypocotyl, cotyledon and shoot apical meristem) seven and fourteen days after treatment. cDNA was prepared using random primers and the presence of the GFP sequence was determined and quantified by real-time PCR as described in Example 10. A significant difference in GFP level was observed in plants seven days after treatment with GFP234Mir390, GFP234Mir390_Helicase or GFP234Mir390_Mir4376 dsRNAs compared to treatment with GFP234, TAS3Mir390 or EDTA. The GFP level detected was between 5.9 fold (following treatment with GFP234Mir390 Mir4376) to 25 fold (following treatment with GFP234Mir390_Helicase) higher compared to plants treated with GFP234, with a p-value<0.05 (t-test). No significant difference was detected when comparing GFP234Mir390X2 to GFP234 treatment (FIGS. 12A-E). The analyses were performed as described for FIGS. 11C-F of Example 10. FIG. 12A shows normalized Ct values for all treatments. FIG. 12B shows a comparison between GFP234 and GFP234Mir390 treatments. FIG. 12C shows a comparison between GFP234 and GFP234Mir390_Helicase treatments. FIG. 12D shows a comparison between GFP234 and GFP234Mir390_Mir4376treatments. FIG. 12E shows a comparison between GFP234 and GFP234Mir390X2 treatments.
[0421] A significant, 33 fold difference in GFP level was observed in plants 14 days after treatment with GFP234Mir390X2 dsRNA compared to treatment with GFP234 (t-test, p-value<0.05). Higher levels of GFP were also detected for GFP234Mir390, GFP234Mir390 Helicase and GFP234Mir390 Mir4376 treatments compared to GFP234 treatment, but with no significant difference (FIGS. 13A-B). The analyses were performed as described for FIGS. 11C-F. FIG. 13A shows normalized Ct values for all treatments. FIG. 13B shows a comparison between GFP234 and GFP234Mir390X2 treatments.
[0422] The same cDNA prepared from RNA extracted from seven-day old seedlings, was used in a second real-time PCR, where the expression levels of TAS3, ARF3 and ARF4 was determined as described in Example 10 (except that random primers, and not oligo-dT primers were used in the cDNA reaction). No significant difference in TAS3, ARF3 or ARF4 expression was detected in seedlings when comparing between TAS3 and TAS3Mir390 treatments (t-test, p-value>0.05).
Example 12: Detection of GFP Sequence in Plants Following Seed Treatment with ta-si dsRNA Constructs
[0423] Tomato seeds were treated with dsRNA molecules corresponding to GFP234 (Trigger#1), GFP234Mir390_Helicase (Trigger#6) and GFP234Mir390_Mir4376 (Trigger#9), as provided above in Example 10 and Table 4, according to the protocol described in Example 1. A final concentration of 50 .mu.g/ml dsRNA diluted with 0.1 mM EDTA was used. Treatment was performed by gently shaking the seeds in the solution for 24 hours in a dark growth chamber at 15.degree. C. followed by washing with water three times for one minute. After treatment, seeds were germinated on wet paper and grown at about 25.degree. C. with 16 hours photoperiod. Seeds that were treated with EDTA solution alone were germinated and grown alongside the treated plants as a control.
[0424] Total RNA was extracted from shoots (including hypocotyl, cotyledon and shoot apical meristem) seven days after treatment. cDNA was prepared using random primers and the presence of the GFP sequence was determined and quantified by real-time PCR as described in Example 10. No significant difference in GFP levels was observed when comparing the GFP234 treated plants to the GFP234Mir390_Helicase or GFP234Mir390_Mir4376 treated plants (Dunnett's test).
Example 13: Detection of GFP Sequence in Plants Following Seed Treatment with ta-si dsRNA Constructs
[0425] Tomato seeds were treated with dsRNA molecules corresponding to GFP234 (Trigger#1), GFP234Mir390 (Trigger#2), GFP234Mir390X2 (Trigger#3), GFP234Mir390 Helicase (Trigger#6), GFP234Mir390_Mir4376 (Trigger#9), TAS3 (Trigger#10) and TAS3Mir390 (Trigger#11) as provided above in Example 10 and Table 5, according to the protocol described in Example 1. A final concentration of 50 .mu.g/ml dsRNA diluted with 0.1 mM EDTA was used. Treatment was performed by gently shaking the seeds in the solution for 24 hours in a dark growth chamber at 15.degree. C. followed by washing with water three times for one minute. After treatment, seeds were germinated on wet paper and grown at about 25.degree. C. with 16 hours photoperiod. Seeds that were treated with EDTA solution alone were germinated and grown alongside the treated plants as a control.
[0426] Total RNA was extracted from shoots (including hypocotyl, cotyledon and shoot apical meristem) seven and 14 days after treatment. cDNA was prepared using random primers and the presence of the GFP sequence was determined and quantified by real-time PCR as described in Example 10. Seven days after treatment, a significant, 10-fold difference in GFP level was observed in plants treated with GFP234Mir390 compared to plants treated with GFP234 (Dunnett's test, p-value=0.035). A 6.6-fold difference in GFP level was observed in plants following treatment with GFP234Mir390_Helicase compared to treatment with GFP234 (Dunnett's test, p-value=0.11). For 14 days old plants, a significant, 34-fold difference in GFP level was observed in plants treated with GFP234Mir390_Helicase compared to plants treated with GFP234 (Dunnett's test, p-value=0.0004). An 8.5-fold difference in GFP level was observed in plants following treatment with GFP234Mir390 compared to treatment with GFP234 (Dunnett's test, p-value=0.058). FIG. 14A shows normalized Ct values seven days after treatment. FIG. 14B shows normalized Ct values 14 days after treatment. The analysis was performed essentially as described for FIG. 11C-F, except that instead of subtracting the normalized Ct values from the number 50 to assign larger values for higher GFP levels, an inverse y-axis is presented.
[0427] RNA extracted from seven-day old shoots treated with TAS3 (Trigger#10) and TAS3Mir390 (Trigger#11) dsRNAs was used in a second real-time PCR to determine the expression levels of TAS3, ARF3 and ARF4 as described in Example 11. A significant, 1.3-fold up-regulation in ARF4 expression was detected in plants following treatment with TAS3Mir390 compared to treatment with TAS3 (Dunnett's test, p-value=0.05). No significant difference in TAS3 or ARF3 expression was detected in those plants.
Example 14: Detection of Gus Sequence in Plants Following Seed Treatment with ta-si dsRNA Constructs
[0428] Tomato seeds were treated with dsRNA molecules corresponding to GUS234 (Trigger#12) and GUS234Mir390 (Trigger#13), as provided above in Example 10 and Table 5, according to the protocol described in Example 1. A final concentration of 50 .mu.g/ml dsRNA diluted with 0.1 mM EDTA was used. Treatment was performed by gently shaking the seeds in the solution for 24 hours in a dark growth chamber at 15.degree. C. followed by washing with water three times for one minute. After treatment, seeds were germinated on wet paper and grown at about 25.degree. C. with 16 hours photoperiod. Seeds that were treated with EDTA solution alone were germinated and grown alongside the treated plants as a control.
[0429] Total RNA was extracted from shoots (including hypocotyl, cotyledon and shoot apical meristem) seven and 14 days after treatment. cDNA was prepared using random primers and the presence of the GUS sequence was determined and quantified by real-time PCR as described in Example 10. A significant increase in GUS levels was observed in plants seven days after treatment with GUS234Mir390 compared to treatment with GUS234 (Dunnett's test, p-value=0.0005). Most of the RNA samples extracted from GUS234-treated plants gave no Ct value, meaning GUS was not detected in those samples. Accordingly, these samples were assigned a value of 40. The resulting difference in the average Ct value between the GUS234 and the GUS234Mir390 treatments was calculated to be about 10, translating into a 994-fold difference in GUS levels. For 14 days old plants, a 12.2-fold difference in GUS levels was observed (Dunnett's test, p-value=0.03865). FIG. 15A shows normalized Ct values seven days after treatment with the two dsRNA constructs. FIG. 15B shows normalized Ct values 14 days after treatment. The analysis was performed as described for FIGS. 14A-B.
Example 15: Small RNA Deep Sequencing of Plants Following Seed Treatment with ta-si dsRNA Constructs
[0430] RNA samples from the seven day old shoots described in Examples 11 and 14 were further analyzed by small RNA deep sequencing. cDNA libraries were prepared with an Illumina TruSeq.TM. Small RNA kit according to the manufacturer's protocol, and sequenced by Illumina MiSeq.RTM. instrument. Each cDNA library was prepared from RNA pooled from three plants originating from the same treatment. For GFP-based dsRNAs, a total of ten libraries were prepared. Two libraries (representing a total of six plants) were prepared from GFP234 treated plants, three libraries (nine plants) were prepared from GFP234Mir390 treated plants, one library (three plants) was prepared from GFP234Mir390X2 treated plants, two libraries (six plants) were prepared from GFP234Mir390_Helicase treated plants and two libraries (six plants) were prepared from TAS3Mir390 treated plants. Table 6 shows the average Ct value of each of the pooled RNA samples, according to the real-time PCR analysis shown in FIG. 12A. The values presented in the table were normalized by subtracting the average Ct value of RNA pooled from the TAS3Mir390 treatment. Low quality reads and reads that contain adaptor sequences were filtered out from the raw sequencing data. Table 6 summarizes the number of reads from each library that were mapped to GFP. In accordance with the real-time PCR results, more reads were mapped to the GFP sequence in the GFP234Mir390 and GFP234Mir390_Helicase treatments compared to the GFP234 treatment.
