Attenuated Mannheimia Haemolytica Strains
Briggs; Robert E. ; et al.
U.S. patent application number 16/017314 was filed with the patent office on 2019-01-03 for attenuated mannheimia haemolytica strains. The applicant listed for this patent is Biotechnology Research and Development Corporation, The United States of America, as represented by the Secretary of Agriculture, The United States of America, as represented by the Secretary of Agriculture. Invention is credited to Robert E. Briggs, Fred M. Tatum.
| Application Number | 20190000954 16/017314 |
| Document ID | / |
| Family ID | 50685318 |
| Filed Date | 2019-01-03 |




| United States Patent Application | 20190000954 |
| Kind Code | A1 |
| Briggs; Robert E. ; et al. | January 3, 2019 |
ATTENUATED MANNHEIMIA HAEMOLYTICA STRAINS
Abstract
The present invention provides attenuated M. haemolitica strains that elicit an immune response in animal against M. haemolitica, compositions comprising said strains, methods of vaccination against M. haemolitica, and kits for use with such methods and compositions. The invention further provides multi-valent vaccines, which provide protective immunity when administered in an effective amount to animals susceptible to "shipping fever" or bovine respiratory disease.
| Inventors: | Briggs; Robert E.; (Boone, IA) ; Tatum; Fred M.; (Nevada, IA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Family ID: | 50685318 | ||||||||||
| Appl. No.: | 16/017314 | ||||||||||
| Filed: | June 25, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15281154 | Sep 30, 2016 | |||
| 16017314 | ||||
| 14934737 | Nov 6, 2015 | |||
| 15281154 | ||||
| 14075169 | Nov 8, 2013 | 9370561 | ||
| 14934737 | ||||
| 61723979 | Nov 8, 2012 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | A61K 2039/522 20130101; A61K 2039/552 20130101; C07K 14/285 20130101; A61P 31/04 20180101; A61K 2039/543 20130101; A61K 2039/70 20130101; C12N 1/36 20130101; A61K 39/102 20130101 |
| International Class: | A61K 39/102 20060101 A61K039/102; C12N 1/36 20060101 C12N001/36; C07K 14/285 20060101 C07K014/285 |
Claims
1-56. (canceled)
57. A bacterial composition comprising an attenuated Mannheimia haemolytica (M. haemolytica) A1 strain and an attenuated M. haemolytica A6 strain; wherein both the A1 and A6 strains contain nucleic acid deletions in their respective leukotoxin A (lktA) genes, which deletions have rendered the strains attenuated relative to the virulent parental strains A1 and A6 from which the attenuated strains A1 and A6 were produced.
58. The bacterial composition of claim 57, consisting essentially of the attenuated strains.
59. The bacterial composition of claim 57, further comprising an adjuvant.
60. The bacterial composition of claim 57, wherein said bacterial composition comprises from about 1.19.times.10.sup.6 to 1.19.times.10.sup.7 CFU of the attenuated M. haemolytica A1 strain and from about 9.2.times.10.sup.5 to 9.2.times.10.sup.6 CFU of the attenuated M. haemolytica A6 strain.
61. The bacterial composition of claim 57, further comprising a vehicle, diluent or excipient and wherein said bacterial composition comprises from about 1.19.times.10.sup.6 to 1.19.times.10.sup.7 CFU of the attenuated M. haemolytica A1 strain and from about 9.2.times.10.sup.5 to 9.2.times.10.sup.6 CFU of the attenuated M. haemolytica A6 strain.
62. The bacterial composition of claim 61, further comprising an adjuvant.
63. The bacterial composition of claim 62, wherein the adjuvant is inactivated bacteria, inactivated virus, fractions of inactivated bacteria, bacterial lipopolysaccharides, bacterial toxins, or derivatives or combinations thereof.
64. The bacterial composition of claim 57, further comprising at least one additional bovine pathogen other than M. haemolytica.
65. The bacterial composition of claim 64, wherein said at least one additional bovine pathogen other than M. haemolytica is selected from the group consisting of Pasteurella multocida, Histophilus somni, and a combination thereof.
Description
[0001] This application incorporates by reference the contents of a 61.8 kb text file created on Sep. 30, 2016 and named "00848800006sequencelisting.txt," which is the sequence listing for this application.
FIELD OF THE INVENTION
[0002] The present invention relates generally to attenuated bacterial vaccines, particularly those providing broad, safe, and effective protection to production animals against infections/disease caused by gram-negative bacteria, including Mannheimia (Pasteurella) haemolytica. The invention further relates to methods of producing the attenuated bacteria, and to PCR methods for differentiating among M. haemolytica serotypes AI and A6, in vivo.
[0003] The invention accordingly relates to immunogenic or vaccine compositions comprising the bacteria of the invention; e.g., live attenuated bacteria. The bacteria also could be inactivated in the compositions, but it may be advantageous that the bacteria are live attenuated M. haemolytica bacteria, either alone, or combined with other bacteria such as Haemophilus somnus and/or Pasteurella multocida. The invention therefore further relates to methods for preparing and/or formulating such compositions; e.g., culturing or growing or propagating the bacteria on or in suitable medium, harvesting the bacteria, optionally inactivating the bacteria, and optionally admixing the bacteria with a suitable veterinarily or pharmaceutically acceptable carrier, excipient, diluent or vehicle and/or an adjuvant and/or stabilizer. Thus, the invention also relates to the use of the bacteria in formulating such compositions.
BACKGROUND OF THE INVENTION
[0004] M. haemolytica is a gram negative bacterium normally found in the upper respiratory tract of healthy cattle, sheep and wild sheep. M. haemolytica descends into the lungs when cattle experience stress such as shipping, weaning, overcrowding, or viral infections and causes fibrinous and necrotizing bronchopneumonia, a chief component of the bovine respiratory disease complex (BRDC). Economic losses due to BRDC in North America is >$1 billion annually (Bowland and Shewen, 2000). M. haemolytica is the bacterium most commonly isolated from the lungs of cattle affected with BRDC. M. haemolytica serotype AI is responsible for approximately 60% of shipping fever, whereas serotypes A6 and A2 account for 26% and 7% respectively (Al-Ghamdi et al., 2000; Purdy et al., 1997). Both M. haemolytica A1 and A6 account for >85% of BRDC cases involving bacterial pathogens.
[0005] The vaccines currently available in the market against M. haemolytica infections are only moderately protective against shipping fever of beef cattle but generally ineffective against neonatal dairy calf pneumonia (Virtala et al., 1996; Rice et al., 2007). The major cause of severe bacterial pneumonia in feedlot and neonatal dairy cattle is M. haemolytica serotype A1 followed by serotype A6 (Schreuer et al., 2000, Rice et al., 2007).
[0006] Experimental evaluation of all the commercial M. haemolytica A1 vaccines used in feedlot showed only partial protection in 50% of the studies (Perino and Hunsaker, 1997). Furthermore, cross-protection against M. haemolytica serotypes (either A6 or A2) has been difficult to achieve using conventional vaccine preparations (Purdy et al., 1993; Sabri et al., 2000). Therefore, an efficacious vaccine against M. haemolytica serotypes A1 and A6 could significantly improve dairy/beef production.
[0007] Effective immunity against M. haemolytica is multifaceted. Neutralizing Antibodies against exotoxin leukotoxin A (LktA) and surface antigens are necessary for protective immunity against M. haemolytica (Shewen and Wilkie, 1988). Due to the complex genetic machinery involved in controlling the expression of various M. haemolytica virulence factors, the specific surface antigens that are important in stimulating immunity have not been clearly determined (Lawrence et al, 2010). However, M. haemolytica outer membrane proteins (OMPs) have been implicated in stimulating immunity against surface antigens (Confer et al., 2003, Morton et al., 1995; Potter et al., 1999).
[0008] Intranasal immunization of cattle has been pursued for a while using bovine herpesvirus-1 (BoHV-1), bovine respiratory syncytial virus (BRSV) and infectious bovine rhinotracheitis virus (IBR) (Ellis et al., 2007; Muylkens et al., 2007). Commercially available Pfizer's INFORCE 3 when administered intranasally claims to prevent BRSV and also aids in the prevention of respiratory disease caused by IBR and bovine parainfluenza virus type 3 (PI3).
In an experimental study when a modified live leukotoxin deficient M. haemolytica mutant was administered intranasally in weaned beef feedlot calves, it resulted in reduced nasopharyngeal colonization with wild type M. haemolytica compared to non-vaccinated control calves (Frank et al., 2003). Although intranasal vaccination and leukotoxin deficient M. haemolytica are known, inventors are aware of no M. haemolytica vaccines successfully combining these concepts.
SUMMARY OF THE INVENTION
[0009] An object of the present disclosure is to provide effective vaccines comprising attenuated M. haemolytica serotypes A1 & A6. Relative to a parent M. haemolytica serotype A1 or A6 strain, the attenuated strains may have genomic modifications, including deletions, substitutions, and additions, and whose presence (or absence) is associated with reduced virulence. In an embodiment, a wildtype M. haemolytica (serotype A1 D153) may be modified to contain a partial gene deletion of the leukotoxin CA (lktCA) genomic locus, resulting in an attenuated bacterium, which secretes a truncated, noncytotoxic form of LktA protein. The vaccines ideally provide safe, effective, and broad protective immunity.
[0010] Another object of the disclosure is to provide multi-valent vaccines, comprising the attenuated M. haemolytica in combination with other bacteria, including P. multocida, M. haemolytica serotype A6, and Histophilus somni (H. somni). Thus, the invention encompasses a 4-way avirulent, modified live vaccine useful against bovine respiratory disease.
[0011] A further object of this invention is to provide methods for treatment and prophylaxis of infection bovine respiratory disease, comprising the steps of administering effective amounts of the inventive vaccines to susceptible bovine animals.
[0012] In one embodiment, the attenuated vaccines further comprises an adjuvant. The adjuvant may be any substance which increases and/or augments the elicited immune response, as compared to attenuated vaccine alone. Mucosal adjuvants, including chitosans and derivatives thereof, are particularly useful for the disclosed oral attenuated vaccines.
[0013] The invention further provides methods for inducing an immunological (or immunogenic) or protective response against M. haemolytica, as well as methods for preventing or treating M. haemolytica, or disease state(s) caused by M. haemolytica, comprising administering the attenuated bacteria, or a composition comprising the attenuated bacteria to animals in need thereof.
[0014] In addition, the disclosure provides PCR methods and reagents useful for diagnosing and/or discriminating between M. haemolytica serotypes A1 and A6. Comparative genomic sequence analysis, further described below, revealed A1- and A6-specific genes, which provide the basis for the methods and reagents provided in this disclosure.
[0015] Kits comprising at least the attenuated M. haemolytica strain and instructions for use are also provided.
[0016] These and other embodiments are disclosed or are obvious from and encompassed by, the following Detailed Description.
BRIEF DESCRIPTION OF THE DRAWINGS
[0017] A full and enabling disclosure of the present invention, including the best mode thereof, to one of ordinary skill in the art, is set forth more particularly in the remainder of the specification, including reference to the accompanying figures, wherein:
[0018] FIG. 1A and FIG. 1B present the scheme used to produce the pCT109GA189.DELTA.lktCA-Kan plasmid (replacement plasmid). The final product for vaccine manufacture incorporated a consensus ribosome-binding site (AGGAGG, rbs) upstream of the start codon which replaced the poor lktC rbs and increased expression of leukotoxoid. The native lktA gene, deleted in the vaccine strain, uses a strong rbs (AGGAGA). For this product, the lktRBSr primer was used in-lieu of the lktCAdelr primer. The consensus site is underlined in FIG. 1A;
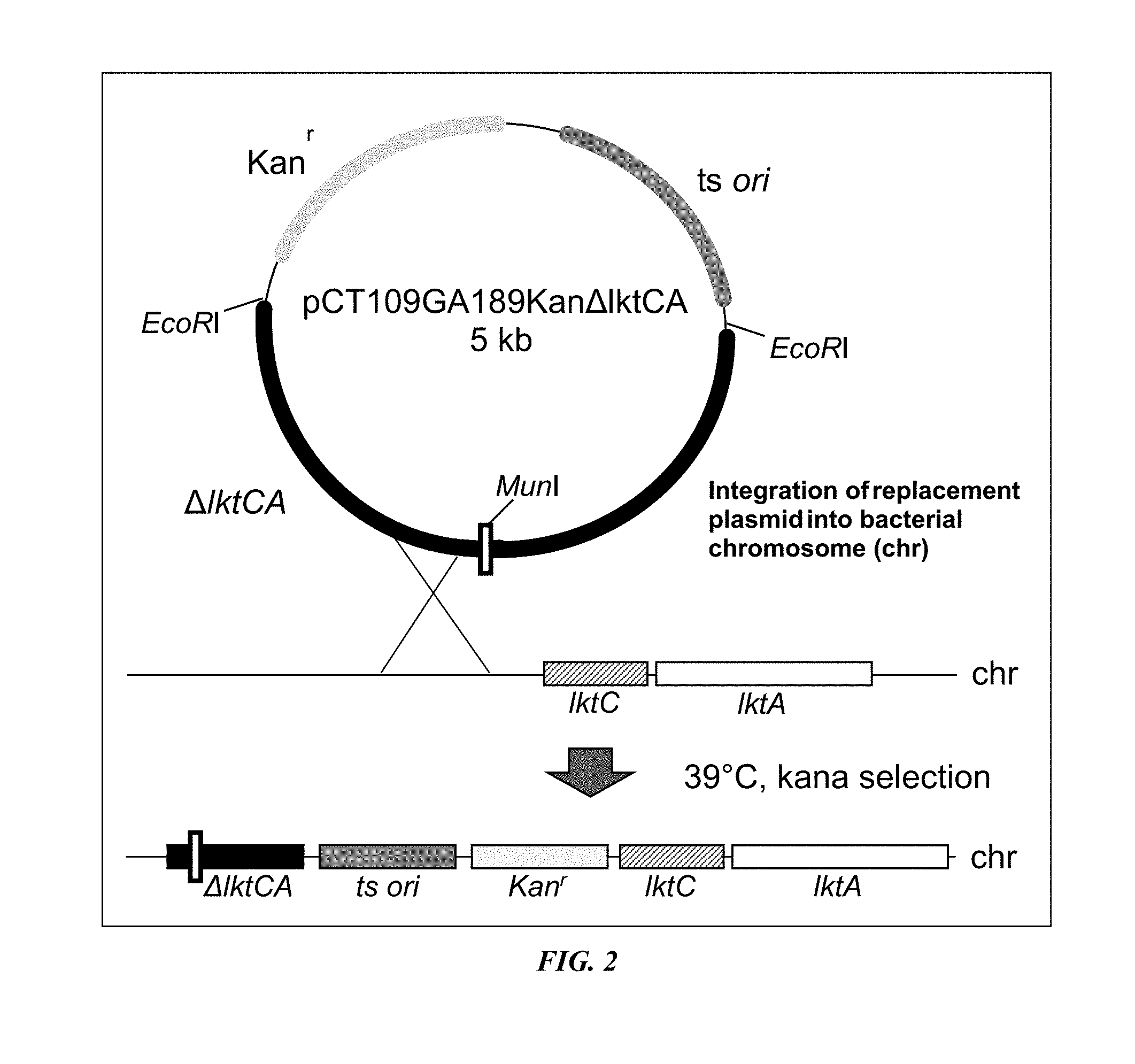
[0019] FIG. 2 illustrates integration of the replacement plasmid into the bacterial genome;
[0020] FIG. 3 depicts resolution/excision of the replacement plasmid, leaving behind only the desired .DELTA.lktCA sequence, stably integrated into the bacterial genome, and encoding the truncated LktA protein;
[0021] FIG. 4A agarose gel electrophoresis of PCR products from M. haemolytica LktCABD operon showing truncated lktCA (lane 2) and wildtype lktCA (lane 3);
[0022] FIG. 4B Western blot analysis of truncated LktA expressed by M haemolytica D153/1-1-PKL, vaccine strain. Lanes: 1-marker; 2-5 .mu.l of culture supernatant containing truncated LktA (*=27 kDa, M. haemolytica D153.DELTA.-1-PKL); 3-5 .mu.l of culture supernatant containing wildtype LktA (*=102 kDa, M. haemolytica D 53 parent strain);
[0023] FIG. 5 is a Venn diagram representing the unique and overlapping genes present in five M. haemolytica isolates.
DETAILED DESCRIPTION OF THE INVENTION
[0024] The present invention relates to a M. haemolytica vaccine or composition which may comprise an attenuated M. haemolytica strain and a pharmaceutically or veterinarily acceptable carrier, excipient, or vehicle, which elicits, induces or stimulates a response in an animal.
[0025] In order to develop an effective M. haemolytica intranasal vaccine, which protects bovines against serogroups A1/A6, inventors used M. haemolytica having a partially deleted lktA gene. This bacterium does not cause cytolysis, but is able to elicit neutralizing antibodies. Prior to the instant disclosure, it was not known whether intranasal administration (or administration via any route) would elicit in bovines a protective immune response.
[0026] Although there are serological methods to distinguish M. haemolytica A1 and A6 these methods are not always reliable and developing strong antisera against A6 is particularly difficult. To overcome this problem, inventors sequenced both A1 and A6 genomes, performed a comparative genomic analysis and developed a real time quantitative polymerase chain reaction (RT-QPCR) method to distinguish between A1 and A6 field isolates and to track our intranasal vaccine combination (M. haemolytica, M. somnus, and P. multocida).
[0027] Thus, an embodiment of this disclosure provides useful RT-QPCR methods, which enable at least the following activities: a) identification of field isolates of M. haemolytica A1 and A6 quickly and screen large number of colonies; b) monitoring of vaccination/colonization of A1 and A6 in nasal cavities; c) elimination of the need for developing high titer antisera; and d) development of rapid, automated diagnostic test kits.
[0028] The present invention further provides attenuated M. haemolytica strains having a deletion in at least one virulence gene. In an embodiment, the deletion is within lktCA, a locus that encodes an enzyme acylase (LktC) and leukotoxin A (LktA), the chief cytotoxin. This deletion may be amplified by polymerase chain reaction (PCR) and the secretion of a truncated LktA can be detected on a Western blot to determine if the bacterium is the mutant or wildtype.
[0029] Deletion of genomic sequence(s) from virulent parental bacteria to produce avirulent, attenuated mutant bacteria is accomplished through novel and non-obvious inventive activity. Such mutant bacteria, also referred to herein as modified-live microorganisms (MLM) are useful for the production of immunogenic compositions or vaccines having both a high degree of immunogenicity and a low (to non-existent) degree of pathogenicity.
[0030] These mutants are also useful as vectors which can be useful for expression in vitro of expression products, as well as for reproduction or replication of nucleotide sequences (e.g., replication of DNA), and for in vivo expression products.
[0031] Engineering of the deletion mutations provides novel and nonobvious nucleotide sequences and genes, as well as novel and nonobvious gene products encoded by the nucleotide sequences and genes. Such gene products provide antigens, immunogens and epitopes, and are useful as isolated gene products. Such isolated gene products, as well as epitopes thereof, are also useful for generating antibodies, which are useful in diagnostic applications.
[0032] Such gene products, which can provide or generate epitopes, antigens or immunogens, are also useful for immunogenic or immunological compositions, as well as vaccines.
[0033] In an aspect, the invention provides bacteria containing an attenuating mutation in a nucleotide sequence or a gene wherein the mutation modifies, reduces or abolishes the expression and/or the biological activity of a polypeptide or protein encoded by a gene, resulting in attenuated virulence of the bacterium. In a particular embodiment, the mutation is an in-frame deletion resulting in the bacterium secreting a truncated leukotoxin. In a particular embodiment, the truncated leukotoxin migrates at about 27 kD on a typical SDS gel.
[0034] Attenuation reduces or abolishes the pathogenicity of the bacteria and the gravity of the clinical signs or lesions, decreases the growth rate of the bacteria, and prevents the death from the bacteria.
[0035] In particular, the present invention encompasses attenuated M. haemolytica strains and vaccines comprising the same, which elicit an immunogenic response in an animal, particularly the attenuated M. haemolytica strains that elicit, induce or stimulate a response in a bovine.
[0036] Particular M. haemolytica attenuated strains of interest have mutations in genes, relative to wild type virulent parent strain, which are associated with virulence. It is recognized that, in addition to strains having the disclosed mutations, attenuated strains having any number of mutations in the disclosed virulence genes can be used in the practice of this invention.
[0037] In another aspect, the novel attenuated M. haemolytica strains are formulated into safe, effective vaccine against M. haemolytica and infections/diseases cause by M. haemolytica.
[0038] In an embodiment, the M. haemolytica vaccines further comprise an adjuvant. In a particular embodiment, the adjuvant is a mucosal adjuvant, such as chitosan, methylated chitosan, trimethylated chitosan, or derivatives or combinations thereof.
[0039] As defined herein, the term "gene" will be used in a broad sense, and shall encompass both coding and non-coding sequences (i.e. upstream and downstream regulatory sequences, promoters, 5'/3' UTR, introns, and exons). Where reference to only a gene's coding sequence is intended, the term "gene's coding sequence" or "CDS" will be used interchangeably throughout this disclosure.
[0040] By "antigen" or "immunogen" means a substance that induces a specific immune response in a host animal. The antigen may comprise a whole organism, killed, attenuated or live; a subunit or portion of an organism; a recombinant vector containing an insert with immunogenic properties; a piece or fragment of DNA capable of inducing an immune response upon presentation to a host animal; a polypeptide, an epitope, a hapten, or any combination thereof. Alternately, the immunogen or antigen may comprise a toxin or antitoxin.
[0041] The terms "protein", "peptide", "polypeptide" and "polypeptide fragment" are used interchangeably herein to refer to polymers of amino acid residues of any length. The polymer can be linear or branched, it may comprise modified amino acids or amino acid analogs, and it may be interrupted by chemical moieties other than amino acids. The terms also encompass an amino acid polymer that has been modified naturally or by intervention; for example disulfide bond formation, glycosylation, lipidation, acetylation, phosphorylation, or any other manipulation or modification, such as conjugation with a labeling or bioactive component.
[0042] The term "immunogenic or antigenic polypeptide" as used herein includes polypeptides that are immunologically active in the sense that once administered to the host, it is able to evoke an immune response of the humoral and/or cellular type directed against the protein. Preferably the protein fragment is such that it has substantially the same immunological activity as the total protein. Thus, a protein fragment according to the invention comprises or consists essentially of or consists of at least one epitope or antigenic determinant. An "immunogenic" protein or polypeptide, as used herein, includes the full-length sequence of the protein, analogs thereof, or immunogenic fragments thereof. By "immunogenic fragment" is meant a fragment of a protein which includes one or more epitopes and thus elicits the immunological response described above. Such fragments can be identified using any number of epitope mapping techniques, well known in the art. See, e.g., Epitope Mapping Protocols in Methods in Molecular Biology, Vol. 66 (Glenn E. Morris, Ed., 1996). For example, linear epitopes may be determined by e.g., concurrently synthesizing large numbers of peptides on solid supports, the peptides corresponding to portions of the protein molecule, and reacting the peptides with antibodies while the peptides are still attached to the supports. Such techniques are known in the art and described in, e.g., U.S. Pat. No. 4,708,871; Geysen et al., 1984. Proc. Natl. Acad. Sci. U.S.A. 81:3998-4002; Geysen et al., 1986. Mol. Immunol. 23:709-715. Similarly, conformational epitopes are readily identified by determining spatial conformation of amino acids such as by, e.g., x-ray crystallography and 2-dimensional nuclear magnetic resonance. See, e.g., Epitope Mapping Protocols, supra. Methods especially applicable to the proteins of T. parva are fully described in PCT/US2004/022605 incorporated herein by reference in its entirety.
[0043] As discussed herein, the invention encompasses active fragments and variants of the antigenic polypeptide. Thus, the term "immunogenic or antigenic polypeptide" further contemplates deletions, additions and substitutions to the sequence, so long as the polypeptide functions to produce an immunological response as defined herein. The term "conservative variation" denotes the replacement of an amino acid residue by another biologically similar residue, or the replacement of a nucleotide in a nucleic acid sequence such that the encoded amino acid residue does not change or is another biologically similar residue. In this regard, particularly preferred substitutions will generally be conservative in nature, i.e., those substitutions that take place within a family of amino acids. For example, amino acids are generally divided into four families: (1) acidic-aspartate and glutamate; (2) basic-lysine, arginine, histidine; (3) non-polar-alanine, valine, leucine, isoleucine, proline, phenylalanine, methionine, tryptophan; and (4) uncharged polar-glycine, asparagine, glutamine, cystine, serine, threonine, tyrosine. Phenylalanine, tryptophan, and tyrosine are sometimes classified as aromatic amino acids. Examples of conservative variations include the substitution of one hydrophobic residue such as isoleucine, valine, leucine or methionine for another hydrophobic residue, or the substitution of one polar residue for another polar residue, such as the substitution of arginine for lysine, glutamic acid for aspartic acid, or glutamine for asparagine, and the like; or a similar conservative replacement of an amino acid with a structurally related amino acid that will not have a major effect on the biological activity. Proteins having substantially the same amino acid sequence as the reference molecule but possessing minor amino acid substitutions that do not substantially affect the immunogenicity of the protein are, therefore, within the definition of the reference polypeptide. All of the polypeptides produced by these modifications are included herein. The term "conservative variation" also includes the use of a substituted amino acid in place of an unsubstituted parent amino acid provided that antibodies raised to the substituted polypeptide also immunoreact with the unsubstituted polypeptide.
[0044] The term "epitope" refers to the site on an antigen or hapten to which specific B cells and/or T cells respond. The term is also used interchangeably with "antigenic determinant" or "antigenic determinant site". Antibodies that recognize the same epitope can be identified in a simple immunoassay showing the ability of one antibody to block the binding of another antibody to a target antigen.
[0045] An "immunological response" to a composition or vaccine is the development in the host of a cellular and/or antibody-mediated immune response to a composition or vaccine of interest. Usually, an "immunological response" includes but is not limited to one or more of the following effects: the production of antibodies, B cells, helper T cells, and/or cytotoxic T cells, directed specifically to an antigen or antigens included in the composition or vaccine of interest. Preferably, the host will display either a therapeutic or protective immunological response such that resistance to new infection will be enhanced and/or the clinical severity of the disease reduced. Such protection will be demonstrated by either a reduction or lack of symptoms and/or clinical disease signs normally displayed by an infected host, a quicker recovery time and/or a lowered viral titer in the infected host.
[0046] By "animal" is intended mammals, birds, and the like. Animal or host as used herein includes mammals and human. The animal may be selected from the group consisting of equine (e.g., horse), canine (e.g., dogs, wolves, foxes, coyotes, jackals), feline (e.g., lions, tigers, domestic cats, wild cats, other big cats, and other felines including cheetahs and lynx), ovine (e.g., sheep), bovine (e.g., cattle), porcine (e.g., pig), avian (e.g., chicken, duck, goose, turkey, quail, pheasant, parrot, finches, hawk, crow, ostrich, emu and cassowary), primate (e.g., prosimian, tarsier, monkey, gibbon, ape), ferrets, seals, and fish. The term "animal" also includes an individual animal in all stages of development, including newborn, embryonic and fetal stages.
[0047] Unless otherwise explained, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this disclosure belongs. The singular terms "a", "an", and "the" include plural referents unless context clearly indicates otherwise. Similarly, the word "or" is intended to include "and" unless the context clearly indicate otherwise.
[0048] It is noted that in this disclosure and particularly in the claims and/or paragraphs, terms such as "comprises", "comprised", "comprising" and the like can have the meaning attributed to it in U.S. patent law; e.g., they can mean "includes", "included", "including", and the like; and that terms such as "consisting essentially of" and "consists essentially of" have the meaning ascribed to them in U.S. patent law, e.g., they allow for elements not explicitly recited, but exclude elements that are found in the prior art or that affect a basic or novel characteristic of the invention.
[0049] The term "nucleic acid" and "polynucleotide" refers to RNA or DNA that is linear or branched, single or double stranded, or a hybrid thereof. The term also encompasses RNA/DNA hybrids. The following are non-limiting examples of polynucleotides: a gene or gene fragment, exons, introns, mRNA, tRNA, rRNA, ribozymes, cDNA, recombinant polynucleotides, branched polynucleotides, plasmids, vectors, isolated DNA of any sequence, isolated RNA of any sequence, nucleic acid probes and primers. A polynucleotide may comprise modified nucleotides, such as methylated nucleotides and nucleotide analogs, uracyl, other sugars and linking groups such as fluororibose and thiolate, and nucleotide branches. The sequence of nucleotides may be further modified after polymerization, such as by conjugation, with a labeling component. Other types of modifications included in this definition are caps, substitution of one or more of the naturally occurring nucleotides with an analog, and introduction of means for attaching the polynucleotide to proteins, metal ions, labeling components, other polynucleotides or solid support. The polynucleotides can be obtained by chemical synthesis or derived from a microorganism.
[0050] The term "gene" is used broadly to refer to any segment of polynucleotide associated with a biological function. Thus, genes include introns and exons as in genomic sequence, or just the coding sequences as in cDNAs and/or the regulatory sequences required for their expression. For example, gene also refers to a nucleic acid fragment that expresses mRNA or functional RNA, or encodes a specific protein, and which includes regulatory sequences.
[0051] An "isolated" biological component (such as a nucleic acid or protein or organelle) refers to a component that has been substantially separated or purified away from other biological components in the cell of the organism in which the component naturally occurs, for instance, other chromosomal and extra-chromosomal DNA and RNA, proteins, and organelles. Nucleic acids and proteins that have been "isolated" include nucleic acids and proteins purified by standard purification methods. The term also embraces nucleic acids and proteins prepared by recombinant technology as well as chemical synthesis.
[0052] The term "conservative variation" denotes the replacement of an amino acid residue by another biologically similar residue, or the replacement of a nucleotide in a nucleic acid sequence such that the encoded amino acid residue does not change or is another biologically similar residue. In this regard, particularly preferred substitutions will generally be conservative in nature, as described above.
[0053] The term "recombinant" means a polynucleotide with semisynthetic, or synthetic origin which either does not occur in nature or is linked to another polynucleotide in an arrangement not found in nature.
[0054] "Heterologous" means derived from a genetically distinct entity from the rest of the entity to which it is being compared. For example, a polynucleotide may be placed by genetic engineering techniques into a plasmid or vector derived from a different source, and is a heterologous polynucleotide. A promoter removed from its native coding sequence and operatively linked to a coding sequence other than the native sequence is a heterologous promoter.
[0055] The polynucleotides of the invention may comprise additional sequences, such as additional encoding sequences within the same transcription unit, controlling elements such as promoters, ribosome binding sites, 5'UTR, 3'UTR, transcription terminators, polyadenylation sites, additional transcription units under control of the same or a different promoter, sequences that permit cloning, expression, homologous recombination, and transformation of a host cell, and any such construct as may be desirable to provide embodiments of this invention.
Methods of Use and Article of Manufacture
[0056] The present invention includes the following method embodiments. In an embodiment, a method of vaccinating an animal comprising administering a composition comprising an attenuated M. haemolytica strain and a pharmaceutical or veterinarily acceptable carrier, excipient, or vehicle to an animal is disclosed. In one aspect of this embodiment, the animal is a bovine.
[0057] The dose volume of compositions for target species that are mammals, e.g., the dose volume of pig or swine compositions, based on bacterial antigens, is generally between about 0.1 to about 2.0 ml, between about 0.1 to about 1.0 ml, and between about 0.5 ml to about 1.0 ml.
[0058] The efficacy of the vaccines may be tested about 2 to 4 weeks after the last immunization by challenging animals, such as bovine, with a virulent strain of M. haemolytica. Both homologous and heterologous strains are used for challenge to test the efficacy of the vaccine. The animal may be challenged by IM or SC injection, spray, intra-nasally, intra-ocularly, intra-tracheally, and/or orally. Samples from joints, lungs, brain, and/or mouth may be collected before and post-challenge and may be analyzed for the presence of M. haemolytica-specific antibody.
[0059] The compositions comprising the attenuated bacterial strains of the invention used in the prime-boost protocols are contained in a pharmaceutically or veterinary acceptable vehicle, diluent or excipient. The protocols of the invention protect the animal from M. haemolytica and/or prevent disease progression in an infected animal.
[0060] The various administrations are preferably carried out 1 to 6 weeks apart. Preferred time interval is 3 to 5 weeks, and optimally 4 weeks according to one embodiment, an annual booster is also envisioned. The animals, for example pigs, may be at least 3-4 weeks of age at the time of the first administration.
[0061] It should be understood by one of skill in the art that the disclosure herein is provided by way of example and the present invention is not limited thereto. From the disclosure herein and the knowledge in the art, the skilled artisan can determine the number of administrations, the administration route, and the doses to be used for each injection protocol, without any undue experimentation.
[0062] Another embodiment of the invention is a kit for performing a method of eliciting or inducing an immunological or protective response against M. haemolytica in an animal comprising an attenuated M. haemolytica immunological composition or vaccine and instructions for performing the method of delivery in an effective amount for eliciting an immune response in the animal.
[0063] Another embodiment of the invention is a kit for performing a method of inducing an immunological or protective response against M. haemolytica in an animal comprising a composition or vaccine comprising an attenuated M. haemolytica strain of the invention, and instructions for performing the method of delivery in an effective amount for eliciting an immune response in the animal.
[0064] Yet another aspect of the present invention relates to a kit for prime-boost vaccination according to the present invention as described above. The kit may comprise at least two vials: a first vial containing a vaccine or composition for the prime-vaccination according to the present invention, and a second vial containing a vaccine or composition for the boost-vaccination according to the present invention. The kit may advantageously contain additional first or second vials for additional prime-vaccinations or additional boost-vaccinations.
[0065] The pharmaceutically or veterinarily acceptable carriers or vehicles or excipients are well known to the one skilled in the art. For example, a pharmaceutically or veterinarily acceptable carrier or vehicle or excipient can be a 0.9% NaCl (e.g., saline) solution or a phosphate buffer. Other pharmaceutically or veterinarily acceptable carrier or vehicle or excipients that can be used for methods of this invention include, but are not limited to, poly-(L-glutamate) or polyvinylpyrrolidone. The pharmaceutically or veterinarily acceptable carrier or vehicle or excipients may be any compound or combination of compounds facilitating the administration of the vector (or protein expressed from an inventive vector in vitro); advantageously, the carrier, vehicle or excipient may facilitate transfection and/or improve preservation of the vector (or protein). Doses and dose volumes are herein discussed in the general description and can also be determined by the skilled artisan from this disclosure read in conjunction with the knowledge in the art, without any undue experimentation.
[0066] The immunological compositions and vaccines according to the invention may comprise or consist essentially of one or more adjuvants. Suitable adjuvants for use in the practice of the present invention are (1) polymers of acrylic or methacrylic acid, maleic anhydride and alkenyl derivative polymers. (2) immunostimulating sequences (ISS), such as oligodeoxyribonucleotide sequences having one or more non-methylated CpG units (Klinman et al., 1996; WO98/16247), (3) an oil in water emulsion, such as the SPT emulsion described on page 147 of "Vaccine Design, The Subunit and Adjuvant Approach" published by M. Powell, M. Newman, Plenum Press 1995, and the emulsion MF59 described on page 183 of the same work, (4) cationic lipids containing a quaternary ammonium salt, e.g., DDA (5) cytokines, (6) aluminum hydroxide or aluminum phosphate, (7) saponin or (8) other adjuvants discussed in any document cited and incorporated by reference into the instant application, or (9) any combinations or mixtures thereof.
[0067] In an embodiment, adjuvants include those which promote improved absorption through mucosal linings. Some examples include MPL, LTK63, toxins, PLG microparticles and several others (Vajdy, M. Immunology and Cell Biology (2004) 82, 617-627). In an embodiment, the adjuvant may be a chitosan (Van der Lubben et al. 2001. Adv. Drug Delivery Reviews 52:139-144; Patel et al. 2005. AAPS PharmSciTech 6(1):E49-E55; Majithiya et al. 2008. Drug Deliv. Technol. 8:40-45; U.S. Pat. No. 5,980,912).