TABLE-US-00011 TABLE 6 Small RNA MiSeq analysis of RNA extracted from plants following seed treatment with ta-si-GFP dsRNA constructs GFP234Mir Treatment GFP234 GFP234Mir390 GFP234Mir390_Helicase 390X2 TAS3Mir390 RT-PCR 1.3 2.8 3.2 6.6 9.5 7.9 5.9 2.7 0.0 4.0 normalized Ct value Libraries size 1.23 1.24 2.09 1.46 1.4 1.59 1.23 1 1.61 1.07 ratio Total # of reads 36 12 74 130 273 122 117 20 0 2 mapped to GFP234 sequence Normalized # of 29 10 35 89 195 77 95 20 0 2 reads mapped to GFP234 sequence
[0431] For GUS-based dsRNAs, two libraries were prepared, one library (representing a total of three plants) was prepared from GUS234 treated plants and one library (three plants) was prepared from GUS234Mir390 treated plants. Table 7 shows the average Ct value of each of the pooled RNA samples, according to the real-time PCR analysis shown in FIG. 15A. The values presented in the Table were normalized by subtracting the average Ct value of RNA pooled from the GUS234 treatment. Table 7 summarizes the number of reads from each library that were mapped to GUS. Data was analyzed as described for Table 6. In accordance with the real-time PCR results, no reads were mapped to the GUS sequence in the GUS234 treatment while some reads were mapped to GUS in the GUS234Mir390 treatment.
TABLE-US-00012 TABLE 7 Small RNA MiSeq analysis of RNA extracted from plants following seed treatment with ta-si-GUS dsRNA constructs. Treatment GUS234 GUS234Mir390 RT-PCR 0 7.6 normalized Ct value Libraries size 1 1.77 ratio Total # of reads 0 27 mapped to GUS234 Normalized # of 0 15 reads mapped to GUS234 sequence
Example 16: Additional dsRNA Constructs
[0432] Additional dsRNA constructs based on the constructs and sequences provided in Examples 9 and 10 are provided. Deep-Sequencing analysis described in Example 15 indicated that the first nucleotide in the small RNA reads mapped to the endogenous TAS3 transcript is predominantly "T". The most abundant reads mapped to the TAS3 transcript were located at positions 37, 38, 79, 100, 101, 103, 184 and 185 of the 234nt sequence and with the exception of the reads mapped to position 185, where the first nucleotide was A, the first nucleotide was T. Therefore, the inclusion of a "T" or "A" in the sequence of the target gene of interest that is flanked by the two miR390 binding sites is expected to improve the efficiency of cleavage and direct it to specific sites within the sequence or alternatively improves the interaction of the resulting small RNAs with downstream effectors.
[0433] Trigger #14 (SEQ ID No. 314), is designed as a modified GFP234Mir390 sequence (based on Example 9B and trigger #2), where the nucleotides at positions 37, 38, 79, 100, 101, 103 and 184 are "T", and the nucleotide at position 185 is A (only positions 37, 79 and 101 were changed. Sequence appears in 5'-3' orientation, the mentioned positions are in lowercase, bold).
TABLE-US-00013 SEQ ID No. 314: GGTGCTATCCTACCTGAGCTTTTTCCGTCCTCCTTGAAATCAATTCCCTT AAGCTCGttCCTGTTGACGAGGGTGTCTCCCTCAAACTTGACTTCAGCAt GTGTCTTGTAGTTCCCGTCGttCtTGAAAGAGATGGTCCTCTCCTGCACG TATCCCTCAGGCATGGCGCTCTTGAAGAAGTCGTGCCGCTTCATATGATC TGGGtaTCTTGAAAAGCATTGAACACCATAAGAGAAAGTAGTGACAAGTG TTGGCCCTTGTCTATCCCTCCTGAGCT
[0434] Trigger #15 (SEQ ID No. 315) is designed as a modified TAS3Mir390 sequence (based on Example 91 and trigger #11). In this sequence, four 21 nucleotides segments that begin with "TT" are selected from the GFP234 sequence and used to replace the original 21 nucleotide segments from the TAS3Mir390 sequence at positions 37, 79, 100 and 184. These positions are in phase with the miR390-guided cleavage site (sequence appears in 5'-3' orientation, the mentioned four segments are in lowercase, bold).
TABLE-US-00014 SEQ ID No. 315: GGTGCTATCCTACCTGAGCTTTTTCTCACCGCTTTTTTTTTTCTGTTGTG TATTCTCtttccgtcctccttgaaatcaGTTCCTCTACCTACCCCATTCt tcccttaagctcgatcctgtttgacttcagcacgtgtcttgGTGTTATCT CTTACGTCTTTATGTTTTGTTTTTTTGCAAATCTTACGTCATGACTTCTT CATGttcccgtcgtccttgaaagagCTTCTTTCCTACTCAACTCTCGTTC TCCTTCCTTGTCTATCCCTCCTGAGCT
[0435] Trigger #16 (SEQ ID No. 316) is designed as a modified GFP234Mir390 sequence (based on Example 9B and trigger #2), where "T" appears every 21 nucleotides. The most 3' "T" is located at position 226 of the GFP234 sequence, 21 nucleotides upstream to the miR390-guided cleavage site and all other "T" positions are in phase with this site (sequence appears in 5'-3' orientation, only mutated nucleotides are in lowercase, bold).
TABLE-US-00015 SEQ ID No. 316: GGTGCTATCCTACCTGAGCTTTTTCCGTCCTCCTTGtAATCAATTCCCTT AAGCTCGtTCCTGTTGACGAGGGTGTCTtCCTCAAACTTGACTTCAGCAt GTGTCTTGTAGTTCCCGTCGTCCTTGAAAGAGATGGTCCTCTCCTGCACG TATCCCTCAGGCtTGGCGCTCTTGAAGAAGTCGTGCCGCTTCATATGATC TGGGTATCTTGAAAAGCATTGAACAtCATAAGAGAAAGTAGTGACAtGTG TTGGCCCTTGTCTATCCCTCCTGAGCT
[0436] Trigger #17 (SEQ ID No. 317) is designed as a modified GFP234Mir390 sequence (based on Example 9B and trigger #2), where ten 21 nucleotide segments that originate from the full-length GFP sequence and begins with a "T" are placed in tandem to produce the 234 nucleotide sequence. The first 15 and the last 9 nucleotides are from the endogenous TAS3 sequence, in order to position the ten segments in-phase with the miR390-guided cleavage site (sequence appears in 5'-3' orientation, the first "T" in each segment is in lowercase, bold, and the TAS3 sequence is underlined).
TABLE-US-00016 SEQ ID No. 317: GGTGCTATCCTACCTGAGCTTTTTCTCACCGCTTTTtAATGGTTGTCTGG TAAAAGGtCGCCAATTGGAGTATTTTGTtGATAATGATCAGCGAGTTGCt CTTCGATGTTGTGGCGGGTCtTGAAGTTGGCTTTGATGCCGtTCTTTTGC TTGTCGGCCATGtGTATACGTTGTGGGAGTTGTtTGTATTCCAACTTGTG GCCGtGTTTCCGTCCTCCTTGAAATtTCCCTTAAGCTCGATCCTGTGTTC TCCTTCCTTGTCTATCCCTCCTGAGCT
[0437] Tomato seeds are treated with dsRNA triggers 14, 15, 16 and 17 as described in Example 10. The seeds are germinated on paper and the seedlings is harvested at 7, 14 and 30 days post treatment and total RNA is extracted from whole seedlings, leaves or roots of the germinated seeds. cDNA is prepared using oligo-dT primers and the expression levels of TAS3, ARF3 and ARF4 are determined by real-time PCR with SYBR Green (Quanta BioSciences). ARF3 and ARF4 genes are regulated by the TAS3 system and their expression is predicted to decrease following trigger treatment.
[0438] Although the disclosure has been described in conjunction with specific embodiments thereof, it is evident that many alternatives, modifications and variations will be apparent to those skilled in the art. Accordingly, it is intended to embrace all such alternatives, modifications and variations that fall within the spirit and broad scope of the appended claims.
[0439] All publications, patents and patent applications mentioned in this specification are herein incorporated in their entirety by reference into the specification, to the same extent as if each individual publication, patent or patent application was specifically and individually indicated to be incorporated herein by reference. In addition, citation or identification of any reference in this application shall not be construed as an admission that such reference is available as prior art to the present disclosure. To the extent that section headings are used, they should not be construed as necessarily limiting.