[0068] In an embodiment, the adjuvant may be inactivated bacteria, an inactivated virus, fractions of inactivated bacteria, bacterial lipopolysaccharides, bacterial toxins, or derivatives or References. [0069] Ackermann, M. R, Brogden, K. A. 2000. Response of the ruminant respiratory tract to Mannheimia (Pasteurella) haemolytica. Microbes Infect. 2:1079-1088. [0070] Al-Ghamdi, G. M., et al, 2000. Serotyping of Mannheimia (Pasteurella) haemolytica isolates from the upper Midwest United States. J. Vet. Diagn. Invest. 12, 576-578. [0071] Bowland, S., Shewen, P., 2000. Bovine respiratory disease: commercial vaccines currently available in Canda. Can. Vet. J. 41, 33-38. [0072] Briggs R. E, Tatum F. M. 2005. Generation and molecular characterization of new temperature--sensitive plasmids intended for genetic engineering of Pasteurellaceae. Appl Environ Micobiol 71:7187-7195. [0073] Burriel, A. R. 1997. News & Notes: Isolation of Pasteurella haemolytica from Grass, Drinking Water, and Straw Bedding Used by Sheep. Curr. Microbiol. 35: 316-318 [0074] Confer, A. W., et al., 2003. Immunogenicity of recombinant Mannheimia haemolytica serotype 1 outer membrane protein PlpE and augmentation of a commercial vaccine. Vaccine 21, 2821-2829. [0075] Davies, R. L, et al. 2002. Mosaic structure and molecular evolution of the leukotoxin operon (lktCABD) in Mannheimia (Pasteurella) haemolytica, Mannheimia glucosida, and Pasteurella trehalosi. J Bacteriol. 184(1):266-277. [0076] Davies, R. L, et al. 2001. Sequence diversity and molecular evolution of the leukotoxin (lktA) gene in bovine and ovine strains of Mannheimia (Pasteurella) haemolytica. J Bacteriol. 183(4):1394-1404. [0077] Ellis, J., et al., 2007. Response of calves to challenge exposure with virulent bovine respiratory syncytial virus following intranasal administration of vaccines formulated for parenteral administration. J. Am. Vet. Med. Assoc. 230, 233-243. [0078] Frank, G. H, et al. 2003. Effect of intranasal exposure to leukotoxin-deficient Mannheimia haemolytica at the time of arrival at the feedyard on subsequent isolation of M. haemolytica from nasal secretions of calves. Am J Vet Res. 64(5):580-585. [0079] Gioia. J. et al. 2006. The genome sequence of Mannheimia haemolytica A1: insights into virulence, natural competence, and Pasteurellaceae phylogeny. J Bacteriol. 188(20):7257-7266. [0080] Lawrence, P. K., et al., 2010. Three-way comparative genomic analysis of two Mannheimia haemolytica isolates. BMC Genomics. 11:535 (Open access). [0081] Morton. R. J., et al., 1995. Vaccination of cattle with outer membrane protein-enriched fractions of Pasteurella haemolytica and resistance against experimental challenge exposure. Am. J. Vet. Res. 56, 875-879. [0082] Miller, M. W. 2001. Pasteurellosis, In E. S. Williams and I. K. Barker (ed.), Infectious diseases of wild mammals. Iowa State University. Press, Ames, Iowa p. 330-339 [0083] Mosier, D. A. 1997. Bacterial pneumonia. Vet. Clin. N. Am. Food Anim. Pract. 13:483-493. [0084] Muylkens, B., et al., 2007. Bovine herpesvirus I infection and infectious bovine rhinotracheitis. Vet. Res. 38, 181-209. [0085] Potter, A. A., et al., 1999. Protective capacity of the Pasteurella haemolytica transferrin-binding proteins TbpA and TbpB in cattle. Microb Pathog 27, 197-206. [0086] Perino, L. J., Hunsaker, B. D., 1997. A review of bovine respiratory disease vaccine field efficacy. [0087] The Bovine Practitioner 31, 59-66. [0088] Purdy, C. W., et al, 1993. Efficacy of Pasteurella haemolytica subunit antigens in a goat model of pasteurellosis. Am. J. Vet. Res. 54, 1637-1647. [0089] Purdy, C. W., et al., 1997. Efficacy of a subcutaneously administered, ultraviolet light-killed Pasteurella multocida A:3-containing bacterin against transthoracic challenge exposure in goats. Am. J. Vet. Res. 58, 841-847. [0090] Rehmtulla, A. J, Thomson, R. G. 1981. A review of the lesions in shipping fever of cattle. Can. Vet. J. 22:1 [0091] Rice, J. A., et al., 2007. Mannheimia haemolytica and bovine respiratory disease. Anim. Health Res. Rev. 8, 117-128. [0092] Sabri, M. Y., et al., 2000. Efficacy of an outer membrane protein of Pasteurella haemolytica A2, A7 or A9-enriched vaccine against intratracheal challenge exposure in sheep. Vet. Microbiol. 73, 13-23. [0093] Schreuer, D., et al. 2000. Evaluation of the efficacy of a new combined (Pasteurella) Mannheimia haemolytica serotype A1 and A6 vaccine in pre-ruminant calves by virulent challenge. Journal Cattle Practice Vol. 8 No. 1 pp. 9-12 [0094] Shewen, P. E., Wilkie, B. N., 1988. Vaccination of calves with leukotoxic culture supernatant from Pasteurella haemolytica. Can. J. Vet. Res. 52, 30-36. [0095] Virtala, A. M., et al., 1996. Epidemiologic and pathologic characteristics of respiratory tract disease in dairy heifers during the first three months of life. J. Am. Vet. Med. Assoc. 208, 2035-2042.
[0096] The invention will now be further described by way of the following non-limiting examples.
EXAMPLES
Example 1--Production of Attenuated M. haemolytica
[0097] M. haemolytica is a commensal organism of the upper respiratory tract of calves and other ruminants. Under stress and in immunocompromised animals M. haemolytica descends into lungs and causes severe systemic disease resulting in pneumonic pasteurellosis or "shipping fever". The pathogen can be spread by nose to nose contact. To attenuate the bacterium, we deleted nucleotides within the LktCA locus, which encodes an enzyme acylase (LktC) and leukotoxin A (LktA), the bacterium's chief cytotoxin. This deletion can be amplified by polymerase chain reaction (PCR) and the secretion of a truncated LktA can be detected on a Western blot to determine if the bacterium is the mutant or wildtype. The genetic engineering is summarized in FIGS. 1A-3. All reagents, including the shuttle vectors pCR2.1, pBC SK, pSK, and pCT109GA189 ts ori, and the E. coli DH11S host cell, are well-known to and accessible by persons skilled in the art.
[0098] Construction of lktCA Deletion.
[0099] pCT109GA 189-Kan.DELTA.lktCA and pCT109GA 189-Kan.DELTA.lktCArbs were constructed as outlined in FIGS. 1A-3. Briefly, two DNA fragments, upstream (1.06 kb, SEQ ID NO:6) and downstream (1.29 kb, SEQ ID NO:7) were PCR amplified from M. haemolytica strain NADC D153 (FIG. 1A). Whole cells were used as template using the primer sets, lktCAf (SEQ ID NO: 1)/lktCAdelr (SEQ ID NO:4) and lktCAr (SEQ ID NO:2)/lktCAdelf (SEQ ID NO:3). The PCR products were phenol-chloroform-extracted to inactivate Taq polymerase and then digested with MunI prior to ligation. The ligation products were PCR amplified with primer pair lktCAf/lktCAr and the products were cloned using a commercially available vector (PCR2.1, Invitrogen, Carlsbad, Calif.) according to manufacturer instructions.
[0100] A product containing an approximately 2.3 kb insert was selected and proper sequence across the deletion was confirmed by DNA sequencing and designated pTA.DELTA.lktCA. A kanamycin cassette derived from pUC4K was placed into the Sal1 site of pBC SK--(Stratagene Inc.) to generate pBCKan. The 2.3 kb deleted leukotoxin insert in pTA.DELTA.lktCA was transferred into pBCKan by digestion with EcoRI and ligation into the unique EcoRI site to form pBCKan.DELTA.lktCA.
[0101] This product was amplified by PCR using primer pair lktCAdelf (SEQ ID NO:3) and lktRBSr (SEQ ID NO:5) to replace the native lktC ribosome binding site (RBS) with a consensus RBS (FIG. 1A). The product was digested with MunI and ligated onto itself to form pBCKan.DELTA.lktCArbs. Proper sequence adjacent to the deletion was confirmed by DNA sequencing. Finally the pBC plasmid backbone of both pBCKan.DELTA.lktCA and pBCKan.DELTA.lktCArbs was replaced with the temperature-sensitive plasmid origin of replication from pCT109GA189 (Briggs and Tatum, 2005) by ligating BssHII-digested preparations of each to generate pCT109GA 189Kan.DELTA.lktCA and pCT109GA189Kan.DELTA.lktCArbs.
[0102] Electrocompetent M. haemolytica serotype AI D153 cells (virulent parental strain) were transformed with pCT109GA189Kan lktCA and pCT109GA189Kan.DELTA.lktCArbs by previously described methods except unmethylated ligation product was directly introduced into the competent cells. (Briggs and Tatum, 2005) Briefly, cells were made electrocompetent by growing them to logarithmic phase in 100 ml of Columbia broth (Difco Laboratories, Detroit, Mich.) at 37.degree. C. with gentle shaking. The cells were pelleted by centrifugation at 5,000 .mu.g and washed in 100 ml of 272 mM sucrose at 0.degree. C., and the pellet was suspended in an equal volume of 272 mM sucrose at 0.degree. C. After electroporation, cells recovered overnight in 10 ml Columbia broth at 30.degree. C. Growth (50 .mu.l) was spread onto Columbia agar plates containing 50 .mu.g/ml kanamycin, which were then incubated 36 hours at 30.degree. C. Individual colonies were passed to broth containing 50 .mu.g/ml kanamycin and incubated overnight at 30.degree. C. Growth (100 .mu.l) was passed again to Columbia agar plates with kanamycin which were incubated overnight at 39.degree. C. Individual colonies were passed to trypticase soy agar (TSA) plates containing 5% defibrinated sheep blood (BA plates, incubated overnight at 39.degree. C.) and to Columbia broth without selection (incubated overnight at 30.degree. C.). Growth in broth was streaked for isolation on BA plates and passed again in broth at 30.degree. C. Non-hemolytic colonies which were kanamycin-sensitive were detected on BA plates after 1 to 3 passages without selection. Representative colonies from each recipient strain and replacement plasmid were selected for further study.
[0103] Because the temperature-sensitive plasmid origin functions poorly in E. coli cloning hosts, these final ligation products were introduced directly into M. haemolytica. Prior cloning steps used E. coli DH11S (Life Technologies, Rockville, Md.) as the cloning host.
[0104] Non-hemolytic mutants were grown in Columbia broth at 37.degree. C. for 3 hours and harvested in late logarithmic growth. Supernatants were dotted onto nitrocellulose along with supernatants from the wild-type parent and a leukotoxin-negative isogenic mutant. After appropriate blocking and washing, the blot was probed with monoclonal anti-leukotoxin antibody 2C9-1E8 (neutralizing antibody produced by NADC, Ames, Iowa). Mutant products containing the native ribosome binding site were found to express low levels of protein reactive to monoclonal antibody, less than that produced by the wild-type parent strain. Products which contained the new ribosome binding site produced much higher levels of reactive protein. Supernatants of two products expressing high levels of leukotoxin were concentrated 15-fold on a 10,000 MW filter (Centriprep, Amicon). The concentrates (1.5 .mu.l) were subjected to SDS-PAGE, blotted to nitrocellulose, and probed with antibody 2C9-IE8. Western blot analysis indicated a new protein reactive with neutralizing anti-leukotoxin monoclonal antibody at an apparent molecular weight consistent with the 27 kDa predicted protein (truncated LktA) product. These representative mutants and single-crossover controls were analyzed by PCR to demonstrate the absence of temperature-sensitive origin and kanamycin-resistance cassette (Step G). The mutant M. haemolytica serotype A1 was designated as DI53.DELTA.lktCA4-707, which refers to the amino acid positions in LktC and LktA respectively where the deleted region begins and ends. Gene insertion was characterized by PCR amplification using LktCAf (SEQ ID NO:1) and LktCAr (SEQ ID NO:2) primers, which flank the deletion site. As indicated by the gel image, PCR amplification yielded the expected 2.3 kb for truncated LktCA, and 5.0 kb for the wildtype bacterium (FIG. 4A). Finally, PCR performed with primers (SEQ ID NOs: 1 & 2) flanking ts ori and kanamycin resistance genes confirmed those elements were no longer present in the final LktCA mutant for Master Seed (MS). Five microliters of the concentrated culture supernatant was run on a SDS-PAGE system, blotted onto PVDF membrane and probed using mouse anti-LktA, neutralizing antibody 2C9-IE8 (1:1000) as primary antibody. Goat anti-mouse IgG (1:4000) coupled with alkaline phosphatase was used as secondary antibody and developed in a substrate solution containing NBT/BCIP for I-5 min (FIG. 4B). The lack of functional acylase prevents the activation of LktA, and furthermore, the N-terminal deletion of LktA prevents it from forming pores on host animal neutrophils or macrophages.
Example 2--Efficacy of Attenuated M. haemolytica in Calves
[0105] Calves were randomly assigned to one of three groups, each receiving either 10.sup.6 or 10.sup.7 CFU of the MH A1+A6 vaccine, or the control RPMI (diluent). Lyophilized Mannheimia haemolytica (MH) serotypes A1 and A6 were resuspended and administered intranasally, I mL to each nostril, of nine calves, aged 5-6 weeks. The calves were observed for feed intake and rectal temperatures taken morning and evening for 3 days post vaccination. Nasal colonization of M. haemolytica A 1 and A6 following vaccination was analyzed by RT-QPCR (differentiated among M. haemolytica A1 and A6 throughout the study). Vaccines were plated on TSA for exact CFU/ml count on each vaccine the following day.
[0106] Challenge.
[0107] A fresh glycerol stock of virulent MH A1 was grown O/N in BHI medium, plated (TSA) the next day and incubated at 37.degree. C. The following day, plates were scraped and diluted into RPMI medium supplemented with 2% inactivated fetal bovine serum. The inoculum was grown at 37.degree. C./200 rpm until desired OD.sub.600 was achieved, and the culture was diluted to the desired CFU/challenge dose and dilution plated to enumerate the exact CFU/ml the following day. The remaining inoculum was immediately dilution plated in the lab. Calves were challenged on DAY via trans-tracheal administration of 2.4.times.10.sup.9 CFU in 20 ml RPMI, chased with 60 ml RPMI. The calves were monitored for change in behavior including lethargy, coughing, nasal discharge and scored as shown in Table 3. Rectal temperatures were monitored for calves showing clinical signs. The lungs were scored for pneumonic lesions and recorded as % lesion on each lobe, and lung tissue was also collected for histopathology. Swabs were taken from lung lesions and trachea to recover the challenge organism. Table 1 presents the study schedule.
TABLE-US-00001 TABLE 1 Study schedule Age Day Event 5-6 0 Day 0-Bleed, Swab and vaccinate intra-nasally weeks 7 7 days post vax-Bleed and swab old 14 14 days post vax-Bleed and swab 21 21 days post vax-Bleed and swab 22 22 days post vax-Bleed, swab & Challenge with M. haemolytica A1 22-29 Observe clinical signs starting 8/7, euthanize any calves if necessary. Euthanize and necropsy all on 8/13 *Calves were observed for feed intake and rectal temperatures (morning/evening) for 3 days, post vaccination.
Samples from each calf were tested using whole cell, Lkt ELISA and RT-QPCR.
TABLE-US-00002 TABLE 2 Clinical signs criteria 0 = Normal 1 = Depression, Anorexia, Cough, Nasal Discharge, Dyspnea 2 = Severely Depressed, Unable to Rise or Walk, Euthanized for Humane Reasons 3 = Dead On Arrival (DOA)
[0108] Results.
[0109] Three days post challenge one of the control calves showed severe signs of pneumonia and was euthanized (36.92% typical M. haemolytica lesions). The remaining 8 calves were euthanized on day 6 and their percent lung involvement is described in Table 3. The results clearly indicate that the vaccine affords protection when administered intranasally. As indicated in Table 4 intranasal vaccination of M. haemolytica A1/A6 combo significantly reduced (62.0% and 76.7% for 6 log and 7 log group respectively) the lung lesions when compared to sham.
[0110] Furthermore, histopathological analysis clearly indicated typical necrotizing bronchopneumonia characteristic of M. haemolytica.
TABLE-US-00003 TABLE 4 Average % reduction in lung Average lesion Actual vaccine dose Lung lung compared Animal # A1/A6 CFU/animal lesion (%) lesion (%) to sham 125 1.19 .times. 10.sup.6/9.2 .times. 10.sup.5 24.03* 176 1.19 .times. 10.sup.6/9.2 .times. 10.sup.5 0.0 188 1.19 .times. 10.sup.6/9.2 .times. 10.sup.5 6.40 10.43 62.0 179 1.19 .times. 10.sup.7/9.2 .times. 10.sup.6 0.87 185 1.19 .times. 10.sup.7/9.2 .times. 10.sup.6 1.837 189 1.19 .times. 10.sup.7/9.2 .times. 10.sup.6 14.91* 6.48 76.7 122 Sham 8.85 177 Sham 37.75 182 Sham 36.92 27.84 *The lesions (gross pathology) were due to typical Mycoplasma bovis chronic infection
Example 3--Development of RT-QPCR Method for Distinguishing Between A1/A6 Serotypes
[0111] The efficacy of intranasal colonization of M. haemolytica A1/A6 was followed during the course of experiment by a novel QPCR method. Briefly, the genomes of above-described A1 and A6 serotype bacteria were compared against one A1 and two A2 genomes available in GenBank. The comparison revealed 63 genes specific for A1 (D153) and 42 genes specific for A6 (D174). Out of these 105 genes we picked a S6 family IgA-specific metalloendopeptidase (SEQ ID NO: 14) specific for A1 and BCCT family betaine/carnitine/choline transporter gene (SEQ ID NO:12) specific for A6 respectively for differential real time PCR. These gene sequences were amplified by using gene specific primers, sequenced by standard Sanger method and verified. Next, we designed real time PCR primers and tagged the probes with two different dyes (A1-5'6 FAM/ZEN/3 and A6-5'Cy5/3'IBRQ) within each gene. To verify the efficacy our assay method we picked M. haemolytica colonies from nasal swabs obtained from calves maintained in our facilities 7 days post vaccination. The individual colonies were amplified by multiplex real time colony PCR using QuantiTect Probe PCR kit mastermix (Qiagen) following the manufacturer's instruction in a MX3000P qPCR machine (Stratagene). A1 and A6 colonies verified by serotyping were used as positive controls for multiplex real time quantitative PCR (RT-QPCR). The ct values were set at machine default setting and each colony verified by multiplex real time PCR was confirmed by leukotoxin (lktA) specific PCR. The RT-QPCR results 7 days post vaccination indicated a preferential colonization of A1 over A6 (Table 5), which was further confirmed by leukotoxin gene specific deletion PCR (Table 6). But 14 and 21 days post vaccination indicated essentially exclusive colonization of A1 (Tables 7 & 8).
TABLE-US-00004 TABLE 5 RT-QPCR results for nasal swabs from D7 Post Vaccination ID A1 A6 .DELTA. lkt 151-1 17 11 + 151-2 15 -- + 151-3 16 -- + 151-4 17 -- + 151-5 15 -- + 154-1 -- -- 154-2 -- 39 154-3 -- -- 154-4 -- -- 154-5 -- 22 157-1 15 -- + 157-2 22 -- + 157-3 17 -- + 157-4 15 33 + 157-5 16 -- + 160-1 18 13 + 160-2 -- 12 + 160-3 -- 12 + 160-4 -- 12 + 160-5 -- 11 + 178-1 -- -- 178-2 -- -- 178-3 -- -- 178-4 -- 24 178-5 -- 31 181-1 15 15 + 181-2 17 -- + 181-3 -- 13 + 181-4 17 -- + 181-5 15 -- + 183-1 16 12 + 183-2 -- 35 183-3 17 -- + 183-4 16 -- + 183-5 -- 17 + 186-1 -- 42 186-2 -- 43 186-3 -- -- 186-4 -- -- 186-5 -- 20 190-1 -- -- 190-2 -- -- 190-3 -- 10 190-4 -- -- 190-5 -- -- 193-1 15 38 + 193-2 15 -- + 193-3 -- 36 193-4 16 20 + 193-5 -- -- A1 mut. Vx 15 -- + A6 mut. Vx -- 11 + Neg -- --
TABLE-US-00005 TABLE 6 PCR results for nasal swabs from D7 Post Vaccination ID/colony A1 A6 .DELTA.lkt ~2300 bp 122-1 -- -- 122-2 -- -- 122-3 -- -- 122-4 -- -- 122-5 -- -- 125-1 16 -- + Y 125-2 17 -- + Y 125-3 17 -- + Y 125-4 16 -- + Y 125-5 17 -- + Y 176-1 17 -- + Y 176-2 17 -- + Y 176-3 16 -- + Y 176-4 16 -- + Y 176-5 16 -- + Y 177-1 -- -- 177-2 -- -- 177-3 -- -- 177-4 -- -- 177-5 -- -- 179-1 17 -- + Y 179-2 16 -- + Y 179-3 -- -- 179-4 16 -- + Y 179-5 29 -- + Y 182-1 -- -- 182-2 -- -- 182-3 -- -- 182-4 -- -- 182-5 -- -- 185-1 -- 15 + Y 185-2 18 -- + Y 185-3 16 -- + Y 185-4 -- -- + Y 185-5 22 -- + Y 188-1 -- -- 188-2 -- -- 188-3 -- -- 188-4 -- -- 188-5 -- -- 189-1 16 -- + Y 189-2 16 -- + Y 189-3 21 -- + Y 189-4 16 -- + Y 189-5 17 -- + Neg -- --
TABLE-US-00006 TABLE 7 PCR results for nasal swabs from D14 Post Vaccination ID-colony # A1 A6 .DELTA.lkt PCR Lkt .DELTA. 122-1 (Con. 0 0 Neg 122-2 (Con. 0 0 Neg 122-3 (Con. 0 0 Neg 125-1 (6 log) 15 0 Pos Y 125-2 (6 log) 16 0 Pos Y 125-3 (6 log) 16 0 Pos Y 176-1 (6 log) 0 0 Neg 176-2 (6 log) 0 0 Neg 176-3 (6 log) 0 0 Neg 177-1 (Con. 0 0 Neg 177-2 (Con. 0 0 Neg 177-3 (Con. 0 0 Neg 179-1 (7 log) 0 0 Neg 179-2 (7 log) 0 0 Neg 179-3 (7 log) 0 0 Neg 182-1 (Con.) 0 0 Neg 182-2 (Con.) 0 0 Neg 182-3 (Con.) 0 0 Neg 185-1 (7 log) 0 0 Neg 185-2 (7 log) 0 0 Neg 185-3 (7 log) 0 0 Neg 188-1 (6 log) 0 0 Neg 188-2 (6 log) 0 0 Neg 188-3 (log) 0 0 Neg 189-1 (7 log) 15 0 Pos Y 189-2 (7 log) 15 0 Pos Y 189-3 (7 log) 15 0 Pos Y A1 Mutant Pos 15 0 Pos Y A6 Mutant Pos 0 0 Pos Y Neg Con. 0 0 Neg
TABLE-US-00007 TABLE 8 PCR results for nasal swabs from D21 Post Vaccination ID-colony # A1 A6 .DELTA.lkt PCR .DELTA.lkt 122-1 (Con.) 0 0 122-2 (Con.) 0 0 122-3 (Con.) 0 0 125-1 (6 log 14 0 + Y 125-2 (6 log 15 0 + Y 125-3 (6 log 15 0 + Y 176-1 (6 log 15 0 + Y 176-2 (6 log 15 0 + Y 176-3 (6 log 15 0 + Y 177-1 (Con.) 0 0 177-2 (Con.) 0 0 177-3 (Con.) 0 0 179-1 (7 log 0 0 179-2 (7 log 0 0 179-3 (7 log 0 0 181-1 (Con.) 0 0 182-2 (Con.) 0 0 182-3 (Con.) 0 0 185-1 (7 log) 15 0 + Y 185-2 (7 log) 14 0 + Y 185-3 (7 log) 15 0 + Y 188-1 (6 log) 14 0 + Y 188-2 (6 log) 15 0 + Y 188-3 (6 log) 14 0 + Y 189-1 (7 log) 16 0 + Y 189-2 (7 log) 17 0 + Y 189-3 (7 log) 15 0 + Y A1 Mutant Pos 15 0 + Y A6 Mutant Pos 0 16 + Y Neg Control 0 0 neg Pre Challenge A1 Wt 15 0 + WT Pre Challenge A1 Wt 16 0 + WT
Example 4--Intranasal Vaccination of Calves Using Mannheimia haemolytica A1 & A6 Vaccines Followed by Virulent Challenge
[0112] Fifteen calves, 4 weeks of age and housed in 3 different pens/5 calves per pen, were randomly assigned to one of the two treatment group. Calves were vaccinated intranasally with modified live Mannheimia haemolytica serotypes A1 and A6 (reconstituted from lyophilized, Table 9), and intranasal colonization of A1 and A6 was monitored by real time PCR. Calves were finally challenged with virulent M. haemolytica A6 (wild type) to determine vaccine efficacy.
TABLE-US-00008 TABLE 9 Treatment Groups. Total Dose/CFU Group Treatment per animal Route/volume Calf Id # 1 M. haemolytica 10.sup.7 (1.43 .times. 10.sup.6 + Intranasal 1 ml 2, 4, 6 A1 + A6 8.63 .times. 10.sup.5)* per nostril 8, 10 2 M. haemolytica 10.sup.8 (1.43 .times. 10.sup.7 + Intranasal 1 ml 1, 3, 5, A1 + A6 8.63 .times. 10.sup.6)* per nostril 7, 9 3 Control- control Intranasal 1 ml 162, 166, Lyophilized per nostril 170, 174, RPMI + 175 stabilizer *Actual CFU/ml based on plate count
[0113] Vaccination.
[0114] Lyophilized cultures of M. haemolytica A1 and A6 were enumerated from a batch stored at 4.degree. C. On vaccination day, the vaccines were diluted in RPMI (colorless) to required CFU/ml for each isolate. Similarly, the sham vaccine (lyophilized RPMI in stabilizer) was diluted in RPMI. The vaccines were plated on TSA to determine the exact CFU/ml count on each vaccine the following day. The vaccines were mixed and administered 1 mL/nostril using a repeat syringe attached with a cannula according to the dose in Table 9. The control group was vaccinated first, followed by the lowest to highest log group. Following vaccination, the samples were collected as described in Table 10, and the calves were observed for feed intake and rectal temperatures taken morning and evening for 3 days post vaccination. Nasal colonization of M. haemolytica A1 and A6 following vaccination was analyzed by Q-PCR as described above.
[0115] M. haemolytica A6 challenge culture. A fresh glycerol stock of M. haemolytica A6 was grown O/N in BHI medium, plated (TSA) the next day and incubated at 37.degree. C. The following day, plates were scraped and diluted into RPMI medium supplemented with 2% inactivated fetal bovine serum. The inoculum was grown at 37.degree. C./200 rpm until desired OD.sub.600 was achieved. The culture was diluted to desired CFU/challenge dose and dilution plated to enumerate the exact CFU/ml the following day. The inoculum was transported on ice and kept on ice during challenge, and administered trans-tracheally using a 14G.times.1 inch needle. The dose was 1.09.times.10.sup.9 CFU/animal in 20 ml RPMI, chased with 60 ml RPMI. Once completed, the remaining inoculum was immediately dilution plated. The calves were monitored for behavior changes including lethargy, coughing, and nasal discharge and scored as shown in Table 11. Rectal temperatures were monitored for calves showing clinical signs. The lungs were scored for pneumonic lesions and recorded as % lesion on each lobe, and tissues were collected for histopathology. Swabs were also taken from lungs (lesions) and trachea to recover the challenge organism.
TABLE-US-00009 TABLE 10 Study Schedule. Age Date Event 4 weeks old 0 Day 0-Bleed, Swab and vaccinate intra-nasally 7 days post vax 7 days post vax-Bleed and swab 15 days post vax 15 days post vax-Bleed and swab & Challenge with M. haemolytica A6 15 to 20 days post Observe clinical signs starting day 15; vax euthanized any calves when necessary. Euthanized and necropsy all on day 20 * Feed intake (daily) and rectal temperatures (twice daily) were monitored for 3 days post vaccination.
TABLE-US-00010 TABLE 11 Clinical signs. Criteria for Post Challenge Observations 0 = Normal 1 = Depression, Anorexia, Cough, Nasal Discharge, Dyspnea 2 = Severely Depressed, Unable to Rise or Walk, Euthanized for Humane Reasons 3 = Dead On Arrival (DOA)
Results.
[0116] Two days post challenge calf #5 and 174 showed severe signs of pneumonia and were euthanized. Calf #7 died on day 3, post challenge. The remaining 12 calves were euthanized on day 5 and their % lung involvement is described in Table 4. The results indicate that 80% of vaccinates were protected by the modified live M. haemolytica A1/A6 vaccine. From the 7 log group, three (1, 3 and 9) animals were protected while the other two animals (5, 7) had significantly large lesions compared to controls. The large lesions could have been caused by an existing Mannheimia, mycoplasma or viral infection, which had been exacerbated by challenge. Overall, 80% of vaccinates (1, 2, 3, 4, 6, 8, 9 and 10) had significantly (89.55% reduction) reduced lung lesion as compared to control, and histopathological analysis indicated typical necrotizing bronchopneumonia in the control animals.
TABLE-US-00011 TABLE 12 Dosage groups. Average % reduction in lung lesion Average compared Actual A1/A6 Lung lung to Animal vaccine dose lesion lesion sham Group # CFU/animal (%) (%) vaccine 10.sup.7 2 1.43 .times. 10.sup.6/8.63 .times. 10.sup.5 0.0 4 1.43 .times. 10.sup.6/8.63 .times. 10.sup.5 8.67 6 1.43 .times. 10.sup.6/8.63 .times. 10.sup.5 5.92 8 1.43 .times. 10.sup.6/8.63 .times. 10.sup.5 4.83 10 1.43 .times. 10.sup.6/8.63 .times. 10.sup.5 0.0 3.88 85.04 10.sup.8 1 1.43 .times. 10.sup.7/8.63 .times. 10.sup.6 0.0 3 1.43 .times. 10.sup.7/8.63 .times. 10.sup.6 0.0 5 1.43 .times. 10.sup.7/8.63 .times. 10.sup.6 41.58 7 1.43 .times. 10.sup.7/8.63 .times. 10.sup.6 64.47 9 1.43 .times. 10.sup.7/8.63 .times. 10.sup.6 2.295 21.66 14.47 162 Sham 37.11 166 Sham 29.82 170 Sham 11.235 174 Sham 25.54 175 Sham 25.97 25.93
[0117] The efficacy of intranasal colonization of M. haemolytica A1 A6 was followed during the course of experiment by above-described QPCR methods. Results for 7 and 15 days post-vaccination indicated vaccinates had a preferential colonization of A1 over A6 which was further confirmed by leukotoxin gene specific deletion PCR (Tables 13 & 14).
TABLE-US-00012 TABLE 13 Day 7 Post Vaccination Sample # Animal # FAM MHA1 MHA1? CY5 MHA6 MHA6? 1 1 No Ct 16.5 + 2 1 No Ct 38.26 + 3 1 No Ct 16.53 + 4 1 No Ct 25 + 5 2 No Ct No Ct 6 2 No Ct No Ct 7 2 No Ct No Ct 8 2 17.01 + No Ct 9 3 No Ct 15.87 + 10 3 25.11 + 20.81 + 11 3 21.91 + 19.69 + 12 3 22.35 + 21.8 + 13 4 16.52 + No Ct 14 4 17.11 + No Ct 15 4 16.26 + No Ct 16 4 16 + No Ct 17 5 39.07 + 41.17 * Plot was bad ~NEG 18 5 15.98 + No Ct 19 5 16.4 + No Ct 20 5 16.44 + No Ct 21 6 17.08 + No Ct 22 6 18.24 + No Ct 23 6 16.8 + No Ct 24 6 17.94 + No Ct 25 7 17.98 + No Ct 26 7 No Ct 16.34 + 27 7 26.57 + 15.46 + 28 7 16.7 + 17.52 + 29 8 16.7 + No Ct 30 8 16.71 + No Ct 31 8 16.1 + No Ct 32 8 15.16 + No Ct 33 9 16.32 + No Ct 34 9 17.03 + No Ct 35 9 16.63 + No Ct 36 9 16.04 + No Ct 37 10 No Ct No Ct 38 10 No Ct No Ct 39 10 No Ct No Ct 40 10 No Ct No Ct 41 162 No Ct No Ct 42 162 No Ct No Ct 43 162 No Ct No Ct 44 162 No Ct No Ct 45 166 No Ct No Ct 46 166 No Ct No Ct 47 166 No Ct No Ct 48 166 No Ct No Ct 49 170 No Ct No Ct 50 170 No Ct No Ct 51 170 No Ct No Ct 52 170 No Ct No Ct 53 174 No Ct No Ct 54 174 No Ct No Ct 55 174 No Ct No Ct 56 174 No Ct No Ct 57 175 No Ct No Ct 58 175 No Ct No Ct 59 175 No Ct No Ct 60 175 No Ct No Ct 61 A1 mut + 16.66 No Ct 62 A6 mut + No Ct 13.85 63 A1 Wt + 15.87 No Ct 64 Neg 40.77 No Ct
TABLE-US-00013 TABLE 14 Day 15 Post Vaccination Animal # FAM MHA1 MHA1? CY5 MHA6 MHA6? Lkt del PCR 1 No Ct 40.53 1 No Ct No Ct 1 No Ct No Ct 1 No Ct No Ct 1 No Ct No Ct 2 No Ct No Ct 2 No Ct No Ct 2 No Ct No Ct 2 No Ct No Ct 2 No Ct No Ct 3 No Ct 15.1 + Mutant 3 No Ct 15.08 + Mutant 3 No Ct 15.19 + Mutant 3 No Ct 15.3 + Mutant 3 No Ct 15.1 + Mutant 4 15.82 No Ct Mutant 4 No Ct No Ct 4 No Ct No Ct 4 No Ct No Ct 4 No Ct No Ct 5 16.13 + No Ct Mutant 5 15.27 + No Ct Mutant 5 17.03 + No Ct Mutant 5 16.49 + No Ct Mutant 5 18.06 + No Ct Mutant 6 No Ct No Ct 6 No Ct No Ct 6 No Ct No Ct 6 40.05 No Ct 6 No Ct No Ct 7 No Ct 16.83 + Mutant 7 No Ct No Ct + 7 No Ct 14.92 + Mutant 7 No Ct 15.21 + Mutant 7 No Ct 16.16 + Mutant 8 No Ct No Ct 8 No Ct No Ct 8 No Ct No Ct 8 No Ct No Ct 8 No Ct No Ct 9 No Ct No Ct 9 No Ct No Ct 9 No Ct No Ct 9 No Ct No Ct 9 No Ct No Ct 10 15.94 + No Ct Mutant 10 No Ct + No Ct 10 No Ct + No Ct 10 23.82 + No Ct Mutant 10 30.04 + No Ct Mutant 162 No Ct No Ct 162 No Ct No Ct 162 No Ct No Ct 162 No Ct No Ct 162 No Ct No Ct 166 No Ct No Ct 166 No Ct No Ct 166 No Ct No Ct 166 No Ct No Ct 166 No Ct No Ct 170 No Ct No Ct 170 No Ct No Ct 170 No Ct No Ct 170 No Ct No Ct 170 No Ct No Ct 174 No Ct No Ct 174 No Ct No Ct 174 No Ct No Ct 174 No Ct No Ct 174 No Ct No Ct 175 16.24 + No Ct Mutant 175 No Ct + No Ct 175 16.54 + No Ct Mutant 175 No Ct + No Ct Mutant 175 23.06 + No Ct Mutant
[0118] Having thus described in detail preferred embodiments of the present invention, it is to be understood that the invention defined by the above paragraphs is not to be limited to particular details set forth in the above description as many apparent variations thereof are possible without departing from the spirit or scope of the present invention.