Sequence CWU 1
1
332120DNAArtificial SequenceDescription of Artificial Sequence
Synthetic T7 primer sequence 1taatacgact cactataggg
20219DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 2ggtgctctga acgtggatg
19322DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 3catcatcgcc atcctcattc
tc 22440DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 4taatacgact cactataggg
gaagaccctc gaaactaagc 40540DNAArtificial SequenceDescription of
Artificial Sequence Synthetic single strand DNA oligonucleotide
5taatacgact cactataggg ggtaagcggc attctaaacc 40620DNAArtificial
SequenceDescription of Artificial Sequence Synthetic single strand
DNA oligonucleotide 6actcagcagt cgtaggattg 20720DNAArtificial
SequenceDescription of Artificial Sequence Synthetic single strand
DNA oligonucleotide 7cttcttatgt tcccgtcagg 208616DNAArtificial
SequenceDescription of Artificial Sequence Synthetic CGMVV dsRNA
product 1 polynucleotide 8taatacgact cactataggg ggtaagcggc
attctaaacc tccaaatcgg aggttggact 60ctgcttctga agagtccagt tctgtttctt
ttgaagatgg cttacaatcc gatcacacct 120agcaaactta ttgcgtttag
tgcttcttat gttcccgtca ggactttact taattttcta 180gttgcttcac
aaggtaccgc tttccagact caagcgggaa gagattcttt ccgcgagtcc
240ctgtctgcgt taccctcgtc tgtcgtagat attaattcta gattcccaga
tgcgggtttt 300tacgctttcc tcaacggtcc tgtgttgagg cctatcttcg
tttcgcttct cagctccacg 360gatacgcgta atagggtcat tgaggttgta
gatcctagca atcctacgac tgctgagtcg 420cttaacgccg taaagcgtac
tgatgacgcg tctacggccg ctagggctga gatagataat 480ttaatagagt
ctatttctaa gggttttgat gtttacgata gggcttcatt tgaagccgcg
540ttttcggtag tctggtcaga ggctaccacc tcgaaagctt agtttcgagg
gtcttcccct 600atagtgagtc gtatta 616940DNAArtificial
SequenceDescription of Artificial Sequence Synthetic single strand
DNA oligonucleotide 9taatacgact cactataggg catcaccatc gaccctaaac
401040DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 10taatacgact cactataggg
gctttaccgc cactaagaac 4011598DNAArtificial SequenceDescription of
Artificial Sequence Synthetic CGMVV dsRNA product 2 polynucleotide
11taatacgact cactataggg gctttaccgc cactaagaac tctgtacact cccttgcggg
60tggtctgagg cttcttgaat tggaatatat gatgatgcaa gtgccctacg gctcaccttg
120ttatgacatc ggcggtaact atacgcagca cttgttcaaa ggtagatcat
atgtgcattg 180ctgcaatccg tgcctagatc ttaaagatgt tgcgaggaat
gtgatgtaca acgatatgat 240cacgcaacat gtacagaggc acaagggatc
tggcgggtgc agacctcttc caactttcca 300gatagatgca ttcaggaggt
acgatagttc tccctgtgcg gtcacctgtt cagacgtttt 360ccaagagtgt
tcctatgatt ttgggagtgg tagggataat catgcagtct cgttgcattc
420aatctacgat atcccttatt cttcgatcgg acctgctctt cataggaaaa
atgtgcgagt 480ttgttatgca gcctttcatt tctcggaggc attgctttta
ggttcgcctg taggtaattt 540aaatagtatt ggggctcagt ttagggtcga
tggtgatgcc ctatagtgag tcgtatta 5981219DNAArtificial
SequenceDescription of Artificial Sequence Synthetic single strand
DNA oligonucleotide 12ggtgctctga acgtggatg 191322DNAArtificial
SequenceDescription of Artificial Sequence Synthetic single strand
DNA oligonucleotide 13catcatcgcc atcctcattc tc 2214166DNAArtificial
SequenceDescription of Artificial Sequence Synthetic backbone
sequence with two smRNA complementary sites and a helicase binding
site 14gcatcctcat cttaatctcg gtgctatcct acctgagctt gatatctagg
cgaagcagcc 60cgaatgctgc accctagatg gcgaaagtcc agtagcgata tcgaattcct
cgagggatcc 120aagcttcctt gtctatccct cctgagctgt tgattttatt ccatgt
1661543DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 15taatacgact cactataggg
agcattcccg gcgggatagt ctg 431643DNAArtificial SequenceDescription
of Artificial Sequence Synthetic single strand DNA oligonucleotide
16taatacgact cactataggg agcattcccg gcgggatagt ctg
431720DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 17cagcgcgaag tctttatacc
201820DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 18ctttgccgta atgagtgacc
201920DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 19ccataaccct ggaggttgag
202020DNAArtificial SequenceDescription of Artificial Sequence
Synthetic single strand DNA oligonucleotide 20atcagacgct gctggtctgg
2021443DNAArtificial SequenceDescription of Artificial Sequence
Synthetic GUS dsRNA product polynucleotide 21taatacgact cactataggg
agatcgacgg cctgtgggca ttcagtctgg atcgcgaaaa 60ctgtggaatt gatcagcgtt
ggtgggaaag cgcgttacaa gaaagccggg ctattgctgt 120gccaggcagt
tttaacgatc agttcgccga tgcagatatt cgtaattatg cgggcaacgt
180ctggtatcag cgcgaagtct ttataccgaa aggttgggca ggccagcgta
tcgtgctgcg 240tttcgatgcg gtcactcatt acggcaaagt gtgggtcaat
aatcaggaag tgatggagca 300tcagggcggc tatacgccat ttgaagccga
tgtcacgccg tatgttattg ccgggaaaag 360tgtacgtatc accgtttgtg
tgaacaacga actgaactgg cagactatcc cgccgggaat 420gctccctata
gtgagtcgta tta 44322116DNAArtificial SequenceDescription of
Artificial Sequence Synthetic backbone sequence with two smRNA
complementary sites, without the helicase binding site 22gcatcctcat
cttaatctcg gtgctatcct acctgagctt gatatcgata tcgaattcct 60cgagggatcc
aagcttcctt gtctatccct cctgagctgt tgattttatt ccatgt
11623167DNAArtificial SequenceDescription of Artificial Sequence
Synthetic backbone sequence with two smRNA complementary sites and
an helicase binding site 23gcatcctcat cttaatctcg tgatttttct
ctacaagcga agatatctag gcgaagcagc 60ccgaatgctg caccctagat ggcgaaagtc
cagtagcgat atcgaattcc tcgagggatc 120caagctttct tgctcaaatg
agtattccag ttgattttat tccatgt 16724117DNAArtificial
SequenceDescription of Artificial Sequence Synthetic backbone
sequence with two smRNA complementary sites, without the helicase
binding site 24gcatcctcat cttaatctcg tgatttttct ctacaagcga
agatatcgat atcgaattcc 60tcgagggatc caagctttct tgctcaaatg agtattccag
ttgattttat tccatgt 1172521RNAUnknownDescription of Unknown miR390
oligonucleotide 25aagcucagga gggauagcgc c
212621RNAUnknownDescription of Unknown miR161.1 oligonucleotide
26uugaaaguga cuacaucggg g 212721RNAUnknownDescription of Unknown
miR400 oligonucleotide 27uaugagagua uuauaaguca c
212821RNAUnknownDescription of Unknown TAS2 3'D6(-) smRNA
oligonucleotide 28auaucccauu ucuaccaucu g