Sequence CWU 1
1
26126DNAArtificial SequencelktCAf primer 1gcattgaatt gatcaactaa
tacttg 26225DNAArtificial SequencelktCAr primer 2caaggtttct
agaaagattt ttcgg 25331DNAArtificial SequencelktCAdelf primer
3gatcaattga aagctgttga agaaattatc g 31429DNAArtificial
SequencelktCAdelr primer 4atacaattga ttcataattt gcactcgat
29548DNAArtificial SequencelktRBSr primer 5caacaattga ttcataattt
gcctcctata attattctaa attaggtc 4861068DNAArtificial Sequence5'
deltalktCA PCR fragment 6gcattgaatt gatcaactaa tacttggttt
ttcaagtgag ttgcaatgcc taaaccatca 60ccaaaatagt ttggattatt gattttctcc
cctacaaaat ctagcccttc gtgttttctt 120gccatctcag ccaataccgg
cacatcgcca aaaatagcat caattcgccc attttgcaca 180tctaaaatag
cattttgata agaggcataa gatttcacat tgtactcttt tttctctttt
240gctaaatagt gttggtaagt agtcccattt tgcacaccaa tcgttttcac
cttagcaaaa 300tctgtatctt ttttcgcaat gaaggcagca gagcttggaa
agtaaggctc gctaaataat 360acttgtttct tacgtggttc cgtaataccc
atacctgaaa ttgcagcatc aaattgtttt 420tgttttaggc tttggattaa
gctatcaaaa ggttggctat ggaatgtaca atttgcattc 480atctctttac
agatagcatt tgcaatatcc acatcaaaac cgataatttc tcccttctct
540tcggtcattt caaatggagg atagcttggc tccatcacaa atttgatatc
ttgtgcctgc 600gcagtaacca cacacccgaa taaaagggtc aaaagtgttt
ttttcataaa aagtccctgt 660gttttcatta taaggattac cactttaacg
cagttacttt cttaaaaaaa gtcttctttt 720cataaagttt gttttatgtc
atacaaacac atcaaattga gatgtagttt ctcaatcctc 780ttgattcctc
tatctcaaaa aaacaaccca aaagaaaaaa gaaaagtata tgttacatta
840atattacaat gtaattattt tgtttaattt ccctacattt tgtataactt
taaaacactc 900ctttttctct tctgattata taaaagacaa aaaatacaat
ttaagctaca aaaaacaaca 960aaaaacaaca aaaaacacga caataagatc
gagtaatgat tatattatgt tataattttt 1020gacctaattt agaataatta
tcgagtccaa attatgaatc aattgtat 106871295DNAArtificial Sequence3'
deltalktCA PCR fragment 7caattgaaag ctgttgaaga aattatcggt
acatcacata acgatatctt taaaggtagt 60aagttcaatg atgcctttaa cggtggtgat
ggtgtcgata ctattgacgg taacgacggc 120aatgaccgct tatttggtgg
taaaggcgat gatattctcg atggtggaaa tggtgatgat 180tttatcgatg
gcggtaaagg caacgaccta ttacacggtg gcaagggcga tgatattttc
240gttcaccgta aaggcgatgg taatgatatt attaccgatt ctgacggcaa
tgataaatta 300tcattctctg attcgaactt aaaagattta acatttgaaa
aagttaaaca taatcttgtc 360atcacgaata gcaaaaaaga gaaagtgacc
attcaaaact ggttccgaga ggctgatttt 420gctaaagaag tgcctaatta
taaagcaact aaagatgaga aaatcgaaga aatcatcggt 480caaaatggcg
agcggatcac ctcaaagcaa gttgatgatc ttatcgcaaa aggtaacggc
540aaaattaccc aagatgagct atcaaaagtt gttgataact atgaattgct
caaacatagc 600aaaaatgtga caaacagctt agataagtta atctcatctg
taagtgcatt tacctcgtct 660aatgattcga gaaatgtatt agtggctcca
acttcaatgt tggatcaaag tttatcttct 720cttcaatttg ctagagcagc
ttaattttta atgattggca actctatatt gtttcacaca 780ttatagagtt
gccgttttat tttataaaag gagacaatat ggaagctaac catcaaagga
840atgatcttgg tttagttgcc ctcactatgt tggcacaata ccataatatt
tcgcttaatc 900cggaagaaat aaaacataaa tttgatcttg acggaaaagg
gctttcttta actgcttggc 960ttttagctgc aaaatcgtta gcgttgaaag
cgaaacacat taaaaaagag atttcccgct 1020tacacttggt gaatttaccg
gcattagttt ggcaagataa cggtaaacat tttttattgg 1080taaaagtgga
taccgataat aaccgctatt taacttacaa tttggaacaa gatgctccac
1140aaattctgtc acaagacgaa tttgaagcct gctatcaagg gcagttaatt
ttggtcacgt 1200ccagagcttc cgtagtaggt caattagcaa agttcgattt
cacctggttt attccggcgg 1260tgatcaaata ccgaaaaatc tttctagaaa ccttg
1295827DNAArtificial SequenceBCCT FAMILY-BETAINE-CARNITINE-CHOLINE
Transporter forward primer 8atgttattcg ccgccggaat ggggatc
27925DNAArtificial SequenceBCCT FAMILY-BETAINE-CARNITINE-CHOLINE
Transporter reverse primer 9acctgcatca ccccaaagcc aagtg
251026DNAArtificial SequenceIGA SPECIFIC SERINE
METALLO-ENDOPEPTIDASE forward primer 10atgaagacca aaacatttac tcgttc
261125DNAArtificial SequenceIGA SPECIFIC SERINE
METALLO-ENDOPEPTIDASE reverse primer 11agcgcttgtg tccctgaacc agcac
25122007DNAArtificial SequenceA6 specific gene-BCCT
FAMILY/BETAINE/CARNITINE/ CHOLINE transporter 12ttggatttaa
tcaaaaaatt aaacacagga agtaccttta gggtaccgat tttcctaccg 60agtttactct
ttgtcagctt tgttgccgtt ttctgtatca tctttccaca gcaagcacaa
120acctcacttg ataccatcaa aaatagtctc ttccaacatt ttagctggtt
ctatattttt 180gcaggctcta tctttttcct gtttctaatt tttctctctt
tcagccgatt gggtgatatt 240aaattagggg cagataccga tgagcctgaa
tttggttttg gctcttggat tgcgatgtta 300ttcgccgccg gaatggggat
cgggttaatg tattttgggg tagcagaacc tattttgcat 360taccttaaac
ccgtccaaca aaatttaact gagccggagc gtatgaaaga agcgatgatg
420acaacgttct atcattgggg tattcacgct tgggcaattt atggtgtgat
tgccttagct 480cttgcttatt ttggcttcag atataagtta gcactcacta
ttcgttccgg attttatccc 540ttactaaaac atcgtatttc aggcttctgg
gggcatttaa ttgatattat tgccctttgt 600agcacgattt tcggtttaac
gactacactt ggctttgggg tgatgcaggt cagtgctggc 660tttaacaatc
taggtttaat tgaacagagc aattttactg ttcttgcgat tatcgtaaca
720gtagcaatgg ctcttgccgt gttatctgcc gtttcgggcg taggcaaagg
ggttaaaatc 780ttaagtgaaa tcaatctcac attagccgga ttgctactta
tttttgtgat aatcaccggc 840ccaactctat tacttttctc aagcttcacc
gaaaatttag gctattattt tagctcgctg 900cttgagatga gtttccgtac
cttcgcttat gaaccggaac atcaaggctg gctaagcggc 960tggacggtcc
tttattgggc atggtgggca tcttgggcgc catttgttgg tttgtttatt
1020gccaagatct ctaaaggcag aaccattcgt gaatttattt taggggtgct
atttgttcca 1080tcgctgttta acattttatg gatgaccagc ttcggcagct
ctgccatttg gttcgatcaa 1140caaactgccg gtgctttagc tgaagtcagc
ggcaataccg aacaactgtt atttaccttt 1200tttgagcaat taccgtttgg
ctctattgcc tctttcgttg ccgtcattgt tatcagtatt 1260ttctttatca
cctctgccga ctcggggatt tttgttctca acagcattgc ttcacaaggc
1320gaagaaaatg caccgaaatg gcaaagcgtg ctttggggag cattattagc
catcttagcg 1380ttatcactac tctattcggg tggcttggct tctctgcaaa
caatgacact gattatcgcc 1440ttaccattta ccttcattat gctgattctc
tgtatcggct tatggaaagg attaatggta 1500gataaccaat acttcaacaa
aaaattctcg caaggtagcc aacattgggc gggtaaagat 1560tggaaacaac
gcttggagaa aatcatcaac ccaagcaata agcaagatgt ccgtcacttc
1620tttattaaag ttgccagacc agcattttta gaacttatcg aggaatttga
aagctatggc 1680ttaatcgcta aaatgaattt caccaacgaa caaaacccga
aattagagtt tgaagtggtg 1740aaagaaaatt tacgcaattt catttacggc
attgaaagtg tgccacggga attatcggat 1800ttggtggtag gtgacgacaa
cctaccgaac attgagcaaa ataccattta cgagccgatt 1860acttatttct
tagacgggcg gaaaggttat gatgtgcaat atatgaccaa agaagagttg
1920attgccgacg tgctgcaaca gtatgaacgc tttatcaatt tagcgatgga
caactcgcac 1980gacttaatga cggctgattt caatcac 200713669PRTArtificial
SequenceA6 specific BCCT FAMILY/BETAINE/CARNITINE/ CHOLINE
transporter 13Leu Asp Leu Ile Lys Lys Leu Asn Thr Gly Ser Thr Phe
Arg Val Pro 1 5 10 15 Ile Phe Leu Pro Ser Leu Leu Phe Val Ser Phe
Val Ala Val Phe Cys 20 25 30 Ile Ile Phe Pro Gln Gln Ala Gln Thr
Ser Leu Asp Thr Ile Lys Asn 35 40 45 Ser Leu Phe Gln His Phe Ser
Trp Phe Tyr Ile Phe Ala Gly Ser Ile 50 55 60 Phe Phe Leu Phe Leu
Ile Phe Leu Ser Phe Ser Arg Leu Gly Asp Ile 65 70 75 80 Lys Leu Gly
Ala Asp Thr Asp Glu Pro Glu Phe Gly Phe Gly Ser Trp 85 90 95 Ile
Ala Met Leu Phe Ala Ala Gly Met Gly Ile Gly Leu Met Tyr Phe 100 105
110 Gly Val Ala Glu Pro Ile Leu His Tyr Leu Lys Pro Val Gln Gln Asn
115 120 125 Leu Thr Glu Pro Glu Arg Met Lys Glu Ala Met Met Thr Thr
Phe Tyr 130 135 140 His Trp Gly Ile His Ala Trp Ala Ile Tyr Gly Val
Ile Ala Leu Ala 145 150 155 160 Leu Ala Tyr Phe Gly Phe Arg Tyr Lys
Leu Ala Leu Thr Ile Arg Ser 165 170 175 Gly Phe Tyr Pro Leu Leu Lys
His Arg Ile Ser Gly Phe Trp Gly His 180 185 190 Leu Ile Asp Ile Ile
Ala Leu Cys Ser Thr Ile Phe Gly Leu Thr Thr 195 200 205 Thr Leu Gly
Phe Gly Val Met Gln Val Ser Ala Gly Phe Asn Asn Leu 210 215 220 Gly
Leu Ile Glu Gln Ser Asn Phe Thr Val Leu Ala Ile Ile Val Thr 225 230
235 240 Val Ala Met Ala Leu Ala Val Leu Ser Ala Val Ser Gly Val Gly
Lys 245 250 255 Gly Val Lys Ile Leu Ser Glu Ile Asn Leu Thr Leu Ala
Gly Leu Leu 260 265 270 Leu Ile Phe Val Ile Ile Thr Gly Pro Thr Leu
Leu Leu Phe Ser Ser 275 280 285 Phe Thr Glu Asn Leu Gly Tyr Tyr Phe
Ser Ser Leu Leu Glu Met Ser 290 295 300 Phe Arg Thr Phe Ala Tyr Glu
Pro Glu His Gln Gly Trp Leu Ser Gly 305 310 315 320 Trp Thr Val Leu
Tyr Trp Ala Trp Trp Ala Ser Trp Ala Pro Phe Val 325 330 335 Gly Leu
Phe Ile Ala Lys Ile Ser Lys Gly Arg Thr Ile Arg Glu Phe 340 345 350
Ile Leu Gly Val Leu Phe Val Pro Ser Leu Phe Asn Ile Leu Trp Met 355
360 365 Thr Ser Phe Gly Ser Ser Ala Ile Trp Phe Asp Gln Gln Thr Ala
Gly 370 375 380 Ala Leu Ala Glu Val Ser Gly Asn Thr Glu Gln Leu Leu
Phe Thr Phe 385 390 395 400 Phe Glu Gln Leu Pro Phe Gly Ser Ile Ala
Ser Phe Val Ala Val Ile 405 410 415 Val Ile Ser Ile Phe Phe Ile Thr
Ser Ala Asp Ser Gly Ile Phe Val 420 425 430 Leu Asn Ser Ile Ala Ser
Gln Gly Glu Glu Asn Ala Pro Lys Trp Gln 435 440 445 Ser Val Leu Trp
Gly Ala Leu Leu Ala Ile Leu Ala Leu Ser Leu Leu 450 455 460 Tyr Ser
Gly Gly Leu Ala Ser Leu Gln Thr Met Thr Leu Ile Ile Ala 465 470 475
480 Leu Pro Phe Thr Phe Ile Met Leu Ile Leu Cys Ile Gly Leu Trp Lys
485 490 495 Gly Leu Met Val Asp Asn Gln Tyr Phe Asn Lys Lys Phe Ser
Gln Gly 500 505 510 Ser Gln His Trp Ala Gly Lys Asp Trp Lys Gln Arg
Leu Glu Lys Ile 515 520 525 Ile Asn Pro Ser Asn Lys Gln Asp Val Arg
His Phe Phe Ile Lys Val 530 535 540 Ala Arg Pro Ala Phe Leu Glu Leu
Ile Glu Glu Phe Glu Ser Tyr Gly 545 550 555 560 Leu Ile Ala Lys Met
Asn Phe Thr Asn Glu Gln Asn Pro Lys Leu Glu 565 570 575 Phe Glu Val
Val Lys Glu Asn Leu Arg Asn Phe Ile Tyr Gly Ile Glu 580 585 590 Ser
Val Pro Arg Glu Leu Ser Asp Leu Val Val Gly Asp Asp Asn Leu 595 600
605 Pro Asn Ile Glu Gln Asn Thr Ile Tyr Glu Pro Ile Thr Tyr Phe Leu
610 615 620 Asp Gly Arg Lys Gly Tyr Asp Val Gln Tyr Met Thr Lys Glu
Glu Leu 625 630 635 640 Ile Ala Asp Val Leu Gln Gln Tyr Glu Arg Phe
Ile Asn Leu Ala Met 645 650 655 Asp Asn Ser His Asp Leu Met Thr Ala
Asp Phe Asn His 660 665 144017DNAArtificial SequenceA1 specific
gene - IGA specific serine metallo- endopeptidase 14atgaagacca
aaacatttac tcgttcttat cttgcttctt ttgtaacaat cgtattaagt 60ttacctgctg
tagcatctgt tgtacgtaat gatgtggact atcaatactt ccgcgatttt
120gccgaaaata aaggaccatt ttcagttggt tcaatgaata ttgatattaa
agacaacaat 180ggacaacttg taggcacgat gcttcataat ttaccaatgg
ttgattttag tgctatggta 240agaggtggat attctacttt aattgcacca
caatatttag ttagtgttgc acataatact 300ggatataaaa atgttcaatt
tggtgctgca ggttataacc ctgattcaca tcactatact 360tataaaattg
ttgaccgcaa tgattatgaa aaggttcaag gagggttgca cccagactat
420catactcctc gattaaataa attagtaaca gaagttgtgc ctgccgcagt
caccaatgca 480ggtacatcta ttaaacccta cttaaatgaa gaacgcttcc
ctatgtttct tcgtgctggt 540tcagggacac aagcgctaag aggaaaagaa
agtaataaaa caactggaat cgctggtgct 600tatgaatatc ttactggcgg
taccacatta caattatcta aaagctcccc tgatcactgg 660ttagattatt
caagtaacct ttatcaagta agctatggac cactttcaac ctatgcacta
720cctggtgata gtggttcagg ttcttacgcc tatgatatga acgaaaaacg
atgggtatta 780gttggtgtgc tcaatttcta taatggtatg gataatcaat
tcaaccgctc tgcgattatc 840cgtaaagatt tccacgagaa aaaatttgcc
gaagatattg caggaacaat caataatacc 900gtacaaaatg cacaattcaa
ttggactgct caaggtaaat ccagctctct tagtcaatca 960aataatgtgc
aaaaactcaa cgttgatcta aaagatagta gcattgcaaa ccaaaacact
1020tctctgccac aagaaaatca cggtaaaacc attaatttta atggtaaaga
tgcaactatt 1080gtactaaaac aggatattga ccaaggtgca ggtgcattaa
atctgaacgc taatctcact 1140attcgtcctg aaacagacca aacttggcaa
ggtgcaggta ttatcgtcgg taaagataaa 1200aaagtgaatt ggcaagtaaa
aaatccacaa ggcgatcgtt tatctaaact cggggaagga 1260acactctatg
taaatggacg tggacagaat cttggcgata tcagtgtagg tgatggtatt
1320gtaatactta accagcaagc cgatcaccaa ggaagaaaac aggcctttaa
tacagtagga 1380atcgtaagtg gtcgccccac tgttgtgcta ggtagtgcag
atcaagttaa tcccgataat 1440atttactttg gatttcgcgg aggtcgttta
gacctaaacg gtaacagcat cgcctttaaa 1500cgtattcaaa acagcgataa
acatgctcgt attgtaaacc acaatcgcga tcacatttct 1560accttaataa
tacaaggcca agatcctctc actagtaatg atcttatatg gggaaaatgg
1620gcaagtaata gcccagcaga catttacgaa tataccaatc cttatcaaaa
taaacgcaaa 1680gattacttcc gtctgaaagg taattcgaga gtatattatc
caacgaatgc tacaagtaac 1740gatcactggg aatttctttc cagtaaccgc
gagcaagcaa tacagaaaat cctagatgcc 1800aaaaacttaa gacagcgcta
tgacacgttt aatggtttta taggggaaga tgcttccaat 1860aaaactaatg
ggatattaaa tgtcgtgttt gatacaaaaa cagaagtaaa tacagaacaa
1920gataaattaa agaatatcta cacaatgtcg ggaggattta accttaatgg
tgaactcacc 1980cttaaaggtg gtacattgtt gctttctggt cacccaacgc
cacacgctta tgatattaag 2040aataagcatg atgttgtgcg tgaaaacgat
tggcaagaca gccattttac tgctaaaaat 2100atcacggtaa ataaaatggc
acaactctat atcgggagaa atgtcaatga agtaaatagt 2160cactttactg
cgactgataa agccaaactc aatttaggat ttattaatcg ttcaacgcca
2220agttgctatg attctgaata cacaggcact acacattgtg aagtgcaagc
ggtcatttcc 2280gataatattt ttgcaaatct agcaacaacc gccattaaag
gtaatgttaa attacaaaac 2340catagccaat taaatttagg caaagcaaac
ctcactggtt ctgtacaagc tgatcaaaca 2400actcatatca ctttagcaaa
tcacagtcac tggttaaaca atggtacgag ccagattggg 2460catcttacaa
tggaaaaagg gtcgatcctt agcctaaacg ataaatttgc taccacggaa
2520atcccagtcc gattcaacaa gatgatcatc caaggtaatc taaaaggtaa
tggacgaatt 2580aactataccg caaatttagc caagggcgaa tctgatcatc
tccaagttga cggtattgct 2640gaaggaaatt ttgtccttgc cgttagaaat
agcacaactg aagcaaatcc aaaaagctca 2700ttaaacctac taagcttaaa
aaatagcaac caagaaggca ataaagcttc tatttctcta 2760gaaaataatt
atgttgatct aggtacttat cgttatgtat tagaaaatcg taatcacaat
2820taccatttat ttaatccatt aataccaaat tcaacctcta aagagatgaa
tgctacatct 2880gtatcctcta ttccaaaaaa ggaatctgtt actaatgttc
ctactttaga taagaaagaa 2940actgaacaaa atcttactca actacaaaaa
gatttttcag cacaccaatt agaaaatcaa 3000aaagcaaaac aatctatgat
aaatgctcaa tctgagctaa gacgactcaa ttcacaactg 3060aatgtattgc
aaaaatatgt gaattctcgt cgcttaggtt actatactca gcaggcagtt
3120ttagaacaaa ttagcattat tcaaaataaa attaaacaaa cacaaacaat
atttaatgac 3180gctaatgcaa ctgtaaaact cacagatcaa aagctagaag
aagccaaatt agctctaggc 3240tctgtaaacg atcttgtatt aataaaagcc
tctgctccag caatgcaagc aactaatcaa 3300gatacgagta tgatgaatat
tattcaagca gattggataa gccaatacgc taacacagca 3360ctttctgaac
tctcggcaca ggctaattct gctctgcaaa tcagtaatag cttagatcgc
3420caactcttca aacaaagcga taaattcaac gtatggagca gcgtcgaaca
tcagaaaacc 3480gagcataaat cagatttata ccgcccgtat aaacaacaaa
ccaacctgac ccaactgggc 3540atacaaatgc cgatagataa cggtttaatg
tttggagttg cattatctaa aaaccacgct 3600aacgcggaat ttaacgaggg
tgtaaacggt aaatcgaatc tactaatggc aagcctatat 3660ggtaagtggc
aatctcaaca aggcactttt atcagccttg atggcagcta cggtaaagca
3720aaaaaccaac tctacctatt tggtgaaaac cactttaccc gccgaatttc
ctctattggt 3780gctaacattg gacatcaatt tgacctcgca ggagttcaaa
ttcagccaac aataggagca 3840agatactacc atttcagcgg ccaagactat
acactaggag gagcgaaaat cagctcacca 3900aatacccact ttatgacata
tcaagcgggt ctaaaagcta gtaaaacttt tcattggatg 3960actggaaagt
tgaaccaagc attacaaccc actatgtgga tgcaagtaac aaacgct
4017151339PRTArtificial SequenceA1 specific IGA specific serine
metallo- endopeptidase 15Met Lys Thr Lys Thr Phe Thr Arg Ser Tyr
Leu Ala Ser Phe Val Thr 1 5 10 15 Ile Val Leu Ser Leu Pro Ala Val
Ala Ser Val Val Arg Asn Asp Val 20 25 30 Asp Tyr Gln Tyr Phe Arg
Asp Phe Ala Glu Asn Lys Gly Pro Phe Ser 35 40 45 Val Gly Ser Met
Asn Ile Asp Ile Lys Asp Asn Asn Gly Gln Leu Val 50 55 60 Gly Thr
Met Leu His Asn Leu Pro Met Val Asp Phe Ser Ala Met Val 65 70 75 80
Arg Gly Gly Tyr Ser Thr Leu Ile Ala Pro Gln Tyr Leu Val Ser Val 85
90 95 Ala His Asn Thr Gly Tyr Lys Asn Val Gln Phe Gly Ala Ala Gly
Tyr 100 105 110
Asn Pro Asp Ser His His Tyr Thr Tyr Lys Ile Val Asp Arg Asn Asp 115
120 125 Tyr Glu Lys Val Gln Gly Gly Leu His Pro Asp Tyr His Thr Pro
Arg 130 135 140 Leu Asn Lys Leu Val Thr Glu Val Val Pro Ala Ala Val
Thr Asn Ala 145 150 155 160 Gly Thr Ser Ile Lys Pro Tyr Leu Asn Glu
Glu Arg Phe Pro Met Phe 165 170 175 Leu Arg Ala Gly Ser Gly Thr Gln
Ala Leu Arg Gly Lys Glu Ser Asn 180 185 190 Lys Thr Thr Gly Ile Ala
Gly Ala Tyr Glu Tyr Leu Thr Gly Gly Thr 195 200 205 Thr Leu Gln Leu
Ser Lys Ser Ser Pro Asp His Trp Leu Asp Tyr Ser 210 215 220 Ser Asn
Leu Tyr Gln Val Ser Tyr Gly Pro Leu Ser Thr Tyr Ala Leu 225 230 235
240 Pro Gly Asp Ser Gly Ser Gly Ser Tyr Ala Tyr Asp Met Asn Glu Lys
245 250 255 Arg Trp Val Leu Val Gly Val Leu Asn Phe Tyr Asn Gly Met
Asp Asn 260 265 270 Gln Phe Asn Arg Ser Ala Ile Ile Arg Lys Asp Phe
His Glu Lys Lys 275 280 285 Phe Ala Glu Asp Ile Ala Gly Thr Ile Asn
Asn Thr Val Gln Asn Ala 290 295 300 Gln Phe Asn Trp Thr Ala Gln Gly
Lys Ser Ser Ser Leu Ser Gln Ser 305 310 315 320 Asn Asn Val Gln Lys
Leu Asn Val Asp Leu Lys Asp Ser Ser Ile Ala 325 330 335 Asn Gln Asn
Thr Ser Leu Pro Gln Glu Asn His Gly Lys Thr Ile Asn 340 345 350 Phe
Asn Gly Lys Asp Ala Thr Ile Val Leu Lys Gln Asp Ile Asp Gln 355 360
365 Gly Ala Gly Ala Leu Asn Leu Asn Ala Asn Leu Thr Ile Arg Pro Glu
370 375 380 Thr Asp Gln Thr Trp Gln Gly Ala Gly Ile Ile Val Gly Lys
Asp Lys 385 390 395 400 Lys Val Asn Trp Gln Val Lys Asn Pro Gln Gly
Asp Arg Leu Ser Lys 405 410 415 Leu Gly Glu Gly Thr Leu Tyr Val Asn
Gly Arg Gly Gln Asn Leu Gly 420 425 430 Asp Ile Ser Val Gly Asp Gly
Ile Val Ile Leu Asn Gln Gln Ala Asp 435 440 445 His Gln Gly Arg Lys
Gln Ala Phe Asn Thr Val Gly Ile Val Ser Gly 450 455 460 Arg Pro Thr
Val Val Leu Gly Ser Ala Asp Gln Val Asn Pro Asp Asn 465 470 475 480
Ile Tyr Phe Gly Phe Arg Gly Gly Arg Leu Asp Leu Asn Gly Asn Ser 485
490 495 Ile Ala Phe Lys Arg Ile Gln Asn Ser Asp Lys His Ala Arg Ile
Val 500 505 510 Asn His Asn Arg Asp His Ile Ser Thr Leu Ile Ile Gln
Gly Gln Asp 515 520 525 Pro Leu Thr Ser Asn Asp Leu Ile Trp Gly Lys
Trp Ala Ser Asn Ser 530 535 540 Pro Ala Asp Ile Tyr Glu Tyr Thr Asn
Pro Tyr Gln Asn Lys Arg Lys 545 550 555 560 Asp Tyr Phe Arg Leu Lys
Gly Asn Ser Arg Val Tyr Tyr Pro Thr Asn 565 570 575 Ala Thr Ser Asn
Asp His Trp Glu Phe Leu Ser Ser Asn Arg Glu Gln 580 585 590 Ala Ile
Gln Lys Ile Leu Asp Ala Lys Asn Leu Arg Gln Arg Tyr Asp 595 600 605
Thr Phe Asn Gly Phe Ile Gly Glu Asp Ala Ser Asn Lys Thr Asn Gly 610
615 620 Ile Leu Asn Val Val Phe Asp Thr Lys Thr Glu Val Asn Thr Glu
Gln 625 630 635 640 Asp Lys Leu Lys Asn Ile Tyr Thr Met Ser Gly Gly
Phe Asn Leu Asn 645 650 655 Gly Glu Leu Thr Leu Lys Gly Gly Thr Leu
Leu Leu Ser Gly His Pro 660 665 670 Thr Pro His Ala Tyr Asp Ile Lys
Asn Lys His Asp Val Val Arg Glu 675 680 685 Asn Asp Trp Gln Asp Ser
His Phe Thr Ala Lys Asn Ile Thr Val Asn 690 695 700 Lys Met Ala Gln
Leu Tyr Ile Gly Arg Asn Val Asn Glu Val Asn Ser 705 710 715 720 His
Phe Thr Ala Thr Asp Lys Ala Lys Leu Asn Leu Gly Phe Ile Asn 725 730
735 Arg Ser Thr Pro Ser Cys Tyr Asp Ser Glu Tyr Thr Gly Thr Thr His
740 745 750 Cys Glu Val Gln Ala Val Ile Ser Asp Asn Ile Phe Ala Asn
Leu Ala 755 760 765 Thr Thr Ala Ile Lys Gly Asn Val Lys Leu Gln Asn
His Ser Gln Leu 770 775 780 Asn Leu Gly Lys Ala Asn Leu Thr Gly Ser
Val Gln Ala Asp Gln Thr 785 790 795 800 Thr His Ile Thr Leu Ala Asn
His Ser His Trp Leu Asn Asn Gly Thr 805 810 815 Ser Gln Ile Gly His
Leu Thr Met Glu Lys Gly Ser Ile Leu Ser Leu 820 825 830 Asn Asp Lys
Phe Ala Thr Thr Glu Ile Pro Val Arg Phe Asn Lys Met 835 840 845 Ile
Ile Gln Gly Asn Leu Lys Gly Asn Gly Arg Ile Asn Tyr Thr Ala 850 855
860 Asn Leu Ala Lys Gly Glu Ser Asp His Leu Gln Val Asp Gly Ile Ala
865 870 875 880 Glu Gly Asn Phe Val Leu Ala Val Arg Asn Ser Thr Thr
Glu Ala Asn 885 890 895 Pro Lys Ser Ser Leu Asn Leu Leu Ser Leu Lys
Asn Ser Asn Gln Glu 900 905 910 Gly Asn Lys Ala Ser Ile Ser Leu Glu
Asn Asn Tyr Val Asp Leu Gly 915 920 925 Thr Tyr Arg Tyr Val Leu Glu
Asn Arg Asn His Asn Tyr His Leu Phe 930 935 940 Asn Pro Leu Ile Pro
Asn Ser Thr Ser Lys Glu Met Asn Ala Thr Ser 945 950 955 960 Val Ser
Ser Ile Pro Lys Lys Glu Ser Val Thr Asn Val Pro Thr Leu 965 970 975
Asp Lys Lys Glu Thr Glu Gln Asn Leu Thr Gln Leu Gln Lys Asp Phe 980
985 990 Ser Ala His Gln Leu Glu Asn Gln Lys Ala Lys Gln Ser Met Ile
Asn 995 1000 1005 Ala Gln Ser Glu Leu Arg Arg Leu Asn Ser Gln Leu
Asn Val Leu 1010 1015 1020 Gln Lys Tyr Val Asn Ser Arg Arg Leu Gly
Tyr Tyr Thr Gln Gln 1025 1030 1035 Ala Val Leu Glu Gln Ile Ser Ile
Ile Gln Asn Lys Ile Lys Gln 1040 1045 1050 Thr Gln Thr Ile Phe Asn
Asp Ala Asn Ala Thr Val Lys Leu Thr 1055 1060 1065 Asp Gln Lys Leu
Glu Glu Ala Lys Leu Ala Leu Gly Ser Val Asn 1070 1075 1080 Asp Leu
Val Leu Ile Lys Ala Ser Ala Pro Ala Met Gln Ala Thr 1085 1090 1095
Asn Gln Asp Thr Ser Met Met Asn Ile Ile Gln Ala Asp Trp Ile 1100
1105 1110 Ser Gln Tyr Ala Asn Thr Ala Leu Ser Glu Leu Ser Ala Gln
Ala 1115 1120 1125 Asn Ser Ala Leu Gln Ile Ser Asn Ser Leu Asp Arg
Gln Leu Phe 1130 1135 1140 Lys Gln Ser Asp Lys Phe Asn Val Trp Ser
Ser Val Glu His Gln 1145 1150 1155 Lys Thr Glu His Lys Ser Asp Leu
Tyr Arg Pro Tyr Lys Gln Gln 1160 1165 1170 Thr Asn Leu Thr Gln Leu
Gly Ile Gln Met Pro Ile Asp Asn Gly 1175 1180 1185 Leu Met Phe Gly
Val Ala Leu Ser Lys Asn His Ala Asn Ala Glu 1190 1195 1200 Phe Asn
Glu Gly Val Asn Gly Lys Ser Asn Leu Leu Met Ala Ser 1205 1210 1215
Leu Tyr Gly Lys Trp Gln Ser Gln Gln Gly Thr Phe Ile Ser Leu 1220
1225 1230 Asp Gly Ser Tyr Gly Lys Ala Lys Asn Gln Leu Tyr Leu Phe
Gly 1235 1240 1245 Glu Asn His Phe Thr Arg Arg Ile Ser Ser Ile Gly
Ala Asn Ile 1250 1255 1260 Gly His Gln Phe Asp Leu Ala Gly Val Gln
Ile Gln Pro Thr Ile 1265 1270 1275 Gly Ala Arg Tyr Tyr His Phe Ser
Gly Gln Asp Tyr Thr Leu Gly 1280 1285 1290 Gly Ala Lys Ile Ser Ser
Pro Asn Thr His Phe Met Thr Tyr Gln 1295 1300 1305 Ala Gly Leu Lys
Ala Ser Lys Thr Phe His Trp Met Thr Gly Lys 1310 1315 1320 Leu Asn
Gln Ala Leu Gln Pro Thr Met Trp Met Gln Val Thr Asn 1325 1330 1335
Ala 162354DNAArtificial SequenceComplete deltaLKTCA with original
RBS 16gcattgaatt gatcaactaa tacttggttt ttcaagtgag ttgcaatgcc
taaaccatca 60ccaaaatagt ttggattatt gattttctcc cctacaaaat ctagcccttc
gtgttttctt 120gccatctcag ccaataccgg cacatcgcca aaaatagcat
caattcgccc attttgcaca 180tctaaaatag cattttgata agaggcataa
gatttcacat tgtactcttt tttctctttt 240gctaaatagt gttggtaagt
agtcccattt tgcacaccaa tcgttttcac cttagcaaaa 300tctgtatctt
ttttcgcaat gaaggcagca gagcttggaa agtaaggctc gctaaataat
360acttgtttct tacgtggttc cgtaataccc atacctgaaa ttgcagcatc
aaattgtttt 420tgttttaggc tttggattaa gctatcaaaa ggttggctat
ggaatgtaca atttgcattc 480atctctttac agatagcatt tgcaatatcc
acatcaaaac cgataatttc tcccttctct 540tcggtcattt caaatggagg
atagcttggc tccatcacaa atttgatatc ttgtgcctgc 600gcagtaacca
cacacccgaa taaaagggtc aaaagtgttt ttttcataaa aagtccctgt
660gttttcatta taaggattac cactttaacg cagttacttt cttaaaaaaa
gtcttctttt 720cataaagttt gttttatgtc atacaaacac atcaaattga
gatgtagttt ctcaatcctc 780ttgattcctc tatctcaaaa aaacaaccca
aaagaaaaaa gaaaagtata tgttacatta 840atattacaat gtaattattt
tgtttaattt ccctacattt tgtataactt taaaacactc 900ctttttctct
tctgattata taaaagacaa aaaatacaat ttaagctaca aaaaacaaca
960aaaaacaaca aaaaacacga caataagatc gagtaatgat tatattatgt
tataattttt 1020gacctaattt agaataatta tcgagtccaa attatgaatc
aattgaaagc tgttgaagaa 1080attatcggta catcacataa cgatatcttt
aaaggtagta agttcaatga tgcctttaac 1140ggtggtgatg gtgtcgatac
tattgacggt aacgacggca atgaccgctt atttggtggt 1200aaaggcgatg
atattctcga tggtggaaat ggtgatgatt ttatcgatgg cggtaaaggc
1260aacgacctat tacacggtgg caagggcgat gatattttcg ttcaccgtaa
aggcgatggt 1320aatgatatta ttaccgattc tgacggcaat gataaattat
cattctctga ttcgaactta 1380aaagatttaa catttgaaaa agttaaacat
aatcttgtca tcacgaatag caaaaaagag 1440aaagtgacca ttcaaaactg
gttccgagag gctgattttg ctaaagaagt gcctaattat 1500aaagcaacta
aagatgagaa aatcgaagaa atcatcggtc aaaatggcga gcggatcacc
1560tcaaagcaag ttgatgatct tatcgcaaaa ggtaacggca aaattaccca
agatgagcta 1620tcaaaagttg ttgataacta tgaattgctc aaacatagca
aaaatgtgac aaacagctta 1680gataagttaa tctcatctgt aagtgcattt
acctcgtcta atgattcgag aaatgtatta 1740gtggctccaa cttcaatgtt
ggatcaaagt ttatcttctc ttcaatttgc tagagcagct 1800taatttttaa
tgattggcaa ctctatattg tttcacacat tatagagttg ccgttttatt
1860ttataaaagg agacaatatg gaagctaacc atcaaaggaa tgatcttggt
ttagttgccc 1920tcactatgtt ggcacaatac cataatattt cgcttaatcc
ggaagaaata aaacataaat 1980ttgatcttga cggaaaaggg ctttctttaa
ctgcttggct tttagctgca aaatcgttag 2040cgttgaaagc gaaacacatt
aaaaaagaga tttcccgctt acacttggtg aatttaccgg 2100cattagtttg
gcaagataac ggtaaacatt ttttattggt aaaagtggat accgataata
2160accgctattt aacttacaat ttggaacaag atgctccaca aattctgtca
caagacgaat 2220ttgaagcctg ctatcaaggg cagttaattt tggtcacgtc
cagagcttcc gtagtaggtc 2280aattagcaaa gttcgatttc acctggttta