212921RNAUnknownDescription of Unknown TAS1b 3'D4(-) smRNA
oligonucleotide 29uucuucuacc auccuaucaa u
213021RNAUnknownDescription of Unknown TAS3 5'D7(+) smRNA
oligonucleotide 30uucuugaccu uguaagaccc c
213121RNAUnknownDescription of Unknown TAS3 5'D8(+) smRNA
oligonucleotide 31uucuugaccu uguaaggccu u
213221RNAUnknownDescription of Unknown miR168 oligonucleotide
32ucgcuuggug caggucggga a 213322RNAUnknownDescription of Unknown
miR828 oligonucleotide 33ucuugcuuaa augaguauuc ca
223421RNAUnknownDescription of Unknown miR393 oligonucleotide
34uccaaaggga ucgcauugau c 213522RNAUnknownDescription of Unknown
plant derived smRNA oligonucleotide 35uucgcuugca gagagaaauc ac
2236173DNAArtificial SequenceDescription of Artificial Sequence
Synthetic Endovirus 5' UTR polynucleotide 36tgaaatgtct tgtacgacca
tttcaaattt atgtaaattg aacgcagcaa caacaggggg 60ggggcggaga cgcccccccc
cttttaaaaa taaaaatgat caaatcaagt acgatcttgg 120tttgatcaaa
tcaaaaaccc ctgttataaa agggtttttg aaaagaaggt acc
1733768DNAArtificial SequenceDescription of Artificial Sequence
Synthetic Endovirus 3' UTR polynucleotide 37aatattataa ctaactctgt
tttgtcaatt tatttttaaa aggatggggc acccctccca 60aaccccgg
6838571DNAArtificial SequenceDescription of Artificial Sequence
Synthetic Endovirus RNA Dependent RNA Polymerase (RDRP) coding
sequence and cloning sites 38atgatagtat ggcaaaggaa agcagtttgt
agtttatttg ccaaattgtt tgtaagatgc 60aaagacagac tgaaaacttt acttgtggat
catatacttt acgtagatgg attgagacca 120gatgaaatat cagccaaatt
aagacaaata tctgatgtat ttggattttt tgaaaacgac 180ctgactaagc
aagatagaca aactgacaaa cccattttag aagtggaaat gttgatgtat
240cttatgttgg gcgttcatcc taacatcata tctagttggc gttcaagtca
tgatgattgg 300agattcaaat ctacaaatta ttggggtaag agcacggcaa
tgagattaac gggacaagct 360acaaccgcac taggaaattg tatcactaat
atgcaagtac actcaaaatt tgtaatcaaa 420aataaatatt ggttaaagtt
tgctttattt cttggggatg atatgtgtat gggtttctca 480cacaagccaa
acacacagca cttacgccag gatatagctt gtaaatttaa tatgcaaagt
540aaagtgagaa ttcctcgagg gatccaagct t 5713936DNAArtificial
SequenceDescription of Artificial Sequence Synthetic T7 Promoter
and restriction sites for cloning 39gagctcctaa tacgactcac
tatagggaga gggccc 364024DNAArtificial SequenceDescription of
Artificial Sequence Synthetic T7 Promoter reverse complement
oligonucleotide sequence 40tctccctata gtgagtcgta ttag
244121RNACitrus x paradisi 41ggugcuaucc uaccugagcu u
214221RNAManihot esculenta 42ggugcuaucc uaccugagcu u
214321RNAPopulus tremula x Populus tremuloides 43ggugcuaucc
uaccugagcu u 214421RNAPinus taeda 44gacgcuaucc ccucugagcu u
214521RNAPopulus trichocarpa 45ggugcuaucc uaccugagcu u
214621RNAPinus taeda 46gacgcuaucc ccucugagcu u 214721RNAZea mays
47gguguuaucc ugauugagcu u 214821RNAGossypium raimondii 48ggugcuaucc
caccugagcu u 214921RNAVitis vinifera 49ggugcuaucc uaccugagcu u
215021RNAEuphorbia esula 50ggugcuaucc uaucugagcu u
215121RNALycopersicon esculentum 51ggugcuaucc uaccugagcu u
215221RNAZea mays 52gguguuaucc cgacugaacu u 215321RNAVitis vinifera
53ggugcuaucc uaccugagcu u 215421RNAGossypium raimondii 54ggugcuaucc
caccugagcu u 215521RNATheobroma cacao 55ggugcuaucc uaccugagcu u
215621RNAPopulus euphratica 56ggugcuaucc uaccugagcu u
215721RNAHordeum vulgare 57gguguuaucc cgaaugagcu u 215821RNAManihot
esculenta 58ggugcuaucc uaccugagcu u 215921RNAMalus x domestica
59ggugcuaucc uaccugagcu u 216021RNAGlycine max 60ggugcuaucc
uaucugagcu u 216121RNASorghum bicolor 61ggcguuaucc ugauugagcu u
216221RNAPopulus trichocarpa 62ggugcuaucc uaccugagcu u
216321RNAVitis vinifera 63ggugcuaucc uaccugagcu u 216421RNACitrus x
paradisi x Poncirus trifoliata 64ggugcuaucc uaccugagcu u
216521RNABruguiera gymnorhiza 65ggugcuaucc uaccugagcu u
216621RNASorghum bicolor 66ggcguuaucc ugauugagcu u
216721RNATriticum aestivum 67gguguuaucc ugauugagcu u 216821RNAMalus
x domestica 68ggugcuaucc uagcugagcu u 216921RNAOryza sativa
69gguguuaucc uaacugagcu u 217021RNAZea mays 70gguguuaucc ugauugagcu
u 217121RNAPopulus tremula 71ggugcuaucc uaccugagcu u
217221RNAAntirrhinum majus 72ggugcuaucc uaucugagcu u
217321RNASorghum bicolor 73ggcguuaucc ugauugagcu u 217421RNAPopulus
trichocarpa x Populus deltoides 74ggugcuaucc uaccugagcu u
217521RNAMesembryanthemum crystallinum 75ggugcuaucc uaccugagcu u
217621RNAZea mays 76ggcguuaucc uaauugagcu u 217721RNAPopulus
trichocarpa 77ggugcuaucc uaccugagcu u 217821RNAPopulus trichocarpa
78ggugcuaucc uaccugagcu u 217921RNAAntirrhinum majus 79ggugcuaucc
uaucugagcu u 218021RNAZea mays 80ggcguuaucc uaauugagcu u
218121RNAPopulus deltoides 81ggugcuaucc uaccugagcu u
218221RNATriticum aestivum 82gguguuaucc ugauugagcu u
218321RNACitrus clementina 83ggugcuaucc uaccugagcu u
218421RNAGlycine soja 84ggugcuaucc uaucugagcu u 218521RNAManihot
esculenta 85ggugcuaucc uaccugagcu u 218621RNAPopulus trichocarpa x
Populus deltoides 86ggugcuaucc uaccugagcu u 218721RNAPopulus
deltoides 87ggugcuaucc uaccugagcu u 218821RNASorghum bicolor
88ggcguuaucc ugauugagcu u 218921RNASaccharum officinarum
89gguguuaucc caauugagcu u 219021RNAZea mays 90gguguuaucc ugauugagcu
u 219121RNAPopulus trichocarpa 91ggugcuaucc uaccugagcu u
219221RNAZea mays 92ggcguuaucc uaauugagcu u 219321RNAZea mays
93gguguuaucc cgacuaaacu u 219421RNATriticum aestivum 94gguguuaucc
ugauugagcu u 219521RNAPopulus trichocarpa 95ggugcuaucc uaccugagcu u
219621RNAPopulus deltoides 96ggugcuaucc uaccugagcu u
219721RNASaccharum officinarum 97gguguuaucc ugaucgagcu u
219821RNAArabidopsis thaliana 98gguguuaucc uaucugagcu u
219921RNAArabidopsis thaliana 99uccaaaugua gucacuuuca g
2110021RNAArabidopsis thaliana 100ccccaauguu guuacuuuca a
2110121RNAArabidopsis thaliana 101cccggaugua aucacuuuca g
2110221RNAArabidopsis thaliana 102cccugauguu guuacuuuca g
2110321RNAArabidopsis thaliana 103uccaaaugua gucacuuuca g
2110421RNAArabidopsis thaliana 104uccaaaugua gucacuuuca g
2110521RNAArabidopsis thaliana 105uccaaaugua gucacuuuca a
2110621RNAArabidopsis thaliana 106uccggaugua gucacuuuua g
2110721RNAArabidopsis thaliana 107uccaaaugua gucacuuuca a