ttccggcggt gatcaaatac cgaaaaatct 2340ttctagaaac cttg
2354172354DNAArtificial SequenceComplete deltalktCA (with consensus
RBS) 17gcattgaatt gatcaactaa tacttggttt ttcaagtgag ttgcaatgcc
taaaccatca 60ccaaaatagt ttggattatt gattttctcc cctacaaaat ctagcccttc
gtgttttctt 120gccatctcag ccaataccgg cacatcgcca aaaatagcat
caattcgccc attttgcaca 180tctaaaatag cattttgata agaggcataa
gatttcacat tgtactcttt tttctctttt 240gctaaatagt gttggtaagt
agtcccattt tgcacaccaa tcgttttcac cttagcaaaa 300tctgtatctt
ttttcgcaat gaaggcagca gagcttggaa agtaaggctc gctaaataat
360acttgtttct tacgtggttc cgtaataccc atacctgaaa ttgcagcatc
aaattgtttt 420tgttttaggc tttggattaa gctatcaaaa ggttggctat
ggaatgtaca atttgcattc 480atctctttac agatagcatt tgcaatatcc
acatcaaaac cgataatttc tcccttctct 540tcggtcattt caaatggagg
atagcttggc tccatcacaa atttgatatc ttgtgcctgc 600gcagtaacca
cacacccgaa taaaagggtc aaaagtgttt ttttcataaa aagtccctgt
660gttttcatta taaggattac cactttaacg cagttacttt cttaaaaaaa
gtcttctttt 720cataaagttt gttttatgtc atacaaacac atcaaattga
gatgtagttt ctcaatcctc 780ttgattcctc tatctcaaaa aaacaaccca
aaagaaaaaa gaaaagtata tgttacatta 840atattacaat gtaattattt
tgtttaattt ccctacattt tgtataactt taaaacactc 900ctttttctct
tctgattata taaaagacaa aaaatacaat ttaagctaca aaaaacaaca
960aaaaacaaca aaaaacacga caataagatc gagtaatgat tatattatgt
tataattttt 1020gacctaattt agaataatta taggaggcaa attatgaatc
aattgaaagc tgttgaagaa 1080attatcggta catcacataa cgatatcttt
aaaggtagta agttcaatga tgcctttaac 1140ggtggtgatg gtgtcgatac
tattgacggt aacgacggca atgaccgctt atttggtggt 1200aaaggcgatg
atattctcga tggtggaaat ggtgatgatt ttatcgatgg cggtaaaggc
1260aacgacctat tacacggtgg caagggcgat gatattttcg ttcaccgtaa
aggcgatggt 1320aatgatatta ttaccgattc tgacggcaat gataaattat
cattctctga ttcgaactta 1380aaagatttaa catttgaaaa agttaaacat
aatcttgtca tcacgaatag caaaaaagag 1440aaagtgacca ttcaaaactg
gttccgagag gctgattttg ctaaagaagt gcctaattat 1500aaagcaacta
aagatgagaa aatcgaagaa atcatcggtc aaaatggcga gcggatcacc
1560tcaaagcaag ttgatgatct tatcgcaaaa ggtaacggca aaattaccca
agatgagcta 1620tcaaaagttg ttgataacta tgaattgctc aaacatagca
aaaatgtgac aaacagctta 1680gataagttaa tctcatctgt aagtgcattt
acctcgtcta atgattcgag aaatgtatta 1740gtggctccaa cttcaatgtt
ggatcaaagt ttatcttctc ttcaatttgc tagagcagct 1800taatttttaa
tgattggcaa ctctatattg tttcacacat tatagagttg ccgttttatt
1860ttataaaagg agacaatatg gaagctaacc atcaaaggaa tgatcttggt
ttagttgccc 1920tcactatgtt ggcacaatac cataatattt cgcttaatcc
ggaagaaata aaacataaat 1980ttgatcttga cggaaaaggg ctttctttaa
ctgcttggct tttagctgca aaatcgttag 2040cgttgaaagc gaaacacatt
aaaaaagaga tttcccgctt acacttggtg aatttaccgg 2100cattagtttg
gcaagataac ggtaaacatt ttttattggt aaaagtggat accgataata
2160accgctattt aacttacaat ttggaacaag atgctccaca aattctgtca
caagacgaat 2220ttgaagcctg ctatcaaggg cagttaattt tggtcacgtc
cagagcttcc gtagtaggtc 2280aattagcaaa gttcgatttc acctggttta
ttccggcggt gatcaaatac cgaaaaatct 2340ttctagaaac cttg
235418249PRTArtificial SequenceTranslation of deltaLKTCA 18Met Asn
Gln Leu Lys Ala Val Glu Glu Ile Ile Gly Thr Ser His Asn 1 5 10 15
Asp Ile Phe Lys Gly Ser Lys Phe Asn Asp Ala Phe Asn Gly Gly Asp 20
25 30 Gly Val Asp Thr Ile Asp Gly Asn Asp Gly Asn Asp Arg Leu Phe
Gly 35 40 45 Gly Lys Gly Asp Asp Ile Leu Asp Gly Gly Asn Gly Asp
Asp Phe Ile 50 55 60 Asp Gly Gly Lys Gly Asn Asp Leu Leu His Gly
Gly Lys Gly Asp Asp 65 70 75 80 Ile Phe Val His Arg Lys Gly Asp Gly
Asn Asp Ile Ile Thr Asp Ser 85 90 95 Asp Gly Asn Asp Lys Leu Ser
Phe Ser Asp Ser Asn Leu Lys Asp Leu 100 105 110 Thr Phe Glu Lys Val
Lys His Asn Leu Val Ile Thr Asn Ser Lys Lys 115 120 125 Glu Lys Val
Thr Ile Gln Asn Trp Phe Arg Glu Ala Asp Phe Ala Lys 130 135 140 Glu
Val Pro Asn Tyr Lys Ala Thr Lys Asp Glu Lys Ile Glu Glu Ile 145 150
155 160 Ile Gly Gln Asn Gly Glu Arg Ile Thr Ser Lys Gln Val Asp Asp
Leu 165 170 175 Ile Ala Lys Gly Asn Gly Lys Ile Thr Gln Asp Glu Leu
Ser Lys Val 180 185 190 Val Asp Asn Tyr Glu Leu Leu Lys His Ser Lys
Asn Val Thr Asn Ser 195 200 205 Leu Asp Lys Leu Ile Ser Ser Val Ser
Ala Phe Thr Ser Ser Asn Asp 210 215 220 Ser Arg Asn Val Leu Val Ala
Pro Thr Ser Met Leu Asp Gln Ser Leu 225 230 235 240 Ser Ser Leu Gln
Phe Ala Arg Ala Ala 245 19372DNAArtificial Sequenced153_00985
d153_00985 Leukotoxin-activating lysine-acyltransferase lktC
serotype A1 (EC 2.3.1.) (Toxin-activating protein C) (Leukotoxin C)
- LKTC 19atgctactta tagataacgg tattccgatc gcttattgta gttgggcaga
tttaaacctt 60gagactgagg tgaaatatat taaggatatt aattcgttaa caccagaaga
atggcagtct 120ggtgacagac gctggattat tgattgggta gcaccattcg
gacattctca attactttat 180aaaaaaatgt
gtcagaaata ccctgatatg atcgtcagat ctatacgctt ttatccaaag
240cagaaagaat taggcaaaat tgcctacttt aaaggaggta aattagataa
aaaaacagca 300aaaaaacgtt ttgatacata tcaagaagag ctggcaacag
cacttaaaaa tgaatttaat 360tttattaaaa aa 37220124PRTArtificial
SequenceLeukotoxin C 20Met Leu Leu Ile Asp Asn Gly Ile Pro Ile Ala
Tyr Cys Ser Trp Ala 1 5 10 15 Asp Leu Asn Leu Glu Thr Glu Val Lys
Tyr Ile Lys Asp Ile Asn Ser 20 25 30 Leu Thr Pro Glu Glu Trp Gln
Ser Gly Asp Arg Arg Trp Ile Ile Asp 35 40 45 Trp Val Ala Pro Phe
Gly His Ser Gln Leu Leu Tyr Lys Lys Met Cys 50 55 60 Gln Lys Tyr
Pro Asp Met Ile Val Arg Ser Ile Arg Phe Tyr Pro Lys 65 70 75 80 Gln
Lys Glu Leu Gly Lys Ile Ala Tyr Phe Lys Gly Gly Lys Leu Asp 85 90
95 Lys Lys Thr Ala Lys Lys Arg Phe Asp Thr Tyr Gln Glu Glu Leu Ala
100 105 110 Thr Ala Leu Lys Asn Glu Phe Asn Phe Ile Lys Lys 115 120
212859DNAArtificial Sequenced153_00984 d153_00984 bifunctional
hemolysin- adenylate cyclase precursor - LKTA 21atgggaacta
gacttacaac cctatcaaat gggctaaaaa acactttaac ggcaaccaaa 60agtggcttac
ataaagccgg tcaatcatta acccaagccg gcagttcttt aaaaactggg
120gcaaaaaaaa ttatcctcta tattccccaa aattaccaat atgatactga
acaaggtaat 180ggtttacagg atttagtcaa agcggccgaa gagttgggga
ttgaggtaca aagagaagaa 240cgcaataata ttgcaacagc tcaaaccagt
ttaggcacga ttcaaaccgc tattggctta 300actgagcgtg gcattgtgtt
atccgctcca caaattgata aattgctaca gaaaactaaa 360gcaggccaag
cattaggttc tgccgaaagc attgtacaaa atgcaaataa agccaaaact
420gtattatctg gcattcaatc tattttaggc tcagtattgg ctggaatgga
tttagatgag 480gccttacaga ataacagcaa ccaacatgct cttgctaaag
ctggcttgga gctaacaaat 540tcattaattg aaaatattgc taattcagta
aaaacacttg acgaatttgg tgagcaaatt 600agtcaatttg gttcaaaact
acaaaatatc aaaggcttag ggactttagg agacaaactc 660aaaaatatcg
gtggacttga taaagctggc cttggtttag atgttatctc agggctatta
720tcgggcgcaa cagctgcact tgtacttgca gataaaaatg cttcaacagc
taaaaaagtg 780ggtgcgggtt ttgaattggc aaaccaagtt gttggtaata
ttaccaaagc cgtttcttct 840tacattttag cccaacgtgt tgcagcaggt
ttatcttcaa ctgggcctgt ggctgcttta 900attgcttcta ctgtttctct
tgcgattagc ccattagcat ttgccggtat tgccgataaa 960tttaatcatg
caaaaagttt agagagttat gccgaacgct ttaaaaaatt aggctatgac
1020ggagataatt tattagcaga atatcagcgg ggaacaggga ctattgatgc
atcggttact 1080gcaattaata ccgcattggc cgctattgct ggtggtgtgt
ctgctgctgc agccggctcg 1140gttattgctt caccgattgc cttattagta
tctgggatta ccggtgtaat ttctacgatt 1200ctgcaatatt ctaaacaagc
aatgtttgag cacgttgcaa ataaaattca taacaaaatt 1260gtagaatggg
aaaaaaataa tcacggtaag aactactttg aaaatggtta cgatgcccgt
1320tatcttgcga atttacaaga taatatgaaa ttcttactga acttaaacaa
agagttacag 1380gcagaacgtg tcatcgctat tactcagcag caatgggata
acaacattgg tgatttagct 1440ggtattagcc gtttaggtga aaaagtcctt
agtggtaaag cctatgtgga tgcgtttgaa 1500gaaggcaaac acattaaagc
cgataaatta gtacagttgg attcggcaaa cggtattatt 1560gatgtgagta
attcgggtaa agcgaaaact cagcatatct tattcagaac gccattattg
1620acgccgggaa cagagcatcg tgaacgcgta caaacaggta aatatgaata
tattaccaag 1680ctcaatatta accgtgtaga tagctggaaa attacagatg
gtgcagcaag ttctaccttt 1740gatttaacta acgttgttca gcgtattggt
attgaattag acaatgctgg aaatgtaact 1800aaaaccaaag aaacaaaaat
tattgccaaa cttggtgaag gtgatgacaa cgtatttgtt 1860ggttctggta
cgacggaaat tgatggcggt gaaggttacg accgagttca ctatagccgt
1920ggaaactatg gtgctttaac tattgatgca accaaagaga ccgagcaagg
tagttatacc 1980gtaaatcgtt tcgtagaaac cggtaaagca ctacacgaag
tgacttcaac ccataccgca 2040ttagtgggca accgtgaaga aaaaatagaa
tatcgtcata gcaataacca gcaccatgcc 2100ggttattaca ccaaagatac
cttgaaagct gttgaagaaa ttatcggtac atcacataac 2160gatatcttta
aaggtagtaa gttcaatgat gcctttaacg gtggtgatgg tgtcgatact
2220attgacggta acgacggcaa tgaccgctta tttggtggta aaggcgatga
tattctcgat 2280ggtggaaatg gtgatgattt tatcgatggc ggtaaaggca
acgacctatt acacggtggc 2340aagggcgatg atattttcgt tcaccgtaaa
ggcgatggta atgatattat taccgattct 2400gacggcaatg ataaattatc
attctctgat tcgaacttaa aagatttaac atttgaaaaa 2460gttaaacata
atcttgtcat cacgaatagc aaaaaagaga aagtgaccat tcaaaactgg
2520ttccgagagg ctgattttgc taaagaagtg cctaattata aagcaactaa
agatgagaaa 2580atcgaagaaa tcatcggtca aaatggcgag cggatcacct
caaagcaagt tgatgatctt 2640atcgcaaaag gtaacggcaa aattacccaa
gatgagctat caaaagttgt tgataactat 2700gaattgctca aacatagcaa
aaatgtgaca aacagcttag ataagttaat ctcatctgta 2760agtgcattta
cctcgtctaa tgattcgaga aatgtattag tggctccaac ttcaatgttg
2820gatcaaagtt tatcttctct tcaatttgct agagcagct
285922953PRTArtificial SequenceLeukotoxin C 22Met Gly Thr Arg Leu
Thr Thr Leu Ser Asn Gly Leu Lys Asn Thr Leu 1 5 10 15 Thr Ala Thr
Lys Ser Gly Leu His Lys Ala Gly Gln Ser Leu Thr Gln 20 25 30 Ala
Gly Ser Ser Leu Lys Thr Gly Ala Lys Lys Ile Ile Leu Tyr Ile 35 40
45 Pro Gln Asn Tyr Gln Tyr Asp Thr Glu Gln Gly Asn Gly Leu Gln Asp
50 55 60 Leu Val Lys Ala Ala Glu Glu Leu Gly Ile Glu Val Gln Arg
Glu Glu 65 70 75 80 Arg Asn Asn Ile Ala Thr Ala Gln Thr Ser Leu Gly
Thr Ile Gln Thr 85 90 95 Ala Ile Gly Leu Thr Glu Arg Gly Ile Val
Leu Ser Ala Pro Gln Ile 100 105 110 Asp Lys Leu Leu Gln Lys Thr Lys
Ala Gly Gln Ala Leu Gly Ser Ala 115 120 125 Glu Ser Ile Val Gln Asn
Ala Asn Lys Ala Lys Thr Val Leu Ser Gly 130 135 140 Ile Gln Ser Ile
Leu Gly Ser Val Leu Ala Gly Met Asp Leu Asp Glu 145 150 155 160 Ala
Leu Gln Asn Asn Ser Asn Gln His Ala Leu Ala Lys Ala Gly Leu 165 170
175 Glu Leu Thr Asn Ser Leu Ile Glu Asn Ile Ala Asn Ser Val Lys Thr
180 185 190 Leu Asp Glu Phe Gly Glu Gln Ile Ser Gln Phe Gly Ser Lys
Leu Gln 195 200 205 Asn Ile Lys Gly Leu Gly Thr Leu Gly Asp Lys Leu
Lys Asn Ile Gly 210 215 220 Gly Leu Asp Lys Ala Gly Leu Gly Leu Asp
Val Ile Ser Gly Leu Leu 225 230 235 240 Ser Gly Ala Thr Ala Ala Leu
Val Leu Ala Asp Lys Asn Ala Ser Thr 245 250 255 Ala Lys Lys Val Gly
Ala Gly Phe Glu Leu Ala Asn Gln Val Val Gly 260 265 270 Asn Ile Thr
Lys Ala Val Ser Ser Tyr Ile Leu Ala Gln Arg Val Ala 275 280 285 Ala
Gly Leu Ser Ser Thr Gly Pro Val Ala Ala Leu Ile Ala Ser Thr 290 295
300 Val Ser Leu Ala Ile Ser Pro Leu Ala Phe Ala Gly Ile Ala Asp Lys
305 310 315 320 Phe Asn His Ala Lys Ser Leu Glu Ser Tyr Ala Glu Arg
Phe Lys Lys 325 330 335 Leu Gly Tyr Asp Gly Asp Asn Leu Leu Ala Glu
Tyr Gln Arg Gly Thr 340 345 350 Gly Thr Ile Asp Ala Ser Val Thr Ala
Ile Asn Thr Ala Leu Ala Ala 355 360 365 Ile Ala Gly Gly Val Ser Ala
Ala Ala Ala Gly Ser Val Ile Ala Ser 370 375 380 Pro Ile Ala Leu Leu
Val Ser Gly Ile Thr Gly Val Ile Ser Thr Ile 385 390 395 400 Leu Gln
Tyr Ser Lys Gln Ala Met Phe Glu His Val Ala Asn Lys Ile 405 410 415
His Asn Lys Ile Val Glu Trp Glu Lys Asn Asn His Gly Lys Asn Tyr 420
425 430 Phe Glu Asn Gly Tyr Asp Ala Arg Tyr Leu Ala Asn Leu Gln Asp
Asn 435 440 445 Met Lys Phe Leu Leu Asn Leu Asn Lys Glu Leu Gln Ala
Glu Arg Val 450 455 460 Ile Ala Ile Thr Gln Gln Gln Trp Asp Asn Asn
Ile Gly Asp Leu Ala 465 470 475 480 Gly Ile Ser Arg Leu Gly Glu Lys
Val Leu Ser Gly Lys Ala Tyr Val 485 490 495 Asp Ala Phe Glu Glu Gly
Lys His Ile Lys Ala Asp Lys Leu Val Gln 500 505 510 Leu Asp Ser Ala
Asn Gly Ile Ile Asp Val Ser Asn Ser Gly Lys Ala 515 520 525 Lys Thr
Gln His Ile Leu Phe Arg Thr Pro Leu Leu Thr Pro Gly Thr 530 535 540
Glu His Arg Glu Arg Val Gln Thr Gly Lys Tyr Glu Tyr Ile Thr Lys 545
550 555 560 Leu Asn Ile Asn Arg Val Asp Ser Trp Lys Ile Thr Asp Gly
Ala Ala 565 570 575 Ser Ser Thr Phe Asp Leu Thr Asn Val Val Gln Arg
Ile Gly Ile Glu 580 585 590 Leu Asp Asn Ala Gly Asn Val Thr Lys Thr
Lys Glu Thr Lys Ile Ile 595 600 605 Ala Lys Leu Gly Glu Gly Asp Asp
Asn Val Phe Val Gly Ser Gly Thr 610 615 620 Thr Glu Ile Asp Gly Gly