2110821RNAArabidopsis thaliana 108cccugauguu gucacuuuca c
2110921RNAArabidopsis thaliana 109cccugaugua uuuacuuuca a
2111021RNAArabidopsis thaliana 110accugaugua aucacuuuca a
2111121RNAArabidopsis thaliana 111cccugaugua uucacuuuca g
2111221RNAArabidopsis thaliana 112gugacuuaca auacucuuau a
2111321RNAArabidopsis thaliana 113gugacuuaua auacucucau a
2111421RNAArabidopsis thaliana 114guuacauaua auacucucau a
2111521RNAArabidopsis thaliana 115gugacuuaca auacucuuau a
2111621RNAArabidopsis thaliana 116gauacauaua auacucucau a
2111721RNAArabidopsis thaliana 117gugacuuaca auacucuuau u
2111821RNAArabidopsis thaliana 118gugacauaua acacucucau u
2111921RNAArabidopsis thaliana 119guaacuuaua guauucucau u
2112021RNAArabidopsis thaliana 120guggcuuaua cuucucucau a
2112121RNAArabidopsis thaliana 121gugacuuaua auacgcuuau a
2112221RNAArabidopsis thaliana 122uucccgagcu gcaucaagcu a
2112322RNAArabidopsis thaliana 123gagacaaugc gaucccuuug ga
2212422RNAArabidopsis thaliana 124gagaccaugc gaucccuuug ga
2212522RNAArabidopsis thaliana 125gaaacaaugc gaucccuuug ga
2212622RNAArabidopsis thaliana 126ggucagagcg aucccuuugg ca
2212722RNAArabidopsis thaliana 127gaaacaaugc gaucccuuug ga
2212821RNAAcorus americanus 128aaacaaugcg aucccuuugg a
2112921RNAAnanas comosus 129aaacaaugcg aucccuuugg a
2113021RNAAnanas comosus 130aaacaaugcg aucccuuugg a
2113121RNAAquilegia formosa x Aquilegia pubescens 131aaacaaugcg
aucccuuugg a 2113221RNABrassica rapa subsp. pekinesis 132aaacaaugcg
aucccuuugg a 2113321RNACitrus sinensis 133aaucuaugag aucacuuugg a
2113421RNACoffea arabica 134aaacaaugcg aucccuuugg a
2113521RNACoffea canephora 135aaacaaugcg aucccuuugg a
2113621RNACoffea canephora 136aaacaaugcg aucccuuugg a
2113721RNACoffea canephora 137aaaccaugcu aucccuuugg a
2113821RNACoffea canephora 138aaacaaugcg aucccuuugg a
2113921RNACoffea canephora 139aaacaaugcg aucccuuugg a
2114021RNAEucalyptus tereticornis 140aaacaaugcg aucccuuugg a
2114121RNAGlycine max 141aaacaaugcg aucccuuugg a 2114221RNAGlycine
max 142aaacgaugcg aucccuuugg a 2114321RNAGlycine max 143aaacaaugcg
aucccuuugg a 2114421RNAGlycine max 144aaacaaugcg aucccuuugg a
2114521RNAGlycine max 145aaacaaugcg aucccuuugg a 2114621RNAGlycine
max 146aaacaaugcg aucccuuugg a 2114721RNAGlycine max 147aaacaaugcg
aucccuuugg a 2114821RNAGlycine max 148aaacaaugcg aucccuuugg a
2114921RNAGlycine max 149aaacaaugcg aucccuuugg a 2115021RNAGlycine
max 150aaacaaugcg aucccuuugg a 2115121RNAGlycine max 151aaacaaugcg
aucccuuugg a 2115221RNAGlycine max 152aaacaaugcg aucccuuugg a
2115321RNAGlycine max 153aaacaaugcg aucccuuugg a 2115421RNAGlycine
max 154aaacaaugcg aucccuuugg a 2115521RNAGlycine max 155aaacaaugcg
aucccuuugg a 2115621RNAGlycine max 156aaacaaugcg aucccuuugg a
2115721RNAGlycine max 157aaacaaugcg aucccuuugg a 2115821RNAGlycine
max 158aaacaaugcg aucccuuugg a 2115921RNAGlycine max 159aaacaaugcg
aucccuuugg a 2116021RNAGlycine soja 160aaacaaugcg aucccuuugg a
2116121RNAGossypium arboreum 161aaacaaugcg aucccuuugg a
2116221RNAGossypium hirsutum 162aaacaaugcg aucccuuugg a
2116321RNAGossypium hirsutum 163aaacaaugcg aucccuuugg a
2116421RNAGossypium hirsutum 164aaacaaugcg aucccuuugg a
2116521RNAHelianthus argophyllus 165agucaaugcg aucccugugg a
2116621RNAHelianthus argophyllus 166agucaaugcg aucccugugg a
2116721RNAHelianthus petiolaris 167aaucaaugag gucucuuugg a
2116821RNALactuca sativa 168aaucaaugag gucucuuugg a
2116921RNALactuca sativa 169aaucaaugag gucucuuugg a 2117021RNALotus
japonicus 170gcacgaggcg aucccuuugg a 2117121RNAMalus x domestica
171aagcaaugcg aucccuuugg a 2117221RNAMalus x domestica
172aaacaaugcg aucccuuugg a 2117321RNAMalus x domestica
173aagcaaugcg aucccuuugg a 2117421RNAMalus x domestica
174aagcaaugcg aucccuuugg a 2117521RNAMalus x domestica
175aagcaaugcg aucccuuugg a 2117621RNAMalus x domestica
176aaacaaugcg aucccuuugg a 2117721RNAMalus x domestica
177aagcaaugcg aucccuuugg a 2117821RNAMalus x domestica
178aagcaaugcg aucccuuugg a 2117921RNAMalus x domestica
179aagcaaugcg aucccuuugg a 2118021RNAMalus x domestica
180aaacaaugcg aucccuuugg a 2118121RNAMalus x domestica
181aaacacugcg aucccuuugg a 2118221RNAMalus x domestica
182aaacaaugcg aucccuuugg a 2118321RNAMalus x domestica
183aagcaaugcg aucccuuugg a 2118421RNAMalus x domestica
184aagcaaugcg aucccuuugg a 2118521RNAMalus x domestica
185aaacaaugcg aucccuuugg a 2118621RNAMalus x domestica
186aagcaaugcg aucccuuugg a 2118721RNAMalus x domestica
187aagcaaugcg aucccuuugg a 2118821RNAManihot esculenta
188aaacaaugcg aucccuuugg a 2118921RNAMedicago truncatula
189aaacaaugcg aucccuuugg a 2119021RNAMedicago truncatula
190aaacaaugcg aucccuuugg a 2119121RNAMedicago truncatula
191aaacaaugcg aucccuuugg a 2119221RNAMedicago truncatula
192aaacaaugcg aucccuuugg a 2119321RNAMedicago truncatula
193aaacaaugcg aucccuuugg a 2119421RNAMedicago truncatula
194aaacaaugcg aucccuuugg a 2119521RNAMedicago truncatula
195aaacaaugcg aucccuuugg a 2119621RNAPicea glauca 196aaucaaugcg
aucccuuugg a 2119721RNAPinus taeda 197aaucaaugcg aucccuuugg a
2119821RNAPinus taeda 198aaucaaugcg aucccuuugg a 2119921RNAPinus
taeda 199aaucaaugcg aucccuuugg a 2120021RNAPinus taeda
200aaucaaugcg aucccuuugg a 2120121RNAPinus taeda 201aaucaaugcg
aucccuuugg a 2120221RNAPoncirus trifoliata 202gaucagagcg aucccuuuga
a 2120321RNAPopulus tremula x Populus tremuloides 203aaacaaugcg
aucccuuugg a 2120421RNAPopulus tremula x Populus tremuloides
204aaacaaugcg aucccuuugg a 2120521RNAPopulus tremula x Populus
tremuloides 205aaacaaugcg aucccuuugg a 2120621RNAPopulus
tremuloides 206aaacaaugcg aucccuuugg a 2120721RNAPopulus
trichocarpa 207aaacaaugcg aucccuuugg a 2120821RNAPopulus
trichocarpa x Populus deltoides 208aaacaaugcg aucccuuugg a
2120921RNAPopulus trichocarpa x Populus deltoides 209aaacaaugcg
aucccuuugg a 2121021RNAPopulus x canadensis 210aaacaaugcg
aucccuuugg a 2121121RNAPrunus persica 211aaucuaugag aucacuuugg a
2121221RNARicinus communis 212aaacaaugcg aucccuuugg a
2121321RNASaruma henryi 213aaacaaugcg aucccuuugg a
2121421RNAZingiber officinale 214aaacaaugcg aucccuuugg a
2121521RNAZingiber officinale 215aaacaaugcg aucccuuugg a
2121621RNAArabidopsis thaliana 216cagauggugg aaaugggaua u
2121721RNAArabidopsis thaliana 217caaauggucg aaaugggaua u
2121821RNAArabidopsis thaliana 218caaauggugg aaauggggua u
2121921RNAArabidopsis thaliana 219caaauggucg aaaugggaua u
2122021RNAArabidopsis thaliana 220cagauggugg aaaugggaua u
2122121RNAArabidopsis thaliana 221caaauggugg aaauggggua u
2122221RNAArabidopsis thaliana 222caaauggugg aaauggggua u
2122321RNAArabidopsis thaliana 223caaauggugg gaaugggaua u
2122421RNAArabidopsis thaliana 224guugaucgua ugguagaaga a
2122521RNAArabidopsis thaliana 225aaggucuugc aaggucaaga a
2122621RNAArabidopsis thaliana 226agggucuugc aaggucaaga a
2122721RNACitrus x paradisi 227uugucuaucc cuccugagcu g
2122821RNAManihot esculenta 228uugucuaucc cuccugagcu a
2122921RNAPopulus tremula x Populus tremuloides 229uugucuaucc
cuccugagcu g 2123021RNAPinus taeda 230uacucuaucu cuccugagcu a
2123121RNAPopulus trichocarpa 231uugucuaucc cuccugagcu a
2123221RNAPinus taeda 232uacucuaucu cuccugagcu a 2123321RNAZea mays
233agcucuaucc cuucugagcu g 2123421RNAGossypium raimondii
234cugucuaucc cuccugagcu a 2123521RNAVitis vinifera 235uugucuaucc
cuccugagcu a 2123621RNAEuphorbia esula 236uugucuaucc cuccugagcu u
2123721RNALycopersicon esculentum 237uugucuaucc cuccugagcu g
2123821RNAZea mays 238ccuucuaucc cuccugagcu a 2123921RNAVitis
vinifera 239uugucuaucc cuccugagcu a 2124021RNAGossypium raimondii
240cugucuaucc cuccugagcu a 2124121RNATheobroma cacao 241cuugcuaucc
cuccugagcu g 2124221RNAPopulus euphratica 242uuggcuaucc cuccugagcu
g 2124321RNAHordeum vulgare 243cuuucuaucc cuccugagcu a
2124421RNAManihot esculenta 244uugucuaucc cuccugagcu a
2124521RNAMalus x domestica 245uugucuaucc cuccugagcu g
2124621RNAGlycine max 246uugucuaucc cuccugagcu g 2124721RNASorghum
bicolor 247augucuaucc cuucugagcu g 2124821RNAPopulus trichocarpa
248uugucuaucc cuccugagcu a 2124921RNAVitis vinifera 249uugucuaucc
cuccugagcu a 2125021RNACitrus x paradise x Poncirus trifoliata
250uugucuaucc cuccugagcu g 2125121RNABruguiera gymnorhiza
251uugucuaucc cuccugagcu g 2125221RNASorghum bicolor 252augucuaucc
cuucugagcu g 2125321RNATriticum aestivum 253ccaucuaucc cuccugagcu a
2125421RNAMalus x domestica 254uugucuaucc cuccugagcu g
2125521RNAOryza sativa 255cggucuaucc cuccugagcu g 2125621RNAZea
mays 256agcucuaucc cuucugagcu u 2125721RNAPopulus tremula
257uugucuaucc cuccugagcu g 2125821RNAAntirrhinum majus
258uugucuaucc cuccugagcu a 2125921RNASorghum bicolor 259augucuaucc
cuucugagcu g 2126021RNAPopulus trichocarpa x Populus deltoides
260uuaucuaucc cuccugagcu a 2126121RNAMesembryanthemum crystallinum
261gugucuaucc cuccugagcu a 2126221RNAZea mays 262augucuaucc
cuucugagcu g 2126321RNAPopulus trichocarpa 263uuaucuaucc cuccugagcu
a 2126421RNAPopulus trichocarpa 264uugucuaucc cuccugagcu a
2126521RNAAntirrhinum majus 265uugucuaucc cuccugagcu a
2126621RNAZea mays 266augucuaucc cuucugagcu g 2126721RNAPopulus
deltoides 267uugucuaucc cuccugagcu a 2126821RNATriticum aestivum
268ccaucuaucc cuccggagcu a 2126921RNACitrus clementina
269uugucuaucc cuccugagcu g 2127021RNAGlycine soja 270uugucuaucc
cuccugagcu g 2127121RNAManihot esculenta 271uugucuaucc cuccugagcu a
2127222RNAPopulus trichocarpa x Populus
deltoidesmodified_base(3)..(5)a, c, u, g, unknown or
othermisc_feature(3)..(5)n is a, c, g, or u 272uunnnuaucc
cuccugagcu au 2227321RNAPopulus deltoides 273uuaucuaucc cuccugagcu
g 2127421RNASorghum bicolor 274augucuaucc cuucugagcu g
2127521RNASaccharum officinarum 275augucuaucc cuucugagcu g
2127621RNAZea mays 276agcucuaucc cuucugagcu g 2127721RNAPopulus
trichocarpa 277uuaucuaucc cuccugagcu a 2127821RNAZea mays
278augucuaucc cuucugaacu g 2127921RNAZea mays 279ccuucuaucc
cuccugagcu a 2128021RNATriticum aestivum 280ccaucuaucc cuccugagcu a
2128121RNAPopulus trichocarpa 281uuaucuaucc cuccugagcu a
2128221RNAPopulus deltoides 282uugucuaucc cuccugagcu a
2128321RNASaccharum officinarum 283augucuaucc
cuucugagcu a 2128421RNAArabidopsis thaliana 284uugucuaucc
cuccugagcu a 2128521RNAArabidopsis thaliana 285guuacuuaca
acacucucau a 2128621RNAArabidopsis thaliana 286guuacuuaca
acacucucau a 2128721RNAArabidopsis thaliana 287agggucuugc
aaggucaaga a 2128821RNAArabidopsis thaliana 288aaggucuugc
aaggucaaga a 21289282DNAUnknownNot known 289ctaatacgac tcactatagg
gagatttccg tcctccttga aatcaattcc cttaagctcg 60atcctgttga cgagggtgtc
tccctcaaac ttgacttcag cacgtgtctt gtagttcccg 120tcgtccttga
aagagatggt cctctcctgc acgtatccct caggcatggc gctcttgaag
180aagtcgtgcc gcttcatatg atctgggtat cttgaaaagc attgaacacc
ataagagaaa 240gtagtgacaa gtgttggctc tccctatagt gagtcgtatt ag
282290325DNAUnknownNot known 290ctaatacgac tcactatagg gagaggtgct
atcctacctg agctttttcc gtcctccttg 60aaatcaattc ccttaagctc gatcctgttg
acgagggtgt ctccctcaaa cttgacttca 120gcacgtgtct tgtagttccc
gtcgtccttg aaagagatgg tcctctcctg cacgtatccc 180tcaggcatgg
cgctcttgaa gaagtcgtgc cgcttcatat gatctgggta tcttgaaaag
240cattgaacac cataagagaa agtagtgaca agtgttggcc cttgtctatc
cctcctgagc 300ttctccctat agtgagtcgt attag 325291368DNAUnknownNot
known 291ctaatacgac tcactatagg gagaagctca ggagggatag acaaggggtg
ctatcctacc 60tgagcttttt ccgtcctcct tgaaatcaat tcccttaagc tcgatcctgt
tgacgagggt 120gtctccctca aacttgactt cagcacgtgt cttgtagttc
ccgtcgtcct tgaaagagat 180ggtcctctcc tgcacgtatc cctcaggcat
ggcgctcttg aagaagtcgt gccgcttcat 240atgatctggg tatcttgaaa
agcattgaac accataagag aaagtagtga caagtgttgg 300caagctcagg
taggatagca ccccttgtct atccctcctg agcttctccc tatagtgagt 360cgtattag
368292301DNAUnknownNot known 292ctaatacgac tcactatagg gagaggtgct
atcctacctg agctttttcc gtcctccttg 60aaatcaattc ccttaagctc gatcctgttg
acgagggtgt ctccctcaaa cttgacttca 120gcacgtgtct tgtagttccc
gtcgtccttg aaagagatgg tcctctcctg cacgtatccc 180tcaggcatgg
cgctcttgaa gaagtcgtgc cgcttcatat gatctgggta tcttgaaaag
240cattgaacac cataagagaa agtagtgaca agtgttggcc cttgtctatc