Glu Gly Tyr Asp Arg Val His Tyr Ser Arg 625 630 635 640 Gly Asn Tyr
Gly Ala Leu Thr Ile Asp Ala Thr Lys Glu Thr Glu Gln 645 650 655 Gly
Ser Tyr Thr Val Asn Arg Phe Val Glu Thr Gly Lys Ala Leu His 660 665
670 Glu Val Thr Ser Thr His Thr Ala Leu Val Gly Asn Arg Glu Glu Lys
675 680 685 Ile Glu Tyr Arg His Ser Asn Asn Gln His His Ala Gly Tyr
Tyr Thr 690 695 700 Lys Asp Thr Leu Lys Ala Val Glu Glu Ile Ile Gly
Thr Ser His Asn 705 710 715 720 Asp Ile Phe Lys Gly Ser Lys Phe Asn
Asp Ala Phe Asn Gly Gly Asp 725 730 735 Gly Val Asp Thr Ile Asp Gly
Asn Asp Gly Asn Asp Arg Leu Phe Gly 740 745 750 Gly Lys Gly Asp Asp
Ile Leu Asp Gly Gly Asn Gly Asp Asp Phe Ile 755 760 765 Asp Gly Gly
Lys Gly Asn Asp Leu Leu His Gly Gly Lys Gly Asp Asp 770 775 780 Ile
Phe Val His Arg Lys Gly Asp Gly Asn Asp Ile Ile Thr Asp Ser 785 790
795 800 Asp Gly Asn Asp Lys Leu Ser Phe Ser Asp Ser Asn Leu Lys Asp
Leu 805 810 815 Thr Phe Glu Lys Val Lys His Asn Leu Val Ile Thr Asn
Ser Lys Lys 820 825 830 Glu Lys Val Thr Ile Gln Asn Trp Phe Arg Glu
Ala Asp Phe Ala Lys 835 840 845 Glu Val Pro Asn Tyr Lys Ala Thr Lys
Asp Glu Lys Ile Glu Glu Ile 850 855 860 Ile Gly Gln Asn Gly Glu Arg
Ile Thr Ser Lys Gln Val Asp Asp Leu 865 870 875 880 Ile Ala Lys Gly
Asn Gly Lys Ile Thr Gln Asp Glu Leu Ser Lys Val 885 890 895 Val Asp
Asn Tyr Glu Leu Leu Lys His Ser Lys Asn Val Thr Asn Ser 900 905 910
Leu Asp Lys Leu Ile Ser Ser Val Ser Ala Phe Thr Ser Ser Asn Asp 915
920 925 Ser Arg Asn Val Leu Val Ala Pro Thr Ser Met Leu Asp Gln Ser
Leu 930 935 940 Ser Ser Leu Gln Phe Ala Arg Ala Ala 945 950
232124DNAArtificial Sequenced153_00983 d153_00983 ABC-type
bacteriocin/ lantibiotic exporters, contain an N-terminal
double-glycine peptidase domain - LKTB 23atggaagcta accatcaaag
gaatgatctt ggtttagttg ccctcactat gttggcacaa 60taccataata tttcgcttaa
tccggaagaa ataaaacata aatttgatct tgacggaaaa 120gggctttctt
taactgcttg gcttttagct gcaaaatcgt tagcgttgaa agcgaaacac
180attaaaaaag agatttcccg cttacacttg gtgaatttac cggcattagt
ttggcaagat 240aacggtaaac attttttatt ggtaaaagtg gataccgata
ataaccgcta tttaacttac 300aatttggaac aagatgctcc acaaattctg
tcacaagacg aatttgaagc ctgctatcaa 360gggcagttaa ttttggtcac
gtccagagct tccgtagtag gtcaattagc aaagttcgat 420ttcacctggt
ttattccggc ggtgatcaaa taccgaaaaa tctttctaga aaccttgatt
480gtttcgatct ttttgcaaat ttttgcccta attacaccgc tattcttcca
agttgttatg 540gataaagtac tggtgcatcg aggtttttca accttgaata
tcattacggt tgccttagct 600attgtgatca tctttgaaat tgtactaagt
ggtttgagaa cctatgtttt ttctcatagc 660actagccgta ttgatgttga
attaggcgct aaattatttc gacatttatt atcactaccc 720atttcttatt
ttgaaaacag acgagttgga gatacagtcg ctagggttag agaattagat
780caaattcgta atttccttac cggacaagca ttaacctcgg tgttagatct
cttattctct 840tttatctttt ttgccgtaat gtggtattac agcccaaaat
taaccttggt aattcttggt 900tcattgccct gctatatttt atggtcaatt
tttattagtc cgattttaag acggcgttta 960gatgagaaat ttgcccgaag
tgctgataac caagcattct tagttgagtc ggtaacagcc 1020atcaatatga
ttaaagcgat ggcggttgct ccacaaatga cggatacatg ggataaacag
1080ctggcaagct atgtttcatc aagtttccgt gtcaccgtat tagcaaccat
tgggcaacaa 1140ggtgtacaac ttattcaaaa aaccgttatg gtgattaacc
tttggttagg ggcacactta 1200gttatttcag gcgatctgag tattgggcaa
ttaattgcct ttaatatgct atcagggcaa 1260gtgattgcac cggtgattcg
gctggctcag ctctggcaag atttccaaca agttgggatt 1320tccgtcactc
gcttaggtga tgttttaaac tctccaaccg aacaatatca aggcaaatta
1380tcactaccag aaataaaagg cgatatctca tttaaaaata tccgctttag
atataaacca 1440gatgcaccaa ctattttaaa taatgtgaat ttagaaatta
ggcaaggaga agtgattggg 1500attgttggac gttccggttc aggcaaaagt
actctgacta aattactgca acgtttttat 1560attcctgaaa atgggcaggt
tttgattgat ggacatgatc tagccttagc tgatccaaac 1620tggctacgcc
gtcaaatagg tgtagtgctg caagataatg tgttattaaa ccgcagtatc
1680cgagaaaata ttgcgctatc agatccagga atgccaatgg agcgagtaat
ttatgcagca 1740aaattagcag gggctcacga ttttatttca gaattgcgtg
aaggttataa caccattgtg 1800ggtgaacaag gagcggggct ttcaggcggg
caacgccaac ggattgcgat tgctcgagct 1860ttggtaaaca acccgaaaat
cctgattttt gatgaggcaa ccagtgccct cgattacgaa 1920tctgagcata
ttattatgca aaatatgcaa aaaatatgcc aaggcagaac cgtgattttg
1980attgcacatc gtttatcgac cgtcaaaaat gcggatcgaa ttattgtgat
ggaaaagggg 2040gaaattgttg agcaaggcaa gcaccacgaa ttactgcaaa
acagtaacgg actttattcc 2100tacttacacc aattacaact taat
212424708PRTArtificial SequenceLeukotoxin B 24Met Glu Ala Asn His
Gln Arg Asn Asp Leu Gly Leu Val Ala Leu Thr 1 5 10 15 Met Leu Ala
Gln Tyr His Asn Ile Ser Leu Asn Pro Glu Glu Ile Lys 20 25 30 His
Lys Phe Asp Leu Asp Gly Lys Gly Leu Ser Leu Thr Ala Trp Leu 35 40
45 Leu Ala Ala Lys Ser Leu Ala Leu Lys Ala Lys His Ile Lys Lys Glu
50 55 60 Ile Ser Arg Leu His Leu Val Asn Leu Pro Ala Leu Val Trp
Gln Asp 65 70 75 80 Asn Gly Lys His Phe Leu Leu Val Lys Val Asp Thr
Asp Asn Asn Arg 85 90 95 Tyr Leu Thr Tyr Asn Leu Glu Gln Asp Ala
Pro Gln Ile Leu Ser Gln 100 105 110 Asp Glu Phe Glu Ala Cys Tyr Gln
Gly Gln Leu Ile Leu Val Thr Ser 115 120 125 Arg Ala Ser Val Val Gly
Gln Leu Ala Lys Phe Asp Phe Thr Trp Phe 130 135 140 Ile Pro Ala Val
Ile Lys Tyr Arg Lys Ile Phe Leu Glu Thr Leu Ile 145 150 155 160 Val
Ser Ile Phe Leu Gln Ile Phe Ala Leu Ile Thr Pro Leu Phe Phe 165 170
175 Gln Val Val Met Asp Lys Val Leu Val His Arg Gly Phe Ser Thr Leu
180 185 190 Asn Ile Ile Thr Val Ala Leu Ala Ile Val Ile Ile Phe Glu
Ile Val 195 200 205 Leu Ser Gly Leu Arg Thr Tyr Val Phe Ser His Ser
Thr Ser Arg Ile 210 215 220 Asp Val Glu Leu Gly Ala Lys Leu Phe Arg
His Leu Leu Ser Leu Pro 225 230 235 240 Ile Ser Tyr Phe Glu Asn Arg
Arg Val Gly Asp Thr Val Ala Arg Val 245 250 255 Arg Glu Leu Asp Gln
Ile Arg Asn Phe Leu Thr Gly Gln Ala Leu Thr 260 265 270 Ser Val Leu
Asp Leu Leu Phe Ser Phe Ile Phe Phe Ala Val Met Trp 275 280 285 Tyr
Tyr Ser Pro Lys Leu Thr Leu Val Ile Leu Gly Ser Leu Pro Cys 290 295
300 Tyr Ile Leu Trp Ser Ile Phe Ile Ser Pro Ile Leu Arg Arg Arg Leu
305 310 315 320 Asp Glu Lys Phe Ala Arg Ser Ala Asp Asn Gln Ala Phe
Leu Val Glu 325 330 335 Ser Val Thr Ala Ile Asn Met Ile Lys Ala Met
Ala Val Ala Pro Gln 340 345 350 Met Thr Asp Thr Trp Asp Lys Gln Leu
Ala Ser Tyr Val Ser Ser Ser 355 360
365 Phe Arg Val Thr Val Leu Ala Thr Ile Gly Gln Gln Gly Val Gln Leu
370 375 380 Ile Gln Lys Thr Val Met Val Ile Asn Leu Trp Leu Gly Ala
His Leu 385 390 395 400 Val Ile Ser Gly Asp Leu Ser Ile Gly Gln Leu
Ile Ala Phe Asn Met 405 410 415 Leu Ser Gly Gln Val Ile Ala Pro Val
Ile Arg Leu Ala Gln Leu Trp 420 425 430 Gln Asp Phe Gln Gln Val Gly
Ile Ser Val Thr Arg Leu Gly Asp Val 435 440 445 Leu Asn Ser Pro Thr
Glu Gln Tyr Gln Gly Lys Leu Ser Leu Pro Glu 450 455 460 Ile Lys Gly
Asp Ile Ser Phe Lys Asn Ile Arg Phe Arg Tyr Lys Pro 465 470 475 480
Asp Ala Pro Thr Ile Leu Asn Asn Val Asn Leu Glu Ile Arg Gln Gly 485
490 495 Glu Val Ile Gly Ile Val Gly Arg Ser Gly Ser Gly Lys Ser Thr
Leu 500 505 510 Thr Lys Leu Leu Gln Arg Phe Tyr Ile Pro Glu Asn Gly
Gln Val Leu 515 520 525 Ile Asp Gly His Asp Leu Ala Leu Ala Asp Pro
Asn Trp Leu Arg Arg 530 535 540 Gln Ile Gly Val Val Leu Gln Asp Asn
Val Leu Leu Asn Arg Ser Ile 545 550 555 560 Arg Glu Asn Ile Ala Leu
Ser Asp Pro Gly Met Pro Met Glu Arg Val 565 570 575 Ile Tyr Ala Ala
Lys Leu Ala Gly Ala His Asp Phe Ile Ser Glu Leu 580 585 590 Arg Glu
Gly Tyr Asn Thr Ile Val Gly Glu Gln Gly Ala Gly Leu Ser 595 600 605
Gly Gly Gln Arg Gln Arg Ile Ala Ile Ala Arg Ala Leu Val Asn Asn 610
615 620 Pro Lys Ile Leu Ile Phe Asp Glu Ala Thr Ser Ala Leu Asp Tyr
Glu 625 630 635 640 Ser Glu His Ile Ile Met Gln Asn Met Gln Lys Ile
Cys Gln Gly Arg 645 650 655 Thr Val Ile Leu Ile Ala His Arg Leu Ser
Thr Val Lys Asn Ala Asp 660 665 670 Arg Ile Ile Val Met Glu Lys Gly
Glu Ile Val Glu Gln Gly Lys His 675 680 685 His Glu Leu Leu Gln Asn
Ser Asn Gly Leu Tyr Ser Tyr Leu His Gln 690 695 700 Leu Gln Leu Asn
705 251434DNAArtificial Sequenced153_00982 d153_00982 Microcin H47
secretion protein - LKTD 25atgaaaatat ggcttagtgg tatttatgaa
tttttcctac gctataaaaa catttgggca 60gaagtatgga aaattcgtaa agaattagac
cacccaaaca gaaaaaaaga cgaaagtgaa 120tttttaccgg cacatttaga
actgattgaa accccggttt ctaaaaaacc acgtctaatt 180gcttatttga
ttatgctatt tttagttgtg gcaattgtgc ttgccagtgt aagcaaagtt
240gaaattgtgg cgactgctcc cggtaaatta acttttagtg gcagaagtaa
agaaattaaa 300ccgattgaaa acgccattgt acaagaaatt ttcgttaaag
atgggcagtt tgtggaaaaa 360gggcaattat tagtcagctt aactgcattg
ggttctgatg cagatatcaa aaagaccatg 420gcttcacttt ctttagctaa
actggagaac tatcgctacc aaactttgct tactgccatt 480gaaaaagagt
ccttgccggt gattgattta tctagaaccg aatttaaaga ttcatcggaa
540gaagatcgac tacgtattaa acacttaatt gaggagcaat acaccacttg
gcaaaaacaa 600aaaacacaga aaactttagc gtataagcgt aaagaggctg
aaaaacaaac aatatttgcc 660tatgtccgta aatatgaagg tgcaacacgt
attgaacaag aaaaattaaa agactttaag 720gcactttata aacagaagtc
tttatctaag cacgaacttc ttgcgcaaga aaataaatta 780attgaggctc
agaatgagct agctgtttat cgctcaaaat taaatgaatt agaaaatgat
840ctactcaatg taaaagaaga acttgaattg atcacgcaat tctttaaaag
cgatgtgttg 900gaaaaattaa agcaacatat tgaaaatgaa cgccaacttc
ggctcgagtt agaaaaaaat 960aatcaacgca gacaggcctc gatgatcaga
gcaccggttt ccggtacggt tcagcaactg 1020aaaattcaca ctataggtgg
tgttgttacg actgctgaaa ccttgatgat cattgtgccg 1080gaagacgatg
tgttagaggc caccgctctg gttccaaaca aagatatcgg ctttgttgca
1140gcagggcagg aggtgattat taaagtggaa actttccctt atacacgcta
tggttatcta 1200actggtcgaa ttaaacatat tagcccggat gcgattgaac
aacctaatgt aggcttagtt 1260tttaatgcaa ctatagctat agataggaag
aatctaacat cgcctgatgg gcgaaaaatt 1320gatttgagtt caggtatgac
aataactgct gaaatcaaaa ccggtgaacg gagtgtaatg 1380agttatttac
tcagcccatt agaagaatct gtcacagaaa gtttaaggga acgc
143426478PRTArtificial SequenceLeukotoxin D 26Met Lys Ile Trp Leu
Ser Gly Ile Tyr Glu Phe Phe Leu Arg Tyr Lys 1 5 10 15 Asn Ile Trp
Ala Glu Val Trp Lys Ile Arg Lys Glu Leu Asp His Pro 20 25 30 Asn
Arg Lys Lys Asp Glu Ser Glu Phe Leu Pro Ala His Leu Glu Leu 35 40
45 Ile Glu Thr Pro Val Ser Lys Lys Pro Arg Leu Ile Ala Tyr Leu Ile
50 55 60 Met Leu Phe Leu Val Val Ala Ile Val Leu Ala Ser Val Ser
Lys Val 65 70 75 80 Glu Ile Val Ala Thr Ala Pro Gly Lys Leu Thr Phe
Ser Gly Arg Ser 85 90 95 Lys Glu Ile Lys Pro Ile Glu Asn Ala Ile
Val Gln Glu Ile Phe Val 100 105 110 Lys Asp Gly Gln Phe Val Glu Lys
Gly Gln Leu Leu Val Ser Leu Thr 115 120 125 Ala Leu Gly Ser Asp Ala
Asp Ile Lys Lys Thr Met Ala Ser Leu Ser 130 135 140 Leu Ala Lys Leu
Glu Asn Tyr Arg Tyr Gln Thr Leu Leu Thr Ala Ile 145 150 155 160 Glu
Lys Glu Ser Leu Pro Val Ile Asp Leu Ser Arg Thr Glu Phe Lys 165 170
175 Asp Ser Ser Glu Glu Asp Arg Leu Arg Ile Lys His Leu Ile Glu Glu
180 185 190 Gln Tyr Thr Thr Trp Gln Lys Gln Lys Thr Gln Lys Thr Leu
Ala Tyr 195 200 205 Lys Arg Lys Glu Ala Glu Lys Gln Thr Ile Phe Ala
Tyr Val Arg Lys 210 215 220 Tyr Glu Gly Ala Thr Arg Ile Glu Gln Glu
Lys Leu Lys Asp Phe Lys 225 230 235 240 Ala Leu Tyr Lys Gln Lys Ser
Leu Ser Lys His Glu Leu Leu Ala Gln 245 250 255 Glu Asn Lys Leu Ile
Glu Ala Gln Asn Glu Leu Ala Val Tyr Arg Ser 260 265 270 Lys Leu Asn
Glu Leu Glu Asn Asp Leu Leu Asn Val Lys Glu Glu Leu 275 280 285 Glu
Leu Ile Thr Gln Phe Phe Lys Ser Asp Val Leu Glu Lys Leu Lys 290 295
300 Gln His Ile Glu Asn Glu Arg Gln Leu Arg Leu Glu Leu Glu Lys Asn
305 310 315 320 Asn Gln Arg Arg Gln Ala Ser Met Ile Arg Ala Pro Val
Ser Gly Thr 325 330 335 Val Gln Gln Leu Lys Ile His Thr Ile Gly Gly
Val Val Thr Thr Ala 340 345 350 Glu Thr Leu Met Ile Ile Val Pro Glu
Asp Asp Val Leu Glu Ala Thr 355 360 365 Ala Leu Val Pro Asn Lys Asp
Ile Gly Phe Val Ala Ala Gly Gln Glu 370 375 380 Val Ile Ile Lys Val
Glu Thr Phe Pro Tyr Thr Arg Tyr Gly Tyr Leu 385 390 395 400 Thr Gly
Arg Ile Lys His Ile Ser Pro Asp Ala Ile Glu Gln Pro Asn 405 410 415
Val Gly Leu Val Phe Asn Ala Thr Ile Ala Ile Asp Arg Lys Asn Leu 420
425 430 Thr Ser Pro Asp Gly Arg Lys Ile Asp Leu Ser Ser Gly Met Thr
Ile 435 440 445 Thr Ala Glu Ile Lys Thr Gly Glu Arg Ser Val Met Ser
Tyr Leu Leu 450 455 460 Ser Pro Leu Glu Glu Ser Val Thr Glu Ser Leu
Arg Glu Arg 465 470 475
D00001

D00002

D00003

D00004

D00005

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.