cctcctgagc 300t 301293258DNAUnknownNot known 293ctaatacgac
tcactatagg gagagccaac acttgtcact actttctctt atggtgttca 60atgcttttca
agatacccag atcatatgaa gcggcacgac ttcttcaaga gcgccatgcc
120tgagggatac gtgcaggaga ggaccatctc tttcaaggac gacgggaact
acaagacacg 180tgctgaagtc aagtttgagg gagacaccct cgtcaacagg
atcgagctta agggaattga 240tttcaaggag gacggaaa 258294375DNAUnknownNot
known 294ctaatacgac tcactatagg gagaggtgct atcctacctg agctttttcc
gtcctccttg 60aaatcaattc ccttaagctc gatcctgttg acgagggtgt ctccctcaaa
cttgacttca 120gcacgtgtct tgtagttccc gtcgtccttg aaagagatgg
tcctctcctg cacgtatccc 180tcaggcatgg cgctcttgaa gaagtcgtgc
cgcttcatat gatctgggta tcttgaaaag 240cattgaacac cataagagaa
agtagtgaca agtgttggcg ctactggact ttcgccatct 300agggtgcagc
attcgggctg cttcgcctac cttgtctatc cctcctgagc ttctccctat
360agtgagtcgt attag 375295344DNAUnknownNot known 295ctaatacgac
tcactatagg gagaagctca ggagggatag acaaggggtg ctatcctacc 60tgagcttttt
ccgtcctcct tgaaatcaat tcccttaagc tcgatcctgt tgacgagggt
120gtctccctca aacttgactt cagcacgtgt cttgtagttc ccgtcgtcct
tgaaagagat 180ggtcctctcc tgcacgtatc cctcaggcat ggcgctcttg
aagaagtcgt gccgcttcat 240atgatctggg tatcttgaaa agcattgaac
accataagag aaagtagtga caagtgttgg 300caagctcagg taggatagca
ccccttgtct atccctcctg agct 344296394DNAUnknownNot known
296ctaatacgac tcactatagg gagaagctca ggagggatag acaaggggtg
ctatcctacc 60tgagcttgcc aacacttgtc actactttct cttatggtgt tcaatgcttt
tcaagatacc 120cagatcatat gaagcggcac gacttcttca agagcgccat
gcctgaggga tacgtgcagg 180agaggaccat ctctttcaag gacgacggga
actacaagac acgtgctgaa gtcaagtttg 240agggagacac cctcgtcaac
aggatcgagc ttaagggaat tgatttcaag gaggacggaa 300aaagctcagg
taggatagca ccccttgtct atccctcctg agcttaggcg aagcagcccg
360aatgctgcac cctagatggc gaaagtccag tagc 394297347DNAUnknownNot
known 297ctaatacgac tcactatagg gagaggtgct atcctacctg agctttttcc
gtcctccttg 60aaatcaattc ccttaagctc gatcctgttg acgagggtgt ctccctcaaa
cttgacttca 120gcacgtgtct tgtagttccc gtcgtccttg aaagagatgg
tcctctcctg cacgtatccc 180tcaggcatgg cgctcttgaa gaagtcgtgc
cgcttcatat gatctgggta tcttgaaaag 240cattgaacac cataagagaa
agtagtgaca agtgttggct cgcaggagag atgacaccag 300accttgtcta
tccctcctga gcttctccct atagtgagtc gtattag 347298282DNALycopersicon
esculentum 298ctaatacgac tcactatagg gagatttctc accgcttttt
tttttctgtt gtgtattctc 60ttttttgact tgttgccttt cgttcctcta cctaccccat
tcttcttgac cttgtaagac 120cttttcttga ccttgtaaga ccccgtgtta
tctcttacgt ctttatgttt tgtttttttg 180caaatcttac gtcatgactt
cttcatgtaa gctttgtttg gtctccttct tctttcctac 240tcaactctcg
ttctcctttc tccctatagt gagtcgtatt ag 282299325DNAUnknownNot known
299ctaatacgac tcactatagg gagaggtgct atcctacctg agctttttct
caccgctttt 60ttttttctgt tgtgtattct cttttttgac ttgttgcctt tcgttcctct
acctacccca 120ttcttcttga ccttgtaaga ccttttcttg accttgtaag
accccgtgtt atctcttacg 180tctttatgtt ttgttttttt gcaaatctta
cgtcatgact tcttcatgta agctttgttt 240ggtctccttc ttctttccta
ctcaactctc gttctccttc cttgtctatc cctcctgagc 300ttctccctat
agtgagtcgt attag 32530021DNALycopersicon esculentum 300aactacattt
ctcccttcca g 2130120DNALycopersicon esculentum 301tcacaacaaa
cacctgctac 2030219DNALycopersicon esculentum 302cgaaagaacc
atctactcc 1930319DNALycopersicon esculentum 303aaagcctctc caactcaac
1930421DNALycopersicon esculentum 304caacccaaag gctgcaaaaa c
2130521DNALycopersicon esculentum 305ggatgcgaca ctcatcgtta g
2130620DNAAequorea victoria 306cgtcgtcctt gaaagagatg
2030720DNAAequorea victoria 307gagccaacac ttgtcactac
2030822DNALycopersicon esculentum 308gctaagaacg ctggacctaa tg
2230920DNALycopersicon esculentum 309agaatagcat ccggtctcag
2031024DNALycopersicon esculentum 310aacaggtggt gctcgactat gact
2431123DNALycopersicon esculentum 311tgctttcgac agtttcactt cca
23312236DNAArtificial SequenceSynthetic Construct 312gccacttgca
aagtcccgct agtgccttgt ccagttgcaa ccacctgttg atccgcatca 60cgcagttcaa
cgctgacatc accattggcc accacctgcc agtcaacaga cgcgtggtta
120cagtcttgcg cgacatgcgt caccacggtg atatcgtcca cccaggtgtt
cggcgtggtg 180tagagcatta cgctgcgatg gattccggca tagttaaaga
aatcatggaa gtaagc 236313278DNAArtificial SequenceSynthetic
Construct 313ggtgctatcc tacctgagct tccacttgca aagtcccgct agtgccttgt
ccagttgcaa 60ccacctgttg atccgcatca cgcagttcaa cgctgacatc accattggcc
accacctgcc 120agtcaacaga cgcgtggtta cagtcttgcg cgacatgcgt
caccacggtg atatcgtcca 180cccaggtgtt cggcgtggtg tagagcatta
cgctgcgatg gattccggca tagttaaaga 240aatcatggaa gtaagccttg
tctatccctc ctgagctc 278314277DNAArtificial SequenceSynthetic
Construct 314ggtgctatcc tacctgagct ttttccgtcc tccttgaaat caattccctt
aagctcgttc 60ctgttgacga gggtgtctcc ctcaaacttg acttcagcat gtgtcttgta
gttcccgtcg 120ttcttgaaag agatggtcct ctcctgcacg tatccctcag
gcatggcgct cttgaagaag 180tcgtgccgct tcatatgatc tgggtatctt
gaaaagcatt gaacaccata agagaaagta 240gtgacaagtg ttggcccttg
tctatccctc ctgagct 277315277DNAArtificial SequenceSynthetic
Construct 315ggtgctatcc tacctgagct ttttctcacc gctttttttt ttctgttgtg
tattctcttt 60ccgtcctcct tgaaatcagt tcctctacct accccattct tcccttaagc
tcgatcctgt 120ttgacttcag cacgtgtctt ggtgttatct cttacgtctt
tatgttttgt ttttttgcaa 180atcttacgtc atgacttctt catgttcccg
tcgtccttga aagagcttct ttcctactca 240actctcgttc tccttccttg
tctatccctc ctgagct 277316277DNAArtificial SequenceSynthetic
Construct 316ggtgctatcc tacctgagct ttttccgtcc tccttgtaat caattccctt
aagctcgttc 60ctgttgacga gggtgtcttc ctcaaacttg acttcagcat gtgtcttgta
gttcccgtcg 120tccttgaaag agatggtcct ctcctgcacg tatccctcag
gcttggcgct cttgaagaag 180tcgtgccgct tcatatgatc tgggtatctt
gaaaagcatt gaacatcata agagaaagta 240gtgacatgtg ttggcccttg
tctatccctc ctgagct 277317277DNAArtificial SequenceSynthetic
Construct 317ggtgctatcc tacctgagct ttttctcacc gctttttaat ggttgtctgg
taaaaggtcg 60ccaattggag tattttgttg ataatgatca gcgagttgct cttcgatgtt
gtggcgggtc 120ttgaagttgg ctttgatgcc gttcttttgc ttgtcggcca
tgtgtatacg ttgtgggagt 180tgtttgtatt ccaacttgtg gccgtgtttc
cgtcctcctt gaaatttccc ttaagctcga 240tcctgtgttc tccttccttg
tctatccctc ctgagct 27731821DNALycopersicon esculentum 318ggtgctatcc
tacctgagct t 2131922DNALycopersicon esculentum 319ccttgtctat
ccctcttgag ct 22320282DNAAequorea victoria 320ctaatacgac tcactatagg
gagatttccg tcctccttga aatcaattcc cttaagctcg 60atcctgttga cgagggtgtc
tccctcaaac ttgacttcag cacgtgtctt gtagttcccg 120tcgtccttga
aagagatggt cctctcctgc acgtatccct caggcatggc gctcttgaag
180aagtcgtgcc gcttcatatg atctgggtat cttgaaaagc attgaacacc
ataagagaaa 240gtagtgacaa gtgttggctc tccctatagt gagtcgtatt ag
282321325DNAArtificial SequenceLycopersicon esculentum and Aequorea
victoria 321ctaatacgac tcactatagg gagaggtgct atcctacctg agctttttcc
gtcctccttg 60aaatcaattc ccttaagctc gatcctgttg acgagggtgt ctccctcaaa
cttgacttca 120gcacgtgtct tgtagttccc gtcgtccttg aaagagatgg
tcctctcctg cacgtatccc 180tcaggcatgg cgctcttgaa gaagtcgtgc
cgcttcatat gatctgggta tcttgaaaag 240cattgaacac cataagagaa
agtagtgaca agtgttggcc cttgtctatc cctcctgagc 300ttctccctat
agtgagtcgt attag 325322368DNAArtificial SequenceLycopersicon
esculentum and Aequorea victoria 322ctaatacgac tcactatagg
gagaagctca ggagggatag acaaggggtg ctatcctacc 60tgagcttttt ccgtcctcct
tgaaatcaat tcccttaagc tcgatcctgt tgacgagggt 120gtctccctca
aacttgactt cagcacgtgt cttgtagttc ccgtcgtcct tgaaagagat
180ggtcctctcc tgcacgtatc cctcaggcat ggcgctcttg aagaagtcgt
gccgcttcat 240atgatctggg tatcttgaaa agcattgaac accataagag
aaagtagtga caagtgttgg 300caagctcagg taggatagca ccccttgtct
atccctcctg agcttctccc tatagtgagt 360cgtattag 368323301DNAArtificial
SequenceLycopersicon esculentum and Aequorea victoria 323ctaatacgac
tcactatagg gagaggtgct atcctacctg agctttttcc gtcctccttg 60aaatcaattc
ccttaagctc gatcctgttg acgagggtgt ctccctcaaa cttgacttca
120gcacgtgtct tgtagttccc gtcgtccttg aaagagatgg tcctctcctg
cacgtatccc 180tcaggcatgg cgctcttgaa gaagtcgtgc cgcttcatat
gatctgggta tcttgaaaag 240cattgaacac cataagagaa agtagtgaca
agtgttggcc cttgtctatc cctcctgagc 300t 301324258DNAArtificial
SequenceLycopersicon esculentum and Aequorea victoria 324ctaatacgac
tcactatagg gagagccaac acttgtcact actttctctt atggtgttca 60atgcttttca
agatacccag atcatatgaa gcggcacgac ttcttcaaga gcgccatgcc
120tgagggatac gtgcaggaga ggaccatctc tttcaaggac gacgggaact
acaagacacg 180tgctgaagtc aagtttgagg gagacaccct cgtcaacagg
atcgagctta agggaattga 240tttcaaggag gacggaaa 258325375DNAArtificial
SequenceLycopersicon esculentum and Aequorea victoria 325ctaatacgac
tcactatagg gagaggtgct atcctacctg agctttttcc gtcctccttg 60aaatcaattc
ccttaagctc gatcctgttg acgagggtgt ctccctcaaa cttgacttca
120gcacgtgtct tgtagttccc gtcgtccttg aaagagatgg tcctctcctg
cacgtatccc 180tcaggcatgg cgctcttgaa gaagtcgtgc cgcttcatat
gatctgggta tcttgaaaag 240cattgaacac cataagagaa agtagtgaca
agtgttggcg ctactggact ttcgccatct 300agggtgcagc attcgggctg
cttcgcctac cttgtctatc cctcctgagc ttctccctat 360agtgagtcgt attag
375326344DNAArtificial SequenceLycopersicon esculentum and Aequorea
victoria 326ctaatacgac tcactatagg gagaagctca ggagggatag acaaggggtg
ctatcctacc 60tgagcttttt ccgtcctcct tgaaatcaat tcccttaagc tcgatcctgt
tgacgagggt 120gtctccctca aacttgactt cagcacgtgt cttgtagttc
ccgtcgtcct tgaaagagat 180ggtcctctcc tgcacgtatc cctcaggcat
ggcgctcttg aagaagtcgt gccgcttcat 240atgatctggg tatcttgaaa
agcattgaac accataagag aaagtagtga caagtgttgg 300caagctcagg
taggatagca ccccttgtct atccctcctg agct 344327394DNAArtificial
SequenceLycopersicon esculentum and Aequorea victoria 327ctaatacgac
tcactatagg gagaagctca ggagggatag acaaggggtg ctatcctacc 60tgagcttgcc
aacacttgtc actactttct cttatggtgt tcaatgcttt tcaagatacc
120cagatcatat gaagcggcac gacttcttca agagcgccat gcctgaggga
tacgtgcagg 180agaggaccat ctctttcaag gacgacggga actacaagac
acgtgctgaa gtcaagtttg 240agggagacac cctcgtcaac aggatcgagc
ttaagggaat tgatttcaag gaggacggaa 300aaagctcagg taggatagca
ccccttgtct atccctcctg agcttaggcg aagcagcccg 360aatgctgcac
cctagatggc gaaagtccag tagc 394328347DNAArtificial
SequenceLycopersicon esculentum and Aequorea victoria 328ctaatacgac
tcactatagg gagaggtgct atcctacctg agctttttcc gtcctccttg 60aaatcaattc
ccttaagctc gatcctgttg acgagggtgt ctccctcaaa cttgacttca
120gcacgtgtct tgtagttccc gtcgtccttg aaagagatgg tcctctcctg
cacgtatccc 180tcaggcatgg cgctcttgaa gaagtcgtgc cgcttcatat
gatctgggta tcttgaaaag 240cattgaacac cataagagaa agtagtgaca
agtgttggct cgcaggagag atgacaccag 300accttgtcta tccctcctga
gcttctccct atagtgagtc gtattag 347329282DNALycopersicon esculentum
329ctaatacgac tcactatagg gagatttctc accgcttttt tttttctgtt
gtgtattctc 60ttttttgact tgttgccttt cgttcctcta cctaccccat tcttcttgac
cttgtaagac 120cttttcttga ccttgtaaga ccccgtgtta tctcttacgt
ctttatgttt tgtttttttg 180caaatcttac gtcatgactt cttcatgtaa
gctttgtttg gtctccttct tctttcctac 240tcaactctcg ttctcctttc
tccctatagt gagtcgtatt ag 282330325DNALycopersicon esculentum
330ctaatacgac tcactatagg gagaggtgct atcctacctg agctttttct
caccgctttt 60ttttttctgt tgtgtattct cttttttgac ttgttgcctt tcgttcctct
acctacccca 120ttcttcttga ccttgtaaga ccttttcttg accttgtaag
accccgtgtt atctcttacg 180tctttatgtt ttgttttttt gcaaatctta
cgtcatgact tcttcatgta agctttgttt 240ggtctccttc ttctttccta
ctcaactctc gttctccttc cttgtctatc cctcctgagc 300ttctccctat
agtgagtcgt attag 325331236DNAArtificial SequenceEscherichia coli
and Lycopersicon esculentum 331gccacttgca aagtcccgct agtgccttgt
ccagttgcaa ccacctgttg atccgcatca 60cgcagttcaa cgctgacatc accattggcc
accacctgcc agtcaacaga cgcgtggtta 120cagtcttgcg cgacatgcgt
caccacggtg atatcgtcca cccaggtgtt cggcgtggtg 180tagagcatta
cgctgcgatg gattccggca tagttaaaga aatcatggaa gtaagc
236332278DNAArtificial SequenceE. coli and Lycopersicon esculentum
332ggtgctatcc tacctgagct tccacttgca aagtcccgct agtgccttgt
ccagttgcaa 60ccacctgttg atccgcatca cgcagttcaa cgctgacatc accattggcc
accacctgcc 120agtcaacaga cgcgtggtta cagtcttgcg cgacatgcgt
caccacggtg atatcgtcca 180cccaggtgtt cggcgtggtg tagagcatta
cgctgcgatg gattccggca tagttaaaga 240aatcatggaa gtaagccttg
tctatccctc ctgagctc 278
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015
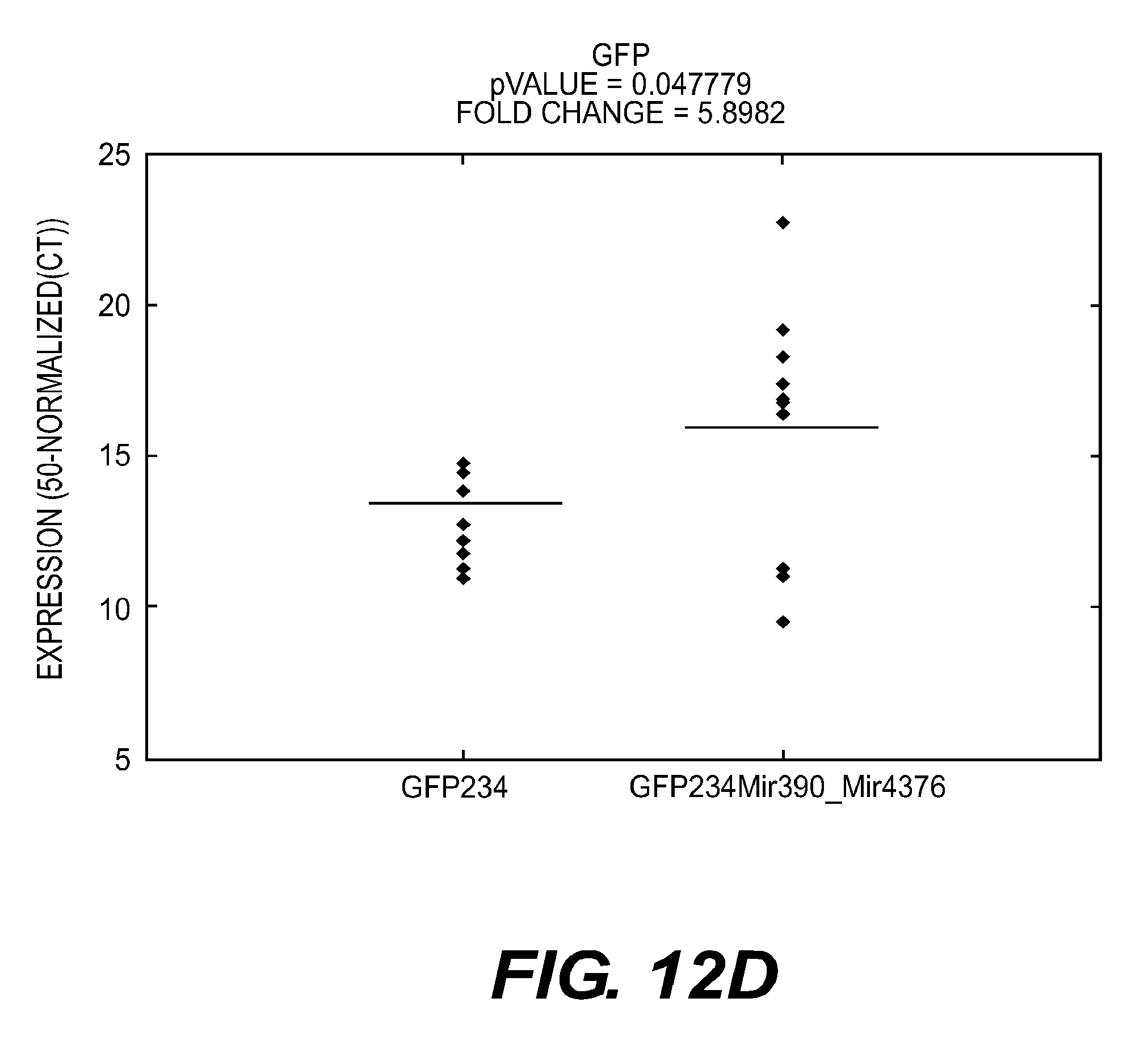
D00016

D00017

D00018

D00019

D00020

D00021

D00022

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.